Leghemoglobin In Soybean
CHO; HYEON-JE ; et al.
U.S. patent application number 17/451913 was filed with the patent office on 2022-04-28 for leghemoglobin in soybean. This patent application is currently assigned to PIONEER HI-BRED INTERNATIONAL, INC.. The applicant listed for this patent is PIONEER HI-BRED INTERNATIONAL, INC.. Invention is credited to HYEON-JE CHO, JOHN D EVERARD, ANTHONY J KINNEY, ZHAN-BIN LIU, KNUT MEYER, THOMAS G PATTERSON, KEVIN G RIPP, BO SHEN.
| Application Number | 20220127631 17/451913 |
| Document ID | / |
| Family ID | 1000005970205 |
| Filed Date | 2022-04-28 |









View All Diagrams
| United States Patent Application | 20220127631 |
| Kind Code | A1 |
| CHO; HYEON-JE ; et al. | April 28, 2022 |
LEGHEMOGLOBIN IN SOYBEAN
Abstract
Soybean plants producing soybean seeds comprising leghemoglobin are produced by modifying the genome of the soybean plant. Soybean plants, soybean seeds and soy protein compositions comprising leghemoglobin are provided. Soybean plants, soybean seeds and soy protein compositions comprising leghemoglobin and additionally one or more of high oleic acid, low linolenic acid, high protein, low stachyose, low raffinose and low protease inhibitors are provided. Protein compositions comprising leghemoglobin, such as soy isolates and concentrates can be made from the soybean seeds. Additionally, methods for generating and using plants, seeds and protein compositions comprising leghemoglobin are disclosed.
| Inventors: | CHO; HYEON-JE; (ANKENY, IA) ; EVERARD; JOHN D; (GRIMES, IA) ; KINNEY; ANTHONY J; (WILMINGTON, DE) ; LIU; ZHAN-BIN; (CLIVE, IA) ; MEYER; KNUT; (WILMINGTON, DE) ; PATTERSON; THOMAS G; (WESTFIELD, IN) ; RIPP; KEVIN G; (DES MOINES, IA) ; SHEN; BO; (JOHNSTON, IA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | PIONEER HI-BRED INTERNATIONAL,
INC. JOHNSTON IA |
||||||||||
| Family ID: | 1000005970205 | ||||||||||
| Appl. No.: | 17/451913 | ||||||||||
| Filed: | October 22, 2021 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 63106519 | Oct 28, 2020 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 14/415 20130101; C12N 15/8251 20130101 |
| International Class: | C12N 15/82 20060101 C12N015/82; C07K 14/415 20060101 C07K014/415 |
Claims
1. A soybean seed comprising a leghemoglobin protein in an amount of at least 0.5% of total protein in the soybean seed, wherein the soybean seed does not contain a recombinant construct comprising a leghemoglobin coding sequence integrated into its genome.
2. The soybean seed of claim 1, wherein the soybean seed genome has been modified to introduce an insertion, deletion or substitution into a native leghemoglobin gene.
3. The soybean seed of claim 1, wherein the soybean seed genome has been modified to introduce an insertion, and the insertion comprises a targeting sequence operably linked to a leghemoglobin coding sequence of the leghemoglobin gene, the targeting sequence targeting the leghemoglobin to an intracellular compartment.
4. The soybean seed of claim 5, wherein the targeting sequence comprises a polynucleotide encoding a polypeptide having at least 95% identity to SEQ ID NO: 32.
5. The soybean seed of claim 1, wherein the genome of the soybean seed has been modified to replace all or part of a coding sequence of a seed storage protein with a leghemoglobin coding sequence.
6. The soybean seed of claim 1, wherein the soybean seed further comprises a characteristic selected from the group consisting of: (i) an oleic acid content of at least 50% of the total seed fatty acids; (ii) a linolenic acid content of less than 3% of the total seed fatty acids; (iii) a protein content of at least 37% of the total weight of the soybean measured at or adjusted to 13% moisture, and (iv) any combination thereof.
7. A plant which produces the soybean seed of claim 1.
8. A soybean seed comprising a genomic modification, the genomic modification comprising an insertion of a leghemoglobin coding sequence into a native seed storage protein gene such that the leghemoglobin coding sequence replaces all or part of the native storage protein gene coding sequence, and wherein the leghemoglobin protein is expressed in the soybean seed in an amount sufficient to impart a pink color to the soybean seed in a transverse section of the seed.
9. The soybean seed of claim 8, wherein the leghemoglobin protein is expressed in an amount of at least 0.5% of the total seed protein.
10. The soybean seed of claim 9, wherein the native seed storage protein gene encodes glycinin or conglycinin.
11. The soybean seed of claim 8, wherein the leghemoglobin coding sequence (i) encodes a polypeptide having at least 95% identity to SEQ ID NO: 2 or (ii) has at least 95% identity to SEQ ID NO: 1.
12. The soybean seed of claim 8, wherein the soybean seed further comprises a characteristic selected from the group consisting of: (i) an oleic acid content of at least 50% of the total seed fatty acids; (ii) a linolenic acid content of less than 3% of the total seed fatty acids; (iii) a protein content of at least 37% of the total weight of the soybean measured at or adjusted to 13% moisture, and (iv) any combination thereof.
13. The soybean seed of claim 8, wherein the soybean seed further comprises a recombinant construct integrated into its genome, the recombinant construct comprising a leghemoglobin coding sequence.
14. The soybean seed of claim 8, wherein the soybean seed comprises at least 37% protein by weight when measured or adjusted to 13% moisture.
15. The soybean seed of claim 14, wherein the soybean seed further comprises a modification to enhance seed protein content, the modification being in a gene encoding at least one of (i) a CCT-domain containing protein, (ii) a reticulon, (iii) a trehalose phosphate synthase, (iv) a HECT ubiquitin Ligase, (v) a MFT (mother of flowering) polypeptide, and (vi) a raffinose synthase.
16. A plant which produces the soybean seed of claim 8.
17. Meal extracted from the soybean seed of claim 8, wherein the meal comprises at least 0.1% leghemoglobin by weight of total protein.
18. A soybean seed comprising a modified leghemoglobin gene, wherein the modified leghemoglobin gene comprises at least one deletion, insertion or substitution into the native leghemoglobin gene, and wherein the leghemoglobin protein is expressed in the soybean seed in an amount sufficient to impart a pink color to the soybean seeds in a transverse section or the seed.
19. The soybean seed of claim 18, wherein the leghemoglobin protein is expressed in an amount of at least 0.5% of the total seed protein.
20. The soybean seed of claim 18, wherein the modified leghemoglobin gene comprises an insertion of a promoter or a regulatory enhancing sequence.
Description
REFERENCE TO SEQUENCE LISTING SUBMITTED ELECTRONICALLY
[0001] The official copy of the sequence listing is submitted electronically via EFS-Web as an ASCII formatted sequence listing with a file named 8429-US-PSP_SequenceListing_ST25.txt created on Oct. 28, 2020 and having a size of 94 kilobytes and is filed concurrently with the specification. The sequence listing comprised in this ASCII formatted document is part of the specification and is herein incorporated by reference in its entirety.
