Modified Pseudomonas Exotoxin A
Pastan; Ira H. ; et al.
U.S. patent application number 16/429829 was filed with the patent office on 2020-06-11 for modified pseudomonas exotoxin a. The applicant listed for this patent is The United States of America, as represented by the Secretary,Department of Health and Human Service Hoffmann-La Roche Inc.. Invention is credited to Ulrich Brinkmann, Guy Georges, Sabine Imhof-Jung, Byungkook Lee, Ronit Mazor, Gerhard Niederfellner, Masanori Onda, Ira H. Pastan, Werner Scheuer.
| Application Number | 20200181280 16/429829 |
| Document ID | / |
| Family ID | 51794961 |
| Filed Date | 2020-06-11 |












View All Diagrams
| United States Patent Application | 20200181280 |
| Kind Code | A1 |
| Pastan; Ira H. ; et al. | June 11, 2020 |
MODIFIED PSEUDOMONAS EXOTOXIN A
Abstract
The invention provides a Pseudomonas exotoxin A (PE) comprising an amino acid sequence having a substitution of one or more B-cell and/or T-cell epitopes. The invention further provides related chimeric molecules, as well as related nucleic acids, recombinant expression vectors, host cells, populations of cells, and pharmaceutical compositions. Methods of treating or preventing cancer in a mammal, methods of inhibiting the growth of a target cell, methods of producing the PE, and methods of producing the chimeric molecule are further provided by the invention.
| Inventors: | Pastan; Ira H.; (Potomac, MD) ; Mazor; Ronit; (Rockville, MD) ; Onda; Masanori; (Rockville, MD) ; Lee; Byungkook; (Potomac, MD) ; Niederfellner; Gerhard; (Oberhausen, DE) ; Imhof-Jung; Sabine; (Planegg, DE) ; Brinkmann; Ulrich; (Weilheim, DE) ; Scheuer; Werner; (Penzberg, DE) ; Georges; Guy; (Habach, DE) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 51794961 | ||||||||||
| Appl. No.: | 16/429829 | ||||||||||
| Filed: | June 3, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15191392 | Jun 23, 2016 | |||
| 16429829 | ||||
| 14505590 | Oct 3, 2014 | 9388222 | ||
| 15191392 | ||||
| 62052665 | Sep 19, 2014 | |||
| 61982051 | Apr 21, 2014 | |||
| 61908464 | Nov 25, 2013 | |||
| 61887418 | Oct 6, 2013 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61P 35/00 20180101; C07K 16/40 20130101; A61K 38/00 20130101; C07K 2319/33 20130101; C07K 2319/50 20130101; A61K 39/39558 20130101; C07K 2317/94 20130101; C07K 2319/30 20130101; C07K 16/30 20130101; A61K 47/6803 20170801; A61K 31/337 20130101; A61K 47/6871 20170801; C07K 16/1214 20130101; A61K 2039/505 20130101; C07K 2317/55 20130101; A61K 47/6859 20170801; A61K 31/337 20130101; C07K 2317/624 20130101; A61K 39/39558 20130101; C07K 2317/24 20130101; A61K 47/6829 20170801; A61K 2300/00 20130101; A61K 2300/00 20130101; C07K 14/21 20130101; C07K 2317/92 20130101; A61K 47/6851 20170801; C07K 2317/73 20130101; C07K 16/303 20130101; C07K 2319/55 20130101 |
| International Class: | C07K 16/30 20060101 C07K016/30; C07K 14/21 20060101 C07K014/21; C07K 16/12 20060101 C07K016/12; C07K 16/40 20060101 C07K016/40; A61K 31/337 20060101 A61K031/337; A61K 47/68 20060101 A61K047/68; A61K 39/395 20060101 A61K039/395 |
Claims
1. A Pseudomonas exotoxin A (PE) comprising a PE amino acid sequence wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted, wherein the PE optionally has: (i) a further substitution of one or more amino acid residues within one or more B cell epitopes, and the further substitution for an amino acid within one or more B-cell epitopes is a substitution of, independently, one or more of amino acid residues D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, Q592, and L597 as defined by reference to SEQ ID NO: 1, (ii) a further substitution of one or more amino acid residues within one or more T-cell epitopes, (iii) a deletion of one or more continuous amino acid residues of residues 1-273 and 285-394 as defined by SEQ ID NO: 1, or (iv) a combination of any one, two, or three of (i)-(iii).
2. An isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula: R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain II wherein: n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3, R.sup.1=1 to 10 amino acid residues FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end, R.sup.2=1 to 10 amino acid residues; R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and, PE functional domain III=residues 395-613 of SEQ ID NO:1, wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted, wherein the PE optionally has: (i) a further substitution of one or more amino acid residues within one or more B cell epitopes, and the further substitution for an amino acid within one or more B-cell epitopes is a substitution of, independently, one or more of amino acid residues D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, Q592, and L597 as defined by reference to SEQ ID NO: 1, (ii) a further substitution of one or more amino acid residues within one or more T-cell epitopes, or (iii) both (i) and (ii).
3. The mutated PE of claim 2, further wherein the FCS is represented by the formula P4-P3-P2-P1, wherein P4 is an amino acid residue at the amino end, P1 is an amino acid residue at the carboxyl end, P1 is an arginine or a lysine residue, and the sequence is cleavable at the carboxyl end of P1 by furin.
4. The mutated PE of claim 3, further wherein the FCS (i) further comprises amino acid residues represented by P6-P5 at the amino end, (ii) further comprises amino acid residues represented by P1'-P2' at the carboxyl end, (iii) wherein if P1 is an arginine or a lysine residue, P2' is tryptophan, and P4 is arginine, valine or lysine, provided that if P4 is not arginine, then P6 and P2 are basic residues, and (iv) the sequence is cleavable at the carboxyl end of P1 by furin.
5. The mutated PE of any one of claims 2-4, further wherein the PE functional domain III consists of the sequence of residues 395 to 613 of SEQ ID NO: 1.
6. The mutated PE of any one of claims 2-5, wherein the mutated PE comprises one or more contiguous residues of residues 365-394 of SEQ ID NO: 1 between the FCS and the PE domain III.
7. The mutated PE of any one of claims 2-6, wherein n is 1 for R.sup.1 and R.sup.2.
8. The mutated PE of any one of claims 2-7, wherein the FCS is SEQ ID NO: 8.
9. The mutated PE of any one of claims 2-8, wherein R.sup.1=a linker of the amino acid sequence of SEQ ID NO: 282, R.sup.2=a linker of the amino acid sequence SEQ ID NO: 284, and the FCS=SEQ ID NO: 8.
10. The mutated PE of any one of claims 2-9, wherein n is 0 for R.sup.3.
11. The mutated PE of any one of claims 2-10, wherein PE functional domain III comprises the amino acid sequence of SEQ ID NO: 37.
12. The mutated PE of any one of claims 2-11, wherein R.sup.1.sub.n-FCS-R.sup.2.sub.n=SEQ ID NO: 36.
13. The PE of any one of claims 1-12, wherein the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of, independently, alanine, glutamic acid, histidine, or asparagine in place of one or more of amino acid residues F443, R456, L477, R494, and L552.
14. The PE of any one of claims 1-13, wherein the substitution of L552 is a substitution of glutamic acid or asparagine in place of L552 and the substitution of L4A77 is a substitution of histidine in place of L477.
15. The PE of any one of claims 1-14, wherein the further substitution of an amino acid within one or more B-cell epitopes is a substitution of, independently, alanine, glycine, serine, or glutamine in place of one or more of amino acid residues E282, E285, P290, R313, N314, P319, D324, E327, E331, Q332, D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, K590, Q592, and L597, as defined by reference to SEQ ID NO: 1.
16. The PE of any one of claims 1-15, wherein the PE has the further substitution of an amino acid within one or more T-cell epitopes, and the further substitution of an amino acid within one or more T-cell epitopes is a substitution of, independently, alanine, glycine, serine, or glutamine in place of one or more of amino acid residues R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 as defined by reference to SEQ ID NO: 1.
17. The PE of any one of claims 1-16, wherein the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of alanine in place of amino acid residue R456; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
18. The PE of any one of claims 1-16, wherein the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue R456, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
19. The PE of any one of claims 1-16, wherein the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of alanine in place of amino acid residue R456; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of asparagine in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
20. The mutated PE of claim 2, wherein one or more of amino acid residues F443, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted.
21. The mutated PE of claim 20, wherein the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443, a substitution of histidine in place of amino acid residue L477, a substitution of alanine in place of amino acid residue R494, and a substitution of glutamic acid or asparagine in place of amino acid residue L552.
22. An isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula: R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III wherein: n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3, R.sup.1=1 to 10 amino acid residues FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end, R.sup.2=1 to 10 amino acid residues; R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and, PE functional domain III=residues 395-613 of SEQ ID NO:1, wherein the PE includes an arginine at position 458, as defined by reference to SEQ ID NO: 1, and wherein the PE has: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538.
23. A chimeric molecule comprising (a) a targeting moiety conjugated or fused to (b) the PE of any one of claims 1-22.
24. The chimeric molecule of claim 23, wherein the targeting moiety is a monoclonal antibody or an antigen binding portion of the monoclonal antibody.
25. The chimeric molecule of claim 24, wherein the monoclonal antibody or antigen binding portion of the monoclonal antibody specifically binds to a cell surface marker selected from the group consisting of CD19, CD21, CD22, CD25, CD30, CD79b, transferrin receptor, epidermal growth factor (EGF) receptor, mesothelin, cadherin, Lewis Y, glypican-3, FAP (fibroblast activation protein alpha), PSMA (prostate specific membrane antigen), CA9=CAIX (carbonic anhydrase IX), LICAM (neural cell adhesion molecule L1), endosialin, HER3 (activated conformation of epidermal growth factor receptor family member 3), Alkl/BMP9 complex (anaplastic lymphoma kinase 1/bone morphogenetic protein 9), TPBG=5T4 (trophoblast glycoprotein), CD33 (sialic acid binding Ig-like lectin 3, myeloid cell surface antigen), CD123 (interleukin 3 receptor alpha), MUC1 (tumor-associated epithelial mucin), ROR1 (receptor tyrosine kinase-like surface antigen), HER1 (activated conformation of epidermal growth factor receptor), and CLL1 (C-type lectin domain family 12, member A).
26. The chimeric molecule of claim 23, wherein the targeting moiety is selected from the group consisting of B3, RFB4, SS, SS1, MN, MB, HN1, HN2, HB21, MORAb-009, antigen binding portions thereof, and the antigen binding portion of HA22.
27. The chimeric molecule of claim 23, wherein the targeting moiety is a humanized SS1 or an antigen binding portion of the humanized SS1.
28. The chimeric molecule of claim 23, wherein the targeting moiety comprises: (a) SEQ ID NOs: 31 and 34; (b) SEQ ID NOs: 45 and 46; (c) SEQ ID NOs: 61 and 62; (d) SEQ ID NOs: 77 and 78; (e) SEQ ID NOs: 93 and 94; (f) SEQ ID NOs: 109 and 110; (g) SEQ ID NOs: 125 and 126; (h) SEQ ID NOs: 141 and 142; (i) SEQ ID NOs: 157 and 158; (j) SEQ ID NOs: 173 and 174; (k) SEQ ID NOs: 49, 50, 53, 54, 57, and 58; (l) SEQ ID NOs: 65, 66, 69, 70, 73, and 74; (m) SEQ ID NOs: 81, 82, 85, 86, 89, and 90; (n) SEQ ID NOs: 97, 98, 101, 102, 105, and 106; (o) SEQ ID NOs: 113, 114, 117, 118, 121, and 122; (p) SEQ ID NOs: 129, 130, 133, 134, 137, and 138; (q) SEQ ID NOs: 145, 146, 149, 150, 153, and 154; (r) SEQ ID NOs: 161, 162, 165, 166, 169, and 170; (s) SEQ ID NOs: 177, 178, 181, 182, 185, and 186; (t) SEQ ID NOs: 31-32 and 34-36; (u) SEQ ID NOs: 33 and 38; or (v) SEQ ID NOs: 93 and 290.
29. The chimeric molecule of any one of claims 23-28, wherein the chimeric molecule comprises a linker comprising SEQ ID NO: 36.
30. The chimeric molecule of any one of claims 23-28, comprising (a) SEQ ID NOs: 39 and 40; (b) SEQ ID NOs: 41 and 42; (c) SEQ ID NOs: 43 and 44; (d) SEQ ID NOs: 291 and 293; (e) SEQ ID NOs: 291 and 294; (f) SEQ ID NOs: 292 and 294; (g) SEQ ID NOs: 295 and 297; or (h) SEQ ID NOs:296 and 297.
31. A nucleic acid comprising a nucleotide sequence encoding the PE of any one of claims 1-22 or the chimeric molecule of any one of claims 23-30.
32. A recombinant expression vector comprising the nucleic acid of claim 31.
33. A host cell comprising the recombinant expression vector of claim 32.
34. A population of cells comprising at least one host cell of claim 33.
35. A pharmaceutical composition comprising (a) the PE of any one of claims 1-22, the chimeric molecule of any one of claims 23-30, the nucleic acid of claim 31, the recombinant expression vector of claim 32, the host cell of claim 33, or the population of cells of claim 34, and (b) a pharmaceutically acceptable carrier.
36. The PE of any one of claims 1-22, the chimeric molecule of any one of claims 23-30, the nucleic acid of claim 31, the recombinant expression vector of claim 32, the host cell of claim 33, the population of cells of claim 34, or the pharmaceutical composition of claim 35, for use in treating or preventing cancer in the mammal.
37. The PE of any one of claims 1-22, the chimeric molecule of any one of claims 23-30, the nucleic acid of claim 31, the recombinant expression vector of claim 32, the host cell of claim 33, the population of cells of claim 34, or the pharmaceutical composition of claim 35, for use in inhibiting growth of a target cell.
38. The PE, chimeric molecule, nucleic acid, recombinant expression vector, host cell, population of cells, or pharmaceutical composition for the use of claim 37, wherein the target cell is a cancer cell.
39. The PE, chimeric molecule, nucleic acid, recombinant expression vector, host cell, population of cells, or pharmaceutical composition for the use of claim 37, wherein the target cell expresses a cell surface marker selected from the group consisting of CD19, CD21, CD22, CD25, CD30, CD79b, transferrin receptor, EGF receptor, mesothelin, cadherin, Lewis Y, glypican-3, FAP (fibroblast activation protein alpha), PSMA (prostate specific membrane antigen), CA9=CAIX (carbonic anhydrase IX), LICAM (neural cell adhesion molecule L1), Endosialin, HER3 (activated conformation of epidermal growth factor receptor family member 3), Alkl/BMP9 complex (anaplastic lymphoma kinase 1/bone morphogenetic protein 9), TPBG=5T4 (trophoblast glycoprotein), CD33 (sialic acid binding Ig-like lectin 3, myeloid cell surface antigen), CD123 (interleukin 3 receptor alpha), MUC1 (tumor-associated epithelial mucin), ROR1 (receptor tyrosine kinase-like surface antigen), HER1 (activated conformation of epidermal growth factor receptor), and CLL1 (C-type lectin domain family 12, member A).
40. A method of producing the PE of any one of claims 1-22 comprising (a) recombinantly expressing the PE and (b) purifying the PE.
41. A method of producing the chimeric molecule of any one of claims 23-30 comprising (a) recombinantly expressing the chimeric molecule and (b) purifying the chimeric molecule.
42. A method of producing the chimeric molecule of any one of claims 23-30 comprising (a) recombinantly expressing the PE of any one of claims 1-22, (b) purifying the PE, and (c) covalently linking a targeting moiety to the purified PE.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This patent application claims the benefit of U.S. Provisional Patent Application Nos. 61/887,418, filed Oct. 6, 2013; 61/908,464, filed Nov. 25, 2013; 61/982,051, filed Apr. 21, 2014; and 62/052,665, filed Sep. 19, 2014, each of which is incorporated herein by reference in its entirety.
INCORPORATION-BY-REFERENCE OF MATERIAL SUBMITTED ELECTRONICALLY
[0002] Incorporated by reference in its entirety herein is a computer-readable nucleotide/amino acid sequence listing submitted concurrently herewith and identified as follows: one 144,473 Byte ASCII (Text) file named "718352_ST25.txt," dated Oct. 1, 2014.
BACKGROUND OF THE INVENTION
[0003] Pseudomonas exotoxin A (PE) is a bacterial toxin with cytotoxic activity that may be effective for destroying or inhibiting the growth of undesirable cells, e.g., cancer cells. Accordingly, PE may be useful for treating or preventing diseases such as, e.g., cancer. However, PE may be highly immunogenic. Accordingly, PE administration may stimulate an anti-PE immune response including, for example, the production of anti-PE antibodies and/or T-cells, that undesirably neutralizes the cytotoxic activity of PE. Such immunogenicity may reduce the amount of PE that can be given to the patient which may, in turn, reduce the effectiveness of the PE for treating the disease, e.g., cancer. Thus, there is a need for improved PE.
[0004] Several deimmunized Pseudomonas exotoxins (PE) are known in art. The domain II deleted versions (for example, PE24) may be less immunogenic and may cause fewer side effects (such as, for example, capillary leak syndrome and hepatotoxicity) as compared to PE38, which contains domain II. Without being bound to a particular theory, it is believed that the reduced immunogenicity and fewer side effects of PE24 could, at least in part, be due to the reduced size of PE24, which disadvantageously results in a shorter serum half life. Different furin cleavable linkers may be employed in PE24 variants. PE immunoconjugates have mostly used dsFv fragments as targeting moieties. Such deimmunized Pseudomonas exotoxins (PE) are described in, for example, International Patent Application Publications WO2005052006, WO2007016150, WO2007014743, WO2007031741, WO200932954, WO201132022, WO2012/154530, and WO 2012/170617.
[0005] Previous immunotoxins have many disadvantages. For example, deimmunization of previous immuntoxins has been incomplete with respect to the human B-cell epitopes because immunogenic reactions still occurred. In addition, the deimmunization of previous immunotoxins was accompanied by a reduced cytotoxic potency. For example, a LO10 deimmunized PE variant described in WO 2012/170617 provided a loss of potency of at least 40% compared to wild type (WT) PE and other PE variants. In International Patent Application Publication WO2013/040141, Pseudomonas exotoxins with less immunogenic B-cell epitopes have been described. In the PE variant LRO10, all B-cell epitopes were removed. This, however, also led to a reduction of cytotoxicity towards tumor cells.
[0006] In addition, fusion of a dsFv with domain II deleted versions of PE (PE24) have a shorter serum half life due to their reduced overall size as compared to dsFv fusions with PE38. The linkers of previous immunoxins also contained T-cell epitopes and poor developability such as, for example, a poor stability at 37.degree. C. In addition, previous anti-mesothelin (MSLN) immunotoxins have only used mouse-derived dsFv fragments fused to PE, which may further contribute to immunogenicity. International Patent Application Publication WO 2012/154530 refers to Pseudomonas exotoxin variant chimeric molecules with short flexible linkers which improve the cytotoxicity towards tumor cells.
BRIEF SUMMARY OF THE INVENTION
[0007] The invention relates to deimmunized Pseudomonas exotoxins and Fab fusions thereof (e.g., humanized anti-MSLN), methods for the treatment of cancer, stabilized pharmaceutical formulations, methods for the reduction of side effects and methods for enhancing the serum half life and optimizing treatment schedule.
[0008] An embodiment of the invention provides a Pseudomonas exotoxin A (PE) comprising a PE amino acid sequence, wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted, wherein the PE optionally has:
[0009] (i) a further substitution of one or more amino acid residues within one or more B cell epitopes, and the further substitution for an amino acid within one or more B-cell epitopes is a substitution of, independently, one or more of amino acid residues D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, Q592, and L597 as defined by reference to SEQ ID NO: 1,
[0010] (ii) a further substitution of one or more amino acid residues within one or more T-cell epitopes,
[0011] (iii) a deletion of one or more continuous amino acid residues of residues 1-273 and 285-394 as defined by SEQ ID NO: 1, or
[0012] (iv) a combination of any one, two, or three of (i)-(iii).
[0013] Another embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0014] wherein:
[0015] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3,
[0016] R.sup.1=1 to 10 amino acid residues
[0017] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0018] R.sup.2=1 to 10 amino acid residues;
[0019] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO:1; and,
[0020] PE functional domain III=residues 395-613 of SEQ ID NO:1, wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted, wherein the PE optionally has:
[0021] (i) a further substitution of one or more amino acid residues within one or more B cell epitopes, and the further substitution for an amino acid within one or more B-cell epitopes is a substitution of, independently, one or more of amino acid residues D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, Q592, and L597 as defined by reference to SEQ ID NO: 1,
[0022] (ii) a further substitution of one or more amino acid residues within one or more T-cell epitopes, or
[0023] (iii) both (i) and (ii).
[0024] Another embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0025] wherein:
[0026] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3,
[0027] R.sup.1=1 to 10 amino acid residues
[0028] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0029] R.sup.2=1 to 10 amino acid residues;
[0030] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO:1; and,
[0031] PE functional domain III=residues 395-613 of SEQ ID NO:1, wherein the PE includes an arginine at position 458, as defined by reference to SEQ ID NO: 1, and
[0032] wherein the PE has:
[0033] (a) a substitution of alanine for amino acid residue R427;
[0034] (b) a substitution of alanine for amino acid residue R463;
[0035] (c) a substitution of alanine for amino acid residue R467;
[0036] (d) a substitution of alanine for amino acid residue R490;
[0037] (e) a substitution of alanine for amino acid residue R505; and
[0038] (f) a substitution of alanine for amino acid residue R538.
[0039] Additional embodiments of the invention provide related chimeric molecules, as well as related nucleic acids, recombinant expression vectors, host cells, populations of cells, and pharmaceutical compositions.
[0040] Still another embodiment of the invention provides a method of treating or preventing cancer in a mammal comprising administering to the mammal the inventive PE, chimeric molecule, nucleic acid, recombinant expression vector, host cell, population of cells, or pharmaceutical composition, in an amount effective to treat or prevent cancer in the mammal.
[0041] Another embodiment of the invention provides a method of inhibiting the growth of a target cell comprising contacting the cell with the inventive PE, chimeric molecule, nucleic acid, recombinant expression vector, host cell, population of cells, or pharmaceutical composition, in an amount effective to inhibit growth of the target cell.
[0042] Additional embodiments of the invention provide methods of producing the inventive PE and methods of producing the inventive chimeric molecule.
BRIEF DESCRIPTION OF THE SEVERAL VIEWS OF THE DRAWINGS
[0043] FIGS. 1A and 1B are T cell epitope heat maps showing the strongest (>20%, black squares), medium (10-20%, dark grey squares), weak (3%-10%, light grey squares) and negative (absence of response; <3%, white squares) responses for naive donors (n=50) (A) and previously treated patients (n=16) (B). The responses are shown as a percentage of responsive spots for each donor. Responses were clustered using automatic sorting based on the responsiveness of the pools.
[0044] FIGS. 2A-2D are graphs showing the response of three donor samples (A-C) and one HCL patient sample (D) to one of 22 peptide pools, control pool (CEFT), or no peptide after stimulation with HA22 (shaded bars) or LR-R494A (unshaded bars) as measured in spot-forming cells (SFCs) per 10.times.10.sup.6 cells. * indicates statistical significance between I-HA22 and LR-R494A (p<0.01).
[0045] FIGS. 3A-3C are graphs showing the response of two donor samples (A-B) and one HCL patient sample (C) to one of 22 peptide pools, CEFT, or no peptide after stimulation with HA22 (shaded bars) or LR-R505A (unshaded bars) as measured in SFCs per 10.times.10.sup.6 cells. * indicates statistical significance (p<0.01).
[0046] FIGS. 4A-4B are graphs showing the response of two donor samples (A-B) to one of 22 peptide pools, CEFT, or no peptide after stimulation with HA22 (shaded bars) or LR-R551A (unshaded bars) as measured in SFCs per 10.times.10.sup.6 cells. * indicates statistical significance in student T test (p<0.01).
[0047] FIGS. 5A-5D are graphs showing the response of two donor samples (C-D) and two mesothelioma patient samples (A-B) after stimulation with recombinant immunotoxin (RIT) and restimulation with peptide 93 or 94 with either the wild-type (WT) amino acid sequence (black bars), L552N (dark grey bars), or L552E (unshaded bars), or treatment with no peptide (light grey bars). * indicates statistical significance in student T test (p<0.05).
[0048] FIGS. 6A-6C are graphs showing the response of three donor samples to 22 peptide pools after stimulation with either HA22-LR (WT) (shaded bars) or LR-R427A (unshaded bars) and restimulation with the appropriate peptides as measured in SFCs per 10.times.10.sup.6 cells. * indicates statistical significance (p<0.05).
[0049] FIGS. 7A-7C are graphs showing the response of two patient samples (A-B) and one donor (C) to one of pools 8-22 after stimulation with either HA22-LR (WT) (shaded bars) or LR-F443A (unshaded bars) and restimulation with the appropriate peptides as measured in SFCs per 10.times.10.sup.6 cells. * indicates statistical significance (p<0.05).
[0050] FIG. 8 is a graph showing the fold change in EC50 of HA22-LR-GGS (circles), HA22 (vertical dashes), or HA22-LR-T18 (triangles) at various temperatures.
[0051] FIGS. 9A-9D are graphs showing the response of four donor samples after stimulation with RIT and restimulation with no peptide, WT peptide 67, or peptide 67 with either a valine or alanine substitution at position 471 as measured in SFCs per 10.times.10.sup.6 cells.
[0052] FIGS. 10A-10E are graphs showing the response of three donor samples (A-C), an HCL patient sample (D), and a mesothelioma patient sample (E) after stimulation with peptide 67 or 68 that contains an alanine mutation (white bars) or histidine mutation (grey bars) at position 477 or no mutation (WT) (black bars).
[0053] FIGS. 11A and 11B are graphs showing the response of samples from a meothelioma patient (A) and a hairy cell leukemia (HCL) patient (B) after stimulation with RIT and restimulation with no peptide, peptide 93, 94 or 95 with either the WT amino acid sequence (shaded bars) or L556V (unshaded bars).
[0054] FIG. 12 is a graph showing the aggregation (% Area) of HMW cFP-0170 (Fab-LO10R-456A short linker; diagonally striped bars) or HMW cFP-0171 (Fab-LO10R-456A elongated liker; horizontally striped bars) as measured by size exclusion chromatograph (SEC) after incubation at 33.degree. C.
[0055] FIG. 13 is a graph showing the aggregation (% Area) of HMW cFP-0170 (Fab-LO10R-456A short linker; diamonds) or HMW cFP-0171 (Fab-LO10R-456A elongated liker; squares) as measured by size exclusion chromatograph (SEC) after incubation at 33.degree. C.
[0056] FIG. 14 is a graph showing the aggregation (radius in nm) of cFP-0166 (Fab-LO10R short linker; diamonds) or cFP-0174 (Fab-LO10R elongated linker; circles) as measured by dynamic light scattering (DLS) at various temperatures in a range from 25.degree. C. to 50.degree. C.
[0057] FIG. 15 is a graph showing the aggregation (radius in nm) of cFP-0170 (Fab-LO10R-456A short linker; diamonds) or cFP-0171 (Fab-LO10R-456A elongated linker; circles) as measured by DLS at various temperatures in a range from 25.degree. C. to 50.degree. C.
[0058] FIG. 16 is a graph showing the aggregation (radius in nm) of cFP-0172 (Fab-LO10R-456A-551A short linker; diamonds) or cFP-0173 (Fab-LO10R-456A-551A elongated linker; circles) as measured by DLS at various temperatures in a range from 25.degree. C. to 50.degree. C.
[0059] FIG. 17 is a graph showing the response of patient sera clone 9H3 to (10.sup.7) or (10.sup.8) phages/well or (10.sup.7) or (10.sup.8) phages/well with antigen pre-incubation incubated with huSS1Fab-PE24LRO10 (with 458A mutation) (left diagonally striped bars), huSS1Fab-PE24LRO10R (with 458R backmutation) (horizontally striped bars), or huSS1Fab-PE24LRO10R-456A (with 458R backmutation and 456A mutation) (right diagonally striped bars) as measured in optical density at 450 nm.
[0060] FIGS. 18A and 18B are graphs showing the antigenicity of the chimeric molecules SS1P (T1), SS1-dsFv-LR-LO10R (T2), SS1-dsFv-LR-LO10R456A (T3), SS1-FABLO10R (Roche 116, T4), SS1-FABLO10R456ALongLinker (Roche 171, T5), and SS1-FABLO10RLongLinker (Roche 174, T6) with respect to 20 patient sera. Two representative examples (A and B) are shown. The Y axis is relative IC50(%).
[0061] FIG. 19 is a graph showing plasma concentration (ng/ml) of chimeric molecules SS1P38 (0.4 mg/kg (diamonds)), SS1P38 (0.2 mg/kg (squares)), Fab-PE24 (0.231 mg/kg (triangles)), (dsFv-PE24 (0.16 mg/kg (x)) administered to mice over a period of time (hours (h)).
[0062] FIG. 20 is a graph showing the serum half life cFP (ng/ml) of 0.3 mg/kg cFP_0205 (squares), 0.3 mg/kg SS1P (circles), or 0.3 mg/kg cFP_0205 (dashed lines) over time (hours) in cyno monkeys.
[0063] FIG. 21 is a graph showing the body weight change of of mice treated with control (vehicle; circles); cFP 0205 3 mg/kg i.v. 3q7d (squares on solid line); cFP 0205 2 mg/kg i.v. 3q7d (squares on larger dashed line); cFP 0205 1 mg/kg i.v. 3q7d (squares on short dashed line); cFP 0205 0.5 mg/kg i.v. 3q7d (squares on dotted line) over time (days after cell inoculation).
[0064] FIGS. 22A-F are fluorescence images of sections of lung (A-B), spleen (C-D) or liver (E-F) of mice treated with labeled cFP0205 at 10.times. (A, C, E) or 40.times. (B, D, F) magnification.
[0065] FIGS. 22G-L are fluorescence images of sections of lung (G-H), spleen (I-J) or liver (K-L) of mice treated with labeled SS1P at 10.times. (G, I, K) or 40.times. (H, J, L) magnification.
[0066] FIG. 23 is a graph showing the tumor size of mice that were untreated (open diamonds) or treated with paclitaxel alone, (grey diamonds) RG7787 (also referred to as R205 or cFP 0205) alone (black diamonds), or a combination of RG7787 and paclitaxel (black and white diamonds) over time measured in days.
[0067] FIG. 24 is a graph showing the tumor size of mice that were untreated (vehicle) (circles) or treated with RG7787 (squares) over time measured in days. Each data point represents the average of mean tumor volume for n=9 animals treated with RG7787 and n=8 control animals. Error bars show standard deviations.
[0068] FIG. 25 is a graph showing the tumor size of mice that were treated with vehicle (control) (circles), RG7787 alone (squares), paclitaxel alone (triangles), or a combination of RG7787 and paclitaxel (diamonds) over time measured in days. Error bars show standard deviations.
[0069] FIG. 26 is a graph showing the tumor size of untreated (UT) (x) mice or mice treated with R205 alone (diamonds), a combination of R205 and taxol (squares), or taxol alone (triangles) over time measured in days.
[0070] FIG. 27A is a graph showing cell viability (%) of CD22-expressing cell lines treated with various concentrations of T18 or MP RIT (ng/ml).
[0071] FIG. 27B is a graph showing the relative cytotoxic activity (%) of T18 or MP RIT that had been heated to one of various temperatures (.degree. C.).
[0072] FIGS. 27C-D are graphs showing the cytotoxic activity (IC50 (ng/ml)) of MP or T18 RIT on cells from hairy cell leukemia (HCL) (C) or chronic lymphocytic leukemia (CLL) (D) patients.
[0073] FIG. 27E is a graph showing the effect of LMB-T18 on tumor size in a xenograft mouse model after four injections of 5 mg/kg (squares) of LMB-T18, three injections of 7.5 mg/kg (triangles) OF LMB-T18, or PBS-0.2% human serum albumin (circles). Arrows represent days of injection for all dose groups. Broken arrow indicates additional injection of 5 mg/ml group. (*) P>0.01 in one-way ANOVA. Error bars indicate SD.
[0074] FIG. 27F is a graph showing the % binding of MP, HA22-LR, and LMB-T18 to serum from patients with neutralizing antibodies to MP.
DETAILED DESCRIPTION OF THE INVENTION
[0075] Pseudomonas exotoxin A ("PE") is a bacterial toxin (molecular weight 66 kD) secreted by Pseudomonas aeruginosa. The native, wild-type PE sequence (SEQ ID NO: 1) is set forth in U.S. Pat. No. 5,602,095, which is incorporated herein by reference. Native, wild-type PE includes three structural domains that contribute to cytotoxicity. Domain Ia (amino acids 1-252) mediates cell binding, domain II (amino acids 253-364) mediates translocation into the cytosol, and domain III (amino acids 400-613) mediates ADP ribosylation of elongation factor 2. While the structural boundary of domain III of PE is considered to start at residue 400, it is contemplated that domain III may require a segment of domain Ib to retain ADP-ribosylating activity. Accordingly, functional domain III is defined as residues 395-613 of PE. The function of domain Ib (amino acids 365-399) remains undefined. Without being bound by a particular theory or mechanism, it is believed that the cytotoxic activity of PE occurs through the inhibition of protein synthesis in eukaryotic cells, e.g., by the inactivation of the ADP-ribosylation of elongation factor 2 (EF-2).
[0076] Substitutions of PE are defined herein by reference to the amino acid sequence of PE. Thus, substitutions of PE are described herein by reference to the amino acid residue present at a particular position, followed by the amino acid with which that residue has been replaced in the particular substitution under discussion. In this regard, the positions of the amino acid sequence of a particular embodiment of a PE are referred to herein as the positions of the amino acid sequence of the particular embodiment or as the positions as defined by SEQ ID NO: 1. When the positions are as defined by SEQ ID NO: 1, then the actual positions of the amino acid sequence of a particular embodiment of a PE are defined relative to the corresponding positions of SEQ ID NO: 1 and may represent different residue position numbers than the residue position numbers of SEQ ID NO: 1. Thus, for example, substitutions refer to a replacement of an amino acid residue in the amino acid sequence of a particular embodiment of a PE corresponding to the indicated position of the 613-amino acid sequence of SEQ ID NO: 1 with the understanding that the actual positions in the respective amino acid sequences may be different. For example, when the positions are as defined by SEQ ID NO: 1, the term "R490" refers to the arginine normally present at position 490 of SEQ ID NO: 1, "R490A" indicates that the arginine normally present at position 490 of SEQ ID NO: 1 is replaced by an alanine, while "K590Q" indicates that the lysine normally present at position 590 of SEQ ID NO: 1 has been replaced with a glutamine. In the event of multiple substitutions at two or more positions, the two or more substitutions may be the same or different, i.e., each amino acid residue of the two or more amino acid residues being substituted can be substituted with the same or different amino acid residue unless explicitly indicated otherwise.
[0077] The terms "Pseudomonas exotoxin" and "PE" as used herein include PE that has been modified from the native protein to reduce or to eliminate immunogenicity. Such modifications may include, but are not limited to, elimination of domain Ia, various amino acid deletions in domains Ib, II, and III, single amino acid substitutions and the addition of one or more sequences at the carboxyl terminus such as DEL and REDL (SEQ ID NO: 7). See Siegall et al., J. Biol. Chem., 264: 14256-14261 (1989). Such modified PEs may be further modified to include any of the inventive substitution(s) for one or more amino acid residues within one or more T-cell and/or B-cell epitopes described herein. In an embodiment, the modified PE may be a cytotoxic fragment of native, wild-type PE. Cytotoxic fragments of PE may include those which are cytotoxic with or without subsequent proteolytic or other processing in the target cell (e.g., as a protein or pre-protein). In a preferred embodiment, the cytotoxic fragment of PE retains at least about 20%, preferably at least about 40%, more preferably at least about 50%, even more preferably at least about 75%, more preferably at least about 90%, and still more preferably at least about 95% of the cytotoxicity of native PE. In particularly preferred embodiments, the cytotoxic fragment has at least the cytotoxicity of native PE, and preferably has increased cytotoxicity as compared to native PE.
[0078] Modified PE that reduces or eliminates immunogenicity includes, for example, PE4E, PE40, PE38, PE25, PE38QQR, PE38KDEL, and PE35. In an embodiment, the PE may be any of PE4E, PE40, PE38, PE25, PE38QQR (in which PE38 has the sequence QQR added at the C-terminus), PE38KDEL (in which PE38 has the sequence KDEL (SEQ ID NO: 5) added at the C-terminus), PE-LR (resistance to lysosomal degradation)(also referred to as PE24), PE24-LO10, and PE35.
[0079] In an embodiment, the PE has been modified to reduce immunogenicity by deleting domain Ia as described in in U.S. Pat. No. 4,892,827, which is incorporated herein by reference. The PE may also be modified by substituting certain residues of domain Ia. In an embodiment, the PE may be PE4E, which is a substituted PE in which domain Ia is present but in which the basic residues of domain Ia at positions 57, 246, 247, and 249 are replaced with acidic residues (e.g., glutamic acid), as disclosed in U.S. Pat. No. 5,512,658, which is incorporated herein by reference.
[0080] PE40 is a truncated derivative of PE (Pai et al., Proc. Nat'lAcad. Sci. USA, 88: 3358-62 (1991) and Kondo et al., Biol. Chem., 263: 9470-9475 (1988)). PE35 is a 35 kD carboxyl-terminal fragment of PE in which amino acid residues 1-279 have been deleted and the molecule commences with a Met at position 280 followed by amino acids 281-364 and 381-613 of native PE. PE35 and PE40 are disclosed, for example, in U.S. Pat. Nos. 5,602,095 and 4,892,827, each of which is incorporated herein by reference. PE25 contains the 11-residue fragment from domain II and all of domain III. In some embodiments, the PE contains only domain III.
[0081] In a preferred embodiment, the PE is PE38. PE38 contains the translocating and ADP ribosylating domains of PE but not the cell-binding portion (Hwang J. et al., Cell, 48: 129-136 (1987)). PE38 is a truncated PE pro-protein composed of amino acids 253-364 and 381-613 which is activated to its cytotoxic form upon processing within a cell (see e.g., U.S. Pat. No. 5,608,039, which is incorporated herein by reference, and Pastan et al., Biochim. Biophys. Acta, 1333: C1-C6 (1997)).
[0082] In another preferred embodiment, the PE is PE-LR. PE-LR contains a deletion of domain II except for a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a deletion of amino acid residues 365-394 of domain Ib. Thus, PE-LR contains amino acid residues 274-284 and 395-613 of SEQ ID NO: 1. PE-LR is described in International Patent Application Publication WO 2009/032954, which is incorporated herein by reference. The PE-LR may, optionally, additionally comprise a GGS (SEQ ID NO: 283) linking peptide between the FCS and amino acid residues 395-613 of SEQ ID NO: 1.
[0083] As noted above, alternatively or additionally, some or all of domain Ib may be deleted with the remaining portions joined by a bridge or directly by a peptide bond. Alternatively or additionally, some of the amino portion of domain II may be deleted. Alternatively or additionally, the C-terminal end may contain the native sequence of residues 609-613 (REDLK) (SEQ ID NO: 6), or may contain a variation that may maintain the ability of the PE to translocate into the cytosol, such as KDEL (SEQ ID NO: 5) or REDL (SEQ ID NO: 7), and repeats of these sequences. See, e.g., U.S. Pat. Nos. 5,854,044; 5,821,238; and 5,602,095 and International Patent Application Publication WO 1999/051643, which are incorporated herein by reference. Any form of PE in which immunogenicity has been eliminated or reduced can be used in combination with any of the inventive substitution(s) for one or more amino acid residues within one or more T-cell and/or B-cell epitopes described herein so long as it remains capable of cytotoxicity to targeted cells, e.g., by translocation and EF-2 ribosylation in a targeted cell.
[0084] An embodiment of the invention provides a Pseudomonas exotoxin A (PE) comprising a PE amino acid sequence wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted, wherein the PE optionally has:
[0085] (i) a further substitution of one or more amino acid residues within one or more B cell epitopes, and the further substitution for an amino acid within one or more B-cell epitopes is a substitution of, independently, one or more of amino acid residues D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, Q592, and L597 as defined by reference to SEQ ID NO: 1,
[0086] (ii) a further substitution of one or more amino acid residues within one or more T-cell epitopes,
[0087] (iii) a deletion of one or more continuous amino acid residues of residues 1-273 and 285-394 as defined by SEQ ID NO: 1, or
[0088] (iv) a combination of any one, two, or three of (i)-(iii).
[0089] It has been discovered that amino acid residues F443, L477, R494, and L552 are located within one or more T-cell epitopes of PE. Thus, a substitution of one or more of amino acid residues F443, L477, R494, and L552 may, advantageously, remove one or more T cell epitope(s). Accordingly, the inventive PEs may, advantageously, be less immunogenic than an unsubstituted (e.g., wild-type) PE.
[0090] A preferred embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0091] wherein:
[0092] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3
[0093] R.sup.1=1 to 10 amino acid residues
[0094] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0095] R.sup.2=1 to 10 amino acid residues;
[0096] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0097] PE functional domain III=residues 395-613 of SEQ ID NO: 1, wherein one or more of amino acid residues F443, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently substituted;
[0098] and the PE comprises optionally a further substitution of an amino acid within one or more B-cell epitopes.
[0099] The substitution of one or more of amino acid residues F443, L477, R494, and L552 may be a substitution of any amino acid residue for one or more of amino acid residues F443, L477, R494, and L552. In an embodiment of the invention, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of, independently, alanine, glutamic acid, histidine, or asparagine in place of one or more of amino acid residues F443, L477, R494, and L552. In an embodiment of the invention, the substitution of L552 is a substitution of glutamic acid or asparagine in place of L552 and the substitution of L477 is a substitution of histidine in place of L477.
[0100] In an embodiment of the invention, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid or asparagine in place of amino acid residue L552.
[0101] In addition to the substitution(s) for one or more amino acid residues within one or more PE T-cell epitopes described herein, the inventive PE may, optionally, also include additional substitution(s) for one or more amino acid residues within one or more B-cell epitopes of SEQ ID NO: 1. In this regard, in an embodiment of the invention, the PE has a substitution of one or more amino acids within one or more B-cell epitopes of SEQ ID NO: 1. In a preferred embodiment of the invention, the substitution of one or more amino acid residues within one or more B-cell epitopes of SEQ ID NO: 1 includes a substitution of alanine, glycine, serine, or glutamine for one or more amino acids within one or more B-cell epitopes of SEQ ID NO: 1. The substitution(s) within one or more B-cell epitopes may, advantageously, further reduce immunogenicity by the removal of one or more B-cell epitopes. The substitution(s) may be located within any suitable PE B-cell epitope. Exemplary B-cell epitopes are disclosed in, for example, International Patent Application Publications WO 2007/016150, WO 2009/032954, and WO 2011/032022, each of which is incorporated herein by reference. In a preferred embodiment, the substitution of one or more amino acids within one or more B-cell epitopes of SEQ ID NO: 1 is a substitution of alanine, glycine, serine, or glutamine, independently, in place of one or more of amino acid residues E282, E285, P290, R313, N314, P319, D324, E327, E331, Q332, D403, D406, R412, R427, E431, R432, D461, D463, R467, Y481, R490, R505, R513, L516, E522, R538, E548, R551, R576, K590, Q592, and L597, wherein the amino acid residues E282, E285, P290, R313, N314, P319, D324, E327, E331, Q332, D403, D406, R412, R427, E431, R432, D461, D463, R467, Y481, R490, R505, R513, L516, E522, R538, E548, R551, R576, K590, Q592, and L597 are defined by reference to SEQ ID NO: 1.
[0102] In an embodiment of the invention, the further substitution of an amino acid within one or more B-cell epitopes is a substitution of, independently, alanine, glycine, serine, or glutamine in place of one or more of amino acid residues E282, E285, P290, R313, N314, P319, D324, E327, E331, Q332, D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, K590, Q592, and L597, as defined by reference to SEQ ID NO: 1. Preferably, the further substitution of an amino acid within one or more B-cell epitopes is a substitution of, independently, alanine, glycine, or serine in place of one or more of amino acid residues R427, R505, and R551. In an especially preferred embodiment, the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443, a substitution of histidine in place of L477, a substitution of alanine in place of R494, and a substitution of glutamic acid in place of L552, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; and (b) a substitution of alanine for amino acid residue R505, as defined by reference to SEQ ID NO: 1.
[0103] In an embodiment of the invention, any of the PEs described herein may have an arginine at position 458, with reference to SEQ ID NO: 1. Without being bound to a particular theory or mechanism, it is believed that an arginine at position 458 provides enhanced cytotoxicity.
[0104] In an embodiment of the invention, the PE has an arginine residue at position 458, as defined by reference to SEQ ID NO: 1. In a preferred embodiment, the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue R456, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; and (e) a substitution of alanine for amino acid residue R505; (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
[0105] In an embodiment of the invention, the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of alanine in place of amino acid residue R456; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
[0106] In an embodiment of the invention, the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue R456, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
[0107] In another embodiment of the invention, the substitution of one or more of amino acid residues F443, R456, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of alanine in place of amino acid residue R456; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of asparagine in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1.
[0108] In a preferred embodiment of the invention, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R456; (c) a substitution of alanine for amino acid residue R463; (d) a substitution of alanine for amino acid residue R467; (e) a substitution of alanine for amino acid residue R490; (f) a substitution of alanine for amino acid residue R505; and (g) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. A preferred embodiment is a PE comprising SEQ ID NO: 285 (T14-L010R+456A).
[0109] In a preferred embodiment of the invention, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of asparagine in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R456; (c) a substitution of alanine for amino acid residue R463; (d) a substitution of alanine for amino acid residue R467; (e) a substitution of alanine for amino acid residue R490; (f) a substitution of alanine for amino acid residue R505; and (g) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. A preferred embodiment is a PE comprising SEQ ID NO: 286 (T15-L010R+456A).
[0110] In a preferred embodiment of the invention, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. A preferred embodiment is a PE comprising SEQ ID NO: 287 (T14-L010R).
[0111] In a preferred embodiment of the invention, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of asparagine in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. A preferred embodiment is a PE comprising SEQ ID NO: 288 (T15-L010R).
[0112] In a preferred embodiment, the FCS is a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)).
[0113] In a preferred embodiment,
[0114] n=1 for R.sup.1 and R.sup.2,
[0115] R.sup.1=a linker of the amino acid sequence of SEQ ID NO: 282 (DKTHKASGG),
[0116] R.sup.2=a linker of the amino acid sequence of SEQ ID NO: 284 (GGGGGS), and
[0117] FCS=furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)). In an especially preferred embodiment, n is 0 for R.sup.3.
[0118] In an embodiment of the invention, the PE has the further substitution of an amino acid within one or more T-cell epitopes. In this regard, the PE may comprise an amino acid sequence having a further substitution of any amino acid in place of one or more amino acid residues at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 as defined by reference to SEQ ID NO: 1. In an embodiment of the invention, the further substitution of any amino acid in place of one or more amino acid residues at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 is a substitution of one or more amino acid residues at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, Y470, 1471, A472, P475, A476, L477, 1493, R494, N495, L498, L499, R500, V501, Y502, V503, R505, L508, P509, R551, L552, T554, 1555, L556, and W558.
[0119] The substitution of one or more amino acid residues at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 of SEQ ID NO: 1 may be a substitution of any amino acid residue in place of an amino acid residue at any one or more of positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 of SEQ ID NO: 1. The substitution of one or more amino acid residues at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 of SEQ ID NO: 1 may include, e.g., a substitution of alanine, glycine, serine, or glutamine in place of one or more amino acid residues at position 421, 422, 423, 425, 427, 429, 439, 440, 443, 444, 446, 447, 450, 463, 464, 465, 466, 467, 468, 469, 470, 471, 472, 473, 474, 475, 476, 477, 478, 479, 480, 481, 482, 483, 484, 485, 486, 487, 488, 489, 490, 491, 492, 493, 494, 495, 496, 497, 498, 499, 500, 501, 502, 503, 504, 505, 506, 507, 508, 509, 510, 511, 512, 513, 514, 515, 516, 517, 518, 519, 551, 552, 554, 555, 556, and 558 of SEQ ID NO: 1. In a preferred embodiment, the substitution of one or more amino acid residues at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 of SEQ ID NO: 1 is a substitution of alanine, glycine, serine, or glutamine in place of one or more of amino acid residues R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, Y470, 1471, A472, P475, A476, L477, 1493, R494, N495, L498, L499, R500, V501, Y502, V503, R505, L508, P509, R551, L552, T554, 1555, L556, and W558. One or more substitutions in one or more T cell epitopes located at positions R421, L422, L423, A425, R427, L429, Y439, H440, F443, L444, A446, A447, 1450, 463-519, R551, L552, T554, 1555, L556, and W558 of PE as defined by reference to SEQ ID NO: 1 may further reduce immunogenicity of PE. In an embodiment, the amino acid sequence does not have a substitution of one or more amino acid residues at positions 427, 467, 485, 490, 505, 513, 516, and 551.
