Rnai Agents, Compositions And Methods Of Use Thereof For Treating Transthyretin (ttr) Associated Diseases
Rajeev; Kallanthottathil G. ; et al.
U.S. patent application number 16/738014 was filed with the patent office on 2020-10-08 for rnai agents, compositions and methods of use thereof for treating transthyretin (ttr) associated diseases. The applicant listed for this patent is Alnylam Pharmaceuticals, Inc.. Invention is credited to Klaus Charisse, Satyanarayana Kuchimanchi, Martin A. Maier, Muthiah Manoharan, Kallanthottathil G. Rajeev, Tracy Zimmermann.
| Application Number | 20200318111 16/738014 |
| Document ID | / |
| Family ID | 1000004914760 |
| Filed Date | 2020-10-08 |








View All Diagrams
| United States Patent Application | 20200318111 |
| Kind Code | A1 |
| Rajeev; Kallanthottathil G. ; et al. | October 8, 2020 |
RNAI AGENTS, COMPOSITIONS AND METHODS OF USE THEREOF FOR TREATING TRANSTHYRETIN (TTR) ASSOCIATED DISEASES
Abstract
The present invention provides RNAi agents, e.g., double stranded RNAi agents, that target the transthyretin (TTR) gene and methods of using such RNAi agents for treating or preventing TTR-associated diseases.
| Inventors: | Rajeev; Kallanthottathil G.; (Wayland, MA) ; Zimmermann; Tracy; (Winchester, MA) ; Manoharan; Muthiah; (Weston, MA) ; Maier; Martin A.; (Belmont, MA) ; Kuchimanchi; Satyanarayana; (Acton, MA) ; Charisse; Klaus; (Acton, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000004914760 | ||||||||||
| Appl. No.: | 16/738014 | ||||||||||
| Filed: | January 9, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15188317 | Jun 21, 2016 | 10570391 | ||
| 16738014 | ||||
| 14358972 | May 16, 2014 | 9399775 | ||
| PCT/US2012/065691 | Nov 16, 2012 | |||
| 15188317 | ||||
| 61680098 | Aug 6, 2012 | |||
| 61615618 | Mar 26, 2012 | |||
| 61561710 | Nov 18, 2011 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 2310/322 20130101; C12N 15/1136 20130101; C12N 2310/14 20130101; A61K 31/713 20130101; C12N 2310/343 20130101; C12N 2320/30 20130101; C12N 2310/351 20130101; C07H 21/02 20130101; C12N 15/113 20130101; C12N 2310/321 20130101; A61K 31/7125 20130101; C12N 2310/346 20130101; C12N 2310/315 20130101 |
| International Class: | C12N 15/113 20060101 C12N015/113; A61K 31/7125 20060101 A61K031/7125; A61K 31/713 20060101 A61K031/713; C07H 21/02 20060101 C07H021/02 |
Claims
1. A double stranded RNAi agent comprising a sense strand complementary to an antisense strand, wherein said antisense strand comprises a region complementary to part of an mRNA encoding transthyretin (TTR), wherein each strand is independently 15 to 30 nucleotides in length, wherein said double stranded RNAi agent is represented by formula (III): sense: 5'n.sub.p-N.sub.a-(XXX).sub.i-N.sub.b-YYY-N.sub.b-(ZZZ).sub.j-N.sub.a-n.s- ub.q3' antisense: 3'n.sub.p'-N.sub.a'-(X'X'X').sub.k-N.sub.b'-Y'Y'Y'-N.sub.b'-(Z'Z'Z').sub.- i-N.sub.a'-n.sub.q'5' (III) wherein: i, j, k, and l are each independently 0 or 1; p, p', q, and q' are each independently 0-6; each N.sub.a and N.sub.a' independently represents an oligonucleotide sequence comprising 0-25 nucleotides which are either modified or unmodified or combinations thereof, each sequence comprising at least two differently modified nucleotides; each N.sub.b and N.sub.b' independently represents an oligonucleotide sequence comprising 0-10 nucleotides which are either modified or unmodified or combinations thereof; each n.sub.p, n.sub.p', n.sub.q, and n.sub.q' independently represents an overhang nucleotide; XXX, YYY, ZZZ, X'X'X', Y'Y'Y', and Z'Z'Z' each independently represent one motif of three identical modifications on three consecutive nucleotides; modifications on N.sub.b differ from the modification on Y and modifications on N.sub.b' differ from the modification on Y'; and wherein the sense strand is conjugated to at least one ligand.
2. The RNAi agent of claim 1, wherein i is 1; j is 1; or both i and j are 1; or wherein k is 1; l is 1; or both k and l are 1.
3. (canceled)
4. The RNAi agent of claim 1, wherein XXX is complementary to X'X'X', YYY is complementary to Y'Y'Y', and ZZZ is complementary to Z'Z'Z'.
5.-10. (canceled)
11. The RNAi agent of claim 1, wherein the duplex region is 15-30 nucleotide pairs in length 17-23 nucleotide pairs in length; 17-25 nucleotide pairs in length; 23-27 nucleotide pairs in length 19-21 nucleotide pairs in length; or 21-23 nucleotide pairs in length.
12.-17. (canceled)
18. The RNAi agent of claim 1, wherein the modifications on the nucleotides are selected from the group consisting of LNA, HNA, CeNA, 2'-methoxyethyl, 2'-O-alkyl, 2'-O-allyl, 2'-C-allyl, 2'-fluoro, 2'-deoxy, 2'-hydroxyl, and combinations thereof.
19. The RNAi agent of claim 18, wherein the modifications on the nucleotides are 2'-O-methyl, 2'-fluoro or both.
20. The RNAi agent of claim 1, wherein the ligand is one or more GalNAc derivatives attached through a bivalent or trivalent branched linker.
21. The RNAi agent of claim 1, wherein the ligand is ##STR00014## and wherein the ligand is attached to the 3' end of the sense strand.
22. (canceled)
23. (canceled)
24. The RNAi agent of claim 1 further comprising at least one phosphorothioate or methylphosphonate internucleotide linkage.
25.-41. (canceled)
42. A pharmaceutical composition comprising the RNAi agent of claim 1.
43.-49. (canceled)
50. A method of inhibiting expression of a transthyretin (TTR) in a cell comprising contacting said cell with the RNAi agent of claim 1 or with the pharmaceutical composition of claim 42 in an amount effective to inhibit expression of said TTR in said cell, thereby inhibiting expression of said transthyretin (TTR) in said cell.
51. (canceled)
52. (canceled)
53. The method of claim 50, wherein said cell is present within a subject.
54. The method of claim 53, wherein said subject is a human.
55.-78. (canceled)
79. A method of treating or preventing a TTR-associated disease in a subject, comprising administering to said subject a therapeutically effective amount or a prophylactically effective amount of the RNAi agent of claim 1 or the pharmaceutical composition of claim 42, thereby treating or preventing said TTR-associated disease in said subject.
80. (canceled)
81. The method of claim 79, wherein said subject is a human.
82. The method of claim 79, wherein said subject is a subject suffering from a TTR-associated disease.
83. (canceled)
84. The method of claim 79, wherein said subject carries a TTR gene mutation that is associated with the development of a TTR-associated disease.
85. The method of claim 79, wherein said TTR-associated disease is selected from the group consisting of senile systemic amyloidosis (SSA), systemic familial amyloidosis, familial amyloidotic polyneuropathy (FAP), familial amyloidotic cardiomyopathy (FAC), leptomeningeal/Central Nervous System (CNS) amyloidosis, and hyperthyroxinemia.
86. (canceled)
87. (canceled)
88. The method of claim 79, wherein said RNAi agent is administered to said subject via subcutaneous, intramuscular or intravenous administration.
89.-96. (canceled)
97. The method of claim 79, further comprising assessing the level of TTR mRNA expression or TTR protein expression in a sample derived from the subject.
98.-113. (canceled)
Description
RELATED APPLICATIONS
[0001] This application is a continuation of U.S. patent application Ser. No. 15/188,317, filed on Jun. 21, 2016, which is a continuation of U.S. patent application Ser. No. 14/358,972, filed on May 16, 2014, now U.S. Pat. No. 9,399,775, issued on Jul. 26, 2016, which is a 35 U.S.C. .sctn. 371 national stage filing of International Application No. PCT/US2012/065691, filed on Nov. 16, 2012, which claims priority to U.S. Provisional Application No. 61/561,710, filed on Nov. 18, 2011, U.S. Provisional Application No. 61/615,618, filed on Mar. 26, 2012, and U.S. Provisional Application No. 61/680,098, filed on Aug. 6, 2012. The entire contents of each of the foregoing application are hereby incorporated herein by reference.
SEQUENCE LISTING
[0002] The instant application contains a Sequence Listing which has been submitted in ASCII format via EFS-Web and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Jan. 8, 2020, is named 121301_00105_SL.txt and is 541,680 bytes in size.
BACKGROUND OF THE INVENTION
[0003] Transthyretin (TTR) (also known as prealbumin) is found in serum and cerebrospinal fluid (CSF). TTR transports retinol-binding protein (RBP) and thyroxine (T4) and also acts as a carrier of retinol (vitamin A) through its association with RBP in the blood and the CSF. Transthyretin is named for its transport of thyroxine and retinol. TTR also functions as a protease and can cleave proteins including apoA-I (the major HDL apolipoprotein), amyloid .beta.-peptide, and neuropeptide Y. See Liz, M. A. et al. (2010) IUBMB Life, 62(6):429-435.
[0004] TTR is a tetramer of four identical 127-amino acid subunits (monomers) that are rich in beta sheet structure. Each monomer has two 4-stranded beta sheets and the shape of a prolate ellipsoid. Antiparallel beta-sheet interactions link monomers into dimers. A short loop from each monomer forms the main dimer-dimer interaction. These two pairs of loops separate the opposed, convex beta-sheets of the dimers to form an internal channel.
[0005] The liver is the major site of TTR expression. Other significant sites of expression include the choroid plexus, retina (particularly the retinal pigment epithelium) and pancreas.
[0006] Transthyretin is one of at least 27 distinct types of proteins that is a precursor protein in the formation of amyloid fibrils. See Guan, J. et al. (Nov. 4, 2011) Current perspectives on cardiac amyloidosis, Am J Physiol Heart Circ Physiol, doi:10.1152/ajpheart.00815.2011. Extracellular deposition of amyloid fibrils in organs and tissues is the hallmark of amyloidosis. Amyloid fibrils are composed of misfolded protein aggregates, which may result from either excess production of or specific mutations in precursor proteins. The amyloidogenic potential of TTR may be related to its extensive beta sheet structure; X-ray crystallographic studies indicate that certain amyloidogenic mutations destabilize the tetrameric structure of the protein. See, e.g., Saraiva M. J. M. (2002) Expert Reviews in Molecular Medicine, 4(12):1-11.
[0007] Amyloidosis is a general term for the group of amyloid diseases that are characterized by amyloid deposits. Amyloid diseases are classified based on their precursor protein; for example, the name starts with "A" for amyloid and is followed by an abbreviation of the precursor protein, e.g., ATTR for amloidogenic transthyretin. Ibid.
[0008] There are numerous TTR-associated diseases, most of which are amyloid diseases. Normal-sequence TTR is associated with cardiac amyloidosis in people who are elderly and is termed senile systemic amyloidosis (SSA) (also called senile cardiac amyloidosis (SCA) or cardiac amyloidosis). SSA often is accompanied by microscopic deposits in many other organs. TTR amyloidosis manifests in various forms. When the peripheral nervous system is affected more prominently, the disease is termed familial amyloidotic polyneuropathy (FAP). When the heart is primarily involved but the nervous system is not, the disease is called familial amyloidotic cardiomyopathy (FAC). A third major type of TTR amyloidosis is leptomeningeal amyloidosis, also known as leptomeningeal or meningocerebrovascular amyloidosis, central nervous system (CNS) amyloidosis, or amyloidosis VII form.
[0009] Mutations in TTR may also cause amyloidotic vitreous opacities, carpal tunnel syndrome, and euthyroid hyperthyroxinemia, which is a non-amyloidotic disease thought to be secondary to an increased association of thyroxine with TTR due to a mutant TTR molecule with increased affinity for thyroxine. See, e.g., Moses et al. (1982) J. Clin. Invest., 86, 2025-2033.
[0010] Abnormal amyloidogenic proteins may be either inherited or acquired through somatic mutations. Guan, J. et al. (Nov. 4, 2011) Current perspectives on cardiac amyloidosis, Am J Physiol Heart Circ Physiol, doi:10.1152/ajpheart.00815.2011. Transthyretin associated ATTR is the most frequent form of hereditary systemic amyloidosis. Lobato, L. (2003) J. Nephrol., 16:438-442. TTR mutations accelerate the process of TTR amyloid formation and are the most important risk factor for the development of ATTR. More than 85 amyloidogenic TTR variants are known to cause systemic familial amyloidosis. TTR mutations usually give rise to systemic amyloid deposition, with particular involvement of the peripheral nervous system, although some mutations are associated with cardiomyopathy or vitreous opacities. Ibid.
[0011] The V30M mutation is the most prevalent TTR mutation. See, e.g., Lobato, L. (2003) JNephrol, 16:438-442. The V122I mutation is carried by 3.9% of the African American population and is the most common cause of FAC. Jacobson, D. R. et al. (1997) N. Engl. J. Med. 336 (7): 466-73. It is estimated that SSA affects more than 25% of the population over age 80. Westermark, P. et al. (1990) Proc. Natl. Acad. Sci. U.S.A. 87 (7): 2843-5.
[0012] Accordingly, there is a need in the art for effective treatments for TTR-associated diseases.
SUMMARY OF THE INVENTION
[0013] The present invention provides RNAi agents, e.g., double stranded RNAi agents, targeting the Transthyretin (TTR) gene. The present invention also provides methods of inhibiting expression of TTR and methods of treating or preventing a TTR-associated disease in a subject using the RNAi agents, e.g. double stranded RNAi agents, of the invention. The present invention is based, at least in part, on the discovery that RNAi agents that comprise particular chemical modifications show a superior ability to inhibit expression of TTR. Agents including a certain pattern of chemical modifications (e.g., an alternating pattern) and a ligand are shown herein to be effective in silencing the activity of the TTR gene. Furthermore, agents including one or more motifs of three identical modifications on three consecutive nucleotides, including one such motif at or near the cleavage site of the agents, show surprisingly enhanced TTR gene silencing activity. When a single such chemical motif is present in the agent, it is preferred to be at or near the cleavage region for enhancing of the gene silencing activity. Cleavage region is the region surrounding the cleavage site, i.e., the site on the target mRNA at which cleavage occurs.
[0014] Accordingly, in one aspect, the present invention features RNAi agents, e.g., double stranded RNAi agents, for inhibiting expression of a transthyretin (TTR). The double stranded RNAi agent includes a sense strand complementary to an antisense strand. The antisense strand includes a region complementary to a part of an mRNA encoding transthyretin. Each strand has 14 to 30 nucleotides, and the double stranded RNAi agent is represented by formula (III):
sense: 5'n.sub.p-N.sub.a-(XXX).sub.i-N.sub.b-YYY-N.sub.b-(ZZZ).sub.j-N.s- ub.a-n.sub.q3'
antisense: 3'n.sub.p'-N.sub.a'-(X'X'X').sub.k-N.sub.b'-Y'Y'Y'-N.sub.b'-(Z'Z'Z').sub.- i-N.sub.a'-n.sub.q'5' (III).
[0015] In Formula III, i, j, k, and l are each independently 0 or 1; p, p', q, and q' are each independently 0-6; each N.sub.a and N.sub.a' independently represents an oligonucleotide sequence including 0-25 nucleotides which are either modified or unmodified or combinations thereof, each sequence including at least two differently modified nucleotides; each N.sub.b and N.sub.b' independently represents an oligonucleotide sequence including 0-10 nucleotides which are either modified or unmodified or combinations thereof, each n.sub.p, n.sub.p', n.sub.q, and n.sub.q' independently represents an overhang nucleotide; XXX, YYY, ZZZ, X'X'X', Y'Y'Y', and Z'Z'Z' each independently represents one motif of three identical modifications on three consecutive nucleotides; modifications on N.sub.b differ from the modification on Y and modifications on N.sub.b' differ from the modification on Y'. In some embodiments, the sense strand is conjugated to at least one ligand, e.g., at least one ligand, e.g., at least one ligand attached to the 3' end of the sense strand. In other embodiments, the ligand may be conjugated to the antisense strand.
[0016] In some embodiments, i is 1; j is 1; or both i and j are 1.
[0017] In some embodiments, k is 1; l is 1; or both k and l are 1.
[0018] In some embodiments, i is 0; j is 1.
[0019] In some embodiments, i is 1, j is 0.
[0020] In some embodiments, k is 0; l is 1.
[0021] In some embodiments, k is 1; l is 0.
[0022] In some embodiments, XXX is complementary to X'X'X', YYY is complementary to Y'Y'Y', and ZZZ is complementary to Z'Z'Z'.
[0023] In some embodiments, the YYY motif occurs at or near the cleavage site of the sense strand.
[0024] In some embodiments, the Y'Y'Y' motif occurs at the 11, 12 and 13 positions of the antisense strand from the 5'-end.
[0025] In some embodiments, the Y' is 2'-O-methyl.
[0026] In some embodiments, the Y' is 2'-fluoro.
[0027] In some embodiments, formula (III) is represented as formula (IIIa):
sense: 5'n.sub.p-N.sub.a-YYY-N.sub.b-ZZZ-N.sub.a-n.sub.q3'
antisense: 3'n.sub.p'-N.sub.a'-Y'Y'Y'-N.sub.b'-Z'Z'Z'-N.sub.a'n.sub.q'5' (IIIa).
In formula IIIa, each N.sub.b and N.sub.b' independently represents an oligonucleotide sequence including 1-5 modified nucleotides.
[0028] In some embodiments, formula (III) is represented as formula (IIIb):
sense: 5'n.sub.p-N.sub.a-XXX-N.sub.b-YYY-N.sub.a-n.sub.q3'
antisense: 3'n.sub.p'-N.sub.a'-X'X'X'-N.sub.b'-Y'Y'Y'-N.sub.a'-n.sub.q'5' (IIIb).
In formula IIIb each N.sub.b and N.sub.b' independently represents an oligonucleotide sequence including 1-5 modified nucleotides.
[0029] In some embodiments, formula (III) is represented as formula (IIIc):
sense: 5'n.sub.p-N.sub.a-XXX-N.sub.b-YYY-N.sub.b-ZZZ-N.sub.a-n.sub.q3'
antisense: 3'n.sub.p'-N.sub.a'-X'X'X'-N.sub.b'-Y'Y'Y'-N.sub.b'-Z'Z'Z'-N.sub.a'-n.sub- .q'5' (IIIc).
In formula IIIc, each N.sub.b and -N.sub.b' independently represents an oligonucleotide sequence including 1-5 modified nucleotides and each N.sub.a and N.sub.a' independently represents an oligonucleotide sequence including 2-10 modified nucleotides.
[0030] In many embodiments, the duplex region is 15-30 nucleotide pairs in length. In some embodiments, the duplex region is 17-23 nucleotide pairs in length, 17-25 nucleotide pairs in length, 23-27 nucleotide pairs in length, 19-21 nucleotide pairs in length, or 21-23 nucleotide pairs in length.
[0031] In certain embodiments, each strand has 15-30 nucleotides.
[0032] In some embodiments, the modifications on the nucleotides are selected from the group consisting of LNA, HNA, CeNA, 2'-methoxyethyl, 2'-O-alkyl, 2'-O-allyl, 2'-C-allyl, 2'-fluoro, 2'-deoxy, 2'-hydroxyl, and combinations thereof. In some preferred embodiments, the modifications on the nucleotides are 2'-O-methyl or 2'-fluoro.
[0033] In some embodiments, the ligand is one or more N-acetylgalactosamine (GalNAc) derivatives attached through a bivalent or trivalent branched linker. In particular embodiments, the ligand is
##STR00001##
[0034] In some embodiments, the ligand is attached to the 3' end of the sense strand.
[0035] In some embodiments, the RNAi agent is conjugated to the ligand as shown in the following schematic
##STR00002##
[0036] wherein X is O or S.
[0037] In some embodiments, the RNAi agent is conjugated to the ligand as shown in the following schematic
##STR00003##
[0038] In some embodiments, the RNAi agent further includes at least one phosphorothioate or methylphosphonate internucleotide linkage. In some embodiments, the phosphorothioate or methylphosphonate internucleotide linkage is at the 3'-terminal of one strand. In some embodiments, the strand is the antisense strand. In other embodiments, the strand is the sense strand.
[0039] In certain embodiments, the base pair at the 1 position of the 5'-end of the duplex is an AU base pair.
[0040] In some embodiments, the Y nucleotides contain a 2'-fluoro modification.
[0041] In some embodiments, the Y' nucleotides contain a 2'-O-methyl modification.
[0042] In some embodiments, p'>0. In some such embodiments, each n is complementary to the target mRNA. In other such embodiments, each n is non-complementary to the target mRNA. In some embodiments, p, p', q and q' are 1-6. In some preferred embodiments, p'=1 or 2. In some preferred embodiments, p'=2. In some such embodiments, q'=0, p=0, q=0, and p' overhang nucleotides are complementary to the target mRNA. In other such embodiments, q'=0, p=0, q=0, and p' overhang nucleotides are non-complementary to the target mRNA.
[0043] In some embodiments, the sense strand has a total of 21 nucleotides and the antisense strand has a total of 23 nucleotides.
[0044] In certain embodiments, linkages between n.sub.p' include phosphorothioate linkages. In some such embodiments, the linkages between n.sub.p' are phosphorothioate linkages.
[0045] In some embodiments, the RNAi agent is selected from the group of agents listed in Table 1.
[0046] In preferred embodiments, the RNAi agent is selected from the group consisting of AD-51544, AD-51545, AD-51546, and AD-51547.
[0047] In an even more preferred embodiment, the RNAi agent is AD-51547 having the following structure:
TABLE-US-00001 sense: (SEQ ID NO: 2) 5'- UfgGfgAfuUfuCfAfUfgUfaacCfaAfgAfL96-3' antisense: (SEQ ID NO: 3) 5'- uCfuUfgGfUfUfaCfaugAfaAfuCfcCfasUfsc-3'
wherein lowercase nucleotides (a, u, g, c) indicate 2'-O-methyl nucleotides; Nf (e.g., Af) indicates a 2'-fluoro nucleotide; s indicates a phosphothiorate linkage; L96 indicates a GalNAc.sub.3 ligand.
[0048] In another aspect, the present invention features a cell containing the RNAi agent for inhibiting expression of TTR.
[0049] In a further aspect, the present invention features a pharmaceutical composition comprising an RNAi agent for inhibiting expression of TTR. In some embodiments, the pharmaceutical composition is a solution comprising the RNAi agent. In some embodiments, the solution comprising the RNAi agent is an unbuffered solution, e.g., saline solution or water. In other embodiments, the solution is a buffered solution, e.g., a solution of phosphate buffered saline (PBS). In other embodiments, the pharmaceutical composition is a liposome or a lipid formulation. In some embodiments, the lipid formulation comprises a XTC or MC3.
[0050] In yet another aspect, the present invention features methods of inhibiting expression of transthyretin (TTR) in a cell. The methods include contacting a cell with an RNAi agent, e.g., a double stranded RNAi agent, in an amount effective to inhibit expression of TTR in the cell, thereby inhibiting expression of TTR in the cell.
[0051] In some embodiments, the expression of TTR is inhibited by at least about 10%, at least about 20%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 70%, at least about 80%, or at least about 90%.
[0052] In other embodiments, the cell is contacted in vitro with the RNAi agent. In other embodiments, the cell is present within a subject. In preferred embodiments, the subject is a human.
[0053] In further embodiments, the subject is a subject suffering from a TTR-associated disease and the effective amount is a therapeutically effective amount. In other embodiments, the subject is a subject at risk for developing a TTR-associated disease and the effective amount is a prophylactically effective amount. In some embodiments, a subject at risk for developing a TTR-associated disease is a subject who carries a TTR gene mutation that is associated with the development of a TTR-associated disease.
[0054] In certain embodiments, the TTR-associated disease is selected from the group consisting of senile systemic amyloidosis (SSA), systemic familial amyloidosis, familial amyloidotic polyneuropathy (FAP), familial amyloidotic cardiomyopathy (FAC), leptomeningeal/Central Nervous System (CNS) amyloidosis, and hyperthyroxinemia.
[0055] In some embodiments, the subject has a TTR-associated amyloidosis and the method reduces an amyloid TTR deposit in the subject.
[0056] In other embodiments, the RNAi agent is administered to the subject by an administration means selected from the group consisting of subcutaneous, intravenous, intramuscular, intrabronchial, intrapleural, intraperitoneal, intraarterial, lymphatic, cerebrospinal, and any combinations thereof. In certain embodiments, the RNAi agent is administered to the subject via subcutaneous or intravenous administration. In preferred embodiments, the RNAi agent is administered to the subject via subcutaneous administration. In some such embodiments, the subcutaneous administration includes administration via a subcutaneous pump or subcutaneous depot.
[0057] In certain embodiments, the RNAi agent is administered to the subject such that the RNAi agent is delivered to a specific site within the subject. In some embodiments, the site is selected from the group consisting of liver, choroid plexus, retina, and pancreas. In preferred embodiments, the site is the liver. In some embodiments, the delivery of the RNAi agent is mediated by asialoglycoprotein receptor (ASGP-R) present in hepatocytes.
[0058] In some embodiments, the RNAi agent is administered at a dose of between about 0.25 mg/kg to about 50 mg/kg, e.g., between about 0.25 mg/kg to about 0.5 mg/kg, between about 0.25 mg/kg to about 1 mg/kg, between about 0.25 mg/kg to about 5 mg/kg, between about 0.25 mg/kg to about 10 mg/kg, between about 1 mg/kg to about 10 mg/kg, between about 5 mg/kg to about 15 mg/kg, between about 10 mg/kg to about 20 mg/kg, between about 15 mg/kg to about 25 mg/kg, between about 20 mg/kg to about 30 mg/kg, between about 25 mg/kg to about 35 mg/kg, or between about 40 mg/kg to about 50 mg/kg.
[0059] In some embodiments, the RNAi agent is administered at a dose of about 0.25 mg/kg, about 0.5 mg/kg, about 1 mg/kg, about 2 mg/kg, about 3 mg/kg, about 4 mg/kg, about 5 mg/kg, about 6 mg/kg, about 7 mg/kg, about 8 mg/kg, about 9 mg/kg, about 10 mg/kg, about 11 mg/kg, about 12 mg/kg, about 13 mg/kg, about 14 mg/kg, about 15 mg/kg, about 16 mg/kg, about 17 mg/kg, about 18 mg/kg, about 19 mg/kg, about 20 mg/kg, about 21 mg/kg, about 22 mg/kg, about 23 mg/kg, about 24 mg/kg, about 25 mg/kg, about 26 mg/kg, about 27 mg/kg, about 28 mg/kg, about 29 mg/kg, 30 mg/kg, about 31 mg/kg, about 32 mg/kg, about 33 mg/kg, about 34 mg/kg, about 35 mg/kg, about 36 mg/kg, about 37 mg/kg, about 38 mg/kg, about 39 mg/kg, about 40 mg/kg, about 41 mg/kg, about 42 mg/kg, about 43 mg/kg, about 44 mg/kg, about 45 mg/kg, about 46 mg/kg, about 47 mg/kg, about 48 mg/kg, about 49 mg/kg or about 50 mg/kg.
[0060] In some embodiments, the RNAi agent is administered in two or more doses. In particular embodiments, the RNAi agent is administered at intervals selected from the group consisting of once every about 2 hours, once every about 3 hours, once every about 4 hours, once every about 6 hours, once every about 8 hours, once every about 12 hours, once every about 24 hours, once every about 48 hours, once every about 72 hours, once every about 96 hours, once every about 120 hours, once every about 144 hours, once every about 168 hours, once every about 240 hours, once every about 336 hours, once every about 504 hours, once every about 672 hours and once every about 720 hours.
[0061] In other embodiments, the method further includes assessing the level of TTR mRNA expression or TTR protein expression in a sample derived from the subject.
[0062] In preferred embodiments, administering the RNAi agent does not result in an inflammatory response in the subject as assessed based on the level of a cytokine or chemokine selected from the group consisting of G-CSF, IFN-.gamma., IL-10, IL-12 (p70), IL1.beta., IL-1ra, IL-6, IL-8, IP-10, MCP-1, MIP-1.alpha., MIP-1.beta., TNF.alpha., and any combinations thereof, in a sample from the subject.
[0063] In some embodiments, the RNAi agent is administered using a pharmaceutical composition
[0064] In preferred embodiments, the RNAi agent is administered in a solution. In some such embodiments, the siRNA is administered in an unbuffered solution. In one embodiment, the siRNA is administered in water. In other embodiments, the siRNA is administered with a buffer solution, such as an acetate buffer, a citrate buffer, a prolamine buffer, a carbonate buffer, or a phosphate buffer or any combination thereof. In some embodiments, the buffer solution is phosphate buffered saline (PBS).
[0065] In another embodiment, the pharmaceutical composition is a liposome or a lipid formulation comprising SNALP or XTC. In one embodiment, the lipid formulation comprises an MC3.
[0066] In another aspect, the invention provides methods of treating or preventing a TTR-associated disease in a subject. The methods include administering to the subject a therapeutically effective amount or prophylactically effective amount of an RNAi agent, e.g., a double stranded RNAi agent, thereby treating or preventing the TTR-associated disease in the subject.
[0067] In some embodiments, TTR expression in a sample derived from the subject is inhibited by at least about 10%, at least about 20%, at least about 30%, at least about 40%, at least about 50%, at least about 60% or at least about 70% at least about 80%, or at least about 90%.
[0068] In some embodiments, the subject is a human.
[0069] In some embodiments, the subject is a subject suffering from a TTR-associated disease. In other embodiments, the subject is a subject at risk for developing a TTR-associated disease.
[0070] In some embodiments, the subject is a subject who carries s a TTR gene mutation that is associated with the development of a TTR-associated disease.
[0071] In certain embodiments, the TTR-associated disease is selected from the group consisting of senile systemic amyloidosis (SSA), systemic familial amyloidosis, familial amyloidotic polyneuropathy (FAP), familial amyloidotic cardiomyopathy (FAC), leptomeningeal/Central Nervous System (CNS) amyloidosis, and hyperthyroxinemia.
[0072] In some embodiments, the subject has a TTR-associated amyloidosis and the method reduces an amyloid TTR deposit in the subject.
[0073] In some embodiments, the RNAi agent is administered to the subject by an administration means selected from the group consisting of subcutaneous, intravenous, intramuscular, intrabronchial, intrapleural, intraperitoneal, intraarterial, lymphatic, cerebrospinal, and any combinations thereof. In certain embodiments, the RNAi agent is administered to the subject via subcutaneous or intravenous administration. In preferred embodiments, the RNAi agent is administered to the subject via subcutaneous administration. In some such embodiments, the subcutaneous administration includes administration via a subcutaneous pump or subcutaneous depot.
[0074] In certain embodiments, the RNAi agent is administered to the subject such that the RNAi agent is delivered to a specific site within the subject. In some such embodiments, the site is selected from the group consisting of liver, choroid plexus, retina, and pancreas. In preferred embodiments, the site is the liver. In some embodiments, the delivery of the RNAi agent is mediated by asialoglycoprotein receptor (ASGP-R) present in hepatocytes.
[0075] In some embodiments, the RNAi agent is administered at a dose of between about 0.25 mg/kg to about 50 mg/kg, e.g., between about 0.25 mg/kg to about 0.5 mg/kg, between about 0.25 mg/kg to about 1 mg/kg, between about 0.25 mg/kg to about 5 mg/kg, between about 0.25 mg/kg to about 10 mg/kg, between about 1 mg/kg to about 10 mg/kg, between about 5 mg/kg to about 15 mg/kg, between about 10 mg/kg to about 20 mg/kg, between about 15 mg/kg to about 25 mg/kg, between about 20 mg/kg to about 30 mg/kg, between about 25 mg/kg to about 35 mg/kg, or between about 40 mg/kg to about 50 mg/kg.
[0076] In some embodiments, the RNAi agent is administered at a dose of about 0.25 mg/kg, about 0.5 mg/kg, about 1 mg/kg, about 2 mg/kg, about 3 mg/kg, about 4 mg/kg, about 5 mg/kg, about 6 mg/kg, about 7 mg/kg, about 8 mg/kg, about 9 mg/kg, about 10 mg/kg, about 11 mg/kg, about 12 mg/kg, about 13 mg/kg, about 14 mg/kg, about 15 mg/kg, about 16 mg/kg, about 17 mg/kg, about 18 mg/kg, about 19 mg/kg, about 20 mg/kg, about 21 mg/kg, about 22 mg/kg, about 23 mg/kg, about 24 mg/kg, about 25 mg/kg, about 26 mg/kg, about 27 mg/kg, about 28 mg/kg, about 29 mg/kg, 30 mg/kg, about 31 mg/kg, about 32 mg/kg, about 33 mg/kg, about 34 mg/kg, about 35 mg/kg, about 36 mg/kg, about 37 mg/kg, about 38 mg/kg, about 39 mg/kg, about 40 mg/kg, about 41 mg/kg, about 42 mg/kg, about 43 mg/kg, about 44 mg/kg, about 45 mg/kg, about 46 mg/kg, about 47 mg/kg, about 48 mg/kg, about 49 mg/kg or about 50 mg/kg.
[0077] In some embodiments, the RNAi agent is administered in two or more doses. In particular embodiments, the RNAi agent is administered at intervals selected from the group consisting of once every about 2 hours, once every about 3 hours, once every about 4 hours, once every about 6 hours, once every about 8 hours, once every about 12 hours, once every about 24 hours, once every about 48 hours, once every about 72 hours, once every about 96 hours, once every about 120 hours, once every about 144 hours, once every about 168 hours, once every about 240 hours, once every about 336 hours, once every about 504 hours, once every about 672 hours and once every about 720 hours.
[0078] In other embodiments, the method further includes assessing the level of TTR mRNA expression or TTR protein expression in a sample derived from the subject.
[0079] In preferred embodiments, administering the RNAi agent does not result in an inflammatory response in the subject as assessed based on the level of a cytokine or chemokine selected from the group consisting of G-CSF, IFN-.gamma., IL-10, IL-12 (p70), IL1.beta., IL-1ra, IL-6, IL-8, IP-10, MCP-1, MIP-1.alpha., MIP-1.beta., TNF.alpha., and any combinations thereof, in a sample from the subject.
[0080] In some embodiments, the RNAi agent is administered using a pharmaceutical composition, e.g., a liposome.
[0081] In some embodiments, the RNAi agent is administered in a solution. In some such embodiments, the siRNA is administered in an unbuffered solution. In one embodiment, the siRNA is administered in saline or water. In other embodiments, the siRNA is administered with a buffer solution, such as an acetate buffer, a citrate buffer, a prolamine buffer, a carbonate buffer, or a phosphate buffer or any combination thereof. In some embodiments, the buffer solution is phosphate buffered saline (PBS).
[0082] In another aspect, the present invention provides a method of inhibiting expression of transthyretin (TTR) in a cell, including contacting a cell with an RNAi agent, e.g., a double stranded RNAi agent, in an amount effective to inhibit expression of TTR in the cell. In one aspect, the double stranded RNAi agent is selected from the group of agents listed in Table 1, thereby inhibiting expression of transthyretin (TTR) in the cell.
[0083] In another aspect, the present invention provides a method of inhibiting expression of transthyretin (TTR) in a cell, including contacting a cell with an RNAi agent, e.g., a double stranded RNAi agent, in an amount effective to inhibit expression of TTR in the cell. In one aspect, the double stranded RNAi agent is selected from the group consisting of AD-51544, AD-51545, AD-51546, and AD-51547, thereby inhibiting expression of transthyretin (TTR) in the cell.
[0084] In a further aspect, the present invention provides a method of treating or preventing a TTR-associated disease in a subject, including administering to the subject a therapeutically effective amount or a prophylactically effective amount of an RNAi agent, e.g., a double stranded RNAi agent. In one aspect, the double stranded RNAi agent is selected from the group of agents listed in Table 1, thereby treating or preventing a TTR-associated disease in the subject.
[0085] In yet another aspect, the present invention provides a method of treating or preventing a TTR-associated disease in a subject, including administering to the subject a therapeutically effective amount or a prophylactically effective amount of an RNAi agent, e.g., a double stranded RNAi agent. In one aspect, the double stranded RNAi agent is selected from the group consisting of AD-51544, AD-51545, AD-51546, and AD-51547, thereby treating or preventing a TTR-associated disease in the subject.
[0086] In further aspects, the invention provides kits for performing the methods of the invention. In one aspect, the invention provides a kit for performing a method of inhibiting expression of transthyretin (TTR) in a cell comprising contacting a cell with an RNAi agent, e.g., a double stranded RNAi agent, in an amount effective to inhibit expression of said TTR in said cell, thereby inhibiting the expression of TTR in the cell. The kit comprises an RNAi agent and instructions for use and, optionally, means for administering the RNAi agent to the subject.
[0087] The present invention is further illustrated by the following detailed description and drawings.
BRIEF DESCRIPTION OF THE DRAWINGS
[0088] FIG. 1 is a graph depicting that administering to mice a single subcutaneous dose of a GalNAc-conjugated RNAi agent targeting TTR resulted in dose-dependent suppression of TTR mRNA.
[0089] FIG. 2 is a graph depicting that administering to mice a single subcutaneous dose of 7.5 mg/kg or 30 mg/kg of a GalNAc conjugated RNAi agent targeting TTR resulted in long lasting suppression of TTR mRNA.
[0090] FIG. 3 depicts the human TTR mRNA sequence.
[0091] FIG. 4 is a graph depicting improved silencing activity of RNAi agents modified relative to the parent AD-45163.
[0092] FIG. 5 is a graph depicting improved silencing activity of RNAi agents modified relative to the parent AD-45165.
[0093] FIG. 6 is a graph depicting improved free uptake silencing following 4 hour incubation with RNAi agents modified relative to the parent AD-45163.
[0094] FIG. 7 is a graph depicting improved free uptake silencing following 24 hour incubation with RNAi agents modified relative to the parent AD-45163.
[0095] FIG. 8 is a graph depicting improved free uptake silencing following 4 hour incubation with RNAi agents modified relative to the parent AD-45165.
[0096] FIG. 9 is a graph depicting improved free uptake silencing following 24 hour incubation with RNAi agents modified relative to the parent AD-45165.
[0097] FIG. 10A is a graph depicting silencing of TTR mRNA in transgenic mice that express hTTR V30M following administration of a single subcutaneous dose of RNAi agents AD-51544, AD-51545, or AD-45163.
[0098] FIG. 10B is a graph depicting silencing of TTR mRNA in transgenic mice that express hTTR V30M following administration of a single subcutaneous dose of RNAi agents AD-51546, AD-51547, or AD-45165.
[0099] FIG. 11 is a graph depicting TTR protein suppression in transgenic mice that express hTTR V30M following administration of a single subcutaneous dose of 5 mg/kg or 1 mg/kg of RNAi agents AD-51544, AD-51545, or AD-45163.
[0100] FIG. 12 is a graph depicting TTR protein suppression in transgenic mice that express hTTR V30M following administration of a single subcutaneous dose of 5 mg/kg or 1 mg/kg of RNAi agents AD-51546, AD-51547, or AD-45165.
[0101] FIG. 13 depicts the protocol for post-dose blood draws in monkeys that received 5.times.5 mg/kg RNAi agent (top line) or 1.times.25 mg/kg RNAi agent (bottom line).
[0102] FIG. 14A is a graph depicting suppression of TTR protein in non-human primates following subcutaneous administration of five 5 mg/kg doses of AD-45163, AD-51544, AD-51545, AD-51546, or AD-51547.
[0103] FIG. 14B is a graph depicting suppression of TTR protein in non-human primates following subcutaneous administration of a single 25 mg/kg dose of AD-45163, AD-51544, AD-51545, AD-51546, or AD-51547.
[0104] FIG. 15 is a graph depicting suppression of TTR protein in non-human primates following subcutaneous administration of AD-51547 at 2.5 mg/kg (white squares), 5 mg/kg (black squares) or 10 mg/kg (patterned squares) per dose, or administration of PBS as a negative control (gray squares).
DETAILED DESCRIPTION OF THE INVENTION
[0105] The present invention provides RNAi agents, e.g., double stranded RNAi agents, and compositions targeting the Transthyretin (TTR) gene. The present invention also provides methods of inhibiting expression of TTR and methods of treating or preventing a TTR-associated disease in a subject using the RNAi agents, e.g., double stranded RNAi agents, of the invention. The present invention is based, at least in part, on the discovery that RNAi agents that comprise particular chemical modifications show a superior ability to inhibit expression of TTR. Agents including a certain pattern of chemical modifications (e.g., an alternating pattern) and a ligand are shown herein to be effective in silencing the activity of the TTR gene. Furthermore, agents including one or more motifs of three identical modifications on three consecutive nucleotides, including one such motif at or near the cleavage site of the agents, show surprisingly enhanced TTR gene silencing activity. When a single such chemical motif is present in the agent, it is preferred to be at or near the cleavage region for enhancing of the gene silencing activity. Cleavage region is the region surrounding the cleavage site, i.e., the site on the target mRNA at which cleavage occurs.
I. Definitions
[0106] As used herein, each of the following terms has the meaning associated with it in this section.
[0107] The term "including" is used herein to mean, and is used interchangeably with, the phrase "including but not limited to".
[0108] The term "or" is used herein to mean, and is used interchangeably with, the term "and/or," unless context clearly indicates otherwise.
[0109] As used herein, a "transthyretin" ("TTR") refers to the well known gene and protein. TTR is also known as prealbumin, HsT2651, PALB, and TBPA. TTR functions as a transporter of retinol-binding protein (RBP), thyroxine (T4) and retinol, and it also acts as a protease. The liver secretes TTR into the blood, and the choroid plexus secretes TTR into the cerebrospinal fluid. TTR is also expressed in the pancreas and the retinal pigment epithelium. The greatest clinical relevance of TTR is that both normal and mutant TTR protein can form amyloid fibrils that aggregate into extracellular deposits, causing amyloidosis. See, e.g., Saraiva M. J. M. (2002) Expert Reviews in Molecular Medicine, 4(12):1-11 for a review. The molecular cloning and nucleotide sequence of rat transthyretin, as well as the distribution of mRNA expression, was described by Dickson, P. W. et al. (1985) J. Biol. Chem. 260(13)8214-8219. The X-ray crystal structure of human TTR was described in Blake, C C. et al. (1974) J Mol Biol 88, 1-12. The sequence of a human TTR mRNA transcript can be found at National Center for Biotechnology Information (NCBI) RefSeq accession number NM_000371. The sequence of mouse TTR mRNA can be found at RefSeq accession number NM_013697.2, and the sequence of rat TTR mRNA can be found at RefSeq accession number NM_012681.1 As used herein, "target sequence" refers to a contiguous portion of the nucleotide sequence of an mRNA molecule formed during the transcription of a TTR gene, including mRNA that is a product of RNA processing of a primary transcription product.
[0110] As used herein, the term "strand comprising a sequence" refers to an oligonucleotide comprising a chain of nucleotides that is described by the sequence referred to using the standard nucleotide nomenclature.
[0111] "G," "C," "A" and "U" each generally stand for a nucleotide that contains guanine, cytosine, adenine, and uracil as a base, respectively. "T" and "dT" are used interchangeably herein and refer to a deoxyribonucleotide wherein the nucleobase is thymine, e.g., deoxyribothymine, 2'-deoxythymidine or thymidine. However, it will be understood that the term "ribonucleotide" or "nucleotide" or "deoxyribonucleotide" can also refer to a modified nucleotide, as further detailed below, or a surrogate replacement moiety. The skilled person is well aware that guanine, cytosine, adenine, and uracil may be replaced by other moieties without substantially altering the base pairing properties of an oligonucleotide comprising a nucleotide bearing such replacement moiety. For example, without limitation, a nucleotide comprising inosine as its base may base pair with nucleotides containing adenine, cytosine, or uracil. Hence, nucleotides containing uracil, guanine, or adenine may be replaced in the nucleotide sequences of the invention by a nucleotide containing, for example, inosine. Sequences comprising such replacement moieties are embodiments of the invention.
[0112] A "double stranded RNAi agent," double-stranded RNA (dsRNA) molecule, also referred to as "dsRNA agent," "dsRNA", "siRNA", "iRNA agent," as used interchangeably herein, refers to a complex of ribonucleic acid molecules, having a duplex structure comprising two anti-parallel and substantially complementary, as defined below, nucleic acid strands. In general, the majority of nucleotides of each strand are ribonucleotides, but as described in detail herein, each or both strands can also include one or more non-ribonucleotides, e.g., a deoxyribonucleotide and/or a modified nucleotide. In addition, as used in this specification, an "RNAi agent" may include ribonucleotides with chemical modifications; an RNAi agent may include substantial modifications at multiple nucleotides. Such modifications may include all types of modifications disclosed herein or known in the art. Any such modifications, as used in a siRNA type molecule, are encompassed by "RNAi agent" for the purposes of this specification and claims.
[0113] In another embodiment, the RNAi agent may be a single-stranded siRNA that is introduced into a cell or organism to inhibit a target mRNA. Single-stranded RNAi agents bind to the RISC endonuclease Argonaute 2, which then cleaves the target mRNA. The single-stranded siRNAs are generally 15-30 nucleotides and are chemically modified. The design and testing of single-stranded siRNAs are described in U.S. Pat. No. 8,101,348 and in Lima et al., (2012) Cell 150: 883-894, the entire contents of each of which are hereby incorporated herein by reference. Any of the antisense nucleotide sequences described herein may be used as a single-stranded siRNA as described herein or as chemically modified by the methods described in Lima et al., (2012) Cell 150:883-894.
[0114] The two strands forming the duplex structure may be different portions of one larger RNA molecule, or they may be separate RNA molecules. Where the two strands are part of one larger molecule, and therefore are connected by an uninterrupted chain of nucleotides between the 3'-end of one strand and the 5'-end of the respective other strand forming the duplex structure, the connecting RNA chain is referred to as a "hairpin loop." Where the two strands are connected covalently by means other than an uninterrupted chain of nucleotides between the 3'-end of one strand and the 5'-end of the respective other strand forming the duplex structure, the connecting structure is referred to as a "linker." The RNA strands may have the same or a different number of nucleotides. The maximum number of base pairs is the number of nucleotides in the shortest strand of the dsRNA minus any overhangs that are present in the duplex. In addition to the duplex structure, an RNAi agent may comprise one or more nucleotide overhangs. The term "siRNA" is also used herein to refer to an RNAi agent as described above.
[0115] In another aspect, the agent is a single-stranded antisense RNA molecule. An antisense RNA molecule is complementary to a sequence within the target mRNA. Antisense RNA can inhibit translation in a stoichiometric manner by base pairing to the mRNA and physically obstructing the translation machinery, see Dias, N. et al., (2002) Mol Cancer Ther 1:347-355. The antisense RNA molecule may have about 15-30 nucleotides that are complementary to the target mRNA. For example, the antisense RNA molecule may have a sequence of at least 15, 16, 17, 18, 19, 20 or more contiguous nucleotides from one of the antisense sequences of Table 1.
[0116] As used herein, a "nucleotide overhang" refers to the unpaired nucleotide or nucleotides that protrude from the duplex structure of an RNAi agent when a 3'-end of one strand of the RNAi agent extends beyond the 5'-end of the other strand, or vice versa. "Blunt" or "blunt end" means that there are no unpaired nucleotides at that end of the double stranded RNAi agent, i.e., no nucleotide overhang. A "blunt ended" RNAi agent is a dsRNA that is double-stranded over its entire length, i.e., no nucleotide overhang at either end of the molecule. The RNAi agents of the invention include RNAi agents with nucleotide overhangs at one end (i.e., agents with one overhang and one blunt end) or with nucleotide overhangs at both ends.
[0117] The term "antisense strand" refers to the strand of a double stranded RNAi agent which includes a region that is substantially complementary to a target sequence (e.g., a human TTR mRNA). As used herein, the term "region complementary to part of an mRNA encoding transthyretin" refers to a region on the antisense strand that is substantially complementary to part of a TTR mRNA sequence. Where the region of complementarity is not fully complementary to the target sequence, the mismatches are most tolerated in the terminal regions and, if present, are generally in a terminal region or regions, e.g., within 6, 5, 4, 3, or 2 nucleotides of the 5' and/or 3' terminus.
[0118] The term "sense strand," as used herein, refers to the strand of a dsRNA that includes a region that is substantially complementary to a region of the antisense strand.
[0119] As used herein, the term "cleavage region" refers to a region that is located immediately adjacent to the cleavage site. The cleavage site is the site on the target at which cleavage occurs. In some embodiments, the cleavage region comprises three bases on either end of, and immediately adjacent to, the cleavage site. In some embodiments, the cleavage region comprises two bases on either end of, and immediately adjacent to, the cleavage site. In some embodiments, the cleavage site specifically occurs at the site bound by nucleotides 10 and 11 of the antisense strand, and the cleavage region comprises nucleotides 11, 12 and 13.
[0120] As used herein, and unless otherwise indicated, the term "complementary," when used to describe a first nucleotide sequence in relation to a second nucleotide sequence, refers to the ability of an oligonucleotide or polynucleotide comprising the first nucleotide sequence to hybridize and form a duplex structure under certain conditions with an oligonucleotide or polynucleotide comprising the second nucleotide sequence, as will be understood by the skilled person. Such conditions can, for example, be stringent conditions, where stringent conditions may include: 400 mM NaCl, 40 mM PIPES pH 6.4, 1 mM EDTA, 50.degree. C. or 70.degree. C. for 12-16 hours followed by washing. Other conditions, such as physiologically relevant conditions as may be encountered inside an organism, can apply. The skilled person will be able to determine the set of conditions most appropriate for a test of complementarity of two sequences in accordance with the ultimate application of the hybridized nucleotides.
