Methods And Compositions For Improving Plant Traits
Temme; Karsten ; et al.
U.S. patent application number 16/159542 was filed with the patent office on 2019-02-07 for methods and compositions for improving plant traits. The applicant listed for this patent is Pivot Bio, Inc.. Invention is credited to Sarah Bloch, Rosemary Clark, Austin Davis-Richardson, Kevin Hammill, Douglas Higgins, Alvin Tamsir, Karsten Temme, Emily Tung.
| Application Number | 20190039964 16/159542 |
| Document ID | / |
| Family ID | 62839550 |
| Filed Date | 2019-02-07 |







View All Diagrams
| United States Patent Application | 20190039964 |
| Kind Code | A1 |
| Temme; Karsten ; et al. | February 7, 2019 |
METHODS AND COMPOSITIONS FOR IMPROVING PLANT TRAITS
Abstract
Methods and systems are provided for generating and utilizing a bacterial composition that comprises at least one genetically engineered bacterial strain that fixes atmospheric nitrogen in an agricultural system that has been fertilized with more than 20 lbs of Nitrogen per acre.
| Inventors: | Temme; Karsten; (Oakland, CA) ; Tamsir; Alvin; (San Francisco, CA) ; Bloch; Sarah; (Emeryville, CA) ; Clark; Rosemary; (El Cerrito, CA) ; Tung; Emily; (Milbrae, CA) ; Hammill; Kevin; (Danville, CA) ; Higgins; Douglas; (Berkeley, CA) ; Davis-Richardson; Austin; (San Francisco, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 62839550 | ||||||||||
| Appl. No.: | 16/159542 | ||||||||||
| Filed: | October 12, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| PCT/US2018/013671 | Jan 12, 2018 | |||
| 16159542 | ||||
| 62577147 | Oct 25, 2017 | |||
| 62566199 | Sep 29, 2017 | |||
| 62467032 | Mar 3, 2017 | |||
| 62447889 | Jan 18, 2017 | |||
| 62445557 | Jan 12, 2017 | |||
| 62445570 | Jan 12, 2017 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 15/70 20130101; C12N 15/743 20130101; C12R 1/22 20130101; C05F 11/08 20130101; C12R 1/07 20130101; C12N 15/52 20130101; A01N 63/10 20200101; C12N 15/111 20130101; A01H 6/4684 20180501; A01N 63/00 20130101; C12N 1/20 20130101; A01H 3/00 20130101; C12N 1/04 20130101; A01H 5/06 20130101; C12R 1/065 20130101; C12R 1/01 20130101; C12R 1/025 20130101 |
| International Class: | C05F 11/08 20060101 C05F011/08; A01H 6/46 20060101 A01H006/46; A01H 5/06 20060101 A01H005/06; C12R 1/22 20060101 C12R001/22; C12R 1/025 20060101 C12R001/025; C12R 1/065 20060101 C12R001/065; C12R 1/07 20060101 C12R001/07; C12N 1/04 20060101 C12N001/04; C12N 15/74 20060101 C12N015/74; C12N 15/11 20060101 C12N015/11; C12N 15/52 20060101 C12N015/52 |
Goverment Interests
STATEMENT AS TO FEDERALLY SPONSORED RESEARCH
[0002] This invention was made with the support of the United States government under SBIR grant 1520545 awarded by the National Science Foundation. The government has certain rights in the disclosed subject matter.
Claims
1. A method of increasing nitrogen fixation in a non-leguminous plant, comprising: applying to the plant a plurality of non-intergeneric bacteria, said plurality comprising non-intergeneric bacteria that: i. have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and ii. produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and wherein the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant, and wherein each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen.
2. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue.
3. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise bacteria that: produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour.
4. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour.
5. The method according to claim 1, wherein the at least one genetic variation comprises an introduced control sequence operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network.
6. The method according to claim 1, wherein the at least one genetic variation comprises a promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network.
7. The method according to claim 1, wherein the at least one genetic variation comprises an inducible promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network.
8. The method according to claim 1, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene of the nitrogen fixation or assimilation genetic regulatory network.
9. The method according to claim 1, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene in the nif gene cluster.
10. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation.
11. The method according to claim 1, wherein the plurality of non-intergeneric bacteria applied to the plant do not stimulate an increase in the uptake of exogenous non-atmospheric nitrogen.
12. The method according to claim 1, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight.
13. The method according to claim 1, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight, and wherein the nitrogen-containing fertilizer comprises ammonium or an ammonium containing molecule.
14. The method according to claim 1, wherein the exogenous nitrogen is selected from fertilizer comprising one or more of: glutamine, ammonia, ammonium, urea, nitrate, nitrite, ammonium-containing molecules, nitrate-containing molecules, and nitrite-containing molecules.
15. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant.
16. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant.
17. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, fix atmospheric nitrogen in non-nitrogen-limiting conditions.
18. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation.
19. The method according to claim 1, wherein the fixed nitrogen produced by the plurality of non-intergeneric bacteria is measured through dilution of enriched fertilizer by atmospheric N.sub.2 gas in plant tissue.
20. The method according to claim 1, wherein the at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network are selected from the group consisting of: nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme.
21. The method according to claim 1, wherein the at least one genetic variation is a mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD.
22. The method according to claim 1, wherein the at least one genetic variation is selected from: (A) a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion.
23. The method according to claim 1, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene.
24. The method according to claim 1, wherein the at least one genetic variation is a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain.
25. The method according to claim 1, wherein the at least one genetic variation is a mutated amtB gene that results in the lack of expression of said amtB gene.
26. The method according to claim 1, wherein the at least one genetic variation is selected from: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof.
27. The method according to claim 1, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain.
28. The method according to claim 1, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated amtB gene that results in the lack of expression of said amtB gene.
29. The method according to claim 1, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome.
30. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are formulated into a composition.
31. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are formulated into a composition comprising an agriculturally acceptable carrier.
32. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are applied into furrows in which seeds of said plant are planted.
33. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are formulated into a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter.
34. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are applied onto a seed of said plant.
35. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating and are applied onto a seed of said plant.
36. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating and are applied onto a seed of said plant, at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed.
37. The method according to claim 1, wherein the plant is a cereal crop.
38. The method according to claim 1, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale.
39. The method according to claim 1, wherein the plant is corn.
40. The method according to claim 1, wherein the plant is an agricultural crop plant.
41. The method according to claim 1, wherein the plant is a genetically modified organism.
42. The method according to claim 1, wherein the plant is not a genetically modified organism.
43. The method according to claim 1, wherein the plant has been genetically engineered or bred for efficient nitrogen use.
44. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise at least two different species of bacteria.
45. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise at least two different strains of the same species of bacteria.
46. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof.
47. The method according to claim 1, wherein the plurality of non-intergeneric bacteria are endophytic, epiphytic, or rhizospheric.
48. The method according to claim 1, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: a bacteria deposited as PTA-122293, a bacteria deposited as PTA-122294, a bacteria deposited as NCMA 201701002, a bacteria deposited as NCMA 201708004, a bacteria deposited as NCMA 201708003, a bacteria deposited as NCMA 201708002, a bacteria deposited as NCMA 201712001, a bacteria deposited as NCMA 201712002, and combinations thereof.
49. The method according to claim 1, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen.
50. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant.
51. The method accordingly to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant.
52. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant.
53. The method according to claim 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant.
54. The method according to claim 1, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per gram of fresh weight of plant root tissue per hour.
55. The method according to claim 1, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-6 mmol N per gram of fresh weight of plant root tissue per hour.
Description
CROSS-REFERENCE
[0001] This application claims the benefit of U.S. Provisional Application No. 62/445,570, filed Jan. 12, 2017; U.S. Provisional Application No. 62/445,557, filed Jan. 12, 2017; U.S. Provisional Application No. 62/447,889, filed Jan. 18, 2017; U.S. Provisional Application No. 62/467,032, filed Mar. 3, 2017; U.S. Provisional Application No. 62/566,199, filed Sep. 29, 2017; and U.S. Provisional Application No. 62/577,147, filed Oct. 25, 2017, which applications are incorporated herein by reference.
STATEMENT REGARDING SEQUENCE LISTING
[0003] The instant application contains a Sequence Listing which has been submitted electronically in ASCII format and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Jan. 3, 2018, is named 47736-707_601_SL.txt and is .apprxeq.599 kb in size.
BACKGROUND OF THE INVENTION
[0004] Plants are linked to the microbiome via a shared metabolome. A multidimensional relationship between a particular crop trait and the underlying metabolome is characterized by a landscape with numerous local maxima. Optimizing from an inferior local maximum to another representing a better trait by altering the influence of the microbiome on the metabolome may be desirable for a variety of reasons, such as for crop optimization. Economically-, environmentally-, and socially-sustainable approaches to agriculture and food production are required to meet the needs of a growing global population. By 2050 the United Nations' Food and Agriculture Organization projects that total food production must increase by 70% to meet the needs of the growing population, a challenge that is exacerbated by numerous factors, including diminishing freshwater resources, increasing competition for arable land, rising energy prices, increasing input costs, and the likely need for crops to adapt to the pressures of a drier, hotter, and more extreme global climate.
[0005] One area of interest is in the improvement of nitrogen fixation. Nitrogen gas (N.sub.2) is a major component of the atmosphere of Earth. In addition, elemental nitrogen (N) is an important component of many chemical compounds which make up living organisms. However, many organisms cannot use N.sub.2 directly to synthesize the chemicals used in physiological processes, such as growth and reproduction. In order to utilize the N.sub.2, the N.sub.2 must be combined with hydrogen. The combining of hydrogen with N.sub.2 is referred to as nitrogen fixation. Nitrogen fixation, whether accomplished chemically or biologically, requires an investment of large amounts of energy. In biological systems, an enzyme known as nitrogenase catalyzes the reaction which results in nitrogen fixation. An important goal of nitrogen fixation research is the extension of this phenotype to non-leguminous plants, particularly to important agronomic grasses such as wheat, rice, and maize. Despite enormous progress in understanding the development of the nitrogen-fixing symbiosis between rhizobia and legumes, the path to use that knowledge to induce nitrogen-fixing nodules on non-leguminous crops is still not clear. Meanwhile, the challenge of providing sufficient supplemental sources of nitrogen, such as in fertilizer, will continue to increase with the growing need for increased food production.
SUMMARY OF THE INVENTION
[0006] An aspect of the invention provides a bacterial composition that comprises at least one genetically engineered bacterial strain that fixes atmospheric nitrogen in an agricultural system that has been fertilized with more than 20 lbs of Nitrogen per acre.
[0007] Another aspect of the invention provides a bacterial composition that comprises at least one bacterial strain that has been bred to fix atmospheric nitrogen in an agricultural system that has been fertilized with more than 20 lbs of Nitrogen per acre.
[0008] An additional aspect of the invention provides a bacterial composition that comprises at least one genetically engineered bacterial strain that fixes atmospheric nitrogen, the at least one genetically engineered bacterial strain comprising exogenously added DNA wherein said exogenously added DNA shares at least 80% identity to a corresponding native bacterial strain.
[0009] A further aspect of the invention provides a seed composition comprising a seed of a plant that is inoculated with a bacterial composition.
[0010] Another aspect of the invention provides a method of growing a crop using a plurality of seeds having a seed composition that is inoculated with a bacterial composition.
[0011] An additional aspect of the invention provides a method of applying a bacterial composition to a field.
[0012] A further aspect of the invention provides a fertilizer composition comprising a bacterial composition.
[0013] Another aspect of the invention provides a method of maintaining soil nitrogen levels. The method comprises planting, in soil of a field, a crop inoculated by a genetically engineered bacterium that fixes atmospheric nitrogen. The method also comprises harvesting said crop, wherein no more than 90% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting.
[0014] An additional aspect of the invention provides a method of delivering a probiotic supplement to a crop plant. The method comprises coating a crop seed with a seed coating, seed treatment, or seed dressing. Said seed coating, seed dressing, or seed treatment comprises living representatives of said probiotic. Additionally, the method comprises applying, in soil of a field, said crop seeds.
[0015] In a further aspect of the invention, the genetically engineered bacterial strain is a genetically engineered Gram-positive bacterial strain. In some cases, the genetically engineered Gram-positive bacterial strain has an altered expression level of a regulator of a Nif cluster. In some cases, the genetically engineered Gram-positive bacterial strain expresses a decreased amount of a negative regulator of a Nif cluster. In some cases, the genetically engineered bacterial strain expresses a decreased amount of GlnR. In some cases, the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 75% identity to a sequence from the Zehr lab NifH database. In some cases, the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 85% identity to a sequence from the Zehr lab NifH database. In some cases, the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 75% identity to a sequence from the Buckley lab NifH database. In some cases, the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 85% identity to a sequence from the Buckley lab NifH database.
[0016] Another aspect of the invention provides a method of increasing nitrogen fixation in a non-leguminous plant. The method comprises applying to the plant a plurality of non-intergeneric bacteria, said plurality comprising non-intergeneric bacteria that (i) have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and (ii) produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. Additionally, the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant. Further, each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen.
[0017] An additional aspect of the invention provides a method of increasing nitrogen fixation in a non-leguminous plant. The method comprises applying to the plant a plurality of non-intergeneric bacteria, said plurality comprising non-intergeneric bacteria that (i) have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and/or (ii) produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. Additionally, the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant. Further, each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen.
[0018] A further aspect of the invention provides a method of breeding microbial strains to improve specific traits of agronomic relevance. The method comprises providing a plurality of microbial strains that have the ability to colonize a desired crop. The method also comprises improving regulatory networks influencing the trait through intragenomic rearrangement. Further, the method comprises assessing microbial strains within the plurality of microbial strains to determine a measure of the trait. Additionally, the method comprises selecting one or more microbial strains of the plurality of microbial strains that generate an improvement in the trait in the presence of the desired crop.
[0019] Another aspect of the invention provides a method of breeding microbial strains to improve specific traits of agronomic relevance. The method comprises providing a plurality of microbial strains that have the ability to colonize a desired crop. The method also comprises introducing genetic diversity into the plurality of microbial strains. Additionally, the method comprises assessing microbial strains within the plurality of microbial strains to determine a measure of the trait. Further, the method comprises selecting one or more microbial strains of the plurality of microbial strains that generate an improvement in the trait in die presence of the desired crop.
[0020] Another aspect of the invention provides a method of increasing the amount of atmospheric derived nitrogen in a non-leguminous plant. The method comprises exposing said non-leguminous plant to engineered non-intergeneric microbes, said engineered non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network or at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network.
[0021] A further aspect of the invention provides a method of increasing an amount of atmospheric derived nitrogen in a corn plant. The method comprises exposing said corn plant to engineered non-intergeneric microbes comprising engineered genetic variations within at least two genes selected from the group consisting of nifL, glnB, glnE, and amtB.
[0022] Another aspect of the invention provides a method of increasing an amount of atmospheric derived nitrogen in a corn plant. The method comprises exposing said corn plant to engineered non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network, wherein said engineered non-intergeneric microbes, in planta, produce at least 5% of fixed nitrogen in said corn plant as measured by dilution of 15N in crops grown in fields treated with fertilizer containing 1.2% 15N.
[0023] An additional aspect of the invention provides a method of increasing nitrogen fixation in a non-leguminous plant. The method comprises applying to the plant a plurality of non-intergeneric bacteria, said plurality comprising non-intergeneric bacteria that (i) have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and (ii) produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. Additionally, the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.5.times.10.sup.-8 mmol N per grain of fresh weight of plant root tissue per hour. Further, the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant. Additionally, each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen.
[0024] Another aspect of the invention provides a method of increasing nitrogen fixation in a non-leguminous plant. The method comprises applying to the plant a plurality of bacteria, said plurality comprising bacteria that (i) have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and/or (ii) produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. Additionally, the plurality of bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant.
[0025] An additional aspect of the invention provides a non-intergeneric bacterial population capable of increasing nitrogen fixation in a non-leguminous plant, comprising a plurality of non-intergeneric bacteria that (i) have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and/or (ii) produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. Additionally, the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in a plant grown in the presence of the plurality of non-intergeneric bacteria. Further, each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in die presence of exogenous nitrogen.
[0026] A further aspect of the invention provides a bacterial population capable of increasing nitrogen fixation in a non-leguminous plant, the bacterial population comprising a plurality of bacteria that (i) have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or (ii) produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. Additionally, the plurality of bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant.
[0027] In another aspect of the invention, a bacterium is provided that (i) has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and/or (ii) produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour.
[0028] In a further aspect of the invention, a non-intergeneric bacterium is provided that comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacterium is capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen, and wherein said bacterium (i) has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and/or (ii) produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour.
[0029] In an additional aspect of the invention provides a method for increasing nitrogen fixation in a plant, comprising administering to the plant an effective amount of a composition that comprises a purified population of bacteria that comprises bacteria with a 16S nucleic acid sequence that is at least about 97% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, 277-283; a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, 284-295; and/or a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260, 296-303; and wherein the plant administered the effective amount of the composition exhibits an increase in nitrogen fixation, as compared to a plant not having been administered the composition.
[0030] A further aspect of the invention provides an isolated bacteria comprising a 16S nucleic acid sequence that is at least about 97% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, 277-283; a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, 284-295; and/or a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260, 296-303.
[0031] Another aspect of the invention provides a method of detecting a non-native junction sequence, comprising: amplifying a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 372-405.
[0032] An additional aspect of the invention provides a method of detecting a non-native junction sequence, comprising: amplifying a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to at least a 10 contiguous base pair fragment contained in SEQ ID NOs: 372-405, said contiguous base pair fragment being comprised of nucleotides at the intersection of: an upstream sequence comprising SEQ ID NOs: 304-337 and downstream sequence comprising SEQ ID NOs: 338-371.
[0033] A further aspect of the invention provides a non-native junction sequence comprising a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 884/0, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 372-405.
[0034] An additional aspect of the invention provides a non-native junction sequence comprising a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to at least a 10 contiguous base pair fragment contained in SEQ ID NOs: 372-405, said contiguous base pair fragment being comprised of nucleotides at the intersection of: an upstream sequence comprising SEQ ID NOs: 304-337 and downstream sequence comprising SEQ ED NOs: 338-371.
[0035] A further aspect of the invention provides a bacterial composition comprising at least one remodeled bacterial strain that fixes atmospheric nitrogen, the at least one remodeled bacterial strain comprising exogenously added DNA wherein said exogenously added DNA shares at least 80% identity to a corresponding native bacterial strain.
[0036] An additional aspect of the invention provides a method of maintaining soil nitrogen levels. The method comprises planting, in soil of a field, a crop inoculated by a remodeled bacterium that fixes atmospheric nitrogen. The method also comprises harvesting said crop, wherein no more than 90% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting.
[0037] Another aspect of the invention provides a method of delivering a probiotic supplement to a crop plant. The method comprises coating a crop seed with a seed coating, seed treatment, or seed dressing, wherein said seed coating, seed dressing, or seed treatment comprise living representatives of said probiotic. The method also comprises applying said crop seeds in soil of a field.
[0038] An additional aspect of the invention provides a method of increasing the amount of atmospheric derived nitrogen in a non-leguminous plant. The method comprises exposing said non-leguminous plant to remodeled non-intergeneric microbes, said remodeled non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network or at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network.
[0039] A further aspect of the invention provides a method of increasing an amount of atmospheric derived nitrogen in a corn plant. The method comprises exposing said corn plant to remodeled non-intergeneric microbes comprising remodeled genetic variations within at least two genes selected from the group consisting of nifL, glnB, glnE, and amtB.
[0040] Another aspect of the invention provides a method of increasing an amount of atmospheric derived nitrogen in a corn plant. The method comprises exposing said corn plant to remodeled non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network, wherein said remodeled non-intergeneric microbes, in planta, produce at least 5% of fixed nitrogen in said corn plant as measured by dilution of 15N in crops grown in fields treated with fertilizer containing 1.2% 15N.
[0041] Additional aspects of the invention provide genus of microbes that are evolved and optimized for in planta nitrogen fixation in non-leguminous crops. In particular, methods of increasing nitrogen fixation in a non-leguminous plant are disclosed. The methods can comprise exposing the plant to a plurality of bacteria. Each member of the plurality comprises one or more genetic variations introduced into one or more genes of non-coding polynucleotides of the bacteria's nitrogen fixation or assimilation genetic regulatory network, such that the bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen. The bacteria are not intergeneric microorganisms. Additionally, the bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant.
[0042] Further aspects of the invention provide beneficial isolated microbes and microbial compositions. In particular, isolated and biologically pure microorganisms that have applications, inter alia, in increasing nitrogen fixation in a crop are provided. The disclosed microorganism can be utilized in their isolated and biologically pure states, as well as being formulated into compositions. Furthermore, the disclosure provides microbial compositions containing at least two members of the disclosed microorganisms, as well as methods of utilizing said microbial compositions.
INCORPORATION BY REFERENCE
[0043] All publications, patents, and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication, patent, or patent application was specifically and individually indicated to be incorporated by reference.
BRIEF DESCRIPTION OF THE DRAWINGS
[0044] The novel features of the invention are set forth with particularity in the appended claims. A better understanding of the features and advantages of the present invention will be obtained by reference to the following detailed description that sets forth illustrative embodiments, in which the principles of the invention are utilized, and the accompanying drawings of which:
[0045] FIGS. 1A-B depict enrichment and isolation of nitrogen fixing bacteria. (A) Nfb agar plate was used to isolate single colonies of nitrogen fixing bacteria. (B) Semi-solid Nfb agar casted in Balch tube. The arrow points to pellicle of enriched nitrogen fixing bacteria.
[0046] FIG. 2 depicts a representative nifH PCR screen. Positive bands were observed at .about.350 bp for two colonies in this screen. Lower bands represent primer-dimers.
[0047] FIG. 3 depicts an example of a PCR screen of colonies from CRISPR-Cas-selected mutagenesis. CI006 colonies were screened with primers specific for the nifL locus. The wild type PCR product is expected at .about.2.2 kb, whereas the mutant is expected at .about.1.1 kb. Seven of ten colonies screened unambiguously show the desired deletion.
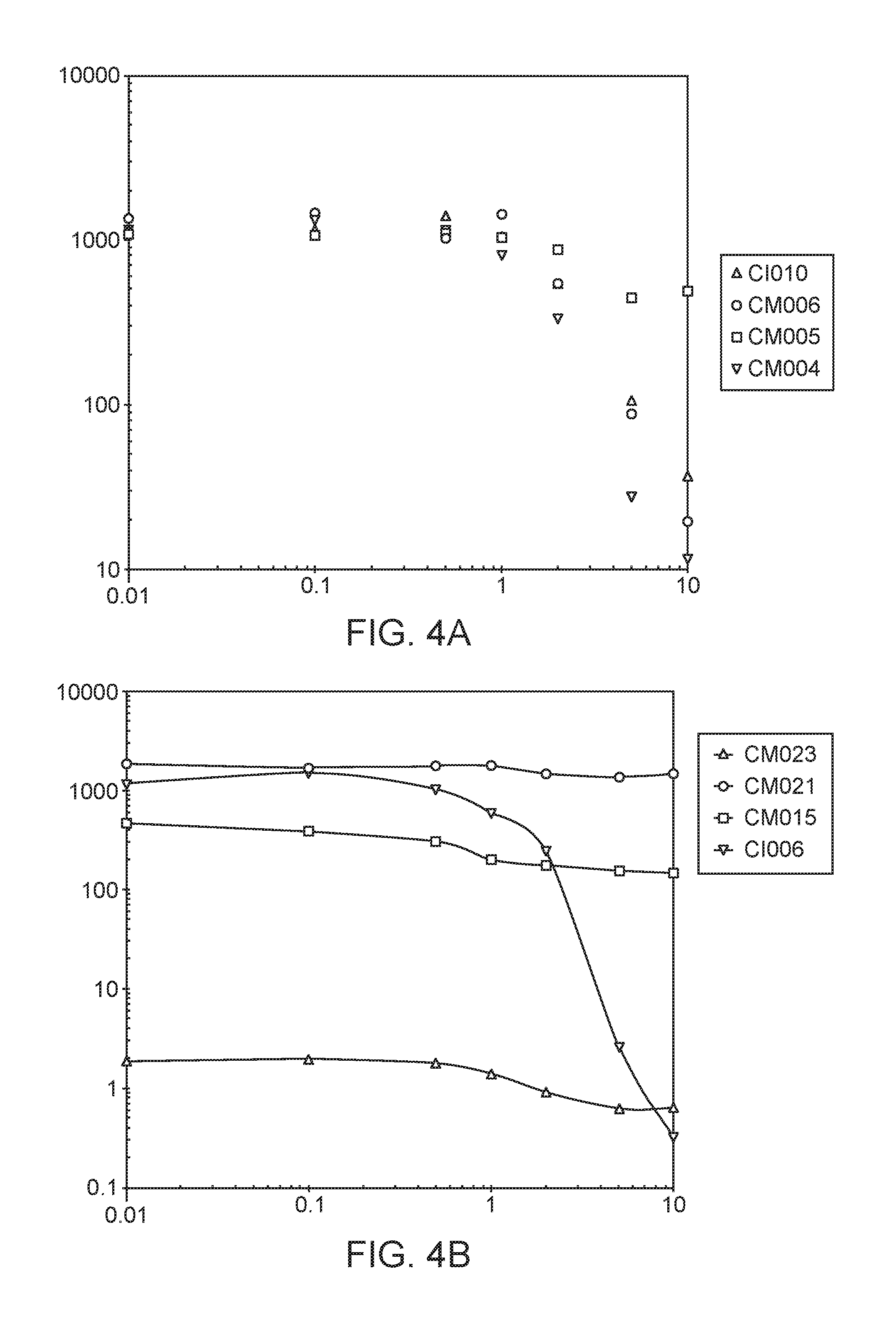
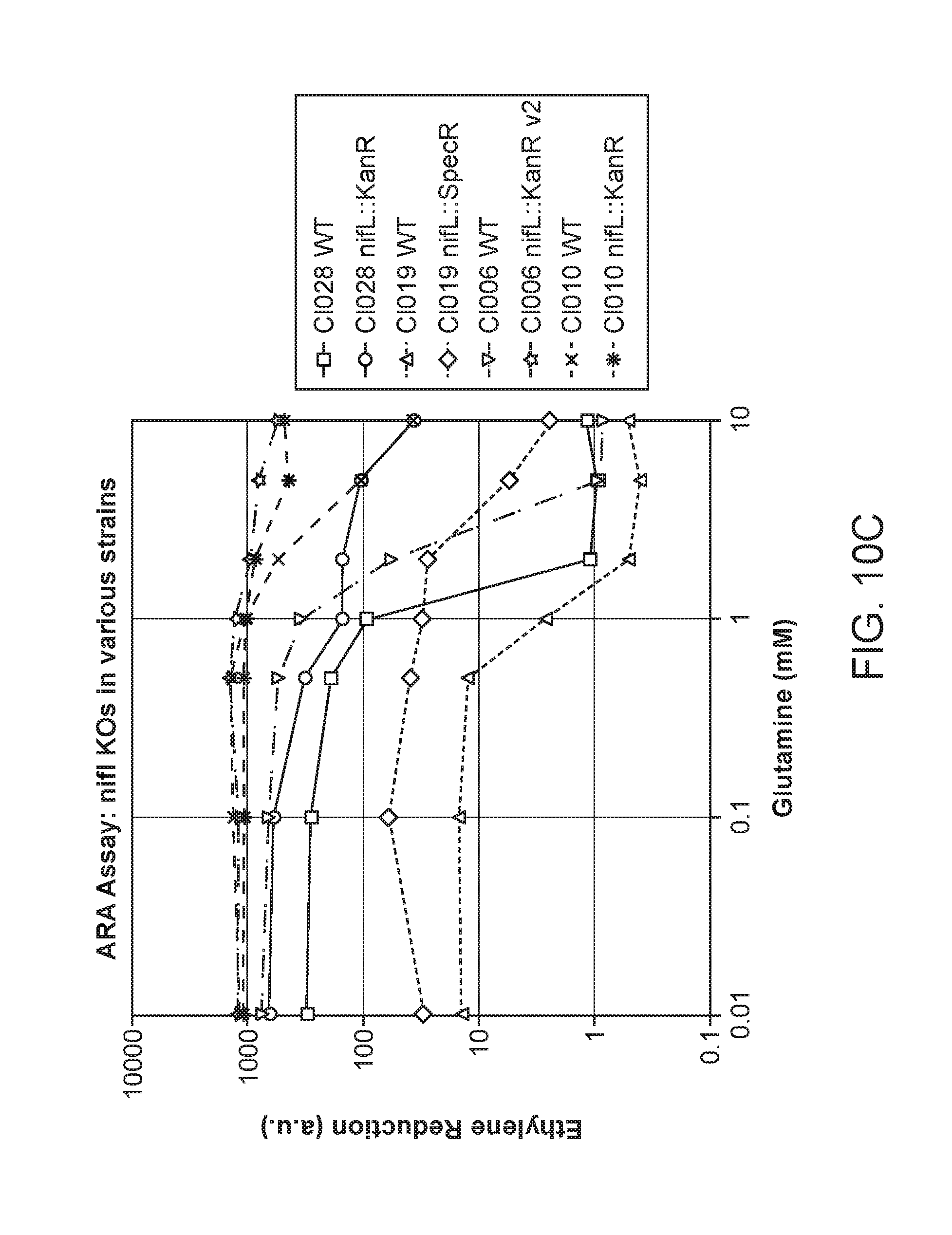
[0048] FIGS. 4A-D depict in vitro phenotypes of various strains. The Acetylene Reduction Assay (ARA) activities of mutants of strain CI010 (FIG. 4A) and mutants of strain CI006 (FIG. 4B) grown in nitrogen fixation media supplemented with 0 to 10 mM glutamine. ARA activities of additional strains are shown in FIG. 4C, and the ammonium excretion profile across time of two strains is shown in FIG. 4D.
[0049] FIG. 5 depicts in culture expression profile of 9 different genes in strains CI006 involved in diazaotrophic nitrogen fixation. Numbers represent counts of each transcript. Various conditions (0, 1, 10 mM Glutamine and 0%, 10%, 20% atmospheric air in N2) are indicated.
[0050] FIG. 6 depicts CI006 colonization of corn roots. Corn seedlings were inoculated with CI006 harboring an RFP expression plasmid. After two weeks of growth and plasmid maintenance through watering with the appropriate antibiotic, roots were harvested and imaged through fluorescence microscopy. Colonization of the root intercellular space is observed.
[0051] FIG. 7 depicts nitrogen derived from microbe level in WT (CI050) and optimized (CM002) strain.
[0052] FIG. 8 shows an experimental setup for a Micro-Tom fruiting mass assay.
[0053] FIG. 9 shows a screen of 10 strains for increase in Micro-Tom plant fruit mass. Results for six replicates are presented. For column 3, p=0.07. For column 7, p=0.05.
[0054] FIGS. 10A-C depict additional results for ARA activities of candidate microbes and counterpart candidate mutants grown in nitrogen fixation media supplemented with 0 to 10 mM glutamine.
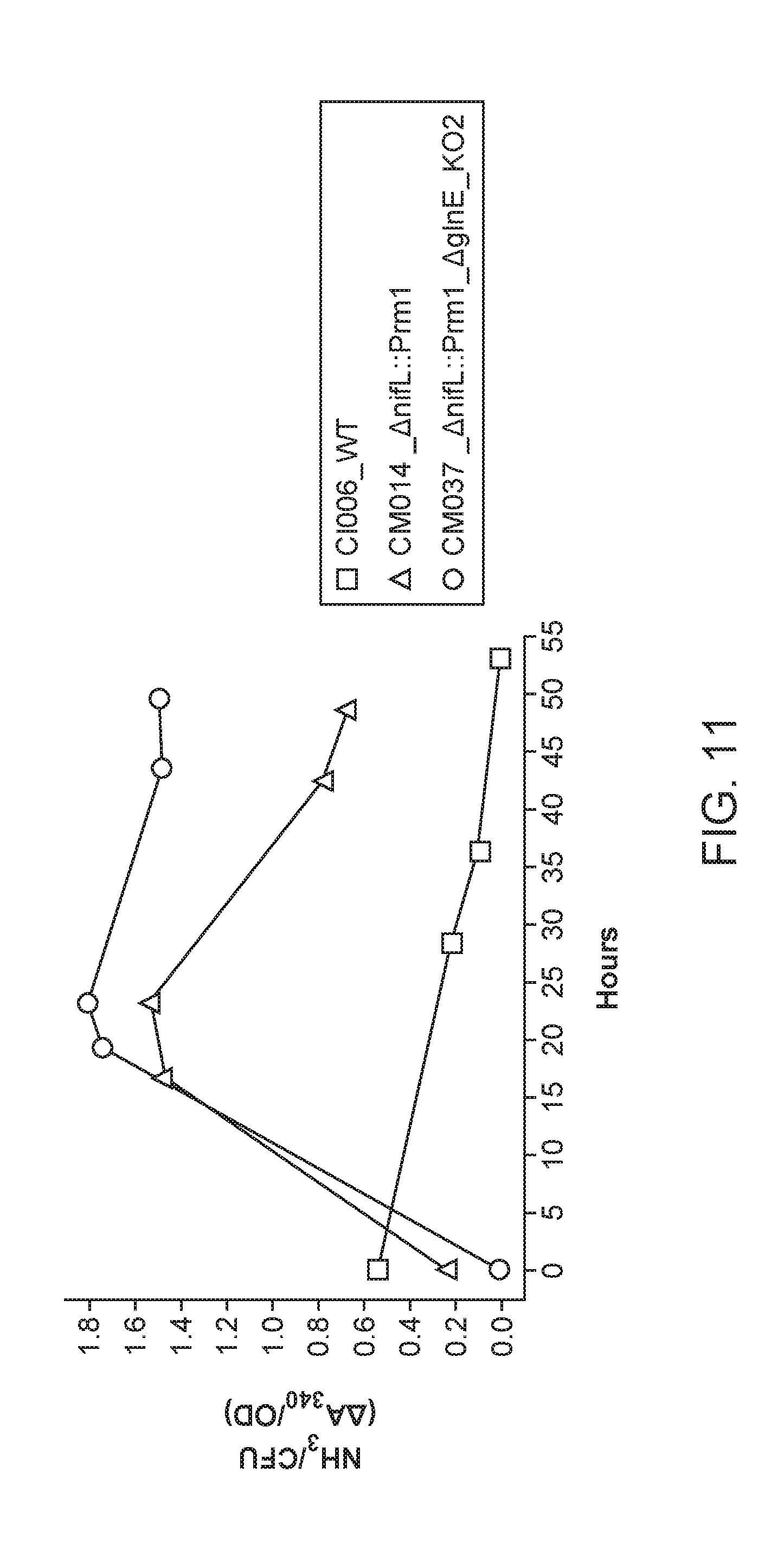
[0055] FIG. 11 depicts a double mutant that exhibits higher ammonia excretion than the single mutant from which it was derived.
[0056] FIG. 12 depicts NDFA obtained from 15N Gas Uptake experiment (extrapolated back using days exposed) to measure NDFA in Corn plants in fertilized condition.
[0057] FIG. 13 depicts NDFA value obtained from 15N Gas Uptake experiment (extrapolated back using days exposed) to measure NDFA in Setaria plants in fertilized condition.
[0058] FIG. 14A depicts rate of incorporation of 15N gas. Plants inoculated with evolved strain showed increase in 151 gas incorporation compared to uninoculated plants.
[0059] FIG. 14B depicts 4 weeks after planting, up to 7% of the nitrogen in plants inoculated with an evolved strain is derived from microbially fixed nitrogen.
[0060] FIG. 14C depicts leaf area (and other biomass measurement, data not shown) is increased in plants inoculated with an evolved strain when compared to uninoculated or wild type inoculated plants.
[0061] FIG. 15A depicts evolved strains that show significantly higher nifH production in the root tissue, as measured by in planta transcriptomic study.
[0062] FIG. 15B depicts that rate of fixed nitrogen found in plant tissue is correlated with the rate in which that particular plant is colonized by HoME optimized strain.
[0063] FIG. 16A depicts a soil texture map of various field soils tested for colonization. Soils in which a few microbes were originally source from are indicated as stars.
[0064] FIG. 16B depicts the colonization rate of Strain 1 and Strain 5 that are tested across four different soil types (circles). Both strains showed relatively robust colonization profile across diverse soil types.
[0065] FIG. 16C depicts colonization of Strain 1 as tested in a field trial over the span of a growing season. Strain 1 persists in the corn tissue up to week 12 after planting and starts to show decline in colonization after that time.
[0066] FIG. 17A depicts a schematic of microbe breeding, in accordance with embodiments.
[0067] FIG. 17B depicts an expanded view of the measurement of microbiome composition as shown in FIG. 17A.
[0068] FIG. 17C depicts sampling of roots utilized in Example 7.
[0069] FIG. 18 depicts the lineage of modified strains that were derived from strain CI006.
[0070] FIG. 19 depicts the lineage of modified strains that were derived from strain CI019.
[0071] FIG. 20 depicts a heatmap of the pounds of nitrogen delivered per acre-season by microbes of the present disclosure recorded as a function of microbes per g-fresh weight by mmol of nitrogen/microbe-hr. Below the thin line that transects the larger image are the microbes that deliver less than one pound of nitrogen per acre-season, and above the line are the microbes that deliver greater than one pound of nitrogen per acre-season. The table below the heatmap gives the precise value of mmol N produced per microbe per hour (mmol N/Microbe hr) along with the precise CFU per gram of fresh weight (CFU/g fw) for each microbe shown in the heatmap. The microbes utilized in the heatmap were assayed for N production in corn. For the WT strains CI006 and CI019, corn root colonization data was taken from a single field site. For the remaining strains, colonization was assumed to be the same as the % VT field level. N-fixation activity was determined using an in vitro ARA assay at 5 mM glutamine.
[0072] FIG. 21 depicts the plant yield of plants having been exposed to strain CI006. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0073] FIG. 22 depicts the plant yield of plants having been exposed to strain CM029. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0074] FIG. 23 depicts the plant yield of plants having been exposed to strain CM038. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0075] FIG. 24 depicts the plant yield of plants having been exposed to strain CI019. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0076] FIG. 25 depicts die plant yield of plants having been exposed to strain CM081. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0077] FIG. 26 depicts the plant yield of plants having been exposed to strains CM029 and CM081. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0078] FIG. 27 depicts the plant yield of plants as the aggregated bushel gain/loss. The area of the circles corresponds to the relative yield, while the shading corresponds to the particular MRTN treatment. The x-axis is the p value and the y-axis is the win rate.
[0079] FIGS. 28A-28E illustrate derivative microbes that fix and excrete nitrogen in vitro under conditions similar to high nitrate agricultural soils. FIG. 28A illustrates the regulatory network controlling nitrogen fixation and assimilation in PBC6.1 is shown, including the key nodes NifL, NifA, GS, GlnE depicted as the two-domain ATase-AR enzyme, and AmtB. FIG. 28B illustrates the genome of Kosakonia sacchari isolate PBC6.1 is shown. The three tracks circumscribing the genome convey transcription data from PBC6.1, PBC6.38, and the differential expression between the strains respectively. FIG. 28C illustrates the nitrogen fixation gene cluster and transcription data is expanded for finer detail. FIG. 28D illustrates nitrogenase activity under varying concentrations of exogenous nitrogen is measured with the acetylene reduction assay. The wild type strain exhibits repression of nitrogenase activity as glutamine concentrations increase, while derivative strains show varying degrees of robustness. Error bars represent standard error of the mean of at least three biological replicates. FIG. 28E illustrates temporal excretion of ammonia by derivative strains is observed at mM concentrations. Wild type strains are not observed to excrete fixed nitrogen, and negligible ammonia accumulates in the media. Error bars represent standard error of the mean.
[0080] FIGS. 29A-29C illustrate greenhouse experiments demonstrate microbial nitrogen fixation in corn. FIG. 29A illustrates microbe colonization six weeks after inoculation of corn plants by PBC6.1 derivative strains. Error bars show standard error of the mean of at least eight biological replicates. FIG. 29B illustrates in planta transcription of nifH measured by extraction of total RNA from roots and subsequent Nanostring analysis. Only derivative strains show WIN transcription in the root environment. Error bars show standard error of the mean of at least 3 biological replicates. FIG. 29C illustrates microbial nitrogen fixation measured by the dilution of isotopic tracer in plant tissues. Derivative microbes exhibit substantial transfer of fixed nitrogen to the plant. Error bars show standard error of the mean of at least ten biological replicates.
[0081] FIG. 30 illustrates PBC6.1 colonization to nearly 21% abundance of the root-associated microbiota in corn roots. Abundance data is based on 16S amplicon sequencing of the rhizosphere and endosphere of corn plants inoculated with PBC6.1 and grown in greenhouse conditions.
[0082] FIG. 31 illustrates transcriptional rates of nifA in derivative strains of PBC6.1 correlated with acetylene reduction rates. An ARA assay was performed as described in the Methods, after which cultures were sampled and subjected to qPCR analysis to determine nifA transcript levels. Error bars show standard error of the mean of at least three biological replicates in each measure.
[0083] FIG. 32 illustrates results from a summer 2017 field testing experiment. The yield results obtained demonstrate that the microbes of the disclosure can serve as a potential fertilizer replacement. For instance, the utilization of a microbe of the disclosure (i.e. 6-403) resulted in a higher yield than the wild type strain (WT) and a higher yield than the untreated control (UTC). The "-25 lbs N" treatment utilizes 25 lbs less N per acre than standard agricultural practices of the region. The "100% N" UTC treatment is meant to depict standard agricultural practices of the region, in which 100% of the standard utilization of N is deployed by the farmer. The microbe "6-403" was deposited as NCMA 201708004 and can be found in Table A. This is a mutant Kosakonia sacchari (also called CM037) and is a progeny mutant strain from CI006 WT.
[0084] FIG. 33 illustrates results from a summer 2017 field testing experiment. The yield results obtained demonstrate that the microbes of the disclosure perform consistently across locations. Furthermore, the yield results demonstrate that the microbes of the disclosure perform well in both a nitrogen stressed environment, as well as an environment that has sufficient supplies of nitrogen. The microbe "6-881" (also known as CM094, PBC6.94), and which is a progeny mutant Kosakonia sacchari strain from CI006 WT, was deposited as NCMA 201708002 and can be found in Table A. The microbe "137-1034," which is a progeny mutant Klebsiella variicola strain from CI137 WT, was deposited as NCMA 201712001 and can be found in Table A. The microbe "137-1036," which is a progeny mutant Klebsiella variicola strain from CI137 WT, was deposited as NCMA 201712002 and can be found in Table A. The microbe "6-404" (also known as CM38, PBC6.38), and which is a progeny mutant Kosakonia sacchari strain from CI006 WT, was deposited as NCMA 201708003 and can be found in Table A. The "Nutrient Stress" condition corresponds to the 0% nitrogen regime. The "Sufficient Fertilizer" condition corresponds to the 100% nitrogen regime.
[0085] FIG. 34 depicts the lineage of modified strains that were derived from strain CI006 (also termed "6", Kosakonia sacchari WT).
[0086] FIG. 35 depicts the lineage of modified strains that were derived from strain CI019 (also termed "19", Rahnella aquatilis WT).
[0087] FIG. 36 depicts the lineage of modified strains that were derived from strain CI137 (also termed ("137", Klebsiella variicola WT).
[0088] FIG. 37 depicts the lineage of modified strains that were derived from strain 1021 (Kosakonia pseudosacchari WT).
[0089] FIG. 38 depicts the lineage of modified strains that were derived from strain 910 (Kluyvera intermedia WT).
[0090] FIG. 39 depicts the lineage of modified strains that were derived from strain 63 (Rahnella aquatilis WT).
[0091] FIG. 40 depicts a heatmap of the pounds of nitrogen delivered per acre-season by microbes of the present disclosure recorded as a function of microbes per g-fresh weight by mmol of nitrogen/microbe-hr. Below the thin line that transects the larger image are the microbes that deliver less than one pound of nitrogen per acre-season, and above the line are the microbes that deliver greater than one pound of nitrogen per acre-season. The Table C in Example 12 gives the precise value of mmol N produced per microbe per hour (mmol N/Microbe hr) along with the precise CFU per gram of fresh weight (CFU/g fw) for each microbe shown in the heatmap. The data in FIG. 40 is derived from microbial strains assayed for N production in corn in field conditions. Each point represents 1b N/acre produced by a microbe using corn root colonization data from a single field site. N-fixation activity was determined using in vitro ARA assay at 5 mM N in the form of glutamine or ammonium phosphate.
[0092] FIG. 41 depicts a heatmap of the pounds of nitrogen delivered per acre-season by microbes of the present disclosure recorded as a function of microbes per g-fresh weight by mmol of nitrogen/microbe-hr. Below the thin line that transects the larger image are the microbes that deliver less than one pound of nitrogen per acre-season, and above the line are the microbes that deliver greater than one pound of nitrogen per acre-season. The Table Din Example 12 gives the precise value of mmol N produced per microbe per hour (mmol N/Microbe hr) along with the precise CFU per gram of fresh weight (CFU/g fw) for each microbe shown in the heatmap. The data in FIG. 41 is derived from microbial strains assayed for N production in corn in laboratory and greenhouse conditions. Each point represents 1b N/acre produced by a single strain. White points represent strains in which corn root colonization data was gathered in greenhouse conditions. Black points represent mutant strains for which corn root colonization levels are derived from average field corn root colonization levels of the wild-type parent strain. Hatched points represent the wild type parent strains at their average field corn root colonization levels. In all cases, N-fixation activity was determined by in vitro ARA assay at 5 mM N in the form of glutamine or ammonium phosphate.
DETAILED DESCRIPTION OF THE INVENTION
[0093] While various embodiments of the invention have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions may occur to those skilled in the art without departing from the invention. It should be understood that various alternatives to the embodiments of the invention described herein may be employed.
[0094] Increased fertilizer utilization brings with it environmental concerns and is also likely not possible for many economically stressed regions of the globe. Furthermore, many industry players in the microbial arena are focused on creating intergeneric microbes. However, there is a heavy regulatory burden placed on engineered microbes that are characterized/classified as intergeneric. These intergeneric microbes face not only a higher regulatory burden, which makes widespread adoption and implementation difficult, but they also face a great deal of public perception scrutiny.
[0095] Currently, there are no engineered microbes on the market that are non-intergeneric and that are capable of increasing nitrogen fixation in non-leguminous crops. This dearth of such a microbe is a missing element in helping to usher in a truly environmentally friendly and more sustainable 21.sup.st century agricultural system.
[0096] The present disclosure solves the aforementioned problems and provides a non-intergeneric microbe that has been engineered to readily fix nitrogen in crops. These microbes are not characterized/classified as intergeneric microbes and thus will not face the steep regulatory burdens of such. Further, the taught non-intergeneric microbes will serve to help 21.sup.st century farmers become less dependent upon utilizing ever increasing amounts of exogenous nitrogen fertilizer.
Definitions
[0097] The terms "polynucleotide", "nucleotide", "nucleotide sequence", "nucleic acid" and "oligonucleotide" are used interchangeably. They refer to a polymeric form of nucleotides of any length, either deoxyribonucleotides or ribonucleotides, or analogs thereof. Polynucleotides may have any three dimensional structure, and may perform any function, known or unknown. The following are non-limiting examples of polynucleotides: coding or non-coding regions of a gene or gene fragment, loci (locus) defined from linkage analysis, exons, introns, messenger RNA (mRNA), transfer RNA (tRNA), ribosomal RNA (rRNA), short interfering RNA (siRNA), short-hairpin RNA (shRNA), micro-RNA (miRNA), ribozymes, cDNA, recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes, and primers. A polynucleotide may comprise one or more modified nucleotides, such as methylated nucleotides and nucleotide analogs. If present, modifications to the nucleotide structure may be imparted before or after assembly of the polymer. The sequence of nucleotides may be interrupted by non-nucleotide components. A polynucleotide may be further modified after polymerization, such as by conjugation with a labeling component.
[0098] "Hybridization" refers to a reaction in which one or more polynucleotides react to form a complex that is stabilized via hydrogen bonding between the bases of the nucleotide residues. The hydrogen bonding may occur by Watson Crick base pairing, Hoogstein binding, or in any other sequence specific manner according to base complementarity. The complex may comprise two strands forming a duplex structure, three or more strands forming a multi stranded complex, a single self-hybridizing strand, or any combination of these. A hybridization reaction may constitute a step in a more extensive process, such as the initiation of PCR, or the enzymatic cleavage of a polynucleotide by an endonuclease. A second sequence that is complementary to a first sequence is referred to as the "complement" of the first sequence. The term "hybridizable" as applied to a polynucleotide refers to the ability of the polynucleotide to form a complex that is stabilized via hydrogen bonding between the bases of the nucleotide residues in a hybridization reaction.
[0099] "Complementarity" refers to the ability of a nucleic acid to form hydrogen bond(s) with another nucleic acid sequence by either traditional Watson-Crick or other non-traditional types. A percent complementarity indicates the percentage of residues in a nucleic acid molecule which can form hydrogen bonds (e.g., Watson-Crick base pairing) with a second nucleic acid sequence (e.g., 5, 6, 7, 8, 9, 10 out of 10 being 50%, 60%, 70%, 80%, 90%, and 100% complementary, respectively). "Perfectly complementary" means that all the contiguous residues of a nucleic acid sequence will hydrogen bond with the same number of contiguous residues in a second nucleic acid sequence. "Substantially complementary" as used herein refers to a degree of complementarity that is at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99%, or 100% over a region of 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 30, 35, 40, 45, 50, or more nucleotides, or refers to two nucleic acids that hybridize under stringent conditions. Sequence identity, such as for the purpose of assessing percent complementarity, may be measured by any suitable alignment algorithm, including but not limited to the Needleman-Wunsch algorithm (see e.g. the EMBOSS Needle aligner available at www.ebi.ac.uk/Tools/psa/emboss_needle/nucleotide.html, optionally with default settings), the BLAST algorithm (see e.g. the BLAST alignment tool available at blast.ncbi.nlm.nih.gov/Blast.cgi, optionally with default settings), or the Smith-Waterman algorithm (see e.g. the EMBOSS Water aligner available at www.ebi.ac.uk/Tools/psa/emboss_water/nucleotide.html, optionally with default settings). Optimal alignment may be assessed using any suitable parameters of a chosen algorithm, including default parameters.
[0100] In general, "stringent conditions" for hybridization refer to conditions under which a nucleic acid having complementarity to a target sequence predominantly hybridizes with a target sequence, and substantially does not hybridize to non-target sequences. Stringent conditions are generally sequence-dependent and vary depending on a number of factors. In general, the longer the sequence, the higher the temperature at which the sequence specifically hybridizes to its target sequence. Non-limiting examples of stringent conditions are described in detail in Tijssen (1993), Laboratory Techniques In Biochemistry And Molecular Biology-Hybridization With Nucleic Acid Probes Part I, Second Chapter "Overview of principles of hybridization and the strategy of nucleic acid probe assay", Elsevier, N.Y.
[0101] As used herein, "expression" refers to the process by which a polynucleotide is transcribed from a DNA template (such as into and mRNA or other RNA transcript) and/or the process by which a transcribed mRNA is subsequently translated into peptides, polypeptides, or proteins. Transcripts and encoded polypeptides may be collectively referred to as "gene product." If the polynucleotide is derived from genomic DNA, expression may include splicing of the mRNA in a eukaryotic cell.
[0102] The terms "polypeptide", "peptide" and "protein" are used interchangeably herein to refer to polymers of amino acids of any length. The polymer may be linear or branched, it may comprise modified amino acids, and it may be interrupted by non-amino acids. The terms also encompass an amino acid polymer that has been modified; for example, disulfide bond formation, glycosylation, lipidation, acetylation, phosphorylation, or any other manipulation, such as conjugation with a labeling component. As used herein the term "amino acid" includes natural and/or unnatural or synthetic amino acids, including glycine and both the D or L optical isomers, and amino acid analogs and peptidomimetics.
[0103] As used herein, the term "about" is used synonymously with the term "approximately." Illustratively, the use of the term "about" with regard to an amount indicates that values slightly outside the cited values, e.g., plus or minus 0.1% to 10%.
[0104] The term "biologically pure culture" or "substantially pure culture" refers to a culture of a bacterial species described herein containing no other bacterial species in quantities sufficient to interfere with the replication of the culture or be detected by normal bacteriological techniques.
[0105] "Plant productivity" refers generally to any aspect of growth or development of a plant that is a reason for which the plant is grown. For food crops, such as grains or vegetables, "plant productivity" can refer to the yield of grain or fruit harvested from a particular crop. As used herein, improved plant productivity refers broadly to improvements in yield of grain, fruit, flowers, or other plant parts harvested for various purposes, improvements in growth of plant parts, including stems, leaves and roots, promotion of plant growth, maintenance of high chlorophyll content in leaves, increasing fruit or seed numbers, increasing fruit or seed unit weight, reducing NO.sub.2 emission due to reduced nitrogen fertilizer usage and similar improvements of the growth and development of plants.
[0106] Microbes in and around food crops can influence the traits of those crops. Plant traits that may be influenced by microbes include: yield (e.g., grain production, biomass generation, fruit development, flower set); nutrition (e.g., nitrogen, phosphorus, potassium, iron, micronutrient acquisition); abiotic stress management (e.g., drought tolerance, salt tolerance, heat tolerance); and biotic stress management (e.g., pest, weeds, insects, fungi, and bacteria). Strategies for altering crop traits include: increasing key metabolite concentrations; changing temporal dynamics of microbe influence on key metabolites; linking microbial metabolite production/degradation to new environmental cues; reducing negative metabolites; and improving the balance of metabolites or underlying proteins.
[0107] As used herein, a "control sequence" refers to an operator, promoter, silencer, or terminator.
[0108] As used herein, "in planta" refers to in the plant, and wherein the plant further comprises plant parts, tissue, leaves, roots, stems, seed, ovules, pollen, flowers, fruit, etc.
[0109] In some embodiments, native or endogenous control sequences of genes of the present disclosure are replaced with one or more intrageneric control sequences.
[0110] As used herein, "introduced" refers to the introduction by means of modern biotechnology, and not a naturally occurring introduction.
[0111] In some embodiments, the bacteria of the present disclosure have been modified such that they are not naturally occurring bacteria.
[0112] In some embodiments, the bacteria of the present disclosure are present in the plant in an amount of at least 10.sup.3 cfu, 10.sup.4 cfu, 10.sup.5 cfu, 10.sup.6 cfu, 10.sup.7 cfu, 10.sup.8 cfu, 10.sup.9 cfu, 10.sup.10 cfu, 10.sup.11 cfu, or 10.sup.12 cfu per gram of fresh or dry weight of the plant. In some embodiments, the bacteria of the present disclosure are present in the plant in an amount of at least about 10.sup.3 cfu, about 10.sup.4 cfu, about 10.sup.5 cfu, about 10.sup.6 cfu, about 10.sup.7 cfu, about 10.sup.8 cfu, about 10.sup.9 cfu, about 10.sup.10 cfu, about 10.sup.11 du, or about 10.sup.12 cfu per gram of fresh or dry weight of the plant. In some embodiments, the bacteria of the present disclosure are present in the plant in an amount of at least 10.sup.3 to 10.sup.9, 10.sup.3 to 10.sup.7, 10.sup.3 to 10.sup.5, 10.sup.5 to 10.sup.9, 10.sup.5 to 10.sup.7, 10.sup.6 to 10.sup.10, 10.sup.6 to 10.sup.7 cfu per gram of fresh or dry weight of the plant.
[0113] Fertilizers and exogenous nitrogen of the present disclosure may comprise the following nitrogen-containing molecules: ammonium, nitrate, nitrite, ammonia, glutamine, etc. Nitrogen sources of the present disclosure may include anhydrous ammonia, ammonia sulfate, urea, diammonium phosphate, urea-form, monoammonium phosphate, ammonium nitrate, nitrogen solutions, calcium nitrate, potassium nitrate, sodium nitrate, etc.
[0114] As used herein, "exogenous nitrogen" refers to non-atmospheric nitrogen readily available in the soil, field, or growth medium that is present under non-nitrogen limiting conditions, including ammonia, ammonium, nitrate, nitrite, urea, uric acid, ammonium acids, etc.
[0115] As used herein, "non-nitrogen limiting conditions" refers to non-atmospheric nitrogen available in the soil, field, media at concentrations greater than about 4 mM nitrogen, as disclosed by Kant et al. (2010. J. Exp. Biol. 62(4):1499-1509), which is incorporated herein by reference.
[0116] As used herein, an "intergeneric microorganism" is a microorganism that is formed by the deliberate combination of genetic material originally isolated from organisms of different taxonomic genera. An "intergeneric mutant" can be used interchangeably with "intergeneric microorganism". An exemplary "intergeneric microorganism" includes a microorganism containing a mobile genetic element which was first identified in a microorganism in a genus different from the recipient microorganism. Further explanation can be found, inter alia, in 40 C.F.R. .sctn. 725.3.
[0117] In aspects, microbes taught herein are "non-intergeneric," which means that the microbes are not intergeneric.
[0118] As used herein, an "intrageneric microorganism" is a microorganism that is formed by the deliberate combination of genetic material originally isolated from organisms of the same taxonomic genera. An "intrageneric mutant" can be used interchangeably with "intrageneric microorganism".
[0119] As used herein, "introduced genetic material" means genetic material that is added to, and remains as a component of, die genome of the recipient.
[0120] In some embodiments, the nitrogen fixation and assimilation genetic regulatory network comprises polynucleotides encoding genes and non-coding sequences that direct, modulate, and/or regulate microbial nitrogen fixation and/or assimilation and can comprise polynucleotide sequences of the nif cluster (e.g., nifA, niFB, nifC, . . . , nifZ), polynucleotides encoding nitrogen regulatory protein C, polynucleotides encoding nitrogen regulatory protein B, polynucleotide sequences of the gin cluster (e.g. glnA and glnD), draT, and ammonia transporters/permeases. In some cases, the Nif cluster may comprise NifB, NifH, NifD, NifK, NifE, NifN, NifX, hesa, and NifV. In some cases, the Nif cluster may comprise a subset of NifB, NifH, NifD, NifK, NifE, NifN, NifX, hesa, and NifV.
[0121] In some embodiments, fertilizer of the present disclosure comprises at least 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 21%, 22%, 23%, 24%, 25%, 26%, 27%, 28%, 29%, 30%, 31%, 32%, 33%, 34%, 35%, 36%, 37%, 38%, 39%, 40%, 41%, 42%, 43%, 44%, 45%, 46%, 47%, 48%, 49%, 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 854/0, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% nitrogen by weight.
[0122] In some embodiments, fertilizer of the present disclosure comprises at least about 5%, about 6%, about 7%, about 8%, about 9%, about 10%, about 11%, about 12%, about 13%, about 14%, about 15%, about 16%, about 17%, about 18%, about 19%, about 20%, about 21%, about 22%, about 23%, about 24%, about 25%, about 26%, about 27%, about 28%, about 29%, about 30%, about 31%, about 32%, about 33%, about 34%, about 35%, about 36%, about 37%, about 38%, about 39%, about 40%, about 41%, about 42%, about 43%, about 44%, about 45%, about 46%, about 47%, about 48%, about 49%, about 50%, about 51%, about 52%, about 53%, about 54%, about 55%, about 56%, about 57%, about 58%, about 59%, about 60%, about 61%, about 62%, about 63%, about 64%, about 65%, about 66%, about 67%, about 68%, about 69%, about 70%, about 71%, about 72%, about 73%, about 74%, about 75%, about 76%, about 77%, about 78%, about 79%, about 80%, about 81%, about 82%, about 83%, about 84%, about 85%, about 86%, about 87%, about 88%, about 89%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, or about 99% nitrogen by weight.
[0123] In some embodiments, fertilizer of the present disclosure comprises about 5% to 50%, about 5% to 75%, about 10% to 50%, about 10% to 75%, about 15% to 50%, about 15% to 75%, about 20% to 50%, about 20% to 75%, about 25% to 50%, about 25% to 75%, about 30% to 50%, about 30% to 75%, about 35% to 50%, about 35% to 75%, about 40% to 50%, about 40% to 75%, about 45% to 50%, about 45% to 75%, or about 50% to 75% nitrogen by weight.
[0124] In some embodiments, the increase of nitrogen fixation and/or the production of 1% or more of the nitrogen in the plant are measured relative to control plants, which have not been exposed to the bacteria of the present disclosure. All increases or decreases in bacteria are measured relative to control bacteria. All increases or decreases in plants are measured relative to control plants.
[0125] As used herein, a "constitutive promoter" is a promoter, which is active under most conditions and/or during most development stages. There are several advantages to using constitutive promoters in expression vectors used in biotechnology, such as: high level of production of proteins used to select transgenic cells or organisms; high level of expression of reporter proteins or storable markers, allowing easy detection and quantification; high level of production of a transcription factor that is part of a regulatory transcription system; production of compounds that requires ubiquitous activity in the organism; and production of compounds that are required during all stages of development. Non-limiting exemplary constitutive promoters include, CaMV 35S promoter, opine promoters, ubiquitin promoter, alcohol dehydrogenase promoter, etc.
[0126] As used herein, a "non-constitutive promoter" is a promoter which is active under certain conditions, in certain types of cells, and/or during certain development stages. For example, tissue specific, tissue preferred, cell type specific, cell type preferred, inducible promoters, and promoters under development control are non-constitutive promoters. Examples of promoters under developmental control include promoters that preferentially initiate transcription in certain tissues.
[0127] As used herein, "inducible" or "repressible" promoter is a promoter which is under chemical or environmental factors control. Examples of environmental conditions that may affect transcription by inducible promoters include anaerobic conditions, certain chemicals, the presence of light, acidic or basic conditions, etc.
[0128] As used herein, a "tissue specific" promoter is a promoter that initiates transcription only in certain tissues. Unlike constitutive expression of genes, tissue-specific expression is the result of several interacting levels of gene regulation. As such, in the art sometimes it is preferable to use promoters from homologous or closely related species to achieve efficient and reliable expression of transgenes in particular tissues. This is one of the main reasons for the large amount of tissue-specific promoters isolated from particular tissues found in both scientific and patent literature.
[0129] As used herein, the term "operably linked" refers to the association of nucleic acid sequences on a single nucleic acid fragment so that the function of one is regulated by the other. For example, a promoter is operably linked with a coding sequence when it is capable of regulating the expression of that coding sequence (i.e., that the coding sequence is under the transcriptional control of the promoter). Coding sequences can be operably linked to regulatory sequences in a sense or antisense orientation. In another example, the complementary RNA regions of the disclosure can be operably linked, either directly or indirectly, 5' to the target mRNA, or 3' to the target mRNA, or within the target mRNA, or a first complementary region is 5' and its complement is 3' to the target mRNA.
[0130] In aspects, "applying to the plant a plurality of non-intergeneric bacteria," includes any means by which the plant (including plant parts such as a seed, root, stem, tissue, etc.) is made to come into contact (i.e. exposed) with said bacteria at any stage of the plant's life cycle. Consequently, "applying to the plant a plurality of non-intergeneric bacteria," includes any of the following means of exposing the plant (including plant parts such as a seed, root, stem, tissue, etc.) to said bacteria: spraying onto plant, dripping onto plant, applying as a seed coat, applying to a field that will then be planted with seed, applying to a field already planted with seed, applying to a field with adult plants, etc.
[0131] As used herein "MRTN" is an acronym for maximum return to nitrogen and is utilized as an experimental treatment in the Examples. MRTN was developed by Iowa State University and information can be found at: http://cnrc.agron.iastate.edu/ The MRTN is the nitrogen rate where the economic net return to nitrogen application is maximized. The approach to calculating the MRTN is a regional approach for developing corn nitrogen rate guidelines in individual states. The nitrogen rate trial data was evaluated for Illinois, Iowa, Michigan, Minnesota, Ohio, and Wisconsin where an adequate number of research trials were available for corn plantings following soybean and corn plantings following corn. The trials were conducted with spring, sidedress, or split preplant/sidedress applied nitrogen, and sites were not irrigated except for those that were indicated for irrigated sands in Wisconsin. MRTN was developed by Iowa State University due to apparent differences in methods for determining suggested nitrogen rates required for corn production, misperceptions pertaining to nitrogen rate guidelines, and concerns about application rates. By calculating the MRTN, practitioners can determine the following: (1) the nitrogen rate where the economic net return to nitrogen application is maximized, (2) the economic optimum nitrogen rate, which is the point where the last increment of nitrogen returns a yield increase large enough to pay for the additional nitrogen, (3) the value of corn grain increase attributed to nitrogen application, and the maximum yield, which is the yield where application of more nitrogen does not result in a corn yield increase. Thus the MRTN calculations provide practitioners with the means to maximize corn crops in different regions while maximizing financial gains from nitrogen applications.
[0132] The term mmol is an abbreviation for millimole, which is a thousandth (10.sup.-3) of a mole, abbreviated herein as mol.
[0133] As used herein the terms "microorganism" or "microbe" should be taken broadly. These terms, used interchangeably, include but are not limited to, the two prokaryotic domains, Bacteria and Archaea. The term may also encompass eukaryotic fungi and protists.
[0134] The term "microbial consortia" or "microbial consortium" refers to a subset of a microbial community of individual microbial species, or strains of a species, which can be described as carrying out a common function, or can be described as participating in, or leading to, or correlating with, a recognizable parameter, such as a phenotypic trait of interest.
[0135] The term "microbial community" means a group of microbes comprising two or more species or strains. Unlike microbial consortia, a microbial community does not have to be carrying out a common function, or does not have to be participating in, or leading to, or correlating with, a recognizable parameter, such as a phenotypic trait of interest.
[0136] As used herein, "isolate," "isolated," "isolated microbe," and like terms, are intended to mean that the one or more microorganisms has been separated from at least one of the materials with which it is associated in a particular environment (for example soil, water, plant tissue, etc.). Thus, an "isolated microbe" does not exist in its naturally occurring environment; rather, it is through the various techniques described herein that the microbe has been removed from its natural setting and placed into a non-naturally occurring state of existence. Thus, the isolated strain or isolated microbe may exist as, for example, a biologically pure culture, or as spores (or other forms of the strain). In aspects, the isolated microbe may be in association with an acceptable carrier, which may be an agriculturally acceptable carrier.
[0137] In certain aspects of the disclosure, the isolated microbes exist as "isolated and biologically pure cultures." It will be appreciated by one of skill in the art, that an isolated and biologically pure culture of a particular microbe, denotes that said culture is substantially free of other living organisms and contains only the individual microbe in question. The culture can contain varying concentrations of said microbe. The present disclosure notes that isolated and biologically pure microbes often "necessarily differ from less pure or impure materials." See, e.g. In re Bergstrom, 427 F.2d 1394, (CCPA 1970)(discussing purified prostaglandins), see also, In re Bergy, 596 F.2d 952 (CCPA 1979)(discussing purified microbes), see also, Parke-Davis & Co. v. H.K. Mulford & Co., 189 F. 95 (S.D.N.Y. 1911) (Learned Hand discussing purified adrenaline), aff'd in part, rev'd in part, 196 F. 496 (2d Cir. 1912), each of which are incorporated herein by reference. Furthermore, in some aspects, the disclosure provides for certain quantitative measures of the concentration, or purity limitations, that must be found within an isolated and biologically pure microbial culture. The presence of these purity values, in certain embodiments, is a further attribute that distinguishes the presently disclosed microbes from those microbes existing in a natural state. See, e.g., Merck & Co. v. Olin Mathieson Chemical Corp., 253 F.2d 156 (4th Cir. 1958) (discussing purity limitations for vitamin B12 produced by microbes), incorporated herein by reference.
[0138] As used herein, "individual isolates" should be taken to mean a composition, or culture, comprising a predominance of a single genera, species, or strain, of microorganism, following separation from one or more other microorganisms.
[0139] Microbes of the present disclosure may include spores and/or vegetative cells. In some embodiments, microbes of the present disclosure include microbes in a viable but non-culturable (VBNC) state. As used herein, "spore" or "spores" refer to structures produced by bacteria and fungi that are adapted for survival and dispersal. Spores are generally characterized as dormant structures; however, spores are capable of differentiation through the process of germination. Germination is the differentiation of spores into vegetative cells that are capable of metabolic activity, growth, and reproduction. The germination of a single spore results in a single fungal or bacterial vegetative cell. Fungal spores are units of asexual reproduction, and in some cases are necessary structures in fungal life cycles. Bacterial spores are structures for surviving conditions that may ordinarily be nonconducive to the survival or growth of vegetative cells.
[0140] As used herein, "microbial composition" refers to a composition comprising one or more microbes of the present disclosure. In some embodiments, a microbial composition is administered to plants (including various plant parts) and/or in agricultural fields.
[0141] As used herein, "carrier," "acceptable carrier," or "agriculturally acceptable carrier" refers to a diluent, adjuvant, excipient, or vehicle with which the microbe can be administered, which does not detrimentally effect the microbe.
Regulation of Nitrogen Fixation
[0142] In some cases, nitrogen fixation pathway may act as a target for genetic engineering and optimization. One trait that may be targeted for regulation by the methods described herein is nitrogen fixation. Nitrogen fertilizer is the largest operational expense on a farm and the biggest driver of higher yields in row crops like corn and wheat. Described herein are microbial products that can deliver renewable forms of nitrogen in non-leguminous crops. While some endophytes have the genetics necessary for fixing nitrogen in pure culture, the fundamental technical challenge is that wild-type endophytes of cereals and grasses stop fixing nitrogen in fertilized fields. The application of chemical fertilizers and residual nitrogen levels in field soils signal the microbe to shut down the biochemical pathway for nitrogen fixation.
[0143] Changes to the transcriptional and post-translational levels of components of the nitrogen fixation regulatory network may be beneficial to the development of a microbe capable of fixing and transferring nitrogen to corn in the presence of fertilizer. To that end, described herein is Host-Microbe Evolution (HoME) technology to precisely evolve regulatory networks and elicit novel phenotypes. Also described herein are unique, proprietary libraries of nitrogen-fixing endophytes isolated from corn, paired with extensive omics data surrounding the interaction of microbes and host plant under different environmental conditions like nitrogen stress and excess. In some embodiments, this technology enables precision evolution of the genetic regulatory network of endophytes to produce microbes that actively fix nitrogen even in the presence of fertilizer in the field. Also described herein are evaluations of the technical potential of evolving microbes that colonize corn root tissues and produce nitrogen for fertilized plants and evaluations of the compatibility of endophytes with standard formulation practices and diverse soils to determine feasibility of integrating the microbes into modern nitrogen management strategies.
[0144] In order to utilize elemental nitrogen (N) for chemical synthesis, life forms combine nitrogen gas (N.sub.2) available in the atmosphere with hydrogen in a process known as nitrogen fixation. Because of the energy-intensive nature of biological nitrogen fixation, diazotrophs (bacteria and archaea that fix atmospheric nitrogen gas) have evolved sophisticated and fight regulation of the nif gene cluster in response to environmental oxygen and available nitrogen. Nif genes encode enzymes involved in nitrogen fixation (such as the nitrogenase complex) and proteins that regulate nitrogen fixation. Shamseldin (2013. Global J. Biotechnol. Biochem. 8(4):84-94) discloses detailed descriptions of nif genes and their products, and is incorporated herein by reference. Described herein are methods of producing a plant with an improved trait comprising isolating bacteria from a first plant, introducing a genetic variation into a gene of the isolated bacteria to increase nitrogen fixation, exposing a second plant to the variant bacteria, isolating bacteria from the second plant having an improved trait relative to the first plant, and repeating the steps with bacteria isolated from the second plant.
[0145] In Proteobacteria, regulation of nitrogen fixation centers around the .sigma..sub.54-dependent enhancer-binding protein NifA, the positive transcriptional regulator of the nif cluster. Intracellular levels of active NifA are controlled by two key factors: transcription of the nifLA operon, and inhibition of NifA activity by protein-protein interaction with NifL. Both of these processes are responsive to intraceullar glutamine levels via the PII protein signaling cascade. This cascade is mediated by GlnD, which directly senses glutamine and catalyzes the uridylylation or deuridylylation of two PII regulatory proteins--GlnB and GlnK--in response the absence or presence, respectively, of bound glutamine. Under conditions of nitrogen excess, unmodified GlnB signals the deactivation of the nifLA promoter. However, under conditions of nitrogen limitation, GlnB is post-translationally modified, which inhibits its activity and Leads to transcription of the nifLA operon. In this way, nifLA transcription is tightly controlled in response to environmental nitrogen via the PII protein signaling cascade. On the post-translational level of NifA regulation, GlnK inhibits the NifL/NifA interaction in a matter dependent on the overall level of free GlnK within the cell.
[0146] NifA is transcribed from the nifLA operon, whose promoter is activated by phosphorylated NtrC, another .sigma..sub.54-dependent regulator. The phosphorylation state of NtrC is mediated by the histidine kinase NtrB, which interacts with deuridylylated GlnB but not undylylated GlnB. Under conditions of nitrogen excess, a high intracellular level of glutamine leads to deuridylylation of GlnB, which then interacts with NtrB to deactivate its phosphorylation activity and activate its phosphatase activity, resulting in dephosphorylation of NtrC and the deactivation of the nifLA promoter. However, under conditions of nitrogen limitation, a low level of intracellular glutamine results in uridylylation of GlnB, which inhibits its interaction with NtrB and allows the phosphorylation of NtrC and transcription of the nifLA operon. In this way, nifLA expression is tightly controlled in response to environmental nitrogen via the PII protein signaling cascade. nifA, ntrB, ntrC, and glnB, are all genes that can be mutated in the methods described herein. These processes may also be responsive to intracellular or extracellular levels of ammonia, urea or nitrates.
[0147] The activity of NifA is also regulated post-translationally in response to environmental nitrogen, most typically through Nil-mediated inhibition of NifA activity. In general, the interaction of NifL and NifA is influenced by the PII protein signaling cascade via GlnK, although the nature of the interactions between GlnK and NifL/NifA varies significantly between diazotrophs. In Klebsiella pneumoniae, both forms of GlnK inhibit the NifL/NifA interaction, and the interaction between GlnK and NifL/NifA is determined by the overall level of free GlnK within the cell. Under nitrogen-excess conditions, deuridylylated GlnK interacts with the ammonium transporter AmtB, which serves to both block ammonium uptake by AmtB and sequester GlnK to the membrane, allowing inhibition of NifA by NifL. On the other hand, in Azotobacter vinelandii, interaction with deuridylylated GlnK is required for the NifL/NifA interaction and NifA inhibition, while uridylylation of GlnK inhibits its interaction with Nil. In diazotrophs lacking the nifL gene, there is evidence that NifA activity is inhibited directly by interaction with the deuridylylated forms of both GlnK and GlnB under nitrogen-excess conditions. In some bacteria the Nif cluster may be regulated by glnR, and further in some cases this may comprise negative regulation. Regardless of the mechanism, post-translational inhibition of NifA is an important regulator of the nif cluster in most known diazotrophs. Additionally, nifL, amtB, glnK, and glnR are genes that can be mutated in the methods described herein.
[0148] In addition to regulating the transcription of the nif gene cluster, many diazotrophs have evolved a mechanism for the direct post-translational modification and inhibition of the nitrogenase enzyme itself, known as nitrogenase shutoff. This is mediated by ADP-ribosylation of the Fe protein (NifH) under nitrogen-excess conditions, which disrupts its interaction with the MoFe protein complex (NifDK) and abolishes nitrogenase activity. DraT catalyzes the ADP-ribosylation of the Fe protein and shutoff of nitrogenase, while DraG catalyzes the removal of ADP-ribose and reactivation of nitrogenase. As with nifLA transcription and NifA inhibition, nitrogenase shutoff is also regulated via the PII protein signaling cascade. Under nitrogen-excess conditions, deuridylylated GlnB interacts with and activates DraT, while deuridylylated GlnK interacts with both DraG and AmtB to form a complex, sequestering DraG to the membrane. Under nitrogen-limiting conditions, the uridylylated forms of GlnB and GlnK do not interact with DraT and DraG, respectively, leading to the inactivation of DraT and the diffusion of DraG to the Fe protein, where it removes the ADP-ribose and activates nitrogenase. The methods described herein also contemplate introducing genetic variation into the nifH, nifD, nifK, and draT genes.
[0149] Although some endophytes have the ability to fix nitrogen in vitro, often the genetics are silenced in the field by high levels of exogenous chemical fertilizers. One can decouple the sensing of exogenous nitrogen from expression of the nitrogenase enzyme to facilitate field-based nitrogen fixation. Improving the integral of nitrogenase activity across time further serves to augment the production of nitrogen for utilization by the crop. Specific targets for genetic variation to facilitate field-based nitrogen fixation using the methods described herein include one or more genes selected from the group consisting of nifA, ntrB, ntrC, glnA, glnB, glnK, draT, amtB, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, and nifQ.
[0150] An additional target for genetic variation to facilitate field-based nitrogen fixation using the methods described herein is the NifA protein. The NifA protein is typically the activator for expression of nitrogen fixation genes. Increasing the production of NifA (either constitutively or during high ammonia condition) circumvents the native ammonia-sensing pathway. In addition, reducing the production of NifL proteins, a known inhibitor of NifA, also leads to an increased level of freely active NifA. In addition, increasing the transcription level of the nifAL operon (either constitutively or during high ammonia condition) also leads to an overall higher level of NifA proteins. Elevated level of nifAL expression is achieved by altering the promoter itself or by reducing the expression of NtrB (part of ntrB and ntrC signaling cascade that originally would result in the shutoff of nifAL operon during high nitrogen condition). High level of NifA achieved by these or any other methods described herein increases the nitrogen fixation activity of the endophytes.
[0151] Another target for genetic variation to facilitate field-based nitrogen fixation using the methods described herein is the glnD/GlnB/GlnK PII signaling cascade. The intracellular glutamine level is sensed through the GlnD/GlnB/GlnK PII signaling cascade. Active site mutations in GlnD that abolish the uridylyl-removing activity of GlnD disrupt the nitrogen-sensing cascade. In addition, reduction of the GlnB concentration short circuits the glutamine-sensing cascade. These mutations "trick" the cells into perceiving a nitrogen-limited state, thereby increasing the nitrogen fixation level activity. These processes may also be responsive to intracellular or extracellular levels of ammonia, urea or nitrates.
[0152] The amtB protein is also a target for genetic variation to facilitate field-based nitrogen fixation using the methods described herein. Ammonia uptake from the environment can be reduced by decreasing the expression level of amtB protein. Without intracellular ammonia, the endophyte is not able to sense the high level of ammonia, preventing the down-regulation of nitrogen fixation genes. Any ammonia that manages to get into the intracellular compartment is converted into glutamine. Intracellular glutamine level is the major currency of nitrogen sensing. Decreasing the intracellular glutamine level prevents the cells from sensing high ammonium levels in the environment. This effect can be achieved by increasing the expression level of glutaminase, an enzyme that converts glutamine into glutamate. In addition, intracellular glutamine can also be reduced by decreasing glutamine synthase (an enzyme that converts ammonia into glutamine). In diazotrophs, fixed ammonia is quickly assimilated into glutamine and glutamate to be used for cellular processes. Disruptions to ammonia assimilation may enable diversion of fixed nitrogen to be exported from the cell as ammonia. The fixed ammonia is predominantly assimilated into glutamine by glutamine synthetase (GS), encoded by glnA, and subsequently into glutamine by glutamine oxoglutarate aminotransferase (GOGAT). In some examples, glnS encodes a glutamine synthetase. GS is regulated post-translationally by GS adenylyl transferase (GlnE), a bi-functional enzyme encoded by glnE that catalyzes both the adenylylation and deadenylylation of GS through activity of its adenylyl-transferase (AT) and adenylyl-removing (AR) domains, respectively. Under nitrogen limiting conditions, glnA is expressed, and GlnE's AR domain de-adynylylates GS, allowing it to be active. Under conditions of nitrogen excess, glnA expression is turned off, and GlnE's AT domain is activated allosterically by glutamine, causing the adenylylation and deactivation of GS.
[0153] Furthermore, the draT gene may also be a target for genetic variation to facilitate field-based nitrogen fixation using the methods described herein. Once nitrogen fixing enzymes are produced by the cell, nitrogenase shut-off represents another level in which cell downregulates fixation activity in high nitrogen condition. This shut-off could be removed by decreasing the expression level of DraT.
[0154] Methods for imparting new microbial phenotypes can be performed at the transcriptional, translational, and post-translational levels. The transcriptional level includes changes at the promoter (such as changing sigma factor affinity or binding sites for transcription factors, including deletion of all or a portion of the promoter) or changing transcription terminators and attenuators. The translational level includes changes at the ribosome binding sites and changing mRNA degradation signals. The post-translational level includes mutating an enzyme's active site and changing protein-protein interactions. These changes can be achieved in a multitude of ways. Reduction of expression level (or complete abolishment) can be achieved by swapping the native ribosome binding site (RBS) or promoter with another with lower strength/efficiency. ATG start sites can be swapped to a GTG, TTG, or CTG start codon, which results in reduction in translational activity of the coding region. Complete abolishment of expression can be done by knocking out (deleting) the coding region of a gene. Frameshifting the open reading frame (ORF) likely will result in a premature stop codon along the ORF, thereby creating a non-functional truncated product. Insertion of in-frame stop codons will also similarly create a non-functional truncated product. Addition of a degradation tag at the N or C terminal can also be done to reduce the effective concentration of a particular gene.
[0155] Conversely, expression level of the genes described herein can be achieved by using a stronger promoter. To ensure high promoter activity during high nitrogen level condition (or any other condition), a transcription profile of the whole genome in a high nitrogen level condition could be obtained and active promoters with a desired transcription level can be chosen from that dataset to replace the weak promoter. Weak start codons can be swapped out with an ATG start codon for better translation initiation efficiency. Weak ribosomal binding sites (RBS) can also be swapped out with a different RBS with higher translation initiation efficiency. In addition, site specific mutagenesis can also be performed to alter the activity of an enzyme.
[0156] Increasing the level of nitrogen fixation that occurs in a plain can lead to a reduction in the amount of chemical fertilizer needed for crop production and reduce greenhouse gas emissions (e.g., nitrous oxide).
Generation of Bacterial Populations
Isolation of Bacteria
[0157] Microbes useful in methods and compositions disclosed herein can be obtained by extracting microbes from surfaces or tissues of native plants. Microbes can be obtained by grinding seeds to isolate microbes. Microbes can be obtained by planting seeds in diverse soil samples and recovering microbes from tissues. Additionally, microbes can be obtained by inoculating plants with exogenous microbes and determining which microbes appear in plant tissues. Non-limiting examples of plant tissues may include a seed, seedling, leaf cutting, plant, bulb, or tuber.
[0158] A method of obtaining microbes may be through the isolation of bacteria from soils. Bacteria may be collected from various soil types. In some example, the soil can be characterized by traits such as high or low fertility, levels of moisture, levels of minerals, and various cropping practices. For example, the soil may be involved in a crop rotation where different crops are planted in the same soil in successive planting seasons. The sequential growth of different crops on the same soil may prevent disproportionate depletion of certain minerals. The bacteria can be isolated from the plants growing in the selected soils. The seedling plants can be harvested at 2-6 weeks of growth. For example, at least 400 isolates can be collected in a round of harvest. Soil and plant types reveal the plant phenotype as well as the conditions, which allow for the downstream enrichment of certain phenotypes.
[0159] Microbes can be isolated from plant tissues to assess microbial traits. The parameters for processing tissue samples may be varied to isolate different types of associative microbes, such as rhizopheric bacteria, epiphytes, or endophytes. The isolates can be cultured in nitrogen-free media to enrich for bacteria that perform nitrogen fixation. Alternatively, microbes can be obtained from global strain banks.
[0160] In planta analytics are performed to assess microbial traits. In some embodiments, the plant tissue can be processed for screening by high throughput processing for DNA and RNA. Additionally, non-invasive measurements can be used to assess plant characteristics, such as colonization. Measurements on wild microbes can be obtained on a plant-by-plant basis. Measurements on wild microbes can also be obtained in the field using medium throughput methods. Measurements can be done successively over time. Model plant system can be used including, but not limited to, Setaria.
[0161] Microbes in a plant system can be screened via transcriptional profiling of a microbe in a plant system. Examples of screening through transcriptional profiling are using methods of quantitative polymerase chain reaction (qPCR), molecular barcodes for transcript detection, Next Generation Sequencing, and microbe tagging with fluorescent markers. Impact factors can be measured to assess colonization in the greenhouse including, but not limited to, microbiome, abiotic factors, soil conditions, oxygen, moisture, temperature, inoculum conditions, and root localization. Nitrogen fixation can be assessed in bacteria by measuring 15N gas/fertilizer (dilution) with IRMS or NanoSIMS as described herein NanoSIMS is high-resolution secondary ion mass spectrometry. The NanoSIMS technique is a way to investigate chemical activity from biological samples. The catalysis of reduction of oxidation reactions that drive the metabolism of microorganisms can be investigated at the cellular, subcellular, molecular and elemental level. NanoSIMS can provide high spatial resolution of greater than 0.1 .mu.m. NanoSIMS can detect the use of isotope tracers such as .sup.13C, .sup.15N, and .sup.18O. Therefore, NanoSIMS can be used to the chemical activity nitrogen in the cell.
[0162] Automated greenhouses can be used for planta analytics. Plant metrics in response to microbial exposure include, but are not limited to, biomass, chloroplast analysis, CCD camera, volumetric tomography measurements.
[0163] One way of enriching a microbe population is according to genotype. For example, a polymerase chain reaction (PCR) assay with a targeted primer or specific primer. Primers designed for the nifH gene can be used to identity diazotrophs because diazotrophs express the nifH gene in the process of nitrogen fixation. A microbial population can also be enriched via single-cell culture-independent approaches and chemotaxis-guided isolation approaches. Alternatively, targeted isolation of microbes can be performed by culturing the microbes on selection media. Premeditated approaches to enriching microbial populations for desired traits can be guided by bioinformatics data and are described herein.
Enriching for Microbes with Nitrogen Fixation Capabilities Using Bioinformatics
[0164] Bioinformatic tools can be used to identify and isolate plant growth promoting rhizobacteria (PGPRs), which are selected based on their ability to perform nitrogen fixation. Microbes with high nitrogen fixing ability can promote favorable traits in plants. Bioinformatic modes of analysis for the identification of PGPRs include, but are not limited to, genomics, metagenomics, targeted isolation, gene sequencing, transcriptome sequencing, and modeling.
[0165] Genomics analysis can be used to identify PGPRs and confirm the presence of mutations with methods of Next Generation Sequencing as described herein and microbe version control.
[0166] Metagenomics can be used to identify and isolate PGPR using a prediction algorithm for colonization. Metadata can also be used to identify the presence of an engineered strain in environmental and greenhouse samples.
[0167] Transcriptomic sequencing can be used to predict genotypes leading to PGPR phenotypes. Additionally, transcriptomic data is used to identify promoters for altering gene expression. Transcriptomic data can be analyzed in conjunction with the Whole Genome Sequence (WGS) to generate models of metabolism and gene regulatory networks.
Domestication of Microbes
[0168] Microbes isolated from nature can undergo a domestication process wherein the microbes are converted to a form that is genetically trackable and identifiable. One way to domesticate a microbe is to engineer it with antibiotic resistance. The process of engineering antibiotic resistance can begin by determining the antibiotic sensitivity in the wild type microbial strain. If the bacteria are sensitive to the antibiotic, then the antibiotic can be a good candidate for antibiotic resistance engineering. Subsequently, an antibiotic resistant gene or a counterselectable suicide vector can be incorporated into the genome of a microbe using recombineering methods. A counterselectable suicide vector may consist of a deletion of the gene of interest, a selectable marker, and the counterselectable marker sacB. Counterselection can be used to exchange native microbial DNA sequences with antibiotic resistant genes. A medium throughput method can be used to evaluate multiple microbes simultaneously allowing for parallel domestication. Alternative methods of domestication include the use of homing nucleases to prevent the suicide vector sequences from looping out or from obtaining intervening vector sequences.
[0169] DNA vectors can be introduced into bacteria via several methods including electroporation and chemical transformations. A standard library of vectors can be used for transformations. An example of a method of gene editing is CRISPR preceded by Cas9 testing to ensure activity of Cas9 in the microbes.
Non-Transgenic Engineering of Microbes
[0170] A microbial population with favorable traits can be obtained via directed evolution. Direct evolution is an approach wherein the process of natural selection is mimicked to evolve proteins or nucleic acids towards a user-defined goal. An example of direct evolution is when random mutations are introduced into a microbial population, the microbes with the most favorable traits are selected, and the growth of the selected microbes is continued. The most favorable traits in growth promoting rhizobacteria (PGPRs) may be in nitrogen fixation. The method of directed evolution may be iterative and adaptive based on the selection process after each iteration.
[0171] Plant growth promoting rhizobacteria (PGPRs) with high capability of nitrogen fixation can be generated. The evolution of PGPRs can be carried out via the introduction of genetic variation. Genetic variation can be introduced via polymerase chain reaction mutagenesis, oligonucleotide-directed mutagenesis, saturation mutagenesis, fragment shuffling mutagenesis, homologous recombination, CRISPR/Cas9 systems, chemical mutagenesis, and combinations thereof. These approaches can introduce random mutations into the microbial population. For example, mutants can be generated using synthetic DNA or RNA via oligonucleotide-directed mutagenesis. Mutants can be generated using tools contained on plasmids, which are later cured. Genes of interest can be identified using libraries from other species with improved traits including, but not limited to, improved PGPR properties, improved colonization of cereals, increased oxygen sensitivity, increased nitrogen fixation, and increased ammonia excretion. Intrageneric genes can be designed based on these libraries using software such as Geneious or Platypus design software. Mutations can be designed with the aid of machine learning, Mutations can be designed with the aid of a metabolic model. Automated design of the mutation can be done using a la Platypus and will guide RNAs for Cas-directed mutagenesis.
[0172] The intra-generic genes can be transferred into the host microbe. Additionally, reporter systems can also be transferred to the microbe. The reporter systems characterize promoters, determine the transformation success, screen mutants, and act as negative screening tools.
[0173] The microbes carrying the mutation can be cultured via serial passaging. A microbial colony contains a single variant of the microbe. Microbial colonies are screened with the aid of an automated colony picker and liquid handler. Mutants with gene duplication and increased copy number express a higher genotype of the desired trait.
Selection of Plant Growth Promoting Microbes Based on Nitrogen Fixation
[0174] The microbial colonies can be screened using various assays to assess nitrogen fixation. One way to measure nitrogen fixation is via a single fermentative assay, which measures nitrogen excretion. An alternative method is the acetylene reduction assay (ARA) with in-line sampling over time. ARA can be performed in high throughput plates of microtube arrays. ARA can be performed with live plants and plant tissues. The media formulation and media oxygen concentration can be varied in ARA assays. Another method of screening microbial variants is by using biosensors. The use of NanoSIMS and Raman microspectroscopy can be used to investigate the activity of the microbes. In some cases, bacteria can also be cultured and expanded using methods of fermentation in bioreactors. The bioreactors are designed to improve robustness of bacteria growth and to decrease the sensitivity of bacteria to oxygen. Medium to high TP plate-based microfermentors are used to evaluate oxygen sensitivity, nutritional needs, nitrogen fixation, and nitrogen excretion. The bacteria can also be co-cultured with competitive or beneficial microbes to elucidate cryptic pathways. Flow cytometry can be used to screen for bacteria that produce high levels of nitrogen using chemical, colorimetric, or fluorescent indicators. The bacteria may be cultured in the presence or absence of a nitrogen source. For example, the bacteria may be cultured with glutamine, ammonia, urea or nitrates.
Microbe Breeding
[0175] Microbe breeding is a method to systematically identify and improve the role of species within the crop microbiome. The method comprises three steps: 1) selection of candidate species by mapping plant-microbe interactions and predicting regulatory networks linked to a particular phenotype, 2) pragmatic and predictable improvement of microbial phenotypes through intra-species crossing of regulatory networks and gene clusters, and 3) screening and selection of new microbial genotypes that produce desired crop phenotypes. To systematically assess the improvement of strains, a model is created that links colonization dynamics of the microbial community to genetic activity by key species. The model is used to predict genetic targets breeding and improve the frequency of selecting improvements in microbiome-encoded traits of agronomic relevance. See, FIG. 17A for a graphical representation of an embodiment of the process. In particular, FIG. 17A depicts a schematic of microbe breeding, in accordance with embodiments. As illustrated in FIG. 17A, rational improvement of the crop microbiome may be used to increase soil biodiversity, tune impact of keystone species, and/or alter timing and expression of important metabolic pathways. To this end, the inventors have developed a microbe breeding pipeline to identify and improve the role of strains within the crop microbiome. The method comprises three steps: 1) selection of candidate species by mapping plant-microbe interactions and predicting regulatory networks linked to a particular phenotype, 2) pragmatic and predictable improvement of microbial phenotypes through intragenomic crossing of gene regulatory networks and gene clusters, and 3) screening and selection of new microbial genotypes that produce desired crop phenotypes. To systematically assess the improvement of strains, the inventors employ a model that links colonization dynamics of the microbial community to genetic activity by key species. This process represents a methodology for breeding and selecting improvements in microbiome-encoded traits of agronomic relevance.
[0176] Production of bacteria to improve plant traits (e.g., nitrogen fixation) can be achieved through serial passage. The production of this bacteria can be done by selecting plants, which have a particular improved trait that is influenced by the microbial flora, in addition to identifying bacteria and/or compositions that are capable of imparting one or more improved traits to one or more plants. One method of producing a bacteria to improve a plant trait includes the steps of: (a) isolating bacteria from tissue or soil of a first plant; (b) introducing a genetic variation into one or more of the bacteria to produce one or more variant bacteria; (c) exposing a plurality of plants to the variant bacteria; (d) isolating bacteria from tissue or soil of one of the plurality of plants, wherein the plant from which the bacteria is isolated has an improved trait relative to other plants in the plurality of plants; and (e) repeating steps (b) to (d) with bacteria isolated from the plant with an improved trait (step (d)). Steps (b) to (d) can be repeated any number of times (e.g., once, twice, three times, four times, five times, ten times, or more) until the improved trait in a plant reaches a desired level. Further, the plurality of plants can be more than two plants, such as 10 to 20 plants, or 20 or more, 50 or more, 100 or more, 300 or more, 500 or more, or 1000 or more plants.
[0177] In addition to obtaining a plant with an improved trait, a bacterial population comprising bacteria comprising one or more genetic variations introduced into one or more genes (e.g., genes regulating nitrogen fixation) is obtained. By repeating the steps described above, a population of bacteria can be obtained that include the most appropriate members of the population that correlate with a plant trait of interest. The bacteria in this population can be identified and their beneficial properties determined, such as by genetic and/or phenotypic analysis. Genetic analysis may occur of isolated bacteria in step (a). Phenotypic and/or genotypic information may be obtained using techniques including: high through-put screening of chemical components of plant origin, sequencing techniques including high throughput sequencing of genetic material, differential display techniques (including DDRT-PCR, and DD-PCR), nucleic acid microarray techniques, RNA-sequencing (Whole Transcriptome Shotgun Sequencing), and qRT-PCR (quantitative real time PCR). Information gained can be used to obtain community profiling information on the identity and activity of bacteria present, such as phylogenetic analysis or microarray-based screening of nucleic acids coding for components of rRNA operons or other taxonomically informative loci. Examples of taxonomically informative loci include 16S rRNA gene, 23S rRNA gene, 5S rRNA gene, 5.85 rRNA gene, 12S rRNA gene, 18S rRNA gene, 28S rRNA gene, gyrB gene, rpoB gene, fusA gene, recA gene, coxl gene, nifD gene. Example processes of taxonomic profiling to determine taxa present in a population are described in US20140155283. Bacterial identification may comprise characterizing activity of one or more genes or one or more signaling pathways, such as genes associated with the nitrogen fixation pathway. Synergistic interactions (where two components, by virtue of their combination, increase a desired effect by more than an additive amount) between different bacterial species may also be present in the bacterial populations.
Genetic Variation--Locations and Sources of Genomic Alteration
[0178] The genetic variation may be a gene selected from the group consisting of: nifA, nifL, ntrB, ntrC, glnA, glnB, glnK, draT, amtB, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, and nifQ. The genetic variation may be a variation in a gene encoding a protein with functionality selected from the group consisting of: glutamine synthetase, glutaminase, glutamine synthetase adenylyltransferase, transcriptional activator, anti-transcriptional activator, pyruvate flavodoxin oxidoreductase, flavodoxin, or NAD+-dinitrogen-reductase aDP-D-ribosyltransferase. The genetic variation may be a mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. Introducing a genetic variation may comprise insertion and/or deletion of one or more nucleotides at a target site, such as 1, 2, 3, 4, 5, 10, 25, 50, 100, 250, 500, or more nucleotides. The genetic variation introduced into one or more bacteria of the methods disclosed herein may be a knock-out mutation (e.g. deletion of a promoter, insertion or deletion to produce a premature stop codon, deletion of an entire gene), or it may be elimination or abolishment of activity of a protein domain (e.g. point mutation affecting an active site, or deletion of a portion of a gene encoding the relevant portion of the protein product), or it may alter or abolish a regulatory sequence of a target gene. One or more regulatory sequences may also be inserted, including heterologous regulatory sequences and regulatory sequences found within a genome of a bacterial species or genus corresponding to the bacteria into which the genetic variation is introduced. Moreover, regulatory sequences may be selected based on the expression level of a gene in a bacterial culture or within a plant tissue. The genetic variation may be a pre-determined genetic variation that is specifically introduced to a target site. The genetic variation may be a random mutation within the target site. The genetic variation may be an insertion or deletion of one or more nucleotides. In some cases, a plurality of different genetic variations (e.g. 2, 3, 4, 5, 10, or more) are introduced into one or more of the isolated bacteria before exposing the bacteria to plants for assessing trait improvement. The plurality of genetic variations can be any of the above types, the same or different types, and in any combination. In some cases, a plurality of different genetic variations are introduced serially, introducing a first genetic variation after a first isolation step, a second genetic variation after a second isolation step, and so forth so as to accumulate a plurality of genetic variations in bacteria imparting progressively improved traits on the associated plants.
Genetic Variation--Methods of Introducing Genomic Alteration
[0179] In general, the term "genetic variation" refers to any change introduced into a polynucleotide sequence relative to a reference polynucleotide, such as a reference genome or portion thereof, or reference gene or portion thereof. A genetic variation may be referred to as a "mutation," and a sequence or organism comprising a genetic variation may be referred to as a "genetic variant" or "mutant". Genetic variations can have any number of effects, such as the increase or decrease of some biological activity, including gene expression, metabolism, and cell signaling. Genetic variations can be specifically introduced to a target site, or introduced randomly. A variety of molecular tools and methods are available for introducing genetic variation. For example, genetic variation can be introduced via polymerase chain reaction mutagenesis, oligonucleotide-directed mutagenesis, saturation mutagenesis, fragment shuffling mutagenesis, homologous recombination, recombineering, lambda red mediated recombination, CRISPR/Cas9 systems, chemical mutagenesis, and combinations thereof. Chemical methods of introducing genetic variation include exposure of DNA to a chemical mutagen, e.g., ethyl methanesulfonate (EMS), methyl methanesulfonate (MMS), N-nitrosourea (EN U), N-methyl-N-nitro-N'-nitrosoguanidine, 4-nitroquinoline N-oxide, diethylsulfate, benzopyrene, cyclophosphamide, bleomycin, trimethylmelamine, acrylamide monomer, nitrogen mustard, vincristine, diepoxyalkanes (for example, diepoxybutane), ICR-170, formaldehyde, procarbazine hydrochloride, ethylene oxide, dimethylnitrosamine, 7,12 dimethylbenz(a)anthracene, chlorambucil, hexamethylphosphoramide, bisulfan, and the like. Radiation mutation-inducing agents include ultraviolet radiation, .gamma.-irradiation, X-rays, and fast neutron bombardment. Genetic variation can also be introduced into a nucleic acid using, e.g., trimethylpsoralen with ultraviolet light. Random or targeted insertion of a mobile DNA element, e.g., a transposable element, is another suitable method for generating genetic variation. Genetic variations can be introduced into a nucleic acid during amplification in a cell-free in vitro system, e.g., using a polymerase chain reaction (PCR) technique such as error-prone PCR. Genetic variations can be introduced into a nucleic acid in vitro using DNA shuffling techniques (e.g., exon shuffling, domain swapping, and the like). Genetic variations can also be introduced into a nucleic acid as a result of a deficiency in a DNA repair enzyme in a cell, e.g., the presence in a cell of a mutant gene encoding a mutant DNA repair enzyme is expected to generate a high frequency of mutations (i.e., about 1 mutation/100 genes-1 mutation/10,000 genes) in the genome of the cell. Examples of genes encoding DNA repair enzymes include but are not limited to Mut H, Mut S, Mut L, and Mut U, and the homologs thereof in other species (e.g., MSH 1 6, PMS 1 2, MLH 1, GTBP, ERCC-1, and the like). Example descriptions of various methods for introducing genetic variations are provided in e.g., Stemple (2004) Nature 5:1-7; Chiang et al. (1993) PCR Methods Appl 2(3): 210-217; Stemmer (1994) Proc. Natl. Acad. Sci. USA 91:10747-10751; and U.S. Pat. Nos. 6,033,861, and 6,773,900.
[0180] Genetic variations introduced into microbes may be classified as transgenic, cisgenic, intragenomic, intrageneric, intergeneric, synthetic, evolved, rearranged, or SNPs.
[0181] Genetic variation may be introduced into numerous metabolic pathways within microbes to elicit improvements in the traits described above. Representative pathways include sulfur uptake pathways, glycogen biosynthesis, the glutamine regulation pathway, the molybdenum uptake pathway, the nitrogen fixation pathway, ammonia assimilation, ammonia excretion or secretion, nNitrogen uptake, glutamine biosynthesis, annamox, phosphate solubilization, organic acid transport, organic acid production, agglutinins production, reactive oxygen radical scavenging genes, Indole Acetic Acid biosynthesis, trehalose biosynthesis, plant cell wall degrading enzymes or pathways, root attachment genes, exopolysaccharide secretion, glutamate synthase pathway, iron uptake pathways, siderophore pathway, chitinase pathway, ACC deaminase, glutathione biosynthesis, phosphorous signaling genes, quorum quenching pathway, cytochrome pathways, hemoglobin pathway, bacterial hemoglobin-like pathway, small RNA rsmZ, rhizobitoxine biosynthesis, lapA adhesion protein, AHL quorum sensing pathway, phenazine biosynthesis, cyclic lipopeptide biosynthesis, and antibiotic production.
[0182] CRISPR/Cas9 (Clustered regularly interspaced short palindromic repeats)/CRISPR-associated (Cas) systems can be used to introduce desired mutations. CRISPR/Cas9 provide bacteria and archaea with adaptive immunity against viruses and plasmids by using CRISPR RNAs (crRNAs) to guide the silencing of invading nucleic acids. The Cas9 protein (or functional equivalent and/or variant thereof, i.e., Cas9-like protein) naturally contains DNA endonuclease activity that depends on the association of the protein with two naturally occurring or synthetic RNA molecules called crRNA and tracrRNA (also called guide RNAs). In some cases, the two molecules are covalently link to form a single molecule (also called a single guide RNA ("sgRNA"). Thus, the Cas9 or Cas9-like protein associates with a DNA-targeting RNA (which term encompasses both the two-molecule guide RNA configuration and the single-molecule guide RNA configuration), which activates the Cas9 or Cas9-like protein and guides the protein to a target nucleic acid sequence. If the Cas9 or Cas9-like protein retains its natural enzymatic function, it will cleave target DNA to create a double-stranded break, which can lead to genome alteration (i.e., editing: deletion, insertion (when a donor polynucleotide is present), replacement, etc.), thereby altering gene expression. Some variants of Cas9 (which variants are encompassed by the term Cas9-like) have been altered such that they have a decreased DNA cleaving activity (in some cases, they cleave a single strand instead of both strands of the target DNA, while in other cases, they have severely reduced to no DNA cleavage activity). Further exemplary descriptions of CRISPR systems for introducing genetic variation can be found in, e.g. U.S. Pat. No. 8,795,965.
[0183] As a cyclic amplification technique, polymerase chain reaction (PCR) mutagenesis uses mutagenic primers to introduce desired mutations. PCR is performed by cycles of denaturation, annealing, and extension. After amplification by PCR, selection of mutated DNA and removal of parental plasmid DNA can be accomplished by: 1) replacement of dCTP by hydroxymethylated-dCTP during PCR, followed by digestion with restriction enzymes to remove non-hydroxymethylated parent DNA only; 2) simultaneous mutagenesis of both an antibiotic resistance gene and the studied gene changing the plasmid to a different antibiotic resistance, the new antibiotic resistance facilitating the selection of the desired mutation thereafter; 3) after introducing a desired mutation, digestion of the parent methylated template DNA by restriction enzyme Dpnl which cleaves only methylated DNA, by which the mutagenized unmethylated chains are recovered; or 4) circularization of the mutated PCR products in an additional ligation reaction to increase the transformation efficiency of mutated DNA. Further description of exemplary methods can be found in e.g. U.S. Pat. No. 7,132,265, U.S. Pat. No. 6,713,285, U.S. Pat. No. 6,673,610, U.S. Pat. No. 6,391,548, U.S. Pat. No. 5,789,166, U.S. Pat. No. 5,780,270, U.S. Pat. No. 5,354,670, U.S. Pat. No. 5,071,743, and US20100267147.
[0184] Oligonucleotide-directed mutagenesis, also called site-directed mutagenesis, typically utilizes a synthetic DNA primer. This synthetic primer contains the desired mutation and is complementary to the template DNA around the mutation site so that it can hybridize with the DNA in the gene of interest. The mutation may be a single base change (a point mutation), multiple base changes, deletion, or insertion, or a combination of these. The single-strand primer is then extended using a DNA polymerase, which copies the rest of the gene. The gene thus copied contains the mutated site, and may then be introduced into a host cell as a vector and cloned. Finally, mutants can be selected by DNA sequencing to check that they contain the desired mutation.
[0185] Genetic variations can be introduced using error-prone PCR. In this technique the gene of interest is amplified using a DNA polymerase under conditions that are deficient in the fidelity of replication of sequence. The result is that the amplification products contain at least one error in the sequence. When a gene is amplified and the resulting product(s) of the reaction contain one or more alterations in sequence when compared to the template molecule, the resulting products are mutagenized as compared to the template. Another means of introducing random mutations is exposing cells to a chemical mutagen, such as nitrosoguanidine or ethyl methanesulfonate (Nestmann, Mutat Res 1975 June; 28(3):323-30), and the vector containing the gene is then isolated from the host.
[0186] Saturation mutagenesis is another form of random mutagenesis, in which one tries to generate all or nearly all possible mutations at a specific site, or narrow region of a gene. In a general sense, saturation mutagenesis is comprised of mutagenizing a complete set of mutagenic cassettes (wherein each cassette is, for example, 1-500 bases in length) in defined polynucleotide sequence to be mutagenized (wherein the sequence to be mutagenized is, for example, from 15 to 100, 000 bases in length). Therefore, a group of mutations (e.g. ranging from 1 to 100 mutations) is introduced into each cassette to be mutagenized. A grouping of mutations to be introduced into one cassette can be different or the same from a second grouping of mutations to be introduced into a second cassette during the application of one round of saturation mutagenesis. Such groupings are exemplified by deletions, additions, groupings of particular codons, and groupings of particular nucleotide cassettes.
[0187] Fragment shuffling mutagenesis, also called DNA shuffling, is a way to rapidly propagate beneficial mutations. In an example of a shuffling process, DNAse is used to fragment a set of parent genes into pieces of e.g. about 50-100 bp in length. This is then followed by a polymerase chain reaction (PCR) without primers--DNA fragments with sufficient overlapping homologous sequence will anneal to each other and are then be extended by DNA polymerase. Several rounds of this PCR extension are allowed to occur, after some of the DNA molecules reach the size of the parental genes. These genes can then be amplified with another PCR, this time with the addition of primers that are designed to complement the ends of the strands. The primers may have additional sequences added to their 5' ends, such as sequences for restriction enzyme recognition sites needed for ligation into a cloning vector. Further examples of shuffling techniques are provided in US20050266541.
[0188] Homologous recombination mutagenesis involves recombination between an exogenous DNA fragment and the targeted polynucleotide sequence. After a double-stranded break occurs, sections of DNA around the 5' ends of the break are cut away in a process called resection. In the strand invasion step that follows, an overhanging 3' end of the broken DNA molecule then "invades" a similar or identical DNA molecule that is not broken. The method can be used to delete a gene, remove exons, add a gene, and introduce point mutations. Homologous recombination mutagenesis can be permanent or conditional. Typically, a recombination template is also provided. A recombination template may be a component of another vector, contained in a separate vector, or provided as a separate polynucleotide. In some embodiments, a recombination template is designed to serve as a template in homologous recombination, such as within or near a target sequence nicked or cleaved by a site-specific nuclease. A template polynucleotide may be of any suitable length, such as about or more than about 10, 15, 20, 25, 50, 75, 100, 150, 200, 500, 1000, or more nucleotides in length. In some embodiments, the template polynucleotide is complementary to a portion of a polynucleotide comprising the target sequence. When optimally aligned, a template polynucleotide might overlap with one or more nucleotides of a target sequences (e.g. about or more than about 1, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 60, 70, 80, 90, 100 or more nucleotides). In some embodiments, when a template sequence and a polynucleotide comprising a target sequence are optimally aligned, the nearest nucleotide of the template polynucleotide is within about 1, 5, 10, 15, 20, 25, 50, 75, 100, 200, 300, 400, 500, 1000, 5000, 10000, or more nucleotides from the target sequence. Non-limiting examples of site-directed nucleases useful in methods of homologous recombination include zinc finger nucleases, CRISPR nucleases, TALE nucleases, and meganuclease. For a further description of the use of such nucleases, see e.g. U.S. Pat. No. 8,795,965 and US20140301990.
[0189] Mutagens that create primarily point mutations and short deletions, insertions, transversions, and/or transitions, including chemical mutagens or radiation, may be used to create genetic variations. Mutagens include, but are not limited to, ethyl methanesulfonate, methylmethane sulfonate, N-ethyl-N-nitrosurea, triethylmelamine, N-methyl-N-nitrosourea, procarbazine, chlorambucil, cyclophosphamide, diethyl sulfate, acrylamide monomer, melphalan, nitrogen mustard, vincristine, dimethylnitrosamine, N-methyl-N'-nitro-Nitrosoguanidine, nitrosoguanidine, 2-aminopurine, 7,12 dimethyl-benz(a)anthracene, ethylene oxide, hexamethylphosphoramide, bisulfan, diepoxyalkanes (diepoxyoctane, diepoxybutane, and the like), 2-methoxy-6-chloro-9[3-(ethyl-2-chloro-ethyl)aminopropylamino]acridine dihydrochloride and formaldehyde.
[0190] Introducing genetic variation may be an incomplete process, such that some bacteria in a treated population of bacteria carry a desired mutation while others do not. In some cases, it is desirable to apply a selection pressure so as to enrich for bacteria carrying a desired genetic variation. Traditionally, selection for successful genetic variants involved selection for or against some functionality imparted or abolished by the genetic variation, such as in the case of inserting antibiotic resistance gene or abolishing a metabolic activity capable of converting a non-lethal compound into a lethal metabolite. It is also possible to apply a selection pressure based on a polynucleotide sequence itself, such that only a desired genetic variation need be introduced (e.g. without also requiting a selectable marker). In this case, the selection pressure can comprise cleaving genomes lacking the genetic variation introduced to a target site, such that selection is effectively directed against the reference sequence into which the genetic variation is sought to be introduced. Typically, cleavage occurs within 100 nucleotides of the target site (e.g. within 75, 50, 25, 10, or fewer nucleotides from the target site, including cleavage at or within the target site). Cleaving may be directed by a site-specific nuclease selected from the group consisting of a Zinc Finger nuclease, a CRISPR nuclease, a TALE nuclease (TALEN), or a meganuclease. Such a process is similar to processes for enhancing homologous recombination at a target site, except that no template for homologous recombination is provided. As a result, bacteria lacking the desired genetic variation are more likely to undergo cleavage that, left unrepaired, results in cell death. Bacteria surviving selection may then be isolated for use in exposing to plants for assessing conferral of an improved trait.
[0191] A CRISPR nuclease may be used as the site-specific nuclease to direct cleavage to a target site. An improved selection of mutated microbes can be obtained by using Cas9 to kill non-mutated cells. Plants are then inoculated with the mutated microbes to re-confirm symbiosis and create evolutionary pressure to select for efficient symbionts. Microbes can then be re-isolated from plant tissues. CRISPR nuclease systems employed for selection against non-variants can employ similar elements to those described above with respect to introducing genetic variation, except that no template for homologous recombination is provided. Cleavage directed to the target site thus enhances death of affected cells.
[0192] Other options for specifically inducing cleavage at a target site are available, such as zinc finger nucleases, TALE nuclease (TALEN) systems, and meganuclease. Zinc-finger nucleases (ZFNs) are artificial DNA endonucleases generated by fusing a zinc finger DNA binding domain to a DNA cleavage domain. ZFNs can be engineered to target desired DNA sequences and this enables zinc-finger nucleases to cleave unique target sequences. When introduced into a cell, ZFNs can be used to edit target DNA in the cell (e.g., the cell's genome) by inducing double stranded breaks. Transcription activator-like effector nucleases (TALENs) are artificial DNA endonucleases generated by fusing a TAL (Transcription activator-like) effector DNA binding domain to a DNA cleavage domain. TALENS can be quickly engineered to bind practically any desired DNA sequence and when introduced into a cell, TALENs can be used to edit target DNA in the cell (e.g., the cell's genome) by inducing double strand breaks. Meganucleases (homing endonuclease) are endodeoxyribonucleases characterized by a large recognition site (double-stranded DNA sequences of 12 to 40 base pairs. Meganucleases can be used to replace, eliminate or modify sequences in a highly targeted way. By modifying their recognition sequence through protein engineering, the targeted sequence can be changed. Meganucleases can be used to modify all genome types, whether bacterial, plant or animal and are commonly grouped into four families: the LAGLIDADG family (SEQ ID NO: 1), the GIY-YIG family, the His-Cyst box family and the HNH family. Exemplary homing endonucleases include I-SceI, I-CeuI, PI-PspI, PI-Sce, I-SceIV, I-CsmI, I-PanI, I-SceII, I-PpoI, I-SceIII, I-CreI, I-TevI, I-TevII and I-TevIII.
Genetic Variation--Methods of Identification
[0193] The microbes of the present disclosure may be identified by one or more genetic modifications or alterations, which have been introduced into said microbe. One method by which said genetic modification or alteration can be identified is via reference to a SEQ ID NO that contains a portion of the microbe's genomic sequence that is sufficient to identify the genetic modification or alteration.
[0194] Further, in the case of microbes that have not had a genetic modification or alteration (e.g. a wild type, WT) introduced into their genomes, the disclosure can utilize 16S nucleic acid sequences to identify said microbes. A 16S nucleic acid sequence is an example of a "molecular marker" or "genetic marker," which refers to an indicator that is used in methods for visualizing differences in characteristics of nucleic acid sequences. Examples of other such indicators are restriction fragment length polymorphism (RFLP) markers, amplified fragment length polymorphism (AFLP) markers, single nucleotide polymorphisms (SNPs), insertion mutations, microsatellite markers (SSRs), sequence-characterized amplified regions (SCARs), cleaved amplified polymorphic sequence (CAPS) markers or isozyme markers or combinations of the markers described herein which defines a specific genetic and chromosomal location. Markers further include polynucleotide sequences encoding 16S or 18S rRNA, and internal transcribed spacer (ITS) sequences, which are sequences found between small-subunit and large-subunit rRNA genes that have proven to be especially useful in elucidating relationships or distinctions when compared against one another. Furthermore, the disclosure utilizes unique sequences found in genes of interest (e.g. nif H,D,K,L,A, glnE, amtB, etc.) to identify microbes disclosed herein.
[0195] The primary structure of major rRNA subunit 16S comprise a particular combination of conserved, variable, and hypervariable regions that evolve at different rates and enable the resolution of both very ancient lineages such as domains, and more modern lineages such as genera. The secondary structure of the 16S subunit include approximately 50 helices which result in base pairing of about 67% of the residues. These highly conserved secondary structural features are of great functional importance and can be used to ensure positional homology in multiple sequence alignments and phylogenetic analysis. Over the previous few decades, the 16S rRNA gene has become the most sequenced taxonomic marker and is the cornerstone for the current systematic classification of bacteria and archaea (Yarza et al. 2014. Nature Rev. Micro. 12:635-45).
[0196] Thus, in certain aspects, the disclosure provides for a sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any sequence in Tables E, F, G, or H.
[0197] Thus, in certain aspects, the disclosure provides for a microbe that comprises a sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 62-303. These sequences and their associated descriptions can be found in Tables F, G, and H.
[0198] In some aspects, the disclosure provides for a microbe that comprises a 16S nucleic acid sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, 277-283. These sequences and their associated descriptions can be found in Tables G and H.
[0199] In some aspects, the disclosure provides for a microbe that comprises a nucleic acid sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, 284-295. These sequences and their associated descriptions can be found in Tables G and H.
[0200] In some aspects, the disclosure provides for a microbe that comprises a nucleic acid sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 177-260, 296-303. These sequences and their associated descriptions can be found in Tables G and H.
[0201] In some aspects, the disclosure provides for a microbe that comprises, or primer that comprises, or probe that comprises, or non-native junction sequence that comprises, a nucleic acid sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 304-424. These sequences and their associated descriptions can be found in Table E.
[0202] In some aspects, the disclosure provides for a microbe that comprises a non-native junction sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 372-405. These sequences and their associated descriptions can be found in Table E.
[0203] In some aspects, the disclosure provides for a microbe that comprises an amino acid sequence, which shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 77, 78, 81, 82, or 83. These sequences and their associated descriptions can be found in Tables F and H.
Genetic Variation--Methods of Detection: Primers, Probes, and Assays
[0204] The present disclosure teaches primers, probes, and assays that are useful for detecting the microbes taught herein. In some aspects, the disclosure provides for methods of detecting the WT parental strains. In other aspects, the disclosure provides for methods of detecting the non-intergeneric engineered microbes derived from the WT strains. In aspects, the present disclosure provides methods of identifying non-intergeneric genetic alterations in a microbe.
[0205] In aspects, the genomic engineering methods of the present disclosure lead to the creation of non-natural nucleotide "junction" sequences in the derived non-intergeneric microbes. These non-naturally occurring nucleotide junctions can be used as a type of diagnostic that is indicative of the presence of a particular genetic alteration in a microbe taught herein.
[0206] The present techniques are able to detect these non-naturally occurring nucleotide junctions via the utilization of specialized quantitative PCR methods, including uniquely designed primers and probes. In some aspects, the probes of the disclosure bind to the non-naturally occurring nucleotide junction sequences. In some aspects, traditional PCR is utilized. In other aspects, real-time PCR is utilized. In some aspects, quantitative PCR (qPCR) is utilized.
[0207] Thus, the disclosure can cover the utilization of two common methods for the detection of PCR products in real-time: (1) non-specific fluorescent dyes that intercalate with any double-stranded DNA, and (2) sequence-specific DNA probes consisting of oligonucleotides that are labelled with a fluorescent reporter which permits detection only after hybridization of the probe with its complementary sequence. In some aspects, only the non-naturally occurring nucleotide junction will be amplified via the taught primers, and consequently can be detected via either a non-specific dye, or via the utilization of a specific hybridization probe. In other aspects, the primers of the disclosure are chosen such that the primers flank either side of a junction sequence, such that if an amplification reaction occurs, then said junction sequence is present.
[0208] Aspects of the disclosure involve non-naturally occurring nucleotide junction sequence molecules per se, along with other nucleotide molecules that are capable of binding to said non-naturally occurring nucleotide junction sequences under mild to stringent hybridization conditions. In some aspects, the nucleotide molecules that are capable of binding to said non-naturally occurring nucleotide junction sequences under mild to stringent hybridization conditions are termed "nucleotide probes."
[0209] In aspects, genomic DNA can be extracted from samples and used to quantify the presence of microbes of the disclosure by using qPCR. The primers utilized in the qPCR reaction can be primers designed by Primer Blast (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) to amplify unique regions of the wild-type genome or unique regions of the engineered non-intergeneric mutant strains. The qPCR reaction can be carried out using the SYBR GreenER qPCR SuperMix Universal (Thermo Fisher P/N 11762100) kit, using only forward and reverse amplification primers; alternatively, the Kapa Probe Force kit (Kapa Biosystems P/N KK4301) can be used with amplification primers and a TaqMan probe containing a RAM dye label at the 5' end, an internal ZEN quencher, and a minor groove binder and fluorescent quencher at the 3' end (Integrated DNA Technologies).
[0210] Certain primer, probe, and non-native junction sequences are listed in Table E. qPCR reaction efficiency can be measured using a standard curve generated from a known quantity of gDNA from the target genome. Data can be normalized to genome copies per g fresh weight using the tissue weight and extraction volume.
[0211] Quantitative polymerase chain reaction (qPCR) is a method of quantifying, in real time, the amplification of one or more nucleic acid sequences. The real time quantification of the PCR assay permits determination of the quantity of nucleic acids being generated by the PCR amplification steps by comparing the amplifying nucleic acids of interest and an appropriate control nucleic acid sequence, which may act as a calibration standard.
[0212] TaqMan probes are often utilized in qPCR assays that require an increased specificity for quantifying target nucleic acid sequences. TaqMan probes comprise a oligonucleotide probe with a fluorophore attached to the 5' end and a quencher attached to the 3' end of the probe. When the TaqMan probes remain as is with the 5' and 3' ends of the probe in close contact with each other, the quencher prevents fluorescent signal transmission from the fluorophore. TaqMan probes are designed to anneal within a nucleic acid region amplified by a specific set of primers. As the Taq polymerase extends the primer and synthesizes the nascent strand, the 5' to 3' exonuclease activity of the Taq polymerase degrades the probe that annealed to the template. This probe degradation releases the fluorophore, thus breaking the close proximity to the quencher and allowing fluorescence of the fluorophore. Fluorescence detected in the qPCR assay is directly proportional to the fluorophore released and the amount of DNA template present in the reaction.
[0213] The features of qPCR allow the practitioner to eliminate the labor-intensive post-amplification step of gel electrophoresis preparation, which is generally required for observation of the amplified products of traditional PCR assays. The benefits of qPCR over conventional PCR are considerable, and include increased speed, ease of use, reproducibility, and quantitative ability
Improvement of Traits
[0214] Methods of the present disclosure may be employed to introduce or improve one or more of a variety of desirable traits. Examples of traits that may introduced or improved include: root biomass, root length, height, shoot length, leaf number, water use efficiency, overall biomass, yield, fruit size, grain size, photosynthesis rate, tolerance to drought, heat tolerance, salt tolerance, resistance to nematode stress, resistance to a fungal pathogen, resistance to a bacterial pathogen, resistance to a viral pathogen, level of a metabolite, and proteome expression. The desirable traits, including height, overall biomass, root and/or shoot biomass, seed germination, seedling survival, photosynthetic efficiency, transpiration rate, seed/fruit number or mass, plant grain or fruit yield, leaf chlorophyll content, photosynthetic rate, root length, or any combination thereof, can be used to measure growth, and compared with the growth rate of reference agricultural plants (e.g., plants without the improved traits) grown under identical conditions.
[0215] A preferred trait to be introduced or improved is nitrogen fixation, as described herein. In some cases, a plant resulting from the methods described herein exhibits a difference in the trait that is at least about 5% greater, for example at least about 5%, at least about 8%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 75%, at least about 80%, at least about 80%, at least about 90%, or at least 100%, at least about 200%, at least about 300%, at least about 400% or greater than a reference agricultural plant grown under the same conditions in the soil. In additional examples, a plant resulting from the methods described herein exhibits a difference in the trait that is at least about 5% greater, for example at least about 5%, at least about 8%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 75%, at least about 80%, at least about 80%, at least about 90%, or at least 100%, at least about 200%, at least about 300%, at least about 400% or greater than a reference agricultural plant grown under similar conditions in the soil.
[0216] The trait to be improved may be assessed under conditions including the application of one or more biotic or abiotic stressors. Examples of stressors include abiotic stresses (such as heat stress, salt stress, drought stress, cold stress, and low nutrient stress) and biotic stresses (such as nematode stress, insect herbivory stress, fungal pathogen stress, bacterial pathogen stress, and viral pathogen stress).
[0217] The trait improved by methods and compositions of the present disclosure may be nitrogen fixation, including in a plant not previously capable of nitrogen fixation. In some cases, bacteria isolated according to a method described herein produce 1% or more (e.g. 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 15%, 20%, or more) of a plant's nitrogen, which may represent an increase in nitrogen fixation capability of at least 2-fold (e.g. 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 50-fold, 100-fold, 1000-fold, or more) as compared to bacteria isolated from the first plant before introducing any genetic variation. In some cases, the bacteria produce 5% or more of a plant's nitrogen. The desired level of nitrogen fixation may be achieved after repeating the steps of introducing genetic variation, exposure to a plurality of plants, and isolating bacteria from plants with an improved trait one or more times (e.g. 1, 2, 3, 4, 5, 10, 15, 25, or more times). In some cases, enhanced levels of nitrogen fixation are achieved in the presence of fertilizer supplemented with glutamine, ammonia, or other chemical source of nitrogen. Methods for assessing degree of nitrogen fixation are known, examples of which are described herein.
[0218] Microbe breeding is a method to systematically identify and improve the role of species within the crop microbiome. The method comprises three steps: 1) selection of candidate species by mapping plant-microbe interactions and predicting regulatory networks linked to a particular phenotype, 2) pragmatic and predictable improvement of microbial phenotypes through intra-species crossing of regulatory networks and gene clusters, and 3) screening and selection of new microbial genotypes that produce desired crop phenotypes. To systematically assess the improvement of strains, a model is created that links colonization dynamics of the microbial community to genetic activity by key species. The model is used to predict genetic targets breeding and improve the frequency of selecting improvements in microbiome-encoded traits of agronomic relevance.
Measuring Nitrogen Delivered in an Agriculturally Relevant Field Context
[0219] In the field, the amount of nitrogen delivered can be determined by the function of colonization multiplied by the activity.
Nitrogen delivered = .intg. Time & Space Colonization .times. Activity ##EQU00001##
[0220] The above equation requires (1) the average colonization per unit of plant tissue, and (2) the activity as either the amount of nitrogen fixed or the amount of ammonia excreted by each microbial cell. To convert to pounds of nitrogen per acre, corn growth physiology is tracked over time, e.g., size of the plant and associated root system throughout the maturity stages.
[0221] The pounds of nitrogen delivered to a crop per acre-season can be calculated by the following equation:
Nitrogen delivered=.intg.Plant Tissue(t).times.Colonization(t).times.Activity(t)dt
[0222] The Plant Tissue(t) is the fresh weight of corn plant tissue over the growing time (t). Values for reasonably making the calculation are described in detail in the publication entitled Roots, Growth and Nutrient Uptake (Mengel. Dept. of Agronomy Pub.# AGRY-95-08 (Rev. May-95. p. 1-8.).
[0223] The Colonization (t) is the amount of the microbes of interest found within the plant tissue, per gram fresh weight of plant tissue, at any particular time, t, during the growing season. In the instance of only a single timepoint available, the single timepoint is normalized as the peak colonization rate over the season, and the colonization rate of the remaining timepoints are adjusted accordingly.
[0224] Activity(t) is the rate of which N is fixed by the microbes of interest per unit time, at any particular time, t, during the growing season. In the embodiments disclosed herein, this activity rate is approximated by in vitro acetylene reduction assay (ARA) in ARA media in the presence of 5 mM glutamine or Ammonium excretion assay in ARA media in the presence of 5 mM ammonium ions.
[0225] The Nitrogen delivered amount is then calculated by numerically integrating the above function. In cases where the values of the variables described above are discretely measured at set timepoints, the values in between those timepoints are approximated by performing linear interpolation.
Nitrogen Fixation
[0226] Described herein are methods of increasing nitrogen fixation in a plant, comprising exposing the plant to bacteria comprising one or more genetic variations introduced into one or more genes regulating nitrogen fixation, wherein the bacteria produce 1% or more of nitrogen in the plant (e.g. 2%, 5%, 10%, or more), which may represent a nitrogen-fixation capability of at least 2-fold as compared to the plant in the absence of the bacteria. The bacteria may produce the nitrogen in the presence of fertilizer supplemented with glutamine, urea, nitrates or ammonia. Genetic variations can be any genetic variation described herein, including examples provided above, in any number and any combination. The genetic variation may be introduced into a gene selected from the group consisting of nifA, nifL, ntrB, ntrC, glutamine synthetase, glnA, glnB, glnK, draT, amtB, glutaminase, glnD, glnE, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifW, nifZ, nifM, nifF, nifB, and nifQ. The genetic variation may be a mutation that results in one or more of: increased expression or activity of nifA or glutaminase; decreased expression or activity of nifL, ntrB, glutamine synthetase, glnB, glnK, draT, amtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. The genetic variation introduced into one or more bacteria of the methods disclosed herein may be a knock-out mutation or it may abolish a regulatory sequence of a target gene, or it may comprise insertion of a heterologous regulatory sequence, for example, insertion of a regulatory sequence found within the genome of the same bacterial species or genus. The regulatory sequence can be chosen based on the expression level of a gene in a bacterial culture or within plant tissue. The genetic variation may be produced by chemical mutagenesis. The plants grown in step (c) may be exposed to biotic or abiotic stressors.
[0227] The amount of nitrogen fixation that occurs in the plants described herein may be measured in several ways, for example by an acetylene-reduction (AR) assay. An acetylene-reduction assay can be performed in vitro or in vivo. Evidence that a particular bacterium is providing fixed nitrogen to a plant can include: I) total plant N significantly increases upon inoculation, preferably with a concomitant increase in N concentration in the plant; 2) nitrogen deficiency symptoms are relieved under N-limiting conditions upon inoculation (which should include an increase in dry matter); 3) N.sub.2 fixation is documented through the use of an .sup.15N approach (which can be isotope dilution experiments, .sup.15N.sub.2 reduction assays, or .sup.15N natural abundance assays); 4) fixed N is incorporated into a plant protein or metabolite; and 5) all of these effects are not be seen in non-inoculated plants or in plants inoculated with a mutant of the inoculum strain.
[0228] The wild-type nitrogen fixation regulatory cascade can be represented as a digital logic circuit where the inputs O.sub.2 and NH.sub.4.sup.+ pass through a NOR gate, the output of which enters an AND gate in addition to ATP. In some embodiments, the methods disclosed herein disrupt the influence of NH.sub.4.sup.+ on this circuit, at multiple points in the regulatory cascade, so that microbes can produce nitrogen even in fertilized fields. However, the methods disclosed herein also envision altering the impact of ATP or O.sub.2 on the circuitry, or replacing the circuitry with other regulatory cascades in the cell, or altering genetic circuits other than nitrogen fixation. Gene clusters can be re-engineered to generate functional products under the control of a heterologous regulatory system. By eliminating native regulatory elements outside of, and within, coding sequences of gene clusters, and replacing them with alternative regulatory systems, the functional products of complex genetic operons and other gene clusters can be controlled and/or moved to heterologous cells, including cells of different species other than the species from which the native genes were derived. Once re-engineered, the synthetic gene clusters can be controlled by genetic circuits or other inducible regulatory systems, thereby controlling the products' expression as desired. The expression cassettes can be designed to act as logic gates, pulse generators, oscillators, switches, or memory devices. The controlling expression cassette can be linked to a promoter such that the expression cassette functions as an environmental sensor, such as an oxygen, temperature, touch, osmotic stress, membrane stress, or redox sensor.
[0229] As an example, the nifL, nifA, nifT, and nifX genes can be eliminated from the nif gene cluster. Synthetic genes can be designed by codon randomizing the DNA encoding each amino acid sequence. Codon selection is performed, specifying that codon usage be as divergent as possible from the codon usage in the native gene. Proposed sequences are scanned for any undesired features, such as restriction enzyme recognition sites, transposon recognition sites, repetitive sequences, sigma 54 and sigma 70 promoters, cryptic ribosome binding sites, and rho independent terminators. Synthetic ribosome binding sites are chosen to match the strength of each corresponding native ribosome binding site, such as by constructing a fluorescent reporter plasmid in which the 150 bp surrounding a gene's start codon (from -60 to +90) is fused to a fluorescent gene. This chimera can be expressed under control of the Ptac promoter, and fluorescence measured via flow cytometry. To generate synthetic ribosome binding sites, a library of reporter plasmids using 150 bp (-60 to +90) of a synthetic expression cassette is generated. Briefly, a synthetic expression cassette can consist of a random DNA spacer, a degenerate sequence encoding an RBS library, and the coding sequence for each synthetic gene. Multiple clones are screened to identify the synthetic ribosome binding site that best matched the native ribosome binding site. Synthetic operons that consist of the same genes as the native operons are thus constructed and tested for functional complementation. A further exemplary description of synthetic operons is provided in US20140329326.