[0120] Preferably, the PE comprises one or more substitutions that increase cytoxicity as disclosed, for example, in International Patent Application Publication WO 2007/016150, which is incorporated herein by reference. In this regard, an embodiment of the invention provides PE with a substitution of an amino acid within one or more B-cell epitopes of SEQ ID NO: 1 and the substitution of an amino acid within one or more B-cell epitopes of SEQ ID NO: 1 is a substitution of valine, leucine, or isoleucine in place of amino acid residue R490, wherein the amino acid residue R490 is defined by reference to SEQ ID NO: 1. In an embodiment of the invention, substitution of one or more amino acid residues at positions 313, 327, 331, 332, 431, 432, 505, 516, 538, and 590 defined by reference to SEQ ID NO: 1 with alanine or glutamine may provide a PE with an increased cytotoxicity as disclosed, for example, in International Patent Application Publication WO 2007/016150, which is incorporated herein by reference. Increased cytotoxic activity and decreased immunogenicity can occur simultaneously, and are not mutually exclusive. Substitutions that both increase cytotoxic activity and decrease immunogenicity, such as substitutions of R490 to glycine or, more preferably, alanine, are especially preferred.
[0121] In an embodiment of the invention, another embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0122] wherein:
[0123] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3,
[0124] R.sup.1=1 to 10 amino acid residues
[0125] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0126] R.sup.2=1 to 10 amino acid residues;
[0127] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0128] PE functional domain III=residues 395-613 of SEQ ID NO:1, wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted, wherein the PE optionally has:
[0129] (i) a further substitution of one or more amino acid residues within one or more B cell epitopes, and the further substitution for an amino acid within one or more B-cell epitopes is a substitution of, independently, one or more of amino acid residues D403, D406, R412, R427, E431, R432, D461, R463, R467, R490, R505, R513, E522, R538, E548, R551, R576, Q592, and L597 as defined by reference to SEQ ID NO: 1,
[0130] (ii) a further substitution of one or more amino acid residues within one or more T-cell epitopes, or
[0131] (iii) both (i) and (ii).
[0132] Another embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0133] wherein:
[0134] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3,
[0135] R.sup.1=1 to 10 amino acid residues
[0136] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0137] R.sup.2=1 to 10 amino acid residues;
[0138] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0139] PE functional domain III=residues 395-613 of SEQ ID NO:1, wherein the PE includes an arginine residue at position 458, as defined by reference to SEQ ID NO: 1, and
[0140] wherein the PE has:
[0141] (a) a substitution of alanine for amino acid residue R427;
[0142] (b) a substitution of alanine for amino acid residue R463;
[0143] (c) a substitution of alanine for amino acid residue R467;
[0144] (d) a substitution of alanine for amino acid residue R490;
[0145] (e) a substitution of alanine for amino acid residue R505; and
[0146] (f) a substitution of alanine for amino acid residue R538.
[0147] In an embodiment of the invention, n is 0 for R.sup.1 and R.sup.2 of Formula I. In another embodiment of the invention, n is 1 for R.sup.1 and R.sup.2. In an embodiment of the invention, when n is 0 for R.sup.1 and R.sup.2, the PE of Formula I may further comprise a GGS (SEQ ID NO: 283) linking peptide between the furin cleavage sequence (FCS) and PE functional domain III.
[0148] Without being bound by a particular theory or mechanism, it is believed that PEs containing the FCS undergo proteolytic processing inside target cells, thereby activating the cytotoxic activity of the toxin. The FCS of the inventive PEs may comprise any suitable furin cleavage sequence of amino acid residues, which sequence is cleavable by furin. Exemplary furin cleavage sequences are described in Duckert et al., Protein Engineering, Design & Selection, 17(1): 107-112 (2004) and International Patent Application Publication WO 2009/032954, each of which is incorporated herein by reference. In an embodiment of the invention, FCS comprises residues 274-284 of SEQ ID NO: 1 (i.e., RHRQPRGWEQL (SEQ ID NO: 8)), wherein the substitution of an amino acid within one or more B-cell epitopes of SEQ ID NO: 1 is a substitution of alanine, glycine, serine, or glutamine for amino acid residue E282 of SEQ ID NO: 1. Other suitable FCS amino acid sequences include, but are not limited to: R-X.sub.1-X.sub.2-R, wherein X.sub.1 is any naturally occurring amino acid and X.sub.2 is any naturally occurring amino acid (SEQ ID NO: 9), RKKR (SEQ ID NO: 10), RRRR (SEQ ID NO: 11), RKAR (SEQ ID NO: 12), SRVARS (SEQ ID NO: 13), TSSRKRRFW (SEQ ID NO: 14), ASRRKARSW (SEQ ID NO: 15), RRVKKRFW (SEQ ID NO: 16), RNVVRRDW (SEQ ID NO: 17), TRAVRRRSW (SEQ ID NO: 18), RQPR (SEQ ID NO: 19), RHRQPRGW (SEQ ID NO: 20), RHRQPRGWE (SEQ ID NO: 21), HRQPRGWEQ (SEQ ID NO: 22), RQPRGWE (SEQ ID NO: 23), RHRSKRGWEQL (SEQ ID NO: 24), RSKR (SEQ ID NO: 25), RHRSKRGW (SEQ ID NO: 26), HRSKRGWE (SEQ ID NO: 27), RSKRGWEQL (SEQ ID NO: 28), HRSKRGWEQL (SEQ ID NO: 29), RHRSKR (SEQ ID NO: 30), and R-X.sub.1-X.sub.2-R, wherein X.sub.1 is any naturally occurring amino acid and X.sub.2 is arginine or lysine (SEQ ID NO: 4).
[0149] In still another embodiment of the invention, PE functional domain III comprises residues 395-613 of SEQ ID NO: 1, wherein one or more of amino acid residues F443, R456, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently, substituted. Amino acid residues F443, R456, L477, R494, and L552 may be substituted as described herein with respect to other aspects of the invention.
[0150] In an embodiment of the invention, the FCS is represented by the formula P4-P3-P2-P1, wherein P4 is an amino acid residue at the amino end, P1 is an amino acid residue at the carboxyl end, P1 is an arginine or a lysine residue, and the sequence is cleavable at the carboxyl end of P1 by furin.
[0151] In another embodiment of the invention, the FCS (i) further comprises amino acid residues represented by P6-P5 at the amino end, (ii) further comprises amino acid residues represented by P1'-P2' at the carboxyl end, (iii) wherein if P is an arginine or a lysine residue, P2' is tryptophan, and P4 is arginine, valine or lysine, provided that if P4 is not arginine, then P6 and P2 are basic residues, and (iv) the sequence is cleavable at the carboxyl end of P1 by furin.
[0152] In still another embodiment of the invention, the PE functional domain III consists of the sequence of residues 395 to 613 of SEQ ID NO: 1.
[0153] In still another embodiment of the invention, the mutated PE comprises one or more contiguous residues of residues 365-394 of SEQ ID NO: 1 between the FCS and the PE domain III.
[0154] Aspects for the development of Pseudomonas exotoxin chimeric molecules as anti-cancer agents include their cytotoxicity towards tumor cells, their immunogenicity towards human B-cells and human T-cells, and their thermal stability. Thermal stability may be useful for the development of pharmaceutical formulations or compositions.
[0155] Therefore, in one aspect of the invention, it has been discovered that by introducing the mutation R456A instead of R458A, it may be possible to remove all B-cell epitopes from a PE without substantially reducing their cytotoxicity towards tumor cells (in case no further T-cell epitopes are removed by further substitutions). Thus, for the first time, a PE is provided in which all B-cell epitopes have been fully removed and which retains its cytotoxic activity.
[0156] An embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
wherein:
[0157] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3
[0158] R.sup.1=1 to 10 amino acid residues
[0159] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0160] R.sup.2=1 to 10 amino acid residues;
[0161] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0162] PE functional domain III=residues 395-613 of SEQ ID NO: 1, wherein the PE includes an arginine at position 458, as defined by reference to SEQ ID NO: 1, and wherein the PE has:
[0163] (a) a substitution of alanine for amino acid residue R427;
[0164] (b) a substitution of alanine for amino acid residue R463;
[0165] (c) a substitution of alanine for amino acid residue R467;
[0166] (d) a substitution of alanine for amino acid residue R490;
[0167] (e) a substitution of alanine for amino acid residue R505;
[0168] (f) a substitution of alanine for amino acid residue R538; and
[0169] (g) a substitution of alanine for amino acid residue R456.
[0170] In a preferred embodiment, the FCS=a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)).
[0171] In a preferred embodiment,
[0172] n=1 for R.sup.1 and R.sup.2,
[0173] R.sup.1=a linker of the amino acid sequence of SEQ ID NO: 282 (DKTHKASGG),
[0174] R.sup.2=a linker of the amino acid sequence of SEQ ID NO: 284 (GGGGGS), and
[0175] FCS=furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)). In an especially preferred embodiment, n is 0 for R.sup.3.
[0176] In a preferred embodiment, PE functional domain III comprises the amino acid sequence of SEQ ID NO: 37.
[0177] An embodiment of the invention provides an isolated, mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0178] wherein:
[0179] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3
[0180] R.sup.1=1 to 10 amino acid residues
[0181] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0182] R.sup.2=1 to 10 amino acid residues;
[0183] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0184] PE functional domain III=residues 395-613 of SEQ ID NO: 1, wherein the PE includes an arginine at position 458, as defined by reference to SEQ ID NO: 1.
[0185] The inventive PE may be less immunogenic than an unsubstituted PE in accordance with the invention if the immune response to the inventive PE is diminished, quantitatively or qualitatively, as compared to the immune response to an unsubstituted PE. A quantitative decrease in immunogenicity encompasses a decrease in the magnitude or degree of the immune response. The magnitude or degree of immunogenicity can be measured on the basis of any number of known parameters, such as a decrease in the level of cytokine (e.g., antigen-specific cytokine) production (cytokine concentration), a decrease in the number of lymphocytes activated (e.g., proliferation of lymphocytes (e.g., antigen-specific lymphocytes)) or recruited, and/or a decrease in the production of antibodies (antigen-specific antibodies), etc. A qualitative decrease in immunogenicity encompasses any change in the nature of the immune response that renders the immune response less effective at mediating the reduction of the cytotoxic activity of the PE. Methods of measuring immunogenicity are known in the art. For example, measuring the types and levels of cytokines produced can measure immunogenicity. Alternatively or additionally, measuring the binding of PE to antibodies (e.g., antibodies previously exposed to PE) and/or measuring the ability of the PE to induce antibodies when administered to a mammal (e.g., humans, mice, and/or mice in which the mouse immune system is replaced with a human immune system) can measure immunogenicity. A less immunogenic PE may be characterized by a decrease in the production of cytokines such as any one or more of IFN-.gamma., TNF-.alpha., and granzyme B, and/or a reduced stimulation of a cell-mediated immune response, such as a decrease in the proliferation and activation of T-cells and/or macrophages specific for PE as compared to that obtained with an unsubstituted PE. Alternatively or additionally, less immunogenic PE may be characterized by an increase in the production of TGF-beta and/or IL-10 as compared to that obtained with an unsubstituted PE. In a preferred embodiment, reduced immunogenicity is characterized by any one or more of a decrease in T cell stimulation, a decrease in T cell proliferation, and a decrease in T cell IFN.gamma. and/or granzyme B secretion. Alternatively or additionally, a less immunogenic PE may be characterized by a decrease in the stimulation and/or activation of B-cells specific for PE as compared to that obtained with an unsubstituted PE. For example, less immunogenic PE may be characterized by a decrease in the differentiation of B cells into antibody-secreting plasma cells and/or memory cells as compared to that obtained with an unsubstituted PE. Reduced immunogenicity may be characterized by any one or more of a decrease in B cell stimulation, a decrease in B cell proliferation, and a decrease in anti-PE antibody secretion. Qualitative and quantitative diminishment of immunogenicity can occur simultaneously and are not mutually exclusive.
[0186] One of ordinary skill in the art will readily appreciate that the inventive PEs can be modified in any number of ways, such that the therapeutic or prophylactic efficacy of the inventive PEs is increased through the modification. For instance, the inventive PEs can be conjugated or fused either directly or indirectly through a linker to a targeting moiety. In this regard, an embodiment of the invention provides a chimeric molecule comprising (a) a targeting moiety conjugated or fused to (b) any of the inventive PEs described herein. The practice of conjugating compounds, e.g., inventive PEs, to targeting moieties is known in the art. See, for instance, Wadwa et al., J. Drug Targeting, 3: 111 (1995), and U.S. Pat. No. 5,087,616. In an embodiment, any of the inventive PEs described herein may lack a targeting moiety. In an embodiment, any of the inventive PEs described herein may have any of the targeting moieties having any of the sequences described herein.
[0187] The term "targeting moiety" as used herein, refers to any molecule or agent that specifically recognizes and binds to a cell-surface marker, such that the targeting moiety directs the delivery of the inventive PE to a population of cells on which surface the receptor is expressed. Targeting moieties include, but are not limited to, antibodies (e.g., monoclonal antibodies), or fragments thereof, peptides, hormones, growth factors, cytokines, and any other natural or non-natural ligands such as, e.g., scaffold antigen binding proteins.
[0188] Scaffold antigen binding proteins are known in the art. For example, fibronectin and designed ankyrin-repeat proteins (DARPins) have been used as alternative scaffolds for antigen-binding domains, see, e.g., Gebauer and Skerra, Curr. Opin. Chem. Biol., 13:245-255 (2009) and Stumpp et al., Drug Discov. Today, 13:695-701 (2008), both of which are incorporated herein by reference in their entirety.
[0189] In an embodiment, a scaffold antigen binding protein is selected from the group consisting of CTLA-4 (Evibody); lipocalin; Protein A-derived molecules such as the Z-domain of Protein A (Affibody, SpA), A-domain (Avimer/Maxibody); heat shock proteins such as GroE1 and GroES; transferrin (trans-body); ankyrin repeat protein (DARPin); peptide aptamer; C-type lectin domain (Tetranectin); human .gamma.-crystallin and human ubiquitin (affilins); PDZ domains; scorpion toxinkunitz-type domains of human protease inhibitors; and fibronectin (adnectin), which has been subjected to protein engineering in order to obtain binding to a ligand other than the natural ligand.
[0190] CTLA-4 (Cytotoxic T Lymphocyte-associated Antigen 4) is a CD28-family receptor expressed mainly by CD4+ T-cells. Its extracellular domain has a variable domain-like Ig fold. Loops corresponding to CDRs of antibodies can be substituted with a heterologous sequence to confer different binding properties. CTLA-4 molecules engineered to have different binding specificities are also referred to as Evibodies. For further details, see J Immunol. Methods, 248(1-2): 31-45 (2001).
[0191] Lipocalins are a family of extracellular proteins which transport small hydrophobic molecules such as steroids, bilins, retinoids and lipids. They have a rigid beta-sheet secondary structure with a number of loops at the open end of the conical structure which can be engineered to bind to different target antigens. Anticalins are between 160-180 amino acids in size, and are derived from lipocalins. For further details, see Biochim. Biophys. Acta., 1482: 337-350 (2000), U.S. Pat. No. 7,250,297B1 and US20070224633.
[0192] An affibody is a scaffold derived from Protein A of Staphylococcus aureus which can be engineered to bind to an antigen. The domain includes a three-helical bundle of approximately 58 amino acids. Libraries have been generated by the randomization of surface residues. For further details, see Protein Eng. Des. Sel., 17: 455-462 (2004) and EP 1641818A1. Avimers are multidomain proteins derived from the A-domain scaffold family. The native domains of approximately 35 amino acids adopt a defined disulphide bonded structure. Diversity is generated by shuffling the natural variation exhibited by the family of A-domains. For further details, see Nature Biotechnology, 23(12): 1556-1561 (2005) and Expert Opinion on Investigational Drugs, 16(6): 909-917 (2007).
[0193] A transferrin is a monomeric serum transport glycoprotein. Transferrins can be engineered to bind to different target antigens by the insertion of peptide sequences in a permissive surface loop. Examples of engineered transferrin scaffolds include the Trans-body. For further details, see J. Biol. Chem., 274: 24066-24073 (1999).
[0194] Designed Ankyrin Repeat Proteins (DARPins) are derived from Ankyrin which is a family of proteins that mediate attachment of integral membrane proteins to the cytoskeleton. A single ankyrin repeat is a 33 residue motif consisting of two alpha-helices and a beta-turn. They can be engineered to bind to different target antigens by randomizing residues in the first alpha-helix and a beta-turn of each repeat. Their binding interface can be increased by increasing the number of modules (a method of affinity maturation). For further details, see J. Mol Biol., 332: 489-503 (2003); PNAS, 100(4): 1700-1705 (2003); J. Mol. Biol., 369, 1015-1028 (2007); and U.S. Patent Application Publication 20040132028A1.
[0195] Fibronectin is a scaffold which can be engineered to bind to antigen. Adnectins contain a backbone of the natural amino acid sequence of the 10th domain of the 15 repeating units of human fibronectin type III (FN3). Three loops at one end of the beta-sandwich can be engineered to enable an Adnectin to specifically recognize a therapeutic target of interest. For further details see Protein Eng. Des. Sel., 18: 435-444 (2005), U.S. Patent Application Publication 20080139791, International Patent Application Publication WO 2005056764 and U.S. Pat. No. 6,818,418B1.
[0196] Peptide aptamers are combinatorial recognition molecules that include a constant scaffold protein, typically thioredoxin (TrxA), which contains a constrained variable peptide loop inserted at the active site. For further details, see Expert Opin. Biol. Ther., 5: 783-797 (2005).
[0197] Microbodies are derived from naturally occurring microproteins of 25-50 amino acids in length which contain 3-4 cysteine bridges. Examples of microproteins include KalataBI, conotoxin, and knottins. The microproteins have a loop which can be engineered to include up to 25 amino acids without affecting the overall fold of the microprotein. For further details of engineered knottin domains, see International Patent Application Publication WO 2008098796.
[0198] Other antigen binding proteins include proteins which have been used as a scaffold to engineer different target antigen binding properties, including human gamma-crystallin and human ubiquitin (affilins), kunitz type domains of human protease inhibitors, PDZ-domains of the Ras-binding protein AF-6, scorpion toxins (charybdotoxin), and C-type lectin domain (tetranectins). See Chapter 7--Non-Antibody Scaffolds from Handbook of Therapeutic Antibodies (2007, edited by Stefan Dubel) and Protein Science, 15:14-27 (2006). Epitope binding domains of the present invention could be derived from any of these alternative protein domains.
[0199] The term "antibody," as used herein, refers to whole (also known as "intact") antibodies or antigen binding portions thereof that retain antigen recognition and binding capability. The antibody or antigen binding portions thereof can be a naturally-occurring antibody or antigen binding portion thereof, e.g., an antibody or antigen binding portion thereof isolated and/or purified from a mammal, e.g., mouse, rabbit, goat, horse, chicken, hamster, human, etc. The antibody or antigen binding portion thereof can be in monomeric or polymeric form. Also, the antibody or antigen binding portion thereof can have any level of affinity or avidity for the cell surface marker. Desirably, the antibody or antigen binding portion thereof is specific for the cell surface marker, such that there is minimal cross-reaction with other peptides or proteins.
[0200] The antibody may be monoclonal or polyclonal and of any isotype, e.g., IgM, IgG (e.g. IgG, IgG2, IgG3 or IgG4), IgD, IgA or IgE. Complementarity determining regions (CDRs) of an antibody or single chain variable fragments (Fvs) of an antibody against a target cell surface marker can be grafted or engineered into an antibody of choice to confer specificity for the target cell surface marker upon that antibody. For example, the CDRs of an antibody against a target cell surface marker can be grafted onto a human antibody framework of a known three dimensional structure (see, e.g., International Patent Application Publications WO 1998/045322 and WO 1987/002671; U.S. Pat. Nos. 5,859,205; 5,585,089; and 4,816,567; European Patent Application Publication 0173494; Jones et al., Nature, 321:522 (1986); Verhoeyen et al., Science, 239: 1534 (1988), Riechmann et al., Nature, 332:323 (1988); and Winter & Milstein, Nature, 349: 293 (1991)) to form an antibody that may raise little or no immunogenic response when administered to a human. In a preferred embodiment, the targeting moiety is a monoclonal antibody or an antigen binding portion of the monoclonal antibody.
[0201] The antigen binding portion can be any portion that has at least one antigen binding site, such as, e.g., the variable regions or CDRs of the intact antibody. Examples of antigen binding portions of antibodies include, but are not limited to, a heavy chain, a light chain, a variable or constant region of a heavy or light chain, a single chain variable fragment (scFv), or an Fe, Fab, Fab', Fv, or F(ab).sub.2' fragment; single domain antibodies (see, e.g., Wesolowski, Med Microbiol Immunol., 198(3): 157-74 (2009); Saerens et al., Curr. Opin. Pharmacol., 8(5):600-8 (2008); Harmsen and de Haard, Appl. Microbiol. Biotechnol., 77(1): 13-22 (2007), helix-stabilized antibodies (see, e.g., Arndt et al., J. Mol. Biol., 312: 221-228 (2001); triabodies; diabodies (European Patent Application Publication 0404097; International Patent Application Publication WO 1993/011161; and Hollinger et al., Proc. Natl. Acad. Sci. USA, 90: 6444-6448 (1993)); single-chain antibody molecules ("scFvs," see, e.g., U.S. Pat. No. 5,888,773); disulfide stabilized antibodies ("dsFvs," see, e.g., U.S. Pat. Nos. 5,747,654 and 6,558,672), and domain antibodies ("dAbs," see, e.g., Holt et al., Trends Biotech, 21(11):484-490 (2003), Ghahroudi et al., FEBSLett., 414:521-526 (1997), Lauwereys et al., EMBO J 17:3512-3520 (1998), Reiter et al., J. Mol. Biol. 290:685-698 (1999); and Davies and Riechmann, Biotechnology, 13:475-479 (2001)).
[0202] Methods of testing antibodies or antigen binding portions thereof for the ability to bind to any cell surface marker are known in the art and include any antibody-antigen binding assay, such as, for example, radioimmunoassay (RIA), ELISA, Western blot, immunoprecipitation, and competitive inhibition assays (see, e.g., Janeway et al., infra, and U.S. Patent Application Publication 2002/0197266 A1).
[0203] Suitable methods of making antibodies are known in the art. For instance, standard hybridoma methods are described in, e.g., Kohler and Milstein, Eur. J. Immunol., 5, 511-519 (1976), Harlow and Lane (eds.), Antibodies: A Laboratory Manual, CSH Press (1988), and C. A. Janeway et al. (eds.), Immunobiology, 5.sup.th Ed., Garland Publishing, New York, N.Y. (2001)). Alternatively, other methods, such as EBV-hybridoma methods (Haskard and Archer, J. Immunol. Methods, 74(2), 361-67 (1984), and Roder et al., Methods Enzymol., 121, 140-67 (1986)), and bacteriophage vector expression systems (see, e.g., Huse et al., Science, 246, 1275-81 (1989)) are known in the art. Further, methods of producing antibodies in non-human animals are described in, e.g., U.S. Pat. Nos. 5,545,806, 5,569,825, and 5,714,352, and U.S. Patent Application Publication 2002/0197266 A1.
[0204] Phage display also can be used to generate the antibody that may be used in the chimeric molecules of the invention. In this regard, phage libraries encoding antigen-binding variable (V) domains of antibodies can be generated using standard molecular biology and recombinant DNA techniques (see, e.g., Sambrook et al. (eds.), Molecular Cloning, A Laboratory Manual, 3.sup.rd Edition, Cold Spring Harbor Laboratory Press, New York (2001)). Phage encoding a variable region with the desired specificity are selected for specific binding to the desired antigen, and a complete or partial antibody is reconstituted comprising the selected variable domain. Nucleic acid sequences encoding the reconstituted antibody are introduced into a suitable cell line, such as a myeloma cell used for hybridoma production, such that antibodies having the characteristics of monoclonal antibodies are secreted by the cell (see, e.g., Janeway et al., supra, Huse et al., supra, and U.S. Pat. No. 6,265,150).
[0205] Alternatively, antibodies can be produced by transgenic mice that are transgenic for specific heavy and light chain immunoglobulin genes. Such methods are known in the art and described in, for example U.S. Pat. Nos. 5,545,806 and 5,569,825, and Janeway et al., supra.
[0206] Alternatively, the antibody can be a genetically-engineered antibody, e.g., a humanized antibody or a chimeric antibody. Humanized antibodies advantageously provide a lower risk of side effects and can remain in the circulation longer. Methods for generating humanized antibodies are known in the art and are described in detail in, for example, Janeway et al., supra, U.S. Pat. Nos. 5,225,539, 5,585,089 and 5,693,761, European Patent 0239400 B1, and United Kingdom Patent 2188638. Humanized antibodies can also be generated using the antibody resurfacing technology described in, for example, U.S. Pat. No. 5,639,641 and Pedersen et al., J. Mol. Biol., 235, 959-973 (1994).
[0207] The targeting moiety may specifically bind to any suitable cell surface marker. The choice of a particular targeting moiety and/or cell surface marker may be chosen depending on the particular cell population to be targeted. Cell surface markers are known in the art (see, e.g., Mufson et al., Front. Biosci., 11:337-43 (2006); Frankel et al., Clin. Cancer Res., 6:326-334 (2000); and Kreitman et al., AAPS Journal, 8(3): E532-E551 (2006)) and may be, for example, a protein or a carbohydrate. In an embodiment of the invention, the targeting moiety is a ligand that specifically binds to a receptor on a cell surface. Exemplary ligands include, but are not limited to, vascular endothelial growth factor (VEGF), Fas, TNF-related apoptosis-inducing ligand (TRAIL), a cytokine (e.g., IL-2, IL-15, IL-4, IL-13), a lymphokine, a hormone, and a growth factor (e.g., transforming growth factor (TGFa), neuronal growth factor, epidermal growth factor).
[0208] The cell surface marker can be, for example, a cancer antigen. The term "cancer antigen" as used herein refers to any molecule (e.g., protein, peptide, lipid, carbohydrate, etc.) solely or predominantly expressed or over-expressed by a tumor cell or cancer cell, such that the antigen is associated with the tumor or cancer. The cancer antigen can additionally be expressed by normal, non-tumor, or non-cancerous cells. However, in such cases, the expression of the cancer antigen by normal, non-tumor, or non-cancerous cells is not as robust as the expression by tumor or cancer cells. In this regard, the tumor or cancer cells can over-express the antigen or express the antigen at a significantly higher level, as compared to the expression of the antigen by normal, non-tumor, or non-cancerous cells. Also, the cancer antigen can additionally be expressed by cells of a different state of development or maturation. For instance, the cancer antigen can be additionally expressed by cells of the embryonic or fetal stage, which cells are not normally found in an adult host. Alternatively, the cancer antigen can be additionally expressed by stem cells or precursor cells, which cells are not normally found in an adult host.
[0209] The cancer antigen can be an antigen expressed by any cell of any cancer or tumor, including the cancers and tumors described herein. The cancer antigen may be a cancer antigen of only one type of cancer or tumor, such that the cancer antigen is associated with or characteristic of only one type of cancer or tumor. Alternatively, the cancer antigen may be a cancer antigen (e.g., may be characteristic) of more than one type of cancer or tumor. For example, the cancer antigen may be expressed by both breast and prostate cancer cells and not expressed at all by normal, non-tumor, or non-cancer cells.
[0210] Exemplary cancer antigens to which the targeting moiety may specifically bind include, but are not limited to mucin 1 (MUC1; tumor-associated epithelial mucin), melanoma associated antigen (MAGE), preferentially expressed antigen of melanoma (PRAME), carcinoembryonic antigen (CEA), prostate-specific antigen (PSA), prostate specific membrane antigen (PSMA), granulocyte-macrophage colony-stimulating factor receptor (GM-CSFR), CD56, human epidermal growth factor receptor 2 (HER2/neu) (also known as erbB-2), CD5, CD7, tyrosinase tumor antigen, tyrosinase related protein (TRP)1, TRP2, NY-ESO-1, telomerase, and p53. In a preferred embodiment, the cell surface marker, to which the targeting moiety specifically binds, is selected from the group consisting of cluster of differentiation (CD) 19, CD21, CD22, CD25, CD30, CD33 (sialic acid binding Ig-like lectin 3, myeloid cell surface antigen), CD79b, CD123 (interleukin 3 receptor alpha), transferrin receptor, EGF receptor, mesothelin, cadherin, Lewis Y, Glypican-3, FAP (fibroblast activation protein alpha), PSMA (prostate specific membrane antigen), CA9=CAIX (carbonic anhydrase IX), L CAM (neural cell adhesion molecule L1), Endosialin, HER3 (activated conformation of epidermal growth factor receptor family member 3), Alkl/BMP9 complex (anaplastic lymphoma kinase 1/bone morphogenetic protein 9), TPBG=5T4 (trophoblast glycoprotein), ROR1 (receptor tyrosine kinase-like surface antigen), HER1 (activated conformation of epidermal growth factor receptor), and CLL1 (C-type lectin domain family 12, member A). Mesothelin is expressed in, e.g., ovarian cancer, mesothelioma, non-small cell lung cancer, lung adenocarcinoma, fallopian tube cancer, head and neck cancer, cervical cancer, and pancreatic cancer. CD22 is expressed in, e.g., hairy cell leukemia, chronic lymphocytic leukemia (CLL), prolymphocytic leukemia (PLL), non-Hodgkin's lymphoma, small lymphocytic lymphoma (SLL), and acute lymphatic leukemia (ALL). CD25 is expressed in, e.g., leukemias and lymphomas, including hairy cell leukemia and Hodgkin's lymphoma. Lewis Y antigen is expressed in, e.g., bladder cancer, breast cancer, ovarian cancer, colorectal cancer, esophageal cancer, gastric cancer, lung cancer, and pancreatic cancer. CD33 is expressed in, e.g., acute myeloid leukemia (AML), chronic myelomonocytic leukemia (CML), and myeloproliferative disorders.
[0211] In an embodiment of the invention, the targeting moiety is an antibody that specifically binds to a cancer antigen. Exemplary antibodies that specifically bind to cancer antigens include, but are not limited to, antibodies against the transferrin receptor (e.g., HB21 and variants thereof), antibodies against CD22 (e.g., RFB4 and variants thereof), antibodies against CD25 (e.g., anti-Tac and variants thereof), antibodies against mesothelin (e.g., SS1, MORAb-009, SS, HN1, HN2, MN, MB, and variants thereof) and antibodies against Lewis Y antigen (e.g., B3 and variants thereof). In this regard, the targeting moiety may be an antibody selected from the group consisting of B3, RFB4, SS, SS1, MN, MB, HN1, HN2, HB21, and MORAb-009, and antigen binding portions thereof. Further exemplary targeting moieties suitable for use in the inventive chimeric molecules are disclosed e.g., in U.S. Pat. No. 5,242,824 (anti-transferrin receptor); U.S. Pat. No. 5,846,535 (anti-CD25); U.S. Pat. No. 5,889,157 (anti-Lewis Y); U.S. Pat. No. 5,981,726 (anti-Lewis Y); U.S. Pat. No. 5,990,296 (anti-Lewis Y); U.S. Pat. No. 7,081,518 (anti-mesothelin); U.S. Pat. No. 7,355,012 (anti-CD22 and anti-CD25); U.S. Pat. No. 7,368,110 (anti-mesothelin); U.S. Pat. No. 7,470,775 (anti-CD30); U.S. Pat. No. 7,521,054 (anti-CD25); and U.S. Pat. No. 7,541,034 (anti-CD22); U.S. Patent Application Publication 2007/0189962 (anti-CD22); Frankel et al., Clin. Cancer Res., 6: 326-334 (2000), and Kreitman et al., AAPS Journal, 8(3): E532-E551 (2006), each of which is incorporated herein by reference. In another embodiment, the targeting moiety may include the targeting moiety of immunotoxins known in the art. Exemplary immunotoxins include, but are not limited to, LMB-2 (Anti-Tac(Fv)-PE38), BL22 and HA22 (RFB4(dsFv)-PE38), SS1P (SS1 (dsFv)-PE38), HB21-PE40, and variants thereof. In a preferred embodiment, the targeting moiety is the antigen binding portion of HA22. HA22 comprises a disulfide-linked Fv anti-CD22 antibody fragment conjugated to PE38. HA22 and variants thereof are disclosed in International Patent Application Publications WO 2003/027135 and WO 2009/032954, which are incorporated herein by reference.
[0212] The antigen binding portion of the targeting moiety may comprise a light chain variable region and/or a heavy chain variable region. In an embodiment of the invention, the heavy chain variable region comprises a heavy chain CDR1 region, a heavy chain CDR2 region, and a heavy chain CDR3 region. In this regard, the antigen binding domain may comprise one or more of a heavy chain CDR1 region comprising SEQ ID NO: 49, 65, 81, 97, 113, 129, 145, 161, or 177; a heavy chain CDR2 region comprising SEQ ID NO: 53, 69, 85, 101, 117, 133, 149, 165, or 181; and a heavy chain CDR3 region comprising SEQ ID NO: 57, 73, 89, 105, 121, 137, 153, 169, or 185. Preferably, the heavy chain comprises all of SEQ ID NOs: (a) 49, 53, and 57 (anti-mesothelin heavy chain CDR1-CDR3, respectively); (b) SEQ ID NOs: 65, 69, and 73 (anti-glypican-3 gc33 heavy chain CDR1-CDR3, respectively); (c) SEQ ID NOs: 81, 85, and 89 (anti-glypican 3 ab acidic heavy chain CDR1-CDR3, respectively); (d) SEQ ID NOs: 97, 101, and 105 (anti-Fap heavy chain CDR1-CDR3, respectively); (c) SEQ ID NOs: 113, 117, 121 (anti-PSMA heavy chain CDR1-CDR3, respectively); (f) SEQ ID NOs: 129, 133, and 137 (anti-CAIX heavy chain CDR1-CDR3, respectively); (g) SEQ ID NOs: 145, 149, and 153 (anti-L1CAM-(1) heavy chain CDR1-CDR3, respectively); (h) SEQ ID NOs: 161, 165, 169 (anti-L1CAM (2) heavy chain CDR1-CDR3, respectively); or (i) SEQ ID NOs: 177, 181, and 185 (anti-L1CAM(3) heavy chain CDR1-CDR3, respectively).
[0213] In an embodiment of the invention, the light chain variable region may comprise a light chain CDR1 region, a light chain CDR2 region, and a light chain CDR3 region. In this regard, the antigen binding domain may comprise one or more of a light chain CDR1 region comprising SEQ ID NO: 50, 66, 82, 98, 114, 130, 146, 162, or 178; a light chain CDR2 region comprising SEQ ID NO: 54, 70, 86, 102, 118, 134, 150, 166, or 182; and a light chain CDR3 region comprising SEQ ID NO: 58, 74, 90, 106, 122, 138, 154, 170, or 186. Preferably, the light chain comprises all of (a) SEQ ID NOs: 50, 54, and 58 (anti-mesothelin light chain CDR1-CDR3, respectively); (b) SEQ ID NOs: 66, 70, and 74 (anti-glypican-3 ab gc33 light chain CDR1-CDR-3, respectively); (c) SEQ ID NOs: 82, 86, and 90 (anti-glypican 3 ab acidic light chain CDR1-CDR3, respectively); (d) SEQ ID NOs: 98, 102, and 106 (anti-FAP light chain CDR1-CDR3, respectively); (e) SEQ ID NOs: 114, 118, and 122 (anti-PSMA light chain CDR1-3, respectively); (f) SEQ ID NOs: 130, 134, and 138 (anti-CAIX light chain CDR1-CDR3, respectively); (g) SEQ ID NOs: 146, 150, and 154 (anti-L1CAM-(1) light chain CDR1-CDR3, respectively); (h) SEQ ID NOs: 162, 166, and 170 (anti-L1CAM-(2) light chain CDR1-CDR3, respectively); or (i) SEQ ID NOs: 178, 182, and 186 (anti-L1CAM-(3) light chain CDR1-CDR3, respectively).
[0214] In an especially preferred embodiment, the antigen binding portion comprises the CDR1, CDR2, CDR3 regions of the light chain and the CDR1, CDR2, CDR3 regions of the heavy chain. In this regard, the antigen binding portion comprises (a) SEQ ID NOs: 49, 50, 53, 54, 57, and 58; (b) SEQ ID NOs: 65, 66, 69, 70, 73, and 74; (c) SEQ ID NOs: 81, 82, 85, 86, 89, and 90; (d) SEQ ID NOs: 97, 98, 101, 102, 105, and 106; (e) SEQ ID NOs: 113, 114, 117, 118, 121, and 122; (f) SEQ ID NOs: 129, 130, 133, 134, 137, and 138; (g) SEQ ID NOs: 145, 146, 149, 150, 153, and 154; (h) SEQ ID NOs: 161, 162, 165, 166, 169, and 170; or (i) SEQ ID NOs: 177, 178, 181, 182, 185, and 186.
[0215] In an embodiment, the antigen binding portion comprises framework regions FR1, FR2, FR3, FR4 of the heavy chain and FR1, FR2, FR3, FR4 of the light chain in addition to the CDR regions described above. In this regard, the antigen binding portion may comprise a heavy chain FR1 region comprising SEQ ID NO: 47, 63, 79, 95, 111, 127, 143, 159, or 175; a heavy chain FR2 region comprising SEQ ID NO: 51, 67, 83, 99, 115, 131, 147, 163, or 179; a heavy chain FR3 region comprising SEQ ID NO: 55, 71, 87, 103, 119, 135, 151, 167, 183; and a heavy chain FR4 region comprising SEQ ID NO: 59, 75, 91, 107, 123, 139, 155, 171, or 187. The antigen binding portion may comprise a light chain FR1 region comprising SEQ ID NO: 48, 64, 80, 96, 112, 128, 144, 160, 176; a light chain FR2 region comprising SEQ ID NO: 52, 68, 84, 100, 116, 132, 148, 164, or 180; a FR3 region comprising SEQ ID NO: 56, 72, 88, 104, 120, 136, 152, 168, or 184; and a FR4 region comprising SEQ ID NO: 60, 76, 92, 108, 124, 140, 156, 172, or 188. Preferably, the antigen binding portion comprises the FR1, FR2, FR3, F4, CDR1, CDR2, and CDR3 regions of the light chain and the FR1, FR2, FR3, F4, CDR1, CDR2, and CDR3 regions of the heavy chain. In this regard, the antigen binding portion comprises (a) SEQ ID NOs: 47-60 (anti-mesothelin); (b) SEQ ID NOs: 63-76 (anti-glypican-3 ab gc33); (c) SEQ ID NOs: 79-92 (anti-glypican 3 ab acidic); (d) SEQ ID NOs: 95-108 (anti-FAP); (e) SEQ ID NOs: 111-124 (anti-PSMA); (f) SEQ ID NOs: 127-140 (anti-CAIX); (g) SEQ ID NOs: 143-156 (anti-L1CAM(1)); (h) SEQ ID NOs: 159-172 (anti-L1CAM(2)); or (i) SEQ ID NOs: 175-188 (anti-L1CAM(3)).
[0216] In an embodiment, the light chain variable region of the antigen binding domain may comprise SEQ ID NO: 46, 62, 78, 94, 110, 126, 142, 158, or 174. In an embodiment, the heavy chain variable region of the antigen binding domain may comprise SEQ ID NO: 45, 61, 77, 93, 109, 125, 141, 157, or 173. In a preferred embodiment, the antigen binding domain comprises both (a) SEQ ID NOs: 45 and 46 (anti-mesothelin); (b) SEQ ID NOs: 61 and 62 (anti-glypican-3 ab gc33); (c) SEQ ID NOs: 77 and 78 (anti-gc33 acidic); (d) SEQ ID NOs: 93 and 94 (anti-FAP); (e) SEQ ID NOs: 109 and 110 (anti-PSMA); (f) SEQ ID NOs: 125 and 126 (anti-CAIX); (g) SEQ ID NOs: 141 and 142 (anti-L1CAM(1)); (h) SEQ ID NOs: 157 and 158 (anti-L1CAM(2)); (i) SEQ ID NOs: 173 and 174 (anti-L1CAM(3)); or (j) SEQ ID NOs: 93 and 290 (anti-FAP).
[0217] Another aspect of the invention is a humanized anti-mesothelin antibody comprising the comprising the variable heavy chain domain VH of SEQ ID NO: 45 and the variable light chain domain of SEQ ID NO: 46. The humanized antibody was generated by using mouse anti-mesothelin antibody SS1 as a starting material and generating specific combinations of certain framework regions with the CDR regions of mouse SS1 antibody. The humanized antibody advantageously provides good binding properties, stability, and developability.
[0218] In an embodiment of the invention, the targeting moiety may be humanized. In an embodiment of the invention, the targeting moiety is a humanized SS1 or an antigen binding portion of the humanized SS1. In this regard, the targeting moiety may comprise a light chain comprising SEQ ID NO: 33 (cFp-0199) and a heavy chain comprising SEQ ID NO: 38 (cFp-0200). In another embodiment, the targeting moiety may comprise a light chain comprising SEQ ID NOs: 31 (cFp-0199 humanized variable light chain domain VL) and SEQ ID NO: 32 (cFp-0199 kappa constant light chain domain). In an embodiment, the targeting moiety may comprise a heavy chain comprising SEQ ID NOs: 34 (cFp-0200 humanized variable heavy chain domain VH) and SEQ ID NO: 35 (cFp-0200 constant heavy chain domain CH1). In an embodiment, the targeting moiety comprises all of SEQ ID NOs: 31, 32, 34, and 35.
[0219] In an embodiment of the invention, the chimeric molecule comprises a linker. The term "linker" as used herein, refers to any agent or molecule that connects the inventive PE to the targeting moiety. One of ordinary skill in the art recognizes that sites on the inventive PE, which are not necessary for the function of the inventive PE, are ideal sites for attaching a linker and/or a targeting moiety, provided that the linker and/or targeting moiety, once attached to the inventive PE, do(es) not interfere with the function of the inventive PE, i.e., cytotoxic activity, inhibit growth of a target cell, or to treat or prevent cancer. The linker may be capable of forming covalent bonds to both the PE and the targeting moiety. Suitable linkers are known in the art and include, but are not limited to, straight or branched-chain carbon linkers, heterocyclic carbon linkers, and peptide linkers. Where the PE and the targeting moiety are polypeptides, the linker may be joined to the amino acids through side groups (e.g., through a disulfide linkage to cysteine). Preferably, the linkers will be joined to the alpha carbon of the amino and carboxyl groups of the terminal amino acids.
[0220] In another aspect of the invention, it has been discovered that by introducing specific amino acids at both sides of the furin cleavage sequence (FCS), a more stable chimeric molecule of the Pseudomonas exotoxin with a targeting moiety has been provided. In addition, further potential T-cell epitopes in the linker region have been removed, which may be particularly advantageous when using Fab fragments of an antibody as a targeting moiety. These chimeric molecules provide improved thermodynamic stability compared to chimeric molecules without these elongated linkers and retain their cytotoxicity towards tumor cells. Accordingly, in an embodiment of the invention, the linker is an elongated linker. The elongated linker may comprise SEQ ID NO: 36.
[0221] In a preferred embodiment, R.sup.1.sub.n-FCS-R.sup.2.sub.n=a linker of the amino acid sequence of SEQ ID NO: 36. The furin cleavage site is located between amino acid residues 15 and 16 of SEQ ID NO: 36.
[0222] An embodiment of the invention provides a chimeric molecule comprising (a) a targeting moiety conjugated or fused to (b) any of the PEs described herein.
[0223] An embodiment of the invention provides an isolated chimeric molecule comprising (a) a targeting moiety conjugated or fused to (b) a mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0224] wherein:
[0225] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3
[0226] R.sup.1=1 to 10 amino acid residues
[0227] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0228] R.sup.2=1 to 10 amino acid residues;
[0229] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0230] PE functional domain III=residues 395-613 of SEQ ID NO: 1, wherein the PE includes an arginine at position 458, as defined by reference to SEQ ID NO: 1, and wherein the PE has:
[0231] (a) a substitution of alanine for amino acid residue R427;
[0232] (b) a substitution of alanine for amino acid residue R463;
[0233] (c) a substitution of alanine for amino acid residue R467;
[0234] (d) a substitution of alanine for amino acid residue R490;
[0235] (e) a substitution of alanine for amino acid residue R505;
[0236] (f) a substitution of alanine for amino acid residue R538; and
[0237] (g) a substitution of alanine for amino acid residue R456.
[0238] In a preferred embodiment of the chimeric molecule, the FCS=furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)).
[0239] In a preferred embodiment of the chimeric molecule,
[0240] n=1 for R.sup.1 and R.sup.2,
[0241] R.sup.1=a linker of the amino acid sequence of SEQ ID NO: 282 (DKTHKASGG),
[0242] R.sup.2=a linker of the amino acid sequence of SEQ ID NO: 284 (GGGGGS), and
[0243] FCS=furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)). In an especially preferred embodiment, n is 0 for R.sup.3.
[0244] In a preferred embodiment of the chimeric molecule, R.sup.1.sub.n-FCS-R.sup.2.sub.n=a linker of the amino acid sequence of SEQ ID NO: 36 and the targeting moiety comprises the Fab fragment of an antibody.
[0245] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-mesothelin antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (LO10R-456A). In one preferred embodiment, the antigen binding portion is the Fab fragment of an anti-mesothelin antibody comprising the variable heavy chain domain VH of SEQ ID NO: 45 and the variable light chain domain of SEQ ID NO: 46. In this regard, the chimeric molecule may comprise (a) SEQ ID NOs: 39 and 40; (b) SEQ ID NOs: 41 and 42; or (c) SEQ ID NOs: 43 and 44.
[0246] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 61 and the variable light chain domain of SEQ ID NO: 62.
[0247] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 77 and the variable light chain domain of SEQ ID NO: 78.
[0248] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a FCS corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 93 (SEQ ID NOs: 97, 101, and 105, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 94 (SEQ ID NOs: 98, 102, and 106, respectively). In one preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 94. In another preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain VL of SEQ ID NO: 290. In this regard, the chimeric molecule may comprise (a) SEQ ID NOs: 291 and 293; (b) SEQ ID NOs: 291 and 294; or (c) SEQ ID NOs: 292 and 294.
[0249] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a FCS corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (L010R-456A). In s preferred embodiment, the antigen binding portion is the Fab fragment of an anti-PSMA antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 109 (SEQ ID NOs: 113, 117, and 121, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 110 (SEQ ID NOs: 114, 118, and 122, respectively).
[0250] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a FCS corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-CAIX antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 125 (SEQ ID NOs: 129, 133, and 137, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 126 (SEQ ID NOs: 130, 134, and 138, respectively). In this regard, the chimeric molecule may comprise (a) SEQ ID NOs: 295 and 297 or (b) SEQ ID NOs: 296 and 297.
[0251] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a FCS corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1 CAM antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (L010R-456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 141 (SEQ ID NOs: 145, 149, and 153, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 142 (SEQ ID NOs: 146, 150, and 154, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 157 and the variable light chain domain of SEQ ID NO: 158. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 173 and the variable light chain domain of SEQ ID NO: 174.
[0252] In a preferred embodiment, the invention provides an isolated chimeric molecule comprising (a) a targeting moiety conjugated or fused to (b) a mutated Pseudomonas exotoxin A (PE), comprising a sequence of the following formula:
R.sup.1.sub.n-FCS-R.sup.2.sub.n--R.sup.3.sub.n-PE functional domain III
[0253] wherein:
[0254] n=0 or 1 independently for each of R.sup.1, R.sup.2 and R.sup.3
[0255] R.sup.1=1 to 10 amino acid residues
[0256] FCS=a furin cleavage sequence of amino acid residues, which sequence is cleavable by furin and has an amino end and a carboxyl end,
[0257] R.sup.2=1 to 10 amino acid residues;
[0258] R.sup.3=1 or more contiguous residues of residues 365-394 of SEQ ID NO: 1; and,
[0259] PE functional domain III=residues 395-613 of SEQ ID NO: 1,
[0260] wherein one or more of amino acid residues F443, L477, R494, and L552 as defined by reference to SEQ ID NO: 1 are, independently substituted;
[0261] and the PE comprises optionally a further substitution of an amino acid within one or more B-cell epitopes.
[0262] It has been discovered that a substitution of histidine in place of amino acid residue L477 leads to a strong reduction of T cell responses compared to substitutions with other amino acids (see, for example, Example 4) while at the same time improving the cytotoxic efficacy of the PE compared to substitutions with other amino acids (see, for example, Examples 3, 5 and 6). Therefore, in a preferred embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of histidine in place of amino acid residue L477. The substitution of glutamic acid or asparagine in place of amino acid residue L552 also leads to further improved toxicity while reducing T cell responses. In a preferred embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of histidine in place of amino acid residue L477 and a substitution of glutamic acid or asparagine in place of amino acid residue L552. In an embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid or asparagine in place of amino acid residue L552.
[0263] In a preferred embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427 and (b) a substitution of alanine for amino acid residue R505 as defined by reference to SEQ ID NO: 1. In a preferred embodiment of the invention, the PE of the chimeric molecule comprises the amino acid sequence of SEQ ID NO: 289 (T18/T20).
[0264] In a preferred embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R456; (c) a substitution of alanine for amino acid residue R463; (d) a substitution of alanine for amino acid residue R467; (e) a substitution of alanine for amino acid residue R490; (f) a substitution of alanine for amino acid residue R505; and (g) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. In a preferred embodiment, the PE of the chimeric molecule comprises the amino acid sequence of SEQ ID NO: 285 (T14-L010R+456A).
[0265] In a preferred embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of asparagine in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R456; (c) a substitution of alanine for amino acid residue R463; (d) a substitution of alanine for amino acid residue R467; (e) a substitution of alanine for amino acid residue R490; (f) a substitution of alanine for amino acid residue R505; and (g) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. In a preferred embodiment, the PE of the chimeric molecule comprises the amino acid sequence of SEQ ID NO: 286 (T15-L010R+456A).