BACKGROUND
[0002] Animal-based meat replacement with plant-based proteins is becoming an industrial trend in food applications. Soy legume hemoglobin, or leghemoglobin, is a globin protein found in the nitrogen-fixing root nodules of leguminous plants. It carries heme, an iron-containing molecule, and functions to protect the nitrogenase enzyme from oxygen inactivation and to facilitate oxygen flow to the nitrogen-fixing bacteria. Leghemoglobin can be fermented from engineered yeast and has use in meat replacements by mimicking the flavor contributed by hemoglobin in meat. Compositions and methods to express leghemoglobin in soybeans are provided.
SUMMARY
[0003] Soybean seeds are provided which contain a leghemoglobin protein in an amount of at least 0.5% of total protein in the soybean seed, without the leghemoglobin being expressed from a recombinant construct comprising a leghemoglobin coding sequence integrated into the soybean genome. The soybean seed genome which does not comprise a recombinant construct containing a leghemoglobin coding sequence can be modified to introduce an insertion, deletion or substitution into a native leghemoglobin gene, such as the coding sequence or regulatory sequence or modified to replace all or part of a coding sequence of a seed storage protein with a leghemoglobin coding sequence.
[0004] Soybean seeds are provided in which the soybean seed genome is modified to introduce an insertion, deletion or substitution into a native leghemoglobin gene or modified to replace all or part of a coding sequence of a seed storage protein with a leghemoglobin coding sequence.
[0005] In some embodiments, a targeting sequence such as a transit peptide is operably linked to a leghemoglobin coding sequence to direct the leghemoglobin to an intracellular compartment such as a plastid.
[0006] In some embodiments, a soybean seed comprising a genomic modification is provided in which the leghemoglobin protein is expressed in the soybean seed in an amount sufficient to impart a pink color to the soybean seed in a transverse section of the seed or in an amount of at least 0.1% of the total seed protein.
[0007] In some embodiments the soybean is modified to directly express leghemoglobin specifically in the seed plastids.
[0008] In some embodiments, the soybean seed having one or more of a modification in the native leghemoglobin gene, an insertion of the leghemoglobin gene under the control of a different native promotor in the nuclear genome, or inclusion of the leghemoglobin sequence in the plastid genome, further comprises a recombinant construct comprising a leghemoglobin coding sequence integrated into its genome.
[0009] In some embodiments, the soybean seed contains further modifications such as nucleotide insertions, deletions, or substitutions into one or more genes encoding a glutamyl tRNA reductase, a ferrochetalase, a glutamyl tRNA reductase binding protein and an aminolevulinic acid synthase. In some embodiments, the soybean seed contains one or more recombinant constructs containing a coding sequence for a glutamyl tRNA reductase, a ferrochetalase, a glutamyl tRNA reductase binding protein and/or an aminolevulinic acid synthase.
[0010] Provided are soybean seeds which contain a leghemoglobin protein in an amount of at least 0.5% of total seed protein and which have one or more of the following characteristics: (i) an oleic acid content of at least 50% of the total seed fatty acids; (ii) a linolenic acid content of less than 3% of the total seed fatty acids; (iii) a protein content of at least 37% of the total weight of the soybean measured at or adjusted to 13% moisture; (iv) a Kunitz Trypsin protease inhibitor activity of less than 5% of that of a control unmodified soybean; (v) a Bowman-Burke protease inhibitors activity of less than 5% of a control unmodified soybean; (vi) a stachyose content of less than 1% weight percent at 13% moisture; and (vii) a raffinose content of less than 0.5% weight percent at 13% moisture.
[0011] In some embodiments, transgenic soybean seeds are provided which contain a recombinant construct comprising a polynucleotide encoding a leghemoglobin protein having at least 95% identity to SEQ ID NO: 2, in which the construct does not comprise a protein storage vesicle targeting sequence, and wherein the soybean does not contain (i) a recombinant construct comprising a sequence encoding a glutamyl tRNA reductase, or a truncated portion thereof, (ii) a recombinant construct comprising a sequence encoding a ferrochetalase, (iii) a recombinant construct comprising a glutamyl tRNA reductase binding protein and (iv) a recombinant construct comprising an aminolevulinic acid synthase, and wherein the soybean seed comprises leghemoglobin in the seed in an amount of at least 0.5% total seed protein.
[0012] In some embodiments, soybean seeds containing leghemoglobin protein in an amount of at least 0.5% of total protein have a genomic modification which includes at least one of (i) a nucleic acid insertion of a soybean genomic sequence which insertion excludes a non-soybean genomic sequence, (ii) one or more nucleic acid substitutions, (iii) one or more nucleic acid deletions, and (iv) any combination thereof, wherein the genomic modification comprises (a) a modification made to the native leghemoglobin gene or (b) an insertion comprising at least a portion of the native leghemoglobin gene.
[0013] In some embodiments, the soybean expresses leghemoglobin and further comprises a different modification to reduce or prevent expression of one or more seed storage coding sequences, such as a glycinin or a conglycinin.
[0014] In some embodiments, the soybean expresses leghemoglobin and further comprises one or more of high oleic acid, low linolenic acid, at least 37% total seed protein at 13% moisture.
[0015] In some embodiments, plants and plant parts grown from the modified soybean seeds containing leghemoglobin are provided.
[0016] In some embodiments, methods for processing soybean meal extracted from the modified soybean seeds which express leghemoglobin are provided in which the meal is contacted with at least one of a cellulase, a hemicellulase, and a pectinase under conditions sufficient to degrade the polysaccharides in the meal and the permeant is filtered from the residue. Meal extracted from the modified soybean seeds is provided containing at least 0.1%, 0.2%, 0.3%, 0.4% or 0.5% leghemoglobin by wt total protein.
[0017] In some embodiments, soybean isolate comprising at least 0.2% leghemoglobin by weight of total protein, with at least about 50% of the leghemoglobin being hemelated with an iron group is provided, which is produced from the modified seeds expressing leghemoglobin.
[0018] In some embodiments, a method is provided for producing soybean meal or isolate from modified soybean seeds comprising leghemoglobin and soybean seeds comprising high oleic acid in which the beans are processed to produce meal or isolate, comprising high oleic acid and leghemoglobin in which at least about 50% of the leghemoglobin is hemelated with an iron group.
BRIEF DESCRIPTION OF THE DRAWINGS AND THE SEQUENCE LISTING
[0019] The disclosure can be more fully understood from the following detailed description and the accompanying drawings and Sequence Listing, which form a part of this application.
