Compositions And Methods For Modulating Pkk Expression
Prakash; Thazha P. ; et al.
U.S. patent application number 16/363969 was filed with the patent office on 2020-02-20 for compositions and methods for modulating pkk expression. This patent application is currently assigned to Ionis Pharmaceuticals, Inc.. The applicant listed for this patent is Ionis Pharmaceuticals, Inc.. Invention is credited to Huynh-Hoa Bui, Susan M. Freier, Thazha P. Prakash, Punit P. Seth, Eric E. Swayze.
| Application Number | 20200056185 16/363969 |
| Document ID | / |
| Family ID | 54359499 |
| Filed Date | 2020-02-20 |












View All Diagrams
| United States Patent Application | 20200056185 |
| Kind Code | A1 |
| Prakash; Thazha P. ; et al. | February 20, 2020 |
COMPOSITIONS AND METHODS FOR MODULATING PKK EXPRESSION
Abstract
Disclosed herein are antisense compounds and methods for decreasing PKK mRNA and protein expression. Such methods, compounds, and compositions are useful to treat, prevent, or ameliorate PKK-associated diseases, disorders, and conditions.
| Inventors: | Prakash; Thazha P.; (Carlsbad, CA) ; Seth; Punit P.; (Carlsbad, CA) ; Swayze; Eric E.; (Encinitas, CA) ; Freier; Susan M.; (San Diego, CA) ; Bui; Huynh-Hoa; (San Diego, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Ionis Pharmaceuticals, Inc. Carlsbad CA |
||||||||||
| Family ID: | 54359499 | ||||||||||
| Appl. No.: | 16/363969 | ||||||||||
| Filed: | March 25, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15308027 | Oct 31, 2016 | 10294477 | ||
| PCT/US2015/028765 | May 1, 2015 | |||
| 16363969 | ||||
| 62088459 | Dec 5, 2014 | |||
| 62058629 | Oct 1, 2014 | |||
| 61987478 | May 1, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 2310/341 20130101; A61P 43/00 20180101; C07H 21/00 20130101; A61P 11/00 20180101; C12N 2310/3341 20130101; C12N 2310/351 20130101; A61P 29/00 20180101; A61P 7/10 20180101; C12N 15/1137 20130101; C12N 2310/315 20130101; A61P 9/10 20180101; A61P 7/02 20180101 |
| International Class: | C12N 15/113 20060101 C12N015/113; C07H 21/00 20060101 C07H021/00 |
Claims
1.-219. (canceled)
220. A compound comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising a portion of at least 15 contiguous nucleobases that is at least 90% complementary to an equal length portion of nucleobases 33183-33242 of SEQ ID NO: 10, and wherein the conjugate group comprises at least one N-Acetylgalactosamine (GalNAc).
221. The compound of claim 220, wherein the portion of at least 15 contiguous nucleobases is 100% complementary to the equal length portion of nucleobases 33183-33242 of SEQ ID NO: 10.
222. The compound of claim 220, wherein the sequence of the modified oligonucleotide is SEQ ID NO: 705.
223. The compound of claim 220, wherein the conjugate group comprises three GalNAcs.
224. The compound of claim 220, wherein the conjugate group consists of: ##STR00280##
225. The compound of claim 224, consisting of the modified oligonucleotide and the conjugate group.
226. The compound of claim 220, wherein the modified oligonucleotide consists of 20 linked nucleosides.
227. The compound of claim 220, wherein the modified oligonucleotide is at least 90% complementary to SEQ ID NO: 10.
228. The compound of claim 220, wherein at least one internucleoside linkage of the modified oligonucleotide is a phosphorothioate linkage.
229. The compound of claim 220, wherein each cytosine of the modified oligonucleotide is a 5'-methylcytosine.
230. The compound of claim 220, wherein the modified oligonucleotide is single-stranded.
231. The compound of claim 220, comprising at least one 2'-O-methoxyethyl nucleoside, 2'-O-methyl nucleoside, constrained ethyl nucleoside, LNA nucleoside, or 3'-fluoro-HNA nucleoside.
232. The compound of claim 220, wherein the modified oligonucleotide is a gapmer.
233. The compound of claim 232, wherein the modified oligonucleotide comprises: a gap segment consisting of 10 linked deoxynucleosides; a 5' wing segment consisting of 5 linked nucleosides; and a 3' wing segment consisting of 5 linked nucleosides; wherein the gap segment is positioned between the 5' wing segment and the 3' wing segment and wherein each nucleoside of each wing segment comprises a modified sugar.
234. The compound of claim 220, wherein the compound is in the form of a salt.
235. A pharmaceutical composition comprising the compound of claim 220 and a pharmaceutically acceptable carrier or diluent.
236. The pharmaceutical composition of claim 235, wherein the pharmaceutically acceptable carrier or diluent is phosphate buffered saline (PBS).
237. The pharmaceutical composition of claim 235, wherein the pharmaceutical composition consists essentially of the compound and PBS.
238. A method comprising administering the compound of claim 220 to a subject in need thereof.
239. The method of claim 238, wherein administering the compound prevents, treats, or ameliorates a PKK associated disease, disorder or condition.
240. A compound comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising a portion of at least 15 contiguous nucleobases that is at least 90% identical to an equal length portion of a sequence selected from SEQ ID NOs: 155, 156, 157, 158, 159, 160, 261, 702, 703, 704, 705, 706, and 707, and wherein the conjugate group comprises at least one N-Acetylgalactosamine (GalNAc).
241. The compound of claim 240, wherein the portion of at least 15 contiguous nucleobases is 100% identical to the sequence selected from SEQ ID NOs: 155, 156, 157, 158, 159, 160, 261, 702, 703, 704, 705, 706, and 707.
242. The compound of claim 240, wherein the sequence of the modified oligonucleotide is selected from SEQ ID NOs: 155, 156, 157, 158, 159, 160, 261, 702, 703, 704, 705, 706, and 707.
243. The compound of claim 240, wherein the conjugate group comprises three GalNAcs.
244. The compound of claim 240, wherein the conjugate group consists of: ##STR00281##
245. The compound of claim 244, consisting of the modified oligonucleotide and the conjugate group.
246. The compound of claim 240, wherein the modified oligonucleotide consists of 20 linked nucleosides.
247. The compound of claim 240, wherein the modified oligonucleotide is at least 90% complementary to SEQ ID NO: 10.
248. The compound of claim 240, wherein at least one internucleoside linkage of the modified oligonucleotide is a phosphorothioate linkage.
249. The compound of claim 240, wherein each cytosine of the modified oligonucleotide is a 5'-methylcytosine.
250. The compound of claim 240, wherein the modified oligonucleotide is single-stranded.
251. The compound of claim 240, comprising at least one 2'-O-methoxyethyl nucleoside, 2'-O-methyl nucleoside, constrained ethyl nucleoside, LNA nucleoside, or 3'-fluoro-HNA nucleoside.
252. The compound of claim 240, wherein the modified oligonucleotide is a gapmer.
253. The compound of claim 252, wherein the modified oligonucleotide comprises: a gap segment consisting of 10 linked deoxynucleosides; a 5' wing segment consisting of 5 linked nucleosides; and a 3' wing segment consisting of 5 linked nucleosides; wherein the gap segment is positioned between the 5' wing segment and the 3' wing segment and wherein each nucleoside of each wing segment comprises a modified sugar.
254. The compound of claim 240, wherein the compound is in the form of a salt.
255. A pharmaceutical composition comprising the compound of claim 240 and a pharmaceutically acceptable carrier or diluent.
256. The pharmaceutical composition of claim 255, wherein the pharmaceutically acceptable carrier or diluent is phosphate buffered saline (PBS).
257. The pharmaceutical composition of claim 255, wherein the pharmaceutical composition consists essentially of the compound and PBS.
258. A method comprising administering the compound of claim 240 to a subject in need thereof.
259. The method of claim 258, wherein administering the compound prevents, treats, or ameliorates a PKK associated disease, disorder or condition.
260. A compound consisting of a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide is described by the following chemical notation: mCes mCes mCes mCes mCes Tds Tds mCds Tds Tds Tds Ads Tds Ads Gds mCes mCes Aes Ges mCe; wherein, A=an adenine, mC=a 5-methylcytosine; G=a guanine, T=a thymine, e=a 2'-O-methoxyethyl modified nucleoside, d=a 2'-deoxynucleoside, and s=a phosphorothioate internucleoside linkage, and wherein the conjugate moiety is described by the following chemical structure: ##STR00282## and wherein the 5' end of the modified oligonucleotide is directly linked to the conjugate moiety.
261. A pharmaceutical composition comprising the compound of claim 260 and at least one of a pharmaceutically acceptable carrier or diluent.
262. The pharmaceutical composition of claim 261, wherein the pharmaceutically acceptable carrier or diluent is PBS.
263. The pharmaceutical composition of claim 261, wherein the pharmaceutical composition consists essentially of the compound and PBS.
264. A method comprising administering the compound of claim 260 to a subject in need thereof.
265. The method of claim 264, wherein administering the compound prevents, treats, or ameliorates a PKK associated disease, disorder or condition.
Description
SEQUENCE LISTING
[0001] The present application is being filed along with a Sequence Listing in electronic format. The Sequence Listing is provided as a file entitled BIOL0252USC1SEQ_ST25.txt created Mar. 25, 2019, which is approximately 636 KB in size. The information in the electronic format of the sequence listing is incorporated herein by reference in its entirety.
FIELD
[0002] Provided are compounds, compositions, and methods for reducing expression of human plasma prekallikrein (PKK) mRNA and protein in an animal. Such compositions and methods are useful to treat, prevent, or ameliorate inflammatory and thromboembolic conditions.
BACKGROUND
[0003] Plasma prekallikrein (PKK) is the precursor of plasma kallikrein (PK), which is encoded by the KLKB1 gene. PKK is a glycoprotein that participates in the surface-dependent activation of blood coagulation, fibrinolysis, kinin generation, and inflammation. PKK is converted to PK by Factor XIIa by the cleavage of an internal Arg-Ile peptide bond. PK liberates kinins from kininogens and also generates plasmin from plasminogen. PK is a member of the kinin-kallikrein pathway, which consists of several proteins that play a role in inflammation, blood pressure control, coagulation, and pain.
SUMMARY
[0004] Provided herein are compounds, compositions, and methods for modulating expression of PKK mRNA and protein. In certain embodiments, compounds useful for modulating expression of PKK mRNA and protein are antisense compounds. In certain embodiments, the antisense compounds are antisense oligonucleotides.
[0005] In certain embodiments, modulation can occur in a cell or tissue. In certain embodiments, the cell or tissue is in an animal. In certain embodiments, the animal is a human. In certain embodiments, PKK mRNA levels are reduced. In certain embodiments, PKK protein levels are reduced. Such reduction can occur in a time-dependent manner or in a dose-dependent manner.
[0006] Also provided are compounds, compositions, and methods useful for preventing, treating, and ameliorating diseases, disorders, and conditions associated with PKK. In certain embodiments, such PKK associated diseases, disorders, and conditions are inflammatory diseases. In certain embodiments, the inflammatory disease may be an acute or chronic inflammatory disease. In certain embodiments, such inflammatory diseases may include hereditary angioedema (HAE), edema, angioedema, swelling, angioedema of the lids, ocular edema, macular edema, and cerebral edema. In certain embodiments, such PKK associated diseases, disorders, and conditions are thromboembolic diseases. In certain embodiments, such thromboembolic diseases may include thrombosis, embolism, thromboembolism, deep vein thrombosis, pulmonary embolism, myocardial infarction, stroke, and infarct.
[0007] Such diseases, disorders, and conditions can have one or more risk factors, causes, or outcomes in common.
[0008] Certain risk factors and causes for development of an inflammatory disease include genetic predisposition to an inflammatory disease and environmental factors. In certain embodiments, the subject has a mutated complement 1 esterase inhibitor (C1-INH) gene or mutated Factor 12 gene. In certain embodiments, the subject has taken or is on angiotensin-converting enzyme inhibitors (ACE inhibitors) or angiotensin II receptor blockers (ARBs). In certain embodiments, the subject has had an allergic reaction leading to angioedema. In certain embodiments, the subject has type I HAE. In certain embodiments, the subject has type II HAE. In certain embodiments, the subject has type III HAE.
[0009] Certain outcomes associated with development of an inflammatory disease include edema/swelling in various body parts including the extremities (i.e., hands, feet, arms, legs), the intestines (abdomen), the face, the genitals, the larynx (i.e., voice box); vascular permeability; vascular leakage; generalized inflammation; abdominal pain; bloating; vomiting; diarrhea; itchy skin; respiratory (asthmatic) reactions; rhinitis; anaphylaxis; bronchoconstriction; hypotension; coma; and death.
[0010] Certain risk factors and causes for development of a thromboembolic disease include genetic predisposition to a thromboembolic disease, immobility, surgery (particularly orthopedic surgery), malignancy, pregnancy, older age, use of oral contraceptives, atrial fibrillation, previous thromboembolic condition, chronic inflammatory disease, and inherited or acquired prothrombotic clotting disorders. Certain outcomes associated with development of a thromboembolic condition include decreased blood flow through an affected vessel, death of tissue, and death.
[0011] In certain embodiments, methods of treatment include administering a PKK antisense compound to an individual in need thereof. In certain embodiments, methods of treatment include administering a PKK antisense oligonucleotide to an individual in need thereof.
DETAILED DESCRIPTION
[0012] It is to be understood that both the foregoing general description and the following detailed description are exemplary and explanatory only and are not restrictive of the invention, as claimed. Herein, the use of the singular includes the plural unless specifically stated otherwise. As used herein, the use of "or" means "and/or" unless stated otherwise. Furthermore, the use of the term "including" as well as other forms, such as "includes" and "included", is not limiting. Also, terms such as "element" or "component" encompass both elements and components comprising one unit and elements and components that comprise more than one subunit, unless specifically stated otherwise.
[0013] Unless specific definitions are provided, the nomenclature used in connection with, and the procedures and techniques of, analytical chemistry, synthetic organic chemistry, and medicinal and pharmaceutical chemistry described herein are those well known and commonly used in the art. Standard techniques may be used for chemical synthesis, and chemical analysis. Certain such techniques and procedures may be found for example in "Carbohydrate Modifications in Antisense Research" Edited by Sangvi and Cook, American Chemical Society, Washington D.C., 1994; "Remington's Pharmaceutical Sciences," Mack Publishing Co., Easton, Pa., 21.sup.st edition, 2005; and "Antisense Drug Technology, Principles, Strategies, and Applications" Edited by Stanley T. Crooke, CRC Press, Boca Raton, Fla.; and Sambrook et al., "Molecular Cloning, A laboratory Manual," 2.sup.nd Edition, Cold Spring Harbor Laboratory Press, 1989, which are hereby incorporated by reference for any purpose. Where permitted, all patents, applications, published applications and other publications and other data referred to throughout in the disclosure are incorporated by reference herein in their entirety.
[0014] The section headings used herein are for organizational purposes only and are not to be construed as limiting the subject matter described. All documents, or portions of documents, cited in this application, including, but not limited to, patents, patent applications, articles, books, and treatises, are hereby expressly incorporated by reference for the portions of the document discussed herein, as well as in their entirety.
Definitions
[0015] Unless specific definitions are provided, the nomenclature utilized in connection with, and the procedures and techniques of, analytical chemistry, synthetic organic chemistry, and medicinal and pharmaceutical chemistry described herein are those well known and commonly used in the art. Standard techniques may be used for chemical synthesis, and chemical analysis. Where permitted, all patents, applications, published applications and other publications, GENBANK Accession Numbers and associated sequence information obtainable through databases such as National Center for Biotechnology Information (NCBI) and other data referred to throughout in the disclosure herein are incorporated by reference for the portions of the document discussed herein, as well as in their entirety.
[0016] Unless otherwise indicated, the following terms have the following meanings:
[0017] "2'-O-methoxyethyl" (also 2'-MOE and 2'-OCH.sub.2CH.sub.2--OCH.sub.3 and MOE) refers to an O-methoxyethyl modification of the 2' position of a furanose ring. A 2'-O-methoxyethyl modified sugar is a modified sugar.
[0018] "2'-O-methoxyethyl modified nucleoside" (also "2'-MOE nucleoside") means a nucleoside comprising a 2'-MOE modified sugar moiety.
[0019] "2'-substituted nucleoside" means a nucleoside comprising a substituent at the 2'-position of the furanose ring other than H or OH. In certain embodiments, 2' substituted nucleosides include nucleosides with bicyclic sugar modifications.
[0020] "2'-deoxynucleoside" means a nucleoside comprising a hydrogen at the 2' position of the sugar portion of the nucleoside.
[0021] "3' target site" refers to the nucleotide of a target nucleic acid which is complementary to the 3'-most nucleotide of a particular antisense compound.
[0022] "5' target site" refers to the nucleotide of a target nucleic acid which is complementary to the 5'-most nucleotide of a particular antisense compound.
[0023] "5-methylcytosine" means a cytosine modified with a methyl group attached to the 5 position. A 5-methylcytosine is a modified nucleobase.
[0024] "About" means within +7% of a value. For example, if it is stated, "the compounds affected at least about 70% inhibition of PKK", it is implied that the PKK levels are inhibited within a range of 63% and 77%.
[0025] "Administered concomitantly" refers to the co-administration of two pharmaceutical agents in any manner in which the pharmacological effects of both are manifest in the patient at the same time. Concomitant administration does not require that both pharmaceutical agents be administered in a single pharmaceutical composition, in the same dosage form, or by the same route of administration. The effects of both pharmaceutical agents need not manifest themselves at the same time. The effects need only be overlapping for a period of time and need not be coextensive.
[0026] "Administering" means providing a pharmaceutical agent to an animal, and includes, but is not limited to administering by a medical professional and self-administering.
[0027] "Alkyl," as used herein, means a saturated straight or branched hydrocarbon radical containing up to twenty four carbon atoms. Examples of alkyl groups include without limitation, methyl, ethyl, propyl, butyl, isopropyl, n-hexyl, octyl, decyl, dodecyl and the like. Alkyl groups typically include from 1 to about 24 carbon atoms, more typically from 1 to about 12 carbon atoms (C.sub.1-C.sub.12 alkyl) with from 1 to about 6 carbon atoms being more preferred.
[0028] As used herein, "alkenyl," means a straight or branched hydrocarbon chain radical containing up to twenty four carbon atoms and having at least one carbon-carbon double bond. Examples of alkenyl groups include without limitation, ethenyl, propenyl, butenyl, 1-methyl-2-buten-1-yl, dienes such as 1,3-butadiene and the like. Alkenyl groups typically include from 2 to about 24 carbon atoms, more typically from 2 to about 12 carbon atoms with from 2 to about 6 carbon atoms being more preferred. Alkenyl groups as used herein may optionally include one or more further substituent groups.
[0029] As used herein, "alkynyl," means a straight or branched hydrocarbon radical containing up to twenty four carbon atoms and having at least one carbon-carbon triple bond. Examples of alkynyl groups include, without limitation, ethynyl, 1-propynyl, 1-butynyl, and the like. Alkynyl groups typically include from 2 to about 24 carbon atoms, more typically from 2 to about 12 carbon atoms with from 2 to about 6 carbon atoms being more preferred. Alkynyl groups as used herein may optionally include one or more further substituent groups.
[0030] As used herein, "acyl," means a radical formed by removal of a hydroxyl group from an organic acid and has the general Formula --C(O)--X where X is typically aliphatic, alicyclic or aromatic. Examples include aliphatic carbonyls, aromatic carbonyls, aliphatic sulfonyls, aromatic sulfinyls, aliphatic sulfinyls, aromatic phosphates, aliphatic phosphates and the like. Acyl groups as used herein may optionally include further substituent groups.
[0031] As used herein, "alicyclic" means a cyclic ring system wherein the ring is aliphatic. The ring system can comprise one or more rings wherein at least one ring is aliphatic. Preferred alicyclics include rings having from about 5 to about 9 carbon atoms in the ring. Alicyclic as used herein may optionally include further substituent groups.
[0032] As used herein, "aliphatic" means a straight or branched hydrocarbon radical containing up to twenty four carbon atoms wherein the saturation between any two carbon atoms is a single, double or triple bond. An aliphatic group preferably contains from 1 to about 24 carbon atoms, more typically from 1 to about 12 carbon atoms with from 1 to about 6 carbon atoms being more preferred. The straight or branched chain of an aliphatic group may be interrupted with one or more heteroatoms that include nitrogen, oxygen, sulfur and phosphorus. Such aliphatic groups interrupted by heteroatoms include without limitation, polyalkoxys, such as polyalkylene glycols, polyamines, and polyimines. Aliphatic groups as used herein may optionally include further substituent groups.
[0033] As used herein, "alkoxy" means a radical formed between an alkyl group and an oxygen atom wherein the oxygen atom is used to attach the alkoxy group to a parent molecule. Examples of alkoxy groups include without limitation, methoxy, ethoxy, propoxy, isopropoxy, n-butoxy, sec-butoxy, tert-butoxy, n-pentoxy, neopentoxy, n-hexoxy and the like. Alkoxy groups as used herein may optionally include further substituent groups.
[0034] As used herein, "aminoalkyl" means an amino substituted C.sub.1-C.sub.12 alkyl radical. The alkyl portion of the radical forms a covalent bond with a parent molecule. The amino group can be located at any position and the aminoalkyl group can be substituted with a further substituent group at the alkyl and/or amino portions.
[0035] As used herein, "aralkyl" and "arylalkyl" mean an aromatic group that is covalently linked to a C.sub.1-C.sub.12 alkyl radical. The alkyl radical portion of the resulting aralkyl (or arylalkyl) group forms a covalent bond with a parent molecule. Examples include without limitation, benzyl, phenethyl and the like. Aralkyl groups as used herein may optionally include further substituent groups attached to the alkyl, the aryl or both groups that form the radical group.
[0036] As used herein, "aryl" and "aromatic" mean a mono- or polycyclic carbocyclic ring system radicals having one or more aromatic rings. Examples of aryl groups include without limitation, phenyl, naphthyl, tetrahydronaphthyl, indanyl, idenyl and the like. Preferred aryl ring systems have from about 5 to about 20 carbon atoms in one or more rings. Aryl groups as used herein may optionally include further substituent groups.
[0037] "Amelioration" refers to a lessening, slowing, stopping, or reversing of at least one indicator of the severity of a condition or disease. The severity of indicators may be determined by subjective or objective measures, which are known to those skilled in the art.
[0038] "Animal" refers to a human or non-human animal, including, but not limited to, mice, rats, rabbits, dogs, cats, pigs, and non-human primates, including, but not limited to, monkeys and chimpanzees.
[0039] "Antisense activity" means any detectable or measurable activity attributable to the hybridization of an antisense compound to its target nucleic acid. In certain embodiments, antisense activity is a decrease in the amount or expression of a target nucleic acid or protein encoded by such target nucleic acid. "Antisense compound" means an oligomeric compound that is is capable of undergoing hybridization to a target nucleic acid through hydrogen bonding. Examples of antisense compounds include single-stranded and double-stranded compounds, such as, antisense oligonucleotides, siRNAs, shRNAs, ssRNAs, and occupancy-based compounds.
[0040] "Antisense compound" means an oligomeric compound that is capable of undergoing hybridization to a target nucleic acid through hydrogen bonding. Examples of antisense compounds include single-stranded and double-stranded compounds, such as, antisense oligonucleotides, siRNAs, shRNAs, ssRNAs, and occupancy-based compounds.
[0041] "Antisense inhibition" means reduction of target nucleic acid levels in the presence of an antisense compound complementary to a target nucleic acid compared to target nucleic acid levels or in the absence of the antisense compound. "Antisense mechanisms" are all those mechanisms involving hybridization of a compound with target nucleic acid, wherein the outcome or effect of the hybridization is either target degradation or target occupancy with concomitant stalling of the cellular machinery involving, for example, transcription or splicing.
[0042] "Antisense mechanisms" are all those mechanisms involving hybridization of a compound with a target nucleic acid, wherein the outcome or effect of the hybridization is either target degradation or target occupancy with concomitant stalling of the cellular machinery involving, for example, transcription or splicing.
[0043] "Antisense oligonucleotide" means a single-stranded oligonucleotide having a nucleobase sequence that permits hybridization to a corresponding segment of a target nucleic acid. "Base complementarity" refers to the capacity for the precise base pairing of nucleobases of an antisense oligonucleotide with corresponding nucleobases in a target nucleic acid (i.e., hybridization), and is mediated by Watson-Crick, Hoogsteen or reversed Hoogsteen hydrogen binding between corresponding nucleobases.
[0044] "Base complementarity" refers to the capacity for the precise base pairing of nucleobases of an antisense oligonucleotide with corresponding nucleobases in a target nucleic acid (i.e., hybridization), and is mediated by Watson-Crick, Hoogsteen or reversed Hoogsteen hydrogen binding between corresponding nucleobases.
[0045] "Bicyclic sugar" means a furanose ring modified by the bridging of two atoms. A bicyclic sugar is a modified sugar.
[0046] "Bicyclic nucleoside" (also BNA) means a nucleoside having a sugar moiety comprising a bridge connecting two carbon atoms of the sugar ring, thereby forming a bicyclic ring system. In certain embodiments, the bridge connects the 4'-carbon and the 2'-carbon of the sugar ring.
[0047] "Cap structure" or "terminal cap moiety" means chemical modifications, which have been incorporated at either terminus of an antisense compound.
[0048] "Carbohydrate" means a naturally occurring carbohydrate, a modified carbohydrate, or a carbohydrate derivative.
[0049] "Carbohydrate cluster" means a compound having one or more carbohydrate residues attached to a scaffold or linker group. (see, e.g., Maier et al., "Synthesis of Antisense Oligonucleotides Conjugated to a Multivalent Carbohydrate Cluster for Cellular Targeting," Bioconjugate Chemistry, 2003, (14): 18-29, which is incorporated herein by reference in its entirety, or Rensen et al., "Design and Synthesis of Novel N-Acetylgalactosamine-Terminated Glycolipids for Targeting of Lipoproteins to the Hepatic Asiaglycoprotein Receptor," J Med. Chem. 2004, (47): 5798-5808, for examples of carbohydrate conjugate clusters).
[0050] "Carbohydrate derivative" means any compound which may be synthesized using a carbohydrate as a starting material or intermediate.
[0051] "cEt" or "constrained ethyl" means a bicyclic nucleoside having a sugar moiety comprising a bridge connecting the 4'-carbon and the 2'-carbon, wherein the bridge has the formula: 4'-CH(CH.sub.3)--O-2'.
[0052] "cEt modified nucleoside" (also "constrained ethyl nucleoside") means a nucleoside comprising a bicyclic sugar moiety comprising a 4'-CH(CH.sub.3)--O-2' bridge.
[0053] "Chemically distinct region" refers to a region of an antisense compound that is in some way chemically different than another region of the same antisense compound. For example, a region having 2'-O-methoxyethyl nucleosides is chemically distinct from a region having nucleosides without 2'-O-methoxyethyl modifications.
[0054] "Chemical modification" means a chemical difference in a compound when compared to a naturally occurring counterpart. Chemical modifications of oligonucleotides include nucleoside modifications (including sugar moiety modifications and nucleobase modifications) and internucleoside linkage modifications. In reference to an oligonucleotide, chemical modification does not include differences only in nucleobase sequence.
[0055] "Chimeric antisense compound" means an antisense compound that has at least two chemically distinct regions, each position having a plurality of subunits.
[0056] "Cleavable bond" means any chemical bond capable of being split. In certain embodiments, a cleavable bond is selected from among: an amide, a polyamide, an ester, an ether, one or both esters of a phosphodiester, a phosphate ester, a carbamate, a di-sulfide, or a peptide.
[0057] "Cleavable moiety" means a bond or group that is capable of being split under physiological conditions. In certain embodiments, a cleavable moiety is cleaved inside a cell or sub-cellular compartments, such as a lysosome. In certain embodiments, a cleavable moiety is cleaved by endogenous enzymes, such as nucleases. In certain embodiments, a cleavable moiety comprises a group of atoms having one, two, three, four, or more than four cleavable bonds.
[0058] "Co-administration" means administration of two or more pharmaceutical agents to an individual. The two or more pharmaceutical agents may be in a single pharmaceutical composition, or may be in separate pharmaceutical compositions. Each of the two or more pharmaceutical agents may be administered through the same or different routes of administration. Co-administration encompasses parallel or sequential administration.
[0059] "Complementarity" means the capacity for pairing between nucleobases of a first nucleic acid and a second nucleic acid.
[0060] "Comprise," "comprises," and "comprising" will be understood to imply the inclusion of a stated step or element or group of steps or elements but not the exclusion of any other step or element or group of steps or elements.
[0061] "Conjugate" or "conjugate group" means an atom or group of atoms bound to an oligonucleotide or oligomeric compound. In general, conjugate groups modify one or more properties of the compound to which they are attached, including, but not limited to pharmacodynamic, pharmacokinetic, binding, absorption, cellular distribution, cellular uptake, charge and/or clearance properties.
"conjugate linker" or "linker" in the context of a conjugate group means a portion of a conjugate group comprising any atom or group of atoms and which covalently link (1) an oligonucleotide to another portion of the conjugate group or (2) two or more portions of the conjugate group.
[0062] Conjugate groups are shown herein as radicals, providing a bond for forming covalent attachment to an oligomeric compound such as an antisense oligonucleotide. In certain embodiments, the point of attachment on the oligomeric compound is the 3'-oxygen atom of the 3'-hydroxyl group of the 3' terminal nucleoside of the oligomeric compound. In certain embodiments the point of attachment on the oligomeric compound is the 5'-oxygen atom of the 5'-hydroxyl group of the 5' terminal nucleoside of the oligomeric compound. In certain embodiments, the bond for forming attachment to the oligomeric compound is a cleavable bond. In certain such embodiments, such cleavable bond constitutes all or part of a cleavable moiety.
[0063] In certain embodiments, conjugate groups comprise a cleavable moiety (e.g., a cleavable bond or cleavable nucleoside) and a carbohydrate cluster portion, such as a GalNAc cluster portion. Such carbohydrate cluster portion comprises: a targeting moiety and, optionally, a conjugate linker. In certain embodiments, the carbohydrate cluster portion is identified by the number and identity of the ligand. For example, in certain embodiments, the carbohydrate cluster portion comprises 3 GalNAc groups and is designated "GalNAc.sub.3". In certain embodiments, the carbohydrate cluster portion comprises 4 GalNAc groups and is designated "GalNAc.sub.4". Specific carbohydrate cluster portions (having specific tether, branching and conjugate linker groups) are described herein and designated by Roman numeral followed by subscript "a". Accordingly "GalNac3-1.sub.a" refers to a specific carbohydrate cluster portion of a conjugate group having 3 GalNac groups and specifically identified tether, branching and linking groups. Such carbohydrate cluster fragment is attached to an oligomeric compound via a cleavable moiety, such as a cleavable bond or cleavable nucleoside.
[0064] "Conjugate compound" means any atoms, group of atoms, or group of linked atoms suitable for use as a conjugate group. In certain embodiments, conjugate compounds may possess or impart one or more properties, including, but not limited to pharmacodynamic, pharmacokinetic, binding, absorption, cellular distribution, cellular uptake, charge and/or clearance properties.
[0065] "Contiguous nucleobases" means nucleobases immediately adjacent to each other.
[0066] "Designing" or "Designed to" refer to the process of creating an oligomeric compound that specifically hybridizes with a selected nucleic acid molecule.
[0067] "Diluent" means an ingredient in a composition that lacks pharmacological activity, but is pharmaceutically necessary or desirable. For example, in drugs that are injected, the diluent may be a liquid, e.g. saline solution.
[0068] "Dose" means a specified quantity of a pharmaceutical agent provided in a single administration, or in a specified time period. In certain embodiments, a dose may be administered in one, two, or more boluses, tablets, or injections. For example, in certain embodiments where subcutaneous administration is desired, the desired dose requires a volume not easily accommodated by a single injection, therefore, two or more injections may be used to achieve the desired dose. In certain embodiments, the pharmaceutical agent is administered by infusion over an extended period of time or continuously. Doses may be stated as the amount of pharmaceutical agent per hour, day, week, or month.
[0069] "Downstream" refers to the relative direction toward the 3' end or C-terminal end of a nucleic acid.
[0070] "Effective amount" in the context of modulating an activity or of treating or preventing a condition means the administration of that amount of pharmaceutical agent to a subject in need of such modulation, treatment, or prophylaxis, either in a single dose or as part of a series, that is effective for modulation of that effect, or for treatment or prophylaxis or improvement of that condition. The effective amount may vary among individuals depending on the health and physical condition of the individual to be treated, the taxonomic group of the individuals to be treated, the formulation of the composition, assessment of the individual's medical condition, and other relevant factors.
[0071] "Efficacy" means the ability to produce a desired effect.
[0072] "Expression" includes all the functions by which a gene's coded information is converted into structures present and operating in a cell. Such structures include, but are not limited to the products of transcription and translation.
[0073] "Fully complementary" or "100% complementary" means each nucleobase of a first nucleic acid has a complementary nucleobase in a second nucleic acid. In certain embodiments, a first nucleic acid is an antisense compound and a target nucleic acid is a second nucleic acid.
[0074] "Gapmer" means a chimeric antisense compound in which an internal region having a plurality of nucleosides that support RNase H cleavage is positioned between external regions having one or more nucleosides, wherein the nucleosides comprising the internal region are chemically distinct from the nucleoside or nucleosides comprising the external regions. The internal region may be referred to as a "gap" and the external regions may be referred to as the "wings."
[0075] "Halo" and "halogen," mean an atom selected from fluorine, chlorine, bromine and iodine.
[0076] "Heteroaryl," and "heteroaromatic," mean a radical comprising a mono- or poly-cyclic aromatic ring, ring system or fused ring system wherein at least one of the rings is aromatic and includes one or more heteroatoms. Heteroaryl is also meant to include fused ring systems including systems where one or more of the fused rings contain no heteroatoms. Heteroaryl groups typically include one ring atom selected from sulfur, nitrogen or oxygen. Examples of heteroaryl groups include without limitation, pyridinyl, pyrazinyl, pyrimidinyl, pyrrolyl, pyrazolyl, imidazolyl, thiazolyl, oxazolyl, isooxazolyl, thiadiazolyl, oxadiazolyl, thiophenyl, furanyl, quinolinyl, isoquinolinyl, benzimidazolyl, benzooxazolyl, quinoxalinyl and the like. Heteroaryl radicals can be attached to a parent molecule directly or through a linking moiety such as an aliphatic group or hetero atom. Heteroaryl groups as used herein may optionally include further substituent groups.
[0077] "Hybridization" means the annealing of complementary nucleic acid molecules. In certain embodiments, complementary nucleic acid molecules include, but are not limited to, an antisense compound and a target nucleic acid. In certain embodiments, complementary nucleic acid molecules include, but are not limited to, an antisense oligonucleotide and a nucleic acid target.
[0078] "Identifying an animal having an inflammatory disease" means identifying an animal having been diagnosed with an inflammatory disease or predisposed to develop an inflammatory disease. Individuals predisposed to develop an inflammatory disease include those having one or more risk factors for developing an inflammatory disease including environmental factors, having a personal or family history, or genetic predisposition to one or more inflammatory disease. Such identification may be accomplished by any method including evaluating an individual's medical history and standard clinical tests or assessments, such as genetic testing.
[0079] "Identifying an animal having a PKK associated disease" means identifying an animal having been diagnosed with a PKK associated disease or predisposed to develop a PKK associated disease. Individuals predisposed to develop a PKK associated disease include those having one or more risk factors for developing a PKK associated disease including having a personal or family history, or genetic predisposition of one or more PKK associated diseases. Such identification may be accomplished by any method including evaluating an individual's medical history and standard clinical tests or assessments, such as genetic testing.
[0080] "Identifying an animal having a thromboembolic disease" means identifying an animal having been diagnosed with a thromboembolic disease or predisposed to develop a thromboembolic disease. Individuals predisposed to develop a thromboembolic disease include those having one or more risk factors for developing a thromboembolic disease including having a personal or family history, or genetic predisposition of one or more thromboembolic diseases, immobility, surgery (particularly orthopedic surgery), malignancy, pregnancy, older age, use of oral contraceptives, atrial fibrillation, previous thromboembolic condition, chronic inflammatory disease, and inherited or acquired prothrombotic clotting disorders. Such identification may be accomplished by any method including evaluating an individual's medical history and standard clinical tests or assessments, such as genetic testing.
[0081] "Immediately adjacent" means there are no intervening elements between the immediately adjacent elements. "Individual" means a human or non-human animal selected for treatment or therapy.
[0082] "Individual" means a human or non-human animal selected for treatment or therapy.
[0083] "Inhibiting PKK" means reducing the level or expression of a PKK mRNA and/or protein. In certain embodiments, PKK mRNA and/or protein levels are inhibited in the presence of an antisense compound targeting PKK, including an antisense oligonucleotide targeting PKK, as compared to expression of PKK mRNA and/or protein levels in the absence of a PKK antisense compound, such as an antisense oligonucleotide.
[0084] "Inhibiting the expression or activity" refers to a reduction or blockade of the expression or activity and does not necessarily indicate a total elimination of expression or activity.
[0085] "Internucleoside linkage" refers to the chemical bond between nucleosides.
[0086] "Internucleoside neutral linking group" means a neutral linking group that directly links two nucleosides.
[0087] "Internucleoside phosphorus linking group" means a phosphorus linking group that directly links two nucleosides.
[0088] "Linkage motif" means a pattern of linkage modifications in an oligonucleotide or region thereof. The nucleosides of such an oligonucleotide may be modified or unmodified. Unless otherwise indicated, motifs herein describing only linkages are intended to be linkage motifs. Thus, in such instances, the nucleosides are not limited.
[0089] "Linked nucleosides" means adjacent nucleosides linked together by an internucleoside linkage.
[0090] "Locked nucleic acid" or "LNA" or "LNA nucleosides" means nucleic acid monomers having a bridge connecting two carbon atoms between the 4' and 2'position of the nucleoside sugar unit, thereby forming a bicyclic sugar. Examples of such bicyclic sugar include, but are not limited to A) .alpha.-L-Methyleneoxy (4'-CH.sub.2--O-2') LNA, (B) .beta.-D-Methyleneoxy (4'-CH.sub.2--O-2') LNA, (C) Ethyleneoxy (4'-(CH.sub.2).sub.2--O-2') LNA, (D) Aminooxy (4'-CH.sub.2--O--N(R)-2') LNA and (E) Oxyamino (4'-CH.sub.2--N(R)--O-2') LNA, as depicted below.
##STR00001##
[0091] As used herein, LNA compounds include, but are not limited to, compounds having at least one bridge between the 4' and the 2' position of the sugar wherein each of the bridges independently comprises 1 or from 2 to 4 linked groups independently selected from --[C(R.sub.1)(R.sub.2)].sub.n--, --C(R.sub.1).dbd.C(R.sub.2)--, --C(R.sub.1).dbd.N--, --C(.dbd.NR.sub.1)--, --C(.dbd.O)--, --C(.dbd.S)--, --O--, --Si(R.sub.1).sub.2--, --S(.dbd.O)-- and --N(R.sub.1)--; wherein: x is 0, 1, or 2; n is 1, 2, 3, or 4; each R.sub.1 and R.sub.2 is, independently, H, a protecting group, hydroxyl, C.sub.1-C.sub.12 alkyl, substituted C.sub.1-C.sub.12 alkyl, C.sub.2-C.sub.12 alkenyl, substituted C.sub.2-C.sub.12 alkenyl, C.sub.2-C.sub.12 alkynyl, substituted C.sub.2-C.sub.12 alkynyl, C.sub.5-C.sub.20 aryl, substituted C.sub.5-C.sub.20 aryl, a heterocycle radical, a substituted heterocycle radical, heteroaryl, substituted heteroaryl, C.sub.5-C.sub.7 alicyclic radical, substituted C.sub.5-C.sub.7 alicyclic radical, halogen, OJ.sub.1, NJ.sub.1J.sub.2, SJ.sub.1, N.sub.3, COOJ.sub.1, acyl (C(.dbd.O)--H), substituted acyl, CN, sulfonyl (S(.dbd.O).sub.2-J.sub.1), or sulfoxyl (S(.dbd.O)-J.sub.1); and each J.sub.1 and J.sub.2 is, independently, H, C.sub.1-C.sub.12 alkyl, substituted C.sub.1-C.sub.12 alkyl, C.sub.2-C.sub.12 alkenyl, substituted C.sub.2-C.sub.12 alkenyl, C.sub.2-C.sub.12 alkynyl, substituted C.sub.2-C.sub.12 alkynyl, C.sub.5-C.sub.20 aryl, substituted C.sub.5-C.sub.20 aryl, acyl (C(.dbd.O)--H), substituted acyl, a heterocycle radical, a substituted heterocycle radical, C.sub.1-C.sub.12 aminoalkyl, substituted C.sub.1-C.sub.12 aminoalkyl or a protecting group.
[0092] Examples of 4'-2' bridging groups encompassed within the definition of LNA include, but are not limited to one of formulae: --[C(R.sub.1)(R.sub.2)].sub.n--, --[C(R.sub.1)(R.sub.2)].sub.n--O--, --C(R.sub.1R.sub.2)--N(R.sub.1)--O-- or --C(R.sub.1R.sub.2)--O--N(R.sub.1)--. Furthermore, other bridging groups encompassed with the definition of LNA are 4'-CH.sub.2-2',4'-(CH.sub.2).sub.2-2',4'-(CH.sub.2).sub.3-2',4'-CH.sub.2-- -O-2',4'-(CH.sub.2).sub.2--O-2',4'-CH.sub.2--O--N(R.sub.1)-2' and 4'-CH.sub.2--N(R.sub.1)--O-2'- bridges, wherein each R.sub.1 and R.sub.2 is, independently, H, a protecting group or C.sub.1-C.sub.12 alkyl.
[0093] Also included within the definition of LNA according to the invention are LNAs in which the 2'-hydroxyl group of the ribosyl sugar ring is connected to the 4' carbon atom of the sugar ring, thereby forming a methyleneoxy (4'-CH.sub.2--O-2') bridge to form the bicyclic sugar moiety. The bridge can also be a methylene (--CH.sub.2--) group connecting the 2' oxygen atom and the 4' carbon atom, for which the term methyleneoxy (4'-CH.sub.2--O-2') LNA is used. Furthermore; in the case of the bicylic sugar moiety having an ethylene bridging group in this position, the term ethyleneoxy (4'-CH.sub.2CH.sub.2--O-2') LNA is used. .alpha.-L-methyleneoxy (4'-CH.sub.2--O-2'), an isomer of methyleneoxy (4'-CH.sub.2--O-2') LNA is also encompassed within the definition of LNA, as used herein.
[0094] "Mismatch" or "non-complementary nucleobase" refers to the case when a nucleobase of a first nucleic acid is not capable of pairing with the corresponding nucleobase of a second or target nucleic acid.
[0095] "Modified internucleoside linkage" refers to a substitution or any change from a naturally occurring internucleoside bond (i.e. a phosphodiester internucleoside bond).
[0096] "Modified nucleobase" means any nucleobase other than adenine, cytosine, guanine, thymidine (also known as 5-methyluracil), or uracil. An "unmodified nucleobase" means the purine bases adenine (A) and guanine (G), and the pyrimidine bases thymine (T), cytosine (C), and uracil (U).
[0097] "Modified nucleoside" means a nucleoside having, independently, a modified sugar moiety and/or modified nucleobase.
[0098] "Modified nucleotide" means a nucleotide having, independently, a modified sugar moiety, modified internucleoside linkage, and/or modified nucleobase.
[0099] "Modified oligonucleotide" means an oligonucleotide comprising at least one modified internucleoside linkage, modified sugar, and/or modified nucleobase.
[0100] "Modified sugar" means substitution and/or any change from a natural sugar moiety.
[0101] "Mono or polycyclic ring system" is meant to include all ring systems selected from single or polycyclic radical ring systems wherein the rings are fused or linked and is meant to be inclusive of single and mixed ring systems individually selected from aliphatic, alicyclic, aryl, heteroaryl, aralkyl, arylalkyl, heterocyclic, heteroaryl, heteroaromatic and heteroarylalkyl. Such mono and poly cyclic structures can contain rings that each have the same level of saturation or each, independently, have varying degrees of saturation including fully saturated, partially saturated or fully unsaturated. Each ring can comprise ring atoms selected from C, N, O and S to give rise to heterocyclic rings as well as rings comprising only C ring atoms which can be present in a mixed motif such as for example benzimidazole wherein one ring has only carbon ring atoms and the fused ring has two nitrogen atoms. The mono or polycyclic ring system can be further substituted with substituent groups such as for example phthalimide which has two .dbd.O groups attached to one of the rings. Mono or polycyclic ring systems can be attached to parent molecules using various strategies such as directly through a ring atom, fused through multiple ring atoms, through a substituent group or through a bifunctional linking moiety.
[0102] "Monomer" means a single unit of an oligomer. Monomers include, but are not limited to, nucleosides and nucleotides, whether naturally occurring or modified.
[0103] "Motif" means the pattern of unmodified and modified nucleosides in an antisense compound.
[0104] "Natural sugar moiety" means a sugar moiety found in DNA (2'-H) or RNA (2'-OH).
[0105] "Naturally occurring internucleoside linkage" means a 3' to 5' phosphodiester linkage.
[0106] "Neutral linking group" means a linking group that is not charged. Neutral linking groups include without limitation phosphotriesters, methylphosphonates, MMI (--CH.sub.2--N(CH.sub.3)--O--), amide-3 (--CH.sub.2--C(.dbd.O)--N(H)--), amide-4 (--CH.sub.2--N(H)--C(.dbd.O)--), formacetal (--O--CH.sub.2--O--), and thioformacetal (--S--CH.sub.2--O--). Further neutral linking groups include nonionic linkages comprising siloxane (dialkylsiloxane), carboxylate ester, carboxamide, sulfide, sulfonate ester and amides (See for example: Carbohydrate Modifications in Antisense Research; Y. S. Sanghvi and P. D. Cook Eds. ACS Symposium Series 580; Chapters 3 and 4, (pp. 40-65)). Further neutral linking groups include nonionic linkages comprising mixed N, O, S and CH.sub.2 component parts.
[0107] "Non-complementary nucleobase" refers to a pair of nucleobases that do not form hydrogen bonds with one another or otherwise support hybridization.
[0108] "Non-internucleoside neutral linking group" means a neutral linking group that does not directly link two nucleosides. In certain embodiments, a non-internucleoside neutral linking group links a nucleoside to a group other than a nucleoside. In certain embodiments, a non-internucleoside neutral linking group links two groups, neither of which is a nucleoside.
[0109] "Non-internucleoside phosphorus linking group" means a phosphorus linking group that does not directly link two nucleosides. In certain embodiments, a non-internucleoside phosphorus linking group links a nucleoside to a group other than a nucleoside. In certain embodiments, a non-internucleoside phosphorus linking group links two groups, neither of which is a nucleoside.
[0110] "Nucleic acid" refers to molecules composed of monomeric nucleotides. A nucleic acid includes, but is not limited to, ribonucleic acids (RNA), deoxyribonucleic acids (DNA), single-stranded nucleic acids, double-stranded nucleic acids, small interfering ribonucleic acids (siRNA), and microRNAs (miRNA).
[0111] "Nucleobase" means a heterocyclic moiety capable of pairing with a base of another nucleic acid.
[0112] "Nucleobase complementarity" refers to a nucleobase that is capable of base pairing with another nucleobase. For example, in DNA, adenine (A) is complementary to thymine (T). For example, in RNA, adenine (A) is complementary to uracil (U). In certain embodiments, complementary nucleobase refers to a nucleobase of an antisense compound that is capable of base pairing with a nucleobase of its target nucleic acid. For example, if a nucleobase at a certain position of an antisense compound is capable of hydrogen bonding with a nucleobase at a certain position of a target nucleic acid, then the position of hydrogen bonding between the oligonucleotide and the target nucleic acid is considered to be complementary at that nucleobase pair.
[0113] "Nucleobase modification motif" means a pattern of modifications to nucleobases along an oligonucleotide. Unless otherwise indicated, a nucleobase modification motif is independent of the nucleobase sequence.
[0114] "Nucleobase sequence" means the order of contiguous nucleobases independent of any sugar, linkage, and/or nucleobase modification.
[0115] "Nucleoside" means a nucleobase linked to a sugar.
[0116] "Nucleoside mimetic" includes those structures used to replace the sugar or the sugar and the base and not necessarily the linkage at one or more positions of an oligomeric compound such as for example nucleoside mimetics having morpholino, cyclohexenyl, cyclohexyl, tetrahydropyranyl, bicyclo, or tricyclo sugar mimetics, e.g., non furanose sugar units. Nucleotide mimetic includes those structures used to replace the nucleoside and the linkage at one or more positions of an oligomeric compound such as for example peptide nucleic acids or morpholinos (morpholinos linked by --N(H)--C(.dbd.O)--O-- or other non-phosphodiester linkage). Sugar surrogate overlaps with the slightly broader term nucleoside mimetic but is intended to indicate replacement of the sugar unit (furanose ring) only. The tetrahydropyranyl rings provided herein are illustrative of an example of a sugar surrogate wherein the furanose sugar group has been replaced with a tetrahydropyranyl ring system. "Mimetic" refers to groups that are substituted for a sugar, a nucleobase, and/or internucleoside linkage. Generally, a mimetic is used in place of the sugar or sugar-internucleoside linkage combination, and the nucleobase is maintained for hybridization to a selected target.
[0117] "Nucleoside motif" means a pattern of nucleoside modifications in an oligonucleotide or a region thereof. The linkages of such an oligonucleotide may be modified or unmodified. Unless otherwise indicated, motifs herein describing only nucleosides are intended to be nucleoside motifs. Thus, in such instances, the linkages are not limited.
[0118] "Nucleotide" means a nucleoside having a phosphate group covalently linked to the sugar portion of the nucleoside.
[0119] "Off-target effect" refers to an unwanted or deleterious biological effect associated with modulation of RNA or protein expression of a gene other than the intended target nucleic acid.
[0120] "Oligomeric compound" or "oligomer" means a polymer of linked monomeric subunits which is capable of hybridizing to at least a region of a nucleic acid molecule.
[0121] "Oligonucleotide" means a polymer of linked nucleosides each of which can be modified or unmodified, independent one from another.
[0122] "Parenteral administration" means administration through injection (e.g., bolus injection) or infusion. Parenteral administration includes subcutaneous administration, intravenous administration, intramuscular administration, intraarterial administration, intraperitoneal administration, or intracranial administration, e.g., intrathecal or intracerebroventricular administration.
[0123] "Peptide" means a molecule formed by linking at least two amino acids by amide bonds. Without limitation, as used herein, peptide refers to polypeptides and proteins.
[0124] "Pharmaceutical agent" means a substance that provides a therapeutic benefit when administered to an individual. For example, in certain embodiments, an antisense oligonucleotide targeted to PKK is a pharmaceutical agent.
[0125] "Pharmaceutical composition" means a mixture of substances suitable for administering to a subject. For example, a pharmaceutical composition may comprise an antisense oligonucleotide and a sterile aqueous solution.
[0126] "Pharmaceutically acceptable derivative" encompasses pharmaceutically acceptable salts, conjugates, prodrugs or isomers of the compounds described herein.
[0127] "Pharmaceutically acceptable salts" means physiologically and pharmaceutically acceptable salts of antisense compounds, i.e., salts that retain the desired biological activity of the parent oligonucleotide and do not impart undesired toxicological effects thereto.
[0128] "Phosphorothioate linkage" means a linkage between nucleosides where the phosphodiester bond is modified by replacing one of the non-bridging oxygen atoms with a sulfur atom. A phosphorothioate linkage is a modified internucleoside linkage.
[0129] "Phosphorus linking group" means a linking group comprising a phosphorus atom. Phosphorus linking groups include without limitation groups having the formula:
##STR00002##
wherein:
[0130] R.sub.a and R.sub.d are each, independently, O, S, CH.sub.2, NH, or NJ.sub.1 wherein J.sub.1 is C.sub.1-C.sub.6 alkyl or substituted C.sub.1-C.sub.6 alkyl;
[0131] R.sub.b is O or S;
[0132] R.sub.c is OH, SH, C.sub.1-C.sub.6 alkyl, substituted C.sub.1-C.sub.6 alkyl, C.sub.1-C.sub.6 alkoxy, substituted C.sub.1-C.sub.6 alkoxy, amino or substituted amino; and
[0133] J.sub.1 is R.sub.b is O or S.
Phosphorus linking groups include without limitation, phosphodiester, phosphorothioate, phosphorodithioate, phosphonate, phosphoramidate, phosphorothioamidate, thionoalkylphosphonate, phosphotriesters, thionoalkylphosphotriester and boranophosphate.
[0134] "PKK" means mammalian plasma prekallikrein, including human plasma prekallikrein. Plasma prekallikrein (PKK) is the precursor of plasma kallikrein (PK), which is encoded by the KLKB1 gene.
[0135] "PKK associated disease" means any disease associated with any PKK nucleic acid or expression product thereof. Such diseases may include an inflammatory disease or a thromboembolic disease. Such diseases may include hereditary angioedema (HAE).
[0136] "PKK mRNA" means any messenger RNA expression product of a DNA sequence encoding PKK.
[0137] "PKK nucleic acid" means any nucleic acid encoding PKK. For example, in certain embodiments, a PKK nucleic acid includes a DNA sequence encoding PKK, an RNA sequence transcribed from DNA encoding PKK (including genomic DNA comprising introns and exons), and an mRNA sequence encoding PKK. "PKK mRNA" means an mRNA encoding a PKK protein.
[0138] "PKK protein" means the polypeptide expression product of a PKK nucleic acid.
[0139] "Portion" means a defined number of contiguous (i.e., linked) nucleobases of a nucleic acid. In certain embodiments, a portion is a defined number of contiguous nucleobases of a target nucleic acid. In certain embodiments, a portion is a defined number of contiguous nucleobases of an antisense compound.
[0140] "Prevent" or "preventing" refers to delaying or forestalling the onset or development of a disease, disorder, or condition for a period of time from minutes to days, weeks to months, or indefinitely.
[0141] "Prodrug" means a therapeutic agent that is prepared in an inactive form that is converted to an active form (i.e., drug) within the body or cells thereof by the action of endogenous enzymes or other chemicals and/or conditions.
[0142] "Prophylactically effective amount" refers to an amount of a pharmaceutical agent that provides a prophylactic or preventative benefit to an animal.
[0143] "Protecting group" means any compound or protecting group known to those having skill in the art. Non-limiting examples of protecting groups may be found in "Protective Groups in Organic Chemistry", T. W. Greene, P. G. M. Wuts, ISBN 0-471-62301-6, John Wiley & Sons, Inc, New York, which is incorporated herein by reference in its entirety.
[0144] "Region" is defined as a portion of the target nucleic acid having at least one identifiable structure, function, or characteristic.
[0145] "Ribonucleotide" means a nucleotide having a hydroxy at the 2' position of the sugar portion of the nucleotide. Ribonucleotides may be modified with any of a variety of substituents.
[0146] "RISC based antisense compound" means an antisense compound wherein at least some of the antisense activity of the antisense compound is attributable to the RNA Induced Silencing Complex (RISC).
[0147] "RNase H based antisense compound" means an antisense compound wherein at least some of the antisense activity of the antisense compound is attributable to hybridization of the antisense compound to a target nucleic acid and subsequent cleavage of the target nucleic acid by RNase H.
[0148] "Salts" mean a physiologically and pharmaceutically acceptable salts of antisense compounds, i.e., salts that retain the desired biological activity of the parent oligonucleotide and do not impart undesired toxicological effects thereto.
[0149] "Segments" are defined as smaller or sub-portions of regions within a target nucleic acid.
[0150] "Separate regions" means portions of an oligonucleotide wherein the chemical modifications or the motif of chemical modifications of any neighboring portions include at least one difference to allow the separate regions to be distinguished from one another.
[0151] "Sequence motif" means a pattern of nucleobases arranged along an oligonucleotide or portion thereof. Unless otherwise indicated, a sequence motif is independent of chemical modifications and thus may have any combination of chemical modifications, including no chemical modifications.
[0152] "Side effects" means physiological responses attributable to a treatment other than desired effects. In certain embodiments, side effects include, without limitation, injection site reactions, liver function test abnormalities, renal function abnormalities, liver toxicity, renal toxicity, central nervous system abnormalities, and myopathies.
[0153] "Single-stranded oligonucleotide" means an oligonucleotide which is not hybridized to a complementary strand.
[0154] "Sites," as used herein, are defined as unique nucleobase positions within a target nucleic acid.
[0155] "Specifically hybridizable" or "specifically hybridizes" refers to an antisense compound having a sufficient degree of complementarity between an antisense oligonucleotide and a target nucleic acid to induce a desired effect, while exhibiting minimal or no effects on non-target nucleic acids under conditions in which specific binding is desired, i.e., under physiological conditions in the case of in vivo assays and therapeutic treatments.
[0156] "Stringent hybridization conditions" or "stringent conditions" refer to conditions under which an oligomeric compound will hybridize to its target sequence, but to a minimal number of other sequences.
[0157] "Subject" means a human or non-human animal selected for treatment or therapy.
[0158] "Substituent" and "substituent group," means an atom or group that replaces the atom or group of a named parent compound. For example a substituent of a modified nucleoside is any atom or group that differs from the atom or group found in a naturally occurring nucleoside (e.g., a modified 2'-substuent is any atom or group at the 2'-position of a nucleoside other than H or OH). Substituent groups can be protected or unprotected. In certain embodiments, compounds of the present disclosure have substituents at one or at more than one position of the parent compound. Substituents may also be further substituted with other substituent groups and may be attached directly or via a linking group such as an alkyl or hydrocarbyl group to a parent compound. Likewise, as used herein, "substituent" in reference to a chemical functional group means an atom or group of atoms that differs from the atom or a group of atoms normally present in the named functional group. In certain embodiments, a substituent replaces a hydrogen atom of the functional group (e.g., in certain embodiments, the substituent of a substituted methyl group is an atom or group other than hydrogen which replaces one of the hydrogen atoms of an unsubstituted methyl group). Unless otherwise indicated, groups amenable for use as substituents include without limitation, halogen, hydroxyl, alkyl, alkenyl, alkynyl, acyl (--C(O)R.sub.aa), carboxyl (--C(O)O--R.sub.aa), aliphatic groups, alicyclic groups, alkoxy, substituted oxy (--O--R.sub.aa), aryl, aralkyl, heterocyclic radical, heteroaryl, heteroarylalkyl, amino (--N(R.sub.bb)(R.sub.cc)), imino (.dbd.NR.sub.bb), amido (--C(O)N--(R.sub.bb)(R.sub.cc) or --N(R.sub.bb)C(O)R.sub.aa), azido (--N.sub.3), nitro (--NO.sub.2), cyano (--CN), carbamido (--OC(O)N(R.sub.bb)(R.sub.c) or --N(R.sub.bb)C(O)OR.sub.aa), ureido (--N(R.sub.bb)C(O)N(R.sub.bb)(R.sub.cc)), thioureido (--N(R.sub.bb)C(S)N(R.sub.bb)(R.sub.cc)), guanidinyl (--N(R.sub.bb)C(.dbd.NR.sub.bb)N(R.sub.bb)(R.sub.cc)), amidinyl (--C(.dbd.NR.sub.bb)N(R.sub.bb)(R.sub.c) or --N(R.sub.bb)C(.dbd.NR.sub.bb)(R.sub.aa)), thiol (--SR.sub.bb), sulfinyl (--S(O)R.sub.bb), sulfonyl (--S(O).sub.2R.sub.bb) and sulfonamidyl (--S(O).sub.2N(R.sub.bb)(R.sub.cc) or --N(R.sub.bb)S(O).sub.2R.sub.bb). Wherein each R.sub.aa, R.sub.bb and R.sub.cc is, independently, H, an optionally linked chemical functional group or a further substituent group with a preferred list including without limitation, alkyl, alkenyl, alkynyl, aliphatic, alkoxy, acyl, aryl, aralkyl, heteroaryl, alicyclic, heterocyclic and heteroarylalkyl. Selected substituents within the compounds described herein are present to a recursive degree.
[0159] "Substituted sugar moiety" means a furanosyl that is not a naturally occurring sugar moiety. Substituted sugar moieties include, but are not limited to furanosyls comprising substituents at the 2'-position, the 3'-position, the 5'-position and/or the 4'-position. Certain substituted sugar moieties are bicyclic sugar moieties.
[0160] "Sugar moiety" means a naturally occurring sugar moiety or a modified sugar moiety of a nucleoside.
[0161] "Sugar motif" means a pattern of sugar modifications in an oligonucleotide or a region thereof.
[0162] "Sugar surrogate" means a structure that does not comprise a furanosyl and that is capable of replacing the naturally occurring sugar moiety of a nucleoside, such that the resulting nucleoside sub-units are capable of linking together and/or linking to other nucleosides to form an oligomeric compound which is capable of hybridizing to a complementary oligomeric compound. Such structures include rings comprising a different number of atoms than furanosyl (e.g., 4, 6, or 7-membered rings); replacement of the oxygen of a furanosyl with a non-oxygen atom (e.g., carbon, sulfur, or nitrogen); or both a change in the number of atoms and a replacement of the oxygen. Such structures may also comprise substitutions corresponding to those described for substituted sugar moieties (e.g., 6-membered carbocyclic bicyclic sugar surrogates optionally comprising additional substituents). Sugar surrogates also include more complex sugar replacements (e.g., the non-ring systems of peptide nucleic acid). Sugar surrogates include without limitation morpholinos, cyclohexenyls and cyclohexitols.
[0163] "Target" refers to a protein, the modulation of which is desired.
[0164] "Target gene" refers to a gene encoding a target.
[0165] "Targeting" or "targeted" means the process of design and selection of an antisense compound that will specifically hybridize to a target nucleic acid and induce a desired effect.
[0166] "Target nucleic acid," "target RNA," and "target RNA transcript" and "nucleic acid target" all mean a nucleic acid capable of being targeted by antisense compounds.
[0167] "Target region" means a portion of a target nucleic acid to which one or more antisense compounds is targeted.
[0168] "Target segment" means the sequence of nucleotides of a target nucleic acid to which an antisense compound is targeted. "5' target site" refers to the 5'-most nucleotide of a target segment. "3' target site" refers to the 3'-most nucleotide of a target segment.
[0169] "Terminal group" means one or more atom attached to either, or both, the 3' end or the 5' end of an oligonucleotide. In certain embodiments a terminal group is a conjugate group. In certain embodiments, a terminal group comprises one or more terminal group nucleosides.
[0170] "Terminal internucleoside linkage" means the linkage between the last two nucleosides of an oligonucleotide or defined region thereof.
[0171] "Therapeutically effective amount" means an amount of a pharmaceutical agent that provides a therapeutic benefit to an individual.
[0172] "Treat" or "treating" or "treatment" refers to administering a composition to effect an improvement of the disease or condition.
[0173] "Type of modification" in reference to a nucleoside or a nucleoside of a "type" means the chemical modification of a nucleoside and includes modified and unmodified nucleosides. Accordingly, unless otherwise indicated, a "nucleoside having a modification of a first type" may be an unmodified nucleoside.
[0174] "Unmodified nucleobases" mean the purine bases adenine (A) and guanine (G), and the pyrimidine bases thymine (T), cytosine (C) and uracil (U).
[0175] "Unmodified nucleotide" means a nucleotide composed of naturally occurring nucleobases, sugar moieties, and internucleoside linkages. In certain embodiments, an unmodified nucleotide is an RNA nucleotide (i.e. (3-D-ribonucleosides) or a DNA nucleotide (i.e. 13-D-deoxyribonucleoside).
[0176] "Upstream" refers to the relative direction toward the 5' end or N-terminal end of a nucleic acid.
[0177] "Wing segment" means a plurality of nucleosides modified to impart to an oligonucleotide properties such as enhanced inhibitory activity, increased binding affinity for a target nucleic acid, or resistance to degradation by in vivo nucleases.
CERTAIN EMBODIMENTS
[0178] Certain embodiments provide compounds, compositions, and methods for inhibiting plasma prekallikrein (PKK) mRNA and protein expression. Certain embodiments provide compounds, compositions, and methods for decreasing PKK mRNA and protein levels.
[0179] Certain embodiments provide antisense compounds targeted to a plasma prekallikrein (PKK) nucleic acid. In certain embodiments, the PKK nucleic acid is the sequence set forth in GENBANK Accession No. NM_000892.3 (incorporated herein as SEQ ID NO: 1), GENBANK Accession No. DC412984.1 (incorporated herein as SEQ ID NO: 2), GENBANK Accession No. CN265612.1 (incorporated herein as SEQ ID NO: 3), GENBANK Accession No. AK297672.1 (incorporated herein as SEQ ID NO: 4), GENBANK Accession No. DC413312.1 (incorporated herein as SEQ ID NO: 5), GENBANK Accession No. AV688858.2 (incorporated herein as SEQ ID NO: 6), GENBANK Accession No. CD652077.1 (incorporated herein as SEQ ID NO: 7), GENBANK Accession No. BC143911.1 (incorporated herein as SEQ ID NO: 8), GENBANK Accession No. CB162532.1 (incorporated herein as SEQ ID NO: 9), GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000 (incorporated herein as SEQ ID NO: 10), GENBANK Accession No. NM_008455.2 (incorporated herein as SEQ ID NO: 11), GENBANK Accession No. BB598673.1 (incorporated herein as SEQ ID NO: 12), GENBANK Accession No. NT_039460.7 truncated from nucleobases 6114001 to U.S. Pat. No. 6,144,000 (incorporated herein as SEQ ID NO: 13), GENBANK Accession No. NM_012725.2 (incorporated herein as SEQ ID NO: 14), GENBANK Accession No. NW_047473.1 truncated from nucleobases 10952001 to Ser. No. 10/982,000 (incorporated herein as SEQ ID NO: 15), GENBANK Accession No. XM_002804276.1 (incorporated herein as SEQ ID NO: 17), and GENBANK Accession No. NW_001118167.1 truncated from nucleobases 2358000 to U.S. Pat. No. 2,391,000 (incorporated herein as SEQ ID NO: 18).
[0180] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 30-2226.
[0181] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases of the nucleobase sequence of SEQ ID NO: 570.
[0182] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases of the nucleobase sequence of SEQ ID NO: 705.
[0183] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of the nucleobase sequence of SEQ ID NO: 1666.
[0184] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 20 linked nucleosides and has the nucleobase sequence of SEQ ID NO: 570.
[0185] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 20 linked nucleosides and has the nucleobase sequence of SEQ ID NO: 705.
[0186] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 16 linked nucleosides and has the nucleobase sequence of SEQ ID NO: 1666.
[0187] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 62, 72, 103, 213, 312, 334-339, 344, 345, 346, 348, 349, 351, 369, 373, 381, 382, 383, 385, 387-391, 399, 411, 412, 414, 416, 444, 446-449, 452, 453, 454, 459, 460, 462-472, 473, 476, 477, 479, 480, 481, 484, 489-495, 497, 500, 504, 506, 522, 526, 535, 558, 559, 560, 564, 566, 568-571, 573, 576, 577, 578, 587, 595, 597-604, 607, 608, 610, 613, 615, 618, 619, 622, 623, 624, 633, 635, 636, 638, 639, 640, 642, 643, 645, 652, 655-658, 660, 661, 670, 674-679, 684, 685, 698, 704, 705, 707, 708, 713, 716, 717, 728, 734, 736, 767, 768, 776, 797, 798, 800, 802, 810, 815, 876, 880, 882, 883, 886, 891, 901-905, 908-911, 922, 923, 924, 931, 942, 950-957, 972, 974, 978, 979, 980, 987-991, 1005, 1017-1021, 1025, 1026, 1029, 1030, 1032, 1034, 1035, 1037, 1040, 1041, 1045, 1046, 1051, 1054, 1059, 1060, 1061, 1064, 1065, 1066, 1075, 1076, 1087, 1089, 1111, 1114, 1116, 1117, 1125, 1133, 1153, 1169, 1177, 1181, 1182, 1187, 1196, 1200, 1214, 1222, 1267, 1276, 1277, 1285, 1286, 1289, 1290, 1291, 1303, 1367, 1389, 1393, 1398-1401, 1406, 1407, 1408, 1411, 1419-1422, 1426, 1430, 1431, 1432, 1434-1437, 1439, 1440, 1443, 1444, 1451, 1452, 1471, 1516, 1527, 1535, 1537, 1538, 1539, 1540, 1541, 1563, 1564, 1567, 1568, 1616, 1617, 1623, 1629, 1664, 1665, 1666, 1679, 1687, 1734, 1804, 1876, 1886, 1915, 2008, 2018, 2100, 2101, 2115, and 2116. In certain embodiments, the modified oligonucleotide achieves at least 80% mRNA inhibition of PKK.
[0188] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 62, 72, 103, 213, 334-339, 344, 346, 348, 349, 351, 381, 382, 383, 385, 389, 390, 391, 446, 448, 452, 453, 454, 466-473, 476, 481, 484, 491, 492, 494, 495, 497, 504, 526, 558, 559, 566, 568-571, 576, 578, 587, 595, 597, 598, 600-604, 607, 610, 613, 618, 619, 624, 635, 638, 639, 645, 652, 656, 657, 658, 660, 674, 675, 676, 684, 698, 704, 705, 707, 713, 716, 768, 876, 880, 901-905, 908-911, 922, 923, 924, 931, 942, 951, 954-957, 972, 974, 978, 979, 987, 988, 990, 1005, 1019, 1020, 1021, 1025, 1032, 1037, 1040, 1041, 1045, 1054, 1059, 1060, 1061, 1064, 1065, 1066, 1075, 1111, 1116, 1117, 1125, 1133, 1153, 1169, 1177, 1200, 1222, 1267, 1285, 1290, 1291, 1303, 1367, 1398, 1399, 1401, 1406, 1408, 1411, 1419, 1420, 1421, 1426, 1430, 1431, 1432, 1434-1437, 1440, 1443, 1444, 1451, 1537-1540, 1563, 1616, 1679, 1687, 1804, 2008, 2101, 2115, and 2116. In certain embodiments, the modified oligonucleotide achieves at least 85% mRNA inhibition of PKK.
[0189] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 334, 346, 351, 382, 390, 391, 446, 448, 452, 453, 468, 469, 470, 471, 472, 476, 481, 491, 495, 504, 558, 566, 568, 570, 571, 578, 587, 597, 598, 600, 604, 613, 635, 638, 645, 656, 658, 660, 674, 675, 684, 704, 705, 880, 901-905, 909, 922, 931, 951, 954, 956, 990, 1005, 1020, 1032, 1037, 1040, 1041, 1045, 1054, 1075, 1111, 1125, 1133, 1153, 1200, 1267, 1291, 1303, 1398, 1399, 1401, 1406, 1420, 1426, 1430, 1431, 1434, 1435, 1436, 1440, 1443, 1451, 1537-1540, 2115, and 2116. In certain embodiments, the modified oligonucleotide achieves at least 90% mRNA inhibition of PKK.
[0190] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 334, 391, 448, 468, 469, 568, 570, 598, 635, 658, 674, 684, 705, 901, 903, 904, 922, 990, 1267, 1291, 1420, 1430, 1431, 1434, 1435, 1436, 1537, 1538, and 1540. In certain embodiments, the modified oligonucleotide achieves at least 95% mRNA inhibition of PKK.
[0191] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 334, 338, 346, 349, 382, 383, 390, 448, 452, 453, 454, 495, 526, 559, 570, 587, 598, 635, 660, 705, 901, 903, 904, 908, 923, 931, 955, 974, 988, 990, 1020, 1039, 1040, 1111, 1117, 1267, 1291, 1349, 1352, 1367, 1389, 1393, 1399, 1401, 1408, 1411, 1426, 1499, 1516, 1535, 1544, 1548, 1563, 1564, 1568, 1569, 1598, 1616, 1617, 1623, 1624, 1643, 1661, 1665, 1666, 1673, 1679, 1695, 1720, 1804, 1817, 1876, 1881, 1886, 1940, 1947, 2008, 2018, 2019, 2031, 2044, 2100, 2101, 2115, and 2116. In certain embodiments, the modified oligonucleotide achieves an IC.sub.50 (.mu.M) of 0.4 or less.
[0192] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 334, 346, 349, 382, 453, 454, 495, 526, 570, 587, 598, 635, 660, 901, 903, 904, 931, 955, 990, 1020, 1111, 1267, 1349, 1352, 1367, 1389, 1399, 1408, 1411, 1426, 1516, 1535, 1544, 1548, 1563, 1564, 1568, 1569, 1598, 1616, 1617, 1623, 1643, 1661, 1665, 1666, 1673, 1695, 1804, 1876, 1881, 2019, 2044, 2100, 2101, 2115, and 2116. In certain embodiments, the modified oligonucleotide achieves an IC.sub.50 (.mu.M) of 0.3 or less.
[0193] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 334, 346, 382, 453, 495, 526, 570, 587, 598, 635, 901, 904, 931, 955, 1020, 1111, 1349, 1352, 1389, 1426, 1516, 1535, 1544, 1548, 1564, 1569, 1598, 1616, 1617, 1665, 1666, 1804, 1876, 1881, 2019, 2044, 2101, and 2116. In certain embodiments, the modified oligonucleotide achieves an IC.sub.50 (.mu.M) of 0.2 or less.
[0194] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and has a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, or at least 16 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 334, 495, 587, 598, 635, 1349, 1352, 1389, 1516, 1544, 1548, 1569, 1598, 1617, 1665, 1666, 1804, 1881, and 2019. In certain embodiments, the modified oligonucleotide achieves an IC.sub.50 (.mu.M) of less than 0.2.
[0195] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 27427-27466 of SEQ ID NO: 10.
[0196] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 33183-33242 of SEQ ID NO: 10.
[0197] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 30570-30610 of SEQ ID NO: 10.
[0198] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 27427-27520 of SEQ ID NO: 10.
[0199] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 33085-33247 of SEQ ID NO: 10.
[0200] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 30475-30639 of SEQ ID NO: 10.
[0201] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 27362-27524 of SEQ ID NO: 10.
[0202] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 33101-33240 of SEQ ID NO: 10.
[0203] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of nucleobases 30463-30638 of SEQ ID NO: 10.
[0204] Certain embodiments provide compounds, comprising a modified oligonucleotide and a conjugate group, wherein the modified oligonucleotide consists of 12 to 30 linked nucleosides and comprises a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases complementary to an equal length portion of exon 9, exon 12, or exon 14 of a PKK nucleic acid.
[0205] In certain embodiments the nucleobase sequence of the modified oligonucleotide is at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% complementary to SEQ ID NO: 10.
[0206] In certain embodiments, the compound consists of a single-stranded modified oligonucleotide.
[0207] In certain embodiments, at least one internucleoside linkage of the modified oligonucleotide is a modified internucleoside linkage.
[0208] In certain embodiments, at least one modified internucleoside linkage of the modified oligonucleotide is a phosphorothioate internucleoside linkage.
[0209] In certain embodiments, the modified oligonucleotide comprises at least 1, 2, 3, 4, 5, 6, or 7 phosphodiester internucleoside linkages.
[0210] In certain embodiments, each internucleoside linkage of the modified oligonucleotide is selected from a phosphodiester internucleoside linkage and a phosphorothioate internucleoside linkage.
[0211] In certain embodiments, each internucleoside linkage of the modified oligonucleotide is a phosphorothioate linkage.
[0212] In certain embodiments, at least one nucleoside of the modified oligonucleotide comprises a modified nucleobase.
[0213] In certain embodiments, the modified nucleobase is a 5-methylcytosine.
[0214] In certain embodiments, the modified oligonucleotide comprises at least one modified sugar.
[0215] In certain embodiments, the modified sugar is a 2' modified sugar, a BNA, or a THP.
[0216] In certain embodiments, the modified sugar is any of a 2'-O-methoxyethyl, 2'-O-methyl, a constrained ethyl, a LNA, or a 3'-fluoro-HNA.
[0217] In certain embodiments, the compound comprises at least one 2'-O-methoxyethyl nucleoside, 2'-O-methyl nucleoside, constrained ethyl nucleoside, LNA nucleoside, or 3'-fluoro-HNA nucleoside.
[0218] In certain embodiments, the modified oligonucleotide comprises:
[0219] a gap segment consisting of 10 linked deoxynucleosides;
[0220] a 5' wing segment consisting of 5 linked nucleosides; and
[0221] a 3' wing segment consisting of 5 linked nucleosides;
[0222] wherein the gap segment is positioned between the 5' wing segment and the 3' wing segment and wherein each nucleoside of each wing segment comprises a modified sugar.
[0223] In certain embodiments, the modified oligonucleotide consists of 20 linked nucleosides.
[0224] In certain embodiments, the modified oligonucleotide consists of 19 linked nucleosides.
[0225] In certain embodiments, the modified oligonucleotide consists of 18 linked nucleosides.
[0226] Certain embodiments provide compounds consisting of a conjugate group and a modified oligonucleotide according to the following formula: Tes Ges mCes Aes Aes Gds Tds mCds Tds mCds Tds Tds Gds Gds mCds Aes Aes Aes mCes Ae; wherein,
[0227] A=an adenine,
[0228] mC=a 5'-methylcytosine
[0229] G=a guanine,
[0230] T=a thymine,
[0231] e=a 2'-O-methoxyethyl modified nucleoside,
[0232] d=a 2'-deoxynucleoside, and
[0233] s=a phosphorothioate internucleoside linkage.
[0234] Certain embodiments provide compounds consisting of a conjugate group and a modified oligonucleotide according to the following formula: mCes mCes mCes mCes mCes Tds Tds mCds Tds Tds Tds Ads Tds Ads Gds mCes mCes Aes Ges mCe; wherein,
[0235] A=an adenine,
[0236] mC=a 5'-methylcytosine;
[0237] G=a guanine,
[0238] T=a thymine,
[0239] e=a 2'-O-methoxyethyl modified nucleoside,
[0240] d=a 2'-deoxynucleoside, and
[0241] s=a phosphorothioate internucleoside linkage.
[0242] Certain embodiments provide compounds consisting of a conjugate group and a modified oligonucleotide according to the following formula: mCes Ges Aks Tds Ads Tds mCds Ads Tds Gds Ads Tds Tds mCks mCks mCe; wherein,
[0243] A=an adenine,
[0244] mC=a 5'-methylcytosine;
[0245] G=a guanine,
[0246] T=a thymine,
[0247] e=a 2'-O-methoxyethyl modified nucleoside,
[0248] k=a cEt modified nucleoside,
[0249] d=a 2'-deoxynucleoside, and
[0250] s=a phosphorothioate internucleoside linkage.
[0251] In certain embodiments, the conjugate group is linked to the modified oligonucleotide at the 5' end of the modified oligonucleotide. In certain embodiments, the conjugate group is linked to the modified oligonucleotide at the 3' end of the modified oligonucleotide. In certain embodiments, the conjugate group comprises at least one N-Acetylgalactosamine (GalNAc), at least two N-Acetylgalactosamines (GalNAcs), or at least three N-Acetylgalactosamines (GalNAcs).
[0252] Certain embodiments provide compounds according to the following formula:
##STR00003##
[0253] Certain embodiments provide compounds according to the following formula:
##STR00004##
[0254] Certain embodiments provide compounds according to the following formula:
##STR00005##
[0255] In certain embodiments, a compound can comprise or consist of any modified oligonucleotide described herein and a conjugate group. In certain embodiments, a compound can comprise or consist of a modified oligonucleotide consisting of 12 to 30 linked nucleosides and having a nucleobase sequence comprising at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 consecutive nucleobases of any of the nucleobase sequences of SEQ ID NOs: 30-2226, and a conjugate group.
[0256] In certain embodiments, a compound having the following chemical structure comprises or consists of ISIS 721744 with a 5'-X, wherein X is a conjugate group comprising GalNAc as described herein:
##STR00006##
[0257] In certain embodiments, a compound having the following chemical structure comprises or consists of ISIS 546254 with a 5'-X, wherein X is a conjugate group comprising GalNAc as described herein:
##STR00007##
[0258] Certain embodiments provide a compound comprising or consisting of the following formula:
##STR00008##
[0259] Certain embodiments provide a compound comprising or consisting of the following formula:
##STR00009##
Certain embodiments provide a compound comprising or consisting of the following formula:
##STR00010##
wherein either R.sup.1 is --OCH.sub.2CH.sub.2OCH.sub.3 (MOE) and R.sup.2 is H; or R.sup.1 and R.sup.2 together form a bridge, wherein R.sup.1 is --O-- and R.sup.2 is --CH.sub.2--, --CH(CH.sub.3)--, or --CH.sub.2CH.sub.2--, and R.sup.1 and R.sup.2 are directly connected such that the resulting bridge is selected from: --O--CH.sub.2--, --O--CH(CH.sub.3)--, and --O--CH.sub.2CH.sub.2--; and for each pair of R.sup.3 and R.sup.4 on the same ring, independently for each ring: either R.sup.3 is selected from H and --OCH.sub.2CH.sub.2OCH.sub.3 and R.sup.4 is H; or R.sup.3 and R.sup.4 together form a bridge, wherein R.sup.3 is --O--, and R.sup.4 is --CH.sub.2--, --CH(CH.sub.3)--, or --CH.sub.2CH.sub.2-- and R.sup.3 and R.sup.4 are directly connected such that the resulting bridge is selected from: --O--CH.sub.2--, --O--CH(CH.sub.3)--, and --O--CH.sub.2CH.sub.2--; and R.sup.5 is selected from H and --CH.sub.3; and Z is selected from S.sup.- and O.sup.-.
[0260] Certain embodiments provide compositions comprising the compound of any preceding claim or salt thereof and at least one of a pharmaceutically acceptable carrier or diluent.
[0261] Certain embodiments provide methods comprising administering to an animal the compound or composition of any preceding claim.
[0262] In certain embodiments, the animal is a human.
[0263] In certain embodiments, administering the compound prevents, treats, or ameliorates a PKK associated disease, disorder or condition.
[0264] In certain embodiments, the PKK associated disease, disorder or condition is a hereditary angioedema (HAE), edema, angioedema, swelling, angioedema of the lids, ocular edema, macular edema, cerebral edema, thrombosis, embolism, thromboembolism, deep vein thrombosis, pulmonary embolism, myocardial infarction, stroke, or infarct.
[0265] Certain embodiments provide use of the compound or composition of any preceding claim for the manufacture of a medicament for treating an inflammatory disease or a thromboembolic disease.
Antisense Compounds
[0266] Oligomeric compounds include, but are not limited to, oligonucleotides, oligonucleosides, oligonucleotide analogs, oligonucleotide mimetics, antisense compounds, antisense oligonucleotides, and siRNAs. An oligomeric compound may be "antisense" to a target nucleic acid, meaning that is is capable of undergoing hybridization to a target nucleic acid through hydrogen bonding.
[0267] In certain embodiments, an antisense compound has a nucleobase sequence that, when written in the 5' to 3' direction, comprises the reverse complement of the target segment of a target nucleic acid to which it is targeted. In certain such embodiments, an antisense oligonucleotide has a nucleobase sequence that, when written in the 5' to 3' direction, comprises the reverse complement of the target segment of a target nucleic acid to which it is targeted.
[0268] In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 12 to 30 subunits in length. In certain embodiments, an antisense compound targeted to PKK nucleic acid is 12 to 25 subunits in length. In certain embodiments, an antisense compound targeted to PKK nucleic acid is 12 to 22 subunits in length. In certain embodiments, an antisense compound targeted to PKK nucleic acid is 14 to 20 subunits in length. In certain embodiments, an antisense compound targeted to PKK nucleic acid is 15 to 25 subunits in length. In certain embodiments, an antisense compound targeted to PKK nucleic acid is 18 to 22 subunits in length. In certain embodiments, an antisense compound targeted to PKK nucleic acid is 19 to 21 subunits in length. In certain embodiments, the antisense compound is 8 to 80, 12 to 50, 13 to 30, 13 to 50, 14 to 30, 14 to 50, 15 to 30, 15 to 50, 16 to 30, 16 to 50, 17 to 30, 17 to 50, 18 to 30, 18 to 50, 19 to 30, 19 to 50, or 20 to 30 linked subunits in length.
[0269] In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 12 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 13 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 14 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 15 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 16 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 17 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 18 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 19 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 20 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 21 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 22 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 23 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 24 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 25 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 26 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 27 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 28 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 29 subunits in length. In certain embodiments, an antisense compound targeted to a PKK nucleic acid is 30 subunits in length. In certain embodiments, the antisense compound targeted to a PKK nucleic acid is 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, or 80 linked subunits in length, or a range defined by any two of the above values. In certain embodiments the antisense compound is an antisense oligonucleotide, and the linked subunits are nucleosides.
[0270] In certain embodiments antisense oligonucleotides targeted to a PKK nucleic acid may be shortened or truncated. For example, a single subunit may be deleted from the 5' end (5' truncation), or alternatively from the 3' end (3' truncation). A shortened or truncated antisense compound targeted to a PKK nucleic acid may have two subunits deleted from the 5' end, or alternatively may have two subunits deleted from the 3' end, of the antisense compound. Alternatively, the deleted nucleosides may be dispersed throughout the antisense compound, for example, in an antisense compound having one nucleoside deleted from the 5' end and one nucleoside deleted from the 3' end.
[0271] When a single additional subunit is present in a lengthened antisense compound, the additional subunit may be located at the 5' or 3' end of the antisense compound. When two or more additional subunits are present, the added subunits may be adjacent to each other, for example, in an antisense compound having two subunits added to the 5' end (5' addition), or alternatively to the 3' end (3' addition), of the antisense compound. Alternatively, the added subunits may be dispersed throughout the antisense compound, for example, in an antisense compound having one subunit added to the 5' end and one subunit added to the 3' end.
[0272] It is possible to increase or decrease the length of an antisense compound, such as an antisense oligonucleotide, and/or introduce mismatch bases without eliminating activity. For example, in Woolf et al. (Proc. Natl. Acad. Sci. USA 89:7305-7309, 1992), a series of antisense oligonucleotides 13-25 nucleobases in length were tested for their ability to induce cleavage of a target RNA in an oocyte injection model. Antisense oligonucleotides 25 nucleobases in length with 8 or 11 mismatch bases near the ends of the antisense oligonucleotides were able to direct specific cleavage of the target mRNA, albeit to a lesser extent than the antisense oligonucleotides that contained no mismatches. Similarly, target specific cleavage was achieved using 13 nucleobase antisense oligonucleotides, including those with 1 or 3 mismatches.
[0273] Gautschi et al (J. Natl. Cancer Inst. 93:463-471, March 2001) demonstrated the ability of an oligonucleotide having 100% complementarity to the bcl-2 mRNA and having 3 mismatches to the bcl-xL mRNA to reduce the expression of both bcl-2 and bcl-xL in vitro and in vivo. Furthermore, this oligonucleotide demonstrated potent anti-tumor activity in vivo.
[0274] Maher and Dolnick (Nuc. Acid. Res. 16:3341-3358,1988) tested a series of tandem 14 nucleobase antisense oligonucleotides, and a 28 and 42 nucleobase antisense oligonucleotides comprised of the sequence of two or three of the tandem antisense oligonucleotides, respectively, for their ability to arrest translation of human DHFR in a rabbit reticulocyte assay. Each of the three 14 nucleobase antisense oligonucleotides alone was able to inhibit translation, albeit at a more modest level than the 28 or 42 nucleobase antisense oligonucleotides.
Antisense Compound Motifs
[0275] In certain embodiments, antisense compounds targeted to a PKK nucleic acid have chemically modified subunits arranged in patterns, or motifs, to confer to the antisense compounds properties such as enhanced inhibitory activity, increased binding affinity for a target nucleic acid, or resistance to degradation by in vivo nucleases.
[0276] Chimeric antisense compounds typically contain at least one region modified so as to confer increased resistance to nuclease degradation, increased cellular uptake, increased binding affinity for the target nucleic acid, and/or increased inhibitory activity. A second region of a chimeric antisense compound may optionally serve as a substrate for the cellular endonuclease RNase H, which cleaves the RNA strand of an RNA:DNA duplex.
[0277] Antisense compounds having a gapmer motif are considered chimeric antisense compounds. In a gapmer an internal region having a plurality of nucleotides that supports RNaseH cleavage is positioned between external regions having a plurality of nucleotides that are chemically distinct from the nucleosides of the internal region. In the case of an antisense oligonucleotide having a gapmer motif, the gap segment generally serves as the substrate for endonuclease cleavage, while the wing segments comprise modified nucleosides. In certain embodiments, the regions of a gapmer are differentiated by the types of sugar moieties comprising each distinct region. The types of sugar moieties that are used to differentiate the regions of a gapmer may in some embodiments include .beta.-D-ribonucleosides, .beta.-D-deoxyribonucleosides, 2'-modified nucleosides (such 2'-modified nucleosides may include 2'-MOE, and 2'-O--CH.sub.3, among others), and bicyclic sugar modified nucleosides (such bicyclic sugar modified nucleosides may include those having a 4'-(CH.sub.2).sub.n--O-2' bridge, where n=1 or n=2 and 4'-CH.sub.2--O--CH.sub.2-2'). In certain embodiments, wings may include several modified sugar moieties, including, for example 2'-MOE. In certain embodiments, wings may include several modified and unmodified sugar moieties. In certain embodiments, wings may include various combinations of 2'-MOE nucleosides and 2'-deoxynucleosides.
[0278] Each distinct region may comprise uniform sugar moieties, variant, or alternating sugar moieties. The wing-gap-wing motif is frequently described as "X--Y--Z", where "X" represents the length of the 5' wing, "Y" represents the length of the gap, and "Z" represents the length of the 3' wing. "X" and "Z" may comprise uniform, variant, or alternating sugar moieties. In certain embodiments, "X" and "Y" may include one or more 2'-deoxynucleosides. "Y" may comprise 2'-deoxynucleosides. As used herein, a gapmer described as "X--Y--Z" has a configuration such that the gap is positioned immediately adjacent to each of the 5' wing and the 3' wing. Thus, no intervening nucleotides exist between the 5' wing and gap, or the gap and the 3' wing. Any of the antisense compounds described herein can have a gapmer motif. In certain embodiments, "X" and "Z" are the same; in other embodiments they are different.
[0279] In certain embodiments, gapmers provided herein include, for example 20-mers having a motif of 5-10-5.
Target Nucleic Acids, Target Regions and Nucleotide Sequences
[0280] Nucleotide sequences that encode human plasma prekallikrein (PKK) include, without limitation, the following: GENBANK Accession No. NM_000892.3 (incorporated herein as SEQ ID NO: 1), GENBANK Accession No. DC412984.1 (incorporated herein as SEQ ID NO: 2), GENBANK Accession No. CN265612.1 (incorporated herein as SEQ ID NO: 3), GENBANK Accession No. AK297672.1 (incorporated herein as SEQ ID NO: 4), GENBANK Accession No. DC413312.1 (incorporated herein as SEQ ID NO: 5), GENBANK Accession No. AV688858.2 (incorporated herein as SEQ ID NO: 6), GENBANK Accession No. CD652077.1 (incorporated herein as SEQ ID NO: 7), GENBANK Accession No. BC143911.1 (incorporated herein as SEQ ID NO: 8), GENBANK Accession No. CB162532.1 (incorporated herein as SEQ ID NO: 9), GENBANK Accession No. NT 016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000 (incorporated herein as SEQ ID NO: 10), GENBANK Accession No. NM_008455.2 (incorporated herein as SEQ ID NO: 11), GENBANK Accession No. BB598673.1 (incorporated herein as SEQ ID NO: 12), GENBANK Accession No. NT_039460.7 truncated from nucleobases 6114001 to U.S. Pat. No. 6,144,000 (incorporated herein as SEQ ID NO: 13), GENBANK Accession No. NM_012725.2 (incorporated herein as SEQ ID NO: 14), GENBANK Accession No. NW_047473.1 truncated from nucleobases 10952001 to Ser. No. 10/982,000 (incorporated herein as SEQ ID NO: 15), GENBANK Accession No. XM_002804276.1 (incorporated herein as SEQ ID NO: 17), and GENBANK Accession No. NW_001118167.1 truncated from nucleobases 2358000 to 2391000 (incorporated herein as SEQ ID NO: 18).
[0281] It is understood that the sequence set forth in each SEQ ID NO in the Examples contained herein is independent of any modification to a sugar moiety, an internucleoside linkage, or a nucleobase. As such, antisense compounds defined by a SEQ ID NO may comprise, independently, one or more modifications to a sugar moiety, an internucleoside linkage, or a nucleobase. Antisense compounds described by Isis Number (Isis No) indicate a combination of nucleobase sequence and motif.
[0282] In certain embodiments, a target region is a structurally defined region of the target nucleic acid. For example, a target region may encompass a 3' UTR, a 5' UTR, an exon, an intron, an exon/intron junction, a coding region, a translation initiation region, translation termination region, or other defined nucleic acid region. The structurally defined regions for PKK can be obtained by accession number from sequence databases such as NCBI and such information is incorporated herein by reference. In certain embodiments, a target region may encompass the sequence from a 5' target site of one target segment within the target region to a 3' target site of another target segment within the same target region.
[0283] Targeting includes determination of at least one target segment to which an antisense compound hybridizes, such that a desired effect occurs. In certain embodiments, the desired effect is a reduction in mRNA target nucleic acid levels. In certain embodiments, the desired effect is reduction of levels of protein encoded by the target nucleic acid or a phenotypic change associated with the target nucleic acid.
[0284] A target region may contain one or more target segments. Multiple target segments within a target region may be overlapping. Alternatively, they may be non-overlapping. In certain embodiments, target segments within a target region are separated by no more than about 300 nucleotides. In certain embodiments, target segments within a target region are separated by a number of nucleotides that is, is about, is no more than, is no more than about, 250, 200, 150, 100, 90, 80, 70, 60, 50, 40, 30, 20, or 10 nucleotides on the target nucleic acid, or is a range defined by any two of the preceeding values. In certain embodiments, target segments within a target region are separated by no more than, or no more than about, 5 nucleotides on the target nucleic acid. In certain embodiments, target segments are contiguous. Contemplated are target regions defined by a range having a starting nucleic acid that is any of the 5' target sites or 3' target sites listed herein.
[0285] Suitable target segments may be found within a 5' UTR, a coding region, a 3' UTR, an intron, an exon, or an exon/intron junction. Target segments containing a start codon or a stop codon are also suitable target segments. A suitable target segment may specifically exclude a certain structurally defined region such as the start codon or stop codon.
[0286] The determination of suitable target segments may include a comparison of the sequence of a target nucleic acid to other sequences throughout the genome. For example, the BLAST algorithm may be used to identify regions of similarity amongst different nucleic acids. This comparison can prevent the selection of antisense compound sequences that may hybridize in a non-specific manner to sequences other than a selected target nucleic acid (i.e., non-target or off-target sequences).
[0287] There may be variation in activity (e.g., as defined by percent reduction of target nucleic acid levels) of the antisense compounds within an active target region. In certain embodiments, reductions in PKK mRNA levels are indicative of inhibition of PKK expression. Reductions in levels of a PKK protein are also indicative of inhibition of target mRNA expression. Further, phenotypic changes are indicative of inhibition of PKK expression. For example, reduced or prevented inflammation can be indicative of inhibition of PKK expression. In another example, reduced or prevented edema/swelling can be indicative of inhibition of PKK expression. In another example, reduced or prevented vascular permeability can be indicative of inhibition of PKK expression. In another example, reduced or prevented vascular leakage can be indicative of inhibition of PKK expression. In certain embodiments, vascular permeability is measured by quantification of a dye, such as Evans Blue.
Hybridization
[0288] In some embodiments, hybridization occurs between an antisense compound disclosed herein and a target nucleic acid. The most common mechanism of hybridization involves hydrogen bonding (e.g., Watson-Crick, Hoogsteen or reversed Hoogsteen hydrogen bonding) between complementary nucleobases of the nucleic acid molecules.
[0289] Hybridization can occur under varying conditions. Stringent conditions are sequence-dependent and are determined by the nature and composition of the nucleic acid molecules to be hybridized.
[0290] Methods of determining whether a sequence is specifically hybridizable to a target nucleic acid are well known in the art. In certain embodiments, the antisense compounds provided herein are specifically hybridizable with a target nucleic acid.
Complementarity
[0291] An antisense compound and a target nucleic acid are complementary to each other when a sufficient number of nucleobases of the antisense compound can hydrogen bond with the corresponding nucleobases of the target nucleic acid, such that a desired effect will occur (e.g., antisense inhibition of a target nucleic acid, such as a PKK nucleic acid).
[0292] Non-complementary nucleobases between an antisense compound and a PKK nucleic acid may be tolerated provided that the antisense compound remains able to specifically hybridize to a target nucleic acid. Moreover, an antisense compound may hybridize over one or more segments of a PKK nucleic acid such that intervening or adjacent segments are not involved in the hybridization event (e.g., a loop structure, mismatch or hairpin structure).
[0293] In certain embodiments, the antisense compounds provided herein, or a specified portion thereof, are, or are at least, 70%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% complementary to an PKK nucleic acid, a target region, target segment, or specified portion thereof. Percent complementarity of an antisense compound with a target nucleic acid can be determined using routine methods.
[0294] For example, an antisense compound in which 18 of 20 nucleobases of the antisense compound are complementary to a target region, and would therefore specifically hybridize, would represent 90 percent complementarity. In this example, the remaining noncomplementary nucleobases may be clustered or interspersed with complementary nucleobases and need not be contiguous to each other or to complementary nucleobases. As such, an antisense compound which is 18 nucleobases in length having four noncomplementary nucleobases which are flanked by two regions of complete complementarity with the target nucleic acid would have 77.8% overall complementarity with the target nucleic acid and would thus fall within the scope of the present invention. Percent complementarity of an antisense compound with a region of a target nucleic acid can be determined routinely using BLAST programs (basic local alignment search tools) and PowerBLAST programs known in the art (Altschul et al., J. Mol. Biol., 1990, 215, 403 410; Zhang and Madden, Genome Res., 1997, 7, 649 656). Percent homology, sequence identity or complementarity, can be determined by, for example, the Gap program (Wisconsin Sequence Analysis Package, Version 8 for Unix, Genetics Computer Group, University Research Park, Madison Wis.), using default settings, which uses the algorithm of Smith and Waterman (Adv. Appl. Math., 1981, 2, 482 489).
[0295] In certain embodiments, the antisense compounds provided herein, or specified portions thereof, are fully complementary (i.e. 100% complementary) to a target nucleic acid, or specified portion thereof. For example, an antisense compound may be fully complementary to a plasma prekallikrein nucleic acid, or a target region, or a target segment or target sequence thereof. As used herein, "fully complementary" means each nucleobase of an antisense compound is capable of precise base pairing with the corresponding nucleobases of a target nucleic acid. For example, a 20 nucleobase antisense compound is fully complementary to a target sequence that is 400 nucleobases long, so long as there is a corresponding 20 nucleobase portion of the target nucleic acid that is fully complementary to the antisense compound. Fully complementary can also be used in reference to a specified portion of the first and/or the second nucleic acid. For example, a 20 nucleobase portion of a 30 nucleobase antisense compound can be "fully complementary" to a target sequence that is 400 nucleobases long. The 20 nucleobase portion of the 30 nucleobase oligonucleotide is fully complementary to the target sequence if the target sequence has a corresponding 20 nucleobase portion wherein each nucleobase is complementary to the 20 nucleobase portion of the antisense compound. At the same time, the entire 30 nucleobase antisense compound may or may not be fully complementary to the target sequence, depending on whether the remaining 10 nucleobases of the antisense compound are also complementary to the target sequence.
[0296] The location of a non-complementary nucleobase may be at the 5' end or 3' end of the antisense compound. Alternatively, the non-complementary nucleobase or nucleobases may be at an internal position of the antisense compound. When two or more non-complementary nucleobases are present, they may be contiguous (i.e. linked) or non-contiguous. In one embodiment, a non-complementary nucleobase is located in the wing segment of a gapmer antisense oligonucleotide.
[0297] In certain embodiments, antisense compounds that are, or are up to 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 nucleobases in length comprise no more than 4, no more than 3, no more than 2, or no more than 1 non-complementary nucleobase(s) relative to a target nucleic acid or specified portion thereof.
[0298] In certain embodiments, antisense compounds that are, or are up to 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 nucleobases in length comprise no more than 6, no more than 5, no more than 4, no more than 3, no more than 2, or no more than 1 non-complementary nucleobase(s) relative to a target nucleic acid or specified portion thereof.
[0299] The antisense compounds provided also include those which are complementary to a portion of a target nucleic acid. As used herein, "portion" refers to a defined number of contiguous (i.e. linked) nucleobases within a region or segment of a target nucleic acid. A "portion" can also refer to a defined number of contiguous nucleobases of an antisense compound. In certain embodiments, the antisense compounds, are complementary to at least an 8 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least a 9 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least a 10 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least an 11 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least a 12 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least a 13 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least a 14 nucleobase portion of a target segment. In certain embodiments, the antisense compounds are complementary to at least a 15 nucleobase portion of a target segment. Also contemplated are antisense compounds that are complementary to at least a 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, or more nucleobase portion of a target segment, or a range defined by any two of these values.
Identity
[0300] The antisense compounds provided herein may also have a defined percent identity to a particular nucleotide sequence, SEQ ID NO, or compound represented by a specific Isis number, or portion thereof. As used herein, an antisense compound is identical to the sequence disclosed herein if it has the same nucleobase pairing ability. For example, a RNA which contains uracil in place of thymidine in a disclosed DNA sequence would be considered identical to the DNA sequence since both uracil and thymidine pair with adenine. Shortened and lengthened versions of the antisense compounds described herein as well as compounds having non-identical bases relative to the antisense compounds provided herein also are contemplated. The non-identical bases may be adjacent to each other or dispersed throughout the antisense compound. Percent identity of an antisense compound is calculated according to the number of bases that have identical base pairing relative to the sequence to which it is being compared.
[0301] In certain embodiments, the antisense compounds, or portions thereof, are at least 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99% or 100% identical to one or more of the antisense compounds or SEQ ID NOs, or a portion thereof, disclosed herein.
[0302] In certain embodiments, a portion of the antisense compound is compared to an equal length portion of the target nucleic acid. In certain embodiments, an 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, or 25 nucleobase portion is compared to an equal length portion of the target nucleic acid.
[0303] In certain embodiments, a portion of the antisense oligonucleotide is compared to an equal length portion of the target nucleic acid. In certain embodiments, an 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, or 25 nucleobase portion is compared to an equal length portion of the target nucleic acid.
Modifications
[0304] A nucleoside is a base-sugar combination. The nucleobase (also known as base) portion of the nucleoside is normally a heterocyclic base moiety. Nucleotides are nucleosides that further include a phosphate group covalently linked to the sugar portion of the nucleoside. For those nucleosides that include a pentofuranosyl sugar, the phosphate group can be linked to the 2', 3' or 5' hydroxyl moiety of the sugar. Oligonucleotides are formed through the covalent linkage of adjacent nucleosides to one another, to form a linear polymeric oligonucleotide. Within the oligonucleotide structure, the phosphate groups are commonly referred to as forming the internucleoside linkages of the oligonucleotide.
[0305] Modifications to antisense compounds encompass substitutions or changes to internucleoside linkages, sugar moieties, or nucleobases. Modified antisense compounds are often preferred over native forms because of desirable properties such as, for example, enhanced cellular uptake, enhanced affinity for nucleic acid target, increased stability in the presence of nucleases, or increased inhibitory activity.
[0306] Chemically modified nucleosides may also be employed to increase the binding affinity of a shortened or truncated antisense oligonucleotide for its target nucleic acid. Consequently, comparable results can often be obtained with shorter antisense compounds that have such chemically modified nucleosides.
Modified Internucleoside Linkages
[0307] The naturally occurring internucleoside linkage of RNA and DNA is a 3' to 5' phosphodiester linkage. Antisense compounds having one or more modified, i.e. non-naturally occurring, internucleoside linkages are often selected over antisense compounds having naturally occurring internucleoside linkages because of desirable properties such as, for example, enhanced cellular uptake, enhanced affinity for target nucleic acids, and increased stability in the presence of nucleases.
[0308] Oligonucleotides having modified internucleoside linkages include internucleoside linkages that retain a phosphorus atom as well as internucleoside linkages that do not have a phosphorus atom. Representative phosphorus containing internucleoside linkages include, but are not limited to, phosphodiesters, phosphotriesters, methylphosphonates, phosphoramidate, and phosphorothioates. Methods of preparation of phosphorous-containing and non-phosphorous-containing linkages are well known.
[0309] In certain embodiments, antisense compounds targeted to a plasma prekallikrein nucleic acid comprise one or more modified internucleoside linkages. In certain embodiments, the modified internucleoside linkages are phosphorothioate linkages. In certain embodiments, each internucleoside linkage of an antisense compound is a phosphorothioate internucleoside linkage.
[0310] In certain embodiments, oligonucleotides comprise modified internucleoside linkages arranged along the oligonucleotide or region thereof in a defined pattern or modified internucleoside linkage motif. In certain embodiments, internucleoside linkages are arranged in a gapped motif. In such embodiments, the internucleoside linkages in each of two wing regions are different from the internucleoside linkages in the gap region. In certain embodiments the internucleoside linkages in the wings are phosphodiester and the internucleoside linkages in the gap are phosphorothioate. The nucleoside motif is independently selected, so such oligonucleotides having a gapped internucleoside linkage motif may or may not have a gapped nucleoside motif and if it does have a gapped nucleoside motif, the wing and gap lengths may or may not be the same.
[0311] In certain embodiments, oligonucleotides comprise a region having an alternating internucleoside linkage motif. In certain embodiments, oligonucleotides of the present invention comprise a region of uniformly modified internucleoside linkages. In certain such embodiments, the oligonucleotide comprises a region that is uniformly linked by phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide is uniformly linked by phosphorothioate. In certain embodiments, each internucleoside linkage of the oligonucleotide is selected from phosphodiester and phosphorothioate. In certain embodiments, each internucleoside linkage of the oligonucleotide is selected from phosphodiester and phosphorothioate and at least one internucleoside linkage is phosphorothioate.
[0312] In certain embodiments, the oligonucleotide comprises at least 6 phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide comprises at least 8 phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide comprises at least 10 phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide comprises at least one block of at least 6 consecutive phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide comprises at least one block of at least 8 consecutive phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide comprises at least one block of at least 10 consecutive phosphorothioate internucleoside linkages. In certain embodiments, the oligonucleotide comprises at least block of at least one 12 consecutive phosphorothioate internucleoside linkages. In certain such embodiments, at least one such block is located at the 3 end of the oligonucleotide. In certain such embodiments, at least one such block is located within 3 nucleosides of the 3' end of the oligonucleotide.
[0313] In certain embodiments, oligonucleotides comprise one or more methylphosponate linkages. In certain embodiments, oligonucleotides having a gapmer nucleoside motif comprise a linkage motif comprising all phosphorothioate linkages except for one or two methylphosponate linkages. In certain embodiments, one methylphosponate linkage is in the central gap of an oligonucleotide having a gapmer nucleoside motif.
[0314] In certain embodiments, it is desirable to arrange the number of phosphorothioate internucleoside linkages and phosphodiester internucleoside linkages to maintain nuclease resistance. In certain embodiments, it is desirable to arrange the number and position of phosphorothioate internucleoside linkages and the number and position of phosphodiester internucleoside linkages to maintain nuclease resistance. In certain embodiments, the number of phosphorothioate internucleoside linkages may be decreased and the number of phosphodiester internucleoside linkages may be increased. In certain embodiments, the number of phosphorothioate internucleoside linkages may be decreased and the number of phosphodiester internucleoside linkages may be increased while still maintaining nuclease resistance. In certain embodiments it is desirable to decrease the number of phosphorothioate internucleoside linkages while retaining nuclease resistance. In certain embodiments it is desirable to increase the number of phosphodiester internucleoside linkages while retaining nuclease resistance.
Modified Sugar Moieties
[0315] Antisense compounds can optionally contain one or more nucleosides wherein the sugar group has been modified. Such sugar modified nucleosides may impart enhanced nuclease stability, increased binding affinity, or some other beneficial biological property to the antisense compounds. In certain embodiments, nucleosides comprise chemically modified ribofuranose ring moieties. Examples of chemically modified ribofuranose rings include without limitation, addition of substitutent groups (including 5' and 2' substituent groups, bridging of non-geminal ring atoms to form bicyclic nucleic acids (BNA), replacement of the ribosyl ring oxygen atom with S, N(R), or C(R.sub.1)(R.sub.2) (R, R.sub.1 and R.sub.2 are each independently H, C.sub.1-C.sub.12 alkyl or a protecting group) and combinations thereof. Examples of chemically modified sugars include 2'-F-5'-methyl substituted nucleoside (see PCT International Application WO 2008/101157 Published on Aug. 21, 2008 for other disclosed 5',2'-bis substituted nucleosides) or replacement of the ribosyl ring oxygen atom with S with further substitution at the 2'-position (see published U.S. Patent Application US2005-0130923, published on Jun. 16, 2005) or alternatively 5'-substitution of a BNA (see PCT International Application WO 2007/134181 Published on Nov. 22, 2007 wherein LNA is substituted with for example a 5'-methyl or a 5'-vinyl group).