[0121] Sequences can be "fully complementary" with respect to each when there is base-pairing of the nucleotides of the first nucleotide sequence with the nucleotides of the second nucleotide sequence over the entire length of the first and second nucleotide sequences. However, where a first sequence is referred to as "substantially complementary" with respect to a second sequence herein, the two sequences can be fully complementary, or they may form one or more, but generally not more than 4, 3 or 2 mismatched base pairs upon hybridization, while retaining the ability to hybridize under the conditions most relevant to their ultimate application. However, where two oligonucleotides are designed to form, upon hybridization, one or more single stranded overhangs, such overhangs shall not be regarded as mismatches with regard to the determination of complementarity. For example, a dsRNA comprising one oligonucleotide 21 nucleotides in length and another oligonucleotide 23 nucleotides in length, wherein the longer oligonucleotide comprises a sequence of 21 nucleotides that is fully complementary to the shorter oligonucleotide, may yet be referred to as "fully complementary" for the purposes described herein.
[0122] "Complementary" sequences, as used herein, may also include, or be formed entirely from, non-Watson-Crick base pairs and/or base pairs formed from non-natural and modified nucleotides, in as far as the above requirements with respect to their ability to hybridize are fulfilled. Such non-Watson-Crick base pairs includes, but not limited to, G:U Wobble or Hoogstein base pairing.
[0123] The terms "complementary," "fully complementary" and "substantially complementary" herein may be used with respect to the base matching between the sense strand and the antisense strand of a dsRNA, or between the antisense strand of a dsRNA and a target sequence, as will be understood from the context of their use.
[0124] As used herein, a polynucleotide that is "substantially complementary to at least part of" a messenger RNA (mRNA) refers to a polynucleotide that is substantially complementary to a contiguous portion of the mRNA of interest (e.g., an mRNA encoding TTR) including a 5' UTR, an open reading frame (ORF), or a 3' UTR. For example, a polynucleotide is complementary to at least a part of a TTR mRNA if the sequence is substantially complementary to a non-interrupted portion of an mRNA encoding TTR.
[0125] The term "inhibiting," as used herein, is used interchangeably with "reducing," "silencing," "downregulating," "suppressing" and other similar terms, and includes any level of inhibition.
[0126] The phrase "inhibiting expression of a TTR," as used herein, includes inhibition of expression of any TTR gene (such as, e.g., a mouse TTR gene, a rat TTR gene, a monkey TTR gene, or a human TTR gene) as well as variants or mutants of a TTR gene. Thus, the TTR gene may be a wild-type TTR gene, a mutant TTR gene (such as a mutant TTR gene giving rise to systemic amyloid deposition), or a transgenic TTR gene in the context of a genetically manipulated cell, group of cells, or organism.
[0127] "Inhibiting expression of a TTR gene" includes any level of inhibition of a TTR gene, e.g., at least partial suppression of the expression of a TTR gene, such as an inhibition of at least about 5%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 91%, at least about 92%, at least about 93%, at least about 94%. at least about 95%, at least about 96%, at least about 97%, at least about 98%, or at least about 99%.
[0128] The expression of a TTR gene may be assessed based on the level of any variable associated with TTR gene expression, e.g., TTR mRNA level, TTR protein level, retinol binding protein level, vitamin A level, or the number or extent of amyloid deposits. Inhibition may be assessed by a decrease in an absolute or relative level of one or more of these variables compared with a control level. The control level may be any type of control level that is utilized in the art, e.g., a pre-dose baseline level, or a level determined from a similar subject, cell, or sample that is untreated or treated with a control (such as, e.g., buffer only control or inactive agent control).
[0129] The phrase "contacting a cell with an RNAi agent," as used herein, includes contacting a cell by any possible means. Contacting a cell with an RNAi agent, e.g., a double stranded RNAi agent, includes contacting a cell in vitro with the RNAi agent or contacting a cell in vivo with the RNAi agent. The contacting may be done directly or indirectly. Thus, for example, the RNAi agent may be put into physical contact with the cell by the individual performing the method, or alternatively, the RNAi agent may be put into a situation that will permit or cause it to subsequently come into contact with the cell.
[0130] Contacting a cell in vitro may be done, for example, by incubating the cell with the RNAi agent. Contacting a cell in vivo may be done, for example, by injecting the RNAi agent into or near the tissue where the cell is located, or by injecting the RNAi agent into another area, e.g., the bloodstream or the subcutaneous space, such that the agent will subsequently reach the tissue where the cell to be contacted is located. For example, the RNAi agent may contain and/or be coupled to a ligand, e.g., a GalNAc.sub.3 ligand, that directs the RNAi agent to a site of interest, e.g., the liver. Combinations of in vitro and in vivo methods of contacting are also possible. In connection with the methods of the invention, a cell might also be contacted in vitro with an RNAi agent and subsequently transplanted into a subject.
[0131] A "patient" or "subject," as used herein, is intended to include either a human or non-human animal, preferably a mammal, e.g., a monkey. Most preferably, the subject or patient is a human.
[0132] A "TTR-associated disease," as used herein, is intended to include any disease associated with the TTR gene or protein. Such a disease may be caused, for example, by excess production of the TTR protein, by TTR gene mutations, by abnormal cleavage of the TTR protein, by abnormal interactions between TTR and other proteins or other endogenous or exogenous substances. A "TTR-associated disease" includes any type of TTR amyloidosis (ATTR) wherein TTR plays a role in the formation of abnormal extracellular aggregates or amyloid deposits. TTR-associated diseases include senile systemic amyloidosis (SSA), systemic familial amyloidosis, familial amyloidotic polyneuropathy (FAP), familial amyloidotic cardiomyopathy (FAC), leptomeningeal/Central Nervous System (CNS) amyloidosis, amyloidotic vitreous opacities, carpal tunnel syndrome, and hyperthyroxinemia. Symptoms of TTR amyloidosis include sensory neuropathy (e.g., paresthesia, hypesthesia in distal limbs), autonomic neuropathy (e.g., gastrointestinal dysfunction, such as gastric ulcer, or orthostatic hypotension), motor neuropathy, seizures, dementia, myelopathy, polyneuropathy, carpal tunnel syndrome, autonomic insufficiency, cardiomyopathy, vitreous opacities, renal insufficiency, nephropathy, substantially reduced mBMI (modified Body Mass Index), cranial nerve dysfunction, and corneal lattice dystrophy.
[0133] "Therapeutically effective amount," as used herein, is intended to include the amount of an RNAi agent that, when administered to a patient for treating a TTR associated disease, is sufficient to effect treatment of the disease (e.g., by diminishing, ameliorating or maintaining the existing disease or one or more symptoms of disease). The "therapeutically effective amount" may vary depending on the RNAi agent, how the agent is administered, the disease and its severity and the history, age, weight, family history, genetic makeup, stage of pathological processes mediated by TTR expression, the types of preceding or concomitant treatments, if any, and other individual characteristics of the patient to be treated.
[0134] "Prophylactically effective amount," as used herein, is intended to include the amount of an RNAi agent that, when administered to a subject who does not yet experience or display symptoms of a TTR-associated disease, but who may be predisposed to the disease, is sufficient to prevent or ameliorate the disease or one or more symptoms of the disease. Symptoms that may be ameliorated include sensory neuropathy (e.g., paresthesia, hypesthesia in distal limbs), autonomic neuropathy (e.g., gastrointestinal dysfunction, such as gastric ulcer, or orthostatic hypotension), motor neuropathy, seizures, dementia, myelopathy, polyneuropathy, carpal tunnel syndrome, autonomic insufficiency, cardiomyopathy, vitreous opacities, renal insufficiency, nephropathy, substantially reduced mBMI (modified Body Mass Index), cranial nerve dysfunction, and corneal lattice dystrophy. Ameliorating the disease includes slowing the course of the disease or reducing the severity of later-developing disease. The "prophylactically effective amount" may vary depending on the RNAi agent, how the agent is administered, the degree of risk of disease, and the history, age, weight, family history, genetic makeup, the types of preceding or concomitant treatments, if any, and other individual characteristics of the patient to be treated.
[0135] A "therapeutically-effective amount" or "prophylacticaly effective amount" also includes an amount of an RNAi agent that produces some desired local or systemic effect at a reasonable benefit/risk ratio applicable to any treatment. RNAi gents employed in the methods of the present invention may be administered in a sufficient amount to produce a reasonable benefit/risk ratio applicable to such treatment.
[0136] The term "sample," as used herein, includes a collection of similar fluids, cells, or tissues isolated from a subject, as well as fluids, cells, or tissues present within a subject. Examples of biological fluids include blood, serum and serosal fluids, plasma, cerebrospinal fluid, ocular fluids, lymph, urine, saliva, and the like. Tissue samples may include samples from tissues, organs or localized regions. For example, samples may be derived from particular organs, parts of organs, or fluids or cells within those organs. In certain embodiments, samples may be derived from the liver (e.g., whole liver or certain segments of liver or certain types of cells in the liver, such as, e.g., hepatocytes), the retina or parts of the retina (e.g., retinal pigment epithelium), the central nervous system or parts of the central nervous system (e.g., ventricles or choroid plexus), or the pancreas or certain cells or parts of the pancreas. In some embodiments, a "sample derived from a subject" refers tocerebrospinal fluid obtained from the subject. In preferred embodiments, a "sample derived from a subject" refers to blood or plasma drawn from the subject. In further embodiments, a "sample derived from a subject" refers to liver tissue (or subcomponents thereof) or retinal tissue (or subcomponents thereof) derived from the subject.
II. RNAi Agents
[0137] The present invention provides RNAi agents with superior gene silencing activity. It is shown herein and in Provisional Application No. 61/561,710 (to which the present application claims priority) that a superior result may be obtained by introducing one or more motifs of three identical modifications on three consecutive nucleotides into a sense strand and/or antisense strand of a RNAi agent, particularly at or near the cleavage site. The sense strand and antisense strand of the RNAi agent may otherwise be completely modified. The introduction of these motifs interrupts the modification pattern, if present, of the sense and/or antisense strand. The RNAi agent also optionally conjugates with a GalNAc derivative ligand, for instance on the sense strand. The resulting RNAi agents present superior gene silencing activity.
[0138] The inventors surprisingly discovered that when the sense strand and antisense strand of the RNAi agent are completely modified, having one or more motifs of three identical modifications on three consecutive nucleotides at or near the cleavage site of at least one strand of a RNAi agent superiorly enhanced the gene silencing activity of the RNAi agent.
[0139] Accordingly, the invention provides RNAi agents, e.g., double stranded RNAi agents, capable of inhibiting the expression of a target gene (i.e., a TTR gene) in vivo. The RNAi agent comprises a sense strand and an antisense strand. Each strand of the RNAi agent can range from 12-30 nucleotides in length. For example, each strand can be between 14-30 nucleotides in length, 17-30 nucleotides in length, 25-30 nucleotides in length, 27-30 nucleotides in length, 17-23 nucleotides in length, 17-21 nucleotides in length, 17-19 nucleotides in length, 19-25 nucleotides in length, 19-23 nucleotides in length, 19-21 nucleotides in length, 21-25 nucleotides in length, or 21-23 nucleotides in length.
[0140] The sense strand and antisense strand typically form a duplex double stranded RNA ("dsRNA"), also referred to herein as an "RNAi agent." The duplex region of an RNAi agent may be 12-30 nucleotide pairs in length. For example, the duplex region can be between 14-30 nucleotide pairs in length, 17-30 nucleotide pairs in length, 27-30 nucleotide pairs in length, 17-23 nucleotide pairs in length, 17-21 nucleotide pairs in length, 17-19 nucleotide pairs in length, 19-25 nucleotide pairs in length, 19-23 nucleotide pairs in length, 19-21 nucleotide pairs in length, 21-25 nucleotide pairs in length, or 21-23 nucleotide pairs in length. In another example, the duplex region is selected from 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, and 27.
[0141] In one embodiment, the RNAi agent may contain one or more overhang regions and/or capping groups of RNAi agent at 3'-end, or 5'-end or both ends of a strand. The overhang can be 1-6 nucleotides in length, for instance 2-6 nucleotides in length, 1-5 nucleotides in length, 2-5 nucleotides in length, 1-4 nucleotides in length, 2-4 nucleotides in length, 1-3 nucleotides in length, 2-3 nucleotides in length, or 1-2 nucleotides in length. The overhangs can be the result of one strand being longer than the other, or the result of two strands of the same length being staggered. The overhang can form a mismatch with the target mRNA or it can be complementary to the gene sequences being targeted or can be other sequence. The first and second strands can also be joined, e.g., by additional bases to form a hairpin, or by other non-base linkers.
[0142] The RNAi agents provided by the present invention include agents with chemical modifications as disclosed, for example, in U.S. Provisional Application No. 61/561,710, filed on Nov. 18, 2011, International Application No. PCT/US2011/051597, filed on Sep. 15, 2010, and PCT Publication WO 2009/073809, the entire contents of each of which are incorporated herein by reference.
[0143] In one embodiment, the nucleotides in the overhang region of the RNAi agent can each independently be a modified or unmodified nucleotide including, but no limited to 2'-sugar modified, such as, 2-F, 2'-O-methyl, thymidine (T), 2'-O-methoxyethyl-5-methyluridine (Teo), 2'-O-methoxyethyladenosine (Aeo), 2'-O-methoxyethyl-5-methylcytidine (m5Ceo), and any combinations thereof. For example, TT can be an overhang sequence for either end on either strand. The overhang can form a mismatch with the target mRNA or it can be complementary to the gene sequences being targeted or can be other sequence.
[0144] The 5'- or 3'-overhangs at the sense strand, antisense strand or both strands of the RNAi agent may be phosphorylated. In some embodiments, the overhang region contains two nucleotides having a phosphorothioate between the two nucleotides, where the two nucleotides can be the same or different. In one embodiment, the overhang is present at the 3'-end of the sense strand, antisense strand or both strands. In one embodiment, this 3'-overhang is present in the antisense strand. In one embodiment, this 3'-overhang is present in the sense strand.
[0145] The RNAi agent may contain only a single overhang, which can strengthen the interference activity of the RNAi, without affecting its overall stability. For example, the single-stranded overhang is located at the 3-terminal end of the sense strand or, alternatively, at the 3-terminal end of the antisense strand. The RNAi may also have a blunt end, located at the 5'-end of the antisense strand (or the 3'-end of the sense strand) or vice versa. Generally, the antisense strand of the RNAi has a nucleotide overhang at the 3'-end, and the 5'-end is blunt. While the Applicants are not bound by theory, the theoretical mechanism is that the asymmetric blunt end at the 5'-end of the antisense strand and 3'-end overhang of the antisense strand favor the guide strand loading into RISC process.
[0146] In one embodiment, the RNAi agent is a double ended bluntmer of 19 nt in length, wherein the sense strand contains at least one motif of three 2'-F modifications on three consecutive nucleotides at positions 7, 8, 9 from the 5' end. The antisense strand contains at least one motif of three 2'-O-methyl modifications on three consecutive nucleotides at positions 11, 12, 13 from the 5' end.
[0147] In one embodiment, the RNAi agent is a double ended bluntmer of 20 nt in length, wherein the sense strand contains at least one motif of three 2'-F modifications on three consecutive nucleotides at positions 8, 9, 10 from the 5' end. The antisense strand contains at least one motif of three 2'-O-methyl modifications on three consecutive nucleotides at positions 11, 12, 13 from the 5' end.
[0148] In one embodiment, the RNAi agent is a double ended bluntmer of 21 nt in length, wherein the sense strand contains at least one motif of three 2'-F modifications on three consecutive nucleotides at positions 9, 10, 11 from the 5' end. The antisense strand contains at least one motif of three 2'-O-methyl modifications on three consecutive nucleotides at positions 11, 12, 13 from the 5' end.
[0149] In one embodiment, the RNAi agent comprises a 21 nucleotides (nt) sense strand and a 23 nucleotides (nt) antisense strand, wherein the sense strand contains at least one motif of three 2'-F modifications on three consecutive nucleotides at positions 9, 10, 11 from the 5' end; the antisense strand contains at least one motif of three 2'-O-methyl modifications on three consecutive nucleotides at positions 11, 12, 13 from the 5' end, wherein one end of the RNAi agent is blunt, while the other end comprises a 2 nt overhang. Preferably, the 2 nt overhang is at the 3'-end of the antisense. Optionally, the RNAi agent further comprises a ligand (preferably GalNAc.sub.3).
[0150] In one embodiment, the RNAi agent comprises a sense and an antisense strand, wherein the sense strand is 25-30 nucleotide residues in length, wherein starting from the 5' terminal nucleotide (position 1) positions 1 to 23 of the first strand comprise at least 8 ribonucleotides; antisense strand is 36-66 nucleotide residues in length and, starting from the 3' terminal nucleotide, comprises at least 8 ribonucleotides in the positions paired with positions 1-23 of sense strand to form a duplex; wherein at least the 3' terminal nucleotide of antisense strand is unpaired with sense strand, and up to 6 consecutive 3' terminal nucleotides are unpaired with sense strand, thereby forming a 3' single stranded overhang of 1-6 nucleotides; wherein the 5' terminus of antisense strand comprises from 10-30 consecutive nucleotides which are unpaired with sense strand, thereby forming a 10-30 nucleotide single stranded 5' overhang; wherein at least the sense strand 5' terminal and 3' terminal nucleotides are base paired with nucleotides of antisense strand when sense and antisense strands are aligned for maximum complementarity, thereby forming a substantially duplexed region between sense and antisense strands; and antisense strand is sufficiently complementary to a target RNA along at least 19 ribonucleotides of antisense strand length to reduce target gene expression when the double stranded nucleic acid is introduced into a mammalian cell; and wherein the sense strand contains at least one motif of three 2'-F modifications on three consecutive nucleotides, where at least one of the motifs occurs at or near the cleavage site. The antisense strand contains at least one motif of three 2'-O-methyl modifications on three consecutive nucleotides at or near the cleavage site.
[0151] In one embodiment, the RNAi agent comprises sense and antisense strands, wherein the RNAi agent comprises a first strand having a length which is at least 25 and at most 29 nucleotides and a second strand having a length which is at most 30 nucleotides with at least one motif of three 2'-O-methyl modifications on three consecutive nucleotides at position 11, 12, 13 from the 5' end; wherein the 3' end of the first strand and the 5' end of the second strand form a blunt end and the second strand is 1-4 nucleotides longer at its 3' end than the first strand, wherein the duplex region which is at least 25 nucleotides in length, and the second strand is sufficiently complementary to a target mRNA along at least 19 nt of the second strand length to reduce target gene expression when the RNAi agent is introduced into a mammalian cell, and wherein dicer cleavage of the RNAi agent preferentially results in an siRNA comprising the 3' end of the second strand, thereby reducing expression of the target gene in the mammal. Optionally, the RNAi agent further comprises a ligand.
[0152] In one embodiment, the sense strand of the RNAi agent contains at least one motif of three identical modifications on three consecutive nucleotides, where one of the motifs occurs at the cleavage site in the sense strand.
[0153] In one embodiment, the antisense strand of the RNAi agent can also contain at least one motif of three identical modifications on three consecutive nucleotides, where one of the motifs occurs at or near the cleavage site in the antisense strand
[0154] For RNAi agent having a duplex region of 17-23 nt in length, the cleavage site of the antisense strand is typically around the 10, 11 and 12 positions from the 5'-end. Thus, the motifs of three identical modifications may occur at the 9, 10, 11 positions; 10, 11, 12 positions; 11, 12, 13 positions; 12, 13, 14 positions; or 13, 14, 15 positions of the antisense strand, the count starting from the 1.sup.st nucleotide from the 5'-end of the antisense strand, or, the count starting from the 1.sup.st paired nucleotide within the duplex region from the 5'-end of the antisense strand. The cleavage site in the antisense strand may also change according to the length of the duplex region of the RNAi from the 5'-end.
[0155] The sense strand of the RNAi agent may contain at least one motif of three identical modifications on three consecutive nucleotides at the cleavage site of the strand; and the antisense strand may have at least one motif of three identical modifications on three consecutive nucleotides at or near the cleavage site of the strand. When the sense strand and the antisense strand form a dsRNA duplex, the sense strand and the antisense strand can be so aligned that one motif of the three nucleotides on the sense strand and one motif of the three nucleotides on the antisense strand have at least one nucleotide overlap, i.e., at least one of the three nucleotides of the motif in the sense strand forms a base pair with at least one of the three nucleotides of the motif in the antisense strand. Alternatively, at least two nucleotides may overlap, or all three nucleotides may overlap.
[0156] In one embodiment, the sense strand of the RNAi agent may contain more than one motif of three identical modifications on three consecutive nucleotides. The first motif should occur at or near the cleavage site of the strand and the other motifs may be wing modifications. The term "wing modification" herein refers to a motif occurring at another portion of the strand that is separated from the motif at or near the cleavage site of the same strand. The wing modification is either adjacent to the first motif or is separated by at least one or more nucleotides. When the motifs are immediately adjacent to each other than the chemistry of the motifs are distinct from each other and when the motifs are separated by one or more nucleotide than the chemistries can be the same or different. Two or more wing modifications may be present. For instance, when two wing modifications are present, each wing modification may occur at one end relative to the first motif which is at or near cleavage site or on either side of the lead motif.
[0157] Like the sense strand, the antisense strand of the RNAi agent may contain at least two motifs of three identical modifications on three consecutive nucleotides, with at least one of the motifs occurring at or near the cleavage site of the strand. This antisense strand may also contain one or more wing modifications in an alignment similar to the wing modifications that is present on the sense strand.
[0158] In one embodiment, the wing modification on the sense strand or antisense strand of the RNAi agent typically does not include the first one or two terminal nucleotides at the 3'-end, 5'-end or both ends of the strand.
[0159] In another embodiment, the wing modification on the sense strand or antisense strand of the RNAi agent typically does not include the first one or two paired nucleotides within the duplex region at the 3'-end, 5'-end or both ends of the strand.
[0160] When the sense strand and the antisense strand of the RNAi agent each contain at least one wing modification, the wing modifications may fall on the same end of the duplex region, and have an overlap of one, two or three nucleotides.
[0161] When the sense strand and the antisense strand of the RNAi agent each contain at least two wing modifications, the sense strand and the antisense strand can be so aligned that two modifications each from one strand fall on one end of the duplex region, having an overlap of one, two or three nucleotides; two modifications each from one strand fall on the other end of the duplex region, having an overlap of one, two or three nucleotides; two modifications one strand fall on each side of the lead motif, having an overlap of one, two or three nucleotides in the duplex region.
[0162] In one embodiment, every nucleotide in the sense strand and antisense strand of the RNAi agent, including the nucleotides that are part of the motifs, may be modified. Each nucleotide may be modified with the same or different modification which can include one or more alteration of one or both of the non-linking phosphate oxygens and/or of one or more of the linking phosphate oxygens; alteration of a constituent of the ribose sugar, e.g., of the 2' hydroxyl on the ribose sugar; wholesale replacement of the phosphate moiety with "dephospho" linkers; modification or replacement of a naturally occurring base; and replacement or modification of the ribose-phosphate backbone.
[0163] As nucleic acids are polymers of subunits, many of the modifications occur at a position which is repeated within a nucleic acid, e.g., a modification of a base, or a phosphate moiety, or a non-linking O of a phosphate moiety. In some cases the modification will occur at all of the subject positions in the nucleic acid but in many cases it will not. By way of example, a modification may only occur at a 3' or 5' terminal position, may only occur in a terminal region, e.g., at a position on a terminal nucleotide or in the last 2, 3, 4, 5, or 10 nucleotides of a strand. A modification may occur in a double strand region, a single strand region, or in both. A modification may occur only in the double strand region of a RNA or may only occur in a single strand region of a RNA. For example, a phosphorothioate modification at a non-linking O position may only occur at one or both termini, may only occur in a terminal region, e.g., at a position on a terminal nucleotide or in the last 2, 3, 4, 5, or 10 nucleotides of a strand, or may occur in double strand and single strand regions, particularly at termini. The 5' end or ends can be phosphorylated.
[0164] It may be possible, e.g., to enhance stability, to include particular bases in overhangs, or to include modified nucleotides or nucleotide surrogates, in single strand overhangs, e.g., in a 5' or 3' overhang, or in both. For example, it can be desirable to include purine nucleotides in overhangs. In some embodiments all or some of the bases in a 3' or 5' overhang may be modified, e.g., with a modification described herein. Modifications can include, e.g., the use of modifications at the 2' position of the ribose sugar with modifications that are known in the art, e.g., the use of deoxyribonucleotides, 2'-deoxy-2'-fluoro (2'-F) or 2'-O-methyl modified instead of the ribosugar of the nucleobase, and modifications in the phosphate group, e.g., phosphorothioate modifications. Overhangs need not be homologous with the target sequence.
[0165] In one embodiment, each residue of the sense strand and antisense strand is independently modified with LNA, HNA, CeNA, 2'-methoxyethyl, 2'-O-methyl, 2'-O-allyl, 2'-C-allyl, 2'-deoxy, 2'-hydroxyl, or 2'-fluoro. The strands can contain more than one modification. In one embodiment, each residue of the sense strand and antisense strand is independently modified with 2'-O-methyl or 2'-fluoro.
[0166] At least two different modifications are typically present on the sense strand and antisense strand. Those two modifications may be the 2'-O-methyl or 2'-fluoro modifications, or others.
[0167] In one embodiment, the N.sub.a and/or N.sub.b comprise modifications of an alternating pattern. The term "alternating motif" as used herein refers to a motif having one or more modifications, each modification occurring on alternating nucleotides of one strand. The alternating nucleotide may refer to one per every other nucleotide or one per every three nucleotides, or a similar pattern. For example, if A, B and C each represent one type of modification to the nucleotide, the alternating motif can be "ABABABABABAB . . . ," "AABBAABBAABB . . . ," "AABAABAABAAB . . . ," "AAABAAABAAAB . . . ," "AAABBBAAABBB . . . ," or "ABCABCABCABC . . . ," etc.
[0168] The type of modifications contained in the alternating motif may be the same or different. For example, if A, B, C, D each represent one type of modification on the nucleotide, the alternating pattern, i.e., modifications on every other nucleotide, may be the same, but each of the sense strand or antisense strand can be selected from several possibilities of modifications within the alternating motif such as "ABABAB . . . " "ACACAC . . . " "BDBDBD . . . " or "CDCDCD . . . ," etc.
[0169] In one embodiment, the RNAi agent of the invention comprises the modification pattern for the alternating motif on the sense strand relative to the modification pattern for the alternating motif on the antisense strand is shifted. The shift may be such that the modified group of nucleotides of the sense strand corresponds to a differently modified group of nucleotides of the antisense strand and vice versa. For example, the sense strand when paired with the antisense strand in the dsRNA duplex, the alternating motif in the sense strand may start with "ABABAB" from 5'-3' of the strand and the alternating motif in the antisense strand may start with "BABABA" from 5'-3' of the strand within the duplex region. As another example, the alternating motif in the sense strand may start with "AABBAABB" from 5'-3' of the strand and the alternating motif in the antisenese strand may start with "BBAABBAA" from 5'-3' of the strand within the duplex region, so that there is a complete or partial shift of the modification patterns between the sense strand and the antisense strand.
[0170] In one embodiment, the RNAi agent comprises the pattern of the alternating motif of 2'-O-methyl modification and 2'-F modification on the sense strand initially has a shift relative to the pattern of the alternating motif of 2'-O-methyl modification and 2'-F modification on the antisense strand initially, i.e., the 2'-O-methyl modified nucleotide on the sense strand base pairs with a 2'-F modified nucleotide on the antisense strand and vice versa. The 1 position of the sense strand may start with the 2'-F modification, and the 1 position of the antisense strand may start with the 2'-O-methyl modification.
[0171] The introduction of one or more motifs of three identical modifications on three consecutive nucleotides to the sense strand and/or antisense strand interrupts the initial modification pattern present in the sense strand and/or antisense strand. This interruption of the modification pattern of the sense and/or antisense strand by introducing one or more motifs of three identical modifications on three consecutive nucleotides to the sense and/or antisense strand surprisingly enhances the gene silencing activity to the target gene.
[0172] In one embodiment, when the motif of three identical modifications on three consecutive nucleotides is introduced to any of the strands, the modification of the nucleotide next to the motif is a different modification than the modification of the motif. For example, the portion of the sequence containing the motif is " . . . N.sub.aYYYN.sub.b . . . ," where "Y" represents the modification of the motif of three identical modifications on three consecutive nucleotide, and "N.sub.a" and "N.sub.b" represent a modification to the nucleotide next to the motif "YYY" that is different than the modification of Y, and where N.sub.a and N.sub.b can be the same or different modifications. Alternatively, N.sub.a and/or N.sub.b may be present or absent when there is a wing modification present.
[0173] The RNAi agent may further comprise at least one phosphorothioate or methylphosphonate internucleotide linkage. The phosphorothioate or methylphosphonate internucleotide linkage modification may occur on any nucleotide of the sense strand or antisense strand or both in any position of the strand. For instance, the internucleotide linkage modification may occur on every nucleotide on the sense strand or antisense strand; each internucleotide linkage modification may occur in an alternating pattern on the sense strand or antisense strand; or the sense strand or antisense strand may contain both internucleotide linkage modifications in an alternating pattern. The alternating pattern of the internucleotide linkage modification on the sense strand may be the same or different from the antisense strand, and the alternating pattern of the internucleotide linkage modification on the sense strand may have a shift relative to the alternating pattern of the internucleotide linkage modification on the antisense strand.
[0174] In one embodiment, the RNAi comprises the phosphorothioate or methylphosphonate internucleotide linkage modification in the overhang region. For example, the overhang region may contain two nucleotides having a phosphorothioate or methylphosphonate internucleotide linkage between the two nucleotides. Internucleotide linkage modifications also may be made to link the overhang nucleotides with the terminal paired nucleotides within duplex region. For example, at least 2, 3, 4, or all the overhang nucleotides may be linked through phosphorothioate or methylphosphonate internucleotide linkage, and optionally, there may be additional phosphorothioate or methylphosphonate internucleotide linkages linking the overhang nucleotide with a paired nucleotide that is next to the overhang nucleotide. For instance, there may be at least two phosphorothioate internucleotide linkages between the terminal three nucleotides, in which two of the three nucleotides are overhang nucleotides, and the third is a paired nucleotide next to the overhang nucleotide. Preferably, these terminal three nucleotides may be at the 3'-end of the antisense strand.
[0175] In one embodiment, the RNAi agent comprises mismatch(es) with the target, within the duplex, or combinations thereof. The mistmatch can occur in the overhang region or the duplex region. The base pair can be ranked on the basis of their propensity to promote dissociation or melting (e.g., on the free energy of association or dissociation of a particular pairing, the simplest approach is to examine the pairs on an individual pair basis, though next neighbor or similar analysis can also be used). In terms of promoting dissociation: A:U is preferred over G:C; G:U is preferred over G:C; and I:C is preferred over G:C (I=inosine). Mismatches, e.g., non-canonical or other than canonical pairings (as described elsewhere herein) are preferred over canonical (A:T, A:U, G:C) pairings; and pairings which include a universal base are preferred over canonical pairings.
[0176] In one embodiment, the RNAi agent comprises at least one of the first 1, 2, 3, 4, or 5 base pairs within the duplex regions from the 5'-end of the antisense strand can be chosen independently from the group of: A:U, G:U, I:C, and mismatched pairs, e.g., non-canonical or other than canonical pairings or pairings which include a universal base, to promote the dissociation of the antisense strand at the 5'-end of the duplex.
[0177] In one embodiment, the nucleotide at the 1 position within the duplex region from the 5'-end in the antisense strand is selected from the group consisting of A, dA, dU, U, and dT. Alternatively, at least one of the first 1, 2 or 3 base pair within the duplex region from the 5'-end of the antisense strand is an AU base pair. For example, the first base pair within the duplex region from the 5'-end of the antisense strand is an AU base pair.
[0178] In one embodiment, the sense strand sequence may be represented by formula (I):
5'n.sub.p-N.sub.a-(XXX).sub.i-N.sub.b-YYY-N.sub.b-(ZZZ)j-N.sub.a-n.sub.q- 3' (I)
[0179] wherein:
[0180] i and j are each independently 0 or 1;
[0181] p and q are each independently 0-6;
[0182] each N.sub.a independently represents an oligonucleotide sequence comprising 0-25 modified nucleotides, each sequence comprising at least two differently modified nucleotides;
[0183] each N.sub.b independently represents an oligonucleotide sequence comprising 0-10 modified nucleotides;
[0184] each n.sub.p and n.sub.q independently represent an overhang nucleotide;
[0185] wherein N.sub.b and Y do not have the same modification; and
[0186] XXX, YYY and ZZZ each independently represent one motif of three identical modifications on three consecutive nucleotides. Preferably YYY is all 2'-F modified nucleotides.
[0187] In one embodiment, the N.sub.a and/or N.sub.b comprise modifications of alternating pattern.
[0188] In one embodiment, the YYY motif occurs at or near the cleavage site of the sense strand. For example, when the RNAi agent has a duplex region of 17-23 nucleotides in length, the YYY motif can occur at or the vicinity of the cleavage site (e.g.: can occur at positions 6, 7, 8, 7, 8, 9, 8, 9, 10, 9, 10, 11, 10, 11, 12 or 11, 12, 13) of--the sense strand, the count starting from the 1.sup.st nucleotide, from the 5'-end; or optionally, the count starting at the 1.sup.st paired nucleotide within the duplex region, from the 5'-end.
[0189] In one embodiment, i is 1 and j is 0, or i is 0 and j is 1, or both i and j are 1. The sense strand can therefore be represented by the following formulas:
5'n.sub.p-N.sub.a--YYY-N.sub.b-ZZZ-N.sub.a-n.sub.q3' (Ia);
5'n.sub.p-N.sub.a--XXX-N.sub.b-YYY-N.sub.a-n.sub.q3' (Ib); or
5'n.sub.p-N.sub.a--XXX-N.sub.b-YYY-N.sub.b-ZZZ-N.sub.a-n.sub.q3' (Ic).
[0190] When the sense strand is represented by formula (Ia), N.sub.b represents an oligonucleotide sequence comprising 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a independently can represent an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0191] When the sense strand is represented as formula (Ib), N.sub.b represents an oligonucleotide sequence comprising 0-10, 0-7, 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a can independently represent an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0192] When the sense strand is represented as formula (Ic), each N.sub.b independently represents an oligonucleotide sequence comprising 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Preferably, N.sub.b is 0, 1, 2, 3, 4, 5 or 6 Each N.sub.a can independently represent an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0193] Each of X, Y and Z may be the same or different from each other.
[0194] In one embodiment, the antisense strand sequence of the RNAi may be represented by formula (II):
5'n.sub.q'-N.sub.a'-(Z'Z'Z').sub.k-N.sub.b'-Y'Y'Y'-N.sub.b'-(X'X'X').sub- .i-N'.sub.a-n.sub.p'3' (II)
[0195] wherein:
[0196] k and l are each independently 0 or 1;
[0197] p' and q' are each independently 0-6;
[0198] each N.sub.a' independently represents an oligonucleotide sequence comprising 0-25 modified nucleotides, each sequence comprising at least two differently modified nucleotides;
[0199] each N.sub.b' independently represents an oligonucleotide sequence comprising 0-10 modified nucleotides;
[0200] each n.sub.p' and n.sub.q' independently represent an overhang nucleotide;
[0201] wherein N.sub.b' and Y' do not have the same modification;
[0202] and
[0203] X'X'X', Y'Y'Y' and Z'Z'Z' each independently represent one motif of three identical modifications on three consecutive nucleotides.
[0204] In one embodiment, the N.sub.a' and/or N.sub.b' comprise modifications of alternating pattern.
[0205] The Y'Y'Y' motif occurs at or near the cleavage site of the antisense strand. For example, when the RNAi agent has a duplex region of 17-23 nt in length, the Y'Y'Y' motif can occur at positions 9, 10, 11; 10, 11, 12; 11, 12, 13; 12, 13, 14; or 13, 14, 15 of the antisense strand, with the count starting from the 1.sup.st nucleotide, from the 5'-end; or optionally, the count starting at the 1.sup.st paired nucleotide within the duplex region, from the 5'-end. Preferably, the Y'Y'Y' motif occurs at positions 11, 12, 13.
[0206] In one embodiment, Y'Y'Y' motif is all 2'-OMe modified nucleotides.
[0207] In one embodiment, k is 1 and l is 0, or k is 0 and l is 1, or both k and l are 1.
[0208] The antisense strand can therefore be represented by the following formulas:
5'n.sub.q'-N.sub.a'-Z'Z'Z'-N.sub.b'-Y'Y'Y'-N.sub.a'-n.sub.p'3' (IIa);
5'n.sub.q'-N.sub.a'-Y'Y'Y'-N.sub.b'-X'X'X'-n.sub.p'3' (Ib); or
5'n.sub.q'-N.sub.a'-Z'Z'Z'-N.sub.b'-Y'Y'Y'-N.sub.b'-X'X'X'-N.sub.a'-n.su- b.p'3' (IIc).
[0209] When the antisense strand is represented by formula (IIa), N.sub.b' represents an oligonucleotide sequence comprising 0-10, 0-7, 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a' independently represents an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0210] When the antisense strand is represented as formula (IIb), N.sub.b' represents an oligonucleotide sequence comprising 0-10, 0-7, 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a' independently represents an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0211] When the antisense strand is represented as formula (IIc), each N.sub.b' independently represents an oligonucleotide sequence comprising 0-10, 0-7, 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a' independently represents an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides. Preferably, N.sub.b is 0, 1, 2, 3, 4, 5 or 6.
[0212] Each of X', Y' and Z' may be the same or different from each other.
[0213] Each nucleotide of the sense strand and antisense strand may be independently modified with LNA, HNA, CeNA, 2'-methoxyethyl, 2'-O-methyl, 2'-O-allyl, 2'-C-allyl, 2'-hydroxyl, 2'-deoxy or 2'-fluoro. For example, each nucleotide of the sense strand and antisense strand is independently modified with 2'-O-methyl or 2'-fluoro. Each X, Y, Z, X', Y' and Z', in particular, may represent a 2'-O-methyl modification or a 2'-fluoro modification.
[0214] In one embodiment, the sense strand of the RNAi agent may contain YYY motif occurring at 9, 10 and 11 positions of the strand when the duplex region is 21 nt, the count starting from the 1.sup.st nucleotide from the 5'-end, or optionally, the count starting at the 1.sup.st paired nucleotide within the duplex region, from the 5'-end; and Y represents 2'-F modification. The sense strand may additionally contain XXX motif or ZZZ motifs as wing modifications at the opposite end of the duplex region; and XXX and ZZZ each independently represents a 2'-OMe modification or 2'-F modification.
[0215] In one embodiment the antisense strand may contain Y'Y'Y' motif occurring at positions 11, 12, 13 of the strand, the count starting from the 1.sup.st nucleotide from the 5'-end, or optionally, the count starting at the 1.sup.st paired nucleotide within the duplex region, from the 5'-end; and Y' represents 2'-O-methyl modification. The antisense strand may additionally contain X'X'X' motif or Z'Z'Z' motifs as wing modifications at the opposite end of the duplex region; and X'X'X' and Z'Z'Z' each independently represents a 2'-OMe modification or 2'-F modification.
[0216] The sense strand represented by any one of the above formulas (Ia), (Ib) and (Ic) forms a duplex with a antisense strand being represented by any one of formulas (IIa), (IIb) and (IIc), respectively.
[0217] Accordingly, the RNAi agents of the invention may comprise a sense strand and an antisense strand, each strand having 14 to 30 nucleotides, the RNAi duplex represented by formula (III):
sense: 5'n.sub.p-N.sub.a-(XXX).sub.i-N.sub.b-YYY-N.sub.b-(ZZZ)j-N.sub.a-- n.sub.q3'
antisense: 3'n.sub.p'-N.sub.a'-(X'X'X').sub.k-N.sub.b'-Y'Y'Y'-N.sub.b'-(Z'Z'Z').sub.- i-N.sub.a'-n.sub.q'5' (III)
[0218] wherein:
[0219] i, j, k, and l are each independently 0 or 1;
[0220] p, p', q, and q' are each independently 0-6;
[0221] each N.sub.a and N.sub.a' independently represents an oligonucleotide sequence comprising 0-25 modified nucleotides, each sequence comprising at least two differently modified nucleotides;
[0222] each N.sub.b and N.sub.b' independently represents an oligonucleotide sequence comprising 0-10 modified nucleotides;
[0223] wherein
[0224] each n.sub.p', n.sub.p, n.sub.q', and n.sub.q independently represents an overhang nucleotide; and
[0225] XXX, YYY, ZZZ, X'X'X', Y'Y'Y', and Z'Z'Z' each independently represent one motif of three identical modifications on three consecutive nucleotides.
[0226] In one embodiment, i is 1 and j is 0; or i is 0 and j is 1; or both i and j are 1. In another embodiment, k is 1 and l is 0; k is 0 and l is 1; or both k and l are 1.
[0227] Exemplary combinations of the sense strand and antisense strand forming a RNAi duplex include the formulas below:
5'n.sub.p-N.sub.a-YYY-N.sub.b-ZZZ-N.sub.a-n.sub.q3'
3'n.sub.p'-N.sub.a'-Y'Y'Y'-N.sub.b'-Z'Z'Z'-N.sub.a'n.sub.q'5' (IIIa)
5'n.sub.p-N.sub.a-XXX-N.sub.b-YYY-N.sub.a-n.sub.q3'
3'n.sub.p'-N.sub.a'-X'X'X'-N.sub.b'-Y'Y'Y'-N.sub.a'-n.sub.q'5' (IIIb)
5'n.sub.p-N.sub.a-XXX-N.sub.b-YYY-N.sub.b-ZZZ-N.sub.a-n.sub.q3'
3'n.sub.p'-N.sub.a'-X'X'X'-N.sub.b'-Y'Y'Y'-N.sub.b'-Z'Z'Z'-N.sub.a-n.sub- .q'5' (IIIc)
[0228] When the RNAi agent is represented by formula (IIIa), each N.sub.b independently represents an oligonucleotide sequence comprising 1-10, 1-7, 1-5 or 1-4 modified nucleotides. Each N.sub.a independently represents an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0229] When the RNAi agent is represented as formula (IIIb), each N.sub.b, N.sub.b' independently represents an oligonucleotide sequence comprising 0-10, 0-7, 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a independently represents an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides.
[0230] When the RNAi agent is represented as formula (IIIc), each N.sub.b, N.sub.b' independently represents an oligonucleotide sequence comprising 0-10, 0-7, 0-10, 0-7, 0-5, 0-4, 0-2 or 0 modified nucleotides. Each N.sub.a, N.sub.a' independently represents an oligonucleotide sequence comprising 2-20, 2-15, or 2-10 modified nucleotides. Each of N.sub.a, N.sub.a', N.sub.b and N.sub.b' independently comprises modifications of alternating pattern.
[0231] Each of X, Y and Z in formulas (III), (IIIa), (IIIb) and (IIIc) may be the same or different from each other.
[0232] When the RNAi agent is represented by formula (III), (IIIa), (IIIb) or (IIIc), at least one of the Y nucleotides may form a base pair with one of the Y' nucleotides. Alternatively, at least two of the Y nucleotides form base pairs with the corresponding Y' nucleotides; or all three of the Y nucleotides all form base pairs with the corresponding Y' nucleotides.
[0233] When the RNAi agent is represented by formula (IIIa) or (IIIc), at least one of the Z nucleotides may form a base pair with one of the Z' nucleotides. Alternatively, at least two of the Z nucleotides form base pairs with the corresponding Z' nucleotides; or all three of the Z nucleotides all form base pairs with the corresponding Z' nucleotides.
[0234] When the RNAi agent is represented as formula (IIIb) or (IIIc), at least one of the X nucleotides may form a base pair with one of the X' nucleotides. Alternatively, at least two of the X nucleotides form base pairs with the corresponding X' nucleotides; or all three of the X nucleotides all form base pairs with the corresponding X' nucleotides.
[0235] In one embodiment, the modification on the Y nucleotide is different than the modification on the Y' nucleotide, the modification on the Z nucleotide is different than the modification on the Z' nucleotide, and/or the modification on the X nucleotide is different than the modification on the X' nucleotide.
[0236] In one embodiment, the RNAi agent is a multimer containing at least two duplexes represented by formula (III), (IIIa), (IIIb) or (IIIc), wherein the duplexes are connected by a linker. The linker can be cleavable or non-cleavable. Optionally, the multimer further comprise a ligand. Each of the duplexes can target the same gene or two different genes; or each of the duplexes can target same gene at two different target sites.
[0237] In one embodiment, the RNAi agent is a multimer containing three, four, five, six or more duplexes represented by formula (III), (IIIa), (IIIb) or (IIIc), wherein the duplexes are connected by a linker. The linker can be cleavable or non-cleavable. Optionally, the multimer further comprises a ligand. Each of the duplexes can target the same gene or two different genes; or each of the duplexes can target same gene at two different target sites.
[0238] In one embodiment, two RNAi agents represented by formula (III), (IIIa), (IIIb) or (IIIc) are linked to each other at the 5' end, and one or both of the 3' ends of the are optionally conjugated to to a ligand. Each of the agents can target the same gene or two different genes; or each of the agents can target same gene at two different target sites.
[0239] Various publications describe multimeric RNAi agents. Such publications include WO2007/091269, U.S. Pat. No. 7,858,769, WO2010/141511, WO2007/117686, WO2009/014887 and WO2011/031520 the entire contents of which are hereby incorporated herein by reference.