Bacterial Species
[0230] Microbes useful in the methods and compositions disclosed herein may be obtained from any source. In some cases, microbes may be bacteria, archaea, protozoa or fungi. The microbes of this disclosure may be nitrogen fixing microbes, for example a nitrogen fixing bacteria, nitrogen fixing archaea, nitrogen fixing fungi, nitrogen fixing yeast, or nitrogen fixing protozoa. Microbes useful in the methods and compositions disclosed herein may be spore forming microbes, for example spore forming bacteria. In some cases, bacteria useful in the methods and compositions disclosed herein may be Gram positive bacteria or Gram negative bacteria. In some cases, the bacteria may be an endospore forming bacteria of the Firmicute phylum. In some cases, the bacteria may be a diazatroph. In some cases, the bacteria may not be a diazotroph.
[0231] The methods and compositions of this disclosure may be used with an archaea, such as, for example, Methanothermobacter thermoautotrophicus.
[0232] In some cases, bacteria which may be useful include, but are not limited to, Agrobacterium radiobacter, Bacillus acidocaldarius, Bacillus acidoterrestris, Bacillus agri, Bacillus aizawai, Bacillus albolactis, Bacillus alcalophilus, Bacillus alvei, Bacillus aminoglucosidicus, Bacillus aminovorans; Bacillus amyloyticus (also known as Paenibacillus amylolyticus) Bacillus amyloliquefaciens, Bacillus aneurinolyticus, Bacillus atrophaeus, Bacillus azotoformans, Bacillus badius, Bacillus cereus (synonyms: Bacillus endorhythmos, Bacillus medusa), Bacillus chitinosporus, Bacillus circulans, Bacillus coagulans, Bacillus endoparasiticus Bacillus fastidiosus, Bacillus firmus, Bacillus kurstaki, Bacillus lacticola, Bacillus lactimorbus, Bacillus lactis, Bacillus laterosporus (also known as Brevibacillus laterosporus), Bacillus lautus, Bacillus lentimorbus, Bacillus lentus, Bacillus licheniformis, Bacillus maroccanus, Bacillus megaterium, Bacillus metiens, Bacillus mycoides, Bacillus natio, Bacillus nematocida, Bacillus nigrificans, Bacillus nigrum, Bacillus pantothenticus, Bacillus popillae, Bacillus psychrosaccharolyticus, Bacillus pumilus, Bacillus sianensis, Bacillus smithii, Bacillus sphaericus, Bacillus subtilis, Bacillus thuringiensis, Bacillus uniflagellatus, Bradyrhizobium japonicum, Brevibacillus brevis Brevibacillus laterosporus (formerly Bacillus laterosporus), Chromobacterium subtsugae, Delftia acidovorans, Lactobacillus acidophilus, Lysobacter antibioticus, Lysobacter enzymogenes, Paenibacillus alvei, Paenibacillus polymyxa, Paenibacillus popilliae (formerly Bacillus popilliae), Pantoea agglomerans, Pasteuria penetrans (formerly Bacillus penetrates), Pasteuria usgae, Pectobacterium carotovorum (formerly Erwinia carotovora), Pseudomonas aeruginosa, Pseudomonas aureofaciens, Pseudomonas cepacia (formerly known as Burkholderia cepacia), Pseudomonas chlororaphis, Pseudomonas fluorescens, Pseudomonas proradix, Pseudomonas putida, Pseudomonas syringae, Serratia entomophila, Serratia marcescens, Streptomyces colombiensis, Streptomyces galbus, Streptomyces goshikiensis, Streptomyces griseoviridis, Streptomyces lavendulae, Streptomyces prasinus, Streptomyces saraceticus, Streptomyces venezuelae, Xanthomonas campestris, Xenorhabdus luminescens, Xenorhabdus nematophila, Rhodococcus globerulus AQ719 (NRRL Accession No. B-21663), Bacillus sp. AQ175 (ATCC Accession No. 55608), Bacillus sp. AQ 177 (ATCC Accession No. 55609), Bacillus sp. AQ178 (ATCC Accession NO. 53522), and Streptomyces sp. strain NRRL Accession No. 13-30145. In some cases the bacterium may be Azotobacter chroococcum, Methanosarcina barker, Klesiella pneumoniae, Azotobacter vinelandii, Rhodobacter spharoides, Rhodobacter capsulatus, Rhodobcter palustris, Rhodosporillum rubrum, Rhizobium leguminosarum or Rhizobium etli.
[0233] In some cases the bacterium may be a species of Clostridium, for example Clostridium pasteurianum, Clostridium beijerinckii, Clostridium perfringens, Clostridium tetani, Clostridium acetobutylicum.
[0234] In some cases, bacteria used with the methods and compositions of the present disclosure may be cyanobacteria. Examples of cyanobacterial genuses include Anabaena (for example Anagaena sp. PCC7120), Nostoc (for example Nostoc punctiforme), or Synechocystis (for example Synechocystis sp. PCC6803).
[0235] In some cases, bacteria used with the methods and compositions of the present disclosure may belong to the phylum Chlorobi, for example Chlorobium tepidum.
[0236] In some cases, microbes used with the methods and compositions of the present disclosure may comprise a gene homologous to a known NifH gene. Sequences of known NifH genes may be found in, for example, the Zehr lab NifH database, (https://wwwzehr.pmc.aese.edu/nifH_Database_Public/, Apr. 4, 2014), or the Buckley lab NifH database (http://www.css.cornell.edu/faculty/buckley/nifh.htm, and Gaby, John Christian, and Daniel H. Buckley. "A comprehensive aligned nifH gene database: a multipurpose tool for studies of nitrogen-fixing bacteria." Database 2014 (2014): bau001.). In some cases, microbes used with the methods and compositions of the present disclosure may comprise a sequence which encodes a polypeptide with at least 60%, 70%, 80%, 85%, 90%, 95%, 96%, 96%, 98%, 99% or more than 99% sequence identity to a sequence from the Zehr lab NifH database, (https://wwwzehr.pmc.ucsc.edu/nifH_Database_Public/, Apr. 4, 2014). In some cases, microbes used with the methods and compositions of the present disclosure may comprise a sequence which encodes a polypeptide with at least 60%, 70%, 80%, 85%, 90%, 95%, 96%, 96%, 98%, 99% or more than 99% sequence identity to a sequence from the Buckley lab NifH database, (Gaby, John Christian, and Daniel H. Buckley. "A comprehensive aligned nifH gene database: a multipurpose tool for studies of nitrogen-fixing bacteria." Database 2014 (2014): bau001.).
[0237] Microbes useful in the methods and compositions disclosed herein can be obtained by extracting microbes from surfaces or tissues of native plants; grinding seeds to isolate microbes; planting seeds in diverse soil samples and recovering microbes from tissues; or inoculating plants with exogenous microbes and determining which microbes appear in plant tissues. Non-limiting examples of plant tissues include a seed, seedling, leaf, cutting, plant, bulb or tuber. In some cases, bacteria are isolated from a seed. The parameters for processing samples may be varied to isolate different types of associative microbes, such as rhizospheric, epiphytes, or endophytes. Bacteria may also be sourced from a repository, such as environmental strain collections, instead of initially isolating from a first plant. The microbes can be genotyped and phenotyped, via sequencing the genomes of isolated microbes; profiling the composition of communities in planta; characterizing the transcriptomic functionality of communities or isolated microbes; or screening microbial features using selective or phenotypic media (e.g., nitrogen fixation or phosphate solubilization phenotypes). Selected candidate strains or populations can be obtained via sequence data; phenotype data; plant data (e.g., genome, phenotype, and/or yield data); soil data (e.g., pH, N/P/K content, and/or bulk soil biotic communities); or any combination of these.
[0238] The bacteria and methods of producing bacteria described herein may apply to bacteria able to self-propagate efficiently on the leaf surface, root surface, or inside plant tissues without inducing a damaging plant defense reaction, or bacteria that are resistant to plant defense responses. The bacteria described herein may be isolated by culturing a plant tissue extract or leaf surface wash in a medium with no added nitrogen. However, the bacteria may be unculturable, that is, not known to be culturable or difficult to culture using standard methods known in the art. The bacteria described herein may be an endophyte or an epiphyte or a bacterium inhabiting the plant rhizosphere (rhizospheric bacteria). The bacteria obtained after repeating the steps of introducing genetic variation, exposure to a plurality of plants, and isolating bacteria from plants with an improved trait one or more times (e.g. 1, 2, 3, 4, 5, 10, 15, 25, or more times) may be endophytic, epiphytic, or rhizospheric. Endophytes are organisms that enter the interior of plants without causing disease symptoms or eliciting the formation of symbiotic structures, and are of agronomic interest because they can enhance plant growth and improve the nutrition of plants (e.g., through nitrogen fixation). The bacteria can be a seed-borne endophyte. Seed-borne endophytes include bacteria associated with or derived from the seed of a grass or plant, such as a seed-borne bacterial endophyte found in mature, dry, undamaged (e.g., no cracks, visible fungal infection, or prematurely germinated) seeds. The seed-borne bacterial endophyte can be associated with or derived from the surface of the seed; alternatively, or in addition, it can be associated with or derived from the interior seed compartment (e.g., of a surface-sterilized seed). In some cases, a seed-borne bacterial endophyte is capable of replicating within the plant tissue, for example, the interior of the seed. Also, in some cases, the seed-borne bacterial endophyte is capable of surviving desiccation.
[0239] The bacterial isolated according to methods of the disclosure, or used in methods or compositions of the disclosure, can comprise a plurality of different bacterial taxa in combination. By way of example, the bacteria may include Proteobacteria (such as Pseudomonas, Enterobacter, Stenotrophomonas, Burkholderia, Rhizobium, Herbaspirillum, Pantoea, Serratia, Rahnella, Azospirillum, Azorhizobium, Azotobacter, Duganella, Delftia, Bradyrhizobiun, Sinorhizobium and Halomonas), Firmicutes (such as Bacillus, Paenibacillus, Lactobacillus, Mycoplasma, and Acetabacterium), and Actinobacteria (such as Streptomyces, Rhodacoccus, Microbacterium, and Curtobacterium). The bacteria used in methods and compositions of this disclosure may include nitrogen fixing bacterial consortia of two or more species. In some cases, one or more bacterial species of the bacterial consortia may be capable of fixing nitrogen. In some cases, one or more species of the bacterial consortia may facilitate or enhance the ability of other bacteria to fix nitrogen. The bacteria which fix nitrogen and the bacteria which enhance the ability of other bacteria to fix nitrogen may be the same or different. In some examples, a bacterial strain may be able to fix nitrogen when in combination with a different bacterial strain, or in a certain bacterial consortia, but may be unable to fix nitrogen in a monoculture. Examples of bacterial genuses which may be found in a nitrogen fixing bacterial consortia include, but are not limited to, Herbaspirillum, Azospirillum, Enterobacter, and Bacillus.
[0240] Bacteria that can be produced by the methods disclosed herein include Azotobacter sp., Bradyrhizobium sp., Klebsiella sp., and Sinorhizobium sp. In some cases, the bacteria may be selected from the group consisting of: Azotobacter vinelandii, Bradyrhizobium japonicum, Klebsiella pneumoniae, and Sinorhizobium meliloti. In some cases, the bacteria may be of the genus Enterobacter or Rahnella. In some cases, the bacteria may be of the genus Frankia, or Clostridium. Examples of bacteria of the genus Clostridium include, but are not limited to, Clostridium acetobutilicum, Clostridium pasteurianum, Clostridium beijerinckii, Clostridium perfringens, and Clostridium tetani. In some cases, the bacteria may be of the genus Paenibacillus, for example Paenibacillus azotofixans, Paenibacillus borealis, Paenibacillus durus, Paenibacillus macerans, Paenibacillus polymyxa, Paenibacillus alvei, Paenibacillus amylolyticus, Paenibacillus campinasensis, Paenibacillus chibensis, Paenibacillus glucanolyticus, Paenibacillus illinoisensis, Paenibacillus larvae subsp. Larvae, Paenibacillus larvae subsp. Pulvifaciens, Paenibacillus lautus, Paenibacillus macerans, Paenibacillus macquariensis, Paenibacillus macquariensis, Paenibacillus pabuli, Paenibacillus peoriae, or Paenibacillus polymyxa.
[0241] In some examples, bacteria isolated according to methods of the disclosure can be a member of one or more of the following taxa: Achronobacter, Acidithiobacillus, Acidovorax, Acidovoraz, Acinetobacter, Actinoplanes, Adlercreutzia, Aerococcus, Aeromonas, Afipia, Agromyces, Ancylobacter, Arthrobacter, Atopostipes, Azospirillum, Bacillus, Bdellovibrio, Beijerinckia, Bosea, Bradyrhizobium, Brevibacillus, Brevundimonas, Burkholderia, Candidatus Haloredivivus, Caulobacter, Cellulomonas, Cellvibrio, Chryseobacterium, Citrobacter, Clostridium, Coraliomargarita, Corynebacterium, Cupriavidus, Curtobacterium, Curvibacter, Deinococcus, Delftia, Desemzia, Devosia, Dokdonella, Dyella, Enhydrobacter, Enterobacter, Enterococcus, Erwinia, Escherichia, Escherichia/Shigella, Exiguobacterium, Ferroglobus, Filimonas, Finegoldia, Flavisolibacter, Flavobacterium, Frigoribacterium, Gluconacetobacter, Hafnia, Halobaculum, Halomonas, Halosimplex, Herbaspirillum, Hymenobacter, Klebsiella, Kocuria, Kosakonia, Lactobacillus, Leclercia, Lentzea, Luteibacter, Luteimonas, Massilia, Mesorhizobium, Methylobacterium, Microbacterium, Micrococcus, Microvirga, Mycobacterium, Neisseria, Nocardia, Oceanibaculum, Ochrobactrum, Okibacterium, Oligotropha, Oryzihumus, Oxalophagus, Paenibacillus, Panteoa, Pantoea, Pelomonas, Perlucidibaca, Plantibacter, Polynucleobacter, Propionibacterium, Propioniciclava, Pseudoclavibacter, Pseudomonas, Pseudonocardia, Pseudoxanthomonas, Psychrobacter, Rahnella, Ralstonia, Rheinheimera, Rhizobium, Rhodococcus, Rhodopseudomonas, Roseateles, Ruminococcus, Sebaldella, Sediminibacillus, Sediminibacterium, Serratia, Shigella, Shigella, Sinorhizobium, Sinosporangium, Sphingobacterium, Sphingomonas, Sphingopyxis, Sphingosinicella, Staphylococcus, 25 Stenotrophomonas, Strenotrophomonas, Streptococcus, Streptomyces, Stygiolobus, Sulfurisphaera, Tatumella, Tepidimonas, Thermomonas, Thiobacillus, Variovorax, WPS-2 genera incertae sedis, Xanthomonas, and Zimmermanella.
[0242] In some cases, a bacterial species selected from at least one of the following genera are utilized: Enterobacter, Klebsiella, Kosakonia, and Rahnella. In some cases, a combination of bacterial species from the following genera are utilized: Enterobacter, Klebsiella, Kosakonia, and Rahnella. In some cases, the species utilized can be one or more of: Enterobacter sacchari, Klebsiella variicola, Kosakonia sacchari, and Rahnella aquatilis.
[0243] In some cases, a Gram positive microbe may have a Molybdenum-Iron nitrogenase system comprising: nifH, nifD, nifK, nifB, nifE, nifN, nifX, hesA, nifV, nifU, nifW, nifU, nifS, nifl1, and nifl2. In some cases, a Gram positive microbe may have a vanadium nitrogenase system comprising: vnfDG, vnfK, vnfE, vnfN, vupC, vupB, vupA, vnfV, vnfR1, vnfH, vnfR2, vnfA, (transcriptional regulator). In some cases, a Gram positive microbe may have an iron-only nitrogenase system comprising: anfK, anfG, anfD, anfH, anfA (transcriptional regulator). In some cases a Gram positive microbe may have a nitrogenase system comprising glnB, and glnK (nitrogen signaling proteins). Some examples of enzymes involved in nitrogen metabolism in Gram positive microbes include glnA (glutamine synthetase), gdh (glutamate dehydrogenase), bdh (3-hydroxybutyrate dehydrogenase), glutaminase, gltAB/gltB/gltS (glutamate synthase), asnA/asnB (aspartate-ammonia ligase/asparagine synthetase), and ansA/ansZ (asparaginase). Some examples of proteins involved in nitrogen transport in Gram positive microbes include amtB (ammonium transporter), glnK (regulator of ammonium transport), glnPHQ/glnQHMP (ATP-dependent glutamine/glutamate transporters), glnT/alsT/yrbD/yflA (glutamine-like proton symport transporters), and gltP/gltT/yhcl/nqt (glutamate-like proton symport transporters).
[0244] Examples of Gram positive microbes which may be of particular interest include Paenibacillus polymixa, Paenibacillus riograndensis, Paenibacillus sp., Frankia sp., Heliobacterium sp., Heliobacterium chlorum, Heliobacillus sp., Heliophilum sp., Heliorestis sp., Clostridium acetobutylicum, Clostridium sp., Mycobacterium flaum, Mycobacterium sp., Arthrobacter sp., Agromyces sp., Corynebacterium autitrophicum, Corynebacterium sp., Micromonspora sp., Propionibacteria sp., Streptomyces sp., and Microbacterium sp.
[0245] Some examples of genetic alterations which may be make in Gram positive microbes include: deleting glnR to remove negative regulation of BNF in the presence of environmental nitrogen, inserting different promoters directly upstream of the nif cluster to eliminate regulation by GlnR in response to environmental nitrogen, mutating glnA to reduce the rate of ammonium assimilation by the GS-GOGAT pathway, deleting amtB to reduce uptake of ammonium from the media, mutating glnA so it is constitutively in the feedback-inhibited (FBI-GS) state, to reduce ammonium assimilation by the GS-GOGAT pathway.
[0246] In some cases, glnR is the main regulator of N metabolism and fixation in Paenibacillus species. In some cases, the genome of a Paenibacillus species may not contain a gene to produce glnR. In some cases, the genome of a Paenibacillus species may not contain a gene to produce glnE or glnD. In some cases, the genome of a Paenibacillus species may contain a gene to produce glnB or glnK. For example Paenibacillus sp. WLY78 doesn't contain a gene for glnB, or its homologs found in the archaeon Methanococcus maripaludis, nifI1 and nifI2. In some cases, the genomes of Paenibacillus species may be variable. For example, Paenibacillus polymixa E681 lacks glnK and gdh, has several nitrogen compound transporters, but only amtB appears to be controlled by GlnR. In another example, Paenibacillus sp. JDR2 has glnK, gdh and most other central nitrogen metabolism genes, has many fewer nitrogen compound transporters, but does have glnPHQ controlled by GlnR. Paenibacillus riograndensis SBR5 contains a standard glnRA operon, an fdx gene, a main nil operon, a secondary nif operon, and an anf operon (encoding iron-only nitrogenase). Putative glnR/tnrA sites were found upstream of each of these operons. GlnR may regulate all of the above operons, except the anf operon. GlnR may bind to each of these regulatory sequences as a dimer.
[0247] Paenibacillus N-fixing strains may fall into two subgroups: Subgroup I, which contains only a minimal nif gene cluster and subgroup II, which contains a minimal cluster, plus an uncharacterized gene between nifX and hesA, and often other clusters duplicating some of the nif genes, such as nifH, nifHDK, nifBEN, or clusters encoding vanadaium nitrogenase (vnf) or iron-only nitrogenase (anf) genes.
[0248] In some cases, the genome of a Paenibacillus species may not contain a gene to produce g/n/3 or &K. In some cases, the genome of a Paenibacillus species may contain a minimal nif cluster with 9 genes transcribed from a sigma-70 promoter. In some cases a Paenibacillus nif cluster may be negatively regulated by nitrogen or oxygen. In some cases, the genome of a Paenibacillus species may not contain a gene to produce sigma-54. For example, Paenibacillus sp. WLY78 does not contain a gene for sigma-54. In some cases, a nif cluster may be regulated by glnR, and/or TnrA. In some cases, activity of a nif cluster may be altered by altering activity of glnR, and/or TnrA.
[0249] In Bacilli, glutamine synthetase (GS) is feedback-inhibited by high concentrations of intracellular glutamine, causing a shift in confirmation (referred to as FBI-GS). Nif clusters contain distinct binding sites for the regulators GlnR and TnrA in several Bacilli species. GlnR binds and represses gene expression in the presence of excess intracellular glutamine and AMP. A role of GlnR may be to prevent the influx and intracellular production of glutamine and ammonium under conditions of high nitrogen availability. TnrA may bind and/or activate (or repress) gene expression in the presence of limiting intracellular glutamine, and/or in the presence of FBI-GS. In some cases the activity of a Bacilli nif cluster may be altered by altering the activity of GlnR.
[0250] Feedback-inhibited glutamine synthetase (FBI-GS) may bind GlnR and stabilize binding of GlnR to recognition sequences. Several bacterial species have a GlnR/TnrA binding site upstream of the nif cluster. Altering the binding of FBI-GS and GlnR may alter the activity of the nif pathway.
Sources of Microbes
[0251] The bacteria (or any microbe according to the disclosure) may be obtained from any general terrestrial environment, including its soils, plants, fungi, animals (including invertebrates) and other biota, including the sediments, water and biota of lakes and rivers; from the marine environment, its biota and sediments (for example, sea water, marine muds, marine plants, marine invertebrates (for example, sponges), marine vertebrates (for example, fish)); the terrestrial and marine geosphere (regolith and rock, for example, crushed subterranean rocks, sand and clays); the cryosphere and its meltwater; the atmosphere (for example, filtered aerial dusts, cloud and rain droplets); urban, industrial and other man-made environments (for example, accumulated organic and mineral matter on concrete, roadside gutters, roof surfaces, and road surfaces).
[0252] The plants from which the bacteria (or any microbe according to the disclosure) are obtained may be a plant having one or more desirable traits, for example a plant which naturally grows in a particular environment or under certain conditions of interest. By way of example, a certain plant may naturally grow in sandy soil or sand of high salinity, or under extreme temperatures, or with little water, or it may be resistant to certain pests or disease present in the environment, and it may be desirable for a commercial crop to be grown in such conditions, particularly if they are, for example, the only conditions available in a particular geographic location. By way of further example, the bacteria may be collected from commercial crops gown in such environments, or more specifically from individual crop plants best displaying a trait of interest amongst a crop grown in any specific environment: for example the fastest-growing plants amongst a crop grown in saline-limiting soils, or the least damaged plants in crops exposed to severe insect damage or disease epidemic, or plants having desired quantities of certain metabolites and other compounds, including fiber content, oil content, and the like, or plants displaying desirable colors, taste or smell. The bacteria may be collected from a plant of interest or any material occurring in the environment of interest, including fungi and other animal and plant biota, soil, water, sediments, and other elements of the environment as referred to previously.
[0253] The bacteria (or any microbe according to the disclosure) may be isolated from plant tissue. This isolation can occur from any appropriate tissue in the plant, including for example root, stem and leaves, and plant reproductive tissues. By way of example, conventional methods for isolation from plants typically include the sterile excision of the plant material of interest (e.g. root or stem lengths, leaves), surface sterilization with an appropriate solution (e.g. 2% sodium hypochlorite), after which the plant material is placed on nutrient medium for microbial growth. Alternatively, the surface-sterilized plant material can be crushed in a sterile liquid (usually water) and the liquid suspension, including small pieces of the crushed plant material spread over the surface of a suitable solid agar medium, or media, which may or may not be selective (e.g. contain only phytic acid as a source of phosphorus). This approach is especially useful for bacteria which form isolated colonies and can be picked off individually to separate plates of nutrient medium, and further purified to a single species by well-known methods. Alternatively, the plant root or foliage samples may not be surface sterilized but only washed gently thus including surface-dwelling epiphytic microorganisms in the isolation process, or the epiphytic microbes can be isolated separately, by imprinting and lifting off pieces of plant roots, stem or leaves onto the surface of an agar medium and then isolating individual colonies as above. This approach is especially useful for bacteria, for example. Alternatively, the roots may be processed without washing off small quantities of soil attached to the roots, thus including microbes that colonize the plant rhizosphere. Otherwise, soil adhering to the roots can be removed, diluted and spread out onto agar of suitable selective and non-selective media to isolate individual colonies of rhizospheric bacteria.
Budapest Treaty on the International Recognition of the Deposit of Microorganisms for the Purpose of Patent Procedures
[0254] The microbial deposits of the present disclosure were made under the provisions of the Budapest Treaty on the International Recognition of the Deposit of Microorganisms for the Purpose of Patent Procedure (Budapest Treaty).
[0255] Applicants state that pursuant to 37 C.F.R. .sctn. 1.808(a)(2) "all restrictions imposed by the depositor on the availability to the public of the deposited material will be irrevocably removed upon the granting of the patent." This statement is subject to paragraph (b) of this section (i.e. 37 C.F.R. .sctn. 1.808(b)).
[0256] Biologically pure cultures of Rahnella aquatilis and Enterobacter sacchari were deposited on Jul. 14, 2015 with the American Type Culture Collection (ATCC; an International Depositary Authority), 10801 University Blvd., Manassas, Va. 20110, USA, and assigned ATTC Patent Deposit Designation numbers PTA-122293 and PTA-122294, respectively. The applicable deposit information is found below in Table A.
[0257] The Enterobacter sacchari has now been reclassified as Kosakonia sacchari, the name for the organism may be used interchangeably throughout the manuscript.
[0258] Many microbes of the present disclosure are derived from two wild-type strains, as depicted in FIG. 18 and FIG. 19. Strain CI006 is a bacterial species previously classified in the genus Enterobacter (see aforementioned reclassification into Kosakonia), and FIG. 19 identities the lineage of the mutants that have been derived from CI006. Strain CI019 is a bacterial species classified in the genus Rahnella, and FIG. 19 identifies the lineage of the mutants that have been derived from CI019. With regard to FIG. 18 and FIG. 19, it is noted that strains comprising CM in the name are mutants of the strains depicted immediately to the left of said CM strain. The deposit information for the CI006 Kosakonia wild type (WT) and CI019 Rahnella WT are found in the below Table A.
[0259] Some microorganisms described in this application were deposited on Jan. 6, 2017 or Aug. 11, 2017 with the Bigelow National Center for Marine Algae and Microbiota (NCMA), located at 60 Bigelow Drive, East Boothbay, Me. 04544, USA. As aforementioned, all deposits were made under the terms of the Budapest Treaty on the International Recognition of the Deposit of Microorganisms for the Purposes of Patent Procedure. The Bigelow National Center for Marine Algae and Microbiota accession numbers and dates of deposit for the aforementioned Budapest Treaty deposits are provided in Table A.
[0260] Biologically pure cultures of Kosakonia sacchari (WT), Rahnella aquatilis (WT), and a variant Kosakonia sacchari strain were deposited on Jan. 6, 2017 with the Bigelow National Center for Marine Algae and Microbiota (NCMA), located at 60 Bigelow Drive, East Boothbay, Me. 04544, USA, and assigned NCMA Patent Deposit Designation numbers 201701001, 201701003, and 201701002, respectively. The applicable deposit information is found below in Table A.
[0261] Biologically pure cultures of variant Kosakonia sacchari strains were deposited on Aug. 11, 2017 with the Bigelow National Center for Marine Algae and Microbiota (NCMA), located at 60 Bigelow Drive, East Boothbay, Me. 04544, USA, and assigned NCMA Patent Deposit Designation numbers 201708004, 201708003, and 201708002, respectively. The applicable deposit information is found below in Table A.
[0262] A biologically pure culture of Klebsiella variicola (WV was deposited on Aug. 11, 2017 with the Bigelow National Center for Marine Algae and Microbiota (NCMA), located at 60 Bigelow Drive, East Boothbay, Me. 04544, USA, and assigned NCMA Patent Deposit Designation number 201708001. Biologically pure cultures of two Klebsiella variicola variants were deposited on Dec. 20, 2017 with the Bigelow National Center for Marine Algae and Microbiota (NCMA), located at 60 Bigelow Drive, East Boothbay, Me. 04544, USA, and assigned NCMA Patent Deposit Designation numbers 201712001 and 201712002, respectively. The applicable deposit information is found below in Table A.
TABLE-US-00001 TABLE A Microorganisms Deposited under the Budapest Treaty Pivot Strain Designation (some strains Depos- have multiple Accession Date of itory designations) Taxonomy Number Deposit ATCC Rahnella aquatilis PTA-122293 Jul. 14, 2015 ATCC Enterobacter PTA-122294 Jul. 14, sacchari 2015 (taxonomically re- classified after de- posit as Kosakonia sacchari) NCMA CI006, Kosakonia sacchari 201701001 Jan. 6, PBC6.1, 6 (WT) 2017 NCMA CI019, 19 Rahnella aquatilis 201701003 Jan. 6, (WT) 2017 NCMA CM029, Kosakonia sacchari 201701002 Jan. 6, 6-412 2017 NCMA 6-403 Kosakonia sacchari 201708004 Aug. 11, CM037 2017 NCMA 6-404, Kosakonia sacchari 201708003 Aug. 11, CM38, 2017 PBC6.38 NCMA CM094, Kosakonia sacchari 201708002 Aug. 11, 6-881, 2017 PBC6.94 NCMA CI137, 137, Klebsiella variicola 201708001 Aug. 11, PB137 (WT) 2017 NCMA 137-1034 Klebsiella variicola 201712001 Dec. 20, 2017 NCMA 137-1036 Klebsiella variicola 201712002 Dec. 20, 2017
Isolated and Biologically Pure Microorganisms
[0263] The present disclosure, in certain embodiments, provides isolated and biologically pure microorganisms that have applications, infer alia, in agriculture. The disclosed microorganisms can be utilized in their isolated and biologically pure states, as well as being formulated into compositions (see below section for exemplary composition descriptions). Furthermore, the disclosure provides microbial compositions containing at least two members of the disclosed isolated and biologically pure microorganisms, as well as methods of utilizing said microbial compositions. Furthermore, the disclosure provides for methods of modulating nitrogen fixation in plants via the utilization of the disclosed isolated and biologically pure microbes.
[0264] In some aspects, the isolated and biologically pure microorganisms of the disclosure are those from Table A. In other aspects, the isolated and biologically pure microorganisms of the disclosure are derived from a microorganism of Table A. For example, a strain, child, mutant, or derivative, of a microorganism from Table A are provided herein. The disclosure contemplates all possible combinations of microbes listed in Table A, said combinations sometimes forming a microbial consortia. The microbes from Table A, either individually or in any combination, can be combined with any plant, active (synthetic, organic, etc.), adjuvant, carrier, supplement, or biological, mentioned in the disclosure.
Compositions
[0265] Compositions comprising bacteria or bacterial populations produced according to methods described herein and/or having characteristics as described herein can be in the form of a liquid, a foam, or a dry product. Compositions comprising bacteria or bacterial populations produced according to methods described herein and/or having characteristics as described herein may also be used to improve plant traits. In some examples, a composition comprising bacterial populations may be in the form of a dry powder, a slurry of powder and water, or a flowable seed treatment. The compositions comprising bacterial populations may be coated on a surface of a seed, and may be in liquid form.
[0266] The composition can be fabricated in bioreactors such as continuous stirred tank reactors, batch reactors, and on the farm. In some examples, compositions can be stored in a container, such as a jug or in mini bulk. In some examples, compositions may be stored within an object selected from the group consisting of a bottle, jar, ampule, package, vessel, bag, box, bin, envelope, carton, container, silo, shipping container, truck bed, and/or case.
[0267] Compositions may also be used to improve plant traits. In some examples, one or more compositions may be coated onto a seed. In some examples, one or more compositions may be coated onto a seedling. In some examples, one or more compositions may be coated onto a surface of a seed. In some examples, one or more compositions may be coated as a layer above a surface of a seed. In some examples, a composition that is coated onto a seed may be in liquid form, in dry product form, in foam form, in a form of a slurry of powder and water, or in a flowable seed treatment. In some examples, one or more compositions may be applied to a seed and/or seedling by spraying, immersing, coating, encapsulating, and/or dusting the seed and/or seedling with the one or more compositions. In some examples, multiple bacteria or bacterial populations can be coated onto a seed and/or a seedling of the plant. In some examples, at least two, at least three, at least four, at least five, at least six, at least seven, at least eight, at least nine, at least ten, or more than ten bacteria of a bacterial combination can be selected from one of the following genera: Acidovorax, Agrobacterium, Bacillus, Burkholderia, Chryseobacterium, Curtobacterium, Enterobacter, Escherichia, Methylobacterium, Paenibacillus, Pantoea, Pseudomonas, Ralstonia, Saccharibacillus, Sphingomonas, and Stenotrophomonas.
[0268] In some examples, at least two, at least three, at least four, at least five, at least six, at least seven, at least eight, at least nine, at least ten, or more than ten bacteria and bacterial populations of an endophytic combination are selected from one of the following families: Bacillaceae, Burkholderiaceae, Comamonadaceae, Enterobacteriaceae, Flavobacteriaceae, Methylobacteriaceae, Microbacteriaceae, Paenibacillileae, Pseudomonnaceae, Rhizobiaceae, Sphingomonadaceae, Xanthomonadaceae, Cladosporiaceae, Gnomoniaceae, Incertae sedis, Lasiosphaeriaceae, Netriaceae, and Pleosporaceae.
[0269] In some examples, at least two, at least three, at least four, at least five, at least six, at least seven, at least eight, at least night, at least ten, or more than ten bacteria and bacterial populations of an endophytic combination are selected from one of the following families: Bacillaceae, Burkholderiaceae, Comamonadaceae, Enterobacteriaceae, Flavobacteriaceae, Methylobacteriaceae, Microbacteriaceae, Paenibacillileae, Pseudomonnaceae, Rhizobiaceae, Sphingomonadaceae, Xanthomonadaceae, Cladosporiaceae, Gnomoniaceae, Incertae sedis, Lasiosphaeriaceae, Netriaceae, Pleosporaceae.
[0270] Examples of compositions may include seed coatings for commercially important agricultural crops, for example, sorghum, canola, tomato, strawberry, barley, rice, maize, and wheat. Examples of compositions can also include seed coatings for corn, soybean, canola, sorghum, potato, rice, vegetables, cereals, and oilseeds. Seeds as provided herein can be genetically modified organisms (GMO), non-GMO, organic, or conventional. In some examples, compositions may be sprayed on the plant aerial parts, or applied to the roots by inserting into furrows in which the plant seeds are planted, watering to the soil, or dipping the roots in a suspension of the composition. In some examples, compositions may be dehydrated in a suitable manner that maintains cell viability and the ability to artificially inoculate and colonize host plants. The bacterial species may be present in compositions at a concentration of between 10.sup.8 to 10.sup.10 CFU/ml. In some examples, compositions may be supplemented with trace metal ions, such as molybdenum ions, iron ions, manganese ions, or combinations of these ions. The concentration of ions in examples of compositions as described herein may between about 0.1 mM and about 50 mM. Some examples of compositions may also be formulated with a carrier, such as beta-glucan, carboxylmethyl cellulose (CMC), bacterial extracellular polymeric substance (EPS), sugar, animal milk, or other suitable carriers. In some examples, peat or planting materials can be used as a carrier, or biopolymers in which a composition is entrapped in the biopolymer can be used as a carrier. The compositions comprising the bacterial populations described herein can improve plant traits, such as promoting plant growth, maintaining high chlorophyll content in leaves, increasing fruit or seed numbers, and increasing fruit or seed unit weight.
[0271] The compositions comprising the bacterial populations described herein may be coated onto the surface of a seed. As such, compositions comprising a seed coated with one or more bacteria described herein are also contemplated. The seed coating can be formed by mixing the bacterial population with a porous, chemically inert granular carrier. Alternatively, the compositions may be inserted directly into the furrows into which the seed is planted or sprayed onto the plant leaves or applied by dipping the roots into a suspension of the composition. An effective amount of the composition can be used to populate the sub-soil region adjacent to the roots of the plant with viable bacterial growth, or populate the leaves of the plant with viable bacterial growth. In general, an effective amount is an amount sufficient to result in plants with improved traits (e.g. a desired level of nitrogen fixation).
[0272] Bacterial compositions described herein can be formulated using an agriculturally acceptable carrier. The formulation useful for these embodiments may include at least one member selected from the group consisting of a tackifier, a microbial stabilizer, a fungicide, an antibacterial agent, a preservative, a stabilizer, a surfactant, an anti-complex agent, an herbicide, a nematicide, an insecticide, a plant growth regulator, a fertilizer, a rodenticide, a dessicant, a bactericide, a nutrient, or any combination thereof. In some examples, compositions may be shelf-stable. For example, any of the compositions described herein can include an agriculturally acceptable carrier (e.g., one or more of a fertilizer such as a non-naturally occurring fertilizer, an adhesion agent such as a non-naturally occurring adhesion agent, and a pesticide such as a non-naturally occurring pesticide). A non-naturally occurring adhesion agent can be, for example, a polymer, copolymer, or synthetic wax. For example, any of the coated seeds, seedlings, or plants described herein can contain such an agriculturally acceptable carrier in the seed coating. In any of the compositions or methods described herein, an agriculturally acceptable carrier can be or can include a non-naturally occurring compound (e.g., a non-naturally occurring fertilizer, a non-naturally occurring adhesion agent such as a polymer, copolymer, or synthetic wax, or a non-naturally occurring pesticide). Non-limiting examples of agriculturally acceptable carriers are described below. Additional examples of agriculturally acceptable carriers are known in the art.
[0273] In some cases, bacteria are mixed with an agriculturally acceptable carrier. The carrier can be a solid carrier or liquid carrier, and in various forms including microspheres, powders, emulsions and the like. The carrier may be any one or more of a number of carriers that confer a variety of properties, such as increased stability, wettability, or dispersability. Wetting agents such as natural or synthetic surfactants, which can be nonionic or ionic surfactants, or a combination thereof can be included in the composition. Water-in-oil emulsions can also be used to formulate a composition that includes the isolated bacteria (see, for example, U.S. Pat. No. 7,485,451). Suitable formulations that may be prepared include wettable powders, granules, gels, agar strips or pellets, thickeners, and the like, microencapsulated particles, and the like, liquids such as aqueous flowables, aqueous suspensions, water-in-oil emulsions, etc. The formulation may include grain or legume products, for example, ground grain or beans, broth or flour derived from grain or beans, starch, sugar, or oil.
[0274] In some embodiments, the agricultural carrier may be soil or a plant growth medium. Other agricultural carriers that may be used include water, fertilizers, plant-based oils, humectants, or combinations thereof. Alternatively, the agricultural carrier may be a solid, such as diatomaceous earth, loam, silica, alginate, clay, bentonite, vermiculite, seed cases, other plant and animal products, or combinations, including granules, pellets, or suspensions. Mixtures of any of the aforementioned ingredients are also contemplated as carriers, such as but not limited to, pesta (flour and kaolin clay), agar or flour-based pellets in loam, sand, or clay, etc. Formulations may include food sources for the bacteria, such as barley, rice, or other biological materials such as seed, plant parts, sugar cane bagasse, hulls or stalks from grain processing, ground plant material or wood from building site refuse, sawdust or small fibers from recycling of paper, fabric, or wood.
[0275] For example, a fertilizer can be used to help promote the growth or provide nutrients to a seed, seedling, or plant. Non-limiting examples of fertilizers include nitrogen, phosphorous, potassium, calcium, sulfur, magnesium, boron, chloride, manganese, iron, zinc, copper, molybdenum, and selenium (or a salt thereof). Additional examples of fertilizers include one or more amino acids, salts, carbohydrates, vitamins, glucose, NaCl, yeast extract, NH.sub.4H.sub.2PO.sub.4, (NH.sub.4).sub.2SO.sub.4, glycerol, valine, L-leucine, lactic acid, propionic acid, succinic acid, malic acid, citric acid, KH tartrate, xylose, lyxose, and lecithin. In one embodiment, the formulation can include a tackifier or adherent (referred to as an adhesive agent) to help bind other active agents to a substance (e.g., a surface of a seed). Such agents are useful for combining bacteria with carriers that can contain other compounds (e.g., control agents that are not biologic), to yield a coating composition. Such compositions help create coatings around the plant or seed to maintain contact between the microbe and other agents with the plant or plant part. In one embodiment, adhesives are selected from the group consisting of: alginate, gums, starches, lecithins, formononetin, polyvinyl alcohol, alkali formononetinate, hesperetin, polyvinyl acetate, cephalins, Gum Arabic, Xanthan Gum, Mineral Oil, Polyethylene Glycol (PEG), Polyvinyl pyrrolidone (PVP), Arabino-galactan, Methyl Cellulose, PEG 400, Chitosan, Polyacrylamide, Polyacrylate, Polyacrylonitrile, Glycerol, Triethylene glycol, Vinyl Acetate, Gellan Gum, Polystyrene, Polyvinyl, Carboxymethyl cellulose, Gum Ghatti, and polyoxyethylene-polyoxybutylene block copolymers.
[0276] In some embodiments, the adhesives can be, e.g. a wax such as carnauba wax, beeswax, Chinese wax, shellac wax, spermaceti wax, candelilla wax, castor wax, ouricury wax, and rice bran wax, a polysaccharide (e.g., starch, dextrins, maltodextrins, alginate, and chitosans), a fat, oil, a protein (e.g., gelatin and zeins), gum arables, and shellacs. Adhesive agents can be non-naturally occurring compounds, e.g., polymers, copolymers, and waxes. For example, non-limiting examples of polymers that can be used as an adhesive agent include: polyvinyl acetates, polyvinyl acetate copolymers, ethylene vinyl acetate (EVA) copolymers, polyvinyl alcohols, polyvinyl alcohol copolymers, celluloses (e.g., ethylcelluloses, methylcelluloses, hydroxymethylcelluloses, hydroxypropylcelluloses, and carboxymethylcelluloses), polyvinylpyrolidones, vinyl chloride, vinylidene chloride copolymers, calcium lignosulfonates, acrylic copolymers, polyvinylacrylates, polyethylene oxide, acylamide polymers and copolymers, polyhydroxyethyl acrylate, methyl acrylamide monomers, and polychloroprene.
[0277] In some examples, one or more of the adhesion agents, anti-fungal agents, growth regulation agents, and pesticides (e.g., insecticide) are non-naturally occurring compounds (e.g., in any combination). Additional examples of agriculturally acceptable carriers include dispersants (e.g., polyvinylpyrrolidone/vinyl acetate PVPIVA S-630), surfactants, binders, and filler agents.
[0278] The formulation can also contain a surfactant. Non-limiting examples of surfactants include nitrogen-surfactant blends such as Prefer 28 (Cenex), Surf-N(US), Inhance (Brandt), P-28 (Wilfann) and Patrol (Helena); esterified seed oils include Sun-It II (AmCy), MSO (UAP), Scoil (Agsco), Hasten (Wilfarm) and Mes-100 (Drexel); and organo-silicone surfactants include Silwet L77 (UAP), Silikin (Terra), Dyne-Amit (Helena), Kinetic (Helena), Sylgard 309 (Wilbur-Ellis) and Century (Precision). In one embodiment, the surfactant is present at a concentration of between 0.01% v/v to 10% v/v. In another embodiment, the surfactant is present at a concentration of between 0.1% v/v to 1% v/v.
[0279] In certain cases, the formulation includes a microbial stabilizer. Such an agent can include a desiccant, which can include any compound or mixture of compounds that can be classified as a desiccant regardless of whether the compound or compounds are used in such concentrations that they in fact have a desiccating effect on a liquid inoculant. Such desiccants are ideally compatible with the bacterial population used, and should promote the ability of the microbial population to survive application on the seeds and to survive desiccation. Examples of suitable desiccants include one or more of trehalose, sucrose, glycerol, and Methylene glycol. Other suitable desiccants include, but are not limited to, non reducing sugars and sugar alcohols (e.g., mannitol or sorbitol). The amount of desiccant introduced into the formulation can range from about 5% to about 50% by weight/volume, for example, between about 10% to about 40%, between about 15% to about 35%, or between about 20% to about 30%. In some cases, it is advantageous for the formulation to contain agents such as a fungicide, an antibacterial agent, an herbicide, a nematicide, an insecticide, a plant growth regulator, a rodenticide, bactericide, or a nutrient. In some examples, agents may include protectants that provide protection against seed surface-borne pathogens. In some examples, protectants may provide some level of control of soil-borne pathogens. In some examples, protectants may be effective predominantly on a seed surface.
[0280] In some examples, a fungicide may include a compound or agent, whether chemical or biological, that can inhibit the growth of a fungus or kill a fungus. In some examples, a fungicide may include compounds that may be fungistatic or fungicidal. In some examples, fungicide can be a protectant, or agents that are effective predominantly on the seed surface, providing protection against seed surface-borne pathogens and providing some level of control of soil-borne pathogens. Non-limiting examples of protectant fungicides include captan, maneb, thiram, or fludioxonil.
[0281] In some examples, fungicide can be a systemic fungicide, which can be absorbed into the emerging seedling and inhibit or kill the fungus inside host plant tissues. Systemic fungicides used for seed treatment include, but are not limited to the following: azoxystrobin, carboxin, mefenoxam, metalaxyl, thiabendazole, trifloxystrobin, and various triazole fungicides, including difenoconazole, ipconazole, tebuconazole, and triticonazole. Mefenoxam and metalaxyl are primarily used to target the water mold fungi Pythium and Phytophthora. Some fungicides are preferred over others, depending on the plant species, either because of subtle differences in sensitivity of the pathogenic fungal species, or because of the differences in the fungicide distribution or sensitivity of the plants. In some examples, fungicide can be a biological control agent, such as a bacterium or fungus. Such organisms may be parasitic to the pathogenic fungi, or secrete toxins or other substances which can kill or otherwise prevent the growth of fungi. Any type of fungicide, particularly ones that are commonly used on plants, can be used as a control agent in a seed composition.
[0282] In some examples, the seed coating composition comprises a control agent which has antibacterial properties. In one embodiment, the control agent with antibacterial properties is selected from the compounds described herein elsewhere. In another embodiment, the compound is Streptomycin, oxytetracycline, oxolinic acid, or gentamicin. Other examples of antibacterial compounds which can be used as part of a seed coating composition include those based on dichlorophene and benzylalcohol hemi formal (Proxel.RTM. from ICI or Acticide.RTM. RS from Thor Chemie and Kathon.RTM. MK 25 from Rohm & Haas) and isothiazolinone derivatives such as alkylisothiazolinones and benzisothiazolinones (Acticide.RTM. NIBS from Thor Chemie).
[0283] In some examples, growth regulator is selected from the group consisting of: Abscisic acid, amidochlor, ancymidol, 6-benzylaminopurine, brassinolide, butralin, chlormequat (chlormequat chloride), choline chloride, cyclanilide, daminozide, dikegulac, dimethipin, 2,6-dimethylpuridine, ethephon, flumetralin, flurprimidol, fluthiacet, forchlorfenuron, gibberellic acid, inabenfide, indole-3-acetic acid, maleic hydrazide, mefluidide, mepiquat (mepiquat chloride), naphthaleneacetic acid, N-6-benzyladenine, paclobutrazol, prohexadione phosphorotrithioate, 2,3,5-tri-iodobenzoic acid, trinexapac-ethyl and uniconazole. Additional non-limiting examples of growth regulators include brassinosteroids, cytokinines (e.g., kinetin and zeatin), auxins (e.g., indolylacetic acid and indolylacetyl aspartate), flavonoids and isoflavanoids (e.g., formononetin and diosmetin), phytoaixins (e.g., glyceolline), and phytoalexin-inducing oligosaccharides (e.g., pectin, chitin, chitosan, polygalacuronic acid, and oligogalacturonic acid), and gibellerins. Such agents are ideally compatible with the agricultural seed or seedling onto which the formulation is applied (e.g., it should not be deleterious to the growth or health of the plant). Furthermore, the agent is ideally one which does not cause safety concerns for human, animal or industrial use (e.g., no safety issues, or the compound is sufficiently labile that the commodity plant product derived from the plant contains negligible amounts of the compound).
[0284] Some examples of nematode-antagonistic biocontrol agents include ARF18; 30 Arthrobotrys spp.; Chaetomium spp.; Cylindrocarpon spp.; Exophilia spp.; Fusarium spp.; Gliocladium spp.; Hirsutella spp.; Lecanicillium spp.; Monacrosporium spp.; Myrothecium spp.; Neocosmospora spp.; Paecilomyces spp.; Pochonia spp.; Stagonospora spp.; vesicular-arbuscular mycorrhizal fungi, Burkholderia spp.; Pasteuria spp., Brevibacillus spp.; Pseudomonas spp.; and Rhizobacteria. Particularly preferred nematode-antagonistic biocontrol agents include ARF18, Arthrobotrys oligospora, Arthrobotrys dactyloides. Chaetomium globosum, Cylindrocarpon heteronema, Exophilia jeanselmei, Exophilia pisciphila, Fusarium aspergilus, Fusarium solani, Gliocladium catenulatum, Gliocladium roseum, Gliocladium vixens, Hirsutella rhossiliensis, Hirsutella minnesotensis, Lecanicillium lecanii, Monacrosporium drechsleri, Monacrosporium gephyropagum, Myrotehcium verrucaria, Neocosmospora vasinfecta, Paecilomyces lilacinus, Pochonia chlamydosporia, Stagonospora heteroderae, Stagonospora phaseoli, vesicular-arbuscular mycorrhizal fungi, Burkholderia cepacia, Pasteuria penetrans, Pasteuria thornei, Pasteuria nishizawae, Pasteuria ramosa, Pastrueia usage, Brevibacillus laterosporus strain G4, Pseudomonas fluorescens and Rhizobacteria.
[0285] Some examples of nutrients can be selected from the group consisting of a nitrogen fertilizer including, but not limited to Urea, Ammonium nitrate, Ammonium sulfate, Non-pressure nitrogen solutions, Aqua ammonia, Anhydrous ammonia, Ammonium thiosulfate, Sulfur-coated urea, Urea-formaldehydes, IBDU, Polymer-coated urea, Calcium nitrate, Ureaform, and Methylene urea, phosphorous fertilizers such as Diammonium phosphate, Monoammonium phosphate, Ammonium polyphosphate, Concentrated superphosphate and Triple superphosphate, and potassium fertilizers such as Potassium chloride, Potassium sulfate, Potassium-magnesium sulfate, Potassium nitrate. Such compositions can exist as free salts or ions within the seed coat composition. Alternatively, nutrients/fertilizers can be complexed or chelated to provide sustained release over time.
[0286] Some examples of rodenticides may include selected from the group of substances consisting of 2-isovalerylindan-1,3-dione, 4-(quinoxalin-2-ylamino) benzenesulfonamide, alpha-chlorohydrin, aluminum phosphide, antu, arsenous oxide, barium carbonate, bisthiosemi, brodifacoum, bromadiolone, bromethalin, calcium cyanide, chloralose, chlorophacinone, cholecalciferol, coumachlor, coumafuryl, coumatetralyl, crimidine, difenacoum, difethialone, diphacinone, ergocalciferol, flocoumafen, fluoroacetamide, flupropadine, flupropadine hydrochloride, hydrogen cyanide, iodomethane, lindane, magnesium phosphide, methyl bromide, norbormide, phosacetim, phosphine, phosphorus, pindone, potassium arsenite, pyrinuron, scilliroside, sodium arsenite, sodium cyanide, sodium fluoroacetate, strychnine, thallium sulfate, warfarin and zinc phosphide.
[0287] In the liquid form, for example, solutions or suspensions, bacterial populations can be mixed or suspended in water or in aqueous solutions. Suitable liquid diluents or carriers include water, aqueous solutions, petroleum distillates, or other liquid carriers.
[0288] Solid compositions can be prepared by dispersing the bacterial populations in and on an appropriately divided solid carrier, such as peat, wheat, bran, vermiculite, clay, talc, bentonite, diatomaceous earth, fuller's earth, pasteurized soil, and the like. When such formulations are used as wettable powders, biologically compatible dispersing agents such as non-ionic, anionic, amphoteric, or cationic dispersing and emulsifying agents can be used.
[0289] The solid carriers used upon formulation include, for example, mineral carriers such as kaolin clay, pyrophyllite, bentonite, montmorillonite, diatomaceous earth, acid white soil, vermiculite, and pearlite, and inorganic salts such as ammonium sulfate, ammonium phosphate, ammonium nitrate, urea, ammonium chloride, and calcium carbonate. Also, organic fine powders such as wheat flour, wheat bran, and rice bran may be used. The liquid carriers include vegetable oils such as soybean oil and cottonseed oil, glycerol, ethylene glycol, polyethylene glycol, propylene glycol, polypropylene glycol, etc.
Application of Bacterial Populations on Crops
[0290] The composition of the bacteria or bacterial population described herein can be applied in furrow, in talc, or as seed treatment. The composition can be applied to a seed package in bulk, mini bulk, in a bag, or in talc.
[0291] The planter can plant the treated seed and grows the crop according to conventional ways, twin row, or ways that do not require tilling. The seeds can be distributed using a control hopper or an individual hopper. Seeds can also be distributed using pressurized air or manually. Seed placement can be performed using variable rate technologies. Additionally, application of the bacteria or bacterial population described herein may be applied using variable rate technologies. In some examples, the bacteria can be applied to seeds of corn, soybean, canola, sorghum, potato, rice, vegetables, cereals, pseudocereals, and oilseeds. Examples of cereals may include barley, Fonio, oats, palmer's grass, rye, pearl millet, sorghum, spelt, teff, triticale, and wheat. Examples of pseudocereals may include breadnut, buckwheat, cattail, chia, flax, grain amaranth, hanza, quinoa, and sesame. In some examples, seeds can be genetically modified organisms (GMO), non-GMO, organic or conventional.
[0292] Additives such as micro-fertilizer, PGR, herbicide, insecticide, and fungicide can be used additionally to treat the crops. Examples of additives include crop protectants such as insecticides, nematicides, fungicide, enhancement agents such as colorants, polymers, pelleting, priming, and disinfectants, and other agents such as inoculant, PGR, softener, and micronutrients. PGRs can be natural or synthetic plant hormones that affect root growth, flowering, or stem elongation. PGRs can include auxins, gibberellins, cytokinins, ethylene, and abscisic acid (ABA).
[0293] The composition can be applied in furrow in combination with liquid fertilizer. In some examples, the liquid fertilizer may be held in tanks. NPK fertilizers contain macronutrients of sodium, phosphorous, and potassium.
[0294] The composition may improve plant traits, such as promoting plant growth, maintaining high chlorophyll content in leaves, increasing fruit or seed numbers, and increasing fruit or seed unit weight. Methods of the present disclosure may be employed to introduce or improve one or more of a variety of desirable traits. Examples of traits that may introduced or improved include: root biomass, root length, height, shoot length, leaf number, water use efficiency, overall biomass, yield, fruit size, grain size, photosynthesis rate, tolerance to drought, heat tolerance, salt tolerance, tolerance to low nitrogen stress, nitrogen use efficiency, resistance to nematode stress, resistance to a fungal pathogen, resistance to a bacterial pathogen, resistance to a viral pathogen, level of a metabolite, modulation in level of a metabolite, proteome expression. The desirable traits, including height, overall biomass, root and/or shoot biomass, seed germination, seedling survival, photosynthetic efficiency, transpiration rate, seed/fruit number or mass, plant grain or fruit yield, leaf chlorophyll content, photosynthetic rate, root length, or any combination thereof, can be used to measure growth, and compared with the growth rate of reference agricultural plants (e.g., plants without the introduced and/or improved traits) grown under identical conditions. In some examples, the desirable traits, including height, overall biomass, root and/or shoot biomass, seed germination, seedling survival, photosynthetic efficiency, transpiration rate, seed/fruit number or mass, plant grain or fruit yield, leaf chlorophyll content, photosynthetic rate, root length, or any combination thereof, can be used to measure growth, and compared with the growth rate of reference agricultural plants (e.g., plants without the introduced and/or improved traits) grown under similar conditions.
[0295] An agronomic trait to a host plant may include, but is not limited to, the following: altered oil content, altered protein content, altered seed carbohydrate composition, altered seed oil composition, and altered seed protein composition, chemical tolerance, cold tolerance, delayed senescence, disease resistance, drought tolerance, ear weight, growth improvement, health enhancement, heat tolerance, herbicide tolerance, herbivore resistance improved nitrogen fixation, improved nitrogen utilization, improved root architecture, improved water use efficiency, increased biomass, increased root length, increased seed weight, increased shoot length, increased yield, increased yield under water-limited conditions, kernel mass, kernel moisture content, metal tolerance, number of ears, number of kernels per ear, number of pods, nutrition enhancement, pathogen resistance, pest resistance, photosynthetic capability improvement, salinity tolerance, stay-green, vigor improvement, increased dry weight of mature seeds, increased fresh weight of mature seeds, increased number of mature seeds per plant, increased chlorophyll content, increased number of pods per plant, increased length of pods per plant, reduced number of wilted leaves per plant, reduced number of severely wilted leaves per plant, and increased number of non-wilted leaves per plant, a detectable modulation in the level of a metabolite, a detectable modulation in the level of a transcript, and a detectable modulation in the proteome, compared to an isoline plant grown from a seed without said seed treatment formulation.
[0296] In some cases, plants are inoculated with bacteria or bacterial populations that are isolated from the same species of plant as the plant element of the inoculated plant. For example, an bacteria or bacterial population that is normally found in one variety of Zea mays (corn) is associated with a plant element of a plant of another variety of Zea mays that in its natural state lacks said bacteria and bacterial populations. In one embodiment, the bacteria and bacterial populations is derived from a plant of a related species of plant as the plant element of the inoculated plant. For example, an bacteria and bacterial populations that is normally found in Zea diploperennis Iltis et al., (diploperennial teosinte) is applied to a Zea mays (corn), or vice versa. In some cases, plants are inoculated with bacteria and bacterial populations that are heterologous to the plant element of the inoculated plant. In one embodiment, the bacteria and bacterial populations is derived from a plant of another species. For example, an bacteria and bacterial populations that is normally found in dicots is applied to a monocot plant (e.g., inoculating corn with a soybean-derived bacteria and bacterial populations), or vice versa. In other cases, the bacteria and bacterial populations to be inoculated onto a plant is derived from a related species of the plant that is being inoculated. In one embodiment, the bacteria and bacterial populations is derived from a related taxon, for example, from a related species. The plant of another species can be an agricultural plant. In another embodiment, the bacteria and bacterial populations is part of a designed composition inoculated into any host plant element.
[0297] In some examples, the bacteria or bacterial population is exogenous wherein the bacteria and bacterial population is isolated from a different plant than the inoculated plant. For example, in one embodiment, the bacteria or bacterial population can be isolated from a different plant of the same species as the inoculated plant. In some cases, the bacteria or bacterial population can be isolated from a species related to the inoculated plant.
[0298] In some examples, the bacteria and bacterial populations described herein are capable of moving from one tissue type to another. For example, the present invention's detection and isolation of bacteria and bacterial populations within the mature tissues of plants after coating on the exterior of a seed demonstrates their ability to move from seed exterior into the vegetative tissues of a maturing plant. Therefore, in one embodiment, the population of bacteria and bacterial populations is capable of moving from the seed exterior into the vegetative tissues of a plant. In one embodiment, the bacteria and bacterial populations that is coated onto the seed of a plant is capable, upon germination of the seed into a vegetative state, of localizing to a different tissue of the plant. For example, bacteria and bacterial populations can be capable of localizing to any one of the tissues in the plant, including: the root, adventitious root, seminal 5 root, root hair, shoot, leaf, flower, bud, tassel, meristem, pollen, pistil, ovaries, stamen, fruit, stolon, rhizome, nodule, tuber, trichome, guard cells, hydathode, petal, sepal, glume, rachis, vascular cambium, phloem, and xylem. In one embodiment, the bacteria and bacterial populations is capable of localizing to the root and/or the root hair of the plant. In another embodiment, the bacteria and bacterial populations is capable of localizing to the photosynthetic tissues, for example, leaves and shoots of the plant. In other cases, the bacteria and bacterial populations is localized to the vascular tissues of the plant, for example, in the xylem and phloem. In still another embodiment, the bacteria and bacterial populations is capable of localizing to the reproductive tissues (flower, pollen, pistil, ovaries, stamen, fruit) of the plant. In another embodiment, the bacteria and bacterial populations is capable of localizing to the root, shoots, leaves and reproductive tissues of the plant. In still another embodiment, the bacteria and bacterial populations colonizes a fruit or seed tissue of the plant. In still another embodiment, the bacteria and bacterial populations is able to colonize the plant such that it is present in the surface of the plant (i.e., its presence is detectably present on the plant exterior, or the episphere of the plant). In still other embodiments, the bacteria and bacterial populations is capable of localizing to substantially all, or all, tissues of the plant. In certain embodiments, the bacteria and bacterial populations is not localized to the root of a plant. In other cases, the bacteria and bacterial populations is not localized to the photosynthetic tissues of the plant.
[0299] The effectiveness of the compositions can also be assessed by measuring the relative maturity of the crop or the crop heating unit (CHU). For example, the bacterial population can be applied to corn, and corn growth can be assessed according to the relative maturity of the corn kernel or the time at which the corn kernel is at maximum weight. The crop heating unit (CHU) can also be used to predict the maturation of the corn crop. The CHU determines the amount of heat accumulation by measuring the daily maximum temperatures on crop growth.
[0300] In examples, bacterial may localize to any one of the tissues in the plant, including: the root, adventitious root, seminal root, root hair, shoot, leaf, flower, bud tassel, meristem, pollen, pistil, ovaries, stamen, fruit, stolon, rhizome, nodule, tuber, trichome, guard cells, hydathode, petal, sepal, glume, rachis, vascular cambium, phloem, and xylem. In another embodiment, the bacteria or bacterial population is capable of localizing to the photosynthetic tissues, for example, leaves and shoots of the plant. In other cases, the bacteria and bacterial populations is localized to the vascular tissues of the plant, for example, in the xylem and phloem. In another embodiment, the bacteria or bacterial population is capable of localizing to reproductive tissues (flower, pollen, pistil, ovaries, stamen, or fruit) of the plant. In another embodiment, the bacteria and bacterial populations is capable of localizing to the root, shoots, leaves and reproductive tissues of the plant. In another embodiment, the bacteria or bacterial population colonizes a fruit or seed tissue of the plant. In still another embodiment, the bacteria or bacterial population is able to colonize the plant such that it is present in the surface of the plant. In another embodiment, the bacteria or bacterial population is capable of localizing to substantially all, or all, tissues of the plant. In certain embodiments, the bacteria or bacterial population is not localized to the root of a plant. In other cases, the bacteria and bacterial populations is not localized to the photosynthetic tissues of the plant.
[0301] The effectiveness of the bacterial compositions applied to crops can be assessed by measuring various features of crop growth including, but not limited to, planting rate, seeding vigor, root strength, drought tolerance, plant height, dry down, and test weight.
Plant Species
[0302] The methods and bacteria described herein are suitable for any of a variety of plants, such as plants in the genera Hordeum, Oryza, Zea, and Triticeae. Other non-limiting examples of suitable plants include mosses, lichens, and algae. In some cases, the plants have economic, social and/or environmental value, such as food crops, fiber crops, oil crops, plants in the forestry or pulp and paper industries, feedstock for biofuel production and/or ornamental plants. In some examples, plants may be used to produce economically valuable products such as a grain, a flour, a starch, a syrup, a meal, an oil, a film, a packaging, a nutraceutical product, a pulp, an animal feed, a fish fodder, a bulk material for industrial chemicals, a cereal product, a processed human-food product, a sugar, an alcohol, and/or a protein. Non-limiting examples of crop plants include maize, rice, wheat, barley, sorghum, millet, oats, rye triticale, buckwheat, sweet corn, sugar cane, onions, tomatoes, strawberries, and asparagus.
[0303] In some examples, plants that may be obtained or improved using the methods and composition disclosed herein may include plants that are important or interesting for agriculture, horticulture, biomass for the production of biofuel molecules and other chemicals, and/or forestry. Some examples of these plants may include pineapple, banana, coconut, lily, grasspeas and grass; and dicotyledonous plants, such as, for example, peas, alfalfa, tomatillo, melon, chickpea, chicory, clover, kale, lentil, soybean, tobacco, potato, sweet potato, radish, cabbage, rape, apple trees, grape, cotton, sunflower, Chale cress, canola, citrus (including orange, mandarin, kumquat, lemon, lime, grapefruit, tangerine, tangelo, citron, and pomelo), pepper, bean, lettuce, Panicum virgatum (switch), Sorghum bicolor (sorghum, sudan), Miscanthus giganteus (miscanthus), Saccharum sp. (energycane), Populus balsamifera (poplar), Zea mays (corn), Glycine max (soybean), Brassica napus (canola), Triticum aestivum (wheat), Gossypium hirsutum (cotton), Oryza sativa (rice), Helianthus annuus (sunflower), Medicago sativa (alfalfa), Beta vulgaris (sugarbeet), Pennisetum glaucum (pearl millet), Panicum spp. Sorghum spp., Miscanthus spp., Saccharum spp., Erianthus spp., Populus spp., Secale cereale (rye), Salix spp. (willow), Eucalyptus spp. (eucalyptus), Triticosecale spp. (triticum-25 wheat X rye), Bamboo, Carthamus tinctorius (safflower), Jatropha curcas (Jatropha), Ricinus communis (castor), Elaeis guineensis (oil palm), Phoenix dactylifera (date palm), Archontophoenix cunninghamiana (king palm), Syagrus romanzoffiana (queen palm), Linum usitatissimum (flax), Brassica juncea, Manihot esculenta (cassaya), Lycopersicon esculentum (tomato), Lactuca saliva (lettuce), Musa paradisiaca (banana), Solanum tuberosum (potato), Brassica oleracea (broccoli, cauliflower, brussel sprouts), Camellia sinensis (tea), Fragaria ananassa (strawberry), Theobroma cacao (cocoa), Coffea arabica (coffee), Vitis vinifera (grape), Ananas comosus (pineapple), Capsicum annum (hot & sweet pepper), Allium cepa (onion), Cucumis melo (melon), Cucumis sativus (cucumber), Cucurbita maxima (squash), Cucurbita moschata (squash), Spinacea oleracea (spinach), Citrullus lanatus (watermelon), Abelmoschus esculentus (okra), Solanum melongena (eggplant), Papaver somniferum (opium poppy), Papaver orientale, Taxus baccata, Taxus brevifolia, Artemisia annua, Cannabis saliva, Camptotheca acuminate, Catharanthus roseus, Vinca rosea, Cinchona officinalis, Coichicum autumnale, Veratrum californica, Digitalis lanata, Digitalis purpurea, Dioscorea 5 spp., Andrographis paniculata, Atropa belladonna, Datura stomonium, Berberis spp., Cephalotaxus spp., Ephedra sinica, Ephedra spp., Erythroxylum coca, Galanthus wornorii, Scopolia spp., Lycopodium serratum (Huperzia serrata), Lycopodium spp., Rauwolfia serpentina, Rauwolfia spp., Sanguinaria canadensis, Hyoscyamus spp., Calendula officinalis, Chrysanthemum parthenium, Coleus forskohlii, Tanacetum parthenium, Parthenium argentatum (guayule), Hevea spp. (rubber), Mentha spicata (mint), Mentha piperita (mint), Bixa orellana, Alstroemeria spp., Rosa spp. (rose), Dianthus caryophyllus (carnation), Petunia spp. (petunia), Poinsettia pulcherrima (poinsettia), Nicotiana tabacum (tobacco), Lupinus albus (lupin), Uniola paniculata (oats), Hordeum vulgare (barley), and Lolium spp. (rye).
[0304] In some examples, a monocotyledonous plant may be used. Monocotyledonous plants belong to the orders of the Alismatales, Arales, Arecales, Bromeliales, Commelinales, Cyclanthales, Cyperales, Eriocaulales, Hydrocharitales, Juncales, Lilliales, Najadales, Orchidales, Pandanales, Poales, Restionales, Triuridales, Typhales, and Zingiberales. Plants belonging to the class of the Gymnospermae are Cycadales, Ginkgoales, Gnetales, and Pinales. In some examples, the monocotyledonous plant can be selected from the group consisting of a maize, rice, wheat, barley, and sugarcane.
[0305] In some examples, a dicotyledonous plant may be used, including those belonging to the orders of the Aristochiales, Asterales, Batales, Campanulales, Capparales, Caryophyllales, Casuarinales, Celastrales, Cornales, Diapensales, Dilleniales, Dipsacales, Ebenales, Ericales, Eucomiales, Euphorbiales, Fabales, Fagales, Gentianales, Geraniales, Haloragales, Hamamelidales, Middles, Juglandales, Lamiales, Laurales, Lecythidales, Leitneriales, Magniolales, Malvales, Myricales, Myrtales, Nymphaeales, Papeverales, Piperales, Plantaginales, Plumbaginales, Podostemales, Polemoniales, Polygalales, Polygonales, Primulales, Proteales, Rafflesiales, Ranunculales, Rhamnales, Rosales, Rubiales, Salicales, Santales, Sapindales, Sarraceniaceae, Scrophulariales, Theales, Trochodendrales, Umbellales, Urticales, and Violates. In some examples, the dicotyledonous plant can be selected from the group consisting of cotton, soybean, pepper, and tomato.
[0306] In some cases, the plant to be improved is not readily amenable to experimental conditions. For example, a crop plant may take too long to grow enough to practically assess an improved trait serially over multiple iterations. Accordingly, a first plant from which bacteria are initially isolated, and/or the plurality of plants to which genetically manipulated bacteria are applied may be a model plant, such as a plant more amenable to evaluation under desired conditions. Non-limiting examples of model plants include Setaria, Brachypodium, and Arabidopsis. Ability of bacteria isolated according to a method of the disclosure using a model plant may then be applied to a plant of another type (e.g. a crop plant) to confirm conferral of the improved trait.
[0307] Traits that may be improved by the methods disclosed herein include any observable characteristic of the plant, including, for example, growth rate, height, weight, color, taste, smell, changes in the production of one or more compounds by the plant (including for example, metabolites, proteins, drugs, carbohydrates, oils, and any other compounds). Selecting plants based on genotypic information is also envisaged (for example, including the pattern of plant gene expression in response to the bacteria, or identifying the presence of genetic markers, such as those associated with increased nitrogen fixation). Plants may also be selected based on the absence, suppression or inhibition of a certain feature or trait (such as an undesirable feature or trait) as opposed to the presence of a certain feature or trait (such as a desirable feature or trait).
Concentrations and Rates of Application of Agricultural Compositions
[0308] As aforementioned, the agricultural compositions of the present disclosure, which comprise a taught microbe, can be applied to plants in a multitude of ways. In two particular aspects, the disclosure contemplates an in-furrow treatment or a seed treatment
[0309] For seed treatment embodiments, the microbes of the disclosure can be present on the seed in a variety of concentrations. For example, the microbes can be found in a seed treatment at a cfu concentration, per seed of: 1.times.10.sup.1, 1.times.10.sup.2, 1.times.10.sup.3, 1.times.10.sup.4, 1.times.10.sup.5, 1.times.10.sup.6, 1.times.10.sup.17, 1.times.10.sup.8, 1.times.10.sup.9, 1.times.10.sup.10, or more. In particular aspects, the seed treatment compositions comprise about 1.times.10.sup.4 to about 1.times.10.sup.8 cfu per seed. In other particular aspects, the seed treatment compositions comprise about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. In other aspects, the seed treatment compositions comprise about 1.times.10.sup.6 cfu per seed.
[0310] In the United States, about 10% of corn acreage is planted at a seed density of above about 36,000 seeds per acre; 1/3 of the corn acreage is planted at a seed density of between about 33,000 to 36,000 seeds per acre; 1/3 of the corn acreage is planted at a seed density of between about 30,000 to 33,000 seeds per acre, and the remainder of the acreage is variable. See, "Corn Seeding Rate Considerations," written by Steve Butzen, available at: https://www.pioneer.com/home/site/us/agronomy/librarykorn-seeding-rate-co- nsiderations/
[0311] Table B below utilizes various cfu concentrations per seed in a contemplated seed treatment embodiment (rows across) and various seed acreage planting densities (1.sup.st column: 15K-41K) to calculate the total amount of cfu per acre, which would be utilized in various agricultural scenarios (i.e. seed treatment concentration per seed.times.seed density planted per acre). Thus, if one were to utilize a seed treatment with 1.times.10.sup.6 cfu per seed and plant 30,000 seeds per acre, then the total cfu content per acre would be 3.times.10.sup.10 (i.e. 30K*1.times.10.sup.6).
TABLE-US-00002 TABLE B Total CFU Per Acre Calculation for Seed Treatment Embodiments Corn Population (i.e, seeds per acre) 1.00E+02 1.00E+03 1.00E+04 1.00E+05 1.00E+06 1.00E+07 1.00E+08 1.00E+09 15,000 1.50E+06 1.50E+07 1.50E+08 1.50E+09 1.50E+10 1.50E+11 1.50E+12 1.50E+13 16,000 1.60E+06 1.60E+07 1.60E+08 1.60E+09 1.60E+10 1.60E+11 1.60E+12 1.60E+13 17,000 1.70E+06 1.70E+07 1.70E+08 1.70E+09 1.70E+10 1.70E+11 1.70E+12 1.70E+13 18,000 1.80E+06 1.80E+07 1.80E+08 1.80E+09 1.80E+10 1.80E+11 1.80E+12 1.80E+13 19,000 1.90E+06 1.90E+07 1.90E+08 1.90E+09 1.90E+10 1.90E+11 1.90E+12 1.90E+13 20,000 2.00E+06 2.00E+07 2.00E+08 2.00E+09 2.00E+10 2.00E+11 2.00E+12 2.00E+13 21,000 2.10E+06 2.10E+07 2.10E+08 2.10E+09 2.10E+10 2.10E+11 2.10E+12 2.10E+13 22,000 2.20E+06 2.20E+07 2.20E+08 2.20E+09 2.20E+10 2.20E+11 2.20E+12 2.20E+13 23,000 2.30E+06 2.30E+07 2.30E+08 2.30E+09 2.30E+10 2.30E+11 2.30E+12 2.30E+13 24,000 2.40E+06 2.40E+07 2.40E+08 2.40E+09 2.40E+10 2.40E+11 2.40E+12 2.40E+13 25,000 2.50E+06 2.50E+07 2.50E+08 2.50E+09 2.50E+10 2.50E+11 2.50E+12 2.50E+13 26,000 2.60E+06 2.60E+07 2.60E+08 2.60E+09 2.60E+10 2.60E+11 2.60E+12 2.60E+13 27,000 2.70E+06 2.70E+07 2.70E+08 2.70E+09 2.70E+10 2.70E+11 2.70E+12 2.70E+13 28,000 2.80E+06 2.80E+07 2.80E+08 2.80E+09 2.80E+10 2.80E+11 2.80E+12 2.80E+13 29,000 2.90E+06 2.90E+07 2.90E+08 2.90E+09 1.90E+10 2.90E+11 2.90E+12 2.90E+13 30,000 3.00E+06 3.00E+07 3.00E+08 3.00E+09 3.00E+10 3.00E+11 3.00E+12 3.00E+13 31,000 3.10E+06 3.10E+07 3.10E+08 3.10E+09 3.10E+10 3.10E+11 3.10E+12 3.10E+13 32,000 3.20E+06 3.20E+07 3.20E+08 3.20E+09 3.20E+10 3.20E+11 3.20E+12 3.20E+13 33,000 3.30E+06 3.30E+07 3.30E+08 3.30E+09 3.30E+10 3.30E+11 3.30E+12 3.30E+13 34,000 3.40E+06 3.40E+07 3.40E+08 3.40E+09 3.40E+10 3.40E+11 3.40E+12 3.40E+13 35,000 3.50E+06 3.50E+07 3.50E+08 3.50E+09 3.50E+10 3.50E+11 3.50E+12 3.50E+13 36,000 3.60E+06 3.60E+07 3.60E+08 3.60E+09 3.60E+10 3.60E+11 3.60E+12 3.60E+13 37,000 3.70E+06 3.70E+07 3.70E+08 3.70E+09 3.70E+10 3.70E+11 3.70E+12 3.70E+13 38,000 3.80E+06 3.80E+07 3.80E+08 3.80E409 3.80E+10 3.80E+11 3.80E+12 3.80E+13 39,000 3.90E+06 3.90E+07 3.90E+08 3.90E+09 3.90E+10 3.90E+11 3.90E+12 3.90E+13 40,000 4.00E+06 4.00E+07 4.00E+08 4.00E+09 4.00E+10 4.00E+11 4.00E+12 4.00E+13 41,000 4.10E+06 4.10E+07 4.10E+08 4.10E+09 4.10E+10 4.10E+11 4.10E+12 4.10E+13
[0312] For in-furrow embodiments, the microbes of the disclosure can be applied at a cfu concentration per acre of: 1.times.10.sup.6, 3.20.times.10.sup.10, 1.60.times.10.sup.11, 3.20.times.10.sup.11, 8.0.times.10.sup.11, 1.6.times.10.sup.12, 3.20.times.10.sup.12, or more. Therefore, in aspects, the liquid in-furrow compositions can be applied at a concentration of between about 1.times.10.sup.6 to about 3.times.10.sup.12 cfu per acre.
[0313] In some aspects, the in-furrow compositions are contained in a liquid formulation. In the liquid in-furrow embodiments, the microbes can be present at a cfu concentration per milliliter of: 1.times.10.sup.1, 1.times.10.sup.2, 1.times.10.sup.3, 1.times.10.sup.4, 1.times.10.sup.5, 1.times.10.sup.6, 1.times.10.sup.7, 1.times.10.sup.8, 1.times.10.sup.9, 1.times.10.sup.10, 1.times.10.sup.11, 1.times.10.sup.12, 1.times.10.sup.13, or more. In certain aspects, the liquid in-furrow compositions comprise microbes at a concentration of about 1.times.10.sup.6 to about 1.times.10.sup.11 cfu per milliliter. In other aspects, the liquid in-furrow compositions comprise microbes at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter. In other aspects, the liquid in-furrow compositions comprise microbes at a concentration of about 1.times.10.sup.8 to about 1.times.10.sup.9 cfu per milliliter. In other aspects, the liquid in-furrow compositions comprise microbes at a concentration of up to about 1.times.10.sup.13 cfu per milliliter.
Examples
[0314] The examples provided herein describe methods of bacterial isolation, bacterial and plant analysis, and plant trait improvement. The examples are for illustrative purposes only and are not to be construed as limiting in any way.
Example 1: Isolation of Microbes from Plant Tissue
[0315] Topsoil was obtained from various agricultural areas in central California. Twenty soils with diverse texture characteristics were collected, including heavy clay, peaty clay loam, silty clay, and sandy loam. Seeds of various field corn, sweet corn, heritage corn and tomato were planted into each soil, as shown in Table 1.
TABLE-US-00003 TABLE 1 Crop Type and Varieties planted into soil with diverse characteristics Crop Type Field Corn Sweet Corn Heritage Corn Tomato Varieties Mo17 Ferry-Morse Victory Seeds Ferry-Morse Roma `Golden Cross `Moseby Prolific` VF Bantam T-51` B73 Ferry-Morse `Silver Victory Seeds `Reid's Stover Roma Queen Hybrid` Yellow Dent` DKC 66- Ferry-Morse `Sugar Victory Seeds Totally Tomatoes 40 Dots` `Hickory King` `Micro Tom Hybrid` DKC 67- Heinz 1015 07 DKC 70- Heinz 2401 01 Heinz 3402 Heinz 5508 Heinz 5608 Heinz 8504
[0316] Plants were uprooted after 2-4 weeks of growth and excess soil on root surfaces was removed with deionized water. Following soil removal, plants were surface sterilized with bleach and rinsed vigorously in sterile water. A cleaned, 1 cm section of root was excised from the plant and placed in a phosphate buffered saline solution containing 3 mm steel beads. A slurry was generated by vigorous shaking of the solution with a Qiagen TissueLyser II.
[0317] The root and saline slurry was diluted and inoculated onto various types of growth media to isolate rhizospheric, endophytic, epiphytic, and other plant-associated microbes. R2A and Nfb agar media were used to obtain single colonies, and semisolid Nfb media slants were used to obtain populations of nitrogen fixing bacteria. After 2-4 weeks incubation in semi-solid Nfb media slants, microbial populations were collected and streaked to obtain single colonies on R2A agar, as shown in FIG. 1A-B. Single colonies were resuspended in a mixture of R2A and glycerol, subjected to PCR analysis, and frozen at -80.degree. C. for later analysis. Approximately 1,000 single colonies were obtained and designated "isolated microbes."
[0318] Isolates were then subjected to a colony PCR screen to detect the presence of the nifH gene in order to identify diazotrophs. The previously-described primer set Ueda 19F/388R, which has been shown to detect over 90% of diazotrophs in screens, was used to probe the presence of the nif cluster in each isolate (Ueda et al. 1995; J. Bacteriol. 177: 1414-1417). Single colonies of purified isolates were picked, resuspended in PBS, and used as a template for colony PCR, as shown in FIG. 2. Colonies of isolates that gave positive PCR bands were re-streaked, and the colony PCR and re-streaking process was repeated twice to prevent false positive identification of diazotrophs. Purified isolates were then designated "candidate microbes."
Example 2: Characterization of Isolated Microbes
Sequencing, Analysis and Phylogenetic Characterization
[0319] Sequencing of 16S rDNA with the 515f-806r printer set was used to generate preliminary phylogenetic identities for isolated and candidate microbes (see e.g. Vernon et al.; BMC Microbiol. 2002 Dec. 23; 2:39.). The microbes comprise diverse genera including: Enterobacter, Burkholderia, Klebsiella, Bradyrhizobium, Rahnella, Xanthomonas, Raoultella, Pantoea, Pseudomonas, Brevundimonas, Agrobacterium, and Paenibacillus, as shown in Table 2.
TABLE-US-00004 TABLE 2 Diversity of microbes isolated from tomato plants as determined by deep 16S rDNA sequencing. Genus Isolates Achromobacter 7 Agrobacterium 117 Agromyces 1 Alicyclobacillus 1 Asticcacaulis 6 Bacillus 131 Bradyrhizobium 2 Brevibacillus 2 Burkholderia 2 Coulobacter 17 Chryseobacterium 42 Comamonas 1 Dyadobacter 2 Flavobacterium 46 Halomonas 3 Leptothrix 3 Lysobacter 2 Neisseria 13 Paenibacillus 1 Paenisporosarcina 3 Pantoea 14 Pedobacter 16 Pimelobacter 2 Pseudomonas 212 Rhizobium 4 Rhodoferax 1 Sphingobacterium 13 Sphingobium 23 Sphingomonas 3 Sphingopyxis 1 Stenotrophomonas 59 Streptococcus 3 Variovorax 37 Xylanimicrobium 1 unidentified 75
[0320] Subsequently, the genomes of 39 candidate microbes were sequenced using Illumina Miseq platform. Genomic DNA from pure cultures was extracted using the QIAmp DNA mini kit (QIAGEN), and total DNA libraries for sequencing were prepared through a third party vendor (SeqMatic, Hayward). Genome assembly was then carried out via the AS pipeline (Tritt et al. 2012; PLoS One 7(9):e42304). Genes were identified and annotated, and those related to regulation and expression of nitrogen fixation were noted as targets for mutagenesis.
Transcriptomic Profiling of Candidate Microbes
[0321] Transcriptomic profiling of strain 0010 was performed to identify promoters that are active in the presence of environmental nitrogen. Strain 0010 was cultured in a defined, nitrogen-free media supplemented with 10 mM glutamine. Total RNA was extracted from these cultures (QIAGEN RNeasy kit) and subjected to RNAseq sequencing via Illumina HiSeq (SeqMatic, Fremont Calif.). Sequencing reads were mapped to CI010 genome data using Geneious, and highly expressed genes under control of proximal transcriptional promoters were identified.
[0322] Tables 3A-C lists genes and their relative expression level as measured through RNASeq sequencing of total RNA. Sequences of the proximal promoters were recorded for use in mutagenesis of nif pathways, nitrogen utilization related pathways, or other genes with a desired expression level.
TABLE-US-00005 TABLE 3A Name Minimum Maximum Length Direction murein lipoprotein CDS 2,929,898 2,930,134 237 forward membrane protein CDS 5,217,517 5,217,843 327 forward zinc/cadmium-binding 3,479,979 3,480,626 648 forward protein CDS acyl carrier protein CDS 4,563,344 4,563,580 237 reverse ompX CDS 4,251,002 4,251,514 513 forward DNA-binding protein HU- 375,156 375,428 273 forward beta CDS sspA CDS 629,998 630,636 639 reverse tatE CDS 3,199,435 3,199,638 204 reverse LexA repressor CDS 1,850,457 1,851,065 609 forward hisS CDS <3999979 4,001,223 >1245 forward
TABLE-US-00006 TABLE 3B RNASeq_ RNASeq_ RNASeq_ RNASeq_ Differential nifL- nifL- WT- WT- Expression Differential Raw Raw Raw Raw Absolute Expression Read Transcript Read Transcript Name Confidence Ratio Count Count Count Count murein 1000 -1.8 12950.5 10078.9 5151.5 4106.8 lipoprotein CDS membrane 1000 -1.3 9522.5 5371.3 5400 3120 protein CDS zinc/cadmium- 3.3 1.1 6461 1839.1 5318 1550.6 binding protein CDS acyl carrier 25.6 1.6 1230.5 957.6 1473.5 1174.7 protein CDS ompX CDS 1.7 1.1 2042 734.2 1687.5 621.5 DNA-binding 6.9 -1.3 1305 881.7 725 501.8 protein HU- beta CDS sspA CDS 0.2 1 654 188.8 504.5 149.2 tatE CDS 1.4 1.3 131 118.4 125 115.8 LexA 0.1 -1.1 248 75.1 164 50.9 repressor CDS hisS CDS 0 -1.1 467 69.2 325 49.3
TABLE-US-00007 TABLE 3C Prm (In Forward SEQ SEQ SEQ direction, -250 ID Expressed ID Neighbor ID Name to +10 region) NO: Sequence NO: Sequence NO: murein GCCTCTCGGGGC 3 ATGAATCGTACT 13 ATGAAAAAGACC 23 lipoprotein GCTTTTTTTTATT AAACTGGTACTG AAAATTGTTTGC CDS CCGGCACTAGCC GGCGCGGTAATC ACCATCGGTCCG GCTATTAATAAA CTGGGTTCTACTC AAAACCGAATCC AATGCAAATCGG TGCTGGCTGGTT GAAGAGATGTTG AATTTACTATTTA GCTCCAGCAATG ACCAAAATGCTG ACGCGAGATTAT CTAAAATCGATC GACGCGGGCATG CTAAGATGAATC AGCTGTCTTCTGA AACGTTATGCGT CGATGGAAGCGC CGTTCAGACTCT CTGAACTTCTCTC GCTGTTTTCACTC GAACGCTAAAGT ACGGTGACTATG GCCTTTTTAAAGT TGACCAGCTGAG CGGAACACGGTC TACGTGATGATTT CAACGACGTGAA AGCGCATCCAGA CGATGCTTCTTTG CGCAATGCGTTC ATCTGCGCAATG AGCGAACGATCA CGACGTTCAGGC TGATGAGTAAAA AAAATAAGCGTA TGCTAAAGATGA CCGGTAAGAAAG TTCAGGTAAAAA CGCAGCTCGCGC CGGCAATCCTGC AATATTCTCATCA TAACCAGCGTCT TGGACACCAAAG CAAAAAAGTTTG GGACAACGCAGC GTCCGGAAATCC TGTAATACTTGTA TACTAAATACCG GTACCATTAAGC ACGCT--- TAAGTAA TGGAAGGCGGCA ACATGGAGATTA ACGACGTCTCCC ACTC TGAAAGCGGGCC AGACCTTCACCTT CACCACCGATAA ATCCGTTGTCGGT AATAACGAAATC GTTGCGGTGACC TATGAAGGCTTC ACCAGCGACCTG AGCGTTGGCAAC ACGGTACTGGTT GACGATGGTCTG ATCGGTATGGAA GTGACCGCTATC GAAGGCAACAAA GTTGTTTGTAAA GTGCTGAACAAC GGCGACCTCGGC GAGAACAAAGGC GTTAACCTGCCG GGCGTATCTATC GCGCTGCCGGCG CTGGCTGAAAAA GACAAACAGGAT CTGATCTTCGGTT GCGAACAGGGCG TTGACTTTGTTGC GGCATCCTTTATC CGTAAGCGTTCT GACGTTGTTGAA ATCCGTGAGCAC CTGAAAGCCCAC GGCGGCGAGAAG ATCCAGATCATC TCCAAAATCGAA AACCAGGAAGGC CTGAACAACTTC GACGAAATCCTC GAAGCCTCTGAC GGCATCATGGTA GCCCGTGGCGAC CTGGGCGTTGAA ATCCCGGTTGAA GAAGTTATCTTC GCGCAGAAGATG ATGATCGAGAAA TGTATCCGCGCG CGTAAAGTCGTT ATCACCGCGACC CAGATGCTGGAT TCCATGATCAAA AACCCGCGTCCG ACCCGTGCGGAA GCAGGCGACGTG GCCAACGCCATC CTCGACGGCACC GACGCAGTTATG CTGTCCGGCGAA TCCGCGAAAGGT AAATACCCGCTG GAAGCGGTCACC ATCATGGCGACC ATCTGCGAACGT ACCGACCGCGTC ATGACCAGCCGT CTTGAGTACAAC AACGACAACCGT AAGCTGCGCATC ACCGAAGCGGTG TGCCGCGGTGCG GTAGAAACGGCT GAAAAACTGGAA GCGCCGCTGATC GTTGTGGCAACC CAGGGCGGTAAA TCCGCGCGCGCC GTACGTAAATAC TTCCCGGATGCC ACTATCCTGGCG CTGACCACCAAC GAAACCACCGCG CGTCAGCTGGTG CTGAGCAAAGGC GTTGTGGCACAG CTGGTTGAAGAT ATCTCCTCTACCG ATGCGTTCTACAT CCAGGGTAAAGA ACTGGCGCTGCA GAGCGGTCTGGC GCGTAAAGGCGA CGTGGTTGTTATG GTTTCCGGCGCG TTAGTCCCGAGC GGAACCACCAAT ACCGCTTCCGTG CACGTGCTGTAA membrane GGTTCACATAAA 4 ATGGCCAACCGA 14 ATGTATTTAAGA 24 protein CATAATTATCGC GCAAACCGCAAC CCCGATGAGGTG CDS CACGGCGATAGC AACGTAGAAGAG GCGCGTGTTCTTG CGTACGCTTTTTG AGCGCTGAAGAT AAAAAGCCGGCT CGTCACAACATC ATCCATAACGAT TCACCATGGATG CATGGTGAAGCC GTCAGCCAATTA TTGTGACGCAAA GGCTTTTTCAAG GCGGATACGCTG AAGCGTACGGCT AACACGCGCCAC GAAGAGGTGCTG ATCGCCGTGGCG CTCATCGGGTCTT AAATCGTGGGGC ATAATTATGTTTA AATACATACTC AGCGACGCCAAA TGTGAACCGTGA ATTCCTCATTATC GACGAAGCGGAG AGCTCGTATGGG TTTTACCGCACGT GCCGCGCGCAAA GCGTACCGCGTT TAACCTTACCTTA AAAGCGCAGGCG AATTATTCATCCG TTCATTAAAGGC CTGCTGAAAGAG GCTTTAAAAGAG AACGCTTTCGGA ACCCGCGCCCGG CGCAGCACAACG ATATTCCATAAA CTTAACGGCAAC CTTGCGGAGCCC GGGCTATTTACA AACCGCGTCCAG GCGTCGGATATC GCATAATTCAAA CAGGCGGCGTGC AAAACCTGCGAT ATCTTGTCCTACA GACGCCATGGGC CATTATGAGCAG CTTATAGACTCA TGCGCTGACAGC TTCCCGCTCTATT ATGGAATTAAGG TACGTGCGCGAC TAGCGCTGGGATG GA AAACCGTGGCAA CTCAACACTCATT AGCGTCGGCGCC ATGGTATTCCAC GCAGCAGCCGTT ACGGGTTCAGTT GGGGTATTTATT CGCGAATGGCGC GGCGTATTACTG TTGAGCGTTTTCT AATTTACGTCGA GAGTGGCCTGTT TAA TGGCGAAACGCA GTATAGCTGA zinc/ GCGCGGAAAATC 5 ATGACCAAAAAG 15 ATGGATAGCGAC 25 cadmium- GACGCATAGCGC ATTTCCGCCCTAG ATTAATCAGGTC binding ATTCTCAGAAGC CGTTTGGCATTG ATTGATTCTTTTG protein CGGCCTGGTCTC GCATGGTAATGG TTAAAGGCCCGG CDS GGTGGAAAAGCG CGAGCAGCCAGG CGGTCGTGGGAA AATCTTTCCCACG CTTTTGCCCACGG AGATTCGCTTTTC ACCGCCGGGCCT TCACCATAGTCA CACCGAGACCAG TTAACAAAAGAA TGGCCCGGCGCT GCCGGCTTCTGA TCAATGACCTGA GACCGAAGCGGA GAATGCGCTATG TTAATGTCGCTAT ACAAAAGGCGAG CGTCGATTTTCCG CCATTCTCTCTCC TGAAGGCATTTTT CGCCTCGAAATC GCGTAATGCGAT GCTGACCAGGAC ATGCTTGCGGGT CTTTTTTCATCAT CTTAAAGGACAGG CAGCTTCACGAT ACCTAACAAACT GCGCTGAGCGAC CCGGCGATTAAA GGCAGAGGGAAA TGGGAGGGGATC GCCGATCGCGCC AGCCGCGCGGTT TGGCAGTCGGTT CAGCTCATGCCG TTTCTGCGAAGT AACCCCTATCTG CACGATGTGCTG GTATTGTAAGAT CTGAACGGGGAT TATATTCCCGCTG TTGTTTGATATGT TTAGATCCGGTTC GCGGATGGAATG TATATCGTAACA TGGAGCAGAAGG ACCCGCAATGGC TATTATTGCAAA CCAAAAAGGCCG TGGCGCCCTCCA CAT GTAAAAGCGTGG CTCTGCTCACTAT CGGAATATCGGG CTTATTTGGTAAA AATATTATAAGA CAGCAGCTGGAA AGGGCTACGCTA TTCGTCCTGCGCC CCGATGTCGACC ACTGGGACGGCA AGATTGGTATCG GCGCGCTTAACG AGGATAACGTCA TGCTGGATAAAC TGGAGTTTCACG AGCAGGTTCCGC TCGGGAAAACCG GCCGCGGTCCCC TCAACGCCTGTA GGGTCGGCTCTTT AGTACAGCTATT TCTGCTGCAGGC CCGGTTACAAAA GCTGAATGAAAT TTCTGACCTACGC GCAGATGCAGCC ATCCGGTAAAAA GCGGGAGCAGCA AGGCGTGCGCTA CACGGCCCGCTT CCTGTTCGAATG TATTGTCACCAG CCAGCAGGCGGA CCTGCTCAGCCA TTCAAAAGCGCC CTGTGCCGATCT GAAGTTTGTTCA GCTGGGCAGCCA GTTTAGCGATCA GGTACAAACCTC CACCATCGCGCC ATCGCGCAGCCA ACGCAAGTCCCA GGCGCTTTTTGA GCATTTCCACATC AGCGATTCGTAA TTTATGGGCAAT GCATATTGACGC GAGTCCCAGGAA CCACTTTGCCGA GCGCTGCTGAAA CCCGTTAACCCG GAGATGGATAAC GGAGTCGGTGGC TGGCCAACCTAC GCAGGCGTTTTA TATCCTTATGCGC CCTCTCGCCAAA TGCATAAAGAGC CTATCTATCCCAC AGATTGTCGACG CTGTTCCAGAAA AAATGCTGCACC TGCGGGCCAATG ACTAA GGCTTTAACGAG TATCTGAATCAC ATCCGCCTGGAG CAGGCCAGAATG CTGTTAAAAGGC CACGATATGAAA GTGAAAGATATC GCCCACGCCTGC GGTTTCGCCGAC AGCAACTACTTC TGCCGCCTGTTTC GCAAAAACACCG AACGCTCGCCGT CGGAGTATCGCC GTCAATATCACA GCCAGCTGACGG AAAAAACAGCCC CGGCAAAAAACT AG acyl CTGACGAAGCGA 6 ATGAGCACTATC 16 ATGAGTTTTGAA 26 carrier GTFACATCACCG GAAGAACGCGTT GGAAAAATCGCG protein GTGAAACTCTGC AAGAAAATTATC CTGGTTACCGGT CDS ACGTCAACGGCG GGCGAACAGCTG GCAAGTCGCGGG GAATGTATATGG GGCGTTAAGCAG ATTGGCCGCGCA TCTGACCGAGAT GAAGAAGTTACC ATCGCTGAAACG TTGCGCAAAACG AACAATGCTTCC CTCGTTGCCCGTG CTCAGGAACCGC TTCGTTGAAGAC GCGCGAAAGTTA GCAGTCTGTGCG CTGGGCGCTGAT TCGGGACTGCGA GTTCACTGTAAT TCTCTTGACACCG CCAGCGAAAGCG GTTTTGTACAAA TTGAGCTGGTAA GCGCGCAGGCGA ATGATTTGCGTTA TGGCTCTGGAAG TCAGCGATTATTT TGAGGGCAAACA AAGAGTTTGATA AGGTGCTAACGG GCCGCAAAATAG CTGAGATTCCGG TAAAGGTCTGCT CGTAAAATCGTG ACGAAGAAGCTG GCTGAATGTGAC GTAAGACCTGCC AGAAAATCACTA CGATCCTGCATCT GGGATTTAGTTG CTGTTCAGGCTG ATTGAATCTGTTC CAAATTTTTCAAC CCATTGATTACAT TGGGAAATATTC ATTTTATACACTA CAACGGCCACCA GCGCAGAATTTG CGAAAACCATCG GGCGTAA GTGAAGTTGATA CGAAAGCGAGTT TCCTGGTGAACA TTGA ATGCCGGGATCA CTCGTGATAACC
TGTTAATGCGCA TGAAAGATGATG AGTGGAACGATA TTATCGAAACCA ACCTGTCATCTGT TTTCCGTCTGTCA AAAGCGGTAATG CGCGCTATGATG AAAAAGCGTCAT GGACGTATTATC ACTATCGGTTCTG TGGTTGGTACCA TGGGAAATGCGG GTCAGGCCAACT ACGCTGCGGCGA AAGCGGGTCTGA TTGGCTTCAGTA AATCACTGGCTC GCGAAGTTGCGT CCCGCGGTATTA CTGTAAACGTTG TTGCTCCGGGCTT TATTGAAACGGA CATGACGCGTGC GCTGACCGATGA GCAGCGTGCGGG TACGCTGGCGGC AGTTCCTGCGGG GCGCCTCGGCTC TCCAAATGAAAT CGCCAGTGCGGT GGCATTTTTAGCC TCTGACGAAGCG AGTTACATCACC GGTGAAACTCTG CACGTCAACGGC GGAATGTATATG GTCTGA ompX ACGCCTGGGGCG 7 ATGAATAAAATT 17 ATGCCCGGCTCG 27 CDS CCGACCAGCGGG GCACGTTTTTCAG TCTCGTAAGGTA AAGAGTGATTTG CACTGGCCGTTG CCGGCATGGTTG GCCAACGAGGCG TTCTGGCTGCATC CCGATACTGGTT CCGCTCTGAATG CGTAGGTACCAC ATTTTAATCGCCA GAAATCATGGCG TGCTTTCGCTGCG TGATTTCCAT ATTAAAATAACC ACTTCTACCGTTA AGTATCGGCAAC CCGGTGGCTACG CATGCCGGTACC CGCAGAGCGACA TTACGAGACGAG TGCAGGGTGAAG CCGGGCATCCTTT CGAACAAAGCTG CTCCTGTCAATTT GCGGTTTCAACC TGTCAAATGCGG TGAAGTACCGCT TAAAGGTTCCAG ACGAGCAAGACA TGTAATTGAATT ACAACCCGCTGG ACCCCGCGCCGG GTGTTATCGGTTC TTGAGCTAATGTT TTTCACCTACACC GAAAAAAAGGGT GAAAAAGATCGT CTTAAAAGCAGT TCTGAATCTGGC ACAATAGGGCGG GTTTACAAAAAA GTCTGAAGATAA GGCCAGTACTAC TTTCA GGCATCACCGCA GGTCCGGCTTAC CGTCTGAACGAC TGGGCTAGCATC TACGGCGTAGTG GGTGTTGGTTAC GGTAAATTCCAG GACAACAGCTAC CCGAACAAATCT GATATGAGCGAC TACGGTTTCTCTT ACGGCGCTGGTC TGCAGTTCAACC CGATCGAAAACG TTGCCCTGGACTT CTCCTACGAGCA GTCTCGCATTCGT AACGTTGACGTT GGCACCTGGATT GCTGGCGTAGGT TACCGCTTCTAA DNA- TCTGATTCCTGAT 8 GTGAATAAATCT 18 ATGAATCCTGAG 28 binding GAAAATAAACGC CAACTGATTGAC CGTTCTGAACGC protein GACCTTGAAGAA AAAATTGCTGCC ATTGAAATCCCC HU-beta ATTCCGGATAAC GGTGCGGACATT GTATTGCCGTTGC CDS GTTATCGCCGATT TCTAAAGCCGCA GCGATGTGGTGG TAGATATCCATC GCTGGACGTGCG TTTATCCGCACAT CGGTGAAACGAA TTAGATGCTTTAA GGTCATACCCCT TCGAGGAAGTTC TCGCTTCTGTTAC GTTTGTAGGGCG TGGCACTTGCGC TGAATCTCTGCA GGAAAAATCTAT TACAGAACGAAC GGCTGGAGATGA CCGTTGTCTCGA CGTTTGGAATGG CGTTGCGCTGGT AGCAGCCATGGA AAGTCGTCACGG AGGGTTTGGTAC CCATGATAAAAA CAAAATAGTGAT TTTTGCTGTTAAA AATCATGCTGGT TTCGCGCAAATA GAGCGCGCTGCC TGCGCAGAAAGA GCGCTAAGAAAA CGTACTGGTCGC AGCCTCGACGGA ATAGGGCTGGTA AATCCGCAAACA TGAGCCGGGTGT AGTAAATTCGTA GGCAAAGAAATC AAACGATCTTTTC CTTGCCAGCCTTT ACCATTGCTGCT ACCGTCGGGACC TTTTGTGTAGCTA GCTAAAGTTCCG GTGGCGTCTATTT ACTTAGATCGCT GGTTTCCGCGCA TGCAAATGCTGA GGCAGGGGGGTC GGTAAAGCGCTG AGCTACCGGACG AATT AAAGACGCGGTA GTACTGTTAAAG AACTGA TGCTGGTCGAAG GTTTGCAGCGCG CGCGCATCTCTG CGCTGTCTGATA ATGGCGAACATT TTTCGGCGAAGG CGGAATACCTTG AATCGCCGGCGA TTGACGAACGCG AGCAGGAAGTGC TGGTTCGTACCG CTATCAGCCAGT TTGAAGGCTACA TCAAGCTGAACA AAAAAATCCCTC CGGAAGTGCTGA CGTCGCTGAATA GCATCGACGATC CGGCGCGTCTGG CGGATACCATCG CTGCGCATATGC CGCTGAAGCTGG CGGACAAACAGT CCGTGCTGGAGA TGTCCGACGTTA ACGAGCGTCTGG AATATCTGATGG CGATGATGGAGT CGGAAATCGATC TGCTGCAGGTGG AGAAGCGTATTC GCAACCGCGTGA AAAAGCAGATGG AGAAATCTCAGC GCGAGTACTATC TGAATGAGCAAA TGAAAGCCATTC AAAAAGAGCTCG GCGAGATGGACG ACCTCCCCGGACG AGAACGAAGCGC TGAAGCGTAAGA TCGACGCGGCGA AAATGCCGAAAG AGGCAAAAGAGA AAACCGAAGCGG AACTGCAAAAAC TGAAAATGATGT CCCCGATGTCGG CGGAAGCGACCG TCGTTCGCGGCT ACATCGACTGGA TGGTGCAGGTAC CGTGGAACGCTC GCAGCAAGGTTA AAAAAGACCTGC GTCAGGCTCAGG AGATCCTCGATA CCGATCACTACG GCCTTGAGCGCG TGAAGGATCGCA TTCTTGAGTACCT CGCGGTGCAGAG CCGTGTTAACAA GCTCAAAGGGCC GATCCTGTGCCT GGTTGGCTCCTCC GGGGGTAGGTAA AACCTCTCTCGG CCAATCCATCGC CAAAGCAACTGG ACGCAAATATGT GCGTATGGCGCT GGGCGGCGTGCG TGATGAAGCGGA AATCCGCGGTCA CCGCCGTACCTA TATTGGCTCAAT GCCGGGCAAACT GATCCAGAAAAT GGCTAAAGTGGG CGTTAAAAACCC GCTGTTCTTGCTG GATGAGATCGAC AAGATGTCTTCT GACATGCGCGGC GATCCGGCCTCG GCGCTGCTGGAG GTGTTGGATCCG GAACAGAACGTG GCCTTTAACGAC CACTATCTGGAA GTGGATTACGAT CTCAGCGACGTG ATGTTCGTTGCG ACCTCTAACTCC ATGAACATCCCG GCGCCGCTGCTG GATCGTATGGAA GTGATCCGCCTCT CCGGCTATACCG AAGATGAGAAGC TAAACATCGCCA AACGCCATCTGC TGTCAAAACAGA TTGAGCGTAACG CGCTCAAGAAAG GCGAGCTGACGG TGGATGACAGCG CGATTATCGGCA TCATTCGCTACTA CACCCGTGAAGC AGGCGTGCGTGG TCTGGAGCGTGA AATCTCGAAACT GTGCCGCAAAGC GGTGAAACAGCT GCTGCTGGATAA GTCGCTGAAACA CATCGAGATTAA CGGCGACAACCT GCACGATTTCCTT GGCGTGCAGCGC TACGACTATGGT CGTGCGGATAGC GAAAACCGCGTA GGTCAGGTGACC GGACTGGCGTGG ACGGAAGTGGGC GGCGATCTGCTG ACCATTGAAACC GCCTGCGTTCCG GGTAAAGGCAAA CTGACCTACACC GGTTCACTGGGT GAAGTCATGCAG GAATCCATCCAG GCGGCGCTGACG GTGGTTCGTTCAC GTGCGGATAAGC TGGGTATTAACT CAGACTTTTACG AAAAACGTGATA TTCACGTTCACGT GCCGGAAGGCGC GACGCCGAAGGA TGGTCCAAGCGC
CGGTATCGCGAT GTGCACCGCGCT GGTTTCCTGTCTG ACGGGTAATCCG GTACGCGCCGAC GTGGCGATGACC GGTGAGATTACC CTCCGTGGCCAG GTATTGCCGATT GGTGGTCTGAAG GAAAAACTGTTG GCCGCGCATCGC GGCGGCATTAAG ACTGTTCTGATTC CTGATGAAAATA AACGCGACCTTG AAGAAATTCCGG ATAACGTTATCG CCGATTTAGATA TCCATCCGGTGA AACGAATCGAGG AAGTTCTGGCAC TTGCGCTACAGA ACGAACCGTTTG GAATGGAAGTCG TCACGGCAAAAT AG sspA CDS GTAAGAAAGTCG 9 ATGGCTGTCGCT 19 ATGGCTGAAAAT 29 GCCTGCGTAAAG GCCAACAAACGT CAATACTACGGC CACGTCGTCGTC TCGGTAATGACG ACCGGTCGCCGC CTCAGTTCTCCAA CTGTTTTCTGGTC AAAAGTTCCGCA ACGTTAATTGTTT CTACTGACATCT GCTCGCGTTTTCA TCTGCTCACGCA ATAGCCATCAGG TCAAACCGGGCA GAACAATTTGCG TCCGCATCGTGCT ACGGTAAAATCG AAAAAACCCGCT GGCCGAAAAAGG TTATCAACCAGC TCGGCGGGTTTTT TGTTAGTTTTGAG GTTCTCTGGAAC TTATGGATAAAT ATAGAGCACGTG AGTACTTCGGTC TTGCCATTTTCCC GAGAAGGACAAC GTGAAACTGCCC TCTACAAACGCC CCGCCTCAGGAT GCATGGTAGTTC CCATTGTTACCAC CTGATTGACCTC GTCAGCCGCTGG TTTTTCAGCATTT AACCCGAATCAA AACTGGTCGACA CCAGAATCCCCT AGCGTACCGACG TGGTTGAGAAAT CACCACAACGTC CTTGTGGATCGT TAGATCTGTACA TTCAAAATCTGG GAGCTCACTCTG TCACCGTTAAAG TAAACTATCATC TGGGAATCTCGC GTGGTGGTATCT CAATTTTCTGCCC ATCATTATGGAA CTGGTCAGGCTG AAATGCAGGTGA TATCTGGATGAG GTGCGATCCGTC TTGTTCATTTTT CGTTTCCCGCATC ACGGTATCACCC CGCCGCTCATGC GCGCTCTGATGG CGGTTTACCCGG AGTACGACGAGT TGGCGCGTGGGG CCCTGCGTGGCG AAAGCCGTCTGT AACTGCGTAAAG ATATGCAGCGTA CTGGTTTCGTTAC TCGAAAAGGACT TCGTGATGCTCGT GGTATTCGTTGAT CAGGTTGAACGT GAATACCATTCA AAGAAAGTCGGC GACCGGTACCGC CTGCGTAAAGCA TGCGCAGGCTGA CGTCGTCGTCCTC TACTGCGCGTAA AGTTCTCCAAAC GCAGCTGCGTGA GTTAA AGAACTACAGGC GATTGCGCCAGT TTTCACCCAGAA GCCCTACTTCCTG AGCGATGAGTTC AGCCTGGTGGAC TGCTACCTGGCA CCACTGCTGTGG CGTCTGCCGGTTC TCGGCGTAGAGC TGGTCGGCGCTG GCGCGAAAGAGC TTAAAGGCTATA TGACTCGCGTATT TGAGCGCGACTC TTTCCTCGCTTCT TTAACTGAAGCC GAACGTGAAATG CGTCTCGGTCGG GGCTAA tatE CDS GTCAAAGCCGTA 10 ATGGGTGAGATT 20 ATGTTTGTTGCTG 30 TTATCGACCCCTT AGTATTACCAAA CCGGACAATTTG AGGGACAACGCT CTGCTGGTAGTC CCGTAACGCCGG TGCCGGGGCGGG GCAGCGCTGATT ACTGGACGGGAA AGAGCGGCCGCA ATCCTGGTGTTTG ACGCGCAGACCT GTTGATTTTTGCC GTACCAAAAAGT GCGTCAGCATGA GAACTTTCAGCT TACGCACGCTGG TGCGCCAGGCCG GATTATATTCAG GTGGAGACCTGG CGGAGCGGGGGG CAGGTACGCGAG GCTCGGCTATCA CGTCGCTTCTGGT CGCCTGCCGGTG AAGGCTTTAAAA TCTGCCTGAGGC TTGCGCAATCGC AAGCCATGAGCG GTTGCTGGCGCG CGCTTTGCGCCA ATGACGATGACA AGACGATAACGA CCGCAATTATTAT GTGCGAAGAAGA TGCGGATTTATC GACGTTTTTTTAA CCAGTGCTGAAG GGTTAAATCCGC ACAAGGCTTGAT AAGCGCCGGCAC CCAGCAGCTGGA TCACCTTGTTACA AGAAGCTCTCTC TGGCGGCTTCTTA GATTGCTATTGTG ATAAAGAGTAA CAGCTCTTGCTG TCCGCGCGTCAA GCGGAGAGCGAA ATAGCCGTTAAT AACAGCGCTTTG TGTATGCGTGTAT ACGACGGTGCTG GATGGCGTATTC ACCCTGCATATC G CCTTCCGGCGAA GGTCGAGCGACG AATACGCTGGTG GCCCTGCGTCAG GGGAAGATTGTG GCGCAATATCAG AAACTGCATCTC TATGATGCGTTC AATATCCAGGAA TCCAGGCTGGTC GATGCCGGGCGG CAAATTCCGCCG CTGATCGAAGTC GACGGGATGCGC GTCGGGCTGATG ACCTGCTACGAT TTACGTTTCCCTG AGCTGGCGCTGT CGTTAGCGCTCA GCGGCGCGCAGC TCATAGTGTTGCC TGCCGCGTGGGT AAAAGGGCCGCT GAAGGAACATCA CTGGGCGACGCT GCTGGCGGCGCG GGCGCTGGATAC AACCTGCTATATT GTCGCCGCAGGA GAGTGCGGGACG CGTAATATCGGT CAAAGCCGTATT ATCGACCCCTTA GGGACAACGCTT GCCGGGGCGGGA GAGCGGCCGCAG TTGATTTTTGCCG AACTTTCAGCTG ATTATATTCAGC AGGTACGCGAGC GCCTGCCGGTGT TGCGCAATCGCC GCTTTGCGCCAC CGCAATTATTAT GA LexA GAGGCGGTGGTT 11 ATGAAAGCGTTA 21 ATGGCCAATAAT 31 repressor GACCGTATCGGT ACGACCAGGCAG ACCACTGGGTTA CDS CCCGAGCATCAT CAAGAGGTGTTT ACCCGAATTATT GAGCTTTCGGGG GATCTCATTCGG AAAGCGGCCGGG CGAGCGAAAGAT GATCATATCAGC TATTCCTGGAAA ATGGGATCGGCG CAGACGGGCATG GGATTCCGTGCG GCGGTACTGCTG CCGCCGACGCGT GCGTGGGTCAAT GCGATTATCATC GCGGAGATTGCT GAGGCCGCATTT GCGCTGATCGCG CAGCGCTTGGGG CGTCAGGAAGGC TGGGGAACGCTG TTTCGCTCCCCAA ATCGCGGCCGTT CTGTGGGCGAAC ACGCGGCGGAAG ATTGCCGTGGCG TACCGCTAAGTC AGCATCTGAAAG ATCGCCTGCTGG TTGTCGTAGCTGC CGCTGGCGCGTA TTGGACGTCGAT TCGCAAAACGGA AAGGCGCAATCG GCCATCACGCGG AAGAAACTCCTG AGATCGTTTCCG GTGCTGCTCATTA ATTTTTGTGTGAA GCGCCTCCCGCG GCTCGGTCCTGTT ATGTGGTTCCAA GTATTCGTCTGCT AGTGATGATAGT AATCACCGTTAG GACGGAAGAAGA TGAAATTATCAA CTGTATATACTCA AACCGGTCTGCC TAGCGCGATTGA CAGCATAACTGT GCTTATTGGCCG GGCGGTGGTTGA ATATACACCCAG CGTCGCGGCAGG CCGTATCGCTCC GGGGC TGAGCCGCTGCT CGAGCATCATGA AGCGCAGCAGCA GCTTTCGGGGCG CATTGAAGGCCA AGCGAAAGATAT CTACCAGGTGGA GGGATCGGCGGC CCCGGCCATGTTT GGTACTGCTGGC AAGCCGAACGCC GATTATCATCGC GATTTTCTGCTGC GCTGATCGCGTG GTGTTAGCGGTA GGGAACGCTGCT TGTCGATGAAGG GTGGGCGAACTA ATATCGGTATTCT CCGCTAA CGATGGCGACCT GCTGGCTGTCCA TAAAACGCAGGA TGTGCGCAATGG TCAGGTGGTTGT GGCGCGTATCGA CGAAGAAGTGAC CGTGAAGCGTCT GAAAAAACAGGG TAACGTCGTGGA ATTGCTGCCGGA AAACAGCGAATT CTCGCCGATCGT GGTCGACCTTCG CGAACAAAGCTT TACTATTGAAGG CCTGGCCGTCGG CGTTATCCGCAA CGGCAACTGGCA ATAA hisS CDS TAAGAAAAGCGG 12 ...ATGAACGATTA 22 ATGCATAACCAG 32 CCTGTACGAAGA TCTGCCGGGCGA GCTCCGATTCAA CGGCGTACGTAA AACCGCTCTCTG CGTAGAAAATCA AGACACTGCTGGA GCAGCGCATTGA AAACGAATTTAC TAACGACGATAT AGGCTCACTGAA GTTGGGAATGTG GATCGATCAGCT GCAGGTGCTTGG CCGATTGGCGAT GGAAGCGCGTAT TAGCTACGGTTA GGCGCCCCCATC TCGCGCTAAAGC CAGCGAAATCCG GCCGTACAGTCG ATCGATGCTGGA TTTGCCGATTGTA ATGACAAACACG TGAGGCGCGTCG GAGCAGACCCCG CGCACCACCGAT TATCGATATCCA TTATTCAAACGC GTGGCGGCGACG GCAGGTTGAAGC GCTATCGGCGAA GTAAATCAAATT GAAATAACGTGT GTGACCGACGTG AAAGCCCTCGAG TGCTGAAGCGATA GTTGAAAAAGAG CGCGTTGGCGCG CGCTTCCCGTGTA ATGTACACCTTTG GATATCGTGCGC TGATTGAACCTG AGGACCGTAACG GTTTCGGTGCCG CGGGCGCGAGGC GCGATAGCCTGA ACGATGGATGCG GCCGGGGTTCAT CTCTACGTCCGG GCGGAAGCGTTC TTTTGTATATATA AAGGCACGGCTG AAACTTATCAAA AAGAGAATAAAC GCTGCGTACGCG CAGCAGGTTAAC GTGGCAAAGAAC CCGGTATCGAAC GTCCCGCTGGTT ATTCAA ATGGTCTCCTGTA GCCGATATCCAC CAATCAAGAACA TTCGATTACCGC GCGCCTGTGGTA ATTGCGCTGAAG CATTGGGCCGAT GTAGCGGAATAC GTTCCGCCACGA GGCGTTGATTGC ACGTCCGCAAAA CTGCGTATTAAC AGGCCGCTACCG CCGGGCAATATC TCAGTTCCACCA GGCAACGAAGAG GATTGGCGCCGA CGTATCCGCATG AGCGTTTGGCCT GTGGTGGACTGC GCAGGGGCCGGA GCTCGCGATAAA TATCGATGCCGA AATATTCCTATCC GCTGATTATGCT GTATCGGGGTAA GACCGCCCGCTG ACGCCGGTTCTCT GTGGCGCGAGCT GGAAAAAGATCT GGGCATCTCCGG CCAGGAAAAATA CCACGTTGCGCT CGGCGAACCGAC GGAGCTGAACTC TCCGCAGGCGCT TATCGGTTCGCTG GCTGGAATCGGC GAGGCTCGCGCT AATGCGCCATGT AACTATCGCGAC TGATCATCTCGAT GCGCTGGTGGCC CGTCTCAACTTCG TATCTTGAGCAG ATCAGTTTAAAG TTTAAAGATAAG TCAGCGTAAAAG CTGGACGAAGAC CCTCCGATGTGTT TGCAAACGCCGC CCTCGCGGTTGA ATGTACACCAAC ATCCTATCGCCTG CCGCTGCGCGTG TTGGCGAAACAG CTGGATTCTAAA ATCGATCAGCCT
AACCCGGACGTC CTGCACCTCGGG CAGGCGCTGCTG ATCACCGAAGCG AACGACGCCCCG GGCGGCGCGCGC ACGCTGGGCGAC AGCGGCGCGGTG TATCTTGATGAA AAGTCCGCGATC GAGTCCAAAACG GGCCTCGGCCTG CATTTTGCCGGG CTGCTGTCTGAA CTGTGCGCGCTG GGGATTGGCGAT CTGGATGATGCC ACGCTGCGCGTC GGTATTCGCTAT TCTCTGGCGGCG ACCGTGAATCAG GATCCCGTTGAA CGTCTGGTACGC GAGATCAAAGTG GGTCTCGACTAC GGCTTCGATATTC TACAACCGCACC TCAAGTCGCTGC GTGTTTGAGTGG GTATTCGCTCTCG GTCACCACCAGC CGGGATCAACTT CTCGGTTCCCAG TATTGCCTGCCCG GGCACCGTCTGC ACCTGTTCACGTC GCCGGAGGCCGT AGGAGTTTGACG TACGATGGTCTG TTATCGGTACCGT GTTGAGCAGCTT TAACGCGCTGGA GGCGGTCGCGCT GCAGCGCCTGGA ACCCCTGGCGTC AGATATCATTAC GGCTTTGCGATG GCCGATGGATAT GGGCTGGAACGT TTCGATCATTGGC CTTGTTTTACTGG TGCGTGGTAAAC TTCAGGCAGTGA GGTCCCGGCGAG ATCCGGAATTTA GCGCTGGTTTCC AAGCCGATCCTG ACCCTCGGCGTA TTGTCGATATATA ACCGGCGGCAAT CCTGGTAGCCTC AAGAAAAGCGGC CGGAACTGACAC CTGTACGAAGAC CCAGTCCGCAGC GGCGTACGTAAA AATGCGTCTGGC GACAGGCTGGAT TGAACAGGTACG AACGACGATATG CGATGCGTTACC ATCGATCAGCTG CGGCGTTAAGCT GAAGCGCGTATT GATGACCAACCA CGCGCTAAAGCA TGGCGGCGGCAA TCGATGCTGGAT CTTTAAGAAGCA GAGGCGCGTCGT GTTTGCGCGCGC ATCGATATCCAG TGATAAATGGGG CAGGTTGAAGCG CGCTCGCGTTGC AAATAA GCTGGTGCTGGG CGAATCAGAAAT CGCCGACGGAAA CGTGGTAGTGAA AGATTTACGCTC AGGTGAGCAAAC TACCGTAACGCA GGATAGCGTTGC TGCGCATTTGCG CACACTTCTGGG TTAA
TABLE-US-00008 Table of Strains First Current Universal Mutagenic DNA Gene 1 Gene 2 Sort Reference Name Name Lineage Description Genotype mutation mutation 1 Application CI006 CI006 Isolated None WT text strain from Enterobacter genera 2 Application CI008 CI008 Isolated None WT text strain from Burkholderia genera 3 Application CI010 CI010 Isolated None WT text strain from Klebsiella genera 4 Application CI019 CI019 Isolated None WT text strain from Rahnella genera 5 Application CI028 CI028 Isolated None WT text strain from Enterobacter genera 6 Application CI050 CI050 Isolated None WT text strain from Klebsiella genera 7 Application CM002 CM002 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID text CI050 gene with a NO: 33 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 8 Application CM011 CM011 Mutant of Disruption of nifL .DELTA.nifL::SpecR SEQ ID text CI019 gene with a NO: 34 spectinomycin resistance expression cassette (SpecR) encoding the streptomycin 3''-O- adenylyltransferase gene aadA inserted. 9 Application CM013 CM013 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID text CI006 gene with a NO: 35 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 10 FIG. 4A CM004 CM004 Mutant of Disruption of amtB .DELTA.amtB::KanR SEQ ID CI010 gene with a NO: 36 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 11 FIG. 4A CM005 CM005 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID CI010 gene with a NO: 37 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 12 FIG. 4B CM015 CM015 Mutant of Disruption of nifL .DELTA.nifL::Prm5 SEQ ID CI006 gene with a fragment NO: 38 of the region upstream of the ompX gene inserted (Prm5). 13 FIG. 4B CM021 CM021 Mutant of Disruption of nifL .DELTA.nifL::Prm2 SEQ ID CI006 gene with a fragment NO: 39 of the region upstream of an unanotated gene and the first 73 bp of that gene inserted (Prm2). 14 FIG. 4B CM023 CM023 Mutant of Disruption of nifL .DELTA.nifL::Prm4 SEQ ID CI006 gene with a fragment NO: 40 of the region upstream of the acpP gene and the first 121 bp of the acpP gene inserted (Prm4). 15 FIG. 10A CM014 CM014 Mutant of Disruption of nifL .DELTA.nifL::Prm1 SEQ ID CI006 gene with a fragment NO: 41 of the region upstream of the lpp gene and the first 29 bp of the lpp gene inserted (Prm1). 16 FIG. 10A CM016 CM016 Mutant of Disruption of nifL .DELTA.nifL::Prm9 SEQ ID CI006 gene with a fragment NO: 42 of the region upstream of the lexA 3 gene and the first 21 bp of the lexA 3 gene inserted (Prm9). 17 FIG. 10A CM022 CM022 Mutant of Disruption of nifL .DELTA.nifL::Prm3 SEQ ID CI006 gene with a fragment NO: 43 of the region upstream of the mntP 1 gene and the first 53 bp of the mntP 1 gene inserted (Prm3). 18 FIG. 10A CM024 CM024 Mutant of Disruption of nifL .DELTA.nifL::Prm7 SEQ ID CI006 gene with a fragment NO: 44 of the region upstream of the sspA gene inserted (Prm7). 19 FIG. 10A CM025 CM025 Mutant of Disruption of nifL .DELTA.nifL::Prm10 SEQ ID CI006 gene with a fragment NO: 45 of the region upstream of the hisS gene and the first 52 bp of the hisS gene inserted (Prm10). 20 FIG. 10B CM006 CM006 Mutant of Disruption of glnB .DELTA.glnB::KanR SEQ ID CI010 gene with a NO: 46 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 21 FIG. 10C CI028 CM017 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID nifL:KanR CI028 gene with a NO: 47 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 22 FIG. 10C CI019 CM011 Mutant of Disruption of nifL .DELTA.nifL::SpecR SEQ ID nifL:SpecR CI019 gene with a NO: 48 spectinomycin resistance expression cassette (SpecR) encoding the streptomycin 3''-O- adenylyltransferase gene aadA inserted. 23 FIG. 10C CI016 CM013 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID nifL:KanR CI006 gene with a NO: 49 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 24 FIG. 10C CI010 CM005 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID nifL:KanR CI010 gene with a NO: 50 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 25 FIG. 4C Strain 2 CI006 Isolated None WT strain from Enterobaeter genera 26 FIG. 4C Strain 4 CI010 Isolated None WT strain from Klebsiella genera 27 FIG. 4C Strain 1 CI019 Isolated None WT strain from Rahnella genera 28 FIG. 4C Strain 3 CI028 Isolated None WT strain from Enterobacter genera 29 FIG. 4B Strain 2 CI006 Isolated None WT strain from Enterobacter genera 30 FIG. 4B High CM014 Mutant of Disruption of nifL .DELTA.nifL::Prm1 SEQ ID CI006 gene with a fragment NO: 51 of the region upstream of the lpp gene and the first 29 bp of the lpp gene inserted (Prm1). 31 FIG. 4B Med CM015 Mutant of Disruption of nifL .DELTA.nifL::Prm5 SEQ ID CI006 gene with a fragment NO: 52 of the region upstream of the ompX gene inserted (Prm5). 32 FIG. 4B Low CM023 Mutant of Disruption of nifL .DELTA.nifL::Prm4 SEQ ID CI006 gene with a fragment NO: 53 of the region upstream of the acpP gene and the first 121 bp of the acpP gene inserted (Prm4). 33 FIG. 4D Strain 2 CI006 Isolated None WT strain from Enterobacter genera 34 FIG. 4D Evolved CM029 Mutant of Disruption of nifL .DELTA.nifL::Prm5 SEQ ID SEQ ID CI006 gene with a fragment .DELTA.glnE- NO: 54 NO: 61 of the region AR_KO1 upstream of the ompX gene inserted (Prm5) and deletion of the 1287 bp after the start codon of the glnE gene containing the adenylyl- removing domain of glutamate-ammonia- ligase adenylyltransferase (.DELTA.glnE-AR_KO1). 35 FIG. 14C Wild CI006 Isolated None WT strain from Enterobacter genera 36 FIG. 14C Evolved CM014 Mutant of Disruption of nifL .DELTA.nifL::Prm1
SEQ ID CI006 gene with a fragment NO: 55 of the region upstream of the lpp gene and the first 29 bp of the lpp gene inserted (Prm1). 37 FIG. 14B Wild CI019 Isolated None WT strain from Rahnella genera 38 FIG. 14B Evolved CM011 Mutant of Disruption of nifL .DELTA.nifL::SpecR SEQ ID CI019 gene with a NO: 56 spectinomycin resistance expression cassette (SpecR) encoding the streptomycin 3''-O- adenylyltransferase gene aadA inserted. 39 FIG. 14A Evolved CM011 Mutant of Disruption of nifL .DELTA.nifL::SpecR SEQ ID CI019 gene with a NO: 57 spectinomycin resistance expression cassette (SpecR) encoding the streptomycin 3''-O- adenylyltransferase gene aadA inserted. 40 FIG. 15A Wild CI006 Isolated None WT strain from Enterobacter genera 41 FIG. 15A Evolved CM013 Mutant of Disruption of nifL .DELTA.nifL::KanR SEQ ID CI006 gene with a NO: 58 kanamycin resistance expression cassette (KanR) encoding the aminoglycoside O- phosphotransferase gene aph1 inserted. 42 FIG. 15B No CM011 Mutant of Disruption of nifL .DELTA.nifL::SpecR SEQ ID name CI019 gene with a NO: 59 spectinomycin resistance expression cassette (SpecR) encoding the streptomycin 3''-O- adenylyltransferase gene aadA inserted. 43 FIG. 16B Strain 5 CI008 Isolated None WT strain from Burkholderia genera 44 FIG. 16B Strain 1 CM011 Mutant of Disruption of nifL .DELTA.nifL::SpecR SEQ ID CI019 gene with a NO: 60 spectinomycin resistance expression cassette (SpecR) encoding the streptomycin 3''-O- adenylyltransferase gene aadA inserted.