[0316] Examples of nucleosides having modified sugar moieties include without limitation nucleosides comprising 5'-vinyl, 5'-methyl (R or S), 4'-S, 2'-F, 2'-OCH.sub.3, 2'-OCH.sub.2CH.sub.3, 2'-OCH.sub.2CH.sub.2F and 2'-O(CH.sub.2).sub.2OCH.sub.3 substituent groups. The substituent at the 2' position can also be selected from allyl, amino, azido, thio, O-allyl, O--C.sub.1-C.sub.10 alkyl, OCF.sub.3, OCH.sub.2F, O(CH.sub.2).sub.2SCH.sub.3, O(CH.sub.2).sub.2--O--N(R.sub.m)(R.sub.n), O--CH.sub.2--C(.dbd.O)--N(R.sub.m)(R.sub.n), and O--CH.sub.2--C(.dbd.O)--N(R)--(CH.sub.2).sub.2--N(R.sub.m)(R.sub.n), where each R.sub.l, R.sub.m and R.sub.n is, independently, H or substituted or unsubstituted C.sub.1-C.sub.10 alkyl.
[0317] As used herein, "bicyclic nucleosides" refer to modified nucleosides comprising a bicyclic sugar moiety. Examples of bicyclic nucleosides include without limitation nucleosides comprising a bridge between the 4' and the 2' ribosyl ring atoms. In certain embodiments, antisense compounds provided herein include one or more bicyclic nucleosides comprising a 4' to 2' bridge. Examples of such 4' to 2' bridged bicyclic nucleosides, include but are not limited to one of the formulae: 4'-(CH.sub.2)--O-2' (LNA); 4'-(CH.sub.2)--S-2'; 4'-(CH.sub.2).sub.2--O-2' (ENA); 4'-CH(CH.sub.3)--O-2' (also referred to as constrained ethyl or cEt) and 4'-CH(CH.sub.2OCH.sub.3)--O-2' (and analogs thereof see U.S. Pat. No. 7,399,845, issued on Jul. 15, 2008); 4'-C(CH.sub.3)(CH.sub.3)--O-2' (and analogs thereof see published International Application WO/2009/006478, published Jan. 8, 2009); 4'-CH.sub.2--N(OCH.sub.3)-2' (and analogs thereof see published International Application WO/2008/150729, published Dec. 11, 2008); 4'-CH.sub.2--O--N(CH.sub.3)-2' (see published U.S. Patent Application US2004-0171570, published Sep. 2, 2004); 4'-CH.sub.2--N(R)--O-2', wherein R is H, C.sub.1-C.sub.12 alkyl, or a protecting group (see U.S. Pat. No. 7,427,672, issued on Sep. 23, 2008); 4'-CH.sub.2--C(H)(CH.sub.3)-2' (see Zhou et al., J. Org. Chem., 2009, 74, 118-134); and 4'-CH.sub.2--C(.dbd.CH.sub.2)-2' (and analogs thereof see published International Application WO 2008/154401, published on Dec. 8, 2008).
[0318] Further reports related to bicyclic nucleosides can also be found in published literature (see for example: Singh et al., Chem. Commun., 1998, 4, 455-456; Koshkin et al., Tetrahedron, 1998, 54, 3607-3630; Wahlestedt et al., Proc. Natl. Acad. Sci. U.S.A, 2000, 97, 5633-5638; Kumar et al., Bioorg. Med. Chem. Lett., 1998, 8, 2219-2222; Singh et al., J. Org. Chem., 1998, 63, 10035-10039; Srivastava et al., J. Am. Chem. Soc., 2007, 129(26) 8362-8379; Elayadi et al., Curr. Opinion Invest. Drugs, 2001, 2, 558-561; Braasch et al., Chem. Biol., 2001, 8, 1-7; and Orum et al., Curr. Opinion Mol. Ther., 2001, 3, 239-243; U.S. Pat. Nos. 6,268,490; 6,525,191; 6,670,461; 6,770,748; 6,794,499; 7,034,133; 7,053,207; 7,399,845; 7,547,684; and 7,696,345; U.S. Patent Publication No. US2008-0039618; US2009-0012281; U.S. Patent Ser. No. 61/026,995 and 61/097,787; Published PCT International applications WO 1999/014226; WO 2004/106356; WO 2005/021570; WO 2007/134181; WO 2008/150729; WO 2008/154401; WO 2009/006478; WO 2010/036698; WO 2011/017521; WO 2009/067647; WO 2009/100320. Each of the foregoing bicyclic nucleosides can be prepared having one or more stereochemical sugar configurations including for example .alpha.-L-ribofuranose and .beta.-D-ribofuranose (see PCT international application PCT/DK98/00393, published on Mar. 25, 1999 as WO 99/14226).
[0319] In certain embodiments, bicyclic sugar moieties of BNA nucleosides include, but are not limited to, compounds having at least one bridge between the 4' and the 2' position of the pentofuranosyl sugar moiety wherein such bridges independently comprises 1 or from 2 to 4 linked groups independently selected from --[C(R.sub.a)(R.sub.b)].sub.n--, --C(R.sub.a).dbd.C(R.sub.b)--, --C(R.sub.a).dbd.N--, --C(.dbd.O)--, --C(.dbd.NR.sub.a)--, --C(.dbd.S)--, --O--, --Si(R.sub.a).sub.2--, --S(.dbd.O).sub.x--, and --N(R.sub.a)--;
[0320] wherein:
[0321] x is 0, 1, or 2;
[0322] n is 1, 2, 3, or 4;
[0323] each R.sub.a and R.sub.b is, independently, H, a protecting group, hydroxyl, C.sub.1-C.sub.12 alkyl, substituted C.sub.1-C.sub.12 alkyl, C.sub.2-C.sub.12 alkenyl, substituted C.sub.2-C.sub.12 alkenyl, C.sub.2-C.sub.12 alkynyl, substituted C.sub.2-C.sub.12 alkynyl, C.sub.5-C.sub.20 aryl, substituted C.sub.5-C.sub.20 aryl, heterocycle radical, substituted heterocycle radical, heteroaryl, substituted heteroaryl, C.sub.5-C.sub.7 alicyclic radical, substituted C.sub.5-C.sub.7 alicyclic radical, halogen, OJ.sub.1, NJ.sub.1J.sub.2, SJ.sub.1, N.sub.3, COOJ.sub.1, acyl (C(.dbd.O)--H), substituted acyl, CN, sulfonyl (S(.dbd.O).sub.2-J.sub.1), or sulfoxyl (S(.dbd.O)-J.sub.1); and each J.sub.1 and J.sub.2 is, independently, H, C.sub.1-C.sub.12 alkyl, substituted C.sub.1-C.sub.12 alkyl, C.sub.2-C.sub.12 alkenyl, substituted C.sub.2-C.sub.12 alkenyl, C.sub.2-C.sub.12 alkynyl, substituted C.sub.2-C.sub.12 alkynyl, C.sub.5-C.sub.20 aryl, substituted C.sub.5-C.sub.20 aryl, acyl (C(.dbd.O)--H), substituted acyl, a heterocycle radical, a substituted heterocycle radical, C.sub.1-C.sub.12 aminoalkyl, substituted C.sub.1-C.sub.12 aminoalkyl or a protecting group.
[0324] In certain embodiments, the bridge of a bicyclic sugar moiety is --[C(R.sub.a)(R.sub.b)].sub.n--, --[C(R.sub.a)(R.sub.b)].sub.n--O--, --C(R.sub.aR.sub.b)--N(R)--O-- or --C(R.sub.aR.sub.b)--O--N(R)--. In certain embodiments, the bridge is 4'-CH.sub.2-2',4'-(CH.sub.2).sub.2-2',4'-(CH.sub.2).sub.3-2',4'-CH.sub.2-- -O-2',4'-(CH.sub.2).sub.2--O-2',4'-CH.sub.2--O--N(R)-2' and 4'-CH.sub.2--N(R)--O-2'- wherein each R is, independently, H, a protecting group or C.sub.1-C.sub.12 alkyl.
[0325] In certain embodiments, bicyclic nucleosides are further defined by isomeric configuration. For example, a nucleoside comprising a 4'-2' methylene-oxy bridge, may be in the .alpha.-L configuration or in the .beta.-D configuration. Previously, .alpha.-L-methyleneoxy (4'-CH.sub.2--O-2') BNA's have been incorporated into antisense oligonucleotides that showed antisense activity (Frieden et al., Nucleic Acids Research, 2003, 21, 6365-6372).
[0326] In certain embodiments, bicyclic nucleosides include, but are not limited to, (A) .alpha.-L-methyleneoxy (4'-CH.sub.2--O-2') BNA, (B) .beta.-D-methyleneoxy (4'-CH.sub.2--O-2') BNA, (C) ethyleneoxy (4'-(CH.sub.2).sub.2--O-2') BNA, (D) aminooxy (4'-CH.sub.2--O--N(R)-2') BNA, (E) oxyamino (4'-CH.sub.2--N(R)--O-2') BNA, and (F) methyl(methyleneoxy) (4'-CH(CH.sub.3)--O-2') BNA, (G) methylene-thio (4'-CH.sub.2--S-2') BNA, (H) methylene-amino (4'-CH.sub.2--N(R)-2') BNA, (I) methyl carbocyclic (4'-CH.sub.2--CH(CH.sub.3)-2') BNA, (J) propylene carbocyclic (4'-(CH.sub.2).sub.3-2') BNA and (K) vinyl BNA as depicted below:
##STR00011## ##STR00012##
[0327] wherein Bx is the base moiety and R is independently H, a protecting group, C.sub.1-C.sub.12 alkyl or C.sub.1-C.sub.12 alkoxy.
[0328] In certain embodiments, bicyclic nucleosides are provided having Formula I:
##STR00013##
wherein:
[0329] Bx is a heterocyclic base moiety;
[0330] -Q.sub.a-Q.sub.b-Q.sub.c- is --CH.sub.2--N(R.sub.c)--CH.sub.2--, --C(.dbd.O)--N(R.sub.c)--CH.sub.2--, --CH.sub.2--O--N(R.sub.c)--, --CH.sub.2--N(R.sub.c)--O-- or --N(R.sub.c)--O--CH.sub.2;
[0331] R.sub.c is C.sub.1-C.sub.12 alkyl or an amino protecting group; and
[0332] T.sub.a and T.sub.b are each, independently H, a hydroxyl protecting group, a conjugate group, a reactive phosphorus group, a phosphorus moiety or a covalent attachment to a support medium.
[0333] In certain embodiments, bicyclic nucleosides are provided having Formula II:
##STR00014##
wherein:
[0334] Bx is a heterocyclic base moiety;
[0335] T.sub.a and T.sub.b are each, independently H, a hydroxyl protecting group, a conjugate group, a reactive phosphorus group, a phosphorus moiety or a covalent attachment to a support medium;
[0336] Z.sub.a is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, substituted C.sub.1-C.sub.6 alkyl, substituted C.sub.2-C.sub.6 alkenyl, substituted C.sub.2-C.sub.6 alkynyl, acyl, substituted acyl, substituted amide, thiol or substituted thio.
[0337] In one embodiment, each of the substituted groups is, independently, mono or poly substituted with substituent groups independently selected from halogen, oxo, hydroxyl, OJ.sub.c, NJ.sub.cJ.sub.d, SJ.sub.c, N.sub.3, OC(.dbd.X)J.sub.c, and NJ.sub.eC(.dbd.X)NJ.sub.cJ.sub.d, wherein each J.sub.c, J.sub.d and J.sub.e is, independently, H, C.sub.1-C.sub.6 alkyl, or substituted C.sub.1-C.sub.6 alkyl and X is O or NJ.sub.c.
[0338] In certain embodiments, bicyclic nucleosides are provided having Formula III:
##STR00015##
wherein:
[0339] Bx is a heterocyclic base moiety;
[0340] T.sub.a and T.sub.b are each, independently H, a hydroxyl protecting group, a conjugate group, a reactive phosphorus group, a phosphorus moiety or a covalent attachment to a support medium;
[0341] Z.sub.b is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, substituted C.sub.1-C.sub.6 alkyl, substituted C.sub.2-C.sub.6 alkenyl, substituted C.sub.2-C.sub.6 alkynyl or substituted acyl (C(.dbd.O)--).
[0342] In certain embodiments, bicyclic nucleosides are provided having Formula IV:
##STR00016##
wherein:
[0343] Bx is a heterocyclic base moiety;
[0344] T.sub.a and T.sub.b are each, independently H, a hydroxyl protecting group, a conjugate group, a reactive phosphorus group, a phosphorus moiety or a covalent attachment to a support medium;
[0345] R.sub.d is C.sub.1-C.sub.6 alkyl, substituted C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, substituted C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl or substituted C.sub.2-C.sub.6 alkynyl;
[0346] each q.sub.a, q.sub.b, q.sub.c and q.sub.d is, independently, H, halogen, C.sub.1-C.sub.6 alkyl, substituted C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, substituted C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl or substituted C.sub.2-C.sub.6 alkynyl, C.sub.1-C.sub.6 alkoxyl, substituted C.sub.1-C.sub.6 alkoxyl, acyl, substituted acyl, C.sub.1-C.sub.6 aminoalkyl or substituted C.sub.1-C.sub.6 aminoalkyl;
[0347] In certain embodiments, bicyclic nucleosides are provided having Formula V:
##STR00017##
wherein:
[0348] Bx is a heterocyclic base moiety;
[0349] T.sub.a and T.sub.b are each, independently H, a hydroxyl protecting group, a conjugate group, a reactive phosphorus group, a phosphorus moiety or a covalent attachment to a support medium;
[0350] q.sub.a, q.sub.b, q.sub.e and q.sub.f are each, independently, hydrogen, halogen, C.sub.1-C.sub.12 alkyl, substituted C.sub.1-C.sub.12 alkyl, C.sub.2-C.sub.12 alkenyl, substituted C.sub.2-C.sub.12 alkenyl, C.sub.2-C.sub.12 alkynyl, substituted C.sub.2-C.sub.12 alkynyl, C.sub.1-C.sub.12 alkoxy, substituted C.sub.1-C.sub.12 alkoxy, OJ.sub.j, SJ.sub.j, SOJ.sub.j, SO.sub.2J.sub.j, N.sub.jj.sub.k, N.sub.3, CN, C(.dbd.O)OJ.sub.j, C(.dbd.O)N.sub.jj.sub.k, C(.dbd.O)J.sub.j, O--C(.dbd.O)N.sub.jj.sub.k, N(H)C(.dbd.NH)NJ.sub.jJ.sub.k, N(H)C(.dbd.O)NJ.sub.jJ.sub.k or N(H)C(.dbd.S)NJ.sub.jJ.sub.k;
[0351] or q.sub.e and q.sub.f together are .dbd.C(q.sub.g)(q.sub.h);
[0352] q.sub.g and q.sub.h are each, independently, H, halogen, C.sub.1-C.sub.12 alkyl or substituted C.sub.1-C.sub.12 alkyl.
[0353] The synthesis and preparation of the methyleneoxy (4'-CH.sub.2--O-2') BNA monomers adenine, cytosine, guanine, 5-methyl-cytosine, thymine and uracil, along with their oligomerization, and nucleic acid recognition properties have been described (Koshkin et al., Tetrahedron, 1998, 54, 3607-3630). BNAs and preparation thereof are also described in WO 98/39352 and WO 99/14226.
[0354] Analogs of methyleneoxy (4'-CH.sub.2--O-2') BNA and 2'-thio-BNAs, have also been prepared (Kumar et al., Bioorg. Med. Chem. Lett., 1998, 8, 2219-2222). Preparation of locked nucleoside analogs comprising oligodeoxyribonucleotide duplexes as substrates for nucleic acid polymerases has also been described (Wengel et al., WO 99/14226). Furthermore, synthesis of 2'-amino-BNA, a novel comformationally restricted high-affinity oligonucleotide analog has been described in the art (Singh et al., J Org. Chem., 1998, 63, 10035-10039). In addition, 2'-amino- and 2'-methylamino-BNA's have been prepared and the thermal stability of their duplexes with complementary RNA and DNA strands has been previously reported.
[0355] In certain embodiments, bicyclic nucleosides are provided having Formula VI:
##STR00018##
wherein:
[0356] Bx is a heterocyclic base moiety;
[0357] T.sub.a and T.sub.b are each, independently H, a hydroxyl protecting group, a conjugate group, a reactive phosphorus group, a phosphorus moiety or a covalent attachment to a support medium;
[0358] each q.sub.i, q.sub.j, q.sub.k and q.sub.l is, independently, H, halogen, C.sub.1-C.sub.12 alkyl, substituted C.sub.1-C.sub.12 alkyl, C.sub.2-C.sub.12 alkenyl, substituted C.sub.2-C.sub.12 alkenyl, C.sub.2-C.sub.12 alkynyl, substituted C.sub.2-C.sub.12 alkynyl, C.sub.1-C.sub.12 alkoxyl, substituted C.sub.1-C.sub.12 alkoxyl, OJ.sub.j, SJ.sub.j, SOJ.sub.j, SO.sub.2J.sub.j, NJ.sub.jJ.sub.k, N.sub.3, CN, C(.dbd.O)OJ.sub.j, C(.dbd.O)NJ.sub.jJ.sub.k, C(.dbd.O)J.sub.j, O--C(.dbd.O)NJ.sub.jJ.sub.k, N(H)C(.dbd.NH)NJ.sub.jJ.sub.k, N(H)C(.dbd.O)NJ.sub.jJ.sub.k or N(H)C(.dbd.S)NJ.sub.jJ.sub.k; and
[0359] q.sub.i and q.sub.j or q.sub.l and q.sub.k together are .dbd.C(q.sub.g)(q.sub.h), wherein q.sub.g and q.sub.h are each, independently, H, halogen, C.sub.1-C.sub.12 alkyl or substituted C.sub.1-C.sub.12 alkyl.
[0360] One carbocyclic bicyclic nucleoside having a 4'-(CH.sub.2).sub.3-2' bridge and the alkenyl analog bridge 4'-CH.dbd.CH--CH.sub.2-2' have been described (Freier et al., Nucleic Acids Research, 1997, 25(22), 4429-4443 and Albaek et al., J Org. Chem., 2006, 71, 7731-7740). The synthesis and preparation of carbocyclic bicyclic nucleosides along with their oligomerization and biochemical studies have also been described (Srivastava et al., J Am. Chem. Soc., 2007, 129(26), 8362-8379).
[0361] As used herein, "4'-2' bicyclic nucleoside" or "4' to 2' bicyclic nucleoside" refers to a bicyclic nucleoside comprising a furanose ring comprising a bridge connecting two carbon atoms of the furanose ring connects the 2' carbon atom and the 4' carbon atom of the sugar ring.
[0362] As used herein, "monocylic nucleosides" refer to nucleosides comprising modified sugar moieties that are not bicyclic sugar moieties. In certain embodiments, the sugar moiety, or sugar moiety analogue, of a nucleoside may be modified or substituted at any position.
[0363] As used herein, "2'-modified sugar" means a furanosyl sugar modified at the 2' position. In certain embodiments, such modifications include substituents selected from: a halide, including, but not limited to substituted and unsubstituted alkoxy, substituted and unsubstituted thioalkyl, substituted and unsubstituted amino alkyl, substituted and unsubstituted alkyl, substituted and unsubstituted allyl, and substituted and unsubstituted alkynyl. In certain embodiments, 2' modifications are selected from substituents including, but not limited to: O[(CH.sub.2).sub.nO].sub.mCH.sub.3, O(CH.sub.2).sub.nNH.sub.2, O(CH.sub.2).sub.nCH.sub.3, O(CH.sub.2).sub.nF, O(CH.sub.2).sub.nONH.sub.2, OCH.sub.2C(.dbd.O)N(H)CH.sub.3, and O(CH.sub.2).sub.nON[(CH.sub.2).sub.nCH.sub.3].sub.2, where n and m are from 1 to about 10. Other 2'-substituent groups can also be selected from: C.sub.1-C.sub.12 alkyl, substituted alkyl, alkenyl, alkynyl, alkaryl, aralkyl, O-alkaryl or O-aralkyl, SH, SCH.sub.3, OCN, Cl, Br, CN, F, CF.sub.3, OCF.sub.3, SOCH.sub.3, SO.sub.2CH.sub.3, ONO.sub.2, NO.sub.2, N.sub.3, NH.sub.2, heterocycloalkyl, heterocycloalkaryl, aminoalkylamino, polyalkylamino, substituted silyl, an RNA cleaving group, a reporter group, an intercalator, a group for improving pharmacokinetic properties, or a group for improving the pharmacodynamic properties of an antisense compound, and other substituents having similar properties. In certain embodiments, modified nucleosides comprise a 2'-MOE side chain (Baker et al., J Biol. Chem., 1997, 272, 11944-12000). Such 2'-MOE substitution have been described as having improved binding affinity compared to unmodified nucleosides and to other modified nucleosides, such as 2'-O-methyl, O-propyl, and O-aminopropyl. Oligonucleotides having the 2'-MOE substituent also have been shown to be antisense inhibitors of gene expression with promising features for in vivo use (Martin, Helv. Chim. Acta, 1995, 78, 486-504; Altmann et al., Chimia, 1996, 50, 168-176; Altmann et al., Biochem. Soc. Trans., 1996, 24, 630-637; and Altmann et al., Nucleosides Nucleotides, 1997, 16, 917-926). As used herein, a "modified tetrahydropyran nucleoside" or "modified THP nucleoside" means a nucleoside having a six-membered tetrahydropyran "sugar" substituted in for the pentofuranosyl residue in normal nucleosides (a sugar surrogate). Modified THP nucleosides include, but are not limited to, what is referred to in the art as hexitol nucleic acid (HNA), anitol nucleic acid (ANA), manitol nucleic acid (MNA) (see Leumann, Bioorg. Med. Chem., 2002, 10, 841-1954) or fluoro HNA (F-HNA) having a tetrahydropyran ring system as illustrated below:
##STR00019##
[0364] In certain embodiments, sugar surrogates are selected having Formula VII:
##STR00020##
wherein independently for each of said at least one tetrahydropyran nucleoside analog of Formula VII:
[0365] Bx is a heterocyclic base moiety;
[0366] T.sub.a and T.sub.b are each, independently, an internucleoside linking group linking the tetrahydropyran nucleoside analog to the antisense compound or one of T.sub.a and T.sub.b is an internucleoside linking group linking the tetrahydropyran nucleoside analog to the antisense compound and the other of T.sub.a and T.sub.b is H, a hydroxyl protecting group, a linked conjugate group or a 5' or 3'-terminal group;
[0367] q.sub.1, q.sub.2, q.sub.3, q.sub.4, q.sub.5, q.sub.6 and q.sub.7 are each independently, H, C.sub.1-C.sub.6 alkyl, substituted C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, substituted C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl or substituted C.sub.2-C.sub.6 alkynyl; and each of R.sub.1 and R.sub.2 is selected from hydrogen, hydroxyl, halogen, substituted or unsubstituted alkoxy, NJ.sub.1J.sub.2, SJ.sub.1, N.sub.3, OC(.dbd.X)J.sub.1, OC(.dbd.X)NJ.sub.1J.sub.2, NJ.sub.3C(.dbd.X)NJ.sub.1J.sub.2 and CN, wherein X is O, S or NJ.sub.1 and each J.sub.1, J.sub.2 and J.sub.3 is, independently, H or C.sub.1-C.sub.6 alkyl.
[0368] In certain embodiments, the modified THP nucleosides of Formula VII are provided wherein q.sub.1, q.sub.2, q.sub.3, q.sub.4, q.sub.5, q.sub.6 and q.sub.7 are each H. In certain embodiments, at least one of q.sub.1, q.sub.2, q.sub.3, q.sub.4, q.sub.5, q.sub.6 and q.sub.7 is other than H. In certain embodiments, at least one of q.sub.1, q.sub.2, q.sub.3, q.sub.4, q.sub.5, q.sub.6 and q.sub.7 is methyl. In certain embodiments, THP nucleosides of Formula VII are provided wherein one of R.sub.1 and R.sub.2 is fluoro. In certain embodiments, R.sub.1 is fluoro and R.sub.2 is H; R.sub.1 is methoxy and R.sub.2 is H, and R.sub.1 is methoxyethoxy and R.sub.2 is H.
[0369] In certain embodiments, sugar surrogates comprise rings having more than 5 atoms and more than one heteroatom. For example nucleosides comprising morpholino sugar moieties and their use in oligomeric compounds has been reported (see for example: Braasch et al., Biochemistry, 2002, 41, 4503-4510; and U.S. Pat. Nos. 5,698,685; 5,166,315; 5,185,444; and 5,034,506). As used here, the term "morpholino" means a sugar surrogate having the following formula:
##STR00021##
In certain embodiments, morpholinos may be modified, for example by adding or altering various substituent groups from the above morpholino structure. Such sugar surrogates are referred to herein as "modified morpholinos."
[0370] Combinations of modifications are also provided without limitation, such as 2'-F-5'-methyl substituted nucleosides (see PCT International Application WO 2008/101157 published on Aug. 21, 2008 for other disclosed 5',2'-bis substituted nucleosides) and replacement of the ribosyl ring oxygen atom with S and further substitution at the 2'-position (see published U.S. Patent Application US2005-0130923, published on Jun. 16, 2005) or alternatively 5'-substitution of a bicyclic nucleic acid (see PCT International Application WO 2007/134181, published on Nov. 22, 2007 wherein a 4'-CH.sub.2--O-2' bicyclic nucleoside is further substituted at the 5' position with a 5'-methyl or a 5'-vinyl group). The synthesis and preparation of carbocyclic bicyclic nucleosides along with their oligomerization and biochemical studies have also been described (see, e.g., Srivastava et al., J. Am. Chem. Soc. 2007, 129(26), 8362-8379).
[0371] In certain embodiments, antisense compounds comprise one or more modified cyclohexenyl nucleosides, which is a nucleoside having a six-membered cyclohexenyl in place of the pentofuranosyl residue in naturally occurring nucleosides. Modified cyclohexenyl nucleosides include, but are not limited to those described in the art (see for example commonly owned, published PCT Application WO 2010/036696, published on Apr. 10, 2010, Robeyns et al., J. Am. Chem. Soc., 2008, 130(6), 1979-1984; Horvath et al., Tetrahedron Letters, 2007, 48, 3621-3623; Nauwelaerts et al., J Am. Chem. Soc., 2007, 129(30), 9340-9348; Gu et al., Nucleosides, Nucleotides & Nucleic Acids, 2005, 24(5-7), 993-998; Nauwelaerts et al., Nucleic Acids Research, 2005, 33(8), 2452-2463; Robeyns et al., Acta Crystallographica, Section F: Structural Biology and Crystallization Communications, 2005, F61(6), 585-586; Gu et al., Tetrahedron, 2004, 60(9), 2111-2123; Gu et al., Oligonucleotides, 2003, 13(6), 479-489; Wang et al., J. Org. Chem., 2003, 68, 4499-4505; Verbeure et al., Nucleic Acids Research, 2001, 29(24), 4941-4947; Wang et al., J. Org. Chem., 2001, 66, 8478-82; Wang et al., Nucleosides, Nucleotides & Nucleic Acids, 2001, 20(4-7), 785-788; Wang et al., J. Am. Chem., 2000, 122, 8595-8602; Published PCT application, WO 06/047842; and Published PCT Application WO 01/049687; the text of each is incorporated by reference herein, in their entirety). Certain modified cyclohexenyl nucleosides have Formula X.
##STR00022##
[0372] wherein independently for each of said at least one cyclohexenyl nucleoside analog of Formula X:
[0373] Bx is a heterocyclic base moiety;
[0374] T.sub.3 and T.sub.4 are each, independently, an internucleoside linking group linking the cyclohexenyl nucleoside analog to an antisense compound or one of T.sub.3 and T.sub.4 is an internucleoside linking group linking the tetrahydropyran nucleoside analog to an antisense compound and the other of T.sub.3 and T.sub.4 is H, a hydroxyl protecting group, a linked conjugate group, or a 5'- or 3'-terminal group; and
[0375] q.sub.1, q.sub.2, q.sub.3, q.sub.4, q.sub.5, q.sub.6, q.sub.7, q.sub.8 and q.sub.9 are each, independently, H, C.sub.1-C.sub.6 alkyl, substituted C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, substituted C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, substituted C.sub.2-C.sub.6 alkynyl or other sugar substituent group.
[0376] As used herein, "2'-modified" or "2'-substituted" refers to a nucleoside comprising a sugar comprising a substituent at the 2' position other than H or OH. 2'-modified nucleosides, include, but are not limited to, bicyclic nucleosides wherein the bridge connecting two carbon atoms of the sugar ring connects the 2' carbon and another carbon of the sugar ring; and nucleosides with non-bridging 2'substituents, such as allyl, amino, azido, thio, O-allyl, O--C.sub.1-C.sub.10 alkyl, --OCF.sub.3, O--(CH.sub.2).sub.2--O--CH.sub.3, 2'-O(CH.sub.2).sub.2SCH.sub.3, O--(CH.sub.2).sub.2--O--N(R.sub.m)(R.sub.n), or O--CH.sub.2--C(.dbd.O)--N(R.sub.m)(R.sub.n), where each R.sub.m and R.sub.n is, independently, H or substituted or unsubstituted C.sub.1-C.sub.10 alkyl. 2'-modified nucleosides may further comprise other modifications, for example at other positions of the sugar and/or at the nucleobase.
[0377] As used herein, "2'-F" refers to a nucleoside comprising a sugar comprising a fluoro group at the 2' position of the sugar ring.
[0378] As used herein, "2'-OMe" or "2'-OCH.sub.3" or "2'-O-methyl" each refers to a nucleoside comprising a sugar comprising an --OCH.sub.3 group at the 2' position of the sugar ring.
[0379] As used herein, "MOE" or "2'-MOE" or "2'-OCH.sub.2CH.sub.2OCH.sub.3" or "2'-O-methoxyethyl" each refers to a nucleoside comprising a sugar comprising a --OCH.sub.2CH.sub.2OCH.sub.3 group at the 2' position of the sugar ring.
[0380] As used herein, "oligonucleotide" refers to a compound comprising a plurality of linked nucleosides. In certain embodiments, one or more of the plurality of nucleosides is modified. In certain embodiments, an oligonucleotide comprises one or more ribonucleosides (RNA) and/or deoxyribonucleosides (DNA).
[0381] Many other bicyclo and tricyclo sugar surrogate ring systems are also known in the art that can be used to modify nucleosides for incorporation into antisense compounds (see for example review article: Leumann, Bioorg. Med. Chem., 2002, 10, 841-1954). Such ring systems can undergo various additional substitutions to enhance activity.
[0382] Methods for the preparations of modified sugars are well known to those skilled in the art. Some representative U.S. patents that teach the preparation of such modified sugars include without limitation, U.S.: 4,981,957; 5,118,800; 5,319,080; 5,359,044; 5,393,878; 5,446,137; 5,466,786; 5,514,785; 5,519,134; 5,567,811; 5,576,427; 5,591,722; 5,597,909; 5,610,300; 5,627,053; 5,639,873; 5,646,265; 5,670,633; 5,700,920; 5,792,847 and 6,600,032 and International Application PCT/US2005/019219, filed Jun. 2, 2005 and published as WO 2005/121371 on Dec. 22, 2005, and each of which is herein incorporated by reference in its entirety.
[0383] In nucleotides having modified sugar moieties, the nucleobase moieties (natural, modified or a combination thereof) are maintained for hybridization with an appropriate nucleic acid target.
[0384] In certain embodiments, antisense compounds comprise one or more nucleosides having modified sugar moieties. In certain embodiments, the modified sugar moiety is 2'-MOE. In certain embodiments, the 2'-MOE modified nucleosides are arranged in a gapmer motif. In certain embodiments, the modified sugar moiety is a bicyclic nucleoside having a (4'-CH(CH.sub.3)--O-2') bridging group. In certain embodiments, the (4'-CH(CH.sub.3)--O-2') modified nucleosides are arranged throughout the wings of a gapmer motif.
Conjugated Antisense Compounds
[0385] In certain embodiments, the present disclosure provides conjugated antisense compounds. In certain embodiments, the present disclosure provides conjugated antisense compounds comprising an antisense oligonucleotide complementary to a nucleic acid transcript. In certain embodiments, the present disclosure provides methods comprising contacting a cell with a conjugated antisense compound comprising an antisense oligonucleotide complementary to a nucleic acid transcript. In certain embodiments, the present disclosure provides methods comprising contacting a cell with a conjugated antisense compound comprising an antisense oligonucleotide and reducing the amount or activity of a nucleic acid transcript in a cell.
[0386] The asialoglycoprotein receptor (ASGP-R) has been described previously. See e.g., Park et al., PNAS vol. 102, No. 47, pp 17125-17129 (2005). Such receptors are expressed on liver cells, particularly hepatocytes. Further, it has been shown that compounds comprising clusters of three N-acetylgalactosamine (GalNAc) ligands are capable of binding to the ASGP-R, resulting in uptake of the compound into the cell. See e.g., Khorev et al., Bioorganic and Medicinal Chemistry, 16, 9, pp 5216-5231 (May 2008). Accordingly, conjugates comprising such GalNAc clusters have been used to facilitate uptake of certain compounds into liver cells, specifically hepatocytes. For example it has been shown that certain GalNAc-containing conjugates increase activity of duplex siRNA compounds in liver cells in vivo. In such instances, the GalNAc-containing conjugate is typically attached to the sense strand of the siRNA duplex. Since the sense strand is discarded before the antisense strand ultimately hybridizes with the target nucleic acid, there is little concern that the conjugate will interfere with activity. Typically, the conjugate is attached to the 3' end of the sense strand of the siRNA. See e.g., U.S. Pat. No. 8,106,022. Certain conjugate groups described herein are more active and/or easier to synthesize than conjugate groups previously described.
[0387] In certain embodiments of the present invention, conjugates are attached to single-stranded antisense compounds, including, but not limited to RNase H based antisense compounds and antisense compounds that alter splicing of a pre-mRNA target nucleic acid. In such embodiments, the conjugate should remain attached to the antisense compound long enough to provide benefit (improved uptake into cells) but then should either be cleaved, or otherwise not interfere with the subsequent steps necessary for activity, such as hybridization to a target nucleic acid and interaction with RNase H or enzymes associated with splicing or splice modulation. This balance of properties is more important in the setting of single-stranded antisense compounds than in siRNA compounds, where the conjugate may simply be attached to the sense strand. Disclosed herein are conjugated single-stranded antisense compounds having improved potency in liver cells in vivo compared with the same antisense compound lacking the conjugate. Given the required balance of properties for these compounds such improved potency is surprising.
[0388] In certain embodiments, conjugate groups herein comprise a cleavable moiety. As noted, without wishing to be bound by mechanism, it is logical that the conjugate should remain on the compound long enough to provide enhancement in uptake, but after that, it is desirable for some portion or, ideally, all of the conjugate to be cleaved, releasing the parent compound (e.g., antisense compound) in its most active form. In certain embodiments, the cleavable moiety is a cleavable nucleoside. Such embodiments take advantage of endogenous nucleases in the cell by attaching the rest of the conjugate (the cluster) to the antisense oligonucleotide through a nucleoside via one or more cleavable bonds, such as those of a phosphodiester linkage. In certain embodiments, the cluster is bound to the cleavable nucleoside through a phosphodiester linkage. In certain embodiments, the cleavable nucleoside is attached to the antisense oligonucleotide (antisense compound) by a phosphodiester linkage. In certain embodiments, the conjugate group may comprise two or three cleavable nucleosides. In such embodiments, such cleavable nucleosides are linked to one another, to the antisense compound and/or to the cluster via cleavable bonds (such as those of a phosphodiester linkage). Certain conjugates herein do not comprise a cleavable nucleoside and instead comprise a cleavable bond. It is shown that that sufficient cleavage of the conjugate from the oligonucleotide is provided by at least one bond that is vulnerable to cleavage in the cell (a cleavable bond).
[0389] In certain embodiments, conjugated antisense compounds are prodrugs. Such prodrugs are administered to an animal and are ultimately metabolized to a more active form. For example, conjugated antisense compounds are cleaved to remove all or part of the conjugate resulting in the active (or more active) form of the antisense compound lacking all or some of the conjugate.
[0390] In certain embodiments, conjugates are attached at the 5' end of an oligonucleotide. Certain such 5'-conjugates are cleaved more efficiently than counterparts having a similar conjugate group attached at the 3' end. In certain embodiments, improved activity may correlate with improved cleavage. In certain embodiments, oligonucleotides comprising a conjugate at the 5' end have greater efficacy than oligonucleotides comprising a conjugate at the 3' end (see, for example, Examples 56, 81, 83, and 84). Further, 5'-attachment allows simpler oligonucleotide synthesis. Typically, oligonucleotides are synthesized on a solid support in the 3' to 5' direction. To make a 3'-conjugated oligonucleotide, typically one attaches a pre-conjugated 3' nucleoside to the solid support and then builds the oligonucleotide as usual. However, attaching that conjugated nucleoside to the solid support adds complication to the synthesis. Further, using that approach, the conjugate is then present throughout the synthesis of the oligonucleotide and can become degraded during subsequent steps or may limit the sorts of reactions and reagents that can be used. Using the structures and techniques described herein for 5'-conjugated oligonucleotides, one can synthesize the oligonucleotide using standard automated techniques and introduce the conjugate with the final (5'-most) nucleoside or after the oligonucleotide has been cleaved from the solid support.
[0391] In view of the art and the present disclosure, one of ordinary skill can easily make any of the conjugates and conjugated oligonucleotides herein. Moreover, synthesis of certain such conjugates and conjugated oligonucleotides disclosed herein is easier and/or requires few steps, and is therefore less expensive than that of conjugates previously disclosed, providing advantages in manufacturing. For example, the synthesis of certain conjugate groups consists of fewer synthetic steps, resulting in increased yield, relative to conjugate groups previously described. Conjugate groups such as GalNAc3-10 in Example 46 and GalNAc3-7 in Example 48 are much simpler than previously described conjugates such as those described in U.S. Pat. No. 8,106,022 or 7,262,177 that require assembly of more chemical intermediates. Accordingly, these and other conjugates described herein have advantages over previously described compounds for use with any oligonucleotide, including single-stranded oligonucleotides and either strand of double-stranded oligonucleotides (e.g., siRNA).
[0392] Similarly, disclosed herein are conjugate groups having only one or two GalNAc ligands. As shown, such conjugates groups improve activity of antisense compounds. Such compounds are much easier to prepare than conjugates comprising three GalNAc ligands. Conjugate groups comprising one or two GalNAc ligands may be attached to any antisense compounds, including single-stranded oligonucleotides and either strand of double-stranded oligonucleotides (e.g., siRNA).
[0393] In certain embodiments, the conjugates herein do not substantially alter certain measures of tolerability. For example, it is shown herein that conjugated antisense compounds are not more immunogenic than unconjugated parent compounds. Since potency is improved, embodiments in which tolerability remains the same (or indeed even if tolerability worsens only slightly compared to the gains in potency) have improved properties for therapy.
[0394] In certain embodiments, conjugation allows one to alter antisense compounds in ways that have less attractive consequences in the absence of conjugation. For example, in certain embodiments, replacing one or more phosphorothioate linkages of a fully phosphorothioate antisense compound with phosphodiester linkages results in improvement in some measures of tolerability. For example, in certain instances, such antisense compounds having one or more phosphodiester are less immunogenic than the same compound in which each linkage is a phosphorothioate. However, in certain instances, as shown in Example 26, that same replacement of one or more phosphorothioate linkages with phosphodiester linkages also results in reduced cellular uptake and/or loss in potency. In certain embodiments, conjugated antisense compounds described herein tolerate such change in linkages with little or no loss in uptake and potency when compared to the conjugated full-phosphorothioate counterpart. In fact, in certain embodiments, for example, in Examples 44, 57, 59, and 86, oligonucleotides comprising a conjugate and at least one phosphodiester internucleoside linkage actually exhibit increased potency in vivo even relative to a full phosphorothioate counterpart also comprising the same conjugate. Moreover, since conjugation results in substantial increases in uptake/potency a small loss in that substantial gain may be acceptable to achieve improved tolerability. Accordingly, in certain embodiments, conjugated antisense compounds comprise at least one phosphodiester linkage.
[0395] In certain embodiments, conjugation of antisense compounds herein results in increased delivery, uptake and activity in hepatocytes. Thus, more compound is delivered to liver tissue. However, in certain embodiments, that increased delivery alone does not explain the entire increase in activity. In certain such embodiments, more compound enters hepatocytes. In certain embodiments, even that increased hepatocyte uptake does not explain the entire increase in activity. In such embodiments, productive uptake of the conjugated compound is increased. For example, as shown in Example 102, certain embodiments of GalNAc-containing conjugates increase enrichment of antisense oligonucleotides in hepatocytes versus non-parenchymal cells. This enrichment is beneficial for oligonucleotides that target genes that are expressed in hepatocytes.
[0396] In certain embodiments, conjugated antisense compounds herein result in reduced kidney exposure. For example, as shown in Example 20, the concentrations of antisense oligonucleotides comprising certain embodiments of GalNAc-containing conjugates are lower in the kidney than that of antisense oligonucleotides lacking a GalNAc-containing conjugate. This has several beneficial therapeutic implications. For therapeutic indications where activity in the kidney is not sought, exposure to kidney risks kidney toxicity without corresponding benefit. Moreover, high concentration in kidney typically results in loss of compound to the urine resulting in faster clearance. Accordingly for non-kidney targets, kidney accumulation is undesired.
[0397] In certain embodiments, the present disclosure provides conjugated antisense compounds represented by the formula:
A-B-C-D.BECAUSE.E-F).sub.q
[0398] wherein
[0399] A is the antisense oligonucleotide;
[0400] B is the cleavable moiety
[0401] C is the conjugate linker
[0402] D is the branching group
[0403] each E is a tether;
[0404] each F is a ligand; and
[0405] q is an integer between 1 and 5.
[0406] In the above diagram and in similar diagrams herein, the branching group "D" branches as many times as is necessary to accommodate the number of (E-F) groups as indicated by "q". Thus, where q=1, the formula is:
A-B-C-D-E-F
[0407] where q=2, the formula is:
##STR00023##
[0408] where q=3, the formula is:
##STR00024##
[0409] where q=4, the formula is:
##STR00025##
[0410] where q=5, the formula is:
##STR00026##
[0411] In certain embodiments, conjugated antisense compounds are provided having the structure:
##STR00027##
[0412] In certain embodiments, conjugated antisense compounds are provided having the structure:
##STR00028##
[0413] In certain embodiments, conjugated antisense compounds are provided having the structure:
##STR00029##
[0414] In certain embodiments, conjugated antisense compounds are provided having the structure:
##STR00030##
[0415] In embodiments having more than one of a particular variable (e.g., more than one "m" or "n"), unless otherwise indicated, each such particular variable is selected independently. Thus, for a structure having more than one n, each n is selected independently, so they may or may not be the same as one another.
[0416] i. Certain Cleavable Moieties
[0417] In certain embodiments, a cleavable moiety is a cleavable bond. In certain embodiments, a cleavable moiety comprises a cleavable bond. In certain embodiments, the conjugate group comprises a cleavable moiety. In certain such embodiments, the cleavable moiety attaches to the antisense oligonucleotide. In certain such embodiments, the cleavable moiety attaches directly to the cell-targeting moiety. In certain such embodiments, the cleavable moiety attaches to the conjugate linker. In certain embodiments, the cleavable moiety comprises a phosphate or phosphodiester. In certain embodiments, the cleavable moiety is a cleavable nucleoside or nucleoside analog. In certain embodiments, the nucleoside or nucleoside analog comprises an optionally protected heterocyclic base selected from a purine, substituted purine, pyrimidine or substituted pyrimidine. In certain embodiments, the cleavable moiety is a nucleoside comprising an optionally protected heterocyclic base selected from uracil, thymine, cytosine, 4-N-benzoylcytosine, 5-methylcytosine, 4-N-benzoyl-5-methylcytosine, adenine, 6-N-benzoyladenine, guanine and 2-N-isobutyrylguanine. In certain embodiments, the cleavable moiety is 2'-deoxy nucleoside that is attached to the 3' position of the antisense oligonucleotide by a phosphodiester linkage and is attached to the linker by a phosphodiester or phosphorothioate linkage. In certain embodiments, the cleavable moiety is 2'-deoxy adenosine that is attached to the 3' position of the antisense oligonucleotide by a phosphodiester linkage and is attached to the linker by a phosphodiester or phosphorothioate linkage. In certain embodiments, the cleavable moiety is 2'-deoxy adenosine that is attached to the 3' position of the antisense oligonucleotide by a phosphodiester linkage and is attached to the linker by a phosphodiester linkage.
[0418] In certain embodiments, the cleavable moiety is attached to the 3' position of the antisense oligonucleotide. In certain embodiments, the cleavable moiety is attached to the 5' position of the antisense oligonucleotide. In certain embodiments, the cleavable moiety is attached to a 2' position of the antisense oligonucleotide. In certain embodiments, the cleavable moiety is attached to the antisense oligonucleotide by a phosphodiester linkage. In certain embodiments, the cleavable moiety is attached to the linker by either a phosphodiester or a phosphorothioate linkage. In certain embodiments, the cleavable moiety is attached to the linker by a phosphodiester linkage. In certain embodiments, the conjugate group does not include a cleavable moiety.
[0419] In certain embodiments, the cleavable moiety is cleaved after the complex has been administered to an animal only after being internalized by a targeted cell. Inside the cell the cleavable moiety is cleaved thereby releasing the active antisense oligonucleotide. While not wanting to be bound by theory it is believed that the cleavable moiety is cleaved by one or more nucleases within the cell. In certain embodiments, the one or more nucleases cleave the phosphodiester linkage between the cleavable moiety and the linker. In certain embodiments, the cleavable moiety has a structure selected from among the following:
##STR00031##
wherein each of Bx, Bx.sub.1, Bx.sub.2, and Bx.sub.3 is independently a heterocyclic base moiety. In certain embodiments, the cleavable moiety has a structure selected from among the following:
##STR00032##
[0420] ii. Certain Linkers
[0421] In certain embodiments, the conjugate groups comprise a linker. In certain such embodiments, the linker is covalently bound to the cleavable moiety. In certain such embodiments, the linker is covalently bound to the antisense oligonucleotide. In certain embodiments, the linker is covalently bound to a cell-targeting moiety. In certain embodiments, the linker further comprises a covalent attachment to a solid support. In certain embodiments, the linker further comprises a covalent attachment to a protein binding moiety. In certain embodiments, the linker further comprises a covalent attachment to a solid support and further comprises a covalent attachment to a protein binding moiety. In certain embodiments, the linker includes multiple positions for attachment of tethered ligands. In certain embodiments, the linker includes multiple positions for attachment of tethered ligands and is not attached to a branching group. In certain embodiments, the linker further comprises one or more cleavable bond. In certain embodiments, the conjugate group does not include a linker.
[0422] In certain embodiments, the linker includes at least a linear group comprising groups selected from alkyl, amide, disulfide, polyethylene glycol, ether, thioether (--S--) and hydroxylamino (--O--N(H)--) groups. In certain embodiments, the linear group comprises groups selected from alkyl, amide and ether groups. In certain embodiments, the linear group comprises groups selected from alkyl and ether groups. In certain embodiments, the linear group comprises at least one phosphorus linking group. In certain embodiments, the linear group comprises at least one phosphodiester group. In certain embodiments, the linear group includes at least one neutral linking group. In certain embodiments, the linear group is covalently attached to the cell-targeting moiety and the cleavable moiety. In certain embodiments, the linear group is covalently attached to the cell-targeting moiety and the antisense oligonucleotide. In certain embodiments, the linear group is covalently attached to the cell-targeting moiety, the cleavable moiety and a solid support. In certain embodiments, the linear group is covalently attached to the cell-targeting moiety, the cleavable moiety, a solid support and a protein binding moiety. In certain embodiments, the linear group includes one or more cleavable bond.
[0423] In certain embodiments, the linker includes the linear group covalently attached to a scaffold group. In certain embodiments, the scaffold includes a branched aliphatic group comprising groups selected from alkyl, amide, disulfide, polyethylene glycol, ether, thioether and hydroxylamino groups. In certain embodiments, the scaffold includes a branched aliphatic group comprising groups selected from alkyl, amide and ether groups. In certain embodiments, the scaffold includes at least one mono or polycyclic ring system. In certain embodiments, the scaffold includes at least two mono or polycyclic ring systems. In certain embodiments, the linear group is covalently attached to the scaffold group and the scaffold group is covalently attached to the cleavable moiety and the linker. In certain embodiments, the linear group is covalently attached to the scaffold group and the scaffold group is covalently attached to the cleavable moiety, the linker and a solid support. In certain embodiments, the linear group is covalently attached to the scaffold group and the scaffold group is covalently attached to the cleavable moiety, the linker and a protein binding moiety. In certain embodiments, the linear group is covalently attached to the scaffold group and the scaffold group is covalently attached to the cleavable moiety, the linker, a protein binding moiety and a solid support. In certain embodiments, the scaffold group includes one or more cleavable bond.
[0424] In certain embodiments, the linker includes a protein binding moiety. In certain embodiments, the protein binding moiety is a lipid such as for example including but not limited to cholesterol, cholic acid, adamantane acetic acid, 1-pyrene butyric acid, dihydrotestosterone, 1,3-Bis-O(hexadecyl)glycerol, geranyloxyhexyl group, hexadecylglycerol, borneol, menthol, 1,3-propanediol, heptadecyl group, palmitic acid, myristic acid, O3-(oleoyl)lithocholic acid, O3-(oleoyl)cholenic acid, dimethoxytrityl, or phenoxazine), a vitamin (e.g., folate, vitamin A, vitamin E, biotin, pyridoxal), a peptide, a carbohydrate (e.g., monosaccharide, disaccharide, trisaccharide, tetrasaccharide, oligosaccharide, polysaccharide), an endosomolytic component, a steroid (e.g., uvaol, hecigenin, diosgenin), a terpene (e.g., triterpene, e.g., sarsasapogenin, friedelin, epifriedelanol derivatized lithocholic acid), or a cationic lipid. In certain embodiments, the protein binding moiety is a C16 to C22 long chain saturated or unsaturated fatty acid, cholesterol, cholic acid, vitamin E, adamantane or 1-pentafluoropropyl.
[0425] In certain embodiments, a linker has a structure selected from among:
##STR00033## ##STR00034## ##STR00035##
wherein each n is, independently, from 1 to 20; and p is from 1 to 6.
[0426] In certain embodiments, a linker has a structure selected from among:
##STR00036## ##STR00037##
[0427] wherein each n is, independently, from 1 to 20.
[0428] In certain embodiments, a linker has a structure selected from among:
##STR00038## ##STR00039##
[0429] wherein n is from 1 to 20.
[0430] In certain embodiments, a linker has a structure selected from among:
##STR00040## ##STR00041## ##STR00042## [0431] wherein each L is, independently, a phosphorus linking group or a neutral linking group; and [0432] each n is, independently, from 1 to 20.
[0433] In certain embodiments, a linker has a structure selected from among:
##STR00043## ##STR00044## ##STR00045## ##STR00046##
[0434] In certain embodiments, a linker has a structure selected from among:
##STR00047## ##STR00048##
[0435] In certain embodiments, a linker has a structure selected from among:
##STR00049## ##STR00050## ##STR00051##
[0436] In certain embodiments, a linker has a structure selected from among:
##STR00052##
[0437] wherein n is from 1 to 20.
[0438] In certain embodiments, a linker has a structure selected from among:
##STR00053##
[0439] In certain embodiments, a linker has a structure selected from among:
##STR00054##
[0440] In certain embodiments, a linker has a structure selected from among:
##STR00055##
[0441] In certain embodiments, the conjugate linker has the structure:
##STR00056##
[0442] In certain embodiments, the conjugate linker has the structure:
##STR00057##
[0443] In certain embodiments, a linker has a structure selected from among:
##STR00058##
[0444] In certain embodiments, a linker has a structure selected from among:
##STR00059##
[0445] wherein each n is independently, 0, 1, 2, 3, 4, 5, 6, or 7.
[0446] iii. Certain Cell-Targeting Moieties In certain embodiments, conjugate groups comprise cell-targeting moieties. Certain such cell-targeting moieties increase cellular uptake of antisense compounds. In certain embodiments, cell-targeting moieties comprise a branching group, one or more tether, and one or more ligand. In certain embodiments, cell-targeting moieties comprise a branching group, one or more tether, one or more ligand and one or more cleavable bond.
[0447] 1. Certain Branching Groups
[0448] In certain embodiments, the conjugate groups comprise a targeting moiety comprising a branching group and at least two tethered ligands. In certain embodiments, the branching group attaches the conjugate linker. In certain embodiments, the branching group attaches the cleavable moiety. In certain embodiments, the branching group attaches the antisense oligonucleotide. In certain embodiments, the branching group is covalently attached to the linker and each of the tethered ligands. In certain embodiments, the branching group comprises a branched aliphatic group comprising groups selected from alkyl, amide, disulfide, polyethylene glycol, ether, thioether and hydroxylamino groups. In certain embodiments, the branching group comprises groups selected from alkyl, amide and ether groups. In certain embodiments, the branching group comprises groups selected from alkyl and ether groups. In certain embodiments, the branching group comprises a mono or polycyclic ring system. In certain embodiments, the branching group comprises one or more cleavable bond. In certain embodiments, the conjugate group does not include a branching group.
[0449] In certain embodiments, a branching group has a structure selected from among:
##STR00060## ##STR00061## ##STR00062##
[0450] wherein each n is, independently, from 1 to 20;
[0451] j is from 1 to 3; and
[0452] m is from 2 to 6.
[0453] In certain embodiments, a branching group has a structure selected from among:
##STR00063## ##STR00064##
[0454] wherein each n is, independently, from 1 to 20; and
[0455] m is from 2 to 6.
[0456] In certain embodiments, a branching group has a structure selected from among:
##STR00065## ##STR00066## ##STR00067##
[0457] In certain embodiments, a branching group has a structure selected from among:
##STR00068## [0458] wherein each A.sub.1 is independently, O, S, C.dbd.O or NH; and [0459] each n is, independently, from 1 to 20.
[0460] In certain embodiments, a branching group has a structure selected from among:
##STR00069## [0461] wherein each A.sub.1 is independently, O, S, C.dbd.O or NH; and [0462] each n is, independently, from 1 to 20.
[0463] In certain embodiments, a branching group has a structure selected from among:
##STR00070## [0464] wherein A.sub.1 is O, S, C.dbd.O or NH; and [0465] each n is, independently, from 1 to 20.
[0466] In certain embodiments, a branching group has a structure selected from among:
##STR00071##
[0467] In certain embodiments, a branching group has a structure selected from among:
##STR00072##
[0468] In certain embodiments, a branching group has a structure selected from among:
##STR00073##
[0469] 2. Certain Tethers
[0470] In certain embodiments, conjugate groups comprise one or more tethers covalently attached to the branching group. In certain embodiments, conjugate groups comprise one or more tethers covalently attached to the linking group. In certain embodiments, each tether is a linear aliphatic group comprising one or more groups selected from alkyl, ether, thioether, disulfide, amide and polyethylene glycol groups in any combination. In certain embodiments, each tether is a linear aliphatic group comprising one or more groups selected from alkyl, substituted alkyl, ether, thioether, disulfide, amide, phosphodiester and polyethylene glycol groups in any combination. In certain embodiments, each tether is a linear aliphatic group comprising one or more groups selected from alkyl, ether and amide groups in any combination. In certain embodiments, each tether is a linear aliphatic group comprising one or more groups selected from alkyl, substituted alkyl, phosphodiester, ether and amide groups in any combination. In certain embodiments, each tether is a linear aliphatic group comprising one or more groups selected from alkyl and phosphodiester in any combination. In certain embodiments, each tether comprises at least one phosphorus linking group or neutral linking group.
[0471] In certain embodiments, the tether includes one or more cleavable bond. In certain embodiments, the tether is attached to the branching group through either an amide or an ether group. In certain embodiments, the tether is attached to the branching group through a phosphodiester group. In certain embodiments, the tether is attached to the branching group through a phosphorus linking group or neutral linking group. In certain embodiments, the tether is attached to the branching group through an ether group. In certain embodiments, the tether is attached to the ligand through either an amide or an ether group. In certain embodiments, the tether is attached to the ligand through an ether group. In certain embodiments, the tether is attached to the ligand through either an amide or an ether group. In certain embodiments, the tether is attached to the ligand through an ether group.
[0472] In certain embodiments, each tether comprises from about 8 to about 20 atoms in chain length between the ligand and the branching group. In certain embodiments, each tether group comprises from about 10 to about 18 atoms in chain length between the ligand and the branching group. In certain embodiments, each tether group comprises about 13 atoms in chain length.
[0473] In certain embodiments, a tether has a structure selected from among:
##STR00074##
[0474] wherein each n is, independently, from 1 to 20; and
[0475] each p is from 1 to about 6.
[0476] In certain embodiments, a tether has a structure selected from among:
##STR00075##
[0477] In certain embodiments, a tether has a structure selected from among:
##STR00076## [0478] wherein each n is, independently, from 1 to 20.
[0479] In certain embodiments, a tether has a structure selected from among:
##STR00077## [0480] wherein L is either a phosphorus linking group or a neutral linking group; [0481] Z.sub.1 is C(.dbd.O)O--R.sub.2; [0482] Z.sub.2 is H, C.sub.1-C.sub.6 alkyl or substituted C.sub.1-C.sub.6 alky; [0483] R.sub.2 is H, C.sub.1-C.sub.6 alkyl or substituted C.sub.1-C.sub.6 alky; and [0484] each m.sub.1 is, independently, from 0 to 20 wherein at least one m.sub.1 is greater than 0 for each tether.
[0485] In certain embodiments, a tether has a structure selected from among:
##STR00078##
[0486] In certain embodiments, a tether has a structure selected from among:
##STR00079## [0487] wherein Z.sub.2 is H or CH.sub.3; and [0488] each m.sub.1 is, independently, from 0 to 20 wherein at least one m.sub.1 is greater than 0 for each tether.
[0489] In certain embodiments, a tether has a structure selected from among:
##STR00080## [0490] wherein each n is independently, 0, 1, 2, 3, 4, 5, 6, or 7. [0491] In certain embodiments, a tether comprises a phosphorus linking group. In certain embodiments, a tether does not comprise any amide bonds. In certain embodiments, a tether comprises a phosphorus linking group and does not comprise any amide bonds.
[0492] 3. Certain Ligands
[0493] In certain embodiments, the present disclosure provides ligands wherein each ligand is covalently attached to a tether. In certain embodiments, each ligand is selected to have an affinity for at least one type of receptor on a target cell. In certain embodiments, ligands are selected that have an affinity for at least one type of receptor on the surface of a mammalian liver cell. In certain embodiments, ligands are selected that have an affinity for the hepatic asialoglycoprotein receptor (ASGP-R). In certain embodiments, each ligand is a carbohydrate. In certain embodiments, each ligand is, independently selected from galactose, N-acetyl galactoseamine, mannose, glucose, glucosamone and fucose. In certain embodiments, each ligand is N-acetyl galactoseamine (GalNAc). In certain embodiments, the targeting moiety comprises 2 to 6 ligands. In certain embodiments, the targeting moiety comprises 3 ligands. In certain embodiments, the targeting moiety comprises 3 N-acetyl galactoseamine ligands.
[0494] In certain embodiments, the ligand is a carbohydrate, carbohydrate derivative, modified carbohydrate, multivalent carbohydrate cluster, polysaccharide, modified polysaccharide, or polysaccharide derivative. In certain embodiments, the ligand is an amino sugar or a thio sugar. For example, amino sugars may be selected from any number of compounds known in the art, for example glucosamine, sialic acid, .alpha.-D-galactosamine, N-Acetylgalactosamine, 2-acetamido-2-deoxy-D-galactopyranose (GalNAc), 2-Amino-3-O--[(R)-1-carboxyethyl]-2-deoxy-.beta.-D-glucopyranose (.beta.-muramic acid), 2-Deoxy-2-methylamino-L-glucopyranose, 4,6-Dideoxy-4-formamido-2,3-di-O-methyl-D-mannopyranose, 2-Deoxy-2-sulfoamino-D-glucopyranose and N-sulfo-D-glucosamine, and N-Glycoloyl-.alpha.-neuraminic acid. For example, thio sugars may be selected from the group consisting of 5-Thio-.beta.-D-glucopyranose, Methyl 2,3,4-tri-O-acetyl-1-thio-6-O-trityl-.alpha.-D-glucopyranoside, 4-Thio-.beta.-D-galactopyranose, and ethyl 3,4,6,7-tetra-O-acetyl-2-deoxy-1,5-dithio-.alpha.-D-gluco-heptopyranoside- .
[0495] In certain embodiments, "GalNac" or "Gal-NAc" refers to 2-(Acetylamino)-2-deoxy-D-galactopyranose, commonly referred to in the literature as N-acetyl galactosamine. In certain embodiments, "N-acetyl galactosamine" refers to 2-(Acetylamino)-2-deoxy-D-galactopyranose. In certain embodiments, "GalNac" or "Gal-NAc" refers to 2-(Acetylamino)-2-deoxy-D-galactopyranose. In certain embodiments, "GalNac" or "Gal-NAc" refers to 2-(Acetylamino)-2-deoxy-D-galactopyranose, which includes both the 3-form: 2-(Acetylamino)-2-deoxy-.beta.-D-galactopyranose and .alpha.-form: 2-(Acetylamino)-2-deoxy-D-galactopyranose. In certain embodiments, both the 0-form: 2-(Acetylamino)-2-deoxy-.beta.-D-galactopyranose and .alpha.-form: 2-(Acetylamino)-2-deoxy-D-galactopyranose may be used interchangeably. Accordingly, in structures in which one form is depicted, these structures are intended to include the other form as well. For example, where the structure for an .alpha.-form: 2-(Acetylamino)-2-deoxy-D-galactopyranose is shown, this structure is intended to include the other form as well. In certain embodiments, In certain preferred embodiments, the .beta.-form 2-(Acetylamino)-2-deoxy-D-galactopyranose is the preferred embodiment.
##STR00081## [0496] 2-(Acetylamino)-2-deoxy-D-galactopyranose
[0496] ##STR00082## [0497] 2-(Acetylamino)-2-deoxy-.beta.-D-galactopyranose
[0497] ##STR00083## [0498] 2-(Acetylamino)-2-deoxy-.alpha.-D-galactopyranose
[0499] In certain embodiments one or more ligand has a structure selected from among:
##STR00084##
[0500] wherein each R.sub.1 is selected from OH and NHCOOH.
[0501] In certain embodiments one or more ligand has a structure selected from among:
##STR00085##
[0502] In certain embodiments one or more ligand has a structure selected from among:
##STR00086##
[0503] In certain embodiments one or more ligand has a structure selected from among:
##STR00087##
[0504] i. Certain Conjugates
[0505] In certain embodiments, conjugate groups comprise the structural features above. In certain such embodiments, conjugate groups have the following structure:
##STR00088##
[0506] wherein each n is, independently, from 1 to 20.
[0507] In certain such embodiments, conjugate groups have the following structure:
##STR00089##
[0508] In certain such embodiments, conjugate groups have the following structure:
##STR00090##
[0509] wherein each n is, independently, from 1 to 20;
[0510] Z is H or a linked solid support;
[0511] Q is an antisense compound;
[0512] X is O or S; and
[0513] Bx is a heterocyclic base moiety.
[0514] In certain such embodiments, conjugate groups have the following structure:
##STR00091##
[0515] In certain such embodiments, conjugate groups have the following structure:
##STR00092##
[0516] In certain such embodiments, conjugate groups have the following structure:
##STR00093##
[0517] In certain such embodiments, conjugate groups have the following structure:
##STR00094##
[0518] In certain such embodiments, conjugate groups have the following structure:
##STR00095##
[0519] In certain such embodiments, conjugate groups have the following structure:
##STR00096##
[0520] In certain such embodiments, conjugate groups have the following structure:
##STR00097##
[0521] In certain such embodiments, conjugate groups have the following structure:
##STR00098##
[0522] In certain embodiments, conjugates do not comprise a pyrrolidine.
[0523] In certain such embodiments, conjugate groups have the following structure:
##STR00099##
[0524] In certain such embodiments, conjugate groups have the following structure:
##STR00100##
[0525] In certain such embodiments, conjugate groups have the following structure:
##STR00101##
[0526] In certain such embodiments, conjugate groups have the following structure:
##STR00102##
[0527] In certain such embodiments, conjugate groups have the following structure:
##STR00103##
[0528] In certain such embodiments, conjugate groups have the following structure:
##STR00104##
[0529] In certain such embodiments, conjugate groups have the following structure:
##STR00105##
[0530] In certain such embodiments, conjugate groups have the following structure:
##STR00106##
[0531] In certain such embodiments, conjugate groups have the following structure:
##STR00107##
[0532] In certain such embodiments, conjugate groups have the following structure:
##STR00108##
In certain such embodiments, conjugate groups have the following structure:
##STR00109##
In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00110##
wherein X is a substituted or unsubstituted tether of six to eleven consecutively bonded atoms. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00111##
wherein X is a substituted or unsubstituted tether often consecutively bonded atoms. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00112##
wherein X is a substituted or unsubstituted tether of four to eleven consecutively bonded atoms and wherein the tether comprises exactly one amide bond. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00113##
wherein Y and Z are independently selected from a C.sub.1-C.sub.12 substituted or unsubstituted alkyl, alkenyl, or alkynyl group, or a group comprising an ether, a ketone, an amide, an ester, a carbamate, an amine, a piperidine, a phosphate, a phosphodiester, a phosphorothioate, a triazole, a pyrrolidine, a disulfide, or a thioether. In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00114##
wherein Y and Z are independently selected from a C.sub.1-C.sub.12 substituted or unsubstituted alkyl group, or a group comprising exactly one ether or exactly two ethers, an amide, an amine, a piperidine, a phosphate, a phosphodiester, or a phosphorothioate. In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00115##
wherein Y and Z are independently selected from a C.sub.1-C.sub.12 substituted or unsubstituted alkyl group.
[0533] In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00116##
wherein m and n are independently selected from 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, and 12.
[0534] In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00117##
wherein m is 4, 5, 6, 7, or 8, and n is 1, 2, 3, or 4. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00118##
wherein X is a substituted or unsubstituted tether of four to thirteen consecutively bonded atoms, and wherein X does not comprise an ether group. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00119##
wherein X is a substituted or unsubstituted tether of eight consecutively bonded atoms, and wherein X does not comprise an ether group. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00120##
wherein X is a substituted or unsubstituted tether of four to thirteen consecutively bonded atoms, and wherein the tether comprises exactly one amide bond, and wherein X does not comprise an ether group. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00121##
wherein X is a substituted or unsubstituted tether of four to thirteen consecutively bonded atoms and wherein the tether consists of an amide bond and a substituted or unsubstituted C.sub.2-C.sub.11 alkyl group. In certain embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00122##
wherein Y is selected from a C.sub.1-C.sub.12 substituted or unsubstituted alkyl, alkenyl, or alkynyl group, or a group comprising an ether, a ketone, an amide, an ester, a carbamate, an amine, a piperidine, a phosphate, a phosphodiester, a phosphorothioate, a triazole, a pyrrolidine, a disulfide, or a thioether. In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00123##
wherein Y is selected from a C.sub.1-C.sub.12 substituted or unsubstituted alkyl group, or a group comprising an ether, an amine, a piperidine, a phosphate, a phosphodiester, or a phosphorothioate. In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00124##
wherein Y is selected from a C.sub.1-C.sub.12 substituted or unsubstituted alkyl group. In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00125##
Wherein n is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, or 12.
[0535] In certain such embodiments, the cell-targeting moiety of the conjugate group has the following structure:
##STR00126##
wherein n is 4, 5, 6, 7, or 8.
[0536] a Certain Conjugated Antisense Compounds
[0537] In certain embodiments, the conjugates are bound to a nucleoside of the antisense oligonucleotide at the 2', 3', of 5' position of the nucleoside. In certain embodiments, a conjugated antisense compound has the following structure:
A-B-C-D E-F).sub.q
[0538] wherein
[0539] A is the antisense oligonucleotide;
[0540] B is the cleavable moiety
[0541] C is the conjugate linker
[0542] D is the branching group
[0543] each E is a tether;
[0544] each F is a ligand; and
[0545] q is an integer between 1 and 5.
[0546] In certain embodiments, a conjugated antisense compound has the following structure:
A-C-D E-F).sub.q
[0547] wherein
[0548] A is the antisense oligonucleotide;
[0549] C is the conjugate linker
[0550] D is the branching group
[0551] each E is a tether;
[0552] each F is a ligand; and
[0553] q is an integer between 1 and 5.
[0554] In certain such embodiments, the conjugate linker comprises at least one cleavable bond.
[0555] In certain such embodiments, the branching group comprises at least one cleavable bond.
[0556] In certain embodiments each tether comprises at least one cleavable bond.
In certain embodiments, the conjugates are bound to a nucleoside of the antisense oligonucleotide at the 2', 3', of 5' position of the nucleoside.
[0557] In certain embodiments, a conjugated antisense compound has the following structure:
A-B-C E-F).sub.q
[0558] wherein
[0559] A is the antisense oligonucleotide;
[0560] B is the cleavable moiety
[0561] C is the conjugate linker
[0562] each E is a tether;
[0563] each F is a ligand; and
[0564] q is an integer between 1 and 5.
In certain embodiments, the conjugates are bound to a nucleoside of the antisense oligonucleotide at the 2', 3', of 5' position of the nucleoside. In certain embodiments, a conjugated antisense compound has the following structure:
A-C E-F).sub.q
[0565] wherein
[0566] A is the antisense oligonucleotide;
[0567] C is the conjugate linker
[0568] each E is a tether;
[0569] each F is a ligand; and
[0570] q is an integer between 1 and 5.
In certain embodiments, a conjugated antisense compound has the following structure:
A-B-D E-F).sub.q
[0571] wherein
[0572] A is the antisense oligonucleotide;
[0573] B is the cleavable moiety
[0574] D is the branching group
[0575] each E is a tether;
[0576] each F is a ligand; and
[0577] q is an integer between 1 and 5.
In certain embodiments, a conjugated antisense compound has the following structure:
A-D E-F).sub.q
[0578] wherein
[0579] A is the antisense oligonucleotide;
[0580] D is the branching group
[0581] each E is a tether;
[0582] each F is a ligand; and
[0583] q is an integer between 1 and 5.
[0584] In certain such embodiments, the conjugate linker comprises at least one cleavable bond.
[0585] In certain embodiments each tether comprises at least one cleavable bond.
[0586] In certain embodiments, a conjugated antisense compound has a structure selected from among the following:
##STR00127##
[0587] In certain embodiments, a conjugated antisense compound has a structure selected from among the following:
##STR00128##
[0588] In certain embodiments, a conjugated antisense compound has a structure selected from among the following:
##STR00129##
[0589] Representative United States patents, United States patent application publications, and international patent application publications that teach the preparation of certain of the above noted conjugates, conjugated antisense compounds, tethers, linkers, branching groups, ligands, cleavable moieties as well as other modifications include without limitation, U.S. Pat. Nos. 5,994,517, 6,300,319, 6,660,720, 6,906,182, 7,262,177, 7,491,805, 8,106,022, 7,723,509, US 2006/0148740, US 2011/0123520, WO 2013/033230 and WO 2012/037254, each of which is incorporated by reference herein in its entirety.
[0590] Representative publications that teach the preparation of certain of the above noted conjugates, conjugated antisense compounds, tethers, linkers, branching groups, ligands, cleavable moieties as well as other modifications include without limitation, BIESSEN et al., "The Cholesterol Derivative of a Triantennary Galactoside with High Affinity for the Hepatic Asialoglycoprotein Receptor: a Potent Cholesterol Lowering Agent" J. Med. Chem. (1995) 38:1846-1852, BIESSEN et al., "Synthesis of Cluster Galactosides with High Affinity for the Hepatic Asialoglycoprotein Receptor" J. Med. Chem. (1995) 38:1538-1546, LEE et al., "New and more efficient multivalent glyco-ligands for asialoglycoprotein receptor of mammalian hepatocytes" Bioorganic & Medicinal Chemistry (2011) 19:2494-2500, RENSEN et al., "Determination of the Upper Size Limit for Uptake and Processing of Ligands by the Asialoglycoprotein Receptor on Hepatocytes in Vitro and in Vivo" J. Biol. Chem. (2001) 276(40):37577-37584, RENSEN et al., "Design and Synthesis of Novel N-Acetylgalactosamine-Terminated Glycolipids for Targeting of Lipoproteins to the Hepatic Asialoglycoprotein Receptor" J. Med. Chem. (2004) 47:5798-5808, SLIEDREGT et al., "Design and Synthesis of Novel Amphiphilic Dendritic Galactosides for Selective Targeting of Liposomes to the Hepatic Asialoglycoprotein Receptor" J. Med. Chem. (1999) 42:609-618, and Valentijn et al., "Solid-phase synthesis of lysine-based cluster galactosides with high affinity for the Asialoglycoprotein Receptor" Tetrahedron, 1997, 53(2), 759-770, each of which is incorporated by reference herein in its entirety.
[0591] In certain embodiments, conjugated antisense compounds comprise an RNase H based oligonucleotide (such as a gapmer) or a splice modulating oligonucleotide (such as a fully modified oligonucleotide) and any conjugate group comprising at least one, two, or three GalNAc groups. In certain embodiments a conjugated antisense compound comprises any conjugate group found in any of the following references: Lee, Carbohydr Res, 1978, 67, 509-514; Connolly et al., J Biol Chem, 1982, 257, 939-945; Pavia et al., IntJPep Protein Res, 1983, 22, 539-548; Lee et al., Biochem, 1984, 23, 4255-4261; Lee et al., Glycoconjugate J, 1987, 4, 317-328; Toyokuni et al., Tetrahedron Lett, 1990, 31, 2673-2676; Biessen et al., J Med Chem, 1995, 38, 1538-1546; Valentijn et al., Tetrahedron, 1997, 53, 759-770; Kim et al., Tetrahedron Lett, 1997, 38, 3487-3490; Lee et al., Bioconjug Chem, 1997, 8, 762-765; Kato et al., Glycobiol, 2001, 11, 821-829; Rensen et al., J Biol Chem, 2001, 276, 37577-37584; Lee et al., Methods Enzymol, 2003, 362, 38-43; Westerlind et al., Glycoconj J, 2004, 21, 227-241; Lee et al., Bioorg Med Chem Lett, 2006, 16(19), 5132-5135; Maierhofer et al., Bioorg Med Chem, 2007, 15, 7661-7676; Khorev et al., Bioorg Med Chem, 2008, 16, 5216-5231; Lee et al., Bioorg Med Chem, 2011, 19, 2494-2500; Kornilova et al., Analyt Biochem, 2012, 425, 43-46; Pujol et al., Angew Chemie Int Ed Engl, 2012, 51, 7445-7448; Biessen et al., J Med Chem, 1995, 38, 1846-1852; Sliedregt et al., J Med Chem, 1999, 42, 609-618; Rensen et al., J Med Chem, 2004, 47, 5798-5808; Rensen et al., Arterioscler Thromb Vasc Biol, 2006, 26, 169-175; van Rossenberg et al., Gene Ther, 2004, 11, 457-464; Sato et al., JAm Chem Soc, 2004, 126, 14013-14022; Lee et al., J Org Chem, 2012, 77, 7564-7571; Biessen et al., FASEB J, 2000, 14, 1784-1792; Rajur et al., Bioconjug Chem, 1997, 8, 935-940; Duff et al., Methods Enzymol, 2000, 313, 297-321; Maier et al., Bioconjug Chem, 2003, 14, 18-29; Jayaprakash et al., Org Lett, 2010, 12, 5410-5413; Manoharan, Antisense Nucleic Acid Drug Dev, 2002, 12, 103-128; Merwin et al., Bioconjug Chem, 1994, 5, 612-620; Tomiya et al., Bioorg Med Chem, 2013, 21, 5275-5281; International applications WO 1998/013381; WO2011/038356; WO1997/046098; WO2008/098788; WO2004/101619; WO2012/037254; WO2011/120053; WO2011/100131; WO2011/163121; WO2012/177947; WO2013/033230; WO2013/075035; WO2012/083185; WO2012/083046; WO2009/082607; WO2009/134487; WO2010/144740; WO2010/148013; WO1997/020563; WO2010/088537; WO2002/043771; WO2010/129709; WO2012/068187; WO2009/126933; WO2004/024757; WO2010/054406; WO2012/089352; WO2012/089602; WO2013/166121; WO2013/165816; U.S. Pat. Nos. 4,751,219; 8,552,163; 6,908,903; 7,262,177; 5,994,517; 6,300,319; 8,106,022; 7,491,805; 7,491,805; 7,582,744; 8,137,695; 6,383,812; 6,525,031; 6,660,720; 7,723,509; 8,541,548; 8,344,125; 8,313,772; 8,349,308; 8,450,467; 8,501,930; 8,158,601; 7,262,177; 6,906,182; 6,620,916; 8,435,491; 8,404,862; 7,851,615; Published U.S. Patent Application Publications US2011/0097264; US2011/0097265; US2013/0004427; US2005/0164235; US2006/0148740; US2008/0281044; US2010/0240730; US2003/0119724; US2006/0183886; US2008/0206869; US2011/0269814; US2009/0286973; US2011/0207799; US2012/0136042; US2012/0165393; US2008/0281041; US2009/0203135; US2012/0035115; US2012/0095075; US2012/0101148; US2012/0128760; US2012/0157509; US2012/0230938; US2013/0109817; US2013/0121954; US2013/0178512; US2013/0236968; US2011/0123520; US2003/0077829; US2008/0108801; and US2009/0203132; each of which is incorporated by reference in its entirety.
Cell Culture and Antisense Compounds Treatment
[0592] The effects of antisense compounds on the level, activity, or expression of PKK nucleic acids can be tested in vitro in a variety of cell types. Cell types used for such analyses are available from commercial vendors (e.g., American Type Culture Collection, Manassas, Va.; Zen-Bio, Inc., Research Triangle Park, N.C.; Clonetics Corporation, Walkersville, Md.) and are cultured according to the vendor's instructions using commercially available reagents (e.g., Life Technologies, Carlsbad, Calif.). Illustrative cell types include, but are not limited to, HepaRG.TM.T cells and mouse primary hepatocytes.
In Vitro Testing of Antisense Oligonucleotides
[0593] Described herein are methods for treatment of cells with antisense oligonucleotides, which can be modified appropriately for treatment with other antisense compounds.
[0594] Cells may be treated with antisense oligonucleotides when the cells reach approximately 60-80% confluency in culture.
[0595] One reagent commonly used to introduce antisense oligonucleotides into cultured cells includes the cationic lipid transfection reagent LIPOFECTIN (Life Technologies, Carlsbad, Calif.). Antisense oligonucleotides may be mixed with LIPOFECTIN in OPTI-MEM 1 (Life Technologies, Carlsbad, Calif.) to achieve the desired final concentration of antisense oligonucleotide and a LIPOFECTIN concentration that may range from 2 to 12 ug/mL per 100 nM antisense oligonucleotide.
[0596] Another reagent used to introduce antisense oligonucleotides into cultured cells includes LIPOFECTAMINE (Life Technologies, Carlsbad, Calif.). Antisense oligonucleotide is mixed with LIPOFECTAMINE in OPTI-MEM 1 reduced serum medium (Life Technologies, Carlsbad, Calif.) to achieve the desired concentration of antisense oligonucleotide and a LIPOFECTAMINE concentration that may range from 2 to 12 ug/mL per 100 nM antisense oligonucleotide.
[0597] Another technique used to introduce antisense oligonucleotides into cultured cells includes electroporation.
[0598] Yet another technique used to introduce antisense oligonucleotides into cultured cells includes free uptake of the oligonucleotides by the cells.
[0599] Cells are treated with antisense oligonucleotides by routine methods. Cells may be harvested 16-24 hours after antisense oligonucleotide treatment, at which time RNA or protein levels of target nucleic acids are measured by methods known in the art and described herein. In general, when treatments are performed in multiple replicates, the data are presented as the average of the replicate treatments.
[0600] The concentration of antisense oligonucleotide used varies from cell line to cell line. Methods to determine the optimal antisense oligonucleotide concentration for a particular cell line are well known in the art. Antisense oligonucleotides are typically used at concentrations ranging from 1 nM to 300 nM when transfected with LIPOFECTAMINE. Antisense oligonucleotides are used at higher concentrations ranging from 625 to 20,000 nM when transfected using electroporation.
RNA Isolation
[0601] RNA analysis can be performed on total cellular RNA or poly(A)+ mRNA. Methods of RNA isolation are well known in the art. RNA is prepared using methods well known in the art, for example, using the TRIZOL Reagent (Life Technologies, Carlsbad, Calif.) according to the manufacturer's recommended protocols.
Analysis of Inhibition of Target Levels or Expression
[0602] Inhibition of levels or expression of a PKK nucleic acid can be assayed in a variety of ways known in the art. For example, target nucleic acid levels can be quantitated by, e.g., Northern blot analysis, competitive polymerase chain reaction (PCR), or quantitative real-time PCR. RNA analysis can be performed on total cellular RNA or poly(A)+ mRNA. Methods of RNA isolation are well known in the art. Northern blot analysis is also routine in the art. Quantitative real-time PCR can be conveniently accomplished using the commercially available ABI PRISM 7600, 7700, or 7900 Sequence Detection System, available from PE-Applied Biosystems, Foster City, Calif. and used according to manufacturer's instructions.
Quantitative Real-Time PCR Analysis of Target RNA Levels
[0603] Quantitation of target RNA levels may be accomplished by quantitative real-time PCR using the ABI PRISM 7600, 7700, or 7900 Sequence Detection System (PE-Applied Biosystems, Foster City, Calif.) according to manufacturer's instructions. Methods of quantitative real-time PCR are well known in the art.
[0604] Prior to real-time PCR, the isolated RNA is subjected to a reverse transcriptase (RT) reaction, which produces complementary DNA (cDNA) that is then used as the substrate for the real-time PCR amplification. The RT and real-time PCR reactions are performed sequentially in the same sample well. RT and real-time PCR reagents may be obtained from Life Technologies (Carlsbad, Calif.). RT real-time-PCR reactions are carried out by methods well known to those skilled in the art.
[0605] Gene (or RNA) target quantities obtained by real time PCR are normalized using either the expression level of a gene whose expression is constant, such as cyclophilin A, or by quantifying total RNA using RIBOGREEN (Life Technologies, Inc. Carlsbad, Calif.). Cyclophilin A expression is quantified by real time PCR, by being run simultaneously with the target, multiplexing, or separately. Total RNA is quantified using RIBOGREEN RNA quantification reagent (Invetrogen, Inc. Eugene, Oreg.). Methods of RNA quantification by RIBOGREEN are taught in Jones, L. J., et al, (Analytical Biochemistry, 1998, 265, 368-374). A CYTOFLUOR 4000 instrument (PE Applied Biosystems) is used to measure RIBOGREEN fluorescence.
[0606] Probes and primers are designed to hybridize to a PKK nucleic acid. Methods for designing real-time PCR probes and primers are well known in the art, and may include the use of software such as PRIMER EXPRESS Software (Applied Biosystems, Foster City, Calif.).
Analysis of Protein Levels
[0607] Antisense inhibition of PKK nucleic acids can be assessed by measuring PKK protein levels. Protein levels of PKK can be evaluated or quantitated in a variety of ways well known in the art, such as immunoprecipitation, Western blot analysis (immunoblotting), enzyme-linked immunosorbent assay (ELISA), quantitative protein assays, protein activity assays (for example, caspase activity assays), immunohistochemistry, immunocytochemistry or fluorescence-activated cell sorting (FACS). Antibodies directed to a target can be identified and obtained from a variety of sources, such as the MSRS catalog of antibodies (Aerie Corporation, Birmingham, Mich.), or can be prepared via conventional monoclonal or polyclonal antibody generation methods well known in the art.
In Vivo Testing of Antisense Compounds
[0608] Antisense compounds, for example, antisense oligonucleotides, are tested in animals to assess their ability to inhibit expression of PKK and produce phenotypic changes.
[0609] In certain embodiments, such phenotypic changes include those associated with an inflammatory disease, such as, reduced inflammation, edema/swelling, vascular permeability, and vascular leakage. In certain embodiments, inflammation is measured by measuring the increase or decrease of edema, temperature, pain, color of tissue, and abdominal function in the animal.
[0610] In certain embodiments, such phenotypic changes include those associated with a thromboembolic disease, such as, prolonged aPTT, prolonged aPTT time in conjunction with a normal PT, decreased quantity of Platelet Factor 4 (PF-4), and reduced formation of thrombus or increased time for thrombus formation.
[0611] Testing may be performed in normal animals, or in experimental disease models. For administration to animals, antisense oligonucleotides are formulated in a pharmaceutically acceptable diluent, such as phosphate-buffered saline. Administration includes parenteral routes of administration, such as intraperitoneal, intravenous, and subcutaneous. Calculation of antisense oligonucleotide dosage and dosing frequency is within the abilities of those skilled in the art, and depends upon factors such as route of administration and animal body weight. Following a period of treatment with antisense oligonucleotides, RNA is isolated from liver tissue and changes in PKK nucleic acid expression are measured.
Certain Indications
[0612] In certain embodiments, the invention provides methods of treating an individual comprising administering one or more pharmaceutical compositions as described herein.
[0613] In certain embodiments, the individual has an inflammatory disease. In certain embodiments, the individual is at risk for developing an inflammatory condition, including, but not limited to hereditary angioedema (HAE), edema, angioedema, swelling, angioedema of the lids, ocular edema, macular edema, and cerebral edema. This includes individuals with an acquired problem, disease, or disorder that leads to a risk of inflammation, for example, genetic predisposition to an inflammatory condition, environmental factors, and exposure to certain medications, including, for example, ACE inhibitors and ARBs. In certain embodiments, the individual has been identified as in need of anti-inflammation therapy. Examples of such individuals include, but are not limited to those having a mutation in the genetic code for complement 1 esterase inhibitor (i.e., C1-INH) or Factor 12. In certain embodiments, an abnormal code can lead to a deficiency in C1-INH (i.e., type I HAE), an inability of existing C1-INH to function properly (type II HAE), or hyperfunctional Factor 12 (i.e., type III HAE).
[0614] In certain embodiments, the individual has a thromboembolic disease. In certain embodiments, the individual is at risk for a blood clotting disorder, including, but not limited to, infarct, thrombosis, embolism, thromboembolism such as deep vein thrombosis, pulmonary embolism, myocardial infarction, and stroke. This includes individuals with an acquired problem, disease, or disorder that leads to a risk of thrombosis, for example, surgery, cancer, immobility, sepsis, atherosclerosis atrial fibrillation, as well as genetic predisposition, for example, antiphospholipid syndrome and the autosomal dominant condition, Factor V Leiden. In certain embodiments, the individual has been identified as in need of anticoagulation therapy. Examples of such individuals include, but are not limited to, those undergoing major orthopedic surgery (e.g., hip/knee replacement or hip fracture surgery) and patients in need of chronic treatment, such as those suffering from arterial fibrillation to prevent stroke.
[0615] In certain embodiments the invention provides methods for prophylactically reducing PKK expression in an individual. Certain embodiments include treating an individual in need thereof by administering to an individual a therapeutically effective amount of an antisense compound targeted to a PKK nucleic acid.
[0616] In one embodiment, administration of a therapeutically effective amount of an antisense compound targeted to a PKK nucleic acid is accompanied by monitoring of PKK levels in the serum of an individual, to determine an individual's response to administration of the antisense compound. An individual's response to administration of the antisense compound is used by a physician to determine the amount and duration of therapeutic intervention.
[0617] In certain embodiments, administration of an antisense compound targeted to a PKK nucleic acid results in reduction of PKK expression by at least 15, 20, 25, 30, 35, 40, 45, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99%, or a range defined by any two of these values. In certain embodiments, pharmaceutical compositions comprising an antisense compound targeted to PKK are used for the preparation of a medicament for treating a patient suffering or susceptible to an inflammatory disease or thromboembolic disease.
Certain Compositions
1. ISIS 546254
[0618] In certain embodiments, ISIS 546254 is characterized as a 5-10-5 MOE gapmer, having a sequence of (from 5' to 3') TGCAAGTCTCTTGGCAAACA (incorporated herein as SEQ ID NO: 570), wherein each internucleoside linkage is a phosphorothioate linkage, each cytosine is a 5'-methylcytosine, each of nucleosides 1-5 and 16-20 are 2'-O-methoxyethyl modified nucleosides, and each of nucleosides 6-15 are 2'-deoxynucleosides.
[0619] In certain embodiments, ISIS 546254 is described by the following chemical notation: Tes Ges mCes Aes Aes Gds Tds mCds Tds mCds Tds Tds Gds Gds mCds Aes Aes Aes mCes Ae; wherein,
[0620] A=an adenine,
[0621] mC=a 5'-methylcytosine
[0622] G=a guanine,
[0623] T=a thymine,
[0624] e=a 2'-O-methoxyethyl modified nucleoside,
[0625] d=a 2'-deoxynucleoside, and
[0626] s=a phosphorothioate internucleoside linkage.
[0627] In certain embodiments, ISIS 546254 is described by the following chemical structure:
##STR00130##
[0628] In certain embodiments, as provided in Example 2 (hereinbelow), ISIS 546254 achieved 95% inhibition of human PKK mRNA in cultured HepaRG.TM. cells (density of 20,000 cells per well) when transfected using electroporation with 5,000 nM antisense oligonucleotide after a treatment period of 24 hours and measured by quantitative real-time PCR using human primer probe set RTS3454 and adjusted according to total RNA content, as measured by RIBOGREEN.RTM..
[0629] In certain embodiments, as provided in Example 5 (see Tables 34 and 41 hereinbelow), ISIS 546254 achieved an IC.sub.50 of 0.2 .mu.M and 0.3 .mu.M in a 4 point dose response curve (0.19 .mu.M, 0.56 .mu.M, 1.67 .mu.M, and 5.0 .mu.M) in cultured HepaRG.TM. cells (density of 20,000 cells per well) when transfected using electroporation after a treatment period of 16 and measured by quantitative real-time PCR using human primer probe set RTS3454 and adjusted according to total RNA content, as measured by RIBOGREEN.RTM..
[0630] In certain embodiments, as provided in Example 7 (hereinbelow), ISIS 546254 achieved 31%, 55%, 84%, and 83% human PKK mRNA inhibition and 0%, 36%, 51%, and 76% human PKK protein inhibition in transgenic mice harboring the human PKK gene sequence when injected subcutaneously twice a week for 3 weeks with 2.5 mg/kg/week, 5.0 mg/kg/week, 10 mg/kg/week or 20 mg/kg/week with ISIS 546254.
[0631] In certain embodiments, as provided in Example 8 (hereinbelow), ISISI 546254 is effective for inhibiting PKK mRNA and protein expression and is tolerable in primates.
2. ISIS 546343
[0632] In certain embodiments, ISIS 546343 is characterized as a 5-10-5 MOE gapmer, having a sequence of (from 5' to 3') CCCCCTTCTTTATAGCCAGC (incorporated herein as SEQ ID NO: 705), wherein each internucleoside linkage is a phosphorothioate linkage, each cytosine is a 5'-methylcytosine, each of nucleosides 1-5 and 16-20 are 2'-O-methoxyethyl modified nucleosides, and each of nucleosides 6-15 are 2'-deoxynucleosides.
[0633] In certain embodiments, ISIS 546343 is described by the following chemical notation: mCes mCes mCes mCes mCes Tds Tds mCds Tds Tds Tds Ads Tds Ads Gds mCes mCes Aes Ges mCe; wherein,
[0634] A=an adenine,
[0635] mC=a 5'-methylcytosine;
[0636] G=a guanine,
[0637] T=a thymine,
[0638] e=a 2'-O-methoxyethyl modified nucleoside,
[0639] d=a 2'-deoxynucleoside, and
[0640] s=a phosphorothioate internucleoside linkage.
[0641] In certain embodiments, ISIS 546343 is described by the following chemical structure:
##STR00131##
[0642] In certain embodiments, as provided in Example 2 (see Tables 9 and 10 hereinbelow), ISIS 546343 achieved 97% and 91% human PKK mRNA inhibition in cultured HepaRG.TM. cells (density of 20,000 cells per well) when transfected using electroporation with 5,000 nM antisense oligonucleotide after a treatment period of 24 hours and measured by quantitative real-time PCR using human primer probe set RTS3454 and adjusted according to total RNA content, as measured by RIBOGREEN.RTM..
[0643] In certain embodiments, as provided twice in Example 5 (see Tables 34 and 41 hereinbelow), ISIS 546343 achieved an IC.sub.50 of 0.4 .mu.M in a 4 point dose response curve (0.19 .mu.M, 0.56 .mu.M, 1.67 .mu.M, and 5.0 M) in cultured HepaRG.TM. cells (density of 20,000 cells per well) when transfected using electroporation after a treatment period of 16 and measured by quantitative real-time PCR using human primer probe set RTS3454 and adjusted according to total RNA content, as measured by RIBOGREEN.RTM..
[0644] In certain embodiments, as provided in Example 7 (hereinbelow), ISIS 546343 achieved 46%, 66%, and 86% human PKK mRNA inhibition and 0%, 38%, and 79% human PKK protein inhibition in transgenic mice harboring the human PKK gene sequence when injected subcutaneously twice a week for 3 weeks with 2.5 mg/kg/week, 5.0 mg/kg/week, 10 mg/kg/week or 20 mg/kg/week with ISIS 546343.
[0645] In certain embodiments, as provided in Example 8 (hereinbelow), ISISI 546343 is effective for inhibiting PKK mRNA and protein expression and is tolerable in primates.
3. ISIS 548048
[0646] In certain embodiments, ISIS 548048 is characterized as a modified antisense oligonucleotide having the nucleobase sequence (from 5' to 3') CGATATCATGATTCCC (incorporated herein as SEQ ID NO: 1666), consisting of a combination of sixteen 2'-deoxynucleosides, 2'-O-methoxyethyl modified nucleosides, and cEt modified nucleosides, wherein each of nucleosides 1, 2, and 16 are 2'-O-methoxyethyl modified nucleosides, wherein each of nucleosides 3, 14, and 15 are cEt modified nucleosides, wherein each of nucleosides 4-13 are 2'-deoxynucleosides, wherein each internucleoside linkage is a phosphorothioate internucleoside linkage, and wherein each cytosine is a 5'-methylcytosine.
[0647] In certain embodiments, ISIS 548048 is described by the following chemical notation: mCes Ges Aks Tds Ads Tds mCds Ads Tds Gds Ads Tds Tds mCks mCks mCe; wherein,
[0648] A=an adenine,
[0649] mC=a 5'-methylcytosine;
[0650] G=a guanine,
[0651] T=a thymine,
[0652] e=a 2'-O-methoxyethyl modified nucleoside,
[0653] k=a cEt modified nucleoside,
[0654] d=a 2'-deoxynucleoside, and
[0655] s=a phosphorothioate internucleoside linkage.
[0656] In certain embodiments, ISIS 548048 is described by the following chemical structure:
##STR00132##
[0657] In certain embodiments, as provided in Example 3 (hereinbelow), ISIS 548048 achieved 84% mRNA inhibition in cultured HepaRG.TM. cells (density of 20,000 cells per well) when transfected using electroporation with 1,000 nM antisense oligonucleotide after a treatment period of 24 hours and measured by quantitative real-time PCR using human primer probe set RTS3454 and adjusted according to total RNA content, as measured by RIBOGREEN.RTM..
[0658] In certain embodiments, as provided in Example 6 (hereinbelow), ISIS 548048 achieved an IC.sub.50 of 0.1 .mu.M in a 4 point dose response curve (0.11 .mu.M, 0.33 .mu.M, 1.00 .mu.M, and 3.00 .mu.M) in cultured HepaRG.TM. cells (density of 20,000 cells per well) when transfected using electroporation after a treatment period of 16 and measured by quantitative real-time PCR using human primer probe set RTS3454 and adjusted according to total RNA content, as measured by RIBOGREEN.
[0659] In certain embodiments, as provided in Example 7 (hereinbelow), ISIS 548048 achieved 7%, 77%, 72% and 80% human PKK mRNA inhibition and 23%, 70%, 89%, and 98% human PKK protein inhibition in transgenic mice harboring the human PKK gene sequence when injected subcutaneously twice a week for 3 weeks with 2.5 mg/kg/week, 5.0 mg/kg/week, 10 mg/kg/week or 20 mg/kg/week with ISIS 548048.
[0660] In certain embodiments, as provided in Example 8 (hereinbelow), ISISI 548048 is effective for inhibiting PKK mRNA and protein expression and is tolerable in primates.
4. ISIS 721744
[0661] In certain embodiments, ISIS 721744 is characterized as a 5-10-5 MOE gapmer, having a sequence of (from 5' to 3') TGCAAGTCTCTTGGCAAACA (incorporated herein as SEQ ID NO: 570), wherein the internucleoside linkages between nucleosides 3 to 4, 4 to 5, 16 to 17, and 17 to 18 are phosphodiester linkages and the internucleoside linkages between nucleosides 1 to 2, 2 to 3, 5 to 6, 6 to 7, 7 to 8, 8 to 9, 9 to 10, 10 to 11, 11 to 12, 12 to 13, 13 to 14, 14 to 15, 15 to 16, 18 to 19, and 19 to 20 are phosphorothioate linkages, each cytosine is a 5'-methylcytosine, each of nucleosides 1-5 and 16-20 are 2'-O-methoxyethyl modified nucleosides, and each of nucleosides 6-15 are 2'-deoxynucleosides.
[0662] In certain embodiments, ISIS 721744 is described by the following chemical notation: GalNAc3-7.sub.a-0 Tes Ges mCeo Aeo Aes Gds Tds mCds Tds mCds Tds Tds Gds Gds mCds Aeo Aeo Aes mCes Ae; wherein,
[0663] A=an adenine,
[0664] mC=a 5'-methylcytosine
[0665] G=a guanine,
[0666] T=a thymine,
[0667] e=a 2'-O-methoxyethyl modified nucleoside,
[0668] d=a 2'-deoxynucleoside,
[0669] o=a phosphodiester internucleoside linkage,
[0670] s=a phosphorothioate internucleoside linkage, and
[0671] GalNAc3-7.sub.a-o=
##STR00133##
[0672] In certain embodiments, ISIS 721744 is described by the following chemical structure:
##STR00134##
Certain Hotspot Regions
1. Nucleobases 27427-27466 of SEQ ID NO: 10
[0673] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 27427-27466 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 27427-27466 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 27427-27466 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense oligonucleotides are linked by phosphorothioate internucleoside linkages.
[0674] In certain embodiments, nucleobases 27427-27466 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 530993, 530994, 530995, 546251, 546252, 546253, 546254, 546255, 546256, 547410, 547411, 547978, 547979, 547980, and 547981.
[0675] In certain embodiments, nucleobases nucleobases 27427-27466 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 94, 95, 96, 566, 567, 568, 569, 570, 571, 572, 573, 1597, 1598, 1599, and 1600.
[0676] In certain embodiments, antisense oligonucleotides targeting nucleobases 27427-27466 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
2. Nucleobases 33183-33242 of SEQ ID NO: 10
[0677] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 33183-33242 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 33183-33242 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 33183-33242 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense oligonucleotides are linked by phosphorothioate internucleoside linkages.
[0678] In certain embodiments, nucleobases 33183-33242 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 531052, 531053, 531054, 531055, 531056, 531057, 531158, 546343, 546345, 547480, 547481, 547482, and 547483.
[0679] In certain embodiments, nucleobases nucleobases 33183-33242 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 155, 156, 157, 158, 159, 160, 261, 702, 703, 704, 705, 706, and 707.
[0680] In certain embodiments, antisense oligonucleotides targeting nucleobases 33183-33242 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK mRNA and/or protein levels in vitro and/or in vivo.
3. Nucleobases 30570-30610 of SEQ ID NO: 10
[0681] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 30570-30610 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 30570-30610 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 30570-30610 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense oligonucleotides are linked by phosphorothioate internucleoside linkages.
[0682] In certain embodiments, nucleobases 30570-30610 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 531026, 546309, 546310, 546311, 546313, 547453, 547454, 547455, 547456, 547457, 547458, 548046, 548047, 548048, 548049, and 548050.
[0683] In certain embodiments, nucleobases nucleobases 30570-30610 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 129, 652, 653, 654, 655, 656, 657, 658, 659, 660, 661, 1664, 1665, 1666, 1667, and 1668.
[0684] In certain embodiments, antisense oligonucleotides targeting nucleobases 30570-30610 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK mRNA and/or protein levels in vitro and/or in vivo.
4. Nucleobases 27427-27520 of SEQ ID NO: 10
[0685] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 27427-27520 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 27427-27520 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 27427-27520 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense oligonucleotides are linked by phosphorothioate internucleoside linkages.
[0686] In certain embodiments, nucleobases 27427-27520 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 530993-530999, 546251-546256, 546258-546260, 546263, 546265-546268, 547410-547417, and 547978-547992.
[0687] In certain embodiments, nucleobases nucleobases 27427-27520 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 94-100, 566-587, and 1597-1611.
[0688] In certain embodiments, antisense oligonucleotides targeting nucleobases 27427-27520 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
5. Nucleobases 33085-33247 of SEQ ID NO: 10
[0689] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 33085-33247 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 33085-33247 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 33085-33247 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense oligonucleotides are linked by phosphorothioate internucleoside linkages.
[0690] In certain embodiments, nucleobases 33085-33247 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 531041-531158, 546336, 546339, 546340, 546343, 546345, 547474-547483, 547778, 548077-548082, and 548677-548678.
[0691] In certain embodiments, nucleobases nucleobases 33085-33247 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 144-160, 261, 693-707, 1256, 1320-1325, 2214, and 2215. In certain embodiments, antisense oligonucleotides targeting nucleobases 33085-33247 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
6. Nucleobases 30475-30639 of SEQ ID NO: 10
[0692] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 30475-30639 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 30475-30639 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 30475-30639 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense olignonucleotides are linked by phosphorothioate internucleoside linkages.
[0693] In certain embodiments, nucleobases 30475-30639 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 531021-531029, 531146, 546297, 546299-546304, 546306-546311, 546313, 546316-546319, 547444-547462, 548031, 548032, and 548034-548056.
[0694] In certain embodiments, nucleobases nucleobases 30475-30639 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 124-132, 249, 633-669, and 1650-1674.
[0695] In certain embodiments, antisense oligonucleotides targeting nucleobases 30475-30639 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
7. Nucleobases 27362-27524 of SEQ ID NO: 10
[0696] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 27362-27524 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 27362-27524 correspond to exon 9 of PKK (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 27362-27524 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 27362-27524 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense olignonucleotides are linked by phosphorothioate internucleoside linkages.
[0697] In certain embodiments, nucleobases 27361-27524 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 530985-530999, 546244, 546247-546256, 546258-546260, 546263, 546265-546268, 547403-547417, 547723, 547968-547970, and 547972-547992.
[0698] In certain embodiments, nucleobases nucleobases 27361-27524 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 86-100, 554-587, 1217, and 1588-1611.
[0699] In certain embodiments, antisense oligonucleotides targeting nucleobases 27362-27524 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
8. Nucleobases 33101-33240 of SEQ ID NO: 10
[0700] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 33101-33240 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 33101-33240 correspond to exon 14 of PKK (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 33101-33240 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 33101-33240 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense olignonucleotides are linked by phosphorothioate internucleoside linkages.
[0701] In certain embodiments, nucleobases 33101-33240 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 531041-531158, 546336, 546339, 546340, 546343, 546345, 547474-547483, 548077-548082, and 548678-548678.
[0702] In certain embodiments, nucleobases nucleobases 33101-33240 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 144-160, 261, 693-707, 1320-1325, and 2215.
[0703] In certain embodiments, antisense oligonucleotides targeting nucleobases 33101-33240 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
9. Nucleobases 30463-30638 of SEQ ID NO: 10
[0704] In certain embodiments, antisense oligonucleotides are designed to target nucleobases 30463-30638 of SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 30463-30638 correspond to exon 12 of PKK (GENBANK Accession No. NT_016354.19 truncated from nucleobases 111693001 to Ser. No. 11/730,000). In certain embodiments, nucleobases 30463-30638 of SEQ ID NO: 10 are a hotspot region. In certain embodiments, nucleobases 30463-30638 of SEQ ID NO: 10 are targeted by antisense oligonucleotides. In certain embodiments, the antisense oligonucleotides are 15, 16, 17, 18, 19, or 20 nucleobases in length. In certain embodiments, the antisense oligonucleotides are gapmers. In certain embodiments, the gapmers are 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. In certain embodiments, the gapmers are 5-10-5 MOE and cEt gapmers, 4-9-4 MOE and cEt gapmers, 4-10-4 MOE and cEt gapmers, 4-10-3 MOE and cEt gapmers, 3-10-4 MOE and cEt gapmers, or 3-10-3 MOE and cEt gapmers. In certain embodiments, the nucleosides of the antisense olignonucleotides are linked by phosphorothioate internucleoside linkages.
[0705] In certain embodiments, nucleobases 30463-30638 of SEQ ID NO: 10 are targeted by the following ISIS numbers: 531021-531029, 531146, 546297, 546299-546304, 546306-546311, 546313, 546316-546319, 547444-547462, 548031, 548032, and 548034-548056.
[0706] In certain embodiments, nucleobases nucleobases 30463-30638 of SEQ ID NO: 10 are targeted by the following SEQ ID NOs: 124-132, 249, 633-669, and 1650-1674.
[0707] In certain embodiments, antisense oligonucleotides targeting nucleobases 30463-30638 of SEQ ID NO: 10 achieve at least 30%, at least 31%, at least 32%, at least 33%, at least 34%, at least 35%, at least 36%, at least 37%, at least 38%, at least 39%, at least 40%, at least 41%, at least 42%, at least 43%, at least 44%, at least 45%, at least 46%, at least 47%, at least 48%, at least 49%, at least 50%, at least 51%, at least 52%, at least 53%, at least 54%, at least 55%, at least 56%, at least 57%, at least 58%, at least 59%, at least 60%, at least 61%, at least 62%, at least 63%, at least 64%, at least 65%, at least 66%, at least 67%, at least 68%, at least 69%, at least 70%, at least 71%, at least 72%, at least 73%, at least 74%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% reduction of PKK and/or protein levels in vitro and/or in vivo.
EXAMPLES
Non-Limiting Disclosure and Incorporation by Reference
[0708] While certain compounds, compositions and methods described herein have been described with specificity in accordance with certain embodiments, the following examples serve only to illustrate the compounds described herein and are not intended to limit the same. Each of the references recited in the present application is incorporated herein by reference in its entirety.
[0709] The following examples illustrate certain embodiments of the present disclosure and are not limiting. Moreover, where specific embodiments are provided, the inventors have contemplated generic application of those specific embodiments. For example, disclosure of an oligonucleotide having a particular motif provides reasonable support for additional oligonucleotides having the same or similar motif. And, for example, where a particular high-affinity modification appears at a particular position, other high-affinity modifications at the same position are considered suitable, unless otherwise indicated.
Example 1: General Method for the Preparation of Phosphoramidites, Compounds 1, 1a and 2
##STR00135##
[0711] Bx is a heterocyclic base;
[0712] Compounds 1, 1a and 2 were prepared as per the procedures well known in the art as described in the specification herein (see Seth et al., Bioorg. Med. Chem., 2011, 21(4), 1122-1125, J. Org. Chem., 2010, 75(5), 1569-1581, Nucleic Acids Symposium Series, 2008, 52(1), 553-554); and also see published PCT International Applications (WO 2011/115818, WO 2010/077578, WO2010/036698, WO2009/143369, WO 2009/006478, and WO 2007/090071), and U.S. Pat. No. 7,569,686).
Example 2: Preparation of Compound 7
##STR00136##
[0714] Compounds 3 (2-acetamido-1,3,4,6-tetra-O-acetyl-2-deoxy-.beta.-Dgalactopyranose or galactosamine pentaacetate) is commercially available. Compound 5 was prepared according to published procedures (Weber et al., J. Med. Chem., 1991, 34, 2692).
Example 3: Preparation of Compound 11
##STR00137##
[0716] Compounds 8 and 9 are commercially available.
Example 4: Preparation of Compound 18
##STR00138## ##STR00139##
[0718] Compound 11 was prepared as per the procedures illustrated in Example 3. Compound 14 is commercially available. Compound 17 was prepared using similar procedures reported by Rensen et al., J. Med. Chem., 2004, 47, 5798-5808.
Example 5: Preparation of Compound 23
##STR00140##
[0720] Compounds 19 and 21 are commercially available.
Example 6: Preparation of Compound 24
##STR00141##
[0722] Compounds 18 and 23 were prepared as per the procedures illustrated in Examples 4 and 5.
Example 7: Preparation of Compound 25
##STR00142##
[0724] Compound 24 was prepared as per the procedures illustrated in Example 6.