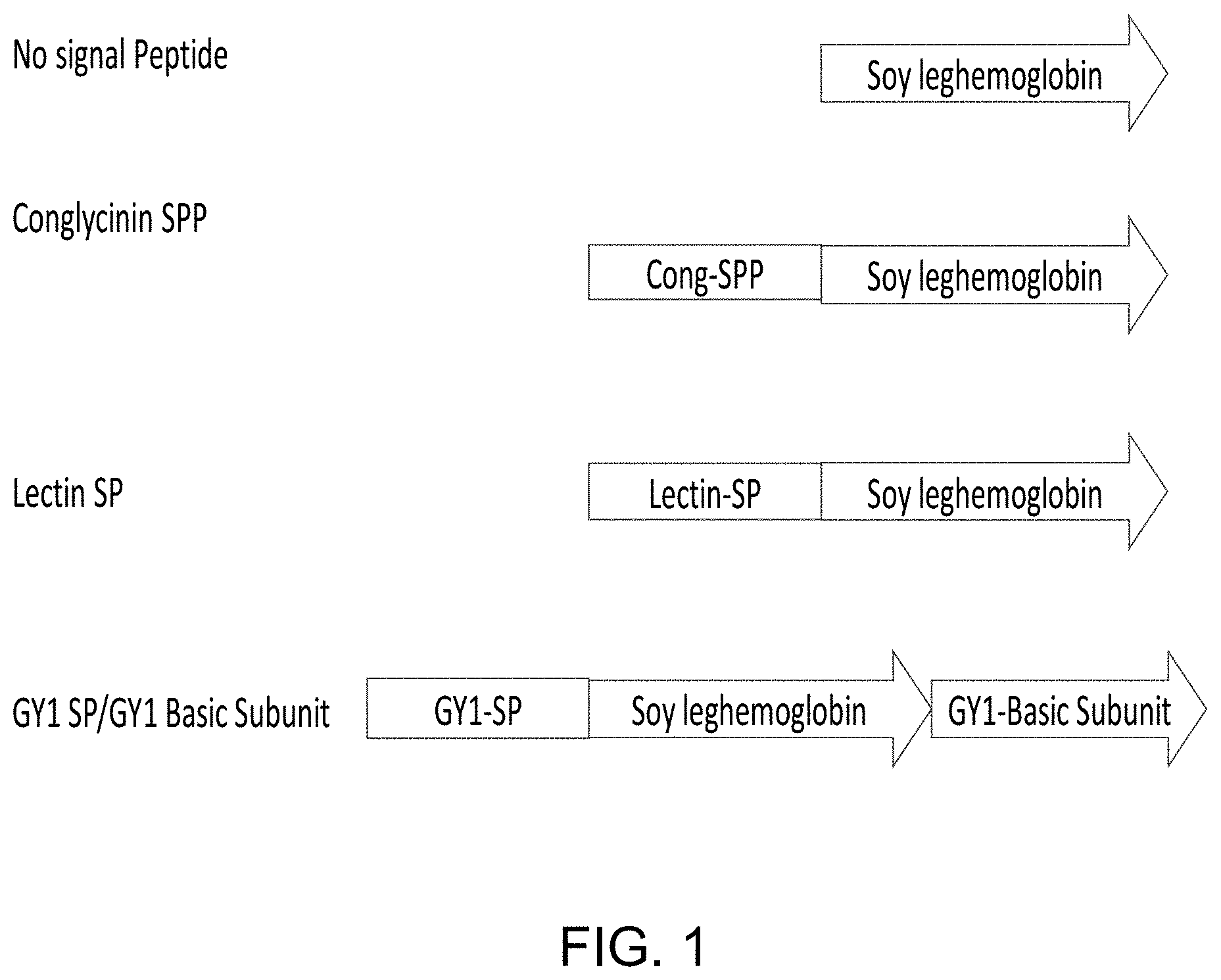
[0020] FIG. 1 is a chart showing construct design for the expression of soybean leghemoglobin with or without different protein targeting sequences
[0021] FIG. 2 is a chart showing construct design for the improvement of soybean leghemoglobin expression level by porphyrin pathway engineering.
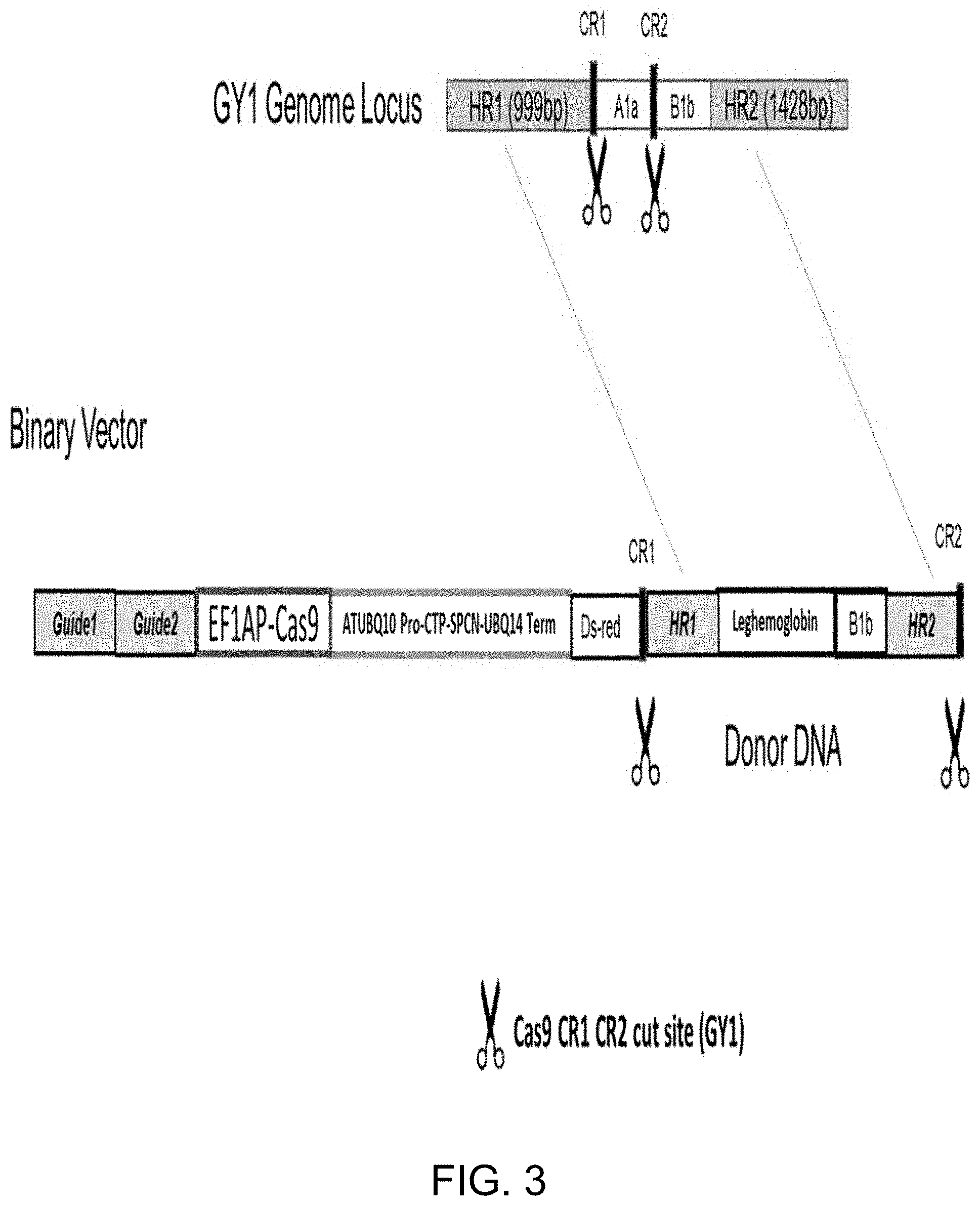
[0022] FIG. 3 is a schematic showing genome engineering of the leghemoglobin gene into the native soybean glycinin gene locus by CR1/CR2 gRNA pair.