[0240] The RNAi agent that contains conjugations of one or more carbohydrate moieties to a RNAi agent can optimize one or more properties of the RNAi agent. In many cases, the carbohydrate moiety will be attached to a modified subunit of the RNAi agent. For example, the ribose sugar of one or more ribonucleotide subunits of a dsRNA agent can be replaced with another moiety, e.g., a non-carbohydrate (preferably cyclic) carrier to which is attached a carbohydrate ligand. A ribonucleotide subunit in which the ribose sugar of the subunit has been so replaced is referred to herein as a ribose replacement modification subunit (RRMS). A cyclic carrier may be a carbocyclic ring system, i.e., all ring atoms are carbon atoms, or a heterocyclic ring system, i.e., one or more ring atoms may be a heteroatom, e.g., nitrogen, oxygen, sulfur. The cyclic carrier may be a monocyclic ring system, or may contain two or more rings, e.g. fused rings. The cyclic carrier may be a fully saturated ring system, or it may contain one or more double bonds.
[0241] The ligand may be attached to the polynucleotide via a carrier. The carriers include (i) at least one "backbone attachment point," preferably two "backbone attachment points" and (ii) at least one "tethering attachment point." A "backbone attachment point" as used herein refers to a functional group, e.g. a hydroxyl group, or generally, a bond available for, and that is suitable for incorporation of the carrier into the backbone, e.g., the phosphate, or modified phosphate, e.g., sulfur containing, backbone, of a ribonucleic acid. A "tethering attachment point" (TAP) in some embodiments refers to a constituent ring atom of the cyclic carrier, e.g., a carbon atom or a heteroatom (distinct from an atom which provides a backbone attachment point), that connects a selected moiety. The moiety can be, e.g., a carbohydrate, e.g. monosaccharide, disaccharide, trisaccharide, tetrasaccharide, oligosaccharide and polysaccharide. Optionally, the selected moiety is connected by an intervening tether to the cyclic carrier. Thus, the cyclic carrier will often include a functional group, e.g., an amino group, or generally, provide a bond, that is suitable for incorporation or tethering of another chemical entity, e.g., a ligand to the constituent ring.
[0242] The RNAi agents may be conjugated to a ligand via a carrier, wherein the carrier can be cyclic group or acyclic group; preferably, the cyclic group is selected from pyrrolidinyl, pyrazolinyl, pyrazolidinyl, imidazolinyl, imidazolidinyl, piperidinyl, piperazinyl, [1,3]dioxolane, oxazolidinyl, isoxazolidinyl, morpholinyl, thiazolidinyl, isothiazolidinyl, quinoxalinyl, pyridazinonyl, tetrahydrofuryl and and decalin; preferably, the acyclic group is selected from serinol backbone or diethanolamine backbone.
[0243] In certain specific embodiments, the RNAi agent of the invention is an agent selected from the group of agents listed in Table 1 and consisting of D1000, D1001, D1002, D1003, D1004, D1005, D1006, D1007, D1008, D1009, D1010, D1011, D1012, D1013, D1014, D1015, D1016, D1017, D1018, D1019, D1020, D1021, D1022, D1023, D1024, D1025, D1026, D1027, D1028, D1029, D1030, D1031, D1032, D1033, D1034, D1035, D1036, D1037, D1038, D1039, D1040, D1041, D1042, D1043, D1044, D1045, D1046, D1047, D1048, D1049, D1050, D1051, D1052, D1053, D1054, D1055, D1056, D1057, D1058, D1059, D1060, D1061, D1062, D1063, D1064, D1065, D1066, D1067, D1068, D1069, D1070, D1071, D1072, D1073, D1074, D1075, D1076, D1077, D1078, D1079, D1080, D1081, D1082, D1083, D1084, D1085, D1086, D1087, D1088, D1089, D1090, D1091, D1092, D1093, D1094, D1095, D1096, D1097, D1098, D1099, D1100, D101, D1102, D1103, D1104, D1105, D1106, D1107, D1108, D1109, D1110, D1111, D112, D1113, D1114, D1115, D1116, D1117, D1118, D1119, D1120, D1121, D1122, D1123, D1124, D1125, D1126, D1127, D1128, D1129, D1130, D1131, D1132, D1133, D1134, D1135, D1136, D1137, D1138, D1139, D1140, D1141, D1142, D1143, D1144, D1145, D1146, D1147, D1148, D1149, D1150, D1151, D1152, D1153, D1154, D1155, D1156, D1157, D1158, D1159, D1160, D1161, D1162, D1163, D1164, D1165, D1166, D1167, D1168, D1169, D1170, D1171, D1172, D1173, D1174, D1175, D1176, D1177, D1178, D1179, D1180, D1181, D1182, D1183, D1184, D1185, D1186, D1187, D1188, D1189, D1190, D1191, D1192, D1193, D1194, D1195, D1196, D1197, D1198, D1199, D1200, D1201, D1202, D1203, D1204, D1205, D1206, D1207, D1208, D1209, D1210, D1211, D1212, D1213, D1214, D1215, D1216, D1217, D1218, D1219, D1220, D1221, D1222, D1223, D1224, D1225, D1226, D1227, D1228, D1229, D1230, D1231, D1232, D1233, D1234, D1235, D1236, D1237, D1238, D1239, D1240, D1241, D1242, D1243, D1244, D1245, D1246, D1247, D1248, D1249, D1250, D1251, D1252, D1253, D1254, D1255, D1256, D1257, D1258, D1259, D1260, D1261, D1262, D1263, D1264, D1265, D1266, D1267, D1268, D1269, D1270, D1271, D1272, D1273, D1274, D1275, D1276, D1277, D1278, D1279, D1280, D1281, D1282, D1283, D1284, D1285, D1286, D1287, D1288, D1289, D1290, D1291, D1292, D1293, D1294, D1295, D1296, D1297, D1298, D1299, D1300, D1301, D1302, D1303, D1304, D1305, D1306, D1307, D1308, D1309, D1310, D1311, D1312, D1313, D1314, D1315, D1316, D1317, D1318, D1319, D1320, D1321, D1322, D1323, D1324, D1325, D1326, D1327, D1328, D1329, D1330, D1331, D1332, D1333, D1334, D1335, D1336, D1337, D1338, D1339, D1340, D1341, D1342, D1343, D1344, D1345, D1346, D1347, D1348, D1349, D1350, D1351, D1352, D1353, D1354, D1355, D1356, D1357, D1358, D1359, D1360, D1361, D1362, D1363, D1364, D1365, D1366, D1367, D1368, D1369, D1370, D1371, D1372, D1373, D1374, D1375, D1376, D1377, D1378, D1379, D1380, D1381, D1382, D1383, D1384, D1385, D1386, D1387, D1388, D1389, D1390, D1391, D1392, D1393, D1394, D1395, D1396, D1397, D1398, D1399, D1400, D1401, D1402, D1403, D1404, D1405, D1406, D1407, D1408, D1409, D1410, D1411, D1412, D1413, D1414, D1415, D1416, D1417, D1418, D1419, D1420, D1421, D1422, D1423, D1424, D1425, D1426, D1427, D1428, D1429, D1430, D1431, D1432, D1433, D1434, D1435, D1436, D1437, D1438, D1439, D1440, D1441, D1442, D1443, D1444, D1445, D1446, D1447, D1448, D1449, D1450, D1451, D1452, D1453, D1454, D1455, D1456, D1457, D1458, D1459, D1460, D1461, D1462, D1463, D1464, D1465, D1466, D1467, D1468, D1469, D1470, D1471, D1472, D1473, D1474, D1475, D1476, D1477, D1478, D1479, D1480, D1481, D1482, D1483, D1484, D1485, D1486, D1487, D1488, D1489, D1490, D1491, D1492, D1493, D1494, D1495, D1496, D1497, D1498, D1499, D1500, D1501, D1502, D1503, D1504, D1505, D1506, D1507, D1508, D1509, D1510, D1511, D1512, D1513, D1514, D1515, D1516, D1517, D1518, D1519, D1520, D1521, D1522, D1523, D1524, D1525, D1526, D1527, D1528, D1529, D1530, D1531, D1532, D1533, D1534, D1535, D1536, D1537, D1538, D1539, D1540, D1541, D1542, D1543, D1544, D1545, D1546, D1547, D1548, D1549, D1550, D1551, D1552, D1553, D1554, D1555, D1556, D1557, D1558, D1559, D1560, D1561, D1562, D1563, D1564, D1565, D1566, D1567, D1568, D1569, D1570, D1571, D1572, D1573, D1574, D1575, D1576, D1577, D1578, D1579, D1580, D1581, D1582, D1583, D1584, D1585, D1586, D1587, D1588, D1589, D1590, D1591, D1592, D1593, D1594, D1595, D1596, D1597, D1598, D1599, D1600, D1601, D1602, D1603, D1604, D1605, D1606, D1607, D1608, D1609, D1610, D1611, D1612, D1613, D1614, D1615, D1616, D1617, D1618, D1619, D1620, D1621, D1622, D1623, D1624, D1625, D1626, D1627, D1628, D1629, D1630, D1631, D1632, D1633, D1634, D1635, D1636, D1637, D1638, D1639, D1640, D1641, D1642, D1643, D1644, D1645, D1646, D1647, D1648, D1649, D1650, D1651, D1652, D1653, D1654, D1655, D1656, D1657, D1658, D1659, D1660, D1661, D1662, D1663, D1664, D1665, D1666, D1667, D1668, D1669, D1670, D1671, D1672, D1673, D1674, D1675, D1676, D1677, D1678, D1679, D1680, D1681, D1682, D1683, D1684, D1685, D1686, D1687, D1688, D1689, D1690, D1691, D1692, D1693, D1694, D1695, D1696, D1697, D1698, D1699, D1700, D1701, D1702, D1703, D1704, D1705, D1706, D1707, D1708, D1709, D1710, D1711, D1712, D1713, D1714, D1715, D1716, D1717, D1718, D1719, D1720, D1721, D1722, D1723, D1724, D1725, D1726, D1727, D1728, D1729, D1730, D1731, D1732, D1733, D1734, D1735, D1736, D1737, D1738, D1739, D1740, D1741, D1742, D1743, D1744, D1745, D1746, D1747, D1748, D1749, D1750, D1751, D1752, D1753, D1754, D1755, D1756, D1757, D1758, D1759, D1760, D1761, D1762, D1763, D1764, D1765, D1766, D1767, D1768, D1769, D1770, D1771, D1772, D1773, D1774, D1775, D1776, D1777, D1778, D1779, D1780, D1781, D1782, D1783, D1784, D1785, D1786, D1787, D1788, D1789, D1790, D1791, D1792, D1793, D1794, D1795, D1796, D1797, D1798, D1799, D1800, D1801, D1802, D1803, D1804, D1805, D1806, D1807, D1808, D1809, D1810, D1811, D1812, D1813, D1814, D1815, D1816, D1817, D1818, D1819, D1820, D1821, D1822, D1823, D1824, D1825, D1826, D1827, D1828, D1829, D1830, D1831, D1832, D1833, D1834, D1835, D1836, D1837, D1838, D1839, D1840, D1841, D1842, D1843, D1844, D1845, D1846, D1847, D1848, D1849, D1850, D1851, D1852, D1853, D1854, D1855, D1856, D1857, D1858, D1859, D1860, D1861, D1862, D1863, D1864, D1865, D1866, D1867, D1868, D1869, D1870, D1871, D1872, D1873, D1874, D1875, D1876, D1877, D1878, D1879, D1880, D1881, D1882, D1883, D1884, D1885, D1886, D1887, D1888, D1889, D1890, D1891, D1892, D1893, D1894, D1895, D1896, D1897, D1898, D1899, D1900, D1901, D1902, D1903, D1904, D1905, D1906, D1907, D1908, D1909, D1910, D1911, D1912, D1913, D1914, D1915, D1916, D1917, D1918, D1919, D1920, D1921, D1922, D1923, D1924, D1925, D1926, D1927, D1928, D1929, D1930, D1931, D1932, D1933, D1934, D1935, D1936, D1937, D1938, D1939, D1940, D1941, D1942, D1943, D1944, D1945, D1946, D1947, D1948, D1949, D1950, D1951, D1952, D1953, D1954, D1955, D1956, D1957, D1958, D1959, D1960, D1961, D1962, D1963, D1964, D1965, D1966, D1967, D1968, D1969, D1970, D1971, D1972, D1973, D1974, D1975, D1976, D1977, D1978, D1979, D1980, D1981, D1982, D1983, D1984, D1985, D1986, D1987, D1988, D1989, D1990, D1991, D1992, D1993, D1994, D1995, D1996, D1997, D1998, D1999, D2000, D2001, D2002, D2003, D2004, D2005, D2006, D2007, D2008, D2009, D2010, D2011, D2012, D2013, D2014, D2015, D2016, D2017, D2018, D2019, D2020, D2021, D2022, D2023, D2024, D2025, D2026, D2027, D2028, D2029, D2030, D2031, D2032, D2033, D2034, D2035, D2036, D2037, D2038, D2039, D2040, D2041, D2042, D2043, D2044, D2045, D2046, D2047, D2048, D2049, D2050, D2051, D2052, D2053, D2054, D2055, D2056, D2057, D2058, D2059, D2060, D2061, D2062, D2063, D2064, D2065, D2066, D2067, D2068, D2069, D2070, D2071, D2072, D2073, D2074, D2075, D2076, D2077, D2078, D2079, D2080, D2081, D2082, D2083, D2084, D2085, D2086, D2087, D2088, D2089, D2090 and D2091.
[0244] These agents may further comprise a ligand, such as a GalNAc ligand.
Ligands
[0245] The RNAi agents of the invention, e.g., double stranded RNAi agents, may optionally be conjugated to one or more ligands. The ligand can be attached to the sense strand, antisense strand or both strands, at the 3'-end, 5'-end or both ends. For instance, the ligand may be conjugated to the sense strand. In preferred embodiments, the ligand is conjugated to the 3'-end of the sense strand. In one preferred embodiment, the ligand is a GalNAc ligand. In particularly preferred embodiments, the ligand is GalNAc.sub.3:
##STR00004##
[0246] A wide variety of entities can be coupled to the RNAi agents of the present invention. Preferred moieties are ligands, which are coupled, preferably covalently, either directly or indirectly via an intervening tether.
[0247] In preferred embodiments, a ligand alters the distribution, targeting or lifetime of the molecule into which it is incorporated. In preferred embodiments a ligand provides an enhanced affinity for a selected target, e.g., molecule, cell or cell type, compartment, receptor e.g., a cellular or organ compartment, tissue, organ or region of the body, as, e.g., compared to a species absent such a ligand. Ligands providing enhanced affinity for a selected target are also termed targeting ligands.
[0248] Some ligands can have endosomolytic properties. The endosomolytic ligands promote the lysis of the endosome and/or transport of the composition of the invention, or its components, from the endosome to the cytoplasm of the cell. The endosomolytic ligand may be a polyanionic peptide or peptidomimetic which shows pH-dependent membrane activity and fusogenicity. In one embodiment, the endosomolytic ligand assumes its active conformation at endosomal pH. The "active" conformation is that conformation in which the endosomolytic ligand promotes lysis of the endosome and/or transport of the composition of the invention, or its components, from the endosome to the cytoplasm of the cell. Exemplary endosomolytic ligands include the GALA peptide (Subbarao et al., Biochemistry, 1987, 26: 2964-2972), the EALA peptide (Vogel et al., J. Am. Chem. Soc., 1996, 118: 1581-1586), and their derivatives (Turk et al., Biochem. Biophys. Acta, 2002, 1559: 56-68). In one embodiment, the endosomolytic component may contain a chemical group (e.g., an amino acid) which will undergo a change in charge or protonation in response to a change in pH. The endosomolytic component may be linear or branched.
[0249] Ligands can improve transport, hybridization, and specificity properties and may also improve nuclease resistance of the resultant natural or modified oligoribonucleotide, or a polymeric molecule comprising any combination of monomers described herein and/or natural or modified ribonucleotides.
[0250] Ligands in general can include therapeutic modifiers, e.g., for enhancing uptake; diagnostic compounds or reporter groups e.g., for monitoring distribution; cross-linking agents; and nuclease-resistance conferring moieties. General examples include lipids, steroids, vitamins, sugars, proteins, peptides, polyamines, and peptide mimics.
[0251] Ligands can include a naturally occurring substance, such as a protein (e.g., human serum albumin (HSA), low-density lipoprotein (LDL), high-density lipoprotein (HDL), or globulin); a carbohydrate (e.g., a dextran, pullulan, chitin, chitosan, inulin, cyclodextrin or hyaluronic acid); or a lipid. The ligand may also be a recombinant or synthetic molecule, such as a synthetic polymer, e.g., a synthetic polyamino acid, an oligonucleotide (e.g., an aptamer). Examples of polyamino acids include polyamino acid is a polylysine (PLL), poly L-aspartic acid, poly L-glutamic acid, styrene-maleic acid anhydride copolymer, poly(L-lactide-co-glycolied) copolymer, divinyl ether-maleic anhydride copolymer, N-(2-hydroxypropyl)methacrylamide copolymer (HMPA), polyethylene glycol (PEG), polyvinyl alcohol (PVA), polyurethane, poly(2-ethylacryllic acid), N-isopropylacrylamide polymers, or polyphosphazine. Example of polyamines include: polyethylenimine, polylysine (PLL), spermine, spermidine, polyamine, pseudopeptide-polyamine, peptidomimetic polyamine, dendrimer polyamine, arginine, amidine, protamine, cationic lipid, cationic porphyrin, quaternary salt of a polyamine, or an alpha helical peptide.
[0252] Ligands can also include targeting groups, e.g., a cell or tissue targeting agent, e.g., a lectin, glycoprotein, lipid or protein, e.g., an antibody, that binds to a specified cell type such as a kidney cell. A targeting group can be a thyrotropin, melanotropin, lectin, glycoprotein, surfactant protein A, Mucin carbohydrate, multivalent lactose, multivalent galactose, N-acetyl-galactosamine, N-acetyl-gulucosamine multivalent mannose, multivalent fucose, glycosylated polyaminoacids, multivalent galactose, transferrin, bisphosphonate, polyglutamate, polyaspartate, a lipid, cholesterol, a steroid, bile acid, folate, vitamin B12, biotin, an RGD peptide, an RGD peptide mimetic or an aptamer.
[0253] Other examples of ligands include dyes, intercalating agents (e.g., acridines), cross-linkers (e.g., psoralene, mitomycin C), porphyrins (TPPC4, texaphyrin, Sapphyrin), polycyclic aromatic hydrocarbons (e.g., phenazine, dihydrophenazine), artificial endonucleases or a chelator (e.g., EDTA), lipophilic molecules, e.g., cholesterol, cholic acid, adamantane acetic acid, 1-pyrene butyric acid, dihydrotestosterone, 1,3-Bis-O(hexadecyl)glycerol, geranyloxyhexyl group, hexadecylglycerol, borneol, menthol, 1,3-propanediol, heptadecyl group, palmitic acid, myristic acid, O3-(oleoyl)lithocholic acid, O3-(oleoyl)cholenic acid, dimethoxytrityl, or phenoxazine) and peptide conjugates (e.g., antennapedia peptide, Tat peptide), alkylating agents, phosphate, amino, mercapto, PEG (e.g., PEG-40K), MPEG, [MPEG].sub.2, polyamino, alkyl, substituted alkyl, radiolabeled markers, enzymes, haptens (e.g., biotin), transport/absorption facilitators (e.g., aspirin, vitamin E, folic acid), synthetic ribonucleases (e.g., imidazole, bisimidazole, histamine, imidazole clusters, acridine-imidazole conjugates, Eu3+ complexes of tetraazamacrocycles), dinitrophenyl, HRP, or AP.
[0254] Ligands can be proteins, e.g., glycoproteins, or peptides, e.g., molecules having a specific affinity for a co-ligand, or antibodies e.g., an antibody, that binds to a specified cell type such as a cancer cell, endothelial cell, or bone cell. Ligands may also include hormones and hormone receptors. They can also include non-peptidic species, such as lipids, lectins, carbohydrates, vitamins, cofactors, multivalent lactose, multivalent galactose, N-acetyl-galactosamine, N-acetyl-gulucosamine multivalent mannose, multivalent fucose, or aptamers. The ligand can be, for example, a lipopolysaccharide, an activator of p38 MAP kinase, or an activator of NF-.kappa.B.
[0255] The ligand can be a substance, e.g., a drug, which can increase the uptake of the iRNA agent into the cell, for example, by disrupting the cell's cytoskeleton, e.g., by disrupting the cell's microtubules, microfilaments, and/or intermediate filaments. The drug can be, for example, taxon, vincristine, vinblastine, cytochalasin, nocodazole, japlakinolide, latrunculin A, phalloidin, swinholide A, indanocine, or myoservin.
[0256] The ligand can increase the uptake of the oligonucleotide into the cell by, for example, activating an inflammatory response. Exemplary ligands that would have such an effect include tumor necrosis factor alpha (TNFalpha), interleukin-1 beta, or gamma interferon.
[0257] In one aspect, the ligand is a lipid or lipid-based molecule. Such a lipid or lipid-based molecule preferably binds a serum protein, e.g., human serum albumin (HSA). An HSA binding ligand allows for distribution of the conjugate to a target tissue, e.g., a non-kidney target tissue of the body. For example, the target tissue can be the liver, including parenchymal cells of the liver. Other molecules that can bind HSA can also be used as ligands. For example, naproxen or aspirin can be used. A lipid or lipid-based ligand can (a) increase resistance to degradation of the conjugate, (b) increase targeting or transport into a target cell or cell membrane, and/or (c) can be used to adjust binding to a serum protein, e.g., HSA.
[0258] A lipid based ligand can be used to modulate, e.g., control the binding of the conjugate to a target tissue. For example, a lipid or lipid-based ligand that binds to HSA more strongly will be less likely to be targeted to the kidney and therefore less likely to be cleared from the body. A lipid or lipid-based ligand that binds to HSA less strongly can be used to target the conjugate to the kidney.
[0259] In a preferred embodiment, the lipid based ligand binds HSA. Preferably, it binds HSA with a sufficient affinity such that the conjugate will be preferably distributed to a non-kidney tissue. However, it is preferred that the affinity not be so strong that the HSA-ligand binding cannot be reversed.
[0260] In another preferred embodiment, the lipid based ligand binds HSA weakly or not at all, such that the conjugate will be preferably distributed to the kidney. Other moieties that target to kidney cells can also be used in place of or in addition to the lipid based ligand.
[0261] In another aspect, the ligand is a moiety, e.g., a vitamin, which is taken up by a target cell, e.g., a proliferating cell. These are particularly useful for treating disorders characterized by unwanted cell proliferation, e.g., of the malignant or non-malignant type, e.g., cancer cells. Exemplary vitamins include vitamin A, E, and K. Other exemplary vitamins include B vitamins, e.g., folic acid, B12, riboflavin, biotin, pyridoxal or other vitamins or nutrients taken up by cancer cells. Also included are HAS, low density lipoprotein (LDL) and high-density lipoprotein (HDL).
[0262] In another aspect, the ligand is a cell-permeation agent, preferably a helical cell-permeation agent. Preferably, the agent is amphipathic. An exemplary agent is a peptide such as tat or antennopedia. If the agent is a peptide, it can be modified, including a peptidylmimetic, invertomers, non-peptide or pseudo-peptide linkages, and use of D-amino acids. The helical agent is preferably an alpha-helical agent, which preferably has a lipophilic and a lipophobic phase.
[0263] The ligand can be a peptide or peptidomimetic. A peptidomimetic (also referred to herein as an oligopeptidomimetic) is a molecule capable of folding into a defined three-dimensional structure similar to a natural peptide. The peptide or peptidomimetic moiety can be about 5-50 amino acids long, e.g., about 5, 10, 15, 20, 25, 30, 35, 40, 45, or 50 amino acids long. A peptide or peptidomimetic can be, for example, a cell permeation peptide, cationic peptide, amphipathic peptide, or hydrophobic peptide (e.g., consisting primarily of Tyr, Trp or Phe). The peptide moiety can be a dendrimer peptide, constrained peptide or crosslinked peptide. In another alternative, the peptide moiety can include a hydrophobic membrane translocation sequence (MTS). An exemplary hydrophobic MTS-containing peptide is RFGF having the amino acid sequence AAVALLPAVLLALLAP (SEQ ID NO:4). An RFGF analogue (e.g., amino acid sequence AALLPVLLAAP) (SEQ ID NO:5) containing a hydrophobic MTS can also be a targeting moiety. The peptide moiety can be a "delivery" peptide, which can carry large polar molecules including peptides, oligonucleotides, and protein across cell membranes. For example, sequences from the HIV Tat protein (GRKKRRQRRRPPQ) (SEQ ID NO:6) and the Drosophila Antennapedia protein (RQIKIWFQNRRMKWKK) (SEQ ID NO:7) have been found to be capable of functioning as delivery peptides. A peptide or peptidomimetic can be encoded by a random sequence of DNA, such as a peptide identified from a phage-display library, or one-bead-one-compound (OBOC) combinatorial library (Lam et al., Nature, 354:82-84, 1991). Preferably the peptide or peptidomimetic tethered to an iRNA agent via an incorporated monomer unit is a cell targeting peptide such as an arginine-glycine-aspartic acid (RGD)-peptide, or RGD mimic. A peptide moiety can range in length from about 5 amino acids to about 40 amino acids. The peptide moieties can have a structural modification, such as to increase stability or direct conformational properties. Any of the structural modifications described below can be utilized. An RGD peptide moiety can be used to target a tumor cell, such as an endothelial tumor cell or a breast cancer tumor cell (Zitzmann et al., Cancer Res., 62:5139-43, 2002). An RGD peptide can facilitate targeting of an iRNA agent to tumors of a variety of other tissues, including the lung, kidney, spleen, or liver (Aoki et al., Cancer Gene Therapy 8:783-787, 2001). Preferably, the RGD peptide will facilitate targeting of an iRNA agent to the kidney. The RGD peptide can be linear or cyclic, and can be modified, e.g., glycosylated or methylated to facilitate targeting to specific tissues. For example, a glycosylated RGD peptide can deliver an iRNA agent to a tumor cell expressing .alpha..sub.V.beta..sub.3 (Haubner et al., Jour. Nucl. Med., 42:326-336, 2001). Peptides that target markers enriched in proliferating cells can be used. For example, RGD containing peptides and peptidomimetics can target cancer cells, in particular cells that exhibit an integrin. Thus, one could use RGD peptides, cyclic peptides containing RGD, RGD peptides that include D-amino acids, as well as synthetic RGD mimics. In addition to RGD, one can use other moieties that target the integrin ligand. Generally, such ligands can be used to control proliferating cells and angiogeneis. Preferred conjugates of this type of ligand target PECAM-1, VEGF, or other cancer gene, e.g., a cancer gene described herein.
[0264] A "cell permeation peptide" is capable of permeating a cell, e.g., a microbial cell, such as a bacterial or fungal cell, or a mammalian cell, such as a human cell. A microbial cell-permeating peptide can be, for example, an .alpha.-helical linear peptide (e.g., LL-37 or Ceropin P1), a disulfide bond-containing peptide (e.g., .alpha.-defensin, .beta.-defensin or bactenecin), or a peptide containing only one or two dominating amino acids (e.g., PR-39 or indolicidin). A cell permeation peptide can also include a nuclear localization signal (NLS). For example, a cell permeation peptide can be a bipartite amphipathic peptide, such as MPG, which is derived from the fusion peptide domain of HIV-1 gp41 and the NLS of SV40 large T antigen (Simeoni et al., Nucl. Acids Res. 31:2717-2724, 2003).
[0265] In one embodiment, a targeting peptide can be an amphipathic .alpha.-helical peptide. Exemplary amphipathic .alpha.-helical peptides include, but are not limited to, cecropins, lycotoxins, paradaxins, buforin, CPF, bombinin-like peptide (BLP), cathelicidins, ceratotoxins, S. clava peptides, hagfish intestinal antimicrobial peptides (HFIAPs), magainines, brevinins-2, dermaseptins, melittins, pleurocidin, H2A peptides, Xenopus peptides, esculentinis-1, and caerins. A number of factors will preferably be considered to maintain the integrity of helix stability. For example, a maximum number of helix stabilization residues will be utilized (e.g., leu, ala, or lys), and a minimum number helix destabilization residues will be utilized (e.g., proline, or cyclic monomeric units. The capping residue will be considered (for example Gly is an exemplary N-capping residue and/or C-terminal amidation can be used to provide an extra H-bond to stabilize the helix. Formation of salt bridges between residues with opposite charges, separated by i.+-.3, or i.+-.4 positions can provide stability. For example, cationic residues such as lysine, arginine, homo-arginine, ornithine or histidine can form salt bridges with the anionic residues glutamate or aspartate.
[0266] Peptide and peptidomimetic ligands include those having naturally occurring or modified peptides, e.g., D or L peptides; .alpha., .beta., or .gamma. peptides; N-methyl peptides; azapeptides; peptides having one or more amide, i.e., peptide, linkages replaced with one or more urea, thiourea, carbamate, or sulfonyl urea linkages; or cyclic peptides.
[0267] The targeting ligand can be any ligand that is capable of targeting a specific receptor. Examples are: folate, GalNAc, galactose, mannose, mannose-6P, clusters of sugars such as GalNAc cluster, mannose cluster, galactose cluster, or an apatamer. A cluster is a combination of two or more sugar units. The targeting ligands also include integrin receptor ligands, Chemokine receptor ligands, transferrin, biotin, serotonin receptor ligands, PSMA, endothelin, GCPII, somatostatin, LDL and HDL ligands. The ligands can also be based on nucleic acid, e.g., an aptamer. The aptamer can be unmodified or have any combination of modifications disclosed herein.
[0268] Endosomal release agents include imidazoles, poly or oligoimidazoles, PEIs, peptides, fusogenic peptides, polycaboxylates, polyacations, masked oligo or poly cations or anions, acetals, polyacetals, ketals/polyketyals, orthoesters, polymers with masked or unmasked cationic or anionic charges, dendrimers with masked or unmasked cationic or anionic charges.
[0269] PK modulator stands for pharmacokinetic modulator. PK modulators include lipophiles, bile acids, steroids, phospholipid analogues, peptides, protein binding agents, PEG, vitamins etc. Examplary PK modulators include, but are not limited to, cholesterol, fatty acids, cholic acid, lithocholic acid, dialkylglycerides, diacylglyceride, phospholipids, sphingolipids, naproxen, ibuprofen, vitamin E, biotin etc. Oligonucleotides that comprise a number of phosphorothioate linkages are also known to bind to serum protein, thus short oligonucleotides, e.g., oligonucleotides of about 5 bases, 10 bases, 15 bases or 20 bases, comprising multiple phosphorothioate linkages in the backbaone are also amenable to the present invention as ligands (e.g., as PK modulating ligands).
[0270] In addition, aptamers that bind serum components (e.g., serum proteins) are also amenable to the present invention as PK modulating ligands.
[0271] Other ligand conjugates amenable to the invention are described in U.S. patent application Ser. No. 10/916,185, filed Aug. 10, 2004; U.S. Ser. No. 10/946,873, filed Sep. 21, 2004; U.S. Ser. No. 10/833,934, filed Aug. 3, 2007; U.S. Ser. No. 11/115,989 filed Apr. 27, 2005 and U.S. Ser. No. 11/944,227 filed Nov. 21, 2007, which are incorporated by reference in their entireties for all purposes.
[0272] When two or more ligands are present, the ligands can all have same properties, all have different properties or some ligands have the same properties while others have different properties. For example, a ligand can have targeting properties, have endosomolytic activity or have PK modulating properties. In a preferred embodiment, all the ligands have different properties.
[0273] Ligands can be coupled to the oligonucleotides at various places, for example, 3'-end, 5'-end, and/or at an internal position. In preferred embodiments, the ligand is attached to the oligonucleotides via an intervening tether, e.g., a carrier described herein. The ligand or tethered ligand may be present on a monomer when the monomer is incorporated into the growing strand. In some embodiments, the ligand may be incorporated via coupling to a "precursor" monomer after the "precursor" monomer has been incorporated into the growing strand. For example, a monomer having, e.g., an amino-terminated tether (i.e., having no associated ligand), e.g., TAP-(CH.sub.2).sub.nNH.sub.2 may be incorporated into a growing oligonucleotide strand. In a subsequent operation, i.e., after incorporation of the precursor monomer into the strand, a ligand having an electrophilic group, e.g., a pentafluorophenyl ester or aldehyde group, can subsequently be attached to the precursor monomer by coupling the electrophilic group of the ligand with the terminal nucleophilic group of the precursor monomer's tether.
[0274] In another example, a monomer having a chemical group suitable for taking part in Click Chemistry reaction may be incorporated, e.g., an azide or alkyne terminated tether/linker. In a subsequent operation, i.e., after incorporation of the precursor monomer into the strand, a ligand having complementary chemical group, e.g. an alkyne or azide can be attached to the precursor monomer by coupling the alkyne and the azide together.
[0275] For double-stranded oligonucleotides, ligands can be attached to one or both strands. In some embodiments, a double-stranded iRNA agent contains a ligand conjugated to the sense strand. In other embodiments, a double-stranded iRNA agent contains a ligand conjugated to the antisense strand.
[0276] In some embodiments, ligand can be conjugated to nucleobases, sugar moieties, or internucleosidic linkages of nucleic acid molecules. Conjugation to purine nucleobases or derivatives thereof can occur at any position including, endocyclic and exocyclic atoms. In some embodiments, the 2-, 6-, 7-, or 8-positions of a purine nucleobase are attached to a conjugate moiety. Conjugation to pyrimidine nucleobases or derivatives thereof can also occur at any position. In some embodiments, the 2-, 5-, and 6-positions of a pyrimidine nucleobase can be substituted with a conjugate moiety. Conjugation to sugar moieties of nucleosides can occur at any carbon atom. Example carbon atoms of a sugar moiety that can be attached to a conjugate moiety include the 2', 3', and 5' carbon atoms. The 1' position can also be attached to a conjugate moiety, such as in an abasic residue. Internucleosidic linkages can also bear conjugate moieties. For phosphorus-containing linkages (e.g., phosphodiester, phosphorothioate, phosphorodithiotate, phosphoroamidate, and the like), the conjugate moiety can be attached directly to the phosphorus atom or to an O, N, or S atom bound to the phosphorus atom. For amine- or amide-containing internucleosidic linkages (e.g., PNA), the conjugate moiety can be attached to the nitrogen atom of the amine or amide or to an adjacent carbon atom.
[0277] Any suitable ligand in the field of RNA interference may be used, although the ligand is typically a carbohydrate e.g. monosaccharide (such as GalNAc), disaccharide, trisaccharide, tetrasaccharide, polysaccharide.
[0278] Linkers that conjugate the ligand to the nucleic acid include those discussed above. For example, the ligand can be one or more GalNAc (N-acetylglucosamine) derivatives attached through a bivalent or trivalent branched linker.
[0279] In one embodiment, the dsRNA of the invention is conjugated to a bivalent and trivalent branched linkers include the structures shown in any of formula (IV)-(VII):
##STR00005##
wherein.
[0280] q.sup.2A, q.sup.2B, q.sup.3A, q.sup.3B q4.sup.A, q.sup.4B, q.sup.5A, q.sup.5B and q.sup.5C represent independently for each occurrence 0-20 and wherein the repeating unit can be the same or different; P.sup.2A, P.sup.2B, P.sup.3A, P.sup.3B, P.sup.4A, P.sup.4B, P.sup.5A, P.sup.5B, P.sup.5C, T.sup.2A, T.sup.2B, T.sup.3A, T.sup.3B, T.sup.4A, T.sup.4B, T.sup.4A, T.sup.5B, T.sup.5C are each independently for each occurrence absent, CO, NH, O, S, OC(O), NHC(O), CH.sub.2, CH.sub.2NH or CH.sub.2O;
[0281] Q.sup.2A, Q.sup.2B, Q.sup.3A, Q.sup.3B, Q.sup.4A, Q.sup.4B, Q.sup.5A, Q.sup.5B, Q.sup.5C are independently for each occurrence absent, alkylene, substituted alkylene wherin one or more methylenes can be interrupted or terminated by one or more of O, S, S(O), SO.sub.2, N(R.sup.N), C(R').dbd.C(R''), C.ident.C or C(O); R.sup.2A, R.sup.2B, R.sup.3A, R.sup.3B, R.sup.4A, R.sup.4B, R.sup.5A, R.sup.5B, R.sup.5C are each independently for each occurrence absent, NH, O, S, CH.sub.2, C(O)O, C(O)NH, NHCH(R.sup.a)C(O), --C(O)--CH(R.sup.a)--NH--, CO, CH.dbd.N--O,
##STR00006##
or heterocyclyl;
[0282] L.sup.2A, L.sup.2B, L.sup.3A, L.sup.3B, L.sup.4A, L.sup.4B, L.sup.5A, L.sup.5B, and L.sup.5C, represent the ligand; i.e. each independently for each occurrence a monosaccharide (such as GalNAc), disaccharide, trisaccharide, tetrasaccharide, oligosaccharide, or polysaccharide; and
[0283] R.sup.a is H or amino acid side chain.
[0284] Trivalent conjugating GalNAc derivatives are particularly useful for use with RNAi agents for inhibiting the expression of a target gene, such as those of formula (VII):
##STR00007##
[0285] wherein L.sup.5A, L.sup.5B and L.sup.5C represent a monosaccharide, such as GalNAc derivative. Examples of suitable bivalent and trivalent branched linker groups conjugating GalNAc derivatives include, but are not limited to, the following compounds:
##STR00008## ##STR00009## ##STR00010##
[0286] In other embodiments, the RNAi agent of the invention is an agent selected from the group consisting of AD-45163, AD-45165, AD-51544, AD-51545, AD-51546, and AD-51547.
III. Pharmaceutical Compositions
[0287] The RNAi agents of the invention may be formulated for administration in any convenient way for use in human or veterinary medicine, by analogy with other pharmaceuticals. The pharmaceutical compositions comprising RNAi agents of the invention may be, for example, solutions with or without a buffer, or compositions containing pharmaceutically acceptable carriers. Such compositions include, for example, aqueous or crystalline compositions, liposomal formulations, micellar formulations, emulsions, and gene therapy vectors.
[0288] In the methods of the invention, the RNAi agent may be administered in a solution. A free RNAi agent may be administered in an unbuffered solution, e.g., in saline or in water. Alternatively, the free siRNA may also be administered in a suitable buffer solution. The buffer solution may comprise acetate, citrate, prolamine, carbonate, or phosphate, or any combination thereof. In a preferred embodiment, the buffer solution is phosphate buffered saline (PBS). The pH and osmolarity of the buffer solution containing the RNAi agent can be adjusted such that it is suitable for administering to a subject.
[0289] In some embodiments, the buffer solution further comprises an agent for controlling the osmolarity of the solution, such that the osmolarity is kept at a desired value, e.g., at the physiologic values of the human plasma. Solutes which can be added to the buffer solution to control the osmolarity include, but are not limited to, proteins, peptides, amino acids, non-metabolized polymers, vitamins, ions, sugars, metabolites, organic acids, lipids, or salts. In some embodiments, the agent for controlling the osmolarity of the solution is a salt. In certain embodiments, the agent for controlling the osmolarity of the solution is sodium chloride or potassium chloride.
[0290] In other embodiments, the RNAi agent is formulated as a composition that includes one or more RNAi agents and a pharmaceutically acceptable carrier. As used herein the language "pharmaceutically acceptable carrier" is intended to include any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents, and the like, compatible with pharmaceutical administration. The use of such media and agents for pharmaceutically active substances is well known in the art. Except insofar as any conventional media or agent is incompatible with the active compound, use thereof in the compositions is contemplated. Supplementary active compounds can also be incorporated into the compositions.
[0291] In one embodiment, the RNAi agent preparation includes at least a second therapeutic agent (e.g., an agent other than an RNA or a DNA). For example, an RNAi agent composition for the treatment of a TTR-associated disease, e.g., a transthyretin-related hereditary amyloidosis (familial amyloid polyneuropathy, FAP), may include a known drug for the amelioration of FAP, e.g., Tafamidis (INN, or Fx-1006A or Vyndagel).
[0292] A formulated RNAi agent composition can assume a variety of states. In some examples, the composition is at least partially crystalline, uniformly crystalline, and/or anhydrous (e.g., it contains less than 80, 50, 30, 20, or 10% of water). In another example, the RNAi agent is in an aqueous phase, e.g., in a solution that includes water.
[0293] The aqueous phase or the crystalline compositions can be incorporated into a delivery vehicle, e.g., a liposome (particularly for the aqueous phase) or a particle (e.g., a microparticle as can be appropriate for a crystalline composition). Generally, the RNAi agent composition is formulated in a manner that is compatible with the intended method of administration, as described herein. For example, in particular embodiments the composition is prepared by at least one of the following methods: spray drying, lyophilization, vacuum drying, evaporation, fluid bed drying, or a combination of these techniques; or sonication with a lipid, freeze-drying, condensation and other self-assembly.
[0294] An RNAi agent preparation can be formulated in combination with another agent, e.g., another therapeutic agent or an agent that stabilizes RNAi agent, e.g., a protein that complexes with the RNAi agent to form an iRNP. Still other agents include chelators, e.g., EDTA (e.g., to remove divalent cations such as Mg.sup.2+), salts, RNAse inhibitors (e.g., a broad specificity RNAse inhibitor such as RNAsin) and so forth.
[0295] In one embodiment, the RNAi agent preparation includes another siRNA compound, e.g., a second RNAi agent that can mediate RNAi with respect to a second gene, or with respect to the same gene. Still other preparation can include at least 3, 5, ten, twenty, fifty, or a hundred or more different RNAi agent species. Such RNAi agents can mediate RNAi with respect to a similar number of different genes.
[0296] The iRNA agents of the invention may be formulated for pharmaceutical use. Pharmaceutically acceptable compositions comprise a therapeutically- or prophylactically effective amount of one or more of the the dsRNA agents in any of the preceding embodiments, taken alone or formulated together with one or more pharmaceutically acceptable carriers (additives), excipient and/or diluents.
[0297] Methods of preparing pharmaceutical compositions of the invention include the step of bringing into association an RNAi agent of the present invention with the carrier and, optionally, one or more accessory ingredients. In general, the compositions are prepared by uniformly and intimately bringing into association an RNAi agent of the present invention with liquid carriers, or finely divided solid carriers, or both, and then, if necessary, shaping the product.
[0298] The pharmaceutical compositions may be specially formulated for administration in solid or liquid form, including those adapted for the following: (1) oral administration, for example, drenches (aqueous or non-aqueous solutions or suspensions), tablets, e.g., those targeted for buccal, sublingual, and systemic absorption, boluses, powders, granules, pastes for application to the tongue; (2) parenteral administration, for example, by subcutaneous, intramuscular, intravenous or epidural injection as, for example, a sterile solution or suspension, or sustained-release formulation; (3) topical application, for example, as a cream, ointment, or a controlled-release patch or spray applied to the skin; (4) intravaginally or intrarectally, for example, as a pessary, cream or foam; (5) sublingually; (6) ocularly; (7) transdermally; or (8) nasally. Delivery using subcutaneous or intravenous methods can be particularly advantageous.
[0299] The phrase "pharmaceutically acceptable" is employed herein to refer to those compounds, materials, compositions, and/or dosage forms which are, within the scope of sound medical judgment, suitable for use in contact with the tissues of human beings and animals without excessive toxicity, irritation, allergic response, or other problem or complication, commensurate with a reasonable benefit/risk ratio.
[0300] The phrase "pharmaceutically-acceptable carrier" as used herein means a pharmaceutically-acceptable material, composition or vehicle, such as a liquid or solid filler, diluent, excipient, manufacturing aid (e.g., lubricant, talc magnesium, calcium or zinc stearate, or steric acid), or solvent encapsulating material, involved in carrying or transporting the subject compound from one organ, or portion of the body, to another organ, or portion of the body. Each carrier must be "acceptable" in the sense of being compatible with the other ingredients of the composition and not injurious to the patient. Some examples of materials which can serve as pharmaceutically-acceptable carriers include: (1) sugars, such as lactose, glucose and sucrose; (2) starches, such as corn starch and potato starch; (3) cellulose, and its derivatives, such as sodium carboxymethyl cellulose, ethyl cellulose and cellulose acetate; (4) powdered tragacanth; (5) malt; (6) gelatin; (7) lubricating agents, such as magnesium state, sodium lauryl sulfate and talc; (8) excipients, such as cocoa butter and suppository waxes; (9) oils, such as peanut oil, cottonseed oil, safflower oil, sesame oil, olive oil, corn oil and soybean oil; (10) glycols, such as propylene glycol; (11) polyols, such as glycerin, sorbitol, mannitol and polyethylene glycol; (12) esters, such as ethyl oleate and ethyl laurate; (13) agar; (14) buffering agents, such as magnesium hydroxide and aluminum hydroxide; (15) alginic acid; (16) pyrogen-free water; (17) isotonic saline; (18) Ringer's solution; (19) ethyl alcohol; (20) pH buffered solutions; (21) polyesters, polycarbonates and/or polyanhydrides; (22) bulking agents, such as polypeptides and amino acids (23) serum component, such as serum albumin, HDL and LDL; and (22) other non-toxic compatible substances employed in pharmaceutical compositions.
[0301] The compositions may conveniently be presented in unit dosage form and may be prepared by any methods well known in the art of pharmacy. The amount of RNAi agent which can be combined with a carrier material to produce a single dosage form will vary depending upon the host being treated, and the particular mode of administration. The RNAi agent which can be combined with a carrier material to produce a single dosage form will generally be that amount of the RNAi agent which produces a desired effect, e.g., therapeutic or prophylactic effect. Generally, out of one hundred percent, this amount will range from about 0.1 percent to about ninety-nine percent of RNAi agent, preferably from about 5 percent to about 70 percent, most preferably from about 10 percent to about 30 percent.
[0302] In certain embodiments, a composition of the present invention comprises an excipient selected from the group consisting of cyclodextrins, celluloses, liposomes, micelle forming agents, e.g., bile acids, and polymeric carriers, e.g., polyesters and polyanhydrides; and an RNAi agent of the present invention. In certain embodiments, an aforementioned composition renders orally bioavailable an RNAi agent of the present invention.
[0303] In some cases, in order to prolong the effect of an RNAi agent, it is desirable to slow the absorption of the agent from subcutaneous or intramuscular injection. This may be accomplished by the use of a liquid suspension of crystalline or amorphous material having poor water solubility. The rate of absorption of the RNAi agent then depends upon its rate of dissolution which, in turn, may depend upon crystal size and crystalline form. Alternatively, delayed absorption of a parenterally-administered RNAi agent may be accomplished by dissolving or suspending the agent in an oil vehicle.
[0304] Liposomes
[0305] An RNAi agent of the invention can be formulated for delivery in a membranous molecular assembly, e.g., a liposome or a micelle. As used herein, the term "liposome" refers to a vesicle composed of amphiphilic lipids arranged in at least one bilayer, e.g., one bilayer or a plurality of bilayers. Liposomes include unilamellar and multilamellar vesicles that have a membrane formed from a lipophilic material and an aqueous interior. The aqueous portion contains the RNAi agent composition. The lipophilic material isolates the aqueous interior from an aqueous exterior, which typically does not include the RNAi agent composition, although in some examples, it may. Liposomes are useful for the transfer and delivery of active ingredients to the site of action. Because the liposomal membrane is structurally similar to biological membranes, when liposomes are applied to a tissue, the liposomal bilayer fuses with bilayer of the cellular membranes. As the merging of the liposome and cell progresses, the internal aqueous contents that include the RNAi agent are delivered into the cell where the RNAi agent can specifically bind to a target RNA and can mediate RNAi. In some cases the liposomes are also specifically targeted, e.g., to direct the RNAi agent to particular cell types.
[0306] A liposome containing an RNAi agent can be prepared by a variety of methods. In one example, the lipid component of a liposome is dissolved in a detergent so that micelles are formed with the lipid component. For example, the lipid component can be an amphipathic cationic lipid or lipid conjugate. The detergent can have a high critical micelle concentration and may be nonionic. Exemplary detergents include cholate, CHAPS, octylglucoside, deoxycholate, and lauroyl sarcosine. The RNAi agent preparation is then added to the micelles that include the lipid component. The cationic groups on the lipid interact with the RNAi agent and condense around the RNAi agent to form a liposome. After condensation, the detergent is removed, e.g., by dialysis, to yield a liposomal preparation of RNAi agent.
[0307] If necessary a carrier compound that assists in condensation can be added during the condensation reaction, e.g., by controlled addition. For example, the carrier compound can be a polymer other than a nucleic acid (e.g., spermine or spermidine). pH can also be adjusted to favor condensation.
[0308] Methods for producing stable polynucleotide delivery vehicles, which incorporate a polynucleotide/cationic lipid complex as structural components of the delivery vehicle, are further described in, e.g., WO 96/37194, the entire contents of which are incorporated herein by reference. Liposome formation can also include one or more aspects of exemplary methods described in Felgner, P. L. et al., Proc. Nat. Acad. Sci., USA 8:7413-7417, 1987; U.S. Pat. Nos. 4,897,355; 5,171,678; Bangham, et al. M. Mol. Biol. 23:238, 1965; Olson, et al. Biochim. Biophys. Acta 557:9, 1979; Szoka, et al. Proc. Nat. Acad. Sci. 75: 4194, 1978; Mayhew, et al. Biochim. Biophys. Acta 775:169, 1984; Kim, et al. Biochim. Biophys. Acta 728:339, 1983; and Fukunaga, et al. Endocrinol. 115:757, 1984. Commonly used techniques for preparing lipid aggregates of appropriate size for use as delivery vehicles include sonication and freeze-thaw plus extrusion (see, e.g., Mayer, et al. Biochim. Biophys. Acta 858:161, 1986). Microfluidization can be used when consistently small (50 to 200 nm) and relatively uniform aggregates are desired (Mayhew, et al. Biochim. Biophys. Acta 775:169, 1984). These methods are readily adapted to packaging RNAi agent preparations into liposomes.
[0309] Liposomes that are pH-sensitive or negatively-charged entrap nucleic acid molecules rather than complex with them. Since both the nucleic acid molecules and the lipid are similarly charged, repulsion rather than complex formation occurs. Nevertheless, some nucleic acid molecules are entrapped within the aqueous interior of these liposomes. pH-sensitive liposomes have been used to deliver DNA encoding the thymidine kinase gene to cell monolayers in culture. Expression of the exogenous gene was detected in the target cells (Zhou et al., Journal of Controlled Release, 19, (1992) 269-274).
[0310] One major type of liposomal composition includes phospholipids other than naturally-derived phosphatidylcholine. Neutral liposome compositions, for example, can be formed from dimyristoyl phosphatidylcholine (DMPC) or dipalmitoyl phosphatidylcholine (DPPC). Anionic liposome compositions generally are formed from dimyristoyl phosphatidylglycerol, while anionic fusogenic liposomes are formed primarily from dioleoyl phosphatidylethanolamine (DOPE). Another type of liposomal composition is formed from phosphatidylcholine (PC) such as, for example, soybean PC, and egg PC. Another type is formed from mixtures of phospholipid and/or phosphatidylcholine and/or cholesterol.
[0311] Examples of other methods to introduce liposomes into cells in vitro and in vivo include U.S. Pat. Nos. 5,283,185; 5,171,678; WO 94/00569; WO 93/24640; WO 91/16024; Felgner, J. Biol. Chem. 269:2550, 1994; Nabel, Proc. Natl. Acad. Sci. 90:11307, 1993; Nabel, Human Gene Ther. 3:649, 1992; Gershon, Biochem. 32:7143, 1993; and Strauss EMBO J. 11:417, 1992.
[0312] In one embodiment, cationic liposomes are used. Cationic liposomes possess the advantage of being able to fuse to the cell membrane. Non-cationic liposomes, although not able to fuse as efficiently with the plasma membrane, are taken up by macrophages in vivo and can be used to deliver RNAi agents to macrophages.