[0266] In a preferred embodiment of the invention with respect to the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of glutamic acid in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. In a preferred embodiment, the PE of the chimeric molecule comprises the amino acid sequence of SEQ ID NO: 287 (T14-L010R).
[0267] In a preferred embodiment of the invention within the chimeric molecule, the substitution of one or more of amino acid residues F443, L477, R494, and L552 is a substitution of alanine in place of amino acid residue F443; a substitution of histidine in place of amino acid residue L477; a substitution of alanine in place of amino acid residue R494; and a substitution of asparagine in place of amino acid residue L552, the PE has an arginine residue at position 458, and the further substitution of an amino acid within one or more B-cell epitopes is: (a) a substitution of alanine for amino acid residue R427; (b) a substitution of alanine for amino acid residue R463; (c) a substitution of alanine for amino acid residue R467; (d) a substitution of alanine for amino acid residue R490; (e) a substitution of alanine for amino acid residue R505; and (f) a substitution of alanine for amino acid residue R538; as defined by reference to SEQ ID NO: 1. In a preferred embodiment, the PE of the chimeric molecule comprises the amino acid sequence of SEQ ID NO: 288 (T15-L010R).
[0268] In a preferred embodiment with respect to the chimeric molecule, the FCS=furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)).
[0269] In a preferred embodiment with respect to the chimeric molecule,
[0270] n=1 for R.sup.1 and R.sup.2,
[0271] R.sup.1=a linker of the amino acid sequence of SEQ ID NO: 282 (DKTHKASGG),
[0272] R.sup.2=a linker of the amino acid sequence of SEQ ID NO: 284 (GGGGGS), and
[0273] FCS=furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)). In an especially preferred embodiment, n is 0 for R.sup.3.
[0274] In a preferred embodiment with respect to the chimeric molecule, R.sup.1.sub.n-FCS-R.sup.2.sub.n=a linker of the amino acid sequence of SEQ ID NO: 36 and the targeting moiety comprises the Fab fragment of an antibody.
[0275] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-mesothelin antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-mesothelin antibody comprising the variable heavy chain domain VH of SEQ ID NO: 45 and the variable light chain domain of SEQ ID NO: 46.
[0276] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-mesothelin antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 286 (T15-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-mesothelin antibody comprising the variable heavy chain domain VH of SEQ ID NO: 45 and the variable light chain domain of SEQ ID NO: 46.
[0277] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-mesothelin antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 287 (T14-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-mesothelin antibody comprising the variable heavy chain domain VH of SEQ ID NO: 45 and the variable light chain domain of SEQ ID NO: 46.
[0278] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-mesothelin antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 288 (T15-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-mesothelin antibody comprising the variable heavy chain domain VH of SEQ ID NO: 45 and the variable light chain domain of SEQ ID NO: 46.
[0279] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 61 (SEQ ID NOs: 65, 69, and 73, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 62 (SEQ ID NOs: 66, 70, and 74, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 77 (SEQ ID NO: 81, 85, and 89, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 78 (SEQ ID NOs: 82, 86, and 90, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 61 and the variable light chain domain of SEQ ID NO: 62. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 77 and the variable light chain domain of SEQ ID NO: 78.
[0280] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 285 (T14-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 61 (SEQ ID NO: 65, 69, and 73, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 62 (SEQ ID NOs: 66, 70, and 74, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 77 (SEQ ID NOs: 81, 85, and 89, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 78 (SEQ ID NOs: 82, 86, and 90, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 61 and the variable light chain domain of SEQ ID NO: 62. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 77 and the variable light chain domain of SEQ ID NO: 78.
[0281] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 286 (T15-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 61 (SEQ ID NO: 65, 69, and 73, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 62 (SEQ ID NOs: 66, 70, and 74, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 77 (SEQ ID NOs: 81, 85, and 89, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 78 (SEQ ID NOs: 82, 86, and 90, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 61 and the variable light chain domain of SEQ ID NO: 62. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 77 and the variable light chain domain of SEQ ID NO: 78.
[0282] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 287 (T14-LO-1OR). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 61 (SEQ ID NOs: 65, 69, and 73, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 62 (SEQ ID NOs: 66, 70, and 74, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 77 (SEQ ID NOs: 81, 85, and 89, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 78 (SEQ ID NOs: 82, 86, and 90, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 61 and the variable light chain domain of SEQ ID NO: 62. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 77 and the variable light chain domain of SEQ ID NO: 78.
[0283] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-glypican-3 antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 288 (T15-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 61 (SEQ ID NOs: 65, 69, and 73, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 62 (SEQ ID NOs: 66, 70, and 74, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 77 (SEQ ID NOs: 81, 85, and 89, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 78 (SEQ ID NOs: 82, 86, and 90, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 61 and the variable light chain domain of SEQ ID NO: 62. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-glypican-3 antibody comprising the variable heavy chain domain VH of SEQ ID NO: 77 and the variable light chain domain of SEQ ID NO: 78. In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHIRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 93 (SEQ ID NOs: 97, 101, and 105, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 94 (SEQ ID NOs: 98, 102, and 106, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 94. In another preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 290.
[0284] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 285 (T14-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 93 (SEQ ID NOs: 97, 101, and 105, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 94 (SEQ ID NOs: 98, 102, and 106, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 94. In another preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 290.
[0285] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 286 (T15-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 93 (SEQ ID NOs: 97, 101, and 105, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 94 (SEQ ID NOs: 98, 102, and 106, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 94. In another preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 290.
[0286] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 287 (T14-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 93 (SEQ ID NOs: 97, 101, and 105, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 94 (SEQ ID NOs: 98, 102, and 106). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 94. In another preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 290.
[0287] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-FAP antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 288 (T15-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 93 (SEQ ID NOs: 97, 101, and 105, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 94 (SEQ ID NOs: 98, 102, and 106, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VH of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 94. In another preferred embodiment, the antigen binding portion is the Fab fragment of an anti-FAP antibody comprising the variable heavy chain domain VII of SEQ ID NO: 93 and the variable light chain domain of SEQ ID NO: 290. In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-PSMA antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 109 (SEQ ID NOs: 113, 117, and 121, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 110 (SEQ ID NOs: 114, 118, and 122, respectively).
[0288] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 285 (T14-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-PSMA antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 109 (SEQ ID NOs: 113, 117, and 121, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 110 (SEQ ID NOs: 114, 118, and 122, respectively).
[0289] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 286 (T15-L010R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-PSMA antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 109 (SEQ ID NOs: 113, 117, and 121, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 110 (SEQ ID NOs: 114, 118, and 122, respectively).
[0290] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 287 (T14-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-PSMA antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 109 (SEQ ID NOs: 113, 117, and 121, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 110 (SEQ ID NOs: 114, 118, and 122, respectively).
[0291] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-PSMA antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-PSMA antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 109 (SEQ ID NOs: 113, 117, and 121, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 110 (SEQ ID NOs: 114, 118, and 122, respectively).
[0292] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-CAIX antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 125 (SEQ ID NOs: 129, 133, and 137, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 126 (SEQ ID NOs: 130, 134, and 138).
[0293] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-CAIX antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 125 (SEQ ID NOs: 129, 133, and 137, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 126 (SEQ ID NOs: 130, 134, and 138, respectively).
[0294] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-CAIX antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 125 (SEQ ID NOs: 129, 133, and 137, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 126 (SEQ ID NOs: 130, 134, and 138, respectively).
[0295] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-CAIX antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 125 (SEQ ID NOs: 129, 133, and 137, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 126 (SEQ ID NOs: 130, 134, and 138, respectively).
[0296] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-CAIX antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-CAIX antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 125 (SEQ ID NOs: 129, 133, and 137, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 126 (SEQ ID NOs: 130, 134, and 138, respectively). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 289 (T18/T20). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1 CAM antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and aPE comprising. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-LICAM antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 141 (SEQ ID NOs: 145, 149, and 153, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 142 (SEQ ID NOs: 146, 150, and 154, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1 CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 157 and the variable light chain domain of SEQ ID NO: 158. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1 CAM antibody comprising the variable heavy chain domain VII of SEQ ID NO: 173 and the variable light chain domain of SEQ ID NO: 174.
[0297] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 285 (T14-LO10R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1 CAM antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 141 (SEQ ID NOs: 145, 149, and 153, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 142 (SEQ ID NOs: 146, 150, and 154, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 157 and the variable light chain domain of SEQ ID NO: 158. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1 CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 173 and the variable light chain domain of SEQ ID NO: 174.
[0298] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1 CAM antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 286 (T15-LO10R+456A). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 141 (SEQ ID NOs: 145, 149, and 153, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 142 (SEQ ID NOs: 146, 150, and 154, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 157 and the variable light chain domain of SEQ ID NO: 158. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 173 and the variable light chain domain of SEQ ID NO: 174.
[0299] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 287 (T14-LO10R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 141 (SEQ ID NOs: 145, 149, and 153, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 142 (SEQ ID NOs: 146, 150, and 154, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 157 and the variable light chain domain of SEQ ID NO: 158. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 173 and the variable light chain domain of SEQ ID NO: 174.
[0300] In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a furin cleavage sequence (FCS) corresponding to amino acid residues 274-284 of SEQ ID NO: 1 (RHRQPRGWEQL (SEQ ID NO: 8)) and a PE comprising SEQ ID NO: 288 (T15-LO10R). In a preferred embodiment, the chimeric molecule comprises the antigen binding portion of an anti-L1CAM antibody, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 288 (T15-L010R). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the CDR1, CDR2 and CDR3 of the variable heavy chain domain VH of SEQ ID NO: 141 (SEQ ID NOs: 145, 149, and 153, respectively) and the CDR1, CDR2 and CDR3 of the variable light chain domain of SEQ ID NO: 142 (SEQ ID NOs: 146, 150, and 154, respectively). In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 157 and the variable light chain domain of SEQ ID NO: 158. In a preferred embodiment, the antigen binding portion is the Fab fragment of an anti-L1CAM antibody comprising the variable heavy chain domain VH of SEQ ID NO: 173 and the variable light chain domain of SEQ ID NO: 174. Included in the scope of the invention are functional portions of the inventive PEs and chimeric molecules described herein. The term "functional portion" when used in reference to a PE or chimeric molecule refers to any part or fragment of the PE or chimeric molecule of the invention, which part or fragment retains the biological activity of the PE or chimeric molecule of which it is a part (the parent PE or chimeric molecule). Functional portions encompass, for example, those parts of a PE or chimeric molecule that retain the ability to specifically bind to and destroy or inhibit the growth of target cells or treat or prevent cancer, to a similar extent, the same extent, or to a higher extent, as the parent PE or chimeric molecule. In reference to the parent PE or chimeric molecule, the functional portion can comprise, for instance, about 10% or more, about 25% or more, about 30% or more, about 50% or more, about 68% or more, about 80% or more, about 90% or more, or about 95% or more, of the parent PE or chimeric molecule.
[0301] The functional portion can comprise additional amino acids at the amino or carboxy terminus of the portion, or at both termini, which additional amino acids are not found in the amino acid sequence of the parent PE or chimeric molecule. Desirably, the additional amino acids do not interfere with the biological function of the functional portion, e.g., specifically binding to and destroying or inhibiting the growth of target cells, having the ability to treat or prevent cancer, etc. More desirably, the additional amino acids enhance the biological activity, as compared to the biological activity of the parent PE or chimeric molecule.
[0302] Included in the scope of the invention are functional variants of the inventive PEs and chimeric molecules described herein. The term "functional variant" as used herein refers to a PE or chimeric molecule having substantial or significant sequence identity or similarity to a parent PE or chimeric molecule, which functional variant retains the biological activity of the PE or chimeric molecule of which it is a variant. Functional variants encompass, for example, those variants of the PE or chimeric molecule described herein (the parent PE or chimeric molecule) that retain the ability to specifically bind to and destroy or inhibit the growth of target cells to a similar extent, the same extent, or to a higher extent, as the parent PE or chimeric molecule. In reference to the parent PE or chimeric molecule, the functional variant can, for instance, be about 30% or more, about 50% or more, about 75% or more, about 80% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, or about 99% or more identical in amino acid sequence to the parent PE or chimeric molecule.
[0303] The functional variant can, for example, comprise the amino acid sequence of the parent PE or chimeric molecule with at least one conservative amino acid substitution. Conservative amino acid substitutions are known in the art and include amino acid substitutions in which one amino acid having certain chemical and/or physical properties is exchanged for another amino acid that has the same chemical or physical properties. For instance, the conservative amino acid substitution can be an acidic amino acid substituted for another acidic amino acid (e.g., Asp or Glu), an amino acid with a nonpolar side chain substituted for another amino acid with a nonpolar side chain (e.g., Ala, Gly, Val, Ile, Leu, Met, Phe, Pro, Trp, Val, etc.), a basic amino acid substituted for another basic amino acid (Lys, Arg, etc.), an amino acid with a polar side chain substituted for another amino acid with a polar side chain (Asn, Cys, Gln, Ser, Thr, Tyr, etc.), etc.
[0304] Alternatively or additionally, the functional variants can comprise the amino acid sequence of the parent PE or chimeric molecule with at least one non-conservative amino acid substitution. In this case, it is preferable for the non-conservative amino acid substitution to not interfere with or inhibit the biological activity of the functional variant. Preferably, the non-conservative amino acid substitution enhances the biological activity of the functional variant, such that the biological activity of the functional variant is increased as compared to the parent PE or chimeric molecule.
[0305] The PE or chimeric molecule of the invention can consist essentially of the specified amino acid sequence or sequences described herein, such that other components of the functional variant, e.g., other amino acids, do not materially change the biological activity of the functional variant.
[0306] The PE or chimeric molecule of the invention (including functional portions and functional variants) of the invention can comprise synthetic amino acids in place of one or more naturally-occurring amino acids. Such synthetic amino acids are known in the art and include, for example, aminocyclohexane carboxylic acid, norleucine, .alpha.-amino n-decanoic acid, homoserine, S-acetylaminomethyl-cysteine, trans-3- and trans-4-hydroxyproline, 4-aminophenylalanine, 4-nitrophenylalanine, 4-chlorophenylalanine, 4-carboxyphenylalanine, .beta.-phenylserine .beta.-hydroxyphenylalanine, phenylglycine, .alpha.-naphthylalanine, cyclohexylalanine, cyclohexylglycine, indoline-2-carboxylic acid, 1,2,3,4-tetrahydroisoquinoline-3-carboxylic acid, aminomalonic acid, aminomalonic acid monoamide, N'-benzyl-N'-methyl-lysine, N',N'-dibenzyl-lysine, 6-hydroxylysine, ornithine, .alpha.-aminocyclopentane carboxylic acid, .alpha.-aminocyclohexane carboxylic acid, .alpha.-aminocycloheptane carboxylic acid, .alpha.-(2-amino-2-norbornane)-carboxylic acid, .alpha.,.gamma.-diaminobutyric acid, .alpha.,.beta.-diaminopropionic acid, homophenylalanine, and .alpha.-tert-butylglycine.
[0307] The PE or chimeric molecule of the invention (including functional portions and functional variants) can be glycosylated, amidated, carboxylated, phosphorylated, esterified, N-acylated, cyclized via, e.g., a disulfide bridge, or converted into an acid addition salt and/or optionally dimerized or polymerized, or conjugated.
[0308] An embodiment of the invention provides a method of producing the inventive PE comprising (a) recombinantly expressing the PE and (b) purifying the PE. The PEs and chimeric molecules of the invention (including functional portions and functional variants) can be obtained by methods of producing proteins and polypeptides known in the art. Suitable methods of de novo synthesizing polypeptides and proteins are described in references, such as Chan et al., Fmoc Solid Phase Peptide Synthesis, Oxford University Press, Oxford, United Kingdom, 2005; Peptide and Protein Drug Analysis, ed. Reid, R., Marcel Dekker, Inc., 2000; Epitope Mapping, ed. Westwood et al., Oxford University Press, Oxford, United Kingdom, 2000; and U.S. Pat. No. 5,449,752. Also, the PEs and chimeric molecules of the invention can be recombinantly expressed using the nucleic acids described herein using standard recombinant methods. See, for instance, Sambrook et al., Molecular Cloning: A Laboratory Manual, 3.sup.rd ed., Cold Spring Harbor Press, Cold Spring Harbor, N.Y. 2001; and Ausubel et al., Current Protocols in Molecular Biology, Greene Publishing Associates and John Wiley & Sons, NY, 1994.
[0309] The method further comprises purifying the PE. Once expressed, the inventive PEs may be purified in accordance with purification techniques known in the art. Exemplary purification techniques include, but are not limited to, ammonium sulfate precipitation, affinity columns, and column chromatography, or by procedures described in, e.g., R. Scopes, Protein Purification, Springer-Verlag, NY (1982).
[0310] Another embodiment of the invention provides a method of producing the inventive chimeric molecule comprising (a) recombinantly expressing the chimeric molecule and (b) purifying the chimeric molecule. The chimeric molecule may be recombinantly expressed and purified as described herein with respect to other aspects of the invention. In an embodiment of the invention, recombinantly expressing the chimeric molecule comprises inserting a nucleotide sequence encoding a targeting moiety and a nucleotide sequence encoding a PE into a vector. The method may comprise inserting the nucleotide sequence encoding the targeting moiety and the nucleotide sequence encoding the PE in frame so that it encodes one continuous polypeptide including a functional targeting moiety region and a functional PE region. In an embodiment of the invention, the method comprises ligating a nucleotide sequence encoding the PE to a nucleotide sequence encoding a targeting moiety so that, upon expression, the PE is located at the carboxyl terminus of the targeting moiety. In an alternative embodiment, the method comprises ligating a nucleotide sequence encoding the PE to a nucleotide sequence encoding a targeting moiety so that, upon expression, the PE is located at the amino terminus of the targeting moiety.
[0311] Still another embodiment of the invention provides a method of producing the inventive chimeric molecule comprising (a) recombinantly expressing the inventive PE, (b) purifying the PE, and (c) covalently linking a targeting moiety to the purified PE. The inventive PE may be recombinantly expressed as described herein with respect to other aspects of the invention. The method further comprises covalently linking a targeting moiety to the purified PE. The method of attaching a PE to a targeting moiety may vary according to the chemical structure of the targeting moiety. For example, the method may comprise reacting any one or more of a variety of functional groups e.g., carboxylic acid (COOH), free amine (--NH.sub.2), or sulfhydryl (--SH) groups present on the PE with a suitable functional group on the targeting moiety, thereby forming a covalent bind between the PE and the targeting moiety. Alternatively or additionally, the method may comprise derivatizing the targeting moiety or PE to expose or to attach additional reactive functional groups. Derivatizing may also include attaching one or more linkers to the targeting moiety or PE.
[0312] In another embodiment of the invention, the inventive PEs and chimeric molecules may be produced using non-recombinant methods. For example, the inventive PEs and chimeric molecules described herein (including functional portions and functional variants) can be commercially synthesized by companies, such as Synpep (Dublin, Calif.), Peptide Technologies Corp. (Gaithersburg, Md.), and Multiple Peptide Systems (San Diego, Calif.). In this respect, the inventive PEs and chimeric molecules can be synthetic, recombinant, isolated, and/or purified.
[0313] It may be desirable, in some circumstances, to free the PE from the targeting moiety when the chimeric molecule has reached one or more target cells. In this regard, the inventive chimeric molecules may comprise a cleavable linker. The linker may be cleavable by any suitable means, e.g., enzymatically. For example, when the target cell is a cancer (e.g., tumor) cell, the chimeric molecule may include a linker cleavable under conditions present at the tumor site (e.g., when exposed to tumor-associated enzymes or acidic pH).
[0314] An embodiment of the invention provides a nucleic acid comprising a nucleotide sequence encoding any of the inventive PEs or the inventive chimeric molecules described herein. The term "nucleic acid," as used herein, includes "polynucleotide," "oligonucleotide," and "nucleic acid molecule," and generally means a polymer of DNA or RNA, which can be single-stranded or double-stranded, which can be synthesized or obtained (e.g., isolated and/or purified) from natural sources, which can contain natural, non-natural or altered nucleotides, and which can contain a natural, non-natural, or altered internucleotide linkage, such as a phosphoroamidate linkage or a phosphorothioate linkage, instead of the phosphodiester found between the nucleotides of an unmodified oligonucleotide. It is generally preferred that the nucleic acid does not comprise any insertions, deletions, inversions, and/or substitutions. However, it may be suitable in some instances, as discussed herein, for the nucleic acid to comprise one or more insertions, deletions, inversions, and/or substitutions.
[0315] Preferably, the nucleic acids of the invention are recombinant. As used herein, the term "recombinant" refers to (i) molecules that are constructed outside living cells by joining natural or synthetic nucleic acid segments, or (ii) molecules that result from the replication of those described in (i) above. For purposes herein, the replication can be in vitro replication or in vivo replication.
[0316] The nucleic acids can be constructed based on chemical synthesis and/or enzymatic ligation reactions using procedures known in the art. See, for example, Sambrook et al., supra, and Ausubel et al., supra. For example, a nucleic acid can be chemically synthesized using naturally occurring nucleotides or variously modified nucleotides designed to increase the biological stability of the molecules or to increase the physical stability of the duplex formed upon hybridization (e.g., phosphorothioate derivatives and acridine substituted nucleotides). Examples of modified nucleotides that can be used to generate the nucleic acids include, but are not limited to, 5-fluorouracil, 5-bromouracil, 5-chlorouracil, 5-iodouracil, hypoxanthine, xanthine, 4-acetylcytosine, 5-(carboxyhydroxymethyl) uracil, 5-carboxymethylaminomethyl-2-thiouridine, 5-carboxymethylaminomethyluracil, dihydrouracil, beta-D-galactosylqueosine, inosine, N.sup.6-isopentenyladenine, 1-methylguanine, 1-methylinosine, 2,2-dimethylguanine, 2-methyladenine, 2-methylguanine, 3-methylcytosine, 5-methylcytosine, N.sup.6-substituted adenine, 7-methylguanine, 5-methylaminomethyluracil, 5-methoxyaminomethyl-2-thiouracil, beta-D-mannosylqueosine, 5'-methoxycarboxymethyluracil, 5-methoxyuracil, 2-methylthio-N.sup.6-isopentenyladenine, uracil-5-oxyacetic acid (v), wybutoxosine, pseudouracil, queosine, 2-thiocytosine, 5-methyl-2-thiouracil, 2-thiouracil, 4-thiouracil, 5-methyluracil, uracil-5-oxyacetic acid methylester, 3-(3-amino-3-N-2-carboxypropyl) uracil, and 2,6-diaminopurine. Alternatively, one or more of the nucleic acids of the invention can be purchased from companies, such as Macromolecular Resources (Fort Collins, Colo.) and Synthegen (Houston, Tex.).
[0317] The invention also provides a nucleic acid comprising a nucleotide sequence which is complementary to the nucleotide sequence of any of the nucleic acids described herein or a nucleotide sequence which hybridizes under stringent conditions to the nucleotide sequence of any of the nucleic acids described herein.
[0318] The nucleotide sequence which hybridizes under stringent conditions preferably hybridizes under high stringency conditions. By "high stringency conditions" is meant that the nucleotide sequence specifically hybridizes to a target sequence (the nucleotide sequence of any of the nucleic acids described herein) in an amount that is detectably stronger than non-specific hybridization. High stringency conditions include conditions which would distinguish a polynucleotide with an exact complementary sequence, or one containing only a few scattered mismatches, from a random sequence that happened to have only a few small regions (e.g., 3-10 bases) that matched the nucleotide sequence. Such small regions of complementarity are more easily melted than a full-length complement of 14-17 or more bases, and high stringency hybridization makes them easily distinguishable. Relatively high stringency conditions would include, for example, low salt and/or high temperature conditions, such as provided by about 0.02-0.1 M NaCl or the equivalent, at temperatures of about 50-70.degree. C. Such high stringency conditions tolerate little, if any, mismatch between the nucleotide sequence and the template or target strand, and are particularly suitable for detecting expression of any of the inventive PEs or chimeric molecules. It is generally appreciated that conditions can be rendered more stringent by the addition of increasing amounts of formamide.
[0319] The invention also provides a nucleic acid comprising a nucleotide sequence that is about 70% or more, e.g., about 80% or more, about 90% or more, about 91% or more, about 92% or more, about 93% or more, about 94% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, or about 99% or more identical to any of the nucleic acids described herein.
[0320] The nucleic acids of the invention can be incorporated into a recombinant expression vector. In this regard, the invention provides recombinant expression vectors comprising any of the nucleic acids of the invention. For purposes herein, the term "recombinant expression vector" means a genetically-modified oligonucleotide or polynucleotide construct that permits the expression of an mRNA, protein, polypeptide, or peptide by a host cell, when the construct comprises a nucleotide sequence encoding the mRNA, protein, polypeptide, or peptide, and the vector is contacted with the cell under conditions sufficient to have the mRNA, protein, polypeptide, or peptide expressed within the cell. The vectors of the invention are not naturally-occurring as a whole. However, parts of the vectors can be naturally-occurring. The inventive recombinant expression vectors can comprise any type of nucleotides, including, but not limited to DNA and RNA, which can be single-stranded or double-stranded, which can be synthesized or obtained in part from natural sources, and which can contain natural, non-natural or altered nucleotides. The recombinant expression vectors can comprise naturally-occurring, non-naturally-occurring internucleotide linkages, or both types of linkages. Preferably, the non-naturally occurring or altered nucleotides or internucleotide linkages does not hinder the transcription or replication of the vector.
[0321] The recombinant expression vector of the invention can be any suitable recombinant expression vector, and can be used to transform or transfect any suitable host cell. Suitable vectors include those designed for propagation and expansion or for expression or for both, such as plasmids and viruses. The vector can be selected from the group consisting of the pUC series (Fermentas Life Sciences), the pBluescript series (Stratagene, LaJolla, Calif.), the pET series (Novagen, Madison, Wis.), the pGEX series (Pharmacia Biotech, Uppsala, Sweden), and the pEX series (Clontech, Palo Alto, Calif.). Bacteriophage vectors, such as .lamda.GT10, .lamda.GT11, .lamda.ZapII (Stratagene), kEMBL4, and XNM1149, also can be used. Examples of plant expression vectors include pBI01, pBI101.2, pBI101.3, pBI121 and pBIN19 (Clontech). Examples of animal expression vectors include pEUK-C1, pMAM, and pMAMneo (Clontech). Preferably, the recombinant expression vector is a viral vector, e.g., a retroviral vector.
[0322] The recombinant expression vectors of the invention can be prepared using standard recombinant DNA techniques described in, for example, Sambrook et al., supra, and Ausubel et al., supra. Constructs of expression vectors, which are circular or linear, can be prepared to contain a replication system functional in a prokaryotic or eukaryotic host cell. Replication systems can be derived, e.g., from ColE1, 2.mu. plasmid, .lamda., SV40, bovine papilloma virus, and the like.
[0323] Desirably, the recombinant expression vector comprises regulatory sequences, such as transcription and translation initiation and termination codons, which are specific to the type of host (e.g., bacterium, fungus, plant, or animal) into which the vector is to be introduced, as appropriate and taking into consideration whether the vector is DNA- or RNA-based.
[0324] The recombinant expression vector can include one or more marker genes, which allow for selection of transformed or transfected hosts. Marker genes include biocide resistance, e.g., resistance to antibiotics, heavy metals, etc., complementation in an auxotrophic host to provide prototrophy, and the like. Suitable marker genes for the inventive expression vectors include, for instance, neomycin/G418 resistance genes, hygromycin resistance genes, histidinol resistance genes, tetracycline resistance genes, and ampicillin resistance genes.
[0325] The recombinant expression vector can comprise a native or nonnative promoter operably linked to the nucleotide sequence encoding the inventive PE or chimeric molecule (including functional portions and functional variants), or to the nucleotide sequence which is complementary to or which hybridizes to the nucleotide sequence encoding the PE or chimeric molecule. The selection of promoters, e.g., strong, weak, inducible, tissue-specific, and developmental-specific, is within the ordinary skill of the artisan. Similarly, the combining of a nucleotide sequence with a promoter is also within the ordinary skill of the artisan. The promoter can be a non-viral promoter or a viral promoter, e.g., a cytomegalovirus (CMV) promoter, an SV40 promoter, an RSV promoter, or a promoter found in the long-terminal repeat of the murine stem cell virus.
[0326] The inventive recombinant expression vectors can be designed for either transient expression, for stable expression, or for both. Also, the recombinant expression vectors can be made for constitutive expression or for inducible expression.
[0327] Another embodiment of the invention further provides a host cell comprising any of the recombinant expression vectors described herein. As used herein, the term "host cell" refers to any type of cell that can contain the inventive recombinant expression vector. The host cell can be a eukaryotic cell, e.g., plant, animal, fungi, or algae, or can be a prokaryotic cell, e.g., bacteria or protozoa. The host cell can be a cultured cell, an adherent cell or a suspended cell, i.e., a cell that grows in suspension. For purposes of producing a recombinant inventive PE or chimeric molecule, the host cell is preferably a prokaryotic cell, e.g., an E. coli cell.
[0328] Also provided by the invention is a population of cells comprising at least one host cell described herein. The population of cells can be a heterogeneous population comprising the host cell comprising any of the recombinant expression vectors described, in addition to at least one other cell, e.g., a host cell which does not comprise any of the recombinant expression vectors. Alternatively, the population of cells can be a substantially homogeneous population, in which the population comprises mainly (e.g., consisting essentially of) host cells comprising the recombinant expression vector. The population also can be a clonal population of cells, in which all cells of the population are clones of a single host cell comprising a recombinant expression vector, such that all cells of the population comprise the recombinant expression vector. In one embodiment of the invention, the population of cells is a clonal population of host cells comprising a recombinant expression vector as described herein.
[0329] The inventive PEs, chimeric molecules (including functional portions and functional variants), nucleic acids, recombinant expression vectors, host cells (including populations thereof), and populations of cells can be isolated and/or purified. The term "isolated" as used herein means having been removed from its natural environment. The term "purified" as used herein means having been increased in purity, wherein "purity" is a relative term, and not to be necessarily construed as absolute purity. For example, the purity can be about 50% or more, about 60% or more, about 70% or more, about 80% or more, about 90% or more, or about 100%. The purity preferably is about 90% or more (e.g., about 90% to about 95%) and more preferably about 98% or more (e.g., about 98% to about 99%).
[0330] The inventive PEs, chimeric molecules (including functional portions and functional variants), nucleic acids, recombinant expression vectors, host cells (including populations thereof), and populations of cells, all of which are collectively referred to as "inventive PE materials" hereinafter, can be formulated into a composition, such as a pharmaceutical composition. In this regard, the invention provides a pharmaceutical composition comprising any of the PEs, chimeric molecules (including functional portions and functional variants), nucleic acids, recombinant expression vectors, host cells (including populations thereof), and populations of cells, and a pharmaceutically acceptable carrier. The inventive pharmaceutical composition containing any of the inventive PE materials can comprise more than one inventive PE material, e.g., a polypeptide and a nucleic acid, or two or more different PEs. Alternatively, the pharmaceutical composition can comprise an inventive PE material in combination with one or more other pharmaceutically active agents or drugs, such as a chemotherapeutic agents, e.g., asparaginase, busulfan, carboplatin, cisplatin, daunorubicin, doxorubicin, fluorouracil, gemcitabine, hydroxyurea, methotrexate, paclitaxel, rituximab, vinblastine, vincristine, etc.
[0331] Preferably, the carrier is a pharmaceutically acceptable carrier. With respect to pharmaceutical compositions, the carrier can be any of those conventionally used and is limited only by chemico-physical considerations, such as solubility and lack of reactivity with the active compound(s), and by the route of administration. The pharmaceutically acceptable carriers described herein, for example, vehicles, adjuvants, excipients, and diluents, are well-known to those skilled in the art and are readily available to the public. It is preferred that the pharmaceutically acceptable carrier be one which is chemically inert to the active agent(s) and one which has no detrimental side effects or toxicity under the conditions of use.
[0332] The choice of carrier will be determined in part by the particular inventive PE material, as well as by the particular method used to administer the inventive PE material. Accordingly, there are a variety of suitable formulations of the pharmaceutical composition of the invention. The following formulations for parenteral (e.g., subcutaneous, intravenous, intraarterial, intramuscular, intradermal, interperitoneal, and intrathecal), oral, and aerosol administration are exemplary and are in no way limiting. More than one route can be used to administer the inventive PE materials, and in certain instances, a particular route can provide a more immediate and more effective response than another route.
[0333] Topical formulations are well-known to those of skill in the art. Such formulations are particularly suitable in the context of the invention for application to the skin.
[0334] Formulations suitable for oral administration can include (a) liquid solutions, such as an effective amount of the inventive PE material dissolved in diluents, such as water, saline, or orange juice; (b) capsules, sachets, tablets, lozenges, and troches, each containing a predetermined amount of the active ingredient, as solids or granules; (c) powders; (d) suspensions in an appropriate liquid; and (e) suitable emulsions. Liquid formulations may include diluents, such as water and alcohols, for example, ethanol, benzyl alcohol, and the polyethylene alcohols, either with or without the addition of a pharmaceutically acceptable surfactant. Capsule forms can be of the ordinary hard- or soft-shelled gelatin type containing, for example, surfactants, lubricants, and inert fillers, such as lactose, sucrose, calcium phosphate, and corn starch. Tablet forms can include one or more of lactose, sucrose, mannitol, corn starch, potato starch, alginic acid, microcrystalline cellulose, acacia, gelatin, guar gum, colloidal silicon dioxide, croscarmellose sodium, talc, magnesium stearate, calcium stearate, zinc stearate, stearic acid, and other excipients, colorants, diluents, buffering agents, disintegrating agents, moistening agents, preservatives, flavoring agents, and other pharmacologically compatible excipients. Lozenge forms can comprise the inventive PE material in a flavor, usually sucrose and acacia or tragacanth, as well as pastilles comprising the inventive PE material in an inert base, such as gelatin and glycerin, or sucrose and acacia, emulsions, gels, and the like additionally containing such excipients as are known in the art.
[0335] The inventive PE material, alone or in combination with other suitable components, can be made into aerosol formulations to be administered via inhalation. These aerosol formulations can be placed into pressurized acceptable propellants, such as dichlorodifluoromethane, propane, nitrogen, and the like. The aerosol formulations also may be formulated as pharmaceuticals for non-pressured preparations, such as in a nebulizer or an atomizer. Such spray formulations also may be used to spray mucosa.
[0336] Formulations suitable for parenteral administration include aqueous and non-aqueous, isotonic sterile injection solutions, which can contain anti-oxidants, buffers, bacteriostats, and solutes that render the formulation isotonic with the blood of the intended recipient, and aqueous and non-aqueous sterile suspensions that can include suspending agents, solubilizers, thickening agents, stabilizers, and preservatives. The inventive PE material can be administered in a physiologically acceptable diluent in a pharmaceutical carrier, such as a sterile liquid or mixture of liquids, including water, saline, aqueous dextrose and related sugar solutions, an alcohol, such as ethanol or hexadecyl alcohol, a glycol, such as propylene glycol or polyethylene glycol, dimethylsulfoxide, glycerol, ketals such as 2,2-dimethyl-1,3-dioxolane-4-methanol, ethers, poly(ethyleneglycol) 400, oils, fatty acids, fatty acid esters or glycerides, or acetylated fatty acid glycerides with or without the addition of a pharmaceutically acceptable surfactant, such as a soap or a detergent, suspending agent, such as pectin, carbomers, methylcellulose, hydroxypropylmethylcellulose, or carboxymethylcellulose, or emulsifying agents and other pharmaceutical adjuvants.
[0337] Oils, which can be used in parenteral formulations include petroleum, animal, vegetable, or synthetic oils. Specific examples of oils include peanut, soybean, sesame, cottonseed, corn, olive, petrolatum, and mineral. Suitable fatty acids for use in parenteral formulations include oleic acid, stearic acid, and isostearic acid. Ethyl oleate and isopropyl myristate are examples of suitable fatty acid esters.
[0338] Suitable soaps for use in parenteral formulations include fatty alkali metal, ammonium, and triethanolamine salts, and suitable detergents include (a) cationic detergents such as, for example, dimethyl dialkyl ammonium halides, and alkyl pyridinium halides, (b) anionic detergents such as, for example, alkyl, aryl, and olefin sulfonates, alkyl, olefin, ether, and monoglyceride sulfates, and sulfosuccinates, (c) nonionic detergents such as, for example, fatty amine oxides, fatty acid alkanolamides, and polyoxyethylenepolypropylene copolymers, (d) amphoteric detergents such as, for example, alkyl-.beta.-aminopropionates, and 2-alkyl-imidazoline quaternary ammonium salts, and (e) mixtures thereof.
[0339] The parenteral formulations will typically contain from about 0.5% to about 25% by weight of the inventive PE material in solution. Preservatives and buffers may be used. In order to minimize or eliminate irritation at the site of injection, such compositions may contain one or more nonionic surfactants having a hydrophile-lipophile balance (HLB) of from about 12 to about 17. The quantity of surfactant in such formulations will typically range from about 5% to about 15% by weight. Suitable surfactants include polyethylene glycol sorbitan fatty acid esters, such as sorbitan monooleate and the high molecular weight adducts of ethylene oxide with a hydrophobic base, formed by the condensation of propylene oxide with propylene glycol. The parenteral formulations can be presented in unit-dose or multi-dose sealed containers, such as ampoules and vials, and can be stored in a freeze-dried (lyophilized) condition requiring only the addition of the sterile liquid excipient, for example, water, for injections, immediately prior to use. Extemporaneous injection solutions and suspensions can be prepared from sterile powders, granules, and tablets of the kind previously described. The requirements for effective pharmaceutical carriers for parenteral compositions are well-known to those of ordinary skill in the art (see, e.g., Pharmaceutics and Pharmacy Practice, J. B. Lippincott Company, Philadelphia, Pa., Banker and Chalmers, eds., pages 238-250 (1982), and ASHP Handbook on Injectable Drugs, Toissel, 4th ed., pages 622-630 (1986)).
[0340] It will be appreciated by one of skill in the art that, in addition to the above-described pharmaceutical compositions, the inventive PE materials of the invention can be formulated as inclusion complexes, such as cyclodextrin inclusion complexes, or liposomes.
[0341] For purposes of the invention, the amount or dose of the inventive PE material administered should be sufficient to effect a desired response, e.g., a therapeutic or prophylactic response, in the mammal over a reasonable time frame. For example, the dose of the inventive PE material should be sufficient to inhibit growth of a target cell or treat or prevent cancer in a period of from about 2 hours or longer, e.g., 12 to 24 or more hours, from the time of administration. In certain embodiments, the time period could be even longer. The dose will be determined by the efficacy of the particular inventive PE material and the condition of the mammal (e.g., human), as well as the body weight of the mammal (e.g., human) to be treated.
[0342] Many assays for determining an administered dose are known in the art. An administered dose may be determined in vitro (e.g., cell cultures) or in vivo (e.g., animal studies). For example, an administered dose may be determined by determining the IC.sub.50 (the dose that achieves a half-maximal inhibition of symptoms), LD.sub.50 (the dose lethal to 50% of the population), the ED.sub.50 (the dose therapeutically effective in 50% of the population), and the therapeutic index in cell culture and/or animal studies. The therapeutic index is the ratio of LD.sub.50 to ED.sub.50 (i.e., LD.sub.50/ED.sub.50).
[0343] The dose of the inventive PE material also will be determined by the existence, nature, and extent of any adverse side effects that might accompany the administration of a particular inventive PE material. Typically, the attending physician will decide the dosage of the inventive PE material with which to treat each individual patient, taking into consideration a variety of factors, such as age, body weight, general health, diet, sex, inventive PE material to be administered, route of administration, and the severity of the condition being treated. By way of example and not intending to limit the invention, the dose of the inventive PE material can be about 0.001 to about 1000 mg/kg body weight of the subject being treated/day, from about 0.01 to about 10 mg/kg body weight/day, about 0.01 mg to about 1 mg/kg body weight/day, from about 1 to about to about 1000 mg/kg body weight/day, from about 5 to about 500 mg/kg body weight/day, from about 10 to about 250 mg/kg body weight/day, about 25 to about 150 mg/kg body weight/day, or about 10 mg/kg body weight/day.
[0344] Alternatively, the inventive PE materials can be modified into a depot form, such that the manner in which the inventive PE material is released into the body to which it is administered is controlled with respect to time and location within the body (see, for example, U.S. Pat. No. 4,450,150). Depot forms of inventive PE materials can be, for example, an implantable composition comprising the inventive PE materials and a porous or non-porous material, such as a polymer, wherein the inventive PE materials is encapsulated by or diffused throughout the material and/or degradation of the non-porous material. The depot is then implanted into the desired location within the body and the inventive PE materials are released from the implant at a predetermined rate.
[0345] The inventive PE materials may be assayed for cytoxicity by assays known in the art. Examples of cytotoxicity assays include a WST assay, which measures cell proliferation using the tetrazolium salt WST-1 (reagents and kits available from Roche Applied Sciences), as described in International Patent Application Publication WO 2011/032022.
[0346] It is contemplated that the inventive pharmaceutical compositions, PEs, chimeric molecules, nucleic acids, recombinant expression vectors, host cells, or populations of cells can be used in methods of treating or preventing cancer. Without being bound by a particular theory or mechanism, it is believed that the inventive PEs destroy or inhibit the growth of cells through the inhibition of protein synthesis in eukaryotic cells, e.g., by the inactivation of the ADP-ribosylation of elongation factor 2 (EF-2). Without being bound to a particular theory or mechanism, the inventive chimeric molecules recognize and specifically bind to cell surface markers, thereby delivering the cytotoxic PE to the population of cells expressing the cell surface marker with minimal or no cross-reactivity with cells that do not express the cell surface marker. In this way, the cytoxicity of PE can be targeted to destroy or inhibit the growth of a particular population of cells, e.g., cancer cells. In this regard, the invention provides a method of treating or preventing cancer in a mammal comprising administering to the mammal any of the PEs, chimeric molecules, nucleic acids, recombinant expression vectors, host cell, population of cells, or pharmaceutical compositions described herein, in an amount effective to treat or prevent cancer in the mammal.
[0347] The terms "treat" and "prevent" as well as words stemming therefrom, as used herein, do not necessarily imply 100% or complete treatment or prevention. Rather, there are varying degrees of treatment or prevention of which one of ordinary skill in the art recognizes as having a potential benefit or therapeutic effect. In this respect, the inventive methods can provide any amount of any level of treatment or prevention of cancer in a mammal. Furthermore, the treatment or prevention provided by the inventive method can include treatment or prevention of one or more conditions or symptoms of the disease, e.g., cancer, being treated or prevented. Also, for purposes herein, "prevention" can encompass delaying the onset of the disease, or a symptom or condition thereof.
[0348] For purposes of the inventive methods, wherein host cells or populations of cells are administered, the cells can be cells that are allogeneic or autologous to the host. Preferably, the cells are autologous to the host.
[0349] With respect to the inventive methods, the cancer can be any cancer, including any of adrenal gland cancer, sarcomas (e.g., synovial sarcoma, osteogenic sarcoma, leiomyosarcoma uteri, angiosarcoma, fibrosarcoma, rhabdomyosarcoma, liposarcoma, myxoma, rhabdomyoma, fibroma, lipoma, and teratoma), lymphomas (e.g., small lymphocytic lymphoma, Hodgkin lymphoma, and non-Hodgkin lymphoma), hepatocellular carcinoma, glioma, head cancers (e.g., squamous cell carcinoma), neck cancers (e.g., squamous cell carcinoma), acute lymphocytic cancer, leukemias (e.g., hairy cell leukemia, myeloid leukemia (acute and chronic), lymphatic leukemia (acute and chronic), prolymphocytic leukemia (PLL), myelomonocytic leukemia (acute and chronic), and lymphocytic leukemia (acute and chronic)), bone cancer (osteogenic sarcoma, fibrosarcoma, malignant fibrous histiocytoma, chondrosarcoma, Ewing's sarcoma, malignant lymphoma (reticulum cell sarcoma), multiple myeloma, malignant giant cell tumor, chordoma, osteochondroma (ostcocartilaginous exostoses), benign chondroma, chondroblastoma, chondromyxoid fibroma, osteoid osteoma, and giant cell tumors), brain cancer (astrocytoma, medulloblastoma, glioma, ependymoma, germinoma (pinealoma), glioblastoma multiforme, oligodendroglioma, schwannoma, and retinoblastoma), fallopian tube cancer, breast cancer, cancer of the anus, anal canal, or anorectum, cancer of the eye, cancer of the intrahepatic bile duct, cancer of the joints, cancer of the neck, gallbladder, or pleura, cancer of the nose, nasal cavity, or middle ear, cancer of the oral cavity, cancer of the vulva (e.g., squamous cell carcinoma, intraepithelial carcinoma, adenocarcinoma, and fibrosarcoma), myeloproliferative disorders (e.g., chronic myeloid cancer), colon cancers (e.g., colon carcinoma), esophageal cancer (e.g., squamous cell carcinoma, adenocarcinoma, leiomyosarcoma, and lymphoma), cervical cancer (cervical carcinoma and pre-invasive cervical dysplasia), gastric cancer, gastrointestinal carcinoid tumor, hypopharynx cancer, larynx cancer, liver cancers (e.g., hepatocellular carcinoma, cholangiocarcinoma, hepatoblastoma, angiosarcoma, hepatocellular adenoma, and hemangioma), lung cancers (e.g., bronchogenic carcinoma (squamous cell, undifferentiated small cell, undifferentiated large cell, and adenocarcinoma), alveolar (bronchiolar) carcinoma, bronchial adenoma, chondromatous hamartoma, small cell lung cancer, non-small cell lung cancer, and lung adenocarcinoma), mesothelioma, skin cancer (e.g., melanoma, basal cell carcinoma, squamous cell carcinoma, Kaposi's sarcoma, nevi, dysplastic nevi, lipoma, angioma, dermatofibroma, and keloids), multiple myeloma, nasopharynx cancer, ovarian cancer (e.g., ovarian carcinoma (serous cystadenocarcinoma, mucinous cystadenocarcinoma, endometrioid carcinoma, and clear cell adenocarcinoma), granulosa-theca cell tumors, Sertoli-Leydig cell tumors, dysgerminoma, and malignant teratoma), pancreatic cancer (e.g., ductal adenocarcinoma, insulinoma, glucagonoma, gastrinoma, carcinoid tumors, and VIPoma), peritoneum, omentum, mesentery cancer, pharynx cancer, prostate cancer (e.g., adenocarcinoma and sarcoma), rectal cancer, kidney cancer (e.g., adenocarcinoma, Wilms tumor (nephroblastoma), and renal cell carcinoma), small intestine cancer (adenocarcinoma, lymphoma, carcinoid tumors, Kaposi's sarcoma, leiomyoma, hemangioma, lipoma, neurofibroma, and fibroma), soft tissue cancer, stomach cancer (e.g., carcinoma, lymphoma, and leiomyosarcoma), testicular cancer (e.g., seminoma, teratoma, embryonal carcinoma, teratocarcinoma, choriocarcinoma, sarcoma, Leydig cell tumor, fibroma, fibroadenoma, adenomatoid tumors, and lipoma), cancer of the uterus (e.g., endometrial carcinoma), thyroid cancer, and urothelial cancers (e.g., squamous cell carcinoma, transitional cell carcinoma, adenocarcinoma, ureter cancer, and urinary bladder cancer).
[0350] As used herein, the term "mammal" refers to any mammal, including, but not limited to, mammals of the order Rodentia, such as mice and hamsters, and mammals of the order Logomorpha, such as rabbits. It is preferred that the mammals are from the order Carnivora, including Felines (cats) and Canines (dogs). It is more preferred that the mammals are from the order Artiodactyla, including Bovines (cows) and Swines (pigs) or of the order Perssodactyla, including Equines (horses). It is most preferred that the mammals are of the order Primates, Ceboids, or Simoids (monkeys) or of the order Anthropoids (humans and apes). An especially preferred mammal is the human.
[0351] Also provided is a method of inhibiting the growth of a target cell comprising contacting the cell with the PE of any of the PEs, chimeric molecules, nucleic acids, recombinant expression vectors, host cell, population of cells, or pharmaceutical compositions described herein, in an amount effective to inhibit growth of the target cell. The growth of the target cell may be inhibited by any amount, e.g., by about 10% or more, about 15% or more, about 20% or more, about 25% or more, about 30% or more, about 35% or more, about 40% or more, about 45% or more, about 50% or more, about 55% or more, about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, or about 100%. The target cell may be provided in a biological sample. A biological sample may be obtained from a mammal in any suitable manner and from any suitable source. The biological sample may, for example, be obtained by a blood draw, leukapheresis, and/or tumor biopsy or necropsy. The contacting can take place in vitro or in vivo with respect to the mammal. Preferably, the contacting is in vitro.
[0352] In an embodiment of the invention, the target cell is a cancer cell. The target cell may be a cancer cell of any of the cancers described herein. In an embodiment of the invention, the target may express a cell surface marker. The cell surface marker may be any cell surface marker described herein with respect to other aspects of the invention. The cell surface marker may be, for example, selected from the group consisting of CD19, CD21, CD22, CD25, CD30, CD33, CD79b, transferrin receptor, EGF receptor, mesothelin, cadherin, and Lewis Y.
[0353] The following examples further illustrate the invention but, of course, should not be construed as in any way limiting its scope.
Example 1
[0354] This example demonstrates the identification of T cell epitopes in domain III of PE38.