[0023] FIG. 4 is a schematic showing genome engineering of the leghemoglobin gene into the native soybean glycinin gene locus by CR1/CR3 gRNA pair
[0024] FIG. 5 is a schematic showing the gene dropout strategy for the conglycinin gene cluster loci.
[0025] FIG. 6 is a photograph of a protein gel showing the seed protein profile of the conglycinin Gm10 gene cluster dropout variants.
[0026] FIG. 7 is a photograph of a protein gel showing the seed protein profile of the conglycinin Gm20 gene cluster dropout variants
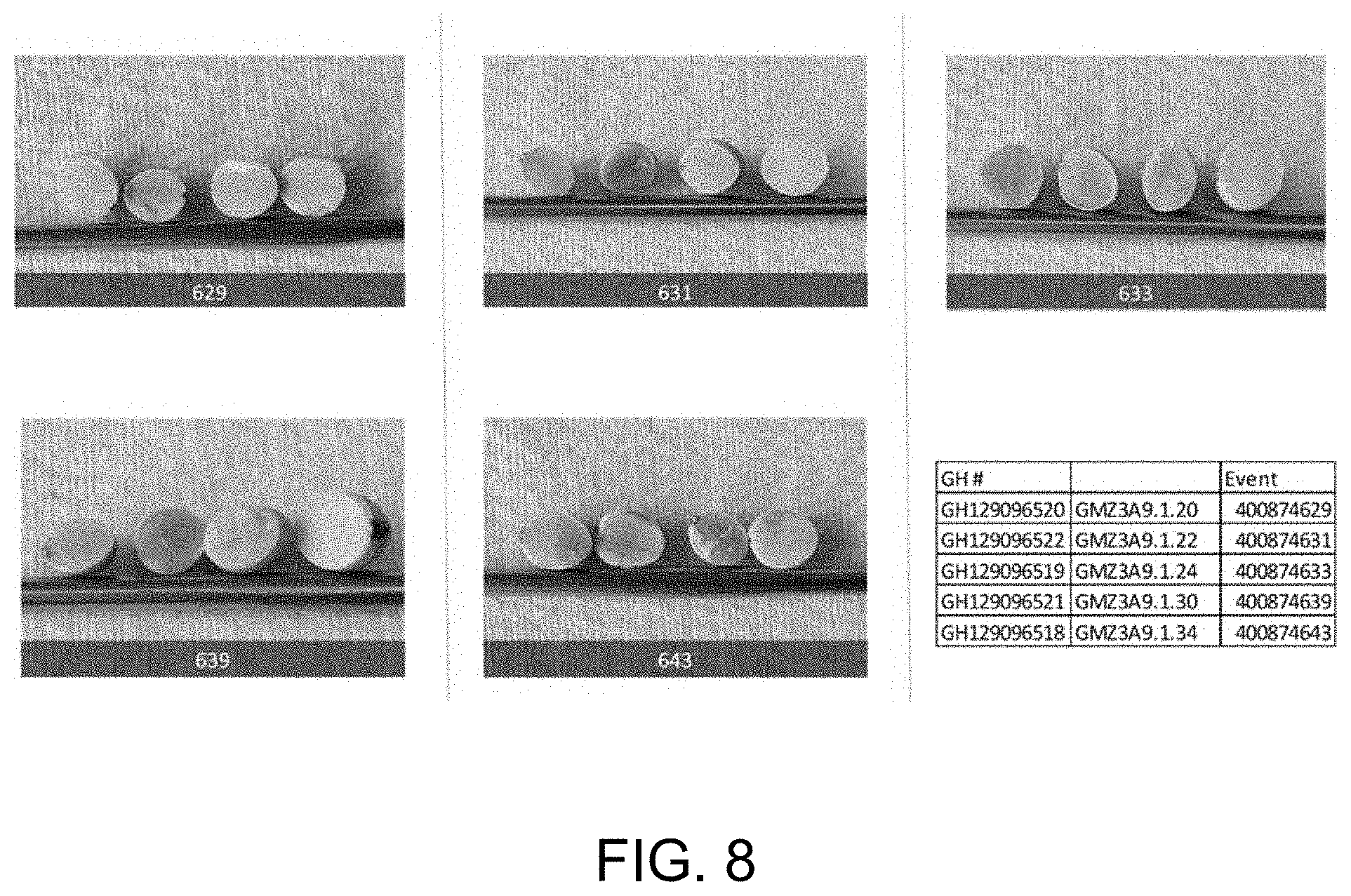
[0027] FIG. 8 is a photograph of seed cross sections of 5 independent events in experiment 1.
[0028] FIG. 9 is a photograph of a Coomassie-stained protein gel showing the presence of 16 KD leghemoglobin protein in experiment 1 (arrowed).
[0029] FIG. 10 is a photograph of seed cross sections of 4 independent events in experiment 5.
[0030] FIG. 11 a photograph of a Coomassie-stained protein gel showing the presence of 16 KD leghemoglobin protein in experiment 5.
[0031] FIG. 12 is a schematic flow chart showing an example of a generalized process for the Enzymatic Soybean Processing (E-SOY) process.
[0032] FIG. 13 is a schematic diagram showing a soybean nuclear transformation binary vector within the T-DNA.
[0033] FIG. 14 is a schematic diagram of soybean chloroplast transformation vector.
DETAILED DESCRIPTION
[0034] The sequence descriptions (Table 1) summarize the Sequence Listing attached hereto, which is hereby incorporated by reference. The Sequence Listing contains one letter codes for nucleotide sequence characters and the single and three letter codes for amino acids as defined in the IUPAC-IUB standards described in Nucleic Acids Research 13:3021-3030 (1985) and in the Biochemical Journal 219(2):345-373 (1984).