TABLE-US-00009 Table of Strains sequences SEQ ID NO: Sequence 33 ATGAGCCATATTCAACGGGAAACGTCTTGCTCCAGGCCGCGATTAAATT CCAACATGGATGCTGATTTATATGGGTATAAATGGGCTCGCGATAATGT CGGGCAATCAGGTGCGACAATCTATCGATTGTATGGGAAGCCCGATGCG CCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTCTCCAATGATGTTA CAGATGAGATGGTCAGACTAAACTGGCTGACGGAATTTATGCCTCTTCC GACCATCAAGCATTTTATCCGTACTCCTGATGATGCATGGTTACTCACCA CTGCGATCCCCGGGAAAACAGCATTCCAGGTATTAGAAGAATATCCTGA TTCAGGTGAAAATATTGTTGATGCGCTGGCAGTGTTCCTGCGCCGGTTGC ATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGCGTATTTCGTC TCGCTCAGGCGCAATCACGAATGAATAACGGTTTGGTTGATGCGAGTGA TTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAAGAA ATGCATAAGCTTTTGCCATTCTCACCGGATTCAGTCGTCACTCATGGTGA TTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTTGTA TTGATGTTGGACGAGTCGGAATCGCAGACCGATACCAGGATCTTGCCAT CCTATGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGCTTT TTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTTCAT TTGATGCTCGATGAGTTTTTCTAATAAGCCTGCCTGGTTCTGCGTTTCCC GCTCTTTAATACCCTGACCGGAGGTGAGCAATGA 34 ATGAGCATCACGGCGTTATCAGCATCATTTCCTGAGGGGAATATCGCCA GCCGCTTGTCGCTGCAACATCCTTCACTGTTTTATACCGTGGTTGAACAA TCTTCGGTGGCGAGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGC TCGCCCCCTCGTCCCGACACTTCCAGATCGCCATAGCGCACAGCGCCTC GAGCGGTGGTAACGGCGCAGTGGCGGTTTTCATGGCTTGTTATGACTGT TTTTTTGGGGTACAGTCTATGCCTCGGGCATCCAAGCAGCAAGCGCGTT ACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGATGTTACGCA GCAGGGCAGTCGCCCTAAAACAAAGTTAAACATCATGAGGGAAGCGGT GATCGCCGAAGTATCGACTCAACTATCAGAGGTAGTTGGCGTCATCGAG CGCCATCTCGAACCGACGTTGCTGGCCGTACATTTGTACGGCTCCGCAG TGGATGGCGGCCTGAAGCCACACAGTGATATTGATTTGCTGGTTACGGT GACCGTAAGGCTTGATGAAACAACGCGGCGAGCTTTGATCAACGACCTT TTGGAAACTTCGGCTTCCCCTGGAGAGAGCGAGATTCTCCGCGCTGTAG AAGTCACCATTGTTGTGCACGACGACATCATTCCGTGGCGTTATCCAGCT AAGCGCGAACTGCAATTTGGAGAATGGCAGCGCAATGACATTCTTGCAG GTATCTTCGAGCCAGCCACGATCGACATTGATCTGGCTATCTTGCTGACA AAAGCAAGAGAACATAGCGTTGCCTTGGTAGGTCCAGCGGCGGAGGAA CTCTTTGATCCGGTTCCTGAACAGGATCTATTTGAGGCGCTAAATGAAA CCTTAACGCTATGGAACTCGCCGCCCGACTGCTGCTGGCGATGAGCGAAA TGTAGTGCTTACGTTGTCCCGCATTTGGTACAGCGCAGTAACCGGCAAA ATCGCGCCGAAGGATGTCGCTGCCGACTGGGCAATGGAGCGCCTGCCGG CCCAGTATCAGCCCGTCATACTTGAAGCTAGACAGGCTTATCTTGGACA AGAAGAAGATCGCTTGGCCTCGCGCGCAGATCAGTTGGAAGAATTTGTC CACTACGTGAAAGGCGAGATCACCAAGGTAGTCGGCAAATAATGTCTA ACAATTCGTTCAAGCCGACGCCGCTTCGCGGCGCGGCTTAACTCAAGCG TTAGATGCACTAAGCACATAATTGCTCACAGCCAAACTATCAGGTCAAG TCTGCTTTTATTATTTTTAAGCGTGCATAATAAGCCCTACACAAATGGTA CCCGACCGGTGGTGAATTTAATCTCGCTGACGTGTAGACATTCCCTTATC CAGACGCTGATCGCCCATCATCGCGGTTCTTTAGATCTCTCGGTCCGCCC TGATGGCGGCACCTTGCTGACGTTACGCCTGCCGGTACAGCAGGTTATC ACCGGAGGCTTAAAATGA 35 CTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGTT GTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCAGGCCGCGATTAAAT TCCAACATGGATGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGATTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCAGACTAAACTGGCTGACGGAATTTATGCCTCTTC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCATGGTTACTCACC ACTGCGATCCCCGGGAAAACAGCATTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTGGCAGTGTTCCTGCGCCGGTT GCATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGCGTATTTC GTCTCCTCTCAGGCGCAATCACGAATGAATAACGGTTTGGTTGATGCGAG TGATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAA GAAATGCATAAGCTTTTGCCATTCTCACCGGATTCAGTCGTCACTCATGG TGATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTT GTATTGATGTTGGACGAGTCGGAATCGCAGACCGATACCAGGATCTTGC CATCCTATGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGC TTTTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTT CATTTGATGCTCGATGAGTTTTTCTAATAAGCCTTGACCCTACGATTCCC GCTATTTCATTCACTGACCGGAGGTTCAAAATGA 36 ATGAAGATAGCAACAATGAAAACAGGTCTGGGAGCGTTGGCTCTTCTTC CCTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGTT GTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCCGTCCGCGCTTAAAC TCCAACATGGACGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGCTTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCCGTCTCAACTGGCTGACGGAGTTTATGCCTCTCC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCGTGGTTACTCACC ACCGCGATTCCTGGGAAAACAGCCTTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTTTTGATGCGCTGGCCGTGTTCCTGCGCCGGTTA CATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGTGTATTTCGT CTTGCTCAGGCGCAATCACGCATGAATAACGGTTTGGTTGATGCGAGTG ATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAAGA AATGCACAAGCTCTTGCCATTCTCACCGGATTCAGTCGTCACTCATGGTG ATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTTGT ATTGATGTTGGACGGGTCGGAATCGCAGACCGTTACCAGGACCTTGCCA TTCTTTGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGCTT TTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTTCA TTTGATGCTCGATGAGTTTTTCTAATAAGCCTGTGAAGGGCTGGACGTA AACAGCCACGGCGAAAACGCCTACAACGCCTGA 37 ATGACCCTGAATATGATGCTCGATAACGCCGTACCCGAGGCGATTGCCG GCTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGT TGTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCCGTCCGCGCTTAAAC TCCAACATGGACGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGCTTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCCGTCTCAACTGGCTGACGGAGTTTATGCCTCTCC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCGTGGTTACTCACC ACCGCGATTCCTGGGAAAACAGCCTTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTGGCCGTGTTCCTGCGCCGGTTA CATTCGATTCCTCTTTTGTAATTGTCCTTTTAACAGCGATCGTGTATTTCGT CTTGCTCAGGCGCAATCACGCATGAATAACGGTTTGGTTGATGCGAGTG ATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAAGA AATGCACAAGCTCTTGCCATTCTCACCGGATTCAGTCGTCACTCATGGTG ATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTTGT ATTGATGTTGGACGGGTCGGAATCGCAGACCGTTACCAGGACCTTGCCA TTCTTTGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGCTT TTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTTCA TTTGATGCTCGATGAGTTTTTCTAATAAGCCTTGGTTCTGCGTTTCCCGCT CTTTAATACCCTGACCGGAGGTGAGCAATGA 38 ATGACCCTGAATATGATGATGGATGCCGGCGGACATCATCGCGACAAAC AATATTAATACCGGCAACCACACCGGCAATTTACGAGACTGCGCAGGCA TCCTTTCTCCCGTCAATTTCTGTCAAATAAAGTAAAAGAGGCAGTCTACT TGAATTACCCCCGGCTGGTTGAGCGTTTGTTGAAAAAAAGTAACTGAAA AATCCGTAGAATAGCGCCACTCTGATGGTTAATTAACCTATTCAATTAA GAATTATCTGGATGAATGTGCCATTAAATGCGCAGCATAATGGTGCGTT GTGCGGGAAAACTGCTTTTTTTTGAAAGGGTTGGTCAGTAGCGGAAACA ACTCACTTCACACCCCGAAGGGGGAAGTTGCCTGACCCTACGATTCCCG CTATTTCATTCACTGACCGGAGGTTCAAAATGA 39 ATGACCCTGAATATGATGATGGATGCCGGCTCACCACGGCGATAACCAT AGGTTTTCGGCGTGGCCACATCCATGGTGAATCCCACTTTTTCCAGCACG CGCGCCACTTCATCGGGTCTTAAATACATAGATTTTCCTCGTCATCTTTC CAAAGCCTCGCCACCTTACATGACTGAGCATGGACCGTGACTCAGAAAA TTCCACAAACGAACCTGAAAGGCGTGATTGCCGTCTGGCCTTAAAAATT ATGGTCTAAACTAAAATTTACATCGAAAACGAGGGAGGATCCTATGTTT AACAAACCGAATCGCCGTGACGTAGATGAAGGTGTTGAGGATATTAACC ACGATGTTAACCAGCTCGAACTCACTTCACACCCCGAAGGGGGAAGTTG CCTGACCCTACGATTCCCGCTATTTCATTCACTGACCGGAGGTTCAAAAT GA 40 ATGACCCTGAATATGATGATGGATGCCGGCTGACGAGGCAGGTTACATC ACTGGTGAAACCCTGCACGTCAATGGCGGAATGTATATGGTTTAACCAC GATGAAAATTATTTGCGTTATTAGGGCGAAAGGCCTCAAAATAGCGTAA AATCGTGGTAAGAACTGCCGGGATTTAGTTGCAAATTTTTCAACATTTTA TACACTACGAAAACCATCGCGAAAGCGAGTTTTGATAGGAAATTTAAGA GTATGAGCACTATCGAAGAACGCGTTAAGAAAATTATCGGCGAACAGCT GGGCGTTAAGCAGGAAGAAGTTACCAACAATGCTTCCTTCGTTGAAGAC CTGGGCGCTGATTCTCTTGACACCGAACTCACTTCACACCCCGAAGGGG GAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACTGACCGGAGGT TCAAAATGA 41 ATGACCCTGAATATGATGATGGATGCCGGCCGTCCTGTAATAATAACCG GACAATTCGGACTGATTAAAAAAGCGCCCTTGTGGCGCTTTTTTTATATT CCCGCCTCCATTTAAAATAAAAAATCCAATCGGATTTCACTATTTAAACT GGCCATTATCTAAGATGAATCCGATGGAAGCTCGCTGTTTTAACACGCG TTTTTTAACCTTTTATTGAAAGTCGGTGCTTCTTTGAGCGAACGATCAAA TTTAAGTGGATTCCCATCAAAAAAATATTCTCAACCTAAAAAAGTTTGT GTAATACTTGTAACGCTACATGGAGATTAACTCAATCTAGAGGGTATTA ATAATGAATCGTACTAAACTGGTACTGGGCGCAACTCACTTCACACCCC GAAGGGGGAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACTGA CCGGAGGTTCAAAATGA 42 ATGACCCTGAATATGATGATGGATGCCGGCATATTGACACCATGACCYCG CGTAATGCTGATTGGTTCTGTGACGCTGGTAATGATTGTCGAAATTCTGA ACAGTGCCATCGAAGCCGTAGTAGACCGTATTGGTGCAGAATTCCATGA ACTTTCCGGGCGGGCGAAGGATATGGGGTCCTGCGGCGGTGCTGATGTCC ATCCTGCTGGCGATGTTTACCTGGATCGCATTACTCTGGTCACATTTTCG ATAACGCTTCCAGAATTCGATAACGCCCTGGTTTTTTGCTTAAATTTGGT TCCAAAATCGCCTTTAGCTGTATATACTCACAGCATAACTGTATATACAC CCAGGGGGCGGGATGAAAGCATTAACGGCCAGGAACTCACTTCACACC CCGAAGGGGGAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACT GACCGGAGGTTCAAAATGA 43 ATGACCCTGAATATGATGATGGATGCCGGCATCATATTGCGCTCCCTGG TTATCATTTGTTACTAAATGAAATGTTATAATATAACAATTATAAATACC ACATCGCTTTCAATTCACCAGCCAAATGAGAGGAGCGCCGTCTGACATA GCCAGCGCTATAAAACATAGCATTATCTATATGTTTATGATTAATAACTG ATTTTTGCGTTTTGGATTTGGCTGTGGCATCCTTGCCGCTCTTTTCGCAGC GTCTGCGTTTTTGCCCTCCGGTCAGGGCATTTAAGGGTCAGCAATGAGTT TTTACGCAATTACGATTCTTGCCTTCGGCATGTCGATGGATGCTTTAACT CACTTCACACCCCGAAGGGGGAAGTTGCCTGACCCTACGATTCCCGCTA TTTCATTCACTGACCGGAGGTTCAAAATGA 44 ATGACCCTGAATATGATGATGGATGCCGGCCGCGTCAGGTTGAACGTAA AAAAGTCGGTCTGCGCAAAGCACGTCGTCGTCCGCAGTTCTCCAAACGT TAATTGGTTTCTGCTTCGGCAGAACGATTGGCGAAAAAACCCGGTGCGA ACCGGGTTTTTTTATGGATAAAGATCGTGTTATCCACAGCAATCCATTGA TTATCTCTTCTTTTTCAGCATTTCCAGAATCCCCTCACCACAAAGCCCGC AAAATCTGGTAAACTATCATCCAATTTTCTGCCCAAATGGCTGGGATTGT TCATTTTTTGTTTGCCTTACAACGAGAGTGACAGTACGCGCGGGTAGTTA ACTCAACATCTGACCGGTCGATAACTCACTTCACACCCCGAAGGGGGAA GTTGCCTGACCCTACGATTCCCGCTATTTCATTCACTGACCGGAGGTTCA AAATGA 45 ATGACCCTGAATATGATGATGGATGCCGGCCCTGTATGAAGATGGCGTG CGCAAAGATCGCCTGGATAACAGCGATATGATTAGCCAGCTTGAAGCCC GCATTCGCGCGAAAGCGTCAATGCTGGACGAAGCGCGTCGTATTCGATGT GCAACAGGTAGAAAAATAAGGTTGCTGGGAAGCGGCAGGCTTCCCGTG TATGATGAACCCGCCCGGCGCGACCCGTTGTTCGTCGCGGCCCCGAGGG TTCATTTTTTGTATTAATAAAGAGAATAAACGTGGCAAAAAATATTCAA GCCATTCGCGGCATGAACGATTATCTGCCTGGCGAACTCACTTCACACC CCGAAGGGGGAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACT GACCGGAGGTTCAAAATGA 46 ATGAAAAAGATTGATGCGATTATTAAACCTTTCAAACTGGATGACGTGC GCTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGT TGTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCCGTCCGCGCTTAAAC TCCAACATGGACGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGCTTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCCGTCTCAACTGGCTGACGGAGTTTATGCCTCTCC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCGTGGTTACTCACC ACCGCGATTCCTGGGAAAACAGCCTTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTCTGCCGTGTTCCTGCGCCGGTTA CATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGTGTATTTCGT CTTGCTCAGGCGCAATCACGCATGAATAACGGTTTGGTTGATGCGAGTG ATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAAGA AATGCACAAGCTCTTGCCATTCTCACCGGATTCAGTCGTCACTCATGGTG ATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTTGT ATTGATGTTGGACGGGTCGGAATCGCAGACCGTTACCAGGACCTTGCCA TTCTTTGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGCTT TTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTTCA TTTGATGCTCGATGAGTTTTTCTAATAAGCCTCGCGCGTGATTCGTATCC GCACCGGCGAAGAAGACGACGCGGCGATTTAA 47 ATGACCATGAACCTGATGACGGATGTCGTCTCAGCCACCGGGATCGCCG GGTTGCTTTCACGACAACACCCGACGCTGTTTTTTACACTAATTGAACAG GCCCCCGTGGCGATCACGCTGACGGATACCGCTGCCCGCATTGTCTATG CCAACCCGGGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGCTCGC CTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGTT GTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCAGGCCGCGATTAAAT TCCAACATGGATGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TTCGGGCAATCAGGTGCGACAATCTATCGATTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCAGACTAAACTGGCTGACGGAATTTATGCCTCTTC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCATGGTTACTCACC ACTGCGATCCCCGGGAAAACAGCATTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTGGCAGTGTTCCTGCGCCGGTT GCATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGCGTATTTC GTCTCGCTCAGGCGCAATCACGAATGAATAACGGTTTGGTTGATGCGAG
TGATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAA GAAATGCATAAGCTTTTGCCATTCTCACCGGATTCAGTCGTCACTCATGG TGATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTT GTATTGATGTTGGACGAGTCGGAATCGCAGACCGATACCAGGATCTTGC CATCCTATGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGC TTTTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTT CATTTGATGCTCGATGAGTTTTTCTAATAAGCCTGACCGGTGGTGAATTT AATCTCGCTGACGTGTAGACATTCATCGATCTGCATCCACGGTCCGGCG GCGGTACCTGCCTGACGCTACGTTTACCGCTCTTTTATGAACTGACCGGA GGCCCAAGATGA 48 ATGAGCATCACGCCGTTATCAGCATCATTTCCTGAGGGGAATATCGCCA GCCGCTTGTCGCTGCAACATCCTTCACTGTTTTATACCGTGGTTGAACAA TCTTCGGTGGCGAGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGC TCGCCCCCTCGTCCCGACACTTCCAGATCGCCATAGCGCACAGCGCCTC GAGCGGTGGTAACGGCGCAGTGGCGGTTTTCATGGCTTGTTATGACTGT TTTTTTGGGGTACAGTCTATGCCTCGGGCATCCAAGCAGCAAGCGCGTT ACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGATGTTACGCA GCAGGGCAGTCGCCCTAAAACAAAGTTAAACATCATGAGGGAAGCGGT GATCGCCGAAGTATCGACTCAACTATCAGAGGTAGTTGGCGTCATCGAG CGCCATCTCGAACCGACGTTGCTGGCCGTACATTTGTACGGCTCCGCAG TGGATGGCGGCCTGAAGCCACACAGTGATATTGATTTGCTGGTTACGGT GACCGTAAGGCTTGATGAAACAACGCGGCGAGCTTTGATCAACGACCTT TTGGAAACTTCGGCTTCCCCTGGAGAGAGCGAGATTCTCCGCGCTGTAG AAGTCACCATTGTTGTGCACGACGACATCATTCCGTGGCGTTATCCAGCT AAGCGCGAACTGCAATTTGGAGAATGGCAGCGCAATGACATTCTTGCAG GTATCTTCGAGCCAGCCACGATCGACATTGATCTGGCTATCTTGCTGACA AAAGCAAGAGAACATAGCGTTGCCTTGGTAGGTCCAGCGGCGGAGGAA CTCTTTGATCCGGTTCCTGAACAGGATCTATTTGAGGCGCTAAATGAAA CCTTAACGCTATGGAACTCGCCGCCCGACTGGGCTGGCGATGAGCGAAA TGTAGTGCTTACGTTGTCCCGCATTTGGTACAGCGCAGTAACCGTGCAAA ATCGCGCCGAAGGATGTCGCTGCCGACTGGGCAATGGAGCGCCTGCCGG CCCAGTATCAGCCCGTCATACTTGAAGCTAGACAGGCTTATCTTGGACA AGAAGAAGATCGCTTGGCCTCGCGCGCAGATCAGTTGGAAGAATTTGTC CACTACGTGAAAGGCGAGATCACCAAGGTAGTCGGCAAATAATGTCTA ACAATTCGTTCAAGCCGACGCCGCTTCGCGGCGCGGCTTAACTCAAGCG TTAGATGCACTAAGCACATAATTGCTCACAGCCAAACTATCAGGTCAAG TCTGCTTTTATTATTTTTAAGCGTGCATAATAAGCCCTACACAAATGGTA CCCGACCGGTGGTGAATTTAATCTCGCTGACGTGTAGACATTCCCTTATC CAGACGCTGATCGCCCATCATCGCGGTTCTTTAGATCTCTCGGTCCGCCC TGATGGCGGCACCTTGCTGACGTTACGCCTGCCGGTACAGCAGGTTATC ACCGGAGGCTTAAAATGA 49 CTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGTT GTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCAGGCCGCGATTAAAT TCCAACATGGATGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGATTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCAGACTAAACTGGCTGACGGAATTTATGCCTCTTC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCATGGTTACTCACC ACTGCGATCCCCGGGAAAACAGCATTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTGGCAGTGTTCCTGCGCCGGTT GCATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGCGTATTTC GTCTCGCTCAGGCGCAATCACGAATGAATAACGGTTTGGTTGATGCGAG TGATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAA GAAATGCATAAGCTTTTGCCATTCTCACCGGATTCAGTCGTCACTCATGG TGATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTT GTATTGATGTTGGACGAGTCGGAATCGCAGACCGATACCAGGATCTTGC CATCCTATGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGC TTTTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTT CATTTGATGCTCGATGAGTTTTTCTAATAAGCCTTGACCCTACGATTCCC GCTATTTCATTCACTGACCGGAGGTTCAAAATGA 50 ATGACCCTGAATATGATGCTCGATAACGCCGTACCCGAGGCGATTGCCG GCTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGT TGTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCCGTCCGCGCTTAAAC TCCAACATGGACGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGCTTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCCGTCTCAACTGGCTGACGGAGTTTATGCCTCTCC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCGTGGTTACTCACC ACCGCGATTCCTGGGAAAACAGCCTTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTGGCCGTGTTCCTGCGCCGGTTA CATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGTGTATTTCGT CTTGCTCAGGCGCAATCACGCATGAATAACGGTTTGGTTGATGCGAGTG ATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAAGA AATGCACAAGCTCTTGCCATTCTCACCGGATTCAGTCGTCACTCATGGTG ATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTTGT ATTGATGTTGGACGGGTCGGAATCGCAGACCGTTACCAGGACCTTGCCA TTCTTTGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGCTT TTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTTCA TTTGATGCTCGATGAGTTTTTCTAATAAGCCTTGGTTCTGCGTTTCCCGCT CTTTAATACCCTGACCGGAGGTGAGCAATGA 51 ATGACCCTGAATATGATGATGGATGCCGGCCGTCCTGTAATAATAACCG GACAATTCGGACTGATTAAAAAAGCGCCCTTGTGGCGCTTTTTTTATATT CCCGCCTCCATTTAAAATAAAAAATCCAATCGGATTTCACTATTTAAACT GGCCATTATCTAAGATGAATCCGATGGAAGCTCGCTGTTTTAACACGCG TTTTTTAACCTTTTATTGAAAGTCGGTGCTTCTTTGAGCGAACGATCAAA TTTAAGTGGATTCCCATCAAAAAAATATTCTCAACCTAAAAAAGTTTGT GTAATACTTGTAACGCTACATGGAGATTAACTCAATCTAGAGGGTATTA ATAATGAATCGTACTAAACTGGTACTGGGCGCAACTCACTTCACACCCC GAAGGGGGAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACTGA CCGGAGGTTCAAAATGA 52 ATGACCCTGAATATGATGATGGATGCCGGCGGACATCATCGCGACAAAC AATATTAATACCGGCAACCACACCGGCAATTTACGAGACTGCGCAGGCA TCCTTTCTCCCGTCAATTTCTGTCAAATAAAGTAAAAGAGGCAGTCTACT TGAATTACCCCCGGCTGGTTGAGCGTTTGTTGAAAAAAAGTAACTGAAA AATCCGTAGAATAGCGCCACTCTGATGGTTAATTAACCTATTCAATTAA GAATTATCTGGATGAATGTGCCATTAAATGCGCAGCATAATGGTGCGTT GTGCGGGAAAACTGCTTTTTTTTGAAAGGGTTGGTCAGTAGCGGAAACA ACTCACTTCACACCCCGAAGGGGGAAGTTGCCTGACCCTACGATTCCCG CTATTTCATTCACTGACCGGAGGTTCAAAATGA 53 ATGACCCTGAATATGATGATGGATGCCGGCTGACGAGGCAGGTTACATC ACTGGTGAAACCCTGCACGTCAATGGCGGAATGTATATGGTTTAACCAC GATGAAAATTATTTGCGTTATTAGGGCGAAAGGCCTCAAAATAGCGTAA AATCGTGGTAAGAACTGCCGGGATTTAGTTGCAAATTTTTCAACATTTTA TACACTACGAAAACCATCGCGAAAGCGAGTTTTGATAGGAAATTTAAGA GTATGAGCACTATCGAAGAACGCGTTAAGAAAATTATCGGCGAACAGCT GGGCGTTAAGCAGGAAGAAGTTACCAACAATGCTTCCTTCGTTGAAGAC CTGGGCGCTGATTCTCTTGACACCGAACTCACTTCACACCCCGAAGGGG GAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACTGACCGGAGGT TCAAAATGA 54 ATGACCCTGAATATGATGATGGATGCCGGCGGACATCATCGCGACAAAC AATATTAATACCGGCAACCACACCGGCAATTTACGAGACTGCGCAGGCA TCCTTTCTCCCGTCAATTTCTGTCAAATAAAGTAAAAGAGGCAGTCTACT TGAATTACCCCCGGCTGGTTGAGCGTTTGTTGAAAAAAAGTAACTGAAA AATCCGTAGAATAGCGCCACTCTGATGGTTAATTAACCTATTCAATTAA GAATTATCTGGATGAATGTGCCATTAAATGCGCAGCATAATGGTGCGTT GTGCGCTGAAAACTGCTTTTTTTTGAAAGGGTTGGTCAGTAGCGGAAACA ACTCACTTCACACCCCGAAGGGGGAAGTTGCCTGACCCTACGATTCCCG CTATTTCATTCACTGACCGGAGGTTCAAAATGA 55 ATGACCCTGAATATGATGATGGATGCCGGCCGTCCTGTAATAATAACCG GACAATTCGGACTGATTAAAAAAGCGCCCTTGTGGCGCTTTTTTTATATT CCCGCCTCCATTTAAAATAAAAAATCCAATCGGATTTCACTATTTAAACT GGCCATTATCTAAGATGAATCCGATGGAAGCTCGCTGTTTTAACACGCG TTTTTTAACCTTTTATTGAAAGTCGGTGCTTCTTTGAGCGAACGATCAAA TTTAAGTGGATTCCCATCAAAAAAATATTCTCAACCTAAAAAAGTTTGT GTAATACTTGTAACGCTACATGGAGATTAACTCAATCTAGAGGGTATTA ATAATGAATCGTACTAAACTGGTACTGGGCGCAACTCACTTCACACCCC GAAGGGGGAAGTTGCCTGACCCTACGATTCCCGCTATTTCATTCACTGA CCGGAGGTTCAAAATGA 56 ATGAGCATCACGGCGTTATCAGCATCATTTCCTGAGGGGAATATCGCCA GCCGCTTGTCGCTGCAACATCCTTCACTGTTTTATACCGTGGTTGAACAA TCTTCGGTGGCGAGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGC TCGCCCCCTCGTCCCGACACTTCCAGATCGCCATAGCGCACAGCGCCTC GAGCGGTGGTAACGGCGCAGTGGCGGTTTTCATGGCTTGTTATGACTGT TTTTTTGGCTGTACAGTCTATGCCTCGGGCATCCAAGCAGCAAGCGCGTT ACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGATGTTACGCA GCAGGGCAGTCGCCCTAAAACAAAGTTAAACATCATGAGGGAAGCGGT GATCGCCGAAGTATCGACTCAACTATCAGAGGTAGTTGGCGTCATCGAG CGCCATCTCGAACCGACGTTGCTGGCCGTACATTTGTACGGCTCCGCAG TGGATGGCGCCCTGAAGCCACACAGTGATATTGATTTGCTGGTTACGGT GACCGTAAGGCTTGATGAAACAACGCGGCGAGCTTTGATCAACGACCTT TTGGAAACTTCGGCTTCCCCTGGAGAGAGCGAGATTCTCCGCGCTGTAG AAGTCACCATTGTTGTGCACGACGACATCATTCCGTGGCGTTATCCAGCT AAGCGCGAACTGCAATTTGGAGAATGGCAGCGCAATGACATTCTTGCAG GTATCTTCGAGCCAGCCACGATCGACATTGATCTGGCTATCTTGCTGACA AAAGCAAGAGAACATAGCGTTGCCTTGGTAGGTCCAGCGGCGGAGGAA CTCTTTGATCCGGTTCCTGAACAGGATCTATTTGAGGCGCTAAATGAAA CCTTAACGCTATGGAACTTCGCCGCCCGACTGGGCTGGCGATGAGCGAAA TGTAGTGCTTACGTTGTCCCGCATTTGGTACAGCGCAGTAACCGGCAAA ATCGCGCCGAAGGATGTCGCTGCCGACTGGGCAATGGAGCGCCTGCCGG CCCAGTATCAGCCCGTCATACTTGAAGCTAGACAGGCTTATCTTGGACA AGAAGAAGATCGCTTGGCCTCGCGCGCAGATCAGTTGGAAGAATTTGTC CACTACGTGAAAGGCGAGATCACCAAGGTAGTCGGCAAATAATGTCTA ACAATTCGTTCAAGCCGACGCCGCTTCGCGGCGCGGCTTAACTCAAGCG TTAGATGCACTAAGCACATAATTGCTCACAGCCAAACTATCAGGTCAAG TCTGCTTTTATTATTTTTAAGCGTGCATAATAAGCCCTACACAAATGGTA CCCGACCGGTGGTGAATTTAATCTCGCTGACGTGTAGACATTCCCTTATC CAGACGCTGATCGCCCATCATCGCGGTTCTTTAGATCTCTCGGTCCGCCC TGATGGCGGCACCTTGCTGACGTTACGCCTGCCGGTACAGCAGGTTATC ACCGGAGGCTTAAAATGA 57 ATGAGCATCACGGCGTTATCAGCATCATTTCCTGAGGGGAATATCGCCA GCCGCTTGTCGCTGCAACATCCTTCACTGTTTTATACCGTGGTTGAACAA TCTTCGGTGGCGAGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGC TCGCCCCCTCGTCCCGACACTTCCAGATCGCCATAGCGCACAGCGCCTC GAGCGGTGGTAACGGCGCAGTGGCGGTTTTCATGGCTTGTTATGACTGT TTTTTTGGGGTACAGTCTATGCCTCGGGCATCCAAGCAGCAAGCGCGTT ACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGATGTTACGCA GCAGGGCAGTCGCCCTAAAACAAAGTTAAACATCATGAGGGAAGCGGT GATCGCCGAAGTATCGACTCAACTATCAGAGGTAGTTGGCGTCATCGAG CGCCATCTCGAACCGACGTTGCTGGCCGTACATTTGTACGGCTCCGCAG TGGATGGCGGCCTGAAGCCACACAGTGATATTGATTTGCTGGTTACGGT GACCGTAAGGCTTGATGAAACAACGCGGCGAGCTTTGATCAACGACCTT TEGGAAACTTCGGCTTCCCCTGGAGAGAGCGAGATTCTCCGCGCTGTAG AAGTCACCATTGTTGTGCACGACGACATCATTCCGTGGCGTTATCCAGCT AAGCGCGAACTGCAATTTGGAGAATGGCAGCGCAATGACATTCTTGCAG GTATCTTCGAGCCAGCCACGATCGACATTGATCTGGCTATCTTGCTGACA AAAGCAAGAGAACATAGCGTTGCCTTGGTAGGTCCAGCGGCGGAGGAA CTCTTTGATCCGGTTCCTGAACAGGATCTATTTGAGGCGCTAAATGAAA CCTTAACGCTATGGAACTCGCCGCCCGACTGGGCTGGCGATGAGCGAAA TGTAGTGCTTACGTTGTCCCGCATTTGGTACAGCGCAGTAACCGGCAAA ATCGCGCCGAAGGATGTCGCTGCCGACTGGGCAATGGAGCGCCTGCCGG CCCAGTATCAGCCCGTCATACTTGAAGCTAGACAGGCTTATCTTGGACA AGAAGAAGATCGCTTGGCCTCGCGCGCAGATCAGTTGGAAGAATTTGTC CACTACGTGAAAGGCGAGATCACCAAGGTAGTCGGCAAATAATGTCTA ACAATTCGTTCAAGCCGACGCCGCTTCGCCTGCGCGGCTTAACTCAACTCG TTAGATGCACTAAGCACATAATTGCTCACAGCCAAACTATCAGGTCAAG TCTGCTTTTATTATTTTTAAGCGTGCATAATAAGCCCTACACAAATGGTA CCCGACCGGTGGTGAATTTAATCTCGCTGACGTGTAGACATTCCCTTATC CAGACGCTGATCGCCCATCATCGCGGTTCTTTAGATCTCTCGGTCCGCCC TGATGGCGGCACCTTGCTGACGTTACGCCTGCCGGTACAGCAGGTTATC ACCGGAGGCTTAAAATGA 58 CTGATCCTTCAACTCAGCAAAAGTTCGATTTATTCAACAAAGCCACGTT GTGTCTCAAAATCTCTGATGTTACATTGCACAAGATAAAAATATATCAT CATGAACAATAAAACTGTCTGCTTACATAAACAGTAATACAAGGGGTGT TATGAGCCATATTCAACGGGAAACGTCTTGCTCCAGGCCGCGATTAAAT TCCAACATGGATGCTGATTTATATGGGTATAAATGGGCTCGCGATAATG TCGGGCAATCAGGTGCGACAATCTATCGATTGTATGGGAAGCCCGATGC GCCAGAGTTGTTTCTGAAACATGGCAAAGGTAGCGTTGCCAATGATGTT ACAGATGAGATGGTCAGACTAAACTGGCTGACGGAATTTATGCCTCTTC CGACCATCAAGCATTTTATCCGTACTCCTGATGATGCATCTGTTACTCACC ACTGCGATCCCCGGGAAAACAGCATTCCAGGTATTAGAAGAATATCCTG ATTCAGGTGAAAATATTGTTGATGCGCTGGCAGTGTTCCTGCGCCGGTT GCATTCGATTCCTGTTTGTAATTGTCCTTTTAACAGCGATCGCGTATTTC GTCTCGCTCAGGCGCAATCACGAATGAATAACGGTTTGGTTGATGCGAG TGATTTTGATGACGAGCGTAATGGCTGGCCTGTTGAACAAGTCTGGAAA GAAATGCATAAGCTTTTGCCATTCTCACCGGATTCAGTCGTCACTCATGG TGATTTCTCACTTGATAACCTTATTTTTGACGAGGGGAAATTAATAGGTT GTATTGATGTTGGACGAGTCGGAATCGCAGACCGATACCAGGATCTTGC CATCCTATGGAACTGCCTCGGTGAGTTTTCTCCTTCATTACAGAAACGGC TTTTTCAAAAATATGGTATTGATAATCCTGATATGAATAAATTGCAGTTT CATTTGATGCTCGATGAGTTTTTCTAATAAGCCTTGACCCTACGATTCCC GCTATTTCATTCACTGACCGGAGGTTCAAAATGA 59 ATGAGCATCACGGCGTTATCAGCATCATTTCCTGAGGGGAATATCGCCA GCCGCTTGTCGCTGCAACATCCTTCACTGTTTTATACCGTGGTTGAACAA TCTTCGGTGGCGAGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGC TCGCCCCCTCGTCCCGACACTTCCAGATCGCCATAGCGCACAGCGCCTC GAGCGGTGGTAACGGCGCAGTGGCGGTTTTCATGGCTTGTTATGACTGT TTTTTTGGGGTACAGTCTATGCCTCGGGCATCCAAGCAGCAAGCGCGTT ACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGATGTTACGCA GCAGGGCAGTCGCCCTAAAACAAAGTTAAACATCATGAGGGAAGCGGT GATCGCCGAAGTATCGACTCAACTATCAGAGGTAGTTGGCGTCATCGAG CGCCATCTCGAACCGACGTTGCTGGCCGTACATTTGTACGGCTCCGCAG TGGATGGCGGCCTGAAGCCACACAGTGATATTGATTTGCTGGTTACGGT GACCGTAAGGCTTGATGAAACAACGCGGCGAGCTTTGATCAACGACCTT TTGGAAACTTCGGCTTCCCCTGGAGAGAGCGAGATTCTCCGCGCTGTAG AAGTCACCATTGTTGTGCACGACGACATCATTCCGTGGCGTTATCCAGCT AAGCGCGAACTGCAATTTGGAGAATGGCAGCGCAATGACATTCTTGCAG GTATCTTCGAGCCAGCCACGATCGACATTGATCTGGCTATCTTGCTGACA AAAGCAAGAGAACATAGCGTTGCCTTGGTAGGTCCAGCGGCGGAGGAA CTCTTTGATCCGGTTCCTGAACAGGATCTATTTGAGGCGCTAAATGAAA CCTTAACGCTATGGAACTCGCCGCCCGACTGGGCTGGCGATGAGCGAAA TGTAGTGCTTACGTTGTCCCGCATTTGGTACAGCGCAGTAACCGGCAAA ATCGCGCCGAAGGATGTCGCTGCCGACTGGGCAATGGAGCGCCTGCCGG CCCAGTATCAGCCCGTCATACTTGAAGCTAGACAGGCTTATCTTGGACA AGAAGAAGATCGCTTGGCCTCGCGCGCAGATCAGTTGGAAGAATTTGTC CACTACGTGAAAGGCGAGATCACCAAGGTAGTCGGCAAATAATGTCTA
ACAATTCGTTCAAGCCGACGCCGCTTCGCGGCGCGGCTTAACTCAAGCG TTAGATGCACTAAGCACATAATTGCTCACAGCCAAACTATCAGGTCAAG TCTGCTTTTATTATTTTTAAGCGTGCATAATAAGCCCTACACAAATGGTA CCCGACCGGTGGTGAATTTAATCTCGCTGACGTGTAGACATTCCCTTATC CAGACGCTGATCGCCCATCATCGCGGTTCTTTAGATCTCTCGGTCCGCCC TGATGGCGGCACCTTGCTGACGTTACGCCTGCCGGTACAGCAGGTTATC ACCCTGAGGCTTAAAATGA 60 ATGAGCATCACGGCGTTATCAGCATCATTTCCTGAGGGGAATATCGCCA GCCGCTTGTCGCTGCAACATCCTTCACTGTTTTATACCGTGGTTGAACAA TCTTCGGTGGCGAGCGTGTTGAGTCATCCTGACTAGCTGAGATGAGGGC TCGCCCCCTCGTCCCGACACTTCCAGATCGCCATAGCGCACAGCGCCTC GAGCGGTGGTAACGGCGCAGTGGCGGTTTTCATGGCTTGTTATGACTGT TTTTTTGGGGTACAGTCTATGCCTCGGGCATCCAAGCACCAAGCGCGTT ACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGATGTTACGCA GCAGGGCAGTCGCCCTAAAACAAAGTTAAACATCATGAGGGAAGCGGT GATCGCCGAAGTATCGACTCAACTATCAGAGGTAGTTGGCGTCATCGAG CGCCATCTCGAACCGACGTTGCTGGCCGTACATTTGTACGGCTCCGCAG TGGATGGCGGCCTGAAGCCACACAGTGATATTGATTTGCTGGTTACGGT GACCGTAAGGCTTGTATGAAACAACGCGGCGAGCTTTGATCAACGACCTT TTGGAAACTTCGGCTTCCCCTGGAGAGAGCGAGATTCTCCGCGCTGTAG AAGTCACCATTGTTGTGCACGACGACATCATTCCGTGGCGTTATCCAGCT AAGCGCGAACTGCAATTTGGAGAATGGCAGCGCAATGACATTCTTGCAG GTATCTTCGAGCCAGCCACGATCGACATTGATCTGGCTATCTTGCTGACA AAAGCAAGAGAACATAGCGTTGCCTTGGTAGGTCCAGCGGCGGAGGAA CTCTTTGATCCGGTTCCTGAACAGGATCTATTTGAGGCGCTAAATGAAA CCTTAACGCTATGGAACTCGCCGCCCGACTGGGCTGGCGATGAGCGAAA TGTAGTGCTTACGTTGTCCCGCATTTGGTACAGCGCAGTAACCGGCAAA ATCGCGCCGAAGGATGTCGCTGCCGACTGGGCAATGGAGCGCCTGCCGG CCCAGTATCAGCCCGTCATACTTGAAGCTAGACAGGCTTATCTTGGACA AGAAGAAGATCGCTTGGCCTCGCGCGCAGATCAGTTGGAAGAATTTGTC CACTACGTGAAAGGCGAGATCACCAAGGTAGTCGGCAAATAATGTCTA ACAATTCGTTCAAGCCGACGCCGCTTCGCGGCGCGGCTTAACTCAAGCG TTAGATGCACTAAGCACATAATTGCTCACAGCCAAACTATCAGGTCAAG TCTGCTTTTATTATTTTTAAGCGTGCATAATAAGCCCTACACAAATGGTA CCCGACCGGTGGTGAATTTAATCTCGCTGACGTGTAGACATTCCCTTATC CAGACGCTGATCGCCCATCATCGCGGTTCTTTAGATCTCTCGGTCCGCCC TGATGGCGGCACCTTGCTGACGTTACGCCTGCCGGTACAGCAGGTTATC ACCGGAGGCTTAAAATGA 61 ATGTTTAACGATCTGATTGGCGATGATGAAACGGATTCGCCGGAAGATG CGCTTTCTGAGAGCTGGCGCGAATTGTGGCAGGATGCGTTGCAGGAGGA GGATTCCACGCCCGTGCTGGCGCATCTCTCAGAGGACGATCGCCGCCGC GTGGTGGCGCTGATTGCCGATTTTCGCAAAGAGTTGGATAAACGCACCA TTGGCCCGCGAGGGCGGCAGGTACTCGATCACTTAATGCCGCATCTGCT CAGCGATGTATGCTCGCGCGACGATGCGCCAGTACCGCTGTCACGCCTG ACGCCGCTGCTCACCGGAATTATTACCCGCACCACTTACCTTGAGCTGCT AAGTGAATTTCCCGGCGCACTGAAACACCTCATTTCCCTGTGTGCCGCGT CGCCGATGGTTGCCAGTCAGCTGGCGCGCTACCCGATCCTGCTTGATGA ATTGCTCGACCCGAATACGCTCTATCAACCGACGGCGATGAATGCCTAT CGCGATGAGCTGCGCCAATACCTGCTGCGCGTGCCGGAAGATGATGAAG AGCAACAGCTTGAGGCGCTGCGGCAGTTTAAGCAGGCGCAGTTGCTGCG CGTGGCGGCGGCGGATATTGCCGGTACGTTGCCAGTAATGAAAGTGAGC GATCACTTAACCTGGCTGGCGGAAGCGATTATTGATGCGGTGGTGCAGC AAGCCTGGGGGCAGATGGTGGCGCGTTATGGCCAGCCAACGCATCTGCA CGATCGCGAAGGGCGCGCTTTTTGCGGTGGTCGGTTATGGCAAGCTGGGC GGCTGGGAGCTGGGTTACAGCTCCGATCTGGATCTGGTATTCCTGCACG ACTGCCCGATGGATGTGATGACCGATGGCGAGCGTGAAATCGATGGTCG CCAGTTCTATTTGCGTCTCGCGCAGCGCGTGATGCACCTGTTTAGCACGC GCACGTCGTCCGGCATCCTTTATGAAGTTGATGCGCGTCTGCGTCCATCT GGCGCTGCGGGGATGCTGGTCACTACTACGGAATCGTTCGCCGATTACC AGCAAAACGAAGCCTGGACGTGGGAACATCAGGCGCTGGCCCGTGCGC GCGTGGTGTACGGCGATCCGCAACTGACCGCCGAATTTGACGCCATTCG CCGCGATATTCTGATGACGCCTCGCGACGGCGCAACGCTGCAAACCGAC GTGCGAGAAATGCGCGAGAAAATGCGTGCCCATCTTGGCAACAAGCAT AAAGACCGCTTCGATCTGAAAGCCGATGAAGGCGGTATCACCGACATCG AGTTTATCGCCCAATATCTGGTGCTGCGCTTTGCCCATGACAACTCCGAA ACTGACGCGCTGGTCGGATAATGTGCGCATTCTCGAAGGGCTGGCGCAA AACCTGCATCATGGAGGAGCAGGAAGCGCAGGCATTGACGCTGGCGTAC ACCACATTGCGTGATGAGCTGCACCACCTGGCGCTGCAAGAGTTGCCGG GACATGTGGCGCTCTCCTGTTTTGTCGCCGAGCGTGCGCTTATTAAAACC AGCTGGGACAAGTGGCTGGTGGAACCGTGCGCCCCGGCGTAA
Assessment of Genetic Tractability
[0323] Candidate microbes were characterized based on transformability and genetic tractability. First, optimal carbon source utilization was determined by growth on a small panel of relevant media as well as a growth curve in both nitrogen-free and rich media. Second, the natural antibiotic resistance of each strain was determined through spot-plating and growth in liquid culture containing a panel of antibiotics used as selective markers for mutagenesis. Third, each strain was tested for its transformability through electroporation of a collection of plasmids. The plasmid collection comprises the combinatorial expansion of seven origins of replication, i.e., p15a, pSC101, CloDF, colA, RK2, pBBR1, and pRO1600 and four antibiotic resistance markers, i.e., CmR, KmR, SpecR, and TetR. This systematic evaluation of origin and resistance marker compatibility was used to identify vectors for plasmid-based mutagenesis in candidate microbes.
Example 3: Mutagenesis of Candidate Microbes
Lambda-Red Mediated Knockouts
[0324] Several mutants of candidate microbes were generated using the plasmid pKD46 or a derivative containing a kanamycin resistance marker (Datsenko et al. 2000; PNAS 97(12): 6640-6645). Knockout cassettes were designed with 250 bp homology flanking the target gene and generated via overlap extension PCR. Candidate microbes were transformed with pKD46, cultured in the presence of arabinose to induce Lambda-Red machinery expression, prepped for electroporation, and transformed with the knockout cassettes to produce candidate mutant strains. Four candidate microbes and one laboratory strain, Klebsiella oxytoca M5A1, were used to generate thirteen candidate mutants of the nitrogen fixation regulatory genes nifL, glnB, and amtB, as shown in Table 4.
TABLE-US-00010 TABLE 4 List of single knockout mutants created through Lambda-red mutagenesis Strain nifL glnB amtB M5A1 X X X CI006 X X X CI010 X X X CI019 X X CI028 X X
[0325] Oligo-Directed Mutagenesis with Cas9 Selection
[0326] Oligo-directed mutagenesis was used to target genomic changes to the rpoB gene in E. coli DH10B, and mutants were selected with a CRISPR-Cas system. A mutagenic oligo (ss1283: "G*T*T*G*ATCAGACCGATGTTCGGACCTTCcaagGTTTCGATCGGACATACGCGAC CGTAGTGGGTCGGGTGTACGTCTCGAACTTCAAAGCC" (SEQ ID NO: 2), where * denotes phosphorothioate bond) was designed to confer rifampicin resistance through a 4-bp mutation to the rpoB gene. Cells containing a plasmid encoding Cas9 were induced for Cas9 expression, prepped for electroporation, and then electroporated with both the mutagenic oligo and a plasmid encoding constitutive expression of a guide RNA (gRNA) that targets Cas9 cleavage of the WT rpoB sequence. Electroporated cells were recovered in nonselective media overnight to allow sufficient segregation of the resulting mutant chromosomes. After plating on selection for the gRNA-encoding plasmid, two out of ten colonies screened were shown to contain the desired mutation, while the rest were shown to be escape mutants generated through protospacer mutation in the gRNA plasmid or Cas9 plasmid loss.
Lambda-Red Mutagenesis with Cas9 Selection
[0327] Mutants of candidate microbes CI006 and CI010 were generated via lambda-red mutagenesis with selection by CRISPR-Cas. Knockout cassettes contained an endogenous promoter identified through transcriptional profiling (as described in Example 2 and depicted in Tables 3A-C) and .about.250 bp homology regions flanking the deletion target. CI006 and CI010 were transformed with plasmids encoding the Lambda-red recombination system (exo, beta, gam genes) under control of an arabinose inducible promoter and Cas9 under control of an IPTG inducible promoter. The Red recombination and Cas9 systems were induced in resulting transformants, and strains were prepared for electroporation. Knockout cassettes and a plasmid-encoded selection gRNA were subsequently transformed into the competent cells. After plating on antibiotics selective for both the Cas9 plasmid and the gRNA plasmid, 7 of the 10 colonies screened showed the intended knockout mutation, as shown in FIG. 3.
Example 4: In Vitro Phenotyping of Candidate Molecules
[0328] The impact of exogenous nitrogen on nitrogenase biosynthesis and activity in various mutants was assessed. The Acetylene Reduction Assay (ARA) (Temme et. al. 2012; 10908): 7085-7090) was used to measure nitrogenase activity in pure culture conditions. Strains were grown in air-tight test tubes, and reduction of acetylene to ethylene was quantified with an Agilent 6890 gas chromatograph. ARA activities of candidate microbes and counterpart candidate mutants grown in nitrogen fixation media supplemented with 0 to 10 mM glutamine are shown in FIGS. 4A-B and FIGS. 10A-C.
[0329] Under anaerobic culture conditions, a range of glutamine and ammonia concentrations was tested to quantify impact on nitrogen fixation activity. In wild-type cells, activity quickly diminished as glutamine concentrations increased. However, in a series of initial knock-out mutations, a class of mutation was validated enabling expression of nitrogen fixation genes under concentrations of glutamine that would otherwise shut off activity in wild type. This profile was generated in four different species of diazotrophs, as seen in FIG. 4C. In addition, by rewiring the regulatory network using genetic parts that have been identified, the nitrogen fixation activity level was tuned predictably. This is seen in FIG. 4B, which illustrates strains CM023, CM021, CM015, and CI006. Strain CM023 is an evolved strain low; strain CM021 is an evolved strain high; strain CM015 is an evolved strain mid; strain CI006 is a wild-type (strain 2). Ammonia excreted into culture supernatants was tested using a enzymatic-based assay (MEGAZYME). The assay measures the amount of NADPH consumed in the absorbance of 340 nm. The assay was conducted on bacterial cultures grown in nitrogen-free, anaerobic environment with a starting density of 1E9 CFU/ml. Across a panel of six evolved strains, one strain excreted up to 100 .mu.M of ammonia over a course of a 48 hour period, as seen in FIG. 4D. Further, a double mutant exhibited higher ammonia excretion than the single mutant from which it was derived, as seen in FIG. 11. This demonstrates a microbial capacity to produce ammonia in excess of its physiological needs.
Transcription Profiling of Pure Cultures
[0330] Transcriptional activity of CI006 was measured using the Nanostring Elements platform. Cells were grown in nitrogen-free media and 10E8 cells were collected after 4 hours incubation. Total RNA was extracted using the Qiagen RNeasy kit. Purified RNA was submitted to Core Diagnostics in Palo Alto, Calif., for probe hybridization and Digital Analyzer analysis, as shown in FIG. 5.
Example 5: In Planta Phenotyping of Candidate Microbes
Colonization of Plants by Candidate Microbes
[0331] Colonization of desired host plants by a candidate microbe was quantified through short-term plant growth experiments. Corn plants were inoculated with strains expressing RFP either from a plasmid or from a Tn5-integrated RFP expression cassette. Plants were grown in both sterilized sand and nonsterile peat medium, and inoculation was performed by pipetting 1 mL of cell culture directly over the emerging plant coleoptile three days post-germination. Plasmids were maintained by watering plants with a solution containing the appropriate antibiotic. After three weeks, plant roots were collected, rinsed three times in sterile water to remove visible soil, and split into two samples. One root sample was analyzed via fluorescence microscopy to identify localization patterns of candidate microbes. Microscopy was performed on 10 mm lengths of the finest intact plant roots, as shown in FIG. 6.
[0332] A second quantitative method for assessing colonization was developed. A quantitative PCR assay was performed on whole DNA preparations from the roots of plants inoculated with the endophytes. Seeds of corn (Dekalb DKC-66-40) were germinated in previously autoclaved sand in a 2.5 inch by 2.5 inch by 10 inch pot. One day after planting, 1 ml of endophyte overnight culture (SOB media) was drenched right at the spot of where the seed was located. 1 mL of this overnight culture is roughly equivalent to about 10 9 cfu, varying within 3-fold of each other, depending on which strain is being used. Each seedling was fertilized 3.times. weekly with 50 mL modified Hoagland's solution supplemented with either 2.5 mM or 0.25 mM ammonium nitrate. At four weeks after planting, root samples were collected for DNA extraction. Soil debris were washed away using pressurized water spray. These tissue samples were then homogenized using QIAGEN Tissuelyzer and the DNA was then extracted using QIAmp DNA Mini Kit (QIAGEN) according to the recommended protocol. qPCR assay was performed using Stratagene Mx3005P RT-PCR on these DNA extracts using primers that were designed (using NCBI's Primer BLAST) to be specific to a loci in each of the endophyte's genome. The presence of the genome copies of the endophytes was quantified. To further confirm the identity of the endophytes, the PCR amplification products were sequenced and are confirmed to have the correct sequence. The summary of the colonization profile of strain CI006 and CI008 from candidate microbes are presented in Table 5. Colonization rate as high as 10 7.times.cfu/g fw of root was demonstrated in strain CI008.
TABLE-US-00011 TABLE 5 Colonization of corn as measured by qPCR Strain Colonization Rate (CFU/g fw) CI006 1.45 .times. 10{circumflex over ( )}5 CI008 1.24 .times. 10{circumflex over ( )}7
In Planta RNA Profiling
[0333] Biosynthesis of nif pathway components in planta was estimated by measuring the transcription of nif genes. Total RNA was obtained from root plant tissue of CI006 inoculated plants (planting methods as described previously). RNA extraction was performed using RNEasy Mini Kit according to the recommended protocol (QIAGEN). Total RNA from these plant tissues was then assayed using Nanostring Elements kits (NanoString Technologies, Inc.) using probes that were specific to the nif genes in the genome of strain CI006. The data of nif gene expression in planta is summarized in Table 6. Expression of nifH genes was detected in plants inoculated by CM013 strains whereas nifH expression was not detectable in CI006 inoculated plants. Strain CM013 is a derivative of strain CI006 in which the nifL gene has been knocked out.
[0334] Highly expressed genes of CM011, ranked by transcripts per kilobase million (TPM), were measured in planta under fertilized condition. The promoters controlling expression of some of these highly expressed genes were used as templates for homologous recombination into targeted nitrogen fixation and assimilation loci. RNA samples from greenhouse grown CM011 inoculated plant were extracted, rRNA removed using Ribo-Zero kit, sequenced using Illumina's Truseq platform and mapped back to the genome of CM011. Highly expressed genes from CM011 are listed in Table 7.
TABLE-US-00012 TABLE 6 Expression of nifH in planta Strains Relative Transcript Expression CI006 9.4 CM013 103.25
TABLE-US-00013 TABLE 7 TPM Raw (Transcripts Gene Read Per Kilobase Gene Name Location Direction Count Million) rpsH CDS 18196-18588 reverse 4841.5 27206.4 rplQ CDS 11650-12039 reverse 4333 24536.2 rpsJ CDS 25013-25324 reverse 3423 24229 rplV CDS 21946-22278 reverse 3367.5 22333 rpsN CDS 18622-18927 reverse 2792 20150.1 rplN CDS 19820-20191 reverse 3317 19691.8 rplF CDS 17649-18182 reverse 4504.5 18628.9 rpsD CDS 13095-13715 reverse 5091.5 18106.6 rpmF CDS 8326-8493 forward 1363.5 17923.8 rplW CDS 23429-23731 reverse 2252 16413.8 rpsM CDS 14153-14509 reverse 2269 14036.2 rplR CDS 17286-17639 reverse 2243.5 13996.1 rplC CDS 24350-24979 reverse 3985 13969.2 rplK CDS 25526-25954 reverse 2648.5 13634.1 rplP CDS 20807-21217 reverse 2423 13019.5 rplX CDS 19495-19809 reverse 1824 12787.8 rpsQ CDS 20362-20616 reverse 1460.5 12648.7 bhsA 3 CDS 79720-79977 reverse 1464 12531.5 rpmC CDS 20616-20807 reverse 998.5 11485 rpoA CDS 12080-13069 reverse 4855 10830.2 rplD CDS 23728-24333 reverse 2916.5 10628.5 bhsA 1 CDS 78883-79140 reverse 1068 9141.9 rpsS CDS 22293-22571 reverse 1138.5 9011.8 rpmA CDS 2210-2467 forward 1028.5 8803.7 rpmD CDS 16585-16764 reverse 694.5 8520.8 rplB CDS 22586-23410 reverse 3132 8384 rpsC CDS 21230-21928 reverse 2574.5 8133.9 rplE CDS 18941-19480 reverse 1972.5 8066.9 rplO CDS 16147-16581 reverse 1551 7874.2 preprotein translocase 14808-16139 reverse 4657 7721.2 subunit SecY CDS rpsE CDS 16771-17271 reverse 1671.5 7368 rpsK CDS 13746-14135 reverse 1223.5 6928.2 tufA CDS 27318-28229 reverse 2850 6901.3 rpmI CDS 38574-38771 forward 615 6859.5 rplU CDS 1880-2191 forward 935.5 6621.7 rplT CDS 38814-39170 forward 1045 6464.4 bhsA 2 CDS 79293-79550 reverse 754 6454.1 rpmB CDS 8391-8627 reverse 682 6355.1 rplJ CDS 23983-24480 reverse 1408 6243.9 fusA 2 CDS 481-2595 reverse 5832 6089.6 rpsA CDS 25062-26771 reverse 4613 5957.6 rpmJ CDS 14658-14774 reverse 314 5926.9 rpsR CDS 52990-53217 forward 603 5840.7 rpsG CDS 2692-3162 reverse 1243 5828.2 rpsI CDS 11354-11746 reverse 980.5 5509.8 cspC 1 CDS 8091-8300 reverse 509 5352.8 rpsF CDS 52270-52662 forward 916 5147.4 rpsT CDS 55208-55471 reverse 602 5035.9 infC CDS 38128-38478 forward 755 4750.3 cspG CDS 30148-30360 forward 446 4624.2
.sup.15N Assay
[0335] The primary method for demonstrating fixation uses the nitrogen isotope 15N, which is found in the atmosphere at a set rate relative to 14N. By supplementing either fertilizer or atmosphere with enriched levels of 15N, one can observe fixation either directly, in heightened amounts of 15N fixed from an atmosphere supplemented with 15N2 gas (Yoshida 1980), or inversely, through dilution of enriched fertilizer by atmospheric N2 gas in plant tissues (Iniguez 2004). The dilution method allows for the observation of cumulative fixed nitrogen over the course of plant growth, while the 15N.sub.2 gas method is restricted to measuring the fixation that occurs over the short interval that a plant can be grown in a contained atmosphere (rate measurement). Therefore, the gas method is superior in specificity (as any elevated 15N.sub.2 levels in the plant above the atmospheric rate can be attributed unambiguously to fixation) but cannot show cumulative activity.
[0336] Both types of assay has been performed to measure fixation activity of improved strains relative to wild-type and uninoculated corn plants, and elevated fixation rates were observed in planta for several of the improved strains (FIG. 12, FIG. 14A, and FIG. 14B). These assays are instrumental in demonstrating that the activity of the strains observed in vitro translates to in vivo results. Furthermore, these assays allow measurement of the impact of fertilizer on strain activity, suggesting suitable functionality in an agricultural setting. Similar results were observed when setaria plants were inoculated with wild-type and improved strains (FIG. 13). In planta fixation activity shown in FIGS. 14A-14C is further backed up by transcriptomic data. Evolved strains exhibit increased nifH transcript level relative to wild-type counterparts. Furthermore, the microbe derived nitrogen level in planta is also correlated with the colonization level on a plant by plant basis. These results (FIG. 12, FIG. 13, FIGS. 14A-14C, FIG. 15A, and FIG. 15B) support the hypothesis that the microbe, through the improved regulation of the nif gene cluster, is the likely reason for the increase in atmospheric derived nitrogen seen in the plant tissue. In addition to measuring fixation directly, the impact of inoculating plants with the improved strains in a nitrogen-stressed plant biomass assay was measured. While plant biomass may be related to many possible microbe interactions with the plant, one would expect that the addition of fixed nitrogen would impact the plant phenotype when nitrogen is limited. Inoculated plants were grown in the complete absence of nitrogen, and significant increases in leaf area, shoot fresh and dry weight, and root fresh and dry weight in inoculated plants relative to untreated controls was observed (FIG. 14C). Although these differences cannot be attributed to nitrogen fixation exclusively, they support the conclusion that the improved strains are actively providing nitrogen to the plant. Corn and setaria plants were grown and inoculated as described above. Fertilizer comprising 1.2% .sup.15N was regularly supplied to plants via watering. Nitrogen fixation by microbes was quantified by measuring the .sup.15N level in the plant tissue. Fourth leaf tissue was collected and dried at 4 weeks after planting. Dried leaf samples were homogenized using beads (QIAGEN Tissuelyzer) and aliquoted out into tin capsules for IRMS (MBL Stable Isotope Laboratory at The Ecosystems Center, Woods Hole, Mass.). Nitrogen derived from the atmosphere (NDFA) was calculated, and nitrogen production by CI050 and CM002 are shown in FIG. 7.
Phytohormone Production Assay
[0337] The dwarf tomato (Solanum lycopersicum) cultivar `Micro-Tom` has previously been used to study the influence of indole-3-acetic acid on fruit ripening through an in vitro assay (Cohen 1996; J Am Soc Hortic Sci 121: 520-524). To evaluate phytohormone production and secretion by candidate microbes, a plate-based screening assay using immature Micro-Tom fruit was developed. Twelve-well tissue culture test plates were prepared by filling wells with agar medium, allowing it to solidify, and spotting 10 uL of overnight microbial cultures onto the agar surface, as shown in FIG. 8. Wells with agar containing increasing amounts of gibberellic acid (GA) but no bacterial culture were used as a positive control and standards. Flowers one day post-anthesis abscised from growing Micro-Tom plants were inserted, stem-first, into the agar at the point of the bacterial spot culture. These flowers were monitored for 2-3 weeks, after which the fruits were harvested and weighed. An increase in plant fruit mass across several replicates indicates production of plant hormone by the inoculant microbe, as shown in FIG. 9.
Example 6: Cyclical Host-Microbe Evolution
[0338] Corn plants were inoculated with CM013 and grown 4 weeks to approximately the V5 growth stage. Those demonstrating improved nitrogen accumulation from microbial sources via .sup.15N analysis were uprooted, and roots were washed using pressurized water to remove bulk soil. A 0.25 g section of root was cut and rinsed in PBS solution to remove fine soil particles and non-adherent microbes. Tissue samples were homogenized using 3 mm steel beads in QIAGEN TissueLyser II. The homogenate was diluted and plated on SOB agar media. Single colonies were resuspended in liquid media and subjected to PCR analysis of 16s rDNA and mutations unique to the inoculating strain. The process of microbe isolation, mutagenesis, inoculation, and re-isolation can be repeated iteratively to improve microbial traits, plant traits, and the colonization capability of the microbe.
Example 7: Compatibility Across Geography
[0339] The ability of the improved microbes to colonize an inoculated plant is critical to the success of the plant under field conditions. While the described isolation methods are designed to select from soil microbes that may have a close relationship with crop plants such as corn, many strains may not colonize effectively across a range of plant genotypes, environments, soil types, or inoculation conditions. Since colonization is a complex process requiring a range of interactions between a microbial strain and host plant, screening for colonization competence has become a central method for selecting priority strains for further development. Early efforts to assess colonization used fluorescent tagging of strains, which was effective but time-consuming and not scalable on a per-strain basis. As colonization activity is not amenable to straightforward improvement, it is imperative that potential product candidates are selected from strains that are natural colonizers.
[0340] An assay was designed to test for robust colonization of the wild-type strains in any given host plant using qPCR and primers designed to be strain-specific in a community sample. This assay is intended to rapidly measure the colonization rate of the microbes from corn tissue samples. Initial tests using strains assessed as probable colonizers using fluorescence microscopy and plate-based techniques indicated that a qPCR approach would be both quantitative and scalable.
[0341] A typical assay is performed as follows: Plants, mostly varieties of maize and wheat, are grown in a peat potting mix in the greenhouse in replicates of six per strain. At four or five days after planting, a 1 mL drench of early stationary phase cultures of bacteria diluted to an OD590 of 0.6-1.0 (approximately 5E+08 CFU/mL) is pipetted over the emerging coleoptile. The plants are watered with tap water only and allowed to grow for four weeks before sampling, at which time, the plants are uprooted and the roots washed thoroughly to remove most peat residues. Samples of clean root are excised and homogenized to create a slurry of plant cell debris and associated bacterial cells. We developed a high-throughput DNA extraction protocol that effectively produced a mixture of plant and bacterial DNA to use as template for qPCR. Based on bacterial cell spike-in experiments, this DNA extraction process provides a quantitative bacterial DNA sample relative to the fresh weight of the roots. Each strain is assessed using strain-specific primers designed using Primer BLAST (Ye 2012) and compared to background amplification from uninoculated plants. Since some primers exhibit off-target amplification in uninoculated plants, colonization is determined either by presence of amplification or elevated amplification of the correct product compared to the background level.
[0342] This assay was used to measure the compatibility of the microbial product across different soil geography. Field soil qualities and field conditions can have a huge influence on the effect of a microbial product. Soil pH, water retention capacity, and competitive microbes are only a few examples of factors in soil that can affect inoculum survival and colonization ability. A colonization assay was performed using three diverse soil types sampled from agricultural fields in California as the plant growth medium (FIG. 16A). An intermediate inoculation density was used to approximate realistic agricultural conditions. Within 3 weeks, Strain 5 colonized all plants at 1E+06 to 1E+07 CFU/g FW. After 7 weeks of plant growth, an evolved version of Strain 1 exhibited high colonization rates (1E+06 CFU/g FW) in all soil types. (FIG. 16B).
[0343] Additionally, to assess colonization in the complexity of field conditions, a 1-acre field trial in San Luis Obispo in June of 2015 was initiated to assess the impacts and colonization of seven of the wild-type strains in two varieties of field corn. Agronomic design and execution of the trial was performed by a contract field research organization, Pacific Ag Research. For inoculation, the same peat culture seed coating technique tested in the inoculation methods experiment was employed. During the course of the growing season, plant samples were collected to assess for colonization in the root and stem interior. Samples were collected from three replicate plots of each treatment at four and eight weeks after planting, and from all six reps of each treatment shortly before harvest at 16 weeks. Additional samples were collected from all six replicate plots of treatments inoculated with Strain 1 and Strain 2, as well as untreated controls, at 12 weeks. Numbers of cells per gram fresh weight of washed roots were assessed as with other colonization assays with qPCR and strain-specific primers. Two strains, Strain 1 and Strain 2, showed consistent and widespread root colonization that peaked at 12 weeks and then declined precipitously (FIG. 16C). While Strain 2 appeared to be present in numbers an order of magnitude lower than Strain 1, it was found in more consistent numbers from plant to plant. No strains appeared to effectively colonize the stem interior. In support of the qPCR colonization data, both strains were successfully re-isolated from the root samples using plating and 16S sequencing to identify isolates of matching sequence.
Example 8: Microbe Breeding
[0344] Examples of microbe breeding can be summarized in the schematic of FIG. 17A. FIG. 17A depicts microbe breeding wherein the composition of the microbiome can be first measured and a species of interest is identified. The metabolism of the microbiome can be mapped and linked to genetics. Afterwards, a targeted genetic variation can be introduced using methods including, but not limited to, conjugation and recombination, chemical mutagenesis, adaptive evolution, and gene editing. Derivative microbes are used to inoculate crops. In some examples, the crops with the best phenotypes are selected.
[0345] As provided in FIG. 17A, the composition of the microbiome can be first measured and a species of interest is identified. FIG. 17B depicts an expanded view of the measurement of the microbiome step. The metabolism of the microbiome can be mapped and linked to genetics. The metabolism of nitrogen can involve the entrance of ammonia (NW) from the rhizosphere into the cytosol of the bacteria via the AmtB transporter. Ammonia and L-glutamate (L-Glu) are catalyzed by glutamine synthetase and ATP into glutamine. Glutamine can lead to the formation of biomass (plant growth), and it can also inhibit expression of the nif operon. Afterwards, a targeted genetic variation can be introduced using methods including, but not limited to, conjugation and recombination, chemical mutagenesis, adaptive evolution, and gene editing. Derivative microbes are used to inoculate crops. The crops with the best phenotypes are selected.
Example 9: Field Trials with Microbes of the Disclosure--Summer 2016
[0346] In order to evaluate the efficacy of strains of the present disclosure on corn growth and productivity under varying nitrogen regimes, field trials were conducted.
[0347] Trials were conducted with (1) seven subplot treatments of six strains plus the control--four main plots comprised 0, 15, 85, and 100% of maximum return to nitrogen (MRTN) with local verification. The control (UTC only) was conducted with 10 100% MRTN plus, 5, 10, or 15 pounds. Treatments had four replications.
[0348] Plots of corn (minimum) were 4 rows of 30 feet in length, with 124 plots per location. All observations were taken from the center two rows of the plots, and all destructive sampling was taken from the outside rows. Seed samples were refrigerated until 1.5 to 2 hours prior to use.
[0349] Local Agricultural Practice:
[0350] The seed was a commercial corn without conventional fungicide and insecticide treatment. All seed treatments were applied by a single seed treatment specialist to assure uniformity. Planting date, seeding rate, weed/insect management, etc. were left to local agricultural practices. With the exception of fungicide applications, standard management practices were followed.
[0351] Soil Characterization:
[0352] Soil texture and soil fertility were evaluated. Soil samples were pre-planted for each replicate to insure residual nitrate levels lower than 50 lbs/Ac. Soil cores were taken from 0 cm to 30 cm. The soil was further characterized for pH, CEC, total K and P.
[0353] Assessments:
[0354] The initial plant population was assessed 14 days after planting (DAP)/acre, and were further assessed for: (1) vigor (1 to 10 scale, w/10=excellent) 14 DAP & V10; (2) recordation of disease ratings any time symptoms are evident in the plots; (3) record any differences in lodging if lodging occurs in the plots; (4) yield (Bu/acre), adjusted to standard moisture pet; (5) test weight; and (6) grain moisture percentage.
[0355] Sampling Requirements:
[0356] The soil was sampled at three timepoints (prior to trial initiation, V10-VT, 1 week post-harvest). All six locations and all plots were sampled at 10 grams per sample (124 plots.times.3 timepoints.times.6 locations).
[0357] Colonization Sampling:
[0358] Colonization samples were collected at two timepoints (V10 and VT) for five locations and six timepoints (V4, V8, V10, VT, R5, and Post-Harvest). Samples were collected as follows: (1) from 0% and 100% MRTN, 60 plots per location; (2) 4 plants per plot randomly selected from the outside rows; (3) 5 grams of root, 8 inches of stalk, and top three leaves--bagged and IDed each separately--12/bags per plot; (4) five locations (60 plots.times.2 timepoints.times.12 bags/plot); and one location (60 plots.times.6 timepoints.times.12 bags/plot. See, FIG. 17C illustrating colonization sampling.
[0359] Normalized difference vegetation index (NDVI) determination was made using a Greenseeker instrument at two timepoints (V4-V6 and VT). Assessed each plot at all six locations (124 plots.times.2 timepoints.times.6 locations).
[0360] Root analysis was performed with Win Rhizo from one location that best illustrated treatment differentiation. Ten plants per plot were randomly sampled (5 adjacent from each outside row; V3-V4 stage plants were preferred) and gently washed to remove as much dirt as reasonable. Ten roots were placed in a plastic bag and labelled. Analyzed with WinRhizo Root Analysis.
[0361] Stalk Characteristics were measured at all six locations between R2 and R5. The stalk diameter of ten plants per plot at the 6'' height were recorded, as was the length of the first internode above the 6'' mark. Ten plants were monitored; five consecutive plants from the center of the two inside rows. Six locations were evaluated (124 plots.times.2 measures.times.6 locations).
[0362] The tissue nitrates were analyzed front all plots and all locations. An 8'' segment of stalk beginning 6'' above the soil when the corn is between one and three weeks after black layer formation; leaf sheaths were removed. All locations and plots were evaluated (6 locations.times.124 plots).
[0363] The following weather data was recorded for all locations from planting to harvest: daily maximum and minimum temperatures, soil temperature at seeding, daily rainfall plus irrigation (if applied), and any unusual weather events such as excessive rain, wind, cold, or heat.
[0364] Yield data across all six locations is presented in Table 8. Nitrogen rate had a significant impact on yield, but strains across nitrogen rates did not. However, at the lowest nitrogen rate, strains CI006, CM029, and CI019 numerically out-yielded the UTC by 4 to 6 bu/acre. Yield was also numerically increased 2 to 4 bu/acre by strains CM029, CI019, and CM081 at 15% MRTN.
TABLE-US-00014 TABLE 8 Yield data across all six locations Stalk Vigor_ Vigor_ Diameter Internode NDVI_ NDVI_ MRTN % YLD (bu) E L (mm) Length (in) Veg Rep 0 143.9 7.0 5.7 18.87 7.18 64.0 70.6 15 165.9 7.2 6.3 19.27 7.28 65.8 72.5 85 196.6 7.1 7.1 20.00 7.31 67.1 74.3 100 197.3 7.2 7.2 20.23 7.37 66.3 72.4 Stalk Vigor_ Vigor_ Diameter Internode NDVI_ NDVI_ Strain YLD (bu) E L (mm) Length (in) Veg Rep CI006 (1) 176.6 7.2 6.6 19.56 18.78 66.1 72.3 CM029 (2) 176.5 7.1 6.5 19.54 18.61 65.4 71.9 CM038 (3) 175.5 7.2 6.5 19.58 18.69 65.7 72.8 CI019 (4) 176.0 7.1 6.6 19.51 18.69 65.5 72.9 CM081 (5) 176.2 7.1 6.6 19.57 18.69 65.8 73.1 CM029/ 174.3 7.1 6.6 19.83 18.79 66.2 72.5 CM081(6) UTC (7) 176.4 7.1 6.6 19.54 18.71 65.9 71.7 Stalk MRTN/ Vigor_ Vigor_ Diameter Internode NDVI_ NDVI_ Strain YLD (bu) E L (mm) Length (in) Veg Rep 0 1 145.6 7.0 5.6 19.07 7.12 63.5 70.3 0 2 147.0 7.0 5.5 18.74 7.16 64.4 70.4 0 3 143.9 7.0 5.5 18.83 7.37 64.6 70.5 0 4 146.0 6.9 5.7 18.86 7.15 63.4 70.7 0 5 141.7 7.0 5.8 18.82 7.05 63.6 70.9 0 6 142.2 7.2 5.8 19.12 7.09 64.7 69.9 0 7 141.2 7.0 5.8 18.64 7.32 64.0 71.4 15 1 164.2 7.3 6.1 19.09 7.21 66.1 71.5 15 2 167.3 7.2 6.3 19.32 7.29 65.5 72.7 15 3 165.6 7.3 6.3 19.36 7.23 64.8 72.5 15 4 167.9 7.3 6.4 19.31 7.51 66.1 72.3 15 5 169.3 7.2 6.2 19.05 7.32 66.0 72.8 15 6 161.9 7.1 6.3 19.45 7.20 66.2 72.2 15 7 165.1 7.3 6.4 19.30 7.18 66.0 73.3 85 1 199.4 7.3 7.2 19.70 7.32 67.2 74.0 85 2 195.1 7.1 7.2 19.99 7.09 66.5 74.4 85 3 195.0 7.0 7.0 20.05 7.26 67.3 74.6 85 4 195.6 7.2 7.1 20.04 7.29 66.4 74.4 85 5 196.4 7.2 7.0 19.87 7.39 67.3 74.5 85 6 195.1 7.0 6.9 20.35 7.34 67.4 74.4 85 7 199.5 6.9 7.2 19.97 7.48 67.4 74.1 100 1 197.1 7.2 7.3 20.38 7.68 67.5 73.4 100 2 196.5 7.0 7.1 20.11 7.21 65.3 70.2 100 3 197.6 7.5 7.3 20.08 7.42 66.3 73.4 100 4 194.6 7.1 7.1 19.83 7.40 66.1 74.1 100 5 197.4 7.2 7.3 20.53 7.36 66.2 74.3 100 6 198.1 7.2 7.4 20.40 7.16 66.6 73.6 100 7 199.9 7.2 7.2 20.26 7.39 66.2 68.1
[0365] Another analysis approach is presented in Table 9. The table comprises the four locations where the response to nitrogen was the greatest which suggests that available residual nitrogen was lowest. This approach does not alter the assessment that the nitrogen rate significantly impacted yield, which strains did not when averaged across all nitrogen rates. However, the numerical yield advantage at the lowest N rate is more pronounced for all strains, particularly CI006, CM029, and CM029/CM081 where yields were increased from 8 to 10 bu/acre. At 15% MRTN, strain CM081 outyielded UTC by 5 bu.
TABLE-US-00015 TABLE 9 Yield data across four locations 4 Location Average - SGS, AgIdea, Bennett, RFR Stalk Internode YLD Diameter Length (bu) Vigor_E Vigor_L (mm) (in) Table 16 MRTN % 0 137.8 7.3 5.84 18.10 5.36 15 162.1 7.5 6.63 18.75 5.40 85 199.2 7.4 7.93 19.58 5.62 100 203.5 7.5 8.14 19.83 5.65 Strain CI006 (1) 175.4 7.5 7.08 19.03 5.59 CM029 (2) 176.1 7.4 7.08 19.09 5.39 CM038 (3) 175.3 7.5 7.05 19.01 5.59 CI019 (4) 174.8 7.5 7.16 19.02 5.45 CM081 (5) 176.7 7.4 7.16 19.00 5.53 CM029/ 175.1 7.4 7.17 19.33 5.46 CM081 (6) UTC (7) 176.0 7.3 7.27 18.98 5.55 MRTN/Strain 0 1 140.0 7.3 5.69 18.32 5.28 0 2 140.7 7.4 5.69 18.19 5.23 0 3 135.5 7.3 5.63 17.95 5.50 0 4 138.8 7.3 5.81 17.99 5.36 0 5 136.3 7.3 6.06 18.05 5.34 0 6 141.4 7.5 6.00 18.43 5.30 0 7 131.9 7.3 6.00 17.75 5.48 15 1 158.0 7.6 6.44 18.53 5.34 15 2 164.1 7.5 6.56 19.13 5.42 15 3 164.3 7.6 6.63 18.68 5.51 15 4 163.5 7.6 6.81 18.84 5.34 15 5 166.8 7.5 6.63 18.60 5.39 15 6 156.6 7.4 6.56 18.86 5.41 15 7 161.3 7.5 6.81 18.62 5.42 85 1 199.4 7.6 8.00 19.15 5.63 85 2 199.0 7.4 8.09 19.49 5.46 85 3 198.2 7.4 7.75 19.88 5.69 85 4 196.8 7.4 8.00 19.65 5.60 85 5 199.5 7.4 7.75 19.26 5.70 85 6 198.7 7.3 7.81 19.99 5.61 85 7 202.8 7.2 8.13 19.66 5.65 100 1 204.3 7.4 8.19 20.11 6.10 100 2 200.6 7.3 8.00 19.53 5.46 100 3 203.3 7.7 8.19 19.55 5.67 100 4 200.2 7.6 8.00 19.59 5.49 100 5 203.9 7.4 8.19 20.08 5.68 100 6 203.8 7.5 8.31 20.05 5.52 100 7 208.1 7.4 8.13 19.90 5.63
[0366] The results from the field trial are also illustrated in FIGS. 21-27. The results indicate that the microbes of the disclosure are able to increase plant yield, which points to the ability of the taught microbes to increase nitrogen fixation in an important agricultural crop, i.e. corn.
[0367] The field based results further validate the disclosed methods of non-intergenerially modifying the genome of selected microbial strains, in order to bring about agriculturally relevant results in a field setting when applying said engineered strains to a crop.
[0368] FIG. 18 depicts the lineage of modified strains that were derived from strain CI006 (WT Kosakonia sacchari). The field data demonstrates that an engineered derivative of the CI006 WT strain, i.e. CM029, is able to bring about numerically relevant results in a field setting. For example, Table 8 illustrates that at 0% MRTN CM029 yielded 147.0 bu/acre compared to untreated control at 141.2 bu/acre (an increase of 5.8 bu/acre). Table 8 also illustrates that at 15% MRTN CM029 yielded 167.3 bu/acre compared to untreated control at 165.1 bu/acre (an increase of 2.2 bu/acre). Table 9 is supportive of these conclusions and illustrates that at 0% MRTN CM029 yielded 140.7 bu/acre compared to untreated control at 131.9 bu/acre (an increase of 8.8 bu/acre). Table 9 also illustrates that at 15% MRTN CM029 yielded 164.1 bu/acre compared to untreated control at 161.3 bu/acre (an increase of 2.8 bu/acre).
[0369] FIG. 19 depicts the lineage of modified strains that were derived from strain CI019 (WT Rahnella aquatilis). The field data demonstrates that an engineered derivative of the CI019 WT strain, i.e. CM081, is able to bring about numerically relevant results in a field setting. For example, Table 8 illustrates that at 15% MRTN CM081 yielded 169.3 bu/acre compared to untreated control at 165.1 bu/acre (an increase of 4.2 bu/acre). Table 9 is supportive of these conclusions and illustrates that at 0% MRTN CM081 yielded 136.3 bu/acre compared to untreated control at 131.9 bu/acre (an increase of 4.4 bu/acre). Table 9 also illustrates that at 15% MRTN CM081 yielded 166.8 bu/acre compared to untreated control at 161.3 bu/acre (an increase of 5.5 bu/acre).
[0370] Further, one can see in Table 9 that the combination of CM029/CM081 at 0% MRTN yielded 141.4 bu/acre compared to untreated control at 131.9 bu/acre (an increase of 9.5 bu/acre).
Example 10: Field Trials with Microbes of the Disclosure
[0371] A diversity of nitrogen fixing bacteria can be found in nature, including in agricultural soils. However, the potential of a microbe to provide sufficient nitrogen to crops to allow decreased fertilizer use may be limited by repression of nitrogenase genes in fertilized soils as well as low abundance in close association with crop roots. Identification, isolation and breeding of microbes that closely associate with key commercial crops might disrupt and improve the regulatory networks linking nitrogen sensing and nitrogen fixation and unlock significant nitrogen contributions by crop-associated microbes. To this end, nitrogen fixing microbes that associate with and colonize the root system of corn were identified.
[0372] Root samples from corn plants grown in agronomically relevant soils were collected, and microbial populations extracted from the rhizosphere and endosphere. Genomic DNA from these samples was extracted, followed by 16S amplicon sequencing to profile the community composition. A Kosakonia sacchari microbe (strain PBC6.1) was isolated and classified through 16S rRNA and whole genome sequencing. This is a particularly interesting nitrogen fixer capable of colonizing to nearly 21% abundance of the root-associated microbiota (FIG. 30). To assess strain sensitivity to exogenous nitrogen, nitrogen fixation rates in pure culture were measured with the classical acetylene reduction assay (ARA) and varying levels of glutamine supplementation. The species exhibited a high level of nitrogen fixing activity in nitrogen-free media, yet exogenous fixed nitrogen repressed nif gene expression and nitrogenase activity (Strain PBC6.1, FIGS. 28C and 28D). Additionally, when released ammonia was measured in the supernatant of PBC6.1 grown in nitrogen-fixing conditions, very little release of fixed nitrogen could be detected (FIG. 28E).
[0373] We hypothesized that PBC6.1 could be a significant contributor of fixed nitrogen in fertilized fields if regulatory networks controlling nitrogen metabolism were rewired to allow optimal nitrogenase expression and ammonia release in the presence of fixed nitrogen. Sufficient genetic diversity should exist within the PBC6.1 genome to enable broad phenotypic remodeling without the insertion of transgenes or synthetic regulatory elements. The isolated strain has a genome of at least 5.4 Mbp and a canonical nitrogen fixation gene cluster. Related nitrogen metabolism pathways in PBC6.1 are similar to those of the model organism for nitrogen fixation, Klebsiella oxytoca m5al.
[0374] Several gene regulatory network nodes were identified which may augment nitrogen fixation and subsequent transfer to a host plant, particularly in high exogenous concentrations of fixed nitrogen (FIG. 28A). The niFL operon directly regulates the rest of the nif cluster through transcriptional activation by NifA and nitrogen- and oxygen-dependent repression of NifA by NifL. Disruption of tuff, can abolish inhibition of NifA and improve nif expression in the presence of both oxygen and exogenous fixed nitrogen. Furthermore, expressing nifA under the control of a nitrogen-independent promoter may decouple nitrogenase biosynthesis from regulation by the NtrB/NtrC nitrogen sensing complex. The assimilation of fixed nitrogen by the microbe to glutamine by glutamine synthetase (GS) is reversibly regulated by the two-domain adenylyltransferase (ATase) enzyme GlnE through the adenylylation and deadenylylation of GS to attenuate and restore activity, respectively. Truncation of the GlnE protein to delete its adenylyl-removing (AR) domain may lead to constitutively adenylylated glutamine synthetase, limiting ammonia assimilation by the microbe and increasing intra- and extracellular ammonia. Finally, reducing expression of AmtB, the transporter responsible for uptake of ammonia, could lead to greater extracellular ammonia. To generate rationally designed microbial phenotypes without the use of transgenes, two approaches were employed: creating markerless deletions of genomic sequences encoding protein domains or whole genes, and rewiring regulatory networks by intragenomic promoter rearrangement. Through an iterative mutagenesis process, several non-transgenic derivative strains of PBC6.1 were generated (Table 10).
TABLE-US-00016 TABLE 10 List of isolated and derivative K. sacchari strains used in this work. Prm, promoter sequence derived from the PBC6.1 genome; .DELTA.glnE.sub.AR1 and .DELTA.glnE.sub.AR2, different truncated versions of glnE gene removing the adenylyl-removing domain sequence. Strain ID Genotype PBC6.1 WT PBC6.14 .DELTA.nifL::Prm1 PBC6.15 .DELTA.nifL::Prm5 PBC6.22 .DELTA.nifL::Prm3 PBC6.37 .DELTA.nifL::Prm1 .DELTA.glnE.sub.AR2 PBC6.38 .DELTA.nifL::Prm1 .DELTA.glnE.sub.AR1 PBC6.93 .DELTA.nifL::Prm1 .DELTA.glnE.sub.AR2 .DELTA.amtB PBC6.94 .DELTA.nifL::Prm1 .DELTA.glnE.sub.AR1 .DELTA.amtB
[0375] Several in vitro assays were performed to characterize specific phenotypes of the derivative strains. The ARA was used to assess strain sensitivity to exogenous nitrogen, in which PBC6.1 exhibited repression of nitrogenase activity at high glutamine concentrations (FIG. 28D). In contrast, most derivative strains showed a derepressed phenotype with varying levels of acetylene reduction observed at high glutamine concentrations. Transcriptional rates of nifA in samples analyzed by qPCR correlated well with acetylene reduction rates (FIG. 31), supporting the hypothesis that nifL disruption and insertion of a nitrogen-independent promoter to drive nifA can lead to nil cluster derepression. Strains with altered GlnE or AmtB activity showed markedly increased ammonium excretion rates compared to wild type or derivative strains without these mutations (FIG. 28E), illustrating the effect of these genotypes on ammonia assimilation and reuptake.
[0376] Two experiments were performed to study the interaction of PBC6.1 derivatives with corn plants and quantify incorporation of fixed nitrogen into plant tissues. First, rates of microbial nitrogen fixation were quantified in a greenhouse study using isotopic tracers. Briefly, plants are grown with 15N labeled fertilizer, and diluted concentrations of 15N in plant tissues indicate contributions of fixed nitrogen from microbes. Corn seedlings were inoculated with selected microbial strains, and plants were grown to the V6 growth stage. Plants were subsequently deconstructed to enable measurement of microbial colonization and gene expression as well as measurement of 15N/14N ratios in plant tissues by isotope ratio mass spectrometry (IRMS). Analysis of the aerial tissue showed a small, nonsignificant contribution by PBC6.38 to plant nitrogen levels, and a significant contribution by PBC6.94 (p=0.011). Approximately 20% of the nitrogen found in above-ground corn leaves was produced by PBC6.94, with the remainder coming from the seed, potting mix, or "background" fixation by other soilborne microbes (FIG. 29C). This illustrates that our microbial breeding pipeline can generate strains capable of making significant nitrogen contributions to plants in the presence of nitrogen fertilizer. Microbial transcription within plant tissues was measured, and expression of the nif gene cluster was observed in derivative strains but not the wild type strain (FIG. 29B), showing the importance of nif derepression for contribution of BNF to crops in fertilized conditions. Root colonization measured by qPCR demonstrated that colonization density is different for each of the strains tested (FIG. 29A). A 50 fold difference in colonization was observed between PBC6.38 and PBC6.94. This difference could be an indication that PBC6.94 has reduced fitness in the rhizosphere relative to PBC6.38 as a result of high levels of fixation and excretion.
Methods
Media
[0377] Minimal medium contains (per liter) 25 g Na.sub.2HPO.sub.4, 0.1 g CaCL.sub.2-2H.sub.2O, 3 g KH.sub.2PO.sub.4, 0.25 g MgSO.sub.4.7H.sub.2O, 1 g NaCl, 2.9 mg FeCl.sub.3, 0.25 mg Na.sub.2MoO.sub.4.2 H.sub.2O, and 20 g sucrose. Growth medium is defined as minimal medium supplemented with 50 ml of 200 mM glutamine per liter.
Isolation of Diazotrophs
[0378] Corn seedlings were grown from seed (DKC 66-40, DeKalb, Ill.) for two weeks in a greenhouse environment controlled from 22.degree. C. (night) to 26.degree. C. (day) and exposed to 16 hour light cycles in soil collected from San Joaquin County, Calif. Roots were harvested and washed with sterile deionized water to remove bulk soil. Root tissues were homogenized with 2 mm stainless steel beads in a tissue lyser (TissueLyser II, Qiagen P/N 85300) for three minutes at setting 30, and the samples were centrifuged for 1 minute at 13,000 rpm to separate tissue from root-associated bacteria. Supernatants were split into two fractions, and one was used to characterize the microbiome through 16S rRNA amplicon sequencing and the remaining fraction was diluted and plated on Nitrogen-free Broth (NfB) media supplemented with 1.5% agar. Plates were incubated at 30.degree. C. for 5-7 days. Colonies that emerged were tested for the presence of the nifH gene by colony PCR with primers Ueda19f and Ueda406r. Genomic DNA from strains with a positive nifH colony PCR was isolated (QIAamp DNA Mini Kit, Cat No. 51306, QIAGEN, Germany) and sequenced (Illumina MiSeq v3, SeqMatic, Fremont, Calif.). Following sequence assembly and annotation, the isolates containing nitrogen fixation gene clusters were utilized in downstream research.
Microbiome Profiling of Isolation Seedlings
[0379] Genomic DNA was isolated from root-associated bacteria using the ZR-96 Genomic DNA I Kit (Zymo Research P/N D3011), and 16S rRNA amplicons were generated using nextera-barcoded primers targeting 799f and 1114r. The amplicon libraries were purified and sequenced with the Illumina MiSeq v3 platform (SeqMatic, Fremont, Calif.). Reads were taxonomically classified using Kraken using the minikraken database (FIG. 30).
Acetylene Reduction Assay (ARA)
[0380] A modified version of the Acetylene Reduction Assay was used to measure nitrogenase activity in pure culture conditions. Strains were propagated from single colony in SOB (RPI, P/N S25040-1000) at 30.degree. C. with shaking at 200 RPM for 24 hours and then subcultured 1:25 into growth medium and grown aerobically for 24 hours (30.degree. C., 200 RPM). 1 ml of the minimal media culture was then added to 4 ml of minimal media supplemented with 0 to 10 mM glutamine in air-tight Hungate tubes and grown anaerobically for 4 hours (30.degree. C., 200 RPM). 10% headspace was removed then replaced by an equal volume of acetylene by injection, and incubation continued for 1 hr. Subsequently, 2 ml of headspace was removed via gas tight syringe for quantification of ethylene production using an Agilent 6850 gas chromatograph equipped with a flame ionization detector (FED).
Ammonium Excretion Assay
[0381] Excretion of fixed nitrogen in the form of ammonia was measured using batch fermentation in anaerobic bioreactors. Strains were propagated from single colony in 1 ml/well of SOB in a 96 well DeepWell plate. The plate was incubated at 30.degree. C. with shaking at 200 RPM for 24 hours and then diluted 1:25 into a fresh plate containing 1 ml/well of growth medium. Cells were incubated for 24 hours (30.degree. C., 200 RPM) and then diluted 1:10 into a fresh plate containing minimal medium. The plate was transferred to an anaerobic chamber with a gas mixture of >98.5% nitrogen, 1.2-1.5% hydrogen and <30 ppM oxygen and incubated at 1350 RPM, room temperature for 66-70 hrs. Initial culture biomass was compared to ending biomass by measuring optical density at 590 nm. Cells were then separated by centrifugation, and supernatant from the reactor broth was assayed for free ammonia using the Megazyme Ammonia Assay kit (P/N K-AMIAR) normalized to biomass at each timepoint.
Extraction of Root-Associated Microbiome
[0382] Roots were shaken gently to remove loose particles, and root systems were separated and soaked in a RNA stabilization solution (Thermo Fisher P/N AM7021) for 30 minutes. The roots were then briefly rinsed with sterile deionized water. Samples were homogenized using bead beating with 1/2-inch stainless steel ball bearings in a tissue lyser (TissueLyser Qiagen P/N 85300) in 2 ml of lysis buffer (Qiagen P/N 79216). Genomic DNA extraction was performed with ZR-96 Quick-gDNA kit (Zymo Research P/N D3010), and RNA extraction using the RNeasy kit (Qiagen P/N 74104).
Root Colonization Assay
[0383] Four days after planting, 1 ml of a bacterial overnight culture (approximately 10.sup.9 cfu) was applied to the soil above the planted seed. Seedlings were fertilized three times weekly with 25 ml modified Hoagland's solution supplemented with 0.5 mM ammonium nitrate. Four weeks after planting, root samples were collected and the total genomic DNA (gDNA) was extracted. Root colonization was quantified using qPCR with primers designed to amplify unique regions of either the wild type or derivative strain genome. QPCR reaction efficiency was measured using a standard curve generated from a known quantity of gDNA from the target genome. Data was normalized to genome copies per g fresh weight using the tissue weight and extraction volume. For each experiment, the colonization numbers were compared to untreated control seedlings.
In Planta Transcriptomics
[0384] Transcriptional profiling of root-associated microbes was measured in seedlings grown and processed as described in the Root Colonization Assay. Purified RNA was sequenced using the Illumina NextSeq platform (SeqMatic, Fremont, Calif.). Reads were mapped to the genome of the inoculated strain using bowtie2 using `--very-sensitive-local` parameters and a minimum alignment score of 30. Coverage across the genome was calculated using samtools. Differential coverage was normalized to housekeeping gene expression and visualized across the genome using Circos and across the nif gene cluster using DNAplotlib. Additionally, the in planta transcriptional profile was quantified via targeted Nanostring analysis. Purified RNA was processed on an nCounter Sprint (Core Diagnostics, Hayward, Calif.).
15N Dilution Greenhouse Study
[0385] 15N fertilizer dilution experiment was performed to assess optimized strain activity in planta. A planting medium containing minimal background N was prepared using a mixture of vermiculite and washed sand (5 rinses in DI 1120). The sand mixture was autoclaved for 1 hour at 122.degree. C. and approximately 600 g measured out into 40 cubic inch (656 mL) pots, which were saturated with sterile DI H.sub.2O and allowed to drain 24 hours before planting. Corn seeds (DKC 66-40) were surface sterilized in 0.625% sodium hypochlorite for 10 minutes, then rinsed five times in sterile distilled water and planted 1 cm deep. The plants were maintained under fluorescent lamps for four weeks with 16-hour day length at room temperatures averaging 22.degree. C. (night) to 26.degree. C. (day).
[0386] Five days after planting, seedlings were inoculated with a 1 ml suspension of cells drenched directly over the emerging coleoptile. Inoculum was prepared from 5 ml overnight cultures in SOB, which were spun down and resuspended twice in 5 ml PBS to remove residual SOB before final dilution to OD of 1.0 (approximately 10.sup.9 CFU/ml). Control plants were treated with sterile PBS, and each treatment was applied to ten replicate plants.
[0387] Plants were fertilized with 25 ml fertilizer solution containing 2% 15N-enriched 2 mM KNO.sub.3 on 5, 9, 14, and 19 days after planting, and the same solution without KNO.sub.3 on 7, 12, 16, and 18 days after planting. The fertilizer solution contained (per liter) 3 mmol CaCl.sub.2), 0.5 mmol KH.sub.2PO.sub.4, 2 mmol MgSO.sub.4, 17.9 .mu.mol FeSO.sub.4, 2.86 mg H.sub.3BO.sub.3, 1.81 mg MnCl.sub.2.4H.sub.2O, 0.22 mg ZnSO.sub.4.7H.sub.2O, 51 .mu.g CuSO.sub.4.5H.sub.2O, 0.12 mg Na.sub.2MoO.sub.4.2H.sub.2O, and 0.14 nmol NiCl.sub.2. All pots were watered with sterile DI H.sub.2O as needed to maintain consistent soil moisture without runoff.
[0388] At four weeks, plants were harvested and separated at the lowest node into samples for root gDNA and RNA extraction and aerial tissue for IRMS. Aerial tissues were wiped as needed to remove sand, placed whole into paper bags and dried for at least 72 hours at 60.degree. C. Once completely dry, total aerial tissue was homogenized by bead beating and 5-7 mg samples were analyzed by isotope ratio mass spectrometry (IRMS) for .delta.15N by the MBL Stable Isotope Laboratory (The Ecosystems Center, Woods Hole, Mass.). Percent NDFA was calculated using the following formula: % NDFA=(.delta.15N of UTC average--.delta.15N of sample)/(.delta.15N of UTC average).times.100.
Example 11: Field Trials with Microbes of the Disclosure--Summer 2017
[0389] In order to evaluate the efficacy of strains of the present disclosure on corn growth and productivity under varying nitrogen regimes, field trials were conducted. The below field data demonstrates that the non-intergeneric microbes of the disclosure are able to fix atmospheric nitrogen and deliver said nitrogen to a plant--resulting in increased yields--in both a nitrogen limiting environment, as well as a non-nitrogen limiting environment.
[0390] Trials were conducted at seven locations across the United states with six geographically diverse Midwestern locations. Five nitrogen regimes were used for fertilizer treatments: 100% of standard agricultural practice of the site/region, 100% minus 25 pounds, 100% minus 50 pounds, 100% minus 75 pounds, and 0%; all per acre. The pounds of nitrogen per acre for the 100% regime depended upon the standard agricultural practices of the site/region. The aforementioned nitrogen regimes ranged from about 153 pounds per acre to about 180 pounds per acre, with an average of about 164 pounds of nitrogen per acre.
[0391] Within each fertilizer regime there were 14 treatments. Each regime had six replications, and a split plot design was utilized. The 14 treatments included: 12 different microbes, 1 UTC with the same fertilizer rate as the main plot, and 1 UTC with 100% nitrogen. In the 100% nitrogen regime the 2.sup.nd UTC is 100 plus 25 pounds.
[0392] Plots of corn, at a minimum, were 4 rows of 30 feet in length (30 inches between rows) with 420 plots per location. All observations, unless otherwise noted, were taken from the center two rows of the plants, and all destructive sampling was taken from the outside rows. Seed samples were refrigerated until 1.5 to 2 hours prior to use.
[0393] Local Agricultural Practice:
[0394] The seed was a commercial corn applied with a commercial seed treatment with no biological co-application. The seeding rate, planting date, weed/insect management, harvest times, and other standard management practices were left to the norms of local agricultural practices for the regions, with the exception of fungicide application (if required).
[0395] Microbe Application:
[0396] The microbes were applied to the seed in a seed treatment over seeds that had already received a normal chemical treatment. The seed were coated with fermentation broth comprising the microbes.
[0397] Soil Characterization:
[0398] Soil texture and soil fertility were evaluated. Standard soil sampling procedures were utilized, which included soil cores of depths from 0-30 cm and 30-60 cm. The standard soil sampling included a determination of nitrate nitrogen, ammonium nitrogen, total nitrogen, organic matter, and CEC. Standard soil sampling further included a determination of pH, total potassium, and total phosphorous. To determine the nitrogen fertilizer levels, preplant soil samples from each location were taken to ensure that die 0-12'' and potentially the 12'' to 24'' soil regions for nitrate nitrogen.
[0399] Prior to planting and fertilization, 2 ml soil samples were collected from 0 to 6-12'' from the UTC. One sample per replicate per nitrogen region was collected using the middle of the row. (5 fertilizer regimes.times.6 replicates=thirty soil samples).
[0400] Post-planting (V4-V6), 2 ml soil samples were collected from 0 to 6-12'' from the UTC. One sample per replicate per nitrogen region was collected using the middle of the row. (5 fertilizer regimes.times.6 replicates=thirty soil samples).
[0401] Post-harvest (V4-V6), 2 ml soil samples were collected from 0 to 6-12'' from the UTC. One sample per replicate per nitrogen region was collected using the middle of the row. Additional post-harvest soil sample collected at 0-12'' from the UTC and potentially 12-24'' from the UTC (5 fertilizer regimes.times.6 replicates=thirty soil samples).
[0402] A V6-V10 soil sample from each fertilizer regime (excluding the treatment of 100% and 100%+25 lbs [in the 100% block] for all fertilizer regimes at 0-12'' and 12-24''. (5 fertilizer regimes.times.2 depths=10 samples per location).
[0403] Post-harvest soil sample from each fertilizer regime (excluding the treatment of 100% and 100%+25 lbs [in the 100% block] for all fertilizer regimes at 0-12'' and 12-24''. (5 fertilizer regimes.times.2 depths=10 samples per location).
[0404] Assessments:
[0405] The initial plant population was assessed at .about.50% UTC and the final plant population was assessed prior to harvest. Assessment included (1) potentially temperature (temperature probe); (2) vigor (1-10 scale with 10=excellent) at V4 and V8-V10; (3) plant height at V8-V10 and V14; (4) yield (bushels/acre) adjusted to standard moisture percentage; (5) test weight; (6) grain moisture percentage; (7) stalk nitrate tests at black layer (420 plots.times.7 locations); (8) colonization with 1 plant per plot in zip lock bag at 0% and 100% fertilizer at V4-V6 (1 plant.times.14 treatments.times.6 replicates.times.2 fertilizer regimes=168 plants); (9) transcriptomics with 1 plant per plot in zip lock bag at 0% and 100% fertilizer at V4-V6 (1 plant.times.14 treatments.times.6 replicates.times.2 fertilizer regimes=168 plants); (10) Normalized difference vegetative index (NDVI) or normalized difference red edge (NDRE) determination using a Greenseeker instrument at two time points (V4-V6 and VT) to assess each plot at all 7 locations (420 plots.times.2 time points.times.7 locations=5,880 data points); (11) stalk characteristics measured at all 7 locations between R2 and R5 by recording the stalk diameter of 10 plants/plot at 6'' height, record length of first internode above the 6'' mark, 10 plants monitored (5 consecutive plants from center of two inside rows) (420 plots.times.10 plants.times.7 locations=29,400 data points).
[0406] Monitoring Schedule:
[0407] Practitioners visited all trials at V3-V4 stage to assess early-season response to treatments and during reproductive growth stage to monitor maturity. Local cooperator visited research trial on an on-going basis.
[0408] Weather Information:
[0409] Weather data spanning from planting to harvest was collected and consisted of daily minimum and maximum temperatures, soil temperature at seeding, daily rainfall plus irrigation (if applied), and unusual weather events such as excessive wind, rain, cold, heat.
[0410] Data Reporting:
[0411] Including the data indicated above, the field trials generated data points including soil textures; row spacing; plot sizes; irrigation; tillage; previous crop; seeding rate; plant population; seasonal fertilizer inputs including source, rate, timing, and placement; harvest area dimensions, method of harvest, such as by hand or machine and measurement tools used (scales, yield monitor, etc.)
[0412] Results:
[0413] Select results from the aforementioned field trial are reported in FIG. 32 and FIG. 33.
[0414] In FIG. 32, it can be seen that a microbe of the disclosure (i.e. 6-403) resulted in a higher yield than the wild type strain (WT) and a higher yield than the untreated control (UTC). The "-25 lbs N" treatment utilizes 25 lbs less N per acre than standard agricultural practices of the region. The "100% N" UTC treatment is meant to depict standard agricultural practices of the region, in which 100% of the standard utilization of N is deployed by the farmer. The microbe "6-403" was deposited as NCMA 201708004 and can be found in Table A. This is a mutant Kosakonia sacchari (also called CM037) and is a progeny mutant strain from CI006 WT.
[0415] In FIG. 33, the yield results obtained demonstrate that the microbes of the disclosure perform consistently across locations. Furthermore, the yield results demonstrate that the microbes of the disclosure perform well in both a nitrogen stressed environment (i.e. a nitrogen limiting environment), as well as an environment that has sufficient supplies of nitrogen (i.e. a non-nitrogen-limiting condition). The microbe "6-881" (also known as CM094, PBC6.94), and which is a progeny mutant Kosakonia sacchari strain from CI006 WT, was deposited as NCMA 201708002 and can be found in Table A. The microbe "137-1034," which is a progeny mutant Klebsiella variicola strain from CI137 WI, was deposited as NCMA 201712001 and can be found in Table A. The microbe "137-1036," which is a progeny mutant Klebsiella variicola strain from CI137 WT, was deposited as NCMA 201712002 and can be found in Table A. The microbe "6-404" (also known as CM38, PBC6.38), and which is a progeny mutant Kosakonia sacchari strain from CI006 WT, was deposited as NCMA 201708003 and can be found in Table A.
Example 12: Genus of Non-Intergeneric Microbes Beneficial for Agricultural Systems
[0416] The microbes of the present disclosure were evaluated and compared against one another for the production of nitrogen produced in an acre across a season. See FIG. 20, FIG. 40, and FIG. 41
[0417] It is hypothesized by the inventors that in order for a population of engineered non-intergeneric microbes to be beneficial in a modern row crop agricultural system, then the population of microbes needs to produce at least one pound or more of nitrogen per acre per season.
[0418] To that end, the inventors have surprisingly discovered a functional genus of microbes that are able to contribute, inter alia, to: increasing yields in non-leguminous crops; and/or lessening a farmer's dependence upon exogenous nitrogen application; and/or the ability to produce at least one pound of nitrogen per acre per season, even in non-nitrogen-limiting environments, said genus being defined by the product of colonization ability.times.mmol of N produced per microbe per hour (i.e. the line partitioning FIGS. 20, 40, and 41).
[0419] With respect to FIGS. 20, 40, and 41, certain data utilizing microbes of the disclosure was aggregated, in order to depict a heatmap of the pounds of nitrogen delivered per acre-season by microbes of the disclosure, which are recorded as a function of microbes per g-fresh weight by mmol of nitrogen/microbe-hr. Below the thin line that transects the larger images are the microbes that deliver less than one pound of nitrogen per acre-season, and above the line are the microbes that deliver greater than one pound of nitrogen per acre-season.
[0420] Field Data & Wild Type Colonization Heatmap:
[0421] The microbes utilized in the FIG. 20 heatmap were assayed for N production in corn. For the WT strains CI006 and CI019, corn root colonization data was taken from a single field site. For the remaining strains, colonization was assumed to be the same as the WT field level. N-fixation activity was determined using an in vitro ARA assay at 5 mM glutamine. The table below the heatmap in FIG. 20 gives the precise value of mmol N produced per microbe per hour (mmol N/Microbe hr) along with the precise CFU per gram of fresh weight (CFU/g fw) for each microbe shown in the heatmap.
[0422] Field Data Heatmap:
[0423] The data utilized in the FIG. 40 heatmap is derived from microbial strains assayed for N production in corn in field conditions. Each point represents lb N/acre produced by a microbe using corn root colonization data from a single field site. N-fixation activity was determined using in vitro ARA assay at 5 mM N in the form of glutamine or ammonium phosphate. The below Table C gives the precise value of mmol N produced per microbe per hour (mmol N/Microbe hr) along with the precise CFU per gram of fresh weight (CFU/g fw) for each microbe shown in the heatmap of FIG. 40.
[0424] Greenhouse & Laboratory Data Heatmap:
[0425] The data utilized in the FIG. 41 heatmap is derived from microbial strains assayed for N production in corn in laboratory and greenhouse conditions. Each point represents lb N/acre produced by a single strain. White points represent strains in which corn root colonization data was gathered in greenhouse conditions. Black points represent mutant strains for which corn root colonization levels are derived from average field corn root colonization levels of the wild-type parent strain. Hatched points represent the wild type parent strains at their average field corn root colonization levels. In all cases, N-fixation activity was determined by in vitro ARA assay at 5 mM N in the form of glutamine or ammonium phosphate. The below Table D gives the precise value of mmol N produced per microbe per hour (mmol N/Microbe hr) along with the precise CFU per gram of fresh weight (CFU/g fw) for each microbe shown in the heatmap of FIG. 41.
TABLE-US-00017 TABLE C FIG. 40 - Field Data Heatmap Peak Activity Coloniza- (mmol tion N Pro- Strain N/Mi- (CFU/ duced/acre Taxonomic Name crobe hr) g fw) season Designation CI006 3.88E-16 1.50E+07 0.24 Kosakonia sacchari 6-404 1.61E-13 3.50E+05 2.28 Kosakonia sacchari 6-848 1.80E-13 2.70E+05 1.97 Kosakonia sacchari 6-881 1.58E-13 5.00E+05 3.20 Kosakonia sacchari 6-412 4.80E-14 1.30E+06 2.53 Kosakonia sacchari 6-403 1.90E-13 1.30E+06 10.00 Kosakonia sacchari CI019 5.33E-17 2.40E+06 0.01 Rahnella aquatilis 19-806 6.65E-14 2.90E+06 7.80 Rahnella aquatilis 19-750 8.90E-14 2.60E+05 0.94 Rahnella aquatilis 19-804 1.72E-14 4.10E+05 0.29 Rahnella aquatilis CI137 3.24E-15 6.50E+06 0.85 Klebsiella variicola 137-1034 1.16E-14 6.30E+06 2.96 Klebsiella variicola 137-1036 3.47E-13 1.30E+07 182.56 Klebsiella variicola 137-1314 1.70E-13 1.99E+04 0.14 Klebsiella variicola 137-1329 1.65E-13 7.25E+04 0.48 Klebsiella variicola 63 3.60E-17 3.11E+05 0.00 Rahnella aquatilis 63-1146 1.90E-14 5.10E+05 0.39 Rahnella aquatilis 1021 1.77E-14 2.69E+07 19.25 Kosakonia pseudosacchari 728 5.56E-14 1445240.09 3.25 Klebsiella variicola
TABLE-US-00018 TABLE D FIG. 41 Greenhouse & Laboratory Data Heatmap N Activity Peak Produced/ Strain (mmol N/ Colonization acre Name Microbe hr) (CFU/g fw) season Taxonomic Designation CI006 3.88E-16 1.50E+07 0.24 Kosakonia sacchari 6-400 2.72E-13 1.79E+05 1.97 Kosakonia sacchari 6-397 1.14E-14 1.79E+05 0.08 Kosakonia sacchari CI137 3.24E-15 6.50E+06 0.85 Klebsiella variicola 137-1586 1.10E-13 1.82E+06 8.10 Klebsiella variicola 137-1382 4.81E-12 1.82E+06 354.60 Klebsiella variicola 1021 1.77E-14 2.69E+07 19.25 Kosakonia pseudosacchari 1021-1615 1.20E-13 2.69E+07 130.75 Kosakonia pseudosacchari 1021-1619 3.93E-14 2.69E+07 42.86 Kosakonia pseudosacchari 1021-1612 1.20E-13 2.69E+07 130.75 Kosakonia pseudosacchari 1021-1623 4.73E-17 2.69E+07 0.05 Kosakonia pseudosacchari 1293 5.44E-17 8.70E+08 1.92 Azospirillum lipoferum 1116 1.05E-14 1.37E+07 5.79 Enterobacter sp. 1113 8.05E-15 4.13E+07 13.45 Enterobacter sp. 910 1.19E-13 1.34E+06 6.46 Kluyvera intermedia 910-1246 2.16E-13 1.34E+06 11.69 Kluyvera intermedia 850 7.2301E-16 1.17E+06 0.03 Achromobacter spiritinus 852 5.96E-16 1.07E+06 0.03 Achromobacter marplatensis 853 6.42E-16 2.55E+06 0.07 Microbacterium murale
[0426] Conclusions:
[0427] The data in FIGS. 20, 40, 41, and Tables C and D, illustrates more than a dozen representative members of the described genus (i.e. microbes to the right of the line in the figures). Further, these numerous representative members come from a diverse array of taxonomic genera, which can be found in the above Tables C and D. Further still, the inventors have discovered numerous genetic attributes that depict a structure/function relationship that is found in many of the microbes. These genetic relationships can be found in the numerous tables of the disclosure setting forth the genetic modifications introduced by the inventors, which include introducing at least one genetic variation into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network.
[0428] Consequently, the newly discovered genus is supported by: (I) a robust dataset, (2) over a dozen representative members, (3) members from diverse taxonomic genera, and (4) classes of genetic modifications that define a structure/function relationship, in the underlying genetic architecture of the genus members.
Example 13: Methods and Assays for Detection of Non-Intergeneric Engineered Microbes
[0429] The present disclosure teaches primers, probes, and assays that are useful for detecting the microbes utilized in the various aforementioned Examples. The assays are able to detect the non-natural nucleotide "junction" sequences in the derived/mutant non-intergeneric microbes. These non-naturally occurring nucleotide junctions can be used as a type of diagnostic that is indicative of the presence of a particular genetic alteration in a microbe.
[0430] The present techniques are able to detect these non-naturally occurring nucleotide junctions via the utilization of specialized quantitative PCR methods, including uniquely designed primers and probes. The probes can bind to the non-naturally occurring nucleotide junction sequences. That is, sequence-specific DNA probes consisting of oligonucleotides that are labelled with a fluorescent reporter, which permits detection only after hybridization of the probe with its complementary sequence can be used. The quantitative methods can ensure that only the non-naturally occurring nucleotide junction will be amplified via the taught primers, and consequently can be detected via either a non-specific dye, or via the utilization of a specific hybridization probe. Another aspect of the method is to choose primers such that the primers flank either side of a junction sequence, such that if an amplification reaction occurs, then said junction sequence is present.
[0431] Consequently, genomic DNA can be extracted from samples and used to quantify the presence of microbes of the disclosure by using qPCR. The primers utilized in the qPCR reaction can be primers designed by Primer Blast (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) to amplify unique regions of the wild-type genome or unique regions of the engineered non-intergeneric mutant strains. The qPCR reaction can be carried out using the SYBR GreenER qPCR SuperMix Universal (Thermo Fisher P/N 11762100) kit, using only forward and reverse amplification primers; alternatively, the Kapa Probe Force kit (Kapa Biosystems P/N KK4301) can be used with amplification primers and a TaqMan probe containing a FAM dye label at the 5' end, an internal ZEN quencher, and a minor groove binder and fluorescent quencher at the 3' end (Integrated DNA Technologies).
[0432] Certain primer, probe, and non-native junction sequences--which can be used in the qPCR methods--are listed in the below Table E. Specifically, the non-native junction sequences can be found in SEQ ID NOs: 372-405.
