Polynucleotide molecules and uses thereof
Fraley , et al. A
U.S. patent number 10,385,088 [Application Number 15/026,836] was granted by the patent office on 2019-08-20 for polynucleotide molecules and uses thereof. This patent grant is currently assigned to ModernaTX, Inc.. The grantee listed for this patent is ModernaTX, Inc.. Invention is credited to Andrew W. Fraley, Atanu Roy, Matthew Stanton.





View All Diagrams
| United States Patent | 10,385,088 |
| Fraley , et al. | August 20, 2019 |
Polynucleotide molecules and uses thereof
Abstract
The present disclosure provides alternative sugar moieties and polynucleotides comprising such sugar moieties, and methods of use thereof.
| Inventors: | Fraley; Andrew W. (Arlington, MA), Roy; Atanu (Stoneham, MA), Stanton; Matthew (Marlton, NJ) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Applicant: |
|
||||||||||
| Assignee: | ModernaTX, Inc. (Cambridge,
MA) |
||||||||||
| Family ID: | 52779294 | ||||||||||
| Appl. No.: | 15/026,836 | ||||||||||
| Filed: | October 2, 2014 | ||||||||||
| PCT Filed: | October 02, 2014 | ||||||||||
| PCT No.: | PCT/US2014/058891 | ||||||||||
| 371(c)(1),(2),(4) Date: | April 01, 2016 | ||||||||||
| PCT Pub. No.: | WO2015/051169 | ||||||||||
| PCT Pub. Date: | April 09, 2015 |
Prior Publication Data
| Document Identifier | Publication Date | |
|---|---|---|
| US 20160237108 A1 | Aug 18, 2016 | |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | Issue Date | ||
|---|---|---|---|---|---|
| 61885979 | Oct 2, 2013 | ||||
| 61896478 | Oct 28, 2013 | ||||
| 61915907 | Dec 13, 2013 | ||||
| 62036944 | Aug 13, 2014 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07H 19/167 (20130101); C07K 14/525 (20130101); C07H 19/20 (20130101); C07H 21/02 (20130101); C07K 14/565 (20130101); C07H 19/067 (20130101); C07H 19/10 (20130101); C07K 14/535 (20130101) |
| Current International Class: | C07H 19/10 (20060101); C07H 19/20 (20060101); C07K 14/525 (20060101); C07K 14/535 (20060101); C07H 21/02 (20060101); C07H 19/167 (20060101); C07H 19/067 (20060101); C07K 14/565 (20060101) |
References Cited [Referenced By]
U.S. Patent Documents
| 5034506 | July 1991 | Summerton et al. |
| 5426180 | June 1995 | Kool |
| 5489677 | February 1996 | Sanghvi et al. |
| 5512439 | April 1996 | Hornes et al. |
| 5591722 | January 1997 | Montgomery et al. |
| 5637459 | June 1997 | Burke et al. |
| 5639873 | June 1997 | Barascut et al. |
| 5789578 | August 1998 | Burton et al. |
| 5808039 | September 1998 | Reddy et al. |
| 5989911 | November 1999 | Fournier et al. |
| 6022715 | February 2000 | Merenkova et al. |
| 6248268 | June 2001 | Cook |
| 6303378 | October 2001 | Bridenbaugh et al. |
| 6423492 | July 2002 | Harbron |
| 6511832 | January 2003 | Guarino et al. |
| 6521411 | February 2003 | Hecker et al. |
| 7691569 | April 2010 | Wohlgemuth et al. |
| 8093367 | January 2012 | Kore et al. |
| 8664194 | March 2014 | de Fougerolles et al. |
| 8680069 | March 2014 | de Fougerolles et al. |
| 8691750 | April 2014 | Constien et al. |
| 8710200 | April 2014 | Schrum et al. |
| 8716465 | May 2014 | Rossi et al. |
| 8802438 | August 2014 | Rossi et al. |
| 8822663 | September 2014 | Schrum et al. |
| 8883506 | November 2014 | Rossi et al. |
| 8898864 | December 2014 | Porter |
| 8969353 | March 2015 | Mahon et al. |
| 8980864 | March 2015 | Hoge et al. |
| 8999380 | April 2015 | Bancel et al. |
| 9050297 | June 2015 | Chakraborty et al. |
| 9061059 | June 2015 | Chakraborty et al. |
| 9089604 | July 2015 | Chakraborty et al. |
| 9095552 | August 2015 | Chakraborty et al. |
| 9107886 | August 2015 | Chakraborty et al. |
| 9114113 | August 2015 | Chakraborty et al. |
| 9149506 | October 2015 | Chakraborty et al. |
| 9428535 | August 2016 | de Fougerolles et al. |
| 9533047 | January 2017 | de Fougerolles et al. |
| 9675668 | June 2017 | Bancel et al. |
| 9751925 | September 2017 | Hoge et al. |
| 9803177 | October 2017 | Rossi et al. |
| 9872900 | January 2018 | Ciaramella |
| 10022435 | July 2018 | Ciaramella |
| 10064935 | September 2018 | Ciaramella |
| 10072057 | September 2018 | Hoge et al. |
| 2001/0025097 | September 2001 | Sheridan et al. |
| 2002/0001812 | January 2002 | Smith et al. |
| 2002/0016450 | February 2002 | Laugharn et al. |
| 2002/0114784 | August 2002 | Li et al. |
| 2002/0130430 | September 2002 | Castor |
| 2002/0153312 | October 2002 | Gjerde et al. |
| 2002/0164635 | November 2002 | Salerno |
| 2003/0120035 | June 2003 | Gao et al. |
| 2003/0170876 | September 2003 | Widner et al. |
| 2003/0170891 | September 2003 | McSwiggen |
| 2003/0180754 | September 2003 | Bergholtz et al. |
| 2003/0180779 | September 2003 | Lofton-Day et al. |
| 2004/0038278 | February 2004 | Tzertzinis et al. |
| 2004/0142433 | July 2004 | Padgett et al. |
| 2004/0220127 | November 2004 | Sternberg et al. |
| 2004/0259097 | December 2004 | De Backer et al. |
| 2005/0003496 | January 2005 | McGall et al. |
| 2005/0130196 | June 2005 | Hofstadler et al. |
| 2006/0058266 | March 2006 | Manoharan et al. |
| 2006/0121441 | June 2006 | Spira |
| 2006/0223081 | October 2006 | Jarrell et al. |
| 2007/0020678 | January 2007 | Ault-Riche et al. |
| 2007/0037148 | February 2007 | Fong et al. |
| 2007/0037770 | February 2007 | Gryaznov et al. |
| 2007/0244062 | October 2007 | Laux et al. |
| 2007/0281336 | December 2007 | Jendrisak et al. |
| 2008/0076910 | March 2008 | Takkellapati et al. |
| 2008/0171711 | July 2008 | Hoerr et al. |
| 2008/0274463 | November 2008 | Chen et al. |
| 2008/0311140 | December 2008 | Lee et al. |
| 2009/0215125 | August 2009 | Reed et al. |
| 2009/0264511 | October 2009 | de Fougerolles et al. |
| 2009/0286852 | November 2009 | Kariko et al. |
| 2010/0015232 | January 2010 | Besenbacher et al. |
| 2010/0047261 | February 2010 | Hoerr et al. |
| 2010/0255574 | October 2010 | Rosen et al. |
| 2010/0261228 | October 2010 | Gharib et al. |
| 2011/0130440 | June 2011 | Manoharan et al. |
| 2011/0218170 | September 2011 | Thottassery et al. |
| 2011/0244026 | October 2011 | Guild et al. |
| 2011/0281938 | November 2011 | Schaub et al. |
| 2012/0046346 | February 2012 | Rossi et al. |
| 2012/0251618 | October 2012 | Schrum et al. |
| 2013/0046083 | February 2013 | Brown et al. |
| 2013/0046084 | February 2013 | Brown et al. |
| 2013/0115272 | May 2013 | de Fougerolles et al. |
| 2013/0115274 | May 2013 | Knopov et al. |
| 2013/0123481 | May 2013 | de Fougerolles et al. |
| 2013/0156849 | June 2013 | de Fougerolles et al. |
| 2013/0165504 | June 2013 | Bancel et al. |
| 2013/0203115 | August 2013 | Schrum et al. |
| 2013/0244282 | September 2013 | Schrum et al. |
| 2013/0245103 | September 2013 | de Fougerolles et al. |
| 2013/0245105 | September 2013 | de Fougerolles et al. |
| 2013/0245106 | September 2013 | de Fougerolles et al. |
| 2013/0251618 | September 2013 | Li et al. |
| 2013/0259923 | October 2013 | Bancel et al. |
| 2013/0259924 | October 2013 | Bancel et al. |
| 2014/0010861 | January 2014 | Bancel et al. |
| 2014/0105964 | April 2014 | Bancel et al. |
| 2014/0105966 | April 2014 | Bancel et al. |
| 2014/0147454 | May 2014 | Chakraborty et al. |
| 2014/0200261 | July 2014 | Hoge et al. |
| 2014/0206752 | July 2014 | Afeyan et al. |
| 2014/0206852 | July 2014 | Hoge et al. |
| 2014/0243399 | August 2014 | Schrum et al. |
| 2014/0273230 | September 2014 | Chen et al. |
| 2014/0275227 | September 2014 | Hoge et al. |
| 2014/0343129 | November 2014 | de Fougerolles et al. |
| 2014/0371302 | December 2014 | Afeyan et al. |
| 2015/0005372 | January 2015 | Hoge et al. |
| 2015/0017211 | January 2015 | de Fougerolles et al. |
| 2015/0030576 | January 2015 | Bancel |
| 2015/0050354 | February 2015 | Bouchon et al. |
| 2015/0050738 | February 2015 | Ozsolak et al. |
| 2015/0051268 | February 2015 | Bancel et al. |
| 2015/0056253 | February 2015 | Bancel et al. |
| 2015/0064235 | March 2015 | Bancel et al. |
| 2015/0064236 | March 2015 | Bancel et al. |
| 2015/0064725 | March 2015 | Schrum et al. |
| 2015/0086614 | March 2015 | Bancel et al. |
| 2015/0111248 | April 2015 | Bancel et al. |
| 2015/0141499 | May 2015 | Bancel et al. |
| 2015/0166616 | June 2015 | Bancel et al. |
| 2015/0167017 | June 2015 | Roy et al. |
| 2015/0211039 | July 2015 | Wang et al. |
| 2015/0291678 | October 2015 | Rudolph et al. |
| 2015/0307542 | October 2015 | Roy et al. |
| 2016/0017313 | January 2016 | Spivak et al. |
| 2016/0024140 | January 2016 | Issa et al. |
| 2016/0024141 | January 2016 | Issa et al. |
| 2016/0024492 | January 2016 | Issa et al. |
| 2016/0024547 | January 2016 | Bancel et al. |
| 2016/0032273 | February 2016 | Shahrokh et al. |
| 2016/0038612 | February 2016 | Hoge et al. |
| 2016/0177295 | June 2016 | Rudolph et al. |
| 2016/0194368 | July 2016 | Hoge et al. |
| 2016/0194625 | July 2016 | Hoge et al. |
| 2016/0237108 | August 2016 | Fraley et al. |
| 2028849 | Sep 1991 | CA | |||
| 2473135 | Jun 2003 | CA | |||
| 0366400 | May 1990 | EP | |||
| 1083232 | Feb 2005 | EP | |||
| 1619254 | Jan 2006 | EP | |||
| 1383556 | Mar 2008 | EP | |||
| 1831160 | Jun 2010 | EP | |||
| 2092064 | Sep 2010 | EP | |||
| 2377938 | Oct 2011 | EP | |||
| 2484770 | Aug 2012 | EP | |||
| 2188379 | Jan 2013 | EP | |||
| 2548960 | Jan 2013 | EP | |||
| 2011-130725 | Jul 2011 | JP | |||
| 2540017 | Jan 2015 | RU | |||
| WO-91/05058 | Apr 1991 | WO | |||
| WO-93/03052 | Feb 1993 | WO | |||
| WO-93/13121 | Jul 1993 | WO | |||
| WO-01/55306 | Aug 2001 | WO | |||
| WO-02/44399 | Jun 2002 | WO | |||
| WO-02/098443 | Dec 2002 | WO | |||
| WO-03/039523 | May 2003 | WO | |||
| WO-03/051881 | Jun 2003 | WO | |||
| WO-2004/020575 | Mar 2004 | WO | |||
| WO-2004/064782 | Aug 2004 | WO | |||
| WO-2006/015445 | Feb 2006 | WO | |||
| WO-2007/024708 | Mar 2007 | WO | |||
| WO-2007/024798 | Mar 2007 | WO | |||
| WO-2007/089607 | Aug 2007 | WO | |||
| WO-2007/120863 | Oct 2007 | WO | |||
| WO-2008/039669 | Apr 2008 | WO | |||
| WO-2008/045505 | Apr 2008 | WO | |||
| WO-2008/083949 | Jul 2008 | WO | |||
| WO-2008/120016 | Oct 2008 | WO | |||
| WO-2009/042971 | Apr 2009 | WO | |||
| WO-2009/051451 | Apr 2009 | WO | |||
| WO-2009/127060 | Oct 2009 | WO | |||
| WO-2009/127230 | Oct 2009 | WO | |||
| WO-2009/147519 | Dec 2009 | WO | |||
| WO-2009/149253 | Dec 2009 | WO | |||
| WO-2010/014895 | Feb 2010 | WO | |||
| WO-2010/017510 | Feb 2010 | WO | |||
| WO-2010/109289 | Sep 2010 | WO | |||
| WO-2010144740 | Dec 2010 | WO | |||
| WO-2011/005850 | Jan 2011 | WO | |||
| WO-2011/012316 | Feb 2011 | WO | |||
| WO-2011/068810 | Jun 2011 | WO | |||
| WO-2011/071931 | Jun 2011 | WO | |||
| WO-2011/127933 | Oct 2011 | WO | |||
| WO-2011/130624 | Oct 2011 | WO | |||
| WO-2011/133868 | Oct 2011 | WO | |||
| WO-2011/140627 | Nov 2011 | WO | |||
| WO-2012/019168 | Feb 2012 | WO | |||
| WO-2012/135805 | Oct 2012 | WO | |||
| WO-2012/138530 | Oct 2012 | WO | |||
| WO-2012/158736 | Nov 2012 | WO | |||
| WO-2012/164565 | Dec 2012 | WO | |||
| WO-2013/039857 | Mar 2013 | WO | |||
| WO-2013/039861 | Mar 2013 | WO | |||
| WO-2013/052523 | Apr 2013 | WO | |||
| WO-2013/090186 | Jun 2013 | WO | |||
| WO-2013/090294 | Jun 2013 | WO | |||
| WO-2013/090648 | Jun 2013 | WO | |||
| WO-2013/090897 | Jun 2013 | WO | |||
| WO-2013/096709 | Jun 2013 | WO | |||
| WO-2013/101690 | Jul 2013 | WO | |||
| WO-2013/113326 | Aug 2013 | WO | |||
| WO-2013/113501 | Aug 2013 | WO | |||
| WO-2013/113502 | Aug 2013 | WO | |||
| WO-2013/130161 | Sep 2013 | WO | |||
| WO-2013/151663 | Oct 2013 | WO | |||
| WO-2013/151664 | Oct 2013 | WO | |||
| WO-2013/151665 | Oct 2013 | WO | |||
| WO-2013/151666 | Oct 2013 | WO | |||
| WO-2013/151667 | Oct 2013 | WO | |||
| WO-2013/151668 | Oct 2013 | WO | |||
| WO-2013/151669 | Oct 2013 | WO | |||
| WO-2013/151670 | Oct 2013 | WO | |||
| WO-2013/151671 | Oct 2013 | WO | |||
| WO-2013/151672 | Oct 2013 | WO | |||
| WO-2013/151736 | Oct 2013 | WO | |||
| WO-2013/184976 | Dec 2013 | WO | |||
| WO-2013/185069 | Dec 2013 | WO | |||
| WO-2014/028429 | Feb 2014 | WO | |||
| WO-2014/081507 | May 2014 | WO | |||
| WO-2014/093574 | Jun 2014 | WO | |||
| WO-2014/093622 | Jun 2014 | WO | |||
| WO-2014/093924 | Jun 2014 | WO | |||
| WO-2014/113089 | Jul 2014 | WO | |||
| WO-2014/144039 | Sep 2014 | WO | |||
| WO-2014/144711 | Sep 2014 | WO | |||
| WO-2014/144767 | Sep 2014 | WO | |||
| WO-2014/152027 | Sep 2014 | WO | |||
| WO-2014/152030 | Sep 2014 | WO | |||
| WO-2014/152031 | Sep 2014 | WO | |||
| WO-2014/152211 | Sep 2014 | WO | |||
| WO-2014/152513 | Sep 2014 | WO | |||
| WO-2014/152540 | Sep 2014 | WO | |||
| WO-2014/152659 | Sep 2014 | WO | |||
| WO-2014/152673 | Sep 2014 | WO | |||
| WO-2014/160243 | Oct 2014 | WO | |||
| WO-2014/160284 | Oct 2014 | WO | |||
| WO-2014/164253 | Oct 2014 | WO | |||
| WO-2015/006747 | Jan 2015 | WO | |||
| WO-2015/034925 | Mar 2015 | WO | |||
| WO-2015/034928 | Mar 2015 | WO | |||
| WO-2015/038892 | Mar 2015 | WO | |||
| WO-2015/048744 | Apr 2015 | WO | |||
| WO-2015/051169 | Apr 2015 | WO | |||
| WO-2015/051173 | Apr 2015 | WO | |||
| WO-2015/051214 | Apr 2015 | WO | |||
| WO-2015/058069 | Apr 2015 | WO | |||
| WO-2015/070413 | May 2015 | WO | |||
| WO-2015/085318 | Jun 2015 | WO | |||
| WO-2015/089511 | Jun 2015 | WO | |||
| WO-2015/101416 | Jul 2015 | WO | |||
| WO-2015/105926 | Jul 2015 | WO | |||
| WO-2015/196118 | Dec 2015 | WO | |||
| WO-2015/196128 | Dec 2015 | WO | |||
| WO-2015/196130 | Dec 2015 | WO | |||
| WO-2016/010840 | Jan 2016 | WO | |||
| WO-2016/011222 | Jan 2016 | WO | |||
| WO-2016/011226 | Jan 2016 | WO | |||
| WO-2016/034620 | Mar 2016 | WO | |||
| WO-2016/036902 | Mar 2016 | WO | |||
| WO-2016/077125 | May 2016 | WO | |||
| WO-2016/118724 | Jul 2016 | WO | |||
| WO-2016/118725 | Jul 2016 | WO | |||
Other References
|
International Search Report and Written Opinion for International Patent Application No. PCT/US2014/058891, dated Apr. 2, 2015 (12 pages). cited by applicant . PubChem Compound Summary for CID 262692, created Mar. 26, 2005. <URL: http://pubchem.ncbi.nlm.nih.gov/compound/262692> (11 pages). cited by applicant . PubChem Compound Summary for CID 479886, created Aug. 1, 2005. <URL: http://pubchem.ncbi.nlm.nih.gov/compound/479886> (12 pages). cited by applicant . Kariko et al., "Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability." Mol Ther. 16(11):1833-40 (2008). cited by applicant . Kore et al., "Synthesis and application of 2'-fluoro-substituted cap analogs." Bioorg Med Chem Letters. 17:5295-9 (2007). cited by applicant . Kormann et al., "Expression of therapeutic proteins after delivery of chemically modified mRNA in mice," Nat Biotechnol. 29(2):154-7 (2011) (6 pages). cited by applicant . Mestas et al., "Of mice and not men: differences between mouse and human immunology," J Immunol. 172(5):2731-8 (2004). cited by applicant . Warren et al., "Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA." Cell Stem Cell. 7(5):618-30 (2010). cited by applicant . Kariko et al., "mRNA is an endogenous ligand for Toll-like receptor 3," J Biol Chem. 279(13): 12542-50 (2004). cited by applicant . Yamamoto et al., "Current prospects for mRNA gene delivery," Eur J Pharm Biopharm. 71(3):484-9 (2009). cited by applicant . Grosjean, Modification and editing of RNA: historical overview and important facts to remember. Fine-Tuning of RNA Functions by Modification and Editing. Grosjean H, 1-22 (2005). cited by applicant . Hikishima et al., "Synthesis of 1,8-naphthyridine C-nucleosides and their base-pairing properties in oligodeoxynucleotides: thermally stable naphthyridine:imidazopyridopyrimidine base-pairing motifs," Angew Chem Int Ed. 44:596-8 (2005). cited by applicant . International Preliminary Report on Patentability for International Application No. PCT/US2014/058891, dated Apr. 14, 2016 (8 pages). cited by applicant . Anderson et al., "Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation," Nucleic Acids Res. 38(17):5884-92 (2010). cited by applicant . Anderson et al., "Nucleoside modifications in RNA limit activation of 2'-5'-oligoadenylate synthetase and increase resistance to cleavage by RNase L," Nucleic Acids Res. 39(21): 9329-38 (2011) (10 pages). cited by applicant . Andries et al., "N1-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice," J Control Release. 217:337-44 (2015). cited by applicant . Bellon et al., "4'-Thio-oligo-beta-D-ribonucleotides: synthesis of beta-4'-thio-oligouridylates, nuclease resistance, base pairing properties, and interaction with HIV-1 reverse transcriptase," Nucleic Acids Res. 21(7):1587-93 (1993). cited by applicant . Bellon et al., "Sugar modified oligonucleotides: synthesis, nuclease resistance and base pairing of oligodeoxynucleotides containing 1-(4'-thio-beta-D-ribofuranosyl)-thymine," Biochem Biophys Res Commun. 184(2):797-803 (1992). cited by applicant . Derrigo et al., "RNA-protein interactions in the control of stability and localization of messenger RNA (review)," Int J Mol Med. 5(2):111-23 (2000). cited by applicant . Extended European Search Report for European Application No. 14850286.7, dated May 2, 2017 (9 pages). cited by applicant . Fath et al., "Multiparameter RNA and codon optimization: a standardized tool to assess and enhance autologous mammalian gene expression," PLoS One 6(3):e17596 (2011) (14 pages). cited by applicant . Haeberli et al., "Syntheses of 4'-thioribonucleosides and thermodynamic stability and crystal structure of RNA oligomers with incorporated 4'-thiocytosine," Nucleic Acids Res. 33(13):3965-75 (2005). cited by applicant . Hansen et al., "Circular RNA and miR-7 in Cancer," Cancer Res. 73(18):5609-12 (2013). cited by applicant . Hansen et al., "Natural RNA circles function as efficient microRNA sponges," Nature. 495(7441):384-8 (2013) (7 pages). cited by applicant . Irier et al., "Translational regulation of GluR2 mRNAs in rat hippocampus by alternative 3' untranslated regions," available in PMC Aug. 17, 2009, published in final edited form as: J Neurochem. 109(2):584-594 (2009) (18 pages). cited by applicant . Kariko et al., "Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protien-encoding mRNA," Nucleic Acids Res. 39(21):e142, DOI: 10.1093/nar/gkr695 (2011) (10 pages). cited by applicant . Kariko et al., "Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA," Immunity. 23(2):165-75 (2005). cited by applicant . Kluiver et al., "Rapid generation of MicroRNA Sponges for MicroRNA Inhibition ," PLoS One. 7(1):E29275(2012) (8 pages). cited by applicant . Kuwahara et al., "Molecular evolution of functional nucleic acids with chemical modifications," Molecules. 15(8):5423-44 (2010). cited by applicant . Melton et al., "Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter," Nucleic Acids Res. 12(18):7035-56 (1984). cited by applicant . Memczak et al., "Circular RNAs are a large class of animal RNAs with regulatory potency," Nature. 495(7441):333-8 (2013) (10 pages). cited by applicant . Qiu et al., "Creating a flexible multiple microRNA expression vector by linking precursor microRNAs," Biochem Biophys Res Commun. 411(2):276-80 (2011). cited by applicant . Tavernier et al., "mRNA as gene therapeutic: how to control protein expression," J Control Release. 150(3):238-47 (2011). cited by applicant . Henke et al., "microRNA-122 stimulates translation of hepatitis C virus RNA," Embo J. 27(24):3300-10 (2008). cited by applicant . Loomis et al., "Strategies for modulating innate immune activation and protein production of in vitro transcribed mRNAs," J Mater Chem B. 4(9):1619-32 (2016). cited by applicant . Nakazato et al., "Purification of messenger RNA and heterogeneous nuclear RNA containing poly(A) sequences," Methods Enzymol. 29:431-443 (1974). cited by applicant . Nielsen et al., "An mRNA is capped by a 2',5' lariat catalyzed by a group I-like ribozyme," Science. 309(5740):1584-7 (2005). cited by applicant . Olesiak et al., "The synthesis of di- and oligo-nucleotides containing a phosphorodithioate internucleotide linkage with one of the sulfur atoms in a 5'-bridging position," Org Biomol Chem. 7(10):2162-9 (2009). cited by applicant . Quabius et al., "Synthetic mRNAs for manipulating cellular phenotypes: an overview," N Biotechnol. 32(1):229-35 (2015). cited by applicant . Rodriguez et al., "Magnetic poly (styrene/divinylbenzene/acrylic acid)-based hybrid microspheres for bio-molecular recognition," Micro Nano Lett. 6(6):349-352 (2011). cited by applicant . Stewart et al., "Effect of azide position on the rate of azido glucose-cyclooctyne cycloaddition," Journal of Carbohydrate Chemistry. 33(7-8):408-19 (2014). cited by applicant . Takita et al., "Precise sequential DNA ligation on a solid substrate: solid-based rapid sequential ligation of multiple DNA molecules," DNA Res. 20(6):583-92 (2013). cited by applicant . Thess et al., "Sequence-engineered mRNA Without Chemical Nucleoside Modifications Enables an Effective Protein Therapy in Large Animals," Mol Ther. 23(9):1456-64 (2015). cited by applicant . Thess et al., Supplementary Material for "Sequence-engineered mRNA Without Chemical Nucleoside Modifications Enables an Effective Protein Therapy in Large Animals," Mol Ther. 23(9):1456-64 (2015), accessed via <https://www.sciencedirect.com/science/article/pii/S1525001616302738#c- esec90> (11 Pages). cited by applicant . Virnekas et al., "Trinucleotide phosphoramidites: ideal reagents for the synthesis of mixed oligonucleotides for random mutagenesis," Nucleic Acids Res. 22(25):5600-7 (1994). cited by applicant . Weissman et al., "mRNA: Fulfilling the Promise of Gene Therapy," Mol Ther. 23(9):1416-7 (2015). cited by applicant. |
Primary Examiner: Crane; Lawrence E
Attorney, Agent or Firm: Clark & Elbing LLP
Claims
The invention claimed is:
1. An mRNA 30 to 5000 nucleotides in length encoding a polypeptide, the mRNA comprising at least one backbone moiety having the structure: ##STR00057## wherein B is a nucleobase; or a salt thereof.
2. The mRNA of claim 1, wherein B has the structure: ##STR00058##
3. The mRNA of claim 1, wherein B has the structure: ##STR00059##
4. The mRNA of claim 1, wherein B has the structure: ##STR00060##
5. The mRNA of claim 1, wherein B has the structure: ##STR00061##
6. The mRNA of claim 1, wherein B has the structure: ##STR00062##
7. The mRNA of claim 1, wherein B has the structure: ##STR00063##
8. The mRNA of claim 1, wherein B has the structure: ##STR00064##
9. The mRNA of claim 1, wherein B has the structure: ##STR00065##
10. The mRNA of claim 1, further comprising: (a) a 5' untranslated region; (b) a 3' untranslated region; and (c) a 5' cap structure.
11. The mRNA of claim 10, wherein the 5' cap structure has the structure: ##STR00066## or a salt thereof.
12. The mRNA of claim 10, further comprising a poly-A tail 100 to 250 nucleotides in length at the 3'-terminus of the 3'-untranslated region.
Description
REFERENCE TO A SEQUENCE LISTING
This application contains a Sequence Listing in computer readable form. The computer readable form is incorporated herein by reference.
BACKGROUND
There are multiple problems with prior methodologies of effecting protein expression. For example, heterologous DNA introduced into a cell can be inherited by daughter cells (whether or not the heterologous DNA has integrated into the chromosome) or by offspring. Introduced DNA can integrate into host cell genomic DNA at some frequency, resulting in alterations and/or damage to the host cell genomic DNA. In addition, multiple steps must occur before a protein is made. Once inside the cell, DNA must be transported into the nucleus where it is transcribed into RNA. The RNA transcribed from DNA must then enter the cytoplasm where it is translated into protein. This need for multiple processing steps creates lag times before the generation of a protein of interest. Further, it is difficult to obtain DNA expression in cells; frequently DNA enters cells but is not expressed or not expressed at reasonable rates or concentrations. This can be a particular problem when DNA is introduced into cells such as primary cells or modified cell lines.
Naturally occurring RNAs are synthesized from four basic ribonucleotides: ATP, CTP, UTP and GTP, but may contain post-transcriptionally modified nucleotides. Further, approximately one hundred different nucleoside modifications have been identified in RNA (Rozenski, J, Crain, P, and McCloskey, J. (1999). The RNA Modification Database: 1999 update. Nucl Acids Res 27: 196-197).
There is a need in the art for biological modalities to address the modulation of intracellular translation of polynucleotides. The present invention solves this problem by providing new mRNA molecules incorporating chemical alternatives which impart properties which are advantageous to therapeutic development.
SUMMARY OF THE INVENTION
The present disclosure provides nucleosides, nucleotides, and polynucleotides having an alternative nucleobase, sugar, or backbone and polynucleotides containing the same.
The present invention provides polynucleotides which may be isolated and/or purified. These polynucleotides may encode one or more polypeptides of interest and comprise a sequence of n number of linked nucleosides or nucleotides comprising at least one alternative sugar moiety as compared to ribose. The polynucleotides may also contain a 5' UTR comprising at least one Kozak sequence, a 3' UTR, and at least one 5' cap structure. The isolated polynucleotides may further contain a poly-A tail and may be purified. Such polynucleotides may also be codon optimized.
In a first aspect, the invention features a compound of Formula I:
##STR00001##
wherein the dotted line represents an optional double bond;
B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, O, or NR.sup.7;
R.sup.1 is hydrogen or fluorine;
R.sup.2 is hydrogen, fluorine, cyano, azido, or optionally substituted C.sub.1-C.sub.6 alkyl;
R.sup.3 and R.sup.4 are independently hydrogen, optionally substituted hydroxyl, or fluorine;
R.sup.5 and R.sup.6 are independently hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.5 and R.sup.6 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.5 and R.sup.6 is absent when the dotted line is a double bond;
R.sup.7 is hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl, or optionally substituted amino;
each Y.sup.2 is independently hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
each Y.sup.3 is independently absent, O, or S;
each Y.sup.5 is independently O, NH, or CR.sup.8R.sup.9;
each Y.sup.6 is O or S;
each Y.sup.7 is O or NH; and
each R.sup.8 and R.sup.9 is independently hydrogen, fluorine, or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.8 and R.sup.9 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.8 and R.sup.9 is absent when the dotted line is a double bond;
wherein if n is 0, X is O, R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, and Y.sup.5 is O, then at least one of Y.sup.1 and Y.sup.4 is optionally substituted amino, and, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, Y.sup.7 is O, X is O, and R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, then Y.sup.4 is optionally substituted amino; or a salt thereof.
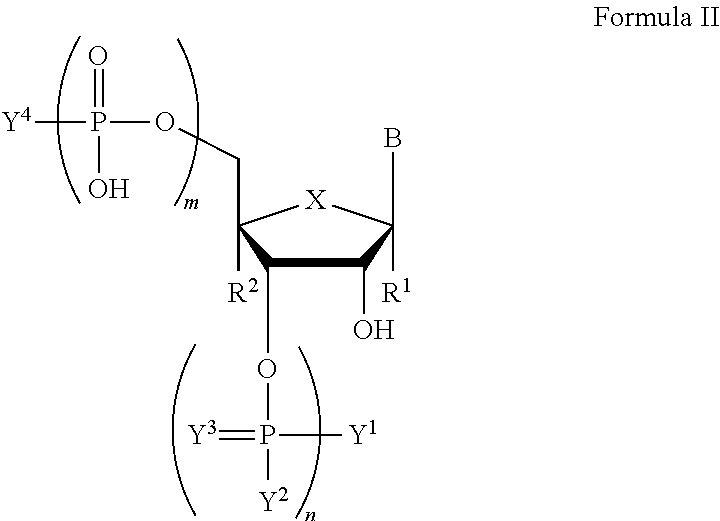
In another aspect, the invention features a compound of Formula II:
##STR00002##
wherein B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, or O;
R.sup.1 and R.sup.2 are independently hydrogen or fluorine;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl (e.g., dimethoxytrityl), or optionally substituted amino;
Y.sup.2 is hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl (e.g., optionally substituted C.sub.1-C.sub.6 alkoxy such as .beta.-cyanoethoxy);
Y.sup.3 is absent or O;
wherein if n is 0, X is O, R.sup.1 and R.sup.2 are hydrogen, then at least one of Y.sup.1 and Y.sup.4 is not hydroxyl or protected hydroxyl, and, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, X is O, and R.sup.1 and R.sup.2 are hydrogen, then Y.sup.4 is not hydroxyl or protected hydroxyl;
or a salt thereof.
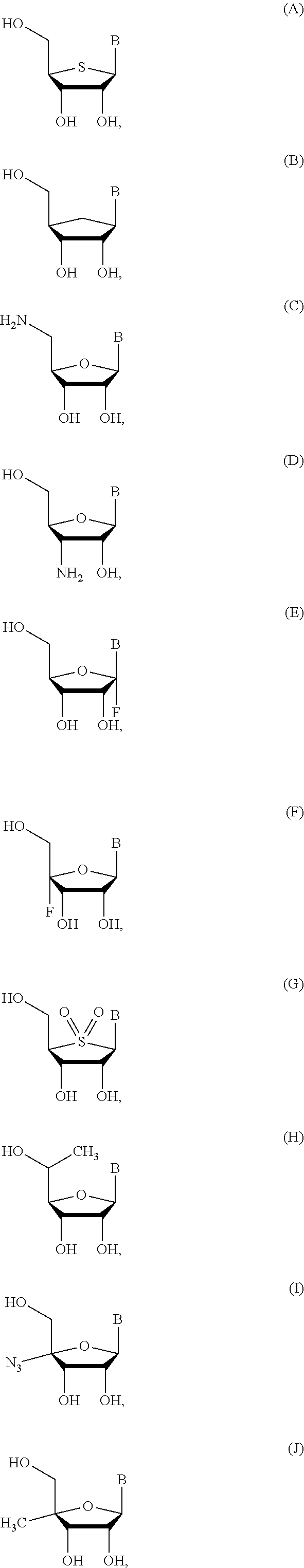
In some embodiments, the compound has the structure:
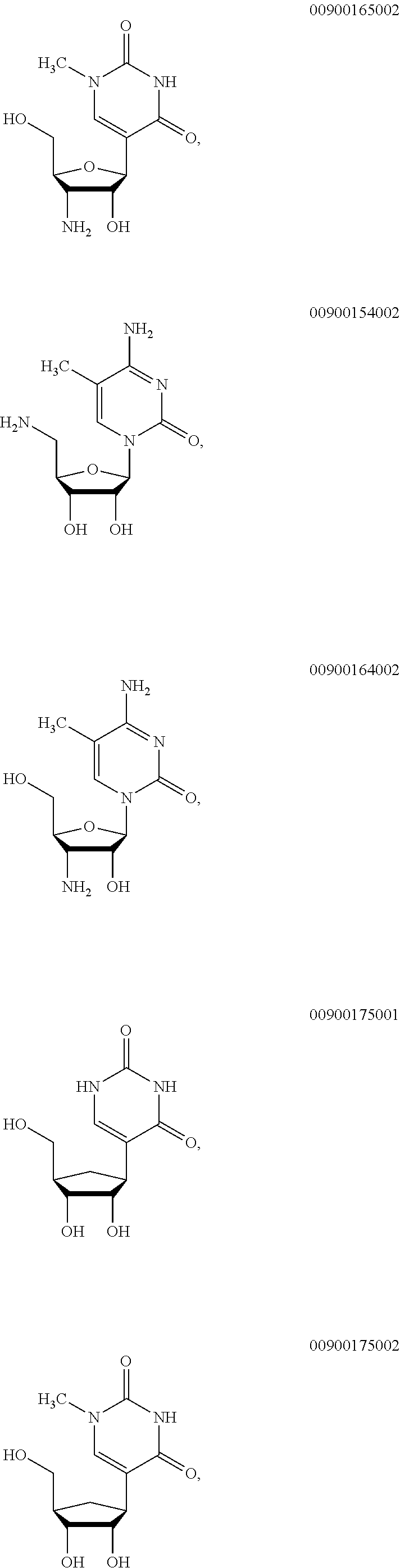
##STR00003## ##STR00004##
wherein m' is an integer from 0 to 2.
In certain embodiments of sugar A, B is uracil. In other embodiments of sugar A, B is pseudouracil. In other embodiments of sugar A, B is 1-methylpseudouracil. In other embodiments of sugar A, B is 5-methoxyuracil. In other embodiments of sugar A, B is cytosine. In other embodiments of sugar A, B is 5-methylcytosine. In other embodiments of sugar A, B is guanine. In other embodiments of sugar A, B is adenine.
In certain embodiments of sugar B, B is uracil. In other embodiments of sugar B, B is pseudouracil. In other embodiments of sugar B, B is 1-methylpseudouracil. In other embodiments of sugar B, B is 5-methoxyuracil. In other embodiments of sugar B, B is cytosine. In other embodiments of sugar B, B is 5-methylcytosine. In other embodiments of sugar B, B is guanine. In other embodiments of sugar B, B is adenine.
In certain embodiments of sugar C, B is uracil. In other embodiments of sugar C, B is pseudouracil. In other embodiments of sugar C, B is 1-methylpseudouracil. In other embodiments of sugar C, B is 5-methoxyuracil. In other embodiments of sugar C, B is cytosine. In other embodiments of sugar C, B is 5-methylcytosine. In other embodiments of sugar C, B is guanine. In other embodiments of sugar C, B is adenine.
In certain embodiments of sugar D, B is uracil. In other embodiments of sugar D, B is pseudouracil. In other embodiments of sugar D, B is 1-methylpseudouracil. In other embodiments of sugar D, B is 5-methoxyuracil. In other embodiments of sugar D, B is cytosine. In other embodiments of sugar D, B is 5-methylcytosine. In other embodiments of sugar D, B is guanine. In other embodiments of sugar D, B is adenine.
In certain embodiments of sugar E, B is uracil. In other embodiments of sugar E, B is pseudouracil. In other embodiments of sugar E, B is 1-methylpseudouracil. In other embodiments of sugar E, B is 5-methoxyuracil. In other embodiments of sugar E, B is cytosine. In other embodiments of sugar E, B is 5-methylcytosine. In other embodiments of sugar E, B is guanine. In other embodiments of sugar E, B is adenine.
In certain embodiments of sugar F, B is uracil. In other embodiments of sugar F, B is pseudouracil. In other embodiments of sugar F, B is 1-methylpseudouracil. In other embodiments of sugar F, B is 5-methoxyuracil. In other embodiments of sugar F, B is cytosine. In other embodiments of sugar F, B is 5-methylcytosine. In other embodiments of sugar F, B is guanine. In other embodiments of sugar F, B is adenine.
In certain embodiments of sugar G, B is uracil. In other embodiments of sugar G, B is pseudouracil. In other embodiments of sugar G, B is 1-methylpseudouracil. In other embodiments of sugar G, B is 5-methoxyuracil. In other embodiments of sugar G, B is cytosine. In other embodiments of sugar G, B is 5-methylcytosine. In other embodiments of sugar G, B is guanine. In other embodiments of sugar G, B is adenine.
In certain embodiments of sugar H, B is uracil. In other embodiments of sugar H, B is pseudouracil. In other embodiments of sugar H, B is 1-methylpseudouracil. In other embodiments of sugar H, B is 5-methoxyuracil. In other embodiments of sugar H, B is cytosine. In other embodiments of sugar H, B is 5-methylcytosine. In other embodiments of sugar H, B is guanine. In other embodiments of sugar H, B is adenine.
In certain embodiments of sugar I, B is uracil. In other embodiments of sugar I, B is pseudouracil. In other embodiments of sugar I, B is 1-methylpseudouracil. In other embodiments of sugar I, B is 5-methoxyuracil. In other embodiments of sugar I, B is cytosine. In other embodiments of sugar I, B is 5-methylcytosine. In other embodiments of sugar I, B is guanine. In other embodiments of sugar I, B is adenine.
In certain embodiments of sugar J, B is uracil. In other embodiments of sugar J, B is pseudouracil. In other embodiments of sugar J, B is 1-methylpseudouracil. In other embodiments of sugar J, B is 5-methoxyuracil. In other embodiments of sugar J, B is cytosine. In other embodiments of sugar J, B is 5-methylcytosine. In other embodiments of sugar J, B is guanine. In other embodiments of sugar J, B is adenine.
In certain embodiments of sugar K, B is uracil. In other embodiments of sugar K, B is pseudouracil. In other embodiments of sugar K, B is 1-methylpseudouracil. In other embodiments of sugar K, B is 5-methoxyuracil. In other embodiments of sugar K, B is cytosine. In other embodiments of sugar K, B is 5-methylcytosine. In other embodiments of sugar K, B is guanine. In other embodiments of sugar K, B is adenine.
In certain embodiments of sugar L, B is uracil. In other embodiments of sugar L, B is pseudouracil. In other embodiments of sugar L, B is 1-methylpseudouracil. In other embodiments of sugar L, B is 5-methoxyuracil. In other embodiments of sugar L, B is cytosine. In other embodiments of sugar L, B is 5-methylcytosine. In other embodiments of sugar L, B is guanine. In other embodiments of sugar L, B is adenine.
In certain embodiments of sugar M, B is uracil. In other embodiments of sugar M, B is pseudouracil. In other embodiments of sugar M, B is 1-methylpseudouracil. In other embodiments of sugar M, B is 5-methoxyuracil. In other embodiments of sugar M, B is cytosine. In other embodiments of sugar M, B is 5-methylcytosine. In other embodiments of sugar M, B is guanine. In other embodiments of sugar M, B is adenine.
In other embodiments, the compound has the structure:
##STR00005##
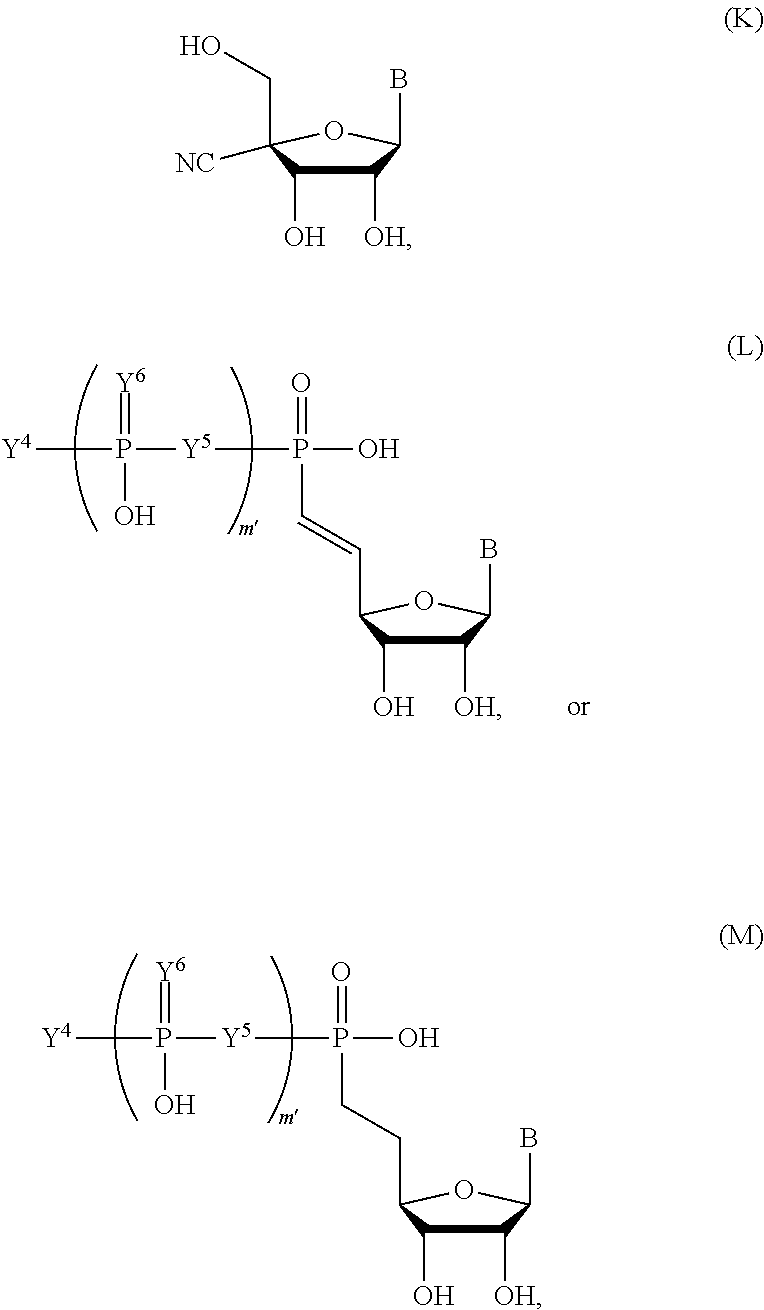
In some embodiments, the compound has the structure:
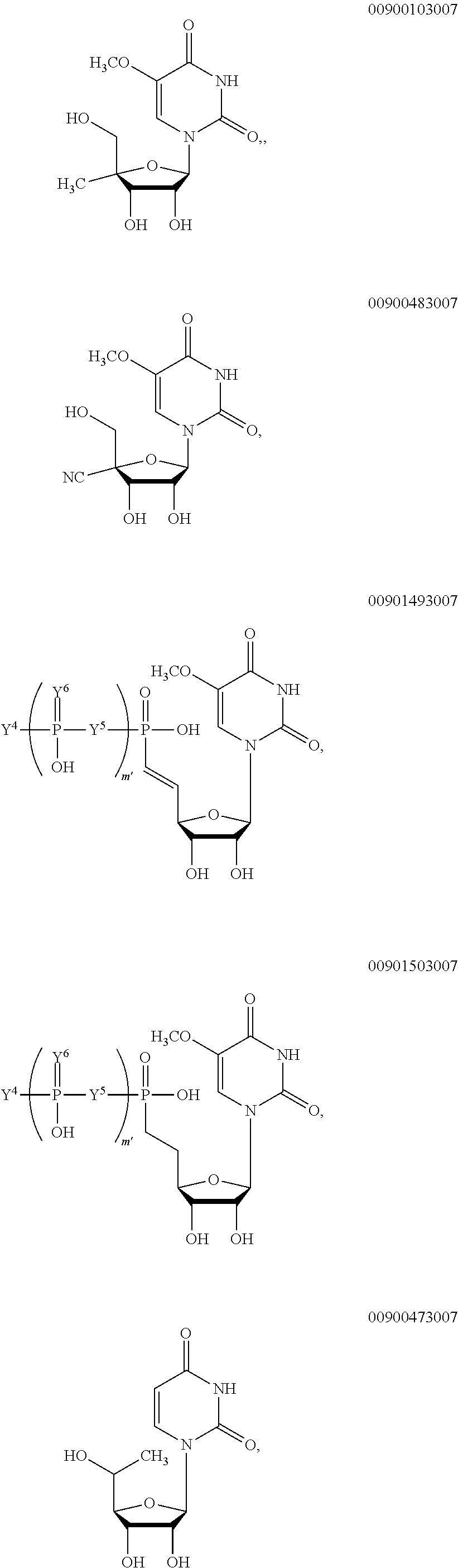
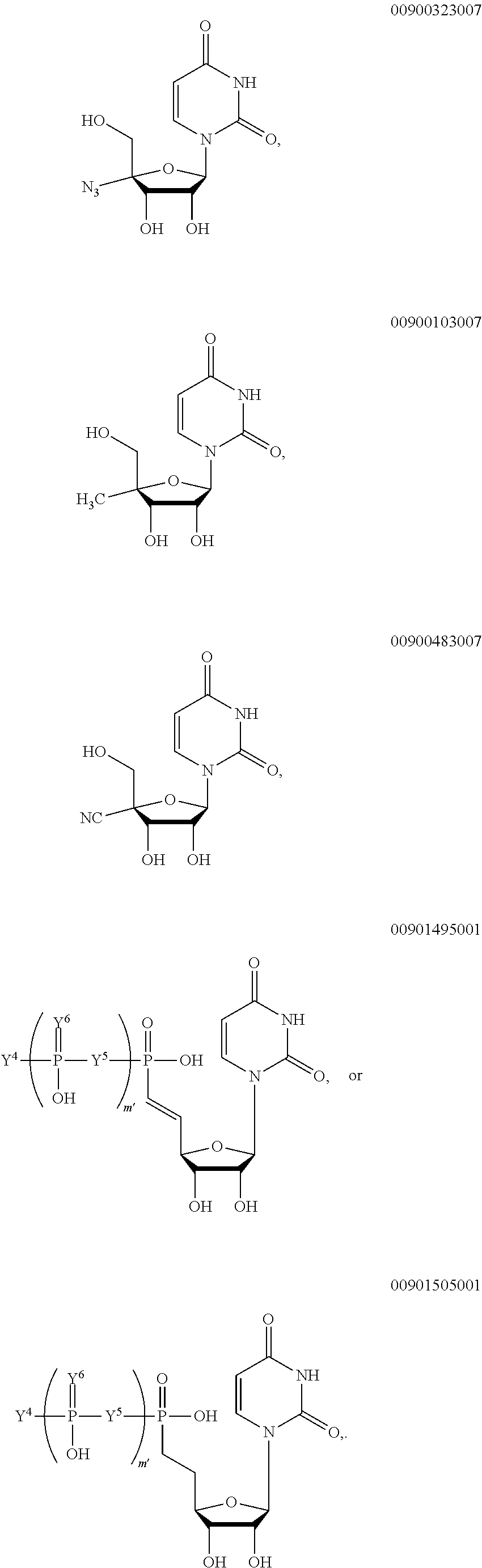
##STR00006## ##STR00007## ##STR00008## ##STR00009## ##STR00010##
wherein m' is an integer from 0 to 2.
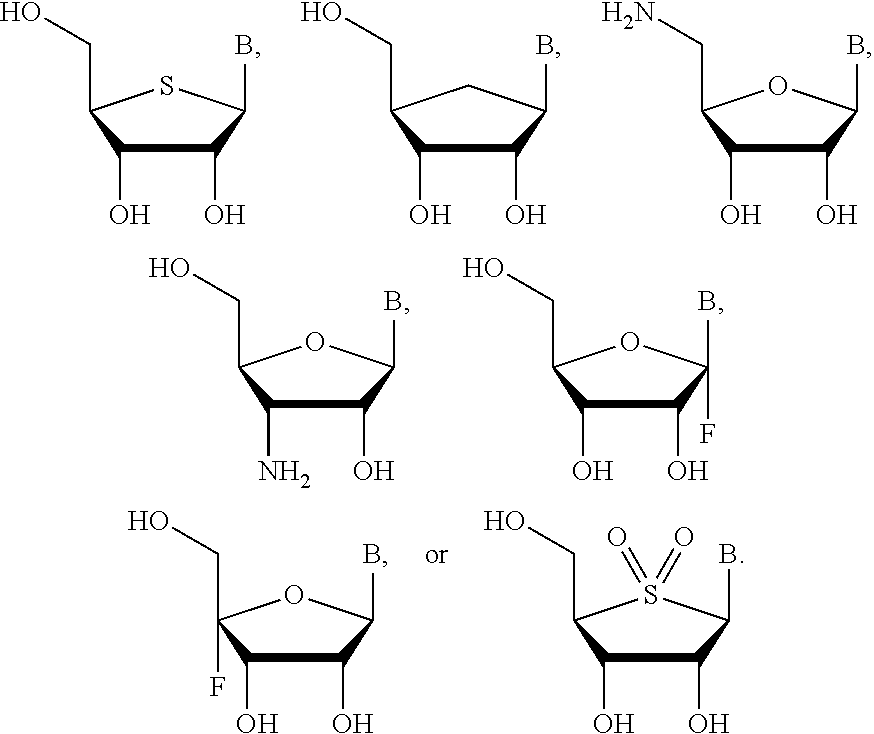
In certain embodiments, the compound has the structure:
##STR00011## ##STR00012## ##STR00013##
In another aspect, the invention features a compound of Formula IA:
##STR00014##
wherein the dotted line represents an optional double bond;
B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, O, or NR.sup.7;
R.sup.1 is hydrogen or fluorine;
R.sup.2 is hydrogen, fluorine, cyano, azido, or optionally substituted C.sub.1-C.sub.6 alkyl;
R.sup.3 and R.sup.4 are independently hydrogen, optionally substituted hydroxyl, or fluorine;
R.sup.5 and R.sup.6 are independently hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.5 and R.sup.6 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.5 and R.sup.6 is absent when the dotted line is a double bond;
R.sup.7 is hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl, or optionally substituted amino;
each Y.sup.2 is independently hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
each Y.sup.3 is independently absent, O, or S;
each Y.sup.5 is independently O, NH, or CR.sup.8R.sup.9;
each Y.sup.6 is O or S;
each Y.sup.7 is O or NH and
each R.sup.8 and R.sup.9 is independently hydrogen, fluorine, or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.8 and R.sup.9 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.8 and R.sup.9 is absent when the dotted line is a double bond;
or a salt thereof.
In some embodiments, if n is 0, X is O, R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, and Y.sup.5 is O, then at least one of Y.sup.1 and Y.sup.4 is optionally substituted amino, and, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, Y.sup.7 is O, X is O, R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, and R.sup.3 is hydroxyl, then Y.sup.4 is optionally substituted amino.
In another aspect, the invention features a compound of Formula IIA:
##STR00015##
wherein B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, or O;
R.sup.1 and R.sup.2 are independently hydrogen or fluorine;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl (e.g., dimethoxytrityl), or optionally substituted amino;
Y.sup.2 is hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl (e.g., optionally substituted C.sub.1-C.sub.6 alkoxy such as .beta.-cyanoethoxy);
Y.sup.3 is absent or O; or a salt thereof.
In certain embodiments, if n is 0, X is O, R.sup.1 and R.sup.2 are hydrogen, then at least one of Y.sup.1 and Y.sup.4 is optionally substituted amino, or, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, X is O, and R.sup.1 and R.sup.2 are hydrogen, then Y.sup.4 is optionally substituted amino.
In some embodiments, the compound has the structure:
##STR00016## ##STR00017##
wherein m' is an integer from 0 to 2.
In certain embodiments of sugar A', B is uracil. In other embodiments of sugar A, B is pseudouracil. In other embodiments of sugar A', B is 1-methylpseudouracil. In other embodiments of sugar A', B is 5-methoxyuracil. In other embodiments of sugar A', B is cytosine. In other embodiments of sugar A', B is 5-methylcytosine. In other embodiments of sugar A', B is guanine. In other embodiments of sugar A', B is adenine.
In certain embodiments of sugar B', B is uracil. In other embodiments of sugar B', B is pseudouracil. In other embodiments of sugar B', B is 1-methylpseudouracil. In other embodiments of sugar B', B is 5-methoxyuracil. In other embodiments of sugar B', B is cytosine. In other embodiments of sugar B', B is 5-methylcytosine. In other embodiments of sugar B', B is guanine. In other embodiments of sugar B', B is adenine.
In certain embodiments of sugar C', B is uracil. In other embodiments of sugar C', B is pseudouracil. In other embodiments of sugar C', B is 1-methylpseudouracil. In other embodiments of sugar C', B is 5-methoxyuracil. In other embodiments of sugar C', B is cytosine. In other embodiments of sugar C', B is 5-methylcytosine. In other embodiments of sugar C', B is guanine. In other embodiments of sugar C', B is adenine.
In certain embodiments of sugar D', B is uracil. In other embodiments of sugar D', B is pseudouracil. In other embodiments of sugar D', B is 1-methylpseudouracil. In other embodiments of sugar D', B is 5-methoxyuracil. In other embodiments of sugar D', B is cytosine. In other embodiments of sugar D', B is 5-methylcytosine. In other embodiments of sugar D', B is guanine. In other embodiments of sugar D', B is adenine.
In certain embodiments of sugar E', B is uracil. In other embodiments of sugar E', B is pseudouracil. In other embodiments of sugar E', B is 1-methylpseudouracil. In other embodiments of sugar E', B is 5-methoxyuracil. In other embodiments of sugar E', B is cytosine. In other embodiments of sugar E', B is 5-methylcytosine. In other embodiments of sugar E', B is guanine. In other embodiments of sugar E', B is adenine.
In certain embodiments of sugar F', B is uracil. In other embodiments of sugar F', B is pseudouracil. In other embodiments of sugar F', B is 1-methylpseudouracil. In other embodiments of sugar F', B is 5-methoxyuracil. In other embodiments of sugar F', B is cytosine. In other embodiments of sugar F', B is 5-methylcytosine. In other embodiments of sugar F', B is guanine. In other embodiments of sugar F', B is adenine.
In certain embodiments of sugar G', B is uracil. In other embodiments of sugar G', B is pseudouracil. In other embodiments of sugar G', B is 1-methylpseudouracil. In other embodiments of sugar G', B is 5-methoxyuracil. In other embodiments of sugar G', B is cytosine. In other embodiments of sugar G', B is 5-methylcytosine. In other embodiments of sugar G', B is guanine. In other embodiments of sugar G', B is adenine.
In certain embodiments of sugar H', B is uracil. In other embodiments of sugar H', B is pseudouracil. In other embodiments of sugar H', B is 1-methylpseudouracil. In other embodiments of sugar H', B is 5-methoxyuracil. In other embodiments of sugar H', B is cytosine. In other embodiments of sugar H', B is 5-methylcytosine. In other embodiments of sugar H', B is guanine. In other embodiments of sugar H', B is adenine.
In certain embodiments of sugar I', B is uracil. In other embodiments of sugar I', B is pseudouracil. In other embodiments of sugar I', B is 1-methylpseudouracil. In other embodiments of sugar I', B is 5-methoxyuracil. In other embodiments of sugar I', B is cytosine. In other embodiments of sugar I', B is 5-methylcytosine. In other embodiments of sugar I', B is guanine. In other embodiments of sugar I', B is adenine.
In certain embodiments of sugar J', B is uracil. In other embodiments of sugar J', B is pseudouracil. In other embodiments of sugar J', B is 1-methylpseudouracil. In other embodiments of sugar J', B is 5-methoxyuracil. In other embodiments of sugar J', B is cytosine. In other embodiments of sugar J', B is 5-methylcytosine. In other embodiments of sugar J', B is guanine. In other embodiments of sugar J', B is adenine.
In certain embodiments of sugar K', B is uracil. In other embodiments of sugar K', B is pseudouracil. In other embodiments of sugar K', B is 1-methylpseudouracil. In other embodiments of sugar K', B is 5-methoxyuracil. In other embodiments of sugar K', B is cytosine. In other embodiments of sugar K', B is 5-methylcytosine. In other embodiments of sugar K', B is guanine. In other embodiments of sugar K', B is adenine.
In certain embodiments of sugar L', B is uracil. In other embodiments of sugar L', B is pseudouracil. In other embodiments of sugar L', B is 1-methylpseudouracil. In other embodiments of sugar L', B is 5-methoxyuracil. In other embodiments of sugar L', B is cytosine. In other embodiments of sugar L', B is 5-methylcytosine. In other embodiments of sugar L', B is guanine. In other embodiments of sugar L', B is adenine.
In certain embodiments of sugar M', B is uracil. In other embodiments of sugar M', B is pseudouracil. In other embodiments of sugar M', B is 1-methylpseudouracil. In other embodiments of sugar M', B is 5-methoxyuracil. In other embodiments of sugar M', B is cytosine. In other embodiments of sugar M', B is 5-methylcytosine. In other embodiments of sugar M', B is guanine. In other embodiments of sugar M', B is adenine.
In some embodiments, the compound has the structure:
##STR00018##
In certain embodiments of compounds of Formula I, Formula II, Formula IA, or Formula IIA, the nucleobase is uracil, pseudouracil, 1-methylpseudouracil, 5-methoxyuracil, cytosine, 5-methylcytosine, guanine, or adenine. Such nucleobases may also be protected with protecting groups as is known in the art. In other embodiments, the compound of Formula I, Formula II, Formula IA, or Formula IIA is a 5' mono-, di-, or triphosphate, and n is 0. In other embodiments, the compound of Formula I, Formula II, Formula IA, or Formula IIA is a 3' phosphoramidite, i.e., n is 1, Y.sup.3 is absent, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl (e.g., .beta.-cyanoethoxy), Y.sup.1 is dialkyl substituted amino, (e.g., diisopropylamino), and m is 0.
In other embodiments, the compound is a 5' mono-, di-, or triphosphate of any of the nucleosides provided herein.
In another aspect, the invention features a polynucleotide comprising at least one backbone moiety of Formula III:
##STR00019##
wherein the dotted line represents an optional double bond;
B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, O, or NR.sup.7;
R.sup.1 is hydrogen or fluorine;
R.sup.2 is hydrogen, fluorine, cyano, azido, or optionally substituted C.sub.1-C.sub.6 alkyl;
R.sup.3 and R.sup.4 are independently hydrogen, optionally substituted hydroxyl, or fluorine;
R.sup.5 and R.sup.6 are independently hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.5 and R.sup.6 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.5 and R.sup.6 is absent when the dotted line is a double bond;
R.sup.7 is hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl;
each Y.sup.2 is independently hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
each Y.sup.3 is independently absent, O, or S;
each Y.sup.5 is independently O, NH, or CR.sup.8R.sup.9;
each Y.sup.6 is independently O or S;
each Y.sup.7 is independently O or NH; and
each R.sup.8 and R.sup.9 is independently hydrogen, fluorine, or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.8 and R.sup.9 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.8 and R.sup.9 is absent when the dotted line is a double bond;
wherein if X is O and R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, then at least one of Y.sup.5 and Y.sup.7 is NH or Y.sup.5 is CR.sup.8R.sup.9;
provided that m and n are both 1 when the backbone moiety is not a 3' or 5' terminal moiety;
or a salt thereof.
In another aspect, the invention features a polynucleotide comprising at least one backbone moiety of Formula IV:
##STR00020##
wherein B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, or O;
R.sup.1 and R.sup.2 are independently hydrogen or fluorine;
Y.sup.2 is hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
Y.sup.3 is absent, O, or S;
wherein if X is O, then at least one of R.sup.1 and R.sup.2 is not hydrogen;
provided that m and n are both 1 when the backbone moiety is not a 3' or 5' terminal moiety;
or a salt thereof.
In certain embodiments, the backbone moiety comprises:
##STR00021## ##STR00022##
In certain embodiments of backbone moiety A, B is uracil. In other embodiments of backbone moiety A, B is pseudouracil. In other embodiments of backbone moiety A, B is 1-methylpseudouracil. In other embodiments of backbone moiety A, B is 5-methoxyuracil. In other embodiments of backbone moiety A, B is cytosine. In other embodiments of backbone moiety A, B is 5-methylcytosine. In other embodiments of backbone moiety A, B is guanine. In other embodiments of backbone moiety A, B is adenine.
In certain embodiments of backbone moiety B, B is uracil. In other embodiments of backbone moiety B, B is pseudouracil. In other embodiments of backbone moiety B, B is 1-methylpseudouracil. In other embodiments of backbone moiety B, B is 5-methoxyuracil. In other embodiments of backbone moiety B, B is cytosine. In other embodiments of backbone moiety B, B is 5-methylcytosine. In other embodiments of backbone moiety B, B is guanine. In other embodiments of backbone moiety B, B is adenine.
In certain embodiments of backbone moiety C, B is uracil. In other embodiments of backbone moiety C, B is pseudouracil. In other embodiments of backbone moiety C, B is 1-methylpseudouracil. In other embodiments of backbone moiety C, B is 5-methoxyuracil. In other embodiments of backbone moiety C, B is cytosine. In other embodiments of backbone moiety C, B is 5-methylcytosine. In other embodiments of backbone moiety C, B is guanine. In other embodiments of backbone moiety C, B is adenine.
In certain embodiments of backbone moiety D, B is uracil. In other embodiments of backbone moiety D, B is pseudouracil. In other embodiments of backbone moiety D, B is 1-methylpseudouracil. In other embodiments of backbone moiety D, B is 5-methoxyuracil. In other embodiments of backbone moiety D, B is cytosine. In other embodiments of backbone moiety D, B is 5-methylcytosine. In other embodiments of backbone moiety D, B is guanine. In other embodiments of backbone moiety D, B is adenine.
In certain embodiments of backbone moiety E, B is uracil. In other embodiments of backbone moiety E, B is pseudouracil. In other embodiments of backbone moiety E, B is 1-methylpseudouracil. In other embodiments of backbone moiety E, B is 5-methoxyuracil. In other embodiments of backbone moiety E, B is cytosine. In other embodiments of backbone moiety E, B is 5-methylcytosine. In other embodiments of backbone moiety E, B is guanine. In other embodiments of backbone moiety E, B is adenine.
In certain embodiments of backbone moiety F, B is uracil. In other embodiments of backbone moiety F, B is pseudouracil. In other embodiments of backbone moiety F, B is 1-methylpseudouracil. In other embodiments of backbone moiety F, B is 5-methoxyuracil. In other embodiments of backbone moiety F, B is cytosine. In other embodiments of backbone moiety F, B is 5-methylcytosine. In other embodiments of backbone moiety F, B is guanine. In other embodiments of backbone moiety F, B is adenine.
In certain embodiments of backbone moiety G, B is uracil. In other embodiments of backbone moiety G, B is pseudouracil. In other embodiments of backbone moiety G, B is 1-methylpseudouracil. In other embodiments of backbone moiety G, B is 5-methoxyuracil. In other embodiments of backbone moiety G, B is cytosine. In other embodiments of backbone moiety G, B is 5-methylcytosine. In other embodiments of backbone moiety G, B is guanine. In other embodiments of backbone moiety G, B is adenine.
In certain embodiments of backbone moiety H, B is uracil. In other embodiments of backbone moiety H, B is pseudouracil. In other embodiments of backbone moiety H, B is 1-methylpseudouracil. In other embodiments of backbone moiety H, B is 5-methoxyuracil. In other embodiments of backbone moiety H, B is cytosine. In other embodiments of backbone moiety H, B is 5-methylcytosine. In other embodiments of backbone moiety H, B is guanine. In other embodiments of backbone moiety H, B is adenine.
In certain embodiments of backbone moiety I, B is uracil. In other embodiments of backbone moiety I, B is pseudouracil. In other embodiments of backbone moiety I, B is 1-methylpseudouracil. In other embodiments of backbone moiety I, B is 5-methoxyuracil. In other embodiments of backbone moiety I, B is cytosine. In other embodiments of backbone moiety I, B is 5-methylcytosine. In other embodiments of backbone moiety I, B is guanine. In other embodiments of backbone moiety I, B is adenine.
In certain embodiments of backbone moiety J, B is uracil. In other embodiments of backbone moiety J, B is pseudouracil. In other embodiments of backbone moiety J, B is 1-methylpseudouracil. In other embodiments of backbone moiety J, B is 5-methoxyuracil. In other embodiments of backbone moiety J, B is cytosine. In other embodiments of backbone moiety J, B is 5-methylcytosine. In other embodiments of backbone moiety J, B is guanine. In other embodiments of backbone moiety J, B is adenine.
In certain embodiments of backbone moiety K, B is uracil. In other embodiments of backbone moiety K, B is pseudouracil. In other embodiments of backbone moiety K, B is 1-methylpseudouracil. In other embodiments of backbone moiety K, B is 5-methoxyuracil. In other embodiments of backbone moiety K, B is cytosine. In other embodiments of backbone moiety K, B is 5-methylcytosine. In other embodiments of backbone moiety K, B is guanine. In other embodiments of backbone moiety K, B is adenine.
In certain embodiments of backbone moiety L, B is uracil. In other embodiments of backbone moiety L, B is pseudouracil. In other embodiments of backbone moiety L, B is 1-methylpseudouracil. In other embodiments of backbone moiety L, B is 5-methoxyuracil. In other embodiments of backbone moiety L, B is cytosine. In other embodiments of backbone moiety L, B is 5-methylcytosine. In other embodiments of backbone moiety L, B is guanine. In other embodiments of backbone moiety L, B is adenine.
In certain embodiments of backbone moiety M, B is uracil. In other embodiments of backbone moiety M, B is pseudouracil. In other embodiments of backbone moiety M, B is 1-methylpseudouracil. In other embodiments of backbone moiety M, B is 5-methoxyuracil. In other embodiments of backbone moiety M, B is cytosine. In other embodiments of backbone moiety M, B is 5-methylcytosine. In other embodiments of backbone moiety M, B is guanine. In other embodiments of backbone moiety M, B is adenine.
In some embodiments, the backbone moiety comprises:
##STR00023##
In another aspect, the invention features a polynucleotide comprising at least one backbone moiety of Formula IIIA:
##STR00024##
wherein the dotted line represents an optional double bond;
B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, O, or NR.sup.7;
R.sup.1 is hydrogen or fluorine;
R.sup.2 is hydrogen, fluorine, cyano, azido, or optionally substituted C.sub.1-C.sub.6 alkyl;
R.sup.3 and R.sup.4 are independently hydrogen, optionally substituted hydroxyl, or fluorine;
R.sup.5 and R.sup.6 are independently hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.5 and R.sup.6 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.5 and R.sup.6 is absent when the dotted line is a double bond;
R.sup.7 is hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl;
each Y.sup.2 is independently hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl
each Y.sup.3 is independently absent, O, or S;
each Y.sup.5 is independently O, NH, or CR.sup.8R.sup.9;
each Y.sup.6 is independently O or S;
each Y.sup.7 is independently O or NH; and
each R.sup.8 and R.sup.9 is independently hydrogen, fluorine, or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.8 and R.sup.9 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.8 and R.sup.9 is absent when the dotted line is a double bond;
provided that m and n are both 1 when the backbone moiety is not a 3' or 5' terminal moiety;
or a salt thereof.
In another aspect, the invention features a polynucleotide comprising at least one backbone moiety of Formula IVA:
##STR00025##
wherein B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, or O;
R.sup.1 and R.sup.2 are independently hydrogen or fluorine;
Y.sup.2 is hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
Y.sup.3 is absent, O, or S;
provided that m and n are both 1 when the backbone moiety is not a 3' or 5' terminal moiety;
or a salt thereof.
In some embodiments, the backbone moiety comprises:
##STR00026## ##STR00027##
In certain embodiments of backbone moiety A', B is uracil. In other embodiments of backbone moiety A, B is pseudouracil. In other embodiments of backbone moiety A', B is 1-methylpseudouracil. In other embodiments of backbone moiety A', B is 5-methoxyuracil. In other embodiments of backbone moiety A', B is cytosine. In other embodiments of backbone moiety A', B is 5-methylcytosine. In other embodiments of backbone moiety A', B is guanine. In other embodiments of backbone moiety A', B is adenine.
In certain embodiments of backbone moiety B', B is uracil. In other embodiments of backbone moiety B', B is pseudouracil. In other embodiments of backbone moiety B', B is 1-methylpseudouracil. In other embodiments of backbone moiety B', B is 5-methoxyuracil. In other embodiments of backbone moiety B', B is cytosine. In other embodiments of backbone moiety B', B is 5-methylcytosine. In other embodiments of backbone moiety B', B is guanine. In other embodiments of backbone moiety B', B is adenine.
In certain embodiments of backbone moiety C', B is uracil. In other embodiments of backbone moiety C', B is pseudouracil. In other embodiments of backbone moiety C', B is 1-methylpseudouracil. In other embodiments of backbone moiety C', B is 5-methoxyuracil. In other embodiments of backbone moiety C', B is cytosine. In other embodiments of backbone moiety C', B is 5-methylcytosine. In other embodiments of backbone moiety C', B is guanine. In other embodiments of backbone moiety C', B is adenine.
In certain embodiments of backbone moiety D', B is uracil. In other embodiments of backbone moiety D', B is pseudouracil. In other embodiments of backbone moiety D', B is 1-methylpseudouracil. In other embodiments of backbone moiety D', B is 5-methoxyuracil. In other embodiments of backbone moiety D', B is cytosine. In other embodiments of backbone moiety D', B is 5-methylcytosine. In other embodiments of backbone moiety D', B is guanine. In other embodiments of backbone moiety D', B is adenine.
In certain embodiments of backbone moiety E', B is uracil. In other embodiments of backbone moiety E', B is pseudouracil. In other embodiments of backbone moiety E', B is 1-methylpseudouracil. In other embodiments of backbone moiety E', B is 5-methoxyuracil. In other embodiments of backbone moiety E', B is cytosine. In other embodiments of backbone moiety E', B is 5-methylcytosine. In other embodiments of backbone moiety E', B is guanine. In other embodiments of backbone moiety E', B is adenine.
In certain embodiments of backbone moiety F', B is uracil. In other embodiments of backbone moiety F', B is pseudouracil. In other embodiments of backbone moiety F', B is 1-methylpseudouracil. In other embodiments of backbone moiety F', B is 5-methoxyuracil. In other embodiments of backbone moiety F', B is cytosine. In other embodiments of backbone moiety F', B is 5-methylcytosine. In other embodiments of backbone moiety F', B is guanine. In other embodiments of backbone moiety F', B is adenine.
In certain embodiments of backbone moiety G', B is uracil. In other embodiments of backbone moiety G', B is pseudouracil. In other embodiments of backbone moiety G', B is 1-methylpseudouracil. In other embodiments of backbone moiety G', B is 5-methoxyuracil. In other embodiments of backbone moiety G', B is cytosine. In other embodiments of backbone moiety G', B is 5-methylcytosine. In other embodiments of backbone moiety G', B is guanine. In other embodiments of backbone moiety G', B is adenine.