[0313] Further advantages of liposomes include: liposomes obtained from natural phospholipids are biocompatible and biodegradable; liposomes can incorporate a wide range of water and lipid soluble drugs; liposomes can protect encapsulated RNAi agents in their internal compartments from metabolism and degradation (Rosoff, in "Pharmaceutical Dosage Forms," Lieberman, Rieger and Banker (Eds.), 1988, volume 1, p. 245). Important considerations in the preparation of liposome formulations are the lipid surface charge, vesicle size and the aqueous volume of the liposomes.
[0314] A positively charged synthetic cationic lipid, N-[1-(2,3-dioleyloxy)propyl]-N,N,N-trimethylammonium chloride (DOTMA) can be used to form small liposomes that interact spontaneously with nucleic acid to form lipid-nucleic acid complexes which are capable of fusing with the negatively charged lipids of the cell membranes of tissue culture cells, resulting in delivery of RNAi agent (see, e.g., Felgner, P. L. et al., Proc. Natl. Acad. Sci., USA 8:7413-7417, 1987 and U.S. Pat. No. 4,897,355 for a description of DOTMA and its use with DNA).
[0315] A DOTMA analogue, 1,2-bis(oleoyloxy)-3-(trimethylammonia)propane (DOTAP) can be used in combination with a phospholipid to form DNA-complexing vesicles. Lipofectin.TM. Bethesda Research Laboratories, Gaithersburg, Md.) is an effective agent for the delivery of highly anionic nucleic acids into living tissue culture cells that comprise positively charged DOTMA liposomes which interact spontaneously with negatively charged polynucleotides to form complexes. When enough positively charged liposomes are used, the net charge on the resulting complexes is also positive. Positively charged complexes prepared in this way spontaneously attach to negatively charged cell surfaces, fuse with the plasma membrane, and efficiently deliver functional nucleic acids into, for example, tissue culture cells. Another commercially available cationic lipid, 1,2-bis(oleoyloxy)-3,3-(trimethylammonia)propane ("DOTAP") (Boehringer Mannheim, Indianapolis, Ind.) differs from DOTMA in that the oleoyl moieties are linked by ester, rather than ether linkages.
[0316] Other reported cationic lipid compounds include those that have been conjugated to a variety of moieties including, for example, carboxyspermine which has been conjugated to one of two types of lipids and includes compounds such as 5-carboxyspermylglycine dioctaoleoylamide ("DOGS") (Transfectam.TM., Promega, Madison, Wis.) and dipalmitoylphosphatidylethanolamine 5-carboxyspermyl-amide ("DPPES") (see, e.g., U.S. Pat. No. 5,171,678).
[0317] Another cationic lipid conjugate includes derivatization of the lipid with cholesterol ("DC-Chol") which has been formulated into liposomes in combination with DOPE (See, Gao, X. and Huang, L., Biochim. Biophys. Res. Commun. 179:280, 1991). Lipopolylysine, made by conjugating polylysine to DOPE, has been reported to be effective for transfection in the presence of serum (Zhou, X. et al., Biochim. Biophys. Acta 1065:8, 1991). For certain cell lines, these liposomes containing conjugated cationic lipids, are said to exhibit lower toxicity and provide more efficient transfection than the DOTMA-containing compositions. Other commercially available cationic lipid products include DMRIE and DMRIE-HP (Vical, La Jolla, Calif.) and Lipofectamine (DOSPA) (Life Technology, Inc., Gaithersburg, Md.). Other cationic lipids suitable for the delivery of oligonucleotides are described in WO 98/39359 and WO 96/37194.
[0318] Liposomal formulations are particularly suited for topical administration, liposomes present several advantages over other formulations. Such advantages include reduced side effects related to high systemic absorption of the administered drug, increased accumulation of the administered drug at the desired target, and the ability to administer RNAi agent into the skin. In some implementations, liposomes are used for delivering RNAi agent to epidermal cells and also to enhance the penetration of RNAi agent into dermal tissues, e.g., into skin. For example, the liposomes can be applied topically. Topical delivery of drugs formulated as liposomes to the skin has been documented (see, e.g., Weiner et al., Journal of Drug Targeting, 1992, vol. 2,405-410 and du Plessis et al., Antiviral Research, 18, 1992, 259-265; Mannino, R. J. and Fould-Fogerite, S., Biotechniques 6:682-690, 1988; Itani, T. et al. Gene 56:267-276. 1987; Nicolau, C. et al. Meth. Enz. 149:157-176, 1987; Straubinger, R. M. and Papahadjopoulos, D. Meth. Enz. 101:512-527, 1983; Wang, C. Y. and Huang, L., Proc. Natl. Acad. Sci. USA 84:7851-7855, 1987).
[0319] Non-ionic liposomal systems have also been examined to determine their utility in the delivery of drugs to the skin, in particular systems comprising non-ionic surfactant and cholesterol. Non-ionic liposomal formulations comprising Novasome I (glyceryl dilaurate/cholesterol/polyoxyethylene-10-stearyl ether) and Novasome II (glyceryl distearate/cholesterol/polyoxyethylene-10-stearyl ether) were used to deliver a drug into the dermis of mouse skin. Such formulations with RNAi agent are useful for treating a dermatological disorder.
[0320] Liposomes that include RNAi agent can be made highly deformable. Such deformability can enable the liposomes to penetrate through pore that are smaller than the average radius of the liposome. For example, transfersomes are a type of deformable liposomes. Transferosomes can be made by adding surface edge activators, usually surfactants, to a standard liposomal composition. Transfersomes that include RNAi agent can be delivered, for example, subcutaneously by infection in order to deliver RNAi agent to keratinocytes in the skin. In order to cross intact mammalian skin, lipid vesicles must pass through a series of fine pores, each with a diameter less than 50 nm, under the influence of a suitable transdermal gradient. In addition, due to the lipid properties, these transferosomes can be self-optimizing (adaptive to the shape of pores, e.g., in the skin), self-repairing, and can frequently reach their targets without fragmenting, and often self-loading.
[0321] Other formulations amenable to the present invention are described in U.S. provisional application Ser. No. 61/018,616, filed Jan. 2, 2008; 61/018,611, filed Jan. 2, 2008; 61/039,748, filed Mar. 26, 2008; 61/047,087, filed Apr. 22, 2008 and 61/051,528, filed May 8, 2008. PCT application no PCT/US2007/080331, filed Oct. 3, 2007 also describes formulations that are amenable to the present invention.
[0322] Surfactants
[0323] Surfactants find wide application in formulations such as emulsions (including microemulsions) and liposomes (see above). RNAi agent (or a precursor, e.g., a larger dsiRNA which can be processed into a siRNA, or a DNA which encodes a siRNA or precursor) compositions can include a surfactant. In one embodiment, the siRNA is formulated as an emulsion that includes a surfactant. The most common way of classifying and ranking the properties of the many different types of surfactants, both natural and synthetic, is by the use of the hydrophile/lipophile balance (HLB). The nature of the hydrophilic group provides the most useful means for categorizing the different surfactants used in formulations (Rieger, in "Pharmaceutical Dosage Forms," Marcel Dekker, Inc., New York, N.Y., 1988, p. 285).
[0324] If the surfactant molecule is not ionized, it is classified as a nonionic surfactant. Nonionic surfactants find wide application in pharmaceutical products and are usable over a wide range of pH values. In general their HLB values range from 2 to about 18 depending on their structure. Nonionic surfactants include nonionic esters such as ethylene glycol esters, propylene glycol esters, glyceryl esters, polyglyceryl esters, sorbitan esters, sucrose esters, and ethoxylated esters. Nonionic alkanolamides and ethers such as fatty alcohol ethoxylates, propoxylated alcohols, and ethoxylated/propoxylated block polymers are also included in this class. The polyoxyethylene surfactants are the most popular members of the nonionic surfactant class.
[0325] If the surfactant molecule carries a negative charge when it is dissolved or dispersed in water, the surfactant is classified as anionic. Anionic surfactants include carboxylates such as soaps, acyl lactylates, acyl amides of amino acids, esters of sulfuric acid such as alkyl sulfates and ethoxylated alkyl sulfates, sulfonates such as alkyl benzene sulfonates, acyl isethionates, acyl taurates and sulfosuccinates, and phosphates. The most important members of the anionic surfactant class are the alkyl sulfates and the soaps.
[0326] If the surfactant molecule carries a positive charge when it is dissolved or dispersed in water, the surfactant is classified as cationic. Cationic surfactants include quaternary ammonium salts and ethoxylated amines. The quaternary ammonium salts are the most used members of this class.
[0327] If the surfactant molecule has the ability to carry either a positive or negative charge, the surfactant is classified as amphoteric. Amphoteric surfactants include acrylic acid derivatives, substituted alkylamides, N-alkylbetaines and phosphatides.
[0328] The use of surfactants in drug products, formulations and in emulsions has been reviewed (Rieger, in "Pharmaceutical Dosage Forms," Marcel Dekker, Inc., New York, N.Y., 1988, p. 285).
[0329] Micelles and other Membranous Formulations
[0330] The RNAi agents of the invention can also be provided as micellar formulations. "Micelles" are defined herein as a particular type of molecular assembly in which amphipathic molecules are arranged in a spherical structure such that all the hydrophobic portions of the molecules are directed inward, leaving the hydrophilic portions in contact with the surrounding aqueous phase. The converse arrangement exists if the environment is hydrophobic.
[0331] A mixed micellar formulation suitable for delivery through transdermal membranes may be prepared by mixing an aqueous solution of the siRNA composition, an alkali metal C.sub.8 to C.sub.22 alkyl sulphate, and a micelle forming compound. Exemplary micelle forming compounds include lecithin, hyaluronic acid, pharmaceutically acceptable salts of hyaluronic acid, glycolic acid, lactic acid, chamomile extract, cucumber extract, oleic acid, linoleic acid, linolenic acid, monoolein, monooleates, monolaurates, borage oil, evening of primrose oil, menthol, trihydroxy oxo cholanyl glycine and pharmaceutically acceptable salts thereof, glycerin, polyglycerin, lysine, polylysine, triolein, polyoxyethylene ethers and analogues thereof, polidocanol alkyl ethers and analogues thereof, chenodeoxycholate, deoxycholate, and mixtures thereof. The micelle forming compounds may be added at the same time or after addition of the alkali metal alkyl sulphate. Mixed micelles will form with substantially any kind of mixing of the ingredients but vigorous mixing in order to provide smaller size micelles.
[0332] In one method a first micellar composition is prepared which contains the siRNA composition and at least the alkali metal alkyl sulphate. The first micellar composition is then mixed with at least three micelle forming compounds to form a mixed micellar composition. In another method, the micellar composition is prepared by mixing the siRNA composition, the alkali metal alkyl sulphate and at least one of the micelle forming compounds, followed by addition of the remaining micelle forming compounds, with vigorous mixing.
[0333] Phenol and/or m-cresol may be added to the mixed micellar composition to stabilize the formulation and protect against bacterial growth. Alternatively, phenol and/or m-cresol may be added with the micelle forming ingredients. An isotonic agent such as glycerin may also be added after formation of the mixed micellar composition.
[0334] For delivery of the micellar formulation as a spray, the formulation can be put into an aerosol dispenser and the dispenser is charged with a propellant. The propellant, which is under pressure, is in liquid form in the dispenser. The ratios of the ingredients are adjusted so that the aqueous and propellant phases become one, i.e., there is one phase. If there are two phases, it is necessary to shake the dispenser prior to dispensing a portion of the contents, e.g., through a metered valve. The dispensed dose of pharmaceutical agent is propelled from the metered valve in a fine spray.
[0335] Propellants may include hydrogen-containing chlorofluorocarbons, hydrogen-containing fluorocarbons, dimethyl ether and diethyl ether. In certain embodiments, HFA 134a (1,1,1,2 tetrafluoroethane) may be used.
[0336] The specific concentrations of the essential ingredients can be determined by relatively straightforward experimentation. For absorption through the oral cavities, it is often desirable to increase, e.g., at least double or triple, the dosage for through injection or administration through the gastrointestinal tract.
[0337] Particles
[0338] In another embodiment, an RNAi agent of the invention may be incorporated into a particle, e.g., a microparticle. Microparticles can be produced by spray-drying, but may also be produced by other methods including lyophilization, evaporation, fluid bed drying, vacuum drying, or a combination of these techniques.
IV. Methods for Inhibiting TTR Expression
[0339] The present invention also provides methods of inhibiting expression of a transthyretin (TTR) in a cell. The methods include contacting a cell with an RNAi agent, e.g., double stranded RNAi agent, in an amount effective to inhibit expression of TTR in the cell, thereby inhibiting expression of TTR in the cell.
[0340] Contacting of a cell with an RNAi agent, e.g., a double stranded RNAi agent, may be done in vitro or in vivo. Contacting a cell in vivo with the RNAi agent includes contacting a cell or group of cells within a subject, e.g., a human subject, with the RNAi agent. Combinations of in vitro and in vivo methods of contacting a cell are also possible. Contacting a cell may be direct or indirect, as discussed above. Furthermore, contacting a cell may be accomplished via a targeting ligand, including any ligand described herein or known in the art. In preferred embodiments, the targeting ligand is a carbohydrate moiety, e.g., a GalNAc.sub.3 ligand, or any other ligand that directs the RNAi agent to a site of interest, e.g., the liver of a subject.
[0341] The term "inhibiting," as used herein, is used interchangeably with "reducing," "silencing," "downregulating", "suppressing", and other similar terms, and includes any level of inhibition.
[0342] The phrase "inhibiting expression of a TTR" is intended to refer to inhibition of expression of any TTR gene (such as, e.g., a mouse TTR gene, a rat TTR gene, a monkey TTR gene, or a human TTR gene) as well as variants or mutants of a TTR gene. Thus, the TTR gene may be a wild-type TTR gene, a mutant TTR gene (such as a mutant TTR gene giving rise to amyloid deposition), or a transgenic TTR gene in the context of a genetically manipulated cell, group of cells, or organism.
[0343] "Inhibiting expression of a TTR gene" includes any level of inhibition of a TTR gene, e.g., at least partial suppression of the expression of a TTR gene. The expression of the TTR gene may be assessed based on the level, or the change in the level, of any variable associated with TTR gene expression, e.g., TTR mRNA level, TTR protein level, or the number or extent of amyloid deposits. This level may be assessed in an individual cell or in a group of cells, including, for example, a sample derived from a subject.
[0344] Inhibition may be assessed by a decrease in an absolute or relative level of one or more variables that are associated with TTR expression compared with a control level. The control level may be any type of control level that is utilized in the art, e.g., a pre-dose baseline level, or a level determined from a similar subject, cell, or sample that is untreated or treated with a control (such as, e.g., buffer only control or inactive agent control).
[0345] In some embodiments of the methods of the invention, expression of a TTR gene is inhibited by at least about 5%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 91%, at least about 92%, at least about 93%, at least about 94%. at least about 95%, at least about 96%, at least about 97%, at least about 98%, or at least about 99%.
[0346] Inhibition of the expression of a TTR gene may be manifested by a reduction of the amount of mRNA expressed by a first cell or group of cells (such cells may be present, for example, in a sample derived from a subject) in which a TTR gene is transcribed and which has or have been treated (e.g., by contacting the cell or cells with an RNAi agent of the invention, or by administering an RNAi agent of the invention to a subject in which the cells are or were present) such that the expression of a TTR gene is inhibited, as compared to a second cell or group of cells substantially identical to the first cell or group of cells but which has not or have not been so treated (control cell(s)). In preferred embodiments, the inhibition is assessed by expressing the level of mRNA in treated cells as a percentage of the level of mRNA in control cells, using the following formula:
( mRNA in control cells ) - ( mRNA in treated cells ) ( mRNA in control cells ) .cndot.100 % ##EQU00001##
[0347] Alternatively, inhibition of the expression of a TTR gene may be assessed in terms of a reduction of a parameter that is functionally linked to TTR gene expression, e.g., TTR protein expression, retinol binding protein level, vitamin A level, or presence of amyloid deposits comprising TTR. TTR gene silencing may be determined in any cell expressing TTR, either constitutively or by genomic engineering, and by any assay known in the art. The liver is the major site of TTR expression. Other significant sites of expression include the choroid plexus, retina and pancreas.
[0348] Inhibition of the expression of a TTR protein may be manifested by a reduction in the level of the TTR protein that is expressed by a cell or group of cells (e.g., the level of protein expressed in a sample derived from a subject). As explained above for the assessment of mRNA suppression, the inhibition of protein expression levels in a treated cell or group of cells may similarly be expressed as a percentage of the level of protein in a control cell or group of cells.
[0349] A control cell or group of cells that may be used to assess the inhibition of the expression of a TTR gene includes a cell or group of cells that has not yet been contacted with an RNAi agent of the invention. For example, the control cell or group of cells may be derived from an individual subject (e.g., a human or animal subject) prior to treatment of the subject with an RNAi agent.
[0350] The level of TTR mRNA that is expressed by a cell or group of cells, or the level of circulating TTR mRNA, may be determined using any method known in the art for assessing mRNA expression. In one embodiment, the level of expression of TTR in a sample is determined by detecting a transcribed polynucleotide, or portion thereof, e.g., mRNA of the TTR gene. RNA may be extracted from cells using RNA extraction techniques including, for example, using acid phenol/guanidine isothiocyanate extraction (RNAzol B; Biogenesis), RNeasy RNA preparation kits (Qiagen) or PAXgene (PreAnalytix, Switzerland). Typical assay formats utilizing ribonucleic acid hybridization include nuclear run-on assays, RT-PCR, RNase protection assays (Melton et al., Nuc. Acids Res. 12:7035), Northern blotting, in situ hybridization, and microarray analysis. Circulating TTR mRNA may be detected using methods the described in PCT/US2012/043584, the entire contents of which are hereby incorporated herein by reference.
[0351] In one embodiment, the level of expression of TTR is determined using a nucleic acid probe. The term "probe", as used herein, refers to any molecule that is capable of selectively binding to a specific TTR. Probes can be synthesized by one of skill in the art, or derived from appropriate biological preparations. Probes may be specifically designed to be labeled. Examples of molecules that can be utilized as probes include, but are not limited to, RNA, DNA, proteins, antibodies, and organic molecules.
[0352] Isolated mRNA can be used in hybridization or amplification assays that include, but are not limited to, Southern or Northern analyses, polymerase chain reaction (PCR) analyses and probe arrays. One method for the determination of mRNA levels involves contacting the isolated mRNA with a nucleic acid molecule (probe) that can hybridize to TTR mRNA. In one embodiment, the mRNA is immobilized on a solid surface and contacted with a probe, for example by running the isolated mRNA on an agarose gel and transferring the mRNA from the gel to a membrane, such as nitrocellulose. In an alternative embodiment, the probe(s) are immobilized on a solid surface and the mRNA is contacted with the probe(s), for example, in an Affymetrix gene chip array. A skilled artisan can readily adapt known mRNA detection methods for use in determining the level of TTR mRNA.
[0353] An alternative method for determining the level of expression of TTR in a sample involves the process of nucleic acid amplification and/or reverse transcriptase (to prepare cDNA) of for example mRNA in the sample, e.g., by RT-PCR (the experimental embodiment set forth in Mullis, 1987, U.S. Pat. No. 4,683,202), ligase chain reaction (Barany (1991) Proc. Nat. Acad. Sci. USA 88:189-193), self sustained sequence replication (Guatelli et al. (1990) Proc. Nat. Acad. Sci. USA 87:1874-1878), transcriptional amplification system (Kwoh et al. (1989) Proc. Nat. Acad. Sci. USA 86:1173-1177), Q-Beta Replicase (Lizardi et al. (1988) Bio/Technology 6:1197), rolling circle replication (Lizardi et al., U.S. Pat. No. 5,854,033) or any other nucleic acid amplification method, followed by the detection of the amplified molecules using techniques well known to those of skill in the art. These detection schemes are especially useful for the detection of nucleic acid molecules if such molecules are present in very low numbers. In particular aspects of the invention, the level of expression of TTR is determined by quantitative fluorogenic RT-PCR (i.e., the TaqMan.TM. System).
[0354] The expression levels of TTR mRNA may be monitored using a membrane blot (such as used in hybridization analysis such as Northern, Southern, dot, and the like), or microwells, sample tubes, gels, beads or fibers (or any solid support comprising bound nucleic acids). See U.S. Pat. Nos. 5,770,722, 5,874,219, 5,744,305, 5,677,195 and 5,445,934, which are incorporated herein by reference. The determination of TTR expression level may also comprise using nucleic acid probes in solution.
[0355] In preferred embodiments, the level of mRNA expression is assessed using branched DNA (bDNA) assays or real time PCR (qPCR). The use of these methods is described and exemplified in the Examples presented herein.
[0356] The level of TTR protein expression may be determined using any method known in the art for the measurement of protein levels. Such methods include, for example, electrophoresis, capillary electrophoresis, high performance liquid chromatography (HPLC), thin layer chromatography (TLC), hyperdiffusion chromatography, fluid or gel precipitin reactions, absorption spectroscopy, a colorimetric assays, spectrophotometric assays, flow cytometry, immunodiffusion (single or double), immunoelectrophoresis, Western blotting, radioimmunoassay (RIA), enzyme-linked immunosorbent assays (ELISAs), immunofluorescent assays, electrochemiluminescence assays, and the like.
[0357] In some embodiments, the efficacy of the methods of the invention can be monitored by detecting or monitoring a reduction in an amyloid TTR deposit. Reducing an amyloid TTR deposit, as used herein, includes any decrease in the size, number, or severity of TTR deposits, or to a prevention or reduction in the formation of TTR deposits, within an organ or area of a subject, as may be assessed in vitro or in vivo using any method known in the art. For example, some methods of assessing amyloid deposits are described in Gertz, M. A. & Rajukumar, S. V. (Editors) (2010), Amyloidosis: Diagnosis and Treatment, New York: Humana Press. Methods of assessing amyloid deposits may include biochemical analyses, as well as visual or computerized assessment of amyloid deposits, as made visible, e.g., using immunohistochemical staining, fluorescent labeling, light microscopy, electron microscopy, fluorescence microscopy, or other types of microscopy. Invasive or noninvasive imaging modalities, including, e.g., CT, PET, or NMR/MRI imaging may be employed to assess amyloid deposits.
[0358] The methods of the invention may reduce TTR deposits in any number of tissues or regions of the body including but not limited to the heart, liver, spleen, esophagus, stomach, intestine (ileum, duodenum and colon), brain, sciatic nerve, dorsal root ganglion, kidney and retina.
[0359] The term "sample" as used herein refers to a collection of similar fluids, cells, or tissues isolated from a subject, as well as fluids, cells, or tissues present within a subject. Examples of biological fluids include blood, serum and serosal fluids, plasma, lymph, urine, cerebrospinal fluid, saliva, ocular fluids, and the like. Tissue samples may include samples from tissues, organs or localized regions. For example, samples may be derived from particular organs, parts of organs, or fluids or cells within those organism. In certain embodiments, samples may be derived from the liver (e.g., whole liver or certain segments of liver or certain types of cells in the liver, such as, e.g., hepatocytes), the retina or parts of the retina (e.g., retinal pigment epithelium), the central nervous system or parts of the central nervous system (e.g., ventricles or choroid plexus), or the pancreas or certain cells or parts of the pancreas. In preferred embodiments, a "sample derived from a subject" refers to blood or plasma drawn from the subject. In further embodiments, a "sample derived from a subject" refers to liver tissue or retinal tissue derived from the subject.
[0360] In some embodiments of the methods of the invention, the RNAi agent is administered to a subject such that the RNAi agent is delivered to a specific site within the subject. The inhibition of expression of TTR may be assessed using measurements of the level or change in the level of TTR mRNA or TTR protein in a sample derived from fluid or tissue from the specific site within the subject. In preferred embodiments, the site is selected from the group consisting of liver, choroid plexus, retina, and pancreas. The site may also be a subsection or subgroup of cells from any one of the aforementioned sites (e.g., hepatocytes or retinal pigment epithelium). The site may also include cells that express a particular type of receptor (e.g., hepatocytes that express the asialogycloprotein receptor).
V. Methods for Treating or Preventing a TTR-Associated Disease
[0361] The present invention also provides methods for treating or preventing a TTR-associated disease in a subject. The methods include administering to the subject a therapeutically effective amount or prophylactically effective amount of an RNAi agent of the invention.
[0362] As used herein, a "subject" includes either a human or a non-human animal, preferably a vertebrate, and more preferably a mammal. A subject may include a transgenic organism. Most preferably, the subject is a human, such as a human suffering from or predisposed to developing a TTR-associated disease.
[0363] In some embodiments, the subject is suffering from a TTR-associated disease. In other embodiments, the subject is a subject at risk for developing a TTR-associated disease, e.g., a subject with a TTR gene mutation that is associated with the development of a TTR associated disease, a subject with a family history of TTR-associated disease, or a subject who has signs or symptoms suggesting the development of TTR amyloidosis.
[0364] A "TTR-associated disease," as used herein, includes any disease caused by or associated with the formation of amyloid deposits in which the fibril precurosors consist of variant or wild-type TTR protein. Mutant and wild-type TTR give rise to various forms of amyloid deposition (amyloidosis). Amyloidosis involves the formation and aggregation of misfolded proteins, resulting in extracellular deposits that impair organ function. Clinical syndromes associated with TTR aggregation include, for example, senile systemic amyloidosis (SSA); systemic familial amyloidosis; familial amyloidotic polyneuropathy (FAP); familial amyloidotic cardiomyopathy (FAC); and leptomeningeal amyloidosis, also known as leptomeningeal or meningocerebrovascular amyloidosis, central nervous system (CNS) amyloidosis, or amyloidosis VII form.
[0365] In some embodiments of the methods of the invention, RNAi agents of the invention are administered to subjects suffering from familial amyloidotic cardiomyopathy (FAC) and senile systemic amyloidosis (SSA). Normal-sequence TTR causes cardiac amyloidosis in people who are elderly and is termed senile systemic amyloidosis (SSA) (also called senile cardiac amyloidosis (SCA) or cardiac amyloidosis). SSA often is accompanied by microscopic deposits in many other organs. TTR mutations accelerate the process of TTR amyloid formation and are the most important risk factor for the development of clinically significant TTR amyloidosis (also called ATTR (amyloidosis-transthyretin type)). More than 85 amyloidogenic TTR variants are known to cause systemic familial amyloidosis.
[0366] In some embodiments of the methods of the invention, RNAi agents of the invention are administered to subjects suffering from transthyretin (TTR)-related familial amyloidotic polyneuropathy (FAP). Such subjects may suffer from ocular manifestations, such as vitreous opacity and glaucoma. It is known to one of skill in the art that amyloidogenic transthyretin (ATTR) synthesized by retinal pigment epithelium (RPE) plays important roles in the progression of ocular amyloidosis. Previous studies have shown that panretinal laser photocoagulation, which reduced the RPE cells, prevented the progression of amyloid deposition in the vitreous, indicating that the effective suppression of ATTR expression in RPE may become a novel therapy for ocular amyloidosis (see, e.g., Kawaji, T., et al., Ophthalmology. (2010) 117: 552-555). The methods of the invention are useful for treatment of ocular manifestations of TTR related FAP, e.g., ocular amyloidosis. The RNAi agent can be delivered in a manner suitable for targeting a particular tissue, such as the eye. Modes of ocular delivery include retrobulbar, subcutaneous eyelid, subconjunctival, subtenon, anterior chamber or intravitreous injection (or internal injection or infusion). Specific formulations for ocular delivery include eye drops or ointments.
[0367] Another TTR-associated disease is hyperthyroxinemia, also known as "dystransthyretinemic hyperthyroxinemia" or "dysprealbuminemic hyperthyroxinemia". This type of hyperthyroxinemia may be secondary to an increased association of thyroxine with TTR due to a mutant TTR molecule with increased affinity for thyroxine. See, e.g., Moses et al. (1982) J. Cin. Invest., 86, 2025-2033.
[0368] The RNAi agents of the invention may be administered to a subject using any mode of administration known in the art, including, but not limited to subcutaneous, intravenous, intramuscular, intraocular, intrabronchial, intrapleural, intraperitoneal, intraarterial, lymphatic, cerebrospinal, and any combinations thereof. In preferred embodiments, the agents are administered subcutaneously.
[0369] In some embodiments, the administration is via a depot injection. A depot injection may release the RNAi agent in a consistent way over a prolonged time period. Thus, a depot injection may reduce the frequency of dosing needed to obtain a desired effect, e.g., a desired inhibition of TTR, or a therapeutic or prophylactic effect. A depot injection may also provide more consistent serum concentrations. Depot injections may include subcutaneous injections or intramuscular injections. In preferred embodiments, the depot injection is a subcutaneous injection.
[0370] In some embodiments, the administration is via a pump. The pump may be an external pump or a surgically implanted pump. In certain embodiments, the pump is a subcutaneously implanted osmotic pump. In other embodiments, the pump is an infusion pump. An infusion pump may be used for intravenous, subcutaneous, arterial, or epidural infusions. In preferred embodiments, the infusion pump is a subcutaneous infusion pump. In other embodiments, the pump is a surgically implanted pump that delivers the RNAi agent to the liver.
[0371] Other modes of administration include epidural, intracerebral, intracerebroventricular, nasal administration, intraarterial, intracardiac, intraosseous infusion, intrathecal, and intravitreal, and pulmonary. The mode of administration may be chosen based upon whether local or systemic treatment is desired and based upon the area to be treated. The route and site of administration may be chosen to enhance targeting. In some embodiments, the RNAi agent is administered to a subject in an amount effective to inhibit TTR expression in a cell within the subject. The amount effective to inhibit TTR expression in a cell within a subject may be assessed using methods discussed above, including methods that involve assessment of the inhibition of TTR mRNA, TTR protein, or related variables, such as amyloid deposits.
[0372] In some embodiments, the RNAi agent is administered to a subject in a therapeutically or prophylactically effective amount.
[0373] "Therapeutically effective amount," as used herein, is intended to include the amount of an RNAi agent that, when administered to a patient for treating a TTR associated disease, is sufficient to effect treatment of the disease (e.g., by diminishing, ameliorating or maintaining the existing disease or one or more symptoms of disease). The "therapeutically effective amount" may vary depending on the RNAi agent, how the agent is administered, the disease and its severity and the history, age, weight, family history, genetic makeup, stage of pathological processes mediated by TTR expression, the types of preceding or concomitant treatments, if any, and other individual characteristics of the patient to be treated.
[0374] "Prophylactically effective amount," as used herein, is intended to include the amount of an RNAi agent that, when administered to a subject who does not yet experience or display symptoms of a TTR-associated disease, but who may be predisposed to the disease, is sufficient to prevent or ameliorate the disease or one or more symptoms of the disease. Symptoms that may be ameliorated include sensory neuropathy (e.g., paresthesia, hypesthesia in distal limbs), autonomic neuropathy (e.g., gastrointestinal dysfunction, such as gastric ulcer, or orthostatic hypotension), motor neuropathy, seizures, dementia, myelopathy, polyneuropathy, carpal tunnel syndrome, autonomic insufficiency, cardiomyopathy, vitreous opacities, renal insufficiency, nephropathy, substantially reduced mBMI (modified Body Mass Index), cranial nerve dysfunction, and corneal lattice dystrophy. Ameliorating the disease includes slowing the course of the disease or reducing the severity of later-developing disease. The "prophylactically effective amount" may vary depending on the RNAi agent, how the agent is administered, the degree of risk of disease, and the history, age, weight, family history, genetic makeup, the types of preceding or concomitant treatments, if any, and other individual characteristics of the patient to be treated.
[0375] A "therapeutically-effective amount" or "prophylacticaly effective amount" also includes an amount of an RNAi agent that produces some desired local or systemic effect at a reasonable benefit/risk ratio applicable to any treatment. RNAi agents employed in the methods of the present invention may be administered in a sufficient amount to produce a reasonable benefit/risk ratio applicable to such treatment.
[0376] As used herein, the phrases "therapeutically effective amount" and "prophylactically effective amount" also include an amount that provides a benefit in the treatment, prevention, or management of pathological processes or symptom(s) of pathological processes mediated by TTR expression. Symptoms of TTR amyloidosis include sensory neuropathy (e.g. paresthesia, hypesthesia in distal limbs), autonomic neuropathy (e.g., gastrointestinal dysfunction, such as gastric ulcer, or orthostatic hypotension), motor neuropathy, seizures, dementia, myelopathy, polyneuropathy, carpal tunnel syndrome, autonomic insufficiency, cardiomyopathy, vitreous opacities, renal insufficiency, nephropathy, substantially reduced mBMI (modified Body Mass Index), cranial nerve dysfunction, and corneal lattice dystrophy.
[0377] The dose of an RNAi agent that is administered to a subject may be tailored to balance the risks and benefits of a particular dose, for example, to achieve a desired level of TTR gene suppression (as assessed, e.g., based on TTR mRNA suppression, TTR protein expression, or a reduction in an amyloid deposit, as defined above) or a desired therapeutic or prophylactic effect, while at the same time avoiding undesirable side effects.
[0378] In one embodiment, the RNAi agent is administered at a dose of between about 0.25 mg/kg to about 50 mg/kg, e.g., between about 0.25 mg/kg to about 0.5 mg/kg, between about 0.25 mg/kg to about 1 mg/kg, between about 0.25 mg/kg to about 5 mg/kg, between about 0.25 mg/kg to about 10 mg/kg, between about 1 mg/kg to about 10 mg/kg, between about 5 mg/kg to about 15 mg/kg, between about 10 mg/kg to about 20 mg/kg, between about 15 mg/kg to about 25 mg/kg, between about 20 mg/kg to about 30 mg/kg, between about 25 mg/kg to about 35 mg/kg, or between about 40 mg/kg to about 50 mg/kg.
[0379] In some embodiments, the RNAi agent is administered at a dose of about 0.25 mg/kg, about 0.5 mg/kg, about 1 mg/kg, about 2 mg/kg, about 3 mg/kg, about 4 mg/kg, about 5 mg/kg, about 6 mg/kg, about 7 mg/kg, about 8 mg/kg, about 9 mg/kg, about 10 mg/kg, about 11 mg/kg, about 12 mg/kg, about 13 mg/kg, about 14 mg/kg, about 15 mg/kg, about 16 mg/kg, about 17 mg/kg, about 18 mg/kg, about 19 mg/kg, about 20 mg/kg, about 21 mg/kg, about 22 mg/kg, about 23 mg/kg, about 24 mg/kg, about 25 mg/kg, about 26 mg/kg, about 27 mg/kg, about 28 mg/kg, about 29 mg/kg, 30 mg/kg, about 31 mg/kg, about 32 mg/kg, about 33 mg/kg, about 34 mg/kg, about 35 mg/kg, about 36 mg/kg, about 37 mg/kg, about 38 mg/kg, about 39 mg/kg, about 40 mg/kg, about 41 mg/kg, about 42 mg/kg, about 43 mg/kg, about 44 mg/kg, about 45 mg/kg, about 46 mg/kg, about 47 mg/kg, about 48 mg/kg, about 49 mg/kg or about 50 mg/kg.
[0380] In some embodiments, the RNAi agent is administered in two or more doses. If desired to facilitate repeated or frequent infusions, implantation of a delivery device, e.g., a pump, semi-permanent stent (e.g., intravenous, intraperitoneal, intracisternal or intracapsular), or reservoir may be advisable. In some embodiments, the number or amount of subsequent doses is dependent on the achievement of a desired effect, e.g., the suppression of a TTR gene, or the achievement of a therapeutic or prophylactic effect, e.g., reducing an amyloid deposit or reducing a symptom of a TTR-associated disease. In some embodiments, the RNAi agent is administered according to a schedule. For example, the RNAi agent may be administered twice per week, three times per week, four times per week, or five times per week. In some embodiments, the schedule involves regularly spaced administrations, e.g., hourly, every four hours, every six hours, every eight hours, every twelve hours, daily, every 2 days, every 3 days, every 4 days, every 5 days, weekly, biweekly, or monthly. In other embodiments, the schedule involves closely spaced administrations followed by a longer period of time during which the agent is not administered. For example, the schedule may involve an initial set of doses that are administered in a relatively short period of time (e.g., about every 6 hours, about every 12 hours, about every 24 hours, about every 48 hours, or about every 72 hours) followed by a longer time period (e.g., about 1 week, about 2 weeks, about 3 weeks, about 4 weeks, about 5 weeks, about 6 weeks, about 7 weeks, or about 8 weeks) during which the RNAi agent is not administered. In one embodiment, the RNAi agent is initially administered hourly and is later administered at a longer interval (e.g., daily, weekly, biweekly, or monthly). In another embodiment, the RNAi agent is initially administered daily and is later administered at a longer interval (e.g., weekly, biweekly, or monthly). In certain embodiments, the longer interval increases over time or is determined based on the achievement of a desired effect. In a specific embodiment, the RNAi agent is administered once daily during a first week, followed by weekly dosing starting on the eighth day of administration. In another specific embodiment, the RNAi agent is administered every other day during a first week followed by weekly dosing starting on the eighth day of administration.
[0381] Any of these schedules may optionally be repeated for one or more iterations. The number of iterations may depend on the achievement of a desired effect, e.g., the suppression of a TTR gene, retinol binding protein level, vitamin A level, and/or the achievement of a therapeutic or prophylactic effect, e.g., reducing an amyloid deposit or reducing a symptom of a TTR-associated disease.
[0382] In some embodiments, the RNAi agent is administered with other therapeutic agents or other therapeutic regimens. For example, other agents or other therapeutic regimens suitable for treating a TTR-associated disease may include a liver transplant, which can reduce mutant TTR levels in the body; Tafamidis (Vyndagel), which kinetically stabilizes the TTR tetramer preventing tetramer dissociation required for TTR amyloidogenesis; and diuretics, which may be employed, for example, to reduce edema in TTR amyloidosis with cardiac involvement.
[0383] In one embodiment, a subject is administered an initial dose and one or more maintenance doses of an RNAi agent. The maintenance dose or doses can be the same or lower than the initial dose, e.g., one-half of the initial dose. A maintenance regimen can include treating the subject with a dose or doses ranging from 0.01 g to 15 mg/kg of body weight per day, e.g., 10 mg/kg, 1 mg/kg, 0.1 mg/kg, 0.01 mg/kg, 0.001 mg/kg, or 0.00001 mg/kg of bodyweight per day. The maintenance doses are, for example, administered no more than once every 2 days, once every 5 days, once every 7 days, once every 10 days, once every 14 days, once every 21 days, or once every 30 days. Further, the treatment regimen may last for a period of time which will vary depending upon the nature of the particular disease, its severity and the overall condition of the patient. In certain embodiments the dosage may be delivered no more than once per day, e.g., no more than once per 24, 36, 48, or more hours, e.g., no more than once every 5 or 8 days. Following treatment, the patient can be monitored for changes in his/her condition. The dosage of the RNAi agent may either be increased in the event the patient does not respond significantly to current dosage levels, or the dose may be decreased if an alleviation of the symptoms of the disease state is observed, if the disease state has been ablated, or if undesired side-effects are observed.
VI. Kits
[0384] The present invention also provides kits for performing any of the methods of the invention. Such kits include one or more RNAi agent(s) and instructions for use, e.g., instructions for inhibiting expression of a TTR in a cell by contacting the cell with the RNAi agent(s) in an amount effective to inhibit expression of the TTR. The kits may optionally further comprise means for contacting the cell with the RNAi agent (e.g., an injection device), or means for measuring the inhibition of TTR (e.g., means for measuring the inhibition of TTR mRNA or TTR protein). Such means for measuring the inhibition of TTR may comprise a means for obtaining a sample from a subject, such as, e.g., a plasma sample. The kits of the invention may optionally further comprise means for administering the RNAi agent(s) to a subject or means for determining the therapeutically effective or prophylactically effective amount.
[0385] This invention is further illustrated by the following examples which should not be construed as limiting. The contents of all references and published patents and patent applications cited throughout the application are hereby incorporated herein by reference.
Examples
Example 1: Inhibition of TTR with TTR-GalNAc Conjugates
[0386] A single dose of the TTR RNAi agent AD-43527 was administered to mice subcutaneously and TTR mRNA levels were determined 72 hours post administration.
[0387] The mouse/rat cross-reactive GalNAc-conjugate, AD-43527, was chosen for in vivo evaluation in WT C57BL/6 mice for silencing of TTR mRNA in liver. The sequence of each strand of AD-43527 is shown below.
[0388] Strand: s=sense; as=antisense
TABLE-US-00002 Oligo Duplex # Strand # Sequence 5' to 3' AD-43527 s A- AfaCfaGfuGfuUfcUfuGfcUfcUfaUfaAfL96 (SEQ ID 89592 NO: 8) as A- uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu (SEQ 83989 ID NO: 9 L96 = GalNAc3; lowercase nts (a, u, g, c) are 2'-O-methyl nucleotides, Nf (i.e., Af) is a 2'-fluoro nucleotide
The ligand used was GalNAc.sub.3:
##STR00011##
This GalNAc3 ligand was conjugated to the 3'-end of the sense strand using the linker and tether as shown below:
##STR00012##
The structure of the resulting GalNAc.sub.3 conjugated sense strand is shown in the following schematic:
##STR00013##
Additional RNAi agents that target TTR and have the following sequences and modifications were synthesized and assayed.
Mouse/Rat Cross Reactive TTR RNAi Agents
TABLE-US-00003 [0389] Duplex Sense strand 5'-3' Antisense strand 5'-3' AD- AfaCfaGfuGfuUfcUfuGfcUfcUfaUfaAfQ11L96 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 43528 (SEQ ID NO: 10) (SEQ ID NO: 11)
Human/cyno cross reactive TTR RNAi agents; parent duplex is AD-18328 [having a sense strand 5'-3' sequence of GuAAccAAGAGuAuuccAudTdT (SEQ ID NO: 12) and antisense strand 5' to 3' sequence of AUGGAAuACUCUUGGUuACdTdT (SEQ ID NO: 13) with the following modifications: alternating 2'F/2'OMe w/2 PS on AS.
TABLE-US-00004 Duplex Sense strand 5'-3' Antisense strand 5'-3' AD- AfuGfuAfaCfcAfaGfaGfuAfuUfcCfaUfL96 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 45163 (SEQ ID NO: 14) (SEQ ID NO: 16) AD- AfuGfuAfaCfcAfaGfaGfuAfuUfcCfaUfQ11L96 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 45164 (SEQ ID NO: 15) (SEQ ID NO: 17)
L96=GalNAc.sub.3; lowercase nts (a,u,g,c) are 2'-O-methyl nucleotides, Nf (i.e., Af) is a 2'-fluoro nucleotide; Q11 is cholesterol; s is phosphorothioate.
[0390] AD-43527 was administered to female C57BL/6 mice (6-10 weeks, 5 per group) via subcutaneous injection at a dose volume of 10 .mu.l/g at a dose of 30, 15, 7.5, 3.5, 1.75 or 0.5 mg/kg of AD-43527. Control animals received PBS by subcutaneous injection at the same dose volume.
[0391] After approximately seventy two hours, mice were anesthetized with 200 .mu.l of ketamine, and then exsanguinated by severing the right caudal artery. Liver tissue was collected, flash-frozen and stored at -80.degree. C. until processing.
[0392] Efficacy of treatment was evaluated by measurement of TTR mRNA in the liver at 72 hours post-dose. TTR liver mRNA levels were assayed utilizing the Branched DNA assays--QuantiGene 1.0 (Panomics). Briefly, mouse liver samples were ground and tissue lysates were prepared. Liver lysis mixture (a mixture of 1 volume of lysis mixture, 2 volume of nuclease-free water and 10 .mu.l of Proteinase-K/ml for a final concentration of 20 mg/ml) was incubated at 65.degree. C. for 35 minutes. 5 .mu.l of liver lysate and 95 .mu.l of working probe set (TTR probe for gene target and GAPDH for endogenous control) were added into the Capture Plate. Capture Plates were incubated at 53.degree. C. 1.degree. C. (aprx. 16-20 hrs). The next day, the Capture Plates were washed 3 times with 1.times. Wash Buffer (nuclease-free water, Buffer Component 1 and Wash Buffer Component 2), then dried by centrifuging for 1 minute at 240 g. 100 .mu.l of Amplifier Probe mix per well was added into the Capture Plate, which was sealed with aluminum foil and incubated for 1 hour at 46.degree. C. 1.degree. C. Following a 1 hour incubation, the wash step was repeated, then 100p of Label Probe mix per well was added. Capture plates were incubated at 46.degree. C. 1.degree. C. for 1 hour. The plates were then washed with 1.times. Wash Buffer, dried and 100 .mu.l substrate per well was added into the Capture Plates. Capture Plates were incubated for 30 minutes at 46.degree. C. followed by incubation for 30 minutes at room temperature. Plates were read using the SpectraMax Luminometer following incubation. bDNA data were analyzed by subtracting the average background from each duplicate sample, averaging the resultant duplicate GAPDH (control probe) and TTR (experimental probe) values, and then computing the ratio: (experimental probe-background)/(control probe-background). The average TTR mRNA level was calculated for each group and normalized to the PBS group average to give relative TTR mRNA as a % of the PBS control group.
[0393] The results are shown in FIG. 1. The GalNAc conjugated RNAi agent targeting TTR had an ED.sub.50 of approximately 5 mg/kg for TTR mRNA knockdown. These results demonstrate that GalNAc conjugated RNAi agents that target TTR are effective at inhibiting expression of TTR mRNA.
Example 2: Inhibition of TTR with TTR-GalNAc Conjugates is Durable
[0394] Mice were administered a subcutaneous dose (either 7.5 or 30.0 mg/kg) of AD-43527, a GalNAc conjugated RNAi agent that targets TTR. The TTR mRNA levels in the liver were evaluated at 1, 3, 5, 7, 10, 13, 15, 19, 26, 33, and 41 days post treatment using the method described in Example 1.
[0395] The results are shown in FIG. 2. At day 19, administration of 30.0 mg/kg GalNAc conjugated RNAi agents still showed about 50% silencing. Full recovery of expression occurred at day 41.
[0396] These results demonstrated that the inhibition provided by GalNAc conjugated siRNA targeting TTR is durable, lasting up to 3, 5, 7, 10, 13, 15, 19, 26 or 33 days post treatment.
Example 3. RNA Synthesis and Duplex Annealing
1. Oligonucleotide Synthesis