Example 8: Preparation of Compound 26
##STR00143##
[0726] Compound 24 is prepared as per the procedures illustrated in Example 6.
Example 9: General Preparation of Conjugated ASOs Comprising GalNAc.sub.3-1 at the 3' Terminus, Compound 29
[0727] ##STR00144## ##STR00145## [0728] Wherein the protected GalNAc.sub.3-1 has the structure:
##STR00146##
[0729] The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-1 (GalNAc.sub.3-1.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. Wherein GalNAc.sub.3-1.sub.a has the formula:
##STR00147##
[0730] The solid support bound protected GalNAc.sub.3-1, Compound 25, was prepared as per the procedures illustrated in Example 7. Oligomeric Compound 29 comprising GalNAc.sub.3-1 at the 3' terminus was prepared using standard procedures in automated DNA/RNA synthesis (see Dupouy et al., Angew. Chem. Int. Ed., 2006, 45, 3623-3627). Phosphoramidite building blocks, Compounds 1 and 1a were prepared as per the procedures illustrated in Example 1. The phosphoramidites illustrated are meant to be representative and not intended to be limiting as other phosphoramidite building blocks can be used to prepare oligomeric compounds having a predetermined sequence and composition. The order and quantity of phosphoramidites added to the solid support can be adjusted to prepare gapped oligomeric compounds as described herein. Such gapped oligomeric compounds can have predetermined composition and base sequence as dictated by any given target.
Example 10: General Preparation Conjugated ASOs Comprising GalNAc.sub.3-1 at the 5' Terminus, Compound 34
##STR00148## ##STR00149##
[0732] The Unylinker.TM. 30 is commercially available. Oligomeric Compound 34 comprising a GalNAc.sub.3-1 cluster at the 5' terminus is prepared using standard procedures in automated DNA/RNA synthesis (see Dupouy et al., Angew. Chem. Int. Ed., 2006, 45, 3623-3627). Phosphoramidite building blocks, Compounds 1 and 1a were prepared as per the procedures illustrated in Example 1. The phosphoramidites illustrated are meant to be representative and not intended to be limiting as other phosphoramidite building blocks can be used to prepare an oligomeric compound having a predetermined sequence and composition. The order and quantity of phosphoramidites added to the solid support can be adjusted to prepare gapped oligomeric compounds as described herein. Such gapped oligomeric compounds can have predetermined composition and base sequence as dictated by any given target.
Example 11: Preparation of Compound 39
##STR00150## ##STR00151##
[0734] Compounds 4, 13 and 23 were prepared as per the procedures illustrated in Examples 2, 4, and 5. Compound 35 is prepared using similar procedures published in Rouchaud et al., Eur. J. Org. Chem., 2011, 12, 2346-2353.
Example 12: Preparation of Compound 40
##STR00152##
[0736] Compound 38 is prepared as per the procedures illustrated in Example 11.
Example 13: Preparation of Compound 44
##STR00153## ##STR00154##
[0738] Compounds 23 and 36 are prepared as per the procedures illustrated in Examples 5 and 11. Compound 41 is prepared using similar procedures published in WO 2009082607.
Example 14: Preparation of Compound 45
##STR00155##
[0740] Compound 43 is prepared as per the procedures illustrated in Example 13.
Example 15: Preparation of Compound 47
##STR00156##
[0742] Compound 46 is commercially available.
Example 16: Preparation of Compound 53
##STR00157## ##STR00158##
[0744] Compounds 48 and 49 are commercially available. Compounds 17 and 47 are prepared as per the procedures illustrated in Examples 4 and 15.
Example 17: Preparation of Compound 54
##STR00159##
[0746] Compound 53 is prepared as per the procedures illustrated in Example 16.
Example 18: Preparation of Compound 55
##STR00160##
[0748] Compound 53 is prepared as per the procedures illustrated in Example 16.
Example 19: General Method for the Preparation of Conjugated ASOs Comprising GalNAc.sub.3-1 at the 3' Position Via Solid Phase Techniques (Preparation of ISIS 647535, 647536 and 651900)
[0749] Unless otherwise stated, all reagents and solutions used for the synthesis of oligomeric compounds are purchased from commercial sources. Standard phosphoramidite building blocks and solid support are used for incorporation nucleoside residues which include for example T, A, G, and C residues. A 0.1 M solution of phosphoramidite in anhydrous acetonitrile was used for .beta.-D-2'-deoxyribonucleoside and 2'-MOE.
[0750] The ASO syntheses were performed on ABI 394 synthesizer (1-2 .mu.mol scale) or on GE Healthcare Bioscience KTA oligopilot synthesizer (40-200 .mu.mol scale) by the phosphoramidite coupling method on an GalNAc.sub.3-1 loaded VIMAD solid support (110 .mu.mol/g, Guzaev et al., 2003) packed in the column. For the coupling step, the phosphoramidites were delivered 4 fold excess over the loading on the solid support and phosphoramidite condensation was carried out for 10 min. All other steps followed standard protocols supplied by the manufacturer. A solution of 6% dichloroacetic acid in toluene was used for removing dimethoxytrityl (DMT) group from 5'-hydroxyl group of the nucleotide. 4,5-Dicyanoimidazole (0.7 M) in anhydrous CH.sub.3CN was used as activator during coupling step. Phosphorothioate linkages were introduced by sulfurization with 0.1 M solution of xanthane hydride in 1:1 pyridine/CH.sub.3CN for a contact time of 3 minutes. A solution of 20% tert-butylhydroperoxide in CH.sub.3CN containing 6% water was used as an oxidizing agent to provide phosphodiester internucleoside linkages with a contact time of 12 minutes.
[0751] After the desired sequence was assembled, the cyanoethyl phosphate protecting groups were deprotected using a 1:1 (v/v) mixture of triethylamine and acetonitrile with a contact time of 45 minutes. The solid-support bound ASOs were suspended in aqueous ammonia (28-30 wt %) and heated at 55.degree. C. for 6 h.
[0752] The unbound ASOs were then filtered and the ammonia was boiled off. The residue was purified by high pressure liquid chromatography on a strong anion exchange column (GE Healthcare Bioscience, Source 30Q, 30 .mu.m, 2.54.times.8 cm, A=100 mM ammonium acetate in 30% aqueous CH.sub.3CN, B=1.5 M NaBr in A, 0-40% of B in 60 min, flow 14 mL min-1, .lamda.=260 nm). The residue was desalted by HPLC on a reverse phase column to yield the desired ASOs in an isolated yield of 15-30% based on the initial loading on the solid support. The ASOs were characterized by ion-pair-HPLC coupled MS analysis with Agilent 1100 MSD system.
[0753] Antisense oligonucleotides not comprising a conjugate were synthesized using standard oligonucleotide synthesis procedures well known in the art.
[0754] Using these methods, three separate antisense compounds targeting ApoC III were prepared. As summarized in Table 17, below, each of the three antisense compounds targeting ApoC III had the same nucleobase sequence; ISIS 304801 is a 5-10-5 MOE gapmer having all phosphorothioate linkages; ISIS 647535 is the same as ISIS 304801, except that it had a GalNAc.sub.3-1 conjugated at its 3'end; and ISIS 647536 is the same as ISIS 647535 except that certain internucleoside linkages of that compound are phosphodiester linkages. As further summarized in Table 17, two separate antisense compounds targeting SRB-1 were synthesized. ISIS 440762 was a 2-10-2 cEt gapmer with all phosphorothioate internucleoside linkages; ISIS 651900 is the same as ISIS 440762, except that it included a GalNAc.sub.3-1 at its 3'-end.
TABLE-US-00001 TABLE 17 Modified ASO targeting ApoC III and SRB-1 SEQ CalCd Observed ID ASO Sequence (5' to 3') Target Mass Mass No. ISIS A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC- .sub.dsT.sub.esT.sub.esT.sub.esA.sub.esT.sub.e ApoC 7165.4 7164.4 2248 304801 III ISIS A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC- .sub.dsT.sub.esT.sub.esT.sub.esA.sub.esT.sub.eoA.sub.do'- ApoC 9239.5 9237.8 2249 647535 GalNAc.sub.3-1.sub.a III ISIS A.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC- .sub.dsT.sub.eoT.sub.eoT.sub.esA.sub.esT.sub.eoA.sub.do'- ApoC 9142.9 9140.8 2249 647536 GalNAc.sub.3-1.sub.a III ISIS T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT- .sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k SRB-1 4647.0 4646.4 2250 440762 ISIS T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT- .sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.koA.sub.d- o'-GalNAc.sub.3-1.sub.a SRB-1 6721.1 6719.4 2251 651900 Subscripts: "e" indicates 2'-M0E modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonuclcoside; "k" indicates 6'-(S)-CH.sub.3 bicyclic nucleoside (e.g. cEt); "s" indicates phosphorothioate intemuclcosidc linkages (PS); "o" indicates phosphodiester intemucleoside linkages (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Superscript "m" indicates 5-mcthylcytosines. "GalNAc.sub.3-1" indicates a conjugate group having the structure shown previously in Example 9. Note that GalNAc.sub.3-1 comprises a cleavable adenosine which links the ASO to remainder of the conjugate, which is designated "GalNAc.sub.3-1.sub.a." This nomenclature is used in the above table to show the full nuclcobase sequence, including the adenosine, which is part of the conjugate. Thus, in the above table, the sequences could also be listed as ending with "GalNAc.sub.3-1" with the "A.sub.do" omitted. This convention of using the subscript "a" to indicate the portion of a conjugate group lacking a cleavable nucleoside or cleavable moiety is used throughout these Examples. This portion of a conjugate group lacking the cleavable moiety is referred to herein as a "cluster" or "conjugate cluster" or "GalNAc.sub.3 cluster." In certain instances it is convenient to describe a conjugate group by separately providing its cluster and its cleavable moiety.
Example 20: Dose-Dependent Antisense Inhibition of Human ApoC III in huApoC III Transgenic Mice
[0755] ISIS 304801 and ISIS 647535, each targeting human ApoC III and described above, were separately tested and evaluated in a dose-dependent study for their ability to inhibit human ApoC III in human ApoC III transgenic mice.
Treatment
[0756] Human ApoCIII transgenic mice were maintained on a 12-hour light/dark cycle and fed ad libitum Teklad lab chow. Animals were acclimated for at least 7 days in the research facility before initiation of the experiment. ASOs were prepared in PBS and sterilized by filtering through a 0.2 micron filter. ASOs were dissolved in 0.9% PBS for injection.
[0757] Human ApoC III transgenic mice were injected intraperitoneally once a week for two weeks with ISIS 304801 or 647535 at 0.08, 0.25. 0.75, 2.25 or 6.75 .mu.mol/kg or with PBS as a control. Each treatment group consisted of 4 animals. Forty-eight hours after the administration of the last dose, blood was drawn from each mouse and the mice were sacrificed and tissues were collected.
ApoC III mRNA Analysis
[0758] ApoC III mRNA levels in the mice's livers were determined using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. ApoC III mRNA levels were determined relative to total RNA (using Ribogreen), prior to normalization to PBS-treated control. The results below are presented as the average percent of ApoC III mRNA levels for each treatment group, normalized to PBS-treated control and are denoted as "% PBS". The half maximal effective dosage (ED.sub.50) of each ASO is also presented in Table 18, below.
[0759] As illustrated, both antisense compounds reduced ApoC III RNA relative to the PBS control. Further, the antisense compound conjugated to GalNAc.sub.3-1 (ISIS 647535) was substantially more potent than the antisense compound lacking the GalNAc.sub.3-1 conjugate (ISIS 304801).
TABLE-US-00002 TABLE 18 Effect of ASO treatment on ApoC III mRNA levels in human ApoC III transgenic mice Dose ED.sub.50 Internucleoside SEQ (.mu.mol/ % (.mu.mol/ 3' linkage/ ID ASO kg) PBS kg) Conjugate Length No. PBS 0 100 -- -- -- ISIS 0.08 95 0.77 None PS/20 2248 304801 0.75 42 2.25 32 6.75 19 ISIS 0.08 50 0.074 GalNAc.sub.3-1 PS/20 2249 647535 0.75 15 2.25 17 6.75 8
ApoC III Protein Analysis (Turbidometric Assay)
[0760] Plasma ApoC III protein analysis was determined using procedures reported by Graham et al, Circulation Research, published online before print Mar. 29, 2013.
[0761] Approximately 100 .mu.l of plasma isolated from mice was analyzed without dilution using an Olympus Clinical Analyzer and a commercially available turbidometric ApoC III assay (Kamiya, Cat # KAI-006, Kamiya Biomedical, Seattle, Wash.). The assay protocol was performed as described by the vendor.
[0762] As shown in the Table 19 below, both antisense compounds reduced ApoC III protein relative to the PBS control. Further, the antisense compound conjugated to GalNAc.sub.3-1 (ISIS 647535) was substantially more potent than the antisense compound lacking the GalNAc.sub.3-1 conjugate (ISIS 304801).
TABLE-US-00003 TABLE 19 Effect of ASO treatment on ApoC III plasma protein levels in human ApoC III transgenic mice Dose ED.sub.50 Internucleoside SEQ (.mu.mol/ % (.mu.mol/ 3' Linkage/ ID ASO kg) PBS kg) Conjugate Length No. PBS 0 100 -- -- -- ISIS 0.08 86 0.73 None PS/20 2248 304801 0.75 51 2.25 23 6.75 13 ISIS 0.08 72 0.19 GalNAc.sub.3-1 PS/20 2249 647535 0.75 14 2.25 12 6.75 11
[0763] Plasma triglycerides and cholesterol were extracted by the method of Bligh and Dyer (Bligh, E. G. and Dyer, W. J. Can. J. Biochem. Physiol. 37: 911-917, 1959)(Bligh, E and Dyer, W, Can J Biochem Physiol, 37, 911-917, 1959)(Bligh, E and Dyer, W, Can J Biochem Physiol, 37, 911-917, 1959) and measured by using a Beckmann Coulter clinical analyzer and commercially available reagents.
[0764] The triglyceride levels were measured relative to PBS injected mice and are denoted as "% PBS". Results are presented in Table 20. As illustrated, both antisense compounds lowered triglyceride levels. Further, the antisense compound conjugated to GalNAc.sub.3-1 (ISIS 647535) was substantially more potent than the antisense compound lacking the GalNAc.sub.3-1 conjugate (ISIS 304801).
TABLE-US-00004 TABLE 20 Effect of ASO treatment on triglyceride levels in transgenic mice Dose ED.sub.50 Internucleoside SEQ (.mu.mol/ % (.mu.mol/ 3' Linkage/ ID ASO kg) PBS kg) Conjugate Length No. PBS 0 100 -- -- -- ISIS 0.08 87 0.63 None PS/20 2248 304801 0.75 46 2.25 21 6.75 12 ISIS 0.08 65 0.13 GalNAc.sub.3-1 PS/20 2249 647535 0.75 9 2.25 8 6.75 9
[0765] Plasma samples were analyzed by HPLC to determine the amount of total cholesterol and of different fractions of cholesterol (HDL and LDL). Results are presented in Tables 21 and 22. As illustrated, both antisense compounds lowered total cholesterol levels; both lowered LDL; and both raised HDL. Further, the antisense compound conjugated to GalNAc.sub.3-1 (ISIS 647535) was substantially more potent than the antisense compound lacking the GalNAc.sub.3-1 conjugate (ISIS 304801). An increase in HDL and a decrease in LDL levels is a cardiovascular beneficial effect of antisense inhibition of ApoC III.
TABLE-US-00005 TABLE 21 Effect of ASO treatment on total cholesterol levels in transgenic mice Total Internucleoside Dose Cholesterol 3' Linkage/ SEQ ASO (.mu.mol/kg) (mg/dL) Conjugate Length ID No. PBS 0 257 -- -- ISIS 0.08 226 None PS/20 2248 304801 0.75 164 2.25 110 6.75 82 ISIS 0.08 230 GalNAc.sub.3-1 PS/20 2249 647535 0.75 82 2.25 86 6.75 99
TABLE-US-00006 TABLE 22 Effect of ASO treatment on HDL and LDL cholesterol levels in transgenic mice Dose HDL LDL Internucleoside SEQ (.mu.mol/ (mg/ (mg/ 3' Linkage/ ID ASO kg) dL) dL) Conjugate Length No. PBS 0 17 28 -- -- ISIS 0.08 17 23 None PS/20 2248 304801 0.75 27 12 2.25 50 4 6.75 45 2 ISIS 0.08 21 21 GalNAc.sub.3-1 PS/20 2249 647535 0.75 44 2 2.25 50 2 6.75 58 2
Pharmacokinetics Analysis (PK)
[0766] The PK of the ASOs was also evaluated. Liver and kidney samples were minced and extracted using standard protocols. Samples were analyzed on MSD1 utilizing IP-HPLC-MS. The tissue level (.mu.g/g) of full-length ISIS 304801 and 647535 was measured and the results are provided in Table 23. As illustrated, liver concentrations of total full-length antisense compounds were similar for the two antisense compounds. Thus, even though the GalNAc.sub.3-1-conjugated antisense compound is more active in the liver (as demonstrated by the RNA and protein data above), it is not present at substantially higher concentration in the liver. Indeed, the calculated EC.sub.50 (provided in Table 23) confirms that the observed increase in potency of the conjugated compound cannot be entirely attributed to increased accumulation. This result suggests that the conjugate improved potency by a mechanism other than liver accumulation alone, possibly by improving the productive uptake of the antisense compound into cells.
[0767] The results also show that the concentration of GalNAc.sub.3-1 conjugated antisense compound in the kidney is lower than that of antisense compound lacking the GalNAc conjugate. This has several beneficial therapeutic implications. For therapeutic indications where activity in the kidney is not sought, exposure to kidney risks kidney toxicity without corresponding benefit. Moreover, high concentration in kidney typically results in loss of compound to the urine resulting in faster clearance. Accordingly, for non-kidney targets, kidney accumulation is undesired. These data suggest that GalNAc.sub.3-1 conjugation reduces kidney accumulation.
TABLE-US-00007 TABLE 23 PK analysis of ASO treatment in transgenic mice Inter- Dose Liver nucleoside SEQ (.mu.mol/ Liver Kidney EC.sub.50 3' Linkage/ ID ASO kg) (.mu.g/g) (.mu.g/g) (.mu.g/g) Conjugate Length No. ISIS 0.1 5.2 2.1 53 None PS/20 2248 304801 0.8 62.8 119.6 2.3 142.3 191.5 6.8 202.3 337.7 ISIS 0.1 3.8 0.7 3.8 GalNAc.sub.3-1 PS/20 2249 647535 0.8 72.7 34.3 2.3 106.8 111.4 6.8 237.2 179.3
[0768] Metabolites of ISIS 647535 were also identified and their masses were confirmed by high resolution mass spectrometry analysis. The cleavage sites and structures of the observed metabolites are shown below. The relative % of full length ASO was calculated using standard procedures and the results are presented in Table 23a. The major metabolite of ISIS 647535 was full-length ASO lacking the entire conjugate (i.e. ISIS 304801), which results from cleavage at cleavage site A, shown below. Further, additional metabolites resulting from other cleavage sites were also observed. These results suggest that introducing other cleabable bonds such as esters, peptides, disulfides, phosphoramidates or acyl-hydrazones between the GalNAc.sub.3-1 sugar and the ASO, which can be cleaved by enzymes inside the cell, or which may cleave in the reductive environment of the cytosol, or which are labile to the acidic pH inside endosomes and lyzosomes, can also be useful.
TABLE-US-00008 TABLE 23a Observed full length metabolites of ISIS 647535 Metabolite ASO Cleavage site Relative % 1 ISIS 304801 A 36.1 2 ISIS 304801 + dA B 10.5 3 ISIS 647535 minus [3 GalNAc] C 16.1 4 ISIS 647535 minus [3 GalNAc + 1 5-hydroxy-pentanoic acid tether] D 17.6 5 ISIS 647535 minus [2 GalNAc + 2 5-hydroxy-pentanoic acid tether] D 9.9 6 ISIS 647535 minus [3 GalNAc + 3 5-hvdroxy-pentanoic acid tether] D 9.8 ##STR00161## ##STR00162## ##STR00163## ##STR00164## ##STR00165## ##STR00166## ##STR00167##
Example 21: Antisense Inhibition of Human ApoC III in Human ApoC III Transgenic Mice in Single Administration Study
[0769] ISIS 304801, 647535 and 647536 each targeting human ApoC III and described in Table 17, were further evaluated in a single administration study for their ability to inhibit human ApoC III in human ApoC III transgenic mice.
Treatment
[0770] Human ApoCIII transgenic mice were maintained on a 12-hour light/dark cycle and fed ad libitum Teklad lab chow. Animals were acclimated for at least 7 days in the research facility before initiation of the experiment. ASOs were prepared in PBS and sterilized by filtering through a 0.2 micron filter. ASOs were dissolved in 0.9% PBS for injection.
[0771] Human ApoC III transgenic mice were injected intraperitoneally once at the dosage shown below with ISIS 304801, 647535 or 647536 (described above) or with PBS treated control. The treatment group consisted of 3 animals and the control group consisted of 4 animals. Prior to the treatment as well as after the last dose, blood was drawn from each mouse and plasma samples were analyzed. The mice were sacrificed 72 hours following the last administration.
[0772] Samples were collected and analyzed to determine the ApoC III mRNA and protein levels in the liver; plasma triglycerides; and cholesterol, including HDL and LDL fractions were assessed as described above (Example 20). Data from those analyses are presented in Tables 24-28, below. Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. The ALT and AST levels showed that the antisense compounds were well tolerated at all administered doses.
[0773] These results show improvement in potency for antisense compounds comprising a GalNAc.sub.3-1 conjugate at the 3' terminus (ISIS 647535 and 647536) compared to the antisense compound lacking a GalNAc.sub.3-1 conjugate (ISIS 304801). Further, ISIS 647536, which comprises a GalNAc.sub.3-1 conjugate and some phosphodiester linkages was as potent as ISIS 647535, which comprises the same conjugate and all internucleoside linkages within the ASO are phosphorothioate.
TABLE-US-00009 TABLE 24 Effect of ASO treatment on ApoC III mRNA levels in human ApoC III transgenic mice Dose Internucleoside SEQ (mg/ % ED.sub.50 3' linkage/ ID ASO kg) PBS (mg/kg) Conjugate Length No. PBS 0 99 -- -- -- ISIS 1 104 13.2 None PS/20 2248 304801 3 92 10 71 30 40 ISIS 0.3 98 1.9 GalNAc.sub.3-1 PS/20 2249 647535 1 70 3 33 10 20 ISIS 0.3 103 1.7 GalNAc.sub.3-1 PS/PO/20 2249 647536 1 60 3 31 10 21
TABLE-US-00010 TABLE 25 Effect of ASO treatment on ApoC III plasma protein levels in human ApoC III transgenic mice Dose Internucleoside SEQ (mg/ % ED.sub.50 3' Linkage/ ID ASO kg) PBS (mg/kg) Conjugate Length No. PBS 0 99 -- -- -- ISIS 1 104 23.2 None PS/20 2248 304801 3 92 10 71 30 40 ISIS 0.3 98 2.1 GalNAc.sub.3-1 PS/20 2249 647535 1 70 3 33 10 20 ISIS 0.3 103 1.8 GalNAc.sub.3-1 PS/PO/20 2249 647536 1 60 3 31 10 21
TABLE-US-00011 TABLE 26 Effect of ASO treatment on triglyceride levels in transgenic mice Dose Internucleoside SEQ (mg/ % ED.sub.50 3' Linkage/ ID ASO kg) PBS (mg/kg) Conjugate Length No. PBS 0 98 -- -- ISIS 1 80 29.1 None PS/20 2248 304801 3 92 10 70 30 47 ISIS 0.3 100 2.2 GalNAc.sub.3-1 PS/20 2249 647535 1 70 3 34 10 23 ISIS 0.3 95 1.9 GalNAc.sub.3-1 PS/PO/20 2249 647536 1 66 3 31 10 23
TABLE-US-00012 TABLE 27 Effect of ASO treatment on total cholesterol levels in transgenic mice Dose 3' Internucleoside SEQ ASO (mg/kg) % PBS Conjugate Linkage/Length ID No. PBS 0 96 -- -- ISIS 1 104 None PS/20 2248 304801 3 96 10 86 30 72 ISIS 0.3 93 GalNAc.sub.3-1 PS/20 2249 647535 1 85 3 61 10 53 ISIS 0.3 115 GalNAc.sub.3-1 PS/PO/20 2249 647536 1 79 3 51 10 54
TABLE-US-00013 TABLE 28 Effect of ASO treatment on HDL and LDL cholesterol levels in transgenic mice HDL LDL Internucleoside SEQ Dose % % 3' Linkage/ ID ASO (mg/kg) PBS PBS Conjugate Length No. PBS 0 131 90 -- -- ISIS 1 130 72 None PS/20 2248 304801 3 186 79 10 226 63 30 240 46 ISIS 0.3 98 86 GalNAc.sub.3-1 PS/20 2249 647535 1 214 67 3 212 39 10 218 35 ISIS 0.3 143 89 GalNAc.sub.3-1 PS/PO/20 2249 647536 1 187 56 3 213 33 10 221 34
[0774] These results confirm that the GalNAc.sub.3-1 conjugate improves potency of an antisense compound. The results also show equal potency of a GalNAc.sub.3-1 conjugated antisense compounds where the antisense oligonucleotides have mixed linkages (ISIS 647536 which has six phosphodiester linkages) and a full phosphorothioate version of the same antisense compound (ISIS 647535).
[0775] Phosphorothioate linkages provide several properties to antisense compounds. For example, they resist nuclease digestion and they bind proteins resulting in accumulation of compound in the liver, rather than in the kidney/urine. These are desirable properties, particularly when treating an indication in the liver. However, phosphorothioate linkages have also been associated with an inflammatory response. Accordingly, reducing the number of phosphorothioate linkages in a compound is expected to reduce the risk of inflammation, but also lower concentration of the compound in liver, increase concentration in the kidney and urine, decrease stability in the presence of nucleases, and lower overall potency. The present results show that a GalNAc.sub.3-1 conjugated antisense compound where certain phosphorothioate linkages have been replaced with phosphodiester linkages is as potent against a target in the liver as a counterpart having full phosphorothioate linkages. Such compounds are expected to be less proinflammatory (See Example 24 describing an experiment showing reduction of PS results in reduced inflammatory effect).
Example 22: Effect of GalNAc.sub.3-1 Conjugated Modified ASO Targeting SRB-1 In Vivo
[0776] ISIS 440762 and 651900, each targeting SRB-1 and described in Table 17, were evaluated in a dose-dependent study for their ability to inhibit SRB-1 in Balb/c mice.
Treatment
[0777] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 440762, 651900 or with PBS treated control. Each treatment group consisted of 4 animals. The mice were sacrificed 48 hours following the final administration to determine the SRB-1 mRNA levels in liver using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. SRB-1 mRNA levels were determined relative to total RNA (using Ribogreen), prior to normalization to PBS-treated control. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to PBS-treated control and is denoted as "% PBS".
[0778] As illustrated in Table 29, both antisense compounds lowered SRB-1 mRNA levels. Further, the antisense compound comprising the GalNAc.sub.3-1 conjugate (ISIS 651900) was substantially more potent than the antisense compound lacking the GalNAc.sub.3-1 conjugate (ISIS 440762). These results demonstrate that the potency benefit of GalNAc.sub.3-1 conjugates are observed using antisense oligonucleotides complementary to a different target and having different chemically modified nucleosides, in this instance modified nucleosides comprise constrained ethyl sugar moieties (a bicyclic sugar moiety).
TABLE-US-00014 TABLE 29 Effect of ASO treatment on SRB-1 mRNA levels in Balb/c mice Internucleoside SEQ Dose Liver ED.sub.50 3' linkage/ ID ASO (mg/kg) % PBS (mg/kg) Conjugate Length No. PBS 0 100 -- -- ISIS 0.7 85 2.2 None PS/14 2250 440762 2 55 7 12 20 3 ISIS 0.07 98 0.3 GalNAc.sub.3-1 PS/14 2251 651900 0.2 63 0.7 20 2 6 7 5
Example 23: Human Peripheral Blood Mononuclear Cells (hPBMC) Assay Protocol
[0779] The hPBMC assay was performed using BD Vautainer CPT tube method. A sample of whole blood from volunteered donors with informed consent at US HealthWorks clinic (Faraday & El Camino Real, Carlsbad) was obtained and collected in 4-15 BD Vacutainer CPT 8 ml tubes (VWR Cat. # BD362753). The approximate starting total whole blood volume in the CPT tubes for each donor was recorded using the PBMC assay data sheet.
[0780] The blood sample was remixed immediately prior to centrifugation by gently inverting tubes 8-10 times. CPT tubes were centrifuged at rt (18-25.degree. C.) in a horizontal (swing-out) rotor for 30 min. at 1500-1800 RCF with brake off (2700 RPM Beckman Allegra 6R). The cells were retrieved from the buffy coat interface (between Ficoll and polymer gel layers); transferred to a sterile 50 ml conical tube and pooled up to 5 CPT tubes/50 ml conical tube/donor. The cells were then washed twice with PBS (Ca.sup.++, Mg.sup.++ free; GIBCO). The tubes were topped up to 50 ml and mixed by inverting several times. The sample was then centrifuged at 330.times.g for 15 minutes at rt (1215 RPM in Beckman Allegra 6R) and aspirated as much supernatant as possible without disturbing pellet. The cell pellet was dislodged by gently swirling tube and resuspended cells in RPMI+10% FBS+pen/strep (.about.1 ml/10 ml starting whole blood volume). A 60 .mu.l sample was pipette into a sample vial (Beckman Coulter) with 600 .mu.l VersaLyse reagent (Beckman Coulter Cat # A09777) and was gently vortexed for 10-15 sec. The sample was allowed to incubate for 10 min. at rt and being mixed again before counting. The cell suspension was counted on Vicell XR cell viability analyzer (Beckman Coulter) using PBMC cell type (dilution factor of 1:11 was stored with other parameters). The live cell/ml and viability were recorded. The cell suspension was diluted to 1.times.10.sup.7 live PBMC/ml in RPMI+10% FBS+pen/strep. The cells were plated at 5.times.10.sup.5 in 50 l/well of 96-well tissue culture plate (Falcon Microtest). 50 l/well of 2.times. concentration oligos/controls diluted in RPMI+10% FBS+pen/strep. was added according to experiment template (100 .mu.l/well total). Plates were placed on the shaker and allowed to mix for approx. 1 min. After being incubated for 24 hrs at 37.degree. C.; 5% CO.sub.2, the plates were centrifuged at 400.times.g for 10 minutes before removing the supernatant for MSD cytokine assay (i.e. human IL-6, IL-10, IL-8 and MCP-1).
Example 24: Evaluation of Proinflammatory Effects in hPBMC Assay for GalNAc.sub.3-1 Conjugated ASOs
[0781] The antisense oligonucleotides (ASOs) listed in Table 30 were evaluated for proinflammatory effect in hPBMC assay using the protocol described in Example 23. ISIS 353512 is an internal standard known to be a high responder for IL-6 release in the assay. The hPBMCs were isolated from fresh, volunteered donors and were treated with ASOs at 0, 0.0128, 0.064, 0.32, 1.6, 8, 40 and 200 M concentrations. After a 24 hr treatment, the cytokine levels were measured.
[0782] The levels of IL-6 were used as the primary readout. The EC.sub.50 and E.sub.max was calculated using standard procedures. Results are expressed as the average ratio of E.sub.max/EC.sub.50 from two donors and is denoted as "E.sub.max/EC.sub.50." The lower ratio indicates a relative decrease in the proinflammatory response and the higher ratio indicates a relative increase in the proinflammatory response.
[0783] With regard to the test compounds, the least proinflammatory compound was the PS/PO linked ASO (ISIS 616468). The GalNAc.sub.3-1 conjugated ASO, ISIS 647535 was slightly less proinflammatory than its non-conjugated counterpart ISIS 304801. These results indicate that incorporation of some PO linkages reduces proinflammatory reaction and addition of a GalNAc.sub.3-1 conjugate does not make a compound more proinflammatory and may reduce proinflammatory response. Accordingly, one would expect that an antisense compound comprising both mixed PS/PO linkages and a GalNAc.sub.3-1 conjugate would produce lower proinflammatory responses relative to full PS linked antisense compound with or without a GalNAc.sub.3-1 conjugate. These results show that GalNAc.sub.3_1 conjugated antisense compounds, particularly those having reduced PS content are less proinflammatory.
[0784] Together, these results suggest that a GalNAc.sub.3-1 conjugated compound, particularly one with reduced PS content, can be administered at a higher dose than a counterpart full PS antisense compound lacking a GalNAc.sub.3-1 conjugate. Since half-life is not expected to be substantially different for these compounds, such higher administration would result in less frequent dosing. Indeed such administration could be even less frequent, because the GalNAc.sub.3-1 conjugated compounds are more potent (See Examples 20-22) and re-dosing is necessary once the concentration of a compound has dropped below a desired level, where such desired level is based on potency.
TABLE-US-00015 TABLE 30 Modified ASOs SEQ ID ASO Sequence (5' to 3') Target No. ISIS G.sub.es.sup.mC.sub.esT.sub.esG.sub.esA.sub.esT.sub.dsT.sub.dsA.sub.d- sG.sub.dsA.sub.dsG.sub.ds TNF.alpha. 2252 104838 A.sub.dsG.sub.dsA.sub.dsG.sub.dsG.sub.esT.sub.es.sup.mC.sub.es.sup.- mC.sub.es.sup.mC.sub.e ISIS T.sub.es.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.dsA.sub.dsT.sub.dsT.s- ub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds CRP 2253 353512 G.sub.dsA.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsT.sub.e- sG.sub.esG.sub.e ISIS A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds ApoC III 2248 304801 .sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT- .sub.esT.sub.esA.sub.esT.sub.e ISIS A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds ApoC III 2249 647535 .sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT- .sub.esT.sub.esA.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a ISIS A.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds ApoC III 2248 616468 .sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.eoT- .sub.eoT.sub.esA.sub.esT.sub.e Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "k" indicates 6'-(S)-CH.sub.3 bicyclic nucleoside (e.g. cEt); "s" indicates phosphorothioate internucleoside linkages (PS); "o" indicates phosphodiester internucleoside linkages PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Superscript "m" indicates 5-methylcytosines. "A.sub.do'-GalNAc.sub.3-1.sub.a" indicats a conjugate having the structure GalNAc.sub.3-1 shown in Example 9 attached to the 3'-end of the antisense olignucleotide, as indicated
TABLE-US-00016 TABLE 31 Proinflammatory Effect of ASOs targeting ApoC III in hPBMC assay Inter- nucleoside SEQ EC.sub.50 E.sub.max E.sub.max/ 3' Linkage/ ID ASO .mu.M) (.mu.M) EC.sub.50 Conjugate Length No. ISIS 353512 0.01 265.9 26,590 None PS/20 2253 (high responder) ISIS 304801 0.07 106.55 1,522 None PS/20 2248 ISIS 647535 0.12 138 1,150 GalNAc.sub.3-1 PS/20 2249 ISIS 616468 0.32 71.52 224 None PS/PO/20 2248
Example 25: Effect of GalNAc.sub.3-1 Conjugated Modified ASO Targeting Human ApoC III In Vitro
[0785] ISIS 304801 and 647535 described above were tested in vitro. Primary hepatocyte cells from transgenic mice at a density of 25,000 cells per well were treated with 0.03, 0.08, 0.24, 0.74, 2.22, 6.67 and 20 M concentrations of modified oligonucleotides. After a treatment period of approximately 16 hours, RNA was isolated from the cells and mRNA levels were measured by quantitative real-time PCR and the hApoC III mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.
[0786] The IC.sub.50 was calculated using the standard methods and the results are presented in Table 32. As illustrated, comparable potency was observed in cells treated with ISIS 647535 as compared to the control, ISIS 304801.
TABLE-US-00017 TABLE 32 Modified ASO targeting human ApoC III in primary hepatocytes Intenucleoside SEQ ASO IC.sub.50 (.mu.M) 3' Conjugate linkage/Length ID No. ISIS 0.44 None PS/20 2248 304801 ISIS 0.31 GalNAc.sub.3-1 PS/20 2249 647535
[0787] In this experiment, the large potency benefits of GalNAc.sub.3-1 conjugation that are observed in vivo were not observed in vitro. Subsequent free uptake experiments in primary hepatocytes in vitro did show increased potency of oligonucleotides comprising various GalNAc conjugates relative to oligonucleotides that lacking the GalNAc conjugate. (see Examples 60, 82, and 92)
Example 26: Effect of PO/PS Linkages on ApoC III ASO Activity
[0788] Human ApoC III transgenic mice were injected intraperitoneally once at 25 mg/kg of ISIS 304801, or ISIS 616468 (both described above) or with PBS treated control once per week for two weeks. The treatment group consisted of 3 animals and the control group consisted of 4 animals. Prior to the treatment as well as after the last dose, blood was drawn from each mouse and plasma samples were analyzed. The mice were sacrificed 72 hours following the last administration.
[0789] Samples were collected and analyzed to determine the ApoC III protein levels in the liver as described above (Example 20). Data from those analyses are presented in Table 33, below.
[0790] These results show reduction in potency for antisense compounds with PO/PS (ISIS 616468) in the wings relative to full PS (ISIS 304801).
TABLE-US-00018 TABLE 33 Effect of ASO treatment on ApoC III protein levels in human ApoC III transgenic mice Dose 3' Internucleoside SEQ ID ASO (mg/kg) % PBS Conjugate linkage/Length No. PBS 0 99 -- -- ISIS 25 24 None Full PS 2248 304801 mg/kg/wk for 2 wks ISIS 25 40 None 14 PS/6 PO 2248 616468 mg/kg/wk for 2 wks
Example 27: Compound 56
##STR00168##
[0792] Compound 56 is commercially available from Glen Research or may be prepared according to published procedures reported by Shchepinov et al., Nucleic Acids Research, 1997, 25(22), 4447-4454.
Example 28: Preparation of Compound 60
##STR00169##
[0794] Compound 4 was prepared as per the procedures illustrated in Example 2. Compound 57 is commercially available. Compound 60 was confirmed by structural analysis.
[0795] Compound 57 is meant to be representative and not intended to be limiting as other monoprotected substituted or unsubstituted alkyl diols including but not limited to those presented in the specification herein can be used to prepare phosphoramidites having a predetermined composition.
Example 29: Preparation of Compound 63
##STR00170##
[0797] Compounds 61 and 62 are prepared using procedures similar to those reported by Tober et al., Eur. J Org. Chem., 2013, 3, 566-577; and Jiang et al., Tetrahedron, 2007, 63(19), 3982-3988.
[0798] Alternatively, Compound 63 is prepared using procedures similar to those reported in scientific and patent literature by Kim et al., Synlett, 2003, 12, 1838-1840; and Kim et al., published PCT International Application, WO 2004063208.
Example 30: Preparation of Compound 63b
##STR00171##
[0800] Compound 63a is prepared using procedures similar to those reported by Hanessian et al., Canadian Journal of Chemistry, 1996, 74(9), 1731-1737.
Example 31: Preparation of Compound 63d
##STR00172##
[0802] Compound 63c is prepared using procedures similar to those reported by Chen et al., Chinese Chemical Letters, 1998, 9(5), 451-453.
Example 32: Preparation of Compound 67
##STR00173##
[0804] Compound 64 was prepared as per the procedures illustrated in Example 2. Compound 65 is prepared using procedures similar to those reported by Or et al., published PCT International Application, WO 2009003009. The protecting groups used for Compound 65 are meant to be representative and not intended to be limiting as other protecting groups including but not limited to those presented in the specification herein can be used.
Example 33: Preparation of Compound 70
##STR00174##
[0806] Compound 64 was prepared as per the procedures illustrated in Example 2. Compound 68 is commercially available. The protecting group used for Compound 68 is meant to be representative and not intended to be limiting as other protecting groups including but not limited to those presented in the specification herein can be used.
Example 34: Preparation of Compound 75a
##STR00175##
[0808] Compound 75 is prepared according to published procedures reported by Shchepinov et al., Nucleic Acids Research, 1997, 25(22), 4447-4454.
Example 35: Preparation of Compound 79
##STR00176##
[0810] Compound 76 was prepared according to published procedures reported by Shchepinov et al., Nucleic Acids Research, 1997, 25(22), 4447-4454.
Example 36: Preparation of Compound 79a
##STR00177##
[0812] Compound 77 is prepared as per the procedures illustrated in Example 35.
Example 37: General Method for the Preparation of Conjugated Oligomeric Compound 82 Comprising a Phosphodiester Linked GalNAc.sub.3-2 Conjugate at 5' Terminus Via Solid Support (Method I)
##STR00178##
[0813] wherein GalNAc.sub.3-2 has the structure:
##STR00179##
[0814] The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-2 (GalNAc.sub.3-2.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. Wherein GalNAc.sub.3-2.sub.a has the formula:
##STR00180##
[0815] The VIMAD-bound oligomeric compound 79b was prepared using standard procedures for automated DNA/RNA synthesis (see Dupouy et al., Angew. Chem. Int. Ed., 2006, 45, 3623-3627). The phosphoramidite Compounds 56 and 60 were prepared as per the procedures illustrated in Examples 27 and 28, respectively. The phosphoramidites illustrated are meant to be representative and not intended to be limiting as other phosphoramidite building blocks including but not limited those presented in the specification herein can be used to prepare an oligomeric compound having a phosphodiester linked conjugate group at the 5' terminus. The order and quantity of phosphoramidites added to the solid support can be adjusted to prepare the oligomeric compounds as described herein having any predetermined sequence and composition.
Example 38: Alternative Method for the Preparation of Oligomeric Compound 82 Comprising a Phosphodiester Linked GalNAc.sub.3-2 Conjugate at 5' Terminus (Method II)
##STR00181##
[0817] The VIMAD-bound oligomeric compound 79b was prepared using standard procedures for automated DNA/RNA synthesis (see Dupouy et al., Angew. Chem. Int. Ed., 2006, 45, 3623-3627). The GalNAc.sub.3-2 cluster phosphoramidite, Compound 79 was prepared as per the procedures illustrated in Example 35. This alternative method allows a one-step installation of the phosphodiester linked GalNAc.sub.3-2 conjugate to the oligomeric compound at the final step of the synthesis. The phosphoramidites illustrated are meant to be representative and not intended to be limiting, as other phosphoramidite building blocks including but not limited to those presented in the specification herein can be used to prepare oligomeric compounds having a phosphodiester conjugate at the 5' terminus. The order and quantity of phosphoramidites added to the solid support can be adjusted to prepare the oligomeric compounds as described herein having any predetermined sequence and composition.
Example 39: General Method for the Preparation of Oligomeric Compound 83h Comprising a GalNAc.sub.3-3 Conjugate at the 5' Terminus (GalNAc.sub.3-1 Modified for 5' End Attachment) Via Solid Support
##STR00182## ##STR00183## ##STR00184##
[0819] Compound 18 was prepared as per the procedures illustrated in Example 4. Compounds 83a and 83b are commercially available. Oligomeric Compound 83e comprising a phosphodiester linked hexylamine was prepared using standard oligonucleotide synthesis procedures. Treatment of the protected oligomeric compound with aqueous ammonia provided the 5'-GalNAc.sub.3-3 conjugated oligomeric compound (83h). Wherein GalNAc.sub.3-3 has the structure:
##STR00185##
[0820] The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-3 (GalNAc.sub.3-3.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. Wherein GalNAc.sub.3-3.sub.a has the formula:
##STR00186##
Example 40: General Method for the Preparation of Oligomeric Compound 89 Comprising a Phosphodiester Linked GalNAc.sub.3-4 Conjugate at the 3' Terminus Via Solid Support
##STR00187## ##STR00188##
[0821] Wherein GalNAc.sub.3-4 has the structure:
##STR00189##
[0822] Wherein CM is a cleavable moiety. In certain embodiments, cleavable moiety is:
##STR00190##
[0823] The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-4 (GalNAc.sub.3-4.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. Wherein GalNAc.sub.3-4.sub.a has the formula:
##STR00191##
[0824] The protected Unylinker functionalized solid support Compound 30 is commercially available. Compound 84 is prepared using procedures similar to those reported in the literature (see Shchepinov et al., Nucleic Acids Research, 1997, 25(22), 4447-4454; Shchepinov et al., Nucleic Acids Research, 1999, 27, 3035-3041; and Hornet et al., Nucleic Acids Research, 1997, 25, 4842-4849).
[0825] The phosphoramidite building blocks, Compounds 60 and 79a are prepared as per the procedures illustrated in Examples 28 and 36. The phosphoramidites illustrated are meant to be representative and not intended to be limiting as other phosphoramidite building blocks can be used to prepare an oligomeric compound having a phosphodiester linked conjugate at the 3' terminus with a predetermined sequence and composition. The order and quantity of phosphoramidites added to the solid support can be adjusted to prepare the oligomeric compounds as described herein having any predetermined sequence and composition.
Example 41: General Method for the Preparation of ASOs Comprising a Phosphodiester Linked GalNAc.sub.3-2 (See Example 37, Bx is Adenine) Conjugate at the 5' Position Via Solid Phase Techniques (Preparation of ISIS 661134)
[0826] Unless otherwise stated, all reagents and solutions used for the synthesis of oligomeric compounds are purchased from commercial sources. Standard phosphoramidite building blocks and solid support are used for incorporation nucleoside residues which include for example T, A, G, and .sup.mC residues. Phosphoramidite compounds 56 and 60 were used to synthesize the phosphodiester linked GalNAc.sub.3-2 conjugate at the 5' terminus. A 0.1 M solution of phosphoramidite in anhydrous acetonitrile was used for .beta.-D-2'-deoxyribonucleoside and 2'-MOE.
[0827] The ASO syntheses were performed on ABI 394 synthesizer (1-2 .mu.mol scale) or on GE Healthcare Bioscience KTA oligopilot synthesizer (40-200 .mu.mol scale) by the phosphoramidite coupling method on VIMAD solid support (110 .mu.mol/g, Guzaev et al., 2003) packed in the column. For the coupling step, the phosphoramidites were delivered at a 4 fold excess over the initial loading of the solid support and phosphoramidite coupling was carried out for 10 min. All other steps followed standard protocols supplied by the manufacturer. A solution of 6% dichloroacetic acid in toluene was used for removing the dimethoxytrityl (DMT) groups from 5'-hydroxyl groups of the nucleotide. 4,5-Dicyanoimidazole (0.7 M) in anhydrous CH.sub.3CN was used as activator during the coupling step. Phosphorothioate linkages were introduced by sulfurization with 0.1 M solution of xanthane hydride in 1:1 pyridine/CH.sub.3CN for a contact time of 3 minutes. A solution of 20% tert-butylhydroperoxide in CH.sub.3CN containing 6% water was used as an oxidizing agent to provide phosphodiester internucleoside linkages with a contact time of 12 minutes.
[0828] After the desired sequence was assembled, the cyanoethyl phosphate protecting groups were deprotected using a 20% diethylamine in toluene (v/v) with a contact time of 45 minutes. The solid-support bound ASOs were suspended in aqueous ammonia (28-30 wt %) and heated at 55.degree. C. for 6 h.
The unbound ASOs were then filtered and the ammonia was boiled off. The residue was purified by high pressure liquid chromatography on a strong anion exchange column (GE Healthcare Bioscience, Source 30Q, 30 .mu.m, 2.54.times.8 cm, A=100 mM ammonium acetate in 30% aqueous CH.sub.3CN, B=1.5 M NaBr in A, 0-40% of B in 60 min, flow 14 mL min-1, X=260 nm). The residue was desalted by HPLC on a reverse phase column to yield the desired ASOs in an isolated yield of 15-30% based on the initial loading on the solid support. The ASOs were characterized by ion-pair-HPLC coupled MS analysis with Agilent 1100 MSD system.
TABLE-US-00019 TABLE 34 ASO comprising a phosphodiester linked GalNAc.sub.3-2 conjugate at the 5' position targeting SRB-1 Observed SEQ ID ISIS No. Sequence (5' to 3') CalCd Mass Mass No. 661134 GalNAc.sub.3-2.sub.a-.sub.o'A.sub.doT.sub.ks.sup.mC.sub.ksA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.ds 6482.2 6481.6 2254 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "k" indicates 6'-(S)-CH.sub.3 bicyclic nucleoside (e.g. cEt); "s" indicates phosphorothioate internucleoside linkages (PS); "o" indicates phosphodiester internucleoside linkages PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Superscript "m" indicates 5-methylcytosines. The structure of GalNAc.sub.3-2.sub.a is shown in Example 37.
Example 42: General Method for the Preparation of ASOs Comprising a GalNAc.sub.3-3 Conjugate at the 5' Position Via Solid Phase Techniques (Preparation of ISIS 661166)
[0829] The synthesis for ISIS 661166 was performed using similar procedures as illustrated in Examples 39 and 41.
[0830] ISIS 661166 is a 5-10-5 MOE gapmer, wherein the 5' position comprises a GalNAc.sub.3-3 conjugate. The ASO was characterized by ion-pair-HPLC coupled MS analysis with Agilent 1100 MSD system.
TABLE-US-00020 TABLE 34a ASO comprising a GalNAc.sub.3-3 conjugate at the 5' position via a hexylamino phosphodiester linkage targeting Malat-1 ISIS Calcd Observed No. Sequence (5' to 3') Conjugate Mass Mass SEQ ID No. 661166 5'-GalNAc.sub.3-3.sub.a-o'.sup.mC.sub.esG.sub.esG.sub.esT.sub.esG.s- ub.es 5'-GalNAc.sub.3-3 8992.16 8990.51 2255 .sup.mC.sub.dsA.sub.dsA.sub.dsG.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .dsA.sub.dsG.sub.ds G.sub.esA.sub.esA.sub.esT.sub.esT.sub.e Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "s" indicates phosphorothioate internucleoside linkages (PS); "o" indicates phosphodiester internucleoside linkages (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Superscript "m" indicates 5-methylcytosines. The structureof "5'-GalNAc.sub.3-3a" is shown in Example 39.
Example 43: Dose-Dependent Study of Phosphodiester Linked GalNAc.sub.3-2 (See Examples 37 and 41, Bx is Adenine) at the 5' Terminus Targeting SRB-1 In Vivo
[0831] ISIS 661134 (see Example 41) comprising a phosphodiester linked GalNAc.sub.3-2 conjugate at the 5' terminus was tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Unconjugated ISIS 440762 and 651900 (GalNAc.sub.3-1 conjugate at 3' terminus, see Example 9) were included in the study for comparison and are described previously in Table 17.
Treatment
[0832] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 440762, 651900, 661134 or with PBS treated control. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. SRB-1 mRNA levels were determined relative to total RNA (using Ribogreen), prior to normalization to PBS-treated control. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to PBS-treated control and is denoted as "% PBS". The ED.sub.50s were measured using similar methods as described previously and are presented below.
[0833] As illustrated in Table 35, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. Indeed, the antisense oligonucleotides comprising the phosphodiester linked GalNAc.sub.3-2 conjugate at the 5' terminus (ISIS 661134) or the GalNAc.sub.3-1 conjugate linked at the 3' terminus (ISIS 651900) showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 440762). Further, ISIS 661134, which comprises the phosphodiester linked GalNAc.sub.3-2 conjugate at the 5' terminus was equipotent compared to ISIS 651900, which comprises the GalNAc.sub.3-1 conjugate at the 3' terminus.
TABLE-US-00021 TABLE 35 ASOs containing GalNAc.sub.3-1 or GalNAc.sub.3-2 targeting SRB-1 ISIS Dosage SRB-1 mRNA ED.sub.50 SEQ No. (mg/kg) levels (% PBS) (mg/kg) Conjugate ID No. PBS 0 100 -- -- 440762 0.2 116 2.58 No conjugate 2250 0.7 91 2 69 7 22 20 5 651900 0.07 95 0.26 3' GalNAc.sub.3-1 2251 0.2 77 0.7 28 2 11 7 8 661134 0.07 107 0.25 5' GalNAc.sub.3-2 2254 0.2 86 0.7 28 2 10 7 6 Structures for 3' GalNAc.sub.3-1 and 5' GalNAc.sub.3-2 were described previously in Examples 9 and 37.
Pharmacokinetics Analysis (PK)
[0834] The PK of the ASOs from the high dose group (7 mg/kg) was examined and evaluated in the same manner as illustrated in Example 20. Liver sample was minced and extracted using standard protocols. The full length metabolites of 661134 (5' GalNAc.sub.3-2) and ISIS 651900 (3' GalNAc.sub.3-1) were identified and their masses were confirmed by high resolution mass spectrometry analysis. The results showed that the major metabolite detected for the ASO comprising a phosphodiester linked GalNAc.sub.3-2 conjugate at the 5' terminus (ISIS 661134) was ISIS 440762 (data not shown). No additional metabolites, at a detectable level, were observed. Unlike its counterpart, additional metabolites similar to those reported previously in Table 23a were observed for the ASO having the GalNAc.sub.3-1 conjugate at the 3' terminus (ISIS 651900). These results suggest that having the phosphodiester linked GalNAc.sub.3-1 or GalNAc.sub.3-2 conjugate may improve the PK profile of ASOs without compromising their potency.
Example 44: Effect of PO/PS Linkages on Antisense Inhibition of ASOs Comprising GalNAc.sub.3-1 Conjugate (See Example 9) at the 3' Terminus Targeting SRB-1
[0835] ISIS 655861 and 655862 comprising a GalNAc.sub.3-1 conjugate at the 3' terminus each targeting SRB-1 were tested in a single administration study for their ability to inhibit SRB-1 in mice. The parent unconjugated compound, ISIS 353382 was included in the study for comparison.
[0836] The ASOs are 5-10-5 MOE gapmers, wherein the gap region comprises ten 2'-deoxyribonucleosides and each wing region comprises five 2'-MOE modified nucleosides. The ASOs were prepared using similar methods as illustrated previously in Example 19 and are described Table 36, below.
TABLE-US-00022 TABLE 36 Modified ASOs comprising GalNAc.sub.3-1 conjugate at the 3' terminus targeting SRB-1 SEQ ID ISIS No. Sequence (5' to 3') Chemistry No. 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds Full PS no conjugate 2256 (parent) .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.e- sT.sub.e 655861 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds Full PS with 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1.sub.a GalNAc.sub.3-1 conjugate 655862 G.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.eoA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds Mixed PS/PO with 2257 .sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1.sub.a GalNAc.sub.3-1 conjugate Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "s" indicates phosphorothioate internucleoside linkages (PS); "o" indicates phosphodiester internucleoside linkages (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Superscript "m" indicates 5-methylcytosines. The structure of "GalNAc.sub.3-1" is shown in Example 9.
Treatment
[0837] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 353382, 655861, 655862 or with PBS treated control. Each treatment group consisted of 4 animals. Prior to the treatment as well as after the last dose, blood was drawn from each mouse and plasma samples were analyzed. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. SRB-1 mRNA levels were determined relative to total RNA (using Ribogreen), prior to normalization to PBS-treated control. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to PBS-treated control and is denoted as "% PBS". The ED.sub.50s were measured using similar methods as described previously and are reported below.
[0838] As illustrated in Table 37, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner compared to PBS treated control. Indeed, the antisense oligonucleotides comprising the GalNAc.sub.3-1 conjugate at the 3' terminus (ISIS 655861 and 655862) showed substantial improvement in potency comparing to the unconjugated antisense oligonucleotide (ISIS 353382). Further, ISIS 655862 with mixed PS/PO linkages showed an improvement in potency relative to full PS (ISIS 655861).
TABLE-US-00023 TABLE 37 Effect of PO/PS linkages on antisense inhibition of ASOs comprising GalNAc.sub.3-1 conjugate at 3' terminus targeting SRB-1 ISIS Dosage SRB-1 mRNA ED.sub.50 SEQ No. (mg/kg) levels (% PBS) (mg/kg) Chemistry ID No. PBS 0 100 -- -- 353382 3 76.65 10.4 Full PS 2256 (parent) 10 52.40 without 30 24.95 conjugate 655861 0.5 81.22 2.2 Full PS 2257 1.5 63.51 with 5 24.61 GalNAc.sub.3-1 15 14.80 conjugate 655862 0.5 69.57 1.3 Mixed 2257 1.5 45.78 PS/PO with 5 19.70 GalNAc.sub.3-1 15 12.90 conjugate
[0839] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Organ weights were also evaluated. The results demonstrated that no elevation in transaminase levels (Table 38) or organ weights (data not shown) were observed in mice treated with ASOs compared to PBS control. Further, the ASO with mixed PS/PO linkages (ISIS 655862) showed similar transaminase levels compared to full PS (ISIS 655861).
TABLE-US-00024 TABLE 38 Effect of PO/PS linkages on transaminase levels of ASOs comprising GalNAc.sub.3-1 conjugate at 3' terminus targeting SRB-1 ISIS Dosage ALT AST SEQ No. (mg/kg) (U/L) (U/L) Chemistry ID No. PBS 0 28.5 65 -- 353382 3 50.25 89 Full PS without 2256 (parent) 10 27.5 79.3 conjugate 30 27.3 97 655861 0.5 28 55.7 Full PS with 2257 1.5 30 78 GalNAc.sub.3-1 5 29 63.5 15 28.8 67.8 655862 0.5 50 75.5 Mixed PS/PO with 2257 1.5 21.7 58.5 GalNAc.sub.3-1 5 29.3 69 15 22 61
Example 45: Preparation of PFP Ester, Compound 110a
##STR00192## ##STR00193##
[0841] Compound 4 (9.5 g, 28.8 mmoles) was treated with compound 103a or 103b (38 mmoles), individually, and TMSOTf (0.5 eq.) and molecular sieves in dichloromethane (200 mL), and stirred for 16 hours at room temperature. At that time, the organic layer was filtered thru celite, then washed with sodium bicarbonate, water and brine. The organic layer was then separated and dried over sodium sulfate, filtered and reduced under reduced pressure. The resultant oil was purified by silica gel chromatography (2%-->10% methanol/dichloromethane) to give compounds 104a and 104b in >80% yield. LCMS and proton NMR was consistent with the structure.
[0842] Compounds 104a and 104b were treated to the same conditions as for compounds 100a-d (Example 47), to give compounds 105a and 105b in >90% yield. LCMS and proton NMR was consistent with the structure.
[0843] Compounds 105a and 105b were treated, individually, with compound 90 under the same conditions as for compounds 901a-d, to give compounds 106a (80%) and 106b (20%). LCMS and proton NMR was consistent with the structure.
[0844] Compounds 106a and 106b were treated to the same conditions as for compounds 96a-d (Example 47), to give 107a (60%) and 107b (20%). LCMS and proton NMR was consistent with the structure. Compounds 107a and 107b were treated to the same conditions as for compounds 97a-d (Example 47), to give compounds 108a and 108b in 40-60% yield. LCMS and proton NMR was consistent with the structure. Compounds 108a (60%) and 108b (40%) were treated to the same conditions as for compounds 100a-d (Example 47), to give compounds 109a and 109b in >80% yields. LCMS and proton NMR was consistent with the structure.
[0845] Compound 109a was treated to the same conditions as for compounds 101a-d (Example 47), to give Compound 110a in 30-60% yield. LCMS and proton NMR was consistent with the structure. Alternatively, Compound 110b can be prepared in a similar manner starting with Compound 109b.
Example 46: General Procedure for Conjugation with PFP Esters (Oligonucleotide 111); Preparation of ISIS 666881 (GalNAc.sub.3-10)
[0846] A 5'-hexylamino modified oligonucleotide was synthesized and purified using standard solid-phase oligonucleotide procedures. The 5'-hexylamino modified oligonucleotide was dissolved in 0.1 M sodium tetraborate, pH 8.5 (200 .mu.L) and 3 equivalents of a selected PFP esterified GalNAc.sub.3 cluster dissolved in DMSO (50 .mu.L) was added. If the PFP ester precipitated upon addition to the ASO solution DMSO was added until all PFP ester was in solution. The reaction was complete after about 16 h of mixing at room temperature. The resulting solution was diluted with water to 12 mL and then spun down at 3000 rpm in a spin filter with a mass cut off of 3000 Da. This process was repeated twice to remove small molecule impurities. The solution was then lyophilized to dryness and redissolved in concentrated aqueous ammonia and mixed at room temperature for 2.5 h followed by concentration in vacuo to remove most of the ammonia. The conjugated oligonucleotide was purified and desalted by RP-HPLC and lyophilized to provide the GalNAc.sub.3 conjugated oligonucleotide.
##STR00194##
[0847] Oligonucleotide 111 is conjugated with GalNAc.sub.3-10. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-10 (GalNAc.sub.3-10.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)-- as shown in the oligonucleotide (ISIS 666881) synthesized with GalNAc.sub.3-10 below. The structure of GalNAc.sub.3-10 (GalNAc.sub.3-10.sub.a-CM-) is shown below:
##STR00195##
[0848] Following this general procedure ISIS 666881 was prepared. 5'-hexylamino modified oligonucleotide, ISIS 660254, was synthesized and purified using standard solid-phase oligonucleotide procedures. ISIS 660254 (40 mg, 5.2 .mu.mol) was dissolved in 0.1 M sodium tetraborate, pH 8.5 (200 .mu.L) and 3 equivalents PFP ester (Compound 110a) dissolved in DMSO (50 .mu.L) was added. The PFP ester precipitated upon addition to the ASO solution requiring additional DMSO (600 .mu.L) to fully dissolve the PFP ester. The reaction was complete after 16 h of mixing at room temperature. The solution was diluted with water to 12 mL total volume and spun down at 3000 rpm in a spin filter with a mass cut off of 3000 Da. This process was repeated twice to remove small molecule impurities. The solution was lyophilized to dryness and redissolved in concentrated aqueous ammonia with mixing at room temperature for 2.5 h followed by concentration in vacuo to remove most of the ammonia. The conjugated oligonucleotide was purified and desalted by RP-HPLC and lyophilized to give ISIS 666881 in 90% yield by weight (42 mg, 4.7 .mu.mol).
TABLE-US-00025 GalNAc.sub.3-10 conjugated oligonucleotide SEQ ASO Sequence (5' to 3') 5' group ID No. ISIS 660254 NH.sub.2(CH.sub.2).sub.6-.sub.oA.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.su- b.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds Hexylamine 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.esT.sub- .ds.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666881 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.- es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds GalNAc.sub.3-10 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
Example 47: Preparation of Oligonucleotide 102 Comprising GalNAc.sub.3-8
##STR00196## ##STR00197## ##STR00198##
[0850] The triacid 90 (4 g, 14.43 mmol) was dissolved in DMF (120 mL) and N,N-Diisopropylethylamine (12.35 mL, 72 mmoles). Pentafluorophenyl trifluoroacetate (8.9 mL, 52 mmoles) was added dropwise, under argon, and the reaction was allowed to stir at room temperature for 30 minutes. Boc-diamine 91a or 91b (68.87 mmol) was added, along with N,N-Diisopropylethylamine (12.35 mL, 72 mmoles), and the reaction was allowed to stir at room temperature for 16 hours. At that time, the DMF was reduced by >75% under reduced pressure, and then the mixture was dissolved in dichloromethane. The organic layer was washed with sodium bicarbonate, water and brine. The organic layer was then separated and dried over sodium sulfate, filtered and reduced to an oil under reduced pressure. The resultant oil was purified by silica gel chromatography (2%-->10% methanol/dichloromethane) to give compounds 92a and 92b in an approximate 80% yield. LCMS and proton NMR were consistent with the structure.
[0851] Compound 92a or 92b (6.7 mmoles) was treated with 20 mL of dichloromethane and 20 mL of trifluoroacetic acid at room temperature for 16 hours. The resultant solution was evaporated and then dissolved in methanol and treated with DOWEX-OH resin for 30 minutes. The resultant solution was filtered and reduced to an oil under reduced pressure to give 85-90% yield of compounds 93a and 93b.
[0852] Compounds 7 or 64 (9.6 mmoles) were treated with HBTU (3.7 g, 9.6 mmoles) and N,N-Diisopropylethylamine (5 mL) in DMF (20 mL) for 15 minutes. To this was added either compounds 93a or 93b (3 mmoles), and allowed to stir at room temperature for 16 hours. At that time, the DMF was reduced by >75% under reduced pressure, and then the mixture was dissolved in dichloromethane. The organic layer was washed with sodium bicarbonate, water and brine. The organic layer was then separated and dried over sodium sulfate, filtered and reduced to an oil under reduced pressure. The resultant oil was purified by silica gel chromatography (5%-->20% methanol/dichloromethane) to give compounds 96a-d in 20-40% yield. LCMS and proton NMR was consistent with the structure.
[0853] Compounds 96a-d (0.75 mmoles), individually, were hydrogenated over Raney Nickel for 3 hours in Ethanol (75 mL). At that time, the catalyst was removed by filtration thru celite, and the ethanol removed under reduced pressure to give compounds 97a-d in 80-90% yield. LCMS and proton NMR were consistent with the structure.
[0854] Compound 23 (0.32 g, 0.53 mmoles) was treated with HBTU (0.2 g, 0.53 mmoles) and N,N-Diisopropylethylamine (0.19 mL, 1.14 mmoles) in DMF (30 mL) for 15 minutes. To this was added compounds 97a-d (0.38 mmoles), individually, and allowed to stir at room temperature for 16 hours. At that time, the DMF was reduced by >75% under reduced pressure, and then the mixture was dissolved in dichloromethane. The organic layer was washed with sodium bicarbonate, water and brine. The organic layer was then separated and dried over sodium sulfate, filtered and reduced to an oil under reduced pressure. The resultant oil was purified by silica gel chromatography (2%-->20% methanol/dichloromethane) to give compounds 98a-d in 30-40% yield. LCMS and proton NMR was consistent with the structure.
[0855] Compound 99 (0.17 g, 0.76 mmoles) was treated with HBTU (0.29 g, 0.76 mmoles) and N,N-Diisopropylethylamine (0.35 mL, 2.0 mmoles) in DMF (50 mL) for 15 minutes. To this was added compounds 97a-d (0.51 mmoles), individually, and allowed to stir at room temperature for 16 hours. At that time, the DMF was reduced by >75% under reduced pressure, and then the mixture was dissolved in dichloromethane. The organic layer was washed with sodium bicarbonate, water and brine. The organic layer was then separated and dried over sodium sulfate, filtered and reduced to an oil under reduced pressure. The resultant oil was purified by silica gel chromatography (5%-->20% methanol/dichloromethane) to give compounds 100a-d in 40-60% yield. LCMS and proton NMR was consistent with the structure.
[0856] Compounds 100a-d (0.16 mmoles), individually, were hydrogenated over 10% Pd(OH).sub.2/C for 3 hours in methanol/ethyl acetate (1:1, 50 mL). At that time, the catalyst was removed by filtration thru celite, and the organics removed under reduced pressure to give compounds 101a-d in 80-90% yield. LCMS and proton NMR was consistent with the structure.
[0857] Compounds 101a-d (0.15 mmoles), individually, were dissolved in DMF (15 mL) and pyridine (0.016 mL, 0.2 mmoles). Pentafluorophenyl trifluoroacetate (0.034 mL, 0.2 mmoles) was added dropwise, under argon, and the reaction was allowed to stir at room temperature for 30 minutes. At that time, the DMF was reduced by >75% under reduced pressure, and then the mixture was dissolved in dichloromethane. The organic layer was washed with sodium bicarbonate, water and brine. The organic layer was then separated and dried over sodium sulfate, filtered and reduced to an oil under reduced pressure. The resultant oil was purified by silica gel chromatography (2%-->5% methanol/dichloromethane) to give compounds 102a-d in an approximate 80% yield. LCMS and proton NMR were consistent with the structure.
##STR00199##
[0858] Oligomeric Compound 102, comprising a GalNAc.sub.3-8 conjugate group, was prepared using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-8 (GalNAc.sub.3-8.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In a preferred embodiment, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--.
[0859] The structure of GalNAc.sub.3-8 (GalNAc.sub.3-8.sub.a-CM-) is shown below:
##STR00200##
Example 48: Preparation of Oligonucleotide 119 Comprising GalNAc.sub.3-7
##STR00201## ##STR00202##
[0861] Compound 112 was synthesized following the procedure described in the literature (J. Med. Chem. 2004, 47, 5798-5808).
[0862] Compound 112 (5 g, 8.6 mmol) was dissolved in 1:1 methanol/ethyl acetate (22 mL/22 mL). Palladium hydroxide on carbon (0.5 g) was added. The reaction mixture was stirred at room temperature under hydrogen for 12 h. The reaction mixture was filtered through a pad of celite and washed the pad with 1:1 methanol/ethyl acetate. The filtrate and the washings were combined and concentrated to dryness to yield Compound 105a (quantitative). The structure was confirmed by LCMS.
[0863] Compound 113 (1.25 g, 2.7 mmol), HBTU (3.2 g, 8.4 mmol) and DIEA (2.8 mL, 16.2 mmol) were dissolved in anhydrous DMF (17 mL) and the reaction mixture was stirred at room temperature for 5 min. To this a solution of Compound 105a (3.77 g, 8.4 mmol) in anhydrous DMF (20 mL) was added. The reaction was stirred at room temperature for 6 h. Solvent was removed under reduced pressure to get an oil. The residue was dissolved in CH.sub.2Cl.sub.2 (100 mL) and washed with aqueous saturated NaHCO.sub.3 solution (100 mL) and brine (100 mL). The organic phase was separated, dried (Na.sub.2SO.sub.4), filtered and evaporated. The residue was purified by silica gel column chromatography and eluted with 10 to 20% MeOH in dichloromethane to yield Compound 114 (1.45 g, 30%). The structure was confirmed by LCMS and .sup.1H NMR analysis.
[0864] Compound 114 (1.43 g, 0.8 mmol) was dissolved in 1:1 methanol/ethyl acetate (4 mL/4 mL). Palladium on carbon (wet, 0.14 g) was added. The reaction mixture was flushed with hydrogen and stirred at room temperature under hydrogen for 12 h. The reaction mixture was filtered through a pad of celite. The celite pad was washed with methanol/ethyl acetate (1:1). The filtrate and the washings were combined together and evaporated under reduced pressure to yield Compound 115 (quantitative). The structure was confirmed by LCMS and .sup.1H NMR analysis.
[0865] Compound 83a (0.17 g, 0.75 mmol), HBTU (0.31 g, 0.83 mmol) and DIEA (0.26 mL, 1.5 mmol) were dissolved in anhydrous DMF (5 mL) and the reaction mixture was stirred at room temperature for 5 min. To this a solution of Compound 115 (1.22 g, 0.75 mmol) in anhydrous DMF was added and the reaction was stirred at room temperature for 6 h. The solvent was removed under reduced pressure and the residue was dissolved in CH.sub.2Cl.sub.2. The organic layer was washed aqueous saturated NaHCO.sub.3 solution and brine and dried over anhydrous Na.sub.2SO.sub.4 and filtered. The organic layer was concentrated to dryness and the residue obtained was purified by silica gel column chromatography and eluted with 3 to 15% MeOH in dichloromethane to yield Compound 116 (0.84 g, 61%). The structure was confirmed by LC MS and .sup.1H NMR analysis.
##STR00203##
[0866] Compound 116 (0.74 g, 0.4 mmol) was dissolved in 1:1 methanol/ethyl acetate (5 mL/5 mL). Palladium on carbon (wet, 0.074 g) was added. The reaction mixture was flushed with hydrogen and stirred at room temperature under hydrogen for 12 h. The reaction mixture was filtered through a pad of celite. The celite pad was washed with methanol/ethyl acetate (1:1). The filtrate and the washings were combined together and evaporated under reduced pressure to yield compound 117 (0.73 g, 98%). The structure was confirmed by LCMS and .sup.1H NMR analysis.
[0867] Compound 117 (0.63 g, 0.36 mmol) was dissolved in anhydrous DMF (3 mL). To this solution N,N-Diisopropylethylamine (70 .mu.L, 0.4 mmol) and pentafluorophenyl trifluoroacetate (72 .mu.L, 0.42 mmol) were added. The reaction mixture was stirred at room temperature for 12 h and poured into a aqueous saturated NaHCO.sub.3 solution. The mixture was extracted with dichloromethane, washed with brine and dried over anhydrous Na.sub.2SO.sub.4. The dichloromethane solution was concentrated to dryness and purified with silica gel column chromatography and eluted with 5 to 10% MeOH in dichloromethane to yield compound 118 (0.51 g, 79%). The structure was confirmed by LCMS and .sup.1H and .sup.1H and .sup.19F NMR.
##STR00204##
[0868] Oligomeric Compound 119, comprising a GalNAc.sub.3-7 conjugate group, was prepared using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-7 (GalNAc.sub.3-7.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--.
[0869] The structure of GalNAc.sub.3-7 (GalNAc.sub.3-7.sub.a-CM-) is shown below:
##STR00205##
Example 49: Preparation of Oligonucleotide 132 Comprising GalNAc.sub.3-5
##STR00206##
[0871] Compound 120 (14.01 g, 40 mmol) and HBTU (14.06 g, 37 mmol) were dissolved in anhydrous DMF (80 mL). Triethylamine (11.2 mL, 80.35 mmol) was added and stirred for 5 min. The reaction mixture was cooled in an ice bath and a solution of compound 121 (10 g, mmol) in anhydrous DMF (20 mL) was added. Additional triethylamine (4.5 mL, 32.28 mmol) was added and the reaction mixture was stirred for 18 h under an argon atmosphere. The reaction was monitored by TLC (ethyl acetate:hexane; 1:1; Rf=0.47). The solvent was removed under reduced pressure. The residue was taken up in EtOAc (300 mL) and washed with 1M NaHSO.sub.4 (3.times.150 mL), aqueous saturated NaHCO.sub.3 solution (3.times.150 mL) and brine (2.times.100 mL). Organic layer was dried with Na.sub.2SO.sub.4. Drying agent was removed by filtration and organic layer was concentrated by rotary evaporation. Crude mixture was purified by silica gel column chromatography and eluted by using 35-50% EtOAc in hexane to yield a compound 122 (15.50 g, 78.13%). The structure was confirmed by LCMS and .sup.1H NMR analysis. Mass m/z 589.3 [M+H].sup.+.
[0872] A solution of LiOH (92.15 mmol) in water (20 mL) and THF (10 mL) was added to a cooled solution of Compound 122 (7.75 g, 13.16 mmol) dissolved in methanol (15 mL). The reaction mixture was stirred at room temperature for 45 min. and monitored by TLC (EtOAc:hexane; 1:1). The reaction mixture was concentrated to half the volume under reduced pressure. The remaining solution was cooled an ice bath and neutralized by adding concentrated HCl. The reaction mixture was diluted, extracted with EtOAc (120 mL) and washed with brine (100 mL). An emulsion formed and cleared upon standing overnight. The organic layer was separated dried (Na.sub.2SO.sub.4), filtered and evaporated to yield Compound 123 (8.42 g). Residual salt is the likely cause of excess mass. LCMS is consistent with structure. Product was used without any further purification. M.W.cal: 574.36; M.W.fd: 575.3 [M+H].sup.+.
##STR00207##
[0873] Compound 126 was synthesized following the procedure described in the literature (J. Am. Chem. Soc. 2011, 133, 958-963).
##STR00208## ##STR00209##
[0874] Compound 123 (7.419 g, 12.91 mmol), HOBt (3.49 g, 25.82 mmol) and compound 126 (6.33 g, 16.14 mmol) were dissolved in and DMF (40 mL) and the resulting reaction mixture was cooled in an ice bath. To this N,N-Diisopropylethylamine (4.42 mL, 25.82 mmol), PyBop (8.7 g, 16.7 mmol) followed by Bop coupling reagent (1.17 g, 2.66 mmol) were added under an argon atmosphere. The ice bath was removed and the solution was allowed to warm to room temperature. The reaction was completed after 1 h as determined by TLC (DCM:MeOH:AA; 89:10:1). The reaction mixture was concentrated under reduced pressure. The residue was dissolved in EtOAc (200 mL) and washed with 1 M NaHSO.sub.4 (3.times.100 mL), aqueous saturated NaHCO.sub.3 (3.times.100 mL) and brine (2.times.100 mL). The organic phase separated dried (Na.sub.2SO.sub.4), filtered and concentrated. The residue was purified by silica gel column chromatography with a gradient of 50% hexanes/EtOAC to 100% EtOAc to yield Compound 127 (9.4 g) as a white foam. LCMS and .sup.1H NMR were consistent with structure. Mass m/z 778.4 [M+H].sup.+.
[0875] Trifluoroacetic acid (12 mL) was added to a solution of compound 127 (1.57 g, 2.02 mmol) in dichloromethane (12 mL) and stirred at room temperature for 1 h. The reaction mixture was co-evaporated with toluene (30 mL) under reduced pressure to dryness. The residue obtained was co-evaporated twice with acetonitrile (30 mL) and toluene (40 mL) to yield Compound 128 (1.67 g) as trifluoro acetate salt and used for next step without further purification. LCMS and .sup.1H NMR were consistent with structure. Mass m/z 478.2 [M+H].sup.+.
[0876] Compound 7 (0.43 g, 0.963 mmol), HATU (0.35 g, 0.91 mmol), and HOAt (0.035 g, 0.26 mmol) were combined together and dried for 4 h over P.sub.2O.sub.5 under reduced pressure in a round bottom flask and then dissolved in anhydrous DMF (1 mL) and stirred for 5 min. To this a solution of compound 128 (0.20 g, 0.26 mmol) in anhydrous DMF (0.2 mL) and N,N-Diisopropylethylamine (0.2 mL) was added. The reaction mixture was stirred at room temperature under an argon atmosphere. The reaction was complete after 30 min as determined by LCMS and TLC (7% MeOH/DCM). The reaction mixture was concentrated under reduced pressure. The residue was dissolved in DCM (30 mL) and washed with 1 M NaHSO.sub.4 (3.times.20 mL), aqueous saturated NaHCO.sub.3 (3.times.20 mL) and brine (3.times.20 mL). The organic phase was separated, dried over Na.sub.2SO.sub.4, filtered and concentrated. The residue was purified by silica gel column chromatography using 5-15% MeOH in dichloromethane to yield Compound 129 (96.6 mg). LC MS and .sup.1H NMR are consistent with structure. Mass m/z 883.4 [M+2H].sup.+.
[0877] Compound 129 (0.09 g, 0.051 mmol) was dissolved in methanol (5 mL) in 20 mL scintillation vial. To this was added a small amount of 10% Pd/C (0.015 mg) and the reaction vessel was flushed with H.sub.2 gas. The reaction mixture was stirred at room temperature under H.sub.2 atmosphere for 18 h. The reaction mixture was filtered through a pad of Celite and the Celite pad was washed with methanol. The filtrate washings were pooled together and concentrated under reduced pressure to yield Compound 130 (0.08 g). LCMS and .sup.1H NMR were consistent with structure. The product was used without further purification. Mass m/z 838.3 [M+2H].sup.+.
[0878] To a 10 mL pointed round bottom flask were added compound 130 (75.8 mg, 0.046 mmol), 0.37 M pyridine/DMF (200 .mu.L) and a stir bar. To this solution was added 0.7 M pentafluorophenyl trifluoroacetate/DMF (100 .mu.L) drop wise with stirring. The reaction was completed after 1 h as determined by LC MS. The solvent was removed under reduced pressure and the residue was dissolved in CHCl.sub.3 (.about.10 mL). The organic layer was partitioned against NaHSO.sub.4 (1 M, 10 mL), aqueous saturated NaHCO.sub.3 (10 mL) and brine (10 mL) three times each. The organic phase separated and dried over Na.sub.2SO.sub.4, filtered and concentrated to yield Compound 131 (77.7 mg). LCMS is consistent with structure. Used without further purification. Mass m/z 921.3 [M+2H].sup.+.
##STR00210##
[0879] Oligomeric Compound 132, comprising a GalNAc.sub.3-5 conjugate group, was prepared using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-5 (GalNAc.sub.3-5.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--.
[0880] The structure of GalNAc.sub.3-5 (GalNAc.sub.3-5.sub.a-CM-) is shown below:
##STR00211##
Example 50: Preparation of Oligonucleotide 144 Comprising GalNAc.sub.4-11
##STR00212## ##STR00213##
[0882] Synthesis of Compound 134. To a Merrifield flask was added aminomethyl VIMAD resin (2.5 g, 450 mol/g) that was washed with acetonitrile, dimethylformamide, dichloromethane and acetonitrile. The resin was swelled in acetonitrile (4 mL). Compound 133 was pre-activated in a 100 mL round bottom flask by adding 20 (1.0 mmol, 0.747 g), TBTU (1.0 mmol, 0.321 g), acetonitrile (5 mL) and DIEA (3.0 mmol, 0.5 mL). This solution was allowed to stir for 5 min and was then added to the Merrifield flask with shaking. The suspension was allowed to shake for 3 h. The reaction mixture was drained and the resin was washed with acetonitrile, DMF and DCM. New resin loading was quantitated by measuring the absorbance of the DMT cation at 500 nm (extinction coefficient=76000) in DCM and determined to be 238 .mu.mol/g. The resin was capped by suspending in an acetic anhydride solution for ten minutes three times.
[0883] The solid support bound compound 141 was synthesized using iterative Fmoc-based solid phase peptide synthesis methods. A small amount of solid support was withdrawn and suspended in aqueous ammonia (28-30 wt %) for 6 h. The cleaved compound was analyzed by LC-MS and the observed mass was consistent with structure. Mass m/z 1063.8 [M+2H].sup.+.
[0884] The solid support bound compound 142 was synthesized using solid phase peptide synthesis methods.
##STR00214##
[0885] The solid support bound compound 143 was synthesized using standard solid phase synthesis on a DNA synthesizer.
[0886] The solid support bound compound 143 was suspended in aqueous ammonia (28-30 wt %) and heated at 55.degree. C. for 16 h. The solution was cooled and the solid support was filtered. The filtrate was concentrated and the residue dissolved in water and purified by HPLC on a strong anion exchange column. The fractions containing full length compound 144 were pooled together and desalted. The resulting GalNAc.sub.4-11 conjugated oligomeric compound was analyzed by LC-MS and the observed mass was consistent with structure.
[0887] The GalNAc.sub.4 cluster portion of the conjugate group GalNAc.sub.4-11 (GalNAc.sub.4-11.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--.
[0888] The structure of GalNAc.sub.4-11 (GalNAc.sub.4-11.sub.a-CM) is shown below:
##STR00215##
Example 51: Preparation of Oligonucleotide 155 Comprising GalNAc.sub.3-6
##STR00216##
[0890] Compound 146 was synthesized as described in the literature (Analytical Biochemistry 1995, 229, 54-60).
##STR00217##
[0891] Compound 4 (15 g, 45.55 mmol) and compound 35b (14.3 grams, 57 mmol) were dissolved in CH.sub.2Cl.sub.2 (200 ml). Activated molecular sieves (4 .ANG.. 2 g, powdered) were added, and the reaction was allowed to stir for 30 minutes under nitrogen atmosphere. TMS-OTf was added (4.1 ml, 22.77 mmol) and the reaction was allowed to stir at room temp overnight. Upon completion, the reaction was quenched by pouring into solution of saturated aqueous NaHCO.sub.3 (500 ml) and crushed ice (.about.150 g). The organic layer was separated, washed with brine, dried over MgSO.sub.4, filtered, and was concentrated to an orange oil under reduced pressure. The crude material was purified by silica gel column chromatography and eluted with 2-10% MeOH in CH.sub.2Cl.sub.2 to yield Compound 112 (16.53 g, 63%). LCMS and .sup.1H NMR were consistent with the expected compound.
[0892] Compound 112 (4.27 g, 7.35 mmol) was dissolved in 1:1 MeOH/EtOAc (40 ml). The reaction mixture was purged by bubbling a stream of argon through the solution for 15 minutes. Pearlman's catalyst (palladium hydroxide on carbon, 400 mg) was added, and hydrogen gas was bubbled through the solution for 30 minutes. Upon completion (TLC 10% MeOH in CH.sub.2Cl.sub.2, and LCMS), the catalyst was removed by filtration through a pad of celite. The filtrate was concentrated by rotary evaporation, and was dried briefly under high vacuum to yield Compound 105a (3.28 g). LCMS and 1H NMR were consistent with desired product.
[0893] Compound 147 (2.31 g, 11 mmol) was dissolved in anhydrous DMF (100 mL). N,N-Diisopropylethylamine (DIEA, 3.9 mL, 22 mmol) was added, followed by HBTU (4 g, 10.5 mmol). The reaction mixture was allowed to stir for .about.15 minutes under nitrogen. To this a solution of compound 105a (3.3 g, 7.4 mmol) in dry DMF was added and stirred for 2 h under nitrogen atmosphere. The reaction was diluted with EtOAc and washed with saturated aqueous NaHCO.sub.3 and brine. The organics phase was separated, dried (MgSO.sub.4), filtered, and concentrated to an orange syrup. The crude material was purified by column chromatography 2-5% MeOH in CH.sub.2C2 to yield Compound 148 (3.44 g, 73%). LCMS and .sup.1H NMR were consistent with the expected product.
[0894] Compound 148 (3.3 g, 5.2 mmol) was dissolved in 1:1 MeOH/EtOAc (75 ml). The reaction mixture was purged by bubbling a stream of argon through the solution for 15 minutes. Pearlman's catalyst (palladium hydroxide on carbon) was added (350 mg). Hydrogen gas was bubbled through the solution for 30 minutes. Upon completion (TLC 10% MeOH in DCM, and LCMS), the catalyst was removed by filtration through a pad of celite. The filtrate was concentrated by rotary evaporation, and was dried briefly under high vacuum to yield Compound 149 (2.6 g). LCMS was consistent with desired product. The residue was dissolved in dry DMF (10 ml) was used immediately in the next step.
##STR00218##
[0895] Compound 146 (0.68 g, 1.73 mmol) was dissolved in dry DMF (20 ml). To this DIEA (450 .mu.L, 2.6 mmol, 1.5 eq.) and HBTU (1.96 g, 0.5.2 mmol) were added. The reaction mixture was allowed to stir for 15 minutes at room temperature under nitrogen. A solution of compound 149 (2.6 g) in anhydrous DMF (10 mL) was added. The pH of the reaction was adjusted to pH=9-10 by addition of DIEA (if necessary). The reaction was allowed to stir at room temperature under nitrogen for 2 h. Upon completion the reaction was diluted with EtOAc (100 mL), and washed with aqueous saturated aqueous NaHCO.sub.3, followed by brine. The organic phase was separated, dried over MgSO.sub.4, filtered, and concentrated. The residue was purified by silica gel column chromatography and eluted with 2-10% MeOH in CH.sub.2Cl.sub.2 to yield Compound 150 (0.62 g, 20%). LCMS and 1H NMR were consistent with the desired product.
[0896] Compound 150 (0.62 g) was dissolved in 1:1 MeOH/EtOAc (5 L). The reaction mixture was purged by bubbling a stream of argon through the solution for 15 minutes. Pearlman's catalyst (palladium hydroxide on carbon) was added (60 mg). Hydrogen gas was bubbled through the solution for 30 minutes. Upon completion (TLC 10% MeOH in DCM, and LCMS), the catalyst was removed by filtration (syringe-tip Teflon filter, 0.45 .mu.m). The filtrate was concentrated by rotary evaporation, and was dried briefly under high vacuum to yield Compound 151 (0.57 g). The LCMS was consistent with the desired product. The product was dissolved in 4 mL dry DMF and was used immediately in the next step.
##STR00219##
[0897] Compound 83a (0.11 g, 0.33 mmol) was dissolved in anhydrous DMF (5 mL) and N,N-Diisopropylethylamine (75 .mu.L, 1 mmol) and PFP-TFA (90 .mu.L, 0.76 mmol) were added. The reaction mixture turned magenta upon contact, and gradually turned orange over the next 30 minutes. Progress of reaction was monitored by TLC and LCMS. Upon completion (formation of the PFP ester), a solution of compound 151 (0.57 g, 0.33 mmol) in DMF was added. The pH of the reaction was adjusted to pH=9-10 by addition of N,N-Diisopropylethylamine (if necessary). The reaction mixture was stirred under nitrogen for .about.30 min. Upon completion, the majority of the solvent was removed under reduced pressure. The residue was diluted with CH.sub.2Cl.sub.2 and washed with aqueous saturated NaHCO.sub.3, followed by brine. The organic phase separated, dried over MgSO.sub.4, filtered, and concentrated to an orange syrup. The residue was purified by silica gel column chromatography (2-10% MeOH in CH.sub.2Cl.sub.2) to yield Compound 152 (0.35 g, 55%). LCMS and .sup.1H NMR were consistent with the desired product.
[0898] Compound 152 (0.35 g, 0.182 mmol) was dissolved in 1:1 MeOH/EtOAc (10 mL). The reaction mixture was purged by bubbling a stream of argon thru the solution for 15 minutes. Pearlman's catalyst (palladium hydroxide on carbon) was added (35 mg). Hydrogen gas was bubbled thru the solution for 30 minutes. Upon completion (TLC 10% MeOH in DCM, and LCMS), the catalyst was removed by filtration (syringe-tip Teflon filter, 0.45 .mu.m). The filtrate was concentrated by rotary evaporation, and was dried briefly under high vacuum to yield Compound 153 (0.33 g, quantitative). The LCMS was consistent with desired product.
[0899] Compound 153 (0.33 g, 0.18 mmol) was dissolved in anhydrous DMF (5 mL) with stirring under nitrogen. To this N,N-Diisopropylethylamine (65 .mu.L, 0.37 mmol) and PFP-TFA (35 .mu.L, 0.28 mmol) were added. The reaction mixture was stirred under nitrogen for .about.30 min. The reaction mixture turned magenta upon contact, and gradually turned orange. The pH of the reaction mixture was maintained at pH=9-10 by adding more N,-Diisopropylethylamine. The progress of the reaction was monitored by TLC and LCMS. Upon completion, the majority of the solvent was removed under reduced pressure. The residue was diluted with CH.sub.2Cl.sub.2 (50 mL), and washed with saturated aqueous NaHCO.sub.3, followed by brine. The organic layer was dried over MgSO.sub.4, filtered, and concentrated to an orange syrup. The residue was purified by column chromatography and eluted with 2-10% MeOH in CH.sub.2Cl.sub.2 to yield Compound 154 (0.29 g, 79%). LCMS and 1H NMR were consistent with the desired product.
##STR00220##
[0900] Oligomeric Compound 155, comprising a GalNAc.sub.3-6 conjugate group, was prepared using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-6 (GalNAc.sub.3-6.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--.
[0901] The structure of GalNAc.sub.3-6 (GalNAc.sub.3-6.sub.a-CM-) is shown below:
##STR00221##
Example 52: Preparation of Oligonucleotide 160 Comprising GalNAc.sub.3-9
##STR00222##
[0903] Compound 156 was synthesized following the procedure described in the literature (J. Med. Chem. 2004, 47, 5798-5808).
[0904] Compound 156, (18.60 g, 29.28 mmol) was dissolved in methanol (200 mL). Palladium on carbon (6.15 g, 10 wt %, loading (dry basis), matrix carbon powder, wet) was added. The reaction mixture was stirred at room temperature under hydrogen for 18 h. The reaction mixture was filtered through a pad of celite and the celite pad was washed thoroughly with methanol. The combined filtrate was washed and concentrated to dryness. The residue was purified by silica gel column chromatography and eluted with 5-10% methanol in dichloromethane to yield Compound 157 (14.26 g, 89%). Mass m/z 544.1 [M-H].sup.-.
[0905] Compound 157 (5 g, 9.17 mmol) was dissolved in anhydrous DMF (30 mL). HBTU (3.65 g, 9.61 mmol) and N,N-Diisopropylethylamine (13.73 mL, 78.81 mmol) were added and the reaction mixture was stirred at room temperature for 5 minutes. To this a solution of compound 47 (2.96 g, 7.04 mmol) was added. The reaction was stirred at room temperature for 8 h. The reaction mixture was poured into a saturated NaHCO.sub.3 aqueous solution. The mixture was extracted with ethyl acetate and the organic layer was washed with brine and dried (Na.sub.2SO.sub.4), filtered and evaporated. The residue obtained was purified by silica gel column chromatography and eluted with 50% ethyl acetate in hexane to yield compound 158 (8.25 g, 73.3%). The structure was confirmed by MS and .sup.1H NMR analysis.
[0906] Compound 158 (7.2 g, 7.61 mmol) was dried over P.sub.2O.sub.5 under reduced pressure. The dried compound was dissolved in anhydrous DMF (50 mL). To this 1H-tetrazole (0.43 g, 6.09 mmol) and N-methylimidazole (0.3 mL, 3.81 mmol) and 2-cyanoethyl-N,N,N',N'-tetraisopropyl phosphorodiamidite (3.65 mL, 11.50 mmol) were added. The reaction mixture was stirred t under an argon atmosphere for 4 h. The reaction mixture was diluted with ethyl acetate (200 mL). The reaction mixture was washed with saturated NaHCO.sub.3 and brine. The organic phase was separated, dried (Na.sub.2SO.sub.4), filtered and evaporated. The residue was purified by silica gel column chromatography and eluted with 50-90% ethyl acetate in hexane to yield Compound 159 (7.82 g, 80.5%). The structure was confirmed by LCMS and .sup.31P NMR analysis.
##STR00223##
[0907] Oligomeric Compound 160, comprising a GalNAc.sub.3-9 conjugate group, was prepared using standard oligonucleotide synthesis procedures. Three units of compound 159 were coupled to the solid support, followed by nucleotide phosphoramidites. Treatment of the protected oligomeric compound with aqueous ammonia yielded compound 160. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-9 (GalNAc.sub.3-9.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-9 (GalNAc.sub.3-9.sub.a-CM) is shown below:
##STR00224##
Example 53: Alternate Procedure for Preparation of Compound 18 (GalNAc.sub.3-1a and GalNAc.sub.3-3a)
##STR00225##
[0909] Lactone 161 was reacted with diamino propane (3-5 eq) or Mono-Boc protected diamino propane (1 eq) to provide alcohol 162a or 162b. When unprotected propanediamine was used for the above reaction, the excess diamine was removed by evaporation under high vacuum and the free amino group in 162a was protected using CbzCl to provide 162b as a white solid after purification by column chromatography. Alcohol 162b was further reacted with compound 4 in the presence of TMSOTf to provide 163a which was converted to 163b by removal of the Cbz group using catalytic hydrogenation. The pentafluorophenyl (PFP) ester 164 was prepared by reacting triacid 113 (see Example 48) with PFPTFA (3.5 eq) and pyridine (3.5 eq) in DMF (0.1 to 0.5 M). The triester 164 was directly reacted with the amine 163b (3-4 eq) and DIPEA (3-4 eq) to provide Compound 18. The above method greatly facilitates purification of intermediates and minimizes the formation of byproducts which are formed using the procedure described in Example 4.
Example 54: Alternate Procedure for Preparation of Compound 18 (GalNAc.sub.3-1a and GalNAc.sub.3-3a)
##STR00226##
[0911] The triPFP ester 164 was prepared from acid 113 using the procedure outlined in example 53 above and reacted with mono-Boc protected diamine to provide 165 in essentially quantitative yield. The Boc groups were removed with hydrochloric acid or trifluoroacetic acid to provide the triamine which was reacted with the PFP activated acid 166 in the presence of a suitable base such as DIPEA to provide Compound 18.
[0912] The PFP protected Gal-NAc acid 166 was prepared from the corresponding acid by treatment with PFPTFA (1-1.2 eq) and pyridine (1-1.2 eq) in DMF. The precursor acid in turn was prepared from the corresponding alcohol by oxidation using TEMPO (0.2 eq) and BAIB in acetonitrile and water. The precursor alcohol was prepared from sugar intermediate 4 by reaction with 1,6-hexanediol (or 1,5-pentanediol or other diol for other n values) (2-4 eq) and TMSOTf using conditions described previously in example 47.
Example 55: Dose-Dependent Study of Oligonucleotides Comprising Either a 3' or 5'-Conjugate Group (Comparison of GalNAc.sub.3-1, 3, 8 and 9) Targeting SRB-1 In Vivo
[0913] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Unconjugated ISIS 353382 was included as a standard. Each of the various GalNAc.sub.3 conjugate groups was attached at either the 3' or 5' terminus of the respective oligonucleotide by a phosphodiester linked 2'-deoxyadenosine nucleoside (cleavable moiety).
TABLE-US-00026 TABLE 39 Modified ASO targeting SRB-1 SEQ ID ASO Sequence (5' to 3') Motif Conjugate No. ISIS 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 none 2256 (parent) .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.e- sT.sub.e ISIS 655861 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 GalNAc.sub.3-1 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1.sub.a ISIS 664078 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 GalNAc.sub.3-9 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-9.sub.a ISIS 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.do 5/10/5 GalNAc.sub.3-3 2258 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 665001 GalNAc.sub.3-8.sub.a-.sub.o'A.sub.do 5/10/5 GalNAc.sub.3-8 2258 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0914] The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9. The structure of GalNAc.sub.3-9 was shown previously in Example 52. The structure of GalNAc.sub.3-3 was shown previously in Example 39. The structure of GalNAc.sub.3-8 was shown previously in Example 47.
Treatment
[0915] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 353382, 655861, 664078, 661161, 665001 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0916] As illustrated in Table 40, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. Indeed, the antisense oligonucleotides comprising the phosphodiester linked GalNAc.sub.3-1 and GalNAc.sub.3-9 conjugates at the 3' terminus (ISIS 655861 and ISIS 664078) and the GalNAc.sub.3-3 and GalNAc.sub.3-8 conjugates linked at the 5' terminus (ISIS 661161 and ISIS 665001) showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 353382). Furthermore, ISIS 664078, comprising a GalNAc.sub.3-9 conjugate at the 3' terminus was essentially equipotent compared to ISIS 655861, which comprises a GalNAc.sub.3-1 conjugate at the 3' terminus. The 5' conjugated antisense oligonucleotides, ISIS 661161 and ISIS 665001, comprising a GalNAc.sub.3-3 or GalNAc.sub.3-9, respectively, had increased potency compared to the 3' conjugated antisense oligonucleotides (ISIS 655861 and ISIS 664078).
TABLE-US-00027 TABLE 40 ASOs containing GalNAc.sub.3-1, 3, 8 or 9 targeting SRB-1 Dosage SRB-1 mRNA ISIS No. (mg/kg) (% Saline) Conjugate Saline n/a 100 353382 3 88 none 10 68 30 36 655861 0.5 98 GalNAc.sub.3-1 (3') 1.5 76 5 31 15 20 664078 0.5 88 GalNAc.sub.3-9 (3') 1.5 85 5 46 15 20 661161 0.5 92 GalNAc.sub.3-3 (5') 1.5 59 5 19 15 11 665001 0.5 100 GalNAc.sub.3-8 (5') 1.5 73 5 29 15 13
[0917] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin and BUN were also evaluated. The change in body weights was evaluated with no significant change from the saline group. ALTs, ASTs, total bilirubin and BUN values are shown in the table below.
TABLE-US-00028 TABLE 41 Dosage Total ISIS No. mg/kg ALT AST Bilirubin BUN Conjugate Saline 24 59 0.1 37.52 353382 3 21 66 0.2 34.65 none 10 22 54 0.2 34.2 30 22 49 0.2 33.72 655861 0.5 25 62 0.2 30.65 GalNac.sub.3-1 (3') 1.5 23 48 0.2 30.97 5 28 49 0.1 32.92 15 40 97 0.1 31.62 664078 0.5 40 74 0.1 35.3 GalNac.sub.3-9 (3') 1.5 47 104 0.1 32.75 5 20 43 0.1 30.62 15 38 92 0.1 26.2 661161 0.5 101 162 0.1 34.17 GalNac.sub.3-3 (5') 1.5 g 42 100 0.1 33.37 5 g 23 99 0.1 34.97 15 53 83 0.1 34.8 665001 0.5 28 54 0.1 31.32 GalNac.sub.3-8 (5') 1.5 42 75 0.1 32.32 5 24 42 0.1 31.85 15 32 67 0.1 31.
Example 56: Dose-Dependent Study of Oligonucleotides Comprising Either a 3' or 5'-Conjugate Group (Comparison of GalNAc.sub.3-1, 2, 3, 5, 6, 7 and 10) Targeting SRB-1 In Vivo
[0918] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Unconjugated ISIS 353382 was included as a standard. Each of the various GalNAc.sub.3 conjugate groups was attached at the 5' terminus of the respective oligonucleotide by a phosphodiester linked 2'-deoxyadenosine nucleoside (cleavable moiety) except for ISIS 655861 which had the GalNAc.sub.3 conjugate group attached at the 3' terminus.
TABLE-US-00029 TABLE 42 Modified ASO targeting SRB-1 SEQ ASO Sequence (5' to 3') Motif Conjugate ID No. ISIS 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 no conjugate 2256 (parent) .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.e- sT.sub.e ISIS 655861 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 GalNAc.sub.3-1 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1.sub.a ISIS 664507 GalNAc.sub.3-2.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-2 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.do 5/10/5 GalNAc.sub.3-3 2258 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666224 GalNAc.sub.3-5.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-5 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666961 GalNAc.sub.3-6.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-6 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666981 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-7 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666881 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.- es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-10 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0919] The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9. The structure of GalNAc.sub.3-2.sub.a was shown previously in Example 37. The structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39. The structure of GalNAc.sub.3-5.sub.a was shown previously in Example 49. The structure of GalNAc.sub.3-6.sub.a was shown previously in Example 51. The structure of GalNAc.sub.3-7.sub.a was shown previously in Example 48. The structure of GalNAc.sub.3-10.sub.a was shown previously in Example 46.
Treatment
[0920] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 353382, 655861, 664507, 661161, 666224, 666961, 666981, 666881 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0921] As illustrated in Table 43, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. Indeed, the conjugated antisense oligonucleotides showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 353382). The 5' conjugated antisense oligonucleotides showed a slight increase in potency compared to the 3' conjugated antisense oligonucleotide.
TABLE-US-00030 TABLE 43 Dosage SRB-1 mRNA ISIS No. (mg/kg) (% Saline) Conjugate Saline n/a 100.0 353382 3 96.0 none 10 73.1 30 36.1 655861 0.5 99.4 GalNAc.sub.3-1 (3') 1.5 81.2 5 33.9 15 15.2 664507 0.5 102.0 GalNAc.sub.3-2 (5') 1.5 73.2 5 31.3 15 10.8 661161 0.5 90.7 GalNAc.sub.3-3 (5') 1.5 67.6 5 24.3 15 11.5 666224 0.5 96.1 GalNAc.sub.3-5 (5') 1.5 61.6 5 25.6 15 11.7 666961 0.5 85.5 GalNAc.sub.3-6 (5') 1.5 56.3 5 34.2 15 13.1 666981 0.5 84.7 GalNAc.sub.3-7 (5') 1.5 59.9 5 24.9 15 8.5 666881 0.5 100.0 GalNAc.sub.3-10 (5') 1.5 65.8 5 26.0 15 13.0
[0922] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin and BUN were also evaluated. The change in body weights was evaluated with no significant change from the saline group. ALTs, ASTs, total bilirubin and BUN values are shown in Table 44 below.
TABLE-US-00031 TABLE 44 ISIS Dosage Total No. mg/kg ALT AST Bilirubin BUN Conjugate Saline 26 57 0.2 27 353382 3 25 92 0.2 27 10 23 40 0.2 25 none 30 29 54 0.1 28 655861 0.5 25 71 0.2 34 GalNac.sub.3-1 (3') 1.5 28 60 0.2 26 5 26 63 0.2 28 15 25 61 0.2 28 664507 0.5 25 62 0.2 25 GalNac.sub.3-2 (5') 1.5 24 49 0.2 26 5 21 50 0.2 26 15 59 84 0.1 22 661161 0.5 20 42 0.2 29 GalNac.sub.3-3 (5') 1.5 g 37 74 0.2 25 5 g 28 61 0.2 29 15 21 41 0.2 25 666224 0.5 34 48 0.2 21 GalNac.sub.3-5 (5') 1.5 23 46 0.2 26 5 24 47 0.2 23 15 32 49 0.1 26 666961 0.5 17 63 0.2 26 GalNAc.sub.3-6 (5') 1.5 23 68 0.2 26 5 25 66 0.2 26 15 29 107 0.2 28 666981 0.5 24 48 0.2 26 GalNAc.sub.3-7 (5') 1.5 30 55 0.2 24 5 46 74 0.1 24 15 29 58 0.1 26 666881 0.5 20 65 0.2 27 GalNAc.sub.3-10 (5') 1.5 23 59 0.2 24 5 45 70 0.2 26 15 21 57 0.2 24
Example 57: Duration of Action Study of Oligonucleotides Comprising a 3'-Conjugate Group Targeting ApoC III In Vivo
[0923] Mice were injected once with the doses indicated below and monitored over the course of 42 days for ApoC-III and plasma triglycerides (Plasma TG) levels. The study was performed using 3 transgenic mice that express human APOC-III in each group.
TABLE-US-00032 TABLE 45 Modified ASO targeting ApoC III SEQ ID ASO Sequence (5' to 3') Linkages No. ISIS A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds PS 2248 304801 .sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT- .sub.esT.sub.esA.sub.esT.sub.e ISIS A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.ds PS 2249 647535 A.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT.sub.esT.sub.esA.sub.esT.sub- .eoA.sub.do'-GalNAc.sub.3-1.sub.a ISIS A.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.dsT.sub.dsT- .sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.ds PO/PS 2249 647536 A.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.eoT.sub.eoT.sub.esA.sub.esT.sub- .eoA.sub.do'-GalNAc.sub.3-1.sub.a Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0924] The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9.
TABLE-US-00033 TABLE 46 ApoC III mRNA (% Saline on Day 1) and Plasma TG Levels (% Saline on Day 1) ASO Dose Target Day 3 Day 7 Day 14 Day 35 Day 42 Saline 0 mg/kg ApoC-III 98 100 100 95 116 ISIS 304801 30 mg/kg ApoC-III 28 30 41 65 74 ISIS 647535 10 mg/kg ApoC-III 16 19 25 74 94 ISIS 647536 10 mg/kg ApoC-III 18 16 17 35 51 Saline 0 mg/kg Plasma TG 121 130 123 105 109 ISIS 304801 30 mg/kg Plasma TG 34 37 50 69 69 ISIS 647535 10 mg/kg Plasma TG 18 14 24 18 71 ISIS 647536 10 mg/kg Plasma TG 21 19 15 32 35
[0925] As can be seen in the table above the duration of action increased with addition of the 3'-conjugate group compared to the unconjugated oligonucleotide. There was a further increase in the duration of action for the conjugated mixed PO/PS oligonucleotide 647536 as compared to the conjugated full PS oligonucleotide 647535.
Example 58: Dose-Dependent Study of Oligonucleotides Comprising a 3'-Conjugate Group (Comparison of GalNAc.sub.3-1 and GalNAc.sub.4-11) Targeting SRB-1 In Vivo
[0926] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Unconjugated ISIS 440762 was included as an unconjugated standard. Each of the conjugate groups were attached at the 3' terminus of the respective oligonucleotide by a phosphodiester linked 2'-deoxyadenosine nucleoside cleavable moiety.
[0927] The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9. The structure of GalNAc.sub.3-11.sub.a was shown previously in Example 50.
Treatment
[0928] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 440762, 651900, 663748 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0929] As illustrated in Table 47, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. The antisense oligonucleotides comprising the phosphodiester linked GalNAc.sub.3-1 and GalNAc.sub.4-11 conjugates at the 3' terminus (ISIS 651900 and ISIS 663748) showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 440762). The two conjugated oligonucleotides, GalNAc.sub.3-1 and GalNAc.sub.4-11, were equipotent.
TABLE-US-00034 TABLE 47 Modified ASO targeting SRB-1 % Saline SEQ ID ASO Sequence (5' to 3') Dose mg/kg control No. Saline 100 ISIS 440762 T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub- .dsG.sub.dsA.sub.ds 0.6 73.45 2250 .sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k 2 59.66 6 23.50 ISIS 651900 T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub- .dsG.sub.dsA.sub.ds 0.2 62.75 2251 .sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.koA.sub.do'-GalNAc.sub.3-1.sub.- a 0.6 29.14 2 8.61 6 5.62 ISIS 663748 T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub- .dsG.sub.dsA.sub.ds 0.2 63.99 2251 .sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.koA.sub.do'-GalNAc.sub.4-11.sub- .a 0.6 33.53 2 7.58 6 5.52 Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "k" indicates 6'-(S)-CH.sub.3 bicyclic nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0930] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin and BUN were also evaluated. The change in body weights was evaluated with no significant change from the saline group. ALTs, ASTs, total bilirubin and BUN values are shown in Table 48 below.