In certain embodiments of backbone moiety H', B is uracil. In other embodiments of backbone moiety H', B is pseudouracil. In other embodiments of backbone moiety H', B is 1-methylpseudouracil. In other embodiments of backbone moiety H', B is 5-methoxyuracil. In other embodiments of backbone moiety H', B is cytosine. In other embodiments of backbone moiety H', B is 5-methylcytosine. In other embodiments of backbone moiety H', B is guanine. In other embodiments of backbone moiety H', B is adenine.
In certain embodiments of backbone moiety I', B is uracil. In other embodiments of backbone moiety I', B is pseudouracil. In other embodiments of backbone moiety I', B is 1-methylpseudouracil. In other embodiments of backbone moiety I', B is 5-methoxyuracil. In other embodiments of backbone moiety I', B is cytosine. In other embodiments of backbone moiety I', B is 5-methylcytosine. In other embodiments of backbone moiety I', B is guanine. In other embodiments of backbone moiety I', B is adenine.
In certain embodiments of backbone moiety J', B is uracil. In other embodiments of backbone moiety J', B is pseudouracil. In other embodiments of backbone moiety J', B is 1-methylpseudouracil. In other embodiments of backbone moiety J', B is 5-methoxyuracil. In other embodiments of backbone moiety J', B is cytosine. In other embodiments of backbone moiety J', B is 5-methylcytosine. In other embodiments of backbone moiety J', B is guanine. In other embodiments of backbone moiety J', B is adenine.
In certain embodiments of backbone moiety K', B is uracil. In other embodiments of backbone moiety K', B is pseudouracil. In other embodiments of backbone moiety K', B is 1-methylpseudouracil. In other embodiments of backbone moiety K', B is 5-methoxyuracil. In other embodiments of backbone moiety K', B is cytosine. In other embodiments of backbone moiety K', B is 5-methylcytosine. In other embodiments of backbone moiety K', B is guanine. In other embodiments of backbone moiety K', B is adenine.
In certain embodiments of backbone moiety L', B is uracil. In other embodiments of backbone moiety L', B is pseudouracil. In other embodiments of backbone moiety L', B is 1-methylpseudouracil. In other embodiments of backbone moiety L', B is 5-methoxyuracil. In other embodiments of backbone moiety L', B is cytosine. In other embodiments of backbone moiety L', B is 5-methylcytosine. In other embodiments of backbone moiety L', B is guanine. In other embodiments of backbone moiety L', B is adenine.
In certain embodiments of backbone moiety M', B is uracil. In other embodiments of backbone moiety M', B is pseudouracil. In other embodiments of backbone moiety M', B is 1-methylpseudouracil. In other embodiments of backbone moiety M', B is 5-methoxyuracil. In other embodiments of backbone moiety M', B is cytosine. In other embodiments of backbone moiety M', B is 5-methylcytosine. In other embodiments of backbone moiety M', B is guanine. In other embodiments of backbone moiety M', B is adenine.
In some embodiments, the backbone moiety comprises:
##STR00028##
In certain embodiments of the polynucleotides, the nucleobase is uracil, pseudouracil, 1-methylpseudouracil, 5-methoxyuracil, cytosine, 5-methylcytosine, guanine, or adenine.
In some embodiments, the backbone moiety comprises:
##STR00029## ##STR00030## ##STR00031## ##STR00032## ##STR00033##
In certain embodiments, the backbone moiety comprises:
##STR00034## ##STR00035## ##STR00036##
In some embodiments, the polynucleotide further includes:
(a) a 5' UTR comprising at least one Kozak sequence;
(b) a 3' UTR; and
(c) at least one 5' cap structure.
In other embodiments, the at least one 5' cap structure is Cap0, Cap1, ARCA, inosine, N1-methyl-guanosine, 2'-fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino-guanosine, LNA-guanosine, or 2-azido-guanosine.
In some embodiments, the polynucleotide further includes a poly-A tail.
In certain embodiments, the polynucleotide encodes a protein of interest.
In some embodiments, the polynucleotide is purified.
The present invention also provides for pharmaceutical compositions comprising the polynucleotides described herein. These may also further include one or more pharmaceutically acceptable excipients selected from a solvent, aqueous solvent, non-aqueous solvent, dispersion media, diluent, dispersion, suspension aid, surface active agent, isotonic agent, thickening or emulsifying agent, preservative, lipid, lipidoids liposome, lipid nanoparticle, core-shell nanoparticles, polymer, lipoplexe peptide, protein, cell, hyaluronidase, and mixtures thereof.
Methods of using the polynucleotides of the invention are also provided. In this instance, the polynucleotides may be formulated by any means known in the art or administered via any of several routes including injection by intradermal, subcutaneous or intramuscular means.
Administration of the polynucleotides of the invention may be via two or more equal or unequal split doses. In some embodiments, the level of the polypeptide produced by the subject by administering split doses of the polynucleotide is greater than the levels produced by administering the same total daily dose of polynucleotide as a single administration.
Detection of the polynucleotides of the invention or the encoded polypeptides may be performed in the bodily fluid of the subject or patient where the bodily fluid is selected from the group consisting of peripheral blood, serum, plasma, ascites, urine, cerebrospinal fluid (CSF), sputum, saliva, bone marrow, synovial fluid, aqueous humor, amniotic fluid, cerumen, breast milk, broncheoalveolar lavage fluid, semen, prostatic fluid, cowper's fluid or pre-ejaculatory fluid, sweat, fecal matter, hair, tears, cyst fluid, pleural and peritoneal fluid, pericardial fluid, lymph, chyme, chyle, bile, interstitial fluid, menses, pus, sebum, vomit, vaginal secretions, mucosal secretion, stool water, pancreatic juice, lavage fluids from sinus cavities, bronchopulmonary aspirates, blastocyl cavity fluid, and umbilical cord blood.
In some embodiments, administration is according to a dosing regimen which occurs over the course of hours, days, weeks, months, or years and may be achieved by using one or more devices selected from multi-needle injection systems, catheter or lumen systems, and ultrasound, electrical or radiation based systems.
Chemical Terms
The names of nucleobases correspond to the name given to the base when part of a nucleoside or nucleotide. For example "pseudo-uracil" refers to the nucleobase of pseudouridine.
As used herein, the term "compound," is meant to include all stereoisomers, geometric isomers, tautomers, and isotopes of the structures depicted.
The compounds described herein can be asymmetric (e.g., having one or more stereocenters). All stereoisomers, such as enantiomers and diastereomers, are intended unless otherwise indicated. Compounds of the present disclosure that contain asymmetrically substituted carbon atoms can be isolated in optically active or racemic forms. Methods on how to prepare optically active forms from optically active starting materials are known in the art, such as by resolution of racemic mixtures or by stereoselective synthesis. Many geometric isomers of olefins, C.dbd.N double bonds, and the like can also be present in the compounds described herein, and all such stable isomers are contemplated in the present disclosure. Cis and trans geometric isomers of the compounds of the present disclosure are described and may be isolated as a mixture of isomers or as separated isomeric forms.
Compounds of the present disclosure also include tautomeric forms. Tautomeric forms result from the swapping of a single bond with an adjacent double bond and the concomitant migration of a proton. Tautomeric forms include prototropic tautomers which are isomeric protonation states having the same empirical formula and total charge. Examples prototropic tautomers include ketone-enol pairs, amide-imidic acid pairs, lactam-lactim pairs, amide-imidic acid pairs, enamine-imine pairs, and annular forms where a proton can occupy two or more positions of a heterocyclic system, such as, 1H- and 3H-imidazole, 1H-, 2H- and 4H-1,2,4-triazole, 1H- and 2H-isoindole, and 1H- and 2H-pyrazole. Tautomeric forms can be in equilibrium or sterically locked into one form by appropriate substitution.
Compounds of the present disclosure also include all of the isotopes of the atoms occurring in the intermediate or final compounds. "Isotopes" refers to atoms having the same atomic number but different mass numbers resulting from a different number of neutrons in the nuclei. For example, isotopes of hydrogen include tritium and deuterium.
The compounds and salts of the present disclosure can be prepared in combination with solvent or water molecules to form solvates and hydrates by routine methods.
At various places in the present specification, substituents of compounds of the present disclosure are disclosed in groups or in ranges. It is specifically intended that the present disclosure include each and every individual subcombination of the members of such groups and ranges. For example, the term "C.sub.1-6 alkyl" is specifically intended to individually disclose methyl, ethyl, C.sub.3 alkyl, C.sub.4 alkyl, C.sub.5 alkyl, and C.sub.6 alkyl. Herein a phrase of the form "optionally substituted X" (e.g., optionally substituted alkyl) is intended to be equivalent to "X, wherein X is optionally substituted" (e.g., "alkyl, wherein said alkyl is optionally substituted"). It is not intended to mean that the feature "X" (e.g. alkyl) per se is optional.
The term "acyl," as used herein, represents a hydrogen or an alkyl group (e.g., a haloalkyl group), as defined herein, that is attached to the parent molecular group through a carbonyl group, as defined herein, and is exemplified by formyl (i.e., a carboxyaldehyde group), acetyl, trifluoroacetyl, propionyl, butanoyl and the like. Exemplary unsubstituted acyl groups include from 1 to 7, from 1 to 11, or from 1 to 21 carbons. In some embodiments, the alkyl group is further substituted with 1, 2, 3, or 4 substituents as described herein.
Non-limiting examples of optionally substituted acyl groups include, alkoxycarbonyl, alkoxycarbonylacyl, arylalkoxycarbonyl, aryloyl, carbamoyl, carboxyaldehyde, (heterocyclyl) imino, and (heterocyclyl)oyl:
The "alkoxycarbonyl" group, which as used herein, represents an alkoxy, as defined herein, attached to the parent molecular group through a carbonyl atom (e.g., --C(O)--OR, where R is H or an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group). Exemplary unsubstituted alkoxycarbonyl include from 1 to 21 carbons (e.g., from 1 to 11 or from 1 to 7 carbons). In some embodiments, the alkoxy group is further substituted with 1, 2, 3, or 4 substituents as described herein.
The "alkoxycarbonylacyl" group, which as used herein, represents an acyl group, as defined herein, that is substituted with an alkoxycarbonyl group, as defined herein (e.g., --C(O)-alkyl-C(O)--OR, where R is an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group). Exemplary unsubstituted alkoxycarbonylacyl include from 3 to 41 carbons (e.g., from 3 to 10, from 3 to 13, from 3 to 17, from 3 to 21, or from 3 to 31 carbons, such as C.sub.1-6 alkoxycarbonyl-C.sub.1-6 acyl, C.sub.1-10 alkoxycarbonyl-C.sub.1-10 acyl, or C.sub.1-20 alkoxycarbonyl-C.sub.1-20 acyl). In some embodiments, each alkoxy and alkyl group is further independently substituted with 1, 2, 3, or 4 substituents, as described herein (e.g., a hydroxyl group) for each group.
The "arylalkoxycarbonyl" group, which as used herein, represents an arylalkoxy group, as defined herein, attached to the parent molecular group through a carbonyl (e.g., --C(O)--O-alkyl-aryl). Exemplary unsubstituted arylalkoxy groups include from 8 to 31 carbons (e.g., from 8 to 17 or from 8 to 21 carbons, such as C.sub.6-10 aryl-C.sub.1-6 alkoxy-carbonyl, C.sub.6-10 aryl-C.sub.1-10 alkoxy-carbonyl, or C.sub.6-10 aryl-C.sub.1-20 alkoxy-carbonyl). In some embodiments, the arylalkoxycarbonyl group can be substituted with 1, 2, 3, or 4 substituents as defined herein.
The "aryloyl" group, which as used herein, represents an aryl group, as defined herein, that is attached to the parent molecular group through a carbonyl group. Exemplary unsubstituted aryloyl groups are of 7 to 11 carbons. In some embodiments, the aryl group can be substituted with 1, 2, 3, or 4 substituents as defined herein.
The "carbamoyl" group, which as used herein, represents --C(O)--N(R.sup.N1).sub.2, where the meaning of each R.sup.N1 is found in the definition of "amino" provided herein.
The "carboxyaldehyde" group, which as used herein, represents an acyl group having the structure --CHO.
The "(heterocyclyl) imino" group, which as used herein, represents a heterocyclyl group, as defined herein, attached to the parent molecular group through an imino group. In some embodiments, the heterocyclyl group can be substituted with 1, 2, 3, or 4 substituent groups as defined herein.
The "(heterocyclyl)oyl" group, which as used herein, represents a heterocyclyl group, as defined herein, attached to the parent molecular group through a carbonyl group. In some embodiments, the heterocyclyl group can be substituted with 1, 2, 3, or 4 substituent groups as defined herein.
The term "alkyl," as used herein, is inclusive of both straight chain and branched chain saturated groups from 1 to 20 carbons (e.g., from 1 to 10 or from 1 to 6), unless otherwise specified. Alkyl groups are exemplified by methyl, ethyl, n- and iso-propyl, n-, sec-, iso- and tert-butyl, neopentyl, and the like, and may be optionally substituted with one, two, three, or, in the case of alkyl groups of two carbons or more, four substituents independently selected from the group consisting of: (1) C.sub.1-6 alkoxy; (2) C.sub.1-6 alkylsulfinyl; (3) amino, as defined herein (e.g., unsubstituted amino (i.e., --NH.sub.2) or a substituted amino (i.e., --N(R.sup.N1).sub.2, where R.sup.N1 is as defined for amino); (4) C.sub.6-10 aryl-C.sub.1-6 alkoxy; (5) azido; (6) halo; (7) (C.sub.2-9 heterocyclyl)oxy; (8) hydroxyl, optionally substituted with an O-protecting group; (9) nitro; (10) oxo (e.g., carboxyaldehyde or acyl); (11) C.sub.1-7 spirocyclyl; (12) thioalkoxy; (13) thiol; (14) --CO.sub.2R.sup.A', optionally substituted with an O-protecting group and where R.sup.A' is selected from the group consisting of (a) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c) C.sub.6-10 aryl, (d) hydrogen, (e) C.sub.1-6 alk-C.sub.6-10 aryl, (f) amino-C.sub.1-20 alkyl, (g) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1(CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; (15) --C(O)NR.sup.B'R.sup.C', where each of R.sup.B' and R.sup.C' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (16) --SO.sub.2R.sup.D', where R.sup.D' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) C.sub.1-6 alk-C.sub.6-10 aryl, and (d) hydroxyl; (17) --SO.sub.2NR.sup.E'R.sup.F', where each of R.sup.E' and R.sup.F' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (18) --C(O)R.sup.G', where R.sup.G' is selected from the group consisting of (a) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c) C.sub.6-10 aryl, (d) hydrogen, (e) C.sub.1-6 alk-C.sub.6-10 aryl, (f) amino-C.sub.1-20 alkyl, (g) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1(CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; (19) --NR.sup.H'C(O)R.sup.I', wherein R.sup.H' is selected from the group consisting of (a1) hydrogen and (b1) C.sub.1-6 alkyl, and R.sup.I' is selected from the group consisting of (a2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b2) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c2) C.sub.6-10 aryl, (d2) hydrogen, (e2) C.sub.1-6 alk-C.sub.6-10 aryl, (f2) amino-C.sub.1-20 alkyl, (g2) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1(CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h2) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; (20) --NR.sup.J'C(O)OR.sup.K', wherein R.sup.J' is selected from the group consisting of (a1) hydrogen and (b1) C.sub.1-6 alkyl, and R.sup.K' is selected from the group consisting of (a2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b2) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c2) C.sub.6-10 aryl, (d2) hydrogen, (e2) C.sub.1-6 alk-C.sub.6-10 aryl, (f2) amino-C.sub.1-20 alkyl, (g2) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1(CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h2) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; and (21) amidine. In some embodiments, each of these groups can be further substituted as described herein. For example, the alkylene group of a C.sub.1-alkaryl can be further substituted with an oxo group to afford the respective aryloyl substituent.
The term "alkylene" and the prefix "alk-," as used herein, represent a saturated divalent hydrocarbon group derived from a straight or branched chain saturated hydrocarbon by the removal of two hydrogen atoms, and is exemplified by methylene, ethylene, isopropylene, and the like. The term "C.sub.x-y alkylene" and the prefix "C.sub.x-y alk-" represent alkylene groups having between x and y carbons. Exemplary values for x are 1, 2, 3, 4, 5, and 6, and exemplary values for y are 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 14, 16, 18, or 20 (e.g., C.sub.1-6, C.sub.1-10, C.sub.2-20, C.sub.2-6, C.sub.2-10, or C.sub.2-20 alkylene). In some embodiments, the alkylene can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein for an alkyl group.
Non-limiting examples of optionally substituted alkyl and alkylene groups include acylaminoalkyl, acyloxyalkyl, alkoxyalkyl, alkoxycarbonylalkyl, alkylsulfinyl, alkylsulfinylalkyl, aminoalkyl, carbamoylalkyl, carboxyalkyl, carboxyaminoalkyl, haloalkyl, hydroxyalkyl, perfluoroalkyl, and sulfoalkyl:
The "acylaminoalkyl" group, which as used herein, represents an acyl group, as defined herein, attached to an amino group that is in turn attached to the parent molecular group through an alkylene group, as defined herein (i.e., -alkyl-N(R.sup.N1)--C(O)--R, where R is H or an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group (e.g., haloalkyl) and R.sup.N1 is as defined herein). Exemplary unsubstituted acylaminoalkyl groups include from 1 to 41 carbons (e.g., from 1 to 7, from 1 to 13, from 1 to 21, from 2 to 7, from 2 to 13, from 2 to 21, or from 2 to 41 carbons). In some embodiments, the alkylene group is further substituted with 1, 2, 3, or 4 substituents as described herein, and/or the amino group is --NH.sub.2 or --NHR.sup.N1, wherein R.sup.N1 is, independently, OH, NO.sub.2, NH.sub.2, NR.sup.N2.sub.2, SO.sub.2OR.sup.N2, SO.sub.2R.sup.N2, SOR.sup.N2, alkyl, aryl, acyl (e.g., acetyl, trifluoroacetyl, or others described herein), or alkoxycarbonylalkyl, and each R.sup.N2 can be H, alkyl, or aryl.
The "acyloxyalkyl" group, which as used herein, represents an acyl group, as defined herein, attached to an oxygen atom that in turn is attached to the parent molecular group though an alkylene group (i.e., -alkyl-O--C(O)--R, where R is H or an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group). Exemplary unsubstituted acyloxyalkyl groups include from 1 to 21 carbons (e.g., from 1 to 7 or from 1 to 11 carbons). In some embodiments, the alkylene group is, independently, further substituted with 1, 2, 3, or 4 substituents as described herein.
The "alkoxyalkyl" group, which as used herein, represents an alkyl group that is substituted with an alkoxy group. Exemplary unsubstituted alkoxyalkyl groups include between 2 to 40 carbons (e.g., from 2 to 12 or from 2 to 20 carbons, such as C.sub.1-6 alkoxy-C.sub.1-6 alkyl, C.sub.1-10 alkoxy-C.sub.1-10 alkyl, or C.sub.1-20 alkoxy-C.sub.1-20 alkyl). In some embodiments, the alkyl and the alkoxy each can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein for the respective group.
The "alkoxycarbonylalkyl" group, which as used herein, represents an alkyl group, as defined herein, that is substituted with an alkoxycarbonyl group, as defined herein (e.g., -alkyl-C(O)--OR, where R is an optionally substituted C.sub.1-20, C.sub.1-10, or C.sub.1-6 alkyl group). Exemplary unsubstituted alkoxycarbonylalkyl include from 3 to 41 carbons (e.g., from 3 to 10, from 3 to 13, from 3 to 17, from 3 to 21, or from 3 to 31 carbons, such as C.sub.1-6 alkoxycarbonyl-C.sub.1-6 alkyl, C.sub.1-10 alkoxycarbonyl-C.sub.1-10 alkyl, or C.sub.1-20 alkoxycarbonyl-C.sub.1-20 alkyl). In some embodiments, each alkyl and alkoxy group is further independently substituted with 1, 2, 3, or 4 substituents as described herein (e.g., a hydroxyl group).
The "alkylsulfinylalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with an alkylsulfinyl group. Exemplary unsubstituted alkylsulfinylalkyl groups are from 2 to 12, from 2 to 20, or from 2 to 40 carbons. In some embodiments, each alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein.
The "aminoalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with an amino group, as defined herein. The alkyl and amino each can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for the respective group (e.g., CO.sub.2R.sup.A', where R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl, e.g., carboxy, and/or an N-protecting group).
The "carbamoylalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with a carbamoyl group, as defined herein. The alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein.
The "carboxyalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with a carboxy group, as defined herein. The alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein, and the carboxy group can be optionally substituted with one or more O-protecting groups.
The "carboxyaminoalkyl" group, which as used herein, represents an aminoalkyl group, as defined herein, substituted with a carboxy, as defined herein. The carboxy, alkyl, and amino each can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for the respective group (e.g., CO.sub.2R.sup.A', where R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl, e.g., carboxy, and/or an N-protecting group, and/or an O-protecting group).
The "haloalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with a halogen group (i.e., F, Cl, Br, or I). A haloalkyl may be substituted with one, two, three, or, in the case of alkyl groups of two carbons or more, four halogens. Haloalkyl groups include perfluoroalkyls (e.g., --CF.sub.3), --CHF.sub.2, --CH.sub.2F, --CCl.sub.3, --CH.sub.2CH.sub.2Br, --CH.sub.2CH(CH.sub.2CH.sub.2Br)CH.sub.3, and --CHICH.sub.3. In some embodiments, the haloalkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for alkyl groups.
The "hydroxyalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with one to three hydroxyl groups, with the proviso that no more than one hydroxyl group may be attached to a single carbon atom of the alkyl group, and is exemplified by hydroxymethyl, dihydroxypropyl, and the like. In some embodiments, the hydroxyalkyl group can be substituted with 1, 2, 3, or 4 substituent groups (e.g., O-protecting groups) as defined herein for an alkyl.
The "perfluoroalkyl" group, which as used herein, represents an alkyl group, as defined herein, where each hydrogen radical bound to the alkyl group has been replaced by a fluoride radical. Perfluoroalkyl groups are exemplified by trifluoromethyl, pentafluoroethyl, and the like.
The "sulfoalkyl" group, which as used herein, represents an alkyl group, as defined herein, substituted with a sulfo group of --SO.sub.3H. In some embodiments, the alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein, and the sulfo group can be further substituted with one or more O-protecting groups (e.g., as described herein).
The term "alkenyl," as used herein, represents monovalent straight or branched chain groups of, unless otherwise specified, from 2 to 20 carbons (e.g., from 2 to 6 or from 2 to 10 carbons) containing one or more carbon-carbon double bonds and is exemplified by ethenyl, 1-propenyl, 2-propenyl, 2-methyl-1-propenyl, 1-butenyl, 2-butenyl, and the like. Alkenyls include both cis and trans isomers. Alkenyl groups may be optionally substituted with 1, 2, 3, or 4 substituent groups that are selected, independently, from amino, aryl, cycloalkyl, or heterocyclyl (e.g., heteroaryl), as defined herein, or any of the exemplary alkyl substituent groups described herein.
Non-limiting examples of optionally substituted alkenyl groups include, alkoxycarbonylalkenyl, aminoalkenyl, and hydroxyalkenyl:
The "alkoxycarbonylalkenyl" group, which as used herein, represents an alkenyl group, as defined herein, that is substituted with an alkoxycarbonyl group, as defined herein (e.g., -alkenyl-C(O)--OR, where R is an optionally substituted C.sub.1-20, C.sub.1-10, or C.sub.1-6 alkyl group). Exemplary unsubstituted alkoxycarbonylalkenyl include from 4 to 41 carbons (e.g., from 4 to 10, from 4 to 13, from 4 to 17, from 4 to 21, or from 4 to 31 carbons, such as C.sub.1-6 alkoxycarbonyl-C.sub.2-6 alkenyl, C.sub.1-10 alkoxycarbonyl-C.sub.2-10 alkenyl, or C.sub.1-20 alkoxycarbonyl-C.sub.2-20 alkenyl). In some embodiments, each alkyl, alkenyl, and alkoxy group is further independently substituted with 1, 2, 3, or 4 substituents as described herein (e.g., a hydroxyl group).
The "aminoalkenyl" group, which as used herein, represents an alkenyl group, as defined herein, substituted with an amino group, as defined herein. The alkenyl and amino each can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for the respective group (e.g., CO.sub.2R.sup.A', where R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl, e.g., carboxy, and/or an N-protecting group).
The "hydroxyalkenyl" group, which as used herein, represents an alkenyl group, as defined herein, substituted with one to three hydroxyl groups, with the proviso that no more than one hydroxyl group may be attached to a single carbon atom of the alkyl group, and is exemplified by dihydroxypropenyl, hydroxyisopentenyl, and the like. In some embodiments, the hydroxyalkenyl group can be substituted with 1, 2, 3, or 4 substituent groups (e.g., O-protecting groups) as defined herein for an alkyl.
The term "alkynyl," as used herein, represents monovalent straight or branched chain groups from 2 to 20 carbon atoms (e.g., from 2 to 4, from 2 to 6, or from 2 to 10 carbons) containing a carbon-carbon triple bond and is exemplified by ethynyl, 1-propynyl, and the like. Alkynyl groups may be optionally substituted with 1, 2, 3, or 4 substituent groups that are selected, independently, from aryl, cycloalkyl, or heterocyclyl (e.g., heteroaryl), as defined herein, or any of the exemplary alkyl substituent groups described herein.
Non-limiting examples of optionally substituted alkynyl groups include alkoxycarbonylalkynyl, aminoalkynyl, and hydroxyalkynyl:
The "alkoxycarbonylalkynyl" group, which as used herein, represents an alkynyl group, as defined herein, that is substituted with an alkoxycarbonyl group, as defined herein (e.g., -alkynyl-C(O)--OR, where R is an optionally substituted C.sub.1-20, C.sub.1-10, or C.sub.1-6 alkyl group). Exemplary unsubstituted alkoxycarbonylalkynyl include from 4 to 41 carbons (e.g., from 4 to 10, from 4 to 13, from 4 to 17, from 4 to 21, or from 4 to 31 carbons, such as C.sub.1-6 alkoxycarbonyl-C.sub.2-6 alkynyl, C.sub.1-10 alkoxycarbonyl-C.sub.2-10 alkynyl, or C.sub.1-20 alkoxycarbonyl-C.sub.2-20 alkynyl). In some embodiments, each alkyl, alkynyl, and alkoxy group is further independently substituted with 1, 2, 3, or 4 substituents as described herein (e.g., a hydroxyl group).
The "aminoalkynyl" group, which as used herein, represents an alkynyl group, as defined herein, substituted with an amino group, as defined herein. The alkynyl and amino each can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for the respective group (e.g., CO.sub.2R.sup.A', where R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl, e.g., carboxy, and/or an N-protecting group).
The "hydroxyalkynyl" group, which as used herein, represents an alkynyl group, as defined herein, substituted with one to three hydroxyl groups, with the proviso that no more than one hydroxyl group may be attached to a single carbon atom of the alkyl group. In some embodiments, the hydroxyalkynyl group can be substituted with 1, 2, 3, or 4 substituent groups (e.g., O-protecting groups) as defined herein for an alkyl.
The term "amino," as used herein, represents --N(R.sup.N1).sub.2, wherein each R.sup.N1 is, independently, H, OH, NO.sub.2, N(R.sup.N2).sub.2, SO.sub.2OR.sup.N2, SO.sub.2R.sup.N2, SOR.sup.N2, an N-protecting group, alkyl, alkenyl, alkynyl, alkoxy, aryl, alkaryl, cycloalkyl, alkcycloalkyl, carboxyalkyl (e.g., optionally substituted with an O-protecting group, such as optionally substituted arylalkoxycarbonyl groups or any described herein), sulfoalkyl, acyl (e.g., acetyl, trifluoroacetyl, or others described herein), alkoxycarbonylalkyl (e.g., optionally substituted with an O-protecting group, such as optionally substituted arylalkoxycarbonyl groups or any described herein), heterocyclyl (e.g., heteroaryl), or alkheterocyclyl (e.g., alkheteroaryl), wherein each of these recited R.sup.N1 groups can be optionally substituted, as defined herein for each group; or two R.sup.N1 combine to form a heterocyclyl or an N-protecting group, and wherein each R.sup.N2 is, independently, H, alkyl, or aryl. The amino groups of the invention can be an unsubstituted amino (i.e., --NH.sub.2) or a substituted amino (i.e., --N(R.sup.N1).sub.2). In a preferred embodiment, amino is --NH.sub.2 or --NHR.sup.N1, wherein R.sup.N1 is, independently, OH, NO.sub.2, NH.sub.2, NR.sup.N2.sub.2, SO.sub.2OR.sup.N2, SO.sub.2R.sup.N2, SOR.sup.N2, alkyl, carboxyalkyl, sulfoalkyl, acyl (e.g., acetyl, trifluoroacetyl, or others described herein), alkoxycarbonylalkyl (e.g., t-butoxycarbonylalkyl) or aryl, and each R.sup.N2 can be H, C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), or C.sub.6-10 aryl.
Non-limiting examples of optionally substituted amino groups include acylamino and carbamyl:
The "acylamino" group, which as used herein, represents an acyl group, as defined herein, attached to the parent molecular group though an amino group, as defined herein (i.e., --N(R.sup.N1)--C(O)--R, where R is H or an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group (e.g., haloalkyl) and R.sup.N1 is as defined herein). Exemplary unsubstituted acylamino groups include from 1 to 41 carbons (e.g., from 1 to 7, from 1 to 13, from 1 to 21, from 2 to 7, from 2 to 13, from 2 to 21, or from 2 to 41 carbons). In some embodiments, the alkyl group is further substituted with 1, 2, 3, or 4 substituents as described herein, and/or the amino group is --NH.sub.2 or --NHR.sup.N1, wherein R.sup.N1 is, independently, OH, NO.sub.2, NH.sub.2, NR.sup.N2.sub.2, SO.sub.2OR.sup.N2, SO.sub.2R.sup.N2, SOR.sup.N2, alkyl, aryl, acyl (e.g., acetyl, trifluoroacetyl, or others described herein), or alkoxycarbonylalkyl, and each R.sup.N2 can be H, alkyl, or aryl.
The "carbamyl" group, which as used herein, refers to a carbamate group having the structure --NR.sup.N1C(.dbd.O)OR or --OC(.dbd.O)N(R.sup.N1).sub.2, where the meaning of each R.sup.N1 is found in the definition of "amino" provided herein, and R is alkyl, cycloalkyl, alkcycloalkyl, aryl, alkaryl, heterocyclyl (e.g., heteroaryl), or alkheterocyclyl (e.g., alkheteroaryl), as defined herein.
The term "amino acid," as described herein, refers to a molecule having a side chain, an amino group, and an acid group (e.g., a carboxy group of --CO.sub.2H or a sulfo group of --SO.sub.3H), wherein the amino acid is attached to the parent molecular group by the side chain, amino group, or acid group (e.g., the side chain). In some embodiments, the amino acid is attached to the parent molecular group by a carbonyl group, where the side chain or amino group is attached to the carbonyl group. Exemplary side chains include an optionally substituted alkyl, aryl, heterocyclyl, alkaryl, alkheterocyclyl, aminoalkyl, carbamoylalkyl, and carboxyalkyl. Exemplary amino acids include alanine, arginine, asparagine, aspartic acid, cysteine, glutamic acid, glutamine, glycine, histidine, hydroxynorvaline, isoleucine, leucine, lysine, methionine, norvaline, ornithine, phenylalanine, proline, pyrrolysine, selenocysteine, serine, taurine, threonine, tryptophan, tyrosine, and valine. Amino acid groups may be optionally substituted with one, two, three, or, in the case of amino acid groups of two carbons or more, four substituents independently selected from the group consisting of: (1) C.sub.1-6 alkoxy; (2) C.sub.1-6 alkylsulfinyl; (3) amino, as defined herein (e.g., unsubstituted amino (i.e., --NH.sub.2) or a substituted amino (i.e., --N(R.sup.N1).sub.2, where R.sup.N1 is as defined for amino); (4) C.sub.6-10 aryl-C.sub.1-6 alkoxy; (5) azido; (6) halo; (7) (C.sub.2-9 heterocyclyl)oxy; (8) hydroxyl; (9) nitro; (10) oxo (e.g., carboxyaldehyde or acyl); (11) C.sub.1-7 spirocyclyl; (12) thioalkoxy; (13) thiol; (14) --CO.sub.2R.sup.A', where R.sup.A' is selected from the group consisting of (a) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c) C.sub.6-10 aryl, (d) hydrogen, (e) C.sub.1-6 alk-C.sub.6-10 aryl, (f) amino-C.sub.1-20 alkyl, (g) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1(CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; (15) --C(O)NR.sup.B'R.sup.C', where each of R.sup.B' and R.sup.C' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (16) --SO.sub.2R.sup.D', where R.sup.D' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) C.sub.1-6 alk-C.sub.6-10 aryl, and (d) hydroxyl; (17) --SO.sub.2NR.sup.E'R.sup.F', where each of R.sup.E' and R.sup.F' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (18) --C(O)R.sup.G', where R.sup.G' is selected from the group consisting of (a) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c) C.sub.6-10 aryl, (d) hydrogen, (e) C.sub.1-6 alk-C.sub.6-10 aryl, (f) amino-C.sub.1-20 alkyl, (g) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1 (CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; (19) --NR.sup.H'C(O)R.sup.I', wherein R.sup.H' is selected from the group consisting of (a1) hydrogen and (b1) C.sub.1-6 alkyl, and R.sup.I' is selected from the group consisting of (a2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b2) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c2) C.sub.6-10 aryl, (d2) hydrogen, (e2) C.sub.1-6 alk-C.sub.6-10 aryl, (f2) amino-C.sub.1-20 alkyl, (g2) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1 (CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h2) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; (20) --NR.sup.J'C(O)OR.sup.K', wherein R.sup.J' is selected from the group consisting of (a1) hydrogen and (b1) C.sub.1-6 alkyl, and R.sup.K' is selected from the group consisting of (a2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl), (b2) C.sub.2-20 alkenyl (e.g., C.sub.2-6 alkenyl), (c2) C.sub.6-10 aryl, (d2) hydrogen, (e2) C.sub.1-6 alk-C.sub.6-10 aryl, (f2) amino-C.sub.1-20 alkyl, (g2) polyethylene glycol of --(CH.sub.2).sub.s2(OCH.sub.2CH.sub.2).sub.s1 (CH.sub.2).sub.s3OR', wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and R' is H or C.sub.1-20 alkyl, and (h2) amino-polyethylene glycol of --NR.sup.N1(CH.sub.2).sub.s2(CH.sub.2CH.sub.2O).sub.s1(CH.sub.2).sub.s3NR- .sup.N1, wherein s1 is an integer from 1 to 10 (e.g., from 1 to 6 or from 1 to 4), each of s2 and s3, independently, is an integer from 0 to 10 (e.g., from 0 to 4, from 0 to 6, from 1 to 4, from 1 to 6, or from 1 to 10), and each R.sup.N1 is, independently, hydrogen or optionally substituted C.sub.1-6 alkyl; and (21) amidine. In some embodiments, each of these groups can be further substituted as described herein.
The term "aryl," as used herein, represents a mono-, bicyclic, or multicyclic carbocyclic ring system having one or two aromatic rings and is exemplified by phenyl, naphthyl, 1,2-dihydronaphthyl, 1,2,3,4-tetrahydronaphthyl, anthracenyl, phenanthrenyl, fluorenyl, indanyl, indenyl, and the like, and may be optionally substituted with 1, 2, 3, 4, or 5 substituents independently selected from the group consisting of: (1) C.sub.1-7 acyl (e.g., carboxyaldehyde); (2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl, C.sub.1-6 alkoxy-C.sub.1-6 alkyl, C.sub.1-6 alkylsulfinyl-C.sub.1-6 alkyl, amino-C.sub.1-6 alkyl, azido-C.sub.1-6 alkyl, (carboxyaldehyde)-C.sub.1-6 alkyl, halo-C.sub.1-6 alkyl (e.g., perfluoroalkyl), hydroxy-C.sub.1-6 alkyl, nitro-C.sub.1-6 alkyl, or C.sub.1-6 thioalkoxy-C.sub.1-6 alkyl); (3) C.sub.1-20 alkoxy (e.g., C.sub.1-6 alkoxy, such as perfluoroalkoxy); (4) C.sub.1-6 alkylsulfinyl; (5) C.sub.6-10 aryl; (6) amino; (7) C.sub.1-6 alk-C.sub.6-10 aryl; (8) azido; (9) C.sub.3-8 cycloalkyl; (10) C.sub.1-6 alk-C.sub.3-8 cycloalkyl; (11) halo; (12) C.sub.1-12 heterocyclyl (e.g., C.sub.1-12 heteroaryl); (13) (C.sub.1-12 heterocyclyl)oxy; (14) hydroxyl; (15) nitro; (16) C.sub.1-20 thioalkoxy (e.g., C.sub.1-6 thioalkoxy); (17) --(CH.sub.2).sub.qCO.sub.2R.sup.A', where q is an integer from zero to four, and R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (18) --(CH.sub.2).sub.qCONR.sup.B'R.sup.C', where q is an integer from zero to four and where R.sup.B' and R.sup.C' are independently selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (19) --(CH.sub.2).sub.qSO.sub.2R.sup.D', where q is an integer from zero to four and where R.sup.D' is selected from the group consisting of (a) alkyl, (b) C.sub.6-10 aryl, and (c) alk-C.sub.6-10 aryl; (20) --(CH.sub.2).sub.qSO.sub.2NR.sup.E'R.sup.F', where q is an integer from zero to four and where each of R.sup.E' and R.sup.F' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (21) thiol; (22) C.sub.6-10 aryloxy; (23) C.sub.3-8 cycloalkoxy; (24) C.sub.6-10 aryl-C.sub.1-6 alkoxy; (25) C.sub.1-6 alk-C.sub.1-12 heterocyclyl (e.g., C.sub.1-6 alk-C.sub.1-12 heteroaryl); (26) C.sub.2-20 alkenyl; and (27) C.sub.2-20 alkynyl. In some embodiments, each of these groups can be further substituted as described herein. For example, the alkylene group of a C.sub.1-alkaryl or a C.sub.1-alkheterocyclyl can be further substituted with an oxo group to afford the respective aryloyl and (heterocyclyl)oyl substituent group.
The "arylalkyl" group, which as used herein, represents an aryl group, as defined herein, attached to the parent molecular group through an alkylene group, as defined herein. Exemplary unsubstituted arylalkyl groups are from 7 to 30 carbons (e.g., from 7 to 16 or from 7 to 20 carbons, such as C.sub.1-6 alk-C.sub.6-10 aryl, C.sub.1-10 alk-C.sub.6-10 aryl, or C.sub.1-20 alk-C.sub.6-10 aryl). In some embodiments, the alkylene and the aryl each can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein for the respective groups. Other groups preceded by the prefix "alk-" are defined in the same manner, where "alk" refers to a C.sub.1-6 alkylene, unless otherwise noted, and the attached chemical structure is as defined herein.
The term "azido" represents an --N.sub.3 group, which can also be represented as --N.dbd.N.dbd.N.
The term "bicyclic," as used herein, refer to a structure having two rings, which may be aromatic or non-aromatic. Bicyclic structures include spirocyclyl groups, as defined herein, and two rings that share one or more bridges, where such bridges can include one atom or a chain including two, three, or more atoms. Exemplary bicyclic groups include a bicyclic carbocyclyl group, where the first and second rings are carbocyclyl groups, as defined herein; a bicyclic aryl groups, where the first and second rings are aryl groups, as defined herein; bicyclic heterocyclyl groups, where the first ring is a heterocyclyl group and the second ring is a carbocyclyl (e.g., aryl) or heterocyclyl (e.g., heteroaryl) group; and bicyclic heteroaryl groups, where the first ring is a heteroaryl group and the second ring is a carbocyclyl (e.g., aryl) or heterocyclyl (e.g., heteroaryl) group. In some embodiments, the bicyclic group can be substituted with 1, 2, 3, or 4 substituents as defined herein for cycloalkyl, heterocyclyl, and aryl groups.
The term "boranyl," as used herein, represents --B(R.sup.B1).sub.3, where each R.sup.B1 is, independently, selected from the group consisting of H and optionally substituted alkyl. In some embodiments, the boranyl group can be substituted with 1, 2, 3, or 4 substituents as defined herein for alkyl.
The terms "carbocyclic" and "carbocyclyl," as used herein, refer to an optionally substituted C.sub.3-12 monocyclic, bicyclic, or tricyclic structure in which the rings, which may be aromatic or non-aromatic, are formed by carbon atoms. Carbocyclic structures include cycloalkyl, cycloalkenyl, and aryl groups.
The term "carbonyl," as used herein, represents a C(O) group, which can also be represented as C.dbd.O.
The term "carboxy," as used herein, means --CO.sub.2H.
The term "cyano," as used herein, represents an --CN group.
The term "cycloalkyl," as used herein represents a monovalent saturated or unsaturated non-aromatic cyclic hydrocarbon group from three to eight carbons, unless otherwise specified, and is exemplified by cyclopropyl, cyclobutyl, cyclopentyl, cyclohexyl, cycloheptyl, bicycle heptyl, and the like. When the cycloalkyl group includes one carbon-carbon double bond, the cycloalkyl group can be referred to as a "cycloalkenyl" group. Exemplary cycloalkenyl groups include cyclopentenyl, cyclohexenyl, and the like. The cycloalkyl groups of this invention can be optionally substituted with: (1) C.sub.1-7 acyl (e.g., carboxyaldehyde); (2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl, C.sub.1-6 alkoxy-C.sub.1-6 alkyl, C.sub.1-6 alkylsulfinyl-C.sub.1-6 alkyl, amino-C.sub.1-6 alkyl, azido-C.sub.1-6 alkyl, (carboxyaldehyde)-C.sub.1-6 alkyl, halo-C.sub.1-6 alkyl (e.g., perfluoroalkyl), hydroxy-C.sub.1-6 alkyl, nitro-C.sub.1-6 alkyl, or C.sub.1-6 thioalkoxy-C.sub.1-6 alkyl); (3) C.sub.1-20 alkoxy (e.g., C.sub.1-6 alkoxy, such as perfluoroalkoxy); (4) C.sub.1-6 alkylsulfinyl; (5) C.sub.6-10 aryl; (6) amino; (7) C.sub.1-6 alk-C.sub.6-10 aryl; (8) azido; (9) C.sub.3-8 cycloalkyl; (10) C.sub.1-6 alk-C.sub.3-8 cycloalkyl; (11) halo; (12) C.sub.1-12 heterocyclyl (e.g., C.sub.1-12 heteroaryl); (13) (C.sub.1-12 heterocyclyl)oxy; (14) hydroxyl; (15) nitro; (16) C.sub.1-20 thioalkoxy (e.g., C.sub.1-6 thioalkoxy); (17) --(CH.sub.2).sub.qCO.sub.2R.sup.A', where q is an integer from zero to four, and R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (18) --(CH.sub.2).sub.qCONR.sup.B'R.sup.C', where q is an integer from zero to four and where R.sup.B' and R.sup.C' are independently selected from the group consisting of (a) hydrogen, (b) C.sub.6-10 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (19) --(CH.sub.2).sub.qSO.sub.2R.sup.D', where q is an integer from zero to four and where R.sup.D' is selected from the group consisting of (a) C.sub.6-10 alkyl, (b) C.sub.6-10 aryl, and (c) C.sub.1-6 alk-C.sub.6-10 aryl; (20) --(CH.sub.2).sub.qSO.sub.2NR.sup.E'R.sup.F', where q is an integer from zero to four and where each of R.sup.E' and R.sup.F' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.6-10 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (21) thiol; (22) C.sub.6-10 aryloxy; (23) cycloalkoxy; (24) C.sub.6-10 aryl-C.sub.1-6 alkoxy; (25) C.sub.1-6 alk-C.sub.1-12 heterocyclyl (e.g., C.sub.1-6 alk-C.sub.1-12 heteroaryl); (26) oxo; (27) C.sub.2-20 alkenyl; and (28) C.sub.2-20 alkynyl. In some embodiments, each of these groups can be further substituted as described herein. For example, the alkylene group of a C.sub.1-alkaryl or a C.sub.1-alkheterocyclyl can be further substituted with an oxo group to afford the respective aryloyl and (heterocyclyl)oyl substituent group.
The "cycloalkylalkyl" group, which as used herein, represents a cycloalkyl group, as defined herein, attached to the parent molecular group through an alkylene group, as defined herein (e.g., an alkylene group of from 1 to 4, from 1 to 6, from 1 to 10, or form 1 to 20 carbons). In some embodiments, the alkylene and the cycloalkyl each can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein for the respective group.
The term "diastereomer," as used herein means stereoisomers that are not mirror images of one another and are non-superimposable on one another.
The term "enantiomer," as used herein, means each individual optically active form of a compound of the invention, having an optical purity or enantiomeric excess (as determined by methods standard in the art) of at least 80% (i.e., at least 90% of one enantiomer and at most 10% of the other enantiomer), preferably at least 90% and more preferably at least 98%.
The term "halo," as used herein, represents a halogen selected from bromine, chlorine, iodine, or fluorine.
The term "heteroalkyl," as used herein, refers to an alkyl group, as defined herein, in which one or two of the constituent carbon atoms have each been replaced by nitrogen, oxygen, or sulfur. In some embodiments, the heteroalkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for alkyl groups. The terms "heteroalkenyl" and heteroalkynyl," as used herein refer to alkenyl and alkynyl groups, as defined herein, respectively, in which one or two of the constituent carbon atoms have each been replaced by nitrogen, oxygen, or sulfur. In some embodiments, the heteroalkenyl and heteroalkynyl groups can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for alkyl groups.
Non-limiting examples of optionally substituted heteroalkyl, heteroalkenyl, and heteroalkynyl groups include acyloxy, alkenyloxy, alkoxy, alkoxyalkoxy, alkoxycarbonylalkoxy, alkynyloxy, aminoalkoxy, arylalkoxy, carboxyalkoxy, cycloalkoxy, haloalkoxy, (heterocyclyl)oxy, perfluoroalkoxy, thioalkoxy, and thioheterocyclylalkyl:
The "acyloxy" group, which as used herein, represents an acyl group, as defined herein, attached to the parent molecular group though an oxygen atom (i.e., --O--C(O)--R, where R is H or an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group). Exemplary unsubstituted acyloxy groups include from 1 to 21 carbons (e.g., from 1 to 7 or from 1 to 11 carbons). In some embodiments, the alkyl group is further substituted with 1, 2, 3, or 4 substituents as described herein.
The "alkenyloxy" group, which as used here, represents a chemical substituent of formula --OR, where R is a C.sub.2-20 alkenyl group (e.g., C.sub.2-6 or C.sub.2-10 alkenyl), unless otherwise specified. Exemplary alkenyloxy groups include ethenyloxy, propenyloxy, and the like. In some embodiments, the alkenyl group can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein (e.g., a hydroxyl group).
The "alkoxy" group, which as used herein, represents a chemical substituent of formula --OR, where R is a C.sub.1-20 alkyl group (e.g., C.sub.1-6 or C.sub.1-10 alkyl), unless otherwise specified. Exemplary alkoxy groups include methoxy, ethoxy, propoxy (e.g., n-propoxy and isopropoxy), t-butoxy, and the like. In some embodiments, the alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein (e.g., hydroxyl or alkoxy).
The "alkoxyalkoxy" group, which as used herein, represents an alkoxy group that is substituted with an alkoxy group. Exemplary unsubstituted alkoxyalkoxy groups include between 2 to 40 carbons (e.g., from 2 to 12 or from 2 to 20 carbons, such as C.sub.1-6 alkoxy-C.sub.1-6 alkoxy, C.sub.1-10 alkoxy-C.sub.1-10 alkoxy, or C.sub.1-20 alkoxy-C.sub.1-20 alkoxy). In some embodiments, the each alkoxy group can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein.
The "alkoxycarbonylalkoxy" group, which as used herein, represents an alkoxy group, as defined herein, that is substituted with an alkoxycarbonyl group, as defined herein (e.g., --O-alkyl-C(O)--OR, where R is an optionally substituted C.sub.1-6, C.sub.1-10, or C.sub.1-20 alkyl group). Exemplary unsubstituted alkoxycarbonylalkoxy include from 3 to 41 carbons (e.g., from 3 to 10, from 3 to 13, from 3 to 17, from 3 to 21, or from 3 to 31 carbons, such as C.sub.1-6 alkoxycarbonyl-C.sub.1-6 alkoxy, C.sub.1-10 alkoxycarbonyl-C.sub.1-10 alkoxy, or C.sub.1-20 alkoxycarbonyl-C.sub.1-20 alkoxy). In some embodiments, each alkoxy group is further independently substituted with 1, 2, 3, or 4 substituents, as described herein (e.g., a hydroxyl group).
The "alkynyloxy" group, which as used herein, represents a chemical substituent of formula --OR, where R is a C.sub.2-20 alkynyl group (e.g., C.sub.2-6 or C.sub.2-10 alkynyl), unless otherwise specified. Exemplary alkynyloxy groups include ethynyloxy, propynyloxy, and the like. In some embodiments, the alkynyl group can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein (e.g., a hydroxyl group).
The "aminoalkoxy" group, which as used herein, represents an alkoxy group, as defined herein, substituted with an amino group, as defined herein. The alkyl and amino each can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for the respective group (e.g., CO.sub.2R.sup.A', where R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl, e.g., carboxy).
The "arylalkoxy" group, which as used herein, represents an alkaryl group, as defined herein, attached to the parent molecular group through an oxygen atom. Exemplary unsubstituted arylalkoxy groups include from 7 to 30 carbons (e.g., from 7 to 16 or from 7 to 20 carbons, such as C.sub.6-10 aryl-C.sub.1-6 alkoxy, C.sub.6-10 aryl-C.sub.1-10 alkoxy, or C.sub.6-10 aryl-C.sub.1-20 alkoxy). In some embodiments, the arylalkoxy group can be substituted with 1, 2, 3, or 4 substituents as defined herein.
The "aryloxy" group, which as used herein, represents a chemical substituent of formula --OR', where R' is an aryl group of 6 to 18 carbons, unless otherwise specified. In some embodiments, the aryl group can be substituted with 1, 2, 3, or 4 substituents as defined herein.
The "carboxyalkoxy" group, which as used herein, represents an alkoxy group, as defined herein, substituted with a carboxy group, as defined herein. The alkoxy group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for the alkyl group, and the carboxy group can be optionally substituted with one or more O-protecting groups.
The "cycloalkoxy" group, which as used herein, represents a chemical substituent of formula --OR, where R is a C.sub.3-8 cycloalkyl group, as defined herein, unless otherwise specified. The cycloalkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein. Exemplary unsubstituted cycloalkoxy groups are from 3 to 8 carbons. In some embodiment, the cycloalkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein.
The "haloalkoxy" group, which as used herein, represents an alkoxy group, as defined herein, substituted with a halogen group (i.e., F, Cl, Br, or I). A haloalkoxy may be substituted with one, two, three, or, in the case of alkyl groups of two carbons or more, four halogens. Haloalkoxy groups include perfluoroalkoxys (e.g., --OCF.sub.3), --OCHF.sub.2, --OCH.sub.2F, --OCCl.sub.3, --OCH.sub.2CH.sub.2Br, --OCH.sub.2CH(CH.sub.2CH.sub.2Br)CH.sub.3, and --OCHICH.sub.3. In some embodiments, the haloalkoxy group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein for alkyl groups.
The "(heterocyclyl)oxy" group, which as used herein, represents a heterocyclyl group, as defined herein, attached to the parent molecular group through an oxygen atom. In some embodiments, the heterocyclyl group can be substituted with 1, 2, 3, or 4 substituent groups as defined herein.
The "perfluoroalkoxy" group, which as used herein, represents an alkoxy group, as defined herein, where each hydrogen radical bound to the alkoxy group has been replaced by a fluoride radical. Perfluoroalkoxy groups are exemplified by trifluoromethoxy, pentafluoroethoxy, and the like.
The "alkylsulfinyl" group, which as used herein, represents an alkyl group attached to the parent molecular group through an --S(O)-- group. Exemplary unsubstituted alkylsulfinyl groups are from 1 to 6, from 1 to 10, or from 1 to 20 carbons. In some embodiments, the alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein.
The "thioarylalkyl" group, which as used herein, represents a chemical substituent of formula --SR, where R is an arylalkyl group. In some embodiments, the arylalkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein.
The "thioalkoxy" group as used herein, represents a chemical substituent of formula --SR, where R is an alkyl group, as defined herein. In some embodiments, the alkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein.
The "thioheterocyclylalkyl" group, which as used herein, represents a chemical substituent of formula --SR, where R is an heterocyclylalkyl group. In some embodiments, the heterocyclylalkyl group can be further substituted with 1, 2, 3, or 4 substituent groups as described herein.
The term "heteroaryl," as used herein, represents that subset of heterocyclyls, as defined herein, which are aromatic: i.e., they contain 4n+2 pi electrons within the mono- or multicyclic ring system. Exemplary unsubstituted heteroaryl groups are of 1 to 12 (e.g., 1 to 11, 1 to 10, 1 to 9, 2 to 12, 2 to 11, 2 to 10, or 2 to 9) carbons. In some embodiment, the heteroaryl is substituted with 1, 2, 3, or 4 substituents groups as defined for a heterocyclyl group.
The term "heteroarylalkyl" refers to a heteroaryl group, as defined herein, attached to the parent molecular group through an alkylene group, as defined herein. Exemplary unsubstituted heteroarylalkyl groups are from 2 to 32 carbons (e.g., from 2 to 22, from 2 to 18, from 2 to 17, from 2 to 16, from 3 to 15, from 2 to 14, from 2 to 13, or from 2 to 12 carbons, such as C.sub.1-6 alk-C.sub.1-12 heteroaryl, C.sub.1-10 alk-C.sub.1-12 heteroaryl, or C.sub.1-20 alk-C.sub.1-12 heteroaryl). In some embodiments, the alkylene and the heteroaryl each can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein for the respective group. Heteroarylalkyl groups are a subset of heterocyclylalkyl groups.
The term "heterocyclyl," as used herein represents a 5-, 6- or 7-membered ring, unless otherwise specified, containing one, two, three, or four heteroatoms independently selected from the group consisting of nitrogen, oxygen, and sulfur. The 5-membered ring has zero to two double bonds, and the 6- and 7-membered rings have zero to three double bonds. Exemplary unsubstituted heterocyclyl groups are of 1 to 12 (e.g., 1 to 11, 1 to 10, 1 to 9, 2 to 12, 2 to 11, 2 to 10, or 2 to 9) carbons. The term "heterocyclyl" also represents a heterocyclic compound having a bridged multicyclic structure in which one or more carbons and/or heteroatoms bridges two non-adjacent members of a monocyclic ring, e.g., a quinuclidinyl group. The term "heterocyclyl" includes bicyclic, tricyclic, and tetracyclic groups in which any of the above heterocyclic rings is fused to one, two, or three carbocyclic rings, e.g., an aryl ring, a cyclohexane ring, a cyclohexene ring, a cyclopentane ring, a cyclopentene ring, or another monocyclic heterocyclic ring, such as indolyl, quinolyl, isoquinolyl, tetrahydroquinolyl, benzofuryl, benzothienyl and the like. Examples of fused heterocyclyls include tropanes and 1,2,3,5,8,8a-hexahydroindolizine. Heterocyclics include pyrrolyl, pyrrolinyl, pyrrolidinyl, pyrazolyl, pyrazolinyl, pyrazolidinyl, imidazolyl, imidazolinyl, imidazolidinyl, pyridyl, piperidinyl, homopiperidinyl, pyrazinyl, piperazinyl, pyrimidinyl, pyridazinyl, oxazolyl, oxazolidinyl, isoxazolyl, isoxazolidiniyl, morpholinyl, thiomorpholinyl, thiazolyl, thiazolidinyl, isothiazolyl, isothiazolidinyl, indolyl, indazolyl, quinolyl, isoquinolyl, quinoxalinyl, dihydroquinoxalinyl, quinazolinyl, cinnolinyl, phthalazinyl, benzimidazolyl, benzothiazolyl, benzoxazolyl, benzothiadiazolyl, furyl, thienyl, thiazolidinyl, isothiazolyl, triazolyl, tetrazolyl, oxadiazolyl (e.g., 1,2,3-oxadiazolyl), purinyl, thiadiazolyl (e.g., 1,2,3-thiadiazolyl), tetrahydrofuranyl, dihydrofuranyl, tetrahydrothienyl, dihydrothienyl, dihydroindolyl, dihydroquinolyl, tetrahydroquinolyl, tetrahydroisoquinolyl, dihydroisoquinolyl, pyranyl, dihydropyranyl, dithiazolyl, benzofuranyl, isobenzofuranyl, benzothienyl, and the like, including dihydro and tetrahydro forms thereof, where one or more double bonds are reduced and replaced with hydrogens. Still other exemplary heterocyclyls include: 2,3,4,5-tetrahydro-2-oxo-oxazolyl; 2,3-dihydro-2-oxo-1H-imidazolyl; 2,3,4,5-tetrahydro-5-oxo-1H-pyrazolyl (e.g., 2,3,4,5-tetrahydro-2-phenyl-5-oxo-1H-pyrazolyl); 2,3,4,5-tetrahydro-2,4-dioxo-1H-imidazolyl (e.g., 2,3,4,5-tetrahydro-2,4-dioxo-5-methyl-5-phenyl-1H-imidazolyl); 2,3-dihydro-2-thioxo-1,3,4-oxadiazolyl(e.g., 2,3-dihydro-2-thioxo-5-phenyl-1,3,4-oxadiazolyl); 4,5-dihydro-5-oxo-1H-triazolyl (e.g., 4,5-dihydro-3-methyl-4-amino 5-oxo-1H-triazolyl); 1,2,3,4-tetrahydro-2,4-dioxopyridinyl (e.g., 1,2,3,4-tetrahydro-2,4-dioxo-3,3-diethylpyridinyl); 2,6-dioxo-piperidinyl (e.g., 2,6-dioxo-3-ethyl-3-phenylpiperidinyl); 1,6-dihydro-6-oxopyridiminyl; 1,6-dihydro-4-oxopyrimidinyl (e.g., 2-(methylthio)-1,6-dihydro-4-oxo-5-methylpyrimidin-1-yl); 1,2,3,4-tetrahydro-2,4-dioxopyrimidinyl (e.g., 1,2,3,4-tetrahydro-2,4-dioxo-3-ethylpyrimidinyl); 1,6-dihydro-6-oxo-pyridazinyl (e.g., 1,6-dihydro-6-oxo-3-ethylpyridazinyl); 1,6-dihydro-6-oxo-1,2,4-triazinyl (e.g., 1,6-dihydro-5-isopropyl-6-oxo-1,2,4-triazinyl); 2,3-dihydro-2-oxo-1H-indolyl (e.g., 3,3-dimethyl-2,3-dihydro-2-oxo-1H-indolyl and 2,3-dihydro-2-oxo-3,3'-spiropropane-1H-indol-1-yl); 1,3-dihydro-1-oxo-2H-iso-indolyl; 1,3-dihydro-1,3-dioxo-2H-iso-indolyl; 1H-benzopyrazolyl (e.g., 1-(ethoxycarbonyl)-1H-benzopyrazolyl); 2,3-dihydro-2-oxo-1H-benzimidazolyl (e.g., 3-ethyl-2,3-dihydro-2-oxo-1H-benzimidazolyl); 2,3-dihydro-2-oxo-benzoxazolyl (e.g., 5-chloro-2,3-dihydro-2-oxo-benzoxazolyl); 2,3-dihydro-2-oxo-benzoxazolyl; 2-oxo-2H-benzopyranyl; 1,4-benzodioxanyl; 1,3-benzodioxanyl; 2,3-dihydro-3-oxo, 4H-1,3-benzothiazinyl; 3,4-dihydro-4-oxo-3H-quinazolinyl (e.g., 2-methyl-3,4-dihydro-4-oxo-3H-quinazolinyl); 1,2,3,4-tetrahydro-2,4-dioxo-3H-quinazolyl (e.g., 1-ethyl-1,2,3,4-tetrahydro-2,4-dioxo-3H-quinazolyl); 1,2,3,6-tetrahydro-2,6-dioxo-7H-purinyl (e.g., 1,2,3,6-tetrahydro-1,3-dimethyl-2,6-dioxo-7H-purinyl); 1,2,3,6-tetrahydro-2,6-dioxo-1H-purinyl (e.g., 1,2,3,6-tetrahydro-3,7-dimethyl-2,6-dioxo-1H-purinyl); 2-oxobenz[c,d]indolyl; 1,1-dioxo-2H-naphth[1,8-c,d]isothiazolyl; and 1,8-naphthylenedicarboxamido. Additional heterocyclics include 3,3a,4,5,6,6a-hexahydro-pyrrolo[3,4-b]pyrrol-(2H)-yl, and 2,5-diazabicyclo[2.2.1]heptan-2-yl, homopiperazinyl (or diazepanyl), tetrahydropyranyl, dithiazolyl, benzofuranyl, benzothienyl, oxepanyl, thiepanyl, azocanyl, oxecanyl, and thiocanyl. Heterocyclic groups also include groups of the formula