TABLE-US-00019 TABLE E Microbial Detection up/down SEQ 100 bp SEQ 100 bp SEQ Junction SEQ F R base Junction stream ID upstream ID downstream ID ''/'' indicating Junction primer primer Probe CI Name junction NO of junction NO junction NO junction des. SEQ SEQ SEQ 1021 ds1131 up 304 TGGTGTCCGGGC 338 TTCTTGGTTCTCT 372 5'- disrupted N/A N/A N/A GAACGTCGCCAG GGAGCGCTTTAT TGGTGTCCGGGC nifL gene/ GTGGCACAAATT CGGCATCCTGAC GAACGTCGCCAG PinfC GTCAGAACTACG TGAAGAATTTGC GTGGCACAAATT ACACGACTAACC AGGCTTCTTCCCA GTCAGAACTACG GACCGCAGGAGT ACCTGGCTTGCA ACACGACTAACC GTGCGATGACCC CCCGTGCAGGTA GACCGCAGGAGT TGAATATGATGA GTTGTGATGAAC GTGCGATGACCC TGGA AT TGAATATGATGA TGGA/ TTCTTGGTTCTCT GGAGCGCTTTAT CGGCATCCTGAC TGAAGAATTTGC AGGCTTCTTCCCA ACCTGGCTTGCA CCCGTGCAGGTA GTTGTGATGAAC AT-3' 1021 ds1131 down 305 CGGAAAACGAGT 339 GCGATAGAACTC 373 5'- PinfC/ N/A N/A N/A TCAAACGGCGCG ACTTCACGCCCC CGGAAAACGAGT disrupted TCCCAATCGTATT GAAGGGGGAAGC TCAAACGGCGCG nifL gene AATGGCGAGATT TGCCTGACCCTAC TCCCAATCGTATT CGCGCCACGGAA GATTCCCGCTATT AATGGCGAGATT GTTCGCTTAACAG TCATTCACTGACC CGCGCCACGGAA GTCTGGAAGGCG GGAGGTTCAAAA GTTCGTTAACA AGCACCTTGGTA TGACCCAGCGAA GGTCTGGAAGGC TT C GAGCAGCTTGGT ATT/ GCGATAGAACTC ACTTCACGCCCC GAAGGGGGAAGC TGCCTGACCCTA CGATTCCCGCTAT TTCATTCACTGAC CGGAGGTTCAAA ATGACCCAGCGA AC-3' 1021 ds1133 N/A 306 CGCCAGAGAGTT 340 TCCCTGTGCGCCG 374 5'- 5' UTR N/A N/A N/A GAAATCGAACAT CGTCGCCGATGG CGCCAGAGAGTF and ATG/ TTCCGTAATACCG TGGCCAGCCAAC GAAATCGAACAT truncated CCATTACCCAGG TGGCGCGCTACC TTCCGTAATACC glnE gene AGCCGTTCTGGTT CGATCCTGCTCG GCCATTACCCAG GCACAGCGGAAA ATGAACTGCTCG GAGCCGTTCTGG ACGTTAACGAAA ACCCGAACACGC TTGCACAGCGGA GGATATTTCGCAT TCTATCAACCGA AAACGTTAACGA G CGG AAGGATATTTCG CATG/ TCCCTGTGCGCC GCGTCGCCGATG GTGGCCAGCCAA CTGGCGCGCTAC CCGATCCTGCTC GATGAACTGCTC GACCCGAACACG CTCTATCAACCG ACGG-3' 1021 ds1145 up 307 CGGGCGAACGTC 341 CGTTCTGTAATAA 375 5'- disrupted N/A N/A N/A GCCAGGTGGCAC TAACCGGACAAT CGGGCGAACGTC nifL gene/ AAATTGTCAGAA TCGGACTGATTA GCCAGGTGGCAC Prm1 CTACGACACGAC AAAAAGCGCCCT AAATTGTCAGAA TAACCGACCGCA CGCGGCGCTTTTT CTACGACACGAC GGAGTGTGCGAT TTATATTCTCGAC TAACCGACCGCA GACCCTGAATAT TCCATTTAAAATA GGAGTGTGCGAT GATGATGGATGC AAAAATCCAATC GACCCTGAATAT CAGC GATGATGGATGC CAGC/ CGTTCTGTAATA ATAACCGGACAA TFCGGACTGATT AAAAAAGCGCCC TCGCGGCGCTTTT TTTATATTCTCGA CTCCATTTAAAAT AAAAAATCCAAT C-3' 1021 ds1145 down 308 TCAACCTAAAAA 342 AACTCACTTCAC 376 5'- Prm1/ N/A N/A N/A AGTTTGTGTAATA GCCCCGAAGGGG TCAACCTAAAAA disrupted CTTGTAACGCTAC GAAGCTGCCTGA AGTTTGTGTAAT nifL gene ATGGAGATTAAC CCCTACGATTCCC ACTTGTAACGCT TCAATCTAGAGG GCTATTTCATTCA ACATGGAGATTA GTATTAATAATG CTGACCGGAGGT ACTCAATCTAGA AATCGTACTAAA TCAAAATGACCC GGGTATTAATAA CTGGTACTGGGC AGCGAACCGAGT TGAATCGTACTA GC CG AACTGGTACTGG GCGC/ AACTCACTTCAC GCCCCGAAGGGG GAAGCTGCCTGA CCCTACGATTCCC GCTATTTCATTCA CTGACCGGAGGT TCAAAATGACCC AGCGAACCGAGT CG-3' 1021 ds1148 up 309 CGGGCGAACGTC 343 CGCGTCAGGTTG 377 5'- disrupted N/A N/A N/A GCCAGGTGGCAC AACGTAAAAAAG CGGGCGAACGTC nifL gene/ AAATTGTCAGAA TCGGTCTGCGCA GCCAGGTGGCAC Prm7 CTACGACACGAC AAGCACGTCGTC AAATTGTCAGAA TAACCGACCGCA GTCCGCAGTTCTC CTACGACACGAC GGAGTGTGCGAT CAAACGTTAATT TAACCGACCGCA GACCCTGAATAT GGTTTCTGCTFCG GGAGTGTGCGAT GATGATGGATGC GCAGAACGATTG GACCCTGAATAT CAGC GC GATGATGGATGC CAGC/ CGCGTCAGGTTG AACGTAAAAAAG TCGGTCTGCGCA AAGCACGTCGTC GTCCGCAGTTCTC CAAACGTTAATT GGTTTCTGCTTCG GCAGAACGATTG GC-3' 1021 ds1148 down 310 AATTTTCTGCCCA 344 AACTCACTTCAC 378 5'- Prm4/ N/A N/A N/A AATGGCTGGGAT GCCCCGAAGGGG AATGGCTGCCCA disrupted TGTTCATTTTTTG GAAGCTGCCFGA AATGGCTGGGAT nifL gene TTTGCCTTACAAC CCCTACGATTCCC TGTTCATTTTTTG GAGAGTGACAGT GCTATTTCATTCA TTTGCCTTACAAC ACGCGCGGGTAG CTGACCGGAGGT GAGAGTGACAGT TTAACTCAACATC TCAAAATGACCC ACGCGCGGGTAG TGACCGGTCGAT AGCGAACCGAGT TTAACTCAACAT CG CTGACCGGTCGA T/ AACTCACTTCAC GCCCCGAAGGGG GAAGCTGCCTGA CCCTACGATTCCC GCTATTTCATTCA CTGACCGGAGGT TCAAAATGACCC AGCGAACCGAGT CG-3' CI006 ds126 N/A 311 GTAACCAATAAA 345 CCGATCCCCATC 379 5'- 5' UTR up N/A N/A N/A GGCCACCACGCC ACTGTGTGTCTTG GTAACCAATAAA to ATG- AGACCACACGAT TATTACAGTGCC GGCCACCACGCC 4 bp of AGTGATGGCAAC GCTTCGTCGGCTT AGACCACACGAT amtB gene/ ACTTTCCAGCTGC CGCCGGTACGAA AGTGATGGCAAC disrupted ACCAGCACCTGA TACGAATGACGC ACTTTCCAGCTGC amtB gene TGGCCCATGGTC GTTGCAGCTCAG ACCAGCACCTGA ACACCTTCAGCG CAACGAAAATTT TGGCCCATGGTC AAA TG ACACCTTCAGCG AAA/ CCGATCCCCATC ACTGTGTGTCTTG TATTACAGTGCC GCTTCGTCGGCTT CGCCGGTACGAA TACGAATGACGC CAACGAAAATTT TG-3' CI019 ds172 down 312 TGGTATTGTCAGT 346 CCGTCTCTGAAG 380 5'- Pnn1.2/ SEQ SEQ N/A CTGAATGAAGCT CTCTCGGTGAAC TGGTATTGTCAGT disrupted ID ID CTTGAAAAAGCT ATTGTTGCGAGG CTGAATGAAGCT nifL gene NO: NO: GAGGAAGCGGGC CAGGATGCGAGC CTTGAAAAAGCT 406 407 GTCGATTTAGTAG TGGTTGTGTMG GAGGAAGCGGGC CAAG TGCC AAATCAGTCCGA ACATTACCGATA GTCGATTTAGTA AAGT TCGC ATGCCGAGCCGC ATGTGCCGCGTG GAAATCAGTCCG TCGC AACA CAGTTTGTCGAAT AACGGGTGCGTT AATGCCGAGCCG CTCA ATGT C ATG CCAGTTTGTCGA CAGG TCAC ATC/ CCGTCTCTGAAG CTCTCGGTGAAC ATTGTTGCGAGG CAGGATGCGAGC TGGTTGTGTTTTG ACATTACCGATA ATGTGCCGCGTG AACGGGTGCGTT ATG-3' CI019 ds172 up 313 ACCGATCCGCAG 347 TGAACATCACTG 381 5'- disrupted N/A N/A N/A GCGCGCATTTGTT ATGCACAAGCTA ACCGATCCGCAG nifL gene/ ATGCCAATCCGG CCTATGTCGAAG GCGCGCATTTGTT Prm1.2 CATTCTGCCGCCA AATTAACTAAAA ATGCCAATCCGG GACGGGTTTTGC AACTGCAAGATG CATTCTGCCGCC ACTTGAGACACTT CAGGCATTCGCG AGACGGGTTTTG TTGGGCGAGAAC TTAAAGCCGACT CACTTGAGACAC CACCGTCTGCTGG TGAGAAATGAGA TTTTGGGCGAGA AGAT ACCACCGTCTGC TGG/ TGAACATCACTG ATGCACAAGCTA CCTATGTCGAAG AATTAACTAAAA AACTGCAAGATG CAGGCATTCGCG TTAAAGCCGACT TGAGAAATGAGA AGAT-3' CI019 ds175 down 314 CGGGAACCGGTG 348 CCGTCTCTGAAG 382 5'- Prm3.1/ SEQ SEQ SEQ TTATAATGCCGCG CTCTCGGTGAAC CGGGAACCGGTG disrupted ID ID ID CCCTCATATTGTG ATTGTTGCGAGG TTATAATGCCGC nifL gene NO: NO: NO: GGGATTTCTTAAT CAGGATGCGAGC GCCCTCATATTGT 408 409 410 GACCTATCCTGG TGGTTGTGTTTTG GGGGATTTCTTA CGCC GGCA /56- GTCCTAAAGTTGT ACATTACCGATA ATGACCTATCCT CTCA TAAC FAM/ AGTTGACATTAG ATGTGCCGCGTG GGGTCCTAAAGT TATT GCAC TA CGGAGCACTAAC AACGGGTGCGTT TGTAGTTGACATT GTGG CCGT ACC ATG AGCGGAGCACTA GGAT TCA CGT AC/ C/ CCGTCTCTGAAG ZEN/T CTCTCGGTGAAC CTG ATTGTTGCGAGG AAG CAGGATGCGAGC CTC TGGTTGTGTTTTG TCG ACATTACCGATA GT/ ATGTGCCGCGTG 3IABkFQ/ AACGGGTGCGTT ATG-3' CI019 ds175 up 315 ACCGATCCGCAG 349 TACAGTAGCGCC 383 5c disrupted N/A N/A N/A GCGCGCATTTGTT TCTCAAAAATAG ACCGATCCGCAG nifL gene/ ATGCCAATCCGG ATAAACGGCTCA GCGCGCATTTGTT Prm3.1 CATTCTGCCGCCA TGTACGTGGGCC ATGCCAATCCGG GACGGGTTTTGC GTTTATTTTTTT CATTCTGCCGCC ACTTGAGACACTT ACCCATAATCGG AGACGGGTTTTG TTGGGCGAGAAC GAACCGGTGTTA CACTTGAGACAC CACCGTCTGCTGG TAATGCCGCGCC TTTTGGGCGAGA CTC ACCACCGTCTGC TGG/ TACAGTAGCGCC
TCTCAAAAATAG ATAAACGGCTCA TGTACGTGGGCC GTTTATTTTTTCT ACCCATAATCGG GAACCGGTGTTA TAATGCCGCGCC CTC-3' CI006 ds20 down 316 TCAACCTAAAAA 350 AACTCACTTCAC 384 5'- Prm1/ SEQ SEQ SEQ AGTTTGTGTAATA ACCCCGAAGGGG TCAACCTAAAAA disrupted ID ID ID CTTGTAACGCTAC GAAGTTGCCTGA AGTTTGTGTAAT nifL gene NO: NO: NO: ATGGAGATTAAC CCCTACGATTCCC ACTTGTAACGCT 411 412 413 TCAATCTAGAGG GCTATTTCATTCA ACATGGAGATTA TAAA CAAA 56- GTATTAATAATG CTGACCGGAGGT ACTCAATCTAGA CTGG TCGA FAM AATCGTACTAAA TCAAAATGACCC GGGTATTAAXAA TACT AGCG AAG CTGGTACTGGGC AGCGAACCGAGT TGAATCGTACTA GGGC CCAG TTGC GC CG AACTGGTACTGG GCAA ACGG CT/ GCGC/ CT TAT ZEN/G AACTCACTTCAC ACC ACCCCGAAGGGG CTAC GAAGTTGCCTGA GATT CCCTACGATTCCC CCC/ GCTATTTCATTCA 3IABkFQ/ CTGACCGGAGGT TCAAAATGACCC AGCGAACCGAGT CG-3' CI006 ds20 up 317 GGGCGACAAACG 351 CGTCCTGTAATA 385 5'- disrupted N/A N/A N/A GCCTGGTGGCAC ATAACCGGACAA GGGCGACAAACG nifL gene/ AAATTGTCAGAA TTCGGACTGATTA GCCTGGTGGCAC Prm1 CTACGACACGAC AAAAAGCGCCCT AAATTGTCAGAA TAACTGACCGCA TGTGGCGCTTTTT CTACGACACGAC GGAGTGTGCGAT TTATATTCCCGCC TAACTGACCGCA GACCCTGAATAT TCCATTTAAAATA GGAGTGTGCGAT GATGATGGATGC AAAAATCCAATC GACCCTGAATAT CGGC GATGATGGATGC CGGC/ CGTCCTGTAATA ATAACCGGACAA TTCGGACTGATT AAAAAAGCGCCC TTGTGGCGCTTTT TTTATATTCCCGC CTCCATTTAAAAT AAAAAATCCAAT C-3' CI006 ds24 up 318 GGGCGACAAACG 352 GGACATCATCGC 386 5'- disrupted SEQ SEQ SEQ GCCTGGTGGCAC GACAAACAATAT GGGCGACAAACG nifL gene/ ID ID ID AAATTGTCAGAA TAATACCGGCAA GCCTGGTGGCAC Prm5 NO: NO: NO: CTACGACACGAC CCACACCGGCAA AAATTGTCAGAA 414 415 416 TAACTGACCGCA TTTACGAGACTG CTACGACACGAC GGTG GCGC /56- GGAGTGTGCGAT CGCAGGCATCCT TAACTGACCGCA CACT AGTC FAM/ GACCCTGAATAT TTCTCCCGTCAAT GGAGTGTGCGAT CTTT TCGT CA GATGATGGATGC TTCTGTCAAATAA GACCCTGAATAT GCAT AAAT GGA CGGC AG GATGATGGATGC GGTT TGCC GTG CGGC/ T/ GGACATCATCGC ZEN/G GACAAACAATAT CGA TAATACCGGCAA TGA CCACACCGGCAA CCC TTTACGAGACTG TGA CGCAGGCATCCT AT/ TTCTCCCGTCAAT 3IABkFQ TTCTGTCAAATA AAG-3' CI006 ds24 down 319 TAAGAATTATCTG 353 AACTCACTTCAC 387 5'- Prm5/ N/A N/A N/A GATGAATGTGCC ACCCCGAAGGGG TAAGAATTATCT disrupted ATTAAATGCGCA GAAGTTGCCTGA GGATGAATGTGC nifL gene GCATAATGGTGC CCCTACGATTCCC CATTAAATGCGC GTTGTGCGGGAA GCTATTTCATTCA AGCATAATGGTG AACTGCTTTTTTT CTGACCGGAGGT CGTTGTGCGGGA TGAAAGGGTTGG TCAAAATGACCC AAACTGCTTTTTT TCAGTAGCGGAA AGCGAACCGAGT TTGAAAGGGTTG AC CG GTCAGTAGCGGA AAC/ AACTCACTTCAC ACCCCGAAGGGG GAAGTTGCCTGA CCCTACGATTCCC GCTATTTCATTCA CTGACCGGAGGT TCAAAATGACCC AGCGAACCGAGT CG-3' CI006 ds30 N/A 320 CGCCAGAGAGTC 354 TTTAACGATCTGA 388 5'- 5' UTR N/A N/A N/A GAAATCGAACAT TTGGCGATGATG CGCCAGAGAGTC and ATG/ TTCCGTAATACCG AAACGGATTCGC GAAATCGAACAT truncated CGATTACCCAGG CGGAAGATGCGC TTCCGTAATACC glnE gene AGCCGTTCTGGTT TTTCTGAGAGCTG GCGATTACCCAG GCACAGCGGAAA GCGCGAATTGTG GAGCCGTTCTGG ACGTTAACGAAA GCAGGATGCGTT TTGCACAGCGGA GGATATTTCGCAT GCAGGAGGAGGA AAACGTTAACGA G TT AAGGATATTTCG CATG/ TTTAACGATCTG ATTGGCGATGAT GAAACGGATTCG CCGGAAGATGCG CTTTCTGAGAGCT GGCGCGAATTGT GGCAGGATGCGT TGCAGGAGGAGG ATT-3' CI006 ds31 N/A 321 CGCCAGAGAGTC 355 GCACTGAAACAC 389 5'- 5' UTR N/A N/A N/A GAAATCGAACAT CTCATTTCCCTGT CGCCAGAGAGTC and ATG/ TTCCGTAATACCG GTGCCGCGTCGC GAAATCGAACAT truncated CGATTACCCAGG CGATGGTTGCCA TTCCGTAATACC glnE gene AGCCGTTCTGGTT GTCAGCTGGCGC CFCGATTACCCAG GCACAGCGGAAA GCTACCCGATCCT GAGCCGTTCTGG ACGTTAACGAAA GCTTGATGAATT TTGCACAGCGGA GGATATTTCGCAT GCTCGACCCGAA AAACGTTAACGA G TA AAGGATATTTCG CATG/ GCACTGAAACAC CTCATTTCCCTGT GTGCCGCGTCGC CGATGGTTGCCA GTCAGCTGGCGC GCTACCCGATCC TGCTTGATGAATT GCTCGACCCGAA TA-3' CI019 ds34 N/A 322 GATGATGGATGC 356 GCGCTCAAACAG 390 5'- 5' UTR N/A N/A N/A TTTCTGGTTAAAC TTAATCCGTCTGT GATGATGGATGC and ATG/ GGGCAACCTCGT GTGCCGCCTCGC TTTCTGGTTAAAC truncated TAACTGACTGACT CGATGGTCGCGA GGGCAACCTCGT glnE gene AGCCTGGGCAAA CACAACTTGCAC TAACTGACTGAC CTGCCCGGGCTTT GTCATCCTTTATT TAGCCTGGGCAA TTTTTGCAAGGAA GCTCGATGAACT ACTGCCCGGGCT TCTGATTTCATG GCTCGACCCGCG TTTTTTTGCAAGG CA AATCTGATTTCAT G/ GCGCTCAAACAG TTAATCCGTCTGT GTGCCGCCTCGC CGATGGTCGCGA CACAACTTGCAC GTCATCCTTTATT GCTCGATGAACT GCTCGACCCGCG CA-3' CI019 ds70 up 323 ACCGATCCGCAG 357 AGTCTGAACTCA 391 5'- disrupted N/A N/A N/A GCGCGCATTTGTT TCCTGCGGCAGT ACCGATCCGCAG nifL gene/ ATGCCAATCCGG CGGTGAGACGTA GCGCGCATTTGTT Prm4 CATTCTGCCGCCA TTTTTGACCAAAG ATGCCAATCCGG GACGGGTTTTGC AGTGATCTACAT CATTCTGCCGCC ACTTGAGACACTT CACGGAATTTTGT AGACGGGTTTTG TTGGGCGAGAAC GGTTGTTGCTGCT CACTTGAGACAC CACCGTCTGCTGG TAAAAGGGCAAA TTTTGGGCGAGA T ACCACCGTCTGC TGG/ AGTCTGAACTCA TCCTGCGGCAGT CGGTGAGACGTA TTTTTGACCAAA GAGTGATCTACA TCACGGAATTTT GTGGTTGTTGCTG CTTAAAAGGGCA AAT-3' CI019 ds70 down 324 CATCGGACACCA 358 CCGTCTCTGAAG 392 5'- Prm4/ N/A N/A N/A CCAGCTTACAAA CTCTCGGTGAAC CATCGGACACCA disrupted TTGCCTGATTGCG ATTGTTGCGAGG CCAGCTTACAAA nifL gene GCCCCGATGGCC CAGGATGCGAGC TTGCCTGATTGCG GGTATCACTGAC TGGTTGTGTTTTG GCCCCGATGGCC CGACCATTTCGTG ACATTACCGATA GGTATCACTGAC CCTTATGTCATGC ATGTGCCGCGTG CGACCATTTCGT GATGGGGGCTGG AACGGGTGCGTT GCCTTATGTCATG G ATG CGATGGGGGCTG GG/ CCGTCTCTGAAG CTCTCGGTGAAC ATTGTTGCGAGG CAGGATGCGAGC TGGTTGTGTTTTG ACATTACCGATA ATGTGCCGCGTG AACGGGTGCGTT ATG-3' 137 ds799 down 325 TCTTCAACAACTG 359 GCCATTGAGCTG 393 5'- PinfC/ SEQ SEQ SEQ GAGGAATAAGGT GCTTCCCGACCG TCTTCAACAACT disrupted ID ID ID ATTAAAGGCGGA CAGGGCGGCACC GGAGGAATAAGG nifL gene NO: NO: NO: AAACGAGTTCAA TGCCTGACCCTGC TATTAAAGGCGG 417 418 419 ACGGCACGTCCG GTTTCCCGCTGTT AAAACGAGTTCA CTCG AGGG /56- AATCGTATCAAT TAACACCCTGAC AACGGCACGTCC GCAG TGTT FAM/ GGCGAGATTCGC CGGAGGTGAAGC GAATCGTATCAA CATG AAAC AA GCCCTGGAAGTT ATGATCCCTGAA TGGCGAGATTTCG GACG AGCG CGG CGC TC CGCCCTGGAAGT TAA GGAA CAC TCGC/ A G/ GCCATTGAGCTG ZEN/T GCTTCCCGACCG CCG CAGGGCGGCACC AAT TGCCTGACCCTG CGT CGTTTCCCGCTGT ATC TTAACACCCTGA AA/ CCGGAGGTGAAG 3IABkFQ/ CATGATCCCTGA ATC-3' 137 ds799 up 326 TCCGGGTTCGGCT 360 AGCGTCAGGTAC 394 5'- disrupted N/A N/A N/A TACCCCGCCGCGT CGGTCATGATTC TCCGGGTTCGGC nifL gene/ TTTGCGCACGGTG ACCGTGCGATTCT TTACCCCGCCGC PinfC TCGGACAATTTGT CGGTTCCCTGGA GTTTTGCGCACG CATAACTGCGAC GCGCTTCATTGGC GTGTCGGACAAT ACAGGAGTTTGC ATCCTGACCGAA TTGTCATAACTGC GATGACCCTGAA GAGTTCGCTGGC GACACAGGAGTT TATGATGCTCGA TTCTTCCCAACCT TGCGATGACCCT G GAATATGATGCT CGA/ AGCGTCAGGTAC CGGTCATGATTC ACCGTGCGATTC TCGGTTCCCTGG AGCGCTTCATTG GCATCCTGACCG AAGAGTTCGCTG GCTTCTTCCCAAC CTG-3' 137 ds809 N/A 327 ATCGCAGCGTCTT 361 GCGCTGAAGCAC 395 5'- 5' UTR SEQ SEQ SEQ TGAATATTTCCGT CTGATCACGCTCT ATCGCAGCGTCT and ATG/ ID ID ID CGCCAGGCGCTG GCGCGGCGTCGC TTGAATATTTCCG truncated NO: NO: NO: GCTGCCGAGCCG CGATGGTCGCCA TCGCCAGGCGCT glnE gene 420 421 422 TTCTGGCTGCATA GCCAGCTGGCGC GGCTGCCGAGCC GAGC GCCG /56- GTGGAAAACGAT GCCACCCGCTGC GTTCTGGCTGCAT CGTT TCGG FAM AATTTCAGGCCA TGCTGGATGAGC AGTGGAAAACGA CTGG CTGA TTAT GGGAGCCCTTAT TGCTGGATCCCA TAATTTCAGGCC CTGC TAGA GGC G ACA AGGGAGCCCTTA ATAG GG GC/ TG/ ZEN/T
GCGCTGAAGCAC GAA CTGATCACGCTCT GCA GCGCGGCGTCGC CCTG CGATGGTCGCCA ATC GCCAGCTGGCGC A/ GCCACCCGCTGC 3IABkFQ/ TGCTGGATGAGC TGCTGGATCCCA ACA-3' 137 ds843 up 328 TCCGGGTTCGGCT 362 GCCCGCTGACCG 396 5'- disrupted N/A N/A N/A TACCCCGCCGCGT ACCAGAACTTCC TCCGGGTTCGGC nifL gene/ TTTGCGCACGGTG ACCTTGGACTCG TTACCCCGCCGC Prm1.2 TCGGACAATTTGT GCTATACCCTTGG GTTTTGCGCACG CATAACTGCGAC CGTGACGGCGCG GTGTCGGACAAT ACAGGAGTTTGC CGATAACTGGGA TTGTCATAACTGC GATGACCCTGAA CTACATCCCCATT GACACAGGAGTT TATGATGCTCGA CCGGTGATCTTAC TGCGATGACCCT C GAATATGATGCT CGA/ GCCCGCTGACCG ACCAGAACTTCC ACCTTGGACTCG GCTATACCCTTG GCGTGACGGCGC GCGATAACTGGG ACTACATCCCCA TTCCGGTGATCTT ACC-3' 137 ds843 down 329 TCACTTTTTAGCA 363 GCCATTGAGCTG 397 5'- Prm1.2/ N/A N/A N/A AAGTTGCACTGG GCTTCCCGACCG TCACTTTTCAGCA disrupted ACAAAAGGTACC CAGGGCGGCACC AAGTTGCACTGG nifL gene ACAATTGGTGTA TGCCTGACCCTGC ACAAAAGGTACC CTGATACTCGAC GTTTCCCGCTGTT ACAATTGGTGTA ACAGCATTAGTG TAACACCCTGAC CTGATACTCGAC TCGATTTTTCATA CGGAGGTGAAGC ACAGCATTAGTG TAAAGGTAATTTT ATGATCCCTGAA TCGATTTTTCATA G TC TAAAGGTAATTT TG/ GCCATTGAGCTG GCTTCCCGACCG CAGGGCGGCACC TGCCTGACCCTG CGTTTCCCGCTGT TTAACACCCTGA CCGGAGGTGAAG CATGATCCCTGA ATC-3' 137 ds853 up 330 TCCGGGTTCGGCT 364 GCTAAAGTTCTC 398 5'- disrupted N/A N/A N/A TACCCCGCCGCGT GGCTAATCGCTG TCCGGGTTCGGC nifL gene/ TTTGCGCACGGTG ATAACATTTGAC TTACCCCGCCGC Prm6.2 TCGGACAATTTGT GCAATGCGCAAT GTTTTGCGCACG CATAACTGCGAC AAAAGGGCATCA GTGTCGGACAAT ACAGGAGTTTGC TTTGATGCCCTTT TTGTCATAACTGC GATGACCCTGAA TTGCACGCTTTCA GACACAGGAGTT TATGATGCTCGA TACCAGAACCTG TGCGATGACCCT GC GAATATGATGCT CGA/ GCTAAAGTTCTC GGCTAATCGCTG ATAACATTDGAC GCAATGCGCAAT AAAAGGGCATCA TTTGATGCCCTTT TTGCACGCTTTCA TACCAGAACCTG GC-3' 137 ds853 down 331 GTTCTCCTTTGCA 365 GCCATTGAGCTG 399 5'- Prm6.2/ N/A N/A N/A ATAGCAGGGAAG GCTTCCCGACCG GTTCTCCTTTGCA disrupted AGGCGCCAGAAC CAGGGCGGCACC ATAGCAGGGAAG nifL gme CGCCAGCGTTGA TGCCTGACCCTGC AGGCGCCAGAAC AGCAGTTTGAAC GTTTCCCGCTGTT CGCCAGCGTTGA GCGTTCAGTGTAT TAACACCCTGAC AGCAGTTTGAAC AATCCGAAACTT CGGAGGTGAAGC GCGTTCAGTGTA AATTTCGGTTTGG ATGATCCCTGAA TAATCCGAAACT A TC TAATTTCGGTTTG GA/ GCCATTGAGCTG GCTTCCCGACCG CAGGGCGGCACC TGCCTGACCCTG CGTTTCCCGCTGT TTAACACCCTGA CCGGAGGTGAAG CATGATCCCTGA ATC-3' 137 ds857 up 332 TCCGGGTTCGGCT 366 CGCCGTCCTCGC 400 5'- disrupted N/A N/A N/A TACCCCGCCGCGT AGTACCATTGCA TCCGGGTTCGGC nifL gene/ TTTGCGCACGGTG ACCGACTTTACA TTACCCCGCCGC Prm8.2 TCGGACAATTTGT GCAAGAAGTGAT GTTTTGCGCACG CATAACTGCGAC TCTGGCACGCAT GTGTCGGACAAT ACAGGAGTTTGC GGAACAAATTCT TTGTCATAACTGC GATGACCCTGAA TGCCAGTCGGGC GACACAGGAGTT TATGATGCTCGA TTTATCCGATGAC TGCGATGACCCT GAA GAATATGATGCT CGA/ CGCCGTCCTCGC ACTACCATTGCA ACCGACTTTACA GCAAGAAGTGAT TCTGGCACGCAT GGAACAAATTCT TGCCAGTCGGGC TTTATCCGATGAC GAA-3' 137 ds857 down 333 GATATGCCTGAA 367 GCCATTGAGCTG 401 5'- Prm8.2/ N/A N/A N/A GTATTCAATTACT GCTTCCCGACCG GATATGCCTGAA disrupted TAGGCATTTACTT CAGGGCGGCACC GTATTCAATTACT nifL gene AACGCAGGCAGG TGCCTGACCCTGC TAGGCATTTACTT CAATTTTGATGCT GTTTCCCGCTGTT AACGCAGGCAGG GCCTATGAAGCG TAACACCCTGAC CAATTTTGATGCT TTTGATTCTGTAC CGGAGGTGAAGC GCCTATGAAGCG TTGAGCTTGATC ATGATCCCTGAA TTTGATTCTGTAC TC TTGAGCTTGATC/ GCCATTGAGCTG GCTTCCCGACCG CAGGGCGGCACC TGCCTGACCCTG CGTTTCCCGCTGT TTAACACCCTGA CCGGAGGTGAAG CATGATCCCTGA ATC-3' 63 ds908 down 334 TGGTATTGTCAGT 368 TCTTTAGATCTCT 402 5'- PinfC/ SEQ SEQ N/A CTGAATGAAGCT CGGTCCGCCCTG TGGTATTGTCAGT disrupted ID ID CTTGAAAAAGCT ATGGCGGCACCT CTGAATGAAGCT nifL gene NO: NO: GAGGAAGCGGGC TGCTGACGTTAC CTTGAAAAAGCT 423 424 GTCGATTTAGTAG GCCTGCCGGTAC GAGGAAGCGGGC GGAA GGGC AAATCAGTCCGA AGCAGGTTATCA GTCGATTTAGTA AACG GGAC ATGCCGAGCCGC CCGGAGGCTTAA GAAATCAGTCCG AGTT CGAG CAGTTTGTCGAAT AATGACCCAGTT AATGCCGAGCCG CAAC AGAT C ACC CCAGTTTGTCGA CGGC CTAA ATC/ TCTTTAGATCTCT CGGTCCGCCCTG ATGGCGGCACCT TGCTGACGTTAC GCCTGCCGGTAC AGCAGGTTATCA CCGGAGGCTTAA AATGACCCAGTT ACC-3' 63 ds908 up 335 TGCAAATTGCAC 369 TGAATATCACTG 403 5'- disrupted N/A N/A N/A GGTTATTCCGGGT ACTCACAAGCTA TGCAAATTGCAC nifL gene/ GAGTATATGTGT CCTATGTCGAAG GGTFATTCCGGG PinfC GATTTGGGTTCCG AATTAACTAAAA TGAGTATATGTG GCATTGCGCAAT AACTGCAAGATG TGATTTGGGTTCC AAAGGGGAGAAA CAGGCATTCGCG GGCATTGCGCAA GACATGAGCATC TTAAAGCCGACT TAAAGGGGAGAA ACGGCGTTATCA TGAGAAATGAGA AGACATGAGCAT GC AGAT CACGGCGTTATC AGC/ TGAATATCACTG ACTCACAAGCTA CCTATGTCGAAG AATTAACTAAAA AACTGCAAGATG CAGGCATTCGCG TTAAAGCCGACT TGAGAAATGAGA AGAT-3' 910 ds960 up 336 TCAGGGCTGCGG 370 CTGGGGTCACTG 404 5'- disrupted N/A N/A N/A ATGTCGGGCGTTT GAGCGCTTTATC TCAGGGCTGCGG nifL gene/ CACAACACAAAA GGCATCCTGACC ATGTCGGGCGTT PinfC TGTTGTAAATGCG GAAGAATTTGCC TCACAACACAAA ACACAGCCGGGC GGTTTCTTCCCGA ATGTTGTAAATG CTGAAACCAGGA CCTGGCTGGCCC CGACACAGCCGG GCGTGTGATGAC CTGTTCAGGTTGT GCCTGAAACCAG CTTTAATATGATG GGTGATGAATAT GAGCGTGTGATG C CA ACCTTTAATATG ATGC/ CTGGGGTCACTG GAGCGCTTTATC GGCATCCTGACC GAAGAATTTGCC GGTTTCTTCCCGA CCTGGCTGGCCC CTGTTCAGGTTGT GGTGATGAATAT CA-3' 910 ds960 down 337 CGGAAAACGAGT 371 GCAATAGAACTA 405 5'- PinfC/ N/A N/A N/A TCAAACGGCACG ACTACCCGCCCT CGGAAAACGAGT disrupted TCCGAATCGTATC GAAGGCGGTACC TCAAACGGCACG nifL gene AATGGCGAGATT TGCCTGACCCTGC TCCGAATCGTAT CGCGCCCAGGAA GATTCCCGTTATT CAATGGCGAGAT GTTCGCTTAACTG TCATTCACTGACC TCGCGCCCAGGA GTCTGGAAGGTG GGAGGCCCACGA AGTTCGCTTAACT AGCAGCTGGGTA TGACCCAGCGAC GGTCTGGAAGGT TT C GAGCAGCTGGGT ATT/ GCAATAGAACTA ACTACCCGCCCT GAAGGCGGTACC TGCCTGACCCTG CGATTCCCGTTAT TTCATTCACTGAC CGGAGGCCCACG CC-3'
TABLE-US-00020 TABLE F Engineered Non-intergeneric Microbes Strain Name Genotype SEQ ID NO CI006 16S rDNA-contig 5 62 CI006 16S rDNA-contig 8 63 CI019 16S rDNA 64 CI006 nifH 65 CI006 nifD 66 CI006 nifK 67 CI006 nifL 63 CI006 nifA 69 CI019 nifH 70 CI019 nifD 71 CI019 nifK 72 CI019 nifL 73 CI019 nifA 74 CI006 Prm5 with 500 bp 75 flanking regions CI006 nifLA operon-upstream 76 intergenic region plus nifL and nifA CDSs CI006 nifL (Amino Acid) 77 CI006 nifA (Amino Acid) 78 CI006 glnE 79 CI006 glnE_KO1 80 CI006 glnE (Amino Acid) 81 CI006 glnE_KO1 (Amino Acid) 82 CI006 GME ATase domain 83 (Amino Acid) CM029 Prm5 inserted into nifL 84 region
TABLE-US-00021 TABLE G Engineered Non-intergeneric Microbes Associated Novel Strain SEQ ID Junction If Strain ID NO Genotype Description Applicable CI63; 63 SEQ ID 16S N/A N/A CI063 NO 85 CI63; 63 SEQ ID nifH N/A N/A CI063 NO 86 CI63; 63 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A CI063 NO 87 in 63 genome CI63; 63 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A CI063 NO 88 in 63 genome CI63; 63 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A CI063 NO 89 in 63 genome CI63; 63 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A CI063 NO 90 in 63 genome CI63; 63 SEQ ID nifL N/A N/A CI063 NO 91 CI63; 63 SEQ ID nifA N/A N/A CI063 NO 92 CI63; 63 SEQ ID glnE N/A N/A CI063 NO 93 CI63; 63 SEQ ID amtB N/A N/A CI063 NO 94 CI63; 63 SEQ ID PinfC 500 bp immediately upstrea of the ATG N/A CI063 NO 95 start codon of the infC gene CI137 137 SEQ ID 16S N/A N/A NO 96 CI137 137 SEQ ID nifH1 1 of 2 unique genes annotated as nifH N/A NO 97 in 137 genome CI137 137 SEQ ID nifH2 2 of 2 unique genes annotated as nifH N/A NO 98 in 137 genome CI137 137 SEQ ID nifDl 1 of 2 unique genes annotated as nifD N/A NO 99 in 137 genome CI137 137 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 100 in 137 genorne CI137 137 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A NO 101 in 137 genome CI137 137 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A NO 102 in 137 genome CI137 137 SEQ ID nifL N/A N/A NO 103 CI137 137 SEQ ID nifA N/A N/A NO 104 CI137 137 SEQ ID glnE N/A N/A NO 105 CI137 137 SEQ ID PinfC 500 bp immediately upstream of the N/A NO 106 TTG start codon of infC CI137 137 SEQ ID amtB N/A N/A NO 107 CI137 137 SEQ ID Prm8.2 internal promoter located in nfpI gene; N/A NO 108 299 bp starting at 81 bp after the A of the ATG of the nlpI gene CI137 137 SEQ ID Prm6.2 300 bp upstream of the secE gene N/A NO 109 starting at 57 bp upstream of the A of the ATG of secE CI137 137 SEQ ID Prm1.2 400 bp immediately upstream of the N/A No 110 ATG of cspE gene none 728 SEQ ID 16S N/A N/A No 111 none 728 SEQ ID nifH N/A N/A NO 112 none 728 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A NO 113 in 728 genome none 728 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 114 in 728 genome none 728 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A NO 115 in 728 genome none 728 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A NO 116 in 728 genome none 728 SEQ ID nifL N/A N/A NO 117 none 728 SEQ ID nifA N/A N/A NO 118 none 728 SEQ ID glnE N/A N/A NO 119 none 728 SEQ ID amtB N/A N/A NO 120 none 850 SEQ ID 16S N/A N/A NO 121 none 852 SEQ ID 16S N/A N/A NO 122 none 853 SEQ ID 16S N/A N/A NO 123 none 910 SEQ ID 16S N/A N/A NO 124 none 910 SEQ ID nifH N/A N/A NO 125 none 910 SEQ ID Dinitrogenase iron- N/A N/A NO 126 molybdenum cofactor CDS none 910 SEQ ID nifD1 N/A N/A NO 127 none 910 SEQ ID nifD2 N/A N/A NO 128 none 910 SEQ ID nifK1 N/A N/A NO 129 none 910 SEQ ID nifK2 N/A N/A NO 130 none 910 SEQ ID nifL N/A N/A NO 131 none 910 SEQ ID nifA N/A N/A NO 132 none 910 SEQ ID glnE N/A N/A NO 133 MC 910 SEQ ID amtB N/A N/A NO 134 none 910 SEQ ID PinfC 498 bp immediately upstream of the N/A NO 135 ATG of the infC gene none 1021 SEQ ID 16S N/A N/A NO 136 none 1021 SEQ ID nifH N/A N/A NO 137 none 1021 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A NO 138 in 910 genome none 1021 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 139 in 910 genome none 1021 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A NO 140 in 910 genome none 1021 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A NO 141 in 910 genome none 1021 SEQ ID nifL N/A N/A NO 142 none 1021 SEQ ID nifA N/A N/A NO 143 none 1021 SEQ ID glnE N/A N/A NO 144 none 1021 SEQ ID amtB N/A N/A NO 145 none 1021 SEQ ID PinfC 500 bp immediately upstream of the N/A NO 146 ATG start codon of the infC gene none 1021 SEQ ID Prm1 348 bp includes the 319 bp immediately N/A NO 147 upstream of the ATG start codon of the lpp gene and the first 29 bp of the lpp gene none 1021 SEQ ID Prm7 339 bp upstream of the sspA gene, N/A NO 148 ending at 46 bp upstream of the ATG of the sspA gene none 1113 SEQ ID 16S N/A N/A NO 149 none 1113 SEQ ID nifH N/A N/A NO 150 none 1113 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A NO 151 in 1113 genome none 1113 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 152 in 1113 genome none 1113 SEQ ID nifK N/A N/A NO 153 none 1113 SEQ ID nifL N/A N/A NO 154 none 1113 SEQ ID nifA partial gene due to a gap in the sequence assembly, N/A NO 155 we can only identify a partial gene from the 1113 genome none 1113 SEQ ID glnE N/A N/A NO 156 none 1116 SEQ ID 16S N/A NO 157 none 1116 SEQ ID nifH N/A NO 158 none 1116 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A NO 159 in 1116 genome none 1116 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 160 in 1116 genome none 1116 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A NO 161 in 1116 genome none 1116 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A NO 162 in 1116 genome none 1116 SEQ ID nifL N/A N/A NO 163 none 1116 SEQ ID nifA N/A N/A NO 164 none 1116 SEQ ID glnE N/A N/A NO 165 none 1116 SEQ ID amtB N/A N/A NO 166 none 1293 SEQ ID 16S N/A N/A NO 167 none 1293 SEQ ID nifH N/A N/A NO 168 none 1293 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A NO 169 in 1293 genome none 1293 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 170 in 1293 genome none 1293 SEQ ID nifK 1 of 2 unique genes annotated as nifK N/A NO 171 in 1293 genome none 1293 SEQ ID nifK1 2 of 2 unique genes annotated as nifK N/A NO 172 in 1293 genome none 1293 SEQ ID nifA N/A N/A NO 173 none 1293 SEQ ID glnE N/A N/A NO 174 none 1293 SEQ ID amtB1 1 of 2 unique genes annotated as amtB N/A NO 175 in 1293 genome none 1293 SEQ ID amtB2 2 of 2 unique genes annotated as amtB N/A NO 176 in 1293 genome none 1021-1612 SEQ ID .DELTA.nifL::PinfC starting at 24 bp after the A of the ATG ds1131 NO 177 start codon, 1375 bp of nifL, have been deleted and replaced with the 1021 PinfC promoter sequence none 1021-1612 SEQ ID .DELTA.nifL::PinfC with starting at 24 bp after the A of the ATG ds1131 NO 178 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the 1021 PinfC promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 1021-1612 SEQ ID glnE.DELTA.AR-2 glnE gene with 1673 bp immediately ds1133 NO 179 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain none 1021-1612 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1673 bp immediately ds1133 NO 180 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included none 1021-1615 SEQ ID .DELTA.nifL::Prm1 starting at 24 bp after the A of the ATG ds1145 NO 181 start codon, 1375 bp of nifL have been deleted and replaced with the 1021 Prm1 promoter sequence none 1021-1615 SEQ ID .DELTA.nifL:Prm1 with starting at 24 bp after the A of the ATG ds1145 NO 182 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the 1021 rm1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 1021-1615 SEQ ID glnE.DELTA.AR-2 glnE gene with 1673 bp immediately ds1133 NO 183 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain none 1021-1615 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1673 bp immediately ds1133 NO 184 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE
gene upstream and downstream are included none 1021-1619 SEQ ID .DELTA.nifL::Prm1 starting at 24 bp after the A of the ATG ds1145 NO 185 start codon, 1375 bp of nifL have been deleted and replaced with the 1021 Prm1 promoter sequence none 1021-1619 SEQ ID .DELTA.nifL:Prm1 with starting at 24 bp after the A of the ATG ds1145 NO 186 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the 1021 rm1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 1021-1623 SEQ ID glnE.DELTA.AR-2 glnE gene with 1673 bp immediately ds1133 NO 187 downstream of the ATG start codon deleted, resulting in a truncated glnE, protein lacking the adenylyl-removing (AR) domain none 1021-1623 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1673 bp immediately ds1133 NO 188 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included none 1021-1623 SEQ ID .DELTA.nifL::Prm7 starting at 24 bp after the A of the ATG ds1148 NO 189 start codon, 1375 bp of nifL have been deleted and replaced with the 1021 Prm7 promoter sequence none 1021-1623 SEQ ID .DELTA.nifL::Prm7 with starting at 24 bp after the A of the ATG ds1148 NO 190 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the 1021 rm7 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 137-1034 SEQ ID glnE.DELTA.AR-2 glnE gene with 1290 bp immediately ds809 NO 191 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain none 137-1034 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1290 bp immediately ds809 NO 192 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included none 137-1036 SEQ ID .DELTA.nifL::PinfC starting at 24 bp after the A of the ATG ds799 NO 193 start codon, 1372 bp of nifL have been deleted and replaced with the 137 PinfC promoter sequence none 137-1036 SEQ ID .DELTA.nifL::PinfC with starting at 24 bp after the A of the ATG ds799 NO 194 500 bp flank start codon, 1372 bp of nifL have been deleted and replaced with the 137 PinfC promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 137-1314 SEQ ID glnE.DELTA.AR-2 36 bp glnE gene with 1290 bp immediately none NO 195 deletion downstream of the ATG start codon deleted AND 36 bp deleted beginning at 1472 bp downstream of the start codon, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain none 137-1314 SEQ ID glnE.DELTA.AR-2 36 bp glnE gene with 1290 bp immediately none NO 196 deletion downstream of the ATG start codon deleted AND 36 bp deleted beginning at 1472 bp downstream of the start codon, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the nifL gene upstream and downstream are included none 137-1314 SEQ ID .DELTA.nifL::Prm8.2 starting at 24 bp after the A of the ATG ds857 NO 197 start codon, 1372 bp of nifL have been deleted and replaced with the 137 Prm8.2 promotor sequence none 137-1314 SEQ ID .DELTA.nifL::Prm8.2 with starting at 24 bp after the A of the ATG ds857 NO 198 500 bp flank start codon, 1372 bp of nifL have been deleted and replaced with the 137 Prm8.2 promoter sequence: 500 bp flanking the nifL gene upstream and downstream are included none 137-1329 SEQ ID glnE.DELTA.AR-2 36 bp glnE gene with 1290 bp immediately none NO 199 deletion downstream of the ATG start codon deleted AND 36 bp deleted beginning at 1472 bp downstream of the start codon resulting in a truncated gluE protein lacking the adenylyl-removing (AR) domain none 137-1329 SEQ ID glnE.DELTA.AR-2 36 bp glnE gene with 1290 bp immediately none NO 200 deletion downstream of the ATG start codon deleted AND 36 bp deleted beginning at 1472 bp downstream of the start codon, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the nifL gene upstream and downstream are included none 137-1329 SEQ ID .DELTA.nifL::Prm6.2 starting at 24 bp after the A of the ATG ds853 NO 201 start codon, 1372 bp of nifL have been deleted and replaced with the 137 Prm6.2 promoter sequence none 137-1329 SEQ ID .DELTA.nifL::Prm6.2 with starting at 24 bp after the A of the ATG ds853 NO 202 500 bp flank start codon, 1372 bp of nifL have been deleted and replaced with the 137 Prm6.2 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 137-1382 SEQ ID .DELTA.nifL::Prm1.2 starting at 24 bp after the A of the ATG NO 203 start codon, 1372 bp of nifL have been ds843 deleted and replaced with the 137 Prm1.2 promoter sequence none 137-1382 SEQ ID .DELTA.nifL::Prm1.2 with starting at 24 bp after the A of the ATG ds843 NO 204 500 bp flank start codon, 1372 bp of nifL have been deleted and replaced with the 137 Prm1.2 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 137-1382 SEQ ID glnE.DELTA.AR-2 36 bp glnE gene with 1290 bp immediately none NO 205 deletion downstream of the ATG start codon deleted AND 36 bp deleted beginning at 1472 bp downstream of the start codon, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain none 137-1382 SEQ ID glnE.DELTA.AR-2 36 bp glnE gene with 1290 bp immediately none NO 206 deletion downstream of the ATG start codon deleted AND 36 bp deleted beginning at 1472 bp downstream of the start codon, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the nifL gene upstream and downstream are included none 137-1586 SEQ ID .DELTA.nifL::PinfC starting at 24 bp after the A of the ATG ds799 NO 207 start codon, 1372 bp of nifL have been deleted and replaced with the 137 PinfC promoter sequence none 137-1586 SEQ ID .DELTA.nifL::PinfC with starting at 24 bp after the A of the ATG ds799 NO 208 500 bp flank start codon, 1372 bp of nifL have been deleted and replaced with the 137 PinfC promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 137-1586 SEQ ID glnE.DELTA.AR-2 glnE gene with 1290 bp immediately NO 209 downstream of the ATG start codon deleted, resulting in a truncated glnE ds809 protein lacking the adenylyl-removing (AR) domain none 137-1586 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1290 bp immediately ds809 NO 210 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included none 19-594 SEQ ID glnE.DELTA.AR-2 glnE gene with 1650 bp immediately ds34 NO 211 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain none 19-594 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1650 bp immediately ds34 NO 212 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included none 19-594 SEQ ID .DELTA.nifL::Prm6.1 starting at 221 bp after the A of the ds180 NO 213 ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm6.1 promoter sequence none 19-594 SEQ ID .DELTA.nifL::Prm6.1 with starting at 221 bp after the A of the ds180 NO 214 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm6.1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 19-714 SEQ ID .DELTA.nifL::Prm6.1 starting at 221 bp after the A of the ds180 NO 215 ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm6.1 promoter sequence none 19-714 SEQ ID .DELTA.nifL::Prm6.1 with starting at 221 bp after the A of the ds180 NO 216 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm6.1promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included none 19-715 SEQ ID .DELTA.nifL::Prm7.1 starting at 221 bp after the A of the ds181 NO 217 ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm7.1 promoter sequence none 19-715 SEQ ID .DELTA.nifL::Prm7.1 with starting at 221 bp after the A of the ds181 NO 218 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm76.1promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included 19-713 19-750 SEQ ID .DELTA.nifL::Prm1.2 starting at 221 bp after the A of the ds172 NO 219 ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm1.2 promoter sequence 19-713 19-750 SEQ ID .DELTA.nifL::Prm1.2 with starting at 22l bp after the A of the ds172 NO 220 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm1.2 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included 17-724 19-804 SEQ ID .DELTA.nifL::Prm1.2 starting at 221 bp after the A of the ds172 NO 221 ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm1.2 promoter sequence 17-724 19-804 SEQ ID .DELTA.nifL::Prm1.2 with starting at 221 bp after the A of the ds172 NO 222 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm1.2 promoter sequence;
500 bp flanking the nifL gene upstream and downstream are included 17-724 19-804 SEQ ID glnE.DELTA.AR-2 glnE gene with 1650 bp immediately ds34 NO 223 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain 17-724 19-804 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1650 bp immediately ds34 NO 224 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included 19-590 19-806 SEQ ID .DELTA.nifL::Prm3.1 starting at 22l bp after the A of the ds175 NO 225 ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm3.1 promoter sequence 19-590 19-806 SEQ ID .DELTA.nifL::Prm3.1 with starting at 221 bp after the A of the ds175 NO 226 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the CI019 Prm3.1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included 19-590 19-806 SEQ ID glnE.DELTA.AR-2 glnE gene with 1650 bp immediately ds34 NO 227 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain 19-590 19-806 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1650 bp immediately ds34 NO 228 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included none 63-1146 SEQ ID .DELTA.nifL::PinfC starting at 24 bp after the A of the ATG ds908 NO 229 start codon, 1375 bp of nifL have been deleted and replaced with the 63 PinfC promoter sequence none 63-1146 SEQ ID .DELTA.nifL::PinfC with starting at 24 bp after the A of the ATG ds908 NO 230 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the 63 PinfC 500 bp flank sequence; 500 bp flanking the nifL gene upstream and downstream are included CM015; 6-397 SEQ ID .DELTA.nifL::Prm5 starting at 31 bp after the A of the ATG ds24 PBC6.15 NO 231 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm5 promoter sequence CM015; 6-397 SEQ ID .DELTA.nifL::Prm5 with starting at 31 bp after the A of the ATG ds24 PBC6.15 NO 232 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm5 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included CM014 6-400 SEQ ID .DELTA.nifL:Prm1 starting at 31 bp after the A of the ATG ds20 NO 233 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence CM014 6-400 SEQ ID .DELTA.nifL::Prm1 with starting at 31 bp after the A of the ATG ds20 NO 234 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included CM037; 6-403 SEQ ID .DELTA.nifL::Prm1 starting at 31 bp after the A of the ATG ds20 PBC6.37 NO 235 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence CM037; 6-403 SEQ ID .DELTA.nifL::Prm1 with starting at 3l bp after the A of the ATG ds20 PBC6.38 NO 236 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence: 500 bp flanking the nifL gene upstream and downstream are included CM037; 6-403 SEQ ID glnE.DELTA.AR-2 glnE gene with 1644 bp immediately ds31 PBC6.3-9 NO 237 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain CM037; 6-403 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1644 bp immediately ds31 PBC6.40 NO 238 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included CM038; 6-404 SEQ ID glnE.DELTA.AR-1 glnE gene with 1287 bp immediately ds30 PBC6.38 NO 239 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain CM038; 6-404 SEQ ID .DELTA.nifL::Prm1 starting at 31 bp after the A of the ATG ds20 PBC6.38 NO 240 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence CM038; 6-404 SEQ ID .DELTA.nifL:Prm1 with starting at 31 bp after the A of the ATG ds20 PBC6.38 NO 241 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included CM038; 6-404 SEQ ID glnE.DELTA.AR4 with glnE gene with 1287 bp immediately ds30 PBC6.38 NO 242 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included CM029; 6-412 SEQ ID glnE.DELTA.AR-1 glnE gene with 1287 bp immediately ds30 PBC6.29 No 243 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain CM029; 6-412 SEQ ID glnE.DELTA.AR-1 with glnE gene with 1287 bp immediately ds30 PBC6.29 NO 244 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included CM029; 6-412 SEQ ID .DELTA.nifL::Prm5 starting at 31 bp after the A of the ATG ds24 PBC6.29 NO 245 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm5 promoter sequence CM029; 6-412 SEQ ID .DELTA.nifL::Prm5 with starting at 31 bp after the A of the ATG ds24 PBC6.29 NO 246 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm5 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included CM093; 6-848 SEQ ID .DELTA.nifL::Prm1 starting at 31 bp after the A of the ATG ds20 PBC6.93 NO 247 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence CM093; 6-848 SEQ ID .DELTA.nifL::Prm1 with starting at 31 bp after the A of the ATG ds20 PBC6.93 NO 248 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included CM093; 6-848 SEQ ID glnE.DELTA.AR-2 glnE gene with 1644 bp immediately ds31 PBC6.93 NO 249 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain CM093; 6-848 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1644 bp immediately ds31 PBC6.93 NO 250 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included CM093; 6-848 SEQ ID .DELTA.amtB First 1088 bp of amtB gene and 4 bp ds126 PBC6.93 NO 251 upstream of start codon deleted; 199 bp of gene remaining lacks a start endow no amtB protein is translated CM093; 6-848 SEQ ID .DELTA.amtB with 500 bp First 1088 bp of amtB gene and 4 bp ds126 PBC6.93 NO 252 flank upstream of start codon deleted; 199 bp of gene remaining lacks a start codon; no amtB protein is translated CM094; 6-881 SEQ ID glnE.DELTA.AR-1 glnE gene with 1287 bp immediately ds30 PBC6.94 NO 253 downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain CM094; 6-881 SEQ ID glnE.DELTA.AR-1 with glnE gene with 1287 bp immediately ds30 PBC6.94 NO 254 500 bp flank downstream of the ATG start codon deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the gthE gene upstream and downstream are included CM094; 6-881 SEQ ID .DELTA.nifL::Prm1 starting at 31 bp after the A of the ATG ds20 PBC6.94 NO 255 start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence CM094; 6-881 SEQ ID .DELTA.nifL::Prm1 with starting at 31 bp after the A of the ATG ds20 PBC6.94 NO 256 500 bp flank start codon, 1375 bp of nifL have been deleted and replaced with the CI006 Prm1 promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included CM094; 6-881 SEQ ID .DELTA.amtB First 1088 bp of amtB gene and 4 bp ds126 PBC6.94 NO 257 upstream of start codon deleted; 199 bp of gene remaining lacks a start codon; no amtB protein is translated CM094; 6-881 SEQ ID .DELTA.amtB with 500 bp First 1088 bp of amtB gene and 4 bp ds126 PBC6.94 NO 258 flank upstream of start codon deleted; 199 bp of gene remaining lacks a start codon; no amtB protein is translated none 910-1246 SEQ ID .DELTA.nifL::PinfC starting at 20 bp after the A of the ATG ds960 NO 259 start codon, 1379 bp of nifL have been deleted and replaced with the 910 PinfC promoter sequence none 910-1246 SEQ ID .DELTA.nifL::PinfC with starting at 20 bp after the A of the ATG ds960 NO 260 500 bp flank start codon, 1379 bp of nifL have been deleted and replaced with the 910 PinfC promoter sequence; 500 bp flanking the nifL gene upstream and downstream are included PBC6.1, CI006 SEQ ID 16S-1 1 of 3 unique 16S rDNA genes in the N/A 6, CI6 NO 261 CI006 genome PBC6.1, CI006 SEQ ID 16S-2 2 of 3 unique 16S rDNA genes in the N/A 6, CI6 NO 262 CI006 genome PBC6.1, CI006 SEQ ID nifH N/A N/A 6, CI6 NO 263 PBC6.1, CI006 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A 6, CI6 NO 264 in CI006 genome PBC6.1, CI006 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A 6, CI6 NO 265 in CI006 genome PBC6.1, CI006 SEQ ID nifL N/A N/A
6, CI6 NO 266 PBC6.1, CI006 SEQ ID nifA N/A N/A 6, CI6 NO 267 PBC6.1, CI006 SEQ ID glnE N/A N/A 6, CI6 NO 268 PBC6.1, CI006 SEQ ID 16S-3 3 of 3 unique 16S rDNA genes in the N/A 6, CI6 NO 269 CI006 genome PBC6.1, CI006 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A 6, CI6 NO 270 in CI006 genome PBC6.1, CI006 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A 6, CI6 NO 271 in CI006 genome PBC6.1, CI006 SEQ ID amtB N/A N/A 6, CI6 NO 272 PBC6.1, CI006 SEQ ID Prm1 348 bp includes the 319 bp immediately N/A 6, CI6 NO 273 upstream of the ATG start codon of the lpp gene and the first 29 bp of the lpp gene PBC6.1, CI006 SEQ ID Prm5 313 bp starting at 432 bp upstream of the N/A 6, CI6 NO 274 ATG start codon of the ompX gene and ending 119 bp upstream of the ATG start codon of the ompX gene 19, CI19 CI019 SEQ ID nifL N/A N/A NO 275 19, CI19 CI019 SEQ ID nifA N/A N/A NO 276 19, CI19 CI019 SEQ ID 16S-1 1 of 7 unique 16S rDNA genes in the N/A NO 277 CI019 genome 19, CI19 CI019 SEQ ID 16S-2 2 of 7 unique 16S rDNA genes in the N/A NO 278 CI019 genome 19, CI19 CI019 SEQ ID 16S-3 3 of 7 unique 16S rDNA genes in the N/A NO 279 CI019 genome 19, CI19 CI019 SEQ ID 16S-4 4 of 7 unique 16S rDNA genes in the N/A NO 280 CI019 genome 19, CI19 CI019 SEQ ID 16S-5 5 of 7 unique 16S rDNA genes in the N/A NO 281 CI019 genome 19, CI19 CI019 SEQ ID 16S-6 6 of 7 unique 16S rDNA genes in the N/A NO 282 CI019 genome 19, CI19 CI019 SEQ ID 16S-7 7 of 7 unique 16S rDNA genes in the N/A NO 283 CI019 genome 19, CI19 CI019 SEQ ID nifH1 1 of 2 unique genes annotated as nifH N/A NO 284 in CI019 genome 19, CI19 CI019 SEQ ID nifH2 2 of 2 unique genes annotated as nifH N/A NO 285 in CI019 genome 19, CI19 CI019 SEQ ID nifD1 1 of 2 unique genes annotated as nifD N/A NO 286 in CI019 genome 19, CI19 CI019 SEQ ID nifD2 2 of 2 unique genes annotated as nifD N/A NO 287 in CI019 genome 19, CI19 CI019 SEQ ID nifK1 1 of 2 unique genes annotated as nifK N/A NO 288 in CI019 19, CI19 CI019 SEQ ID nifK2 2 of 2 unique genes annotated as nifK N/A NO 289 in CI019 genome 19, CI19 CI019 SEQ ID glnE N/A N/A NO 290 19, CI19 CI019 SEQ ID prm4 449 bp immediately upstream of the N/A NO 291 ATG of the dscC 2 gene 19, CI19 CI019 SEQ ID Prm1.2 500 bp immediately upstream of the N/A NO 292 TTG start codon of the infC gene 19, CI19 CI019 SEQ ID Prm3.1 170 bp immediately upstream of the N/A NO 293 ATG start codon of the rplN gene 19, CI20 CI020 SEQ ID Prm6.1 142 bp immediately upstream of the N/A NO 294 ATG of a highly-expressed hypothetical protein (annotated as PROKKA_00662 in CI019 assembly 82) 19, CI21 CI021 SEQ ID Prm7.1 293 bp immediately upstream of the N/A NO 295 ATG of the lpp gene 19-375, CM67 SEQ ID glnE.DELTA.AR-2 glnE gene with 1650 bp immediately ds34 19-417, NO 296 downstream of the ATG start codon CM067 deleted, resulting in a truncated glnE protein lacking the adenylyl-removing (AR) domain 19-375, CM67 SEQ ID glnE.DELTA.AR-2 with glnE gene with 1650 bp immediately ds34 19-417, NO 297 500 bp flank downstream of the ATG start codon CM067 deleted, resulting in a truncated g1nE protein lacking the adenylyl-removing (AR) domain; 500 bp flanking the glnE gene upstream and downstream are included 19-375, CM67 SEQ ID .DELTA.nifL::null-v1 starting at 221 bp after the A of the none 19-417, NO 298 ATG start codon, 845 bp of nifL have CM067 been deleted and replaced with the 31 bp sequence "GGAGTCTGAACTCATCCTGCGA TGGGGGCTG" 19-375, CM467 SEQ ID .DELTA.nifL::null-v1 with starting at 221 bp after the A of the none 19-417, NO 299 500 bp flank ATG start codon, 845 bp of nifL have CM067 been deleted and replaced with the 31 bp sequence "GGAGTCTGAACTCATCCTGCGA TGGGGGCTG"; 500 bp flanking the nifL gene upstream and downstream are included 19-377, CM69 SEQ ID .DELTA.nifL::null-v2 starting at 221 bp after the A of the none CM069 NO 300 ATG start codon, 845 bp of nifL have been deleted and replaced with the 5 bp sequence "TTAAA" 19-377 CM69 SEQ ID .DELTA.nifL::null-v2 with starting at 221 bp after the A of the none CM069 NO 301 500 bp flank ATG start codon, 845 bp of nifL have been deleted and replaced with the 5 bp sequence "TTAAA"; 500 bp flanking the nifL gene upstream and downstream are included 19-389, CM81 SEQ ID .DELTA.nifL::Prm4 starting at 221 bp after the A of the ds70 19-418, NO 302 ATG start codon, 845 bp of nifL have CM081 been deleted and replaced with the CI19 Prm4 sequence 19-389, CM81 SEQ ID .DELTA.nifL::Prm4 with starting at 221 bp after the A of the ds70 19-418, NO 303 500 bp flank ATG start codon, 845 bp of nifL have CM081 been deleted and replaced with the CI19 Prm4 sequence; 500 bp flanking the nifL gene upstream and downstream are included
TABLE-US-00022 TABLE H SEQ ID NO: Sequence 61 atgttttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctgagagctggcgcg- aattgtggcaggatgcgttgcaggagga ggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgctgattgccgatttt- cgcaaagagttggataaacgcacc attggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgtatgctcgcgcgacg- atgcgccagtaccgctgtcacgcc tgacgccgctgctcaccggaattattacccgcaccacttaccttgagagctaagtgaatttcccggcgcactg- aaacacctcatttccctgtgtgccgc gtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcgacccgaatacgctc- tatcaaccgacggcgatgaatgccta tcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagcttgaggcgctgcgg- cagtttaagcaggcgcagttgct gcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcacttaacctggctggcg- gaagcgattattgatgcggtggtg cagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgcacgatcgcgaagggcgcggtt- ttgcggtggtcggttatggcaag ctgggcggctgggagctgggttacagctccgatctggatctggtattcctgcacgactgcccgatggatgtga- tgaccgatggcgagcgtgaaatcg atggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgcacgtcgtccggcat- cctttatgaagttgatgcgcgtctgcgt ccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagcaaaacgaagcctgga- cgtgggaacatcaggcgctggcc cgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgccgcgatattctgatga- cgcctcgcgacggcgcaacgctgc aaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaagaccgcttcgatct- gaaagccgatgaaggcggtatca ccgacatcgagtttatcgcccaatatctggtgctgugctttgcccatgacaagccgaaactgacgcgctggtc- ggataatgcgcattctcgaagggc tggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctggcgtacaccacattgcgtgatga- gctgcaccacctggcgctgcaa gagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttattaaaaccagctgggacaagt- ggctggtggaaccgtgcgccccggc gtaa 62 ttgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgaacggtagca- cagagagcttgctctcgggtgacgagt ggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaatacc- gcataacgtcgcaagaccaaaga gggggaccttcgggcctcttgccatcagaggcccagatgggattagctagtaggtggggtaacggctcaccta- ggcgacgatccctagctggtag agaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatattgc- acaatgggcgcaagcctgatgc agccatgccgcggtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaagggagtaaggttaa- taaccttattcattgacgttacccg cagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaattac- tgggcgtaaagcgcacgcaggc ggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcatccgaaactggcaggcttgagtct- cgtagagggaggtagaattccagg tgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggcctcctggacgaagactgacgc- tcaggtgcgaaagcgtgggga gcaaacaggattagataccctggtagtccacgccgtaaacgatgtctatttggaggttgtgcccttgaggcgt- ggcttccggagctaacgcgttaaata gaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggcgagcat- gtggtttaattcgatgcaacgc gaagaaccttacctggtcttgacatccacagaactttccagagatggattggtgccttcgggaactgtgagac- aggtgctgcatggctgtcgtcagctc gtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcggtccggccgcg- aactcaaaggagactgccagtgataa actggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccagggctacacacgtgctacaatggc- gcatacaaagagaagcgacctcg cgagagtaagcggacctcataaagtgcgtcgtagtccggattggagtctgcaactcgactccatgaagtcgga- atcgctagtaatcgtggatcagaat gccacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaagaagta- ggtagcttaaccttcgggagggc gcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttggatc- acctcctt 63 ttgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgaacggtagca- cagagagcttgctctcgggtgacgagt ggcggacgggtgagtaatgtcgggaaactgcctgatggagggggataactactggaaacggtagctaataccg- cataacgtcgcaagaccaaaga gggggaccttcgggcctcttgccatcagatgtgcccagatgggattagctagtaggtggggtaacggctcacc- taggcgacgatccctagctggtctg agaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatattgc- acaatgggcgcaagcctgatgc agccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaggnantanggttaa- taacctgtgttnattgacgttacc cgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaatt- actgggcgtaaagcgcacgcag gcggtctgtcaagtcggatgtgaaatccccgggctcaacctgggactgcatccgaaactggcaggcttgagtc- tcgtagagggaggtagaattcca ggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggcctcctggacgaagactgac- gctcaggtgcgaaagcgtggg gagcaaacaggattagataccctggtagtccacgccgtaaacgatgtctatttggaggttgtgcccttgaggc- gtggcttccggagctaacgcgttaaa tagaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtggag- catgtggtttaattcgatgcaac gcgaagaaccttacctggtcttgacatccacagaacttagcagagatgctttggtgccttcgggaactgtgag- acaggtgctgcatggctgtcgtcagc tcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcggttaggccg- ggaactcaaaggagactgccagtgat aaactggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccacggctacacacgtgctacaatg- gcgcatacaaagagaagcgacct cgcgagagtaagcggacctcataaagtgcgtcgtagtccggattggagtctgcaactcgactccatgaagtcg- gaatcgctagtaatcgtggatcaga atgccacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaagaag- taggtagcttaaccttcgggaggg cgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttggat- cacctcctt 64 attgaagagtttgatcatggctcagattgaacgctggcggtcaggcctaacacatgcaagtcgagcggcan- cgggaagtagcttgctactttgccggc gagcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaa- taccgcatgacctcgaaagagc aaagtggggatcttcggacctcacgccatcggatgtgcccagatgggattagctagtaggtgaggtaatggct- nacctaggcgacgatccctagct ggtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaa- tattgcacaatgggcgcaagc ctgatgcagccatgccgcgtgtgtgaagaaggccttagggttgtaaagcactttcagcgaggaggaggcanca- nacttaatacgtgtgntgattgac gttactcgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatc- ggaattactgggcgtaaagcgca cgcaggcggtttgttaagtcagatgtgaaatccccgagcttaacttcgggaactgcatttgaaactggcaagc- tagagtcttgtagaggggggtagaatt ccaggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagact- gacgctcaggtgcgaaagcgt ggggagcaattcaggattagataccctggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttg- aggcgtggcttccggagctaacgcgt taagtcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggt- ggagcatgtggtttaattcgatgc aacgcgaagaaccttacctactcttgacatccagagaatttgccagagatggcgaagtgccttcgggaactct- gagacaggtgctgcatggctgcgt cagctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcacgtna- tggtgggaactcaaaggagactgccg gtgataaaccggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacgtgcta- caatggcatatacaaagagaagc gaactcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtagcaactcgactccatgaa- gtcggaatcgctagtaacgtaga tcagaatgctacggtgaatacgttccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaa- gaagtaggtagcttaaccttcggg agggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggtt- ggatcacctcctt 65 atgaccatgcgtcaatgcgccatttacggcaaaggtgggatcggcaaatcgaccaccacacagaacctggt- cgccgcgctggcggagatgggtaaa aaagtcatgattgtcggctgtgacccgaaagccgattccacgcgtttgatcctgcatgcgaaagcgcagaaca- ccattatggagatggctgctgaagt cggctccgtggaagacctggagttagaagacgtgctgcaaatcggttacggcggcgtgcgctgcgcagagtcc- ggcggcccggagccaggcgtg ggctgtgccggtcgcggggtgatcaccgcgattaacttcctcgaagaagaaggcgcttacgtgccggatctcg- attttgttttctacgacgtgctgggc gacgtggtatgcggtggtttcgccatgccgattcgtgaaaacaaagcgcaggagatctacatcgtttgctctg- gcgaaatgatggcgatgtacgccgc caacaacatctccaaaggcatcgtgaaatacgccaaatccggtaaagtgcgcctcggcgggctgatttgtaac- tcgcgccagaccgaccgtgaagat gaactgatcattgcgctggcagaaaaactcggcacgcagatgatccactttgttcccgcgacaacattgtgca- gcgtgcggaaatccgccgtatgac ggttatcgaatatgacccgacctgcaatcaggcgaacgaatatcgcagccttgccagcaaaatcgtcaacaac- accaaaatggtggtgcccaccccc tgcaccatggatgaactggaagaactgctgatggagttcggcattatggatgtggaagacaccagcatcattg- gtaaaaccgccgccgaagaaaacg ccgtctga 66 atgagcaatgcaacaggcgaacgcaacctggagataatcgagcaggtgctcgaggttttcccggagaagac- gcgcaaagaacgcagaaaacacat gatggtgacggacccggagcaggaaagcgtcggtaagtgcatcatctctaaccgcaaatcgcagccaggcgtg- atgaccgtgcgcggctgctcgta tgccggttcgaaaggggtggtatttgggccaatcaaggatatggcgcatatctcgcatggcccaatcggctgc- ggccaatactcccgcgccgggcgg cggactactacaccggcgtcagcggcgtggacagcttcggcacgctcaacttcacctccgattttcaggagcg- cgacatcgtgtttggcggcgataa aaagctcgccaaactgattgaagagctggaagagctgttcccgctgaccaaaggcatttcgattcagtcggaa- tgcccggtcggcctgattggcgatg acattgaggccgtcgcgaacgccagccgcaaagccatcaacaaaccggttattccggtgcgttgcgaaggctt- tcgcggcgtgtcgcaatccctcgg tcaccatattgccaacgatgtgatccgcgactgggtgctggataaccgcgaaggcaaaccgttcgaatccacc- ccttacgatgtggcgatcatcggcg attacaacatcggcggcgatgcctgggcttcgagcattttgctcgaagagatgggcttgcgggtggtggcaca- gtggtctggcgacggtacgctggt ggagatggaaaacacgccgttcgtcaaactgaacctggtgcattgttaccgctcaatgaactacatctcgcgc- catatggaggagaagcacggtattc cgtggatggaatacaacttcttggtccgacgaaaatcgcggaatcgctgcgcaaaatcgccgaccagtttgac- gacaccattcgcgccaacgccga agcggtgatcgccagataccaggcgcaaaacgacgccattatcgccaaatatcgcccgcgtctggaggggcgc- aaagtgctgctttatatgggcgg gctgcgtccgcgccatgtgattggcgcctatgaagacctgggaatggagatcatcgctgccggttatgagttc- ggtcataacgatgattacgaccgca ccttgccggatctgaaagagggcacgctgctgtttgatgatgccagcagttatgagaggaggcgttcgtcaac- gcgctgaaaccggatctcatcggtt ccggcatcaaagagaagtacatctttcagaaaatgggcgtgccgtttcgccagatgcactcctgggattactc- cggcccgtaccacggctatgacggc ttcgccatcttcgcccgcgatatggatatgacgctcaacaaccccgcgtggggccagttgaccgcgccgtggc- tgaaatccgcctga 67 atgagccagactgctgagaaaatacagaattgccatcccctgtttgaacaggatgcttaccagacgctgtt- tgccggtaaacgggcactcgaagaggc gcactcgccggagcgggtgcaggagtgtttcaatggaccactaccccggaatatgaagcgctgaactttaaac- gcgaagcgctgactatccaccc ggcaaaagcctgccagccgctgggcgcggtgctctgttcgctggggtttgccaataccctaccgtatgtgcac- gcttcacaggttgcgtggcctattt ccgcacgtactttaaccgccactttaaagaaccggtggcctgcgtgtcggattcaatgacggaagacgcggcg- gtgttcggcgggaataacaacctc aacaccggcttacaaaacgccagcgcgctgtataaaccggagattatcgccgtctctaccacctgtatggcgg- aagtgatcggtgatgatttgcaggc ctttatcgccaacgccaaaaaagatggttttctcgatgccgccatccccgtgccctacgcgcacacccccagt- tttatcggcagccatatcaccggctg ggataacatgtttgaaggttttgcccggacctttacggagaccatgaagctcagcccggcaaactttcacgca- tcaacctggtgaccgggtttgaaac ctatctcggcaatttccgcgtgctgaaacgcatgatggaacaaatggaggtgccggcgagtggctctccgatc- cgtcggaagtgctgcatactcccg ccaacgggcattaccagatgtacgcgggcgggacgacgcagcaagagatgcgcgaggcgccggatgctatcga- caccctgttcctgcagccctg gcaactggtgaaaagcaaaaaagtggtgcaggagatgtggaatcagcccgccaccgaggtttctgttcccgtt- gggctggcaggaacacacgaact gttgatggcgattagccagttaaccggcaaggccattcccgattcactggcgctggagcgccggcggctggtc- gatatgatgctcgattcccacacct ggttgcacggtaaaaaattcggcctgaggcgatccggattttgtcatgggattgacccgtttcctgctggagc- tgggctgcgaaccgaccgttatcctc tgccacaacggtaacaaacgctggcagaaagcaatgaagaaaatgcttgacgcctcgccgtacggccaggaga- gcgaagtttatcaactgcgatt tgtggcatttccgctcgctgatgtttacccgccagccggattttatgattggcaactcgtacggcaagttcat- tcagcgcgacaccttagccaaaggcga gcagtttgaagttccgctgatccgcctcggttttccccttcgaccgccaccatctgcaccgccagaccacctg- gggctacgagggcgccatgagca ttctcactacccttgtgaatgcggtactggagaaagtggacaaagagaccatcaagctcggcaaaaccgacta- cagcttcgatcttatccgttaa 68 atgaccctgaatatgatgatggatgccggcgcgcccgaggcaatccccggtgcgctttcgcgacaccatcc- tgggctgttttttaccatcgttgaagaa gcgcccgtcgccatttcgctgactgatgccgacgcacgcattgtctatgccaacccggctttctgccgccaga- ccggctatgaactagaagcgttgttg cagcaaaatccccgcctgatgcaagtcgccaaaccccacgggaaatctatcaggatatgtggcacaccttgtt- acaacgccgaccgtggcgcgggc aattgattaaccgccaccgcgacggcagcctgatctggtcgagatcgatatcaccccggtgattaacccgttt- ggcgaactggaacactacctggca atgcagcgcgatatcagcgccagttatgcgctggagcagcggttgcccaatcacatgacgctgaccgaagcgg- tgctgaataacattccggcggcg gtggttgtagtggatgaacgcgatcatgtggttatggataaccttgcctacaaaacgttctgtgccgactgcg- gcggaaaagagctcctgagcgaactc aatttttcagcccgaaaagcggagctggcaaacggccagctcttaccggtggtgctgcgcggtgaggtgcgct- ggttgtcggtgacctgctgggcgc tgccgcgcgtcagcgaagaagccagcgctactttattgataacaggctgacgcgcacgctggggtgatcaccg- acgacacccaacaacgccagc agcaggaacagggccgacttgaccgccttaaacagcagatgaccaacggcaaactactggcagcgatccgcga- agcgcttgacgccgcgctgatc cagcttaactgccccatcaatatgctggcggcggcgcgacgtttaaacggcagtgataacaacaatgtggcgc- tcgacgccgcgtggcgcgaaggt gaagaggcgatggcgcggctgaaacgttgccgcccgtcgctggaactggaaaggcggccgtctggccgctgca- acccattttgacgatctgcgc gcgctttatcttcacccgctacgagcaggggaaaaatttgcaggtcacgctggattcccatcatctggtggga- tttggtcagcgtacgcaactgttagcct gcctgagtctgtggctcgatcgcacgctggatattgccgccgggctgggtgatttcaccgcgcaaacgcagat- ttacgcccgcgaagaagagggctg gctactttgtatatcactgacaatgtgccgagatcccgctgcgccacacccactcgccggatgcgcttaacgc-
tccgggaaaaggcatggagctgc gcctgatccagacgctggtggcacaccaccacggcgcaatagaactcacttcacaccccgaagggggaagttg- cctgaccctacgattcccgctatt tcattcactgaccgcaggttcaaaatga 69 atgacccagcgaaccgagtcgggtaataccgtctggcgatcgatttgtcccagcagttcactgcgatgcag- cgcataaggtggtactcagccggg cgaccgaggtcgatcagacgctccagcaagtgctgtgcgtattgcacaatgacgcctttttgcagcacggcat- gatctgctgtacgacagccagcag gcgattttgaatattgaagcgttgcaggaagccgatcagcagttaatccccggcagctcgcaaatccgctatc- gtccgggcgaagggctggtcggga cggtgctttcgcagggccaatcattaggctggcgcgcgttgctgacgatcagcgctttcttgaccggctcggg- ttgtatgattacaacctgccgtttatc gccgtgccgctgatagggccagatgcgcagactttcggtgtgagacggcacaacccatggcgcgttacgaaga- gcgattacccgcctgcacccgc tttctggaaacggtcgctaacctggtcgcgcaaaccgtgcgtttgatggcaccaccggcagtgcgcccttccc- cgcgcgccgccataacacaggccg ccagcccgattatcctgcacggcctcacgcgcatttggttttgaaaatatggtcggtaacagtccggcgatgc- gccagaccatggagattatccgcag gtttcgcgctgggacaccaccgttctggtacgcggcgagagtggcaccggcaaggagctgattgccaacgcca- tccaccaccattcgccgcgtgcc ggtgcgccatttgtgaaattcaactgtgcggcgctgccggacacactgctggaaagcgaattgttcggtcacg- agaaaggggcatttaccggcgcgg tacgccagcgtaaaggccgttttgagctggccgatggcggcacgctgtttcttgacgagatcggcgagagtag- cgcctcgtttcaggctaagctgctg cgcattttgcaggaaggcgaagggaacgcgtcggcggcgacgagacattgcaagtgatttgtgcgcattattg- ccgcgacgaaccgcaatcttgaa gatgaagtccggctggggcactttcgcgaagatctctattatcgcctgaatgtgatgcccatcgccctgccgc- cactacgcgaacgccaggaggacat tgccgagctggcgcactactggtgcgtaaaatcgcccataaccagagccgtacgctgcgcattagcgagggcg- ctatccgcctgctgatgagctaca actggcccggtaatgtgcgcgaactggaaaactgcatgagcgctcagcggtgatgtcggagaacggtctgatc- gatcgggatgtgattttgtttaatc atcgcgaccagccagccaaaccgccagttatcaggtctcgcatgatgataactggctcgataacaaccttgac- gagcgccagcggctgattgcggc gctggaaaaagcgggatgggtacaagccaaagccgcgcgcttgctggggatgacgccgcgccaggtcgcctat- cgtattcagacgatggatataac cctgccaaggctataa 70 atggcaatgcgtcaatgtgcaatctacgggaaagggggtattggtaaatccaccactacccaaaaccttgt- agcggctctggccgaaatgaataagaa ggtcatgatcgtcggctgtgaccctaaggctgattcaacccgactcattctgcatgcgaaagcacagaacacc- atcatggaaatggccgctgaagtgg gctccgtggtagatctggagctggaagatgtgatgcaaatcggctatggcggcgtgcgctggcggaatcaggc- ggccctgagcctggtgtgggtt gtgccggacgcgggggtgatcaccgccatcaacttcctcgaagaagaaggcgcgtatgtgccggatctggatt- tcgtgttttacgacgtattgggcgat gtggtctgtggcggtttcgcgatgccaattcgcgaaaacaaagcgcaggaaatctacatcgtatgctccggtg- aaatgatggcgatgtatgccgccaa caacatttccaaaggcatcgtgaaatacgcgaaatcgggcaaagttcgcctggccgggctgatctgtaactcc- cgccagacggatcgcgaagatgaa ctgatcatcgcgaggctgaaaaacttggcacgcaaatgatccacttcgtgccgcgtgacaacattgtgcaacg- cgctgaaatccgccgcatgacggt catcgaatacgacccgacttgtgcgcaggcagatcagtcgtgcactggcgaacaaaatcgtcaacaacaccaa- aatggtggtgccgacaccggtc accatggatgagctggaagccctgttaatggaatttggcattatggaagaagaagacctgaccatcgtcggtc- gtaccgccgccgaagaggcgtga 71 atgaccagtgaaacacgcgaacgtaacgaggcattgatccaggaagtgctggagatcttccccgagaaggc- gcttaaagatcgtaagaaacacatg atgaccaccgacccggcgatggaatctgtcggcaaggtattgtctcaaaccgcaaatcacagccgggcgtgat- gaccgtgcgaggctgcgcttacg ccggttccaaaggcgtggtctttggcccgatcaaagacatggcgcatatctcccacggcccggttggttgcgg- ccagtattctcgtgccggacgccgt aactattacaccggctggagggcgtgaacagctttggcaccctcaacttcaccagtgattttcaggaacggga- catcgtatttggcggcgataaaaag ctcgacaaactgatcgacgaactggagatgttgtcccgctgaccaaaggcatttcggtacagtcggaatgtcc- ggtcggtctgatcggcgatgacatt tctgccgtcgccaaagccagcagcgccaaaatcggtaagccggtcgtgccggtacgctgcgaggggttccgcg- gtgtgtcgcaatcgctcggccat cacattgctaacgatgtcatccgcgactgggtgctggataaccgcgaaggcaatgaatttgaaaccacgcctt- acgacgtggcgattatcggcgacta caacatcggcggtgacgcctgggcctcacgtattctgctcgaagaaatggggctgcgtgtggtggcgcagtgg- tccggcgacggcacgctggtgga gatggaaaacaccccgaaagtcgcactcaatctggtgcactgctaccgctcgatgaactacatctcccgcata- tggaagaaaaacacggcattccgt ggatggaatacaacttctttggcccgtccaaaattgcggaatctctgcgcgaaatcgcggcgcgttttgacga- taccatccggaaaaacgccgaagc ggtgattgaaaaatatcaggcgcaaacgcaggcggtgatcgacaaataccgtccgcgtaggaaggcaaaaagg- tgctgttgtatctcggcggtttac gtccgcgccacatcatcggggcgtatgaagatctgggaatggaaatcatcggtaccggctatgaattcggtca- taacgatgattacgaccgcaccttac cgatgctcaaagaaggcacgttgctgttcgatgacctgtgcagttatgagctggaagcgttcgttaaagcgct- gaaaccggatcttgtcgggtcaggc atcaaagaaaaatacattttccacaaaatgggcgtgccgttccgccagatgcactcctgggattattccggcc- cttatcacggctacgacggtttcggca tttttgcccgtgacatggacatgacgctgaacaatccgggctggagtcagctgaccgccccctggttgaaatc- ggcctga 72 atgagtcaagatcttggcaccccaaaatcctgtttcccgctgacgagcaggatgaataccagagtatgttt- acccacaaacgcgcgaggaagaagca cacggcgaggcgatagtgcgggaagtgtttgaatggaccaccacgcaggaatatcaggatctgaacttctcgc- gtgaagcgctgaccgtcgacccg gcgaaagcctgccagccgttaggcgcggtactttgcgcgctgggttttgccaacacgttgccgtatgtccacg- gttcacaaggctgtgtggcgtatttc cgtacctattttaatcctcatttcaaagagccggtggcctgtgtttccgactcaatgaccgaagatgccgccg- tttttggcggaaataacaacatgaatgtc ggtctggaaaacgccagcgcgctgtacaagccgganattattgctgtctccaccacctgtatggcggaagtga- tcggtgatgacctgcaggcttttatc gccaacgccaaaaaagacggatttgtggatgccgggatgccaatcccgtatgcccatacaccgagtttctggg- cagtcatgtcaccggctgggacaa catgtttgaaggcttcgcccgtacctttaccaccgacgccacgcgggaatatcagccgggcaaacttgccaaa- ctgaacgtggtgaccggttttgaaa cttatctcggcaactaccgggttattcaccgcatgatgagccagatgggggtcgaatgcagcgtcttgtccga- tccgtctgaagtgctcgacaccccgg ctgacggccaataccgcatgtatgccggcggcaccacgcaaaccgaaatgcgtgatgcaccggatgccatcga- caccttgctgctgcaaccgtggc aattgcagaaaaccaaaaaagtggtgcagggcgactggaatcagccgggcaccgaagtcagtgtaccgattgg- cctggcggcgaccgatgccttg ctgatgacggtangcgaactgaccggcaaaccgatagctgacacgctggcgactgaacgtggccgtctggtgg- acatgatgctcgattcccacacct ggctgcatggcaagcgtttcggtctctacggtgacccggattttgtgatgggcatgaccgcattcctgctgga- actgggctgtgaaccgaccaccattct cagccataacggcaacaaacgctggcagaaagccatgaagaaaatgctggctgattcgccttacgggcaggac- agcgaagtgtattgtgaactgcga tctgtggcatttccgctcgctgatgtttacccgtaaaccggactttatgatcggcaactcaacggaaaattca- ttcagcgtgacacgctggccaaaggcg aacagttcgaagtgccgctgatccgcatcggttttccgatttttgaccggcaccatttgcaccgtcagaccac- ctggggatacgaaggggcgatgagc atactgacgcaactggtgaatgccgtacttgaacaactggatcgcgaaaccatgaagctcggcaaaaccgact- acaacttcgtcctgttccgctaa 73 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatcc- ttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcanctgccgccagacggg- ttttgcacttgagacacttttgggcg agaaccaccgtctgctggccagccagcagacgccgaaacatatctatgacgaaatgtggcgcactttgagcag- ggcaaatcctggaacggccaact gatcaaccggcgtaataaccgttcgctttatctggcggatgtcactatcacgcctgttttaggcgggacgggc- aggtggagcattacctcggcatgca caaagatatcagcgagaaatacgcgctggaacagcggttgcgcaaccacatcaccttgttcacggaggtgctg- aacaatattcccgccgccgtggtg gtggtggatgagcaggacaatgtggtgatggacaatctggcctacaaaaccctttgcgcggactgcggcggca- aagagctgctggctgaaatgggc tatccgcaactcaaagagatgctcancagtggcgaaccggtgccggtttccatgcgcggcaacgtacgctggt- tttctttcggtcaatggttattcagg gcgttaatgattgaggccagccgcttttttaccggcattaccgcgccgggaaaactgattgttctgaccgact- gcaccgatcagcatcaccggcagcag cagggttatcttgaccggcttaagcaaaaactcaccaacggcaaattattggcggccatccgtgagtcgctcg- atgccgcgcttatccagctcaacgg gccaatcaatatgctggcggctccgcgtcgtcttaacggcgaagaaggcaacaacatggcgctggaattcgcc- tggcgcgaaggcgagcaggcgg tgagtcgcttacaggcctgccgtccgtcgctggattttgagccgcaggcagaatggccggtcagtgaattctt- tgacgatctgagcgcgctgtacgaca gccattttctcagtgacggtgaattgcgttacgtggtcatgccatctgatctgcacgctgtcgggcaacgaac- gcaaatccttaccgcgctgagcttatg gattgatcacacgctgtcacaggcgcaggccatgccgtctctgaagctctcggtgaacattgcgcgaggcagg- atgcgagctggttgtgttttgacatt accgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaagcggcgttttcccgtccggggaatggca- tggagctgcgccttatccagacgc tgatcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggcaccttgctgacgttacgcct- gccggtacagcaggttatcaccggag gcttaaatga 74 atgacccagttcctaccgcgggcccggttatccggcgctttgatatgtctgcccagtttacggcgctttat- cgcatcagcgtggcgctgagtcaggaa agcaacaccgggcgcgcactggcggcgatcctcgaagtgcttcacgatcatgcatttatgcaatacggcatgg- tgtgtctgtttgataaagaacgcaat gcactctttgtggaatccctgcatggcatcgacggcgaaaggaaaaaagagacccgccatgtccgttaccgca- tgggggaaggcgtgatcggcgcg gtgatgagccagcgtcaggcgctcgtgttaccgcgcatttcagacgatcagcgttttctcgaccgcctgaata- tttacgattacagccttccgttgattgg cgtgcccgatccccggtgcggataatcagccatcgggcgtgctggtggcacagccgatggcgttgcacgaaga- ccggctgactgccagtacgcggtt tttagaaatggtcgccaatctcatcagccagccactgcgttctgccacgcccccggaatcattgcctgctcaa- acgccggtccggtgcagtgttccgcg ccagtttggtttcgagcagatggtcgggaaaagtcaggcgatgcgccagacgatggacattttacggcaggtt- tccaaatgggataccacggttctggt gcgtggtgaaagcggcaccggcaaggaacttatcgccaatgccattcattacaactcaccccgtgcggccgcg- ccatttgtgaaattcaactgcgccg cgctgccggataacctgctggaaagcgaactgttcggtcatgaaaaaggggccttcaccggcgctatccgtac- ccgtaaaggccgctttgaactggc ggacgggggcacgttattcctcgatgaaatcggcgaatcgagcgcgtcgtttcaggccaaattgctgcgcatt- ttgcaggaaggtgaaatggaacgg gtcggcggcgataccacgctgaaagttgatgtgcgcattattgctgccaccaaccgtaatcttgaagaggaag- tgcgtgccgggaattttcgcgaaga cctgtattatcgcctgaacgtgatgccggtttcgctgcctgcactgcgtgaaaggctggatgatatcgccgat- ctggcgccgtttaggtcaattaagatt gcgctgcgtcaggggcgggaactgcgcatcagcgacggtgcggtgcgtctgctgatgacctacagctggccag- gcaacgtgcgtgaactggaaaa ctgtctcgaacgggcgtcggtaatgaccgatgaagggctgatcgaccgcgacgtgatcctgttcaatcaccat- gaatccccggcgctgtccgtcaaac ccggcctgccgctcgcgacagatgaaagctggctggatcaggaactcgacgaacgccagcgggtgattgccgc- actggagaaaaccggctgggt gcaggccaaagcggcccgactgctgggcatgacaccgcgccagattgcctaccgtatccagattatggacatc- aacatgcaccgtatctga 75 aaaactaccgccgcaattaatgaacccaacgctactgttgccgggccatgctcttccccggcgcgctgccc- ggaaaggatatagattgcccagcacg cgccagcaccaagcgcgaacgccgcgccagtgagatcaacatgtgaaacattttcgcccagcggcagcagata- caagaggccaagtaccgccag gatcacccagatgaaatccaccgggcggcgtgaggcaaaaagcgccaccgccagcgggccggtaaattccagc- gccaccgcaacgccgagcgg tatcgtctggatcgataaatagaacatatagttcatggcgccgagcgacaggccataaaacagcagtggcagg- cgttgttcacgggtaaaatgtaaac gccagggcttgaacactacgaccaaaataagggtgccaagtgcgagacgcagcgcggtgacgccgggtgcgcc- aacaatcggaaacagtgatttc gccagcgacgcgcctccctgaatggacatcatcgcgacaaacaatattaataccggcaaccacaccggcaatt- tacgagactgcgcaggcatccttt ctcccgtcaatttctgtcaaataaagtaaaagaggcagtctacttgaattacccccggctggttgagcgttgt- tgaaaaaaagtaactgaaaaatccgta gaatagcgccactctgatggttaattaacctattcaattaagaattatctggatgaatgtgccattaaatgcg- cagcataatggtgcgttgtgcgggaaaa ctgcttttttttgaaagggttggtcagtagcggaaactttctgttacatcaaatggcgctttagaccccaatt- cccgcaaagagtttcttaactaattttgatat atttaaacgcgtaggacgtaggatttacttgaagcacatttgaggtggattatgaaaaaaattgcatgtcttt- cagcactggccgcacttctggcggtttct gcaggttccgcagtagcagcaacttcaaccgtaactggcggctacgctcagagcgacgctcagggtattgcta- acaaaactaacggtttcaacctgaa atatcgctacgagcaggacaacaacccgctgggtgttatcggttcctttacttacactgaaaaagatcgcacc- gaaagcagcgtttataacaaagcgca gtactacggcatcaccgcaggcccggcttaccgcatcaacgactgggcgagcatctacggtgttgtgggtgta- ggttacggtaaattccagcagattg tagacaccgctaaagtgtctgacaccagcgactacg 76 aacacacgctcctgttganaaagagatcccgccgggaaatgcggtgancgtgtctgatattgcgaagagtg- tgccagttttggcgcgggcaaaacct gcaccagtttggtattaatgcaccagtctggcgctttttttcgcgagtttctcctcgctaatgcccgccaggc- gcggctttggcgctgatagcgcgctg aataccgatctggatcaaggttttgtcgggttatcagccaaaaggtgcactctttgcatggttatacgtgcct- gacatgttgtccgggcgacaaacggcct ggtggcacaaattgtcagaactacgacacgactaactgaccgcaggagtgtgcgatgaccctgaatatgatga- tggatgccggcgcgcccgaggca atcgccggtgcgctttcgcgacaccatcctgggctgttttttaccatcgttgaagaagcgcccgtcgccattt- cgctgactgatgccgacgcacgcattg tctatgccaacccggctttctgccgccagaccggctatgaactagaagcgttgttgcagcaaaatccccgcct- gcttgcaagtcgccaaaccccacgg gaaatctatcaggatatgtggcacaccattgttacaacgccgaccgtggcgcgggcaattgattaaccgccac- cgcgacggcagcctgtatctggtcga gatcgatatcaccccggtgattaacccgtttggcgaactggaacactacctggcaatgcagcgcgatatcagc- gccagttatgcgctggagcagggt tgcgcaatcacatgacgctgaccgaagcggtgctgaataacauccggcggcggtggngtagtggatgaacgcg- atcatgtggttatggataaccttg cctacaaaacgttctgtgccgactgcggcggaaaagagctcctgaggaactcaatttttcagcccgaaaaggg- agctggcaaacggccaggtctt accggtggtgctgcgcggtgaggtgcgctggttgtcggtgacctgctgggcgctgccgggcgtcagcgaagaa- gccagttcgctactttattgataac aggctgacgcgcacgctggtggtgatcaccgacgacacccaacaacgccagcagcaggaacagggccgacttg- accgccttaaacagcagatga ccaacggcaaactactggcagcgatccgcgaagcgcttgacgccgcgctgatccagataactgccccatcaat- atgctggcggcggcgcgacgttt aaacggcagtgataacaacaatgtggcgctcgacgccgcgtggcgcgaaggtgaagtggcgatggcgcggctg- aaacgttgccgcccgtcgctg gaactggaaagtgcggccgtctggccgctgcaaccctttttgacgatctgcgcgcgctttatcacacccgcta- cgagcaggggaaaaatttgcaggt cacgctggattcccatcatctggtgggatttggtcagcgtacgcaactgttagcctgcctgagtctgtggctc- gatcgcacgctggatattgccgccgg gctgggtgatttcaccgcgcaaacgcagatttacgcccgcgaagaagagggctggctctctttgtatatcact- gacaatgtgccgctgatcccgctgcg ccacacccactcgccggatgcgcttaacgctccgggaaaaggcatgcacctgcgcctgatccagacgctggtg- gcacaccaccacggcgcaatag
aactcacttcacaccccgaagggggaagttgcctgaccctacgattcccgctattcattcactgaccggaggt- tcaaatgacccagcgaaccgagt cgggtaataccgtctggcgcttcgatttgtcccagcagttcactgcgatgcagcgcataagcgtggtactcag- ccgggcgaccgaggtcgatcagac gctccagcaagtgctgtgcgtattgcacaatgacgcctttttgcagcacggcatgatctgtctgtacgacagc- cagcaggcgattttgaatattgaagcg ttgcaggaagccgatcagcagttaatccccggcagctcgcaaatccgctatcgtccgggcgaagggctggtcg- ggacggtgctttcgcagggccaa tcattagtgctggcgcgcgttgctgacgatcaggctttcttgaccggctcgggttgtatgattacaacctgcc- gtttatcgccgtgccgctgatagggcc agatgcgcagactttcggtgtgctgacggcacaacccatggcgcgttacgaagagcgattacccgcctgcacc- cgctttctggaaacggtcgctaac ctggtcgcgcaaaccgtgcgtttgatggcaccaccggcagtgcgcccttccccgcgcgccgccataacacagg- ccgccagcccgaaatcctgcac ggcctcacgcgcatttggttttgaaaatatggtcggtaacagtccggcgatgcgccagaccatggagattatc- cgtcaggtttcgcgctgggacaccac cgttctggtacgcggcgagagtggcaccggcaaggagctgattgccaacgccatccaccaccattcgccgcgt- gccggtgcgccatttgtgaaattc aactgtgcggcgctgccggacacactgctggaaagcgaattgttcggtcacgagaaaggggcatttaccggcg- cggtacgccagcgtaaaggccg ttttgagctggccgatggcggcacgctgtttcttgacgagatcggcgagagtagcgcctcgtttcaggctaag- ctgctgcgcattttgcaggaaggcga aatggaacgcgtcggcggcgacgagacattgcaagtgaatgtgcgcattattgccgcgacgaaccgcaatctt- gaagatgaagtccggctggcgca ctttcgcgaagatctctattatcgcctgaatgtgatgcccatcgccctgccgccactacgcgaacgccaggag- gacattgccgagctggcgcactttct ggtgcgtaaaatcgcccataaccagagccgtacgctgcgcattagcgagggcgctatccgcctgctgatgagc- tacaactggcccggtaatgtgcgc gaactggaaaactgccttgagcgctcagcggtgatgtcggagaacggtctgatcgatcgggatgtgattttgt- ttaatcatcgcgaccagccagccaaa ccgccagttatcagcgtctcgcatgatgataactggctcgataacaaccttgacgagcgccagggctgattgc- ggcgctggaaaaaggggatgg gtacaagccaaagccgcgcgcttgctggggatgacgccgcgccaggtcgcctatcgtattcagacgatggata- taaccctgccaaggctataa 77 MTLNMMMDAGAPEAIAGALSRHHPGLFFTIVEEAPVAISLTDADARIVYANPAFcRQTGYELE ALLQQNPRLLASRQTPREIYQDMWHTLLQRRPWRGQLINRHRDGSLYLVEIDITPVINPFGELE HYLAMQRDISASYALEQRLRNHMTLTEAVLNNIPAAVVVVDERDHVVMDNLAYKTFcADcG GKELLSELNFSARKAELANGQVLPVVLRGEVRWLSVTcWALPGVSEEASRYFIDNRLTRTLVV ITDDTQDRQQQEQGRLDRLKQQMTNGKLLAAIREALDAALIQLNcPINMLAAARRLNGSDNN NVALDAAWREGEEAMARLKRcRPSLELESAAVWPLQPFFDDLRALYHTRYEQGKNLQVTLD SHHLVGFGQRTQLLAcLSLWLDRTLDIAAGLGDFTAQTQIYAREEEGWLSLYITDNVPLIPLRH THSPDALNAPGKGMELRLIQTLVAHHHGAIELTSHPEGGScLTLRFPLFHSLTGGSK 78 MTQRTESGNTVWRFDLSQQFTAMQRISVVLSRATEVDQTLQQVLcVLHNDAFLQHGMIcLYD SQQAILNIEALQEADQQLIPGSSQIRYRPGEGLVGTVLSQGQSLVLARVADDQRFLDRLGLYDY NLPFIAVPLIGPDAQTFGVLTAQPMARYEERLPAcTRFLETVANLVAQTVRLMAPPAVRPSPRA AITQAASPKScTASRAFGFENMVGNSPAMRQTMEIIRQVSRWDTTVLVRGESGTGKELIANAIH HHSPRAGAPFVKFNcAALPDTLLESELFGHEKGAFTGAVRQRKGRFELADGGTLFLDEIGESSA SFQAKLLRILQEGEMERVGGDETLQVNVRIIAATNRNLEDEVRLGHFREDLYYRLNVMPIALPP LRERQEDIAELAHFLVRKIAHNQSRTLRISEGAIRLLMSYNWPGNVRELENcLERSAVMSENGL IDRDVILFNHRDQPAKPPVISVSHDDNWLDNNLDERQRLIAALEKAGWVQAKAARLLGMTPR QVAYRIQTMDITLPRL 79 atgccgcaccacgcagcattgtcgcagcactggcaaacggtattttctcgtctgccggaatcgctcaccgc- gcagccattgagcgcgcaggcgcagt cagtgctcacttttagtgattttgttcaggacagcatcatcgcgcatcctgagtggctggcagagcttgaaag- cgcgccgccgcctgcgaacgaatggc aacactatgcgcaatggctgcaagcggcgctggatggcgtcaccgatgaagcctcgctgatgcgcgcgctgcg- gctgtttcgccgtcgcatcatggt gcgcatcgcctggagccaggcgttacagttggtggcggaagaagatatcctgcaacagcttagcgtgctggcg- gaaaccctgatcgtcgccgcgcg cgactggctttatgaggcctgctgccgtgagtggggaacgccgagcaatccacaaggcgtggcgcagccgatg- ctggtactcggcatgggcaaact gggtggcggcgaactcaatttctcatccgatatcgatttgattttcgcctggccggaaaatggcgcaacgcgc- ggtggacgccgtgagctggataacg cgcaatttttcactcgccttggtcaacggctgattaaagtcctcgaccagccaacgcaggatggctttgtcta- ccgcgtcgatatgcgcttgcgcccgttt ggcgacagcggcccgctggtgctgagctttgccgcgctggaagattactaccaggagcaggggcgcgattggg- aacgctacgcgatggtgaaagc gcgcattatgggcgataacgacggcgaccatgcgcgggagttgcgcgcaatgctgcgcccgtttgttttccgc- cgtatatcgacttcagcgtgaaca gtccctgcgtaacatgaaaggcatgattgcccgcgaaggtgcgtcgccgtggcctgaaggacaacattaagct- cggcgcgggcgggatccgcgaaat agaatttatcgtccaggttttccagagattcgcggcggtcgcgagcctgcactgcaatcgcgacactgttgcc- gacgcttgctgccatagatcaactg catctgctgccggatggcgacgcaacccggctgcgcgaggcgtatttgtggctgcgacggctggagaacctgc- tgcaaagcatcaatgacgaacag acacagacgctgccgggcgatgaactgaatcgcgcgcgcctcgcctggggaatgggcaaagatagctgggaag- cgctctgcgaaacgctggaag cgcatatgtcggcggtgcgtcagatatttaacgatctgattggcgatgatgaaacggattcgccggaagatgc- gctttctgagagctggcgcgaattgt ggcaggatgcgttgcaggaggaggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgt- ggtggcgctgattgccgattttcg caaagagttggataaacgcaccattggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgaca- gcgatgtatgctcgcgcgacgat gcgccagtaccgctgtcacgcctgacgccgctgacaccggaattattacccgcaccacttaccttgagctgct- aagtgaatttcccggcgcactgaaa cacctcatttccctgtgtgccgcgtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatg- aattgctcgacccgaatacgctctatc aaccgacggcgatgaatgcctatcgcgatgagagcgccaatacctgctgcgcgtgccggaagatgatgaagag- caacagcttgaggcgctgcgg cagtttaagcaggcgcagttgctgcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtga- gcgatcacttaacctggctggcgg aagcgattattgatgcggtggtgcagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatct- gcacgatcgcgaagggcgcggtt ttgcggtggtcggttatggcaagctgggcggctgggagctgggttacagctccgatctggatctggtattcct- gcacgactgcccgatggatgtgatga ccgatggcgagcgtgaaatcgatggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttag- cacgcgcacgtcgtccggcatccttt atgaagttgatgcgcgtctgcgtccatctggcgctgcggggatgctggtcactactacggaatcgttcgccga- ttaccagcaaaacgaagcctggacg tgggaacatcaggcgctggcccgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgcca- ttcgccgcgatattctgatcacgc ctcgcgacggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaa- gcataaagaccgcttcgatctga aagccgatgaaggcgftatcaccgacatcgagtttatcgcccaatataggtgaggctttgcccatgacaagcc- gaaactgacgcgctggtcggat aatgtgcgcattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctgg- cgtacaccacattgcgtgatga gctgcaccacctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgctt- attaaaaccagctgggacaagtggc tggtggaaccgtgcgccccggcgtaa 80 atgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctgagagctggcgcga- attgtggcaggatgcgttgcaggagga ggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgctgattgccgatttt- cgcaaagagttggataaacgcacc attggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgtatgctcgcgcgacg- atgcgccagtaccgctgtcacgcc tgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgctaagtgaatttcccggcgcact- gaaacacctcatttccctgtgtgccgc gtcgccgaggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcgacccgaatacgctct- atcaaccgacggcgatgaatgccta tcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagcttgaggcgctgcgg- cagtttaagcaggcgcagttgct gcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcacttaacctggctggcg- gaagcgattattgatgcggtggtg cagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgcacgatcgcgaagggcgcggtt- ttgcggtggtcggttatggcaag ctgggcggctgggagctgggttacagctccgatctggatctggattcctgcacgactgcccgatggatgtgat- gaccgatggcgagcgtgaaatcg atggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgcacgtcgtccggcat- cctttatgaagttgatgcgcgtctgcgt ccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagcaaaacgaagcctgga- cgtgggaacatcaggcgctggcc cgtgcgcgcgtggtgtacggcgaccgcaactgaccgccgaatttgacgccattcgccgcgatattctgatgac- gcctcgcgacggcgcaacgctgc aaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaagaccgcttcgatct- ganagccgatgaaggcggtatca ccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaagccgaaactgacgcgctggtc- ggataatgtgcgcattctcgaagggc tggcgcaaaacggcatcatggaggagcaggaagcgcaggcattcacgctggcgtacaccacattgcgtgatga- gctgcaccacctggcgctgcaa gagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttattaaaaccagctgggacaagt- gcctggtggaaccgtgcgccccggc gtaa 81 MPHHAGLSQHWQTVFSRLPESLTAQPLSAQAQSVLTFSDFVQDSIIAHPEWLAELESAPPPANE WQHYAQWLQAALDGVTDEASLMRALRLFRRRIMVRIAWSQALQLVAEEDILQQLSVLAETLI VAARDWLYEAccREWGTPSNPQGVAQPMLVLGMGKLGGGELNFSSDIDLIFAWPENGATRG GTRELDNAQFFTRLGQRLIKVLDQPTQDGFVYRVDMRLRPFGDSGPLVLSFAALEDYYQEQGR DWERYAMVKARIMGDNDGDHARELRAMLRPFVFRRYIDFSVIQSLRNMKGMIAREVRRRGL KDNIKLGAGGIREIEFIVQVFQLIRGGREPALQSRSLLPTLAAIDQLHLLPDGDATRLREAYLWL RRLENLLQSINDEQTQTLPGDELNRARLAWGMGKDSWEALcETLEAHMSAVRQIFNDLIGDD ETDSPEDALSESWRELWQDALQEEDSTPVLAHLSEDDRRRVVALIADFRKELDKRTIGPRGRQ VLDHLMPHLLSDVcSRDDAPVPLSRLTPLLTGIITRTTYLELLSEFPGALKHLISLcAASPMVAS QLARYPILLDELLDPNTLYQPTAMNAYRDELRQYLLRVPEDDEEQQLEALRQFKQAQLLRVA AADIAGTLPVMKVSDHLTWLAEAIIDAVVQQAWGQMVARYGQPTHLHDREGRGFAVVGYG KLGGWELGYSSDLDLVFLHDcPMDVMTDGEREIDGRQFYLRLAQRVMHLFSTRTSSGILYEV DARLRPSGAAGMLVTTTESFADYQQNEAWTWEHQALARARVVYGDPQLTAEFDAIRRDILM TPRDGATLQTDVREMREKMRAHLGNKHKDRFDLKADEGGITTDIEFIAQYLVLRFAHDKPKLT RWSDNVRILEOLAQNGIMEEQEAQALTLAYTTLRDELHHLALQELPGHVALScFVAERALIKT SWDKWLVEPcAPA 82 MFNDLIGDDETDSPEDALSESWRELWQDALQEEDSTPVLAHLSEDDRRRVVALIADFRKELDK RTIGPRGRQVLDHLMPHLLSDVcSRDDAPVPLSRLTPLLTGIITRTTYLELLSEFPGALKHLISLc AASPMVASQLARYPILLDELLDPNTLYQPTAMNAYRDELRQYLLRVPEDDEEQQLEALRQFKQ AQLLRVAAADIAGTLPVMKVSDHLTWLAEAIIDAVVQQAWGQMVARYGQPTHLHDREGRGF AVVGYGKLGGWELGYSSDLDLVFLHDcPMDVMTDGEREIDGRQFYLRLAQRVMHLFSTRTSS GILYEVDARLRPSGAAGMLVTTTESFADYQQNEAWTWEHQALARARVVYGDYQLTAEFDAIR RDILMTPRDGATLQTDVREMEREKMRAHLGNKHKDRFDLKADEGGITDIEFIAQYLVLRFAHD KPKLTRWSDNVRILEGLAQNGIMEEQEAQALTLAYTTLRDELHHLALQELPGHVALScFVAER ALIKTSWDKWLVEPcAPA 83 EEQQLEALRQFKQAQLLRVAAADIAGTLPVMKVSDHLTWLAEAIIDAVVQQAWGQMVARYG QPTHLHDREGRGFAVVGYGKLGGWELGYSSDLDLVFLHDcPMDVMTDGEREIDGRQFYLRL AQRVMHLFSTRTSSGILYEVDARLRPSGAAGMLVTTTESFADYQQNEAWTWEHQALARARV VYGDPQLTAEFDAIRRDILMTPRDGATLQTDVREMREKMRAHLGNKHKDRFDLKADEGGITD IEFIAQYLVLRFAHDKPKLTRWSDNVRILEGLAQNGIMEEQEAQALTLAYTTLRDELHHLALQE LPGHVALScFVAERALIKTSWDKWLVEPcAPA 84 ccgagcgtcggggtgcctaatatcagcaccggatacgagagaaaagtgtctacatcggttcggttgatatt- gaccggcgcatccgccagcccgccca gtttctggtggatctgtttggcgattttgcgggtcttgccggtgtcggtgccgaaaaaaataccaatatttgc- cataacacacgctcctgttgaaaaagag atcccgccgggaaatgcggtgaacgtgtctgatattgcgaagaggtgccagttttatcgcgggcaaaacctgc- accagtttggttattaatgcacca gtctggcgctttttttcgccgagtttctcctcgctaatgcccgccaggcgcggctttggcgctgatagcgcgc- tgaataccgatctggatcaaggttttgtc gggttatcagccaaaaggtgcactctttgcatggttatacgtgcctgacatgttgtccgggcgacaaacggcc- tggtggcacaaattattgtcagaactacg acacgactaactgaccgcaggagtgtgcgatgaccctgaatatgatgatggatgccggcggacatcatcgcga- caaacaatattaataccggcaacc acaccggcaatttacgagactgcgcaggcatcctttctcccgtcaatttctgtcaaataaagtaaaagaggca- gtctacttgaattacccccggctggttg agcgtttgttgaaaaaaagtaactgaaaaatccgtagaatagcgccactctgatggttaattaacctattcaa- ttaagaattatctggatgaatgtgccatta aatgcgcagcataatggtgcgttgtgcgggaaaactgcttttttttgaaagggttggtcagtagcggaaacaa- ctcacttcacaccccgaagggggaa gttgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcg- ggtaataccgtctggcgcttcgatttgt cccagcagttcactgcgatgcagcgcataagcgtggtactcagccgggcgaccgaggtcgatcagacgctcca- gcaagtgctgtgcgtattgcaca atgacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgaatattgaagcgtt- gcaggaagccgatcagcagttaatccc cggcagctcgcaaatccgctatcgtccgggcgaagggctggtcgggacggtgctttcgcagggccaatcatta- gtgctggcgcgcgttgctgacgat cagcgctttcttgaccggctcgggttgtatgattacaacctgccgtttatc 85 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcggcagc- gggaagtagatgctactttgccggc gagcggcggacgggtgagtaatgtctgggtatctgcctgatggagggggataactactggaaacggtagctaa- taccgcatgacctcgaaagagc aaagtgggggatcttcggacctcacgccatcggatgtgcccagatgggattagctagtaggtgaggtaatggc- tcacctaggcgacgatccctagct ggtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaa- tattgcacaatgggcgcaagc ctgatgcagccatgccgcgtgtgtgaagaaggccttagggttgtaaagcactttcagcgaggaggaaggcatc- anacttaatacgtcggtgattgac gttactcgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatc- ggaattactgggcgtaaagcgca cgcaggcggtttgttaagtcagatgtgaaatccccgcgcttaacgtgggaactgcatttgaaactggcaagct- agagtcttgagaggggggtagaatt ccaggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagact- gacgctcaggtgcgaaagcgt ggggagcaaacaggattagataccctggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttga- ggcgtggcttccggagctaacgcgt taagtcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggt- ggagcatgtggtttaattcgatgc aacgcgaagaaccttacctactcttgacatccacgcaattcgccagagatggcttagtgccttcgggaaccgt- ganacaggtgctgcatggctgtcgt cagctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcnngtna- tgnngggaactcaaaggagactgcc ggtgataaaccggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacgtgct- acaatggcatatacaaacagaag cgaactcgcgagg 86 atggcaatgcgtcaatgcgcaatctacgggaaagggggtattgggaaatccaccactacccaaaaccttgt- agcggctctggccgaaatgaataaga aggtcatgatcgtcggctgtgaccctaaggctgattcaacccgcctcattctgcatgcgaaagcacagaacac- catcatggaaatggccgctgaagtg ggctccgtggaagatctggagctggaagatgtgatgcaaatcggctatggcggcgtgcgctgtgcggaatcag- gcggccctgagcctggtgtgggt tgtgccggacgcggggtgatcaccgccatcaacttcctcgaagaagaaggcgcgtatgtgccggatctggatt- ttgtgttttacgacgtattgggcgat gtggtctgtggcggtttcgcgatgccaattcgcgaaaacaaagcgcaggaaatctacatcgtgtgctccggtg- aaatgatggcgatgtatgccgccaa caacatttccaaaggcatcgtgaaatacgcgaaatcgggcaaagttcgcctggccgggctgatctgtaactcc- cgccagacggatcgcgaagatgaa ctgatcatcgcgctggctgaaaaacttacacgcaatttgatccacttcgtgccgcgtgacaacattgtgcaac-
gcgctgaaatccgccgcatgacggt catcgaatacgacccgacttgtgcgcaggcagatcagtatcgtgcactggcgaacaaaatcgtcaacaacacc- aaaatggtggtgccgacaccggtc accatggatgagctggaagccctgttaatggaatttggcattatggaagaagaagacctcgccatcgtcggtc- gtaccgccgccgaagaggcgtga 87 atgaaggcaaaagagattctggcgctgattgatgagccagcctgtgagcataaccacaagcagaagtcggg- ttgcagcctgccgaaaccgggcgc gacggcaggcggttgtgcgtttgatggcgcgcagattgcgctgctgccggtcgcggacgtcgcgcatctggtg- cacggcccgattggctgtaccgg cagttcatgggacaaccgtggcagccgcagttccgggccttccatcaaccgcatgggcttcaccaccgacatg- agcgagcaggatgtgattatgggg cgcggcgagcgacgcttatttcacgccgtgcagcacatcgtcagccattaccatccggtggcggtctttattt- acaacacctgcgtacccgcgatggaa ggggatgacgttgaagccgtgtgtcgcgccgcatcggccgctgccggtgtgccggttatttcagtcgatgccg- ccggtttctacggcagcaaaaatct cggtaaccgcattgccggggacgtgatggtcaaaaaggtgatcggccagcgcgaacccgcgccgtggccggaa- aactcaccgatccccgccgga caccgccacagcatcagcctgattggcgaattcaatattgccggcgagttctggcacgttctgccgctgctcg- atgagctcgggatccgcgtgctgtgc agcctttccggggattcccgttttgctgaaatccagactatgaaccgtggcgaagccaacatgctggtgtgct- cgcgggcgctgatcaacgtcgcccg aaaaatggaagagcgttaccagatcccatggtttgaaggcagtttttatggcctgcgttccatggctgattcc- ctgcgcacgatcgccgtgctgctcaaa gacccggatttacaggcgcgcacagaacgtctgattgagcgcgaggaggcggcgacacatcttgcgcttgcgc- cttaccgtgcgcggctcagcgg gcgcaaggcgctgctgtataccggtggcgtgaaatcctggtcggtggtctcggcgttacaggatttaggcatc- acggtggtggcgaccggcacccga aaatcaaccgaagaagacaagcagcgtattcgcgaactgatgggtgaagacgtgctgatgctcgacgaaggca- atgccagaaccttgctcgacacc ctctatcgtttcggcggcgacatcatgatcgccgggggccgcaacatgtataccgcgtacaaagccgcctgcc- gttcctggatatcaatcaggagcg cgagcatgcgtttgccggatatcacgggctggtaaatctggccgaacagttgtgtatcaccctggaaagcccg- gtctgggcgcaggtcaaccgtctg gcgccgtggcgctaa 88 atgaccagtgaaacacgcgaacgtaacgaggcattgatccaggaagtgctggagatcttccccgagaaggc- gcttaaagatcgtaagaaacacatg atgaccaccgacccggcgatggaatctgtcggcaagtgtattgtctcaaaccgcaaatcacagccgggcgtga- tgaccgtgcgaggctgcgcttacg ccggttccaaaggcgtggtctttggcccgatcaaagacatggcgcatatctcccacggcccggttggttgcgg- ccagtattcccgtgccggacgccgt aactattacaccggctggagcggcgtgaacagctttggcaccctcaacttcaccagtgattttcaggaacggg- acatcgtatttggcggcgataaaaag ctcgacaaattgatcgatgaactggagatgttgttcccgctgagcaaaggcatttcggtgcagtcggaatgtc- cggtcggtctgatcggcgatgacattt ctgccgtcgccaaagccagcagcgccaaaatcggtaagccggtcgtgccggtacgctgcgaggggttccgcgg- tgtgtcgcaatcgctcggccatc acattgctaacgatgtcatccgcgactgggtgctggataaccgcgaaggcaatgaatttgaaaccacgcctta- cgacgtggcgattatcggcgactac aacatcggcggtgacgcctggccctcacgtattctgctcgaagaaatggggctgcgcgtggtggcgcagtggt- ccggcgacggcacgctggtgga gatggaaaacaccccgaaagtcgcgctcaatctggtgcactgctaccgctcgatgaactacatctcccgtcat- atggaagaaaaacacggcattccgt ggatggaatacaacttctttggcccgaccaaaattgcggaatctctgcgcgaaatcgcggcggttttgacgat- accatccggaaaaacgccgaagc ggtgattgaaaaatatcaggcgcaaacgcaggcggtgatcgacaaataccgtccgcgtctggaaggcaaaaag- gtgctgttgtatctcggcggtttac gtccgcgccacatcatcggggcgtatgaagatctgggaatggaaatcatcggtaccggctatgaattcggtca- taacgatgattacgaccgcaccttac cgatgctcaaagaaggcacgttgctgttcgatgacctgagcagttatgagctggaagcgttcgttaaagcgct- gaaaccggatcttgtcgggtcaggta tcaaagaaaaatacattttccagaaaatgggcgtgccgttccgccagatgcactcctgggattattccggccc- ttatcacggctacgacggtttcggcat ttttgcccgtgacatggacatgacgctgaacaatccgggctggagtcagctgaccgccccctggttgaaaagc- ctga 89 atggctcaaattctgcgtaatgccaagccgcttgccaccacgcctgtcaaaagcgggcaaccgctcgcggc- gatcctggccagtcaggggctggaa aattgcatcccgctggttcacggcggcaaggttgtagcgcgttcgccaaagttttcttcatccagcattttca- cgatccgatcccgttgcagtccacggc gatggaatcgaccacgactatcatgggctcggatggcaacgtcagtactgcgttgaccacgttgtgtcagcgc- agtaatccaaaagccattgtgattttg agcaccggactgtcagaagcgcagggcagtgatttgtcgatggcgctgcgtgagtttcgcgacaaagaaccgc- gctttaatgccatcgctattctgac cgttaacacgccggatttttacggctcgctggaaaacggctacagcgcgctgatggaaagcgtgatcactcag- tgggtgccggaaaagccgccgac cggcatgcgtaacaagcgcgtgaacctgctggtgagccatctgctgacgccgggggatctggaattactgcgc- agctatgtcgaagcctttggcctg caaccggtgatcctgccggatttatcacagtcgctggacggacatctggcgaatggcgatttcaatccggtca- cgcagggcggcacgtcgcaacgcc agattgaacaaatggggcagagcctgaccaccattaccattggcagttcgctcaactgcgccgccagtctgat- ggcgatgcgcagccgtggcatggc gctgaacctgccgcacctgatgacgctggaaaacatggacagtctgatccgccatctgcatcaggtgtcaggc- cgcgaggtaccggcatggattgag cgccagcgcgggcaactgaggacgccatgatcgactgccatacctggctgcagtcacagcgtattgcgctggc- ggcagaagcggatttgctggtg gcgtggtgcgatttcgctcagagccagggaatgcgcgtcgggccggtgattgcgccggttaatcagcagtcac- tggccgggctgccggtcgaacag gtggtgatcggcgatctggaagatttacaaacccggctcgacagctacccggtttcactgctggtggcgaact- cccacgctgcaccactggcggaaa aaaacggtatcgcgctggtacgtgccggtttcccgctttacgaccgtctcggggaatttcgccgcgtgcggca- gggctatgcgggtattcgcgacacc ttgttcgaactcgcgaacctgatgaggcgcgccatcacatgctgacggcgtatcactcaccgcttaggcaggt- gttcggcctgagcccggtaccgg aggccagtcatgaggcggctaa 90 atgagtcaagatcttggcaccccaaaatcctgtttcccgctcttccagcaggatgaataccagaatatgtt- tacccacaaacgcgcgctggaagaagca cacggcgaggcgaaagtgcgggaagtgtttgaatggaccaccacgcaggaatatcaggatctgaacttctcgc- gtgaagcgctgaccgtcgacccg gcgaaagcctgccagccgttaggcgccgtactttgcgcgctgggttttaccaacacgttgccgtatgtccatg- gttcacaaggctgtgtggcctatttcc gtacctattttaatcgtcatttcaaagagccggtggcctgtgtttccgactcaatgaccgaagatgccgccgt- ttttggcggaaataacaacatgaatgtc ggtctggaaaacgccagcgcgctgtacaagccggaaattattgcggtctccaccacctgtatggcggaagtga- tcggtgatgacctgcaggcttttatc gccaacgccaaaaaagacggatttgtggatgcggtatgccaatcccgtatgcccatacaccgagttttctggg- cagtcatgtcaccggctgggacaa catgtttgaaggcttcgcccgtacctttaccaccgacgccacgcgggaatatcagccgggcaaacttgccaaa- ctgaacgtggtgaccggttttgaaa cttatctcggcaactaccgggttattcaccgcatgatgagccagatgggggtcgaatgcagcgtcttgtccga- tccgtctgaagtgctcgacaccccgg ctgacggccaataccgcatgtatgccgacggcaccacgcaaaccgaaatgcgtgatgaccggatgccatcgac- accttgctgctgcaaccgtggc aattacagaaaaccaaaaaggtggtgcagggcgactggaatcagccgggcaccgaagtcagtgtaccgattgg- cctggcggcgaccgatgccttg ctgatgacggtaaggaactgaccggcaaaccgatagctgacgcgctggcgactgaacgtggccgtctggtgga- catgatgctcgattctcacacct ggctgcacggcaagcgtttcggtctctacggtgacccggattttgtgatgggcatgaccgcattcctgctgga- actgggctgtgaaccgaccaccattc tcagccataacggcaacaaacgctggcagaaagccatgaagaaaatgctggctgattcgccttacggacagga- cagcgaagtgtatgtgaactgcg atctgtggcatttccgctcgctgatgtttacccgtaaaccggactttatgatcggcaactatacggaaaattc- attcagcgtgacacgctggccaaaggc gaacagttcgaagtgccgctgatccgtatcggattcccgatttttgaccggcaccatttgcaccgtcagacca- cctggggatacgagggcgcgatgag catcctgacgcaactggtgaatgcggtgctcgaacagctggatcgcgaaaccatgaagctcggcaaaaccgac- tacaacttcgatctgatccgctaa 91 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagcgcttgtcgctgcaacatcct- tcactgttttataccgtggttgaacaatca tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggccagccagcagacgccgaaacatatctatgacgaaatgtggcgcactttgttgca- gggctaatcctggaacggccaact gatcaaccggcgtaataaccgttcgctttatctggcggatgtcactatcacgcctgttttaggcgcggacggg- caggtggagcattacctcggcatgca caaagatatcagcgagaaatacgcgctgcagcagggttgcgcaaccacatcaccttgttcacggaggtgctga- acatttattcccgccgccgtggtg gtggtggatgagaggacaatgtggtgatggacaatctggcctacaaaaccctgtgcgctgactgcggcgcaaa- agagctgttggccgaaatgggc tatccgcaactcaaagagatgctcaacagtggcgaaccggtgccggtttccatgcgcggcaacgtacgctggt- tttctttcggtcagtggtcattgcag ggcgttaatgaagaggccagccgcttttttaccggcattaccgcgccgggaaaactgattgttctcaccgact- gaccgatcagcatcaccggcagca gcagggttatcttgaccggctcaagcaaaaacttaccaacggcaaattgctggcagccatccgcgagtcgctt- gatgccgcgctgattcagctcaacg ggccattttaatatgctggcggctgcgcgtcgtcttaacggcgaagaaggcaacaacatggcgctggaattcg- cctggcgcgaaggcgagcaggcg gtgagtcgcttacaggcctgccgtccgtcgctggattttgagccgcaggcagaatggccggtcagtgaattct- tcgacgatctgagcgcgctgtacga cagccattttctcagtgacggtgaattgcgtacgtggtcatgccatctgatctgcacgctgtcgggcaacgaa- cgcaaatccttaccgcgctgagcttat ggattgatcacacgctgtcacaggcgcaggccatgccgtctctgaagctctcggtgaacattgttgcgaagca- ggatgcgagctggttgtgttttgaca ttaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaagcggcgttttcccgtccgggcaatgg- catggagctgcgccttatccagac gctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggcaccttgagacgttacgcc- tgccggtacagcaggttatcaccgga ggcttaaaatga 92 atgacccagttacctaccgcgggcccggttatccggcgctttgatatgtntgcccagtttacggcgcttta- tcgcatcagcgtggcgctgagtcaggaa agcaataccgcgcgcgcactggcggcgatcctcgaagtgcttcacgatcatgcatttatgcaatacggcatgg- tgtgtctgttcgataaagaacgcaat gcactgtttgtggaatccctgcatggcatcgacggcgaaaggaaaaaagagacccgccatgtccgttaccgca- tgggggaaggcgtgatcggcgc ggtgatgagccagcgtcaggcgctggtgtaccgcgcatttcagacgatcagcgttttctcgaccgcctgaata- tttacgattacagcctgccgatgatt ggtgtgccgatccccggtgcggataatcagcctgcgggtggctggtggcacagccgatggcgttgcacgaaga- ccggctggctgccagtacgcg gtttttagaaatggcgccaatctcatcagccagccactgcgttctgccacgcccccggaatcattgcctgctc- aaacgccggtccggtgcagtgttccg cgccagtttggttttgagcagatggtcgggaaagtcaggcgatgcgccagacgatggacattttacggcaggt- ttccaaatgggataccacggttctg gtgcgtggtgaaagcggcaccggcaaggaacttatcgccaatgccattcattacaactcaccccgtgcggccg- cgccatttgtgaaattcaactgcgc cgcgctgccggataacctgctggaaagcgaactgttcggtcatgaaaaaggggccttcaccggcgctatacgc- acccgaaaaggccgctttgaact ggcggacgggggcacgttattcctcgatgaaatcggcgaatcgagcgcgtcgtttcaggccaaattgctgcgc- attttgcaggaaggtgaaatggaa cgggtcggcggcgataccacgctgaaagttgatgtgcgcattattgctgccaccaaccgtaatcttgaggagg- aagtgcgtgccgggaattttcgcga agacctgtattatcgcctcaacgtgatgccggtttcgctgcctgcactgcgtgaaaggctggatgatatcgcc- gatctggcgccgtttctggtcaaaaag attgcgctgcgtcaggggcgggaactgcgcatcagtgatggtgcggtgcgtctgctgatgacctacagctggc- caggcaacgtgcgtgaactggaa aactgcctcgaacgggcgtcggtaatgaccgatgaagggctgatcgaccgcgacgtgatcctgttcaatcacc- atgagtccccggcgctgtccgtca aacccggtctgccgctcgcgacagatgaaagctggctggatcaggaactcgacgaacgccagcgggtgattgc- tgcactggagaaaaccggctgg gtgcaggccaaagcggcccgactgagggcatgacaccgcgccagattgcctaccgtatccagattatggacat- caacatgcaccgtatctga 93 atgttgccactttcttctgttttgcaaagccacgcgcagagtttgcctgaacgctggcatgaacatcctga- aaacctgcccctccccgatgatgaacagct ggctgtgctgagcagcagtgaattcatgacggacagtttgctggcttttccgcagtggtggcatgaaattgtc- caaaatccccctcaggcgcaggagtg gcaactttaccgtcagtggctggatgaatcactgacgcaggtgactgacgaagccgggttaatgaaagctttg- cgtctgttccgccgccgtattctgac ccgcattgcgtggtcacagtccgcgcaaaccagcgaagcaaaagatacgcttcaccagctgagtgaactggcg- gaattattgattgtcagcgcccgt gactggctgtatgccgcttgctgtcgcgagttcggtacgccggtcaatgccgcaggggaaccgcagagaatgc- tgatcctcgggatgggcaaactc ggcggtggcgagctgaatttcatcggacatcgacctgacctgatttttgcttatccggaaaatggccagacac- gcggcggtcggcgtgaactggataacgc acaattttcacccggctcggccagcgtctgatcaaagcgctggatcagcccactatcgacggttttgtctatc- gcgtggacatgcgtttgcgtccgttcg gcgacagtggcccgctggtgatgagcttcccggcactggaagattattatcaggaacaggggcgcgactggga- acgctacgcaatggtgaaagcg cgtctgatgggcggcgcggaggacatcagcagtcaggaattgcgtaaaatgctgatgcttttgtcttccgccg- ttatatcgatttcagtgtgatccagtc cctgcgtaacatgaaaggcatgatcgcccgcgaagtacgccgccgtggtctgaaagacaacatcaaactcggc- gcaggcggtattcgtgaaattgaa tttatcgtgcaggtatttcagctgatccgtggcggtcgtgaaccggcattgcagcagcgtgcgttgttgccaa- cgcttcaggcgctggaaaatctgggg ctgctgccggtagagcaggtgttgcagttgcgtaacagctatctgttcctgcgacgtctggaaaacctgttgc- aggccattgctgacgagcaaacgcaa accttaccgtccgatgagctgaatcaggcgcgtctggcgtgggggatgaattacgctggctggccgcagctta- ggatgcagtgaatgctcacatgca ggccgtacgcgcggtatttaacgatctgattggcgatgacacgccagatgccgaagatgacgtgcaactctcc- cggttcagcagtttatggattgatac gcttgagcctgacgagctggctccgctggtgccgcaacttgacgaaaatgcgcaacggcatgttttacatcag- attgctgattttcgccgtgacgtggat aaacgcacgatagggccacgtgggcgtgatcagttggatttgctgatgccgcgtttactggcccaggtctgca- cctataaaaatgcggatgtgacgtta cagcgcctgatgcagttgctgctcaatatcgtcacgcgcacgacgtatatcgagctgctggtggaatatcccg- gtgcgctcaaacagttaatacgtctgt gcgctgcctcgccgatggtggcgacgcaacttgcgcgtcatcctttattgctcgacgaactgctcgacccgcg- cacgctttaccagccgattgagccg ggcgcgtaccgtgatgaactgcggcaatatctgatgcgggtgccaaccgaagacgaagaacaacagcttgaag- ccgtgcgccagttcaaacaggc acagcatttacgtattgcggccggggatatttccggtgcgttgccggtgatgaaagtcagtgaccatttaacc- taccttgcggaggccattctgacgtt gtggttgcaacaggcgtgggaacaaatggtcgtaaaatacggtcagccaacccatcttcagcaccgtaaaggg- cgcggttttgccgtggtgggttacg gaaaactcggtggctgggagctgggttacagctcggatctggatctggtcttcctgctcgattgcgcgccgga- agtcatgaccgacggcgaacgcag cattgacgggcgtcagttttatctgcggctggcgcagcgcatcatgcatttattagcacccgtacgtcgtcag- gcattctttatgaggttgacccgcgtc tgcggccttccggtgcttccggcatgctggtcagcaccatcgaagcttttgcggattatcaggccaacgaagc- ctggacatgggagcatcaggcgctg gttcgcgcgcgtgtggtttatggtgatccgcaactgacgcagcaatttaatgccacgcgtcgcgacattcttt- gccgccagcgcgatgccgacggcttg cgtaaggaagtccgtgaaatgcgcgagaaaatgtatgcccatctgggcagcaaaagagccgacgagtttgatc- tgaaagccgatccgggtggcata acggatattgaattcatcgcacaatatctggttctgcgtttcgcgcatgatgagccgaagagacccgctggta- gataacgtgcggattttcgaactgat ggcgcgacatgacatcatgccggaagaggaagcacgccatctgacgcaggcttacgtgacattgcgcgatgaa- attcatcatctggcgtgcaggaa