TABLE-US-00001 TABLE 1 Sequence Listing Description SEQ ID NO: Name Type 1 Glyma.20g191200 Leghemoglobin CDS DNA 2 Glyma.20g191200 Leghemoglobin peptide PRT 3 Beta-conglycinin SPP-Leghemoglobin CDS DNA 4 Beta-conglycinin SPP-Leghemoglobin peptide PRT 5 Lectin SP-Leghemoglobin CDS DNA 6 Lectin SP-Leghemoglobin Peptide PRT 7 GY1 SP-leghemoglobin-GY1 basic subunit CDS DNA 8 GY1 SP-leghemoglobin-GY1 basic subunit peptide PRT 9 beta-conglycinin promoter DNA 10 glycinin promoter DNA 11 Glyma.04g0898000 CDS DNA 12 Glyma.04g0898000 peptide PRT 13 Glyma.04g050400 CDS DNA 14 Glyma.04g050400 Peptide PRT 15 Phaseolin promoter DNA 16 Napin Promoter DNA 17 GM-GY1-CR1 RNA 18 GM-GY1-CR2 RNA 19 GM-GY1-CR3 RNA 20 glyma.03g163500 Glycinin 1 gene DNA 21 glyma.03g163500 Glycinin 1 peptide PRT 22 Donor DNA for GM-GY1-CR1/CR2 design DNA 23 Donor DNA for GM-GY1-CR1/CR3 design DNA 24 GM-CONG-gRNA1 RNA 25 GM-CONG-gRNA2 RNA 26 GM-CONG-gRNA3 RNA 27 GM-CONG-gRNA4 RNA 28 GM-CONG-gRNA5 RNA 29 GM-CONG-gRNA6 RNA 30 GM-CONG-gRNA7 RNA 31 Rubisco SSUSP::LH DNA 32 Rubisco SSUSP::LH PRT 33 Globulin peptide PRT 34 Globulin peptide PRT 35 Glyma.04G037000.1 CDS urophorphyrinogen III DNA synthase 36 Glyma.04G037000.1 polypeptide urophorphyrinogen PRT III synthase 37 glutamate-1-semialdehyde 2,1-aminomutase DNA Glyma.04G002900.1 38 glutamate-1-semialdehyde 2,1-aminomutase PRT Glyma.04G002900.1 39 Glutamyl-tRNA reductase-binding protein DNA Glyma.08G222600 40 Glutamyl-tRNA reductase-binding protein PRT Glyma.08G222600 41 chloroplast signal particle 43 Glyma.11G097200 DNA 42 chloroplast signal particle 43 Glyma.11G097200 PRT 43 Glyma.20g191200 leghemoglobin genomic sequence DNA 44 Disis GG artificial sequence DNA 45 Maize PPR protein CDS coding sequence DNA 46 Maize PPR protein polypetide sequence PRT 47 Soybean PPR protein CDS coding sequence DNA Glyma.15g162500 48 Soybean PPR protein polypetide sequence PRT Glyma.15g162500
[0035] The present disclosure describes modified soybean seeds expressing the leghemoglobin protein, the leghemoglobin complex or a combination thereof. Leghemoglobin is a protein synthesized in soy root nodules upon colonization by nitrogen-fixing bacteria. As used herein, "leghemoglobin protein" or "leghemoglobin" refer to the globulin protein or polypeptide, whether unfolded or folded into a monomer and which may or may not have associated with it a heme group (porphyrin bound to iron). As used herein "leghemoglobin complex" or "leghemoglobin protein complex" refers particularly to the complex which includes the leghemoglobin protein associated with a heme group (porphyrin bound to iron). Such a complex, when present in sufficient quantities can impart a red or pink color to the cells or tissue containing the complex, detectable to the eye, such as in the transverse section of a soybean seed expressing leghemoglobin complex. As used herein with respect to the color of a soybean in the transverse section, pink color means any shade of pink or red.
[0036] The soybean seeds can be modified to increase expression of leghemoglobin which forms a heme complex without the need to target expression of the leghemoglobin to a protein storage vesicle or other targeted cellular compartment.
[0037] In some embodiments the leghemoglobin without a heme group, the leghemoglobin complex, or a combination of both forms can be present in soybean seeds at at least 0.01%, 0.05%, 0.5%, 1%, 1.5%, 2%, 2.5%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10% or more and less than 75%, 50%, 25%, 20%, 15%, 10%, 5%, 4% or 3% of the total seed protein.
[0038] Suitably at least 5, 10, 15, 20, 25, 30, 40, 50, 60, 70, 80, 90 or 95 percent and less than 100, 99.9, 95, 90, 85, 80, 70, 60 or 50 percent of the total leghemoglobin forms a complex with a heme group in the soybean seed.
[0039] Soybean seeds disclosed herein, as well as plant parts, plant cells, tissue cultures and plants grown therefrom are provided.
[0040] In certain embodiments, the soybean seeds have introduced a recombinant construct comprising a leghemoglobin coding sequence operably linked to a promoter functional in a soybean seed cell. As used herein a recombinant construct is a construct that includes a promoter sequence operably linked to a polynucleotide encoding a polypeptide, and optionally other regulatory sequences wherein the recombinant construct is exogenous to the plant, plant cell or seed. As used herein, the term "exogenous" with respect to a nucleic acid indicates that the nucleic acid is not in its natural genomic location. Such plants containing recombinant constructs are referred to as transgenic plants. A regulatory sequence can be a sequence in or around a gene which promotes transcription or termination of polypeptide coding sequences.
[0041] Non-limiting examples of recombinant DNA constructs include a polynucleotide of interest operably linked to heterologous sequences, also referred to as "regulatory elements," which aid in the expression, autologous replication, and/or genomic insertion of the sequence of interest. Such regulatory elements include, for example, promoters, termination sequences, enhancers, etc., or any component of an expression cassette; a plasmid, cosmid, virus, autonomously replicating sequence, phage, or linear or circular single-stranded or double-stranded DNA or RNA nucleotide sequence; and/or sequences that encode heterologous polypeptides.
[0042] The recombinant DNA constructs or recombinant constructs provided comprise at least one regulatory element that when integrated into the genome is not present in its native location in the soybean genome or is from the genome of another species. In certain embodiments, the at least one regulatory element of the recombinant DNA construct comprises a promoter, preferably a heterologous promoter that drives expression of the leghemoglobin in seeds, such as a glycinin or conglycinin promotor.
[0043] In an embodiment, the soybean seed containing a recombinant construct comprising a leghemoglobin coding sequence do not contain one or more or all of the following (i) a recombinant construct comprising a sequence encoding a glutamyl tRNA reductase, or a truncated portion thereof, (ii) a recombinant construct comprising a sequence encoding a ferrochetalase, (iii) a recombinant construct comprising a glutamyl tRNA reductase binding protein and (iv) a recombinant construct comprising an aminolevulinic acid synthase. A truncated coding sequence is a sequence that has had the N' or C' terminus, or both, of the coding sequence removed such that a polypeptide is synthesized from the coding sequence that is shorter than the native non-truncated polypeptide and lacks a number of amino acids at the C' terminus the N' terminus or both. The inventors discovered that high levels of leghemoglobin and leghemoglobin complex could be expressed in soybean seeds in amounts detectable by eye through conferring a pink color without the need to augment expression with these additional recombinant constructs and without the need to include a targeting sequence that would direct the leghemoglobin to a particular cellular compartment, such as a protein storage vesicle.
[0044] In some embodiments the native leghemoglobin gene is modified. The genomic sequence of the leghemoglobin gene is provided in SEQ ID NO:43 and modifications may be made to or include all or part of this sequence or to a sequence corresponding to SEQ ID NO: 43 in the soybean genome, including to specific regions identified herein. With respect to SEQ ID NO: 43, the regulatory region, including the promotor and 5' UTR, is from nucleotide position 1 to position 2058, exon 1 is from position 2059 to position 2156, intron 1 is from position 2157 to position 2275, exon 2 is from position 2276 to position 2384, intron 2 is from position 2385 to position 2574, exon 3 is from position 2575 to position 2679, intron 3 is from position 2680 to position 2876, exon 4 is from position 2877 to position 3002, the terminator, including the 3' UTR, is from position 3003 to position 5214.