TABLE-US-00035 TABLE 48 Dosage Total ISIS No. mg/kg ALT AST Bilirubin BUN Conjugate Saline 30 76 0.2 40 440762 0.60 32 70 0.1 35 none 2 26 57 0.1 35 6 31 48 0.1 39 651900 0.2 32 115 0.2 39 GalNac.sub.3-1 (3') 0.6 33 61 0.1 35 2 30 50 0.1 37 6 34 52 0.1 36 663748 0.2 28 56 0.2 36 GalNac.sub.4-11 (3') 0.6 34 60 0.1 35 2 44 62 0.1 36 6 38 71 0.1 33
Example 59: Effects of GalNAc.sub.3-1 Conjugated ASOs Targeting FXI In Vivo
[0931] The oligonucleotides listed below were tested in a multiple dose study for antisense inhibition of FXI in mice. ISIS 404071 was included as an unconjugated standard. Each of the conjugate groups was attached at the 3' terminus of the respective oligonucleotide by a phosphodiester linked 2'-deoxyadenosine nucleoside
TABLE-US-00036 TABLE 49 Modified ASOs targeting FXI SEQ ID ASO Sequence (5' to 3') Linkages No. ISIS T.sub.esG.sub.esG.sub.esT.sub.esA.sub.esA.sub.dsT.sub.ds.sup.mC.sub.d- s.sup.mC.sub.dsA.sub.ds.sup.mC.sub.ds PS 2259 404071 T.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.esG.sub.esA.sub.esG.sub- .esG.sub.e ISIS T.sub.esG.sub.esG.sub.esT.sub.esA.sub.esA.sub.dsT.sub.ds.sup.mC.sub.d- s.sup.mC.sub.dsA.sub.ds.sup.mC.sub.ds PS 2260 656172 T.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.esG.sub.esA.sub.esG.sub- .esG.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a ISIS T.sub.esG.sub.eoG.sub.eoT.sub.eoA.sub.eoA.sub.dsT.sub.ds.sup.mC.sub.d- s.sup.mC.sub.dsA.sub.ds.sup.mC.sub.ds PO/PS 2260 656173 T.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.eoG.sub.eoA.sub.esG.sub- .esG.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0932] The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9.
Treatment
[0933] Six week old male Balb/c mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously twice a week for 3 weeks at the dosage shown below with ISIS 404071, 656172, 656173 or with PBS treated control. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver FXI mRNA levels using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. Plasma FXI protein levels were also measured using ELISA. FXI mRNA levels were determined relative to total RNA (using RIBOGREEN.RTM.), prior to normalization to PBS-treated control. The results below are presented as the average percent of FXI mRNA levels for each treatment group. The data was normalized to PBS-treated control and is denoted as "% PBS". The ED.sub.50s were measured using similar methods as described previously and are presented below.
TABLE-US-00037 TABLE 50 Factor XI mRNA (% Saline) Dose ASO mg/kg % Control Conjugate Linkages Saline 100 none ISIS 3 92 none PS 404071 10 40 30 15 ISIS 0.7 74 GalNAc.sub.3-1 PS 656172 2 33 6 9 ISIS 0.7 49 GalNAc.sub.3-1 PO/PS 656173 2 22 6 1
[0934] As illustrated in Table 50, treatment with antisense oligonucleotides lowered FXI mRNA levels in a dose-dependent manner. The oligonucleotides comprising a 3'-GalNAc.sub.3-1 conjugate group showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 404071). Between the two conjugated oligonucleotides an improvement in potency was further provided by substituting some of the PS linkages with PO (ISIS 656173).
[0935] As illustrated in Table 50a, treatment with antisense oligonucleotides lowered FXI protein levels in a dose-dependent manner. The oligonucleotides comprising a 3'-GalNAc.sub.3-1 conjugate group showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 404071). Between the two conjugated oligonucleotides an improvement in potency was further provided by substituting some of the PS linkages with PO (ISIS 656173).
TABLE-US-00038 TABLE 50a Factor XI protein (% Saline) Dose Protein (% ASO mg/kg Control) Conjugate Linkages Saline 100 none ISIS 3 127 none PS 404071 10 32 30 3 ISIS 0.7 70 GalNAc.sub.3-1 PS 656172 2 23 6 1 ISIS 0.7 45 GalNAc.sub.3-1 PO/PS 656173 2 6 6 0
[0936] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin, total albumin, CRE and BUN were also evaluated. The change in body weights was evaluated with no significant change from the saline group. ALTs, ASTs, total bilirubin and BUN values are shown in the table below.
TABLE-US-00039 TABLE 51 ISIS No. Dosage mg/kg ALT AST Total Albumin Total Bilirubin CRE BUN Conjugate Saline 71.8 84.0 3.1 0.2 0.2 22.9 404071 3 152.8 176.0 3.1 0.3 0.2 23.0 none 10 73.3 121.5 3.0 0.2 0.2 21.4 30 82.5 92.3 3.0 0.2 0.2 23.0 656172 0.7 62.5 111.5 3.1 0.2 0.2 23.8 GalNac.sub.3-1 (3') 2 33.0 51.8 2.9 0.2 0.2 22.0 6 65.0 71.5 3.2 0.2 0.2 23.9 656173 0.7 54.8 90.5 3.0 0.2 0.2 24.9 2 85.8 71.5 3.2 0.2 0.2 21.0 GalNac.sub.3-1 (3') 6 114.0 101.8 3.3 0.2 0.2 22.7
Example 60: Effects of Conjugated ASOs Targeting SRB-1 In Vitro
[0937] The oligonucleotides listed below were tested in a multiple dose study for antisense inhibition of SRB-1 in primary mouse hepatocytes. ISIS 353382 was included as an unconjugated standard. Each of the conjugate groups were attached at the 3' or 5' terminus of the respective oligonucleotide by a phosphodiester linked 2'-deoxyadenosine nucleoside cleavable moiety.
TABLE-US-00040 TABLE 52 Modified ASO targeting SRB-1 SEQ ASO Sequence (5' to 3') Motif Conjugate ID No. ISIS 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 none 2256 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 655861 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 GalNAc.sub.3-1 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1.sub.a ISIS 655862 G.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 GalNAc.sub.3-1 2257 .sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1.sub.a ISIS 661161 GalNAc.sub.3-3.sub.a-o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup- .mC.sub.esA.sub.dsG.sub.ds 5/10/5 GalNAc.sub.3-3 2258 T.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 665001 GalNAc.sub.3-8.sub.a-o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup- .mC.sub.esA.sub.dsG.sub.ds 5/10/5 GalNAc.sub.3-8 2258 T.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 664078 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub- .ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds 5/10/5 GalNAc.sub.3-9 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-9.sub.a ISIS 666961 GalNAc.sub.3-6.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.ds 5/10/5 GalNAc.sub.3-6 2258 T.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 664507 GalNAc.sub.3-2.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-2 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666881 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.- es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-10 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666224 GalNAc.sub.3-5.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-5 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e ISIS 666981 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT.sub.e- s.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds 5/10/5 GalNAc.sub.3-7 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0938] The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9. The structure of GalNAc.sub.3-3a was shown previously in Example 39. The structure of GalNAc.sub.3-8a was shown previously in Example 47. The structure of GalNAc.sub.3-9a was shown previously in Example 52. The structure of GalNAc.sub.3-6a was shown previously in Example 51. The structure of GalNAc.sub.3-2a was shown previously in Example 37. The structure of GalNAc.sub.3-10a was shown previously in Example 46. The structure of GalNAc.sub.3-5a was shown previously in Example 49. The structure of GalNAc.sub.3-7a was shown previously in Example 48.
Treatment
[0939] The oligonucleotides listed above were tested in vitro in primary mouse hepatocyte cells plated at a density of 25,000 cells per well and treated with 0.03, 0.08, 0.24, 0.74, 2.22, 6.67 or 20 nM modified oligonucleotide. After a treatment period of approximately 16 hours, RNA was isolated from the cells and mRNA levels were measured by quantitative real-time PCR and the SRB-1 mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.RTM..
[0940] The IC.sub.50 was calculated using standard methods and the results are presented in Table 53. The results show that, under free uptake conditions in which no reagents or electroporation techniques are used to artificially promote entry of the oligonucleotides into cells, the oligonucleotides comprising a GalNAc conjugate were significantly more potent in hepatocytes than the parent oligonucleotide (ISIS 353382) that does not comprise a GalNAc conjugate.
TABLE-US-00041 TABLE 53 Internucleoside SEQ ID ASO IC.sub.50 (nM) linkages Conjugate No. ISIS 353382 190.sup.a PS none 2256 ISIS 655861 11.sup.a PS GalNAc.sub.3-1 2257 ISIS 655862 3 PO/PS GalNAc.sub.3-1 2257 ISIS 661161 15.sup.a PS GalNAc.sub.3-3 2258 ISIS 665001 20 PS GalNAc.sub.3-8 2258 ISIS 664078 55 PS GalNAc.sub.3-9 2257 ISIS 666961 22.sup.a PS GalNAc.sub.3-6 2258 ISIS 664507 30 PS GalNAc.sub.3-2 2258 ISIS 666881 30 PS GalNAc.sub.3-10 2258 ISIS 666224 30.sup.a PS GalNAc.sub.3-5 2258 ISIS 666981 40 PS GalNAc.sub.3-7 2258 .sup.aAverage of multiple runs.
Example 61: Preparation of Oligomeric Compound 175 Comprising GalNAc.sub.3-12
##STR00227## ##STR00228## ##STR00229##
[0941] Compound 169 is commercially available. Compound 172 was prepared by addition of benzyl (perfluorophenyl) glutarate to compound 171. The benzyl (perfluorophenyl) glutarate was prepared by adding PFP-TFA and DIEA to 5-(benzyloxy)-5-oxopentanoic acid in DMF. Oligomeric compound 175, comprising a GalNAc.sub.3-12 conjugate group, was prepared from compound 174 using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-12 (GalNAc.sub.3-12.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In a certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-12 (GalNAc.sub.3-12.sub.a-CM-) is shown below:
##STR00230##
Example 62: Preparation of Oligomeric Compound 180 Comprising GalNAc.sub.3-13
##STR00231## ##STR00232##
[0942] Compound 176 was prepared using the general procedure shown in Example 2. Oligomeric compound 180, comprising a GalNAc.sub.3-13 conjugate group, was prepared from compound 177 using the general procedures illustrated in Example 49. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-13 (GalNAc.sub.3-13.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In a certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-13 (GalNAc.sub.3-13.sub.a-CM-) is shown below:
##STR00233##
Example 63: Preparation of Oligomeric Compound 188 Comprising GalNAc.sub.3-14
##STR00234## ##STR00235##
[0943] Compounds 181 and 185 are commercially available. Oligomeric compound 188, comprising a GalNAc.sub.3-14 conjugate group, was prepared from compound 187 using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-14 (GalNAc.sub.3-14.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-14 (GalNAc.sub.3-14.sub.a-CM-) is shown below:
##STR00236##
Example 64: Preparation of Oligomeric Compound 197 Comprising GalNAc.sub.3-15
##STR00237## ##STR00238##
[0944] Compound 189 is commercially available. Compound 195 was prepared using the general procedure shown in Example 31. Oligomeric compound 197, comprising a GalNAc.sub.3-15 conjugate group, was prepared from compounds 194 and 195 using standard oligonucleotide synthesis procedures. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-15 (GalNAc.sub.3-15.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-15 (GalNAc.sub.3-15.sub.a-CM-) is shown below:
##STR00239##
Example 65: Dose-Dependent Study of Oligonucleotides Comprising a 5'-Conjugate Group (Comparison of GalNAc.sub.3-3, 12, 13, 14, and 15) Targeting SRB-1 In Vivo
[0945] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Unconjugated ISIS 353382 was included as a standard. Each of the GalNAc.sub.3 conjugate groups was attached at the 5' terminus of the respective oligonucleotide by a phosphodiester linked 2'-deoxyadenosine nucleoside (cleavable moiety).
TABLE-US-00042 TABLE 54 Modified ASOs targeting SRB-1 SEQ ISIS ID No. Sequences (5' to 3') Conjugate No. 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e none 2256 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- sG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.ds GalNAc.sub.3-3 2258 T.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 671144 GalNAc.sub.3-12.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.ds GalNAc.sub.3-12 2258 T.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 670061 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.ds GalNAc.sub.3-13 2258 T.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 671261 GalNAc.sub.3-14.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.ds GalNAc.sub.3-14 2258 T.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 671262 GalNAc.sub.3-15.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.ds GalNAc.sub.3-15 2258 T.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0946] The structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39. The structure of GalNAc.sub.3-12a was shown previously in Example 61. The structure of GalNAc.sub.3-13a was shown previously in Example 62. The structure of GalNAc.sub.3-14a was shown previously in Example 63. The structure of GalNAc.sub.3-15a was shown previously in Example 64.
Treatment
[0947] Six to eight week old C57bl6 mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once or twice at the dosage shown below with ISIS 353382, 661161, 671144, 670061, 671261, 671262, or with saline. Mice that were dosed twice received the second dose three days after the first dose. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0948] As illustrated in Table 55, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. No significant differences in target knockdown were observed between animals that received a single dose and animals that received two doses (see ISIS 353382 dosages 30 and 2.times.15 mg/kg; and ISIS 661161 dosages 5 and 2.times.2.5 mg/kg). The antisense oligonucleotides comprising the phosphodiester linked GalNAc.sub.3-3, 12, 13, 14, and 15 conjugates showed substantial improvement in potency compared to the unconjugated antisense oligonucleotide (ISIS 335382).
TABLE-US-00043 TABLE 55 SRB-1 mRNA (% Saline) Dosage SRB-1 mRNA ED.sub.50 ISIS No. (mg/kg) (% Saline) (mg/kg) Conjugate Saline n/a 100.0 n/a n/a 353382 3 85.0 22.4 none 10 69.2 30 34.2 2 .times. 15 36.0 661161 0.5 87.4 2.2 GalNAc.sub.3-3 1.5 59.0 5 25.6 2 .times. 2.5 27.5 15 17.4 671144 0.5 101.2 3.4 GalNAc.sub.3-12 1.5 76.1 5 32.0 15 17.6 670061 0.5 94.8 2.1 GalNAc.sub.3-13 1.5 57.8 5 20.7 15 13.3 671261 0.5 110.7 4.1 GalNAc.sub.3-14 1.5 81.9 5 39.8 15 14.1 671262 0.5 109.4 9.8 GalNAc.sub.3-15 1.5 99.5 5 69.2 15 36.1
[0949] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin and BUN were also evaluated. The changes in body weights were evaluated with no significant differences from the saline group (data not shown). ALTs, ASTs, total bilirubin and BUN values are shown in Table 56 below.
TABLE-US-00044 TABLE 56 Total Dosage ALT Bilirubin BUN ISIS No. (mg/kg) (U/L) AST (U/L) (mg/dL) (mg/dL) Conjugate Saline n/a 28 60 0.1 39 n/a 353382 3 30 77 0.2 36 none 10 25 78 0.2 36 30 28 62 0.2 35 2 .times. 15 22 59 0.2 33 661161 0.5 39 72 0.2 34 GalNAc.sub.3-3 1.5 26 50 0.2 33 5 41 80 0.2 32 2 .times. 2.5 24 72 0.2 28 15 32 69 0.2 36 671144 0.5 25 39 0.2 34 GalNAc.sub.3-12 1.5 26 55 0.2 28 5 48 82 0.2 34 15 23 46 0.2 32 670061 0.5 27 53 0.2 33 GalNAc.sub.3-13 1.5 24 45 0.2 35 5 23 58 0.1 34 15 24 72 0.1 31 671261 0.5 69 99 0.1 33 GalNAc.sub.3-14 1.5 34 62 0.1 33 5 43 73 0.1 32 15 32 53 0.2 30 671262 0.5 24 51 0.2 29 GalNAc.sub.3-15 1.5 32 62 0.1 31 5 30 76 0.2 32 15 31 64 0.1 32
Example 66: Effect of Various Cleavable Moieties on Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising a 5'-GalNAc.sub.3 Cluster
[0950] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Each of the GalNAc.sub.3 conjugate groups was attached at the 5' terminus of the respective oligonucleotide by a phosphodiester linked nucleoside (cleavable moiety (CM)).
TABLE-US-00045 TABLE 57 Modified ASOs targeting SRB-1 ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-3a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 670699 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.doG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-3a T.sub.d 2261 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.e- sT.sub.esT.sub.e 670700 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.eoG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-3a A.sub.e 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.e- sT.sub.esT.sub.e 670701 GalNAC.sub.3-3.sub.a-.sub.o'T.sub.eoG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-3a T.sub.e 2261 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.e- sT.sub.esT.sub.e 671165 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.eoT.sub.eo- T.sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- ds GalNAc.sub.3-13a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.e- sT.sub.esT.sub.e Capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0951] The structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39. The structure of GalNAc.sub.3-13a was shown previously in Example 62.
Treatment
[0952] Six to eight week old C57bl6 mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with ISIS 661161, 670699, 670700, 670701, 671165, or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the liver SRB-1 mRNA levels using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0953] As illustrated in Table 58, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. The antisense oligonucleotides comprising various cleavable moieties all showed similar potencies.
TABLE-US-00046 TABLE 58 SRB-1 mRNA (% Saline) Dosage SRB-1 mRNA GalNAc.sub.3 ISIS No. (mg/kg) (% Saline) Cluster CM Saline n/a 100.0 n/a n/a 661161 0.5 87.8 GalNAc.sub.3-3a A.sub.d 1.5 61.3 5 33.8 15 14.0 670699 0.5 89.4 GalNAc.sub.3-3a T.sub.d 1.5 59.4 5 31.3 15 17.1 670700 0.5 79.0 GalNAc.sub.3-3a A.sub.e 1.5 63.3 5 32.8 15 17.9 670701 0.5 79.1 GalNAc.sub.3-3a T.sub.e 1.5 59.2 5 35.8 15 17.7 671165 0.5 76.4 GalNAc.sub.3-13a A.sub.d 1.5 43.2 5 22.6 15 10.0
[0954] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin and BUN were also evaluated. The changes in body weights were evaluated with no significant differences from the saline group (data not shown). ALTs, ASTs, total bilirubin and BUN values are shown in Table 56 below.
TABLE-US-00047 TABLE 59 Total Dosage ALT AST Bilirubin BUN GalNAc.sub.3 ISIS No. (mg/kg) (U/L) (U/L) (mg/dL) (mg/dL) Cluster CM Saline n/a 24 64 0.2 31 n/a n/a 661161 0.5 25 64 0.2 31 GalNAc.sub.3-3a A.sub.d 1.5 24 50 0.2 32 5 26 55 0.2 28 15 27 52 0.2 31 670699 0.5 42 83 0.2 31 GalNAc.sub.3-3a T.sub.d 1.5 33 58 0.2 32 5 26 70 0.2 29 15 25 67 0.2 29 670700 0.5 40 74 0.2 27 GalNAc.sub.3-3a A.sub.e 1.5 23 62 0.2 27 5 24 49 0.2 29 15 25 87 0.1 25 670701 0.5 30 77 0.2 27 GalNAc.sub.3-3a T.sub.e 1.5 22 55 0.2 30 5 81 101 0.2 25 15 31 82 0.2 24 671165 0.5 44 84 0.2 26 GalNAc.sub.3-13a A.sub.d 1.5 47 71 0.1 24 5 33 91 0.2 26 15 33 56 0.2 29
Example 67: Preparation of Oligomeric Compound 199 Comprising GalNAc.sub.3-16
##STR00240##
[0956] Oligomeric compound 199, comprising a GalNAc.sub.3-16 conjugate group, is prepared using the general procedures illustrated in Examples 7 and 9. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-16 (GalNAc.sub.3-16.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-16 (GalNAc.sub.3-16.sub.a-CM-) is shown below:
##STR00241##
Example 68: Preparation of Oligomeric Compound 200 Comprising GalNAc.sub.3-17
##STR00242##
[0957] Oligomeric compound 200, comprising a GalNAc.sub.3-17 conjugate group, was prepared using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-17 (GalNAc.sub.3-17.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-17 (GalNAc.sub.3-17.sub.a-CM-) is shown below:
##STR00243##
Example 69: Preparation of Oligomeric Compound 201 Comprising GalNAc.sub.3-18
##STR00244##
[0958] Oligomeric compound 201, comprising a GalNAc.sub.3-18 conjugate group, was prepared using the general procedures illustrated in Example 46. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-18 (GalNAc.sub.3-18.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-18 (GalNAc.sub.3-18.sub.a-CM-) is shown below:
##STR00245##
Example 70: Preparation of Oligomeric Compound 204 Comprising GalNAc.sub.3-19
##STR00246##
[0959] Oligomeric compound 204, comprising a GalNAc.sub.3-19 conjugate group, was prepared from compound 64 using the general procedures illustrated in Example 52. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-19 (GalNAc.sub.3-19.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-19 (GalNAc.sub.3-19.sub.a-CM-) is shown below:
##STR00247##
Example 71: Preparation of Oligomeric Compound 210 Comprising GalNAc.sub.3-20
##STR00248## ##STR00249##
[0961] Compound 205 was prepared by adding PFP-TFA and DIEA to 6-(2,2,2-trifluoroacetamido)hexanoic acid in acetonitrile, which was prepared by adding triflic anhydride to 6-aminohexanoic acid. The reaction mixture was heated to 80.degree. C., then lowered to rt. Oligomeric compound 210, comprising a GalNAc.sub.3-20 conjugate group, was prepared from compound 208 using the general procedures illustrated in Example 52. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-20 (GalNAc.sub.3-20.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-20 (GalNAc.sub.3-20.sub.a-CM-) is shown below:
##STR00250##
Example 72: Preparation of Oligomeric Compound 215 Comprising GalNAc.sub.3-21
##STR00251## ##STR00252##
[0962] Compound 211 is commercially available. Oligomeric compound 215, comprising a GalNAc.sub.3-21 conjugate group, was prepared from compound 213 using the general procedures illustrated in Example 52. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-21 (GalNAc.sub.3-21a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-21 (GalNAc.sub.3-21.sub.a-CM-) is shown below:
##STR00253##
Example 73: Preparation of Oligomeric Compound 221 Comprising GalNAc.sub.3-22
##STR00254## ##STR00255##
[0963] Compound 220 was prepared from compound 219 using diisopropylammonium tetrazolide. Oligomeric compound 221, comprising a GalNAc.sub.3-21 conjugate group, is prepared from compound 220 using the general procedure illustrated in Example 52. The GalNAc.sub.3 cluster portion of the conjugate group GalNAc.sub.3-22 (GalNAc.sub.3-22.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the cleavable moiety is --P(.dbd.O)(OH)-A.sub.d-P(.dbd.O)(OH)--. The structure of GalNAc.sub.3-22 (GalNAc.sub.3-22.sub.a-CM-) is shown below:
##STR00256##
Example 74: Effect of Various Cleavable Moieties on Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising a 5'-GalNAc.sub.3 Conjugate
[0964] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice. Each of the GalNAc.sub.3 conjugate groups was attached at the 5' terminus of the respective oligonucleotide.
TABLE-US-00048 TABLE 60 Modified ASOs targeting SRB-1 ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es n/a n/a 2256 .sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-3a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 666904 GalNAc.sub.3-3.sub.a-.sub.o'G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.- sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.ds GalNAc.sub.3-3a PO 2256 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 675441 GalNAc.sub.3-17.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- ds GalNAc.sub.3-17a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 675442 GalNAc.sub.3-18.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsC.sub.dsA.sub.- dsT.sub.ds GalNAc.sub.3-18a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.es.sup.mC.sub.esT.sub.e- sT.sub.e In all tables, capital letters indicate the nucleobase for each nucleoside and .sup.mC indicates a 5-methyl cytosine. Subscripts: "e" indicates a 2'-MOE modified nucleoside; "d" indicates a .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); "o" indicates a phosphodiester internucleoside linkage (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Conjugate groups are in bold.
[0965] The structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39. The structure of GalNAc.sub.3-17a was shown previously in Example 68, and the structure of GalNAc.sub.3-18a was shown in Example 69.
Treatment
[0966] Six to eight week old C57BL/6 mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with an oligonucleotide listed in Table 60 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the SRB-1 mRNA levels using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0967] As illustrated in Table 61, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner. The antisense oligonucleotides comprising a GalNAc conjugate showed similar potencies and were significantly more potent than the parent oligonucleotide lacking a GalNAc conjugate.
TABLE-US-00049 TABLE 61 SRB-1 mRNA (% Saline) Dosage SRB-1 mRNA GalNAc.sub.3 ISIS No. (mg/kg) (% Saline) Cluster CM Saline n/a 100.0 n/a n/a 353382 3 79.38 n/a n/a 10 68.67 30 40.70 661161 0.5 79.18 GalNAc.sub.3-3a A.sub.d 1.5 75.96 5 30.53 15 12.52 666904 0.5 91.30 GalNAc.sub.3-3a PO 1.5 57.88 5 21.22 15 16.49 675441 0.5 76.71 GalNAc.sub.3-17a A.sub.d 1.5 63.63 5 29.57 15 13.49 675442 0.5 95.03 GalNAc.sub.3-18a A.sub.d 1.5 60.06 5 31.04 15 19.40
[0968] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were measured relative to saline injected mice using standard protocols. Total bilirubin and BUN were also evaluated. The change in body weights was evaluated with no significant change from the saline group (data not shown). ALTs, ASTs, total bilirubin and BUN values are shown in Table 62 below.
TABLE-US-00050 TABLE 62 Total Dosage ALT AST Bilirubin BUN GalNAc.sub.3 ISIS No. (mg/kg) (U/L) (U/L) (mg/dL) (mg/dL) Cluster CM Saline n/a 26 59 0.16 42 n/a n/a 353382 3 23 58 0.18 39 n/a n/a 10 28 58 0.16 43 30 20 48 0.12 34 661161 0.5 30 47 0.13 35 GalNAc.sub.3-3a A.sub.d 1.5 23 53 0.14 37 5 26 48 0.15 39 15 32 57 0.15 42 666904 0.5 24 73 0.13 36 GalNAc.sub.3-3a PO 1.5 21 48 0.12 32 5 19 49 0.14 33 15 20 52 0.15 26 675441 0.5 42 148 0.21 36 GalNAc.sub.3-17a A.sub.d 1.5 60 95 0.16 34 5 27 75 0.14 37 15 24 61 0.14 36 675442 0.5 26 65 0.15 37 GalNAc.sub.3-18a A.sub.d 1.5 25 64 0.15 43 5 27 69 0.15 37 15 30 84 0.14 37
Example 75: Pharmacokinetic Analysis of Oligonucleotides Comprising a 5'-Conjugate Group
[0969] The PK of the ASOs in Tables 54, 57 and 60 above was evaluated using liver samples that were obtained following the treatment procedures described in Examples 65, 66, and 74. The liver samples were minced and extracted using standard protocols and analyzed by IP-HPLC-MS alongside an internal standard. The combined tissue level (.mu.g/g) of all metabolites was measured by integrating the appropriate UV peaks, and the tissue level of the full-length ASO missing the conjugate ("parent," which is Isis No. 353382 in this case) was measured using the appropriate extracted ion chromatograms (EIC).
TABLE-US-00051 TABLE 63 PK Analysis in Liver Total Parent ASO Tissue Tissue Level Level Dosage by UV by EIC GalNAc.sub.3 ISIS No. (mg/kg) (.mu.g/g) (.mu.g/g) Cluster CM 353382 3 8.9 8.6 n/a n/a 10 22.4 21.0 30 54.2 44.2 661161 5 32.4 20.7 GalNAc.sub.3-3a A.sub.d 15 63.2 44.1 671144 5 20.5 19.2 GalNAc.sub.3-12a A.sub.d 15 48.6 41.5 670061 5 31.6 28.0 GalNAc.sub.3-13a A.sub.d 15 67.6 55.5 671261 5 19.8 16.8 GalNAc.sub.3-14a A.sub.d 15 64.7 49.1 671262 5 18.5 7.4 GalNAc.sub.3-15a A.sub.d 15 52.3 24.2 670699 5 16.4 10.4 GalNAc.sub.3-3a T.sub.d 15 31.5 22.5 670700 5 19.3 10.9 GalNAc.sub.3-3a A.sub.e 15 38.1 20.0 670701 5 21.8 8.8 GalNAc.sub.3-3a T.sub.e 15 35.2 16.1 671165 5 27.1 26.5 GalNAc.sub.3-13a A.sub.d 15 48.3 44.3 666904 5 30.8 24.0 GalNAc.sub.3-3a PO 15 52.6 37.6 675441 5 25.4 19.0 GalNAc.sub.3-17a A.sub.d 15 54.2 42.1 675442 5 22.2 20.7 GalNAc.sub.3-18a A.sub.d 15 39.6 29.0
[0970] The results in Table 63 above show that there were greater liver tissue levels of the oligonucleotides comprising a GalNAc.sub.3 conjugate group than of the parent oligonucleotide that does not comprise a GalNAc.sub.3 conjugate group (ISIS 353382) 72 hours following oligonucleotide administration, particularly when taking into consideration the differences in dosing between the oligonucleotides with and without a GalNAc.sub.3 conjugate group. Furthermore, by 72 hours, 40-98% of each oligonucleotide comprising a GalNAc.sub.3 conjugate group was metabolized to the parent compound, indicating that the GalNAc.sub.3 conjugate groups were cleaved from the oligonucleotides.
Example 76: Preparation of Oligomeric Compound 230 Comprising GalNAc.sub.3-23
##STR00257##
[0972] Compound 222 is commercially available. 44.48 ml (0.33 mol) of compound 222 was treated with tosyl chloride (25.39 g, 0.13 mol) in pyridine (500 mL) for 16 hours. The reaction was then evaporated to an oil, dissolved in EtOAc and washed with water, sat. NaHCO.sub.3, brine, and dried over Na.sub.2SO.sub.4. The ethyl acetate was concentrated to dryness and purified by column chromatography, eluted with EtOAc/hexanes (1:1) followed by 10% methanol in CH.sub.2Cl.sub.2 to give compound 223 as a colorless oil. LCMS and NMR were consistent with the structure. 10 g (32.86 mmol) of 1-Tosyltriethylene glycol (compound 223) was treated with sodium azide (10.68 g, 164.28 mmol) in DMSO (100 mL) at room temperature for 17 hours. The reaction mixture was then poured onto water, and extracted with EtOAc. The organic layer was washed with water three times and dried over Na.sub.2SO.sub.4. The organic layer was concentrated to dryness to give 5.3 g of compound 224 (92%). LCMS and NMR were consistent with the structure. 1-Azidotriethylene glycol (compound 224, 5.53 g, 23.69 mmol) and compound 4 (6 g, 18.22 mmol) were treated with 4 A molecular sieves (5 g), and TMSOTf (1.65 ml, 9.11 mmol) in dichloromethane (100 mL) under an inert atmosphere. After 14 hours, the reaction was filtered to remove the sieves, and the organic layer was washed with sat. NaHCO.sub.3, water, brine, and dried over Na.sub.2SO.sub.4. The organic layer was concentrated to dryness and purified by column chromatography, eluted with a gradient of 2 to 4% methanol in dichloromethane to give compound 225. LCMS and NMR were consistent with the structure. Compound 225 (11.9 g, 23.59 mmol) was hydrogenated in EtOAc/Methanol (4:1, 250 mL) over Pearlman's catalyst. After 8 hours, the catalyst was removed by filtration and the solvents removed to dryness to give compound 226. LCMS and NMR were consistent with the structure.
[0973] In order to generate compound 227, a solution of nitromethanetrispropionic acid (4.17 g, 15.04 mmol) and Hunig's base (10.3 ml, 60.17 mmol) in DMF (100 mL) were treated dropwise with pentaflourotrifluoro acetate (9.05 ml, 52.65 mmol). After 30 minutes, the reaction was poured onto ice water and extracted with EtOAc. The organic layer was washed with water, brine, and dried over Na.sub.2SO.sub.4. The organic layer was concentrated to dryness and then recrystallized from heptane to give compound 227 as a white solid. LCMS and NMR were consistent with the structure. Compound 227 (1.5 g, 1.93 mmol) and compound 226 (3.7 g, 7.74 mmol) were stirred at room temperature in acetonitrile (15 mL) for 2 hours. The reaction was then evaporated to dryness and purified by column chromatography, eluting with a gradient of 2 to 10% methanol in dichloromethane to give compound 228. LCMS and NMR were consistent with the structure. Compound 228 (1.7 g, 1.02 mmol) was treated with Raney Nickel (about 2 g wet) in ethanol (100 mL) in an atmosphere of hydrogen. After 12 hours, the catalyst was removed by filtration and the organic layer was evaporated to a solid that was used directly in the next step. LCMS and NMR were consistent with the structure. This solid (0.87 g, 0.53 mmol) was treated with benzylglutaric acid (0.18 g, 0.8 mmol), HBTU (0.3 g, 0.8 mmol) and DIEA (273.7 .mu.l, 1.6 mmol) in DMF (5 mL). After 16 hours, the DMF was removed under reduced pressure at 65.degree. C. to an oil, and the oil was dissolved in dichloromethane. The organic layer was washed with sat. NaHCO.sub.3, brine, and dried over Na.sub.2SO.sub.4. After evaporation of the organic layer, the compound was purified by column chromatography and eluted with a gradient of 2 to 20% methanol in dichloromethane to give the coupled product. LCMS and NMR were consistent with the structure. The benzyl ester was deprotected with Pearlman's catalyst under a hydrogen atmosphere for 1 hour. The catalyst was them removed by filtration and the solvents removed to dryness to give the acid. LCMS and NMR were consistent with the structure. The acid (486 mg, 0.27 mmol) was dissolved in dry DMF (3 mL). Pyridine (53.61 .mu.l, 0.66 mmol) was added and the reaction was purged with argon. Pentaflourotriflouro acetate (46.39 .mu.l, 0.4 mmol) was slowly added to the reaction mixture. The color of the reaction changed from pale yellow to burgundy, and gave off a light smoke which was blown away with a stream of argon. The reaction was allowed to stir at room temperature for one hour (completion of reaction was confirmed by LCMS). The solvent was removed under reduced pressure (rotovap) at 70.degree. C. The residue was diluted with DCM and washed with 1N NaHSO.sub.4, brine, saturated sodium bicarbonate and brine again. The organics were dried over Na.sub.2SO.sub.4, filtered, and were concentrated to dryness to give 225 mg of compound 229 as a brittle yellow foam. LCMS and NMR were consistent with the structure.
[0974] Oligomeric compound 230, comprising a GalNAc.sub.3-23 conjugate group, was prepared from compound 229 using the general procedure illustrated in Example 46. The GalNAc.sub.3 cluster portion of the GalNAc.sub.3-23 conjugate group (GalNAc.sub.3-23.sub.a) can be combined with any cleavable moiety to provide a variety of conjugate groups. The structure of GalNAc.sub.3-23 (GalNAc.sub.3-23.sub.a-CM) is shown below:
##STR00258##
Example 77: Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising a GalNAc.sub.3 Conjugate
[0975] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice.
TABLE-US-00052 TABLE 64 Modified ASOs targeting SRB-1 ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-3a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 666904 GalNAc.sub.3-3.sub.a-.sub.o'G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.- sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.ds GalNAc.sub.3-3a PO 2256 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 673502 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.eoT.sub.eo- T.sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- ds GalNAc.sub.3-10a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.e- sT.sub.esT.sub.e 677844 GalNAc.sub.3-9.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d- s GalNAc.sub.3-9a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 677843 GalNAc.sub.3-23.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.- ds GalNAc.sub.3-23a A.sub.d 2258 G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e- sT.sub.esT.sub.e 655861 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es GalNAc.sub.3-1a A.sub.d 2257 .sup.mC.sub.esT.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a 677841 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es GalNAc.sub.3-19a A.sub.d 2257 .sup.mC.sub.esT.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-19.sub.a 677842 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es GalNAc.sub.3-20a A.sub.d 2257 .sup.mC.sub.esT.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-20.sub.a The structure of GalNac.sub.3-1.sub.a was shown previously in Example 9, GalNAc.sub.3-3.sub.a was shown in Example 39, GalNAc.sub.3-9a was shown in Examble 52, GalNac.sub.3-10a was shown in Example 46, GalNAc.sub.3-19.sub.a was shown in Example 70, GalNAc.sub.3-20.sub.a was shown in Example 71, and GalNAc.sub.3-23.sub.a was shown in Example 76.
Treatment
[0976] Six to eight week old C57BL/6 mice (Jackson Laboratory, Bar Harbor, Me.) were each injected subcutaneously once at a dosage shown below with an oligonucleotide listed in Table 64 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration to determine the SRB-1 mRNA levels using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0977] As illustrated in Table 65, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner.
TABLE-US-00053 TABLE 65 SRB-1 mRNA (% Saline) Dosage SRB-1 mRNA GalNAc.sub.3 ISIS No. (mg/kg) (% Saline) Cluster CM Saline n/a 100.0 n/a n/a 661161 0.5 89.18 GalNAc.sub.3-3a A.sub.d 1.5 77.02 5 29.10 15 12.64 666904 0.5 93.11 GalNAc.sub.3-3a PO 1.5 55.85 5 21.29 15 13.43 673502 0.5 77.75 GalNAc.sub.3-10a A.sub.d 1.5 41.05 5 19.27 15 14.41 677844 0.5 87.65 GalNAc.sub.3-9a A.sub.d 1.5 93.04 5 40.77 15 16.95 677843 0.5 102.28 GalNAc.sub.3-23a A.sub.d 1.5 70.51 5 30.68 15 13.26 655861 0.5 79.72 GalNAc.sub.3-1a A.sub.d 1.5 55.48 5 26.99 15 17.58 677841 0.5 67.43 GalNAc.sub.3-19a A.sub.d 1.5 45.13 5 27.02 15 12.41 677842 0.5 64.13 GalNAc.sub.3-20a A.sub.d 1.5 53.56 5 20.47 15 10.23
[0978] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in serum were also measured using standard protocols. Total bilirubin and BUN were also evaluated. Changes in body weights were evaluated, with no significant change from the saline group (data not shown). ALTs, ASTs, total bilirubin and BUN values are shown in Table 66 below.
TABLE-US-00054 TABLE 66 Total Dosage ALT AST Bilirubin BUN GalNAc.sub.3 ISIS No. (mg/kg) (U/L) (U/L) (mg/dL) (mg/dL) Cluster CM Saline n/a 21 45 0.13 34 n/a n/a 661161 0.5 28 51 0.14 39 GalNAc.sub.3-3a A.sub.d 1.5 23 42 0.13 39 5 22 59 0.13 37 15 21 56 0.15 35 666904 0.5 24 56 0.14 37 GalNAc.sub.3-3a PO 1.5 26 68 0.15 35 5 23 77 0.14 34 15 24 60 0.13 35 673502 0.5 24 59 0.16 34 GalNAc.sub.3-10a A.sub.d 1.5 20 46 0.17 32 5 24 45 0.12 31 15 24 47 0.13 34 677844 0.5 25 61 0.14 37 GalNAc.sub.3-9a A.sub.d 1.5 23 64 0.17 33 5 25 58 0.13 35 15 22 65 0.14 34 677843 0.5 53 53 0.13 35 GalNAc.sub.3-23a A.sub.d 1.5 25 54 0.13 34 5 21 60 0.15 34 15 22 43 0.12 38 655861 0.5 21 48 0.15 33 GalNAc.sub.3-1a A.sub.d 1.5 28 54 0.12 35 5 22 60 0.13 36 15 21 55 0.17 30 677841 0.5 32 54 0.13 34 GalNAc.sub.3-19a A.sub.d 1.5 24 56 0.14 34 5 23 92 0.18 31 15 24 58 0.15 31 677842 0.5 23 61 0.15 35 GalNAc.sub.3-20a A.sub.d 1.5 24 57 0.14 34 5 41 62 0.15 35 15 24 37 0.14 32
Example 78: Antisense Inhibition In Vivo by Oligonucleotides Targeting Angiotensinogen Comprising a GalNAc.sub.3 Conjugate
[0979] The oligonucleotides listed below were tested in a dose-dependent study for antisense inhibition of Angiotensinogen (AGT) in normotensive Sprague Dawley rats.
TABLE-US-00055 TABLE 67 Modified ASOs targeting AGT ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 552668 .sup.mC.sub.esA.sub.es.sup.mC.sub.esT.sub.esG.sub.esA.sub.dsT.sub.d- sT.sub.dsT.sub.dsT.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.ds.sup.mC.sub.ds.sup.- mC.sub.dsA.sub.esG.sub.es n/a n/a 2262 G.sub.esA.sub.esT.sub.e 669509 .sup.mC.sub.esA.sub.es.sup.mC.sub.esT.sub.esG.sub.esA.sub.dsT.sub.d- sT.sub.dsT.sub.dsT.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.ds.sup.mC.sub.ds.sup.- mC.sub.dsA.sub.esG.sub.es GalNAc.sub.3-1.sub.a A.sub.d 2263 G.sub.esA.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a
The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9.
Treatment
[0980] Six week old, male Sprague Dawley rats were each injected subcutaneously once per week at a dosage shown below, for a total of three doses, with an oligonucleotide listed in Table 67 or with PBS. Each treatment group consisted of 4 animals. The rats were sacrificed 72 hours following the final dose. AGT liver mRNA levels were measured using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. AGT plasma protein levels were measured using the Total Angiotensinogen ELISA (Catalog # JP27412, IBL International, Toronto, ON) with plasma diluted 1:20,000. The results below are presented as the average percent of AGT mRNA levels in liver or AGT protein levels in plasma for each treatment group, normalized to the PBS control.
[0981] As illustrated in Table 68, treatment with antisense oligonucleotides lowered AGT liver mRNA and plasma protein levels in a dose-dependent manner, and the oligonucleotide comprising a GalNAc conjugate was significantly more potent than the parent oligonucleotide lacking a GalNAc conjugate.
TABLE-US-00056 TABLE 68 AGT liver mRNA and plasma protein levels AGT liver AGT plasma ISIS Dosage mRNA protein GalNAc.sub.3 No. (mg/kg) (% PBS) (% PBS) Cluster CM PBS n/a 100 100 n/a n/a 552668 3 95 122 n/a n/a 10 85 97 30 46 79 90 8 11 669509 0.3 95 70 GalNAc.sub.3-1a A.sub.d 1 95 129 3 62 97 10 9 23
[0982] Liver transaminase levels, alanine aminotransferase (ALT) and aspartate aminotransferase (AST), in plasma and body weights were also measured at time of sacrifice using standard protocols. The results are shown in Table 69 below.
TABLE-US-00057 TABLE 69 Liver transaminase levels and rat body weights Body Dosage ALT AST Weight (% GalNAc.sub.3 ISIS No. (mg/kg) (U/L) (U/L) of baseline) Cluster CM PBS n/a 51 81 186 n/a n/a 552668 3 54 93 183 n/a n/a 10 51 93 194 30 59 99 182 90 56 78 170 669509 0.3 53 90 190 GalNAc.sub.3-1a A.sub.d 1 51 93 192 3 48 85 189 10 56 95 189
Example 79: Duration of Action In Vivo of Oligonucleotides Targeting APOC-III Comprising a GalNAc.sub.3 Conjugate
[0983] The oligonucleotides listed in Table 70 below were tested in a single dose study for duration of action in mice.
TABLE-US-00058 TABLE 70 Modified ASOs targeting APOC-III ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 304801 A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.d- sT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.- mC.sub.dsT.sub.esT.sub.es n/a n/a 2248 T.sub.esA.sub.esT.sub.e 647535 A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.d- sT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.- mC.sub.dsT.sub.esT.sub.es GalNAc.sub.3-1a A.sub.d 2249 T.sub.esA.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a 663083 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doA.sub.esG.sub.es.sup.mC.sub.esT- .sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.d- s GalNAc.sub.3-3a A.sub.d 2264 .sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT.sub.esT.sub.esA.sub- .esT.sub.e 674449 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doA.sub.esG.sub.es.sup.mC.sub.esT- .sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.d- s GalNAc.sub.3-7a A.sub.d 2264 .sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT.sub.esT.sub.esA.sub- .esT.sub.e 674450 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doA.sub.esG.sub.es.sup.mC.sub.es- T.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.- ds GalNAc.sub.3-10a A.sub.d 2264 .sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT.sub.esT.sub.esA.sub- .esT.sub.e 674451 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doA.sub.esG.sub.es.sup.mC.sub.es- T.sub.esT.sub.es.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.- ds GalNAc.sub.3-13a A.sub.d 2264 .sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.esT.sub.esT.sub.esA.sub- .esT.sub.e
The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9, GalNAc.sub.3-3.sub.a was shown in Example 39, GalNAc.sub.3-7.sub.a was shown in Example 48, GalNAc.sub.3-10.sub.a was shown in Example 46, and GalNAc.sub.3-13.sub.a was shown in Example 62.
Treatment
[0984] Six to eight week old transgenic mice that express human APOC-III were each injected subcutaneously once with an oligonucleotide listed in Table 70 or with PBS. Each treatment group consisted of 3 animals. Blood was drawn before dosing to determine baseline and at 72 hours, 1 week, 2 weeks, 3 weeks, 4 weeks, 5 weeks, and 6 weeks following the dose. Plasma triglyceride and APOC-III protein levels were measured as described in Example 20. The results below are presented as the average percent of plasma triglyceride and APOC-III levels for each treatment group, normalized to baseline levels, showing that the oligonucleotides comprising a GalNAc conjugate group exhibited a longer duration of action than the parent oligonucleotide without a conjugate group (ISIS 304801) even though the dosage of the parent was three times the dosage of the oligonucleotides comprising a GalNAc conjugate group.
TABLE-US-00059 TABLE 71 Plasma triglyceride and APOC-III protein levels in transgenic mice Time point Tri- (days glycerides APOC-III ISIS Dosage post- (% protein (% GalNAc.sub.3 No. (mg/kg) dose) baseline) baseline) Cluster CM PBS n/a 3 97 102 n/a n/a 7 101 98 14 108 98 21 107 107 28 94 91 35 88 90 42 91 105 304801 30 3 40 34 n/a n/a 7 41 37 14 50 57 21 50 50 28 57 73 35 68 70 42 75 93 647535 10 3 36 37 GalNAc.sub.3-1a A.sub.d 7 39 47 14 40 45 21 41 41 28 42 62 35 69 69 42 85 102 663083 10 3 24 18 GalNAc.sub.3-3a A.sub.d 7 28 23 14 25 27 21 28 28 28 37 44 35 55 57 42 60 78 674449 10 3 29 26 GalNAc.sub.3-7a A.sub.d 7 32 31 14 38 41 21 44 44 28 53 63 35 69 77 42 78 99 674450 10 3 33 30 GalNAc.sub.3-10a A.sub.d 7 35 34 14 31 34 21 44 44 28 56 61 35 68 70 42 83 95 674451 10 3 35 33 GalNAc.sub.3-13a A.sub.d 7 24 32 14 40 34 21 48 48 28 54 67 35 65 75 42 74 97
Example 80: Antisense Inhibition In Vivo by Oligonucleotides Targeting Alpha-1 Antitrypsin (A1AT) Comprising a GalNAc.sub.3 Conjugate
[0985] The oligonucleotides listed in Table 72 below were tested in a study for dose-dependent inhibition of A1AT in mice.
TABLE-US-00060 TABLE 72 Modified ASOs targeting A1AT ISIS GalNAc.sub.3 SEQ ID No. Sequences (5' to 3') Cluster CM No. 476366 A.sub.es.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.esA.sub.esA.sub.dsT- .sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.dsA.sub.dsA.sub.dsG.sub.dsG.sub- .dsA.sub.esA.sub.es n/a n/a 2265 G.sub.esG.sub.esA.sub.e 656326 A.sub.es.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.esA.sub.esA.sub.dsT- .sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.dsA.sub.dsA.sub.dsG.sub.dsG.sub- .dsA.sub.esA.sub.es GalNAc.sub.3-1a A.sub.d 2266 G.sub.esG.sub.esA.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a 678381 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doA.sub.es.sup.mC.sub.es.sup.mC.s- ub.es.sup.mC.sub.esA.sub.esA.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsG- .sub.dsA.sub.ds GalNAc.sub.3-3a A.sub.d 2267 A.sub.dsG.sub.dsG.sub.dsA.sub.esA.sub.esG.sub.esG.sub.esA.sub.e 678382 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doA.sub.es.sup.mC.sub.es.sup.mC.s- ub.es.sup.mC.sub.esA.sub.esA.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsG- .sub.dsA.sub.ds GalNAc.sub.3-7a A.sub.d 2267 A.sub.dsG.sub.dsG.sub.dsA.sub.esA.sub.esG.sub.esG.sub.esA.sub.e 678383 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doA.sub.es.sup.mC.sub.es.sup.mC.- sub.es.sup.mC.sub.esA.sub.esA.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.ds- G.sub.ds GalNAc.sub.3-10a A.sub.d 2267 A.sub.dsA.sub.dsG.sub.dsG.sub.dsA.sub.esA.sub.esG.sub.esG.sub.esA.sub.e 678384 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doA.sub.es.sup.mC.sub.es.sup.mC.- sub.es.sup.mC.sub.esA.sub.esA.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.ds- G.sub.ds GalNAc.sub.3-13a A.sub.d 2267 A.sub.dsA.sub.dsG.sub.dsG.sub.dsA.sub.esA.sub.esG.sub.esG.sub.esA.sub.e
The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9, GalNAc.sub.3-3.sub.a was shown in Example 39, GalNAc.sub.3-7.sub.a was shown in Example 48, GalNAc.sub.3-10.sub.a was shown in Example 46, and GalNAc.sub.3-13.sub.a was shown in Example 62.
Treatment
[0986] Six week old, male C57BL/6 mice (Jackson Laboratory, Bar Harbor, Me.) were each injected subcutaneously once per week at a dosage shown below, for a total of three doses, with an oligonucleotide listed in Table 72 or with PBS. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration. A1AT liver mRNA levels were determined using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. A1AT plasma protein levels were determined using the Mouse Alpha 1-Antitrypsin ELISA (catalog #41-A1AMS-E01, Alpco, Salem, N.H.). The results below are presented as the average percent of A1AT liver mRNA and plasma protein levels for each treatment group, normalized to the PBS control.
[0987] As illustrated in Table 73, treatment with antisense oligonucleotides lowered A1AT liver mRNA and A1AT plasma protein levels in a dose-dependent manner. The oligonucleotides comprising a GalNAc conjugate were significantly more potent than the parent (ISIS 476366).
TABLE-US-00061 TABLE 73 A1AT liver mRNA and plasma protein levels A1AT liver A1AT plasma ISIS Dosage mRNA protein GalNAc.sub.3 No. (mg/kg) (% PBS) (% PBS) Cluster CM PBS n/a 100 100 n/a n/a 476366 5 86 78 15 73 61 n/a n/a 45 30 38 656326 0.6 99 90 GalNAc.sub.3-1a A.sub.d 2 61 70 6 15 30 18 6 10 678381 0.6 105 90 GalNAc.sub.3-3a A.sub.d 2 53 60 6 16 20 18 7 13 678382 0.6 90 79 GalNAc.sub.3-7a A.sub.d 2 49 57 6 21 27 18 8 11 678383 0.6 94 84 GalNAc.sub.3-10a A.sub.d 2 44 53 6 13 24 18 6 10 678384 0.6 106 91 GalNAc.sub.3-13a A.sub.d 2 65 59 6 26 31 18 11 15
[0988] Liver transaminase and BUN levels in plasma were measured at time of sacrifice using standard protocols. Body weights and organ weights were also measured. The results are shown in Table 74 below. Body weight is shown as % relative to baseline. Organ weights are shown as % of body weight relative to the PBS control group.
TABLE-US-00062 TABLE 74 Body Liver Kidney Spleen ISIS Dosage ALT AST BUN weight (% weight (Rel weight (Rel weight (Rel No. (mg/kg) (U/L) (U/L) (mg/dL) baseline) % BW) % BW) % BW) PBS n/a 25 51 37 119 100 100 100 476366 5 34 68 35 116 91 98 106 15 37 74 30 122 92 101 128 45 30 47 31 118 99 108 123 656326 0.6 29 57 40 123 100 103 119 2 36 75 39 114 98 111 106 6 32 67 39 125 99 97 122 18 46 77 36 116 102 109 101 678381 0.6 26 57 32 117 93 109 110 2 26 52 33 121 96 106 125 6 40 78 32 124 92 106 126 18 31 54 28 118 94 103 120 678382 0.6 26 42 35 114 100 103 103 2 25 50 31 117 91 104 117 6 30 79 29 117 89 102 107 18 65 112 31 120 89 104 113 678383 0.6 30 67 38 121 91 100 123 2 33 53 33 118 98 102 121 6 32 63 32 117 97 105 105 18 36 68 31 118 99 103 108 678384 0.6 36 63 31 118 98 103 98 2 32 61 32 119 93 102 114 6 34 69 34 122 100 100 96 18 28 54 30 117 98 101 104
Example 81: Duration of Action In Vivo of Oligonucleotides Targeting A1AT Comprising a GalNAc.sub.3 Cluster
[0989] The oligonucleotides listed in Table 72 were tested in a single dose study for duration of action in mice.
Treatment
[0990] Six week old, male C57BL/6 mice were each injected subcutaneously once with an oligonucleotide listed in Table 72 or with PBS. Each treatment group consisted of 4 animals. Blood was drawn the day before dosing to determine baseline and at 5, 12, 19, and 25 days following the dose. Plasma A1AT protein levels were measured via ELISA (see Example 80). The results below are presented as the average percent of plasma A1AT protein levels for each treatment group, normalized to baseline levels. The results show that the oligonucleotides comprising a GalNAc conjugate were more potent and had longer duration of action than the parent lacking a GalNAc conjugate (ISIS 476366). Furthermore, the oligonucleotides comprising a 5'-GalNAc conjugate (ISIS 678381, 678382, 678383, and 678384) were generally even more potent with even longer duration of action than the oligonucleotide comprising a 3'-GalNAc conjugate (ISIS 656326).
TABLE-US-00063 TABLE 75 Plasma A1AT protein levels in mice Time point ISIS Dosage (days post- A1AT (% GalNAc.sub.3 No. (mg/kg) dose) baseline) Cluster CM PBS n/a 5 93 n/a n/a 12 93 19 90 25 97 476366 100 5 38 n/a n/a 12 46 19 62 25 77 656326 18 5 33 GalNAc.sub.3-1a A.sub.d 12 36 19 51 25 72 678381 18 5 21 GalNAc.sub.3-3a A.sub.d 12 21 19 35 25 48 678382 18 5 21 GalNAc.sub.3-7a A.sub.d 12 21 19 39 25 60 678383 18 5 24 GalNAc.sub.3-10a A.sub.d 12 21 19 45 25 73 678384 18 5 29 GalNAc.sub.3-13a A.sub.d 12 34 19 57 25 76
Example 82: Antisense Inhibition In Vitro by Oligonucleotides Targeting SRB-1 Comprising a GalNAc.sub.3 Conjugate
[0991] Primary mouse liver hepatocytes were seeded in 96 well plates at 15,000 cells/well 2 hours prior to treatment. The oligonucleotides listed in Table 76 were added at 2, 10, 50, or 250 nM in Williams E medium and cells were incubated overnight at 37.degree. C. in 5% CO.sub.2. Cells were lysed 16 hours following oligonucleotide addition, and total RNA was purified using RNease 3000 BioRobot (Qiagen). SRB-1 mRNA levels were determined using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. IC.sub.50 values were determined using Prism 4 software (GraphPad). The results show that oligonucleotides comprising a variety of different GalNAc conjugate groups and a variety of different cleavable moieties are significantly more potent in an in vitro free uptake experiment than the parent oligonucleotides lacking a GalNAc conjugate group (ISIS 353382 and 666841).
TABLE-US-00064 TABLE 76 Inhibition of SRB-1 expression in vitro ISIS GalNAc IC.sub.50 SEQ No. Sequence (5' to 3') Linkages cluster CM (nM) ID No. 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds PS n/a n/a 250 2256 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 655861 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds PS GalNAc.sub.3- A.sub.d 40 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-1a 1.sub.a 661161 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 40 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 3.sub.a 661162 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- A.sub.d 8 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e 3.sub.a 664078 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds PS GalNAc.sub.3- A.sub.d 20 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-9.sub.a 9.sub.a 665001 GalNAc.sub.3-8.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 70 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 8.sub.a 666224 GalNAc.sub.3-5.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 80 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 5.sub.a 666841 G.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds PO/PS n/a n/a >250 2256 .sup.mC.sub.dsT.sub.dsT.sub.eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e 666881 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 30 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 10.sub.a 666904 GalNAc.sub.3-3.sub.a-.sub.o'G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.- sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds PS GalNAc.sub.3- PO 9 2256 A.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub- .es.sup.mC.sub.esT.sub.esT.sub.e 3.sub.a 666924 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- T.sub.d 15 2261 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 3.sub.a 666961 GalNAc.sub.3-6.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 150 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 6.sub.a 666981 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.esT- .sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 20 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 7.sub.a 670061 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 30 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 13.sub.a 670699 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.doG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- T.sub.d 15 2261 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e 3.sub.a 670700 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.eoG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- A.sub.e 30 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT 3.sub.a 670701 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.eoG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- T.sub.e 25 2261 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e 3.sub.a 671144 GalNAc.sub.3-12.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 40 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 12.sub.a 671165 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.eoT.sub.eo- T.sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- A.sub.d 8 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT 13.sub.a 671261 GalNAc.sub.3-14.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d >250 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 14.sub.a 671262 GalNAc.sub.3-15.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d >250 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 15.sub.a 673501 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.eoT.sub.eoT- .sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- A.sub.d 30 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e 7.sub.a 673502 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.eoT.sub.eo- T.sub.eo.sup.mC.sub.eoA.sub.dsG.sub.dsT.sub.ds PO/PS GalNAc.sub.3- A.sub.d 8 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .eo.sup.mC.sub.eo.sup.mC.sub.esT.sub.esT.sub.e 10.sub.a 675441 GalNAc.sub.3-17.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 30 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 17.sub.a 675442 GalNAc.sub.3-18.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 20 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 18.sub.a 677841 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds PS GalNAc.sub.3- A.sub.d 40 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-19.sub.a 19.sub.a 677842 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds PS GalNAc.sub.3- A.sub.d 30 2257 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e- oA.sub.do'-GalNAc.sub.3-20.sub.a 20.sub.a 677843 GalNAc.sub.3-23.sub.a-.sub.o'A.sub.doG.sub.es.sup.mC.sub.esT.sub.es- T.sub.es.sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds PS GalNAc.sub.3- A.sub.d 40 2258 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub- .es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 23.sub.a
The structure of GalNAc.sub.3-1a was shown previously in Example 9, GalNAc.sub.3-3.sub.a was shown in Example 39, GalNAc.sub.3-5.sub.a was shown in Example 49, GalNAc.sub.3-6.sub.a was shown in Example 51, GalNAc.sub.3-7.sub.a was shown in Example 48, GalNAc.sub.3-8.sub.a was shown in Example 47, GalNAc.sub.3-9.sub.a was shown in Example 52, GalNAc.sub.3-10.sub.a was shown in Example 46, GalNAc.sub.3-12.sub.a was shown in Example 61, GalNAc.sub.3-13.sub.a was shown in Example 62, GalNAc.sub.3-14.sub.a was shown in Example 63, GalNAc.sub.3-15.sub.a was shown in Example 64, GalNAc.sub.3-17.sub.a was shown in Example 68, GalNAc.sub.3-18.sub.a was shown in Example 69, GalNAc.sub.3-19.sub.a was shown in Example 70, GalNAc.sub.3-20.sub.a was shown in Example 71, and GalNAc.sub.3-23.sub.a was shown in Example 76.
Example 83: Antisense Inhibition In Vivo by Oligonucleotides Targeting Factor XI Comprising a GalNAc.sub.3 Cluster
[0992] The oligonucleotides listed in Table 77 below were tested in a study for dose-dependent inhibition of Factor XI in mice.
TABLE-US-00065 TABLE 77 Modified oligonucleotides targeting Factor XI ISIS GalNAc SEQ No. Sequence (5' to 3') cluster CM ID No. 404071 T.sub.esG.sub.esG.sub.esT.sub.esA.sub.esA.sub.dsT.sub.ds.sup.mC.sub- .ds.sup.mC.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsT.sub.ds.sup.mC.sub- .dsA.sub.esG.sub.es n/a n/a 2259 A.sub.esG.sub.esGe 656173 T.sub.esG.sub.eoG.sub.eoT.sub.eoA.sub.eoA.sub.dsT.sub.ds.sup.mC.sub- .ds.sup.mC.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsT.sub.ds.sup.mC.sub- .dsA.sub.eoG.sub.eo GalNAc.sub.3-1.sub.a A.sub.d 2260 A.sub.esG.sub.esG.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a 663086 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doT.sub.esG.sub.eoG.sub.eoT.sub.e- oA.sub.eoA.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.ds.sup.mC.sub.d- sT.sub.ds GalNAc.sub.3-3.sub.a A.sub.d 2268 T.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.eoG.sub.eoA.sub.esG.sub.esG.sub.e 678347 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.doT.sub.esG.sub.eoG.sub.eoT.sub.e- oA.sub.eoA.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.ds.sup.mC.sub.d- sT.sub.ds GalNAc.sub.3-7.sub.a A.sub.d 2268 T.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.eoG.sub.eoA.sub.esG.sub.esG.sub.e 678348 GalNAc.sub.3-10.sub.a-.sub.o'A.sub.doT.sub.esG.sub.eoG.sub.eoT.sub.- eoA.sub.eoA.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.ds.sup.mC.sub.- ds GalNAc.sub.3-10.sub.a A.sub.d 2268 T.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.eoG.sub.eoA.sub.esG.sub.esG.s- ub.e 678349 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doT.sub.esG.sub.eoG.sub.eoT.sub.- eoA.sub.eoA.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.ds.sup.mC.sub.- ds GalNAc.sub.3-13.sub.a A.sub.d 2268 T.sub.dsT.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.eoG.sub.eoA.sub.esG.sub.esG.s- ub.e
The structure of GalNAc.sub.3-1a was shown previously in Example 9, GalNAc.sub.3-3.sub.a was shown in Example 39, GalNAc.sub.3-7.sub.a was shown in Example 48, GalNAc.sub.3-10.sub.a was shown in Example 46, and GalNAc.sub.3-13.sub.a was shown in Example 62.
Treatment
[0993] Six to eight week old mice were each injected subcutaneously once per week at a dosage shown below, for a total of three doses, with an oligonucleotide listed below or with PBS. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final dose. Factor XI liver mRNA levels were measured using real-time PCR and normalized to cyclophilin according to standard protocols. Liver transaminases, BUN, and bilirubin were also measured. The results below are presented as the average percent for each treatment group, normalized to the PBS control.
[0994] As illustrated in Table 78, treatment with antisense oligonucleotides lowered Factor XI liver mRNA in a dose-dependent manner. The results show that the oligonucleotides comprising a GalNAc conjugate were more potent than the parent lacking a GalNAc conjugate (ISIS 404071). Furthermore, the oligonucleotides comprising a 5'-GalNAc conjugate (ISIS 663086, 678347, 678348, and 678349) were even more potent than the oligonucleotide comprising a 3'-GalNAc conjugate (ISIS 656173).
TABLE-US-00066 TABLE 78 Factor XI liver mRNA, liver transaminase, BUN, and bilirubin levels ISIS Dosage Factor XI ALT AST BUN Bilirubin GalNAc.sub.3 SEQ No. (mg/kg) mRNA (% PBS) (U/L) (U/L) (mg/dL) (mg/dL) Cluster ID No. PBS n/a 100 63 70 21 0.18 n/a n/a 404071 3 65 41 58 21 0.15 n/a 2259 10 33 49 53 23 0.15 30 17 43 57 22 0.14 656173 0.7 43 90 89 21 0.16 GalNAc.sub.3-1a 2260 2 9 36 58 26 0.17 6 3 50 63 25 0.15 663086 0.7 33 91 169 25 0.16 GalNAc.sub.3-3a 2268 2 7 38 55 21 0.16 6 1 34 40 23 0.14 678347 0.7 35 28 49 20 0.14 GalNAc.sub.3-7a 2268 2 10 180 149 21 0.18 6 1 44 76 19 0.15 678348 0.7 39 43 54 21 0.16 GalNAc.sub.3-10a 2268 2 5 38 55 22 0.17 6 2 25 38 20 0.14 678349 0.7 34 39 46 20 0.16 GalNAc.sub.3-13a 2268 2 8 43 63 21 0.14 6 2 28 41 20 0.14
Example 84: Duration of Action In Vivo of Oligonucleotides Targeting Factor XI Comprising a GalNAc.sub.3 Conjugate
[0995] The oligonucleotides listed in Table 77 were tested in a single dose study for duration of action in mice.
Treatment
[0996] Six to eight week old mice were each injected subcutaneously once with an oligonucleotide listed in Table 77 or with PBS. Each treatment group consisted of 4 animals. Blood was drawn by tail bleeds the day before dosing to determine baseline and at 3, 10, and 17 days following the dose. Plasma Factor XI protein levels were measured by ELISA using Factor XI capture and biotinylated detection antibodies from R & D Systems, Minneapolis, Minn. (catalog # AF2460 and # BAF2460, respectively) and the OptEIA Reagent Set B (Catalog #550534, BD Biosciences, San Jose, Calif.). The results below are presented as the average percent of plasma Factor XI protein levels for each treatment group, normalized to baseline levels. The results show that the oligonucleotides comprising a GalNAc conjugate were more potent with longer duration of action than the parent lacking a GalNAc conjugate (ISIS 404071). Furthermore, the oligonucleotides comprising a 5'-GalNAc conjugate (ISIS 663086, 678347, 678348, and 678349) were even more potent with an even longer duration of action than the oligonucleotide comprising a 3'-GalNAc conjugate (ISIS 656173).
TABLE-US-00067 TABLE 79 Plasma Factor XI protein levels in mice Time point Factor SEQ ISIS Dosage (days XI (% GalNAc.sub.3 ID No. (mg/kg) post-dose) baseline) Cluster CM No. PBS n/a 3 123 n/a n/a n/a 10 56 17 100 404071 30 3 11 n/a n/a 2259 10 47 17 52 656173 6 3 1 GalNAc.sub.3-1a A.sub.d 2260 10 3 17 21 663086 6 3 1 GalNAc.sub.3-3a A.sub.d 2268 10 2 17 9 678347 6 3 1 GalNAc.sub.3-7a A.sub.d 2268 10 1 17 8 678348 6 3 1 GalNAc.sub.3-10a A.sub.d 2268 10 1 17 6 678349 6 3 1 GalNAc.sub.3-13a A.sub.d 2268 10 1 17 5
Example 85: Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising a GalNAc.sub.3 Conjugate
[0997] Oligonucleotides listed in Table 76 were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice.
Treatment
[0998] Six to eight week old C57BL/6 mice were each injected subcutaneously once per week at a dosage shown below, for a total of three doses, with an oligonucleotide listed in Table 76 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 48 hours following the final administration to determine the SRB-1 mRNA levels using real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. The results below are presented as the average percent of liver SRB-1 mRNA levels for each treatment group, normalized to the saline control.
[0999] As illustrated in Tables 80 and 81, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner.
TABLE-US-00068 TABLE 80 SRB-1 mRNA in liver Dosage SRB-1 mRNA GalNAc.sub.3 ISIS No. (mg/kg) (% Saline) Cluster CM Saline n/a 100 n/a n/a 655861 0.1 94 GalNAc.sub.3-1a A.sub.d 0.3 119 1 68 3 32 661161 0.1 120 GalNAc.sub.3-3a A.sub.d 0.3 107 1 68 3 26 666881 0.1 107 GalNAc.sub.3-10a A.sub.d 0.3 107 1 69 3 27 666981 0.1 120 GalNAc.sub.3-7a A.sub.d 0.3 103 1 54 3 21 670061 0.1 118 GalNAc.sub.3-13a A.sub.d 0.3 89 1 52 3 18 677842 0.1 119 GalNAc.sub.3-20a A.sub.d 0.3 96 1 65 3 23
TABLE-US-00069 TABLE 81 SRB-1 mRNA in liver Dosage SRB-1 mRNA GalNAc.sub.3 ISIS No. (mg/kg) (% Saline) Cluster CM 661161 0.1 107 GalNAc.sub.3-3a A.sub.d 0.3 95 1 53 3 18 677841 0.1 110 GalNAc.sub.3-19a A.sub.d 0.3 88 1 52 3 25
[1000] Liver transaminase levels, total bilirubin, BUN, and body weights were also measured using standard protocols. Average values for each treatment group are shown in Table 82 below.
TABLE-US-00070 TABLE 82 ISIS Dosage ALT AST Bilirubin BUN Body Weight GalNAc.sub.3 No. (mg/kg) (U/L) (U/L) (mg/dL) (mg/dL) (% baseline) Cluster CM Saline n/a 19 39 0.17 26 118 n/a n/a 655861 0.1 25 47 0.17 27 114 GalNAc.sub.3-1a A.sub.d 0.3 29 56 0.15 27 118 1 20 32 0.14 24 112 3 27 54 0.14 24 115 661161 0.1 35 83 0.13 24 113 GalNAc.sub.3-3a A.sub.d 0.3 42 61 0.15 23 117 1 34 60 0.18 22 116 3 29 52 0.13 25 117 666881 0.1 30 51 0.15 23 118 GalNAc.sub.3-10a A.sub.d 0.3 49 82 0.16 25 119 1 23 45 0.14 24 117 3 20 38 0.15 21 112 666981 0.1 21 41 0.14 22 113 GalNAc.sub.3-7a A.sub.d 0.3 29 49 0.16 24 112 1 19 34 0.15 22 111 3 77 78 0.18 25 115 670061 0.1 20 63 0.18 24 111 GalNAc.sub.3-13a A.sub.d 0.3 20 57 0.15 21 115 1 20 35 0.14 20 115 3 27 42 0.12 20 116 677842 0.1 20 38 0.17 24 114 GalNAc.sub.3-20a A.sub.d 0.3 31 46 0.17 21 117 1 22 34 0.15 21 119 3 41 57 0.14 23 118
Example 86: Antisense Inhibition In Vivo by Oligonucleotides Targeting TTR Comprising a GalNAc.sub.3 Cluster
[1001] Oligonucleotides listed in Table 83 below were tested in a dose-dependent study for antisense inhibition of human transthyretin (TTR) in transgenic mice that express the human TTR gene.
Treatment
[1002] Eight week old TTR transgenic mice were each injected subcutaneously once per week for three weeks, for a total of three doses, with an oligonucleotide and dosage listed in the tables below or with PBS. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration. Tail bleeds were performed at various time points throughout the experiment, and plasma TTR protein, ALT, and AST levels were measured and reported in Tables 85-87. After the animals were sacrificed, plasma ALT, AST, and human TTR levels were measured, as were body weights, organ weights, and liver human TTR mRNA levels. TTR protein levels were measured using a clinical analyzer (AU480, Beckman Coulter, CA). Real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) were used according to standard protocols to determine liver human TTR mRNA levels. The results presented in Tables 84-87 are the average values for each treatment group. The mRNA levels are the average values relative to the average for the PBS group. Plasma protein levels are the average values relative to the average value for the PBS group at baseline. Body weights are the average percent weight change from baseline until sacrifice for each individual treatment group. Organ weights shown are normalized to the animal's body weight, and the average normalized organ weight for each treatment group is then presented relative to the average normalized organ weight for the PBS group.
[1003] In Tables 84-87, "BL" indicates baseline, measurements that were taken just prior to the first dose. As illustrated in Tables 84 and 85, treatment with antisense oligonucleotides lowered TTR expression levels in a dose-dependent manner. The oligonucleotides comprising a GalNAc conjugate were more potent than the parent lacking a GalNAc conjugate (ISIS 420915). Furthermore, the oligonucleotides comprising a GalNAc conjugate and mixed PS/PO internucleoside linkages were even more potent than the oligonucleotide comprising a GalNAc conjugate and full PS linkages.
TABLE-US-00071 TABLE 83 Oligonucleotides targeting human TTR GalNAc SEQ Isis No. Sequence 5' to 3' Linkages cluster CM ID No. 420915 T.sub.es.sup.mC.sub.esT.sub.esT.sub.esG.sub.esG.sub.dsT.sub.dsT.sub- .dsA.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.ds PS n/a n/a 2269 A.sub.esT.sub.es.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.e 660261 T.sub.es.sup.mC.sub.esT.sub.esT.sub.esG.sub.esG.sub.dsT.sub.dsT.sub- .dsA.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.ds PS GalNAc.sub.3-1a A.sub.d 2270 A.sub.esT.sub.es.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.eoA.sub.do'-GalNA- c.sub.3-1.sub.a 682883 GalNAc.sub.3-3.sub.a-o'T.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eoG.sub.- eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds.sup.mC.sub.dsA.sub.ds PS/PO GalNAc.sub.3-3a PO 2269 T.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.sup.mC.sub.es.sup.mC.sub- .es.sup.mC.sub.e 682884 GalNAc.sub.3-7.sub.a-o'T.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eoG.sub.- eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds.sup.mC.sub.dsA.sub.ds PS/PO GalNAc.sub.3-7a PO 2269 T.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.sup.mC.sub.es.sup.mC.sub- .es.sup.mC.sub.e 682885 GalNAc.sub.3-10.sub.a-o'T.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eoG.sub- .eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds.sup.mC.sub.ds PS/PO GalNAc.sub.3-10a PO 2269 A.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.sup.mC.sub.es.su- p.mC.sub.es.sup.mC.sub.e 682886 GalNAc.sub.3-13.sub.a-o'T.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eoGe.su- b.eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds.sup.mC.sub.ds PS/PO GalNAc.sub.3-13a PO 2269 A.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.sup.mC.sub.es.su- p.mC.sub.es.sup.mC.sub.e 684057 T.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eoG.sub.eoG.sub.dsT.sub.dsT.sub- .dsA.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.ds PS/PO GalNAc.sub.3-19a A.sub.d 2270 A.sub.eoT.sub.eo.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.eoA.sub.do'-GalNA- c.sub.3-19.sub.a
The legend for Table 85 can be found in Example 74. The structure of GalNAc.sub.3-1 was shown in Example 9. The structure of GalNAc.sub.3-3.sub.a was shown in Example 39. The structure of GalNAc.sub.3-7.sub.a was shown in Example 48. The structure of GalNAc.sub.3-10.sub.a was shown in Example 46. The structure of GalNAc.sub.3-13.sub.a was shown in Example 62. The structure of GalNAc.sub.3-19.sub.a was shown in Example 70.
TABLE-US-00072 TABLE 84 Antisense inhibition of human TTR in vivo TTR Plasma TTR SEQ Isis Dosage mRNA protein GalNAc ID No. (mg/kg) (% PBS) (% PBS) cluster CM No. PBS n/a 100 100 n/a n/a 420915 6 99 95 n/a n/a 2269 20 48 65 60 18 28 660261 0.6 113 87 GalNAc.sub.3-1a A.sub.d 2270 2 40 56 6 20 27 20 9 11
TABLE-US-00073 TABLE 85 Antisense inhibition of human TTR in vivo TTR Plasma TTR protein (% PBS at BL) SEQ Dosage mRNA Day 17 GalNAc ID Isis No. (mg/kg) (% PBS) BL Day 3 Day 10 (After sac) cluster CM No. PBS n/a 100 100 96 90 114 n/a n/a 420915 6 74 106 86 76 83 n/a n/a 2269 20 43 102 66 61 58 60 24 92 43 29 32 682883 0.6 60 88 73 63 68 GalNAc.sub.3-3a PO 2269 2 18 75 38 23 23 6 10 80 35 11 9 682884 0.6 56 88 78 63 67 GalNAc.sub.3-7a PO 2269 2 19 76 44 25 23 6 15 82 35 21 24 682885 0.6 60 92 77 68 76 GalNAc.sub.3-10a PO 2269 2 22 93 58 32 32 6 17 85 37 25 20 682886 0.6 57 91 70 64 69 GalNAc.sub.3-13a PO 2269 2 21 89 50 31 30 6 18 102 41 24 27 684057 0.6 53 80 69 56 62 GalNAc.sub.3-19a A.sub.d 2270 2 21 92 55 34 30 6 11 82 50 18 13
TABLE-US-00074 TABLE 86 Transaminase levels, body weight changes, and relative organ weights Dosage ALT (U/L) AST (U/L) Body Liver Spleen Kidney SEQ (mg / Day Day Day Day Day Day (% (% (% (% ID Isis No. kg) BL 3 10 17 BL 3 10 17 BL) PBS) PBS) PBS) No. PBS n/a 33 34 33 24 58 62 67 52 105 100 100 100 n/a 420915 6 34 33 27 21 64 59 73 47 115 99 89 91 2269 20 34 30 28 19 64 54 56 42 111 97 83 89 60 34 35 31 24 61 58 71 58 113 102 98 95 660261 0.6 33 38 28 26 70 71 63 59 111 96 99 92 2270 2 29 32 31 34 61 60 68 61 118 100 92 90 6 29 29 28 34 58 59 70 90 114 99 97 95 20 33 32 28 33 64 54 68 95 114 101 106 92
TABLE-US-00075 TABLE 87 Transaminase levels, body weight changes, and relative organ weights Dosage ALT (U/L) AST (U/L) Body Liver Spleen Kidney SEQ (mg/ BL Day Day Day Day Day Day (% (% (% (% ID Isis No. kg) 3 10 17 BL 3 10 17 BL) PBS) PBS) PBS) No. PBS n/a 32 34 37 41 62 78 76 77 104 100 100 100 n/a 420915 6 32 30 34 34 61 71 72 66 102 103 102 105 2269 20 41 34 37 33 80 76 63 54 106 107 135 101 60 36 30 32 34 58 81 57 60 106 105 104 99 682883 0.6 32 35 38 40 53 81 74 76 104 101 112 95 2269 2 38 39 42 43 71 84 70 77 107 98 116 99 6 35 35 41 38 62 79 103 65 105 103 143 97 682884 0.6 33 32 35 34 70 74 75 67 101 100 130 99 2269 2 31 32 38 38 63 77 66 55 104 103 122 100 6 38 32 36 34 65 85 80 62 99 105 129 95 682885 0.6 39 26 37 35 63 63 77 59 100 109 109 112 2269 2 30 26 38 40 54 56 71 72 102 98 111 102 6 27 27 34 35 46 52 56 64 102 98 113 96 682886 0.6 30 40 34 36 58 87 54 61 104 99 120 101 2269 2 27 26 34 36 51 55 55 69 103 91 105 92 6 40 28 34 37 107 54 61 69 109 100 102 99 684057 0.6 35 26 33 39 56 51 51 69 104 99 110 102 2270 2 33 32 31 40 54 57 56 87 103 100 112 97 6 39 33 35 40 67 52 55 92 98 104 121 108
Example 87: Duration of Action In Vivo by Single Doses of Oligonucleotides Targeting TTR Comprising a GalNAc.sub.3 Cluster
[1004] ISIS numbers 420915 and 660261 (see Table 83) were tested in a single dose study for duration of action in mice. ISIS numbers 420915, 682883, and 682885 (see Table 83) were also tested in a single dose study for duration of action in mice.
Treatment
[1005] Eight week old, male transgenic mice that express human TTR were each injected subcutaneously once with 100 mg/kg ISIS No. 420915 or 13.5 mg/kg ISIS No. 660261. Each treatment group consisted of 4 animals. Tail bleeds were performed before dosing to determine baseline and at days 3, 7, 10, 17, 24, and 39 following the dose. Plasma TTR protein levels were measured as described in Example 86. The results below are presented as the average percent of plasma TTR levels for each treatment group, normalized to baseline levels.
TABLE-US-00076 TABLE 88 Plasma TTR protein levels Time point SEQ ISIS Dosage (days TTR GalNAc.sub.3 ID No. (mg/kg) post-dose) (% baseline) Cluster CM No. 420915 100 3 30 n/a n/a 2269 7 23 10 35 17 53 24 75 39 100 660261 13.5 3 27 GalNAc.sub.3-1a A.sub.d 2270 7 21 10 22 17 36 24 48 39 69
Treatment
[1006] Female transgenic mice that express human TTR were each injected subcutaneously once with 100 mg/kg ISIS No. 420915, 10.0 mg/kg ISIS No. 682883, or 10.0 mg/kg 682885. Each treatment group consisted of 4 animals. Tail bleeds were performed before dosing to determine baseline and at days 3, 7, 10, 17, 24, and 39 following the dose. Plasma TTR protein levels were measured as described in Example 86. The results below are presented as the average percent of plasma TTR levels for each treatment group, normalized to baseline levels.
TABLE-US-00077 TABLE 89 Plasma TTR protein levels Time point SEQ ISIS Dosage (days TTR GalNAc.sub.3 ID No. (mg/kg) post-dose) (% baseline) Cluster CM No. 420915 100 3 48 n/a n/a 2269 7 48 10 48 17 66 31 80 682883 10.0 3 45 GalNAc.sub.3-3a PO 2269 7 37 10 38 17 42 31 65 682885 10.0 3 40 GalNAc.sub.3-10a PO 2269 7 33 10 34 17 40 31 64
The results in Tables 88 and 89 show that the oligonucleotides comprising a GalNAc conjugate are more potent with a longer duration of action than the parent oligonucleotide lacking a conjugate (ISIS 420915).
Example 88: Splicing Modulation In Vivo by Oligonucleotides Targeting SMN Comprising a GalNAc.sub.3 Conjugate
[1007] The oligonucleotides listed in Table 90 were tested for splicing modulation of human survival of motor neuron (SMN) in mice.
TABLE-US-00078 TABLE 90 Modified ASOs targeting SMN ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 387954 A.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.es.sup.mC.sub.esT.sub.e- sT.sub.esT.sub.es.sup.mC.sub.esA.sub.esT.sub.esA.sub.esA.sub.esT.sub.esG.s- ub.es.sup.mC.sub.esT.sub.esG.sub.es n/a n/a 2271 G.sub.e 699819 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.esT.sub.esT.sub.es.sup.mC.sub.esA- .sub.es.sup.mC.sub.esT.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.esT.sub.e- sA.sub.esA.sub.es GalNAc.sub.3-7a PO 2271 T.sub.esG.sub.es.sup.mC.sub.esT.sub.esG.sub.esG.sub.e 699821 GalNAc.sub.3-7.sub.a-.sub.o'A.sub.esT.sub.eoT.sub.eo.sup.mC.sub.eoA- .sub.eo.sup.mC.sub.eoT.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.eoA.sub.eoT.sub.e- oA.sub.eo GalNAc.sub.3-7a PO 2271 A.sub.eoT.sub.eoG.sub.eo.sup.mC.sub.eoT.sub.esG.sub.esG.sub.e 700000 A.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.es.sup.mC.sub.esT.sub.e- sT.sub.esT.sub.es.sup.mC.sub.esA.sub.esT.sub.esA.sub.esA.sub.esT.sub.esG.s- ub.es.sup.mC.sub.esT.sub.esG.sub.es GalNAc.sub.3-1a A.sub.d 2272 G.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a 703421 X-ATT.sup.mCA.sup.mCTTT.sup.mCATAATG.sup.mCTGG n/a n/a 2271 703422 GalNAc.sub.3-7.sub.b-X-ATT.sup.mCA.sup.mCTTT.sup.mCATAATG.sup.mCTGG GalNAc.sub.3-7b n/a 2271
The structure of GalNAc.sub.3-7.sub.a was shown previously in Example 48. "X" indicates a 5' primary amine generated by Gene Tools (Philomath, Oreg.), and GalNAc.sub.3-7.sub.b indicates the structure of GalNAc.sub.3-7.sub.a lacking the --NH--C.sub.6--O portion of the linker as shown below:
##STR00259##
ISIS numbers 703421 and 703422 are morphlino oligonucleotides, wherein each nucleotide of the two oligonucleotides is a morpholino nucleotide.
Treatment
[1008] Six week old transgenic mice that express human SMN were injected subcutaneously once with an oligonucleotide listed in Table 91 or with saline. Each treatment group consisted of 2 males and 2 females. The mice were sacrificed 3 days following the dose to determine the liver human SMN mRNA levels both with and without exon 7 using real-time PCR according to standard protocols. Total RNA was measured using Ribogreen reagent. The SMN mRNA levels were normalized to total mRNA, and further normalized to the averages for the saline treatment group. The resulting average ratios of SMN mRNA including exon 7 to SMN mRNA missing exon 7 are shown in Table 91. The results show that fully modified oligonucleotides that modulate splicing and comprise a GalNAc conjugate are significantly more potent in altering splicing in the liver than the parent oligonucleotides lacking a GlaNAc conjugate. Furthermore, this trend is maintained for multiple modification chemistries, including 2'-MOE and morpholino modified oligonucleotides.
TABLE-US-00079 TABLE 91 Effect of oligonucleotides targeting human SMN in vivo ISIS Dose +Exon GalNAc.sub.3 SEQ No. (mg/kg) 7/-Exon 7 Cluster CM ID No. Saline n/a 1.00 n/a n/a n/a 387954 32 1.65 n/a n/a 2271 387954 288 5.00 n/a n/a 2271 699819 32 7.84 GalNAc.sub.3-7a PO 2271 699821 32 7.22 GalNAc.sub.3-7a PO 2271 700000 32 6.91 GalNAc.sub.3-1a A.sub.d 2272 703421 32 1.27 n/a n/a 2271 703422 32 4.12 GalNAc.sub.3-7b n/a 2271
Example 89: Antisense Inhibition In Vivo by Oligonucleotides Targeting Apolipoprotein A (Apo(a)) Comprising a GalNAc.sub.3 Conjugate
[1009] The oligonucleotides listed in Table 92 below were tested in a study for dose-dependent inhibition of Apo(a) in transgenic mice.
TABLE-US-00080 TABLE 92 Modified ASOs targeting Apo(a) ISIS GalNAc.sub.3 SEQ ID No. Sequences (5' to 3') Cluster CM No. 494372 T.sub.esG.sub.es.sup.mC.sub.esT.sub.es.sup.mC.sub.es.sup.mC.sub.dsG- .sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.ds n/a n/a 2281 T.sub.dsT.sub.esG.sub.esT.sub.esT.sub.es.sup.mC.sub.e 681257 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.- sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-7a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eoG.sub.eoT.sub.esT.sub.es.su- p.mC.sub.e
[1010] The structure of GalNAc.sub.3-7.sub.a was shown in Example 48.
Treatment
[1011] Eight week old, female C57BL/6 mice (Jackson Laboratory, Bar Harbor, Me.) were each injected subcutaneously once per week at a dosage shown below, for a total of six doses, with an oligonucleotide listed in Table 92 or with PBS. Each treatment group consisted of 3-4 animals. Tail bleeds were performed the day before the first dose and weekly following each dose to determine plasma Apo(a) protein levels. The mice were sacrificed two days following the final administration. Apo(a) liver mRNA levels were determined using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) according to standard protocols. Apo(a) plasma protein levels were determined using ELISA, and liver transaminase levels were determined. The mRNA and plasma protein results in Table 93 are presented as the treatment group average percent relative to the PBS treated group. Plasma protein levels were further normalized to the baseline (BL) value for the PBS group. Average absolute transaminase levels and body weights (% relative to baseline averages) are reported in Table 94.
[1012] As illustrated in Table 93, treatment with the oligonucleotides lowered Apo(a) liver mRNA and plasma protein levels in a dose-dependent manner. Furthermore, the oligonucleotide comprising the GalNAc conjugate was significantly more potent with a longer duration of action than the parent oligonucleotide lacking a GalNAc conjugate. As illustrated in Table 94, transaminase levels and body weights were unaffected by the oligonucleotides, indicating that the oligonucleotides were well tolerated.
TABLE-US-00081 TABLE 93 Apo(a) liver mRNA and plasma protein levels ISIS Dosage Apo(a) mRNA Apo(a) plasma protein (% PBS) No. (mg/kg) (% PBS) BL Week 1 Week 2 Week 3 Week 4 Week 5 Week 6 PBS n/a 100 100 120 119 113 88 121 97 494372 3 80 84 89 91 98 87 87 79 10 30 87 72 76 71 57 59 46 30 5 92 54 28 10 7 9 7 681257 0.3 75 79 76 89 98 71 94 78 1 19 79 88 66 60 54 32 24 3 2 82 52 17 7 4 6 5 10 2 79 17 6 3 2 4 5
TABLE-US-00082 TABLE 94 Dosage Body weight ISIS No. (mg/kg) ALT (U/L) AST (U/L) (% baseline) PBS n/a 37 54 103 494372 3 28 68 106 10 22 55 102 30 19 48 103 681257 0.3 30 80 104 1 26 47 105 3 29 62 102 10 21 52 107
Example 90: Antisense Inhibition In Vivo by Oligonucleotides Targeting TTR Comprising a GalNAc.sub.3 Cluster
[1013] Oligonucleotides listed in Table 95 below were tested in a dose-dependent study for antisense inhibition of human transthyretin (TTR) in transgenic mice that express the human TTR gene.
Treatment
[1014] TTR transgenic mice were each injected subcutaneously once per week for three weeks, for a total of three doses, with an oligonucleotide and dosage listed in Table 96 or with PBS. Each treatment group consisted of 4 animals. Prior to the first dose, a tail bleed was performed to determine plasma TTR protein levels at baseline (BL). The mice were sacrificed 72 hours following the final administration. TTR protein levels were measured using a clinical analyzer (AU480, Beckman Coulter, CA). Real-time PCR and RIBOGREEN RNA quantification reagent (Molecular Probes, Inc. Eugene, Oreg.) were used according to standard protocols to determine liver human TTR mRNA levels. The results presented in Table 96 are the average values for each treatment group. The mRNA levels are the average values relative to the average for the PBS group. Plasma protein levels are the average values relative to the average value for the PBS group at baseline. "BL" indicates baseline, measurements that were taken just prior to the first dose. As illustrated in Table 96, treatment with antisense oligonucleotides lowered TTR expression levels in a dose-dependent manner. The oligonucleotides comprising a GalNAc conjugate were more potent than the parent lacking a GalNAc conjugate (ISIS 420915), and oligonucleotides comprising a phosphodiester or deoxyadenosine cleavable moiety showed significant improvements in potency compared to the parent lacking a conjugate (see ISIS numbers 682883 and 666943 vs 420915 and see Examples 86 and 87).
TABLE-US-00083 TABLE 95 Oligonucleotides targeting human TTR GalNAc SEQ Isis No. Sequence 5' to 3' Linkages cluster CM ID No. 420915 T.sub.es.sup.mC.sub.esT.sub.esT.sub.esG.sub.esG.sub.dsT.sub.dsT.sub- .dsA.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.ds PS n/a n/a 2269 A.sub.esT.sub.es.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.e 682883 GalNAc.sub.3-3.sub.a-o'T.sub.es.sup.mC.sub.eoT.sub.eoT.sub.eoG.sub.- eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds.sup.mC.sub.dsA.sub.ds PS/PO GalNAc.sub.3-3a PO 2269 T.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.sup.mC.sub.es.sup.mC.sub- .es.sup.mC.sub.e 666943 GalNAc.sub.3-3.sub.a-o'A.sub.doT.sub.es.sup.mC.sub.eoT.sub.eoT.sub.- eoG.sub.eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds PS/PO GalNAc.sub.3-3a A.sub.d 2273 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.ds A.sub.eoT.sub.eo.sup.mC.sub.es.sup.mC.sub.es.sup.mC.sub.e 682887 GalNAc.sub.3-7.sub.a-o'A.sub.doT.sub.es.sup.mC.sub.eoT.sub.eoT.sub.- eoG.sub.eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds PS/PO GalNAc.sub.3-7a A.sub.d 2273 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.su- p.mC.sub.es.sup.mC.sub.es.sup.mC.sub.e 682888 GalNAc.sub.3-10.sub.a-o'A.sub.doT.sub.es.sup.mC.sub.eoT.sub.eoT.sub- .eoG.sub.eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds PS/PO GalNAc.sub.3-10a A.sub.d 2273 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.su- p.mC.sub.es.sup.mC.sub.es.sup.mC.sub.e 682889 GalNAc.sub.3-13.sub.a-o'A.sub.doT.sub.es.sup.mC.sub.eoT.sub.eoT.sub- .eoG.sub.eoG.sub.dsT.sub.dsT.sub.dsA.sub.ds PS/PO GalNAc.sub.3-13a A.sub.d 2273 .sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.dsA.sub.dsA.sub.eoT.sub.eo.su- p.mC.sub.es.sup.mC.sub.es.sup.mC.sub.e
The legend for Table 95 can be found in Example 74. The structure of GalNAc.sub.3-3.sub.a was shown in Example 39. The structure of GalNAc.sub.3-7.sub.a was shown in Example 48. The structure of GalNAc.sub.3-10.sub.a was shown in Example 46. The structure of GalNAc.sub.3-13.sub.a was shown in Example 62.
TABLE-US-00084 TABLE 96 Antisense inhibition of human TTR in vivo Dosage TTR mRNA TTR protein GalNAc Isis No. (mg/kg) (% PBS) (% BL) cluster CM PBS n/a 100 124 n/a n/a 420915 6 69 114 n/a n/a 20 71 86 60 21 36 682883 0.6 61 73 GalNAc.sub.3-3a PO 2 23 36 6 18 23 666943 0.6 74 93 GalNAc.sub.3-3a A.sub.d 2 33 57 6 17 22 682887 0.6 60 97 GalNAc.sub.3-7a A.sub.d 2 36 49 6 12 19 682888 0.6 65 92 GalNAc.sub.3-10a A.sub.d 2 32 46 6 17 22 682889 0.6 72 74 GalNAc.sub.3-13a A.sub.d 2 38 45 6 16 18
Example 91: Antisense Inhibition In Vivo by Oligonucleotides Targeting Factor VII Comprising a GalNAc.sub.3 Conjugate in Non-Human Primates
[1015] Oligonucleotides listed in Table 97 below were tested in a non-terminal, dose escalation study for antisense inhibition of Factor VII in monkeys.
Treatment
[1016] Non-naive monkeys were each injected subcutaneously on days 0, 15, and 29 with escalating doses of an oligonucleotide listed in Table 97 or with PBS. Each treatment group consisted of 4 males and 1 female. Prior to the first dose and at various time points thereafter, blood draws were performed to determine plasma Factor VII protein levels. Factor VII protein levels were measured by ELISA. The results presented in Table 98 are the average values for each treatment group relative to the average value for the PBS group at baseline (BL), the measurements taken just prior to the first dose. As illustrated in Table 98, treatment with antisense oligonucleotides lowered Factor VII expression levels in a dose-dependent manner, and the oligonucleotide comprising the GalNAc conjugate was significantly more potent in monkeys compared to the oligonucleotide lacking a GalNAc conjugate.
TABLE-US-00085 TABLE 97 Oligonucleotides targeting Factor VII GalNAc SEQ Isis No. Sequence 5' to 3' Linkages cluster CM ID No. 407935 A.sub.esT.sub.esG.sub.es.sup.mC.sub.esA.sub.esT.sub.dsG.sub.dsG.sub- .dsT.sub.dsG.sub.dsA.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.ds PS n/a n/a 2274 T.sub.es.sup.mC.sub.esT.sub.esG.sub.esA.sub.e 686892 GalNAc.sub.3-10.sub.a-o'A.sub.esT.sub.esG.sub.es.sup.mC.sub.esA.sub- .esT.sub.dsG.sub.dsG.sub.dsT.sub.dsG.sub.ds PS GalNAc.sub.3-10a PO 2274 A.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.esT.sub- .esG.sub.esA.sub.e
The legend for Table 97 can be found in Example 74. The structure of GalNAc.sub.3-10.sub.a was shown in Example 46.
TABLE-US-00086 TABLE 98 Factor VII plasma protein levels ISIS No. Day Dose (mg/kg) Factor VII (% BL) 407935 0 n/a 100 15 10 87 22 n/a 92 29 30 77 36 n/a 46 43 n/a 43 686892 0 3 100 15 10 56 22 n/a 29 29 30 19 36 n/a 15 43 n/a 11
Example 92: Antisense Inhibition in Primary Hepatocytes by Antisense Oligonucleotides Targeting ApoC-III Comprising a GalNAc.sub.3 Conjugate
[1017] Primary mouse hepatocytes were seeded in 96-well plates at 15,000 cells per well, and the oligonucleotides listed in Table 99, targeting mouse ApoC-III, were added at 0.46, 1.37, 4.12, or 12.35, 37.04, 111.11, or 333.33 nM or 1.00 M. After incubation with the oligonucleotides for 24 hours, the cells were lysed and total RNA was purified using RNeasy (Qiagen). ApoC-III mRNA levels were determined using real-time PCR and RIBOGREEN.RTM. RNA quantification reagent (Molecular Probes, Inc.) according to standard protocols. IC.sub.50 values were determined using Prism 4 software (GraphPad). The results show that regardless of whether the cleavable moiety was a phosphodiester or a phosphodiester-linked deoxyadensoine, the oligonucleotides comprising a GalNAc conjugate were significantly more potent than the parent oligonucleotide lacking a conjugate.
TABLE-US-00087 TABLE 99 Inhibition of mouse APOC-III expression in mouse primary hepatocytes ISIS IC.sub.50 SEQ No. Sequence (5' to 3') CM (nM) ID No. 440670 .sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.dsT.sub.d- sA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.sub.dsA.sub.ds.sup.mC.s- ub.esA.sub.esG.sub.es.sup.mC.sub.esA.sub.e n/a 13.20 2275 661180 .sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.dsT.sub.d- sA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.sub.dsA.sub.ds.sup.mC.s- ub.es A.sub.d 1.40 2276 A.sub.esG.sub.es.sup.mC.sub.esA.sub.eoA.sub.do'-GalNAc.sub.3-1.sub.a 680771 GalNAc.sub.3-3.sub.a-o'.sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.es- T.sub.esT.sub.dsT.sub.dsA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.- sub.dsA.sub.ds.sup.mC.sub.es PO 0.70 2275 A.sub.esG.sub.es.sup.mC.sub.esA.sub.e 680772 GalNAc.sub.3-7.sub.a-o'.sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.es- T.sub.esT.sub.dsT.sub.dsA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.- sub.dsA.sub.ds.sup.mC.sub.es PO 1.70 2275 A.sub.esG.sub.es.sup.mC.sub.esA.sub.e 680773 GalNAc.sub.3-10.sub.a-o'.sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.e- sT.sub.esT.sub.dsT.sub.dsA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG- .sub.dsA.sub.ds.sup.mC.sub.es PO 2.00 2275 A.sub.esG.sub.es.sup.mC.sub.esA.sub.e 680774 GalNAc.sub.3-13.sub.a-o'.sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.e- sT.sub.esT.sub.dsT.sub.dsA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG- .sub.dsA.sub.ds.sup.mC.sub.es PO 1.50 2275 A.sub.esG.sub.es.sup.mC.sub.esA.sub.e 681272 GalNAc.sub.3-3.sub.a-o'.sup.mC.sub.esA.sub.eoG.sub.eo.sup.mC.sub.eo- T.sub.eoT.sub.dsT.sub.dsA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.- sub.dsA.sub.ds.sup.mC.sub.eo PO <0.46 2275 A.sub.eoG.sub.es.sup.mC.sub.esA.sub.e 681273 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.do.sup.mC.sub.esA.sub.esG.sub.es.- sup.mC.sub.esT.sub.esT.sub.dsT.sub.dsA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub- .dsG.sub.dsG.sub.dsA.sub.ds A.sub.d 1.10 2277 .sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.esA.sub.e 683733 .sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.dsT.sub.d- sA.sub.dsT.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.sub.dsA.sub.ds.sup.mC.s- ub.es A.sub.d 2.50 2276 A.sub.esG.sub.es.sup.mC.sub.esA.sub.eoA.sub.do'-GalNAc.sub.3-19.sub.a
The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9, GalNAc.sub.3-3.sub.a was shown in Example 39, GalNAc.sub.3-7.sub.a was shown in Example 48, GalNAc.sub.3-10.sub.a was shown in Example 46, GalNAc.sub.3-13.sub.a was shown in Example 62, and GalNAc.sub.3-19.sub.a was shown in Example 70.
Example 93: Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising Mixed Wings and a 5'-GalNAc.sub.3 Conjugate
[1018] The oligonucleotides listed in Table 100 were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice.
TABLE-US-00088 TABLE 100 Modified ASOs targeting SRB-1 ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 449093 T.sub.ksT.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.d- sA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub- .ks.sup.mC.sub.k n/a n/a 2278 699806 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.ksT.sub.ks.sup.mC.sub.ksA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.ds A.sub.dsT.sub.ds G.sub.dsA.sub.ds.sup.mC.sub.ds GalNAc.sub.3-3a PO 2278 T.sub.dsT.sub.ks.sup.mC.sub.ks.sup.mC.sub.k 699807 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.ksT.sub.ks.sup.mC.sub.ksA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.ds A.sub.dsT.sub.ds G.sub.dsA.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7a PO 2278 T.sub.dsT.sub.ks.sup.mC.sub.ks.sup.mC.sub.k 699809 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.ksT.sub.ks.sup.mC.sub.ksA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.ds A.sub.dsT.sub.ds G.sub.ds A.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7a PO 2278 T.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.e 699811 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.ds A.sub.dsT.sub.ds G.sub.dsA.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7a PO 2278 T.sub.dsT.sub.ks.sup.mC.sub.ks.sup.mC.sub.k 699813 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.ksT.sub.ds.sup.mC.sub.ksA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.ds A.sub.dsT.sub.ds G.sub.dsA.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7a PO 2278 T.sub.dsT.sub.ks.sup.mC.sub.ds.sup.mC.sub.k 699815 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esT.sub.ks.sup.mC.sub.ksA.sub.dsG- .sub.dsT.sub.ds.sup.mC.sub.ds A.sub.dsT.sub.ds G.sub.dsA.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7a PO 2278 T.sub.dsT.sub.ks.sup.mC.sub.ks.sup.mC.sub.e
The structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39, and the structure of GalNAc.sub.3-7a was shown previously in Example 48. Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "k" indicates 6'-(S)--CH.sub.3 bicyclic nucleoside (cEt); "s" indicates phosphorothioate internucleoside linkages (PS); "o" indicates phosphodiester internucleoside linkages (PO). Superscript "m" indicates 5-methylcytosines.
Treatment
[1019] Six to eight week old C57BL/6 mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once at the dosage shown below with an oligonucleotide listed in Table 100 or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration. Liver SRB-1 mRNA levels were measured using real-time PCR. SRB-1 mRNA levels were normalized to cyclophilin mRNA levels according to standard protocols. The results are presented as the average percent of SRB-1 mRNA levels for each treatment group relative to the saline control group. As illustrated in Table 101, treatment with antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner, and the gapmer oligonucleotides comprising a GalNAc conjugate and having wings that were either full cEt or mixed sugar modifications were significantly more potent than the parent oligonucleotide lacking a conjugate and comprising full cEt modified wings.
[1020] Body weights, liver transaminases, total bilirubin, and BUN were also measured, and the average values for each treatment group are shown in Table 101. Body weight is shown as the average percent body weight relative to the baseline body weight (% BL) measured just prior to the oligonucleotide dose.
TABLE-US-00089 TABLE 101 SRB-1 mRNA, ALT, AST, BUN, and total bilirubin levels and body weights SRB-1 Body ISIS Dosage mRNA ALT AST weight No. (mg/kg) (% PBS) (U/L) (U/L) Bil BUN (% BL) PBS n/a 100 31 84 0.15 28 102 449093 1 111 18 48 0.17 31 104 3 94 20 43 0.15 26 103 10 36 19 50 0.12 29 104 699806 0.1 114 23 58 0.13 26 107 0.3 59 21 45 0.12 27 108 1 25 30 61 0.12 30 104 699807 0.1 121 19 41 0.14 25 100 0.3 73 23 56 0.13 26 105 1 24 22 69 0.14 25 102 699809 0.1 125 23 57 0.14 26 104 0.3 70 20 49 0.10 25 105 1 33 34 62 0.17 25 107 699811 0.1 123 48 77 0.14 24 106 0.3 94 20 45 0.13 25 101 1 66 57 104 0.14 24 107 699813 0.1 95 20 58 0.13 28 104 0.3 98 22 61 0.17 28 105 1 49 19 47 0.11 27 106 699815 0.1 93 30 79 0.17 25 105 0.3 64 30 61 0.12 26 105 1 24 18 41 0.14 25 106
Example 94: Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising 2'-Sugar Modifications and a 5'-GalNAc.sub.3 Conjugate
[1021] The oligonucleotides listed in Table 102 were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice.
TABLE-US-00090 TABLE 102 Modified ASOs targeting SRB-1 ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 353382 G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.esA.sub.dsG.sub.d- sT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.es.sup.mC.sub.es.sup.mC.sub.es n/a n/a 2256 T.sub.esT.sub.e 700989 G.sub.msC.sub.msU.sub.msU.sub.msC.sub.msA.sub.dsG.sub.dsT.sub.ds.su- p.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsU.sub.msC- .sub.msC.sub.ms n/a n/a 2279 U.sub.msU.sub.m 666904 GalNAc.sub.3-3.sub.a-.sub.o'G.sub.es.sup.mC.sub.esT.sub.esT.sub.es.- sup.mC.sub.esA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.d- sA.sub.ds GalNAc.sub.3-3a PO 2256 .sup.mC.sub.dsT.sub.dsT.sub.es.sup.mC.sub.es.sup.mC.sub.esT.sub.esT.sub.e 700991 GalNAc.sub.3-7.sub.a-.sub.o'G.sub.msC.sub.msU.sub.msU.sub.msC.sub.m- sA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.ds GalNAc.sub.3-7a PO 2279 A.sub.ds.sup.mC.sub.dsT.sub.dsU.sub.msC.sub.msC.sub.msU.sub.msU.sub.m
Subscript "m" indicates a 2'-O-methyl modified nucleoside. See Example 74 for complete table legend. The structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39, and the structure of GalNAc.sub.3-7a was shown previously in Example 48.
Treatment
[1022] The study was completed using the protocol described in Example 93. Results are shown in Table 103 below and show that both the 2'-MOE and 2'-OMe modified oligonucleotides comprising a GalNAc conjugate were significantly more potent than the respective parent oligonucleotides lacking a conjugate. The results of the body weights, liver transaminases, total bilirubin, and BUN measurements indicated that the compounds were all well tolerated.
TABLE-US-00091 TABLE 103 SRB-1 mRNA ISIS No. Dosage (mg/kg) SRB-1 mRNA (% PBS) PBS n/a 100 353382 5 116 15 58 45 27 700989 5 120 15 92 45 46 666904 1 98 3 45 10 17 700991 1 118 3 63 10 14
Example 95: Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising Bicyclic Nucleosides and a 5'-GalNAc.sub.3 Conjugate
[1023] The oligonucleotides listed in Table 104 were tested in a dose-dependent study for antisense inhibition of SRB-1 in mice.
TABLE-US-00092 TABLE 104 Modified ASOs targeting SRB-1 ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No 440762 T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.d- sT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k n/a n/a 2250 666905 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT- .sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.d- sT.sub.ks.sup.mC.sub.k GalNAc.sub.3-3.sub.a PO 2250 699782 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT- .sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.d- sT.sub.ks.sup.mC.sub.k GalNAc.sub.3-7.sub.a PO 2250 699783 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.ls.sup.mC.sub.lsA.sub.dsG.sub.dsT- .sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.d- sT.sub.ls.sup.mC.sub.l GalNAc.sub.3-3.sub.a PO 2250 653621 T.sub.ls.sup.mC.sub.lsA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.d- sT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ls.sup.mC.sub.loA.sub- .do'-GalNAc.sub.3-1.sub.a GalNAc.sub.3-1.sub.a A.sub.d 2251 439879 T.sub.gs.sup.mC.sub.gsA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.d- sT.sub.d G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.gs.sup.mC.sub.g n/a n/a 2250 699789 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.gs.sup.mC.sub.gsA.sub.dsG.sub.dsT- .sub.ds.sup.mC.sub.dsA.sub.dsT.sub.d G.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.gs.sup.mC.sub.g GalNAc.sub.3-3.sub.a PO 2250
Subscript "g" indicates a fluoro-HNA nucleoside, subscript "1" indicates a locked nucleoside comprising a 2'-O--CH.sub.2-4' bridge. See the Example 74 table legend for other abbreviations. The structure of GalNAc.sub.3-1.sub.a was shown previously in Example 9, the structure of GalNAc.sub.3-3.sub.a was shown previously in Example 39, and the structure of GalNAc.sub.3-7a was shown previously in Example 48.
Treatment
[1024] The study was completed using the protocol described in Example 93. Results are shown in Table 105 below and show that oligonucleotides comprising a GalNAc conjugate and various bicyclic nucleoside modifications were significantly more potent than the parent oligonucleotide lacking a conjugate and comprising bicyclic nucleoside modifications. Furthermore, the oligonucleotide comprising a GalNAc conjugate and fluoro-HNA modifications was significantly more potent than the parent lacking a conjugate and comprising fluoro-HNA modifications. The results of the body weights, liver transaminases, total bilirubin, and BUN measurements indicated that the compounds were all well tolerated.
TABLE-US-00093 TABLE 105 SRB-1 mRNA, ALT, AST, BUN, and total bilirubin levels and body weights ISIS No. Dosage (mg/kg) SRB-1 mRNA (% PBS) PBS n/a 100 440762 1 104 3 65 10 35 666905 0.1 105 0.3 56 1 18 699782 0.1 93 0.3 63 1 15 699783 0.1 105 0.3 53 1 12 653621 0.1 109 0.3 82 1 27 439879 1 96 3 77 10 37 699789 0.1 82 0.3 69 1 26
Example 96: Plasma Protein Binding of Antisense Oligonucleotides Comprising a GalNAc.sub.3 Conjugate Group
[1025] Oligonucleotides listed in Table 70 targeting ApoC-III and oligonucleotides in Table 106 targeting Apo(a) were tested in an ultra-filtration assay in order to assess plasma protein binding.