##STR00037## where
E' is selected from the group consisting of --N-- and --CH--; F' is selected from the group consisting of --N.dbd.CH--, --NH--CH.sub.2--, --NH--C(O)--, --NH--, --CH.dbd.N--, --CH.sub.2--NH--, --C(O)--NH--, --CH.dbd.CH--, --CH.sub.2--, --CH.sub.2CH.sub.2--, --CH.sub.2O--, --OCH.sub.2--, --O--, and --S--; and G' is selected from the group consisting of --CH-- and --N--. Any of the heterocyclyl groups mentioned herein may be optionally substituted with one, two, three, four or five substituents independently selected from the group consisting of: (1) C.sub.1-7 acyl (e.g., carboxyaldehyde); (2) C.sub.1-20 alkyl (e.g., C.sub.1-6 alkyl, C.sub.1-6 alkoxy-C.sub.1-6 alkyl, C.sub.1-6 alkylsulfinyl-C.sub.1-6 alkyl, amino-C.sub.1-6 alkyl, azido-C.sub.1-6 alkyl, (carboxyaldehyde)-C.sub.1-6 alkyl, halo-C.sub.1-6 alkyl (e.g., perfluoroalkyl), hydroxy-C.sub.1-6 alkyl, nitro-C.sub.1-6 alkyl, or C.sub.1-6 thioalkoxy-C.sub.1-6 alkyl); (3) C.sub.1-20 alkoxy (e.g., C.sub.1-6 alkoxy, such as perfluoroalkoxy); (4) C.sub.1-6 alkylsulfinyl; (5) C.sub.6-10 aryl; (6) amino; (7) C.sub.1-6 alk-C.sub.6-10 aryl; (8) azido; (9) C.sub.3-8 cycloalkyl; (10) C.sub.1-6 alk-C.sub.3-8 cycloalkyl; (11) halo; (12) C.sub.1-12 heterocyclyl (e.g., C.sub.2-12 heteroaryl); (13) (C.sub.1-12 heterocyclyl)oxy; (14) hydroxyl; (15) nitro; (16) C.sub.1-20 thioalkoxy (e.g., C.sub.1-6 thioalkoxy); (17) --(CH.sub.2).sub.qCO.sub.2R.sup.A', where q is an integer from zero to four, and R.sup.A' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, (c) hydrogen, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (18) --(CH.sub.2).sub.qCONR.sup.B'R.sup.C', where q is an integer from zero to four and where R.sup.B' and R.sup.C' are independently selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (19) --(CH.sub.2).sub.qSO.sub.2R.sup.D', where q is an integer from zero to four and where R.sup.D' is selected from the group consisting of (a) C.sub.1-6 alkyl, (b) C.sub.6-10 aryl, and (c) C.sub.1-6 alk-C.sub.6-10 aryl; (20) --(CH.sub.2).sub.qSO.sub.2NR.sup.E'R.sup.F', where q is an integer from zero to four and where each of R.sup.E' and R.sup.F' is, independently, selected from the group consisting of (a) hydrogen, (b) C.sub.1-6 alkyl, (c) C.sub.6-10 aryl, and (d) C.sub.1-6 alk-C.sub.6-10 aryl; (21) thiol; (22) C.sub.6-10 aryloxy; (23) C.sub.3-8 cycloalkoxy; (24) arylalkoxy; (25) C.sub.1-6 alk-C.sub.1-12 heterocyclyl (e.g., C.sub.1-6 alk-C.sub.1-12 heteroaryl); (26) oxo; (27) (C.sub.1-12 heterocyclyl)imino; (28) C.sub.2-20 alkenyl; and (29) C.sub.2-20 alkynyl. In some embodiments, each of these groups can be further substituted as described herein. For example, the alkylene group of a C.sub.1-alkaryl or a C.sub.1-alkheterocyclyl can be further substituted with an oxo group to afford the respective aryloyl and (heterocyclyl)oyl substituent group.
The "heterocyclylalkyl" group, which as used herein, represents a heterocyclyl group, as defined herein, attached to the parent molecular group through an alkylene group, as defined herein. Exemplary unsubstituted heterocyclylalkyl groups are from 2 to 32 carbons (e.g., from 2 to 22, from 2 to 18, from 2 to 17, from 2 to 16, from 3 to 15, from 2 to 14, from 2 to 13, or from 2 to 12 carbons, such as C.sub.1-6 alk-C.sub.1-12 heterocyclyl, C.sub.1-10 alk-C.sub.1-12 heterocyclyl, or C.sub.1-20 alk-C.sub.1-12 heterocyclyl). In some embodiments, the alkylene and the heterocyclyl each can be further substituted with 1, 2, 3, or 4 substituent groups as defined herein for the respective group.
The term "hydrocarbon," as used herein, represents a group consisting only of carbon and hydrogen atoms.
The term "hydroxyl," as used herein, represents an --OH group. In some embodiments, the hydroxyl group can be substituted with a substituent group (e.g., optionally substituted alkyl or an O-protecting group).
The term "isomer," as used herein, means any tautomer, stereoisomer, enantiomer, or diastereomer of any compound of the invention. It is recognized that the compounds of the invention can have one or more chiral centers and/or double bonds and, therefore, exist as stereoisomers, such as double-bond isomers (i.e., geometric E/Z isomers) or diastereomers (e.g., enantiomers (i.e., (+) or (-)) or cis/trans isomers). According to the invention, the chemical structures depicted herein, and therefore the compounds of the invention, encompass all of the corresponding stereoisomers, that is, both the stereomerically pure form (e.g., geometrically pure, enantiomerically pure, or diastereomerically pure) and enantiomeric and stereoisomeric mixtures, e.g., racemates. Enantiomeric and stereoisomeric mixtures of compounds of the invention can typically be resolved into their component enantiomers or stereoisomers by well-known methods, such as chiral-phase gas chromatography, chiral-phase high performance liquid chromatography, crystallizing the compound as a chiral salt complex, or crystallizing the compound in a chiral solvent. Enantiomers and stereoisomers can also be obtained from stereomerically or enantiomerically pure intermediates, reagents, and catalysts by well-known asymmetric synthetic methods.
The term "N-protected amino," as used herein, refers to an amino group, as defined herein, to which is attached one or two N-protecting groups, as defined herein.
The term "N-protecting group," as used herein, represents those groups intended to protect an amino group against undesirable reactions during synthetic procedures. Commonly used N-protecting groups are disclosed in Greene, "Protective Groups in Organic Synthesis," 3.sup.rd Edition (John Wiley & Sons, New York, 1999), which is incorporated herein by reference. N-protecting groups include acyl, aryloyl, or carbamyl groups such as formyl, acetyl, propionyl, pivaloyl, t-butylacetyl, 2-chloroacetyl, 2-bromoacetyl, trifluoroacetyl, trichloroacetyl, phthalyl, o-nitrophenoxyacetyl, .alpha.-chlorobutyryl, benzoyl, 4-chlorobenzoyl, 4-bromobenzoyl, 4-nitrobenzoyl, and chiral auxiliaries such as protected or unprotected D, L or D, L-amino acids such as alanine, leucine, phenylalanine, and the like; sulfonyl-containing groups such as benzenesulfonyl, p-toluenesulfonyl, and the like; carbamate forming groups such as benzyloxycarbonyl, p-chlorobenzyloxycarbonyl, p-methoxybenzyloxycarbonyl, p-nitrobenzyloxycarbonyl, 2-nitrobenzyloxycarbonyl, p-bromobenzyloxycarbonyl, 3,4-dimethoxybenzyloxycarbonyl, 3,5-dimethoxybenzyloxycarbonyl, 2,4-dimethoxybenzyloxycarbonyl, 4-methoxybenzyloxycarbonyl, 2-nitro-4,5-dimethoxybenzyloxycarbonyl, 3,4,5-trimethoxybenzyloxycarbonyl, 1-(p-biphenylyl)-1-methylethoxycarbonyl, .alpha.,.alpha.-dimethyl-3,5-dimethoxybenzyloxycarbonyl, benzhydryloxy carbonyl, t-butyloxycarbonyl, diisopropylmethoxycarbonyl, isopropyloxycarbonyl, ethoxycarbonyl, methoxycarbonyl, allyloxycarbonyl, 2,2,2,-trichloroethoxycarbonyl, phenoxycarbonyl, 4-nitrophenoxy carbonyl, fluorenyl-9-methoxycarbonyl, cyclopentyloxycarbonyl, adamantyloxycarbonyl, cyclohexyloxycarbonyl, phenylthiocarbonyl, and the like, alkaryl groups such as benzyl, triphenylmethyl, benzyloxymethyl, and the like and silyl groups, such as trimethylsilyl, and the like. Preferred N-protecting groups are formyl, acetyl, benzoyl, pivaloyl, t-butylacetyl, alanyl, phenylsulfonyl, benzyl, t-butyloxycarbonyl (Boc), and benzyloxycarbonyl (Cbz).
The term "nitro," as used herein, represents an --NO.sub.2 group.
The term "O-protecting group," as used herein, represents those groups intended to protect an oxygen containing (e.g., phenol, hydroxyl, or carbonyl) group against undesirable reactions during synthetic procedures. Commonly used O-protecting groups are disclosed in Greene, "Protective Groups in Organic Synthesis," 3.sup.rd Edition (John Wiley & Sons, New York, 1999), which is incorporated herein by reference. Exemplary O-protecting groups include acyl, aryloyl, or carbamyl groups, such as formyl, acetyl, propionyl, pivaloyl, t-butylacetyl, 2-chloroacetyl, 2-bromoacetyl, trifluoroacetyl, trichloroacetyl, phthalyl, o-nitrophenoxyacetyl, .alpha.-chlorobutyryl, benzoyl, 4-chlorobenzoyl, 4-bromobenzoyl, t-butyldimethylsilyl, tri-iso-propylsilyloxymethyl, 4,4'-dimethoxytrityl, isobutyryl, phenoxyacetyl, 4-isopropylpehenoxyacetyl, dimethylformamidino, and 4-nitrobenzoyl; alkylcarbonyl groups, such as acyl, acetyl, propionyl, pivaloyl, and the like; optionally substituted arylcarbonyl groups, such as benzoyl; silyl groups, such as trimethylsilyl (TMS), tert-butyldimethylsilyl (TBDMS), tri-iso-propylsilyloxymethyl (TOM), triisopropylsilyl (TIPS), and the like; ether-forming groups with the hydroxyl, such methyl, methoxymethyl, tetrahydropyranyl, benzyl, p-methoxybenzyl, trityl, and the like; alkoxycarbonyls, such as methoxycarbonyl, ethoxycarbonyl, isopropoxycarbonyl, n-isopropoxycarbonyl, n-butyloxycarbonyl, isobutyloxycarbonyl, sec-butyloxycarbonyl, t-butyloxycarbonyl, 2-ethylhexyloxycarbonyl, cyclohexyloxycarbonyl, methyloxycarbonyl, and the like; alkoxyalkoxycarbonyl groups, such as methoxymethoxycarbonyl, ethoxymethoxycarbonyl, 2-methoxyethoxycarbonyl, 2-ethoxyethoxycarbonyl, 2-butoxyethoxycarbonyl, 2-methoxyethoxymethoxycarbonyl, allyloxycarbonyl, propargyloxycarbonyl, 2-butenoxycarbonyl, 3-methyl-2-butenoxycarbonyl, and the like; haloalkoxycarbonyls, such as 2-chloroethoxycarbonyl, 2-chloroethoxycarbonyl, 2,2,2-trichloroethoxycarbonyl, and the like; optionally substituted arylalkoxycarbonyl groups, such as benzyloxycarbonyl, p-methylbenzyloxycarbonyl, p-methoxybenzyloxycarbonyl, p-nitrobenzyloxycarbonyl, 2,4-dinitrobenzyloxycarbonyl, 3,5-dimethylbenzyloxycarbonyl, p-chlorobenzyloxycarbonyl, p-bromobenzyloxy-carbonyl, fluorenylmethyloxycarbonyl, and the like; and optionally substituted aryloxycarbonyl groups, such as phenoxycarbonyl, p-nitrophenoxycarbonyl, o-nitrophenoxycarbonyl, 2,4-dinitrophenoxycarbonyl, p-methylphenoxycarbonyl, m-methylphenoxycarbonyl, o-bromophenoxycarbonyl, 3,5-dimethylphenoxycarbonyl, p-chlorophenoxycarbonyl, 2-chloro-4-nitrophenoxy-carbonyl, and the like); substituted alkyl, aryl, and alkaryl ethers (e.g., trityl; methylthiomethyl; methoxymethyl; benzyloxymethyl; siloxymethyl; 2,2,2,-trichloroethoxymethyl; tetrahydropyranyl; tetrahydrofuranyl; ethoxyethyl; 1-[2-(trimethylsilyl)ethoxy]ethyl; 2-trimethylsilylethyl; t-butyl ether; p-chlorophenyl, p-methoxyphenyl, p-nitrophenyl, benzyl, p-methoxybenzyl, and nitrobenzyl); silyl ethers (e.g., trimethylsilyl; triethylsilyl; triisopropylsilyl; dimethylisopropylsilyl; t-butyldimethylsilyl; t-butyldiphenylsilyl; tribenzylsilyl; triphenylsilyl; and diphenymethylsilyl); carbonates (e.g., methyl, methoxymethyl, 9-fluorenylmethyl; ethyl; 2,2,2-trichloroethyl; 2-(trimethylsilyl)ethyl; vinyl, allyl, nitrophenyl; benzyl; methoxybenzyl; 3,4-dimethoxybenzyl; and nitrobenzyl); carbonyl-protecting groups (e.g., acetal and ketal groups, such as dimethyl acetal, 1,3-dioxolane, and the like; acylal groups; and dithiane groups, such as 1,3-dithianes, 1,3-dithiolane, and the like); carboxylic acid-protecting groups (e.g., ester groups, such as methyl ester, benzyl ester, t-butyl ester, orthoesters, and the like; and oxazoline groups.
The term "oxo" as used herein, represents .dbd.O.
The prefix "perfluoro," as used herein, represents anyl group, as defined herein, where each hydrogen radical bound to the alkyl group has been replaced by a fluoride radical. For example, perfluoroalkyl groups are exemplified by trifluoromethyl, pentafluoroethyl, and the like.
The term "protected hydroxyl," as used herein, refers to an oxygen atom bound to an O-protecting group.
The term "spirocyclyl," as used herein, represents a C.sub.2-7 alkylene diradical, both ends of which are bonded to the same carbon atom of the parent group to form a spirocyclic group, and also a C.sub.1-6 heteroalkylene diradical, both ends of which are bonded to the same atom. The heteroalkylene radical forming the spirocyclyl group can containing one, two, three, or four heteroatoms independently selected from the group consisting of nitrogen, oxygen, and sulfur. In some embodiments, the spirocyclyl group includes one to seven carbons, excluding the carbon atom to which the diradical is attached. The spirocyclyl groups of the invention may be optionally substituted with 1, 2, 3, or 4 substituents provided herein as optional substituents for cycloalkyl and/or heterocyclyl groups.
The term "stereoisomer," as used herein, refers to all possible different isomeric as well as conformational forms which a compound may possess (e.g., a compound of any formula described herein), in particular all possible stereochemically and conformationally isomeric forms, all diastereomers, enantiomers and/or conformers of the basic molecular structure. Some compounds of the present invention may exist in different tautomeric forms, all of the latter being included within the scope of the present invention.
The term "sulfonyl," as used herein, represents an --S(O).sub.2-- group.
The term "thiol," as used herein represents an --SH group.
Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Methods and materials are described herein for use in the present disclosure; other, suitable methods and materials known in the art can also be used. The materials, methods, and examples are illustrative only and not intended to be limiting. All publications, patent applications, patents, sequences, database entries, and other references mentioned herein are incorporated by reference in their entirety. In case of conflict, the present specification, including definitions, will control.
Other features and advantages of the present disclosure will be apparent from the following detailed description and figures, and from the claims.
DETAILED DESCRIPTION OF THE INVENTION
The present disclosure provides, alternative sugar moieties and polynucleotides including these alternatives that may exhibit improved therapeutic properties including, but not limited to, a reduced innate immune response when introduced into a population of cells.
As there remains a need in the art for therapeutic modalities to address the myriad barriers surrounding the efficacious modulation of intracellular translation and processing of polynucleotides encoding polypeptides or fragments thereof, certain mRNA sequences containing alternative sugar moieties may have the potential as therapeutics with benefits beyond just evading, avoiding or diminishing the immune response.
The present invention addresses this need by providing polynucleotides which encode a polypeptide of interest (e.g., unnatural mRNA) and which have structural and/or chemical features that preferably avoid one or more of the problems in the art, for example, features which are useful for optimizing polynucleotide-based therapeutics while retaining structural and functional integrity, overcoming the threshold of expression, improving expression rates, half life and/or protein concentrations, optimizing protein localization, and avoiding deleterious bio-responses such as the immune response and/or degradation pathways.
Polypeptides of interest, according to the present invention may be selected from any of those disclosed in US 2013/0259924, US 2013/0259923, WO 2013/151663, WO 2013/151669, WO 2013/151670, WO 2013/151664, WO 2013/151665, WO 2013/151736, U.S. Provisional Patent Application No. 61/618,862, U.S. Provisional Patent Application No. 61/681,645, U.S. Provisional Patent Application No. 61/618,873, U.S. Provisional Patent Application No. 61/681,650, U.S. Provisional Patent Application No. 61/618,878, U.S. Provisional Patent Application No. 61/681,654, U.S. Provisional Patent Application No. 61/618,885, U.S. Provisional Patent Application No. 61/681,658, U.S. Provisional Patent Application No. 61/618,911 s, U.S. Provisional Patent Application No. 61/681,667, U.S. Provisional Patent Application No. 61/618,922, U.S. Provisional Patent Application No. 61/681,675, U.S. Provisional Patent Application No. 61/618,935, U.S. Provisional Patent Application No. 61/681,687, U.S. Provisional Patent Application No. 61/618,945, U.S. Provisional Patent Application No. 61/681,696, U.S. Provisional Patent Application No. 61/618,953, and U.S. Provisional Patent Application No. 61/681,704, the contents of which are incorporated herein by reference in their entirety.
Provided herein, in part, are polynucleotides encoding polypeptides of interest which contain one or more of alternative sugar moieties of the nucleotide compared to the natural counterpart, to improve one or more of the stability and/or clearance in tissues, receptor uptake and/or kinetics, cellular access by the compositions, engagement with translational machinery, mRNA half-life, translation efficiency, immune evasion, protein production capacity, secretion efficiency (when applicable), accessibility to circulation, protein half-life and/or modulation of a cell's status, function and/or activity.
The alternative sugar moieties of the invention, including the combination of alternatives taught herein may have superior properties making them more suitable as therapeutic modalities.
It has been determined that the "all or none" model in the art is sorely insufficient to describe the biological phenomena associated with the therapeutic utility of mRNA containing alternative nucleotides. To improve protein production, one may consider the nature of the alternative nucleoside, nucleotide, or polynucleotide, or combination of alternative groups, the percent incorporation of the alternatives and survey more than one cytokine or metric to determine the efficacy and risk profile of a particular unnatural mRNA.
In one aspect of the invention, methods of determining the effectiveness of an mRNA containing alternative sugar moieties as compared to natural mRNA involves the measure and analysis of one or more cytokines whose expression is triggered by the administration of the exogenous polynucleotide of the invention. These values are compared to administration of a natural polynucleotide or to a standard metric such as cytokine response, PolyIC, R-848 or other standard known in the art.
One example of a standard metric developed herein is the measure of the ratio of the level or amount of encoded polypeptide (protein) produced in the cell, tissue or organism to the level or amount of one or more (or a panel) of cytokines whose expression is triggered in the cell, tissue or organism as a result of administration or contact with the unnatural polynucleotide. Such ratios are referred to herein as the Protein: Cytokine Ratio or "PC" Ratio. The higher the PC ratio, the more efficacious the unnatural polynucleotide (polynucleotide encoding the protein measured). Preferred PC Ratios, by cytokine, of the present invention may be greater than 1, greater than 10, greater than 100, greater than 1000, greater than 10,000 or more. Alternative polynucleotides having higher PC Ratios than an alternative polynucleotide of a different or natural construct are preferred.
The PC ratio may be further qualified by the percentage of alternative sugar moieties present in the polynucleotide. For example, normalized to a 100% alternative polynucleotide, the protein production as a function of cytokine (or risk) or cytokine profile can be determined.
In one embodiment, the present invention provides a method for determining, across chemistries, cytokines or percentage of alternative nucleotides, the relative efficacy of any particular polynucleotide by comparing the PC Ratio of the alternative polynucleotide to the natural counterpart.
In another embodiment, the mRNA of the invention are substantially non toxic and non mutagenic.
In one embodiment, the alternative sugar moieties and polynucleotides can disrupt interactions, which may cause innate immune responses. Further, these alternative sugar moieties and polynucleotides can be used to deliver a payload, e.g., detectable or therapeutic agent, to a biological target. For example, the polynucleotides can be covalently linked to a payload, e.g. a detectable or therapeutic agent, through a linker attached to the nucleobase or the sugar moiety. The compositions and methods described herein can be used, in vivo and in vitro, both extracellularly or intracellularly, as well as in assays such as cell free assays.
In another aspect, the present disclosure provides alternative sugar moieties that may reduce the cellular innate immune response, as compared to the cellular innate immune induced by a corresponding natural polynucleotide.
In another aspect, the present disclosure provides compositions comprising a compound as described herein. In some embodiments, the composition is a reaction mixture. In some embodiments, the composition is a pharmaceutical composition. In some embodiments, the composition is a cell culture. In some embodiments, the composition further comprises an RNA polymerase and a cDNA template. In some embodiments, the composition further comprises a nucleotide selected from the group consisting of adenosine, cytosine, guanosine, and uracil.
In a further aspect, the present disclosure provides methods of making a pharmaceutical formulation comprising a physiologically active secreted protein, comprising transfecting a first population of human cells with the pharmaceutical polynucleotide made by the methods described herein, wherein the secreted protein is active upon a second population of human cells.
In some embodiments, the secreted protein is capable of interacting with a receptor on the surface of at least one cell present in the second population.
In certain embodiments, provided herein are combination therapeutics containing one or more alternative polynucleotides containing translatable regions that preferably encode for a protein or proteins that boost a mammalian subject's immunity along with a protein that induces antibody dependent cellular toxicity.
In one embodiment, it is intended that the compounds of the present disclosure are stable. It is further appreciated that certain features of the present disclosure, which are, for clarity, described in the context of separate embodiments, can also be provided in combination in a single embodiment. Conversely, various features of the present disclosure which are, for brevity, described in the context of a single embodiment, can also be provided separately or in any suitable subcombination.
Alternative Nucleotides, Nucleosides and Polynucleotides of the Invention
Herein, in a sugar moiety or a polynucleotide (such as the polynucleotides of the invention, e.g., mRNA molecule), the term "alternative" refers to a compound differing chemically with respect to ribose. Generally, herein, this term is not intended to refer to the ribonucleotide modifications in naturally occurring 5'-terminal mRNA cap moieties. In a polypeptide, the term "modification" refers to a modification as compared to the canonical set of 20 amino acids.
The alternatives may be various. In some embodiments, where the polynucleotide is an mRNA, the coding region, the flanking regions and/or the terminal regions may contain one, two, or more (optionally different) alternative sugar moieties. In some embodiments, an alternative polynucleotide introduced to a cell may exhibit reduced degradation in the cell, as compared to a natural polynucleotide.
As described herein, the polynucleotides of the invention preferably do not substantially induce an innate immune response of a cell into which the polynucleotide (e.g., mRNA) is introduced. Features of an induced innate immune response include 1) increased expression of pro-inflammatory cytokines, 2) activation of intracellular PRRs (RIG-I, MDA5, etc, and/or 3) termination or reduction in protein translation.
In certain embodiments, it may desirable for an alternative polynucleotide molecule introduced into the cell to be degraded intracellularly. For example, degradation of an alternative polynucleotide molecule may be preferable if precise timing of protein production is desired. Thus, in some embodiments, the invention provides an alternative polynucleotide molecule containing a degradation domain, which is capable of being acted on in a directed manner within a cell.
The polynucleotides can optionally include other agents (e.g., RNAi-inducing agents, RNAi agents, siRNAs, shRNAs, miRNAs, antisense RNAs, ribozymes, catalytic DNA, tRNA, RNAs that induce triple helix formation, aptamers, vectors, etc.). In some embodiments, the polynucleotides may include one or more messenger RNAs (mRNAs) having one or more alternative nucleoside or nucleotides (i.e., unnatural mRNA molecules). Details for these polynucleotides follow.
Polynucleotides
In some embodiments, the polynucleotides of the invention include a first region of linked nucleosides encoding a polypeptide of interest, a first flanking region located at the 5' terminus of the first region, and a second flanking region located at the 3' terminus of the first region.
Sugar Alternatives
The alternative nucleosides and nucleotides (e.g., building block molecules), which may be incorporated into a polynucleotide (e.g., RNA or mRNA, as described herein), can include an alternative sugar.
Generally, RNA includes the sugar group ribose, which is a 5-membered ring having an oxygen. Exemplary alternative sugar moieties include compounds having the structure of Formula I:
##STR00038##
wherein the dotted line represents an optional double bond;
B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, O, or NR.sup.7;
R.sup.1 is hydrogen or fluorine;
R.sup.2 is hydrogen, fluorine, cyano, azido, or optionally substituted C.sub.1-C.sub.6 alkyl;
R.sup.3 and R.sup.4 are independently hydrogen, optionally substituted hydroxyl, or fluorine;
R.sup.5 and R.sup.6 are independently hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.5 and R.sup.6 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.5 and R.sup.6 is absent when the dotted line is a double bond;
R.sup.7 is hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl, or optionally substituted amino;
each Y.sup.2 is independently hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
each Y.sup.3 is independently absent, O, or S;
each Y.sup.5 is independently O, NH, or CR.sup.8R.sup.9;
each Y.sup.6 is O or S;
each Y.sup.7 is O or NH; and
each R.sup.8 and R.sup.9 is independently hydrogen, fluorine, or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.8 and R.sup.9 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.8 and R.sup.9 is absent when the dotted line is a double bond;
wherein if n is 0, X is O, R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, and Y.sup.5 is O, then at least one of Y.sup.1 and Y.sup.4 is optionally substituted amino, and, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, Y.sup.7 is O, X is O, and R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, then Y.sup.4 is optionally substituted amino;
or a salt thereof.
Additional alternative sugar moieties include compounds having the structure of Formula II:
##STR00039##
wherein B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, or O;
R.sup.1 and R.sup.2 are independently hydrogen or fluorine;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl, or optionally substituted amino;
Y.sup.2 is hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl (e.g., .beta.-cyanoethyl);
Y.sup.3 is absent or O;
wherein if n is 0, X is O, R.sup.1 and R.sup.2 are hydrogen, then at least one of Y.sup.1 and Y.sup.4 is not hydroxyl or protected hydroxy, and, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, X is O, and R.sup.1 and R.sup.2 are hydrogen, then Y.sup.4 is not hydroxyl or protected hydroxyl;
or a salt thereof.
Additional alternative sugar moieties include compounds having the structure of Formula IA:
##STR00040##
wherein the dotted line represents an optional double bond;
B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, O, or NR.sup.7;
R.sup.1 is hydrogen or fluorine;
R.sup.2 is hydrogen, fluorine, cyano, azido, or optionally substituted C.sub.1-C.sub.6 alkyl;
R.sup.3 and R.sup.4 are independently hydrogen, optionally substituted hydroxyl, or fluorine;
R.sup.5 and R.sup.6 are independently hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.5 and R.sup.6 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.5 and R.sup.6 is absent when the dotted line is a double bond;
R.sup.7 is hydrogen or optionally substituted C.sub.1-C.sub.6 alkyl;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl, or optionally substituted amino;
each Y.sup.2 is independently hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl;
each Y.sup.3 is independently absent, O, or S;
each Y.sup.5 is independently O, NH, or CR.sup.8R.sup.9;
each Y.sup.6 is O or S;
each Y.sup.7 is O or NH and
each R.sup.8 and R.sup.9 is independently hydrogen, fluorine, or optionally substituted C.sub.1-C.sub.6 alkyl, or R.sup.8 and R.sup.9 are combined to form an optionally substituted C.sub.3-C.sub.6 cycloalkyl, provided that one of R.sup.8 and R.sup.9 is absent when the dotted line is a double bond;
or a salt thereof.
In some embodiments, if n is 0, X is O, R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, and Y.sup.5 is O, then at least one of Y.sup.1 and Y.sup.4 is optionally substituted amino, and, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, Y.sup.7 is O, X is O, R.sup.1, R.sup.2, R.sup.4, R.sup.5, and R.sup.6 are hydrogen, and R.sup.3 is hydroxyl, then Y.sup.4 is optionally substituted amino.
Additional alternative sugar moieties include compounds having the structure of Formula IIA:
##STR00041##
wherein B is a nucleobase;
m and n are independently an integer from 0 to 3;
X is S, CH.sub.2, SO.sub.2, or O;
R.sup.1 and R.sup.2 are independently hydrogen or fluorine;
Y.sup.1 and Y.sup.4 are independently hydroxyl, protected hydroxyl (e.g., dimethoxytrityl), or optionally substituted amino
Y.sup.2 is hydroxyl or optionally substituted C.sub.1-C.sub.6 heteroalkyl (e.g., optionally substituted C.sub.1-C.sub.6 alkoxy such as .beta.-cyanoethoxy);
Y.sup.3 is absent or O; or a salt thereof.
In certain embodiments, if n is 0, X is O, R.sup.1 and R.sup.2 are hydrogen, then at least one of Y.sup.1 and Y.sup.4 is optionally substituted amino, or, if m is 0, n is 1, Y.sup.1 is optionally substituted amino, Y.sup.2 is optionally substituted C.sub.1-C.sub.6 heteroalkyl, Y.sup.3 is absent, X is O, and R.sup.1 and R.sup.2 are hydrogen, then Y.sup.4 is optionally substituted amino.
Synthesis of Polynucleotide Molecules
The polynucleotide molecules for use in accordance with the invention may be prepared according to any useful technique, as described herein. The alternative sugar moieties used in the synthesis of polynucleotide molecules disclosed herein can be prepared from readily available starting materials using the following general methods and procedures. Where typical or preferred process conditions (e.g., reaction temperatures, times, mole ratios of reactants, solvents, pressures, etc.) are provided, a skilled artisan would be able to optimize and develop additional process conditions. Optimum reaction conditions may vary with the particular reactants or solvent used, but such conditions can be determined by one skilled in the art by routine optimization procedures.
The processes described herein can be monitored according to any suitable method known in the art. For example, product formation can be monitored by spectroscopic means, such as nuclear magnetic resonance spectroscopy (e.g., .sup.1H or .sup.13C) infrared spectroscopy, spectrophotometry (e.g., UV-visible), or mass spectrometry, or by chromatography such as high performance liquid chromatography (HPLC) or thin layer chromatography.
Preparation of polynucleotide molecules of the present invention can involve the protection and deprotection of various chemical groups. The need for protection and deprotection, and the selection of appropriate protecting groups can be readily determined by one skilled in the art. The chemistry of protecting groups can be found, for example, in Greene, et al., Protective Groups in Organic Synthesis, 2d. Ed., Wiley & Sons, 1991, which is incorporated herein by reference in its entirety.
The reactions of the processes described herein can be carried out in suitable solvents, which can be readily selected by one of skill in the art of organic synthesis. Suitable solvents can be substantially nonreactive with the starting materials (reactants), the intermediates, or products at the temperatures at which the reactions are carried out, i.e., temperatures which can range from the solvent's freezing temperature to the solvent's boiling temperature. A given reaction can be carried out in one solvent or a mixture of more than one solvent. Depending on the particular reaction step, suitable solvents for a particular reaction step can be selected.
Resolution of racemic mixtures of unnatural polynucleotides (e.g., polynucleotides or unnatural mRNA molecules) can be carried out by any of numerous methods known in the art. An example method includes fractional recrystallization using a "chiral resolving acid" which is an optically active, salt-forming organic acid. Suitable resolving agents for fractional recrystallization methods are, for example, optically active acids, such as the D and L forms of tartaric acid, diacetyltartaric acid, dibenzoyltartaric acid, mandelic acid, malic acid, lactic acid or the various optically active camphorsulfonic acids. Resolution of racemic mixtures can also be carried out by elution on a column packed with an optically active resolving agent (e.g., dinitrobenzoylphenylglycine). Suitable elution solvent composition can be determined by one skilled in the art.
Different sugar alternatives may exist at various positions in the polynucleotide. One of ordinary skill in the art will appreciate that the alternative sugar moieties may be located at any position(s) of a polynucleotide such that the function of the polynucleotide is not substantially decreased. A polynucleotide may also include a 5' or 3' terminal alternative. The polynucleotide may contain from about 1% to about 100% alternative sugar moieties (either in relation to overall sugar content, or in relation to one or more types of sugar) or any intervening percentage (e.g., from 1% to 20%, from 1% to 25%, from 1% to 50%, from 1% to 60%, from 1% to 70%, from 1% to 80%, from 1% to 90%, from 1% to 95%, from 10% to 20%, from 10% to 25%, from 10% to 50%, from 10% to 60%, from 10% to 70%, from 10% to 80%, from 10% to 90%, from 10% to 95%, from 10% to 100%, from 20% to 25%, from 20% to 50%, from 20% to 60%, from 20% to 70%, from 20% to 80%, from 20% to 90%, from 20% to 95%, from 20% to 100%, from 50% to 60%, from 50% to 70%, from 50% to 80%, from 50% to 90%, from 50% to 95%, from 50% to 100%, from 70% to 80%, from 70% to 90%, from 70% to 95%, from 70% to 100%, from 80% to 90%, from 80% to 95%, from 80% to 100%, from 90% to 95%, from 90% to 100%, and from 95% to 100%).
Other components of polynucleotides are optional, and are beneficial in some embodiments. For example, a 5' untranslated region (UTR) and/or a 3'UTR are provided, wherein either or both may independently contain one or more different nucleotide alternatives. In such embodiments, sugar alternatives may also be present in the translatable region. Also provided are polynucleotides containing a Kozak sequence.
Alternative Polynucleotides
The present disclosure provides polynucleotides, including RNAs such as mRNAs that contain one or more alternative sugar moieties (termed "alternative polynucleotides") as described herein, which may have useful properties including the lack of a substantial induction of the innate immune response of a cell into which the mRNA is introduced. Because these alternative polynucleotides may enhance the efficiency of protein production, intracellular retention of polynucleotides, and viability of contacted cells, as well as possess reduced immunogenicity, these polynucleotides having these properties are also termed "enhanced polynucleotides" herein.
The term "polynucleotide," in its broadest sense, includes any compound that an oligonucleotide chain of two or more nucleotides. Exemplary polynucleotides for use in accordance with the present disclosure include, but are not limited to, one or more of DNA, RNA including messenger mRNA (mRNA), hybrids thereof, RNAi-inducing agents, RNAi agents, siRNAs, shRNAs, miRNAs, antisense RNAs, ribozymes, catalytic DNA, RNAs that induce triple helix formation, aptamers, vectors, etc., described in detail herein.
Provided are alternative polynucleotides containing a translatable region and one, two, or more than two different sugar alternatives. In some embodiments, the alternative polynucleotide exhibits reduced degradation in a cell into which the polynucleotide is introduced, relative to a corresponding natural polynucleotide. Exemplary polynucleotides include ribonucleic acids (RNAs) and deoxyribonucleic acids (DNAs), or a hybrid thereof. In preferred embodiments, the alternative polynucleotide includes messenger RNAs (mRNAs). As described herein, the polynucleotides of the present disclosure preferably do not substantially induce an innate immune response of a cell into which the mRNA is introduced.
In certain embodiments, it is desirable to intracellularly degrade an alternative polynucleotide introduced into the cell, for example if precise timing of protein production is desired. Thus, the present disclosure provides an alternative polynucleotide containing a degradation domain, which is capable of being acted on in a directed manner within a cell.
Other components of polynucleotides are optional, and are beneficial in some embodiments. For example, a 5' untranslated region (UTR) and/or a 3'UTR are provided, wherein either or both may independently contain one or more different sugar alternatives. In such embodiments, sugar alternatives may also be present in the translatable region. Also provided are polynucleotides containing a Kozak sequence.
Further, provided are polynucleotides containing an internal ribosome entry site (IRES). An IRES may act as the sole ribosome binding site, or may serve as one of multiple ribosome binding sites of an mRNA. An mRNA containing more than one functional ribosome binding site may encode several peptides or polypeptides that are translated independently by the ribosomes ("multicistronic mRNA"). When polynucleotides are provided with an IRES, further optionally provided is a second translatable region. Examples of IRES sequences that can be used according to the present disclosure include without limitation, those from picornaviruses (e.g. FMDV), pest viruses (CFFV), polio viruses (PV), encephalomyocarditis viruses (ECMV), foot-and-mouth disease viruses (FMDV), hepatitis C viruses (HCV), classical swine fever viruses (CSFV), murine leukemia virus (MLV), simian immune deficiency viruses (SIV) or cricket paralysis viruses (CrPV).
Major Groove Interacting Partners
As described herein, the phrase "major groove interacting partner" refers RNA recognition receptors that detect and respond to RNA ligands through interactions, e.g. binding, with the major groove face of a nucleotide or polynucleotide. As such, RNA ligands comprising alternative sugar or polynucleotides as described herein decrease interactions with major groove binding partners, and therefore decrease an innate immune response, or expression and secretion of pro-inflammatory cytokines, or both.
Example major groove interacting, e.g. binding, partners include, but are not limited to the following nucleases and helicases. Within membranes, TLRs (Toll-like Receptors) 3, 7, and 8 can respond to single- and double-stranded RNAs. Within the cytoplasm, members of the superfamily 2 class of DEX(D/H) helicases and ATPases can sense RNAs to initiate antiviral responses. These helicases include the RIG-I (retinoic acid-inducible gene I) and MDA5 (melanoma differentiation-associated gene 5). Other examples include laboratory of genetics and physiology 2 (LGP2), HIN-200 domain containing proteins, or Helicase-domain containing proteins.
Prevention or Reduction of Innate Cellular Immune Response
The term "innate immune response" includes a cellular response to exogenous single stranded polynucleotides, generally of viral or bacterial origin, which involves the induction of cytokine expression and release, particularly the interferons, and cell death. Protein synthesis is also reduced during the innate cellular immune response. While it is advantageous to eliminate the innate immune response in a cell which is triggered by introduction of exogenous polynucleotides, the present disclosure provides alternative polynucleotides such as mRNAs that may substantially reduce the immune response, including interferon signaling, without entirely eliminating such a response. In some embodiments, the immune response is reduced by 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, 99%, 99.9%, or greater than 99.9% as compared to the immune response induced by a corresponding natural polynucleotide. Such a reduction can be measured by expression or activity level of Type 1 interferons or the expression of interferon-regulated genes such as the toll-like receptors (e.g., TLR7 and TLR8). Reduction or lack of induction of innate immune response can also be measured by decreased cell death following one or more administrations of unnatural RNAs to a cell population; e.g., cell death is 10%, 25%, 50%, 75%, 85%, 90%, 95%, or over 95% less than the cell death frequency observed with a corresponding natural polynucleotide. Moreover, cell death may affect fewer than 50%, 40%, 30%, 20%, 10%, 5%, 1%, 0.1%, 0.01% or fewer than 0.01% of cells contacted with the alternative polynucleotides.
In some embodiments, the alternative polynucleotides, including mRNA molecules preferably do not induce, or induce only minimally, an immune response by the recipient cell or organism. Such evasion or avoidance of an immune response trigger or activation may be a novel feature of the unnatural polynucleotides of the present invention.
The present disclosure provides for the repeated introduction (e.g., transfection) of alternative polynucleotides into a target cell population, e.g., in vitro, ex vivo, or in vivo. The step of contacting the cell population may be repeated one or more times (such as two, three, four, five or more than five times). In some embodiments, the step of contacting the cell population with the alternative polynucleotides is repeated a number of times sufficient such that a predetermined efficiency of protein translation in the cell population is achieved. Given the preferable reduced cytotoxicity of the target cell population provided by the polynucleotide alternatives, such repeated transfections may be achievable in a diverse array of cell types in vitro and/or in vivo.
Polypeptide Variants
Provided are polynucleotides that encode variant polypeptides, which have a certain identity with a reference polypeptide sequence. The term "identity" as known in the art, refers to a relationship between the sequences of two or more peptides, as determined by comparing the sequences. In the art, "identity" also means the degree of sequence relatedness between peptides, as determined by the number of matches between strings of two or more amino acid residues. "Identity" measures the percent of identical matches between the smaller of two or more sequences with gap alignments (if any) addressed by a particular mathematical model or computer program (i.e., "algorithms"). Identity of related peptides can be readily calculated by known methods. Such methods include, but are not limited to, those described in Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D. W., ed., Academic Press, New York, 1993; Computer Analysis of Sequence Data, Part 1, Griffin, A. M., and Griffin, H. G., eds., Humana Press, New Jersey, 1994; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M. Stockton Press, New York, 1991; and Carillo et al., SIAM J. Applied Math. 48, 1073 (1988).
In some embodiments, the polypeptide variant preferably has the same or a similar activity as the reference polypeptide. Alternatively, the variant may have an altered activity (e.g., increased or decreased) relative to a reference polypeptide. Generally, variants of a particular polynucleotide or polypeptide of the present disclosure will have at least about 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to that particular reference polynucleotide or polypeptide as determined by sequence alignment programs and parameters described herein and known to those skilled in the art.
As recognized by those skilled in the art, protein fragments, functional protein domains, and homologous proteins are also considered to be within the scope of this present disclosure. For example, provided herein is any protein fragment of a reference protein (meaning a polypeptide sequence at least one amino acid residue shorter than a reference polypeptide sequence but otherwise identical) 10, 20, 30, 40, 50, 60, 70, 80, 90, 100 or greater than 100 amino acids in length. In another example, any protein that includes a stretch of about 20, about 30, about 40, about 50, or about 100 amino acids which are about 40%, about 50%, about 60%, about 70%, about 80%, about 90%, about 95%, or about 100% identical to any of the sequences described herein can be utilized in accordance with the present disclosure. In certain embodiments, a protein sequence to be utilized in accordance with the present disclosure includes 2, 3, 4, 5, 6, 7, 8, 9, 10, or more mutations as shown in any of the sequences provided or referenced herein.
Polypeptide Libraries
Also provided are polynucleotide libraries containing alternative nucleosides, wherein the polynucleotides individually contain a first polynucleotide sequence encoding a polypeptide, such as an antibody, protein binding partner, scaffold protein, and other polypeptides known in the art. Preferably, the polynucleotides are mRNA in a form suitable for direct introduction into a target cell host, which in turn synthesizes the encoded polypeptide.
In certain embodiments, multiple variants of a protein, each with different amino acid modification(s), are produced and tested to determine the best variant in terms of pharmacokinetics, stability, biocompatibility, and/or biological activity, or a biophysical property such as expression level. Such a library may contain 10, 10.sup.2, 10.sup.3, 10.sup.4, 10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9, or over 10.sup.9 possible variants (including substitutions, deletions of one or more residues, and insertion of one or more residues).
Polypeptide-Polynucleotide Complexes
Proper protein translation involves the physical aggregation of a number of polypeptides and polynucleotides associated with the mRNA. Provided by the present disclosure are protein-polynucleotide complexes, containing a translatable mRNA having one or more alternative sugars (e.g., at least two different alternative sugars) and one or more polypeptides bound to the mRNA. Generally, the proteins are provided in an amount effective to prevent or reduce an innate immune response of a cell into which the complex is introduced.
Untranslatable Alternative Polynucleotides
As described herein, provided are mRNAs having sequences that are substantially not translatable. Such mRNA is may be effective as a vaccine when administered to a mammalian subject.
Also provided are alternative polynucleotides that contain one or more noncoding regions. Such alternative polynucleotides are generally not translated, but may be capable of binding to and sequestering one or more translational machinery component such as a ribosomal protein or a transfer RNA (tRNA), thereby effectively reducing protein expression in the cell. The alternative polynucleotide may contain a small nucleolar RNA (sno-RNA), micro RNA (miRNA), small interfering RNA (sRNA) or Piwi-interacting RNA (piRNA).
Synthesis of Alternative Polynucleotides
Polynucleotides for use in accordance with the present disclosure may be prepared according to any available technique including, but not limited to chemical synthesis, enzymatic synthesis, which is generally termed in vitro transcription, enzymatic or chemical cleavage of a longer precursor, etc. Methods of synthesizing RNAs are known in the art (see, e.g., Gait, M. J. (ed.) Oligonucleotide synthesis: a practical approach, Oxford [Oxfordshire], Washington, D.C.: IRL Press, 1984; and Herdewijn, P. (ed.) Oligonucleotide synthesis: methods and applications, Methods in Molecular Biology, v. 288 (Clifton, N.J.) Totowa, N.J.: Humana Press, 2005; both of which are incorporated herein by reference).
Different sugar alternatives and/or backbone structures may exist at various positions in the polynucleotide. One of ordinary skill in the art will appreciate that the sugar alternative(s) may be located at any position(s) of a polynucleotide such that the function of the polynucleotide is not substantially decreased. The 5' or 3' terminus may also include an alternative. The polynucleotides may contain at a minimum one and at maximum 100% alternative sugar, or any intervening percentage, such as at least 5% alternative sugars, at least 10% alternative sugars, at least 25% alternative sugars, at least 50% alternative sugars, at least 80% alternative sugars, or at least 90% alternative sugars.
Generally, the shortest length of an unnatural mRNA of the present disclosure can be the length of an mRNA sequence that is sufficient to encode for a dipeptide. In another embodiment, the length of the mRNA sequence is sufficient to encode for a tripeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for a tetrapeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for a pentapeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for a hexapeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for a heptapeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for an octapeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for a nonapeptide. In another embodiment, the length of an mRNA sequence is sufficient to encode for a decapeptide.
Examples of dipeptides that the alternative polynucleotide sequences can encode for include, but are not limited to, carnosine and anserine.
In a further embodiment, the mRNA is greater than 30 nucleotides in length. In another embodiment, the RNA molecule is greater than 35 nucleotides in length. In another embodiment, the length is at least 40 nucleotides. In another embodiment, the length is at least 45 nucleotides. In another embodiment, the length is at least 55 nucleotides. In another embodiment, the length is at least 60 nucleotides. In another embodiment, the length is at least 60 nucleotides. In another embodiment, the length is at least 80 nucleotides. In another embodiment, the length is at least 90 nucleotides. In another embodiment, the length is at least 100 nucleotides. In another embodiment, the length is at least 120 nucleotides. In another embodiment, the length is at least 140 nucleotides. In another embodiment, the length is at least 160 nucleotides. In another embodiment, the length is at least 180 nucleotides. In another embodiment, the length is at least 200 nucleotides. In another embodiment, the length is at least 250 nucleotides. In another embodiment, the length is at least 300 nucleotides. In another embodiment, the length is at least 350 nucleotides. In another embodiment, the length is at least 400 nucleotides. In another embodiment, the length is at least 450 nucleotides. In another embodiment, the length is at least 500 nucleotides. In another embodiment, the length is at least 600 nucleotides. In another embodiment, the length is at least 700 nucleotides. In another embodiment, the length is at least 800 nucleotides. In another embodiment, the length is at least 900 nucleotides. In another embodiment, the length is at least 1000 nucleotides. In another embodiment, the length is at least 1100 nucleotides. In another embodiment, the length is at least 1200 nucleotides. In another embodiment, the length is at least 1300 nucleotides. In another embodiment, the length is at least 1400 nucleotides. In another embodiment, the length is at least 1500 nucleotides. In another embodiment, the length is at least 1600 nucleotides. In another embodiment, the length is at least 1800 nucleotides. In another embodiment, the length is at least 2000 nucleotides. In another embodiment, the length is at least 2500 nucleotides. In another embodiment, the length is at least 3000 nucleotides. In another embodiment, the length is at least 4000 nucleotides. In another embodiment, the length is at least 5000 nucleotides, or greater than 5000 nucleotides.
For example, the alternative polynucleotides described herein can be prepared using methods that are known to those skilled in the art of polynucleotide synthesis.
5' Capping
The 5' cap structure of an mRNA is involved in nuclear export, increasing mRNA stability and binds the mRNA Cap Binding Protein (CBP), which is responsible for mRNA stability in the cell and translation competency through the association of CBP with poly(A) binding protein to form the mature cyclic mRNA species. The cap further assists the removal of 5' proximal introns removal during mRNA splicing.
Endogenous mRNA molecules may be 5'-end capped generating a 5'-ppp-5'-triphosphate linkage between a terminal guanosine cap residue and the 5'-terminal transcribed sense nucleotide of the mRNA. This 5'-guanylate cap may then be methylated to generate an N7-methyl-guanylate residue. The ribose sugars of the terminal and/or anteterminal transcribed nucleotides of the 5' end of the mRNA may optionally also be 2'-O-methylated. 5'-decapping through hydrolysis and cleavage of the guanylate cap structure may target a nucleic acid molecule, such as an mRNA molecule, for degradation.
Modifications to the nucleic acids of the present invention may generate a non-hydrolyzable cap structure preventing decapping and thus increasing mRNA half-life. Because cap structure hydrolysis requires cleavage of 5'-ppp-5' phosphorodiester linkages, modified nucleotides may be used during the capping reaction. For example, a Vaccinia Capping Enzyme from New England Biolabs (Ipswich, Mass.) may be used with .alpha.-thio-guanosine nucleotides according to the manufacturer's instructions to create a phosphorothioate linkage in the 5'-ppp-5' cap. Additional modified guanosine nucleotides may be used such as .alpha.-methyl-phosphonate and seleno-phosphate nucleotides.
Additional modifications include, but are not limited to, 2'-O-methylation of the ribose sugars of 5'-terminal and/or 5'-anteterminal nucleotides of the mRNA (as mentioned above) on the 2'-hydroxyl group of the sugar ring. Multiple distinct 5'-cap structures can be used to generate the 5'-cap of a nucleic acid molecule, such as an mRNA molecule.
5' Cap structures include those described in International Patent Publication Nos. WO2008127688, WO 2008016473, and WO 2011015347, each of which is incorporated herein by reference in its entirety.
Cap analogs, which herein are also referred to as synthetic cap analogs, chemical caps, chemical cap analogs, or structural or functional cap analogs, differ from natural (i.e. endogenous, wild-type or physiological) 5'-caps in their chemical structure, while retaining cap function. Cap analogs may be chemically (i.e. non-enzymatically) or enzymatically synthesized and/linked to a nucleic acid molecule.