cacagcgggaaagtggccgcagacagctttgccactgagcgcgcccaaatccgcgccagctgggcaaactggc- ttggctga 94 atgaaaaaacttttatccatgatggggcttggtgcagtggctttgctaccttcgcttgccatggcagcacc- accagcagcggcaaacggtgctgataac gcctttatgatgatttgtaccgcgctggtattgttcatgaccgtacccggtgtggcgttgttctacggcggct- tactgcgttctaaaaacgttttgtccatgct gactcaggttattgttacctttgctctggtctgcgtcctgtggatcctctacggttacagccttgccttcagt- gaaggtaacgcgttcttcggtggtttcagca acgtaataatgaaaggcattggcctggattctgtgactggcaccttctcgcagatgatccacgttgcattcca- gtgttcatttgcctgcatcactgtagcgc tgatcgtaggtggtattgctgaacgtgtgcgtttctcagcagttctgattttcactgtgatctggctgacttt- ctcttatattccgatggctcacatggtatggg caggcggtttcctggctgctgacggtgcgctggactttgccggtggtaccgttgttcatatcaatgccgcaat- tgctggcctggtaggggcttatctgctg ggtaaacgcgccggttttggcaaagaagctttcaaaccacacaacctgccaatggtcttcactggcgcctcaa- tcctgtatgtgggctggttcggcttca atgcgggttcagcaagtgccgcaagctctgttgccgcgctggctttcctgaacactgtcattgctactgctgg- cgcaatcctgtcctggacgctggttga gtggatggtgcgcggtaagccctcactgctgggcgcaagctccggtgctatcgcaggtctggtggctatcacg- cctgcatgtggtacggtcggcgta ggtggtgctctgattatcggtctggtaggcggtatcactggtctgtggggggttgttaccctgaaaaaatggc- tgcgtgttgatgacacctgtgatgtgtt cggtgttcatggcgtgtgcggtatcgtaggttgtctgctgacgggtgtattcacgtccagttcacttggcggc- gtgggctacgcagaaggcgtgaccat gggccatcaggtttgggtgcagttcttcagcgtgtgcgtaacattggtctggtcaggcgttgttgccttcatc- ggttacaaagtggctgacatgatcgtag gtctgcgtgttcctgaagaacaagaacgcgaaggtctggacgttaacagccacggcgaaaacgcttacaacca- ataa 95 tgaatatcactgactcacaagctacctatgtcgaagaattaactaaaaaactgcaagatgcaggcattcgc- gttaaagccgacttgagaaatgagaaga ttggctttaaaattcgcgaacacacgctacgccgtgttccttatatgttagtttgtggcgataaagaggtcga- agcaggcaaagttgctgttcgtactcgtc gcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaactgctggcggaaatccgcagcagaag- tcatcatcaactggaggaataaa gtattaaaggcggaaaacgagttcaaccggcgtcctaatcgcattaacaaagagattcgcgcgcaagaagttc- gcctcaccggcgtcgatggcga gcagattggtattgtcagtctgaatgaagctcttgaaaaagctgaggaagcgggcgtcgatttagtagaaatc- agtccgaatgccgagccgccagtttg tcgaatc 96 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgatagtcgagcggtagc- acagagagcttgctctcgggtgacga gcggcggaccggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaata- ccgcataacgtcgcaagaccaaa gtgggggaccttcgggcctcatgccatcagatgtgcccagatgggattagctagtaggtggggtaacggctca- cctaggcgacgatccctagctggt ctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatat- tgcacaatgggcgcaagcctga tgcagccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaaggcggtgagg- ttaataacctcaccgattgacgtta cccgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaa- ttactgggcgtaaagcgcacgc acgcggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcattcgaaactggcaggctaga- gtcttgtagaggggggtagaattc caggtgtagggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactga- cgctcaggtgcgaaagcgtg gggagcaaacaggattagataccctggtagtccacgctgtaaacgatgtcgatttggaggttgtgcccttgag- gcgtggcttccggagctaacgcgtta aatcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtgg- agcatgtggtttaattcgatgca acgcgaagaaccttacctggtcttgacatccacagaactttccagagatggattggtgccttcgggaactgtg- agacaggtgctgcatggctgtcgtca gctcgtgttgtgaaatgttggg 97 atgaccatgcgtcaatgcgctatctacggtaaaggcggtatcggtaaatccaccaccacccagaatctcgt- cgcggccctcgccgagatgggtaaga aagtgatgatcgtcggctgcgatccgaaagcggattccacccgtctgatcctccacgctaaagcccagaacac- catcatggagatggcggcggaagt gggctcggtcgaggatctggagctcgaagacgttctgcaaatcggctatggcgatgtccgttgcgccgaatcc- ggcggcccggagccaggcgtcg gctgcgccggacgcggggtgatcaccgccatcaacttcctcgaggaagaaggcgcctatgaagaagatttgga- tttcgtcttctatgacgtcctcggc gacgtcgtctgcggcggcttcgctatgccgatccgcgaaaacaaagcccaggagatctacatcgtctgctccg- gcgagatgatggcgatgtatgccg ccaacaatatctccaaagggatcgtgaagtacgccaaatccggcaaggtgcgcctcggcggcctgatctgtaa- ctcgcaaaaccgaccgggaag acgaactgatcatcgccctggcggagaagcttggcacgcagatgatccacttcgttccccgcgacaacattgt- gcagcgcgcggagatccgccgga tgacggtgatcgagtacgacccgacctgtcagcaggcgaatgaatatcgtcaactggcgcagaagatcgtcaa- taacaccaaaaaagtggtgccga cgccgtgcaccatggacgagctggaatcgctgctgatggagttccgcatcatggaagaagaagacaccagcat- cattggtaaaaccgccgctgaag aaaacgcggcctga 98 atggttaggaaaagtagaagtaaaaatacaaatatagaactaactgaacatgaccatttattaataagtca- aataaaaaagcttaaaacacaaaccacttg cttttttaataataaaggaggggttgggaagactacattagtagcaaatttaggagcagagctatcaataaac- tttagtgcaaaagttcttattgtggatgcc gaccctcaatgtaatctcacgcagtatgtatttagtgatgaagaaactcaggacttatatgggcaagaaaatc- cagatagtatttatacagtaataagacc actatcctttggtaaaggatatgaaagtgacctccctataaggcatgtagagaatttcggttttgacataatt- gtcggtgaccctagacttgctttacaggaa gaccttttagctggagactggcgagatgccaaaggcggtgggatgcgaggaattaggacaacttttgtatttg- cagagttaattaagaaagctcgtgag ctaaattatgattttgttttctttgacatgggaccatcattaggcgcaatcaacagggcagtattactggcaa- tggaattctttgtcgtcccaatgtcaatcga tgtattttcactatgggctattaaaaatattggctccacggtttcaatatggaaaaaagaattagacacaggg- attcggctctcagaggaacctagcgaatt atcacaattatcacctcaaggaaaactaaagtttctcggttacgtcacccaacaacataaagaacgctctgga- tacgatacaattcagcttgagaatactg aggaagaaataaaatcgaaacgtcgggtaaaggcgtatgaagacattggagaggtgtttccttctaaaattac- tgagcatctttctaaactttatgcatca aaagatatgaacccacaccttggagatatacgtcatttaggtagtttagctccgaaatcacaatcacaacacg- ttccgatgatatcagtgtctggtacagg aaattacaccagacttagaaaaagcgcgcgtgaactttatcgagatattgcaagaagatacttagagaacatt- cagactgctaatggcgagaaatag 99 atgaagggaaaggaaattctggcgctgctggacgaacccgcctgcgagcacaaccagaagcaaaaatccgg- ctgcagcgcccctaagcccggcg ctaccgccggcggttgcgccttcgacggcgcgcagataacgctcctgcccatcgccgacgtcgcgcacctggt- gcacggccccatcggctgcgcg ggcagctcgtgggataaccgcggcagcgtcagcgccggcccggccctcaaccggctcggctttaccaccgatc- ttaacgaacaggttgtgattatg ggccgcggcgaacgccgcctgttccacgccgtgcgtcacatcgtcgaccgctatcatccggcggcggtcttta- tctacaacacctgcgtaccggcga tggagggcgatgacatcgaggcggtctgccaggccgcacagaccgccaccggcgtcccggtcatcgctattga- cgccgccggtttctacggcagta aaaatcttggcaaccgaatggcgggcgacgtgatgctcaggcaggtgattggccagcgcgaaccggccccgtg- gccagacaacacgccctttgcc ccggcccagcgccacgatatcggcctgattggcgaattcaatatcgccggcgagttctggcaggtccagccgc- tgctcgacgagctggggatccgc gtcctcggcagcctctccggcgacggccgctttgccgagatccagaccctgcaccgggcgcaggccaatatgc- tggtgtgctcgcgcgcgctgatc aacgtcgcccgggggctggagctgcgctacggcacgccgtggtttgaaggatgcttctacgggatccgcgcca- cctccgacgccttgcgccagct ggcgacgctgctgggggatgacgacctgcgccgccgcaccgaggcgctgatcgcccgcgaagagcaggcggcg- gagcaggctcttgcgccgt ggcgtgagcagctccgcgggcgcaaagtgctgctctataccggcggcgtgaaatcctggtcggtggtatcggc- cctgcaggatctcggcatgaccg tggtggccaccggcacgcgcaaatccaccgaggaggacaaacagcggatccgtgagctgatgggcgacgaggc- ggtgatgcttgaggagggca atgcccgcaccctgctcgacgtggtgtaccgctatcaggccgacctgatgatcgccggcggacgcaatatgta- caccgcctggaaagcccggctgc cgttctcgatatcaatcaggagcgcgagcacgcctacgccggctatcagggcatcatcaccctcgcccgccag- ctctgtctgaccctcgccagcccc gtctggccgcaaacgcatacccgcgccccgtggcgctag 100 atgaccaacgcaacaggcgaacgtaaccttgcgctcatccaggaagtcctggaggtgtttcccgaaaccg- cgcgcaaagagcgcagaaagcacat gatgatcagcgatccgcagatggagagcgtcggcaagtgcattatctcgaaccgtaaatcgcagcccggggtg- atgaccgtgcgaggctgcgccta tgcgggctcgaaaggggtggtgtttgggccaatcaaagacatggcccatatctcgcacggccccatcggctgc- ggccagtattcccgcgccggacg gcgcaactactataccggcgtcagcggtgtcgacagcttcggcaccctgaacttcacctctgattttcaggag- cgcgatattgttttcggcggcgataaa aagctgaccaaactgatcgaagagatggagctgctgttcccgctgaccaaagggatcaccatccagtcggagt- gcccggtgggcctgatcggcgat gacatcagcgccgtagccaacgccagcagcaaggcgctggataaaccggtgatcccggtgcgctgcgaaggct- ttcgcggcgtatcgcaatcgct gggccaccatatcgccaacgacgtggtgcgcgactgggtgctgaacaatcgcgaagggcagccgtttgccagc- accccgtacgatgttgccatcatt ggcgattacaacatcggcggcgacgcctgggcctcgcgcattctgctggaagagatgggcctgcgcgtagtgg- cgcagtggtccggcgacggca ccctggtggagatggagaacaccccattcgttaagcttaacctcgtccactgctaccgttcgatgaactatat- cgcccgccatatggaggagaaacatc agatcccatggatggaatataacttcttcggcccgaccaaaatcgccgaatcgctgcgcaagatcgccgatca- atttgatgacaccattcgcgccaatg cggaagcggtgatcgcataatatgaggggcagatggcggccatcatcgccaaatatcgcccgcggctggaggg- gcgcaaagtgctgctgtacatg ggggggctgcggccgcgccacgtcatcggcgcctatgaggatctcgggatggagatcatcgccgccggctacg- agtttgcccataacgatgattac gaccgcaccctgccggacctaaagagggcaccctgctgtttgacgatgccagcagctatgagctgtgaggcct- tcgtcaaagcgctgaaacctgac ctcatcggctccgggatcaaagagaaatatatcttccagaaaatgggggtgccgttccgccagatgcactcct- gggactattccggcccctatcacgg ctatgacggcttcgccatctttgcccgcgatatggatatgaccctgaacaatccggcgtggaacgaactgact- gccccgtggctgaagtctgcgtga 101 atggcagatattatccgcagtgaaaaaccgctggcatgagcccgattaaaaccgggcaaccgctcggggc- gatcctcgccagcctcgggctggc ccaggccatcccgctggtccacggcgcccagggctgcagcgccttcgccaaagttttctttattcagcatttc- catgacccggtgccgctgcagtcgac ggccatggatccgaccgccacgatcatgggggccgacggcaatatcttcaccgcgctcgacaccctctgccag- cgccacagcccgcaggccatcg tgctgctcagcaccggtctggcggaagcgcagggcagcgatatcgcccgggtggtgcgccagtttcgcgaggc- gcatccgcgccataacggcgtg gcgatcctcaccgtcaataccccggatttttttggctctatggaaaacggctacagcgcggtgatcgagagcg- tgatcgagcagtgggtcgcgccgac gccgcgtccggggcagcggccccggcgggtcaacctgctggtcagccacctctgttcgccaggggatatcgaa- tggctgggccgctgcgtggag gcctttggcctgcagcccgtgatcctgccggacctctcgcagtcaatggatggccacctcggtgaaggggatt- ttacgcccctgacccagggcggcg cctcgctgcgccagattgcccagatgggccagagtctgggcagcttcgccattggcgtgtcgctccagcgggc- ggcatcgctcctgacccaacgca gccgaggcgacgtgatcgccctgccgcatctgatgaccctcgaccattgcgatacctttatccatcagctggc- gaagatgtccggacgccgcgtacc ggcctggattgagcgccagcgtggccagctgcaggatgcgatgatcgactgccatatgtggcttcagggccag- cgcatggcgatggcggcggagg gcgacctgctggcggcgtggtgtgatttcgcccgcagccaggggatgcagcccggcccgctggtcgcccccac- cagccaccccagcctgcgcca gctgccggtcgagcaagtcgtgccgggggatcttgaggatctgcagcagctgctgagccaccaacccgccgat- ctgctggtggctaactctcacgc ccgcgatctggcggagcagtttgccctgccgctgatccgcgtcggttttcccctcttcgaccggctcggtgag- tttcgtcgcgtccgccaggggtacgc cggtatgcgagatacgctgtttgaactggccaatctgctgcgcgaccgccatcaccacaccgccctctaccgc- tcgccgcttcgccagggcgccgac ccccagccggcttcaggagacgcttatgccgcccattaa 102 atgagccaaacgatcgataaaattcacagctgttatccgctgtttgaacaggatgaataccagaccctgt- tccagaataaaaagacccttgaagaggcg cacgacgcgcagcgtgtgcaggaggtttttgcctggaccaccaccgccgagtatgaagcgctgaaatccagcg- cgaggcgctgaccgtcgaccc ggccaaagcctgccagccgctcggcgccgtactctgcgcgctggggttcgccggcaccctgccctacgtgcac- ggctcccagggctgcgtcgcct attttcgcacctactttaaccgccattttaaagagccggtcgcctgcgtctccgactccatgaccgaggacgc- ggcggtgttcggcggcaacaacaac atgaatctgggcctgcagaatgccagcgcgctgtataaacccgagattatcgccgtctccaccacctgtatgg- ccgaggtgatcggcgacgatctgca ggcgtttatcgccaacgccaaaaaagagggatttgttgacgaccgcatcgccattccttacgcccataccccc- agctttatcggcagccatgtcaccgg ctgggacaatatgttcgaagggttcgcgaagacctttaccgctgactacgccgggcagccgggcaaacagcaa- aagctcaatctggtgaccggattt gagacctatctcggcaacttccgcgtgctgaagggatgatggcgcagatggatgtcccgtgcagcctgctctc- cgacccatcagaggtgctcgaca cccccgccgacggccattaccggatgtacgccggcggcaccagccagcaggagatcaaaaccgcgccggacgc- cattgacaccctgctgctgca gccgtggcagctggtgaaaagcaaaaaggtggttcaggagatgtggaaccagcccgccaccgaggtggccgtt- ccgctgggcctggccgccacc gacgcgctgctgatgaccgtcagtcagctgaccggcaaaccgatcgccgacgactgaccaggagcgcggccgg- ctggtcgacatgatgctggat tcccacacctggctgcatggcaaaaaattcggcctctacggcgatccggatttcgtgatggggctgacgcgct- tcctgctggagctgggctgcgagcc gacggtgatcctcagtcataacgccaataaacgctggcaaaaagcgatgaagaaaatgctcgatgcctcgccg- tacggtcaggaaagcgaagtgttc atcaactgcgacctgtggcacttccggtcgctgatgttcacccgtcagccggactttatgatcggtaactcct- acggcaagtttatccaggcgataccc tggcaaagggcaaagccttcgaagtgccgctgatccgtctgggctttccgctgttcgaccgccatcatctgca- ccgccagaccacctggggctatgaa ggcgcaatgaacatcgtcacgacgctggtgaacgccgtgctggaaaaactggaccacgacaccagccagttgg- gcaaaaccgattacagcttcgac ctcgttcgttaa 103 atgaccctgaatatgatgctcgataacgccgcgccggaggccatcgccggcgcgctgactcaacaacatc- cggggctgttttttaccatggtggaaca ggcctcggtggccatctccctcaccgatgccagcgccaggatcatttacgccaacccggcgttttgccgccag- accggctattcgctggcgcaattgtt aaaccagaacccgcgcctgctggccagcagccagacgccgcgcgagatctatcaggagatgtggcataccctg- ctccagcgtcagccctggcgcg gtcagctgattaatcagcgtcgggacggcggcctgtacctggtggagattgacatcaccccggtgcttagccc- gcaaggggaactggagcattatct ggcgatgcagcgggatatcagcgtcagctacaccctcgaacagcggctgcgcaaccatatgaccctgatggag- gcggtgctgaataatatccccgc cgccgtggtagtggtggacgagcaggatcgggtggtgatggacaacctcgcctacaaaaccttctgcgctgac- tgcggcggccgggagctgctcac cgagctgcaggtctcccctggccggatgacgcccggcgtggaggcgatcctgccggtggcgctgcgcggggcc- gcgcgctggctgtcggtaacc tgctggccgttacccgccgtcagtgaagaggccagccgctactttatcgacagcgcgctggcgcggaccctgg- tggtgatcgccgactgtacccag cagcgtcagcagcaggagcaagggcgccttgaccggctgaagcagcaaatgaccgccggcanctgctggcggc-
gatccgcgagtcgctggac gccgcgctgatccagctgaactgcccgattaatatgctggcggcagcccgtcggctgaacggcgagggaagcg- ggaatgtggcgctggaggccg cctggcgtgaaggggaagaggcgatggcgcggctccagcgctgtcgcccatcgctggaactcgaaaaccccgc- cgtctggccgctgcagccctttt tcgacgatctgtgcgccctctaccgtacacgcttcgatcccgacgggctgcaggtcgacatggcctcaccgca- tctgatcggctttggccagcgcacc ccactgctggcgtgcttaagcctgtggctcgatcgcaccctggccctcgccgccgaactcccctccgtgccgc- tggcgatgcagctctacgccgagg agaacgacggaggctgtcgctgtatctgactgacaacgtaccgctgctgcaggtgcgctacgctcactccccc- gacgcgctganctcgccgggca aaggcatggagctgcggctgatccagaccctggtggcgcaccatcgcgagccattgagctggcttcccgaccg- cagggcggcacctgcctgacc ctgcgtttcccgctgtttaacaccctgaccggaggtgaagcatga 104 atgatccctgaatccgacccggacaccaccgtcagacgcttcgacctctctcagcagttcaccgccatga- gcggataagcgtggtgctgagccggg ccaccgaggccagcaaaacgctgcaggaggtgctcagcgtattacacaacgatgcctttatgcagcacgggat- gatctgcctgtacgacagcgagc aggagatcctcagtatcgaagcgctgcagcaaaccggccagcagcccctccccggcagcacgcagatccgcta- tcgccccggcgagggactggt ggggaccgtgctggcccaggggcagtcgctggtgctgccccgggtcgccgacgatcagcgttttctcgaccgc- ctgagcactacgattacgatctg ccgtttatcgccgtaccgttgatggggcccaacgcccggccaataggggtgctggcggcccagccgatggcgc- gccaggaagagcggctgccgg cctgcacccgttttctcgaaaccgtcgccaacctcgtcgcccagaccatccggctgatgatccttccggcctc- acccgccctgtcgagccgccagccg ccgaaggtggaacggccgccggcctgctcgtcgtcgcgcggcgtgggccttgacaatatggtcggcaagagcc- cggcgatgcgccagatcgtgg aggtgatccgtcaggtttcgcgctgggacaccaccgtgctggtacgcggcgaaagcggcaccgggaaagagct- gatcgccaacgccatccatcac cattcgccacgggctggcgccgccttcgtcaaatttaaactgcgcggcgctgccggacaccctgctggaaagc- gaactgttcggccatgagaaaggc gcctttaccgagcggtgcgtcagcgtaaaggacgttttgagctggcggatggcggcaccctgttcctcgatga- gattggtganagcagcgcctcg ccaggccaagctgctgcgatcctccaggagggggagatggagcgggtcggcggcgatgagaccctgcgggtga- atgtccgcatcatcgccgcc accaaccgtcacctggaggaggaggtccggctgggccatttccgcgaggatctctactatcgctgaacgtgat- gccatcgccctgcccccgagc gcgagcgtcaggaggacatcgccgagctggcgcacttcctggtgcgcaaaatcggccagcatcaggggcgcac- gctgcggatcagcgagggcg cgatccgcctgctgatggagtacagctggccgggtaacgttcgcgaactggagaactgcctcgaacgatcggc- ggtgatgcggagagtggcctga tcgatcgcgacgtgatcctcttcactcaccaggatcgtcccgccaaagccctgcctgccctgcgggccagcgg- aagacagctggctggacaacagcc tggacgaacgtcagcgactgatcgccgcgctggaaaaagccggctgggtgcaggccaagccggcacggctgct- ggggatgacgccgcgccagg tcgcttatcggatccagatcatggatatcaccctgccgcgtctgtag 105 atgatgccgctttctccgcaattacagcagcactggcagacggtcgctgaccgtctgccagcggattttc- ccattgccgaactgagcccacaggccag gtccgtcatggcgttcagcgattttgtcgaacagagtgtgatcgcccagccgggctggctgaatgagcttgcg- gactcctcgccggaggcggaagag tggcggcattacgaggcctggctgcaggatcgcctgcaggccgtcactgacgaagcggggttgatgcgagagc- tgcgtctcttccgccgccagatg atggtccgcatcgcctgggcgcaggcgctgtcgctggtgagcgaagaagagactctgcagcagctgagcgtcc- tggcggagaccctgattgtcgcc gcccgcgactggctgtacgccgcctgctgtaaggagtggggaacgccatgcaatgccgagggccagccgcagc- cgctgctgatcctcgggatgg gaaaagctgggcggcggcgagctgaacttctcttccgatatcgatctgatctttgcctggcctgagcatggcg- ccacccgcggcggccgccgcgagct ggataacgcccaggctttacccgtctggggcagcggctgatcaaggcccttgaccagccgacgcaggacggct- ttgtctatcgggttgacatgcgcc tgcggccgtttggcgacagtgggccgctggtactcagttttgcggcgctggaagattattaccaggagcaggg- tcgggactgggaacgctatgcgat ggtgaaagcgcggatcatgggcgataacgacggcgtgtacgccagcgagttgcgcgcgatgctccgtcctttc- gtcttccgccgttatatcgacttca gcgtgatccagtcgctgcgtaacatgaaaggcatgatcgcccgcgaagtgcggcgtcgcgggctgaaagacaa- catcaagctcggcgccggcgg gatccgtgaaattgagtttatcgttcaggtctttcaactgatccgcggtggtcgcgaacctgcactgcagcag- gcgccctgctgccgacgctggcgg cgattgatgagctacatctgctgccggaaggcgacgcggcgctgctgcgcgaggcctatctgttcctgcgccg- gctggaaaacctgctgcaaagcat caacgatgagcagacccagaccctgccgcaggatgaacttaaccgcgccaggctggcgtgggggatgcatacc- gaagactgggagacgctgagc gcgcagctggcgagccagatggccaacgtgcggcgagtgtttaatgaactgatcggcgatgatgaggatcagt- ccccggatgagcaactggccga gtactggcgcgagctgtggcaggatgcgctggaagaagatgacgccagcccggcgctggcgcatttaaacgat- accgaccgccgtagcgtgctgg cgctgattgccgattttcgtaaagagctggatcggcgcaccatcggcccgcgggccgccaggtgctggatcag- ctgatgccgcatctgctgagcga aatctgctcgcgcgccgatgcgccgctgcctctggcggatcacgccgctgttgaccgggatcgtcacccgtac- cacctatcttgagctgctgagc gaattccccggcgcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgc- gccacccgctgctgctggatga gctcgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacct- gctgcgcgtgccggaagaggatga agagcagcagctggaggcgttgcgccagtttaagcaggcgcagcagagcatatcgcggcggcggatatcgctg- gtaccctgccggtgatgaagg tcagcgatcatttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggt- cgctcgctacggccagccgaccc acctgcacgatcgccagggtcgcggcttcgccgtcgtcggctacggtaagcttggcggctgggagctgggcta- cagctccgatctcgatctggtgttc ctccatgactgcccggcggaggtgatgaccgacggcgagcgggagattgacggccgtcagttctacctgcggc- tggcccagcggatcatgcacct gttcagcacccgcacctcgccggtattctctacgaagtggacgcccggctgcgtccttctggcgcggcgggga- tgctggtcaccaccgccgacgc gtttgctgactatcagcagaacgaagcctggacgtgggaacatcaggcgctggtgcgcgcccgcgtggtctat- ggcgacccggcgctgcaggcgc gctttgacgccattcgtcgcgatatcctgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcga- gatgcgcgagaagatgcgcgcc caccttggcaacaaacatcccgatcgtttgatatcaaagccgatgccggcgggatcaccgatattgaatttat- tactcagtatctggtcctacgctatgcc agtgacaagccgaagctgacccgctggtctgacaacgtgcgtattcttgagctgctggcgcagaagacatcat- ggacgaggaggaggcgcgcgc cttaacgcatgcgtacaccaccttgcgtgatgcgctccatcacctggccctgcaggagcagccgggacacgtg- gcgccagaggccttcagccggga gcgtcagcaggtcagcgccagctggcagaagtggctgatggcttaa 106 agcgtcaggtaccggtcatgattcaccgtgcgattctcggttccctggagcgcttcattggcatcctgac- cgaagagttcgctggcttcttcccaacctg gattgcaccagtgcaggtagtggtcatgaatattaccgattctcaggctgaatacgttaacgaattgacgcgt- aaactacaaaatgcgggcattcgtgta aaagcagacttgagaaatgagaagattggctttaaaatccgcgagcacactttacgtcgtgcccgtatatgtt- ggtctgtggcgacaaagaagtcgaa gccggcaaagtggccgtgcgcacccgtcgcgggaaagacctcggcagcatggacgtaagtgaagtgattgaga- agctgcaacaagagattcgca gccgcagtcttcaacaactggaggaataaggtattaaaggcggaaaacgagttcaaacggcacgtccgaatcg- tatcaatggcgagattcgcgccct ggaagttcgc 107 atgaaaatggcaacaatgaaatcgggtctgggggcattagcccttcttccgggactggcaatggccgcgc- ccgcagtggcggacaaagccgataac gcgtttatgatgatttgcaccgcgctggttctgtttatgaccatcccggggatcgcgctgttttacggcggcc- tgatccgcggcaaaaacgtcctttccat gctgactcaggtgattgtgacctttggcctcgtctgcgtactgtgggtgatttatggctataccctggccttc- ggcaccggcggcagcttcttcgctagttt tgactgggtgatgctgaaaaatattgaactgaaagcgctgatgggcaccttctatcagtacatccacgtggca- tccagggctcgttcgcctgtatcacc gtcgggctgatcgtgcgggcgctggctgagcgtattcgtttctccgccgtgctgatttttgtggtggtgtgga- tgacgctctcttatgttccgattgcgcac atggtctggggcggcggtctgctggcgacccacggcgcgctggacttcgcgggcggcaccgttgtacacatca- acgccgcggttgccgggctggt gggtgcgtacatgatgggcaaacgtggggcttcggcaaagaagcgttcaaaccgcacaatctgccgatggtgt- tcaccggaaccgccatcctctac gtgggctggttcggcttcaacgccggctccgccagcgcagcgaacgaaattgccgcattggctttcgtcaaca- ccgtcgtcgccacagcggctgcca tcctggcgtggacctttggcgaatgggccctgcgcggtaaaccttcactgctgggcgcctgctccggggcgat- tgccggtctggttggcgtcacacca gcctgtgggtatatcggtgtcggtgggcgttgattgtgggtatcgcatctggtctggcgggcatctggggcgt- aacggcgctgaaacgctggctgcg ggttgatgacccttgcgacgtcttcggcgtccacggcgtctgcggcatcgtcggctgtatcctgaccggtatc- ttcgcggccacctctctgggcggcgt gggttatgcagaaggcgtcaccatgggccatcagctgctggtgcaactcgagagtatcgcgattaccatcgtc- tggtcgggcgttgtcgctttcattgg ctacaaagtggcggacatgaccgtggggctgcgcgtaccagaagagcaggagcgcgaaggactggacgtcaac- agccatggcgaaaacgccta caacgcctga 108 cgccgtcctcgcagtaccattgcaaccgactttacagcaagaagtgattctggcacgcatggaacaaatt- cttgccagtcgggctttatccgatgacga acgcgcacagcttttatatgagcgcggagtgttgtatgatagtctcggtctgagggcattagcgcgaaatgat- ttttcacaagcgctggcaatccgaccc gatatgcctgaagtattcaattacttaggcatttacttaacgcaggcaggcaattttgatgctgcctatgaag- cgtttgattctgtacttgagcttgatc 109 gctaaagttctcggctaatcgctgataacatttgacgcaatgcgcaataaaagggcatcatttgatgccc- tttttgcacgctttcataccagaacctggctc atcagtgattttttttgtcataatcattgctgagacaggctctgaagagggcgtttatacaccaaaccattcg- agcggtagcgcgacggcaagtcagcgtt ctcctttgcaatagcagggaagaggcgccagaaccgccagcgttgaagcagtttgaacgcgttcagtgtataa- tccgaaacttaatttcggtttgga 110 gcccgctgaccgaccagaacttccaccttggactcggctatacccttggcgtgacggcgcgcgataactg- ggactacatccccattccggtgatctta ccattggcgtcaataggttacggtccggcgactttccagatgacctatattcccggcacctacaataacggta- acgtttacttcgcctgggctcgataca gttttaattcgctaagtcttagcaataaatgagataagcggtgtgtcttgtggaaaaacaaggactaaagcgt- tacccactaaaaaagatagcgacttttat cactttttagcaaagttgcactggacaaaaggtaccacaattggtgtactgatactcgacacagcattagtgt- cgatttttcatataaaggtaattttg 111 ttgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcggtagc- acagagagcttgctctcgggtgacgag cggcggacgggtgagtaatgtagggaaactgcctgatggagggggataactactggaaacggtagctaatacc- gcataacgtcgcaagaccaaag tgggggaccttcgggcctcatgccatcagatgtgcccagatgggattagctagtaggtggggtaacggctcac- ctaggcgacgatccctagctggtct gagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatattg- cacaatgggcgcaagcctgat gcagccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaaggcgntnaggt- taataaccttgtcgattgacgttac ccgcagaagaagcaccggctaactccggccagcagccgcggtaatacggagggtgcaagcgttaatcggaatt- actgggcgtaaagcgcacgca ggcggtagtcaagtcggatgtgaaatccccgggctcaacctgggaactgcattcgaaactggcaggctagagt- cttgtagaggggggtagaattcc agggtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactgac- gctcaggtgcgaaagcgtgg ggagcaaacaggattagaccctggtagtccacgctgtaaacgatgtcgatttggaggttgtgcccttgaggcg- tggcttccggagctagcgcgttaa atcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtgga- gcatgtggtttaattcgatgcaa cgcgaagaaccttacctggtcttgacatccacagaactttccagagatggattggtgccttcgggaactgtga- gacaggtgctgcatggctgtcgtcag ctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcggtnnggcc- gggaactcaaaggagactgccagtg ataaactggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccagggctacacacgtgctacaa- tggcatatacaaagagaagcgac ctcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtctgcaactcgactccatgaagt- cggaatcgctagtaatcgtagatca gaatgctacggtgaatacgttcccgggccttgtacacaccgcccgtacaccatgggagtgggttgcaaaagaa- gtaggtagcttaaccttcgggag ggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttgg- atcacctcctt 112 atgaccatgcgtcaatgcgctatctacggtaaaggcggtatcggtaaatccaccaccacccagaatctcg- tcgcggcctcgccgagatgggtaaga aagtgatgatcgtcggctgcgatccgaaagcggactccacccgtctgatccttcacgctaaagcccagaacac- catcatggagatggcggcggaagt gggctcggtcgaggatctggagctcgaagacgttctgcaaatcggctatggcgatgtccgttgcgccgaatcc- ggcggcccggagccaggcgtcg gctgcgccggacgcggggtgatcaccgccatcaacttcctcgaggaagaaggcgcctatgaggaagatttgga- tttcgtcttctatgacgtcctcggc gacgtagtctgcggcggcttcgccatgccgatccgcgaaaacaaagcccaggagatctacatcgtctgctccg- gcgagatgatggcgatgtatgccg ccaacaatatctccaaggggatcgtgaagtacgcgaaatctggcaaggtgcgcctcggcggcctgatctgtaa- ctcgcgcaaaaccgaccgggaag acgaactgatcatcgccctggcggagaagcttggcacgcagatgatccacttcgttccccgcgacaacattgt- gcagcgcgcggagatccgccgga tgacggtgatcgagtacgacccgacctgtcagcaggcgaatgaatatcgtcaactggcgcagaagatcgtcaa- taacaccaaaaaagtggtgccaa cgccgtgcaccatggacgagctggaatcgctgctgatggagttcggcatcatggaagaagaagacaccagcat- cattggtaaaaccgccgctgaag aaaacgcggcctga 113 atgaccaacgcaacaggcgaacgtaaccttgcgctcatccaggaagtcctggaggtgtttcccgaaaccg- cgcgcaaagagcgcagaaagcacat gatgatcagcgatccgcagatggagagcgtcggcaagtgcattatctcgaaccgtaaatcgcagcccggggtg- atgaccgtgcgtggctgcgcctat gcgggcttcgaaaggggtggtgtttgggccaatcaaagacatggcccatatctcgcacggccccatcggctgc- ggccagtactgcgcgccggacg gcgcaactactataccggcgtcagcggtgtcgacagcttcggcaccctgaacttcacctctgattttcaggag- cgcgatattgttttcggcggcgataaa aagctgaccaaactgatcgaagagatggagctgcttcccgctgctccaaagggatcaccatccaatcggagtg- cccggtgggcctgatcggcgat gacatcagcgccgtggccaacgccagcagcaaggcgctggataaaccggtgatcccggtgcgctgcgaaggct- ttcgcggcgtatcgcaatcgct gggccaccatatcgccaacgacgtggtgcgcgactgggtgctgaacaatcgcgaagggcagccgtttgccagc- accccgtatgatgttgccatcatt ggcgattacaacatcggcggcgacgcctgggcctcgcgcattctgctggaagagatggggctgcgcgtagtgg- cgcagtggtccggcgacggca ccctggttgagatggagaacaccccattcgttaagcttaacctcgccactgctaccgacgatgaactatatcg- cccgccatatggaggagaaacatc agatcccgtggatggaatataacttcttcggcccgaccaaaatcgccgaatcgctgcgcaagatcgccgatca- atttgatgacaccattcgcgccaatg cggaagcggtgatcgccaaatatgaggggcagatggcggccatcatcgccaaatatcgcccgcggctggaggg- gcgcaaagtgctgctgtacatg gagggctgcggccgcgccacgtcatcggcgcctatgaggatctcgggatggagatcatcgccgccggctacga- gtttgcccataacgatgattac gaccgcaccctgccggacctgaaagagggcaccctgctgtttgacgatgccagcagctatgagctggaggcct- tcgtcaaagcgctgaaacctgac
ctcatcggctccgggatcaaagagaaatatatcttccagaaaatgggggtgccgttccgccagatgcactcct- gggactattccggcccctatcacgg ctatgacggcttcgccatctttgcccgccatatggatatgaccctgaacaatccggcgtggaacgaactgact- gccccgtggctgaagtctgcgtga 114 atgaagggaaaggaaattaggcgctgctggacgaacccgcctgcgagcacaaccagaagcaaaaatccgg- ctgcagcgctcctaagcccggcg caaccgccggcggctgcgccttcgacggcgcgcagataacgctcctgcccatcgccgacgtcgcgcacctggt- gcacggccccatcggctgcgc gggcagctcgtgggataaccgcggcagcgtcagcgccggcccggccctcaaccggctcggctttaccaccgat- cttaacgaacaggatgtgattat gggccgcggcgaacgccgcctgttccacgccgtccgtcacatcgtcgaccgctatcatccggcggcggtcttt- atctacaacacctgcgtaccggcg atggagggggatgacctggaggccgtctgccaggccgcacagaccgccaccggcgtcccggtcatcgccattg- acgccgccggtttctacggcag taaaaatcttggcaaccgaatggcgggcgacgtgatgctcaggcaggtgattggccagcgcgaaccggccccg- tggccagacaacacgccctttg ccccggcccagcgccacgatatcggcctgattggcgaattcaatatcgccggcgagttctggcaggtccagcc- gctgctcgacgagctggggatcc gcgtcctcggcagcctctccggcgacggccgctttgccgagatccagaccctgcaccgggcgcaggccaatat- gctggtgtgctcgcgcgcgctga tcaacgtcgcccgggggctggagctgcgctacggcacgccgtggtttgaaggcagcttctacgggatccgcgc- cacctccgacgccttgcgccagc tggcggcgctgctgggaatgacgacctgtgccgccgcaccgaggcgctgatcgcccgcgaagagcaggcggcg- gagcaggcgctggcgccg tggcgcgagcagctccgtgggcgcaaagtgttgctctacaccggcggcgtgaaatcctggtcggtggtatcag- ccctgcaggatctcggcatgacc gtggtggccaccggcacgcggaaatccaccgaggaggacaaacagggatccgtgagctgatgggcgacgaggc- ggtgatgcttgaggagggc aatgctcgcaccctgctcgacgtggtgaccgctatcaggccgacctgatgatcgccggcggacgcaatatgta- caccgcctggaaagcccggctg ccgtttctcgatatcaatcaggagcgcgagcacgcctacgccggctatcagggcatcatcaccctcgcccgcc- agactgtctgaccctcgccagtcc cgtctggccgcaaacgcatacccgcgccccgtggcgctag 115 atggcagacattatccgcagtgaaaaaccgctggcggtgagcccgattaaaaccgggcaaccgctcgggg- cgatcctcgccagcctcgggctggc ccaggccatcccgctggtccacggcgcccagggctgcagcgccttcgccaaagttttctttattcagcatttc- catgacccggtgccgctgcagtcgac ggccatggatccgaccgccacgatcatgggggccgacggcaatatcttcaccgcgctcgacaccctctgccag- cgccacagcccgcaggccatcg tgctgctcagcaccggtctggcggaagcgcagggcagcgatatcgcccgggtggtgcgccagtttcgtgaggc- gcatccgcgccataacggcgtg gcgatcctcaccgtcaataccccggatttttttggctcgatggaaaacggctacagcgcggtgatcgagagcg- tgatcgagcagtgggcgcgccga cgccgcgtccggggcagcggccccggcgggtcaacctgctggtcagccacctctgttcgccaggggatatcga- atggctgggccgctgcgtggag gcctttggcctgcagccggtgatcctgccggacctctcgcagtcaatggatggccacctcggtgaaggggatt- ttacgcccctgacccagggcggcg cctcgctgcgccagattgcccagatgggccagagtctgggcagcttcgccattggcgtgtcgctccagcgggc- ggcatcgctcctgacccaacgca gccgcggcgacgtgatcgccctgccgcatctgatgaccctcgaccattgcgatacctttatccatcagctggc- gaagatgtccggacgccgcgtacc ggcctggattgagcgccagcgcggccagctgcaggatgcgatgatcgactgccatatgtggcttcagggccag- cgcatggcgatggcggcggag ggcgacctgaggcggcgtggtgtgatttcgcccgcagccaggggatgcagcccggcccgctggtcgcccccac- cagccaccccagcctgagcc agctgccggtcgatcaggtcgtgccgggggatcttgaggatctgcagcagctgctgagccaccaacccgccga- tctgctggtggctaactctcacgc ccgctgatctggcggagcagtttgccctgccgctgatccgcgtcggttttcccctatcgaccggctcggtgag- atcgtcgcgtccgccaggcgtacgc cggtatgcgagatacgctgtttgagctggccaatctgctgcgcgaccgccatcaccacaccgccctctaccgc- tcgccgcttcgccagggcgccgac cccctgccggcttcaggagacgcttatgccgcccattaa 116 gtgccgctgatccgtctgggctttccgctgttcgaccgccatcatctgcaccgccagaccacctggggct- atgaaggcgcaatgaacatcgtcacgac gctggtgaacgccgtgctggaaaaactggaccacgacaccagttgggcaaaaccgattacagcttcgacctcg- ttcgttaa 117 atgaccctgaatatgatgctcgataacgccgcaccggaggccatcgccggcgcgctgactcaacaacatc- cggggctgttttttaccatggtggaaca ggcctcggtggccatatccctcaccgatgccagcgccaggatcatttacgccaacccagcgtttgccgccaga- ccggctattcgctggcgcaattgt aaaccagaacccgcgcctgctggccagcagccagacgccgcgcgcgatctatcaggagatgtggcataccctg- ctccagcgtcagccctggcgcg gtcagctgattaatcagcgtcgggacggcggcctgtgcctggtggagattgacatcaccccggtgcttagccc- gcaaggggaactggagcattatct ggcgatgcagcgggatatcagcgtcagctacaccctcgaacaacggctgcgcaaccatatgaccctgatggag- gcggtgctgaataatatccccgc cgccgtggtggtggtggacgagcaggatcgggtggtgatggacaacctcgcctacataaccttctgcgctgac- tgcggcggccgggagctgctca ccgagctgcaggtctcccctggccggatgacgcccggcgtggaggcgatcctgccggtagcgctgcgcggggc- cgcgcgctggctgtcggtaac ctgctggccgttgcccggcgtcagtgaagaggccagccgctactttatcgacagcgcgctggcgcggaccctg- gtggtgatcgccgactgtaccca gcagcgtcagcagcaggagcaaggacgccttgaccggctgaagcagcaaatgaccgccggcaagctgctggcg- gcgatccgcgagtcgctgga cgccgcgctgatccagctgaactgcccgattaatatgaggcggcagcccgtcggctgaacggcgagggaagcg- ggaatgtggcgctggaggcc gcctggcgtgaagggcaagaggcgatggcgcggctccagcgctgtcgcccatcgctggaactcgaaaaccccg- ccgtctggccgctgcagccctt tttcgacgatctgtgcgccctctaccgtacccgcttcgatcccgacgcgctgcaggtcgacatggcctcaccg- catctgatcggctttggccagcgcac cccgctgctgccgtgcttaagcctgtggctcgaccgcaccctggccctcgccgccgaattgccctccatgcgc- tggcgatgcagctctatgccgag gagaacgacggctggctgtcgctgtacctgactgataacgtaccgctgttgcagggcgctacgcccactcccc- cgacgcgctgaactcgccgggta aaggcatggagctgcgcctgatccagaccctggtggcgcaccatcgcggggccattgagctggcttcccgacc- gcagggcggcacctgcctgacc ctgcgtttcccgctgtttaacaccctgaccggaggtgaagcatga 118 atgatccctgaatccgacccggacaccaccgtcagacgcttcgacctctacagcagttcaccgccatgca- gcggataagcgtggtgctgagccggg ccaccgaggccagcaaaacgctgcaggaggtactcactgtattgcacaacgatgccatatgcagcacgggatg- atctgcctgtacgacagcgagca ggagatcctcagtatcgaagcgctgcagcaaaccggccagcagcccctccccggcagcacgcagatccgctat- cgccccggcgagggactggtg gggaccgtgctggcccaggggcagtcgctggtgctgccccgggtcgccgacgatcagcgttttctcgaccgcc- tgagcctctacgattacgatctgc cgtttatcgccgtaccgttgatggggcccaacgcccggccaataggggtgctggcggcccagccgatggcgcg- ccaggaagagcggctgccggc ctgcacccgttttacgaaaccgtcgccaacacgtcgcccagaccatccggctgatgatccttccggcctcacc- cgccagtcgagccgccagccgc cgaaggtggaacggccgccggcctgctcgtcgtcgcgcggcgtgggccttgacaatatggtcggcaagagccc- ggcgatgcgccagatcgtgga ggtgatccgtcaggtttcgcgctgggacaccaccgtgctggtgcgcggtgaaagcggcaccgggaaagagctg- atcgccaacgccatccatcacc attcgccacgggctggcgccgccttcgtcaaatttaactgcgcggcgctgccggacaccctgctggaaagcga- actgttcggccatgagaaaggcg cctttaccggggcggtgcgtcagcgtaaaggacgttttgagctggccgatcgcggcaccctgttcctcgatga- gattggtgaaagcagcgcctcgttc caggccaagctgctgcgtatcctccaggagggggagatggagcgggtcggcggcgatgagaccctgcgggtga- atgtccgcatcatcgccgcca ccaaccgtcacctggaggaggaggtccggctgggccatttccgcgaggatctctattatcgtctgaacgtgat- gcccatcgccctgcccccgctgcgc gagcgtcaggaggacatcgccgagctggcgcacttcctggtgcgcaaaatcggccagcatcaggggcgcacgc- tgcggatcagcgagggcgcg atccgcctgctgatggagtacagctggccgggtaacgttcgcgaactggagaactgcctcgaacgatcggcgg- tgatgtcggagagtggcctgatc gatcgcgacgtgatcctcttcactcaccaggatcgtcccgccaaagccctgcctgccaggggccagcggaaga- cagctggctggacaacagcct ggacgaacgtcagcgactgatcgccgcgctggaaaaagccggctgggtgcaggccaaggcgccacggctgctg- gggatgacgccgcgccaggt cgcttaccggatccagatcatggatatcaccctgccgcgtctgtag 119 atgatgccgctttctccgcaattacagcagcactggcagacggtcgctgaccgtctgccagcggattttc- ccattgcagaactgagcccacaggccag gtcggtcatggcgttcagcgattttgtcgaacagagtgtgatcgcccagccgggctggctgaatgagatgcgg- actcctcgccggaggcggaagag tggcggcattacgaggcctggctgcaggatcgcctgcaggccgtcactgacgaagcggggttgatgcgagagc- tgcgtctatccgccgccagatg atggtccgcatcgcagggcgcaggcgctgtcgctggtgagcgaagaagagaccctgagcagctgagcgccctg- gcggagaccctgattgtcgc cgcccgcgactggctctacgccgcctgctgtaaggagtggggaacgccatgcaatgccgagggccagccgcag- ccgctgctgatcctcgggatgg gaaagctgggcggcggcgagctgaacttctatccgatatcgatagatctttgcctggcctgagcatggcgcca- cccgcggcggccgccgcgagct ggataacgcccagttctttacccgtctggggcagcggctgatcaaggcccttgaccagccgacgcaggacggc- tttgtctatcgggttgacatgcgcc tgcggccgtttggcgacagtgggccgctggtactcagctttgcggcactggaagattattaccaggagcaggg- tcgggactgggaacgctatgcgat ggtgtaagcgcggatcatgggcgataacgacggcgtgtacgccagcgagttgcgcgcgatgctccgtcctttc- gtcttccgccgttatatcgacttca gcgtgatccagtcgctgcgtaacatgaaaggcatgatogcccgcgaagtgcggcgtcgcgggctgaaagacaa- catcaagctcggcgccggcgg gatccgtgaaattgagtttatcgttcaggtctttcagctgatccgcggtggtcgcgaacctgcactgcagcag- cgcgccctgctgccgacgctggcgg cgattgatgagctacatctgctgccggaaggcgacgcggcgctgctgcgcgaggcctatctgttcctgcgccg- gctggaaaacctgctgcaaagcat caacgatgattcagacccagaccctgccgcaggatgaacttaaccgcgccaggctggcgtgggggatgcatac- cgaagactgggagacgctgagc gcgcagctggcgagccagatggccaacgtgcggcgagtgtttaatgaactgatcggcgatgatgaggatcagt- ccccggatgagcaactggccga gtactggcgcgagctgtggcaggatgcgctggaagaagatgacgccagcccggcgctggcgcatttaaacgat- accgaccgccgtagcgtgctgg cgctgattgccgattttcgtaaagagctggatcggcgcaccatcggcccgcgcggccgccaggtgctggatca- gctgatgccgcatctgagagcga aatctgctcgcgtgccgatgcgccgagcctctggcgcggatcacgccgctgttgaccgggatcgtcacccgta- ccacctatcttgagctgctgagcg aattccccggcgcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgcg- ccacccgctgctgctggatgag ctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctgc- tgcgcgtgccggaagaggatcaa gagcagcagctggaggcgtgcgccagtttaagcaggcgcagagctgcatatcgcggcggcggatatcgctggt- accctgccggtgatgaaggt caggatcacttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggtcg- ctcgctacggtcagccgaccca cctgcacgatcgccagggtcgcggatcgccgttgtcggctacggtaagctcggcgcctgggagctggactaca- gctccgatctcgatctggtgttcc tccatgactgcccggcggaggtgatgaccgacggcgaggggagattgacggccgtcagttctacctgcggctg- gcccagcggatcatgcacctgt tcagcacccgcacctcgtccggtattactacgaagtggacgcccggctgcgtcatctggcgcggcaggttgct- ggtcaccaccgccgacgcgtt tgctgactatcagagaacgaagcctggacgtgggaacatcaggcgctggtgcgcgcccgcgtggtctatggcg- acccggcgctgcaggcgcgct ttgacgccattcgtcgcgatatcctgaccaccccgcgggaggggacgaccctgcagaccgaggttcgcaagat- gcgcgagaagatgcgcgcccac cttggcaacaaacatcccgatcgattgatatcaaagccgatgccggcgggatcaccgatattgaatttattac- tcagtatctggtcctacgctatgccagt gacaagccgaagctgacccgctggtctgacaacgtgcgtattcttgagctgctggcgcaggacgacatcatgg- acgaggaggaggcgcgcgcctta acgcatgcatacaccaccttgcgtgatgcgctccatcacctggccctgcaggagcagccgggacacgtggcgc- cagaggccttcagccgggagg tcagcaggtcagcgccagctggcagaagtggctgatggcttaa 120 atgaaaatggcaacaatgaaatcgggtctgggggcattagcccttcttccgggactggcaatggccgcgc- ccgcagtggcggacaaagccgataac gcgtttatgatgatttgcaccgcgaggttctgtttatgaccatcccggggatcgcgctgattacggcggcctg- atccgcggcaaaaacgtcctttccat gctgactcaggtgattgtgacctttggcctggtntgcgtactgtgggtgatttatggctataccctggccttc- ggaaccggcggcagcttcttcggtagctt tgactgggtgatgctgaaaaatattgaactgaaagcgctgatgggcaccttctatcagtacatccacgtggcc- ttccagggctcgttcgcctgtatcacc gtcgggctgatcgtgggggcgctggctgagcgtattcgtttctccgccgtgctgattttcgtggtggtgtgga- tgacgctacttatgttccgattgcgca catggtctggggcggcggtctgctggcgacccacggcgcgctggacttcgcgggcggcaccgttgtacacatc- aacgccgcggttgccgggctgg tgggtgcgtatatgatgggcaaacgtgtgggcttcggcaaagaagcgttcaaaccgcacaatctgccgatggt- gttcaccggaaccgccatcctctac gtgggctggttcggcttcaacgccggctccgccagcgcagcgaacgaaattgccgcactggctttcgtcaaca- ccgtcgtcgccacagcggcagcc atcctggcctggacctttggcgaatgggctctgcgcggcaaaccttcactgctgggcgcctgctccggggcga- ttgccggtctggttggcgtcacacc agcctgtgggtatatcggtgtcggtggggcgttgattgtgggtatcgcatctggtctggcgggcatctggggc- gtaacggcgctgaaacgctggctgc gggttgatgacccttgcgacgtcttcggcgtccacggcgtctgcggtcgtcggctgtatcctgaccggtatct- tcgcggccacctctctgggcggc gtgggttatgcagaaggcgtcaccatgggccatcagctgctggtgcaactcgagagtatcgcgattaccatcg- tctggtcgggcgttgtcgattcattg gctacaaagtggcggacatgaccgtggggctgcgcgtaccagaagagaggagcgcgaaggactggacgtcaac- agccatggcgaaaacgcct acaacgcctga 121 ctgaagagtttgatcctggctcagattgaacgctagcgggatgccttacacatgcaagtcgaacggcagc- acggacttcggtctggtggcgagtggc gaacgggtgagtaatgtatcggaacgtgcctagtagcgggggataactacgcgaaagcgtagctaataccgca- tacgccctacgggggaaagcag gggatcgcaagaccttgcactattagagcggccgatatcggattagatagttggtggggtaanggatcaccaa- ggcgacgatccgtagctggattgag aggacgaccagccacactgggactgagacacggcccagactcctacgggaggcagcagtggggaattttggac- aatgggggaaaccctgatcca gccatcccgcgtgtgcgatgaaggccttcgggatgtaaagcacttttggcaggaaagaaacgtcatgggntaa- taccccgtgaaactgacggtacctg cagaataagcaccggctaactacgtgccagcagccgcggtaatacgtagggtgcaagcgttaatcggaattac- tgggcgtaaagcgtgcgcaggcg gttcggaaagaaagatgtgaaatcccagagcttaactttggaactgcatttttaactaccgggctagagtgtg- tcagagggaggtggaattccgcgtgta gcagtgaaatgcgtagatatgcggaggaacaccgatggcgaaggcagcctcctgggataacactgacgctcat- gcacgaaagcgtggggagcaa acaggattagataccctgctagtccacgccctaaacgatgtcaactagctgttggggccttcgggccttagta- gcgcagataacgcgtgaagttgacc gcctggggagtacggtcgcaagattaaaactcaaaggaattgacggggacccgcacaagcggtggatgatgtg- gattaattcgatgcaacgcgaaa aaccttacctacccttgacatgtctggaattctgaagagattcggaagtgctcgcaagagaaccggaacacag- gtgctgcatggctgtcgtatgatcgt gtcgtgagatgttgggttaagtcccgcaacgagcgcaacccttgtcattagttgctacgaaagggcactctaa- tgagactgccggtgacaaaccggag gaaggtggggatgacgtcaagtcctcatggcccttatgggtagggcttcacacgtcatacaatggtcgggaca- gagggtcgccaacccgcgagggg gagccaatcccagaaacccgatcgtagtccggatcgcagtctgcaactcgactgcgtgaagtcggaatcgcta-
gtaatcgcggatcagcatgtcgcg gtgaatacgttcccgggtcttgtacacaccgcccgtcacaccatgggagtgggttttaccagaagtagttagc- ctaacc 122 ctgaagagtttgatcctcgctcagattgaacgctaggggatgccttacacatgcaagtcgaacggcagca- cggacttcggtctggtggcgagtggc gaacgggtgagtaatgtatcggaacgtgcctagtagcgggggataactacgcgaaagcgtagctaataccgca- tacgccctacgggggaaagcag gggatcgcaagaccttgcactattagagcggccgatatcggattagatagttggtggggtaaaggctcaccaa- ggcgacgatccgtagatggtttgag aggacgaccagccacactgggactgagacacggcccagactcctacgggaggcagcagtggggaattttggac- aatgggcgaaaccctgatcca gccatcccgcgtgtgcgatgaaggccttcgggttgtaaagcacttttggcaggaaagaaacgtcatgggttaa- taccccgtgaaactgacggtacctg cagaataagcaccggctaactacgtgccagcagccgcggtaatacgtagggtgcaagcgttaatcggaattac- tgggcgtaaagcgtgcgcaggcg gttcggaaagaaagatgtgaaatcccagagcttaactttggaactgcatttttaactaccgggctagagtgtg- tcagagggaggtggaattccgcgtgta gcagtgaaatgcgtagatatgcggaggaacaccgatggcgaaggcagcctcctgagataacactgacgctcat- gcacgaaagcgtggggagcaa acaggattagataccctggtagtccacgccctaaacgatgtcaactagctgttggggcatcgggccttagtag- cgcagctaacgcgtgaagttgacc gcctggggagtacggtcgcaagattaaaactcaaaggaattgacggggacccgcacaagcggtggatgatgtg- gattaattcgatgcaacgcgaaa aaccttacctacccttgacatgtctggaattcngaagagattnggaagtgctcgcaagagaaccggaacacag- gtgctgcatggctgtcgtcagctcg tgtcgtgagatgttgggttaagtcccgcaacgagcgcaacccttgtcattagttgatacgattagggcactct- aatgagactgccggtgacaaaccgga ggaaggtgggcatgacgtcaagtcctcatggcccttatgggtagggcttcacacgtcatacaatggtcgggac- agagggtcgccaacccgcgaggg ggagccaatcccagaaacccgatcgtagtccggatcgcagtctgcaactcgactgcgtgaagtcggaatcgct- agtaatcgcggatcagcatgtcgc ggtgaatacgttcccgggtcttgtacacaccgcccgtcacaccatgggagtgggttttaccagaagtagttag- cctaaccgnaaggggggcgattacc acggtaggattcatgactggggtgaagtcgtaacaaggtagccgtatcggaaggtgaggctggatcacctcct- tt 123 tacggagagtttgatcctggctcaggatgaacgctcgcggcctgcttaacacatgcaagtcgaacggttg- aacacggagcttgctctctgggatcagtg gcgaacgggtgagtaacacgtcagcaacctgcccctgactctgggataagcgctggaaacggcgtctaatact- ggatatgtgacgtggccgcatgga ctgcgtctggaaagaatttcggttggggatgggatcgcggcctatatgcttgttggtgaggtaatggctcacc- aaggcgtcgacgggtagccggcctg agagggtgaccggccacactgggactgagacacggcccagactcctacgggaggcagcagtggggaatattgc- acaatgggcgcaagcctgatg cagcaacgccgcgtgagggatgacggccttcgggagtaaacctcttttagcagggaagaaggaaagtgacggt- acctgcagaaaaagcgccgg ctaactacgtgccagcagccgcggtaatacgtagggcgcaagcgttatccggaattattgggcggaaagagct- cgtaggcggtttgtcgcgtctgctgt gaaatccggaggctcaacctccggcctgcagtgggtacgggcagactagagtgcggtaggggagattggaatt- cctggtgtagcggtggaatgcgc agatatcaggaggaacaccgatggcgaaggcagatctctgggccgtaactgacgctgaggagcgaaagggtgg- ggagcaaacaggcttagatac cctggtagtccaccccgtaaacgttgggaactagttgtggggtccattccacggattccgtgacgcagataac- gcattaagttccccgcctggggagta cggccgcaaggctaaaactcaaaggaattgacggggacccgcacaagcgacggagcatgcggattaattcgat- gcaacgcgaagaaccttaccaa ggcttgacatatacgagaacgggccagaaatggtcaactctttggacactcgtaaacaggtggtgcatggttg- tcgtcagatcgtgtcgtgagatgttg ggttaagtcccgcaacgagcgcaaccctcgttctatgttgccagcacctaatggtgggaactcatgggatact- gccggggtcaactcggaggaaggt ggggatgacgtcaaatcatcatcccccttatgtcttgggcttcacgcatgctacaatggccggtacaaagggc- tgcaataccgcgaggtggaccgaat cccaaaaagccggtcccagttcggattgaggtctgcaactcgacctcatgaagtcggagtcgctagtaatcgc- agatcagcaacgctgcggtgaata cgttcccgggtcttgtacacaccgcccgtcaagtcatgaaagtcggtaacacctgaagccggtggcctaaccc- ttgtggagggagccgtcgaaggtg ggatcggtaattaggactaagtcgtaacaaggtagccgtaccggaaggtgcggctggatcacctccttt 124 attgaagagtttgatcatggctcagattgaacgctggcggcagccctaacacatgcaagtcgaacggtag- cacagagagcttgctctcgggtgacga gtggcggacgggtgagtaatgtctgggaaactgcccgatggagggggataactactggaaacggtagctaata- ccgcataatgtcgcaagaccaaa gagggggaccttcggccctcttgccatcggatgtgcccagatgggattagattgatggtgaggtaatggatca- ccaaggcgacgatccctagatggtc tgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatatt- gcacaatgggcgcaagcctgat gcagccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaaggcgatncggt- taataaccgtgttgattgacgttac ccgcagaagaagcaccggctaactccgtgccagcagccccggtaatacggagggtgcaagagataatcggaat- tactgggcgtaaagcgcacgca ggcggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcattcgaaactggcaggcttgag- tcttgtagaggggggtagaattcca ggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactgac- gctcaggtgcgaaagcgtggg gagcaaacaggattagataccctggtagtccacgccgtaaacgatgtcgacttggaggttgtgcccttgaggc- gtggcttccggagctaacgcgttaa gtcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtgga- gcatgtggtttaattcgatgcaa cgcgaagaaccttacctggtcttgacatccacggaattnggcagagatgccttagtgccttcgggaaccgtga- gacaggtgctgcatggctgtcgtca gctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcggtccggc- cgggaactcaaaggagactgccagt gataaactggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccagggctacacacgtgctaca- atggcatatacaaagagaagcga cctcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtctgcaactcgactccatgaag- tcggaatcgctagtaatcgtggatc agaatgccacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaag- aagtaggtagcttaaccttcggga gggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttg- gatcacctcctt 125 atgaccatgcgtcaatgcgccatttatggcaaaggtgggatcggcaaatccaccaccacgcaaaacctcg- tcgccgctctcgcggaaatgggtaaaa aagtgatgatcgtcggctgcgacccgaaagggactccacccgtctgatcctgcatgcgaaagcacagaacacc- attatggagatggccgccgaag tgggttcagtggaagaccttgaactggaagatgtgctgcaaatcggttacggcggcgtgcgttgtgcagaatc- cggcggcccggagccaggcgtgg gttgtgcaggccgcggcgttattaccgccattaacttccttgaagaagaaggcgcctatgtcagcgacctcga- ctttgtcttctatgacgtcctcggtgac gtggtctgcggcgggttcgccatgccgattcgtgaaaacaaagcgcaagagatctatatcgtctgctccgggg- aaatgatggcgatgtatgccgctaa caacataccaaaggcatcgtgaaatacgctaaatccggcaaggtgcgcctgggcgggctgatttgtaactccc- gtcagaccgaccgcgaagatgaa ctgatcatcgcgctggcagaaaaactgggcacccagattgattcactttgtgccacgcgacaacatcgtccag- cgcgcggaaattcgccgtatgacgg ttatcgaatatgacccgaaatgcaaccaggccgacgaataccgcgcgctggcgaacaagatcgtcaacaacac- cctgatggtcgtcccgaccccttg caccatggatgaactggaagagctgctgatggaattcggcattatggatgtggaagacgccagcatcatcggt- aaaaccgccgccgaagaaaacgc ggcctga 126 atgaacgataacgatgtccttttctggcgcatgctgccgctatttcagtgtctgccggaactgcaacccg- cgcagatcctggcctggctgacaggagaa cgcgacgacgccttaaccccggcgtacctcgataagcttaacgtccgcgaactggaagcgaccttcccgtctg- aaacggcgatgatgtcgcccgca cgctggagccgcgttaacgcgtgccttcacggtacgctgcccgcacacctgcaggtaaaaagcaccactcgtc- aggggcaattacgggtagcattt gttcacaggatggattgctgatcaatggtcattttggtcaggggcggctgttttttatctacgcctttgatga- acagggcggatggctacacgcgttacgc cgtcttccctcggccccgcaaacccaggagccgaatgaagttcgcgcgcagctcctgagtgattgccacctgc- tgttttgtgaagccattggcggccc tgcggcggcccggctgattcgtcacaatatccacccgatgaaagtgtcgccagggatgtccattgccgcccag- tgtgatgccattaccgcactgctga gcggacgtctgccaccgtggctggcaaaacgtcttgagaaagccaacccgctggagaagggtgttttaa 127 atgaagggaaatgacattctcgcgctgctggatgaacccgcctgcgaacacaatcacaaacagaaatccg- gctgtagcgcccctaaacccggtgcc acggcgggcggttgcgcgttcgacggcgcgcaaatcaccctgttgccgctgtcggtagtggcgcacctggtcc- acggaccgattggctgcacggg aagctcctgggataaccggggcagtatgagctccggccccagtctcaaccggctcggctttaccaccgacctg- aacgagcaggatgtcaaatgggg cgcggcgaacggcggcttttccacgcggtgcgtcatatcgtcaaccgttatcaccctgccgccgtgtttatct- ataacacctgcgttccggcgatggag ggtgatgatattgacgccgtctgtcaggcggcggaaaccgccaccggcgtgccagtgattgccgttcatgccg- ccgggttctatggcagcaaaaacc ttggcaaccgtctcgcgggtgaagtgatggttaacaaggtcattggacggcgcccgcccgccccctggccgga- cgatacccccttcgcgccggaac accgccacgatatcggcctgattggcgaatttaatatcgccggggagttctggcacgttcagccgctgctcga- tgagctgggtattcgcgtgctgggca gcctttccggggatggccgttttagtgaaatccagaccctgcaccacgcgcaggtcaatatgctggtctgctc- aagagcgctgatcaatgttgcccgca ccctggaacagcgctatggcaccccctggtttgagggcagtttttacggcgtgcgcgctacctccgatgccct- gcgtcaactggcatccctgcttggcg acagcgatctgattgcccccaccgaagccgttattgcccgcgaagaagccacggcaaatcagccgctcgcccc- gtggcgcgaacggctacagggt cgcaaagtgctgctctataccggtggcgtgaaatcaggtcggtgatctccgcattgcaggatttagggatgac- cctcgtgccgactggcacccgcaa atctaccgaagaagataagcagcgtattcgcgaattaatgggcgatgacgcgctaatgctggaagaaggcaac- gcccgcaccctgctggatgtggtg taccgctatcaggcggatttgatgatcgctggggggcgtaacatgtataccgcgtacaaagcgggctgccgtt- tctggatatcaaccaggagcgtga acacgcctttgcgggttatcgcggcatcgtcaccctcgcccaacagctttgccagactattgaaagccccgtc- tggccgcaaacacacgcccgcgcg ccgtgccaataa 128 atgagcaatgcaacaggcgaacgtaatctggaaattatccaggaagtgctggagatctttcccgaaaaaa- cgcgcaaagaacgcagaaagcacatg atggttaccgacccggagatggaaagcgtcgggaaatgcatcatctctaaccgcaaatcgcagccgggtgtga- tgactgtccgcggctgacctacg ccgggtcgaaaggcgtggtttttgggccgattaaagatatggcccacatctcccacggcccgatcggctgtgg- gcagtactcccgtgccgggcggc gcaactactacaccggggtcagcggcgttgattccttcgggacgctgaactttacctctgattttcaggagcg- cgatatcgtcttcggcggcgataaaaa gctcaccaaactgattgaggagatggaggaactgttcccgctgaccaaaggcatctccattcagtcggagtgc- ccggtaggtttaatcggtgacgatat cgaagcggtggcgaatgccagtaaaaaagcgctcaacaagccggtgatcccggtgcgttgcgaaggctttcgc- ggcgtgtcgcagtcgctcggtca ccatatcgccaacgacgttatccgcgactgggtgctggataaccgcgaagggaagcccttcgaatctaccccc- tatgacgtggccatcatcggcgatt acaacatcgggggggatgcctgggcgtcgcgcattctgcttgaagagatggggttacgcgtggtggcgcagtg- gtccggtgacggcacgctggtag agatggtaaacaccccgttcgtcaagctgaacaggtgcactgctaccgctctatgaactacatctctcgccat- atggaagagaaacacggtatcccgt ggatggagtacaacttcttcggcccgaccaattatcgccgaatcgagcgtaagatcgccgatcaatttgacga- caccatccgcgccaatgcggaagc ggtgatcgccaaatatcaggcgcaaattcgatgcgattatcgccaaataccgcccgcgtctcgaaggccgcaa- ggtgctgctctatatgggtggcctg cgtcctcgccacgtgattggcgcgtatgaggatttgggcatggagattgtcgccgccgggtatgaatttgccc- ataacgacgattacgaccgcaccct gccggacctcaaagagggcacgctgttgttcgacgatgccagcagttatgaactggaagccttcgtgaaggcg- attaagccggacctcattggctca ggcatcaaggaaaaatacattttccagaaaatgggggtaccgtttcgccagatgcactcctgggattactccg- gcccgtatcacggctatgacggcttt gccatctttgcccgcaatatggacatgacgctcaacaatcccgcctggggcgagttgaccgcaccctggctga- aatcagcctga 129 atggcagatatcatccgtaatcagaaaccgctggcggtaagcccggtaaaaagcggccagccgttaggcg- ccattctggcgagcctcggctttgtgc acagattccactggtgcacggtgcgcagggatgcagcgcgttcgccaaagtgttttttatccaacattttcat- gaccctattccgctgcaatccacggcg atggaccccacctcaacggtcatgggggcggacggcaatatccttgccgcgctcaatacgctgtgccagcgca- ttcaccccgaaagctatcgtcctgt tgagtaccggcctgtctgaggcgcagggcagcgatatcagccgcgtggtacgtcagtttcgtgaggattttcc- ccgccacaaaaatatcgccctcctg acggtcaacaccccggatttttacggcacgctggagaacggctttagtgcggtggtggaaagcgtcatcgaac- agtgggtgccggaaagcctcag catggcctgcgtaaccggcgggtcaacttgttgttaagtcacctgctgacgcccggtgatgttgagttgctgc- gcagctacgtcgaggcttttggcctgc aaccggtgatcgtgccggatctttcacagtcgctggatggtcacctggcaagcggtgatttttcgccggtcac- tcaggggggaacgcccctgcgcatt atcgaacagatgggacagagcctgtgcacgtttgctattggcgtgtcgctgtcccgtgcggcatcgagctggc- acagcgtagccgtggcgangtga tcgtgcttccccatctgatgaccatggaacattgcgaccgttttattcatcaactgaagatcatttccgggcg- cgaggttcccgcctggattgagcgccag cgcggacaattgcaggatgcgatgatcgattgtcatatgtggttgcaggatacccggctcgcgctggccgccg- agggcgatctgctggcgggctggt gtgatttcgcccgtagccagggcatgctccccggccccgttgtggcgccggtcagccagccgggcctgcgaca- gcttcccgtggagaaagtggtca ttggcgatctggaagatatgcaggatttactctgcgctatacctgctgacctgctggtcgccaactcccatgc- cgcagacctggccgaacaattctccat cccgctgatccgcgccgggttccctatcttcgacaggcttggcgaatttcgtcgcgtgcgtcagggatacccc- ggcattcgcgacacgctgtttgagct ggcgaacctgatgcgcgaacgtcatcaccacctgcccgtctaccgctcccocctgcgccagcaatttgcccag- gacgctgacggaggccgctatgc aacatgttaa 130 atgagccaaactgctgagaaaattgtcacctgtcatccgctgtttgaacaggacgaataccagacgctgt- ttcgcaataagcgcggtctggaagaggc gcacgacccgcagcgcgtgaagaggtttttgaatggaccaccacggcggagtatgaagcgctgaactttaagc- gtgaagcgttaaccgtcgatccg gcaaaggcctgccagcctttaggatcggtactctgctcgctgggttttgccaatacgctgccttagtgcacgg- ttcccagggctgtgtggcctatttccg cacctattttaaccgtcatttcaaagagccgatcgcttgcgttccgactctatgacagaggatgcggtcttcg- gcgccggcaacaacaacctaacacc gggttgcaaaatgccagcgccctgacaaaccggaaattgtcgctgtgctccactacctgtatggcggaggtca- tcggcgatgacctgcaggcctttatc gccaacgccaaaaaggacgggtttattgatgccgccattccggtgccctacgcccatacgccaagttttatcg- gtagccacatcaccggctgggacaa catgtttgaaggtttcgcccgggcatttaccgccgatcacgtggcgcaaccgggcaaactggcgaagctaaac- ctggtgaccggttttgaaacctatct tggcaattaccgcgtgctcaaacgcatgatggcccagatggaggtgccctgtagcctgctgtctgacccgtct- gagggttagatacgccagccgacg gccactatcgcatgtatgcgggcggcacaacgcaacaagagatgcgcgacgcccccgatgctatcgacaccca- gctgctgctgcaaccctggcatctgg tgaagagtaaaaaagtggtgcaggagtcctggggccagcccgccacagaagtgtccatcccaatgggactgac- cgggaccgacgaactgctgatg gcagtcagtcagttaaccggcaaaccggtggccgatgaactgacgctggagcgtgggcgcctggtggatatga- ttctcgattcacacacctggctgc acggtaagaaattcggtctctacggcgatccggatttgtgatggggctgacgcgtttcctgctggaactgggc- tgcgagccgacggttatcctctgtca
taacggtagcaagcgctggcagaaagcgatgaagaaaatgcttgaggcatcgccctacggtcaggagagcgaa- gtgttcatcaactgcgatctgtg gcatttccgctcgctgatgtttacccgcaaaccggactttatgatcggcaactcgtacgccaaattcatccag- cgtgacacgctggcgaaaggcgaaca gtttgaagttccgctgatccgtcttggcttcccgttgttcgaccgccaccacctgcatcgccagaccacatgg- ggttatgaaggggcgatgaatatcgtc accaccctggtcaacgccgtgctggaaaaagtcgaccgcgataccatcaaactgggcaaaacggactacagct- tccgaccttgtccgctaa 131 atgacctttaatatgatgctggagaccagcgcaccgcagcacattgcgggcaacctctcacttcaacatc- ccggactgttttccacgatggttgaacag gctccgatcgcgatttcgctgaccgacccggacgcgaggattctgtacgctaatccggccttttgtcgccaga- ccggttatagcctggaagagctgctc aaccagaaccatcgcatactggcaagccancagacgccgcgcagcatttatcaggaactgtggcaaacgctgc- tgcaacagatgccctggcgcggt cagctcatcaatcgccgtcgggatggcagcctttatctggctgaggtcgatatcaccccggtcgtcaacaaac- agggcgaactggaacactacctcgc catgcaacgtgatatcagcgccagctatgcgctcgaacagcgattgcgcaatcacaccaccatgagcgaggcg- gtgctgaacaacattcctgccgcc gtggtggtggtcaacgagcaggaccaggtagtcatggacaacctcgcctacaaaaccttctgtgccgactgcg- gtggcaaggagctgctcaccgaa ctggatttctcccggcgcaaaagcgatctctatgccgggcaaatactgcctgtggtgctgcgcggcgccgtgc- gctggactctgtcacctgaggac cttgccgggggtgagcgaagaagccagccgctactttattgataccgcgagccccgcaccctggtggtgatca- ccgactgcacccagcaacaaca acaggcccgaacagggccgtctcgatcgtctcaaacaggagatgaccaccgggaagctgctggccgcgatccg- tgaatcgttggatgccgcgctggt tcagctaaactgccccatcaatatgctggcggcggcgcgacgtctcaacggtgaagataaccataacgtggcg- ctggatgccgcgtggcgcgaggg ggaagaggcgctggcccgcctgcaacgctgccgcccttctctcgatctggaagagagcgcgctgtggcctctg- caaccgctgtttgacgacctgcg cgccctttaccatacccgctataacaatggcgaaaatctgcacgttgaaatggcctctccgcatctggcgggg- tttggtcagcgcacgcagatccttgc ctgctcagtttgtggctcgaccgtacgctggccctcgccgccgcgctaccggacagaacgctgcatacccagc- tttacgcccgtgaagaagatggct ggctgtgtccatttggctgacagataatggccgctcatccatgtgcgatacgcccactcccccgatgccctga- acgcccccggcaaagggatggagct gcgattgattcaaaccctggttgcccatcatcgcggcgcaatagaactaactacccgccagatggcggtacct- gcctgaccctgcgattcccgttattt cattcactgacccgaggcccacgatga 132 atgacccagcgacccgagtcgggcaccaccgtctggcgttttgatctctcacagcaatttaccgccatgc- agcgcatcagcgtggtgttgagtcgcgc aaccgagataagccagacgctgcaggaggtgctgtgtgttctgcataatgacgcatttatgcaacacggcatg- ctgtgtctgtatgacaaccagcagg aaattctgagtattgaagccttgcaggaggcagaccaacatctgatccccggcagctcgcaaattcgctatcg- ccctggcgaagggctggtaggagc cgtactgtcccagggacaatctcttgtgctgccgcgtgtcgccgacgatcaacgctttctcgacaggcttggc- atctatgattacacctgccgtttatcg ccgtccccttaatggggccaggcgcgcagacgattggcgtgctcgccgcgcagccgatggcgcgtctggagga- gcggcttccttctgtacgcgct ttctggaaaccgtcgccaatctggtcgcacagacagtccggctgatgaccccgcctgccgccgccacaccgcg- cgccgcgattgcccagaccgaa cgccagcgcaactgtggcactcctcgccccttcggctttgagaatatggtgggcaaaagcccggccatgcagc- agacaatggacattatccgccagg tttcgcgctgggataccacggtactggtgcgcggcgaaagcggcaccggtaaagaacttatcgccaatgctat- tcatcacaactcccctcgcgccgcc gcgccctttgtgaaatttaactgcgcggcgctaccggatacgctggagagagcgaattgttcggccatgaaaa- aggggcgttcaccggcgcggttc gccagcgtaaaggacgtttgaactggccgatggcggcacactgtttcttgatgaaattggcgaaagcagcgcc- tcgttccaggccaaactgctgcgt attttgcaggagggtgaaatggagcgcgttggcggcgacgaaaccctgcgcgtcaatgtgcgtatcatcgccg- ccaccaaccggaatctggaagaa gaggtgcggatgggcaatttccgcgaggatctctattatcgcctcaacgtaatgcccatctccctgcccccgc- tgcgtgaacgtcaggaggacattgc cgagctggcgcactttctggtgcgcaaaatcgcccataaccaggggcgtacgctgcgcatcagtgatggcgcc- atccgtctgctgatgggttacaact ggcccggtaacgtgctgtgagctggaaaattgcctggaacgttcggcagtgatgtcagaaaacggcctgatcg- accgcgatgtggtgctctttaaccac cgtgagaacacgccaaaactcgctatcgccgccgcgccaaaagaggatagctggcttgatcaaacgctggatg- aacgtcaacggctgattgccgcg ctggaaaaagccgggtgggtgcaggccaaagcggcgcgtctgctgggtatgacgccccgtcaggtcgcctatc- ggatacaaattatggatatcagc atgcccaggatgtga 133 atgatgccgcactctccacagctacagcagcactggcaaactgtactggcccgcttgcctgagtcattca- gtgaaacaccgcttagtgaacaagcgca gttagtgcttactttcagtgattttgtgcaggatagccttgccgcgcatcctgactggctggctgagctggaa- agcgcaccgccacaggcggacgagtg gaagcagtatgcgcaaacccttcgcgaatcgctggaaaggtgtgggagatgaggcatcattaatgcgtgcgct- gcgcctgttccgtcgccatatgatgg tgcgcattgcctgggcgcagtcgctggcgctggtggcagaagatgagacgttgcagcagttgagcgtactggc- ggagaccctgatcgtcgctgcac gcgactggctttacgatgcctgctgtcgcgagtggggaacgccgtgcaatcagcagggggaaccgcagccgtt- gctgatcctgggcatgggcaag ctgggtggcggggagcttaacttttcgtccgatatcgatctgatttttgcctggccggaaaacggttcaacgc- gcggtgggcgacgcgaacttgataac gcccagttttttactcgcttgggacagcgcctgatcaaagtgctcgaccagccgacgcaggatggctttgtct- atcgcgtggatatgcggctgcgcccg tttggcgacagcggtccgctggtgctgagttttgccgcgctggaagattattatcaggagcaggggcgcgact- gggaacgttatgcgatggtgaaagc ccgcattatgggcgataaggacgatgtttacgctggcgaattacgggccatgctgcggccgttcgtcttccgt- cgctatatcgatttcagcgttattcagt ctctgcgtaacatgaaagggatgattgcccgcgaagtgcgccgccgtggtctgaaagataacattaagctggg- cgcgggcggcatccgtgagattga gtttatcgttcaggtgttccagttgatacgcggtgggcgcgagccgtcgttgcagtcccgttcactgttaccg- acgctggacgctatcgataagctgggt ttgctgccgcctggcgatgcaccggcgttacgccaggcctatttgtatctgcgccgtctggaaaaacctgctg- caaagcattaacgacgaacaaacgca gacgctgccgacagatgaactcaatcgcgcgcgtctggcctgggggatgcgggtcgcagactgggaaaccctg- accgctgagcttgaaaagcaga tgtctgccgtacgagggatattcaacaccctgattggcgatgacgaagccgaagagcagggggatgcgctctg- cgggcaatggagtgagttgtggc aggatgcgtttcaggaagatgacagcacgcctgtgctggcgcacctttctgacgatgatcgccgccgcgtggt- cgcgatgattgctgattttcgcaaag agctggataaacgcaccattggcccacgcggccgccaggtgctcgaccatctgatgccgcatctgttgagtga- tgtctgctcccgtgaggatgcccct gtaccgttgtctcgcgtgacgccgctgttaacgggaattgtcacgcgtacgacgtatcttgagctgctcagcg- agtttcctggtgcgcgtaagcatctga tttcactctgtgccgcctcgccgatggtggccagtaagctggcgcgctatccgttattgctggatgagttgct- cgatccgaataccctttatcagcccacg gcgatgaatgcctaccgggatgagctacgtcagtatctgctgcgtgtgccggatgacgatgaagagcagcaac- tggaggcgttacgccagtttaaac aggctcaattgttgcgtgtggcggcagcagatctggcaggcacactccccgtgatgaaagtgagcgatcactt- aacatggcttgccgaagccatcatt gaagccgtggtacaacaggcgtggagcctgatggtatcgcgttatgggcagccgaaacacttacgcgaccgtg- aaggccgtgggtttgcagtggtc ggttacggcaaactgggcggttgggagctgggctatagttccgatctggatttgattttccttcatgactgtc- cggtggacgtgatgactgacggcgagc gggaaatcgatggccgccaattttatctgcgccttgcccagcgcgtgatgcacctgttcagtacgcgcacctc- atccgggatcctgtatgaggtagacg cgcgcttgcgcccgtccggtgcggcgggaatgctggtgacctcaaccgaatcctttgccgactaccagcgcac- cgaagcctggacctgggaacatc aggcgctggttcgcgcccgcgttgtctatggcgatccacaattaaacgcgcaatttgatgccatccgccgcga- tatcaccatgaccgtgcgtaatggtg caacgttacaaaccgaggtgcgcgagatgcgcgaaaaaatgcgcgcccacttgagcaataagcacaaggatcg- ctttgatattaaagccgatgagg gtggaattaccgatatcgaatttatcacccagtatctggtgctgcgttatgcccatgccaaaccgaaactgac- gcgctggtcggacaatgtccgcattct ggaagggctggcgcaaaacggcattatggaagagcaggaagcgcaggcacttaccaccgcctatacaacgttg- cgtgatgagctgcatcacctgg cgctacaggagctgccaggacatgttccggaggcatgttttgtcgctgaacgcgcgatggtgcgagcctgctg- gaacaagtggttggtggagccgtg cgaggacgcgtaa 134 atgaagaaagcactattaaaagcgggtctggcctcgctggcattactgccgtgtctggctatggcagccg- atccggttgtcgtcgataaagccgacaat gcctttatgatgatttgcaccgcgctggtgctgtttatgtcaattccgggcatcgccctgttctatggtggtt- taatccgcggtaaaaacgtcctttctatgct gacacaggttgcggttacgttcgcactggtgtgcgtgctgtgggtggtttacggctactctctggcctttggc- actggcggcagcttcttcggtagcttcg actgggtgatgctgaaaaatattgagctgaaagcgctgatgggcaccatctatcagtacattcacgttgcgtt- ccagggctcgtttgcctgtattaccgtc ggcctgattgtcggtgcgctggcagaacgtatccgtttctccgcagtactgattttcgtcgtggtatggctga- cgctgtcctacgtgccgatcgcacacat ggtctggggcggcggtctgctggcaacccatggcgccatggattttgcgggcggtacagtcgttcacatcaac- gcagccgttgcaggcctggtgggt gcttacctgattggcaaacgtgtcggtttcggtaaagaagcgtttaaaccgcacaacctgccgatggtgttta- ccggtacggcaatcctctactttggctg gttcggattcaacgcgggttctgcaagcgcggcgaacgaaattgcgggtctggcttttgttaacaccgtcgtg- gcaacagcgggtgcaatcctctcctg ggtcttcggtgagtgggcgctgcgcggcaaaccgtctctgttgggtgcctgttctggtgcgattgctggcctc- gtgggtatcaccccggcgtgtggtta cgttggtgtgggtggcgcgctgatcgtgggcatcgttgcaggcctggcgggtctgtggggcgttaccgcgctg- aaacgctggctgcgtgttgacgac ccgtgtgatgtcttcggtgttcacggcgtgtgcggtatcgtaggttgtatcatgacaggtatcttcgcagcca- cttcactgggcggcgtgggttatgccga aggcgtgaccatgggccatcaggttctggtacaactggaaagtatcgccattactatcgtatggtctggtatc- gtcgcctttatcggttacaaactggctg atatgacagtgggtctgcgtgttccggaagatcaggaacgcgaagggctggacgtcaacagccacggcgagaa- cgcctacaacgcctga 135 ctggggtcactggagcgctttatcggcatcctgaccgaagaatttgccggtttcttcccgacctggctgg- cccctgttcaggttgtggtgatgaatatcac tgattctcaagctgaatatgtcaacgaattgacccgtaaattgcaaaatgcgggcattcgtgtaaaagcggac- ttgagaaacgagaagattggctttaaa atccgcgagcacactttacgtcgtgtcccttatatgttggtctgtggtgataaagaggtggaagcaggcaaag- tggccgttcgcacccgccgcggtaa agacctgggcagcctggacgtaagtgaagtgattgagaagctgcaacaagagattcgcagccgcagtcttcaa- caactggaggaataaggtattaaa ggcggaaaacgagttcaaacggcacgtccgaatcgtatcaatggcgagattcgcgcccaggaagacgcttaac- tggtctggaaggtgagcagctg ggtatt 136 attgaagagtttcatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgaacggtag- cacagagagcttgctctcgggtgacga gtggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaata- ccgcataacgtcgcaagaccaaa gagggggaccttcgggcctcttgccatcagatgtgcccagatgggattagctagtaggtgggctaacggctca- cctaggcgacgatccctagctggt ctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatat- tgcacaatgggcgcaagcctga tgcagccatgccgcgtgtatgaagaaggccttcgggttgtaaagtactttcagcggggaggaaggcganacgg- ttaataaccgtgttgattgacgtta cccgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgaagcgttaatcggaat- tactgggcgtaaagcgcacgc aggcggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcatccgaaactggcaggcttga- gtacgtagagggaggtagaattc caggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggcctcctggacgaagactg- acgctcaggtgcgaaagcgtg gggagcaaacaggattagataccctggtagtccacgccgtaaacgatgtctatttggaggttgtgcccttgag- gcgtggcttccggagctaacgcgtta aatagaccgcctggggagtacggccgcaggttaaaactcaaatgaattgacgggggcccgcacaaccggtgga- gcatgtggtttaattcgatgca acgcgaagaaccttacctggtcttgacatccacagaacttgccagagatggcttggtgccttcgggaactgtg- agacaggtgctgcatggctgtcgtca gctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcggtccggc- cgggaactcaaaggagactgccagt gataaactggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccagggctacacacgtgctaca- atggcgcatacaaagaagcga cctcgcgagagcaagcggacctcataaagtgcgtcgtagtcccgattggagtctgcaactcgactccatgaag- tcggaatcgctagtaatcgtggatc agaatgccacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaag- aagtaggtagcttaaccttcggga gggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttg- gatcacctcctt 137 atgaccatgcgtcaatgcgccatttacggcaaaggtgggatcggtaaatcgaccaccacacagaacctgg- tcgccgcgctggcggagatgggtaag aaagtcatgatcgtcggctgcgatccgaaagccgactccacgcgtttgatcctgcatgcgaaagcgcagaaca- ccattatggagatggccgccgaag tcggctccgtcgaagacctggaattagaagacgtgctgcaaatcggttacggcggcgtgcgctgcgcggaatc- cggtggcccggagccaggtgtg ggttgtgccggtcgtggcgtgatcaccgcgattaacttcctcgaagaagaaggcgcttacgtgccggatctgg- attttgttttctacgacgtgctgggcg acgtggtatgcggtggtttcgccatgccgattcgtgaaaacaaagcgcaggagatctacatcgtttgctctgg- cgaaatgatggcgatgtacgccgcc aataacatctccaaaggcatcgtgaaatatgccaaatccggtaaagtgcgcctcggcgggctgatttgtaact- cgcgccagaccgaccgcgaagatg aactcatcattgcgctggcggaaaaactcggcacgcaaatgatccactttgttccccgcgacaacattgtgca- gcgtgcggaaatccgccgtatgacg gttatcgaatatgacccgacctgcaatcaggccaacgaatatcgcagccttgccagcaaaatcgtcaacaaca- ccaaaatggtggtaccaaccccctg caccatggatgaactggaagaactgctgatggagttcggcattatggatgtggaagacgccagcatcattggt- aaaaccgccgccgaagaaaacgc cgtctga 138 atgagcaatgcaacaggcgaacgtaacctggaaatcatcgagcaggtgctggaggttttcccggaaaaga- cgcgcaaagagcgcagaaaacacat gatggtgacggacccggagcaggagagcgtcggcaagtgcatcatctctaaccgcaaatcgcagccgggcgtg- atgaccgtgcgtggctgctcgt atgccggatcaaaaggggtggtatttgggccaatcaaagatatggcgcatatctcccacggcccgatcggctg- cgggcagtactcccgcgccgggc ggcgtaactactataccggcgtcagcggcgtggacagtttcggcacgctcaacttcacctccgatttccagga- gcgcgacatcgtgtttggcggcgac aaaaagctcgccaaactgattgaagagctggaagaactgtttccgctgaccaaaggcatttcgattcagtcgg- aatgcccggtcggcctgattggcga tgatattgaagccgtggcgaacgccagccgcaaagcgatcaacaaaccggttattccggtgcgttgcgaaggc- tttcgcggcgtgtcgcaatccctc ggtcaccatattgccaacgatgtgatccgcgactgggtactggataaccgcgaaggcaaaccgtttgaatcca- ccccttacgatgtggcgatcatcgg cgattacaacatcggtggcgacgcctgggcctcgcgcattttgctcgaagagatggggttgcgggtggtcgcg- cagtggtccggcgacatacgct ggtgcagatggaaaacacgccgttcgtcaaactgaacctggtgcactgctaccgctcgatgaactacatctcg- cgccatatggaggagaagcacggt attccgtggatggaatacaacttctttggcccgacgaaaatcgcggaatcgctgcgcaaaatcgccgacctgt- tcgacgacaccattcgcgccaacgc cgaagcggtgatcgcccgataccaggcgcagaacgacgccattatcgccaaatatcgcccacgtctggagggt- cgcaaagtgttgctctatatgggc gggctgcgtccgcgccatgtgattggcgcctatgaagatctgggaatggagatcatcgccgccggttatgagt- ttggtcataacgacgattacgaccg
caccctgccggatctgaaagagggcacgctgctgtttgatgacgccagcagctatgagctggaggcgtttgtc- aacgcgctgaaaccggatctcatc ggttccggcatcaaagagaagtacatctttcagaaaatgggcgtgccgtttcgccagatgcactcctgggatt- actccggcccgtaccacggctatgac ggcttcgccatcttcgcccgcgatatggatatgacgctcaacaaccccgcctggggtcagttgaccgcgccgt- ggcttaaatccgcctga 139 atgaaggggaacgacatcctggctctgctcgatgaaccagcctgcgagcataaccataaacagaaaaccg- gctgtagcgcgccaaacccggcgc caccgccggaggctgcgccttcgacggcgcacagatcaccctgctgccactttccgatgtggcgcatctggta- catggcccgattggctgcgccggc agctcatgggataaccgtggcagcctgagttctggcccgctgattaaccgactcggattcaccactgatttga- acgaacaggatgtcatcatggggcg cggcgagcggcggttgtttcacgcggtgcgccatattgtcgagcgctatcacccggcggcggtatttatttac- aacacctgcgttccggctatggaagg cgatgacattgacgcggtctgccaggccgccgcgaccgccaccggtgtgcccgtgattgccgtagatgtggcc- ggtttttacggtagcaaaaacctg ggtaaccgcctcgcgggcgaggtgatggtgaaaaaagttatcggcgggcgcgaacccgcgccgtggccggaca- atacaccttttgccccggcgca ccgccatgacataggcctgattggcgaatttaacatcgccggcgagttctggcatatccagccgctgcttgat- gagctgggtattcgcgtccttggctcc ctttccggcgacgggcgctttgccgagatccagacgttgcaccgcgcgcaggtcaatatgctggtgtgctcca- gggcgctgattaatgtcgccagatc gcttgaacaacgttatggcacaccctggtttgaaggcagtttttatggcgttcgcgccacctccgatgccctg- cgccagctggcaacactcaccggcga tagcgatttaatggcgcgaaccgaacggctgatcgcacgtgaagagcaagccacagaacaggcgctagcaccg- ctgcgtgaacggttacacggcc ggaaagtgctgctctataccggtggcgtgaaatcctggtcggtggtttcggcgctgcaggatctcggcatgac- ggtcgttgctaccggaacgcgcaaa tccaccgaagaggataaacaacgcatccgtgaactgatgggcgatgacgccatcatgctggatgaaggcaatg- cccgcgccttgctggatgtggtct atcgctacaaagccgacatgatgatcgcgggcgggcgcaacatgtacaccgcctataaagcgcgtctgccctt- tctggatatcaaccaggagcgtga acacgcgtttgccggttatcgcggcatcatcacgcttgccgaacaactttgtcagacgctggaaagcccggtc- tggccgcaaacacatgcccgcgcc ccgtggcaataa 140 atgagccagactgctgagaaaatacagaattgccatcccctgtttgaacaggacgcctaccagacactat- ttgccggtaaacgggcactcgaagagg ctcactcgccggagcgggtgcaggaagtgtttcaatggaccaccaccccggaatacgaagcgctgaacttcaa- acgcgaagcgctgactatcgacc cggcaaaagcctgccagccgctgggggcggtgctagttcgctggggtttgccaacaccctgccgtatgtgcac- ggttcacagggttgtgtggcctat ttccgtacgtactttaaccgccacttcaaagaaccggtggcctgcgtgtcggattcgatgacggaagacgcgg- ccgtgttcggcgggaataacaacct caacaccgattacaaaacgccagcgcactgtataaaccggtgattatcgccgtctctaccacctgtatggcgg- aagtgatcggtgatgatttacagg cgtttatcgccaacgccaaaaaagatggttttctcgatgccgccatccccgtgccctacgcccacacccccag- ttttatcggtagccatatcaccggctg ggacaacatgtttgaaggttttgcccgtacctttaccgcaaaccatcagccacagcccggtaaactttcacgc- ctgaacctggtgaccgggtttgaaac ctatctcggcaatttccgcgtgagaaacgcatgatggaacaaatggaggtgcaggcgagtgtgctctccgatc- cgtcggaggtgctggacaccccc gccaatggccattaccagatgtacgcgggcggtacgacgcagcaagagatgcgcgaggcaccggatgccatcg- acaccctgctgctgcaaccgtg gcagctggtgaaaagcaaaaaagtggtgcaggagatgtggaatcagcccgccaccgaggttgccattcccgtc- gggctggcaggcacagacgaa ctgttgatggcgattagccagttaaccggcaaagccattcccgattcgctggcgctggagcgcgggcggctgg- tcgatatgatgctcgactcccacac ctggttacacggtaaaaaattcggtctgtttggcgatccggattttgtcatgggattgacccgcttcctgctg- gaactgggctgtgaacctgccgtcatcct ctgccataacggtaacaaacgctggcaaaaagcgatgaagaaaatgctcgatgcttcaccgtacggccaggag- agcgaagtgtttatcaactgcgac ttgtggcatttccgctcgctgatgttcacccgccagccggattttatgattggcaactcgtacgccaagttta- ttcagcgcgacaccttagccaagggcga acagtttgaagtcccgctgatccgcctcggttttccgctgttcgaccgtcaccatctgcaccgccagaccacc- tggggctacgagggcgcgatgagca ttctcacgacgctggtgaatgcggtactggagaaagtggacaaagagaccatcaagctcggcaaaaccgacta- cagcttcgatcttatccgttaa 141 atggctgatattgttcgtagtaaaaaaccgctggcggtgagcccgataaaaagcggccagccgctggggg- cgatcctggcaagcctgggtttcgaac agtgcataccgctggtacacggcgctcaggggtgcagcgcgttcgcgaaagtgttctttattcaacattttca- cgacccgatcccgctgcaatcgacgg cgatggacccgacttccaccattatgggcgccgatgaaaacatttttaccgcgctcaatgttctctgccagcg- caacgccgcgaaagccatcgtgctgc tcagcaccgggctgtcagaagcccagggcagcgatatttcacgagtggtgcgccagtttcgtgatgactttcc- gcggcataaaaacgtggcgctgctc accgtcaacaccccggatttctacggctcgctggaaaacggctacagcgccgtgctggaaagcatgattgaac- agtgggtgcccgcgcagcccgcc gccagcctgcgcaaccgtcgcgtcaacctgctggtcagccatttactgacgccgggcgatatcgaactgttac- gcagttatgtggaagcattcggtctg caaccggtgattgtgccggatctatcgcagtcgctggacggacatctggccaacggtgatttttcgcccgtca- cccaggggggaacaccgctgcgca tgattgaacagatgggccaaaacctggccacttttgtgattggccactcgctggggcgggcggcggcgttact- ggcgcagcgcagccgtggcgagg tgatcgccctgccgcatctgatgacgcttgatgcgtgcgacacctttatccatcgcctgaaaaccctctccgg- gcgcgacgtgcccgcgtggattgag cgccagcgcgggcaagtgcaggatgcgatgatcgattgccatatgtggttgcagggcgcggctatcgccatgg- ccgcagaaggcgatcacctggc ggcatggtgcgatttcgcccgcagccagggcatgatccccggcccggttgtcgcgccggtcagccagccgggg- ttgcaaaatctgccggttgaaat ggtggttcatcggcgatctggaagatatgcaggcttcggctttgcgcgacgcccgccgcgttactggtggcca- attctcatgccgccgatctcgccacgc agtttgatatgtcgcttatccgcgccggttttccggtgtatgaccggctgggggaatttcgtcggctgcgcca- ggggtatagcggcattcgtgacacgc tgtttgagctggcgaatgtgatgcgcgaacgccattgcccgcttgcaacctaccgctcgccgctgcgtcagcg- cttcggcgacaacgttacgccagg agatcggtatgccgcatgttaa 142 atgaccctgaatatgatgatggatgccagcgcgcccgaggccatcgccggtgcgctttcgcaacaacatc- ctgggctgttttttaccatcgttgaagaa gccccgtcgctatttcactaaccgatgccgaggcacgtattgtctatgccaacccggcattctgccgccagac- cggctatgagcttgaggagttgttg cagcaaaatccccgcctgcttgccagtcagcagaccccacgggaaatctaccaggatatgtggcacaccctgt- tacaacgtcgaccatggcgcggg caattgatcaaccgccaccgtgacggcagcctttttctggttgagatcgatatcaccccggtgattaacccgt- ttggcgaactggaacactacctggcca tgcagcgcgatatcagcgccggttatgcgctggagcagcggttgcgtaatcacatggcgctgaccgaagcggt- gctgaataacattccggcggcgg tggtcgtggtcgatgaacgcgatcgtgtggttatggataacctcgcctataaaactttctgtgctgattgcgg- cggaaaagagctactgagcgaactcca tttttcagcccgtaaagcggagctggcaaacggccaggtcttaccggtggtgctgcgcggcgcggtgcgctgg- ttgtcggtcacctgctgggcgctg ccaggcgtcagcgaagaagccagtcgctactttattgataataccttgacgcgcacgctggtggtcatcaccg- acgacacccagcagcgccagcag caagagcaaggacggcttgaccgccttaaacagcagatgaccagcggcaaactgctggcggcgatccgcgaag- cgcttgacgccgcgctgatcc agcttaactgccccatcaatatgctggcggcggcgcggcgtttaaacggcagtgataacagcaacgtagcgct- ggacgccgcgtggcgcgaaggt gaagaagcgatggcgcggctgaaacggtgccgcccgtcgctggagctggaaagtgccgccgtctggccgctgc- aacccttttttgacgacttgcgc gcgctttatcacacccgctacgagcagggtaaaaatttgcaggtcacgctggattcgacgcatctggtgggat- ttggtcagcgaacccaactgctggc ctgcctgagtctgtggctcgatcgcacgctggatattgccgtcgggctgcgtgatttcaccgcccaaacgcag- atttacgcccgggaagaagcgggct ggctctcgttgtatatcactgacaatgtgccgttgattccgctgcgccatacccattcgccggatgcgcttaa- cgcaccgggaaaaggtatggagttgc ggctgatccagacgctggtagcgcatcacaacggcgcgatagaactcacttcacgccccgaagggggaagctg- cctgaccctacgattcccgctatt tcattcactgaccggaggttcaaaatga 143 atgacccagcgaaccgagtcgggtaataccgtctggcgcttcgatttatcccagcagttcaccgcgatga- gcggataagcgtggttctcagccggg cgaccgaggttgaacagacactccagcaggtgtgtgcgtattgcacaatgacgcctttttgcagcacggcatg- atctgtctgtacgacagccagcag gcgattttgactattgaagcgttgcaggaagccgatcagcagttgatccccggcagctcgcaaattcgctacc- gtccgggtgaagggctggtcggga cggtgctttcgcaggggcaatcgttagtgctggcgcgtgtggctgacgatcagcgctttcttgaccgcctggg- actgtatgattacaacctgccgtttatc gccgtgccgctgatagggccggatgcgcagacttttggcgtgctgacggcgcaaccgatggcgcgttacgaag- agcggttacccgcctgcacccg ctttctggaaacggtcgcgaatctggtggcgcagaccgtgcgtttgatgacgccgccggctgcacgcccttcc- ccacgcgctgccatcacgccaacc gccagcccgaaatcgtgcagtacttcacgcgcgttcggcttcgaaaatatggtcggcaacagcccggcaatgc- gccagaccatggagattatccgtc aggtttcgcgctgggataccaccgttctggtgcgcggcgagagcggcaccggcaaggaactgattgccaacgc- catccatcacaattcgccgcgcg ccagtgcgccatttgtgaaattcaactgtgcggcgctgccggacacattgcttgaaagcgaattatttggtca- tgaaaaaggcgcctttaccggcgcgg tacgccagcgtaaaggccgttttgagctggccgatggcggcacgctgtttcttgacgaaattggggaaagcag- cgcctcgtttcaggctaagctgctg cgtattttgcaggagggcgaaatggaacgcgtcggtggtgacgagacattgcaagtgaatgtgcgcatcattg- ccgcgacgaaccgcaaccttgaag atgaagtacgcctgggacattttcgcgaagatctctattaccgcctgaatgtgatgcccatcgccctgccgcc- gctgcgcgaacgccaggaccacatc gccgaactggcacattttctggtgcgtaaaatcgcccacaaccagaaccgcacgctgcgcattagcgagggcg- ctatccgcctgctgatgagctaca gctggcccggcaatgtgcgcgaactggaaaactgccttgagcgctctgcggtgatgtoggaaaacggtctgat- cgatcgggacgtgattttatttaatc atcgcgaccagccagccaaaccgccggttatcagcgtcacgcccgacgataactggctcgataacacccttga- cgagcgccagcggctgattgccg cgctgaaaaaagcgggatgggtacaagccaaagccgcccgcttgctggggatgacgccgcgccaggtcgctta- tcgtattcagaccatggatatca ccctgccaaggctataa 144 atgccgcaccacgcaggattgtcgcagcactggcaaacggttttttctcgtctgccggaagcgctcaccg- cgcaaccattgagcgcgcaggcgcagt cagtgctcacttttagtgattttgttcaggacagcatcatcgtgcatcctgagtggctggcagagcttgaaag- cgcaccgccgccagcgaacgagtggc aacactacgcgcaatggctgcaagcggcgctggagggcgtcaccgatgaaacctcgctgatgcgcacgctgcg- gctgtttcgccgtcgcattatggt gcgcatcgcctggagtcaggcgctacagttggtggcggaagaggatatcctgcaacagctcagcgtgctggcg- gaaactctgatcgtcgccgcgcg cgactggctctatgacgcctgctgccgtgagtggggaacgccgtgcaatccgcaaggcgtcgcgcagccgatg- ctggtgctcggcatgggcaaact tggcggcggcgaactcaatttctcatccgatatcgatttgatttttgcctggccggaaaatggcaccacgcgc- ggcggacgccgtgaactggataacg cgcagttttttacccgccttggtcaacggctaattaaagtcctcgaccagcccacgcaggatggctttgtcta- ccgcgtcgatatgcgcttgcgtcccttt ggcgacagcggcccgctggtgctgagttttgccgcgctggaagattactaccaggagcaggggcgcgactggg- aacgatacgcgatggtgaaagc gcgcattatgggggacaacgacggcgaccatgcgcgagagttgcgcgccatgctgcgcccgttcgttttccgc- cgctatatcgacttcagcgtgatcc agtctctgcgcaacatgaaaggcatgattgcccgcgaagtgcggcgtcgcggcctgaaggacaacataaaact- cggcgcgggcggtattcgcgaa atagagtttatcgtgcaggttttccagttgattcgcggcggtcgcgagcctgcgctgcaatcgcgttcgctgt- tgccgacgcttgctgccattgatcaact acatctgctgccagatggtgatgcaccccggctgcgcgaggcgtatttgtggctgcgacggctggaaaacttg- ctgaaagcattaatgacgaacag acacagacgctgccggccgatgatttgaatcgcgcgcgcctcgcctggggaatgggcaaagagagctgggaag- cgctctgcgaaacgctggaag cgcatatgtcggcggtgggcagattttcaacgatctgattggcgatgatgaaacggattcgccggaagatgcg- ctttctgagggctggcgcgaattgt ggcaggatgcgttgcaggaagaggactctacgcccgtgctggcgcatctttccgaggacgatcgccgccgcgt- ggtggcgctgattgctgattttcg caaagagctggataaacgcaccattggcccgcgcgggcgacaggtactcgatcacttaatgccgcatctgctc- agcgatgtatgctcgcgtgacgat gcgccagtgccgctgtcgcgtctgacgccgctgctcaccggtattattacgcgcaccacttaccttgagctgc- tgagtgaattccccggtgcgctgaaa cacctcatttccctgtgcgccgcgtcgccgatggtggccagccaactggcgcgctacccgatcctgctcgatg- aactgctcgacccgaacacgctcta tcaaccgacggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagaggatgaa- gagcagcaactggaggcgctac ggcagtttaagcacgcgcagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatgaaagt- gagcgatcacttaacctggctggc ggaagcgattatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccgacgcat- ctgcacgatcgcgaagggcgc ggtttcgccgtggtcggttacggcaaacttggcggctgggaattaggttacagctccgatctggatctggtgt- tcctgcacgactgccccatggatgtga tgaccgatggcgagcgtgaaatcgatggccgccagttctatttgcgcctcgcgcagcgcgtcatgcacctgtt- cagcacgcgcacgtcgtccggcatt ctttatgaagtcgatgcgcgtttgcgcccgtccggcgcggccggaatgctggtgaccactgcggaagcgttcg- ccgattatcaaaaaaatgaagcctg gacatgggagcatcaggcgctggcgcgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgac- gccattcgccgcgatatcctgat gacctcccgcgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatcttagt- aacaagcacaaagaccgtttcga tctgaaagccgatgaaggcggtatcaccgatattgagtttatcgctcagtatctggtgctgcgctttgcccat- gagaagccgaaactgacgcgctggtc ggataatgtgcgcatcctcgaagggctggcgcaaaacggcatcatggatgagcaggaagagcaggcattgacg- ctggcgtacaccacgttgcgtg atgagctgcaccacctggcgctgcaagagctgccaggacatgtggcgctctcctgttttgtcgccgagcgtgc- gcttatcaaaaccagctgggacaa gtggctggtggaaccgtgcgccccggcgtaa 145 atgaaaaacacaacattaaaaacggctcttgctttcgctggcgttgctgccaggcctggcgatggcggct- cccgctgtggcggataaagccgacaacg gctttatgatgatttgcaccgcgctggtgctgtttatgaccattccgggcattgcgagttctacggcggtttg- atccgcggtaaaaacgtgctgtcgatgc tgacgcaggttgccgtcaccttcgctctggtgtgcatcctgtgggtggtttacggctactctctggcatttgg- cgagggcaacagcttcttcggcagtttc aactgggcgatgttgaaaaacatcgaattgaaagccgtgatgggcagcatttatcagtacatccacgtggcgt- tccagggctcctttgcttgtatcaccg ttggcctgattgtcggtgcgctggctgagcgtattcgcttctctgcggtgctgatttttgtggtggtatggct- gacgctttcttatgtgccgattgcgcacat ggtctggggtggcggtctgctggcaacccacggcgcgctggatttcgcgcgcggtacggttgttcacatcaac- gccgcgatcgcaggtctggtggg ggcttacctgattggcaaacgcgtgggctttggcaaagaagcgttcaaaccgcataacctgccgatggtcttc- accggcaccgcgatcctctatgttgg ctggtttggcttcaacgccggctctgcaagctcggcgaacgaaatcgctgcgctggctttcgtgaacacggtt- gttgccactgcggccgctattctggc gtgggtatttggcgagtgggcaatgcgcggtaagccgtctctgctcggtgcctgttctggtgccatcgcgggt- ctggttggtatcaccctcggcgtgcg gttatgtgggtgtcggcggcgcgctgattgtgggtctgattgccggtctggcagggctgtggggcgttactgc- actgaaacgtatgttggtgttgatga cccatgcgatgtcttcggtgtgcacggcgtgtgcggcatcgtgggttgtatcctgaccggtatcttcgcgtct- acgtcgctgggcggtgtcggtttcgct gaaggggtgaccatgggccatcaggtactggtacagctggaaagcgttgccatcactatcgtgtggtctggcg- tggtggcctttatcggttacaaactg gcggatatgacggtaggcctgcgcgtaccggaagagcaagagcgtgaagggctggatgtgaacagccacggcg-
aaaatgcgtataacgcctga 146 ttcttggttctctggagcgctttatcggcatcctgactgaagaatttgcaggcttcttcccaacctggct- tcacccgtgcaggtagttgtgatgaacatca ctgattcgcaggctgaatacgttaacgaattgacccgtaaactgcaaaatgcgggcattcgtgtaaaagcaga- cttgagaaacgagaagattggcttta aaatccgcgagcacactttacgtcgtgtcccttatatgctggtttgtggtgacaaagaggtcgaagccggcaa- agttgctgtgcgtacccgtcgcggta aagacctgggtagcctggacgtaaatgatgttatcgagaagctgcaacaagagattcgcagccgcagtcttca- acaactggaggaataaggtattaaa ggcggaaaacgagttcaaacggcgcgtcccaatcgtattaatggcgagattcgcgccacggaagttcgcttaa- caggtaggaaggcgagcagctt ggtatt 147 cgttctgtaataataaccggacaattcggactgattaaaaaagcgccctcgcggcgctttttttatattc- tcgactccatttaaaataaaaaatccaatcgga tttcactatttaaactggccattatctaagatgaatccgatggaagctcgctgttttaacacgcgttttttaa- ccattattgaaagtcggtgcttctttgagcga acgatcaaatttaagtcgattcccatcaaaaaaatattctcaacctaaaaaagtttgtgtaatacttgtaacg- ctacatggagattaactcaatctagagggt attaataatgaatcgtactaaactggtactgggcgc 148 cgcgtcaggttgaacgtaaaaaagtcggtctgcgcaaagcacgtcgtcgtccgcagttctccaaacgtta- attggtttctgcttcggcagaacgattggc gaaaaaacccggtgcgaaccgggtttttttatggataaagatcgtgttatccacagcaatccattgattatct- cttctttttcagcatttccagaatcccctca ccacaaagcccgcaaaatctggtaaactatcatccaattttctgcccaaatggctgggattgttcattttttg- tttgccttacaacgagagtgacagtacgc gcgggtagttaactcaacatctgaccggtcgat 149 ttgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgngcggaagc- acaggagagcttgctctctgggtgacg agcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaat- accgcataacgtcgcaagaccaa agagggggaccttcgggcctcttgccatcagatgtgcccagatgggattagctagtaggtggggtaacggcnc- acctaggcgacgatccctagctg gtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaat- attgcacaatgggcgcaagcct gatgcagccatgccgcgtgtatgaagaaggccttcgggttgtttaagtactttcagcggggaggaaggtgttg- nggttaataaccncagcaattgacgt tacccgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggaggntgcaagcgttanncgg- aatnantgggcgtaaagcgtn cgcaggcggtntgtnaagtcggatgtgaaatccccgggctcaacctgggaactgcattcgaaactggcaggct- agagtnnngtagaggngggtag aattccnggtgtagcggtgaaatgcgtagagatcnggangaanaccngtggcgaaggcggcccnctggacaaa- gactgacgctnaggngcgaa agcgggggagcaaacaggattagataccctngtagtccacgccgtaaacgatgtcgacttggaggttgtgccc- ttgaggcgtggcttccggagcta acgcgttaagtcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcaca- agcggtggagcatgtggtttaatt cgatgcaacgcgaagaaccttacctactcttgacatccagagaacttnncagagatgnnttggtgccttcggg- aactctgagacaggtgctgcatggc tgtcgtcagctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagc- ggtncggccgggaactcaaaggagac tgccagtgataaactggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacg- tgctacaatggcgcatacaaagag aagcgacctcgcgagagcaagcggacctcataaagtgcgtcgtagtccggattggagtctgcaactcgactcc- atgaagtcggaatcgctagtaatc gtagatcagaatgctacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggtt- gcaaaagaagtaggtagcttaacct tcgggagggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctg- cggttggatcacctcctt 150 atgaccatgcgtcaatgtgccatttacggcaaaggtggtatcggtaaatccactaccacgcaaaacctgg- tcgccgcgctggcggagatgggcaaga aagtaatgatcgtcggctgcgacccgaaagcagactccactcgtctgatcctgcatgcgaaagcgcagaacac- cattatggagatggcggctgaagt cggctccgtggaagaccttgaactggaagatgtgctgcaaatcggttacggcgacgtacgctgcgcagaatcc- ggcggcccggaaccaggcgttg gctgtgctggtcgcggggtaattaccgccatcaacttcctggaagaagaaggcgcctatgttcccgacctcga- tttcgtcttttacgacgtgttgggcga cgtggtgtgcggggggttcgccatgccgattcgcgaaaacaaagcgcaggagatctacatcgtctgctccggc- gaaatgatggcgatgtacgccgc caacaacatctctaaaggcatcgtgaaatacgccaaatccggcaaaggcgccttggcgggctgatctgtaact- cccgtcagaccgaccgcgaagat gagctgatcatagcgctggcggaaaaactcggcacccagatgatccacttcgtgccgcgcgacaacatcgtgc- aacgcgctgaaatccgccgtatg acggtgattgagtacgatccgaaatgcaaccaggccaatgaataccgcacgctggcgaacaagatcgtcaact- tacaccaaaatggtcgtgccaacg cccatcaccatggacgaactggaagagctgttgatggaattcggcattatggatgtggaagacaccagcatta- tcggtaaaaccgccgcagaagaaa acgcggtttga 151 atgagcaatgcaacaggcgaacgtaataggagatcatccaggaagtgctggagatctttccggaaaaaac- gcgcaaagaacgcagaaagcacatg atggtgagcgacccggagatggaaagcgtcgggaaatgcatcatctccaaccgtaagtcgcagcccggcgtaa- tgaccgtgcgcggttgctcttac gccggttctaaaggggtggtattcgggccgatcaaagatatggcccatatttcccacggcccggtcggctgcg- gtcagtactcccgcgccgggcggc gtaactactacaccggcgtcagcggtgtggatagcttcggtacgctcaactttacctccgattttcaggagcg- cgatatcgtgtttggcggcgataaaaa gctgaccaaactgattgaagagatggagacgctgttcccgctgaccaaagggatctccattcagtccgaatgc- ccggtcggcctgattggcgacgac attgaagccgttgccaacgccagccgcaaagccatcaataaaccggtcattccggtgcgctgcgaaggttttc- gcggcgtttcccagtcactcggtca ccacattgccaacgacgtgatccgcgactgggtactggataaccgcgaaggcaagccgtttgaggccggtcct- tatgacgtggcgatcatcgccgat tacaacatcggcggcgatgcctgggcgtcgcgcattttgctcgaagagatgggcctgcgcgtggtggcgcagt- ggtccggcgacggcacgctggtt gagatggagaccacgccgttcgtcaaactcaaccttgtgcactgctaccgctcaatgaactatatctcccgcc- atatggaggagaaacacggtattccg tggatggagtacaacttcttcggtccgaccaaagtcgccgaatcgttgcgcaaaatcgccgatatgtttgatg- acaccattcgcgccaacgccgaagc ggtgatcgccaaatatcaggcgcagaacgacgccatcatcgccaaataccgtccgcgtctggaaggccgcaaa- gtgctgctgtatatgggcggttta cgtcctcgccatgtgattggcgcttatgaagatctgggaatggaaattatcgctgcgggttatgaattcgccc- acaacgatgactacgaccgcaccctg ccggatctgaaagaaggcaccttgctgttcgacgatgccagcagttatgaactggaagcctttgtcaaagcgc- tgaagccggatctgatcggctccgg cattaaagagaagtacatatccagaaaatgggcgtgccgtttcgccagatgcactcctgggattactccggcc- cctatcacggttatgacggctttgcc atcttcgcccgcgatatggatatgacgatcaacaaccccgcgtggggccagttgaccgcgccgtggctgaaat- ccgcctga 152 atgggacgcggcgagcgccgcctgttccatgccgtgcgccacatcgtcaaccgctaccacccggccgccg- tctttatctataacacctgcgttcccgc gatggagggcgacgatatcgaagccgtctgccaggcggcagaaaccgccatccgcgtaccggtgattgccgtt- gatgtcgccgggttttacggcag caaaaatctcggcaaccggttggccggtgaagtgatggtgaaaaaaggtgattggcgggcgtgaacccgcgcc- gtggccggaagataccccttttgc cccggcgcaccgccacgatatcgggctgattggcgaattcaatattgccggagagttctggcatattcagccg- ctgctcgatgagctgggtattcgcgt gctcggcagcctctccggcgacgggcgcttcagtgaaatccagacgctgcaccgggcgcaggtcaatatgctg- gctcctccagggcgctgatcaa cgtcgcccgctcgctggagcagcgctacggcacgccgtggtttgaaggcagtttttatggtgttcgcgccacc- tctgacgccctgcgccaactggcg gcgctgaccggagaccgcgatctgatgcagcgcaccgattcagctcattgcccgcgaagagcagcaaacagag- caggcgctggccccgctgcgc gagcgcctgcgcgggcgcaaagcgctgctctatnccggcggcgtgaaatcctggacggtggtttcggcgcttc- aggatctgggcatggaagtggtg gcgaccggcacgcgcaaatccaccgaagaggataaacagcgcatccgcgaactgatgggcgccgacgcgctga- tgcttgatgaaggtaacgccc gctcgctgctggacgggtttaccgctacaaggcggacatgatgatcgccgggggacgcaatatgtcaccgcct- acaaagcgcggctgccgttcct cgatatcaatcaggagcgcgagcacgcctttgccggctaccgcggcattgtcaccctggccgaacagctctgc- ctgaccatggaaagcccggtctg gccgcaaacccattcccgcgcaccgtggcaataa 153 atgatggagcaaatggacgtgccgtccagcctgctttccgatccctccgaagtgctggataccccggctg- acgggcattaccacatgtatgcggggcg gtacgacccagcaggagatgcgcgaagcgcctgacgctatcgacaccctgctgctgcaaccctggcaactggt- gtaaaccaaaaaagtggtgcag gaaagctggaaccagcccgctaccgaggtgcaaatcccaatggggctggccggaaccgacgagctgctgatga- cggtaagccagttaaccggca aagccattccggatagcttagcgctggaacgcggtcggctggtggatatgatgctcgactcccacacctggct- gcacggcaagaaattcggcctgttc ggtgacccggattttgtcatggggctgacccgcttcctgctggaactgggctgcgaaccgacggtgattctgt- gccataacggcagcaagcgctggc agaaagcgatgaagaaaatgcttgaagcctcgccgtacgcgaaagagagcgaagtctttatcaactgcgattt- gtggcatttccgctcgctgatttac ccgtcagccggactttatgatcggcaactcctacgccaagtttatccaccgcgatacgctggcgaagggtgag- cagtttgaagtgccgctgatccgcc tggggttcccgctgttcgatcgccaccatctgcaccgccagaccacctggggttacgaaggggccatgagtat- cctcaccacgctggttaatgcggtg ctggagaaagtcgacagagagaccatcaagctcggcaaaaccgactacagcttcgatcttatccgttaa 154 atgcagcgcgacatcagcaccagctacgcgctggaacaacggctgcgcaatcatatgacgctgaccgaag- ccgtcttgaataacattccggcggcg gttgtagtggtggatgaacgcgatcgggtggtgatggataacctcgcctacaaaaccttttgcgccgattgcg- gcggtaaagaactactcaccgaaatc aacttttccgcccataaggcggagctggcgcagggcctggtactgccggtagtgctgcgcggcaccgtgcgct- ggttgtccgttacctgttgggcgct gccgggcgtcagcgaagaagcaggccgctactttattgatagcgccgtgccgcgcacgctggtggtgatcacc- gataatactcagcagcagcaaca acaggagcaggggcgtcttgatcgtctgaagcagcagataaccagcggtaaattgctggcggcgatccgcgaa- tcgctggacgccgcgctggtac aactcaattgcccaattaatatgctggccgccgcacgccgcttaaatggcgacgagcatagcaatctggcgct- ggatgccgcatggcgtgaaggcga agaagcgatggcgcggttgcagcgctgccgcccgtcgctggaactggaaagcccggcagtctggccgctccag- ccgttccttgacgatctgcgtgc cctgtatcacacccgatataaccagggcgaaaacctgcaaattgagctggaatcccccgacctggtgggcttt- ggccagcgaacacaactgcttgcct gcctgagcctgtggctcgacagaaccctggatattgccgcggagctacgtgatttcacggtacagactcaact- ttacgcccgcgaagagagcggctg gctgtcgttctatttaaacgacaatgtgccgctgattcaggtgcgctacacccattcacccgatgcactnaat- gcgcccggtaaaggcatggagctgcg gctgatccagacgctggtcgcccaccatcgaggcgcaatagaactgacctcacgccctcagggaggcacctgt- ctgatcctgcgtttcccattatttta ctcgctgacaggaggctcactatga 155 atgactcagcgaaccgagtcgggtacaaccgtctggcgctttgacctctcccaacagtttacagccatgc- agcgtatcagtgtggtgttaagccgcgc gacggagatcgggcagacgctacaggaagtgctgtgcgtgctgcacaacgatgcctttatgcagcacgggatg- atctgtccgtacgcgcgggtgcg cgtcttcgcgagcgtatggctttga 156 atgcgcgtggaagactggtcaacgctgaccgaacggctcgatgcccatatggcaggcgtgcgccgaatct- ttaacgaactgatcggtgatgacgaaa gtgagtcgcaggacgatgcgctctccgagcactggcgcgagctgtggcaggacgcgcttcaggaagatgacac- cacgccggtgagacgcactta accgacgacgcgcgccatcgcgtggtggcgctgatcgctganttccgtcttgagagaacaaacgcgccatcgg- cccgcgtngcgccaggtgctg gatcacctgatgccgcacctgagagcgaagtctgctcgcgtgccgatgcgccggtgccgctgtcgcggatgat- gcccctgagaggggattatca cccgtactacctaccttgaactcctgagcgagttccctggcgcgcttaagcacctgatttcactctgcgccgc- gtcgccgatggtggccaacaagctgg cgcgttacccgctgctgctggatgagctgctcgatccgaataccctttatcaaccgacggcgaccgacgccta- ccgggacgaactgcgtcagtatctg ctgcgcgtgccggaagaagacgaagagcaacagctcggaggcgctgcgtcagtttaagcaggcccagatgctg- cgcgtggcggccgcagatattg ccggaacgctgccggtgatgaaagtgagcgatcacttaacctggcttgcggaagcgattatcgacgcggtggt- gcatcaggcctgggtgcagatggt ggcgcgctatggccagccgaaacatctggctgaccgtgatggtcgcggcttcgctggtggtgggttacggtaa- gctcggcggttgggagctgggcta tagctccgatctggatttaatcttcctccacgactgcccggttgatgtgatgaccgacggcgagcgcgagatt- gacgggcgtcagttctacctgcgcct ggcgcagcgcatcatgcacctgttcagcacccgcacctcgtcgggcattttgtatgaagtggatgcccgtagc- gcccgtccggcgcggcgggcatg ctggtcacctcgacggagtccttcgctgattaccagaagaatgaagcctggacgtgggagcatcaggcgctgg- tgcgcgcccggtggtgtatggcg atccgctgctgaaaacgcagtttgacgtgattcgtaaggaagtcatgaccaccgtgcgcgatggcagcacgct- gcaaacggaagtgcgcgaaatgc gcgagaaaatgcgcgcgcacttaggcaataaacatcgcgatcgctttgatattaaagccgatgagggcggtat- taccgatattgagtttattacccagta tctggtgttgctgcacgcgcatgacaagccgaagctgacgcgctggtcggataacgtgcgcattaggaactgc- tggcgcaaaacgacattatggac gagcaggaggcgcaggccttaacccgtgcctatacaacgcttcgcgatgagctccatcatctggcgttgcagg- agcagcngggncacgtggcgct ggactgtttcaccgctgaacgcgctcaggtaacggccagctggcagaagtggctggtggaaccgtgcgtaaca- aatcaagtgtga 157 agatgtgcccagatgggattagctagtaggtggggtaacggcncacctaggcgacgatccctagctggtc- tgagaggatgaccagccacactggaa ctgagacacggtccagactcctacgggaggcagcagtggggaatattgcacaatgggcgcaagcctgatgcag- ccatgccgcgtgtatgaagaag gccttcgggttgtaaagtactttcagcggggaggaaggtgttgtggttaataaccncagcaattgacgttacc- cgcagaagaagcaccggctaactcc gtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaattactgggcgtaaagcgcacgcaggcg- gtctgtcaagtcggatgtgaaat ccccgggctcaacctgggaactgcattcgaaactggcaggctagagtcttgtagaggggggtagaattccagg- tgtagcggtgaaatgcgtagagat ctggaggaataccggtggcgaaggcggccccctggacaaagactgacgctcaggtgcgaaagcgtggggagca- aacaggattagataccctggt agtccacgccgtaaacgatgtcgacttggaggttgtgcccttgaggcgtggcttccggagctaacgcgttaag- tcgaccgcctggggagtacggccg caaggttaaacctcaaatgaattgacgggggcccgcacaagcggtggagcatgtggtttaattcgatgcaacg- cgaagaaccttacctactcttgacat ccagagaacttnncagagatgnnttggtgccttcgggaacctgagacaggtctgcatggctgtcgtcagctcg- tgttgtgaaatgttgggttaagtc ccgcaacgagcgcaacccttatcctttgttgccagggtccggccgggaactcaaaggagactgccagtgataa- actggaggaaggtggggatgac gtcaagtcatcatggcccttacgagtagggctacacacgtgctacaatggcgcatacaaagagaagcgacctc- gcgagagcaagggacctcataa agtgcgtcgtagtccggattggagtctgcaactcgactccatgaagtcggaatcgctagtaatcgtagatcag- aatgctacggtgaatacgttcccggg ccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaagaagtaggtagcttaaccttcgggagggc- gcttaccactttgtgattcatgactg gggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttggatcacctcctt 158 atgaccatgcgcaatgtgccatttacggcaaaggtggtatcggtaaatccactaccacgcaaaacctggt- cgccgcgctggcggagatgggcaaga
aagtaatgatcgtcggctgcgacccgaaagcagactccactcgtctgatcctgcatgcgaaagcgcagaacac- cattatggagatggcggctgaagt cggctccgtggaagaccttgaactggaagatgtgctgcaaatcggttacggcgacgtacgctgcgcagaatcc- ggcggcccggaaccaggcgttg gagtgctggtcgcggggtaattaccgccatcaacttcctggaagaagaaggcgcctatgttcccgacctcgat- ttcgtcttttacgacgtgttgggcga cgtggtgtgcggggggttcgccatgccgattcgcgaaaacaaagcgcaggagatctacatcgtctgctccggc- gaaatgatggcgatgtacgccgc caacaacatctctaaaggcatcgtgaaatacgccaaatccggcaaagtgcgccttggcgggctgatctgtaac- tcccgtcagaccgaccgcgaagat gagctgatcatagcgctggcggaaaaactcggcacccagatgatccacttcgtgccgcgcgacaacatcgtgc- aacgcgctgaaatccgccgtatg acggtgattgagtacgatccgaaatgcaaccaggccaatgaataccgcacgctggcgaacaagatcgtcaaca- acaccaaaatggtcgtgccaacg cccatcaccatggacgaactggaagagctgttgatggaattcggcattatggatgtggaagacaccagcatta- tcggtaaaaccgccgcagaagaaa acgcggtttga 159 atgagcaatgcaacaggcgaacgtaatctggagatcatccaggaagtgctggagatctttccggaaaaaa- cgcgcaaagaacgcagaaagcacatg atggtgagcgacccggagatggaaagcgtcgggaaatgcatcatctccaaccgtaagtcgcagcccggcgtaa- tgaccgtgcgcggttgctcttac gccggttctaaaggggtggtattcgggccgatcaaagatatggcccatatttcccacggcccggtcggctgcg- gcagtactcccgcgccgggcggc gtaactactacaccggcgtcagcggtgtggatagcttcggtacgctcaactttacctccgattttcaggagcg- cgatatcgtgtttggcggcgataaaaa gctgaccaaactgattgaagtgatggagacgctgttcccgctgaccaaagggatctccattcagtccgaatgc- ccggtcggcctgattggcgacgac attgaagccgttgccaacgccagccgcaaagccatcaataaaccggtcattccggtgcgctgcgaaggttttc- gcggcgtttcccagtcactcggtca ccacattgccaacgacgtgatccgcgactgggtactggataaccgcgaaggcaagccgtttgaggccggtcct- tatgacgtggcgatcatcggcgat tacaacatcggcggcgatgcctgggcgtcgcgcattttgctcgaagagatgggcctgcgcgtggtggcgcagt- ggtccggcgacggcacgctggtt gagatggagaacacgccgttcgtcaaactcaaccttgtgcactgctaccgctcaatgaactatatctcccgcc- atatggaggagaaacacggtattccg tggatggagtacaacttcttcggtccgaccaaagtcgccgaatcgttgcgcaaaatcgccgatatgtttgatg- acaccattcgcgccaacgccgaagc ggtgatcgccaaatatcaggcgcagaacgacgccatcatcgccaaataccgtccgcgtctggaaggccgcaaa- gtgctgctgtatatgggcggttta cgtcctcgccatgtgattggcgcttatgaagatctggggatggaaattatcgctgcgggttatgaattcgccc- acaacgatgactacgaccgcaccctg ccggatctgaaagaaggcaccttgctgttcgacgatgccagcagttatgaactggaagcctttgtcaaaggct- gaagccggatctgatcggctccgg cattaaagagaagtacatcttccagaaaatgggcgtgccgtttcgccagatgcactcctgggattactccggc- ccctatcacggttatgacggctttgcc atcttcgcccgcgatatggatatgacgatcaacaaccccgcgtggggccagttgaccgcgccgtggctgaaat- ccgcctga 160 atgaaggggaacgagatcctggctttgctcgatgaacctgcctgcgagcacaaccataaacagaaatccg- gctgcagcgcgccgaaacccggcgc gacagcgggcggctgcgcctttgacggtgcgcagatcaccctgctgccactctccgatgttgcccacctggta- cacggccccattggttgtaccggta gctcatgggataaccgtggcagcttcagttccggcccgacgatcaaccggctgggttaccaccgatctgagcg- aacaggatgtgatcatgggacg cggcgagcgccgcctgttccatgccgtgcgccacatcgtcaaccgctaccacccggccgccgtctttatctat- aacacctgcgttcccgcgatggagg gcgacgatatcgaagccgtctgccaggcggcagaaaccgccatcggcgtaccggtgattgccgttgatgtcgc- cgggttttacggcagcaaaaatct cggcaccggttggccggtgaagtgatggtgaaaaaggtgattggcgggcgtgaacccgcgccgtggccggaag- ataccccttttgccccggcgc accgccacgatatcgggctgattggcgaattcaatattgccggagagttctggcatattcagccgctgctcga- tgagctgggtattcgcgtgctcggca gcctctccggcgacgggcgcttcagtgaaatccagacgctgcaccgggcgcaggtcaatatgctggtctgctc- cagggcgctgatcaacgtcgccc gctcgctggagcagcgctacggcacgccgtggtttgaaggcagtttttatggtgttcgcgccacctctgacgc- cctgcgccaactggcggcgctgac cggagaccgcgatctgatgcagcgcaccgaacagctcattgcccgcgaagagcagcaaacagagcaggcgctg- gccccgctgcgcgagcgcct gcggggcgcaaagcgctgctctataccggcggcgtgaaatcaggtcggggtttcggcgcttcaggatctgggc- atggaagtggtggcgaccgg cacgcgcaaatccaccgaagaggataaacagcgcatccgcgaactgatgggcgccgacgcgctgatgcttgat- gaaggtaacgcccgctcgctgc tggacgtggtttaccgctacaaggcggacatgatgatcgccgggggacgcaatatgtacaccgcctacaaagc- gcggctgccgtcctcgatatcaat caggagcgcgagcacgcctttgccggctaccgcggcattgtcaccctggccgaacagctctgcctgaccatgg- aaagcccggtctcgccgcaaac ccattcccgcgcaccgtggcaataa 161 atgagccaaagtgctgagaaaattcaaaactgtcatccgctgtttgaacaggatgcgtaccagatgctga- taaagataaacggcaactggaagaggc ccacgatccggcgcgcgtgcaggaggtctttcaatggaccaccaccgccgagtatgaagcgcttaactttcaa- cgcgaagcgctgactatcgatccg gccaaagcctgccagcgctgggtgcggtactgtgctcgctgggctttgccaataccctgccctatgttcacga- ctcccaggggtgcgtggcctatttc cgcacctattttaaccgtcactttaaagagccgattgcctgtgtttctgactcgatgacggaagatgcggcag- tattcggcggcaacaacaacctgaaca ccgggttgcagaacgccaggccctctacaagccggaaatcattgccgtctccaccacctgtatggcggaggtc- atcggcgacgacctgcaggcgt ttattgctaacgccaaagaagacggctttatcgacgcggcgatcccggtgccttacgcgcacacgccaagctt- tatcggcagccatatcaccggctgg gacaatatgtttgagggcttcgcccgtacctttaccgccgattacagcggacaaccgggcaaattaccgcgta- tcaatctggtcagcggatttgaaacct atctcggtaatttccgcgtgctgaaacgcatgatggagcaaatggacgtgccgtgcagcctgctttccgatcc- ctccgaagtgctggataccccggctg acgggcattaccacatgtatgcgggcggtacgacccagcaggagatgcgcgaagcgcctgacgctatcgacac- cctgctgctgcaaccctggcaa ctggttgaaaaccaaaaaaagtggtgcaggaaagctggaaccagcccgctaccgaggtgataatcccaatggg- gctggccggaaccgacgagagc tgatgacggtaagccagttaaccggcaaagccattccggatagcttagcgaggaacgcggtcggctggtggat- atgatgctcgactcccacacctg gctgcacggcaagaaattcggcctgttcggtgacccggattttgtcatggggctgacccgcttcctgctggaa- ctgggctgcgaaccgacggtgattct gtgccatagcggcagcaagcgaggcagaaagcgatgaagaaaatgcttgaagcctcgccgtacgggaaagaga- gcgaagtctttatcaaagcga tttgtggcatttccgctcgctgatgtttacccgtcagccggactttatgatcggcaactcctacgccaagttt- atccagcgcgatacgctggcgaagggtg agcagtttgaagtgccgctgatccgcctggggttcccgctgttcgatcgccaccatctgcaccgccagaccac- ctggggttacgaaggggccatgag tatcctcaccacgctggttaatgcggtgctggagaaagtcgacagagagaccatcaagctcggcaaaaccgac- tacagcttcgatcttatccgttaa 162 atgccagaaattatccgtagttaaaaagccgctggccgtcagcccggtaaaaagtggccagccgctgggc- gcgattctggcgagcatgggctttgaa cagagcattccgctggttcatggcgctcacgggtgcagcgccttcgcgaaggtcttttttatccagcattttc- acgatccgatcccgctgcaatcgacgg caatggacccgacatcgaccattatgggtgccgatgagaacatctttaccgcgctgaatgtgctgtgttcacg- caacaacccgaaagcgattgttctgct gagcactggcctttccgaggcgcagggaagcgatatttcgcgcgtggtgcgccagttccgcgatgaatatccg- cgccataaaggggtggcgctgct gaccgtcaacacgccggatttttacggcagcctggaaaacggctacagcgcggtgctggagagcatggttgaa- cagtgggtgccggaaaaaccgc agccgggcgtgcgcaatcgccgcgtgaacctgctgctcagccatttgcttacgccgggcgacattgagctgct- gcgaagttatgtcgaggcatttggc ctgcagccggtgatggtccggatctttcccagtcgctggatggccatctcgccagcggggatttctcgccaat- tacccagggcggcagcagcctgc ggctgattgaacagatgggacagagtcttggcacgttcgccattggcgtatccctctcccgcgccgcgcaatt- gctggcgcagcgcagccatgcgga agtggtcaccctgccgcatctgatgaccatgagccagtgcgatacgtttattcatcaactgaagcgcctctcc- gggcgcgatgttccggcgtggatcga acgccagcgcgggcaactgcaggatgcgatgatcgattgtcatatgtggttgcagggcgcgcctgtcgcgctg- gccgccgagggcgatctgctcgc cgcctggtgcgatttcgcctgcgatatgggcatggtgcccggcccggtggtggcgccggtgagccagaaaggg- ttgcaggatctgccggtcgaaaa agtcattatcggcgatctggaggatatgcaggatctgttgtgtgaaacgcctgcatcgctgctcgtctctaat- tctcacgccgctgatttggccgggcagt tcgacattccgctggtgcgcgccggtttccccctgttcgaccgtctgggcgagtttcgccgcgtgcgccaggg- ttacgccgggatgcgcgacaccttg tttgagctggcgaatgccctgcgcgatcgccatcatcatcttgccgcttatcactcgccgctgcgccagcgtt- tttacgaacccgcatcttcgggaggtg actatgcaacatgttaa 163 atgaccctgaatatgatgatggacgccaccgcgcccgccgagatcgccggagcgctctcacaacagcatc- ccggattgtttttcaccatggttgaaca ggcgcccgtcgcgatttcactgaccgatgccgatgcccacattctctacgccaaccccgcgttttgtcgccag- tcggggtatgaactggaagagttgtt gcagcaaaacccgcgcctgcttgccagtaagaagacgccgcgtgaaatctaccaggaaatgtggcacaccctg- ctgcaacaccgtccgtggcgcg gacaactgatcaaccgtcgccgcgacggcagcctgtttctggtggaaatcgacatcaccccactgtttgatgc- gttcggcaaactcgaacattacctgg ccatgagcgcgacatcagcaccagctacgcgctggaacaacggctgcgcaatcatatgacgctgaccgaagcc- gtcttgaataacattccggcgg cggttgtagtggtggatgaacgcgatcgggtggtgatggataacctcgcctacaaaaccttttgcgccgattg- cggcggtaaagaactactcaccgaa atcaacttttccgcccataaggcggagctggcgcagggcctggtactgccggtagtgctgcgggcaccgtgcg- ctggttgtccgttacctgttgggc gctgccgggcgtcagcgaagaagaggccgctactttattgatagcgccgtgccgcgcacgctggtggtgatca- ccgataatactcagcagcagca acaacaggagcaggggcgtcttgatcgtctgaagcagagataaccagcggtaaattgctggcggcgatccgcg- aatcgctggacgccgcgctgg tacaactcaattgcccaattaatatgctggccgccgcacgccgcttaaatggcgacgagcatagcaatctggc- gctggatgccgcatggcgtgaagg cgaagaagcgatggcgcggttgcagcgctgccgcccgtcgctggaactggaaagcccggcagtctggccgctc- cagccgttccttgacgatctgc gtgccctgtatcacacccgatataaccagggcgaaaacctgcaaattgagctggaatcccccgacctggtggg- ctttggccagcgaacacaactgctt gcctgcctgagcctgtggctcgacagaaccctggatattgccgcggagctacgtgatttcacggtacagactc- aactttacgcccgcgaagagagcg gctggctgtcgttctatttaaacgacaatgtgccgctgattcaggtgcgctacacccattcacccgatgcact- caatgcgcccggtaaaggcatggagc tgcggctgatccagacgctggtcgcccaccatcgaggcgcaatagaactgacctcacgccctcagggaggcac- ctgtctgatcctgcgtttcccatta ttttactcgctgacaggaggctcactatga 164 atgactcagcgaaccgagtcggggacaaccgtctggcgctttgacctctcccaacagtttacagccatgc- agcgtatcagtgtggtgttaaaccgcgc gacggagatcgggcagacgctacaggaagtgctgtgcgtgctgcacaacgatgcctttatgcagcacgggatg- atctgtctgtacgacagtaagcaa gcgatcctttccattgaagccttgcatgaggccgatcagagttaattcccggcagttcacagattcgctaccg- tccgggcgaagggctggtaggcac ggtgctttcacagggacagtcgctggtactgccctgtgtctccgacgatcggcgttttctcgatcgcctggga- ttgtatgattacagcttgccgtttatcgc cgtgccgctgatggggccaaactcgcagcctatcggcgtgctggccgcccagcctatggcgcgttacgaggag- ggctgcccgcctgcacgcgttt tcttgaaaccgtcgccaatctggtggcgcaaaccgttcgcctgatgacaccgcccagcgtcgcgtctccaccc- cgtgctgctgccgcgcagattgcca gccagcgcgggtgcgcgtcttcgcgagcgtatggctttgaaaacatggtcggtaaaagcgcggctatgcgtca- gacgctggaaattattcgccaggt atcacgctgggacaccaccgtgctggtgcgtggcgaaagcggaaccggtaaagagttgatagccaacgctatc- caccacaattcaccgcgcgccg ccgcgccgtttgtcaaattcaactgcgcggcgctgcccgatacgctgaggagagtgaactcttcggtcatgaa- aaaggcgcgtttaccggcgcggtg cgccagcgcaaaggccgtttcgaactggcggatggcggtacgctgtttcttgatgagatcggcgaaagtagcg- cctcgtttcaggcgaaattgctgcg tatcttgcaggaaggcgaaatggaacgcgtcggcggcgacgaaacgctgcgggtgaatgtacggatcattgcc- gccaccaaccgcaatctggaag aggaagtgcggctgggtaattttcgcgaagatctctactatcgccttaatgtgatgccgatctccctgccccc- gctccgcgagcgtcaggaggacatcg tcgagctggcgcattttctggtgcgcaaaatcgcgcaaaaccagaaccgcacgctgcgcatcagcgatggcgc- gatccgtttgttgatgagctatagc tggcctggaaacgtgcgtgagctggaaaactgccttgagcgatcggcggtgatgtcggaaaacgggctgatcg- atcgcgacgtgattttgtttcacca cagggaaaatctgcaaaaacgccacagaccagtgcgccgcgcgaagagagctggctcgatcagaacctcgatg- aggacaaagattgatcgcc gcgctggagaaagccggttgggtacaggcaaaagccgcgcgcctgagggaatgaccccgcgccaggtggccta- tcgtattcagacgatggacat tgccatgccgagattgtag 165 atgccgctttcttcgcaggttacagcagcagtggcagaccgtttgcgaacgtctgcctgagtcattaccg- gcgtcatcgttaagcgagcaggcaaagag cgtgctcgtcttcagtgattttgtgcaggaaagtatcaccgccaacccgaactggctggcggaacttgagaac- gcaccaccgcaggcagaagagtgg cggcactatgctggctggctgcaaactgtactcgaagacgttacggatgaggccacgctgatgcgcgtcctgc- gccagttccgtcgtcggctgatggt ggcattgcctgggctcaggcgctggaactggtgagcgaagagagtacgctgcagcagttaagcgagctggcgc- aaacgttgattgtcgccgcgc gagactggctctatgcctgctgtaaagagtggggcacgccgtgcagcgaggaaggggttcctcagccgctgtt- gattctgggcatgggaaagc tgggcggctgcgagctgaacttctcctctgatatcgacctgatttttgcctggccggagaacggctccacgcg- cggaggccgccgcgagctgaacaa cgcgcagttctttacccgtctcggccagcgcctgattaaagcgctggatcagcccacgcaggacggttttgtt- taccgcgtggacatgcgcctgcgtcc gtttggcgacagcgggccgctggtgctgagctttgcggcgctggaagattattaccaggagcaaggtcgcgac- tgggagcgttacgcgatggtcaa agcgcggatcatgggcaacagcgacgacgcttatgccaacgagctgcgcgccatgctgcgtccgttcgtgttc- cgtcgctatatcgacttcagcgtca tccagtccctgcgaaatatgaaagggatgattgcccgcgaggtgcgccgccgtgggctgaaagacaatatcaa- gctcggtgcgggcggcatccgc gaaatcgaatttatcgtccaggtcttccagcttattcgcggcggacgcgagccgtcgctgcagtcccgttcct- tattaccgacgctgagcgccattgcgc agctgcatctcctgccggacggcgacgcgcaaaccctgcgcgaggcctatcttttcctgcgtcgtctggaaaa- cctgctgcaaagcattaatgacgaa cagacccaaaccctgccgggcgacgaccttaaccgggcgcgtctggcctggggaatgcgcgtggaagactggt- caacgctgaccgaacggctcg atgcccatatggcaggcgtgcgccgaatctttaacgaactgatcggtgatgacgaaagtgagtcgcaggacga- tgcgctctccgagcactggcgcg agctgtggcaggacgcgcttcaggaagatgacaccacgccggtgctgacgcacttaaccgacgacgcgcgcca- tcgcgtggtggcgctgatcgct gatttccgtcttgagctgaacaaacgcgccatcggcccgcgtggtcgccaggtgctggatcacctgatgccgc- acctgctgagcgaagtctgctcgc gtgccgatgcgccggtgccgctgtcgcggatgatgcccctgctgaggggattatcacccgtactacctacctt- gaactcctgagcgagttccctggc gcgcttaagcacctgatttcactctgcgccgcgtcgccgatggtggccaacaagctggcgcgttacccgctgc- tgctggatgagctgctcgatccgaa taccctttatcaaccgacggcgaccgacgcctaccgggacgaactgcgtcagtatctgctgcgcgtgccggaa- gaagacgaagagcaacagctgg aggcgctgcgtcagtttaagcaggcccagatgctgcgcgtggcggccgcagatattgccggaacgctgccggt- gatgaaagtgagcgatcacttaa cctggcttgcggaagcgattatcgacgcggtggtgcatcaggcctgggtgcagatggtggcgcgctatggcca- gccgaaacatctggctgaccgtg atggtcgcggcttcgcggtggtgggttacggtaagctcggcggttgggagctgggctatagctccgatctgga-