TABLE-US-00094 TABLE 106 Modified oligonucleotides targeting Apo(a) ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No 494372 T.sub.esG.sub.es.sup.mC.sub.esT.sub.es.sup.mC.sub.es.sup.mC.sub.dsG- .sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.esG.sub.esT.sub.es n/a n/a 2281 T.sub.es.sup.mC.sub.e 693401 T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.sup.mC.sub.eo.sup.mC.sub.dsG- .sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.eoG.sub.eoT.sub.es n/a n/a 2281 T.sub.es.sup.mC.sub.e 681251 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.es.sup.mC.sub.esT.sub.es.- sup.mC.sub.es.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.d- sG.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7.sub.a PO 2281 T.sub.dsT.sub.esG.sub.esT.sub.esT.sub.es.sup.mC.sub.e 681257 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.- sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.d- sG.sub.ds.sup.mC.sub.ds GalNAc.sub.3-7.sub.a PO 2281 T.sub.dsT.sub.eoG.sub.eoT.sub.esT.sub.es.sup.mC.sub.e
See the Example 74 for table legend. The structure of GalNAc.sub.3-7a was shown previously in Example 48.
[1026] Ultrafree-MC ultrafiltration units (30,000 NMWL, low-binding regenerated cellulose membrane, Millipore, Bedford, Mass.) were pre-conditioned with 300 .mu.L of 0.5% Tween 80 and centrifuged at 2000 g for 10 minutes, then with 300 .mu.L of a 300 .mu.g/mL solution of a control oligonucleotide in H.sub.2O and centrifuged at 2000 g for 16 minutes. In order to assess non-specific binding to the filters of each test oligonucleotide from Tables 70 and 106 to be used in the studies, 300 .mu.L of a 250 ng/mL solution of oligonucleotide in H.sub.2O at pH 7.4 was placed in the pre-conditioned filters and centrifuged at 2000 g for 16 minutes. The unfiltered and filtered samples were analyzed by an ELISA assay to determine the oligonucleotide concentrations. Three replicates were used to obtain an average concentration for each sample. The average concentration of the filtered sample relative to the unfiltered sample is used to determine the percent of oligonucleotide that is recovered through the filter in the absence of plasma (% recovery).
[1027] Frozen whole plasma samples collected in K3-EDTA from normal, drug-free human volunteers, cynomolgus monkeys, and CD-1 mice, were purchased from Bioreclamation LLC (Westbury, N.Y.). The test oligonucleotides were added to 1.2 mL aliquots of plasma at two concentrations (5 and 150 .mu.g/mL). An aliquot (300 .mu.L) of each spiked plasma sample was placed in a pre-conditioned filter unit and incubated at 37.degree. C. for 30 minutes, immediately followed by centrifugation at 2000 g for 16 minutes. Aliquots of filtered and unfiltered spiked plasma samples were analyzed by an ELISA to determine the oligonucleotide concentration in each sample. Three replicates per concentration were used to determine the average percentage of bound and unbound oligonucleotide in each sample. The average concentration of the filtered sample relative to the concentration of the unfiltered sample is used to determine the percent of oligonucleotide in the plasma that is not bound to plasma proteins (% unbound). The final unbound oligonucleotide values are corrected for non-specific binding by dividing the % unbound by the % recovery for each oligonucleotide. The final % bound oligonucleotide values are determined by subtracting the final % unbound values from 100. The results are shown in Table 107 for the two concentrations of oligonucleotide tested (5 and 150 .mu.g/mL) in each species of plasma. The results show that GalNAc conjugate groups do not have a significant impact on plasma protein binding. Furthermore, oligonucleotides with full PS internucleoside linkages and mixed PO/PS linkages both bind plasma proteins, and those with full PS linkages bind plasma proteins to a somewhat greater extent than those with mixed PO/PS linkages.
TABLE-US-00095 TABLE 107 Percent of modified oligonucleotide bound to plasma proteins ISIS Human plasma Monkey plasma Mouse plasma No. 5 .mu.g/mL 150 .mu.g/mL 5 .mu.g/mL 150 .mu.g/mL 5 .mu.g/mL 150 .mu.g/mL 304801 99.2 98.0 99.8 99.5 98.1 97.2 663083 97.8 90.9 99.3 99.3 96.5 93.0 674450 96.2 97.0 98.6 94.4 94.6 89.3 494372 94.1 89.3 98.9 97.5 97.2 93.6 693401 93.6 89.9 96.7 92.0 94.6 90.2 681251 95.4 93.9 99.1 98.2 97.8 96.1 681257 93.4 90.5 97.6 93.7 95.6 92.7
Example 97: Modified Oligonucleotides Targeting TTR Comprising a GalNAc.sub.3 Conjugate Group
[1028] The oligonucleotides shown in Table 108 comprising a GalNAc conjugate were designed to target TTR.
TABLE-US-00096 TABLE 108 Modified oligonucleotides targeting TTR GalNAc.sub.3 SEQ ID ISIS No. Sequences (5' to 3') Cluster CM No 666941 GalNAc.sub.3-3.sub.a-o'A.sub.do T.sub.es .sup.m C.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds GalNAc.sub.3-3 A.sub.d 2273 A.sub.ds T.sub.ds G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 666942 T.sub.es .sup.m C.sub.eo T.sub.eo T.sub.eo G.sub.eo G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds A.sub.ds GalNAc.sub.3-1 A.sub.d 2270 A.sub.eo T.sub.eo .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.eo A.sub.do'-GalNAc.sub.3-3.sub.a 682876 GalNAc.sub.3-3.sub.a-o'T.sub.es .sup.mC.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.3-3 PO 2269 G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 682877 GalNAc.sub.3-7.sub.a-o'T.sub.es .sup.m C.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.3-7 PO 2269 G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 682878 GalNAc.sub.3-10.sub.a-o'T.sub.es .sup.mC.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds A.sub.ds GalNAc.sub.3-10 PO 2269 T.sub.ds G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 682879 GalNAc.sub.3-13.sub.a-o'T.sub.es .sup.mC.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds A.sub.ds GalNAc.sub.3-13 PO 2269 T.sub.ds G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 682880 GalNAc.sub.3-7.sub.a-o'A.sub.do T.sub.es .sup.mC.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds GalNAc.sub.3-7 A.sub.d 2273 A.sub.ds T.sub.ds G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 682881 GalNAc.sub.3-10.sub.a-o'A.sub.do T.sub.es .sup.mC.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds GalNAc.sub.3-10 A.sub.d 2273 A.sub.ds T.sub.ds G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 682882 GalNAc.sub.3-13.sub.a-o'A.sub.do T.sub.es .sup.mC.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds GalNAc.sub.3-13 A.sub.d 2273 A.sub.ds T.sub.ds G.sub.ds A.sub.ds A.sub.ds A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.e 684056 T.sub.es .sup.m C.sub.es T.sub.es T.sub.es G.sub.es G.sub.ds T.sub.ds T.sub.ds A.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds A.sub.ds GalNAc.sub.3-19 A.sub.d 2270 A.sub.es T.sub.es .sup.mC.sub.es .sup.mC.sub.es .sup.mC.sub.eo A.sub.do'-GalNAc.sub.3-19.sub.a
The legend for Table 108 can be found in Example 74. The structure of GalNAc.sub.3-1 was shown in Example 9. The structure of GalNAc.sub.3-3.sub.a was shown in Example 39. The structure of GalNAc.sub.3-7.sub.a was shown in Example 48. The structure of GalNAc.sub.3-10.sub.a was shown in Example 46. The structure of GalNAc.sub.3-13.sub.a was shown in Example 62. The structure of GalNAc.sub.3-19.sub.a was shown in Example 70.
Example 98: Evaluation of Pro-Inflammatory Effects of Oligonucleotides Comprising a GalNAc Conjugate in hPMBC Assay
[1029] The oligonucleotides listed in Table 109 and were tested for pro-inflammatory effects in an hPMBC assay as described in Examples 23 and 24. (See Tables 30, 83, 95, and 108 for descriptions of the oligonucleotides.) ISIS 353512 is a high responder used as a positive control, and the other oligonucleotides are described in Tables 83, 95, and 108. The results shown in Table 109 were obtained using blood from one volunteer donor. The results show that the oligonucleotides comprising mixed PO/PS internucleoside linkages produced significantly lower pro-inflammatory responses compared to the same oligonucleotides having full PS linkages. Furthermore, the GalNAc conjugate group did not have a significant effect in this assay.
TABLE-US-00097 TABLE 109 ISIS No. E.sub.max/EC.sub.50 GalNAc.sub.3 cluster Linkages CM 353512 3630 n/a PS n/a 420915 802 n/a PS n/a 682881 1311 GalNAc.sub.3-10 PS A.sub.d 682888 0.26 GalNAc.sub.3-10 PO/PS A.sub.d 684057 1.03 GalNAc.sub.3-19 PO/PS A.sub.d
Example 99: Binding Affinities of Oligonucleotides Comprising a GalNAc Conjugate for the Asialoglycoprotein Receptor
[1030] The binding affinities of the oligonucleotides listed in Table 110 (see Table 76 for descriptions of the oligonucleotides) for the asialoglycoprotein receptor were tested in a competitive receptor binding assay. The competitor ligand, .alpha.1-acid glycoprotein (AGP), was incubated in 50 mM sodium acetate buffer (pH 5) with 1 U neuraminidase-agarose for 16 hours at 37.degree. C., and >90% desialylation was confirmed by either sialic acid assay or size exclusion chromatography (SEC). Iodine monochloride was used to iodinate the AGP according to the procedure by Atsma et al. (see J Lipid Res. 1991 January; 32(1): 173-81.) In this method, desialylated .alpha.1-acid glycoprotein (de-AGP) was added to 10 mM iodine chloride, Na.sup.125I, and 1 M glycine in 0.25 M NaOH. After incubation for 10 minutes at room temperature, .sup.125I-labeled de-AGP was separated from free .sup.125I by concentrating the mixture twice utilizing a 3 KDMWCO spin column. The protein was tested for labeling efficiency and purity on a HPLC system equipped with an Agilent SEC-3 column (7.8.times.300 mm) and a -RAM counter. Competition experiments utilizing .sup.125I-labeled de-AGP and various GalNAc-cluster containing ASOs were performed as follows. Human HepG2 cells (10.sup.6 cells/ml) were plated on 6-well plates in 2 ml of appropriate growth media. MEM media supplemented with 10% fetal bovine serum (FBS), 2 mM L-Glutamine and 10 mM HEPES was used. Cells were incubated 16-20 hours @ 37.degree. C. with 5% and 10% CO.sub.2 respectively. Cells were washed with media without FBS prior to the experiment. Cells were incubated for 30 min @37.degree. C. with 1 ml competition mix containing appropriate growth media with 2% FBS, 10.sup.-8 M .sup.125I-labeled de-AGP and GalNAc-cluster containing ASOs at concentrations ranging from 10.sup.-11 to 10.sup.-5 M. Non-specific binding was determined in the presence of 10.sup.-2 M GalNAc sugar. Cells were washed twice with media without FBS to remove unbound .sup.125I-labeled de-AGP and competitor GalNAc ASO. Cells were lysed using Qiagen's RLT buffer containing 1% -mercaptoethanol. Lysates were transferred to round bottom assay tubes after a brief 10 min freeze/thaw cycle and assayed on a .gamma.-counter. Non-specific binding was subtracted before dividing .sup.125I protein counts by the value of the lowest GalNAc-ASO concentration counts. The inhibition curves were fitted according to a single site competition binding equation using a nonlinear regression algorithm to calculate the binding affinities (K.sub.D's).
[1031] The results in Table 110 were obtained from experiments performed on five different days. Results for oligonucleotides marked with superscript "a" are the average of experiments run on two different days. The results show that the oligonucleotides comprising a GalNAc conjugate group on the 5'-end bound the asialoglycoprotein receptor on human HepG2 cells with 1.5 to 16-fold greater affinity than the oligonucleotides comprising a GalNAc conjugate group on the 3'-end.
TABLE-US-00098 TABLE 110 Asialoglycoprotein receptor binding assay results Oligonucleotide end to GalNAc which GalNAc ISIS No. conjugate conjugate is attached K.sub.D (nM) 661161.sup.a GalNAc.sub.3-3 5' 3.7 666881.sup.a GalNAc.sub.3-10 5' 7.6 666981 GalNAc.sub.3-7 5' 6.0 670061 GalNAc.sub.3-13 5' 7.4 655861.sup.a GalNAc.sub.3-1 3' 11.6 677841.sup.a GalNAc.sub.3-19 3' 60.8
Example 100: Antisense Inhibition In Vivo by Oligonucleotides Comprising a GalNAc Conjugate Group Targeting Apo(a) In Vivo
[1032] The oligonucleotides listed in Table 111a below were tested in a single dose study for duration of action in mice.
TABLE-US-00099 TABLE 111a Modified ASOs targeting APO(a) ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 681251 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.es.sup.mC.sub.esT.sub.es.- sup.mC.sub.es.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-7a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.esG.sub.es T.sub.esT.sub.es.sup.mC.sub.e 681257 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.- sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-7a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eoG.sub.eo T.sub.esT.sub.es.sup.mC.sub.e
The structure of GalNAc.sub.3-7.sub.a was shown in Example 48.
Treatment
[1033] Female transgenic mice that express human Apo(a) were each injected subcutaneously once per week, for a total of 6 doses, with an oligonucleotide and dosage listed in Table 111b or with PBS. Each treatment group consisted of 3 animals. Blood was drawn the day before dosing to determine baseline levels of Apo(a) protein in plasma and at 72 hours, 1 week, and 2 weeks following the first dose. Additional blood draws will occur at 3 weeks, 4 weeks, 5 weeks, and 6 weeks following the first dose. Plasma Apo(a) protein levels were measured using an ELISA. The results in Table 111b are presented as the average percent of plasma Apo(a) protein levels for each treatment group, normalized to baseline levels (% BL), The results show that the oligonucleotides comprising a GalNAc conjugate group exhibited potent reduction in Apo(a) expression. This potent effect was observed for the oligonucleotide that comprises full PS internucleoside linkages and the oligonucleotide that comprises mixed PO and PS linkages.
TABLE-US-00100 TABLE 111b Apo(a) plasma protein levels Apo(a) at Apo(a) at Apo(a) at 72 hours 1 week 3 weeks ISIS No. Dosage (mg/kg) (% BL) (% BL) (% BL) PBS n/a 116 104 107 681251 0.3 97 108 93 1.0 85 77 57 3.0 54 49 11 10.0 23 15 4 681257 0.3 114 138 104 1.0 91 98 54 3.0 69 40 6 10.0 30 21 4
Example 101: Antisense Inhibition by Oligonucleotides Comprising a GalNAc Cluster Linked Via a Stable Moiety
[1034] The oligonucleotides listed in Table 112 were tested for inhibition of mouse APOC-III expression in vivo. C57Bl/6 mice were each injected subcutaneously once with an oligonucleotide listed in Table 112 or with PBS. Each treatment group consisted of 4 animals. Each mouse treated with ISIS 440670 received a dose of 2, 6, 20, or 60 mg/kg. Each mouse treated with ISIS 680772 or 696847 received 0.6, 2, 6, or 20 mg/kg. The GalNAc conjugate group of ISIS 696847 is linked via a stable moiety, a phosphorothioate linkage instead of a readily cleavable phosphodiester containing linkage. The animals were sacrificed 72 hours after the dose. Liver APOC-III mRNA levels were measured using real-time PCR. APOC-III mRNA levels were normalized to cyclophilin mRNA levels according to standard protocols. The results are presented in Table 112 as the average percent of APOC-III mRNA levels for each treatment group relative to the saline control group. The results show that the oligonucleotides comprising a GalNAc conjugate group were significantly more potent than the oligonucleotide lacking a conjugate group. Furthermore, the oligonucleotide comprising a GalNAc conjugate group linked to the oligonucleotide via a cleavable moiety (ISIS 680772) was even more potent than the oligonucleotide comprising a GalNAc conjugate group linked to the oligonucleotide via a stable moiety (ISIS 696847).
TABLE-US-00101 TABLE 112 Modified oligonucleotides targeting mouse APOC-III APOC-III ISIS Dosage mRNA (% SEQ No. Sequences (5' to 3') CM (mg/kg) PBS) ID No. 440670 .sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.dsT.sub.d- sA.sub.dsT.sub.dsT.sub.dsA.sub.ds n/a 2 92 2275 G.sub.dsG.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.esA.sub.esG.sub.es .sup.mC.sub.esA.sub.e 6 86 20 59 60 37 680772 GalNAc.sub.3-7.sub.a-o'.sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.es- T.sub.esT.sub.dsT.sub.dsA.sub.ds PO 0.6 79 2275 T.sub.dsT.sub.dsA.sub.dsG.sub.dsG.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.esA.s- ub.esG.sub.es.sup.mC.sub.esA.sub.e 2 58 6 31 20 13 696847 GalNAc.sub.3-7.sub.a-s'.sup.mC.sub.esA.sub.esG.sub.es.sup.mC.sub.es- T.sub.esT.sub.dsT.sub.dsA.sub.dsT.sub.ds n/a (PS) 0.6 83 2275 T.sub.dsA.sub.dsG.sub.dsG.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.esA.sub.esG.s- ub.es.sup.mC.sub.esA.sub.e 2 73 6 40 20 28
The structure of GalNAc.sub.3-7a was shown in Example 48.
Example 102: Distribution in Liver of Antisense Oligonucleotides Comprising a GalNAc Conjugate
[1035] The liver distribution of ISIS 353382 (see Table 36) that does not comprise a GalNAc conjugate and ISIS 655861 (see Table 36) that does comprise a GalNAc conjugate was evaluated. Male balb/c mice were subcutaneously injected once with ISIS 353382 or 655861 at a dosage listed in Table 113. Each treatment group consisted of 3 animals except for the 18 mg/kg group for ISIS 655861, which consisted of 2 animals. The animals were sacrificed 48 hours following the dose to determine the liver distribution of the oligonucleotides. In order to measure the number of antisense oligonucleotide molecules per cell, a Ruthenium (II) tris-bipyridine tag (MSD TAG, Meso Scale Discovery) was conjugated to an oligonucleotide probe used to detect the antisense oligonucleotides. The results presented in Table 113 are the average concentrations of oligonucleotide for each treatment group in units of millions of oligonucleotide molecules per cell. The results show that at equivalent doses, the oligonucleotide comprising a GalNAc conjugate was present at higher concentrations in the total liver and in hepatocytes than the oligonucleotide that does not comprise a GalNAc conjugate. Furthermore, the oligonucleotide comprising a GalNAc conjugate was present at lower concentrations in non-parenchymal liver cells than the oligonucleotide that does not comprise a GalNAc conjugate. And while the concentrations of ISIS 655861 in hepatocytes and non-parenchymal liver cells were similar per cell, the liver is approximately 80% hepatocytes by volume. Thus, the majority of the ISIS 655861 oligonucleotide that was present in the liver was found in hepatocytes, whereas the majority of the ISIS 353382 oligonucleotide that was present in the liver was found in non-parenchymal liver cells.
TABLE-US-00102 TABLE 113 Concentration Concentration in Concentration in in whole hepatocytes non-parenchymal liver (molecules* liver cells ISIS Dosage (molecules*10{circumflex over ( )}6 10{circumflex over ( )}6 (molecules*10{circumflex over ( )}6 No. (mg/kg) per cell) per cell) per cell) 353382 3 9.7 1.2 37.2 10 17.3 4.5 34.0 20 23.6 6.6 65.6 30 29.1 11.7 80.0 60 73.4 14.8 98.0 90 89.6 18.5 119.9 655861 0.5 2.6 2.9 3.2 1 6.2 7.0 8.8 3 19.1 25.1 28.5 6 44.1 48.7 55.0 18 76.6 82.3 77.1
Example 103: Duration of Action In Vivo of Oligonucleotides Targeting APOC-III Comprising a GalNAc.sub.3 Conjugate
[1036] The oligonucleotides listed in Table 114 below were tested in a single dose study for duration of action in mice.
TABLE-US-00103 TABLE 114 Modified ASOs targeting APOC-III ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 304801 A.sub.esG.sub.es.sup.mC.sub.esT.sub.esT.sub.es.sup.mC.sub.dsT.sub.d- sT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.- mC.sub.dsT.sub.esT.sub.es n/a n/a 2248 T.sub.esA.sub.esT.sub.e 663084 GalNAc.sub.3-3.sub.a-.sub.o'A.sub.doA.sub.esG.sub.eo.sup.mC.sub.eoT- .sub.eoT.sub.eo.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.d- s GalNAc.sub.3-3a A.sub.d 2264 .sup.mC.sub.dsA.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.eoT.sub.eoT.sub.esA.sub- .esT.sub.e 679241 A.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eoT.sub.eo.sup.mC.sub.dsT.sub.d- sT.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsA.sub.dsG.sub.ds.sup.- mC.sub.dsT.sub.eoT.sub.eo GalNAc.sub.3-19a A.sub.d 2249 T.sub.esA.sub.esT.sub.eoA.sub.do'-GalNAc.sub.3-19.sub.a
The structure of GalNAc.sub.3-3.sub.a was shown in Example 39, and GalNAc.sub.3-19.sub.a was shown in Example 70.
Treatment
[1037] Female transgenic mice that express human APOC-III were each injected subcutaneously once with an oligonucleotide listed in Table 114 or with PBS. Each treatment group consisted of 3 animals. Blood was drawn before dosing to determine baseline and at 3, 7, 14, 21, 28, 35, and 42 days following the dose. Plasma triglyceride and APOC-III protein levels were measured as described in Example 20. The results in Table 115 are presented as the average percent of plasma triglyceride and APOC-III levels for each treatment group, normalized to baseline levels. A comparison of the results in Table 71 of example 79 with the results in Table 115 below show that oligonucleotides comprising a mixture of phosphodiester and phosphorothioate internucleoside linkages exhibited increased duration of action than equivalent oligonucleotides comprising only phosphorothioate internucleoside linkages.
TABLE-US-00104 TABLE 115 Plasma triglyceride and APOC-III protein levels in transgenic mice Time point APOC-III (days protein ISIS Dosage post- Triglycerides (% GalNAc.sub.3 No. (mg/kg) dose) (% baseline) baseline) Cluster CM PBS n/a 3 96 101 n/a n/a 7 88 98 14 91 103 21 69 92 28 83 81 35 65 86 42 72 88 304801 30 3 42 46 n/a n/a 7 42 51 14 59 69 21 67 81 28 79 76 35 72 95 42 82 92 663084 10 3 35 28 GalNAc.sub.3-3a A.sub.d 7 23 24 14 23 26 21 23 29 28 30 22 35 32 36 42 37 47 679241 10 3 38 30 GalNAc.sub.3-19a A.sub.d 7 31 28 14 30 22 21 36 34 28 48 34 35 50 45 42 72 64
Example 104: Synthesis of Oligonucleotides Comprising a 5'-GalNAc.sub.2 Conjugate
##STR00260## ##STR00261##
[1039] Compound 120 is commercially available, and the synthesis of compound 126 is described in Example 49. Compound 120 (1 g, 2.89 mmol), HBTU (0.39 g, 2.89 mmol), and HOBt (1.64 g, 4.33 mmol) were dissolved in DMF (10 mL. and N,N-diisopropylethylamine (1.75 mL, 10.1 mmol) were added. After about 5 min, aminohexanoic acid benzyl ester (1.36 g, 3.46 mmol) was added to the reaction. After 3 h, the reaction mixture was poured into 100 mL of 1 M NaHSO4 and extracted with 2.times.50 mL ethyl acetate. Organic layers were combined and washed with 3.times.40 mL sat NaHCO.sub.3 and 2.times. brine, dried with Na.sub.2SO.sub.4, filtered and concentrated. The product was purified by silica gel column chromatography (DCM:EA:Hex, 1:1:1) to yield compound 231. LCMS and NMR were consistent with the structure. Compounds 231 (1.34 g, 2.438 mmol) was dissolved in dichloromethane (10 mL) and trifluoracetic acid (10 mL) was added. After stirring at room temperature for 2 h, the reaction mixture was concentrated under reduced pressure and co-evaporated with toluene (3.times.10 mL). The residue was dried under reduced pressure to yield compound 232 as the trifluoracetate salt. The synthesis of compound 166 is described in Example 54. Compound 166 (3.39 g, 5.40 mmol) was dissolved in DMF (3 mL). A solution of compound 232 (1.3 g, 2.25 mmol) was dissolved in DMF (3 mL) and N,N-diisopropylethylamine (1.55 mL) was added. The reaction was stirred at room temperature for 30 minutes, then poured into water (80 mL) and the aqueous layer was extracted with EtOAc (2.times.100 mL). The organic phase was separated and washed with sat. aqueous NaHCO.sub.3 (3.times.80 mL), 1 M NaHSO.sub.4 (3.times.80 mL) and brine (2.times.80 mL), then dried (Na.sub.2SO.sub.4), filtered, and concentrated. The residue was purified by silica gel column chromatography to yield compound 233. LCMS and NMR were consistent with the structure. Compound 233 (0.59 g, 0.48 mmol) was dissolved in methanol (2.2 mL) and ethyl acetate (2.2 mL). Palladium on carbon (10 wt % Pd/C, wet, 0.07 g) was added, and the reaction mixture was stirred under hydrogen atmosphere for 3 h. The reaction mixture was filtered through a pad of Celite and concentrated to yield the carboxylic acid. The carboxylic acid (1.32 g, 1.15 mmol, cluster free acid) was dissolved in DMF (3.2 mL). To this N,N-diisopropylethylamine (0.3 mL, 1.73 mmol) and PFPTFA (0.30 mL, 1.73 mmol) were added. After 30 min stirring at room temperature the reaction mixture was poured into water (40 mL) and extracted with EtOAc (2.times.50 mL). A standard work-up was completed as described above to yield compound 234. LCMS and NMR were consistent with the structure. Oligonucleotide 235 was prepared using the general procedure described in Example 46. The GalNAc.sub.2 cluster portion (GalNAc.sub.2-24.sub.a) of the conjugate group GalNAc.sub.2-24 can be combined with any cleavable moiety present on the oligonucleotide to provide a variety of conjugate groups. The structure of GalNAc.sub.2-24 (GalNAc.sub.2-24.sub.a-CM) is shown below:
##STR00262##
Example 105: Synthesis of Oligonucleotides Comprising a GalNAc.sub.1-25 Conjugate
##STR00263##
[1041] The synthesis of compound 166 is described in Example 54. Oligonucleotide 236 was prepared using the general procedure described in Example 46.
[1042] Alternatively, oligonucleotide 236 was synthesized using the scheme shown below, and compound 238 was used to form the oligonucleotide 236 using procedures described in Example 10.
##STR00264##
The GalNAc.sub.1 cluster portion (GalNAc.sub.1-25.sub.a) of the conjugate group GalNAc.sub.1-25 can be combined with any cleavable moiety present on the oligonucleotide to provide a variety of conjugate groups. The structure of GalNAc.sub.1-25 (GalNAc.sub.1-25.sub.a-CM) is shown below:
##STR00265##
Example 106: Antisense Inhibition In Vivo by Oligonucleotides Targeting SRB-1 Comprising a 5'-GalNAc.sub.2 or a 5'-GalNAc.sub.3 Conjugate
[1043] Oligonucleotides listed in Tables 116 and 117 were tested in dose-dependent studies for antisense inhibition of SRB-1 in mice.
Treatment
[1044] Six to week old, male C57BL/6 mice (Jackson Laboratory, Bar Harbor, Me.) were injected subcutaneously once with 2, 7, or 20 mg/kg of ISIS No. 440762; or with 0.2, 0.6, 2, 6, or 20 mg/kg of ISIS No. 686221, 686222, or 708561; or with saline. Each treatment group consisted of 4 animals. The mice were sacrificed 72 hours following the final administration. Liver SRB-1 mRNA levels were measured using real-time PCR. SRB-1 mRNA levels were normalized to cyclophilin mRNA levels according to standard protocols. The antisense oligonucleotides lowered SRB-1 mRNA levels in a dose-dependent manner, and the ED.sub.50 results are presented in Tables 116 and 117. Although previous studies showed that trivalent GalNAc-conjugated oligonucleotides were significantly more potent than divalent GalNAc-conjugated oligonucleotides, which were in turn significantly more potent than monovalent GalNAc conjugated oligonucleotides (see, e.g., Khorev et al., Bioorg. & Med. Chem., Vol. 16, 5216-5231 (2008)), treatment with antisense oligonucleotides comprising monovalent, divalent, and trivalent GalNAc clusters lowered SRB-1 mRNA levels with similar potencies as shown in Tables 116 and 117.
TABLE-US-00105 TABLE 116 Modified oligonucleotides targeting SRB-1 ISIS ED.sub.50 SEQ No. Sequences (5' to 3') GalNAc Cluster (mg/kg) ID No 440762 T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.d- sT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k n/a 4.7 2250 686221 GalNAc.sub.2-24.sub.a-.sub.o'A.sub.doT.sub.ks.sup.mC.sub.ksA.sub.ds- G.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds GalNAc.sub.2-24.sub.a 0.39 2254 .sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k 686222 GalNAc.sub.3-13.sub.a-.sub.o'A.sub.doT.sub.ks.sup.mC.sub.ksA.sub.ds- G.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds GalNAc.sub.3-13.sub.a 0.41 2254 .sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k
See Example 93 for table legend. The structure of GalNAc.sub.3-13a was shown in Example 62, and the structure of GalNAc.sub.2-24a was shown in Example 104.
TABLE-US-00106 TABLE 117 Modified oligonucleotides targeting SRB-1 ISIS ED.sub.50 SEQ No. Sequences (5' to 3') GalNAc Cluster (mg/kg) ID No 440762 T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.dsT.sub.ds.sup.mC.sub.dsA.sub.d- sT.sub.dsG.sub.dsA.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k n/a 5 2250 708561 GalNAc.sub.1-25.sub.a-.sub.o'T.sub.ks.sup.mC.sub.ksA.sub.dsG.sub.ds- T.sub.ds.sup.mC.sub.dsA.sub.dsT.sub.dsG.sub.dsA.sub.ds GalNAc.sub.1-25.sub.a 0.4 2250 .sup.mC.sub.dsT.sub.dsT.sub.ks.sup.mC.sub.k
[1045] See Example 93 for table legend. The structure of GalNAc.sub.1-25a was shown in Example 105. The concentrations of the oligonucleotides in Tables 116 and 117 in liver were also assessed, using procedures described in Example 75. The results shown in Tables 117a and 117b below are the average total antisense oligonucleotide tissues levels for each treatment group, as measured by UV in units of .mu.g oligonucleotide per gram of liver tissue. The results show that the oligonucleotides comprising a GalNAc conjugate group accumulated in the liver at significantly higher levels than the same dose of the oligonucleotide lacking a GalNAc conjugate group. Furthermore, the antisense oligonucleotides comprising one, two, or three GalNAc ligands in their respective conjugate groups all accumulated in the liver at similar levels. This result is surprising in view of the Khorev et al. literature reference cited above and is consistent with the activity data shown in Tables 116 and 117 above.
TABLE-US-00107 TABLE 117a Liver concentrations of oligonucleotides comprising a GalNAc.sub.2 or GalNAc.sub.3 conjugate group Dosage [Antisense GalNAc ISIS No. (mg/kg) oligonucleotide] (.mu.g) cluster CM 440762 2 2.1 n/a n/a 7 13.1 20 31.1 686221 0.2 0.9 GalNAc.sub.2-24.sub.a A.sub.d 0.6 2.7 2 12.0 6 26.5 686222 0.2 0.5 GalNAc.sub.3-13.sub.a A.sub.d 0.6 1.6 2 11.6 6 19.8
TABLE-US-00108 TABLE 117b Liver concentrations of oligonucleotides comprising a GalNAc.sub.1 conjugate group Dosage [Antisense GalNAc ISIS No. (mg/kg) oligonucleotide] (.mu.g) cluster CM 440762 2 2.3 n/a n/a 7 8.9 20 23.7 0.2 0.4 708561 0.6 1.1 GalNAc.sub.1-25.sub.a PO 2 5.9 6 23.7 20 53.9
##STR00266##
[1046] Oligonucleotide 239 is synthesized via coupling of compound 47 (see Example 15) to acid 64 (see Example 32) using HBTU and DIEA in DMF. The resulting amide containing compound is phosphitylated, then added to the 5'-end of an oligonucleotide using procedures described in Example 10. The GalNAc.sub.1 cluster portion (GalNAc.sub.1-26.sub.a) of the conjugate group GalNAc.sub.1-26 can be combined with any cleavable moiety present on the oligonucleotide to provide a variety of conjugate groups. The structure of GalNAc.sub.1-26 (GalNAc.sub.1-26.sub.a-CM) is shown below:
##STR00267##
[1047] In order to add the GalNAc.sub.1 conjugate group to the 3'-end of an oligonucleotide, the amide formed from the reaction of compounds 47 and 64 is added to a solid support using procedures described in Example 7. The oligonucleotide synthesis is then completed using procedures described in Example 9 in order to form oligonucleotide 240.
##STR00268##
The GalNAc.sub.1 cluster portion (GalNAc.sub.1-27.sub.a) of the conjugate group GalNAc.sub.1-27 can be combined with any cleavable moiety present on the oligonucleotide to provide a variety of conjugate groups. The structure of GalNAc.sub.1-27 (GalNAc.sub.1-27.sub.a-CM) is shown below:
##STR00269##
Example 108: Antisense Inhibition In Vivo by Oligonucleotides Comprising a GalNAc Conjugate Group Targeting Apo(a) In Vivo
[1048] The oligonucleotides listed in Table 118 below were tested in a single dose study in mice.
TABLE-US-00109 TABLE 118 Modified ASOs targeting APO(a) ISIS GalNAc.sub.3 SEQ No. Sequences (5' to 3') Cluster CM ID No. 494372 T.sub.esG.sub.es.sup.mC.sub.esT.sub.es.sup.mC.sub.es.sup.mC.sub.dsG- .sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.ds n/a n/a 2281 T.sub.dsT.sub.esG.sub.esT.sub.esT.sub.es.sup.mC.sub.e 681251 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.es.sup.mC.sub.esT.sub.es.- sup.mC.sub.es.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-7a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.esG.sub.esT.sub.esT.sub.es.su- p.mC.sub.e 681255 GalNAc.sub.3-3.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.- sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-3a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eoG.sub.eoT.sub.esT.sub.es.su- p.mC.sub.e 681256 GalNAc.sub.3-10.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo- .sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-10a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eoG.sub.eoT.sub.esT.sub.es.su- p.mC.sub.e 681257 GalNAc.sub.3-7.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.- sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-7a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eoG.sub.eoT.sub.esT.sub.es.su- p.mC.sub.e 681258 GalNAc.sub.3-13.sub.a-.sub.o'T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo- .sup.mC.sub.eo.sup.mC.sub.dsG.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds GalNAc.sub.3-13a PO 2281 T.sub.dsG.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.eoG.sub.eoT.sub.esT.sub.es.su- p.mC.sub.e 681260 T.sub.esG.sub.eo.sup.mC.sub.eoT.sub.eo.sup.mC.sub.eo.sup.mC.sub.dsG- .sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.dsT.sub.dsG.sub.ds.sup.mC.sub.dsT.sub- .dsT.sub.eoG.sub.eo GalNAc.sub.3-19a A.sub.d 2280 T.sub.esT.sub.es.sup.mC.sub.eoA.sub.do'-GalNAc.sub.3-19
The structure of GalNAc.sub.3-7.sub.a was shown in Example 48.
Treatment
[1049] Male transgenic mice that express human Apo(a) were each injected subcutaneously once with an oligonucleotide and dosage listed in Table 119 or with PBS. Each treatment group consisted of 4 animals. Blood was drawn the day before dosing to determine baseline levels of Apo(a) protein in plasma and at 1 week following the first dose. Additional blood draws will occur weekly for approximately 8 weeks. Plasma Apo(a) protein levels were measured using an ELISA. The results in Table 119 are presented as the average percent of plasma Apo(a) protein levels for each treatment group, normalized to baseline levels (% BL), The results show that the antisense oligonucleotides reduced Apo(a) protein expression. Furthermore, the oligonucleotides comprising a GalNAc conjugate group exhibited even more potent reduction in Apo(a) expression than the oligonucleotide that does not comprise a conjugate group.
TABLE-US-00110 TABLE 119 Apo(a) plasma protein levels Dosage Apo(a) at 1 week ISIS No. (mg/kg) (% BL) PBS n/a 143 494372 50 58 681251 10 15 681255 10 14 681256 10 17 681257 10 24 681258 10 22 681260 10 26
Example 109: Synthesis of Oligonucleotides Comprising a GalNAc.sub.1-28 or GalNAc.sub.1-29 Conjugate
##STR00270##
[1051] Oligonucleotide 241 is synthesized using procedures similar to those described in Example 71 to form the phosphoramidite intermediate, followed by procedures described in Example 10 to synthesize the oligonucleotide. The GalNAc.sub.1 cluster portion (GalNAc.sub.1-28.sub.a) of the conjugate group GalNAc.sub.1-28 can be combined with any cleavable moiety present on the oligonucleotide to provide a variety of conjugate groups. The structure of GalNAc.sub.1-28 (GalNAc.sub.1-28.sub.a-CM) is shown below:
##STR00271##
[1052] In order to add the GalNAc.sub.1 conjugate group to the 3'-end of an oligonucleotide, procedures similar to those described in Example 71 are used to form the hydroxyl intermediate, which is then added to the solid support using procedures described in Example 7. The oligonucleotide synthesis is then completed using procedures described in Example 9 in order to form oligonucleotide 242.
##STR00272##
The GalNAc.sub.1 cluster portion (GalNAc.sub.1-29.sub.a) of the conjugate group GalNAc.sub.1-29 can be combined with any cleavable moiety present on the oligonucleotide to provide a variety of conjugate groups. The structure of GalNAc.sub.1-29 (GalNAc.sub.1-29-CM) is shown below:
##STR00273##
Example 110: Synthesis of Oligonucleotides Comprising a GalNAc.sub.1-30 Conjugate
##STR00274##
[1054] Oligonucleotide 246 comprising a GalNAc.sub.1-30 conjugate group, wherein Y is selected from O, S, a substituted or unsubstituted C.sub.1-C.sub.10 alkyl, amino, substituted amino, azido, alkenyl or alkynyl, is synthesized as shown above. The GalNAc.sub.1 cluster portion (GalNAc.sub.1-30.sub.a) of the conjugate group GalNAc.sub.1-30 can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, Y is part of the cleavable moiety. In certain embodiments, Y is part of a stable moiety, and the cleavable moiety is present on the oligonucleotide. The structure of GalNAc.sub.1-30.sub.a is shown below:
##STR00275##
Example 111: Synthesis of Oligonucleotides Comprising a GalNAc.sub.2-31 or GalNAc.sub.2-32 Conjugate
##STR00276##
[1056] Oligonucleotide 250 comprising a GalNAc.sub.2-31 conjugate group, wherein Y is selected from O, S, a substituted or unsubstituted C.sub.1-C.sub.10 alkyl, amino, substituted amino, azido, alkenyl or alkynyl, is synthesized as shown above. The GalNAc.sub.2 cluster portion (GalNAc.sub.2-31.sub.a) of the conjugate group GalNAc.sub.2-31 can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the Y-containing group directly adjacent to the 5'-end of the oligonucleotide is part of the cleavable moiety. In certain embodiments, the Y-containing group directly adjacent to the 5'-end of the oligonucleotide is part of a stable moiety, and the cleavable moiety is present on the oligonucleotide. The structure of GalNAc.sub.2-31.sub.a is shown below:
##STR00277##
[1057] The synthesis of an oligonucleotide comprising a GalNAc.sub.2-32 conjugate is shown below.
##STR00278##
[1058] Oligonucleotide 252 comprising a GalNAc.sub.2-32 conjugate group, wherein Y is selected from O, S, a substituted or unsubstituted C.sub.1-C.sub.10 alkyl, amino, substituted amino, azido, alkenyl or alkynyl, is synthesized as shown above. The GalNAc.sub.2 cluster portion (GalNAc.sub.2-32.sub.a) of the conjugate group GalNAc.sub.2-32 can be combined with any cleavable moiety to provide a variety of conjugate groups. In certain embodiments, the Y-containing group directly adjacent to the 5'-end of the oligonucleotide is part of the cleavable moiety. In certain embodiments, the Y-containing group directly adjacent to the 5'-end of the oligonucleotide is part of a stable moiety, and the cleavable moiety is present on the oligonucleotide. The structure of GalNAc.sub.2-32a is shown below:
##STR00279##
Example 112: Modified Oligonucleotides Comprising a GalNAc.sub.1 Conjugate
[1059] The oligonucleotides in Table 120 targeting SRB-1 were synthesized with a GalNAc.sub.1 conjugate group in order to further test the potency of oligonucleotides comprising conjugate groups that contain one GalNAc ligand.
TABLE-US-00111 TABLE 120 GalNAc SEQ ISIS No. Sequence (5' to 3') cluster CM ID NO. 711461 GalNAc.sub.1-25.sub.a-o'A.sub.do G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds GalNAc.sub.1-25.sub.a A.sub.d 2258 T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.e 711462 GalNAc.sub.1-25.sub.a-o'G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.1-25.sub.a PO 2256 G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.e 711463 GalNAc.sub.1-25.sub.a-o'G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.1-25.sub.a PO 2256 G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.e 711465 GalNAc.sub.1-26.sub.a-o'A.sub.do G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds GalNAc.sub.1-26.sub.a A.sub.d 2258 T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.e 711466 GalNAc.sub.1-26.sub.a-o'G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.1-26.sub.a PO 2256 G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.e 711467 GalNAc.sub.1-26.sub.a-o'G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.1-26.sub.a PO 2256 G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.e 711468 GalNAc.sub.1-28.sub.a-o'A.sub.do G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds GalNAc.sub.1-28.sub.a A.sub.d 2258 T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.e 711469 GalNAc.sub.1-28.sub.a-o'G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.1-28.sub.a PO 2256 G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.e 711470 GalNAc.sub.1-28.sub.a-o'G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds GalNAc.sub.1-28.sub.a PO 2256 G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.e 713844 G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-27.sub.a PO 2256 T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.eo'-GalNAc.sub.1-27.sub.a 713845 G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-27.sub.a PO 2256 T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.eo'-GalNAc.sub.1-27.sub.a 713846 G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-27.sub.a A.sub.d 2257 T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.eo A.sub.do'-GalNAc.sub.1-27.sub.a 713847 G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-29.sub.a PO 2256 T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.eo'-GalNAc.sub.1-29.sub.a 713848 G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-29.sub.a PO 2256 T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.eo'-GalNAc.sub.1-29.sub.a 713849 G.sub.es .sup.mC.sub.es T.sub.es T.sub.es .sup.mC.sub.es A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-29.sub.a A.sub.d 2257 T.sub.es .sup.mC.sub.es .sup.mC.sub.es T.sub.es T.sub.eo A.sub.do'-GalNAc.sub.1-29.sub.a 713850 G.sub.es .sup.mC.sub.eo T.sub.eo T.sub.eo .sup.mC.sub.eo A.sub.ds G.sub.ds T.sub.ds .sup.mC.sub.ds A.sub.ds T.sub.ds G.sub.ds A.sub.ds .sup.mC.sub.ds T.sub.ds GalNAc.sub.1-29.sub.a A.sub.d 2257 T.sub.eo .sup.mC.sub.eo .sup.mC.sub.es T.sub.es T.sub.eo A.sub.do'-GalNAc.sub.1-29.sub.a
Example 113: Antisense Oligonucleotides Targeting Kallikrein B, Plasma (Fletcher Factor) 1 Comprising a GalNAc Cluster
[1060] The oligonucleotides in Table 121 were designed to target human kallikrein B, plasma (Fletcher factor) 1, or prekallikrein (PKK).
TABLE-US-00112 TABLE 121 Sequences (5' to 3') SEQ ID No. GalNAc.sub.3-3-T.sub.esG.sub.es.sup.mC.sub.esA.sub.esA.sub.esG.sub.dsT.sub- .ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.su- p.mC.sub.dsA.sub.esA.sub.esA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-3-T.sub.esG.sub.eo.sup.mC.sub.eoA.sub.eoA.sub.eoG.sub.dsT.sub- .ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.su- p.mC.sub.dsA.sub.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-7-T.sub.esG.sub.es.sup.mC.sub.esA.sub.esA.sub.esG.sub.dsT.sub- .ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.su- p.mC.sub.dsA.sub.esA.sub.esA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-7-T.sub.esG.sub.eo.sup.mC.sub.eoA.sub.eoA.sub.eoG.sub.dsT.sub- .ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.su- p.mC.sub.dsA.sub.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-10-T.sub.esG.sub.es.sup.mC.sub.esA.sub.esA.sub.esG.sub.dsT.su- b.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.s- up.mC.sub.dsA.sub.esA.sub.esA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-10-T.sub.esG.sub.eo.sup.mC.sub.eoA.sub.eoA.sub.eoG.sub.dsT.su- b.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.s- up.mC.sub.dsA.sub.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-13-T.sub.esG.sub.es.sup.mC.sub.esA.sub.esA.sub.esG.sub.dsT.su- b.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.s- up.mC.sub.dsA.sub.esA.sub.esA.sub.es.sup.mC.sub.esA.sub.e 570 GalNAc.sub.3-13-T.sub.esG.sub.eo.sup.mC.sub.eoA.sub.eoA.sub.eoG.sub.dsT.su- b.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.s- up.mC.sub.dsA.sub.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e 570 T.sub.esG.sub.es.sup.mC.sub.esA.sub.esA.sub.esG.sub.dsT.sub.ds.sup.mC.sub.- dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.sup.mC.sub.dsA.su- b.esA.sub.esA.sub.es.sup.mC.sub.esA.sub.e-GalNAc.sub.3-19 570 T.sub.esG.sub.eo.sup.mC.sub.eoA.sub.eoA.sub.eoG.sub.dsT.sub.ds.sup.mC.sub.- dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds.sup.mC.sub.dsA.su- b.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e-GalNAc.sub.3-19 570 GalNAc.sub.3-7.sub.a-o'T.sub.esG.sub.es.sup.mC.sub.eoA.sub.eoA.sub.es G.sub.ds T.sub.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.su- b.dsG.sub.ds.sup.mC.sub.dsA.sub.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e 570
Example 114: Antisense Inhibition of Human PKK in HepaRG.TM..sup.T Cells by Antisense Oligonucleotides with 2'-MOE Sugar Modifications
[1061] Antisense oligonucleotides were designed targeting a PKK nucleic acid and were tested for their effects on PKK mRNA in vitro. HepaRG.TM. cells, which are terminally differentiated hepatic cells derived from a human hepatic progenitor cell line and retain many characteristics of primary human hepatocytes (Lubberstedt M. et al., J. Pharmacol. Toxicol. Methods 2011 63: 59-68), were used in the screen.
[1062] The chimeric antisense oligonucleotides in the tables below were designed as 5-10-5 MOE gapmers. The gapmers are 20 nucleosides in length, wherein the central gap segment comprises often 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising five nucleosides each. Each nucleoside in the 5' wing segment and each nucleoside in the 3' wing segment has a 2'-O-methoxyethyl modification. The internucleoside linkages throughout each gapmer are phosphorothioate linkages. All cytosine residues throughout each gapmer are 5-methylcytosines. "Start site" indicates the 5'-most nucleoside to which the gapmer is targeted in the human gene sequence. "Stop site" indicates the 3'-most nucleoside to which the gapmer is targeted in the human gene sequence. Each gapmer listed in the tables below is targeted to either the human PKK mRNA, designated herein as SEQ ID NO: 1 (GENBANK Accession No. NM_000892.3) or the human PKK genomic sequence, designated herein as SEQ ID NO: 10 (GENBANK Accession No. NT_016354.19 truncated from nucleotides 111693001 to Ser. No. 11/730,000). `n/a` indicates that the antisense oligonucleotide does not target that particular gene sequence.
[1063] Cultured HepaRG.TM. cells at a density of 20,000 cells per well were transfected using electroporation with 3,000 nM antisense oligonucleotide. After a treatment period of approximately 24 hours, RNA was isolated from the cells and PKK mRNA levels were measured by quantitative real-time PCR. Human primer probe set RTS3454 (forward sequence CCAAAAAAGGTGCACCAGTAACA, designated herein as SEQ ID NO: 20; reverse sequence CCTCCGGGACTGTACTTTAATAGG, designated herein as SEQ ID NO: 21; probe sequence CACGCAAACATTTCACAAGGCAGAGTACC, designated herein as SEQ ID NO: 22) was used to measure mRNA levels. PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.RTM.. The antisense oligonucleotides were tested in a series of experiments that had similar culture conditions. The results for each experiment are presented in separate tables shown below. Results are presented as percent inhibition of PKK, relative to untreated control cells.
TABLE-US-00113 TABLE 1 SEQ SEQ ID ID SEQ ID SEQ ID NO: 1 NO: 1 NO: 10 NO: 10 SEQ Start Stop % Start Stop ID ISIS NO Site Site Sequence inhibition Site Site NO 530929 1 20 AACGGTCTTCAAGCTGTTCT 59 3393 3412 30 530930 6 25 AAATGAACGGTCTTCAAGCT 17 3398 3417 31 530931 11 30 CTTAAAAATGAACGGTCTTC 29 3403 3422 32 530932 16 35 TGTCACTTAAAAATGAACGG 52 3408 3427 33 530933 31 50 TGGAGGTGAGTCTCTTGTCA 76 3423 3442 34 530934 36 55 CTTCTTGGAGGTGAGTCTCT 54 3428 3447 35 530935 68 87 GCTTGAATAAAATCATTCTG 0 n/a n/a 36 530936 73 92 TGCTTGCTTGAATAAAATCA 27 4072 4091 37 530937 78 97 TAAGTTGCTTGCTTGAATAA 0 4077 4096 38 530938 88 107 GGAAATGAAATAAGTTGCTT 11 4087 4106 39 530939 93 112 AACAAGGAAATGAAATAAGT 0 4092 4111 40 530940 98 117 TAGCAAACAAGGAAATGAAA 7 4097 4116 41 530941 103 122 AACTGTAGCAAACAAGGAAA 22 4102 4121 42 530942 108 127 CAGGAAACTGTAGCAAACAA 22 4107 4126 43 530943 113 132 ATCCACAGGAAACTGTAGCA 56 n/a n/a 44 530944 118 137 CAGACATCCACAGGAAACTG 0 n/a n/a 45 530945 157 176 ATCCCCACCTCTGAAGAAGG 0 8029 8048 46 530946 160 179 TACATCCCCACCTCTGAAGA 0 8032 8051 47 530947 165 184 GAAGCTACATCCCCACCTCT 27 8037 8056 48 530948 170 189 ACATGGAAGCTACATCCCCA 35 8042 8061 49 530949 175 194 GGTGTACATGGAAGCTACAT 31 8047 8066 50 530950 221 240 ACCTTGGGTGGAATGTGCAC 47 8093 8112 51 530951 226 245 CAAACACCTTGGGTGGAATG 49 8098 8117 52 530952 234 253 CTGAATAGCAAACACCTTGG 38 8106 8125 53 530953 239 258 GAAAACTGAATAGCAAACAC 7 8111 8130 54 530954 244 263 TGGAAGAAAACTGAATAGCA 47 8116 8135 55 530955 278 297 CAAACCTTTTCTCCATGTCA 55 n/a n/a 56 530956 300 319 ACACTATCTTTCAAGAAGCA 57 9834 9853 57 530957 386 405 GGCAAGCACTTATTTGATGA 56 n/a n/a 58 530958 432 451 TTAAAATTGACTCCTCTCAT 60 12688 12707 59 530959 456 475 TCAACACTGCTAACCTTAGA 60 12712 12731 60 530960 461 480 ATTCTTCAACACTGCTAACC 58 12717 12736 61 530961 466 485 TTGGCATTCTTCAACACTGC 88 12722 12741 62 530962 472 491 CCTTTTTTGGCATTCTTCAA 64 12728 12747 63 530963 479 498 TGGTGCACCTTTTTTGGCAT 78 12735 12754 64 530964 628 647 CTTCAGTGAGAATCCAGATT 44 14199 14218 65 530965 637 656 GGCACAGGGCTTCAGTGAGA 73 14208 14227 66 530966 649 668 AATTTCTGAAAGGGCACAGG 58 14220 14239 67 530967 654 673 CAACCAATTTCTGAAAGGGC 69 n/a n/a 68 530968 680 699 CAAGATGCTGGAAGATGTTC 18 26128 26147 69 530969 846 865 GTGCCACTTTCAGATGTTTT 0 27110 27129 70 530970 851 870 TTGGTGTGCCACTTTCAGAT 74 27115 27134 71 530971 856 875 GGAACTTGGTGTGCCACTTT 85 27120 27139 72 530972 861 880 GTAGAGGAACTTGGTGTGCC 42 27125 27144 73 530973 866 885 GAGGAGTAGAGGAACTTGGT 52 27130 27149 74 530974 871 890 TTCTTGAGGAGTAGAGGAAC 18 27135 27154 75 530975 876 895 GTGTTTTCTTGAGGAGTAGA 41 27140 27159 76 530976 881 900 ATATGGTGTTTTCTTGAGGA 26 27145 27164 77 530977 886 905 TCCAGATATGGTGTTTTCTT 55 27150 27169 78 530978 891 910 CTATATCCAGATATGGTGTT 0 27155 27174 79 530979 901 920 GGTTAAAAGGCTATATCCAG 35 27165 27184 80 530980 906 925 TTGCAGGTTAAAAGGCTATA 29 27170 27189 81 530981 911 930 TTCTTTTGCAGGTTAAAAGG 0 27175 27194 82 530982 916 935 TAAAGTTCTTTTGCAGGTTA 0 27180 27199 83 530983 931 950 ATGGCAGGGTTCAGGTAAAG 9 n/a n/a 84 530984 936 955 TTAGAATGGCAGGGTTCAGG 25 n/a n/a 85 530985 941 960 AAATTTTAGAATGGCAGGGT 32 27363 27382 86 530986 946 965 CGGGTAAATTTTAGAATGGC 62 27368 27387 87 530987 951 970 ACTCCCGGGTAAATTTTAGA 0 27373 27392 88 530988 961 980 TCCAAAGTCAACTCCCGGGT 76 27383 27402 89 530989 966 985 TCTCCTCCAAAGTCAACTCC 28 27388 27407 90 530990 971 990 ATTCTTCTCCTCCAAAGTCA 32 27393 27412 91 530991 976 995 ATTCAATTCTTCTCCTCCAA 43 27398 27417 92 530992 981 1000 GTCACATTCAATTCTTCTCC 70 27403 27422 93 530993 1005 1024 CAAACATTCACTCCTTTAAC 30 27427 27446 94 530994 1010 1029 CTTGGCAAACATTCACTCCT 50 27432 27451 95 530995 1015 1034 AGTCTCTTGGCAAACATTCA 49 27437 27456 96 530996 1038 1057 TGACAGCGAATCATCTTTGT 51 27460 27479 97 530997 1043 1062 AAAACTGACAGCGAATCATC 39 27465 27484 98 530998 1048 1067 AGTGAAAAACTGACAGCGAA 0 27470 27489 99 530999 1071 1090 CAGTCTTCTGGGAGTAAAGA 31 27493 27512 100 531000 1098 1117 AAGAAACACTTACACTTCTC 1 n/a n/a 101 531001 1108 1127 AGATAATCTTAAGAAACACT 44 27629 27648 102 531002 1155 1174 GAGCTCCCTTGTGTCCCATA 85 27676 27695 103 531003 1160 1179 AACCAGAGCTCCCTTGTGTC 49 27681 27700 104 531004 1165 1184 AGAGTAACCAGAGCTCCCTT 76 27686 27705 105 531005 1170 1189 CTCAAAGAGTAACCAGAGCT 76 27691 27710 106 531006 1216 1235 GCTTGTTTTTGTTGTGCAGA 49 27892 27911 107
TABLE-US-00114 TABLE 123 SEQ ID SEQ ID SEQ ID SEQ ID NO: 1 NO: 1 NO: 10 NO: 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence inhibition Site Site NO 482586 1608 1627 ACCCAACAGTTGGTATAAAT 0 31914 31933 108 486847 1563 1582 AGGCATATTGGTTTTTGGAA 78 31869 31888 109 531007 46 65 AACACAATTGCTTCTTGGAG 51 3438 3457 110 531008 675 694 TGCTGGAAGATGTTCATGTG 51 26123 26142 111 531009 1239 1258 TTTGTTCCTCCAACAATGCG 65 27915 27934 112 531010 1244 1263 AAGAGTTTGTTCCTCCAACA 52 27920 27939 113 531011 1249 1268 CCAAGAAGAGTTTGTTCCTC 0 27925 27944 114 531012 1254 1273 TCTCCCCAAGAAGAGTTTGT 48 27930 27949 115 531013 1264 1283 CCAGGGCCACTCTCCCCAAG 56 27940 27959 116 531014 1287 1306 AGCTTCACCTGCAGGCTCAC 0 27963 27982 117 531015 1324 1343 TATGAGTGACCCTCCACACA 52 28000 28019 118 531016 1329 1348 TGTCCTATGAGTGACCCTCC 39 28005 28024 119 531017 1334 1353 ACTGGTGTCCTATGAGTGAC 31 28010 28029 120 531018 1339 1358 GACCCACTGGTGTCCTATGA 54 28015 28034 121 531019 1344 1363 GTGAGGACCCACTGGTGTCC 28 28020 28039 122 531020 1369 1388 AAGCCCATCAAAGCAGTGGG 0 n/a n/a 123 531021 1420 1439 GTCTGACAGATTTAAAATGC 50 30498 30517 124 531022 1425 1444 GTAATGTCTGACAGATTTAA 74 30503 30522 125 531023 1430 1449 CTTTTGTAATGTCTGACAGA 71 30508 30527 126 531024 1452 1471 TTTATTTGTGAGAAAGGTGT 69 30530 30549 127 531025 1457 1476 TCTCTTTTATTTGTGAGAAA 34 30535 30554 128 531026 1501 1520 ATCATGATTCCCTTCTGAGA 73 30579 30598 129 531027 1530 1549 AAAGGAGCCTGGAGTTTTAT 0 30608 30627 130 531028 1535 1554 AATTCAAAGGAGCCTGGAGT 56 30613 30632 131 531029 1540 1559 AGTGTAATTCAAAGGAGCCT 59 30618 30637 132 531030 1545 1564 AATTCAGTGTAATTCAAAGG 24 n/a n/a 133 531031 1550 1569 TTTGGAATTCAGTGTAATTC 59 n/a n/a 134 531032 1555 1574 TGGTTTTTGGAATTCAGTGT 67 n/a n/a 135 531033 1557 1576 ATTGGTTTTTGGAATTCAGT 53 n/a n/a 136 531034 1560 1579 CATATTGGTTTTTGGAATTC 36 31866 31885 137 531035 1565 1584 GTAGGCATATTGGTTTTTGG 46 31871 31890 138 531036 1581 1600 GTGTCACCTTTGGAAGGTAG 71 31887 31906 139 531037 1604 1623 AACAGTTGGTATAAATTGTG 35 31910 31929 140 531038 1605 1624 CAACAGTTGGTATAAATTGT 22 31911 31930 141 531039 1609 1628 TACCCAACAGTTGGTATAAA 36 31915 31934 142 531040 1632 1651 TCCTTCGAGAAGCCCCATCC 27 31938 31957 143 531041 1677 1696 AAAGGAATATTTACCTTTTG 68 33121 33140 144 531042 1682 1701 TTACCAAAGGAATATTTACC 11 33126 33145 145 531043 1687 1706 ATTTGTTACCAAAGGAATAT 27 33131 33150 146 531044 1697 1716 GGCATTCTTCATTTGTTACC 68 33141 33160 147 531045 1702 1721 TTTCTGGCATTCTTCATTTG 37 33146 33165 148 531046 1709 1728 GATATCTTTTCTGGCATTCT 54 33153 33172 149 531047 1714 1733 ATCTTGATATCTTTTCTGGC 68 33158 33177 150 531048 1719 1738 TTATAATCTTGATATCTTTT 42 33163 33182 151 531049 1724 1743 TTATTTTATAATCTTGATAT 2 33168 33187 152 531050 1729 1748 TTGGGTTATTTTATAATCTT 18 33173 33192 153 531051 1734 1753 ATCCGTTGGGTTATTTTATA 51 33178 33197 154 531052 1739 1758 AGACCATCCGTTGGGTTATT 60 33183 33202 155 531053 1744 1763 AGCACAGACCATCCGTTGGG 49 33188 33207 156 531054 1754 1773 CTTTATAGCCAGCACAGACC 48 33198 33217 157 531055 1759 1778 CCCTTCTTTATAGCCAGCAC 68 33203 33222 158 531056 1764 1783 TTTCCCCCTTCTTTATAGCC 45 33208 33227 159 531057 1769 1788 CATCTTTTCCCCCTTCTTTA 48 33213 33232 160 531058 1779 1798 CCCTTACAAGCATCTTTTCC 60 n/a n/a 161 531059 1820 1839 ACATTCCATTGTGTTTGCAA 55 33919 33938 162 531060 1841 1860 TGGTGATGCCCACCAAACGC 35 33940 33959 163 531061 1872 1891 TGCTCCCTGCGGGCACAGCC 52 33971 33990 164 531062 1877 1896 CAGGTTGCTCCCTGCGGGCA 39 33976 33995 165 531063 1882 1901 GACACCAGGTTGCTCCCTGC 51 33981 34000 166 531064 1887 1906 GTGTAGACACCAGGTTGCTC 56 33986 34005 167 531065 1892 1911 CTTTGGTGTAGACACCAGGT 57 33991 34010 168 531066 1897 1916 AGCGACTTTGGTGTAGACAC 67 33996 34015 169 531067 1902 1921 TACTCAGCGACTTTGGTGTA 31 34001 34020 170 531068 1907 1926 CCATGTACTCAGCGACTTTG 59 34006 34025 171 531069 1912 1931 CCAGTCCATGTACTCAGCGA 56 34011 34030 172 531070 1930 1949 CTGTGTTTTCTCTAAAATCC 68 34029 34048 173 531071 1935 1954 CTGCTCTGTGTTTTCTCTAA 73 34034 34053 174 531072 2026 2045 GCTCAGAATTTGACTTGAAC 64 34125 34144 175 531073 2031 2050 CCCAGGCTCAGAATTTGACT 51 34130 34149 176 531074 2049 2068 CTTTGCAGATGAGGACCCCC 67 34148 34167 177 531075 2054 2073 CCATGCTTTGCAGATGAGGA 64 34153 34172 178 531076 2059 2078 ACTCTCCATGCTTTGCAGAT 68 34158 34177 179 531077 2064 2083 ATGCCACTCTCCATGCTTTG 51 34163 34182 180 531078 2111 2130 AGCAGCTCTGAGTGCACTGT 77 34210 34229 181 531079 2116 2135 TCCTCAGCAGCTCTGAGTGC 58 34215 34234 182 531080 2121 2140 CATTGTCCTCAGCAGCTCTG 55 34220 34239 183 531081 n/a n/a TGGTTTTTGGAATTCTGAAA 14 31861 31880 184 531082 n/a n/a ATATTGGTTTTTGGAATTCT 31 31865 31884 185
TABLE-US-00115 TABLE 124 SEQ ID SEQ ID SEQ ID SEQ ID NO: 1 NO: 1 NO: 10 NO: 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence inhibition Site Site NO 531083 n/a n/a TGTACTAGTTTCCTATAACT 60 14738 14757 186 14809 14828 14880 14899 14939 14958 15071 15090 15214 15233 15286 15305 15345 15364 15477 15496 15549 15568 15607 15626 15679 15698 15809 15828 15881 15900 15939 15958 531084 n/a n/a ATAGGGACACAACCAAGGAA 25 16296 16315 187 531085 n/a n/a AGGCACAGAGCCAGCACCCA 9 16495 16514 188 531086 n/a n/a CCTGCCTCCTGGCAGCCTTC 48 16696 16715 189 531087 n/a n/a CCAGGTGTGGACAGCAGCTG 52 16821 16840 190 531088 n/a n/a GGTTTTGTTTGTAAAATTAG 27 17159 17178 191 531089 n/a n/a AAAACACCATTAAATCCATT 45 17306 17325 192 531090 n/a n/a ACAGAAACCATGATGTTGCT 59 17644 17663 193 531091 n/a n/a TCAGCCCAATGTCCTAACCT 35 17793 17812 194 531092 n/a n/a CCTTCACTGACTCTCTTTTC 24 17922 17941 195 531093 n/a n/a TTCTCCTGGCTCAGAAGCTC 60 18053 18072 196 23315 23334 531094 n/a n/a GAATGTCAGGCCTCTGGGCC 48 18181 18200 197 531095 n/a n/a CTAACAACCCCACAATATCA 20 18390 18409 198 531096 n/a n/a CCCAATTCTTAGTCCTTTAA 45 18523 18542 199 531097 n/a n/a ACCAAGCTCAGCCTCCAACT 41 18648 18667 200 531098 n/a n/a TTATTAGTCAAATCACCCAA 19 18773 18792 201 531099 n/a n/a TGGATGGGTAGAGGCCTTTC 64 18898 18917 202 531100 n/a n/a CCCCCTCCCTTCCCTACACA 0 19023 19042 203 531101 n/a n/a ATGTAAGTTACAAGCCACTA 37 19153 19172 204 531102 n/a n/a TGCCTCTTTAATAAAAACTC 42 19484 19503 205 531103 n/a n/a ACTCATTGCCTTAACTCAGG 40 19636 19655 206 531104 n/a n/a ACTTGACCTTACTGTTTTAG 20 19886 19905 207 531105 n/a n/a CTCCTCCCCAGGCTGCTCCT 16 22092 22111 208 531106 n/a n/a AAGATCTAGATAATTCTTGT 31 22332 22351 209 531107 n/a n/a TCAACTCACACCTGACCTAA 30 22457 22476 210 531108 n/a n/a TGAACCCAAAACTCTGGCAC 50 22771 22790 211 531109 n/a n/a AGCCCAAGGAACATCTCACC 52 22959 22978 212 531110 n/a n/a GCCTGTTTGGTGGTCTCTTC 86 23110 23129 213 531111 n/a n/a CTTCTCCTGGCTCAGAAGCT 68 18054 18073 214 23316 23335 531112 n/a n/a ATGTATGATTCTAAGAACTT 14 23479 23498 215 531113 n/a n/a AACAGACACATTATTTATAT 0 23604 23623 216 531114 n/a n/a AGAGTCAAGTCCACAGACAT 40 24246 24265 217 531115 n/a n/a TCCTAAATAGGAACAAAGTA 0 24372 24391 218 531116 n/a n/a TTGTTAAGGTTGTAGAGAGA 23 24688 24707 219 531117 n/a n/a ACCCAATTATTTTTAATGGC 62 24876 24895 220 531118 n/a n/a GCCTAAATGTAAGAGCTAAA 26 25157 25176 221 531119 n/a n/a TAAACTCTTACATTTATAGA 0 25293 25312 222 531120 n/a n/a AAATAAAAGCACTCAGACTG 0 25418 25437 223 531121 n/a n/a TTGGTCTACAGATTCAATGC 72 25550 25569 224 531122 n/a n/a TAACAAAAATGCCTTGTGCC 33 25710 25729 225 531123 n/a n/a TCCCAGCTCCAGTCACCACC 74 25866 25885 226 531124 n/a n/a GTACTAAACATCCTAAGTGA 2 25992 26011 227 531125 n/a n/a ACTCGCCTTTGTGACTCGAT 23 26264 26283 228 531126 n/a n/a TTTTGAATCTTCATTCAAAG 0 26551 26570 229 531127 n/a n/a CAGAGCCTTGATCAGAATAA 12 26676 26695 230 531128 n/a n/a AAGTTCCACCTTCTAACTGG 18 26831 26850 231 531129 n/a n/a AGCAGCTCACACCCAAAAAG 0 27005 27024 232 531130 n/a n/a TTCTGTGTCAATTATAAACA 0 27344 27363 233 531131 n/a n/a TAGAAAGAGTAAGCCTTCAC 0 27587 27606 234 531132 n/a n/a AGTGAGGTTACTCACCAGAG 0 27732 27751 235 531133 n/a n/a TTTTGTTGTGCAGACTGAAA 19 27886 27905 236 531134 n/a n/a TTACCCATCAAAGCAGTGGG 6 28045 28064 237 531135 n/a n/a AATGTTGTGAATACCATCCC 16 28174 28193 238 531136 n/a n/a TAACATTTCTATGGGCCTGA 6 28670 28689 239 531137 n/a n/a TGTCTACTATTTGACCAATA 19 28795 28814 240 531138 n/a n/a TTTAAATGTGTCACTTAATC 0 28987 29006 241 531139 n/a n/a TCACTAAAACAAAAATACTT 0 29156 29175 242 531140 n/a n/a TCTTCCAGGCCAACCACCTT 22 29321 29340 243 531141 n/a n/a TGCAAGGCATGTGTGCACAA 47 29532 29551 244 531142 n/a n/a TGTTTAAAATATCTCTATAC 8 30008 30027 245 531143 n/a n/a CATGGAAAAATTAAGCTCAT 0 30133 30152 246 531144 n/a n/a TGAAGATTCTATTTAACAAA 0 30266 30285 247 531145 n/a n/a GCCTAGGAGAGAAAAATAAA 0 30445 30464 248 531146 n/a n/a CCAGTGTAATTCAAAGGAGC 40 30620 30639 249 531147 n/a n/a CCATTATTTCCATCACCTGC 18 30871 30890 250 531148 n/a n/a TACCCAAATTATACCTGGAA 8 31015 31034 251 531149 n/a n/a AGAGGTAAAGCAACTTGCCC 45 31429 31448 252 531150 n/a n/a TCCTTAATAGTCATAGCAGG 48 31558 31577 253 531151 n/a n/a TCACCACCATTTTTCACATG 44 31683 31702 254 531152 n/a n/a GTTATGGATATAGACTTTAA 0 31808 31827 255 531153 n/a n/a CTAGAAGCAATATTTAAAGC 0 31974 31993 256 531154 n/a n/a ATGAAGTAAGATGCTTAAAA 16 32162 32181 257 531155 n/a n/a CTTCTTGTCTCAGATTACCA 79 32464 32483 258 531156 n/a n/a TCTGAAAAGCCCTCCGAGCT 0 32589 32608 259 531157 n/a n/a AAGTGAATCAGAGCAGTGTA 46 32961 32980 260 531158 n/a n/a ACCTTACAAGCATCTTTTCC 41 33223 33242 261 531159 n/a n/a ATTTGTTAAAAGTTGCTTAT 0 33368 33387 262 531160 n/a n/a TGATATCATCATCCCAATGA 13 33510 33529 263
TABLE-US-00116 TABLE 125 SEQ ID SEQ ID SEQ ID SEQ ID NO: 1 NO: 1 NO: 10 NO: 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence inhibition Site Site NO 531083 n/a n/a TGTACTAGTTTCCTATAACT 68 14738 14757 264 14809 14828 14880 14899 14939 14958 15071 15090 15214 15233 15286 15305 15345 15364 15477 15496 15549 15568 15607 15626 15679 15698 15809 15828 15881 15900 15939 15958 531161 n/a n/a CAGACACCTTCTTCACAAGG 40 898 917 264 531162 n/a n/a AATTTCCCAGATGTATTAGT 43 1054 1073 265 531163 n/a n/a TCAGCAGAAATCATGTAGGC 60 1181 1200 266 531164 n/a n/a TTAAATATAAAGAGATCCTC 38 1609 1628 267 531165 n/a n/a GTAATAAAAGGAATGATAAA 0 1825 1844 268 531166 n/a n/a AGACAGTAAACAAAATCAGG 12 2046 2065 269 531167 n/a n/a CAAGAAACCACCAAAGGAAG 37 2176 2195 270 531168 n/a n/a ACCCCAACAGACAGCCCACC 55 2314 2333 271 531169 n/a n/a TGGGCTCACCCCAGTGGACC 54 2580 2599 272 531170 n/a n/a GCCTGGCCCCCAAGACTCTA 54 2743 2762 273 531171 n/a n/a AGGCCTGCCACAGGCCAGAC 40 2873 2892 274 531172 n/a n/a TTCAAGCCTGGGCAGCACAG 71 3004 3023 275 531173 n/a n/a AAAATAACTTCACTAGAGCT 22 3131 3150 276 531174 n/a n/a TGTTAAGTATATTAACTATT 10 3256 3275 277 531175 n/a n/a TACTCAGGAAATTAGAATAT 25 3550 3569 278 531176 n/a n/a TTATGAAACCTCTTGATTTG 0 3753 3772 279 531177 n/a n/a TTCTTGTAAATGTCTGAATT 61 3971 3990 280 531178 n/a n/a ACCACAGGAAACTGTAGCAA 72 4111 4130 281 531179 n/a n/a GATTGGACCCAGACACTATA 57 4506 4525 282 531180 n/a n/a CCTCTTAAGTCACCATAGAC 45 4785 4804 283 531181 n/a n/a GGTTGAGGGACAGACACAGG 36 4940 4959 284 531182 n/a n/a ATAATCATGATTTATTTTGC 34 5099 5118 285 531183 n/a n/a CATAAGAATGTGCACACAAA 39 5382 5401 286 531184 n/a n/a ACTCTTATTAGCTGGTAGAA 74 5538 5557 287 531185 n/a n/a GGACCAAAACTGAGAGGCAG 63 5663 5682 288 531186 n/a n/a CCATTACTCTCAAGCTCCAC 75 5890 5909 289 531187 n/a n/a ATCTATTGGTTCAGGAGCCA 72 6015 6034 290 531188 n/a n/a GTTAAAACAACTAGAAGCCA 67 6146 6165 291 531189 n/a n/a AGGTGTTCTTGCTTATCCTC 63 6484 6503 292 531190 n/a n/a GCAGTCACTCCTCTTCCAGC 59 6659 6678 293 531191 n/a n/a AAGTGTATTGCCTAGATTTC 37 6784 6803 294 531192 n/a n/a GAGTGCCATCTTCTCTGCAC 61 6968 6987 295 531193 n/a n/a TTATTCCCAGCTCTAAAATA 23 7274 7293 296 531194 n/a n/a CTCACAATTCTGTAAGGGAA 64 7596 7615 297 531195 n/a n/a ATAAAATATATTAAGGCAAC 61 7846 7865 298 531196 n/a n/a TTGAGTCAGACATCCTGTGA 38 7996 8015 299 531197 n/a n/a TACCTTTTCTCCATGTCATT 42 8148 8167 300 531198 n/a n/a GGGATTTTGCTGAAGCTGGT 73 8273 8292 301 531199 n/a n/a CTTTGAATAGAAAATGACTA 1 8415 8434 302 531200 n/a n/a CAAAATCACAAGTTCTAGAT 51 8617 8636 303 531201 n/a n/a TTTCCAATACTTTTACAAAT 52 8760 8779 304 531202 n/a n/a ATTAATAAGCATCTCTCTGA 31 9109 9128 305 531203 n/a n/a TGACTATCCAATTTCTAGTT 67 9253 9272 306 531204 n/a n/a CTTGTAGTCTGCACTTAATG 60 9418 9437 307 531205 n/a n/a ACATTTTTTAAGTACAGGAA 0 9602 9621 308 531206 n/a n/a GAAATGTCTAGCATTTTCTA 28 9755 9774 309 531207 n/a n/a CCACTTATTTGATGACCACA 64 9915 9934 310 531208 n/a n/a TCCAGAATACTGCCCCATCT 23 10050 10069 311 531209 n/a n/a TGGATTCATTTTCTGCAAAT 81 10175 10194 312 531210 n/a n/a AGACATTGTCAAATGTCCCC 60 10322 10341 313 531211 n/a n/a TTGATGTCAGCACTGTTGAC 77 10480 10499 314 531212 n/a n/a ACATCAGTAGCTTCAGATGT 56 10618 10637 315 531213 n/a n/a CAAAATTAATTGTGCATAAT 13 10820 10839 316 531214 n/a n/a TTTTTCTTTAAATTTTGCTA 37 11120 11139 317 531215 n/a n/a TAGAGATTTTATGTACTTGG 63 11245 11264 318 531216 n/a n/a AAACACAGGAATTTGCAGAC 33 11408 11427 319 531217 n/a n/a GTGGAATAAACCATAATCTA 47 11579 11598 320 531218 n/a n/a GATAATTCTTTTCACAGACA 72 12028 12047 321 531219 n/a n/a CTTCTCTATCTCCCAGTGTT 61 12227 12246 322 531220 n/a n/a CAATACAGGTAAATTTCACG 56 12374 12393 323 531221 n/a n/a AAGGGATTTAAAATTTTTAT 0 12507 12526 324 531222 n/a n/a GGCAAGCTGTACAAGAAAAA 19 12642 12661 325 531223 n/a n/a TGTACTCACCGGTACTCTGC 58 12805 12824 326 531224 n/a n/a AAGAGAATGCTCAGAAATGG 25 13435 13454 327 531225 n/a n/a ACACTTGTACCCCATACATC 45 13560 13579 328 531226 n/a n/a GACAGTAGAGACTGGGAAGG 12 13708 13727 329 531227 n/a n/a TACCAATTTCTGAAAGGGCA 72 14224 14243 330 531228 n/a n/a CAGAGTAAACTCCCCATCTC 33 14387 14406 331 531229 n/a n/a CTTCAAAGCCAGCAGTGTAA 69 14514 14533 332 531230 n/a n/a CTTACTGGGCTAAAATCAAG 46 14639 14658 333 531231 n/a n/a TATCACTGTACTAGTTTCCT 94 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 531232 n/a n/a CTGTACTAGTTTCCTATAAC 85 14739 14758 335 14810 14829 14881 14900 14940 14959 15000 15019 15072 15091 15215 15234 15287 15306 15346 15365 15406 15425 15478 15497 15550 15569 15608 15627 15680 15699 15810 15829 15882 15901 15940 15959 531233 n/a n/a ACTGTACTAGTTTCCTATAA 86 14740 14759 336 14811 14830 14882 14901 14941 14960 15001 15020 15073 15092 15216 15235 15288 15307 15347 15366 15407 15426 15479 15498 15551 15570 15609 15628 15681 15700 15811 15830 15883 15902 15941 15960 531234 n/a n/a CACTGTACTAGTTTCCTATA 86 14741 14760 337 14812 14831 14883 14902 14942 14961 15002 15021 15074 15093 15217 15236 15289 15308 15348 15367 15408 15427 15480 15499 15552 15571 15610 15629 15682 15701 15812 15831 15884 15903 15942 15961 531235 n/a n/a TCACTGTACTAGTTTCCTAT 86 14742 14761 338 14813 14832 14884 14903 14943 14962 15003 15022 15075 15094 15218 15237 15290 15309 15349 15368 15409 15428 15481 15500 15553 15572 15611 15630 15683 15702
15813 15832 15885 15904 15943 15962 531236 n/a n/a ATCACTGTACTAGTTTCCTA 87 14743 14762 339 14814 14833 14885 14904 14944 14963 15004 15023 15076 15095 15219 15238 15291 15310 15350 15369 15410 15429 15482 15501 15554 15573 15612 15631 15684 15703 15814 15833 15886 15905 15944 15963 531237 n/a n/a GTGGAATGTCATGGCAATTT 56 16399 16418 340
Example 115: Antisense Inhibition of Human PKK in HepaRG.TM. Cells by Antisense Oligonucleotides with 2'-MOE Sugar Modifications
[1064] Additional antisense oligonucleotides were designed targeting a PKK nucleic acid and were tested for their effects on PKK mRNA in vitro.
[1065] The chimeric antisense oligonucleotides in the tables below were designed as 5-10-5 MOE gapmers, 4-9-4 MOE gapmers, 4-10-4 MOE gapmers, 4-10-3 MOE gapmers, 3-10-4 MOE gapmers, or 3-10-3 MOE gapmers. The 5-10-5 MOE gapmers are 20 nucleosides in length, wherein the central gap segment comprises often 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising five nucleosides each. The 4-9-4 MOE gapmers are 17 nucleosides in length, wherein the central gap segment comprises of nine 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising four nucleosides each. The 4-10-4 MOE gapmers are 18 nucleosides in length, wherein the central gap segment comprises often 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising four nucleosides each. The 4-10-3 MOE gapmers are 17 nucleosides in length, wherein the central gap segment comprises often 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising four and three nucleosides respectively. The 3-10-4 MOE gapmers are 17 nucleosides in length, wherein the central gap segment comprises often 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising three and four nucleosides respectively. The 3-10-3 MOE gapmers are 16 nucleosides in length, wherein the central gap segment comprises often 2'-deoxynucleosides and is flanked by wing segments on the 5' direction and the 3' direction comprising three nucleosides each. Each nucleoside in the 5' wing segment and each nucleoside in the 3' wing segment has a 2'-O-methoxyethyl modification. The internucleoside linkages throughout each gapmer are phosphorothioate linkages. All cytosine residues throughout each gapmer are 5-methylcytosines. "Start site" indicates the 5'-most nucleoside to which the gapmer is targeted in the human gene sequence. "Stop site" indicates the 3'-most nucleoside to which the gapmer is targeted in the human gene sequence. Each gapmer listed in the tables below is targeted to either SEQ ID NO: 1 or SEQ ID NO: 10. `n/a` indicates that the antisense oligonucleotide does not target that particular gene sequence.
[1066] Cultured HepaRG.TM. cells at a density of 20,000 cells per well were transfected using electroporation with 5,000 nM antisense oligonucleotide. After a treatment period of approximately 24 hours, RNA was isolated from the cells and PKK mRNA levels were measured by quantitative real-time PCR. Human primer probe set RTS3454 was used to measure mRNA levels. PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.RTM.. The antisense oligonucleotides were tested in a series of experiments that had similar culture conditions. The results for each experiment are presented in separate tables shown below. Results are presented as percent inhibition of PKK, relative to untreated control cells.