[0397] Oligonucleotides were synthesized on an AKTAoligopilot synthesizer or an ABI 394 synthsizer. Commercially available controlled pore glass solid support (dT-CPG, 500 .ANG., Prime Synthesis) and RNA phosphoramidites with standard protecting groups, 5'-O-dimethoxytrityl N6-benzoyl-2'-t-butyldimethylsilyl-adenosine-3'-O--N,N'-diisopropyl-2-cya- noethylphosphoramidite, 5'-O-dimethoxytrityl-N4-acetyl-2'-t-butyldimethylsilyl-cytidine-3'-O--N,N- '-diisopropyl-2-cyanoethylphosphoramidite, 5'-O-dimethoxytrityl-N2-isobutryl-2'-t-butyldimethylsilyl-guanosine-3'-O-- -N,N'-diisopropyl-2-cyanoethylphosphoramidite, and 5'-O-dimethoxytrityl-2'-t-butyldimethylsilyl-uridine-3'-O--N,N'-diisoprop- yl-2-cyanoethylphosphoramidite (Pierce Nucleic Acids Technologies) were used for the oligonucleotide synthesis unless otherwise specified. The 2'-F phosphoramidites, 5'-O-dimethoxytrityl-N4-acetyl-2'-fluro-cytidine-3'-O--N,N'-diisopropyl-2- -cyanoethyl-phosphoramidite and 5'-O-dimethoxytrityl-2'-fluro-uridine-3'-O--N,N'-diisopropyl-2-cyanoethyl- -phosphoramidite were purchased from (Promega). All phosphoramidites were used at a concentration of 0.2M in acetonitrile (CH.sub.3CN) except for guanosine which was used at 0.2M concentration in 10% THF/ANC (v/v). Coupling/recycling time of 16 minutes was used. The activator was 5-ethyl thiotetrazole (0.75M, American International Chemicals), for the PO-oxidation Iodine/Water/Pyridine was used and the PS-oxidation PADS (2%) in 2,6-lutidine/ACN (1:1 v/v) was used.
[0398] Ligand conjugated strands were synthesized using a solid support containing the corresponding ligand. For example, the introduction of a carbohydrate moiety/ligand (for e.g., GalNAc) at the 3'-end of a sequence was achieved by starting the synthesis with the corresponding carbohydrate solid support. Similarly a cholesterol moiety at the 3'-end was introduced by starting the synthesis on the cholesterol support. In general, the ligand moiety was tethered to trans-4-hydroxyprolinol via a tether of choice as described in the previous examples to obtain a hydroxyprolinol-ligand moiety. The hydroxyprolinol-ligand moiety was then coupled to a solid support via a succinate linker or was converted to phosphoramidite via standard phosphitylation conditions to obtain the desired carbohydrate conjugate building blocks. Fluorophore labeled siRNAs were synthesized from the corresponding phosphoramidite or solid support, purchased from Biosearch Technologies. The oleyl lithocholic (GalNAc).sub.3 polymer support made in house at a loading of 38.6 .mu.mol/gram. The Mannose (Man).sub.3 polymer support was also made in house at a loading of 42.0 .mu.mol/gram.
[0399] Conjugation of the ligand of choice at the desired position, for example at the 5'-end of the sequence, was achieved by coupling of the corresponding phosphoramidite to the growing chain under standard phosphoramidite coupling conditions unless otherwise specified. An extended 15 minute coupling of 0.1M solution of phosphoramidite in anhydrous CH.sub.3CN in the presence of 5-(ethylthio)-1H-tetrazole activator to a solid bound oligonucleotide. Oxidation of the internucleotide phosphite to the phosphate was carried out using standard iodine-water as reported in Beaucage, S. L. (2008) Solid-phase synthesis of siRNA oligonucleotides. Curr. Opin. Drug Discov. Devel., 11, 203-216; Mueller, S., Wolf, J. and Ivanov, S. A. (2004) Current Strategies for the Synthesis of RNA. Curr. Org. Synth., 1, 293-307; Xia, J., Noronha, A., Toudjarska, I., Li, F., Akinc, A., Braich, R., Frank-Kamenetsky, M., Rajeev, K. G., Egli, M. and Manoharan, M. (2006) Gene Silencing Activity of siRNAs with a Ribo-difluorotoluyl Nucleotide. ACS Chem. Biol., 1, 176-183 or by treatment with tert-butyl hydroperoxide/acetonitrile/water (10:87:3) with a 10 minute oxidation wait time conjugated oligonucleotide. Phosphorothioate was introduced by the oxidation of phosphite to phosphorothioate by using a sulfur transfer reagent such as DDTT (purchased from AM Chemicals), PADS and or Beaucage reagent The cholesterol phosphoramidite was synthesized in house, and used at a concentration of 0.1 M in dichloromethane. Coupling time for the cholesterol phosphoramidite was 16 minutes.
2. Deprotection-I (Nucleobase Deprotection)
[0400] After completion of synthesis, the support was transferred to a 100 ml glass bottle (VWR). The oligonucleotide was cleaved from the support with simultaneous deprotection of base and phosphate groups with 80 mL of a mixture of ethanolic ammonia [ammonia:ethanol (3:1)] for 6.5 h at 55.degree. C. The bottle was cooled briefly on ice and then the ethanolic ammonia mixture was filtered into a new 250 ml bottle. The CPG was washed with 2.times.40 mL portions of ethanol/water (1:1 v/v). The volume of the mixture was then reduced to 30 ml by roto-vap. The mixture was then frozen on dry ice and dried under vacuum on a speed vac.
3. Deprotection-I (Removal of 2' TBDMS Group)
[0401] The dried residue was resuspended in 26 ml of triethylamine, triethylamine trihydrofluoride (TEA.3HF) or pyridine-HF and DMSO (3:4:6) and heated at 60.degree. C. for 90 minutes to remove the tert-butyldimethylsilyl (TBDMS) groups at the 2' position. The reaction was then quenched with 50 ml of 20 mM sodium acetate and pH adjusted to 6.5, and stored in freezer until purification.
4. Analysis
[0402] The oligonucleotides were analyzed by high-performance liquid chromatography (HPLC) prior to purification and selection of buffer and column depends on nature of the sequence and or conjugated ligand.
5. HPLC Purification
[0403] The ligand conjugated oligonucleotides were purified by reverse phase preparative HPLC. The unconjugated oligonucleotides were purified by anion-exchange HPLC on a TSK gel column packed in house. The buffers were 20 mM sodium phosphate (pH 8.5) in 10% CH.sub.3CN (buffer A) and 20 mM sodium phosphate (pH 8.5) in 10% CH.sub.3CN, 1M NaBr (buffer B). Fractions containing full-length oligonucleotides were pooled, desalted, and lyophilized. Approximately 0.15 OD of desalted oligonucleotidess were diluted in water to 150 .mu.l and then pipetted in special vials for CGE and LC/MS analysis. Compounds were finally analyzed by LC-ESMS and CGE.
6. RNAi Agent Preparation
[0404] For the preparation of an RNAi agent, equimolar amounts of sense and antisense strand were heated in 1.times.PBS at 95.degree. C. for 5 minutes and slowly cooled to room temperature. The integrity of the duplex was confirmed by HPLC analysis. Table 1 below reflects the RNAi agents which target human or rodent TTR mRNA.
TABLE-US-00005 TABLE 1 RNAi Agents and Results of In Vitro Screening % of mRNA remained SEQ SEQ conc. of siRNA Duplex ID ID 0.1 0.01 IC50 ID S ID NO: Sense strand (S) AS ID NO: Antisense strand (AS) 1 nM nM nM (nM) D1000 S1000 18 AfuGfuAfaCfcAfAfGfaGfuAfuUfcCfasu AS1000 1110 AfUfgGfaAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.03 0.1 0.47 0.006 D1001 S1001 19 AfsuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1001 1111 aUfsgGfAfAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.03 0.10 0.49 0.0065 D1002 S1002 20 AfuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1002 1112 aUfgGfAfAfuAfcUfcuuGfgsUfuAfcAfusGfsa 0.04 0.10 0.46 0.0068 D1003 S1003 21 AfuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1003 1113 aUfgGfAfAfuAfcUfcuuGfgUfsuAfcAfusGfsa 0.05 0.12 0.56 0.0073 D1004 S1004 22 aUGuaACccAGagUAuuCCasu AS1004 1114 AUggAAuaCUcuUGguUAcaUsGsa 0.07 0.13 0.44 0.008 D1005 S1005 23 AfuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1005 1115 aUfgGfAfAfuAfcUfcuuGfgsUfsuAfcAfusGfsa 0.06 0.11 0.53 0.0093 D1006 S1006 24 AfuGfuAfAfccAfAfGfaGfuAfuUfcCfasUf AS1006 1116 aUfgGfaAfuAfcUfcuuGfGfuuAfcAfusGfsa 0.05 0.16 0.55 0.0095 D1007 S1007 25 AfuGfuAfAfCfcAfAfGfaGfuAfuUfcCfasUf AS1007 1117 aUfgGfaAfuAfcUfcuuGfguuAfcAfusGfsa 0.05 0.14 0.48 0.0098 D1008 S1008 26 auguaaccaadGadGudAudAcdGasu AS1008 1118 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.07 0.11 0.33 0.010 D1009 S1009 27 UfgGfGfAfuUfuCfAfUfgUfaAfcCfAfAfgsAf AS1009 1119 uCfuugGfuUfaCfaugAfaAfuccCfasUfsc 0.03 0.14 0.56 0.0101 D1010 S1010 28 UfgGfgauUfuCfAfUfgUfaAfcCfaAfgsAf AS1010 1120 uCfuUfgGfuUfaCfaugAfaAfUfCfcCfasUfsc 0.03 0.14 0.65 0.0101 D1011 S1011 29 aUfGfuAfAfccAfAfGfaGfuAfuUfcCfasUf AS1011 1121 aUfgGfaAfuAfcUfcuuGfGfuuAfcaUfsgsa 0.06 0.10 0.55 0.011 D1012 S1012 30 UfgGfgAfuUfuCfAfUfgUfaacCfaAfgsAf AS1012 1122 uCfuUfgGfUfUfaCfaugAfaAfuCfcCfasUfsc 0.04 0.13 0.54 0.0114 D1013 S1013 31 auguaaccaadGadGudAudAcdGasu AS1013 1123 aUfgGfaAfuAfcUfcUfugdGudTadCadTsgsa 0.11 0.19 0.49 0.011 D1014 S1014 32 AfuGfuaaCfcAfAfGfaGfuAfuUfcCfasUf AS1014 1124 aUfgGfaAfuAfcUfcuuGfgUfUfAfcAfusGfsa 0.04 0.16 0.59 0.013 D1015 S1015 33 AfuguAfaccAfaGfdAGfdTAdTudCcdAsu AS1015 1125 dAUdGgdAadTAfdCUfcUfuGfgUfuAfcAfusGfsa 0.07 0.15 0.51 0.013 D1016 S1016 34 auGfuAfaCfcAfAfGfaGfuAfuUfcCfasUf AS1016 1126 aUfgGfaAfuAfcUfcuuGfgUfuAfcAfUfsGfsa 0.05 0.14 0.64 0.013 D1017 S1017 35 UfGfggAfuUfuCfAfUfgUfAfAfcCfaAfgsAf AS1017 1127 uCfuUfgGfuuaCfaugAfaAfuCfCfcasUfsc 0.09 0.41 0.74 0.0133 D1018 S1018 36 AfuguAfaCfcAfAfGfaGfuAfuUfcCfasUf AS1018 1128 aUfgGfaAfuAfcUfcuuGfgUfuAfCfAfusGfsa 0.03 0.14 0.61 0.014 D1019 S1019 37 AfuGfuAfaccAfAfGfaGfuAfuUfcCfasUf AS1019 1129 aUfgGfaAfuAfcUfcuuGfGfUfuAfcAfusGfsa 0.02 0.2 0.7 0.014 D1020 S1020 38 AfsuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1020 1130 asUfsgGfAfAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.04 0.16 0.67 0.0156 D1021 S1021 39 aUfguAfAfccAfAfgagUfaUfuCfcasUf AS1021 1131 aUfGfgAfaUfaCfUfCfuuGfGfuuAfCfaUfsgsa 0.11 0.24 0.64 0.016 D1022 S1022 40 dTdGggdAdTuudCdAugdTdAacdCdAagsdA AS1022 1132 udCdTugdGdTuadCdAugdAdAaudCdCcasdTsc 0.08 0.27 0.64 0.0161 D1023 S1023 41 AfsuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1023 1133 aUfgsGfAfAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.03 0.19 0.63 0.0163 D1024 S1024 42 UfgGfgAfuUfuCfAfUfguaAfcCfaAfgsAf AS1024 1134 uCfuUfgGfuUfAfCfaugAfaAfuCfcCfasUfsc 0.05 0.25 0.69 0.0164 D1025 S1025 43 UfgGfgAfuUfuCfAfUfgUfAfAfcCfaAfgsAf AS1025 1135 uCfuUfgGfuuaCfaugAfaAfuCfcCfasUfsc 0.04 0.18 0.75 0.0166 D1026 S1026 44 UfgGfgAfuUfuCfAfUfgUfaAfcCfaAfgsAf AS1026 1136 uCfuUfgGfuUfaCfaugAfaAfuCfcCfasUfsc 0.04 0.19 0.66 0.0178 D1027 S1027 45 UfgGfgAfuUfuCfAfUfgUfaAfccaAfgsAf AS1027 1137 uCfuUfGfGfuUfaCfaugAfaAfuCfcCfasUfsc 0.04 0.19 0.69 0.018 D1028 S1028 46 dAdTgudAdAccdAdAgadGdTaudTdCcasdT AS1028 1138 adTdGgadAdTacdTdCuudGdGuudAdCausdGsa 0.15 0.29 0.72 0.018 D1029 S1029 47 AdTGdTAdACdCAdAGdAGdTAdTUdCCdAsU AS1029 1139 dAUdGGdAAdTAdCUdCUdTGdGUdTAdCAdTsGsdA 0.1 0.27 0.61 0.018 D1030 S1030 48 UfgGfGfAfuuuCfAfUfgUfaAfcCfaAfgsAf AS1030 1140 uCfuUfgGfuUfaCfaugAfAfAfuccCfasUfsc 0.04 0.21 0.64 0.0187 D1031 S1031 49 AfuGfuAfAfccAfAfGfAfGfuAfuuccAfsu AS1031 1141 AfUfGfGfAfAfuAfCfUfCfUfuGfGfuuAfcAfusGfsa 0.06 0.15 0.62 0.019 D1032 S1032 50 AfsuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1032 1142 asUfgGfAfAfuAfcUfcuuGfgUfsuAfcAfusGfsa 0.09 0.34 0.78 0.021 D1033 S1033 51 UfgGfgAfuUfuCfaUfGfUfaacCfaAfgsAf AS1033 1143 uCfuUfgGfUfUfacaUfgAfaAfuCfcCfasUfsc 0.06 0.26 0.57 0.0212 D1034 S1034 52 AfuGfuAfAfccAfaGfaGfuAfuUfcCfasUf AS1034 1144 aUfgGfaAfuAfcUfcUfuGfGfuuAfcAfusGfsa 0.11 0.39 0.82 0.0216 D1035 S1035 53 UfgGfgAfuuuCfAfUfgUfaAfcCfaAfgsAf AS1035 1145 uCfuUfgGfuUfaCfaugAfAfAfuCfcCfasUfsc 0.04 0.16 0.56 0.0222 D1036 S1036 54 UfgGfGfAfuUfuCfaUfgUfaAfcCfAfAfgsAf AS1036 1146 uCfuugGfuUfaCfaUfgAfaAfuccCfasUfsc 0.06 0.31 0.78 0.0234 D1037 S1037 55 UfgGfGfAfuUfuCfAfUfgUfaAfcCfaAfgsAf AS1037 1147 uCfuUfgGfuUfaCfaugAfaAfuccCfasUfsc 0.03 0.14 0.62 0.0235 D1038 S1038 56 UfGfggAfUfuuCfAfugUfAfacCfAfagsAf AS1038 1148 uCfUfugGfUfuaCfAfugAfAfauCfCfcasUfsc 0.09 0.39 0.78 0.0239 D1039 S1039 57 AfuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1039 1149 aUfgGfAfAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.03 0.14 0.59 0.025 D1040 S1040 58 AfuGfuAfaCfcAfAfGfaGfuAfuUfccasUf AS1040 1150 aUfGfGfaAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.03 0.13 0.56 0.025 D1041 S1041 59 AfsuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1041 1151 asUfgGfAfAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.06 0.27 0.79 0.0252 D1042 S1042 60 UfgGfgAfuuuCfAfUfgUfAfAfcCfaAfgsAf AS1042 1152 uCfuUfgGfuuaCfaugAfAfAfuCfcCfasUfsc 0.05 0.27 0.67 0.0259 D1043 S1043 61 AfuGfuAfaCfcAfAfGfaGfuauUfcCfasUf AS1043 1153 aUfgGfaAfUfAfcUfcuuGfgUfuAfcAfusGfsa 0.02 0.16 0.63 0.027 D1044 S1044 62 AfsuGfuAfaCfcAfAfGfaGfuAfuucCfasUf AS1044 1154 asUfgGfAfAfuAfcUfcuuGfgsUfsuAfcAfusGfsa 0.06 0.30 0.81 0.0271 D1045 S1045 63 aUfguAfAfccAfAfgaGfGfauUfCfcasUf AS1045 1155 aUfGfgaAfUfacUfCfuuGfGfuuAfCfaUfsgsa 0.12 0.29 0.8 0.028 D1046 S1046 64 AfuGfuAfaCfcAfAfGfaguAfuUfcCfasUf AS1046 1156 aUfgGfaAfuAfCfUfcuuGfgUfuAfcAfusGfsa 0.03 0.15 0.59 0.030 D1047 S1047 65 UfgGfGfAfuUfuCfaUfgUfAfAfcCfaAfgsAf AS1047 1157 uCfuUfgGfuuaCfaUfgAfaAfuccCfasUfsc 0.08 0.44 0.83 0.0324 D1048 S1048 66 AfuGfuAfaCfcAfAfGfaGfuAfuUfcCfasUf AS1048 1158 aUfgGfaAfuAfcUfcuuGfgUfuAfcAfusGfsa 0.07 0.23 0.67 0.036 D1049 S1049 67 AfuGfuAfAfccAfAfGfAfGfuAfuuccAfsu AS1049 1159 AfUfGfGfAfAfuAfCfUfCfUfUfGfGfUfuAfCfAfusGfsa 0.08 0.23 0.73 0.037 D1050 S1050 68 UfgGfgAfuuuCfaUfgUfaAfcCfAfAfgsAf AS1050 1160 uCfuugGfuUfaCfaUfgAfAfAfuCfcCfasUfsc 0.06 0.29 0.78 0.0372 D1051 S1051 69 AfuGfuAfaccaagaguAfuUfcCfasUf AS1051 1161 aUfgGfaAfudAcdTcdTudGgdTuAfcAfusgsa 0.12 0.41 0.86 0.040 D1052 S1052 70 AfuguAfaccAfaGfdAGfdTAdTUdCcdAsu AS1052 1162 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.1 0.22 0.72 0.042 D1053 S1053 71 AfuguAfaccAfaGfdAGfdTAdTUdCcdAsu AS1053 1163 dAUdGGdAAfuAfcUfcUfuGfGfUfuAfCfAfusGfsa 0.09 0.31 0.69 0.044 D1054 S1054 72 AfuGfuAfaCfcAfaGfadGdTAfuUfcdCdAsUf AS1054 1164 adTdGGfaAfudAdCUfcUfuGfgUfuAfcAfusGfsa 0.1 0.45 0.75 0.047 D1055 S1055 73 AfuguAfaccAfaGfaGfdTAdTUdCcdAsu AS1055 1165 dAUdGGdAadTAfcUfcUfuGfgUfuAfcAfusGfsa 0.12 0.26 0.7 0.049 D1056 S1056 74 AuGuAaCcAaGaGuAuUcCasU AS1056 1166 aUgGaAuAcUcUuGgUuAcAusGsa 0.08 0.24 0.65 0.050 D1057 S1057 75 AfuguAfaccAfagaGfuauUfccasUf AS1057 1167 aUfGfGfaAfUfAfcUfCfUfuGfGfUfuAfCfAfusGfsa 0.14 0.42 0.62 0.051 D1058 S1058 76 AfuGfuAfaccaagaguAfuUfcCfasUf AS1058 1168 aUfgGfaAfudAcdTcdTudGgdTuAfcAfusGfsa 0.12 0.36 0.86 0.053 D1059 S1059 77 AfuguAfaccAfaGfdAGfdTAdTUdCcdAsu AS1059 1169 dAUdGGdAadTAfdCUfcUfuGfgUfuAfcAfusGfsa 0.09 0.27 0.7 0.054 D1060 S1060 78 adTgudAdAccdAdAgagdTadTudCcasdT AS1060 1170 adTdGgdAadTadCdTdCuudGdGuudAdCadTsgsa 0.11 0.37 0.66 0.056 D1061 S1061 79 AfuGfuAfaCfcAfaGfdAdGuAfuUfcdCdAsUf AS1061 1171 adTdGGfaAfuAfdCdTcUfuGfgUfuAfcAfusGfsa 0.1 0.31 0.77 0.059 D1062 S1062 80 AfuguAfaccAfaGfdAGfdTAdTudCcdAsu AS1062 1172 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.1 0.27 0.65 0.059 D1063 S1063 81 adTdGuadAdCccdAdGagdTdAuudCdCasu AS1063 1173 dAdTggdAdAuadCdTcudTdGgudTdAcadTsdGsa 0.12 0.44 0.82 0.064 D1064 S1064 82 AfuGfuAfaCfcAfaGfaGfdTdAuUfcdCdAsUf AS1064 1174 adTdGGfaAfdTdAcUfcUfuGfgUfuAfcAfusGfsa 0.12 0.32 0.83 0.064 D1065 S1065 83 AfuguAfaccAfaGfaGfdTAdTudCcdAsu AS1065 1175 dAUdGgdAadTAfcUfcUfuGfgUfuAfcAfusGfsa 0.13 0.34 0.72 0.066 D1066 S1066 84 AfuGfuAfaCfcAfaGfaGfudAdTUfcdCdAsUf AS1066 1176 adTdGGfadAdTAfcUfcUfuGfgUfuAfcAfusGfsa 0.11 0.33 0.72 0.067 D1067 S1067 85 AfuguAfaccAfaGfaGfdTAdTUdCcdAsu AS1067 1177 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.11 0.37 0.62 0.070 D1068 S1068 86 AfuguAfaccAfaGfaGfdTAdTUdCcdAsu AS1068 1178 dAUdGGdAAuAfcUfcUfuGfGfUfuAfCfAfusGfsa 0.16 0.33 0.64 0.072 D1069 S1069 87 aUfGfuaAfCfccAfGfagUfAfuuCfCfasu AS1069 1179 AfUfggAfAfuaCfUfcuUfGfguUfAfcaUfsGfsa 0.14 0.43 0.73 0.074 D1070 S1070 88 AfuGfuAfaCfCfAfaGfaguAfuUfcCfasUf AS1070 1180 aUfgGfaAfuAfCfUfcUfuggUfuAfcAfusGfsa 0.08 0.42 0.94 0.075 D1071 S1071 89 UfgGfgAfuuuCfaUfgUfaAfcCfaAfgsAf AS1071 1181 uCfuUfgGfuUfaCfaUfgAfAfAfuCfcCfasUfsc 0.14 0.28 0.83 0.0797 D1072 S1072 90 AfuGfuAfaCfcAfaGfAfGfuauUfcCfasUf AS1072 1182 aUfgGfaAfUfAfcucUfuGfgUfuAfcAfusGfsa 0.05 0.26 0.8 0.082 D1073 S1073 91 AfuGfuAfaCfcAfaGfadGdTdAdTUfcCfasUf AS1073 1183 aUfgGfadAdTdAdCUfcUfuGfgUfuAfcAfusGfsa 0.12 0.41 0.73 0.083 D1074 S1074 92 AfUfguAfAfccAfAfgaGfUfauUfCfcasUf AS1074 1184 aUfGfgaAfUfacUfCfuuGfGfuuAfCfausGfsa 0.14 0.44 0.75 0.086 D1075 S1075 93 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1075 1185 aUfgGfdAdAdTdAcUfcUfuGfgUfuAfcAfusGfsa 0.1 0.41 0.72 0.088 D1076 S1076 94 AfuGfuAfaCfcAfaGfaGfudAdTdTdCCfasUf AS1076 1186 aUfgdGdAdAdTAfcUfcUfuGfgUfuAfcAfusGfsa 0.15 0.45 0.86 0.088 D1077 S1077 95 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasu AS1077 1187 AfUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.08 0.46 0.95 0.092 D1078 S1078 96 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1078 1188 dAUdGGdAadTAfcUfcUfuGfgUfuAfcAfusGfsa 0.09 0.32 0.76 0.093 D1079 S1079 97 AfuguAfaccAfaGfaGfdTadTudCcdAsu AS1079 1189 dAudGgdAadTAfcUfcUfuGfgUfuAfcAfusGfsa 0.14 0.38 0.76 0.095 D1080 S1080 98 AfuGfuAfaCfcAfaGfAfGfuAfuucCfasUf AS1080 1190
aUfgGfAfAfuAfcucUfuGfgUfuAfcAfusGfsa 0.05 0.42 0.86 0.099 D1081 S1081 99 AfuGfuAfaCfcAfaGfaGfuAfuUfdCdCdAsdT AS1081 1191 dAdTdGdGaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.17 0.47 0.9 0.105 D1082 S1082 100 AfuGfuAfaccaagaguAfuUfcCfasUf AS1082 1192 aUfgGfaAfudACfudCUfudGGfudTAfcAfusgsa 0.12 0.44 0.83 0.106 D1083 S1083 101 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1083 1193 adTdGGfaAfdTdAcUfcUfuGfgUfuAfcAfusGfsa 0.11 0.34 0.74 0.109 D1084 S1084 102 AfuGfuAfAfCfcAfaGfaGfuauUfcCfasUf AS1084 1194 aUfgGfaAfUfAfcUfcUfuGfguuAfcAfusGfsa 0.1 0.45 0.93 0.117 D1085 S1085 103 AfuGfUfAfaCfcAfaGfaGfuauUfcCfasUf AS1085 1195 aUfgGfaAfUfAfcUfcUfuGfgUfuacAfusGfsa 0.07 0.42 0.78 0.120 D1086 S1086 104 aUfguAfAfccAfAfgaGfuAfuUfcCfasUf AS1086 1196 aUfgGfaAfuAfcUfCfuuGfGfuuAfCfaUfsgsa 0.17 0.45 0.83 0.1197 D1087 S1087 105 AfuGfuAfaCfcAfaGfaGfUfAfuUfcCfasu AS1087 1197 AfUfgGfaAfuacUfcUfuGfgUfuAfcAfusGfsa 0.05 0.3 0.7 0.120 D1088 S1088 106 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1088 1198 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusgsa 0.11 0.46 0.8 0.120 D1089 S1089 107 AfuGfuAfaCfcAfaGfaGfUfAfuUfcCfasUf AS1089 1199 aUfgGfaAfuacUfcUfuGfgUfuAfcAfusGfsa 0.14 0.49 0.85 0.122 D1090 S1090 108 AfuGfuAfaCfcAfaGfaGfuauUfcCfasUf AS1090 1200 aUfgGfaAfUfAfcUfcUfuGfgUfuAfcAfusGfsa 0.1 0.41 0.85 0.125 D1091 S1091 109 AfuguAfaccAfaGfaGfdTAdTudCcdAsu AS1091 1201 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.16 0.38 0.77 0.125 D1092 S1092 110 AfuGfuAfaCfcAfaGfAfGfuAfuUfcCfasu AS1092 1202 AfUfgGfaAfuAfcucUfuGfgUfuAfcAfusGfsa 0.05 0.31 0.93 0.126 D1093 S1093 111 auGfuAfaCfcAfaGfAfGfuAfuUfcCfasUf AS1093 1203 aUfgGfaAfuAfcucUfuGfgUfuAfcAfUfsGfsa 0.06 0.33 0.9 0.135 D1094 S1094 112 AfuGfuAfaCfcAfaGfaGfUfAfuUfccasUf AS1094 1204 aUfGfGfaAfuacUfcUfuGfgUfuAfcAfusGfsa 0.07 0.39 0.85 0.142 D1095 S1095 113 AfuGfuAfaCfcAfaGfAfGfuAfuUfcCfasUf AS1095 1205 aUfgGfaAfuAfcucUfuGfgUfuAfcAfusGfsa 0.09 0.39 0.76 0.146 D1096 S1096 114 AfuGfuAfaCfcAfaGfaGfUfAfuucCfasUf AS1096 1206 aUfgGfAfAfuacUfcUfuGfgUfuAfcAfusGfsa 0.06 0.38 0.85 0.147 D1097 S1097 115 AfuGfUfAfaCfcAfaGfaGfuAfuucCfasUf AS1097 1207 aUfgGfAfAfuAfcUfcUfuGfgUfuacAfusGfsa 0.12 0.47 0.87 0.147 D1098 S1098 116 AfuGfuAfaCfcAfaGfaGfuAfUfUfccasUf AS1098 1208 aUfGfGfaauAfcUfcUfuGfgUfuAfcAfusGfsa 0.06 0.42 0.85 0.151 D1099 S1099 117 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1099 1209 dAUdGGdAadTAfdCUfcUfuGfgUfuAfcAfusGfsa 0.16 0.41 0.85 0.152 D1100 S1100 118 AfuguAfaccAfaGfaGfuAfuUfcCfasUf AS1100 1210 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.15 0.48 0.72 0.152 D1101 S1101 119 AfuGfuAfaCfcAfaGfAfGfuAfuUfccasUf AS1101 1211 aUfGfGfaAfuAfcucUfuGfgUfuAfcAfusGfsa 0.06 0.38 0.94 0.158 D1102 S1102 120 AfuGfuAfaccaagaguAfuUfcCfasUf AS1102 1212 aUfgGfaAfuAfdCuCfdTuGfdGuUfacAfusGfsa 0.21 0.45 0.89 0.162 D1103 S1103 121 AfuGfuaaCfCfAfaGfaGfuAfuUfcCfasUf AS1103 1213 aUfgGfaAfuAfcUfcUfuggUfUfAfcAfusGfsa 0.14 0.49 0.95 0.163 D1104 S1104 122 AfuGfuAfaccAfaGfaGfUfAfuUfcCfasUf AS1104 1214 aUfgGfaAfuacUfcUfuGfGfUfuAfcAfusGfsa 0.06 0.36 0.92 0.163 D1105 S1105 123 AfuGfuAfaCfcAfaGfaGfuAfuucCfasUf AS1105 1215 aUfgGfAfAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.1 0.45 0.84 0.167 D1106 S1106 124 AfuGfuaaCfcAfaGfAfGfuAfuUfcCfasUf AS1106 1216 aUfgGfaAfuAfcucUfuGfgUfUfAfcAfusGfsa 0.09 0.43 0.91 0.170 D1107 S1107 125 AfuGfuAfaccAfaGfAfGfuAfuUfcCfasUf AS1107 1217 aUfgGfaAfuAfcucUfuGfGfUfuAfcAfusGfsa 0.09 0.46 1 0.171 D1108 S1108 126 AfuguAfaccAfaGfaGfdTadTudCcdAsu AS1108 1218 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.11 0.39 0.71 0.176 D1109 S1109 127 AfuGfUfAfaCfcAfaGfaGfuAfuUfccasUf AS1109 1219 aUfGfGfaAfuAfcUfcUfuGfgUfuacAfusGfsa 0.1 0.43 0.9 0.180 D1110 S1110 128 AfuGfuAfaCfcAfaGfaguAfUfUfcCfasUf AS1110 1220 aUfgGfaauAfCfUfcUfuGfgUfuAfcAfusGfsa 0.06 0.42 0.88 0.182 D1111 S1111 129 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1111 1221 dAUdGGdAAuAfcUfcUfuGfGfUfuAfCfAfusGfsa 0.18 0.49 0.79 0.183 D1112 S1112 130 AfuGfUfAfaccAfaGfaGfuAfuUfcCfasUf AS1112 1222 aUfgGfaAfuAfcUfcUfuGfGfUfuacAfusGfsa 0.14 0.48 0.85 0.195 D1113 S1113 131 AfuGfuAfaCfcAfaGfaguAfuUfcCfasUf AS1113 1223 aUfgGfaAfuAfCfUfcUfuGfgUfuAfcAfusGfsa 0.09 0.41 0.85 0.201 D1114 S1114 132 auGfuAfaCfcAfaGfaGfUfAfuUfcCfasUf AS1114 1224 aUfgGfaAfuacUfcUfuGfgUfuAfcAfUfsGfsa 0.05 0.44 0.94 0.201 D1115 S1115 133 AfuguAfaCfcAfaGfaGfUfAfuUfcCfasUf AS1115 1225 aUfgGfaAfuacUfcUfuGfgUfuAfCfAfusGfsa 0.08 0.41 0.96 0.204 D1116 S1116 134 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1116 1226 adTdGGfadAdTAfcUfcUfuGfgUfuAfcAfusGfsa 0.15 0.47 0.79 0.208 D1117 S1117 135 AfuGfuaaCfcAfaGfaGfUfAfuUfcCfasUf AS1117 1227 aUfgGfaAfuacUfcUfuGfgUfUfAfcAfusGfsa 0.08 0.42 0.92 0.224 D1118 S1118 136 auguaaccaagaguauuccasu AS1118 1228 AfUfGfGfAfAfUfAfCfUfCfUfUfGfGfUfUfAfCfAfUfsgsa 0.19 0.5 0.87 0.303 D1119 S1119 137 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1119 1229 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.14 0.55 0.89 D1120 S1120 138 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1120 1230 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.19 0.63 0.72 D1121 S1121 139 AfuGfuAfaccAfaGfaGfuAfuUfcCfasUf AS1121 1231 aUfgGfaAfuAfcUfcUfuGfGfUfuAfcAfusGfsa 0.14 0.61 0.91 D1122 S1122 140 AfUfGfuAfaCfcAfaGfaGfuAfuUfccasUf AS1122 1232 aUfGfGfaAfuAfcUfcUfuGfgUfuAfcausGfsa 0.14 0.54 0.95 D1123 S1123 141 auGfuAfAfCfcAfaGfaGfuAfuUfcCfasUf AS1123 1233 aUfgGfaAfuAfcUfcUfuGfguuAfcAfUfsGfsa 0.13 0.61 0.97 D1124 S1124 142 AfuGfuAfaCfcAfaGfaGfuAfUfUfcCfasUf AS1124 1234 aUfgGfaauAfcUfcUfuGfgUfuAfcAfusGfsa 0.14 0.56 0.94 D1125 S1125 143 AfuGfuAfaCfcaaGfaGfuAfuUfcCfasUf AS1125 1235 aUfgGfaAfuAfcUfcUfUfGfgUfuAfcAfusGfsa 0.21 0.74 0.95 D1126 S1126 144 AfUfGfuAfaCfcAfaGfaGfuAfuucCfasUf AS1126 1236 aUfgGfAfAfuAfcUfcUfuGfgUfuAfcausGfsa 0.2 0.69 0.91 D1127 S1127 145 AfuguAfAfCfcAfaGfaGfuAfuUfcCfasUf AS1127 1237 aUfgGfaAfuAfcUfcUfuGfguuAfCfAfusGfsa 0.17 0.7 0.96 D1128 S1128 146 AfUfGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1128 1238 aUfgGfaAfuAfcUfcUfuGfgUfuAfcausGfsa 0.19 0.62 0.85 D1129 S1129 147 AfuGfuAfaCfcAfaGfaGfuAfuUfCfCfasUf AS1129 1239 aUfggaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.23 0.76 0.98 D1130 S1130 148 AfuGfuAfaCfcAfagaGfuAfuUfcCfasUf AS1130 1240 aUfgGfaAfuAfcUfCfUfuGfgUfuAfcAfusGfsa 0.21 0.64 0.9 D1131 S1131 149 AfuGfuAfAfCfcaaGfaGfuAfuUfcCfasUf AS1131 1241 aUfgGfaAfuAfcUfcUfUfGfguuAfcAfusGfsa 0.17 0.7 1.01 D1132 S1132 150 AfuGfUfAfaCfcAfaGfaGfuAfuUfcCfasUf AS1132 1242 aUfgGfaAfuAfcUfcUfuGfgUfuacAfusGfsa 0.17 0.58 0.87 D1133 S1133 151 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfAfsUf AS1133 1243 augGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.33 0.89 1.05 D1134 S1134 152 AfUfGfuAfaCfcAfaGfaguAfuUfcCfasUf AS1134 1244 aUfgGfaAfuAfCfUfcUfuGfgUfuAfcausGfsa 0.16 0.64 0.96 D1135 S1135 153 AfuGfUfAfaCfcAfaGfaguAfuUfcCfasUf AS1135 1245 aUfgGfaAfuAfCfUfcUfuGfgUfuacAfusGfsa 0.12 0.53 0.96 D1136 S1136 154 AfuGfuAfAfCfcAfagaGfuAfuUfcCfasUf AS1136 1246 aUfgGfaAfuAfcUfCfUfuGfguuAfcAfusGfsa 0.16 0.58 0.98 D1137 S1137 155 AfuGfuAfAfCfcAfaGfaGfuAfuUfcCfasUf AS1137 1247 aUfgGfaAfuAfcUfcUfuGfguuAfcAfusGfsa 0.16 0.6 0.91 D1138 S1138 156 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1138 1248 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsAf 0.1 0.54 0.91 D1139 S1139 157 AfUfGfuAfaCfcAfagaGfuAfuUfcCfasUf AS1139 1249 aUfgGfaAfuAfcUfCfUfuGfgUfuAfcausGfsa 0.24 0.68 0.98 D1140 S1140 158 AfuGfUfAfaCfcAfagaGfuAfuUfcCfasUf AS1140 1250 aUfgGfaAfuAfcUfCfUfuGfgUfuacAfusGfsa 0.13 0.75 0.9 D1141 S1141 159 AfuGfuAfAfCfcAfaGfaguAfuUfcCfasUf AS1141 1251 aUfgGfaAfuAfCfUfcUfuGfguuAfcAfusGfsa 0.15 0.52 1.05 D1142 S1142 160 AfuGfuAfaCfCfAfaGfaGfuAfuUfcCfasUf AS1142 1252 aUfgGfaAfuAfcUfcUfuggUfuAfcAfusGfsa 0.16 0.66 0.89 D1143 S1143 161 auGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1143 1253 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfUfsGfsa 0.12 0.51 0.89 D1144 S1144 162 AfUfGfuAfaCfcaaGfaGfuAfuUfcCfasUf AS1144 1254 aUfgGfaAfuAfcUfcUfUfGfgUfuAfcausGfsa 0.25 0.71 0.95 D1145 S1145 163 AfuGfUfAfaCfcaaGfaGfuAfuUfcCfasUf AS1145 1255 aUfgGfaAfuAfcUfcUfUfGfgUfuacAfusGfsa 0.17 0.74 0.98 D1146 S1146 164 AfuguAfaCfcAfaGfaGfuAfuUfcCfasUf AS1146 1256 aUfgGfaAfuAfcUfcUfuGfgUfuAfCfAfusGfsa 0.11 0.51 0.86 D1147 S1147 165 AfuGfuAfaCfcAfaGfaGfuAfuUfccasUf AS1147 1257 aUfGfGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.1 0.52 0.83 D1148 S1148 166 AfUfGfuAfaccAfaGfaGfuAfuUfcCfasUf AS1148 1258 aUfgGfaAfuAfcUfcUfuGfGfUfuAfcausGfsa 0.14 0.63 0.98 D1149 S1149 167 AfuGfuAfAfCfcAfaGfaGfuAfuucCfasUf AS1149 1259 aUfgGfAfAfuAfcUfcUfuGfguuAfcAfusGfsa 0.13 0.58 0.88 D1150 S1150 168 AfuGfuaaCfcAfaGfaGfuAfuUfcCfasUf AS1150 1260 aUfgGfaAfuAfcUfcUfuGfgUfUfAfcAfusGfsa 0.15 0.62 0.94 D1151 S1151 169 AfUfGfuaaCfcAfaGfaGfuAfuUfcCfasUf AS1151 1261 aUfgGfaAfuAfcUfcUfuGfgUfUfAfcausGfsa 0.18 0.73 0.94 D1152 S1152 170 auGfUfAfaCfcAfaGfaGfuAfuUfcCfasUf AS1152 1262 aUfgGfaAfuAfcUfcUfuGfgUfuacAfUfsGfsa 0.13 0.53 0.97 D1153 S1153 171 AfuGfuAfAfCfcAfaGfaGfuAfuUfccasUf AS1153 1263 aUfGfGfaAfuAfcUfcUfuGfguuAfcAfusGfsa 0.13 0.53 0.98 D1154 S1154 172 UfgGfgAfuUfuCfaUfgUfaAfcCfaAfgsAf AS1154 1264 uCfuUfgGfuUfaCfaUfgAfaAfuCfcCfasUfsc 0.09 0.5 0.78 D1155 S1155 173 UfgGfGfAfuuuCfaUfgUfAfAfcCfaAfgsAf AS1155 1265 uCfuUfgGfuuaCfaUfgAfAfAfuccCfasUfsc 0.13 0.62 0.89 D1156 S1156 174 UfgGfgAfuuuCfaUfGfUfaAfcCfaAfgsAf AS1156 1266 uCfuUfgGfuUfacaUfgAfAfAfuCfcCfasUfsc 0.12 0.65 0.85 D1157 S1157 175 UfgGfgAfuUfuCfaUfgUfAfAfcCfaAfgsAf AS1157 1267 uCfuUfgGfuuaCfaUfgAfaAfuCfcCfasUfsc 0.11 0.54 0.85 D1158 S1158 176 UfgGfgAfuuuCfaUfgUfAfAfcCfaAfgsAf AS1158 1268 uCfuUfgGfuuaCfaUfgAfAfAfuCfcCfasUfsc 0.13 0.53 0.8 D1159 S1159 177 UfGfggAfUfuUfcAfuGfuAfAfccAfAfgsAf AS1159 1269 uCfuuGfGfuuAfcAfuGaAfauCfCfcasUfsc 0.59 0.89 0.81 D1160 S1160 178 UfGfggAfUfuuCfaUfgUfAfAfcCfaAfgsAf AS1160 1270 uCfuUfgGfuuaCfaUfgAfAfauCfCfcasUfsc 0.16 0.72 0.9 D1161 S1161 179 UfgGfgAfuUfucaUfGfUfaAfcCfaAfgsAf AS1161 1271 uCfuUfgGfuUfacaUfGfAfaAfuCfcCfasUfsc 0.27 0.69 0.86 D1162 S1162 180 AfuGfuAfaCfcaaGfaGfUfAfuUfcCfasUf AS1162 1272 aUfgGfaAfuacUfcUfUfGfgUfuAfcAfusGfsa 0.12 0.6 0.95 D1163 S1163 181 AfuGfuAfaccAfaGfaGfuAfUfUfcCfasUf AS1163 1273 aUfgGfaauAfcUfcUfuGfGfUfuAfcAfusGfsa 0.05 0.56 1.02
D1164 S1164 182 AfuGfuAfaCfcAfagaGfUfAfuUfcCfasUf AS1164 1274 aUfgGfaAfuacUfCfUfuGfgUfuAfcAfusGfsa 0.13 0.55 1 D1165 S1165 183 AfuGfuAfaCfcaaGfaGfuAfUfUfcCfasUf AS1165 1275 aUfgGfaauAfcUfcUfUfGfgUfuAfcAfusGfsa 0.09 0.6 0.97 D1166 S1166 184 AfuguAfaCfCfAfaGfaGfuAfuUfcCfasUf AS1166 1276 aUfgGfaAfuAfcUfcUfuggUfuAfCfAfusGfsa 0.15 0.59 0.91 D1167 S1167 185 AfuGfuAfaCfcAfagaGfuAfUfUfcCfasUf AS1167 1277 aUfgGfaauAfcUfCfUfuGfgUfuAfcAfusGfsa 0.11 0.59 1 D1168 S1168 186 AfuGfuAfaCfCfAfagaGfuAfuUfcCfasUf AS1168 1278 aUfgGfaAfuAfcUfCfUfuggUfuAfcAfusGfsa 0.13 0.57 0.94 D1169 S1169 187 auGfuAfaCfcAfaGfaGfuAfUfUfcCfasUf AS1169 1279 aUfgGfaauAfcUfcUfuGfgUfuAfcAfUfsGfsa 0.08 0.5 0.9 D1170 S1170 188 AfuguAfaCfcAfaGfaGfuAfUfUfcCfasUf AS1170 1280 aUfgGfaauAfcUfcUfuGfgUfuAfCfAfusGfsa 0.06 0.53 0.91 D1171 S1171 189 auGfuAfaCfcAfaGfaGfuAfuUfCfCfasUf AS1171 1281 aUfggaAfuAfcUfcUfuGfgUfuAfcAfUfsGfsa 0.07 0.56 0.89 D1172 S1172 190 AfuGfuAfaCfCfAfaGfaGfuAfuucCfasUf AS1172 1282 aUfgGfAfAfuAfcUfcUfuggUfuAfcAfusGfsa 0.13 0.59 0.98 D1173 S1173 191 AfuGfuAfaCfcaaGfAfGfuAfuUfcCfasUf AS1173 1283 aUfgGfaAfuAfcucUfUfGfgUfuAfcAfusGfsa 0.2 0.65 1.03 D1174 S1174 192 AfuGfuaaCfcAfaGfaGfuAfUfUfcCfasUf AS1174 1284 aUfgGfaauAfcUfcUfuGfgUfUfAfcAfusGfsa 0.07 0.51 0.95 D1175 S1175 193 AfuguAfaCfcAfaGfaGfuAfuUfCfCfasUf AS1175 1285 aUfggaAfuAfcUfcUfuGfgUfuAfCfAfusGfsa 0.2 0.53 0.76 D1176 S1176 194 auGfuAfaCfcAfaGfaGfuAfuUfcCfAfsUf AS1176 1286 augGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.74 0.98 0.81 D1177 S1177 195 AfuGfuAfaCfcAfaGfaGfuAfuucCfAfsUf AS1177 1287 augGfAfAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.43 0.64 0.88 D1178 S1178 196 auguaaccAfaGfaGfuAfuUfcCfasUf AS1178 1288 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.17 0.49 0.81 D1179 S1179 197 AfuGfuaaCfcAfaGfaGfuAfuUfCfCfasUf AS1179 1289 aUfggaAfuAfcUfcUfuGfgUfUfAfcAfusGfsa 0.22 0.65 0.73 D1180 S1180 198 AfuguAfaCfcAfaGfaGfuAfuUfcCfAfsUf AS1180 1290 augGfaAfuAfcUfcUfuGfgUfuAfcAfUfsGfsa 0.6 1.09 0.8 D1181 S1181 199 auGfuAfaCfcAfaGfaGfuAfuUfccasu AS1181 1291 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.3 0.78 0.78 D1182 S1182 200 auguaaccaaGfaGfuAfuUfcCfasUf AS1182 1292 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.35 0.73 0.84 D1183 S1183 201 AfuGfuAfaccAfaGfaGfuAfuUfCfCfasUf AS1183 1293 aUfggaAfuAfcUfcUfuGfGfUfuAfcAfusGfsa 0.19 0.6 0.94 D1184 S1184 202 AfuGfuaaCfcAfaGfaGfuAfuUfcCfAfsUf AS1184 1294 augGfaAfuAfcUfcUfuGfgUfuAfCfAfusGfsa 0.61 1.08 0.8 D1185 S1185 203 auGfuAfaCfcAfaGfaGfuAfuuccasu AS1185 1295 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.16 0.52 0.72 D1186 S1186 204 auguaaccaagaGfuAfuUfcCfasUf AS1186 1296 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.2 0.53 0.74 D1187 S1187 205 AfuGfuAfaCfcaaGfaGfuAfuUfCfCfasUf AS1187 1297 aUfggaAfuAfcUfcUfUfGfgUfuAfcAfusGfsa 0.34 0.66 0.85 D1188 S1188 206 AfuGfuAfaccAfaGfaGfuAfuUfcCfAfsUf AS1188 1298 augGfaAfuAfcUfcUfuGfgUfUfAfcAfusGfsa 0.61 0.98 1.02 D1189 S1189 207 AfuGfuAfaCfcAfaGfaGfuAfuuccasu AS1189 1299 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.3 0.73 0.85 D1190 S1190 208 auguaaccaagaguauuccasu AS1190 1300 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.28 0.69 0.78 D1191 S1191 209 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1191 1301 aUfgGfaAfuAfcUfcUfugdGudTadCadTsgsa 0.33 0.88 0.64 D1192 S1192 210 AfuGfuAfaCfcAfagaGfuAfuUfCfCfasUf AS1192 1302 aUfggaAfuAfcUfCfUfuGfgUfuAfcAfusGfsa 0.31 0.64 0.83 D1193 S1193 211 AfuGfuAfaCfcaaGfaGfuAfuUfcCfAfsUf AS1193 1303 augGfaAfuAfcUfcUfuGfGfUfuAfcAfusGfsa 0.64 0.82 0.92 D1194 S1194 212 AfuGfuAfaCfcAfaGfaGfuauuccasu AS1194 1304 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.21 0.62 0.77 D1195 S1195 213 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1195 1305 aUfgGfaAfuAfcUfcUfuGfGfUfuAfCfAfusGfsa 0.17 0.7 0.95 D1196 S1196 214 AfuGfuAfaCfcAfaGfaguAfuUfCfCfasUf AS1196 1306 aUfggaAfuAfCfUfcUfuGfgUfuAfcAfusGfsa 0.19 0.71 0.65 D1197 S1197 215 AfuGfuAfaCfcAfagaGfuAfuUfcCfAfsUf AS1197 1307 augGfaAfuAfcUfcUfUfGfgUfuAfcAfusGfsa 0.64 0.82 0.93 D1198 S1198 216 auguAfaCfcAfaGfaGfuAfuUfccasu AS1198 1308 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.19 0.65 0.72 D1199 S1199 217 AfuGfuAfaCfcAfaGfaGfuauUfCfCfasUf AS1199 1309 aUfggaAfUfAfcUfcUfuGfgUfuAfcAfusGfsa 0.15 0.52 0.64 D1200 S1200 218 AfuGfuAfaCfcAfaGfaguAfuUfcCfAfsUf AS1200 1310 augGfaAfuAfcUfCfUfuGfgUfuAfcAfusGfsa 0.48 0.74 0.92 D1201 S1201 219 auguAfaCfcAfaGfaGfuAfuUfcCfasu AS1201 1311 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.17 0.71 0.77 D1202 S1202 220 AfuGfuAfaCfcAfaGfaGfuauUfcCfAfsUf AS1202 1312 augGfaAfuAfCfUfcUfuGfgUfuAfcAfusGfsa 0.43 0.69 0.85 D1203 S1203 221 auguaaCfcAfaGfaGfuAfuUfcCfasUf AS1203 1313 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.14 0.61 0.76 D1204 S1204 222 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1204 1314 adTdGGfaAfudAdCUfcUfuGfgUfuAfcAfusGfsa 0.16 0.56 0.89 D1205 S1205 223 AfuGfuAfaCfcAfaGfaGfdTdAdTdTcCfasUf AS1205 1315 aUfgGfdAdAdTdAcUfcUfuGfgUfuAfcAfusGfsa 0.13 0.57 0.9 D1206 S1206 224 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1206 1316 adTdGdGdAAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.29 0.73 0.89 D1207 S1207 225 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1207 1317 adTdGGfaAfuAfdCdTcUfuGfgUfuAfcAfusGfsa 0.16 0.56 0.78 D1208 S1208 226 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1208 1318 aUfdGdGdAdAuAfcUfcUfuGfgUfuAfcAfusGfsa 0.22 0.67 0.89 D1209 S1209 227 AfuguAfaccAfaGfaGfuAfuUfcCfasUf AS1209 1319 aUfgGfaAfuAfcUfcUfuGfGfUfuAfCfAfusGfsa 0.14 0.55 0.78 D1210 S1210 228 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1210 1320 aUfgdGdAdAdTAfcUfcUfuGfgUfuAfcAfusGfsa 0.14 0.5 0.84 D1211 S1211 229 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1211 1321 aUfgGfadAdTdAdCUfcUfuGfgUfuAfcAfusGfsa 0.14 0.59 0.72 D1212 S1212 230 auguaaccaaGfaGfuAfuUfcCfasUf AS1212 1322 aUfgGfaAfuAfcUfcUfugdGudTadCadTsgsa 0.21 0.74 0.77 D1213 S1213 231 AfuGfuAfaCfcAfaGfaGfuAfudTdCdCdAsUf AS1213 1323 adTdGdGdAAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.15 0.53 0.91 D1214 S1214 232 aUfgUfaAfcCfaAfgAfgUfaUfuCfcAfsu AS1214 1324 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.12 0.71 0.87 D1215 S1215 233 AfuGfuAfaCfcAfaGfaGfuAfdTdTdCdCasUf AS1215 1325 aUfdGdGdAdAuAfcUfcUfuGfgUfuAfcAfusGfsa 0.18 0.67 0.97 D1216 S1216 234 AfuGfuAfaccaagaguAfuUfcCfasUf AS1216 1326 aUfgGfaAfuacucuuggUfuAfcAfusgsa 0.36 0.87 1.07 D1217 S1217 235 AfuGfuAfaccaagaguAfuUfcCfasUf AS1217 1327 aUfgGfaAfuAfCfUfCfUfuGfGfuuAfcAfusgsa 0.37 0.73 1.03 D1218 S1218 236 AfUfguAfAfccAfAfgaGfUfauUfCfcasUf AS1218 1328 aUfGfgaAfUfacUfCfuuGfGfuuAfCfausGfsa 0.23 0.42 0.84 D1219 S1219 237 AfuGfuAfaccaagaguAfuUfcCfasUf AS1219 1329 aUfgGfaAfuaCfUfcUfUfgGfuuAfcAfusgsa 0.43 0.71 1.03 D1220 S1220 238 AfuGfuAfaccaagaguAfuUfcCfasUf AS1220 1330 aUfgGfaAfuAfcUfCfUfuGfGfuuAfcAfusgsa 0.37 0.63 0.99 D1221 S1221 239 AfuGfuAfaccaagaguAfuUfcCfasUf AS1221 1331 aUfgGfaAfuAfcUfCfUfuGfGfuUfacAfusgsa 0.29 0.84 0.88 D1222 S1222 240 AfuGfuAfaccaagaguAfuUfcCfasUf AS1222 1332 aUfgGfaAfuaCfuCfuUfgGfuuAfcAfusgsa 0.31 0.8 0.99 D1223 S1223 241 auGfuAfAfccAfaGfagUfaUfUfcCfasUf AS1223 1333 aUfgGfaaUfaCfUfcUfuGfGfuuAfcAfAfsgsa 0.09 0.52 0.82 D1224 S1224 242 AfuGfuAfaccaagaguAfuUfcCfasUf AS1224 1334 aUfgGfaAfuadCudCudTgdGuuAfcAfusgsa 0.22 0.79 1 D1225 S1225 243 auGfuaAfccAfagAfguAfuuCfcasUf AS1225 1335 aUfGfgAfAfuAfCfuCfUfuGfGfuUfAfcAfUfsGfsa 0.31 0.76 0.84 D1226 S1226 244 AfuGfuAfaccaagaguAfuUfcCfasUf AS1226 1336 aUfgGfaAfuadCUfcdTUfgdGuuAfcAfusgsa 0.26 0.64 0.87 D1227 S1227 245 augUfaacCfaagAfguaUfuccAfsu AS1227 1337 aUfgGfAfaUfAfCfuCfUfUfgGfUfUfaCfAfUfsGfsa 0.33 0.79 0.81 D1228 S1228 246 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1228 1338 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.464 0.932 0.978 D1229 S1229 247 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1229 1339 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.453 1.047 1.178 D1230 S1230 248 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1230 1340 aUfgGfaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.831 0.967 1.151 D1231 S1231 249 auGfuAfAfCfcAfaGfaGfuAfuUfcCfasu AS1231 1341 AfUfgGfaAfuAfcUfcUfuGfguuAfcAfUfsGfsa 0.09 0.5 1.07 D1232 S1232 250 AfuGfuAfaCfCfAfaGfaGfuAfuUfcCfasu AS1232 1342 AfUfgGfaAfuAfcUfcUfuggUfuAfcAfusGfsa 0.11 0.54 1.1 D1233 S1233 251 AfuGfuAfaCfcAfaGfaGfuAfuUfCfCfasu AS1233 1343 AfUfggaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.19 0.61 0.74 D1234 S1234 252 aUfgUfaAfcCfaAfgAfgUfaUfuCfcAfsu AS1234 1344 AfuGfgAfaUfaCfuCfuUfgGfuUfaCfaUfsgsAf 0.22 0.61 0.98 D1235 S1235 253 aUfgUfaAfcCfaAfgAfgUfaUfuCfcAfsu AS1235 1345 AfuGfgAfaUfaCfuCfuUfgGfuUfaCfaUfsgsAf 0.27 0.69 0.92 D1236 S1236 254 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1236 1346 AfuGfgAfaUfaCfuCfuUfgGfuUfaCfaUfsgsAf 0.54 1.08 0.8 D1237 S1237 255 augUfaAfccaAfgAfguaUfuCfcasu AS1237 1347 AfUfGfgAfaUfAfCfuCfuUfGfGfuUfaCfAfUfsgsa 0.29 0.61 0.79 D1238 S1238 256 AfugUfaAfccaAfgAfguaUfuCfcasu AS1238 1348 AfUfGfgAfaUfAfCfuCfuUfGfGfuUfaCfAfusgsa 0.31 0.6 0.88 D1239 S1239 257 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1239 1349 dAUdGGdAauAfcUfcUfuGfgUfuAfcAfusGfsa 0.2 0.67 0.85 D1240 S1240 258 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1240 1350 dAUdGgdAauAfcUfcUfuGfgUfuAfcAfusGfsa 0.23 0.58 0.68 D1241 S1241 259 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1241 1351 dAudGgdAauAfcUfcUfuGfgUfuAfcAfusGfsa 0.25 0.65 0.78 D1242 S1242 260 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1242 1352 dAUdGgdAadTAfcUfcUfuGfgUfuAfcAfusGfsa 0.18 0.64 0.84 D1243 S1243 261 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1243 1353 dAUdGGdAAfuAfcUfcUfuGfGfUfuAfCfAfusGfsa 0.19 0.72 0.87 D1244 S1244 262 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1244 1354 dAUdGgdAadTAfdCUfcUfuGfgUfuAfcAfusGfsa 0.16 0.55 0.8 D1245 S1245 263 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1245 1355 dAUdGGdAAuAfcUfcUfuGfgUfuAfcAfusGfsa 0.22 0.51 0.9 D1246 S1246 264 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1246 1356 dAudGgdAadTAfcUfcUfuGfgUfuAfcAfusGfsa 0.27 0.78 0.66 D1247 S1247 265 AfuGfuAfaCfcAfaGfaGfuAfuUfcCfasUf AS1247 1357 dAdTdGdGaAfuAfcUfcUfuGfgUfuAfcAfusGfsa 0.16 0.57 0.97
D1248 S1248 266 AfacaAfuguUfcUfuGfdCUdCudAudAsa AS1248 1358 dTUdAudAgdAGfcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.09 0.36 0.0047 D1249 S1249 267 AfaCfaGfuGfuUfcUfuGfCfUfcUfaUfasa AS1249 1359 UfUfaUfaGfagcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.10 0.47 0.005 D1250 S1250 268 AfaCfaGfuGfuUfcUfugcUfcUfAfUfasAf AS1250 1360 uUfauaGfaGfCfAfaGfaAfcAfcUfgUfusUfsu 0.07 0.14 0.55 0.005 D1251 S1251 269 AfaCfaGfuGfuUfcUfuGfcucUfAfUfasAf AS1251 1361 uUfauaGfAfGfcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.14 0.49 0.006 D1252 S1252 270 cAGuGuucuuGcucuAuAAdTdT AS1252 1362 UuAuAGAGcAAGAAcACUGdTdT 0.006 D1253 S1253 271 AfaCfaGfuGfuUfcUfugcUfCfUfaUfasAf AS1253 1363 uUfaUfagaGfCfAfaGfaAfcAfcUfgUfusUfsu 0.05 0.12 0.43 0.006 D1254 S1254 272 AfaCfaGfuGfuUfCfUfuGfcUfcUfaUfasa AS1254 1364 UfUfaUfaGfaGfcAfagaAfcAfcUfgUfusUfsu 0.06 0.13 0.39 0.006 D1255 S1255 273 AfaCfaGfuGfuUfcUfuGfcUfCfUfaUfasa AS1255 1365 UfUfaUfagaGfcAfaGfaAfcAfcUfgUfusUfsu 0.08 0.17 0.48 0.007 D1256 S1256 274 AfaCfaGfuGfuUfcUfUfGfcUfcUfaUfasa AS1256 1366 UfUfaUfaGfaGfcaaGfaAfcAfcUfgUfusUfsu 0.08 0.14 0.40 0.007 D1257 S1257 275 AfaCfaGfuGfuUfcUfuGfcUfCfUfaUfasAf AS1257 1367 uUfaUfagaGfcAfaGfaAfcAfcUfgUfusUfsUf 0.07 0.12 0.40 0.007 D1258 S1258 276 AfaCfaguGfuUfCfUfuGfcUfcUfaUfasAf AS1258 1368 uUfaUfaGfaGfcAfagaAfcAfCfUfgUfusUfsu 0.08 0.13 0.41 0.007 D1259 S1259 277 AfaCfAfGfuGfuUfcUfuGfcucUfaUfasAf AS1259 1369 uUfaUfaGfAfGfcAfaGfaAfcAfcugUfusUfsu 0.05 0.11 0.35 0.008 D1260 S1260 278 AfacaGfuGfuUfCfUfuGfcUfcUfaUfasAf AS1260 1370 uUfaUfaGfaGfcAfagaAfcAfcUfGfUfusUfsu 0.06 0.12 0.40 0.008 D1261 S1261 279 AfacaGfuGfuUfcUfuGfcUfCfUfaUfasAf AS1261 1371 uUfaUfagaGfcAfaGfaAfcAfcUfGfUfusUfsu 0.06 0.13 0.42 0.008 D1262 S1262 280 AfaCfaGfuGfuUfcUfuGfcucUfaUfasAf AS1262 1372 uUfaUfaGfAfGfcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.13 0.37 0.008 D1263 S1263 281 cAGuGuucuuGcucuAuAAdTdT AS1263 1373 UuAuAGAGcAAGAAcACUGdTdT 0.008 D1264 S1264 282 AfaCfaGfuGfuUfcUfuGfCfUfcUfauasAf AS1264 1374 uUfAfUfaGfagcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.12 0.50 0.008 D1265 S1265 283 AfaCfaGfuguUfCfUfuGfcUfcUfaUfasAf AS1265 1375 uUfaUfaGfaGfcAfagaAfCfAfcUfgUfusUfsu 0.12 0.13 0.48 0.009 D1266 S1266 284 AfacaGfuGfuUfcUfuGfcUfcUfAfUfasAf AS1266 1376 uUfauaGfaGfcAfaGfaAfcAfcUfGfUfusUfsu 0.07 0.15 0.51 0.009 D1267 S1267 285 AfacaAfuguUfcUfuGfdCudCudAudAsa AS1267 1377 dTudAudAgdAGfcAfaGfaAfcAfcAfgUfusUfsu 0.06 0.14 0.48 0.0088 D1268 S1268 286 AfaCfaGfuGfuUfCfUfuGfcucUfaUfasAf AS1268 1378 uUfaUfaGfAfGfcAfagaAfcAfcUfgUfusUfsu 0.05 0.09 0.35 0.009 D1269 S1269 287 cAGuGuucuuGcucuAuAAdTdT AS1269 1379 UuAuAGAGcAAGAAcACUGdTdT 0.009 D1270 S1270 288 aaCfaGfuGfuUfcUfuGfcUfCfUfaUfasAf AS1270 1380 uUfaUfagaGfcAfaGfaAfcAfcUfgUfUfsUfsu 0.07 0.14 0.49 0.009 D1271 S1271 289 AfaCfaGfUfGfuUfcUfuGfcucUfaUfasAf AS1271 1381 uUfaUfaGfAfGfcAfaGfaAfcacUfgUfusUfsu 0.06 0.10 0.36 0.009 D1272 S1272 290 cAGuGuucuuGcucuAuAAdTdT AS1272 1382 UuAuAGAGcAAGAAcACUGdTdT 0.009 D1273 S1273 291 AfaCfaGfUfGfuUfcUfuGfcUfcUfaUfasAf AS1273 1383 uUfaUfaGfaGfcAfaGfaAfcacUfgUfusUfsUf 0.06 0.13 0.51 0.009 D1274 S1274 292 AfaCfaGfuGfuUfCfUfuGfcUfcuaUfasAf AS1274 1384 uUfaUfAfGfaGfcAfagaAfcAfcUfgUfusUfsu 0.06 0.12 0.46 0.010 D1275 S1275 293 cAGuGuucuuGcucuAuAAdTdT AS1275 1385 UuAuAGAGcAAGAAcACUGdTdT 0.010 D1276 S1276 294 AfaCfaGfuGfuUfCfUfuGfcUfcUfauasAf AS1276 1386 uUfAfUfaGfaGfcAfagaAfcAfcUfgUfusUfsu 0.06 0.14 0.47 0.010 D1277 S1277 295 AfaCfaguGfuUfcUfuGfcUfCfUfaUfasAf AS1277 1387 uUfaUfagaGfcAfaGfaAfcAfCfUfgUfusUfsu 0.07 0.15 0.50 0.010 D1278 S1278 296 AfaCfaGfuGfuUfCfUfugcUfcUfaUfasAf AS1278 1388 uUfaUfaGfaGfCfAfagaAfcAfcUfgUfusUfsu 0.06 0.13 0.43 0.010 D1279 S1279 297 cAGuGuucuuGcucuAuAAdTdT AS1279 1389 UuAuAGAGcAAGAAcACUGdTdT 0.010 D1280 S1280 298 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasa AS1280 1390 UfUfaUfaGfaGfcAfaGfaAfcAfcUfgUfususu 0.06 0.14 0.45 0.010 D1281 S1281 299 AfaCfAfGfuGfuUfcUfuGfcUfcUfaUfasa AS1281 1391 UfUfaUfaGfaGfcAfaGfaAfcAfcugUfusUfsu 0.07 0.18 0.46 0.011 D1282 S1282 300 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1282 1392 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.15 0.55 0.011 D1283 S1283 301 AfaCfaGfuGfuUfcUfuGfcucUfaUfasAf AS1283 1393 uUfaUfaGfAfGfcAfaGfaAfcAfcUfgUfususu 0.07 0.12 0.45 0.011 D1284 S1284 302 AfacaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1284 1394 uUfaUfaGfaGfcAfaGfaAfcAfcUfGfUfusUfsu 0.06 0.13 0.48 0.011 D1285 S1285 303 AfAfCfaGfuGfuUfcUfuGfcucUfaUfasAf AS1285 1395 uUfaUfaGfAfGfcAfaGfaAfcAfcUfguusUfsu 0.06 0.11 0.40 0.011 D1286 S1286 304 AfaCfAfGfuGfuUfcUfuGfcUfcUfauasAf AS1286 1396 uUfAfUfaGfaGfcAfaGfaAfcAfcugUfusUfsu 0.06 0.16 0.47 0.011 D1287 S1287 305 AfaCfaGfuGfuUfcUfugcUfcUfaUfasAf AS1287 1397 uUfaUfaGfaGfCfAfaGfaAfcAfcUfgUfususu 0.07 0.19 0.46 0.012 D1288 S1288 306 AfaCfaGfuGfuUfcUfugcUfcUfaUfasAf AS1288 1398 uUfaUfaGfaGfCfAfaGfaAfcAfcUfgUfusUfsu 0.06 0.17 0.46 0.012 D1289 S1289 307 AfaCfaGfuGfuUfcUfUfGfcucUfaUfasAf AS1289 1399 uUfaUfaGfAfGfcaaGfaAfcAfcUfgUfusUfsu 0.05 0.09 0.31 0.012 D1290 S1290 308 AfAfCfaGfuGfuUfcUfuGfcUfcUfaUfasa AS1290 1400 UfUfaUfaGfaGfcAfaGfaAfcAfcUfguusUfsu 0.06 0.16 0.49 0.013 D1291 S1291 309 AfaCfaGfuGfuUfCfUfuGfcUfcUfaUfasAf AS1291 1401 uUfaUfaGfaGfcAfagaAfcAfcUfgUfusUfsUf 0.06 0.11 0.32 0.013 D1292 S1292 310 AfaCfAfGfuGfuUfcUfugcUfcUfaUfasAf AS1292 1402 uUfaUfaGfaGfCfAfaGfaAfcAfcugUfusUfsu 0.06 0.14 0.44 0.013 D1293 S1293 311 AfaCfaGfUfGfuUfcUfuGfcUfcUfaUfasa AS1293 1403 UfUfaUfaGfaGfcAfaGfaAfcacUfgUfusUfsu 0.07 0.16 0.39 0.013 D1294 S1294 312 AfaCfAfGfuGfuUfcUfuGfcUfcuaUfasAf AS1294 1404 uUfaUfAfGfaGfcAfaGfaAfcAfcugUfusUfsu 0.07 0.18 0.41 0.014 D1295 S1295 313 AfaCfaGfUfGfuUfcUfuGfcUfcuaUfasAf AS1295 1405 uUfaUfAfGfaGfcAfaGfaAfcacUfgUfusUfsu 0.07 0.18 0.47 0.014 D1296 S1296 314 adAdCagdTdGuudCdTugdCdTcudAdTasa AS1296 1406 dTdTaudAdGagdCdAagdAdAcadCdTgudTsdTsu 0.12 0.21 0.68 0.0146 D1297 S1297 315 AfacaGfUfGfuUfcUfuGfcUfcUfaUfasAf AS1297 1407 uUfaUfaGfaGfcAfaGfaAfcacUfGfUfusUfsu 0.06 0.15 0.50 0.016 D1298 S1298 316 AfaCfaGfUfGfuUfcUfuGfcUfcUfauasAf AS1298 1408 uUfAfUfaGfaGfcAfaGfaAfcacUfgUfusUfsu 0.08 0.17 0.50 0.016 D1299 S1299 317 AfaCfaguGfuUfcUfuGfcUfcUfaUfasAf AS1299 1409 uUfaUfaGfaGfcAfaGfaAfcAfCfUfgUfususu 0.07 0.16 0.50 0.018 D1300 S1300 318 AfaCfaGfuGfuUfcUfUfGfcUfcUfauasAf AS1300 1410 uUfAfUfaGfaGfcaaGfaAfcAfcUfgUfusUfsu 0.06 0.12 0.43 0.020 D1301 S1301 319 AfaCfaGfUfGfuUfcUfugcUfcUfaUfasAf AS1301 1411 uUfaUfaGfaGfCfAfaGfaAfcacUfgUfusUfsu 0.07 0.17 0.45 0.021 D1302 S1302 320 AfaCfaGfuguUfcUfUfGfcUfcUfaUfasAf AS1302 1412 uUfaUfaGfaGfcaaGfaAfCfAfcUfgUfusUfsu 0.06 0.14 0.49 0.021 D1303 S1303 321 AfAfCfaguGfuUfcUfuGfcUfcUfaUfasAf AS1303 1413 uUfaUfaGfaGfcAfaGfaAfcAfCfUfguusUfsu 0.07 0.24 0.51 0.022 D1304 S1304 322 AfaCfaGfuGfuucUfuGfcUfcUfaUfasAf AS1304 1414 uUfaUfaGfaGfcAfaGfAfAfcAfcUfgUfususu 0.09 0.27 0.47 0.033 D1305 S1305 323 aadCdAgudGdTucdTdTgcdTdCuadTdAsa AS1305 1415 udTadTdAgadGdCaadGdAacdAdCugdTdTsusu 0.19 0.36 0.86 0.045 D1306 S1306 324 AfacaGfuguUfcUfuGfdCUdCUdAudAsa AS1306 1416 dTUdAUdAGfaGfcAfaGfaAfCfAfcUfGfUfusUfsu 0.08 0.22 0.61 D1307 S1307 325 AfacaGfuguUfcUfdTGfdCUdCUdAudAsa AS1307 1417 dTUdAUdAGfaGfcAfaGfaAfCfAfcUfGfUfusUfsu 0.13 0.39 0.84 D1308 S1308 326 AfacaGfuguUfcUfuGfdCUdCUdAudAsa AS1308 1418 dTUdAUdAgdAGfcAfaGfaAfcAfcUfgUfusUfsu 0.09 0.13 0.48 D1309 S1309 327 AfacaGfuguUfcUfdTGfdCUdCUdAudAsa AS1309 1419 dTUdAUdAgdAGfdCAfaGfaAfcAfcUfgUfusUfsu 0.07 0.13 0.58 D1310 S1310 328 AfacaAfuguUfcUfdTGfdCUdCudAudAsa AS1310 1420 dTUdAudAgdAGfdCAfaGfaAfcAfcAfgUfusUfsu 0.07 0.14 0.55 D1311 S1311 329 AfaCfaAfuGfuUfcUfuGfcUfcUfdAdTdAsdA AS1311 1421 dTdTdAdTaGfaGfcAfaGfaAfcAfcAfgUfusUfsu 0.10 0.30 0.66 D1312 S1312 330 AfacaGfuguUfcUfuGfdCUdCUdAudAsa AS1312 1422 dTUdAUdAgdAGfcAfaGfaAfcAfcUfgUfusUfsu 0.09 0.13 0.48 D1313 S1313 331 AfAfCfaGfuGfuucUfuGfcUfcUfaUfasAf AS1313 1423 uUfaUfaGfaGfcAfaGfAfAfcAfcUfguusUfsu 0.14 0.38 0.74 D1314 S1314 332 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1314 1424 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.19 0.54 D1315 S1315 333 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1315 1425 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.15 0.55 D1316 S1316 334 AfaCfaGfuGfuUfcUfuGfcUfcUfauasAf AS1316 1426 uUfAfUfaGfaGfcAfaGfaAfcAfcUfgUfususu 0.07 0.16 0.53 D1317 S1317 335 AfacaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1317 1427 uUfaUfaGfaGfcAfaGfaAfcAfcUfGfUfususu 0.07 0.16 0.55 D1318 S1318 336 AfAfCfaGfuguUfcUfuGfcUfcUfaUfasAf AS1318 1428 uUfaUfaGfaGfcAfaGfaAfCfAfcUfguusUfsu 0.10 0.32 0.61 D1319 S1319 337 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1319 1429 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfususu 0.08 0.16 0.53 D1320 S1320 338 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1320 1430 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfususu 0.08 0.16 0.61 D1321 S1321 339 AfaCfaGfuGfuUfcUfuGfcUfCfUfaUfasAf AS1321 1431 uUfaUfagaGfcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.14 0.58 D1322 S1322 340 AfaCfaGfuGfuUfcuuGfcUfcUfaUfasAf AS1322 1432 uUfaUfaGfaGfcAfAfGfaAfcAfcUfgUfusUfsu 0.15 0.49 0.84 D1323 S1323 341 AfaCfaGfuGfuUfcUfuGfcUfcuaUfasAf AS1323 1433 uUfaUfAfGfaGfcAfaGfaAfcAfcUfgUfususu 0.07 0.20 0.62 D1324 S1324 342 AfAfCfaGfuGfuUfcUfuGfcUfcUfaUfasAf AS1324 1434 uUfaUfaGfaGfcAfaGfaAfcAfcUfguusUfsu 0.08 0.25 0.78 D1325 S1325 343 AfAfCfaGfuGfuUkUfuGfcUfcUfaUfasAf AS1325 1435 uUfaUfaGfaGfcAfaGfaAfcAfcUfguusUfsu 0.08 0.18 0.80 D1326 S1326 344 AfaCfaGfuGfuUfcUfuGfcUfcUfAfUfasAf AS1326 1436 uUfauaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.21 0.66 D1327 S1327 345 AfaCfaGfuGfuucUfuGfcUfcUfaUfasAf AS1327 1437 uUfaUfaGfaGfcAfaGfAfAfcAfcUfgUfusUfsu 0.10 0.31 0.70 D1328 S1328 346 AfAfCfaGfuGfuUfcUfuGfcUfcUfauasAf AS1328 1438 uUfAfUfaGfaGfcAfaGfaAfcAfcUfguusUfsu 0.07 0.15 0.55 D1329 S1329 347 AfaCfAfGfuGfuUfcUfuGfcUfcUfaUfasAf AS1329 1439 uUfaUfaGfaGfcAfaGfaAfcAfcugUfusUfsu 0.08 0.19 0.71 D1330 S1330 348 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfAfsAf AS1330 1440 uuaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.09 0.27 0.76 D1331 S1331 349 AfaCfaGfuguUfcUfuGfcUfcUfaUfasAf AS1331 1441
uUfaUfaGfaGfcAfaGfaAfCfAfcUfgUfusUfsu 0.07 0.21 0.65 D1332 S1332 350 AfAfCfaGfuGfuUfcUfuGfcUfcuaUfasAf AS1332 1442 uUfaUfAfGfaGfcAfaGfaAfcAfcUfguusUfsu 0.07 0.17 0.53 D1333 S1333 351 AfaCfaGfUfGfuUfcUfuGfcUfcUfaUfasAf AS1333 1443 uUfaUfaGfaGfcAfaGfaAfcacUfgUfusUfsu 0.08 0.25 0.73 D1334 S1334 352 AfaCfaguGfuUfcUfuGfcUfcUfaUfasAf AS1334 1444 uUfaUfaGfaGfcAfaGfaAfcAfCfUfgUfusUfsu 0.07 0.18 0.54 D1335 S1335 353 AfaCfaGfuGfuUfcuuGfcUfcUfaUfasAf AS1335 1445 uUfaUfaGfaGfcAfAfGfaAfcAfcUfgUfususu 0.14 0.38 0.57 D1336 S1336 354 AfaCfaGfuGfUfUfcUfuGfcUfcUfaUfasAf AS1336 1446 uUfaUfaGfaGfcAfaGfaacAfcUfgUfusUfsu 0.16 0.50 0.96 D1337 S1337 355 AfaCfaGfuGfuUfcUfuGfcUfcUfauasAf AS1337 1447 uUfAfUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.08 0.19 0.54 D1338 S1338 356 AfAfCfaGfuGfuUfcUfugcUfcUfaUfasAf AS1338 1448 uUfaUfaGfaGfCfAfaGfaAfcAfcUfguusUfsu 0.08 0.20 0.69 D1339 S1339 357 AfaCfaGfuGfuUfCfUfuGfcUfcUfaUfasAf AS1339 1449 uUfaUfaGfaGfcAfagaAfcAfcUfgUfusUfsu 0.07 0.16 0.55 D1340 S1340 358 AfaCfaGfuGfuUfcUfuGfcUfcuaUfasAf AS1340 1450 uUfaUfAfGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.08 0.17 0.57 D1341 S1341 359 AfaCfaGfuguUfcUfuGfcUfcUfaUfasAf AS1341 1451 uUfaUfaGfaGfcAfaGfaAfCfAfcUfgUfususu 0.08 0.22 0.63 D1342 S1342 360 AfAfCfaGfuGfuUfcuuGfcUfcUfaUfasAf AS1342 1452 uUfaUfaGfaGfcAfAfGfaAfcAfcUfguusUfsu 0.21 0.56 0.86 D1343 S1343 361 AfacaGfuGfUfUfcUfuGfcUfcUfaUfasAf AS1343 1453 uUfaUfaGfaGfcAfaGfaacAfcUfGfUfusUfsu 0.14 0.37 0.73 D1344 S1344 362 AfaCfaGfuGfuucUfUfGfcUfcUfaUfasAf AS1344 1454 uUfaUfaGfaGfcaaGfAfAfcAfcUfgUfusUfsu 0.08 0.20 0.66 D1345 S1345 363 AfaCfAfGfuGfuUfcuuGfcUfcUfaUfasAf AS1345 1455 uUfaUfaGfaGfcAfAfGfaAfcAfcugUfusUfsu 0.12 0.34 0.73 D1346 S1346 364 AfaCfaGfuGfUfUfcUfuGfcUfcUfauasAf AS1346 1456 uUfAfUfaGfaGfcAfaGfaacAfcUfgUfusUfsu 0.16 0.42 0.90 D1347 S1347 365 AfaCfaGfuGfUfUfcUfuGfcUfcUfaUfasAf AS1347 1457 uUfaUfaGfaGfcAfaGfaacAfcUfgUfusUfsUf 0.17 0.43 0.85 D1348 S1348 366 AfaCfAfGfuGfuucUfuGfcUfcUfaUfasAf AS1348 1458 uUfaUfaGfaGfcAfaGfAfAfcAfcugUfusUfsu 0.08 0.21 0.58 D1349 S1349 367 AfaCfaGfuGfUfUfcUfuGfcUfcuaUfasAf AS1349 1459 uUfaUfAfGfaGfcAfaGfaacAfcUfgUfusUfsu 0.21 0.39 0.88 D1350 S1350 368 AfaCfaguGfuUfcUfUfGfcUfcUfaUfasAf AS1350 1460 uUfaUfaGfaGfcaaGfaAfcAfCfUfgUfusUfsu 0.06 0.13 0.52 D1351 S1351 369 AfaCfAfGfuguUfcUfuGfcUfcUfaUfasAf AS1351 1461 uUfaUfaGfaGfcAfaGfaAfCfAfcugUfusUfsu 0.08 0.21 0.58 D1352 S1352 370 AfaCfaGfUfGfuUfcuuGfcUfcUfaUfasAf AS1352 1462 uUfaUfaGfaGfcAfAfGfaAfcacUfgUfusUfsu 0.18 0.49 0.84 D1353 S1353 371 AfaCfaGfuGfUfUfcUfuGfcucUfaUfasAf AS1353 1463 uUfaUfaGfAfGfcAfaGfaacAfcUfgUfusUfsu 0.11 0.25 0.68 D1354 S1354 372 AfacaGfuGfuUfcUfUfGfcUfcUfaUfasAf AS1354 1464 uUfaUfaGfaGfcaaGfaAfcAfcUfGfUfusUfsu 0.07 0.15 0.52 D1355 S1355 373 AfaCfaGfUfGfuucUfuGfcUfcUfaUfasAf AS1355 1465 uUfaUfaGfaGfcAfaGfAfAfcacUfgUfusUfsu 0.10 0.26 0.63 D1356 S1356 374 AfaCfaGfuGfUfUfcUfugcUfcUfaUfasAf AS1356 1466 uUfaUfaGfaGfCfAfaGfaacAfcUfgUfusUfsu 0.16 0.33 0.79 D1357 S1357 375 AfaCfAfGfuGfuUfcUfuGfcUfcUfaUfasAf AS1357 1467 uUfaUfaGfaGfcAfaGfaAfcAfcugUfusUfsUf 0.09 0.19 0.51 D1358 S1358 376 AfaCfaGfuGfUfUfcuuGfcUfcUfaUfasAf AS1358 1468 uUfaUfaGfaGfcAfAfGfaacAfcUfgUfusUfsu 0.22 0.48 0.71 D1359 S1359 377 AfaCfaGfuGfuUfcUfUfGfcUfcUfaUfasAf AS1359 1469 uUfaUfaGfaGfcaaGfaAfcAfcUfgUfusUfsUf 0.10 0.17 0.61 D1360 S1360 378 AfaCfaguGfUfUfcUfuGfcUfcUfaUfasAf AS1360 1470 uUfaUfaGfaGfcAfaGfaacAfCfUfgUfusUfsu 0.14 0.40 0.87 D1361 S1361 379 AfaCfaGfuGfuUfcUfUfGfcUfcuaUfasAf AS1361 1471 uUfaUfAfGfaGfcaaGfaAfcAfcUfgUfusUfsu 0.07 0.14 0.52 D1362 S1362 380 aaCfaGfuGfuUfcUfuGfCfUfcUfaUfasAf AS1362 1472 uUfaUfaGfagcAfaGfaAfcAfcUfgUfUfsUfsu 0.10 0.28 0.81 D1363 S1363 381 AfaCfaGfuGfuucUfuGfcUfcUfAfUfasAf AS1363 1473 uUfauaGfaGfcAfaGfAfAfcAfcUfgUfusUfsu 0.06 0.16 0.68 D1364 S1364 382 AfaCfaGfuGfuUfcUfugcUfcUfaUfAfsAf AS1364 1474 uuaUfaGfaGfCfAfaGfaAfcAfcUfgUfusUfsu 0.09 0.26 0.67 D1365 S1365 383 aacaguguucuugcucuauasa AS1365 1475 uUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.20 0.59 0.95 D1366 S1366 384 AfaCfaGfuGfuUfcUfuGfCfUfcUfauasAf AS1366 1476 uUfAfUfaGfagcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.13 0.53 D1367 S1367 385 AfaCfaGfuGfuUfcUfuGfCfUfcUfaUfasAf AS1367 1477 uUfaUfaGfagcAfaGfaAfcAfcUfgUfusUfsUf 0.08 0.16 0.53 D1368 S1368 386 AfaCfaGfuguUfcUfuGfcUfcUfAfUfasAf AS1368 1478 uUfauaGfaGfcAfaGfaAfCfAfcUfgUfusUfsu 0.07 0.15 0.54 D1369 S1369 387 AfaCfaGfuGfuUfcuuGfcUfcUfaUfAfsAf AS1369 1479 uuaUfaGfaGfcAfAfGfaAfcAfcUfgUfusUfsu 0.23 0.56 0.89 D1370 S1370 388 AfaCfaGfuGfuUfcUfuGfCfUfcuaUfasAf AS1370 1480 uUfaUfAfGfagcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.12 0.55 D1371 S1371 389 AfaCfaGfuGfuUfcUfuGfCfUfcuaUfasAf AS1371 1481 uUfaUfAfGfagcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.18 0.58 D1372 S1372 390 AfaCfaguGfuUfcUfuGfcUfcUfAfUfasAf AS1372 1482 uUfauaGfaGfcAfaGfaAfcAfCfUfgUfusUfsu 0.06 0.15 0.56 D1373 S1373 391 AfaCfaGfuGfuucUfuGfcUfcUfaUfAfsAf AS1373 1483 uuaUfaGfaGfcAfaGfAfAfcAfcUfgUfusUfsu 0.21 0.51 0.89 D1374 S1374 392 AfacaGfuguUfcUfuGfcUfcUfaUfasAf AS1374 1484 uUfaUfaGfaGfcAfaGfaAfCfAfcUfGfUfusUfsu 0.08 0.21 0.64 D1375 S1375 393 AfaCfaGfuGfuUfcuuGfCfUfcUfaUfasAf AS1375 1485 uUfaUfaGfagcAfAfGfaAfcAfcUfgUfusUfsu 0.15 0.40 0.94 D1376 S1376 394 AfaCfaGfuGfuUfcuuGfCfUfcUfaUfasAf AS1376 1486 uUfaUfaGfagcAfAfGfaAfcAfcUfgUfusUfsu 0.13 0.40 0.96 D1377 S1377 395 AfaCfaGfuGfuUfcUfuGfcUfCfUfauasAf AS1377 1487 uUfAfUfagaGfcAfaGfaAfcAfcUfgUfusUfsu 0.08 0.17 0.64 D1378 S1378 396 AfaCfaGfuguUfcUfuGfcUfcUfaUfAfsAf AS1378 1488 uuaUfaGfaGfcAfaGfaAfCfAfcUfgUfusUfsu 0.18 0.50 0.97 D1379 S1379 397 AfaCfaGfuGfuucUfuGfCfUfcUfaUfasAf AS1379 1489 uUfaUfaGfagcAfaGfAfAfcAfcUfgUfusUfsu 0.08 0.24 0.79 D1380 S1380 398 aaCfaGfuGfuUfcUfuGfcUfcUfAfUfasAf AS1380 1490 uUfauaGfaGfcAfaGfaAfcAfcUfgUfUfsUfsu 0.07 0.14 0.58 D1381 S1381 399 AfaCfaguGfuUfcUfuGfcUfcUfaUfAfsAf AS1381 1491 uuaUfaGfaGfcAfaGfaAfcAfCfUfgUfusUfsu 0.11 0.34 0.96 D1382 S1382 400 AfaCfaGfuguUfcUfuGfCfUfcUfaUfasAf AS1382 1492 uUfaUfaGfagcAfaGfaAfCfAfcUfgUfusUfsu 0.08 0.18 0.69 D1383 S1383 401 AfaCfaGfuGfuUfcuuGfcUfCfUfaUfasAf AS1383 1493 uUfaUfagaGfcAfAfGfaAfcAfcUfgUfusUfsu 0.14 0.38 0.85 D1384 S1384 402 AfaCfaGfuGfuUfcUfuGfcUfcUfAfUfasAf AS1384 1494 uUfauaGfaGfcAfaGfaAfcAfcUfgUfusUfsUf 0.07 0.16 0.54 D1385 S1385 403 AfacaGfuGfuUfcUfuGfcUfcUfaUfAfsAf AS1385 1495 uuaUfaGfaGfcAfaGfaAfcAfcUfGfUfusUfsu 0.08 0.20 0.75 D1386 S1386 404 aacaguguucUfuGfcUcUaudAsa AS1386 1496 uUfdAUdAGfaGfcAfaGfaadCadCudGdTdTsusu 0.25 0.56 0.90 D1387 S1387 405 AfaCfaguGfuUfcUfuGfCfUfcUfaUfasAf AS1387 1497 uUfaUfaGfagcAfaGfaAfcAfCfUfgUfusUfsu 0.08 0.19 0.70 D1388 S1388 406 AfaCfaGfuGfuucUfuGfcUfCfUfaUfasAf AS1388 1498 uUfaUfagaGfcAfaGfAfAfcAfcUfgUfusUfsu 0.08 0.14 0.60 D1389 S1389 407 AfaCfaGfuGfuUfcUfuGfcUfcuaUfAfsAf AS1389 1499 uuaUfAfGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.08 0.19 0.62 D1390 S1390 408 aaCfaGfuGfuUfcUfuGfcUfcUfaUfAfsAf AS1390 1500 uuaUfaGfaGfcAfaGfaAfcAfcUfgUfUfsUfsu 0.08 0.27 0.76 D1391 S1391 409 aacaguguucdTudGcdTcdTadTasa AS1391 1501 uUfdAUdAGfaGfcAfaGfaadCadCudGudTsusu 0.18 0.36 0.81 D1392 S1392 410 AfacaGfuGfuUfcUfuGfCfUfcUfaUfasAf AS1392 1502 uUfaUfaGfagcAfaGfaAfcAfcUfGfUfusUfsu 0.07 0.17 0.55 D1393 S1393 411 AfaCfaGfuguUfcUfuGfcUfCfUfaUfasAf AS1393 1503 uUfaUfagaGfcAfaGfaAfCfAfcUfgUfusUfsu 0.07 0.15 0.57 D1394 S1394 412 AfaCfaGfuGfuUfcuuGfcUfcUfAfUfasAf AS1394 1504 uUfauaGfaGfcAfAfGfaAfcAfcUfgUfusUfsu 0.26 0.68 1.06 D1395 S1395 413 AfaCfaGfuGfuUfcUfuGfcucUfaUfAfsAf AS1395 1505 uuaUfaGfAfGfcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.18 0.58 D1396 S1396 414 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfAfsAf AS1396 1506 uuaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsUf 0.09 0.27 0.73 D1397 S1397 415 AfaCfaAfuGfuUfcUfuGfcdAdCdTdAUfasAf AS1397 1507 uUfadTdAdGdAGfcAfaGfaGfcAfcAfgUfusUfsu 0.20 0.51 0.73 D1398 S1398 416 AfacaGfuguUfcuuGfcucUfauasAf AS1398 1508 uUfAfUfaGfAfGfcAfAfGfaAfCfAfcUfGfUfusUfsu 0.13 0.34 0.86 D1399 S1399 417 dAacadGugudTcuudGcucdTauasdA AS1399 1509 udTdAdTadGdAdGcdAdAdGadAdCdAcdTdGdTusdTsu 0.24 0.42 0.82 D1400 S1400 418 AfaCfaAfuGfuUfcUfuGfdCdAdCdTaUfasAf AS1400 1510 uUfaUfdAdGdAdGcAfaGfaGfcAfcAfgUfusUfsu 0.49 0.85 0.78 D1401 S1401 419 AfaCfaAfuGfuUfcUfudGdCdAdCUfaUfasAf AS1401 1511 uUfaUfadGdAdGdCAfaGfaGfcAfcAfgUfusUfsu 0.67 0.83 0.85 D1402 S1402 420 aaCfAfguGfUfucUfUfgcUfCfuaUfAfsa AS1402 1512 uUfaUfAfgaGfCfaaGfAfacAfCfugUfUfsusu 0.18 0.47 0.80 D1403 S1403 421 AfaCfaAfuGfuUfcUfuGfcdAdCUfadTdAsAf AS1403 1513 udTdAUfadGdAGfcAfaGfaGfcAfcAfgUfusUfsu 0.73 0.89 0.77 D1404 S1404 422 aacAgugUucuUgcuCuauAsa AS1404 1514 uUaUAgAGCaAGAaCACuGUUsusu 0.12 0.39 0.79 D1405 S1405 423 AacaGuguUcuuGcucUauasA AS1405 1515 uUAUaGAGcAAGaACAcUGUusUsu 0.12 0.37 0.77 D1406 S1406 424 AfaCfaAfuGfuUfcUfudGdCAfcUfadTdAsAf AS1406 1516 udTdAUfaGfadGdCAfaGfaGfcAfcAfgUfusUfsu 0.59 0.93 0.89 D1407 S1407 425 aACagUGuuCUugCUcuAUasa AS1407 1517 UUauAGagCAagAAcaCUguUsUsu 0.09 0.16 0.55 D1408 S1408 426 AfaCfaAfuGfuUfcUfuGfcAfcdTdAdTdAsAf AS1408 1518 udTdAdTdAGfaGfcAfaGfaAfcAfcAfgUfusUfsu 0.22 0.64 0.86 D1409 S1409 427 aaCAguGUucUUgcUCuaUAsa AS1409 1519 uUaUAgaGCaaGAacACugUUsusu 0.13 0.31 0.76 D1410 S1410 428 AfaCfaAfuGfuUfcUfuGfcAfdCdTdAdTdAsAf AS1410 1520 udTdAdTdAdGaGfcAfaGfaGfcAfcAfgUfusUfsu 0.77 0.94 0.93 D1411 S1411 429 aacAfgugUfucuUfgcuCfuauAfsa AS1411 1521 uUfaUfAfgAfGfCfaAfGfAfaCfAfCfuGfUfUfsusu 0.23 0.53 1.04 D1412 S1412 430 aacdAgugdTucudTgcudCuaudAsa AS1412 1522 udTadTdAgdAdGdCadAdGdAadCdAdCudGdTdTsusu 0.30 0.64 0.90 D1413 S1413 431 AfaCfaGfuGfuUfcUfuGfcUfcUfaUfasa AS1413 1523 UfUfaUfaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.09 0.19 0.63 D1414 S1414 432 AfaCfaGfuGfUfUfcUfuGfcUfcUfaUfasa AS1414 1524 UfUfaUfaGfaGfcAfaGfaacAfcUfgUfusUfsu 0.11 0.28 0.66