tttaatcttcctccacgactgcccgg ttgatgtgatgaccgacggcgagcgcgagattgacgggcgtcagttctacctgcgcctggcgcagcgcatcat- gcacctgttcagcacccgcacctc gtcgggcattgtatgaagtggatgcccgtctgcgcccgtccggcgcggcgggcatgctggtcacctcgacgga- gtccttcgctgattaccagaaga atgaagcctggacgtgggtgcatcaggcgctggtgcgcgcccgtgtggtgtatggcgatccgctgctgaaacg- cagtttgacgtgattcgtaagga agtcatgaccaccgtgcgcgatggcagcacgctgaaacggaaggcgcgaaatgcgcgagaaaatgcgcgcgca- cttaggcaataaacatcgc gatcgctttgatattaaagccgatgagggcggtattaccgatattgagtttattacccagtatctggtgttgc- tgcacgcgcatgacaagccgaagctgac gcgctggtcggataacgtgcgcattctggaactgctggcgcaaaacgacattatggacgagcaggaggcgcag- gccttaacccgtgcctatacaac gcttcgcgatgagctccatcatctggcgttgcaggagcagccgggacacgtggcgctggactgtttcaccgct- gaacgcgctcaggtaacggccag ctggcagaagtggctggtggaaccgtgcgtaacaaatcaagtgtga 166 atgaagatagcaacacttaaaacgggtctgggttcgctggcactgctgccgggcctggcgctggctgctg- cacctgcggtggcagacaaagccgat aacgcctttatgatgatcagcaccgcgctggtgctgttcatgtccattccgggcattgcgctgttctatggcg- gcctgatccgtggcaaaaacgttctctc catgctgacgcaggttgccgtaacgttcgcgctggtctgcgtactgtgggtggtttacggttactcgctggct- ttcggcacgggcaacgcgttctttggta acttcgactgggtgatgctgaaaaatattgaactgaccgcgctgatgggcagtttctaccagtatattcacgt- tgctttccagggctcgttcgcctgcatta ccgtcgggctgattgtaggcgcgcttgccgagcgtattcgtttctctgcggtcctgatcttcgtggtggtctg- gctgacgctctcctatgtgccgattgcg cacatggtctggggtggcggctgctggcgacgcatggcgcgctggacttcgcgggcggtaccgttgtgcacat- taacgccgcatagcgggtctg gttggcgcatacctgattggcaaacgcgtgggcttcggtaaagaagcgttcaaaccgcacaacctgccgatgg- tcttcaccggtaccgcgatcctcta ctttggctggtttggtttcaacgccggctcagcaagtgccgcgaacgaaatcgccgcgctggccttcgtgaat- accgttgtggccacggcaggtgcaa tcctctcctgggtctttggcgagtgcgctgtgcgcggtaaaccttctctgctgggtgcctgttcgggggcgat- tgctggtctggtcggtatcaccccagc atgtggttatgtcggtgtgggtggcgcgctgctggtcggcctggtgtcaggtctggcgggtctgtggggcgtg- acggcgctgaaacgtattctgcgcg ttgatgacccttgcgatgtgtttggcgtgcacggcgtgtgcggcatcgtcggctgtatcatgaccggtatctt- tgcagcgaaatcgctgggtggcgtgg gctacgcagaaggcgtcaccatggcccatcaggtgctggtgcagctggaaagtattgctgtcaccgtggtgtg- gtctgccgttgtcgctttcattggcta caaactggcggacatgacggttggtctgcgcgtgccggaagagcaggaacgcgaaggtctggacgtcaacagc- cacggcgagaatgcgtataac gcatga 167 gccgagagaggggcccgcgtcggattaggtagttggtgaggtaatggctcaccaagccttcgatccgtag- ctggtctgagaggatgatcagccaca ctgggactgagacacggcccagactcctacgggaggcagcagtggggaatattggacaatgggcgcaagcctg- atccagcaatgccgcgtgagtg atgaaggccttagggttgtaaagctctttcgcacgcgacgatgatgacggtagcgtgagaagaagccccggct- aacttcgtgccagcagccgcggta atacgnagggagcnagcgttnntcggaattactgggcgtaaagngcgcgtaggcggcntgttnagtcagaagt- gaaagccccgggctcaacctgg gaatagcttttgatactggcaggcttgagttccggagaggatggtggaattcccngtgtagnggtgaaatncg- tagatattgggangaacaccngtgg cgaaggcggcnatctggacgganactgacgctgaggcgcgaaagcgtggggagcaaacaggattagataccct- ngtagtccacgccgtaaacga tgaatgctagacgtcggggtgcatgacttcggtgtcgccgctaacgcattaagcattccgcctggggagtacg- gccgcaaggttaaaactcaaagg aattgacgggggcccgcacaagcggtggagcatgtggtttaattcgaagcaacgcgcagaaccttaccaaccc- ttgacatgtccactttgggctcgag agatngggtccttcagttcggctgggtggaacacaggtgctgcatggctgtcgtcagctcgtgtcgtgagatg- ttgggttaagtcccgcaacgagcgc aacccctaccgtcagttgccatcattcagttgggcactctggtggaaccgccggtgacaagccggaggaaggc- ggggatgacgtcaagtcctcatg gcccttatgggttgggctacacacgtgctacaatggcggtgacagtgggaagcgaagtcgcgagatggagcaa- atccccaaaagccgtctcagttc ggatcgcactctgcaactcgagtgcgtgaagttggcaatcgctagtaatcgcggatcagcacgccgcggtgaa- tacgttcccgggccttgtacacacc gcccgtcacaccatgggagttggttttacccgaaggtggtgcgctaaccgcaaggaggcagccaaccacggta- aggtcagcgactggggtgaagt cgtaacaaggtagccgtagggcaacctgcggctggatcacctccttt 168 atggccaaagcgcctctgcgtcagatcgccttttacggcaagggcggtatcggcaagtccaccacctctc- agaacacgctggccgcgctggtcgag ctggatcagaggatcctgatcgtcggctgcgacccgaaggccgactcgacccgcctgatcctgcacgcaaagg- cccaggacaccgtcctgcatctg gccgccgaggccggctcggtcgaggatctggagctcgaggacgttctcaagatcggctacaagaacatcaagt- gcgtcgagtccggcggtccgga gccgggggtcggctgcgccggccgcggcgtcatcacctcgatcaacttcctggaagagaacggcgcctacgac- gacgtggactatgtgtcctacga cgtgctgggcgacgtggtctgcggcggcttctccatgccgatccgcgagaacaaggcccaggaaatctacatc- gtcatgccggcgagatgatggc gctgtacgccgccaacaacatcgccaagggcatcctgaagtacgcgcacagcggcggcgtccgtctcggcggc- ctgatagcaacgagcgccag accgacaaggaatgggatctggccgacgcgctggccaaccgcctgggctccaagctgatccacttcgtgccgc- gcgacaacatcgtccagcacgc cgagagcgccgcatgacgctcatcgagtacgccccggacagcaagcaggccggcgaataccgcgcgctcgcca- acaagatccatgcgaactcc ggccagggttgcatcccgaccccgatcaccatggaagagctggaagagatgctgatggacttcggcatcatga- agaccgaggagcagcagctcgc cgagctcgccgccaaggaagcggcgaaggccggcgcctga 169 atgagcctgtccgagaacaccacggtcgacgtcaagaacctcgtcaacgaagtcctcgaagcctatcccg- aaaaatcccgcaagcgccgcgccaa gcacctgaacgtgctggaggccgaggccaaggaagggcgtcaagtcgaacgtcaagtccatccccggcgtcat- gaccatccgcggctgcgcct atgccggctccaagggcgtggtgtggggtccgatcaaggacatgatccacatctcccacggtccggtcggctg- cggctactactcaggtccggccg ccgcaactactacatcggcgacaccggtgtggacagctggggcacgatgcacttcacctccgacttccaggag- aaggacatcgtcttcggcggcga caagaagctgcacaaggtcatcgaggaaatcaacgagctgttcccgctggtgaacggcatctcgatccagtcg- gaatgcccgatcggcctgatcgg cgacgacatcgaggctgtcgcccgcgccaagtcggcggaaatcggcaagccggtcatccccgtgcgctgcgaa- ggcttccgcggcgtgtcccagt cgctgggccaccacatcgccaacgacgccatccgagactgggtgttcgagaagacggaacccaaggccggctt- cgtctccaccccctatgacgtca ccatcatcggcgactacaacatcggcggcgacgcaggtcgtcccgcatcctgctggaggagatcggcctgcgc- gtgatcgcccagtggtcgggc gacggcacgctcgccgaactggagaacacgccgaaggccaaggtcaacctgatccactgctaccgctcgatga- actacatcgcgcgccacatgga agagaagttcaacattccttggatggaatacaacttcttcggcccgagccagatcgccgaatccctgcgcaag- atcgccgctctcttcgacgacaagat caaggagaacgccgagaaggtcatcgcccgctaccagccgatggtcgatgcggtcatcgccaagtacaagccg- cggctcgaaggcaagaaggtc atgatctacgtcggcggcctgcgtccccgccacgtcgtcgatgcctaccatgacctcggcatggagatcaccg- gcaccggctacgagttcgcccaca acgacgactatcagcgcacgcagcactacgtgaaggaaggcacgctgatctacgacgacgtcaccgcgttcga- actggagaagttcgtcgaggcg atgcgtcccgacctcgtcgcgtcgggcatcaaggaaaagtacgtgttccagaagatgggcctgccgttccgcc- agatgcacagctgggactactccg gcccgtaccacggctatgacggcttcgcgatcttcgcccgcgacatggacctggccatcaacaaccccgtctg- gggcgtgatgaaggccccgttctg a 170 atgctccaggacaagatccaggatgtcttcaacgaaccgggctgcgcgaccaaccaagccaaatcggcca- aggagaagaagaagggctgcacca agtcgctgaaaccgggggcggcagccggcggctgagcctatgacggggcgatgatcgtgctccagccgatcgc- cgacgccgcccatctggtccat ggccccatcgcctgcctcggaaacagttgggacaaccgcggctccaaatcctccggctcgcagctctaccgca- ccggcttcaccaccgatctgtcgg aactggacgtcatcggcggcggcgagaagaagctctaccgcgccatcaaggagatcgttcagcaatacgaccc- gccggccgtcttcgtctatcaga cctgcgtgcccgccatgaccggcgacgacatcgccgcattgcaagttcgccacgcagaagctgagcaagccgg- tgatcacggtggactcgcc gggcttcgtcgggtcgaagaatctcggcaacaagctggccggcgaagccctgctggagcatgtcatcggcacg- gtcgaaccggactacaccaccc cgaccgacgtctgcatcatcggcgaatacaaccttgccggcgagctgtggctggtcaagccgctgctggacga- gatcggcatccgcctcctgtcctg catttccggcgacggccgctaccgggaggtggcgcaggcccaccgcgcccgcgtcaccatgatggtgtgcagc- caggcgctggtgaatgtcggg cgcaagatggaggagcgctacggcatcctctatttcgaggggtccttctacggcgtgtccgacatgtcggaca- ccctgcgcaccatgacccgcatgc tggtggagcgggcgccgacaagggcctgatcgaccgggcggagggcgtgatcgcgcgggaggaaagccgggtc- tggcgccggctggaaccc tacaagccgcgcttcgacggcaagcgcgtccttctcttcaccggcggcgtcaagagctggtcgatggtcagcg- cgctggagggtgcggggctgacc atcctcggcacctccaccaagaaatcgaccagggaggacaaggagcgcatcaagaagatgaagggcgaagagt- tccaccagtgggacgatttgaa gccgcgcgacatctacaggatgctggccgacgatcaggccgacatcatgatgtccggcggccgctcgcagttc- atctcgctgaaggccaaggttcc ctggctcgacatcaaccaggagcgccaccacgcctatgccggctatgacggcatcgtcaatctctgcgaggag- atcgacaaaacgctgtcgaatcc gatctggcgtcaggtgcgtcagccggcaccgtgggagtccggcgcgtcctccacccttctggcttcctcgatg- gcggcggagtga 171 atgtcccacatccagcgcttcccctccgccgccaaggccgcctccaccaacccgctgaagatgagccagc- cgctgggtgcggctctggcctatctcg gcgtcgaccgctgcctgccgctgttccatggctcgcagggctgcaccgccttcgggctggtcctgctggtgcg- ccatttccgcgaggcgatcccgct ccagaccacggcgatggatcaggtcgccaccatcctcggcggctacgacaatctggagcaggcgatccgcacc- atcgtcgagcgcaaccagccc gccatgatcgccgtcgccaccaccggcgtcaccgagaccaagggcgaggatatggccggacagtacacgctgt- tccgccagcgcaaccccgactt ggccgacacggccctggtcttcgccaacacccccgacttcgccggcggcttcgaggacggcttcgccgccgcg- gtcaccgcgatggtcgagcggt tggtcgaaccgtcgccggtgcgcatcccgacccaggtcaacgtgctggccggctgccatctgtcccccggcga- cgtggaggaactgcgcgacatc atcgaaggcttcggcctgtcgccgatcttcctgaccgacctgtcgctgtcgatggcgggccgccagccggccg- acttcaccgccacctcgctgggcg gcgtgaccgtcgatcagatccgcgccatgggcgcttcggccctcaccatcgtggtcggtgagcatatgcgggt- ggccggtaacgcgctggagctga agaccgacgtgcccagccatttatcaaccgcctgaccgggctggaggcgacggacaagctggtccggctgctg- atggagttgtcgggcaagccg gcgcccgcccggctgcggcgccagcgcgaaagcctggtcgatgccatgctcgacgggcatttcttctacagcc- gcaagcgcatcgccgtcgcgct ggagcccgacctgctctatgccgtcaccggcttcctcgccgacatgggggccgaggtgatcgccgcggtgtcc- ccgacgcagagcccggtgctgg agcggttgaaggccgccaccatcatggtcggcgatcattccgacgtggagacgctggcccgcgacgccgacct- gatcgctccaactcgcacggg cggcagggagccgcgcggatcggcgtggctctgcaccgcatgggcctgccgctgttcgaccggctgggggccg- gcctgcgcgtccaggtcggct accgaggcacgcgggaactgctgtgcgacatcggcaacctgttcctcgcccgcgagatggaccacgagcacgg- gcacgagagccacgaccacg gggaatcccacggctgccgaggcggatcatgcggatgcaacgccgtctga 172 atgaccgacaagctttcgcagagcgccgacaaggtcctcgaccactacaccctcttccggcagcccgaat- acgcggcgatgttcgagaagaagaag accgagttcgagtacggccattcggacgaggaagtcgcccgcgtgtccgaatggaccaagtccgaggactaca- aggcgaagaacttcgcccgtga agcggtcgtcatcaacccgaccaaggcctgccagccgatcggcgcaatgttcgccgcccagggcttcgaaggc- accctgcccttcgtccacggctc ccagggctgcgtcgcctattaccgcacccacctgacccgtcacttcaaggagccgaacagcgcggtctcctcg- tcgagacggaggacgcggcggt gttcggcggcctgaacaacatgatcgacggcctggcgaacgcctatgcgctctacaagccgaagatgatcgag- gtgatgaccacctgcatggccga agtcatcggcgacgatttgcagggcttcatcgccaatgcgaagaccaaggacaggtcccggccgacttcccgg- tcccctacgcccacaccccggc cttcgtcggcagccacatcgtcggctacgacaacatgatcaaggggatcctgaccaacttctggggtacgtcg- gagaatttcgacacacccaagacc gagcagatcaacctgatcccgggattcgacggcttcgccgtcggcaacaaccgcgaactgaagcgcatcgccg- gcgaattcggcgtgaagctgca aatcctgtccgacgtgtccgacaatttcgacacgccgatgaatggcgagtaccgcatgtatgacggcggcacc- accatcgaggagaccaaggaggc cctgcacgccaaggccaccatctccatgcaggagtacaacacgacccagaccctgcaattctgcaaggagaag- ggtcaggaagtcgccaagttcaa ctacccgatgggcgtcaccggcaccgacgagctgctgctgaagctcgccgaactgtcgggcaagccggtcccg- gccagcctgaagctggagcgc ggccgtctggtcgacgccatcgccgacagccacacccacatgcacggcaagcgcttcgccgtctatggcgacc- cggacttctgcctgggcatgtcc aagttcctgctggagctgggtgcggagccggtgcacatcctgtcgacgtcgggctccaagaaggggagaagca- ggtccagaaggtgctggacgg ctcgcccttcggcgcctcgggcaaggcccatggcggcaaggatctgtggcacctgcgttcgctgatcttcacc- gacaaggtggactacatcatcggc aacagctacggcaagtatctggagcgcgacaccaaggttccgctgatccgcctgacctacccgatcttcgacc- gccaccaccaccaccgctacccg acctggggctaccagggcgcgctgaacgtgctggtacggatcctaaccggatcttcgaggacatcgacgccaa- caccaacatcgtcggccagac cgactactcgttcgacctgatccgctga 173 atgttgacctctgatattgttggcaaattgcgctgcatcgcagcagaccccaaagcgggcatcgcaaggg- gcctcgacaccgggacgacgaagatc ggtcccgtttgggagggtgacgtgggcgacaccgtggatttcgaagcgctgcgccagcgggcggtccactccc- tgttcgaacatctggaatccatgt gcgtcagcgccgtcgccgtcgaccacaccggccgcatcgcctggatggacgagaagtacaaggctctgctggg- cgttcccgacgacccgcgcgg ccggcaggtggaggacgtcatccccaacagccagctgcgccgggtgatcgacagcggccagccgcagccgatg- gacatcatggagttcgacgac cggtccttcgtggtgacgcgcatgccgctgttcggcaccgacggttcgatcatcggcgccatcggcttcgtgc- tgttcgaccgcgccgaatatctccg cccgctggtccgcaaatacgagaagatgcaggaggagctggcccgcacccagcaggagctggcgcatgagcgc- cgcgccaaatactccttctcg cagttcctgggcgccagcgaatcgatccgcgagatcaagcggctggggcgccgcgccgcccagatggattcga- ccgtcctgctgctgggcgaaac cgggaccggcaaggagctgctggcccaggccatccattccgccagcccgcaggcgtccaagcccttcgtcggc- gtcaatgtcgccgccattccgg aaaccctgctggaggcggagttcttcggcgtcgoccccggcgccttcaccggcgccgaccgccgccaccgcga- cggcaagttccagctcgccaac ggcggcaccctgttcctcgacgagatcggcgacatgccgctgccggtgcaggccaagcttctgcgcgtgctgc- aggagcgggagatcgagccgct cggctccaacaaggtggtgcgggtcgatgtccgcatcatcgccgccaccagccgtgacctgcacgccctggtg- cgtgagaagcagttccgcgccg acctctattaccggctgaatgtggtgccgatcaccctgccgccgctgcgcgaccggccggaggacatcgagag- catcgccgaccgcatcctggaac agctggcgatccagcagggcacgccgcogcgcgagctgctggaatcggcggtgcaggtgctgcgcgactatga- ctggcccggcaatgtgcgcga gctttacaacacgctggaacgggtggtggcgctgaccgatgcgccgatcctgaccgcgccgcacatccgcagc- gtgctgcccggccagcatccgg ccggcgcgtcggccctgccgctggcggccggcgcgcggccgttgcaggaggtgctgacgccgccgagcgccac- gccatcgccgcggcgcttg aggaggcgaacggcgtcaaggcgcgggcggcgaagctgctgggcatttcgcgcgcgtcgctgtacgaacgcat-
ggtgacgctggggttggggg cgacgcagtag 174 atgccgagtcccatcgcgttctcaagccccttgccgaagcctttcgacagcgcgcaggggcgctggggat- ggaggctggcgccagcaggccgc cgcggcggagccggagacccgcgcctgggcggaagccttcgccgattcggagaccggccgggcgctgatcggg- gcggtgtgcggcaacagcc cgtatctcggccacagcctgacgcgggagttgcccttcgtcgcccgtacagtgcaggacggcttcgacgacac- cttcgccgcgctgatcgccgctct ccatgccgagcatggcgaggagaaatcgatggaccggctgatggccggcctgcgggtggcgaagcggcgggcg- gcgagctgatcgcgctggc cgacatcgccgggcgtggccgctgttccgcgtcaccggcgccctgtcggagctggcggagacgggggtgcagc- tggccgcgaatttcctgctgc gccgcgccagggaggccgggacgctgacgctgccggatccgcagcgaccgtgggtcggttcgggcctgatcgt- tttgggcatgggtaagcttggc gggcgcgaactcaactattccagcgacatcgacctgatcgtcctgtatgacgacgctgttgtgcagacgcccc- agccggacaacctcgcgcgaacct tcatcaggctcgcacgcgatcttgtccgcattatggatgaacggaccaaggacggctacgtcttccgcaccga- ccttcggcttaggcccgatcccggc gccacgccgctggcggtttccgtctccgcagccgaaatttattacggcagcgtcggtcagaactgggaacgcg- cggcgatgatcaaggcccgtccc atcgccggcgatctggaggcgggcgcgcctcctttgtccgcttcctggagcccttcgtctggcgccgcaacct- ggatttcgccgccatccaggacatcca ttcgatcaaacgccagatcaacgcccacaagggccaccgcgaggtgacggtcaacggccacgacatcaaggtc- ggccgcggcggcatccgcga gatcgagacttcgcccagacccagcagctgatcttcggcgggcgcgacccgcgcgtgcgaatcgctccgaccc- tgatggcgaacgaggcgctgc gcgacgtcggccgcgtgccgccgcagacggtggaagagcttgccggggcctatcatttcctgcgccgtgtcga- acatcgcatccagatgatcgacg accagcagacccatcgtattcccgccgacgatgccggggtggcgcatttggccaccttcctcggctatgacga- ccccgccgccttccgggcggaact gctggcgacgctggggcaggtggaggaccgctattccgagctgttcgaggaggcgccgtcgctttccggcccc- ggcaatctggtcttcaccggca ccgaccccgatccgggcacgatggagacgctgaagggcatgggcttcgccgatccggccgcgtcatcagcgtg- gtgtcggctggcatcgcgg ccgctaccgcgccacccggtcgggccgggcgcgggagctgctgacggagctgacgccggccctgctgagtgcg- ctgaccaagaccccggcccc cgattcggcgctgatgaacttcgacgatttcctcggcaagctgccggccggcgtcggtctgttctcgctgttc- gtcgccaatccctggctcctggagctg gtggcggagatcatgggcatcgcgccgcagatggcgcagacgctgtcgcgcaacccgtcgctgctcgacgccg- tgctgtcgccggacttcttcgac ccgctgccgggcaaggaggacgggctggccgacgaccacgcccgcgtgatggcgccggcccgcgatttcgagg- atgcgctgaccctgtcgcgg cgctggaccaacgaccagcgcttccgcgccggggtgcatatcctgcgcggcatcaccgatggcgaccgctgcg- gcgccttcctggccgatctggc cgacatcgtcgtccccgaccttgcccgccgggtggaggaggagttcgcccagcgccacggccatattcccggc- ggcgcctgggtggtggtggcga tgggcaagtcggcagccggcagctgaccatcacgtccgacatcgacctgatcgtcatctacgatgtggcgccc- ggccaagggggcgggggcgg tccccgcttgtcggatggtgccaagccgctgtcgcccaacgagtattacatcaagtgactcagcgtctgacca- acgccattaccgcgccgatgcgc gacggccggctctacgaggtcgacatgcggctgcgcccgtcgggcaacgccgggccgctcgccacctcgctgg- acgctttcctgaaatatcaggc gaccgatgcctggacctgggagcatatggccctgacccgcgcccgggtgatcggcggtgatgcggagctggcc- gggcgggtgtcggcagcgatc cgctcggtgctgacggcgccgcgcgatgccgaccggctgctgtgggacgtggccgacatgcggcggcggatcg- agaaggagttcgggacgacc aatgtctggaacgtcaaatacgcccgcggcggcctgatcgacatcgagttcatcgcccagtacctgaactgcg- ccatggtcacgagcggccggac atcctgcacatcggcaccgccaaggcgctgggctgcgccgcccggacgggcgcgctggcgccggaggtggcgg- aggatctggagacgacgct gcggctgtggcggcgggtgcagggctttctgcggttgaccaccgccggggtgctcgatcccaatcaggtgtcg- cccagcctgctggccgggctggt ccgcgccgcctttcctgctgactttcagggcgagcgtgagcctcgactgttgacttccccgaactggaccaca- aaatccgtgccgtcgccgcccgc gcccatggtcatttcaagaccctggtcgaggaaccggcgggccgtctggccccacccgccaccacgcctccag- cctga 175 atgaaccgtctgttccaatggccgcaccgatgatggcggttgctctgggcgcggtcggcatgccggccgc- agcccttgcccaggatccggcggctg ccgccgctgccgcggctgcggctgcggagccgccgctgctgccgcaccggcggctccggcgctgaatggcggc- gacaccgcctggatgctcat ctccaccgcgctggtgctgatgatgaccatccccggcctggcgctgttctacggcggcatggtccgcaagatg- aacgtgctgtcgacggtgatgcag agcttcgccatcacctgcctgatcagcgtcctgtggtacgtcatcggctacagcctggccttcaccggcaccg- gtgcctatgtcggcggtctcgaccg gctgttcctcaacgggctcgacttcacgaaggccttcgtgctgggcgaggcgaccgggtcgggcgtcccgacg- accatccccgagccggtcttcat gatgttccagatgacctttgcgatcatcaccccggccctgatcaccggcgccttcgccgaccgatgaagttct- cctccctgctggtcttcaccgcgctg tggtcgatcgtggtctatgcgccgatcgcccactgggtctggtacccgtcgggcttcctgttcggcctagcgt- gaggacttcgccggcggcacggt cgtgcacatcaaccgccggcgtcgccggcctggtcgccgcgctggtgatcggcaagcgcaagggctacccgaa- ggaagccttcatgccgcacaac ctggtgctgtcgctgatcggcgcctcgctgctgtgggtcggctggttcggcttcaacgccggttcggccctga- ccgccggtccgcgtgccggcatgg cgctggccgccacgcacatcgccaccgccggtgccgccatgggctggctgttcgcggagtggatcgtcaaggg- caagccgtcgatcctcggcatc atctccggcgccgtcgccggcctggtcgggtgaccccggccgccggcttcgtcgacccgacgggcgccatcgt- catcggcatcgtcgcccgcgt ggtctgcttctggtcggccaccagcctcaagcacatgctgggctatgacgacagcctggacgccttcggcgtg- cacggcgtcggcggcctgatcgg cgccatcctgaccggcgtcttcgccaagatgtcggtgtccaacagcgaaggcggcttcgcctccgtcctgcag- gccgacccgaaggccacgctgg gcctgctggaaggcaacgccgccgccgtctggatccaggtccagggcgtcctctacaccatggtctggtgcgc- catcgccaccttcgtcctgctgaa gatcgtcgatgtggtcatgggcctgcgcgtcgaagaggatgtggagcgcgacggtctcgacctcgccctgcat- ggcgagagcatccactaa 176 atggatgcggcaaagacgggtggcgacgtccttttcgtgctgatgggcgcggtgatggtgctggcgatgc- attgcggcttcgccctgctggaggtcg ggacggtccggcgcaagaatcaggtcaacgcgctggtgaagatcctgtcggacttcgccatgtcgaccatcgc- ctattttttcgtcggttatgccgtgg cctacggcatcgacttcttcgccgacgcccacacgctggtcggcaagggaagcggcgggttcgcggcctatgg- ctacgatctggtgaagttcttcttc ctggcgaccttcgccgccgcggtgccggccatcgtctcgggcggcatcgccgagcgtgctaggttctggccgc- aggccgccgccacgctggcgct gatcgcgctgttctatccattgctggaaggcacggtctggggcacccgcttcggcctgcaaagctggatggcc- gcgaccttcggccagcctttccacg acttcgccggatctgtggtggtgcatgccttcggcggctgggtggcgctgggtgccgtgctgaacctcggcaa- ccgccgcggccgctaccgtccga acggctcgctgatcgccattccgccgtcgaacatccccttcctggcgctgggcgcctgggtgctgtgcgtggg- gtggttcggcttcaacgtgatgagc gcccaggtgctggatggcgtgacgggtctggtggcgctgaactcgctgatggcgatggtcggcggcatcgtca- cctcgctggtgatcagccgcacc gatcccggcttcgtccacaacggcgcgctggccggtctggtggcggtctgcgccgggtccgacgtgatgcacc- cgctgggcgcgctggtcaccgg cggcatcgccgggctgctgttcgtctgggccttcaacaaatgccagatcgactggaagatcgacgacgtgctg- ggcgtctggccgctgcacggctg tgcggcctgaccggcggcctgctggccggcgtcttcgggcaggaggcactgggcggccttggcggcgtgtcga- tcctcagccagatcgtcggcac ggcaagcggcgccagcttcggattcgctacgggtctggcggtctaccgcctgctgcgcgtcaccgtccgcatc- cgcctcgatcccgaccaggagta caagggcgccgacttgtcgttgcaccatatcaccgcgtacccggaagaggacgcgccgaccctgtaa 177 atgaccctgaatatgatgatggattcttggttctctggagcgctttatcggcatcctgactgaagatttg- caggcttcttcccaacctggcttgcacccgt gcaggtagttgtgatgaacatcactgattcgcaggctgaatacgttaacgaattgacccgtaaactgcaaaat- gcgggcattcgtgtaaaagcagactt gagaaacgagaagattggctttaaaatccgcgagcacactttacgtcgtgtcccttatatgctggtttgtggt- gacaaagaggtcgaagccggcaaagt tgctgtgcgtacccgtcgcggtaaagacctgggtagcctggacgtaaatgatgatcgagaagctgcaacaaga- gattcgcagccgcagtcttcaac aactggaggaataaggtattaaaggcggaaaacgagttcaaacggcgcgtcccaacgtattaatggcgagatt- cgcgccacggaagttcgcttaac aggtctggaaggcgagcagcttggtattgcgatagaactcacttcacgccccgaagggggaagctgcctgacc- ctacgattcccgctatttcattcact gaccggaggttcaaaatga 178 accggataagagagaaaagtgcgacgtcggtccggttgatattgaccggcgcatccgccagctcgcccag- tttttggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccganaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgcggggaaaatgcggtgaacat gtcagctattgcgaagagtgtgccagttttgctcacgggcaaaagctgcaccagaatgggattaatgcaccag- cctggcgctttttttcgcggcacgtc ccctcgctaatgcccgtctggcgcggctttgacgctgataaggcgctgaataccgatctggatcaaggtttgt- cgggttatcgtccaaaaggtgcactc ttgcatggttataagtgcctgacatggtgtccgggcgaacgtcgccaggtggcacaaattgtcagaactacga- cacgactaaccgaccgcaggagt gtgcgatgaccctgaatatgatgatggattatggttctctggagcgctttatcggcatcctgactgaagaatt- tgcaggcttcttcccaacctggcttgca cccgtgcaggtagttgtgatgaacatcactgattcgcaggctgaatacgttacgaattgacccgtaaactgca- aaatgcgggcattcgtgtaaaagca gacttgagaaacgagaagattggctttaaaatccgcgagcacactttacgtcgtgtcccttatatgctggttt- gtggtgacaaagaggtcgaagccggc aaagttgctgtgcgtacccgtcgcggtaaagacctgggtagcctggacgtaaatgatgttatcgagaagctgc- aacaagagattcgcagccgcagtct tcaacaactggaggaataaggtattaaaggcggaaaacgagttcaaacggcgcgtcccaatcgtattaatggc- gagattcgcgccacggaagttcgc ttaacaggtctggaaggcgagcagcttggtattgcgatagaactcacttcacgccccgaagggggaagctgcc- tgaccctacgattcccgctatttcat tcactgaccggaggttcaaaatgacccagcgaaccgagtcgggtaataccgtctggcgcttcgatttatccca- gcagttcaccgcgatgcagcggata agcgtggttctcagcgggcgaccgaggttgaacagacactccagcaggtgctgtgcgtattgcacaatgacgc- ctttttgcagcacggcatgatctg tctgtacgacagccagcaggcgattttgactattgaagcgttgcaggaagccgatcagcagttgatccccggc- agctcgcaaattcgctaccgtccgg gtgaagggaggtcgggacggtgctttcgcaggggcaatcgttagtgctggcgcgtgtggctgacgatcagcgc- tttcttgaccgcctgggactgtat gattacaacctgccgtttatcgccgtgccgctgatagggccggatgcgcagacttttggcgtgctgacggcgc- aaccgatggcgcgttacgaagagc ggttacccgcctgcacccgctttctggaaacggtc 179 tccctgtgcgccgcgtcgccgatggtgaccagcaactggcgcgctacccgatcctgctcgagaactgctc- gacccgaacacgctctatcaaccga cggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagaggatgaagagcagca- actggaggcgctacggcagttta agcaggcgcagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatgaaagtgagcgatca- cttaacctggctggcggaagcga ttatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccgacgcatctgcacga- tcgcgaagggcgcggtttcgccg tggtcggttacggcaaacttacggctgggaattaggttacagctccgatctggatctggtgttcctgcacgac- tcccccatggatgtgatgaccgatg gcgagcgtgaaatcgatggcccccagttctatttgcgcctcgcgcagcgcgtgatgcacctgttcagcacgcg- cacgtcgtccggcattctttatgaag tcgatgcgcgtttgcgcccgtccgggcggccggaatgctggtgaccactgcggaagcgttcgccgattatcaa- aaaaatgaagcctggacatggg agcatcaggcgctggcgcgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcg- ccgcgatatcctgatgacctcccg cgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatcttggtaacaagcac- aaagaccgtttcgatctgaaagc cgatgaaggcggatcaccgatattgagtttatcgctcagtatctggtgctgcgctttgcccatgagaagccga- aactgacgcgctggtcggataatgg cgcatcctcgaagggctggcgcaaaacggcatcatggatgagcaggaagcgcaggcattgacgctggcgtaca- ccacgttgcgtgatgagctgca ccacctggcgctgcaagagctgccaggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttatcaaa- accagctgggacaagtggctggtg gaaccgtgcgccccggcgtaa 180 taaagcgagcgctcacttacgtgatctgttgacgcagtccgaagcgaccattacttcagccgtttcagca- gatacggcggtgtggagtgcgcaatcag ccctggcgaaactggtgctcaccgagtggttagtgacgcagggctggcgaaccttccttgatgaaaaagcgca- ggctaagtttgccgactcctttaaa cgctttgctgacgttcatctgtcacgcagcgccgccgagctgaaaaaagcctttgcccagccgctgggcgaca- gctatcgcgaccagttaccgcggc tggcgcgtgatatcgacagcgcgttattgctggccggacattacgatcgcgcgcgcgccgtggagtggctgga- aaactggcaggggcttcagcacg ctattgaaacgcgccagagagttgaaatcgaacatttccgtaataccgccattacccaggagccgttctggtt- gcacagggaaaacgttaacgaaag gatatttcgcatgtccctgtgcgccgcgtcgccgatggtggccagccaactggcgcgctacccgatcagctcg- atgaactgctcgacccgaacacg ctctatcaaccgacggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagagg- atgaagagcagcaactcgaggc gctacggcagtttaagcaggcgcagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatg- aaagtgagcgatcacttaacctgg ctggcggaagcgattatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccga- cgcatctgcacgatcgcgaagg gcgcggtttcgccgtggtcggttacgccaaacttggcggctgggaattaggttacagctccgatctggatctg- gtgttcctgcacgactgccccatgga tgtgatgaccgatggcgagcgtgaaatcgatggccgccagttctatttgcgcctcgcgcagcgcgtgatgcac- ctgttcagcacgcgcacgtcgtccg gcattctttatgaagtcgatgcgcgtttgcgcccgtccggcgcggccggaatgctggtgaccactgcggaagc- gttcgccgattatcaaaaaaatgaa gcctggacatgggagcatcaggcgctggcgcgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaat- ttgacgccattcgccgcgatatc ctgatgacctcccgcgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatc- ttggtaacaagcacaaagaccgt ttcgatctgaaagccgatgaaggcggtatcaccgatattgagtttatcgctcagatctggtgctgcgctttgc- ccatgagaagccgaaactgacgcgct ggtcggataatgtgcgcatcctcgaagggctggcgcaaaacggcatcatggatgagcaggaagcgcaggcatt- gacgctggcgtacaccacgttg cgtgatgagctgcaccacctggcgctgcaagagctgccaggacatgtggcgactctcctgttttgtcgccgag- cgtgcgcttatcaaaaccagctggga caagtggctggtggaaccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaattttgtatctctcagga- gacaggaatgaaagttacgctgcca gagttcaatcaagccggtgtcatggtggtgggtgatgtgatgctggatcgctactggtacggcccaaccagcc- gcatttctccggaagcgccagttcc ggttgttaaagtcgatactattgaagagcgaccgggcggtgcggcaaacgtggcgatgaacattgcctcgctg- ggcgcaacggcgcgtctggttggc ctgactggcattgatgatgcggcgcgcgcgctgagcaaagcgctggcggatgttaatgttaaatgtgacttcg- tttctgttccgactcaccccaccatca ctaagctgcgcgtgctgtcgcgtaaccagcaactgattcgc 181 atgaccctgaatatgatgatggatgccagccgttctgtaataataaccggacaattcggactgattaaaa- aagcgccctcgcggcgctttttttatattctc gactccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccgatg- gaagctcgctgttttaacacgcgtttttaa ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatattct- caacctaaaaaagtttgtgtaatacttgtaac gctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaactca- cttcacgccccgaagggagaagctgc
ctgaccctacgattcccgctatttcattcactgaccggaggttcacaatga 182 accggataagagagaaaagtgtcgacgtcggtccggttgatattgaccggcgcatccgccagctcgccca- gtttttggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgcggggaaaatgcggtgaacat gtcagctattgcgaagactgtgccagttttgctcacgggcaaaagctgcaccagaatgggtattaatgcacca- gcctggcgctttttttcgcggcacgtc ccctcgctaatgcccgtctggcgcggctttgacgctgataaggcgctgaataccgatctggatcaaggttttg- tcgggttatcgtccaaaaggtgcactc tttgcatggttataagtgcctgacatggtgtccgggcgaacgtcgccaggtgccacaaattgtcagaactacg- acacgactaaccgaccgcaggagt gtgcgatgaccctgaatatgatgatggatgccagccgttctgtaataatcaccggacaattcggactgattaa- aaaagcgccctcgcggcgctttttttat attctcgactccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaat- ccgatggaagctcgctgttttaacacgcgtt ttttaaccttttattgaaagtcgggcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaat- attctcaacctaaaaaagtttgtgtaatactt gtaacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgca- actcacttcacgccccgaagggggaa gctgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcg- ggtaataccgtctggcgcttcgattta tcccagcagttcaccgcgatgcagcggataagcgtggttctcagccggccgaccgaggttgaacagacactcc- agcaggtgctgtgcgtattgcaca atgacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgactattgaagcgtt- gcaggaagccgatcagcagttgatccc cggcagctcgcaaattcgctaccgtccgggtgaagggctggtcgggacggtgctttcgcaggggcaatcgtta- gtgctggcgcgtgtggctgacgat cagcgctttcttgaccgcctgggactgtatgattacaacctgccgtttatcgccgtgccgctgatagggccgg- atgcgcagacttttggcgtgctgacg gcgcaaccgatggcgcgttacgaagagcggttacccgcctgcacccgctttctggaaacggtc 183 tccctgtgcgccgcgtcgccgatggtggccagccaactggcgcgctacccgatcctgctcgatgaactgc- tcgacccgaacacgctctatcaaccga cggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagaggatgaagagcagca- actggaggcgctacggcagttta agcaggcgcagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatgaaagtgagcgatca- cttaacctggctggcggaagcga ttatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccgacgcatctgcacga- tcgcgaagggcgcggtttcgccg tggtcggttacggcaaacttggcggctgggaattaggttacagctccgatctggatctggtgttcctgcacga- ctgccccatggatgtgatgaccgatg gcgagcgtgaaatcgatggcccccagttctatttgcgcctcgcgcagcgcgtgatgcacctgttcagcacgcg- cacgtcgtccggcattctttatgaag tcgatgcgcgtttgcgcccgtccggcgggccggaatgctggtgaccactgggaagcgttcgccgattatcaaa- aaaatgaagcctggacatggg agcatcaggcgctggcgcgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcg- ccgcgatatcctgatgacctcccg cgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatcttggtaacaagcac- aaagaccgtttcgatctgaaagc cgatgaaggcggtatcaccgatattgagtttatcgctcagtatctgggctgcgctttgcccatgagaagccga- aactgacgcgctggtcggataatgtg cgcatcctcgaagggctggcgcaaaacggcatcatggatgagcagcaagcgcaggcattgacgctggcgtaca- ccacgttgcgtgatgagctgca ccacctggcgctgcaagagctgccaggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttatcaaa- accagctgggacaagtggctggtg gaaccgtgcgccccggcgtaa 184 taaagcgagcgctcacttacgtgatctgttgacgcagtccgaagcgaccattacttcagccgtttcagca- gatacggcggtgtggagtgcgcaatcag ccctggcgaaactggtgctcaccgagtggttagtgacgcagggctggcgaaccttccttgatgaaaaagcgca- ggctaagtttgccgactcctttaaa cgctttgctgacgttcatctgtcactcagcgccgccgagctgtaaaaagcctttgcccagccgctgggcgaca- gctattgcgaccagttaccgcggc tggcgcgtgatatcgacagcgcgttattgctggccggacattacgatcgcgcgcgcgccgtggagtggctgga- aaactggcaggggcttcagcacg ctattgaaacgcgccagagagttgaaatcgaacatttccgtaataccgccattacccaggagccgttctggtt- gcacagcggaaaacgttacgaaag gatatttcgcatgtccctgtgcgccgcgtcgccgatggtggccagccaactggcgcgctacccgatcctgctc- gatgaactgctcgacccgaacacg ctctatcsaccgacggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagagg- atgaagagcagcaactggaggc gctacggcagtttaagcaggcccagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatg- aaagtgagcgatcacttaacctgg ctggcggaagcgattatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccga- cgcatctgcacgatcgcgaagg gcgcggtttcgccgtggtcggttacggcaaacttggcggctgggaattaggttacagctccgataggatctgg- tgttcctgcacgactgccccatgga tgtgatgaccgatggccagcctgaaatcgatggccgccagttctatttgcgcctcgcgcagcgcgtgatgcac- ctgttcagcacgcgcacgtcgtccg gcattctttatgaagtcgatgcgcgtttgcgcccgtccggcgcggccggaatgctggtgaccactgcggaagc- gttcgccgattatcaaaaaaatgaa gcctggacatgggagcatcagccgtagcgcgtgcgcgcgtggtgtaccgcgatccgcaactgaccgccgaatt- tgacgccattcgccgcagatatc ctgatgacctcccgcgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatc- ttggtaacaagcacaaagaccgt ttcgatctgaaagccgatgaaggcggtatcaccgatattgagtttatcgctcagtatctggtgctgcgctttg- cccatgagaagccgaaactgacgcgct ggtcggataatgtgcgcatcctcgaagggctggcgcaaaacggcatggatggatgagcaggaagcgcaggcat- tgacgctggcgtacaccacgttg cgtgatgagctgcaccacctggcgctgcaagagctgccaggacatgtggcgctctcctgttttgtcgccgagc- gtgcgatatcaaaaccagctggga caagtggctggtggaaccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaattttgtatctctcagga- gacaggaatgaaagttacgctgcca gagttcaatcaagccggtgtcatggtggtgggtgatgtgatgctggatcgctactatacggcccaaccagocg- catttctccggaagcgccagcc ggttgttaaagtcgatactattgaagagcgaccgggcggtgcggcaaacgtggcgatgaacattgcctcgctg- ggcgcaacggcgcgtctggttggc ctgactggcattgatgatgcggcgcgcgcgctgagcaaagcgctggcggagttaatgttaaatgtgacttcgt- ttctgttccgactcaccccaccatca ctaagctgcgcgtgctgtcgcgtaaccagcaactgattcgc 185 atgaccctgaatatgatgatggatgccagccgttctgaataataaccggacaattcggactgattaaaaa- agcgccctcgcggcgctttttttatattctc gactccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccgatg- gaagacgctgattttaacacgcgttttttaa ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttattgtggattcccatcaaaaaaatattc- tcaacctaaaaaagtttgtgtaatacttgtaac gctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaactca- cttcacgccccgaagggggaagctgc ctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatga 186 accggataagagagaaaagtgtcgacgtcggtccggttgatattgaccggcgcatccgccagctcgccca- gtttttggtggatctgtttggcgattttgc gggtcttgccggtgtcgctgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgcggggaaaatgcggtgaacat gtcagctattgcgaagagtgtgccagttttgctcacgggcaaaagctgcaccagaatgggtattaatgcacca- gcctggcgctttttttcgcggcacgtc ccctcgctaatgcccgtctggcgcggctttgacgctgataaggcgctgaataccgatctggatcaaggttttg- tcgggttatcgtccaaaaggtgcactc tttgcatggttataagtgcctgacatggtgtccgggcgaacgtcgccaggtggcacaaattgtcagaactacg- acacgactaaccgaccgcaggagt gtgcgatgaccctgaatatgatgatggatgccagccgttctgtaataataaccggacaattcggactgattaa- aaaagcgccctcgcggcgctttttat attctcgactccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaat- ccgatggaagctcgctgtttaacacgcgtt ttttaaccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcagaaaaa- tattctcaacctaaaaaagtttgtgtaatactt gtaacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgca- actcacttcacgccccgaagggggaa gctgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcg- ggtaataccgtctggcgcttcgattta tcccagcagtcaccgcgatgcagcggataagcgtggttctcagccgggcgaccgaggttgaacagacactcca- gcaggtgctgtgcgtattgcaca atgacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgactattgaagcgtt- gcaggaagccgatcagcagttgatccc cggcagctcgcaaattcgctaccgtccgggtgaagggctggtcgggacggtgctttcgcaggggcaatcgtta- gtgaggcgcgtgtggctgacgat cagcgctttcttgaccgcctgggactgtatgattacaacctgccgtttatcgccgtgccgctgatagggccgg- atgcgcagacttttggcgtgctgacg gcgcaaccgatggcgcgttacgaagagcggttacccgcctgcacccgctttctggaaacggtc 187 tccctgtgcgccgcgtcgccgatggtggccagccaactggcgcgctacccgatcctgctcgatgaactgc- tcgacccgaacacgctctatcaaccga cggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagaggatgaagagcagca- actggaggcgctacggcagttta agcaggcgcagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatgaaagtgagcgatca- cttaacctggctggcggaagcga ttatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccgacgcatctgcacga- tcgcgaagggcgcggtttcgccg tggtcggttacggcaaacttggcggctgggaattaggttacagctccgatctggatctggtgttcctgcacga- ctgccccatggatggatgaccgatg gcgagcgtgaaatcgatggcccccagttctatttgcgcctcgcgcagcgcgtgatgcacctgttcagcacgcg- cacgtcgtccggcattctttatgaag tcgatgcgcgtttgcgcccgtccggcgcggccggaatgctggtgaccactgcggaagcgttcgccgattatca- aaaaaatgaagcctggacatggg agcatcaggcgctggcgcgtcgcggtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgcc- gcgatatcctgatgacctcccg cgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatcttggtaacaagcac- aaagaccgtttcgatctgaaagc cgatgaaggcggtatcaccgatattgagtttatcgctcagtatctggtgctgcgctttgcccatgagaagccg- aaactgacgcgctggtcggataatgtg cgcatcctcgaagggctggcgcaaaacggcatcatggatgagcaggaagcgcaggcattgacgctggcgtaca- ccacgttgcgtgatgagctgca ccacctggcgctgcaagagagccaggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttatcaaaa- ccagctgggacaagtggctggtg gaaccgtgcgccccggcgtaa 188 taaagcgagcgctcacttacgtgatctgttgacgcagtccgaagcgaccattacttcagccgtttcagca- gatacggcggtgtggagtgcgcaatcag ccctggcgaaactggtgctcaccgagtggttagtgacgcagggctggcgaaccttccttgatgaanagcgcag- gctaagtttgccgactcctttaaa cgctttgctgacgttcatctgtcacgcagcgccgccgagctgaaaaaagcctttgcccagccgctgggcgaca- gctatcgcgaccagttaccgcggc tggcgcgtgatatcgacagcgcgttattgctggccggacattacgatcgcgcgcgcgccgtggagtggctgga- aaactggcaggggcttcagcacg ctattgaaacgcgccagagagttgaaatcgaacatttccgtaataccgccattacccaggagccgttctggtt- gcacagcggaaaacgttaacgaaag gatatttcgcatgtccctgtgcgccgcgtcgccgatggtggccagccaactggcgcgctacccgatcctgctc- gatgaactgctcgacccgaacacg ctctatcaaccgacggcgatgaacgcctatcgcgatgaactgcgacaatacctgttgcgcgtgccggaagagg- atgaagagcagcaactggaggc gctacggcagtttaagcaggcccagttgttgcgcgtagcggcggcggatatcgccggtacgttacccgtcatg- aaagtgagcgatcacttaacctgg ctggcggaagcgattatcgatgcggtggtgcagcaagcctggaaccagatggtggcgcgttacggccagccga- cgcatctgcacgatcgcgaagg gcgcggtttcgccgtggtcggttacggaaacttggcggctgggaattaggttacagctccgatctggatctgg- tgttcctgcacgactgccccatgga tgtgatgaccgatggcgagcgtgaaatcgatggccgccagttctatttgcgcctcgcgcagcgcgtgatgcac- ctgttcagcacgcgcacgtcgtccg gcattctttatgaagtcgatgcgcgtttgcgcccgtccggcgcgccggaatgctggtgaccactgcggaagcg- ttcgccgattatcaaaaaaatgaa gcctggacatgggagcatcaggctggcgcgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaattt- gacgccattcgccgcgatatc ctgatgacctcccgcgatgccgctaccctgcaaaccgaagtgcgggaaatgcgtgagaaaatgcgcgcccatc- ttggtaacaagcacaaagaccgt ttcgatctgaaagccgatgaaggcggtatcaccgatattgagtttatcgctcagtatctggtgctgcgctttg- cccatgagaagccgaaactgacgcgct ggtcggataagtgcgcatcctcgaagggctggcgcaaaacggcatcatggatgagcacgaagcgcaggcattg- acgctggcgacaccacgttg cgtgatgagagcaccacctggcgctgcaagagctgccaggacatgtggcgactcctgttttgtcgccgagcgg- cgcttatcaaaaccagctggga caagtggctggtggaaccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaattttgtatctctcagga- gacaggaatgaaagttacgctgcca gagttcaatcaagccggtgtcatggtggtgggtgatgtgatgctggatcgctactggtacggcccaaccagcc- gcatttctccggaagcgccagttcc ggttgttaaagtcgatactattgaagagcgaccgggcggtgcggcaaacgtggcgatgaacattgcctcgctg- ggcgcaacggcgcgtctggttggc ctgactggcattgatgatgcggcgcgcgcgctgagcaaagcgctggcggatgttaatgttaaatgtgacttcg- tttctgttccgactcaccccaccatca ctaagctgcgcgtgctgtcgcgtaaccagcaactgattcgc 189 atgaccctgaatatgatgatggatgccagccgcgtcaggttgaacgtaaaaaagtcggtctgcgcaaagc- acgtcgtcgtccgcagttctccaaacgtt aattggtttctgcttcggcagaacgattggcgaaaaaacccggtgcgaaccgggtttttttatggataaagat- cgtgttatccacagcaatccattgattat ctcttctttttcagcatttccagaatcccctcaccacaaagcccgcaaaatctggtaaactatcatccaattt- tctgcccaaatggctgggattgttcattttt gtttgcatacaacgagagtgacagtacgcgcgggtagttaactcaacatctgaccggtcgataactcacttca- cgccccgaagggggaagctgcctg accctacgattcccgctatttcattcactgaccggaggttcaaaatga 190 accggataagagagaaaagtgtcgacgtcggtccggttgatattgaccggcgcatccgccagctcgccca- gtttttggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgcggggaaaatgcgggaacat gtcagctattgcgaagagtgtgccagttttgctcacgggcaaaagctgcaccagaatgggtattaatgcacca- gcctggcgctttttttcgcggcacgtc ccctcgctaatgcccgtctggcgcggctttgacgctgataaggcgctgaataccgatctggatcaaggttttg- tcgggttatcgtccaaaaggtgcactc tttgcatggttataagtgcctgacatggtgtccgggcgaacgtcgccaggtggcacaaattgtcagaactacg- acacgactaaccgaccgcaggagt gtgcgatgaccctgaatatgatgatggatgccagccgcgtcaggttgaacgtaaaaaagtcggtctgcgcaaa- gcacgtcgtcgtccgcagttctcca aacgttaattggtttctgcttcggcagaacgattggcgaaaaaacccggtgcgaaccgggtttttttatggat- aaagatcgtgttatccacagcaatccatt gattatctcttctttttcagcatttccagaatcccctcaccacaaagcccgcaaaatctggtaaactatcatc- caattttctgcccaaatggctgggattgttc attttttgtttgccttacaacgagagtgacagtacgcgcgggtagttaactcaacatctgaccggcgataact- cacttcacgccccgaagggggaagct gcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcgggt- aataccgtctggcgcttcgatttatcc cagcagttcaccgcgatgcagcggataagcgtggttctcagccgggcgaccgaggttgaacagacactccagc- aggtgctgtgcgtattgcacaat gacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgactattgaagcgttgc- aggaagccgatcagcagttgatcccc ggcagctcgcaaattcgctaccgtccgggtgaagggctggtcgggacggtgctttcgcaggggcaatcgttag- tgctggcgcgtgtggctgacgatc agcgctttcttgaccgcctgggactgtatgattacaacctgccgtttatcgccgtgccgctgatagggccgga- tgcgcagacttttggcgtgctgacgg cgcaaccgatggcgcgttacgaagagcggttacccgcctgcacccgctttctggaaacggt
191 atggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgcgccacc- cgctgctgctggatgagctgctggat cccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctgctgcgcgtgc- cggaagaggatgaagagcagca gctggaggcgttgcgccagtttaagcaggcgcagcagctgcatatcgcggcggcggatatcgctggtaccctg- ccggtgatgaaggtcagcgatca cttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggtcgctcgctac- ggccagccgacccacctgcacg atcgccagggtcgcggcttcgccgtcgtcggctacggtaagcttggcggctgggagctgggctacagctccga- tctcgatctggtgttcctccatgac tgcccggcggaggtgatgaccgacggcgagcgggagattgacgtccgtcagttctacctgcggctggcccagc- ggatcatgcacctgttgcagcac ccgcacctcgtccggtattctctacgaagtggacgcccggctgcgtccttctggcgcggcggggatgctgtgg- tcaccaccgccgacgcgtttgctgact atcagcagaacgaagcctggacgtgggaacatcaggcgctggtgcgcgcccgcgtggtctatggcgacccggc- gctgcaggcgcgctttgacgc cattcgtcgcgatatcctgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcgagatgcgcgag- aagatgcgcgcccaccttggca acaaacatcccgatcgttttgatatcaaagccgatgccggcgggatcaccgatattgaatttattactcagta- tctggtcctacgctatgccagtgacaag ccgaagctgacccgctggtctgacaacgtgcgtattcttgagctgctggcgcagaacgacatcatggacgagg- aggaggcgcgcgccttaacgcat gcgtacaccaccttgcgtgatgcgctccatcacctggccctgcaggaccagccgggacacgtggcgccagagg- ccttcagccgggagcgtcagca ggtcagcgccagctggcagaagtggctgatggcttaa 192 cgtaaggcgaccacccagctccgcgcgttgctgaacgacgctgaagccgttctgctggccgcggacaccg- ccgacgaggcgttattccgcaccga ggtcgtcggcgccaaactggccctgactgaatggctggtccagcgcggctggcgtccgttcctcaacgaggca- ggagagaaaaaatagccggat cgttcaaacggtttgccgatattaacctctcgcgggtggcggccgagctgcgcagcgccgtgcagcatctggc- ggttgaagatgccgccgaccagtt gccgaagctgcccgcgacatcgacagcgtccagctgctggcgggcgcctatggcgacgccgtcgcgccgtggc- tggagaactggcaggagctt caccgtgcaatagcacatgacgatcgcagcgtctttgaatatttccgtcgccaggcgctggctgccgagccgt- tctggctgcatagtggaaaacgata atttcaggccagggagcccttatggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgcca- gccagctggcgcgccacccgctg ctgctggatgagctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgc- gccagtacctgctgcgcgtgccg gaagaggatgaagagcagcagctggaggcgttgcgccagtttaagcaggcgcagcagctgcatatcgcggcgg- cggatatcgctggtaccctgcc ggtgatgaaggtcagcgatcacttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgg- gggcagatggtcgctcgctacgg ccagccgacccacctgcacgatcgccagggtcgcggcttcgccgtcgtcggctacggtaagcttggcggctgg- gagctgggctacagctccgatct cgatctggtgttcaccatgactgcccggcggaggtgatgaccgacggcgagcgggagattgacggccgcagtt- ctacctgcggctggcccagcg gatcatgcacctgttcagcacccgcacctcgtccggtattctctacgaagtggacgcccggctgcgtccttct- ggcgcggcggggatgctggtcacca ccgccgacgcgtttgctgactatcagcagaacgaagcctggacgtgggaacatcaggcgctggtgcgcgcccg- cgtggtctatggcgacccggcg ctgcaggcgcgctttgacgccattcgtcgcgatatcctgaccaccccccaaggggatgaccctgcagaccgag- gttcgcgagatgcgcgagaa gatgcgcgcccaccttggcaacaaacatcccgatcgttttgatatcaaagccgatgccggcgggatcaccgat- attgaatttattactcagtatctggtcc tacgctatgccagtgacaagccgaagctgacccgctggtctgacagcgtgcgtattcttgagctgctggcgca- gaqacgacatcatgcacgaggagga ggcgcgcgccttaacgcatgcgtacaccaccttgcgtgatgcgctccatcacctggccctgcaggagcagccg- ggacacgtggcgccagaggcct tcagccgggagcgtcagcaggtcagcgccagctggcagaagtggctgatggcttaactataaaatcgggtgtg- ctatatcgcgcgcaaagtttgcgt ctcgcaggagagagtcatgaaagtaacgctgccggagtttgaacgtgcaggagtgttggtggtgggtgatgtg- atgctggaccgctactggtacggc cccaccagtcgtatttcccggaagccccggtgccggtggtgaaggtggaaaatatcgaagaacgtcctggcgg- cgcggcaaacgtagcgatgaa catcgcctccctgggggcaacgtcgcgcctggtgggattgaccgggattgatgacgctgcccgcgctgagcca- ggcgctggccaatgtgaatgt gaagtgcgacttcgtctccgtcccgactcacccgaccatcaccaagctgcgggtgctgtcgcgcaatcagcag- ctgatccgcctcgactttgaagag ggcttctccggcgtggatccgcagccgatgcatgagcgcattcagcaggcgctgggagccattggcgcactgg 193 atgaccctgaatatgatgctagaagcgtcaggtaccggtcatgattcaccgtgcgattctcggttccctg- gagcgcttcattggcatcctgaccgaagag ttcgctggcttcttcccaacctggattgcaccagtgcaggtagtggtcatgaatattaccgattctcaggctg- aatacgttaacgaattgacgcgtaaacta caaaatgcgggcattcgtgtaaaagcagacttgagaaatgagaagattggctttaaaatccgcgagcacactt- tacgtcgtgtcccgtatatgttggtct gtggcgacaaagaagtcgaagccggcaaagtggccgtgcgcacccgtcgcgggaaagacctcggcagcatgga- cgtaagtgaagtgattgagaa gctgcaacaagagattcgcagccgcagtcttcaacaactggaggaataaggtattaaaggcggaaaacgagtt- caaacggcacgtccgaatcgtatc aatggcgagattcgcgccctggaagttcgcgccattgagctggcttcccgaccgcagggcggcacagcctgac- cctgcgtttcccgctgtttaacac cctgaccggaggtgaagcatga 194 ggccgtcgcccagcgtcggcgtccccaacagcagggcgggtaggccagcaggtccgccagcgtggcgcgg- ttaatattgaccggggcggcgg cggcctcccccagctgcttgtggatcattttcgcgatcttgcgggtttttaccggtatcggtaccaaagaaaa- tgccaatgttcgccatagtacgctcctgt cggnatggtgttgaaaaaaggaatgacgacagaggtattgcgaaggctgtgccaggttgccctgcaccgcgac- ggcccatccctgccccatcagga tcgcttcgcatcacgatgccgcgcgccaaaggcgcacccggcggggcgaaaggtaaaaatccgtgaattttcc- ccctgtcggatcaatgtttcgcgtg gtcgttccgataagggcgcacactttgcatggttatccgggttcggcttaccccgccgagttttgcgcacggt- gtcggacaatttgtcataactgcgaca caggagtttgcgatgaccctgaatatgatgctagaagcgcaggtaccggtcatgattcaccgtgcgattctcg- gttccctggagcgcttcattggcatc ctgaccgaagagttcgctggcttcttcccaacctggattgcaccagtgcaggtagtggtcatgaatattaccg- attctcaggctgaatacgttaacgaatt gacgcgtaaactacaaaatgcgggcattcgtgtaaaagcagacttgagaaatcagaagattggctttaaaatc- cgcgagcacactttacgtcgtgtccc gtatatgttggtctgtgccgacaaagaagtcgaagccggcnaagtggccgtgcgcacccgtcgcgggaaagac- ctcggcagcatggacgtaagtg aagtgattgagaagctgcaaccagagattcgcagccgcagtcttcaacaactggaggaataaggtattaaagg- cggaaaacgagttcaaacggcac gtccgaatcgtatcaatggcgagattcgcgccctggaagttcgcgccattgagctggcttcccgaccgcaggg- cggcacctgcctgaccctgcgtttc ccgctgtttaacaccctgaccggaggtgaagcatgatccctgaatccgacccggacaccaccgtcagacgctt- cgacctctctcagcagttcaccgcc atgcagcggataaggtggtgagagccgggccaccgaggccagcaaaacgctgcaggaggtgctcagcgtatta- cacaacgatgcctttatgcag cacgggatgatctgcctctacgacagcgagcaggagatcctcagtatcgaagcgctgcagcaaaccggccagc- agcccctccccggcagcacgc agatccgctatcgccccggcgagggactggtggggaccgtgctggcccaggggcagtcgctggtgctgccccg- ggtcgccgacgatcagcgtttt ctcgaccgcctgagcctctacgattacgatctgccgtttatcgccgtaccgttgatggggcccaacgcccggc- caataggggtgctggcggcccagc cgatggcgcgccaggaagagcggctgccggcctgcacccgttttctcgaaaccgtc 195 atggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgcgccacc- cgctgctgaggatgagctgctggat cccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctgctgcgcgtgc- cggaagaggatgaagagcagca gctgcatatcgcggcggcggatatcgctggtaccctgccggtgatgaaggtcagcgatcacttaacctggctt- gccgaagcgatcctcgacgcggtg gtgcagcaggcatgggggcagatggtcgctcgctacggccagccgacccacctgcacgatcgccagggtcgcg- gcttcgccgtcgtcggctacg gtaagcttggcggctgggagctgggctacagctccgatctcgatctggtgttcctccatgactgcccggcgga- ggtgatgaccgacggcgagcggg agattgacggccgtcagttctacctgcggctggcccagcggatcatgcacctgttcagcacccgcacctcgtc- cggtattctctacgaagtggacgcc cggctgcgtccttaggcgcggcggggatgctggtcaccaccgccgacgcgtttgctgactatcagcagaacga- agcctggacgtgggaacatcag gcgctggtgcgcgcccgcgtggtctatggcgacccggcgctgcaggcgcgctttgacgccattcgtcgcgata- tcctgaccaccccgcgggaggg gatgaccctgcagaccgaggttcgcgagatgcgcgagaagatgcgcgcccaccttggcaacaaacatcccgat- cgttttgatatcaaagccgatgcc ggcgggatcaccgatattgaatttattactcagtatctggtcctacgctatgccagtgacaagccgaagctga- cccgctggtctgacancgtgcgtattct tgagctgctggcgcagaacgacatcatggacgaggaggaggcgcgcgccttaacgcatgcgtacaccaccttg- cgtgatgcgctccatcacctggc cctgcaggagcagccgggacacgtggcgccagaggccttcagccgggagcgtcagcaggtcagcgccagctgg- cagaagtggctgatggcttaa 196 cgtaaggcgaccacccagctccgcgcgttgctgaacgacgctgaagccgttctgctaccgcggacaccgc- cgacgaggcgttattccgcaccga ggtcgtcggcgccaaactggccctgactgaatggctggtccagcgcggctggcgtccgttcctcaacgaggca- ggagagaaaaaaatagccggat cgttcaaacggtttgccgatattaacctctcgcgggtggcggccgagctgcgcagcgccgtgcagcatctggc- ggttgaagatgccgccgaccagtt gccgaagctgtcccgcgacatcgacagcgtccagctgctggcgggcgcctatggcgacgccgtcgcgccgtgg- ctggagaactggcaggagctt caccgtgcaatagcacatgaccatcgcagcgtctttgaatatttccgtcgccaggcgctggctgccgagccgt- tctggctgcatagtggaaaacgata atttcaggccagggagcccttatggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgcca- gccagctggcgcgccacccgctg ctgctggatgagctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagagcg- ccagtacctgctgcgcgtgccg gaagaggatgaagagcaggctgcatatcgcggcggcggatatcgctggtaccctgccggtgatgaaggtcagc- gatcacttaacctggcttgcc gaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggtcgctcgctacggccagccgacccacc- tgcacgatcgccagggtcgcg gcttcgccgtcgtcggctacggtaagcttggcggctgggagctgggctacagctccgatctcgatctggtgtt- cctccatgactgcccggcggaggtg atgaccgtcggcgagcgggagattgacggccgtcagttctacctgcggctggcccagcggatcatgcacctgt- tcagcacccgcacctcgtccggt attctctacgaagtggacgcccggctgcgtccttctggcgcggcggggatgctggtcaccaccgccgacgcgt- ttgagactatcagcagaacgaag cctggacgtgggattcatcaggcgctggtgcgcgcccgcgtggtctatggcgacccggcgctgcaggcgcgct- ttgacgccattcgtcgcgatatcc tgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcgagatgcgcgagaagatgcgcgcccacct- tggcaacaaacatcccgatcg ttttgatatcaaagccgatgccggcgggatcaccgatattgaatttattactcagtatctggtcctacgctat- gccagtgacaagccgaagctgacccgct ggtctgacaacgtgcgtattcttgagctgctggcgcagaacgacatcatggacgaggaggaggcgcgcgcctt- aacgcatgcgtacaccaccttgc gtgatgcgctccatcacctggccctgcaggagcagccgggacacgtggccccagaggccttcagccgggagcg- tcagcaggtcagcgccagctg gcagaagtggctgatggcttaactataaaatcgggtgtgctattatcgcgcgcaaagtttgcgtctcgcagga- gagagtcatgaaagtaacgctgccg gagtttgaacgtgcaggagtgttggtggtgggtgatgtgatgctggaccgctactggtacggccccaccagtc- gtatttccccggaagcccoggtgcc ggtggtgaaggtggtaaatatcgaagaacgtcctggcggcgcggcaaacgtagcgatgaacatcgcctccctg- ggggcaacgtcgcgcctggtgg gattgaccgggattgatgacgctgcccgcgcgctgagccaggcgctggccaatgtgaatgtgaagtgcgactt- cgtctccgtcccgactcacccgac catcaccaagctgcgggtgctgtcgcgcaatcagcagctgatccgcctcgactttgaagagggcttctccggc- gtggatccgcagccgatgcatgag cgcattcagcaggcgctgggagccattggcgcactgg 197 atgaccctgaatataagctcgacgccgtcctcgcagtaccattgcaaccgactttacagcaagaagtgat- tctggcacgcatggaacaaattcttgcca gtcgggctttatccgatgacgaacgcgcacagcttttatatgagcgcggagtgttgtatgatagtctcggtct- gagggcattagcgcgaaatgatttttca caagcgctggcaatccgacccgatatgcctgaagtattcaattacttaggcatttacttaacgcaggcaggca- attttgatgctgcctatgaagcgtttgat tctgtacttgagcttgatcgccattgagctggcttcccgaccgcagggcggcacctgcctgaccctgcgtttc- ccgctgtttaacaccctgaccggaggt gaagcatga 198 cccaacagcagggccgggtaggccagcaggtccgccagcgtggcgcggttaatattgaccggggcggcgg- cggcctcccccagctgcttgtgga tcattttcgcgatcttgcgggttttaccggtatcggtaccaaagaaaatgccaatgttcgccatagtacgctc- ctgtcggaatggtgttgaaaaaaggaat gacgacagaggtattgcgaaggctgtgccaggttgccctgcaccgcgacggcccatccctgccccatcaggat- cgcttcgcatcacgatgccgcgc gccaaaggcgcacccggcggggcgaaaggtaaaaatccgtgaattttccccctgtcggatcaatgatcgcgtg- gtcgttccgataagggcgcacac tttgcatggttatccgggttcggcttaccccgccgcgttttgcgcacggtgtcggacaatttgtcataactgc- gacacaggagtttgcgatgaccctgaat atgatgctcgacgccgtcctcgcagtaccattgcaaccgactttacagcaagaagtgattctggcacgcatgg- aacaaattcttgccagtcgggctttat ccgatgacgaacgcgcacagcattatatgagcgcggagtgttgtatgatagtctcggtctgagggcattagcg- cgaaatgatttttcacaagcgctggc aatccgacccgatatgcctgaagtattcaattacttaggcatttacttaacgcaggaggcaattttgatgctg- cctatgaagcgtttgattctgtacttgag cttgatcgccattgagaggcttcccgaccgcagggcggcacctgcctgaccctgcgtttcccgctgtttaaca- ccctgaccggaggtgaagcatgat ccctgaatccgacccggacaccaccgtcagacgcttcgacctctctcagcagttcaccgccatgcagcggata- agcgtggtgctgagccgggccac cgaggccagcaaaacgctgcaggaggtgctcagcgtattacacaacgatgcctttatgcagcacgggatgatc- tgcctgtacgacagcgagcagga gatcctcagtatcgaagcgctgcagcaaaccggccagcagcccctccccggcagcacgcagatccgctatcgc- cccggcgagggactggtgggg accgtgctggcccaggggcagtcgctggtgctgccccgggtcgccgacgatcagcgttttctcgaccgcctga- gcctctacgattacgatctgccgtt tatcgccgtaccgttgatggggcccaacgcccggccaataggggtgctggcggcccagccgatggcgcgccag- gaagagcggctgccggcctgc acccgttttctcgaaaccgtc 199 atggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgcgccacc- cgctgctgctggatgagctgctggat cccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctgctccgcgtgc- cggaagaggatgaagagcagca gctgcatatcgcggcggcggatatcgctggtaccctgccggtgatgaaggtcagcgatcacttaacctggctt- gccgaagcgatcctcgacgcggtg gtgcagcaggcatgggggcagatggtcgctcgctacggccagccgacccacctgcacgatcgccagggtcgcg- gcttcgccgtcgtcggctacg gtaagcttggcggctgggagctgggctacagctccgatctcgatctggtgttcctccatgactgcccggcgga- ggtgatgaccgacggcgagcggg agattgacggccgtcagttctacctgcggctggcccagcggatcatgcacctgttcagcacccgcacctcgtc- cggtattctctacgaagtggacgcc cggctgcgtccttctggcgcggcggggatgctggtcaccaccgccgacgcgtttgctgactatcagcagaacg- aagcctggacgtgggaacatcag gcgctggtgcgcgcccgcgtggtctatggcgacccggcgctgcaggcgcgctttgacgccattcgtcgcgata- tcctgaccaccccgcgggaggg gatgaccctgcagaccgaggttcgcgagatgcgcgagaagatgcgcgcccaccttggcaacaaacatcccgat- cgttttgatatcaaagccgatgcc
ggcgggatcaccgatattgaatttattactcagtatctggtcctacgctatgccagtgacaagccgaagctga- cccgctggtctgacaacgtgcgtattct tgagctgctggcgcagaacgacatcatggacgaggaggaggcgcgcgccttaacgcatgcgtacaccaccttg- cgtgatgcgctccatcacctggc cctgcaggagcagccgggacacgtggcgccagaggccttcagccgggagcgtcagcaggtcagcgccagctgg- cagaagtggctgatggcttaa 200 cgtaaggcgaccacccagctccgcgcgttgctgaacgacgctgaagccgttctgctggccgcggacaccg- ccgacgaggcgttattccgcaccga ggtcgtcggcgccaaactggccctgactgaatggctggtccagcgcggctggcgtccgttcctcaacgaggca- ggagagaaaaaaatagccggat cgttcaaacggtttgccgatattaacctctcgcgggtggcggccgagctgcgcagcgccgtgcagcatctggc- ggttgaagatgccgccgaccagtt gccgaagctgtcccgcgacatcgacagcgtccagctgctggcgggcgcctatggcgacgccgtcgcgccgtgg- ctggagaactggcaggagctt caccgtgcaatagcacatgacgatcgcagcgtctttgaatatttccgtcgccaggcgctggctgccgagccgt- tctggctgcatagtgggaaaacgata atttcaggccagggagcccttatggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgcca- gccagctggcgcgccacccgctg ctgctggatgagctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgc- gccagtacctgctgcgcgtgccg gaagaggatgaagagcagcagctgcatatcgcggcggcggatatcgctggtaccctgccggtgatgaaggtca- gcgacacttaacctggcttgcc gaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggtcgctcgctacggccagccgacccacc- tgcacgatcgccagggtcgcg gcttcgccgtcgtcggctacggtaagcttggcggctgagagctgggctacagctccgatctcgatctggtgtt- cctccatgactgcccggcggaggtg atgaccgacggcgagcgggagattgacggccgtcagttctacctccggctggcccagcggatcatgcacctgt- tcagcacccgcacctcgtccggt attctctacgaagtggacgcccggctgcgtccttctggcgcggcggggatgctggtcaccaccgccgacgcgt- ttgctgactatcagcagaacgaag cctggacgtgggaacatcaggcgctggtgcgcgcccgcgtggtctatggcgacccggcgctgcaggcgcgctt- tgacgccattcgtcgcgatatcc tgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcgagatgcgcgagaagatgcgcgcccacct- tggcaacaaacatcccgatcg ttttgatatcaaagccgatgccggcgggatcaccgatattgaatttattactcagtatctggtcctacgctat- gccagtgacaagccgaagctgacccgct ggtctgacaacgtgcgtattcttgagctgctggcgcagaacgacatcatggacgaggaggaggcgcgcgcctt- aacgcatgcgtacaccaccttgc gtgatgcgctccatcacctggccctgcaggagcagccgggacacgtggcgccagaggccttcagccgggagcg- tcagcaggtcagcgccagctg gcagaagtggctgatggcttaactataaaatcgggtgtgctattatcgcgcgcaaagtttgcgtctcgcagga- gagagtcatgaaactaacgctgccg gagtttgaacgtgcaggagtgttggtggtgggtgatgtgatgctggaccgctactggtacggccccaccagtc- gtatttccccggaagccccggtgcc ggtggtgaaggtggaaaatatcgaagtacgtcctggcggcgcggcaaacgtagcgatgaacatcgcctccctg- ggggcaacgtcgcgcctggtgg gattgaccgggattgatgacgctgcccgcgcgctgagccaggcgctggccaatgtgaatgtgaagtgcgactt- cgtctccgtcccgactcacccgac catcaccaagctgcgggtgctgtcgcgcaatcagcagctgatccgcctcgactttgaagagggcttctccggc- gtggatccgcagccgatgcatgag cgcattcagcaggcgctgggagccattggcgcactgg 201 atgaccctgaatatgatgctcgagctaaagttctcggctaatcgctgattacatttgacgcaatgcgcaa- taaaagggcatcatttgatgccctttttgcac gctttcataccagaacctggctcatcagtgattttttttgtcataatcattgctgagacaggctctgaagagg- gcgtttatacaccaaaccattcgagcggt agcgcgacggcaagtcagcgttctcctttgcaatagcagggaagaggcgccagaaccgccagcgttgaagcag- tttgaacgcgttcagtgtataatc cgaaacttaatttcggtttggagccattgagctggcttcccgaccgcagggcggcacctgcctgaccctgcgt- ttcccgctgtttaacaccctgaccgg aggtgaagcatga 202 ggccgtcgcccagcgtcggcgtccccaacagcagggccgggtaggccagcaggtccgccagcgtggcgcg- gttaatattgaccggggcggcgg cggcctcccccagctgcttgtggatcattttcgcgatcttgcgggttttaccggtatcggtaccaaagaaaat- gccaatgttcgccatagtacgctcctgt cggaatggtgttgaaaaaaggaatgacgacagtggtattgcgaaggctgtgccaggttgccctgcaccgcgac- ggcccatccctgccccatcagga tcgcttcgcatcacgatgccgcgcgccaaaggcgcacccggcggggcgaaaggtaaaaatccgtgaattttcc- ccctgtcggatcaatgtttcgcgtg gtcgttccgataagggcgcacactttgcatggttatccgggttcggcttaccccgccgcgttttgcgcacggt- gtcggacaatttgtcataactgcgaca caggagtttgcgatgaccctgaatatgatgctcgagctaaagttctcggctaatcgctgataacatttgacgc- aatgcgcaataaaagggcatcatttgat gcccttttgcacgctttcataccagaacctggctcatcagtgattttttttgtcataatcattgctgagacag- gctctgaagagggcgtttatacaccaaac cattcgagcggtagcgcgacggcaagtcagcgttctcctttgcaatagcagggaagaggcgccagaaccgcca- gcgttgaagcagtttgaacgcgt tcagtgtataatccgaaacttaatttcggtttggagccattgagctggcttcccgaccgcagggcggcacctg- cctgaccctgcgtttcccgctgtttaac accctgaccggaggtgtagcatgatccctgaatccgacccagacaccaccgtcagacgcttcgacctctctca- gcagttcaccgccatgcagcggat aagcgtggtgctgagccgggccaccgaggccagcaaaacgctgcaggaggtgctcagcgtattacacaacgat- gcctttatgcagcacgggatgat ctgcctgtacgacagcgagcaggagatcctcagtatcgaagcgctgcagcaaaccggccagcagcccctcccc- ggcagcacgcagatccgctatc gccccggcgagggactggtggggaccgtgctggcccaggggcagtcgctggtgctgccccgggtcgccgacga- tcagcgttttctccaccgcctg agcctctacgattacgatctgccgtttatcgccgtaccgttgatggggcccaacgcccggccaataggggtgc- tggcggcccagccgatggcgcgc caggaagagcggctgccggcctgcacccgttttctcgaaaccgtc 203 atgaccctgaatatgatgctcgagcccgctgaccgaccagaacttccaccttggactcggctataccctt- ggcgtgacggcgcgcgataactgggact acatccccattccggtgatcttaccattggcgtcaataggttacggtccggcgactttccagatgacctatat- tcccggcacctacaataacggtaacgttt acttcgcctgggctcgtatacagttttaattcgctaagtcttagcaataaatgagataagcggtgtgtcttgt- ggaaaaacaaggactaaagcgttaccca ctaaaaaagatagcgacttttatcactttttagcaaagttgcactggacaaaaggtaccacaattggtgtact- gatactcgacacagcattagtgtcgatttt tcatataaaggtaattttggccattgagctggcttcccgaccgcagggcggcacctgcctgaccctgcgtttc- ccgctgtttaacaccctgaccggaggt gaagcatga 204 ggccgtcgcccagcgtcggcgtccccaacagcagggccgggtaggccagcaggtccgccagcgtggcgcg- cttaatattgaccggggcggcgg cggcctcccccagctgcttgtggatcattttcgcgatcttgcgggttttaccggtatcggtaccaaagaaaat- gccaatgttcgccatagtacgctcctgt cggaatggtgttgaaaaaaggaatgacgacagaggattgcgaaggctgtgccaggttgccctgcaccgcgacg- gcccatccctgccccatcagga tcgcttcgcatcacgatgccgcgcgccaaaggcgcacccggcggggcgaaaggtaaaaatccgtgaattttcc- ccctgtcggatcaatgtttcgcgtg gtcgttccgataagggcgcacactttgcatggttatccgggttcggcttaccccgccgcgttttgcgcacggt- gtcggacaatttgtcataactgcgaca caggagtttgcgatgaccctgaatatgatgctcgagcccgctgaccgaccagaacttccaccttggactcggc- tatacccttggcgtgacggcgcgcg ataactgggactacatccccattccggtgatcttaccattggcgtcaataggttacggtccggcgactttcca- gatgacctatattcccggcacctacaat aacggtaacgtttacttcgcctgggctcgtatacagttttaattcgctaagtcttagcaataaatgagataag- cggtgtgtcttgtggaaaaacaaggacta aagcgttacccactaaaaaagatagcgacttttatcactttttagcaaagttgcactggacaaaaggtaccac- aattggtgtactgatactcgacacagca ttagtgtcgatttttcatataaaggtaattttggccattgagctggcttcccgaccgcagggcggcacctgcc- tgaccctgcgtttcccgctgtttaacacc ctgaccggaggtgaagcatgatccctgaatccgacccggacaccaccgtcagacgatcgacctctctcagcag- ttcaccgccatgcagcggataag cgtggtgctgagccgggccaccgaggccagcaaaacgctgcaggaggtgctcagcgtattacacaacgatgcc- tttatgcagcacgggatgatctg cctgtacgacagcgagcaggagatcctcagtatcgaagcgctgcagcaaaccggccagcagcccctccccggc- agcacgcagatccgctatcgcc ccggcgagggactggtggggaccgtgctggcccaggggcagtcgctggtgctgccccgggtcgccgacgatca- gcgttttctcgaccgcctgagc ctctacgattacgatctgccgtttatcgccgtaccgttgatggggcccaacgcccggccaataggggtgctgg- cggcccagccgatggcgcgccag gaagagcggctgccggcctgcacccgttttctcgaaaccgtc 205 atggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgcgccacc- cgctgctgctggatgagctgctggat cccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctgctgcgcgtgc- cggaagaggatgaagagcagca gctgcatatcgcggcggcggatatcgctggtaccctgccggtgatgaaggtcagcgatcacttaacctggctt- gccgaagcgatcctcgacgcggtg gtgcagcaggcatgggggcagatggtcgctcgctacggccagccgacccacctgcacgatcgccagggtcgcg- ccttcgccgtcgtcggctacg gtaagcttggcggctgggagagggctacagctccgatctcgatctggtgttcctccatgactgcccggcggag- gtgatgaccgacggcgagggg agattgacggccgtcagttctacctgcggctggcccagcggatcatgcacctgttcagcacccgcacctcgtc- cggtattctctacgaagtggacgcc cggctgcgtccttctggcgcggcggggatgctggtcaccaccgccgacgcgtttgctgactatcagcagaacg- aagcctggacgtgggaacatcag gcgctggtgcgcgcccgcgtggtctatggcgacccggcgctgcaggcgcgctttgacgccattcgtcgcgata- tcctgaccaccccgcgggaggg gatgaccctgcagaccgaggttcgcgagatgcgcgagaagatgcgcgcccaccttggcaacaaacatcccgat- cgttttgatatcaaagccgatgcc ggcgggatcaccgatattgaatttattactcagatctggtcctacgctatgccagtgacaagccgaagctgac- ccgctggtctgacaacgtgcgtattct tgagctgctggcgcagaacgacatcatggacgaggaggaggcgcgcgccttaacgcatgcgtacaccaccttg- cgtgatgcgctccatcacctggc cctgcaggagcagccgcgacacgtggcgccagaggccttcagccgggagcgtcagcaggtcagcgccagctgg- cagaagtggctgatggcttaa 206 cgtaaggcgaccacccagctccgcgcgttgctgaacgacgctgaagccgttctgctggccgcggacaccg- ccgacgaggcgttattccgcaccga ggtcgtcggcgccaaactggccctgactgaatggctggtccagcgcggctggcgtccgttcctcaacgaggca- ggagagaaaaaaatagccggat cgttcaacggtttgccgatattaacctctcgcaggtggcggccgagctgcgcagcgccgtgcagcatctggcg- gttgaagatgccgccgaccagtt gccgaagctgtcccgcgacatcgacagcgtccagctgctggcgggcgcctatggcgacgccgtcgcgccgtgg- ctggagaactggcaggagctt caccgtgcaatagcacatgacgatcgcagcgtctttgaatatttccgtcgccaggcgctggctgccgagccgt- tctggctgcatagggaaaacgata atttcaggccagggagcccttatggcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgcca- gccagctggcgcgccacccgctg ctgctggatgagctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgc- gccagacctgctgcgcgtgccg gaagaggatgaagagcagcagctgcatatcgcggcggcggatatcgctggtaccctgccggtgatgaaggtca- gcgatcacttaacctggcttgcc gaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggtcgctcgctacggccagccgacccacc- tgcacgatcgccagggtcgcg gcttcgccgtcgtcggctacggtaagcttggcggctgggagctgggctacagctccgatctcgatctggtgtt- cctccatgactgcccggcggaggtg atgaccgacggcgaggggagattgacggccgcagttctacctgcggctggcccagcggatcatgcacctgttc- agcacccgcacctcgccggt attctctacgaagtggacgcccggctgcgtccttctggcgcggcgcggatgctggtcaccaccgccgacgcgt- ttgctgactatcagcagaacgaag cctggacgtgggaacatcaggcgctggtgcgcgcccgcgtggtctattgcgacccggcgctgcaggcgcgctt- tgacgccattcgtcgcgatatcc tgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcgagatgcgcgagaagatgcgcgcccacct- tggcaacaaacatcccgatcg ttttgatatcaaagccgatgccggcgggatcaccgatattgaatttattactcagtatctggtcctacgctat- gccagtgacaagccgaagctgacccgct ggtctgacaacgtgcgtattcttgagctgctggcgcagaacgacatcatggacgaggaggaggcgcgcgcctt- aacgcatgcgtacaccaccttgc gtgatgcgctccatcacctggccctgcaggagcagccgggacacgtggcgccagaggccttcagccgagagcg- tcagcaggtcagcgccagctg gcagaagtggctgatggcttaactataaaatcgggtgtgctattatcgcgcgcaaagtttgcgtctcgcagga- gagagtcatgaaagtaacgctgccg gagtttgaacgtgcaggagttggtggtgggtgatgtgatgctggaccgctactggtacggccccaccagtcgt- atttccccggaagccccggtgcc ggtggtgaaggtggaaaatatcgaagaacgtcctggcggcgcggcaaacgtagcgatgaacatcgcctccctg- ggggcaacgtcgcgcctggtgg gattgaccgggattgatgacgctgcccgcgcgctgagccaggcgctggccaatgtgaatgtgaagtgcgactt- cgtctccgtcccgactcacccgac catcaccaagctgcgggtgctgtcgcgcaatcagcagctgatccgcctcgactttgaagagggcttctccggc- gtggatccgcagccgatgcatgag cgcattcagcaggcgctgggagccattggcgcactgg 207 atgaccctgaatatgatgctagaagcgtcaggtaccggtcatgattcaccgtgcgattctcggttccctg- gagcgcttcattggcatcctgaccgaagag ttcgctggcttcttcccaacctggattgcaccagtgcaggtagtggtcatgaatattaccgattctcaggctg- aatacgttaccgaangacgcgtaaacta caaaatgcgggcattcgtgtaaaagcagacttgagaaatgagaagattggctttaaaatccgcgagcacactt- tacgtcgtgtcccgtatatgtggtct gtggcgacaaagaagtcgaagccggcaaagtggccgtgcgcacccgtcgcgggaaagacctcggcagcatgga- cgtaagtgaagtgattgagaa gctgcaacaagagattcgcagccgcagtcttcaacaactggaggaataaggtattaaaggcggaaaacgagtt- caaacggcacgtccgaatcgtatc aatgccgagattcgcgccctcgaagttcgcgccattgagctggcttcccgaccgcagggcgccacctgcctga- ccctgcgtttcccgctgtttaacac cctgaccggaggtgaagcatga 208 ggccgtcgcccagcgtcggcgtccccaacagcagggccgggtaggccagcaggccgccagcgtggcgcgg- ttaatattgaccggggcggcgg cggcctcccccagagcttgtggatcattttcgcgatcttgcgggttttaccggtatcgaaccaattgaaaatg- ccaatgttcgccatagtacgctcctgt cggaatggtgttgaaaaaaggaatgacgacagaggtattgcgaaggctggccaggttgccctgcaccgcgacg- gcccatccctgccccatcagga tcgcttcgcatcacgatgccgcgcgccaaaggcgcacccggcggggcgaaaggtaaaaatccgtgaattttcc- ccctgtcggatcaatgtttcgcgtg gtcgttccgataagggcgcacactttgcatggttatccgggttcggcttaccccgccgcgttttgcgcacggt- gtcggacaatttgtcataactgcgaca caggagtttgcgatgaccctgaatatgatgctagaagcgtcaggtaccggtcatgattcaccgtgcgattctc- ggttccctggagcgatcattggcatc ctgaccgaagagttcgctggcttcttcccaacctggattgcaccagtgcaggtagtggtcatgaatattaccg- attctcaggctgaatacgttaacgaatt gacgcgtaaactacaaaatgcggcattcgtgtaaaagcagacttgagaaatgagaagattggctttaaaatcc- gcgagcacactttacgtcgtgtccc gtatatgttggtctgtggcgacaaagaagtcgaagccggcaaagtggccgtgcgcacccgtcgcgggaaagac- ctcggcagcatggacgtaagtg aagtgattgagaagctgcaacaagagattcgcagccgcagtcttcaacaactggaggaataaggtattaaagg- cggaaancgagttcaaacggcac gtccgaatcgtatcaatggcgagattcgcgccctggaagttcgcgccattgagctggcttcccgaccgcaggg- cggcacctgcctgaccctgcgtttc ccgctgtttaacaccctgaccggaggtgaagcatgatccctgaatccgacccggacaccaccgtcagacgctt- cgacctctctcagcagttcaccgcc atgcagcggataagcgtggtgctgagccgggccaccgaggccagcaaaacgctgcaggtggtgctcagcgtat- tacacaacgatgcctttatgcag cacgggatgatctgcctgtacgacagcgagcaggagatcctcagtatcgaagcgctgcagcaaaccggccagc- agcccctccccggcagcacgc agatccgctatcgccccggcgagggactggtggggaccgtgctggcccaggggcagtcgctggtgctgccccg-
ggtcgccgacgatcagcgtttt ctcgaccgcctgagcctctacgattacgatctgccgtttatcgccgtaccgttgatggggcccaacgcccggc- caataggggtgctggcggcccagc cgatggcgcgccaggaagagcggctgccggcctgcacccgttttctcgaaaccgtc 209 atggcgctgaagcacctgatcacgactgcgcggcgtcgccgatggtcgccagccagctcgcgcgccaccc- gctgctgctggatgagctgctggat cccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctgctgcgcgtgc- cggaagaggatgaagagcagca gctggaggcgttgcgccagtttaagcaggcgcagcagctgcatatcgcggcggcggatatcgctggtaccctg- ccggtgatgaaggtcagcgatca cttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggtcgctcgctac- ggccagccgacccacctgcacg atcgccagggtcgcggcttcgccgtcgtcggctacggtaagcttacggctgggagctgggctacagctccgat- ctcgatctggtgttcctccatgac tgcccggcggaggtgatgaccgacggcgagcggcagattgacggccgtcagttctacctgcggctggcccagc- ggatcatgcacctgttcagcac ccgcacctcgtccggtattctctacgaagtggacgcccggctgcgtccttctggcgcggcggggatgctggtc- accaccgccgacgcgtttgctgact atcagcagaacgaagcctggacgtgggaacatcaggcgctggtgcgcgcccgcggtctatggcgacccggcgc- tgcaggcgcgctttgacgc cattcgtcgcgatatcctgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcgagatgcgcgag- aagatgcgcgcccaccttggca acaaacatcccgatcgttttgatatcaaagccgatgccggcgggatcaccgatattgaatttattactcagta- tctggtcctacgctatgccagtgacang ccgaagctgacccgctggtagacaacgtgcgtattcttgagctgctggcgcagaacgacatcatggacgagga- ggaggcgcgcgccttaacgcat gcgtacaccaccttgcgtcatgcgctccatcacctggccctgcaggagcagccgggacacgtggcgccagagg- ccttcagccgggagcgtcagca ggtcagcgccagctggcagaagtggctgatggcttaa 210 cgtaaggcgaccacccagctccgcgcgttgctgaacgacgctgaagccgttctgctggccgcggacaccg- ccgacgaggcgttattccgcaccga ggtcgtcggcgccaaactggccctgactgaatggctggtccagcgcggctggcgtccgttcctcaacgaggca- ggagagaaaaaaatagccggat cgttcaaacggtttgccgatattaacctctcgcgggtggcggccgagctgcgcagcgccgtgcagcatctggc- ggttgaagatgcgccgaccagtt gccgaagctgtcccgcgacatcgacagcgtccagctgctggcgggcgcctatggcgacgccgtcgcgccgtgg- ctggagaactggcaggagctt caccgtgcaatagcacatgacgatcgcagcgtctttgaatatttccgcgccaggcgctggctgccgagccgtt- ctggcttcatagtggaaaacgata atttcaggccagggagcccttatggcgctgaagcacctgatcacgctagcgcggcgtcgccgatggtcgccag- ccagctggcgcgccacccgctg ctgctggatgagctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgc- gccagtacctgctgcgcgtgccg gaagaggatgaagagcagcagctggaggcgttgcgccagtttagcaggcgcagcagctgcatatcgcggcggc- ggatatcgctggtaccctgcc ggtgatgaaggtcagcgatcacttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgg- gggcagatggtcgctcgctacgg ccagccgacccacctgcacgatcgccagcgtcgcggcttcgccgtcgtcggctacggtaagcttggcggctgg- gagctgggctacagctccgatct cgatctggtgttcctccatgtctgcccggcggaggtgatgaccgacggcgagcgggagattgacggccgtcag- ttctacctgcggctggcccagcg gatcatgcacctgttcagcacccgcacctcgtccggtattctctacgaagtggacgcccggctgcgtccttct- ggcgcggcggggatgctggtcacca ccgccgacgcgtttgctgactatcagcagaacgaagcctggacgtgggaacatcaggcgctggtgcgcgcccg- cgtggtctatgccgacccggcg ctgcaggcgcgctttgacgccattcgtcgcgatatcctgaccaccccgcgggaggggatgaccctgcagaccg- aggttcgcgagatgcgcgagaa gatgcgcgcccaccttggcaacaaacatcccgatcgttttgatatcaaagccgatgccggcgggatcaccgat- attgaatttattactcagtatctggtcc tacgctatgccagtgacaagccgaagctgacccgctggtctgacaacgtgcgtattcttgagctgctggcgca- gaacgacatcatggacgaggagga ggcgcgcgccttaacgcatgcgtacaccaccttgcgtgatgcgctccatcacctggccctgcaggagcagccg- ggacacgtggcgccagaggcct tcagccgggagcgtcagcaggtcagcgccagctggcagaagtggctgatcgcttaactataaaatcgggtgtg- ctattatcgcgcgcaaagtttgcgt ctcgcaggagagagtcatgaaagtaacgctgccggagttgaacgtgcaggagtgttggtggtggggatgtgat- gctggaccgctactggtacggc cccaccagtcgtatttccccggaagccccggtgccggtggtgaaggtggaaaatatcgaagaacgtcctggcg- gcgcggcaaacgtagcgatgaa catcgcctccctgggggcaacgtcgcgcctggtgggattgaccgggattgatgacgctgcccgcgcgctgagc- caggcgctggccaatggaatgt gaagtgcgacttcgtctccgtcccgactcacccgaccatcaccaagctgcgggtgctgtcgcgcaatcagcag- ctgatccgcctcgactttgaagag ggcttctccggcgtggatccgcagccgatgcatgagcgcattcagcaggcgctgggagccattggcgcactgg 211 atggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtcgcgacaacttgcacgtcatcct- ttattgctcgatgaactgctcgacccgc gcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatagatgcgggtgccaaca- gaagacgaagaacagcagcttg aagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccggt- gatgaaagtcagtgaccatttaacc taccttgccgaggccattctcgatgtcgtggtgcagcatgcgtgggaacaaatggtcgtaaaatacgggcagc- ccgcgcatcttcagcaccgtgagg ggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggatct- ggtcttcctgctcgattgcgcgccgg aggtgatgacggacggcgaacgcagcatcgacggacgtcagttttatatcggctggcgcagcgcattatgcac- ttattcagcacccggacatcgtca ggcattctttacgaggttgatccgcgtagcgaccttccggcgcatccggcatgctggtcagtaccattgaagc- gtttgcagattatcaggccaatgaag cctggacgtgggagcatcaggcgctggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaatt- taacgccacgcgtcgcgacattct ttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccatctg- gggagtaaaaaagcccacgagtt tgatctgaaagccgatccgggtggcatcacggatattgaattcattgcaccatacctggttctgcgtttcgcg- cntgatgagccgaagctgacgcgctg gtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgccggaagaggaagcgcgccatctg- acgcaggcttatgtgacgctgcgcg atgaaattcatcatctggcgttgcaggaacacagcgggaagtggccgcggacagctttgctactgagcgcgcg- cagatccgtgccagctgggcaa agtggctcggctga 212 cggtactggaacagaaatcggcggatgcgtaggagatttgttatgacacggcctgtctgaagtgcaagta- gtgcttacttcctggctggcaacctcag gctggacgccgtttattgatgataaatctgcgaagaaactggacgcttccttcaaacgttttgctgacatcat- gctcggtcgtaccgcagcggatctgaaa gaagcctttgcgcagccactgacggaagaaggttatcgcgatcagctggcgcgcctgaaacgccagatcatta- ccttccatttgcttgccggtgcttac cctgaaaaagacgtcgatgcgtatattgccggctgggtggacctgcaacaggccatcgttcagcagcaacacg- cctgggaggattcggcccgttctc acgcggtgatgatggatgattctggttaaacgggcaacctcgtaactgactgactagcctgggcaaactgccc- gggcttttttttgcaaggaatctgat ttcatggcgctcaaacagttaatccgctgtggccgcctcgccgatggtcgcgacacaacttgcacgtcatcct- ttattgctcgatgaactgctcgaccc gcgcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatctgatgcgggtgcca- acagaagacgaagaacagcagct tgaagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccg- gtgatgaaagtcagtgaccatttaa cctaccttgccgaggccattctcgatgtcgtggtgcagcatgcgtgggaacaaatggtcgtaaaatacgggca- gcccgcgcatcttcagcaccgtga ggggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggat- ctggtcttcctgctcgattgcgcgcc ggaggtgatgacggacggcgaacgcagcatcgacggacgtcagttttatcttcggctggcgcagcgcattatg- cacttattcagcacccggacatcgt caggcattctttacgaggttgatccgcgctgcgaccttccggcgcatccggcatgaggtcagtaccattgaag- cgtttgcagattatcaggccaatga agcctggacgtgggagcatcaggcgaggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaat- ttaacgccacgcgtcgcgacat tctttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccat- ctggggagtaaaaaagcccacga gtttgatctgaaagccgatccgggtggcatcacggatattgaattcattgcacaatacctggttctgcgtttc- gcgcatgatgagccgaagctgacgcgc tggtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgcccgaagaggaagcgcgccatc- tgacgcaggcttatgtgacgctgcg cgatgaaattcatcatctggcgttgcaggaacacagcgggaaagtggccgcggacagctttgctactgagcgc- gcgcagatccgtgccagctgggc aaagtggctcggctgagggtttttattcggctaacaggcgcttgtgatattatccggcgcattgtatttaccc- gatttgatttatctgttttggagtcttgggat gaaagtgactttgcctgattttcaccgcgcaggtgtgctggttgtcggtgacgtaatgttagaccgttactgg- tatggcccgaccaatcgtatttctccgga agctccggtgccggtggtgaaggtcagtaccattgaagagcggcctggcggtgagctaacgtggcgatgaaca- tttcatctctgggcgcctcttcct gtctgatcggcctgaccggcgtagacgacgctgcgtgccctcagtgagcgtctggcagaagtgaaagaaactg- cgatttcgtcgcactatccaca catcctaccatcaccaaactgcgaattttgtcccgtaaccagcaactgatccgcctcgactttgaggaaggtt- ttgaaggcgttgatctcgagccgatgct gaccaaaataga 213 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggaattttttttcacaaaggtagcgttattgaatcgcacattttaaactgttggccg- ctgtggaaggaatattggtgaaaggtg cggttttaaggcctttttctttgactctctgtcgttacaaagttaatatgcgcgccctccgtctctgaagctc- tcggtgaacattgttgcgaggcaggatgcg agctggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaagcggcgt- tttcccgtccggggaatggcatggag ctgcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggca- ccttgctgacgttacgcctgccggtac agcaggttatcaccggaggcttaaaatga 214 tgtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttgttgtg- cgaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgcccgttttgacgaaaaatgcacataaattgcctg- cgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggaattttttttcacaaagcgtagcgttattgaatcgcacattttaaactgt- tggccgctgtggaagcgaatattggtga aaggtgcggttttaaggcctttttctttgactctctgtcgttacaaagttaatatgcgcgccctccgtctctg- aagctctcggtgaacattgttgcgaggcag gatgcgagctggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaag- cggcgttttcccgtccggggaatggc atggagctgcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatg- gcggcaccttgctgacgttacgcctgc cggtacagcaggttatcaccggaggcttaaaatgacccagttacctaccgcgggcccggttatccggcgcttt- gatatgtctgcccagtttacggcgctt tatcgcatcagcgtggcgctgagtcaggaaagcaacaccgggcgcgcactggcggcgatcctcgaagtgcttc- acgatcatgcatttatgcaatacg gcatggtgtgtctgtttgataaagaacgcaatgcactctttgtggaatccctgcatggcatcgacggcgaaag- gaaaaaagagacccgccatgtccgtt accgcatgggggaaggcgtgatcggcgcggtgatgagccagcgtcaggcgctggtgttaccgcgcatttcaga- cgatcagcgttttctcgaccgcct gaatatttacgattacagcctgccgttgattggcgtgccgatccccggtgcggataatcagccatcgggcgtg- ctggtggcacagccgatggcgttgc acgaagaccggctgactgccagtacgcggtttttagaaatggtc 215 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggaattttttttcacaaagcgtagcgttattgaatcgcacattttgatgttggccgc- tgtggaagcgaatattggtgaaaggtg cggttttaaggcctttttctttgactctctgtcgttacaaagttaatatgcgcgccctccgtctctgaagctc- tcggtgaacattgttgcgaggcaggatgcg agctggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggttgcggcgt- tttcccgtccggggaatggcatggag ctgcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggca- ccttgctgacgttacgcctgccggtac agcaggttatcaccggaggcttaaaatga 216 tgtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttgttgtg- cgaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgcccgttttgcacgaaaaatgcacataaattgcct- gcgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggaattttttttcacaaagcgtagcgttattgaatcgcacattttaaactgt- tggccgctgtggaagcgaatattggtga aaggtgcggttttaaggcctttttctttgactctctgtcgttacaaagttaatatgcgcgccctccgtctctg- aagctctcggtgagcattgttgcgaggcag gatgcgagctggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaag- cggcgttttcccgtccggggaatggc atggagctgcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatg- gcggcaccttgctgacgttacgcctgc cggtacagcaggttatcaccggaggcttaaaatgacccagttacctaccgcgggcccggttatcccgcgcttt- gatatgtctgcccagtttacggcgctt tatcgcatcagcgtggcgctgagtcaggaaagcaacaccgggcgcgcactggcggcgatcctcgaagtgcttc- acgatcatgcatttatgcaatacg gcatggtgtgtctgtttgataaagaacgcaatgcactctttgtggaatccctgcatggcatcgacggcgaaag- gaaaaaagagacccgccatgtccgtt accgcatgggggaaggcgtgatcggcgcggtgatgagccagcgtcaggcgctggtgttaccgcgcatttcaga- cgatcagcgttttctcgaccgcct gaatatttacgattacagcctgccgttgtttggcgtgccgatccccggtgcggataatcagccatcgggcgtg- ctggtggcacagccgatggcgttgc acgaagaccggctgactgccagtacgcggtttttagaaatggtc 217 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggttaaaaacgtgaccacgagcattaataaacgccacgaaatgtggcgtttatttat-
tcaaaaagtatcttctttcataaaaagtg ctaaatgcagtagcagcaaaattgggataagtcccatggaatacggctgttttcgctgcaatttttaactttt- tcgtaaaaaaagatgtttctttgagcgaac gatcaaaatataggttaaccggcaaaaaattattctcattagaaaatagtttgtgtaatacttgtaacgctac- atggagattaacttaatctagagggtttta taccgtctctgaagctacggtgaacattgttgcgaggcaggatgcgagctggttgtgttttgacattaccgat- aatgtgccgcgtgaacgggtgcgttat gcccgcccggaagcggcgttttcccgtccggggaatggcatggagagcgccttatccagacgctgatcgccca- tcatcgcggttctttagatctctcg gtccgccctgatggcggcaccttgctgacgttacgcctgccggtacagcaggttatcaccggaggcttaaaat- ga 218 gtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttgttgtgc- gaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttatgttgatgcagacgggttaatgcccgttttgcacgaaaaatgcacataaattgcctg- cgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtagctggttaaaaacgtgaccacgagcattaataaacgccacgaaatgtggcgtttat- ttattcaaaaagtatatctttcataa aaagtgctaaggcagtagcagcaaaattgggataagtcccatggaatacggctgttttcgctgcaatttttaa- ctttttcgtaaaaaaagatgtttctttgag cgaacgatcaaaatatagcgttaaccggcaaaaaattattacattagaaaatagtttgtgtaatacttgtaac- gctacatggagattaacttaatctagagg gttttataccgtctctgaagctctcggtgaacattgttgcgaggcaggatgcgagaggttgtgttttgacatt- accgataatgtgccgcgtgaacgcgtg cgttatgcccgcccggaagcggcgttttcccgtccggggaatggcatggagctgcgccttatccagacgctga- tcgcccatcatcgcggttctttagat ctctcggtccgccctgatggcggcaccttgctgacgttacgcctgccggtacagcaggttatcaccggaggct- taaaatgacccagttacctaccgcg ggcccggttatccggcgctttgatatgtctgcccagtttacggcgctttatcgcatcagcgtggcgctgagtc- aggaaagcaacaccgggcgcgcact ggcggcgatcctcgaagtgcttcacgatcatgcatttatgcaatacggcatggtgtgtctgtttgataaagaa- cgcaatgcactctttgtggaatccctgc atggcatcgacggcgaaaggaaaaaagagacccgccatgtccgttaccgcatgggggaaggcgtgatcggcgc- ggtgatgagccagcgtcaggc gctggtgttaccgcgcatttcagacgatcagcgttttctcgaccgcctgaatatttacgattacagcctgccg- ttgattggcgtgccgatccccggtgcgg ataatcagccatcgggcgtgctggtggcacagccgatggcgttgcacgaagaccggctgactgccagtacgcg- ctttttagaaatggtcg 219 atgagcatcacggcgttatcagactcatttcctgagggaatatcgccagccgcttgtcgctgcaacatcc- ttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggtgaacatcactgatgcacaagctacctatgtcgaagaattaactaaaaaactgaa- gatgcaggcattcgcgttaaagcc gacttgagaaatgagaagattggctttaaaattcgcgaacacacgctacgccgtgttccttatatgttagttt- gtggcgataaagaggtcgaagcaggca aagttgctgttcgtacccgccgcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaactgct- ggcggaaatccgcagcagaagtctt catcaactggaggaataaagtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcattaacaaag- agattcgcgcgcaagaagttcgcc tcacaggcgtcgatggcgagcagattggtattgtcagtctgaatgaagctcttgaaaaagctgaggaagcggg- cgtcgatttagtagaaatcagtccg aatgccgagccgccagtttgtcgaatcccgtctagaagctctcggtgaacattgttgcgaggcaggatgcgag- ctggttgtgttttgacattaccgata atggccgcgtgaacgggtgcgttatgcccgcccggaagcggcgttttcccgtccggggaatggcatggagctg- cgccttatccagacgctgatcgc ccatcatcgcggttctttagatctctcggtccgccctgatggcggcaccttgctgacgttacgcctgccggta- cagcaggttatcaccggaggcttaaaa tga 220 tgtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttggtgcg- aacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgcccgttttgcacgaaaaatgcacataaattgcct- gcgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggtgaacatcactgatgcacaagctacctatgtcgaagaattaactaaaaaa- ctgcaagatgcaggcattcgcgtta aagccgacttgagaaatgagaagattggctttaaaattcgcgaacacacgctacgccgtgttccttatatgtt- agtttgtggcgataaagaggtcgaagc aggcaaagttgctgttcgtacccgccgcggcattagacttaggaagcatggatgttagcgaagtcgttgacaa- actgctggcggaaatccgcagcaga agtcttcatcaactggaggaataaagtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcatta- acaaagagattcgcgcgcaagaagt tcgcctcacaggcgtcgatggcgagcagattggtattgtcagtctgaatgaagctcttgaaaaagctgaggaa- gcgggcgtcgatttagtagaaatca gtccgaatgccgacccgccagtttgtcgaatcccgtctctgaagctctcggtgaacattgttgcgaggcagga- tgcgagctggttgtgttttgacattac cgataatgtgccgcgtgaacgggtgcgttatgcccgcccggagggcgttttcccgtccggggaatggcatgga- gctgcgccttatccagacgctg atcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggcaccttgctgacgttacgcctgc- cggtacagcaggttatcaccggaggct taaaatgacccagttacctaccgcgggcccggttatccggcgctttgatatgtctgcccagtttacggcgctt- tatcgcatcagcgtggcgctgagtcag gaaagcaacaccgggcgcgcactggcggcgatcctcgaagtgcttcacgatcatgcatttatgcaatacggca- tggtgtgtctgtttgataaagaacg caatgcactctttgtggaatccctgcatggcatcgacggcgaaaggaaaaaagagacccgccatgtccgttac- cgcatgggggaaggcgtgatcgg cgcggtgatgagccagcgtcaggcgctggtgttaccgcgcatttcagacgatcagcgttttctcgaccgcctg- aatatttacgattacagcctgccgttg attggcgtgccgatccccggtgcggataatcagccatcgggcgtgctggtggcacagccgatggcgttgcacg- aagaccggctgactgccagtacg cggtttttagaaatggtc 221 atgagcatcaccgcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggtgaacatcactgatgcacaagctacctatgtcgaagaattaactaaaaaactgca- agatgcaggcattcgcgttaaagcc gacttgagaaatgagaagattggctttaaaattcgcgaacacacgctacgccgtgttccttatatgttagtgg- tggcgataaagaggtcgaagcaggca aagttgctgttcgtacccgccgcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaactgct- ggcggaaatccgcagcagaagtctt catcaactggaggaataaagtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcattaacaaag- agattcgcgcgcaagaagttcgcc tcacaggcgtcgatggcgagcagattggtattgtcagtctgaatgaagctcttgaaaaagctgaggaagcggg- cgtcgatttagtagaaatcagtccg aatgccgagccgccagtttgtcgaatcccgtctctgaagctctcggtgaacattgttgcgaggcaggatgcga- gctggagtgttttgacattaccgata atgtgccgcgtgaacgggtgcgttatgcccccccggaagcgccgttttcccgtccggggaatggcatggagct- gcgccttatccagacgctgatcgc ccatcatcgcggttctttagatctctcggtccgccctgatggcggcaccttgctgacgttacgcctgcggtac- agcaggttatcaccggacgcttaaaa tga 222 tgtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttgttgtg- cgaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgccgttttgcacgaaaaatgcacataaattgcctg- cgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggtgaacatcactgatgcacaagctacctatgtcgaagaattaactaaaaaa- ctgcaagatgcaggcattcgcgtta aagccgacttgagaaatgagaagattggctttaaaattcgcgaacacacgctacgccgtgttccttatatgtt- agtttgtggcgataaagaggtcgaagc aggcaaagttgctgttcgtacccgccgcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaa- ctgctggcggaaatccgcagcaga agtcttcatcaactggaggaataaagtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcatta- acaaagagattcgcgcgcaagaagt tcgcctcacaggcgtcgatggcgagcagattggtattgtcagtctgaatgaagctcttgaaaaagctgaggaa- ggggcgtcgatttagtagaaatca gtccgaatgccgagccgccagtttgtcgaatcccgtctctgaagctctcggtgaacattgttgcgaggcagga- tgcgagctggttgtgttttgacattac cgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaagggcgttttcccgtccggggaatggcatgg- agctgcgccttatccagacgctg atcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggcaccttgctgacgttacgcctgc- cggtacagcaggttatcaccggaggct taaaatgacccagttacctaccgcgggcccggttatccggcgctttgatatgtctgcccagtttacggcgctt- tatcgcatcagcgtggcgctgagtcag gaaagcaacaccgggcgcgcactggcggcgatcctcgaagtgcttcacgatcatgcatttatgcaatacggca- tggtgtgtctgtttgataaagaacg caatgcactctttgtggaatccctgcatggcatcgacggcgaaaggaaaaaagagacccgccatgtccgttac- cgcatgggggaaggcgtgatcgg cgcggtgatgagccaggtcaggcgctggtgttaccgcgcatttcagacgatcagcgttttctcgaccgcctga- atatttacgattacagcctgccgttg attggcgtgccgatccccggtgcggataatcagccatcgggcgtgctggtggcacagccgatggcgttgcacg- aagaccggctgactgccagtacg cggtttttagaaatggtc 223 atggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtcgcgacacaacttgcacgtcatc- ctttattgctcgatgaactgctcgacccgc gcacgctttaccagccgattgagccgggcgcttaccgcgacgaactccgtcagtatctgatgcgggtgccaac- agaagacgaagaacagcagcttg aagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccggt- gatgaaagtcagtgaccatttaacc taccttcccgaggccattctcgatgtcgtggtgcagcatgcgtgggaacaaatggtcgtaaaatacgggcagc- ccgcgcatcttcagcaccgtgagg ggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggatct- ggtcttcctgctcgattgcgcgccgg acgtgatgacggacggcgaacgcagcatcgacggacgtcagttttatcttcggctggcgcagcgcattatgca- cttattcagcacccggacatcgca ggcattctttacgaggttgatccgcgtctgcgaccttccggcgcatccggcatgctggtcagtaccattgaag- cgtttgcagattatcaggccaatgaag cctggacgtgggagcatcaggcgctggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaatt- taacgccacgcgtcgcgacattct ttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccatagg- ggagtaaaaaagcccacgagtt tgatctgaaagccgatccgggtggcatcacggatattgaattcattgcacaatacctggttctgcgtttcgcg- catgatgagccgaagctgacgcgctg gtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgccggaagaggaagcgcgccatctg- acgcaggcttatgtgacgctgcgcg atgaaattcatcatctggcgttgcaggaacacagcgggaaagtggcgcggacagctttgctactgagcgcgcg- cagatccgtgccagctgggcaa agtggctcggctga 224 cggtactggaacagaaatcggcggatgcgcaggaaatttgttatgacacggcctgtctgaagtgcaagtt- agtgcttacttcctggctggcaacctcag gctggacgccgtttattgatgataaatctgcgaagaaactggacgcttccttcaaacgttttgctgacatcat- gctcggtcgtaccgcagggatctgaaa gaagcctttgcgcagccactgacggaagaaggttatcgcgatcagctggcgcgcctgaaacgccagatcatta- ccttccatttgcttgccggtgcttac cctgaaaaagacgtcgatgcgtatattgccggctgggtggacctgcaacaggccatcgttcagcagcaacacg- cctgggaggattcggcccgttctc acgcggtgatgatggatgctttctggttaaacgggcaacctcgttaactgactgactagcctgggcaaactgc- ccgggcttttttttgcaaggaatctgat ttcatggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtcgcgacacaacttgcacgtcatc- ctttattgctcgatgaactgctcgaccc gcgcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatctgatgcgcgggtgc- caacagaagacgaagaacagcagct tgaagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccg- gtgatgaaagtcagtgaccatttaa cctaccttgccgaggccattctcgatgtcgtggtgcagcatgcgtgggaacaaatggtcgtaaaatacgggca- gcccgcgcatcttcagcaccgtga ggggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggat- ctggtcttcctgctcgattgcgcgcc ggaggtgatgacggacggccaacgcagcatcgacggtcgtcagttttatcttcggctggcgcaggcattatgc- acttattcagcacccggacatcgt caggcattctttacgaggttgatccgcgtctccgaccttccggcgcatccggcatgctggtcagtaccattga- aggtttgcagattatcaggccaatga agcctggacgtgggagcatcaggcgctggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaa- tttaacgccacgcgtcgcgacat tctttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccat- ctggggagtaaaaaagcccacga gtttgatctgaaagccgatccgggtggcatcacggatattgaattcattgacaatacctggttctgcgtttcg- cgcatgatgagccgaagctgacgcgc tggtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgccggaagaggaagagcgccatc- tgacgcaggcttatgtgacgctgcg cgatgaaattcatcatctggcgttgcaggaacacagcgggaaagtggccgcggacagctttgctactgagcgc- gcgcagatccgtgccagctgggc aaagtggctcggctgagggtttttattcggctaacaggcgcttgtgatattatccggcgcattgtatttaccc- gatttgatttatctgttttggagtcttgggat gaaagtgactttgcctgattttcaccgcgcaggtgtgctggttgtcggtgacgtaatgttagaccgttactgg- tatggcccgaccaatcgtatttctccgga agctccggtgccggtggtgaagctcagtaccattgaagagcggcctggcggtgcagctaacgtggcgatgaac- atttcatctctgggcgcctcttcct gtctgatcggcctgaccggcgtagacgacgctgcgcgtgccctcagtgagcgtctggcagaagtgattagtta- actgcgatttcgtcgcactatccaca catcctaccatcaccaaactgcgaattttgtcccgtaaccagcaactgatccgcctcgactttgaggaaggtt- ttgaaggcgttgatctcgagccgatgct gaccaaaataga 225 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgagcaacatcc- ttcactgttttataccgtattgaacaatct
tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggtacagtaggcctctcaaaaatagataaacggctcatgtacgtgggccgtttattt- tttctacccataatcgggaaccggtgt tataatgccgcgccctcatattgtggggatttcttaatgacctatcctgggcctaaagagtagttgacattag- cggagcactaacccgtctagaagctct cggtgaacattgttgcgaggcaggatgcgagctggttgtgttttgacattaccgataatgtgccgcgtgaacg- ggtgcgttatgcccgcccggaagcg gcgttttcccgtccggggaatggcatggagctgcgccttatccagacgctgatcgcccatcatcgcggttctt- tagatctctcggtccgccctgatggcg gcaccttgctgacgttacgcctgccggtacagcaggttatcaccggaggcttaaaatga 226 tgtttcgtctcgaggccgggcaactgagcggccccgttaaaccgacctgggctggcatctgttgttgtgc- gaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacaggaatcattatcagcgccagt- ggctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgcccgttttgcacgaaaaatgcacataaattgcct- gcgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gataaaacaccccgtttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- caggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggcgaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtgcttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggtacagtagcgcctctcaaaaatagataaacggctcatgtacgtgggccgt- ttattttttctacccataatcgggaac cggtgttataatgccgcgccctcatattgtggggatttcttaatgacctatcctgggtcctaaagttgtagtt- gacattagcggagcactaacccgtctctg aagctctcggtgaacattgttgcgaggcaggatgcgagctggttgtgttttgacattaccgataatgtgccgc- gtgaacgggtgcgttatgcccgcccg gaagcggcgttttcccgtccggggaatggcatggagctgcgccttatccagacgctgatcgcccatcatcgcg- gttctttagatctctcggtccgccct gatggcggcaccttgctgacgttacgcctgccggtacagcaggttatcaccggaggcttaaaatgacccagtt- acctaccgcgggcccggttatccg gcgctttgatatgtctgcccagtttacggcgctttatcgcatcagcgtggcgctgagtcaggaaagcaacacc- gggcgcgcactggcggcgatcctcg aagtgcttcacgatcatgcatttatgcaatacggcatggtgtgtctgtttgataaagaacgcaatgcactctt- tgtggaatccctgcatggcatcgacggc gaaaggaaaaaagagacccgccatgtccgttaccgcatgggggaaggcgtgatcggcgcggtgatgagccagc- gtcaggcgctggtgttaccgc gcattttagacgatcagcgttttctcgaccgcctgaatatttacgattacagcctgccgttgattggcgtgcc- gatccccggtgcggataatcagccatcg ggcgtgctggtggcacagccgatggcgttgcacgaagaccggctgactgccagtacgcggtttttagaaatgg- tc 227 atggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtagcgacacaacttgcacgtcatc- ctttattgctcgatgaactgctcgacccgc gcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatctgatgcgggtgccaac- agaagacgaagaacagcagcttg aagccgtgcgccagttcaaacaggcccagcatttgcgtatagcagccggggatatttccggggcattgccggt- gatgaaagtcagtgaccatttaacc taccttgccgaggccattctcgatgtcgtggtgcagcatgcgtgggaacaaatggtcgtaaaatacgggcagc- ccgcgcatcttcagcaccgtgagg ggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggatct- ggtcttcagctcgattgcgcgccgg aggtgatgacggacggcgaacgcagcatcgacggacgtcagttttatcttcggctggcgcagcgcattatgca- cttattcagcacccggacatcgtca ggcattctttacgaggttgatccgcgtctgcgaccttccggcgcatccggcatgctggtcagtaccattgaag- cgtttgcagattatcaggccaatgaag cctggacgtgggagcatcaggcgctggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaatt- taacgccacgcgtcgcgacattct ttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccatctg- gggagtaaaaaagcccacgagtt tgatctgaaagccgatcccggtggcatcacggatattgaattcattgacaatacctggttctgcgtttcgcgc- atcatgagccgaagctgacgcgctg gtctgataacgtgcggatttttgtactgatggcacgatatgacatcatgcggaagaggaagcgcgccatctga- cgcaggcttatgtgacgctgcgcg atgaaattcatcatctggcgttgcacgaacacagcgggaaagtggccgcggacagctttgctactgagcgcgc- gcagatccgtgccagctgggcaa agtggctcggctga 228 cggtactggaacagaaatcggcggatgcgcaggaaatttgttatgacacggcctgtctgaagtgcaagtt- agtgcttacttcctggctggcaacctcag gctggacgccgtttattgatgataaatctgcgaagaaactggacgcttccttcaaacgttttgctgacatcat- gctcggtcgtaccgcagcggatctgaaa gaagcctttgcgcagccactgacggaagaaggttatcgcgatcagctggcgcgcctgaaacgccagatcatta- ccttccatttgcttgccggtgcttac cctgaaaaagacgtcgatgcgtatattgccggctgggtggacctgcaacaggccatcgttcagcagcaacacg- cctgggaggattcggcccgttctc acgcggtgatgatggatgctttctggttacgggcaacctcgttaactgactgactagcctgggcaaactgccc- gggcttttttttgcaaggaatctgat ttcatggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtcgcgacacaacttgcacgtcatc- ctttattgctcgatgaactgctcgaccc gcgcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatctgatgcgggtgcca- acagaagacgaagaacagcagct tgaagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccg- gtgatgaaagtcagtgaccatttaa cctaccttgccgaggccattctcgatgtcgtggtgcagaatgcgtgggaacaaatggtcgtaaaatacgggca- gcccgcgcatcttcagcaccgtga ggggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggat- ctggtcttcctgctcgattgcgcgcc ggaggtgatgacggacggcgaacgcagcatcgacggacgtcagttttcttcggaggcgcagcgcattatgcac- ttattcagcacccggacatcgt caggcattctttacgaggttgatccgcgtctgcgaccttccggcgcatccggcatgctggtcagtaccattga- agcgtttgcagattatcaggccaatga agcctggacgtgggagcatcaggcgctgtgtcgcgcgcgcgtggtttacggggatccgcaactgacacagcaa- tttaacgccacgcgtagcgacat tctttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccat- ctggggagtaaaaaagcccacga gtttgatctgaaagccgatccgggtggcatcacggatattgaattcattgcacaatacctggttctgcgtttc- gcgcatgatgagccgaagctgacgcgc tggtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgccggaagaggaaggcgccatct- gacgcaggcttatgtgacgctgcg cgatgaaattcatcatctggcgttgcaggaacacagcgggaaagtggccgcggacagctttgctactgagcgc- gcgcagatccgtgccagctgggc aaagtggctcggctgagggtttttattcggctaacaggcgcttgtgatattatccggcgcattgtatttaccc- gatttgatttatctgttttggagtcttgggat gaaagtgactttgcctgattttcaccgcgcaggtgtgctggttgtcggtgacgtaatgttagaccgttactgg- tatggcccgaccaatcgtatttctccgga agctccggtgccggtggtgaaggtcagtaccattgaagagcggcctggcggtgcagctaacgtggcgatgaac- attcatctctggacgcctcttcct gtctgatcggcctgaccggcgtagacgacgctgcgcgtgccctcagtgagcgtctggcagaagtgaaagttaa- ctgcgatttcgtcgcactatccaca catcctaccatcaccaaactgcgaattttgtcccgtaaccagcaactgatccgcctcgactttgaggaaggtt- ttgaaggcgttgatctcgagccgatgct gaccaaaataga 229 atgagcatcacggcgttatcagctgaatatcactgactcacaagctacctatgtcgaagaattaactaaa- aaactgcaagatgcaggcattcgcgttaaa gcgacttgagaaatgagaagattggctttaaaattcgcgaacacacgctacgccgtgtccttatatgttagtt- tgtggcgataaagaggtcgaagcag gcaaagttgctgttcgtactcgtcgcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaact- gctggcggaaatccgcagcagaagt catcatcaactggaggaataaagtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcattaaca- aagagattcgcgcgcaagaagttc gcctcaccggcgtcgatggcgagcagattggtattgcagtctgaatgaagctcttgaaaaagctgaggaagcg- ggcgtcgatttagtagaaatcagt ccgaatgccgagccgccagtttgtcgaatctctttagatctctcggtccgccctgatggcggcaccttgctga- cgttacgcctgccggtacagcaggtta tcaccggaggcttaaaatga 230 tgtttcgtcgaagccgggcaactgagcagccccgttgaaaccgaactgggctggcatctgttgttgtgcg- aacaaattcgcctgcgcaacccttgc cgaaagccgaggccttaacgcgggtgcgtcagcaactgattgcccggcaacagaatcattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttggtgcagacgggttaatgcccgttttgcacgaaaaatgcacataaactgcct- tcgctgccttataacagcgcatggaaatc ctgcctcctgccttgtgccacgccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgatg- tttaaaacaccccgttcagatcaaccttt gggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatatgtgatttgggttcc- ggcattgcgcaataaaggggagaaa gacatgagcatcacggcgttatcagctgaatatcactgactcacaagctacctatgtcgaagaattaactaaa- aaactgcaggatgcaggcattcgcgtt aaagccgacttgagaaatgagaagattggctttaaaattcgcgaacacacgctacgccgtgttccttatatgt- tagtttgtggcgataaagaggtcgaag caggcaaagtgctgttcgtactcgtcgcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaa- ctgctggcggagatccgcagcag aagtcatcatcaactggaggaataaagtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcatt- aacaaagagattcgcgcgcaagaa gttcgcctcaccggcgtcgatggcgagcagattggtattgtcagtctgaatgaagacttgaaaaagctgagga- agcgggcgtcgatttagtagaaatc agtccgaatgccgagccgccagtttgtcgaatctctttagatctctcggtccgccctgatggcggcaccttgc- tgacgttacgcctgccggtacagcag gttatcaccggaggcttaaaatgacccagttacctaccgcgggcccggttatccggcgctttgatatgtctgc- ccagtttacggcgctttatcgcatcagc gtggcgctgagtcaggagagcaataccgcgcgcgcactggcggcgatcctcgaagtgcttcacgatcatgcat- ttatgcaatacggcatggtgtgtct gttcgataaagaacgcaatgcactgtttgtgcaatccctgcatggcatcgacggcgaaaggaaaaaagaaacc- cgccatgtccgttaccgcatgggg gaaggcgtgatcggcgcggtgatgagccagcgtcaggcgtggtgttaccgcgcatttcagacgatcagcgttt- tctcgaccgcctgaatatttacgat tacagcctgccgctgattggtgtgccgatccccggtgcggataatcagcctgcgggtgtgctggtggcacagc- cgatggcgttgcacgaagaccgg ctggctgccactacgcgctttttagaaatggtc 231 atgaccctgaatatgatgatggatgccggcggacatcatcgcgacaaacaatattaataccggcaaccac- accggcaatttacgagactgcgcaggc atcctttacccgtcaatttctgtcaaataaagtaaaagaggcagtctacttgaattacccccggctggttgag- cgtttgttgaaaaaaagtaactgaaaa tccgtagaatagcgccactctgatggttaattaacctattcaattaagaattatctggatgaatgtgccatta- aatgcgcagcataatggtgcgttgtgcgg gaaaactgcttttttttgaaagggttggtcagtagggaaacaactcacttcacaccccgaagggggaagttgc- ctgaccctacgattcccgctatttcat tcactgaccggaggttcaaaatga 232 accggatacgagagaaaagtgtctacatttggttcggttgatattgaccggcgcatccgccagcccgccc- agtttctggggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaaacctgcaccagtttggttattaatgcaccag- tctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggttttgtcgg- gttatcagccaaaaggtgcactcttt gcatggttatacgtgcctgacatgttgtccgggcgacaaacggcctggtggcacaaattgtcagaactacgac- acgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgcccgcggacatcatcgcgacaaacaatattaataccggcaaccac- accggcaatttacgagactgcgca ggcatcctttctcccgtcaatttctgtcaaataaagtaaaagaggcagtctacttgaattacccccggctggt- tgagcgtttgttgaaaaaaagtaactgaa aaatccgtagaatagcgccactctgatggttaattaacctattcaattaagaattatctggatgaatgtgcca- ttaaatgcgcagattaatggtgcgttgtg cgggaaaactgctttttttgaaagggttggtcagtagcggaaacaactcacttcacaccccgaagggggaagt- tgcctgaccctacgattcccgctatt tcattcactgaccggaggttcaaaatgacccagcgaaccgagtcgggtaataccgtctggcgcttcgatttgt- cccagcagttcactgcgatgcagcgc ataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccagcaagtgctgtgcgtattgcacaatg- acgcctttttgcagcacggcatgat ctgtctgtacgacagccagcaggcgattttgaatattgaagcgttgcaggaagccgatcagcagttaatcccc- ggcagctcgcaaatccgctatcgtcc gggcgaagggctggtcgggacggtgctttcgcagggccaatcattagtgcttgcgcgcgttgctgacgatcag- cgctttcttgaccggctcgggttgt atgattacaacctgccgtttatcgccgtgccgctgatagggccagatgcgcagactttcggtggctgacggca- caacccatggcgcgttacgaagag cgattacccgcctgcacccgctttctggaaacggtc 233 atgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatattccc gcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccgatg- gaagctcgctgttttaacacgcgttttttaa ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatattct- caacctaaaaaagtttgtgtaatacttgtaac gctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggactgggcgcaactcac- ttcacaccccgaagggggaagttgc ctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatga 234 accggatacgagagaaaagtgtctacatcggttcggttgatattgaccggcgcatccgccagcccgccca- gtttctggtggatctgtttggcgattttgc gggtcttgccggtgcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagatc- ccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaaacctgcaccagtttggttattaatgcaccag- tctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggttttgtcgg- gttatcagccaaaaggtgcactcttt gcatggttatacgtgcctgacatgttgtccgggcgacaaacggcctggtggcacaaattgtcagaactacgac- acgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgccggccgtcctgtaataataaccgcacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatatt cccgcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccg- atggaagctcgctgttttaacacgcgttttt taaccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatat- tctcaacctaaaaaagtttgtgtaatacttgt aacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaac- tcacttcacaccccgaagggggaagt tgcctgaccctacgattcccgctatttcattcactgaccggaggttcataatgacccagcgaaccgagtcggg- taataccgtctggcgcttcgatttgtc ccagcagttcactgcgatgcagcgcataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccag- caagtgctgtgcgtattgcacaat gacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgaatattgaagcgttgc- aggaagccgatcagcagttaatcccc ggcagctcgcaaatccgctatcgtccgggcgaagggctggtcgggacggtgtttcgcagggcaatcattagtg- ctggcgcgcgttgctgacgatc agcgctttcttgaccggctcgggttgtatgattacaacctgccgtttatcgccgtgccgctgatagggccaga- tgcgcagactttcggtgtgctgacggc acaacccatggcgcgttacgaagagcgattacccgcctgcacccgctttctggaaacggtc 235 atgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgcatttttttattccc gcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccgatg- gaagctcgctgttttaacacgcgtttttaa
ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatattct- caacctaaaaaagtttgtgtaatacttgtaac gctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaactca- cttcacaccccgaagggggaagttgc ctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatga 236 accggatacgagagaaaagtgtctacatcggttcggttgatattgaccggcgcatccgccagcccgccca- gtttctggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaaacctgcaccagtttggttattaatgcaccag- tctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggttttgtcgg- gttatcagccaaaaggtgcactcttt gcatggttatacgtgcctgacatgttgtccgggcgacaaacggcctggtggcacaaattgtcagaactacgac- acgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatatt cccgcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccg- atggaagctcgctgttttaacacgcgttttt taaccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatat- tctcaacctaaaaaagtttgtgtaatacttgt aacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaac- tcacttcacaccccgaagggggaagt tgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcggg- taataccgtctggcgatcgatttgtc ccagcagttcactgcgatgcagcgcataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccag- caagtgctgtgcgtattgcacaat gacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgaatattgaagcgttgc- aggaagccgatcagcagttaatcccc ggcagctcgcaaatccgctatcgtccgggcgaagggctggtcgggacggtgctttcgcagggccaatcattag- tgctggcgcgcgttgctgacgatc agcgctttcttgaccggctcgggttgtatgattacaacctgccgtttatcgccgtgccgctgatagggccaga- tgcgcagactttcggtgtgctgacggc acaacccatggcgcgttacgaagagcgattacccgcctgcacccgctttctggaaacggtc 237 atggcactgaaacacctcatttccctgtgtgccgcgtcgccgatggttgccagtcagctggcgcgctacc- cgatcctgcttgatgaattgctcgacccg aatacgctctatcaaccgacggcgatgaatgcctatcgcgatgagctgcgccaatacctgctgcgcgtgccgg- aagatgatgaagagcaacagcttg aggcgctgcggcagtttaagcaggcgcagttgctgcgcgtggcggcggcggatattgccggtacgttgccagt- aatgaaagtgagcgatcacttaac ctggctggcggaagcgattattgatgcggtggtgcagcaagcctgggggcagatggtggcgcgttatggccag- ccaacgcatctgcacgatcgcga agggcgggttttgcggtggtcggttatggcaagctgggcggctgggagctgggttacagctccgatctggatc- tggtattcctgcacgactgcccga tggatgtgatgaccgatggcgagcgtgaaatcgatggtcgccagttctatttgcgtctcgcgcagcgcgtgat- gcacctgtttagcacgcgcacgtcgt ccggcatcctttatgaagttgatgcgcgtctgcgtccatctggcgctgcggggatgctggtcactactacgga- atcgttcgccgattaccagcaaaacg aagcctggacgtgggaacatcaggcgctggcccgtgcgcgcgtggtgtacggcgatccgcaactgaccgccga- atttgacgccattcgccgcgata ttctgatgacgcctcgcgacggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagaaaatgcgtgccca- tcttggcaacaagcataaagacc gcttcgatctgaaagccgatgaaggcggtatcaccgacatcgagtttatcgcccaatatctggtgctgcgctt- tgcccatgacaagccgaaactgacgc gctggtcggataatgtgcgcattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggc- attgacgctggcgtacaccaca ttgcgtgatgagctgcaccacctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccg- agcgtgcgcttattaaaaccagctgg gacaagtggctggtggaa 238 gcgcaaagcgagtgctcacttacgtgatctgttgacacaatctgaagcgaccataacttctgccgtttca- gcgaatacggcggtgtggagcgcacaatc agccctggcgaagctggtgctcaccgagtggctagtgacgcagggctggcgaaccttccttgatgaaaaagcg- caggccaaattcgccgactccttt aaacgctttgctgacatccatctgtcacgcagcgccgccgagctgaaaaaaggcctttgcccaaccgctgggc- gacagctatcgcgaccagttgccgc gcctggcgcgtgatatcgactgcgcgttactgctggccgggcattacgatcgcgcgcgcgccgtggaatggct- ggaaaactggcaggggcttcagc acgccattgaaacgcgccagagagtcgaaatcgaacatttccgtaataccgcgattacccaggagccgttctg- gttgcacagcggaaaacgttaacg aaaggatatttcgcatggcactgaaacacctcatttccctgtgtgccgcgtcgccgatggttgccagtcagct- ggcgcgctacccgatcctgcttgatga attgctcgacccgaatacgctctatataccgacggcgatgaatgcctatcgcgatgagctgcgcatatacctg- ctgcgcgtgccggaagatgatgaa gagcaacagcttgaggcgctgcggcagtttaagcaggcgcagttgctgcgcgtggcggcggcggatattgccg- gtacgttgccagtaatgaaagtg agcgatcacttaacctggctggcggaagcgattattgatgcggtggtgcagcaagcctgggggcagatggtgg- cgcgttatggccagccaacgcat ctgcacgatcgcgaagggcgcggttttgcggtggtcggttatggcaagctgggcggctgggagctgggttaca- gctccgatctggatctggtattcct gcacgactgcccgatggatgtgatgaccgatggcgagcgtgaaatcgatggtcgccagttctatttgcgtctc- gcgcagcgcgtgatgcacctgtttag cacgcgcacgtcgtccggcatcctttatgaagttgatgcgcgtctgcgtccatctggcgctgcggggatgctg- gtcactactacggaatcgttcgccga ttaccagcaaaacgaagcctgcacgtgggaacatcaggcgctggcccgtgcgcgcgtggtgtacggcgatccg- caactgaccgccgaatttgacg ccattcgccgcgatattctgatgacgcctcgcgacggcgcaacgctgcaaaccgacgtgcgagaaatgcgcga- gaaaatgcgtgcccatcttggca acaagcataaagaccgcttcgatctgaaagccgatgaaggcggtatcaccgacatcgagtttatcgcccaata- tctggtgctgcgctttgcccatgaca aaccgaaactgacgcgctggtcggataatgtgcgcattctcgaagggctggcgcaaaacggcatcatggagga- gcaggaagcgcaggcattgacg ctggcgtacaccacattgcgtgatgagagcaccacctggcgctgcaagagttgccgggacatgtggcgctctc- ctgttttgtcgccgagcgtgcgctt attaaaaccagctgggacaagtggctggtggaaccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaa- ttttgtatctctcaggagacaggaa tgaaagtgacgagccagagtttaagcaagccggtgtaatggtggtgggtgatgtgatgctggatcgttactgg- tatggcccaaccagccgtatctctc cggaagcgccagtcccggttgttaaagtcgataccattgaagagcgtcctggcggcgcggcaaacgtggcgat- gaatatcgcctcactgggcgcca cggcgcgtctggttggcctgactggcattgacgatgcggcgcgcgcgctgagcaaagcgctggccgatgttaa- cgttaaatgtgacttcgtttctgttc cgacgcatcccaccatcactaagctgcgcgtgctgtcgcgtaaccagcagctgattcgcctggactttgaaga- gggttttgaaggagtcgatccgcaa ccgatgcatcgaacgcatcagccaggcgcttggtaatattggcgcgctggtgctgtcggatt 239 atgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctgagagctggcgcg- aattgtggcaggatgcgttgcaggagga ggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgctgattgccgatttt- cgcaaagagttggataaacgcacc attggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgtatgctcgcgcgacg- atgcgccagtaccgctgtcacgcc tgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgctaagtgaatttcccggcgcact- gaaacacctcatttccctgtgtgccgc gtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcgacccgaatacgctc- tatcaaccgacggcgatgaatgccta tcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagcttgaggcgctgcgg- cagtttaagcaggcgcagttgct gcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcacttaacctggctggcg- gaagcgattattgatgcggtggtg cagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgcacgatcgcgaagggcgcggtt- ttgcggtggtcggttatggcaag ctgggcggctgggagctgggttacagctccgatctggatctggtattcctgcacgactgcccgatggatgtga- tgaccgatggcgagcgtgaaatcg atggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgcacgtcgtccggcat- cctttatgaagttgatacgcgtctgcgt ccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagcaaaacgaagcctgga- cgtgggaacatcaggcgctggcc cgtgcacgcgtgggtacggcgatccgcaactgaccgccgaatttgacgccattcgccgcgatattctgatgac- gcctcgcgacggcgcaacgctgc aaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaagaccgcttcgatct- gaaagccgatgaaggcggtatca ccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaagccgaaactgacgcgctggtc- ggataatgtgcgcattctcgaagggc tggcgcaaanggcatcatggaggagcaggaagcgcaggcattgacgctggcgtacaccacattgcgtgatgag- ctgcaccacctggcgctgcaa gagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttattaaaaccagctgggacaagt- ggctggtggaaccgtgcgccccggc gtaa 240 atgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatattccc gcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccgatg- gaagctcgctgtttaacacgcgttttttaa ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatattct- caacctaaaaaagtttgtgtaatacttgtaac gctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaactca- cttcacaccccgaagggggaagttgc ctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatga 241 accggatacgagagaaaagtgtctacatcgatcggttgatattgaccggcgcatccgccagcccgcccag- tttctggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaaacctgcaccagtttggttattaatgcaccag- tctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggttttgtcgg- gttatcagccaaaaggtgcactcttt gcatggttatacgtgcctgacatgttgtccggccgacaaacggcctggtggcacaaattgtcagaactacgac- acgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatatt cccgcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccg- atggaagctcgctgttttaacacgcgttttt taaccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatat- tctcaacctaaaaaagtttgtgtaatacttgt aacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaac- tcacttcacaccccgaaggggcaagt tgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcggg- taataccgtctggcgcttcgatttgtc ccagcagttcactgcgatgcagcgcataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccag- caagtgctgtgcgtattgcacaat gacgcctttttgcagcacggcatgatctgtctgtacgacagccagcagccgattttgaatattgaagcgttgc- aggaagccgatcagcagttaatcccc ggcagctcgcaaatccgtatcgtccgggcgaagggctggtcgggacggtgctttcgcagggccaatcattagt- gctggcgcgcgttgctgacgatc agcgctttcttgaccggctcgggttgtatgattacaacctgccgtttatcgccgtgccgctgatagggccaga- tgcgcagactttcggtgtgctgacggc acaacccatggcgcgttacgaagagcgattacccgcctgcacccgctttctggaaacggtc 242 gcgcaaagcgagtgctcacttacgtgatctgttgacacaatctgaagcgaccataacttctgccgtttca- gcgaatacggcggtgtggagcgcacaatc agccctggcgaagctggtgctcaccgagtggctagtgacgcagggctggcgaaccttccttgatgaaaaagcg- caggccaaattcgccgactccttt aaacgctttgctgacatccatctgtcacgcagcgccgccgagctgaaaaaagcctttgcccaaccgctgggcg- acagctatcgcgaccagttgccgc gcctggcgcgtgatatcgactgcgcgttactgctggccgggcattacgatcgcgcgcgcgccgtggaatggct- ggaaaactggcaggggcttcagc acgccattgaaacgcgccagagagtcgaaatcgaacatttccgtaataccgcgattacccaggagccgttctg- gttgcacagcggaaaacgttaacg aaaggatatttcgcatgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctga- gagctggcgcgaattgtggcaggatg cgttgcaggaggaggattccacgcccgtgaggcgcatctctcagaggacgatcgccgccgcgtggtggcgctg- attgccgattttcgcaaagagtt ggataaacgcaccattggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgta- tgacgcgcgacgatgcgccagta ccgctgtcacgcctgacgccgctgacaccggaattattacccgcaccacttaccttgagctgctaagtgaatt- tcccggcgcactgaaacacctcattt ccctgtgtgccgcgtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcga- cccgaatacgctctatcaaccgacg gcgatgaatgcctatcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagc- ttgaggcgctgcggcagtttaag caggcgcagttgctgcgcgtcgcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcact- taacctggctggcggaagcgatta ttgatgcggtggtgcagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgcacgatcg- cgaagggcgcggttttgcggtgg tcggttatggcaagctgggcggctgggagctgggttacagctccgatctggatctggtattcctgcacgactg- cccgatggatgtgatgaccgatggc gagcgtgaaatcgatggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgca- cgtcgccggcatcctttatgaagttg atgcgcgctgcgtccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagcaa- aacgaagcctggacgtgggaaca tcaggcgctggcccgtgcgcgcgggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgccgcg- atattctgatgacgcctcgcgac ggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaag- accgcttcgatctgaaagccgat gaaggcggtatcaccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaagccgaaac- tgacgcgctggtcggataatgtgcg cattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctggcgtacacc- acattgcgtgatgagctgcacc acctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttattaaaac- cagctgggacaagtggctggtggaa ccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaattttgtatctctcaggagacaggaatgaaagtg- acgctgccagagtttaagcaagccg gtgtaatggtggtgggtgatgtgatgctggatcgttactggtatggcccaaccagccgtatctctccggaagc- gccagtcccggttgttaaagtcgatac cattgaagagcgtcctggcgccgcggcaaacgtggcgatgaatatcgcctcactgggcgccacggcgcgtctg- gttggcctgactggcattgacga tgcggcgcgcgcgctgagcaaagcgctggccgatgttaacgttaaatgtgacttcgtttctgttccgacgcat- cccaccatcactaagctgcgcgtgct gtcgcgtaaccagcagctgattcgcctggactttgaagagggttttgaaggagtcgatccgcaaccgatgcat- gaacgcatcagccaggcgcttggta atattggcgcgctggtgctgtcggatt 243 atgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctgagagctggcgcg- aattgtggcaggatgcgttgcaggagga ggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgctgattgccgatttt- cgcaaagagttggataaacgcacc attggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgtatgctcgcgcgacg- atgcgccagtaccgctgcacgcc tgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgctaagtgaatttcccggcgcact- gattacacctcatttccctgtgtgccgc gtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcgacccgaatacgctc- tatcaaccgacggcgatgaatgccta tcggatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagcttgaggcgctgcggc- agtttaagcaggcgcagttgct gcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcacttaacctggctggcg- gaagcgattattgatgcggtggtg cagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgacgatcgcgaagggcgcggttt-
gcggggtcggttatggcaag ctgggcggctgggagctgggtacagctccgatctggatctggtattcctgcacgactgcccgatggatgtgat- gaccgatggcgagcgtgaaatcg atggtcgccagtctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgacgtcgtccggcatcc- tttatgaagttgatgcgcgtctgcgt ccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagcaaaacgaagcctgga- cgtgggaacatcaggcgctggcc cgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgccgcgatattctgatga- cgcctcgcgacggcgcaacgctgc aaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaagaccgcttcgatct- gaaagccgatgaaggcggtatca ccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaagccgaaactgacgcgctggtc- ggataatgtgcgcattctcgaagggc tggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctggcgtacaccacattgcgtgatga- gctgcaccacctgggctgcaa gagttgccgggacatgtggcgctctcctgtttgtcgccgagcgtgcgcttattaaaaccagctgggacaagtg- gctggtggaaccgtgcgccccggc gtaa 244 gcgcaaagcgagtgctcacttacgtgatctgttgacacaatctgaagcgaccataacttctgccgtttca- gcgaatacggcggtgtggagcgcacaatc agccctggcgaagctggtgctcaccgagtggctagtgacgcagggctggcgaaccttccttgatgaaaaagcg- caggccaaattcgccgactccttt aaacgctttgctgacatccatctgtcacgcagcgccgccgagctgaaaaaagcctttgcccaaccgctgggcg- acagctatcgcgaccagttgccgc gcctggcgcgtgatatcgactgcgcgttactgctggccgggcattacgatcgcgcgcgcgccgtggaatggct- ggaaaactggcaggggcttcagc acgccattgaaacgcgccagagagtcgaaatcgaacatttccgtaataccgcgattacccaggagccgttctg- gttgcacagcggaaaacgttaacg aaaggatatttcgcatgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctga- gagctggcgcgaattgtggcaggatg cgttgcaggaggaggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgct- gattgccgattttcgcaaagagtt ggataaacgcaccattggcccgcgagggcggaaggtactcgatcacttaatgccgcatctgctcagcgatgta- tgctcgcgcgacgatgcgccagta ccgctgtcacgcctgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgctaagtgaat- ttcccggcgcactgaaacacctcattt ccctgtgtgccgcgtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcga- cccgaatacgctctatcaaccgacg gcgatgaatgcctatcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagc- ttgaggcgctgcggcagtttaag caggcgcagttgctgcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcact- taacctggctggcggaagcgatta ttgatgcggtgggcagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatagcacgatcgcg- aagggcgcggttttgcggtgg tcggttatggcaagctgggcggctgggagctgggttacagctccgatctggatctggtattcctgacgactgc- ccgatggatgtgatgaccgatggc gagcgtgaaatcgatggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgca- cgtcgtccggcatcctttatgaagttg atgcgcgtctgcgtccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagca- aaacgaagcctggacgtgggaaca tcagccgctggcccgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgccgc- gatattctgatgacgcctcgcgac ggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagagaaaatgcgtgcccatcttggcaacaagcataa- agaccgcttcgatctgaaagccgat gaaggcggtatcaccgacatcgagtttatcgcccaatatctgggctgcgctttgcccatgacaagccgaaact- gacgcgctggcggataatgtgcg cattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctggcgtacacc- acattgcgtgatgagctgcacc acctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttattaaaac- cagctgggacaagtggctggtggaa ccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaattttgtatctctcaggagacagcaatgaaagtg- acgctgccagagtttaagcaagccg gtgtaatggtggtgggtgatgtgatgctggatcgttactggtatggcccaaccagccgtatctctccggaagc- gccagtcccggttgttaaagtcgatac cattgaagagcgtcctggcggcgcggcaaacgtggcgatgaatatcgcctcactgggcgccacggcgcgtctg- gttggcctgactggcattgacga tgcggcgcgcgcgctgagcaaagcgctggccgatgttaacgtaaatgtgacttcgtttctgttccgacgcatc- ccaccatcactaagctgcgcgtgct gtcgcgtaaccagcagctgattcgcctggactttgaagagggttttgaaggagtcgatccgcaaccgatgcat- gaacgcatcagccaggcgcttggta atattggcgcgctggtgctgcggatt 245 atgaccctgaatatgatgatggatgccggcggacatcatcgcgacaaacaatattaataccggcaaccac- accggcaatttacgagactgcgcaggc atcctttctcccgtcaatttctgtcaaataaagtaaaagaggcagtctacttgaattacccccggctggttca- gcgtttgttgaaaaaaagtaactgaaaaa tccgtagaatagcgccactctgatggtaattacctattcaattaagaattatctggatgaatgtgccattaaa- tgcgcagcataatggtggttgtgcgg gaaaactgcttttttttgaaagggttggtcagtagcggaaacaactcacttcacaccccgaagggggaagttg- cctgaccctacgattcccgctatttcat tcactgaccggaggttcaaaatga 246 accggatacgagagaaaagtgtctacatcggttcggttgatattgaccggcgcatccgccagcccgccca- gtttctggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaanataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaacctgcaccagtttggttattaatgcaccagt- ctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggttttgtcgg- gttatcagccaaaggtgcactcttt gcatggtatacgtgcctgacatgttgtccgggcgacaaacggcctggtggcacaaattgtcagaactacgaca- cgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgccggcggacatcatcgcgacaaacaatattaataccggcaaccac- accggcaatttacgagactgcgca ggcatcctttctcccgtcaatttctgcaaataaagtaaaagaggcagtctacttgaattacccccggctggtt- gagcgtttgttgaaaaaaagtaactgaa aaatccgtagaatagcgccactctgatggttaattaacctattcaattaagaattatctggatgaatgtgcca- ttaaatgcgcagcataatggtgcgttgtg cgggaaaactgcttttttttgaaagggttggtcagtagcggaaacaactcacttcacaccccgaagggggaag- ttgcctgaccctacgattcccgctatt tcattcactgaccggaggttcaaaatgacccagcgaaccgagtcgggtaataccgtaggcgcttcgatttgtc- ccagcagttcactgcgatgcagcgc ataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccagcaagtgctgtgcgtattgacaatga- cgcctttttgcagcacggcatgat ctgctgtacgacagccaccaggcgattttgaatattgaagcgttgcaggaagccgatcagcagttaatccccg- gcagctcgcaaatccgctatcgtcc gggcgaagggctggtcgggacggtgctttcgcagggccaatcattagtgctggcgcgcgttgctgacgatcag- cgctttcttgaccggctcggttgt atgattacaacctgccgtttatcgccgtgccgctgatagggccagatgcgcagactttcggtgtgctgacggc- acaacccatggcgcgttacgaagag cgattacccgcctgcacccgctttctggaaacggtc 247 atgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatattccc gcctccatttaaaataaaaaatccaatcggattcactatttaaactggccattatctaagatgaatccgatgg- aagctcgctgttttaacacgcgttttttaa ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatattct- caacctaaaaaagtttgtgtaatacttgtaac gctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaactca- cttcacaccccgaagggggaagttgc ctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatga 248 accggatacgagagaaaagtgctacatcggttcggttgatattgaccggcgcatccgccagcccgcccag- tttctggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaaacctgcaccagtttggttattaatgcaccag- tctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggtttgtcggg- ttatcagccaaaaggtgcactcttt gcatggttatacgtgcctgacatgttgtccgggcgacaaacggcctggtggcacaaattgtcagaactacgac- acgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatatt cccgcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccg- atggaagctcgctgttttaacacgcttttt taaccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatat- tctcaacctaaaaaagtttgtgtaatacttgt aacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaac- tcacttcacaccccgaagggggaagt tgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcggg- gtaataccgtctggcgcttcgatttgtc ccagcagttcactgcgatgcagcgcataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccag- caagtgctgtgcgtattgcacaat gacgccttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgaatattgaagcgttgca- ggaagccgatcagcagttaatcccc ggcagctcgcaaatccgctatcgtccgggcgaagggctggtcgggacggtgctttcgcagggccaatcattag- tgctggcgcgcgttgctgacgatc agcgctttcttgaccggctcgggttgatgattacaacctgccgtttatcgccgtgccgctgatagggccagat- gcgcagactttcggtgtgctgacggc acaacccatggcgcgttacgaagagcgattcccgcctgcacccgctttctggaaacggtc 249 atggcactgaaacacctcattccctgtgtgccgcgtcgccgatggttgccagtcagctggcgcgctaccc- gatcctgcttgatgaattgctcgacccg aatacgctctatcaaccgacggcgatgaatgcctatcgcgatgagagcgccaatacctgctgcgcgtgccgga- agatgatgaagagcaacagcttg aggcgctgcggcagtttaagcaggcgcagttgctgcgcgtggcggcggcggatattgccggtacgttgccagt- aatgaaagtgagcgatcacttaac ctggctggcggaggcgattattgatgcggtggtgcagcaagcctgggggcagatggtggcgcgttatggccag- ccaacgcatctgcacgatcgcga agggcgcggttttgcggtggtcggttatggcaagctgggcggctgggagctgggttacagctccgatctggat- ctggtattcctgcacgactgcccga tggatgtgatgaccgatggcgagcgtgaaatcgatggtcaccagttctatttgcgtctcgcgcagcgcgtgat- gcacctgtttagcacgcgcacgtcgt ccggcatcctttatgaagttgatgcgcgtctgtccatctggcgctgcggatgctggtcactactacggaatcg- ttcgccgattaccagcaaaacg aagcctggacgtgggaacatcaggcgctggcccgtgcgcgcgtggtgtacggcgatccgcaactgaccgccga- attgacgccattcgccgcgata ttctgatgacgcctcgcgacggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagaaaatgcgtgccca- tcttggcaacaagcataaagacc gcttcgatctgaaagccgatgaaggcggtatcaccgacatcgagtttatcgcccaatatctggtgctgcgctt- tgcccatgacaagccgaaactgacgc gctggtcggataatgtgcgcattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggc- attgacgctggcgtacaccaca ttgcgtgatgagctgcaccacctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccg- agcgtgcgcttattaaaaccagctgg gacaagtggctggtggaa 250 gcgcaaagcgagtgctcacttacgtgatctgttgacacaatctgaagcgaccataacttctgccgtttca- gcgaatacggcggtgtggagcgcacaatc agccctggcgaagctggtgctcaccgagtggctagtgacgcagggctggcgaaccttccttgatgaaaaagcg- caggccaaatcgccgactccttt aaacgctttgctgacatccatctgtcacgcagcgccgccgagctgaaaaaagcctttgcccaaccgctgggcg- acagctatcgcgaccagttgccgc gcctggcgcgtgatatcgactgcgcgttactgctggccgggcattacgatcgcgcgcgcgccgtggaatggct- ggaaaactggcaggggcttcagc acgccattgaaacgcgccagagagtcgaaatcgaacatttccgtaataccgcgattacccaggagccgttctg- gttgcacagcggaaaacgtttacg aaaggatatttcgcatggcactgaaacacctcatttccctgtgtgccgcgtcgccgatggttgccagtcagct- ggcgcgctacccgatcctgcttgatga attgctcgacccgaatacgctctatcaaccgacggcgatgaatgcctatcgcgatgagctgcgccaatacctg- ctgcgcgtgccggaagatgatgaa gagcaacagcttgaggcgctgcggcagtttaagcaggcgcagttgctgcgcgtggcggcgggatattgccggt- acgttgccagtaatgaaagtg agcgatcacttaacctggctggcggaagcgattattgatgcggtggtgcagcaagcctgggggcagatggtgg- cgcgttatggccagccaacgcat ctgcacgatcgcgaagggcgcggttttgcggtggtcggttatggcaagagggcggctgggagctgggttacag- ctccgatctggatctggtattcct gcacgactgcccgatggatgtgatgaccgatggcgagcgtgaaatcgatggtcgcagttctatttgcgtctcg- cgcagcgcgtgatgcacctgttag cacgcgcacgtcgtccggcatcctttatgaagttgatgcgcgtctgcgtccatctagcgctgcggggatgctg- gtcactactacggaatcgttcgccga ttaccagcaaaacgaagcctggacgtgggaacatcaggcgctggcccgtgcgcgtggtgtacggcgatccgca- actgaccgccgaatttgacg ccattcgccgcgatattctgatgacgcctcgcgacggcgcaacgctgcaaaccgacgtgcgagaaatgcgcga- gaaaatgcgtgcccatcttggca acaagcataaagaccgcttcgatctgaaagccgatgaaggcggtatcaccgacatcgagtttatcgcccaata- tctggtgctgcgctttgcccatgaca agccgaaactgacgcgctggtcggataatgtgcgcattctcgaacggctggcgcaaaacggcatcatggagga- gcaggaagcgcaggcattcacg ctggcgtacaccacattgcgtgatgagctgcaccacctggcgctgcaagagttgccgggacatgtggcgctct- cctgttttgtcgccgagcgtgcgctt attaaaaccagctgggacaagtggctggtggaaccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaa- ttttgtatctctcaggagacaggaa tgaaagtgacgctgccagagtttaagcaagccggtgtaatggtggtgggtgatgtgatgctggatcgttactg- gtatggcccaaccagccgtatctctc cggaaggccagtcccggttgttaaagtcgataccattgaagagcgtcctggcggcgcggcaaacgtggcgatg- aatatcgcctcactgggcgcca cggcgcgctggttggcctgactggcattgacgatgcggcgcgcgcgctgagcaaagcgctggccgatgttaac- gttaaatgtgacttcgtttctgttc cgacgcatcccaccatcactaagctgcgcgtgctgtcggtaaccagcagctgattcgcctggactttgaagag- ggttttgaaggagtcgatccgcaa ccgatgcatgaacgcatcagccaggcgcttggtaatattggcgcgctggtgctgtcggatt 251 tttcgctgaaggtgtgaccatgggccatcaggtgctggtgcagctggaaagtgttgccatcactatcgtg- tggtctggcgtggtggcctttattggttaca aactggcggacatgacggtaggcctgcgcgtaccggaagaacaagaacgtgaagggctggatgaaacagccac- ggcgaaaacgcctataacgc ctga 252 tttcctttctgactctgcccgtccgggcgcactaacggcctgaaatactccctcttttcattcctggcac- aacgattaaatgtagttgcgtgttagctgcg gccattatcgaattcgactggagggggatctatgaagctggttaccgtggtgattaagccattcaaacttgaa- gacgtgcgtgaagcgctttcttctattg gtattcaagggttgaccgtaactgaagtgaaaggctttggccgtcagaagggtcacgctgagctgtaccgcgg- tgcggaatatagcgttaatttcctgc cgaaagtgaaaattgatgtggcgatcgctgacgatcaactcgatgaagtaatcgatgtgatcagcaaagcggc- ctacaccggaaaaattggcgacgg caaaattttcgttgctgagctgcaacgcgtcattcgtattcgtaccggcgaagccgacgaagcggcactgtaa- tacaagacacacagtgatggggatc ggtttcgctgaaggtgtgaccatgggccatcaggtgctggtgcagctggaaagtgttgccatcactatcgtgt- ggtctggcgtggtggcctttattggtta caaactggcggacatgacggtaggcctgcgcgtaccggaagaacaagaacgtgaagggctggatgtaaacagc- cacggcgaaaacgcctataac gcctgattgcgttgagttatctcctgagcataaaaaagcctccattcggaggcttttctttttttaagtttaa- aggcggttagttgcgattgcgcatgacgcc ttcctgcacgctggacgcgaccagcacaccctcttgcgtatagaactcgccgcgcacattaaccgcgagcgct- ggaggctgacgtgctttccacactg tagagcagccattcgttcatattaaacgggcgatggaaccacatggagtggtcaatggtggcaacctgcatac- cgcgctcaaggaagcccacgccgt
gcggctgaagtgcaaccggcaggaagttaaagtctgaggcatatccaagcagatattgatgtacgcgaaaatc- gtccacaccgtgccgtttgcgcg gatccatacctggcgggtgggatcggcaacgtggcctttcagcgggttatgaaactcaaccgggcggatctcc- agtggtttatcactaagaaacttctc tttggcctgcgg 253 atgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctgagagctggcgcg- aattgtggcaggatgcgttgcaggagga ggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgctgattgccgatttt- cgcaaagagttggataaacgcacc attggcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgtatgctcgcgcgacg- atgcgccagtaccgctgtcacgcc tgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgctaagtgaatttcccggcgcact- gaaacacctcatttccctgtgtgccgc gtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcgacccgaatacgctc- tatcaaccgacggcgatgaatgccta tcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagcttgaggcgctgcgg- cagtttaagcaggcgcagttgct gcgcgggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcacttaacctggctggcgg- aagcgattattgatgcggtggtg cagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgcacgatcgcgaagggcgcggtt- ttgcggtggtcggttatggcaag ctgggcggctgggagctgggttacagatccgatctggatctggtattcctgcacgactgcccgatggatgtga- tgaccgatggcgagcgtgaaatcg atggtcgccagttctatttgcgtctcccgcagcgcgtgatgcacctgtttagcacgcgcacgtcgtccggcat- cctttatgaagttgatgcgcgtctgcgt ccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagcaaaacgaagcctgga- cgtgggaacatcaggcgctggcc cgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgccgcgatattctgatga- cgcctcgcgacggcgcaacgctgc aaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaagaccgcttcgatct- gaaagccgatgaaggcggtatca ccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaagccgaaactgacgcgctggtc- ggataatgtgcgcattctcgaagggc tggcgcaaaacggcatcatggacgagcaggaagcgcaggcattgacgctggcgtacaccacattgcgtgatga- gctgaccacctggcgctgcaa gagttgccgggacatgggcgctctcctgttttgtcgccgagcgtgcgcttattaaaaccagctgggacaagtg- gctggtggaaccgtgcgccccggc gtaa 254 gcgcaaagcgagtgctcacttacgtgatctgttgacacaatctgaaggaccataacttctgccgtttcag- cgaatacggcggtgtggaggcacaatc agccctggcgaagctggtgctcaccgagtggctagtgacgcagggctggcgaaccttccttgatgaaaaagcg- caggccaaattcgccgactccttt aaacgctttgctgacatccatctgtcacgcagcgccgccgagctgaaaaaagcctttgcccaaccgctgggcg- acagctatcgcgaccagttgccgc gcctgccgcgtgatatcgactgcgcgttactgctggccgggcattacgatcgcgcgcgcgccgtggaatggct- ggaaaactggcaggggcttcagc acgccattgaaacgcgccagagagtcgaaatcgaacatttccgtaataccgcgattacccaggagccgttctg- gttgcacagcggaaaacgttaacg aaaggatatttcgcatgtttaacgatctgattggcgatgatgaaacggattcgccggaagatgcgctttctga- gagctggcgcgaattgtggcaggatg cgttgcaggaggaggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtggtggcgct- gattgccgattttcgcaaagagtt ggataaacgcaccattcgcccgcgagggcggcaggtactcgatcacttaatgccgcatctgctcagcgatgta- tgctcgcgcgacgatgcgccagta ccgctgtcacgcctgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgctaagtgaat- ttcccggcgcactgaaacacctcattt ccctgtgtgccgcgtcgccgatggttgccagtcagctggcgcgctacccgatcctgcttgatgaattgctcga- cccgaatacgctctatcaaccgacg gcgatgaatgcctatcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaagagcaacagc- ttgaggcgtgggcagtttaag caggcgcagttgctgcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtgagcgatcact- taacctggctggcggaagcgatta ttgatgcggtggtgcagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatctgcacgatcg- cgaagggcgcggttttgcggtgg tcggttatggcaagctgggcggctgggagctgggttacagctccgatctggatctggtattcctgcacgactg- cccgatggatgtgatgaccgatggc gagcgtgaaatcgatggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttagcacgcgca- cgtcgtccggcatcctttatgaagttg atgcgcgtctgcgtccatctggcgctgcggggatgctggtcactactacggaatcgttcgccgattaccagca- aaacgaagcctggacgtgggaaca tcaggcgctggcccgtgcgcgcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccattcgccgc- gatattctgatgacgcctcgcgac ggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaagcataaag- accgcttcgatctgaaagccgat gaaggcggtatcaccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaagccgaaac- tgacgcgctggtcggataatgtgcg cattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctggcgtacacc- acattgcgtgatgagctgcacc acctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgcttattaaaac- cagctgggacaagtggctggtggaa ccgtgcgccccggcgtaagtgtggtatcatcgcgcgcaaattttgtatctctcaggagacaggaatgaaagtg- acgctgccagagtttaagcaagccg gtgtaatggtggtgggtgatgtgatgctggatcgttactggtatggcccaaccagccgtatctctccggaagc- gccagtcccggttgttaaagtcgatac cattgaagagcgtcctggcggcgcggcaaacgtggcgatgaatatcgcctcactgggcgccacggcgcgtctg- gttggcctgactggcattgacga tgcggcgcgcgcgctgagcaaagcgctggccgatgttaacgttaaatgtgacttcgtttctgttccgacgcat- cccaccatcactaagctgcgcgtgct gtcgcgtaaccagcagctgattcgcctggactttgaagagggttttgaaggagtcgatccgcaaccgatgcat- gaacgcatcagccaggcgcttggta atattggcgcgctggtgctgtcggatt 255 atgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattggactgattaaaaa- agcgcccttgtggcgattttttatattccc gcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccgatg- gaagctcgctgttttaacacgcgttttttaa ccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatattct- caacctaaaaaagtttgtgtaatacttgtaac gcttcatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaactca- cttcacaccccgaaggcgaagttgc ctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatga 256 accggatacgagagaaaagtgtctacatcggttcggttgatattgaccggcgcatccgccagcccgccca- gtttctggtggatctgtttggcgattttgc gggtcttgccggtgtcggtgccgaaaaaaataccaatatttgccataacacacgctcctgttgaaaaagagat- cccgccgggaaatgcggtgaacgtg tctgatattgcgaagagtgtgccagttttggtcgcgggcaaaacctgcaccagtttggttattaatgcaccag- tctggcgctttttttcgccgagtttctcct cgctaatgcccgccaggcgcggctttggcgctgatagcgcgctgaataccgatctggatcaaggttttgtcgg- gttatcagccaaaaggtgcactcttt gcatggttatacgtgcctgacatgttgtccgggcgacaaacggcctggtggcacaaattgtcagaactacgac- acgactaactgaccgcaggagtgt gcgatgaccctgaatatgatgatggatgccggccgtcctgtaataataaccggacaattcggactgattaaaa- aagcgcccttgtggcgctttttttatatt cccgcctccatttaaaataaaaaatccaatcggatttcactatttaaactggccattatctaagatgaatccg- atggaagctcgctgttttancacgcgttttt taaccttttattgaaagtcggtgcttctttgagcgaacgatcaaatttaagtggattcccatcaaaaaaatat- tcttcaacctaaaaaagtttgtgtaatacttgt aacgctacatggagattaactcaatctagagggtattaataatgaatcgtactaaactggtactgggcgcaac- tcacttcacaccccgaagggggaagt tgcctgaccctacgattcccgctatttcattcactgaccggaggttcaaaatgacccagcgaaccgagtcggg- taataccgtctggcgcttcgatttgtc ccagcagttcactgcgatgcagcgcataagcgtggtactcagccgggcgaccgaggtcgatcagacgctccag- caagtgctgtgcgtattgcacaat gacgcctttttgcagcacggcatgatctgtctgtacgacagccagcaggcgattttgaatattgaagcgttgc- aggaagccgatcagcagttaatcccc ggcagctcgcaaatccgctatcgtccgggcgaagggctggtcgggacggtgctttcgcagggccaatcattag- tgctggcgcgcgttgctgacgatc agcgctttcttgaccggctcgggttgtatgattacaacctgccgtttatcgccgtgccgctgatagggccaga- tgcgcagactttcggtgtgctgacggc acaacccatggcgcgttacgaagagcgattacccgcctgcacccgctttctggaaacggtc 257 tttcgctgaaggtgtgaccatgggccatcaggtgctggtgcagctggaaagtgttgccatcactatcgtg- tggtctggcgtggtggcctttattggttaca aactggcggacatgacggtaggcctgcgcgtaccggaagaacaagaacgtgaagggctggatgtaaacagcca- cggcgaaaacgcctataacgc ctga 258 tttcctttctgactctgcccgtccgggcgcactaacggcctgaaatactccctcttttcattcctggcac- aacgattgcaatgtctgttgcgtgttagctgcg gccattatcgaattcgactggagggggatctatgaagctggttaccgtggtgattaagccattcaaacttgaa- gacgtgcgtgaagcgctttcttctattg gtattcaagggttgaccgtaactgaagtgaaaggctttggccgtcagaagggtcacgctgagctgtaccgcgg- tgcggaatatagcgttaatttcctgc cgaaagtgaaaattgatgtggcgatcgctgacgatcaactcgatgaagtaatcgatgtgatcagcaaagcggc- ctacaccggaaaaattggcgacgg caaaattttcgttgctgagctgcaacgcgtcattcgtattcgtaccggcgaagccgacgaagggcactgtaat- acaagacacacagtgatggggatc ggtttcgctgaaggtgtgaccatgggccatcaggtgctggtgcagctggaaagtgttgccatcactatcgtgt- ggtctggcgtggtggcctttattggtta caaactggcggacatgacggtaggcctgcgcgtaccggaagaacaagaacgtgaagggctggatgtaaacagc- cacgacgaaacgcctataac gcctgattgcgttgagttatctcctgagcataaaaaagcctccattcggaggcttttctttttttaagtttaa- agcgcggttagttgcgattgcgcatgacgcc ttcctgcacgctggacgcgaccagcacaccctcttgcgtatagaactcgccgcgcacaaaaccgcgagcgctg- gaggctgacgtgctttccacactg tagagcagccattcgttcatattaaacgggcgatggaaccacatggagtggtcaatggtggcaacctgcatac- cgcgctcaaggaagcccacgccgt gcggctgaagtgcaaccggcaggaagttaaagtctgaggcatatccaagcagatattgatgtacgcgaaaatc- gtccggcaccgtgccgtttgcgcg gatccatacctggcgggtggcatcggcaacgtggcctttcagcgggttatgaaactcaaccgggcggatctcc- agtggtttatcactaagaaacttctc tttggcctgcgg 259 atgacctttaatatgatgcctggggtcactggagcgcttttatcggcatcctgaccgaagaatttgccgg- tttcttcccgacctggctggcccctgttcagg ttgtggtgatgaatatcactgattctcaagctgaatatgtcaacgaattgacccgtaaattgataaatgcggg- cattcgtgtaaaagcggacttgagaaac gagaagattggctttaaaatccgcgagcacactttacgtcgtgtcccttatatgttggtctgtggtgataaag- aggtggaagcaggcaaagtggccgttc gcacccgccgcggtaaagacctgggcagcctggacgtaaggaagtgattgagaagctgcaacaacagattcgc- agccgcagtcttcaacaactgg aggaataaggtattaaaggcggaatacgagttcaaacggcacgtccgaatcgtatcaatggcgagattcgcgc- ccaggaagttcgcttaactggtctg gaaggtgagcagctgggtattgcaatagaactaactacccgccctgaaggcggtacctgcctgaccctgcgat- tcccgttatttcattcactgaccgga ggcccacgatga 260 ggtacgacaaaaacgtctccagcgacgtgcggttaatattgactggcgcatccgccacatcccccagttt- ttgctggatcagtttggcgattttgcgggtt tttcccgtgtcactgccaaaaaaaataccaatgttagccatgtcgcgctcctgttgagaaagaataaggccgc- ctgcaaacggcggatatcccttctcct gttgcgaaggctgtgccaggtttttttaaggccttctgtgcactgaaatgcgtgaaaaaatgactcttttttg- tgcaggcaccgtcctctctccgctatccag acctgctttgaaggcctctgagggccaaatcagggccaaaacacgaatcacgatcaatgtttcggcgcgttac- ctgttcgaaaggtgcactctttgcat ggttaatcacacccaatcaggcctgcggatgtcgggcgtttcacaacacaaaatgttgtaaatgcgacacagc- cgggcctgaaaccaggagcgtgtg atgacctttaatatgatgcctggggtcactggagcgctttatcggcatcctgaccgaagaatttgccggtttc- ttcccgacctggctggcccctgttcagg ttgtggtgatgaatatcactgattctcaagctgaatatgtcaacgaattgacccgtaaattgcaaaatgcggg- cattcgtgtaaaagcggacttgagaaac gagaagattggctttaaaatccgcgagcacactttacgtcgtgtcccttatatgttggtctgtggtgataaag- aggtggaagcaggcaaagtggccgttc gcacccgccgcggtaaagacctgggcagcctggacgtaagtgaagtgattgagaagctgcaacaagagattcg- cagccgcagtcttcaacaactgg aggaataaggtattaaaggcggaaaacgagttcaaacggcacgtccgaatcgtatcaatggcgagattcgcgc- ccaggaagttcgcttaactggtctg gaaggtgagcagctgggtattgcaatagaactaactacccgccctgaaggcggtacctgcctgaccctgcgat- tcccgttatttcattcactgaccgga ggcccacgatgacccaccgacccgagtcgggcaccaccgtctggcgttttgatctctcacagcaatttaccgc- catgcagcgcatcagcgtggtgttg agtcgcgcaaccgagataagccagacgctgcaggaggtgctgtgtgttctgcataatgacgcatttatgcaac- acggcatgctgtgtctgtatgacaac cagcaggaaattctgagtattgaagccttgcaggaggcagaccaacatctgatccccggcagctcgcaaattc- gctatcgccctggcgaagggctgg taggagccgtactgtcccagggacaatctcttgtgctgccgcgtgtcgccgacgatcaacgctttctcgacag- gcttggcatctatgattacaacctgcc gtttatcgccgtccccttaatggggccaggcgcgcagacgattggcgtgctcgccgcgcagccgatggcgcgt- ctggaggagcggcttccttcctgt acgcgctttctggaaaccgtc 261 ttgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgaacggtagc- acagagagcttgctctcgggtgacgagt ggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaatacc- gcataacgtcgcaagaccaaaga ggcggaccttcgggcctcttgccatcagatgtgcccagatgggattagctagtaggtggggtaacggctcacc- taggcgacgatccctagctggtctg agaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatattgc- acaatgggcgcaagcctgatgc agccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaagggagtaaggtta- ataaccttattcattgacgttacccg cagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaattac- tgggcgtaaaccgcacgcaggc ggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcatccgaaactggcaggcttgagtct- cgtagagggaggtagaattccagg tgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggcctcctggacgaagactgacgc- tcaggtgcgaaagcgtgggga gcaaacaggattagataccctggtagtccacgccgtaaacgatgtctatttggaggttgtgcccttgaggcgt- ggcttccggagctaacgcgttaaata gaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtggagca- tgtggtttaattcgatgcaacgc gaagaaccttacctggtcttgacatccacagaactttccagagatggattggtgccttcgggaactgtgagac- aggtgctgcatggctgtcgtcagctc gtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcggtccggccggg- aactcaaaggagactgccagtgataa actggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccagggctacacacgtgctacaatggc- gcatacaaagagaagcgacctcg cgagagtaagcggacctcataaagtgcgtcgtagtccggattggagtctgcaactcgactccatgaagtcgga- atcgctagtaatcgtggatcagaat gccacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaagaagta- ggtagcttaaccttcgggagggc gcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttggatc- acctcctt 262 ttgagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgaacggtagca- cagagagcttgctctcgggtgacgagt ggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaatacc- gcataacgtcgcaagaccaaaga
gggggaccttcgggcctcttgccatcagatgtgcccagatgggattagctagtaggtggggtaacggctcacc- taggcgacgatccctagctggtctg agaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatattcc- acaatgggcgcaagcctgatgc agccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaaggnantanggtta- ataacctgtgttnattgacgttacc cgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaatt- actgggcgtaaagcgcacgcag gcggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcatccgaaactggcaggcttgagt- ctcgtagagggaggtagaattcca ggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggcctcctggacgaagactgac- gctcaggtgcgaaagcgtggg gagcaaacaggattagataccctggtagtccacgccgtaaacgatgtctatttggaggttgtgcccttgaggc- gtggcttccggagctaacgcgttaaa tagaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtggag- catgtggtttaattcgatgcaac gcgaagaaccttacctggtcttgacatccacagaacttagcagagatgctttggtgccttcgggaactgtgag- acaggtgctgcatggctgtcgtcagc tcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaaccatatcctttgttgccagcggttaggccgg- gaactcaaaggagactgccagtgat aaactggaggaaggtggggatgacgtcaagtcatcatggcccttacgaccagggctacacacgtgctacaatg- gcgcatacaaagagaagcgacct cgcgagagtaagcggacctcataaagtccgtcgtagtccggattggagtctgcaactcgactccatgaagtcg- gaatcgctagtaatcgtggatcaga atgccacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaagaag- taggtagcttaaccttcgggaggg cgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttggat- cacctcctt 263 atgaccatgcgtcaatgcgccatttacggcaaaggtgggatcggcaaatcgaccaccacacagaacctgg- tcgccgcgctggcggagatgggtaaa aaagtcatgattgtcggctgtgacccgaaagccgattccacgcgtttgatcctgcatgcgaaagcgcagaaca- ccattatggagatggctgagaagt cggctccgtggaagacctggagttagaagacgtgctgcaaatcggttacggcggcgtgcgctgcgcagagtcc- ggcggcccggagccaggcgtg ggctgtgccggtcgccgggtgatcaccgcgattaacttcctcgaagaagaaggcgcttacgtgccggatctcg- attttgttttctacgacgtgctgggc gacgtggtatgcggtggtttcgccatcccgattcgtgaaaacaaagcgcaggagatctacatcgtttgctctc- gcgaaatgatggcgatgtacgccgc caacaacatctccaaaggcatcgtgaaatacgccaaatccggtaaagtgcgcctcggcgggctgatttgtaac- tcgcgccagaccgaccgtgaagat gaactgatcattgcgctggcagaaaaactcggcacgcagatgatccactttgttccccgcgacaacattgtgc- agcgtgcggaaatccgccgtatgac ggttatcgaatatgacccgacctgcaatcaggcgaacgaatatcgcagccttgccagcaaaatcgtcaacaac- accaaaatggtggtgcccaccccc tgcaccatggatgaactggaagaactgctgatggagttcggcattatggatgtggaagacaccagcatcattg- gtaaaaccgccgccgaagaaaacg ccgtctga 264 atgagcaatgcaacaggcgaacgcaacctggagataatcgagcaggtgctcgaggttttcccggagaaga- cgcgcaaagaacgcagaaaacacat gatggtgacggacccggagcaggaaagcgtcggtaagtgcatcatctctaaccgcaaatcgcagccaggcgtg- atgaccgtgcgcggctgctcgta tgccggttcgaaaggggtggtatttgggccaatcaaggatatggcgcatatctcgcatggcccaatcggctgc- ggccaatactcccgcgccgggcgg cggaactactacaccggcgtcagcggcgtggacagcttcggcacgctcaacttcacctccgattttcaggagc- gcgacatcgtgtttggcggcgataa aaagctcgccaaactgattgaagagctggaagagctgttcccgctgaccaaaggcatttcgattcagtcggaa- tgcccggtcggcctgattggcgatg acattgaggccgtcgcgaacgccagccgcaaagccatcaacaaaccggttattccggtgcgttgcgaaggctt- tcgcggcgtgtcgcaatccctcgg tcaccatattgccaacgatgtgatccgcgactgggtgctggataaccgcgaaggcaaaccgttcgaatccacc- ccttacgatgtggcgatcatcggcg attacaacatcggcggcgatgcctgggcttcgcgcattttgctcgaagagatgggcttgcgggggtggcacag- tggtctggcgacggtacgctggt ggagatggaaaacacgccgttcgtcaaactgaacctggtgcattgttaccgctcaatgaactacatctcgcgc- catatggaggagaagcacggtattc cgtggatggaatacaacttctttggtccgacgaaaatcgcggaatcgctgcgcaaaatcgccgaccagtttga- cgacaccattcgcgccaacgccga agcggtgatcgccagataccagccgcaaaacgacgccattatcgccaaatatcgcccgcgtctggaggggcgc- aaagtgctgctttatatgggcgg gctgcgtccgcgccatgtgattgccgcctatgaagacctgggaatggagatcatcgctgccgctatgagttcg- gtcataacgatgattacgaccgca ccttgccggatctgaaagagggcacgctgctgtttgagatgccagcagttatgagctggaggcgttcgtcaac- gcgctgaaaccggatctcatcggtt ccggcatcaaagagaagtacatctttcagaaaatgggcgtgccgtttcgccagatgcactcctgggattactc- cggcccgtaccacggctatgacggc ttcgccatcttcgcccgcgatatggatatgacgctcaacaaccccgcgtggggccagttgaccgcgccgtggc- tgaaatccgcctga 265 atgagccagactgctgagaaaatacagaattgccatcccctgtttgaacaggatgcttaccagacgctgt- ttgccggtaaacgggcactcgaagaggc gcactcgccggagcgggtgcaggaagtgtttcaatggaccactaccccggaatatgaagcgctgaactttaaa- cgcgaagcgctgactatcgaccc ggcaaaagcctgccagccgctgggcgcggtgctctgttcgctggggtttgccaataccctaccgtatgtgcac- ggttcacagggttgcgtggcctattt ccgcacgtactttaaccgccactttaaagaaccggtggcctgcgtgtcggattcaatgacggaagacgcggcc- gtgttcggcgggaataacaacctc aacaccggcttacaaaacgccagcgcgctgtataaaccggagattatcgccgtctctaccacctgtatggcgg- aagtgatcggtgatgatttgcaggc ctttatcgccaacgccaaaaaagatggttttctcgatgccgccatccccgtgccctacgcgcacacccccagt- tttatcggcagccatatcaccggctg ggataacatgtttgaaggttttgcccggacctttacggcagaccatgaagctcagcccggcaaactttcacgc- atcaacctggtgaccgggtttgaaac ctatctcggcaatttccgcgtgctgaaacgcatgatggaacaaatggaggtgccggcgagtgtgctctccgat- ccgtcggaagtgctggatactcccg ccaacgggcattaccagatgtacgcgggcgggacgacgcagcaagagatgcgcgaggcgccggatgctatcga- caccctgttgctgcagccctg gcaactggtgaaaagcaaaaaagtggtgcaggagatgtggaatcagcccgccaccgaggtttctgttcccgtt- gggctggcaggaacagacgaact gttgatggcgattagccagttaaccggcaaggccattcccgattcactggcgctggagcgcgggcggctggtc- gatatgatgctcgattcccacacct ggttgcacggtaaaaaattcggcctgtttggcgatccggattttgtcatgggattgacccgtttcctgctgga- gctgggctgcgaaccgaccgttatcctc tgccacaacggtaacaaacgctggcagaaagcaatgaagaaaatgcttgacgcctcgccgtacggccaggaga- gcgaagtgtttatcaactgcgatt tgtggcatttccgctcgctgatgtttacccgccagccggattttatgattggcaactcgtacggcaagttcat- tcagcgcgacaccttagccaaaggcga gcagtttgaagttccgctgatccgcctcggttttcccctgttcgaccgccaccatctgcaccgccagaccacc- tggggctacgagggcgccatgagca ttctcactacccttgtgaatgcggtactggagagagtggacaaagagaccatcaagctcggcaaaaccgacta- cagcttcgatatcttatccgttaa 266 atgaccctgaatatgatgatggatgccggcgcgcccgaggcaatcgccggtgcgctttcgcgacaccatc- ctgggctgttttttaccatcgtgaagaa gcgcccgtcgccatttcgctgactgatgccgacgcacgcattgtctatgccaacccggctttctgccgccaga- ccggctatgaactagaagcgttgttg cagcaaaatccccgcctgcttgcaagtcgccaaaccccacgggaaatctatcaggatatgtggcacaccttgt- tacaacgccgaccgtggcgcgggc aattgattaaccgccaccgcgacggcagcctgtatctggtcgagatcgatatcaccccggtgattaacccgtt- tggcgaactggaacactacctggca atgcagcgcgatatcagcgccagttatgcgctggagcagcggttgcgcaatcacatgacgctgaccgaagcgg- tgctgaataacattccgccggcg gtggttgtagtggatgaacgcgatcatgtggttatggataaccttgcctacaaaacgttctgtgccgactgcg- gcggaaaagagctcctgagcgaactc aatttttcagcccgaaaagcggagctggcaaacggccaggtcttaccggtggtgctgcgcggtgaggtgcgct- ggttgtcggtgacctgctgggcgc tgccgcgcgtcagcgaagaagccagtcgctactttattgataacaggctgacgcgcacgctggtggtgatcac- cgacgacacccaacaacgccagc agcaggaacagggccgacttgaccgccttaaacagcagatgaccaacggcaaactactggcagcgatccgcga- aggcttgacgccgcgctgatc cagcttaactgccccatcaatatgctggcggcggcgcgacgtttaaacggcagtgataacaacaatgtggcgc- tcgacgccgcgtacgcgaaggt gaagaggcgatggcgcggctgaaacgttgccgcccgtcgctggaactggaaagtgcggccgtctggccgctgc- aacccttttttgacgatctgcgc gcgctttatcacacccgctacgagcaggggaaaaatttgcaggtcacgctggattcccatcatctggtgggat- ttggtcagcgtacgcaactgttagcct gcctgagtctgtggctcgatcgcacgctggatattgccgccgggctgggtgatttcaccgcgcaaacgcagat- ttacgcccgcgaagaagagggctg gctctctttgtatatcactgacaatgtgccgctgatcccgctgcgccacacccactcgccggatgcgcttaac- gctccgggaaaaggcatggagctgc gcctgatccagacgctggtggcacaccaccacggcgcaatagaactcacttcacaccccgaagggggaagttg- cctgaccctacgattcccgctatt tcattcactgaccggaggttcaaaatga 267 atgacccagcgaaccgagtcgggtaataccgtctggcgatcgatttggcccagcagttcactgcgatgca- gcgcataagcgtggtactcagccggg cgaccgaggtcgatcagacgctccagcaagtgctggcgtattgcacaatgacgcctttttgcagcacggcatg- atctgtctgtacgacagccagcag gcgattttgaatattgaagcgttgcaggaagccgatcagcagttaatccccggcagctcgcaaatccgctatc- gtccgggcgaagggctggtcggga cggtgctttcgcagggccaatcattagtgctggcgcgcgttgctgacgatcagcgctttcttgaccggctcgg- gttgtatgattacaacctgccgtttatc gccgtgccgctgatagggccagatgcgcagactttcggtgtgctgacggcacaacccatggcgcgttacgaag- agcgattacccgcctgcacccgc tttctggaaacggtcgctaacctggtcgcgcaaaccgtgcgtttgatggcaccaccggcagtgcgcccttccc- cgcgcgccgccataacacaggccg ccagcccgaaatcctgcacggcctcacgcgcatttggttttgaaaatatggtcggtaacagtccggcgatgcg- ccagaccatggagattatccgtcag gtttcgcgctgggacaccaccgttctggtacgcggcgagagtggcaccggcaaggagctgattgccaacgcca- tccaccaccattcgccgcgtgcc ggtgcgccatttgtgaaattcaactgtgcggcgctgccggacacactgctggaaagcgaattgttcggtcacg- agaaaggggcatttaccggcgcgg tacgccagcgtaaaggccgttttgagctggccgatggcggcacgctgtttcttgacgagatcggcgagtgtag- cgcctcgtttcaggctaagctgctg cgcattttgcaggaaggcgaaatggaacgcgtcggcggcgacgagacattgcaagtgaatgtgcgcattattg- ccgcgacgaaccgcaatcttgaa gatgaagtccggctggggcactttcgcgaagatctctattatcgcctgaatgtgatgcccatcgccctgccgc- cactacgcgaacgccaggaggacat tgccgagctggcgcactttctggtgcgtaaaatcgcccataaccagagccgtacgctgcgcattagcgagggc- gctatccgcctgctgatgagctaca actggcccggtaatgtgcgcgaactggaaaactgccttgagcgctcagcggtgatgtcggagaacggtctgat- cgatcgggatgtgattttgtttaatc atcgcgaccagccagccaaaccgccagttatcagcgtctcgcatgatgataactggctcgataacaaccttga- cgagcgccagcggctgattgcggc gctggaaaaagcgggatgggtacaagccaaagccgcgcgcttgctggggatgacgccgcgccaggtcgcctat- cgtattcagacgatggatataac cctgccaaggctataa 268 atgccgcaccacgcaggattgtcgcagcactggcaaacggtattttctcgtctgccggaatcgctcaccg- cgcagccattgagcgcgcaggcgcagt cagtgctcacttttagtgattttgttcaggacagcatcatcgcgcatcctgagtggctggcagagcttgaaag- cgcgccgccgcctgcgaacgaatggc aacactatgcgcaatggctgcaagcggcgctggatggcgtcaccgatgaagcctcgagatgcgcgcgctgcgg- ctgtttcgccgtcgcatcatggt gcgcatcgcctggagccaggcgttacagttggtggcggaagaagatatcctgcaacagcttagcgtgctggcg- gaaaccctgatcgtcgccgcgcg cgactggctttatgaggcctgctgccgtgagtggggaacgccgagcaatccacaaggcgtggcgcagccgatg- ctggtactcggcatgggcaaact gggtggcggcgaactcaatttctcatccgatatcgatttgattttcgcctgggacggaaaatggcgcaacgcg- cggtggacgccgtgagctggataacg cgcaatttttcactcgccttggtcaacggctgattaaagtcctcgaccagccaacgcaggatggctttgtcta- ccgcgtcgatatgcgcttgcgcccgttt ggcgacagcggcccgctggtgctgagctttgccgcgctggaagattactaccaggagcaggggcgcgattggg- aacgctacgcgatggtgaaagc gcgcattatgggcgataacgacggcgaccatgcgcgggagttgcgcgcaatgctgcgcccgtttgttttccgc- cgttatatcgacttcagcgtgattca gtccctgcgtaacatgaaaggcatgattgcccgcgaagtgcgtcgccgtggcctgaagacaacattaagctcg- gcgcgggcgggatccgcgaaat agaatttatcgtccaggttttccagctgattcgcggcggtcgcgagcctgcactgcaatcgcgttcactgttg- ccgacgcttgctgccatagatcaactg catctgctgccggatggcgacgcaacccggctgcgcgaggcgtatttgtggctgcgacggctggagaacctgc- tgcaaagcatcaatgacgaacag acacagacgctgccgggcgatgaactgaatcgcgcgcgcctcgcctggggaatgggcaaagatagctgggaag- cgctctgcgaaacgctggaag cgcatatgtcggcggtgcgtcagatatttaacgatctgattggcgatgatgaaacggattcgccggaagatgc- gctttctgagagctggcgcgaattgt ggcaggatgcgagcaggaggaggattccacgcccgtgctggcgcatctctcagaggacgatcgccgccgcgtg- gtggcgctgattgccgattttcg caaagagttggataaacgcaccattggcccgcgagggoggcaggtactcgatcacttaatgccgcatctgctc- agcgatgtatgctcgcgcgacgat gcgccagtaccgctgtcacgcctgacgccgctgctcaccggaattattacccgcaccacttaccttgagctgc- taagtgaatttcccggcgcactgaaa cacctcatttccctgtgtgccgcgtcgccgatggttgccagtcagaggcgcgctacccgatcctgcttgatga- attgctcgacccgaatacgctctatc aaccgacggcgatgaatgcctatcgcgatgagctgcgccaatacctgctgcgcgtgccggaagatgatgaaga- gcaacagcttgaggcgctgcgg cagtttaagcaggcgcagttgctgcgcgtggcggcggcggatattgccggtacgttgccagtaatgaaagtga- gcgatcacttaacctggctggcgg aagcgattattgatgcggtggtgcagcaagcctgggggcagatggtggcgcgttatggccagccaacgcatct- gcacgatcgcgaagggcgcggtt ttgcggtggtcggttatggcaagctgggcggctgggagctgggttacagctccgatctggatctggtattcct- gcacgactgcccgatggatgtgatga ccgatggcgagcgtgaaatcgatggtcgccagttctatttgcgtctcgcgcagcgcgtgatgcacctgtttag- cacgcgcacgtcgtccggcatccttt atgaagttgatgcgcgtctgcgtccatctggcgctgcggggatgctggtcactactacggaatcgttcgccga- ttaccagcaaaacgaagcctggacg tgggaacatcaggcgctggcccggcggcgtggtgtacggcgatccgcaactgaccgccgaatttgacgccatt- cgccgcgatattctgatgacgc ctcgcgacggcgcaacgctgcaaaccgacgtgcgagaaatgcgcgagaaaatgcgtgcccatcttggcaacaa- gcataaagaccgcttcgatctga aagccgatgaaggcggtatcaccgacatcgagtttatcgcccaatatctggtgctgcgctttgcccatgacaa- gccgaaactgacgcgctggtcggat aatgtgcgcattctcgaagggctggcgcaaaacggcatcatggaggagcaggaagcgcaggcattgacgctgg- cgtacaccacattgcgtgatga gctgcaccacctggcgctgcaagagttgccgggacatgtggcgctctcctgttttgtcgccgagcgtgcgctt- attaaaaccagctgggacaagtggc tggtggaaccgtgcgccccggcgtaa 269 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgaacggtag- cacagagagcttgactctcgggttgacga gtggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaata- ccgcataacgtcgcaagaccaaa gagggggaccttcgggcctcttgccatcagatgtgcccagatgggattagctagtaggtggggtaacggctca- cctaggcgacgatccctagctggt ctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatat- tgcacaatgggcgcaagcctga tgcagccatgccgcgtgtgtgaagaaggccttcgggttgtaaagcactttcagcggggaggaagggagtaagg- ttaataaccttgctcattgacgttat ccgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcggaat- tactgggcgtaaagcgcacgca
ggcggtctgtcaagtcggatgtgaaatccccgggctcaacctgggaactgcatccgaaactggcaggcttgag- tctcgtagagggaggtagaattcc aggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggcctcctggacgaagactga- cgctcaggtgcgaaagnnnnn nnnnnnaacaggattagataccctggtagtccatgccgtaaacgatgtctactagccgttggggcctttgagg- ctttagtggcgcagctaacgcgata agtagaccgcctggggagtacggtcgcaagactaaanctcaaatgaattgacgggggcccgcacaagcggtgg- agcatgtggtttaattcgatgca acgcgaagaaccttacctggccttgacatagtaagaattttccagagatggattggtgccttcgggaacttac- atacaggtgctgcatggctgtcgtcag ctcgtgtcgtgagatgttgggttaagtcccgcaacgagcgcaacccttgtcattagttgctacatttagttgg- gcactctaatgagactgccggtgacaaa ccggaggaaggtggggatgacgtcaagtcctcatggcccttataggtggggctacacacgtcatacaatggct- ggtacaaagggttgccaacccgc gagggggagctaatcccataaaaccagtcgtagtccggatcgcagtctgcaactcgactgcgtgaagtcggaa- tcgctagtaatcgtggatcagaat gtcacggtgaatacgttcccgggtcttgtacacaccgcccgtcacaccatgggagcgggttctgccagaagta- gttagcttaaccgcaaggagggcg attaccacggcagggttcgtgactggggtgaagtcgtaacaaggtagccgtatcggaaggtgcggctgcatca- cctccttt 270 atgaccatgcgtcaatgcgccatttacggcaaaggtgggatcggcaaatcgaccaccacacagaacctgg- tcgccgcgctggcggagatgggtaaa aaagtcatgattgtcggctgtgacccgaaagccgattccacgcgtttgatcctgcatgcgaaagcgcagaaca- ccattatggagatggctgctgaagt cggctccgtggaagacctggagttagaagacgtgctgcaaatcggttacggcggcgtgcgctgcgcagagtcc- ggcggcccggagccaggcgtg ggctgtgccggtcgcggggtgatcaccgcgattaacttcctcgaagaagaaggcgcttacgtgccggatctcg- attttgttttctacgacgtgctgggc gacgtggtatgcggtggtttcgccatgccgattcgtgaaaacaaagcgcaggagatctacatcgtttgctctg- gcgaaatgatggcgatgtacgccgc caacaacatctccaaaggcatcgtgaaatacgccaaatccggtaaagtgcgcctcggcgggctgatttgtaac- tcgcgccagaccgaccgtgaagat gaactgatcattgcgctggcagaaaaactcggcacgcagatgatccactttgttccccgcgacaacattgtgc- agcgtgcggaaatccgccgtatgac ggttatcgaatatgacccgacctgcaatcaggcgaacgaatatcgcagccttgccagcaaaatcgtcaacaac- accatnatggtggtgcccaccccc tgcaccatggatgaactggaagaactgctgatggagttcggcattatggatgtggaagacaccagcatcattg- gtaaaaccgccgccgaagaaaacg ccgtctga 271 atggccgaaattctgcgcagtaaaaaaccgctggcggtcagcccgataaaaagggccagccgctgggggc- gatcctcgcaagcctgggtgtcga acagtgcataccgctggtacacggcgcacagggatgtagcgcgttcgcgaaggtgttctttattctacatttt- cacgatccgatcccgctgcaatcgac ggcgatggatccgacttccaccattatgggcgccgatgaaaacatttttaccgcgctcaatgtgctctgccag- cgcaacgccgcgaaagccattgtgct gctcagcaccgggctttcagaagcccagggcagcgacatttcgcgggtggtgcgccagtttcgtgatgatttt- ccccggcataaaggcgttgcgctgc tcaccgtcaacacacccgatttctacggctcgctggaaaacggctacagcgccgtgctggaaagcatgattga- acagtgggtacccgcacagcccg ccgccagcctgcgcaaccgccgtgtcaacctgctggtcagccatttactgacaccaggcgatatcgaactgtt- gcgcagttatgttgaagccttcggcc tgcaaccggtgattgtgccggatctgtcgctgtcgctggacgggcatctggcagacggtgatttttcgcctgt- tacccaagggggaacatcgctgcgc atgattgaacagatggggcaaaacctggccacctttgtgattggcgcctcgctgggccgtgcggcggcgttac- tggcgcagcgcagccgtggcgag gtgatcgccctgccgcatctgatgacgcttgcagcctgcgacacgtttattcatcgactgaaaaccctctccg- ggcgcgatgtccccgcgtggattgag cgccagcgcggccaagttcaggatgcgatgatcgattgccatatgtggctgcagggtgcggctatcgccatgg- cagcagaaggcgatcacctggcg gcatggtgcgatttcgcccgcagccagggcatgatccccggcccgattgtcgcaccggtcagccagccggggt- tgcaaaatctgccggttgaaacc gtggttatcggcgatctggaagatatgcaggatcggctttgcgcgacgcccgccgcgttactggtggccaatt- ctcatgccgccgatctcgccacgca gtttgatttgtcacttatccgcgccgggttcccggtgtatgaccggctgggggaatttcgtcgcctgcgccag- gggtacagcggcattcgtgacacgct gtttgagctgggaatgtgatgcgcgagcgccatcacocgcttgcaacctaccgctcgccgctgcgccagcacg- ccgacgacaacgttacgcctgg agatctgtatgccgcatgttaa 272 atgaaaaacacaacattaaaaacagcgcttgcttcgctggcgttactgcctggcctggcgatggcggctc- ccgctgtggcggataaagccgacaacg gctttatgatgatttgcaccgcgctggtgctgtttatgaccattccgggcattgcgctgttctacggcggttt- gatccgcggtaaaaacgtgctgtcgatgc tgacgcaggttgccgtcaccttcgcactggtgtgcattctgtgggtggtgtatggctactcgctggcatttgg- cgagggcaacagcttcttcgggagtttt aactgggcgatgttgaaaaacatcgaactgaaagccgtgatgggcagcatttatcagtatatccacgtggcgt- tccagggttccttcgcctgtatcaccg ttggcctgattgtcggtgcactggctgagcgtattcgcttactgcggtgctgatttttgtggtggtatggctg- acgctttcttacgtgccgattgcacacat ggtggggggcggcggtctgctggcaacccacggtgcgctggatttcgcaggcggtacggttgttcacatcaac- gctgcgattgcaggtctggtgggg gcttacctgattggcaaacgcgtgggctttggcaaagaagcattcaaaccgcataacctgccgatggtcttca- ctggcaccgctatcctgatgttggct ggtttggtttcaacgccggctccgcaagctcggcgaacgaaattgctgcgctggccttcgtgaacactgtcgt- tgccactgctgccgctattctggcgtg ggtatttggcgaatgggcaatgcgcggcaagccgtctctgctcggtgcctgttctggtgccatcgcgggtctg- gttggtatcacccccgcctgtggttat gtgggtgtcggcggtgcgctgattgtgggtctgattgccggtctggctgggctgtggggcgttactgcgctga- aacgtatgttgcgtgtcgatgacccg tgtgacgtattcggtgtgcacggcgtgtgcggcatcgtgggctgtatcctgacgggtatcttcgcctctacgt- cgctgggtggtgtcggtttcgctgaag gtgtgaccatgggccatcaggtgctggtgcagctggaaagtgttgccatcactatcgtgtggtctggcgtggt- ggcctttattggttacaaactggcgga catgacggtaggcctgcgcgtaccggaagaacaagaacgtgaagggctggatgtaaacagccacggcgaaaac- gcctataacgcctga 273 cgtcctgtaataataaccggacaattcggactgattaaaaaagcgcccttgtggcgctttttttatattc- ccgcctccatttaaaataaaaaatccaatcgga tttcactatttaaactggccattatctaagatgaatccgatggaagacgctgttttaacacgcgttttttaac- cttttattgaaagtcggtgcttctttgagcga acgatcaaatttaaatggattcccatcaaaaaaatattctcaacctaaaaaagtttgtgtaatacttgtaacg- ctacatggagattaactcaatctagagggt attaataatgaatcgtactaaactggtactgggcgc 274 ggacatcatcgcgacaaacaatattaataccggcaaccacaccggcaatttacgagactgcgcaggcatc- ctttctcccgtcaatttctgtcaaataaag taaaagaggcagtctacttgaattacccccggctggttgagcgtttgttgaaaaaaagtaactgaaaaatccg- tagaatagcgccactctgatggttaatt aacctattcaattaagaattatctggatgaatgtgccattaaatgcgcagcataatggtgcgttgtgcgggaa- tactgcttttttttgaaagggttagtcagt agcggaaac 275 atgaccctgaatatgatgctcgataacgccgcgccggacgccatcgccggcgcgctgactcaacaacatc- cggggagttttttaccatggtggaaca ggcctcggtggccatctccctcaccgatgccagcgccaggatcatttacgccaacccggcgttttgccgccag- accggctattcgctggcgcaattgtt aaaccagaacccgcgcctgctggccagcagccagacgccgcgcgagatctatcaggagatgtggcataccctg- accagcgtcagccctggcgcg gtcagctgattaatcagcgtcgggacggcggcctgtacctggtggagattgacatcaccccggtgcttagccc- gcaaggggaactggagcattatct ggcgatgcagcgggatatcagcgtcagctacaccctcgaacagcggctgcgcaaccatatgaccctgatggag- gcggtgctgaataatatccccgc cgccgtggtagtggtggacgagcaggatcgggtggtgatggacaacctcgcctacaaaaccttctgcgctgac- tgcggcggccgggagctgctcac cgagctgcaggtctcccctggccggatgacgcccggcgtggaggcgatcctgccggtggcgctgcgcggggcc- gcgcgctggctgtcggtaacc tgctggccgttgcccggcgtcagtgaagaggccagccgctactttatcgacagcgcgctggcgcggaccctgg- tggtgatcgccgactgtacccag cagcgtcagcagcaggagcaagggcgccttgaccggctgaagcagcaaatgaccgccggcaagctgctggcgg- cgatccgcgagtcgctggac gccgcgctgatccagctgaactgcccgattaatatgctggcggcagcccgtcggctgaacggcgagggaagcg- ggaatgtggcgctggaggccg cctggcgtgaaggggaagaggcgatggcgcggctccagcgctgtcgcccatcgctggaactcgaaaaccccgc- cgtctggccgctgcagccctttt tcgacgatctgtgcgccctctaccgtacacgcttcgatcccgacgggctgcaggtcgacatggcctcaccgca- tctgatcggctttggccagcgcacc ccactgctggcgtgcttaagcctgtggctcgatcgcaccctggccctcgccgccgaactcccctccgtgccgc- tggcgatgcagctctacgccgagg agaacgacggctggctgtcgctgtatctgactgacaacgtaccgctgctgcagggcgctacgctcactccccc- gacgcgctgaactcgccgggca aaggcatggagctgaggctgatccagaccctggtggcgcaccatcgcggggccattgagctggcttcccgacc- gcagggcggcacctgcctgacc ctgcgtttcccgctgtttaacaccctgaccggaggtgaagcatga 276 atgatccctgaatccgacccggacaccaccgtcagacgcttcgacctctctcagcagttcaccgccatgc- agcggataagcgtggtgctgagccggg ccaccgaggccagcaaaacgctgcaggaggtgctcagcgtattacacaacgatgcctttatgcagcacgggat- gatctgcctgtacgacagcgagc aggagatcctcagtatcgaagcgctgcagcaaaccggccagcagcccctccccggcagcacgcagatccgcta- tcgccccggcgagcgactggt ggggaccgtgctggcccaggggcagtcgctggtgctgccccgggtcgccgacgatcagcgttttctcgaccgc- ctgagcctctacgattacgatctg ccgtttatcgccgaccgttgatgcggcccaacgcccggccaataggggtgctggcggcccagccgatggcgcg- ccaggaagagcggctgccgg cctgcacccgttttctcgaaaccgtcgccaacctcgtcgcccagaccatccggctgatgatccttccggcctc- acccgccctgtcgagccgccagccg ccgaaggtggaacggccgccggcctgctcgtcgtcgcgcggcgtgggccttgacaatatggtcggcaagagcc- cggcgatgcgccagatcgtgg aggtgatccgtcaggtttcgcgctgggacaccaccgtgctggtacgcggcgaaagcggcaccgggaaagagct- gatcgccaacgccatccatcac cattcgccacgggctggcgccgccttcgtcaaatttaactgcgcggcgctgccggacaccctgctggaaagcg- aactgttcggccatgagaaaggc gcctttaccggggcggtgcgtcagcgtaaaggacgttttgagctggcggatggcggcaccctgttcctcgatg- agattggtgaaagcagcgcctcgtt ccaggccaagagctgcgtatcctccaggagggggagatggagcgggtcggcggcgatgagaccctgcgggtga- atgtccgcatcatcgccgcc accaaccgtcacctggaggaggaggtccggctgggccatttccgcgaggatctctactatcgtctgaacgtga- tgcccatcgccctgcccccgctgc gcgagcgtcaggaggacatcgccgagctggcgcacttcctggtgcgcaaaatcggccagcatcaggggcgcac- gctgcggatcagcgagggcg cgatccgcctgctgatggagtacagctggccgggtaacgttcgcgaactggagaactgcctcgaacgatcggc- ggtgatgtcggagagtggcctga tcgatcgcgacgtgatcctcttcactcaccaggatcgtcccgcaaagccctgcctgccagcgggccagcggaa- gacagctggctggacaacagcc tggacgaacgtcagcgactgatcgccgcgctggaaaaagccggctgggtgcaggccaaggcggcacggctgct- ggggatgacgccgcgccagg tcgcttatcggatccagatcatggatatcaccctgccgcgtctgtag 277 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcggcag- cgggaagtagcttgctactttgccggc gagcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaa- taccgcatgacctcgaaagagc aaagtgggggatcttcggacctcacgccatcggatgtgcccagatgggattagctagtaggtgaggtaatggc- tcacctaggcgacgatccctagct ggtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaa- tattgcacaatgggcgcaagc ctgatgcagccatgccgcggtgtgaagaaggccttagggtttgtaaagcactttcagcgaggaggaaggcatc- atacttaatacgtgtggtgattgtcg ttactcgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcg- gaattactgggcgtaaagcgcac gcaggcggtttgttaagtcagatgtgaaatccccgagcttaacttgggaactgcatttgaaactggcaagcta- gagtcttgtagaggggggtagaattc caggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactg- acgctcaggtgcgaaagcgtg gggagcaaacaggattagataccctggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttgag- gcgtggcttcggagctaacgcgtt aagtcgaccgcctggggagtacggccgcaagtttaaaactcaaatgaattgacgggggcccgcacaaagcggt- ggagcatgtggtttaattcgatgc aacgcgaagaaccttacctactcttgacatccagagaatttgccagagatggcgaagtgccttcgggaactct- gagacaggtgctgcatggctgtcgt cagctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcacgtaa- tggtgggaactcaaaggagactgccg gtgataaaccggaggaaggtggggatgacgtcaagtcatcattgcccttacgagtagggctacacacgtgcta- caatggcatatacaaagagaagc gaactcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtctgcaactcgactccatga- agtcggaatcgctagtaatcgtaga tcagaatgctacggtgaatacgttccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaa- gaagtaggtagcttaaccttcggg agggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggtt- ggatcacctcctt 278 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcggcat- cgggaagtagcttgctactttgccggcg agcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaat- accgcatgacctcgaaagagca aagtgggggatcttcggacctcacgccatcggatgtgcccagatgggattagctagtaggtgaggtaatggct- nacctaggcgacgatccctagctg gtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaat- attgcacaatcggcgcaagcct gatgcagccatgccgcgtgtgtgaagaaggccttaggggttaaagcactttcagcgaggaggaaggcatcata- cttaatacgtgtggtgattgacgtt actcgcagaagaagcaccggctaactccgtgccagcagacgggtaatacggagggtgcaagcgttaatcggaa- ttactgggcgtaaagcgcacg caggcggtttgttaagtcagatgtgaaatccccgagcttaacttgggaactgcatttgaaactggcaagctag- agtcttgtagaggggggtagaattcca ggtgtagggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagtctgacg- ctcaggtgcgaaagcgtggg gagcaaacaggattagataccctggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttgaggc- gtggcttccggagctaacgcgttaa gtcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtgga- gcatgtggtttaattcgatgcaa cgcgaagaaccttacctactcttgacatccagagaatttgccagagatggcgaagtgccttcgggaactctga- gacaggtgctgcatggctgtcgtca gctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcacgtgatg- gtgggaactcaaaggagactgccggt gataaaccggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacgtgctaca- atggcatatacaaagagaagcga actcgcgagagcaagggacctcataaagtatgtcgtagtccggattggagtctgcaactcgactccatgaagt- cggaatcgctagtaatcgtagatca gaatgctacggtgaatacgttcccgggccttgtacacaccgcccgtcacacattgggagtgggttgcaaaaga- agtaggtagcttaaccttcgggag ggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttgg- atcacctcctt 279 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcggcag- gggaagtagcttgctactttgccggc gagcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaa- taccgcatgacctcgaaagagc aaagttgggggatcttcggacctcacgccatcggatgtgcccagatgggattagctagtaggtgaggttaatg-
gctcacctaggcgacgatccctagct ggtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaa- tattgcacaatgggcgcaagc ctgatgcagccatgccgcgtgtgtgaagaaggccttagggttgtaaagcactttcagcgaggaggaaggcanc- atacttaatacgtgtggtgattgac gttactcgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatc- ggaattactgggcgtaaagcgca cgcaggcggtttgttaagtcagatgtgaaatccccgagcttaacttgggaactgcatttgaaactggcaagct- agagtcttgtagaggggggtagaatt ccaggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagact- gacgctcaggtgcgaaaggcgt ggggagcaaacaggattagataccctggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttga- ggcgtggcttccggagctaacgcgt taagtcgaccgcctggggagtacggccgcaaggttaaactcaaatgaattgacgggggcccgcacaagcggtg- gagcatgtggtttaattcgatgc aacgcgaagaaccttacctactcttgacatccagagaatttgccagagatggcgaagtgccttcgggaactct- gagacaggtgctgcatggagtcgt cagctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcacgtga- tggtgggaactcaaaggagactgccg gtgataaaccggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacgtgcta- caatggcatatacaaagagaagc gaactcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtagcaactcgactccatgaa- gtcggaatcgctagtaatcgtaga tcagaatgctacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaa- agaagtaggtagcttaaccttcggg agggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggtt- ggatcacctcctt 280 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcgacan- cgggaagtagcttgctactttgccggc gagcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaa- taccgcatgacctcgaaagagc aaagtgggggatcttcggacctcacgccatcggatgtgcctagatgggattagctagtaggtgaggtaatggc- ttacctaggcgacgatccctagctg gctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagggggaatat- tgcacaatgggcgcaagcct gatgcagccatgccgcgtgtgtgaagaaggccttagggttgtaaagcactttcagcgaggaggaaggcatcac- acttaatacgtgggtgattgacgtt actcgcagaagaagcaccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcgttaatcgga- attactgggcgtaaagcgcacg caggcggtttgttaagtcagatgtgaaatccccgagcttaacttgggaactgcatttgaaactggcaagctag- agtcttgtagagggggtgagaattcca ggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactgac- gctcaggtgcgaaagcgtggg gagcaaacaggattagataccctggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttgaggc- gtggcttccggagctaacgcgttaa gtcgaccgactggggagtacggccgaaaggtttaaaactcaaatgaattgacgggggcccgcacaagcggtgg- agcatgtggtttaattcgatgcaa cgcgaagaaccttacctactcttgacatccagagaatttgccagagatggcgaaggccttcgggaactctgag- acaggtgctgcatggctgtcgtca gctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgcaacccttatcctttgttgccagcacgtgatg- gtgggaactcaaaggagactgccggt gataaaccggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacggctacaa- tggcatatacaaagagaagcga actcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtctgcaactcgactccatgaag- tcggaatcgctagtaatcgtagatca gaatgctactgtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaaga- agtaggtagcttaaccttcgggag ggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttgg- atcacctcctt 281 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgcaagtcgagcggcag- cgggaagaagcttgctactttgccggc gagcggcggacgggtgagtaatgtctgggaaactgcctgatggagggggataactactggaaacggtagctaa- taccgcatgacctcgaaagagc aaagtgggggatcttcggacctcacgccatcggatgtgcccagatgggattagctagaggtgaggtaatggct- cacctaggcgacgatccctagct ggtctgagaggatgaccagccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaa- tattgcacaatgggcgcaagc ctgatgcagccatgccgcgtgtgtgaagaaggccttagggttgtaaagcactttcagcgaggaggaaggcatc- acacttaatacgtgtgttgattgacg ttactcgcagaagaagcaccggctaactccgtgccagcagccgcgattaatacggagggtgcaagcgttaatc- ggaattactgggcgtaaagcgcac gcaggcggtttgttaagtcagatgtgaaatccccgagcttaacttgggaactgcatttgaaactggcaagcta- gagtcttgtagaggggggtagaattc caggtgtagcggtgaaatgcgtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactg- acgctcaggtgctgaaagcgtg gggagcaaacaggattagataccaggtagtccacgctgtaaacgatgtcgacttggaggttgtgcccttgagg- cgtggcttccggagctaacgcgtt aagtcgaccgcctggggagtacggccgcaaggttaaaactcaaatgaattgacgggggcccgcacaagcggtg- gagcatgtggtttaattcgatgc aacgcgaagaaccttacctactcttgacatccagagaatttgccagagatggcgaagtgccttcgggaactct- gagacaggtgctgcatggctgtcgt cagctcgtgttgtgaaatgttgggttaagtacgcaacgagcgcaacccttatcctttgttgccagcacgtgat- ggtgggaactcaaaggagactgccg gtgataaaccggaggaaggtggggatgacgtcaagtcatcatggcccttacgagtagggctacacacgtgcta- caatggcatatacaaagagaagc gaactcgcgagagcaagcggacctcataaagtatgtcgtagtccggattggagtctgcaactcgactccatga- agtcggaatcgctagtaatcgtaga tcagaatgctacggtgaatacgttcccgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaa- agaagtaggtagcttaaccttcggg agggcgcttaccactttgtgattcatgactggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggtt- ggatcacctcctt 282 gtagctaataccgcatgacctcgaaagagcaaagtgggggatcttcggacctcacgccatcggatgtgcc- cagatgggattagctagtaggtgaggt aatggctcacctaggcgacgatccctagctggtctgagaggatgaccagccacactggaactgagacacggtc- cagactcctacgggaggcagca gtggggaatattgcacaatgggcgcaagcctgatgcagccatgccgcgtgtgtgaagaaggccttagggttgt- aaagcactttcagcgaggaggaa ggcatcanacttaatacgtgtgntgattgacgttactcgcagaagaagcaccggctaactccgtgccagcagc- cgcggtaatacggagggtgcaagc gttaatcggaattactgcgcgtaaagcgcacgcaggcggtttgttaagtcagatgtgaaatccccgagcttaa- cttgggaactgcatttgaaactggca agctagagtcttgtagaggggggtagaattccaggtgtagcggtgaaatgcgtagagatctggaggaataccg- gtggcgaaggcggccccctggac aaagactgacgctcaggtgcgaaagcgtgggagcaaacaggattagataccctggtagtccacgctaaacgat- gtcgacttggaggttgtgccc ttgaggcgtggcttccggagctaacgcgttaagtcgaccgcctggggagtacggccgcaaggttaaaactcaa- atgaattgacgggggcccgcaca agcggtggagcatgtggtttaattcgatgcaacgcgaagaaccttacctactcttgacatccagagaatttgc- cagagatggcgaagtgccttcgggaa ctctgagacaggtgctgcatggctgtcgtcagctcgtgttgtgaaatgttgggttaagtcccgcaacgagcgc- aacccttatcctttgttgccagcacgt natggtgggaactcaaaggagactgccggtgataaac 283 attgaagagtttgatcatggctcagattgaacgctggcggcaggcctaacacatgaagtcgagcggcagc- gggaagtagcttgctactttgccggc gagcggcggacgggtgagtaatgtcctgatggaggggataactactggaacggtagctaataccgcacctcga- aagagcaaagtgggggatcttcg gacctcacgccatcggatgtgcccagatgggattagctagtaggtgaggtaatggctcacctaggcgacgatc- cctagctggtctgagaggatgacc agccacactggaactgagacacggtccagactcctacgggaggcagcagtggggaatattgcacaatgggcgc- aagcctgatgcagccatgccgc gtgtgtgaagaaggccttagggttctaaagcactttcagcgagcaggaaggcatcatacttaatacgtgtggt- gattgacgttactcgcagaagaagca ccggctaactccgtgccagcagccgcggtaatacggagggtgcaagcttsatcggaattactgggcgtaaagc- gcacgcaggcggttgttaagtca gatgtgaaatccccgagcttaacttgggaactgcatttgaaactggcaagctagagtcttgtttgaggggggt- agaattccaggtgtagcggtgaaatgc gtagagatctggaggaataccggtggcgaaggcggccccctggacaaagactgacgctcaggtgcgaaagcgg- gggagcaaacaggattaata ccctggtagtccacgctgtaacgatgtcgacttggaggttgtgccctgaggcgtggcttccggagctaacgcg- ttaagtcgaccgcctggggagtacg gccgcaaggtaaaactaaatgaattgacgggggcccgcacaagcggtggagcatgtggtttaattcgatgcaa- cgcgaagaaccttacctactctt gacatccagagaatttgccagagatggcgaagtgccttcgggaactctgagacaggtgctgcatggctgtcgt- cagctcgtgttgtgaaatgttgggtt aagtcccgcaacgagcgcaacccttatcctttgttgccagcacgtaatggtgggaactcaaaggagactgccg- gtgataaaccggaggaaggtggg gatgacgtcaagtcatcatggcccttacgagtagggctacacacgtgctacaatggcatatacaaagagaagc- gaactcgcgagagcaagcggacc tcataaagtatgtcgtagtccggattggagtctgcgactcgactccatgaagtcggaatcgctagaatcgtag- atcagaatgctacggtgaatacgttcc cgggccttgtacacaccgcccgtcacaccatgggagtgggttgcaaaagaagtaggtagcttaacatcgggag- ggcgcttaccactttgtgattcatg actggggtgaagtcgtaacaaggtaaccgtaggggaacctgcggttggatcacctcctt 284 atgaccatgcgtcaatgcgctatctacggtaaaggcggtatcggtaaatccaccaccacccagaatctcg- tcgcggccctcgccgagatgggtaaga aagtgatgatcgtcggctgcgatccgaaagcggattccacccgtctgatcctccacgctaaagcccagaacac- catcatggagatggcggcggaagt gggctcggtcgaggatctggagctcgaagacgttctgcaaatcggctatggcgatgtccgttgcgccgaatcc- ggcggcccggagccaggcgtcg gctgcgccggacgcggggtgatcaccgccatcaacttcctcgaggaaggcgcctatgaagaagatttggattt- cgtcttctatgacgtcctcggc gacgtggtctgcggcggcttcgctatgccgatccgcgaaaacaaagcccaggagatctacatcgtctgctccg- gcgagatgatggcgatgtatgccg ccaacaatatctccaaagggatcgtgaagtacgccaaatccggcaaggtgcgcctcggcggcctgatctgtaa- ctcgcgcaaaaccgaccgggaag acgaactgatcatcgccctggcggagaagcttggcacgcagatgatccacttcgttccccgcgacaacattgt- gcagcgcgcggagatccgccgga tgacggtgatcgagtacgacccgacctgtcagcaggcgaggaatatcgtcaactggcgcagaagatcgtcaat- aacaccaaaaaagtggtgccga cgccgtgcaccatggacgagctggaatcgctgctgatggagttcggcatcatggaagaagaagacaccagcat- cattggtaaaaccgccgctgaag aaaacgcggcctga 285 atggttaggaaaagtagaagtaaaaatacaaatatagaactaactgaacatgaccatttattaataagtc- aaataaaaaagcttaaaacacaaaccacttg cttttttaataataaaggaggggttgggaagactacattagtagcaaatttaggagcagagctatcaataaac- tttagtgcaaaagttcttattgtggatgcc gaccctcaatgtaatctcacgcagtatgtattaagtgatgaagaaactcaggacttatatgggcaagaaaatc- cagatagtatttatacagtaataagacc actatcctttggtaaaggatatgaaagtgacctccctataaggcatgtagagaatttcggttttgacataatt- gtcggtgaccctagacttgctttacaggaa gaccttttagctggagactggcgagatgccaaaggcggtgggatgcgaggaattaggacaacttttgtatttg- cagagttaattaagaaagctcgtgag ctaaattatgattttgttttctttgacatgggaccatcattaggcgcaatcaacagggcagtattactggcaa- tggaattctttgtcgtcccaatttcaatcga tgtattttcactatgggctattaaaaatattggctccacggtttcaatatggaaaaaagaattagacacaggg- attcggctctcagaggaacctagcgaatt atcacaattatcttcctcaaggaaaactaaagtttctcggttacgtcacccaacaacataaagaacgctctgg- atacgatacaattcagcttgagaatactg aggaagaaataaaatcgaaacgtcgggtaaaggcgtatgaagacattggagaggtgtttccttctaaaattac- tgagcatctttctaaactttatgatca aaagatatgaacccacaccttggagatatacgtcatttaggtagtttagctccgaaatcacaatcacaacacg- ttccgatgatatcagtgtctggtacagg aaattacaccagacttagaaaaagcgcgcgtgaactttatcgagatattgcaagaagatacttagagaacatt- cagactgctaatggcgagaaatag 286 atgaagggaaaggaaattctggcgctgctggacgaacccgcctgcgagcacaaccagaagcaaaaatccg- gctgcagcgcccctaagcccggcg ctaccgccggcggttgcgccttcgacggcgcgcagataacgctcctgcccatcgccgacgtcgcgcacctggt- gcacggccccatcggctgcgcg ggcagctcgtgggataaccgcggcagcgtcagcgccggcccggccctcaaccggctcggctttaccaccgatc- ttaacgaacaggatgtgattatg ggccgcggcgaacgccgcctgttccacgccgtgcgtcacatcgtcgaccgctatcatccggcggcggtcttta- tctacaacacctgcgtaccggcga tggagggcgatgacatcgaggcggtctgccaggccgcacagaccgccaccggcgtcccggtcatcgctattga- cgccgccggtttctacggcagta aaaatcttggcaaccgaatggcgggcgacgtgatgctcaggcaggtgattggccagcgcgaaccggccccgtg- gccagacaacacgccctttgcc ccggcccagcgccacgatatcggcctgattggcgaattcaatatcgccggcgagttctggcaggtccagccgc- tgacgacgagctggagatccgc gtcctcggcaccctctccggcgacggccgctttgccgagatccagaccctgcaccgggcgcaggccaatatgc- tggtgtgctcgcgcgcgctgatc aacgtcgcccgggggctggagctgcgctacggcacgccgtggtttgaaggcagcttctacgggatccgcgcca- cctccgacgccttgcgccagct ggcgacgctgctgggggatgacgacctgcgccgccgcaccgaggcgctgatcgcccgcgaagagcaggcggcg- gagcaggctcttgcgccgt ggcgtgagcagctccgcgggcgcaaatgctgctctataccggcggcgtgaaatcctggtcggtggtatcggcc- ctgcaggatctcggcatgaccg tggtggccaccggcacgcgcaaatccaccgaggaggacaaacagcggatccgtgagctgatgggcgacgaggc- ggtgatgcttgaggagggca atgcccgcaccctgctcgacgtggtgtaccgctatcaggccgacctgatgatcgccggcggacgcaatatgta- caccgcctggaaagcccggctgc cgtttctcgatatcaatcaggagcgcgagcacgcctacgccggctatcagggcatcatcaccctcgcccgcca- gctctgtctgaccctcgccagcccc gtctggccgcaaacgcatacccgcgccccgtggcgctag 287 atgaccaacgcaacaggcgaacgtaaccttgcgctcatccaggaagtcctggaggtgtttcccgaaaccg- cgcgcaaagagcgcagaaagcacat gatgatcagcgatccgagatggagagcgtcggcaagtgcattatctcgaaccgtaaatcgcagcccggggtga- tgaccgtgcgcggctgcgccta tgcgggctcgaaaggggtggtgtttgggccaatcaaagacatggcccatatctcgcacggccccatcggctgc- ggccagtattcccgcgccggacg gcgcaactactataccggcgtcagcggtgtcgacagcttcggcaccctgaacttcacctctgattttcaggag- cgcgatattgttttcggcggcgataaa aagctgaccaaactgatcgaagagatggagagctgttcccgctgaccaaagggatcaccatccagtcggagtg- cccggtgggcctgatcggcgat gacatcagcgccgtagccaacgccagcagcaaggcgctggataaaccggtgatcccggtgcgctgcgaaggct- ttcgcggcgtatcgcaatcgct gggccaccatatcgccaacgacgtggtgcgcgactgggtgctgaacaatcgcgaagggcagccgtttgccagc- accccgtacgatgttgccatcatt ggcgattacaacatcggcggcgacgcctgggcctcgcgcattctgctggaagagatggggctgcgcgtagtgg- cgcagtggtccggcgacggca ccctggtggagatggagagcaccccattcgttaagcttaacctcgtccactgctaccgttcgatgaactatat- cgcccgccatatggaggagaaacatc agatcccatggatggaatataacttcttcggcccgaccaaaatcgccgaatcgctgcgcaagatcgccgatca- atttgatgacaccattcgcgccaatg cggaagcggtgatcgccaaatatgaggggcagatggcggccatcatcgccaaatatcgcccgcggctggaggg- gcgcaaagtgctgctgtacatg ggggggctgaggccgcgccacgtcatcggcgcctatgaggatctcgggatggagatcatcgccgccggctacg- agtttgcccataacgatgattac gaccgcaccctgacggacctgaaagagggcaccctgctgtttgacgatgccagcagatatgagctggaggcct- tcgtcaaagcgctgaaacctgac ctcatcggctccgggatcaaagagaaatatatcttccagaaaatgggggtgcgttccgccagatgcactcctg- ggactattccggcccctatcacgg ctatgacggcttcgccatctttgcccgcgatatggatatgaccctgaacaatccggcgtggaacgaactgact- gccccgtggctgaagtctgcgtga
288 atggcagatattatccgcagtgaaaaaccgctggcggtgagcccgattaaaaccgggcaaccgctcgggg- cgatcctcgccagcctcgggctggc ccaggccatcccgctggccacggcgcccagggctgcagcgccttcgccaaagttttctttattcagcatttcc- atgacccggtgccgctgcagtcgac ggccatggatccgaccgccacgatcatgggggccgacggcaatatcttcaccgcgctcgacaccctctgccag- cgccacagcccgcaggccatcg tgctgctcagcaccggtctggcggaagcgcagggcagcgatatcgcccgggtggtgcgccagtttcgcgaggc- gcatccgcgccataacggcgtg gcgatcctcaccgtcaataccccggatttttttggctctatggaaaacggctacagcgcggtgatcgagagcg- tgatcgagaagtgggtcgcgccgac gccgcgtccgggcagcggccccggcgggtcaacctgctggtcagccacctctgttcgccaggggatatcgaat- ggctgggccgctgcgtggag gcctttggcctgcagccggtgatcctgccggacctctcgcagtcaatggatggccacctcggtgaaggggatt- ttacgcccctgacccagggcggcg cctcgctgcgccagattgcccagatgggccagagtagggcagcttcgccattggcgtgtcgctccagagggcg- gcatcgatcctgacccaacgca gccgcggcgacgtgatcgccctgccgatctgatgaccctcgaccattgcgatacctttatccatcagctggcg- aagatgtccggacgccgcgtacc ggcctggattgagcgccagcgtggccagctgcaggatgcgatgatcgactgccatatgtggcttcagggccag- cgcatggcgatggcggcggagg gcgacctgctggcggcgtggtgtgatttcgcccgcagccaggggatgcagcccggcccgctggtcgcccccac- cagccaccccagcctgcgcca gctgccggcgagcaagtcgtgccgggggatcttgaggatctgcagcagctgctgagccaccaacccgccgatc- tgaggtggctaactctcacgc ccgcgatctggcggagcagtttgccctgccgctgatccgcgtcggttttcccctcttcgaccggctcggtgag- tttcgtcgagtccgccaggggtacgc cggtatgcgagatacgctgtttgaactggccaatctgctgcgcgaccgccatcaccacaccgccctctaccgc- tcgccgcttcgccagggcgccgac ccccagccggcttcaggagacgcttatgccgcccattaa 289 atgagccaaacgatcgataaaattcacagctgttatccgatatttgaacaggatgaataccagaccctgt- tccagaataaaaagacccttgaagaggcg cacgacgcgcagcgtgtgcaggaggtttttgcctggaccaccaccgccgagtatgaagcgctgaacttccagc- gcgaggcgctgaccgtcgaccc ggccaaagcctgccagccgctcggcgccgtactctgcgcgctggggttcgccggcaccctgccctacgtgcac- ggctcccagggctgcgtcgcct atttcgcacctactttaaccgccattttaaagagccggtcgcctgcgtctccgactccatgaccgaggacgcg- gcggtgttcggcggcaacaacaac atgaatctgggcctgcacaatgccagcgcgctgtataaacccgagattatcgccgtctccaccacctgtatgg- ccgaggtgatcggcgacgatctgca ggcgtttatcgccaacgccaaaaaagagggatttgttgacgaccgcatcgccattccttacgcccataccccc- agctttatcggcagccatgtcaccgg ctgggacaatatgttcgaagggttcgcgaagacctttaccgctgactacgccgggaagccgggcaaacagaaa- aagctcaatctggtgaccggattt gagacctatctcggcaacttccgcgtgctgaagcggatgatggcgcagatggatgtcccgtgcagcctgctct- ccgacccatcagaggtgctcgaca cccccgccgacggccattaccggatgtacgccggcggcaccagccagcaggagatcaaaaccgcgccggacgc- cattgacaccctgctgctgca gccgtggcagatggtgaaaagcaaaaaggtggttcaggagatgtggaaccagcccgccaccgaggtggccgtt- ccgctgggcctggacgccacc gacgcgctgctgatgaccgtcagtcagctgaccggcaaaccgatcgccgacgctctgaccctggagagaggcc- ggctggtcgacatgatgctggat tcccacacctggctgcatggcaaaaaattcggcctctacggcgatccggatttcgtgatggggctgacgcgct- tcctgctggagctgggctgcgagcc gacggtgatcctcagtcataacgccaataaacgctggcaaaaagcgatgaagaaaatgctcgatgcctcgccg- tacggtcaggaaagcgaagtgttc atcaactgcgacctgtggcacttccggtcgctgatgttcacccgtcagccggactttatgatcggtaactcct- acggcaagtttatccagcgcgataccc tggcaaagggcaaagccttcgaagtgccgatgatccgtctgggctttccgctgttcgaccgccatcatctgca- ccgccagaccacctggggctatgaa ggcgcaatgaacatcgtcacgacgctggtgaacgccgtgctggaaaaactggaccacgacaccagccagttgg- gcaaaaccgattacagcttcgac ctcgttcgttaa 290 atgatgccgctttctccgcaattacagcagcactggcagacggtcgctgaccgtctgccagaggattttc- ccattgccgaactgagcccacaggccag gtcggtcatggcgttcagcgattttgtcgaacagagtgtgatcgcccagccgggctggctgaatgagcttgag- gactcctcgccggaggcggaagag tggcggcattacgaggcctggctgcaggatcgcctgcaggccgtcactgacgaagggggttgatgcgagagct- gcgtctcttccgccgccagatg atggtccgcatcgcctgggcgcaggcgctgtcgctggtgagcgaagaagagactctgcagcagctgagcgtcc- tggcggagaccctgattgtcgcc gcccgcgactggctgtacgccgcctgctgtaaggagtggggaacgccatgcaatgccgagggccagccgcagc- cgctgctgatcctcgggatgg gaaagctgggcggcggcgagctgaacttctcttccgatatcgatctgatctttgcctggcctgagcatggcgc- cacccgcggcggccgccgcgagct ggataacgcccagttctttacccgtctggggcagcggctgatcaaggcccttgaccagccgacgcaggacggc- tttgtctatcgggttgacatgcgcc tgcggccgtttggcgacagtgggccgctggtactcagttttgcggcgctggaagattattaccaggagaaggg- tcgggactgggaacgctatgcgat ggtgaaagcgcggatcatggccgataacgacggcgtgtacgccagcgacttgcgcgcgatgctccgtcctttc- gtcttccgccgttatatcgacttca gcgtgatccagtcgctgcgtaacatgaaaggcatgatcgcccgcgaagtgcggcgtcgcgggctgaaagacaa- catcaagctcggcgccggcgg gatccgtgaaattgagtttatcgttcaggctttcaactgatccgcggtggtcgcgaacctgcactgcagcagc- gcgccctgctgccgacgctggcgg cgattgatgagctacatctgctgccggaaggcgacgcggcgctgctgcgcgaggcctatctgttcctgcgccg- gctggaaaacctgctgcaaagcat caacgatgagcagacccagaccctgccgcaggatgaacttaaccgcgccaggctggcgtgggggatgcatacc- gaagactgggagacgctgagc gcgcagctggcgagccagatggccaacgtgcggcgagtgtttaatgaactgatcggcgatgatgaggatcagt- ccccggatgagcaactggccga gtactggcgcgagctgtggcagcatgcgctggaagaagatgacgccagcccggcgctggcgcatttaaacgat- accgaccgccgtagcgtgctgg cgctgattgccgattttcgtaaagagctggatcggcgcaccatcggcccgcgcggccgccaggtgctggatca- gcttgatgccgcatctgctgagcga aatctgctcgcgcgccgatgcgccgctgcctctggcgcggatcacgccgctgttgaccgggatcgtcacccgt- accacctatcttgagctgctgagc gaattccccggcgcgctgaagcacctgatcacgctctgcgcggcgtcgccgatggtcgccagccagctggcgc- gccacccgctgctgctggatga gctgctggatcccaacaccctctatcagccgacggcgaccgatgcctatcgcgacgagctgcgccagtacctg- ctgcgcgtgccggaagaggatga agagcagcagatggaggcgttgcgccagtttaagcaggcgcagcagctgcatatcgcggcggcggatatcgct- ggtaccctgccggtgatgaagg tcagcgatcacttaacctggcttgccgaagcgatcctcgacgcggtggtgcagcaggcatgggggcagatggt- cgctcgctacggccagccgaccc acctgcacgatcgccagggtcgcggcttcgccgtcgtcggctacgctaagcttggcggctgggagctgggcta- cagctccgatctcgatctggtgttc ctccatgactgcccggcggaggtgatgaccgacggcgagcgggagattgacggccgtcagttctacctgcggc- tggcccagcggatcatgcacct gttcagcacccgcacctcgtccggtattctctacgaagttgacgcccggagcgccttctggcgcggcggggat- gaggtcaccaccgccgacgc gtttgctgactatcagcagaacgaagcaggacgtgggaacatcaggcgctggtgcgcgcccgcgtggtctatg- gcgacccggcgctgcaggcgc gctttgacgccattcgtcgcgatatcctgaccaccccgcgggaggggatgaccctgcagaccgaggttcgcga- gatgcgcgagaagatgcgcgcc caccttggcaacaaacatcccgatcgttttgatatcaaagccgatgccggcgggatcaccgatattgaattta- ttactcagtatctggtcctacgctatgcc agtgacaagccgaagctgacccgctggtctgacaacgtgcgtattcttgagctgctggcgcagaacgacatca- tggaggaggaggaggcgcgcgc cttaacgcatgcgtacaccaccttgcgtgatgcgctccatcacctggccctgcaggagcagccgggacacgtg- gcgccagaggccttcagccggga gcgtcagcaggtcagcgccagctggcagaagtggctgatggcttaa 291 agtctgaactcatcctgcggcagtcggtgagacgtatttttgaccaaagagtgatctacatcacggaatt- ttgtggttgttgctgcttaaaagggcaaatct acccttagaatcaactgttatatcagggggattcagagagatattaggaatttgcacaagcgcacaatttaac- cacatcatgataacgccatgtaaaaca aagataaaaaaacaaaatgcagtgacttacatcgcaagcaaggcattttcttatccaattgctcaaagtttgg- cctttcatatcgcaacgaaaatgcgtaat atacgcgcccttgcggacatcagtatggtcattcctagttcatgcgcatcggacaccaccagcttacaaattg- cctgattgcggccccgatggccggtat cactgaccgaccatttcgtgccttatgtcatgcgatgggggctgg 292 tgaacatcactgatgcacaagctacctatgacgaagaattaactaaaaaactgcaagatgaggcattcgc- gttaaagccgacttgagaaatgagaag attggctttaaaattcgcgaacacacgctacgccgtgttcatatatgttagtttgtggcgataaagaggtcga- agcaggcaaagttgctgttcgtacccg ccgcggcaaagacttaggaagcatggatgttagcgaagtcgttgacaaactgctggcggaaatccgcagcaga- agtcttcatcaactggaggaataa agtattaaaggcggaaaacgagttcaaccggcgcgtcctaatcgcattaacaaagagattcgcgcgcaagaag- ttcgcctcacaggcgtcgatggcg agcagattggtattgtcagtagaatgaagctcttgaaaaagctgaggaaggggcgtcgatttagtagaaatca- gtccgaatgccgagccgccagttt gtcgaatc 293 tacagtagcgcctctcaaaaatagataaacggctcatgtacgtgggccgtttattttttctacctataat- cgggaaccggtgttataatgccgcgccctcat attgtggggattcttaatgacctatcctgggtcctaaagttgtagttgacattagcggagcactaac 294 aattttttttcacaaagcgtagcgttattgaatcgcacattttaaactgttggccgctgtggaagcgaat- attcgtgaaagtgcggttttaaggcctttttctt tgactctctgtcgttacaaagttaatatgcgcgccct 295 ttaaaaacgtgaccacgagcattaataaacgccacgaaatgtggcgtttatttattcaaaaagtatctct- ttcataaaaagtgctaaatgcagtagcagca aaattgggataagtcccatggaatacggctgttttcgctgcaatttttaactttttcgtaaaaaaagatgttt- ctttgagcgaacgatcaaaatatagcgttaa ccggcaaaaaattattctcattagaaaatagtttgtgtaatacttgtaacgctacatggagattaacttaatc- tagagggttttata 296 atggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtcgcgacacaacttgcacgtcatc- ctttattgctcgatgaactgctcgacccgc gcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatctgatgcgggtgccaac- agnagacgaagaacagcagcttg aagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccggt- gatgaaagtcagtgaccatttaacc taccttgccgaggccattctcgatgtcgtggtgcagcatgcgtgggaacaaatggtcgtaaaatacgggcagc- ccgcgcatcttcagcaccgtgagg ggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctggttatagctcagatctggatctg- gtcttcctgctcgattgcgcgccgg aggtgatgacggacggcgaacgcagcatcgacggacgtcagttttatcttcggctggcgcagcgcattatgca- cttattcagcacccggacatcgtca ggcattctttacgaggttgatccgcgtctgcgaccttccggcgcatccggcatgctggtcagtaccattgaag- cgtttgcagattatcaggccaatgaag cctggacgtgggagcatcaggcgctggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaatt- taacgccacgcgtcgcgacattct ttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccatctg- gggagtaaaaaagcccacgagtt tgactgaaagccgatccgggtggcatcacggatattgaattcattgcacaatacctggttctgcgtttcgcgc- atgatgagccgagctgacgcgctg gtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgccggaagaggaagcgcgccatctg- acgcaggcttatgtgacgctgcgog atgaaattcatcatctggcgttgcaggaacacagcgggaaagtggccgcggacagctttgctactgagcgcgc- gcagatccgtgccagctgggcga agtggctcggctga 297 cggtactggaacagaaatcggcggatgcgcaggaaatttgttatgacacggcctgtctgaagtgcaagtt- agtgcttacttcctggctggcaacctcag gctggacgccgtttattgatgataaatctgcgaagaaactggacgcttccttcaaacgttttgctgacatcat- gctcggtcgtaccgcagcggatctgaaa gaggcctttgcgcagccactgacggaagaaggttatcgcgatcagctggcgcgcctgaaacgccagatcatta- ccttccatttgatgccggtgcttac cctgaaaaagacgtcgatgcgtatattgccggctgggtggacctgcaacaggccatcgttcagcagcaacacg- cctgggaggattcggcccgtctc acgcggtgatgatggatgctttctggttaaacgggcaacctcgttaactgactgactagcctgggcaaactgc- ccgggcttttttttgcaaggaatctgat ttcatggcgctcaaacagttaatccgtctgtgtgccgcctcgccgatggtcgcgacacaacttgcacgtcatc- ctttattgctcgatgaactgctcgaccc gcgcacgctttaccagccgattgagccgggcgcttaccgcgacgaactgcgtcagtatctgatgcgggtgcca- acagaagacgaagaacagcagct tgaagccgtgcgccagttcaaacaggcccagcatttgcgtatcgcagccggggatatttccggggcattgccg- gtgatgaangtcagtgaccatttaa cctaccttgccgaggccattctcgatgtcgggtgcagcatgctgtgggaacaaatggtcgtaaaatacgggca- gcccgcgcatcttcagcaccgtga ggggcgcggttttgccgtggtcggttacgggaaactcggtggctgggagctgggttatagctcagatctggat- ctggtcttcctgctcgattgcgcgcc ggaggtgatgacggacggcgaacgcagcatcgacggacgtcagttttatcttcggctggcgcagcgcattatg- cacttattcagcacccggacatcgt caggcattctttacgaggttgatccgcgtctgcgaccttccggcgcatccggcatgctggtcagtaccattga- agcgtttgcagattatcaggccaatga agcctggacgtgggagcatcaggcgctggttcgcgcgcgcgtggtttacggggatccgcaactgacacagcaa- tttaacgccacgcgtcgcgacat tctttgccgccagcgcgatggcgacggcctgcgtaaggaggtccgtgaaatgcgcgagaaaatgtatgcccat- ctggggagtaaaaaagcccacga gtttgatagaaagccgatccgggtggcatcacggatattgaattcattgcacaatacctggttctgcgtttcg- cgcatgatgagccgaagctgacgcgc tggtctgataacgtgcggatttttgaactgatggcacgatatgacatcatgccggaagaggaagcgcgccatc- tgacgcaggcttatgtgacgctgcg cgatgaaattcatcatctggcgttgcaggaacacagcgggaaagtggccgcggacagctttgctactgagcgc- gcgcagatccgtgccagctgggc aaagtggctcggctgagggtttttattcggctaacaggcgcttgtgatattatccggcgcattgtatttaccc- gatttgatttatctgttttggagtcttgggat gaaagtgactttgcagattttcaccgcgcaggtgtgctggttgtcggtgacgtaatgttagaccgttactggt- atggcccgaccaatcgtatttctccgga agctccggtgccggtggtgaaggtcagtaccattgaagagcggcctggcggtgcagctaacgtggcgatgaac- atttcatctctgggcgcctcttcct gtctgatcggcctgaccggcgtagacgacgctgcgcgtgccctcagtgagcgtctggcagaagtgaaagttaa- ctgcgttttcgtcgcactatccaca catcctaccatcaccaaactgcgaattttgtcccgtaaccagcaactgatccgcctcgactttgaggaaggtt- ttgaaggcgttgatctcgagccgatgct gaccaaaataga 298 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggagtctgaactcatcctgcgatgggggctgggccgtctctgaagctctcggtgaac- attgttgcgaggcaggatgcgagct ggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaagggcgttttcc- cgtccggggaatggcatggagctgc gccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatggcggcacctt- gctgacgttacgcctgccggtacagca ggttatcaccggaggcttaaaatga 299 tgtttcgtctcgaggccgggcaactgagggccccgttgaaaccgacctgggctggcatctgttgttgtgc- gaacaaattcgcctgccgcaaccttgc cgaaagcccaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgcccgttttgcacgaaaaatgcacataaattgcct-
gcgttgccttataacaggcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaacgcctgcaaattgcacggttattccgggtgagtatatgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgacgggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgcc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggagtctgaactcatcctgcgatgggggctgggccgtctctgaagctctcgg- tgaacattgttgcgaggcaggatg cgagctggttgtgttttcacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccggaagcggc- gttttcccgtccggggaatggcatcg agctgcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccctgatggcgg- caccttgctgacgttacgcctgccggt acagcaggttatcaccggaggcttaaaatgacccagttacctaccgcgcgcccggttatccggcgctttgata- tgtctgcccagtttacggcgctttatc gcatcagcgtggcgctgagtcaggaaagcaacaccgggcgcgcactggcggcgatcctcgaagtgcttcacga- tcatgcatttatgcaatacggcat ggtgtgtctgtttgataaagaacgcaatgcactctttgtggaatccctgcatggcatcgacggcgaaaggaaa- aaagagacccgccatgtccgttacc gcatgggggaaggcgtgatcggcgcggtgatgagccagcgtcaggcgctggtgttaccgcgcatttcagacga- tcagcgttttctcgaccccctgaa tatttacgattacagcctgccgttgattggcgtgccgatccccggtgcggataatcagccatcgggcgtgctg- gtggcacagccgatggcgttgcacg aagaccggctgactgccagtacgcggtttttagaaatggtc 300 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggttaaagcctgccggtacagcaggttatcaccggaggcttaaatga 301 tgtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttgttgtg- cgaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gagcaacaactgatcaacgcct gagcctgttctccttcttgttgatgcagacgggttaatgcccgttttgcacgaaaaatgcacataaatttgcc- tgcgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatctgtgtgatttgggtt- ccggcattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caattcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgc- cagacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggttaaagcctgccggtacagcaggttatcaccggaggcttaaaatgaccca- gttacctaccgcgggcccggttat ccggcgctttgatatgtctgcccagtttacggcgctttatcgcatcagcgtggcgctgagtcaggaaagcaac- accgggcgcgcactggcggcgatc ctcgaagtgcttcacgatcatgcatttatgcaatacggcatggtgtgtctgtttgataaagaacgcaatgcac- tctttgtggaatccctgcatggcatcgac ggcgaaaggaaaaaagagacccgccatgtccgttaccgcatgggggaaggcgtgatcggcgcggtgatgagcc- agcgtcaggcgctggtgttac cgcgcatttcagacgatcagcgttttctcgaccgcctgaatatttacgattacagcctgccgttgattggcgt- gccgatccccggtgcggataatcagcc atcgggcgtgctggtggcacagccgatggcgttgcacgaagaccggctgactgccagtacgcggtttttagaa- atggtc 302 atgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaacatc- cttcactgttttataccgtggttgaacaatct tcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccgccagacgg- gttttgcacttgagacacttttgggcg agaaccaccgtctgctggagtctgaactcatcctgcggcagtcggtgagacgtatttttgaccaaagagtgat- ctacatcacggaattttgtggttgttgc tgcttaaaagggcaaatctacccttagaatcaactgttatatcagggggattcagagagatattaggaatttg- cacaagcgcacaatttaaccacatcatg ataacgccatgtaaaacaaagataaaaaaacaaaatgcagtgacttacatcgcaagcaaggcattttcttatc- caattgctcaaagtttggcccttcatatc gcaacgaaaatgcgtaatatacgcgcccttgcggacatcagtatggtcattcctagttcatgcgcatcggaca- ccaccagcttacaaattgcctgattgc ggccccgatggccggtatcactgaccgaccatttcgtgccttatgtcatgcgatgggggctggcccgtctctg- aagctctcggtgaacattgttgcgag gcaggatgcgagctggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgcccgcccg- gaagcggcgttttcccgtccgggga atggcatggagagcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtccgccct- gatggcggcaccttgctgacgttacg cctgccggtacagcaggttatcaccggaggcttaaaatga 303 tgtttcgtctcgaggccgggcaactgagcggccccgttgaaaccgacctgggctggcatctgttgttgtg- cgaacaaattcgcctgccgcaacccttgc cgaaagccgaagccttaacgcgggtgcgtcagcaactgattgcccggcaacagaaacattatcagcgccagtg- gctgcaacaactgatcaacgcct gagcctgttctccttcttgttgatgagacgggttaatgcccgttttgcacggaaaatgcacataaattgcctg- cgttgccttataacagcgcagggaaat cctgcctccggccttgtgccacaccgcgctttgcctggtttgtggtaaaaactggcccgctttgcatcctgat- gcttaaaacaccccgttcagatcaacct ttgggcagataagcccgcgaaaggcctgcaaattgcacggttattccgggtgagtatatgtgtcatttgggtt- ccgccattgcgcaataaaggggaga aagacatgagcatcacggcgttatcagcatcatttcctgaggggaatatcgccagccgcttgtcgctgcaaca- tccttcactgttttataccgtggttgaa caatcttcggtggcgatttcgctgaccgatccgcaggcgcgcatttgttatgccaatccggcattctgccccc- agacgggttttgcacttgagacactttt gggcgagaaccaccgtctgctggagtctgaactcatcctgcggcagtcggtgagacgtatttttgaccaaaga- gtgatctacatcacggaattttgtggt tgttgctgcttaaaagggcaaatctacccttagaatcaactgttatatcagggggattcagagagatattagg- aatttgcacaagcgcacaatttaaccac atcatgataacgccatgtaaaacaaagataaaaaaacaaaatgcagtgacttacatcgcaagcaaggcatttt- cttatccaattgctcaaagtttggccttt catatcgcaacgaaaatgcgtaatatacgcgcccttgcggacatcagtatggtcattcctagttcatgcgcat- cggacaccaccagcttacaaattgcct gattgcggccccgatggccggtatcactgaccgaccatttcgtgccttatgtcatgcgatgggggctgggccg- tctctgaagctctcggtgaacattgtt gcgaggcaggatgcgagctggttgtgttttgacattaccgataatgtgccgcgtgaacgggtgcgttatgccc- gcccggaagcggcgttttcccgtcc ggggaatggcatggagctgcgccttatccagacgctgatcgcccatcatcgcggttctttagatctctcggtc- cgccctgatggcggcaccttgctgac gttacgcctgccggtacagcaggttatcaccggaggcttaaaatgacccagttacctaccgcgggcccggtta- tccggcgctttgatatgtctgcccag tttacggcgctttatcgcatcagcgtggcgctgagtcacgaaagcaacaccgggcgcgcactggcgccgatcc- tcgaagtgcttcacgatcatgcatt tatgcaatacggcatggtgtgctgtttgataaagaacgcaatgcactctttgtggaatccctgcatggcatcg- acggcgaaaggaaaaaagagaccc gccatgtccgttaccgcatgggggaaggcgtgatcggcgcggtgatgagccagcgtcaggcgctggtgttacc- gcgcatttcagacgatcagcgttt tctcgaccgcctgaatatatttacgattacagcctgccgttgattggcgtgccgatccccggtgcggataatc- agccatcgggcgtgctggtggcacagcc gatggcgttgcacgaagaccggctgactgccagtacgcggtttttagaaatggtc
[0433] The use of the terms "a" and "an" and "the" and similar referents in the context of describing the invention (especially in the context of the following claims) are to be construed to cover both the singular and the plural, unless otherwise indicated herein or clearly contradicted by context. The terms "comprising," "having," "including," and "containing" are to be construed as open-ended terms (i.e., meaning "including, but not limited to,") unless otherwise noted. Recitation of ranges of values herein are merely intended to serve as a shorthand method of referring individually to each separate value falling within the range, unless otherwise indicated herein, and each separate value is incorporated into the specification as if it were individually recited herein. For example, if the range 10-15 is disclosed, then 11, 12, 13, and 14 are also disclosed. All methods described herein can be performed in any suitable order unless otherwise indicated herein or otherwise clearly contradicted by context. The use of any and all examples, or exemplary language (e.g., "such as") provided herein, is intended merely to better illuminate the invention and does not pose a limitation on the scope of the invention unless otherwise claimed. No language in the specification should be construed as indicating any non-claimed element as essential to the practice of the invention.
CLAUSES
[0434] 1. A method of increasing nitrogen fixation in a non-leguminous plant, comprising: [0435] a. applying to the plant a plurality of non-intergeneric bacteria, said plurality comprising non-intergeneric bacteria that: [0436] i. have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or [0437] ii. produce fixed N of at least about 1.times.10.sup.4' mmol N per bacterial cell per hour, and [0438] wherein the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant, and [0439] wherein each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen. [0440] 2. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0441] 3. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise bacteria that: produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0442] 4. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0443] 5. The method according to clause 1, wherein the at least one genetic variation comprises an introduced control sequence operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0444] 6. The method according to clause 1, wherein the at least one genetic variation comprises a promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0445] 7. The method according to clause 1, wherein the at least one genetic variation comprises an inducible promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0446] 8. The method according to clause 1, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene of the nitrogen fixation or assimilation genetic regulatory network. [0447] 9. The method according to clause 1, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene in the nif gene cluster. [0448] 10. The method according to clause 1, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation. [0449] 11. The method according to clause 1, wherein the plurality of non-intergeneric bacteria applied to the plant do not stimulate an increase in the uptake of exogenous non-atmospheric nitrogen. [0450] 12. The method according to clause 1, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight. [0451] 13. The method according to clause 1, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight, and wherein the nitrogen-containing fertilizer comprises ammonium or an ammonium containing molecule. [0452] 14. The method according to clause 1, wherein the exogenous nitrogen is selected from fertilizer comprising one or more of: glutamine, ammonia, ammonium, urea, nitrate, nitrite, ammonium-containing molecules, nitrate-containing molecules, and nitrite-containing molecules. [0453] 15. The method according to clause 1, wherein the plurality of non-intergeneric bacteria, in plan/a, produce 5% or more of the fixed nitrogen in the plant. [0454] 16. The method according to clause 1, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0455] 17. The method according to clause 1, wherein the plurality of non-intergeneric bacteria, in planta, fix atmospheric nitrogen in non-nitrogen-limiting conditions. [0456] 18. The method according to clause 1, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation. [0457] 19. The method according to clause 1, wherein the fixed nitrogen produced by the plurality of non-intergeneric bacteria is measured through dilution of enriched fertilizer by atmospheric N2 gas in plant tissue. [0458] 20. The method according to clause 1, wherein the at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network are selected from the group consisting of: nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifD, nifK, nifY, nifE, nifL, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme. [0459] 21. The method according to clause 1, wherein the at least one genetic variation is a mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. [0460] 22. The method according to clause 1, wherein the at least one genetic variation is selected from: (A) a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion. [0461] 23. The method according to clause 1, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene. [0462] 24. The method according to clause 1, wherein the at least one genetic variation is a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0463] 25. The method according to clause 1, wherein the at least one genetic variation is a mutated amtB gene that results in the lack of expression of said amtB gene. [0464] 26. The method according to clause 1, wherein the at least one genetic variation is selected from: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof. [0465] 27. The method according to clause 1, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0466] 28. The method according to clause 1, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated amtB gene that results in the lack of expression of said amtB gene. [0467] 29. The method according to clause 1, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome. [0468] 30. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are formulated into a composition. [0469] 31. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are formulated into a composition comprising an agriculturally acceptable carrier. [0470] 32. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are applied into furrows in which seeds of said plant are planted. [0471] 33. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are formulated into a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 du per milliliter. [0472] 34. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are applied onto a seed of said plant. [0473] 35. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating and are applied onto a seed of said plant. [0474] 36. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating and are applied onto a seed of said plant, at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. [0475] 37. The method according to clause 1, wherein the plant is a cereal crop. [0476] 38. The method according to clause 1, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0477] 39. The method according to clause 1, wherein the plant is corn. [0478] 40. The method according to clause 1, wherein the plant is an agricultural crop plant. [0479] 41. The method according to clause 1, wherein the plant is a genetically modified organism. [0480] 42. The method according to clause 1, wherein the plant is not a genetically modified organism. [0481] 43. The method according to clause 1, wherein the plant has been genetically engineered or bred for efficient nitrogen use. [0482] 44. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise at least two different species of bacteria. [0483] 45. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise at least two different strains of the same species of bacteria. [0484] 46. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0485] 47. The method according to clause 1, wherein the plurality of non-intergeneric bacteria are endophytic, epiphytic, or rhizospheric. [0486] 48. The method according to clause 1, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: a bacteria deposited as PTA-122293, a bacteria deposited as PTA-122294, a bacteria deposited as NCMA 201701002, a bacteria deposited as NCMA 201708004, a bacteria deposited as NCMA 201708003, a bacteria deposited as NCMA 201708002, a bacteria deposited as NCMA 201712001, a bacteria deposited as NCMA 201712002, and combinations thereof. [0487] 49. A bacterial composition, comprising: [0488] at least one genetically engineered bacterial strain that fixes atmospheric nitrogen in an agricultural system that has been fertilized with more than 20 lbs of Nitrogen per acre. [0489] 50. A bacterial composition, comprising: [0490] at least one bacterial strain that has been bred to fix atmospheric nitrogen in an agricultural system that has been fertilized with more than 20 lbs of Nitrogen per acre. [0491] 51. The bacterial composition of clause 49 or clause 50, wherein said fertilizer is a chemical fertilizer selected from the group consisting of anhydrous ammonia, ammonia sulfate, urea, diammonium phosphate, urea-form, UAN (urea ammonium nitrate) monoammonium phosphate, ammonium nitrate, nitrogen solutions, calcium nitrate, potassium nitrate, and sodium nitrate. [0492] 52. A bacterial composition, comprising: [0493] at least one genetically engineered bacterial strain that fixes atmospheric nitrogen, the at least one genetically engineered bacterial strain comprising exogenously added DNA wherein said exogenously added DNA shares at least 80% identity to a corresponding native bacterial strain. [0494] 53. The bacterial composition of clause 52, wherein said exogenously added DNA shares at least 85% identity to a corresponding native bacterial strain. [0495] 54. The bacterial composition of clause 52, wherein said exogenously added DNA shares at least 90% identity to a corresponding native bacterial strain. [0496] 55. The bacterial composition of clause 52, wherein said exogenously added DNA shares at least 95% identity to a corresponding native bacterial strain. [0497] 56. The bacterial composition of clause 52, wherein said exogenously added DNA shares at least 99% identity to a corresponding native bacterial strain. [0498] 57. The bacterial composition of clause 52, wherein said exogenously added DNA is derived from a same bacterial strain as said corresponding native bacterial strain. [0499] 58. The bacterial composition of any of the preceding clauses, wherein said bacterial composition is a fertilizing composition. [0500] 59. The bacterial composition of any of the preceding clauses, wherein said at least one genetically engineered bacterial strain comprises at least one variation in a gene or intergenic region within 10,000 bp of a gene selected from the group consisting of: nifA, nifL, ntrB, ntrC, glnA, glnB, glnK, draT, amtB, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, and nifQ. [0501] 60. The bacterial composition of any of the preceding clauses, further comprising at least one additional component selected from the group consisting of a tackifier, a microbial stabilizer, a fungicide, an antibacterial agent, a preservative, a stabilizer, a surfactant, an anti-complex agent, an herbicide, a nematicide, an insecticide, a plant growth regulator, a fertilizer, a rodenticide, a dessicant, a bactericide, and a nutrient. [0502] 61. The bacterial composition of any of the preceding clauses, further comprising at least one additional component selected from the group consisting of a colorant, primer, pellet, disinfectant, plant growth regulator, safener, and a nematicide. [0503] 62. The bacterial composition of any of the preceding clauses, wherein said bacterial composition is formulated to be applied to a field. [0504] 63. The bacterial composition of clause 62, wherein said bacterial composition is formulated to be applied in-furrow.