TABLE-US-00117 TABLE 126 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 98 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546131 4 23 ATGAACGGTCTTCAAGCTGT 5-10-5 75 3396 3415 341 547269 5 24 AATGAACGGTCTTCAAGCTG 5-10-5 56 3397 3416 342 547270 7 26 AAAATGAACGGTCTTCAAGC 5-10-5 68 3399 3418 343 547271 10 29 TTAAAAATGAACGGTCTTCA 5-10-5 60 3402 3421 344 547272 13 32 CACTTAAAAATGAACGGTCT 5-10-5 82 3405 3424 345 547273 25 44 TGAGTCTCTTGTCACTTAAA 5-10-5 93 3417 3436 346 547274 29 48 GAGGTGAGTCTCTTGTCACT 5-10-5 70 3421 3440 347 546136 30 49 GGAGGTGAGTCTCTTGTCAC 5-10-5 86 3422 3441 348 547275 32 51 TTGGAGGTGAGTCTCTTGTC 5-10-5 87 3424 3443 349 546137 40 59 ATTGCTTCTTGGAGGTGAGT 5-10-5 76 3432 3451 350 547276 42 61 CAATTGCTTCTTGGAGGTGA 5-10-5 93 3434 3453 351 547277 44 63 CACAATTGCTTCTTGGAGGT 5-10-5 75 3436 3455 352 547278 45 64 ACACAATTGCTTCTTGGAGG 5-10-5 70 3437 3456 353 546138 47 66 AAACACAATTGCTTCTTGGA 5-10-5 69 3439 3458 354 547279 48 67 AAAACACAATTGCTTCTTGG 5-10-5 69 3440 3459 355 547280 49 68 GAAAACACAATTGCTTCTTG 5-10-5 47 3441 3460 356 547281 70 89 TTGCTTGAATAAAATCATTC 5-10-5 41 4069 4088 357 546140 72 91 GCTTGCTTGAATAAAATCAT 5-10-5 60 4071 4090 358 547282 74 93 TTGCTTGCTTGAATAAAATC 5-10-5 53 4073 4092 359 547283 76 95 AGTTGCTTGCTTGAATAAAA 5-10-5 67 4075 4094 360 546141 82 101 GAAATAAGTTGCTTGCTTGA 5-10-5 56 4081 4100 361 547284 86 105 AAATGAAATAAGTTGCTTGC 5-10-5 26 4085 4104 362 547285 102 121 ACTGTAGCAAACAAGGAAAT 5-10-5 51 4101 4120 363 546143 106 125 GGAAACTGTAGCAAACAAGG 5-10-5 46 4105 4124 364 546144 110 129 CACAGGAAACTGTAGCAAAC 5-10-5 75 4109 4128 365 547286 117 136 AGACATCCACAGGAAACTGT 5-10-5 68 n/a n/a 366 547287 120 139 GTCAGACATCCACAGGAAAC 5-10-5 69 n/a n/a 367 546146 123 142 TGAGTCAGACATCCACAGGA 5-10-5 72 n/a n/a 368 547288 131 150 CATAGAGTTGAGTCAGACAT 5-10-5 80 8003 8022 369 546147 132 151 TCATAGAGTTGAGTCAGACA 5-10-5 76 8004 8023 370 547289 133 152 TTCATAGAGTTGAGTCAGAC 5-10-5 74 8005 8024 371 546148 137 156 CGTTTTCATAGAGTTGAGTC 5-10-5 68 8009 8028 372 546149 155 174 CCCCACCTCTGAAGAAGGCG 5-10-5 83 8027 8046 373 546150 158 177 CATCCCCACCTCTGAAGAAG 5-10-5 58 8030 8049 374 547290 163 182 AGCTACATCCCCACCTCTGA 5-10-5 76 8035 8054 375 546151 166 185 GGAAGCTACATCCCCACCTC 5-10-5 76 8038 8057 376 547291 168 187 ATGGAAGCTACATCCCCACC 5-10-5 74 8040 8059 377 547292 171 190 TACATGGAAGCTACATCCCC 5-10-5 60 8043 8062 378 546152 172 191 GTACATGGAAGCTACATCCC 5-10-5 73 8044 8063 379 546153 176 195 GGGTGTACATGGAAGCTACA 5-10-5 76 8048 8067 380 546154 195 214 TGGCAGTATTGGGCATTTGG 5-10-5 85 8067 8086 381 547293 199 218 CATCTGGCAGTATTGGGCAT 5-10-5 92 8071 8090 382 547294 201 220 CTCATCTGGCAGTATTGGGC 5-10-5 85 8073 8092 383 546155 202 221 CCTCATCTGGCAGTATTGGG 5-10-5 47 8074 8093 384 547295 203 222 ACCTCATCTGGCAGTATTGG 5-10-5 88 8075 8094 385 547296 206 225 TGCACCTCATCTGGCAGTAT 5-10-5 72 8078 8097 386 546156 211 230 GAATGTGCACCTCATCTGGC 5-10-5 81 8083 8102 387 547297 213 232 TGGAATGTGCACCTCATCTG 5-10-5 84 8085 8104 388 546157 216 235 GGGTGGAATGTGCACCTCAT 5-10-5 85 8088 8107 389 547298 218 237 TTGGGTGGAATGTGCACCTC 5-10-5 90 8090 8109 390 546158 219 238 CTTGGGTGGAATGTGCACCT 5-10-5 95 8091 8110 391 546159 229 248 TAGCAAACACCTTGGGTGGA 5-10-5 76 8101 8120 392 546160 235 254 ACTGAATAGCAAACACCTTG 5-10-5 78 8107 8126 393 547299 237 256 AAACTGAATAGCAAACACCT 5-10-5 76 8109 8128 394 546163 250 269 ACTTGCTGGAAGAAAACTGA 5-10-5 42 8122 8141 395 547300 252 271 GAACTTGCTGGAAGAAAACT 5-10-5 37 8124 8143 396 546164 257 276 TGATTGAACTTGCTGGAAGA 5-10-5 33 8129 8148 397 546165 260 279 CATTGATTGAACTTGCTGGA 5-10-5 71 8132 8151 398 547301 261 280 TCATTGATTGAACTTGCTGG 5-10-5 80 8133 8152 399 546166 263 282 TGTCATTGATTGAACTTGCT 5-10-5 70 8135 8154 400 547302 266 285 CCATGTCATTGATTGAACTT 5-10-5 58 8138 8157 401 546167 268 287 CTCCATGTCATTGATTGAAC 5-10-5 73 8140 8159 402 547303 270 289 TTCTCCATGTCATTGATTGA 5-10-5 72 8142 8161 403 547304 273 292 CTTTTCTCCATGTCATTGAT 5-10-5 71 8145 8164 404 547305 280 299 ACCAAACCTTTTCTCCATGT 5-10-5 47 n/a n/a 405 546170 283 302 GCAACCAAACCTTTTCTCCA 5-10-5 54 n/a n/a 406 547306 284 303 AGCAACCAAACCTTTTCTCC 5-10-5 62 n/a n/a 407 547307 286 305 GAAGCAACCAAACCTTTTCT 5-10-5 58 n/a n/a 408 547308 290 309 TCAAGAAGCAACCAAACCTT 5-10-5 66 n/a n/a 409 547309 293 312 CTTTCAAGAAGCAACCAAAC 5-10-5 71 9827 9846 410 547310 295 314 ATCTTTCAAGAAGCAACCAA 5-10-5 81 9829 9848 411 546171 297 316 CTATCTTTCAAGAAGCAACC 5-10-5 81 9831 9850 412 547311 299 318 CACTATCTTTCAAGAAGCAA 5-10-5 71 9833 9852 413 546172 301 320 AACACTATCTTTCAAGAAGC 5-10-5 81 9835 9854 414 547312 325 344 ATGTACTTTTGGCAGGGTTC 5-10-5 46 9859 9878 415 546173 327 346 CGATGTACTTTTGGCAGGGT 5-10-5 84 9861 9880 416 547313 330 349 GTTCGATGTACTTTTGGCAG 5-10-5 73 9864 9883 417
TABLE-US-00118 TABLE 127 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 86 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546174 333 352 CCTGTTCGATGTACTTTTGG 5-10-5 74 9867 9886 418 547314 336 355 GCACCTGTTCGATGTACTTT 5-10-5 73 9870 9889 419 546175 338 357 CTGCACCTGTTCGATGTACT 5-10-5 78 9872 9891 420 547315 340 359 AACTGCACCTGTTCGATGTA 5-10-5 50 9874 9893 421 547316 342 361 GAAACTGCACCTGTTCGATG 5-10-5 75 9876 9895 422 547317 344 363 CAGAAACTGCACCTGTTCGA 5-10-5 75 9878 9897 423 547318 345 364 CCAGAAACTGCACCTGTTCG 5-10-5 74 9879 9898 424 546177 348 367 TGTCCAGAAACTGCACCTGT 5-10-5 75 9882 9901 425 547319 351 370 GAATGTCCAGAAACTGCACC 5-10-5 62 9885 9904 426 547320 353 372 AGGAATGTCCAGAAACTGCA 5-10-5 73 9887 9906 427 547321 356 375 TCAAGGAATGTCCAGAAACT 5-10-5 53 9890 9909 428 547322 358 377 CTTCAAGGAATGTCCAGAAA 5-10-5 65 9892 9911 429 547323 361 380 TTGCTTCAAGGAATGTCCAG 5-10-5 56 9895 9914 430 547324 363 382 CATTGCTTCAAGGAATGTCC 5-10-5 76 9897 9916 431 547325 368 387 GACCACATTGCTTCAAGGAA 5-10-5 67 9902 9921 432 546181 369 388 TGACCACATTGCTTCAAGGA 5-10-5 75 9903 9922 433 547326 370 389 ATGACCACATTGCTTCAAGG 5-10-5 48 9904 9923 434 547327 373 392 TTGATGACCACATTGCTTCA 5-10-5 45 9907 9926 435 547328 375 394 ATTTGATGACCACATTGCTT 5-10-5 40 9909 9928 436 547329 377 396 TTATTTGATGACCACATTGC 5-10-5 24 9911 9930 437 547330 378 397 CTTATTTGATGACCACATTG 5-10-5 60 9912 9931 438 546183 380 399 CACTTATTTGATGACCACAT 5-10-5 69 9914 9933 439 547331 382 401 AGCACTTATTTGATGACCAC 5-10-5 47 n/a n/a 440 546184 384 403 CAAGCACTTATTTGATGACC 5-10-5 65 n/a n/a 441 547332 390 409 CGATGGCAAGCACTTATTTG 5-10-5 44 n/a n/a 442 547333 395 414 TGTCTCGATGGCAAGCACTT 5-10-5 76 n/a n/a 443 546186 396 415 ATGTCTCGATGGCAAGCACT 5-10-5 84 n/a n/a 444 547334 397 416 AATGTCTCGATGGCAAGCAC 5-10-5 74 n/a n/a 445 547335 402 421 TTATAAATGTCTCGATGGCA 5-10-5 93 12658 12677 446 547336 403 422 TTTATAAATGTCTCGATGGC 5-10-5 81 12659 12678 447 546188 407 426 CTCCTTTATAAATGTCTCGA 5-10-5 95 12663 12682 448 547337 409 428 AACTCCTTTATAAATGTCTC 5-10-5 84 12665 12684 449 547338 411 430 TCAACTCCTTTATAAATGTC 5-10-5 71 12667 12686 450 547339 413 432 TATCAACTCCTTTATAAATG 5-10-5 42 12669 12688 451 546190 419 438 CTCTCATATCAACTCCTTTA 5-10-5 92 12675 12694 452 547340 422 441 CTCCTCTCATATCAACTCCT 5-10-5 93 12678 12697 453 547341 424 443 GACTCCTCTCATATCAACTC 5-10-5 87 12680 12699 454 546192 428 447 AATTGACTCCTCTCATATCA 5-10-5 51 12684 12703 455 547342 433 452 ATTAAAATTGACTCCTCTCA 5-10-5 66 12689 12708 456 546193 434 453 CATTAAAATTGACTCCTCTC 5-10-5 57 12690 12709 457 547343 436 455 CACATTAAAATTGACTCCTC 5-10-5 78 12692 12711 458 547344 438 457 GACACATTAAAATTGACTCC 5-10-5 80 12694 12713 459 547345 439 458 AGACACATTAAAATTGACTC 5-10-5 80 12695 12714 460 547346 444 463 ACCTTAGACACATTAAAATT 5-10-5 57 12700 12719 461 546195 448 467 GCTAACCTTAGACACATTAA 5-10-5 83 12704 12723 462 547347 451 470 ACTGCTAACCTTAGACACAT 5-10-5 82 12707 12726 463 546196 452 471 CACTGCTAACCTTAGACACA 5-10-5 83 12708 12727 464 547348 453 472 ACACTGCTAACCTTAGACAC 5-10-5 83 12709 12728 465 547349 458 477 CTTCAACACTGCTAACCTTA 5-10-5 88 12714 12733 466 546198 459 478 TCTTCAACACTGCTAACCTT 5-10-5 85 12715 12734 467 547350 464 483 GGCATTCTTCAACACTGCTA 5-10-5 96 12720 12739 468 546199 465 484 TGGCATTCTTCAACACTGCT 5-10-5 97 12721 12740 469 547351 467 486 TTTGGCATTCTTCAACACTG 5-10-5 92 12723 12742 470 546200 500 519 AAAACTGGCAGCGAATGTTA 5-10-5 91 12756 12775 471 547352 541 560 CCGGTACTCTGCCTTGTGAA 5-10-5 94 12797 12816 472 547354 547 566 ATTGTTCCGGTACTCTGCCT 5-10-5 89 n/a n/a 473 546203 548 567 AATTGTTCCGGTACTCTGCC 5-10-5 76 n/a n/a 474 547355 549 568 CAATTGTTCCGGTACTCTGC 5-10-5 77 n/a n/a 475 546204 555 574 AATAGGCAATTGTTCCGGTA 5-10-5 91 n/a n/a 476 547356 556 575 TAATAGGCAATTGTTCCGGT 5-10-5 83 n/a n/a 477 547357 559 578 CTTTAATAGGCAATTGTTCC 5-10-5 78 14130 14149 478 546205 562 581 GTACTTTAATAGGCAATTGT 5-10-5 83 14133 14152 479 547359 569 588 CGGGACTGTACTTTAATAGG 5-10-5 81 14140 14159 480 546208 605 624 CGTTACTCAGCACCTTTATA 5-10-5 92 14176 14195 481 546209 629 648 GCTTCAGTGAGAATCCAGAT 5-10-5 73 14200 14219 482 546210 651 670 CCAATTTCTGAAAGGGCACA 5-10-5 79 14222 14241 483 547360 653 672 AACCAATTTCTGAAAGGGCA 5-10-5 88 n/a n/a 484 547361 655 674 GCAACCAATTTCTGAAAGGG 5-10-5 46 n/a n/a 485 546211 656 675 GGCAACCAATTTCTGAAAGG 5-10-5 42 n/a n/a 486 546212 678 697 AGATGCTGGAAGATGTTCAT 5-10-5 48 26126 26145 487 547362 701 720 CAACATCCACATCTGAGAAC 5-10-5 47 26149 26168 488 547363 703 722 GGCAACATCCACATCTGAGA 5-10-5 84 26151 26170 489 546213 707 726 CCCTGGCAACATCCACATCT 5-10-5 82 26155 26174 490
TABLE-US-00119 TABLE 128 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 88 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547364 710 729 GAACCCTGGCAACATCCACA 5-10-5 92 26158 26177 491 546214 712 731 GAGAACCCTGGCAACATCCA 5-10-5 88 26160 26179 492 547365 713 732 TGAGAACCCTGGCAACATCC 5-10-5 81 26161 26180 493 547366 717 736 GGAGTGAGAACCCTGGCAAC 5-10-5 86 26165 26184 494 546216 719 738 CTGGAGTGAGAACCCTGGCA 5-10-5 93 26167 26186 495 547367 721 740 ATCTGGAGTGAGAACCCTGG 5-10-5 76 26169 26188 496 547368 723 742 GCATCTGGAGTGAGAACCCT 5-10-5 89 26171 26190 497 547369 725 744 AAGCATCTGGAGTGAGAACC 5-10-5 76 26173 26192 498 547370 728 747 CAAAAGCATCTGGAGTGAGA 5-10-5 73 26176 26195 499 546217 730 749 CACAAAAGCATCTGGAGTGA 5-10-5 83 26178 26197 500 546218 740 759 TGGTCCGACACACAAAAGCA 5-10-5 71 26188 26207 501 547371 741 760 ATGGTCCGACACACAAAAGC 5-10-5 66 26189 26208 502 547372 742 761 GATGGTCCGACACACAAAAG 5-10-5 32 26190 26209 503 547373 745 764 GCAGATGGTCCGACACACAA 5-10-5 90 26193 26212 504 546220 750 769 TAGGTGCAGATGGTCCGACA 5-10-5 71 26198 26217 505 547374 752 771 GATAGGTGCAGATGGTCCGA 5-10-5 81 26200 26219 506 547375 754 773 GTGATAGGTGCAGATGGTCC 5-10-5 72 26202 26221 507 546222 756 775 GGGTGATAGGTGCAGATGGT 5-10-5 12 26204 26223 508 547376 778 797 GAATGTAAAGAAGAGGCAGT 5-10-5 43 26226 26245 509 546224 780 799 TAGAATGTAAAGAAGAGGCA 5-10-5 65 26228 26247 510 547377 788 807 CATTTGTATAGAATGTAAAG 5-10-5 6 26236 26255 511 547378 790 809 TACATTTGTATAGAATGTAA 5-10-5 0 26238 26257 512 546226 793 812 CCATACATTTGTATAGAATG 5-10-5 37 26241 26260 513 547379 802 821 CTCGATTTTCCATACATTTG 5-10-5 37 26250 26269 514 547380 805 824 TGACTCGATTTTCCATACAT 5-10-5 42 26253 26272 515 546228 806 825 GTGACTCGATTTTCCATACA 5-10-5 60 26254 26273 516 547381 807 826 TGTGACTCGATTTTCCATAC 5-10-5 49 26255 26274 517 547382 810 829 CTTTGTGACTCGATTTTCCA 5-10-5 62 26258 26277 518 547383 812 831 TTCTTTGTGACTCGATTTTC 5-10-5 37 n/a n/a 519 546229 816 835 ACATTTCTTTGTGACTCGAT 5-10-5 19 n/a n/a 520 547384 818 837 AAACATTTCTTTGTGACTCG 5-10-5 50 n/a n/a 521 547385 847 866 TGTGCCACTTTCAGATGTTT 5-10-5 80 27111 27130 522 546230 848 867 GTGTGCCACTTTCAGATGTT 5-10-5 70 27112 27131 523 546231 852 871 CTTGGTGTGCCACTTTCAGA 5-10-5 79 27116 27135 524 547386 853 872 ACTTGGTGTGCCACTTTCAG 5-10-5 78 27117 27136 525 546232 857 876 AGGAACTTGGTGTGCCACTT 5-10-5 86 27121 27140 526 547387 878 897 TGGTGTTTTCTTGAGGAGTA 5-10-5 73 27142 27161 527 546233 879 898 ATGGTGTTTTCTTGAGGAGT 5-10-5 69 27143 27162 528 547388 880 899 TATGGTGTTTTCTTGAGGAG 5-10-5 55 27144 27163 529 547389 884 903 CAGATATGGTGTTTTCTTGA 5-10-5 61 27148 27167 530 546234 885 904 CCAGATATGGTGTTTTCTTG 5-10-5 69 27149 27168 531 547390 887 906 ATCCAGATATGGTGTTTTCT 5-10-5 63 27151 27170 532 547391 889 908 ATATCCAGATATGGTGTTTT 5-10-5 32 27153 27172 533 546235 893 912 GGCTATATCCAGATATGGTG 5-10-5 77 27157 27176 534 547392 895 914 AAGGCTATATCCAGATATGG 5-10-5 81 27159 27178 535 546236 900 919 GTTAAAAGGCTATATCCAGA 5-10-5 50 27164 27183 536 546237 903 922 CAGGTTAAAAGGCTATATCC 5-10-5 64 27167 27186 537 547393 905 924 TGCAGGTTAAAAGGCTATAT 5-10-5 73 27169 27188 538 547394 907 926 TTTGCAGGTTAAAAGGCTAT 5-10-5 29 27171 27190 539 546238 909 928 CTTTTGCAGGTTAAAAGGCT 5-10-5 63 27173 27192 540 546239 912 931 GTTCTTTTGCAGGTTAAAAG 5-10-5 47 27176 27195 541 547395 914 933 AAGTTCTTTTGCAGGTTAAA 5-10-5 15 27178 27197 542 546240 917 936 GTAAAGTTCTTTTGCAGGTT 5-10-5 23 27181 27200 543 546241 920 939 CAGGTAAAGTTCTTTTGCAG 5-10-5 69 27184 27203 544 547396 921 940 TCAGGTAAAGTTCTTTTGCA 5-10-5 49 n/a n/a 545 547397 923 942 GTTCAGGTAAAGTTCTTTTG 5-10-5 27 n/a n/a 546 546242 925 944 GGGTTCAGGTAAAGTTCTTT 5-10-5 8 n/a n/a 547 547398 927 946 CAGGGTTCAGGTAAAGTTCT 5-10-5 16 n/a n/a 548 547399 928 947 GCAGGGTTCAGGTAAAGTTC 5-10-5 10 n/a n/a 549 547400 930 949 TGGCAGGGTTCAGGTAAAGT 5-10-5 0 n/a n/a 550 547401 933 952 GAATGGCAGGGTTCAGGTAA 5-10-5 22 n/a n/a 551 546243 934 953 AGAATGGCAGGGTTCAGGTA 5-10-5 16 n/a n/a 552 547402 937 956 TTTAGAATGGCAGGGTTCAG 5-10-5 59 n/a n/a 553 547403 939 958 ATTTTAGAATGGCAGGGTTC 5-10-5 10 27361 27380 554 546244 942 961 TAAATTTTAGAATGGCAGGG 5-10-5 27 27364 27383 555 547404 956 975 AGTCAACTCCCGGGTAAATT 5-10-5 64 27378 27397 556 547405 959 978 CAAAGTCAACTCCCGGGTAA 5-10-5 47 27381 27400 557 546247 960 979 CCAAAGTCAACTCCCGGGTA 5-10-5 90 27382 27401 558 546248 963 982 CCTCCAAAGTCAACTCCCGG 5-10-5 86 27385 27404 559 547406 965 984 CTCCTCCAAAGTCAACTCCC 5-10-5 81 27387 27406 560 546249 968 987 CTTCTCCTCCAAAGTCAACT 5-10-5 68 27390 27409 561 547407 975 994 TTCAATTCTTCTCCTCCAAA 5-10-5 59 27397 27416 562 546250 977 996 CATTCAATTCTTCTCCTCCA 5-10-5 65 27399 27418 563 547408 980 999 TCACATTCAATTCTTCTCCT 5-10-5 84 27402 27421 564 547409 982 1001 AGTCACATTCAATTCTTCTC 5-10-5 67 27404 27423 565 546251 1007 1026 GGCAAACATTCACTCCTTTA 5-10-5 92 27429 27448 566
TABLE-US-00120 TABLE 129 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 95 14744 14763 344 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546252 1011 1030 TCTTGGCAAACATTCACTCC 5-10-5 73 27433 27452 567 546253 1014 1033 GTCTCTTGGCAAACATTCAC 5-10-5 98 27436 27455 568 547410 1017 1036 CAAGTCTCTTGGCAAACATT 5-10-5 88 27439 27458 569 546254 1019 1038 TGCAAGTCTCTTGGCAAACA 5-10-5 95 27441 27460 570 546255 1024 1043 CTTTGTGCAAGTCTCTTGGC 5-10-5 92 27446 27465 571 547411 1027 1046 CATCTTTGTGCAAGTCTCTT 5-10-5 79 27449 27468 572 546256 1028 1047 TCATCTTTGTGCAAGTCTCT 5-10-5 83 27450 27469 573 547412 1029 1048 ATCATCTTTGTGCAAGTCTC 5-10-5 73 27451 27470 574 546258 1036 1055 ACAGCGAATCATCTTTGTGC 5-10-5 74 27458 27477 575 546259 1040 1059 ACTGACAGCGAATCATCTTT 5-10-5 86 27462 27481 576 546260 1045 1064 GAAAAACTGACAGCGAATCA 5-10-5 84 27467 27486 577 547413 1047 1066 GTGAAAAACTGACAGCGAAT 5-10-5 94 27469 27488 578 546263 1061 1080 GGAGTAAAGAATAAGTGAAA 5-10-5 0 27483 27502 579 547414 1063 1082 TGGGAGTAAAGAATAAGTGA 5-10-5 76 27485 27504 580 547415 1065 1084 TCTGGGAGTAAAGAATAAGT 5-10-5 71 27487 27506 581 546265 1069 1088 GTCTTCTGGGAGTAAAGAAT 5-10-5 65 27491 27510 582 546266 1072 1091 ACAGTCTTCTGGGAGTAAAG 5-10-5 63 27494 27513 583 547416 1075 1094 CTTACAGTCTTCTGGGAGTA 5-10-5 79 27497 27516 584 546267 1076 1095 CCTTACAGTCTTCTGGGAGT 5-10-5 72 27498 27517 585 547417 1077 1096 TCCTTACAGTCTTCTGGGAG 5-10-5 68 27499 27518 586 546268 1079 1098 CTTCCTTACAGTCTTCTGGG 5-10-5 93 27501 27520 587 547418 1092 1111 CACTTACACTTCTCTTCCTT 5-10-5 0 n/a n/a 588 546270 1093 1112 ACACTTACACTTCTCTTCCT 5-10-5 32 n/a n/a 589 546271 1097 1116 AGAAACACTTACACTTCTCT 5-10-5 60 n/a n/a 590 547419 1101 1120 CTTAAGAAACACTTACACTT 5-10-5 51 n/a n/a 591 547420 1112 1131 CCATAGATAATCTTAAGAAA 5-10-5 8 27633 27652 592 547421 1115 1134 CATCCATAGATAATCTTAAG 5-10-5 69 27636 27655 593 547422 1117 1136 ACCATCCATAGATAATCTTA 5-10-5 70 27638 27657 594 546275 1119 1138 GAACCATCCATAGATAATCT 5-10-5 87 27640 27659 595 546276 1123 1142 TGGAGAACCATCCATAGATA 5-10-5 74 27644 27663 596 546277 1146 1165 TGTGTCCCATACGCAATCCT 5-10-5 90 27667 27686 597 547423 1150 1169 CCCTTGTGTCCCATACGCAA 5-10-5 95 27671 27690 598 546279 1153 1172 GCTCCCTTGTGTCCCATACG 5-10-5 82 27674 27693 599 547424 1156 1175 AGAGCTCCCTTGTGTCCCAT 5-10-5 90 27677 27696 600 546280 1158 1177 CCAGAGCTCCCTTGTGTCCC 5-10-5 86 27679 27698 601 547425 1161 1180 TAACCAGAGCTCCCTTGTGT 5-10-5 85 27682 27701 602 546281 1162 1181 GTAACCAGAGCTCCCTTGTG 5-10-5 85 27683 27702 603 547426 1164 1183 GAGTAACCAGAGCTCCCTTG 5-10-5 92 27685 27704 604 547427 1166 1185 AAGAGTAACCAGAGCTCCCT 5-10-5 79 27687 27706 605 547428 1169 1188 TCAAAGAGTAACCAGAGCTC 5-10-5 78 27690 27709 606 546283 1171 1190 TCTCAAAGAGTAACCAGAGC 5-10-5 88 27692 27711 607 547429 1173 1192 AATCTCAAAGAGTAACCAGA 5-10-5 81 27694 27713 608 547430 1174 1193 CAATCTCAAAGAGTAACCAG 5-10-5 70 27695 27714 609 546284 1176 1195 CACAATCTCAAAGAGTAACC 5-10-5 89 27697 27716 610 546285 1180 1199 GTTACACAATCTCAAAGAGT 5-10-5 76 27701 27720 611 547431 1184 1203 CAGTGTTACACAATCTCAAA 5-10-5 67 27705 27724 612 547432 1186 1205 CCCAGTGTTACACAATCTCA 5-10-5 90 27707 27726 613 547433 1189 1208 GTCCCCAGTGTTACACAATC 5-10-5 63 27710 27729 614 546287 1192 1211 GTTGTCCCCAGTGTTACACA 5-10-5 82 27713 27732 615 546288 1240 1259 GTTTGTTCCTCCAACAATGC 5-10-5 78 27916 27935 616 547434 1243 1262 AGAGTTTGTTCCTCCAACAA 5-10-5 54 27919 27938 617 547435 1248 1267 CAAGAAGAGTTTGTTCCTCC 5-10-5 85 27924 27943 618 546290 1251 1270 CCCCAAGAAGAGTTTGTTCC 5-10-5 86 27927 27946 619 547436 1253 1272 CTCCCCAAGAAGAGTTTGTT 5-10-5 0 27929 27948 620 547437 1255 1274 CTCTCCCCAAGAAGAGTTTG 5-10-5 50 27931 27950 621 547438 1261 1280 GGGCCACTCTCCCCAAGAAG 5-10-5 82 27937 27956 622 546291 1263 1282 CAGGGCCACTCTCCCCAAGA 5-10-5 81 27939 27958 623 547439 1298 1317 TCTGAGCTGTCAGCTTCACC 5-10-5 85 27974 27993 624 546293 1301 1320 GCCTCTGAGCTGTCAGCTTC 5-10-5 64 27977 27996 625 547440 1327 1346 TCCTATGAGTGACCCTCCAC 5-10-5 67 28003 28022 626 546294 1328 1347 GTCCTATGAGTGACCCTCCA 5-10-5 72 28004 28023 627 547441 1331 1350 GGTGTCCTATGAGTGACCCT 5-10-5 62 28007 28026 628 547442 1332 1351 TGGTGTCCTATGAGTGACCC 5-10-5 42 28008 28027 629 547443 1336 1355 CCACTGGTGTCCTATGAGTG 5-10-5 70 28012 28031 630 546295 1337 1356 CCCACTGGTGTCCTATGAGT 5-10-5 67 28013 28032 631 546296 1370 1389 GAAGCCCATCAAAGCAGTGG 5-10-5 27 n/a n/a 632 546297 1397 1416 TATAGATGCGCCAAACATCC 5-10-5 82 30475 30494 633 547444 1398 1417 CTATAGATGCGCCAAACATC 5-10-5 71 30476 30495 634 547445 1402 1421 GCCACTATAGATGCGCCAAA 5-10-5 97 30480 30499 635 546299 1404 1423 ATGCCACTATAGATGCGCCA 5-10-5 84 30482 30501 636 546300 1424 1443 TAATGTCTGACAGATTTAAA 5-10-5 58 30502 30521 637 546301 1427 1446 TTGTAATGTCTGACAGATTT 5-10-5 93 30505 30524 638 546302 1444 1463 TGAGAAAGGTGTATCTTTTG 5-10-5 87 30522 30541 639 547446 1447 1466 TTGTGAGAAAGGTGTATCTT 5-10-5 84 30525 30544 640 546303 1448 1467 TTTGTGAGAAAGGTGTATCT 5-10-5 77 30526 30545 641 547447 1449 1468 ATTTGTGAGAAAGGTGTATC 5-10-5 80 30527 30546 642
TABLE-US-00121 TABLE 130 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 96 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547448 1451 1470 TTATTTGTGAGAAAGGTGTA 5-10-5 75 30529 30548 643 547449 1453 1472 TTTTATTTGTGAGAAAGGTG 5-10-5 71 30531 30550 644 546304 1454 1473 CTTTTATTTGTGAGAAAGGT 5-10-5 94 30532 30551 645 547450 1456 1475 CTCTTTTATTTGTGAGAAAG 5-10-5 71 30534 30553 646 547451 1471 1490 TTGGTGAATAATAATCTCTT 5-10-5 75 30549 30568 647 546306 1472 1491 TTTGGTGAATAATAATCTCT 5-10-5 65 30550 30569 648 547452 1474 1493 GTTTTGGTGAATAATAATCT 5-10-5 47 30552 30571 649 546307 1478 1497 TATAGTTTTGGTGAATAATA 5-10-5 12 30556 30575 650 546308 1482 1501 ACTTTATAGTTTTGGTGAAT 5-10-5 57 30560 30579 651 546309 1492 1511 CCCTTCTGAGACTTTATAGT 5-10-5 88 30570 30589 652 546310 1496 1515 GATTCCCTTCTGAGACTTTA 5-10-5 78 30574 30593 653 546311 1499 1518 CATGATTCCCTTCTGAGACT 5-10-5 79 30577 30596 654 547453 1500 1519 TCATGATTCCCTTCTGAGAC 5-10-5 81 30578 30597 655 547454 1502 1521 TATCATGATTCCCTTCTGAG 5-10-5 92 30580 30599 656 547455 1503 1522 ATATCATGATTCCCTTCTGA 5-10-5 88 30581 30600 657 547456 1506 1525 GCGATATCATGATTCCCTTC 5-10-5 89 30584 30603 658 546313 1507 1526 GGCGATATCATGATTCCCTT 5-10-5 60 30585 30604 659 547457 1509 1528 AAGGCGATATCATGATTCCC 5-10-5 89 30587 30606 660 547458 1513 1532 TATCAAGGCGATATCATGAT 5-10-5 84 30591 30610 661 547459 1519 1538 GAGTTTTATCAAGGCGATAT 5-10-5 28 30597 30616 662 547460 1522 1541 CTGGAGTTTTATCAAGGCGA 5-10-5 72 30600 30619 663 546316 1524 1543 GCCTGGAGTTTTATCAAGGC 5-10-5 51 30602 30621 664 546317 1528 1547 AGGAGCCTGGAGTTTTATCA 5-10-5 12 30606 30625 665 546318 1534 1553 ATTCAAAGGAGCCTGGAGTT 5-10-5 47 30612 30631 666 547461 1537 1556 GTAATTCAAAGGAGCCTGGA 5-10-5 49 30615 30634 667 547462 1539 1558 GTGTAATTCAAAGGAGCCTG 5-10-5 59 30617 30636 668 546319 1541 1560 CAGTGTAATTCAAAGGAGCC 5-10-5 50 30619 30638 669 547463 1564 1583 TAGGCATATTGGTTTTTGGA 5-10-5 74 31870 31889 670 546320 1566 1585 GGTAGGCATATTGGTTTTTG 5-10-5 72 31872 31891 671 546321 1569 1588 GAAGGTAGGCATATTGGTTT 5-10-5 53 31875 31894 672 546322 1584 1603 CTTGTGTCACCTTTGGAAGG 5-10-5 74 31890 31909 673 547464 1585 1604 GCTTGTGTCACCTTTGGAAG 5-10-5 95 31891 31910 674 546323 1587 1606 GTGCTTGTGTCACCTTTGGA 5-10-5 94 31893 31912 675 547465 1592 1611 AAATTGTGCTTGTGTCACCT 5-10-5 88 31898 31917 676 547466 1596 1615 GTATAAATTGTGCTTGTGTC 5-10-5 82 31902 31921 677 546324 1597 1616 GGTATAAATTGTGCTTGTGT 5-10-5 73 31903 31922 678 547467 1598 1617 TGGTATAAATTGTGCTTGTG 5-10-5 80 31904 31923 679 547468 1600 1619 GTTGGTATAAATTGTGCTTG 5-10-5 61 31906 31925 680 546325 1602 1621 CAGTTGGTATAAATTGTGCT 5-10-5 74 31908 31927 681 546326 1607 1626 CCCAACAGTTGGTATAAATT 5-10-5 62 31913 31932 682 547469 1610 1629 TTACCCAACAGTTGGTATAA 5-10-5 67 31916 31935 683 546327 1612 1631 GGTTACCCAACAGTTGGTAT 5-10-5 95 31918 31937 684 546328 1624 1643 GAAGCCCCATCCGGTTACCC 5-10-5 84 31930 31949 685 547470 1628 1647 TCGAGAAGCCCCATCCGGTT 5-10-5 70 31934 31953 686 546329 1631 1650 CCTTCGAGAAGCCCCATCCG 5-10-5 18 31937 31956 687 546330 1636 1655 TTTCTCCTTCGAGAAGCCCC 5-10-5 55 31942 31961 688 547471 1638 1657 CCTTTCTCCTTCGAGAAGCC 5-10-5 58 31944 31963 689 547472 1641 1660 TCACCTTTCTCCTTCGAGAA 5-10-5 44 n/a n/a 690 546331 1642 1661 TTCACCTTTCTCCTTCGAGA 5-10-5 59 n/a n/a 691 547473 1649 1668 TTTGGATTTCACCTTTCTCC 5-10-5 5 n/a n/a 692 547474 1659 1678 TGTAGAATATTTTGGATTTC 5-10-5 51 33103 33122 693 547475 1686 1705 TTTGTTACCAAAGGAATATT 5-10-5 44 33130 33149 694 547476 1688 1707 CATTTGTTACCAAAGGAATA 5-10-5 75 33132 33151 695 546336 1689 1708 TCATTTGTTACCAAAGGAAT 5-10-5 66 33133 33152 696 547477 1692 1711 TCTTCATTTGTTACCAAAGG 5-10-5 74 33136 33155 697 547478 1695 1714 CATTCTTCATTTGTTACCAA 5-10-5 85 33139 33158 698 546339 1712 1731 CTTGATATCTTTTCTGGCAT 5-10-5 65 33156 33175 699 546340 1716 1735 TAATCTTGATATCTTTTCTG 5-10-5 30 33160 33179 700 547479 1718 1737 TATAATCTTGATATCTTTTC 5-10-5 48 33162 33181 701 547480 1756 1775 TTCTTTATAGCCAGCACAGA 5-10-5 60 33200 33219 702 547481 1758 1777 CCTTCTTTATAGCCAGCACA 5-10-5 71 33202 33221 703 547482 1760 1779 CCCCTTCTTTATAGCCAGCA 5-10-5 90 33204 33223 704 546343 1761 1780 CCCCCTTCTTTATAGCCAGC 5-10-5 97 33205 33224 705 547483 1762 1781 TCCCCCTTCTTTATAGCCAG 5-10-5 71 33206 33225 706 546345 1773 1792 CAAGCATCTTTTCCCCCTTC 5-10-5 86 33217 33236 707 546346 1796 1815 AGGGACCACCTGAATCTCCC 5-10-5 83 33895 33914 708 547484 1799 1818 CTAAGGGACCACCTGAATCT 5-10-5 69 33898 33917 709 546347 1800 1819 ACTAAGGGACCACCTGAATC 5-10-5 28 33899 33918 710 547485 1803 1822 CAAACTAAGGGACCACCTGA 5-10-5 49 33902 33921 711 546348 1804 1823 GCAAACTAAGGGACCACCTG 5-10-5 79 33903 33922 712 547486 1805 1824 TGCAAACTAAGGGACCACCT 5-10-5 89 33904 33923 713 546349 1810 1829 GTGTTTGCAAACTAAGGGAC 5-10-5 48 33909 33928 714 547487 1811 1830 TGTGTTTGCAAACTAAGGGA 5-10-5 72 33910 33929 715 546350 1868 1887 CCCTGCGGGCACAGCCTTCA 5-10-5 88 33967 33986 716 546351 1873 1892 TTGCTCCCTGCGGGCACAGC 5-10-5 82 33972 33991 717 546352 1880 1899 CACCAGGTTGCTCCCTGCGG 5-10-5 75 33979 33998 718 547488 1881 1900 ACACCAGGTTGCTCCCTGCG 5-10-5 71 33980 33999 719
TABLE-US-00122 TABLE 131 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 72 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547448 1451 1470 TTATTTGTGAGAAAGGTGTA 5-10-5 83 30529 30548 643 547449 1453 1472 TTTTATTTGTGAGAAAGGTG 5-10-5 73 30531 30550 644 546304 1454 1473 CTTTTATTTGTGAGAAAGGT 5-10-5 86 30532 30551 645 547450 1456 1475 CTCTTTTATTTGTGAGAAAG 5-10-5 67 30534 30553 646 547451 1471 1490 TTGGTGAATAATAATCTCTT 5-10-5 64 30549 30568 647 546306 1472 1491 TTTGGTGAATAATAATCTCT 5-10-5 71 30550 30569 648 547452 1474 1493 GTTTTGGTGAATAATAATCT 5-10-5 62 30552 30571 649 546307 1478 1497 TATAGTTTTGGTGAATAATA 5-10-5 0 30556 30575 650 546308 1482 1501 ACTTTATAGTTTTGGTGAAT 5-10-5 43 30560 30579 651 546309 1492 1511 CCCTTCTGAGACTTTATAGT 5-10-5 81 30570 30589 652 546310 1496 1515 GATTCCCTTCTGAGACTTTA 5-10-5 67 30574 30593 653 546311 1499 1518 CATGATTCCCTTCTGAGACT 5-10-5 76 30577 30596 654 547453 1500 1519 TCATGATTCCCTTCTGAGAC 5-10-5 81 30578 30597 655 547454 1502 1521 TATCATGATTCCCTTCTGAG 5-10-5 78 30580 30599 656 547455 1503 1522 ATATCATGATTCCCTTCTGA 5-10-5 66 30581 30600 657 547456 1506 1525 GCGATATCATGATTCCCTTC 5-10-5 96 30584 30603 658 546313 1507 1526 GGCGATATCATGATTCCCTT 5-10-5 75 30585 30604 659 547457 1509 1528 AAGGCGATATCATGATTCCC 5-10-5 92 30587 30606 660 547458 1513 1532 TATCAAGGCGATATCATGAT 5-10-5 64 30591 30610 661 547459 1519 1538 GAGTTTTATCAAGGCGATAT 5-10-5 51 30597 30616 662 547460 1522 1541 CTGGAGTTTTATCAAGGCGA 5-10-5 75 30600 30619 663 546316 1524 1543 GCCTGGAGTTTTATCAAGGC 5-10-5 60 30602 30621 664 546317 1528 1547 AGGAGCCTGGAGTTTTATCA 5-10-5 31 30606 30625 665 546318 1534 1553 ATTCAAAGGAGCCTGGAGTT 5-10-5 46 30612 30631 666 547461 1537 1556 GTAATTCAAAGGAGCCTGGA 5-10-5 55 30615 30634 667 547462 1539 1558 GTGTAATTCAAAGGAGCCTG 5-10-5 54 30617 30636 668 546319 1541 1560 CAGTGTAATTCAAAGGAGCC 5-10-5 61 30619 30638 669 547463 1564 1583 TAGGCATATTGGTTTTTGGA 5-10-5 84 31870 31889 670 546320 1566 1585 GGTAGGCATATTGGTTTTTG 5-10-5 69 31872 31891 671 546321 1569 1588 GAAGGTAGGCATATTGGTTT 5-10-5 56 31875 31894 672 546322 1584 1603 CTTGTGTCACCTTTGGAAGG 5-10-5 68 31890 31909 673 547464 1585 1604 GCTTGTGTCACCTTTGGAAG 5-10-5 84 31891 31910 674 546323 1587 1606 GTGCTTGTGTCACCTTTGGA 5-10-5 80 31893 31912 675 547465 1592 1611 AAATTGTGCTTGTGTCACCT 5-10-5 85 31898 31917 676 547466 1596 1615 GTATAAATTGTGCTTGTGTC 5-10-5 43 31902 31921 677 546324 1597 1616 GGTATAAATTGTGCTTGTGT 5-10-5 82 31903 31922 678 547467 1598 1617 TGGTATAAATTGTGCTTGTG 5-10-5 65 31904 31923 679 547468 1600 1619 GTTGGTATAAATTGTGCTTG 5-10-5 46 31906 31925 680 546325 1602 1621 CAGTTGGTATAAATTGTGCT 5-10-5 79 31908 31927 681 546326 1607 1626 CCCAACAGTTGGTATAAATT 5-10-5 64 31913 31932 682 547469 1610 1629 TTACCCAACAGTTGGTATAA 5-10-5 50 31916 31935 683 546327 1612 1631 GGTTACCCAACAGTTGGTAT 5-10-5 84 31918 31937 684 546328 1624 1643 GAAGCCCCATCCGGTTACCC 5-10-5 81 31930 31949 685 547470 1628 1647 TCGAGAAGCCCCATCCGGTT 5-10-5 68 31934 31953 686 546329 1631 1650 CCTTCGAGAAGCCCCATCCG 5-10-5 8 31937 31956 687 546330 1636 1655 TTTCTCCTTCGAGAAGCCCC 5-10-5 67 31942 31961 688 547471 1638 1657 CCTTTCTCCTTCGAGAAGCC 5-10-5 43 31944 31963 689 547472 1641 1660 TCACCTTTCTCCTTCGAGAA 5-10-5 42 n/a n/a 690 546331 1642 1661 TTCACCTTTCTCCTTCGAGA 5-10-5 44 n/a n/a 691 547473 1649 1668 TTTGGATTTCACCTTTCTCC 5-10-5 26 n/a n/a 692 547474 1659 1678 TGTAGAATATTTTGGATTTC 5-10-5 34 33103 33122 693 547475 1686 1705 TTTGTTACCAAAGGAATATT 5-10-5 42 33130 33149 694 547476 1688 1707 CATTTGTTACCAAAGGAATA 5-10-5 71 33132 33151 695 546336 1689 1708 TCATTTGTTACCAAAGGAAT 5-10-5 73 33133 33152 696 547477 1692 1711 TCTTCATTTGTTACCAAAGG 5-10-5 68 33136 33155 697 547478 1695 1714 CATTCTTCATTTGTTACCAA 5-10-5 55 33139 33158 698 546339 1712 1731 CTTGATATCTTTTCTGGCAT 5-10-5 64 33156 33175 699 546340 1716 1735 TAATCTTGATATCTTTTCTG 5-10-5 56 33160 33179 700 547479 1718 1737 TATAATCTTGATATCTTTTC 5-10-5 9 33162 33181 701 547480 1756 1775 TTCTTTATAGCCAGCACAGA 5-10-5 49 33200 33219 702 547481 1758 1777 CCTTCTTTATAGCCAGCACA 5-10-5 77 33202 33221 703 547482 1760 1779 CCCCTTCTTTATAGCCAGCA 5-10-5 65 33204 33223 704 546343 1761 1780 CCCCCTTCTTTATAGCCAGC 5-10-5 91 33205 33224 705 547483 1762 1781 TCCCCCTTCTTTATAGCCAG 5-10-5 77 33206 33225 706 546345 1773 1792 CAAGCATCTTTTCCCCCTTC 5-10-5 80 33217 33236 707 546346 1796 1815 AGGGACCACCTGAATCTCCC 5-10-5 70 33895 33914 708 547484 1799 1818 CTAAGGGACCACCTGAATCT 5-10-5 64 33898 33917 709 546347 1800 1819 ACTAAGGGACCACCTGAATC 5-10-5 22 33899 33918 710 547485 1803 1822 CAAACTAAGGGACCACCTGA 5-10-5 66 33902 33921 711 546348 1804 1823 GCAAACTAAGGGACCACCTG 5-10-5 76 33903 33922 712 547486 1805 1824 TGCAAACTAAGGGACCACCT 5-10-5 78 33904 33923 713 546349 1810 1829 GTGTTTGCAAACTAAGGGAC 5-10-5 35 33909 33928 714 547487 1811 1830 TGTGTTTGCAAACTAAGGGA 5-10-5 61 33910 33929 715 546350 1868 1887 CCCTGCGGGCACAGCCTTCA 5-10-5 74 33967 33986 716 546351 1873 1892 TTGCTCCCTGCGGGCACAGC 5-10-5 60 33972 33991 717 546352 1880 1899 CACCAGGTTGCTCCCTGCGG 5-10-5 74 33979 33998 718 547488 1881 1900 ACACCAGGTTGCTCCCTGCG 5-10-5 72 33980 33999 719
TABLE-US-00123 TABLE 132 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 90 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547489 1883 1902 AGACACCAGGTTGCTCCCTG 5-10-5 34 33982 34001 720 547490 1885 1904 GTAGACACCAGGTTGCTCCC 5-10-5 55 33984 34003 721 546353 1900 1919 CTCAGCGACTTTGGTGTAGA 5-10-5 55 33999 34018 722 546354 1903 1922 GTACTCAGCGACTTTGGTGT 5-10-5 47 34002 34021 723 547491 1906 1925 CATGTACTCAGCGACTTTGG 5-10-5 47 34005 34024 724 547492 1911 1930 CAGTCCATGTACTCAGCGAC 5-10-5 62 34010 34029 725 546356 1913 1932 TCCAGTCCATGTACTCAGCG 5-10-5 60 34012 34031 726 546357 1947 1966 GCTTTTCCATCACTGCTCTG 5-10-5 79 34046 34065 727 546358 1951 1970 CTGAGCTTTTCCATCACTGC 5-10-5 83 34050 34069 728 547493 1952 1971 TCTGAGCTTTTCCATCACTG 5-10-5 72 34051 34070 729 546359 1955 1974 GCATCTGAGCTTTTCCATCA 5-10-5 79 34054 34073 730 546360 1958 1977 ACTGCATCTGAGCTTTTCCA 5-10-5 13 34057 34076 731 547494 1963 1982 TGGTGACTGCATCTGAGCTT 5-10-5 70 34062 34081 732 547495 1965 1984 GCTGGTGACTGCATCTGAGC 5-10-5 61 34064 34083 733 547496 1967 1986 ATGCTGGTGACTGCATCTGA 5-10-5 80 34066 34085 734 546362 1969 1988 TCATGCTGGTGACTGCATCT 5-10-5 71 34068 34087 735 546363 1973 1992 CTTCTCATGCTGGTGACTGC 5-10-5 81 34072 34091 736 547497 1977 1996 ACTGCTTCTCATGCTGGTGA 5-10-5 68 34076 34095 737 546364 1979 1998 GGACTGCTTCTCATGCTGGT 5-10-5 61 34078 34097 738 547498 1981 2000 CTGGACTGCTTCTCATGCTG 5-10-5 44 34080 34099 739 547499 1983 2002 CTCTGGACTGCTTCTCATGC 5-10-5 65 34082 34101 740 546365 1986 2005 AGACTCTGGACTGCTTCTCA 5-10-5 64 34085 34104 741 547500 1989 2008 CCTAGACTCTGGACTGCTTC 5-10-5 65 34088 34107 742 546366 1991 2010 TGCCTAGACTCTGGACTGCT 5-10-5 79 34090 34109 743 547501 1993 2012 ATTGCCTAGACTCTGGACTG 5-10-5 55 34092 34111 744 546367 1997 2016 AAAAATTGCCTAGACTCTGG 5-10-5 61 34096 34115 745 546368 2003 2022 GGTTGTAAAAATTGCCTAGA 5-10-5 44 34102 34121 746 547502 2006 2025 TCAGGTTGTAAAAATTGCCT 5-10-5 64 34105 34124 747 546369 2007 2026 CTCAGGTTGTAAAAATTGCC 5-10-5 51 34106 34125 748 547503 2008 2027 ACTCAGGTTGTAAAAATTGC 5-10-5 66 34107 34126 749 547504 2010 2029 GAACTCAGGTTGTAAAAATT 5-10-5 37 34109 34128 750 546370 2014 2033 ACTTGAACTCAGGTTGTAAA 5-10-5 34 34113 34132 751 547505 2015 2034 GACTTGAACTCAGGTTGTAA 5-10-5 69 34114 34133 752 546372 2021 2040 GAATTTGACTTGAACTCAGG 5-10-5 49 34120 34139 753 546373 2025 2044 CTCAGAATTTGACTTGAACT 5-10-5 59 34124 34143 754 547506 2028 2047 AGGCTCAGAATTTGACTTGA 5-10-5 78 34127 34146 755 547507 2029 2048 CAGGCTCAGAATTTGACTTG 5-10-5 56 34128 34147 756 546374 2030 2049 CCAGGCTCAGAATTTGACTT 5-10-5 50 34129 34148 757 547508 2032 2051 CCCCAGGCTCAGAATTTGAC 5-10-5 69 34131 34150 758 547509 2034 2053 CCCCCCAGGCTCAGAATTTG 5-10-5 58 34133 34152 759 546375 2036 2055 GACCCCCCAGGCTCAGAATT 5-10-5 48 34135 34154 760 547510 2041 2060 ATGAGGACCCCCCAGGCTCA 5-10-5 40 34140 34159 761 547511 2042 2061 GATGAGGACCCCCCAGGCTC 5-10-5 53 34141 34160 762 547512 2045 2064 GCAGATGAGGACCCCCCAGG 5-10-5 74 34144 34163 763 547513 2046 2065 TGCAGATGAGGACCCCCCAG 5-10-5 72 34145 34164 764 546378 2048 2067 TTTGCAGATGAGGACCCCCC 5-10-5 79 34147 34166 765 546379 2056 2075 CTCCATGCTTTGCAGATGAG 5-10-5 69 34155 34174 766 546380 2062 2081 GCCACTCTCCATGCTTTGCA 5-10-5 81 34161 34180 767 547514 2066 2085 AGATGCCACTCTCCATGCTT 5-10-5 85 34165 34184 768 546381 2068 2087 GAAGATGCCACTCTCCATGC 5-10-5 73 34167 34186 769 547515 2069 2088 AGAAGATGCCACTCTCCATG 5-10-5 58 34168 34187 770 546382 2072 2091 CAAAGAAGATGCCACTCTCC 5-10-5 58 34171 34190 771 547516 2076 2095 GATGCAAAGAAGATGCCACT 5-10-5 48 34175 34194 772 546383 2077 2096 GGATGCAAAGAAGATGCCAC 5-10-5 57 34176 34195 773 547517 2079 2098 TAGGATGCAAAGAAGATGCC 5-10-5 57 34178 34197 774 547518 2083 2102 TCCTTAGGATGCAAAGAAGA 5-10-5 51 34182 34201 775 546384 2085 2104 CGTCCTTAGGATGCAAAGAA 5-10-5 81 34184 34203 776 546385 2120 2139 ATTGTCCTCAGCAGCTCTGA 5-10-5 67 34219 34238 777 547519 2126 2145 CCAGACATTGTCCTCAGCAG 5-10-5 76 34225 34244 778 546386 2128 2147 AGCCAGACATTGTCCTCAGC 5-10-5 78 34227 34246 779 547520 2130 2149 TCAGCCAGACATTGTCCTCA 5-10-5 76 34229 34248 780 547521 2132 2151 CTTCAGCCAGACATTGTCCT 5-10-5 58 34231 34250 781 546387 2138 2157 AGCGGGCTTCAGCCAGACAT 5-10-5 77 34237 34256 782 547522 2141 2160 GAAAGCGGGCTTCAGCCAGA 5-10-5 73 34240 34259 783 546388 2143 2162 CTGAAAGCGGGCTTCAGCCA 5-10-5 71 34242 34261 784 546389 2147 2166 CGTGCTGAAAGCGGGCTTCA 5-10-5 71 34246 34265 785 546390 2165 2184 GTCAGCCCCTGGTTACGGCG 5-10-5 70 34264 34283 786 547523 2167 2186 TTGTCAGCCCCTGGTTACGG 5-10-5 69 34266 34285 787 547524 2169 2188 CATTGTCAGCCCCTGGTTAC 5-10-5 58 34268 34287 788 546391 2170 2189 GCATTGTCAGCCCCTGGTTA 5-10-5 54 34269 34288 789 547525 2174 2193 CCTCGCATTGTCAGCCCCTG 5-10-5 78 34273 34292 790 546392 2176 2195 GACCTCGCATTGTCAGCCCC 5-10-5 72 34275 34294 791 547526 2178 2197 GCGACCTCGCATTGTCAGCC 5-10-5 59 34277 34296 792 547527 2185 2204 CTCAGTTGCGACCTCGCATT 5-10-5 58 34284 34303 793 546393 2186 2205 TCTCAGTTGCGACCTCGCAT 5-10-5 77 34285 34304 794 546394 2196 2215 GTCATGGAGATCTCAGTTGC 5-10-5 71 34295 34314 795 547528 2200 2219 CACAGTCATGGAGATCTCAG 5-10-5 78 34299 34318 796
TABLE-US-00124 TABLE 133 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 90 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546403 n/a n/a CCATGAACATCCTATCCGTG 5-10-5 83 3282 3301 797 546406 n/a n/a TGTCCTGTCAACATATTCCA 5-10-5 80 3299 3318 798 546409 n/a n/a GGGTTTCTGCCAACAGTTTC 5-10-5 77 3326 3345 799 546410 n/a n/a GACTTTGGGTTTCTGCCAAC 5-10-5 83 3332 3351 800 546411 n/a n/a ATATTGACTTTGGGTTTCTG 5-10-5 56 3337 3356 801 546412 n/a n/a GGCTTCAATATTGACTTTGG 5-10-5 84 3344 3363 802 546416 n/a n/a CTGCAGGCAATATTTTGCTT 5-10-5 62 3364 3383 803 546418 n/a n/a ATGTGGCACTGCAGGCAATA 5-10-5 72 3372 3391 804 546419 n/a n/a TTCTAATGTGGCACTGCAGG 5-10-5 65 3377 3396 805 546421 n/a n/a TCAAGCTGTTCTAATGTGGC 5-10-5 71 3385 3404 806 546422 n/a n/a ACGGTCTTCAAGCTGTTCTA 5-10-5 72 3392 3411 807 546425 n/a n/a GGTCAATCTGACTAGTGAAT 5-10-5 69 2284 2303 808 546426 n/a n/a TCTCTGGTCAATCTGACTAG 5-10-5 49 2289 2308 809 546429 n/a n/a GCCCACCAACAATCTCTGGT 5-10-5 84 2301 2320 810 546432 n/a n/a GACCCCAACAGACAGCCCAC 5-10-5 62 2315 2334 811 546444 n/a n/a CCAGAATCATGCCTTGTGGG 5-10-5 61 4765 4784 812 546447 n/a n/a GTCACCATAGACCCAGAATC 5-10-5 68 4777 4796 813 546450 n/a n/a GTGGCCCTCTTAAGTCACCA 5-10-5 73 4790 4809 814 546453 n/a n/a CTCATTGTTGTGTGGCCCTC 5-10-5 82 4801 4820 815 546459 n/a n/a GTAGCCATACATCTGAGGAA 5-10-5 46 4830 4849 816 546461 n/a n/a ATGTTTATTGTAGCCATACA 5-10-5 53 4839 4858 817 546492 n/a n/a CTCGCCTTTGTGACTCGATT 5-10-5 61 26263 26282 818 546493 n/a n/a CATACTCGCCTTTGTGACTC 5-10-5 35 26267 26286 819 546494 n/a n/a GCATACTCGCCTTTGTGACT 5-10-5 67 26268 26287 820 546495 n/a n/a TGCATACTCGCCTTTGTGAC 5-10-5 65 26269 26288 821 546395 2209 2228 TTCACAACACACAGTCATGG 5-10-5 72 34308 34327 822 546397 2233 2252 TTTTTTGATCTTTCACCATT 5-10-5 55 n/a n/a 823 546496 n/a n/a ATGCATACTCGCCTTTGTGA 5-10-5 54 26270 26289 824 26301 26320 546497 n/a n/a CATGCATACTCGCCTTTGTG 5-10-5 56 26271 26290 825 26302 26321 546498 n/a n/a CCATGCATACTCGCCTTTGT 5-10-5 65 26272 26291 826 26303 26322 547529 2203 2222 ACACACAGTCATGGAGATCT 5-10-5 49 34302 34321 827 547530 2206 2225 ACAACACACAGTCATGGAGA 5-10-5 63 34305 34324 828 547531 2213 2232 TTATTTCACAACACACAGTC 5-10-5 69 34312 34331 829 546499 n/a n/a TCCATGCATACTCGCCTTTG 5-10-5 20 26273 26292 830 546500 n/a n/a TTCCATGCATACTCGCCTTT 5-10-5 46 26274 26293 831 546501 n/a n/a TTTCCATGCATACTCGCCTT 5-10-5 53 26275 26294 832 546502 n/a n/a GATTTTCCATGCATACTCGC 5-10-5 37 26278 26297 833 546503 n/a n/a GTGATGCGATTTTCCATGCA 5-10-5 53 26285 26304 834 546508 n/a n/a GCAGCAAGTGCTCCCCATGC 5-10-5 43 26317 26336 835 546511 n/a n/a GTGATGAAAGTACAGCAGCA 5-10-5 50 26331 26350 836 546683 n/a n/a TCCTATCCGTGTTCAGCTGT 5-10-5 69 3273 3292 837 546684 n/a n/a TACTCTCTACATACTCAGGA 5-10-5 71 3561 3580 838 546687 n/a n/a TGAGACCTCCAGACTACTGT 5-10-5 76 3847 3866 839 546690 n/a n/a CTCTGCTGGTTTTAGACCAC 5-10-5 44 4027 4046 840 546695 n/a n/a GGGACAATCTCCACCCCCGA 5-10-5 36 4225 4244 841 546698 n/a n/a TGCAGAGTGTCATCTGCGAA 5-10-5 59 4387 4406 842 546700 n/a n/a TGGTTCCCTAGCGGTCCAGA 5-10-5 78 4561 4580 843 546705 n/a n/a CCCCTGTAGTTGGCTGTGGT 5-10-5 66 5046 5065 844 546707 n/a n/a GCAAGTCAAAGAGTGTCCAC 5-10-5 73 5283 5302 845 546710 n/a n/a GAAGCCTGTTAGAGTTGGCC 5-10-5 73 5576 5595 846 546719 n/a n/a CCCCCATGTCCATGGACTTT 5-10-5 55 6329 6348 847 547532 n/a n/a CTGCCAACAGTTTCAACTTT 5-10-5 65 3320 3339 848 547533 n/a n/a TTTTGCTTGGCTTCAATATT 5-10-5 23 3352 3371 849 547534 n/a n/a ATCTGACTAGTGAATGGCTT 5-10-5 72 2279 2298 850 547535 n/a n/a AGACAGCCCACCAACAATCT 5-10-5 28 2306 2325 851 547536 n/a n/a TGCATAGACCCCAACAGACA 5-10-5 48 2321 2340 852 547537 n/a n/a CCTGTGCATAGACCCCAACA 5-10-5 65 2325 2344 853 547538 n/a n/a CCAGCAGAAATCCTGTGCAT 5-10-5 77 2336 2355 854 547539 n/a n/a AGAACTCCAGCAGAAATCCT 5-10-5 43 2342 2361 855 547540 n/a n/a TTGTGTGGCCCTCTTAAGTC 5-10-5 44 4794 4813 856 547541 n/a n/a TATAGATGTTTATTGTAGCC 5-10-5 36 4844 4863 857 547542 n/a n/a ATACTCGCCTTTGTGACTCG 5-10-5 35 26266 26285 858 547543 n/a n/a TTTTCCATGCATACTCGCCT 5-10-5 54 26276 26295 859 547544 n/a n/a TCGCCTTTGTGATGCGATTT 5-10-5 15 26293 26312 860 547545 n/a n/a ATACTCGCCTTTGTGATGCG 5-10-5 43 26297 26316 861 547546 n/a n/a CATACTCGCCTTTGTGATGC 5-10-5 11 26298 26317 862 547547 n/a n/a GCATACTCGCCTTTGTGATG 5-10-5 42 26299 26318 863 547548 n/a n/a TGCATACTCGCCTTTGTGAT 5-10-5 61 26300 26319 864 547549 n/a n/a CCCATGCATACTCGCCTTTG 5-10-5 36 26304 26323 865 547550 n/a n/a CCCCATGCATACTCGCCTTT 5-10-5 53 26305 26324 866 547551 n/a n/a TCCCCATGCATACTCGCCTT 5-10-5 38 26306 26325 867 547552 n/a n/a CTCCCCATGCATACTCGCCT 5-10-5 53 26307 26326 868 547553 n/a n/a TGCTCCCCATGCATACTCGC 5-10-5 64 26309 26328 869 547554 n/a n/a GCTCTGATTGGGTCACCACA 5-10-5 50 5743 5762 870 547555 n/a n/a TGTCTCCTTCCACTTGCTCC 5-10-5 58 5923 5942 871 547556 n/a n/a GCCATTTTATCCCTGAGATT 5-10-5 55 6130 6149 872 547557 n/a n/a CTGTGCTGTATTTTGGAGCC 5-10-5 59 6413 6432 873
TABLE-US-00125 TABLE 134 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: NO: 1 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 85 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546732 n/a n/a GGATTTGGCCCTGAGCCCCA 5-10-5 14 6933 6952 874 546735 n/a n/a CAACCTGTCCATTCCCTGGG 5-10-5 46 7082 7101 875 546739 n/a n/a ATTCGGTGTCTTTACTGGCT 5-10-5 89 7228 7247 876 546746 n/a n/a TCCTGTTGCCTGACATGCTA 5-10-5 65 7694 7713 877 546747 n/a n/a CTCCCACTGACTGACTACTC 5-10-5 64 7904 7923 878 546749 n/a n/a GCTGGTCCTTGAACCCCGTG 5-10-5 53 8259 8278 879 546753 n/a n/a CTGGCTCACTATAGGCCCCA 5-10-5 91 8655 8674 880 546756 n/a n/a ATAAGCATCTCTCTGACCTA 5-10-5 47 9105 9124 881 546763 n/a n/a GCTTCCCCAATACTTGCTGG 5-10-5 84 9695 9714 882 546765 n/a n/a GTGTCCAGAATACTGCCCCA 5-10-5 82 10053 10072 883 546770 n/a n/a GTGGACGACTGCCCTGTGCC 5-10-5 74 10435 10454 884 546773 n/a n/a TCTCTAGCATCCTAGTCCTC 5-10-5 67 10586 10605 885 546780 n/a n/a ATACTGGCTAAGTCAGGCCC 5-10-5 83 10982 11001 886 546784 n/a n/a GGCAGGGAGGTGGATTATTC 5-10-5 58 11440 11459 887 546789 n/a n/a GCTTCTCTATCTCCCAGTGT 5-10-5 79 12228 12247 888 546791 n/a n/a GATGCATGCAGCAATACAGG 5-10-5 52 12385 12404 889 546795 n/a n/a GTCTCGATGGCAAGCTGTAC 5-10-5 72 12650 12669 890 546796 n/a n/a GTACTCACCGGTACTCTGCC 5-10-5 82 12804 12823 891 546799 n/a n/a ATGAAGGGCGAGGCGCAGTG 5-10-5 5 13258 13277 892 546803 n/a n/a CCCCATACATCTATGCAAAT 5-10-5 40 13551 13570 893 546804 n/a n/a ACATGACTCCAGTGATGGAT 5-10-5 57 13632 13651 894 546808 n/a n/a AAAATGACACCAAAATTCGC 5-10-5 0 13841 13860 895 546811 n/a n/a TGGACATCCTTCCCCTCGCA 5-10-5 49 13967 13986 896 546817 n/a n/a GCTCTGAGCCTTCCGCCTCT 5-10-5 77 14472 14491 897 546822 n/a n/a ACTAGTTTCCTATAACTGCT 5-10-5 32 14735 14754 898 546823 n/a n/a TACTAGTTTCCTATAACTGC 5-10-5 44 14736 14755 899 546824 n/a n/a GTACTAGTTTCCTATAACTG 5-10-5 79 14737 14756 900 546825 n/a n/a GTATCACTGTACTAGTTTCC 5-10-5 96 14745 14764 901 14816 14835 14887 14906 14946 14965 15006 15025 15078 15097 15221 15240 15293 15312 15352 15371 15412 15431 15484 15503 15556 15575 15614 15633 15686 15705 15816 15835 15888 15907 15946 15965 546826 n/a n/a AGTATCACTGTACTAGTTTC 5-10-5 90 14746 14765 902 14817 14836 14888 14907 14947 14966 15007 15026 15079 15098 15222 15241 15294 15313 15353 15372 15413 15432 15485 15504 15557 15576 15615 15634 15687 15706 15817 15836 15889 15908 15947 15966 546827 n/a n/a CAGTATCACTGTACTAGTTT 5-10-5 98 14747 14766 903 14818 14837 14889 14908 14948 14967 15008 15027 15080 15099 15152 15171 15223 15242 15295 15314 15354 15373 15414 15433 15486 15505 15558 15577 15616 15635 15688 15707 15818 15837 15890 15909 15948 15967 546828 n/a n/a ACAGTATCACTGTACTAGTT 5-10-5 95 14748 14767 904 14819 14838 14890 14909 14949 14968 15009 15028 15081 15100 15153 15172 15224 15243 15296 15315 15355 15374 15415 15434 15487 15506 15559 15578 15617 15636 15689 15708 15819 15838 15891 15910 15949 15968 546829 n/a n/a AACAGTATCACTGTACTAGT 5-10-5 94 14749 14768 905 14820 14839 14891 14910 14950 14969 15010 15029 15082 15101 15154 15173 15225 15244 15297 15316 15356 15375 15416 15435 15488 15507 15560 15579 15618 15637 15690 15709 15820 15839 15892 15911 15950 15969 546830 n/a n/a TAACAGTATCACTGTACTAG 5-10-5 78 14750 14769 906 14821 14840 14892 14911 14951 14970 15011 15030 15083 15102 15155 15174 15226 15245 15298 15317 15357 15376 15417 15436 15489 15508 15561 15580 15619 15638 15691 15710 15821 15840 15893 15912 15951 15970 546831 n/a n/a TCTAACAGTATCACTGTACT 5-10-5 79 14752 14771 907 14823 14842 14894 14913 15013 15032 15085 15104 15228 15247 15300 15319 15419 15438 15491 15510 15621 15640 15823 15842 15953 15972 546832 n/a n/a CTCTAACAGTATCACTGTAC 5-10-5 88 14753 14772 908 14824 14843 14895 14914 15014 15033 15086 15105 15229 15248 15301 15320 15420 15439 15492 15511 15622 15641 15824 15843 15954 15973 546833 n/a n/a ACTCTAACAGTATCACTGTA 5-10-5 90 14754 14773 909 14825 14844 14896 14915 15015 15034 15087 15106 15230 15249 15302 15321 15421 15440 15493 15512 15623 15642 15825 15844 15955 15974 546834 n/a n/a AACTCTAACAGTATCACTGT 5-10-5 86 14755 14774 910 14826 14845 14897 14916 15016 15035 15088 15107 15231 15250 15303 15322 15422 15441 15494 15513 15624 15643 15826 15845 15956 15975 546835 n/a n/a TAACTCTAACAGTATCACTG 5-10-5 86 14756 14775 911 14827 14846 14898 14917 15017 15036
15089 15108 15232 15251 15304 15323 15423 15442 15495 15514 15625 15644 15827 15846 15957 15976 546836 n/a n/a ATAACTCTAACAGTATCACT 5-10-5 30 14757 14776 912 14828 14847 14899 14918 15018 15037 15090 15109 15233 15252 15305 15324 15424 15443 15496 15515 15626 15645 15828 15847 15958 15977 546837 n/a n/a TATAACTCTAACAGTATCAC 5-10-5 0 14758 14777 913 14829 14848 14900 14919 15019 15038 15091 15110 15234 15253 15306 15325 15425 15444 15497 15516 15627 15646 15829 15848 15959 15978 546838 n/a n/a CTATAACTCTAACAGTATCA 5-10-5 43 14759 14778 914 14830 14849 14901 14920 15020 15039 15092 15111 15235 15254 15307 15326 15426 15445 15498 15517 15628 15647 15830 15849 15960 15979 546839 n/a n/a CCTATAACTCTAACAGTATC 5-10-5 47 14760 14779 915 14831 14850 14902 14921 15021 15040 15093 15112 15236 15255 15308 15327 15427 15446 15499 15518 15629 15648 15831 15850 15961 15980 546840 n/a n/a CTGTCCTATAACTCTAACAG 5-10-5 53 14764 14783 916 14835 14854 546841 n/a n/a CACTGTCCTATAACTCTAAC 5-10-5 38 14766 14785 917 14837 14856 546842 n/a n/a TCACTGTCCTATAACTCTAA 5-10-5 54 14767 14786 918 14838 14857 546843 n/a n/a TATCACTGTCCTATAACTCT 5-10-5 52 14769 14788 919 14840 14859 546844 n/a n/a GTCCTATATCACTGTCCTAT 5-10-5 75 14775 14794 920 14846 14865 15180 15199 15716 15735 546845 n/a n/a TGTCCTATATCACTGTCCTA 5-10-5 75 14776 14795 921 14847 14866 15181 15200 15717 15736 546846 n/a n/a CTGTCCTATATCACTGTCCT 5-10-5 95 14777 14796 922 14848 14867 15182 15201 15718 15737 546847 n/a n/a ACTGTCCTATATCACTGTCC 5-10-5 88 14778 14797 923 14849 14868 15183 15202 15719 15738 546848 n/a n/a TCACTGTCCTATATCACTGT 5-10-5 86 14780 14799 924 14851 14870 14976 14995 15185 15204 15257 15276 15382 15401 15520 15539 15650 15669 15721 15740 15852 15871 15982 16001 547558 n/a n/a CCCCCAGTTCCCATGCAAGG 5-10-5 52 6640 6659 925 547559 n/a n/a GAGCACAGATCTCTTCAAGT 5-10-5 69 6822 6841 926 547560 n/a n/a GACGGTCACCCAGCCCTGAC 5-10-5 42 7459 7478 927 547561 n/a n/a AAGGGAAATTAGAGGCAGGC 5-10-5 57 7583 7602 928 547562 n/a n/a CTTTCTTGAGACAATCCCTT 5-10-5 59 8463 8482 929 547563 n/a n/a GTGGGATCAGAGAATGACTA 5-10-5 48 9267 9286 930 547564 n/a n/a CCCTCTGTCTTAGATGTCCA 5-10-5 94 9390 9409 931 547565 n/a n/a CTTATCAGTCCCAGTCATGT 5-10-5 63 10698 10717 932 547566 n/a n/a AAGAGTTGGGATGCGACTCT 5-10-5 76 11335 11354 933 547567 n/a n/a TCCACTCCTAAGAAGTATGG 5-10-5 60 11546 11565 934 547568 n/a n/a GCACCCTTTTCATTGAGATT 5-10-5 70 12070 12089 935 547569 n/a n/a ACTACCATTTGGGTTGGTAG 5-10-5 9 12571 12590 936 547570 n/a n/a AAGCCCTGTTTGGTTTTTAG 5-10-5 18 12900 12919 937 547571 n/a n/a AAATGACACCAAAATTGAGT 5-10-5 14 13744 13763 938 547572 n/a n/a AAATGACACCAAAATTCGCT 5-10-5 40 13840 13859 939 547573 n/a n/a TAAGCAAGGCCTATGTGTGG 5-10-5 2 13880 13899 940 547574 n/a n/a ACACGCACAGGTCCCAGGGC 5-10-5 51 14314 14333 941 547575 n/a n/a GGGAAACTCTTTCCTCGCCC 5-10-5 89 14583 14602 942 547576 n/a n/a CTAGTTTCCTATAACTGCTG 5-10-5 29 14734 14753 943 547577 n/a n/a CTAACAGTATCACTGTACTA 5-10-5 79 14751 14770 944 14822 14841 14893 14912 15012 15031 15084 15103 15227 15246 15299 15318 15418 15437 15490 15509 15620 15639 15822 15841 15952 15971 547578 n/a n/a GTCCTATAACTCTAACAGTA 5-10-5 30 14762 14781 945 14833 14852 547579 n/a n/a TGTCCTATAACTCTAACAGT 5-10-5 0 14763 14782 946 14834 14853 547580 n/a n/a ATCACTGTCCTATAACTCTA 5-10-5 61 14768 14787 947 14839 14858 547581 n/a n/a ATATCACTGTCCTATAACTC 5-10-5 60 14770 14789 948 14841 14860 547582 n/a n/a TATATCACTGTCCTATAACT 5-10-5 22 14771 14790 949 14842 14861 15176 15195 15712 15731 16160 16179 547583 n/a n/a CACTGTCCTATATCACTGTC 5-10-5 80 14779 14798 950 14850 14869 15184 15203 15720 15739
TABLE-US-00126 TABLE 135 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 85 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546849 n/a n/a ATCACTGTCCTATATCACTG 5-10-5 93 14781 14800 951 14852 14871 14977 14996 15186 15205 15258 15277 15383 15402 15521 15540 15651 15670 15722 15741 15853 15872 15983 16002 546850 n/a n/a TATCACTGTCCTATATCACT 5-10-5 80 14782 14801 952 14853 14872 14978 14997 15116 15135 15187 15206 15259 15278 15384 15403 15522 15541 15652 15671 15723 15742 15854 15873 15984 16003 546851 n/a n/a AGTATCACTGTCCTATATCA 5-10-5 81 14784 14803 953 14980 14999 15118 15137 15386 15405 15524 15543 15986 16005 546852 n/a n/a CAGTATCACTGTCCTATATC 5-10-5 94 14785 14804 954 14981 15000 15119 15138 15387 15406 15525 15544 15987 16006 546853 n/a n/a ACAGTATCACTGTCCTATAT 5-10-5 86 14786 14805 955 14982 15001 15120 15139 15388 15407 15526 15545 15988 16007 546854 n/a n/a TAACAGTATCACTGTCCTAT 5-10-5 90 14788 14807 956 14984 15003 15050 15069 15122 15141 15390 15409 15456 15475 15528 15547 15990 16009 546855 n/a n/a ATAACAGTATCACTGTCCTA 5-10-5 87 14789 14808 957 14985 15004 15051 15070 15123 15142 15391 15410 15457 15476 15529 15548 15991 16010 546856 n/a n/a AACTATAACAGTATCACTGT 5-10-5 54 14793 14812 958 15055 15074 15127 15146 15160 15179 15461 15480 15533 15552 15566 15585 15696 15715 15898 15917 15995 16014 546857 n/a n/a TATAACTATAACAGTATCAC 5-10-5 7 14796 14815 959 15058 15077 15130 15149 15163 15182 15464 15483 15536 15555 15569 15588 15699 15718 15770 15789 15998 16017 546858 n/a n/a CTATAACTATAACAGTATCA 5-10-5 21 14797 14816 960 15059 15078 15131 15150 15164 15183 15465 15484 15537 15556 15570 15589 15700 15719 15771 15790 15999 16018 546859 n/a n/a TTTCCTATAACTATAACAGT 5-10-5 7 14801 14820 961 15063 15082 15469 15488 15541 15560 546860 n/a n/a CTAGTTTCCTATAACTATAA 5-10-5 36 14805 14824 962 14876 14895 14935 14954 15067 15086 15210 15229 15282 15301 15341 15360 15473 15492 15545 15564 15603 15622 15675 15694 15746 15765 15805 15824 15877 15896 15935 15954 546861 n/a n/a TAACAATATCACTGTCCTAT 5-10-5 68 14859 14878 963 15193 15212 15265 15284 15586 15605 15658 15677 15729 15748 15860 15879 16086 16105 16183 16202 16234 16253 546862 n/a n/a AACTATAACAATATCACTGT 5-10-5 0 14864 14883 964 14923 14942 15198 15217 15270 15289 15329 15348 15591 15610 15663 15682 15734 15753 15793 15812 15865 15884 15923 15942 16066 16085 16091 16110 16144 16163 16239 16258 546863 n/a n/a TAACTATAACAATATCACTG 5-10-5 21 14865 14884 965 14924 14943 15199 15218 15271 15290 15330 15349 15592 15611 15664 15683 15735 15754 15794 15813 15866 15885 15924 15943 16067 16086 16092 16111 16145 16164 16240 16259 546864 n/a n/a ATAACTATAACAATATCACT 5-10-5 0 14866 14885 966 14925 14944 15200 15219 15272 15291 15331 15350 15593 15612 15665 15684 15736 15755 15795 15814 15867 15886 15925 15944 16068 16087 16093 16112 16146 16165 16241 16260 546865 n/a n/a TATAACTATAACAATATCAC 5-10-5 0 14867 14886 967 14926 14945 15201 15220 15273 15292 15332 15351 15594 15613 15666 15685 15737 15756 15796 15815 15868 15887 15926 15945 16069 16088 16094 16113 16147 16166 16242 16261 546866 n/a n/a GTTTCCTATAACTATAACAA 5-10-5 35 14873 14892 968 14932 14951 15207 15226 15279 15298 15338 15357 15600 15619 15672 15691 15743 15762 15802 15821 15874 15893 15932 15951 546867 n/a n/a ACCTATAACTCTAACAGTAT 5-10-5 40 14903 14922 969 15022 15041 15094 15113 15237 15256 15309 15328 15428 15447 15500 15519 15630 15649 15832 15851 15962 15981 546868 n/a n/a TACCTATAACTCTAACAGTA 5-10-5 51 14904 14923 970 15023 15042 15095 15114 15238 15257 15310 15329 15429 15448
15501 15520 15631 15650 15833 15852 15963 15982 546869 n/a n/a TGTACCTATAACTCTAACAG 5-10-5 53 14906 14925 971 15025 15044 15240 15259 15312 15331 15431 15450 15503 15522 15633 15652 15835 15854 15965 15984 546870 n/a n/a CTGTACCTATAACTCTAACA 5-10-5 87 14907 14926 972 15026 15045 15241 15260 15313 15332 15432 15451 15504 15523 15634 15653 15836 15855 15966 15985 546871 n/a n/a ACTGTACCTATAACTCTAAC 5-10-5 73 14908 14927 973 15027 15046 15242 15261 15314 15333 15433 15452 15505 15524 15635 15654 15837 15856 15967 15986 546872 n/a n/a CACTGTACCTATAACTCTAA 5-10-5 87 14909 14928 974 15028 15047 15243 15262 15315 15334 15434 15453 15506 15525 15636 15655 15838 15857 15968 15987 546873 n/a n/a CAATATCACTGTACCTATAA 5-10-5 34 14915 14934 975 15321 15340 15785 15804 546874 n/a n/a ATAACAATATCACTGTACCT 5-10-5 68 14919 14938 976 15325 15344 15789 15808 16062 16081 16140 16159 546875 n/a n/a ACTATAACAATATCACTGTA 5-10-5 33 14922 14941 977 15328 15347 15792 15811 16065 16084 16143 16162 546876 n/a n/a GTCCTATATCACTGTACCTG 5-10-5 87 14971 14990 978 546877 n/a n/a CACTGTCCTATATCACTGTA 5-10-5 88 14975 14994 979 15256 15275 15381 15400 15519 15538 15649 15668 15851 15870 15981 16000 546878 n/a n/a CCTATAACAGTATCACTGTC 5-10-5 81 14988 15007 980 15394 15413 546879 n/a n/a TTTCCTATAACAGTATCACT 5-10-5 42 14991 15010 981 15397 15416 546880 n/a n/a GTTTCCTATAACAGTATCAC 5-10-5 41 14992 15011 982 15398 15417 546881 n/a n/a AGTTTCCTATAACAGTATCA 5-10-5 49 14993 15012 983 15399 15418 546882 n/a n/a TAGTTTCCTATAACAGTATC 5-10-5 24 14994 15013 984 15400 15419 546883 n/a n/a CTAGTTTCCTATAACAGTAT 5-10-5 19 14995 15014 985 15401 15420 546884 n/a n/a ACTAGTTTCCTATAACAGTA 5-10-5 6 14996 15015 986 15402 15421 547584 n/a n/a GTATCACTGTCCTATATCAC 5-10-5 85 14783 14802 987 14979 14998 15117 15136 15385 15404 15523 15542 15985 16004 547585 n/a n/a AACAGTATCACTGTCCTATA 5-10-5 85 14787 14806 988 14983 15002 15121 15140 15389 15408 15527 15546 15989 16008 547586 n/a n/a TATAACAGTATCACTGTCCT 5-10-5 82 14790 14809 989 14986 15005 15052 15071 15124 15143 15392 15411 15458 15477 15530 15549 15992 16011 547587 n/a n/a CTATAACAGTATCACTGTCC 5-10-5 96 14791 14810 990 14987 15006 15053 15072 15125 15144 15393 15412 15459 15478 15531 15550 15993 16012 547588 n/a n/a ACTATAACAGTATCACTGTC 5-10-5 83 14792 14811 991 15054 15073 15126 15145 15460 15479 15532 15551 15994 16013 547589 n/a n/a TAACTATAACAGTATCACTG 5-10-5 36 14794 14813 992 15056 15075 15128 15147 15161 15180 15462 15481 15534 15553 15567 15586 15697 15716 15996 16015 547590 n/a n/a ATAACTATAACAGTATCACT 5-10-5 0 14795 14814 993 15057 15076 15129 15148 15162 15181 15463 15482 15535 15554 15568 15587 15698 15717 15997 16016 547591 n/a n/a CCTATAACTATAACAGTATC 5-10-5 23 14798 14817 994 15060 15079 15165 15184 15466 15485 15538 15557 15571 15590 15701 15720 15772 15791 16000 16019 547592 n/a n/a TCCTATAACTATAACAGTAT 5-10-5 27 14799 14818 995 15061 15080 15166 15185 15467 15486 15539 15558 15572 15591 15702 15721 16001 16020 547593 n/a n/a TTCCTATAACTATAACAGTA 5-10-5 29 14800 14819 996 15062 15081 15468 15487 15540 15559 547594 n/a n/a GTTTCCTATAACTATAACAG 5-10-5 19 14802 14821 997 15064 15083 15470 15489 15542 15561 547595 n/a n/a ACTAGTTTCCTATAACTATA 5-10-5 21 14806 14825 998 14877 14896 14936 14955 15068 15087 15211 15230 15283 15302 15342 15361 15474 15493 15546 15565 15604 15623 15676 15695 15747 15766 15806 15825 15878 15897 15936 15955 547596 n/a n/a TACTAGTTTCCTATAACTAT 5-10-5 14 14807 14826 999 14878 14897 14937 14956 15069 15088 15212 15231 15284 15303 15343 15362 15475 15494 15547 15566 15605 15624 15677 15696 15748 15767 15807 15826 15879 15898 15937 15956 547597 n/a n/a CAATATCACTGTCCTATATC 5-10-5 29 14856 14875 1000 15190 15209 15262 15281 15655 15674 15726 15745 15857 15876 547598 n/a n/a ACTATAACAATATCACTGTC 5-10-5 59 14863 14882 1001 15197 15216 15269 15288 15590 15609 15662 15681 15733 15752 15864 15883 15922 15941 16090 16109 16238 16257 547599 n/a n/a TTCCTATAACTATAACAATA 5-10-5 4 14871 14890 1002 14930 14949 15205 15224 15277 15296 15336 15355 15598 15617 15670 15689 15741 15760 15800 15819 15872 15891 15930 15949 547600 n/a n/a TTTCCTATAACTATAACAAT 5-10-5 26 14872 14891 1003 14931 14950 15206 15225 15278 15297 15337 15356 15599 15618 15671 15690 15742 15761 15801 15820
15873 15892 15931 15950 547601 n/a n/a GTACCTATAACTCTAACAGT 5-10-5 75 14905 14924 1004 15024 15043 15239 15258 15311 15330 15430 15449 15502 15521 15632 15651 15834 15853 15964 15983 547602 n/a n/a TCACTGTACCTATAACTCTA 5-10-5 93 14910 14929 1005 15029 15048 15244 15263 15316 15335 15435 15454 15507 15526 15637 15656 15839 15858 15969 15988 547603 n/a n/a TATCACTGTACCTATAACTC 5-10-5 41 14912 14931 1006 15246 15265 15318 15337 15509 15528 15639 15658 15841 15860 15971 15990 547604 n/a n/a ATATCACTGTACCTATAACT 5-10-5 0 14913 14932 1007 15247 15266 15319 15338 15510 15529 15640 15659 15783 15802 15842 15861 15972 15991 547605 n/a n/a ACAATATCACTGTACCTATA 5-10-5 43 14916 14935 1008 15322 15341 15786 15805 16137 16156 547606 n/a n/a AACAATATCACTGTACCTAT 5-10-5 43 14917 14936 1009 15323 15342 15787 15806 16138 16157 547607 n/a n/a TAACAATATCACTGTACCTA 5-10-5 49 14918 14937 1010 15324 15343 15788 15807 16139 16158 547608 n/a n/a TATAACAATATCACTGTACC 5-10-5 35 14920 14939 1011 15326 15345 15790 15809 16063 16082 16141 16160 547609 n/a n/a CTATAACAATATCACTGTAC 5-10-5 23 14921 14940 1012 15327 15346 15791 15810 16064 16083 16142 16161 547610 n/a n/a TGTAACAGTATCACTGTACT 5-10-5 45 14953 14972 1013 547611 n/a n/a CTGTAACAGTATCACTGTAC 5-10-5 71 14954 14973 1014 547612 n/a n/a CCTGTAACAGTATCACTGTA 5-10-5 68 14955 14974 1015 547613 n/a n/a CTATATCACTGTACCTGTAA 5-10-5 39 14968 14987 1016 547614 n/a n/a CCTATATCACTGTACCTGTA 5-10-5 81 14969 14988 1017 547615 n/a n/a TCCTATATCACTGTACCTGT 5-10-5 84 14970 14989 1018 547616 n/a n/a TGTCCTATATCACTGTACCT 5-10-5 86 14972 14991 1019 15253 15272 15378 15397 15516 15535 15646 15665 15848 15867 15978 15997 547617 n/a n/a CTGTCCTATATCACTGTACC 5-10-5 91 14973 14992 1020 15254 15273 15379 15398 15517 15536 15647 15666 15849 15868 15979 15998 547618 n/a n/a ACTGTCCTATATCACTGTAC 5-10-5 87 14974 14993 1021 15255 15274 15380 15399 15518 15537 15648 15667 15850 15869 15980 15999 547619 n/a n/a TCCTATAACAGTATCACTGT 5-10-5 70 14989 15008 1022 15395 15414 547620 n/a n/a TTCCTATAACAGTATCACTG 5-10-5 65 14990 15009 1023 15396 15415 547621 n/a n/a TACTAGTTTCCTATAACAGT 5-10-5 12 14997 15016 1024 15403 15422 547622 n/a n/a GTCACTGTACCTATAACTCT 5-10-5 88 15030 15049 1025 15436 15455 547623 n/a n/a TGTCACTGTACCTATAACTC 5-10-5 81 15031 15050 1026 15437 15456 547624 n/a n/a ATGTCACTGTACCTATAACT 5-10-5 64 15032 15051 1027 15438 15457
TABLE-US-00127 TABLE 136 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ ID NO Site Site Sequence inhibition Motif Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 93 5-10-5 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546885 n/a n/a TATGTCACTGTACCTATAAC 46 5-10-5 15033 15052 1028 15439 15458 546886 n/a n/a CTATGTCACTGTACCTATAA 80 5-10-5 15034 15053 1029 15440 15459 546887 n/a n/a CCTATGTCACTGTACCTATA 82 5-10-5 15035 15054 1030 15441 15460 546888 n/a n/a TCCTATGTCACTGTACCTAT 78 5-10-5 15036 15055 1031 15442 15461 546889 n/a n/a GTCCTATGTCACTGTACCTA 93 5-10-5 15037 15056 1032 15443 15462 546890 n/a n/a TGTCCTATGTCACTGTACCT 78 5-10-5 15038 15057 1033 15444 15463 546891 n/a n/a CTGTCCTATGTCACTGTACC 81 5-10-5 15039 15058 1034 15445 15464 546892 n/a n/a ACTGTCCTATGTCACTGTAC 82 5-10-5 15040 15059 1035 15446 15465 546893 n/a n/a CACTGTCCTATGTCACTGTA 70 5-10-5 15041 15060 1036 15447 15466 546894 n/a n/a TCACTGTCCTATGTCACTGT 91 5-10-5 15042 15061 1037 15448 15467 546895 n/a n/a TATCACTGTCCTATGTCACT 77 5-10-5 15044 15063 1038 15450 15469 546896 n/a n/a GTATCACTGTCCTATGTCAC 75 5-10-5 15045 15064 1039 15451 15470 546897 n/a n/a AGTATCACTGTCCTATGTCA 90 5-10-5 15046 15065 1040 15452 15471 546898 n/a n/a AACAGTATCACTGTCCTATG 91 5-10-5 15049 15068 1041 15455 15474 546899 n/a n/a CTACCTATAACTCTAACAGT 27 5-10-5 15096 15115 1042 546901 n/a n/a ACTGTCCTATAACTATAACA 56 5-10-5 15170 15189 1043 15576 15595 15706 15725 16005 16024 16076 16095 16101 16120 16154 16173 546902 n/a n/a CACTGTCCTATAACTATAAC 71 5-10-5 15171 15190 1044 15577 15596 15707 15726 16006 16025 16077 16096 16102 16121 16155 16174 546903 n/a n/a CCTATATCACTGTACCTATA 91 5-10-5 15250 15269 1045 15375 15394 15513 15532 15643 15662 15845 15864 15975 15994 546904 n/a n/a TCCTATATCACTGTACCTAT 80 5-10-5 15251 15270 1046 15376 15395 15514 15533 15644 15663 15846 15865 15976 15995 546905 n/a n/a TACCTATAACAGTATCACTG 65 5-10-5 15363 15382 1047 546907 n/a n/a ATAACTATAACAGTATCACC 37 5-10-5 15769 15788 1048 546908 n/a n/a TCACTGTACCTATAACTATA 77 5-10-5 15780 15799 1049 16252 16271 546909 n/a n/a AACAATATCACTGTACCTTT 44 5-10-5 16060 16079 1050 546910 n/a n/a TAACAATATCACTGTACCTT 82 5-10-5 16061 16080 1051 546911 n/a n/a GTCCTATAACTATAACAATA 52 5-10-5 16073 16092 1052 16098 16117 16151 16170 547625 n/a n/a CAGTATCACTGTCCTATGTC 79 5-10-5 15047 15066 1053 15453 15472 547626 n/a n/a ACAGTATCACTGTCCTATGT 91 5-10-5 15048 15067 1054 15454 15473 547627 n/a n/a TCTACCTATAACTCTAACAG 71 5-10-5 15097 15116 1055 547628 n/a n/a CTCTACCTATAACTCTAACA 34 5-10-5 15098 15117 1056 547629 n/a n/a ACTCTACCTATAACTCTAAC 0 5-10-5 15099 15118 1057 547630 n/a n/a ACTGTCCTATATCACTCTAC 76 5-10-5 15112 15131 1058 547631 n/a n/a CACTGTCCTATATCACTCTA 85 5-10-5 15113 15132 1059 547632 n/a n/a TCACTGTCCTATATCACTCT 87 5-10-5 15114 15133 1060 547633 n/a n/a ATCACTGTCCTATATCACTC 87 5-10-5 15115 15134 1061 547634 n/a n/a ATCACTGTACTAGTTTTCTA 72 5-10-5 15148 15167 1062 547635 n/a n/a TATCACTGTACTAGTTTTCT 53 5-10-5 15149 15168 1063 547636 n/a n/a GTATCACTGTACTAGTTTTC 86 5-10-5 15150 15169 1064 547637 n/a n/a AGTATCACTGTACTAGTTTT 88 5-10-5 15151 15170 1065 547638 n/a n/a ATAACAGTATCACTGTACTA 87 5-10-5 15156 15175 1066 15358 15377 15562 15581 15692 15711 15894 15913 547639 n/a n/a GTCCTATAACTATAACAGTA 72 5-10-5 15167 15186 1067 15573 15592 15703 15722 16002 16021 547640 n/a n/a TGTCCTATAACTATAACAGT 13 5-10-5 15168 15187 1068 15574 15593 15704 15723 16003 16022 547641 n/a n/a CTGTCCTATAACTATAACAG 43 5-10-5 15169 15188 1069 15575 15594 15705 15724 16004 16023 547642 n/a n/a TCACTGTCCTATAACTATAA 72 5-10-5 15172 15191 1070 15578 15597 15708 15727 16007 16026 16078 16097 16103 16122 16156 16175 547643 n/a n/a ATCACTGTCCTATAACTATA 72 5-10-5 15173 15192 1071 15579 15598 15709 15728 16008 16027 16079 16098 16104 16123 16157 16176 16176 16195 547644 n/a n/a TATCACTGTCCTATAACTAT 51 5-10-5 15174 15193 1072 15580 15599 15710 15729 16009 16028 16080 16099 16158 16177 16177 16196 16228 16247 547645 n/a n/a ATATCACTGTCCTATAACTA 60 5-10-5 15175 15194 1073 15581 15600 15711 15730 16010 16029 16081 16100 16159 16178 16178 16197 16229 16248 547646 n/a n/a CTATATCACTGTACCTATAA 23 5-10-5 15249 15268 1074 15374 15393 15512 15531 15642 15661 15844 15863 15974 15993 547647 n/a n/a GTCCTATATCACTGTACCTA 92 5-10-5 15252 15271 1075 15377 15396 15515 15534 15645 15664 15847 15866 15977 15996 547648 n/a n/a CCTATAACAGTATCACTGTA 83 5-10-5 15361 15380 1076 547649 n/a n/a ACCTATAACAGTATCACTGT 73 5-10-5 15362 15381 1077 547650 n/a n/a GTACCTATAACAGTATCACT 32 5-10-5 15364 15383 1078 547651 n/a n/a TGTACCTATAACAGTATCAC 48 5-10-5 15365 15384 1079 547652 n/a n/a TCACTGTACCTATAACAGTA 59 5-10-5 15369 15388 1080 547653 n/a n/a ATCACTGTACCTATAACAGT 57 5-10-5 15370 15389 1081 547654 n/a n/a TATCACTGTACCTATAACAG 53 5-10-5 15371 15390 1082 547655 n/a n/a AATATCACTGTCCTATAACT 37 5-10-5 15582 15601 1083 16011 16030 16082 16101 16179 16198 16230 16249 547656 n/a n/a CAATATCACTGTCCTATAAC 42 5-10-5 15583 15602 1084 16083 16102 16180 16199 16231 16250 547657 n/a n/a ACAATATCACTGTCCTATAA 43 5-10-5 15584 15603 1085 16084 16103 16181 16200 16232 16251 547658 n/a n/a CGTACTAGTTTCCTATAACT 68 5-10-5 15750 15769 1086 547659 n/a n/a ACTATAACAGTATCACCGTA 80 5-10-5 15766 15785 1087 547660 n/a n/a AACTATAACAGTATCACCGT 68 5-10-5 15767 15786 1088
547661 n/a n/a TAACTATAACAGTATCACCG 80 5-10-5 15768 15787 1089 547662 n/a n/a ACCTATAACTATAACAGTAT 0 5-10-5 15773 15792 1090 547663 n/a n/a TACCTATAACTATAACAGTA 10 5-10-5 15774 15793 1091 547664 n/a n/a GTACCTATAACTATAACAGT 2 5-10-5 15775 15794 1092 547665 n/a n/a TGTACCTATAACTATAACAG 10 5-10-5 15776 15795 1093 547666 n/a n/a ATCACTGTACCTATAACTAT 71 5-10-5 15781 15800 1094 16253 16272 547667 n/a n/a TATCACTGTACCTATAACTA 55 5-10-5 15782 15801 1095 547668 n/a n/a CAACTATAACAGTATCACTG 44 5-10-5 15899 15918 1096 547669 n/a n/a ACAACTATAACAGTATCACT 0 5-10-5 15900 15919 1097 547670 n/a n/a TACAACTATAACAGTATCAC 0 5-10-5 15901 15920 1098 547671 n/a n/a CTACAACTATAACAGTATCA 0 5-10-5 15902 15921 1099 547672 n/a n/a CAATATCACTGTCCTACAAC 36 5-10-5 15915 15934 1100 547673 n/a n/a GAATATCACTGTCCTATAAC 21 5-10-5 16012 16031 1101 547674 n/a n/a ACAATATCACTGTACCTTTA 53 5-10-5 16059 16078 1102 547675 n/a n/a TGTCCTATAACTATAACAAT 10 5-10-5 16074 16093 1103 16099 16118 16152 16171 547676 n/a n/a CTGTCCTATAACTATAACAA 41 5-10-5 16075 16094 1104 16100 16119 16153 16172
TABLE-US-00128 TABLE 137 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 93 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546529 n/a n/a GCACCTGGCAGAACAGTACC 5-10-5 65 26419 26438 1105 546578 n/a n/a GACAGTGGGCCAGAGCCTTG 5-10-5 73 26686 26705 1106 546912 n/a n/a ACATCACTGTCCTATAACTA 5-10-5 26 16106 16125 1107 546913 n/a n/a GTACCTATATCACTGTAACT 5-10-5 38 16126 16145 1108 546914 n/a n/a ATATCACTGTACCTATATCA 5-10-5 52 16134 16153 1109 546915 n/a n/a TCACTGTCCTATAACTATAT 5-10-5 39 16175 16194 1110 546916 n/a n/a CGTCACTGTACCTATAACTG 5-10-5 92 16203 16222 1111 546917 n/a n/a ATCACTGTCCTATAACTATT 5-10-5 63 16227 16246 1112 546918 n/a n/a AACATCACTGTACCTATAAC 5-10-5 14 16256 16275 1113 546926 n/a n/a GCCATCCAGGGTGCTCTCCC 5-10-5 81 16839 16858 1114 546931 n/a n/a GCCCCCGGAGCACCTTCACT 5-10-5 58 17205 17224 1115 546935 n/a n/a CGTGGTTAGCCTGACATCTC 5-10-5 86 17412 17431 1116 546939 n/a n/a GCCATCTGGTTAGCCTCCGA 5-10-5 89 17664 17683 1117 546942 n/a n/a TACACTGAACCCCCTTAGGC 5-10-5 56 18570 18589 1118 546943 n/a n/a CAGTTTGGCCTTTCCATCTC 5-10-5 54 18819 18838 1119 546944 n/a n/a GCCACTAACCCACCTCTTAA 5-10-5 42 19140 19159 1120 546946 n/a n/a ACTCCCATCTACTCCCCCAT 5-10-5 41 19291 19310 1121 546954 n/a n/a CTGCTGATTGTGTCTGGCTC 5-10-5 71 20235 20254 1122 546955 n/a n/a ACAAGGCTTCGAGGACAGCC 5-10-5 49 20339 20358 1123 546964 n/a n/a GCGATTCCTTGCCTCTGCTG 5-10-5 53 21550 21569 1124 546967 n/a n/a CACCGCGCGAATGCCTGCCT 5-10-5 93 22657 22676 1125 546969 n/a n/a ATCCAACCTCTCTCCCTATC 5-10-5 53 22901 22920 1126 546970 n/a n/a GCCCAAGCCTACATGCATAC 5-10-5 61 23426 23445 1127 546975 n/a n/a GGCCTGGATACAGCCTTTCT 5-10-5 70 23825 23844 1128 546977 n/a n/a GTCCCGAAGAGTCAAGTCCA 5-10-5 76 24253 24272 1129 546979 n/a n/a ACTGTTGTCCATAGCAGCAT 5-10-5 71 24504 24523 1130 546980 n/a n/a AGCCCTCAATTGTTGCTGGT 5-10-5 79 24664 24683 1131 546983 n/a n/a GATGACCTGCAGATGCACAG 5-10-5 74 24978 24997 1132 546986 n/a n/a CAGGATAGAACTGATGGTCC 5-10-5 91 25318 25337 1133 546990 n/a n/a AGAACAGGAGACAATCCACT 5-10-5 49 25680 25699 1134 546994 n/a n/a GTTCATGTGGCAACCTGTGA 5-10-5 58 26112 26131 1135 547677 n/a n/a CATCACTGTCCTATAACTAT 5-10-5 62 16105 16124 1136 547678 n/a n/a TACCTATATCACTGTAACTA 5-10-5 21 16125 16144 1137 547679 n/a n/a TGTACCTATATCACTGTAAC 5-10-5 28 16127 16146 1138 547680 n/a n/a TATCACTGTACCTATATCAC 5-10-5 41 16133 16152 1139 547681 n/a n/a AATATCACTGTACCTATATC 5-10-5 6 16135 16154 1140 547682 n/a n/a CAATATCACTGTACCTATAT 5-10-5 20 16136 16155 1141 547683 n/a n/a ACTATATCACTGTCCTATAA 5-10-5 33 16162 16181 1142 547684 n/a n/a TAACTATATCACTGTCCTAT 5-10-5 43 16164 16183 1143 547685 n/a n/a ATAACTATATCACTGTCCTA 5-10-5 35 16165 16184 1144 547686 n/a n/a CTGTCCTATAACTATATCAC 5-10-5 36 16172 16191 1145 547687 n/a n/a ACTGTCCTATAACTATATCA 5-10-5 41 16173 16192 1146 547688 n/a n/a CACTGTCCTATAACTATATC 5-10-5 47 16174 16193 1147 547689 n/a n/a GTAACAATATCACTGTCCTA 5-10-5 73 16184 16203 1148 547690 n/a n/a CTGTAACAATATCACTGTCC 5-10-5 76 16186 16205 1149 547691 n/a n/a ACTGTAACAATATCACTGTC 5-10-5 36 16187 16206 1150 547692 n/a n/a CACTGTACCTATAACTGTAA 5-10-5 47 16200 16219 1151 547693 n/a n/a TCACTGTACCTATAACTGTA 5-10-5 61 16201 16220 1152 547694 n/a n/a GTCACTGTACCTATAACTGT 5-10-5 92 16202 16221 1153 547695 n/a n/a ACTGTCCTATAACTATTACA 5-10-5 31 16224 16243 1154 547696 n/a n/a CACTGTCCTATAACTATTAC 5-10-5 26 16225 16244 1155 547697 n/a n/a TCACTGTCCTATAACTATTA 5-10-5 63 16226 16245 1156 547698 n/a n/a ACCTATAACTATAACAATAT 5-10-5 0 16245 16264 1157 547699 n/a n/a TACCTATAACTATAACAATA 5-10-5 10 16246 16265 1158 547700 n/a n/a GTACCTATAACTATAACAAT 5-10-5 0 16247 16266 1159 547701 n/a n/a CATCACTGTACCTATAACTA 5-10-5 49 16254 16273 1160 547702 n/a n/a ACATCACTGTACCTATAACT 5-10-5 44 16255 16274 1161 547703 n/a n/a CAACATCACTGTACCTATAA 5-10-5 25 16257 16276 1162 547704 n/a n/a ACATCTTGTCATTAACATCC 5-10-5 61 16435 16454 1163 547705 n/a n/a GCACCCAATACAGGGCCAGG 5-10-5 69 16512 16531 1164 547706 n/a n/a TGCCTCCTGGCAGCCTTCAA 5-10-5 73 16694 16713 1165 547707 n/a n/a TGAAAAGCCACGCCCTTAGC 5-10-5 32 16975 16994 1166 547708 n/a n/a GCCAGGAGACAGCCCTACTC 5-10-5 67 17055 17074 1167 547709 n/a n/a AGCCCAATGTCCTAACCTGT 5-10-5 76 17791 17810 1168 547710 n/a n/a TGCGGTTATATGGGCTGAAG 5-10-5 85 19540 19559 1169 547711 n/a n/a CCTTTAGCCACTCCTCTTGC 5-10-5 45 20061 20080 1170 547712 n/a n/a CCCCATGGTACCAAAGCCAT 5-10-5 79 20528 20547 1171 547713 n/a n/a CTCAATGCCACCCTTTCCCC 5-10-5 37 20880 20899 1172 547714 n/a n/a CTGTCTAACTGGCCTGGCTG 5-10-5 19 21326 21345 1173 547715 n/a n/a GGTCAGAAGGCCTCTTATTC 5-10-5 21 21750 21769 1174 547716 n/a n/a CCATCTGTCCCCTCAATCCC 5-10-5 9 22197 22216 1175 547717 n/a n/a ACTCTGGCACTGGTCATGGA 5-10-5 54 22761 22780 1176 547718 n/a n/a ATAAAGTGCGATTAAGCCCC 5-10-5 86 23515 23534 1177 547719 n/a n/a TACCAAGCTTGTAGAAGGGA 5-10-5 69 23633 23652 1178 547720 n/a n/a GAAAGACGGCCAATGGGAAA 5-10-5 8 24177 24196 1179 547721 n/a n/a CTCTATCAAAATCCTGCTGC 5-10-5 68 25527 25546 1180 547722 n/a n/a CTCCAGTCACCACCATTGCC 5-10-5 80 25860 25879 1181
TABLE-US-00129 TABLE 138 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Motif inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 91 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 546599 n/a n/a AAGAGTAAGCCTTCACAGGG 5-10-5 82 27583 27602 1182 546606 n/a n/a CTCACCAGAGTTGTCCCCAG 5-10-5 0 27722 27741 1183 546999 n/a n/a GCAGCTCACACCCAAAAAGC 5-10-5 29 27004 27023 1184 547000 n/a n/a TCTGTTACCTTGAGGATTGT 5-10-5 63 27276 27295 1185 547006 n/a n/a CGCCATCTGCCCTGTACAGA 5-10-5 39 28248 28267 1186 547008 n/a n/a TTGGTGGTGGGATTGGTGGT 5-10-5 81 28333 28352 1187 28388 28407 28443 28462 28608 28627 28620 28639 547009 n/a n/a AATTGGTGGTGGGATTGGTG 5-10-5 73 28335 28354 1188 547010 n/a n/a GAATTGGTGGTGGGATTGGT 5-10-5 39 28336 28355 1189 547011 n/a n/a GGCAGGATTGGTGGTGGAAT 5-10-5 22 28352 28371 1190 547013 n/a n/a TGAGATTGGTGGTGGGTGGC 5-10-5 0 28369 28388 1191 547015 n/a n/a GGTGGTGGGATTGGTGCTGA 5-10-5 55 28429 28448 1192 547016 n/a n/a GTAGGTGGTGGGATTGGTGG 5-10-5 62 28456 28475 1193 28535 28554 547017 n/a n/a GGTAGGTGGTGGGATTGGTG 5-10-5 61 28457 28476 1194 28536 28555 547018 n/a n/a GGTGGCGGGATTGGTGGTGG 5-10-5 58 28477 28496 1195 28556 28575 547019 n/a n/a GATCGGTGGTGGGATTGGTC 5-10-5 83 28500 28519 1196 28579 28598 547020 n/a n/a GGATCGGTGGTGGGATTGGT 5-10-5 47 28501 28520 1197 28580 28599 547021 n/a n/a TTGGTGGCGGGATCGGTGGT 5-10-5 57 28510 28529 1198 28589 28608 547022 n/a n/a ATTGGTGGCGGGATCGGTGG 5-10-5 69 28511 28530 1199 547023 n/a n/a GATTGGTGGCGGGATCGGTG 5-10-5 91 28512 28531 1200 547024 n/a n/a GGATTGGTGGCGGGATCGGT 5-10-5 56 28513 28532 1201 547025 n/a n/a TGGTGGTGGGATTGGTGGTT 5-10-5 72 28607 28626 1202 547029 n/a n/a TCTTCTAGGGCCACACCTCT 5-10-5 50 28891 28910 1203 547035 n/a n/a TGGTCCCAAATTGGAGTGCA 5-10-5 40 29383 29402 1204 547039 n/a n/a TCTCTATACAGCTGGGCACA 5-10-5 0 29997 30016 1205 547049 n/a n/a CACTTCCCAGCAACCCTCAC 5-10-5 20 30765 30784 1206 547055 n/a n/a GCTCCTGGCAGCAATGACCC 5-10-5 70 31104 31123 1207 547059 n/a n/a GGGTATCTTCACTGTTCCAG 5-10-5 12 31540 31559 1208 547063 n/a n/a CGTCATGCTTACCTTTCTCC 5-10-5 23 31955 31974 1209 547069 n/a n/a GCCCTCCGAGCTTTGGCAAC 5-10-5 35 32581 32600 1210 547071 n/a n/a GCAGCCCCCCAGAAATCCCA 5-10-5 27 32708 32727 1211 547076 n/a n/a TCTCAAGCAGCCTATTGTGT 5-10-5 14 33263 33282 1212 547080 n/a n/a GTGCAAGACCTTGCTTGCCA 5-10-5 54 33657 33676 1213 547081 n/a n/a CTGTAGTCCACTACACAGCA 5-10-5 83 33801 33820 1214 547082 n/a n/a TCTCCCTGAGTCACAGTGGA 5-10-5 64 33881 33900 1215 547085 n/a n/a CCAGGTGCAGCACGGAGAGG 5-10-5 44 34479 34498 1216 547723 n/a n/a TAGAATGGCAGGGTTCTGTG 5-10-5 53 27357 27376 1217 547724 n/a n/a GATGCATCCAACACTTACCC 5-10-5 16 28059 28078 1218 547725 n/a n/a ATTGGTGGTGGGATTGGTGG 5-10-5 26 28334 28353 1219 28389 28408 28444 28463 28523 28542 28609 28628 28621 28640 547726 n/a n/a GCAGGATTGGTGGTGGAATT 5-10-5 0 28351 28370 1220 547727 n/a n/a TGGCAGGATTGGTGGTGGAA 5-10-5 0 28353 28372 1221 547728 n/a n/a GAGATTGGTGGTGGGTGGCA 5-10-5 88 28368 28387 1222 547729 n/a n/a GTGAGATTGGTGGTGGGTGG 5-10-5 45 28370 28389 1223 547730 n/a n/a GATTGGTGGTGGGATTGGTG 5-10-5 60 28390 28409 1224 28433 28452 28445 28464 28524 28543 28610 28629 28622 28641 547731 n/a n/a GGATTGGTGGTGGGATTGGT 5-10-5 49 28391 28410 1225 28434 28453 28446 28465 28525 28544 28611 28630 28623 28642 547732 n/a n/a AGGATTGGTGGTGGGATTGG 5-10-5 0 28392 28411 1226 547733 n/a n/a TAGGATTGGTGGTGGGATTG 5-10-5 0 28393 28412 1227 547734 n/a n/a GTAGGATTGGTGGTGGGATT 5-10-5 14 28394 28413 1228 547735 n/a n/a GGTAGGATTGGTGGTGGGAT 5-10-5 39 28395 28414 1229 547736 n/a n/a TGGTAGGATTGGTGGTGGGA 5-10-5 54 28396 28415 1230 547737 n/a n/a TGGTGGTGGGATTGGTGCTG 5-10-5 59 28430 28449 1231 547738 n/a n/a TTGGTGGTGGGATTGGTGCT 5-10-5 41 28431 28450 1232 547739 n/a n/a ATTGGTGGTGGGATTGGTGC 5-10-5 12 28432 28451 1233 547740 n/a n/a AGGTGGTGGGATTGGTGGTG 5-10-5 30 28454 28473 1234 28533 28552 547741 n/a n/a TAGGTGGTGGGATTGGTGGT 5-10-5 47 28455 28474 1235 28534 28553 547742 n/a n/a ATCGGTGGTGGGATTGGTCG 5-10-5 57 28499 28518 1236 28578 28597 547743 n/a n/a GGTGGTGGGATTGGTGGCGG 5-10-5 61 28520 28539 1237 547744 n/a n/a TGGTGGTGGGATTGGTGGCG 5-10-5 65 28521 28540 1238 547745 n/a n/a TTGGTGGTGGGATTGGTGGC 5-10-5 55 28522 28541 1239 547746 n/a n/a GTTGGTGGCGGGATCGGTGG 5-10-5 0 28590 28609 1240 547748 n/a n/a GGTTGGTGGCGGGATCGGTG 5-10-5 78 28591 28610 1241 547750 n/a n/a TGGTTGGTGGCGGGATCGGT 5-10-5 41 28592 28611 1242 547752 n/a n/a GTGGTTGGTGGCGGGATCGG 5-10-5 41 28593 28612 1243 547754 n/a n/a GGGATTGGTGGTTGGTGGCG 5-10-5 47 28600 28619 1244 547756 n/a n/a GGGTCTTGCTCCACCCACAT 5-10-5 49 29244 29263 1245 547758 n/a n/a CCAAGTAGTGCAAGGCATGT 5-10-5 24 29540 29559 1246 547760 n/a n/a ATCATGCTTACTGCAAGTGA 5-10-5 19 30219 30238 1247 547762 n/a n/a TGAAACTGGGCAGTCCTTCC 5-10-5 0 30417 30436 1248 547764 n/a n/a CCACCTTCTTACATATGCTA 5-10-5 24 30644 30663 1249 547766 n/a n/a GCCTCTCAGACGGCACAGAC 5-10-5 0 30902 30921 1250 547768 n/a n/a TTGCCCTCACACATTCGAAT 5-10-5 0 30977 30996 1251 547770 n/a n/a TGCTTTCTGCCCAACCTCTA 5-10-5 48 31727 31746 1252 547772 n/a n/a CTGTGCTCCCGGCCATTAGC 5-10-5 0 32312 32331 1253 547774 n/a n/a GAGACAGTTTGGCAAGCTAC 5-10-5 46 32389 32408 1254 547776 n/a n/a GGAGAGAGACGGCACCCTGT 5-10-5 48 32828 32847 1255 547778 n/a n/a TCACCTGTGAGTAACCAATA 5-10-5 53 33085 33104 1256 547780 n/a n/a CCCCTCTTAAATAGCACATG 5-10-5 67 33441 33460 1257 547782 n/a n/a CCAAGTATCTCATGTGCCTG 5-10-5 67 33580 33599 1258
TABLE-US-00130 TABLE 139 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Motif inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT 5-10-5 90 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 548706 n/a n/a CTAGTTTCCTATAACT 3-10-3 0 14738 14753 1259 14809 14824 14880 14895 14939 14954 15071 15086 15214 15229 15286 15301 15345 15360 15477 15492 15549 15564 15607 15622 15679 15694 15750 15765 15809 15824 15881 15896 15939 15954 548707 n/a n/a ACTAGTTTCCTATAAC 3-10-3 10 14739 14754 1260 14810 14825 14881 14896 14940 14955 15000 15015 15072 15087 15215 15230 15287 15302 15346 15361 15406 15421 15478 15493 15550 15565 15608 15623 15680 15695 15751 15766 15810 15825 15882 15897 15940 15955 548708 n/a n/a TACTAGTTTCCTATAA 3-10-3 0 14740 14755 1261 14811 14826 14882 14897 14941 14956 15001 15016 15073 15088 15216 15231 15288 15303 15347 15362 15407 15422 15479 15494 15551 15566 15609 15624 15681 15696 15752 15767 15811 15826 15883 15898 15941 15956 548709 n/a n/a GTACTAGTTTCCTATA 3-10-3 0 14741 14756 1262 14812 14827 14883 14898 14942 14957 15002 15017 15074 15089 15217 15232 15289 15304 15348 15363 15408 15423 15480 15495 15552 15567 15610 15625 15682 15697 15753 15768 15812 15827 15884 15899 15942 15957 548710 n/a n/a TGTACTAGTTTCCTAT 3-10-3 0 14742 14757 1263 14813 14828 14884 14899 14943 14958 15003 15018 15075 15090 15218 15233 15290 15305 15349 15364 15409 15424 15481 15496 15553 15568 15611 15626 15683 15698 15813 15828 15885 15900 15943 15958 548711 n/a n/a CTGTACTAGTTTCCTA 3-10-3 21 14743 14758 1264 14814 14829 14885 14900 14944 14959 15004 15019 15076 15091 15219 15234 15291 15306 15350 15365 15410 15425 15482 15497 15554 15569 15612 15627 15684 15699 15814 15829 15886 15901 15944 15959 548712 n/a n/a ACTGTACTAGTTTCCT 3-10-3 9 14744 14759 1265 14815 14830 14886 14901 14945 14960 15005 15020 15077 15092 15220 15235 15292 15307 15351 15366 15411 15426 15483 15498 15555 15570 15613 15628 15685 15700 15815 15830 15887 15902 15945 15960 548713 n/a n/a CACTGTACTAGTTTCC 3-10-3 33 14745 14760 1266 14816 14831 14887 14902 14946 14961 15006 15021 15078 15093 15221 15236 15293 15308 15352 15367 15412 15427 15484 15499 15556 15571 15614 15629 15686 15701 15816 15831 15888 15903 15946 15961 548714 n/a n/a TCACTGTACTAGTTTC 3-10-3 15 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548715 n/a n/a ATCACTGTACTAGTTT 3-10-3 0 14747 14762 1268 14818 14833 14889 14904 14948 14963 15008 15023 15080 15095 15152 15167 15223 15238 15295 15310 15354 15369 15414 15429 15486 15501 15558 15573 15616 15631 15688 15703 15818 15833 15890 15905 15948 15963 548716 n/a n/a TATCACTGTACTAGTT 3-10-3 10 14748 14763 1269 14819 14834 14890 14905 14949 14964 15009 15024 15081 15096 15153 15168 15224 15239 15296 15311 15355 15370 15415 15430 15487 15502 15559 15574 15617 15632 15689 15704 15819 15834 15891 15906 15949 15964 548717 n/a n/a ACTAGTTTCCTATAACT 3-10-4 0 14738 14754 1270 14809 14825 14880 14896 14939 14955 15071 15087 15214 15230 15286 15302 15345 15361 15477 15493 15549 15565 15607 15623 15679 15695 15750 15766 15809 15825 15881 15897 15939 15955 548718 n/a n/a TACTAGTTTCCTATAAC 3-10-4 0 14739 14755 1271 14810 14826 14881 14897
14940 14956 15000 15016 15072 15088 15215 15231 15287 15303 15346 15362 15406 15422 15478 15494 15550 15566 15608 15624 15680 15696 15751 15767 15810 15826 15882 15898 15940 15956 548719 n/a n/a GTACTAGTTTCCTATAA 3-10-4 0 14740 14756 1272 14811 14827 14882 14898 14941 14957 15001 15017 15073 15089 15216 15232 15288 15304 15347 15363 15407 15423 15479 15495 15551 15567 15609 15625 15681 15697 15752 15768 15811 15827 15883 15899 15941 15957 548720 n/a n/a TGTACTAGTTTCCTATA 3-10-4 0 14741 14757 1273 14812 14828 14883 14899 14942 14958 15002 15018 15074 15090 15217 15233 15289 15305 15348 15364 15408 15424 15480 15496 15552 15568 15610 15626 15682 15698 15812 15828 15884 15900 15942 15958 548721 n/a n/a CTGTACTAGTTTCCTAT 3-10-4 27 14742 14758 1274 14813 14829 14884 14900 14943 14959 15003 15019 15075 15091 15218 15234 15290 15306 15349 15365 15409 15425 15481 15497 15553 15569 15611 15627 15683 15699 15813 15829 15885 15901 15943 15959 548722 n/a n/a ACTGTACTAGTTTCCTA 3-10-4 26 14743 14759 1275 14814 14830 14885 14901 14944 14960 15004 15020 15076 15092 15219 15235 15291 15307 15350 15366 15410 15426 15482 15498 15554 15570 15612 15628 15684 15700 15814 15830 15886 15902 15944 15960 548723 n/a n/a CACTGTACTAGTTTCCT 3-10-4 62 14744 14760 1276 14815 14831 14886 14902 14945 14961 15005 15021 15077 15093 15220 15236 15292 15308 15351 15367 15411 15427 15483 15499 15555 15571 15613 15629 15685 15701 15815 15831 15887 15903 15945 15961 548724 n/a n/a TCACTGTACTAGTTTCC 3-10-4 61 14745 14761 1277 14816 14832 14887 14903 14946 14962 15006 15022 15078 15094 15221 15237 15293 15309 15352 15368 15412 15428 15484 15500 15556 15572 15614 15630 15686 15702 15816 15832 15888 15904 15946 15962 548725 n/a n/a ATCACTGTACTAGTTTC 3-10-4 32 14746 14762 1278 14817 14833 14888 14904 14947 14963 15007 15023 15079 15095 15222 15238 15294 15310 15353 15369 15413 15429 15485 15501 15557 15573 15615 15631 15687 15703 15817 15833 15889 15905 15947 15963 548726 n/a n/a TATCACTGTACTAGTTT 3-10-4 21 14747 14763 1279 14818 14834 14889 14905 14948 14964 15008 15024 15080 15096 15152 15168 15223 15239 15295 15311 15354 15370 15414 15430 15486 15502 15558 15574 15616 15632 15688 15704 15818 15834 15890 15906 15948 15964 548727 n/a n/a ACTAGTTTCCTATAACT 4-10-3 0 14738 14754 1270 14809 14825 14880 14896 14939 14955 15071 15087 15214 15230 15286 15302 15345 15361 15477 15493 15549 15565 15607 15623 15679 15695 15750 15766 15809 15825 15881 15897 15939 15955 548728 n/a n/a TACTAGTTTCCTATAAC 4-10-3 0 14739 14755 1271 14810 14826 14881 14897 14940 14956 15000 15016 15072 15088 15215 15231 15287 15303 15346 15362 15406 15422 15478 15494 15550 15566 15608 15624 15680 15696 15751 15767 15810 15826 15882 15898 15940 15956 548729 n/a n/a GTACTAGTTTCCTATAA 4-10-3 13 14740 14756 1272 14811 14827 14882 14898 14941 14957 15001 15017 15073 15089 15216 15232 15288 15304 15347 15363 15407 15423 15479 15495 15551 15567 15609 15625 15681 15697 15752 15768 15811 15827 15883 15899 15941 15957 548730 n/a n/a TGTACTAGTTTCCTATA 4-10-3 0 14741 14757 1273 14812 14828 14883 14899 14942 14958 15002 15018 15074 15090 15217 15233 15289 15305 15348 15364 15408 15424 15480 15496 15552 15568 15610 15626 15682 15698 15812 15828 15884 15900 15942 15958 548731 n/a n/a CTGTACTAGTTTCCTAT 4-10-3 49 14742 14758 1274 14813 14829 14884 14900 14943 14959 15003 15019 15075 15091 15218 15234 15290 15306 15349 15365 15409 15425 15481 15497 15553 15569 15611 15627 15683 15699 15813 15829 15885 15901
15943 15959 548732 n/a n/a ACTGTACTAGTTTCCTA 4-10-3 36 14743 14759 1275 14814 14830 14885 14901 14944 14960 15004 15020 15076 15092 15219 15235 15291 15307 15350 15366 15410 15426 15482 15498 15554 15570 15612 15628 15684 15700 15814 15830 15886 15902 15944 15960 548733 n/a n/a CACTGTACTAGTTTCCT 4-10-3 84 14744 14760 1276 14815 14831 14886 14902 14945 14961 15005 15021 15077 15093 15220 15236 15292 15308 15351 15367 15411 15427 15483 15499 15555 15571 15613 15629 15685 15701 15815 15831 15887 15903 15945 15961 548734 n/a n/a TCACTGTACTAGTTTCC 4-10-3 51 14745 14761 1277 14816 14832 14887 14903 14946 14962 15006 15022 15078 15094 15221 15237 15293 15309 15352 15368 15412 15428 15484 15500 15556 15572 15614 15630 15686 15702 15816 15832 15888 15904 15946 15962 548735 n/a n/a ATCACTGTACTAGTTTC 4-10-3 48 14746 14762 1278 14817 14833 14888 14904 14947 14963 15007 15023 15079 15095 15222 15238 15294 15310 15353 15369 15413 15429 15485 15501 15557 15573 15615 15631 15687 15703 15817 15833 15889 15905 15947 15963 548736 n/a n/a TATCACTGTACTAGTTT 4-10-3 21 14747 14763 1279 14818 14834 14889 14905 14948 14964 15008 15024 15080 15096 15152 15168 15223 15239 15295 15311 15354 15370 15414 15430 15486 15502 15558 15574 15616 15632 15688 15704 15818 15834 15890 15906 15948 15964 548737 n/a n/a ACTAGTTTCCTATAACT 4-9-4 11 14738 14754 1270 14809 14825 14880 14896 14939 14955 15071 15087 15214 15230 15286 15302 15345 15361 15477 15493 15549 15565 15607 15623 15679 15695 15750 15766 15809 15825 15881 15897 15939 15955 548738 n/a n/a TACTAGTTTCCTATAAC 4-9-4 0 14739 14755 1271 14810 14826 14881 14897 14940 14956 15000 15016 15072 15088 15215 15231 15287 15303 15346 15362 15406 15422 15478 15494 15550 15566 15608 15624 15680 15696 15751 15767 15810 15826 15882 15898 15940 15956 548739 n/a n/a GTACTAGTTTCCTATAA 4-9-4 0 14740 14756 1272 14811 14827 14882 14898 14941 14957 15001 15017 15073 15089 15216 15232 15288 15304 15347 15363 15407 15423 15479 15495 15551 15567 15609 15625 15681 15697 15752 15768 15811 15827 15883 15899 15941 15957 548740 n/a n/a TGTACTAGTTTCCTATA 4-9-4 0 14741 14757 1273 14812 14828 14883 14899 14942 14958 15002 15018 15074 15090 15217 15233 15289 15305 15348 15364 15408 15424 15480 15496 15552 15568 15610 15626 15682 15698 15812 15828 15884 15900 15942 15958 548741 n/a n/a CTGTACTAGTTTCCTAT 4-9-4 69 14742 14758 1274 14813 14829 14884 14900 14943 14959 15003 15019 15075 15091 15218 15234 15290 15306 15349 15365 15409 15425 15481 15497 15553 15569 15611 15627 15683 15699 15813 15829 15885 15901 15943 15959 548742 n/a n/a ACTGTACTAGTTTCCTA 4-9-4 50 14743 14759 1275 14814 14830 14885 14901 14944 14960 15004 15020 15076 15092 15219 15235 15291 15307 15350 15366 15410 15426 15482 15498 15554 15570 15612 15628 15684 15700 15814 15830 15886 15902 15944 15960 548743 n/a n/a CACTGTACTAGTTTCCT 4-9-4 80 14744 14760 1276 14815 14831 14886 14902 14945 14961 15005 15021 15077 15093 15220 15236 15292 15308 15351 15367 15411 15427 15483 15499 15555 15571 15613 15629 15685 15701 15815 15831 15887 15903 15945 15961 548744 n/a n/a TCACTGTACTAGTTTCC 4-9-4 83 14745 14761 1277 14816 14832 14887 14903 14946 14962 15006 15022 15078 15094 15221 15237 15293 15309 15352 15368 15412 15428 15484 15500 15556 15572 15614 15630 15686 15702 15816 15832 15888 15904 15946 15962 548745 n/a n/a ATCACTGTACTAGTTTC 4-9-4 71 14746 14762 1278 14817 14833 14888 14904 14947 14963 15007 15023 15079 15095 15222 15238 15294 15310 15353 15369 15413 15429 15485 15501 15557 15573 15615 15631
15687 15703 15817 15833 15889 15905 15947 15963 548746 n/a n/a TATCACTGTACTAGTTT 4-9-4 40 14747 14763 1279 14818 14834 14889 14905 14948 14964 15008 15024 15080 15096 15152 15168 15223 15239 15295 15311 15354 15370 15414 15430 15486 15502 15558 15574 15616 15632 15688 15704 15818 15834 15890 15906 15948 15964 548747 n/a n/a TACTAGTTTCCTATAACT 4-10-4 2 14738 14755 1280 14809 14826 14880 14897 14939 14956 15071 15088 15214 15231 15286 15303 15345 15362 15477 15494 15549 15566 15607 15624 15679 15696 15750 15767 15809 15826 15881 15898 15939 15956 548748 n/a n/a GTACTAGTTTCCTATAAC 4-10-4 0 14739 14756 1281 14810 14827 14881 14898 14940 14957 15000 15017 15072 15089 15215 15232 15287 15304 15346 15363 15406 15423 15478 15495 15550 15567 15608 15625 15680 15697 15751 15768 15810 15827 15882 15899 15940 15957 548749 n/a n/a TGTACTAGTTTCCTATAA 4-10-4 0 14740 14757 1282 14811 14828 14882 14899 14941 14958 15001 15018 15073 15090 15216 15233 15288 15305 15347 15364 15407 15424 15479 15496 15551 15568 15609 15626 15681 15698 15811 15828 15883 15900 15941 15958 548750 n/a n/a CTGTACTAGTTTCCTATA 4-10-4 62 14741 14758 1283 14812 14829 14883 14900 14942 14959 15002 15019 15074 15091 15217 15234 15289 15306 15348 15365 15408 15425 15480 15497 15552 15569 15610 15627 15682 15699 15812 15829 15884 15901 15942 15959 548751 n/a n/a ACTGTACTAGTTTCCTAT 4-10-4 53 14742 14759 1284 14813 14830 14884 14901 14943 14960 15003 15020 15075 15092 15218 15235 15290 15307 15349 15366 15409 15426 15481 15498 15553 15570 15611 15628 15683 15700 15813 15830 15885 15902 15943 15960 548752 n/a n/a CACTGTACTAGTTTCCTA 4-10-4 89 14743 14760 1285 14814 14831 14885 14902 14944 14961 15004 15021 15076 15093 15219 15236 15291 15308 15350 15367 15410 15427 15482 15499 15554 15571 15612 15629 15684 15701 15814 15831 15886 15903 15944 15961 548753 n/a n/a TCACTGTACTAGTTTCCT 4-10-4 82 14744 14761 1286 14815 14832 14886 14903 14945 14962 15005 15022 15077 15094 15220 15237 15292 15309 15351 15368 15411 15428 15483 15500 15555 15572 15613 15630 15685 15702 15815 15832 15887 15904 15945 15962 548754 n/a n/a ATCACTGTACTAGTTTCC 4-10-4 77 14745 14762 1287 14816 14833 14887 14904 14946 14963 15006 15023 15078 15095 15221 15238 15293 15310 15352 15369 15412 15429 15484 15501 15556 15573 15614 15631 15686 15703 15816 15833 15888 15905 15946 15963 548755 n/a n/a TATCACTGTACTAGTTTC 4-10-4 20 14746 14763 1288 14817 14834 14888 14905 14947 14964 15007 15024 15079 15096 15222 15239 15294 15311 15353 15370 15413 15430 15485 15502 15557 15574 15615 15632 15687 15704 15817 15834 15889 15906 15947 15964 548756 n/a n/a GTATCACTGTACTAGTT 4-9-4 81 14748 14764 1289 14819 14835 14890 14906 14949 14965 15009 15025 15081 15097 15153 15169 15224 15240 15296 15312 15355 15371 15415 15431 15487 15503 15559 15575 15617 15633 15689 15705 15819 15835 15891 15907 15949 15965 548757 n/a n/a AGTATCACTGTACTAGT 4-9-4 87 14749 14765 1290 14820 14836 14891 14907 14950 14966 15010 15026 15082 15098 15154 15170 15225 15241 15297 15313 15356 15372 15416 15432 15488 15504 15560 15576 15618 15634 15690 15706 15820 15836 15892 15908 15950 15966 548758 n/a n/a CAGTATCACTGTACTAG 4-9-4 97 14750 14766 1291 14821 14837 14892 14908 14951 14967 15011 15027 15083 15099 15155 15171 15226 15242 15298 15314 15357 15373 15417 15433 15489 15505 15561 15577 15619 15635 15691 15707 15821 15837 15893 15909 15951 15967 548759 n/a n/a AACAGTATCACTGTACT 4-9-4 68 14752 14768 1292 14823 14839 14894 14910 14953 14969 15013 15029 15085 15101 15157 15173 15228 15244
15300 15316 15359 15375 15419 15435 15491 15507 15563 15579 15621 15637 15693 15709 15823 15839 15895 15911 15953 15969 548760 n/a n/a TAACAGTATCACTGTAC 4-9-4 53 14753 14769 1293 14824 14840 14895 14911 14954 14970 15014 15030 15086 15102 15158 15174 15229 15245 15301 15317 15360 15376 15420 15436 15492 15508 15564 15580 15622 15638 15694 15710 15824 15840 15896 15912 15954 15970 548761 n/a n/a CTAACAGTATCACTGTA 4-9-4 49 14754 14770 1294 14825 14841 14896 14912 15015 15031 15087 15103 15230 15246 15302 15318 15421 15437 15493 15509 15623 15639 15825 15841 15955 15971 548762 n/a n/a TCTAACAGTATCACTGT 4-9-4 16 14755 14771 1295 14826 14842 14897 14913 15016 15032 15088 15104 15231 15247 15303 15319 15422 15438 15494 15510 15624 15640 15826 15842 15956 15972 548763 n/a n/a CTCTAACAGTATCACTG 4-9-4 44 14756 14772 1296 14827 14843 14898 14914 15017 15033 15089 15105 15232 15248 15304 15320 15423 15439 15495 15511 15625 15641 15827 15843 15957 15973 548764 n/a n/a TATCACTGTCCTATAAC 4-9-4 31 14772 14788 1297 14843 14859 15177 15193 15583 15599 15713 15729 16012 16028 16083 16099 16161 16177 16180 16196 16231 16247 548765 n/a n/a ATATCACTGTCCTATAA 4-9-4 0 14773 14789 1298 14844 14860 15178 15194 15584 15600 15714 15730 16013 16029 16084 16100 16162 16178 16181 16197 16232 16248 548766 n/a n/a TATATCACTGTCCTATA 4-9-4 36 14774 14790 1299 14845 14861 15179 15195 15715 15731 16163 16179 548767 n/a n/a TATCACTGTCCTATATC 4-9-4 59 14785 14801 1300 14856 14872 14981 14997 15119 15135 15190 15206 15262 15278 15387 15403 15525 15541 15655 15671 15726 15742 15857 15873 15987 16003 548768 n/a n/a GTATCACTGTCCTATAT 4-9-4 56 14786 14802 1301 14982 14998 15120 15136 15388 15404 15526 15542 15988 16004 548769 n/a n/a AGTATCACTGTCCTATA 4-9-4 64 14787 14803 1302 14983 14999 15121 15137 15389 15405 15527 15543 15989 16005 548770 n/a n/a TAACAGTATCACTGTCC 4-9-4 92 14791 14807 1303 14987 15003 15053 15069 15125 15141 15393 15409 15459 15475 15531 15547 15993 16009 548771 n/a n/a ATAACAGTATCACTGTC 4-9-4 62 14792 14808 1304 14988 15004 15054 15070 15126 15142 15394 15410 15460 15476 15532 15548 15994 16010 548772 n/a n/a TATAACAGTATCACTGT 4-9-4 0 14793 14809 1305 14989 15005 15055 15071 15127 15143 15160 15176 15362 15378 15395 15411 15461 15477 15533 15549 15566 15582 15696 15712 15898 15914 15995 16011 548773 n/a n/a CTATAACAGTATCACTG 4-9-4 0 14794 14810 1306 14990 15006 15056 15072 15128 15144 15161 15177 15363 15379 15396 15412 15462 15478 15534 15550 15567 15583 15697 15713 15899 15915 15996 16012 548774 n/a n/a CCTATAACTATAACAGT 4-9-4 0 14801 14817 1307 15063 15079 15168 15184 15469 15485 15541 15557 15574 15590 15704 15720 15775 15791 16003 16019 548775 n/a n/a TCCTATAACTATAACAG 4-9-4 0 14802 14818 1308 15064 15080 15169 15185 15470 15486 15542 15558 15575 15591 15705 15721 16004 16020 548776 n/a n/a CCTATAACTATAACAAT 4-9-4 0 14872 14888 1309 14931 14947 15206 15222 15278 15294 15337 15353 15599 15615 15671 15687 15742 15758 15801 15817 15873 15889 15931 15947 16074 16090 16099 16115 16152 16168 16247 16263 548777 n/a n/a GTAACAGTATCACTGTA 4-9-4 41 14955 14971 1310 548778 n/a n/a ATAACAGTATCACTGTA 4-9-4 20 15159 15175 1311 15361 15377 15565 15581 15695 15711 15897 15913 548779 n/a n/a GTCCTATAACTATAACA 4-9-4 0 15170 15186 1312 15576 15592 15706 15722 16005 16021 16076 16092 16101 16117 16154 16170 548780 n/a n/a TGTCCTATAACTATAAC 4-9-4 22 15171 15187 1313 15577 15593 15707 15723 16006 16022 16077 16093 16102 16118 16155 16171 548781 n/a n/a ACCTATAACTATAACAG 4-9-4 0 15776 15792 1314 548782 n/a n/a TACCTATAACTATAACA 4-9-4 0 15777 15793 1315 16249 16265 548783 n/a n/a ACCTATAACTATAACAA 4-9-4 0 16248 16264 1316
Example 116: Antisense Inhibition of Human PKK in HepaRG.TM. Cells by Antisense Oligonucleotides with MOE, Deoxy and cEt Sugar Modifications
[1067] Additional antisense oligonucleotides were designed targeting a PKK nucleic acid and were tested for their effects on PKK mRNA in vitro.