D1415 S1415 433 AfaCfaGfuGfuUfcUfuGfCfUfcUfaUfasa AS1415 1525 UfUfaUfaGfagcAfaGfaAfcAfcUfgUfusUfsu 0.06 0.13 0.53 D1416 S1416 434 aacaguguucuugcucuauasa AS1416 1526 UfUfAfUfAfGfAfGfCfAfAfGfAfAfCfAfCfUfGfUfUfsusu 0.20 0.53 0.99 D1417 S1417 435 AfaCfaGfuGfuUfcUfuGfcUfcUfAfUfasa AS1417 1527 UfUfauaGfaGfcAfaGfaAfcAfcUfgUfusUfsu 0.07 0.17 0.53 D1418 S1418 436 aAfCfagUfGfuuCfUfugCfUfcuAfUfasa AS1418 1528 UfUfauAfGfagCfAfagAfAfcaCfUfguUfsUfsu 0.08 0.20 0.70 D1419 S1419 437 AfaCfAfGfuGfuUfcUfuGfcUfcUfaUfasAf AS1419 1529 uUfaUfaGfaGfcAfaGfaAfcAfcugUfusUfsUf 0.08 0.20 0.70 D1420 S1420 438 GfaCfuUfcUfcCfUfCfcAfgugGfaCfcUfL96 AS1420 1530 aGfgUfcCfAfCfuGfgagGfaGfaAfgUfcsCfsc D1421 S1421 439 GfaCfuUfcUfcCfUfCfcAfgUfGfGfaCfcUfL96 AS1421 1531 aGfgUfccaCfuGfgagGfaGfaAfgUfcsCfsc D1422 S1422 440 AfcUfuCfuCfcUfCfCfaGfuggAfcCfuGfL96 AS1422 1532 cAfgGfuCfCfAfcUfggaGfgAfgAfaGfusCfsc D1423 S1423 441 AfcUfuCfuCfcUfCfCfaGfuGfGfAfcCfuGfL96 AS1423 1533 cAfgGfuccAfcUfggaGfgAfgAfaGfusCfsc D1424 S1424 442 CfuUfcUfcCfuCfCfAfgUfggaCfcUfgAfL96 AS1424 1534 uCfaGfgUfCfCfaCfuggAfgGfaGfaAfgsUfsc D1425 S1425 443 CfuUfcUfcCfuCfCfAfgUfgGfAfCfcUfgAfL96 AS1425 1535 uCfaGfgucCfaCfuggAfgGfaGfaAfgsUfsc D1426 S1426 444 UfuCfuCfcUfcCfAfGfuGfgacCfuGfaAfL96 AS1426 1536 uUfcAfgGfUfCfcAfcugGfaGfgAfgAfasGfsu D1427 S1427 445 UfuCfuCfcUfcCfAfGfuGfgAfCfCfuGfaAfL96 AS1427 1537 uUfcAfgguCfcAfcugGfaGfgAfgAfasGfsu D1428 S1428 446 UfcUfcCfuCfcAfGfUfgGfaccUfgAfaGfL96 AS1428 1538 cUfuCfaGfGfUfcCfacuGfgAfgGfaGfasAfsg D1429 S1429 447 UfcUfcCfuCfcAfGfUfgGfaCfCfUfgAfaGfL96 AS1429 1539 cUfuCfaggUfcCfacuGfgAfgGfaGfasAfsg D1430 S1430 448 CfuCfcUfcCfaGfUfGfgAfccuGfaAfgGfL96 AS1430 1540 cCfuUfcAfGfGfuCfcacUfgGfaGfgAfgsAfsa D1431 S1431 449 CfuCfcUfcCfaGfUfGfgAfcCfUfGfaAfgGfL96 AS1431 1541 cCfuUfcagGfuCfcacUfgGfaGfgAfgsAfsa D1432 S1432 450 UfcCfuCfcAfgUfGfGfaCfcugAfaGfgAfL96 AS1432 1542 uCfcUfuCfAfGfgUfccaCfuGfgAfgGfasGfsa D1433 S1433 451 UfcCfuCfcAfgUfGfGfaCfcUfGfAfaGfgAfL96 AS1433 1543 uCfcUfucaGfgUfccaCfuGfgAfgGfasGfsa D1434 S1434 452 CfcUfcCfaGfuGfGfAfcCfugaAfgGfaCfL96 AS1434 1544 gUfcCfuUfCfAfgGfuccAfcUfgGfaGfgsAfsg D1435 S1435 453 CfcUfcCfaGfuGfGfAfcCfuGfAfAfgGfaCfL96 AS1435 1545 gUfcCfuucAfgGfuccAfcUfgGfaGfgsAfsg D1436 S1436 454 CfuCfcAfgUfgGfAfCfcUfgaaGfgAfcGfL96 AS1436 1546 cGfuCfcUfUfCfaGfgucCfaCfuGfgAfgsGfsa D1437 S1437 455 CfuCfcAfgUfgGfAfCfcUfgAfAfGfgAfcGfL96 AS1437 1547 cGfuCfcuuCfaGfgucCfaCfuGfgAfgsGfsa D1438 S1438 456 UfcCfaGfuGfgAfCfCfuGfaagGfaCfgAfL96 AS1438 1548 uCfgUfcCfUfUfcAfgguCfcAfcUfgGfasGfsg D1439 S1439 457 UfcCfaGfuGfgAfCfCfuGfaAfGfGfaCfgAfL96 AS1439 1549 uCfgUfccuUfcAfgguCfcAfcUfgGfasGfsg D1440 S1440 458 CfcAfgUfgGfaCfCfUfgAfaggAfcGfaGfL96 AS1440 1550 cUfcGfuCfCfUfuCfaggUfcCfaCfuGfgsAfsg D1441 S1441 459 CfcAfgUfgGfaCfCfUfgAfaGfGfAfcGfaGfL96 AS1441 1551 cUfcGfuccUfuCfaggUfcCfaCfuGfgsAfsg D1442 S1442 460 CfaGfuGfgAfcCfUfGfaAfggaCfgAfgGfL96 AS1442 1552 cCfuCfgUfCfCfuUfcagGfuCfcAfcUfgsGfsa D1443 S1443 461 CfaGfuGfgAfcCfUfGfaAfgGfAfCfgAfgGfL96 AS1443 1553 cCfuCfgucCfuUfcagGfuCfcAfcUfgsGfsa D1444 S1444 462 AfgUfgGfaCfcUfGfAfaGfgacGfaGfgGfL96 AS1444 1554 cCfcUfcGfUfCfcUfucaGfgUfcCfaCfusGfsg D1445 S1445 463 AfgUfgGfaCfcUfGfAfaGfgAfCfGfaGfgGfL96 AS1445 1555 cCfcUfcguCfcUfucaGfgUfcCfaCfusGfsg D1446 S1446 464 GfuGfgAfcCfuGfAfAfgGfacgAfgGfgAfL96 AS1446 1556 uCfcCfuCfGfUfcCfuucAfgGfuCfcAfcsUfsg D1447 S1447 465 GfuGfgAfcCfuGfAfAfgGfaCfGfAfgGfgAfL96 AS1447 1557 uCfcCfucgUfcCfuucAfgGfuCfcAfcsUfsg D1448 S1448 466 UfgGfaCfcUfgAfAfGfgAfcgaGfgGfaUfL96 AS1448 1558 aUfcCfcUfCfGfuCfcuuCfaGfgUfcCfasCfsu D1449 S1449 467 UfgGfaCfcUfgAfAfGfgAfcGfAfGfgGfaUfL96 AS1449 1559 aUfcCfcucGfuCfcuuCfaGfgUfcCfasCfsu D1450 S1450 468 GfgAfcCfuGfaAfGfGfaCfgagGfgAfuGfL96 AS1450 1560 cAfuCfcCfUfCfgUfccuUfcAfgGfuCfcsAfsc D1451 S1451 469 GfgAfcCfuGfaAfGfGfaCfgAfGfGfgAfuGfL96 AS1451 1561 cAfuCfccuCfgUfccuUfcAfgGfuCfcsAfsc D1452 S1452 470 GfaCfcUfgAfaGfGfAfcGfaggGfaUfgGfL96 AS1452 1562 cCfaUfcCfCfUfcGfuccUfuCfaGfgUfcsCfsa D1453 S1453 471 GfaCfcUfgAfaGfGfAfcGfaGfGfGfaUfgGfL96 AS1453 1563 cCfaUfcccUfcGfuccUfuCfaGfgUfcsCfsa D1454 S1454 472 AfcCfuGfaAfgGfAfCfgAfgggAfuGfgGfL96 AS1454 1564 cCfcAfuCfCfCfuCfgucCfuUfcAfgGfusCfsc D1455 S1455 473 AfcCfuGfaAfgGfAfCfgAfgGfGfAfuGfgGfL96 AS1455 1565 cCfcAfuccCfuCfgucCfuUfcAfgGfusCfsc D1456 S1456 474 CfcUfgAfaGfgAfCfGfaGfggaUfgGfgAfL96 AS1456 1566 uCfcCfaUfCfCfcUfcguCfcUfuCfaGfgsUfsc D1457 S1457 475 CfcUfgAfaGfgAfCfGfaGfgGfAfUfgGfgAfL96 AS1457 1567 uCfcCfaucCfcUfcguCfcUfuCfaGfgsUfsc D1458 S1458 476 CfuGfaAfgGfaCfGfAfgGfgauGfgGfaUfL96 AS1458 1568 aUfcCfcAfUfCfcCfucgUfcCfuUfcAfgsGfsu D1459 S1459 477 CfuGfaAfgGfaCfGfAfgGfgAfUfGfgGfaUfL96 AS1459 1569 aUfcCfcauCfcCfucgUfcCfuUfcAfgsGfsu D1460 S1460 478 UfgAfaGfgAfcGfAfGfgGfaugGfgAfuUfL96 AS1460 1570 aAfuCfcCfAfUfcCfcucGfuCfcUfuCfasGfsg D1461 S1461 479 UfgAfaGfgAfcGfAfGfgGfaUfGfGfgAfuUfL96 AS1461 1571 aAfuCfccaUfcCfcucGfuCfcUfuCfasGfsg D1462 S1462 480 GfaAfgGfaCfgAfGfGfgAfuggGfaUfuUfL96 AS1462 1572 aAfaUfcCfCfAfuCfccuCfgUfcCfuUfcsAfsg D1463 S1463 481 GfaAfgGfaCfgAfGfGfgAfuGfGfGfaUfuUfL96 AS1463 1573 aAfaUfcccAfuCfccuCfgUfcCfuUfcsAfsg D1464 S1464 482 AfaGfgAfcGfaGfGfGfaUfgggAfuUfuCfL96 AS1464 1574 gAfaAfuCfCfCfaUfcccUfcGfuCfcUfusCfsa D1465 S1465 483 AfaGfgAfcGfaGfGfGfaUfgGfGfAfuUfuCfL96 AS1465 1575 gAfaAfuccCfaUfcccUfcGfuCfcUfusCfsa D1466 S1466 484 AfgGfaCfgAfgGfGfAfuGfggaUfuUfcAfL96 AS1466 1576 uGfaAfaUfCfCfcAfuccCfuCfgUfcCfusUfsc D1467 S1467 485 AfgGfaCfgAfgGfGfAfuGfgGfAfUfuUfcAfL96 AS1467 1577 uGfaAfaucCfcAfuccCfuCfgUfcCfusUfsc D1468 S1468 486 GfgAfcGfaGfgGfAfUfgGfgauUfuCfaUfL96 AS1468 1578 aUfgAfaAfUfCfcCfaucCfcUfcGfuCfcsUfsu D1469 S1469 487 GfgAfcGfaGfgGfAfUfgGfgAfUfUfuCfaUfL96 AS1469 1579 aUfgAfaauCfcCfaucCfcUfcGfuCfcsUfsu D1470 S1470 488 GfaCfgAfgGfgAfUfGfgGfauuUfcAfuGfL96 AS1470 1580 cAfuGfaAfAfUfcCfcauCfcCfuCfgUfcsCfsu D1471 S1471 489 GfaCfgAfgGfgAfUfGfgGfaUfUfUfcAfuGfL96 AS1471 1581 cAfuGfaaaUfcCfcauCfcCfuCfgUfcsCfsu D1472 S1472 490 AfcGfaGfgGfaUfGfGfgAfuuuCfaUfgUfL96 AS1472 1582 aCfaUfgAfAfAfuCfccaUfcCfcUfcGfusCfsc D1473 S1473 491 AfcGfaGfgGfaUfGfGfgAfuUfUfCfaUfgUfL96 AS1473 1583 aCfaUfgaaAfuCfccaUfcCfcUfcGfusCfsc D1474 S1474 492 CfgAfgGfgAfuGfGfGfaUfuucAfuGfuAfL96 AS1474 1584 uAfcAfuGfAfAfaUfcccAfuCfcCfuCfgsUfsc D1475 S1475 493 CfgAfgGfgAfuGfGfGfaUfuUfCfAfuGfuAfL96 AS1475 1585 uAfcAfugaAfaUfcccAfuCfcCfuCfgsUfsc D1476 S1476 494 GfaGfgGfaUfgGfGfAfuUfucaUfgUfaAfL96 AS1476 1586 uUfaCfaUfGfAfaAfuccCfaUfcCfcUfcsGfsu D1477 S1477 495 GfaGfgGfaUfgGfGfAfuUfuCfAfUfgUfaAfL96 AS1477 1587 uUfaCfaugAfaAfuccCfaUfcCfcUfcsGfsu D1478 S1478 496 AfgGfgAfuGfgGfAfUfuUfcauGfuAfaCfL96 AS1478 1588 gUfuAfcAfUfGfaAfaucCfcAfuCfcCfusCfsg D1479 S1479 497 AfgGfgAfuGfgGfAfUfuUfcAfUfGfuAfaCfL96 AS1479 1589 gUfuAfcauGfaAfaucCfcAfuCfcCfusCfsg D1480 S1480 498 GfgGfaUfgGfgAfUfUfuCfaugUfaAfcCfL96 AS1480 1590 gGfuUfaCfAfUfgAfaauCfcCfaUfcCfcsUfsc D1481 S1481 499 GfgGfaUfgGfgAfUfUfuCfaUfGfUfaAfcCfL96 AS1481 1591 gGfuUfacaUfgAfaauCfcCfaUfcCfcsUfsc D1482 S1482 500 GfgAfuGfgGfaUfUfUfcAfuguAfaCfcAfL96 AS1482 1592 uGfgUfuAfCfAfuGfaaaUfcCfcAfuCfcsCfsu D1483 S1483 501 GfgAfuGfgGfaUfUfUfcAfuGfUfAfaCfcAfL96 AS1483 1593 uGfgUfuacAfuGfaaaUfcCfcAfuCfcsCfsu D1484 S1484 502 GfaUfgGfgAfuUfUfCfaUfguaAfcCfaAfL96 AS1484 1594 uUfgGfuUfAfCfaUfgaaAfuCfcCfaUfcsCfsc D1485 S1485 503 GfaUfgGfgAfuUfUfCfaUfgUfAfAfcCfaAfL96 AS1485 1595 uUfgGfuuaCfaUfgaaAfuCfcCfaUfcsCfsc D1486 S1486 504 AfuGfgGfaUfuUfCfAfuGfuaaCfcAfaGfL96 AS1486 1596 cUfuGfgUfUfAfcAfugaAfaUfcCfcAfusCfsc D1487 S1487 505 AfuGfgGfaUfuUfCfAfuGfuAfAfCfcAfaGfL96 AS1487 1597 cUfuGfguuAfcAfugaAfaUfcCfcAfusCfsc D1488 S1488 506 UfgGfgAfuUfuCfAfUfgUfaacCfaAfgAfL96 AS1488 1598 uCfuUfgGfUfUfaCfaugAfaAfuCfcCfasUfsc D1489 S1489 507 UfgGfgAfuUfuCfAfUfgUfaAfCfCfaAfgAfL96 AS1489 1599 uCfuUfgguUfaCfaugAfaAfuCfcCfasUfsc D1490 S1490 508 GfgGfaUfuUfcAfUfGfuAfaccAfaGfaGfL96 AS1490 1600 cUfcUfuGfGfUfuAfcauGfaAfaUfcCfcsAfsu D1491 S1491 509 GfgGfaUfuUfcAfUfGfuAfaCfCfAfaGfaGfL96 AS1491 1601 cUfcUfuggUfuAfcauGfaAfaUfcCfcsAfsu D1492 S1492 510 GfgAfuUfuCfaUfGfUfaAfccaAfgAfgUfL96 AS1492 1602 aCfuCfuUfGfGfuUfacaUfgAfaAfuCfcsCfsa D1493 S1493 511 GfgAfuUfuCfaUfGfUfaAfcCfAfAfgAfgUfL96 AS1493 1603 aCfuCfuugGfuUfacaUfgAfaAfuCfcsCfsa D1494 S1494 512 GfaUfuUfcAfuGfUfAfaCfcaaGfaGfuAfL96 AS1494 1604 uAfcUfcUfUfGfgUfuacAfuGfaAfaUfcsCfsc D1495 S1495 513 GfaUfuUfcAfuGfUfAfaCfcAfAfGfaGfuAfL96 AS1495 1605 uAfcUfcuuGfgUfuacAfuGfaAfaUfcsCfsc D1496 S1496 514 AfuUfuCfaUfgUfAfAfcCfaagAfgUfaUfL96 AS1496 1606 aUfaCfuCfUfUfgGfuuaCfaUfgAfaAfusCfsc D1497 S1497 515 AfuUfuCfaUfgUfAfAfcCfaAfGfAfgUfaUfL96 AS1497 1607 aUfaCfucuUfgGfuuaCfaUfgAfaAfusCfsc D1498 S1498 516 UfuUfcAfuGfuAfAfCfcAfagaGfuAfuUfL96 AS1498 1608 aAfuAfcUfCfUfuGfguuAfcAfuGfaAfasUfsc
D1499 S1499 517 UfuUfcAfuGfuAfAfCfcAfaGfAfGfuAfuUfL96 AS1499 1609 aAfuAfcucUfuGfguuAfcAfuGfaAfasUfsc D1500 S1500 518 UfuCfaUfgUfaAfCfCfaAfgagUfaUfuCfL96 AS1500 1610 gAfaUfaCfUfCfuUfgguUfaCfaUfgAfasAfsu D1501 S1501 519 UfuCfaUfgUfaAfCfCfaAfgAfGfUfaUfuCfL96 AS1501 1611 gAfaUfacuCfuUfgguUfaCfaUfgAfasAfsu D1502 S1502 520 UfcAfuGfuAfaCfCfAfaGfaguAfuUfcCfL96 AS1502 1612 gGfaAfuAfCfUfcUfuggUfuAfcAfuGfasAfsa D1503 S1503 521 UfcAfuGfuAfaCfCfAfaGfaGfUfAfuUfcCfL96 AS1503 1613 gGfaAfuacUfcUfuggUfuAfcAfuGfasAfsa D1504 S1504 522 CfaUfgUfaAfcCfAfAfgAfguaUfuCfcAfL96 AS1504 1614 uGfgAfaUfAfCfuCfuugGfuUfaCfaUfgsAfsa D1505 S1505 523 CfaUfgUfaAfcCfAfAfgAfgUfAfUfuCfcAfL96 AS1505 1615 uGfgAfauaCfuCfuugGfuUfaCfaUfgsAfsa D1506 S1506 524 AfuGfuAfaCfcAfAfGfaGfuauUfcCfaUfL96 AS1506 1616 aUfgGfaAfUfAfcUfcuuGfgUfuAfcAfusGfsa D1507 S1507 525 AfuGfuAfaCfcAfAfGfaGfuAfUfUfcCfaUfL96 AS1507 1617 aUfgGfaauAfcUfcuuGfgUfuAfcAfusGfsa D1508 S1508 526 UfgUfaAfcCfaAfGfAfgUfauuCfcAfuUfL96 AS1508 1618 aAfuGfgAfAfUfaCfucuUfgGfuUfaCfasUfsg D1509 S1509 527 UfgUfaAfcCfaAfGfAfgUfaUfUfCfcAfuUfL96 AS1509 1619 aAfuGfgaaUfaCfucuUfgGfuUfaCfasUfsg D1510 S1510 528 GfuAfaCfcAfaGfAfGfuAfuucCfaUfuUfL96 AS1510 1620 aAfaUfgGfAfAfuAfcucUfuGfgUfuAfcsAfsu D1511 S1511 529 GfuAfaCfcAfaGfAfGfuAfuUfCfCfaUfuUfL96 AS1511 1621 aAfaUfggaAfuAfcucUfuGfgUfuAfcsAfsu D1512 S1512 530 UfaAfcCfaAfgAfGfUfaUfuccAfuUfuUfL96 AS1512 1622 aAfaAfuGfGfAfaUfacuCfuUfgGfuUfasCfsa D1513 S1513 531 UfaAfcCfaAfgAfGfUfaUfuCfCfAfuUfuUfL96 AS1513 1623 aAfaAfuggAfaUfacuCfuUfgGfuUfasCfsa D1514 S1514 532 AfaCfcAfaGfaGfUfAfuUfccaUfuUfuUfL96 AS1514 1624 aAfaAfaUfGfGfaAfuacUfcUfuGfgUfusAfsc D1515 S1515 533 AfaCfcAfaGfaGfUfAfuUfcCfAfUfuUfuUfL96 AS1515 1625 aAfaAfaugGfaAfuacUfcUfuGfgUfusAfsc D1516 S1516 534 AfcCfaAfgAfgUfAfUfuCfcauUfuUfuAfL96 AS1516 1626 uAfaAfaAfUfGfgAfauaCfuCfuUfgGfusUfsa D1517 S1517 535 AfcCfaAfgAfgUfAfUfuCfcAfUfUfuUfuAfL96 AS1517 1627 uAfaAfaauGfgAfauaCfuCfuUfgGfusUfsa D1518 S1518 536 CfcAfaGfaGfuAfUfUfcCfauuUfuUfaCfL96 AS1518 1628 gUfaAfaAfAfUfgGfaauAfcUfcUfuGfgsUfsu D1519 S1519 537 CfcAfaGfaGfuAfUfUfcCfaUfUfUfuUfaCfL96 AS1519 1629 gUfaAfaaaUfgGfaauAfcUfcUfuGfgsUfsu D1520 S1520 538 CfaAfgAfgUfaUfUfCfcAfuuuUfuAfcUfL96 AS1520 1630 aGfuAfaAfAfAfuGfgaaUfaCfuCfuUfgsGfsu D1521 S1521 539 CfaAfgAfgUfaUfUfCfcAfuUfUfUfuAfcUfL96 AS1521 1631 aGfuAfaaaAfuGfgaaUfaCfuCfuUfgsGfsu D1522 S1522 540 AfaGfaGfuAfuUfCfCfaUfuuuUfaCfuAfL96 AS1522 1632 uAfgUfaAfAfAfaUfggaAfuAfcUfcUfusGfsg D1523 S1523 541 AfaGfaGfuAfuUfCfCfaUfuUfUfUfaCfuAfL96 AS1523 1633 uAfgUfaaaAfaUfggaAfuAfcUfcUfusGfsg D1524 S1524 542 AfgAfgUfaUfuCfCfAfuUfuuuAfcUfaAfL96 AS1524 1634 uUfaGfuAfAfAfaAfuggAfaUfaCfuCfusUfsg D1525 S1525 543 AfgAfgUfaUfuCfCfAfuUfuUfUfAfcUfaAfL96 AS1525 1635 uUfaGfuaaAfaAfuggAfaUfaCfuCfusUfsg D1526 S1526 544 GfaGfuAfuUfcCfAfUfuUfuuaCfuAfaAfL96 AS1526 1636 uUfuAfgUfAfAfaAfaugGfaAfuAfcUfcsUfsu D1527 S1527 545 GfaGfuAfuUfcCfAfUfuUfuUfAfCfuAfaAfL96 AS1527 1637 uUfuAfguaAfaAfaugGfaAfuAfcUfcsUfsu D1528 S1528 546 AfgUfaUfuCfcAfUfUfuUfuacUfaAfaGfL96 AS1528 1638 cUfuUfaGfUfAfaAfaauGfgAfaUfaCfusCfsu D1529 S1529 547 AfgUfaUfuCfcAfUfUfuUfuAfCfUfaAfaGfL96 AS1529 1639 cUfuUfaguAfaAfaauGfgAfaUfaCfusCfsu D1530 S1530 548 GfuAfuUfcCfaUfUfUfuUfacuAfaAfgCfL96 AS1530 1640 gCfuUfuAfGfUfaAfaaaUfgGfaAfuAfcsUfsc D1531 S1531 549 GfuAfuUfcCfaUfUfUfuUfaCfUfAfaAfgCfL96 AS1531 1641 gCfuUfuagUfaAfaaaUfgGfaAfuAfcsUfsc D1532 S1532 550 UfaUfuCfcAfuUfUfUfuAfcuaAfaGfcAfL96 AS1532 1642 uGfcUfuUfAfGfuAfaaaAfuGfgAfaUfasCfsu D1533 S1533 551 UfaUfuCfcAfuUfUfUfuAfcUfAfAfaGfcAfL96 AS1533 1643 uGfcUfuuaGfuAfaaaAfuGfgAfaUfasCfsu D1534 S1534 552 AfuUfcCfaUfuUfUfUfaCfuaaAfgCfaGfL96 AS1534 1644 cUfgCfuUfUfAfgUfaaaAfaUfgGfaAfusAfsc D1535 S1535 553 AfuUfcCfaUfuUfUfUfaCfuAfAfAfgCfaGfL96 AS1535 1645 cUfgCfuuuAfgUfaaaAfaUfgGfaAfusAfsc D1536 S1536 554 UfuCfcAfuUfuUfUfAfcUfaaaGfcAfgUfL96 AS1536 1646 aCfuGfcUfUfUfaGfuaaAfaAfuGfgAfasUfsa D1537 S1537 555 UfuCfcAfuUfuUfUfAfcUfaAfAfGfcAfgUfL96 AS1537 1647 aCfuGfcuuUfaGfuaaAfaAfuGfgAfasUfsa D1538 S1538 556 UfcCfaUfuUfuUfAfCluAfaagCfaGfuGfL96 AS1538 1648 cAfcUfgCfUfUfuAfguaAfaAfaUfgGfasAfsu D1539 S1539 557 UfcCfaUfuUfuUfAfCluAfaAfGfCfaGfuGfL96 AS1539 1649 cAfcUfgcuUfuAfguaAfaAfaUfgGfasAfsu D1540 S1540 558 CfcAfuUfuUfuAfCfUfaAfagcAfgUfgUfL96 AS1540 1650 aCfaCfuGfCfUfuUfaguAfaAfaAfuGfgsAfsa D1541 S1541 559 CfcAfuUfuUfuAfCfUfaAfaGfCfAfgUfgUfL96 AS1541 1651 aCfaCfugcUfuUfaguAfaAfaAfuGfgsAfsa D1542 S1542 560 CfaUfuUfuUfaCfUfAfaAfgcaGfuGfuUfL96 AS1542 1652 aAfcAfcUfGfCfuUfuagUfaAfaAfaUfgsGfsa D1543 S1543 561 CfaUfuUfuUfaCfUfAfaAfgCfAfGfuGfuUfL96 AS1543 1653 aAfcAfcugCfuUfuagUfaAfaAfaUfgsGfsa D1544 S1544 562 AfuUfuUfuAfcUfAfAfaGfcagUfgUfuUfL96 AS1544 1654 aAfaCfaCfUfGfcUfuuaGfuAfaAfaAfusGfsg D1545 S1545 563 AfuUfuUfuAfcUfAfAfaGfcAfGfUfgUfuUfL96 AS1545 1655 aAfaCfacuGfcUfuuaGfuAfaAfaAfusGfsg D1546 S1546 564 UfuUfuUfaCfuAfAfAfgCfaguGfuUfuUfL96 AS1546 1656 aAfaAfcAfCfUfgCfuuuAfgUfaAfaAfasUfsg D1547 S1547 565 UfuUfuUfaCfuAfAfAfgCfaGfUfGfuUfuUfL96 AS1547 1657 aAfaAfcacUfgCfuuuAfgUfaAfaAfasUfsg D1548 S1548 566 UfuUfuAfcUfaAfAfGfcAfgugUfuUfuCfL96 AS1548 1658 gAfaAfaCfAfCfuGfcuuUfaGfuAfaAfasAfsu D1549 S1549 567 UfuUfuAfcUfaAfAfGfcAfgUfGfUfuUfuCfL96 AS1549 1659 gAfaAfacaCfuGfcuuUfaGfuAfaAfasAfsu D1550 S1550 568 UfuUfaCfuAfaAfGfCfaGfuguUfuUfcAfL96 AS1550 1660 uGfaAfaAfCfAfcUfgcuUfuAfgUfaAfasAfsa D1551 S1551 569 UfuUfaCfuAfaAfGfCfaGfuGfUfUfuUfcAfL96 AS1551 1661 uGfaAfaacAfcUfgcuUfuAfgUfaAfasAfsa D1552 S1552 570 UfuAfcUfaAfaGfCfAfgUfguuUfuCfaCfL96 AS1552 1662 gUfgAfaAfAfCfaCfugcUfuUfaGfuAfasAfsa D1553 S1553 571 UfuAfcUfaAfaGfCfAfgUfgUfUfUfuCfaCfL96 AS1553 1663 gUfgAfaaaCfaCfugcUfuUfaGfuAfasAfsa D1554 S1554 572 UfaCfuAfaAfgCfAfGfuGfuuuUfcAfcCfL96 AS1554 1664 gGfuGfaAfAfAfcAfcugCfuUfuAfgUfasAfsa D1555 S1555 573 UfaCfuAfaAfgCfAfGfuGfuUfUfUfcAfcCfL96 AS1555 1665 gGfuGfaaaAfcAfcugCfuUfuAfgUfasAfsa D1556 S1556 574 AfcUfaAfaGfcAfGfUfgUfuuuCfaCfcUfL96 AS1556 1666 aGfgUfgAfAfAfaCfacuGfcUfuUfaGfusAfsa D1557 S1557 575 AfcUfaAfaGfcAfGfUfgUfuUfUfCfaCfcUfL96 AS1557 1667 aGfgUfgaaAfaCfacuGfcUfuUfaGfusAfsa D1558 S1558 576 CfuAfaAfgCfaGfUfGfuUfuucAfcCfuCfL96 AS1558 1668 gAfgGfuGfAfAfaAfcacUfgCfuUfuAfgsUfsa D1559 S1559 577 CfuAfaAfgCfaGfUfGfuUfuUfCfAfcCfuCfL96 AS1559 1669 gAfgGfugaAfaAfcacUfgCfuUfuAfgsUfsa D1560 S1560 578 UfaAfaGfcAfgUfGfUfuUfucaCfcUfcAfL96 AS1560 1670 uGfaGfgUfGfAfaAfacaCfuGfcUfuUfasGfsu D1561 S1561 579 UfaAfaGfcAfgUfGfUfuUfuCfAfCfcUfcAfL96 AS1561 1671 uGfaGfgugAfaAfacaCfuGfcUfuUfasGfsu D1562 S1562 580 AfaAfgCfaGfuGfUfUfuUfcacCfuCfaUfL96 AS1562 1672 aUfgAfgGfUfGfaAfaacAfcUfgCfuUfusAfsg D1563 S1563 581 AfaAfgCfaGfuGfUfUfuUfcAfCfCfuCfaUfL96 AS1563 1673 aUfgAfgguGfaAfaacAfcUfgCfuUfusAfsg D1564 S1564 582 AfaGfcAfgUfgUfUfUfuCfaccUfcAfuAfL96 AS1564 1674 uAfuGfaGfGfUfgAfaaaCfaCfuGfcUfusUfsa D1565 S1565 583 AfaGfcAfgUfgUfUfUfuCfaCfCfUfcAfuAfL96 AS1565 1675 uAfuGfaggUfgAfaaaCfaCfuGfcUfusUfsa D1566 S1566 584 AfgCfaGfuGfuUfUfUfcAfccuCfaUfaUfL96 AS1566 1676 aUfaUfgAfGfGfuGfaaaAfcAfcUfgCfusUfsu D1567 S1567 585 AfgCfaGfuGfuUfUfUfcAfcCfUfCfaUfaUfL96 AS1567 1677 aUfaUfgagGfuGfaaaAfcAfcUfgCfusUfsu D1568 S1568 586 GfcAfgUfgUfuUfUfCfaCfcucAfuAfuGfL96 AS1568 1678 cAfuAfuGfAfGfgUfgaaAfaCfaCfuGfcsUfsu D1569 S1569 587 GfcAfgUfgUfuUfUfCfaCfcUfCfAfuAfuGfL96 AS1569 1679 cAfuAfugaGfgUfgaaAfaCfaCfuGfcsUfsu D1570 S1570 588 CfaGfuGfuUfuUfCfAfcCfucaUfaUfgCfL96 AS1570 1680 gCfaUfaUfGfAfgGfugaAfaAfcAfcUfgsCfsu D1571 S1571 589 CfaGfuGfuUfuUfCfAfcCfuCfAfUfaUfgCfL96 AS1571 1681 gCfaUfaugAfgGfugaAfaAfcAfcUfgsCfsu D1572 S1572 590 AfgUfgUfuUfuCfAfCfcUfcauAfuGfcUfL96 AS1572 1682 aGfcAfuAfUfGfaGfgugAfaAfaCfaCfusGfsc D1573 S1573 591 AfgUfgUfuUfuCfAfCfcUfcAfUfAfuGfcUfL96 AS1573 1683 aGfcAfuauGfaGfgugAfaAfaCfaCfusGfsc D1574 S1574 592 GfuGfuUfuUfcAfCfCfuCfauaUfgCluAfL96 AS1574 1684 uAfgCfaUfAfUfgAfgguGfaAfaAfcAfcsUfsg D1575 S1575 593 GfuGfuUfuUfcAfCfCfuCfaUfAfUfgCluAfL96 AS1575 1685 uAfgCfauaUfgAfgguGfaAfaAfcAfcsUfsg D1576 S1576 594 UfgUfuUfuCfaCfCfUfcAfuauGfcUfaUfL96 AS1576 1686 aUfaGfcAfUfAfuGfaggUfgAfaAfaCfasCfsu D1577 S1577 595 UfgUfuUfuCfaCfCfUfcAfuAfUfGfcUfaUfL96 AS1577 1687 aUfaGfcauAfuGfaggUfgAfaAfaCfasCfsu D1578 S1578 596 GfuUfuUfcAfcCfUfCfaUfaugCluAfuGfL96 AS1578 1688 cAfuAfgCfAfUfaUfgagGfuGfaAfaAfcsAfsc D1579 S1579 597 GfuUfuUfcAfcCfUfCfaUfaUfGfCluAfuGfL96 AS1579 1689 cAfuAfgcaUfaUfgagGfuGfaAfaAfcsAfsc D1580 S1580 598 UfuUfuCfaCfcUfCfAfuAfugcUfaUfgUfL96 AS1580 1690 aCfaUfaGfCfAfuAfugaGfgUfgAfaAfasCfsa D1581 S1581 599 UfuUfuCfaCfcUfCfAfuAfuGfCfUfaUfgUfL96 AS1581 1691 aCfaUfagcAfuAfugaGfgUfgAfaAfasCfsa D1582 S1582 600 UfuUfcAfcCfuCfAfUfaUfgcuAfuGfuUfL96 AS1582 1692
aAfcAfuAfGfCfaUfaugAfgGfuGfaAfasAfsc D1583 S1583 601 UfuUfcAfcCfuCfAfUfaUfgClUfAfuGfuUfL96 AS1583 1693 aAfcAfuagCfaUfaugAfgGfuGfaAfasAfsc D1584 S1584 602 UfuCfaCfcUfcAfUfAfuGfcuaUfgUfuAfL96 AS1584 1694 uAfaCfaUfAfGfcAfuauGfaGfgUfgAfasAfsa D1585 S1585 603 UfuCfaCfcUfcAfUfAfuGfcUfAfUfgUfuAfL96 AS1585 1695 uAfaCfauaGfcAfuauGfaGfgUfgAfasAfsa D1586 S1586 604 UfcAfcCfuCfaUfAfUfgCluauGfuUfaGfL96 AS1586 1696 cUfaAfcAfUfAfgCfauaUfgAfgGfuGfasAfsa D1587 S1587 605 UfcAfcCfuCfaUfAfUfgCluAfUfGfuUfaGfL96 AS1587 1697 cUfaAfcauAfgCfauaUfgAfgGfuGfasAfsa D1588 S1588 606 CfaCfcUfcAfuAfUfGfcUfaugUfuAfgAfL96 AS1588 1698 uCfuAfaCfAfUfaGfcauAfuGfaGfgUfgsAfsa D1589 S1589 607 CfaCfcUfcAfuAfUfGfcUfaUfGfUfuAfgAfL96 AS1589 1699 uCfuAfacaUfaGfcauAfuGfaGfgUfgsAfsa D1590 S1590 608 AfcCfuCfaUfaUfGfCluAfuguUfaGfaAfL96 AS1590 1700 uUfcUfaAfCfAfuAfgcaUfaUfgAfgGfusGfsa D1591 S1591 609 AfcCfuCfaUfaUfGfCluAfuGfUfUfaGfaAfL96 AS1591 1701 uUfcUfaacAfuAfgcaUfaUfgAfgGfusGfsa D1592 S1592 610 CfcUfcAfuAfuGfCfUfaUfguuAfgAfaGfL96 AS1592 1702 cUfuCluAfAfCfaUfagcAfuAfuGfaGfgsUfsg D1593 S1593 611 CfcUfcAfuAfuGfCfUfaUfgUfUfAfgAfaGfL96 AS1593 1703 cUfuCfuaaCfaUfagcAfuAfuGfaGfgsUfsg D1594 S1594 612 CfuCfaUfaUfgClUfAfuGfuuaGfaAfgUfL96 AS1594 1704 aCfuUfcUfAfAfcAfuagCfaUfaUfgAfgsGfs u D1595 S1595 613 CfuCfaUfaUfgClUfAfuGfuUfAfGfaAfgUfL96 AS1595 1705 aCfuUfcuaAfcAfuagCfaUfaUfgAfgsGfsu D1596 S1596 614 UfcAfuAfuGfcUfAfUfgUfuagAfaGfuCfL96 AS1596 1706 gAfcUfuCfUfAfaCfauaGfcAfuAfuGfasGfsg D1597 S1597 615 UfcAfuAfuGfcUfAfUfgUfuAfGfAfaGfuCfL96 AS1597 1707 gAfcUfucuAfaCfauaGfcAfuAfuGfasGfsg D1598 S1598 616 CfaUfaUfgCluAfUfGfuUfagaAfgUfcCfL96 AS1598 1708 gGfaCfuUfCfUfaAfcauAfgCfaUfaUfgsAfsg D1599 S1599 617 CfaUfaUfgCluAfUfGfuUfaGfAfAfgUfcCfL96 AS1599 1709 gGfaCfuucUfaAfcauAfgCfaUfaUfgsAfsg D1600 S1600 618 AfuAfuGfcUfaUfGfUfuAfgaaGfuCfcAfL96 AS1600 1710 uGfgAfcUfUfCfuAfacaUfaGfcAfuAfusGfsa D1601 S1601 619 AfuAfuGfcUfaUfGfUfuAfgAfAfGfuCfcAfL96 AS1601 1711 uGfgAfcuuCluAfacaUfaGfcAfuAfusGfsa D1602 S1602 620 UfaUfgCluAfuGfUfUfaGfaagUfcCfaGfL96 AS1602 1712 cUfgGfaCfUfUfcUfaacAfuAfgCfaUfasUfsg D1603 S1603 621 UfaUfgCluAfuGfUfUfaGfaAfGfUfcCfaGfL96 AS1603 1713 cUfgGfacuUfcUfaacAfuAfgCfaUfasUfsg D1604 S1604 622 AfuGfcUfaUfgUfUfAfgAfaguCfcAfgGfL96 AS1604 1714 cCfuGfgAfCfUfuCfuaaCfaUfaGfcAfusAfsu D1605 S1605 623 AfuGfcUfaUfgUfUfAfgAfaGfUfCfcAfgGfL96 AS1605 1715 cCfuGfgacUfuCluaaCfaUfaGfcAfusAfsu D1606 S1606 624 UfgCluAfuGfuUfAfGfaAfgucCfaGfgCfL96 AS1606 1716 gCfcUfgGfAfCfuUfcuaAfcAfuAfgCfasUfsa D1607 S1607 625 UfgCluAfuGfuUfAfGfaAfgUfCfCfaGfgCfL96 AS1607 1717 gCfcUfggaCfuUfcuaAfcAfuAfgCfasUfsa D1608 S1608 626 GfcUfaUfgUfuAfGfAfaGfuccAfgGfcAfL96 AS1608 1718 uGfcCfuGfGfAfcUfucuAfaCfaUfaGfcsAfsu D1609 S1609 627 GfcUfaUfgUfuAfGfAfaGfuCfCfAfgGfcAfL96 AS1609 1719 uGfcCfuggAfcUfucuAfaCfaUfaGfcsAfsu D1610 S1610 628 CfuAfuGfuUfaGfAfAfgUfccaGfgCfaGfL96 AS1610 1720 cUfgCfcUfGfGfaCfuucUfaAfcAfuAfgsCfsa D1611 S1611 629 CfuAfuGfuUfaGfAfAfgUfcCfAfGfgCfaGfL96 AS1611 1721 cUfgCfcugGfaCfuucUfaAfcAfuAfgsCfsa D1612 S1612 630 UfaUfgUfuAfgAfAfGfuCfcagGfcAfgAfL96 AS1612 1722 uCfuGfcCfUfGfgAfcuuCfuAfaCfaUfasGfsc D1613 S1613 631 UfaUfgUfuAfgAfAfGfuCfcAfGfGfcAfgAfL96 AS1613 1723 uCfuGfccuGfgAfcuuCfuAfaCfaUfasGfsc D1614 S1614 632 AfuGfuUfaGfaAfGfUfcCfaggCfaGfaGfL96 AS1614 1724 cUfcUfgCfCfUfgGfacuUfcUfaAfcAfusAfsg D1615 S1615 633 AfuGfuUfaGfaAfGfUfcCfaGfGfCfaGfaGfL96 AS1615 1725 cUfcUfgccUfgGfacuUfcUfaAfcAfusAfsg D1616 S1616 634 UfgUfuAfgAfaGfUfCfcAfggcAfgAfgAfL96 AS1616 1726 uCfuCfuGfCfCfuGfgacUfuCfuAfaCfasUfsa D1617 S1617 635 UfgUfuAfgAfaGfUfCfcAfgGfCfAfgAfgAfL96 AS1617 1727 uCfuCfugcCfuGfgacUfuCfuAfaCfasUfsa D1618 S1618 636 GfuUfaGfaAfgUfCfCfaGfgcaGfaGfaCfL96 AS1618 1728 gUfcUfcUfGfCfcUfggaCfuUfcUfaAfcsAfsu D1619 S1619 637 GfuUfaGfaAfgUfCfCfaGfgCfAfGfaGfaCfL96 AS1619 1729 gUfcUfcugCfcUfggaCfuUfcUfaAfcsAfsu D1620 S1620 638 UfuAfgAfaGfuCfCfAfgGfcagAfgAfcAfL96 AS1620 1730 uGfuCfuCfUfGfcCfuggAfcUfuCfuAfasCfsa D1621 S1621 639 UfuAfgAfaGfuCfCfAfgGfcAfGfAfgAfcAfL96 AS1621 1731 uGfuCfucuGfcCfuggAfcUfuCluAfasCfsa D1622 S1622 640 UfaGfaAfgUfcCfAfGfgCfagaGfaCfaAfL96 AS1622 1732 uUfgUfcUfCfUfgCfcugGfaCfuUfcUfasAfsc D1623 S1623 641 UfaGfaAfgUfcCfAfGfgCfaGfAfGfaCfaAfL96 AS1623 1733 uUfgUfcucUfgCfcugGfaCfuUkUfasAfsc D1624 S1624 642 AfgAfaGfuCfcAfGfGfcAfgagAfcAfaUfL96 AS1624 1734 aUfuGfuCfUfCfuGfccuGfgAfcUfuCfusAfsa D1625 S1625 643 AfgAfaGfuCfcAfGfGfcAfgAfGfAfcAfaUfL96 AS1625 1735 aUfuGfucuCfuGfccuGfgAfcUfuCfusAfsa D1626 S1626 644 GfaAfgUfcCfaGfGfCfaGfagaCfaAfuAfL96 AS1626 1736 uAfuUfgUfCfUfcUfgccUfgGfaCfuUfcsUfsa D1627 S1627 645 GfaAfgUfcCfaGfGfCfaGfaGfAfCfaAfuAfL96 AS1627 1737 uAfuUfgucUfcUfgccUfgGfaCfuUfcsUfsa D1628 S1628 646 AfaGfuCfcAfgGfCfAfgAfgacAfaUfaAfL96 AS1628 1738 uUfaUfuGfUfCfuCfugcCfuGfgAfcUfusCfsu D1629 S1629 647 AfaGfuCfcAfgGfCfAfgAfgAfCfAfaUfaAfL96 AS1629 1739 uUfaUfuguCfuCfugcCfuGfgAfcUfusCfsu D1630 S1630 648 AfgUfcCfaGfgCfAfGfaGfacaAfuAfaAfL96 AS1630 1740 uUfuAfuUfGfUfcUfcugCfcUfgGfaCfusUfsc D1631 S1631 649 AfgUfcCfaGfgCfAfGfaGfaCfAfAfuAfaAfL96 AS1631 1741 uUfuAfuugUfcUfcugCfcUfgGfaCfusUfsc D1632 S1632 650 GfuCfcAfgGfcAfGfAfgAfcaaUfaAfaAfL96 AS1632 1742 uUfuUfaUfUfGfuCfucuGfcCfuGfgAfcsUfsu D1633 S1633 651 GfuCfcAfgGfcAfGfAfgAfcAfAfUfaAfaAfL96 AS1633 1743 uUfuUfauuGfuCfucuGfcCfuGfgAfcsUfsu D1634 S1634 652 UfcCfaGfgCfaGfAfGfaCfaauAfaAfaCfL96 AS1634 1744 gUfuUfuAfUfUfgUfcucUfgCfcUfgGfasCfsu D1635 S1635 653 UfcCfaGfgCfaGfAfGfaCfaAfUfAfaAfaCfL96 AS1635 1745 gUfuUfuauUfgUfcucUfgCfcUfgGfasCfsu D1636 S1636 654 CfcAfgGfcAfgAfGfAfcAfauaAfaAfcAfL96 AS1636 1746 uGfuUfuUfAfUfuGfucuCfuGfcCfuGfgsAfsc D1637 S1637 655 CfcAfgGfcAfgAfGfAfcAfaUfAfAfaAfcAfL96 AS1637 1747 uGfuUfuuaUfuGfucuCfuGfcCfuGfgsAfsc D1638 S1638 656 CfaGfgCfaGfaGfAfCfaAfuaaAfaCfaUfL96 AS1638 1748 aUfgUfuUfUfAfuUfgucUfcUfgCfcUfgsGfsa D1639 S1639 657 CfaGfgCfaGfaGfAfCfaAfuAfAfAfaCfaUfL96 AS1639 1749 aUfgUfuuuAfuUfgucUfcUfgCfcUfgsGfsa D1640 S1640 658 AfgGfcAfgAfgAfCfAfaUfaaaAfcAfuUfL96 AS1640 1750 aAfuGfuUfUfUfaUfuguCfuCfuGfcCfusGfsg D1641 S1641 659 AfgGfcAfgAfgAfCfAfaUfaAfAfAfcAfuUfL96 AS1641 1751 aAfuGfuuuUfaUfuguCfuCfuGfcCfusGfsg D1642 S1642 660 GfgCfaGfaGfaCfAfAfuAfaaaCfaUfuCfL96 AS1642 1752 gAfaUfgUfUfUfuAfuugUfcUfcUfgCfcsUfsg D1643 S1643 661 GfgCfaGfaGfaCfAfAfuAfaAfAfCfaUfuCfL96 AS1643 1753 gAfaUfguuUfuAfuugUfcUfcUfgCfcsUfsg D1644 S1644 662 GfcAfgAfgAfcAfAfUfaAfaacAfuUfcCfL96 AS1644 1754 gGfaAfuGfUfUfuUfauuGfuCfuCfuGfcsCfsu D1645 S1645 663 GfcAfgAfgAfcAfAfUfaAfaAfCfAfuUfcCfL96 AS1645 1755 gGfaAfuguUfuUfauuGfuCfuCfuGfcsCfsu D1646 S1646 664 CfaGfaGfaCfaAfUfAfaAfacaUfuCfcUfL96 AS1646 1756 aGfgAfaUfGfUfuUfuauUfgUfcUfcUfgsCfsc D1647 S1647 665 CfaGfaGfaCfaAfUfAfaAfaCfAfUfuCfcUfL96 AS1647 1757 aGfgAfaugUfuUfuauUfgUfcUfcUfgsCfsc D1648 S1648 666 AfgAfgAfcAfaUfAfAfaAfcauUfcCfuGfL96 AS1648 1758 cAfgGfaAfUfGfuUfuuaUfuGfuCfuCfusGfsc D1649 S1649 667 AfgAfgAfcAfaUfAfAfaAfcAfUfUfcCfuGfL96 AS1649 1759 cAfgGfaauGfuUfuuaUfuGfuCfuCfusGfsc D1650 S1650 668 GfaGfaCfaAfuAfAfAfaCfauuCfcUfgUfL96 AS1650 1760 aCfaGfgAfAfUfgUfuuuAfuUfgUfcUfcsUfsg D1651 S1651 669 GfaGfaCfaAfuAfAfAfaCfaUfUfCfcUfgUfL96 AS1651 1761 aCfaGfgaaUfgUfuuuAfuUfgUfcUfcsUfsg D1652 S1652 670 AfgAfcAfaUfaAfAfAfcAfuucCfuGfuGfL96 AS1652 1762 cAfcAfgGfAfAfuGfuuuUfaUfuGfuCfusCfsu D1653 S1653 671 AfgAfcAfaUfaAfAfAfcAfuUfCfCfuGfuGfL96 AS1653 1763 cAfcAfggaAfuGfuuuUfaUfuGfuCfusCfsu D1654 S1654 672 GfaCfaAfuAfaAfAfCfaUfuccUfgUfgAfL96 AS1654 1764 uCfaCfaGfGfAfaUfguuUfuAfuUfgUfcsUfsc D1655 S1655 673 GfaCfaAfuAfaAfAfCfaUfuCfCfUfgUfgAfL96 AS1655 1765 uCfaCfaggAfaUfguuUfuAfuUfgUfcsUfsc D1656 S1656 674 AfcAfaUfaAfaAfCfAfuUfccuGfuGfaAfL96 AS1656 1766 uUfcAfcAfGfGfaAfuguUfuUfaUfuGfusCfsu D1657 S1657 675 AfcAfaUfaAfaAfCfAfuUfcCfUfGfuGfaAfL96 AS1657 1767 uUfcAfcagGfaAfuguUfuUfaUfuGfusCfsu D1658 S1658 676 CfaAfuAfaAfaCfAfUfuCfcugUfgAfaAfL96 AS1658 1768 uUfuCfaCfAfGfgAfaugUfuUfuAfuUfgsUfsc D1659 S1659 677 CfaAfuAfaAfaCfAfUfuCfcUfGfUfgAfaAfL96 AS1659 1769 uUfuCfacaGfgAfaugUfuUfuAfuUfgsUfsc D1660 S1660 678 AfaUfaAfaAfcAfUfUfcCfuguGfaAfaGfL96 AS1660 1770 cUfuUfcAfCfAfgGfaauGfuUfuUfaUfusGfsu D1661 S1661 679 AfaUfaAfaAfcAfUfUfcCfuGfUfGfaAfaGfL96 AS1661 1771 cUfuUfcacAfgGfaauGfuUfuUfaUfusGfsu D1662 S1662 680 AfuAfaAfaCfaUfUfCfcUfgugAfaAfgGfL96 AS1662 1772 cCfuUfuCfAfCfaGfgaaUfgUfuUfuAfusUfsg D1663 S1663 681 AfuAfaAfaCfaUfUfCfcUfgUfGfAfaAfgGfL96 AS1663 1773 cCfuUfucaCfaGfgaaUfgUfuUfuAfusUfsg D1664 S1664 682 UfaAfaAfcAfuUfCfCfuGfugaAfaGfgCfL96 AS1664 1774 gCfcUfuUfCfAfcAfggaAfuGfuUfuUfasUfsu D1665 S1665 683 UfaAfaAfcAfuUfCfCfuGfuGfAfAfaGfgCfL96 AS1665 1775 gCfcUfuucAfcAfggaAfuGfuUfuUfasUfsu
D1666 S1666 684 AfaAfaCfaUfuCfCfUfgUfgaaAfgGfcAfL96 AS1666 1776 uGfcCfuUfUfCfaCfaggAfaUfgUfuUfusAfsu D1667 S1667 685 AfaAfaCfaUfuCfCfUfgUfgAfAfAfgGfcAfL96 AS1667 1777 uGfcCfuuuCfaCfaggAfaUfgUfuUfusAfsu D1668 S1668 686 AfaAfcAfuUfcCfUfGfuGfaaaGfgCfaCfL96 AS1668 1778 gUfgCfcUfUfUfcAfcagGfaAfuGfuUfusUfsa D1669 S1669 687 AfaAfcAfuUfcCfUfGfuGfaAfAfGfgCfaCfL96 AS1669 1779 gUfgCfcuuUfcAfcagGfaAfuGfuUfusUfsa D1670 S1670 688 AfaCfaUfuCfcUfGfUfgAfaagGfcAfcUfL96 AS1670 1780 aGfuGfcCfUfUfuCfacaGfgAfaUfgUfusUfsu D1671 S1671 689 AfaCfaUfuCfcUfGfUfgAfaAfGfGfcAfcUfL96 AS1671 1781 aGfuGfccuUfuCfacaGfgAfaUfgUfusUfsu D1672 S1672 690 AfcAfuUfcCfuGfUfGfaAfaggCfaCfuUfL96 AS1672 1782 aAfgUfgCfCfUfuUfcacAfgGfaAfuGfusUfsu D1673 S1673 691 AfcAfuUfcCfuGfUfGfaAfaGfGfCfaCfuUfL96 AS1673 1783 aAfgUfgccUfuUfcacAfgGfaAfuGfusUfsu D1674 S1674 692 CfaUfuCfcUfgUfGfAfaAfggcAfcUfuUfL96 AS1674 1784 aAfaGfuGfCfCfuUfucaCfaGfgAfaUfgsUfsu D1675 S1675 693 CfaUfuCfcUfgUfGfAfaAfgGfCfAfcUfuUfL96 AS1675 1785 aAfaGfugcCfuUfucaCfaGfgAfaUfgsUfsu D1676 S1676 694 AfuUfcCfuGfuGfAfAfaGfgcaCfuUfuUfL96 AS1676 1786 aAfaAfgUfGfCfcUfuucAfcAfgGfaAfusGfsu D1677 S1677 695 AfuUfcCfuGfuGfAfAfaGfgCfAfCfuUfuUfL96 AS1677 1787 aAfaAfgugCfcUfuucAfcAfgGfaAfusGfsu D1678 S1678 696 UfuCfcUfgUfgAfAfAfgGfcacUfuUfuCfL96 AS1678 1788 gAfaAfaGfUfGfcCfuuuCfaCfaGfgAfasUfsg D1679 S1679 697 UfuCfcUfgUfgAfAfAfgGfcAfCfUfuUfuCfL96 AS1679 1789 gAfaAfaguGfcCfuuuCfaCfaGfgAfasUfsg D1680 S1680 698 UfcCfuGfuGfaAfAfGfgCfacuUfuUfcAfL96 AS1680 1790 uGfaAfaAfGfUfgCfcuuUfcAfcAfgGfasAfsu D1681 S1681 699 UfcCfuGfuGfaAfAfGfgCfaCfUfUfuUfcAfL96 AS1681 1791 uGfaAfaagUfgCfcuuUfcAfcAfgGfasAfsu D1682 S1682 700 CfcUfgUfgAfaAfGfGfcAfcuuUfuCfaUfL96 AS1682 1792 aUfgAfaAfAfGfuGfccuUfuCfaCfaGfgsAfsa D1683 S1683 701 CfcUfgUfgAfaAfGfGfcAfcUfUfUfuCfaUfL96 AS1683 1793 aUfgAfaaaGfuGfccuUfuCfaCfaGfgsAfsa D1684 S1684 702 CfuGfuGfaAfaGfGfCfaCfuuuUfcAfuUfL96 AS1684 1794 aAfuGfaAfAfAfgUfgccUfuUfcAfcAfgsGfsa D1685 S1685 703 CfuGfuGfaAfaGfGfCfaCfuUfUfUfcAfuUfL96 AS1685 1795 aAfuGfaaaAfgUfgccUfuUfcAfcAfgsGfsa D1686 S1686 704 UfgUfgAfaAfgGfCfAfcUfuuuCfaUfuCfL96 AS1686 1796 gAfaUfgAfAfAfaGfugcCfuUfuCfaCfasGfsg D1687 S1687 705 UfgUfgAfaAfgGfCfAfcUfuUfUfCfaUfuCfL96 AS1687 1797 gAfaUfgaaAfaGfugcCfuUfuCfaCfasGfsg D1688 S1688 706 GfuGfaAfaGfgCfAfCfuUfuucAfuUfcCfL96 AS1688 1798 gGfaAfuGfAfAfaAfgugCfcUfuUfcAfcsAfsg D1689 S1689 707 GfuGfaAfaGfgCfAfCfuUfuUfCfAfuUfcCfL96 AS1689 1799 gGfaAfugaAfaAfgugCfcUfuUfcAfcsAfsg D1690 S1690 708 UfgAfaAfgGfcAfCfUfuUfucaUfuCfcAfL96 AS1690 1800 uGfgAfaUfGfAfaAfaguGfcCfuUfuCfasCfsa D1691 S1691 709 UfgAfaAfgGfcAfCfUfuUfuCfAfUfuCfcAfL96 AS1691 1801 uGfgAfaugAfaAfaguGfcCfuUfuCfasCfsa D1692 S1692 710 GfaAfaGfgCfaCfUfUfuUfcauUfcCfaCfL96 AS1692 1802 gUfgGfaAfUfGfaAfaagUfgCfcUfuUfcsAfsc D1693 S1693 711 GfaAfaGfgCfaCfUfUfuUfcAfUfUfcCfaCfL96 AS1693 1803 gUfgGfaauGfaAfaagUfgCfcUfuUfcsAfsc D1694 S1694 712 AfaAfgGfcAfcUfUfUfuCfauuCfcAfcUfL96 AS1694 1804 aGfuGfgAfAfUfgAfaaaGfuGfcCfuUfusCfsa D1695 S1695 713 AfaAfgGfcAfcUfUfUfuCfaUfUfCfcAfcUfL96 AS1695 1805 aGfuGfgaaUfgAfaaaGfuGfcCfuUfusCfsa D1696 S1696 714 AfaGfgCfaCfuUfUfUfcAfuucCfaCfuUfL96 AS1696 1806 aAfgUfgGfAfAfuGfaaaAfgUfgCfcUfusUfsc D1697 S1697 715 AfaGfgCfaCfuUfUfUfcAfuUfCfCfaCfuUfL96 AS1697 1807 aAfgUfggaAfuGfaaaAfgUfgCfcUfusUfsc D1698 S1698 716 AfgGfcAfcUfuUfUfCfaUfuccAfcUfuUfL96 AS1698 1808 aAfaGfuGfGfAfaUfgaaAfaGfuGfcCfusUfsu D1699 S1699 717 AfgGfcAfcUfuUfUfCfaUfuCfCfAfcUfuUfL96 AS1699 1809 aAfaGfuggAfaUfgaaAfaGfuGfcCfusUfsu D1700 S1700 718 GfgCfaCfuUfuUfCfAfuUfccaCfuUfuAfL96 AS1700 1810 uAfaAfgUfGfGfaAfugaAfaAfgUfgCfcsUfsu D1701 S1701 719 GfgCfaCfuUfuUfCfAfuUfcCfAfCfuUfuAfL96 AS1701 1811 uAfaAfgugGfaAfugaAfaAfgUfgCfcsUfsu D1702 S1702 720 GfcAfcUfuUfuCfAfUfuCfcacUfuUfaAfL96 AS1702 1812 uUfaAfaGfUfGfgAfaugAfaAfaGfuGfcsCfsu D1703 S1703 721 GfcAfcUfuUfuCfAfUfuCfcAfCfUfuUfaAfL96 AS1703 1813 uUfaAfaguGfgAfaugAfaAfaGfuGfcsCfsu D1704 S1704 722 CfaCfuUfuUfcAfUfUfcCfacuUfuAfaCfL96 AS1704 1814 gUfuAfaAfGfUfgGfaauGfaAfaAfgUfgsCfsc D1705 S1705 723 CfaCfuUfuUfcAfUfUfcCfaCfUfUfuAfaCfL96 AS1705 1815 gUfuAfaagUfgGfaauGfaAfaAfgUfgsCfsc D1706 S1706 724 AfcUfuUfuCfaUfUfCfcAfcuuUfaAfcUfL96 AS1706 1816 aGfuUfaAfAfGfuGfgaaUfgAfaAfaGfusGfsc D1707 S1707 725 AfcUfuUfuCfaUfUfCfcAfcUfUfUfaAfcUfL96 AS1707 1817 aGfuUfaaaGfuGfgaaUfgAfaAfaGfusGfsc D1708 S1708 726 CfuUfuUfcAfuUfCfCfaCfuuuAfaCfuUfL96 AS1708 1818 aAfgUfuAfAfAfgUfggaAfuGfaAfaAfgsUfsg D1709 S1709 727 CfuUfuUfcAfuUfCfCfaCfuUfUfAfaCfuUfL96 AS1709 1819 aAfgUfuaaAfgUfggaAfuGfaAfaAfgsUfsg D1710 S1710 728 UfuUfuCfaUfuCfCfAfcUfuuaAfcUfuGfL96 AS1710 1820 cAfaGfuUfAfAfaGfuggAfaUfgAfaAfasGfsu D1711 S1711 729 UfuUfuCfaUfuCfCfAfcUfuUfAfAfcUfuGfL96 AS1711 1821 cAfaGfuuaAfaGfuggAfaUfgAfaAfasGfsu D1712 S1712 730 UfuUfcAfuUfcCfAfCfuUfuaaCfuUfgAfL96 AS1712 1822 uCfaAfgUfUfAfaAfgugGfaAfuGfaAfasAfsg D1713 S1713 731 UfuUfcAfuUfcCfAfCfuUfuAfAfCfuUfgAfL96 AS1713 1823 uCfaAfguuAfaAfgugGfaAfuGfaAfasAfsg D1714 S1714 732 UfuCfaUfuCfcAfCfUfuUfaacUfuGfaUfL96 AS1714 1824 aUfcAfaGfUfUfaAfaguGfgAfaUfgAfasAfsa D1715 S1715 733 UfuCfaUfuCfcAfCfUfuUfaAfCfUfuGfaUfL96 AS1715 1825 aUfcAfaguUfaAfaguGfgAfaUfgAfasAfsa D1716 S1716 734 UfcAfuUfcCfaCfUfUfuAfacuUfgAfuUfL96 AS1716 1826 aAfuCfaAfGfUfuAfaagUfgGfaAfuGfasAfsa D1717 S1717 735 UfcAfuUfcCfaCfUfUfuAfaCfUfUfgAfuUfL96 AS1717 1827 aAfuCfaagUfuAfaagUfgGfaAfuGfasAfsa D1718 S1718 736 CfaUfuCfcAfcUfUfUfaAfcuuGfaUfuUfL96 AS1718 1828 aAfaUfcAfAfGfuUfaaaGfuGfgAfaUfgsAfsa D1719 S1719 737 CfaUfuCfcAfcUfUfUfaAfcUfUfGfaUfuUfL96 AS1719 1829 aAfaUfcaaGfuUfaaaGfuGfgAfaUfgsAfsa D1720 S1720 738 AfuUfcCfaCfuUfUfAfaCfuugAfuUfuUfL96 AS1720 1830 aAfaAfuCfAfAfgUfuaaAfgUfgGfaAfusGfsa D1721 S1721 739 AfuUfcCfaCfuUfUfAfaCfuUfGfAfuUfuUfL96 AS1721 1831 aAfaAfucaAfgUfuaaAfgUfgGfaAfusGfsa D1722 S1722 740 UfuCfcAfcUfuUfAfAfcUfugaUfuUfuUfL96 AS1722 1832 aAfaAfaUfCfAfaGfuuaAfaGfuGfgAfasUfsg D1723 S1723 741 UfuCfcAfcUfuUfAfAfcUfuGfAfUfuUfuUfL96 AS1723 1833 aAfaAfaucAfaGfuuaAfaGfuGfgAfasUfsg D1724 S1724 742 UfcCfaCfuUfuAfAfCfuUfgauUfuUfuUfL96 AS1724 1834 aAfaAfaAfUfCfaAfguuAfaAfgUfgGfasAfsu D1725 S1725 743 UfcCfaCfuUfuAfAfCfuUfgAfUfUfuUfuUfL96 AS1725 1835 aAfaAfaauCfaAfguuAfaAfgUfgGfasAfsu D1726 S1726 744 CfcAfcUfuUfaAfCfUfuGfauuUfuUfuAfL96 AS1726 1836 uAfaAfaAfAfUfcAfaguUfaAfaGfuGfgsAfsa D1727 S1727 745 CfcAfcUfuUfaAfCfUfuGfaUfUfUfuUfuAfL96 AS1727 1837 uAfaAfaaaUfcAfaguUfaAfaGfuGfgsAfsa D1728 S1728 746 CfaCfuUfuAfaCfUfUfgAfuuuUfuUfaAfL96 AS1728 1838 uUfaAfaAfAfAfuCfaagUfuAfaAfgUfgsGfsa D1729 S1729 747 CfaCfuUfuAfaCfUfUfgAfuUfUfUfuUfaAfL96 AS1729 1839 uUfaAfaaaAfuCfaagUfuAfaAfgUfgsGfsa D1730 S1730 748 AfcUfuUfaAfcUfUfGfaUfuuuUfuAfaAfL96 AS1730 1840 uUfuAfaAfAfAfaUfcaaGfuUfaAfaGfusGfsg D1731 S1731 749 AfcUfuUfaAfcUfUfGfaUfuUfUfUfuAfaAfL96 AS1731 1841 uUfuAfaaaAfaUfcaaGfuUfaAfaGfusGfsg D1732 S1732 750 CfuUfuAfaCfuUfGfAfuUfuuuUfaAfaUfL96 AS1732 1842 aUfuUfaAfAfAfaAfucaAfgUfuAfaAfgs Ufsg D1733 S1733 751 CfuUfuAfaCfuUfGfAfuUfuUfUfUfaAfaUfL96 AS1733 1843 aUfuUfaaaAfaAfucaAfgUfuAfaAfgsUfsg D1734 S1734 752 UfuUfaAfcUfuGfAfUfuUfuuuAfaAfuUfL96 AS1734 1844 aAfuUfuAfAfAfaAfaucAfaGfuUfaAfasGfsu D1735 S1735 753 UfuUfaAfcUfuGfAfUfuUfuUfUfAfaAfuUfL96 AS1735 1845 aAfuUfuaaAfaAfaucAfaGfuUfaAfasGfsu D1736 S1736 754 UfuAfaCfuUfgAfUfUfuUfuuaAfaUfuCfL96 AS1736 1846 gAfaUfuUfAfAfaAfaauCfaAfgUfuAfasAfsg D1737 S1737 755 UfuAfaCfuUfgAfUfUfuUfuUfAfAfaUfuCfL96 AS1737 1847 gAfaUfuuaAfaAfaauCfaAfgUfuAfasAfsg D1738 S1738 756 UfaAfcUfuGfaUfUfUfuUfuaaAfuUfcCfL96 AS1738 1848 gGfaAfuUfUfAfaAfaaaUfcAfaGfuUfasAfsa D1739 S1739 757 UfaAfcUfuGfaUfUfUfuUfuAfAfAfuUfcCfL96 AS1739 1849 gGfaAfuuuAfaAfaaaUfcAfaGfuUfasAfsa D1740 S1740 758 AfaCfuUfgAfuUfUfUfuUfaaaUfuCfcCfL96 AS1740 1850 gGfgAfaUfUfUfaAfaaaAfuCfaAfgUfusAfsa D1741 S1741 759 AfaCfuUfgAfuUfUfUfuUfaAfAfUfuCfcCfL96 AS1741 1851 gGfgAfauuUfaAfaaaAfuCfaAfgUfusAfsa D1742 S1742 760 AfcUfuGfaUfuUfUfUfuAfaauUfcCfcUfL96 AS1742 1852 aGfgGfaAfUfUfuAfaaaAfaUfcAfaGfusUfsa D1743 S1743 761 AfcUfuGfaUfuUfUfUfuAfaAfUfUfcCfcUfL96 AS1743 1853 aGfgGfaauUfuAfaaaAfaUfcAfaGfusUfsa D1744 S1744 762 CfuUfgAfuUfuUfUfUfaAfauuCfcCfuUfL96 AS1744 1854 aAfgGfgAfAfUfuUfaaaAfaAfuCfaAfgsUfsu D1745 S1745 763 CfuUfgAfuUfuUfUfUfaAfaUfUfCfcCfuUfL96 AS1745 1855 aAfgGfgaaUfuUfaaaAfaAfuCfaAfgsUfsu D1746 S1746 764 UfuGfaUfuUfuUfUfAfaAfuucCfcUfuAfL96 AS1746 1856 uAfaGfgGfAfAfuUfuaaAfaAfaUfcAfasGfsu D1747 S1747 765 UfuGfaUfuUfuUfUfAfaAfuUfCfCfcUfuAfL96 AS1747 1857 uAfaGfggaAfuUfuaaAfaAfaUfcAfasGfsu D1748 S1748 766 UfgAfuUfuUfuUfAfAfaUfuccCfuUfaUfL96 AS1748 1858 aUfaAfgGfGfAfaUfuuaAfaAfaAfuCfasAfsg D1749 S1749 767 UfgAfuUfuUfuUfAfAfaUfuCfCfCfuUfaUfL96 AS1749 1859 aUfaAfgggAfaUfuuaAfaAfaAfuCfasAfsg
D1750 S1750 768 GfaUfuUfuUfuAfAfAfuUfcccUfuAfuUfL96 AS1750 1860 aAfuAfaGfGfGfaAfuuuAfaAfaAfaUfcsAfsa D1751 S1751 769 GfaUfuUfuUfuAfAfAfuUfcCfCfUfuAfuUfL96 AS1751 1861 aAfuAfaggGfaAfuuuAfaAfaAfaUfcsAfsa D1752 S1752 770 AfuUfuUfuUfaAfAfUfuCfccuUfaUfuGfL96 AS1752 1862 cAfaUfaAfGfGfgAfauuUfaAfaAfaAfusCfsa D1753 S1753 771 AfuUfuUfuUfaAfAfUfuCfcCfUfUfaUfuGfL96 AS1753 1863 cAfaUfaagGfgAfauuUfaAfaAfaAfusCfsa D1754 S1754 772 UfuUfuUfuAfaAfUfUfcCfcuuAfuUfgUfL96 AS1754 1864 aCfaAfuAfAfGfgGfaauUfuAfaAfaAfasUfsc D1755 S1755 773 UfuUfuUfuAfaAfUfUfcCfcUfUfAfuUfgUfL96 AS1755 1865 aCfaAfuaaGfgGfaauUfuAfaAfaAfasUfsc D1756 S1756 774 UfuUfuUfaAfaUfUfCfcCfuuaUfuGfuCfL96 AS1756 1866 gAfcAfaUfAfAfgGfgaaUfuUfaAfaAfasAfsu D1757 S1757 775 UfuUfuUfaAfaUfUfCfcCfuUfAfUfuGfuCfL96 AS1757 1867 gAfcAfauaAfgGfgaaUfuUfaAfaAfasAfsu D1758 S1758 776 UfuUfuAfaAfuUfCfCfcUfuauUfgUfcCfL96 AS1758 1868 gGfaCfaAfUfAfaGfggaAfuUfuAfaAfasAfsa D1759 S1759 777 UfuUfuAfaAfuUfCfCfcUfuAfUfUfgUfcCfL96 AS1759 1869 gGfaCfaauAfaGfggaAfuUfuAfaAfasAfsa D1760 S1760 778 UfuUfaAfaUfuCfCfCfuUfauuGfuCfcCfL96 AS1760 1870 gGfgAfcAfAfUfaAfgggAfaUfuUfaAfasAfsa D1761 S1761 779 UfuUfaAfaUfuCfCfCfuUfaUfUfGfuCfcCfL96 AS1761 1871 gGfgAfcaaUfaAfgggAfaUfuUfaAfasAfsa D1762 S1762 780 UfuAfaAfuUfcCfCfUfuAfuugUfcCfcUfL96 AS1762 1872 aGfgGfaCfAfAfuAfaggGfaAfuUfuAfasAfsa D1763 S1763 781 UfuAfaAfuUfcCfCfUfuAfuUfGfUfcCfcUfL96 AS1763 1873 aGfgGfacaAfuAfaggGfaAfuUfuAfasAfsa D1764 S1764 782 UfaAfaUfuCfcCfUfUfaUfuguCfcCfuUfL96 AS1764 1874 aAfgGfgAfCfAfaUfaagGfgAfaUfuUfasAfsa D1765 S1765 783 UfaAfaUfuCfcCfUfUfaUfuGfUfCfcCfuUfL96 AS1765 1875 aAfgGfgacAfaUfaagGfgAfaUfuUfasAfsa D1766 S1766 784 AfaAfuUfcCfcUfUfAfuUfgucCfcUfuCfL96 AS1766 1876 gAfaGfgGfAfCfaAfuaaGfgGfaAfuUfusAfsa D1767 S1767 785 AfaAfuUfcCfcUfUfAfuUfgUfCfCfcUfuCfL96 AS1767 1877 gAfaGfggaCfaAfuaaGfgGfaAfuUfusAfsa D1768 S1768 786 AfaUfuCfcCfuUfAfUfuGfuccCfuUfcCfL96 AS1768 1878 gGfaAfgGfGfAfcAfauaAfgGfgAfaUfusUfsa D1769 S1769 787 AfaUfuCfcCfuUfAfUfuGfuCfCfCfuUfcCfL96 AS1769 1879 gGfaAfgggAfcAfauaAfgGfgAfaUfusUfsa D1770 S1770 788 AfuUfcCfcUfuAfUfUfgUfcccUfuCfcAfL96 AS1770 1880 uGfgAfaGfGfGfaCfaauAfaGfgGfaAfusUfsu D1771 S1771 789 AfuUfcCfcUfuAfUfUfgUfcCfCfUfuCfcAfL96 AS1771 1881 uGfgAfaggGfaCfaauAfaGfgGfaAfusUfsu D1772 S1772 790 UfuCfcCfuUfaUfUfGfuCfccuUfcCfaAfL96 AS1772 1882 uUfgGfaAfGfGfgAfcaaUfaAfgGfgAfasUfsu D1773 S1773 791 UfuCfcCfuUfaUfUfGfuCfcCfUfUfcCfaAfL96 AS1773 1883 uUfgGfaagGfgAfcaaUfaAfgGfgAfasUfsu D1774 S1774 792 UfcCfcUfuAfuUfGfUfcCfcuuCfcAfaAfL96 AS1774 1884 uUfuGfgAfAfGfgGfacaAfuAfaGfgGfasAfsu D1775 S1775 793 UfcCfcUfuAfuUfGfUfcCfcUfUfCfcAfaAfL96 AS1775 1885 uUfuGfgaaGfgGfacaAfuAfaGfgGfasAfsu D1776 S1776 794 CfcCfuUfaUfuGfUfCfcCfuucCfaAfaAfL96 AS1776 1886 uUfuUfgGfAfAfgGfgacAfaUfaAfgGfgsAfsa D1777 S1777 795 CfcCfuUfaUfuGfUfCfcCfuUfCfCfaAfaAfL96 AS1777 1887 uUfuUfggaAfgGfgacAfaUfaAfgGfgsAfsa D1778 S1778 796 CfcUfuAfuUfgUfCfCfcUfuccAfaAfaAfL96 AS1778 1888 uUfuUfuGfGfAfaGfggaCfaAfuAfaGfgsGfsa D1779 S1779 797 CfcUfuAfuUfgUfCfCfcUfuCfCfAfaAfaAfL96 AS1779 1889 uUfuUfuggAfaGfggaCfaAfuAfaGfgsGfsa D1780 S1780 798 CfuUfaUfuGfuCfCfCfuUfccaAfaAfaAfL96 AS1780 1890 uUfuUfuUfGfGfaAfgggAfcAfaUfaAfgsGfsg D1781 S1781 799 CfuUfaUfuGfuCfCfCfuUfcCfAfAfaAfaAfL96 AS1781 1891 uUfuUfuugGfaAfgggAfcAfaUfaAfgsGfsg D1782 S1782 800 UfuAfuUfgUfcCfCfUfuCfcaaAfaAfaAfL96 AS1782 1892 uUfuUfuUfUfGfgAfaggGfaCfaAfuAfasGfsg D1783 S1783 801 UfuAfuUfgUfcCfCfUfuCfcAfAfAfaAfaAfL96 AS1783 1893 uUfuUfuuuGfgAfaggGfaCfaAfuAfasGfsg D1784 S1784 802 UfaUfuGfuCfcCfUfUfcCfaaaAfaAfaAfL96 AS1784 1894 uUfuUfuUfUfUfgGfaagGfgAfcAfaUfasAfsg D1785 S1785 803 UfaUfuGfuCfcCfUfUfcCfaAfAfAfaAfaAfL96 AS1785 1895 uUfuUfuuuUfgGfaagGfgAfcAfaUfasAfsg D1786 S1786 804 AfuUfgUfcCfcUfUfCfcAfaaaAfaAfaGfL96 AS1786 1896 cUfuUfuUfUfUfuGfgaaGfgGfaCfaAfusAfsa D1787 S1787 805 AfuUfgUfcCfcUfUfCfcAfaAfAfAfaAfaGfL96 AS1787 1897 cUfuUfuuuUfuGfgaaGfgGfaCfaAfusAfsa D1788 S1788 806 UfuGfuCfcCfuUfCfCfaAfaaaAfaAfgAfL96 AS1788 1898 uCfuUfuUfUfUfuUfggaAfgGfgAfcAfasUfsa D1789 S1789 807 UfuGfuCfcCfuUfCfCfaAfaAfAfAfaAfgAfL96 AS1789 1899 uCfuUfuuuUfuUfggaAfgGfgAfcAfasUfsa D1790 S1790 808 UfgUfcCfcUfuCfCfAfaAfaaaAfaGfaGfL96 AS1790 1900 cUfcUfuUfUfUfuUfuggAfaGfgGfaCfasAfsu D1791 S1791 809 UfgUfcCfcUfuCfCfAfaAfaAfAfAfaGfaGfL96 AS1791 1901 cUfcUfuuuUfuUfuggAfaGfgGfaCfasAfsu D1792 S1792 810 GfuCfcCfuUfcCfAfAfaAfaaaAfgAfgAfL96 AS1792 1902 uCfuCfuUfUfUfuUfuugGfaAfgGfgAfcsAfsa D1793 S1793 811 GfuCfcCfuUfcCfAfAfaAfaAfAfAfgAfgAfL96 AS1793 1903 uCfuCfuuuUfuUfuugGfaAfgGfgAfcsAfsa D1794 S1794 812 UfcCfcUfuCfcAfAfAfaAfaaaGfaGfaAfL96 AS1794 1904 uUfcUfcUfUfUfuUfuuuGfgAfaGfgGfasCfsa D1795 S1795 813 UfcCfcUfuCfcAfAfAfaAfaAfAfGfaGfaAfL96 AS1795 1905 uUfcUfcuuUfuUfuuuGfgAfaGfgGfasCfsa D1796 S1796 814 CfcCfuUfcCfaAfAfAfaAfaagAfgAfaUfL96 AS1796 1906 aUfuCfuCfUfUfuUfuuuUfgGfaAfgGfgsAfsc D1797 S1797 815 CfcCfuUfcCfaAfAfAfaAfaAfGfAfgAfaUfL96 AS1797 1907 aUfuCfucuUfuUfuuuUfgGfaAfgGfgsAfsc D1798 S1798 816 CfcUfuCfcAfaAfAfAfaAfagaGfaAfuCfL96 AS1798 1908 gAfuUfcUfCfUfuUfuuuUfuGfgAfaGfgsGfsa D1799 S1799 817 CfcUfuCfcAfaAfAfAfaAfaGfAfGfaAfuCfL96 AS1799 1909 gAfuUfcucUfuUfuuuUfuGfgAfaGfgsGfsa D1800 S1800 818 CfuUfcCfaAfaAfAfAfaAfgagAfaUfcAfL96 AS1800 1910 uGfaUfuCfUfCfuUfuuuUfuUfgGfaAfgsGfsg D1801 S1801 819 CfuUfcCfaAfaAfAfAfaAfgAfGfAfaUfcAfL96 AS1801 1911 uGfaUfucuCfuUfuuuUfuUfgGfaAfgsGfsg D1802 S1802 820 UfuCfcAfaAfaAfAfAfaGfagaAfuCfaAfL96 AS1802 1912 uUfgAfuUfCfUfcUfuuuUfuUfuGfgAfasGfsg D1803 S1803 821 UfuCfcAfaAfaAfAfAfaGfaGfAfAfuCfaAfL96 AS1803 1913 uUfgAfuucUfcUfuuuUfuUfuGfgAfasGfsg D1804 S1804 822 UfcCfaAfaAfaAfAfAfgAfgaaUfcAfaAfL96 AS1804 1914 uUfuGfaUfUfCfuCfuuuUfuUfuUfgGfasAfsg D1805 S1805 823 UfcCfaAfaAfaAfAfAfgAfgAfAfUfcAfaAfL96 AS1805 1915 uUfuGfauuCfuCfuuuUfuUfuUfgGfasAfsg D1806 S1806 824 CfcAfaAfaAfaAfAfGfaGfaauCfaAfaAfL96 AS1806 1916 uUfuUfgAfUfUfcUfcuuUfuUfuUfuGfgsAfsa D1807 S1807 825 CfcAfaAfaAfaAfAfGfaGfaAfUfCfaAfaAfL96 AS1807 1917 uUfuUfgauUfcUfcuuUfuUfuUfuGfgsAfsa D1808 S1808 826 CfaAfaAfaAfaAfGfAfgAfaucAfaAfaUfL96 AS1808 1918 aUfuUfuGfAfUfuCfucuUfuUfuUfuUfgsGfsa D1809 S1809 827 CfaAfaAfaAfaAfGfAfgAfaUfCfAfaAfaUfL96 AS1809 1919 aUfuUfugaUfuCfucuUfuUfuUfuUfgsGfsa D1810 S1810 828 AfaAfaAfaAfaGfAfGfaAfucaAfaAfuUfL96 AS1810 1920 aAfuUfuUfGfAfuUfcucUfuUfuUfuUfusGfsg D1811 S1811 829 AfaAfaAfaAfaGfAfGfaAfuCfAfAfaAfuUfL96 AS1811 1921 aAfuUfuugAfuUfcucUfuUfuUfuUfusGfsg D1812 S1812 830 AfaAfaAfaAfgAfGfAfaUfcaaAfaUfuUfL96 AS1812 1922 aAfaUfuUfUfGfaUfucuCfuUfuUfuUfusUfsg D1813 S1813 831 AfaAfaAfaAfgAfGfAfaUfcAfAfAfaUfuUfL96 AS1813 1923 aAfaUfuuuGfaUfucuCfuUfuUfuUfusUfsg D1814 S1814 832 AfaAfaAfaGfaGfAfAfuCfaaaAfuUfuUfL96 AS1814 1924 aAfaAfuUfUfUfgAfuucUfcUfuUfuUfusUfsu D1815 S1815 833 AfaAfaAfaGfaGfAfAfuCfaAfAfAfuUfuUfL96 AS1815 1925 aAfaAfuuuUfgAfuucUfcUfuUfuUfusUfsu D1816 S1816 834 AfaAfaAfgAfgAfAfUfcAfaaaUfuUfuAfL96 AS1816 1926 uAfaAfaUfUfUfuGfauuCfuCfuUfuUfusUfsu D1817 S1817 835 AfaAfaAfgAfgAfAfUfcAfaAfAfUfuUfuAfL96 AS1817 1927 uAfaAfauuUfuGfauuCfuCfuUfuUfusUfsu D1818 S1818 836 AfaAfaGfaGfaAfUfCfaAfaauUfuUfaCfL96 AS1818 1928 gUfaAfaAfUfUfuUfgauUfcUfcUfuUfusUfsu D1819 S1819 837 AfaAfaGfaGfaAfUfCfaAfaAfUfUfuUfaCfL96 AS1819 1929 gUfaAfaauUfuUfgauUfcUfcUfuUfusUfsu D1820 S1820 838 AfaAfgAfgAfaUfCfAfaAfauuUfuAfcAfL96 AS1820 1930 uGfuAfaAfAfUfuUfugaUfuCfuCfuUfusUfsu D1821 S1821 839 AfaAfgAfgAfaUfCfAfaAfaUfUfUfuAfcAfL96 AS1821 1931 uGfuAfaaaUfuUfugaUfuCfuCfuUfusUfsu D1822 S1822 840 AfaGfaGfaAfuCfAfAfaAfuuuUfaCfaAfL96 AS1822 1932 uUfgUfaAfAfAfuUfuugAfuUfcUfcUfusUfsu D1823 S1823 841 AfaGfaGfaAfuCfAfAfaAfuUfUfUfaCfaAfL96 AS1823 1933 uUfgUfaaaAfuUfuugAfuUfcUfcUfusUfsu D1824 S1824 842 AfgAfgAfaUfcAfAfAfaUfuuuAfcAfaAfL96 AS1824 1934 uUfuGfuAfAfAfaUfuuuGfaUfuCfuCfusUfsu D1825 S1825 843 AfgAfgAfaUfcAfAfAfaUfuUfUfAfcAfaAfL96 AS1825 1935 uUfuGfuaaAfaUfuuuGfaUfuCfuCfusUfsu D1826 S1826 844 GfaGfaAfuCfaAfAfAfuUfuuaCfaAfaGfL96 AS1826 1936 cUfuUfgUfAfAfaAfuuuUfgAfuUkUfcsUfsu D1827 S1827 845 GfaGfaAfuCfaAfAfAfuUfuUfAfCfaAfaGfL96 AS1827 1937 cUfuUfguaAfaAfuuuUfgAfuUkUfcsUfsu D1828 S1828 846 AfgAfaUfcAfaAfAfUfuUfuacAfaAfgAfL96 AS1828 1938 uCfuUfuGfUfAfaAfauuUfuGfaUfuCfusCfsu D1829 S1829 847 AfgAfaUfcAfaAfAfUfuUfuAfCfAfaAfgAfL96 AS1829 1939 uCfuUfuguAfaAfauuUfuGfaUfuCfusCfsu D1830 S1830 848 GfaAfuCfaAfaAfUfUfuUfacaAfaGfaAfL96 AS1830 1940 uUfcUfuUfGfUfaAfaauUfuUfgAfuUfcsUfsc D1831 S1831 849 GfaAfuCfaAfaAfUfUfuUfaCfAfAfaGfaAfL96 AS1831 1941 uUfcUfuugUfaAfaauUfuUfgAfuUfcsUfsc D1832 S1832 850 AfaUfcAfaAfaUfUfUfuAfcaaAfgAfaUfL96 AS1832 1942 aUfuCfuUfUfGfuAfaaaUfuUfuGfaUfusCfsu D1833 S1833 851 AfaUfcAfaAfaUfUfUfuAfcAfAfAfgAfaUfL96 AS1833 1943
aUfuCfuuuGfuAfaaaUfuUfuGfaUfusCfsu D1834 S1834 852 AfuCfaAfaAfuUfUfUfaCfaaaGfaAfuCfL96 AS1834 1944 gAfuUfcUfUfUfgUfaaaAfuUfuUfgAfusUfsc D1835 S1835 853 AfuCfaAfaAfuUfUfUfaCfaAfAfGfaAfuCfL96 AS1835 1945 gAfuUfcuuUfgUfaaaAfuUfuUfgAfusUfsc D1836 S1836 854 UfcAfaAfaUfuUfUfAfcAfaagAfaUfcAfL96 AS1836 1946 uGfaUfuCfUfUfuGfuaaAfaUfuUfuGfasUfsu D1837 S1837 855 UfcAfaAfaUfuUfUfAfcAfaAfGfAfaUfcAfL96 AS1837 1947 uGfaUfucuUfuGfuaaAfaUfuUfuGfasUfsu D1838 S1838 856 CfaAfaAfuUfuUfAfCfaAfagaAfuCfaAfL96 AS1838 1948 uUfgAfuUfCfUfuUfguaAfaAfuUfuUfgsAfsu D1839 S1839 857 CfaAfaAfuUfuUfAfCfaAfaGfAfAfuCfaAfL96 AS1839 1949 uUfgAfuucUfuUfguaAfaAfuUfuUfgsAfsu D1840 S1840 858 AfaAfaUfuUfuAfCfAfaAfgaaUfcAfaAfL96 AS1840 1950 uUfuGfaUfUfCfuUfuguAfaAfaUfuUfusGfsa D1841 S1841 859 AfaAfaUfuUfuAfCfAfaAfgAfAfUfcAfaAfL96 AS1841 1951 uUfuGfauuCfuUfuguAfaAfaUfuUfusGfsa D1842 S1842 860 AfaAfuUfuUfaCfAfAfaGfaauCfaAfaGfL96 AS1842 1952 cUfuUfgAfUfUfcUfuugUfaAfaAfuUfusUfsg D1843 S1843 861 AfaAfuUfuUfaCfAfAfaGfaAfUfCfaAfaGfL96 AS1843 1953 cUfuUfgauUfcUfuugUfaAfaAfuUfusUfsg D1844 S1844 862 AfaUfuUfuAfcAfAfAfgAfaucAfaAfgGfL96 AS1844 1954 cCfuUfuGfAfUfuCfuuuGfuAfaAfaUfusUfsu D1845 S1845 863 AfaUfuUfuAfcAfAfAfgAfaUfCfAfaAfgGfL96 AS1845 1955 cCfuUfugaUfuCfuuuGfuAfaAfaUfusUfsu D1846 S1846 864 AfuUfuUfaCfaAfAfGfaAfucaAfaGfgAfL96 AS1846 1956 uCfcUfuUfGfAfuUfcuuUfgUfaAfaAfusUfsu D1847 S1847 865 AfuUfuUfaCfaAfAfGfaAfuCfAfAfaGfgAfL96 AS1847 1957 uCfcUfuugAfuUfcuuUfgUfaAfaAfusUfsu D1848 S1848 866 UfuUfuAfcAfaAfGfAfaUfcaaAfgGfaAfL96 AS1848 1958 uUfcCfuUfUfGfaUfucuUfuGfuAfaAfasUfsu D1849 S1849 867 UfuUfuAfcAfaAfGfAfaUfcAfAfAfgGfaAfL96 AS1849 1959 uUfcCfuuuGfaUfucuUfuGfuAfaAfasUfsu D1850 S1850 868 UfuUfaCfaAfaGfAfAfuCfaaaGfgAfaUfL96 AS1850 1960 aUfuCfcUfUfUfgAfuucUfuUfgUfaAfasAfsu D1851 S1851 869 UfuUfaCfaAfaGfAfAfuCfaAfAfGfgAfaUfL96 AS1851 1961 aUfuCfcuuUfgAfuucUfuUfgUfaAfasAfsu D1852 S1852 870 UfuAfcAfaAfgAfAfUfcAfaagGfaAfuUfL96 AS1852 1962 aAfuUfcCfUfUfuGfauuCfuUfuGfuAfasAfsa D1853 S1853 871 UfuAfcAfaAfgAfAfUfcAfaAfGfGfaAfuUfL96 AS1853 1963 aAfuUfccuUfuGfauuCfuUfuGfuAfasAfsa D1854 S1854 872 UfaCfaAfaGfaAfUfCfaAfaggAfaUfuCfL96 AS1854 1964 gAfaUfuCfCfUfuUfgauUfcUfuUfgUfasAfsa D1855 S1855 873 UfaCfaAfaGfaAfUfCfaAfaGfGfAfaUfuCfL96 AS1855 1965 gAfaUfuccUfuUfgauUfcUfuUfgUfasAfsa D1856 S1856 874 AfcAfaAfgAfaUfCfAfaAfggaAfuUfcUfL96 AS1856 1966 aGfaAfuUfCfCfuUfugaUfuCfuUfuGfusAfsa D1857 S1857 875 AfcAfaAfgAfaUfCfAfaAfgGfAfAfuUfcUfL96 AS1857 1967 aGfaAfuucCfuUfugaUfuCfuUfuGfusAfsa D1858 S1858 876 CfaAfaGfaAfuCfAfAfaGfgaaUfuCluAfL96 AS1858 1968 uAfgAfaUfUfCfcUfuugAfuUfcUfuUfgsUfsa D1859 S1859 877 CfaAfaGfaAfuCfAfAfaGfgAfAfUfuCluAfL96 AS1859 1969 uAfgAfauuCfcUfuugAfuUfcUfuUfgsUfsa D1860 S1860 878 AfaAfgAfaUfcAfAfAfgGfaauUfcUfaGfL96 AS1860 1970 cUfaGfaAfUfUfcCfuuuGfaUfuCfuUfusGfsu D1861 S1861 879 AfaAfgAfaUfcAfAfAfgGfaAfUfUfcUfaGfL96 AS1861 1971 cUfaGfaauUfcCfuuuGfaUfuCfuUfusGfsu D1862 S1862 880 AfaGfaAfuCfaAfAfGfgAfauuCluAfgAfL96 AS1862 1972 uCluAfgAfAfUfuCfcuuUfgAfuUfcUfusUfsg D1863 S1863 881 AfaGfaAfuCfaAfAfGfgAfaUfUfCluAfgAfL96 AS1863 1973 uCfuAfgaaUfuCfcuuUfgAfuUfcUfusUfsg D1864 S1864 882 AfgAfaUfcAfaAfGfGfaAfuucUfaGfaAfL96 AS1864 1974 uUfcUfaGfAfAfuUfccuUfuGfaUfuCfusUfsu D1865 S1865 883 AfgAfaUfcAfaAfGfGfaAfuUfCfUfaGfaAfL96 AS1865 1975 uUfcUfagaAfuUfccuUfuGfaUfuCfusUfsu D1866 S1866 884 GfaAfuCfaAfaGfGfAfaUfucuAfgAfaAfL96 AS1866 1976 uUfuCfuAfGfAfaUfuccUfuUfgAfuUfcsUfsu D1867 S1867 885 GfaAfuCfaAfaGfGfAfaUfuCfUfAfgAfaAfL96 AS1867 1977 uUfuCfuagAfaUfuccUfuUfgAfuUfcsUfsu D1868 S1868 886 AfaUfcAfaAfgGfAfAfuUfcuaGfaAfaGfL96 AS1868 1978 cUfuUfcUfAfGfaAfuucCfuUfuGfaUfusCfsu D1869 S1869 887 AfaUfcAfaAfgGfAfAfuUfcUfAfGfaAfaGfL96 AS1869 1979 cUfuUfcuaGfaAfuucCfuUfuGfaUfusCfsu D1870 S1870 888 AfuCfaAfaGfgAfAfUfuCluagAfaAfgUfL96 AS1870 1980 aCfuUfuCfUfAfgAfauuCfcUfuUfgAfusUfsc D1871 S1871 889 AfuCfaAfaGfgAfAfUfuCluAfGfAfaAfgUfL96 AS1871 1981 aCfuUfucuAfgAfauuCfcUfuUfgAfusUfsc D1872 S1872 890 UfcAfaAfgGfaAfUfUfcUfagaAfaGfuAfL96 AS1872 1982 uAfcUfuUfCfUfaGfaauUfcCfuUfuGfasUfsu D1873 S1873 891 UfcAfaAfgGfaAfUfUfcUfaGfAfAfaGfuAfL96 AS1873 1983 uAfcUfuucUfaGfaauUfcCfuUfuGfasUfsu D1874 S1874 892 CfaAfaGfgAfaUfUfCfuAfgaaAfgUfaUfL96 AS1874 1984 aUfaCfuUfUfCfuAfgaaUfuCfcUfuUfgsAfsu D1875 S1875 893 CfaAfaGfgAfaUfUfCfuAfgAfAfAfgUfaUfL96 AS1875 1985 aUfaCfuuuCfuAfgaaUfuCfcUfuUfgsAfsu D1876 S1876 894 AfaAfgGfaAfuUfCfUfaGfaaaGfuAfuCfL96 AS1876 1986 gAfuAfcUfUfUfcUfagaAfuUfcCfuUfusGfsa D1877 S1877 895 AfaAfgGfaAfuUfCfUfaGfaAfAfGfuAfuCfL96 AS1877 1987 gAfuAfcuuUfcUfagaAfuUfcCfuUfusGfsa D1878 S1878 896 AfaGfgAfaUfuCfUfAfgAfaagUfaUfcUfL96 AS1878 1988 aGfaUfaCfUfUfuCfuagAfaUfuCfcUfusUfsg D1879 S1879 897 AfaGfgAfaUfuCfUfAfgAfaAfGfUfaUfcUfL96 AS1879 1989 aGfaUfacuUfuCfuagAfaUfuCfcUfusUfsg D1880 S1880 898 AfgGfaAfuUfcUfAfGfaAfaguAfuCfuGfL96 AS1880 1990 cAfgAfuAfCfUfuUfcuaGfaAfuUfcCfusUfsu D1881 S1881 899 AfgGfaAfuUfcUfAfGfaAfaGfUfAfuCfuGfL96 AS1881 1991 cAfgAfuacUfuUfcuaGfaAfuUfcCfusUfsu D1882 S1882 900 GfgAfaUfuCluAfGfAfaAfguaUfcUfgGfL96 AS1882 1992 cCfaGfaUfAfCfuUfucuAfgAfaUfuCfcsUfsu D1883 S1883 901 GfgAfaUfuCluAfGfAfaAfgUfAfUfcUfgGfL96 AS1883 1993 cCfaGfauaCfuUfucuAfgAfaUfuCfcsUfsu D1884 S1884 902 GfaAfuUfcUfaGfAfAfaGfuauCfuGfgGfL96 AS1884 1994 cCfcAfgAfUfAfcUfuucUfaGfaAfuUfcsCfsu D1885 S1885 903 GfaAfuUfcUfaGfAfAfaGfuAfUfCfuGfgGfL96 AS1885 1995 cCfcAfgauAfcUfuucUfaGfaAfuUfcsCfsu D1886 S1886 904 AfaUfuCluAfgAfAfAfgUfaucUfgGfgCfL96 AS1886 1996 gCfcCfaGfAfUfaCfuuuCluAfgAfaUfusCfsc D1887 S1887 905 AfaUfuCluAfgAfAfAfgUfaUfCfUfgGfgCfL96 AS1887 1997 gCfcCfagaUfaCfuuuCluAfgAfaUfusCfsc D1888 S1888 906 AfuUfcUfaGfaAfAfGfuAfucuGfgGfcAfL96 AS1888 1998 uGfcCfcAfGfAfuAfcuuUfcUfaGfaAfusUfsc D1889 S1889 907 AfuUfcUfaGfaAfAfGfuAfuCfUfGfgGfcAfL96 AS1889 1999 uGfcCfcagAfuAfcuuUfcUfaGfaAfusUfsc D1890 S1890 908 UfuCfuAfgAfaAfGfUfaUfcugGfgCfaGfL96 AS1890 2000 cUfgCfcCfAfGfaUfacuUfuCfuAfgAfasUfsu D1891 S1891 909 UfuCluAfgAfaAfGfUfaUfcUfGfGfgCfaGfL96 AS1891 2001 cUfgCfccaGfaUfacuUfuCluAfgAfasUfsu D1892 S1892 910 UfcUfaGfaAfaGfUfAfuCfuggGfcAfgAfL96 AS1892 2002 uCfuGfcCfCfAfgAfuacUfuUfcUfaGfasAfsu D1893 S1893 911 UfcUfaGfaAfaGfUfAfuCfuGfGfGfcAfgAfL96 AS1893 2003 uCfuGfcccAfgAfuacUfuUfcUfaGfasAfsu D1894 S1894 912 CfuAfgAfaAfgUfAfUfcUfgggCfaGfaAfL96 AS1894 2004 uUfcUfgCfCfCfaGfauaCfuUfuCfuAfgsAfsa D1895 S1895 913 CfuAfgAfaAfgUfAfUfcUfgGfGfCfaGfaAfL96 AS1895 2005 uUfcUfgccCfaGfauaCfuUfuCfuAfgsAfsa D1896 S1896 914 UfaGfaAfaGfuAfUfCfuGfggcAfgAfaCfL96 AS1896 2006 gUfuCfuGfCfCfcAfgauAfcUfuUkUfasGfsa D1897 S1897 915 UfaGfaAfaGfuAfUfCfuGfgGfCfAfgAfaCfL96 AS1897 2007 gUfuCfugcCfcAfgauAfcUfuUfcUfasGfsa D1898 S1898 916 AfgAfaAfgUfaUfCfUfgGfgcaGfaAfcGfL96 AS1898 2008 cGfuUfcUfGfCfcCfagaUfaCfuUfuCfusAfsg D1899 S1899 917 AfgAfaAfgUfaUfCfUfgGfgCfAfGfaAfcGfL96 AS1899 2009 cGfuUfcugCfcCfagaUfaCfuUfuCfusAfsg D1900 S1900 918 GfaAfaGfuAfuCfUfGfgGfcagAfaCfgCfL96 AS1900 2010 gCfgUfuCfUfGfcCfcagAfuAfcUfuUfcsUfsa D1901 S1901 919 GfaAfaGfuAfuCfUfGfgGfcAfGfAfaCfgCfL96 AS1901 2011 gCfgUfucuGfcCfcagAfuAfcUfuUfcsUfsa D1902 S1902 920 AfaAfgUfaUfcUfGfGfgCfagaAfcGfcUfL96 AS1902 2012 aGfcGfuUfCfUfgCfccaGfaUfaCfuUfusCfsu D1903 S1903 921 AfaAfgUfaUfcUfGfGfgCfaGfAfAfcGfcUfL96 AS1903 2013 aGfcGfuucUfgCfccaGfaUfaCfuUfusCfsu D1904 S1904 922 AfaGfuAfuCfuGfGfGfcAfgaaCfgCluAfL96 AS1904 2014 uAfgCfgUfUfCfuGfcccAfgAfuAfcUfusUfsc D1905 S1905 923 AfaGfuAfuCfuGfGfGfcAfgAfAfCfgCluAfL96 AS1905 2015 uAfgCfguuCfuGfcccAfgAfuAfcUfusUfsc D1906 S1906 924 AfgUfaUfcUfgGfGfCfaGfaacGfcUfaGfL96 AS1906 2016 cUfaGfcGfUfUfcUfgccCfaGfaUfaCfusUfsu D1907 S1907 925 AfgUfaUfcUfgGfGfCfaGfaAfCfGfcUfaGfL96 AS1907 2017 cUfaGfcguUfcUfgccCfaGfaUfaCfusUfsu D1908 S1908 926 GfuAfuCfuGfgGfCfAfgAfacgCluAfgGfL96 AS1908 2018 cCfuAfgCfGfUfuCfugcCfcAfgAfuAfcsUfsu D1909 S1909 927 GfuAfuCfuGfgGfCfAfgAfaCfGfCluAfgGfL96 AS1909 2019 cCfuAfgcgUfuCfugcCfcAfgAfuAfcsUfsu D1910 S1910 928 UfaUfcUfgGfgCfAfGfaAfcgcUfaGfgAfL96 AS1910 2020 uCfcUfaGfCfGfuUfcugCfcCfaGfaUfasCfsu D1911 S1911 929 UfaUfcUfgGfgCfAfGfaAfcGfCfUfaGfgAfL96 AS1911 2021 uCfcUfagcGfuUfcugCfcCfaGfaUfasCfsu D1912 S1912 930 AfuCfuGfgGfcAfGfAfaCfgcuAfgGfaGfL96 AS1912 2022 cUfcCfuAfGfCfgUfucuGfcCfcAfgAfusAfsc D1913 S1913 931 AfuCfuGfgGfcAfGfAfaCfgClUfAfgGfaGfL96 AS1913 2023 cUfcCfuagCfgUfucuGfcCfcAfgAfusAfsc D1914 S1914 932 UfcUfgGfgCfaGfAfAfcGfcuaGfgAfgAfL96 AS1914 2024 uCfuCfcUfAfGfcGfuucUfgCfcCfaGfasUfsa D1915 S1915 933 UfcUfgGfgCfaGfAfAfcGfcUfAfGfgAfgAfL96 AS1915 2025 uCfuCfcuaGfcGfuucUfgCfcCfaGfasUfsa D1916 S1916 934 CfuGfgGfcAfgAfAfCfgCluagGfaGfaGfL96 AS1916 2026 cUfcUfcCfUfAfgCfguuCfuGfcCfcAfgsAfsu