[0045] In some embodiments, the modification is made from position 1-2058 of SEQ ID NO:43, 100-2058 of SEQ ID NO:43, 200-2058 of SEQ ID NO:43, 300-2058 of SEQ ID NO:43, 400-2058 of SEQ ID NO:43, 500-2058 of SEQ ID NO:43, 600-2058 of SEQ ID NO:43, 700-2058 of SEQ ID NO:43, 800-2058 of SEQ ID NO:43, 900-2058 of SEQ ID NO:43, 1000-2058 of SEQ ID NO:43, 1100-2058 of SEQ ID NO:43, 1200-2058 of SEQ ID NO:43, 1300-2058 of SEQ ID NO:43, 1400-2058 of SEQ ID NO:43, 1500-2058 of SEQ ID NO:43, 1600-2058 of SEQ ID NO:43, 1700-2058 of SEQ ID NO:43, 1800-2058 of SEQ ID NO:43, or 1900-2058 of SEQ ID NO:43.
[0046] In some embodiments, the plastid genome of the soybean seed comprises a modification in which a sequence encoding a leghemoglobin protein is inserted into the plastid genome, such that the leghemoglobin polypeptide is expressed directly in the seed plastids without the need for a transit peptide. Seed-specific plastid transformation can be achieved by inserting a construct comprising the leghemoglobin coding sequence connected to a DicisGG sequence such as SEQ ID NO: 44. The plant is co-transformed to express a PPR protein, such as a PPR10 protein such as SEQ ID NO: 46 or 48, with the sequence encoding the PPR protein under the control of a promotor active in the seed and which is seed-specific, such as a promotor of a seed storage protein such as glycinin or conglycinin. Different seed-specific promotors can be selected to modulate the amount of expression of the PPR protein. Alternatively, expression of the PPR protein can be achieved through genome editing to replace all or part of a native sequence which is expressed in a seed specific fashion, such as at the locus of a seed storage protein. The PPR protein acts as a trigger in the seed for the DicisGG sequence to facilitate direct expression of the leghemoglobin in the seed plastids, without or with little or minimal expression occurring in non-seed plant parts such as the roots, stems, leaves and flowers.
[0047] The plastid expression of leghemoglobin can be combined with expression of leghemoglobin from a nuclear genomic source, such as by transformation of the nuclear genome with a transgenic construct, or by genome editing of a native nuclear gene, such as by inserting, deleting or substituting one or more nucleotides into the native leghemoglobin gene or by genome editing of a gene highly expressed in the soybean seed such as by inserting the leghemoglobin sequence into a seed storage protein gene to replace all or part of the coding sequence of the seed storage protein such that the leghemoglobin is expressed instead of the seed storage protein. The plastid expression of leghemoglobin can be combined with increased or decreased expression of other genes encoding or contributing to the control of the heme biosynthesis pathway, such as disclosed herein.
[0048] In some embodiments, an intracellular targeting sequence or transit sequence such as a plastid targeting sequence is included and operably linked to a sequence encoding leghemoglobin, such as being placed just before the N' terminus of a sequence encoding leghemoglobin, such that the intracellular targeting sequence targets expression of the leghemoglobin to an intracellular compartment such as a protein storage vesicle or a plastid. The targeting sequence and operably linked leghemoglobin sequence, such as occurs in SEQ ID NO: 31 or a polynucleotide encoding SEQ ID NO: 32, can be operably linked to a regulatory sequence in a recombinant construct and used to transform soybean. The targeting sequence can be operably linked to a leghemoglobin sequence, such as occurs in SEQ ID NO: 31, or a sequence encoding SEQ ID NO: 32, and can be inserted through genome editing to replace all or part of the coding sequence of a seed storage protein such as glycinin or conglycinin, such that the native regulatory elements of the seed storage protein direct expression of the targeting sequence and the leghemoglobin coding sequence such that the leghemoglobin protein is expressed with a transit peptide and targeted to an intracellular compartment. The targeting sequence can be inserted into the native leghemoglobin gene, optionally with other insertions, or deletions or substitutions, so that leghemoglobin is expressed in the soybean seed from its native locus with a transit peptide and targeted to an intracellular compartment. In one embodiment the plastid targeting sequence is included at the N' terminus of the coding sequence or polypeptide of interest. One example of a plastid targeting sequence is the Rubisco SSUSP plastid targeting sequence, such as encoded by the nucleotide sequence from position 1 to position 165 of SEQ ID NO: 31, with the corresponding peptide targeting sequence at position 1 to position 55 of SEQ ID NO: 32. The leghemoglobin coding sequence is from position 166 to position 603 of SEQ ID NO: 31 and the corresponding peptide form position 56 to position 200 of SEQ ID NO: 32.
[0049] In some embodiments, soybean seeds are provided which express leghemoglobin from two or more sources, constructs or genomic locations, such as from two or more of (i) a recombinant construct inserted into the genome, (ii) a genome modification in which the leghemoglobin coding sequence replaces all or part of a seed storage protein coding sequence such as described herein (iii) a genome modification in which the native leghemoglobin gene is modified to include one or more of an insertion, deletion or substitution, such as into the regulatory region or coding sequence of the leghemoglobin gene and (iv) a plastid genome modification in which the plastid genome is modified to express a leghemoglobin coding sequence. In some embodiments, the two or more sources include at least one source in which the leghemoglobin coding sequence is operably linked to an intracellular targeting sequence, such as a plastid targeting sequence as described herein, and another source in which the leghemoglobin coding sequence is not operably linked to an intracellular targeting sequence.
[0050] In certain embodiments, the soybean seeds that comprise leghemoglobin and optionally other modifications as described herein can further comprise a modification to increase the amount of leghemoglobin complex in the soybean seed. The modification to increase leghemoglobin complex can include modified expression of one or more of a glutamyl-tRNA reductase, a glutamate-1-semialdehyde 2,1-aminomutase, an aminolevulinate dehydratase (HEMB1), a hydroxymethylbilane synthase (NEMC), a urophorphyrinogen III synthase, a urophorphyrinogen decarboxylase, a coporphyrinogen III oxidase (HEMF,CPDX), a protoporphyrinogen oxidase (PPDX), and/or a ferrochelatase. The modification can include the introduction of a recombinant construct into the genome of the plant, or the modification can include a gene editing modification, such as an insertion, deletion and/or substitution into the genes from which these polypeptides are expressed, such as to enhance transcription of the coding sequences of these genes.
[0051] In some embodiments, the soybean plants, cells and seeds comprise modifications in genes that encode regulatory proteins that modulate expression or activity of enzymes contributing to heme production or hemelation of leghemoglobin. For example, soybean genes encoding proteins that regulate glutamyl-tRNA reductase activity include glutamyl-tRNA reductase-binding protein (Glyma.08G222600), chloroplast signal particle 43(Glyma.11G097200) and FLUORESCENT IN BLUE LIGHT (Glyma.16G010200 and Glyma.07G041700) can be modified, such as by insertion, deletion or substitution to increase or enhance the formation of heme and/or the leghemoglobin complex in soybean.