[1068] The chimeric antisense oligonucleotides in the tables below were designed as deoxy, MOE and cEt gapmers. The gapmers are 16 nucleosides in length wherein the nucleoside have either a MOE sugar modification, a cEt sugar modification, or a deoxy modification. The `Chemistry` column describes the sugar modifications of each oligonucleotide. `k` indicates an cEt sugar modification; the number indicates the number of deoxynucleosides; otherwise, `d` indicates a deoxynucleoside; and `e` indicates a 2'-O-methoxyethyl modification. The internucleoside linkages throughout each gapmer are phosphorothioate linkages. All cytosine residues throughout each oligonucleotide are 5-methylcytosines. "Start site" indicates the 5'-most nucleoside to which the gapmer is targeted in the human gene sequence. "Stop site" indicates the 3'-most nucleoside to which the gapmer is targeted in the human gene sequence. Each gapmer listed in the tables below is targeted to either the human PKK mRNA, designated herein as SEQ ID NO: 1 or the human PKK genomic sequence, designated herein as SEQ ID NO: 10. `n/a` indicates that the antisense oligonucleotide does not target that particular gene sequence.
[1069] Cultured HepaRG.TM. cells at a density of 20,000 cells per well were transfected using electroporation with 1,000 nM antisense oligonucleotide. After a treatment period of approximately 24 hours, RNA was isolated from the cells and PKK mRNA levels were measured by quantitative real-time PCR. Human primer probe set RTS3454 was used to measure mRNA levels. ISIS 531231 was also included in this assay. PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.RTM.. The antisense oligonucleotides were tested in a series of experiments that had similar culture conditions. The results for each experiment are presented in separate tables shown below. Results are presented as percent inhibition of PKK, relative to untreated control cells.
TABLE-US-00131 TABLE 140 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Chemistry inhibition Site Site NO 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 95 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548074 1642 1657 CCTTTCTCCTTCGAGA eekd.sub.10kke 0 31948 31963 1317 548075 1643 1658 ACCTTTCTCCTTCGAG eekd.sub.10kke 0 31949 31964 1318 548076 1644 1659 CACCTTTCTCCTTCGA eekd.sub.10kke 26 n/a n/a 1319 548077 1691 1706 ATTTGTTACCAAAGGA eekd.sub.10kke 51 33135 33150 1320 548078 1696 1711 TCTTCATTTGTTACCA eekd.sub.10kke 36 33140 33155 1321 548079 1762 1777 CCTTCTTTATAGCCAG eekd.sub.10kke 39 33206 33221 1322 548080 1763 1778 CCCTTCTTTATAGCCA eekd.sub.10kke 0 33207 33222 1323 548081 1764 1779 CCCCTTCTTTATAGCC eekd.sub.10kke 64 33208 33223 1324 548082 1776 1791 AAGCATCTTTTCCCCC eekd.sub.10kke 42 33220 33235 1325 548083 1800 1815 AGGGACCACCTGAATC eekd.sub.10kke 0 33899 33914 1326 548084 1801 1816 AAGGGACCACCTGAAT eekd.sub.10kke 0 33900 33915 1327 548085 1802 1817 TAAGGGACCACCTGAA eekd.sub.10kke 8 33901 33916 1328 548086 1803 1818 CTAAGGGACCACCTGA eekd.sub.10kke 36 33902 33917 1329 548087 1804 1819 ACTAAGGGACCACCTG eekd.sub.10kke 24 33903 33918 1330 548088 1805 1820 AACTAAGGGACCACCT eekd.sub.10kke 27 33904 33919 1331 548089 1806 1821 AAACTAAGGGACCACC eekd.sub.10kke 34 33905 33920 1332 548090 1807 1822 CAAACTAAGGGACCAC eekd.sub.10kke 46 33906 33921 1333 548091 1809 1824 TGCAAACTAAGGGACC eekd.sub.10kke 62 33908 33923 1334 548092 1810 1825 TTGCAAACTAAGGGAC eekd.sub.10kke 30 33909 33924 1335 548093 1811 1826 TTTGCAAACTAAGGGA eekd.sub.10kke 0 33910 33925 1336 548094 1812 1827 GTTTGCAAACTAAGGG eekd.sub.10kke 74 33911 33926 1337 548095 1813 1828 TGTTTGCAAACTAAGG eekd.sub.10kke 35 33912 33927 1338 548096 1814 1829 GTGTTTGCAAACTAAG eekd.sub.10kke 23 33913 33928 1339 548097 1876 1891 TGCTCCCTGCGGGCAC eekd.sub.10kke 2 33975 33990 1340 548098 1887 1902 AGACACCAGGTTGCTC eekd.sub.10kke 0 33986 34001 1341 548099 1904 1919 CTCAGCGACTTTGGTG eekd.sub.10kke 55 34003 34018 1342 548100 1905 1920 ACTCAGCGACTTTGGT eekd.sub.10kke 25 34004 34019 1343 548101 1906 1921 TACTCAGCGACTTTGG eekd.sub.10kke 47 34005 34020 1344 548102 1907 1922 GTACTCAGCGACTTTG eekd.sub.10kke 58 34006 34021 1345 548103 1908 1923 TGTACTCAGCGACTTT eekd.sub.10kke 66 34007 34022 1346 548104 1909 1924 ATGTACTCAGCGACTT eekd.sub.10kke 59 34008 34023 1347 548105 1910 1925 CATGTACTCAGCGACT eekd.sub.10kke 49 34009 34024 1348 548106 1911 1926 CCATGTACTCAGCGAC eekd.sub.10kke 79 34010 34025 1349 548107 1912 1927 TCCATGTACTCAGCGA eekd.sub.10kke 76 34011 34026 1350 548108 1953 1968 GAGCTTTTCCATCACT eekd.sub.10kke 61 34052 34067 1351 548109 1959 1974 GCATCTGAGCTTTTCC eekd.sub.10kke 77 34058 34073 1352 548110 1960 1975 TGCATCTGAGCTTTTC eekd.sub.10kke 62 34059 34074 1353 548111 1963 1978 GACTGCATCTGAGCTT eekd.sub.10kke 53 34062 34077 1354 548112 1965 1980 GTGACTGCATCTGAGC eekd.sub.10kke 23 34064 34079 1355 548113 1966 1981 GGTGACTGCATCTGAG eekd.sub.10kke 56 34065 34080 1356 548114 1967 1982 TGGTGACTGCATCTGA eekd.sub.10kke 70 34066 34081 1357 548115 1972 1987 CATGCTGGTGACTGCA eekd.sub.10kke 76 34071 34086 1358 548116 1973 1988 TCATGCTGGTGACTGC eekd.sub.10kke 3 34072 34087 1359 548117 1974 1989 CTCATGCTGGTGACTG eekd.sub.10kke 73 34073 34088 1360 548118 1975 1990 TCTCATGCTGGTGACT eekd.sub.10kke 47 34074 34089 1361 548119 1984 1999 TGGACTGCTTCTCATG eekd.sub.10kke 25 34083 34098 1362 548121 1986 2001 TCTGGACTGCTTCTCA eekd.sub.10kke 64 34085 34100 1363 548122 1987 2002 CTCTGGACTGCTTCTC eekd.sub.10kke 55 34086 34101 1364 548123 1990 2005 AGACTCTGGACTGCTT eekd.sub.10kke 49 34089 34104 1365 548124 1991 2006 TAGACTCTGGACTGCT eekd.sub.10kke 51 34090 34105 1366 548125 1992 2007 CTAGACTCTGGACTGC eekd.sub.10kke 89 34091 34106 1367 548126 1995 2010 TGCCTAGACTCTGGAC eekd.sub.10kke 19 34094 34109 1368 548127 1996 2011 TTGCCTAGACTCTGGA eekd.sub.10kke 60 34095 34110 1369 548128 1997 2012 ATTGCCTAGACTCTGG eekd.sub.10kke 55 34096 34111 1370 548129 2022 2037 TTTGACTTGAACTCAG eekd.sub.10kke 35 34121 34136 1371 548130 2023 2038 ATTTGACTTGAACTCA eekd.sub.10kke 27 34122 34137 1372 548131 2024 2039 AATTTGACTTGAACTC eekd.sub.10kke 45 34123 34138 1373 548132 2025 2040 GAATTTGACTTGAACT eekd.sub.10kke 0 34124 34139 1374 548133 2026 2041 AGAATTTGACTTGAAC eekd.sub.10kke 23 34125 34140 1375 548134 2027 2042 CAGAATTTGACTTGAA eekd.sub.10kke 17 34126 34141 1376 548135 2028 2043 TCAGAATTTGACTTGA eekd.sub.10kke 46 34127 34142 1377 548136 2031 2046 GGCTCAGAATTTGACT eekd.sub.10kke 39 34130 34145 1378 548137 2032 2047 AGGCTCAGAATTTGAC eekd.sub.10kke 62 34131 34146 1379 548138 2036 2051 CCCCAGGCTCAGAATT eekd.sub.10kke 52 34135 34150 1380 548139 2047 2062 AGATGAGGACCCCCCA eekd.sub.10kke 56 34146 34161 1381 548140 2048 2063 CAGATGAGGACCCCCC eekd.sub.10kke 74 34147 34162 1382 548141 2049 2064 GCAGATGAGGACCCCC eekd.sub.10kke 66 34148 34163 1383 548142 2063 2078 ACTCTCCATGCTTTGC eekd.sub.10kke 44 34162 34177 1384 548143 2064 2079 CACTCTCCATGCTTTG eekd.sub.10kke 39 34163 34178 1385 548144 2068 2083 ATGCCACTCTCCATGC eekd.sub.10kke 52 34167 34182 1386 548145 2079 2094 ATGCAAAGAAGATGCC eekd.sub.10kke 63 34178 34193 1387 548146 2088 2103 GTCCTTAGGATGCAAA eekd.sub.10kke 68 34187 34202 1388 548147 2089 2104 CGTCCTTAGGATGCAA eekd.sub.10kke 81 34188 34203 1389 548148 2114 2129 GCAGCTCTGAGTGCAC eekd.sub.10kke 66 34213 34228 1390 548149 2127 2142 GACATTGTCCTCAGCA eekd.sub.10kke 39 34226 34241 1391 548150 2129 2144 CAGACATTGTCCTCAG eekd.sub.10kke 60 34228 34243 1392
TABLE-US-00132 TABLE 141 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Chemistry inhibition Site Site NO 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 84 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 547843 384 399 CACTTATTTGATGACC eekd.sub.10kke 83 9918 9933 1393 547844 385 400 GCACTTATTTGATGAC eekd.sub.10kke 13 n/a n/a 1394 547845 394 409 CGATGGCAAGCACTTA eekd.sub.10kke 0 n/a n/a 1395 547846 395 410 TCGATGGCAAGCACTT eekd.sub.10kke 0 n/a n/a 1396 547847 396 411 CTCGATGGCAAGCACT eekd.sub.10kke 46 n/a n/a 1397 547848 400 415 ATGTCTCGATGGCAAG eekd.sub.10kke 93 12656 12671 1398 547849 401 416 AATGTCTCGATGGCAA eekd.sub.10kke 79 12657 12672 1399 547850 402 417 AAATGTCTCGATGGCA eekd.sub.10kke 51 12658 12673 1400 547851 403 418 TAAATGTCTCGATGGC eekd.sub.10kke 93 12659 12674 1401 547852 404 419 ATAAATGTCTCGATGG eekd.sub.10kke 67 12660 12675 1402 547853 405 420 TATAAATGTCTCGATG eekd.sub.10kke 0 12661 12676 1403 547854 416 431 ATCAACTCCTTTATAA eekd.sub.10kke 10 12672 12687 1404 547855 417 432 TATCAACTCCTTTATA eekd.sub.10kke 59 12673 12688 1405 547856 419 434 CATATCAACTCCTTTA eekd.sub.10kke 93 12675 12690 1406 547858 423 438 CTCTCATATCAACTCC eekd.sub.10kke 82 12679 12694 1407 547859 424 439 CCTCTCATATCAACTC eekd.sub.10kke 77 12680 12695 1408 547860 425 440 TCCTCTCATATCAACT eekd.sub.10kke 71 12681 12696 1409 547861 427 442 ACTCCTCTCATATCAA eekd.sub.10kke 0 12683 12698 1410 547862 428 443 GACTCCTCTCATATCA eekd.sub.10kke 22 12684 12699 1411 547863 429 444 TGACTCCTCTCATATC eekd.sub.10kke 73 12685 12700 1412 547864 430 445 TTGACTCCTCTCATAT eekd.sub.10kke 53 12686 12701 1413 547865 434 449 AAAATTGACTCCTCTC eekd.sub.10kke 3 12690 12705 1414 547866 436 451 TTAAAATTGACTCCTC eekd.sub.10kke 46 12692 12707 1415 547867 447 462 CCTTAGACACATTAAA eekd.sub.10kke 34 12703 12718 1416 547868 448 463 ACCTTAGACACATTAA eekd.sub.10kke 47 12704 12719 1417 547869 449 464 AACCTTAGACACATTA eekd.sub.10kke 45 12705 12720 1418 547870 451 466 CTAACCTTAGACACAT eekd.sub.10kke 89 12707 12722 1419 547871 452 467 GCTAACCTTAGACACA eekd.sub.10kke 96 12708 12723 1420 547872 453 468 TGCTAACCTTAGACAC eekd.sub.10kke 85 12709 12724 1421 547873 454 469 CTGCTAACCTTAGACA eekd.sub.10kke 77 12710 12725 1422 547874 455 470 ACTGCTAACCTTAGAC eekd.sub.10kke 70 12711 12726 1423 547875 456 471 CACTGCTAACCTTAGA eekd.sub.10kke 73 12712 12727 1424 547876 457 472 ACACTGCTAACCTTAG eekd.sub.10kke 78 12713 12728 1425 547877 458 473 AACACTGCTAACCTTA eekd.sub.10kke 81 12714 12729 1426 547879 460 475 TCAACACTGCTAACCT eekd.sub.10kke 69 12716 12731 1427 547880 461 476 TTCAACACTGCTAACC eekd.sub.10kke 69 12717 12732 1428 547881 465 480 ATTCTTCAACACTGCT eekd.sub.10kke 0 12721 12736 1429 547882 500 515 CTGGCAGCGAATGTTA eekd.sub.10kke 91 12756 12771 1430 547883 501 516 ACTGGCAGCGAATGTT eekd.sub.10kke 99 12757 12772 1431 547884 518 533 CGTGGCATATGAAAAA eekd.sub.10kke 87 12774 12789 1432 547885 539 554 CTCTGCCTTGTGAAAT eekd.sub.10kke 45 12795 12810 1433 547886 544 559 CGGTACTCTGCCTTGT eekd.sub.10kke 97 12800 12815 1434 547889 547 562 TTCCGGTACTCTGCCT eekd.sub.10kke 91 n/a n/a 1435 547890 550 565 TTGTTCCGGTACTCTG eekd.sub.10kke 97 n/a n/a 1436 547891 551 566 ATTGTTCCGGTACTCT eekd.sub.10kke 84 n/a n/a 1437 547892 553 568 CAATTGTTCCGGTACT eekd.sub.10kke 29 n/a n/a 1438 547893 554 569 GCAATTGTTCCGGTAC eekd.sub.10kke 81 n/a n/a 1439 547894 555 570 GGCAATTGTTCCGGTA eekd.sub.10kke 92 n/a n/a 1440 547898 563 578 CTTTAATAGGCAATTG eekd.sub.10kke 0 14134 14149 1441 547899 566 581 GTACTTTAATAGGCAA eekd.sub.10kke 49 14137 14152 1442 547900 567 582 TGTACTTTAATAGGCA eekd.sub.10kke 93 14138 14153 1443 547901 568 583 CTGTACTTTAATAGGC eekd.sub.10kke 77 14139 14154 1444 547902 569 584 ACTGTACTTTAATAGG eekd.sub.10kke 20 14140 14155 1445 547903 604 619 CTCAGCACCTTTATAG eekd.sub.10kke 62 14175 14190 1446 547904 605 620 ACTCAGCACCTTTATA eekd.sub.10kke 56 14176 14191 1447 547905 606 621 TACTCAGCACCTTTAT eekd.sub.10kke 20 14177 14192 1448 547906 607 622 TTACTCAGCACCTTTA eekd.sub.10kke 59 14178 14193 1449 547907 652 667 ATTTCTGAAAGGGCAC eekd.sub.10kke 27 14223 14238 1450 547908 654 669 CAATTTCTGAAAGGGC eekd.sub.10kke 94 14225 14240 1451 547909 655 670 CCAATTTCTGAAAGGG eekd.sub.10kke 82 14226 14241 1452 547910 656 671 ACCAATTTCTGAAAGG eekd.sub.10kke 26 14227 14242 1453 547911 661 676 TGGCAACCAATTTCTG eekd.sub.10kke 0 n/a n/a 1454 547912 701 716 ATCCACATCTGAGAAC eekd.sub.10kke 23 26149 26164 1455 547913 706 721 GCAACATCCACATCTG eekd.sub.10kke 71 26154 26169 1456 547914 707 722 GGCAACATCCACATCT eekd.sub.10kke 74 26155 26170 1457 547915 708 723 TGGCAACATCCACATC eekd.sub.10kke 0 26156 26171 1458 547916 710 725 CCTGGCAACATCCACA eekd.sub.10kke 70 26158 26173 1459 547917 712 727 ACCCTGGCAACATCCA eekd.sub.10kke 33 26160 26175 1460 547918 713 728 AACCCTGGCAACATCC eekd.sub.10kke 1 26161 26176 1461 547919 714 729 GAACCCTGGCAACATC eekd.sub.10kke 41 26162 26177 1462
TABLE-US-00133 TABLE 142 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 62 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 88 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 547751 7 22 TGAACGGTCTTCAAGC eekd.sub.10kke 0 3399 3414 1463 547753 8 23 ATGAACGGTCTTCAAG eekd.sub.10kke 3 3400 3415 1464 547755 13 28 TAAAAATGAACGGTCT eekd.sub.10kke 0 3405 3420 1465 547757 28 43 GAGTCTCTTGTCACTT eekd.sub.10kke 69 3420 3435 1466 547759 29 44 TGAGTCTCTTGTCACT eekd.sub.10kke 73 3421 3436 1467 547763 31 46 GGTGAGTCTCTTGTCA eekd.sub.10kke 66 3423 3438 1468 547765 32 47 AGGTGAGTCTCTTGTC eekd.sub.10kke 20 3424 3439 1469 547767 35 50 TGGAGGTGAGTCTCTT eekd.sub.10kke 74 3427 3442 1470 547769 36 51 TTGGAGGTGAGTCTCT eekd.sub.10kke 81 3428 3443 1471 547771 37 52 CTTGGAGGTGAGTCTC eekd.sub.10kke 60 3429 3444 1472 547773 38 53 TCTTGGAGGTGAGTCT eekd.sub.10kke 47 3430 3445 1473 547777 43 58 TTGCTTCTTGGAGGTG eekd.sub.10kke 69 3435 3450 1474 547779 44 59 ATTGCTTCTTGGAGGT eekd.sub.10kke 41 3436 3451 1475 547781 46 61 CAATTGCTTCTTGGAG eekd.sub.10kke 49 3438 3453 1476 547783 48 63 CACAATTGCTTCTTGG eekd.sub.10kke 48 3440 3455 1477 547784 72 87 GCTTGAATAAAATCAT eekd.sub.10kke 46 4071 4086 1478 547785 79 94 GTTGCTTGCTTGAATA eekd.sub.10kke 48 4078 4093 1479 547786 80 95 AGTTGCTTGCTTGAAT eekd.sub.10kke 44 4079 4094 1480 547787 81 96 AAGTTGCTTGCTTGAA eekd.sub.10kke 22 4080 4095 1481 547788 82 97 TAAGTTGCTTGCTTGA eekd.sub.10kke 49 4081 4096 1482 547789 86 101 GAAATAAGTTGCTTGC eekd.sub.10kke 20 4085 4100 1483 547790 87 102 TGAAATAAGTTGCTTG eekd.sub.10kke 23 4086 4101 1484 547791 106 121 ACTGTAGCAAACAAGG eekd.sub.10kke 49 4105 4120 1485 547792 116 131 TCCACAGGAAACTGTA eekd.sub.10kke 31 n/a n/a 1486 547793 117 132 ATCCACAGGAAACTGT eekd.sub.10kke 16 n/a n/a 1487 547794 136 151 TCATAGAGTTGAGTCA eekd.sub.10kke 49 8008 8023 1488 547795 155 170 ACCTCTGAAGAAGGCG eekd.sub.10kke 66 8027 8042 1489 547796 161 176 ATCCCCACCTCTGAAG eekd.sub.10kke 35 8033 8048 1490 547797 167 182 AGCTACATCCCCACCT eekd.sub.10kke 33 8039 8054 1491 547799 169 184 GAAGCTACATCCCCAC eekd.sub.10kke 41 8041 8056 1492 547800 174 189 ACATGGAAGCTACATC eekd.sub.10kke 20 8046 8061 1493 547801 175 190 TACATGGAAGCTACAT eekd.sub.10kke 11 8047 8062 1494 547802 176 191 GTACATGGAAGCTACA eekd.sub.10kke 41 8048 8063 1495 547803 177 192 TGTACATGGAAGCTAC eekd.sub.10kke 0 8049 8064 1496 547804 178 193 GTGTACATGGAAGCTA eekd.sub.10kke 22 8050 8065 1497 547805 180 195 GGGTGTACATGGAAGC eekd.sub.10kke 54 8052 8067 1498 547807 197 212 GCAGTATTGGGCATTT eekd.sub.10kke 75 8069 8084 1499 547808 203 218 CATCTGGCAGTATTGG eekd.sub.10kke 56 8075 8090 1500 547809 204 219 TCATCTGGCAGTATTG eekd.sub.10kke 33 8076 8091 1501 547810 206 221 CCTCATCTGGCAGTAT eekd.sub.10kke 60 8078 8093 1502 547811 207 222 ACCTCATCTGGCAGTA eekd.sub.10kke 49 8079 8094 1503 547812 211 226 GTGCACCTCATCTGGC eekd.sub.10kke 51 8083 8098 1504 547813 219 234 GGTGGAATGTGCACCT eekd.sub.10kke 34 8091 8106 1505 547814 220 235 GGGTGGAATGTGCACC eekd.sub.10kke 60 8092 8107 1506 547815 255 270 AACTTGCTGGAAGAAA eekd.sub.10kke 3 8127 8142 1507 547816 256 271 GAACTTGCTGGAAGAA eekd.sub.10kke 45 8128 8143 1508 547817 257 272 TGAACTTGCTGGAAGA eekd.sub.10kke 18 8129 8144 1509 547818 260 275 GATTGAACTTGCTGGA eekd.sub.10kke 4 8132 8147 1510 547819 264 279 CATTGATTGAACTTGC eekd.sub.10kke 11 8136 8151 1511 547820 265 280 TCATTGATTGAACTTG eekd.sub.10kke 0 8137 8152 1512 547821 282 297 CAAACCTTTTCTCCAT eekd.sub.10kke 44 n/a n/a 1513 547822 287 302 GCAACCAAACCTTTTC eekd.sub.10kke 71 n/a n/a 1514 547823 288 303 AGCAACCAAACCTTTT eekd.sub.10kke 51 n/a n/a 1515 547824 331 346 CGATGTACTTTTGGCA eekd.sub.10kke 82 9865 9880 1516 547825 332 347 TCGATGTACTTTTGGC eekd.sub.10kke 59 9866 9881 1517 547826 333 348 TTCGATGTACTTTTGG eekd.sub.10kke 31 9867 9882 1518 547827 334 349 GTTCGATGTACTTTTG eekd.sub.10kke 47 9868 9883 1519 547828 337 352 CCTGTTCGATGTACTT eekd.sub.10kke 63 9871 9886 1520 547829 338 353 ACCTGTTCGATGTACT eekd.sub.10kke 59 9872 9887 1521 547830 340 355 GCACCTGTTCGATGTA eekd.sub.10kke 74 9874 9889 1522 547831 342 357 CTGCACCTGTTCGATG eekd.sub.10kke 49 9876 9891 1523 547832 343 358 ACTGCACCTGTTCGAT eekd.sub.10kke 59 9877 9892 1524 547833 344 359 AACTGCACCTGTTCGA eekd.sub.10kke 40 9878 9893 1525 547834 345 360 AAACTGCACCTGTTCG eekd.sub.10kke 63 9879 9894 1526 547835 349 364 CCAGAAACTGCACCTG eekd.sub.10kke 81 9883 9898 1527 547836 350 365 TCCAGAAACTGCACCT eekd.sub.10kke 50 9884 9899 1528 547837 352 367 TGTCCAGAAACTGCAC eekd.sub.10kke 51 9886 9901 1529 547838 362 377 CTTCAAGGAATGTCCA eekd.sub.10kke 45 9896 9911 1530 547839 363 378 GCTTCAAGGAATGTCC eekd.sub.10kke 35 9897 9912 1531 547840 365 380 TTGCTTCAAGGAATGT eekd.sub.10kke 36 9899 9914 1532 547841 369 384 CACATTGCTTCAAGGA eekd.sub.10kke 42 9903 9918 1533 547842 375 390 GATGACCACATTGCTT eekd.sub.10kke 10 9909 9924 1534
TABLE-US-00134 TABLE 143 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ ID NO Site Site Sequence Chemistry inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 75 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 91 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 547843 384 399 CACTTATTTGATGACC eekd.sub.10kke 83 9918 9933 1393 547844 385 400 GCACTTATTTGATGAC eekd.sub.10kke 76 n/a n/a 1394 547845 394 409 CGATGGCAAGCACTTA eekd.sub.10kke 64 n/a n/a 1395 547846 395 410 TCGATGGCAAGCACTT eekd.sub.10kke 42 n/a n/a 1396 547847 396 411 CTCGATGGCAAGCACT eekd.sub.10kke 72 n/a n/a 1397 547848 400 415 ATGTCTCGATGGCAAG eekd.sub.10kke 79 12656 12671 1398 547849 401 416 AATGTCTCGATGGCAA eekd.sub.10kke 90 12657 12672 1399 547850 402 417 AAATGTCTCGATGGCA eekd.sub.10kke 80 12658 12673 1400 547851 403 418 TAAATGTCTCGATGGC eekd.sub.10kke 84 12659 12674 1401 547852 404 419 ATAAATGTCTCGATGG eekd.sub.10kke 66 12660 12675 1402 547853 405 420 TATAAATGTCTCGATG eekd.sub.10kke 30 12661 12676 1403 547854 416 431 ATCAACTCCTTTATAA eekd.sub.10kke 9 12672 12687 1404 547855 417 432 TATCAACTCCTTTATA eekd.sub.10kke 38 12673 12688 1405 547856 419 434 CATATCAACTCCTTTA eekd.sub.10kke 51 12675 12690 1406 547857 421 436 CTCATATCAACTCCTT eekd.sub.10kke 84 12677 12692 1535 547858 423 438 CTCTCATATCAACTCC eekd.sub.10kke 76 12679 12694 1407 547859 424 439 CCTCTCATATCAACTC eekd.sub.10kke 88 12680 12695 1408 547860 425 440 TCCTCTCATATCAACT eekd.sub.10kke 70 12681 12696 1409 547861 427 442 ACTCCTCTCATATCAA eekd.sub.10kke 57 12683 12698 1410 547862 428 443 GACTCCTCTCATATCA eekd.sub.10kke 88 12684 12699 1411 547863 429 444 TGACTCCTCTCATATC eekd.sub.10kke 77 12685 12700 1412 547864 430 445 TTGACTCCTCTCATAT eekd.sub.10kke 73 12686 12701 1413 547865 434 449 AAAATTGACTCCTCTC eekd.sub.10kke 61 12690 12705 1414 547866 436 451 TTAAAATTGACTCCTC eekd.sub.10kke 40 12692 12707 1415 547867 447 462 CCTTAGACACATTAAA eekd.sub.10kke 53 12703 12718 1416 547868 448 463 ACCTTAGACACATTAA eekd.sub.10kke 71 12704 12719 1417 547869 449 464 AACCTTAGACACATTA eekd.sub.10kke 77 12705 12720 1418 547870 451 466 CTAACCTTAGACACAT eekd.sub.10kke 83 12707 12722 1419 547871 452 467 GCTAACCTTAGACACA eekd.sub.10kke 77 12708 12723 1420 547872 453 468 TGCTAACCTTAGACAC eekd.sub.10kke 73 12709 12724 1421 547873 454 469 CTGCTAACCTTAGACA eekd.sub.10kke 82 12710 12725 1422 547874 455 470 ACTGCTAACCTTAGAC eekd.sub.10kke 60 12711 12726 1423 547875 456 471 CACTGCTAACCTTAGA eekd.sub.10kke 57 12712 12727 1424 547876 457 472 ACACTGCTAACCTTAG eekd.sub.10kke 59 12713 12728 1425 547877 458 473 AACACTGCTAACCTTA eekd.sub.10kke 93 12714 12729 1426 547878 459 474 CAACACTGCTAACCTT eekd.sub.10kke 62 12715 12730 1536 547879 460 475 TCAACACTGCTAACCT eekd.sub.10kke 65 12716 12731 1427 547880 461 476 TTCAACACTGCTAACC eekd.sub.10kke 59 12717 12732 1428 547881 465 480 ATTCTTCAACACTGCT eekd.sub.10kke 50 12721 12736 1429 547882 500 515 CTGGCAGCGAATGTTA eekd.sub.10kke 96 12756 12771 1430 547883 501 516 ACTGGCAGCGAATGTT eekd.sub.10kke 0 12757 12772 1431 547884 518 533 CGTGGCATATGAAAAA eekd.sub.10kke 49 12774 12789 1432 547885 539 554 CTCTGCCTTGTGAAAT eekd.sub.10kke 57 12795 12810 1433 547886 544 559 CGGTACTCTGCCTTGT eekd.sub.10kke 89 12800 12815 1434 547887 545 560 CCGGTACTCTGCCTTG eekd.sub.10kke 99 12801 12816 1537 547888 546 561 TCCGGTACTCTGCCTT eekd.sub.10kke 99 n/a n/a 1538 547889 547 562 TTCCGGTACTCTGCCT eekd.sub.10kke 97 n/a n/a 1435 547890 550 565 TTGTTCCGGTACTCTG eekd.sub.10kke 90 n/a n/a 1436 547891 551 566 ATTGTTCCGGTACTCT eekd.sub.10kke 88 n/a n/a 1437 547892 553 568 CAATTGTTCCGGTACT eekd.sub.10kke 28 n/a n/a 1438 547893 554 569 GCAATTGTTCCGGTAC eekd.sub.10kke 80 n/a n/a 1439 547894 555 570 GGCAATTGTTCCGGTA eekd.sub.10kke 91 n/a n/a 1440 547895 556 571 AGGCAATTGTTCCGGT eekd.sub.10kke 94 n/a n/a 1539 547896 557 572 TAGGCAATTGTTCCGG eekd.sub.10kke 95 n/a n/a 1540 547897 558 573 ATAGGCAATTGTTCCG eekd.sub.10kke 82 n/a n/a 1541 547898 563 578 CTTTAATAGGCAATTG eekd.sub.10kke 28 14134 14149 1441 547899 566 581 GTACTTTAATAGGCAA eekd.sub.10kke 68 14137 14152 1442 547900 567 582 TGTACTTTAATAGGCA eekd.sub.10kke 68 14138 14153 1443 547901 568 583 CTGTACTTTAATAGGC eekd.sub.10kke 85 14139 14154 1444 547902 569 584 ACTGTACTTTAATAGG eekd.sub.10kke 33 14140 14155 1445 547903 604 619 CTCAGCACCTTTATAG eekd.sub.10kke 6 14175 14190 1446 547904 605 620 ACTCAGCACCTTTATA eekd.sub.10kke 41 14176 14191 1447 547905 606 621 TACTCAGCACCTTTAT eekd.sub.10kke 59 14177 14192 1448 547906 607 622 TTACTCAGCACCTTTA eekd.sub.10kke 70 14178 14193 1449 547907 652 667 ATTTCTGAAAGGGCAC eekd.sub.10kke 27 14223 14238 1450 547908 654 669 CAATTTCTGAAAGGGC eekd.sub.10kke 71 14225 14240 1451 547909 655 670 CCAATTTCTGAAAGGG eekd.sub.10kke 51 14226 14241 1452 547910 656 671 ACCAATTTCTGAAAGG eekd.sub.10kke 34 14227 14242 1453 547911 661 676 TGGCAACCAATTTCTG eekd.sub.10kke 15 n/a n/a 1454 547912 701 716 ATCCACATCTGAGAAC eekd.sub.10kke 53 26149 26164 1455 547913 706 721 GCAACATCCACATCTG eekd.sub.10kke 61 26154 26169 1456 547914 707 722 GGCAACATCCACATCT eekd.sub.10kke 63 26155 26170 1457 547915 708 723 TGGCAACATCCACATC eekd.sub.10kke 62 26156 26171 1458 547916 710 725 CCTGGCAACATCCACA eekd.sub.10kke 56 26158 26173 1459 547917 712 727 ACCCTGGCAACATCCA eekd.sub.10kke 54 26160 26175 1460 547918 713 728 AACCCTGGCAACATCC eekd.sub.10kke 65 26161 26176 1461 547919 714 729 GAACCCTGGCAACATC eekd.sub.10kke 73 26162 26177 1462
TABLE-US-00135 TABLE 144 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 16 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 83 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 547920 716 731 GAGAACCCTGGCAACA eekd.sub.10kke 52 26164 26179 1542 547921 717 732 TGAGAACCCTGGCAAC eekd.sub.10kke 43 26165 26180 1543 547922 722 737 TGGAGTGAGAACCCTG eekd.sub.10kke 79 26170 26185 1544 547923 725 740 ATCTGGAGTGAGAACC eekd.sub.10kke 68 26173 26188 1545 547924 742 757 GTCCGACACACAAAAG eekd.sub.10kke 53 26190 26205 1546 547925 743 758 GGTCCGACACACAAAA eekd.sub.10kke 16 26191 26206 1547 547927 745 760 ATGGTCCGACACACAA eekd.sub.10kke 79 26193 26208 1548 547928 746 761 GATGGTCCGACACACA eekd.sub.10kke 70 26194 26209 1549 547929 747 762 AGATGGTCCGACACAC eekd.sub.10kke 65 26195 26210 1550 547930 757 772 TGATAGGTGCAGATGG eekd.sub.10kke 48 26205 26220 1551 547931 758 773 GTGATAGGTGCAGATG eekd.sub.10kke 58 26206 26221 1552 547932 804 819 CGATTTTCCATACATT eekd.sub.10kke 33 26252 26267 1553 547933 805 820 TCGATTTTCCATACAT eekd.sub.10kke 44 26253 26268 1554 547934 806 821 CTCGATTTTCCATACA eekd.sub.10kke 38 26254 26269 1555 547935 807 822 ACTCGATTTTCCATAC eekd.sub.10kke 27 26255 26270 1556 547936 808 823 GACTCGATTTTCCATA eekd.sub.10kke 44 26256 26271 1557 547937 811 826 TGTGACTCGATTTTCC eekd.sub.10kke 56 26259 26274 1558 547938 812 827 TTGTGACTCGATTTTC eekd.sub.10kke 56 26260 26275 1559 547939 813 828 TTTGTGACTCGATTTT eekd.sub.10kke 70 26261 26276 1560 547940 817 832 TTTCTTTGTGACTCGA eekd.sub.10kke 71 n/a n/a 1561 547941 852 867 GTGTGCCACTTTCAGA eekd.sub.10kke 66 27116 27131 1562 547942 853 868 GGTGTGCCACTTTCAG eekd.sub.10kke 85 27117 27132 1563 547943 854 869 TGGTGTGCCACTTTCA eekd.sub.10kke 83 27118 27133 1564 547944 857 872 ACTTGGTGTGCCACTT eekd.sub.10kke 54 27121 27136 1565 547945 858 873 AACTTGGTGTGCCACT eekd.sub.10kke 62 27122 27137 1566 547946 859 874 GAACTTGGTGTGCCAC eekd.sub.10kke 81 27123 27138 1567 547947 860 875 GGAACTTGGTGTGCCA eekd.sub.10kke 80 27124 27139 1568 547948 861 876 AGGAACTTGGTGTGCC eekd.sub.10kke 77 27125 27140 1569 547949 880 895 GTGTTTTCTTGAGGAG eekd.sub.10kke 6 27144 27159 1570 547950 881 896 GGTGTTTTCTTGAGGA eekd.sub.10kke 49 27145 27160 1571 547951 887 902 AGATATGGTGTTTTCT eekd.sub.10kke 25 27151 27166 1572 547952 888 903 CAGATATGGTGTTTTC eekd.sub.10kke 46 27152 27167 1573 547953 895 910 CTATATCCAGATATGG eekd.sub.10kke 16 27159 27174 1574 547954 902 917 TAAAAGGCTATATCCA eekd.sub.10kke 36 27166 27181 1575 547956 904 919 GTTAAAAGGCTATATC eekd.sub.10kke 13 27168 27183 1576 547957 905 920 GGTTAAAAGGCTATAT eekd.sub.10kke 6 27169 27184 1577 547958 907 922 CAGGTTAAAAGGCTAT eekd.sub.10kke 57 27171 27186 1578 547959 908 923 GCAGGTTAAAAGGCTA eekd.sub.10kke 60 27172 27187 1579 547960 909 924 TGCAGGTTAAAAGGCT eekd.sub.10kke 40 27173 27188 1580 547961 910 925 TTGCAGGTTAAAAGGC eekd.sub.10kke 5 27174 27189 1581 547962 911 926 TTTGCAGGTTAAAAGG eekd.sub.10kke 16 27175 27190 1582 547963 927 942 GTTCAGGTAAAGTTCT eekd.sub.10kke 22 n/a n/a 1583 547964 928 943 GGTTCAGGTAAAGTTC eekd.sub.10kke 0 n/a n/a 1584 547965 929 944 GGGTTCAGGTAAAGTT eekd.sub.10kke 29 n/a n/a 1585 547966 930 945 AGGGTTCAGGTAAAGT eekd.sub.10kke 13 n/a n/a 1586 547967 933 948 GGCAGGGTTCAGGTAA eekd.sub.10kke 25 n/a n/a 1587 547968 940 955 TTAGAATGGCAGGGTT eekd.sub.10kke 37 27362 27377 1588 547969 953 968 TCCCGGGTAAATTTTA eekd.sub.10kke 0 27375 27390 1589 547970 954 969 CTCCCGGGTAAATTTT eekd.sub.10kke 42 27376 27391 1590 547972 958 973 TCAACTCCCGGGTAAA eekd.sub.10kke 49 27380 27395 1591 547973 961 976 AAGTCAACTCCCGGGT eekd.sub.10kke 62 27383 27398 1592 547974 962 977 AAAGTCAACTCCCGGG eekd.sub.10kke 52 27384 27399 1593 547975 963 978 CAAAGTCAACTCCCGG eekd.sub.10kke 44 27385 27400 1594 547976 964 979 CCAAAGTCAACTCCCG eekd.sub.10kke 49 27386 27401 1595 547977 967 982 CCTCCAAAGTCAACTC eekd.sub.10kke 57 27389 27404 1596 547978 1014 1029 CTTGGCAAACATTCAC eekd.sub.10kke 71 27436 27451 1597 547979 1018 1033 GTCTCTTGGCAAACAT eekd.sub.10kke 77 27440 27455 1598 547980 1020 1035 AAGTCTCTTGGCAAAC eekd.sub.10kke 54 27442 27457 1599 547981 1029 1044 TCTTTGTGCAAGTCTC eekd.sub.10kke 76 27451 27466 1600 547982 1034 1049 AATCATCTTTGTGCAA eekd.sub.10kke 54 27456 27471 1601 547983 1035 1050 GAATCATCTTTGTGCA eekd.sub.10kke 56 27457 27472 1602 547984 1036 1051 CGAATCATCTTTGTGC eekd.sub.10kke 55 27458 27473 1603 547985 1037 1052 GCGAATCATCTTTGTG eekd.sub.10kke 63 27459 27474 1604 547986 1039 1054 CAGCGAATCATCTTTG eekd.sub.10kke 63 27461 27476 1605 547987 1040 1055 ACAGCGAATCATCTTT eekd.sub.10kke 64 27462 27477 1606 547988 1042 1057 TGACAGCGAATCATCT eekd.sub.10kke 56 27464 27479 1607 547989 1043 1058 CTGACAGCGAATCATC eekd.sub.10kke 66 27465 27480 1608 547990 1044 1059 ACTGACAGCGAATCAT eekd.sub.10kke 58 27466 27481 1609 547991 1077 1092 TACAGTCTTCTGGGAG eekd.sub.10kke 0 27499 27514 1610 547992 1080 1095 CCTTACAGTCTTCTGG eekd.sub.10kke 17 27502 27517 1611 547993 1113 1128 TAGATAATCTTAAGAA eekd.sub.10kke 26 27634 27649 1612 547994 1120 1135 CCATCCATAGATAATC eekd.sub.10kke 53 27641 27656 1613 547995 1149 1164 GTGTCCCATACGCAAT eekd.sub.10kke 64 27670 27685 1614 547996 1150 1165 TGTGTCCCATACGCAA eekd.sub.10kke 65 27671 27686 1615
TABLE-US-00136 TABLE 145 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Chemistry inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 0 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 80 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 547997 1151 1166 TTGTGTCCCATACGCA eekd.sub.10kke 89 27672 27687 1616 547998 1152 1167 CTTGTGTCCCATACGC eekd.sub.10kke 82 27673 27688 1617 547999 1153 1168 CCTTGTGTCCCATACG eekd.sub.10kke 50 27674 27689 1618 548000 1154 1169 CCCTTGTGTCCCATAC eekd.sub.10kke 54 27675 27690 1619 548001 1163 1178 ACCAGAGCTCCCTTGT eekd.sub.10kke 64 27684 27699 1620 548002 1164 1179 AACCAGAGCTCCCTTG eekd.sub.10kke 56 27685 27700 1621 548003 1165 1180 TAACCAGAGCTCCCTT eekd.sub.10kke 66 27686 27701 1622 548004 1167 1182 AGTAACCAGAGCTCCC eekd.sub.10kke 80 27688 27703 1623 548005 1169 1184 AGAGTAACCAGAGCTC eekd.sub.10kke 77 27690 27705 1624 548006 1172 1187 CAAAGAGTAACCAGAG eekd.sub.10kke 54 27693 27708 1625 548007 1174 1189 CTCAAAGAGTAACCAG eekd.sub.10kke 70 27695 27710 1626 548008 1175 1190 TCTCAAAGAGTAACCA eekd.sub.10kke 71 27696 27711 1627 548009 1184 1199 GTTACACAATCTCAAA eekd.sub.10kke 47 27705 27720 1628 548010 1187 1202 AGTGTTACACAATCTC eekd.sub.10kke 80 27708 27723 1629 548011 1189 1204 CCAGTGTTACACAATC eekd.sub.10kke 14 27710 27725 1630 548012 1192 1207 TCCCCAGTGTTACACA eekd.sub.10kke 3 27713 27728 1631 548013 1193 1208 GTCCCCAGTGTTACAC eekd.sub.10kke 37 27714 27729 1632 548014 1194 1209 TGTCCCCAGTGTTACA eekd.sub.10kke 31 27715 27730 1633 548015 1195 1210 TTGTCCCCAGTGTTAC eekd.sub.10kke 50 27716 27731 1634 548016 1248 1263 AAGAGTTTGTTCCTCC eekd.sub.10kke 55 27924 27939 1635 548017 1252 1267 CAAGAAGAGTTTGTTC eekd.sub.10kke 3 27928 27943 1636 548018 1253 1268 CCAAGAAGAGTTTGTT eekd.sub.10kke 22 27929 27944 1637 548019 1255 1270 CCCCAAGAAGAGTTTG eekd.sub.10kke 24 27931 27946 1638 548020 1256 1271 TCCCCAAGAAGAGTTT eekd.sub.10kke 76 27932 27947 1639 548021 1261 1276 CACTCTCCCCAAGAAG eekd.sub.10kke 0 27937 27952 1640 548022 1262 1277 CCACTCTCCCCAAGAA eekd.sub.10kke 69 27938 27953 1641 548023 1290 1305 GCTTCACCTGCAGGCT eekd.sub.10kke 58 27966 27981 1642 548024 1297 1312 GCTGTCAGCTTCACCT eekd.sub.10kke 79 27973 27988 1643 548025 1300 1315 TGAGCTGTCAGCTTCA eekd.sub.10kke 66 27976 27991 1644 548026 1332 1347 GTCCTATGAGTGACCC eekd.sub.10kke 52 28008 28023 1645 548027 1334 1349 GTGTCCTATGAGTGAC eekd.sub.10kke 18 28010 28025 1646 548028 1335 1350 GGTGTCCTATGAGTGA eekd.sub.10kke 38 28011 28026 1647 548029 1336 1351 TGGTGTCCTATGAGTG eekd.sub.10kke 12 28012 28027 1648 548030 1337 1352 CTGGTGTCCTATGAGT eekd.sub.10kke 52 28013 28028 1649 548031 1397 1412 GATGCGCCAAACATCC eekd.sub.10kke 73 30475 30490 1650 548032 1398 1413 AGATGCGCCAAACATC eekd.sub.10kke 51 30476 30491 1651 548034 1400 1415 ATAGATGCGCCAAACA eekd.sub.10kke 31 30478 30493 1652 548035 1404 1419 CACTATAGATGCGCCA eekd.sub.10kke 44 30482 30497 1653 548036 1405 1420 CCACTATAGATGCGCC eekd.sub.10kke 74 30483 30498 1654 548037 1427 1442 AATGTCTGACAGATTT eekd.sub.10kke 70 30505 30520 1655 548038 1428 1443 TAATGTCTGACAGATT eekd.sub.10kke 67 30506 30521 1656 548039 1445 1460 GAAAGGTGTATCTTTT eekd.sub.10kke 29 30523 30538 1657 548040 1449 1464 GTGAGAAAGGTGTATC eekd.sub.10kke 62 30527 30542 1658 548041 1450 1465 TGTGAGAAAGGTGTAT eekd.sub.10kke 64 30528 30543 1659 548042 1452 1467 TTTGTGAGAAAGGTGT eekd.sub.10kke 63 30530 30545 1660 548043 1453 1468 ATTTGTGAGAAAGGTG eekd.sub.10kke 76 30531 30546 1661 548044 1474 1489 TGGTGAATAATAATCT eekd.sub.10kke 12 30552 30567 1662 548045 1483 1498 TTATAGTTTTGGTGAA eekd.sub.10kke 0 30561 30576 1663 548046 1506 1521 TATCATGATTCCCTTC eekd.sub.10kke 84 30584 30599 1664 548047 1508 1523 GATATCATGATTCCCT eekd.sub.10kke 83 30586 30601 1665 548048 1509 1524 CGATATCATGATTCCC eekd.sub.10kke 84 30587 30602 1666 548049 1510 1525 GCGATATCATGATTCC eekd.sub.10kke 62 30588 30603 1667 548050 1512 1527 AGGCGATATCATGATT eekd.sub.10kke 37 30590 30605 1668 548051 1513 1528 AAGGCGATATCATGAT eekd.sub.10kke 61 30591 30606 1669 548052 1535 1550 CAAAGGAGCCTGGAGT eekd.sub.10kke 43 30613 30628 1670 548053 1538 1553 ATTCAAAGGAGCCTGG eekd.sub.10kke 36 30616 30631 1671 548054 1539 1554 AATTCAAAGGAGCCTG eekd.sub.10kke 45 30617 30632 1672 548055 1541 1556 GTAATTCAAAGGAGCC eekd.sub.10kke 78 30619 30634 1673 548056 1543 1558 GTGTAATTCAAAGGAG eekd.sub.10kke 40 30621 30636 1674 548057 1564 1579 CATATTGGTTTTTGGA eekd.sub.10kke 49 31870 31885 1675 548058 1565 1580 GCATATTGGTTTTTGG eekd.sub.10kke 71 31871 31886 1676 548059 1568 1583 TAGGCATATTGGTTTT eekd.sub.10kke 50 31874 31889 1677 548060 1588 1603 CTTGTGTCACCTTTGG eekd.sub.10kke 76 31894 31909 1678 548061 1589 1604 GCTTGTGTCACCTTTG eekd.sub.10kke 86 31895 31910 1679 548062 1598 1613 ATAAATTGTGCTTGTG eekd.sub.10kke 19 31904 31919 1680 548063 1600 1615 GTATAAATTGTGCTTG eekd.sub.10kke 35 31906 31921 1681 548064 1602 1617 TGGTATAAATTGTGCT eekd.sub.10kke 54 31908 31923 1682 548065 1603 1618 TTGGTATAAATTGTGC eekd.sub.10kke 22 31909 31924 1683 548067 1606 1621 CAGTTGGTATAAATTG eekd.sub.10kke 18 31912 31927 1684 548068 1609 1624 CAACAGTTGGTATAAA eekd.sub.10kke 0 31915 31930 1685 548069 1610 1625 CCAACAGTTGGTATAA eekd.sub.10kke 57 31916 31931 1686 548070 1611 1626 CCCAACAGTTGGTATA eekd.sub.10kke 85 31917 31932 1687 548071 1629 1644 AGAAGCCCCATCCGGT eekd.sub.10kke 55 31935 31950 1688 548072 1640 1655 TTTCTCCTTCGAGAAG eekd.sub.10kke 33 31946 31961 1689 548073 1641 1656 CTTTCTCCTTCGAGAA eekd.sub.10kke 24 31947 31962 1690
TABLE-US-00137 TABLE 146 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 19 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 66 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548151 2139 2154 GGGCTTCAGCCAGACA eekd.sub.10kke 35 34238 34253 1691 548152 2140 2155 CGGGCTTCAGCCAGAC eekd.sub.10kke 32 34239 34254 1692 548153 2149 2164 TGCTGAAAGCGGGCTT eekd.sub.10kke 44 34248 34263 1693 548154 2150 2165 GTGCTGAAAGCGGGCT eekd.sub.10kke 7 34249 34264 1694 548155 2151 2166 CGTGCTGAAAGCGGGC eekd.sub.10kke 76 34250 34265 1695 548156 2168 2183 TCAGCCCCTGGTTACG eekd.sub.10kke 0 34267 34282 1696 548157 2172 2187 ATTGTCAGCCCCTGGT eekd.sub.10kke 7 34271 34286 1697 548158 2174 2189 GCATTGTCAGCCCCTG eekd.sub.10kke 18 34273 34288 1698 548159 2175 2190 CGCATTGTCAGCCCCT eekd.sub.10kke 59 34274 34289 1699 548160 2176 2191 TCGCATTGTCAGCCCC eekd.sub.10kke 60 34275 34290 1700 548161 2177 2192 CTCGCATTGTCAGCCC eekd.sub.10kke 59 34276 34291 1701 548162 2178 2193 CCTCGCATTGTCAGCC eekd.sub.10kke 25 34277 34292 1702 548163 2179 2194 ACCTCGCATTGTCAGC eekd.sub.10kke 46 34278 34293 1703 548164 2180 2195 GACCTCGCATTGTCAG eekd.sub.10kke 40 34279 34294 1704 548165 2181 2196 CGACCTCGCATTGTCA eekd.sub.10kke 53 34280 34295 1705 548166 2182 2197 GCGACCTCGCATTGTC eekd.sub.10kke 0 34281 34296 1706 548167 2183 2198 TGCGACCTCGCATTGT eekd.sub.10kke 36 34282 34297 1707 548168 2184 2199 TTGCGACCTCGCATTG eekd.sub.10kke 61 34283 34298 1708 548169 2185 2200 GTTGCGACCTCGCATT eekd.sub.10kke 7 34284 34299 1709 548170 2186 2201 AGTTGCGACCTCGCAT eekd.sub.10kke 68 34285 34300 1710 548171 2187 2202 CAGTTGCGACCTCGCA eekd.sub.10kke 47 34286 34301 1711 548172 2188 2203 TCAGTTGCGACCTCGC eekd.sub.10kke 0 34287 34302 1712 548173 2189 2204 CTCAGTTGCGACCTCG eekd.sub.10kke 51 34288 34303 1713 548174 2190 2205 TCTCAGTTGCGACCTC eekd.sub.10kke 68 34289 34304 1714 548175 2191 2206 ATCTCAGTTGCGACCT eekd.sub.10kke 0 34290 34305 1715 548176 2192 2207 GATCTCAGTTGCGACC eekd.sub.10kke 38 34291 34306 1716 548177 2193 2208 AGATCTCAGTTGCGAC eekd.sub.10kke 45 34292 34307 1717 548178 2194 2209 GAGATCTCAGTTGCGA eekd.sub.10kke 54 34293 34308 1718 548179 2195 2210 GGAGATCTCAGTTGCG eekd.sub.10kke 52 34294 34309 1719 548180 2199 2214 TCATGGAGATCTCAGT eekd.sub.10kke 79 34298 34313 1720 548181 2200 2215 GTCATGGAGATCTCAG eekd.sub.10kke 55 34299 34314 1721 548182 2201 2216 AGTCATGGAGATCTCA eekd.sub.10kke 55 34300 34315 1722 548183 2202 2217 CAGTCATGGAGATCTC eekd.sub.10kke 43 34301 34316 1723 548184 2203 2218 ACAGTCATGGAGATCT eekd.sub.10kke 73 34302 34317 1724 548185 2208 2223 AACACACAGTCATGGA eekd.sub.10kke 23 34307 34322 1725 548186 2209 2224 CAACACACAGTCATGG eekd.sub.10kke 0 34308 34323 1726 548187 n/a n/a CATCCTATCCGTGTTC eekd.sub.10kke 33 3279 3294 1727 548189 n/a n/a CATGAACATCCTATCC eekd.sub.10kke 24 3285 3300 1728 548190 n/a n/a TATTCCATGAACATCC eekd.sub.10kke 43 3290 3305 1729 548191 n/a n/a GTCAACATATTCCATG eekd.sub.10kke 0 3297 3312 1730 548192 n/a n/a CCTGTCAACATATTCC eekd.sub.10kke 65 3300 3315 1731 548193 n/a n/a TGTCCTGTCAACATAT eekd.sub.10kke 58 3303 3318 1732 548194 n/a n/a GCCAACAGTTTCAACT eekd.sub.10kke 61 3322 3337 1733 548195 n/a n/a TTCTGCCAACAGTTTC eekd.sub.10kke 84 3326 3341 1734 548196 n/a n/a CAATATTGACTTTGGG eekd.sub.10kke 6 3343 3358 1735 548197 n/a n/a TGCTTGGCTTCAATAT eekd.sub.10kke 68 3353 3368 1736 548198 n/a n/a ACTGCAGGCAATATTT eekd.sub.10kke 49 3369 3384 1737 548199 n/a n/a GCACTGCAGGCAATAT eekd.sub.10kke 24 3371 3386 1738 548200 n/a n/a CTAATGTGGCACTGCA eekd.sub.10kke 19 3379 3394 1739 548201 n/a n/a TGTTCTAATGTGGCAC eekd.sub.10kke 67 3383 3398 1740 548202 n/a n/a GCTGTTCTAATGTGGC eekd.sub.10kke 9 3385 3400 1741 548203 n/a n/a TGACTAGTGAATGGCT eekd.sub.10kke 73 2280 2295 1742 548204 n/a n/a TCTGACTAGTGAATGG eekd.sub.10kke 25 2282 2297 1743 548205 n/a n/a TCAATCTGACTAGTGA eekd.sub.10kke 14 2286 2301 1744 548206 n/a n/a GGTCAATCTGACTAGT eekd.sub.10kke 45 2288 2303 1745 548207 n/a n/a CTGGTCAATCTGACTA eekd.sub.10kke 60 2290 2305 1746 548208 n/a n/a CTCTGGTCAATCTGAC eekd.sub.10kke 19 2292 2307 1747 548209 n/a n/a CAATCTCTGGTCAATC eekd.sub.10kke 57 2296 2311 1748 548210 n/a n/a CAACAATCTCTGGTCA eekd.sub.10kke 55 2299 2314 1749 548211 n/a n/a ACCAACAATCTCTGGT eekd.sub.10kke 51 2301 2316 1750 548212 n/a n/a AGCCCACCAACAATCT eekd.sub.10kke 44 2306 2321 1751 548213 n/a n/a GACAGCCCACCAACAA eekd.sub.10kke 70 2309 2324 1752 548214 n/a n/a CAGACAGCCCACCAAC eekd.sub.10kke 55 2311 2326 1753 548215 n/a n/a GCATAGACCCCAACAG eekd.sub.10kke 61 2324 2339 1754 548216 n/a n/a GTGCATAGACCCCAAC eekd.sub.10kke 45 2326 2341 1755 548217 n/a n/a CTGTGCATAGACCCCA eekd.sub.10kke 69 2328 2343 1756 548218 n/a n/a TCCTGTGCATAGACCC eekd.sub.10kke 59 2330 2345 1757 548219 n/a n/a GAAATCCTGTGCATAG eekd.sub.10kke 8 2334 2349 1758 548220 n/a n/a GCAGAAATCCTGTGCA eekd.sub.10kke 69 2337 2352 1759 548221 n/a n/a ACTCCAGCAGAAATCC eekd.sub.10kke 49 2343 2358 1760 548222 n/a n/a AATCATGCCTTGTGGG eekd.sub.10kke 32 4765 4780 1761 548223 n/a n/a TAGACCCAGAATCATG eekd.sub.10kke 50 4774 4789 1762 548224 n/a n/a CCATAGACCCAGAATC eekd.sub.10kke 20 4777 4792 1763 548225 n/a n/a AGTCACCATAGACCCA eekd.sub.10kke 48 4782 4797 1764 548226 n/a n/a TAAGTCACCATAGACC eekd.sub.10kke 39 4784 4799 1765 548227 n/a n/a GTGGCCCTCTTAAGTC eekd.sub.10kke 0 4794 4809 1766
TABLE-US-00138 TABLE 147 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 42 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 80 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548228 n/a n/a GTTGTGTGGCCCTCTT eekd.sub.10kke 37 4799 4814 1767 548229 n/a n/a CATTGTTGTGTGGCCC eekd.sub.10kke 31 4803 4818 1768 548230 n/a n/a TACTCATTGTTGTGTG eekd.sub.10kke 10 4807 4822 1769 548231 n/a n/a AATACTCATTGTTGTG eekd.sub.10kke 11 4809 4824 1770 548232 n/a n/a GCCATACATCTGAGGA eekd.sub.10kke 3 4831 4846 1771 548233 n/a n/a ATTGTAGCCATACATC eekd.sub.10kke 38 4837 4852 1772 548234 n/a n/a TTATTGTAGCCATACA eekd.sub.10kke 17 4839 4854 1773 548235 n/a n/a TCTAGATGACCTGAAG eekd.sub.10kke 0 18147 18162 1774 548236 n/a n/a TACATCTAGATGACCT eekd.sub.10kke 37 18151 18166 1775 548237 n/a n/a GTATACATCTAGATGA eekd.sub.10kke 22 18154 18169 1776 548238 n/a n/a ACTCGCCTTTGTGACT eekd.sub.10kke 31 26268 26283 1777 548239 n/a n/a TACTCGCCTTTGTGAC eekd.sub.10kke 18 26269 26284 1778 548240 n/a n/a ATACTCGCCTTTGTGA eekd.sub.10kke 3 26270 26285 1779 26301 26316 548241 n/a n/a CATACTCGCCTTTGTG eekd.sub.10kke 1 26271 26286 1780 26302 26317 548242 n/a n/a GCATACTCGCCTTTGT eekd.sub.10kke 25 26272 26287 1781 26303 26318 548243 n/a n/a ATGCATACTCGCCTTT eekd.sub.10kke 0 26274 26289 1782 26305 26320 548244 n/a n/a CATGCATACTCGCCTT eekd.sub.10kke 51 26275 26290 1783 26306 26321 548245 n/a n/a CCATGCATACTCGCCT eekd.sub.10kke 31 26276 26291 1784 26307 26322 548246 n/a n/a TTCCATGCATACTCGC eekd.sub.10kke 46 26278 26293 1785 548247 n/a n/a CGATTTTCCATGCATA eekd.sub.10kke 56 26283 26298 1786 548248 n/a n/a TGCGATTTTCCATGCA eekd.sub.10kke 13 26285 26300 1787 548249 n/a n/a TGTGATGCGATTTTCC eekd.sub.10kke 22 26290 26305 1788 548250 n/a n/a CTTTGTGATGCGATTT eekd.sub.10kke 0 26293 26308 1789 548251 n/a n/a GCCTTTGTGATGCGAT eekd.sub.10kke 13 26295 26310 1790 548252 n/a n/a ACTCGCCTTTGTGATG eekd.sub.10kke 33 26299 26314 1791 548253 n/a n/a TACTCGCCTTTGTGAT eekd.sub.10kke 8 26300 26315 1792 548254 n/a n/a CCCATGCATACTCGCC eekd.sub.10kke 39 26308 26323 1793 548255 n/a n/a CCCCATGCATACTCGC eekd.sub.10kke 38 26309 26324 1794 548256 n/a n/a GCTCCCCATGCATACT eekd.sub.10kke 25 26312 26327 1795 548257 n/a n/a AGTGCTCCCCATGCAT eekd.sub.10kke 2 26315 26330 1796 548258 n/a n/a CAAGTGCTCCCCATGC eekd.sub.10kke 0 26317 26332 1797 548259 n/a n/a GTGATGAAAGTACAGC eekd.sub.10kke 45 26335 26350 1798 548260 n/a n/a AGGAGTTTGTCAGAAC eekd.sub.10kke 28 3210 3225 1799 548261 n/a n/a TTCAGGGAGTGATGTC eekd.sub.10kke 36 3241 3256 1800 548262 n/a n/a CCTATCCGTGTTCAGC eekd.sub.10kke 73 3276 3291 1801 548263 n/a n/a CTCTACATACTCAGGA eekd.sub.10kke 62 3561 3576 1802 548264 n/a n/a CAGTCCAAAAATCCCT eekd.sub.10kke 60 3701 3716 1803 548265 n/a n/a CCTCTTGATTTGGGCA eekd.sub.10kke 85 3749 3764 1804 548266 n/a n/a TTGGCCAACTCTGTGG eekd.sub.10kke 44 3816 3831 1805 548267 n/a n/a GACCTCCAGACTACTG eekd.sub.10kke 34 3848 3863 1806 548268 n/a n/a TGTGTCTAGGGAGTTG eekd.sub.10kke 52 3898 3913 1807 548269 n/a n/a AGCACACAATTACTGG eekd.sub.10kke 62 3946 3961 1808 548270 n/a n/a CTGCTGGTTTTAGACC eekd.sub.10kke 28 4029 4044 1809 548271 n/a n/a TTCACTTACCACAGGA eekd.sub.10kke 56 4122 4137 1810 548272 n/a n/a GGTGCCACTTGCTTGG eekd.sub.10kke 54 4178 4193 1811 548273 n/a n/a AATCTCCACCCCCGAA eekd.sub.10kke 5 4224 4239 1812 548274 n/a n/a TACCTGACAAGTGGTC eekd.sub.10kke 0 4287 4302 1813 548275 n/a n/a GTCCCAAGACATTCCT eekd.sub.10kke 40 4350 4365 1814 548276 n/a n/a CAGAGTGTCATCTGCG eekd.sub.10kke 49 4389 4404 1815 548277 n/a n/a GGATTGGACCCAGACA eekd.sub.10kke 57 4511 4526 1816 548278 n/a n/a GGTTCCCTAGCGGTCC eekd.sub.10kke 74 4564 4579 1817 548279 n/a n/a CACCTAGAACTATCCA eekd.sub.10kke 39 4632 4647 1818 548280 n/a n/a CTCCCTCTGTAATGAT eekd.sub.10kke 43 4736 4751 1819 548281 n/a n/a GGTTGAGGGACAGACA eekd.sub.10kke 0 4944 4959 1820 548282 n/a n/a GTGGGTTTGCACATGG eekd.sub.10kke 73 4992 5007 1821 548283 n/a n/a GGCTTATGCTCCTTCT eekd.sub.10kke 56 5017 5032 1822 548284 n/a n/a CCCCCTGTAGTTGGCT eekd.sub.10kke 35 5051 5066 1823 548285 n/a n/a GCTTACTTACATCCCT eekd.sub.10kke 52 5132 5147 1824 548286 n/a n/a GGGACTACATGCAATA eekd.sub.10kke 47 5166 5181 1825 548287 n/a n/a GTCAAAGAGTGTCCAC eekd.sub.10kke 38 5283 5298 1826 548288 n/a n/a GAATAGCAAGCTCCAA eekd.sub.10kke 64 5348 5363 1827 548289 n/a n/a CATGATACCACACCAC eekd.sub.10kke 28 5484 5499 1828 548290 n/a n/a GAGCACTCTTATTAGC eekd.sub.10kke 31 5546 5561 1829 548291 n/a n/a CCTGTTAGAGTTGGCC eekd.sub.10kke 35 5576 5591 1830 548292 n/a n/a AGGACACTGTTTCCAG eekd.sub.10kke 38 5627 5642 1831 548293 n/a n/a GTCACCAGAACCACAT eekd.sub.10kke 44 5683 5698 1832 548294 n/a n/a GTGTGCACTTTCTGGT eekd.sub.10kke 33 5716 5731 1833 548295 n/a n/a CTCTGATTGGGTCACC eekd.sub.10kke 26 5746 5761 1834 548296 n/a n/a ACCAACAACTCAGGCC eekd.sub.10kke 34 5858 5873 1835 548297 n/a n/a ACTCTCAAGCTCCACG eekd.sub.10kke 32 5889 5904 1836 548298 n/a n/a GGACAATATGTCTCCT eekd.sub.10kke 0 5935 5950 1837 548299 n/a n/a CATTGTGCTCAACTGA eekd.sub.10kke 35 5961 5976 1838 548300 n/a n/a GCCCATGGTGAATCTG eekd.sub.10kke 53 5995 6010 1839 548301 n/a n/a CCTAGTACAAAGTGGC eekd.sub.10kke 65 6050 6065 1840 548302 n/a n/a GCCATTTTATCCCTGA eekd.sub.10kke 71 6134 6149 1841 548303 n/a n/a GGGCCCCCATGTCCAT eekd.sub.10kke 0 6336 6351 1842
TABLE-US-00139 TABLE 148 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 72 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 67 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548305 n/a n/a GTTCTTGCTTATCCTC eekd.sub.10kke 55 6484 6499 1843 548306 n/a n/a ATGTGACAGTCAGGGA eekd.sub.10kke 8 6559 6574 1844 548307 n/a n/a TTCTGCAACTGAGCCT eekd.sub.10kke 6 6587 6602 1845 548308 n/a n/a AATGGCAGGTCCTGGC eekd.sub.10kke 9 6616 6631 1846 548309 n/a n/a AGACAGTTGGTGGTTT eekd.sub.10kke 41 6700 6715 1847 548310 n/a n/a GAGGAGTTGGTTTAGT eekd.sub.10kke 0 6750 6765 1848 548311 n/a n/a TGACCACCTCTCGGGT eekd.sub.10kke 10 6860 6875 1849 548312 n/a n/a ATTTGGCCCTGAGCCC eekd.sub.10kke 0 6935 6950 1850 548313 n/a n/a GCCTTTGAGGGAGTGG eekd.sub.10kke 35 7024 7039 1851 548314 n/a n/a ACAACCTGTCCATTCC eekd.sub.10kke 43 7087 7102 1852 548315 n/a n/a GTTGTCAACTGGGACC eekd.sub.10kke 14 7125 7140 1853 548316 n/a n/a CTGTTCAGGTAGCACA eekd.sub.10kke 64 7150 7165 1854 548317 n/a n/a CCGGGAAAGACTGTCT eekd.sub.10kke 42 7190 7205 1855 548318 n/a n/a ACTGCACCCCACATAT eekd.sub.10kke 18 7257 7272 1856 548319 n/a n/a CCTCATCTCAGTATGA eekd.sub.10kke 26 7398 7413 1857 548320 n/a n/a GCACACAGACTTGCCC eekd.sub.10kke 0 7508 7523 1858 548321 n/a n/a CTGCATCTGGACTATG eekd.sub.10kke 38 7559 7574 1859 548322 n/a n/a AGGGAAATTAGAGGCA eekd.sub.10kke 38 7586 7601 1860 548323 n/a n/a CTGTTGCCTGACATGC eekd.sub.10kke 43 7696 7711 1861 548324 n/a n/a ACATAAATTCCCCACA eekd.sub.10kke 29 7741 7756 1862 548325 n/a n/a CCCACTGACTGACTAC eekd.sub.10kke 27 7906 7921 1863 548326 n/a n/a TCCTGTGACAGAACCA eekd.sub.10kke 27 7988 8003 1864 548327 n/a n/a CTACACCTTTCTGCAC eekd.sub.10kke 6 8221 8236 1865 548328 n/a n/a GGTCCTTGAACCCCGT eekd.sub.10kke 68 8260 8275 1866 548329 n/a n/a AGCAGATCTGGGTTGT eekd.sub.10kke 59 8328 8343 1867 548330 n/a n/a GACTAGCTTCTACTAC eekd.sub.10kke 34 8404 8419 1868 548331 n/a n/a ACAATCCCTTAGCCCA eekd.sub.10kke 73 8457 8472 1869 548332 n/a n/a GATGAAATGTGCACCT eekd.sub.10kke 46 8491 8506 1870 548333 n/a n/a GACTGTGCTATCCGCT eekd.sub.10kke 58 8550 8565 1871 548334 n/a n/a GCTCACTATAGGCCCC eekd.sub.10kke 69 8656 8671 1872 548335 n/a n/a TAGCATCATGCCACAG eekd.sub.10kke 51 8684 8699 1873 548336 n/a n/a GCACATTAGGAGGTAG eekd.sub.10kke 1 9039 9054 1874 548337 n/a n/a TACCGCTGGGTGCGGT eekd.sub.10kke 10 9075 9090 1875 548338 n/a n/a ATGAAACTGTGGCTCG eekd.sub.10kke 80 9131 9146 1876 548339 n/a n/a ACATGTGGGATCAGAG eekd.sub.10kke 37 9275 9290 1877 548340 n/a n/a GATGATCCTCACATAC eekd.sub.10kke 35 9316 9331 1878 548341 n/a n/a TAGAACCTTCCTCCAC eekd.sub.10kke 30 9341 9356 1879 548342 n/a n/a GGAAGACTTCCCTCTG eekd.sub.10kke 0 9403 9418 1880 548343 n/a n/a TAGTGATAAGAGCTGG eekd.sub.10kke 78 9472 9487 1881 548344 n/a n/a GGCAACTATGTTCTCA eekd.sub.10kke 76 9536 9551 1882 548345 n/a n/a CTAACTCCATCACTGC eekd.sub.10kke 55 9637 9652 1883 548346 n/a n/a TCCCCAATACTTGCTG eekd.sub.10kke 35 9696 9711 1884 548347 n/a n/a GCTGTTCTAAGCGAGA eekd.sub.10kke 31 9976 9991 1885 548348 n/a n/a TGAGTGATGCCTTCCA eekd.sub.10kke 82 10024 10039 1886 548349 n/a n/a TCCAGAATACTGCCCC eekd.sub.10kke 61 10054 10069 1887 548350 n/a n/a GCGCTAACCTCATAAA eekd.sub.10kke 29 10148 10163 1888 548351 n/a n/a CTGGAAACGAGACACA eekd.sub.10kke 33 10201 10216 1889 548352 n/a n/a GAGAGAGATGTTCCCT eekd.sub.10kke 47 10240 10255 1890 548353 n/a n/a CTGCTGGTTGAGAATC eekd.sub.10kke 48 10287 10302 1891 548354 n/a n/a ATGTCCCCAGTGGAAG eekd.sub.10kke 41 10314 10329 1892 548355 n/a n/a GCATCCTCCCTAGTTG eekd.sub.10kke 47 10362 10377 1893 548356 n/a n/a TGTTGGTCAGCATTCA eekd.sub.10kke 63 10411 10426 1894 548357 n/a n/a GACGACTGCCCTGTGC eekd.sub.10kke 69 10436 10451 1895 548358 n/a n/a ATTTGGGCCTAGTGGT eekd.sub.10kke 0 10515 10530 1896 548359 n/a n/a CCTAGTCCTCAAGTTT eekd.sub.10kke 0 10580 10595 1897 548360 n/a n/a CAAGACATCAGTAGCT eekd.sub.10kke 45 10626 10641 1898 548361 n/a n/a CTTATCAGTCCCAGTC eekd.sub.10kke 52 10702 10717 1899 548362 n/a n/a GACAACCCATCAGTTG eekd.sub.10kke 33 10742 10757 1900 548363 n/a n/a CAGCAGGCTCAAAGTG eekd.sub.10kke 37 10915 10930 1901 548364 n/a n/a TGGCTAAGTCAGGCCC eekd.sub.10kke 30 10982 10997 1902 548365 n/a n/a TGTACTCCACCTCACG eekd.sub.10kke 55 11017 11032 1903 548366 n/a n/a AGCAAGCTAAGTGAGT eekd.sub.10kke 5 11199 11214 1904 548367 n/a n/a GTTCTTGAGTGTAGAG eekd.sub.10kke 52 11260 11275 1905 548368 n/a n/a GTGTTCATACGGAAGC eekd.sub.10kke 59 11299 11314 1906 548369 n/a n/a GTTGGGATGCGACTCT eekd.sub.10kke 50 11335 11350 1907 548370 n/a n/a ACGAAGTCTCTTTCCT eekd.sub.10kke 53 11385 11400 1908 548371 n/a n/a CGATGAGTTGGGCAGG eekd.sub.10kke 57 11454 11469 1909 548372 n/a n/a GATACCTTTCCACTCC eekd.sub.10kke 61 11558 11573 1910 548373 n/a n/a TCCCCAAGATTATGTG eekd.sub.10kke 16 11596 11611 1911 548374 n/a n/a GCACCCTTTTCATTGA eekd.sub.10kke 41 12074 12089 1912 548375 n/a n/a TCGACTTCTCCTGTCT eekd.sub.10kke 27 12199 12214 1913 548376 n/a n/a GCCTTTGACCTTTCGC eekd.sub.10kke 65 12261 12276 1914 548377 n/a n/a GTGTGCTGAGGTTTGC eekd.sub.10kke 80 12297 12312 1915 548378 n/a n/a GCAAGATGCATGCAGC eekd.sub.10kke 49 12393 12408 1916 548379 n/a n/a ATCGAACTCTGCTTGA eekd.sub.10kke 44 12477 12492 1917 548380 n/a n/a GCCCAGTTTTGGCAAC eekd.sub.10kke 7 12540 12555 1918 548381 n/a n/a CCCACTACCATTTGGG eekd.sub.10kke 0 12578 12593 1919
TABLE-US-00140 TABLE 149 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 46 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 64 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548459 n/a n/a CAACTATAACAGTATC eekd.sub.10kke 26 15903 15918 1920 548460 n/a n/a CTATACCACGGTAACT eekd.sub.10kke 0 16036 16051 1921 548461 n/a n/a CCTATATCACTGTAAC eekd.sub.10kke 0 16127 16142 1922 548462 n/a n/a ACCTATATCACTGTAA eekd.sub.10kke 0 16128 16143 1923 548463 n/a n/a TCACTGTACCTATATC eekd.sub.10kke 0 16135 16150 1924 548464 n/a n/a GTCCTATAACTATATC eekd.sub.10kke 0 16174 16189 1925 548465 n/a n/a CTGTACCTATAACTGT eekd.sub.10kke 0 16202 16217 1926 548466 n/a n/a CGTCACTGTACCTATA eekd.sub.10kke 71 16207 16222 1927 548467 n/a n/a CATCACTGTACCTATA eekd.sub.10kke 20 16258 16273 1928 548468 n/a n/a CAACATCACTGTACCT eekd.sub.10kke 6 16261 16276 1929 548469 n/a n/a TTCCCTACCCCTGGTA eekd.sub.10kke 0 16331 16346 1930 548470 n/a n/a GGTGGAATGTCATGGC eekd.sub.10kke 56 16404 16419 1931 548471 n/a n/a GCGGAAAACTGGCCGT eekd.sub.10kke 17 16474 16489 1932 548472 n/a n/a CCCAATACAGGGCCAG eekd.sub.10kke 0 16513 16528 1933 548473 n/a n/a CCAACCTTCCCAATCT eekd.sub.10kke 0 16554 16569 1934 548474 n/a n/a GAAGGTGTGCTGTCGC eekd.sub.10kke 33 16602 16617 1935 548475 n/a n/a ATCGAGTCCTGCCTCC eekd.sub.10kke 17 16707 16722 1936 548476 n/a n/a GCAAATCCTTCCAGCA eekd.sub.10kke 27 16755 16770 1937 548477 n/a n/a GCACGAGCTTGCCTGT eekd.sub.10kke 26 16787 16802 1938 548478 n/a n/a GAGCCATCCAGGGTGC eekd.sub.10kke 53 16845 16860 1939 548479 n/a n/a AGGCCATTTGATCCGA eekd.sub.10kke 68 16913 16928 1940 548480 n/a n/a GCCACGCCCTTAGCAG eekd.sub.10kke 20 16973 16988 1941 548481 n/a n/a GTTCCCTGAGGAACGG eekd.sub.10kke 2 17010 17025 1942 548482 n/a n/a GGCAGTTAGGCCAGGA eekd.sub.10kke 53 17068 17083 1943 548483 n/a n/a CTACAGATCATCCCTA eekd.sub.10kke 5 17102 17117 1944 548484 n/a n/a CCCCGGAGCACCTTCA eekd.sub.10kke 41 17207 17222 1945 548485 n/a n/a GTGACCCAAGGGTCGA eekd.sub.10kke 17 17252 17267 1946 548486 n/a n/a CGTGGTTAGCCTGACA eekd.sub.10kke 68 17416 17431 1947 548487 n/a n/a TCCATGTCAGAGTTGC eekd.sub.10kke 71 17461 17476 1948 548488 n/a n/a CCTCCTTTTGGCTTGA eekd.sub.10kke 63 17530 17545 1949 548489 n/a n/a TTCCCCAGAGGTGATA eekd.sub.10kke 16 17582 17597 1950 548490 n/a n/a TCTGGTTAGCCTCCGA eekd.sub.10kke 58 17664 17679 1951 548491 n/a n/a TGGCCAAGCAACCAGT eekd.sub.10kke 57 17715 17730 1952 548492 n/a n/a GCCCAATGTCCTAACC eekd.sub.10kke 51 17794 17809 1953 548493 n/a n/a CCACCGCTGCCCGCCA eekd.sub.10kke 37 18013 18028 1954 548494 n/a n/a TGTGACCCCCCACCGC eekd.sub.10kke 39 18022 18037 1955 548495 n/a n/a TTGTGACCCCCCACCG eekd.sub.10kke 55 18023 18038 1956 548496 n/a n/a ACTGAACCCCCTTAGG eekd.sub.10kke 0 18571 18586 1957 548497 n/a n/a CCTTCATACCCCTCAC eekd.sub.10kke 26 18725 18740 1958 548498 n/a n/a CCGATAACAGACCGGC eekd.sub.10kke 71 18795 18810 1959 548499 n/a n/a ATACCCGGAGTCAGGA eekd.sub.10kke 56 18955 18970 1960 548500 n/a n/a ATTGCTCAGGCCCCCT eekd.sub.10kke 29 19037 19052 1961 548501 n/a n/a CAAGCCACTAACCCAC eekd.sub.10kke 33 19147 19162 1962 548502 n/a n/a AATTCTTGGACCAAGG eekd.sub.10kke 25 19234 19249 1963 548503 n/a n/a CCATCTACTCCCCCAT eekd.sub.10kke 9 19291 19306 1964 548504 n/a n/a GCAGCGAGCATTCCAA eekd.sub.10kke 28 19352 19367 1965 548505 n/a n/a GGACAATGCCTATGCT eekd.sub.10kke 21 19386 19401 1966 548506 n/a n/a GAAGCCATTCACTGCA eekd.sub.10kke 32 19436 19451 1967 548507 n/a n/a AAACTCCTCTCAAGGC eekd.sub.10kke 53 19474 19489 1968 548508 n/a n/a GCACCACCATGCGGTT eekd.sub.10kke 43 19553 19568 1969 548509 n/a n/a TGCAGGGCTGCGCAGT eekd.sub.10kke 41 19960 19975 1970 548510 n/a n/a TTAGCCACTCCTCTTG eekd.sub.10kke 30 20062 20077 1971 548511 n/a n/a AGCTAGCTGACCCCAA eekd.sub.10kke 16 20092 20107 1972 548512 n/a n/a TCCGCCTTTGGATACT eekd.sub.10kke 49 20155 20170 1973 548513 n/a n/a CCTGCTGATTGTGTCT eekd.sub.10kke 16 20240 20255 1974 548514 n/a n/a TCGAGGACAGCCCCCA eekd.sub.10kke 40 20335 20350 1975 548515 n/a n/a ACCCGTCAGCCTCAGC eekd.sub.10kke 59 20381 20396 1976 548516 n/a n/a CTTGCCTATTCACCCC eekd.sub.10kke 49 20544 20559 1977 548517 n/a n/a CGGACAAGCCTTACAG eekd.sub.10kke 43 20596 20611 1978 548518 n/a n/a CACACTTACCCCGCTC eekd.sub.10kke 12 20741 20756 1979 548519 n/a n/a CCTCCCCTTGTGTGTC eekd.sub.10kke 31 20843 20858 1980 548520 n/a n/a CCGCTTCCCTGACTGT eekd.sub.10kke 43 20919 20934 1981 548521 n/a n/a CAGCTCCCTTACTAGG eekd.sub.10kke 61 20958 20973 1982 548522 n/a n/a AGGTATTGACCGCCAG eekd.sub.10kke 55 21062 21077 1983 548523 n/a n/a GGTAAATCCATCCCCT eekd.sub.10kke 44 21157 21172 1984 548524 n/a n/a GCCCGATCACCTTAGA eekd.sub.10kke 45 21220 21235 1985 548525 n/a n/a GTCTAACTGGCCTGGC eekd.sub.10kke 2 21328 21343 1986 548526 n/a n/a CTAAGCTGTGTCTCAT eekd.sub.10kke 26 21373 21388 1987 548527 n/a n/a TGTTTCAAGTGCCAGA eekd.sub.10kke 50 21434 21449 1988 548528 n/a n/a TGCAGTGGTCAAGCAT eekd.sub.10kke 32 21478 21493 1989 548529 n/a n/a GCGATTCCTTGCCTCT eekd.sub.10kke 56 21554 21569 1990 548530 n/a n/a ATAATAGAGGCAGCCA eekd.sub.10kke 50 21592 21607 1991 548531 n/a n/a GTCAGAAGGCCTCTTA eekd.sub.10kke 21 21753 21768 1992 548532 n/a n/a TATTTATCCGACCTCT eekd.sub.10kke 34 21881 21896 1993 548533 n/a n/a GAGGTGGTTGGAGCTA eekd.sub.10kke 9 21926 21941 1994 548534 n/a n/a CAGATCCCAATTCTTC eekd.sub.10kke 22 22063 22078 1995 548535 n/a n/a GAGTCTTTCCAATCCT eekd.sub.10kke 13 22142 22157 1996
TABLE-US-00141 TABLE 150 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Chemistry inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 46 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 64 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548536 n/a n/a TCTCAATCCCAACCCC eekd.sub.10kke 0 22168 22183 1997 548537 n/a n/a CCTCAATCCCAACCCA eekd.sub.10kke 0 22191 22206 1998 548538 n/a n/a TAGTGGCAAGAACCAC eekd.sub.10kke 0 22627 22642 1999 548539 n/a n/a CGCGCGAATGCCTGCC eekd.sub.10kke 41 22658 22673 2000 548540 n/a n/a GACACCTGCTTGATTA eekd.sub.10kke 7 22704 22719 2001 548541 n/a n/a GGCACTGGTCATGGAC eekd.sub.10kke 39 22760 22775 2002 548542 n/a n/a GCGCCATCCTTCAATC eekd.sub.10kke 7 22857 22872 2003 548543 n/a n/a GATCCACCCATGACCT eekd.sub.10kke 32 22997 23012 2004 548544 n/a n/a GCTGTGACTCAGATCA eekd.sub.10kke 62 23070 23085 2005 548545 n/a n/a CTCTTCGCATGGACAC eekd.sub.10kke 46 23100 23115 2006 548546 n/a n/a GCCCAAGCCTACATGC eekd.sub.10kke 35 23430 23445 2007 548547 n/a n/a GTGCGATTAAGCCCCA eekd.sub.10kke 86 23514 23529 2008 548548 n/a n/a GCTTGTAGAAGGGATT eekd.sub.10kke 54 23631 23646 2009 548549 n/a n/a TGTGCAATCAGGTGGA eekd.sub.10kke 56 23765 23780 2010 548550 n/a n/a CCGGCCTGGATACAGC eekd.sub.10kke 0 23831 23846 2011 548551 n/a n/a CGGCCAATGGGAAAGG eekd.sub.10kke 25 24175 24190 2012 548552 n/a n/a TGGAGGAGTAGGGAAT eekd.sub.10kke 10 24200 24215 2013 548553 n/a n/a CCCGAAGAGTCAAGTC eekd.sub.10kke 46 24255 24270 2014 548554 n/a n/a GTGCTGCATTGCATGA eekd.sub.10kke 42 24290 24305 2015 548555 n/a n/a ACACGCCAGGTGAAAA eekd.sub.10kke 2 24322 24337 2016 548556 n/a n/a ATGCATGCCTACCCAA eekd.sub.10kke 43 24526 24541 2017 548557 n/a n/a GTTACTCTGTGATCCA eekd.sub.10kke 81 24581 24596 2018 548558 n/a n/a AACATTGTGTAGCTGC eekd.sub.10kke 75 24640 24655 2019 548559 n/a n/a GAGACTGAAGCCCTCA eekd.sub.10kke 44 24676 24691 2020 548560 n/a n/a CACTGCCTAGAAAGGC eekd.sub.10kke 16 24734 24749 2021 548561 n/a n/a TGTAGTATCCAGAGTA eekd.sub.10kke 46 24930 24945 2022 548562 n/a n/a AGATGACCTGCAGATG eekd.sub.10kke 50 24983 24998 2023 548563 n/a n/a AAACCATGAATTAGGT eekd.sub.10kke 20 25100 25115 2024 548564 n/a n/a TTGCTACTTTACACCA eekd.sub.10kke 69 25208 25223 2025 548565 n/a n/a GGCATTAGGATAGGCA eekd.sub.10kke 63 25350 25365 2026 548566 n/a n/a CACTCAGACTGTCTGA eekd.sub.10kke 0 25413 25428 2027 548567 n/a n/a AGATCCGGAATAACCA eekd.sub.10kke 67 25459 25474 2028 548568 n/a n/a ATTGACAACCATCCTA eekd.sub.10kke 27 25496 25511 2029 548569 n/a n/a ACTCATTGGTCTACAG eekd.sub.10kke 41 25559 25574 2030 548570 n/a n/a ATGCCTTGTGCCTATT eekd.sub.10kke 74 25706 25721 2031 548571 n/a n/a ACTCTGAGGCCTTAGG eekd.sub.10kke 59 25794 25809 2032 548572 n/a n/a GCATTACTCAGCATGT eekd.sub.10kke 63 25836 25851 2033 548573 n/a n/a CCAGTCACCACCATTG eekd.sub.10kke 65 25862 25877 2034 548574 n/a n/a GGTCTAACTCTAAGGG eekd.sub.10kke 0 25920 25935 2035 548575 n/a n/a TGTCCTTTAAAGTATC eekd.sub.10kke 18 25971 25986 2036 548576 n/a n/a TCATGTGGCAACCTGT eekd.sub.10kke 41 26114 26129 2037 548577 n/a n/a AATCTGCACCTGGCAG eekd.sub.10kke 42 26428 26443 2038 548578 n/a n/a CATGGCTATTGCTTCC eekd.sub.10kke 73 26513 26528 2039 548579 n/a n/a GGGCTATATTGCCAGC eekd.sub.10kke 46 26614 26629 2040 548580 n/a n/a CCAGAGCCTTGATCAG eekd.sub.10kke 36 26681 26696 2041 548581 n/a n/a GGTGGGTTATCTGAGA eekd.sub.10kke 13 26710 26725 2042 548582 n/a n/a TAGCTCCATGCTGTGT eekd.sub.10kke 59 26735 26750 2043 548583 n/a n/a GGGAATTTATGCTGCC eekd.sub.10kke 79 26782 26797 2044 548584 n/a n/a TGATGAAGTTCCACCT eekd.sub.10kke 47 26840 26855 2045 548585 n/a n/a TAGGCACAGACAACCT eekd.sub.10kke 33 26869 26884 2046 548586 n/a n/a TCCAACTACAGGACTC eekd.sub.10kke 39 26943 26958 2047 548587 n/a n/a TTCTGGGAAACTCTCT eekd.sub.10kke 45 26969 26984 2048 548588 n/a n/a AGCTCACACCCAAAAA eekd.sub.10kke 10 27006 27021 2049 548589 n/a n/a TCTGTTACCTTGAGGA eekd.sub.10kke 40 27280 27295 2050 548590 n/a n/a TGGTCATGTCAACTGT eekd.sub.10kke 35 27550 27565 2051 548591 n/a n/a GTAAGCCTTCACAGGG eekd.sub.10kke 3 27583 27598 2052 548592 n/a n/a CTCACCAGAGTTGTCC eekd.sub.10kke 7 27726 27741 2053 548593 n/a n/a CATCCCTGACAGGTCC eekd.sub.10kke 61 27759 27774 2054 548594 n/a n/a CCCTTCTAACCAAGGA eekd.sub.10kke 30 27825 27840 2055 548595 n/a n/a GGATGAGATGCATCCA eekd.sub.10kke 8 28069 28084 2056 548596 n/a n/a ATGGCGGTGAAGCAGC eekd.sub.10kke 20 28127 28142 2057 548597 n/a n/a TGAATACCATCCCCGC eekd.sub.10kke 50 28171 28186 2058 548598 n/a n/a GCGCCATCTGCCCTGT eekd.sub.10kke 50 28253 28268 2059 548599 n/a n/a TGGGTTGGAGGAGTGG eekd.sub.10kke 19 28311 28326 2060 548600 n/a n/a TGGTGGTGGGATTGGT eekd.sub.10kke 53 28336 28351 2061 28391 28406 28434 28449 28446 28461 28525 28540 28611 28626 28623 28638 548601 n/a n/a TTGGTGGTGGGATTGG eekd.sub.10kke 18 28337 28352 2062 28392 28407 28435 28450 28447 28462 28526 28541 28612 28627 28624 28639 548602 n/a n/a GGTGGTGGAATTGGTG eekd.sub.10kke 20 28347 28362 2063 548603 n/a n/a GAGATTGGTGGTGGGT eekd.sub.10kke 35 28372 28387 2064 548604 n/a n/a GTGGTGGGATTGGTGC eekd.sub.10kke 22 28432 28447 2065 548605 n/a n/a TGGCGGGATTGGTGGT eekd.sub.10kke 12 28479 28494 2066 28558 28573 548606 n/a n/a CGGTGGTGGGATTGGT eekd.sub.10kke 41 28501 28516 2067 28580 28595 548607 n/a n/a TCGGTGGTGGGATTGG eekd.sub.10kke 34 28502 28517 2068 28581 28596 548608 n/a n/a ATCGGTGGTGGGATTG eekd.sub.10kke 25 28503 28518 2069 28582 28597 548609 n/a n/a GATCGGTGGTGGGATT eekd.sub.10kke 30 28504 28519 2070 28583 28598 548610 n/a n/a GGATCGGTGGTGGGAT eekd.sub.10kke 2 28505 28520 2071 28584 28599 548611 n/a n/a GCGGGATCGGTGGTGG eekd.sub.10kke 7 28508 28523 2072 28587 28602 548612 n/a n/a GGCGGGATCGGTGGTG eekd.sub.10kke 20 28509 28524 2073 28588 28603
TABLE-US-00142 TABLE 151 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 ISIS Start Stop % Start Stop SEQ NO Site Site Sequence Chemistry inhibition Site Site ID NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 46 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 64 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548382 n/a n/a GAGCAAATACAGTCCA eekd.sub.10kke 19 12620 12635 2074 548383 n/a n/a GTCTCGATGGCAAGCT eekd.sub.10kke 49 12654 12669 2075 548384 n/a n/a CTCACCGGTACTCTGC eekd.sub.10kke 49 12805 12820 2076 548385 n/a n/a TCCTGGAGGCACCAAT eekd.sub.10kke 0 12847 12862 2077 548386 n/a n/a AGCCCTGTTTGGTTTT eekd.sub.10kke 0 12903 12918 2078 548387 n/a n/a TGAAGGGCGAGGCGCA eekd.sub.10kke 22 13261 13276 2079 548388 n/a n/a AAGAGGATGTCAGGCT eekd.sub.10kke 4 13357 13372 2080 548389 n/a n/a TTGAGGAAAGACCTGC eekd.sub.10kke 11 13399 13414 2081 548390 n/a n/a GCTGAGTGTGACTTAA eekd.sub.10kke 43 13455 13470 2082 548391 n/a n/a GTACATGACTCCAGTG eekd.sub.10kke 34 13638 13653 2083 548392 n/a n/a GTAGAGCATGGAGCGA eekd.sub.10kke 31 13730 13745 2084 548393 n/a n/a CGCTTCAGGAAAGCGA eekd.sub.10kke 26 13828 13843 2085 548394 n/a n/a GGCAGGAGACTCCGTG eekd.sub.10kke 25 13919 13934 2086 548395 n/a n/a ATCCTTCCCCTCGCAA eekd.sub.10kke 0 13966 13981 2087 548396 n/a n/a TAATGAGTGGGTTAGG eekd.sub.10kke 0 14007 14022 2088 548397 n/a n/a GGAGCAGTGCAGGTAA eekd.sub.10kke 1 14065 14080 2089 548398 n/a n/a ATAGGCAATTGTTCCT eekd.sub.10kke 55 14129 14144 2090 548399 n/a n/a AGTCCTACAATTACCA eekd.sub.10kke 11 14239 14254 2091 548400 n/a n/a GGGCTCCTATTCCACC eekd.sub.10kke 13 14277 14292 2092 548401 n/a n/a GCCAGCTATGGGAACA eekd.sub.10kke 71 14333 14348 2093 548402 n/a n/a CCCCATCTCGAAGCCC eekd.sub.10kke 45 14380 14395 2094 548403 n/a n/a GAGTACATTGGGCCCA eekd.sub.10kke 25 14418 14433 2095 548404 n/a n/a GAGCCTTCCGCCTCTC eekd.sub.10kke 37 14471 14486 2096 548405 n/a n/a CGGACCTTCATCTTCA eekd.sub.10kke 35 14529 14544 2097 548406 n/a n/a TCTAGAGGCCGCCTGC eekd.sub.10kke 0 14558 14573 2098 548407 n/a n/a CCTATAACTGCTGCTC eekd.sub.10kke 24 14731 14746 2099 548408 n/a n/a TATCACTGTACTAGTT eekd.sub.10kke 47 14748 14763 1269 14819 14834 14890 14905 14949 14964 15009 15024 15081 15096 15153 15168 15224 15239 15296 15311 15355 15370 15415 15430 15487 15502 15559 15574 15617 15632 15689 15704 15819 15834 15891 15906 15949 15964 548409 n/a n/a GTATCACTGTACTAGT eekd.sub.10kke 81 14749 14764 2100 14820 14835 14891 14906 14950 14965 15010 15025 15082 15097 15154 15169 15225 15240 15297 15312 15356 15371 15416 15431 15488 15503 15560 15575 15618 15633 15690 15705 15820 15835 15892 15907 15950 15965 548410 n/a n/a AGTATCACTGTACTAG eekd.sub.10kke 85 14750 14765 2101 14821 14836 14892 14907 14951 14966 15011 15026 15083 15098 15155 15170 15226 15241 15298 15313 15357 15372 15417 15432 15489 15504 15561 15576 15619 15634 15691 15706 15821 15836 15893 15908 15951 15966 548411 n/a n/a CAGTATCACTGTACTA eekd.sub.10kke 72 14751 14766 2102 14822 14837 14893 14908 14952 14967 15012 15027 15084 15099 15156 15171 15227 15242 15299 15314 15358 15373 15418 15433 15490 15505 15562 15577 15620 15635 15692 15707 15822 15837 15894 15909 15952 15967 548412 n/a n/a TAACAGTATCACTGTA eekd.sub.10kke 17 14754 14769 2103 14825 14840 14896 14911 14955 14970 15015 15030 15087 15102 15159 15174 15230 15245 15302 15317 15361 15376 15421 15436 15493 15508 15565 15580 15623 15638 15695 15710 15825 15840 15897 15912 15955 15970 548413 n/a n/a CTAACAGTATCACTGT eekd.sub.10kke 55 14755 14770 2104 14826 14841 14897 14912 15016 15031 15088 15103 15231 15246 15303 15318 15422 15437 15494 15509 15624 15639 15826 15841 15956 15971 548414 n/a n/a TCTAACAGTATCACTG eekd.sub.10kke 20 14756 14771 2105 14827 14842 14898 14913 15017 15032 15089 15104 15232 15247 15304 15319 15423 15438 15495 15510 15625 15640 15827 15842 15957 15972 548415 n/a n/a ATAACTCTAACAGTAT eekd.sub.10kke 0 14761 14776 2106 14832 14847 14903 14918 15022 15037 15094 15109 15237 15252 15309 15324 15428 15443 15500 15515 15630 15645 15832 15847 15962 15977 548416 n/a n/a CTATAACTCTAACAGT eekd.sub.10kke 9 14763 14778 2107 14834 14849 14905 14920 15024 15039 15096 15111 15239 15254 15311 15326 15430 15445 15502 15517 15632 15647 15834 15849 15964 15979 548417 n/a n/a ACTGTCCTATAACTCT eekd.sub.10kke 24 14769 14784 2108 14840 14855 548418 n/a n/a TATATCACTGTCCTAT eekd.sub.10kke 39 14775 14790 2109 14846 14861
15180 15195 15716 15731 16164 16179 548419 n/a n/a CCTATATCACTGTCCT eekd.sub.10kke 52 14777 14792 2110 14848 14863 15182 15197 15718 15733 548420 n/a n/a TCCTATATCACTGTCC eekd.sub.10kke 58 14778 14793 2111 14849 14864 15183 15198 15719 15734 548421 n/a n/a CACTGTCCTATATCAC eekd.sub.10kke 56 14783 14798 2112 14854 14869 14979 14994 15117 15132 15188 15203 15260 15275 15385 15400 15523 15538 15653 15668 15724 15739 15855 15870 15985 16000 548422 n/a n/a GTATCACTGTCCTATA eekd.sub.10kke 69 14787 14802 2113 14983 14998 15121 15136 15389 15404 15527 15542 15989 16004 548423 n/a n/a AGTATCACTGTCCTAT eekd.sub.10kke 72 14788 14803 2114 14984 14999 15050 15065 15122 15137 15390 15405 15456 15471 15528 15543 15990 16005 548424 n/a n/a CAGTATCACTGTCCTA eekd.sub.10kke 90 14789 14804 2115 14985 15000 15051 15066 15123 15138 15391 15406 15457 15472 15529 15544 15991 16006 548425 n/a n/a AACAGTATCACTGTCC eekd.sub.10kke 90 14791 14806 2116 14987 15002 15053 15068 15125 15140 15393 15408 15459 15474 15531 15546 15993 16008 548426 n/a n/a TATAACAGTATCACTG eekd.sub.10kke 14 14794 14809 2117 14990 15005 15056 15071 15128 15143 15161 15176 15363 15378 15396 15411 15462 15477 15534 15549 15567 15582 15697 15712 15899 15914 15996 16011 548427 n/a n/a CTATAACAGTATCACT eekd.sub.10kke 24 14795 14810 2118 14991 15006 15057 15072 15129 15144 15162 15177 15364 15379 15397 15412 15463 15478 15535 15550 15568 15583 15698 15713 15900 15915 15997 16012 548428 n/a n/a TAACTATAACAGTATC eekd.sub.10kke 0 14798 14813 2119 15060 15075 15132 15147 15165 15180 15466 15481 15538 15553 15571 15586 15701 15716 15772 15787 16000 16015 548429 n/a n/a TATAACTATAACAGTA eekd.sub.10kke 0 14800 14815 2120 15062 15077 15134 15149 15167 15182 15468 15483 15540 15555 15573 15588 15703 15718 15774 15789 16002 16017 548430 n/a n/a CCTATAACTATAACAG eekd.sub.10kke 21 14802 14817 2121 15064 15079 15169 15184 15470 15485 15542 15557 15575 15590 15705 15720 15776 15791 16004 16019 548431 n/a n/a TACCTATAACTCTAAC eekd.sub.10kke 9 14908 14923 2122 15027 15042 15099 15114 15242 15257 15314 15329 15433 15448 15505 15520 15635 15650 15837 15852 15967 15982 548432 n/a n/a ACTGTACCTATAACTC eekd.sub.10kke 43 14912 14927 2123 15031 15046 15246 15261 15318 15333 15437 15452 15509 15524 15639 15654 15841 15856 15971 15986 548433 n/a n/a TATCACTGTACCTATA eekd.sub.10kke 33 14916 14931 2124 15250 15265 15322 15337 15375 15390 15513 15528 15643 15658 15786 15801 15845 15860 15975 15990 16137 16152 548434 n/a n/a ACAATATCACTGTACC eekd.sub.10kke 63 14920 14935 2125 15326 15341 15790 15805 16063 16078 16141 16156 548435 n/a n/a AACAATATCACTGTAC eekd.sub.10kke 19 14921 14936 2126 15327 15342 15791 15806 16064 16079 16142 16157 548436 n/a n/a ATATCACTGTACCTGT eekd.sub.10kke 8 14970 14985 2127 548437 n/a n/a TATATCACTGTACCTG eekd.sub.10kke 74 14971 14986 2128 548438 n/a n/a CTATATCACTGTACCT eekd.sub.10kke 38 14972 14987 2129 15253 15268 15378 15393 15516 15531 15646 15661 15848 15863 15978 15993 548439 n/a n/a CCTATATCACTGTACC eekd.sub.10kke 46 14973 14988 2130 15254 15269 15379 15394 15517 15532 15647 15662 15849 15864 15979 15994 548440 n/a n/a CCTATAACAGTATCAC eekd.sub.10kke 32 14992 15007 2131 15365 15380 15398 15413 548441 n/a n/a TCCTATAACAGTATCA eekd.sub.10kke 42 14993 15008 2132 15399 15414 548442 n/a n/a TTCCTATAACAGTATC eekd.sub.10kke 17 14994 15009 2133 15400 15415 548443 n/a n/a GTTTCCTATAACAGTA eekd.sub.10kke 12 14996 15011 2134 15402 15417 548444 n/a n/a CTATGTCACTGTACCT eekd.sub.10kke 43 15038 15053 2135 15444 15459 548445 n/a n/a CCTATGTCACTGTACC eekd.sub.10kke 62 15039 15054 2136 15445 15460 548446 n/a n/a TCCTATGTCACTGTAC eekd.sub.10kke 16 15040 15055 2137 15446 15461 548447 n/a n/a CACTGTCCTATGTCAC eekd.sub.10kke 59 15045 15060 2138 15451 15466 548448 n/a n/a TCACTGTCCTATGTCA eekd.sub.10kke 61 15046 15061 2139 15452 15467 548449 n/a n/a ATCACTGTCCTATGTC eekd.sub.10kke 62 15047 15062 2140 15453 15468 548450 n/a n/a CTACCTATAACTCTAA eekd.sub.10kke 0 15100 15115 2141 548451 n/a n/a GTCCTATAACTATAAC eekd.sub.10kke 0 15171 15186 2142 15577 15592 15707 15722 16006 16021 16077 16092 16102 16117 16155 16170 548452 n/a n/a TATATCACTGTACCTA eekd.sub.10kke 65 15252 15267 2143 15377 15392 15515 15530 15645 15660 15847 15862 15977 15992 548453 n/a n/a TACCTATAACAGTATC eekd.sub.10kke 12 15367 15382 2144 548454 n/a n/a ACTGTACCTATAACAG eekd.sub.10kke 17 15371 15386 2145 548455 n/a n/a CACCGTACTAGTTTCC eekd.sub.10kke 64 15757 15772 2146 548456 n/a n/a TATAACAGTATCACCG eekd.sub.10kke 52 15768 15783 2147 548457 n/a n/a CTATAACAGTATCACC eekd.sub.10kke 13 15769 15784 2148 548458 n/a n/a ACCTATAACTATAACA eekd.sub.10kke 0 15777 15792 2149 16249 16264
TABLE-US-00143 TABLE 152 SEQ SEQ SEQ SEQ ID ID ID ID NO: NO: NO: 1 NO: 1 10 10 SEQ ISIS Start Stop % Start Stop ID NO Site Site Sequence Chemistry inhibition Site Site NO 531231 n/a n/a TATCACTGTACTAGTTTCCT eeeeed.sub.10eeeee 48 14744 14763 334 14815 14834 14886 14905 14945 14964 15005 15024 15077 15096 15220 15239 15292 15311 15351 15370 15411 15430 15483 15502 15555 15574 15613 15632 15685 15704 15815 15834 15887 15906 15945 15964 547747 n/a n/a TCACTGTACTAGTTTC eekd.sub.10kke 88 14746 14761 1267 14817 14832 14888 14903 14947 14962 15007 15022 15079 15094 15222 15237 15294 15309 15353 15368 15413 15428 15485 15500 15557 15572 15615 15630 15687 15702 15817 15832 15889 15904 15947 15962 548613 n/a n/a TGGCGGGATCGGTGGT eekd.sub.10kke 39 28510 28525 2150 28589 28604 548614 n/a n/a TGGTGGCGGGATCGGT eekd.sub.10kke 0 28513 28528 2151 28592 28607 548615 n/a n/a TTGGTGGCGGGATCGG eekd.sub.10kke 10 28514 28529 2152 28593 28608 548616 n/a n/a ATTGGTGGCGGGATCG eekd.sub.10kke 35 28515 28530 2153 548617 n/a n/a GATTGGTGGCGGGATC eekd.sub.10kke 44 28516 28531 2154 548618 n/a n/a GTTGGTGGCGGGATCG eekd.sub.10kke 18 28594 28609 2155 548619 n/a n/a GGTTGGTGGCGGGATC eekd.sub.10kke 19 28595 28610 2156 548620 n/a n/a TGGTTGGTGGCGGGAT eekd.sub.10kke 24 28596 28611 2157 548621 n/a n/a GAACACATCAGGGATT eekd.sub.10kke 33 28638 28653 2158 548622 n/a n/a TTTCTATGGGCCTGAC eekd.sub.10kke 0 28669 28684 2159 548623 n/a n/a GCTGTCACTTAAGCCA eekd.sub.10kke 16 28862 28877 2160 548624 n/a n/a TCTAGGGCCACACCTC eekd.sub.10kke 24 28892 28907 2161 548625 n/a n/a GTTCTACACACAGTAC eekd.sub.10kke 0 29014 29029 2162 548626 n/a n/a GCAGTATGTTCAATCC eekd.sub.10kke 36 29202 29217 2163 548627 n/a n/a CCCACATGTACCACCG eekd.sub.10kke 22 29235 29250 2164 548628 n/a n/a GTATGGCAGAGCCCCT eekd.sub.10kke 9 29285 29300 2165 548629 n/a n/a CCCATCTTGGGACTTT eekd.sub.10kke 44 29341 29356 2166 548630 n/a n/a TGGTCCCAAATTGGAG eekd.sub.10kke 33 29387 29402 2167 548631 n/a n/a CTCACAATACTGAGCC eekd.sub.10kke 55 29421 29436 2168 548632 n/a n/a GGAGATATCAGGTGCA eekd.sub.10kke 45 29499 29514 2169 548633 n/a n/a CAAGGCATGTGTGCAC eekd.sub.10kke 41 29534 29549 2170 548634 n/a n/a GCCTTATTCTGTGCAA eekd.sub.10kke 0 29583 29598 2171 548635 n/a n/a AGGTGTGGCGCGCGCC eekd.sub.10kke 18 29853 29868 2172 548636 n/a n/a CTCTATACAGCTGGGC eekd.sub.10kke 5 30000 30015 2173 548637 n/a n/a GCTGATCTTCTAATGC eekd.sub.10kke 38 30063 30078 2174 548638 n/a n/a CCTCATTGCTCCACTA eekd.sub.10kke 26 30103 30118 2175 548639 n/a n/a TGGGAAGAAACTAGCA eekd.sub.10kke 10 30159 30174 2176 548640 n/a n/a GAATGTTGCTGTCCCA eekd.sub.10kke 32 30194 30209 2177 548641 n/a n/a GCATCATGCTTACTGC eekd.sub.10kke 23 30225 30240 2178 548642 n/a n/a GCGGCAGTAGTGAATC eekd.sub.10kke 23 30288 30303 2179 548643 n/a n/a CCTACCTAATTCCTCC eekd.sub.10kke 0 30329 30344 2180 548644 n/a n/a AACTGGGCAGTCCTTC eekd.sub.10kke 14 30418 30433 2181 548645 n/a n/a CCAGCGCAATTCTGCT eekd.sub.10kke 8 30666 30681 2182 548646 n/a n/a CGTTTCCCTCAACTCC eekd.sub.10kke 24 30750 30765 2183 548647 n/a n/a CACGGCAAGTCGCGGG eekd.sub.10kke 39 30790 30805 2184 548648 n/a n/a CAGTTGTATCCCTCCC eekd.sub.10kke 32 30852 30867 2185 548649 n/a n/a GCCTCTCAGACGGCAC eekd.sub.10kke 0 30906 30921 2186 548650 n/a n/a CTGATCCCACTTGCCC eekd.sub.10kke 21 30991 31006 2187 548651 n/a n/a AGTCTCTTTCCTACCC eekd.sub.10kke 61 31030 31045 2188 548652 n/a n/a CCACGATGCTCTGGCC eekd.sub.10kke 65 31068 31083 2189 548653 n/a n/a TCGGCTCCTGGCAGCA eekd.sub.10kke 46 31111 31126 2190 548654 n/a n/a ACCATTCCTGACCATG eekd.sub.10kke 34 31151 31166 2191 548655 n/a n/a CCCGAGGTCACATAAT eekd.sub.10kke 56 31416 31431 2192 548656 n/a n/a TTACAACAGACCCAGG eekd.sub.10kke 35 31497 31512 2193 548657 n/a n/a AGCAGGGTATCTTCAC eekd.sub.10kke 26 31548 31563 2194 548658 n/a n/a GAAGTTCCTGTGTCTT eekd.sub.10kke 11 31593 31608 2195 548659 n/a n/a CCAACCTCTAAGGCTA eekd.sub.10kke 17 31721 31736 2196 548660 n/a n/a ATGCTTACCTTTCTCC eekd.sub.10kke 0 31955 31970 2197 548661 n/a n/a ACGACCCACTCCATGT eekd.sub.10kke 18 32016 32031 2198 548662 n/a n/a TGCTTAAAAGTCTCCC eekd.sub.10kke 5 32155 32170 2199 548663 n/a n/a GCCCTAGAAGGGCCCA eekd.sub.10kke 20 32219 32234 2200 548664 n/a n/a GCGGGTGGTCTTGCAC eekd.sub.10kke 38 32245 32260 2201 548665 n/a n/a GCTCCCGGCCATTAGC eekd.sub.10kke 8 32312 32327 2202 548666 n/a n/a TCTCCATAGTGAGACG eekd.sub.10kke 1 32342 32357 2203 548667 n/a n/a TGGCAAGCTACCTTCT eekd.sub.10kke 51 32384 32399 2204 548668 n/a n/a GGGAGCTTTCATGGCT eekd.sub.10kke 68 32506 32521 2205 548669 n/a n/a AATGCAGGCCAGCATC eekd.sub.10kke 42 32541 32556 2206 548670 n/a n/a GAAAAGCCCTCCGAGC eekd.sub.10kke 15 32590 32605 2207 548671 n/a n/a CAACAATCCAAAGCCT eekd.sub.10kke 3 32674 32689 2208 548672 n/a n/a CCCCCCAGAAATCCCA eekd.sub.10kke 40 32708 32723 2209 548673 n/a n/a GACCTTGCTTCCATGT eekd.sub.10kke 40 32753 32768 2210 548674 n/a n/a GAGAGACGGCACCCTG eekd.sub.10kke 4 32829 32844 2211 548675 n/a n/a GGGAAGGTAGTGTTAC eekd.sub.10kke 8 32898 32913 2212 548676 n/a n/a GTGAATCAGAGCAGTG eekd.sub.10kke 63 32963 32978 2213 548677 n/a n/a TCACCTGTGAGTAACC eekd.sub.10kke 40 33089 33104 2214 548678 n/a n/a GAGTTACCTTACAAGC eekd.sub.10kke 22 33232 33247 2215 548679 n/a n/a TCTCAAGCAGCCTATT eekd.sub.10kke 0 33267 33282 2216 548680 n/a n/a GCCCCTCTTAAATAGC eekd.sub.10kke 9 33446 33461 2217 548681 n/a n/a GATATCATCATCCCAA eekd.sub.10kke 22 33513 33528 2218 548682 n/a n/a GTATCCCCTTTTCTAT eekd.sub.10kke 0 33556 33571 2219 548683 n/a n/a AGTATCTCATGTGCCT eekd.sub.10kke 46 33581 33596 2220 548684 n/a n/a CAAGACCTTGCTTGCC eekd.sub.10kke 24 33658 33673 2221 548685 n/a n/a TAGTCCACTACACAGC eekd.sub.10kke 24 33802 33817 2222 548686 n/a n/a ACGACAATGGGATTCA eekd.sub.10kke 0 33844 33859 2223 548687 n/a n/a GAATCTCCCTGAGTCA eekd.sub.10kke 20 33888 33903 2224 548688 n/a n/a TAGAGGGATCCCAGGA eekd.sub.10kke 0 34416 34431 2225 548689 n/a n/a CCAGGTGCAGCACGGA eekd.sub.10kke 12 34483 34498 2226
Example 117: Dose-Dependent Antisense Inhibition of Human PKK in HepaRG.TM. Cells
[1070] Gapmers from the studies described above exhibiting significant in vitro inhibition of PKK mRNA were selected and tested at various doses in HepaRG.TM. cells. Cells were plated at a density of 20,000 cells per well and transfected using electroporation with 0.12 M, 0.37 M, 1.11 M, 3.33 M, and 10.00 M concentrations of antisense oligonucleotide. After a treatment period of approximately 16 hours, RNA was isolated from the cells and PKK mRNA levels were measured by quantitative real-time PCR. Human PKK primer probe set RTS3454 was used to measure mRNA levels. PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN. The antisense oligonucleotides were tested in a series of experiments that had similar culture conditions. The results for each experiment are presented in separate tables shown below. Results are presented as percent inhibition of PKK, relative to untreated control cells.