For example, the Anti-Reverse Cap Analog (ARCA) cap contains two guanines linked by a 5'-5'-triphosphate group, wherein one guanine contains an N7 methyl group as well as a 3'-O-methyl group (i.e., N7,3'-O-dimethyl-guanosine-5'-triphosphate-5'-guanosine (m.sup.7G-3'mppp-G; which may equivalently be designated 3' O-Me-m7G(5)ppp(5')G). The 3'-O atom of the other, unmodified, guanine becomes linked to the 5'-terminal nucleotide of the capped nucleic acid molecule (e.g. an mRNA or mmRNA). The N7- and 3'-O-methylated guanine provides the terminal moiety of the capped nucleic acid molecule (e.g. mRNA or mmRNA).
Another exemplary cap is mCAP, which is similar to ARCA but has a 2'-O-methyl group on guanosine (i.e., N7,2'-O-dimethyl-guanosine-5'-triphosphate-5'-guanosine, m.sup.7Gm-ppp-G).
In one embodiment, the cap is a dinucleotide cap analog. As a non-limiting example, the dinucleotide cap analog may be modified at different phosphate positions with a boranophosphate group or a phophoroselenoate group such as the dinucleotide cap analogs described in U.S. Pat. No. 8,519,110, the contents of which are herein incorporated by reference in its entirety.
In another embodiment, the cap analog is a N7-(4-chlorophenoxyethyl) substituted dinucleotide form of a cap analog known in the art and/or described herein. Non-limiting examples of a N7-(4-chlorophenoxyethyl) substituted dinucleotide form of a cap analog include a N7-(4-chlorophenoxyethyl)-G(5)ppp(5')G and a N7-(4-chlorophenoxyethyl)-m.sup.3'-OG(5)ppp(5')G cap analog (See e.g., the various cap analogs and the methods of synthesizing cap analogs described in Kore et al. Bioorganic & Medicinal Chemistry 2013 21:4570-4574; the contents of which are herein incorporated by reference in its entirety). In another embodiment, a cap analog of the present invention is a 4-chloro/bromophenoxyethyl analog.
While cap analogs allow for the concomitant capping of a nucleic acid molecule in an in vitro transcription reaction, up to 20% of transcripts remain uncapped. This, as well as the structural differences of a cap analog from endogenous 5'-cap structures of nucleic acids produced by the endogenous, cellular transcription machinery, may lead to reduced translational competency and reduced cellular stability.
Modified nucleic acids of the invention may also be capped post-transcriptionally, using enzymes, in order to generate more authentic 5'-cap structures. As used herein, the phrase "more authentic" refers to a feature that closely mirrors or mimics, either structurally or functionally, an endogenous or wild type feature. That is, a "more authentic" feature is better representative of an endogenous, wild-type, natural or physiological cellular function and/or structure as compared to synthetic features or analogs, etc., of the prior art, or which outperforms the corresponding endogenous, wild-type, natural or physiological feature in one or more respects. Non-limiting examples of more authentic 5'-cap structures of the present invention are those which, among other things, have enhanced binding of cap binding proteins, increased half life, reduced susceptibility to 5' endonucleases and/or reduced 5' decapping, as compared to synthetic 5'-cap structures known in the art (or to a wild-type, natural or physiological 5'-cap structure). For example, recombinant Vaccinia Virus Capping Enzyme and recombinant 2'-O-methyltransferase enzyme can create a canonical 5'-5'-triphosphate linkage between the 5'-terminal nucleotide of an mRNA and a guanine cap nucleotide wherein the cap guanine contains an N7 methylation and the 5'-terminal nucleotide of the mRNA contains a 2'-O-methyl. Such a structure is termed the Cap1 structure. This cap results in a higher translational-competency and cellular stability and a reduced activation of cellular pro-inflammatory cytokines, as compared, e.g., to other 5'cap analog structures known in the art. Cap structures include 7mG(5')ppp(5')N,pN2p (cap 0), 7mG(5')ppp(5')NImpNp (cap 1), 7mG(5')-ppp(5')NImpN2mp (cap 2) and m(7)Gpppm(3)(6,6,2')Apm(2')Apm(2')Cpm(2)(3,2')Up (cap 4).
Because the modified nucleic acids may be capped post-transcriptionally, and because this process is more efficient, nearly 100% of the modified nucleic acids may be capped. This is in contrast to .about.80% when a cap analog is linked to an mRNA in the course of an in vitro transcription reaction.
According to the present invention, 5' terminal caps may include endogenous caps or cap analogs. According to the present invention, a 5' terminal cap may comprise a guanine analog. Useful guanine analogs include inosine, N1-methyl-guanosine, 2'fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino-guanosine, LNA-guanosine, and 2-azido-guanosine.
In one embodiment, the nucleic acids described herein may contain a modified 5'-cap. A modification on the 5'-cap may increase the stability of mRNA, increase the half-life of the mRNA, and could increase the mRNA translational efficiency. The modified 5'-cap may include, but is not limited to, one or more of the following modifications: modification at the 2' and/or 3' position of a capped guanosine triphosphate (GTP), a replacement of the sugar ring oxygen (that produced the carbocyclic ring) with a methylene moiety (CH.sub.2), a modification at the triphosphate bridge moiety of the cap structure, or a modification at the nucleobase (G) moiety.
The 5'-cap structure that may be modified includes, but is not limited to, the caps described herein such as Cap0 having the substrate structure for cap dependent translation of:
##STR00042## or Cap1 having the substrate structure for cap dependent translation of:
##STR00043##
As a non-limiting example, the modified 5'-cap may have the substrate structure for cap dependent translation of:
##STR00044## ##STR00045## ##STR00046## ##STR00047## ##STR00048## where R.sub.1 and R.sub.2 are defined in Table 1:
TABLE-US-00001 TABLE 1 R.sub.1 and R.sub.2 for CAP-022 to CAP096 Cap Structure Number R.sub.1 R.sub.2 CAP-022 C.sub.2H.sub.5 (Ethyl) H CAP-023 H C.sub.2H.sub.5 (Ethyl) CAP-024 C.sub.2H.sub.5 (Ethyl) C.sub.2H.sub.5 (Ethyl) CAP-025 C.sub.3H.sub.7 (Propyl) H CAP-026 H C.sub.3H.sub.7 (Propyl) CAP-027 C.sub.3H.sub.7 (Propyl) C.sub.3H.sub.7 (Propyl) CAP-028 C.sub.4H.sub.9 (Butyl) H CAP-029 H C.sub.4H.sub.9 (Butyl) CAP-030 C.sub.4H.sub.9 (Butyl) C.sub.4H.sub.9 (Butyl) CAP-031 C.sub.5H.sub.11 (Pentyl) H CAP-032 H C.sub.5H.sub.11 (Pentyl) CAP-033 C.sub.5H.sub.11 (Pentyl) C.sub.5H.sub.11 (Pentyl) CAP-034 H.sub.2C--C.ident.CH (Propargyl) H CAP-035 H H.sub.2C--C.ident.CH (Propargyl) CAP-036 H.sub.2C--C.ident.CH (Propargyl) H.sub.2C--C.ident.CH (Propargyl) CAP-037 CH.sub.2CH.dbd.CH.sub.2 (Allyl) H CAP-038 H CH.sub.2CH.dbd.CH.sub.2 (Allyl) CAP-039 CH.sub.2CH.dbd.CH.sub.2 (Allyl) CH.sub.2CH.dbd.CH.sub.2 (Allyl) CAP-040 CH.sub.2OCH.sub.3 (MOM) H CAP-041 H CH.sub.2OCH.sub.3 (MOM) CAP-042 CH.sub.2OCH.sub.3 (MOM) CH.sub.2OCH.sub.3 (MOM) CAP-043 CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) H CAP-044 H CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) CAP-045 CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) CAP-046 CH.sub.2SCH.sub.3 (MTM) H CAP-047 H CH.sub.2SCH.sub.3 (MTM) CAP-048 CH.sub.2SCH.sub.3 (MTM) CH.sub.2SCH.sub.3 (MTM) CAP-049 CH.sub.2C.sub.6H.sub.5 (Benzyl) H CAP-050 H CH.sub.2C.sub.6H.sub.5 (Benzyl) CAP-051 CH.sub.2C.sub.6H.sub.5 (Benzyl) CH.sub.2C.sub.6H.sub.5 (Benzyl) CAP-052 CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) H CAP-053 H CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) CAP-054 CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) CAP-055 CH.sub.2C.sub.6H.sub.4--OMe (p- H Methoxybenzyl) CAP-056 H CH.sub.2C.sub.6H.sub.4--OMe (p- Methoxybenzyl) CAP-057 CH.sub.2C.sub.6H.sub.4--OMe (p- CH.sub.2C.sub.6H.sub.4--OMe (p- Methoxybenzyl) Methoxybenzyl) CAP-058 CH.sub.2C.sub.6H.sub.4--NO.sub.2 H (p-Nitrobenzyl) CAP-059 H CH.sub.2C.sub.6H.sub.4--NO.sub.2 (p-Nitrobenzyl) CAP-060 CH.sub.2C.sub.6H.sub.4--NO.sub.2 CH.sub.2C.sub.6H.sub.4--NO.sub.2 (p-Nitrobenzyl) (p-Nitrobenzyl) CAP-061 CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) H where X = F, Cl, Br or I CAP-062 H CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) where X = F, Cl, Br or I CAP-063 CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) where X = F, Cl, Br or I where X = F, Cl, Br or I CAP-064 CH.sub.2C.sub.6H.sub.4--N.sub.3 H (p-Azidobenzyl) CAP-065 H CH.sub.2C.sub.6H.sub.4--N.sub.3 (p-Azidobenzyl) CAP-066 CH.sub.2C.sub.6H.sub.4--N.sub.3 CH.sub.2C.sub.6H.sub.4--N.sub.3 (p-Azidobenzyl) (p-Azidobenzyl) CAP-067 CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- H Trifluoromethylbenzyl) CAP-068 H CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- Trifluoromethylbenzyl) CAP-069 CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- Trifluoromethylbenzyl) Trifluoromethylbenzyl) CAP-070 CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- H Trifluoromethoxylbenzyl) CAP-071 H CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- Trifluoromethoxylbenzyl) CAP-072 CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- Trifluoromethoxylbenzyl) Trifluoromethoxylbenzyl) CAP-073 CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- H bis(Trifluoromethyl)benzyl] CAP-074 H CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- bis(Trifluoromethyl)benzyl] CAP-075 CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- bis(Trifluoromethyl)benzyl] bis(Trifluoromethyl)benzyl] CAP-076 Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t- H Butyldiphenylsilyl) CAP-077 H Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t- Butyldiphenylsilyl) CAP-078 Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t- Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t- Butyldiphenylsilyl) Butyldiphenylsilyl) CAP-079 CH.sub.2CH.sub.2CH.dbd.CH.sub.2 H (Homoallyl) CAP-080 H CH.sub.2CH.sub.2CH.dbd.CH.sub.2 (Homoallyl) CAP-081 CH.sub.2CH.sub.2CH.dbd.CH.sub.2 CH.sub.2CH.sub.2CH.dbd.CH.sub.2 (Homoallyl) (Homoallyl) CAP-082 P(O)(OH).sub.2 (MP) H CAP-083 H P(O)(OH).sub.2 (MP) CAP-084 P(O)(OH).sub.2 (MP) P(O)(OH).sub.2 (MP) CAP-085 P(S)(OH).sub.2 (Thio-MP) H CAP-086 H P(S)(OH).sub.2 (Thio-MP) CAP-087 P(S)(OH).sub.2 (Thio-MP) P(S)(OH).sub.2 (Thio-MP) CAP-088 P(O)(CH.sub.3)(OH) H (Methylphophonate) CAP-089 H P(O)(CH.sub.3)(OH) (Methylphophonate) CAP-090 P(O)(CH.sub.3)(OH) P(O)(CH.sub.3)(OH) (Methylphophonate) (Methylphophonate) CAP-091 PN(.sup.IPr).sub.2(OCH.sub.2CH.sub.2CN) H (Phosporamidite) CAP-092 H PN(.sup.IPr).sub.2(OCH.sub.2CH.sub.2CN) (Phosporamidite) CAP-093 PN(.sup.IPr).sub.2(OCH.sub.2CH.sub.2CN) PN(.sup.IPr).sub.2(OCH.sub- .2CH.sub.2CN) (Phosporamidite) (Phosporamidite) CAP-094 SO.sub.2CH.sub.3 H (Methanesulfonic acid) CAP-095 H SO.sub.2CH.sub.3 (Methanesulfonic acid) CAP-096 SO.sub.2CH.sub.3 SO.sub.2CH.sub.3 (Methanesulfonic acid) (Methanesulfonic acid)
or
##STR00049## where R.sub.1 and R.sub.2 are defined in Table 5:
TABLE-US-00002 TABLE 2 R.sub.1 and R.sub.2 for CAP-097 to CAP111 Cap Structure Number R.sub.1 R.sub.2 CAP-097 NH.sub.2 (amino) H CAP-098 H NH.sub.2 (amino) CAP-099 NH.sub.2 (amino) NH.sub.2 (amino) CAP-100 N.sub.3 (Azido) H CAP-101 H N.sub.3 (Azido) CAP-102 N.sub.3 (Azido) N.sub.3 (Azido) CAP-103 X (Halo: F, Cl, Br, I) H CAP-104 H X (Halo: F, Cl, Br, I) CAP-105 X (Halo: F, Cl, Br, I) X (Halo: F, Cl, Br, I) CAP-106 SH (Thiol) H CAP-107 H SH (Thiol) CAP-108 SH (Thiol) SH (Thiol) CAP-109 SCH.sub.3 (Thiomethyl) H CAP-110 H SCH.sub.3 (Thiomethyl) CAP-111 SCH.sub.3 (Thiomethyl) SCH.sub.3 (Thiomethyl)
In Table 1, "MOM" stands for methoxymethyl, "MEM" stands for methoxyethoxymethyl, "MTM" stands for methylthiomethyl, "BOM" stands for benzyloxymethyl and "MP" stands for monophosphonate. In Table 1 and 2, "F" stands for fluorine, "Cl" stands for chlorine, "Br" stands for bromine and "I" stands for iodine.
In a non-limiting example, the modified 5'cap may have the substrate structure for vaccinia mRNA capping enzyme of:
##STR00050## ##STR00051## ##STR00052## ##STR00053## ##STR00054## ##STR00055## where R.sub.1 and R.sub.2 are defined in Table 3:
TABLE-US-00003 TABLE 3 R.sub.1 and R.sub.2 for CAP-136 to CAP-210 Cap Structure Number R.sub.1 R.sub.2 CAP-136 C.sub.2H.sub.5 (Ethyl) H CAP-137 H C.sub.2H.sub.5 (Ethyl) CAP-138 C.sub.2H.sub.5 (Ethyl) C.sub.2H.sub.5 (Ethyl) CAP-139 C.sub.3H.sub.7 (Propyl) H CAP-140 H C.sub.3H.sub.7 (Propyl) CAP-141 C.sub.3H.sub.7 (Propyl) C.sub.3H.sub.7 (Propyl) CAP-142 C.sub.4H.sub.9 (Butyl) H CAP-143 H C.sub.4H.sub.9 (Butyl) CAP-144 C.sub.4H.sub.9 (Butyl) C.sub.4H.sub.9 (Butyl) CAP-145 C.sub.5H.sub.11 (Pentyl) H CAP-146 H C.sub.5H.sub.11 (Pentyl) CAP-147 C.sub.5H.sub.11 (Pentyl) C.sub.5H.sub.11 (Pentyl) CAP-148 H.sub.2C--C.ident.CH (Propargyl) H CAP-149 H H.sub.2C--C.ident.CH (Propargyl) CAP-150 H.sub.2C--C.ident.CH (Propargyl) H.sub.2C--C.ident.CH (Propargyl) CAP-151 CH.sub.2CH.dbd.CH.sub.2 (Allyl) H CAP-152 H CH.sub.2CH.dbd.CH.sub.2 (Allyl) CAP-153 CH.sub.2CH.dbd.CH.sub.2 (Allyl) CH.sub.2CH.dbd.CH.sub.2 (Allyl) CAP-154 CH.sub.2OCH.sub.3 (MOM) H CAP-155 H CH.sub.2OCH.sub.3 (MOM) CAP-156 CH.sub.2OCH.sub.3 (MOM) CH.sub.2OCH.sub.3 (MOM) CAP-157 CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) H CAP-158 H CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) CAP-159 CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3 (MEM) CAP-160 CH.sub.2SCH.sub.3 (MTM) H CAP-161 H CH.sub.2SCH.sub.3 (MTM) CAP-162 CH.sub.2SCH.sub.3 (MTM) CH.sub.2SCH.sub.3 (MTM) CAP-163 CH.sub.2C.sub.6H.sub.5 (Benzyl) H CAP-164 H CH.sub.2C.sub.6H.sub.5 (Benzyl) CAP-165 CH.sub.2C.sub.6H.sub.5 (Benzyl) CH.sub.2C.sub.6H.sub.5 (Benzyl) CAP-166 CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) H CAP-167 H CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) CAP-168 CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) CH.sub.2OCH.sub.2C.sub.6H.sub.5 (BOM) CAP-169 CH.sub.2C.sub.6H.sub.4--OMe (p- H Methoxybenzyl) CAP-170 H CH.sub.2C.sub.6H.sub.4--OMe (p- Methoxybenzyl) CAP-171 CH.sub.2C.sub.6H.sub.4--OMe (p- CH.sub.2C.sub.6H.sub.4--OMe (p- Methoxybenzyl) Methoxybenzyl) CAP-172 CH.sub.2C.sub.6H.sub.4--NO.sub.2 (p- H Nitrobenzyl) CAP-173 H CH.sub.2C.sub.6H.sub.4--NO.sub.2 (p- Nitrobenzyl) CAP-174 CH.sub.2C.sub.6H.sub.4--NO.sub.2 (p- CH.sub.2C.sub.6H.sub.4--NO.sub.2 (p- Nitrobenzyl) Nitrobenzyl) CAP-175 CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) H where X = F, Cl, Br or I CAP-176 H CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) where X = F, Cl, Br or I CAP-177 CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) CH.sub.2C.sub.6H.sub.4--X (p-Halobenzyl) where X = F, Cl, Br or I where X = F, Cl, Br or I CAP-178 CH.sub.2C.sub.6H.sub.4--N.sub.3 H (p-Azidobenzyl) CAP-179 H CH.sub.2C.sub.6H.sub.4--N.sub.3 (p-Azidobenzyl) CAP-180 CH.sub.2C.sub.6H.sub.4--N.sub.3 CH.sub.2C.sub.6H.sub.4--N.sub.3 (p-Azidobenzyl) (p-Azidobenzyl) CAP-181 CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- H Trifluoromethylbenzyl) CAP-182 H CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- Trifluoromethylbenzyl) CAP-183 CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- CH.sub.2C.sub.6H.sub.4--CF.sub.3 (p- Trifluoromethylbenzyl) Trifluoromethylbenzyl) CAP-184 CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- H Trifluoromethoxylbenzyl) CAP-185 H CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- Trifluoromethoxylbenzyl) CAP-186 CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- CH.sub.2C.sub.6H.sub.4--OCF.sub.3 (p- Trifluoromethoxylbenzyl) Trifluoromethoxylbenzyl) CAP-187 CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- H bis(Trifluoromethyl)benzyl] CAP-188 H CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- bis(Trifluoromethyl)benzyl] CAP-189 CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- CH.sub.2C.sub.6H.sub.3--(CF.sub.3).sub.2 [2,4- bis(Trifluoromethyl)benzyl] bis(Trifluoromethyl)benzyl] CAP-190 Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t- H Butyldiphenylsilyl) CAP-191 H Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t-Butyldiphenylsilyl) CAP-192 Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 (t- Si(C.sub.6H.sub.5).sub.2C.sub.4H.sub.9 Butyldiphenylsilyl) (t-Butyldiphenylsilyl) CAP-193 CH.sub.2CH.sub.2CH.dbd.CH.sub.2 H (Homoallyl) CAP-194 H CH.sub.2CH.sub.2CH.dbd.CH.sub.2 (Homoallyl) CAP-195 CH.sub.2CH.sub.2CH.dbd.CH.sub.2 CH.sub.2CH.sub.2CH.dbd.CH.sub.2 (Homoallyl) (Homoallyl) CAP-196 P(O)(OH).sub.2 (MP) H CAP-197 H P(O)(OH).sub.2 (MP) CAP-198 P(O)(OH).sub.2 (MP) P(O)(OH).sub.2 (MP) CAP-199 P(S)(OH).sub.2 (Thio-MP) H CAP-200 H P(S)(OH).sub.2 (Thio-MP) CAP-201 P(S)(OH).sub.2 (Thio-MP) P(S)(OH).sub.2 (Thio-MP) CAP-202 P(O)(CH.sub.3)(OH) H (Methylphophonate) CAP-203 H P(O)(CH.sub.3)(OH) (Methylphophonate) CAP-204 P(O)(CH.sub.3)(OH) P(O)(CH.sub.3)(OH) (Methylphophonate) (Methylphophonate) CAP-205 PN(.sup.IPr).sub.2(OCH.sub.2CH.sub.2CN) H (Phosporamidite) CAP-206 H PN(.sup.IPr).sub.2(OCH.sub.2CH.sub.2CN) (Phosporamidite) CAP-207 PN(.sup.IPr).sub.2(OCH.sub.2CH.sub.2CN) PN(.sup.IPr).sub.2(OCH.sub- .2CH.sub.2CN) (Phosporamidite) (Phosporamidite) CAP-208 SO.sub.2CH.sub.3 H (Methanesulfonic acid) CAP-209 H SO.sub.2CH.sub.3 (Methanesulfonic acid) CAP-210 SO.sub.2CH.sub.3 SO.sub.2CH.sub.3 (Methanesulfonic acid) (Methanesulfonic acid)
or
##STR00056## where R.sub.1 and R.sub.2 are defined in Table 4:
TABLE-US-00004 TABLE 4 R.sub.1 and R.sub.2 for CAP-211 to 225 Cap Structure Number R.sub.1 R.sub.2 CAP-211 NH.sub.2 (amino) H CAP-212 H NH.sub.2 (amino) CAP-213 NH.sub.2 (amino) NH.sub.2 (amino) CAP-214 N.sub.3 (Azido) H CAP-215 H N.sub.3 (Azido) CAP-216 N.sub.3 (Azido) N.sub.3 (Azido) CAP-217 X (Halo: F, Cl, Br, I) H CAP-218 H X (Halo: F, Cl, Br, I) CAP-219 X (Halo: F, Cl, Br, I) X (Halo: F, Cl, Br, I) CAP-220 SH (Thiol) H CAP-221 H SH (Thiol) CAP-222 SH (Thiol) SH (Thiol) CAP-223 SCH.sub.3 (Thiomethyl) H CAP-224 H SCH.sub.3 (Thiomethyl) CAP-225 SCH.sub.3 (Thiomethyl) SCH.sub.3 (Thiomethyl)
In Table 3, "MOM" stands for methoxymethyl, "MEM" stands for methoxyethoxymethyl, "MTM" stands for methylthiomethyl, "BOM" stands for benzyloxymethyl and "MP" stands for monophosphonate. In Table 3 and 4, "F" stands for fluorine, "Cl" stands for chlorine, "Br" stands for bromine and "I" stands for iodine.
In another non-limiting example, of the modified capping structure substrates CAP-112-CAP-225 could be added in the presence of vaccinia capping enzyme with a component to create enzymatic activity such as, but not limited to, S-adenosylmethionine (AdoMet), to form a modified cap for mRNA.
In one embodiment, the replacement of the sugar ring oxygen (that produced the carbocyclic ring) with a methylene moiety (CH.sub.2) could create greater stability to the C--N bond against phosphorylases as the C--N bond is resistant to acid or enzymatic hydrolysis. The methylene moiety may also increase the stability of the triphosphate bridge moiety and thus increasing the stability of the mRNA. As a non-limiting example, the cap substrate structure for cap dependent translation may have the structure such as, but not limited to, CAP-014 and CAP-015 and/or the cap substrate structure for vaccinia mRNA capping enzyme such as, but not limited to, CAP-123 and CAP-124. In another example, CAP-112-CAP-122 and/or CAP-125-CAP-225, can be modified by replacing the sugar ring oxygen (that produced the carbocyclic ring) with a methylene moiety (CH.sub.2).
In another embodiment, the triphophosphate bridge may be modified by the replacement of at least one oxygen with sulfur (thio), a borane (BH.sub.3) moiety, a methyl group, an ethyl group, a methoxy group and/or combinations thereof. This modification could increase the stability of the mRNA towards decapping enzymes. As a non-limiting example, the cap substrate structure for cap dependent translation may have the structure such as, but not limited to, CAP-016-CAP-021 and/or the cap substrate structure for vaccinia mRNA capping enzyme such as, but not limited to, CAP-125-CAP-130. In another example, CAP-003-CAP-015, CAP-022-CAP-124 and/or CAP-131-CAP-225, can be modified on the triphosphate bridge by replacing at least one of the triphosphate bridge oxygens with sulfur (thio), a borane (BH.sub.3) moiety, a methyl group, an ethyl group, a methoxy group and/or combinations thereof.
In one embodiment, CAP-001-134 and/or CAP-136-CAP-225 may be modified to be a thioguanosine analog similar to CAP-135. The thioguanosine analog may comprise additional modifications such as, but not limited to, a modification at the triphosphate moiety (e.g., thio, BH.sub.3, CH.sub.3, C.sub.2H.sub.5, OCH.sub.3, S and S with OCH.sub.3), a modification at the 2' and/or 3' positions of 6-thio guanosine as described herein and/or a replacement of the sugar ring oxygen (that produced the carbocyclic ring) as described herein.
In one embodiment, CAP-001-121 and/or CAP-123-CAP-225 may be modified to be a modified 5'cap similar to CAP-122. The modified 5'cap may comprise additional modifications such as, but not limited to, a modification at the triphosphate moiety (e.g., thio, BH.sub.3, CH.sub.3, C.sub.2H.sub.5, OCH.sub.3, S and S with OCH.sub.3), a modification at the 2' and/or 3' positions of 6-thio guanosine as described herein and/or a replacement of the sugar ring oxygen (that produced the carbocyclic ring) as described herein.
In one embodiment, the 5'cap modification may be the attachment of biotin or conjugation at the 2' or 3' position of a GTP.
In another embodiment, the 5' cap modification may include a CF.sub.2 modified triphosphate moiety.
In another embodiment, the triphosphate bridge of any of the cap structures described herein may be replaced with a tetraphosphate or pentaphosphate bridge. Examples of tetraphosphate and pentaphosphate containing bridges and other cap modifications are described in Jemielity, J. et al. RNA 2003 9:1108-1122; Grudzien-Nogalska, E. et al. Methods Mol. Biol. 2013 969:55-72; and Grudzien, E. et al. RNA, 2004 10:1479-1487, each of which is incorporated herein by reference in its entirety.
Terminal Architecture Modifications: Stem Loop
In one embodiment, the nucleic acids of the present invention may include a stem loop such as, but not limited to, a histone stem loop. The stem loop may be a nucleotide sequence that is about 25 or about 26 nucleotides in length such as, but not limited to, SEQ ID NOs: 7-17 as described in International Patent Publication No. WO2013103659, incorporated herein by reference in its entirety. The histone stem loop may be located 3' relative to the coding region (e.g., at the 3' terminus of the coding region). As a non-limiting example, the stem loop may be located at the 3' end of a nucleic acid described herein.
In one embodiment, the stem loop may be located in the second terminal region. As a non-limiting example, the stem loop may be located within an untranslated region (e.g., 3'UTR) in the second terminal region.
In one embodiment, the nucleic acid such as, but not limited to mRNA, which comprises the histone stem loop may be stabilized by the addition of at least one chain terminating nucleoside. Not wishing to be bound by theory, the addition of at least one chain terminating nucleoside may slow the degradation of a nucleic acid and thus can increase the half-life of the nucleic acid.
In one embodiment, the chain terminating nucleoside may be, but is not limited to, those described in International Patent Publication No. WO2013103659, incorporated herein by reference in its entirety. In another embodiment, the chain terminating nucleosides which may be used with the present invention includes, but is not limited to, 3'-deoxyadenosine (cordycepin), 3'-deoxyuridine, 3'-deoxycytosine, 3'-deoxyguanosine, 3'-deoxythymine, 2',3'-dideoxynucleosides, such as 2',3'-dideoxyadenosine, 2',3'-dideoxyuridine, 2',3'-dideoxycytosine, 2',3'-dideoxyguanosine, 2',3'-dideoxythymine, a 2'-deoxynucleoside, or a --O-methylnucleoside.
In another embodiment, the nucleic acid such as, but not limited to mRNA, which comprises the histone stem loop may be stabilized by a modification to the 3'region of the nucleic acid that can prevent and/or inhibit the addition of oligio(U) (see e.g., International Patent Publication No. WO2013103659, incorporated herein by reference in its entirety).
In yet another embodiment, the nucleic acid such as, but not limited to mRNA, which comprises the histone stem loop may be stabilized by the addition of an oligonucleotide that terminates in a 3'-deoxynucleoside, 2',3'-dideoxynucleoside 3'-0-methylnucleosides, 3'-0-ethylnucleosides, 3'-arabinosides, and other modified nucleosides known in the art and/or described herein.
In one embodiment, the nucleic acids of the present invention may include a histone stem loop, a polyA tail sequence and/or a 5'cap structure. The histone stem loop may be before and/or after the polyA tail sequence. The nucleic acids comprising the histone stem loop and a polyA tail sequence may include a chain terminating nucleoside described herein.
In another embodiment, the nucleic acids of the present invention may include a histone stem loop and a 5'cap structure. The 5'cap structure may include, but is not limited to, those described herein and/or known in the art.
In one embodiment, the conserved stem loop region may comprise a miR sequence described herein. As a non-limiting example, the stem loop region may comprise the seed sequence of a miR sequence described herein. In another non-limiting example, the stem loop region may comprise a miR-122 seed sequence.
In another embodiment, the conserved stem loop region may comprise a miR sequence described herein and may also include a TEE sequence.
In one embodiment, the incorporation of a miR sequence and/or a TEE sequence changes the shape of the stem loop region which may increase and/or decrease translation. (see e.g, Kedde et al. A Pumilio-induced RNA structure switch in p27-3'UTR controls miR-221 and miR-22 accessibility. Nature Cell Biology. 2010, herein incorporated by reference in its entirety).
In one embodiment, the modified nucleic acids described herein may comprise at least one histone stem-loop and a polyA sequence or polyadenylation signal. Non-limiting examples of nucleic acid sequences encoding for at least one histone stem-loop and a polyA sequence or a polyadenylation signal are described in International Patent Publication No. WO2013120497, WO2013120629, WO2013120500, WO2013120627, WO2013120498, WO2013120626, WO2013120499 and WO2013120628, the contents of each of which are incorporated herein by reference in their entirety. In one embodiment, the nucleic acid encoding for a histone stem loop and a polyA sequence or a polyadenylation signal may code for a pathogen antigen or fragment thereof such as the nucleic acid sequences described in International Patent Publication No WO2013120499 and WO2013120628, the contents of both of which are incorporated herein by reference in their entirety. In another embodiment, the nucleic acid encoding for a histone stem loop and a polyA sequence or a polyadenylation signal may code for a therapeutic protein such as the nucleic acid sequences described in International Patent Publication No WO2013120497 and WO2013120629, the contents of both of which are incorporated herein by reference in their entirety. In one embodiment, the nucleic acid encoding for a histone stem loop and a polyA sequence or a polyadenylation signal may code for a tumor antigen or fragment thereof such as the nucleic acid sequences described in International Patent Publication No WO2013120500 and WO2013120627, the contents of both of which are incorporated herein by reference in their entirety. In another embodiment, the nucleic acid encoding for a histone stem loop and a polyA sequence or a polyadenylation signal may code for a allergenic antigen or an autoimmune self-antigen such as the nucleic acid sequences described in International Patent Publication No WO2013120498 and WO2013120626, the contents of both of which are incorporated herein by reference in their entirety.
Terminal Architecture Modifications: 3'UTR and Triple Helices
In one embodiment, nucleic acids of the present invention may include a triple helix on the 3' end of the modified nucleic acid, enhanced modified RNA or ribonucleic acid. The 3' end of the nucleic acids of the present invention may include a triple helix alone or in combination with a Poly-A tail.
In one embodiment, the nucleic acid of the present invention may comprise at least a first and a second U-rich region, a conserved stem loop region between the first and second region and an A-rich region. The first and second U-rich region and the A-rich region may associate to form a triple helix on the 3' end of the nucleic acid. This triple helix may stabilize the nucleic acid, enhance the translational efficiency of the nucleic acid and/or protect the 3' end from degradation. Exemplary triple helices include, but are not limited to, the triple helix sequence of metastasis-associated lung adenocarcinoma transcript 1 (MALAT1), MEN-.beta. and polyadenylated nuclear (PAN) RNA (See Wilusz et al., Genes & Development 2012 26:2392-2407; herein incorporated by reference in its entirety). In one embodiment, the 3' end of the modified nucleic acids, enhanced modified RNA or ribonucleic acids of the present invention comprises a first U-rich region comprising TTTTTCTTTT (SEQ ID NO: 1), a second U-rich region comprising TTTTGCTTTTT (SEQ ID NO: 2) or TTTTGCTTTT (SEQ ID NO: 3), an A-rich region comprising AAAAAGCAAAA (SEQ ID NO: 4). In another embodiment, the 3' end of the nucleic acids of the present invention comprises a triple helix formation structure comprising a first U-rich region, a conserved region, a second U-rich region and an A-rich region.
In one embodiment, the triple helix may be formed from the cleavage of a MALAT1 sequence prior to the cloverleaf structure. While not meaning to be bound by theory, MALAT1 is a long non-coding RNA which, when cleaved, forms a triple helix and a tRNA-like cloverleaf structure. The MALAT1 transcript then localizes to nuclear speckles and the tRNA-like cloverleaf localizes to the cytoplasm (Wilusz et al. Cell 2008 135(5): 919-932; incorporated herein by reference in its entirety).
As a non-limiting example, the terminal end of the nucleic acid of the present invention comprising the MALAT1 sequence can then form a triple helix structure, after RNaseP cleavage from the cloverleaf structure, which stabilizes the nucleic acid (Peart et al. Non-mRNA 3' end formation: how the other half lives; WIREs RNA 2013; incorporated herein by reference in its entirety).
In one embodiment, the nucleic acids or mRNA described herein comprise a MALAT1 sequence. In another embodiment, the nucleic acids or mRNA may be polyadenylated. In yet another embodiment, the nucleic acids or mRNA is not polyadenylated but has an increased resistance to degradation compared to unmodified nucleic acids or mRNA.
In one embodiment, the nucleic acids of the present invention may comprise a MALAT1 sequence in the second flanking region (e.g., the 3'UTR). As a non-limiting example, the MALAT1 sequence may be human or mouse.
In another embodiment, the cloverleaf structure of the MALAT1 sequence may also undergo processing by RNaseZ and CCA adding enzyme to form a tRNA-like structure called mascRNA (MALAT1-associated small cytoplasmic RNA). As a non-limiting example, the mascRNA may encode a protein or a fragment thereof and/or may comprise a microRNA sequence. The mascRNA may comprise at least one chemical modification described herein.
Terminal Architecture Modifications: Poly-A Tails
During RNA processing, a long chain of adenine nucleotides (poly-A tail) is normally added to a messenger RNA (mRNA) molecules to increase the stability of the molecule. Immediately after transcription, the 3' end of the transcript is cleaved to free a 3' hydroxyl. Then poly-A polymerase adds a chain of adenine nucleotides to the RNA. The process, called polyadenylation, adds a poly-A tail that is between 100 and 250 residues long.
Methods for the stabilization of RNA by incorporation of chain-terminating nucleosides at the 3'-terminus include those described in International Patent Publication No. WO2013103659, incorporated herein in its entirety.
Unique poly-A tail lengths may provide certain advantages to the modified RNAs of the present invention.
Generally, the length of a poly-A tail of the present invention is greater than 30 nucleotides in length. In another embodiment, the poly-A tail is greater than 35 nucleotides in length. In another embodiment, the length is at least 40 nucleotides. In another embodiment, the length is at least 45 nucleotides. In another embodiment, the length is at least 55 nucleotides. In another embodiment, the length is at least 60 nucleotides. In another embodiment, the length is at least 60 nucleotides. In another embodiment, the length is at least 80 nucleotides. In another embodiment, the length is at least 90 nucleotides. In another embodiment, the length is at least 100 nucleotides. In another embodiment, the length is at least 120 nucleotides. In another embodiment, the length is at least 140 nucleotides. In another embodiment, the length is at least 160 nucleotides. In another embodiment, the length is at least 180 nucleotides. In another embodiment, the length is at least 200 nucleotides. In another embodiment, the length is at least 250 nucleotides. In another embodiment, the length is at least 300 nucleotides. In another embodiment, the length is at least 350 nucleotides. In another embodiment, the length is at least 400 nucleotides. In another embodiment, the length is at least 450 nucleotides. In another embodiment, the length is at least 500 nucleotides. In another embodiment, the length is at least 600 nucleotides. In another embodiment, the length is at least 700 nucleotides. In another embodiment, the length is at least 800 nucleotides. In another embodiment, the length is at least 900 nucleotides. In another embodiment, the length is at least 1000 nucleotides. In another embodiment, the length is at least 1100 nucleotides. In another embodiment, the length is at least 1200 nucleotides. In another embodiment, the length is at least 1300 nucleotides. In another embodiment, the length is at least 1400 nucleotides. In another embodiment, the length is at least 1500 nucleotides. In another embodiment, the length is at least 1600 nucleotides. In another embodiment, the length is at least 1700 nucleotides. In another embodiment, the length is at least 1800 nucleotides. In another embodiment, the length is at least 1900 nucleotides. In another embodiment, the length is at least 2000 nucleotides. In another embodiment, the length is at least 2500 nucleotides. In another embodiment, the length is at least 3000 nucleotides.
In some embodiments, the nucleic acid or mRNA includes from about 30 to about 3,000 nucleotides (e.g., from 30 to 50, from 30 to 100, from 30 to 250, from 30 to 500, from 30 to 750, from 30 to 1,000, from 30 to 1,500, from 30 to 2,000, from 30 to 2,500, from 50 to 100, from 50 to 250, from 50 to 500, from 50 to 750, from 50 to 1,000, from 50 to 1,500, from 50 to 2,000, from 50 to 2,500, from 50 to 3,000, from 100 to 500, from 100 to 750, from 100 to 1,000, from 100 to 1,500, from 100 to 2,000, from 100 to 2,500, from 100 to 3,000, from 500 to 750, from 500 to 1,000, from 500 to 1,500, from 500 to 2,000, from 500 to 2,500, from 500 to 3,000, from 1,000 to 1,500, from 1,000 to 2,000, from 1,000 to 2,500, from 1,000 to 3,000, from 1,500 to 2,000, from 1,500 to 2,500, from 1,500 to 3,000, from 2,000 to 3,000, from 2,000 to 2,500, and from 2,500 to 3,000).
In one embodiment, the poly-A tail may be 80 nucleotides, 120 nucleotides, 160 nucleotides in length on a modified RNA molecule described herein.
In another embodiment, the poly-A tail may be 20, 40, 80, 100, 120, 140 or 160 nucleotides in length on a modified RNA molecule described herein.
In one embodiment, the poly-A tail is designed relative to the length of the overall modified RNA molecule. This design may be based on the length of the coding region of the modified RNA, the length of a particular feature or region of the modified RNA (such as the mRNA), or based on the length of the ultimate product expressed from the modified RNA. When relative to any additional feature of the modified RNA (e.g., other than the mRNA portion which includes the poly-A tail) the poly-A tail may be 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100% greater in length than the additional feature. The poly-A tail may also be designed as a fraction of the modified RNA to which it belongs. In this context, the poly-A tail may be 10, 20, 30, 40, 50, 60, 70, 80, or 90% or more of the total length of the construct or the total length of the construct minus the poly-A tail.
In one embodiment, engineered binding sites and/or the conjugation of nucleic acids or mRNA for Poly-A binding protein may be used to enhance expression. The engineered binding sites may be sensor sequences which can operate as binding sites for ligands of the local microenvironment of the nucleic acids and/or mRNA. As a non-limiting example, the nucleic acids and/or mRNA may comprise at least one engineered binding site to alter the binding affinity of Poly-A binding protein (PABP) and analogs thereof. The incorporation of at least one engineered binding site may increase the binding affinity of the PABP and analogs thereof.
Additionally, multiple distinct nucleic acids or mRNA may be linked together to the PABP (Poly-A binding protein) through the 3'-end using modified nucleotides at the 3'-terminus of the poly-A tail. Transfection experiments can be conducted in relevant cell lines at and protein production can be assayed by ELISA at 12 hr, 24 hr, 48 hr, 72 hr and day 7 post-transfection. As a non-limiting example, the transfection experiments may be used to evaluate the effect on PABP or analogs thereof binding affinity as a result of the addition of at least one engineered binding site.
In one embodiment, a polyA tail may be used to modulate translation initiation. While not wishing to be bound by theory, the polyA til recruits PABP which in turn can interact with translation initiation complex and thus may be essential for protein synthesis.
In another embodiment, a polyA tail may also be used in the present invention to protect against 3'-5' exonuclease digestion.
In one embodiment, the nucleic acids or mRNA of the present invention are designed to include a polyA-G Quartet. The G-quartet is a cyclic hydrogen bonded array of four guanine nucleotides that can be formed by G-rich sequences in both DNA and RNA. In this embodiment, the G-quartet is incorporated at the end of the poly-A tail. The resultant nucleic acid or mRNA may be assayed for stability, protein production and other parameters including half-life at various time points. It has been discovered that the polyA-G quartet results in protein production equivalent to at least 75% of that seen using a poly-A tail of 120 nucleotides alone.
In one embodiment, the nucleic acids or mRNA of the present invention may comprise a polyA tail and may be stabilized by the addition of a chain terminating nucleoside. The nucleic acids and/or mRNA with a polyA tail may further comprise a 5'cap structure.
In another embodiment, the nucleic acids or mRNA of the present invention may comprise a polyA-G Quartet. The nucleic acids and/or mRNA with a polyA-G Quartet may further comprise a 5'cap structure.
In one embodiment, the chain terminating nucleoside which may be used to stabilize the nucleic acid or mRNA comprising a polyA tail or polyA-G Quartet may be, but is not limited to, those described in International Patent Publication No. WO2013103659, incorporated herein by reference in its entirety. In another embodiment, the chain terminating nucleosides which may be used with the present invention includes, but is not limited to, 3'-deoxyadenosine (cordycepin), 3'-deoxyuridine, 3'-deoxycytosine, 3'-deoxyguanosine, 3'-deoxythymine, 2',3'-dideoxynucleosides, such as 2',3'-dideoxyadenosine, 2',3'-dideoxyuridine, 2',3'-dideoxycytosine, 2',3'-dideoxyguanosine, 2',3'-dideoxythymine, a 2'-deoxynucleoside, or a --O-methylnucleoside.
In another embodiment, the nucleic acid such as, but not limited to mRNA, which comprise a polyA tail or a polyA-G Quartet may be stabilized by a modification to the 3'region of the nucleic acid that can prevent and/or inhibit the addition of oligio(U) (see e.g., International Patent Publication No. WO2013103659, incorporated herein by reference in its entirety).
In yet another embodiment, the nucleic acid such as, but not limited to mRNA, which comprise a polyA tail or a polyA-G Quartet may be stabilized by the addition of an oligonucleotide that terminates in a 3'-deoxynucleoside, 2',3'-dideoxynucleoside 3'-0-methylnucleosides, 3'-0-ethylnucleosides, 3'-arabinosides, and other modified nucleosides known in the art and/or described herein.
5'UTR, 3'UTR and Translation Enhancer Elements (TEEs)
In one embodiment, the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA may include at least one translational enhancer polynucleotide, translation enhancer element, translational enhancer elements (collectively referred to as "TEE"s). As a non-limiting example, the TEE may be located between the transcription promoter and the start codon. The polynucleotides, primary constructs, modified nucleic acids and/or mmRNA with at least one TEE in the 5'UTR may include a cap at the 5'UTR. Further, at least one TEE may be located in the 5'UTR of polynucleotides, primary constructs, modified nucleic acids and/or mmRNA undergoing cap-dependent or cap-independent translation.
The term "translational enhancer element" or "translation enhancer element" (herein collectively referred to as "TEE") refers to sequences that increase the amount of polypeptide or protein produced from an mRNA.
In one aspect, TEEs are conserved elements in the UTR which can promote translational activity of a nucleic acid such as, but not limited to, cap-dependent or cap-independent translation. The conservation of these sequences has been previously shown by Panek et al (Nucleic Acids Research, 2013, 1-10; incorporated herein by reference in its entirety) across 14 species including humans.
In one non-limiting example, the TEEs known may be in the 5'-leader of the Gtx homeodomain protein (Chappell et al., Proc. Natl. Acad. Sci. USA 101:9590-9594, 2004, incorporated herein by reference in their entirety).
In another non-limiting example, TEEs are disclosed as SEQ ID NOs: 1-35 in US Patent Publication No. US20090226470, SEQ ID NOs: 1-35 in US Patent Publication US20130177581, SEQ ID NOs: 1-35 in International Patent Publication No. WO2009075886, SEQ ID NOs: 1-5, and 7-645 in International Patent Publication No. WO2012009644, SEQ ID NO: 1 in International Patent Publication No. WO1999024595, SEQ ID NO: 1 in U.S. Pat. No. 6,310,197, and SEQ ID NO: 1 in U.S. Pat. No. 6,849,405, each of which is incorporated herein by reference in its entirety.
In yet another non-limiting example, the TEE may be an internal ribosome entry site (IRES), HCV-IRES or an IRES element such as, but not limited to, those described in U.S. Pat. No. 7,468,275, US Patent Publication Nos. US20070048776 and US20110124100 and International Patent Publication Nos. WO2007025008 and WO2001055369, each of which is incorporated herein by reference in its entirety. The IRES elements may include, but are not limited to, the Gtx sequences (e.g., Gtx9-nt, Gtx8-nt, Gtx7-nt) described by Chappell et al. (Proc. Natl. Acad. Sci. USA 101:9590-9594, 2004) and Zhou et al. (PNAS 102:6273-6278, 2005) and in US Patent Publication Nos. US20070048776 and US20110124100 and International Patent Publication No. WO2007025008, each of which is incorporated herein by reference in its entirety.
"Translational enhancer polynucleotides" or "translation enhancer polynucleotide sequences" are polynucleotides which include one or more of the specific TEE exemplified herein and/or disclosed in the art (see e.g., U.S. Pat. Nos. 6,310,197, 6,849,405, 7,456,273, 7,183,395, US20090226470, US20070048776, US20110124100, US20090093049, US20130177581, WO2009075886, WO2007025008, WO2012009644, WO2001055371 WO1999024595, and EP2610341A1 and EP2610340A1; each of which is incorporated herein by reference in its entirety) or their variants, homologs or functional derivatives. One or multiple copies of a specific TEE can be present in the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA. The TEEs in the translational enhancer polynucleotides can be organized in one or more sequence segments. A sequence segment can harbor one or more of the specific TEEs exemplified herein, with each TEE being present in one or more copies. When multiple sequence segments are present in a translational enhancer polynucleotide, they can be homogenous or heterogeneous. Thus, the multiple sequence segments in a translational enhancer polynucleotide can harbor identical or different types of the specific TEEs exemplified herein, identical or different number of copies of each of the specific TEEs, and/or identical or different organization of the TEEs within each sequence segment.
In one embodiment, the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA may include at least one TEE that is described in International Patent Publication No. WO1999024595, WO2012009644, WO2009075886, WO2007025008, WO1999024595, European Patent Publication No. EP2610341A1 and EP2610340A1, U.S. Pat. Nos. 6,310,197, 6,849,405, 7,456,273, 7,183,395, US Patent Publication No. US20090226470, US20110124100, US20070048776, US20090093049, and US20130177581 each of which is incorporated herein by reference in its entirety. The TEE may be located in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA.
In another embodiment, the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA may include at least one TEE that has at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95% or at least 99% identity with the TEEs described in US Patent Publication Nos. US20090226470, US20070048776, US20130177581 and US20110124100, International Patent Publication No. WO1999024595, WO2012009644, WO2009075886 and WO2007025008, European Patent Publication No. EP2610341A1 and EP2610340A1, U.S. Pat. Nos. 6,310,197, 6,849,405, 7,456,273, 7,183,395, each of which is incorporated herein by reference in its entirety.
In one embodiment, the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA may include at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18 at least 19, at least 20, at least 21, at least 22, at least 23, at least 24, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 55 or more than 60 TEE sequences. The TEE sequences in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may be the same or different TEE sequences. The TEE sequences may be in a pattern such as ABABAB or AABBAABBAABB or ABCABCABC or variants thereof repeated once, twice, or more than three times. In these patterns, each letter, A, B, or C represent a different TEE sequence at the nucleotide level.
In one embodiment, the 5'UTR may include a spacer to separate two TEE sequences. As a non-limiting example, the spacer may be a 15 nucleotide spacer and/or other spacers known in the art. As another non-limiting example, the 5'UTR may include a TEE sequence-spacer module repeated at least once, at least twice, at least 3 times, at least 4 times, at least 5 times, at least 6 times, at least 7 times, at least 8 times and at least 9 times or more than 9 times in the 5'UTR.
In another embodiment, the spacer separating two TEE sequences may include other sequences known in the art which may regulate the translation of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention such as, but not limited to, miR sequences described herein (e.g., miR binding sites and miR seeds). As a non-limiting example, each spacer used to separate two TEE sequences may include a different miR sequence or component of a miR sequence (e.g., miR seed sequence).
In one embodiment, the TEE in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may include at least 5%, at least 10%, at least 15%, at least 20%, at least 25%, at least 30%, at least 35%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 99% or more than 99% of the TEE sequences disclosed in US Patent Publication Nos. US20090226470, US20070048776, US20130177581 and US20110124100, International Patent Publication No. WO1999024595, WO2012009644, WO2009075886 and WO2007025008, European Patent Publication No. EP2610341A1 and EP2610340A1, U.S. Pat. Nos. 6,310,197, 6,849,405, 7,456,273, and U.S. Pat. No. 7,183,395 each of which is incorporated herein by reference in its entirety. In another embodiment, the TEE in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may include a 5-30 nucleotide fragment, a 5-25 nucleotide fragment, a 5-20 nucleotide fragment, a 5-15 nucleotide fragment, a 5-10 nucleotide fragment of the TEE sequences disclosed in US Patent Publication Nos. US20090226470, US20070048776, US20130177581 and US20110124100, International Patent Publication No. WO1999024595, WO2012009644, WO2009075886 and WO2007025008, European Patent Publication No. EP2610341A1 and EP2610340A1, U.S. Pat. Nos. 6,310,197, 6,849,405, 7,456,273, and U.S. Pat. No. 7,183,395; each of which is incorporated herein by reference in its entirety.
In one embodiment, the TEE in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may include at least 5%, at least 10%, at least 15%, at least 20%, at least 25%, at least 30%, at least 35%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 99% or more than 99% of the TEE sequences disclosed in Chappell et al. (Proc. Natl. Acad. Sci. USA 101:9590-9594, 2004) and Zhou et al. (PNAS 102:6273-6278, 2005), in Supplemental Table 1 and in Supplemental Table 2 disclosed by Wellensiek et al (Genome-wide profiling of human cap-independent translation-enhancing elements, Nature Methods, 2013; DOI:10.1038/NMETH.2522); each of which is herein incorporated by reference in its entirety. In another embodiment, the TEE in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may include a 5-30 nucleotide fragment, a 5-25 nucleotide fragment, a 5-20 nucleotide fragment, a 5-15 nucleotide fragment, a 5-10 nucleotide fragment of the TEE sequences disclosed in Chappell et al. (Proc. Natl. Acad. Sci. USA 101:9590-9594, 2004) and Zhou et al. (PNAS 102:6273-6278, 2005), in Supplemental Table 1 and in Supplemental Table 2 disclosed by Wellensiek et al (Genome-wide profiling of human cap-independent translation-enhancing elements, Nature Methods, 2013; DOI:10.1038/NMETH.2522); each of which is incorporated herein by reference in its entirety.
In one embodiment, the TEE used in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention is an IRES sequence such as, but not limited to, those described in U.S. Pat. No. 7,468,275 and International Patent Publication No. WO2001055369, each of which is incorporated herein by reference in its entirety.
In one embodiment, the TEEs used in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may be identified by the methods described in US Patent Publication No. US20070048776 and US20110124100 and International Patent Publication Nos. WO2007025008 and WO2012009644, each of which is incorporated herein by reference in its entirety.
In another embodiment, the TEEs used in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may be a transcription regulatory element described in U.S. Pat. Nos. 7,456,273 and 7,183,395, US Patent Publication No. US20090093049, and International Publication No. WO2001055371, each of which is incorporated herein by reference in its entirety. The transcription regulatory elements may be identified by methods known in the art, such as, but not limited to, the methods described in U.S. Pat. Nos. 7,456,273 and 7,183,395, US Patent Publication No. US20090093049, and International Publication No. WO2001055371, each of which is incorporated herein by reference in its entirety.
In yet another embodiment, the TEE used in the 5'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention is an oligonucleotide or portion thereof as described in U.S. Pat. Nos. 7,456,273 and 7,183,395, US Patent Publication No. US20090093049, and International Publication No. WO2001055371, each of which is incorporated herein by reference in its entirety.
The 5' UTR comprising at least one TEE described herein may be incorporated in a monocistronic sequence such as, but not limited to, a vector system or a nucleic acid vector. As a non-limiting example, the vector systems and nucleic acid vectors may include those described in U.S. Pat. Nos. 7,456,273 and 7,183,395, US Patent Publication No. US20070048776, US20090093049 and US20110124100 and International Patent Publication Nos. WO2007025008 and WO2001055371, each of which is incorporated herein by reference in its entirety.
In one embodiment, the TEEs described herein may be located in the 5'UTR and/or the 3'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA. The TEEs located in the 3'UTR may be the same and/or different than the TEEs located in and/or described for incorporation in the 5'UTR.
In one embodiment, the 3'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA may include at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18 at least 19, at least 20, at least 21, at least 22, at least 23, at least 24, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 55 or more than 60 TEE sequences. The TEE sequences in the 3'UTR of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention may be the same or different TEE sequences. The TEE sequences may be in a pattern such as ABABAB or AABBAABBAABB or ABCABCABC or variants thereof repeated once, twice, or more than three times. In these patterns, each letter, A, B, or C represent a different TEE sequence at the nucleotide level.
In one embodiment, the 3'UTR may include a spacer to separate two TEE sequences. As a non-limiting example, the spacer may be a 15 nucleotide spacer and/or other spacers known in the art. As another non-limiting example, the 3'UTR may include a TEE sequence-spacer module repeated at least once, at least twice, at least 3 times, at least 4 times, at least 5 times, at least 6 times, at least 7 times, at least 8 times and at least 9 times or more than 9 times in the 3'UTR.
In another embodiment, the spacer separating two TEE sequences may include other sequences known in the art which may regulate the translation of the polynucleotides, primary constructs, modified nucleic acids and/or mmRNA of the present invention such as, but not limited to, miR sequences described herein (e.g., miR binding sites and miR seeds). As a non-limiting example, each spacer used to separate two TEE sequences may include a different miR sequence or component of a miR sequence (e.g., miR seed sequence).
In one embodiment, the incorporation of a miR sequence and/or a TEE sequence changes the shape of the stem loop region which may increase and/or decrease translation. (see e.g, Kedde et al. A Pumilio-induced RNA structure switch in p27-3'UTR controls miR-221 and miR-22 accessibility. Nature Cell Biology. 2010, herein incorporated by reference in its entirety).
Heterologous 5'UTRs
A 5' UTR may be provided as a flanking region to the modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention. 5'UTR may be homologous or heterologous to the coding region found in the modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention. Multiple 5' UTRs may be included in the flanking region and may be the same or of different sequences. Any portion of the flanking regions, including none, may be codon optimized and any may independently contain one or more different structural or chemical modifications, before and/or after codon optimization.
Shown in Lengthy Table 21 in U.S. Provisional Application No. 61/775,509, filed Mar. 9, 2013, entitled Heterologous Untranslated Regions for mRNA and in Lengthy Table 21 and in Table 22 in U.S. Provisional Application No. 61/829,372, filed May 31, 2013, entitled Heterologous Untranslated Regions for mRNA, the contents of each of which are incorporated herein by reference in their entirety, is a listing of the start and stop site of the modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention. In Table 21 each 5'UTR (5'UTR-005 to 5'UTR 68511) is identified by its start and stop site relative to its native or wild type (homologous) transcript (ENST; the identifier used in the ENSEMBL database).
To alter one or more properties of the polynucleotides, primary constructs or mmRNA of the invention, 5'UTRs which are heterologous to the coding region of the modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention are engineered into compounds of the invention. The modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids are then administered to cells, tissue or organisms and outcomes such as protein level, localization and/or half-life are measured to evaluate the beneficial effects the heterologous 5'UTR may have on the modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention. Variants of the 5' UTRs may be utilized wherein one or more nucleotides are added or removed to the termini, including A, T, C or G. 5'UTRs may also be codon-optimized or modified in any manner described herein.