[0505] 64. The bacterial composition of clause 62, wherein said bacterial composition is formulated to be applied as a seed coating, seed dressing, or seed treatment. [0506] 65. The bacterial composition of clause 61 or clause 62, wherein said bacterial composition is formulated to be applied at, prior to, or post planting of the seed. [0507] 66. A seed composition comprising a seed of a plant that is inoculated with a bacterial composition of any of the preceding clauses. [0508] 67. A method of growing a crop using a plurality of seeds having a seed composition of clause 66. [0509] 68. The method of clause 67, further comprising harvesting said crop. [0510] 69. A method of applying a bacterial composition of any of the preceding clauses to a field. [0511] 70. The method of clause 69, wherein said bacterial composition is applied to said field in a form selected from the group consisting of a liquid form, a dry form, a granule, a powder, and a pellet. [0512] 71. The method of clause 69, wherein said bacterial composition is applied to said field as a seed coating, seed dressing, or seed treatment. [0513] 72. The method of clause 69, wherein said bacterial composition is applied to said field as an in-furrow treatment. [0514] 73. The method of clause 69, wherein said bacterial composition is applied to said field at, prior to, or post planting of the seed. [0515] 74. A fertilizer composition comprising a bacterial composition of any of the preceding clauses. [0516] 75. The fertilizer composition of clause 74, wherein said fertilizer composition is a seed coating, seed dressing, or seed treatment composition. [0517] 76. The fertilizer composition of clause 74, wherein said fertilizer composition is an in-furrow composition. [0518] 77. The fertilizer composition of clause 74, wherein said fertilizer composition is provided to a crop at, prior to, or post planting. [0519] 78. The fertilizer composition of clause 74, further comprising a porous carrier. [0520] 79. The fertilizer composition of clause 74, further comprising an additional synergistic component that, when combined with said bacterial composition, increases a fertilizing benefit of said fertilizer composition to a crop that is beyond a cumulative benefit of its individual components. [0521] 80. A method of maintaining soil nitrogen levels, comprising: [0522] planting, in soil of a field, a crop inoculated by a genetically engineered bacterium that fixes atmospheric nitrogen; and [0523] harvesting said crop, wherein no more than 90% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0524] 81. The method of clause 80, wherein no more than 80% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0525] 82. The method of clause 80, wherein no more than 70% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0526] 83. The method of clause 80, wherein no more than 60% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0527] 84. The method of clause 80, wherein no more than 50% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0528] 85. The method of clause 80, wherein no more than 40% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0529] 86. The method of clause 80, wherein said genetically engineered bacterium comprises a bacterial composition of any of the preceding clauses. [0530] 87. The method of clause 80, wherein said genetically engineered bacterium consists of a bacterial composition of any of the preceding clauses. [0531] 88. A method of delivering a probiotic supplement to a crop plant, comprising: [0532] coating a crop seed with a seed coating, seed treatment, or seed dressing, wherein said seed coating, seed dressing, or seed treatment comprises living representatives of said probiotic; and [0533] applying, in soil of a field, said crop seeds. [0534] 89. The method of clause 88, wherein said seed coating, seed dressing, or seed treatment is applied in a single layer to said crop seed. [0535] 90. The method of clause 88, wherein said seed coating is applied in multiple layers to said crop seed. [0536] 91. The method of clause 88, wherein said seed coating is applied in a blend to said crop seed. [0537] 92. The method of clause 88, wherein said crop seed is non-nodulating. [0538] 93. The method of clause 88, wherein said seed coating comprises a bacterial composition of any of the preceding clauses. [0539] 94. The method of any of the proceeding clauses, wherein the genetically engineered bacterial strain is a genetically engineered Gram-positive bacterial strain. [0540] 95. The method of clause 94, wherein the genetically engineered Gram-positive bacterial strain has an altered expression level of a regulator of a Nif cluster. [0541] 96. The method of clause 94, wherein the genetically engineered Gram-positive bacterial strain expresses a decreased amount of a negative regulator of a Nif cluster. [0542] 97. The method of clause 94, wherein the genetically engineered bacterial strain expresses a decreased amount of GlnR. [0543] 98. The method of any one of the proceeding clauses, wherein the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 75% identity to a sequence from the Zehr lab NifH database. [0544] 99. The method of any one of the proceeding clauses, wherein the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 85% identity to a sequence from the Zehr lab NifH database. [0545] 100. The method of any one of the proceeding clauses, wherein the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 75% identity to a sequence from the Buckley lab NifH database. [0546] 101. The method of any one of the proceeding clauses, wherein the genome of the genetically engineered bacterial strain encodes a polypeptide with at least 85% identity to a sequence from the Buckley lab NifH database. [0547] 102. A method of breeding microbial strains to improve specific traits of agronomic relevance, comprising: [0548] providing a plurality of microbial strains that have the ability to colonize a desired crop; [0549] improving regulatory networks influencing the trait through intragenomic rearrangement; [0550] assessing microbial strains within the plurality of microbial strains to determine a measure of the trait; and [0551] selecting one or more microbial strains of the plurality of microbial strains that generate an improvement in the trait in the presence of the desired crop. [0552] 103. The method of clause 102, wherein the specific trait which is improved is the ability of the microbial strain to fix nitrogen. [0553] 104. The method of clause 103, wherein the specific trait which is improved is the ability of the microbial strain to fix atmospheric nitrogen in the presence of N-fertilized growing conditions. [0554] 105. A method of breeding microbial strains to improve specific traits of agronomic relevance, comprising: [0555] providing a plurality of microbial strains that have the ability to colonize a desired crop; [0556] introducing genetic diversity into the plurality of microbial strains; [0557] assessing microbial strains within the plurality of microbial strains to determine a measure of the trait; and [0558] selecting one or more microbial strains of the plurality of microbial strains that generate an improvement in the trait in the presence of the desired crop. [0559] 106. The method of clause 105, wherein the specific trait which is improved is the ability of the microbial strain to fix nitrogen. [0560] 107. The method of clause 106, wherein the specific trait which is improved is the ability of the microbial strain to fix atmospheric nitrogen in the presence of N-fertilized growing conditions. [0561] 108. A method of increasing the amount of atmospheric derived nitrogen in a non-leguminous plant, comprising: [0562] exposing said non-leguminous plant to engineered non-intergeneric microbes, said engineered non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network or at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network. [0563] 109. The method of clause 108, wherein said engineered non-intergeneric microbes comprise at least one genetic variation introduced into said nitrogen fixation genetic regulatory network. [0564] 110. The method of clause 108, wherein said engineered non-intergeneric microbes comprise at least one genetic variation introduced into said nitrogen assimilation genetic regulatory network. [0565] 111. The method of clause 108, wherein said engineered non-intergeneric microbes comprise at least one genetic variation introduced into said nitrogen fixation genetic regulatory network and at least one genetic variation introduced into said nitrogen assimilation genetic regulatory network. [0566] 112. The method of clause 108, wherein said engineered non-intergeneric microbes are applied into furrows in which seeds of said non-leguminous plant are planted. [0567] 113. The method of clause 108, wherein said engineered non-intergeneric microbes are coated onto a seed of said non-leguminous plant. [0568] 114. The method of clause 108, wherein said non-leguminous plant is a non-leguminous agricultural crop plant selected from the group consisting of sorghum, canola, tomato, strawberry, barley, rice, corn, wheat, potato, millet, cereals, grains, and maize. [0569] 115. The method of clause 108, wherein said engineered non-intergeneric microbes colonize at least a root of said non-leguminous plant such that said engineered non-intergeneric microbes are present in said non-leguminous plant in an amount of at least 10.sup.5 colony forming units per gram fresh weight of tissue. [0570] 116. The method of clause 108, wherein said engineered non-intergeneric microbes are capable of fixing atmospheric nitrogen in non-nitrogen-limiting conditions. [0571] 117. The method of clause 108, wherein said engineered non-intergeneric microbes, in planta, excrete nitrogen-containing products of nitrogen fixation. [0572] 118. The method of clause 108, wherein said at least one genetic variation is introduced into a gene selected from the group consisting of nifA, nut, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nit)), nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme. [0573] 119. The method of clause 108, wherein said engineered non-intergeneric microbes, in planta, produce at least 1% of fixed nitrogen in said non-leguminous plant. [0574] 120. The method of clause 119, wherein said fixed nitrogen in said non-leguminous plant produced by said engineered non-intergeneric microbes is measured by dilution of 15N in crops grown in fields treated with fertilizer containing 1.2% 15N. [0575] 121. The method of clause 119, wherein said engineered non-intergeneric microbes, in planta, produce 5% or more of the fixed nitrogen in said non-leguminous plant. [0576] 122. The method of clause 108, wherein said non-intergeneric microbes are engineered using at least one type of engineering selected from the group consisting of directed mutagenesis, random mutagenesis, and directed evolution. [0577] 123. A method of increasing an amount of atmospheric derived nitrogen in a corn plant, comprising [0578] exposing said corn plant to engineered non-intergeneric microbes comprising engineered genetic variations within at least two genes selected from the group consisting of nifL, glnB, glnE, and amtB. [0579] 124. The method of clause 123, wherein said engineered non-intergeneric microbes, in planta, excrete nitrogen-containing products of nitrogen fixation. [0580] 125. The method of clause 123, wherein said engineered non-intergeneric microbes are applied into furrows in which seeds of said corn plant are planted. [0581] 126. The method of clause 123, wherein said engineered non-intergeneric microbes are coated onto a seed of said corn plant. [0582] 127. A method of increasing an amount of atmospheric derived nitrogen in a corn plant, comprising: [0583] exposing said corn plant to engineered non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network, wherein said engineered non-intergeneric microbes, in planta, produce at least 5% of fixed nitrogen in said corn plant as measured by dilution of 15N in crops grown in fields treated with fertilizer containing 1.2% 15N. [0584] 128. A method of increasing nitrogen fixation in a non-leguminous plant, comprising: [0585] a. applying to the plant a plurality of non-intergeneric bacteria, said plurality comprising non-intergeneric bacteria that: [0586] i. have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and [0587] ii. produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and [0588] wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.5.times.10.sup.4 mmol N per gram of fresh weight of plant root tissue per hour, and [0589] wherein the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant, and wherein each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen. [0590] 129. The method according to clause 128, wherein the at least one genetic variation comprises an introduced control sequence operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0591] 130. The method according to clause 128, wherein the at least one genetic variation comprises a promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0592] 131. The method according to clause 128, wherein the at least one genetic variation comprises an inducible promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network.
[0593] 132. The method according to clause 128, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene of the nitrogen fixation or assimilation genetic regulatory network. [0594] 133. The method according to clause 128, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene in the nif gene cluster. [0595] 134. The method according to clause 128, wherein the plurality of non-intergeneric bacteria, in planta, excrete the nitrogen-containing products of nitrogen fixation. [0596] 135. The method according to clause 128, wherein the plurality of non-intergeneric bacteria applied to the plant do not stimulate an increase in the uptake of exogenous non-atmospheric nitrogen. [0597] 136. The method according to clause 128, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight. [0598] 137. The method according to clause 128, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight, and wherein the nitrogen-containing fertilizer comprises ammonium or an ammonium containing molecule. [0599] 138. The method according to clause 128, wherein the exogenous nitrogen is selected from fertilizer comprising one or more of: glutamine, ammonia, ammonium, urea, nitrate, nitrite, ammonium-containing molecules, nitrate-containing molecules, and nitrite-containing molecules. [0600] 139. The method according to clause 128, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0601] 140. The method according to clause 128, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0602] 141. The method according to clause 128, wherein the plurality of non-intergeneric bacteria, in planta, fix atmospheric nitrogen in non-nitrogen-limiting conditions. [0603] 142. The method according to clause 128, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation. [0604] 143. The method according to clause 128, wherein the fixed nitrogen produced by the plurality of non-intergeneric bacteria is measured through dilution of enriched fertilizer by atmospheric N2 gas in plant tissue. [0605] 144. The method according to clause 128, wherein the at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network are selected from the group consisting of: nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifQ, nifA, and a gene associated with biosynthesis of a nitrogenase enzyme. [0606] 145. The method according to clause 128, wherein the at least one genetic variation is a mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. [0607] 146. The method according to clause 128, wherein the at least one genetic variation is selected from: (A) a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion. [0608] 147. The method according to clause 128, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene. [0609] 148. The method according to clause 128, wherein the at least one genetic variation is a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0610] 149. The method according to clause 128, wherein the at least one genetic variation is a mutated amtB gene that results in the lack of expression of said amtB gene. [0611] 150. The method according to clause 128, wherein the at least one genetic variation is selected from: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said n/IL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof. [0612] 151. The method according to clause 128, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0613] 152. The method according to clause 128, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated amtB gene that results in the lack of expression of said amtB gene. [0614] 153. The method according to clause 128, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome. [0615] 154. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are formulated into a composition. [0616] 155. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are formulated into a composition comprising an agriculturally acceptable carrier. [0617] 156. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are applied into furrows in which seeds of said plant are planted. [0618] 157. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are formulated into a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter. [0619] 158. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are applied onto a seed of said plant. [0620] 159. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating and are applied onto a seed of said plant. [0621] 160. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating and are applied onto a seed of said plant, at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. [0622] 161. The method according to clause 128, wherein the plant is a cereal crop. [0623] 162. The method according to clause 128, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0624] 163. The method according to clause 128, wherein the plant is corn. [0625] 164. The method according to clause 128, wherein the plant is an agricultural crop plant. [0626] 165. The method according to clause 128, wherein the plant is a genetically modified organism. [0627] 166. The method according to clause 128, wherein the plant is not a genetically modified organism. [0628] 167. The method according to clause 128, wherein the plant has been genetically engineered or bred for efficient nitrogen use. [0629] 168. The method according to clause 128, wherein the plurality of non-intergeneric bacteria comprise at least two different species of bacteria. [0630] 169. The method according to clause 128, wherein the plurality of non-intergeneric bacteria comprise at least two different strains of the same species of bacteria. [0631] 170. The method according to clause 128, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0632] 171. The method according to clause 128, wherein the plurality of non-intergeneric bacteria are endophytic, epiphytic, or rhizospheric. [0633] 172. The method according to clause 128, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: a bacteria deposited as PTA-122293, a bacteria deposited as PTA-122294, a bacteria deposited as NCMA 201701002, a bacteria deposited as NCMA 201708004, a bacteria deposited as NCMA 201708003, a bacteria deposited as NCMA 201708002, a bacteria deposited as NCMA 201712001, a bacteria deposited as NCMA 201712002, and combinations thereof. [0634] 173. A method of increasing nitrogen fixation in a non-leguminous plant, comprising: [0635] a. applying to the plant a plurality of bacteria, said plurality comprising bacteria that: [0636] i. have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or [0637] ii. produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and [0638] wherein the plurality of bacteria, in planta, produce 1% or more of the fixed nitrogen in the plant. [0639] 174. The method according to clause 173, wherein the plurality of bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0640] 175. The method according to clause 173, wherein the plurality of bacteria comprise bacteria that: produce fixed N of at least about 173.times.10.sup.-17 mmol N per bacterial cell per hour. [0641] 176. The method according to clause 173, wherein the plurality of bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0642] 177. The method according to clause 173, wherein the plurality of bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.5.times.10.sup.-8 mmol N per gram of fresh weight of plant root tissue per hour. [0643] 178. The method according to clause 173, wherein the plurality of bacteria, in planta, excrete the nitrogen-containing products of nitrogen fixation. [0644] 179. The method according to clause 173, wherein the plurality of bacteria applied to the plant do not stimulate an increase in the uptake of exogenous non-atmospheric nitrogen. [0645] 180. The method according to clause 173, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome. [0646] 181. The method according to clause 173, wherein the plurality of bacteria are formulated into a composition. [0647] 182. The method according to clause 173, wherein the plurality of bacteria are formulated into a composition comprising an agriculturally acceptable carrier. [0648] 183. The method according to clause 173, wherein the plurality of bacteria are applied into furrows in which seeds of said plant are planted. [0649] 184. The method according to clause 173, wherein the plurality of bacteria are formulated into a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter. [0650] 185. The method according to clause 173, wherein the plurality of bacteria are applied onto a seed of said plant. [0651] 186. The method according to clause 173, wherein the plurality of bacteria are formulated into a seed coating and are applied onto a seed of said plant. [0652] 187. The method according to clause 173, wherein the plurality of bacteria are formulated into a seed coating and are applied onto a seed of said plant, at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. [0653] 188. The method according to clause 173, wherein the plant is a cereal crop. [0654] 189. The method according to clause 173, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0655] 190. The method according to clause 173, wherein the plant is corn. [0656] 191. The method according to clause 173, wherein the plant is an agricultural crop plant. [0657] 192. The method according to clause 173, wherein the plant is a genetically modified organism. [0658] 193. The method according to clause 173, wherein the plant is not a genetically modified organism. [0659] 194. The method according to clause 173, wherein the plant has been genetically engineered or bred for efficient nitrogen use. [0660] 195. The method according to clause 173, wherein the plurality of bacteria comprise at least two different species of bacteria. [0661] 196. The method according to clause 173, wherein the plurality of bacteria comprise at least two different strains of the same species of bacteria. [0662] 197. The method according to clause 173, wherein the plurality of bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0663] 198. The method according to clause 173, wherein the plurality of bacteria are endophytic, epiphytic, or rhizospheric. [0664] 199. The method according to clause 173, wherein the plurality of bacteria are selected from: a bacteria deposited as NCMA 201701003, a bacteria deposited as NCMA 201701001, and a bacteria deposited as NCMA 201708001. [0665] 200. A non-intergeneric bacterial population capable of increasing nitrogen fixation in a non-leguminous plant, comprising: [0666] a. a plurality of non-intergeneric bacteria, that: [0667] i. have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or [0668] ii. produce fixed N of at least about 1
.times.10.sup.-17 mmol N per bacterial cell per hour, and [0669] wherein the plurality of non-intergeneric bacteria, in planta, produce 1% or more of the fixed nitrogen in a plant grown in the presence of the plurality of non-intergeneric bacteria, and [0670] wherein each member of the plurality of non-intergeneric bacteria comprises at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacteria are capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen. [0671] 201. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0672] 202. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise bacteria that: produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0673] 203. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gam of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0674] 204. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation comprises an introduced control sequence operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0675] 205. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation comprises a promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0676] 206. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation comprises an inducible promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0677] 207. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene of the nitrogen fixation or assimilation genetic regulatory network. [0678] 208. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria do not comprise a constitutive promoter operably linked to a gene in the nif gene cluster. [0679] 209. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation. [0680] 210. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria applied to the plant do not stimulate an increase in the uptake of exogenous non-atmospheric nitrogen. [0681] 211. The non-intergeneric bacterial population according to clause 200, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight. [0682] 212. The non-intergeneric bacterial population according to clause 200, wherein the plant is grown in soil from a field which has been administered at least about 50 lbs of nitrogen-containing fertilizer per acre, and wherein the nitrogen-containing fertilizer comprises at least about 5% nitrogen by weight, and wherein the nitrogen-containing fertilizer comprises ammonium or an ammonium containing molecule. [0683] 213. The non-intergeneric bacterial population according to clause 200, wherein the exogenous nitrogen is selected from fertilizer comprising one or more of: glutamine, ammonia, ammonium, urea, nitrate, nitrite, ammonium-containing molecules, nitrate-containing molecules, and nitrite-containing molecules. [0684] 214. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0685] 215. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0686] 216. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria, in planta, fix atmospheric nitrogen in non-nitrogen-limiting conditions. [0687] 217. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation. [0688] 218. The non-intergeneric bacterial population according to clause 200, wherein the fixed nitrogen produced by the plurality of non-intergeneric bacteria is measured through dilution of enriched fertilizer by atmospheric N.sub.2 gas in plant tissue. [0689] 219. The non-intergeneric bacterial population according to clause 200, wherein the at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network are selected from the group consisting of: nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme. [0690] 220. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is a mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. [0691] 221. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is selected from: (A) a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion. [0692] 222. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is a mutated/tiff, gene that has been altered to comprise a heterologous promoter inserted into said nifL gene. [0693] 223. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0694] 224. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is a mutated amtB gene that results in the lack of expression of said amtB gene. [0695] 225. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is selected from: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof. [0696] 226. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0697] 227. The non-intergeneric bacterial population according to clause 200, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated amtB gene that results in the lack of expression of said amtB gene. [0698] 228. The non-intergeneric bacterial population according to clause 200, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome. [0699] 229. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria are formulated into a composition. [0700] 230. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria are formulated into a composition comprising an agriculturally acceptable carrier. [0701] 231. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria are formulated into a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter. [0702] 232. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria are formulated into a seed coating. [0703] 233. The non-intergeneric bacterial population according to clause 200, wherein the plant is a cereal crop. [0704] 234. The non-intergeneric bacterial population according to clause 200, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0705] 235. The non-intergeneric bacterial population according to clause 200, wherein the plant is corn. [0706] 236. The non-intergeneric bacterial population according to clause 200, wherein the plant is an agricultural crop plant. [0707] 237. The non-intergeneric bacterial population according to clause 200, wherein the plant is a genetically modified organism. [0708] 238. The non-intergeneric bacterial population according to clause 200, wherein the plant is not a genetically modified organism. [0709] 239. The non-intergeneric bacterial population according to clause 200, wherein the plant has been genetically engineered or bred for efficient nitrogen use. [0710] 240. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise at least two different species of bacteria. [0711] 241. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise at least two different strains of the same species of bacteria. [0712] 242. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium morale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0713] 243. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria are endophytic, epiphytic, or rhizospheric. [0714] 244. The non-intergeneric bacterial population according to clause 200, wherein the plurality of non-intergeneric bacteria comprise bacteria selected from: a bacteria deposited as PTA-122293, a bacteria deposited as PTA-122294, a bacteria deposited as NCMA 201701002, a bacteria deposited as NCMA 201708004, a bacteria deposited as NCMA 201708003, a bacteria deposited as NCMA 201708002, a bacteria deposited as NCMA 201712001, a bacteria deposited as NCMA 201712002, and combinations thereof [0715] 245. A bacterial population capable of increasing nitrogen fixation in a non-leguminous plant, comprising: [0716] a. a plurality of bacteria, that: [0717] i. have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or [0718] ii. produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and [0719] wherein the plurality of bacteria, in planta, produce 1% or more of the fixed nitrogen in die plant. [0720] 246. The bacterial population according to clause 245, wherein the plurality of bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0721] 247. The bacterial population according to clause 245, wherein the plurality of bacteria comprise bacteria that: produce fixed N of at least about 245.times.10.sup.-17 mmol N per bacterial cell per hour. [0722] 248. The bacterial population according to clause 245, wherein the plurality of bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0723] 249. The bacterial population according to clause 245, wherein the plurality of bacteria comprise bacteria that: have an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produce fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.5.times.mmol N per gram of fresh weight of plant root tissue per hour. [0724] 250. The bacterial population according to clause 245, wherein the plurality of bacteria, in planta, excrete the nitrogen-containing products of nitrogen fixation. [0725] 251. The bacterial population according to clause 245, wherein the plurality of bacteria applied to the plant do not stimulate an increase in the uptake of exogenous non-atmospheric nitrogen. [0726] 252. The bacterial population according to clause 245, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome. [0727] 253. The bacterial population according to clause 245, wherein the plurality of bacteria are formulated into a composition. [0728] 254. The bacterial population according to clause 245, wherein the plurality of bacteria are formulated into a composition comprising an agriculturally acceptable carrier. [0729] 255. The bacterial population according to clause 245, wherein the plurality of bacteria are applied into furrows in which seeds of said plant are planted.
[0730] 256. The bacterial population according to clause 245, wherein the plurality of bacteria are formulated into a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter. [0731] 257. The bacterial population according to clause 245, wherein the plurality of bacteria are applied onto a seed of said plant. [0732] 258. The bacterial population according to clause 245, wherein the plurality of bacteria are formulated into a seed coating and are applied onto a seed of said plant. [0733] 259. The bacterial population according to clause 245, wherein the plurality of bacteria are formulated into a seed coating and are applied onto a seed of said plant, at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. [0734] 260. The bacterial population according to clause 245, wherein the plant is a cereal crop. [0735] 261. The bacterial population according to clause 245, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0736] 262. The bacterial population according to clause 245, wherein the plant is corn. [0737] 263. The bacterial population according to clause 245, wherein the plant is an agricultural crop plant. [0738] 264. The bacterial population according to clause 245, wherein the plant is a genetically modified organism. [0739] 265. The bacterial population according to clause 245, wherein the plant is not a genetically modified organism. [0740] 266. The bacterial population according to clause 245, wherein the plant has been genetically engineered or bred for efficient nitrogen use. [0741] 267. The bacterial population according to clause 245, wherein the plurality of bacteria comprise at least two different species of bacteria. [0742] 268. The bacterial population according to clause 245, wherein the plurality of bacteria comprise at least two different strains of the same species of bacteria. [0743] 269. The bacterial population according to clause 245, wherein the plurality of bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0744] 270. The bacterial population according to clause 245, wherein the plurality of bacteria are endophytic, epiphytic, or rhizospheric. [0745] 271. The bacterial population according to clause 245, wherein the plurality of bacteria are selected from: a bacteria deposited as NCMA 201701003, a bacteria deposited as NCMA 201701001, and a bacteria deposited as NCMA 201708001. [0746] 272. A bacterium that: [0747] i. has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or [0748] ii. produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0749] 273. A non-intergeneric bacterium, comprising: at least one genetic variation introduced into at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network, such that the non-intergeneric bacterium is capable of fixing atmospheric nitrogen in the presence of exogenous nitrogen, and wherein said bacterium: [0750] i. has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue; and/or [0751] ii. produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0752] 274. The non-intergeneric bacterium according to clause 273, wherein the bacterium has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0753] 275. The non-intergeneric bacterium according to clause 273, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0754] 276. The non-intergeneric bacterium according to clause 273, wherein the bacterium has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue and produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour. [0755] 277. The non-intergeneric bacterium according to clause 273, wherein the bacterium has an average colonization ability per unit of plant root tissue of at least about 1.0.times.10.sup.4 bacterial cells per grain of fresh weight of plant root tissue and produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour, and wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.5.times.10.sup.-8 mmol N per gram of fresh weight of plant root tissue per hour. [0756] 278. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation comprises an introduced control sequence operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0757] 279. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation comprises a promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0758] 280. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation comprises an inducible promoter operably linked to the at least one gene of the nitrogen fixation or assimilation genetic regulatory network. [0759] 281. The non-intergeneric bacterium according to clause 273, wherein the bacterium does not comprise a constitutive promoter operably linked to a gene of the nitrogen fixation or assimilation genetic regulatory network. [0760] 282. The non-intergeneric bacterium according to clause 273, wherein the bacterium does not comprise a constitutive promoter operably linked to a gene in the nif gene cluster. [0761] 283. The non-intergeneric bacterium according to clause 273, wherein the bacterium, in planta, excretes the nitrogen-containing products of nitrogen fixation. [0762] 284. The non-intergeneric bacterium according to clause 273, wherein the at least one gene, or non-coding polynucleotide, of the nitrogen fixation or assimilation genetic regulatory network are selected from the group consisting of: nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme. [0763] 285. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is a mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. [0764] 286. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is selected from: (A) a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion. [0765] 287. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is a mutated MIL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene. [0766] 288. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0767] 289. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is a mutated amtB gene that results in the lack of expression of said amtB gene. [0768] 290. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is selected from: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof. [0769] 291. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0770] 292. The non-intergeneric bacterium according to clause 273, wherein the at least one genetic variation is a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said tuft gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated amtB gene that results in the lack of expression of said amtB gene. [0771] 293. The non-intergeneric bacterium according to clause 273, formulated into a composition. [0772] 294. The non-intergeneric bacterium according to clause 273, formulated into a composition comprising an agriculturally acceptable carrier. [0773] 295. The non-intergeneric bacterium according to clause 273, formulated into a liquid in-furrow composition. [0774] 296. The non-intergeneric bacterium according to clause 273, formulated into a seed coating. [0775] 297. The non-intergeneric bacterium according to clause 273, wherein said bacterium is selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0776] 298. The non-intergeneric bacterium according to clause 273, wherein said bacterium is endophytic, epiphytic, or rhizospheric. [0777] 299. The non-intergeneric bacterium according to clause 273, wherein said bacterium is selected from: a bacteria deposited as PTA-122293, a bacteria deposited as PTA-122294, a bacteria deposited as NCMA 201701002, a bacteria deposited as NCMA 201708004, a bacteria deposited as NCMA 201708003, a bacteria deposited as NCMA 201708002, a bacteria deposited as NCMA 201712001, a bacteria deposited as NCMA 201712002, and combinations thereof. [0778] 300. A method of increasing nitrogen fixation in a plant, comprising administering to the plant an effective amount of a composition comprising: [0779] i. a purified population of bacteria that comprises bacteria with a 16S nucleic acid sequence that is at least about 97% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, 277-283; [0780] ii. a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, 284-295; and/or [0781] iii. a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260, 296-303; and [0782] wherein the plant administered the effective amount of the composition exhibits an increase in nitrogen fixation, as compared to a plant not having been administered the composition. [0783] 301. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a 16S nucleic acid sequence that is at least about 97% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, and 277-283. [0784] 302. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a 16S nucleic acid sequence that is at least about 99% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, and 277-283. [0785] 303. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a 165 nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, and 277-283. [0786] 304. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0787] 305. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 95% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0788] 306. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 99% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0789] 307. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0790] 308. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303. [0791] 309. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 95% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303. [0792] 310. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence that is at least about 99% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303.
[0793] 311. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprises bacteria with a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303. [0794] 312. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with at least one genetic variation introduced into a gene selected from the group consisting of nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifH, nifD, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme. [0795] 313. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with at least one genetic variation introduced into a gene selected from the group consisting of nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme, wherein the genetic variation (A) is a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion. [0796] 314. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with at least one mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, or AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. [0797] 315. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a mutated MIL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene. [0798] 316. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0799] 317. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a mutated amtB gene that results in the lack of expression of said amtB gene. [0800] 318. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with at least one of the following genetic alterations: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof. [0801] 319. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0802] 320. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated am/B gene that results in the lack of expression of said amtB gene. [0803] 321. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a nifL modification that expresses a Nil protein at less than about 50% of a bacteria lacking the nifL modification. [0804] 322. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a nifL modification that expresses a Nil protein at less than about 50% of a bacteria lacking the nifL modification, wherein the modification is a disruption, knockout, or truncation. [0805] 323. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a raft, modification that expresses a Nil protein at less than about 50% of a bacteria lacking the nifL modification, and wherein the bacteria further comprise a promoter operably linked to a nifA sequence. [0806] 324. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a nifL modification that expresses a Nil protein at less than about 50% of a bacteria lacking the nifL modification, and wherein the bacteria lack a nifL homolog. [0807] 325. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification. [0808] 326. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification, wherein the modification is a disruption, knockout, or a truncation. [0809] 327. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification, and wherein the GlnE protein sequence lacks an adenylyl removing (AR) domain. [0810] 328. The method of clause 300, wherein the composition comprises: a purified population of bacteria that comprise bacteria with a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification, and wherein the GlnE protein sequence lacks adenylyl removing (AR) activity. [0811] 329. The method of clause 300, wherein the purified population of bacteria, in planta, provide at least 1% of fixed nitrogen to the plant. [0812] 330. The method of clause 300, wherein the purified population of bacteria, in planta, provide at least 5% of fixed nitrogen to the plant. [0813] 331. The method of clause 300, wherein the purified population of bacteria, in planta, provide at least 10% of fixed nitrogen to the plant. [0814] 332. The method of clause 300, wherein the purified population of bacteria are capable of fixing atmospheric nitrogen in non-nitrogen-limiting conditions. [0815] 333. The method of clause 300, wherein the purified population of bacteria, in planta, excrete nitrogen-containing products of nitrogen fixation. [0816] 334. The method of clause 300, wherein the purified population of bacteria, in planta, provide at least 1% of fixed nitrogen to the plant, and wherein said fixed nitrogen is measured through dilution of enriched fertilizer by atmospheric N2 gas in plant tissue. [0817] 335. The method of clause 300, wherein the purified population of bacteria colonize a root of said plant and are present in an amount of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0818] 336. The method of clause 300, wherein the plant comprises the seed, stalk, flower, fruit, leaves, or rhizome. [0819] 337. The method of clause 300, wherein the composition comprises: an agriculturally acceptable carrier. [0820] 338. The method of clause 300, wherein the composition comprising the purified population of bacteria is administered into furrows in which seeds of said plant are planted. [0821] 339. The method of clause 300, wherein the composition comprising the purified population of bacteria is formulated as a liquid in-furrow composition comprising bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 du per milliliter. [0822] 340. The method of clause 300, wherein the composition comprising the purified population of bacteria is administered onto a seed of said plant. [0823] 341. The method of clause 300, wherein the composition comprising the purified population of bacteria is formulated as a seed coating and is administered onto a seed of said plant. [0824] 342. The method of clause 300, wherein the composition comprising the purified population of bacteria is formulated as a seed coating and is administered onto a seed of said plant, at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. [0825] 343. The method of clause 300, wherein the plant is non-leguminous. [0826] 344. The method of clause 300, wherein the plant is a cereal crop. [0827] 345. The method of clause 300, wherein the plant is selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0828] 346. The method of clause 300, wherein the plant is corn. [0829] 347. The method of clause 300, wherein the plant is a legume. [0830] 348. The method of clause 300, wherein the plant is a grain crop. [0831] 349. The method of clause 300, wherein the purified population of bacteria comprise bacteria selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0832] 350. The method of clause 300, wherein the purified population of bacteria comprise a bacteria of genus Rahnella. [0833] 351. The method of clause 300, wherein the purified population of bacteria comprise a bacteria of species Rahnella aquatilis. [0834] 352. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as PTA-122293. [0835] 353. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201701003. [0836] 354. The method of clause 300, wherein the purified population of bacteria comprise a bacteria of genus Kosakonia. [0837] 355. The method of clause 300, wherein the purified population of bacteria comprise a bacteria of species Kosakonia sacchari. [0838] 356. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as PTA-122294. [0839] 357. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201701001. [0840] 358. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201701002. [0841] 359. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201708004. [0842] 360. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201708003. [0843] 361. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201708002. [0844] 362. The method of clause 300, wherein the purified population of bacteria comprise a bacteria of genus Klebsiella [0845] 363. The method of clause 300, wherein the purified population of bacteria comprise a bacteria of species Klebsiella variicola. [0846] 364. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201708001. [0847] 365. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201712001. [0848] 366. The method of clause 300, wherein the purified population of bacteria comprise a bacteria deposited as NCMA 201712002. [0849] 367. An isolated bacteria, comprising: [0850] i. a 16S nucleic acid sequence that is at least about 97% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, 277-283; [0851] ii. a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, 284-295; and/or [0852] iii. a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260, 296-303. [0853] 368. The isolated bacteria of clause 367, comprising: a 1.65 nucleic acid sequence that is at least about 97% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, and 277-283. [0854] 369. The isolated bacteria of clause 367, comprising: a 16S nucleic acid sequence that is at least about 99% identical to a nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, and 277-283. [0855] 370. The isolated bacteria of clause 367, comprising: a 16S nucleic acid sequence selected from SEQ ID NOs: 85, 96, 111, 121, 122, 123, 124, 136, 149, 157, 167, 261, 262, 269, and 277-283. [0856] 371. The isolated bacteria of clause 367, comprising: a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0857] 372. The isolated bacteria of clause 367, comprising: a nucleic acid sequence that is at least about 95% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0858] 373. The isolated bacteria of clause 367, comprising: a nucleic acid sequence that is at least about 99% identical to a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0859] 374. The isolated bacteria of clause 367, comprising: a nucleic acid sequence selected from SEQ ID NOs: 86-95, 97-110, 112-120, 125-135, 137-148, 150-156, 158-166, 168-176, 263-268, 270-274, 275, 276, and 284-295. [0860] 375. The isolated bacteria of clause 367, comprising: a nucleic acid sequence that is at least about 90% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303. [0861] 376. The isolated bacteria of clause 367, comprising: a nucleic acid sequence that is at least about 95% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303. [0862] 377. The isolated bacteria of clause 367, comprising: a nucleic acid sequence that is at least about 99% identical to a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303.
[0863] 378. The isolated bacteria of clause 367, comprising: a nucleic acid sequence selected from SEQ ID NOs: 177-260 and 296-303. [0864] 379. The isolated bacteria of clause 367, comprising: at least one genetic variation introduced into a gene selected from the group consisting of nifA, nifL, ntrB, ntrC, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme. [0865] 380. The isolated bacteria of clause 367, comprising: at least one genetic variation introduced into a gene selected from the group consisting of nifA, nifL, ntrB, polynucleotide encoding glutamine synthetase, glnA, glnB, glnK, drat, amtB, polynucleotide encoding glutaminase, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, nifQ, and a gene associated with biosynthesis of a nitrogenase enzyme, wherein the genetic variation (A) is a knock-out mutation; (B) alters or abolishes a regulatory sequence of a target gene; (C) comprises the insertion of a heterologous regulatory sequence; or (D) a domain deletion. [0866] 381. The isolated bacteria of clause 367, comprising: at least one mutation that results in one or more of: increased expression or activity of NifA or glutaminase; decreased expression or activity of NifL, NtrB, glutamine synthetase, GlnB, GlnK, DraT, or AmtB; decreased adenylyl-removing activity of GlnE; or decreased uridylyl-removing activity of GlnD. [0867] 382. The isolated bacteria of clause 367, comprising: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene. [0868] 383. The isolated bacteria of clause 367, comprising: a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0869] 384. The isolated bacteria of clause 367, comprising: a mutated amtB gene that results in the lack of expression of said amtB gene. [0870] 385. The isolated bacteria of clause 367, comprising: at least one of the following genetic alterations: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said MIL gene; a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain; a mutated amtB gene that results in the lack of expression of said amtB gene; and combinations thereof. [0871] 386. The isolated bacteria of clause 367, comprising: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain. [0872] 387. The isolated bacteria of clause 367, comprising: a mutated nifL gene that has been altered to comprise a heterologous promoter inserted into said nifL gene and a mutated glnE gene that results in a truncated GlnE protein lacking an adenylyl-removing (AR) domain and a mutated amtB gene that results in the lack of expression of said amtB gene. [0873] 388. The isolated bacteria of clause 367, comprising: a nifL modification that expresses a NifL protein at less than about 50% of a bacteria lacking the nifL modification. [0874] 389. The isolated bacteria of clause 367, comprising: a MIL modification that expresses a nifL protein at less than about 50% of a bacteria lacking the nifL modification, wherein the modification is a disruption, knockout, or truncation. [0875] 390. The isolated bacteria of clause 367, comprising: a nifL modification that expresses a NifL protein at less than about 50% of a bacteria lacking the nifL modification, and wherein the bacteria further comprise a promoter operably linked to a nifA sequence. [0876] 391. The isolated bacteria of clause 367, comprising: a nifL modification that expresses a NifL protein at less than about 50% of a bacteria lacking the nifL modification, and wherein the bacteria lack a nifL homolog. [0877] 392. The isolated bacteria of clause 367, comprising: a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification. [0878] 393. The isolated bacteria of clause 367, comprising: a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification, wherein the modification is a disruption, knockout, or a truncation. [0879] 394. The isolated bacteria of clause 367, comprising: a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification, and wherein the GlnE protein sequence lacks an adenylyl removing (AR) domain. [0880] 395. The isolated bacteria of clause 367, comprising: a glnE modification that expresses a GlnE protein at less than about 50% of a bacteria lacking the glnE modification, and wherein the GlnE protein sequence lacks adenylyl removing (AR) activity. [0881] 396. The isolated bacteria of clause 367, wherein the bacteria, in planta, provide at least 1% of fixed nitrogen to a plant exposed to said bacteria. [0882] 397. The isolated bacteria of clause 367, wherein the bacteria, in planta, provide at least 5% of fixed nitrogen to a plant exposed to said bacteria. [0883] 398. The isolated bacteria of clause 367, wherein the bacteria, in planta, provide at least 10% of fixed nitrogen to a plant exposed to said bacteria. [0884] 399. The isolated bacteria of clause 367, wherein the bacteria is capable of fixing atmospheric nitrogen in non-nitrogen-limiting conditions. [0885] 400. The isolated bacteria of clause 367, wherein the bacteria, in planta, excretes nitrogen-containing products of nitrogen fixation. [0886] 401. The isolated bacteria of clause 367, wherein the bacteria, in planta, provide at least 1% of fixed nitrogen to a plant exposed to said bacteria, and wherein said fixed nitrogen is measured through dilution of enriched fertilizer by atmospheric N.sub.2 gas in plant tissue. [0887] 402. The isolated bacteria of clause 367, wherein the bacteria colonize a root of a plant exposed to said bacteria to a concentration of at least about 1.0.times.10.sup.4 bacterial cells per gram of fresh weight of plant root tissue. [0888] 403. The isolated bacteria of clause 367, formulated into an agricultural composition. [0889] 404. The isolated bacteria of clause 367, formulated into an in-furrow composition. [0890] 405. The isolated bacteria of clause 367, formulated as a liquid in-furrow composition that comprises bacteria at a concentration of about 1.times.10.sup.7 to about 1.times.10.sup.10 cfu per milliliter. [0891] 406. The isolated bacteria of clause 367, formulated as a seed treatment or seed coating. [0892] 407. The isolated bacteria of clause 367, formulated as a seed treatment or seed coating that comprises bacteria at a concentration of about 1.times.10.sup.5 to about 1.times.10.sup.7 cfu per seed. [0893] 408. The isolated bacteria of clause 367, wherein the bacteria is in contact with a plant and increases nitrogen fixation in the plant. [0894] 409. The isolated bacteria of clause 367, disposed on a non-leguminous plant. [0895] 410. The isolated bacteria of clause 367, disposed on a cereal crop. [0896] 411. The isolated bacteria of clause 367, disposed on a plant selected from the group consisting of: corn, rice, wheat, barley, sorghum, millet, oat, rye, and triticale. [0897] 412. The isolated bacteria of clause 367, disposed on corn. [0898] 413. The isolated bacteria of clause 367, disposed on a legume. [0899] 414. The isolated bacteria of clause 367, disposed on a grain crop. [0900] 415. The isolated bacteria of clause 367, selected from: Rahnella aquatilis, Klebsiella variicola, Achromobacter spiritinus, Achromobacter marplatensis, Microbacterium murale, Kluyvera intermedia, Kosakonia pseudosacchari, Enterobacter sp., Azospirillum lipoferum, Kosakonia sacchari, and combinations thereof. [0901] 416. The isolated bacteria of clause 367, wherein the bacteria is of the genus Rahnella. [0902] 417. The isolated bacteria of clause 367, wherein the bacteria is of the species Rahnella aquatilis. [0903] 418. The isolated bacteria of clause 367, deposited as PTA-122293. [0904] 419. The isolated bacteria of clause 367, deposited as NCMA 201701003. [0905] 420. The isolated bacteria of clause 367, wherein the bacteria is of the genus Kosakonia. [0906] 421. The isolated bacteria of clause 367, wherein the bacteria is of the species Kosakonia sacchari. [0907] 422. The isolated bacteria of clause 367, deposited as PTA-122294. [0908] 423. The isolated bacteria of clause 367, deposited as NCMA 201701001. [0909] 424. The isolated bacteria of clause 367, deposited as NCMA 201701002. [0910] 425. The isolated bacteria of clause 367, deposited as NCMA 201708004. [0911] 426. The isolated bacteria of clause 367, deposited as NCMA 201708003. [0912] 427. The isolated bacteria of clause 367, deposited as NCMA 201708002. [0913] 428. The isolated bacteria of clause 367, wherein the bacteria is of the genus Klebsiella. [0914] 429. The isolated bacteria of clause 367, wherein the bacteria is of the species Klebsiella variicola. [0915] 430. The isolated bacteria of clause 367, deposited as NCMA 201708001. [0916] 431. The isolated bacteria of clause 367, deposited as NCMA 201712001. [0917] 432. The isolated bacteria of clause 367, deposited as NCMA 201712002. [0918] 433. A composition comprising any one or more bacteria of clauses 415 to 432. [0919] 434. A method of detecting a non-native junction sequence, comprising: amplifying a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ED NOs: 372-405. [0920] 435. The method according to clause 434, wherein said amplifying is by conducting a polymerase chain reaction. [0921] 436. The method according to clause 434, wherein said amplifying is by conducting a quantitative polymerase chain reaction. [0922] 437. A method of detecting a non-native junction sequence, comprising: amplifying a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to at least a 10 contiguous base pair fragment contained in SEQ ID NOs: 372-405, said contiguous base pair fragment being comprised of nucleotides at the intersection of: an upstream sequence comprising SEQ ID NOs: 304-337 and downstream sequence comprising SEQ ID NOs: 338-371. [0923] 438. The method according to clause 437, wherein said amplifying is by conducting a polymerase chain reaction. [0924] 439. The method according to clause 437, wherein said amplifying is by conducting a quantitative polymerase chain reaction. [0925] 440. A non-native junction sequence, comprising: a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NOs: 372-405. [0926] 441. A non-native junction sequence, comprising: a nucleotide sequence that shares at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 1.00% sequence identity to at least a 10 contiguous base pair fragment contained in SEQ ID NOs: 372-405, said contiguous base pair fragment being comprised of nucleotides at the intersection of: an upstream sequence comprising SEQ ID NOs: 304-337 and downstream sequence comprising SEQ ID NOs: 338-371. [0927] 442. A bacterial composition, comprising: at least one remodeled bacterial strain that fixes atmospheric nitrogen, the at least one remodeled bacterial strain comprising exogenously added DNA wherein said exogenously added DNA shares at least 80% identity to a corresponding native bacterial strain. [0928] 443. A method of maintaining soil nitrogen levels, comprising: [0929] planting, in soil of a field, a crop inoculated by a remodeled bacterium that fixes atmospheric nitrogen; and [0930] harvesting said crop, wherein no more than 90% of a nitrogen dose required for producing said crop is administered to said soil of said field between planting and harvesting. [0931] 444. A method of delivering a probiotic supplement to a crop plant, comprising: [0932] coating a crop seed with a seed coating, seed treatment, or seed dressing, wherein said seed coating, seed dressing, or seed treatment comprising living representatives of said probiotic; and [0933] applying in soil of a field, said crop seeds. [0934] 445. A method of increasing the amount of atmospheric derived nitrogen in a non-leguminous plant, comprising: [0935] exposing said non-leguminous plant to remodeled non-intergeneric microbes, said remodeled non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network or at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network. [0936] 446. A method of increasing an amount of atmospheric derived nitrogen in a corn plant, comprising: [0937] exposing said corn plant to remodeled non-intergeneric microbes comprising remodeled genetic variations within at least two genes selected from the group consisting of nifL, glnB, glnE, and amtB. [0938] 447. A method of increasing an amount of atmospheric derived nitrogen in a corn plant, comprising: [0939] exposing said corn plant to remodeled non-intergeneric microbes comprising at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation genetic regulatory network, wherein said remodeled non-intergeneric microbes, in planta, produce at least 5% of fixed nitrogen in said corn plant as measured by dilution of 15N in crops grown in fields treated with fertilizer containing 1.2% 15N. [0940] 448. The method of any of the previous clauses, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen. [0941] 449. The bacterium of any of the previous clauses, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen. [0942] 450. The bacterial population of any of the previous clauses, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen. [0943] 451. The isolated bacteria of any of the previous clauses, wherein the bacterium produces fixed N of at least about 1
.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen. [0944] 452. The non-intergeneric bacterial population of any of the previous clauses, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen. [0945] 453. The non-intergeneric bacterium of any of the previous clauses, wherein the bacterium produces fixed N of at least about 1.times.10.sup.-17 mmol N per bacterial cell per hour when in the presence of exogenous nitrogen. [0946] 454. The method of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0947] 455. The method of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0948] 456. The bacterium of any of the previous clauses, wherein the plurality of bacteria, in plank; produce 5% or more of the fixed nitrogen in the plant. [0949] 457. The bacterial population of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0950] 458. The isolated bacteria of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0951] 459. The non-intergeneric bacterial population of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0952] 460. The non-intergeneric bacterium of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 5% or more of the fixed nitrogen in the plant. [0953] 461. The method of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0954] 462. The method of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0955] 463. The bacterium of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0956] 464. The bacterial population of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0957] 465. The isolated bacteria of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0958] 466. The non-intergeneric bacterial population of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0959] 467. The non-intergeneric bacterium of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 10% or more of the fixed nitrogen in the plant. [0960] 468. The method of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0961] 469. The method of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0962] 470. The bacterium of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0963] 471. The bacterial population of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0964] 472. The isolated bacteria of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0965] 473. The non-intergeneric bacterial population of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0966] 474. The non-intergeneric bacterium of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 15% or more of the fixed nitrogen in the plant. [0967] 475. The method of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0968] 476. The method of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0969] 477. The bacterium of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0970] 478. The bacterial population of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0971] 479. The isolated bacteria of any of the previous clauses, wherein the plurality of bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0972] 480. The non-intergeneric bacterial population of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0973] 481. The non-intergeneric bacterium of any of the previous clauses, wherein the plurality of non-intergeneric bacteria, in planta, produce 20% or more of the fixed nitrogen in the plant. [0974] 482. The method of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per gram of fresh weight of plant root tissue per hour. [0975] 483. The bacterium of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per gram of fresh weight of plant root tissue per hour. [0976] 484. The bacterial population of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per grain of fresh weight of plant root tissue per hour. [0977] 485. The isolated bacteria of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per gram of fresh weight of plant root tissue per hour. [0978] 486. The non-intergeneric bacterial population of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per gram of fresh weight of plant root tissue per hour. [0979] 487. The non-intergeneric bacterium of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-7 mmol N per gram of fresh weight of plant root tissue per hour. [0980] 488. The method of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-6 mmol N per gram of fresh weight of plant root tissue per hour. [0981] 489. The bacterium of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-6 mmol N per gram of fresh weight of plant root tissue per hour. [0982] 490. The bacterial population of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-6 mmol N per gram of fresh weight of plant root tissue per hour. [0983] 491. The isolated bacteria of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.-6 mmol N per gram of fresh weight of plant root tissue per hour. [0984] 492. The non-intergeneric bacterial population of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.16 mmol N per gram of fresh weight of plant root tissue per hour. [0985] 493. The non-intergeneric bacterium of any of the previous clauses, wherein the product of (i) the average colonization ability per unit of plant root tissue and (ii) produced fixed N per bacterial cell per hour, is at least about 2.0.times.10.sup.0.6 mmol N per grain of fresh weight of plant root tissue per hour. [0986] 494. The method of any of the previous clauses, wherein the plant has been remodeled or bred for efficient nitrogen use. [0987] 495. The bacterial composition of any of the previous clauses, wherein said at least one remodeled bacterial strain comprises at least one variation in a gene or intergenic region within 10,000 bp of a gene selected from the group consisting of: nifA, ntrB, ntrC, glnA, glnB, glnK, draT, amtB, glnD, glnE, nifJ, nifH, nifD, nifK, nifY, nifE, nifN, nifU, nifS, nifV, nifW, nifZ, nifM, nifF, nifB, and nifQ. [0988] 496. The method of any of the previous clauses, wherein said remodeled bacterium comprises a bacterial composition of any of the preceding clauses. [0989] 497. The method of any of the previous clauses, wherein said remodeled bacterium consists of a bacterial composition of any of the preceding clauses. [0990] 498. The method of any of the previous clauses, wherein the remodeled bacterial strain is a remodeled Gram-positive bacterial strain. [0991] 499. The method of any of the previous clauses, wherein the remodeled Gram-positive bacterial strain has an altered expression level of a regulator of a Nif cluster. [0992] 500. The method of any of the previous clauses, wherein the remodeled Gram-positive bacterial strain expresses a decreased amount of a negative regulator of a Nif cluster. [0993] 501. The method of any of the previous clauses, wherein the remodeled bacterial strain expresses a decreased amount of GlnR. [0994] 502. The method of any of the previous clauses, wherein the genome of the remodeled bacterial strain encodes a polypeptide with at least 75% identity to a sequence from the Zehr lab NifH database. [0995] 503. The method of any of the previous clauses, wherein the genome of the remodeled bacterial strain encodes a polypeptide with at least 85% identity to a sequence from the Zehr lab NifH database. [0996] 504. The method of any of the previous clauses, wherein the genome of the remodeled bacterial strain encodes a polypeptide with at least 75% identity to a sequence from the Buckley lab NifH database. [0997] 505. The method of any of the previous clauses, wherein the genome of the remodeled bacterial strain encodes a polypeptide with at least 85% identity to a sequence from the Buckley lab NifH database. [0998] 506. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes comprise at least one genetic variation introduced into said nitrogen fixation genetic regulatory network. [0999] 507. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes comprise at least one genetic variation introduced into said nitrogen assimilation genetic regulatory network. [1000] 508. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes comprise at least one genetic variation introduced into said nitrogen fixation genetic regulatory network and at least one genetic variation introduced into said nitrogen assimilation genetic regulatory network. [1001] 509. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes are applied into furrows in which seeds of said non-leguminous plant are planted. [1002] 510. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes are coated onto a seed of said non-leguminous plant. [1003] 511. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes colonize at least a root of said non-leguminous plant such that said remodeled non-intergeneric microbes are present in said non-leguminous plant in an amount of at least 10.sup.5 colony forming units per gram fresh weight of tissue. [1004] 512. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes are capable of fixing atmospheric nitrogen in non-nitrogen-limiting conditions. [1005] 513. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes, in planta, excrete nitrogen-containing products of nitrogen fixation. 514. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes, in planta, produce at least 1% of fixed nitrogen in said non-leguminous plant. [1006] 515. The method of any of the previous clauses, wherein said fixed nitrogen in said non-leguminous plant produced by said remodeled non-intergeneric microbes is measured by dilution of 15N in crops grown in fields treated with fertilizer containing 1.2% 15N. [1007] 516. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes, in planta, produce 5% or more of the fixed nitrogen in said non-leguminous plant. [1008] 517. The method of any of the previous clauses, wherein said non-intergeneric microbes are remodeled using at least one type of engineering selected from the group consisting of directed mutagenesis, random mutagenesis, and directed evolution. [1009] 518. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes, in planta, excrete nitrogen-containing products of nitrogen fixation. [1010] 519. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes are applied into furrows in which seeds of said corn plant are planted. [1011] 520. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes are coated onto a seed of said corn plant. [1012] 521. The method of any of the previous clauses, wherein said remodeled non-intergeneric microbes comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network.
[1013] 522. The method of any of the previous clauses, wherein said non-intergeneric microbes comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1014] 523. The method of any of the previous clauses, wherein said genetically engineered non-intergeneric microbes comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1015] 524. The method of any of the previous clauses, wherein said remodeled non-intergeneric bacteria comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1016] 525. The method of any of the previous clauses, wherein said non-intergeneric bacteria comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1017] 526. The method of any of the previous clauses, wherein said genetically engineered non-intergeneric bacteria comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1018] 527. The bacterium of any of the previous clauses, wherein said bacterium comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1019] 528. The bacterial population of any of the previous clauses, wherein bacteria within said bacterial population comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1020] 529. The isolated bacteria of any of the previous clauses, wherein said isolated bacteria comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1021] 530. The non-intergeneric bacterial population of any of the previous clauses, wherein non-intergeneric bacteria within said non-intergeneric bacterial population comprise at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1022] 531. The non-intergeneric bacterium of any of the previous clauses, wherein said non-intergeneric bacterium comprises at least one genetic variation introduced into a nitrogen fixation genetic regulatory network and at least one genetic variation introduced into a nitrogen assimilation gentic regulatory network. [1023] 532. A composition comprising any one or more bacteria of any of the previous clauses.
[1024] While preferred embodiments of the present invention have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the invention. It should be understood that various alternatives to the embodiments of the invention described herein may be employed in practicing the invention. It is intended that the following claims define the scope of the invention and that methods and structures within the scope of these claims and their equivalents be covered thereby.
Sequence CWU 0 SQTB SEQUENCE LISTING The patent application
contains a lengthy "Sequence Listing" section. A copy of the
"Sequence Listing" is available in electronic form from the USPTO
web site
(http://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20190039964A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
0 SQTB SEQUENCE LISTING The patent application contains a lengthy
"Sequence Listing" section. A copy of the "Sequence Listing" is
available in electronic form from the USPTO web site
(http://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20190039964A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
References
-
ebi.ac.uk/Tools/psa/emboss_needle/nucleotide.html
-
ebi.ac.uk/Tools/psa/emboss_water/nucleotide.html
-
cnrc.agron.iastate.edu/TheMRTNisthenitrogenratewheretheeconomicnetreturntonitrogenapplicationismaximized
-
ncbi.nlm.nih.gov/tools/primer-blast
-
wwwzehr.pmc.aese.edu/nifH_Database_Public
-
css.cornell.edu/faculty/buckley/nifh.htm
-
wwwzehr.pmc.ucsc.edu/nifH_Database_Public
-
pioneer.com/home/site/us/agronomy/librarykorn-seeding-rate-considerations
-
seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20190039964A1
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
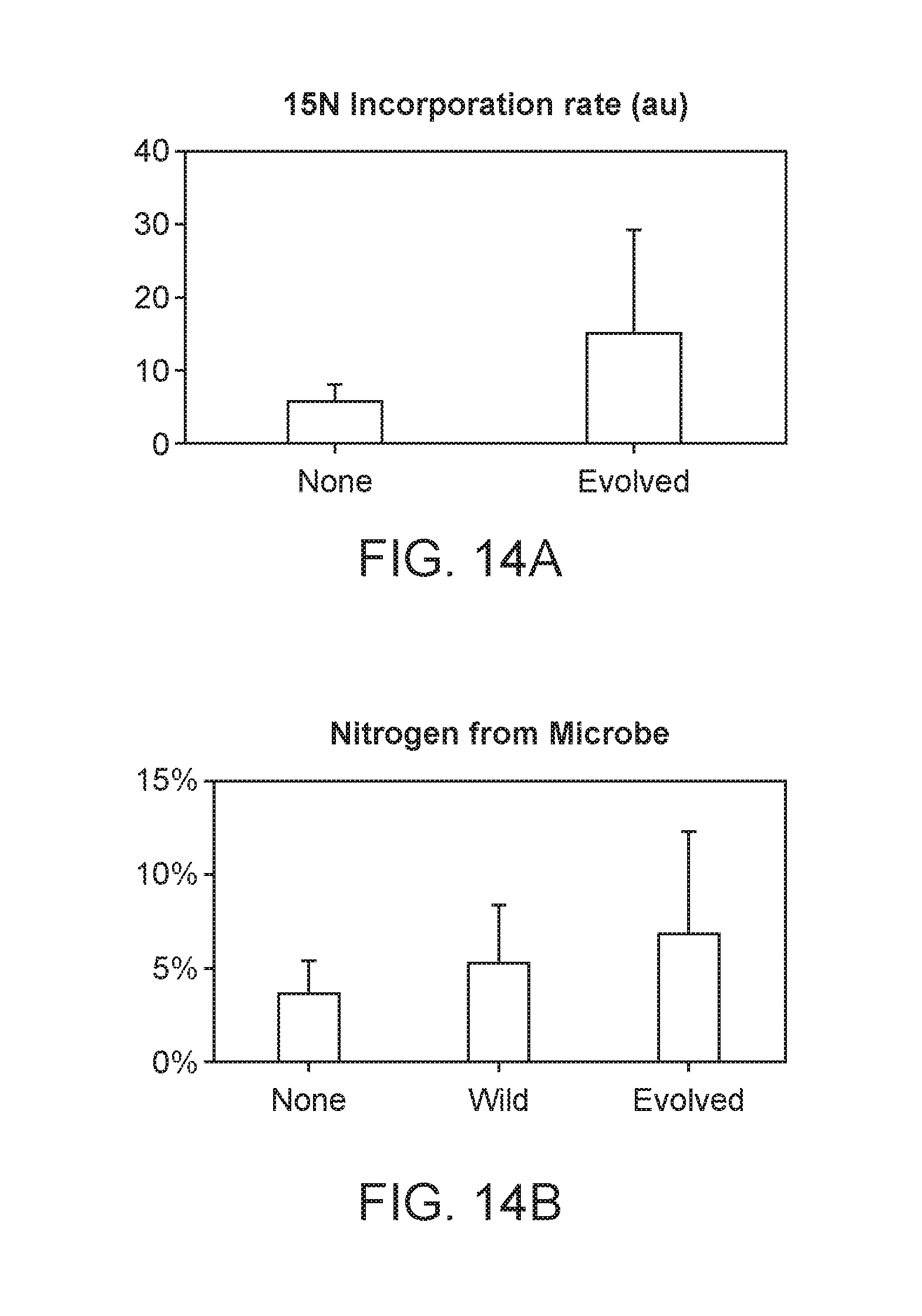

D00014
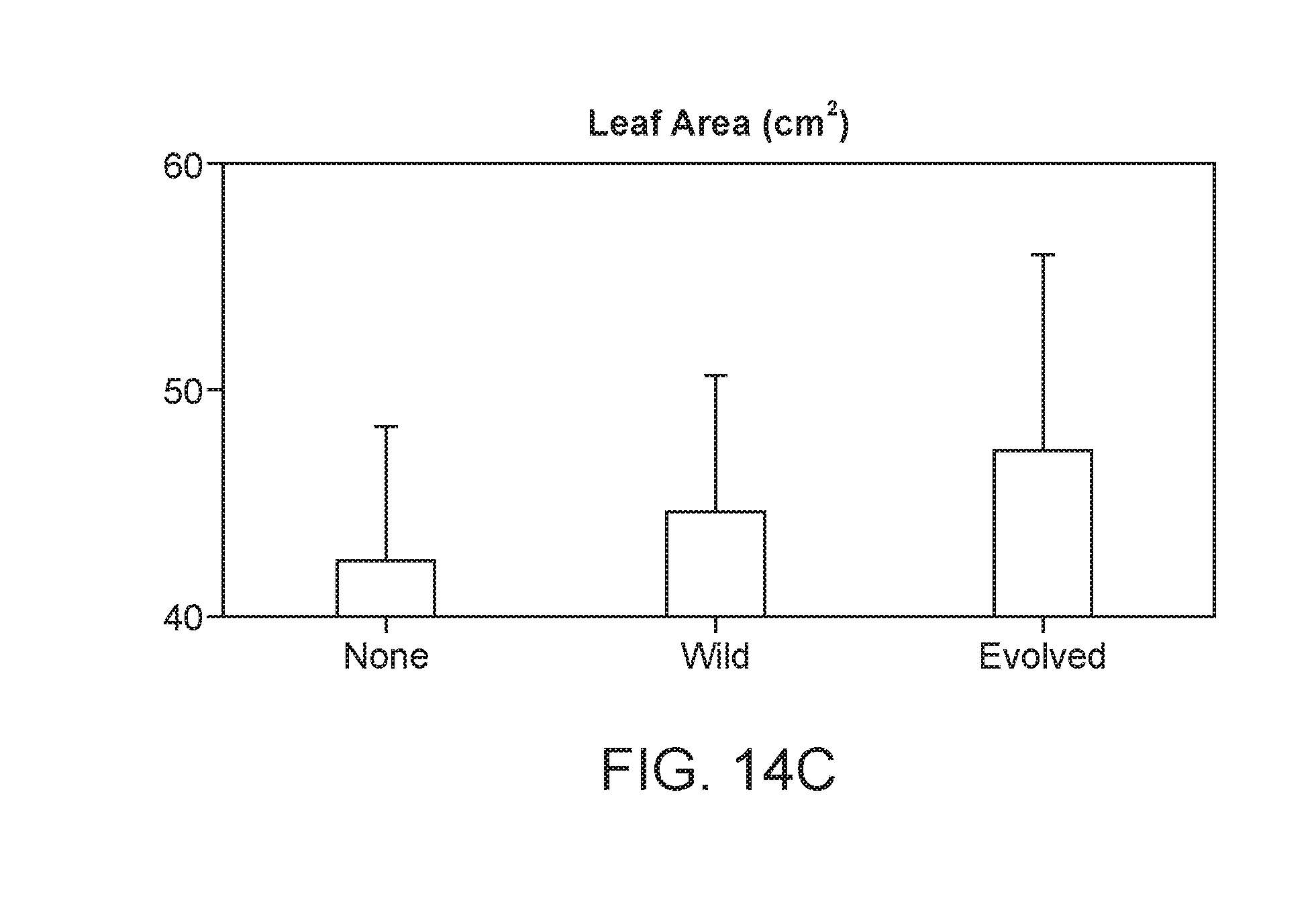
D00015
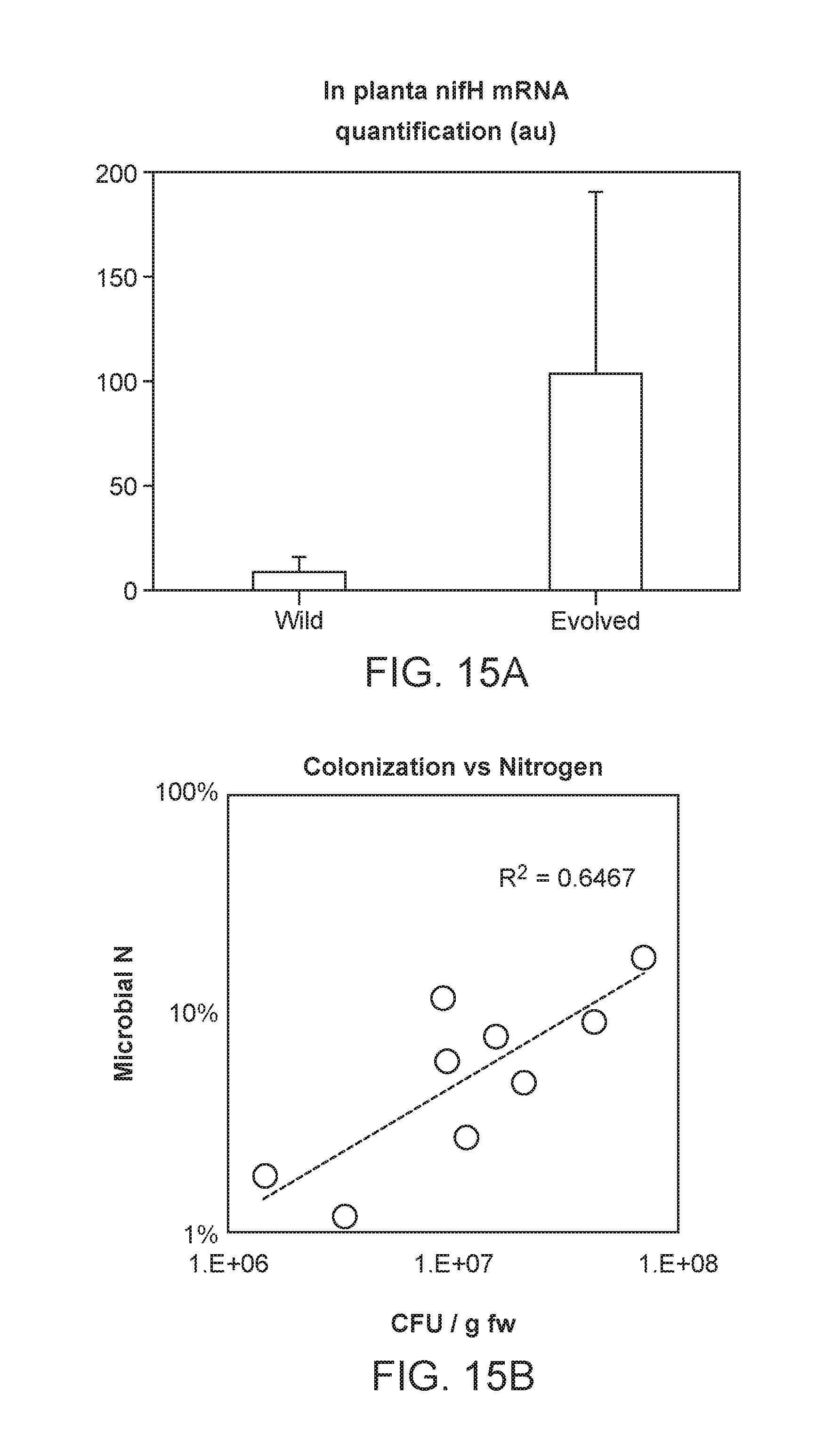
D00016

D00017

D00018
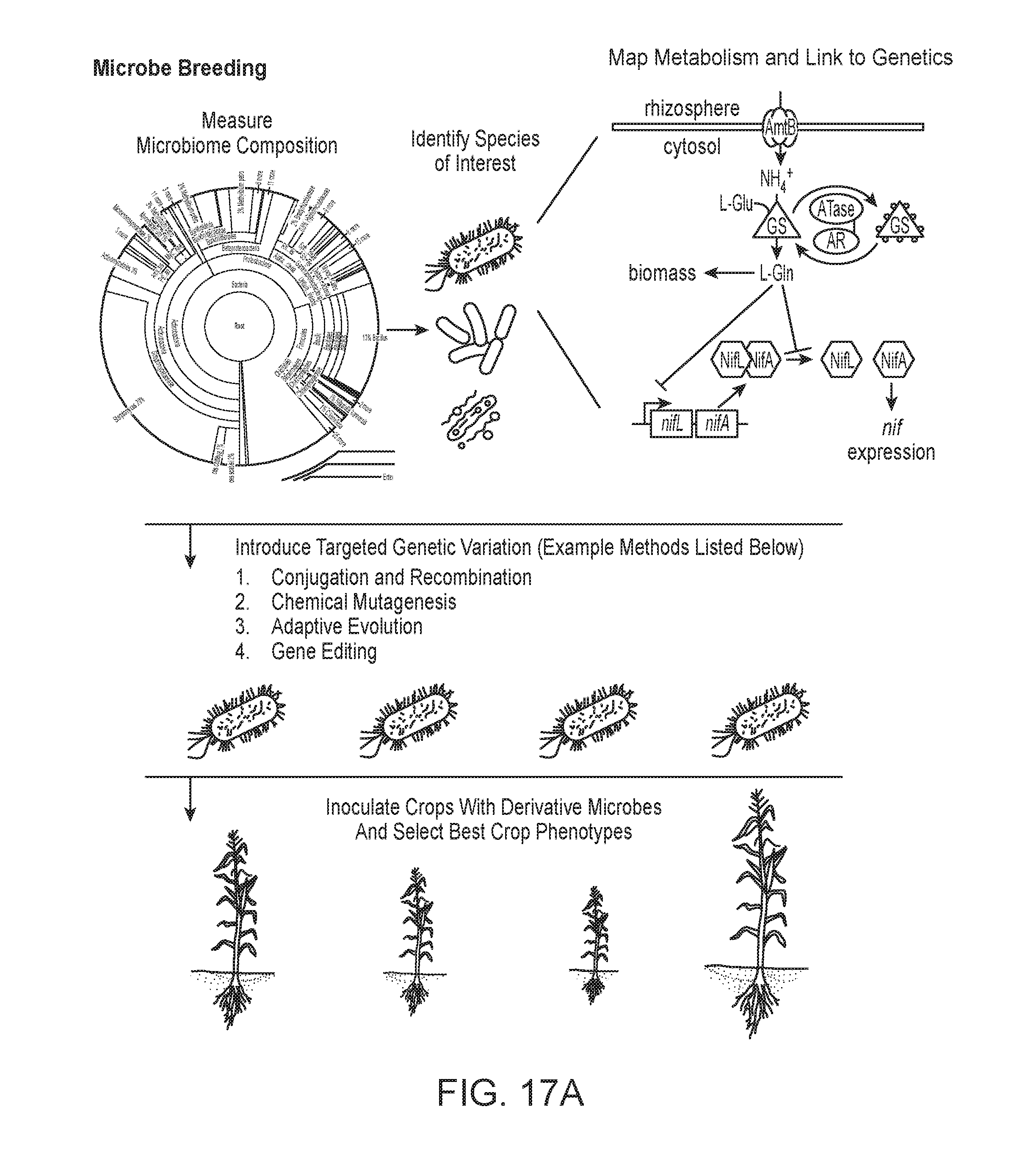
D00019

D00020

D00021
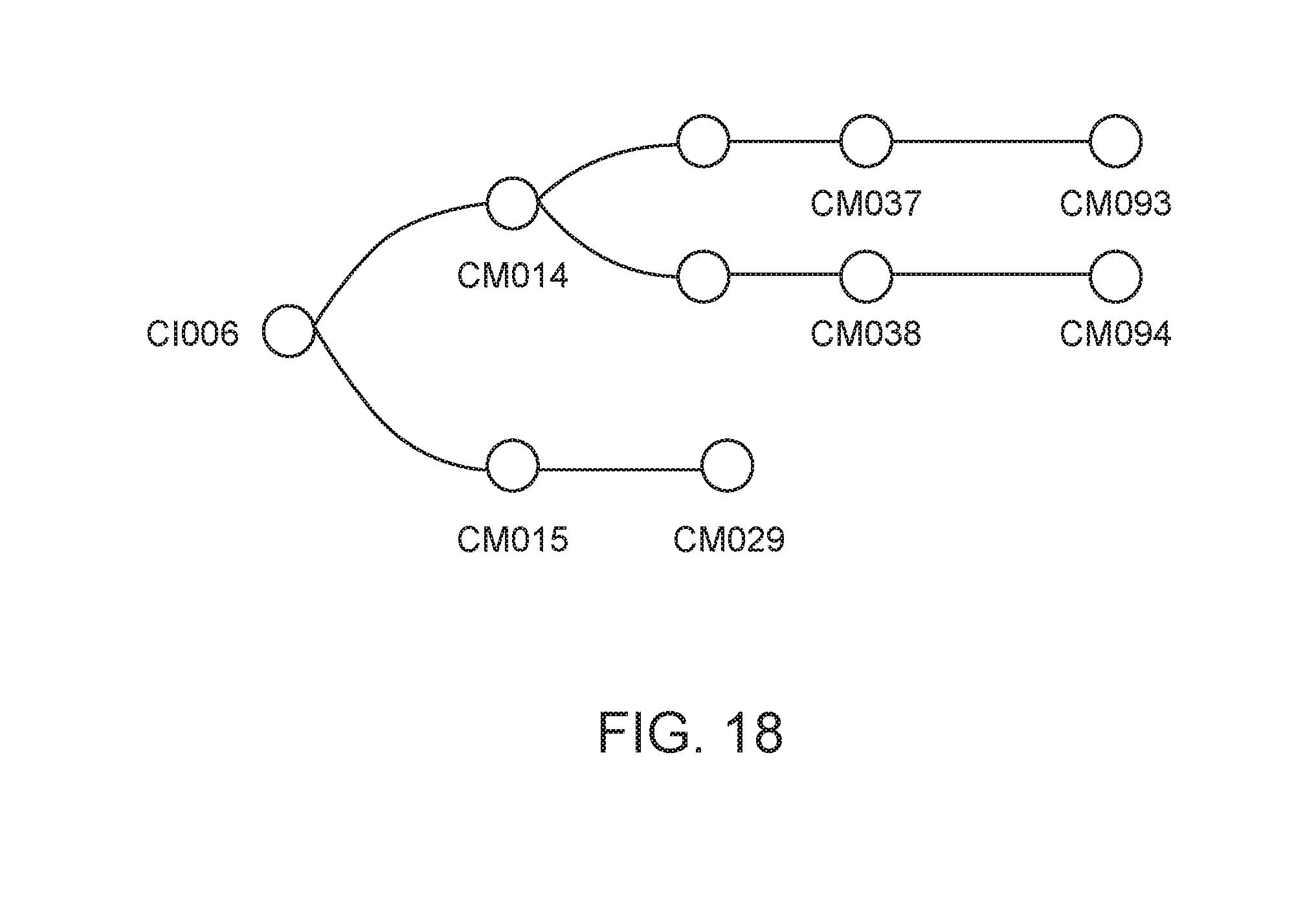
D00022
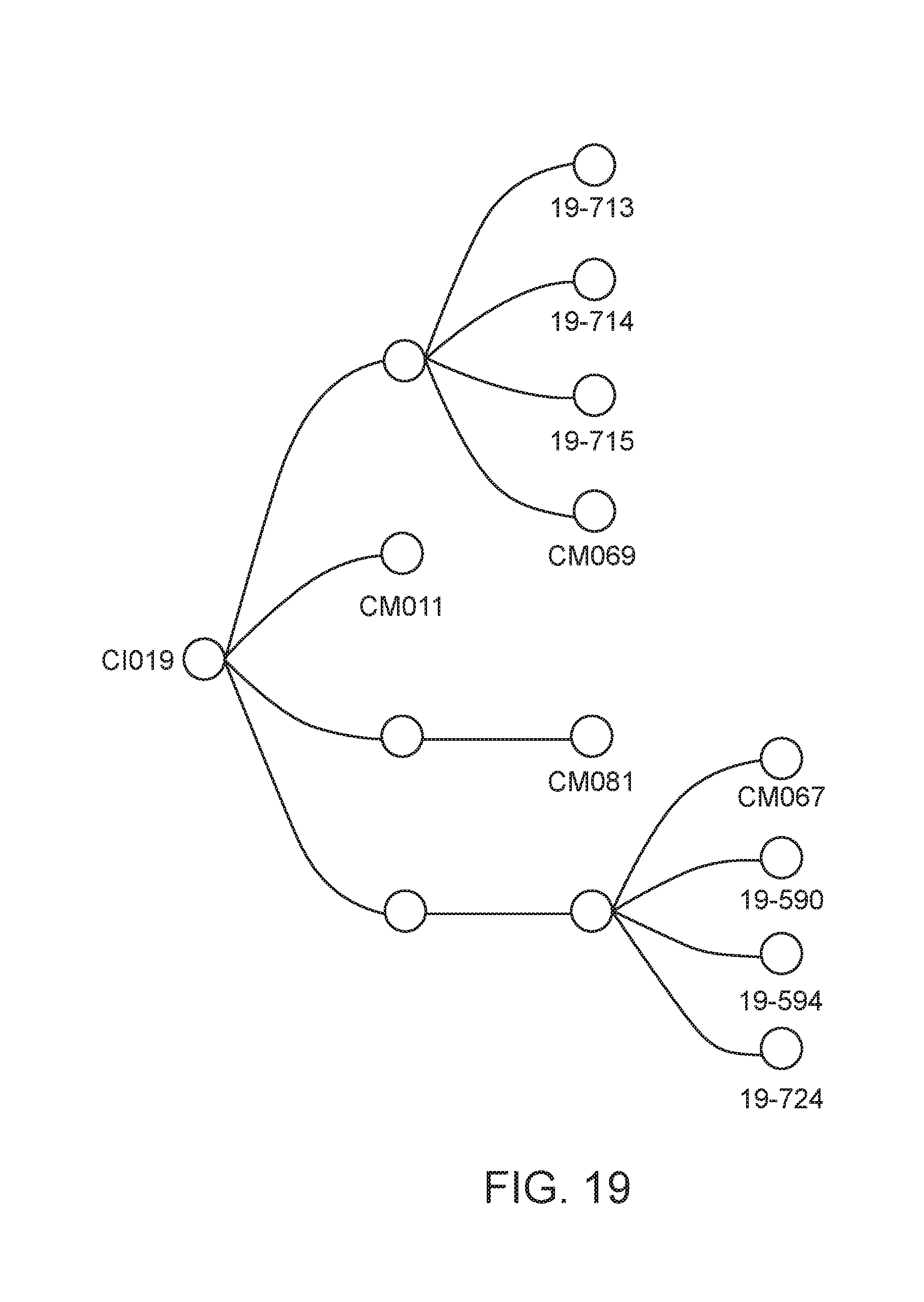
D00023
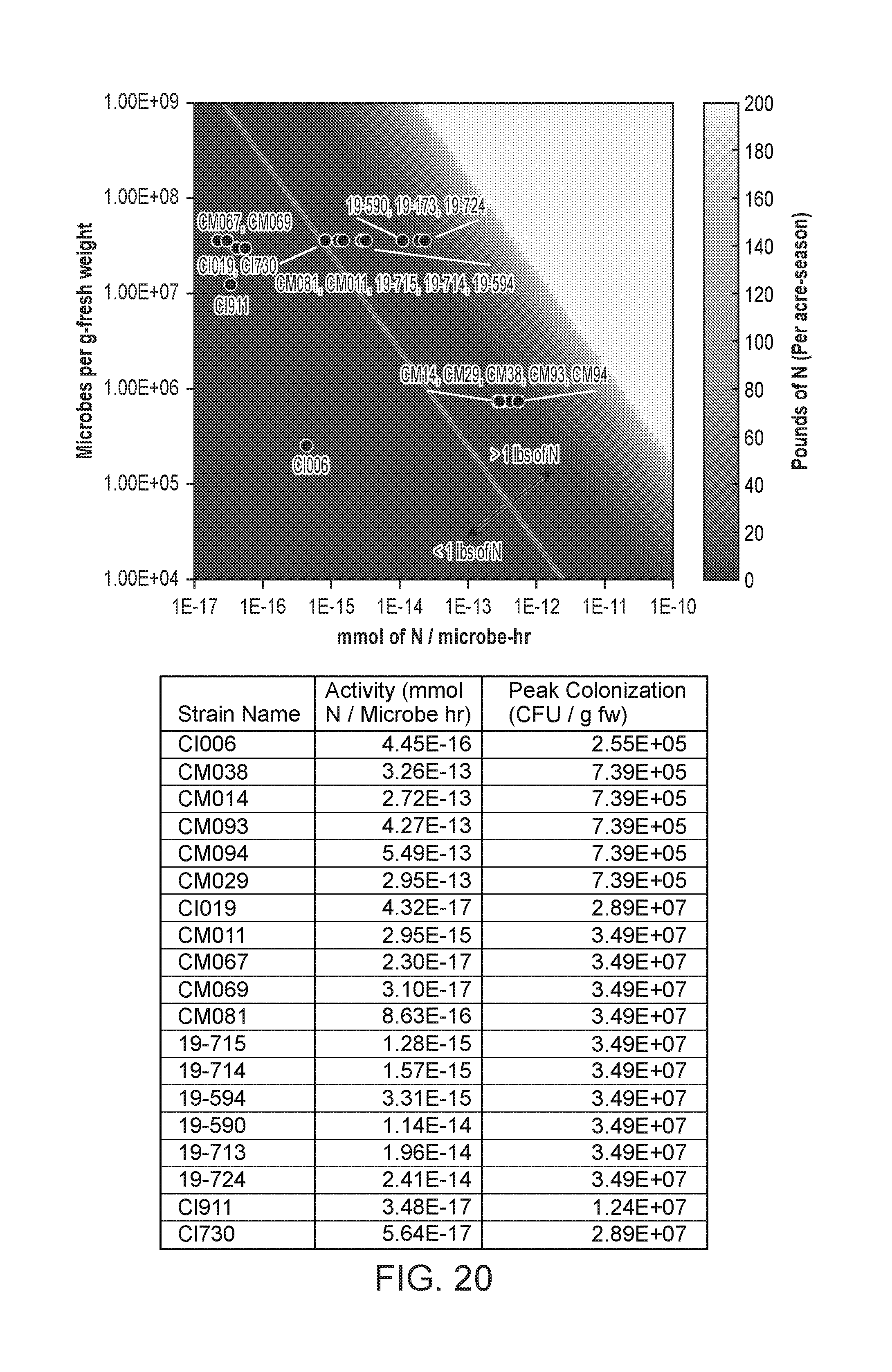
D00024
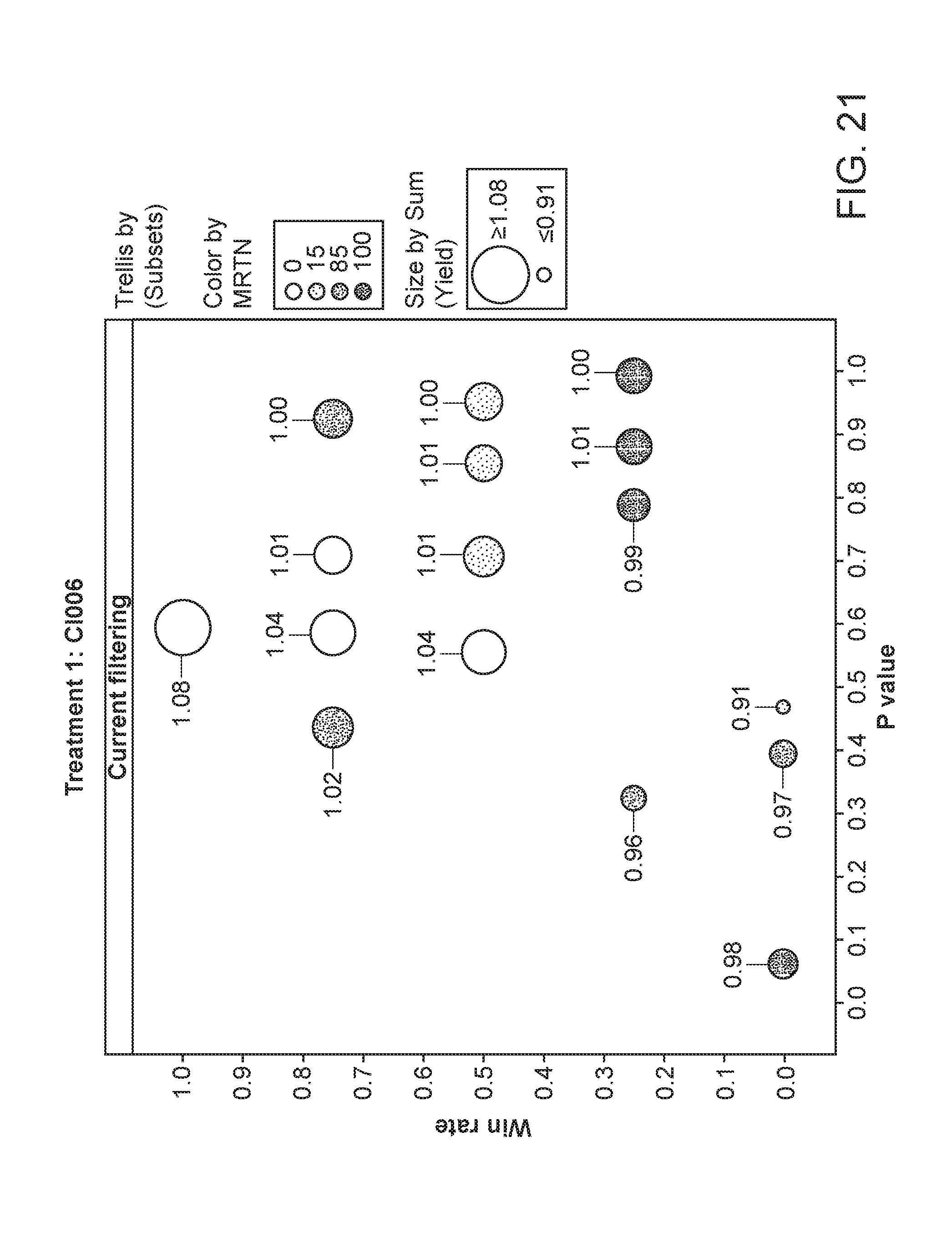
D00025
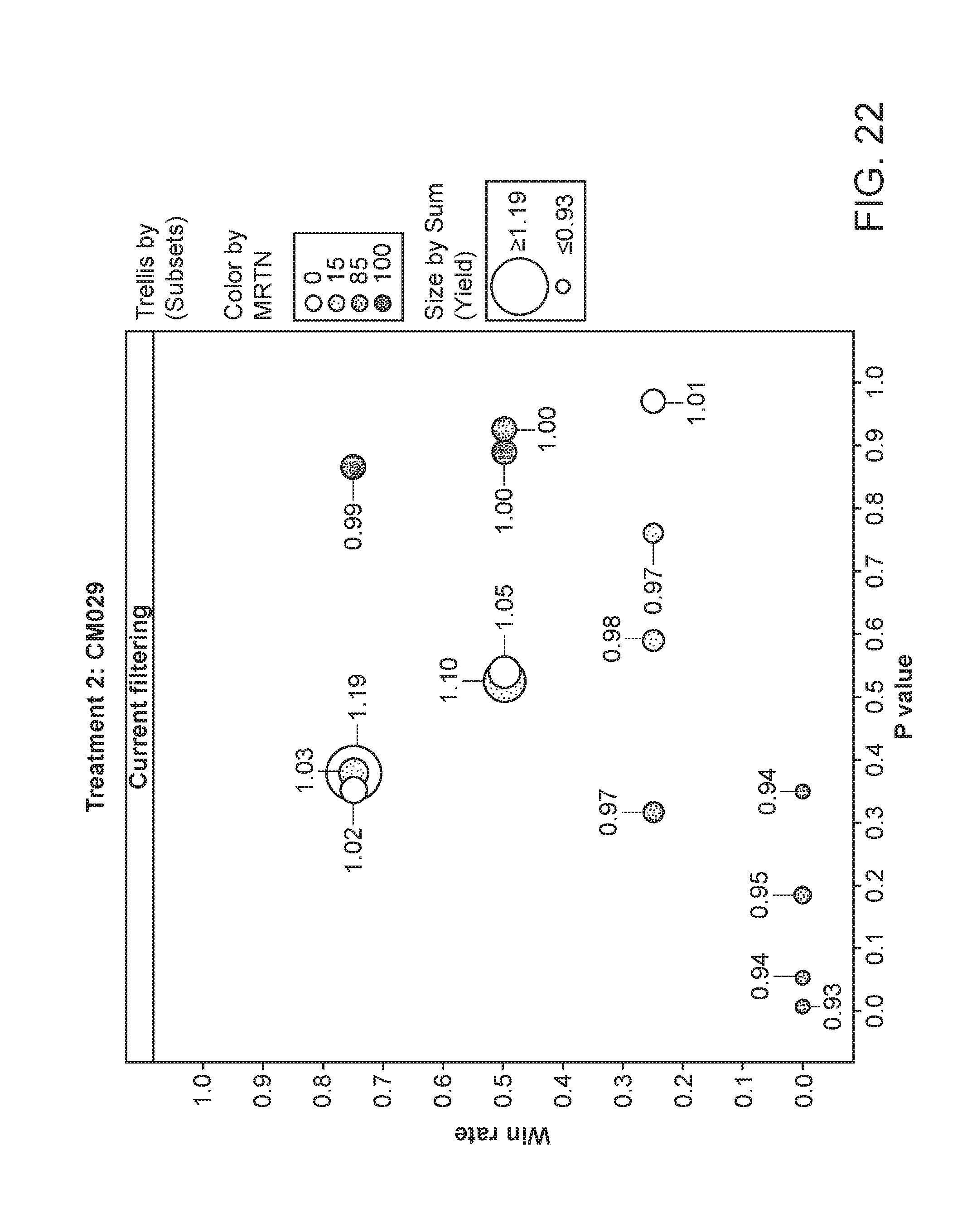
D00026
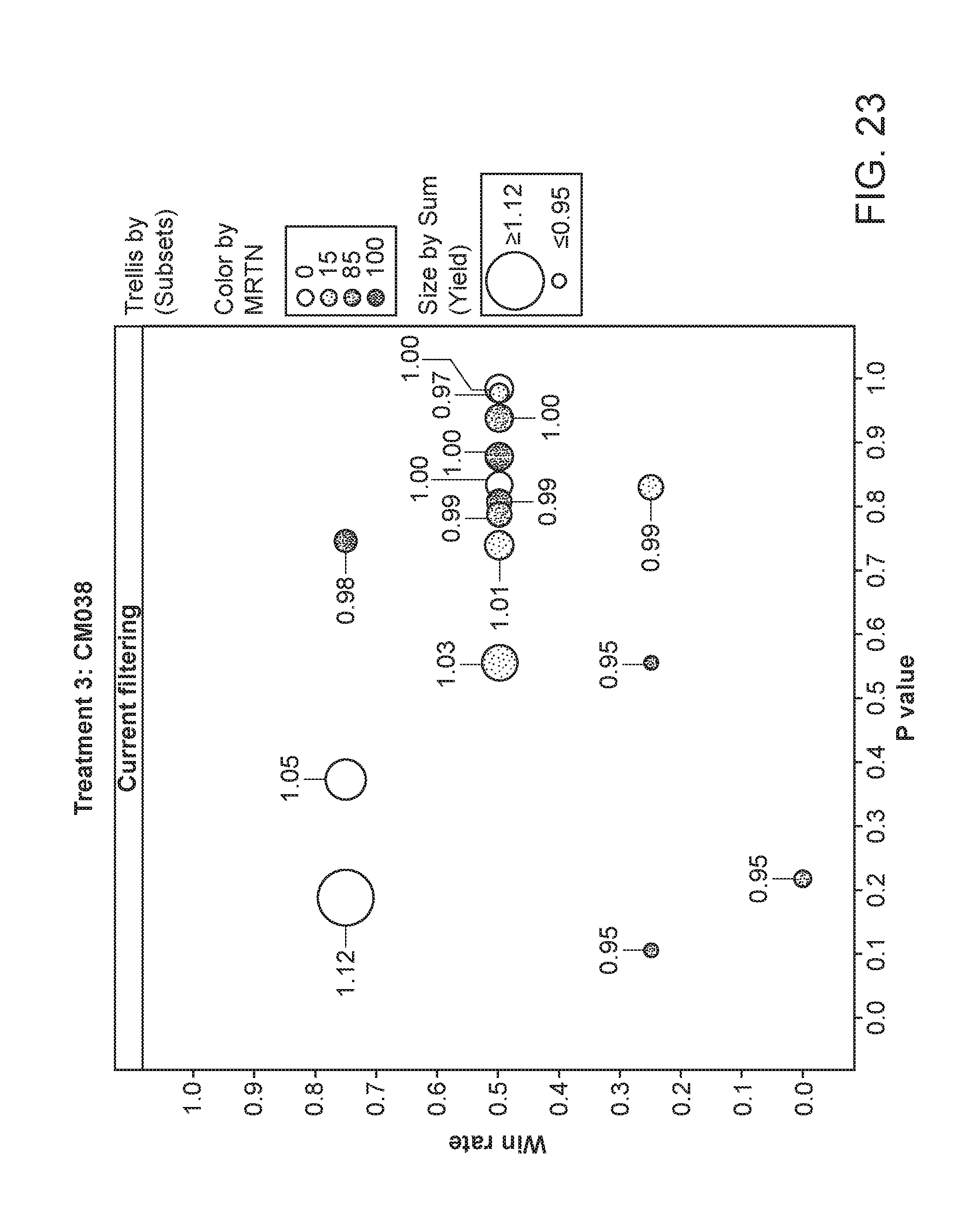
D00027
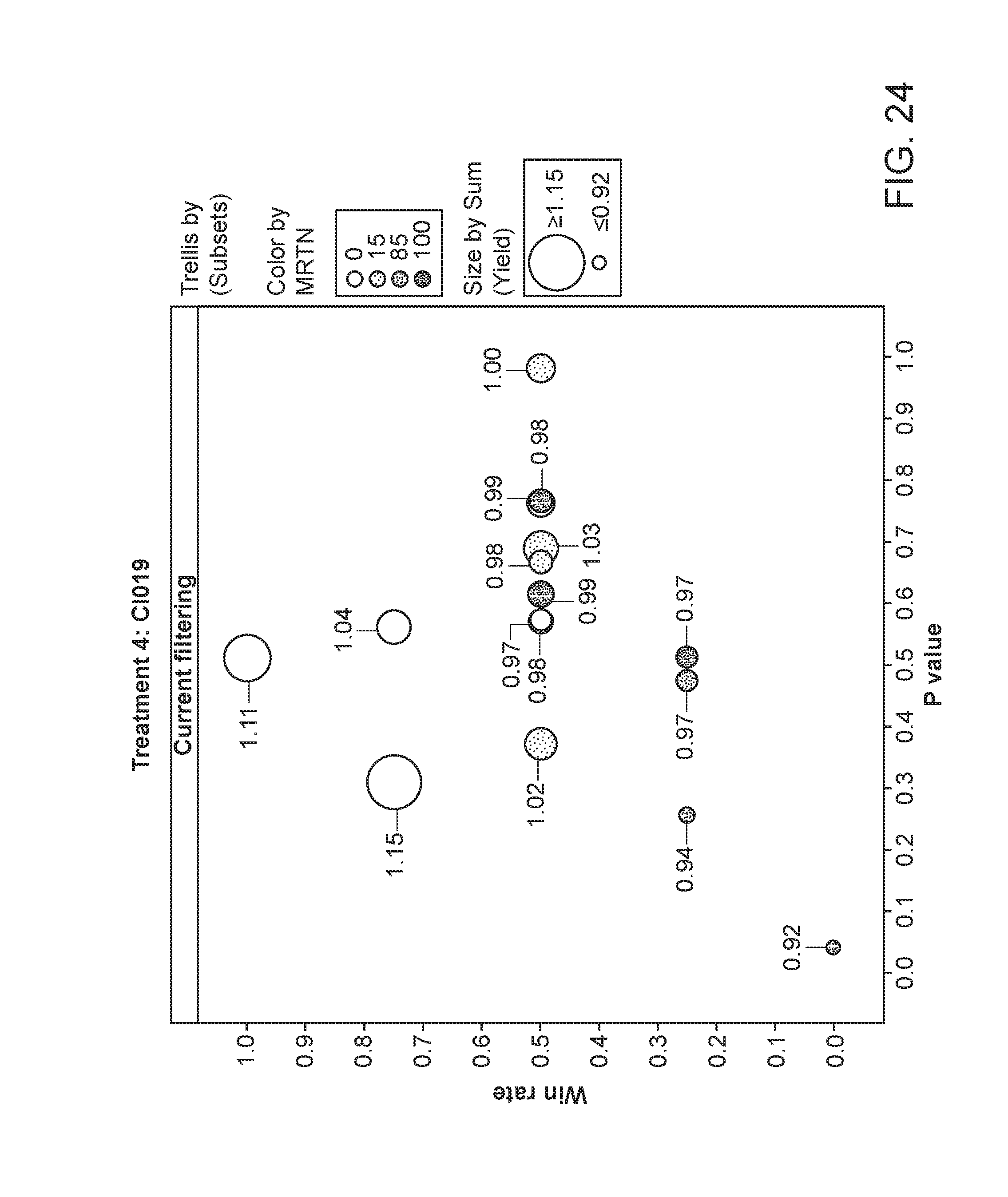
D00028
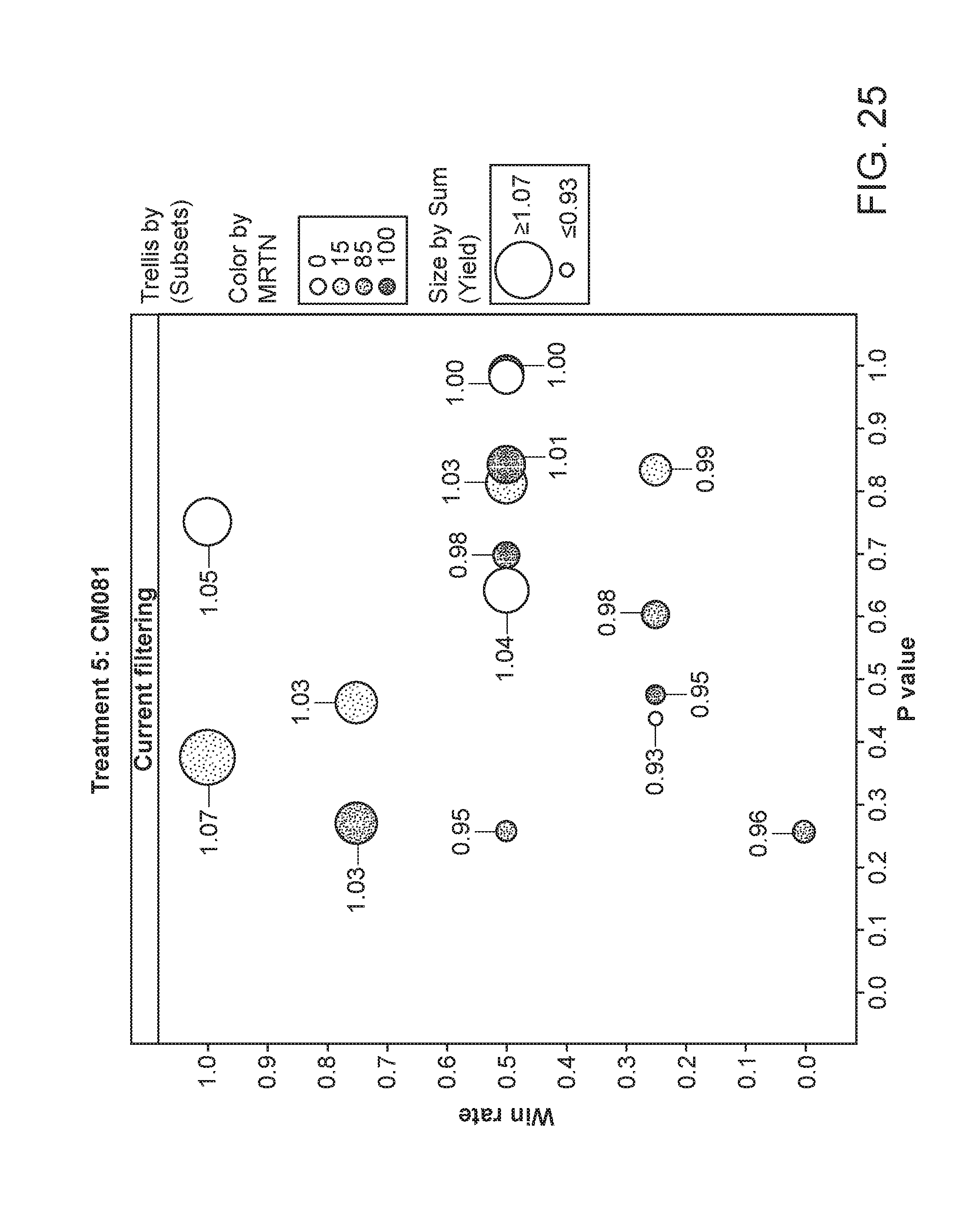
D00029
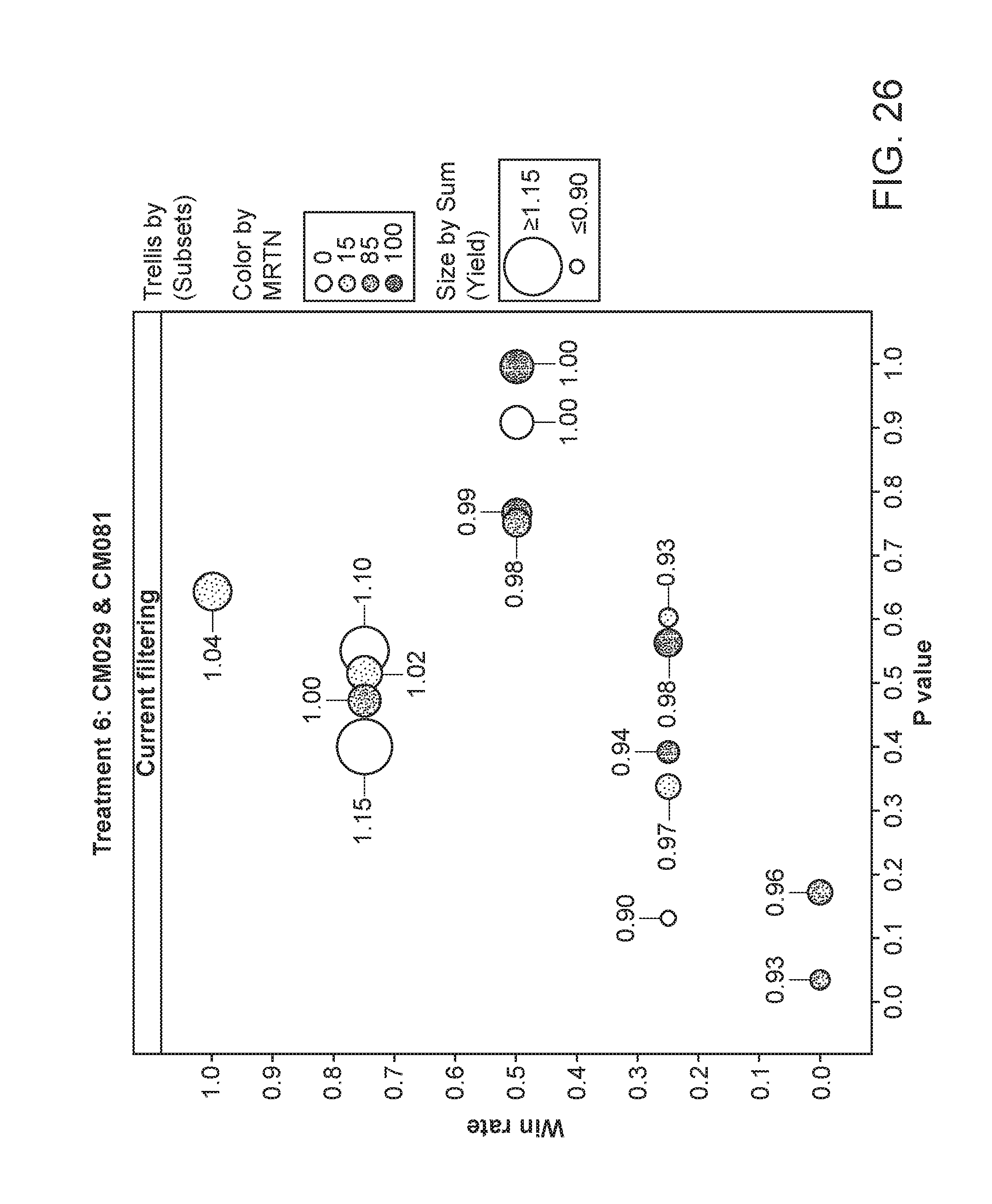
D00030
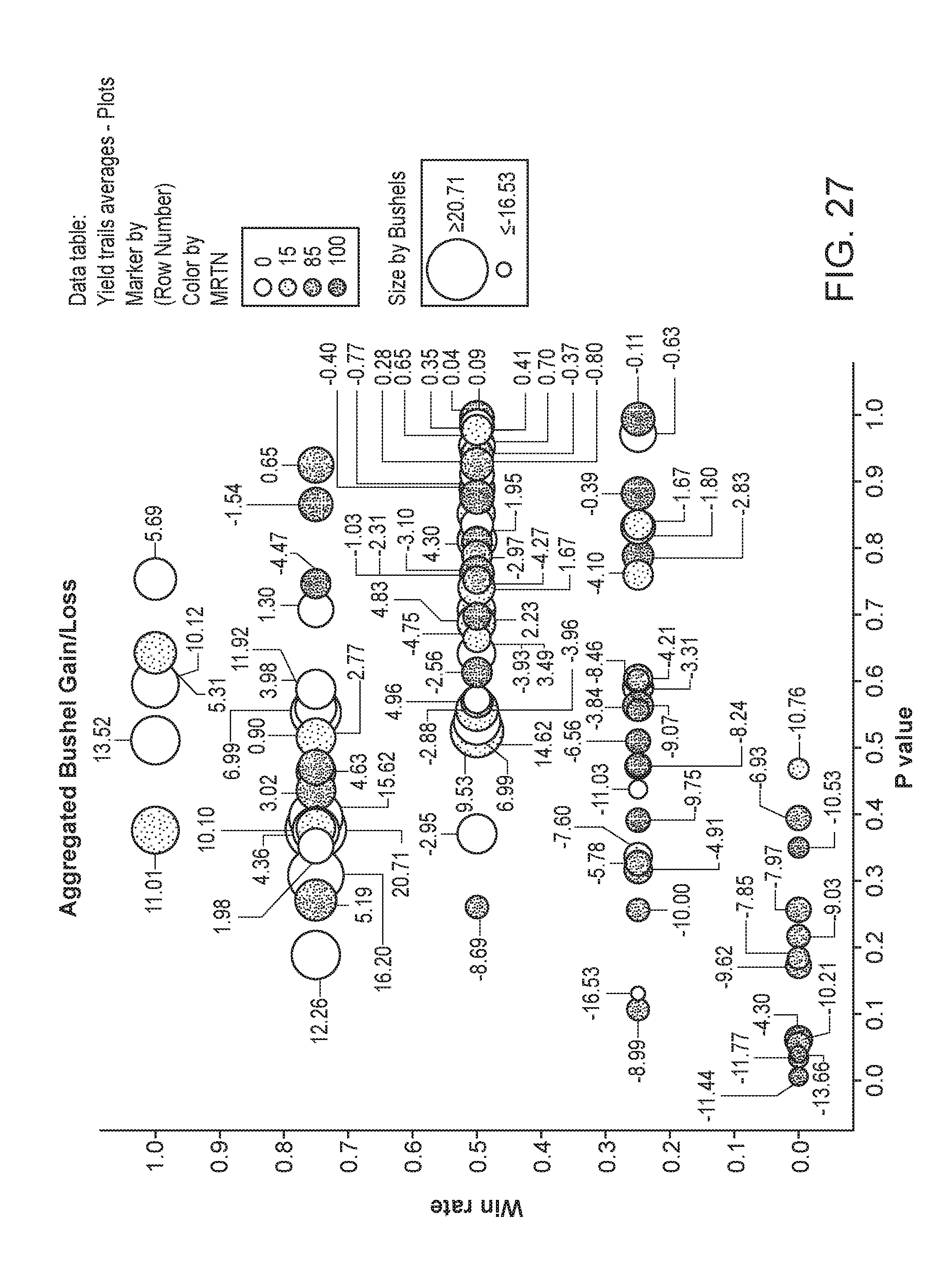
D00031
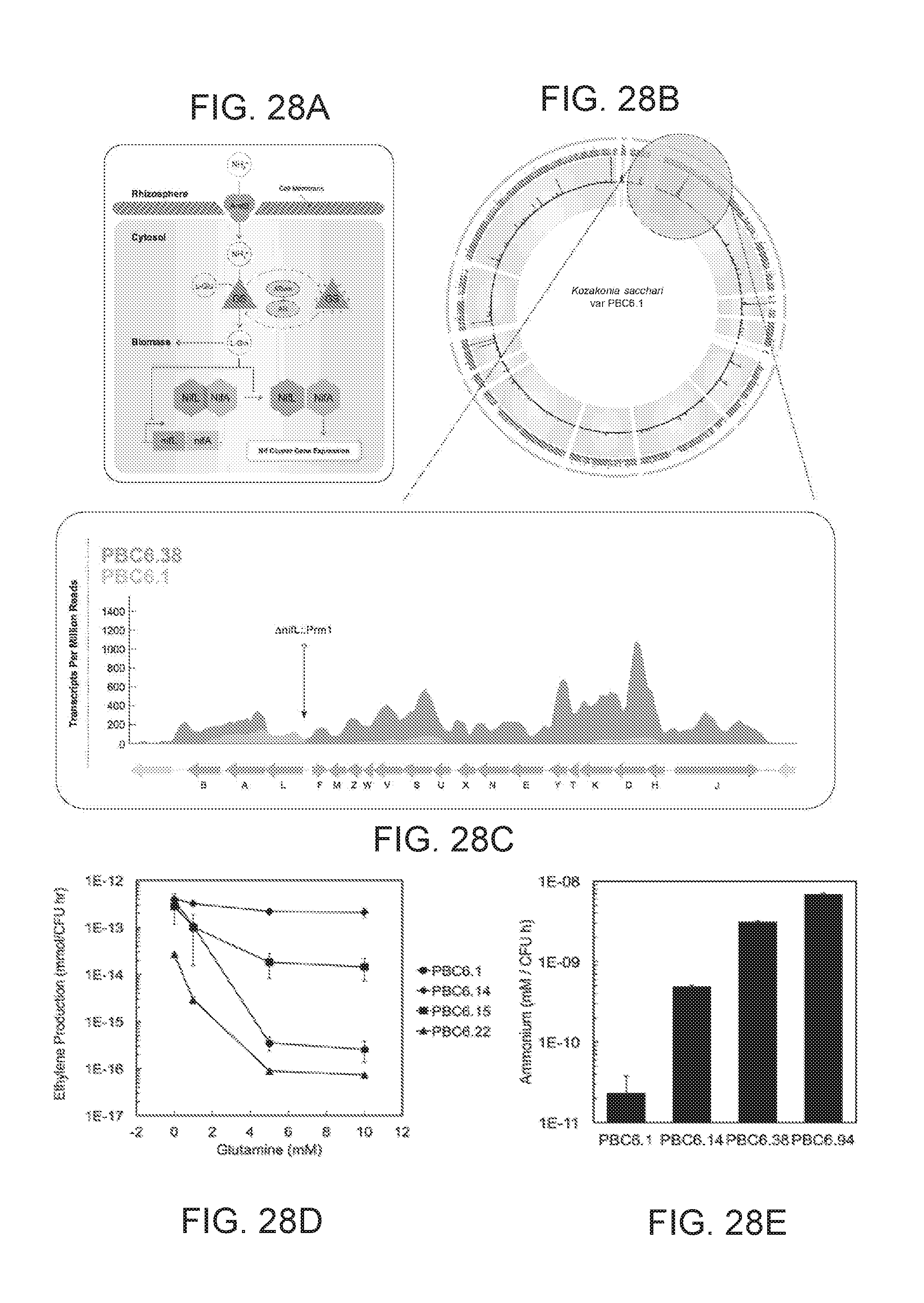
D00032

D00033

D00034

D00035
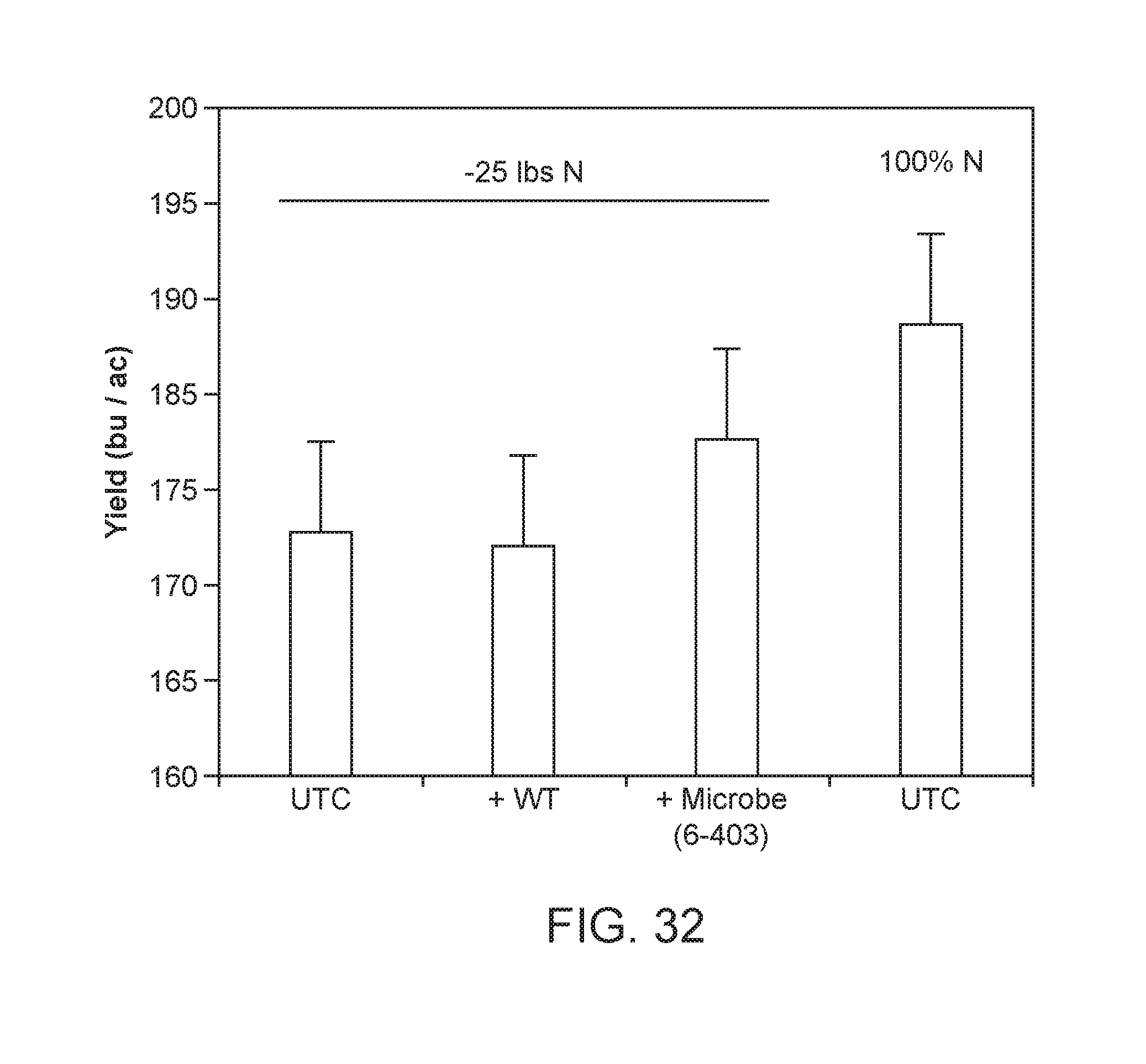
D00036
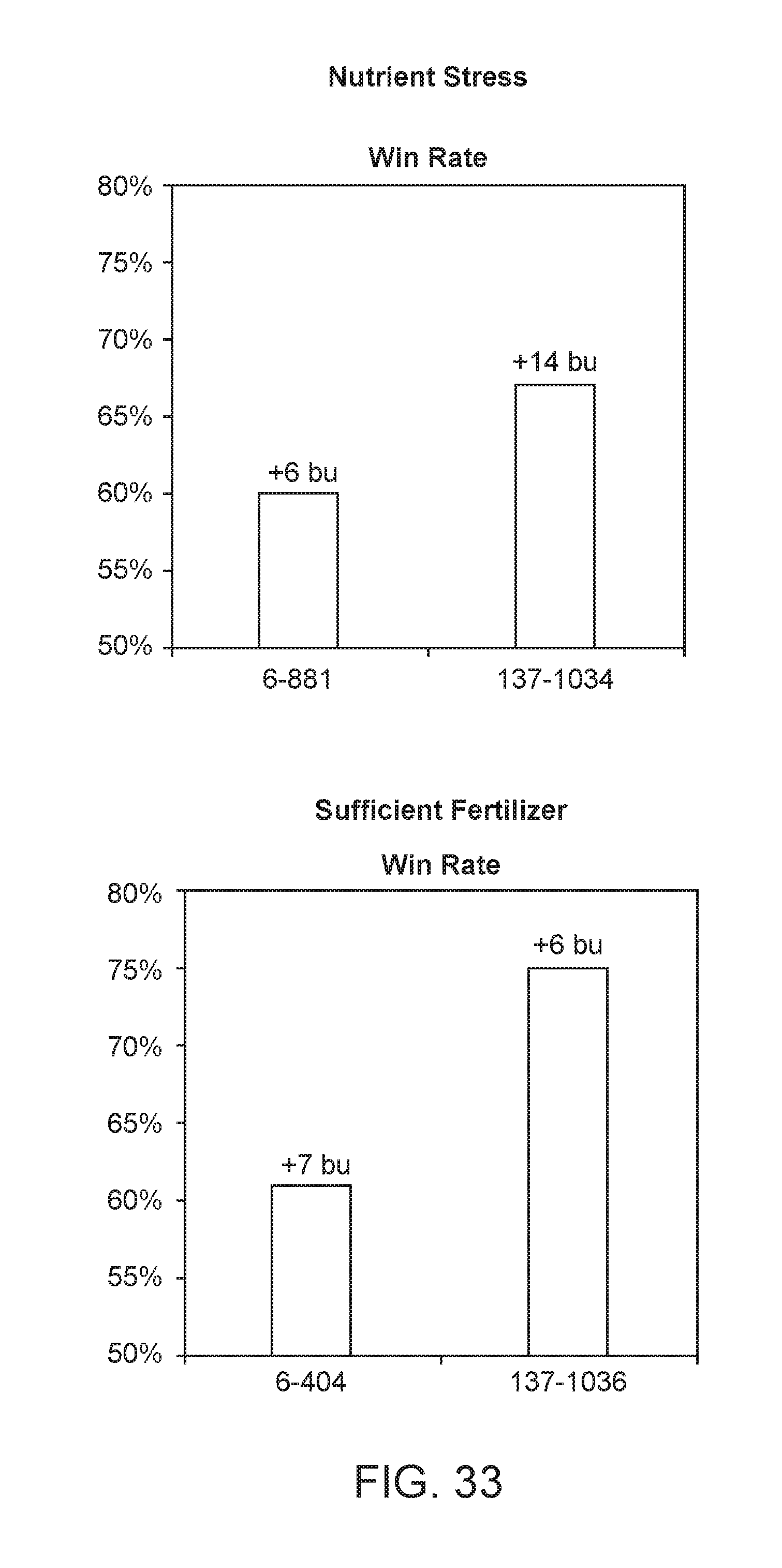
D00037
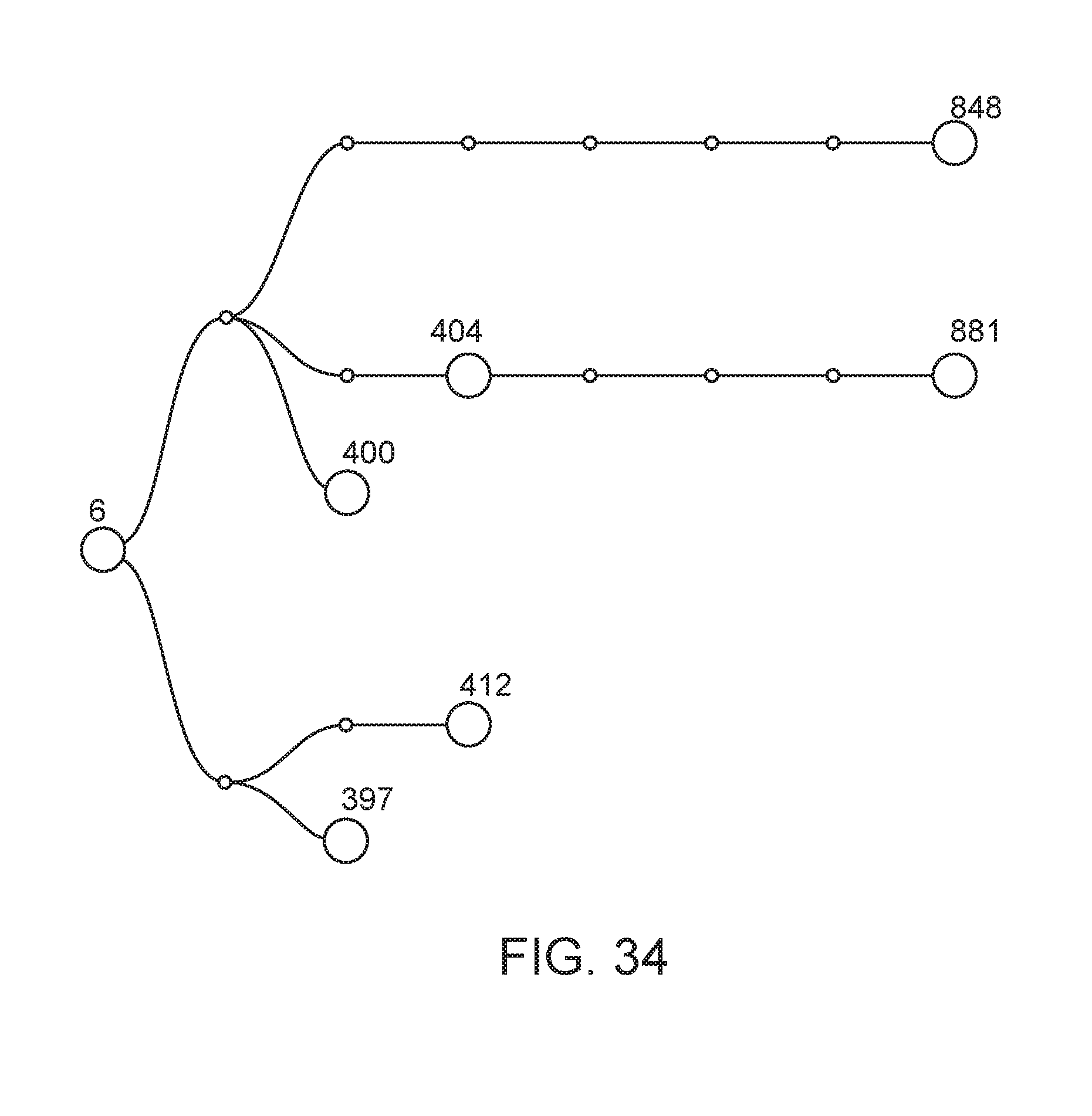
D00038
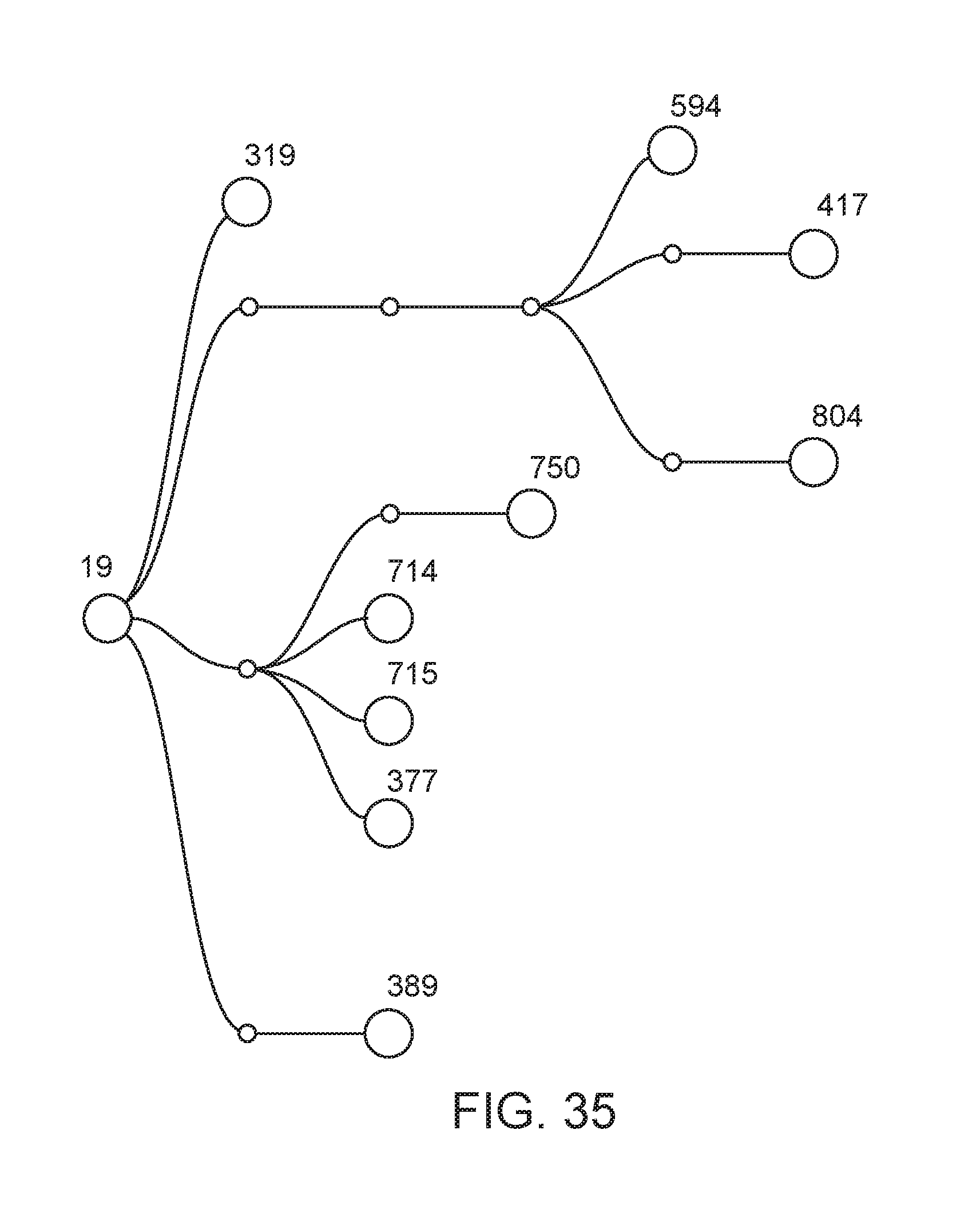
D00039
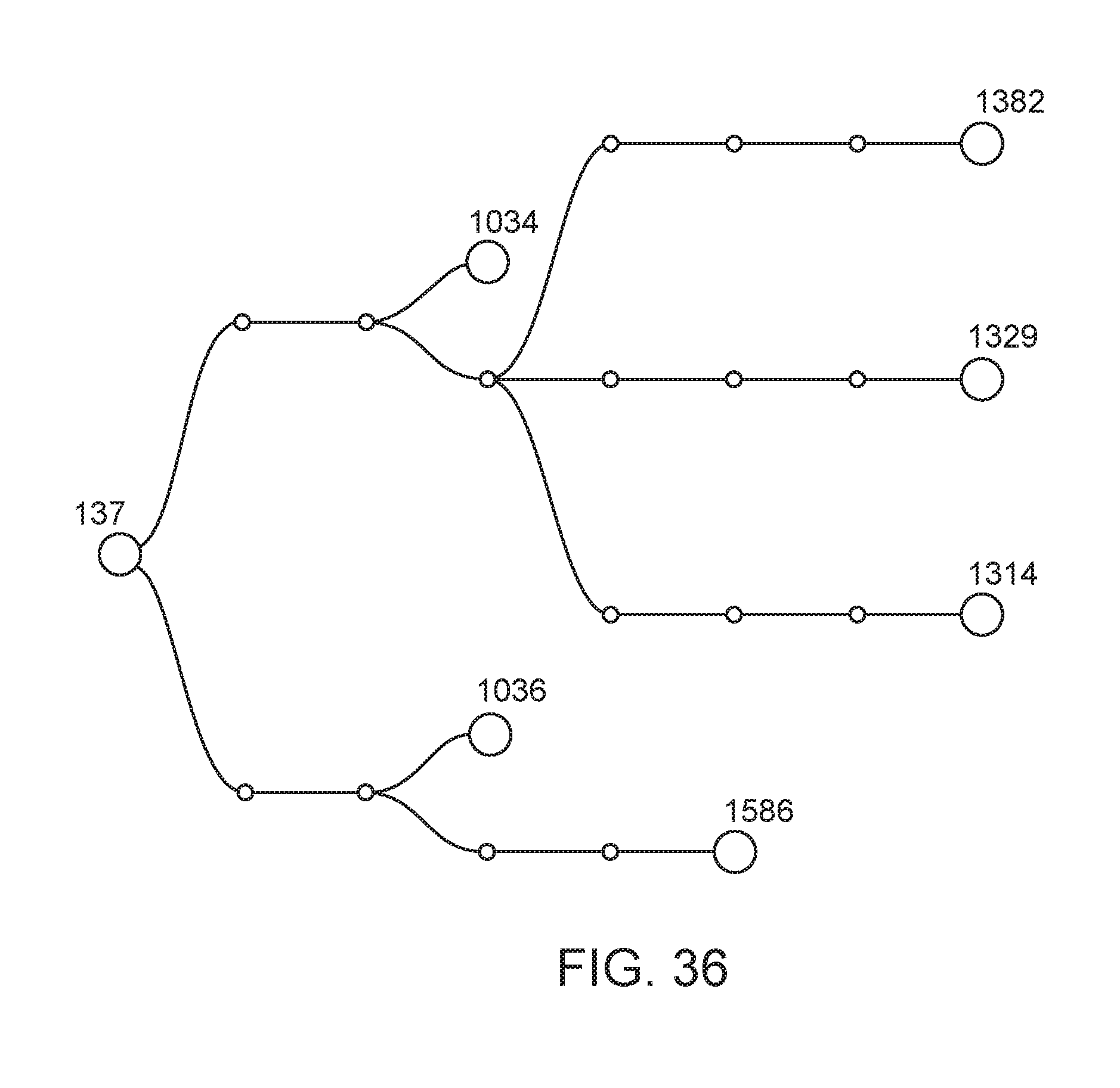
D00040
D00041

D00042

D00043

D00044

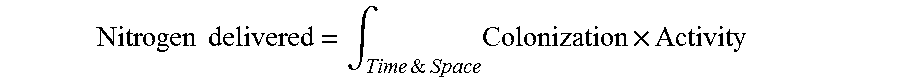
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.