[0355] To identify the T cell epitopes in PE38, peripheral blood mononuclear cells (PBMCs) were first incubated with a recombinant immunotoxin (RIT) for 14 days to allow the RIT to be processed by donor APCs to relevant peptides to be presented to T cells. The enriched PBMCs were subsequently exposed to a peptide library composed of 111 partially overlapping peptides spanning the sequence of PE38 and T cell activation was measured using an ELISpot assay for interleukin-2 (IL-2).
[0356] Samples from 50 normal donors with no recorded previous exposure to PE38 and with a broad distribution of human leukocyte antigen (HLA) alleles were analyzed. The cells of this population contained naive T cell epitope. Samples from nine mesothelioma and seven hairy cell leukemia (IIHCL) patients that were previously treated with PE-based RIT and who had developed high levels of neutralizing antibodies were also analyzed. The cells of this population contained memory T cell epitopes. The responses of the naive donors and the previously treated patients are shown in a heat map format in FIGS. 1A-1B.
[0357] FIG. 1A presents the positive and negative responses of the 50 naive donors. The responses are shown as a percentage of responsive spots for each donor. The strongest positive response is shown as >20%, medium responses are shown as 10%-20%, weak responses are shown as 3%-10%, and the absence of a response (negative) is shown as <3%. The results included a total of 258 positive responses, and 201 of those were weak (3-10%), 44 were medium (10-20%), and 13 were strong (>20%), leaving a total of 5292 negative responses (<3%). There were 122 responses in domain II (peptides 1-38) including 11 strong, 24 medium, and 87 weak responses. There were 136 responses in domain III (peptides 39-111) including two strong, 20 medium and 114 weak responses.
[0358] Peptides 15 and 14 (in pool 3) provided the strongest and most frequent responses in the study with 21 and 18 positive responses, respectively. A total of 36 peptides did not provide any response. Sixteen of the 50 naive donors had a single epitope giving a response, as indicated by one, two, or three responses in overlapping consecutive peptides. Fifteen donors had two separate epitopes, 14 had three epitopes, and five had four epitopes. On 22 occasions, a fine screen of a positive pool did not provide a specific peptide response or was not done, resulting in five consecutive positive responses in the heat map.
[0359] FIG. 1B illustrates the positive and negative responses of the nine mesothelioma and the seven HCL patients. It was found that the mesothelioma patients provided 919 negative responses and 80 responses, including two strong responses (>20%), four medium responses (10-20%), and 74 weak responses (3-10%). The two strong responses were in peptides 13 and 14, and the medium responses were in peptides 15, 35, 57 and 74. Thirty-five of the responses were in domain II (peptides 1-38) and 45 in domain III (peptides 38-111). The hot spot pattern created in the map is similar to the hot spot pattern in the naive donors map. The responsiveness (meaning how many epitopes were found for each patient) was higher than the donor's pattern with one patient having two epitopes, four patients having three epitopes, two patients having four epitopes and two more patients having five epitopes.
[0360] Moreover, it was found that the seven HCL patients provided three strong responses (>20%), nine medium responses (10-20%), 37 weak responses (3-10%), and 728 negative responses (<3%). The strong responses were all in the consecutive peptides 75, 76 and 77. Twenty-two of the responses were in domain 11 and 37 responses were in domain III. Interestingly, the immunodominant epitope in peptides 14 and 15 that was identified in more than 40% of the naive donors and 55% of the mesothelioma patients (five out of nine samples) did not have a single response in the HCL patients. One patient responded to peptides 12 and 13 that have amino acids in common with peptides 14 and 15. However, no responses were found directly to peptides 14 and 15. This may be attributed to the biased HLA distribution of those patients' samples or to the nature of the disease.
[0361] Overall, eight major epitopes were identified in PE38 that account for 93% of the responses. Because all T cell epitopes in domain II could be eliminated using the LR deletion, the five epitopes remaining in domain III were the focus of subsequent study. Table 1A summarizes the epitopes' locations, sequences, response rates, and the mutation that diminishes each with minimal disturbance to cytotoxic activity. The epitope ranking was composed of a number of factors, including the number of naive donors responding to the epitope, the relative strength of the response, the number of mesothelioma patients that responded, and, to a lesser extent, the HCL patients' responses. The epitopes from HCL patients were evaluated to a lesser extent in this analysis because of the HLA bias of the patient cohort and in the HCL patient population, in which HCL patients have an enrichment of BRB1_11 (Annino et al., Leuk. Lymphoma, 14 (Suppl.) 1:63-5 (1994)). Epitope 2 spans five peptides. To simplify analysis, it was divided into two sub-epitopes: peptides 77-78 (2A) and peptides 75-76 (2B). Some donors responded to both 2A and 2B epitopes, while others responded to only one. Epitope 6 (peptides 93-96) and epitope 8 (peptides 56-59) were also divided into two sub-epitopes.
TABLE-US-00001 TABLE 1A Responses Relative Epitope Peptide Donors Mesothelioma HCL cytotoxic ranking # Sequence (n = 50) (n = 9) (n = 7) Mutations Activity* 1 13-15 LVALYLAARLSWNQV 21 6 1 Domain II 100% SEQ ID NO: 189 deletion 2 A 77-78 GALLRVYVPRSSLPG 14.sup.a 3.sup.a 6.sup.a R505A 100% SEQ ID NO: 190 B 74-76 IRNGALLRVYVPRSS 10.sup.a 6.sup.a 5.sup.a R494A 21-36% SEQ ID NO: 191 3 8-9 RQPRGWEQLEQCGYP 9 3 3 Domain II 100% SEQ ID NO: 192 deletion 4 5-6 LPLETFTRHRQPRQW 10 2 0 Domain II 100% SEQ ID NO: 193 deletion 5 67-68 WRGFYIAGDPALAYG 8 2 2 L477H 100% SEQ ID NO: 194 6A + B 93-96 GPEEEGGRLETILGWPLA 8 1 2 L552E 100% SEQ ID NO: 195 7 51-52 TVERLLQAHRQLEER 5 1 0 R427A 100% SEQ ID NO: 196 8A + B 56-59 FVGYHGTFLEAAQSIVFG 5 5 4 F443A >100% SEQ ID NO: 197 *Activity for a single point mutation in HA22 RIT and evaluated in CA46 cell line. .sup.aDonors and patients that responded to epitope 2A overlap with the patients and donors that responded to 2B.
[0362] To compare the results from naive donors to immunotoxin-treated patients, two patient cohorts that made neutralizing antibodies against the RIT were studied. The patients' DRB1 HLA alleles are shown in Table 1B. The same epitopes identified in the donor cohort were also present in the patient cohorts. One cohort was from mesothelioma patients treated with SS1P (anti-mesothelin Fv fused to P38) (Chowdhury et al., Nat. Biotechnol., 17: 568-572 (1999)); the other from leukemia patients treated with moxetumomab pasudotox (MP), a RIT including PE38 fused to an anti-CD22 Fv (Kreitman et al., Clin. Cancer Res., 17: 6398-6405 (2011)). The naive donor epitope responses ranged from 1-4 epitopes per donor, with an average of 2.1, and the patient responses ranged from 1-7 per patient, with a higher average of 3.4 (P<0.001 in Student T test). This suggests that some responses in the naive population were too weak to be detected by this method, and were amplified after exposure to immunotoxin. The patient samples did not identify any major epitopes that were not identified using the donor cohort.
TABLE-US-00002 TABLE 1B HLA haplotype Donor Diagnosis DRB1 71509WBp HCL 04, 11 102609aph HCL 1103, 1303 112309aph HCL 0404, 0701 121809aph HCL 07, 11 050710aph HCL 01, 07 021012b aph HCL 04, 07 071912aph HCL 07, 11 120909 aph Mesothelioma 0410, 1501 011410 aph Mesothelioma 0101, 0801 012810aph Mesothelioma 0301, 1501 050510aph Mesothelioma 0401, 1302 091510 aph Mesothelionla 07, 15 022811aph Mesothelioma 1301, 1302 100711aph Mesothelioma 0103, 03 021012aph Mesothelioma 0101, 0101 031612 aph Mesothelioma 0401, 12
Example 2
[0363] This example demonstrates the elimination of T cell epitopes in domain III.
[0364] For each one of the epitopes in domain III that are described in Table 1, alanine scanning mutagenesis was performed. The alanine mutant was incorporated into the RIT according to the following 11 steps: (1) a list was made of all alanine peptide variants for each epitope; (2) in silico prediction was performed to rule out alanine variants that had increased binding to at least six HLA alleles using an HLA binding algorithm (Immune Epitope Database, IEDB) (Wang et al., PLoS Comput. Biol., 4:e1000048 (2008)) that measured their ability to bind to 13 major HLA groups; (3) alanine variants were assayed using 8-15 donors and patient samples with in vitro expansion and ELIspot; (4) an alanine mutation with diminished T cell activation was identified; (5) the alanine mutation was cloned into the HA22-LR plasmid, and a RIT was constructed with a single point mutation; (6) activity was studied using the WST8 assay.
[0365] If the protein was aggregated or has low cytotxic activity, (7) in silico prediction and crystal structure was used to identify alternative amino acids to alanine and (8) the new mutation was cloned into an HA22-LR plasmid, and a RIT was constructed with a single point mutation and activity was studied. If the protein was aggregated or had low cytotoxic activity, then the next best alanine mutant was identified as in step 4 and step 5 was carried out. If the protein was active, step 10 was carried out.
[0366] If the protein was active, (10) validation was carried out to determine whether the epitope was diminished and that no new epitopes were created due to the mutation; and (11) the successful point mutations were combined into a single RIT.
[0367] The results for all epitopes are shown in Tables 2-7. In Tables 2-7, underlined values represent <10%, D=Donor, Pt=patient, "Meso"=mesothelioma, the position where the amino acid was replaced with alanine is underlined, and the average values are bolded. Alanine variants were assayed using 8-15 donor and patient samples with in vitro expansion and ELIspot.
TABLE-US-00003 TABLE 2 CD4+ T cell response to alanine variant peptides and WT76. donor/ HCL Meso HCL Meso Meso Mean patient Pt Pt Pt Pt Pt D1 D2 D3 D4 D5 n = 16 No peptide 3% 0% 0% 1% 0% 0% 9% 0% 17% 1% 0% WT76 IRNGALLRVYVPRSS 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% SEQ ID NO: 198 I493A ARNGALLRVYVPRSS 2% 13% 0% 36% 52% 6% 20% 110% 71% 76% 25% SEQ ID NO: 199 R494A IANGALLRVYVPRSS 1% 0% 2% 2% 0% 12% 26% 111% 88% 94% 24% SEQ ID NO: 200 N495A IRAGALLRVYVPRSS 95% 25% 2% 31% 12% 35% 40% 102% 72% 72% 34% SEQ ID NO: 201 G496A IRNAALLRVYVPRSS 107% 58% 18% 63% 34% 18% 37% 96% 44% 99% 42% SEQ ID NO: 202 L498A IRNGAALRVYVPRSS 4% 4% 0% 4% 27% 0% 80% 6% 16% 31% 5% SEQ ID NO: 203 L499A IRNGALARVYVPRSS 7% 0% 0% 1% 2% 6% 31% 30% 15% 57% 3% SEQ ID NO: 204 R500A IRNGALLAVYVPRSS 5% 4% 24% 2% 0% 24% 26% 1% 20% 6% 4% SEQ ID NO: 205 V501A IRNGALLRAYVPRSS 5% 0% 7% 13% 29% 0% 46% 25% 27% 48% 11% SEQ ID NO: 206 Y502A IRNGALLRVAVPRSS 29% 4% 18% 23% 70% 12% 23% 5% 20% 0% 19% SEQ ID NO: 207 V503A IRNGALLRVYAPRSS 54% 58% 47% 68% 30% 41% 63% 35% 27% 43% 41% SEQ ID NO: 208 P504A IRNGALLRVYVARSS 91% 125% 49% 122% 117% 206% 60% 35% 26% 18% 74% SEQ ID NO: 209 R505A IRNGALLRVYVPASS 95% 83% 87% 127% 133% 147% 131% 36% 23% 30% 87% SEQ ID NO: 210 S506A IRNGALLRVYVPRAS 92% 138% 44% 107% 100% 312% 214% 101% 55% 82% 106% SEQ ID NO: 211 S507A IRNGALLRVYVPRSA 99% 96% 51% 69% 106% 235% 137% 71% 83% 84% 94% SEQ ID NO: 212
TABLE-US-00004 TABLE 3 T cell response to alanine variant peptides and WT77. HCL HCL HCL HCL Meso Mean D/Pt Pt 1 Pt 2 Pt 3 Pt 4 Pt 1 D1 D2 D3 (n = 8) No peptide 0% 0% 0% 12% 1% 18% 9% 7% 6% wt 77 GALLRVYVPRSSLPG 100% 100% 100% 100% 100% 100% 100% 100% 100% SEQ ID NO: 213 G496A AALLRVYVPRSSLPG 29% 51% 126% 151% 37% 58% 112% 123% 86% SEQ ID NO: 214 A497G GGLLRVYVPRSSLPG 103% 68% 102% 167% 71% 108% 129% 126% 109% SEQ ID NO: 215 L498A GAALRVYVPRSSLPG 61% 38% 7% 166% 43% 20% 13% 140% 61% SEQ ID NO: 216 L499A GALARVYVPRSSLPG 46% 24% 33% 162% 1% 26% 73% 61% 53% SEQ ID NO: 217 R500A GALLAVYVPRSSLPG 59% 31% 1% 189% 6% 16% 1% 124% 53% SEQ ID NO: 218 V501A GALLRAYVPRSSLPG 7% 18% 18% 84% 15% 26% 49% 28% 31% SEQ ID NO: 219 Y502A GALLRVAVPRSSLPG 14% 1% 4% 10% 0% 25% 8% 9% 9% SEQ ID NO: 220 V503A GALLRVYAPRSSLPG 17% 1% 15% 18% 1% 39% 49% 9% 19% SEQ ID NO: 221 P504A GALLRVYVARSSLPG 46% 32% 38% 127% 36% 43% 12% 41% 47% SEQ ID NO: 222 R505A GALLRVYVPASSLPG 7% 1% 28% 15% 2% 24% 39% 22% 17% SEQ ID NO: 223 S506A GALLRVYVPRASLPG 68% 42% 107% 128% 48% 93% 122% 50% 82% SEQ ID NO: 224 S507A GALLRVYVPRSALPG 35% 37% 86% 164% 60% 106% 96% 224% 101% SEQ ID NO: 225 L508A GALLRVYVPRSSAPG 8% 0% 73% 12% 46% 99% 88% 15% 43% SEQ ID NO: 226 P509A GALLRVYVPRSSLAG 10% 4% 87% 57% 72% 101% 119% 16% 58% SEQ ID NO: 227
TABLE-US-00005 TABLE 4 T cell response to alanine variant peptides and WT 67 HCL HCL HCL Meso Meso Mean % from WT Pt 1 Pa 2 Pt 3 Pt 1 Pt 2 D1 D2 D3 D4 D5 (n = 14) No 6% 3% 3% 13% 1% 1% 13% 0% 0% 0% 13% peptide WT 67 WRGFYIAGDPALAYG 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% SEQ ID NO: 228 F469A WRGAYIAGDPALAYG 80% 68% 93% 11% 81% 104% 35% 97% 88% 60% 69% SEQ ID NO: 229 Y470A WRGFAIAGDPALAYG 61% 6% 61% 6% 68% 48% 67% 29% 16% 92% 43% SEQ ID NO: 230 I471A WRGFYAAGDPALAYG 17% 36% 22% 7% 35% 20% 20% 22% 12% 55% 24% SEQ ID NO: 231 A472G WRGFYIGGDPALAYG 37% 81% 38% 20% 60% 84% 75% 46% 19% 76% 48% SEQ ID NO: 232 P475A WRGFYIAGDAALAYG 72% 18% 46% 73% 38% 34% 57% 15% 16% 70% 43% SEQ ID NO: 233 A476G WRGFYIAGDPGLAYG 75% 49% 40% 110% 32% 207% 26% 67% 35% 115% 70% SEQ ID NO: 234 L477A WRGFYIAGDPAAAYG 54% 32% 54% 92% 52% 25% 42% 26% 16% 71% 44% SEQ ID NO: 235 A478G WRGFYIAGDPALGYG 80% 81% 62% 96% 58% 134% 24% 63% 98% 66% 72% SEQ ID NO: 236 Y479A WRGFYIAGDPALAAG 114% 106% 76% 250% 138% 108% 117% 71% 21% 94% 104% SEQ ID NO: 237
TABLE-US-00006 TABLE 5 Alanine scanning for peptides 93-94 GPEEEGGRLETILGWPLA (SEQ ID NO: 238) HCL HCL Meso Meso Meso Mean Pt 1 D1 D2 Pt 2 Pt 1 Pt 2 Pt 3 D3 D4 D5 n = 10 Responder groups 93 93 93 94 94 94 94 94 94 94 No 3% 9% 2% 7% 4% 3% 15% 35% 25% 12% 11% peptide WT 93-94 GPEEEGGRLETILGWPLA 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% SEQ ID NO: 238 E547A GPEAEGGRLETILGWPLA 73% 18% 254% 85% 120% 122% 136% 73% 106% 90% 108% SEQ ID NO: 239 E548A GPEEAGGRLETILGWPLA 58% 24% 170% 112% 98% 86% 137% 102% 207% 87% 108% SEQ ID NO: 240 R551A GPEEEGGALETILGWPLA 62% 6% 2% 95% 70% 42% 54% 40% 67% 49% 49% SEQ ID NO: 241 L552A GPEEEGGRAETILGWPLA 54% 44% 10% 91% 35% 11% 14% 85% 51% 46% 44% SEQ ID NO: 242 T554A GPEEEGGRLEAILGWPLA 77% 9% 5% 94% 133% 130% 129% 110% 95% 112% 89% SEQ ID NO: 243 I555A GPEEEGGRLETALGWPLA 111% 176% 119% 67% 25% 15% 36% 63% 36% 36% 69% SEQ ID NO: 244 L556A GPEEEGGRLETIAGWPLA 24% 235% 3% 40% 10% 13% 43% 35% 83% 29% 51% SEQ ID NO: 245 W558A GPEEEGGRLETILGAPLA 20% 200% 13% 13% 26% 7% 26% 46% 54% 31% 44% SEQ ID NO: 246 P559A GPEEEGGRLETILGWALA 24% 197% 208% 95% 69% 37% 74% 65% 121% 46% 94% SEQ ID NO: 247 L560A GPEEEGGRLETILGWPAA 81% 321% 162% 89% 132% 117% 75% 46% 138% 95% 126% SEQ ID NO: 248
TABLE-US-00007 TABLE 6 Alanine scanning for peptide TVERLLQAHRQLEER (SEQ ID NO: 249) HCL Meso Meso Meso Mean Pt 1 Pt 1 Pt 2 Pt 3 D1 D2 D3 n = 7 No 0% 1% 0% 1% 13% 14% 7% 5% peptide WT51 TVERLLQAHRQLEER 100% 100% 100% 100% 100% 100% 100% 100% SEQ ID NO: 249 R421A TVEALLQAHRQLEER 23% 87% 2% 5% 22% 12% 26% 25% SEQ ID NO: 250 L422A TVERALQAHRQLEER 29% 18% 0% 1% 31% 10% 16% 17% SEQ ID NO: 251 L423A TVERLAQAHRQLEER 47% 21% 11% 3% 8% 10% 6% 14% SEQ ID NO: 252 A425G TVERLLQGHRQLEER 105% 51% 4% 6% 57% 16% 68% 52% SEQ ID NO: 253 R427A TVERLLQAHAQLEER 78% 81% 1% 2% 37% 10% 35% 37% SEQ ID NO: 254 L429A TVERLLQAHRQAEER 63% 64% 38% 36% 28% 19% 124% 67% SEQ ID NO: 255 E430A TVERLLQAHRQLAER 100% 100% 242% 73% 112% 26% 99% 108% SEQ ID NO: 256 R432A TVERLLQAHRQLEEA 105% 92% 87% 69% 65% 57% 142% 106% SEQ ID NO: 257
TABLE-US-00008 TABLE 7 Alanine scanning for 18 mer peptide FVGYHGTFLEAAQSIVFG (57-58) (SEQ ID NO: 258) HCL Meso Meso HCL HCL Meso Meso Pt 1 Pt 1 Pt 2 D1 Pt 2 Pt 3 Pt 3 Pt 4 D2 D3 Avg Responder groups 57 57 57 57 58 58 58 58 58 58 No 0% 0% 0% 4% 0% 1% 1% 0% 10% 9% 4% peptide WT 57-58 FVGYHGTFLEAAQSIVFG 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% 100% SEQ ID NO: 258 F436A AVGYHGTFLEAAQSIVFG 57% 98% 159% 7% 103% 103% 89% 45% 41% 123% 83% SEQ ID NO: 259 V437A FAGYHGTFLEAAQSIVFG 84% 78% 130% 81% 97% 95% 96% 85% 102% 148% 99% SEQ ID NO: 260 G438A FVAYHGTFLEAAQSIVFG 67% 52% 78% 79% 71% 88% 118% 91% 98% 68% 83% SEQ ID NO: 261 Y439A FVGAHGTFLEAAQSIVFG 22% 96% 21% 17% 79% 97% 125% 147% 110% 154% 91% SEQ ID NO: 262 H440A FVGYAGTFLEAAQSIVFG 11% 12% 95% 6% 46% 113% 108% 120% 90% 102% 75% SEQ ID NO: 263 T442A FVGYHGAFLEAAQSIVFG 77% 84% 151% 31% 48% 102% 93% 50% 59% 127% 82% SEQ ID NO: 264 F443A FVGYHGTALEAAQSIVFG 2% 0% 40% 7% 5% 2% 1% 1% 20% 7% 10% SEQ ID NO: 265 L444A FVGYHGTFAEAAQSIVFG 69% 54% 74% 140% 9% 4% 55% 8% 14% 0% 37% SEQ ID NO: 266 A446G FVGYHGTFLEGAQSIVFG 80% 40% 47% 119% 80% 37% 107% 125% 69% 16% 78% SEQ ID NO: 267 A447G FVGYHGTFLEAGQSIVFG 65% 104% 57% 103% 19% 28% 118% 87% 43% 7% 67% SEQ ID NO: 268 5449A FVGYHGTFLEAAQAIVFG 98% 126% 93% 104% 23% 65% 128% 123% 163% 200% 110% SEQ ID NO: 269 I450A FVGYHGTFLEAAQSAVFG 138% 159% 130% 124% 2% 66% 108% 104% 31% 11% 78% SEQ ID NO: 270 V451A FVGYHGTFLEAAQSIAFG 119% 127% 162% 121% 82% 117% 142% 118% 137% 50% 116% SEQ ID NO: 271 L452A FVGYHGTFLEAAQSIVAG 119% 126% 154% 156% 99% 104% 103% 151% 104% 123% 127% SEQ ID NO: 272
[0368] Epitopes 2A and 2B were scanned separately to cover the 9 mer core of all five peptides that gave responses. Y502A diminished the responses of both epitopes (Tables 2 and 3). For epitopes 5 and 7, alanine mutants were compared to peptide 67 and 51, respectively, and I471A and L423A had the lowest T cell response (Tables 4 and 6). To cover all 9 mer cores in epitopes 6 and 8 that contained four positive peptides, 18 mer WT and alanine variants were synthesized. It was found that L552A was the most effective in lowering the response in epitope 6 (Table 5) and F443A was best for epitope 8 (Table 7).
Example 3
[0369] This example demonstrates the cytotoxic activity of point mutation proteins in HA22-LR.
[0370] Step 4 of Example 2 was carried out. The best variants identified in the alanine scan or using further in silico prediction are described in Table 1A.
[0371] Forty mutant RITs were constructed. The cytotoxic activity of point mutation proteins in HA22-LR was evaluated with respect to CA46, Raji, and Daudi cells. Yields of purified protein, calculated accessible surface areas (Lee et al., J. Mol. Biol., 55:379-400 (1971); Roscoe et al., Infect. Immunity, 62:5055-5065 (1994)) of WT amino acids residues and cytotoxic activity of each RIT are shown in Table 8.
TABLE-US-00009 TABLE 8 Protein Protein activity ASA yield Mean relative activity (%) .+-. SD Peptide Mutation (.ANG..sup.2)* (mg) CA46 Raji Daudi 76 R500A 0 A 76 L498A 4 A 76 L499A 0.1 A 76 R494A 35 2.0 21 .+-. 3 33 .+-. 10 36 .+-. 8 76 R494D 35 3.23 11 77 R505A 150 4 78 .+-. 13 185 .+-. 106 76/77 Y502A 0 A 76/77 Y502H 0 A 76/77 Y502F 0 A 76/77 Y502K 0 A 67 I471A 4 1.6 24 67 I471V 4 4 83 108 67 I471M 4 3 3 67 I471H 4 A 67 I471G 4 A 67 I471S 4 1.3 2 67 I471D 4 A 67 Y470A 98 2.3 <1 <1 67 L477A 7 1.4 54 .+-. 21 47 .+-. 13 67 A476G 38 5 67 .+-. 4 84 67 L477G 7 4 14 .+-. 6 14 .+-. 6 67 L477T 7 2.5 <1 <1 67 L477H 7 3 101 .+-. 22 95 .+-. 13 51 L422A 0 2 <1 51 L423A 22 4 30 51 L423N 22 3 5 51 L423T 22 3.2 50 76 51 L423S 22 5 65 51 R421A 16 1 <10 51 R427A 142 0.71 185 .+-. 85 150 .+-. 61 150 .+-. 4 93 R551A 100 4 208 188 .+-. 464 94 W558A 40 1 <1 >2 94 W558D 40 1 <1 94 W558N 40 0.7 <1 94 L556A 0 1.2 2 0 2 .+-. 0.4 94 L556V 0 4.2 95 .+-. 32 77 94 L552N 82 2.4 200 94 L552E 82 2.1 110 57/58 F443A 12 3.6 174 .+-. 110 113 .+-. 60 57/58 F443V 12 2.26 95 A-aggregated *Accessible Surface Areas (ASA) were calculated for the wild type residue.
Example 4
[0372] This example demonstrates that the substitutions in PE domain III diminish the epitope and do not create new epitopes due to the mutation.
[0373] The response of three donor samples and one patient sample to 22 peptide pools after stimulation with either HA22 (wt) or LR-R494A was measured. "LR" denotes the deletion of all of domain II except the furin cleavage sequence. The results are shown in FIGS. 2A-2D. As shown in FIGS. 2A-2D, the epitope in peptide 76 was diminished by the R494A mutation.
[0374] The response of two donor samples and one patient sample to 22 peptide pools after stimulation with either HA22 (wt) or LR-R505A was measured. The results are shown in FIGS. 3A-3C. As shown in FIGS. 3A-3C, the epitope in peptide 77 was diminished by the R505A mutation.
[0375] The response of two donors to 22 peptide pools after stimulation with either HA22-LR (WT) or HA22-R551A and restimulation with the appropriate peptides was also measured. The results are shown in FIGS. 4A-4B. As shown in FIGS. 4A and 4B, the mutation R551A diminished the epitope in peptide 93.
[0376] The response of cells from two donors and two patients after stimulation with RIT and restimulation with peptides 93, 94 with either the WT amino acid sequence, L552E or L552N was measured. The results are shown in FIGS. 5A-5D. As shown in FIGS. 5A-5D, the epitope in peptides 93 and 94 was diminished by the L552N and L552E mutations.
[0377] The response of three donors to 22 peptide pools after stimulation with either HA22-LR (WT) or LR-R427A and restimulation with the appropriate peptides was also measured. The results are shown in FIGS. 6A-6C. As shown in FIGS. 6A-6C, the epitope in peptide 51 was diminished by the R427A mutation.
[0378] The response of two patients and one donor to pools 8-22 after stimulation with either HA22-LR (WT) or LR-F443A and restimulation with the appropriate peptides was measured. The results are shown in FIGS. 7A-7C. As shown in FIGS. 7A-7C, the epitope in pools 11 and 12 was diminished by the F443A mutation.
[0379] Four donor samples were stimulated with RIT and restimulated with peptide 67 containing a valine or alanine mutation at position 1471. The response is shown in FIGS. 9A-9D in Spot Forming Cells (SFC)/10.sup.6 cells. As shown in FIGS. 9A-9D, mutation 1471V did not diminish the epitope in peptide 67.
[0380] T cell activation as a response to stimulation with peptide 67 or 68 that contained an alanine or histidine mutation at position L477 or no mutation (WT) was also measured. The results (SFC/10.sup.6 cells) are shown in Table 9 and FIGS. 10A-10D. As shown in FIGS. 10A-10D and Table 9, the mutation L477H diminished the epitope in peptide 67 and 68.
TABLE-US-00010 TABLE 9 hcl meso 11410 41311 102909 021012 031612 Mean WT 67 100 100 100 100 100 100 67 L477A 81 21 24 133 39 60 67 L477H 0 30 123 65 23 48 WT 68 100 100 100 100 100 100 68 L477A 69 19 31 62 38 44 68 L477H 2 34 46 32 48 32
[0381] Cells from two patients were stimulated with RIT and restimulated with peptide 93, 94 or 95 with either the WT amino acid sequence or L556V. The results are shown in FIGS. 11A and B. Mutation L477V did not diminish the epitope in peptide 94.
Example 5
[0382] This example demonstrates the activity of combinations of mutations in HA22-LR.
[0383] To combine the point mutations that were effective in diminishing the T cell epitopes and maintain good cytotoxic activity, genes were designed that contain different combinations of point mutations. The genes were cloned into the HA22-LR RIT plasmid. Table 10 shows the seven RITs that were constructed and also HA22-LR WT. Table 10 also shows the cytotoxic activity in two cell lines and the calculated remaining T cell response. The calculated remaining response is the sum of the responses shown in the epitope heat map for each of the epitopes, assuming that the mutation will eliminate the responses in the epitope completely. This calculation does not take into account the strength of the response.
[0384] As shown in Table 10, out of the 258 responses identified in the donor screen of PE38, 136 responses were in domain III and are present in HA22-LR. Elimination of the epitopes in peptide (77-78), (51-52) and (57-58) with mutations R505A, R427A and F443A (HA22-LR-T2) reduced the calculated responses to 82 responses and maintained a very active protein with 280% and 135% relative activity in CA46 and Raji cells, respectively, compared to WT. HA22-LR-T3 contains four mutations: R505A, R427A, F443A and R494A and had diminished activity of 55% and 50%. HA22-LR-T7 also has four mutations, (R505A, R427A, F443A and R551A) and had good cytotoxic activity with 113% and 100% relative activity in CA46 and Raji cells. For the construction of HA22-LR-T5, five mutations, R505A, R494A, L427A, R551A and F443A, were inserted which reduced the calculated remaining response to 39 responses. However, the protein's cytotoxic activity was severely reduced to 24% and 19% relative activity with EC.sub.50>1 ng/ml. HA22-LR-T9 also had five point mutations (R505A, R427A, F443A, R494A and L477A). T9 had 46 calculated remaining responses; all 46 remaining responses were weak or medium). It maintained cytotoxic activity with EC.sub.50<1 ng/ml with relative activity of 36% and 26% in CA46 and Raji cells, respectively. HA22-LR-T11 substitutes L477A in T9 with L477H which improved that activity to 0.2 ng/ml with 85% relative activity in CA46. Lastly, T18 (also referred to as "LMB-T18") was the first RIT that contains six mutations and diminished all the major epitopes that were identified. It had good relative activity with IC50<0.3 ng/ml and relative activity of 63% and 104% compared to HA22-LR in CA46 and Raji cells. To improve cytotoxic activity, a Gly-Gly-Ser peptide linker was inserted after the furin cleavage site.
TABLE-US-00011 TABLE 10 HA22- LR T2-HA T3-HA T5-HA T7-HA T9-HA T11-HA T18 RIT backbone HA22- HA22- HA22- HA22- HA22- HA22- HA22-LR LR LR LR LR LR LR R505A 77- + + + + + + + 78 R494A 75- + + + + + 76 L477A 67- + 68 L477H 67- + + 68 R427A 51- + + + + + + + 52 R551A 93- + + 94 L552E + F443A 57- + + + + + + + 58 Cytotoxic Activity CA46 IC50* 0.17 0.06 0.31 0.71 0.15 0.48 0.2 0.27 (ng/ml) CA46 Relative 100% 280% 55% 24% 113% 36% 85% 63% activity Raji IC50* 0.23 0.17 0.46 1.18 0.23 0.88 0.24 (ng/ml) Raji Relative 100% 135% 50% 19% 100% 26% 104% activity Calculated 136/258 82/258 62/258 39/258 59/258 46/258 46/258 19/258 Responses *IC50 values are an average of 2-9 assays, depending on the protein.
[0385] To characterize the cytotoxic activity properties of T3 and T18, WST-8 assays were performed in four cell lines that express CD22: ca46, Raji, Daudi and HAL-01. Each assay was performed three times. Table 11A shows the average EC.sub.50 values for each cell line. Both HA22-LR-T3 and HA22-LR-T18 were very cytotoxic, although somewhat less than the parent molecule HA22-LR-GGS. Compared with HA22-LR-GGS, HA22-LR-T18 showed relative activity that ranged between 28%-55% with IC.sub.50<1 ng/ml in the four cell lines. HA22-LR-T3 was also very active, however, less active than T18.
TABLE-US-00012 TABLE 11A Summary of EC.sub.50 and relative activity of HA22-LR-T3 and HA22-LR-T18 RITs HA22-LR- HA22-LR- Cell Line GGS HA22-LR-T3 T18 CA46 EC.sub.50.+-. 0.1 0.4 0.3 (ng/mL) SD .+-.0.003 .+-.0.08 .+-.0.01 Relative 100 18 28 activity (%) Raji EC.sub.50.+-. 0.1 0.3 0.2 (ng/mL) SD .+-.0.04 .+-.0.15 .+-.0.07 Relative 100 34 55 activity (%) Daudi EC.sub.50.+-. 0.1 0.4 0.3 (ng/mL) SD .+-.0.04 0.08 .+-.0.17 Relative 100 31 38 activity (%) HAL-01 EC.sub.50.+-. 0.3 1.5 0.7 (ng/mL) SD .+-.0.05 .+-.0.47 .+-.0.16 Relative 100 20 42 activity (%)
[0386] Cytotoxicity assays of LMB-T18 were performed on four CD22 expressing cell lines and compared with the cytotoxic activity of MP (FIG. 27A, Table 11B). LMB-T18 was very potent with an EC50<10 pM in all cell lines. Compared to MP, LMB-T18 had a small increase of 53% in activity in CA46 cells, 54% in Daudi cells and >200% in HAL-01 cells (p=0.2, 0.06 and 0.01 respectively in Student T test); however, in Raji cells LMB-T18 had a 52% activity decrease (p=0.3 in Student T test). Without being bound by a particular theory or mechanism, it is believed that the decrease in activity in Raji cells was probably due to the domain II deletion (Weldon et al., Blood, 113: 3792-3800 (2009)). The stability of LMB-T18 was compared with MP by heating samples for 15 minutes at various temperatures and, after cooling, measuring residual cytotoxic activities (FIG. 27B) on Raji cells. MP lost 50% of its activity after 15 minutes incubation at 56.degree. C. Unexpectedly, LMB-T18 was more heat-resistant (p<0.05 in Student T test); it only lost 50% of its activity after a 15-minute incubation at 70.degree. C. Cells from seven HCL and six chronic lymphocytic leukemia (CLL) patients were used to determine activity on patient cells. FIGS. 27C-D show that LMB-T18 is more active than MP on CLL cells though this difference is not seen with HCL cells.
TABLE-US-00013 TABLE 11B EC50 .+-. SD (pM) Relative activity Cell line MP LMB-T18 (%) P value CA46 (n = 4) 3.4 .+-. 1.7 2.2 .+-. 0.4 153 0.2 Raji (n = 4) 1.1 .+-. 0.3 2.3 .+-. 0.6 48 0.3 Daudi (n-4) 3.7 .+-. 1.6 2.4 .+-. 0.8 154 0.06 HAL-01 (n = 4) 25.7 .+-. 3 8.1 .+-. 1 318 0.01
Example 6
[0387] This example demonstrates the relative activity of B and T cell epitope modified RIT.
[0388] B cell epitopes in PE38 were previously identified and the bulky surface residues were mutated with alanine or serine to reduce antigenicity (Liu et al., Protein Eng. Dec. Sel., (25)(1): 1-6 (2012)). The previously identified mutant RIT with low antigenicity was named SS1LO10R and it contained the following mutations: R505A, R427A, R490A, R463A, R467A, R538A.
[0389] To make a minimally immunogenic RIT targeting mesothelin, the LO10 modifications were incorporated into SS1P and then the T cell epitope-point mutations were incorporated one by one. Cytotoxic activity was tested for each one of the variants using mesothelin expressing cell lines (A9/431 and HAY). Two mutations R505A and R427A were found to be effective in diminishing both B and T cell epitopes.
[0390] Table 12 summarizes the cytotoxic activity of the mutated PE in two cell lines and the calculated remaining response of the combination RIT that were constructed. Combination of SS1-LO10 with a single point mutation F443A (SS1-LO10-T2) had high (100%-115%) relative activity that corresponded to the activity of these point mutations in HA22-LR. Combination of R494A and R551A in SS1-LO10-T1 gave a very low activity indicating that the combination of R551A and R494A diminished protein activity in both HA22 and SS1-LO10.
TABLE-US-00014 TABLE 12 Relative activity of B and T cell epitope modified RIT Cytotoxic activity Cal- A431/H9 HAY cu- Rel- Rel- lated RIT R505A R494A L477A R427A R551A F443A IC50* ative IC50* ative re- Back- 77- 75- 67- 51- 93- 57- (ng/ activ- (ng/ activ- spon- bone 78 76 68 L477H 52 94 L552E L552N 58 R456A m) ity m) ity ses SS1 0.3 0.5 136/ LO10R 258 SS1 SS1 + + + + 1.6 19% 50 10% 52/ LO10R- LO10R 258 T1 SS1 SS1 + + + 0.17 176% 0.5 100% 82/ LO10R- LO10R 258 T2 SS1 SS1 + + + + 0.29 103% 0.6 83% 62/ LO10R- LO10R 258 T3 SS1 SS1 + + + + + 2 15% nd 62/ LO10R- LO10R 258 T4 SS1 SS1 + + + + 0.6 50% 2 50% 59/ LO10R- LO10R 258 T7 SS1 SS1 + + + + + 0.9 33% 2.5 20% 59/ LO10R- LO10R 258 T8 SS1 SS1 + + + + + 1.1 27% 6.9 7% 46/ LO10R- LO10R 258 T9 SS1 SS1 + + + + + 0.42 71% 7 7% 46/ LO10R- LO10R 258 T11 SS1 SS1 + + + + + + 0.23 130% 1.5 33% 19/ LO10R- LO10R 258 T14 SS1 SS1 + + + + + + 0.27 111% 1.9 26% 19/ LO10R- LO10R 258 T15 *IC50 values are an average of 1-9 assays, depending on the protein; nd-not done
[0391] SS1 LO10R-T11, which contained R505A, R427A, R494A, F443A and L477H, had 71% activity in H9 cells but only 7% in HAY cells (assay in HAY was done once). T14 and T15, which contained R505A, R427A, R494A, F443A and L477H, with an additional mutation (L552E and L552N, respectively) had >100% relative activity in H9 cells and IC50>1 ng/ml. These two mutants represent the maximal T cell deimmunization done in this project thus far.
[0392] Mutant T14 was selected as the best candidate for clinical development because it includes the removal of B cell and many T cell epitopes, yet provides good cytotoxic activity on mesothelin expressing cell lines.
Example 7
[0393] This example demonstrates the anti-tumor activity of T14.
[0394] Anti-tumor experiments were carried out by injecting saline (control) or T14 into nude mice with A431/H9 xenografts. Tumor size was measured every other day and reported in mm. Animals were weighed at the same time and the weight was reported in grams. The results are shown in Table 13 below. In Table 13, the arrows show the day immunotoxin was injected. As shown in Table 13, doses up to 7 mg/kg.times.3 were well tolerated with no signs of toxicity in the mice. In addition, major tumor regressions were seen at all dose levels of T14.
TABLE-US-00015 TABLE 13 Mouse Days Group # Day 5 .dwnarw. 7 .dwnarw. 9 .dwnarw. 12 .dwnarw. 14 .dwnarw. A 1 108 23 g 114 22 g 74 23 g 46 36 T14 2 100 22 g 100 22 g 56 22 g 40 64 5 mg mg/kgx5 3 97 20 g 92 20 g 31 20 g 0 10 iv QOD 4 101 19 g 94 19 g 60 20 g 52 74 (100 ug/mouse) 5 97 22 g 83 22 g 57 22 g 41 70 6 103 21 g 95 21 g 58 21 g 37 58 Mean 101 96 56 36 52 B 1 100 22 g 91 20 g 81 20 g 86 78 T 14 2 96 23 g 103 23 g 50 23 g 14 66 6 mg/kgx3 3 10021 g 112 21 g 91 22 g 54 88 iv QOD 4 10324 g 117 24 g 53 24 g 55 98 (120 ug/mouse) 5 102 21 g 121 20 g 62 20 g 82 125 Mean 100 109 67 58 91 C 1 100 21 g 98 21 g 36 22 g 53 22 g 69 7 mg/kgx3 2 101 20 g 126 19 g 44 20 g 65 21 g 103 iv GOD Mean 101 112 40 59 86 (140 ug/mouse) D 1 100 239 440 816 Control 2 95 133 234 612 saline 3 104 252 463 813 4 92 138 251 543 5 100 152 231 589 Mean 98 183 324 675 Over 800
Example 8
[0395] This example demonstrates the thermal stability of T18.
[0396] The stability of T18 was investigated by heating the RIT for 15 minutes at various temperatures. Cytotoxic assays were subsequently performed. EC50 of the different heated proteins were calculated. The results are shown in FIG. 8. The fold increase in EC50 is shown. As shown in FIG. 8, it was found that T18 maintained cytotoxic activity in 60.degree. C. and lost 50% of its activity at 68.degree. C. Leaver et al., Methods in Enzymology, 487: 545-74 (2011).
Example 9.1
[0397] This example demonstrates the construction, expression, and purification of anti-mesothelin deimmunized PE chimeric molecules with an elongated linker between the anti-mesothelin-Fab fragment heavy chain and the PE 24 domain and the removal of predicted T cell epitopes in the linker.
[0398] The linker sequence used in WO 2012/154530 and Weldon et al., Mol. Cancer Ther., 12: 48-57 (2013) to fuse the dsFv fragment (via the C-terminus of VH domain) to the deimmunized PE was also used to fuse a Fab fragment (via the C-terminus of CH1 domain) to the deimmunized PE. The resulting peptide region, including parts of the CH1 domain, the linker, and part of the the deimmunized PE (SEQ ID NO: 273) was examined by TEPITOPE analysis (Bian et al., Methods, 34: 468-475 (2004)). Potential T cell epitopes relevant for European (EU) and United States populations were identified within the region between the CH1 domain and the linker and the region between the linker and the PE24 domain. Potential T cell epitopes were characterized by a confidence number of 3 or lower. The results are shown in Table 14. As shown in Table 14, the linker region without the specific elongations (SEQ ID NO: 273) showed seven potential T-cell epitopes. One potential T-cell epitope was located in the transition region WEQLGGSPT (SEQ ID NO: 274), and the other six T-cell epitopes were located in the transition region VEPKSCKAS (SEQ ID NO: 275). Both of these regions are relevant mainly for European (EU) and Asian populations.
[0399] The potential T cell epitopes were destroyed by inserting an amino acid sequence (insert) in each of the transition regions, thereby providing an insert on each of both sides of the furin cleavage site. The sequence DKTH (SEQ ID NO: 276) was inserted into the linker at a location not directly adjacent to the furin cleavage sequence (FCS), and the sequence GGG (SEQ ID NO: 277) was inserted into the linker at a location directly adjacent to the FCS without potential T-cell epitopes. The resulting elongated peptide region, including a) parts of the CH1 domain, b) the linker elongated at the Fab fusion area and elongated at the PE fusion area, and c) part of the of the the deimmunized PE comprised amino acid SEQ ID NO: 278. The elongated peptide region was studied using TEPITOPE analysis, and the results are shown in Table 15A. As shown in Table 15A, the elongated linker did not generate an alert in the TEPITOPE analysis (confidence numbers of 6 or higher) and, at the same time, provided improved stability (see Example 9.2 below).
TABLE-US-00016 TABLE 14 Start Confi- Pro- An- f f(NE f(SE f(SW max Pos Sequence Score dence file chor Inhibitor Allele Pivot Pept. (EU) f(US) Asia) Asia) Asia) (f(all)) 26 WEQLGGSPT 0.3 3 ********* 0 0 HLA- 1 0 8.0% 0.6% 5.7% 0.1% 4.1% 8.0% (SEQ ID NO: 274) DRB1*0101 8 VEPKSCKAS 1.6 2 ***A***** 1 0 HLA- 1 0 0.1% 12.0% 2.6% 0.1% 0.8% 12.0% (SEQ ID NO: 275) DRB1*0802 8 VEPKSCKAS 2.6 2 ***A***** 1 0 HLA- 1 0 0.1% 0.1% 0.6% 0.0% 0.0% 0.6% (SEQ ID NO: 275) DRB1*0804 8 VEPKSCKAS 2.6 3 ***A***** 1 0 HLA- 1 0 0.1% 0.0% 0.1% 0.0% 0.0% 0.1% (SEQ ID NO: 275) DRB1*0806 8 VEPKSCKAS 1.9 3 ********* 0 0 HLA- 1 0 0.2% 0.0% 0.0% 0.0% 1.0% 1.0% (SEQ ID NO: 275) DRB1*1102 8 VEPKSCKAS 1.9 3 ********* 0 0 HLA- 1 0 N.A. N.A. N.A. N.A. N.A. 0.0% (SEQ ID NO: 275) DRB1*1121 8 VEPKSCKAS 1.9 3 ********* 0 0 HLA- 1 0 N.A. N.A. N.A. N.A. N.A. 0.0% (SEQ ID NO: 275) DRB1*1322 all Alerts listed <=3
TABLE-US-00017 TABLE 15A Start Confi- Pro- An- f(NE f(SE f(SW Pos Sequence Score dence file chors Inhibitors Allele Pivot Peptide f(EU) f(US) Asia) Asia) Asia) 30 WEQLGGGGG -1.2 9 ******I** 0 1 HLA-DRB1*0101 1 0 8.0% 0.6% 5.7% 0.1% 4.1% (SEQ ID NO: 279) 33 LGGGGGSPT -0.7 9 ********* 0 0 HLA-DRB1*0102 1 0 0.6% 0.1% 0.0% 0.0% 0.7% (SEQ ID NO: 280) 30 WEQLGGGGG(SEQ -1 9 ***A****I 1 1 HLA-DRB1*1307 1 0 0.0% 0.0% 0.0% 0.0% 0.0% ID NO: 279) 30 WEQLGGGGG(SEQ 1.4 6 ********* 0 0 HLA-DRB5*0101 1 0 N.A. N.A. N.A. N.A. N.A. ID NO: 279) 30 WEQLGGGGG(SEQ 1.4 6 ********* 0 0 HLA-DRB5*0105 1 0 N.A. N.A. N.A. N.A. N.A. ID NO: 279) no Alert <=3
Example 9.2
[0400] This example demonstrates the stability of the deimmunized PE fusion proteins and conjugates having the elongated linker of SEQ ID NO: 36.
[0401] Six Fab-Linker-PE24 variants were produced and compared with respect to their thermal stability and propensity for aggregation. The six variants all contained the same humanized Fab variant H1/L1 of the SS1 antibody, but differed with respect to the linker and the PE24 mutant. For the linker, either the linker described in International Patent Application Publication WO 2012/154530 (kasggrhrqprgweqlggs (SEQ ID NO: 281)) or the elongated linker with the additional amino acids ("long linker"; SEQ ID NO: 36 (dkthkasggrhrqprgweqlgggggs))) was used. As PE24 variants, LRO10R, LRO10R-456A (SEQ ID NO: 37) or LRO10R-456A-551A were employed in the constructs.
[0402] FIGS. 12-13 show the level of aggregation of the two constructs with the PE24 variant LRO10R-456A that was measured in terms of the area under the aggregate peak by size exclusion chromatography (SEC) after incubation at 33.degree. C. The two constructs in FIGS. 12-13 differed only with respect to the linker employed. The short linker SEQ ID NO: 281 was used in construct cFP-0170 and the elongated linker SEQ ID NO: 36 was used in construct cFP-0171.
[0403] FIGS. 14-16 show the thermal stability of constructs as measured by dynamic light scattering (DLS). For dynamic light scattering, samples were prepared at a concentration of 1 mg/mL in 20 mM histidine/histidine chloride, 140 mM NaCl, pH 6.0, transferred into an optical 384-well plate by centrifugation through a 0.4 .mu.m filter plate and covered with paraffin oil. The hydrodynamic radius was measured in a WYATT DYNAPRO Plate Reader II repeatedly by dynamic light scattering while the samples were heated at a rate of 0.05.degree. C./min from 25.degree. C. to 80.degree. C. The aggregation onset temperature was defined as the temperature at which the hydrodynamic radius starts to increase. The results are shown in FIGS. 14-16. The DLS curves for the six constructs showed higher aggregation temperatures for the elongated linker compared to the short published linker, with a strong shift to a higher aggregation temperature for the LRO10R variant (FIGS. 14-16).