Incorporating microRNA Binding Sites
In one embodiment, modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention would not only encode a polypeptide but also a sensor sequence. Sensor sequences include, for example, microRNA binding sites, transcription factor binding sites, structured mRNA sequences and/or motifs, artificial binding sites engineered to act as pseudo-receptors for endogenous nucleic acid binding molecules. Non-limiting examples, of polynucleotides comprising at least one sensor sequence are described in co-pending and co-owned U.S. Provisional Patent Application No. U.S. 61/753,661, filed Jan. 17, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironments, U.S. Provisional Patent Application No. U.S. 61/754,159, filed Jan. 18, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironments, U.S. Provisional Patent Application No. U.S. 61/781,097, filed Mar. 14, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironments, U.S. Provisional Patent Application No. U.S. 61/829,334, filed May 31, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironments, U.S. Provisional Patent Application No. U.S. 61/839,893, filed Jun. 27, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironments, U.S. Provisional Patent Application No. U.S. 61/842,733, filed Jul. 3, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironment, and US Provisional Patent Application No. U.S. 61/857,304, filed Jul. 23, 2013, entitled Signal-Sensor Polynucleotide for the Alteration of Cellular Phenotypes and Microenvironment, the contents of each of which are incorporated herein by reference in their entirety.
In one embodiment, microRNA (miRNA) profiling of the target cells or tissues is conducted to determine the presence or absence of miRNA in the cells or tissues.
microRNAs (or miRNA) are 19-25 nucleotide long noncoding RNAs that bind to the 3'UTR of nucleic acid molecules and down-regulate gene expression either by reducing nucleic acid molecule stability or by inhibiting translation. The modified nucleic acids (mRNA), enhanced modified RNA or ribonucleic acids of the invention may comprise one or more microRNA target sequences, microRNA sequences, or microRNA seeds. Such sequences may correspond to any known microRNA such as those taught in US Publication US2005/0261218 and US Publication US200510059005, the contents of which are incorporated herein by reference in their entirety.
A microRNA sequence comprises a "seed" region, i.e., a sequence in the region of positions 2-8 of the mature microRNA, which sequence has perfect Watson-Crick complementarity to the miRNA target sequence. A microRNA seed may comprise positions 2-8 or 2-7 of the mature microRNA. In some embodiments, a microRNA seed may comprise 7 nucleotides (e.g., nucleotides 2-8 of the mature microRNA), wherein the seed-complementary site in the corresponding miRNA target is flanked by an adenine (A) opposed to microRNA position 1. In some embodiments, a microRNA seed may comprise 6 nucleotides (e.g., nucleotides 2-7 of the mature microRNA), wherein the seed-complementary site in the corresponding miRNA target is flanked by an adenine (A) opposed to microRNA position 1. See for example, Grimson A, Farh K K, Johnston W K, Garrett-Engele P, Lim L P, Bartel D P; Mol Cell. 2007 July 6; 27(1):91-105. The bases of the microRNA seed have complete complementarity with the target sequence. By engineering microRNA target sequences into the 3'UTR of nucleic acids or mRNA of the invention one can target the molecule for degradation or reduced translation, provided the microRNA in question is available. This process will reduce the hazard of off target effects upon nucleic acid molecule delivery. Identification of microRNA, microRNA target regions, and their expression patterns and role in biology have been reported (Bonauer et al., Curr Drug Targets 2010 11:943-949; Anand and Cheresh Curr Opin Hematol 2011 18:171-176; Contreras and Rao Leukemia 2012 26:404-413 (2011 Dec. 20. doi: 10.1038/Ieu.2011.356); Bartel Cell 2009 136:215-233; Landgraf et al, Cell, 2007 129:1401-1414; Gentner and Naldini, Tissue Antigens. 2012 80:393-403 and all references therein; each of which is incorporated herein by reference in its entirety).
For example, if the mRNA is not intended to be delivered to the liver but ends up there, then miR-122, a microRNA abundant in liver, can inhibit the expression of the gene of interest if one or multiple target sites of miR-122 are engineered into the 3'UTR of the modified nucleic acids, enhanced modified RNA or ribonucleic acids. Introduction of one or multiple binding sites for different microRNA can be engineered to further decrease the longevity, stability, and protein translation of a modified nucleic acids, enhanced modified RNA or ribonucleic acids. As used herein, the term "microRNA site" refers to a microRNA target site or a microRNA recognition site, or any nucleotide sequence to which a microRNA binds or associates. It should be understood that "binding" may follow traditional Watson-Crick hybridization rules or may reflect any stable association of the microRNA with the target sequence at or adjacent to the microRNA site.
Conversely, for the purposes of the modified nucleic acids, enhanced modified RNA or ribonucleic acids of the present invention, microRNA binding sites can be engineered out of (i.e. removed from) sequences in which they naturally occur in order to increase protein expression in specific tissues. For example, miR-122 binding sites may be removed to improve protein expression in the liver.
In one embodiment, the modified nucleic acids, enhanced modified RNA or ribonucleic acids of the present invention may include at least one miRNA-binding site in the 3'UTR in order to direct cytotoxic or cytoprotective mRNA therapeutics to specific cells such as, but not limited to, normal and/or cancerous cells (e.g., HEP3B or SNU449).
In another embodiment, the modified nucleic acids, enhanced modified RNA or ribonucleic acids of the present invention may include three miRNA-binding sites in the 3'UTR in order to direct cytotoxic or cytoprotective mRNA therapeutics to specific cells such as, but not limited to, normal and/or cancerous cells (e.g., HEP3B or SNU449).
Regulation of expression in multiple tissues can be accomplished through introduction or removal or one or several microRNA binding sites. The decision of removal or insertion of microRNA binding sites, or any combination, is dependent on microRNA expression patterns and their profilings in diseases.
Examples of tissues where microRNA are known to regulate mRNA, and thereby protein expression, include, but are not limited to, liver (miR-122), muscle (miR-133, miR-206, miR-208), endothelial cells (miR-17-92, miR-126), myeloid cells (miR-142-3p, miR-142-5p, miR-16, miR-21, miR-223, miR-24, miR-27), adipose tissue (let-7, miR-30c), heart (miR-1d, miR-149), kidney (miR-192, miR-194, miR-204), and lung epithelial cells (let-7, miR-133, miR-126).
Specifically, microRNAs are known to be differentially expressed in immune cells (also called hematopoietic cells), such as antigen presenting cells (APCs) (e.g. dendritic cells and macrophages), macrophages, monocytes, B lymphocytes, T lymphocytes, granuocytes, natural killer cells, etc. Immune cell specific microRNAs are involved in immunogenicity, autoimmunity, the immune-response to infection, inflammation, as well as unwanted immune response after gene therapy and tissue/organ transplantation. Immune cells specific microRNAs also regulate many aspects of development, proliferation, differentiation and apoptosis of hematopoietic cells (immune cells). For example, miR-142 and miR-146 are exclusively expressed in the immune cells, particularly abundant in myeloid dendritic cells. It was demonstrated in the art that the immune response to exogenous nucleic acid molecules was shut-off by adding miR-142 binding sites to the 3'UTR of the delivered gene construct, enabling more stable gene transfer in tissues and cells. miR-142 efficiently degrades the exogenous mRNA in antigen presenting cells and suppresses cytotoxic elimination of transduced cells (Annoni A et al., blood, 2009, 114, 5152-5161; Brown B D, et al., Nat med. 2006, 12(5), 585-591; Brown B D, et al., blood, 2007, 110(13): 4144-4152, each of which is incorporated herein by reference in its entirety).
An antigen-mediated immune response can refer to an immune response triggered by foreign antigens, which, when entering an organism, are processed by the antigen presenting cells and displayed on the surface of the antigen presenting cells. T cells can recognize the presented antigen and induce a cytotoxic elimination of cells that express the antigen.
Introducing the miR-142 binding site into the 3'-UTR of a polypeptide of the present invention can selectively repress the gene expression in the antigen presenting cells through miR-142 mediated mRNA degradation, limiting antigen presentation in APCs (e.g. dendritic cells) and thereby preventing antigen-mediated immune response after the delivery of the polynucleotides. The polynucleotides are therefore stably expressed in target tissues or cells without triggering cytotoxic elimination.
In one embodiment, microRNAs binding sites that are known to be expressed in immune cells, in particular, the antigen presenting cells, can be engineered into the polynucleotide to suppress the expression of the sensor-signal polynucleotide in APCs through microRNA mediated RNA degradation, subduing the antigen-mediated immune response, while the expression of the polynucleotide is maintained in non-immune cells where the immune cell specific microRNAs are not expressed. For example, to prevent the immunogenic reaction caused by a liver specific protein expression, the miR-122 binding site can be removed and the miR-142 (and/or mirR-146) binding sites can be engineered into the 3-UTR of the polynucleotide.
To further drive the selective degradation and suppression of mRNA in APCs and macrophage, the polynucleotide may include another negative regulatory element in the 3-UTR, either alone or in combination with mir-142 and/or mir-146 binding sites. As a non-limiting example, one regulatory element is the Constitutive Decay Elements (CDEs).
Immune cells specific microRNAs include, but are not limited to, hsa-let-7a-2-3p, hsa-let-7a-3p, hsa-7a-5p, hsa-let-7c, hsa-let-7e-3p, hsa-let-7e-5p, hsa-let-7g-3p, hsa-let-7g-5p, hsa-let-7i-3p, hsa-let-7i-5p, miR-10a-3p, miR-10a-5p, miR-1184, hsa-let-7f-1-3p, hsa-let-7f-2-5p, hsa-let-7f-5p, miR-125b-1-3p, miR-125b-2-3p, miR-125b-5p, miR-1279, miR-130a-3p, miR-130a-5p, miR-132-3p, miR-132-5p, miR-142-3p, miR-142-5p, miR-143-3p, miR-143-5p, miR-146a-3p, miR-146a-5p, miR-146b-3p, miR-146b-5p, miR-147a, miR-147b, miR-148a-5p, miR-148a-3p, miR-150-3p, miR-150-5p, miR-151b, miR-155-3p, miR-155-5p, miR-15a-3p, miR-15a-5p, miR-15b-5p, miR-15b-3p, miR-16-1-3p, miR-16-2-3p, miR-16-5p, miR-17-5p, miR-181a-3p, miR-181a-5p, miR-181a-2-3p, miR-182-3p, miR-182-5p, miR-197-3p, miR-197-5p, miR-21-5p, miR-21-3p, miR-214-3p, miR-214-5p, miR-223-3p, miR-223-5p, miR-221-3p, miR-221-5p, miR-23b-3p, miR-23b-5p, miR-24-1-5p, miR-24-2-5p, miR-24-3p, miR-26a-1-3p, miR-26a-2-3p, miR-26a-5p, miR-26b-3p, miR-26b-5p, miR-27a-3p, miR-27a-5p, miR-27b-3p, miR-27b-5p, miR-28-3p, miR-28-5p, miR-2909, miR-29a-3p, miR-29a-5p, miR-29b-1-5p, miR-29b-2-5p, miR-29c-3p, miR-29c-5p, miR-30e-3p, miR-30e-5p, miR-331-5p, miR-339-3p, miR-339-5p, miR-345-3p, miR-345-5p, miR-346, miR-34a-3p, miR-34a-5p, miR-363-3p, miR-363-5p, miR-372, miR-377-3p, miR-377-5p, miR-493-3p, miR-493-5p, miR-542, miR-548b-5p, miR548c-5p, miR-548i, miR-548j, miR-548n, miR-574-3p, miR-598, miR-718, miR-935, miR-99a-3p, miR-99a-5p, miR-99b-3p and miR-99b-5p. microRNAs that are enriched in specific types of immune cells are listed in Table 13. Furthermore, novel miroRNAs are discovered in the immune cells in the art through micro-array hybridization and microtome analysis (Jima D D et al, Blood, 2010, 116:e118-e127; Vaz C et al., BMC Genomics, 2010, 11,288, the content of each of which is incorporated herein by reference in its entirety.)
MicroRNAs that are known to be expressed in the liver include, but are not limited to, miR-107, miR-122-3p, miR-122-5p, miR-1228-3p, miR-1228-5p, miR-1249, miR-129-5p, miR-1303, miR-151a-3p, miR-151a-5p, miR-152, miR-194-3p, miR-194-5p, miR-199a-3p, miR-199a-5p, miR-199b-3p, miR-199b-5p, miR-296-5p, miR-557, miR-581, miR-939-3p, miR-939-5p. MicroRNA binding sites from any liver specific microRNA can be introduced to or removed from the polynucleotides to regulate the expression of the polynucleotides in the liver. Liver specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites in order to prevent immune reaction against protein expression in the liver.
MicroRNAs that are known to be expressed in the lung include, but are not limited to, let-7a-2-3p, let-7a-3p, let-7a-5p, miR-126-3p, miR-126-5p, miR-127-3p, miR-127-5p, miR-130a-3p, miR-130a-5p, miR-130b-3p, miR-130b-5p, miR-133a, miR-133b, miR-134, miR-18a-3p, miR-18a-5p, miR-18b-3p, miR-18b-5p, miR-24-1-5p, miR-24-2-5p, miR-24-3p, miR-296-3p, miR-296-5p, miR-32-3p, miR-337-3p, miR-337-5p, miR-381-3p, miR-381-5p. MicroRNA binding sites from any lung specific microRNA can be introduced to or removed from the polynucleotide to regulate the expression of the polynucleotide in the lung. Lung specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites in order to prevent an immune reaction against protein expression in the lung.
MicroRNAs that are known to be expressed in the heart include, but are not limited to, miR-1, miR-133a, miR-133b, miR-149-3p, miR-149-5p, miR-186-3p, miR-186-5p, miR-208a, miR-208b, miR-210, miR-296-3p, miR-320, miR-451a, miR-451b, miR-499a-3p, miR-499a-5p, miR-499b-3p, miR-499b-5p, miR-744-3p, miR-744-5p, miR-92b-3p and miR-92b-5p. MicroRNA binding sites from any heart specific microRNA can be introduced to or removed from the polynucleotides to regulate the expression of the polynucleotides in the heart. Heart specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites to prevent an immune reaction against protein expression in the heart.
MicroRNAs that are known to be expressed in the nervous system include, but are not limited to, miR-124-5p, miR-125a-3p, miR-125a-5p, miR-125b-1-3p, miR-125b-2-3p, miR-125b-5p, miR-1271-3p, miR-1271-5p, miR-128, miR-132-5p, miR-135a-3p, miR-135a-5p, miR-135b-3p, miR-135b-5p, miR-137, miR-139-5p, miR-139-3p, miR-149-3p, miR-149-5p, miR-153, miR-181c-3p, miR-181c-5p, miR-183-3p, miR-183-5p, miR-190a, miR-190b, miR-212-3p, miR-212-5p, miR-219-1-3p, miR-219-2-3p, miR-23a-3p, miR-23a-5p, miR-30a-5p, miR-30b-3p, miR-30b-5p, miR-30c-1-3p, miR-30c-2-3p, miR-30c-5p, miR-30d-3p, miR-30d-5p, miR-329, miR-342-3p, miR-3665, miR-3666, miR-380-3p, miR-380-5p, miR-383, miR-410, miR-425-3p, miR-425-5p, miR-454-3p, miR-454-5p, miR-483, miR-510, miR-516a-3p, miR-548b-5p, miR-548c-5p, miR-571, miR-7-1-3p, miR-7-2-3p, miR-7-5p, miR-802, miR-922, miR-9-3p and miR-9-5p. MicroRNAs enriched in the nervous system further include those specifically expressed in neurons, including, but not limited to, miR-132-3p, miR-132-3p, miR-148b-3p, miR-148b-5p, miR-151a-3p, miR-151a-5p, miR-212-3p, miR-212-5p, miR-320b, miR-320e, miR-323a-3p, miR-323a-5p, miR-324-5p, miR-325, miR-326, miR-328, miR-922 and those specifically expressed in glial cells, including, but not limited to, miR-1250, miR-219-1-3p, miR-219-2-3p, miR-219-5p, miR-23a-3p, miR-23a-5p, miR-3065-3p, miR-3065-5p, miR-30e-3p, miR-30e-5p, miR-32-5p, miR-338-5p, miR-657. MicroRNA binding sites from any CNS specific microRNA can be introduced to or removed from the polynucleotides to regulate the expression of the polynucleotide in the nervous system. Nervous system specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites in order to prevent immune reaction against protein expression in the nervous system.
MicroRNAs that are known to be expressed in the pancreas include, but are not limited to, miR-105-3p, miR-105-5p, miR-184, miR-195-3p, miR-195-5p, miR-196a-3p, miR-196a-5p, miR-214-3p, miR-214-5p, miR-216a-3p, miR-216a-5p, miR-30a-3p, miR-33a-3p, miR-33a-5p, miR-375, miR-7-1-3p, miR-7-2-3p, miR-493-3p, miR-493-5p and miR-944. MicroRNA binding sites from any pancreas specific microRNA can be introduced to or removed from the polynucleotide to regulate the expression of the polynucleotide in the pancreas. Pancreas specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites in order to prevent an immune reaction against protein expression in the pancreas.
MicroRNAs that are known to be expressed in the kidney further include, but are not limited to, miR-122-3p, miR-145-5p, miR-17-5p, miR-192-3p, miR-192-5p, miR-194-3p, miR-194-5p, miR-20a-3p, miR-20a-5p, miR-204-3p, miR-204-5p, miR-210, miR-216a-3p, miR-216a-5p, miR-296-3p, miR-30a-3p, miR-30a-5p, miR-30b-3p, miR-30b-5p, miR-30c-1-3p, miR-30c-2-3p, miR30c-5p, miR-324-3p, miR-335-3p, miR-335-5p, miR-363-3p, miR-363-5p and miR-562. MicroRNA binding sites from any kidney specific microRNA can be introduced to or removed from the polynucleotide to regulate the expression of the polynucleotide in the kidney. Kidney specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites to prevent an immune reaction against protein expression in the kidney.
MicroRNAs that are known to be expressed in the muscle further include, but are not limited to, let-7g-3p, let-7g-5p, miR-1, miR-1286, miR-133a, miR-133b, miR-140-3p, miR-143-3p, miR-143-5p, miR-145-3p, miR-145-5p, miR-188-3p, miR-188-5p, miR-206, miR-208a, miR-208b, miR-25-3p and miR-25-5p. MicroRNA binding sites from any muscle specific microRNA can be introduced to or removed from the polynucleotide to regulate the expression of the polynucleotide in the muscle. Muscle specific microRNAs binding sites can be engineered alone or further in combination with immune cells (e.g. APCs) microRNA binding sites to prevent an immune reaction against protein expression in the muscle.
MicroRNAs are differentially expressed in different types of cells, such as endothelial cells, epithelial cells and adipocytes. For example, microRNAs that are expressed in endothelial cells include, but are not limited to, let-7b-3p, let-7b-5p, miR-100-3p, miR-100-5p, miR-101-3p, miR-101-5p, miR-126-3p, miR-126-5p, miR-1236-3p, miR-1236-5p, miR-130a-3p, miR-130a-5p, miR-17-5p, miR-17-3p, miR-18a-3p, miR-18a-5p, miR-19a-3p, miR-19a-5p, miR-19b-1-5p, miR-19b-2-5p, miR-19b-3p, miR-20a-3p, miR-20a-5p, miR-217, miR-210, miR-21-3p, miR-21-5p, miR-221-3p, miR-221-5p, miR-222-3p, miR-222-5p, miR-23a-3p, miR-23a-5p, miR-296-5p, miR-361-3p, miR-361-5p, miR-421, miR-424-3p, miR-424-5p, miR-513a-5p, miR-92a-1-5p, miR-92a-2-5p, miR-92a-3p, miR-92b-3p and miR-92b-5p. Many novel microRNAs are discovered in endothelial cells from deep-sequencing analysis (Voellenkle C et al., RNA, 2012, 18, 472-484, herein incorporated by reference in its entirety) microRNA binding sites from any endothelial cell specific microRNA can be introduced to or removed from the polynucleotide to modulate the expression of the polynucleotide in the endothelial cells in various conditions.
For further example, microRNAs that are expressed in epithelial cells include, but are not limited to, let-7b-3p, let-7b-5p, miR-1246, miR-200a-3p, miR-200a-5p, miR-200b-3p, miR-200b-5p, miR-200c-3p, miR-200c-5p, miR-338-3p, miR-429, miR-451a, miR-451b, miR-494, miR-802 and miR-34a, miR-34b-5p, miR-34c-5p, miR-449a, miR-449b-3p, miR-449b-5p specific in respiratory ciliated epithelial cells; let-7 family, miR-133a, miR-133b, miR-126 specific in lung epithelial cells; miR-382-3p, miR-382-5p specific in renal epithelial cells and miR-762 specific in corneal epithelial cells. MicroRNA binding sites from any epithelial cell specific MicroRNA can be introduced to or removed from the polynucleotide to modulate the expression of the polynucleotide in the epithelial cells in various conditions.
In addition, a large group of microRNAs are enriched in embryonic stem cells, controlling stem cell self-renewal as well as the development and/or differentiation of various cell lineages, such as neural cells, cardiac, hematopoietic cells, skin cells, osteogenic cells and muscle cells (Kuppusamy K T et al., Curr. Mol Med, 2013, 13(5), 757-764; Vidigal J A and Ventura A, Semin Cancer Biol. 2012, 22(5-6), 428-436; Goff L A et al., PLoS One, 2009, 4:e7192; Morin R D et al., Genome Res, 2008, 18, 610-621; Yoo J K et al., Stem Cells Dev. 2012, 21(11), 2049-2057, each of which is herein incorporated by reference in its entirety). MicroRNAs abundant in embryonic stem cells include, but are not limited to, let-7a-2-3p, let-a-3p, let-7a-5p, let7d-3p, let-7d-5p, miR-103a-2-3p, miR-103a-5p, miR-106b-3p, miR-106b-5p, miR-1246, miR-1275, miR-138-1-3p, miR-138-2-3p, miR-138-5p, miR-154-3p, miR-154-5p, miR-200c-3p, miR-200c-5p, miR-290, miR-301a-3p, miR-301a-5p, miR-302a-3p, miR-302a-5p, miR-302b-3p, miR-302b-5p, miR-302c-3p, miR-302c-5p, miR-302d-3p, miR-302d-5p, miR-302e, miR-367-3p, miR-367-5p, miR-369-3p, miR-369-5p, miR-370, miR-371, miR-373, miR-380-5p, miR-423-3p, miR-423-5p, miR-486-5p, miR-520c-3p, miR-548e, miR-548f, miR-548g-3p, miR-548g-5p, miR-548i, miR-548k, miR-548l, miR-548m, miR-548n, miR-548o-3p, miR-548o-5p, miR-548p, miR-664a-3p, miR-664a-5p, miR-664b-3p, miR-664b-5p, miR-766-3p, miR-766-5p, miR-885-3p, miR-885-5p, miR-93-3p, miR-93-5p, miR-941, miR-96-3p, miR-96-5p, miR-99b-3p and miR-99b-5p. Many predicted novel microRNAs are discovered by deep sequencing in human embryonic stem cells (Morin R D et al., Genome Res, 2008, 18, 610-621; Goff L A et al., PLoS One, 2009, 4:e7192; Bar M et al., Stem cells, 2008, 26, 2496-2505, the content of each of which is incorporated herein by references in its entirety).
In one embodiment, the binding sites of embryonic stem cell specific microRNAs can be included in or removed from the 3-UTR of the polynucleotide to modulate the development and/or differentiation of embryonic stem cells, to inhibit the senescence of stem cells in a degenerative condition (e.g. degenerative diseases), or to stimulate the senescence and apoptosis of stem cells in a disease condition (e.g. cancer stem cells).
Many microRNA expression studies are conducted in the art to profile the differential expression of microRNAs in various cancer cells/tissues and other diseases. Some microRNAs are abnormally over-expressed in certain cancer cells and others are under-expressed. For example, microRNAs are differentially expressed in cancer cells (WO2008/154098, US2013/0059015, US2013/0042333, WO2011/157294); cancer stem cells (US2012/0053224); pancreatic cancers and diseases (US2009/0131348, US2011/0171646, US2010/0286232, U.S. Pat. No. 8,389,210); asthma and inflammation (U.S. Pat. No. 8,415,096); prostate cancer (US2013/0053264); hepatocellular carcinoma (WO2012/151212, US2012/0329672, WO2008/054828, U.S. Pat. No. 8,252,538); lung cancer cells (WO2011/076143, WO2013/033640, WO2009/070653, US2010/0323357); cutaneous T cell lymphoma (WO2013/011378); colorectal cancer cells (WO2011/0281756, WO2011/076142); cancer positive lymph nodes (WO2009/100430, US2009/0263803); nasopharyngeal carcinoma (EP2112235); chronic obstructive pulmonary disease (US2012/0264626, US2013/0053263); thyroid cancer (WO2013/066678); ovarian cancer cells (US2012/0309645, WO2011/095623); breast cancer cells (WO2008/154098, WO2007/081740, US2012/0214699), leukemia and lymphoma (WO2008/073915, US2009/0092974, US2012/0316081, US2012/0283310, WO2010/018563, the content of each of which is incorporated herein by reference in its entirety.)
As a non-limiting example, microRNA sites that are over-expressed in certain cancer and/or tumor cells can be removed from the 3-UTR of the polynucleotide encoding the polypeptide of interest, restoring the expression suppressed by the over-expressed microRNAs in cancer cells, thus ameliorating the corresponsive biological function, for instance, transcription stimulation and/or repression, cell cycle arrest, apoptosis and cell death. Normal cells and tissues, wherein microRNAs expression is not up-regulated, will remain unaffected.
MicroRNA can also regulate complex biological processes such as angiogenesis (miR-132) (Anand and Cheresh Curr Opin Hematol 2011 18:171-176). In the modified nucleic acids, enhanced modified RNA or ribonucleic acids of the invention, binding sites for microRNAs that are involved in such processes may be removed or introduced, in order to tailor the expression of the modified nucleic acids, enhanced modified RNA or ribonucleic acids expression to biologically relevant cell types or to the context of relevant biological processes. In this context, the mRNA are defined as auxotrophic mRNA.
MicroRNA gene regulation may be influenced by the sequence surrounding the microRNA such as, but not limited to, the species of the surrounding sequence, the type of sequence (e.g., heterologous, homologous and artificial), regulatory elements in the surrounding sequence and/or structural elements in the surrounding sequence. The microRNA may be influenced by the 5'UTR and/or the 3'UTR. As a non-limiting example, a non-human 3'UTR may increase the regulatory effect of the microRNA sequence on the expression of a polypeptide of interest compared to a human 3'UTR of the same sequence type.
In one embodiment, other regulatory elements and/or structural elements of the 5'-UTR can influence microRNA mediated gene regulation. One example of a regulatory element and/or structural element is a structured IRES (Internal Ribosome Entry Site) in the 5'UTR, which is necessary for the binding of translational elongation factors to initiate protein translation. EIF4A2 binding to this secondarily structured element in the 5'UTR is necessary for microRNA mediated gene expression (Meijer H A et al., Science, 2013, 340, 82-85, herein incorporated by reference in its entirety). The modified nucleic acids, enhanced modified RNA or ribonucleic acids of the invention can further be modified to include this structured 5'-UTR in order to enhance microRNA mediated gene regulation.
At least one microRNA site can be engineered into the 3' UTR of the modified nucleic acids, enhanced modified RNA or ribonucleic acids of the present invention. In this context, at least two, at least three, at least four, at least five, at least six, at least seven, at least eight, at least nine, at least ten or more microRNA sites may be engineered into the 3' UTR of the ribonucleic acids of the present invention. In one embodiment, the microRNA sites incorporated into the modified nucleic acids, enhanced modified RNA or ribonucleic acids may be the same or may be different microRNA sites. In another embodiment, the microRNA sites incorporated into the modified nucleic acids, enhanced modified RNA or ribonucleic acids may target the same or different tissues in the body. As a non-limiting example, through the introduction of tissue-, cell-type-, or disease-specific microRNA binding sites in the 3' UTR of a modified nucleic acid mRNA, the degree of expression in specific cell types (e.g. hepatocytes, myeloid cells, endothelial cells, cancer cells, etc.) can be reduced.
In one embodiment, a microRNA site can be engineered near the 5' terminus of the 3'UTR, about halfway between the 5' terminus and 3'terminus of the 3'UTR and/or near the 3'terminus of the 3'UTR. As a non-limiting example, a microRNA site may be engineered near the 5' terminus of the 3'UTR and about halfway between the 5' terminus and 3'terminus of the 3'UTR. As another non-limiting example, a microRNA site may be engineered near the 3'terminus of the 3'UTR and about halfway between the 5' terminus and 3'terminus of the 3'UTR. As yet another non-limiting example, a microRNA site may be engineered near the 5' terminus of the 3'UTR and near the 3' terminus of the 3'UTR.
In another embodiment, a 3'UTR can comprise 4 microRNA sites. The microRNA sites may be complete microRNA binding sites, microRNA seed sequences and/or microRNA binding site sequences without the seed sequence.
In one embodiment, a nucleic acid of the invention may be engineered to include at least one microRNA in order to dampen the antigen presentation by antigen presenting cells. The microRNA may be the complete microRNA sequence, the microRNA seed sequence, the microRNA sequence without the seed or a combination thereof. As a non-limiting example, the microRNA incorporated into the nucleic acid may be specific to the hematopoietic system. As another non-limiting example, the microRNA incorporated into the nucleic acid of the invention to dampen antigen presentation is miR-142-3p.
In one embodiment, a nucleic acid may be engineered to include microRNA sites which are expressed in different tissues of a subject. As a non-limiting example, a modified nucleic acid, enhanced modified RNA or ribonucleic acid of the present invention may be engineered to include miR-192 and miR-122 to regulate expression of the modified nucleic acid, enhanced modified RNA or ribonucleic acid in the liver and kidneys of a subject. In another embodiment, a modified nucleic acid, enhanced modified RNA or ribonucleic acid may be engineered to include more than one microRNA sites for the same tissue. For example, a modified nucleic acid, enhanced modified RNA or ribonucleic acid of the present invention may be engineered to include miR-17-92 and miR-126 to regulate expression of the modified nucleic acid, enhanced modified RNA or ribonucleic acid in endothelial cells of a subject.
In one embodiment, the therapeutic window and or differential expression associated with the target polypeptide encoded by the modified nucleic acid, enhanced modified RNA or ribonucleic acid encoding a signal (also referred to herein as a polynucleotide) of the invention may be altered. For example, polynucleotides may be designed whereby a death signal is more highly expressed in cancer cells (or a survival signal in a normal cell) by virtue of the miRNA signature of those cells. Where a cancer cell expresses a lower level of a particular miRNA, the polynucleotide encoding the binding site for that miRNA (or miRNAs) would be more highly expressed. Hence, the target polypeptide encoded by the polynucleotide is selected as a protein which triggers or induces cell death. Neighboring noncancer cells, harboring a higher expression of the same miRNA would be less affected by the encoded death signal as the polynucleotide would be expressed at a lower level due to the effects of the miRNA binding to the binding site or "sensor" encoded in the 3'UTR. Conversely, cell survival or cytoprotective signals may be delivered to tissues containing cancer and non-cancerous cells where a miRNA has a higher expression in the cancer cells--the result being a lower survival signal to the cancer cell and a larger survival signature to the normal cell. Multiple polynucleotides may be designed and administered having different signals according to the previous paradigm.
In one embodiment, the expression of a nucleic acid may be controlled by incorporating at least one sensor sequence in the nucleic acid and formulating the nucleic acid. As a non-limiting example, a nucleic acid may be targeted to an orthotopic tumor by having a nucleic acid incorporating a miR-122 binding site and formulated in a lipid nanoparticle comprising the cationic lipid DLin-KC2-DMA.
According to the present invention, the polynucleotides may be modified as to avoid the deficiencies of other polypeptide-encoding molecules of the art. Hence, in this embodiment the polynucleotides are referred to as modified polynucleotides.
Through an understanding of the expression patterns of microRNA in different cell types, modified nucleic acids, enhanced modified RNA or ribonucleic acids such as polynucleotides can be engineered for more targeted expression in specific cell types or only under specific biological conditions. Through introduction of tissue-specific microRNA binding sites, modified nucleic acids, enhanced modified RNA or ribonucleic acids, could be designed that would be optimal for protein expression in a tissue or in the context of a biological condition.
Transfection experiments can be conducted in relevant cell lines, using engineered modified nucleic acids, enhanced modified RNA or ribonucleic acids and protein production can be assayed at various time points post-transfection. For example, cells can be transfected with different microRNA binding site-engineering nucleic acids or mRNA and by using an ELISA kit to the relevant protein and assaying protein produced at 6 hr, 12 hr, 24 hr, 48 hr, 72 hr and 7 days post-transfection. In vivo experiments can also be conducted using microRNA-binding site-engineered molecules to examine changes in tissue-specific expression of formulated modified nucleic acids, enhanced modified RNA or ribonucleic acids.
Non-limiting examples of cell lines which may be useful in these investigations include those from ATCC (Manassas, Va.) including MRC-5, A549, T84, NCI-H2126 [H2126], NCI-H1688 [H1688], WI-38, WI-38 VA-13 subline 2RA, WI-26 VA4, C3A [HepG2/C3A, derivative of Hep G2 (ATCC HB-8065)], THLE-3, H69AR, NCI-H292 [H292], CFPAC-1, NTERA-2 cl.D1 [NT2/D1], DMS 79, DMS 53, DMS 153, DMS 114, MSTO-211H, SW 1573 [SW-1573, SW1573], SW 1271 [SW-1271, SW1271], SHP-77, SNU-398, SNU-449, SNU-182, SNU-475, SNU-387, SNU-423, NL20, NL20-TA [NL20T-A], THLE-2, HBE135-E6E7, HCC827, HCC4006, NCI-H23 [H23], NCI-H1299, NCI-H187 [H187], NCI-H358 [H-358, H358], NCI-H378 [H378], NCI-H522 [H522], NCI-H526 [H526], NCI-H727 [H727], NCI-H810 [H810], NCI-H889 [H889], NCI-H1155 [H1155], NCI-H1404 [H1404], NCI-N87 [N87], NCI-H196 [H196], NCI-H211 [H211], NCI-H220 [H220], NCI-H250 [H250], NCI-H524 [H524], NCI-H647 [H647], NCI-H650 [H650], NCI-H711 [H711], NCI-H719 [H719], NCI-H740 [H740], NCI-H748 [H748], NCI-H774 [H774], NCI-H838 [H838], NCI-H841 [H841], NCI-H847 [H847], NCI-H865 [H865], NCI-H920 [H920], NCI-H1048 [H1048], NCI-H1092 [H1092], NCI-H1105 [H1105], NCI-H1184 [H1184], NCI-H1238 [H1238], NCI-H1341 [H1341], NCI-H1385 [H1385], NCI-H1417 [H1417], NCI-H1435 [H1435], NCI-H1436 [H1436], NCI-H1437 [H1437], NCI-H1522 [H1522], NCI-H1563 [H1563], NCI-H1568 [H1568], NCI-H1573 [H1573], NCI-H1581 [H1581], NCI-H1618 [H1618], NCI-H1623 [H1623], NCI-H1650 [H-1650, H1650], NCI-H1651 [H1651], NCI-H1666 [H-1666, H1666], NCI-H1672 [H1672], NCI-H1693 [H1693], NCI-H1694 [H1694], NCI-H1703 [H1703], NCI-H1734 [H-1734, H1734], NCI-H1755 [H1755], NCI-H1755 [H1755], NCI-H1770 [H1770], NCI-H1793 [H1793], NCI-H1836 [H1836], NCI-H1838 [H1838], NCI-H1869 [H1869], NCI-H1876 [H1876], NCI-H1882 [H1882], NCI-H1915 [H1915], NCI-H1930 [H1930], NCI-H1944 [H1944], NCI-H1975 [H-1975, H1975], NCI-H1993 [H1993], NCI-H2023 [H2023], NCI-H2029 [H2029], NCI-H2030 [H2030], NCI-H2066 [H2066], NCI-H2073 [H2073], NCI-H2081 [H2081], NCI-H2085 [H2085], NCI-H2087 [H2087], NCI-H2106 [H2106], NCI-H2110 [H2110], NCI-H2135 [H2135], NCI-H2141 [H2141], NCI-H2171 [H2171], NCI-H2172 [H2172], NCI-H2195 [H2195], NCI-H2196 [H2196], NCI-H2198 [H2198], NCI-H2227 [H2227], NCI-H2228 [H2228], NCI-H2286 [H2286], NCI-H2291 [H2291], NCI-H2330 [H2330], NCI-H2342 [H2342], NCI-H2347 [H2347], NCI-H2405 [H2405], NCI-H2444 [H2444], UMC-11, NCI-H64 [H64], NCI-H735 [H735], NCI-H735 [H735], NCI-H1963 [H1963], NCI-H2107 [H2107], NCI-H2108 [H2108], NCI-H2122 [H2122], Hs 573.T, Hs 573.Lu, PLC/PRF/5, BEAS-2B, Hep G2, Tera-1, Tera-2, NCI-H69 [H69], NCI-H128 [H128], ChaGo-K-1, NCI-H446 [H446], NCI-H209 [H209], NCI-H146 [H146], NCI-H441 [H441], NCI-H82 [H82], NCI-H460 [H460], NCI-H596 [H596], NCI-H676B [H676B], NCI-H345 [H345], NCI-H820 [H820], NCI-H520 [H520], NCI-H661 [H661], NCI-H510A [H510A, NCI-H510], SK-HEP-1, A-427, Calu-1, Calu-3, Calu-6, SK-LU-1, SK-MES-1, SW 900 [SW-900, SW900], Malme-3M, and Capan-1.
In some embodiments, modified messenger RNA can be designed to incorporate microRNA binding region sites that either have 100% identity to known seed sequences or have less than 100% identity to seed sequences. The seed sequence can be partially mutated to decrease microRNA binding affinity and as such result in reduced downmodulation of that mRNA transcript. In essence, the degree of match or mis-match between the target mRNA and the microRNA seed can act as a rheostat to more finely tune the ability of the microRNA to modulate protein expression. In addition, mutation in the non-seed region of a microRNA binding site may also impact the ability of a microRNA to modulate protein expression.
In one embodiment, a miR sequence may be incorporated into the loop of a stem loop.
In another embodiment, a miR seed sequence may be incorporated in the loop of a stem loop and a miR binding site may be incorporated into the 5' or 3' stem of the stem loop.
In one embodiment, a TEE may be incorporated on the 5'end of the stem of a stem loop and a miR seed may be incorporated into the stem of the stem loop. In another embodiment, a TEE may be incorporated on the 5'end of the stem of a stem loop, a miR seed may be incorporated into the stem of the stem loop and a miR binding site may be incorporated into the 3'end of the stem or the sequence after the stem loop. The miR seed and the miR binding site may be for the same and/or different miR sequences.
In one embodiment, the incorporation of a miR sequence and/or a TEE sequence changes the shape of the stem loop region which may increase and/or decrease translation. (see e.g, Kedde et al. A Pumilio-induced RNA structure switch in p27-3'UTR controls miR-221 and miR-22 accessibility. Nature Cell Biology. 2010, incorporated herein by reference in its entirety).
In one embodiment, the incorporation of a miR sequence and/or a TEE sequence changes the shape of the stem loop region which may increase and/or decrease translation. (see e.g, Kedde et al. A Pumilio-induced RNA structure switch in p27-3'UTR controls miR-221 and miR-22 accessibility. Nature Cell Biology. 2010, incorporated herein by reference in its entirety).
In one embodiment, the 5'UTR may comprise at least one microRNA sequence. The microRNA sequence may be, but is not limited to, a 19 or 22 nucleotide sequence and/or a microRNA sequence without the seed.
In one embodiment the microRNA sequence in the 5'UTR may be used to stabilize the nucleic acid and/or mRNA described herein.
In another embodiment, a microRNA sequence in the 5'UTR may be used to decrease the accessibility of the site of translation initiation such as, but not limited to a start codon. Matsuda et al (PLoS One. 2010 11(5):e15057; incorporated herein by reference in its entirety) used antisense locked nucleic acid (LNA) oligonucleotides and exon-junction complexes (EJCs) around a start codon (-4 to +37 where the A of the AUG codons is +1) in order to decrease the accessibility to the first start codon (AUG). Matsuda showed that altering the sequence around the start codon with an LNA or EJC the efficiency, length and structural stability of the nucleic acid or mRNA is affected. The nucleic acids or mRNA of the present invention may comprise a microRNA sequence, instead of the LNA or EJC sequence described by Matsuda et al, near the site of translation initiation in order to decrease the accessibility to the site of translation initiation. The site of translation initiation may be prior to, after or within the microRNA sequence. As a non-limiting example, the site of translation initiation may be located within a microRNA sequence such as a seed sequence or binding site. As another non-limiting example, the site of translation initiation may be located within a miR-122 sequence such as the seed sequence or the mir-122 binding site.
In one embodiment, the nucleic acids or mRNA of the present invention may include at least one microRNA in order to dampen the antigen presentation by antigen presenting cells. The microRNA may be the complete microRNA sequence, the microRNA seed sequence, the microRNA sequence without the seed or a combination thereof. As a non-limiting example, the microRNA incorporated into the nucleic acids or mRNA of the present invention may be specific to the hematopoietic system. As another non-limiting example, the microRNA incorporated into the nucleic acids or mRNA of the present invention to dampen antigen presentation is miR-142-3p.
In one embodiment, the nucleic acids or mRNA of the present invention may include at least one microRNA in order to dampen expression of the encoded polypeptide in a cell of interest. As a non-limiting example, the nucleic acids or mRNA of the present invention may include at least one miR-122 binding site in order to dampen expression of an encoded polypeptide of interest in the liver. As another non-limiting example, the nucleic acids or mRNA of the present invention may include at least one miR-142-3p binding site, miR-142-3p seed sequence, miR-142-3p binding site without the seed, miR-142-5p binding site, miR-142-5p seed sequence, miR-142-5p binding site without the seed, miR-146 binding site, miR-146 seed sequence and/or miR-146 binding site without the seed sequence.
In one embodiment, the nucleic acids or mRNA of the present invention may comprise at least one microRNA binding site in the 3'UTR in order to selectively degrade mRNA therapeutics in the immune cells to subdue unwanted immunogenic reactions caused by therapeutic delivery. As a non-limiting example, the microRNA binding site may be the modified nucleic acids more unstable in antigen presenting cells. Non-limiting examples of these microRNA include mir-142-5p, mir-142-3p, mir-146a-5p and mir-146-3p.
In one embodiment, the nucleic acids or mRNA of the present invention comprises at least one microRNA sequence in a region of the nucleic acid or mRNA which may interact with a RNA binding protein.
RNA Motifs for RNA Binding Proteins (RBPs)
RNA binding proteins (RBPs) can regulate numerous aspects of co- and post-transcription gene expression such as, but not limited to, RNA splicing, localization, translation, turnover, polyadenylation, capping, modification, export and localization. RNA-binding domains (RBDs), such as, but not limited to, RNA recognition motif (RR) and hnRNP K-homology (KH) domains, typically regulate the sequence association between RBPs and their RNA targets (Ray et al. Nature 2013. 499:172-177; incorporated herein by reference in its entirety). In one embodiment, the canonical RBDs can bind short RNA sequences. In another embodiment, the canonical RBDs can recognize structure RNAs.
In one embodiment, to increase the stability of the mRNA of interest, an mRNA encoding HuR can be co-transfected or co-injected along with the mRNA of interest into the cells or into the tissue. These proteins can also be tethered to the mRNA of interest in vitro and then administered to the cells together. Poly A tail binding protein, PABP interacts with eukaryotic translation initiation factor eIF4G to stimulate translational initiation. Co-administration of mRNAs encoding these RBPs along with the mRNA drug and/or tethering these proteins to the mRNA drug in vitro and administering the protein-bound mRNA into the cells can increase the translational efficiency of the mRNA. The same concept can be extended to co-administration of mRNA along with mRNAs encoding various translation factors and facilitators as well as with the proteins themselves to influence RNA stability and/or translational efficiency.
In one embodiment, the nucleic acids and/or mRNA may comprise at least one RNA-binding motif such as, but not limited to a RNA-binding domain (RBD).
In one embodiment, the RBD may be any of the RBDs, fragments or variants thereof descried by Ray et al. (Nature 2013. 499:172-177; incorporated herein by reference in its entirety).
In one embodiment, the nucleic acids or mRNA of the present invention may comprise a sequence for at least one RNA-binding domain (RBDs). When the nucleic acids or mRNA of the present invention comprise more than one RBD, the RBDs do not need to be from the same species or even the same structural class.
In one embodiment, at least one flanking region (e.g., the 5'UTR and/or the 3'UTR) may comprise at least one RBD. In another embodiment, the first flanking region and the second flanking region may both comprise at least one RBD. The RBD may be the same or each of the RBDs may have at least 60% sequence identity to the other RBD. As a non-limiting example, at least on RBD may be located before, after and/or within the 3'UTR of the nucleic acid or mRNA of the present invention. As another non-limiting example, at least one RBD may be located before or within the first 300 nucleosides of the 3'UTR.
In another embodiment, the nucleic acids and/or mRNA of the present invention may comprise at least one RBD in the first region of linked nucleosides. The RBD may be located before, after or within a coding region (e.g., the ORF).
In yet another embodiment, the first region of linked nucleosides and/or at least one flanking region may comprise at least on RBD. As a non-limiting example, the first region of linked nucleosides may comprise a RBD related to splicing factors and at least one flanking region may comprise a RBD for stability and/or translation factors.
In one embodiment, the nucleic acids and/or mRNA of the present invention may comprise at least one RBD located in a coding and/or non-coding region of the nucleic acids and/or mRNA.
In one embodiment, at least one RBD may be incorporated into at least one flanking region to increase the stability of the nucleic acid and/or mRNA of the present invention.
In one embodiment, a microRNA sequence in a RNA binding protein motif may be used to decrease the accessibility of the site of translation initiation such as, but not limited to a start codon. The nucleic acids or mRNA of the present invention may comprise a microRNA sequence, instead of the LNA or EJC sequence described by Matsuda et al, near the site of translation initiation in order to decrease the accessibility to the site of translation initiation. The site of translation initiation may be prior to, after or within the microRNA sequence. As a non-limiting example, the site of translation initiation may be located within a microRNA sequence such as a seed sequence or binding site. As another non-limiting example, the site of translation initiation may be located within a miR-122 sequence such as the seed sequence or the mir-122 binding site.
In another embodiment, an antisense locked nucleic acid (LNA) oligonucleotides and exon-junction complexes (EJCs) may be used in the RNA binding protein motif. The LNA and EJCs may be used around a start codon (-4 to +37 where the A of the AUG codons is +1) in order to decrease the accessibility to the first start codon (AUG).
Codon Optimization
The polynucleotides of the invention, their regions or parts or subregions may be codon optimized. Codon optimization methods are known in the art and may be useful in efforts to achieve one or more of several goals. These goals include to match codon frequencies in target and host organisms to ensure proper folding, bias GC content to increase mRNA stability or reduce secondary structures, minimize tandem repeat codons or base runs that may impair gene construction or expression, customize transcriptional and translational control regions, insert or remove protein trafficking sequences, remove/add post translation modification sites in encoded protein (e.g., glycosylation sites), add, remove or shuffle protein domains, insert or delete restriction sites, modify ribosome binding sites and mRNA degradation sites, to adjust translational rates to allow the various domains of the protein to fold properly, or to reduce or eliminate problem secondary structures within the polynucleotide. Codon optimization tools, algorithms and services are known in the art, non-limiting examples include services from GeneArt (Life Technologies), DNA2.0 (Menlo Park Calif.) and/or proprietary methods. In one embodiment, the ORF sequence is optimized using optimization algorithms. Codon options for each amino acid are given in Table 5.
TABLE-US-00005 TABLE 5 Codon Options Single Letter Amino Acid Code Codon Options Isoleucine I ATT, ATC, ATA Leucine L CTT, CTC, CTA, CTG, TTA, TTG Valine V GTT, GTC, GTA, GTG Phenylalanine F TTT, TTC Methionine M ATG Cysteine C TGT, TGC Alanine A GCT, GCC, GCA, GCG Glycine G GGT, GGC, GGA, GGG Proline P CCT, CCC, CCA, CCG Threonine T ACT, ACC, ACA, ACG Serine S TCT, TCC, TCA, TCG, AGT, AGC Tyrosine Y TAT, TAC Tryptophan W TGG Glutamine Q CAA, CAG Asparagine N AAT, AAC Histidine H CAT, CAC Glutamic acid E GAA, GAG Aspartic acid D GAT, GAC Lysine K AAA, AAG Arginine R CGT, CGC, CGA, CGG, AGA, AGG Selenocysteine Sec UGA in mRNA in presence of Selenocystein insertion element (SECIS) Stop codons Stop TAA, TAG, TGA
"Codon optimized" refers to the modification of a starting nucleotide sequence by replacing at least one codon of the starting nucleotide sequence with a codon that is more frequently used in the group of abundant polypeptides of the host organism. Table 6 contains the codon usage frequency for humans (Codon usage database: [[www.]]kazusa.or.jp/codon/cgi-bin/showcodon.cgi?species=9606&aa=1&style=- N).
Codon optimization may be used to increase the expression of polypeptides by the replacement of at least one, at least two, at least three, at least four, at least five, at least six, at least seven, at least eight, at least nine, at least ten or at least 1%, at least 2%, at least 4%, at least 6%, at least 8%, at least 10%, at least 20%, at least 40%, at least 60%, at least 80%, at least 90% or at least 95%, or all codons of the starting nucleotide sequence with more frequently or the most frequently used codons for the respective amino acid as determined for the group of abundant proteins.
In one embodiment of the invention, the modified nucleotide sequences contain for each amino acid the most frequently used codons of the abundant proteins of the respective host cell.
TABLE-US-00006 TABLE 6 Codon usage frequency table for humans. Amino Amino Amino Amino Codon Acid % Codon Acid % Codon Acid % Codon Acid % UUU F 46 UCU S 19 UAU Y 44 UGU C 46 UUC F 54 UCC S 22 UAC Y 56 UGC C 54 UUA L 8 UCA S 15 UAA * 30 UGA * 47 UUG L 13 UCG S 5 UAG * 24 UGG W 100 CUU L 13 CCU P 29 CAU H 42 CGU R 8 CUC L 20 CCC P 32 CAC H 58 CGC R 18 CUA L 7 CCA P 28 CAA Q 27 CGA R 11 CUG L 40 CCG P 11 CAG Q 73 CGG R 20 AUU I 36 ACU T 25 AAU N 47 AGU S 15 AUC I 47 ACC T 36 AAC N 53 AGC S 24 AUA I 17 ACA T 28 AAA K 43 AGA R 21 AUG M 100 ACG T 11 AAG K 57 AGG R 21 GUU V 18 GCU A 27 GAU D 46 GGU G 16 GUC V 24 GCC A 40 GAC D 54 GGC G 34 GUA V 12 GCA A 23 GAA E 42 GGA G 25 GUG V 46 GCG A 11 GAG E 58 GGG G 25
In one embodiment, after a nucleotide sequence has been codon optimized it may be further evaluated for regions containing restriction sites. At least one nucleotide within the restriction site regions may be replaced with another nucleotide in order to remove the restriction site from the sequence but the replacement of nucleotides does alter the amino acid sequence which is encoded by the codon optimized nucleotide sequence.
Features, which may be considered beneficial in some embodiments of the present invention, may be encoded by regions of the polynucleotide and such regions may be upstream (5') or downstream (3') to a region which encodes a polypeptide. These regions may be incorporated into the polynucleotide before and/or after codon optimization of the protein encoding region or open reading frame (ORF). It is not required that a polynucleotide contain both a 5' and 3' flanking region. Examples of such features include, but are not limited to, untranslated regions (UTRs), Kozak sequences, an oligo(dT) sequence, and detectable tags and may include multiple cloning sites which may have XbaI recognition.
In some embodiments, a 5' UTR and/or a 3' UTR region may be provided as flanking regions. Multiple 5' or 3' UTRs may be included in the flanking regions and may be the same or of different sequences. Any portion of the flanking regions, including none, may be codon optimized and any may independently contain one or more different structural or chemical modifications, before and/or after codon optimization.
After optimization (if desired), the polynucleotides components are reconstituted and transformed into a vector such as, but not limited to, plasmids, viruses, cosmids, and artificial chromosomes. For example, the optimized polynucleotide may be reconstituted and transformed into chemically competent E. coli, yeast, neurospora, maize, drosophila, etc. where high copy plasmid-like or chromosome structures occur by methods described herein.
Uses of Alternative Polynucleotides
Therapeutic Agents
The alternative polynucleotides described herein can be used as therapeutic agents. For example, an alternative polynucleotide described herein can be administered to an animal or subject, wherein the alternative polynucleotide is translated in vivo to produce a therapeutic peptide in the animal or subject. Accordingly, provided herein are compositions, methods, kits, and reagents for treatment or prevention of disease or conditions in humans and other mammals. The active therapeutic agents of the present disclosure include alternative polynucleotides, cells containing alternative polynucleotides or polypeptides translated from the alternative polynucleotides, polypeptides translated from alternative polynucleotides, cells contacted with cells containing alternative polynucleotides or polypeptides translated from the alternative polynucleotides, tissues containing cells containing alternative polynucleotides and organs containing tissues containing cells containing alternative polynucleotides.
Provided are methods of inducing translation of a synthetic or recombinant polynucleotide to produce a polypeptide in a cell population using the alternative polynucleotides described herein. Such translation can be in vivo, ex vivo, in culture, or in vitro. The cell population is contacted with an effective amount of a composition containing a polynucleotide that has at least one nucleoside alternative, and a translatable region encoding the polypeptide. The population is contacted under conditions such that the polynucleotide is localized into one or more cells of the cell population and the recombinant polypeptide is translated in the cell from the polynucleotide.
An effective amount of the composition is provided based, at least in part, on the target tissue, target cell type, means of administration, physical characteristics of the polynucleotide (e.g., size, and extent of alternative nucleosides), and other determinants. In general, an effective amount of the composition provides efficient protein production in the cell, preferably more efficient than a composition containing a corresponding natural polynucleotide. Increased efficiency may be demonstrated by increased cell transfection (i.e., the percentage of cells transfected with the polynucleotide), increased protein translation from the polynucleotide, decreased polynucleotide degradation (as demonstrated, e.g., by increased duration of protein translation from an alternative polynucleotide), or reduced innate immune response of the host cell or improve therapeutic utility.
Aspects of the present disclosure are directed to methods of inducing in vivo translation of a recombinant polypeptide in a mammalian subject in need thereof. Therein, an effective amount of a composition containing a polynucleotide that has at least one alternative nucleoside and a translatable region encoding the polypeptide is administered to the subject using the delivery methods described herein. The polynucleotide is provided in an amount and under other conditions such that the polynucleotide is localized into a cell or cells of the subject and the recombinant polypeptide is translated in the cell from the polynucleotide. The cell in which the polynucleotide is localized, or the tissue in which the cell is present, may be targeted with one or more than one rounds of polynucleotide administration.
Other aspects of the present disclosure relate to transplantation of cells containing alternative polynucleotides to a mammalian subject. Administration of cells to mammalian subjects is known to those of ordinary skill in the art, such as local implantation (e.g., topical or subcutaneous administration), organ delivery or systemic injection (e.g., intravenous injection or inhalation), as is the formulation of cells in pharmaceutically acceptable carrier. Compositions containing alternative polynucleotides are formulated for administration intramuscularly, transarterially, intraperitoneally, intravenously, intranasally, subcutaneously, endoscopically, transdermally, or intrathecally. In some embodiments, the composition is formulated for extended release.