D1917 S1917 935 CfuGfgGfcAfgAfAfCfgCluAfGfGfaGfaGfL96 AS1917 2027 cUfcUfccuAfgCfguuCfuGfcCfcAfgsAfsu D1918 S1918 936 UfgGfgCfaGfaAfCfGfcUfaggAfgAfgAfL96 AS1918 2028 uCfuCfuCfCfUfaGfcguUfcUfgCfcCfasGfsa D1919 S1919 937 UfgGfgCfaGfaAfCfGfcUfaGfGfAfgAfgAfL96 AS1919 2029 uCfuCfuccUfaGfcguUfcUfgCfcCfasGfsa D1920 S1920 938 GfgGfcAfgAfaCfGfCfuAfggaGfaGfaUfL96 AS1920 2030 aUfcUfcUfCfCluAfgcgUfuCfuGfcCfcsAfsg D1921 S1921 939 GfgGfcAfgAfaCfGfCluAfgGfAfGfaGfaUfL96 AS1921 2031 aUfcUfcucCfuAfgcgUfuCfuGfcCfcsAfsg D1922 S1922 940 GfgCfaGfaAfcGfCfUfaGfgagAfgAfuCfL96 AS1922 2032 gAfuCfuCfUfCfcUfagcGfuUfcUfgCfcsCfsa D1923 S1923 941 GfgCfaGfaAfcGfCfUfaGfgAfGfAfgAfuCfL96 AS1923 2033 gAfuCfucuCfcUfagcGfuUfcUfgCfcsCfsa D1924 S1924 942 GfcAfgAfaCfgClUfAfgGfagaGfaUfcCfL96 AS1924 2034 gGfaUfcUfCfUfcCfuagCfgUfuCfuGfcsCfsc D1925 S1925 943 GfcAfgAfaCfgClUfAfgGfaGfAfGfaUfcCfL96 AS1925 2035 gGfaUfcucUfcCfuagCfgUfuCfuGfcsCfsc D1926 S1926 944 CfaGfaAfcGfcUfAfGfgAfgagAfuCfcAfL96 AS1926 2036 uGfgAfuCfUfCfuCfcuaGfcGfuUfcUfgsCfsc D1927 S1927 945 CfaGfaAfcGfcUfAfGfgAfgAfGfAfuCfcAfL96 AS1927 2037 uGfgAfucuCfuCfcuaGfcGfuUfcUfgsCfsc D1928 S1928 946 AfgAfaCfgCfuAfGfGfaGfagaUfcCfaAfL96 AS1928 2038 uUfgGfaUfCfUfcUfccuAfgCfgUfuCfusGfsc D1929 S1929 947 AfgAfaCfgCluAfGfGfaGfaGfAfUfcCfaAfL96 AS1929 2039 uUfgGfaucUfcUfccuAfgCfgUfuCfusGfsc D1930 S1930 948 GfaAfcGfcUfaGfGfAfgAfgauCfcAfaAfL96 AS1930 2040 uUfuGfgAfUfCfuCfuccUfaGfcGfuUfcsUfsg D1931 S1931 949 GfaAfcGfcUfaGfGfAfgAfgAfUfCfcAfaAfL96 AS1931 2041 uUfuGfgauCfuCfuccUfaGfcGfuUfcsUfsg D1932 S1932 950 AfaCfgCluAfgGfAfGfaGfaucCfaAfaUfL96 AS1932 2042 aUfuUfgGfAfUfcUfcucCfuAfgCfgUfusCfsu D1933 S1933 951 AfaCfgCluAfgGfAfGfaGfaUfCfCfaAfaUfL96 AS1933 2043 aUfuUfggaUfcUfcucCfuAfgCfgUfusCfsu D1934 S1934 952 AfcGfcUfaGfgAfGfAfgAfuccAfaAfuUfL96 AS1934 2044 aAfuUfuGfGfAfuCfucuCfcUfaGfcGfusUfsc D1935 S1935 953 AfcGfcUfaGfgAfGfAfgAfuCfCfAfaAfuUfL96 AS1935 2045 aAfuUfuggAfuCfucuCfcUfaGfcGfusUfsc D1936 S1936 954 CfgCluAfgGfaGfAfGfaUfccaAfaUfuUfL96 AS1936 2046 aAfaUfuUfGfGfaUfcucUfcCfuAfgCfgsUfsu D1937 S1937 955 CfgCluAfgGfaGfAfGfaUfcCfAfAfaUfuUfL96 AS1937 2047 aAfaUfuugGfaUfcucUfcCfuAfgCfgsUfsu D1938 S1938 956 GfcUfaGfgAfgAfGfAfuCfcaaAfuUfuCfL96 AS1938 2048 gAfaAfuUfUfGfgAfucuCfuCfcUfaGfcsGfsu D1939 S1939 957 GfcUfaGfgAfgAfGfAfuCfcAfAfAfuUfuCfL96 AS1939 2049 gAfaAfuuuGfgAfucuCfuCfcUfaGfcsGfsu D1940 S1940 958 CfuAfgGfaGfaGfAfUfcCfaaaUfuUfcCfL96 AS1940 2050 gGfaAfaUfUfUfgGfaucUfcUfcCfuAfgsCfsg D1941 S1941 959 CfuAfgGfaGfaGfAfUfcCfaAfAfUfuUfcCfL96 AS1941 2051 gGfaAfauuUfgGfaucUfcUfcCfuAfgsCfsg D1942 S1942 960 UfaGfgAfgAfgAfUfCfcAfaauUfuCfcAfL96 AS1942 2052 uGfgAfaAfUfUfuGfgauCfuCfuCfcUfasGfsc D1943 S1943 961 UfaGfgAfgAfgAfUfCfcAfaAfUfUfuCfcAfL96 AS1943 2053 uGfgAfaauUfuGfgauCfuCfuCfcUfasGfsc D1944 S1944 962 AfgGfaGfaGfaUfCfCfaAfauuUfcCfaUfL96 AS1944 2054 aUfgGfaAfAfUfuUfggaUfcUfcUfcCfusAfsg D1945 S1945 963 AfgGfaGfaGfaUfCfCfaAfaUfUfUfcCfaUfL96 AS1945 2055 aUfgGfaaaUfuUfggaUfcUfcUfcCfusAfsg D1946 S1946 964 GfgAfgAfgAfuCfCfAfaAfuuuCfcAfuUfL96 AS1946 2056 aAfuGfgAfAfAfuUfuggAfuCfuCfuCfcsUfsa D1947 S1947 965 GfgAfgAfgAfuCfCfAfaAfuUfUfCfcAfuUfL96 AS1947 2057 aAfuGfgaaAfuUfuggAfuCfuCfuCfcsUfsa D1948 S1948 966 GfaGfaGfaUfcCfAfAfaUfuucCfaUfuGfL96 AS1948 2058 cAfaUfgGfAfAfaUfuugGfaUfcUfcUfcsCfsu D1949 S1949 967 GfaGfaGfaUfcCfAfAfaUfuUfCfCfaUfuGfL96 AS1949 2059 cAfaUfggaAfaUfuugGfaUfcUfcUfcsCfsu D1950 S1950 968 AfgAfgAfuCfcAfAfAfuUfuccAfuUfgUfL96 AS1950 2060 aCfaAfuGfGfAfaAfuuuGfgAfuCfuCfusCfsc D1951 S1951 969 AfgAfgAfuCfcAfAfAfuUfuCfCfAfuUfgUfL96 AS1951 2061 aCfaAfuggAfaAfuuuGfgAfuCfuCfusCfsc D1952 S1952 970 GfaGfaUfcCfaAfAfUfuUfccaUfuGfuCfL96 AS1952 2062 gAfcAfaUfGfGfaAfauuUfgGfaUfcUfcsUfsc D1953 S1953 971 GfaGfaUfcCfaAfAfUfuUfcCfAfUfuGfuCfL96 AS1953 2063 gAfcAfaugGfaAfauuUfgGfaUfcUfcsUfsc D1954 S1954 972 AfgAfuCfcAfaAfUfUfuCfcauUfgUfcUfL96 AS1954 2064 aGfaCfaAfUfGfgAfaauUfuGfgAfuCfusCfsu D1955 S1955 973 AfgAfuCfcAfaAfUfUfuCfcAfUfUfgUfcUfL96 AS1955 2065 aGfaCfaauGfgAfaauUfuGfgAfuCfusCfsu D1956 S1956 974 GfaUfcCfaAfaUfUfUfcCfauuGfuCfuUfL96 AS1956 2066 aAfgAfcAfAfUfgGfaaaUfuUfgGfaUfcsUfsc D1957 S1957 975 GfaUfcCfaAfaUfUfUfcCfaUfUfGfuCfuUfL96 AS1957 2067 aAfgAfcaaUfgGfaaaUfuUfgGfaUfcsUfsc D1958 S1958 976 AfuCfcAfaAfuUfUfCfcAfuugUfcUfuGfL96 AS1958 2068 cAfaGfaCfAfAfuGfgaaAfuUfuGfgAfusCfsu D1959 S1959 977 AfuCfcAfaAfuUfUfCfcAfuUfGfUfcUfuGfL96 AS1959 2069 cAfaGfacaAfuGfgaaAfuUfuGfgAfusCfsu D1960 S1960 978 UfcCfaAfaUfuUfCfCfaUfuguCfuUfgCfL96 AS1960 2070 gCfaAfgAfCfAfaUfggaAfaUfuUfgGfasUfsc D1961 S1961 979 UfcCfaAfaUfuUfCfCfaUfuGfUfCfuUfgCfL96 AS1961 2071 gCfaAfgacAfaUfggaAfaUfuUfgGfasUfsc D1962 S1962 980 CfcAfaAfuUfuCfCfAfuUfgucUfuGfcAfL96 AS1962 2072 uGfcAfaGfAfCfaAfuggAfaAfuUfuGfgsAfsu D1963 S1963 981 CfcAfaAfuUfuCfCfAfuUfgUfCfUfuGfcAfL96 AS1963 2073 uGfcAfagaCfaAfuggAfaAfuUfuGfgsAfsu D1964 S1964 982 CfaAfaUfuUfcCfAfUfuGfucuUfgCfaAfL96 AS1964 2074 uUfgCfaAfGfAfcAfaugGfaAfaUfuUfgsGfsa D1965 S1965 983 CfaAfaUfuUfcCfAfUfuGfuCfUfUfgCfaAfL96 AS1965 2075 uUfgCfaagAfcAfaugGfaAfaUfuUfgsGfsa D1966 S1966 984 AfaAfuUfuCfcAfUfUfgUfcuuGfcAfaGfL96 AS1966 2076 cUfuGfcAfAfGfaCfaauGfgAfaAfuUfusGfsg D1967 S1967 985 AfaAfuUfuCfcAfUfUfgUfcUfUfGfcAfaGfL96 AS1967 2077 cUfuGfcaaGfaCfaauGfgAfaAfuUfusGfsg D1968 S1968 986 AfaUfuUfcCfaUfUfGfuCfuugCfaAfgCfL96 AS1968 2078 gCfuUfgCfAfAfgAfcaaUfgGfaAfaUfusUfsg D1969 S1969 987 AfaUfuUfcCfaUfUfGfuCfuUfGfCfaAfgCfL96 AS1969 2079 gCfuUfgcaAfgAfcaaUfgGfaAfaUfusUfsg D1970 S1970 988 AfuUfuCfcAfuUfGfUfcUfugcAfaGfcAfL96 AS1970 2080 uGfcUfuGfCfAfaGfacaAfuGfgAfaAfusUfsu D1971 S1971 989 AfuUfuCfcAfuUfGfUfcUfuGfCfAfaGfcAfL96 AS1971 2081 uGfcUfugcAfaGfacaAfuGfgAfaAfusUfsu D1972 S1972 990 UfuUfcCfaUfuGfUfCfuUfgcaAfgCfaAfL96 AS1972 2082 uUfgCfuUfGfCfaAfgacAfaUfgGfaAfasUfsu D1973 S1973 991 UfuUfcCfaUfuGfUfCfuUfgCfAfAfgCfaAfL96 AS1973 2083 uUfgCfuugCfaAfgacAfaUfgGfaAfasUfsu D1974 S1974 992 UfuCfcAfuUfgUfCfUfuGfcaaGfcAfaAfL96 AS1974 2084 uUfuGfcUfUfGfcAfagaCfaAfuGfgAfasAfsu D1975 S1975 993 UfuCfcAfuUfgUfCfUfuGfcAfAfGfcAfaAfL96 AS1975 2085 uUfuGfcuuGfcAfagaCfaAfuGfgAfasAfsu D1976 S1976 994 UfcCfaUfuGfuCfUfUfgCfaagCfaAfaGfL96 AS1976 2086 cUfuUfgCfUfUfgCfaagAfcAfaUfgGfasAfsa D1977 S1977 995 UfcCfaUfuGfuCfUfUfgCfaAfGfCfaAfaGfL96 AS1977 2087 cUfuUfgcuUfgCfaagAfcAfaUfgGfasAfsa D1978 S1978 996 CfcAfuUfgUfcUfUfGfcAfagcAfaAfgCfL96 AS1978 2088 gCfuUfuGfCfUfuGfcaaGfaCfaAfuGfgsAfsa D1979 S1979 997 CfcAfuUfgUfcUfUfGfcAfaGfCfAfaAfgCfL96 AS1979 2089 gCfuUfugcUfuGfcaaGfaCfaAfuGfgsAfsa D1980 S1980 998 CfaUfuGfuCfuUfGfCfaAfgcaAfaGfcAfL96 AS1980 2090 uGfcUfuUfGfCfuUfgcaAfgAfcAfaUfgsGfsa D1981 S1981 999 CfaUfuGfuCfuUfGfCfaAfgCfAfAfaGfcAfL96 AS1981 2091 uGfcUfuugCfuUfgcaAfgAfcAfaUfgsGfsa D1982 S1982 1000 AfuUfgUfcUfuGfCfAfaGfcaaAfgCfaCfL96 AS1982 2092 gUfgCfuUfUfGfcUfugcAfaGfaCfaAfusGfsg D1983 S1983 1001 AfuUfgUfcUfuGfCfAfaGfcAfAfAfgCfaCfL96 AS1983 2093 gUfgCfuuuGfcUfugcAfaGfaCfaAfusGfsg D1984 S1984 1002 UfuGfuCfuUfgCfAfAfgCfaaaGfcAfcGfL96 AS1984 2094 cGfuGfcUfUfUfgCfuugCfaAfgAfcAfasUfsg D1985 S1985 1003 UfuGfuCfuUfgCfAfAfgCfaAfAfGfcAfcGfL96 AS1985 2095 cGfuGfcuuUfgCfuugCfaAfgAfcAfasUfsg D1986 S1986 1004 UfgUfcUfuGfcAfAfGfcAfaagCfaCfgUfL96 AS1986 2096 aCfgUfgCfUfUfuGfcuuGfcAfaGfaCfasAfsu D1987 S1987 1005 UfgUfcUfuGfcAfAfGfcAfaAfGfCfaCfgUfL96 AS1987 2097 aCfgUfgcuUfuGfcuuGfcAfaGfaCfasAfsu D1988 S1988 1006 GfuCfuUfgCfaAfGfCfaAfagcAfcGfuAfL96 AS1988 2098 uAfcGfuGfCfUfuUfgcuUfgCfaAfgAfcsAfsa D1989 S1989 1007 GfuCfuUfgCfaAfGfCfaAfaGfCfAfcGfuAfL96 AS1989 2099 uAfcGfugcUfuUfgcuUfgCfaAfgAfcsAfsa D1990 S1990 1008 UfcUfuGfcAfaGfCfAfaAfgcaCfgUfaUfL96 AS1990 2100 aUfaCfgUfGfCfuUfugcUfuGfcAfaGfasCfsa D1991 S1991 1009 UfcUfuGfcAfaGfCfAfaAfgCfAfCfgUfaUfL96 AS1991 2101 aUfaCfgugCfuUfugcUfuGfcAfaGfasCfsa D1992 S1992 1010 CfuUfgCfaAfgCfAfAfaGfcacGfuAfuUfL96 AS1992 2102 aAfuAfcGfUfGfcUfuugCfuUfgCfaAfgsAfsc D1993 S1993 1011 CfuUfgCfaAfgCfAfAfaGfcAfCfGfuAfuUfL96 AS1993 2103 aAfuAfcguGfcUfuugCfuUfgCfaAfgsAfsc D1994 S1994 1012 UfuGfcAfaGfcAfAfAfgCfacgUfaUfuAfL96 AS1994 2104 uAfaUfaCfGfUfgCfuuuGfcUfuGfcAfasGfsa D1995 S1995 1013 UfuGfcAfaGfcAfAfAfgCfaCfGfUfaUfuAfL96 AS1995 2105 uAfaUfacgUfgCfuuuGfcUfuGfcAfasGfsa D1996 S1996 1014 UfgCfaAfgCfaAfAfGfcAfcguAfuUfaAfL96 AS1996 2106 uUfaAfuAfCfGfuGfcuuUfgCfuUfgCfasAfsg D1997 S1997 1015 UfgCfaAfgCfaAfAfGfcAfcGfUfAfuUfaAfL96 AS1997 2107 uUfaAfuacGfuGfcuuUfgCfuUfgCfasAfsg D1998 S1998 1016 GfcAfaGfcAfaAfGfCfaCfguaUfuAfaAfL96 AS1998 2108 uUfuAfaUfAfCfgUfgcuUfuGfcUfuGfcsAfsa D1999 S1999 1017 GfcAfaGfcAfaAfGfCfaCfgUfAfUfuAfaAfL96 AS1999 2109 uUfuAfauaCfgUfgcuUfuGfcUfuGfcsAfsa D2000 S2000 1018 CfaAfgCfaAfaGfCfAfcGfuauUfaAfaUfL96 AS2000 2110 aUfuUfaAfUfAfcGfugcUfuUfgCfuUfgsCfsa
D2001 S2001 1019 CfaAfgCfaAfaGfCfAfcGfuAfUfUfaAfaUfL96 AS2001 2111 aUfuUfaauAfcGfugcUfuUfgCfuUfgsCfsa D2002 S2002 1020 AfaGfcAfaAfgCfAfCfgUfauuAfaAfuAfL96 AS2002 2112 uAfuUfuAfAfUfaCfgugCfuUfuGfcUfusGfsc D2003 S2003 1021 AfaGfcAfaAfgCfAfCfgUfaUfUfAfaAfuAfL96 AS2003 2113 uAfuUfuaaUfaCfgugCfuUfuGfcUfusGfsc D2004 S2004 1022 AfgCfaAfaGfcAfCfGfuAfuuaAfaUfaUfL96 AS2004 2114 aUfaUfuUfAfAfuAfcguGfcUfuUfgCfusUfsg D2005 S2005 1023 AfgCfaAfaGfcAfCfGfuAfuUfAfAfaUfaUfL96 AS2005 2115 aUfaUfuuaAfuAfcguGfcUfuUfgCfusUfsg D2006 S2006 1024 GfcAfaAfgCfaCfGfUfaUfuaaAfuAfuGfL96 AS2006 2116 cAfuAfuUfUfAfaUfacgUfgCfuUfuGfcsUfsu D2007 S2007 1025 GfcAfaAfgCfaCfGfUfaUfuAfAfAfuAfuGfL96 AS2007 2117 cAfuAfuuuAfaUfacgUfgCfuUfuGfcsUfsu D2008 S2008 1026 CfaAfaGfcAfcGfUfAfuUfaaaUfaUfgAfL96 AS2008 2118 uCfaUfaUfUfUfaAfuacGfuGfcUfuUfgsCfsu D2009 S2009 1027 CfaAfaGfcAfcGfUfAfuUfaAfAfUfaUfgAfL96 AS2009 2119 uCfaUfauuUfaAfuacGfuGfcUfuUfgsCfsu D2010 S2010 1028 AfaAfgCfaCfgUfAfUfuAfaauAfuGfaUfL96 AS2010 2120 aUfcAfuAfUfUfuAfauaCfgUfgCfuUfusGfsc D2011 S2011 1029 AfaAfgCfaCfgUfAfUfuAfaAfUfAfuGfaUfL96 AS2011 2121 aUfcAfuauUfuAfauaCfgUfgCfuUfusGfsc D2012 S2012 1030 AfaGfcAfcGfuAfUfUfaAfauaUfgAfuCfL96 AS2012 2122 gAfuCfaUfAfUfuUfaauAfcGfuGfcUfusUfsg D2013 S2013 1031 AfaGfcAfcGfuAfUfUfaAfaUfAfUfgAfuCfL96 AS2013 2123 gAfuCfauaUfuUfaauAfcGfuGfcUfusUfsg D2014 S2014 1032 AfgCfaCfgUfaUfUfAfaAfuauGfaUfcUfL96 AS2014 2124 aGfaUfcAfUfAfuUfuaaUfaCfgUfgCfusUfsu D2015 S2015 1033 AfgCfaCfgUfaUfUfAfaAfuAfUfGfaUfcUfL96 AS2015 2125 aGfaUfcauAfuUfuaaUfaCfgUfgCfusUfsu D2016 S2016 1034 GfcAfcGfuAfuUfAfAfaUfaugAfuCfuGfL96 AS2016 2126 cAfgAfuCfAfUfaUfuuaAfuAfcGfuGfcsUfsu D2017 S2017 1035 GfcAfcGfuAfuUfAfAfaUfaUfGfAfuCfuGfL96 AS2017 2127 cAfgAfucaUfaUfuuaAfuAfcGfuGfcsUfsu D2018 S2018 1036 CfaCfgUfaUfuAfAfAfuAfugaUfcUfgCfL96 AS2018 2128 gCfaGfaUfCfAfuAfuuuAfaUfaCfgUfgsCfsu D2019 S2019 1037 CfaCfgUfaUfuAfAfAfuAfuGfAfUfcUfgCfL96 AS2019 2129 gCfaGfaucAfuAfuuuAfaUfaCfgUfgsCfsu D2020 S2020 1038 AfcGfuAfuUfaAfAfUfaUfgauCfuGfcAfL96 AS2020 2130 uGfcAfgAfUfCfaUfauuUfaAfuAfcGfusGfsc D2021 S2021 1039 AfcGfuAfuUfaAfAfUfaUfgAfUfCfuGfcAfL96 AS2021 2131 uGfcAfgauCfaUfauuUfaAfuAfcGfusGfsc D2022 S2022 1040 CfgUfaUfuAfaAfUfAfuGfaucUfgCfaGfL96 AS2022 2132 cUfgCfaGfAfUfcAfuauUfuAfaUfaCfgsUfsg D2023 S2023 1041 CfgUfaUfuAfaAfUfAfuGfaUfCfUfgCfaGfL96 AS2023 2133 cUfgCfagaUfcAfuauUfuAfaUfaCfgsUfsg D2024 S2024 1042 GfuAfuUfaAfaUfAfUfgAfucuGfcAfgCfL96 AS2024 2134 gCfuGfcAfGfAfuCfauaUfuUfaAfuAfcsGfsu D2025 S2025 1043 GfuAfuUfaAfaUfAfUfgAfuCfUfGfcAfgCfL96 AS2025 2135 gCfuGfcagAfuCfauaUfuUfaAfuAfcsGfsu D2026 S2026 1044 UfaUfuAfaAfuAfUfGfaUfcugCfaGfcCfL96 AS2026 2136 gGfcUfgCfAfGfaUfcauAfuUfuAfaUfasCfsg D2027 S2027 1045 UfaUfuAfaAfuAfUfGfaUfcUfGfCfaGfcCfL96 AS2027 2137 gGfcUfgcaGfaUfcauAfuUfuAfaUfasCfsg D2028 S2028 1046 AfuUfaAfaUfaUfGfAfuCfugcAfgCfcAfL96 AS2028 2138 uGfgCfuGfCfAfgAfucaUfaUfuUfaAfusAfsc D2029 S2029 1047 AfuUfaAfaUfaUfGfAfuCfuGfCfAfgCfcAfL96 AS2029 2139 uGfgCfugcAfgAfucaUfaUfuUfaAfusAfsc D2030 S2030 1048 UfuAfaAfuAfuGfAfUfcUfgcaGfcCfaUfL96 AS2030 2140 aUfgGfcUfGfCfaGfaucAfuAfuUfuAfasUfsa D2031 S2031 1049 UfuAfaAfuAfuGfAfUfcUfgCfAfGfcCfaUfL96 AS2031 2141 aUfgGfcugCfaGfaucAfuAfuUfuAfasUfsa D2032 S2032 1050 UfaAfaUfaUfgAfUfCfuGfcagCfcAfuUfL96 AS2032 2142 aAfuGfgCfUfGfcAfgauCfaUfaUfuUfasAfsu D2033 S2033 1051 UfaAfaUfaUfgAfUfCfuGfcAfGfCfcAfuUfL96 AS2033 2143 aAfuGfgcuGfcAfgauCfaUfaUfuUfasAfsu D2034 S2034 1052 AfaAfuAfuGfaUfCfUfgCfagcCfaUfuAfL96 AS2034 2144 uAfaUfgGfCfUfgCfagaUfcAfuAfuUfusAfsa D2035 S2035 1053 AfaAfuAfuGfaUfCfUfgCfaGfCfCfaUfuAfL96 AS2035 2145 uAfaUfggcUfgCfagaUfcAfuAfuUfusAfsa D2036 S2036 1054 AfaUfaUfgAfuCfUfGfcAfgccAfuUfaAfL96 AS2036 2146 uUfaAfuGfGfCfuGfcagAfuCfaUfaUfusUfsa D2037 S2037 1055 AfaUfaUfgAfuCfUfGfcAfgCfCfAfuUfaAfL96 AS2037 2147 uUfaAfuggCfuGfcagAfuCfaUfaUfusUfsa D2038 S2038 1056 AfuAfuGfaUfcUfGfCfaGfccaUfuAfaAfL96 AS2038 2148 uUfuAfaUfGfGfcUfgcaGfaUfcAfuAfusUfsu D2039 S2039 1057 AfuAfuGfaUfcUfGfCfaGfcCfAfUfuAfaAfL96 AS2039 2149 uUfuAfaugGfcUfgcaGfaUfcAfuAfusUfsu D2040 S2040 1058 UfaUfgAfuCfuGfCfAfgCfcauUfaAfaAfL96 AS2040 2150 uUfuUfaAfUfGfgCfugcAfgAfuCfaUfasUfsu D2041 S2041 1059 UfaUfgAfuCfuGfCfAfgCfcAfUfUfaAfaAfL96 AS2041 2151 uUfuUfaauGfgCfugcAfgAfuCfaUfasUfsu D2042 S2042 1060 AfuGfaUfcUfgCfAfGfcCfauuAfaAfaAfL96 AS2042 2152 uUfuUfuAfAfUfgGfcugCfaGfaUfcAfusAfsu D2043 S2043 1061 AfuGfaUfcUfgCfAfGfcCfaUfUfAfaAfaAfL96 AS2043 2153 uUfuUfuaaUfgGfcugCfaGfaUfcAfusAfsu D2044 S2044 1062 UfgAfuCfuGfcAfGfCfcAfuuaAfaAfaGfL96 AS2044 2154 cUfuUfuUfAfAfuGfgcuGfcAfgAfuCfasUfsa D2045 S2045 1063 UfgAfuCfuGfcAfGfCfcAfuUfAfAfaAfaGfL96 AS2045 2155 cUfuUfuuaAfuGfgcuGfcAfgAfuCfasUfsa D2046 S2046 1064 GfaUfcUfgCfaGfCfCfaUfuaaAfaAfgAfL96 AS2046 2156 uCfuUfuUfUfAfaUfggcUfgCfaGfaUfcsAfsu D2047 S2047 1065 GfaUfcUfgCfaGfCfCfaUfuAfAfAfaAfgAfL96 AS2047 2157 uCfuUfuuuAfaUfggcUfgCfaGfaUfcsAfsu D2048 S2048 1066 AfuCfuGfcAfgCfCfAfuUfaaaAfaGfaCfL96 AS2048 2158 gUfcUfuUfUfUfaAfuggCfuGfcAfgAfusCfsa D2049 S2049 1067 AfuCfuGfcAfgCfCfAfuUfaAfAfAfaGfaCfL96 AS2049 2159 gUfcUfuuuUfaAfuggCfuGfcAfgAfus Cfsa D2050 S2050 1068 UfcUfgCfaGfcCfAfUfuAfaaaAfgAfcAfL96 AS2050 2160 uGfuCfuUfUfUfuAfaugGfcUfgCfaGfasUfsc D2051 S2051 1069 UfcUfgCfaGfcCfAfUfuAfaAfAfAfgAfcAfL96 AS2051 2161 uGfuCfuuuUfuAfaugGfcUfgCfaGfasUfsc D2052 S2052 1070 CfuGfcAfgCfcAfUfUfaAfaaaGfaCfaCfL96 AS2052 2162 gUfgUfcUfUfUfuUfaauGfgCfuGfcAfgsAfsu D2053 S2053 1071 CfuGfcAfgCfcAfUfUfaAfaAfAfGfaCfaCfL96 AS2053 2163 gUfgUfcuuUfuUfaauGfgCfuGfcAfgsAfsu D2054 S2054 1072 UfgCfaGfcCfaUfUfAfaAfaagAfcAfcAfL96 AS2054 2164 uGfuGfuCfUfUfuUfuaaUfgGfcUfgCfasGfsa D2055 S2055 1073 UfgCfaGfcCfaUfUfAfaAfaAfGfAfcAfcAfL96 AS2055 2165 uGfuGfucuUfuUfuaaUfgGfcUfgCfasGfsa D2056 S2056 1074 GfcAfgCfcAfuUfAfAfaAfagaCfaCfaUfL96 AS2056 2166 aUfgUfgUfCfUfuUfuuaAfuGfgCfuGfcsAfsg D2057 S2057 1075 GfcAfgCfcAfuUfAfAfaAfaGfAfCfaCfaUfL96 AS2057 2167 aUfgUfgucUfuUfuuaAfuGfgCfuGfcsAfsg D2058 S2058 1076 CfaGfcCfaUfuAfAfAfaAfgacAfcAfuUfL96 AS2058 2168 aAfuGfuGfUfCfuUfuuuAfaUfgGfcUfgsCfsa D2059 S2059 1077 CfaGfcCfaUfuAfAfAfaAfgAfCfAfcAfuUfL96 AS2059 2169 aAfuGfuguCfuUfuuuAfaUfgGfcUfgsCfsa D2060 S2060 1078 AfgCfcAfuUfaAfAfAfaGfacaCfaUfuCfL96 AS2060 2170 gAfaUfgUfGfUfcUfuuuUfaAfuGfgCfusGfsc D2061 S2061 1079 AfgCfcAfuUfaAfAfAfaGfaCfAfCfaUfuCfL96 AS2061 2171 gAfaUfgugUfcUfuuuUfaAfuGfgCfusGfsc D2062 S2062 1080 GfcCfaUfuAfaAfAfAfgAfcacAfuUfcUfL96 AS2062 2172 aGfaAfuGfUfGfuCfuuuUfuAfaUfgGfcsUfsg D2063 S2063 1081 GfcCfaUfuAfaAfAfAfgAfcAfCfAfuUfcUfL96 AS2063 2173 aGfaAfuguGfuCfuuuUfuAfaUfgGfcsUfsg D2064 S2064 1082 CfcAfuUfaAfaAfAfGfaCfacaUfuCfuGfL96 AS2064 2174 cAfgAfaUfGfUfgUfcuuUfuUfaAfuGfgsCfsu D2065 S2065 1083 CfcAfuUfaAfaAfAfGfaCfaCfAfUfuCfuGfL96 AS2065 2175 cAfgAfaugUfgUfcuuUfuUfaAfuGfgsCfsu D2066 S2066 1084 CfaUfuAfaAfaAfGfAfcAfcauUfcUfgUfL96 AS2066 2176 aCfaGfaAfUfGfuGfucuUfuUfuAfaUfgsGfsc D2067 S2067 1085 CfaUfuAfaAfaAfGfAfcAfcAfUfUfcUfgUfL96 AS2067 2177 aCfaGfaauGfuGfucuUfuUfuAfaUfgsGfsc D2068 S2068 1086 AfuUfaAfaAfaGfAfCfaCfauuCfuGfuAfL96 AS2068 2178 uAfcAfgAfAfUfgUfgucUfuUfuUfaAfusGfsg D2069 S2069 1087 AfuUfaAfaAfaGfAfCfaCfaUfUfCfuGfuAfL96 AS2069 2179 uAfcAfgaaUfgUfgucUfuUfuUfaAfusGfsg D2070 S2070 1088 UfuAfaAfaAfgAfCfAfcAfuucUfgUfaAfL96 AS2070 2180 uUfaCfaGfAfAfuGfuguCfuUfuUfuAfasUfsg D2071 S2071 1089 UfuAfaAfaAfgAfCfAfcAfuUfCfUfgUfaAfL96 AS2071 2181 uUfaCfagaAfuGfuguCfuUfuUfuAfasUfsg D2072 S2072 1090 UfaAfaAfaGfaCfAfCfaUfucuGfuAfaAfL96 AS2072 2182 uUfuAfcAfGfAfaUfgugUfcUfuUfuUfasAfsu D2073 S2073 1091 UfaAfaAfaGfaCfAfCfaUfuCfUfGfuAfaAfL96 AS2073 2183 uUfuAfcagAfaUfgugUfcUfuUfuUfasAfsu D2074 S2074 1092 AfaAfaAfgAfcAfCfAfuUfcugUfaAfaAfL96 AS2074 2184 uUfuUfaCfAfGfaAfuguGfuCfuUfuUfusAfsa D2075 S2075 1093 AfaAfaAfgAfcAfCfAfuUfcUfGfUfaAfaAfL96 AS2075 2185 uUfuUfacaGfaAfuguGfuCfuUfuUfusAfsa D2076 S2076 1094 AfaAfaGfaCfaCfAfUfuCfuguAfaAfaAfL96 AS2076 2186 uUfuUfuAfCfAfgAfaugUfgUfcUfuUfusUfsa D2077 S2077 1095 AfaAfaGfaCfaCfAfUfuCfuGfUfAfaAfaAfL96 AS2077 2187 uUfuUfuacAfgAfaugUfgUfcUfuUfusUfsa D2078 S2078 1096 AfaAfgAfcAfcAfUfUfcUfguaAfaAfaAfL96 AS2078 2188 uUfuUfuUfAfCfaGfaauGfuGfuCfuUfusUfsu D2079 S2079 1097 AfaAfgAfcAfcAfUfUfcUfgUfAfAfaAfaAfL96 AS2079 2189 uUfuUfuuaCfaGfaauGfuGfuCfuUfusUfsu D2080 S2080 1098 AfaGfaCfaCfaUfUfCfuGfuaaAfaAfaAfL96 AS2080 2190 uUfuUfuUfUfAfcAfgaaUfgUfgUfcUfusUfsu D2081 S2081 1099 AfaGfaCfaCfaUfUfCfuGfuAfAfAfaAfaAfL96 AS2081 2191 uUfuUfuuuAfcAfgaaUfgUfgUfcUfusUfsu D2082 S2082 1100 AfgAfcAfcAfuUfCfUfgUfaaaAfaAfaAfL96 AS2082 2192 uUfuUfuUfUfUfaCfagaAfuGfuGfuCfusUfsu D2083 S2083 1101 AfgAfcAfcAfuUfCfUfgUfaAfAfAfaAfaAfL96 AS2083 2193 uUfuUfuuuUfaCfagaAfuGfuGfuCfusUfsu D2084 S2084 1102 GfaCfaCfaUfuCfUfGfuAfaaaAfaAfaAfL96 AS2084 2194
uUfuUfuUfUfUfuAfcagAfaUfgUfgUfcsUfsu D2085 S2085 1103 GfaCfaCfaUfuCfUfGfuAfaAfAfAfaAfaAfL96 AS2085 2195 uUfuUfuuuUfuAfcagAfaUfgUfgUfcsUfsu D2086 S2086 1104 AfcAfcAfuUfcUfGfUfaAfaaaAfaAfaAfL96 AS2086 2196 uUfuUfuUfUfUfuUfacaGfaAfuGfuGfusCfsu D2087 S2087 1105 AfcAfcAfuUfcUfGfUfaAfaAfAfAfaAfaAfL96 AS2087 2197 uUfuUfuuuUfuUfacaGfaAfuGfuGfusCfsu D2088 S2088 1106 CfaCfaUfuCfuGfUfAfaAfaaaAfaAfaAfL96 AS2088 2198 uUfuUfuUfUfUfuUfuacAfgAfaUfgUfgsUfsc D2089 S2089 1107 CfaCfaUfuCfuGfUfAfaAfaAfAfAfaAfaAfL96 AS2089 2199 uUfuUfuuuUfuUfuacAfgAfaUfgUfgsUfsc D2090 S2090 1108 AfcAfuUfcUfgUfAfAfaAfaaaAfaAfaAfL96 AS2090 2200 uUfuUfuUfUfUfuUfuuaCfaGfaAfuGfusGfsu D2091 S2091 1109 AfcAfuUfcUfgUfAfAfaAfaAfAfAfaAfaAfL96 AS2091 2201 uUfuUfuuuUfuUfuuaCfaGfaAfuGfusGfsu Lowercase nucleotides (a, u, g, c) are 2'-O-methyl nucleotides; Nf (e.g., Af) is a 2'-fluoro nucleotide; s is aphosphothiorate linkage; L96 indicates a GalNAc.sub.3 ligand.
Example 4: In Vitro Screening of RNAi Agents
Cell Culture and Transfections
[0405] Human Hep3B cells or rat H.II.4.E cells (ATCC, Manassas, Va.) were grown to near confluence at 37.degree. C. in an atmosphere of 5% CO2 in RPMI (ATCC) supplemented with 10% FBS, streptomycin, and glutamine (ATCC) before being released from the plate by trypsinization. Transfection was carried out by adding 14.8 .mu.l of Opti-MEM plus 0.2 .mu.l of Lipofectamine RNAiMax per well (Invitrogen, Carlsbad Calif. cat #13778-150) to 5 .mu.l of siRNA duplexes per well into a 96-well plate and incubated at room temperature for 15 minutes. 80 .mu.l of complete growth media without antibiotic containing .about.2.times.104 Hep3B cells were then added to the siRNA mixture. Cells were incubated for either 24 or 120 hours prior to RNA purification. Single dose experiments were performed at 10 nM and 0.1 nM final duplex concentration and dose response experiments were done using 8, 4 fold serial dilutions with a maximum dose of 10 nM final duplex concentration.
Total RNA Isolation Using DYNABEADS mRNA Isolation Kit (Invitrogen, Part #: 610-12)
[0406] Cells were harvested and lysed in 150 .mu.l of Lysis/Binding Buffer then mixed for 5 minutes at 850 rpm using an Eppendorf Thermomixer (the mixing speed was the same throughout the process). Ten microliters of magnetic beads and 80 .mu.l Lysis/Binding Buffer mixture were added to a round bottom plate and mixed for 1 minute. Magnetic beads were captured using magnetic stand and the supernatant was removed without disturbing the beads. After removing the supernatant, the lysed cells were added to the remaining beads and mixed for 5 minutes. After removing the supernatant, magnetic beads were washed 2 times with 150 .mu.l Wash Buffer A and mixed for 1 minute. Beads were capture again and supernatant removed. Beads were then washed with 150 .mu.l Wash Buffer B, captured and supernatant was removed. Beads were next washed with 150 .mu.l Elution Buffer, captured and supernatant removed. Beads were allowed to dry for 2 minutes. After drying, 50 .mu.l of Elution Buffer was added and mixed for 5 minutes at 70.degree. C. Beads were captured on magnet for 5 minutes. 40 .mu.l of supernatant was removed and added to another 96 well plate.
cDNA Synthesis Using ABI High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, Calif., Cat #4368813)
[0407] A master mix of 1 .mu.l 10.times. Buffer, 0.4 .mu.l 25.times.dNTPs, 1 .mu.l Random primers, 0.5 .mu.l Reverse Transcriptase, 0.5 .mu.l RNase inhibitor and 1.6 .mu.l of H.sub.2O per reaction were added into 5 .mu.l total RNA. cDNA was generated using a Bio-Rad C-1000 or S-1000 thermal cycler (Hercules, Calif.) through the following steps: 25.degree. C. 10 min, 37.degree. C. 120 min, 85.degree. C. 5 sec, 4.degree. C. hold.
Real Time PCR
[0408] 2 .mu.l of cDNA were added to a master mix containing 0.5l GAPDH TaqMan Probe (Applied Biosystems Cat #4326317E (human) Cat #4308313 (rodent)), 0.5p TTR TaqMan probe (Applied Biosystems cat # HS00174914 ml (human) cat # Rn00562124_ml (rat)) and 5 .mu.l Lightcycler 480 probe master mix (Roche Cat #04887301001) per well in a 384 well plate (Roche cat #04887301001). Real time PCR was done in a Roche LC 480 Real Time PCR machine (Roche). Each duplex was tested in at least two independent transfections and each transfection was assayed in duplicate, unless otherwise noted.
[0409] To calculate relative fold change, real time data were analyzed using the .DELTA..DELTA.Ct method and normalized to assays performed with cells transfected with 10 nM AD-1955, or mock transfected cells. IC.sub.50s were calculated using a 4 parameter fit model using XLFit and normalized to cells transfected with AD-1955 (sense sequence: cuuAcGcuGAGuAcuucGAdTsdT (SEQ ID NO: 2202); antisense sequence: UCGAAGuCUcAGCGuAAGdTsdT (SEQ ID NO: 2203)) or naive cells over the same dose range, or to its own lowest dose. IC.sub.50s were calculated for each individual transfection as well as in combination, where a single IC.sub.50 was fit to the data from both transfections.
[0410] The results of gene silencing of the exemplary siRNA duplex with various motif modifications of the invention are shown in Table 1 above.
Example 5: In Vitro Silencing Activity of Chemically Modified RNAi Agents that Target TTR
[0411] The following experiments demonstrated the beneficial effects of chemical modifications, including the introduction of triplet repeat motifs, together with a GalNAc.sub.3 ligand, on the silencing activity of RNAi agents that target TTR. The sequences of the agents investigated are provided in Table 2 below. The regions of complementarity to the TTR mRNA are as follows: the region of complementarity of RNAi agents AD-45165, AD-51546 and AD-51547 is GGATGGGATTTCATGTAACCAAGA (SEQ ID NO: 2204) and the region or complemetarity of RNAi agents AD-45163, AD-51544, and AD-51545 is TTCATGTAACCAAGAGTATTCCAT (SEQ ID NO: 2205).
Protocol for Assessment of IC.sub.50 in Hep3B Cells
[0412] The IC.sub.50 for each modified siRNA was determined in Hep3B cells (a human hepatoma cell line) by standard reverse transfection using Lipofectamine RNAiMAX. In brief, reverse transfection was carried out by adding 5 .mu.L of Opti-MEM to 5 .mu.L of siRNA duplex per well into a 96-well plate along with 10 .mu.L of Opti-MEM plus 0.5 .mu.L of Lipofectamine RNAiMax per well (Invitrogen, Carlsbad Calif. cat #13778-150) and incubating at room temperature for 15-20 minutes. Following incubation, 100 .mu.L of complete growth media without antibiotic containing 12,000-15,000 Hep3B cells was then added to each well. Cells were incubated for 24 hours at 37.degree. C. in an atmosphere of 5% CO.sub.2 prior to lysis and analysis of TTR and GAPDH mRNA by bDNA (Quantigene). Seven different siRNA concentrations ranging from 10 nM to 0.6 .mu.M were assessed for IC.sub.50 determination and TTR/GAPDH for siRNA transfected cells was normalized to cells transfected with 10 nM Luc siRNA. The results are shown in Table 2.
Protocol for Assessment of Free-Uptake IC.sub.50
[0413] Free uptake silencing in primary cynomolgus hepatocytes was assessed following incubation with TTR siRNA for either 4 hours or 24 hours. Silencing was measured at 24 hours from the initial exposure. In brief, 96-well culture plates were coated with 0.05%-0.1% collagen (Sigma C3867-1VL) at room temperature, 24 hours prior to the start of the experiment. On the day of assay, siRNAs were diluted in pre-warmed Plating Media consisting of DMEM supplemented with GIBCO's Maintenance Media Kit (Serum-Free, Life Technologies CM4000), and added to the collagen-coated 96-well culture plates. Cryopreserved primary cynomolgus hepatocytes were rapidly thawed in a 37.degree. C. water bath, and immediately diluted in Plating Media to a concentration of 360,000 cells/mL. A volume of cell suspension was gently pipetted on top of the pre-plated siRNAs such that the final cell count was 18,000 cells/well. The plate was lightly swirled to mix and spread cells evenly across the wells and placed in a 37.degree. C., 5% CO.sub.2 incubator for 24 hours prior to lysis and analysis of TTR and GAPDH mRNA by bDNA (Quantigene, Affymetrix). In the case of the 4 h incubation with siRNA, the media was decanted after 4 hours of exposure to the cells, and replaced with fresh Plating Media for the remaining 20 hours of incubation. Downstream analysis for TTR and GAPDH mRNA was the same as described above. For a typical dose response curve, siRNAs were titrated from 1 uM to 0.24 nM by 4 fold serial dilution.
TABLE-US-00006 TABLE 2 In vitro Activity Summary for Alternating TTR-GalNAc and Variants with Triplet Motifs Free-Uptake Hep3B Duplex IC50 (.mu.M) IC50 ID S (5'-3') AS (5'-3'') 4 h 24 h (nM) AD-45163 AfuGfuAfaCfcAfaGfaGfu aUfgGfaAfuAfcUfcUfuGfg 0.04101 0.00820 0.0115 AfuUfcCfaUfL96 UfuAfcAfusGfsa (SEQ ID NO: 2206) (SEQ ID NO: 2212) AD-51544 AfuGfuAfaCfcAfAfGfaGf aUfgGfAfAfuAfcUfcuuGfg 0.00346 0.00374 0.0014 uAfuucCfaUfL96 UfuAfcAfusGfsa (SEQ ID NO: 2207) (SEQ ID NO: 2213) AD-51545 AfuGfuAfAfCfcAfAfGfaG aUfgGfaAfuAfcUfcuuGfgu 0.00395 0.00389 0.0018 fuAfuUfcCfaUfL96 uAfcAfusGfsa (SEQ ID NO: 2208) (SEQ ID NO: 2214) AD-45165 UfgGfgAfuUfuCfaUfgUfa uCfuUfgGfuUfaCfaUfgAfa 0.02407 0.00869 0.0112 AfcCfaAfgAfL96 AfuCfcCfasUfsc (SEQ ID NO: 2209) (SEQ ID NO: 2215) AD-51546 UfgGfGfAfuUfuCfAfUfgU uCfuugGfuUfaCfaugAfaAf 0.00317 0.00263 0.0017 faAfcCfAfAfgAfL96 uccCfasUfsc (SEQ ID NO: 2210) (SEQ ID NO: 2216) AD-51547 UfgGfgAfuUfuCfAfUfgUf uCfuUfgGfUfUfaCfaugAfa 0.00460 0.00374 0.0028 aacCfaAfgAfL96 AfuCfcCfasUfsc (SEQ ID NO: 2211) (SEQ ID NO: 2217) Lowercase nucleotides (a, u, g, c) indicate 2'-O-methyl nucleotides; Nf (e.g., Af) indicates a 2'-fluoro nucleotide; s indicates a phosphothiorate linkage; L96 indicates a GalNAc.sub.3 ligand; bold nucleotides indicate changes relative to the corresponding parent agent. Each bold nucleotide is at the center of a triplet motif.
[0414] The results are provided in Table 2 and demonstrate that modified RNAi agents that target TTR provide enhanced silencing activity.
Results: Improved Activity of Modified RNAi Agents
[0415] Parent RNAi agents with alternating chemical modifications and a GalNAc3 ligand provided an IC.sub.50 in Hep3B cells of about 0.01 nM. As shown in FIGS. 4-5 and in Table 2, agents modified relative to the parent agents, for example, by the addition of one or more repeating triplets of 2'-fluoro and 2'-O-methyl modifications, showed unexpectedly enhanced silencing activity, achieving IC.sub.50 values in Hep3B cells that were 5-8 fold better than the corresponding parent agent.
Results: Free Uptake IC.sub.50s in Hep3B Cells
[0416] As shown in Table 2 and FIGS. 6-7, RNAi agents modified relative to the parent AD-45163 also showed enhanced free uptake silencing. The modified agents showed more than double the silencing activity of the parent after a 24 hour incubation period and nearly 10 times the silencing activity of the parent after a 4 hour incubation period.
[0417] As shown in Table 2 and FIGS. 8-9, RNAi agents modified relative to the parent AD-45165 also showed enhanced free uptake silencing. The modified agents showed 2-3 times the silencing activity of the parent after a 24 hour incubation period and 5-8 times the silencing activity of the parent after a 4 hour incubation period.
[0418] Taken collectively, these results demonstrate that the modified RNAi agents presented herein, e.g., AD-51544, AD-51545, AD-51546, and AD-51547, all showed unexpectedly good inhibition of TTR mRNA in in vitro silencing experiments.
Example 6: TTR mRNA Silencing and TTR Protein Suppression in Transgenic Mice
[0419] To assess the efficacy of the RNAi agents AD-45163, AD-51544, AD-51545, AD45165, AD-51546, and AD-51547, these agents were administered to transgenic mice that express human transthyretin with the V30M mutation (see Santos, S D., Fernaandes, R., and Saraiva, M J. (2010) Neurobiology of Aging, 31, 280-289). The V30M mutation is known to cause familial amyloid polyneuropathy type I in humans. See, e.g., Lobato, L. (2003) JNephrol., 16(3):438-42.
[0420] The RNAi agents (in PBS buffer) or PBS control were administered to mice (2 male and 2 female) of 18-24 months of age in a single subcutaneous dose of 5 mg/kg or 1 mg/kg. After approximately 48 hours, mice were anesthetized with 200 .mu.l of ketamine, and then exsanguinated by severing the right caudal artery. Whole blood was isolated and plasma was isolated and stored at -80.degree. C. until assaying. Liver tissue was collected, flash-frozen and stored at -80.degree. C. until processing.
[0421] Efficacy of treatment was evaluated by (i) measurement of TTR mRNA in liver at 48 hours post-dose, and (ii) measurement of TTR protein in plasma at pre-bleed and at 48 hours post-dose. TTR liver mRNA levels were assayed utilizing the Branched DNA assays--QuantiGene 2.0 (Panomics cat #: QS0011). Briefly, mouse liver samples were ground and tissue lysates were prepared. Liver lysis mixture (a mixture of 1 volume of lysis mixture, 2 volume of nuclease-free water and 10 ul of Proteinase-K/ml for a final concentration of 20 mg/ml) was incubated at 65.degree. C. for 35 minutes. 20 .mu.l of Working Probe Set (TTR probe for gene target and GAPDH for endogenous control) and 80 ul of tissue-lysate were then added into the Capture Plate. Capture Plates were incubated at 55.degree. C. 1.degree. C. (aprx. 16-20 hrs). The next day, the Capture Plates were washed 3 times with 1.times. Wash Buffer (nuclease-free water, Buffer Component 1 and Wash Buffer Component 2), then dried by centrifuging for 1 minute at 240 g. 100 .mu.l of pre-Amplifier Working Reagent was added into the Capture Plate, which was sealed with aluminum foil and incubated for 1 hour at 55.degree. C. 1.degree. C. Following 1 hour incubation, the wash step was repeated, then 100 .mu.l of Amplifier Working Reagent was added. After 1 hour, the wash and dry steps were repeated, and 100 .mu.l of Label Probe was added. Capture plates were incubated 50.degree. C. 1.degree. C. for 1 hour. The plate was then washed with 1.times. Wash Buffer, dried and 1001 Substrate was added into the Capture Plate. Capture Plates were read using the SpectraMax Luminometer following a 5 to 15 minute incubation. bDNA data were analyzed by subtracting the average background from each triplicate sample, averaging the resultant triplicate GAPDH (control probe) and TTR (experimental probe) values, and then computing the ratio: (experimental probe-background)/(control probe-background).
[0422] Plasma TTR levels were assayed utilizing the commercially available kit "AssayMax Human Prealbumin ELISA Kit" (AssayPro, St. Charles, Mo., Catalog # EP3010-1) according to manufacturer's guidelines. Briefly, mouse plasma was diluted 1:10,000 in 1.times. mix diluents and added to pre-coated plates along with kit standards, and incubated for 2 hours at room temperature followed by 5.times. washes with kit wash buffer. Fifty microliters of biotinylated prealbumin antibody was added to each well and incubated for 1 hr at room temperature, followed by 5.times. washes with wash buffer. Fifty microliters of streptavidin-peroxidase conjugate was added to each well and plates were incubated for 30 minutes at room temperature followed by washing as previously described. The reaction was developed by the addition of 50 .mu.l/well of chromogen substrate and incubation for 10 minutes at room temperature with stopping of reaction by the addition of 50 .mu.l/well of stop solution. Absorbance at 450 nm was read on a Versamax microplate reader (Molecular Devices, Sunnyvale, Calif.) and data were analyzed utilizing the Softmax 4.6 software package (Molecular Devices).
[0423] The results are shown in FIGS. 10-12. FIGS. 10A and 10B show that the RNAi agents modified relative to the parent agents AD-45163 and AD-45165 showed RNA silencing activity that was similar or more potent compared with that of the parent agents. FIG. 11 shows that the agents AD-51544 and AD-51545 showed dose dependent silencing activity and that the silencing activity of these agents at a dose of 5 mg/kg was similar to that of the corresponding parent AD-45163. FIG. 12 shows that the agents AD-51546 and AD-51547 also showed dose-dependent silencing activity. Furthermore, the silencing activity of AD-51546 and AD-51547 at a dose of 5 mg/kg was superior to that of the corresponding parent AD-45165.
Example 7: Serum and Liver Pharmacokinetic Profiles of RNAi Agents that Target TTR in Mice
[0424] To assess the pharmacokinetic profiles of the RNAi agents AD-45163, AD-51544, AD-51545, AD-51546, and AD-51547, these agents, in PBS buffer, were administered to C57BL/6 mice using a single IV bolus or subcutaneous (SC) administration. The plasma concentrations and liver concentrations of the agents were assessed at various timepoints after the administration.
[0425] The plasma pharmacokinetic parameters are presented in Tables 3 and 4 below. The mean resident time (MRT) in plasma was about 0.2 hours after IV dosing and about 1 hour after SC dosing. At a dose of 25 mg/kg, the agents AD-51544, AD-51545, AD-51546, and AD-51547 showed similar plasma pharmacokinetic properties. Each of these agents had more than 75% bioavailability from the subcutaneous space. Their bioavailability was superior to that of the parent agent AD-45163 that was administered at a higher dose of 30 mg/kg. The subcutaneous bioavailability of AD-51544 and AD-51547 was about 100%, whereas that of AD-51545 was 90% and that of and AD-51546 was 76%.
TABLE-US-00007 TABLE 3 Summary of Plasma PK Parameter Estimates After SC Administration of TTR-GalNAc siRNAs in Mice 30 mpk 25 mpk 25 mpk 25 mpk 25 mpk AD- AD- AD- AD- AD- 45163 51544 51545 51546 51547 (h/c (h/c (h/c (h/c (h/c TTR- TTR- TTR- TTR- TTR- Parameter GalNAc) GalNAc) GalNAc) GalNAc) GalNAc) Plasma 0.25 1 0.5 1 0.5 Tmax (h) Plasma 9.6 11.7 10.9 11.7 12.1 Cmax (.mu.g/mL) Plasma 12.4 21.9 19.9 20.9 25.3 AUC (h*.mu.g/mL) F.sub.sc(%) 79 100 90.1 76.0 99.2
TABLE-US-00008 TABLE 4 Plasma siRNA PK Parameters in Mice after an IV Bolus or SC Dose of AD-51544, 51545, 51546 or 51547 at 25 mg/kg Test Article AD-51544 AD-51545 AD-51546 AD-51547 siRNA Dose (mg/kg) 25 25 25 25 Route of Administration IV SC IV SC IV SC IV SC t.sub.max (h) 0.083 1 0.083 0.5 0.083 1 0.083 0.5 C.sub.max (.mu.g/mL) 96.5.sup.a 11.7 108.sup.a 10.9 128.sup.a 10.9 123.sup.a 12.1 AUC.sub.0-last (h .mu.g/mL) 21.6 21.9 22.1 19.9 27.5 20.9 25.5 25.3 MRT.sub.0-last (h) 0.17 1.2 0.16 1.1 0.22 1.4 0.19 1.3 Apparent t.sub.1/2 .beta. (h).sup.b ND ND ND 0.49 ND 1.2 ND 0.56 F.sub.SC (%).sup.c -- 102 -- 90.1 -- 76.0 -- 99.2 .sup.aConcentration at the 1.sup.st sampling time (5 min) after IV dosing .sup.bApparent elimination half-life (t.sub.1/2 .beta.) could not be determined (ND) for all 4 test articles after IV dosing as the terminal phase of the concentration-time profiles was not well defined, as a result, the t.sub.1/2 .beta.-associated PK parameters (eg, AUC.sub.0-.infin., CL and Vss) were not reported. .sup.cSC bioavailability, calculated as percentage ratio of AUC.sub.0-last after SC and IV dosing at 25 mg/kg
[0426] The results also indicated that the RNAi agents AD-45163, AD-51544, AD-51545, AD-51546, and AD-51547 achieved similar or higher concentrations in the liver when administered subcutaneously than when administered by IV bolus. The liver pharmacokinetic parameters are presented in Tables 5 and 6 below. The peak concentration (C.sub.max) and area under the curve (AUC.sub.0-last) in the liver were two to three times higher after subcutaneous administration as compared with IV administration of the same agent at the same dose. Liver exposures were highest for AD-51547 and lowest for AD-51545. The mean resident time (MRT) and elimination half-life were longer for AD-51546 and AD-51547 compared with AD-51544 and AD-51545. Following subcutaneous administration, the approximate MRTs were 40 hours for AD-51546 and 25 hours for AD-51547, whereas the MRTs for AD-51544 and AD-51545 were lower (about 6-9 hours). The elimination half life of AD-51546 and AD-51547 was also higher (41-53 hours) than was the elimination half life of AD-51544 and AD-51545 (6-10 hours).
TABLE-US-00009 TABLE 5 Summary of Liver PK Parameter Estimates After SC Administration of TTR-GalNAc siRNAs in Mice 30 mpk 25 mpk 25 mpk 25 mpk 25 mpk AD- AD- AD- AD- AD- 45163 51544 51545 51546 51547 (h/c (h/c (h/c (h/c (h/c TTR- TTR- TTR- TTR- TTR- Parameter GalNAc) GalNAc) GalNAc) GalNAc) GalNAc) Liver Tmax 8 4 4 2 8 (h) Liver Cmax 313 126 80 117 174 (.mu.g/g) Liver AUC 4519 1092 763 2131 4583 (h*.mu.g/g)
TABLE-US-00010 TABLE 6 Liver siRNA PK Parameters in Mice after an IV Bolus or SC Dose of AD-51544, 51545, 51546 or 51547 at 25 mg/kg Test Article AD-51544 AD-51545 AD-51546 AD-51547 siRNA Dose (mg/kg) 25 25 25 25 Route of Administration IV SC IV SC IV SC IV SC t.sub.max (h) 1 4 1 4 4 2 2 8 C.sub.max (.mu.g/g) 67.9 126 37.0 80.5 35.3 117 73.8 174 AUC.sub.0-last (h .mu.g/g) 632 1092 324 763 984 2131 1429 4583 MRT.sub.0-last (h) 8.7 6.5 5.9 8.5 45.7 40.2 29.4 25.3 Apparent t.sub.1/2.beta. (h) 8.1 8.2 5.7 10.0 51.1 45.3 41.1 52.7
Example 8: In Vitro Stability of RNAi Agents in Monkey Serum
[0427] The serum stability of RNAi agents AD-51544, AD-51545, AD-51546, and AD-51547 was also assessed in monkeys. The results demonstrated that the antisense and sense strands of AD-51544, AD-51545, and AD-51547 showed serum stability over a period of about 24 hours (data not shown).
Example 9: RNAi Agents Produce Lasting Suppression of TTR Protein in Non-Human Primates
[0428] The RNA silencing activity of RNAi agents AD-45163, AD-51544, AD-51545, AD-51546, and AD-51547 was assessed by measuring suppression of TTR protein in serum of cynomologous monkeys following subcutaneous administration of five 5 mg/kg doses (one dose each day for 5 days) or a single 25 mg/kg dose. Pre-dose TTR protein levels in serum were assessed by averaging the levels at 11 days prior to the first dose, 7 days prior to the first dose, and 1 day prior to the first dose. Post-dose serum levels of TTR protein were assessed by determining the level in serum beginning at 1 day after the final dose (i.e., study day 5 in the 5.times.5 mg/kg group and study day 1 in the 1.times.25 mg/kg group) until 49 days after the last dose (i.e., study day 53 in the 5.times.5 mg/kg group and study day 49 in the 1.times.25 mg/kg group). See FIG. 13.
[0429] TTR protein levels were assessed as described in Example 6. The results are shown in FIGS. 14A and 14B and in Tables 7 and 8.
[0430] A maximal suppression of TTR protein of up to about 50% was achieved in the groups that received 25 mg/kg of AD-45163, AD-51544, AD-51546, and AD-51547 (see Table 8). A greater maximal suppression of TTR protein of about 70% was achieved in the groups that received 5.times.5 mg/kg of AD-45163, AD-51544, AD-51546, and AD-51547 (see Table 7). The agent AD-51545 produced a lesser degree of suppression in both administration protocols. Significant suppression of about 20% or more persisted for up to 49 days after the last dose of AD-51546 and AD-51547 in both the 1.times.25 mg/kg and 5.times.5 mg/kg protocols. Generally, better suppression was achieved in the 5.times.5 mg/kg protocol than in the 1.times.25 mg/kg protocol.
TABLE-US-00011 TABLE 7 Fraction Serum Transthyretin Relative to Pre-dose in Cynomolgus Monkeys (5 mg/kg daily for 5 days) D-11 D-7 D-1 D5 D7 D9 D11 D14 D18 D22 D26 D32 D39 D46 D53 AD-45163 0.98 0.99 1.03 0.71 0.52 0.40 0.34 0.27 0.31 0.39 0.48 0.64 0.68 0.81 0.88 AD-51544 1.02 0.99 0.99 0.60 0.47 0.37 0.35 0.39 0.48 0.58 0.66 0.74 0.83 0.91 0.92 AD-51545 1.03 0.97 1.00 0.73 0.65 0.63 0.69 0.68 0.78 0.87 0.97 1.00 1.03 1.06 1.09 AD-51546 1.01 0.97 1.02 0.59 0.42 0.35 0.30 0.32 0.43 0.58 0.66 0.77 0.92 0.93 0.97 AD-51547 0.99 0.99 1.02 0.74 0.54 0.41 0.34 0.34 0.39 0.49 0.51 0.53 0.65 0.70 0.77
TABLE-US-00012 TABLE 8 Fraction Serum Transthyretin Relative to Pre-dose in Cynomolgus Monkeys (25 mg/kg) D-11 D-7 D-1 D1 D3 D5 D7 D10 D14 D18 D22 D28 D35 D42 D49 AD-45163 1.04 1.01 0.95 0.99 0.84 0.67 0.57 0.44 0.45 0.51 0.58 0.66 0.72 0.78 0.85 AD-51544 1.01 1.04 0.95 0.92 0.69 0.57 0.49 0.48 0.56 0.65 0.69 0.77 0.83 0.87 0.94 AD-51545 0.98 1.02 0.99 0.87 0.77 0.69 0.71 0.72 0.84 0.90 0.92 0.99 1.00 1.00 1.00 AD-51546 1.04 1.03 0.93 0.89 0.71 0.62 0.53 0.50 0.55 0.70 0.70 0.69 0.72 0.79 0.84 AD-51547 0.96 1.03 1.01 1.19 0.90 0.70 0.54 0.48 0.50 0.50 0.52 0.58 0.62 0.70 0.72
Example 10: Tolerability of RNAi Agents that Target TTR
In Cytokine Evaluation in Whole Blood Assay
[0431] To assess the tolerability of RNAi agents that target TTR (including AD-45163, AD-51544, AD-51545, AD-51546, and AD-51547), each agent was tested in a whole blood assay using blood from three human donors. The agents were either 300 nM DOTAP transfected or 1 .mu.M without transfection reagent (free siRNA). There was less than a two fold change for the following cytokines/chemokines: G-CSF, IFN-.gamma., IL-10, IL-12 (p70), IL1.beta., IL-1ra, IL-6, IL-8, IP-10, MCP-1, MIP-1.alpha., MIP-1.beta., TNF.alpha.. (Results not shown).
In Vivo Evaluation
[0432] To assess in vivo tolerability, RNAi agents were injected subcutaneously in CD1 mice at a dose of 125 mg/kg. No cytokine induction was observed at 2, 4, 6, 24, or 48 hours after subcutaneous injection of AD-45163. No significant cytokine induction was observed at 6 or 24 hours after subcutaneous injection of AD-51544, AD-51545, AD-51546, or AD-51547.
[0433] To further assess in vivo tolerability, multiple RNAi agents (including AD-45163, AD-51544, AD-51545, AD-51546, and AD-51547) were tested by subcutaneous injection of 5 and 25 mg in non-human primates (cynomologous monkeys) with dose volumes between 1-2 ml per site. No erythema or edema was observed at injection sites.
Single SC Dose Rat Tolerability Study
[0434] To assess toxicity, rats were injected with a single subcutaneous dose of 100, 250, 500, or 750 mg/kg of AD-45163 (see Table 9). The following assessments were made: clinical signs of toxicity, body weight, hematology, clinical chemistry and coagulation, organ weights (liver & spleen); gross and microscopic evaluation (kidney, liver, lung, lymph node, spleen, testes, thymus, aorta, heart, intestine (small and large).
TABLE-US-00013 TABLE 9 Single SC Dose Rat Tolerability Study: 100, 250, 500 & 750 mg/kg of AD-45163 in Sprague Dawley Rats Dose Dose No. Male Day of Level Volume Route & Sprague Necropsy Group (mg/kg) (ml/kg) Regimen Dawley Rats PBS 0 10 SC Injection 7/group Day 4 AD-45163 100 Day 1 (5 Tox Parent (2 sites) animals, 2 TK animals)
[0435] The results showed no test article-related clinical signs of toxicity, effects on body weight, organ weights, or clinical chemistry. No histopathology was observed in heart, kidneys, testes, spleen, liver, and thymus. There was a non-adverse, slight test article-related increase in WBC (68%, primarily attributed to increase in NEUT and MONO) at 750 mg/kg. These results indicate that a single-dose of up to 750 mg/kg is well tolerated in rats.
Tolerability of Repeated Subcutaneous Administrations in Rats
[0436] To assess the tolerability of repeated subcutaneous administrations of AD-45163, daily subcutaneous injections of 300 mg/kg were given for 5 days, and a necropsy was performed on day 6. The study design is shown in Table 10.
TABLE-US-00014 TABLE 10 Five Day Repeat Dose Tolerability Study in Rat Dose Level Cone No of Tox Group (kmg/kg (mg/mL) Animals Nx Day 6 PBS 0 0 2M, 2F 2M, 2F AD-45163 300 150 2M, 2F 2M, 2F
[0437] The following outcome variables were assessed: clinical signs, body weights, hematology, clinical chemistry and coagulation, organ weights, gross and microscopic evaluation (liver, spleen, kidney, heart, GI tract and first and last injection site). The results showed no test article-related clinical signs, body weight or organ weight effects, and also no test article-related findings in clinical hematology or chemistry. There was a possible slight prolongation of activated partial thromboplastin time (APTT) on day 6 (20.4 vs. 17.4 sec). Histopathology revealed no test article-related findings in the liver, spleen, heart, and GI tract. In the kidney, minimal to slight hypertrophy of the tubular epithelium (not adverse) was observed. At the last injection site, there was minimal multifocal mononuclear infiltration, not adverse. These results indicate that five daily 300 mg/kg doses of the parent RNAi agent AD-45163 are well tolerated in rats.
Example 11: RNAi Agents Produce Lasting Suppression of TTR Protein in Non-Human Primates
[0438] The RNA silencing activity of RNAi agent AD-51547 was assessed by measuring suppression of TTR protein in the serum of cynomologous monkeys following subcutaneous administration of a "loading phase" of the RNAi agent: five daily doses of either 2.5 mg/kg, 5 mg/kg or 10 mg/kg (one dose each day for 5 days) followed by a "maintenance phase" of the RNAi agent: weekly dosing of either 2.5 mg/kg, 5 mg/kg or 10 mg/kg for 4 weeks. Pre-dose TTR protein levels in serum were assessed by averaging the levels at 11 days prior to the first dose, 7 days prior to the first dose, and 1 day prior to the first dose. Post-dose serum levels of TTR protein were assessed by determining the level in serum relative to pre-dose beginning at 1 day after the loading phase was completed until 40 days after the last dose of the maintenance phase (i.e., study day 70).
[0439] TTR protein levels were assessed as described in Example 6. The results are shown in FIG. 15.
[0440] A maximal suppression of TTR protein of up to about 80% was achieved in all of the groups that received either 2.5 mg/kg, 5 mg/kg or 10 mg/kg of AD-51547. Nadir knockdown was achieved in all of the groups by about day 14, the suppression sustained at nadir knockdown levels with a weekly maintenance dose of either 2.5 mg/kg, 5 mg/kg or 10 mg/kg of AD-51547. The levels of TTR had not returned to baseline more than 40 days after the day of administration of the last maintenance dose for the 5 and 2.5 mg/kg dose levels.
EQUIVALENTS
[0441] Those skilled in the art will recognize, or be able to ascertain using no more than routine experimentation, many equivalents to the specific embodiments and methods described herein. Such equivalents are intended to be encompassed by the scope of the following claims.
Sequence CWU 0 SQTB SEQUENCE LISTING The patent application
contains a lengthy "Sequence Listing" section. A copy of the
"Sequence Listing" is available in electronic form from the USPTO
web site
(https://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200318111A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
0 SQTB SEQUENCE LISTING The patent application contains a lengthy
"Sequence Listing" section. A copy of the "Sequence Listing" is
available in electronic form from the USPTO web site
(https://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200318111A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
References