[0404] Table 15B shows the thermal stability of constructs cFP-0170 and cFP-0171 as measured by differential scanning calorimetry (DSC). Differential scanning calorimetry was performed using a MicroCal DSC system (GE Healthcare). The samples were adjusted to a protein concentration of approximately 1 mg/ml in 20 mM histidine/HCl, 140 mM sodium chloride, pH 6.0. The reference cell was filled with a buffer corresponding to the sample buffer. The samples were placed in the sample cell and heated from 4.degree. C. to 110.degree. C. at a heating rate of 60.degree. C./hour. The pre-scan was 15 minutes, the filtering period was 10 seconds, and the feedback mode/gain was set to passive. The midpoint of a thermal transition temperature (Tm, or thermal transition temperature) was obtained by analyzing the data using MicroCal DSC analysis software.
TABLE-US-00018 TABLE 15B T-max of the Fab Construct fragment part T-max of the PE24 part Humanized SS1 Fab long 73.6 41.2 linker-LRO10R, LRO10R- 456A Humanized SS1 Fab long 73.7 43.2 linker-LRO10R, LRO10R
[0405] As shown in FIGS. 12-16 and Table 15B, all chimeric Fab-PE molecules with the elongated linker provided a decreased tendency toward aggregation and a higher aggregation temperature as compared to chimeric Fab-PE molecules with a short linker that was not elongated.
Example 10
[0406] This example demonstrates the ability of deimmunized PE fusion proteins with an extended linker to kill tumor cells and inhibit cellular protein synthesis. This example also demonstrates the antigenicity of the deimmunized PE fusion proteins with an extended linker.
[0407] The potency of different Fab-PE24 variants with respect to the ability to inhibit protein synthesis was compared. A clone of the pancreatic cancer cell line ASPC-1 (ASPC-1 Luc) that was stably transfected with luciferase was used. The luciferase protein in these cells undergoes a high turnover so that the measurable activity drops markedly within 24 hours of inhibition of its resynthesis. Briefly, cells were seeded on 96 well plates. After overnight culture, different concentrations of the cFP variants were added to the medium, and cells were incubated for another 72 hours. At the end of the incubation period, the cells were lysed, and luciferase activity was determined with the STEADY-GLOW assay according to the manufacturer's instructions. In a typical experiment, an 80-90% reduction of luciferase activity was observed in lysates from treated cells as compared to lysates from untreated control cells. EC50 values, i.e., the concentrations that achieve half maximal effects, were determined based on a free four parameter fit. Table 16 compares the relative potencies in a representative experiment by setting the IC50 of huFabLRO10R to 1.
[0408] Untargeted PE24 (variant LR8M) only inhibited at concentrations >3.5 .mu.g/ml, while the previously described LRO10 variant fused to a humanized SS1 Fab fragment was .about.500 fold more potent on a molar basis. The LRO10R backmutation increased potency 10 to 20 fold (EC50 values 1-2 ng/ml versus 18 ng/ml). The 456A mutation had no adverse effect on potency, while the 551A mutation did reduce activity to the level of LRO10 (EC50 10-23 ng/ml). The new extended linker also had no negative impact on the cellular potency of the molecules in a comparison of any of the three pairs of constructs with the extended linker versus those with the previously described shorter linker.
[0409] Using cell viability assessed by CELLTITERGLOW assays, the cytotoxic potency of different Fab-PE24 variants using the same humanized SS1 targeting variant on two different tumor cell lines (A431 H9 and H596) was compared. Briefly, cells were seeded on 96 well plates. After overnight culture, different concentrations of the cFPs were added to the medium, and cells were incubated for 72 hours. At the end of the incubation period, cell viability was determined using a CELLTITERGLOW assay. Table 16 compares the relative potencies by setting the IC50 of huFabLRO10R to 1.
TABLE-US-00019 TABLE 16 Viability assay results AsPC-1 Luciferase A431 H9 H596 Luc assay results Fold potency reduction cFP variant relative to huFabLRO10R huFabLRO10 1.8 2.5 17.5 LR8M 400 450 9452 huFabLRO10R 1 1 1 huFabLRO10R- 1.2 1 2.3 long linker huFabLRO10R- 0.8 1 2.1 456A huFabLRO10R- 0.9 1.2 3 456A- longlinker huFabLRO10R- 2.1 3.3 22.5 456A-551A huFabLRO10R- 2.1 4.5 9.8 456A-551A- longlinker
[0410] Results similar to those in Table 16 were also obtained using a cell viability assay with an 3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide (MTT)-based read-out in different cell lines (Table 17).
TABLE-US-00020 TABLE 17 IC 50s in ng/ml (did not correct for MW differences) Hu-1 Hu2 Hu3 HAY AGS L55 SS1P 2 0.3 2.9 1.0 0.3 3.0 SS1-LR-GGS 0.25 0.01 0.7 -- -- 1.0 SS1-LR-GGS-010R -- 0.01 -- 0.3 1.0 2.0 Fab-LR-GGS-010R Short 0.25 0.08 0.4 2.5 0.6 3.5 Fab-LR-GGS-010R 456A Short 0.25 0.05 0.6 2.5 0.6 3.0 Fab-LR-GGS-010R long 0.35 0.04 1.8 2.5 0.9 5.0 Fab-LR-GGS-010R 456A long 0.25 0.04 2.0 1.4 1.1 5.0 Fab-LR-GGS-010R 456A 0.7 0.18 4.2 4.0 6.0 20.0 551A Short Fab-LR-GGS-010R 456A Also Also Also Also Also Also 551A long low low low low low low
Example 11
[0411] This example demonstrates the removal of B-cell epitopes by 456A mutations in PE.
[0412] A phagemid containing the scFv 9H3 in the vector pCANTAB as described by Liu et al., PNAS, 109: 11782-11787 (2012) was prepared by standard methods as described in Sambrook, J. et al., Molecular cloning: A laboratory manual; Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 1989. The molecular biological reagents were used according to the manufacturer's instructions. The phagemid was transformed into electrocompetent TG1 cells. From single colonies, precultures were grown overnight and used to inoculate 4 ml cultures of 2.times.YT/1% glucose supplemented with the appropriate selective antibiotic ampicillin. At an OD.sub.600 of 0.3-0.8, 10.sup.10 cfu VCSM 13 helper phage and IPTG was added to a final concentration of 1 mM. After incubation for 15 minutes at 37.degree. C., the culture was grown at 37.degree. C. overnight. Two hours after helper phage addition, kanamycin was added to final concentration of 30 .mu.g/ml. The phage-containing supernatant was harvested from the overnight culture by centrifugation, and the scFv displaying phage were precipitated from the supernatant by addition of 1/5 v/v 20% PEG-6000/2.5 M NaCl and isolated by centrifugation. The phage pellet was dissolved in 0.5 ml PBS/1% BSA, and the phage solution was cleared by centrifugation and sterile filtration. The titer of the phage solution was determined by standard methods as described in Barbas, C. et al., Phage Display--A laboratory manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 2001.
[0413] 96 well MaxiSorp microtiter plates were coated with 100 .mu.g/ml Fab-PE24 variants. After blocking with PBS buffer supplemented with 2% BSA, 10.sup.7 and 108 scFv-displaying phage were captured on the plate for one hour. As specific negative controls, scFv-displaying phages were pre-incubated with 50 .mu.g/m of the Fab-PE24 variants for 1 hour at room temperature (RT) before capturing them on the plate. Plates were washed with PBST buffer (PBS+0.5% Tween 20). Binding of scFv-displaying phage was detected with POD conjugated anti-m13 antibody (Progen). After a final wash, the plates were incubated with POD substrate. Absorbance was measured at 405 nm. Results are shown in FIG. 17, where it can be seen that the B-cell epitope contained in the phagemid containing the scFv 9H3 was not removed in LRO10R (with 458R backmutation) while it was eliminated by both mutations 458A and 456A with a slightly stronger effect for the 456A mutation.
[0414] Antigenicity testing was done as described in Weldon et al., Mol. Cancer Ther., 12: 48-57 (2013). As shown in FIG. 18, compared to SS1P (T1), the de-immunized variant LRO10R (T2) had a 100-10,000 fold reduced reactivity with most sera from patients that have been previously exposed to a PE-based therapeutic fusion protein both with the murine dsFv fusion format and the humanized Fab fusion format. Also the introduction of the 456A mutation (T3 and T5) or the new extended linker (T5 and T6) showed a 100-10,000 fold reduced reactivity with most sera from patients that have been previously exposed to a PE-based therapeutic fusion protein.
Example 12
[0415] This example demonstrates the construction, expression and purification of deimmunized PE variants and Fab fusions thereof as well as dsFv fusions for comparison.
Example 12.1: Cloning of Anti-Mesothelin Deimmunized PE Chimeric Molecules cFP-0077 (SEQ ID NOs. 41 and 42) and cFP-0078 (SEQ ID NOs: 43 and 44)
[0416] For the expression of cFP-0077 and cFP-0078, the E. coli host/vector system, which enables an antibiotic-free plasmid selection by complementation of an E. coli auxotrophy (PyrF), was employed (European Patent Application Publication 0 972 838 and U.S. Pat. No. 6,291,245). Standard methods were used to manipulate DNA as described in Sambrook, J. et al., supra. The molecular biological reagents were used according to the manufacturer's instructions. Desired gene segments were made by commercial gene synthesis. The synthesized gene fragments were cloned into a specified expression vector. The DNA sequences of the subcloned gene fragments were confirmed by DNA sequencing.
[0417] The expression plasmids for the production of the light chain and the heavy chain, respectively, were prepared as follows: The light chain (LC) plasmids 15478 (based on the amino acid sequences of cFP-0077 and encoding SEQ ID NO: 41) and 15479 (based on the amino acid sequences of cFP-0078 and encoding SEQ ID NO: 43) are expression plasmids for the expression of an antibody light chain fragment in E. coli. They were generated by ligating the antibody VL domain fragment into the vector using the XhoI/BsiWI restriction sites for 15478, and XhoI/BlpI for 15479.
[0418] The light chain E. coli expression plasmids comprise the following elements: [0419] the origin of replication from the vector pBR322 for replication in E. coli (corresponding to positions 2517-3160 according to Sutcliffe et al., Quant. Biol., 43: 77-90 (1979), [0420] the URA3 gene of Saccharomyces cerevisiae coding for orotidine 5'-phosphate decarboxylase (Rose et al., Gene, 29: 113-124 (1984) which allows plasmid selection by complementation of E. coli pyrF mutant strains (uracil auxotrophy), [0421] the antibody light chain expression cassette comprising: [0422] the T5 hybrid promoter (T5-PN25/03/04 hybrid promoter according to Bujard et al., Methods. Enzymol., 155: 416-433 (1987) and Stueber, et al., Immunol. Methods IV, 121-152 (1990) including a synthetic ribosomal binding site according to Stueber et al. (supra), [0423] the antibody light chain variable domain comprising the CDRs of SS1, and [0424] the human Ck domain for 15478 but not for 15479, [0425] two bacteriophage-derived transcription terminators, the .lamda.-T0 terminator (Schwarz et al., Nature, 272: 410-414 (1978) and the fd-terminator (Beck et al., Gene, 1-3: 35-58 (1981), and [0426] the lad repressor gene from E. coli (Farabaugh, Nature, 274: 765-769 (1978).
[0427] The heavy chain (HC) plasmids 15476 (based on the amino acid sequences of cFP-0077 and encoding SEQ ID NO: 42) and 15477 (based on the amino acid sequences of cFP-0078 and encoding SEQ ID NO: 44) are expression plasmids for the expression of a fusion protein including an antibody heavy chain fragment, a linker containing a furin cleavage site, and a mutant of domain III of Pseudomonas aeruginosa Exotoxin A (PE), in E. coli. It was generated by ligation of the VH domain fragment into the vector using the XhoI/BsrGI restriction sites for 15476, and XhoI/HindIII restriction sites for 15477.
[0428] The cytolytic fusion protein heavy chain E. coli expression plasmids comprise the following elements: [0429] the origin of replication from the vector pBR322 for replication in E. coli (corresponding to positions 2517-3160 according to Sutcliffe et al., Quant. Biol, 43: 77-90 1979), [0430] the URA3 gene of Saccharomyces cerevisiae coding for orotidine 5'-phosphate decarboxylase (Rose et al., Gene, 29: 113-124 (1984) which allows plasmid selection by complementation of E. coli pyrF mutant strains (uracil auxotrophy), [0431] the heavy chain-PE domain III fusion protein expression cassette comprising [0432] the T5 hybrid promoter (T5-PN25/03/04 hybrid promoter according to Bujard et al., Methods. Enzymol., 155: 416-433 (1987) and Stueber et al., Immunol. Methods IV, (1990) 121-152) including a synthetic ribosomal binding site according to Stueber et al. (supra), [0433] the antibody heavy chain variable domain comprising the CDRs of SS1, [0434] the human CH1 domain for 15476 but not for 15477, [0435] the linker comprising a furin cleavage site [0436] the mutant variant of Pseudomonas aeruginosa Exotoxin A domain III [0437] two bacteriophage-derived transcription terminators, the .lamda.-T0 terminator (Schwarz et al., Nature, 272: 410-414 (1978) and the fd-terminator (Beck et al., Gene, 1-3: 35-58 (1981), and [0438] the lad repressor gene from E. coli (Farabaugh, Nature, 274: 765-769 (1978).
Example 12.2: Expression of Anti-Mesothelin Deimmunized PE Chimeric Molecules cFP-0077 and cFP-0078 in E. coli
[0439] All polypeptide chains were expressed separately in the E. coli strain CSPZ-6.
[0440] The E. coli K12 strain CSPZ-6 (thi-1, .DELTA.pyrF) was transformed by electroporation with the expression plasmids 15476, 15477, 15478 and 15479, respectively, resulting in the 4 strains cFP-0019 (based on plasmid 15476), cFP-0020 (based on plasmid 15477), cFP-0040 (based on plasmid 15478) and cFP-0041 (based on plasmid 15479). For each of the four strains, the transformed E. coli cells were first grown at 37.degree. C. on agar plates. A colony picked from this plate was transferred to a 3 mL roller culture and grown at 37.degree. C. to an optical density of 1-2 (measured at 578 nm). Then 1000 .mu.l of culture was mixed with 1000 .mu.l sterile 86%-glycerol and immediately frozen at -80.degree. C. for long time storage. The correct product expression of this clone was first verified in small scale shake flask experiments and analyzed with SDS-Page prior to the transfer to the 10 L fermenter.
Pre Cultivation:
[0441] For pre-fermentation, a chemically defined medium (CD-PCMv2.20) was used: NH.sub.4Cl 1.0 g/l, K.sub.2HPO.sub.4*3H.sub.2O 18.3 g/l, citrate 1.6 g/l, Glycine 0.78 g/l, L-Alanine 0.29 g/l, L-Arginine 0.41 g/l, L-Asparagine*H.sub.2O 0.37 g/l, L-Aspartate 0.05 g/l, L-Cysteine*HCl*H.sub.2O 0.05 g/l, L-Histidine 0.05 g/l, L-Isoleucine 0.31 g/l, L-Leucine 0.38 g/l, L-Lysine*HCl 0.40 g/l, L-Methionine 0.27 g/l, L-Phenylalanine 0.43 g/l, L-Proline 0.36 g/l, L-Serine 0.15 g/l, L-Threonine 0.40 g/l, L-Tryptophan 0.07 g/l, L-Valine 0.33 g/l, L-Tyrosine 0.51 g/l, L-Glutamine 0.12 g/l, Na-L-Glutamate*H.sub.2O 0.82 g/l, Glucose*H.sub.2O 6.0 g/l, trace elements solution 0.5 ml/l, MgSO.sub.4*7H2O 0.86 g/l, and Thiamin*HCl 17.5 mg/l. The trace elements solution contained FeSO.sub.4*7H.sub.2O 10.0 g/l, ZnSO.sub.4*7H.sub.2O 2.25 g/l, MnSO.sub.4*H.sub.2O 2.13 g/l, H.sub.3BO.sub.3 0.50 g/l, (NH.sub.4)6Mo.sub.7O.sub.24*4H.sub.2O 0.3 g/l, CoCl.sub.2*6H.sub.2O 0.42 g/l, and CuSO.sub.4*5H.sub.2O 1.0 g/l dissolved in 0.5M HCl.
[0442] For pre-fermentation, 220 ml of CD-PCMv2.20 medium in a 1000 ml Erlenmeyer-flask with four baffles was inoculated with 1.0 ml out of a research seed bank ampoule. The cultivation was performed on a rotary shaker for 8 hours at 32.degree. C. and 170 rpm until an optical density (578 nm) of 2.9 was obtained. 100 ml of the pre cultivation was used to inoculate the batch medium of the 10 L bioreactor.
Fermentation of cFP-0019 and cFP-0020:
[0443] For fermentation in a 101 Biostat C, DCU3 fermenter (Sartorius, Melsungen, Germany) the following chemically defined batch medium was used: KH.sub.2PO.sub.4 1.59 g/l, (NH.sub.4)2HPO.sub.4 7.45 g/l, K.sub.2HPO.sub.4*3H.sub.2O 13.32 g/l, citrate 2.07 g/l, L-methionine 1.22 g/l, NaHCO.sub.3 0.82 g/l, trace elements solution 7.3 ml/l, MgSO.sub.4*7 H2O 0.99 g/l, thiamine*HCl 20.9 mg/l, glucose*H.sub.2O 29.3 g/l, biotin 0.2 mg/l, and 1.2 ml/l Synperonic 10% anti foam agent. The trace elements solution contained FeSO.sub.4*7H.sub.2O 10 g/l, ZnSO.sub.4*7H.sub.2O 2.25 g/l, MnSO.sub.4*H.sub.2O 2.13 g/l, CuSO.sub.4*5H.sub.2O 1.0 g/l, CoCl.sub.2*6H.sub.2O 0.42 g/l, (NH.sub.4)6Mo.sub.7O.sub.24*4H.sub.2O 0.3 g/l, and H.sub.3BO.sub.3 0.50 g/l solubilized in 0.5M HCl solution.
[0444] The feed 1 solution contained 700 g/l glucose*H.sub.2O, 7.4 g/l MgSO.sub.4*7 H2O and 0.1 g/l FeSO.sub.4*7H.sub.2O. Feed 2 comprises KH.sub.2PO.sub.4 52.7 g/l, K.sub.2HPO.sub.4*3H.sub.2O 139.9 g/1l and (NH.sub.4)2HPO.sub.4 66.0 g/l. All components were dissolved in deionized water. The alkaline solution for pH regulation was an aqueous 12.5% (w/v) NH.sub.3 solution supplemented with 11.25 g/l L-methionine.
[0445] Starting with 4.2 l sterile batch medium plus 100 ml inoculum from the pre-cultivation, the batch fermentation was performed at 31.degree. C., pH 6.9.+-.0.2, 800 mbar back pressure and an initial aeration rate of 10 l/min. The relative value of dissolved oxygen (pO.sup.2) was kept at 50% throughout the fermentation by increasing the stirrer speed up to 1500 rpm. After the initially supplemented glucose was depleted, indicated by a steep increase in dissolved oxygen values, the temperature was shifted to 25.degree. C., and 15 minutes later the fermentation entered the fed-batch mode with the start of both feeds (60 and 14 g/h respectively). The rate of feed of 2 was kept constant, while the rate of feed 1 was increased stepwise with a predefined feeding profile from 60 to finally 160 g/h within 7 hours. When carbon dioxide off gas concentration leveled above 2%, the aeration rate was constantly increased from 10 to 20 l/min within 5 hours. The expression of recombinant protein was induced by the addition of 2.4 g IPTG at an optical density of approx. 120. The target protein was expressed as inclusion bodies within the cytoplasm.
[0446] After 24 hours of cultivation, an optical density of 209 was achieved, and the whole broth was cooled down to 4-8.degree. C. The bacteria were harvested via centrifugation with a flow-through centrifuge (13,000 rpm, 13 l/h) and the obtained biomass was stored at -20.degree. C. until further processing (cell disruption).
Fermentation of cFP-0040 and cFP-0041:
[0447] For fermentation in a 10 l Biostat C, DCU3 fermenter (Sartorius, Melsungen, Germany) the following complex batch medium was used: Bacto-Trypton 20 g/l, yeast extract 15 g/l, KH.sub.2PO.sub.4 1.5 g/l, K2HPO4*3H2O 6.6 g/l, NaCl 1.0 g/l, MgSO.sub.4*7 H2O 0.74 g/1, glucose*H.sub.2O 3.0 g/l, and 0.2 ml/l Synperonic 10% anti foam agent.
[0448] The feed 1 solution contained Bacto-Trypton 250 g/l and yeast extract 175 g/l. pH was controlled using a 75% glucose*H.sub.2O. All components were dissolved in deionized water.
[0449] Starting with 7.6 l sterile batch medium plus 100 ml inoculum from the pre cultivation, the batch fermentation was performed at 37.degree. C., pH 7.0.+-.0.3, 500 mbar back pressure and an initial aeration rate of 10 l/min. The relative value of dissolved oxygen (pO.sup.2) was kept at 50% throughout the fermentation by increasing the stirrer speed up to 1500 rpm. The feeding of feed 1 was started after the culture reached an optical density of 15. When carbon dioxide off gas concentration leveled above 2%, the aeration rate was constantly increased from 10 to 20 l/min within 5 hours. The expression of recombinant protein was induced by the addition of 2.4 g IPTG at an optical density of approx. 20. The target protein was expressed insoluble to inclusion bodies within the cytoplasm.
[0450] After 13 hours of cultivation, an optical density of 100-115 was achieved and the whole broth was cooled down to 4-8.degree. C. The bacteria were harvested via centrifugation with a flow-through centrifuge (13,000 rpm, 13 l/h) and the obtained biomass was stored at -20.degree. C. until further processing (cell disruption and inclusion body preparation).
Analysis of Product Formation (for cFP-0019, -0020-0040 and -0041):
[0451] Samples were drawn from the fermenter, one prior to induction and the others at dedicated time points after induction of protein expression, were analyzed with SDS-polyacrylamide gel electrophoresis. From every sample, the same amount of cells (OD.sub.Target=10) were suspended in 5 mL PBS buffer and disrupted via sonication on ice. Then 100 .mu.L of each suspension were centrifuged (15,000 rpm, 5 minutes), and each supernatant was withdrawn and transferred to a separate vial. This was to discriminate between soluble and insoluble expressed target protein. To each supernatant (=soluble protein fraction) 100 .mu.L and to each pellet (=insoluble protein fraction) 200 .mu.L of SDS sample buffer (Lacmmli, Nature, 227: 680-685 (1970) were added. Samples were heated for 15 minutes at 95.degree. C. under intense mixing to solubilize and reduce all proteins in the samples. After cooling to room temperature, 5 .mu.L of each sample were transferred to a 4-20% TGX Criterion Stain Free polyacrylamide gel (Bio-Rad). Additionally, 5 .mu.l molecular weight standard (Precision Plus Protein Standard, Bio-Rad) were applied.
[0452] The electrophoresis was run for 60 minutes at 200 V and thereafter the gel was transferred the GELDOC EZ Imager (Bio-Rad) and processed for 5 minutes with UV radiation. Gel images were analyzed using IMAGE LAB analysis software (Bio-Rad). Relative quantification of protein expression was done by comparing the volume of the product bands to the volume of the 25 kDa band of the molecular weight standard.
Inclusion Body Preparation:
[0453] The inclusion body preparations (IBP) of the 10 L fermentations were started directly after the harvest of the bacteria with the re-suspension of the harvested bacteria cells in buffer 1 (12.1 g/l Tris, 0.246 g/l MgSO4*7H2O, 12 ml/l 25%-HCl). The buffer volume was calculated in dependence of the dry matter content of the biomass. Lysozyme (100 kU/mg, 0.12 mg/g DCW) and a small amount of benzonase (5 U/g DCW) were added. Then the suspension was homogenized at 900 bar (APV Rannie 5, 1 pass) to disrupt the bacteria cells followed by the addition of further benzonase (30 U/gDCW) and an incubation for 30 minutes at 30-37.degree. C. Then the first wash buffer (60 g/l Brij, 87.6 g/l NaCl, 22.5 g/l EDTA, 6 ml/l 10N NaOH) was added, and again the suspension was incubated for 30 minutes. The following centrifugation step (BP12, Sorvall) led to the inclusion body slurry, which was re-suspended in the second wash buffer (12.1 g/l Tris, 7.4 g/l EDTA, 11 ml/l 25%-HCl) and incubated for 20 minutes. A further separation step harvested the inclusion bodies, which were stored frozen at -20.degree. C. or immediately solubilized for refolding and purification.
Example 12.3: Refolding and Purification of Anti-Mesothelin Deimmunized PE Chimeric Molecules cFP-0077 and cFP-0078
[0454] cFP-0077 and cFP-0078 were both obtained by refolding and purification.
[0455] Renaturation and Purification of Fab-PE24 (cFP-0077):
[0456] Inclusion bodies of HC-PE24 and LC were solubilized separately in 8 M Guanidinium-Hydrochloride, 100 mM Tris/HCl, 1 mM EDTA, pH 8.0.+-.100 mM Dithiothreitol (DTT) overnight at RT (1 g IB in 5 mL). Solubilizates were adjusted to pH 3 and centrifuged, and the pellet was discarded. After extensive dialysis against 8 M Guanidinium-HCl, 10 mM EDTA, pH 3.0 to remove DTT, the total protein concentration was determined using the Biuret method. The purity of HC and LC content was estimated via SDS-PAGE.
[0457] Solubilizates were diluted at a 1:1 molar ratio in renaturation buffer containing 0.5 M arginine, 2 mM EDTA, pH 10.+-.0.9 mM GSH/GSSG, respectively, at 2-10.degree. C. The target protein concentration was increased in two steps to 0.22 g/L, with an incubation time of 4 h between the two doses. Afterwards, the renaturation solution was kept at 2-10.degree. C. overnight.
[0458] The renaturate was diluted with H.sub.2O to <3 mS/cm and pumped onto an anion exchange column (AIEX) equilibrated in 20 mM Tris/HCl, pH 7.4. After washing the column with equilibration buffer, the protein was eluted with a gradient up to 20 mM Tris/HCl, 400 mM NaCl, pH 7.4. Peak fractions containing Fab-PE24 were pooled, concentrated and applied onto a preparative Size Exclusion Column (SEC) in 20 mM Tris, 150 mM NaCl, pH 7.4 to remove aggregates, fragments and E. coli proteins. The final protein pool was adjusted to the required protein concentration and analyzed via SDS-PAGE, analytical SEC and UV.sub.280, and identity was confirmed by mass spectrometry.
[0459] Renaturation and Purification of dsFv-PE24 (cFP-0078):
[0460] Inclusion bodies of HC-PE24 and LC were solubilized separately 8 M Guanidinium-Hydrochloride, 100 mM Tris/HCl, 1 mM EDTA, pH 8.5.+-.100 mM DTT overnight at RT (1 g IB in 5 mL). Solubilisates were adjusted to pH 3 and centrifuged, and the pellet was discarded. After extensive dialysis against 7.2 M Guanidinium-HCl, 10 mM EDTA, pH 3.0 to remove DTT, the total protein concentration was determined using the Biuret method, and the purity of HC and LC content was estimated via SDS-PAGE.
[0461] Solubilizates were diluted 1:100 in renaturation buffer containing 0.5 M arginine, 2 mM EDTA, pH 10.+-.0.9 mM GSH/GSSG, respectively, at a 1:1 molar ratio and kept overnight at 2-10.degree. C.
[0462] The renaturate was diluted with H.sub.2O to <3 mS/cm and pumped onto an anion exchange column (AIEX) equilibrated in 20 mM Tris/HCl, 1 mM EDTA, pH 7.4. After washing the column with equilibration buffer, the protein was eluted with a gradient up to 20 mM Tris/HCl, 1 mM EDTA, 400 mM NaCl, pH 7.4. Peak fractions containing dsFv-PE24 were pooled, concentrated and applied onto a preparative Size Exclusion Column (SEC) in 20 mM Tris/HCl, 150 mM NaCl, pH 7.4 to remove aggregates, fragments and E. coli proteins. The final protein pool was adjusted to the required protein concentration and analyzed via CE-SDS, analytical SEC and UV.sub.280. Identity was confirmed by mass spectrometry.
Example 12.4: Protein Analysis of cFP-0077 and cFP-0078
[0463] Samples were analyzed by OD 280 nm using a UV spectrophotometer to determine the protein concentration in solution. The extinction coefficient required for this was calculated from the amino acid sequence according to Pace et al., Protein Science, 4: 2411-2423 (1995). Size-exclusion chromatography (SE-HPLC) was performed on TSK-Gel300SWXL or Superdex 200 columns with a 0.2 M potassium phosphate buffer, comprising 0.25 M KCl, pH 7.0 as mobile phase in order to determine the content of monomeric, aggregated and degraded species in the samples. Sodium dodecyl sulfate (SDS) polyacrylamide gel electrophoresis (reducing and non-reducing) was performed to analyze the purity of the complex preparations with regard to product-related degradation products and unrelated impurities (for details, see below). Electrospray ionisation mass spectrometry (ESI-MS) was performed with reduced (TCEP) samples to confirm the correct mass/identity of each chain and detect chemical modifications. ESI-MS of the non-reduced samples was carried out to analyze the nature and quality of the fully assembled protein and detect potential product-related side products.
Method for SDS-PAGE and Coomassie Staining
Device: Invitrogen XCELL SURE LOCK Mini-Cell
Gel: 4-20% Tris-Glycine Gel, Invitrogen EC6025BOX
Buffer: Tris-Glycine SDS Running Buffer (10.times.), Invitrogen LC2675-5
[0464] Sample buffer: Tris-Glycine SDS Sample Buffer (2.times.), Invitrogen LC2676 Reducing buffer: NuPAGE Sample Reducing Agent (10.times.), lnvitrogen NP0004
Molecular Weight Marker: Mark 12, MW Standard, Invitrogen LC5677
[0465] The sample was adjusted to a protein concentration of 1 mg/ml with buffer. For sample reduction, the following procedure was carried out: A reduction buffer including 4 ml Sample buffer (2.times.) and 1 ml reducing buffer (10.times.) was prepared. The sample was diluted 1:1 with reduction buffer and incubated for 5 minutes at 70.degree. C.
[0466] The gel electrophoresis was carried out at 125 V for 90 minutes. The gels were stained with SIMPLY BLUE Safe Stain (Invitrogen, Cat. No. LC6065).
Example 12.5: Comparison of Fab Vs dsFv Fusion Protein-Cytotoxic Properties
[0467] The cytotoxic potency of a dsFv-PE24 fusion protein format and a Fab-PE24 format was compared using the same humanized SS1 targeting moiety on three different tumor cell lines (MKN45, H596, and A431H9). Briefly, cells were seeded on 96 well plates. After overnight culture, different concentrations of the cFPs were added to the medium and cells were incubated for 72 h. At the end of incubation period, cell viability was determined using a CELLTITERGLOW assay.
[0468] In each case, the dose-response curves and IC50 values for inhibition of cell viability were similar for the dsFv-PE24 and the Fab-PE24 format. The molar ratio of the IC50 values is listed in Table 18. Depending on the cell line tested, the targeted PE24 fusion proteins were several hundred to one thousand fold more potent than untargeted PE24.
TABLE-US-00021 TABLE 18 Ratio of the molar IC50 values Cell line dsFv-PE24/Fab-PE24 A431H9 0.37 MKN45 2.3 H596 0.38
Example 12.6: Comparison of Fab Versus dsFv Fusion Protein--Serum Kinetics
[0469] The serum kinetics of the following three Pseudomonas exotoxin A (PE)-based anti-mesothelin immunotoxins in SCID beige mice was compared: SS1dsFv-PE38 (MW: 62.5 kDa), SS1dsFv-PE24 (MW: 49.9 kDa) and SS1Fab-PE24 (MW: 72.2 kDa). The murine SS1 antibody moiety was used for all constructs. The kinetics were compared at doses equimolar to 0.2 mg/kg (3.2 nmol/kg) SS1P) corresponding to 0.231 mg/kg SS1Fab-PE24, and 0.160 mg/kg for SS1dsFv-PE24.
[0470] The constructs were administered as a single intravenous dose, and their serum levels were analyzed at 0.08, 0.5, 1, 1.5, 2, 3, 4, 5, and 7 hours after dosing. Nine mice per group were administered the study medication, and blood was collected in a staggered manner at three times per animal by retro-orbital puncture, resulting in nine time points with n=3 samples. Serum samples were analyzed by an ELISA that detects only intact test compounds having the antibody fragment plus the PE part. In this assay format, the cFP is captured on a MSLN-coated surface and the PE moiety is detected with a rabbit polyclonal anti-PE antibody and biotinylated anti-rabbit secondary antibody.
[0471] For each time point, average serum values from three animals were plotted for the treatment groups together with historical data from CD2F1 mice. The SS1dsFv-PE24 construct was very rapidly cleared with a serum half-life of only -12 minutes. In agreement with the historical data, clearance of SS1P was much slower, resulting in a serum half life of between 35 and 44 minutes (values derived from measured and historical data, respectively). The pharmacokinetic properties of the SS1Fab-PE24 construct were very similar to SS1dsFv-PE38 with a serum half life of 43 minutes and similar values were also obtained for clearance rate, volume of distribution, c.sub.max and area under the curve (Table 19 and FIG. 19).
TABLE-US-00022 TABLE 19 Comparison Fab versus dsFv fusion protein-Serum kinetics Parameter Unit SS1P SS1-Fab-PE24 SS1-dsFv-PE24 C.sub.L mL/(min kg) 2.24 1.53 7.06 V.sub.C mL/kg 84.3 75.4 146 V.sub.SS mL/kg 92 80.5 103 T.sub.1/2 h 0.574 0.716 0.201 C.sub.max ng/mL 1990 2730 837 AUC (0-inf) ngh/mL 1490 2520 378
Example 13
[0472] This example demonstrates the humanization of the anti-mesothelin antibody SS1.
Example 13.1: Design of Humanized Heavy and Light Chain Variable Regions
[0473] The structures of the VH and the VL domain of the SS1 antibody were modeled in silico, and the model was compared to a structural database of human VH and VL domains. A panel of the most structurally similar V domains were chosen for grafting the CDRs of SS1 onto the human VH and VL domains. In addition, similarities in the primary sequence were taken into account to narrow down the choice of the human V domains by aligning the primary sequence of the VH and VL domain of SS1 to the human V domain repertoire. Backmutations within the human framework regions to mouse parent residues were introduced in some humanization variants. Similarly, mutations in the CDRs were introduced in some variants, where appropriate, to potentially increase the affinity to the antigen or to maintain the CDR tertiary structure.
[0474] The designed V domain variants were cloned into heavy chain and light chain vectors 6318 and 6319 via unique cloning sites to generate human IgG light and heavy chains. The heavy and light chain vectors were co-transfected into HEK293 suspension cells in microtiter culture plates in a matrix manner to obtain cell cultures expressing full size IgG having all possible light/heavy chain combinations. After 5 days, cultivation at 37.degree. C., the supernatants were harvested and purified by Protein A affinity chromatography in the microtiter scale.
Example 13.2: Cloning of Humanized Heavy and Light Chain Variable Regions
[0475] Standard methods were used to manipulate DNA as described in Sambrook, J. et al., supra. The molecular biological reagents were used according to the manufacturer's instructions. Desired gene segments were prepared by commercial gene synthesis. The synthesized gene fragments were cloned into a specified expression vector. The DNA sequence of the subcloned gene fragments were confirmed by DNA sequencing.
[0476] Expression vector for the antibody heavy chains: The gene segments of the designed humanized antibody heavy chain variable domains were cloned into the specified expression vector via the unique restriction sites HindIII and XhoI. The expression vector was designed to express the antibody heavy chain variable domain in fusion with the human antibody domains CH1, hinge, CH2 and CH3 in HEK293 cells, resulting in a conventional antibody heavy chain. All domains were separated by introns. Besides the expression cassette for the antibody heavy chain, the vector contained: [0477] an origin of replication from the vector pUC 18 which allows replication of this plasmid in E. coli, [0478] a .beta.-lactamase gene which confers ampicillin resistance in E. coli, [0479] an SV40 promotor and origin for expression of the DHFR selection marker, [0480] Murine dihydrofolate reductase (DHFR) as selection marker for antibody expression, and [0481] SV40' early polyadenylation ("early poly A") signal sequence.
[0482] The transcription unit of the antibody heavy chain is composed of the following elements: [0483] the immediate early enhancer and promoter from the human cytomegalovirus, [0484] a 5'-untranslated region of a human antibody germline gene, [0485] a murine immunoglobulin heavy chain signal sequence including a signal sequence intron (signal sequence 1, intron, signal sequence 2 [L1-intron-L2]), [0486] the VH sequence followed by an intron, [0487] human CH1, hinge, CH2, CH3 domains separated by introns, [0488] the bovine growth hormone (bGH) polyadenylation ("poly A") signal sequence, and [0489] the human gastrin transcription terminator (HGT).
[0490] Expression Vector Fort he Antibody Light Chains:
[0491] The gene segments of the designed humanized antibody light chain variable domains were cloned into a specified expression vector via the unique restriction sites BsmI and Cell. The expression vector was designed to place the antibody light chain variable domain in fusion with the human antibody domain Ck in HEK293 cells. Both domains were separated by an intron. Besides the expression cassette for the antibody light chain, the vector contains: [0492] an origin of replication from the vector pUC 18 which allows replication of this plasmid in E. coli, and [0493] a .beta.-lactamase gene which confers ampicillin resistance in E. coli.
[0494] The transcription unit of the antibody light chain is composed of the following elements: [0495] the immediate early enhancer and promoter from the human cytomegalovirus, [0496] a 5'-untranslated region of the human cytomegalovirus, [0497] an Intron A sequence of the human cytomegalovirus, [0498] a 5'-untranslated region of a human antibody germline gene, [0499] a murine immunoglobulin heavy chain signal sequence including a signal sequence intron (signal sequence 1, intron, signal sequence 2 [L1-intron-L2]) and the unique restriction site BsmI at the 3' end of L2, [0500] a VL domain followed by an intron, [0501] human Ck domain, [0502] the bovine growth hormone (bGH) polyadenylation ("poly A") signal sequence, and [0503] Human gastrin transcription termination factor (HGT).
Example 13.3: Expression of Humanized Antibodies
[0504] Transient Transfection of Humanized Antibody Light and Heavy Chains:
[0505] Recombinant humanized antibody variants were generated by transient transfection of HEK293-Freestyle cells (human embryonic kidney cell line 293, Invitrogen) grown in suspension. The transfected cells were cultivated in F17 medium (Gibco) or Freestyle 293 medium (Invitrogen), either one supplemented with 6 mM glutamine, either ultra-glutamine (Biowhittake/Lonza) or L-glutamine (Sigma), with 8% CO.sub.2 at 37.degree. C. in shake flasks in the scale of 2 ml medium in 48-deep-well plates to 1 L medium in shake flasks. 293-Free transfection reagent (Novagen/Merck) was used in a ratio of reagent (l) to DNA (g) of 4:3. Light and heavy chains were expressed from two different plasmids using a molar ratio of light chain to heavy chain encoding plasmid ranging from 1:2 to 2:1, respectively. Humanized antibody containing cell culture supernatants were harvested at day 6 to 8 after transfection. General information regarding the recombinant expression of human immunoglobulins in, e.g., HEK293 cells, is given in: Meissner et al., Biotechnol. Bioeng., 75: 197-203 (2001). The supernatants were purified by affinity chromatography on MabSelect Sure (GE, Protein A) equilibrated in PBS. After application of the filtrated supernatants, the antibodies were eluted with 50 mM sodium acetate, pH 3.2. The pH of the eluates was adjusted immediately to pH>6 with 2 M Tris/HCl, pH 9.0 followed by dialysis into 20 mM histidine, 140 mM NaCl, pH 6.0. Antibodies were analyzed by UV.sub.280, SDS-PAGE and analytical SEC. The sequence was confirmed by mass spectrometry.
Example 13.4: Characterization of Humanized SS1 Variants
[0506] ELISA screening of humanized SS1 variants: All SS1 humanization variants in the IgG format were screened for affinity by ELISA. 384 well MaxiSorp microtiter plates were coated with 0.5 .mu.g/ml anti-His antibody (Novagen). After blocking with PBS buffer supplemented with 2% BSA, 0.1% Tween 20, and 0.2 .mu.g/ml human/cynomolgus/murine mesothelin (in house/R&D Systems), all proteins with a His-tag were captured on the plate for one hour. The plates were washed with PBST Buffer (PBS+0.1% Tween 20) and dilutions of humanized anti-mesothelin antibodies in PBS were incubated for 1 hour at room temperature. Binding of antibodies was detected with HRP conjugated anti human Fe antibody (GE Healthcare). After a final wash, the plates were incubated with HRP substrate. Absorbance was measured at 370 nm on an ENVISION plate reader. EC50 curve fit analysis was performed using XLfit4 analysis plug-in for EXCEL software (model 205).
[0507] To normalize the obtained values to the IgG titer, quantitation of human IgG was performed by sandwich ELISA using streptavidin coated plates. Streptavidin-coated 384 well microtiter plates (Microcoat) were incubated with a mixture of 0.25 g/ml biotinylated anti-human IgG antibody (Jackson Imm. Res.), 0.05 .mu.g/ml anti-human IgG-HRP conjugate (Jackson Imm. Res.), and dilutions of the humanized ant-mesothelin antibodies. After 1.5 hours incubation at room temperature (RT), plates were washed with PBS buffer supplemented with 0.1% Tween 20. HRP substrate was added, and the absorbance was measured at 370 nm on an ENVISION plate reader. The calculation of data was performed using a human IgG reference as a standard (in house) for calibration and XLfit4 analysis plug-in for EXCEL software (model 205) for curve fit analysis.
[0508] In Tables 20-21 below, the ELISA data of all humanization variants are shown. ELISA/BIACORE binding to huMesothelin (EC50 ng/ml) was measured, and the results are shown in Table 20 (in house) and Table 21 (R&D). Normalized to IgG titer, this primary screen identified the variant VL001/VH001 as one of the combinations with the best EC50 value.
[0509] BiaCore screening of humanized SS1 variants: The best variants containing CDRs most similar to SS1 were chosen for kinetics analysis by surface plasmon resonance (SPR). An SPR based assay has been used to determine the kinetic parameters of the binding between several MSLN PE cFP humanization variants and human mesothelin. Therefore, Protein A was immobilized by amine coupling to the surface of the CM5 biosensor chip. The samples were then captured, and human mesothelin was injected. The sensor chip surface was regenerated between each analysis cycle. The equilibrium constant K.sub.D as well as the rate constants k.sub.d and k.sub.a were finally gained by fitting the data to a 1:1 langmuir interaction model. About 175 response units (RU) of Protein A (10 g/ml) were coupled onto the CM5 sensor chip at pH 4.0 by using an amine coupling kit supplied by GE Healthcare (10 minutes activation). The sample and system buffer was HBS-P+ (0.01 M HEPES, 0.15 M NaCl, 0.005% surfactant P20 sterile-filtered, pH 7.4). The flow cell temperature was set to 25.degree. C., and the sample compartment temperature was set to 12.degree. C. The system was primed with running buffer. The protein A binding sites were saturated with IgGs (about 0.03 .mu.g/mL) to generate similar capture levels for each sample, by injecting them for 40 seconds at a flow rate of 10 .mu.l/min. Afterwards, a single 50 nM human mesothelin solution was injected for 120 seconds at a flow rate of 30 l/min, followed by a 180 second dissociation phase. Thereby a relative KD determination allowed a ranking of different MSLN binders. Each cycle was regenerated with two injections of glycine-HCl pH 1.5 (30 seconds, 30 l/min).
[0510] Table 22 below lists the BiaCore data of 12 selected humanization variants. The final humanization variant VL01/VH01 shows one of the lowest Kd values among all humanization variants. Table 22 shows the Biacore results for cFP.15438-15457 and cFP.15438-15459, all comprising humanized VH1. cFP.15438, which comprises the amino acid sequence of SEQ ID NO: 45, displays the highest affinity of the humanized variants of SS1 antibody. Thus, the VH of cFP.15438 provides advantageous binding properties.
[0511] The five best variants were cloned into the Fab-Linker-PE format containing the published linker and LRO10 as the PE24 variant, expressed, refolded and purified. Cloning, expression, refolding and purification were carried out as described below for cFP-0205. The five humanization variants in the cytolytic fusion format were analyzed for stability at 37.degree. C. for 7 days by the method of dynamic light scattering described in Example 9.2, size exclusion chromatography, SDS-PAGE, and mass spectrometry at distinct time points.
[0512] Size-exclusion chromatography (SE-HPLC) was performed on TSK-Gel300SWXL or Superdex 200 columns with a 0.2 M potassium phosphate buffer comprising 0.25 M KCl (pH 7.0) as the mobile phase in order to determine the content of monomeric, aggregated, and degraded species in the samples. Sodium dodecyl sulfate (SDS) polyacrylamide gel electrophoresis (reducing and non-reducing) was performed to analyze the purity of the complex preparations with regard to product-related degradation products and unrelated impurities (for details, see below). Electrospray ionisation mass spectrometry (ESI-MS) was performed with reduced (TCEP) samples to confirm the correct mass/identity of each chain and to detect chemical modifications. ESI-MS of the non-reduced samples was carried out to analyze the nature and quality of the fully assembled protein and to detect potential product-related side products.
[0513] The active concentration of the Fab fragments within the cytolytic fusion format after temperature stress at 37.degree. C. for 7 days was monitored by surface plasmon resonance (SPR). Mesothelin was immobilized onto the surface of a SPR biosensor. By injecting the sample into the flow cells of the SPR spectrometer, it formed a complex with the immobilized mesothelin, resulting in an increased mass on the sensor chip surface and, therefore, a higher response (as 1 RU is defined as 1 .mu.g/mm.sup.2). Afterwards, the sensor chip is regenerated by dissolving the sample-mesothelin-complex. The gained responses are then evaluated relative to the response displayed by the reference standard (which is assumed to be 100% active). First, around 50 resonance units (RU) of human MSLN (0.75 .mu.g/ml) were coupled on a C1 chip (GE Healthcare) at pH 4.5 by using the amine coupling kit of GE Healthcare. The sample and system buffer was HBS-P+(0.01 M HEPES, 0.15 M NaCl, 0.005% surfactant P20 sterile-filtered, pH 7.4). The flow cell temperature was set to 25.degree. C., and the sample compartment temperature was set to 12.degree. C. The system was primed with running buffer. Then, a 5 nM solution of the MSLN PE cFP construct was injected for 60 seconds at a flow rate of 30 .mu.l/min, followed by a 60 second dissociation phase. Then, the sensor chip surface was regenerated by two 20 second long injections of the regeneration solution (0.31M KS ON, 1.22M MgCl.sub.2, 0.61M urea, 1.22M Gua-HCl, 6.7 mM EDTA) at a flow rate of 30 .mu.l/min, followed by an extra wash step with buffer and a 5 second stabilization period.
[0514] Stability data were obtained by incubating the 5 best humanized variants in the Fab-linker-PE24LR010 format at 37.degree. C. for 7 days. The results are shown in Table 23. As shown in Table 23, the variant VH01/VL01 was the only variant that showed no fragmentation after temperature stress. The VL01 promoted recovery as all VL01 combinations formed only aggregates after temperature stress but did not precipitate. Negative results of affinity measurements after temperature stress were due to these aggregates. Accordingly, the humanization variant VH1/VL1 (SEQ ID NO: 45)/(SEQ ID NO: 46) was chosen for further development. VH1 (SEQ ID NO: 45) provided favorable functionality and binding properties. The combination of VH1 (SEQ ID NO: 45)/VL1 (SEQ ID NO: 46) provided favorable developability.