[0052] In certain embodiments, the soybean seeds are edited to contain a leghemoglobin coding sequence inserted into a native gene encoding a seed storage protein and replacing in whole or in part the native seed storage coding sequence. Such an edited construct comprising an exogenous nucleic acid coding sequence operably linked to a native promoter in its native position in the genome would not be considered a recombinant construct, because the promoter and other regulatory elements are not exogenous to their native environment. For example, in an edited genome, the gene structure can remain largely unaltered, with the native seed-storage protein coding sequence being replaced by a different coding sequence, such as with a globulin protein, such as leghemoglobin. Such plants, seeds and cells may be referred to as modified or edited plants, seeds or cells.
[0053] One or more suitable seed storage protein coding sequences can be replaced with a globulin coding sequence using the methods described herein, such as a leghemoglobin coding sequence, including for example a sequence encoding a glycinin, conglycinin, 2S albumin, Kunitz trypsin inhibitor (KTI), a Bowman-Birk inhibitor (BBI), or a combination thereof.
[0054] The Kunitz trypsin inhibitor (KTI) and Bowman-Birk inhibitor (BBI) activity can be reduced to less than 50%, 40%, 30%, 20%, 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3%, 2%, 1%, 0.5%, 0.4%, 0.3%, 0.2% or 0.1% of the activity found in a wild type, null or control soybean which has not had a modification to reduce or prevent expression of a Kunitz trypsin inhibitor (KTI) or Bowman-Birk inhibitor (BBI) coding sequence.
[0055] As used herein "encoding," "encoded," or the like, with respect to a specified nucleic acid, is meant comprising the information for translation into the specified protein. A nucleic acid encoding a protein may comprise non-translated sequences (e.g., introns) within translated regions of the nucleic acid, or may lack such intervening non-translated sequences (e.g., as in cDNA). The information by which a protein is encoded is specified by the use of codons. Typically, the amino acid sequence is encoded by the nucleic acid using the "universal" genetic code. However, variants of the universal code, such as is present in some plant, animal and fungal mitochondria, the bacterium Mycoplasma capricolum (Yamao, et al., (1985) Proc. Natl. Acad. Sci. USA 82:2306-9) or the ciliate Macronucleus, may be used when the nucleic acid is expressed using these organisms.
[0056] When the nucleic acid is prepared or altered synthetically, advantage can be taken of known codon preferences of the intended host where the nucleic acid is to be expressed. For example, although nucleic acid sequences disclosed herein may be expressed in both monocotyledonous and dicotyledonous plant species, sequences can be modified to account for the specific codon preferences and GC content preferences of monocotyledonous plants or dicotyledonous plants as these preferences have been shown to differ (Murray, et al., (1989) Nucleic Acids Res. 17:477-98).
[0057] As used herein, "polynucleotide" includes reference to a deoxyribopolynucleotide, ribopolynucleotide or analogs thereof that have the essential nature of a natural ribonucleotide in that they hybridize, under stringent hybridization conditions, to substantially the same nucleotide sequence as naturally occurring nucleotides and/or allow translation into the same amino acid(s) as the naturally occurring nucleotide(s). A polynucleotide can be full-length or a subsequence of a structural or regulatory gene. Unless otherwise indicated, the term includes reference to the specified sequence as well as the complementary sequence thereof. Thus, DNAs or RNAs with backbones modified for stability or for other reasons are "polynucleotides" as that term is intended herein. Moreover, DNAs or RNAs comprising unusual bases, such as inosine, or modified bases, such as tritylated bases, to name just two examples, are polynucleotides as the term is used herein. It will be appreciated that a great variety of modifications have been made to DNA and RNA that serve many useful purposes known to those of skill in the art. The term polynucleotide as it is employed herein embraces such chemically, enzymatically or metabolically modified forms of polynucleotides, as well as the chemical forms of DNA and RNA characteristic of viruses and cells, including inter alia, simple and complex cells.
[0058] The terms "polypeptide," "peptide" and "protein" are used interchangeably herein to refer to a polymer of amino acid residues. The terms apply to amino acid polymers in which one or more amino acid residue is an artificial chemical analogue of a corresponding naturally occurring amino acid, as well as to naturally occurring amino acid polymers.
[0059] As used herein, "sequence identity" or "identity" in the context of two nucleic acid or polypeptide sequences includes reference to the residues in the two sequences, which are the same when aligned for maximum correspondence over a specified comparison window. When percentage of sequence identity is used in reference to proteins it is recognized that residue positions which are not identical often differ by conservative amino acid substitutions, where amino acid residues are substituted for other amino acid residues with similar chemical properties (e.g., charge or hydrophobicity) and therefore do not change the functional properties of the molecule. Where sequences differ in conservative substitutions, a percent similarity may be used. Sequences, which differ by such conservative substitutions, are said to have "sequence similarity" or "similarity." Means for making this adjustment are well known to those of skill in the art. Typically, this involves scoring a conservative substitution as a partial rather than a full mismatch, thereby increasing the percentage sequence identity. Thus, for example, where an identical amino acid is given a score of 1 and a non-conservative substitution is given a score of zero, a conservative substitution is given a score between zero and 1. The scoring of conservative substitutions is calculated, e.g., according to the algorithm of Meyers and Miller, (1988) Computer Applic. Biol. Sci. 4:11-17, e.g., as implemented in the program PC/GENE (Intelligenetics, Mountain View, Calif., USA).
[0060] As used herein, "percentage of sequence identity" means the value determined by comparing two optimally aligned sequences over a comparison window, wherein the portion of the polynucleotide sequence in the comparison window may comprise additions or deletions (i.e., gaps) as compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. The percentage is calculated by determining the number of positions at which the identical nucleic acid base or amino acid residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison and multiplying the result by 100 to yield the percentage of sequence identity.
[0061] Provided are polynucleotide and polypeptide sequences which have at least or at least about 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.6%, 99.7%, 99.8% or 99.9% and less than 100%, 99%, 95% or 90% identity to the polypeptides and polynucleotides of any one of SEQ ID NOs: 1-48, or to specified sequences within defined positions of any one of SEQ ID NOs: 1-48, such as disclosed herein.
[0062] As used herein, "reference sequence" is a defined sequence used as a basis for sequence comparison. A reference sequence may be a subset or the entirety of a specified sequence; for example, as a segment of a full-length cDNA or gene sequence or the complete cDNA or gene sequence.
[0063] As used herein, "comparison window" means reference to a contiguous and specified segment of a polynucleotide sequence, wherein the polynucleotide sequence may be compared to a reference sequence and wherein the portion of the polynucleotide sequence in the comparison window may comprise additions or deletions (i.e., gaps) compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. Generally, the comparison window is at least 20 contiguous nucleotides in length, and optionally can be 30, 40, 50, 100 or longer. Those of skill in the art understand that to avoid a high similarity to a reference sequence due to inclusion of gaps in the polynucleotide sequence a gap penalty is typically introduced and is subtracted from the number of matches.