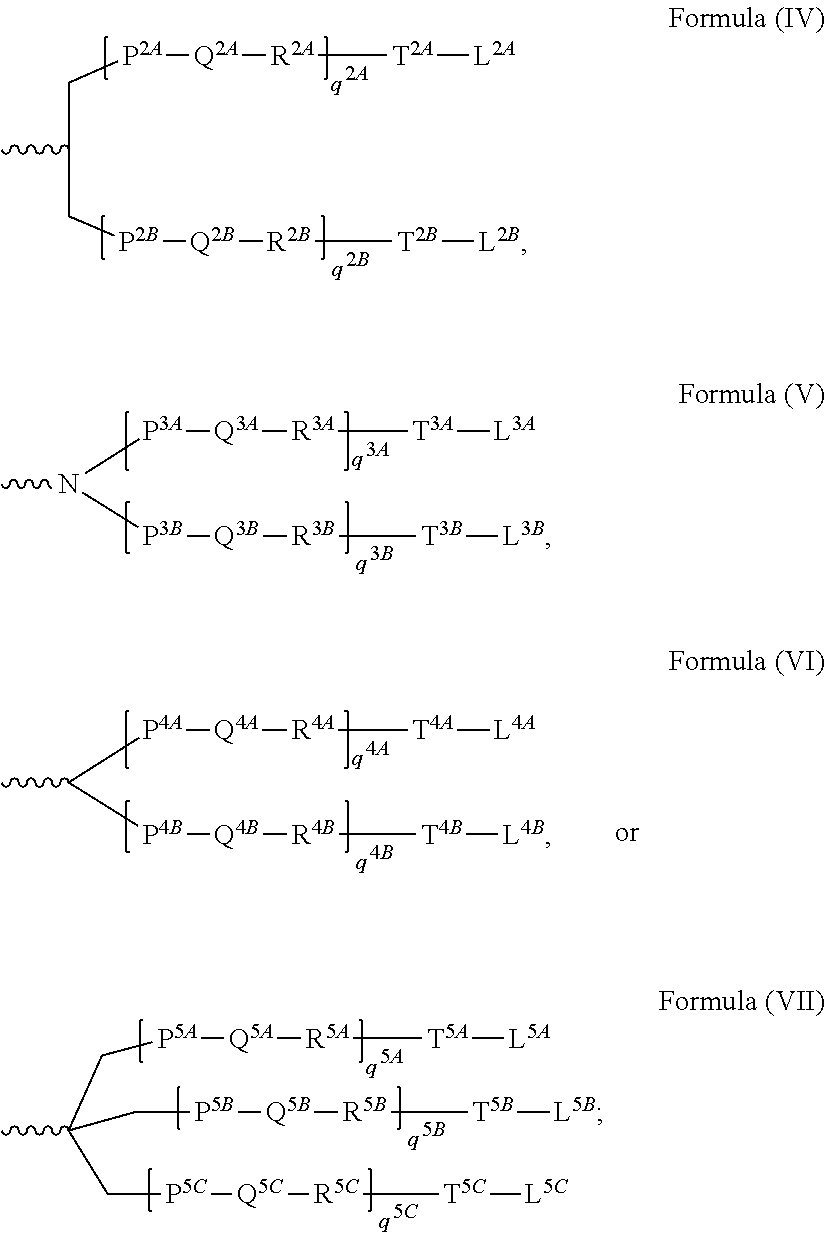



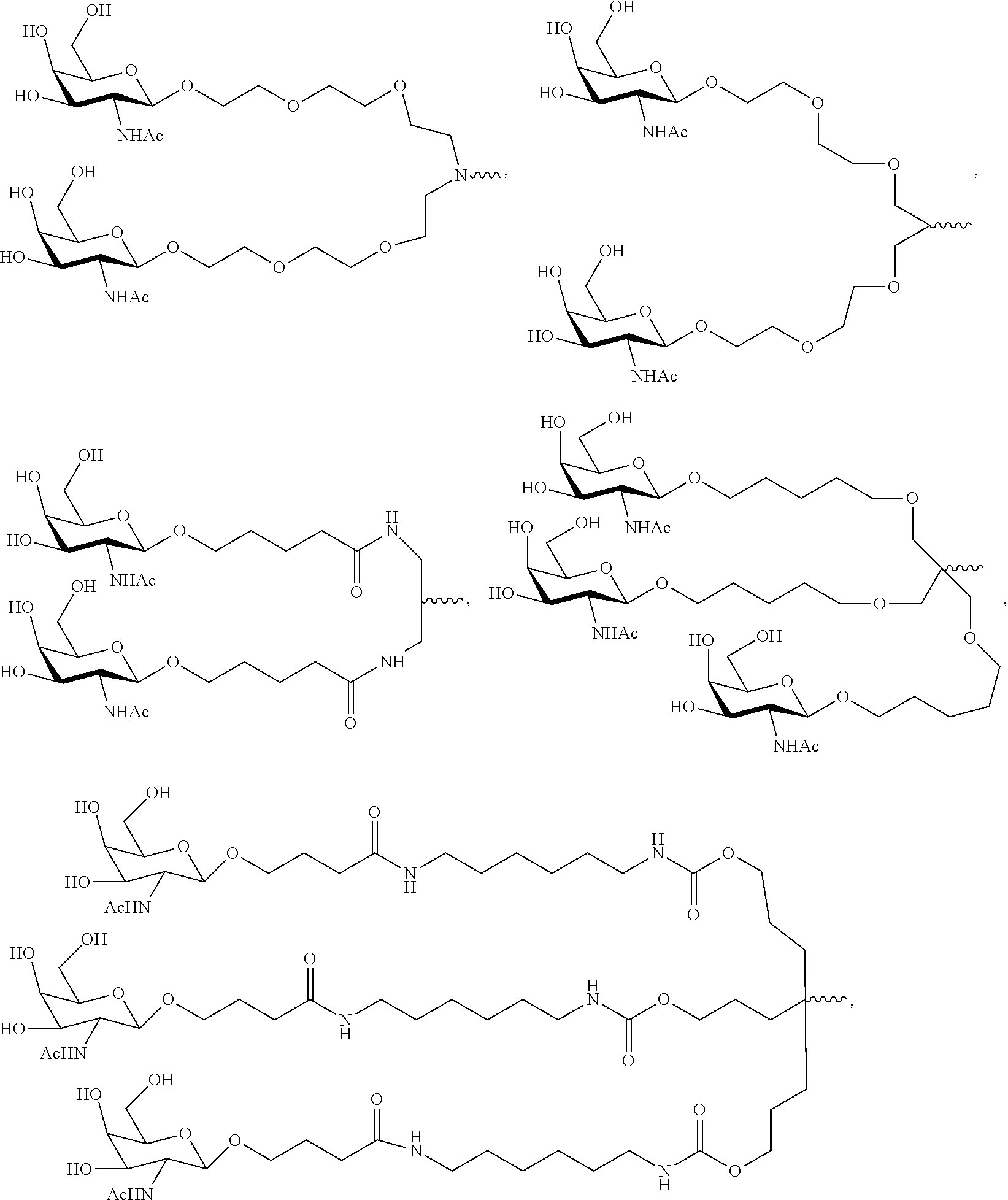



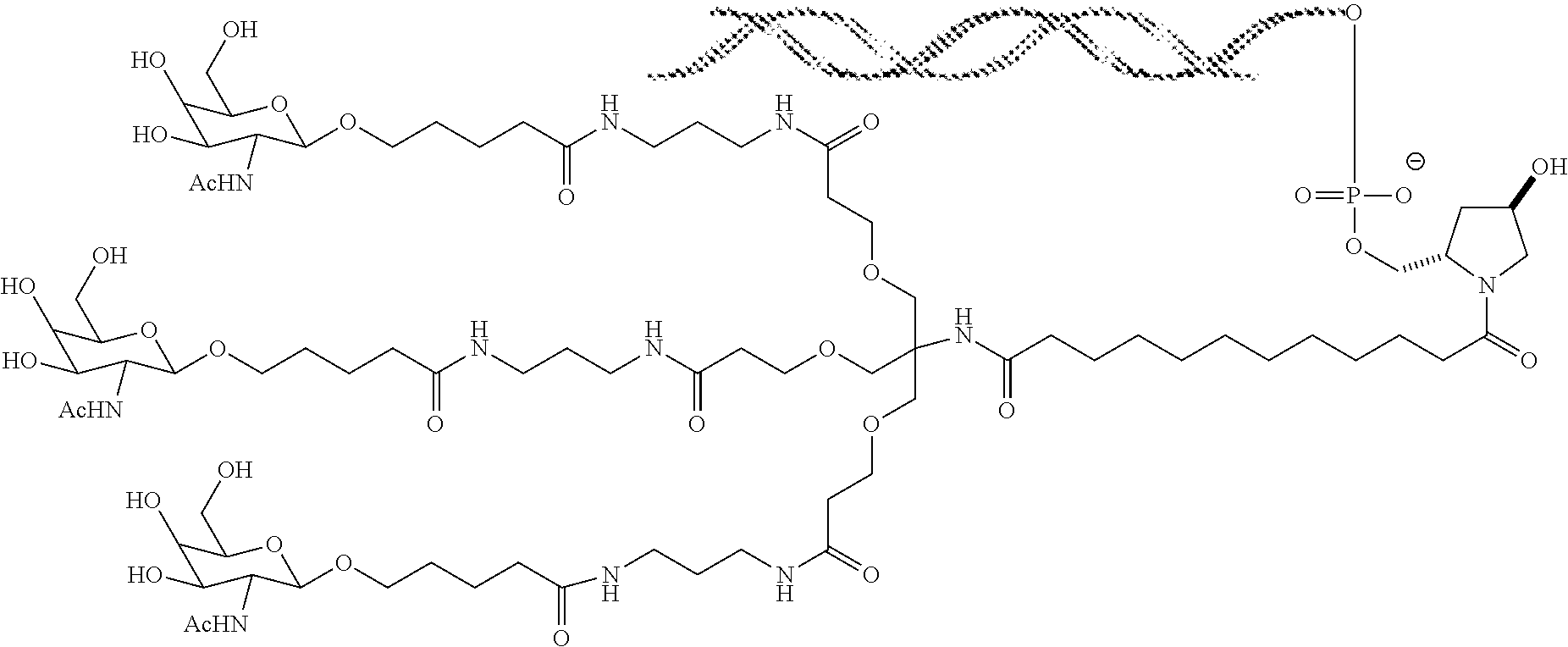

D00001
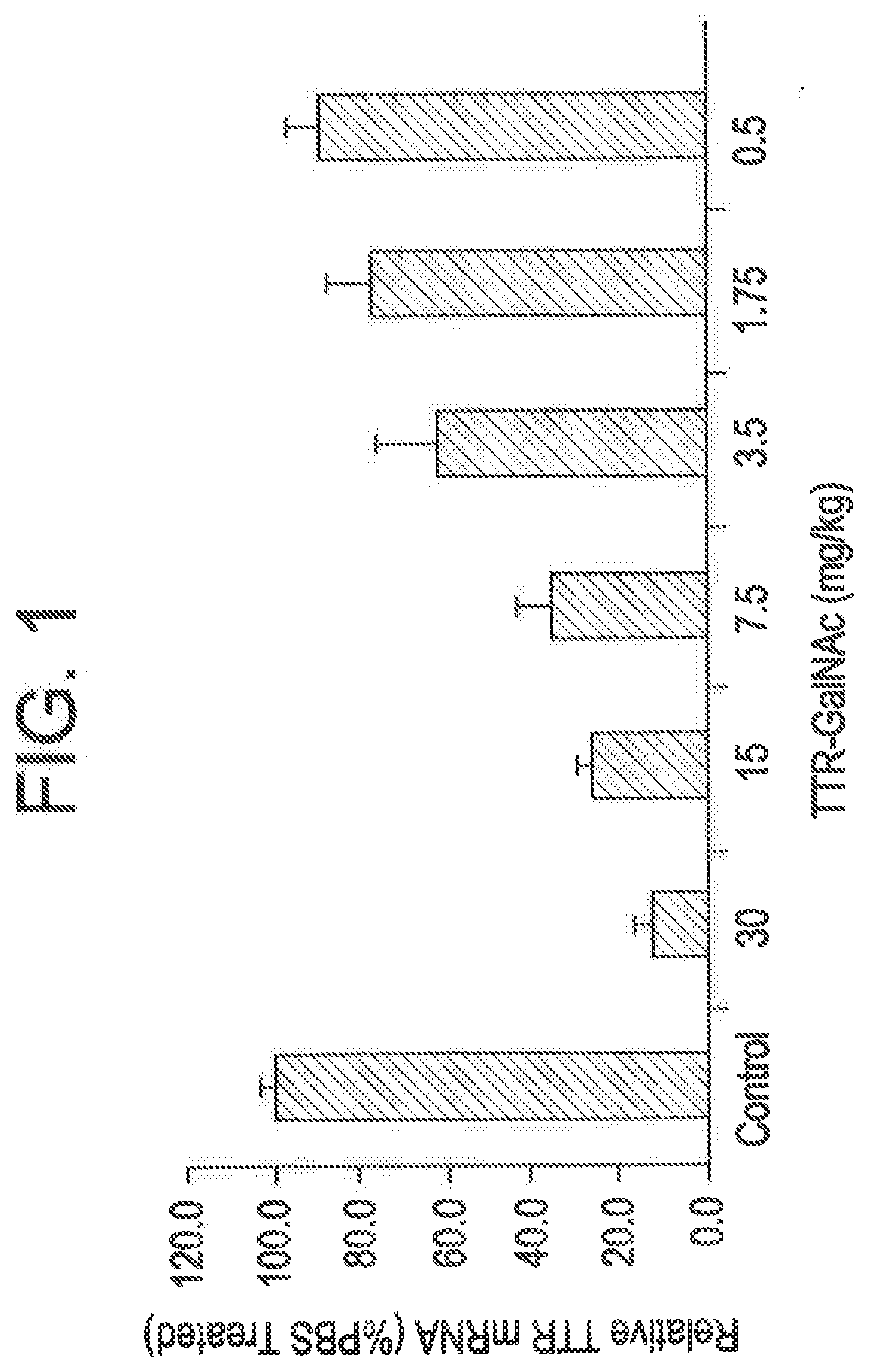
D00002
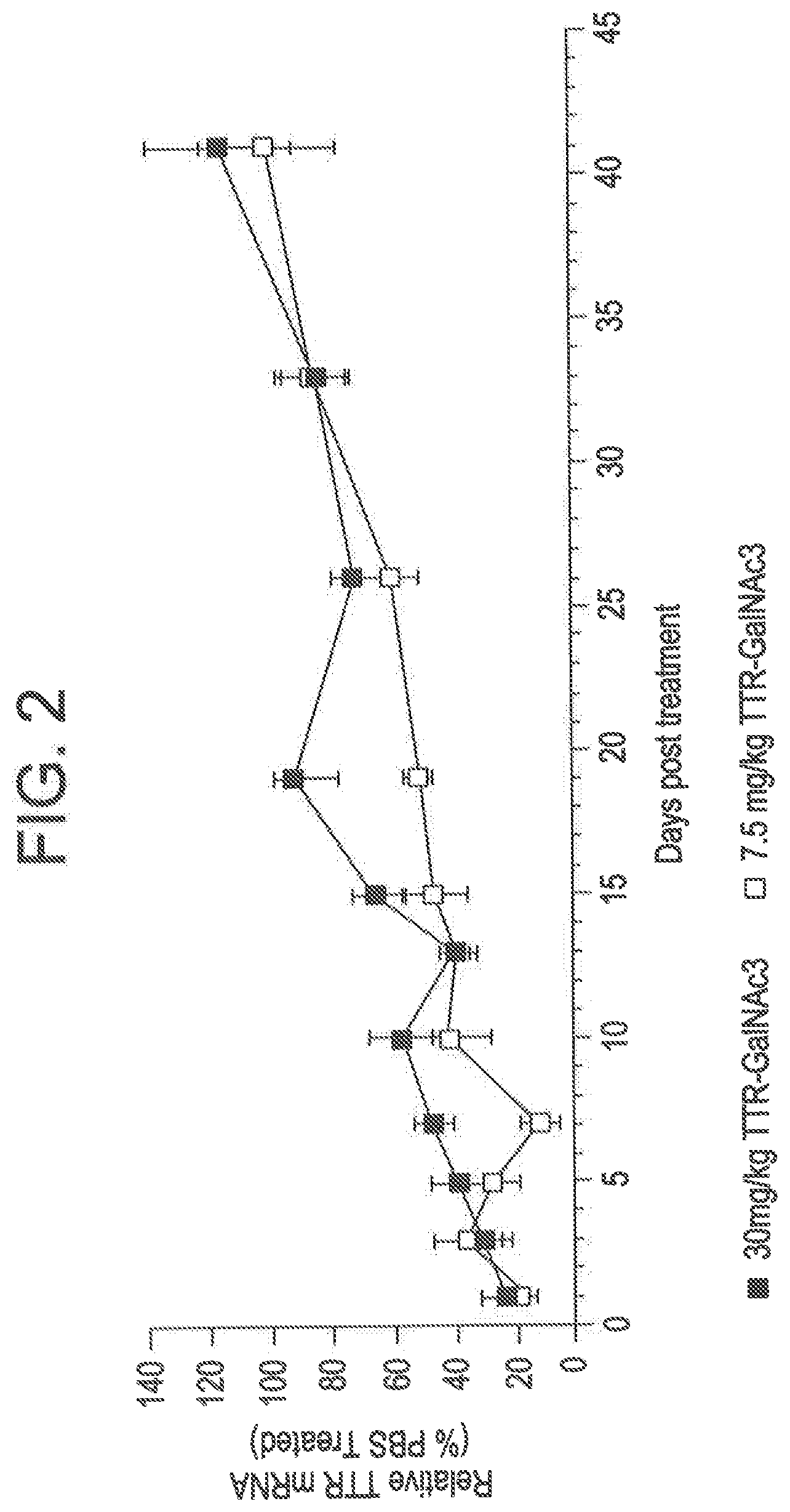
D00003

D00004
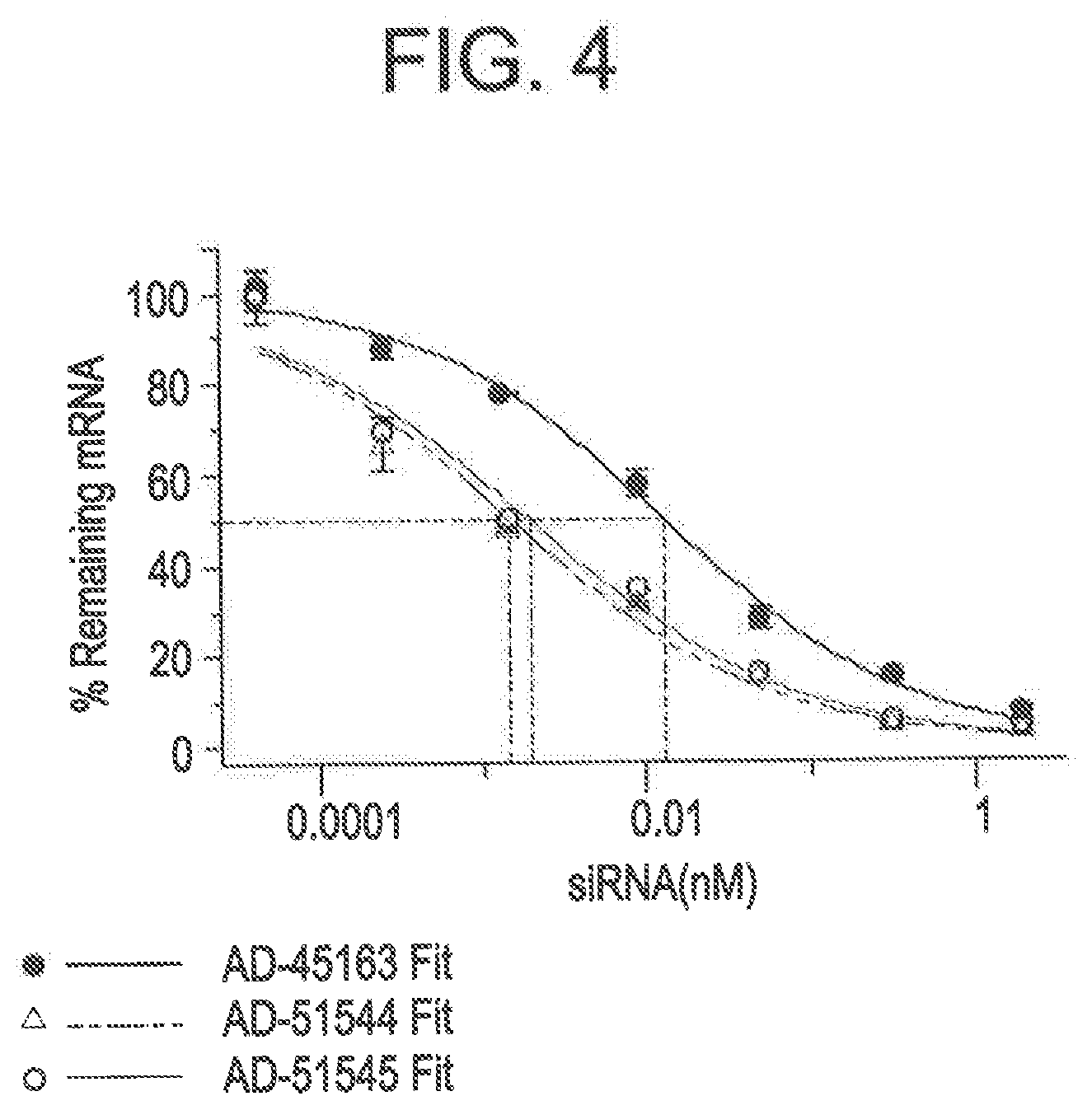
D00005

D00006
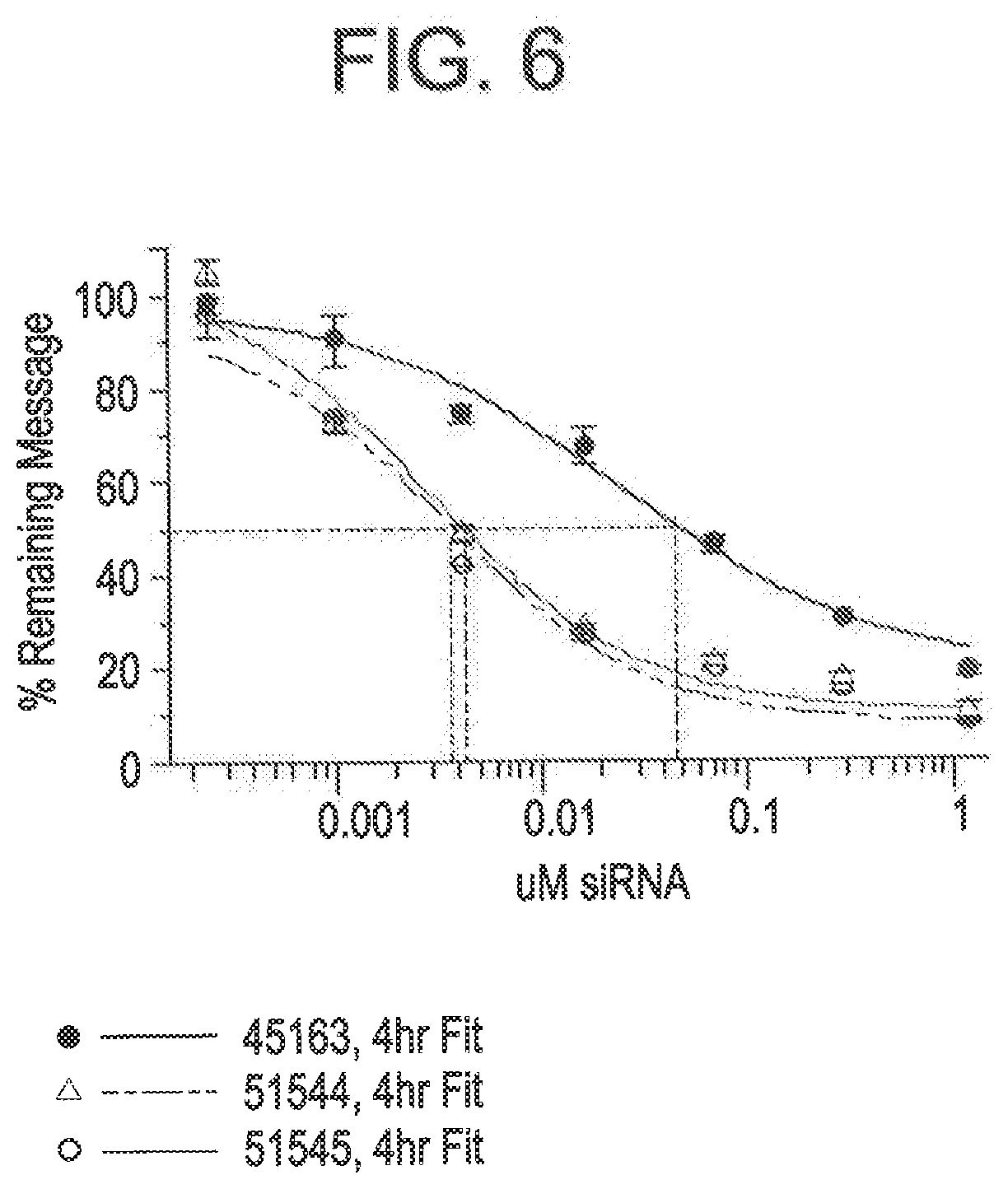
D00007

D00008

D00009

D00010

D00011

D00012
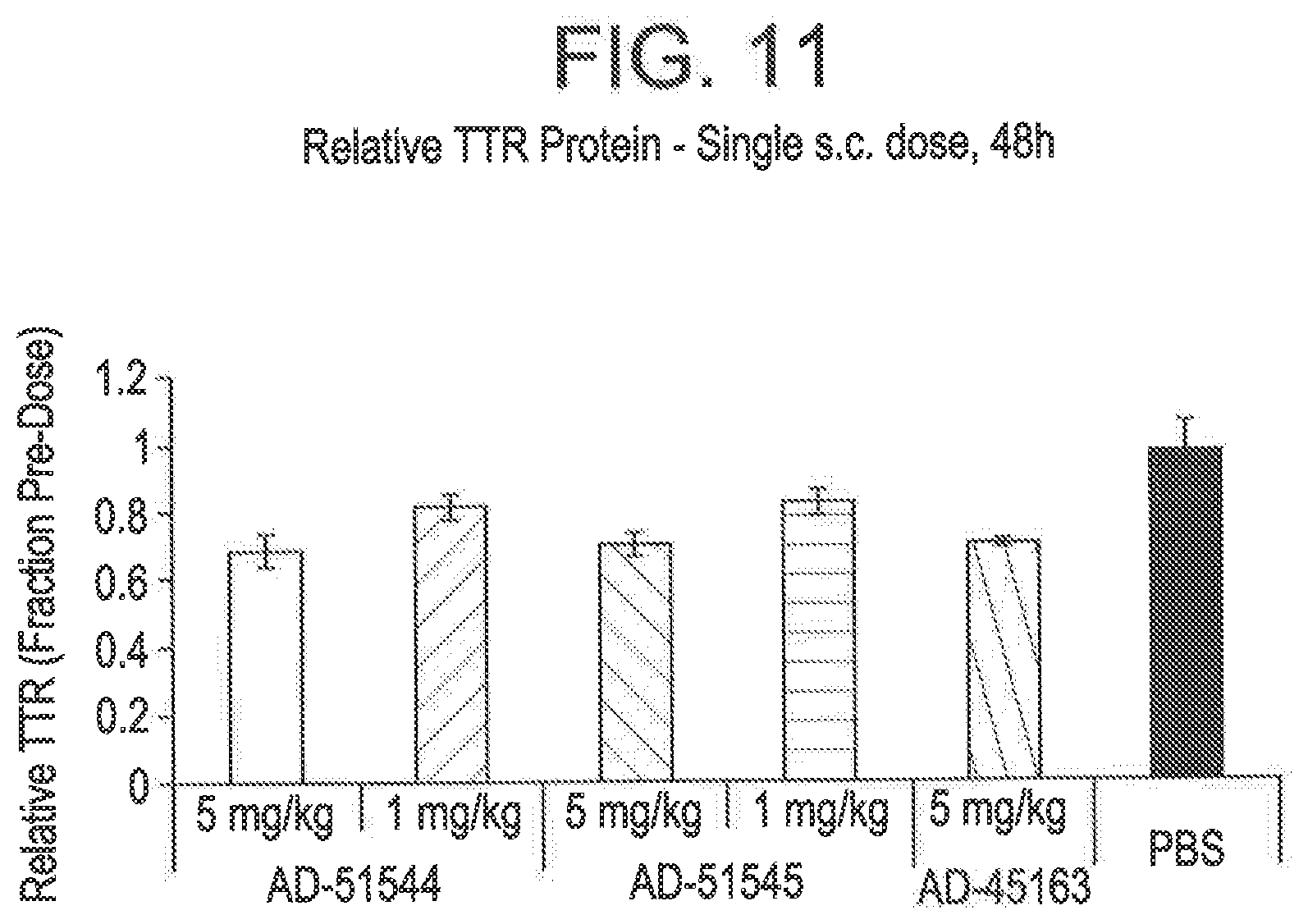
D00013

D00014

D00015

D00016


XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.