TABLE-US-00023 TABLE 20 z a b c d e m1 m2 m3 ss1 001/15457 MSAb- MSAb- MSAb- LC_SS 000/ (SEQ ID 002/ 003/ 004/ 005/ 1/1546 2/1546 3/1546 1/ VL 15456 NO: 46) 15458 15459 15460 15461 2 3 4 15455 mouse humanized mVL IMGT_ IMGT_ IMGT_ IMGT_ Morpho Morpho Morpho Past hVK _ hVK_ hVK_1_ hVK_1_ tek tek tek Pastan VH (C99Q) 1_39 3_11 39 Hercept. 39 PAT PAT PAT PAT Mouse 0 000/15437 mVH Past 20.10 17.80 13.36 20.80 32.72 20.15 16.89 26.38 13.89 17.42 (C44Q) Hu- 001/15438 IMGT_hVH_1_46 32.04 20.27 15.57 22.44 28.95 30.31 33.50 21.78 21.23 27.22 man- 1 (SEQ ID ized NO: 45) 2 002/15439 VBase_VH1_1 88.85 204.74 32.01 48.39 38.63 103.46 181.39 81.92 47.73 271.77 3 003/15440 VBase_VH1_1 37.76 100.18 31.58 39.86 63.66 70.03 26.60 108.18 39.95 230.87 4 004/15441 VBase_VH1_1 39.46 46.18 20.21 52.31 112.42 68.93 73.53 71.82 89.99 270.06 5 005/15442 Herceptin 71.84 432.87 39.39 70.55 94.99 119.52 26.06 326.88 60.33 >500.0 6 006/15443 IMGT_hVH_5_51 196.59 287.08 41.11 144.51 466.56 163.55 >500.0 83.61 78.58 52.29 7 007/15444 IMGT_hVH_5_51 155.21 67.47 137.39 71.03 >500.0 69.42 210.52 89.05 182.10 181.13 8 008/15445 IMGT_hVH_5_51 431.47 44.39 42.37 17.40 143.88 25.90 20.03 16.34 37.09 87.91 9 009/15446 IMGT_hVH_1_8 118.32 62.23 71.58 31.00 230.84 68.65 103.29 97.08 80.73 44.37 10 010/15447 IGHV4-34-05 41.35 93.85 35.06 33.49 135.83 95.84 66.42 65.63 26.84 235.42 11 011/15448 IMGT_hVH_3_21 44.04 52.11 24.52 41.68 54.65 62.08 15.60 124.88 35.08 47.05 12 012/15449 VBase_VH1_1 30.19 37.51 7.86 47.87 36.40 61.27 30.13 35.53 20.68 86.50 13 013/15450 VBase_VH1_1 70.85 >500.0 58.84 129.80 111.42 132.05 16.51 204.88 33.20 >500.0 14 014/15451 VBase_VH1_1 #N/A #N/A #N/A #N/A #N/A #N/A #N/A #N/A #N/A #N/A M1 MSAb- Morphotek 25.65 19.12 13.24 18.82 33.79 21.71 26.82 13.24 17.05 7.37 1/15452 M2 MSAb- Morphotek 23.42 15.51 33.37 18.46 30.84 30.16 39.31 23.75 35.77 30.54 2/15453 M3 MSAb- Morphotek 34.70 62.58 17.71 15.82 64.02 53.63 117.75 19.86 42.60 45.23 3/15454 SS1 HC_SS1/ Pastan 47.15 38.16 26.40 28.85 60.38 37.42 29.65 29.79 14.77 41.13 15436
TABLE-US-00024 TABLE 21 z a b c d e m1 m2 m3 ss1 001/15457 LC_ 000/ (SEQ ID 002/ 003/ 004/ 005/ MSAb- MSAb- MSAb- SS1/ VL 15456 NO: 46) 15458 15459 15460 15461 1/15462 2/15463 3/15464 15455 mouse humanized mVL IMGT_ IMGT_ IMGT_ IMGT_ Past hVK _ hVK_ hVK_1_ hVK_1_ Morph Morph Morph Pastan VH (C99Q) 1_39 3_11 39 Hercept 39 PAT PAT PAT PAT Mouse 0 000/15437 mVHPast (C44Q) 24.93 18.25 14.91 16.97 28.24 19.65 13.30 20.08 16.62 21.59 Hu- 1 001/15438 IMGT_ 22.81 15.55 16.22 21.03 14.77 21.75 33.91 21.69 18.48 20.27 man- (SEQ hVH _1_46 ized ID NO: 45) 2 002/15439 VBase_VH1_1 20.80 30.71 9.67 22.61 18.10 23.88 26.61 30.97 26.83 23.16 3 003/15440 VBase_VH1_1 11.17 64.84 21.95 24.52 27.66 46.86 11.40 33.78 23.40 43.11 4 004/15441 VBase_VH1_1 13.30 18.79 11.38 20.34 20.71 23.06 18.22 20.33 19.35 35.09 5 005/15442 Herceptin 57.64 75.30 21.79 26.38 22.71 32.75 7.46 78.60 41.55 51.40 6 006/15443 IMGT_hVH_5_51 37.98 68.72 17.00 31.68 96.41 77.48 59.94 13.32 37.23 9.70 7 007/15444 IMGT_hVH_5_51 30.92 32.33 50.56 47.93 97.81 21.06 35.60 49.19 35.21 26.52 8 008/15445 IMGT_hVH_5_51 27.84 32.16 13.63 17.70 36.39 14.14 8.96 12.58 15.26 14.19 9 009/15446 IMGT_hVH_1_8 37.15 20.48 23.00 9.24 99.49 45.65 13.29 16.81 20.11 11.92 10 010/15447 IGHV4_34_05 17.19 36.94 18.55 3.79 33.67 29.20 15.63 19.91 14.16 58.47 11 011/15448 IMGT_hVH_3_21 17.85 19.31 12.09 26.23 19.38 25.78 6.71 42.47 25.88 26.04 12 012/15449 VBase_VH1_1 26.50 18.36 4.84 25.45 32.87 32.83 22.30 16.38 21.03 30.01 13 013/15450 VBase_VH1_1 17.11 48.46 26.21 32.81 17.97 55.13 18.76 40.91 16.60 70.42 14 014/15451 VBase_VH1_1 #N/A #N/A #N/A #N/A #N/A #N/A #N/A #N/A #N/A #N/A M1 MSAb- Morphotek 16.76 26.05 10.85 15.40 27.67 15.26 18.45 10.07 14.15 10.22 1/15452 M2 MSAb- Morphotek 18.94 13.24 25.61 18.06 18.24 32.68 49.52 33.94 33.72 31.20 2/15453 M3 MSAb- Morphotek 43.79 80.00 24.10 20.13 57.15 51.29 76.89 21.74 57.13 43.21 3/15454 SS1 HC_SS1/ Pastan 34.77 37.48 30.64 28.76 50.76 48.23 24.34 31.68 17.66 46.14 15436
TABLE-US-00025 TABLE 22 Antibody Antigen Curve Temp (.degree. C.) Fit ka kd KD KD [nM] cFP-0004-0002 Mesothelin Fc = 4-3 25 1:1 Binding 7.88E+05 2.39E-04 3.04E-10 0.30 cFP-0006-0002 Mesothelin Fc = 4-3 25 1:1 Binding 9.29E+05 2.62E-04 2.83E-10 0.28 cFP.15438-15457 Mesothelin Fc = 4-3 25 1:1 Binding 8.39E+05 2.20E-04 2.63E-10 0.26 cFP.15438-15459 Mesothelin Fc = 4-3 25 1:1 Binding 7.13E+05 1.97E-04 2.77E-10 0.28 cFP.15438-15460 Mesothelin Fc = 4-3 25 1:1 Binding 4.92E+05 1.63E-04 3.31E-10 0.33 cFP.15442-15457 Mesothelin Fc = 4-3 25 1:1 Binding 2.15E+05 0.001387 6.45E-09 6.45 cFP.15442-15459 Mesothelin Fc = 4-3 25 1:1 Binding 2.25E+05 0.002445 1.09E-08 10.89 cFP.15442-15460 Mesothelin Fc = 4-3 25 1:1 Binding 1.42E+05 0.002221 1.57E-08 15.70 cFP.15447-15457 Mesothelin Fc = 4-3 25 1:1 Binding 6.31E+05 9.76E-04 1.55E-09 1.55 cFP.15447-15459 Mesothelin Fe = 4-3 25 1:1 Binding 5.14E+05 0.001916 3.73E-09 3.73 cFP.15447-15460 Mesothelin Fc = 4-3 25 1:1 Binding 3.55E+05 0.002069 5.82E-09 5.82 cFP.15448-15457 Mesothelin Fc = 4-3 25 1:1 Binding 1.77E+05 0.001729 9.79E-09 9.79 cFP.15448-15459 Mesothelin Fc = 4-3 25 1:1 Binding 1.73E+05 0.001909 1.11E-08 11.05 cFP.15448-15460 Mesothelin Fc = 4-3 25 1:1 Binding 1.12E+05 0.00216 1.93E-08 19.32
TABLE-US-00026 TABLE 23 T.sub.agggreagetion ESI-MS Target Stress effect on VL VH [.degree. C.] Recovery SEC SDS-PAGE ESI-MS d d&r binding binding VL01 VH10 40 + HMW Fragmentation Fragmentation. Fragmentation. +/- HMW VL03 VH01 ~35 - LMW Fragmentation. Fragmentation. Fragmentation. + +/- VL01 VH01 37 + HMWs No No No + HMW fragmentation fragmentation fragmentation VL03 VH10 37 - HMW +/- Fragmentation. Fragmentation. +/- +/- VL04 VH01 34 - LMW +/- No No + + fragmentation fragmentation
Example 14
[0515] This example demonstrates the cloning, expression, refolding, and purification of chimeric deimmunized PE LOR10-456A as a Fab fusion with humanized anti-mesothelin SS1 (cFP-0205).
[0516] The chimeric PE anti-mesothelin molecule cFP-0205 was obtained after cloning, expression, refolding, and purification. cFP-0205 (also referred to as R205 or RG7787) comprises amino acid sequences SEQ ID NO: 39 and 40.
[0517] Cloning of cFP-0205:
[0518] For the expression of cFP-0205, the E. coli host/vector system which enables an antibiotic-free plasmid selection by complementation of an E. coli auxotrophy (PyrF) was employed (EP 0 972 838 and U.S. Pat. No. 6,291,245). Standard methods were used to manipulate DNA as described in Sambrook, J. et al., supra. The molecular biological reagents were used according to the manufacturer's instructions. Desired gene segments were made by commercial gene synthesis. The synthesized gene fragments were cloned into a specified expression vector. The DNA sequences of the subcloned gene fragments were confirmed by DNA sequencing.
[0519] The expression plasmids for the production of the light chain (LC) and the heavy chain (HC), respectively, were prepared as follows: The LC Plasmid 15496 is an expression plasmid for the expression of an antibody light chain in E. coli. It was generated by ligating the antibody VL domain fragment into the vector using the NdeI/BsiWI restriction sites.
[0520] The light chain E. coli expression plasmid comprises the following elements: [0521] the origin of replication from the vector pBR322 for replication in E. coli (corresponding to positions 2517-3160 according to Sutcliffe et al., Quant. Biol., 43: 77-90 (1979), [0522] the URA3 gene of Saccharomyces cerevisiae coding for orotidine 5'-phosphate decarboxylase (Rose et al., Gene, 29: 113-124 (1984), which allows plasmid selection by complementation of E. coli pyrF mutant strains (uracil auxotrophy), [0523] the antibody light chain expression cassette, comprising [0524] the T5 hybrid promoter (T5-PN25/03/04 hybrid promoter according to Bujard et al., Methods. Enzymol., 155: 416-433 (1987) and Stueber et al., Immunol. Methods IV, 121-152 (1990) including a synthetic ribosomal binding site according to Stueber et al. (supra), [0525] the antibody light chain variable domain comprising the CDRs of SS1, [0526] the human Ck domain, [0527] two bacteriophage-derived transcription terminators, the .lamda.-T0 terminator (Schwarz et al., Nature, 272: 410-414 (1978) and the fd-terminator (Beck et al., Gene, 1-3: 35-58 (1981), and [0528] the lad repressor gene from E. coli (Farabaugh, P. J., Nature, 274 (1978) 765-769).
[0529] The HC Plasmid 18023 is an expression plasmid for the expression of a fusion protein including an antibody heavy chain, a linker containing a furin cleavage site, and a mutant of domain III of Pseudomonas Exotoxin A, in E. coli. It was generated by ligation of the VH domain fragment into the vector using the NdeI/BsrGI restriction sites. The specific PE variant fragment was exchanged using the BsrGI/BsiWI restriction sites.
[0530] The cytolytic fusion protein heavy chain E. coli expression plasmid comprises the following elements: [0531] the origin of replication from the vector pBR322 for replication in E. coli (corresponding to positions 2517-3160 according to Sutcliffe et al., Quant. Biol., 43: 77-90 (1979), [0532] the URA3 gene of Saccharomyces cerevisiae coding for orotidine 5'-phosphate decarboxylase (Rose et al., Gene, 29: 113-124 (1984), which allows plasmid selection by complementation of E. coli pyrF mutant strains (uracil auxotrophy), [0533] the heavy chain-PE domain III fusion protein expression cassette, comprising: [0534] the T5 hybrid promoter (T5-PN25/03/04 hybrid promoter according to Bujard et al., Methods. Enzymol., 155: 416-433 (1987) and Stueber et al., Immunol. Methods IV, 121-152 (1990), including a synthetic ribosomal binding site according to Stueber et al. (supra), [0535] the antibody heavy chain variable domain comprising the CDRs of SS1, [0536] the human CH1 domain, [0537] the linker comprising a furin cleavage site, [0538] the mutant variant of Pseudomonas Exotoxin A domain III, [0539] two bacteriophage-derived transcription terminators, the .lamda.-T0 terminator (Schwarz et al., Nature, 272: 410-414 (1978) and the fd-terminator (Beck et al., Gene, 1-3: 35-58 (1981), and [0540] the lad repressor gene from E. coli (Farabaugh, Nature, 274: 765-769 (1978).
[0541] Expression of cFP-0205 in E. coli:
[0542] The E. coli K12 strains CSPZ-6 (thi-1, .DELTA.pyrF) and CSPZ-13 (thi-1, .DELTA.ompT, .DELTA.pyrF) were transformed by electroporation with the expression plasmid nos. 15496 and 18023, respectively. The transformed E. coli cells were first grown at 37.degree. C. on agar plates. For each transformation, a colony picked from this plate was transferred to a 3 mL roller culture and grown at 37.degree. C. to an optical density of 1-2 (measured at 578 nm). Then 1000 .mu.l of culture were mixed with 1000 .mu.l sterile 86%-glycerol and immediately frozen at -80.degree. C. for long term storage. The correct product expression of these clones was first verified by small scale shake flask experiments and analyzed with SDS-Page prior to the transfer to the 10 L fermenter.
Pre Cultivation:
[0543] For pre-fermentation, a chemically defined medium (CD-PCMv2.60) was used: NH.sub.4Cl 1.25 g/l, K.sub.2HPO.sub.4*3H.sub.2O 18.3 g/l, citrate 1.6 g/l, L-Methionine 0.20 g/l, Glucose*H.sub.2O 8.0 g/l, trace elements solution 0.5 ml/l, MgSO.sub.4*7H.sub.2O 0.86 g/l, and Thiamin*HCl 17.5 mg/l. The trace elements solution contained FeSO.sub.4*7H.sub.2O 10.0 g/l, ZnSO.sub.4*7H.sub.2O 2.25 g/l, MnSO.sub.4*H.sub.2O 2.13 g/l, H.sub.3BO.sub.3 0.50 g/l, (NH.sub.4)6Mo.sub.7O.sub.24*4H.sub.2O 0.3 g/l, CoCl.sub.2*6H.sub.2O 0.42 g/l, CuSO.sub.4*5H.sub.2O 1.0 g/l dissolved in 0.5M HCl.
[0544] For pre-fermentation, five 1000 mL shake-flasks with four baffles filled with 220 mL of medium were inoculated with 1.0 mL each out of a primary seed bank ampoule. The cultivation was performed on a rotary shaker for 11-12 hours at 37.degree. C. and 170 rpm until an optical density (578 nm) of 7 to 9 was obtained. The shake flasks were pooled, and the optical density was determined. The inoculum volume was calculated with Vinoc.=1000 mL*5/ODPC and is dependent on the optical density of the pre cultivation to inoculate the batch medium of each 100 L bioreactor run with an equal amount of cells.
100 L Scale Fermentation:
[0545] For fermentation in a 100 L fermenter (Sartorius, Melsungen, Germany), the following chemically defined batch medium was used: KH.sub.2PO.sub.4 1.59 g/l, (NH.sub.4).sub.2HPO.sub.4 7.45 g/l, K.sub.2HPO.sub.4*3H.sub.2O 13.32 g/l, citrate 2.07 g/l, L-methionine 1.22 g/l, NaHCO.sub.3 0.82 g/l, trace elements solution 7.3 ml/l, MgSO.sub.4*7 H2O 0.99 g/l, thiamine*HCl 20.9 mg/l, glucose*H.sub.2O 29.3 g/l, biotin 0.2 mg/l, 1.2 ml/l Synperonic 10% anti foam agent. The trace elements solution contained FeSO.sub.4*7H.sub.2O 10 g/l, ZnSO.sub.4*7H.sub.2O 2.25 g/l, MnSO.sub.4*H.sub.2O 2.13 g/l, CuSO.sub.4*5H.sub.2O 1.0 g/l, CoCl.sub.2*6H.sub.2O 0.42 g/l, (NH.sub.4)6Mo.sub.7O.sub.24*4H.sub.2O 0.3 g/l, H.sub.3BO.sub.3 0.50 g/l solubilized in a 0.5M HCl solution.
[0546] The feed 1 solution contained 700 g/l glucose*H.sub.2O, 7.4 g/1l MgSO.sub.4*7 H.sub.2O and 0.1 g/l FeSO.sub.4*7H.sub.2O. Feed 2 comprised KH.sub.2PO.sub.4 52.7 g/l, K.sub.2HPO.sub.4*3H.sub.2O 139.9 g/l and (NH.sub.4)2HPO.sub.4 66.0 g/1. All components were dissolved in deionized water. The alkaline solution for pH regulation was an aqueous 12.5% (w/v) NH.sub.3 solution supplemented with 11.25 g/l L-methionine.
[0547] Starting with 42 L of sterile batch medium plus the calculated inoculum of the pre cultivation the batch, fermentation was performed at 31.degree. C., pH 6.9.+-.0.2, 800 mbar back pressure and an initial aeration rate of 50 l/min. The relative value of dissolved oxygen (pO.sup.2) was kept at 50% throughout the fermentation by increasing the stirrer speed up to 1000 rpm. After the initially supplemented glucose was depleted, indicated by a steep increase in dissolved oxygen values, the temperature was shifted to 37.degree. C. when producing the light chain or kept constant when producing the heavy chain construct and 15 minutes later, the fermentation entered the fed-batch mode with the start of both feeds (600 and 140 g/h respectively). The rate of feed 2 was kept constant, while the rate of feed 1 was increased stepwise with a predefined feeding profile from 600 to finally 1400 g/h when producing the light chain or 1600 g/h when producing the heavy chain construct within 6-7 hours. When carbon dioxide off gas concentration leveled above 2%, the aeration rate was stepwise increased by 10 L/min from 50 to 100 l/min within 5 hours. The expression of recombinant target proteins as insoluble inclusion bodies located in the cytoplasm was induced by the addition of 24 g IPTG at an optical density of approx. 40 for the variable light Fab-chain and 120 for the variable heavy Fab-chain.
[0548] After 24 hours of cultivation, the whole broth was cooled down to 4-8.degree. C. and stored overnight in the fermenter vessel. The bacteria were harvested via centrifugation with a flow-through centrifuge (13,000 rpm, 13 l/h) or a separator and the obtained biomass was stored at -20.degree. C. until further processing (cell disruption) or immediately processed for inclusion body isolation.
Analysis of Product formation:
[0549] Samples were drawn from the fermenter, one prior to induction and the others at dedicated time points after induction of protein expression. The samples were analyzed by SDS-Polyacrylamide gel electrophoresis. For every sample, the same amount of cells (OD.sub.Target=10) was suspended in 5 mL PBS buffer and disrupted via sonication on ice. Then, 100 .mu.L of each suspension were centrifuged (15,000 rpm, 5 minutes) and each supernatant was withdrawn and transferred to a separate vial. This was to discriminate between soluble and insoluble expressed target protein. To each supernatant (=soluble protein fraction) 100 L and to each pellet (=insoluble protein fraction) 200 .mu.L of SDS sample buffer (Laemmli, Nature, 227: 680-685 (1970) were added. Samples were heated for 15 minutes at 95.degree. C. under intense mixing to solubilize and reduce all proteins in the samples. After cooling to room temperature, 5 .mu.L of each sample were transferred to a 4-20% TGX Criterion Stain Free polyacrylamide gel (Bio-Rad). Additionally, 5 .mu.l molecular weight standard (Precision Plus Protein Standard, Bio-Rad) and 3 amounts (0.3 .mu.l, 0.6 .mu.l and 0.9 .mu.l) quantification standard with known target protein concentration (0.1 .mu.g/.mu.l) were applied.
[0550] The electrophoresis was run for 60 Minutes at 200 V and, thereafter, the gel was transferred the GELDOC EZ Imager (Bio-Rad) and processed for 5 minutes with UV radiation. Gel images were analyzed using Image Lab analysis software (Bio-Rad). With the three standards, a linear regression curve was calculated with a coefficient of >0.99 and the concentrations of target protein in the original sample were calculated. The yield was up to 3-4 g/L of the Fab-light chain and 10-12 g/L for the Fab-heavy chain PE fusion.
Inclusion Body Preparation:
[0551] The inclusion body preparations (IBP) of the 100 L fermentations were started directly after the harvest of the bacteria with the re-suspension of the harvested bacteria cells in buffer 1 (12.1 g/L Tris, 0.246 g/L MgSO.sub.4*7H.sub.2O, 12 mL/L 25%-HCl). The buffer volume was calculated in dependence of the dry matter content of the biomass. Lysozyme (100 kU/mg, 0.12 mg/gDCW) and a small amount of benzonase (5 U/gDCW) were added. Then, the suspension was homogenized at 900 bar (APV Rannie 5, 1 pass) to disrupt the bacteria cells followed by the addition further benzonase (30 U/gDCW) and an incubation for 30 minutes at 30-37.degree. C. Then, the first wash buffer (60 g/L Brij, 87.6 g/L NaCl, 22.5 g/L EDTA, 6 mL/L 10N NaOH) was added and again the suspension was incubated for 30 minutes. The following separation step (CSC6, Westfalia) led to an inclusion body slurry which was re-suspended in the second wash buffer (12.1 g/L Tris, 7.4 g/L EDTA, 11 mL/L 25%-HCl) and incubated for 20 minutes. A further separation step harvested the inclusion bodies to a single use, sterile, plastic bag, which was immediately transferred to the DSP department.
Results:
[0552] The fermentations of the light chain yielded an optical density measured at 578 nm of 170-190 and a final product yield of 10-12 g/L. The fermentations of the heavy chain toxin fusion yielded an optical density measured at 578 nm of 210-230 and a final product yield of 3-4.5 g/L.
[0553] Refolding and Purification of cFP-0205:
[0554] Inclusion bodies of HC-PE24 and LC were solubilized separately in 8 M guanidinium-hydrochloride, 100 mM Tris/HCl, 1 mM EDTA, pH 8.0.+-.100 mM dithiothreitol (DTT) overnight at RT (1 g IB in 5 mL). Solubilizates were adjusted to pH 3 and centrifuged, and the pellet was discarded. After extensive dialysis against 8 M guanidinium-HCl, 10 mM EDTA, pH 3.0 to remove DTT, the total protein concentration was determined using the Biuret method. The purity of the HC and LC content was estimated via SDS-PAGE.
[0555] Solubilizates were diluted at a 1:1 molar ratio in renaturation buffer containing 0.5 M arginine, 2 mM EDTA, pH 10.+-.1 mM GSH/GSSG, respectively, at 2-10.degree. C. The target protein concentration was increased stepwise from 0.1 g/L up to 0.5 g/L, with an incubation time of 2 hours between each dose. After up to 5 pulses, the renaturation solution was kept at 2-10.degree. C. overnight.
[0556] The renaturate was diluted with H.sub.2O to <3 mS/cm and pumped onto an anion exchange column (AIEX) equilibrated in 20 mM Tris/HCl, pH 7.4. After washing the column with equilibration buffer, the protein was eluted with a gradient up to 20 mM Tris/HCl, 400 mM NaCl, pH 7.4. Peak fractions containing Fab-PE24 were pooled, concentrated and applied onto a preparative Size Exclusion Column (SEC) in 20 mM His, 140 mM NaCl, pH 5.5 or 6.0 to remove aggregates, fragments, and E. coli proteins. The final protein pool was adjusted to the required protein concentration and analyzed via SDS-PAGE or CE-SDS, analytical SEC and UV.sub.280. Identity was confirmed by mass spectrometry.
[0557] Protein Analysis of cFP-0205:
[0558] Sample analysis was carried out as described above.
Characterization of Final Deimmunized PE Variant as Fab Fusion with Humanized Anti-Mesothelin SS1 (cFP0205).
[0559] Cytotoxicity and in vitro tumor cell killing: The cytotoxic potency of cFP0205 was compared to that of SS1P in cell viability assays with different cancer cell lines and primary mesothelioma cells. The results are shown in Table 24. As shown in Table 24, for many cancer cell lines, SS1P and cFP0205 had comparable potencies (difference in IC.sub.50 values <3). However, some cell lines (e.g., Hu-1 and Hu-2) and also the primary mesothelioma cells (RH19, RH21) were significantly more sensitive to cFP0205 than to SS1P (6-10 fold lower IC.sub.50 values).
TABLE-US-00027 TABLE 24 SS1P cFP0205 Cell line (IC50 in ng/ml) (IC50 in ng/ml) Hu-1 2 0.25 Hu-2 0.3 0.04 Hu-3 2.9 2 HAY 1 1.4 AG5 0.3 1.1 L55 3 5 MKN-28 0.5 2 ASPC-1-luc 1.3 2.4 H596 4.6 15.7 RH19 3.7 1.2 RH21 2.3 0.35
Characterization of Final Deimmunized PE Variant as Fab Fusion with Humanized Anti-Mesothelin SS1 (cFP0205).
[0560] Serum half life: A pilot study was performed in cynomolgus monkeys (1 male and 1 female) with cFP0205 in order to assess the pharmacokinetics of single doses. The animals were intravenously dosed with 0.3 mg/kg of cFP0205. For pharmacokinetics, blood samples were taken predose, 0.083 (5 minutes, end of infusion), 1, 2, 3, 5, 8, 24, 48 and 168 hours postdose. Levels of free drug (dashed line) as well as total drug (black squares and solid line) were determined by different ELISA formats (FIG. 20). For determining total drug levels, a one-step acid dissociation was performed before the capturing step on mesothelin coated plates. The free and total drug levels were plotted over time. For comparison, historical data with SS1P were also inserted into FIG. 20 (black circles and solid line). These historical data were generated with a cytotoxic activity assay as a read-out for serum levels of SS1P. The cytotoxicity assay detects free drug, but also detects, at least in part, drug bound to soluble mesothelin which might contribute to the observed activity. The measured free drug levels for cFP0205 were similar to the first phase of the historical data with SS1P. The measured total drug levels for cFP0205 were comparable to the second phase of the kinetic observed for SS1P. In summary, not only in mice, but also in cynomolgus monkey, the serum level kinetics of SS1P and cFP0205 were very comparable.
Characterization of Final Deimmunized PE Variant as Fab Fusion with Humanized Anti Mesothelin SS1 (cFP0205)
[0561] Reduced Off-Target Toxicity:
[0562] SCID beige mice with small subcutaneous tumors of H596 lung cancer cells (average tumor volume 100 mm.sup.3) were dosed with 0.5, 1, 2, and 3 mg/kg cFP0205 3.times./w qod and body weight loss was monitored over one month. The results are shown in FIG. 21.
[0563] As shown in FIG. 21, the maximally tolerated dose of SS1P in mice given intravenously 3.times./w qod was 0.4 mg/kg. cFP0205 given with this same regimen was tolerated in SCID beige mice at doses up to 3 mg/kg. Some body weight loss was observed in all treatment groups. However, the effect of the 0.5 mg/kg dose of cFP0205 was indiscriminable from the vehicle control. Dose-dependently, an increasing loss of body weight occurred with 1, 2, and 3 mg/kg. In the 3 mg/kg group, the maximum weight loss was 15% and this was observed 4 days after the last application of cFP0205. In the 2 and 3 mg/kg treatment groups, no further decline in body weight was observed after the last application. All animals started to recover body weight from day 5 after the last treatment onwards. Initial body weight recovery was most pronounced in the 3 mg/kg group. In summary, despite similar pharmacokinetic properties of both molecules in mice, cFP0205 was tolerated at an almost 10 fold higher dose compared to SS1P.
[0564] A single intravenous dose (short infusion) toxicology study was done in 8 week old female Wistar Furth rats (150-175 g body weight) in order to evaluate and compare the off-target toxicity of SS1P and cFP0205. In particular, the risk of inducing vascular leak (edema) in the lung caused by the test item was investigated. Three animals per group received the active substance or vehicle by an intravenous short infusion at an infusion rate of 0.3 ml/min. Animals were necropsied 24 hours after dosing. Lung with mainstem bronchi and liver were macroscopically evaluated, fluid collection in the lung was assessed, and clinical chemistry assays were performed on blood samples. The results are shown in Table 25.
TABLE-US-00028 TABLE 25 SS1P cFP0205 Dose (i.v.), n = 3 2 mg/kg 10 mg/kg In-life rales (lung), rolling gait, hunched posture, none (clinical signs, bw) piloerection and slight bw loss Necropsy tan-stained thoracic fluid in 2 rats; brown none discoloration of liver in all 3 rats Fluid Smear mesothelial cells and eosinophils none Evaluation Clinical Liver: .uparw..uparw..uparw. in ALT (30x), AST (40x), Liver: Chemistry .gamma.GT, GLDH, SDH; marginal VLS: .dwnarw. Protein, .dwnarw.albumin (serum protein SDH .uparw. loss) Hemolysis ?: .uparw. bilirubin (but no hematology indication) .uparw. BUN: poor physical condition (.uparw. protein catabolism)
The serum half life of SS1P and cFP0205 is expected to be very similar in rats as shown for mice.
[0565] As shown in Table 25, two out of three rats that were dosed intravenously with 2 mg/kg of SS1P showed fluid accumulation in the lungs. All animals in this group showed clear clinical signs of toxicity and, for all animals, clinical chemistry values were indicative of severe liver damage. In contrast to this, all three animals treated with 10 mg/kg of cFP0205 showed no signs of toxicity. Only in one animal was the value for one of the liver enzymes slightly elevated. These findings demonstrate that the cFP0205 molecule causes much less hepatotoxicity and vascular leak syndrome than the SS1P classical immunotoxin format.
[0566] A pilot toxicology study was performed in cynomolgus monkeys (1 male and 1 female) with cFP0205 in order to determine the tolerability of repeated daily doses of the test item following intravenous administration and to check for potential late onset toxicity (up to 72 hrs after last repeat-dose). Postdosing observations of the animals were performed immediately, 0.25, 0.5, 1, 2, and 4 hours after end of dosing. During the dosing phase, the general behavior and appearance of the animals was observed twice daily, while body weight, food consumption and feces were analysed daily.
[0567] Dosing cFP0205 at 1 mg/kg daily for 5 consecutive days was well tolerated by both animals. No hemorrhaging, acute inflammation, or ulceration at the injection site were observed. That the animals showed no clinical signs of toxicity or non-tolerability firmly established 1 mg/kg/d 5.times. as the non-severely toxic dose, while the classical immunotoxin format represented by SS1P had previously been shown to be non-tolerated at a dose of 0.3 mg/kg/d 5.times. with animals showing persistent clinical signs of toxicity like hunched posture with tremors, hypoactivity, and poor appetite. In summary, cFP0205 is much better tolerated than SS1P not only in mice, but also in cynomolgus monkeys, a fully cross-reactive species for the targeting moieties of these immunoconjugates.
Characterization of Final Deimmunized PE Variant as Fab Fusion with Humanized SS1 (cFP0205)
[0568] cFP0205 and SS1P were labeled with Cy5 on free amino groups. Outstaged SCID beige mice with subcutaneous H596 tumors were intravenously injected (4 mice/group) at a dose of 2 mg/kg with the 2 fluorescent-labeled molecules. After 6 hours, the animals were sacricifed. The lung, liver, and spleen were formalin-fixed, embedded and sectioned. Fluorescence images of representative sections of these organs were taken at different magnification. The results are shown in FIGS. 22A-22L.
[0569] As shown in FIGS. 22A-22L, SS1P treatment led to pronounced fluorescence staining of lung and spleen tissue and massively stained what appeared to be the reticuloendothelial system in the liver. In contrast to this, there was no staining of lung tissue observed upon cFP0205 application. Also, staining of the spleen was much less pronounced and the staining pattern was different from that of SS1P treated animals. Also in the liver, the staining with cFP0205 was drastically reduced compared to SS1P. Only isolated single cells that appear to be Kupffer cells were fluorescently stained. This suggests that the lower off-target toxicity of a Fab-PE24 compared to a dsFv-PE38 is due to differences in normal tissue distribution.
Example 15
[0570] This example demonstrates the production of anti-glypican 3-PE variant chimeric molecules.
[0571] An anti-glypican 3-PE variant chimeric molecule comprising a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 37 (LO10R-456A), the variable heavy chain domain VH of SEQ ID NO: 61, and the variable light chain domain of SEQ ID NO: 62 (GPC3-PE24-LR-LO10R-456A-long-linker) was prepared by a method analogous to the method of preparing the anti-mesothelin PE variant chimeric molecule cFP-077 described in Example 12 (from cloning to purification).
[0572] An anti-glypican 3-PE variant chimeric molecule comprising a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 37 (LO10R-456A), the variable heavy chain domain VH of SEQ ID NO: 77, and the variable light chain domain of SEQ ID NO: 78 was also prepared by a method analogous to the method of preparing the anti-mesothelin PE variant chimeric molecule cFP-077 described in Example 12 (from cloning to purification).
Example 16
[0573] This example demonstrates the cytotoxic properties of the anti-glypican 3-PE variant chimeric molecule GPC3-PE24-LR-LO10R-456A-long-linker as compared to GPC3-PE24-LR-LO10R.
[0574] The cytotoxic potency of GPC3-PE24-LR-LO10R was compared to that of GPC3-PE24-LR-LO10R-456A-long-linker on the glypican 3-positive liver cancer cell line HepG2. Briefly, HepG2 cells were seeded at a density of 6000 cells/well on 96 well plates. After overnight culture, different concentrations of the cFPs were added to the medium, and the cells were incubated for 72 hours. At the end of incubation period, cell viability was determined using a CELLTITERGLOW assay. The results are shown in Table 26.
[0575] As shown in Table 26, comparable dose-response curves for the inhibition of cell viability were observed on HepG2 cells for both molecules. The IC.sub.50 value of the more completely de-immunized PE variant with the 456A mutation and the longer linker was even slightly lower than that of the LR-LO10R variant with the short linker. Non-targeted ("free PE24") or mistargeted PE24 ("CD33-Fab-PE"; HepG2 cells do not express CD33) did not show any significant reduction of cell viability at concentrations <30 .mu.M. In summary, the more completely de-immunized GPC3-targeted LR-LO10R-456A-long-linker variant had the same or even slightly better cytotoxic potency as compared to the GPC3-targeted LR-LO10R variant with the shorter linker. Due to the additional 456A mutation within the PE, all B-cell epitopes were removed.
TABLE-US-00029 TABLE 26 IC50 in CELLTITERGLOW assay with HepG2 Fusion protein (expressing glypican-3) GPC3-PE24-LR-LO10R-short-linker 0.68 nM GPC3-PE24-LR-LO10R-456A- 0.54 nM long-linker Non-targeted PE24 >>30 .mu.M CD33-PE24-LR-LO10R-short-linker no effect up to 30 .mu.M
Example 17
[0576] This example demonstrates the cytotoxicity of deimmunized PE fusion proteins having a A458R mutation.
[0577] The cytotoxicity of SS1P, SS1-LR-GGS, SS1-LR-GGS-LO10, SS1-LR-GGS-LO10-A458R, and SS1-LR-GGS-LO10-A458-456A was measured using the WST method. The results are shown in Table 27.
TABLE-US-00030 TABLE 27 A431/H9 HAY M30 Hu-Meso AGS SS1P 0.05(N = 14) 1.49(N = 6) 0.28(N = 5) 0.48(N = 5) 0.36(N = 5) SS1-LR-GGS 0.16(N = 2) ND ND ND 0.3 SS1-LR-GGS-L010 0.17(N = 5) 0.68 3.99 0.11 11(N = 2) SS1-LR-GGS-L010-A458R 0.13(N = 18) 0.39(N = 11) 0.57(N = 3) 0.03(N = 4) 1.05(N = 9) SS1-LR-GGS-L010-A458R- 0.07(N = 4) 0.1(N = 2) 0.58(N = 3) ND 0.5 456A
[0578] As shown in Table 27, the A458R mutation increased the activity of SS1-LR-GGS-L010 on several cell lines. The addition of the 456A mutation further increased activity on several cell lines. A431/H9 is an epidermoid carcinoma expressing mesothelin, and Hay is a mesothelioma line. M30 is also a mesothelioma cell line, and AGS is a stomach cancer cell line. Hu-Meso are cells from a mesothelioma patient that were placed in culture for a few months.
Example 18
[0579] This example demonstrates the treatment of HCC70 tumors using RG7787.
[0580] Female nude mice were innoculated into the intramammary fat pad with HCC70 cells at time 0. HCC70 is a breast cancer cell line. Intravenous treatment with RG7787 (2.5 mg/kg IV) or vehicle was begun on day 6 and continued every other day for a total of 5 doses (arrows). The size of the tumor was measured. The results are shown in FIG. 24.
[0581] As shown in FIG. 24, the tumor size of mice treated with RG7787 was decreased as compared to that of control mice (vehicle). There is a statistically significant difference between the 2 groups beginning at day 10, with p<0.00001 by day 14.
Example 19
[0582] This example demonstrates the treatment of KLM1 tumors in mice using a combination of paclitaxel and RG7787.
[0583] Four week, four day old mice were injected with 4.times.10.sup.6 KLM1 cells. KLM1 is a pancreatic cancer cell line that expresses mesothelin. Thirteen days later, mice were untreated or treated with paclitaxel (50 mg/kg), RG7787 (2.5 mg/kg), or a combination of RG7787 (2.5 mg/kg) and paclitaxel (50 mg/kg). Tumor size was measured for up to 89 days after treatment. The results are shown in FIG. 23.
[0584] As shown in FIG. 23, the tumor size of mice treated with the combination of paclitaxel and RG7787 was decreased as compared to that of control mice (untreated) or that of the mice treated with paclitaxel alone or RG7787 alone.
Example 20
[0585] This example demonstrates the treatment of HCC70 tumors in mice using a combination of paclitaxel and RG7787.
[0586] Female athymic nude mice were inoculated with HCC70 cells at time 0. Animals were treated with vehicle, RG7787 with IP vehicle injection, paclitaxel with IV vehicle injection, or paclitaxel and RG7787 combination. Paclitaxel (50 mg/kg) was administered by IP injection on days marked with long arrows. RG7787 (2.5 mg/kg) was administered IV on days indicated by short arrows. Mean tumor volumes for n=5 vehicle and n=6 mice treated with RG7787, paclitaxel or the combination are indicated by the markers. The results are shown in FIG. 25.
[0587] As shown in FIG. 25, the tumor size of mice treated with the combination of paclitaxel and RG7787 was decreased as compared to that of control mice (vehicle) or that of the mice treated with paclitaxel alone or RG7787 alone.
Example 21
[0588] This example demonstrates the treatment of MKN-28 tumors in mice using a combination of paclitaxel and R205.
[0589] Athymic nude mice were inoculated with MKN-28 cells at time 0. MKN-28 is a gastric cancer cell line which expresses mesothelin. Animals were untreated (UT) or treated with Roche 205 (R205 or 205) alone, paclitaxel (taxol) alone, or a combination of R205 and paclitaxel. Tumor size was measured. The results are shown in FIG. 26. As shown in FIG. 26, the tumor size of mice treated with the combination of paclitaxel and R205 was decreased as compared to that of control mice (untreated) or that of the mice treated with paclitaxel alone or R205 alone.
Example 22
[0590] This example demonstrates the in vivo antitumor activity of LMB-T18. Severe combined immunodeficient (SCID) mice were implanted with CA46 cells. Seven days later, when tumors reached over 100 mm.sup.3 in size, the mice were treated with PBS (control) or LMB-T18 (5 mg/kg.times.4 or 7.5 mg/kg.times.3) intravenously. Mice receiving 5.0 mg/kg were treated four times, on days 7, 9, 11 and 16, and the higher dose group was treated with 7.5 mg/kg three times on days 7, 9 and 11. Marked tumor regressions were observed in all mice (FIG. 27E) with only minor weight loss (average of 6%). 5/7 mice treated with 5.0 mg/kg maintained complete tumor regression on day 33, and 3/7 maintained complete tumor regression in the 7.5 mg/kg group. To assess the nonspecific toxicity of LMB-T18, six tumor bearing mice were treated intravenously with two doses of 10 mg/kg QOD. One mouse showed severe weight loss and was euthanized.
Example 23
[0591] This example demonstrates that LMB-T18 has greatly diminished T cell activation.
[0592] To determine if LMB-T18 had a decrease in T cell stimulation or if new T cell epitopes were created by the mutations, PBMCs were stimulated from the highest responder donors (n=13) and I-CL and mesothelioma patients (n=7) with MP or LMB-T18. Cells were re-stimulated with the 39 novel peptides representing the differences between MP and LMB-T18. A decrease of 90% in donor T cell activation (p<0.0001 in Student T test) was observed. Even in patients with activated T cells, there was an 83% decrease. (p<0.0001 in Student T test). Furthermore, no new epitopes were created by the mutations.
Example 24
[0593] This example demonstrates that LMB-T18 has reduced binding to anti-sera from patients.
[0594] The antigenicity of LMB-T18 was evaluated by comparing the reactivity of MP, HA22-LR and LMB-T18 with serum from patients with neutralizing antibodies to MP. Binding was measured using ICC-ELISA with serum from 13 MP treated patients and is shown in FIG. 27F. It was found that like HA22-LR, LMB-T18 had a significantly reduced binding to serum compared to MP (p<0.001 one-way ANOVA). It was also found that LMB-T18 had significantly lower binding compared to HA22-LR (p<0.001 one-way ANOVA), indicating that the mutations in LMB-T18 reduced the binding to anti-sera.
Example 25
[0595] This example demonstrates the production of anti-FAP-PE variant chimeric molecules.
[0596] Three anti-FAP-PE variant chimeric molecules comprising: [0597] (a) a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 37 (LO10R-456A)), the variable heavy chain domain VH of SEQ ID NO: 93, and the variable light chain domain of SEQ ID NO: 94 (iFAP-PE24-LR-LO10 OR-456A-long-linker) [0598] (b) a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 37 ((LO10R-456A)), the variable heavy chain domain VH of SEQ ID NO: 93, and the variable light chain domain of SEQ ID NO: 294 (LCL4-PE24-LR-LO10 OR-456A-long-linker) (including the VL variant of SEQ ID NO: 290), and [0599] (c) a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 37 (T-20), the variable heavy chain domain VH of SEQ ID NO: 93, and the variable light chain domain of SEQ ID NO: 290 (LCL4-T20); were prepared by a method analogous to the method of preparing the anti-mesothelin PE variant chimeric molecule cFP-077 described in Example 12 (from cloning to purification), except as described below.
[0600] The anti-FAP-PE variant chimeric molecules (b) and (c) included a modified variable light chain domain (VL) in which the methionine at position 4 of the original VL sequence of SEQ ID NO: 94 was substituted with leucine to provide a modified VL sequence of SEQ ID NO: 290 ("LCL4"). The modified variable light chain domain SEQ ID NO: 290 was found to reduce the production of impurities related to N-terminal modifications. These N-terminal modifications provided a mixture of variants with different light chain lengths due to the initiation of expression of the unmodified sequence at either the N-terminal methionine or the methionine at position 4 of of SEQ ID NO: 94.
Pre-Cultivation:
[0601] For pre-fermentation, the same chemically defined medium (CD-PCMv2.20) was used as described in Example 12.
[0602] The cultivation was performed on a rotary shaker for 8 hours at 37.degree. C. and 170 rpm until an optical density (578 nm) of 13.3 or 14.4 was obtained. 150 ml of the pre-cultivation was used to inoculate the batch medium of the 10 L bioreactor.
Fermentation:
[0603] For fermentation in a 101 Biostat C, DCU3 fermenter (Sartorius, Melsungen, Germany), the same chemically-defined batch medium as described in Example 12 was used.
[0604] One feed solution that was used contained 700 g/l glucose*H.sub.2O, 7.4 g/l MgSO.sub.4*7 H.sub.2O and 0.1 g/l FeSO.sub.4*7H.sub.2O. All components were dissolved in deionized water. The alkaline solution for pH regulation was an aqueous 12.5% (w/v) NH.sub.3 solution supplemented with 11.25 g/l L-methionine and 10 g/L Leucine and 10 g/L Threonine.
[0605] Starting with 6.24 L sterile batch medium plus 150 mL inoculum from the pre-cultivation, the batch fermentation was performed at 32.degree. C., pH 6.9.+-.0.2, 800 mbar back pressure and an initial aeration rate of 10 l/min. The relative value of dissolved oxygen (pO.sub.2) was kept at 50% throughout the fermentation by increasing the stirrer speed up to 1500 rpm. After the initially supplemented glucose was depleted, indicated by a steep increase in dissolved oxygen values, the temperature was shifted to 32.degree. C. or 37.degree. C., and 15 minutes later, the fermentation entered the fed-batch mode with the start of both the feeds (60 and 14 g/h respectively). The rate of the feed was increased stepwise with a predefined feeding profile from 90 to finally 210 or 240 g/h within 5.5 or 6.5 hours. When carbon dioxide off gas concentration leveled above 2%, the aeration rate was constantly increased from 10 to 20 l/min within 5 hours. The expression of recombinant protein was induced by the addition of 3.6 g IPTG at an optical density of approx. 40 or 120. The target protein was expressed as inclusion bodies within the cytoplasm.
[0606] After 24 hours of cultivation, an optical density of 179 or 171 was achieved, and the whole broth was cooled down to 4-8.degree. C. The bacteria were harvested via centrifugation with lab centrifuge (4500 rpm, cooling at 4.degree. C., for 1 h) and the obtained biomass was stored at -20.degree. C. until further processing (cell disruption).
Analysis of Product Formation:
[0607] The specific differences in comparison to the method of Example 12 are described below:
[0608] The same amount of cells (OD.sub.Target=10) from every sample were suspended in 5 mL PBS buffer and disrupted via sonication on ice. Then 100 .mu.L of each suspension were centrifuged (8,000 rpm, 5 minutes) and each supernatant was withdrawn and transferred to a separate vial.
[0609] After adding Laemmli buffer, the samples were heated for 45 minutes 40.degree. C. under intense mixing to solubilize and reduce all proteins in the samples.
Inclusion Body Preparation:
[0610] The inclusion body preparations (IBP) of the 10 L fermentations were processed in a manner analogous to the method described in Example 12, with the exception that after disruption of the bacteria cells, further benzoase (30 U/gDCW) was added and incubated for 60 minutes at 25.degree. C.
[0611] Solubilization, renaturation and purification were performed in a manner similar to that described in Example 12. The following sequences were used: [0612] Sequence of iFAP-PE24-LR-LO10R-456A-long-linker and LCL4-PE24-LR-LO10R-456A-long-linker chimeric full length heavy chain comprising the variable domain VH of SEQ ID NO: 93, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37 (SEQ ID NO: 291); [0613] Sequence LCL4-T20 chimeric heavy chain construct comprising the full length heavy chain comprising the variable domain VH of SEQ ID NO: 93, a linker comprising the amino acid sequence of SEQ ID NO: 36 and a PE comprising SEQ ID NO: 289 (T18/T20) (SEQ ID NO: 292); [0614] Sequence of full length iFAP light chain (SEQ ID NO: 293) comprising the variable light chain domain of SEQ ID NO: 94 and constant region; [0615] Sequence of full length LCL4 light chain (SEQ ID NO: 294) comprising the variable light chain domain of SEQ ID NO: 290 and constant region; and [0616] Sequence of the variable light chain domain of SEQ ID NO: 290 (with mutation M4L at position 4 (compared to SEQ ID NO: 94).
Example 26
[0617] This example demonstrates the cytotoxicity of the anti-FAP-PE variant chimeric molecules iFAP-PE24-LR-LO10R-456A-long-linker, LCL4-PE24-LR-LO10R-456A-long-linker, and LCL4-T20.
[0618] The cytotoxicity of iFAP-PE24-LR-LO10R-456A-long-linker was compared to that of LCL4-PE24-LR-LO10R-456A-long-linker and LCL4-T20 ("cFPs") with respect to the FAP-positive fibroblast cell line MRC-5, the FAP-positive desmoplastic melanoma cell line LOX-IMVI, and the FAP-positive sarcoma cell line OsA-CL. Briefly, cells were seeded at a density of 10,000 cells/well on 96-well plates. After overnight culture, different concentrations of the cFPs were added to the medium, and the cells were incubated for 72 hours. At the end of incubation period, cell viability was determined using a CELLTITER GLO assay. The results are shown in Table 28 and Table 29.
[0619] As shown in Table 28, comparable IC50 values for the inhibition of cell viability were observed on all 3 tested cell lines for iFAP-PE24-LR-LO10R-456A-long-linker (3 different batches) and for LCL4-PE24-LR-LO10R-456A-long-linker. As shown in Table 29, LCL4-T20 also showed potent inhibition of cell viability. In summary, iFAP-PE24-LR-LO10R-456A-long-linker, LCL4-PE24-LR-LO10 OR-456A-long-linker, and LCL4-T20 showed potent cytotoxic effects on FAP-positive tumor cell lines and fibroblasts.
TABLE-US-00031 TABLE 28 IC50 in CELLTITER GLO assay with cell lines expressing FAP(ng/ml) Fusion MRC-5 MRC-5 LOX- protein (experiment 1) (experiment 2) IMVI OsA-CL iFAP-PE24- 14.3 8.5 6.6 98.3 LR-LO10R- 456A-long- linker (Batch 1) iFAP-PE24- 14.4 9.3 9.1 46.3 LR-LO10R- 456A-long- linker (Batch 2) iFAP-PE24- 10.0 5.5 5.7 55.3 LR-LO10R- 456A-long- linker (Batch 3) LCL4-PE24- 13.9 9.8 9.7 103.1 LR-LO10R- 456A-long- linker
TABLE-US-00032 TABLE 29 IC50 in CELLTITER GLO assay with cell lines expressing FAP(nM) Fusion protein MRC-5 LOX-IMVI OsA-CL LCL4-PE24-LR-LO10R- 0.15 nM 0.33 nM 0.23 nM 456A-long-linker LCL4-T20 0.98 nM 8.99 nM 0.50 nM
Example 27
[0620] This example demonstrates the production of anti-CAIX-PE variant chimeric molecules.