The subject to whom the therapeutic agent is administered suffers from or is at risk of developing a disease, disorder, or deleterious condition. Provided are methods of identifying, diagnosing, and classifying subjects on these bases, which may include clinical diagnosis, biomarker levels, genome-wide association studies (GWAS), and other methods known in the art.
In certain embodiments, the administered alternative polynucleotide directs production of one or more recombinant polypeptides that provide a functional activity which is substantially absent in the cell in which the recombinant polypeptide is translated. For example, the missing functional activity may be enzymatic, structural, or gene regulatory in nature.
In other embodiments, the administered alternative polynucleotide directs production of one or more recombinant polypeptides that replace a polypeptide (or multiple polypeptides) that is substantially absent in the cell in which the recombinant polypeptide is translated. Such absence may be due to genetic mutation of the encoding gene or regulatory pathway thereof. In other embodiments, the administered alternative polynucleotide directs production of one or more recombinant polypeptides to supplement the amount of polypeptide (or multiple polypeptides) that is present in the cell in which the recombinant polypeptide is translated. Alternatively, the recombinant polypeptide functions to antagonize the activity of an endogenous protein present in, on the surface of, or secreted from the cell. Usually, the activity of the endogenous protein is deleterious to the subject, for example, due to mutation of the endogenous protein resulting in altered activity or localization. Additionally, the recombinant polypeptide antagonizes, directly or indirectly, the activity of a biological moiety present in, on the surface of, or secreted from the cell. Examples of antagonized biological moieties include lipids (e.g., cholesterol), a lipoprotein (e.g., low density lipoprotein), a polynucleotide, a carbohydrate, or a small molecule toxin.
The recombinant proteins described herein are engineered for localization within the cell, potentially within a specific compartment such as the nucleus, or are engineered for secretion from the cell or translocation to the plasma membrane of the cell.
As described herein, a useful feature of the alternative polynucleotides of the present disclosure is the capacity to reduce, evade, avoid or eliminate the innate immune response of a cell to an exogenous polynucleotide. Provided are methods for performing the titration, reduction or elimination of the immune response in a cell or a population of cells. In some embodiments, the cell is contacted with a first composition that contains a first dose of a first exogenous polynucleotide including a translatable region and at least one alternative nucleoside, and the level of the innate immune response of the cell to the first exogenous polynucleotide is determined. Subsequently, the cell is contacted with a second composition, which includes a second dose of the first exogenous polynucleotide, the second dose containing a lesser amount of the first exogenous polynucleotide as compared to the first dose. Alternatively, the cell is contacted with a first dose of a second exogenous polynucleotide. The second exogenous polynucleotide may contain one or more alternative nucleosides, which may be the same or different from the first exogenous polynucleotide or, alternatively, the second exogenous polynucleotide may not contain alternative nucleosides. The steps of contacting the cell with the first composition and/or the second composition may be repeated one or more times. Additionally, efficiency of protein production (e.g., protein translation) in the cell is optionally determined, and the cell may be re-transfected with the first and/or second composition repeatedly until a target protein production efficiency is achieved.
Therapeutics for Diseases and Conditions
Provided are methods for treating or preventing a symptom of diseases characterized by missing or aberrant protein activity, by replacing the missing protein activity or overcoming the aberrant protein activity. Because of the rapid initiation of protein production following introduction of unnatural mRNAs, as compared to viral DNA vectors, the compounds of the present disclosure are particularly advantageous in treating acute diseases such as sepsis, stroke, and myocardial infarction. Moreover, the lack of transcriptional regulation of the unnatural mRNAs of the present disclosure is advantageous in that accurate titration of protein production is achievable. Multiple diseases are characterized by missing (or substantially diminished such that proper protein function does not occur) protein activity. Such proteins may not be present, are present in very low quantities or are essentially non-functional. The present disclosure provides a method for treating such conditions or diseases in a subject by introducing polynucleotide or cell-based therapeutics containing the alternative polynucleotides provided herein, wherein the alternative polynucleotides encode for a protein that replaces the protein activity missing from the target cells of the subject.
Diseases characterized by dysfunctional or aberrant protein activity include, but not limited to, cancer and proliferative diseases, genetic diseases (e.g., cystic fibrosis), autoimmune diseases, diabetes, neurodegenerative diseases, cardiovascular diseases, and metabolic diseases. The present disclosure provides a method for treating such conditions or diseases in a subject by introducing polynucleotide or cell-based therapeutics containing the alternative polynucleotides provided herein, wherein the alternative polynucleotides encode for a protein that antagonizes or otherwise overcomes the aberrant protein activity present in the cell of the subject.
Specific examples of a dysfunctional protein are the missense or nonsense mutation variants of the cystic fibrosis transmembrane conductance regulator (CFTR) gene, which produce a dysfunctional or nonfunctional, respectively, protein variant of CFTR protein, which causes cystic fibrosis.
Thus, provided are methods of treating cystic fibrosis in a mammalian subject by contacting a cell of the subject with an alternative polynucleotide having a translatable region that encodes a functional CFTR polypeptide, under conditions such that an effective amount of the CTFR polypeptide is present in the cell. Preferred target cells are epithelial cells, such as the lung, and methods of administration are determined in view of the target tissue; i.e., for lung delivery, the RNA molecules are formulated for administration by inhalation.
In another embodiment, the present disclosure provides a method for treating hyperlipidemia in a subject, by introducing into a cell population of the subject with an unnatural mRNA molecule encoding Sortilin, a protein recently characterized by genomic studies, thereby ameliorating the hyperlipidemia in a subject. The SORT1 gene encodes a trans-Golgi network (TGN) transmembrane protein called Sortilin. Genetic studies have shown that one of five individuals has a single nucleotide polymorphism, rs12740374, in the 1p13 locus of the SORT1 gene that predisposes them to having low levels of low-density lipoprotein (LDL) and very-low-density lipoprotein (VLDL). Each copy of the minor allele, present in about 30% of people, alters LDL cholesterol by 8 mg/dL, while two copies of the minor allele, present in about 5% of the population, lowers LDL cholesterol 16 mg/dL. Carriers of the minor allele have also been shown to have a 40% decreased risk of myocardial infarction. Functional in vivo studies in mice describes that overexpression of SORT1 in mouse liver tissue led to significantly lower LDL-cholesterol levels, as much as 80% lower, and that silencing SORT1 increased LDL cholesterol approximately 200% (Musunuru K et al. From noncoding variant to phenotype via SORT1 at the 1p13 cholesterol locus. Nature 2010; 466: 714-721).
Methods of Cellular Polynucleotide Delivery
Methods of the present disclosure enhance polynucleotide delivery into a cell population, in vivo, ex vivo, or in culture. For example, a cell culture containing a plurality of host cells (e.g., eukaryotic cells such as yeast or mammalian cells) is contacted with a composition that contains an enhanced polynucleotide having at least one nucleoside alternative and, optionally, a translatable region. The composition also generally contains a transfection reagent or other compound that increases the efficiency of enhanced polynucleotide uptake into the host cells. The enhanced polynucleotide exhibits enhanced retention in the cell population, relative to a corresponding natural polynucleotide. The retention of the enhanced polynucleotide is greater than the retention of the corresponding polynucleotide. In some embodiments, it is at least about 50%, 75%, 90%, 95%, 100%, 150%, 200% or more than 200% greater than the retention of the natural polynucleotide. Such retention advantage may be achieved by one round of transfection with the enhanced polynucleotide, or may be obtained following repeated rounds of transfection.
In some embodiments, the enhanced polynucleotide is delivered to a target cell population with one or more additional polynucleotides. Such delivery may be at the same time, or the enhanced polynucleotide is delivered prior to delivery of the one or more additional polynucleotides. The additional one or more polynucleotides may be alternative polynucleotides or natural polynucleotides. It is understood that the initial presence of the enhanced polynucleotides does not substantially induce an innate immune response of the cell population and, moreover, that the innate immune response will not be activated by the later presence of the natural polynucleotides. In this regard, the enhanced polynucleotide may not itself contain a translatable region, if the protein desired to be present in the target cell population is translated from the natural polynucleotides.
Targeting Moieties
In embodiments of the present disclosure, alternative polynucleotides are provided to express a protein-binding partner or a receptor on the surface of the cell, which functions to target the cell to a specific tissue space or to interact with a specific moiety, either in vivo or in vitro. Suitable protein-binding partners include antibodies and functional fragments thereof, scaffold proteins, or peptides. Additionally, alternative polynucleotides can be employed to direct the synthesis and extracellular localization of lipids, carbohydrates, or other biological moieties.
Permanent Gene Expression Silencing
Methods of the present disclosure include a method for epigenetically silencing gene expression in a mammalian subject, comprising a polynucleotide where the translatable region encodes a polypeptide or polypeptides capable of directing sequence-specific histone H3 methylation to initiate heterochromatin formation and reduce gene transcription around specific genes for the purpose of silencing the gene. For example, a gain-of-function mutation in the Janus Kinase 2 gene is responsible for the family of Myeloproliferative Diseases.
Delivery of a Detectable or Therapeutic Agent to a Biological Target
The alternative nucleosides, alternative nucleotides, and alternative polynucleotides described herein can be used in a number of different scenarios in which delivery of a substance (the "payload") to a biological target is desired, for example delivery of detectable substances for detection of the target, or delivery of a therapeutic agent. Detection methods can include both imaging in vitro and in vivo imaging methods, e.g., immunohistochemistry, bioluminescence imaging (BLI), Magnetic Resonance Imaging (MRI), positron emission tomography (PET), electron microscopy, X-ray computed tomography, Raman imaging, optical coherence tomography, absorption imaging, thermal imaging, fluorescence reflectance imaging, fluorescence microscopy, fluorescence molecular tomographic imaging, nuclear magnetic resonance imaging, X-ray imaging, ultrasound imaging, photoacoustic imaging, lab assays, or in any situation where tagging/staining/imaging is required.
For example, the alternative nucleosides, alternative nucleotides, and alternative polynucleotides described herein can be used in reprogramming induced pluripotent stem cells (iPS cells), which can then be used to directly track cells that are transfected compared to total cells in the cluster. In another example, a drug that is attached to the alternative polynucleotide via a linker and is fluorescently labeled can be used to track the drug in vivo, e.g. intracellularly. Other examples include the use of an alternative polynucleotide in reversible drug delivery into cells.
The alternative nucleosides, alternative nucleotides, and alternative polynucleotides described herein can be used in intracellular targeting of a payload, e.g., detectable or therapeutic agent, to specific organelle. Exemplary intracellular targets can include the nuclear localization for advanced mRNA processing, or a nuclear localization sequence (NLS) linked to the mRNA containing an inhibitor.
In addition, the alternative nucleosides, alternative nucleotides, and alternative nucleic acids described herein can be used to deliver therapeutic agents to cells or tissues, e.g., in living animals. For example, the alternative nucleosides, alternative nucleotides, and alternative nucleic acids described herein can be used to deliver highly polar chemotherapeutics agents to kill cancer cells. The alternative nucleic acids attached to the therapeutic agent through a linker can facilitate member permeation allowing the therapeutic agent to travel into a cell to reach an intracellular target.
In another example, the alternative nucleosides, alternative nucleotides, and alternative nucleic acids can be attached to a viral inhibitory peptide (VIP) through a cleavable linker. The cleavable linker will release the VIP and dye into the cell. In another example, the alternative nucleosides, alternative nucleotides, and alternative nucleic acids can be attached through the linker to a ADP-ribosylate, which is responsible for the actions of some bacterial toxins, such as cholera toxin, diphtheria toxin, and pertussis toxin. These toxin proteins are ADP-ribosyltransferases that modify target proteins in human cells. For example, cholera toxin ADP-ribosylates G proteins, causing massive fluid secretion from the lining of the small intestine, resulting in life-threatening diarrhea.
Pharmaceutical Compositions
The present disclosure provides proteins generated from unnatural mRNAs. Pharmaceutical compositions may optionally comprise one or more additional therapeutically active substances. In accordance with some embodiments, a method of administering pharmaceutical compositions comprising an alternative nucleic acids encoding one or more proteins to be delivered to a subject in need thereof is provided. In some embodiments, compositions are administered to humans. For the purposes of the present disclosure, the phrase "active ingredient" generally refers to a protein, protein encoding or protein-containing complex as described herein.
Although the descriptions of pharmaceutical compositions provided herein are principally directed to pharmaceutical compositions which are suitable for administration to humans, it will be understood by the skilled artisan that such compositions are generally suitable for administration to animals of all sorts. Modification of pharmaceutical compositions suitable for administration to humans in order to render the compositions suitable for administration to various animals is well understood, and the ordinarily skilled veterinary pharmacologist can design and/or perform such modification with merely ordinary, if any, experimentation. Subjects to which administration of the pharmaceutical compositions is contemplated include, but are not limited to, humans and/or other primates; mammals, including commercially relevant mammals such as cattle, pigs, horses, sheep, cats, dogs, mice, and/or rats; and/or birds, including commercially relevant birds such as chickens, ducks, geese, and/or turkeys.
Formulations of the pharmaceutical compositions described herein may be prepared by any method known or hereafter developed in the art of pharmacology. In general, such preparatory methods include the step of bringing the active ingredient into association with an excipient and/or one or more other accessory ingredients, and then, if necessary and/or desirable, shaping and/or packaging the product into a desired single- or multi-dose unit.
A pharmaceutical composition in accordance with the present disclosure may be prepared, packaged, and/or sold in bulk, as a single unit dose, and/or as a plurality of single unit doses. As used herein, a "unit dose" is discrete amount of the pharmaceutical composition comprising a predetermined amount of the active ingredient. The amount of the active ingredient is generally equal to the dosage of the active ingredient which would be administered to a subject and/or a convenient fraction of such a dosage such as, for example, one-half or one-third of such a dosage.
Relative amounts of the active ingredient, the pharmaceutically acceptable excipient, and/or any additional ingredients in a pharmaceutical composition in accordance with the present disclosure will vary, depending upon the identity, size, and/or condition of the subject treated and further depending upon the route by which the composition is to be administered. By way of example, the composition may comprise between 0.1% and 100% (w/w) active ingredient.
Pharmaceutical formulations may additionally comprise a pharmaceutically acceptable excipient, which, as used herein, includes any and all solvents, dispersion media, diluents, or other liquid vehicles, dispersion or suspension aids, surface active agents, isotonic agents, thickening or emulsifying agents, preservatives, solid binders, lubricants and the like, as suited to the particular dosage form desired. Remington's The Science and Practice of Pharmacy, 21.sup.st Edition, A. R. Gennaro (Lippincott, Williams & Wilkins, Baltimore, Md., 2006; incorporated herein by reference) discloses various excipients used in formulating pharmaceutical compositions and known techniques for the preparation thereof. Except insofar as any conventional excipient medium is incompatible with a substance or its derivatives, such as by producing any undesirable biological effect or otherwise interacting in a deleterious manner with any other component(s) of the pharmaceutical composition, its use is contemplated to be within the scope of this present disclosure.
In some embodiments, a pharmaceutically acceptable excipient is at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% pure. In some embodiments, an excipient is approved for use in humans and for veterinary use. In some embodiments, an excipient is approved by United States Food and Drug Administration. In some embodiments, an excipient is pharmaceutical grade. In some embodiments, an excipient meets the standards of the United States Pharmacopoeia (USP), the European Pharmacopoeia (EP), the British Pharmacopoeia, and/or the International Pharmacopoeia.
Pharmaceutically acceptable excipients used in the manufacture of pharmaceutical compositions include, but are not limited to, inert diluents, dispersing and/or granulating agents, surface active agents and/or emulsifiers, disintegrating agents, binding agents, preservatives, buffering agents, lubricating agents, and/or oils. Such excipients may optionally be included in pharmaceutical formulations. Excipients such as cocoa butter and suppository waxes, coloring agents, coating agents, sweetening, flavoring, and/or perfuming agents can be present in the composition, according to the judgment of the formulator.
Exemplary diluents include, but are not limited to, calcium carbonate, sodium carbonate, calcium phosphate, dicalcium phosphate, calcium sulfate, calcium hydrogen phosphate, sodium phosphate lactose, sucrose, cellulose, microcrystalline cellulose, kaolin, mannitol, sorbitol, inositol, sodium chloride, dry starch, cornstarch, powdered sugar, etc., and/or combinations thereof.
Exemplary granulating and/or dispersing agents include, but are not limited to, potato starch, corn starch, tapioca starch, sodium starch glycolate, clays, alginic acid, guar gum, citrus pulp, agar, bentonite, cellulose and wood products, natural sponge, cation-exchange resins, calcium carbonate, silicates, sodium carbonate, cross-linked poly(vinyl-pyrrolidone) (crospovidone), sodium carboxymethyl starch (sodium starch glycolate), carboxymethyl cellulose, cross-linked sodium carboxymethyl cellulose (croscarmellose), methylcellulose, pregelatinized starch (starch 1500), microcrystalline starch, water insoluble starch, calcium carboxymethyl cellulose, magnesium aluminum silicate (Veegum), sodium lauryl sulfate, quaternary ammonium compounds, etc., and/or combinations thereof.
Exemplary surface active agents and/or emulsifiers include, but are not limited to, natural emulsifiers (e.g. acacia, agar, alginic acid, sodium alginate, tragacanth, chondrux, cholesterol, xanthan, pectin, gelatin, egg yolk, casein, wool fat, cholesterol, wax, and lecithin), colloidal clays (e.g. bentonite [aluminum silicate] and Veegum.RTM. [magnesium aluminum silicate]), long chain amino acid derivatives, high molecular weight alcohols (e.g. stearyl alcohol, cetyl alcohol, oleyl alcohol, triacetin monostearate, ethylene glycol distearate, glyceryl monostearate, and propylene glycol monostearate, polyvinyl alcohol), carbomers (e.g. carboxy polymethylene, polyacrylic acid, acrylic acid polymer, and carboxyvinyl polymer), carrageenan, cellulosic derivatives (e.g. carboxymethylcellulose sodium, powdered cellulose, hydroxymethyl cellulose, hydroxypropyl cellulose, hydroxypropyl methylcellulose, methylcellulose), sorbitan fatty acid esters (e.g. polyoxyethylene sorbitan monolaurate [Tween.RTM.20], polyoxyethylene sorbitan [Tween.RTM.60], polyoxyethylene sorbitan monooleate [Tween.RTM.80], sorbitan monopalmitate [Span.RTM.40], sorbitan monostearate [Span.RTM.60], sorbitan tristearate [Span.RTM.65], glyceryl monooleate, sorbitan monooleate [Span.RTM.80]), polyoxyethylene esters (e.g. polyoxyethylene monostearate [Myrj.RTM.45], polyoxyethylene hydrogenated castor oil, polyethoxylated castor oil, polyoxymethylene stearate, and Solutol.RTM.), sucrose fatty acid esters, polyethylene glycol fatty acid esters (e.g. Cremophor.RTM.), polyoxyethylene ethers, (e.g. polyoxyethylene lauryl ether [Brij.RTM.30]), poly(vinyl-pyrrolidone), diethylene glycol monolaurate, triethanolamine oleate, sodium oleate, potassium oleate, ethyl oleate, oleic acid, ethyl laurate, sodium lauryl sulfate, Pluronic.RTM.F 68, Poloxamer.RTM.188, cetrimonium bromide, cetylpyridinium chloride, benzalkonium chloride, docusate sodium, etc. and/or combinations thereof.
Exemplary binding agents include, but are not limited to, starch (e.g. cornstarch and starch paste); gelatin; sugars (e.g. sucrose, glucose, dextrose, dextrin, molasses, lactose, lactitol, mannitol); natural and synthetic gums (e.g. acacia, sodium alginate, extract of Irish moss, panwar gum, ghatti gum, mucilage of isapol husks, carboxymethylcellulose, methylcellulose, ethylcellulose, hydroxyethylcellulose, hydroxypropyl cellulose, hydroxypropyl methylcellulose, microcrystalline cellulose, cellulose acetate, poly(vinyl-pyrrolidone), magnesium aluminum silicate (Veegum.RTM.), and larch arabogalactan); alginates; polyethylene oxide; polyethylene glycol; inorganic calcium salts; silicic acid; polymethacrylates; waxes; water; alcohol; etc.; and combinations thereof.
Exemplary preservatives may include, but are not limited to, antioxidants, chelating agents, antimicrobial preservatives, antifungal preservatives, alcohol preservatives, acidic preservatives, and/or other preservatives. Exemplary antioxidants include, but are not limited to, alpha tocopherol, ascorbic acid, acorbyl palmitate, butylated hydroxyanisole, butylated hydroxytoluene, monothioglycerol, potassium metabisulfite, propionic acid, propyl gallate, sodium ascorbate, sodium bisulfite, sodium metabisulfite, and/or sodium sulfite. Exemplary chelating agents include ethylenediaminetetraacetic acid (EDTA), citric acid monohydrate, disodium edetate, dipotassium edetate, edetic acid, fumaric acid, malic acid, phosphoric acid, sodium edetate, tartaric acid, and/or trisodium edetate. Exemplary antimicrobial preservatives include, but are not limited to, benzalkonium chloride, benzethonium chloride, benzyl alcohol, bronopol, cetrimide, cetylpyridinium chloride, chlorhexidine, chlorobutanol, chlorocresol, chloroxylenol, cresol, ethyl alcohol, glycerin, hexetidine, imidurea, phenol, phenoxyethanol, phenylethyl alcohol, phenylmercuric nitrate, propylene glycol, and/or thimerosal. Exemplary antifungal preservatives include, but are not limited to, butyl paraben, methyl paraben, ethyl paraben, propyl paraben, benzoic acid, hydroxybenzoic acid, potassium benzoate, potassium sorbate, sodium benzoate, sodium propionate, and/or sorbic acid. Exemplary alcohol preservatives include, but are not limited to, ethanol, polyethylene glycol, phenol, phenolic compounds, bisphenol, chlorobutanol, hydroxybenzoate, and/or phenylethyl alcohol. Exemplary acidic preservatives include, but are not limited to, vitamin A, vitamin C, vitamin E, beta-carotene, citric acid, acetic acid, dehydroacetic acid, ascorbic acid, sorbic acid, and/or phytic acid. Other preservatives include, but are not limited to, tocopherol, tocopherol acetate, deteroxime mesylate, cetrimide, butylated hydroxyanisol (BHA), butylated hydroxytoluened (BHT), ethylenediamine, sodium lauryl sulfate (SLS), sodium lauryl ether sulfate (SLES), sodium bisulfite, sodium metabisulfite, potassium sulfite, potassium metabisulfite, Glydant Plus.RTM., Phenonip.RTM., methylparaben, Germall.RTM.115, Germaben.RTM.II, Neolone.TM., Kathon.TM., and/or Euxyl.RTM..
Exemplary buffering agents include, but are not limited to, citrate buffer solutions, acetate buffer solutions, phosphate buffer solutions, ammonium chloride, calcium carbonate, calcium chloride, calcium citrate, calcium glubionate, calcium gluceptate, calcium gluconate, d-gluconic acid, calcium glycerophosphate, calcium lactate, propanoic acid, calcium levulinate, pentanoic acid, dibasic calcium phosphate, phosphoric acid, tribasic calcium phosphate, calcium hydroxide phosphate, potassium acetate, potassium chloride, potassium gluconate, potassium mixtures, dibasic potassium phosphate, monobasic potassium phosphate, potassium phosphate mixtures, sodium acetate, sodium bicarbonate, sodium chloride, sodium citrate, sodium lactate, dibasic sodium phosphate, monobasic sodium phosphate, sodium phosphate mixtures, tromethamine, magnesium hydroxide, aluminum hydroxide, alginic acid, pyrogen-free water, isotonic saline, Ringer's solution, ethyl alcohol, etc., and/or combinations thereof.
Exemplary lubricating agents include, but are not limited to, magnesium stearate, calcium stearate, stearic acid, silica, talc, malt, glyceryl behanate, hydrogenated vegetable oils, polyethylene glycol, sodium benzoate, sodium acetate, sodium chloride, leucine, magnesium lauryl sulfate, sodium lauryl sulfate, etc., and combinations thereof.
Exemplary oils include, but are not limited to, almond, apricot kernel, avocado, babassu, bergamot, black current seed, borage, cade, camomile, canola, caraway, carnauba, castor, cinnamon, cocoa butter, coconut, cod liver, coffee, corn, cotton seed, emu, eucalyptus, evening primrose, fish, flaxseed, geraniol, gourd, grape seed, hazel nut, hyssop, isopropyl myristate, jojoba, kukui nut, lavandin, lavender, lemon, litsea cubeba, macademia nut, mallow, mango seed, meadowfoam seed, mink, nutmeg, olive, orange, orange roughy, palm, palm kernel, peach kernel, peanut, poppy seed, pumpkin seed, rapeseed, rice bran, rosemary, safflower, sandalwood, sasquana, savoury, sea buckthorn, sesame, shea butter, silicone, soybean, sunflower, tea tree, thistle, tsubaki, vetiver, walnut, and wheat germ oils. Exemplary oils include, but are not limited to, butyl stearate, caprylic triglyceride, capric triglyceride, cyclomethicone, diethyl sebacate, dimethicone 360, isopropyl myristate, mineral oil, octyldodecanol, oleyl alcohol, silicone oil, and/or combinations thereof.
Liquid dosage forms for oral and parenteral administration include, but are not limited to, pharmaceutically acceptable emulsions, microemulsions, solutions, suspensions, syrups, and/or elixirs. In addition to active ingredients, liquid dosage forms may comprise inert diluents commonly used in the art such as, for example, water or other solvents, solubilizing agents and emulsifiers such as ethyl alcohol, isopropyl alcohol, ethyl carbonate, ethyl acetate, benzyl alcohol, benzyl benzoate, propylene glycol, 1,3-butylene glycol, dimethylformamide, oils (in particular, cottonseed, groundnut, corn, germ, olive, castor, and sesame oils), glycerol, tetrahydrofurfuryl alcohol, polyethylene glycols and fatty acid esters of sorbitan, and mixtures thereof. Besides inert diluents, oral compositions can include adjuvants such as wetting agents, emulsifying and suspending agents, sweetening, flavoring, and/or perfuming agents. In certain embodiments for parenteral administration, compositions are mixed with solubilizing agents such as Cremophor.RTM., alcohols, oils, modified oils, glycols, polysorbates, cyclodextrins, polymers, and/or combinations thereof.
Injectable preparations, for example, sterile injectable aqueous or oleaginous suspensions may be formulated according to the known art using suitable dispersing agents, wetting agents, and/or suspending agents. Sterile injectable preparations may be sterile injectable solutions, suspensions, and/or emulsions in nontoxic parenterally acceptable diluents and/or solvents, for example, as a solution in 1,3-butanediol. Among the acceptable vehicles and solvents that may be employed are water, Ringer's solution, U.S.P., and isotonic sodium chloride solution. Sterile, fixed oils are conventionally employed as a solvent or suspending medium. For this purpose any bland fixed oil can be employed including synthetic mono- or diglycerides. Fatty acids such as oleic acid can be used in the preparation of injectables.
Injectable formulations can be sterilized, for example, by filtration through a bacterial-retaining filter, and/or by incorporating sterilizing agents in the form of sterile solid compositions which can be dissolved or dispersed in sterile water or other sterile injectable medium prior to use.
In order to prolong the effect of an active ingredient, it is often desirable to slow the absorption of the active ingredient from subcutaneous or intramuscular injection. This may be accomplished by the use of a liquid suspension of crystalline or amorphous material with poor water solubility. The rate of absorption of the drug then depends upon its rate of dissolution which, in turn, may depend upon crystal size and crystalline form. Alternatively, delayed absorption of a parenterally administered drug form is accomplished by dissolving or suspending the drug in an oil vehicle. Injectable depot forms are made by forming microencapsule matrices of the drug in biodegradable polymers such as polylactide-polyglycolide. Depending upon the ratio of drug to polymer and the nature of the particular polymer employed, the rate of drug release can be controlled. Examples of other biodegradable polymers include poly(orthoesters) and poly(anhydrides). Depot injectable formulations are prepared by entrapping the drug in liposomes or microemulsions which are compatible with body tissues.
Compositions for rectal or vaginal administration are typically suppositories which can be prepared by mixing compositions with suitable non-irritating excipients such as cocoa butter, polyethylene glycol or a suppository wax which are solid at ambient temperature but liquid at body temperature and therefore melt in the rectum or vaginal cavity and release the active ingredient.
Solid dosage forms for oral administration include capsules, tablets, pills, powders, and granules. In such solid dosage forms, an active ingredient is mixed with at least one inert, pharmaceutically acceptable excipient such as sodium citrate or dicalcium phosphate and/or fillers or extenders (e.g. starches, lactose, sucrose, glucose, mannitol, and silicic acid), binders (e.g. carboxymethylcellulose, alginates, gelatin, polyvinylpyrrolidinone, sucrose, and acacia), humectants (e.g. glycerol), disintegrating agents (e.g. agar, calcium carbonate, potato or tapioca starch, alginic acid, certain silicates, and sodium carbonate), solution retarding agents (e.g. paraffin), absorption accelerators (e.g. quaternary ammonium compounds), wetting agents (e.g. cetyl alcohol and glycerol monostearate), absorbents (e.g. kaolin and bentonite clay), and lubricants (e.g. talc, calcium stearate, magnesium stearate, solid polyethylene glycols, sodium lauryl sulfate), and mixtures thereof. In the case of capsules, tablets and pills, the dosage form may comprise buffering agents.
Solid compositions of a similar type may be employed as fillers in soft and hard-filled gelatin capsules using such excipients as lactose or milk sugar as well as high molecular weight polyethylene glycols and the like. Solid dosage forms of tablets, dragees, capsules, pills, and granules can be prepared with coatings and shells such as enteric coatings and other coatings well known in the pharmaceutical formulating art. They may optionally comprise opacifying agents and can be of a composition that they release the active ingredient(s) only, or preferentially, in a certain part of the intestinal tract, optionally, in a delayed manner. Examples of embedding compositions which can be used include polymeric substances and waxes. Solid compositions of a similar type may be employed as fillers in soft and hard-filled gelatin capsules using such excipients as lactose or milk sugar as well as high molecular weight polyethylene glycols and the like.
Dosage forms for topical and/or transdermal administration of a composition may include ointments, pastes, creams, lotions, gels, powders, solutions, sprays, inhalants and/or patches. Generally, an active ingredient is admixed under sterile conditions with a pharmaceutically acceptable excipient and/or any needed preservatives and/or buffers as may be required. Additionally, the present disclosure contemplates the use of transdermal patches, which often have the added advantage of providing controlled delivery of a compound to the body. Such dosage forms may be prepared, for example, by dissolving and/or dispensing the compound in the proper medium. Alternatively or additionally, rate may be controlled by either providing a rate controlling membrane and/or by dispersing the compound in a polymer matrix and/or gel.
Suitable devices for use in delivering intradermal pharmaceutical compositions described herein include short needle devices such as those described in U.S. Pat. Nos. 4,886,499; 5,190,521; 5,328,483; 5,527,288; 4,270,537; 5,015,235; 5,141,496; and 5,417,662. Intradermal compositions may be administered by devices which limit the effective penetration length of a needle into the skin, such as those described in PCT publication WO 99/34850 and functional equivalents thereof. Jet injection devices which deliver liquid compositions to the dermis via a liquid jet injector and/or via a needle which pierces the stratum corneum and produces a jet which reaches the dermis are suitable. Jet injection devices are described, for example, in U.S. Pat. Nos. 5,480,381; 5,599,302; 5,334,144; 5,993,412; 5,649,912; 5,569,189; 5,704,911; 5,383,851; 5,893,397; 5,466,220; 5,339,163; 5,312,335; 5,503,627; 5,064,413; 5,520,639; 4,596,556; 4,790,824; 4,941,880; 4,940,460; and PCT publications WO 97/37705 and WO 97/13537. Ballistic powder/particle delivery devices which use compressed gas to accelerate vaccine in powder form through the outer layers of the skin to the dermis are suitable. Alternatively or additionally, conventional syringes may be used in the classical mantoux method of intradermal administration.
Formulations suitable for topical administration include, but are not limited to, liquid and/or semi liquid preparations such as liniments, lotions, oil in water and/or water in oil emulsions such as creams, ointments and/or pastes, and/or solutions and/or suspensions. Topically-administrable formulations may, for example, comprise from about 1% to about 10% (w/w) active ingredient, although the concentration of active ingredient may be as high as the solubility limit of the active ingredient in the solvent. Formulations for topical administration may further comprise one or more of the additional ingredients described herein.
A pharmaceutical composition may be prepared, packaged, and/or sold in a formulation suitable for pulmonary administration via the buccal cavity. Such a formulation may comprise dry particles which comprise the active ingredient and which have a diameter in the range from about 0.5 nm to about 7 nm or from about 1 nm to about 6 nm. Such compositions are conveniently in the form of dry powders for administration using a device comprising a dry powder reservoir to which a stream of propellant may be directed to disperse the powder and/or using a self propelling solvent/powder dispensing container such as a device comprising the active ingredient dissolved and/or suspended in a low-boiling propellant in a sealed container. Such powders comprise particles wherein at least 98% of the particles by weight have a diameter greater than 0.5 nm and at least 95% of the particles by number have a diameter less than 7 nm. Alternatively, at least 95% of the particles by weight have a diameter greater than 1 nm and at least 90% of the particles by number have a diameter less than 6 nm. Dry powder compositions may include a solid fine powder diluent such as sugar and are conveniently provided in a unit dose form.
Low boiling propellants generally include liquid propellants having a boiling point of below 65.degree. F. at atmospheric pressure. Generally the propellant may constitute 50% to 99.9% (w/w) of the composition, and active ingredient may constitute 0.1% to 20% (w/w) of the composition. A propellant may further comprise additional ingredients such as a liquid non-ionic and/or solid anionic surfactant and/or a solid diluent (which may have a particle size of the same order as particles comprising the active ingredient).
Pharmaceutical compositions formulated for pulmonary delivery may provide an active ingredient in the form of droplets of a solution and/or suspension. Such formulations may be prepared, packaged, and/or sold as aqueous and/or dilute alcoholic solutions and/or suspensions, optionally sterile, comprising active ingredient, and may conveniently be administered using any nebulization and/or atomization device. Such formulations may further comprise one or more additional ingredients including, but not limited to, a flavoring agent such as saccharin sodium, a volatile oil, a buffering agent, a surface active agent, and/or a preservative such as methylhydroxybenzoate. Droplets provided by this route of administration may have an average diameter in the range from about 0.1 nm to about 200 nm.
Formulations described herein as being useful for pulmonary delivery are useful for intranasal delivery of a pharmaceutical composition. Another formulation suitable for intranasal administration is a coarse powder comprising the active ingredient and having an average particle from about 0.2 .mu.m to 500 .mu.m. Such a formulation is administered in the manner in which snuff is taken, i.e. by rapid inhalation through the nasal passage from a container of the powder held close to the nose.
Formulations suitable for nasal administration may, for example, comprise from about as little as 0.1% (w/w) and as much as 100% (w/w) of active ingredient, and may comprise one or more of the additional ingredients described herein. A pharmaceutical composition may be prepared, packaged, and/or sold in a formulation suitable for buccal administration. Such formulations may, for example, be in the form of tablets and/or lozenges made using conventional methods, and may, for example, 0.1% to 20% (w/w) active ingredient, the balance comprising an orally dissolvable and/or degradable composition and, optionally, one or more of the additional ingredients described herein. Alternately, formulations suitable for buccal administration may comprise a powder and/or an aerosolized and/or atomized solution and/or suspension comprising active ingredient. Such powdered, aerosolized, and/or aerosolized formulations, when dispersed, may have an average particle and/or droplet size in the range from about 0.1 nm to about 200 nm, and may further comprise one or more of any additional ingredients described herein.
A pharmaceutical composition may be prepared, packaged, and/or sold in a formulation suitable for ophthalmic administration. Such formulations may, for example, be in the form of eye drops including, for example, a 0.1/1.0% (w/w) solution and/or suspension of the active ingredient in an aqueous or oily liquid excipient. Such drops may further comprise buffering agents, salts, and/or one or more other of any additional ingredients described herein. Other opthalmically-administrable formulations which are useful include those which comprise the active ingredient in microcrystalline form and/or in a liposomal preparation. Ear drops and/or eye drops are contemplated as being within the scope of this present disclosure.
General considerations in the formulation and/or manufacture of pharmaceutical agents may be found, for example, in Remington: The Science and Practice of Pharmacy 21.sup.st ed., Lippincott Williams & Wilkins, 2005 (incorporated herein by reference).
Administration
The present disclosure provides methods comprising administering proteins or complexes in accordance with the present disclosure to a subject in need thereof. Proteins or complexes, or pharmaceutical, imaging, diagnostic, or prophylactic compositions thereof, may be administered to a subject using any amount and any route of administration effective for preventing, treating, diagnosing, or imaging a disease, disorder, and/or condition (e.g., a disease, disorder, and/or condition relating to working memory deficits). The exact amount required will vary from subject to subject, depending on the species, age, and general condition of the subject, the severity of the disease, the particular composition, its mode of administration, its mode of activity, and the like. Compositions in accordance with the present disclosure are typically formulated in dosage unit form for ease of administration and uniformity of dosage. It will be understood, however, that the total daily usage of the compositions of the present disclosure will be decided by the attending physician within the scope of sound medical judgment. The specific therapeutically effective, prophylactically effective, or appropriate imaging dose level for any particular patient will depend upon a variety of factors including the disorder being treated and the severity of the disorder; the activity of the specific compound employed; the specific composition employed; the age, body weight, general health, sex and diet of the patient; the time of administration, route of administration, and rate of excretion of the specific compound employed; the duration of the treatment; drugs used in combination or coincidental with the specific compound employed; and like factors well known in the medical arts.
Proteins to be delivered and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof may be administered to animals, such as mammals (e.g., humans, domesticated animals, cats, dogs, mice, rats, etc.). In some embodiments, pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof are administered to humans.
Proteins to be delivered and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof in accordance with the present disclosure may be administered by any route. In some embodiments, proteins and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof, are administered by one or more of a variety of routes, including oral, intravenous, intramuscular, intra-arterial, intramedullary, intrathecal, subcutaneous, intraventricular, transdermal, interdermal, rectal, intravaginal, intraperitoneal, topical (e.g. by powders, ointments, creams, gels, lotions, and/or drops), mucosal, nasal, buccal, enteral, vitreal, intratumoral, sublingual; by intratracheal instillation, bronchial instillation, and/or inhalation; as an oral spray, nasal spray, and/or aerosol, and/or through a portal vein catheter. In some embodiments, proteins or complexes, and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof, are administered by systemic intravenous injection. In specific embodiments, proteins or complexes and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof may be administered intravenously and/or orally. In specific embodiments, proteins or complexes, and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof, may be administered in a way which allows the protein or complex to cross the blood-brain barrier, vascular barrier, or other epithelial barrier.
However, the present disclosure encompasses the delivery of proteins or complexes, and/or pharmaceutical, prophylactic, diagnostic, or imaging compositions thereof, by any appropriate route taking into consideration likely advances in the sciences of drug delivery.
In general the most appropriate route of administration will depend upon a variety of factors including the nature of the protein or complex comprising proteins associated with at least one agent to be delivered (e.g., its stability in the environment of the gastrointestinal tract, bloodstream, etc.), the condition of the patient (e.g., whether the patient is able to tolerate particular routes of administration), etc. The present disclosure encompasses the delivery of the pharmaceutical, prophylactic, diagnostic, or imaging compositions by any appropriate route taking into consideration likely advances in the sciences of drug delivery.
In certain embodiments, compositions in accordance with the present disclosure may be administered at dosage levels sufficient to deliver from about 0.0001 mg/kg to about 100 mg/kg, from about 0.01 mg/kg to about 50 mg/kg, from about 0.1 mg/kg to about 40 mg/kg, from about 0.5 mg/kg to about 30 mg/kg, from about 0.01 mg/kg to about 10 mg/kg, from about 0.1 mg/kg to about 10 mg/kg, or from about 1 mg/kg to about 25 mg/kg, of subject body weight per day, one or more times a day, to obtain the desired therapeutic, diagnostic, prophylactic, or imaging effect. The desired dosage may be delivered three times a day, two times a day, once a day, every other day, every third day, every week, every two weeks, every three weeks, or every four weeks. In certain embodiments, the desired dosage may be delivered using multiple administrations (e.g., two, three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, or more administrations).
Proteins or complexes may be used in combination with one or more other therapeutic, prophylactic, diagnostic, or imaging agents. By "in combination with," it is not intended to imply that the agents must be administered at the same time and/or formulated for delivery together, although these methods of delivery are within the scope of the present disclosure. Compositions can be administered concurrently with, prior to, or subsequent to, one or more other desired therapeutics or medical procedures. In general, each agent will be administered at a dose and/or on a time schedule determined for that agent. In some embodiments, the present disclosure encompasses the delivery of pharmaceutical, prophylactic, diagnostic, or imaging compositions in combination with agents that improve their bioavailability, reduce and/or modify their metabolism, inhibit their excretion, and/or modify their distribution within the body.
It will further be appreciated that therapeutically, prophylactically, diagnostically, or imaging active agents utilized in combination may be administered together in a single composition or administered separately in different compositions. In general, it is expected that agents utilized in combination with be utilized at levels that do not exceed the levels at which they are utilized individually. In some embodiments, the levels utilized in combination will be lower than those utilized individually.
The particular combination of therapies (therapeutics or procedures) to employ in a combination regimen will take into account compatibility of the desired therapeutics and/or procedures and the desired therapeutic effect to be achieved. It will also be appreciated that the therapies employed may achieve a desired effect for the same disorder (for example, a composition useful for treating cancer in accordance with the present disclosure may be administered concurrently with a chemotherapeutic agent), or they may achieve different effects (e.g., control of any adverse effects).
Kits
The present disclosure provides a variety of kits for conveniently and/or effectively carrying out methods of the present disclosure. Typically kits will comprise sufficient amounts and/or numbers of components to allow a user to perform multiple treatments of a subject(s) and/or to perform multiple experiments.
In one aspect, the disclosure provides kits for protein production, comprising a first isolated nucleic acid comprising a translatable region and an alternative nucleic acid, wherein the nucleic acid is capable of evading or avoiding induction of an innate immune response of a cell into which the first isolated nucleic acid is introduced, and packaging and instructions.
In one aspect, the disclosure provides kits for protein production, comprising: a first isolated alternative nucleic acid comprising a translatable region, provided in an amount effective to produce a desired amount of a protein encoded by the translatable region when introduced into a target cell; a second nucleic acid comprising an inhibitory nucleic acid, provided in an amount effective to substantially inhibit the innate immune response of the cell; and packaging and instructions.
In one aspect, the disclosure provides kits for protein production, comprising a first isolated nucleic acid comprising a translatable region and an alternative sugar, wherein the nucleic acid exhibits reduced degradation by a cellular nuclease, and packaging and instructions.
In one aspect, the disclosure provides kits for protein production, comprising a first isolated nucleic acid comprising a translatable region and at least two different alternative sugars, wherein the nucleic acid exhibits reduced degradation by a cellular nuclease, and packaging and instructions.
In one aspect, the disclosure provides kits for protein production, comprising a first isolated nucleic acid comprising a translatable region and at least one alternative sugar, wherein the nucleic acid exhibits reduced degradation by a cellular nuclease; a second nucleic acid comprising an inhibitory nucleic acid; and packaging and instructions.
In another aspect, the disclosure provides compositions for protein production, comprising a first isolated nucleic acid comprising a translatable region and a sugar alternative, wherein the nucleic acid exhibits reduced degradation by a cellular nuclease, and a mammalian cell suitable for translation of the translatable region of the first nucleic acid.
Definitions
About: As used herein, the term "about" means+/-10% of the recited value.
Administered in combination: As used herein, the term "administered in combination" or "combined administration" means that two or more agents are administered to a subject at the same time or within an interval such that there may be an overlap of an effect of each agent on the patient. In some embodiments, they are administered within about 60, 30, 15, 10, 5, or 1 minute of one another. In some embodiments, the administrations of the agents are spaced sufficiently closely together such that a combinatorial (e.g., a synergistic) effect is achieved.
Animal: As used herein, the term "animal" refers to any member of the animal kingdom. In some embodiments, "animal" refers to humans at any stage of development. In some embodiments, "animal" refers to non-human animals at any stage of development. In certain embodiments, the non-human animal is a mammal (e.g., a rodent, a mouse, a rat, a rabbit, a monkey, a dog, a cat, a sheep, cattle, a primate, or a pig). In some embodiments, animals include, but are not limited to, mammals, birds, reptiles, amphibians, fish, and worms. In some embodiments, the animal is a transgenic animal, genetically-engineered animal, or a clone.
Antigens of interest or desired antigens: As used herein, the terms "antigens of interest" or "desired antigens" include those proteins and other biomolecules provided herein that are immunospecifically bound by the antibodies and fragments, mutants, variants, and alterations thereof described herein. Examples of antigens of interest include, but are not limited to, insulin, insulin-like growth factor, hGH, tPA, cytokines, such as interleukins (IL), e.g., IL-1, IL-2, IL-3, IL-4, IL-5, IL-6, IL-7, IL-8, IL-9, IL-10, IL-11, IL-12, IL-13, IL-14, IL-15, IL-16, IL-17, IL-18, interferon (IFN) alpha, IFN beta, IFN gamma, IFN omega or IFN tau, tumor necrosis factor (TNF), such as TNF alpha and TNF beta, TNF gamma, TRAIL; G-CSF, GM-CSF, M-CSF, MCP-1 and VEGF.
Approximately: As used herein, the term "approximately" or "about," as applied to one or more values of interest, refers to a value that is similar to a stated reference value. In certain embodiments, the term "approximately" or "about" refers to a range of values that fall within 25%, 20%, 19%, 18%, 17%, 16%, 15%, 14%, 13%, 12%, 11%, 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3%, 2%, 1%, or less in either direction (greater than or less than) of the stated reference value unless otherwise stated or otherwise evident from the context (except where such number would exceed 100% of a possible value).
Associated with: As used herein, the terms "associated with," "conjugated," "linked," "attached," and "tethered," when used with respect to two or more moieties, means that the moieties are physically associated or connected with one another, either directly or via one or more additional moieties that serves as a linking agent, to form a structure that is sufficiently stable so that the moieties remain physically associated under the conditions in which the structure is used, e.g., physiological conditions. An "association" need not be strictly through direct covalent chemical bonding. It may also suggest ionic or hydrogen bonding or a hybridization based connectivity sufficiently stable such that the "associated" entities remain physically associated.
Biocompatible: As used herein, the term "biocompatible" means compatible with living cells, tissues, organs or systems posing little to no risk of injury, toxicity or rejection by the immune system.
Biodegradable: As used herein, the term "biodegradable" means capable of being broken down into innocuous products by the action of living things.
Biologically active: As used herein, the phrase "biologically active" refers to a characteristic of any substance that has activity in a biological system and/or organism. For instance, a substance that, when administered to an organism, has a biological effect on that organism, is considered to be biologically active. In particular embodiments, a polynucleotide of the present invention may be considered biologically active if even a portion of the polynucleotide is biologically active or mimics an activity considered biologically relevant.
Conserved: As used herein, the term "conserved" refers to nucleotides or amino acid residues of a polynucleotide sequence or polypeptide sequence, respectively, that are those that occur unaltered in the same position of two or more sequences being compared. Nucleotides or amino acids that are relatively conserved are those that are conserved amongst more related sequences than nucleotides or amino acids appearing elsewhere in the sequences.