[1071] The half maximal inhibitory concentration (IC.sub.50) of each oligonucleotide is also presented. PKK mRNA levels were significantly reduced in a dose-dependent manner in antisense oligonucleotide treated cells.
TABLE-US-00144 TABLE 153 ISIS 0.12 0.37 1.11 3.33 10.00 IC.sub.50 No .mu.M .mu.M .mu.M .mu.M .mu.M (.mu.M) 486847 0 34 48 71 87 1.1 530933 15 13 42 67 66 1.7 530959 12 27 53 80 94 0.9 530965 8 5 63 83 91 0.8 530967 30 36 48 82 91 0.7 530970 1 0 66 76 84 1.0 530971 12 40 52 66 70 1.3 530988 0 25 54 86 78 0.9 530992 0 50 63 83 80 0.7 531002 6 28 58 82 86 0.9 531004 0 14 25 71 84 2.1 531005 14 28 61 73 77 0.9 531022 0 0 32 62 77 2.2 531078 10 27 54 69 92 1.1 531231 23 30 76 89 94 0.6
TABLE-US-00145 TABLE 154 ISIS 0.12 0.37 1.11 3.33 10.00 IC.sub.50 No .mu.M .mu.M .mu.M .mu.M .mu.M (.mu.M) 531026 23 26 49 75 85 1.0 531055 3 28 64 76 81 0.9 531069 19 39 48 76 83 0.9 531071 23 37 56 83 83 0.7 531110 14 29 49 76 85 1.1 531121 0 13 47 69 79 1.5 531123 14 43 51 71 64 0.9 531172 0 16 37 60 60 2.1 531198 0 35 62 76 60 0.8 531231 18 0 36 76 84 2.0 531232 15 26 40 62 76 1.7 531233 17 27 50 77 84 1.0 531234 24 21 47 72 82 1.4 531235 27 55 62 84 95 0.4 531236 4 28 59 85 93 0.8
Example 118: Dose-Dependent Antisense Inhibition of Human PKK in HepaRG.TM. Cells
[1072] Gapmers from the studies described above exhibiting significant in vitro inhibition of PKK mRNA were selected and tested at various doses in HepaRG.TM. cells. Cells were plated at a density of 20,000 cells per well and transfected using electroporation with 0.19 M, 0.56 M, 1.67 M, and 5.00 M concentrations of antisense oligonucleotide. After a treatment period of approximately 16 hours, RNA was isolated from the cells and PKK mRNA levels were measured by quantitative real-time PCR. Human PKK primer probe set RTS3454 was used to measure mRNA levels. PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.RTM.. The antisense oligonucleotides were tested in a series of experiments that had similar culture conditions. The results for each experiment are presented in separate tables shown below. Results are presented as percent inhibition of PKK, relative to untreated control cells. `n/a` indicates that there was no measurement done for that particular antisense oligonucleotide for that particular dose.
[1073] The half maximal inhibitory concentration (IC.sub.50) of each oligonucleotide is also presented. PKK mRNA levels were significantly reduced in a dose-dependent manner in antisense oligonucleotide treated cells.
TABLE-US-00146 TABLE 155 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M .mu.M 531231 32 30 73 89 0.5 546158 5 45 79 83 0.7 546188 36 55 81 83 0.4 546253 1 13 46 81 1.7 546254 51 66 80 91 0.2 546343 28 64 87 87 0.4 546825 46 73 86 88 0.2 546827 32 70 84 90 0.3 546828 39 58 87 93 0.3 546829 3 30 73 88 1.0 546846 36 45 71 82 0.5 547413 0 0 41 83 2.2 547423 37 50 92 90 0.4 547445 41 75 82 88 0.2 547456 12 67 66 80 1.0 547464 21 52 67 97 0.6 547564 51 48 82 90 0.2 547587 20 62 84 86 0.5 548758 41 47 82 94 0.4
TABLE-US-00147 TABLE 156 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M .mu.M 531231 25 34 84 92 0.7 546190 33 65 86 n/a 0.4 546208 16 45 79 91 0.7 546216 62 69 88 88 0.1 546255 32 35 78 87 0.5 546268 56 50 82 93 0.1 546301 25 50 53 87 0.8 546849 23 35 83 91 0.7 546852 19 40 78 85 0.8 546889 23 54 78 91 0.6 546916 43 71 79 89 0.2 546967 20 39 76 71 0.7 547273 44 69 87 87 0.2 547276 35 44 71 77 0.6 547335 8 52 85 92 0.7 547340 46 79 88 n/a 0.2 547602 18 53 92 87 0.5 547647 1 70 72 n/a 0.8 547694 0 29 67 90 1.2
TABLE-US-00148 TABLE 157 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 58 64 77 98 0.1 546247 0 29 71 88 1.1 546251 31 60 99 89 0.5 546753 28 47 83 96 0.5 546826 17 40 87 97 0.7 546833 8 33 74 94 0.9 546854 23 39 83 94 0.6 546894 15 47 50 93 0.9 546897 40 56 71 95 0.4 546903 15 37 74 98 0.8 546986 31 49 77 89 0.5 547293 53 57 80 86 0.2 547298 32 61 74 90 0.4 547364 38 47 54 89 0.6 547373 20 7 49 86 1.1 547426 19 50 84 93 0.6 547454 19 40 58 92 0.9 547617 52 66 77 93 0.2 548770 26 54 77 91 0.5
TABLE-US-00149 TABLE 158 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 34 47 72 n/a 0.5 546214 0 0 68 85 1.3 546304 0 6 51 71 2.1 546739 35 55 57 79 0.6 546832 19 38 70 95 0.8 546847 39 57 75 89 0.4 546855 18 7 30 82 2.2 546877 0 19 75 80 1.3 546939 1 66 86 90 0.6 547349 0 8 66 76 1.6 547360 8 27 76 76 0.8 547368 0 0 31 80 2.5 547483 0 9 49 71 2.1 547575 0 34 82 93 1.1 547618 0 0 73 98 1.3 547622 0 47 79 90 0.9 547637 10 21 36 82 1.8 547731 0 0 17 56 5.0 548752 0 0 51 90 1.9
TABLE-US-00150 TABLE 159 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 21 45 67 96 0.7 546195 34 51 79 92 0.5 546198 7 3 45 92 1.3 546287 0 15 39 89 1.7 546358 0 19 71 80 1.3 546403 0 20 37 41 >5.0 546410 13 43 52 75 1.2 546412 0 1 61 62 2.3 546429 6 10 44 69 2.3 546834 1 0 30 83 2.3 547006 0 0 54 77 1.5 547294 28 59 87 86 0.4 547337 23 41 55 79 1.0 547514 18 8 51 80 1.9 547584 26 34 76 86 0.7 547585 42 57 70 95 0.4 547615 20 26 41 84 1.4 547636 0 24 79 94 1.1 548744 14 35 63 83 1.0
TABLE-US-00151 TABLE 160 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 21 39 90 97 0.6 546232 49 50 94 97 0.2 546248 25 66 87 93 0.4 546835 9 35 68 93 0.9 546848 0 18 91 97 1.0 546853 47 64 84 n/a 0.2 546870 35 42 80 95 0.5 546872 32 33 82 94 0.4 546876 0 50 85 95 0.8 547275 34 66 82 95 0.3 547341 36 58 91 95 0.3 547366 0 45 68 91 1.2 547453 25 40 54 92 0.8 547457 41 65 80 85 0.3 547616 26 50 72 89 0.6 547632 44 47 81 97 0.6 547633 12 46 78 n/a 0.7 547718 36 12 69 74 1.6 548757 18 49 82 93 0.6
TABLE-US-00152 TABLE 161 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 6 38 74 95 0.8 546291 22 32 34 72 2.0 546310 0 36 56 80 1.3 546896 0 45 82 97 0.8 546980 0 18 29 80 2.2 547009 0 9 21 63 3.6 547019 0 6 54 86 1.6 547277 2 32 34 62 2.8 547288 0 0 0 38 >5.0 547374 0 15 24 44 >5.0 547493 0 26 64 77 1.3 547520 0 25 66 64 1.1 547712 0 5 21 62 3.8 547722 0 15 32 73 2.4 547728 0 2 16 61 4.4 547780 0 10 36 55 3.9 548743 25 57 73 88 0.5 548753 0 23 49 84 1.5 548756 0 4 16 86 >5.0
TABLE-US-00153 TABLE 162 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 25 55 89 97 0.5 546188 27 69 88 97 0.4 546216 23 78 95 98 <0.2 546254 40 63 84 95 0.3 546268 0 71 92 92 0.5 546343 37 32 83 95 0.4 546825 38 82 n/a 99 0.2 546827 23 74 98 96 0.4 546828 0 64 89 97 0.2 546846 26 49 85 n/a 0.5 546967 22 45 74 92 0.7 547273 0 60 82 83 0.6 547340 34 84 96 n/a 0.3 547423 78 92 n/a n/a <0.2 547445 80 87 98 91 <0.2 547564 46 66 90 97 0.2 547587 38 64 91 97 0.3 547602 1 9 52 93 1.4 548758 0 72 79 n/a 0.6
TABLE-US-00154 TABLE 163 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 7 39 56 97 1.0 546190 21 34 76 98 0.7 546208 5 33 70 97 0.9 546251 19 45 91 97 0.6 546255 5 39 82 96 0.8 546739 4 62 84 86 0.6 546753 17 31 70 91 0.9 546849 13 45 84 98 0.7 546889 25 9 73 92 1.4 546897 16 17 69 97 0.8 546916 0 27 73 97 1.0 546986 7 28 69 86 1.1 547276 6 3 53 68 2.2 547293 0 45 65 70 1.3 547298 0 12 67 87 1.7 547335 0 13 73 95 1.3 547426 18 35 80 95 0.7 547617 17 37 79 98 0.7 548770 9 0 61 92 1.7
TABLE-US-00155 TABLE 164 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 531231 6 56 68 97 0.8 546195 0 27 91 94 0.9 546232 0 74 95 96 0.2 546248 0 59 73 89 0.8 546832 36 49 85 97 0.4 546847 14 44 83 95 0.7 546853 4 49 74 92 0.8 546870 36 34 61 91 1.0 546872 42 13 59 99 1.4 546896 35 60 83 n/a 0.4 546939 16 71 96 95 0.4 547275 56 16 80 86 1.2 547294 4 70 84 91 0.6 547341 45 44 81 95 0.6 547457 33 42 70 83 0.6 547584 0 21 64 92 1.3 547585 0 46 89 93 0.8 547632 0 0 63 91 1.6 548743 22 47 74 96 0.6
Example 119: Dose-Dependent Antisense Inhibition of Human PKK in HepaRG.TM. Cells
[1074] Gapmers from the studies described above exhibiting significant in vitro inhibition of PKK mRNA were selected and tested at various doses in HepaRG.TM. cells. Cells were plated at a density of 20,000 cells per well and transfected using electroporation with 0.11 M, 0.33 M, 1.00 M, and 3.00 M concentrations of antisense oligonucleotide. After a treatment period of approximately 16 hours, RNA was isolated from the cells and PKK mRNA levels were measured by quantitative real-time PCR. Human PKK primer probe set RTS3454 was used to measure mRNA levels. PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN. The antisense oligonucleotides were tested in a series of experiments that had similar culture conditions. The results for each experiment are presented in separate tables shown below. Results are presented as percent inhibition of PKK, relative to untreated control cells. `n/a` indicates that there was no measurement done for that particular antisense oligonucleotide for that particular dose.
[1075] The half maximal inhibitory concentration (IC.sub.50) of each oligonucleotide is also presented. PKK mRNA levels were significantly reduced in a dose-dependent manner in antisense oligonucleotide treated cells.
TABLE-US-00156 TABLE 165 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 547747 24 29 81 89 0.4 547769 12 17 80 96 0.6 547824 45 73 78 n/a 0.1 547835 44 27 53 79 0.9 547843 0 52 80 91 0.4 547857 36 66 77 93 0.2 547870 0 44 80 97 0.6 547943 33 70 87 90 0.2 547946 0 47 74 n/a 0.5 547947 24 58 81 93 0.3 547998 55 73 91 91 0.1 548004 24 47 80 92 0.3 548010 0 11 49 64 1.4 548047 50 62 76 95 0.1 548147 59 94 80 n/a 0.0 548338 41 58 79 95 0.2 548348 19 46 67 91 0.4 548409 21 60 90 93 0.3 548557 5 47 82 95 0.4
TABLE-US-00157 TABLE 166 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 547747 8 61 90 92 0.3 547807 26 71 61 94 0.4 547922 67 75 81 92 0.0 547927 56 64 92 88 0.1 547948 60 80 88 97 0.0 547979 56 58 94 97 0.1 548005 53 49 71 95 0.4 548024 28 57 84 82 0.3 548043 14 60 90 92 0.3 548055 43 57 50 88 0.3 548106 53 54 82 94 0.1 548109 50 92 79 85 0.1 548155 49 50 70 81 0.3 548180 11 59 71 88 0.4 548278 3 59 78 93 0.4 548343 61 67 88 92 0.0 548558 53 61 78 95 0.1 548570 20 40 70 94 0.4 548583 43 46 93 88 0.2
TABLE-US-00158 TABLE 167 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 547747 3 44 72 90 0.5 547849 36 52 67 n/a 0.3 547851 16 46 83 n/a 0.4 547859 29 56 83 78 0.3 547862 26 71 69 n/a 0.3 547877 29 66 83 n/a 0.2 547942 25 51 91 n/a 0.3 547997 39 68 n/a 82 0.2 548046 7 35 64 77 0.7 548048 49 66 86 92 0.1 548061 26 61 59 n/a 0.4 548070 26 35 48 63 1.1 548125 33 50 81 73 0.3 548195 5 23 61 76 0.8 548265 47 69 78 67 0.1 548410 31 58 85 82 0.2 548424 17 67 86 72 0.3 548425 41 57 68 80 0.2 548547 30 41 76 90 0.4
TABLE-US-00159 TABLE 168 0.19 0.56 1.67 5.00 IC.sub.50 ISIS No .mu.M .mu.M .mu.M .mu.M (.mu.M) 547747 16 59 85 96 0.3 547808 19 22 48 71 1.1 547861 7 40 75 84 0.5 548069 6 0 27 66 1.9 548128 14 29 49 66 1.1 548170 0 8 26 65 2.0 548174 20 18 29 62 2.0 548197 33 37 51 75 0.8 548201 0 7 70 85 0.8 548217 22 24 54 71 0.9 548220 0 0 0 6 >3 548247 16 50 62 82 0.5 548422 0 32 71 93 0.7 548479 2 52 82 97 0.4 548486 20 48 77 92 0.4 548521 21 0 3 1 >3 548655 0 0 8 33 >3 548667 0 37 73 86 0.7 548668 10 30 61 84 0.7
Example 120: Efficacy of Antisense Oligonucleotides Targeting Human PKK in Transgenic Mice
[1076] Transgenic mice containing a 37,390 base pair fragment of the human KLKB1 gene sequence (chromosome 4: position 187148672-187179625, accession no: NC_000004.11) were treated with ISIS antisense oligonucleotides selected from studies described above, which were evaluated for efficacy in this model.
Treatment
[1077] Groups of transgenic mice were injected subcutaneously twice a week for 3 weeks with 2.5 mg/kg/week, 5.0 mg/kg/week, 10 mg/kg/week or 20 mg/kg/week of ISIS 546232, ISIS 546251, ISIS 546254, ISIS 546343, ISIS 546828, ISIS 547455, ISIS 547457, ISIS 547927, and ISIS 548048. One group of transgenic mice was injected subcutaneously twice a week for 3 weeks with PBS. Mice were euthanized 48 hours after the last dose, and organs and plasma were harvested for further analysis.
RNA Analysis
[1078] To evaluate the effect of ISIS oligonucleotides on target reduction, RNA was extracted from liver tissue for real-time PCR analysis of human PKK. Results are presented as percent inhibition of PKK mRNA, relative to PBS control. As shown in Table 169, treatment with ISIS antisense oligonucleotides resulted in significant reduction of PKK mRNA in comparison to the PBS control.
TABLE-US-00160 TABLE 169 Percent Inhibition of PKK mRNA in the transgenic mice liver relative to the PBS control % ISIS No Dose inhibition 547927 20 71 10 93 5 52 2.5 35 547455 20 62 10 45 5 69 2.5 0 546232 20 84 10 30 5 53 2.5 57 546254 20 83 10 84 5 55 2.5 31 546343 20 86 10 66 5 n/a 2.5 46 548048 20 80 10 72 5 77 2.5 7 546828 20 83 10 32 5 62 2.5 77 546251 20 79 10 66 5 51 2.5 13 547457 20 62 10 45 5 69 2.5 0
Protein Analysis
[1079] Plasma PKK protein levels were evaluated in all groups. Results are presented as percent inhibition of PKK protein, relative to PBS control. As shown in Table 170, treatment with ISIS antisense oligonucleotides resulted in significant reduction of PKK protein levels in comparison to the PBS control.
TABLE-US-00161 TABLE 170 Percent reduction of PKK protein levels in the transgenic mice relative to the PBS control % ISIS No Dose inhibition 547927 20 80 10 n/a 5 21 2.5 25 547455 20 78 10 32 5 0 2.5 0 546232 20 79 10 33 5 6 2.5 0 546254 20 76 10 51 5 36 2.5 0 546343 20 79 10 38 5 n/a 2.5 0 548048 20 98 10 89 5 70 2.5 23 546828 20 93 10 36 5 25 2.5 0 546251 20 69 10 52 5 30 2.5 22 547457 20 60 10 31 5 4 2.5 0
Example 121: Effect of ISIS Antisense Oligonucleotides Targeting Human PKK in Cynomolgus Monkeys
[1080] Cynomolgus monkeys were treated with ISIS antisense oligonucleotides selected from studies described above. Antisense oligonucleotide efficacy and tolerability were evaluated. The human antisense oligonucleotides tested are cross-reactive with the rhesus genomic sequence (GENBANK Accession No. NW_001118167.1 truncated from nucleotides 2358000 to 2391000 and designated herein as SEQ ID NO: 18). The target start site of each oligonucleotide to SEQ ID NO: 18 is presented in Table 171. `Mismatches` indicates that the number of nucleotides by which the oligonucleotide is mismatched to the rhesus sequence. The greater the complementarity between the human oligonucleotide and the rhesus monkey sequence, the more likely the human oligonucleotide can cross-react with the rhesus monkey sequence. `n/a` indicates that the oligonucleotide is has more than 3 mismatches with the rhesus gene sequence.
TABLE-US-00162 TABLE 171 Antisense oligonucleotides complementary to SEQ ID NO: 18 Target SEQ ID ISIS No Start Site Mismatches Sequence Chemistry NO. 547927 22059 1 ATGGTCCGACACACAA Deoxy, MOE and cEt 1548 546232 n/a n/a AGGAACTTGGTGTGCCACTT 5-10-5 MOE 526 547455 27391 0 ATATCATGATTCCCTTCTGA 5-10-5 MOE 657 546254 23858 1 TGCAAGTCTCTTGGCAAACA 5-10-5 MOE 570 546343 30532 0 CCCCCTTCTTTATAGCCAGC 5-10-5 MOE 705 548048 27397 0 CGATATCATGATTCCC Deoxy, MOE and cEt 1666 546828 13632 1 ACAGTATCACTGTACTAGTT 5-10-5 MOE 904 546251 23846 0 GGCAAACATTCACTCCTTTA 5-10-5 MOE 566 547457 27397 0 AAGGCGATATCATGATTCCC 5-10-5 MOE 660
Treatment
[1081] Prior to the study, the monkeys were kept in quarantine for a 30-day period, during which the animals were observed daily for general health. The monkeys were 2-4 years old and weighed between 2 and 4 kg. Ten groups of four randomly assigned male cynomolgus monkeys each were injected subcutaneously with ISIS oligonucleotide or PBS. PBS solution or ISIS oligonucleotides at a dose of 40 mg/kg were administered initially with a loading regimen consisting of four doses on the first week of the study (days 1, 3, 5, and 7), followed by a maintenance regimen consisting of once weekly administration starting on Day 14 (weeks 2 to 13). Subcutaneous injections were performed in clock-wise rotations at 4 sites on the back; one site per dose. The injection sites were delineated by tattoo, while sedated using ketamine, and were separated by a minimum of 3 cm.
[1082] During the study period, the monkeys were observed a minimum of once daily for signs of illness or distress. Any animal experiencing more than momentary or slight pain or distress due to the treatment, injury or illness was promptly reported to the responsible veterinarian and the Study Director. Any animal in poor health or in a possible moribund condition was identified for further monitoring and possible euthanasia. For example, two monkeys treated with ISIS 547445 were euthanized due to subdued behavior, lateral position, lack of response to stimuli and decreased respiration. The protocols described in the Example were approved by the Institutional Animal Care and Use Committee (IACUC).
Target Reduction
RNA Analysis
[1083] On day 87 or 88, 48 hours after the final dose, RNA was extracted from liver tissue for real-time PCR analysis of PKK using primer probe set RTS3455 (forward sequence CCTGTGTGGAGGGTCACTCA, designated herein as SEQ ID NO: 23; reverse sequence CCACTATAGATGCGCCAAACATC, designated herein as SEQ ID NO: 24; probe sequence CCCACTGCTTTGATGGGCTTCCC, designated herein as SEQ ID NO: 25). The results were normalized to the housekeeping gene, Cyclophilin. Results are presented as percent inhibition of PKK mRNA, relative to PBS control. As shown in Table 172, treatment with ISIS antisense oligonucleotides resulted in significant reduction of PKK mRNA in comparison to the PBS control.
TABLE-US-00163 TABLE 172 Percent Inhibition of PKK mRNA in the cynomolgus monkey liver relative to the PBS control ISIS No % inhibition 546232 88 546251 90 546254 88 546343 74 546828 45 547455 90 547457 89 547927 54 548048 95
Protein Analysis
[1084] Approximately 0.9 mL of blood was collected each time from all available animals at pre-dose, day 17, day 31, day 45, day 59, and day 73, and placed in tubes containing 3.2% sodium citrate. The tubes were centrifuged (3000 rpm for 10 min at room temperature) to obtain plasma. PKK protein levels were measured in the plasma by ELISA. The results are presented in Table 173, expressed as percentage inhibition compared to the PBS control levels. The results indicate that ISIS oligonucleotides significantly reduced PKK protein levels.
TABLE-US-00164 TABLE 173 PKK protein level reduction (%) in the cynomolgus monkey plasma relative to control levels Day 17 Day 31 Day 45 Day 59 Day 73 ISIS 546232 53 58 72 75 70 ISIS 546251 71 75 75 81 77 ISIS 546254 38 51 63 74 73 ISIS 546343 56 74 69 70 70 ISIS 546828 0 8 23 39 39 ISIS 547455 26 33 43 58 58 ISIS 547457 68 75 79 76 80 ISIS 547927 8 0 15 10 18 ISIS 548048 90 93 95 95 95
Tolerability Studies
Liver Function
[1085] To evaluate the effect of ISIS oligonucleotides on hepatic function, the monkeys were fasted overnight. Approximately, 1.5 mL of blood samples were collected from all the study groups. Blood was collected in tubes without anticoagulant for serum separation. The tubes were kept at room temperature for a minimum of 90 min and then centrifuged at 3,000 rpm for 10 min. Levels of various liver function markers were measured using a Toshiba 120FR NEO chemistry analyzer (Toshiba Co., Japan). The results are presented in Table 174 and indicate that antisense oligonucleotides had no effect on liver function outside the expected range for antisense oligonucleotides.
TABLE-US-00165 TABLE 174 Liver function markers in cynomolgus monkey plasma Albumin AST ALT (g/dL) (IU/L) (IU/L) PBS 4.2 48 60 ISIS 546232 4.1 63 140 ISIS 546251 3.7 51 58 ISIS 546254 3.8 68 54 ISIS 546343 4.3 49 76 ISIS 546828 3.7 75 67 ISIS 547455 3.8 56 61 ISIS 547457 4.0 54 52 ISIS 547927 4.2 59 61 ISIS 548048 4.2 44 47
Hematology
[1086] To evaluate any effect of ISIS oligonucleotides in cynomolgus monkeys on hematologic parameters, blood samples of approximately 1.2 mL of blood was collected pre-dose and on day 87 or day 88 from each of the available study animals in tubes containing K.sub.2-EDTA. Samples were analyzed for red blood cell (RBC) count, white blood cells (WBC) count, platelet count, hemoglobin content and hematocrit, using an ADVIA2120i hematology analyzer (SIEMENS, USA). The data is presented in Table 175.
[1087] The data indicate treatment with most of the oligonucleotides did not cause any changes in hematologic parameters outside the expected range for antisense oligonucleotides at this dose.
TABLE-US-00166 TABLE 175 Hematological parameters in cynomolgus monkeys RBC Platelets WBC Hemoglobin HCT (.times.10.sup.6/.mu.L) (.times.10.sup.3/.mu.L) (.times.10.sup.3/.mu.L) (g/dL) (%) PBS 5.4 458 13 13.1 43 ISIS 546232 5.4 391 11 12.9 42 ISIS 546251 5.7 419 8 12.9 43 ISIS 546254 5.3 436 11 12.4 41 ISIS 546343 5.5 373 14 12.6 42 ISIS 546828 6.0 408 11 12.9 43 ISIS 547455 4.5 448 13 10.2 34 ISIS 547457 6.4 367 10 13.8 45 ISIS 547927 5.2 461 45 12.5 41 ISIS 548048 5.9 393 11 13.4 44
Kidney Function
[1088] To evaluate the effect of ISIS oligonucleotides on kidney function, the monkeys were fasted overnight. Approximately, 1.5 mL of blood samples were collected from all the study groups. Blood was collected in tubes without anticoagulant for serum separation. The tubes were kept at room temperature for a minimum of 90 min and then centrifuged at 3,000 rpm for 10 min. Levels of BUN and creatinine were measured using a Toshiba 120FR NEO chemistry analyzer (Toshiba Co., Japan). Results are presented in Table 176, expressed in mg/dL. The plasma chemistry data indicate that most of the ISIS oligonucleotides did not have any effect on the kidney function outside the expected range for antisense oligonucleotides. Specifically, treatment with ISIS 546254 was well tolerated in terms of the kidney function of the monkeys.
[1089] Kidney function was also assessed by urinalysis. Fresh urine from all animals was collected using a clean cage pan on wet ice. Food was removed overnight the day before fresh urine collection was done but water was supplied. The total protein and creatinine levels were measured using a Toshiba 120FR NEO automated chemistry analyzer (Toshiba Co., Japan) and the protein to creatinine ratio was calculated. The results are presented in Table 177.
TABLE-US-00167 TABLE 176 Plasma BUN and creatinine levels (mg/dL) in cynomolgus monkeys BUN Creatinine PBS 22.8 0.9 ISIS 546232 22.7 1.0 ISIS 546251 25.4 1.1 ISIS 546254 25.7 0.9 ISIS 546343 26.2 1.0 ISIS 546828 24.7 0.9 ISIS 547455 29.4 0.9 ISIS 547457 24.3 1.0 ISIS 547927 22.3 1.0 ISIS 548048 21.9 0.9
TABLE-US-00168 TABLE 177 Urine protein/creatinine ratio in cynomolgus monkeys Ratio ISIS 546232 0.03 ISIS 546251 0.12 ISIS 546254 0.04 ISIS 546343 0.01 ISIS 546828 0.03 ISIS 547455 0.70 ISIS 547457 0.03 ISIS 547927 0.04 ISIS 548048 0.03 PBS 0.06
C-Reactive Protein Level Analysis
[1090] To evaluate any inflammatory effect of ISIS oligonucleotides in cynomolgus monkeys, the monkeys were fasted overnight. Approximately, 1.5 mL of blood samples were collected from all the study groups. Blood was collected in tubes without anticoagulant for serum separation. The tubes were kept at room temperature for a minimum of 90 min and then centrifuged at 3,000 rpm for 10 min. C-reactive protein (CRP), which is synthesized in the liver and which serves as a marker of inflammation, was measured using a Toshiba 120FR NEO chemistry analyzer (Toshiba Co., Japan). Complement C3 was also measured similarly, and the data is presented as a percentage of baseline values. The results are presented in Table 178 and indicate that treatment with ISIS oligonucleotides did not cause any inflammation in monkeys.
TABLE-US-00169 TABLE 178 C-reactive protein and C3 levels in cynomolgus monkey plasma CRP C3 (% of (mg/dL) baseline) PBS 0.2 73 ISIS 546232 0.5 50 ISIS 546251 0.7 62 ISIS 546254 0.8 61 ISIS 546343 0.2 60 ISIS 546828 0.6 56 ISIS 547455 1.9 64 ISIS 547457 0.3 53 ISIS 547927 0.2 73 ISIS 548048 0.2 69
Example 122: Antisense Inhibition of Murine PKK mRNA in Mouse Primary Hepatocytes
[1091] Antisense oligonucleotides targeting a murine PKK nucleic acid were designed and tested for their effects on PKK mRNA in vitro. Cultured mouse primary hepatocytes at a density of 10,000 cells per well were transfected using Cytofectin reagent with 12.5 nM, 25.0 nM, 50.0 nM, 100.0 nM, and 200.0 nM of antisense oligonucleotide. After a treatment period of approximately 24 hours, RNA was isolated from the cells and mouse PKK mRNA levels were measured by quantitative real-time PCR using the murine primer probe set RTS3313 (forward sequence TGCCTGCTGTTCAGCTTTCTC, designated herein as SEQ ID NO: 2228; reverse sequence TGGCAAAGTCCCTGTAATGCT, designated herein as SEQ ID NO: 2229; probe sequence CGTGACTCCACCCAAAGAGACAAATAAACG, designated herein as SEQ ID NO: 2230). PKK mRNA levels were adjusted according to total RNA content, as measured by RIBOGREEN.
[1092] The chimeric antisense oligonucleotides were designed as 5-10-5 MOE gapmers. The gapmers are 20 nucleotides in length, wherein the central gap segment is comprised of ten 2'-deoxynucleosides and is flanked on both sides (in the 5' and 3' directions) by wings comprising 5 nucleosides each. Each nucleoside in the 5' wing segment and each nucleoside in the 3' wing segment has a 2'-O-methoxyethyl modification. The internucleoside linkages throughout each gapmer are phosphorothioate linkages. All cytosine residues throughout each gapmer are 5-methylcytosines. Results demonstrate that PKK mRNA levels were significantly reduced in a dose dependent manner.
[1093] In one specific example, ISIS 482584 (GGCATATTGGTTTTTGGAAT; SEQ ID NO: 2244) reduced PKK mRNA in a dose dependent manner yielding a half maximal inhibitory concentration (IC.sub.50) of 84 nM (see Table 179). ISIS 482584 is targeted to SEQ ID NO: 11 (GENBANK Accession No. NM_008455.2) and has a target start site of 1586 and a target stop site of 1605. "Target start site" indicates the 5'-most nucleotide to which the gapmer is targeted. "Target stop site" indicates the 3'-most nucleotide to which the gapmer is targeted.
TABLE-US-00170 TABLE 179 Dose-dependent inhibition of mouse PKK mRNA levels by ISIS 482584 % Dose inhibition 12.5 nM 0 25.0 nM 47 50.0 nM 27 100.0 nM 60 200.0 nM 82
Example 123: Antisense Inhibition of PKK mRNA in BALB/c Mice
[1094] ISIS 482584 was tested for its effect on murine PKK mRNA in vivo.
Treatment
[1095] Six groups of male BALB/c mice each were treated with 2.5 mg/kg, 5.0 mg/kg, 10.0 mg/kg, 20.0 mg/kg, 40.0 mg/kg, or 80.0 mg/kg of ISIS 482584, administered subcutaneously twice a week for 3 weeks (weekly doses of 5.0 mg/kg, 10.0 mg/kg, 20.0 mg/kg, 40.0 mg/kg, 80.0 mg/kg, or 160.0 mg/kg). A control group of BALB/c mice was treated with PBS, administered subcutaneously twice a week for 3 weeks. Two days after the last dose of antisense oligonucleotide or PBS, mice from all groups were anesthetized with 150 mg/kg ketamine mixed with 10 mg/kg xylazine, administered by intraperitoneal injection. Liver was collected for RNA analysis.
RNA Analysis
[1096] RNA was extracted from liver tissue for real-time PCR analysis of PKK. PKK mRNA levels were measured using the murine primer probe set (forward sequence ACAAGTGCATTTTACAGACCAGAGTAC, designated herein as SEQ ID NO: 2231; reverse sequence GGTTGTCCGCTGACTTTATGCT, designated herein as SEQ ID NO: 2232; probe sequence AAGCACAGTGCAAGCGGAACACCC, designated herein as SEQ ID NO: 2233). Results are presented as percent inhibition of PKK, relative to PBS control. As shown in Table 180, treatment with ISIS 482584 resulted in significant dose-dependent reduction of PKK mRNA in comparison to the PBS control.
TABLE-US-00171 TABLE 180 Dose-dependent reduction of PKK mRNA in BALB/c mice liver Dose % (mg/kg/wk) inhibition 5 3 10 42 20 68 40 85 80 91 160 94
Protein Analysis
[1097] Plasma was collected in tubes containing sodium citrate as an anticoagulant. The samples were run on a 4-12% gradient SDS-polyacrylamide gel (Invitrogen), followed by immunoblotting with murine PKK antibody (R&D Systems). Blots were incubated with secondary fluorophore-labeled antibodies (LI-COR) and imaged in an Odyssey Imager (LI-COR). Results are presented as percent inhibition of PKK, relative to PBS control. As shown in Table 181, treatment with ISIS 482584 resulted in significant dose-dependent reduction of PKK plasma protein in comparison to the PBS control.
TABLE-US-00172 TABLE 181 Dose-dependent reduction of PKK protein in BALB/c mice plasma Dose % (mg/kg/wk) inhibition 5 5 10 24 20 47 40 76 80 81 160 n.d. n.d. = no data
Example 124: In Vivo Effect of Antisense Inhibition of Murine PKK in an Angioedema Mouse Model
[1098] Hereditary angioedema (HAE) is characterized by local swelling and increase in vascular permeability in subcutaneous tissues (Morgan, B. P. N. Engl. J. Med. 363: 581-83, 2010). It is caused by a deficiency of the C1 inhibitor, a protein of the complement system. Two mouse models were used in this study including an established mouse model of C1-INH deficiency and a captopril-induced edema model, both of which cause vascular permeability, a hallmark of HAE. Reversal of vascular permeability is accompanied by increased plasma levels of high molecular weight kininogen (HMWK).
[1099] In the first model, angioedema was induced by treatment with Captopril, a known antihypertensive agent, which increases vascular permeability in mice and replicates the pathology of hereditary angioedema.
[1100] In the second model, angioedema was induced by treatment with ISIS 461756, an antisense oligonucleotide which targets murine C1 inhibitor mRNA, which increases vascular permeability in mice and replicates the pathology of hereditary angioedema. ISIS 461756 (SEQ ID NO: 2245; AAAGTGGTTGATACCCTGGG) is a 5-10-5 MOE gapmer targeting nucleosides 1730-1749 of NM_009776.3 (SEQ ID NO: 2243).
[1101] The effect of HOE-140 and ISIS 482584, an antisense oligonucleotide inhibitor of PKK, were evaluated in the Captopril and ISIS 461756-induced mouse models of vascular permeability. Some of the murine groups were treated with HOE-140, a selective antagonist of the bradykinin B2 receptor, which blocks vasodilation and vascular permeability (Cruden and Newby, Expert Opin. Pharmacol. 9: 2383-90, 2008). Other mice were treated with ISIS 482584, which inhibits PKK mRNA expression. The effect of treatment with HOE-140 was compared with the effect of treatment with ISIS 482584.
Treatment
[1102] The various treatment groups for this assay are presented in Table 182.
[1103] Group 1 consisted of 4 C57BL/6J-Tyrc-2J mice treated with PBS administered subcutaneously twice a week for 4 weeks. No other treatment was administered to Group 1 which served as a control group to measure the basal level of vascular permeability.
[1104] Group 2 consisted of 8 C57BL/6J-Tyrc-2J mice treated with PBS administered subcutaneously twice a week for 4 weeks. At the end of the treatment, the mice were intraperitoneally administered 20 .mu.g of captopril. Group 2 served as a PBS control group for captopril-induced vascular permeability.
[1105] Group 3 consisted of 8 C57BL/6J-Tyrc-2J mice treated with PBS administered subcutaneously twice a week for 4 weeks. On day 14, the mice were treated with 50 mg/kg of the antisense oligonucleotide targeting C1 inhibitor, ISIS 461756, administered subcutaneously twice a week for 2 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. Group 3 served as a PBS control group for captopril and ISIS 461756-induced vascular permeability.
[1106] Group 4 consisted of 8 C57BL/6J-Tyrc-2J mice treated with PBS administered subcutaneously twice a week for 4 weeks. On day 14, the mice were treated with 50 mg/kg of the antisense oligonucleotide targeting C1 inhibitor, ISIS 461756, administered subcutaneously twice a week for 2 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. The mice were then also intraperitoneally administered 30 .mu.g of HOE-140. Group 4 served as a positive control for inhibition of vascular permeability with HOE-140.
[1107] Group 5 consisted of 8 C57BL/6J-Tyrc-2J mice treated with 40 mg/kg of control oligonucleotide ISIS 141923, a 5-10-5 MOE gapmer with no known murine target, (CCTTCCCTGAAGGTTCCTCC; SEQ ID NO: 2246) administered subcutaneously twice a week for 4 weeks. On day 14, the mice were treated with 50 mg/kg of the antisense oligonucleotide targeting C1 inhibitor, ISIS 461756, administered subcutaneously twice a week for 2 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. Group 5 served as a control group for captopril and ISIS 461756-induced vascular permeability.
[1108] Group 6 consisted of 8 C57BL/6J-Tyrc-2J mice and was treated with 40 mg/kg of ISIS 482584 administered subcutaneously twice a week for 4 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. Group 6 served as the experimental treatment group for examining the effect of PKK ASO on captopril-induced vascular permeability.
[1109] Group 7 consisted of 8 C57BL/6J-Tyrc-2J mice treated with 40 mg/kg of ISIS 482584 administered subcutaneously twice a week for 4 weeks. On day 14, the mice were treated with 50 mg/kg of the antisense oligonucleotide targeting C1 inhibitor, ISIS 461756, administered subcutaneously twice a week for 2 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. Group 7 served as the experimental treatment group for examining the effect of PKK ASO on captopril and ISIS 461756-induced vascular permeability.
[1110] All the groups were then injected with 30 mg/kg of Evans Blue solution into the tail vein. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, ears, and intestines were harvested. Blood samples were taken through cardiac puncture.
TABLE-US-00173 TABLE 182 Treatment groups Group No. Treatment Captopril ISIS 461756 HOE-140 1. (N = 4) PBS No No No 2. (N = 8) PBS Yes No No 3. (N = 8) PBS Yes Yes No 4. (N = 8) PBS Yes Yes Yes 5. (N = 8) ISIS 141923 Yes Yes No 6. (N = 8) ISIS 482584 Yes No No 7. (N = 8) ISIS 482584 Yes Yes No
Quantification of Vascular Permeability
[1111] The harvested tissues from the feet, colon, ears, and intestines were placed separately in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing ear and feet tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm, and is presented in Table 183. Mice displaying any manifestation of angioedema take up more dye and, therefore, demonstrate high OD values.
[1112] As presented in Table 183, treatment with ISIS 482584 prevents vascular permeability in mice treated with captopril (Group 6) and in mice treated with captopril and ISIS 461756 (Group 7) compared to the respective PBS control groups (Groups 2 and 3). Measures of vascular permeability in mice of Groups 6 and 7 were also reduced in most of the tissues in comparison to the mice treated with the control oligonucleotide, ISIS 141923 (Group 5), where vascular permeability was induced with captopril and ISIS 461756. Measures of vascular permeability in the colon and feet tissues of both the treatment groups (Groups 6 and 7) were comparable to basal levels, as observed in mice treated with only PBS (Group 1). Reduction in vascular permeability in mice treated with ISIS 482584 was comparable to that seen in mice treated with the bradykinin 2 receptor antagonist, HOE140, which served as a positive control in this assay.
[1113] Therefore, antisense inhibition of PKK mRNA may be beneficial for the treatment and prevention of vascular permeability, which is symptomatic of HAE.
TABLE-US-00174 TABLE 183 OD.sub.600nm of Evans Blue dye to measure vascular permeability Group ISIS No. Treatment Captopril 461756 HOE-140 Colons Intestines Feet Ears 1 PBS No No No 0.26 0.16 0.11 0.02 2 PBS Yes No No 0.49 0.29 0.12 0.07 3 PBS Yes Yes No 0.49 0.34 0.11 0.12 4 PBS Yes Yes Yes 0.14 0.18 0.07 0.09 5 ISIS 141923 Yes Yes No 0.44 0.29 0.14 0.08 6 ISIS 482584 Yes No No 0.27 0.30 0.07 0.14 7 ISIS 482584 Yes Yes No 0.21 0.34 0.07 0.06
Quantification of High Molecular Weight Kininogen (HMWK)
[1114] Western blot quantification of HMWK from blood samples shows that Groups 1 and 2 have low levels of HMWK as compared to Groups 6 and 7 indicating that vascular permeability is reversed in Groups 6 and 7. Samples from Groups 1 and 2 have increased HMWK cleavage product as compared to Groups 6 and 7. Thus, lack of HMWK is caused by PKK cleavage of HMWK into cleavage products (including bradykinin and HKa).
Example 125: In Vivo Effect of Antisense Inhibition of Murine PKK on Basal Permeability and Captopril-Induced Permeability in Mice
[1115] Basal permeability is the level of vascular permeability occurring in the tissues of naive, untreated mice. The effect of ISIS 482584 in the prevention of vascular permeability, either basal or captopril-induced, was evaluated.
Treatment
[1116] The various treatment groups for this assay are presented in Table 184.
[1117] Group 1 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 4 weeks. No other treatment was administered to Group 1 which served as a control group to measure the basal levels of vascular permeability.
[1118] Group 2 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 4 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril.
[1119] Group 2 served as the negative control group for captopril-induced vascular permeability.
[1120] Group 3 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 4 weeks. At the end of the treatment period, the mice were intraperitoneally administered 30 .mu.g of HOE-140.
[1121] Group 3 served as a positive control for inhibition of basal vascular permeability.
[1122] Group 4 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 4 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. The mice were also intraperitoneally administered 30 .mu.g of HOE-140. Group 4 served as a positive control for inhibition of captopril-induced vascular permeability.
[1123] Group 5 consisted of 8 mice and was treated with 40 mg/kg of ISIS 482584 administered subcutaneously twice a week for 4 weeks. Group 5 served as an experimental treatment group for examining the effect of ISIS 482584 on basal vascular permeability.
[1124] Group 6 consisted of 8 mice and was treated with 40 mg/kg of ISIS 482584 administered subcutaneously twice a week for 4 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. Group 6 served as an experimental treatment group for examining the effect of ISIS 482584 on captopril-induced vascular permeability.
[1125] All the groups were then injected with 30 mg/kg of Evans Blue solution. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, ears, and intestines were harvested.
TABLE-US-00175 TABLE 184 Treatment groups Group No. Treatment Captopril HOE-140 1. (N = 8) PBS No No 2. (N = 8) PBS Yes No 3. (N = 8) PBS No Yes 4. (N = 8) PBS Yes Yes 5. (N = 8) ISIS 482584 No No 6. (N = 8) ISIS 482584 Yes No
Quantification of Vascular Permeability
[1126] The harvested tissues from the feet, colon, intestine, and ears were placed separately in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing feet and ear tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm, and is presented in Table 185. Mice displaying any manifestation of angioedema take up more dye and, therefore, demonstrate high OD values.
[1127] As presented in Table 185, mice treated with ISIS 482584 demonstrated reduced basal vascular permeability compared to the PBS control (Group 5 vs. Group 1). The reduction in basal vascular permeability by treatment with ISIS 482584 was comparable to that caused by treatment with HOE-140 (Group 3, which served as the positive control). Mice treated with ISIS 482584 also demonstrated reduced captopril-induced vascular permeability in most tissues compared to the PBS control (Group 6 vs. Group 2). The reduction in captopril-induced vascular permeability by treatment with ISIS 482584 was comparable to that caused by treatment with HOE-140 (Group 4, which served as the positive control).
TABLE-US-00176 TABLE 185 OD.sub.600nm of Evans Blue dye to measure vascular permeability Group HOE- No. Treatment Captopril 140 Colon Feet Intestine Ears 1 PBS No No 0.27 0.08 0.23 0.06 2 PBS Yes No 0.61 0.08 0.24 0.01 3 PBS No Yes 0.18 0.06 0.21 0.03 4 PBS Yes Yes 0.29 0.03 0.14 0.00 5 ISIS 482584 No No 0.19 0.07 0.22 0.04 6 ISIS 482584 Yes No 0.37 0.05 0.22 0.00
Example 126: Dose-Dependent Effect of Antisense Inhibition of Murine PKK on Captopril-Induced Vascular Permeability
[1128] The effect of varying doses on ISIS 482584 on captopril-induced vascular permeability was evaluated.
Treatment
[1129] The various treatment groups for this assay are presented in Table 186.
[1130] Group 1 consisted of 4 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. No other treatment was administered to Group 1 which served as a control group to measure the basal levels of vascular permeability.
[1131] Group 2 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril.
[1132] Group 2 served as the control group for captopril-induced vascular permeability.
[1133] Group 3 consisted of 4 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. The mice were also intraperitoneally administered 30 .mu.g of Icatibant (HOE-140). Group 4 served as a positive control for inhibition of captopril-induced vascular permeability.
[1134] Groups 4, 5, 6, 7, 8, and 9 consisted of 8 mice each and were treated with 2.5 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, or 80 mg/kg (corresponding to 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, 80 mg/kg, or 160 mg/kg per week), respectively of ISIS 482584 administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice of all the groups were intraperitoneally administered 20 .mu.g of captopril. Groups 4-9 served as the experimental treatment groups for examining the effect of varying doses of ISIS 482584 on captopril-induced vascular permeability.
[1135] All the groups were then injected with 30 mg/kg of Evans Blue solution in the tail vein. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, ears, and intestines were harvested. Blood samples were taken through cardiac puncture.
TABLE-US-00177 TABLE 186 Treatment groups Group Dose No. Treatment (mg/kg/wk) Captopril HOE-140 1. (N = 4) PBS -- No No 2. (N = 8) PBS -- Yes No 3. (N = 4) PBS -- Yes Yes 4. (N = 8) ISIS 482584 160 Yes No 5. (N = 8) ISIS 482584 80 Yes No 6. (N = 8) ISIS 482584 40 Yes No 7. (N = 8) ISIS 482584 20 Yes No 8. (N = 8) ISIS 482584 10 Yes No 9. (N = 8) ISIS 482584 5 Yes No
Quantification of Vascular Permeability
[1136] The harvested tissues were placed in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing feet and ear tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm, and is presented in Table 187. Mice displaying any manifestation of angioedema take up more dye and, therefore, demonstrate high OD values.
[1137] As presented in Table 187, mice treated with higher doses of ISIS 482584 (Groups 4, 5, and 6) had reduced levels of captopril-induced vascular permeability compared to the corresponding PBS control group (Group 2). The reduction in vascular permeability in mice of these treatment groups (Groups 4 and 5) was comparable to the levels of basal vascular permeability (as shown in Group 1) as well as in mice treated with HOE-140 (Group 3).
TABLE-US-00178 TABLE 187 OD.sub.600nm of Evans Blue dye to measure vascular permeability Group Dose No. Treatment (mg/kg) Captopril HOE-140 Colon Feet Intestine Ears 1 PBS -- No No 0.16 0.07 0.13 0.01 2 PBS -- Yes No 0.39 0.12 0.18 0.07 3 PBS -- Yes Yes 0.15 0.03 0.10 0.04 4 ISIS 482584 160 Yes No 0.26 0.10 0.15 0.05 5 ISIS 482584 80 Yes No 0.21 0.04 0.17 0.03 6 ISIS 482584 40 Yes No 0.36 0.10 0.20 0.05 7 ISIS 482584 20 Yes No 0.40 0.11 0.20 0.07 8 ISIS 482584 10 Yes No 0.41 0.10 0.19 0.05 9 ISIS 482584 5 Yes No 0.41 0.10 0.17 0.05
Quantification of Vascular Leakage
[1138] The blood drawn through cardiac puncture was immediately mixed with 3 times the volume of ice-cold ethanol. The solution was centrifuged at 15,000 g for 20 minutes at 4.degree. C. to remove cell debris and precipitated plasma proteins. The ethanol extracts were further purified by ultra-filtration through a 10 kDa MWCO filter. The color intensity of the ethanol extracted plasma solution was then measured at OD.sub.620 nm. The results are presented in Table 188 as percentage increase or decrease of the OD values of the Group 1 PBS control. It was expected that tissues from mice displaying manifestation of angioedema would leak more dye from the plasma and, therefore, demonstrate low OD values, whereas treatment groups may display higher OD values due to reduced vascular leakage. Mice treated with 160 mg/kg/week and 80 mg/kg/week of ISIS 482584 (Groups 4 and 5) demonstrated less vascular leakage compared to the PBS negative control treated with captopril (Group 2). The results from Groups 4 and 5 were comparable to the positive control treated with HOE-140 (Group 3).
TABLE-US-00179 TABLE 188 Percentage of OD.sub.620 nm of Evans Blue dye compared to the PBS basal control to measure vascular leakage Group Dose HOE- No. Treatment (mg/kg) Captopril 140 Plasma 2 PBS -- Yes No -43 3 PBS -- Yes Yes 5 4 ISIS 482584 160 Yes No 91 5 ISIS 482584 80 Yes No 40 6 ISIS 482584 40 Yes No -31 7 ISIS 482584 20 Yes No -26 8 ISIS 482584 10 Yes No -20 9 ISIS 482584 5 Yes No -23
Example 127: Dose-Dependent Effect of Antisense Inhibition of Murine PKK on Basal Permeability in Mice
[1139] The effect of varying doses on ISIS 482584 on basal vascular permeability was evaluated.
Treatment
[1140] The various treatment groups for this assay are presented in Table 189.
[1141] Group 1 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. No other treatment was administered to Group 1 which served as a control group to measure the basal levels of vascular permeability.
[1142] Group 2 consisted of 4 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice were intraperitoneally administered 30 .mu.g of HOE-140.
[1143] Group 2 served as a positive control for inhibition of basal vascular permeability.
[1144] Groups 3, 4, 5, 6, 7, and 8 consisted of 8 mice each and were treated with 2.5 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, or 80 mg/kg (corresponding to 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, 80 mg/kg, or 160 mg/kg per week), respectively of ISIS 482584 administered subcutaneously twice a week for 3 weeks. Groups 4-9 served as the experimental treatment groups for examining the effect of varying doses of ISIS 482584 on basal vascular permeability.
[1145] All the groups were then injected with 30 mg/kg of Evans Blue solution in the tail vein. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, and ears were harvested and examined for permeability defects. Blood samples were taken through cardiac puncture.
TABLE-US-00180 TABLE 189 Treatment groups Group Dose No. Treatment (mg/kg/week) HOE-140 1. (N = 8) PBS -- No 2. (N = 4) PBS -- Yes 3. (N = 8) ISIS 482584 160 No 4. (N = 8) ISIS 482584 80 No 5. (N = 8) ISIS 482584 40 No 6. (N = 8) ISIS 482584 20 No 7. (N = 8) ISIS 482584 10 No 8. (N = 8) ISIS 482584 5 No
Quantification of Vascular Permeability
[1146] The harvested tissues from the feet, colon, and ears were placed in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing feet and ear tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm, and is presented in Table 190. Higher OD values are associated with higher levels of permeability.
[1147] As presented in Table 190, most of the tissues of mice treated with ISIS 482584 at all doses (Groups 3-8) demonstrated reduced basal vascular permeability compared to the PBS control (Group 1). The reduction in basal vascular permeability of the ISIS oligonucleotide-treated groups was comparable to the same demonstrated in the positive control group treated with HOE-140 (Group 2).
TABLE-US-00181 TABLE 190 OD.sub.600nm of Evans Blue dye to measure vascular permeability Group Dose HOE- No. Treatment (mg/kg/week) 140 Colon Feet Ears 1 PBS -- No 0.27 0.17 0.013 2 PBS -- Yes 0.24 0.09 0.047 3 ISIS 482584 160 No 0.25 0.11 0.019 4 ISIS 482584 80 No 0.24 0.09 0.014 5 ISIS 482584 40 No 0.27 0.11 0.011 6 ISIS 482584 20 No 0.26 0.11 0.009 7 ISIS 482584 10 No 0.31 0.10 0.015 8 ISIS 482584 5 No 0.32 0.11 0.009
Quantification of Vascular Leakage
[1148] The blood drawn through cardiac puncture was immediately mixed with 3 times the volume of ice-cold ethanol. The solution was centrifuged at 15,000 g for 20 minutes at 4.degree. C. to remove cell debris and precipitated plasma proteins. The ethanol extracts were further purified by ultra-filtration through a 10 kDa MWCO filter. The color intensity of the ethanol extracted plasma solution was then measured at OD.sub.620 nm. The results are presented in Table 191 as percentage increase or decrease of the OD values of the Group 1 PBS control. It was expected that treatment groups may display higher OD values due to reduced vascular leakage. All the mice in the ISIS oligonucleotide-treated groups demonstrated significantly reduced vascular leakage compared to the PBS negative control.
TABLE-US-00182 TABLE 191 Percentage of OD.sub.620 nm of Evans Blue dye compared to the PBS basal control to measure vascular leakage Group Dose No. Treatment (mg/kg/week) HOE-140 Plasma 2. (N = 8) ISIS 482584 160 No 95 3. (N = 8) ISIS 482584 80 No 93 4. (N = 8) ISIS 482584 40 No 83 5. (N = 8) ISIS 482584 20 No 56 6. (N = 8) ISIS 482584 10 No 36
Quantification of High Molecular Weight Kininogen (HMWK)
[1149] Western blot quantification of HMWK from blood samples are presented in Tables 192 and 193.
[1150] As shown in Table 192, Groups treated with 482584 have higher levels of HMWK as compared to PBS control, increasing in a dose-dependent manner. Treatment with PKK antisense oligonucleotide results in stabilization of HMWK. Thus, vascular permeability is reduced in ISIS 482584-treated groups in a dose-dependent manner. As shown in Table 193, Groups treated with ISIS 482584 have lower HMWK cleavage product as compared to PBS control, decreasing in a dose-dependent manner. Thus, reduced HMWK is caused by PKK cleavage of HMWK into cleavage products (including bradykinin and HKa). Data are presented in Intensity Units as measured by densitometer.
TABLE-US-00183 TABLE 192 Quantification of HMWK by densitometer Group No Treatment Dose (mg/kg/week) Intensity Units 1 PBS -- 89 3 ISIS 482584 160 21358 4 ISIS 482584 80 7279 5 ISIS 482584 40 873 6 ISIS 482584 20 608 7 ISIS 482584 10 507
TABLE-US-00184 TABLE 193 Quantification of HMWK cleavage product by densitometer Group No Treatment Dose (mg/kg/week) Intensity Units 1 PBS -- 401738 3 ISIS 482584 160 19936 4 ISIS 482584 80 204482 5 ISIS 482584 40 388135 6 ISIS 482584 20 403360 7 ISIS 482584 10 414774
Example 128: Combination Therapy of Antisense Oligonucleotides Targeting PKK and Factor 12 on Captopril-Induced Vascular Permeability in Mice
[1151] Mice were treated varying doses of ISIS 410944, a 5-10-5 MOE gapmer targeting Factor 12 (GCATGGGACAGAGATGGTGC; SEQ ID NO: 2247), and ISIS 482584 in a captopril-induced vascular permeability model.
Treatment
[1152] The various treatment groups for this assay are presented in Table 194.
[1153] Group 1 consisted of 4 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. No other treatment was administered to Group 1 which served as a control group to measure the basal levels of vascular permeability.
[1154] Group 2 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril.
[1155] Group 2 served as the control group for captopril-induced vascular permeability.
[1156] Group 3 consisted of 4 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice were intraperitoneally administered 20 .mu.g of captopril. The mice were also intraperitoneally administered 30 .mu.g of HOE-140. Group 3 served as a positive control for inhibition of captopril-induced vascular permeability.
[1157] Groups 4, 5, 6, 7, and 8 consisted of 8 mice each and were treated with 2.5 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg, or 40 mg/kg (corresponding to 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, or 80 mg/kg per week), respectively of ISIS 482584 and ISIS 410944 each administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice of all the groups were intraperitoneally administered 20 .mu.g of captopril. Groups 4-8 served as the experimental treatment groups for examining the effect of ISIS 410944 and ISIS 482584 on captopril-induced vascular permeability.
[1158] All the groups were then injected with 30 mg/kg of Evans Blue solution in the tail vein. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, ears, and intestines were harvested.
TABLE-US-00185 TABLE 194 Treatment groups Dose Group (mg/kg/wk) No. Treatment of each ASO Captopril HOE-140 1. (N = 4) PBS -- No No 2. (N = 8) PBS -- Yes No 3. (N = 4) PBS -- Yes Yes 4. (N = 8) ISIS 80 Yes No 482584 + ISIS 410944 5. (N = 8) ISIS 40 Yes No 482584 + ISIS 410944 6. (N = 8) ISIS 20 Yes No 482584 + ISIS 410944 7. (N = 8) ISIS 10 Yes No 482584 + ISIS 410944 8. (N = 8) ISIS 5 Yes No 482584 + ISIS 410944
Quantification of Vascular Permeability
[1159] The harvested tissues from the feet, colon, and ears were placed in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing feet and ear tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm, and is presented in Table 195. Higher OD values are associated with higher levels of permeability.
[1160] As presented in Table 195, most of the tissues of mice treated with a combination of ISIS 482584 and ISIS 410944 at all doses (Groups 3-8) demonstrated reduced vascular permeability compared to the PBS control (Group 1). The reduction in vascular permeability of the ISIS oligonucleotide-treated groups was comparable to the same demonstrated in the basal PBS control (Group 1), as well as the positive control group treated with HOE140 (Group 2). Combination of PKK and Factor 12 antisense oligonucleotides results in synergistic decrease in permeability. As expected, a corresponding synergistic decrease in vascular leakage was also observed.
TABLE-US-00186 TABLE 195 OD.sub.600nm of Evans Blue dye to measure vascular permeability Dose Group (mg/kg/wk) No. Treatment of each ASO Captopril HOE-140 Colon Feet Intestines Ears 1 PBS -- No No 0.24 0.11 0.13 0.01 2 PBS -- Yes No 0.38 0.15 0.11 0.05 3 PBS -- Yes Yes 0.23 0.06 0.15 0.04 4 ISIS 482584 + 80 Yes No 0.19 0.07 0.11 0.04 ISIS 410944 5 ISIS 482584 + 40 Yes No 0.19 0.07 0.12 0.03 ISIS 410944 6 ISIS 482584 + 20 Yes No 0.22 0.08 0.12 0.04 ISIS 410944 7 ISIS 482584 + 10 Yes No 0.38 0.13 0.13 0.05 ISIS 410944 8 ISIS 482584 + 5 Yes No 0.53 0.12 0.13 0.03 ISIS 410944
Example 129: Combination Therapy of Antisense Oligonucleotides Targeting PKK and Factor 12 on Basal Vascular Permeability in Mice
[1161] Mice were treated with varying doses of ISIS 410944, an antisense oligonucleotide targeting Factor 12, and ISIS 482584 in a basal vascular permeability model.
Treatment
[1162] The various treatment groups for this assay are presented in Table 196.
[1163] Group 1 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. No other treatment was administered to Group 1 which served as a control group to measure the basal levels of vascular permeability.
[1164] Group 2 consisted of 4 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. At the end of the treatment period, the mice were intraperitoneally administered 30 .mu.g of HOE-140. Group 2 served as a positive control for inhibition of basal vascular permeability.
[1165] Groups 3, 4, 5, 6, and 7 consisted of 8 mice each and were treated with 2.5 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg, or 40 mg/kg (corresponding to 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, or 80 mg/kg per week), respectively of ISIS 482584 and ISIS 410944 each administered subcutaneously twice a week for 3 weeks. Groups 3-7 served as the experimental treatment groups for examining the effect of ISIS 410944 and ISIS 482584 on basal vascular permeability.
[1166] All the groups were then injected with 30 mg/kg of Evans Blue solution in the tail vein. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, ears, and intestines were harvested.
TABLE-US-00187 TABLE 196 Treatment groups Dose Group No. Treatment (mg/kg/wk) HOE-140 1. (N = 8) PBS -- No 2. (N = 4) PBS -- Yes 3. (N = 8) ISIS 482584 + ISIS 80 No 410944 4. (N = 8) ISIS 482584 + ISIS 40 No 410944 5. (N = 8) ISIS 482584 + ISIS 20 No 410944 6. (N = 8) ISIS 482584 + ISIS 10 No 410944 7. (N = 8) ISIS 482584 + ISIS 5 No 410944
Quantification of Vascular Permeability
[1167] The harvested tissues from the feet, colon, intestines, and ears were placed in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing feet and ear tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm, and is presented in Table 197. Higher OD values are associated with higher levels of permeability.
[1168] As presented in Table 197, most of the tissues of mice treated with a combination of ISIS 482584 and ISIS 410944 at all doses (Groups 2-7) demonstrated reduced vascular permeability compared to the PBS control (Group 1). The reduction in vascular permeability of the ISIS oligonucleotide-treated groups was comparable to the same demonstrated in positive control group treated with HOE140 (Group 2). Combination of PKK and Factor 12 antisense oligonucleotides results in synergistic decrease in permeability. As expected, a corresponding synergistic decrease in vascular leakage was also observed.
TABLE-US-00188 TABLE 197 OD.sub.600nm of Evans Blue dye to measure vascular permeability Group Dose No. Treatment (mg/kg/wk) HOE-140 Colon Feet Intestines Ears 1 PBS -- No 0.19 0.08 0.10 0.004 2 PBS -- Yes 0.14 0.04 0.08 0.008 3 ISIS 482584 + 80 No 0.14 0.04 0.09 0.01 ISIS 410944 4 ISIS 482584 + 40 No 0.15 0.05 0.10 0.006 ISIS 410944 5 ISIS 482584 + 20 No 0.15 0.04 0.10 0.007 ISIS 410944 6 ISIS 482584 + 10 No 0.15 0.06 0.10 0.004 ISIS 410944 7 ISIS 482584 + 5 No 0.14 0.05 0.13 0.002 ISIS 410944
Example 130: Inhibition of Factor 12 Protein Activation by ISIS 482584
[1169] The effect of antisense inhibition of PKK mRNA on Factor 12 protein activation was evaluated.
Treatment
[1170] The various treatment groups for this assay are presented in Table 198.
[1171] Group 1 consisted of 8 mice and was treated with PBS administered subcutaneously twice a week for 3 weeks. No other treatment was administered to Group 1 which served as a control group to measure Factor 12 activation.
[1172] Groups 2, 3, 4, 5, and 6 consisted of 8 mice each and were treated with 2.5 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg, or 40 mg/kg (corresponding to 5 mg/kg, 10 mg/kg, 20 mg/kg, 40 mg/kg, or 80 mg/kg per week), respectively of ISIS 482584 administered subcutaneously twice a week for 3 weeks. Groups 2-6 served as the treatment groups for measuring the effect of ISIS 482584 on Factor 12 activation.
[1173] At the end of the treatment period, plasma was harvested from the mice for the Spectrozyme.RTM. Factor 12a based amidolytic assay for Factor 12 in plasma.
TABLE-US-00189 TABLE 198 Treatment groups Group Dose No. Treatment (mg/kg/wk) 1. (N = 8) PBS -- 2. (N = 8) ISIS 482584 80 3. (N = 8) ISIS 482584 40 4. (N = 8) ISIS 482584 20 5. (N = 8) ISIS 482584 10 6. (N = 8) ISIS 482584 5
Assay for Factor 12 Activation in Plasma
[1174] Plasma (5 .mu.L) was added to 85 .mu.L of PBS with 1 ug/ml dextran sulfate (500 kDa) in a 96 well polypropelene microplate and the solution was incubated for 5 minutes at room temperature. Spectrozyme.RTM. FXIIa (10 .mu.L of a 2 mM solution) and 0.2 mM KALLISTOP.TM. solution was added and the absorbance kinetic was measured at 405 nm. Factor 12 activation was measured in the linear phase of absorbance accumulation. The results are presented in Table 199 as a percentage of Factor 12 activation measured in the PBS control sample. As observed in Table 199, inhibition of PKK by ISIS 482584 results in decreased activation of Factor 12 by its substrate, implying the that PKK is required for proper factor 12 activation.
TABLE-US-00190 TABLE 199 Percentage Factor 12 activation compared to the PBS control Dose % F12 (mg/kg/wk) activation 80 14 40 24 20 47 10 63 5 82
Example 131: In Vivo Effect of Antisense Inhibition of Murine PKK on C1-INH Antisense Oligonucleotide-Induced Vascular Permeability
[1175] Vascular permeability induced by ISIS 461756, an antisense oligonucleotide which targets murine C1 inhibitor mRNA, increases vascular permeability in mice and replicates the pathology of hereditary angioedema. The effect of ISIS 482584 on this model was evaluated.
Treatment
[1176] One group of 8 mice was treated with 40 mg/kg ISIS 482584 administered subcutaneously twice a week for 3 weeks (weekly dose of 80 mg/kg). A second group of 8 mice was treated with 40 mg/kg of the control oligonucleotide, ISIS 141923, administered subcutaneously twice a week for 3 weeks (weekly dose of 80 mg/kg). A third group of 8 mice was treated with PBS administered subcutaneously twice a week for 3 weeks. On day 14, all the groups were treated with 12.5 mg/kg ISIS 461756 administered subcutaneously twice a week for 3 weeks (weekly dose of 25 mg/kg). A control group of mice was treated with PBS administered subcutaneously twice a week for 3 weeks but was not administered ISIS 461756.
[1177] At the end of the treatment period, all the groups were injected with 30 mg/kg of Evans Blue solution into the tail vein. The mice were sacrificed 30 min after the Evans Blue solution administration and colons, feet, ears, and intestines were harvested. The liver was also harvested for RNA analysis.
RNA Analysis
[1178] RNA was isolated from the liver for RT-PCR analysis of C1-INH and PKK mRNAs. The primer probe set for C1-INH is RTS3218 (forward sequence GAGTCCCCCAGAGCCTACAGT, designated herein as SEQ ID NO: 2234; reverse sequence TGTCATTTGTTATTGTGATGGCTACA, designated herein as SEQ ID NO: 2235; probe sequence CTGCCCTCTACCTGGCCAACAACCA, designated herein as SEQ ID NO: 2236). The primer probe set for PKK is RTS3287 (forward sequence ACAAGTGCATTTTACAGACCAGAGTAC, designated herein as SEQ ID NO: 2237; reverse sequence GGTTGTCCGCTGACTTTATGCT, designated herein as SEQ ID NO: 2238; probe sequence AAGCACAGTGCAAGCGGAACACCC, designated herein as SEQ ID NO: 2239). The results are presented in Table 200 as percent inhibition compared to the PBS control not treated with ISIS 461756. The data indicates that ISIS 461756 significantly reduced C1-INH mRNA expression and that treatment with ISIS 482584 significantly reduced PKK expression.
TABLE-US-00191 TABLE 200 Percent inhibition of mRNA expression in mice treated with ISIS 461756 compared to the untreated PBS control C1-INH PKK Treatment mRNA mRNA PBS 76 0 ISIS 141923 79 0 ISIS 482584 77 78
Quantification of Vascular Permeability
[1179] The harvested tissues from the feet, colon, and intestines were placed in formamide solution overnight to leach out the Evans Blue dye. The formamide solution containing feet tissue was heated to 55.degree. C. and left overnight. The color intensity of the dye-infused formamide solution was then measured at OD.sub.600 nm. The data is presented in Table 201 as percent increase or reduction compared to the PBS control not treated with ISIS 461756. The data indicates that treatment with ISIS 482584 prevented vascular permeability induced by ISIS 461756.
TABLE-US-00192 TABLE 201 Percent change in vascular permeability in mice treated with ISIS 461756 compared to the untreated PBS control Treatment Colon Feet Intestines PBS 13 70 27 ISIS 141923 2 80 14 ISIS 482584 -23 2 -25
Example 132: In Vivo Effect of Antisense Inhibition of Murine PKK in the FeCl.sub.3-Induced Inferior Vena Cava Thrombosis Model
[1180] ISIS 482584, which demonstrated significant in vitro and in vivo inhibition of PKK, was evaluated in the FeCl.sub.3-induced inferior vena cava thrombosis mouse model.
Treatment
[1181] Three groups of 8 male BALB/c mice were treated with 10 mg/kg, 20 mg/kg, or 40 mg/kg of ISIS 482584, administered subcutaneously twice a week for 3 weeks (weekly doses of 20 mg/kg, 40 mg/kg, or 80 mg/kg). Two control groups of 12 BALB/c mice each were treated with PBS, administered subcutaneously twice a week for 3 weeks. Two days after the last dose of antisense oligonucleotide or PBS, mice from all groups were anesthetized with 150 mg/kg ketamine mixed with 10 mg/kg xylazine, administered by intraperitoneal injection. Thrombus formation was induced with FeCl.sub.3 in all groups of anesthetized mice except the first control group.
[1182] In mice undergoing FeCl.sub.3 treatment, thrombus formation was induced by applying a piece of filter paper (2.times.4 mm) pre-saturated with 10% FeCl.sub.3 solution directly on the vena cava. After 3 minutes of exposure, the filter paper was removed. Thirty minutes after the filter paper application, a fixed length of the vein containing the thrombus was dissected out for platelet analysis. Liver was collected for RNA analysis.
Quantification of Platelet Composition
[1183] Real-time PCR quantification of platelet factor-4 (PF-4) was used to quantify platelets in the vena cava as a measure of thrombus formation. PF-4 mRNA levels were measured using the murine primer probe set mPF4_LTS_00086 (forward sequence AGACCCATTTCCTCAAGGTAGAACT, designated herein as SEQ ID NO: 2240; reverse sequence CGCAGCGACGCTCATG, designated herein as SEQ ID NO: 2241; probe sequence TCTTTGGGTCCAGTGGCACCCTCTT, designated herein as SEQ ID NO: 2242). Results are presented as a percentage of PF-4 in ISIS oligonucleotide-treated mice, as compared to the two PBS-treated control groups. As shown in Table 202, treatment with ISIS 482584 resulted in a significant reduction of PF-4 in comparison to the PBS control. Therefore, reduction of PKK by the compound provided herein is useful for inhibiting thrombus formation.
TABLE-US-00193 TABLE 202 Analysis of thrombus formation by real-time PCR quantification of PF-4 in the FeCl.sub.3 induced venous thrombosis model Dose in mg/kg/wk PF-4 PBS - FeCl.sub.3 -- 0 PBS + FeCl.sub.3 -- 100 ISIS 482584 20 62 40 34 80 25
Example 133: In Vivo Effect of Antisense Inhibition of Murine PKK in a Tail Bleeding Assay
[1184] Tail-bleeding was measured to observe whether treatment with ISIS 482584 causes excess bleeding or hemorrhage in mice.
Treatment
[1185] Groups of 10 male BALB/c mice were treated with 10 mg/kg, 20 mg/kg, or 40 mg/kg of ISIS 482584, administered subcutaneously twice a week for 3 weeks (weekly doses of 20 mg/kg, 40 mg/kg, or 80 mg/kg). A control group of 8 BALB/c mice was treated with PBS, administered subcutaneously twice a week for 3 weeks.
Tail-Bleeding Assay
[1186] Two days after the final treatment of ISIS oligonucleotides or PBS, mice were placed in a tail bleeding chamber. Mice were anesthetized in the chamber with isoflurane. Then, a small piece of tail (approximately 4 mm from the tip) was cut with sterile scissors. The cut tail was immediately placed in a 15 mL Falcon tube filled with approximately 10 mL of 0.9% NaCl buffer solution warmed to 37.degree. C. The blood was collected over the course of 40 minutes. The saline filled tubes were weighed both before and after bleeding. The results are provided in Table 203.
[1187] Treatment with ISIS 482584 did not significantly affect bleeding. These data suggest that the hemorrhagic potential of the compounds provided herein is low. These data taken with the results provided in Example 19 suggest inhibition of PKK with the compounds described herein are useful for providing antithrombotic activity without associated bleeding risk.
TABLE-US-00194 TABLE 203 Tail bleeding assay after treatment with ISIS 482584 Dose Bleeding (mg/kg/wk) (mL) PBS -- 0.03 ISIS 482584 20 0.03 40 0.14 80 0.07
Example 134: In Vivo Effect of Antisense Inhibition of Murine PKK in the FeCl.sub.3 Induced Mesenteric Thrombosis Model
[1188] ISIS 482584 was evaluated in the FeCl.sub.3 induced mesenteric thrombosis mouse model.
Treatment
[1189] A group of 6-8 Swiss-Webster mice was treated with 40 mg/kg of ISIS 482584, administered subcutaneously twice a week for 3 weeks (weekly dose of 80 mg/kg). A control group of 6 Swiss-Webster mice was treated with PBS, administered subcutaneously twice a week for 3 weeks. Two days after the last dose of antisense oligonucleotide or PBS, mice from all groups were anesthetized with 75 mg/kg ketamine mixed with 25 mg/kg xylazine, administered by subcutaneous injection.
[1190] Rhodamine 6G dye at a dosage of 5 mg/kg was injected subcutaneously to stain platelets. Alexa-647-labeled anti-fibrinogen antibody at a dosage of 1 mg/kg was injected via tail vein injection to stain fibrin. The abdomen was opened by a middle incision. The visceral mesentery was spread on a glass coverslip and the mesenteric arterioles (70-120 .mu.m) were located by observation under a microscope. Thrombus formation was induced by applying of cotton threads (2.times.0.3 mm) pre-saturated with 6% FeCl.sub.3 solution directly on the target vessel. After three minutes of exposure, the thread was removed and the color intensities of both the dyes were recorded by fluorescent microscopy (Olympus FluoView 1000 confocal laser scanning microscope) with appropriate filters for 70 min.
[1191] The results for platelet aggregation in the control and treatment groups are presented in Table 204, expressed in arbitrary units (a.u.). Platelet aggregation was reduced in mice treated with ISIS 482584 at a dose of 80 mg/kg/week as compared to mice treated with PBS. The results for fibrin formation in the control and treatment groups are presented in Table 205, also expressed in arbitrary units (a.u.). Fibrin formation was reduced in mice treated with ISIS 482584 at a dose of 80 mg/kg/week as compared to mice treated with PBS. Therefore, these results suggest that ISIS 482584 inhibits thrombus formation.
TABLE-US-00195 TABLE 204 Analysis of platelet aggregation by real-time measurement of fluorescent intensity (a.u.) in a FeCl.sub.3 induced mesenteric thrombus model Time (sec) PBS 80 mg/kg/wk 752 54 74 1018 315 11 1284 485 7 1550 654 0 1815 1079 0 2081 1164 0 2347 1452 0 2613 1440 38 2879 1689 148 3144 1716 129 3410 1845 169 3676 1865 131 3944 2055 87
TABLE-US-00196 TABLE 205 Analysis of fibrin formation by real-time measurement of fluorescent intensity (a.u.) in a FeCl.sub.3 induced mesenteric thrombus model Time (sec) PBS 80 mg/kg/wk 752 9 54 1018 86 7 1284 203 1 1550 319 10 1815 521 16 2081 598 15 2347 831 61 2613 959 88 2879 1157 141 3144 1236 150 3410 1374 173 3676 1629 160 3944 1822 128
Example 135: In Vivo Effect of Antisense Inhibition of Murine PKK in the Stenosis-Induced Inferior Vena Cava Thrombosis Model
[1192] ISIS 482584 was evaluated in the stenosis-induced inferior vena cava (IVC) thrombosis model. Reduced blood flow and endothelial damage are hallmarks of this model, also known as the St. Tomas model.
Treatment
[1193] Four groups of 6-8 BALB/c mice were treated with 5 mg/kg, 10 mg/kg, 20 mg/kg, or 40 mg/kg of ISIS 482584, administered subcutaneously twice a week for 3 weeks (weekly doses of 10 mg/kg, 20 mg/kg, 40 mg/kg, or 80 mg/kg). A control group of 8 BALB/c mice was treated with PBS, administered subcutaneously twice a week for 3 weeks. Two days after the last dose of antisense oligonucleotide or PBS was administered, mice from all groups were anesthetized with 2.5% inhalant isoflurane. The IVC of the mice was exposed via a midline abdominal incision below the left renal vein, and was separated from the abdominal aorta by blunt dissection. A 6-0 silk tie (Ethicon, UK) was placed behind the blood vessel just below the left renal vein and a metal 4-0 suture (Ethicon, UK) was placed longitudinally over the IVC to tie the silk tie on top. The metal suture was then removed. Two neurovascular surgical clips (Braun Medical Inc, PA) were placed at two separate positions below the ligation for 20 seconds each, after which they were removed. The abdominal cavity contents were then replaced and the abdomen was closed. After 24 hrs, the IVC was exposed and checked for thrombi formation. All thrombi formed were collected and fixed in 10% formalin for 24 hrs.
[1194] The thrombi were weighed and the results are presented in Table 206, expressed in milligrams. As demonstrated by the results, treatment with increasing doses of ISIS 482584 resulted in corresponding decrease in thrombus weight. The results indicate that antisense inhibition of PKK is useful for inhibiting thrombus formation.
TABLE-US-00197 TABLE 206 Thrombi weights in the stenosis-induced IVC thrombosis model Dose in Weight mg/kg/wk (mg) PBS -- 10 ISIS 482584 10 8 20 6 40 5 80 3
Example 136: Inhibition of Murine PKK with an Antisense Oligonucleotide Comprising a GalNAc.sub.3 Conjugate Group
[1195] ISIS 482584 and ISIS 722059, shown in the table below, were tested for their effects on murine PKK mRNA in vivo.
TABLE-US-00198 TABLE 207 ISIS 722059, comprising a GalNAc.sub.3 conjugate group and its parent, ISIS 482584 SEQ Isis No. Sequence (5' to 3') Chemistry ID NO. 482584 G.sub.esG.sub.es.sup.mC.sub.esA.sub.esT.sub.esA.sub.dsT.sub.dsT.sub- .dsG.sub.dsG.sub.dsT.sub.dsT.sub.dsT.sub.dsT.sub.dsT.sub.dsG.sub.esG.sub.e- sA.sub.esA.sub.esT.sub.e No conjugate 2244 group and full PS 722059 GalNAc.sub.3-7.sub.a-o'G.sub.esG.sub.es.sup.mC.sub.eoA.sub.eoT.sub.- esA.sub.dsT.sub.dsT.sub.dsG.sub.dsG.sub.ds 5'-GalNAc.sub.3-7 and 2244 T.sub.dsT.sub.dsT.sub.dsT.sub.dsT.sub.dsG.sub.eoG.sub.eoA.sub.esA.sub.esT- .sub.e mixed PS/PO Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "s" indicates phosphorothioate internucleoside linkages (PS); "o" indicates phosphodiester internucleoside linkages (PO); and "o'" indicates --O--P(.dbd.O)(OH)--. Superscript "m" indicates 5-methylcytosines. The structure of "GalNAc.sub.3-7" is shown in Example 48.
Treatment
[1196] Four groups of four C57Bl/6J-Tyr.sup.c-2J mice each were treated with 5.0 mg/kg, 10.0 mg/kg, 20.0 mg/kg, or 40.0 mg/kg of ISIS 482584, administered subcutaneously twice a week for 3 weeks (weekly doses of 10.0 mg/kg, 20.0 mg/kg, 40.0 mg/kg, or 80.0 mg/kg). Four groups of four BALB/c mice each were treated with 1.0 mg/kg, 2.0 mg/kg, 4.0 mg/kg, or 8.0 mg/kg of ISIS 722059, administered subcutaneously twice a week for 3 weeks (weekly doses of 2.0 mg/kg, 4.0 mg/kg, 8.0 mg/kg, or 16.0 mg/kg). A control group of four BALB/c mice was treated with PBS, administered subcutaneously twice a week for 3 weeks. Three days after the last dose of antisense oligonucleotide or PBS, mice from all groups were anesthetized with vaporized isoflurane in air at 2.5% for induction followed by 1-2% isoflurane by nosecone for maintenance. This was followed by cervical dislocation. Following euthanasia, liver was collected for RNA analysis.
RNA Analysis
[1197] RNA was extracted from liver tissue for real-time PCR analysis of PKK. PKK mRNA levels were measured using the murine primer probe set (forward sequence ACAAGTGCATTTTACAGACCAGAGTAC, designated herein as SEQ ID NO: 2231; reverse sequence GGTTGTCCGCTGACTTTATGCT, designated herein as SEQ ID NO: 2232; probe sequence AAGCACAGTGCAAGCGGAACACCC, designated herein as SEQ ID NO: 2233). Results are presented as percent inhibition of PKK, relative to PBS control. As shown in Table 208 below, Isis 722059, comprising a GalNAc.sub.3 conjugate group, reduced PKK mRNA significantly more potently than the parent antisense oligonucleotide, Isis 482584. This result is consistent with the results in the above examples, in which antisense oligonucleotides comprising a GalNAc.sub.3 conjugate group were significantly more potent than their parent antisense oligonucleotides, for many target genes in both mouse and human. Thus, it is expected that human PKK antisense oligonucleotides comprising a GalNAc.sub.3 conjugate group would likewise reduce human PKK mRNA significantly more potently than their parent antisense oligonucleotides that do not comprise a conjugate group.
TABLE-US-00199 TABLE 208 Percent Inhibition of PKK mRNA in liver relative to the PBS control Dose % ED.sub.50 ISIS No. (mg/kg/week) inhibition (mg/kg/week) 482584 10 42.6 17.2 20 53.3 40 71.4 80 90.8 722059 2 50.1 2.09 4 76.7 8 80.8 16 86.1
Example 137: Inhibition of Human PKK with an Antisense Oligonucleotide Comprising a GalNAc.sub.3 Conjugate Group
[1198] ISIS 546254 and ISIS 721744, shown in the table below, were tested for their effects on human PKK mRNA in vitro.
TABLE-US-00200 TABLE 209 ISIS 721744, comprising a GalNAc.sub.3 conjugate group and its parent, ISIS 546254 SEQ Isis No. Sequence (5' to 3') Chemistry ID NO. 546254 T.sub.esG.sub.es.sup.mC.sub.esA.sub.esA.sub.esG.sub.dsT.sub.ds.sup.- mC.sub.dsT.sub.ds.sup.mC.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.sub.ds No conjugate 570 G.sub.d.sup.mC.sub.dsA.sub.esA.sub.esA.sub.es.sup.mC.sub.esA.sub.e group and full PS 721744 GalNAc.sub.3-7.sub.a-o'T.sub.esG.sub.es.sup.mC.sub.eoA.sub.eoA.sub.- esG.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.ds.sup.mC.sub.dsT.sub.dsT.sub.dsG.su- b.ds 5'-GalNAc.sub.3-7.sub.a-o' 570 G.sub.ds.sup.mC.sub.dsA.sub.eoA.sub.eoA.sub.es.sup.mC.sub.esA.sub.e and mixed PS/PO Subscripts: "e" indicates 2'-MOE modified nucleoside; "d" indicates .beta.-D-2'-deoxyribonucleoside; "s" indicates a phosphorothioate internucleoside linkage (PS); and "o" indicates a phosphodiester internucleoside linkage (PO). Superscript "m" indicates 5-methylcytosine. The structure of "GalNAc.sub.3-7" is shown in Example 48, and "GalNAc.sub.3-7.sub.a-o'" indicates a GalNAc.sub.3-7 conjugate group in which the cleavable moiety is --O--P(.dbd.O)(OH)--.
[1199] Primary human hepatocyte co-cultures that include stromal cells in order to mimic the physiological microenviroment of the liver in vitro (HepatoPac kit HPHU-TX-96S, Hepregen, Medford, Mass.) were used according to the manufacturer's instructions. A concentration of Isis oligonucleotide listed in table below or PBS was added to each well in the absence of any transfection reagent. 96 hours later, cells were lysed and RNA was isolated from the cells. PKK mRNA levels were measured by quantitative real-time PCR using primer probe set RTS3454 and normalized to total RNA content, as measured by RIBOGREEN.RTM.. The results are presented in the table below as percent inhibition of PKK mRNA levels, relative to PBS treated cells; and IC.sub.50 values were calculated using a 4 parameter logistic model (JMP Software, Cary, N.C.). The results show that, under free uptake conditions in which no reagents or electroporation techniques were used to artificially promote entry of the oligonucleotides into cells, the oligonucleotide comprising a GalNAc conjugate was significantly more potent than the parent oligonucleotide that does not comprise a GalNAc conjugate.
TABLE-US-00201 TABLE 210 Percent Inhibition of PKK mRNA relative to the PBS control ISIS No. Concentration (.mu.M) Inhibition (%) IC.sub.50 (.mu.M) 546254 0.1 30 2.12 0.3 25 1.0 24 3.0 63 10.0 85 721744 0.03 34 0.07 0.1 52 0.3 81 1.0 92 3.0 98
Sequence CWU 0 SQTB SEQUENCE LISTING The patent application
contains a lengthy "Sequence Listing" section. A copy of the
"Sequence Listing" is available in electronic form from the USPTO
web site
(http://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200056185A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
0 SQTB SEQUENCE LISTING The patent application contains a lengthy
"Sequence Listing" section. A copy of the "Sequence Listing" is
available in electronic form from the USPTO web site
(http://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200056185A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
References






















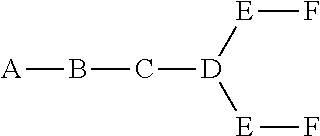




























































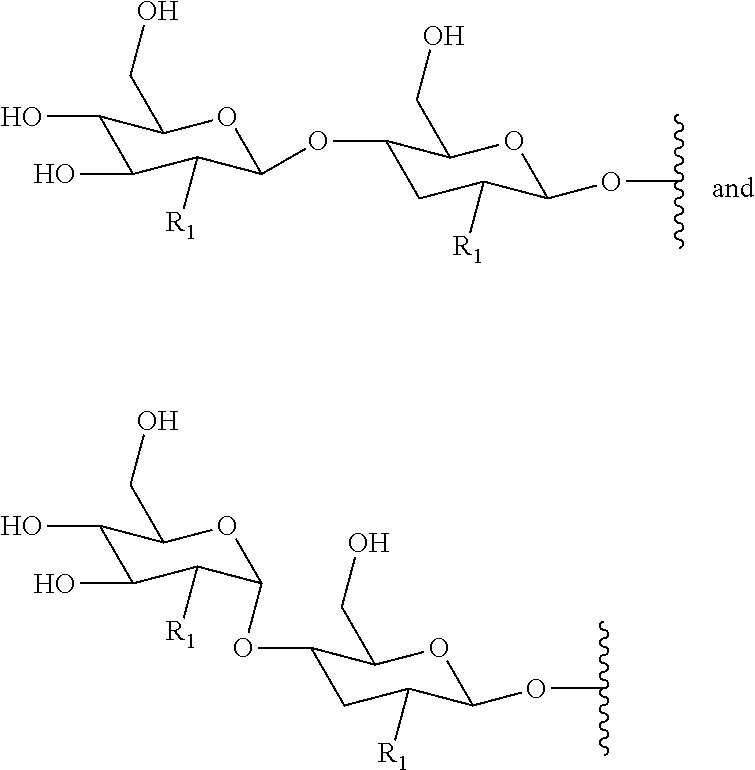



























































































































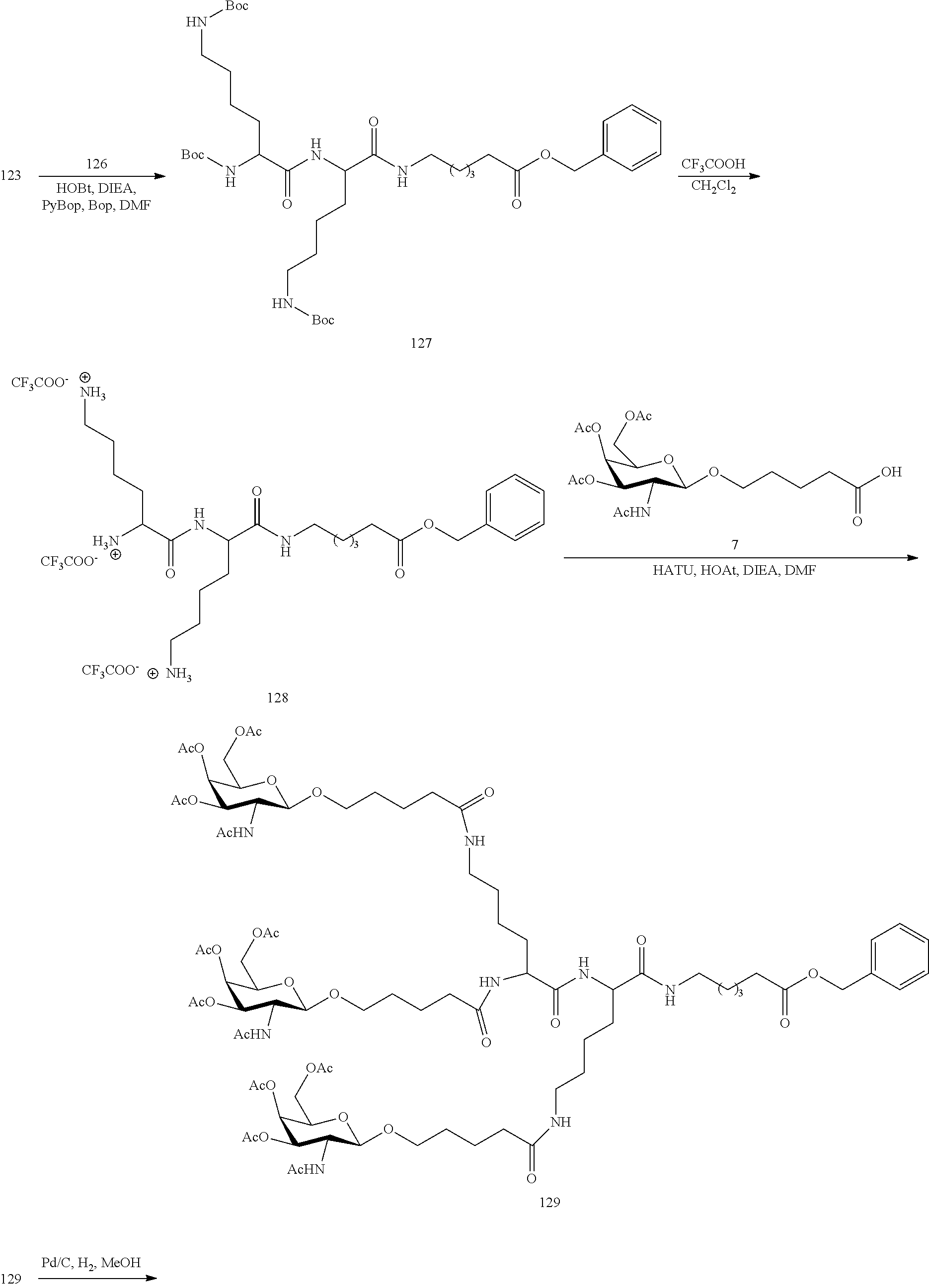








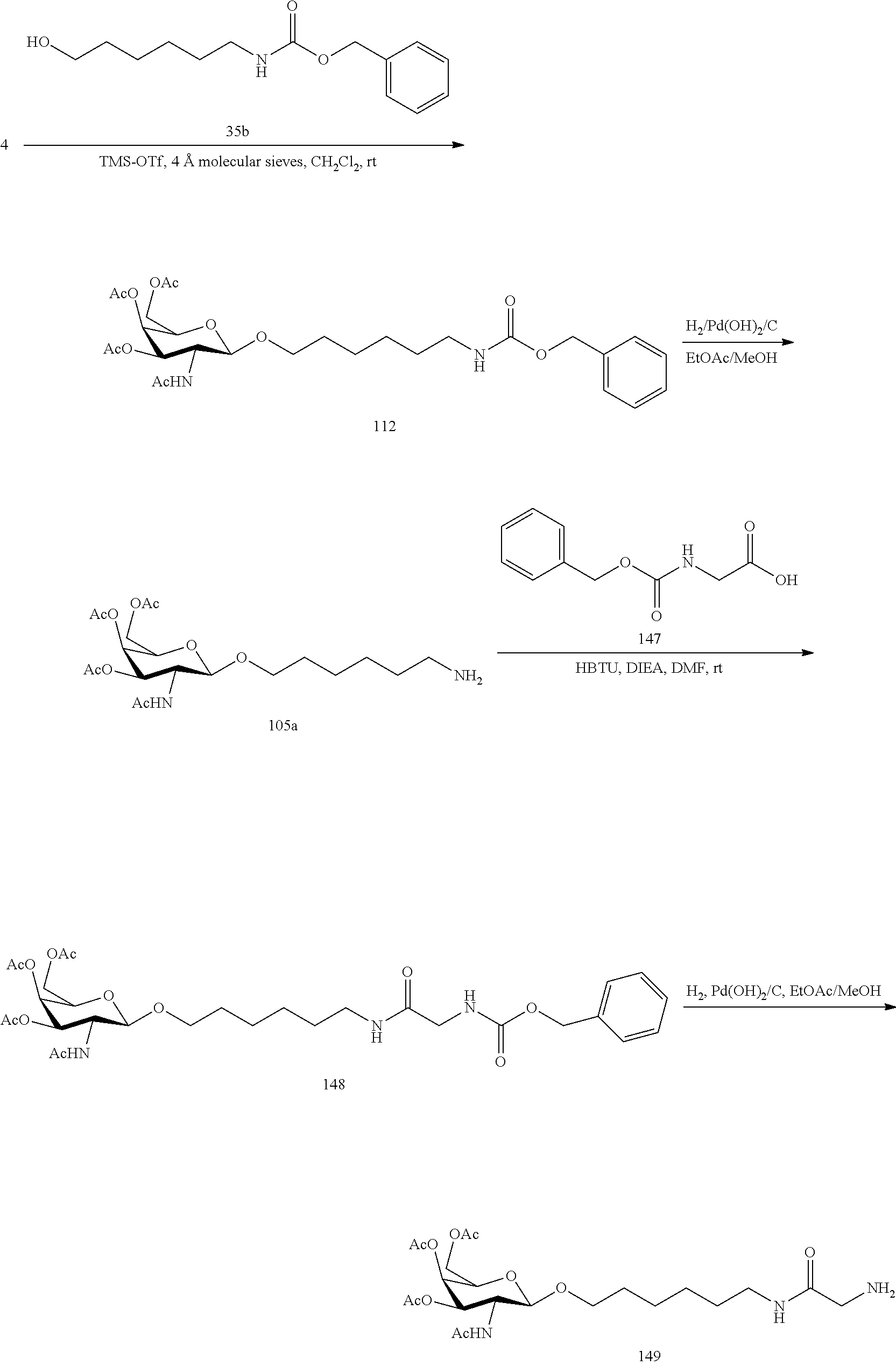























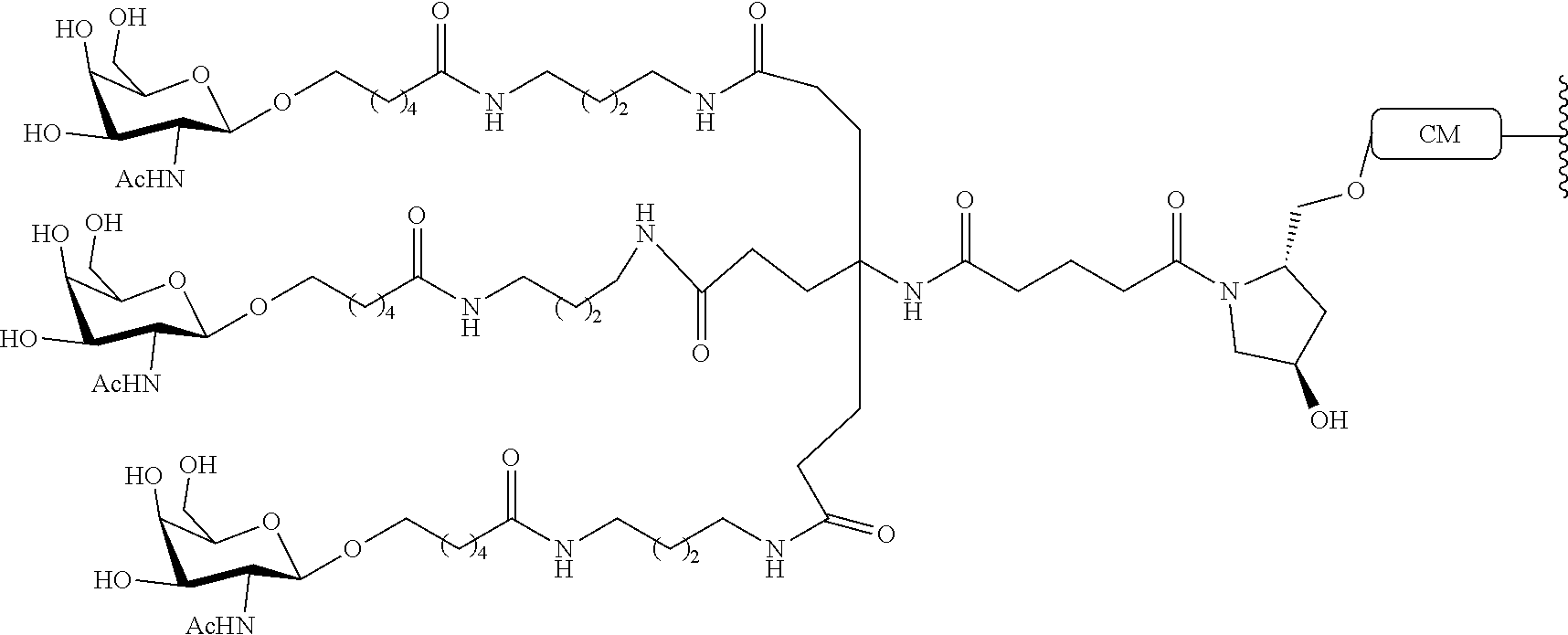









































XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.