[0621] Two anti-CAIX-PE variant chimeric molecules comprising: [0622] (a) a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 37 (LO10R-456A), the variable heavy chain domain VH of SEQ ID NO: 125, and the variable light chain domain of SEQ ID NO: 126; or [0623] (b) a linker comprising the amino acid sequence of SEQ ID NO: 36, a PE comprising SEQ ID NO: 289 (T-20), the variable heavy chain domain VH of SEQ ID NO: 125, and the variable light chain domain of SEQ ID NO: 126; were prepared by a method analogous to that described for the anti-mesothelin PE variant chimeric molecule cFP-077 of Example 12 (from cloning to purification), except as described below.
[0624] Samples were analyzed by OD 280 nm using a UV spectrophotometer to determine the protein concentration in solution. The materials for the SDS-PAGE and Coomassie Staining Device are set forth in Table 30 below.
TABLE-US-00033 TABLE 30 Invitrogen XCell Sure Lock Mini-Cell Gel: 4-12% Bis-Tris Gel, Invitrogen NP0321 Buffer: MES SDS Running Buffer (10x), Invitrogen NP0002 Sample buffer: Tris-Glycine SDS Sample Buffer (2x), Invitrogen LC2676 Reducing buffer: NUPAGE Sample Reducing Agent (10x), Invitrogen NP0004 Molecular Weight Precision Plus KALEIDOSCOPE Standard Marker: 161-0375
[0625] After adding Laemmli buffer, the samples were heated for 45 minutes at 40.degree. C. under intense mixing to solubilize and reduce all proteins in the samples.
[0626] The sample was adjusted to a protein concentration of 1 mg/ml with buffer. Sample reduction used a reduction buffer comprising 4 ml of sample buffer (2.times.) and 1 ml of reducing buffer (10.times.). The samples were reduced by diluting the sample 1:1 with reduction buffer and incubating the sample for 10 minutes at 70.degree. C.
[0627] The gel electrophoresis was carried out at 200 V for 40 minutes. The gels were stained with SIMPLY BLUE Safe Stain (Invitrogen, Cat. No. LC6065).
Inclusion Body Preparation:
[0628] The inclusion body preparations (IBP) of the 10 L fermentations were processed in a manner analogous to that described in Example 12, with the exception that after disruption of the bacteria cells, further benzoase (30U/gDCW) was added and incubated for 60 minutes at 25.degree. C.
[0629] Solubilization, renaturation and purification were performed in a manner similar to that described in Example 12.
[0630] The following sequences were used: [0631] Sequence of CAIX-PE24-LR-LO10R-456A-long-linker chimeric heavy chain construct (SEQ ID NO: 295): comprising the variable heavy chain domain VH of SEQ ID NO: 125, a linker comprising the amino acid sequence of SEQ ID NO: 36, and a PE comprising SEQ ID NO: 37; [0632] Sequence of CAIX-T20 chimeric heavy chain construct (SEQ ID NO: 296) comprising the variable heavy chain domain VH of SEQ ID NO: 125, a linker comprising the amino acid sequence of SEQ ID NO: 36 and a PE comprising SEQ ID NO: 289 (T18/T20); and [0633] Sequence of full length light chain (SEQ ID NO: 297) comprising the variable light chain domain of SEQ ID NO: 126) and constant region.
Example 28
[0634] This example demonstrates the cytotoxicity (in vitro tumor cell killing) of the anti-CAIX-PE variant chimeric molecules CAIX-PE24-LR-LO10R-456A-long-linker and CAIX-T20.
[0635] The cytotoxicity of CAIX-PE24-LR-LO10 OR-456A-long-linker and CAIX-T20 ("cFPs") was compared in cell viability assays using CAIX-expressing RCC-MF cells. Briefly, cells were seeded at a density of 7,500 cells/well on 96 well plates. After overnight culture, different concentrations of the cFPs were added to the medium, and the cells were incubated for 72 hours. At the end of incubation period, cell viability was determined using a CELLTITER GLO assay. The results are shown in Table 31.
[0636] In summary, CAIX-PE24-LR-LO10 OR-456A-long-linker and CAIX-T20 showed potent cytotoxic effects on CAIX-positive tumor cell lines.
TABLE-US-00034 TABLE 31 IC50 in CELLTITER GLO assay with cell lines expressing CAIX (nM) Fusion protein RCC-MF cells CAIX-PE24-LR-LO10R- 0.02 nM 456A-long-linker CAIX-T20 0.048 nM
[0637] All references, including publications, patent applications, and patents, cited herein are hereby incorporated by reference to the same extent as if each reference were individually and specifically indicated to be incorporated by reference and were set forth in its entirety herein.
[0638] The use of the terms "a" and "an" and "the" and "at least one" and similar referents in the context of describing the invention (especially in the context of the following claims) are to be construed to cover both the singular and the plural, unless otherwise indicated herein or clearly contradicted by context. The use of the term "at least one" followed by a list of one or more items (for example, "at least one of A and B") is to be construed to mean one item selected from the listed items (A or B) or any combination of two or more of the listed items (A and B), unless otherwise indicated herein or clearly contradicted by context. The terms "comprising," "having," "including," and "containing" are to be construed as open-ended terms (i.e., meaning "including, but not limited to,") unless otherwise noted. Recitation of ranges of values herein are merely intended to serve as a shorthand method of referring individually to each separate value falling within the range, unless otherwise indicated herein, and each separate value is incorporated into the specification as if it were individually recited herein. All methods described herein can be performed in any suitable order unless otherwise indicated herein or otherwise clearly contradicted by context. The use of any and all examples, or exemplary language (e.g., "such as") provided herein, is intended merely to better illuminate the invention and does not pose a limitation on the scope of the invention unless otherwise claimed. No language in the specification should be construed as indicating any non-claimed element as essential to the practice of the invention.
[0639] Preferred embodiments of this invention are described herein, including the best mode known to the inventors for carrying out the invention. Variations of those preferred embodiments may become apparent to those of ordinary skill in the art upon reading the foregoing description. The inventors expect skilled artisans to employ such variations as appropriate, and the inventors intend for the invention to be practiced otherwise than as specifically described herein. Accordingly, this invention includes all modifications and equivalents of the subject matter recited in the claims appended hereto as permitted by applicable law. Moreover, any combination of the above-described elements in all possible variations thereof is encompassed by the invention unless otherwise indicated herein or otherwise clearly contradicted by context.
Sequence CWU 1
1
2971613PRTPseudomonas aeruginosa 1Ala Glu Glu Ala Phe Asp Leu Trp
Asn Glu Cys Ala Lys Ala Cys Val1 5 10 15Leu Asp Leu Lys Asp Gly Val
Arg Ser Ser Arg Met Ser Val Asp Pro 20 25 30Ala Ile Ala Asp Thr Asn
Gly Gln Gly Val Leu His Tyr Ser Met Val 35 40 45Leu Glu Gly Gly Asn
Asp Ala Leu Lys Leu Ala Ile Asp Asn Ala Leu 50 55 60Ser Ile Thr Ser
Asp Gly Leu Thr Ile Arg Leu Glu Gly Gly Val Glu65 70 75 80Pro Asn
Lys Pro Val Arg Tyr Ser Tyr Thr Arg Gln Ala Arg Gly Ser 85 90 95Trp
Ser Leu Asn Trp Leu Val Pro Ile Gly His Glu Lys Pro Ser Asn 100 105
110Ile Lys Val Phe Ile His Glu Leu Asn Ala Gly Asn Gln Leu Ser His
115 120 125Met Ser Pro Ile Tyr Thr Ile Glu Met Gly Asp Glu Leu Leu
Ala Lys 130 135 140Leu Ala Arg Asp Ala Thr Phe Phe Val Arg Ala His
Glu Ser Asn Glu145 150 155 160Met Gln Pro Thr Leu Ala Ile Ser His
Ala Gly Val Ser Val Val Met 165 170 175Ala Gln Thr Gln Pro Arg Arg
Glu Lys Arg Trp Ser Glu Trp Ala Ser 180 185 190Gly Lys Val Leu Cys
Leu Leu Asp Pro Leu Asp Gly Val Tyr Asn Tyr 195 200 205Leu Ala Gln
Gln Arg Cys Asn Leu Asp Asp Thr Trp Glu Gly Lys Ile 210 215 220Tyr
Arg Val Leu Ala Gly Asn Pro Ala Lys His Asp Leu Asp Ile Lys225 230
235 240Pro Thr Val Ile Ser His Arg Leu His Phe Pro Glu Gly Gly Ser
Leu 245 250 255Ala Ala Leu Thr Ala His Gln Ala Cys His Leu Pro Leu
Glu Thr Phe 260 265 270Thr Arg His Arg Gln Pro Arg Gly Trp Glu Gln
Leu Glu Gln Cys Gly 275 280 285Tyr Pro Val Gln Arg Leu Val Ala Leu
Tyr Leu Ala Ala Arg Leu Ser 290 295 300Trp Asn Gln Val Asp Gln Val
Ile Arg Asn Ala Leu Ala Ser Pro Gly305 310 315 320Ser Gly Gly Asp
Leu Gly Glu Ala Ile Arg Glu Gln Pro Glu Gln Ala 325 330 335Arg Leu
Ala Leu Thr Leu Ala Ala Ala Glu Ser Glu Arg Phe Val Arg 340 345
350Gln Gly Thr Gly Asn Asp Glu Ala Gly Ala Ala Asn Ala Asp Val Val
355 360 365Ser Leu Thr Cys Pro Val Ala Ala Gly Glu Cys Ala Gly Pro
Ala Asp 370 375 380Ser Gly Asp Ala Leu Leu Glu Arg Asn Tyr Pro Thr
Gly Ala Glu Phe385 390 395 400Leu Gly Asp Gly Gly Asp Val Ser Phe
Ser Thr Arg Gly Thr Gln Asn 405 410 415Trp Thr Val Glu Arg Leu Leu
Gln Ala His Arg Gln Leu Glu Glu Arg 420 425 430Gly Tyr Val Phe Val
Gly Tyr His Gly Thr Phe Leu Glu Ala Ala Gln 435 440 445Ser Ile Val
Phe Gly Gly Val Arg Ala Arg Ser Gln Asp Leu Asp Ala 450 455 460Ile
Trp Arg Gly Phe Tyr Ile Ala Gly Asp Pro Ala Leu Ala Tyr Gly465 470
475 480Tyr Ala Gln Asp Gln Glu Pro Asp Ala Arg Gly Arg Ile Arg Asn
Gly 485 490 495Ala Leu Leu Arg Val Tyr Val Pro Arg Ser Ser Leu Pro
Gly Phe Tyr 500 505 510Arg Thr Ser Leu Thr Leu Ala Ala Pro Glu Ala
Ala Gly Glu Val Glu 515 520 525Arg Leu Ile Gly His Pro Leu Pro Leu
Arg Leu Asp Ala Ile Thr Gly 530 535 540Pro Glu Glu Glu Gly Gly Arg
Leu Glu Thr Ile Leu Gly Trp Pro Leu545 550 555 560Ala Glu Arg Thr
Val Val Ile Pro Ser Ala Ile Pro Thr Asp Pro Arg 565 570 575Asn Val
Gly Gly Asp Leu Asp Pro Ser Ser Ile Pro Asp Lys Glu Gln 580 585
590Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser Gln Pro Gly Lys Pro Pro
595 600 605Arg Glu Asp Leu Lys 610212PRTArtificial
SequenceSyntheticMISC_FEATURE(1)..(1)Xaa at position 1 is Leu or
AlaMISC_FEATURE(4)..(4)Xaa at position 4 is Leu or
AlaMISC_FEATURE(5)..(5)Xaa at position 5 is Tyr or
AlaMISC_FEATURE(6)..(6)Xaa at position 6 is Leu or
AlaMISC_FEATURE(9)..(9)Xaa at position 9 is Arg or Ala 2Xaa Val Ala
Xaa Xaa Xaa Ala Ala Xaa Leu Ser Trp1 5 10312PRTPseudomonas
aeruginosa 3Leu Val Ala Leu Tyr Leu Ala Ala Arg Leu Ser Trp1 5
1044PRTArtificial SequenceSyntheticMISC_FEATURE(2)..(2)Xaa at
position 2 is any amino acidMISC_FEATURE(3)..(3)Xaa at position 3
is Arg or Lys 4Arg Xaa Xaa Arg154PRTArtificial SequenceSynthetic
5Lys Asp Glu Leu165PRTPseudomonas aeruginosa 6Arg Glu Asp Leu Lys1
574PRTPseudomonas aeruginosa 7Arg Glu Asp Leu1811PRTPseudomonas
aeruginosa 8Arg His Arg Gln Pro Arg Gly Trp Glu Gln Leu1 5
1094PRTArtificial SequenceSyntheticmisc_feature(2)..(3)Xaa can be
any naturally occurring amino acid 9Arg Xaa Xaa
Arg1104PRTArtificial SequenceSynthetic 10Arg Lys Lys
Arg1114PRTArtificial SequenceSynthetic 11Arg Arg Arg
Arg1124PRTArtificial SequenceSynthetic 12Arg Lys Ala
Arg1136PRTArtificial SequenceSynthetic 13Ser Arg Val Ala Arg Ser1
5149PRTArtificial SequenceSynthetic 14Thr Ser Ser Arg Lys Arg Arg
Phe Trp1 5159PRTArtificial SequenceSynthetic 15Ala Ser Arg Arg Lys
Ala Arg Ser Trp1 5168PRTArtificial SequenceSynthetic 16Arg Arg Val
Lys Lys Arg Phe Trp1 5178PRTArtificial SequenceSynthetic 17Arg Asn
Val Val Arg Arg Asp Trp1 5189PRTArtificial SequenceSynthetic 18Thr
Arg Ala Val Arg Arg Arg Ser Trp1 5194PRTArtificial
SequenceSynthetic 19Arg Gln Pro Arg1208PRTArtificial
SequenceSynthetic 20Arg His Arg Gln Pro Arg Gly Trp1
5219PRTArtificial SequenceSynthetic 21Arg His Arg Gln Pro Arg Gly
Trp Glu1 5229PRTArtificial SequenceSynthetic 22His Arg Gln Pro Arg
Gly Trp Glu Gln1 5237PRTArtificial SequenceSynthetic 23Arg Gln Pro
Arg Gly Trp Glu1 52411PRTArtificial SequenceSynthetic 24Arg His Arg
Ser Lys Arg Gly Trp Glu Gln Leu1 5 10254PRTArtificial
SequenceSynthetic 25Arg Ser Lys Arg1268PRTArtificial
SequenceSynthetic 26Arg His Arg Ser Lys Arg Gly Trp1
5278PRTArtificial SequenceSynthetic 27His Arg Ser Lys Arg Gly Trp
Glu1 5289PRTArtificial SequenceSynthetic 28Arg Ser Lys Arg Gly Trp
Glu Gln Leu1 52910PRTArtificial SequenceSynthetic 29His Arg Ser Lys
Arg Gly Trp Glu Gln Leu1 5 10306PRTArtificial SequenceSynthetic
30Arg His Arg Ser Lys Arg1 531106PRTArtificial SequenceSynthetic
31Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr
Met 20 25 30His Trp Tyr Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu
Ile Tyr 35 40 45Asp Thr Ser Lys Leu Ala Ser Gly Val Pro Ser Arg Phe
Ser Gly Ser 50 55 60Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser
Leu Gln Pro Glu65 70 75 80Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Trp
Ser Lys His Pro Leu Thr 85 90 95Phe Gly Gln Gly Thr Lys Leu Glu Ile
Lys 100 10532107PRTArtificial SequenceSynthetic 32Arg Thr Val Ala
Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys
Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro
Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser
Gly Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55
60Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65
70 75 80Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser
Ser 85 90 95Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 100
10533214PRTArtificial SequenceSynthetic 33Met Asp Ile Gln Met Thr
Gln Ser Pro Ser Ser Leu Ser Ala Ser Val1 5 10 15Gly Asp Arg Val Thr
Ile Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr 20 25 30Met His Trp Tyr
Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Asp Thr
Ser Lys Leu Ala Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Trp Ser Lys His Pro Leu
85 90 95Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 21034119PRTArtificial SequenceSynthetic
34Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Gly
Tyr 20 25 30Thr Met Asn Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Leu Ile Thr Pro Tyr Asn Gly Ala Ser Ser Tyr Asn
Gln Lys Phe 50 55 60Arg Gly Lys Ala Thr Met Thr Val Asp Thr Ser Thr
Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Gly Gly Tyr Asp Gly Arg Gly
Phe Asp Tyr Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val Ser Ser
11535103PRTArtificial SequenceSynthetic 35Ala Ser Thr Lys Gly Pro
Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1 5 10 15Ser Thr Ser Gly Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro
Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His
Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser
Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr65 70 75
80Tyr Ile Cys Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys
85 90 95Lys Val Glu Pro Lys Ser Cys 1003626PRTArtificial
SequenceSynthetic 36Asp Lys Thr His Lys Ala Ser Gly Gly Arg His Arg
Gln Pro Arg Gly1 5 10 15Trp Glu Gln Leu Gly Gly Gly Gly Gly Ser 20
2537219PRTArtificial SequenceSynthetic 37Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly Asp Val Ser Phe Ser1 5 10 15Thr Arg Gly Thr Gln
Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His 20 25 30Ala Gln Leu Glu
Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr 35 40 45Phe Leu Glu
Ala Ala Gln Ser Ile Val Phe Gly Gly Val Ala Ala Arg 50 55 60Ser Gln
Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala Gly Asp65 70 75
80Pro Ala Leu Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Ala
85 90 95Gly Arg Ile Arg Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Ala
Ser 100 105 110Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu Thr Leu Ala
Ala Pro Glu 115 120 125Ala Ala Gly Glu Val Glu Arg Leu Ile Gly His
Pro Leu Pro Leu Ala 130 135 140Leu Asp Ala Ile Thr Gly Pro Glu Glu
Glu Gly Gly Arg Leu Glu Thr145 150 155 160Ile Leu Gly Trp Pro Leu
Ala Glu Arg Thr Val Val Ile Pro Ser Ala 165 170 175Ile Pro Thr Asp
Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser 180 185 190Ile Pro
Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser 195 200
205Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 210
21538223PRTArtificial SequenceSynthetic 38Met Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly1 5 10 15Ala Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Gly 20 25 30Tyr Thr Met Asn
Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp 35 40 45Met Gly Leu
Ile Thr Pro Tyr Asn Gly Ala Ser Ser Tyr Asn Gln Lys 50 55 60Phe Arg
Gly Lys Ala Thr Met Thr Val Asp Thr Ser Thr Ser Thr Val65 70 75
80Tyr Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
85 90 95Cys Ala Arg Gly Gly Tyr Asp Gly Arg Gly Phe Asp Tyr Trp Gly
Gln 100 105 110Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly
Pro Ser Val 115 120 125Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser
Gly Gly Thr Ala Ala 130 135 140Leu Gly Cys Leu Val Lys Asp Tyr Phe
Pro Glu Pro Val Thr Val Ser145 150 155 160Trp Asn Ser Gly Ala Leu
Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175Leu Gln Ser Ser
Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185 190Ser Ser
Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys 195 200
205Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys 210
215 22039214PRTArtificial SequenceSynthetic 39Met Asp Ile Gln Met
Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val1 5 10 15Gly Asp Arg Val
Thr Ile Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr 20 25 30Met His Trp
Tyr Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Asp
Thr Ser Lys Leu Ala Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Trp Ser Lys His Pro Leu
85 90 95Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 21040468PRTArtificial SequenceSynthetic
40Met Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly1
5 10 15Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr
Gly 20 25 30Tyr Thr Met Asn Trp Val Arg Gln Ala Pro Gly Gln Gly Leu
Glu Trp 35 40 45Met Gly Leu Ile Thr Pro Tyr Asn Gly Ala Ser Ser Tyr
Asn Gln Lys 50 55 60Phe Arg Gly Lys Ala Thr Met Thr Val Asp Thr Ser
Thr Ser Thr Val65 70 75 80Tyr Met Glu Leu Ser Ser Leu Arg Ser Glu
Asp Thr Ala
Val Tyr Tyr 85 90 95Cys Ala Arg Gly Gly Tyr Asp Gly Arg Gly Phe Asp
Tyr Trp Gly Gln 100 105 110Gly Thr Leu Val Thr Val Ser Ser Ala Ser
Thr Lys Gly Pro Ser Val 115 120 125Phe Pro Leu Ala Pro Ser Ser Lys
Ser Thr Ser Gly Gly Thr Ala Ala 130 135 140Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val Ser145 150 155 160Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185
190Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys
195 200 205Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser
Cys Asp 210 215 220Lys Thr His Lys Ala Ser Gly Gly Arg His Arg Gln
Pro Arg Gly Trp225 230 235 240Glu Gln Leu Gly Gly Gly Gly Gly Ser
Pro Thr Gly Ala Glu Phe Leu 245 250 255Gly Asp Gly Gly Asp Val Ser
Phe Ser Thr Arg Gly Thr Gln Asn Trp 260 265 270Thr Val Glu Arg Leu
Leu Gln Ala His Ala Gln Leu Glu Glu Arg Gly 275 280 285Tyr Val Phe
Val Gly Tyr His Gly Thr Phe Leu Glu Ala Ala Gln Ser 290 295 300Ile
Val Phe Gly Gly Val Ala Ala Arg Ser Gln Asp Leu Ala Ala Ile305 310
315 320Trp Ala Gly Phe Tyr Ile Ala Gly Asp Pro Ala Leu Ala Tyr Gly
Tyr 325 330 335Ala Gln Asp Gln Glu Pro Asp Ala Ala Gly Arg Ile Arg
Asn Gly Ala 340 345 350Leu Leu Arg Val Tyr Val Pro Ala Ser Ser Leu
Pro Gly Phe Tyr Arg 355 360 365Thr Ser Leu Thr Leu Ala Ala Pro Glu
Ala Ala Gly Glu Val Glu Arg 370 375 380Leu Ile Gly His Pro Leu Pro
Leu Ala Leu Asp Ala Ile Thr Gly Pro385 390 395 400Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro Leu Ala 405 410 415Glu Arg
Thr Val Val Ile Pro Ser Ala Ile Pro Thr Asp Pro Arg Asn 420 425
430Val Gly Gly Asp Leu Asp Pro Ser Ser Ile Pro Asp Lys Glu Gln Ala
435 440 445Ile Ser Ala Leu Pro Asp Tyr Ala Ser Gln Pro Gly Lys Pro
Pro Arg 450 455 460Glu Asp Leu Lys46541214PRTArtificial
SequenceSynthetic 41Met Asp Ile Glu Leu Thr Gln Ser Pro Ala Ile Met
Ser Ala Ser Pro1 5 10 15Gly Glu Lys Val Thr Met Thr Cys Ser Ala Ser
Ser Ser Val Ser Tyr 20 25 30Met His Trp Tyr Gln Gln Lys Ser Gly Thr
Ser Pro Lys Arg Trp Ile 35 40 45Tyr Asp Thr Ser Lys Leu Ala Ser Gly
Val Pro Gly Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Asn Ser Tyr Ser
Leu Thr Ile Ser Ser Val Glu Ala65 70 75 80Glu Asp Asp Ala Thr Tyr
Tyr Cys Gln Gln Trp Ser Lys His Pro Leu 85 90 95Thr Phe Gly Gln Gly
Thr Lys Leu Glu Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val
Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr
Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135
140Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln145 150 155 160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys
21042461PRTArtificial SequenceSynthetic 42Met Gln Val Gln Leu Gln
Gln Ser Gly Pro Glu Leu Glu Lys Pro Gly1 5 10 15Ala Ser Val Lys Ile
Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Gly 20 25 30Tyr Thr Met Asn
Trp Val Lys Gln Ser His Gly Lys Gly Leu Glu Trp 35 40 45Ile Gly Leu
Ile Thr Pro Tyr Asn Gly Ala Ser Ser Tyr Asn Gln Lys 50 55 60Phe Arg
Gly Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala65 70 75
80Tyr Met Asp Leu Leu Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Phe
85 90 95Cys Ala Arg Gly Gly Tyr Asp Gly Arg Gly Phe Asp Tyr Trp Gly
Gln 100 105 110Gly Thr Thr Val Thr Val Ser Ser Ala Ser Thr Lys Gly
Pro Ser Val 115 120 125Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser
Gly Gly Thr Ala Ala 130 135 140Leu Gly Cys Leu Val Lys Asp Tyr Phe
Pro Glu Pro Val Thr Val Ser145 150 155 160Trp Asn Ser Gly Ala Leu
Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175Leu Gln Ser Ser
Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185 190Ser Ser
Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys 195 200
205Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Lys
210 215 220Ala Ser Gly Gly Arg His Arg Gln Pro Arg Gly Trp Glu Gln
Leu Gly225 230 235 240Gly Ser Pro Thr Gly Ala Glu Phe Leu Gly Asp
Gly Gly Asp Val Ser 245 250 255Phe Ser Thr Arg Gly Thr Gln Asn Trp
Thr Val Glu Arg Leu Leu Gln 260 265 270Ala His Ala Gln Leu Glu Glu
Arg Gly Tyr Val Phe Val Gly Tyr His 275 280 285Gly Thr Phe Leu Glu
Ala Ala Gln Ser Ile Val Phe Gly Gly Val Arg 290 295 300Ala Ala Ser
Gln Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala305 310 315
320Gly Asp Pro Ala Leu Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp
325 330 335Ala Ala Gly Arg Ile Arg Asn Gly Ala Leu Leu Arg Val Tyr
Val Pro 340 345 350Ala Ser Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu
Thr Leu Ala Ala 355 360 365Pro Glu Ala Ala Gly Glu Val Glu Arg Leu
Ile Gly His Pro Leu Pro 370 375 380Leu Ala Leu Asp Ala Ile Thr Gly
Pro Glu Glu Glu Gly Gly Arg Leu385 390 395 400Glu Thr Ile Leu Gly
Trp Pro Leu Ala Glu Arg Thr Val Val Ile Pro 405 410 415Ser Ala Ile
Pro Thr Asp Pro Arg Asn Val Gly Gly Asp Leu Asp Pro 420 425 430Ser
Ser Ile Pro Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr 435 440
445Ala Ser Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 450 455
46043107PRTArtificial SequenceSynthetic 43Met Asp Ile Glu Leu Thr
Gln Ser Pro Ala Ile Met Ser Ala Ser Pro1 5 10 15Gly Glu Lys Val Thr
Met Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr 20 25 30Met His Trp Tyr
Gln Gln Lys Ser Gly Thr Ser Pro Lys Arg Trp Ile 35 40 45Tyr Asp Thr
Ser Lys Leu Ala Ser Gly Val Pro Gly Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Asn Ser Tyr Ser Leu Thr Ile Ser Ser Val Glu Ala65 70 75
80Glu Asp Asp Ala Thr Tyr Tyr Cys Gln Gln Trp Ser Lys His Pro Leu
85 90 95Thr Phe Gly Cys Gly Thr Lys Leu Glu Ile Lys 100
10544358PRTArtificial SequenceSynthetic 44Met Gln Val Gln Leu Gln
Gln Ser Gly Pro Glu Leu Glu Lys Pro Gly1 5 10 15Ala Ser Val Lys Ile
Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Gly 20 25 30Tyr Thr Met Asn
Trp Val Lys Gln Ser His Gly Lys Cys Leu Glu Trp 35 40 45Ile Gly Leu
Ile Thr Pro Tyr Asn Gly Ala Ser Ser Tyr Asn Gln Lys 50 55 60Phe Arg
Gly Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala65 70 75
80Tyr Met Asp Leu Leu Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Phe
85 90 95Cys Ala Arg Gly Gly Tyr Asp Gly Arg Gly Phe Asp Tyr Trp Gly
Gln 100 105 110Gly Thr Thr Val Thr Val Ser Ser Lys Ala Ser Gly Gly
Arg His Arg 115 120 125Gln Pro Arg Gly Trp Glu Gln Leu Gly Gly Ser
Pro Thr Gly Ala Glu 130 135 140Phe Leu Gly Asp Gly Gly Asp Val Ser
Phe Ser Thr Arg Gly Thr Gln145 150 155 160Asn Trp Thr Val Glu Arg
Leu Leu Gln Ala His Ala Gln Leu Glu Glu 165 170 175Arg Gly Tyr Val
Phe Val Gly Tyr His Gly Thr Phe Leu Glu Ala Ala 180 185 190Gln Ser
Ile Val Phe Gly Gly Val Arg Ala Ala Ser Gln Asp Leu Ala 195 200
205Ala Ile Trp Ala Gly Phe Tyr Ile Ala Gly Asp Pro Ala Leu Ala Tyr
210 215 220Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Ala Gly Arg Ile
Arg Asn225 230 235 240Gly Ala Leu Leu Arg Val Tyr Val Pro Ala Ser
Ser Leu Pro Gly Phe 245 250 255Tyr Arg Thr Ser Leu Thr Leu Ala Ala
Pro Glu Ala Ala Gly Glu Val 260 265 270Glu Arg Leu Ile Gly His Pro
Leu Pro Leu Ala Leu Asp Ala Ile Thr 275 280 285Gly Pro Glu Glu Glu
Gly Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro 290 295 300Leu Ala Glu
Arg Thr Val Val Ile Pro Ser Ala Ile Pro Thr Asp Pro305 310 315
320Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser Ile Pro Asp Lys Glu
325 330 335Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser Gln Pro Gly
Lys Pro 340 345 350Pro Arg Glu Asp Leu Lys 35545119PRTArtificial
SequenceSynthetic 45Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr
Ser Phe Thr Gly Tyr 20 25 30Thr Met Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly Leu Glu Trp Met 35 40 45Gly Leu Ile Thr Pro Tyr Asn Gly Ala
Ser Ser Tyr Asn Gln Lys Phe 50 55 60Arg Gly Lys Ala Thr Met Thr Val
Asp Thr Ser Thr Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Gly Gly Tyr
Asp Gly Arg Gly Phe Asp Tyr Trp Gly Gln Gly 100 105 110Thr Leu Val
Thr Val Ser Ser 11546106PRTArtificial SequenceSynthetic 46Asp Ile
Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp
Arg Val Thr Ile Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr Met 20 25
30His Trp Tyr Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu Ile Tyr
35 40 45Asp Thr Ser Lys Leu Ala Ser Gly Val Pro Ser Arg Phe Ser Gly
Ser 50 55 60Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln
Pro Glu65 70 75 80Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Trp Ser Lys
His Pro Leu Thr 85 90 95Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100
1054730PRTArtificial SequenceSynthetic 47Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser
Cys Lys Ala Ser Gly Tyr Ser Phe Thr 20 25 304823PRTArtificial
SequenceSynthetic 48Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser
Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
20495PRTArtificial SequenceSynthetic 49Gly Tyr Thr Met Asn1
55010PRTArtificial SequenceSynthetic 50Ser Ala Ser Ser Ser Val Ser
Tyr Met His1 5 105114PRTArtificial SequenceSynthetic 51Trp Val Arg
Gln Ala Pro Gly Gln Gly Leu Glu Trp Met Gly1 5 105215PRTArtificial
SequenceSynthetic 52Trp Tyr Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu
Leu Ile Tyr1 5 10 155317PRTArtificial SequenceSynthetic 53Leu Ile
Thr Pro Tyr Asn Gly Ala Ser Ser Tyr Asn Gln Lys Phe Arg1 5 10
15Gly547PRTArtificial SequenceSynthetic 54Asp Thr Ser Lys Leu Ala
Ser1 55532PRTArtificial SequenceSynthetic 55Lys Ala Thr Met Thr Val
Asp Thr Ser Thr Ser Thr Val Tyr Met Glu1 5 10 15Leu Ser Ser Leu Arg
Ser Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg 20 25
305632PRTArtificial SequenceSynthetic 56Gly Val Pro Ser Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr1 5 10 15Leu Thr Ile Ser Ser Leu
Gln Pro Glu Asp Phe Ala Thr Tyr Tyr Cys 20 25 305710PRTArtificial
SequenceSynthetic 57Gly Gly Tyr Asp Gly Arg Gly Phe Asp Tyr1 5
10589PRTArtificial SequenceSynthetic 58Gln Gln Trp Ser Lys His Pro
Leu Thr1 55911PRTArtificial SequenceSynthetic 59Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Ser1 5 106010PRTArtificial SequenceSynthetic
60Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys1 5 1061115PRTArtificial
SequenceSynthetic 61Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr
Thr Phe Thr Asp Tyr 20 25 30Glu Met His Trp Val Arg Gln Ala Pro Gly
Gln Gly Leu Glu Trp Met 35 40 45Gly Ala Leu Asp Pro Lys Thr Gly Asp
Thr Ala Tyr Ser Gln Lys Phe 50 55 60Lys Gly Arg Val Thr Leu Thr Ala
Asp Lys Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu
Thr Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Thr Arg Phe Tyr Ser
Tyr Thr Tyr Trp Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ser
11562112PRTArtificial SequenceSynthetic 62Asp Val Val Met Thr Gln
Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile
Ser Cys Arg Ser Ser Gln Ser Leu Val His Ser 20 25 30Asn Arg Asn Thr
Tyr Leu His Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln Leu
Leu Ile Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55 60Asp Arg
Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75
80Ser Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Ser Gln Asn
85 90 95Thr His Val Pro Pro Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile
Lys 100 105 1106330PRTArtificial SequenceSynthetic 63Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr 20 25
306423PRTArtificial SequenceSynthetic 64Asp Val Val Met Thr Gln Ser
Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser
Cys 20655PRTArtificial SequenceSynthetic 65Asp Tyr Glu Met His1
56616PRTArtificial SequenceSynthetic 66Arg Ser Ser Gln Ser Leu Val
His Ser Asn Arg Asn Thr Tyr Leu His1 5 10 156714PRTArtificial
SequenceSynthetic 67Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp
Met Gly1 5 106815PRTArtificial SequenceSynthetic 68Trp Tyr Leu Gln
Lys Pro Gly Gln Ser Pro Gln Leu Leu Ile Tyr1 5 10
156917PRTArtificial SequenceSynthetic 69Ala Leu Asp Pro Lys Thr Gly
Asp Thr Ala Tyr Ser Gln Lys Phe Lys1 5 10 15Gly707PRTArtificial
SequenceSynthetic 70Lys Val Ser Asn
Arg Phe Ser1 57132PRTArtificial SequenceSynthetic 71Arg Val Thr Leu
Thr Ala Asp Lys Ser Thr Ser Thr Ala Tyr Met Glu1 5 10 15Leu Ser Ser
Leu Thr Ser Glu Asp Thr Ala Val Tyr Tyr Cys Thr Arg 20 25
307232PRTArtificial SequenceSynthetic 72Gly Val Pro Asp Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr1 5 10 15Leu Lys Ile Ser Arg Val
Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys 20 25 30736PRTArtificial
SequenceSynthetic 73Phe Tyr Ser Tyr Thr Tyr1 5749PRTArtificial
SequenceSynthetic 74Ser Gln Asn Thr His Val Pro Pro Thr1
57511PRTArtificial SequenceSynthetic 75Trp Gly Gln Gly Thr Leu Val
Thr Val Ser Ser1 5 107610PRTArtificial SequenceSynthetic 76Phe Gly
Gln Gly Thr Lys Leu Glu Ile Lys1 5 1077115PRTArtificial
SequenceSynthetic 77Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Ala1 5 10 15Ser Val Thr Val Ser Cys Lys Ala Ser Gly Tyr
Thr Phe Thr Asp Tyr 20 25 30Glu Met His Trp Ile Arg Gln Pro Pro Gly
Glu Gly Leu Glu Trp Ile 35 40 45Gly Ala Ile Asp Pro Lys Thr Gly Asp
Thr Ala Tyr Ser Glu Ser Phe 50 55 60Gln Asp Arg Val Thr Leu Thr Ala
Asp Lys Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu
Thr Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Thr Arg Phe Tyr Ser
Tyr Thr Tyr Trp Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ser
11578112PRTArtificial SequenceSynthetic 78Asp Ile Val Met Thr Gln
Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile
Ser Cys Gln Ala Ser Glu Ser Leu Val His Ser 20 25 30Asn Arg Asn Thr
Tyr Leu His Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln Leu
Leu Ile Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55 60Asp Arg
Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75
80Ser Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Ser Gln Asn
85 90 95Thr His Val Pro Pro Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Glu 100 105 1107930PRTArtificial SequenceSynthetic 79Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val
Thr Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr 20 25
308023PRTArtificial SequenceSynthetic 80Asp Ile Val Met Thr Gln Ser
Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser
Cys 20815PRTArtificial SequenceSynthetic 81Asp Tyr Glu Met His1
58216PRTArtificial SequenceSynthetic 82Gln Ala Ser Glu Ser Leu Val
His Ser Asn Arg Asn Thr Tyr Leu His1 5 10 158314PRTArtificial
SequenceSynthetic 83Trp Ile Arg Gln Pro Pro Gly Glu Gly Leu Glu Trp
Ile Gly1 5 108415PRTArtificial SequenceSynthetic 84Trp Tyr Leu Gln
Lys Pro Gly Gln Ser Pro Gln Leu Leu Ile Tyr1 5 10
158517PRTArtificial SequenceSynthetic 85Ala Ile Asp Pro Lys Thr Gly
Asp Thr Ala Tyr Ser Glu Ser Phe Gln1 5 10 15Asp867PRTArtificial
SequenceSynthetic 86Lys Val Ser Asn Arg Phe Ser1 58732PRTArtificial
SequenceSynthetic 87Arg Val Thr Leu Thr Ala Asp Lys Ser Thr Ser Thr
Ala Tyr Met Glu1 5 10 15Leu Ser Ser Leu Thr Ser Glu Asp Thr Ala Val
Tyr Tyr Cys Thr Arg 20 25 308832PRTArtificial SequenceSynthetic
88Gly Val Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr1
5 10 15Leu Lys Ile Ser Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr
Cys 20 25 30896PRTArtificial SequenceSynthetic 89Phe Tyr Ser Tyr
Thr Tyr1 5909PRTArtificial SequenceSynthetic 90Ser Gln Asn Thr His
Val Pro Pro Thr1 59111PRTArtificial SequenceSynthetic 91Trp Gly Gln
Gly Thr Leu Val Thr Val Ser Ser1 5 109210PRTArtificial
SequenceSynthetic 92Phe Gly Gln Gly Thr Lys Val Glu Ile Glu1 5
1093124PRTArtificial SequenceSynthetic 93Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser
Cys Lys Thr Ser Arg Tyr Thr Phe Thr Glu Tyr 20 25 30Thr Ile His Trp
Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Ile 35 40 45Gly Gly Ile
Asn Pro Asn Asn Gly Ile Pro Asn Tyr Asn Gln Lys Phe 50 55 60Lys Gly
Arg Val Thr Ile Thr Val Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Arg Ile Ala Tyr Gly Tyr Asp Glu Gly His Ala Met
Asp 100 105 110Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115
12094113PRTArtificial SequenceSynthetic 94Asp Ile Val Met Thr Gln
Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile
Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Arg Asn Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys
Leu Leu Ile Phe Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp
Arg Phe Ser Gly Ser Gly Phe Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Tyr Phe Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys9530PRTArtificial SequenceSynthetic 95Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val
Lys Val Ser Cys Lys Thr Ser Arg Tyr Thr Phe Thr 20 25
309623PRTArtificial SequenceSynthetic 96Asp Ile Val Met Thr Gln Ser
Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn
Cys 20975PRTArtificial SequenceSynthetic 97Glu Tyr Thr Ile His1
59817PRTArtificial SequenceSynthetic 98Lys Ser Ser Gln Ser Leu Leu
Tyr Ser Arg Asn Gln Lys Asn Tyr Leu1 5 10 15Ala9914PRTArtificial
SequenceSynthetic 99Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp
Ile Gly1 5 1010015PRTArtificial SequenceSynthetic 100Trp Tyr Gln
Gln Lys Pro Gly Gln Pro Pro Lys Leu Leu Ile Phe1 5 10
1510117PRTArtificial SequenceSynthetic 101Gly Ile Asn Pro Asn Asn
Gly Ile Pro Asn Tyr Asn Gln Lys Phe Lys1 5 10
15Gly1027PRTArtificial SequenceSynthetic 102Trp Ala Ser Thr Arg Glu
Ser1 510332PRTArtificial SequenceSynthetic 103Arg Val Thr Ile Thr
Val Asp Thr Ser Ala Ser Thr Ala Tyr Met Glu1 5 10 15Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg 20 25
3010432PRTArtificial SequenceSynthetic 104Gly Val Pro Asp Arg Phe
Ser Gly Ser Gly Phe Gly Thr Asp Phe Thr1 5 10 15Leu Thr Ile Ser Ser
Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys 20 25
3010515PRTArtificial SequenceSynthetic 105Arg Arg Ile Ala Tyr Gly
Tyr Asp Glu Gly His Ala Met Asp Tyr1 5 10 151069PRTArtificial
SequenceSynthetic 106Gln Gln Tyr Phe Ser Tyr Pro Leu Thr1
510711PRTArtificial SequenceSynthetic 107Trp Gly Gln Gly Thr Leu
Val Thr Val Ser Ser1 5 1010810PRTArtificial SequenceSynthetic
108Phe Gly Gln Gly Thr Lys Val Glu Ile Lys1 5 10109115PRTArtificial
SequenceSynthetic 109Glu Val Gln Leu Gln Gln Ser Gly Pro Glu Leu
Val Lys Pro Gly Thr1 5 10 15Ser Val Arg Ile Ser Cys Lys Thr Ser Gly
Tyr Thr Phe Thr Glu Tyr 20 25 30Thr Ile His Trp Val Lys Gln Ser His
Gly Lys Ser Leu Glu Trp Ile 35 40 45Gly Asn Ile Asn Pro Asn Asn Gly
Gly Thr Thr Tyr Asn Gln Lys Phe 50 55 60Glu Asp Lys Ala Thr Leu Thr
Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Glu Leu Arg Ser
Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala Ala Gly Trp
Asn Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr 100 105 110Val Ser
Ser 115110107PRTArtificial SequenceSynthetic 110Asp Ile Val Met Thr
Gln Ser His Lys Phe Met Ser Thr Ser Val Gly1 5 10 15Asp Arg Val Ser
Ile Ile Cys Lys Ala Ser Gln Asp Val Gly Thr Ala 20 25 30Val Asp Trp
Tyr Gln Gln Lys Pro Gly Gln Ser Pro Lys Leu Leu Ile 35 40 45Tyr Trp
Ala Ser Thr Arg His Thr Gly Val Pro Asp Arg Phe Thr Gly 50 55 60Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Thr Asn Val Gln Ser65 70 75
80Glu Asp Leu Ala Asp Tyr Phe Cys Gln Gln Tyr Asn Ser Tyr Pro Leu
85 90 95Thr Phe Gly Ala Gly Thr Met Leu Asp Leu Lys 100
10511130PRTArtificial SequenceSynthetic 111Glu Val Gln Leu Gln Gln
Ser Gly Pro Glu Leu Val Lys Pro Gly Thr1 5 10 15Ser Val Arg Ile Ser
Cys Lys Thr Ser Gly Tyr Thr Phe Thr 20 25 3011223PRTArtificial
SequenceSynthetic 112Asp Ile Val Met Thr Gln Ser His Lys Phe Met
Ser Thr Ser Val Gly1 5 10 15Asp Arg Val Ser Ile Ile Cys
201135PRTArtificial SequenceSynthetic 113Glu Tyr Thr Ile His1
511411PRTArtificial SequenceSynthetic 114Lys Ala Ser Gln Asp Val
Gly Thr Ala Val Asp1 5 1011514PRTArtificial SequenceSynthetic
115Trp Val Lys Gln Ser His Gly Lys Ser Leu Glu Trp Ile Gly1 5
1011615PRTArtificial SequenceSynthetic 116Trp Tyr Gln Gln Lys Pro
Gly Gln Ser Pro Lys Leu Leu Ile Tyr1 5 10 1511717PRTArtificial
SequenceSynthetic 117Asn Ile Asn Pro Asn Asn Gly Gly Thr Thr Tyr
Asn Gln Lys Phe Glu1 5 10 15Asp1187PRTArtificial SequenceSynthetic
118Trp Ala Ser Thr Arg His Thr1 511932PRTArtificial
SequenceSynthetic 119Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser
Thr Ala Tyr Met Glu1 5 10 15Leu Arg Ser Leu Thr Ser Glu Asp Ser Ala
Val Tyr Tyr Cys Ala Ala 20 25 3012032PRTArtificial
SequenceSynthetic 120Gly Val Pro Asp Arg Phe Thr Gly Ser Gly Ser
Gly Thr Asp Phe Thr1 5 10 15Leu Thr Ile Thr Asn Val Gln Ser Glu Asp
Leu Ala Asp Tyr Phe Cys 20 25 301216PRTArtificial SequenceSynthetic
121Gly Trp Asn Phe Asp Tyr1 51229PRTArtificial SequenceSynthetic
122Gln Gln Tyr Asn Ser Tyr Pro Leu Thr1 512311PRTArtificial
SequenceSynthetic 123Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser1 5
1012410PRTArtificial SequenceSynthetic 124Phe Gly Ala Gly Thr Met
Leu Asp Leu Lys1 5 10125119PRTArtificial SequenceSynthetic 125Asp
Val Lys Leu Val Glu Ser Gly Gly Gly Leu Val Lys Leu Gly Gly1 5 10
15Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30Tyr Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Leu
Val 35 40 45Ala Ala Ile Asn Ser Asp Gly Gly Ile Thr Tyr Tyr Leu Asp
Thr Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn
Thr Leu Tyr65 70 75 80Leu Gln Met Ser Ser Leu Lys Ser Glu Asp Thr
Ala Leu Phe Tyr Cys 85 90 95Ala Arg His Arg Ser Gly Tyr Phe Ser Met
Asp Tyr Trp Gly Gln Gly 100 105 110Thr Ser Val Thr Val Ser Ser
115126107PRTArtificial SequenceSynthetic 126Asp Ile Val Met Thr Gln
Ser Gln Arg Phe Met Ser Thr Thr Val Gly1 5 10 15Asp Arg Val Ser Ile
Thr Cys Lys Ala Ser Gln Asn Val Val Ser Ala 20 25 30Val Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ser Pro Lys Leu Leu Ile 35 40 45Tyr Ser Ala
Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg Phe Thr Gly 50 55 60Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Asn Met Gln Ser65 70 75
80Glu Asp Leu Ala Asp Phe Phe Cys Gln Gln Tyr Ser Asn Tyr Pro Trp