In some embodiments, two or more sequences are said to be "completely conserved" if they are 100% identical to one another. In some embodiments, two or more sequences are said to be "highly conserved" if they are at least 70% identical, at least 80% identical, at least 90% identical, or at least 95% identical to one another. In some embodiments, two or more sequences are said to be "highly conserved" if they are about 70% identical, about 80% identical, about 90% identical, about 95%, about 98%, or about 99% identical to one another. In some embodiments, two or more sequences are said to be "conserved" if they are at least 30% identical, at least 40% identical, at least 50% identical, at least 60% identical, at least 70% identical, at least 80% identical, at least 90% identical, or at least 95% identical to one another. In some embodiments, two or more sequences are said to be "conserved" if they are about 30% identical, about 40% identical, about 50% identical, about 60% identical, about 70% identical, about 80% identical, about 90% identical, about 95% identical, about 98% identical, or about 99% identical to one another. Conservation of sequence may apply to the entire length of an oligonucleotide or polypeptide or may apply to a portion, region or feature thereof.
Cyclic or Cyclized: As used herein, the term "cyclic" refers to the presence of a continuous loop. Cyclic molecules need not be circular, only joined to form an unbroken chain of subunits. Cyclic molecules such as the mRNA of the present invention may be single units or multimers or comprise one or more components of a complex or higher order structure.
Cytostatic: As used herein, "cytostatic" refers to inhibiting, reducing, suppressing the growth, division, or multiplication of a cell (e.g., a mammalian cell (e.g., a human cell)), bacterium, virus, fungus, protozoan, parasite, prion, or a combination thereof.
Cytotoxic: As used herein, "cytotoxic" refers to killing or causing injurious, toxic, or deadly effect on a cell (e.g., a mammalian cell (e.g., a human cell)), bacterium, virus, fungus, protozoan, parasite, prion, or a combination thereof.
Delivery: As used herein, "delivery" refers to the act or manner of delivering a compound, substance, entity, moiety, cargo or payload.
Delivery Agent: As used herein, "delivery agent" refers to any substance which facilitates, at least in part, the in vivo delivery of a polynucleotide to targeted cells.
Destabilized: As used herein, the term "destable," "destabilize," or "destabilizing region" means a region or molecule that is less stable than a starting, wild-type or native form of the same region or molecule.
Detectable label: As used herein, "detectable label" refers to one or more markers, signals, or moieties which are attached, incorporated or associated with another entity that is readily detected by methods known in the art including radiography, fluorescence, chemiluminescence, enzymatic activity, absorbance and the like. Detectable labels include radioisotopes, fluorophores, chromophores, enzymes, dyes, metal ions, ligands such as biotin, avidin, streptavidin and haptens, quantum dots, and the like. Detectable labels may be located at any position in the peptides or proteins disclosed herein. They may be within the amino acids, the peptides, or proteins, or located at the N- or C-termini.
Digest: As used herein, the term "digest" means to break apart into smaller pieces or components. When referring to polypeptides or proteins, digestion results in the production of peptides.
Distal: As used herein, the term "distal" means situated away from the center or away from a point or region of interest.
Effective amount of an agent: As used herein, is that amount sufficient to effect beneficial or desired results, for example, clinical results, and, as such, an "effective amount" depends upon the context in which it is being applied. For example, in the context of administering an agent that treats cancer, an effective amount of an agent is, for example, an amount sufficient to achieve treatment, as defined herein, of cancer, as compared to the response obtained without administration of the agent.
Encoded protein cleavage signal: As used herein, "encoded protein cleavage signal" refers to the nucleotide sequence which encodes a protein cleavage signal.
Engineered: As used herein, embodiments of the invention are "engineered" when they are designed to have a feature or property, whether structural or chemical, that varies from a starting point, wild type or native molecule.
Expression: As used herein, "expression" of a nucleic acid sequence refers to one or more of the following events: (1) production of an RNA template from a DNA sequence (e.g., by transcription); (2) processing of an RNA transcript (e.g., by splicing, editing, 5' cap formation, and/or 3' end processing); (3) translation of an RNA into a polypeptide or protein; and (4) post-translational modification of a polypeptide or protein.
Feature: As used herein, a "feature" refers to a characteristic, a property, or a distinctive element.
Formulation: As used herein, a "formulation" includes at least a polynucleotide and a delivery agent.
Fragment: A "fragment," as used herein, refers to a portion. For example, fragments of proteins may comprise polypeptides obtained by digesting full-length protein isolated from cultured cells.
Functional: As used herein, a "functional" biological molecule is a biological molecule in a form in which it exhibits a property and/or activity by which it is characterized.
Homology: As used herein, the term "homology" refers to the overall relatedness between polymeric molecules, e.g. between polynucleotide molecules (e.g. DNA molecules and/or RNA molecules) and/or between polypeptide molecules. In some embodiments, polymeric molecules are considered to be "homologous" to one another if their sequences are at least 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 99% identical or similar. The term "homologous" necessarily refers to a comparison between at least two sequences (polynucleotide or polypeptide sequences). In accordance with the invention, two polynucleotide sequences are considered to be homologous if the polypeptides they encode are at least about 50%, 60%, 70%, 80%, 90%, 95%, or even 99% for at least one stretch of at least about 20 amino acids. In some embodiments, homologous polynucleotide sequences are characterized by the ability to encode a stretch of at least 4-5 uniquely specified amino acids. For polynucleotide sequences less than 60 nucleotides in length, homology is determined by the ability to encode a stretch of at least 4-5 uniquely specified amino acids. In accordance with the invention, two protein sequences are considered to be homologous if the proteins are at least about 50%, 60%, 70%, 80%, or 90% identical for at least one stretch of at least about 20 amino acids.
Identity: As used herein, the term "identity" refers to the overall relatedness between polymeric molecules, e.g., between oligonucleotide molecules (e.g. DNA molecules and/or RNA molecules) and/or between polypeptide molecules. Calculation of the percent identity of two polynucleotide sequences, for example, can be performed by aligning the two sequences for optimal comparison purposes (e.g., gaps can be introduced in one or both of a first and a second polynucleotide sequences for optimal alignment and non-identical sequences can be disregarded for comparison purposes). In certain embodiments, the length of a sequence aligned for comparison purposes is at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, or 100% of the length of the reference sequence. The nucleotides at corresponding nucleotide positions are then compared. When a position in the first sequence is occupied by the same nucleotide as the corresponding position in the second sequence, then the molecules are identical at that position. The percent identity between the two sequences is a function of the number of identical positions shared by the sequences, taking into account the number of gaps, and the length of each gap, which needs to be introduced for optimal alignment of the two sequences. The comparison of sequences and determination of percent identity between two sequences can be accomplished using a mathematical algorithm. For example, the percent identity between two nucleotide sequences can be determined using methods such as those described in Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D. W., ed., Academic Press, New York, 1993; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; Computer Analysis of Sequence Data, Part I, Griffin, A. M., and Griffin, H. G., eds., Humana Press, New Jersey, 1994; and Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M Stockton Press, New York, 1991; each of which is incorporated herein by reference. For example, the percent identity between two nucleotide sequences can be determined using the algorithm of Meyers and Miller (CABIOS, 1989, 4:11-17), which has been incorporated into the ALIGN program (version 2.0) using a PAM120 weight residue table, a gap length penalty of 12 and a gap penalty of 4. The percent identity between two nucleotide sequences can, alternatively, be determined using the GAP program in the GCG software package using an NWSgapdna.CMP matrix. Methods commonly employed to determine percent identity between sequences include, but are not limited to those disclosed in Carillo, H., and Lipman, D., SIAM J Applied Math., 48:1073 (1988); incorporated herein by reference. Techniques for determining identity are codified in publicly available computer programs. Exemplary computer software to determine homology between two sequences include, but are not limited to, GCG program package, Devereux, J., et al., Nucleic Acids Research, 12(1), 387 (1984)), BLASTP, BLASTN, and FASTA Altschul, S. F. et al., J. Molec. Biol., 215, 403 (1990)).
Inhibit expression of a gene: As used herein, the phrase "inhibit expression of a gene" means to cause a reduction in the amount of an expression product of the gene. The expression product can be an RNA transcribed from the gene (e.g., an mRNA) or a polypeptide translated from an mRNA transcribed from the gene. Typically a reduction in the level of an mRNA results in a reduction in the level of a polypeptide translated therefrom. The level of expression may be determined using standard techniques for measuring mRNA or protein.
In vitro: As used herein, the term "in vitro" refers to events that occur in an artificial environment, e.g., in a test tube or reaction vessel, in cell culture, in a Petri dish, etc., rather than within an organism (e.g., animal, plant, or microbe).
In vivo: As used herein, the term "in vivo" refers to events that occur within an organism (e.g., animal, plant, or microbe or cell or tissue thereof).
Isolated: As used herein, the term "isolated" refers to a substance or entity that has been separated from at least some of the components with which it was associated (whether in nature or in an experimental setting). Isolated substances may have varying levels of purity in reference to the substances from which they have been associated. Isolated substances and/or entities may be separated from at least about 10%, about 20%, about 30%, about 40%, about 50%, about 60%, about 70%, about 80%, about 90%, or more of the other components with which they were initially associated. In some embodiments, isolated agents are more than about 80%, about 85%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or more than about 99% pure. As used herein, a substance is "pure" if it is substantially free of other components. Substantially isolated: By "substantially isolated" is meant that the compound is substantially separated from the environment in which it was formed or detected. Partial separation can include, for example, a composition enriched in the compound of the present disclosure. Substantial separation can include compositions containing at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 90%, at least about 95%, at least about 97%, or at least about 99% by weight of the compound of the present disclosure, or salt thereof. Methods for isolating compounds and their salts are routine in the art.
Linker: As used herein, a linker refers to a group of atoms, e.g., 10-1,000 atoms, and can be comprised of the atoms or groups such as, but not limited to, carbon, amino, alkylamino, oxygen, sulfur, sulfoxide, sulfonyl, carbonyl, and imine. The linker can be attached to an alternative nucleoside or nucleotide on the nucleobase or sugar moiety at a first end, and to a payload, e.g., a detectable or therapeutic agent, at a second end. The linker may be of sufficient length as to not interfere with incorporation into a polynucleotide sequence. The linker can be used for any useful purpose, such as to form multimers (e.g., through linkage of two or more polynucleotides) or conjugates, as well as to administer a payload, as described herein. Examples of chemical groups that can be incorporated into the linker include, but are not limited to, alkyl, alkenyl, alkynyl, amido, amino, ether, thioether, ester, alkylene, heteroalkylene, aryl, or heterocyclyl, each of which can be optionally substituted, as described herein. Examples of linkers include, but are not limited to, unsaturated alkanes, polyethylene glycols (e.g., ethylene or propylene glycol monomeric units, e.g., diethylene glycol, dipropylene glycol, triethylene glycol, tripropylene glycol, tetraethylene glycol, or tetraethylene glycol), and dextran polymers, Other examples include, but are not limited to, cleavable moieties within the linker, such as, for example, a disulfide bond (--S--S--) or an azo bond (--N.dbd.N--), which can be cleaved using a reducing agent or photolysis. Non-limiting examples of a selectively cleavable bond include an amido bond can be cleaved for example by the use of tris(2-carboxyethyl)phosphine (TCEP), or other reducing agents, and/or photolysis, as well as an ester bond can be cleaved for example by acidic or basic hydrolysis.
Naturally occurring: As used herein, "naturally occurring" means existing in nature without artificial aid.
Non-human vertebrate: As used herein, a "non human vertebrate" includes all vertebrates except Homo sapiens, including wild and domesticated species. Examples of non-human vertebrates include, but are not limited to, mammals, such as alpaca, banteng, bison, camel, cat, cattle, deer, dog, donkey, gayal, goat, guinea pig, horse, llama, mule, pig, rabbit, reindeer, sheep water buffalo, and yak.
Off-target: As used herein, "off target" refers to any unintended effect on any one or more target, gene, or cellular transcript.
Open reading frame: As used herein, "open reading frame" or "ORF" refers to a sequence which does not contain a stop codon in a given reading frame.
Operably linked: As used herein, the phrase "operably linked" refers to a functional connection between two or more molecules, constructs, transcripts, entities, moieties or the like.
Paratope: As used herein, a "paratope" refers to the antigen-binding site of an antibody.
Patient: As used herein, "patient" refers to a subject who may seek or be in need of treatment, requires treatment, is receiving treatment, will receive treatment, or a subject who is under care by a trained professional for a particular disease or condition.
Peptide: As used herein, "peptide" is less than or equal to 50 amino acids long, e.g., about 5, 10, 15, 20, 25, 30, 35, 40, 45, or 50 amino acids long.
Pharmaceutically acceptable: The phrase "pharmaceutically acceptable" is employed herein to refer to those compounds, materials, compositions, and/or dosage forms which are, within the scope of sound medical judgment, suitable for use in contact with the tissues of human beings and animals without excessive toxicity, irritation, allergic response, or other problem or complication, commensurate with a reasonable benefit/risk ratio.
Pharmaceutically acceptable excipients: The phrase "pharmaceutically acceptable excipient," as used herein, refers any ingredient other than the compounds described herein (for example, a vehicle capable of suspending or dissolving the active compound) and having the properties of being substantially nontoxic and non-inflammatory in a patient. Excipients may include, for example: antiadherents, antioxidants, binders, coatings, compression aids, disintegrants, dyes (colors), emollients, emulsifiers, fillers (diluents), film formers or coatings, flavors, fragrances, glidants (flow enhancers), lubricants, preservatives, printing inks, sorbents, suspensing or dispersing agents, sweeteners, and waters of hydration. Exemplary excipients include, but are not limited to: butylated hydroxytoluene (BHT), calcium carbonate, calcium phosphate (dibasic), calcium stearate, croscarmellose, crosslinked polyvinyl pyrrolidone, citric acid, crospovidone, cysteine, ethylcellulose, gelatin, hydroxypropyl cellulose, hydroxypropyl methylcellulose, lactose, magnesium stearate, maltitol, mannitol, methionine, methylcellulose, methyl paraben, microcrystalline cellulose, polyethylene glycol, polyvinyl pyrrolidone, povidone, pregelatinized starch, propyl paraben, retinyl palmitate, shellac, silicon dioxide, sodium carboxymethyl cellulose, sodium citrate, sodium starch glycolate, sorbitol, starch (corn), stearic acid, sucrose, talc, titanium dioxide, vitamin A, vitamin E, vitamin C, and xylitol.
Pharmaceutically acceptable salts: The present disclosure also includes pharmaceutically acceptable salts of the compounds described herein. As used herein, "pharmaceutically acceptable salts" refers to derivatives of the disclosed compounds wherein the parent compound is modified by converting an existing acid or base moiety to its salt form (e.g., by reacting the free base group with a suitable organic acid). Examples of pharmaceutically acceptable salts include, but are not limited to, mineral or organic acid salts of basic residues such as amines; alkali or organic salts of acidic residues such as carboxylic acids; and the like. Representative acid addition salts include acetate, adipate, alginate, ascorbate, aspartate, benzenesulfonate, benzoate, bisulfate, borate, butyrate, camphorate, camphorsulfonate, citrate, cyclopentanepropionate, digluconate, dodecylsulfate, ethanesulfonate, fumarate, glucoheptonate, glycerophosphate, hemisulfate, heptonate, hexanoate, hydrobromide, hydrochloride, hydroiodide, 2-hydroxy-ethanesulfonate, lactobionate, lactate, laurate, lauryl sulfate, malate, maleate, malonate, methanesulfonate, 2-naphthalenesulfonate, nicotinate, nitrate, oleate, oxalate, palmitate, pamoate, pectinate, persulfate, 3-phenylpropionate, phosphate, picrate, pivalate, propionate, stearate, succinate, sulfate, tartrate, thiocyanate, toluenesulfonate, undecanoate, valerate salts, and the like. Representative alkali or alkaline earth metal salts include sodium, lithium, potassium, calcium, magnesium, and the like, as well as nontoxic ammonium, quaternary ammonium, and amine cations, including, but not limited to ammonium, tetramethylammonium, tetraethylammonium, methylamine, dimethylamine, trimethylamine, triethylamine, ethylamine, and the like. The pharmaceutically acceptable salts of the present disclosure include the conventional non-toxic salts of the parent compound formed, for example, from non-toxic inorganic or organic acids. The pharmaceutically acceptable salts of the present disclosure can be synthesized from the parent compound which contains a basic or acidic moiety by conventional chemical methods. Generally, such salts can be prepared by reacting the free acid or base forms of these compounds with a stoichiometric amount of the appropriate base or acid in water or in an organic solvent, or in a mixture of the two; generally, nonaqueous media like ether, ethyl acetate, ethanol, isopropanol, or acetonitrile are preferred. Lists of suitable salts are found in Remington's Pharmaceutical Sciences, 17.sup.th ed., Mack Publishing Company, Easton, Pa., 1985, p. 1418, Pharmaceutical Salts: Properties, Selection, and Use, P. H. Stahl and C. G. Wermuth (eds.), Wiley-VCH, 2008, and Berge et al., Journal of Pharmaceutical Science, 66, 1-19 (1977), each of which is incorporated herein by reference in its entirety.
Pharmacokinetic: As used herein, "pharmacokinetic" refers to any one or more properties of a molecule or compound as it relates to the determination of the fate of substances administered to a living organism. Pharmacokinetics is divided into several areas including the extent and rate of absorption, distribution, metabolism and excretion. This is commonly referred to as ADME where: (A) Absorption is the process of a substance entering the blood circulation; (D) Distribution is the dispersion or dissemination of substances throughout the fluids and tissues of the body; (M) Metabolism (or Biotransformation) is the irreversible transformation of parent compounds into daughter metabolites; and (E) Excretion (or Elimination) refers to the elimination of the substances from the body. In rare cases, some drugs irreversibly accumulate in body tissue.
Pharmaceutically acceptable solvate: The term "pharmaceutically acceptable solvate," as used herein, means a compound of the invention wherein molecules of a suitable solvent are incorporated in the crystal lattice. A suitable solvent is physiologically tolerable at the dosage administered. For example, solvates may be prepared by crystallization, recrystallization, or precipitation from a solution that includes organic solvents, water, or a mixture thereof. Examples of suitable solvents are ethanol, water (for example, mono-, di-, and tri-hydrates), N-methylpyrrolidinone (NMP), dimethyl sulfoxide (DMSO), N,N'-dimethylformamide (DMF), N,N'-dimethylacetamide (DMAC), 1,3-dimethyl-2-imidazolidinone (DMEU), 1,3-dimethyl-3,4,5,6-tetrahydro-2-(1H)-pyrimidinone (DMPU), acetonitrile (ACN), propylene glycol, ethyl acetate, benzyl alcohol, 2-pyrrolidone, benzyl benzoate, and the like. When water is the solvent, the solvate is referred to as a "hydrate."
Physicochemical: As used herein, "physicochemical" means of or relating to a physical and/or chemical property.
Preventing: As used herein, the term "preventing" refers to partially or completely delaying onset of an infection, disease, disorder and/or condition; partially or completely delaying onset of one or more symptoms, features, or clinical manifestations of a particular infection, disease, disorder, and/or condition; partially or completely delaying onset of one or more symptoms, features, or manifestations of a particular infection, disease, disorder, and/or condition; partially or completely delaying progression from an infection, a particular disease, disorder and/or condition; and/or decreasing the risk of developing pathology associated with the infection, the disease, disorder, and/or condition.
Prodrug: The present disclosure also includes prodrugs of the compounds described herein. As used herein, "prodrugs" refer to any substance, molecule or entity which is in a form predicate for that substance, molecule or entity to act as a therapeutic upon chemical or physical alteration. Prodrugs may by covalently bonded or sequestered in some way and which release or are converted into the active drug moiety prior to, upon or after administered to a mammalian subject. Prodrugs can be prepared by modifying functional groups present in the compounds in such a way that the modifications are cleaved, either in routine manipulation or in vivo, to the parent compounds. Prodrugs include compounds wherein hydroxyl, amino, sulfhydryl, or carboxyl groups are bonded to any group that, when administered to a mammalian subject, cleaves to form a free hydroxyl, amino, sulfhydryl, or carboxyl group respectively. Preparation and use of prodrugs is discussed in T. Higuchi and V. Stella, "Pro-drugs as Novel Delivery Systems," Vol. 14 of the A.C.S. Symposium Series, and in Bioreversible Carriers in Drug Design, ed. Edward B. Roche, American Pharmaceutical Association and Pergamon Press, 1987, both of which are hereby incorporated by reference in their entirety.
Proliferate: As used herein, the term "proliferate" means to grow, expand or increase or cause to grow, expand or increase rapidly. "Proliferative" means having the ability to proliferate. "Anti-proliferative" means having properties counter to or inapposite to proliferative properties.
Protein cleavage site: As used herein, "protein cleavage site" refers to a site where controlled cleavage of the amino acid chain can be accomplished by chemical, enzymatic or photochemical means.
Protein cleavage signal: As used herein "protein cleavage signal" refers to at least one amino acid that flags or marks a polypeptide for cleavage.
Protein of interest: As used herein, the terms "proteins of interest" or "desired proteins" include those provided herein and fragments, mutants, variants, and alterations thereof.
Proximal: As used herein, the term "proximal" means situated nearer to the center or to a point or region of interest.
Purified: As used herein, "purify," "purified," "purification" means to make substantially pure or clear from unwanted components, material defilement, admixture or imperfection.
Sample: As used herein, the term "sample" or "biological sample" refers to a subset of its tissues, cells or component parts (e.g. body fluids, including but not limited to blood, mucus, lymphatic fluid, synovial fluid, cerebrospinal fluid, saliva, amniotic fluid, amniotic cord blood, urine, vaginal fluid and semen). A sample further may include a homogenate, lysate or extract prepared from a whole organism or a subset of its tissues, cells or component parts, or a fraction or portion thereof, including but not limited to, for example, plasma, serum, spinal fluid, lymph fluid, the external sections of the skin, respiratory, intestinal, and genitourinary tracts, tears, saliva, milk, blood cells, tumors, organs. A sample further refers to a medium, such as a nutrient broth or gel, which may contain cellular components, such as proteins or polynucleotide molecule.
Signal Sequences: As used herein, the phrase "signal sequences" refers to a sequence which can direct the transport or localization of a protein.
Significant or Significantly: As used herein, the terms "significant" or "significantly" are used synonymously with the term "substantially."
Single unit dose: As used herein, a "single unit dose" is a dose of any therapeutic administered in one dose/at one time/single route/single point of contact, i.e., single administration event.
Similarity: As used herein, the term "similarity" refers to the overall relatedness between polymeric molecules, e.g. between polynucleotide molecules (e.g. DNA molecules and/or RNA molecules) and/or between polypeptide molecules. Calculation of percent similarity of polymeric molecules to one another can be performed in the same manner as a calculation of percent identity, except that calculation of percent similarity takes into account conservative substitutions as is understood in the art.
Split dose: As used herein, a "split dose" is the division of single unit dose or total daily dose into two or more doses.
Stable: As used herein "stable" refers to a compound that is sufficiently robust to survive isolation to a useful degree of purity from a reaction mixture, and preferably capable of formulation into an efficacious therapeutic agent.
Stabilized: As used herein, the term "stabilize", "stabilized," "stabilized region" means to make or become stable.
Subject: As used herein, the term "subject" or "patient" refers to any organism to which a composition in accordance with the invention may be administered, e.g., for experimental, diagnostic, prophylactic, and/or therapeutic purposes. Typical subjects include animals (e.g., mammals such as mice, rats, rabbits, non-human primates, and humans) and/or plants.
Substantially: As used herein, the term "substantially" refers to the qualitative condition of exhibiting total or near-total extent or degree of a characteristic or property of interest. One of ordinary skill in the biological arts will understand that biological and chemical phenomena rarely, if ever, go to completion and/or proceed to completeness or achieve or avoid an absolute result. The term "substantially" is therefore used herein to capture the potential lack of completeness inherent in many biological and chemical phenomena.
Substantially equal: As used herein as it relates to time differences between doses, the term means plus/minus 2%.
Substantially simultaneously: As used herein and as it relates to plurality of doses, the term means within 2 seconds.
Suffering from: An individual who is "suffering from" a disease, disorder, and/or condition has been diagnosed with or displays one or more symptoms of a disease, disorder, and/or condition.
Susceptible to: An individual who is "susceptible to" a disease, disorder, and/or condition has not been diagnosed with and/or may not exhibit symptoms of the disease, disorder, and/or condition but harbors a propensity to develop a disease or its symptoms. In some embodiments, an individual who is susceptible to a disease, disorder, and/or condition (for example, cancer) may be characterized by one or more of the following: (1) a genetic mutation associated with development of the disease, disorder, and/or condition; (2) a genetic polymorphism associated with development of the disease, disorder, and/or condition; (3) increased and/or decreased expression and/or activity of a protein and/or polynucleotide associated with the disease, disorder, and/or condition; (4) habits and/or lifestyles associated with development of the disease, disorder, and/or condition; (5) a family history of the disease, disorder, and/or condition; and (6) exposure to and/or infection with a microbe associated with development of the disease, disorder, and/or condition. In some embodiments, an individual who is susceptible to a disease, disorder, and/or condition will develop the disease, disorder, and/or condition. In some embodiments, an individual who is susceptible to a disease, disorder, and/or condition will not develop the disease, disorder, and/or condition.
Synthetic: The term "synthetic" means produced, prepared, and/or manufactured by the hand of man. Synthesis of polynucleotides or polypeptides or other molecules of the present invention may be chemical or enzymatic.
Targeted Cells: As used herein, "targeted cells" refers to any one or more cells of interest. The cells may be found in vitro, in vivo, in situ or in the tissue or organ of an organism. The organism may be an animal, preferably a mammal, more preferably a human and most preferably a patient.
Therapeutic Agent: The term "therapeutic agent" refers to any agent that, when administered to a subject, has a therapeutic, diagnostic, and/or prophylactic effect and/or elicits a desired biological and/or pharmacological effect.
Therapeutically effective amount: As used herein, the term "therapeutically effective amount" means an amount of an agent to be delivered (e.g., polynucleotide, drug, therapeutic agent, diagnostic agent, prophylactic agent, etc.) that is sufficient, when administered to a subject suffering from or susceptible to an infection, disease, disorder, and/or condition, to treat, improve symptoms of, diagnose, prevent, and/or delay the onset of the infection, disease, disorder, and/or condition.
Therapeutically effective outcome: As used herein, the term "therapeutically effective outcome" means an outcome that is sufficient in a subject suffering from or susceptible to an infection, disease, disorder, and/or condition, to treat, improve symptoms of, diagnose, prevent, and/or delay the onset of the infection, disease, disorder, and/or condition.
Total daily dose: As used herein, a "total daily dose" is an amount given or prescribed in 24 hr period. It may be administered as a single unit dose.
Transcription factor: As used herein, the term "transcription factor" refers to a DNA-binding protein that regulates transcription of DNA into RNA, for example, by activation or repression of transcription. Some transcription factors effect regulation of transcription alone, while others act in concert with other proteins. Some transcription factor can both activate and repress transcription under certain conditions. In general, transcription factors bind a specific target sequence or sequences highly similar to a specific consensus sequence in a regulatory region of a target gene. Transcription factors may regulate transcription of a target gene alone or in a complex with other molecules.
Treating: As used herein, the term "treating" refers to partially or completely alleviating, ameliorating, improving, relieving, delaying onset of, inhibiting progression of, reducing severity of, and/or reducing incidence of one or more symptoms or features of a particular infection, disease, disorder, and/or condition. For example, "treating" cancer may refer to inhibiting survival, growth, and/or spread of a tumor. Treatment may be administered to a subject who does not exhibit signs of a disease, disorder, and/or condition and/or to a subject who exhibits only early signs of a disease, disorder, and/or condition for the purpose of decreasing the risk of developing pathology associated with the disease, disorder, and/or condition.
Equivalents and Scope
Those skilled in the art will recognize, or be able to ascertain using no more than routine experimentation, many equivalents to the specific embodiments in accordance with the invention described herein. The scope of the present invention is not intended to be limited to the above Description, but rather is as set forth in the appended claims.
In the claims, articles such as "a," "an," and "the" may mean one or more than one unless indicated to the contrary or otherwise evident from the context. Claims or descriptions that include "or" between one or more members of a group are considered satisfied if one, more than one, or all of the group members are present in, employed in, or otherwise relevant to a given product or process unless indicated to the contrary or otherwise evident from the context. The invention includes embodiments in which exactly one member of the group is present in, employed in, or otherwise relevant to a given product or process. The invention includes embodiments in which more than one, or all of the group members are present in, employed in, or otherwise relevant to a given product or process.
It is also noted that the term "comprising" is intended to be open and permits but does not require the inclusion of additional elements or steps. When the term "comprising" is used herein, the term "consisting of" is thus also encompassed and disclosed.
Where ranges are given, endpoints are included. Furthermore, it is to be understood that unless otherwise indicated or otherwise evident from the context and understanding of one of ordinary skill in the art, values that are expressed as ranges can assume any specific value or subrange within the stated ranges in different embodiments of the invention, to the tenth of the unit of the lower limit of the range, unless the context clearly dictates otherwise.
In addition, it is to be understood that any particular embodiment of the present invention that falls within the prior art may be explicitly excluded from any one or more of the claims. Since such embodiments are deemed to be known to one of ordinary skill in the art, they may be excluded even if the exclusion is not set forth explicitly herein. Any particular embodiment of the compositions of the invention (e.g., any polynucleotide or protein encoded thereby; any method of production; any method of use; etc.) can be excluded from any one or more claims, for any reason, whether or not related to the existence of prior art.
All cited sources, for example, references, publications, databases, database entries, and art cited herein, are incorporated into this application by reference, even if not expressly stated in the citation. In case of conflicting statements of a cited source and the instant application, the statement in the instant application shall control.
EXAMPLES
Example 1: PCR for cDNA Production
PCR procedures for the preparation of cDNA are performed using 2.times.KAPA HIFI.TM. HotStart ReadyMix by Kapa Biosystems (Woburn, Mass.). This system includes 2.times.KAPA ReadyMix12.5 .mu.l; Forward Primer (10 uM) 0.75 .mu.l; Reverse Primer (10 uM) 0.75 .mu.l; Template cDNA 100 ng; and dH.sub.2O diluted to 25.0 .mu.l. The reaction conditions are at 95.degree. C. for 5 min. and 25 cycles of 98.degree. C. for 20 sec, then 58.degree. C. for 15 sec, then 72.degree. C. for 45 sec, then 72.degree. C. for 5 min. then 4.degree. C. to termination.
The reverse primer of the instant invention incorporates a poly-T.sub.120 for a poly-A.sub.120 in the mRNA. Other reverse primers with longer or shorter poly-T tracts can be used to adjust the length of the poly-A tail in the mRNA.
The reaction is cleaned up using Invitrogen's PURELINK.TM. PCR Micro Kit (Carlsbad, Calif.) per manufacturer's instructions (up to 5 .mu.g). Larger reactions will require a cleanup using a product with a larger capacity. Following the cleanup, the cDNA is quantified using the NanoDrop and analyzed by agarose gel electrophoresis to confirm the cDNA is the expected size. The cDNA is then submitted for sequencing analysis before proceeding to the in vitro transcription reaction.
Example 2: In Vitro Transcription (IVT)
A. Materials and Methods
Unnatural mRNAs according to the invention are made using standard laboratory methods and materials for in vitro transcription with the exception that the nucleotide mix contains alternative nucleotides. The open reading frame (ORF) of the gene of interest may be flanked by a 5' untranslated region (UTR) containing a strong Kozak translational initiation signal and an alpha-globin 3' UTR terminating with an oligo(dT) sequence for templated addition of a polyA tail for mRNAs not incorporating adenosine analogs. Adenosine-containing mRNAs are synthesized without an oligo (dT) sequence to allow for post-transcription poly (A) polymerase poly-(A) tailing.
The ORF may also include various upstream or downstream additions (such as, but not limited to, .beta.-globin, tags, etc.) may be ordered from an optimization service such as, but limited to, DNA2.0 (Menlo Park, Calif.) and may contain multiple cloning sites which may have XbaI recognition. Upon receipt of the construct, it may be reconstituted and transformed into chemically competent E. coli.
For the present invention, NEB DH5-alpha Competent E. coli may be used. Transformations are performed according to NEB instructions using 100 ng of plasmid. The protocol is as follows:
Thaw a tube of NEB 5-alpha Competent E. coli cells on ice for 10 minutes.
Add 1-5 .mu.l containing 1 pg-100 ng of plasmid DNA to the cell mixture. Carefully flick the tube 4-5 times to mix cells and DNA. Do not vortex.
Place the mixture on ice for 30 minutes. Do not mix.
Heat shock at 42.degree. C. for exactly 30 seconds. Do not mix.
Place on ice for 5 minutes. Do not mix.
Pipette 950 .mu.l of room temperature SOC into the mixture.
Place at 37.degree. C. for 60 minutes. Shake vigorously (250 rpm) or rotate.
Warm selection plates to 37.degree. C.
Mix the cells thoroughly by flicking the tube and inverting.
Spread 50-100 .mu.l of each dilution onto a selection plate and incubate overnight at 37.degree. C. Alternatively, incubate at 30.degree. C. for 24-36 hours or 25.degree. C. for 48 hours.
A single colony is then used to inoculate 5 ml of LB growth media using the appropriate antibiotic and then allowed to grow (250 RPM, 37.degree. C.) for 5 hours. This is then used to inoculate a 200 ml culture medium and allowed to grow overnight under the same conditions.
To isolate the plasmid (up to 850 .mu.g), a maxi prep is performed using the Invitrogen PURELINK.TM. HiPure Maxiprep Kit (Carlsbad, Calif.), following the manufacturer's instructions.
In order to generate cDNA for In Vitro Transcription (IVT), the plasmid is first linearized using a restriction enzyme such as XbaI. A typical restriction digest with XbaI will comprise the following: Plasmid 1.0 .mu.g; 10.times. Buffer 1.0 .mu.l; XbaI 1.5 .mu.l; dH.sub.2O up to 10 .mu.l; incubated at 37.degree. C. for 1 hr. If performing at lab scale (<5 .mu.g), the reaction is cleaned up using Invitrogen's PURELINK.TM. PCR Micro Kit (Carlsbad, Calif.) per manufacturer's instructions. Larger scale purifications may need to be done with a product that has a larger load capacity such as Invitrogen's standard PURELINK.TM. PCR Kit (Carlsbad, Calif.). Following the cleanup, the linearized vector is quantified using the NanoDrop and analyzed to confirm linearization using agarose gel electrophoresis.
IVT Reaction
The in vitro transcription reaction generates mRNA containing alternative nucleotides or alternative RNA. The input nucleotide triphosphate (NTP) mix is made in-house using natural and unnatural NTPs.
A typical in vitro transcription reaction includes the following:
TABLE-US-00007 Template cDNA 1.0 .mu.g 10x transcription buffer (400 mM Tris-HCl pH 8.0, 2.0 .mu.l 190 mM MgCl2, 50 mM DTT, 10 mM Spermidine) Custom NTPs (25 mM each 7.2 .mu.l RNase Inhibitor 20 U T7 RNA polymerase 3000 U dH.sub.20 up to 20.0 .mu.l
Incubation at 37.degree. C. for 3 hr-5 hrs.
The crude IVT mix may be stored at 4.degree. C. overnight for cleanup the next day. 1 U of RNase-free DNase is then used to digest the original template. After 15 minutes of incubation at 37.degree. C., the mRNA is purified using Ambion's MEGACLEAR.TM. Kit (Austin, Tex.) following the manufacturer's instructions. This kit can purify up to 500 .mu.g of RNA. Following the cleanup, the RNA is quantified using the NanoDrop and analyzed by agarose gel electrophoresis to confirm the RNA is the proper size and that no degradation of the RNA has occurred.
The T7 RNA polymerase may be selected from, T7 RNA polymerase, T3 RNA polymerase and mutant polymerases such as, but not limited to, the novel polymerases able to incorporate alternative NTPs as well as those polymerases described by Liu (Esvelt et al. (Nature (2011) 472(7344):499-503 and U.S. Publication No. 20110177495) which recognize alternate promoters, Ellington (Chelliserrykattil and Ellington, Nature Biotechnology (2004) 22(9):1155-1160) describing a T7 RNA polymerase variant to transcribe 2'-O-methyl RNA and Sousa (Padilla and Sousa, Nucleic Acids Research (2002) 30(24): e128) describing a T7 RNA polymerase double mutant; herein incorporated by reference in their entireties.
B. Agarose Gel Electrophoresis of Unnatural mRNA
Individual unnatural mRNAs (200-400 ng in a 20 .mu.l volume) are loaded into a well on a non-denaturing 1.2% Agarose E-Gel (Invitrogen, Carlsbad, Calif.) and run for 12-15 minutes according to the manufacturer protocol.
C. Agarose Gel Electrophoresis of RT-PCR Products
Individual reverse transcribed-PCR products (200-400 ng) are loaded into a well of a non-denaturing 1.2% Agarose E-Gel (Invitrogen, Carlsbad, Calif.) and run for 12-15 minutes according to the manufacturer protocol.
D. Nanodrop Unnatural mRNA Quantification and UV Spectral Data
Unnatural mRNAs in TE buffer (1 .mu.l) are used for Nanodrop UV absorbance readings to quantitate the yield of each alternative mRNA from an in vitro transcription reaction (UV absorbance traces are not shown).
Example 3: Enzymatic Capping of mRNA
Capping of the mRNA is performed as follows where the mixture includes: IVT RNA 60 .mu.g-180 .mu.g and dH.sub.2O up to 72 .mu.l. The mixture is incubated at 65.degree. C. for 5 minutes to denature RNA, and then is transferred immediately to ice.
The protocol then involves the mixing of 10.times. Capping Buffer (0.5 M Tris-HCl (pH 8.0), 60 mM KCl, 12.5 mM MgCl.sub.2) (10.0 .mu.l); 20 mM GTP (5.0 .mu.l); 20 mM S-Adenosyl Methionine (2.5 .mu.l); RNase Inhibitor (100 U); 2'-O-Methyltransferase (400 U); Vaccinia capping enzyme (Guanylyl transferase) (40 U); dH.sub.2O (Up to 28 .mu.l); and incubation at 37.degree. C. for 30 minutes for 60 .mu.g RNA or up to 2 hours for 180 .mu.g of RNA.
The mRNA is then purified using Ambion's MEGACLEAR.TM. Kit (Austin, Tex.) following the manufacturer's instructions. Following the cleanup, the RNA is quantified using the NANODROP.TM. (ThermoFisher, Waltham, Mass.) and analyzed by agarose gel electrophoresis to confirm the RNA is the proper size and that no degradation of the RNA has occurred. The RNA product may also be sequenced by running a reverse-transcription-PCR to generate the cDNA for sequencing.
Example 4: 5'-Guanosine Capping
A. Materials and Methods
The cloning, gene synthesis and vector sequencing may be performed by DNA2.0 Inc. (Menlo Park, Calif.). The ORF is restriction digested using XbaI and used for cDNA synthesis using tailed- or tail-less-PCR. The tailed-PCR cDNA product is used as the template for the mRNA synthesis reaction using 25 mM each alternative nucleotide mix (all alternative nucleotides may be custom synthesized or purchased from TriLink Biotech, San Diego, Calif. except pyrrolo-C triphosphate which may be purchased from Glen Research, Sterling Va.; unmodified nucleotides are purchased from Epicenter Biotechnologies, Madison, Wis.) and CellScript MEGASCRIPT.TM. (Epicenter Biotechnologies, Madison, Wis.) complete mRNA synthesis kit.
The in vitro transcription reaction is run for 4 hours at 37.degree. C. Alternative mRNAs incorporating adenosine alternatives are poly (A) tailed using yeast Poly (A) Polymerase (Affymetrix, Santa Clara, Calif.). The PCR reaction uses HiFi PCR 2.times. MASTER MIX.TM. (Kapa Biosystems, Woburn, Mass.). Alternative mRNAs are post-transcriptionally capped using recombinant Vaccinia Virus Capping Enzyme (New England BioLabs, Ipswich, Mass.) and a recombinant 2'-O-methyltransferase (Epicenter Biotechnologies, Madison, Wis.) to generate the 5'-guanosine Cap1 structure. Cap 2 structure and Cap 2 structures may be generated using additional 2'-O-methyltransferases. The In vitro transcribed mRNA product is run on an agarose gel and visualized. Alternative mRNA may be purified with Ambion/Applied Biosystems (Austin, Tex.) MEGAClear RNA.TM. purification kit. The PCR uses PURELINK.TM. PCR purification kit (Invitrogen, Carlsbad, Calif.). The product is quantified on NANODROP.TM. UV Absorbance (ThermoFisher, Waltham, Mass.). Quality, UV absorbance quality and visualization of the product was performed on an 1.2% agarose gel. The product is resuspended in TE buffer.
B. 5' Capping Alternative Polynucleotide (mRNA) Structure
5'-capping of alternative mRNA may be completed concomitantly during the in vitro-transcription reaction using the following chemical RNA cap analogs to generate the 5'-guanosine cap structure according to manufacturer protocols: 3'-0-Me-m.sup.7G(5')ppp(5')G (the ARCA cap); G(5')ppp(5')A; G(5')ppp(5')G; m.sup.7G(5')ppp(5')A; m.sup.7G(5')ppp(5')G (New England BioLabs, Ipswich, Mass.). 5'-capping of alternative mRNA may be completed post-transcriptionally using a Vaccinia Virus Capping Enzyme to generate the "Cap 0" structure: m.sup.7G(5')ppp(5')G (New England BioLabs, Ipswich, Mass.). Cap 1 structure may be generated using both Vaccinia Virus Capping Enzyme and a 2'-O methyl-transferase to generate: m7G(5')ppp(5')G-2'-O-methyl. Cap 2 structure may be generated from the Cap 1 structure followed by the 2'-o-methylation of the 5'-antepenultimate nucleotide using a 2'-O methyl-transferase. Cap 3 structure may be generated from the Cap 2 structure followed by the 2'-o-methylation of the 5'-preantepenultimate nucleotide using a 2'-O methyl-transferase. Enzymes are preferably derived from a recombinant source.
When transfected into mammalian cells, the unnatural mRNAs have a stability of 12-18 hours or more than 18 hours, e.g., 24, 36, 48, 60, 72 or greater than 72 hours.
Example 5: PolyA Tailing Reaction
Without a poly-T in the cDNA, a poly-A tailing reaction must be performed before cleaning the final product. This is done by mixing Capped IVT RNA (100 .mu.l); RNase Inhibitor (20 U); 10.times. Tailing Buffer (0.5 M Tris-HCl (pH 8.0), 2.5 M NaCl, 100 mM MgCl.sub.2)(12.0 .mu.l); 20 mM ATP (6.0 .mu.l); Poly-A Polymerase (20 U); dH.sub.2O up to 123.5 .mu.l and incubation at 37.degree. C. for 30 min. If the poly-A tail is already in the transcript, then the tailing reaction may be skipped and proceed directly to cleanup with Ambion's MEGACLEAR.TM. kit (Austin, Tex.) (up to 500 .mu.g). Poly-A Polymerase is preferably a recombinant enzyme expressed in yeast.
For studies performed and described herein, the poly-A tail is encoded in the IVT template to comprise 160 nucleotides in length. However, it should be understood that the processivity or integrity of the poly-A tailing reaction may not always result in exactly 160 nucleotides. Hence poly-A tails of approximately 160 nucleotides, e.g, about 150-165, 155, 156, 157, 158, 159, 160, 161, 162, 163, 164 or 165 are within the scope of the invention.
Example 6: Method of Screening for Protein Expression
A. Electrospray Ionization
A biological sample which may contain proteins encoded by unnatural RNA administered to the subject is prepared and analyzed according to the manufacturer protocol for electrospray ionization (ESI) using 1, 2, 3 or 4 mass analyzers. A biologic sample may also be analyzed using a tandem ESI mass spectrometry system.
Patterns of protein fragments, or whole proteins, are compared to known controls for a given protein and identity is determined by comparison.
B. Matrix-Assisted Laser Desorption/Ionization
A biological sample which may contain proteins encoded by unnatural RNA administered to the subject is prepared and analyzed according to the manufacturer protocol for matrix-assisted laser desorption/ionization (MALDI).
Patterns of protein fragments, or whole proteins, are compared to known controls for a given protein and identity is determined by comparison.
C. Liquid Chromatography-Mass Spectrometry-Mass Spectrometry
A biological sample, which may contain proteins encoded by unnatural RNA, may be treated with a trypsin enzyme to digest the proteins contained within. The resulting peptides are analyzed by liquid chromatography-mass spectrometry-mass spectrometry (LC/MS/MS). The peptides are fragmented in the mass spectrometer to yield diagnostic patterns that can be matched to protein sequence databases via computer algorithms. The digested sample may be diluted to achieve 1 ng or less starting material for a given protein. Biological samples containing a simple buffer background (e.g. water or volatile salts) are amenable to direct in-solution digest; more complex backgrounds (e.g. detergent, non-volatile salts, glycerol) require an additional clean-up step to facilitate the sample analysis.
Patterns of protein fragments, or whole proteins, are compared to known controls for a given protein and identity is determined by comparison.
Example 7: Transfection
A. Reverse Transfection
For experiments performed in a 24-well collagen-coated tissue culture plate, Keratinocytes or other cells are seeded at a cell density of 1.times.10.sup.5. For experiments performed in a 96-well collagen-coated tissue culture plate, Keratinocytes are seeded at a cell density of 0.5.times.10.sup.5. For each alternative mRNA to be transfected, alternative mRNA: RNAIMAX.TM. are prepared as described and mixed with the cells in the multi-well plate within 6 hours of cell seeding before cells had adhered to the tissue culture plate.
B. Forward Transfection
In a 24-well collagen-coated tissue culture plate, cells are seeded at a cell density of 0.7.times.10.sup.5. For experiments performed in a 96-well collagen-coated tissue culture plate, keratinocytes, if used, are seeded at a cell density of 0.3.times.10.sup.5. Cells are then grown to a confluency of >70% for over 24 hours. For each alternative mrna to be transfected, alternative mrna: Rnaimax.TM. are prepared as described and transfected onto the cells in the multi-well plate over 24 hours after cell seeding and adherence to the tissue culture plate.
C. Translation Screen: ELISA
Cells are grown in EpiLife medium with Supplement S7 from Invitrogen at a confluence of >70%. Cells are reverse transfected with 300 ng of the indicated alternative mRNA complexed with RNAIMAX.TM. from Invitrogen. Alternatively, cells are forward transfected with 300 ng alternative mRNA complexed with RNAIMAX.TM. from Invitrogen. The RNA: RNAIMAX.TM. complex is formed by first incubating the RNA with Supplement-free EPILIFE.RTM. media in a 5.times. volumetric dilution for 10 minutes at room temperature.
In a second vial, RNAIMAX.TM. reagent is incubated with Supplement-free EPILIFE.RTM. Media in a 10.times. volumetric dilution for 10 minutes at room temperature. The RNA vial is then mixed with the RNAIMAX.TM. vial and incubated for 20-30 at room temperature before being added to the cells in a drop-wise fashion. Secreted polypeptide concentration in the culture medium is measured at 18 hours post-transfection for each of the unnatural mRNAs in triplicate. Secretion of the polypeptide of interest from transfected human cells is quantified using an ELISA kit from Invitrogen or R&D Systems (Minneapolis, Minn.) following the manufacturers recommended instructions.
D. Dose and Duration: ELISA
Cells are grown in EPILIFE.RTM. medium with Supplement S7 from Invitrogen at a confluence of >70%. Cells are reverse transfected with 0 ng, 46.875 ng, 93.75 ng, 187.5 ng, 375 ng, 750 ng, or 1500 ng alternative mRNA complexed with RNAIMAX.TM. from Invitrogen. The alternative mRNA: RNAIMAX.TM. complex is formed as described. Secreted polypeptide concentration in the culture medium is measured at 0, 6, 12, 24, and 48 hours post-transfection for each concentration of each alternative mRNA in triplicate. Secretion of the polypeptide of interest from transfected human cells is quantified using an ELISA kit from Invitrogen or R&D Systems following the manufacturers recommended instructions.
Example 8: Cellular Innate Immune Response: IFN-Beta ELISA and TNF-Alpha ELISA
An enzyme-linked immunosorbent assay (ELISA) for Human Tumor Necrosis Factor-.alpha. (TNF-.alpha.), Human Interferon-.beta. (IFN-.beta.) and Human Granulocyte-Colony Stimulating Factor (G-CSF) secreted from in vitro-transfected Human Keratinocyte cells is tested for the detection of a cellular innate immune response.
Cells are grown in EPILIFE.RTM. medium with Human Growth Supplement in the absence of hydrocortisone from Invitrogen at a confluence of >70%. Cells are reverse transfected with 0 ng, 93.75 ng, 187.5 ng, 375 ng, 750 ng, 1500 ng or 3000 ng of the indicated chemically alternative mRNA complexed with RNAIMAX.TM. from Invitrogen as described in triplicate. Secreted TNF-.alpha. in the culture medium is measured 24 hours post-transfection for each of the unnatural mRNAs using an ELISA kit from Invitrogen according to the manufacturer protocols.
Secreted IFN-.beta. is measured 24 hours post-transfection for each of the unnatural mRNAs using an ELISA kit from Invitrogen according to the manufacturer protocols. Secreted hu-G-CSF concentration is measured at 24 hours post-transfection for each of the alternative mRNAs. Secretion of the polypeptide of interest from transfected human cells is quantified using an ELISA kit from Invitrogen or R&D Systems (Minneapolis, Minn.) following the manufacturers recommended instructions. These data indicate which unnatural mRNA are capable eliciting a reduced cellular innate immune response in comparison to natural and other alternative polynucleotides or reference compounds by measuring exemplary type 1 cytokines such as TNF-alpha and IFN-beta.
Example 9: Cytotoxicity and Apoptosis
This experiment demonstrates cellular viability, cytotoxity and apoptosis for distinct alternative mRNA-in vitro transfected Human Keratinocyte cells. Keratinocytes are grown in EPILIFE.RTM. medium with Human Keratinocyte Growth Supplement in the absence of hydrocortisone from Invitrogen at a confluence of >70%. Keratinocytes are reverse transfected with 0 ng, 46.875 ng, 93.75 ng, 187.5 ng, 375 ng, 750 ng, 1500 ng, 3000 ng, or 6000 ng of unnatural mRNA complexed with RNAIMAX.TM. from Invitrogen. The unnatural mRNA: RNAIMAX.TM. complex is formed. Secreted huG-CSF concentration in the culture medium is measured at 0, 6, 12, 24, and 48 hours post-transfection for each concentration of each unnatural mRNA in triplicate. Secretion of the polypeptide of interest from transfected human keratinocytes is quantified using an ELISA kit from Invitrogen or R&D Systems following the manufacturers recommended instructions. Cellular viability, cytotoxicity and apoptosis is measured at 0, 12, 48, 96, and 192 hours post-transfection using the APOTOX-GLO.TM. kit from Promega (Madison, Wis.) according to manufacturer instructions.
OTHER EMBODIMENTS
It is to be understood that while the present disclosure has been described in conjunction with the detailed description thereof, the foregoing description is intended to illustrate and not limit the scope of the present disclosure, which is defined by the scope of the appended claims. Other aspects, advantages, and modifications are within the scope of the following claims.
SEQUENCE LISTINGS
1
4110DNAHomo sapiens 1tttttctttt 10211DNAHomo sapiens 2ttttgctttt t 11310DNAHomo sapiens 3ttttgctttt 10411DNAHomo sapiens 4aaaaagcaaa a 11
* * * * *
References
C00001