85 90 95Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys 100
10512730PRTArtificial SequenceSynthetic 127Asp Val Lys Leu Val Glu
Ser Gly Gly Gly Leu Val Lys Leu Gly Gly1 5 10 15Ser Leu Lys Leu Ser
Cys Ala Ala Ser Gly Phe Thr Phe Ser 20 25 3012823PRTArtificial
SequenceSynthetic 128Asp Ile Val Met Thr Gln Ser Gln Arg Phe Met
Ser Thr Thr Val Gly1 5 10 15Asp Arg Val Ser Ile Thr Cys
201295PRTArtificial SequenceSynthetic 129Asn Tyr Tyr Met Ser1
513011PRTArtificial SequenceSynthetic 130Lys Ala Ser Gln Asn Val
Val Ser Ala Val Ala1 5 1013114PRTArtificial SequenceSynthetic
131Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Leu Val Ala1 5
1013215PRTArtificial SequenceSynthetic 132Trp Tyr Gln Gln Lys Pro
Gly Gln Ser Pro Lys Leu Leu Ile Tyr1 5 10 1513317PRTArtificial
SequenceSynthetic 133Ala Ile Asn Ser Asp Gly Gly Ile Thr Tyr Tyr
Leu Asp Thr Val Lys1 5 10 15Gly1347PRTArtificial SequenceSynthetic
134Ser Ala Ser Asn Arg Tyr Thr1 513532PRTArtificial
SequenceSynthetic 135Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn
Thr Leu Tyr Leu Gln1 5 10 15Met Ser Ser Leu Lys Ser Glu Asp Thr Ala
Leu Phe Tyr Cys Ala Arg 20 25 3013632PRTArtificial
SequenceSynthetic 136Gly Val Pro Asp Arg Phe Thr Gly Ser Gly Ser
Gly Thr Asp Phe Thr1 5 10 15Leu Thr Ile Ser Asn Met Gln Ser Glu Asp
Leu Ala Asp Phe Phe Cys 20 25 3013710PRTArtificial
SequenceSynthetic 137His Arg Ser Gly Tyr Phe Ser Met Asp Tyr1 5
101389PRTArtificial SequenceSynthetic 138Gln Gln Tyr Ser Asn Tyr
Pro Trp Thr1 513911PRTArtificial SequenceSynthetic 139Trp Gly Gln
Gly Thr Ser Val Thr Val Ser Ser1 5 1014010PRTArtificial
SequenceSynthetic 140Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys1 5
10141117PRTArtificial SequenceSynthetic 141Gln Val Gln Leu Gln Gln
Pro Gly Ala Glu Leu Val Lys Ser Gly Ala1 5 10 15Ser Val Asn Leu Ser
Cys Arg Ala Ser Gly Tyr Thr Phe Thr Arg Tyr 20 25 30Trp Met Leu Trp
Val Arg Gln Arg Pro Gly His Gly Leu Glu Trp Val 35 40 45Gly Glu Ile
Asn Pro Arg Asn Asp Arg Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys Thr
Lys Ala Thr Leu Thr Val Asp Arg Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Thr Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Phe Cys
85 90 95Ala Leu Gly Gly Gly Tyr Ala Met Asp Tyr Trp Gly Gln Gly Thr
Ser 100 105 110Val Thr Val Ser Ser 115142107PRTArtificial
SequenceSynthetic 142Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu
Ser Ala Phe Leu Gly1 5 10 15Asp Arg Val Thr Ile Ser Cys Arg Ala Ser
Gln Asp Ile Ser Asn Tyr 20 25 30Leu Asn Trp Tyr Gln Gln Lys Pro
Asp Gly Thr Val Lys Leu Leu Ile 35 40 45Tyr Tyr Thr Ser Arg Leu His
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp
Tyr Ser Leu Thr Ile Ser Asn Leu Glu Gln65 70 75 80Glu Asp Phe Ala
Thr Tyr Phe Cys Gln Gln Gly Asn Thr Leu Pro Trp 85 90 95Thr Phe Gly
Gly Gly Thr Lys Leu Glu Ile Lys 100 10514330PRTArtificial
SequenceSynthetic 143Gln Val Gln Leu Gln Gln Pro Gly Ala Glu Leu
Val Lys Ser Gly Ala1 5 10 15Ser Val Asn Leu Ser Cys Arg Ala Ser Gly
Tyr Thr Phe Thr 20 25 3014423PRTArtificial SequenceSynthetic 144Asp
Ile Gln Met Thr Gln Thr Thr Ser Ser Leu Ser Ala Phe Leu Gly1 5 10
15Asp Arg Val Thr Ile Ser Cys 201455PRTArtificial SequenceSynthetic
145Arg Tyr Trp Met Leu1 514611PRTArtificial SequenceSynthetic
146Arg Ala Ser Gln Asp Ile Ser Asn Tyr Leu Asn1 5
1014714PRTArtificial SequenceSynthetic 147Trp Val Arg Gln Arg Pro
Gly His Gly Leu Glu Trp Val Gly1 5 1014815PRTArtificial
SequenceSynthetic 148Trp Tyr Gln Gln Lys Pro Asp Gly Thr Val Lys
Leu Leu Ile Tyr1 5 10 1514917PRTArtificial SequenceSynthetic 149Glu
Ile Asn Pro Arg Asn Asp Arg Thr Asn Tyr Asn Glu Lys Phe Lys1 5 10
15Thr1507PRTArtificial SequenceSynthetic 150Tyr Thr Ser Arg Leu His
Ser1 515132PRTArtificial SequenceSynthetic 151Lys Ala Thr Leu Thr
Val Asp Arg Ser Ser Ser Thr Ala Tyr Met Gln1 5 10 15Leu Thr Ser Leu
Thr Ser Glu Asp Ser Ala Val Tyr Phe Cys Ala Leu 20 25
3015232PRTArtificial SequenceSynthetic 152Gly Val Pro Ser Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Tyr Ser1 5 10 15Leu Thr Ile Ser Asn
Leu Glu Gln Glu Asp Phe Ala Thr Tyr Phe Cys 20 25
301538PRTArtificial SequenceSynthetic 153Gly Gly Gly Tyr Ala Met
Asp Tyr1 51549PRTArtificial SequenceSynthetic 154Gln Gln Gly Asn
Thr Leu Pro Trp Thr1 515511PRTArtificial SequenceSynthetic 155Trp
Gly Gln Gly Thr Ser Val Thr Val Ser Ser1 5 1015610PRTArtificial
SequenceSynthetic 156Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys1 5
10157117PRTArtificial SequenceSynthetic 157Glu Val Gln Leu Val Gln
Ser Gly Gly Gly Leu Val Gln Ser Gly Gly1 5 10 15Ser Leu Arg Leu Ser
Cys Arg Ala Ser Gly Tyr Thr Phe Thr Arg Tyr 20 25 30Trp Met Leu Trp
Val Arg Gln Arg Pro Gly His Gly Leu Glu Trp Val 35 40 45Gly Glu Ile
Asn Pro Arg Asn Asp Arg Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys Thr
Arg Phe Thr Ile Ser Val Asp Arg Ser Lys Ser Thr Ala Tyr65 70 75
80Leu Gln Met Asp Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Phe Cys
85 90 95Ala Leu Gly Gly Gly Tyr Ala Met Asp Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115158107PRTArtificial
SequenceSynthetic 158Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser
Gln Asp Ile Ser Asn Tyr 20 25 30Leu Asn Trp Tyr Gln Gln Lys Pro Gly
Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Tyr Thr Ser Arg Leu His Ser
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Tyr
Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr
Tyr Phe Cys Gln Gln Gly Asn Thr Leu Pro Trp 85 90 95Thr Phe Gly Gly
Gly Thr Lys Leu Glu Ile Lys 100 10515930PRTArtificial
SequenceSyntheti 159Glu Val Gln Leu Val Gln Ser Gly Gly Gly Leu Val
Gln Ser Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Arg Ala Ser Gly Tyr
Thr Phe Thr 20 25 3016023PRTArtificial SequenceSyntheti 160Asp Ile
Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp
Arg Val Thr Ile Thr Cys 201615PRTArtificial SequenceSynthetic
161Arg Tyr Trp Met Leu1 516211PRTArtificial SequenceSynthetic
162Arg Ala Ser Gln Asp Ile Ser Asn Tyr Leu Asn1 5
1016314PRTArtificial SequenceSynthetic 163Trp Val Arg Gln Arg Pro
Gly His Gly Leu Glu Trp Val Gly1 5 1016415PRTArtificial
SequenceSynthetic 164Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys
Leu Leu Ile Tyr1 5 10 1516517PRTArtificial SequenceSynthetic 165Glu
Ile Asn Pro Arg Asn Asp Arg Thr Asn Tyr Asn Glu Lys Phe Lys1 5 10
15Thr1667PRTArtificial SequenceSynthetic 166Tyr Thr Ser Arg Leu His
Ser1 516732PRTArtificial SequenceSynthetic 167Arg Phe Thr Ile Ser
Val Asp Arg Ser Lys Ser Thr Ala Tyr Leu Gln1 5 10 15Met Asp Ser Leu
Arg Ala Glu Asp Thr Ala Val Tyr Phe Cys Ala Leu 20 25
3016832PRTArtificial SequenceSynthetic 168Gly Val Pro Ser Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Tyr Thr1 5 10 15Leu Thr Ile Ser Ser
Leu Gln Pro Glu Asp Phe Ala Thr Tyr Phe Cys 20 25
301698PRTArtificial SequenceSynthetic 169Gly Gly Gly Tyr Ala Met
Asp Tyr1 51709PRTArtificial SequenceSynthetic 170Gln Gln Gly Asn
Thr Leu Pro Trp Thr1 517111PRTArtificial SequenceSynthetic 171Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser1 5 1017210PRTArtificial
SequenceSynthetic 172Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys1 5
10173117PRTArtificial SequenceSynthetic 173Glu Val Gln Leu Val Gln
Ser Gly Gly Gly Leu Val Gln Ser Gly Gly1 5 10 15Ser Leu Arg Leu Ser
Cys Arg Ala Ser Gly Tyr Thr Phe Thr Arg Tyr 20 25 30Trp Met Leu Trp
Val Arg Gln Arg Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Glu Ile
Asn Pro Arg Asn Asp Arg Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys Thr
Arg Phe Thr Ile Ser Val Asp Arg Ser Lys Asn Thr Leu Tyr65 70 75
80Leu Gln Met Asp Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Phe Cys
85 90 95Ala Leu Gly Gly Gly Tyr Ala Met Asp Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115174107PRTArtificial
SequenceSynthetic 174Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser
Gln Asp Ile Ser Asn Tyr 20 25 30Leu Asn Trp Tyr Gln Gln Lys Pro Gly
Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Tyr Thr Ser Arg Leu His Ser
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Tyr
Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr
Tyr Phe Cys Gln Gln Gly Asn Thr Leu Pro Trp 85 90 95Thr Phe Gly Gly
Gly Thr Lys Leu Glu Ile Lys 100 10517530PRTArtificial
SequenceSynthetic 175Glu Val Gln Leu Val Gln Ser Gly Gly Gly Leu
Val Gln Ser Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Arg Ala Ser Gly
Tyr Thr Phe Thr 20 25 3017623PRTArtificial SequenceSynthetic 176Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys 201775PRTArtificial SequenceSynthetic
177Arg Tyr Trp Met Leu1 517811PRTArtificial SequenceSynthetic
178Arg Ala Ser Gln Asp Ile Ser Asn Tyr Leu Asn1 5
1017914PRTArtificial SequenceSynthetic 179Trp Val Arg Gln Arg Pro
Gly Lys Gly Leu Glu Trp Val Ala1 5 1018015PRTArtificial
SequenceSynthetic 180Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys
Leu Leu Ile Tyr1 5 10 1518117PRTArtificial SequenceSynthetic 181Glu
Ile Asn Pro Arg Asn Asp Arg Thr Asn Tyr Asn Glu Lys Phe Lys1 5 10
15Thr1827PRTArtificial SequenceSynthetic 182Tyr Thr Ser Arg Leu His
Ser1 518332PRTArtificial SequenceSynthetic 183Arg Phe Thr Ile Ser
Val Asp Arg Ser Lys Asn Thr Leu Tyr Leu Gln1 5 10 15Met Asp Ser Leu
Arg Ala Glu Asp Thr Ala Val Tyr Phe Cys Ala Leu 20 25
3018432PRTArtificial SequenceSynthetic 184Gly Val Pro Ser Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Tyr Thr1 5 10 15Leu Thr Ile Ser Ser
Leu Gln Pro Glu Asp Phe Ala Thr Tyr Phe Cys 20 25
301858PRTArtificial SequenceSynthetic 185Gly Gly Gly Tyr Ala Met
Asp Tyr1 51869PRTArtificial SequenceSynthetic 186Gln Gln Gly Asn
Thr Leu Pro Trp Thr1 518711PRTArtificial SequenceSynthetic 187Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser1 5 1018810PRTArtificial
SequenceSynthetic 188Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys1 5
1018915PRTArtificial SequenceSynthetic 189Leu Val Ala Leu Tyr Leu
Ala Ala Arg Leu Ser Trp Asn Gln Val1 5 10 1519015PRTArtificial
SequenceSynthetic 190Gly Ala Leu Leu Arg Val Tyr Val Pro Arg Ser
Ser Leu Pro Gly1 5 10 1519115PRTArtificial SequenceSynthetic 191Ile
Arg Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Arg Ser Ser1 5 10
1519215PRTArtificial SequenceSynthetic 192Arg Gln Pro Arg Gly Trp
Glu Gln Leu Glu Gln Cys Gly Tyr Pro1 5 10 1519315PRTArtificial
SequenceSynthetic 193Leu Pro Leu Glu Thr Phe Thr Arg His Arg Gln
Pro Arg Gly Trp1 5 10 1519415PRTArtificial SequenceSynthetic 194Trp
Arg Gly Phe Tyr Ile Ala Gly Asp Pro Ala Leu Ala Tyr Gly1 5 10
1519518PRTArtificial SequenceSynthetic 195Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro1 5 10 15Leu
Ala19615PRTArtificial SequenceSynthetic 196Thr Val Glu Arg Leu Leu
Gln Ala His Arg Gln Leu Glu Glu Arg1 5 10 1519718PRTArtificial
SequenceSynthetic 197Phe Val Gly Tyr His Gly Thr Phe Leu Glu Ala
Ala Gln Ser Ile Val1 5 10 15Phe Gly19815PRTArtificial
SequenceSynthetic 198Ile Arg Asn Gly Ala Leu Leu Arg Val Tyr Val
Pro Arg Ser Ser1 5 10 1519915PRTArtificial SequenceSynthetic 199Ala
Arg Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Arg Ser Ser1 5 10
1520015PRTArtificial SequenceSynthetic 200Ile Ala Asn Gly Ala Leu
Leu Arg Val Tyr Val Pro Arg Ser Ser1 5 10 1520115PRTArtificial
SequenceSynthetic 201Ile Arg Ala Gly Ala Leu Leu Arg Val Tyr Val
Pro Arg Ser Ser1 5 10 1520215PRTArtificial SequenceSynthetic 202Ile
Arg Asn Ala Ala Leu Leu Arg Val Tyr Val Pro Arg Ser Ser1 5 10
1520315PRTArtificial SequenceSynthetic 203Ile Arg Asn Gly Ala Ala
Leu Arg Val Tyr Val Pro Arg Ser Ser1 5 10 1520415PRTArtificial
SequenceSynthetic 204Ile Arg Asn Gly Ala Leu Ala Arg Val Tyr Val
Pro Arg Ser Ser1 5 10 1520515PRTArtificial SequenceSynthetic 205Ile
Arg Asn Gly Ala Leu Leu Ala Val Tyr Val Pro Arg Ser Ser1 5 10
1520615PRTArtificial SequenceSynthetic 206Ile Arg Asn Gly Ala Leu
Leu Arg Ala Tyr Val Pro Arg Ser Ser1 5 10 1520715PRTArtificial
SequenceSynthetic 207Ile Arg Asn Gly Ala Leu Leu Arg Val Ala Val
Pro Arg Ser Ser1 5 10 1520815PRTArtificial SequenceSynthetic 208Ile
Arg Asn Gly Ala Leu Leu Arg Val Tyr Ala Pro Arg Ser Ser1 5 10
1520915PRTArtificial SequenceSynthetic 209Ile Arg Asn Gly Ala Leu
Leu Arg Val Tyr Val Ala Arg Ser Ser1 5 10 1521015PRTArtificial
SequenceSynthetic 210Ile Arg Asn Gly Ala Leu Leu Arg Val Tyr Val
Pro Ala Ser Ser1 5 10 1521115PRTArtificial SequenceSynthetic 211Ile
Arg Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Arg Ala Ser1 5 10
1521215PRTArtificial SequenceSynthetic 212Ile Arg Asn Gly Ala Leu
Leu Arg Val Tyr Val Pro Arg Ser Ala1 5 10 1521315PRTArtificial
SequenceSynthetic 213Gly Ala Leu Leu Arg Val Tyr Val Pro Arg Ser
Ser Leu Pro Gly1 5 10 1521415PRTArtificial SequenceSynthetic 214Ala
Ala Leu Leu Arg Val Tyr Val Pro Arg Ser Ser Leu Pro Gly1 5 10
1521515PRTArtificial SequenceSynthetic 215Gly Gly Leu Leu Arg Val
Tyr Val Pro Arg Ser Ser Leu Pro Gly1 5 10 1521615PRTArtificial
SequenceSynthetic 216Gly Ala Ala Leu Arg Val Tyr Val Pro Arg Ser
Ser Leu Pro Gly1 5 10 1521715PRTArtificial SequenceSynthetic 217Gly
Ala Leu Ala Arg Val Tyr Val Pro Arg Ser Ser Leu Pro Gly1 5 10
1521815PRTArtificial SequenceSynthetic 218Gly Ala Leu Leu Ala Val
Tyr Val Pro Arg Ser Ser Leu Pro Gly1 5 10 1521915PRTArtificial
SequenceSynthetic 219Gly Ala Leu Leu Arg Ala Tyr Val Pro Arg Ser
Ser Leu Pro Gly1 5 10 1522015PRTArtificial SequenceSynthetic 220Gly
Ala Leu Leu Arg Val Ala Val Pro Arg Ser Ser Leu Pro Gly1 5 10
1522115PRTArtificial SequenceSynthetic 221Gly Ala Leu Leu Arg Val
Tyr Ala Pro Arg Ser Ser Leu Pro Gly1 5 10 1522215PRTArtificial
SequenceSynthetic 222Gly Ala Leu Leu Arg Val Tyr Val Ala Arg Ser
Ser Leu Pro Gly1 5 10 1522315PRTArtificial SequenceSynthetic 223Gly
Ala Leu Leu Arg Val Tyr Val Pro Ala Ser Ser Leu Pro Gly1 5 10
1522415PRTArtificial SequenceSynthetic 224Gly Ala Leu Leu Arg Val
Tyr Val Pro Arg Ala Ser Leu Pro Gly1 5 10 1522512PRTArtificial
SequenceSynthetic 225Gly Ala Leu Leu Arg Val Tyr Val Pro Arg Ser
Ala1 5 1022615PRTArtificial SequenceSynthetic 226Gly Ala Leu Leu
Arg Val Tyr Val Pro Arg Ser Ser Ala Pro Gly1 5 10
1522715PRTArtificial SequenceSynthetic 227Gly Ala Leu Leu Arg Val
Tyr Val Pro Arg Ser Ser Leu Ala Gly1 5 10 1522815PRTArtificial
SequenceSynthetic 228Trp Arg Gly Phe Tyr Ile Ala Gly Asp Pro Ala
Leu Ala Tyr Gly1 5 10 1522915PRTArtificial SequenceSynthetic 229Trp
Arg Gly Ala Tyr Ile Ala Gly Asp Pro Ala Leu Ala Tyr Gly1 5 10
1523015PRTArtificial SequenceSynthetic 230Trp Arg Gly Phe Ala Ile
Ala Gly Asp Pro Ala Leu Ala Tyr Gly1 5 10 1523115PRTArtificial
SequenceSynthetic 231Trp Arg Gly Phe Tyr Ala Ala Gly Asp Pro Ala
Leu Ala Tyr Gly1 5 10 1523215PRTArtificial SequenceSynthetic 232Trp
Arg Gly Phe Tyr Ile Gly Gly Asp Pro Ala Leu Ala Tyr Gly1 5 10
1523315PRTArtificial SequenceSynthetic 233Trp Arg Gly Phe Tyr Ile
Ala Gly Asp Ala Ala Leu Ala Tyr Gly1 5 10 1523415PRTArtificial
SequenceSynthetic 234Trp Arg Gly Phe Tyr Ile Ala Gly Asp Pro Gly
Leu Ala Tyr Gly1 5 10 1523515PRTArtificial SequenceSynthetic 235Trp
Arg Gly Phe Tyr Ile Ala Gly Asp Pro Ala Ala Ala Tyr Gly1 5 10
1523615PRTArtificial SequenceSynthetic 236Trp Arg Gly Phe Tyr
Ile Ala Gly Asp Pro Ala Leu Gly Tyr Gly1 5 10 1523715PRTArtificial
SequenceSynthetic 237Trp Arg Gly Phe Tyr Ile Ala Gly Asp Pro Ala
Leu Ala Ala Gly1 5 10 1523818PRTArtificial SequenceSynthetic 238Gly
Pro Glu Glu Glu Gly Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro1 5 10
15Leu Ala23918PRTArtificial SequenceSynthetic 239Gly Pro Glu Ala
Glu Gly Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro1 5 10 15Leu
Ala24018PRTArtificial SequenceSynthetic 240Gly Pro Glu Glu Ala Gly
Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro1 5 10 15Leu
Ala24118PRTArtificial SequenceSynthetic 241Gly Pro Glu Glu Glu Gly
Gly Ala Leu Glu Thr Ile Leu Gly Trp Pro1 5 10 15Leu
Ala24218PRTArtificial SequenceSynthetic 242Gly Pro Glu Glu Glu Gly
Gly Arg Ala Glu Thr Ile Leu Gly Trp Pro1 5 10 15Leu
Ala24318PRTArtificial SequenceSynthetic 243Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Ala Ile Leu Gly Trp Pro1 5 10 15Leu
Ala24418PRTArtificial SequenceSynthetic 244Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ala Leu Gly Trp Pro1 5 10 15Leu
Ala24518PRTArtificial SequenceSynthetic 245Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ile Ala Gly Trp Pro1 5 10 15Leu
Ala24618PRTArtificial SequenceSynthetic 246Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ile Leu Gly Ala Pro1 5 10 15Leu
Ala24718PRTArtificial SequenceSynthetic 247Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ile Leu Gly Trp Ala1 5 10 15Leu
Ala24818PRTArtificial SequenceSynthetic 248Gly Pro Glu Glu Glu Gly
Gly Arg Leu Glu Thr Ile Leu Gly Trp Pro1 5 10 15Ala
Ala24915PRTArtificial SequenceSynthetic 249Thr Val Glu Arg Leu Leu
Gln Ala His Arg Gln Leu Glu Glu Arg1 5 10 1525015PRTArtificial
SequenceSynthetic 250Thr Val Glu Ala Leu Leu Gln Ala His Arg Gln
Leu Glu Glu Arg1 5 10 1525115PRTArtificial SequenceSynthetic 251Thr
Val Glu Arg Ala Leu Gln Ala His Arg Gln Leu Glu Glu Arg1 5 10
1525215PRTArtificial SequenceSynthetic 252Thr Val Glu Arg Leu Ala
Gln Ala His Arg Gln Leu Glu Glu Arg1 5 10 1525315PRTArtificial
SequenceSynthetic 253Thr Val Glu Arg Leu Leu Gln Gly His Arg Gln
Leu Glu Glu Arg1 5 10 1525415PRTArtificial SequenceSynthetic 254Thr
Val Glu Arg Leu Leu Gln Ala His Ala Gln Leu Glu Glu Arg1 5 10
1525515PRTArtificial SequenceSynthetic 255Thr Val Glu Arg Leu Leu
Gln Ala His Arg Gln Ala Glu Glu Arg1 5 10 1525615PRTArtificial
SequenceSynthetic 256Thr Val Glu Arg Leu Leu Gln Ala His Arg Gln
Leu Ala Glu Arg1 5 10 1525715PRTArtificial SequenceSynthetic 257Thr
Val Glu Arg Leu Leu Gln Ala His Arg Gln Leu Glu Glu Ala1 5 10
1525818PRTArtificial SequenceSynthetic 258Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly25918PRTArtificial SequenceSynthetic 259Ala Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26018PRTArtificial SequenceSynthetic 260Phe Ala Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26118PRTArtificial SequenceSynthetic 261Phe Val Ala Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26218PRTArtificial SequenceSynthetic 262Phe Val Gly Ala His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26318PRTArtificial SequenceSynthetic 263Phe Val Gly Tyr Ala Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26418PRTArtificial SequenceSynthetic 264Phe Val Gly Tyr His Gly
Ala Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26518PRTArtificial SequenceSynthetic 265Phe Val Gly Tyr His Gly
Thr Ala Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26618PRTArtificial SequenceSynthetic 266Phe Val Gly Tyr His Gly
Thr Phe Ala Glu Ala Ala Gln Ser Ile Val1 5 10 15Phe
Gly26718PRTArtificial SequenceSynthetic 267Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Gly Ala Gln Ser Ile Val1 5 10 15Phe
Gly26818PRTArtificial SequenceSynthetic 268Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Gly Gln Ser Ile Val1 5 10 15Phe
Gly26918PRTArtificial SequenceSynthetic 269Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ala Ile Val1 5 10 15Phe
Gly27018PRTArtificial SequenceSynthetic 270Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ala Val1 5 10 15Phe
Gly27118PRTArtificial SequenceSynthetic 271Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Ala1 5 10 15Phe
Gly27218PRTArtificial SequenceSynthetic 272Phe Val Gly Tyr His Gly
Thr Phe Leu Glu Ala Ala Gln Ser Ile Val1 5 10 15Ala
Gly27342PRTArtificial SequenceSynthetic 273Asn Thr Lys Val Asp Lys
Lys Val Glu Pro Lys Ser Cys Lys Ala Ser1 5 10 15Gly Gly Arg His Arg
Gln Pro Arg Gly Trp Glu Gln Leu Gly Gly Ser 20 25 30Pro Thr Gly Ala
Glu Phe Leu Gly Asp Gly 35 402749PRTArtificial SequenceSynthetic
274Trp Glu Gln Leu Gly Gly Ser Pro Thr1 52759PRTArtificial
SequenceSynthetic 275Val Glu Pro Lys Ser Cys Lys Ala Ser1
52764PRTArtificial SequenceSynthetic 276Asp Lys Thr
His12773PRTArtificial SequenceSynthetic 277Gly Gly
Gly127849PRTArtificial SequenceSynthetic 278Asn Thr Lys Val Asp Lys
Lys Val Glu Pro Lys Ser Cys Asp Lys Thr1 5 10 15His Lys Ala Ser Gly
Gly Arg His Arg Gln Pro Arg Gly Trp Glu Gln 20 25 30Leu Gly Gly Gly
Gly Gly Ser Pro Thr Gly Ala Glu Phe Leu Gly Asp 35 40
45Gly2799PRTArtificial SequenceSynthetic 279Trp Glu Gln Leu Gly Gly
Gly Gly Gly1 52809PRTArtificial SequenceSynthetic 280Leu Gly Gly
Gly Gly Gly Ser Pro Thr1 528119PRTArtificial SequenceSynthetic
281Lys Ala Ser Gly Gly Arg His Arg Gln Pro Arg Gly Trp Glu Gln Leu1
5 10 15Gly Gly Ser2829PRTArtificial SequenceSynthetic 282Asp Lys
Thr His Lys Ala Ser Gly Gly1 52833PRTArtificial SequenceSynthetic
283Gly Gly Ser12846PRTArtificial SequenceSynthetic 284Gly Gly Gly
Gly Gly Ser1 5285219PRTArtificial SequenceSynthetic 285Pro Thr Gly
Ala Glu Phe Leu Gly Asp Gly Gly Asp Val Ser Phe Ser1 5 10 15Thr Arg
Gly Thr Gln Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His 20 25 30Ala
Gln Leu Glu Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr 35 40
45Ala Leu Glu Ala Ala Gln Ser Ile Val Phe Gly Gly Val Ala Ala Arg
50 55 60Ser Gln Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala Gly
Asp65 70 75 80Pro Ala His Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro
Asp Ala Ala 85 90 95Gly Arg Ile Ala Asn Gly Ala Leu Leu Arg Val Tyr
Val Pro Ala Ser 100 105 110Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu
Thr Leu Ala Ala Pro Glu 115 120 125Ala Ala Gly Glu Val Glu Arg Leu
Ile Gly His Pro Leu Pro Leu Ala 130 135 140Leu Asp Ala Ile Thr Gly
Pro Glu Glu Glu Gly Gly Arg Glu Glu Thr145 150 155 160Ile Leu Gly
Trp Pro Leu Ala Glu Arg Thr Val Val Ile Pro Ser Ala 165 170 175Ile
Pro Thr Asp Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser 180 185
190Ile Pro Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser
195 200 205Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 210
215286219PRTArtificial SequenceSynthetic 286Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly Asp Val Ser Phe Ser1 5 10 15Thr Arg Gly Thr Gln
Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His 20 25 30Ala Gln Leu Glu
Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr 35 40 45Ala Leu Glu
Ala Ala Gln Ser Ile Val Phe Gly Gly Val Ala Ala Arg 50 55 60Ser Gln
Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala Gly Asp65 70 75
80Pro Ala His Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Ala
85 90 95Gly Arg Ile Ala Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Ala
Ser 100 105 110Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu Thr Leu Ala
Ala Pro Glu 115 120 125Ala Ala Gly Glu Val Glu Arg Leu Ile Gly His
Pro Leu Pro Leu Ala 130 135 140Leu Asp Ala Ile Thr Gly Pro Glu Glu
Glu Gly Gly Arg Asn Glu Thr145 150 155 160Ile Leu Gly Trp Pro Leu
Ala Glu Arg Thr Val Val Ile Pro Ser Ala 165 170 175Ile Pro Thr Asp
Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser 180 185 190Ile Pro
Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser 195 200
205Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 210
215287219PRTArtificial SequenceSynthetic 287Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly Asp Val Ser Phe Ser1 5 10 15Thr Arg Gly Thr Gln
Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His 20 25 30Ala Gln Leu Glu
Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr 35 40 45Ala Leu Glu
Ala Ala Gln Ser Ile Val Phe Gly Gly Val Arg Ala Arg 50 55 60Ser Gln
Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala Gly Asp65 70 75
80Pro Ala His Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Ala
85 90 95Gly Arg Ile Ala Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Ala
Ser 100 105 110Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu Thr Leu Ala
Ala Pro Glu 115 120 125Ala Ala Gly Glu Val Glu Arg Leu Ile Gly His
Pro Leu Pro Leu Ala 130 135 140Leu Asp Ala Ile Thr Gly Pro Glu Glu
Glu Gly Gly Arg Glu Glu Thr145 150 155 160Ile Leu Gly Trp Pro Leu
Ala Glu Arg Thr Val Val Ile Pro Ser Ala 165 170 175Ile Pro Thr Asp
Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser 180 185 190Ile Pro
Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser 195 200
205Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 210
215288219PRTArtificial SequenceSynthetic 288Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly Asp Val Ser Phe Ser1 5 10 15Thr Arg Gly Thr Gln
Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His 20 25 30Ala Gln Leu Glu
Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr 35 40 45Ala Leu Glu
Ala Ala Gln Ser Ile Val Phe Gly Gly Val Arg Ala Arg 50 55 60Ser Gln
Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala Gly Asp65 70 75
80Pro Ala His Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Ala
85 90 95Gly Arg Ile Ala Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Ala
Ser 100 105 110Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu Thr Leu Ala
Ala Pro Glu 115 120 125Ala Ala Gly Glu Val Glu Arg Leu Ile Gly His
Pro Leu Pro Leu Ala 130 135 140Leu Asp Ala Ile Thr Gly Pro Glu Glu
Glu Gly Gly Arg Asn Glu Thr145 150 155 160Ile Leu Gly Trp Pro Leu
Ala Glu Arg Thr Val Val Ile Pro Ser Ala 165 170 175Ile Pro Thr Asp
Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser 180 185 190Ile Pro
Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser 195 200
205Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 210
215289219PRTArtificial SequenceSynthetic 289Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly Asp Val Ser Phe Ser1 5 10 15Thr Arg Gly Thr Gln
Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His 20 25 30Ala Gln Leu Glu
Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr 35 40 45Ala Leu Glu
Ala Ala Gln Ser Ile Val Phe Gly Gly Val Arg Ala Arg 50 55 60Ser Gln
Asp Leu Asp Ala Ile Trp Arg Gly Phe Tyr Ile Ala Gly Asp65 70 75
80Pro Ala His Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Arg
85 90 95Gly Arg Ile Ala Asn Gly Ala Leu Leu Arg Val Tyr Val Pro Ala
Ser 100 105 110Ser Leu Pro Gly Phe Tyr Arg Thr Ser Leu Thr Leu Ala
Ala Pro Glu 115 120 125Ala Ala Gly Glu Val Glu Arg Leu Ile Gly His
Pro Leu Pro Leu Arg 130 135 140Leu Asp Ala Ile Thr Gly Pro Glu Glu
Glu Gly Gly Arg Glu Glu Thr145 150 155 160Ile Leu Gly Trp Pro Leu
Ala Glu Arg Thr Val Val Ile Pro Ser Ala 165 170 175Ile Pro Thr Asp
Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser 180 185 190Ile Pro
Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser 195 200
205Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys 210
215290113PRTArtificial SequenceSynthetic 290Asp Ile Val Leu Thr Gln
Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile
Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Arg Asn Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys
Leu Leu Ile Phe Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp
Arg Phe Ser Gly Ser Gly Phe Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Tyr Phe Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys291473PRTArtificial SequenceSynthetic 291Met Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly1 5 10 15Ala
Ser Val Lys Val Ser Cys Lys Thr Ser Arg Tyr Thr Phe Thr Glu 20 25
30Tyr Thr Ile His Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp
35 40 45Ile Gly Gly Ile Asn Pro Asn Asn Gly Ile Pro Asn Tyr Asn Gln
Lys 50 55 60Phe Lys Gly Arg Val Thr Ile Thr Val Asp Thr Ser Ala Ser
Thr Ala65 70 75 80Tyr Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr 85 90 95Cys Ala Arg Arg Arg Ile Ala Tyr Gly Tyr Asp
Glu Gly His Ala Met 100 105 110Asp Tyr Trp Gly Gln Gly Thr Leu Val
Thr Val Ser Ser Ala Ser Thr 115 120 125Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Ser Ser Lys Ser Thr Ser 130 135 140Gly Gly Thr Ala Ala
Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu145 150 155 160Pro Val
Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His 165 170
175Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser
Ser
180 185 190Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr
Ile Cys 195 200 205Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp
Lys Lys Val Glu 210 215 220Pro Lys Ser Cys Asp Lys Thr His Lys Ala
Ser Gly Gly Arg His Arg225 230 235 240Gln Pro Arg Gly Trp Glu Gln
Leu Gly Gly Gly Gly Gly Ser Pro Thr 245 250 255Gly Ala Glu Phe Leu
Gly Asp Gly Gly Asp Val Ser Phe Ser Thr Arg 260 265 270Gly Thr Gln
Asn Trp Thr Val Glu Arg Leu Leu Gln Ala His Ala Gln 275 280 285Leu
Glu Glu Arg Gly Tyr Val Phe Val Gly Tyr His Gly Thr Phe Leu 290 295
300Glu Ala Ala Gln Ser Ile Val Phe Gly Gly Val Ala Ala Arg Ser
Gln305 310 315 320Asp Leu Ala Ala Ile Trp Ala Gly Phe Tyr Ile Ala
Gly Asp Pro Ala 325 330 335Leu Ala Tyr Gly Tyr Ala Gln Asp Gln Glu
Pro Asp Ala Ala Gly Arg 340 345 350Ile Arg Asn Gly Ala Leu Leu Arg
Val Tyr Val Pro Ala Ser Ser Leu 355 360 365Pro Gly Phe Tyr Arg Thr
Ser Leu Thr Leu Ala Ala Pro Glu Ala Ala 370 375 380Gly Glu Val Glu
Arg Leu Ile Gly His Pro Leu Pro Leu Ala Leu Asp385 390 395 400Ala
Ile Thr Gly Pro Glu Glu Glu Gly Gly Arg Leu Glu Thr Ile Leu 405 410
415Gly Trp Pro Leu Ala Glu Arg Thr Val Val Ile Pro Ser Ala Ile Pro
420 425 430Thr Asp Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser
Ile Pro 435 440 445Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr
Ala Ser Gln Pro 450 455 460Gly Lys Pro Pro Arg Glu Asp Leu Lys465
470292473PRTArtificial SequenceSynthetic 292Met Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly1 5 10 15Ala Ser Val Lys Val
Ser Cys Lys Thr Ser Arg Tyr Thr Phe Thr Glu 20 25 30Tyr Thr Ile His
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp 35 40 45Ile Gly Gly
Ile Asn Pro Asn Asn Gly Ile Pro Asn Tyr Asn Gln Lys 50 55 60Phe Lys
Gly Arg Val Thr Ile Thr Val Asp Thr Ser Ala Ser Thr Ala65 70 75
80Tyr Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
85 90 95Cys Ala Arg Arg Arg Ile Ala Tyr Gly Tyr Asp Glu Gly His Ala
Met 100 105 110Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser
Ala Ser Thr 115 120 125Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser
Ser Lys Ser Thr Ser 130 135 140Gly Gly Thr Ala Ala Leu Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu145 150 155 160Pro Val Thr Val Ser Trp
Asn Ser Gly Ala Leu Thr Ser Gly Val His 165 170 175Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser 180 185 190Val Val
Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys 195 200
205Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu
210 215 220Pro Lys Ser Cys Asp Lys Thr His Lys Ala Ser Gly Gly Arg
His Arg225 230 235 240Gln Pro Arg Gly Trp Glu Gln Leu Gly Gly Gly
Gly Gly Ser Pro Thr 245 250 255Gly Ala Glu Phe Leu Gly Asp Gly Gly
Asp Val Ser Phe Ser Thr Arg 260 265 270Gly Thr Gln Asn Trp Thr Val
Glu Arg Leu Leu Gln Ala His Ala Gln 275 280 285Leu Glu Glu Arg Gly
Tyr Val Phe Val Gly Tyr His Gly Thr Ala Leu 290 295 300Glu Ala Ala
Gln Ser Ile Val Phe Gly Gly Val Arg Ala Arg Ser Gln305 310 315
320Asp Leu Asp Ala Ile Trp Arg Gly Phe Tyr Ile Ala Gly Asp Pro Ala
325 330 335His Ala Tyr Gly Tyr Ala Gln Asp Gln Glu Pro Asp Ala Arg
Gly Arg 340 345 350Ile Ala Asn Gly Ala Leu Leu Arg Val Tyr Val Pro
Ala Ser Ser Leu 355 360 365Pro Gly Phe Tyr Arg Thr Ser Leu Thr Leu
Ala Ala Pro Glu Ala Ala 370 375 380Gly Glu Val Glu Arg Leu Ile Gly
His Pro Leu Pro Leu Arg Leu Asp385 390 395 400Ala Ile Thr Gly Pro
Glu Glu Glu Gly Gly Arg Glu Glu Thr Ile Leu 405 410 415Gly Trp Pro
Leu Ala Glu Arg Thr Val Val Ile Pro Ser Ala Ile Pro 420 425 430Thr
Asp Pro Arg Asn Val Gly Gly Asp Leu Asp Pro Ser Ser Ile Pro 435 440
445Asp Lys Glu Gln Ala Ile Ser Ala Leu Pro Asp Tyr Ala Ser Gln Pro
450 455 460Gly Lys Pro Pro Arg Glu Asp Leu Lys465
470293221PRTArtificial SequenceSynthetic 293Met Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu1 5 10 15Gly Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr 20 25 30Ser Arg Asn Gln
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly 35 40 45Gln Pro Pro
Lys Leu Leu Ile Phe Trp Ala Ser Thr Arg Glu Ser Gly 50 55 60Val Pro
Asp Arg Phe Ser Gly Ser Gly Phe Gly Thr Asp Phe Thr Leu65 70 75
80Thr Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln
85 90 95Gln Tyr Phe Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val
Glu 100 105 110Ile Lys Arg Thr Val Ala Ala Pro Ser Val Phe Ile Phe
Pro Pro Ser 115 120 125Asp Glu Gln Leu Lys Ser Gly Thr Ala Ser Val
Val Cys Leu Leu Asn 130 135 140Asn Phe Tyr Pro Arg Glu Ala Lys Val
Gln Trp Lys Val Asp Asn Ala145 150 155 160Leu Gln Ser Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Asp Ser Lys 165 170 175Asp Ser Thr Tyr
Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp 180 185 190Tyr Glu
Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu 195 200
205Ser Ser Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 210 215
220294221PRTArtificial SequenceSynthetic 294Met Asp Ile Val Leu Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu1 5 10 15Gly Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr 20 25 30Ser Arg Asn Gln
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly 35 40 45Gln Pro Pro
Lys Leu Leu Ile Phe Trp Ala Ser Thr Arg Glu Ser Gly 50 55 60Val Pro
Asp Arg Phe Ser Gly Ser Gly Phe Gly Thr Asp Phe Thr Leu65 70 75
80Thr Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln
85 90 95Gln Tyr Phe Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val
Glu 100 105 110Ile Lys Arg Thr Val Ala Ala Pro Ser Val Phe Ile Phe
Pro Pro Ser 115 120 125Asp Glu Gln Leu Lys Ser Gly Thr Ala Ser Val
Val Cys Leu Leu Asn 130 135 140Asn Phe Tyr Pro Arg Glu Ala Lys Val
Gln Trp Lys Val Asp Asn Ala145 150 155 160Leu Gln Ser Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Asp Ser Lys 165 170 175Asp Ser Thr Tyr
Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp 180 185 190Tyr Glu
Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu 195 200
205Ser Ser Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 210 215
220295464PRTArtificial SequenceSynthetic 295Met Asp Val Lys Leu Val
Glu Ser Gly Gly Gly Leu Val Lys Leu Gly1 5 10 15Gly Ser Leu Lys Leu
Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn 20 25 30Tyr Tyr Met Ser
Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Leu 35 40 45Val Ala Ala
Ile Asn Ser Asp Gly Gly Ile Thr Tyr Tyr Leu Asp Thr 50 55 60Val Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu65 70 75
80Tyr Leu Gln Met Ser Ser Leu Lys Ser Glu Asp Thr Ala Leu Phe Tyr
85 90 95Cys Ala Arg His Arg Ser Gly Tyr Phe Ser Met Asp Tyr Trp Gly
Gln 100 105 110Gly Thr Ser Val Thr Val Ser Ser Ala Ser Thr Lys Gly
Pro Ser Val 115 120 125Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser
Gly Gly Thr Ala Ala 130 135 140Leu Gly Cys Leu Val Lys Asp Tyr Phe
Pro Glu Pro Val Thr Val Ser145 150 155 160Trp Asn Ser Gly Ala Leu
Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175Leu Gln Ser Ser
Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185 190Ser Ser
Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys 195 200
205Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Lys
210 215 220Ala Ser Gly Gly Arg His Arg Gln Pro Arg Gly Trp Glu Gln
Leu Gly225 230 235 240Gly Gly Gly Gly Ser Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly 245 250 255Asp Val Ser Phe Ser Thr Arg Gly Thr
Gln Asn Trp Thr Val Glu Arg 260 265 270Leu Leu Gln Ala His Ala Gln
Leu Glu Glu Arg Gly Tyr Val Phe Val 275 280 285Gly Tyr His Gly Thr
Phe Leu Glu Ala Ala Gln Ser Ile Val Phe Gly 290 295 300Gly Val Ala
Ala Arg Ser Gln Asp Leu Ala Ala Ile Trp Ala Gly Phe305 310 315
320Tyr Ile Ala Gly Asp Pro Ala Leu Ala Tyr Gly Tyr Ala Gln Asp Gln
325 330 335Glu Pro Asp Ala Ala Gly Arg Ile Arg Asn Gly Ala Leu Leu
Arg Val 340 345 350Tyr Val Pro Ala Ser Ser Leu Pro Gly Phe Tyr Arg
Thr Ser Leu Thr 355 360 365Leu Ala Ala Pro Glu Ala Ala Gly Glu Val
Glu Arg Leu Ile Gly His 370 375 380Pro Leu Pro Leu Ala Leu Asp Ala
Ile Thr Gly Pro Glu Glu Glu Gly385 390 395 400Gly Arg Leu Glu Thr
Ile Leu Gly Trp Pro Leu Ala Glu Arg Thr Val 405 410 415Val Ile Pro
Ser Ala Ile Pro Thr Asp Pro Arg Asn Val Gly Gly Asp 420 425 430Leu
Asp Pro Ser Ser Ile Pro Asp Lys Glu Gln Ala Ile Ser Ala Leu 435 440
445Pro Asp Tyr Ala Ser Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys
450 455 460296464PRTArtificial SequenceSynthetic 296Met Asp Val Lys
Leu Val Glu Ser Gly Gly Gly Leu Val Lys Leu Gly1 5 10 15Gly Ser Leu
Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn 20 25 30Tyr Tyr
Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Leu 35 40 45Val
Ala Ala Ile Asn Ser Asp Gly Gly Ile Thr Tyr Tyr Leu Asp Thr 50 55
60Val Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu65
70 75 80Tyr Leu Gln Met Ser Ser Leu Lys Ser Glu Asp Thr Ala Leu Phe
Tyr 85 90 95Cys Ala Arg His Arg Ser Gly Tyr Phe Ser Met Asp Tyr Trp
Gly Gln 100 105 110Gly Thr Ser Val Thr Val Ser Ser Ala Ser Thr Lys
Gly Pro Ser Val 115 120 125Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr
Ser Gly Gly Thr Ala Ala 130 135 140Leu Gly Cys Leu Val Lys Asp Tyr
Phe Pro Glu Pro Val Thr Val Ser145 150 155 160Trp Asn Ser Gly Ala
Leu Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175Leu Gln Ser
Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185 190Ser
Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys 195 200
205Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Lys
210 215 220Ala Ser Gly Gly Arg His Arg Gln Pro Arg Gly Trp Glu Gln
Leu Gly225 230 235 240Gly Gly Gly Gly Ser Pro Thr Gly Ala Glu Phe
Leu Gly Asp Gly Gly 245 250 255Asp Val Ser Phe Ser Thr Arg Gly Thr
Gln Asn Trp Thr Val Glu Arg 260 265 270Leu Leu Gln Ala His Ala Gln
Leu Glu Glu Arg Gly Tyr Val Phe Val 275 280 285Gly Tyr His Gly Thr
Ala Leu Glu Ala Ala Gln Ser Ile Val Phe Gly 290 295 300Gly Val Arg
Ala Arg Ser Gln Asp Leu Asp Ala Ile Trp Arg Gly Phe305 310 315
320Tyr Ile Ala Gly Asp Pro Ala His Ala Tyr Gly Tyr Ala Gln Asp Gln
325 330 335Glu Pro Asp Ala Arg Gly Arg Ile Ala Asn Gly Ala Leu Leu
Arg Val 340 345 350Tyr Val Pro Ala Ser Ser Leu Pro Gly Phe Tyr Arg
Thr Ser Leu Thr 355 360 365Leu Ala Ala Pro Glu Ala Ala Gly Glu Val
Glu Arg Leu Ile Gly His 370 375 380Pro Leu Pro Leu Arg Leu Asp Ala
Ile Thr Gly Pro Glu Glu Glu Gly385 390 395 400Gly Arg Glu Glu Thr
Ile Leu Gly Trp Pro Leu Ala Glu Arg Thr Val 405 410 415Val Ile Pro
Ser Ala Ile Pro Thr Asp Pro Arg Asn Val Gly Gly Asp 420 425 430Leu
Asp Pro Ser Ser Ile Pro Asp Lys Glu Gln Ala Ile Ser Ala Leu 435 440
445Pro Asp Tyr Ala Ser Gln Pro Gly Lys Pro Pro Arg Glu Asp Leu Lys
450 455 460297215PRTArtificial SequenceSynthetic 297Met Asp Ile Val
Met Thr Gln Ser Gln Arg Phe Met Ser Thr Thr Val1 5 10 15Gly Asp Arg
Val Ser Ile Thr Cys Lys Ala Ser Gln Asn Val Val Ser 20 25 30Ala Val
Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ser Pro Lys Leu Leu 35 40 45Ile
Tyr Ser Ala Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg Phe Thr 50 55
60Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Asn Met Gln65
70 75 80Ser Glu Asp Leu Ala Asp Phe Phe Cys Gln Gln Tyr Ser Asn Tyr
Pro 85 90 95Trp Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg Thr
Val Ala 100 105 110Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu
Gln Leu Lys Ser 115 120 125Gly Thr Ala Ser Val Val Cys Leu Leu Asn
Asn Phe Tyr Pro Arg Glu 130 135 140Ala Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln Ser Gly Asn Ser145 150 155 160Gln Glu Ser Val Thr
Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu 165 170 175Ser Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val 180 185 190Tyr
Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys 195 200
205Ser Phe Asn Arg Gly Glu Cys 210 215
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025
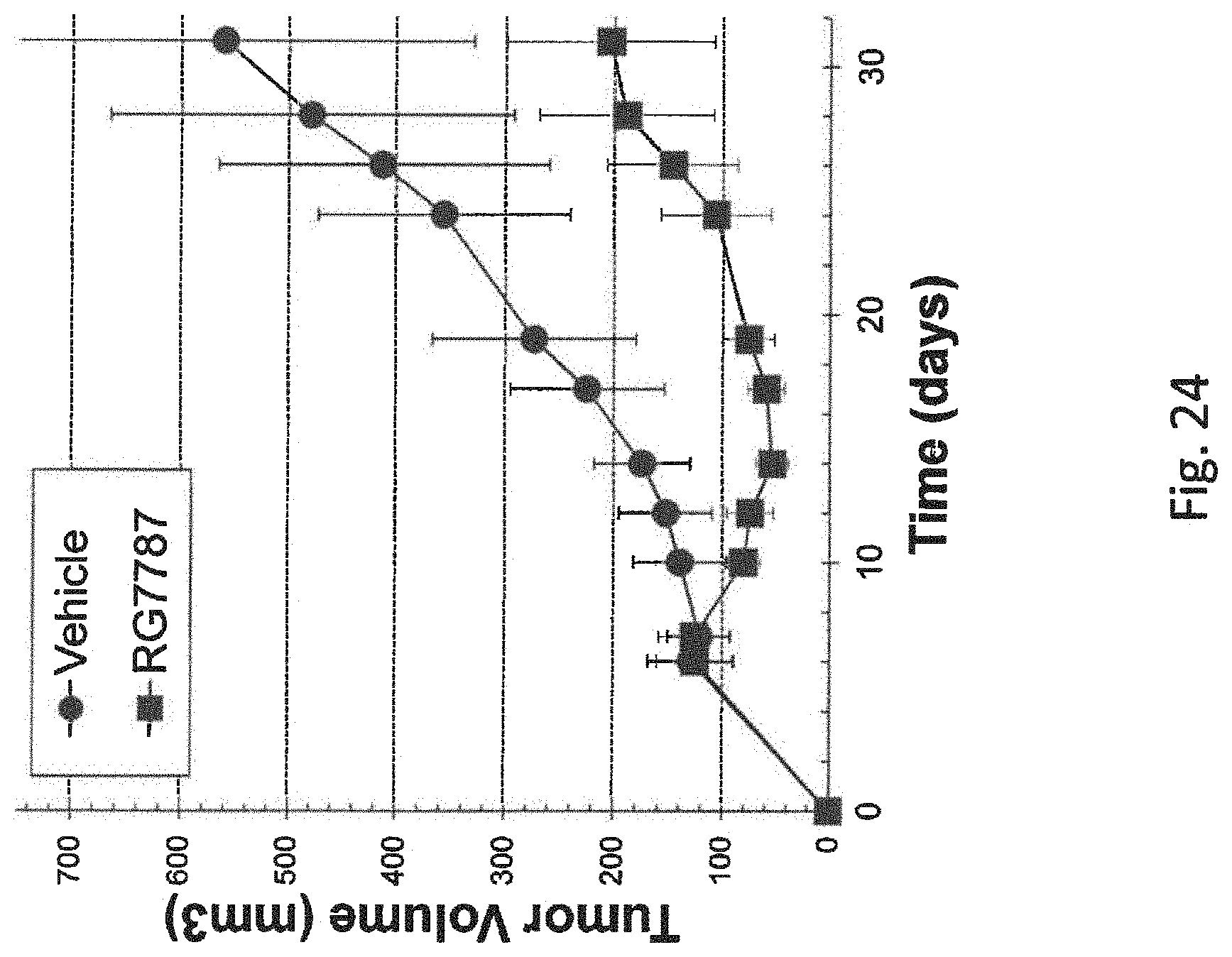
D00026

D00027

D00028

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.