C00002

C00003

C00004

C00005

C00006

C00007

C00008

C00009

C00010

C00011

C00012

C00013

C00014
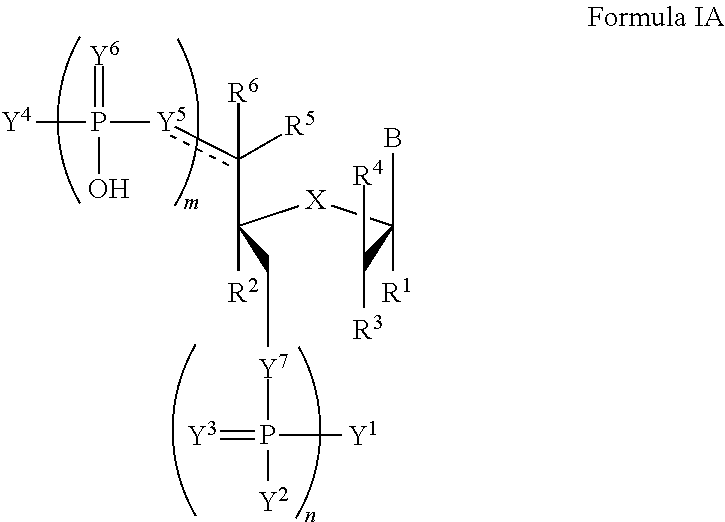
C00015

C00016

C00017

C00018

C00019
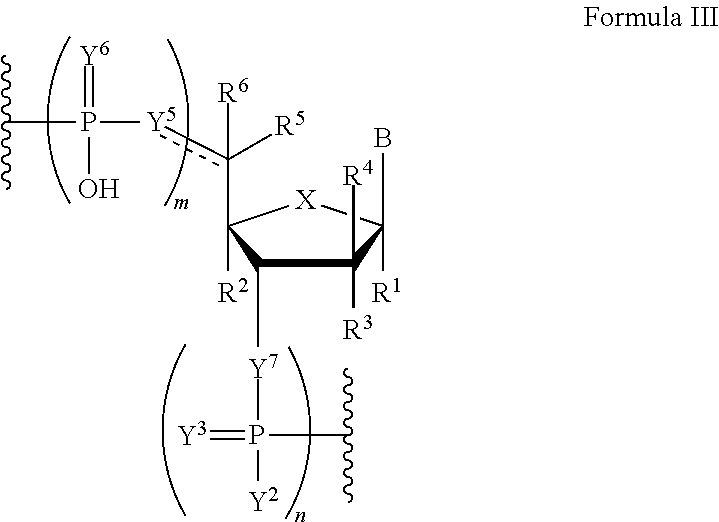
C00020
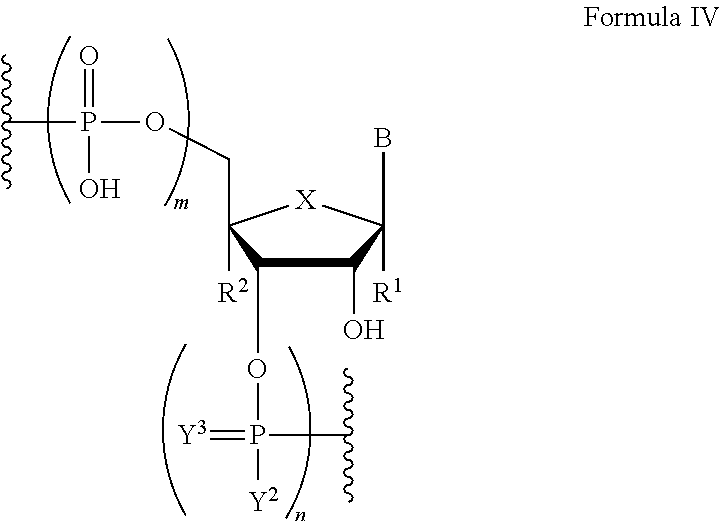
C00021

C00022
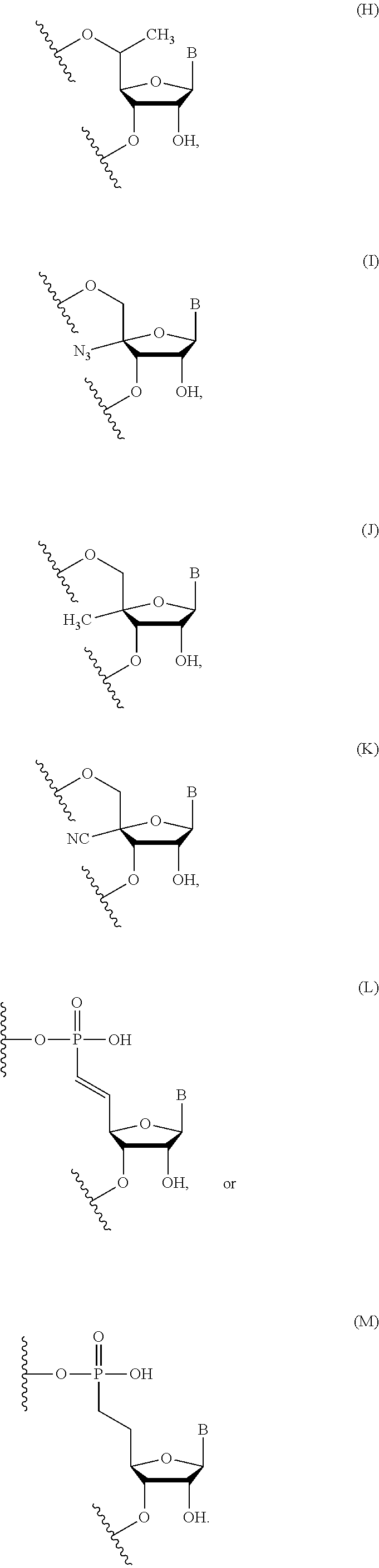
C00023
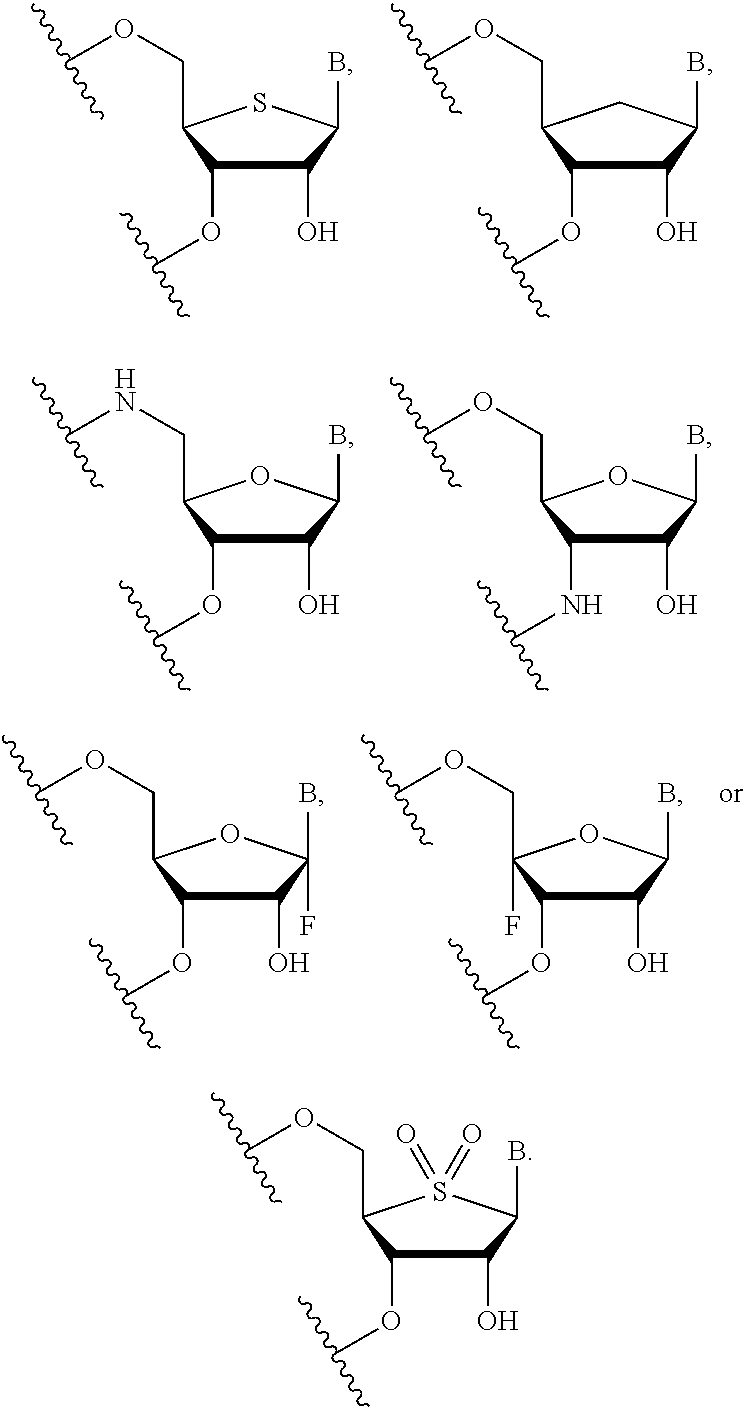
C00024
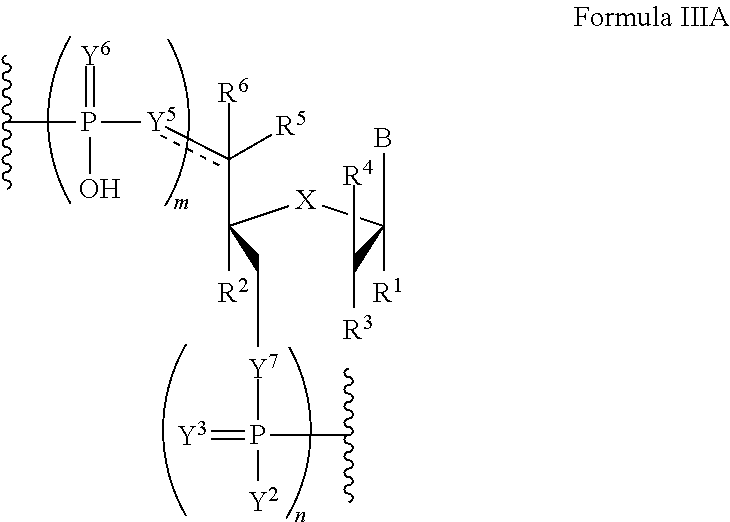
C00025

C00026

C00027
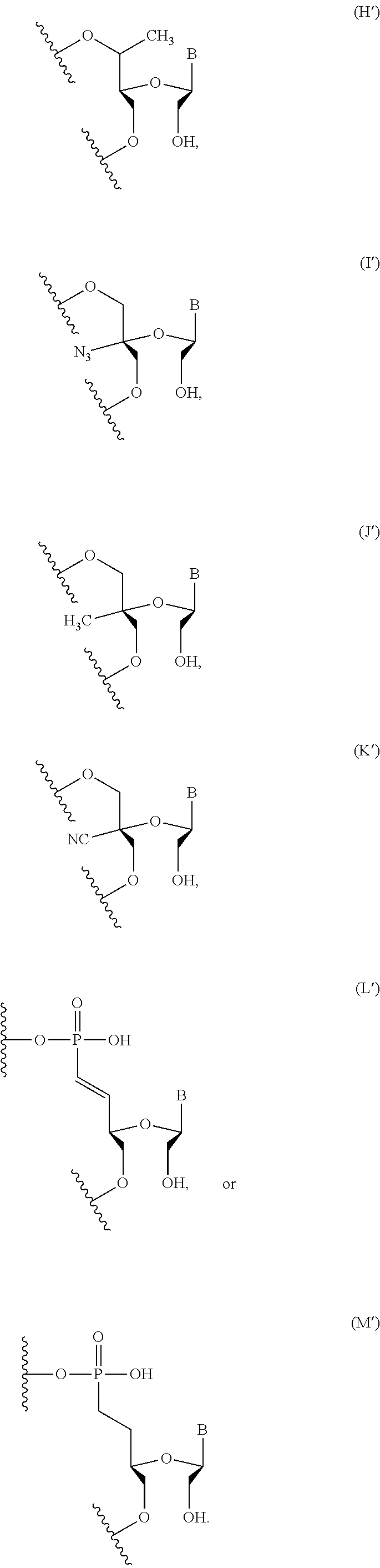
C00028

C00029
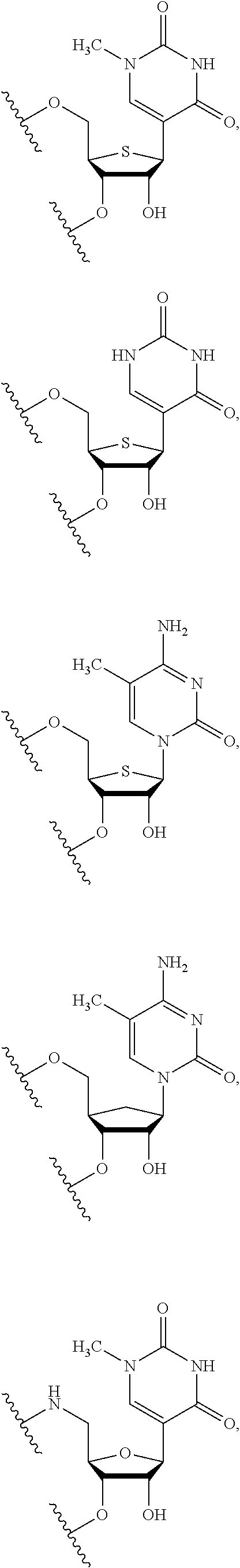
C00030

C00031
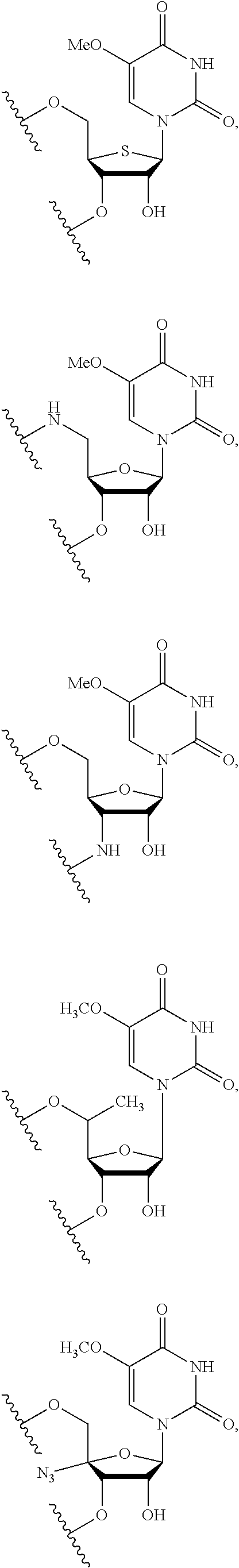
C00032
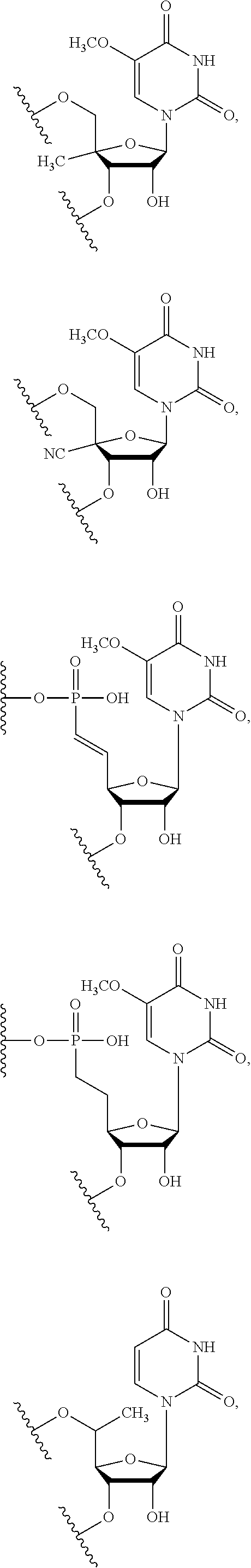
C00033

C00034

C00035

C00036
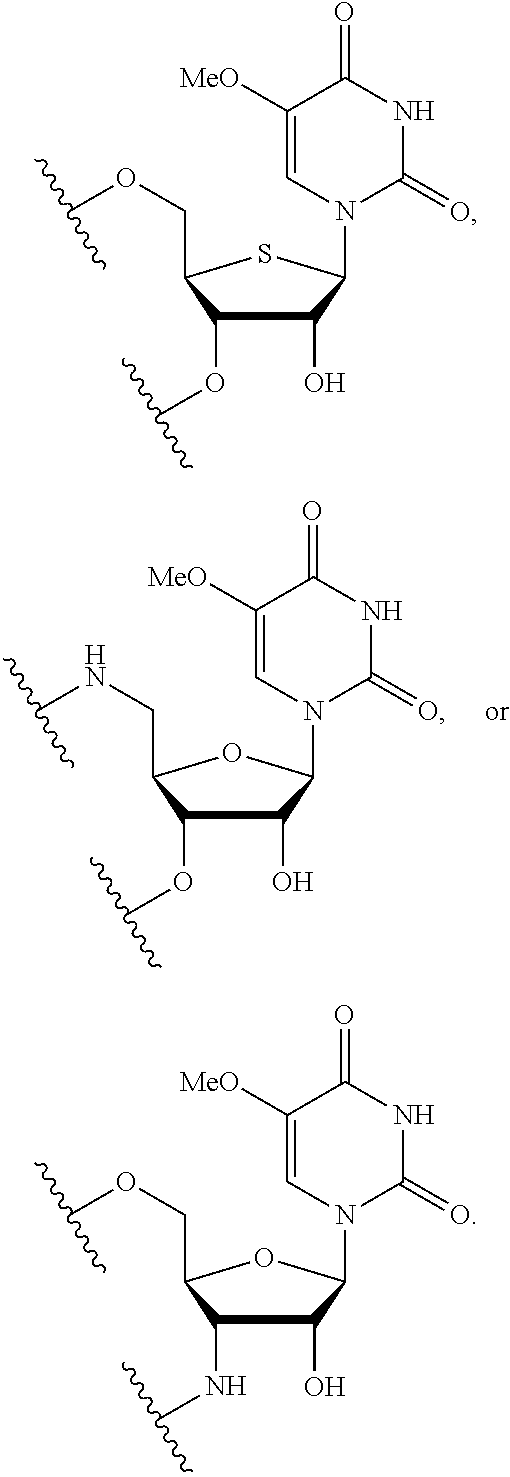
C00037
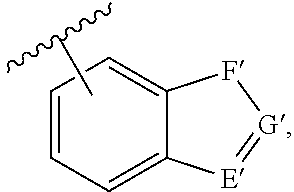
C00038
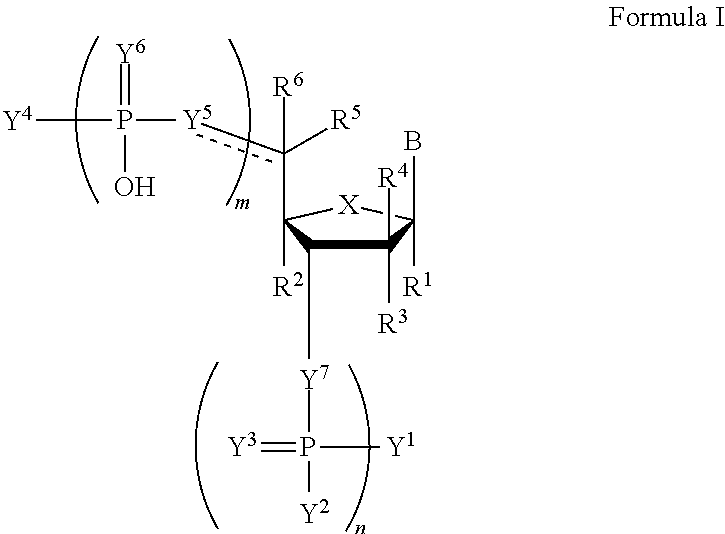
C00039
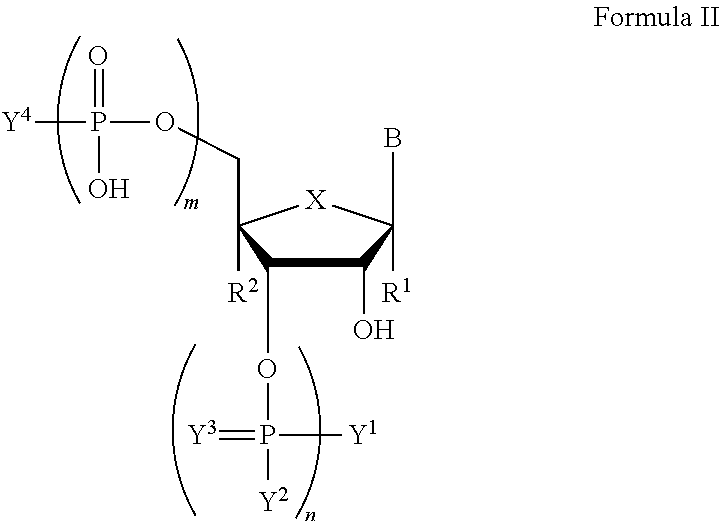
C00040
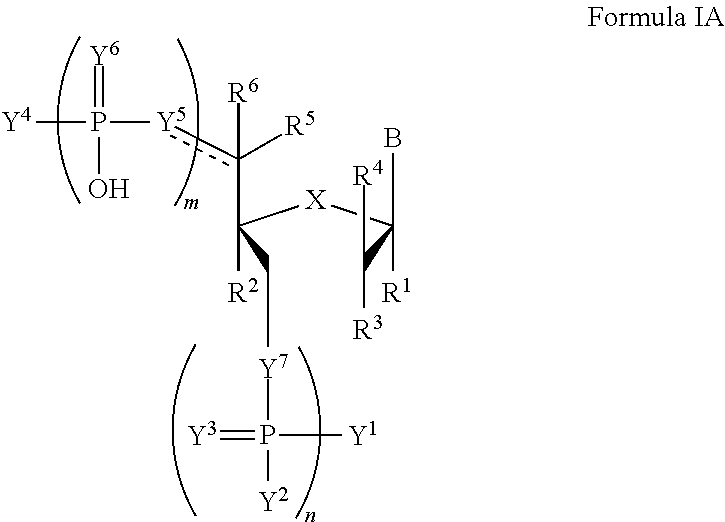
C00041
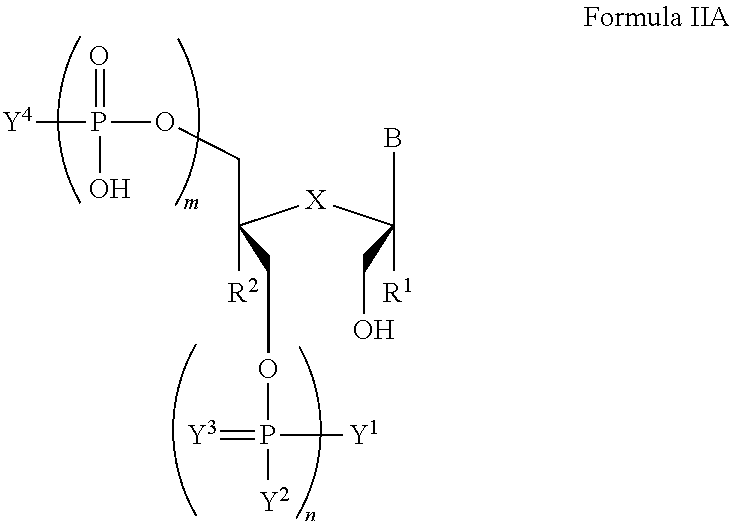
C00042
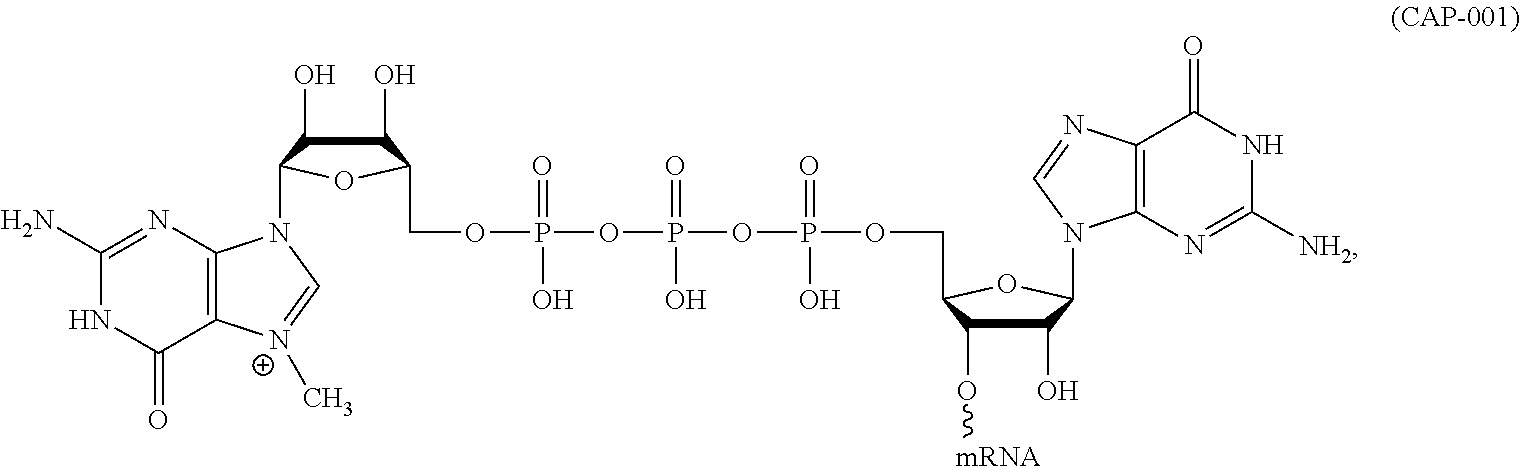
C00043

C00044

C00045

C00046
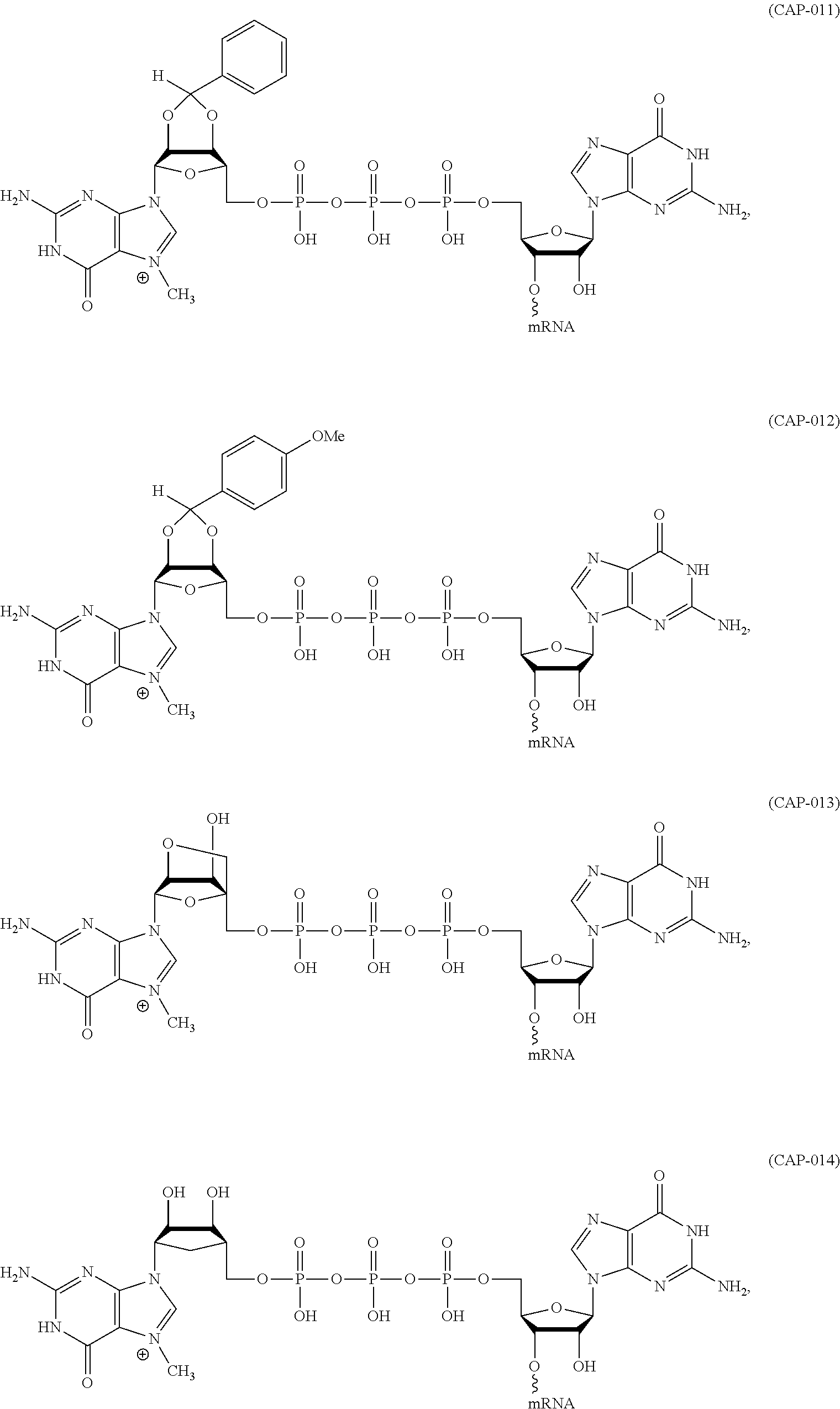
C00047
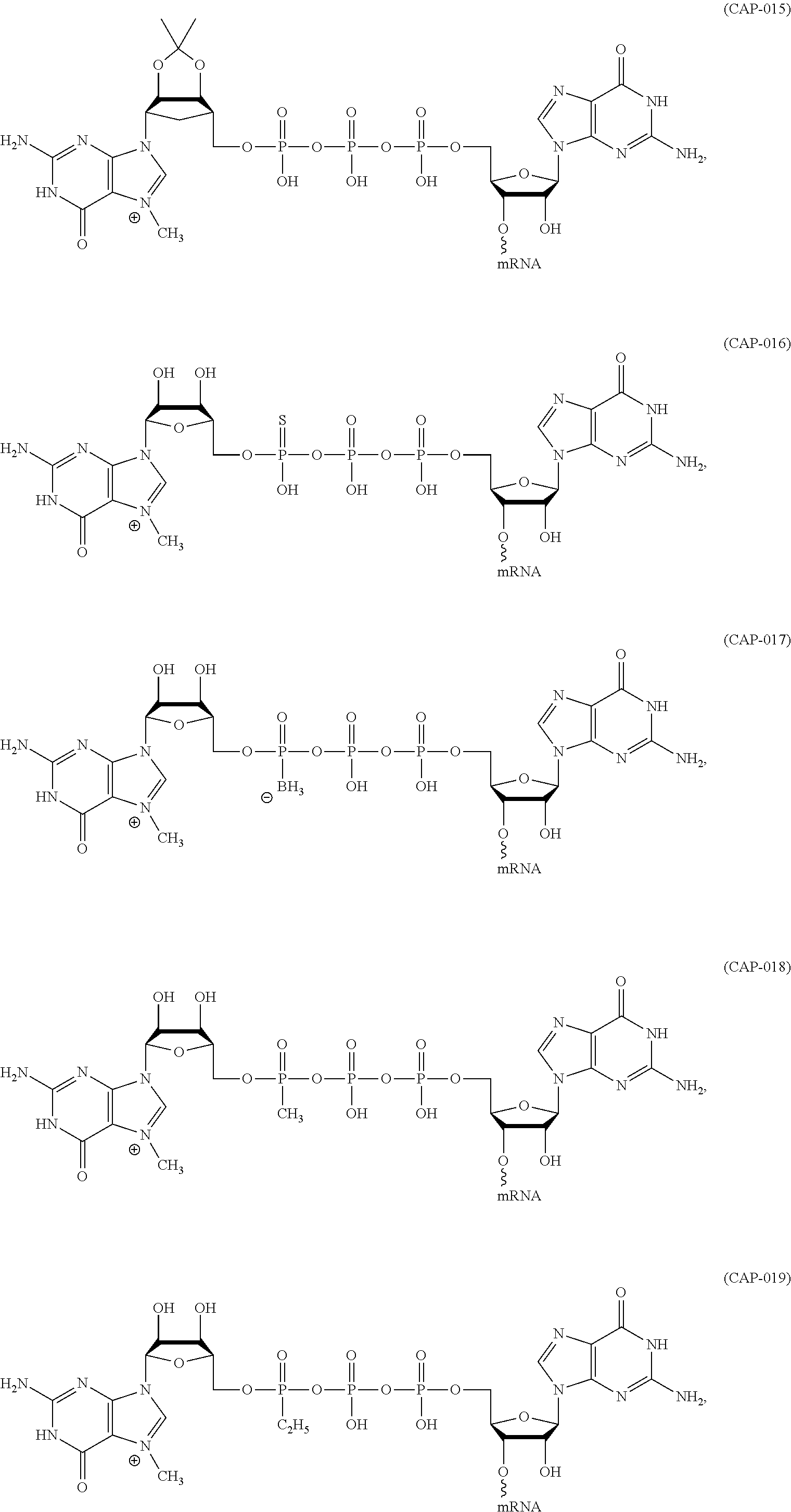
C00048

C00049
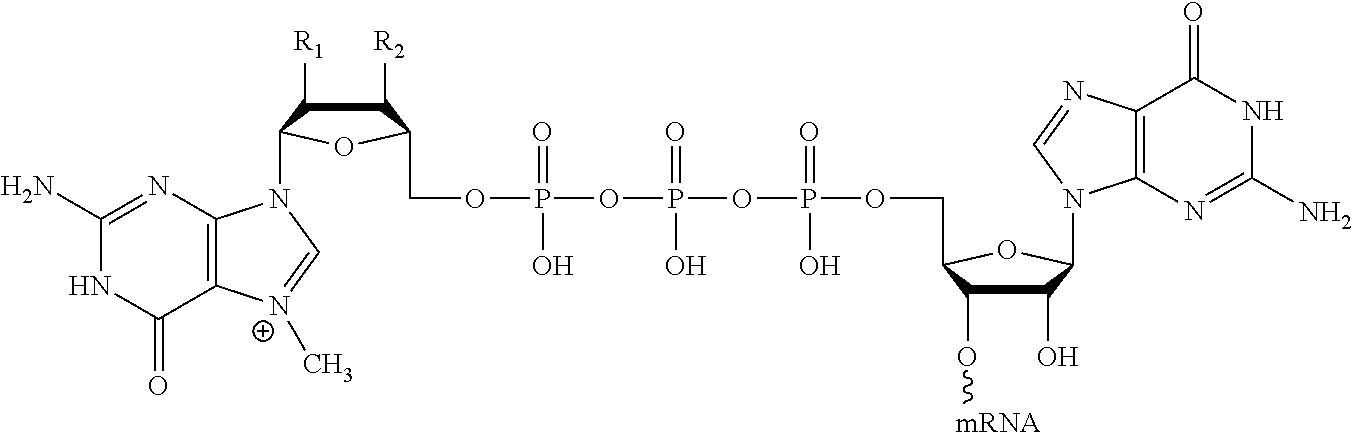
C00050

C00051

C00052
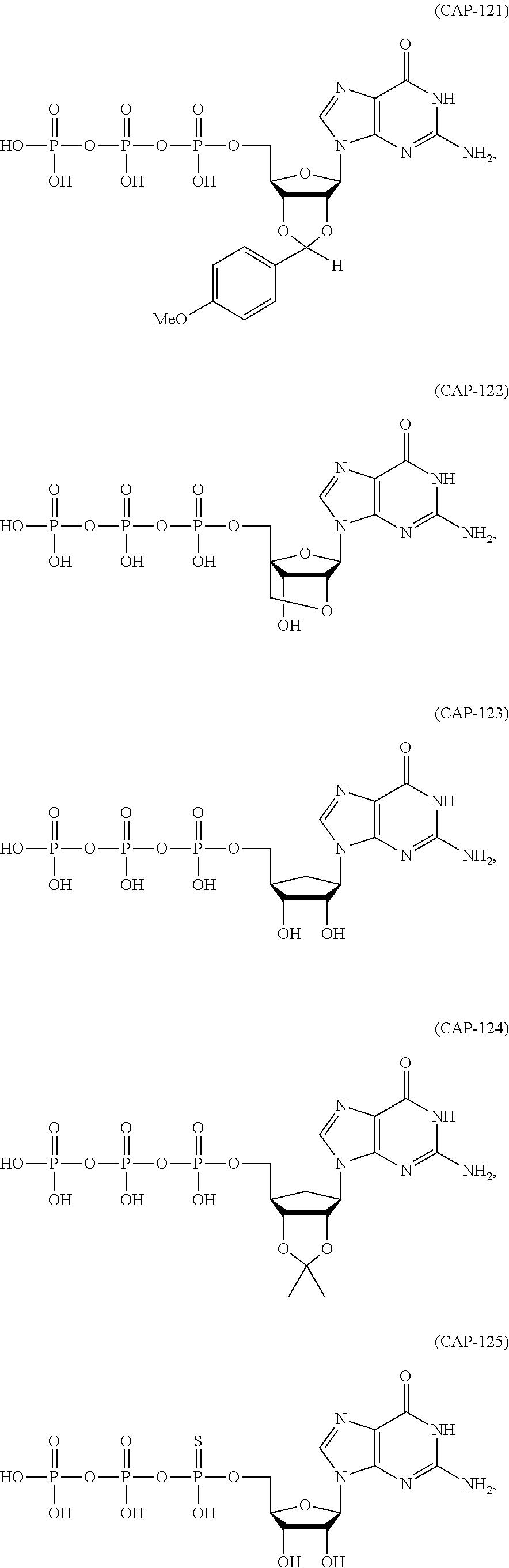
C00053

C00054
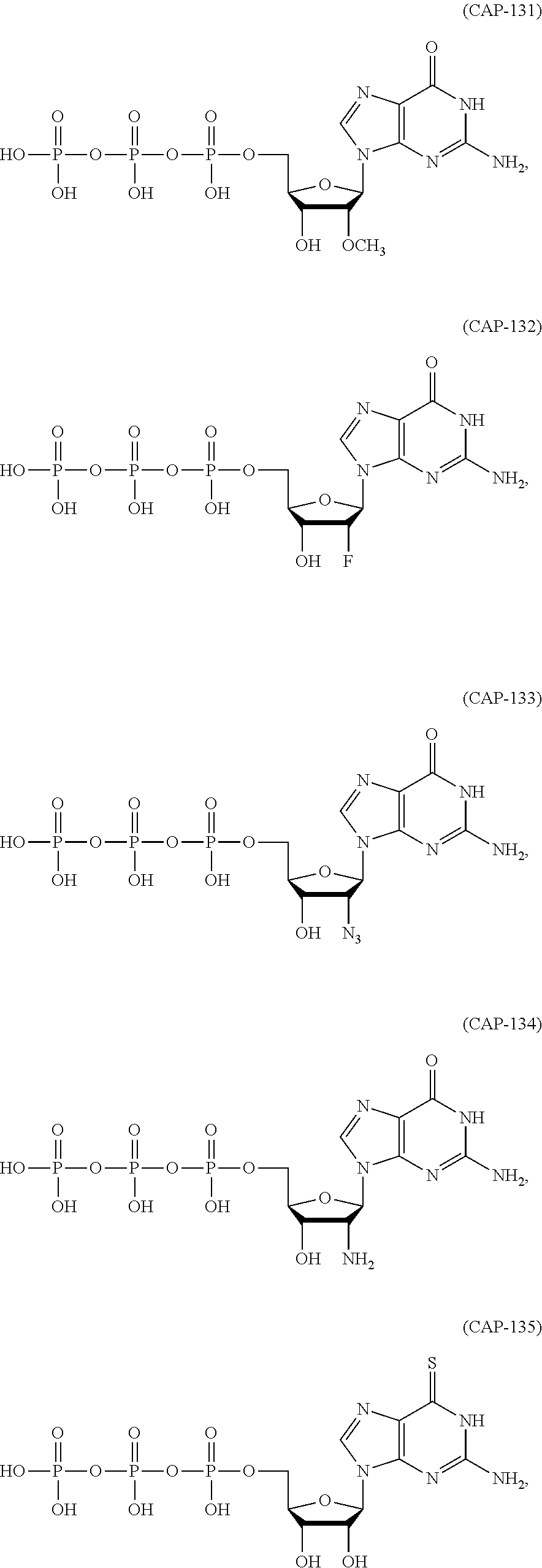
C00055
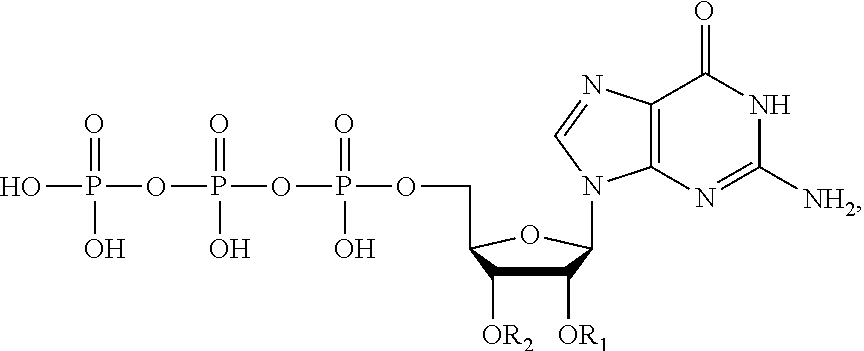
C00056
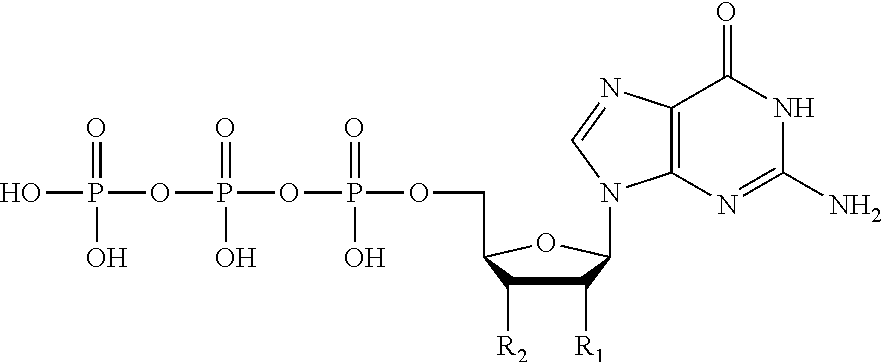
C00057

C00058
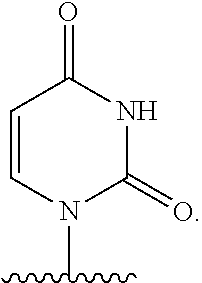
C00059
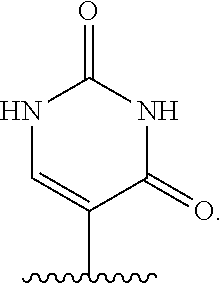
C00060
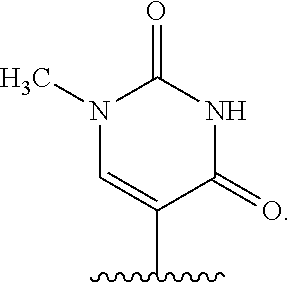
C00061
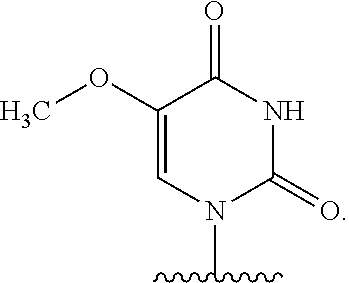
C00062
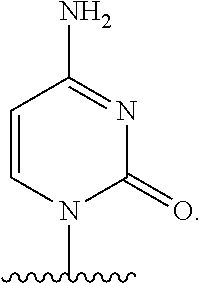
C00063
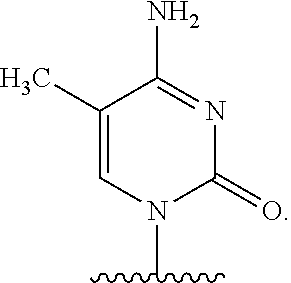
C00064
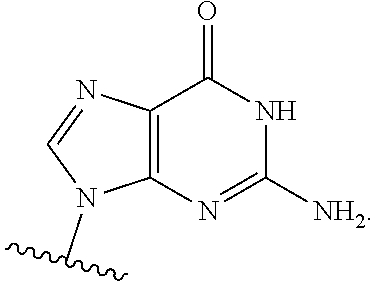
C00065
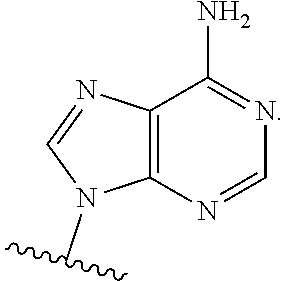
C00066
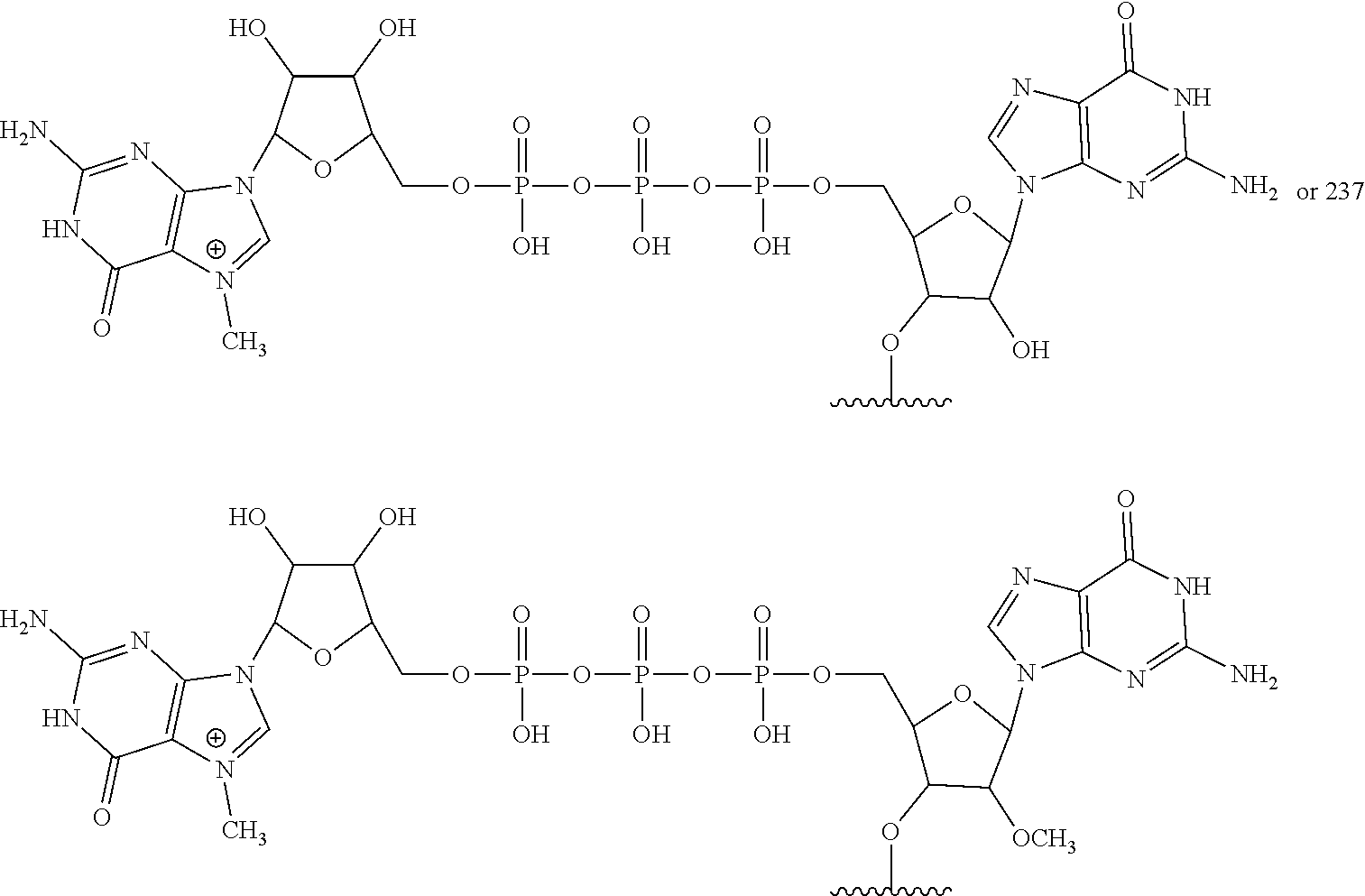
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.