[0064] Methods of alignment of nucleotide and amino acid sequences for comparison are well known in the art. The local homology algorithm (BESTFIT) of Smith and Waterman, (1981) Adv. Appl. Math 2:482, may conduct optimal alignment of sequences for comparison; by the homology alignment algorithm (GAP) of Needleman and Wunsch, (1970) J. Mol. Biol. 48:443-53; by the search for similarity method (Tfasta and Fasta) of Pearson and Lipman, (1988) Proc. Natl. Acad. Sci. USA 85:2444; by computerized implementations of these algorithms, including, but not limited to: CLUSTAL in the PC/Gene program by Intelligenetics, Mountain View, Calif., GAP, BESTFIT, BLAST, FASTA and TFASTA in the Wisconsin Genetics Software Package.RTM., Version 8 (available from Genetics Computer Group (GCG.RTM. programs (Accelrys, Inc., San Diego, Calif.)). The CLUSTAL program is well described by Higgins and Sharp, (1988) Gene 73:237-44; Higgins and Sharp, (1989) CABIOS 5:151-3; Corpet, et al., (1988) Nucleic Acids Res. 16:10881-90; Huang, et al., (1992) Computer Applications in the Biosciences 8:155-65, and Pearson, et al., (1994) Meth. Mol. Biol. 24:307-31. The preferred program to use for optimal global alignment of multiple sequences is PileUp (Feng and Doolittle, (1987) J. Mol. Evol., 25:351-60 which is similar to the method described by Higgins and Sharp, (1989) CABIOS 5:151-53 and hereby incorporated by reference). The BLAST family of programs which can be used for database similarity searches includes: BLASTN for nucleotide query sequences against nucleotide database sequences; BLASTX for nucleotide query sequences against protein database sequences; BLASTP for protein query sequences against protein database sequences; TBLASTN for protein query sequences against nucleotide database sequences; and TBLASTX for nucleotide query sequences against nucleotide database sequences. See, Current Protocols in Molecular Biology, Chapter 19, Ausubel, et al., eds., Greene Publishing and Wiley-Interscience, New York (1995).
[0065] GAP uses the algorithm of Needleman and Wunsch, supra, to find the alignment of two complete sequences that maximizes the number of matches and minimizes the number of gaps. GAP considers all possible alignments and gap positions and creates the alignment with the largest number of matched bases and the fewest gaps. It allows for the provision of a gap creation penalty and a gap extension penalty in units of matched bases. GAP must make a profit of gap creation penalty number of matches for each gap it inserts. If a gap extension penalty greater than zero is chosen, GAP must, in addition, make a profit for each gap inserted of the length of the gap times the gap extension penalty. Default gap creation penalty values and gap extension penalty values in Version 10 of the Wisconsin Genetics Software Package.RTM. are 8 and 2, respectively. The gap creation and gap extension penalties can be expressed as an integer selected from the group of integers consisting of from 0 to 100. Thus, for example, the gap creation and gap extension penalties can be 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, 30, 40, 50 or greater.
[0066] GAP presents one member of the family of best alignments. There may be many members of this family, but no other member has a better quality. GAP displays four figures of merit for alignments: Quality, Ratio, Identity and Similarity. The Quality is the metric maximized in order to align the sequences. Ratio is the quality divided by the number of bases in the shorter segment. Percent Identity is the percent of the symbols that actually match. Percent Similarity is the percent of the symbols that are similar. Symbols that are across from gaps are ignored. A similarity is scored when the scoring matrix value for a pair of symbols is greater than or equal to 0.50, the similarity threshold. The scoring matrix used in Version 10 of the Wisconsin Genetics Software Package.RTM. is BLOSUM62 (see, Henikoff and Henikoff, (1989) Proc. Natl. Acad. Sci. USA 89:10915).
[0067] Unless otherwise stated, sequence identity/similarity values provided herein refer to the value obtained using the BLAST 2.0 suite of programs using default parameters (Altschul, et al., (1997) Nucleic Acids Res. 25:3389-402).
[0068] The leghemoglobin sequences and recombinant constructs described herein can be provided for expression in a plant of interest or an organism of interest. The cassette can include 5' and 3' regulatory sequences operably linked to a leghemoglobin polynucleotide or a modified leghemoglobin polynucleotide. "Operably linked" is intended to mean a functional linkage between two or more elements. For, example, an operable linkage between a polynucleotide of interest and a regulatory sequence (e.g., a promoter) is a functional link that allows for expression of the polynucleotide of interest. Operably linked elements may be contiguous or non-contiguous. When used to refer to the joining of two protein coding regions, operably linked is intended that the coding regions are in the same reading frame. The cassette may additionally contain at least one additional gene to be co-transformed into the organism. Alternatively, the additional gene(s) can be provided on multiple expression cassettes. Such an expression cassette is provided with a plurality of restriction sites and/or recombination sites for insertion of the modified glycinin polynucleotide to be under the transcriptional regulation of the regulatory regions. The expression cassette may additionally contain selectable marker genes.
[0069] The expression cassette can include in the 5'-3' direction of transcription, a transcriptional and translational initiation region (e.g., a promoter), a modified leghemoglobin polynucleotide described herein, and a transcriptional and translational termination region (e.g., termination region) functional in plants. The regulatory regions (e.g., promoters, transcriptional regulatory regions, and translational termination regions) and/or the modified leghemoglobin polynucleotide may be native/analogous to the host cell or to each other. Alternatively, the regulatory regions and/or the modified leghemoglobin polynucleotide may be heterologous to the host cell or to each other.
[0070] As used herein, "heterologous" in reference to a sequence is a sequence that originates from a foreign species, or, if from the same species, is substantially modified from its native form in composition and/or genomic locus by deliberate human intervention. For example, a promoter operably linked to a heterologous polynucleotide that is from a species different from the species from which the polynucleotide was derived, or, if from the same/analogous species, one or both are substantially modified from their original form and/or genomic locus, or the promoter is not the native promoter for the operably linked polynucleotide.
[0071] The termination region may be native with the transcriptional initiation region, with the plant host, or may be derived from another source (i.e., foreign or heterologous) than the promoter, the modified glycinin polynucleotide, the plant host, or any combination thereof.
[0072] The expression cassette may additionally contain a 5' leader sequences. Such leader sequences can act to enhance translation. Translation leaders are known in the art and include viral translational leader sequences.
[0073] In preparing the expression cassette, the various DNA fragments may be manipulated, so as to provide for the DNA sequences in the proper orientation and, as appropriate, in the proper reading frame. Toward this end, adapters or linkers may be employed to join the DNA fragments or other manipulations may be involved to provide for convenient restriction sites, removal of superfluous DNA, removal of restriction sites, or the like. For this purpose, in vitro mutagenesis, primer repair, restriction, annealing, resubstitutions, e.g., transitions and transversions, may be involved.
[0074] As used herein "promoter" refers to a region of DNA upstream from the start of transcription and involved in recognition and binding of RNA polymerase and other proteins to initiate transcription. A "plant promoter" is a promoter capable of initiating transcription in plant cells. Exemplary plant promoters include, but are not limited to, those that are obtained from plants, plant viruses and bacteria which comprise genes expressed in plant cells such Agrobacterium or Rhizobium. Certain types of promoters preferentially initiate transcription in certain tissues, such as leaves, roots, seeds, fibres, xylem vessels, tracheids or sclerenchyma. Such promoters are referred to as "tissue preferred." A "cell type" specific promoter primarily drives expression in certain cell types in one or more organs, for example, vascular cells in roots or leaves. An "inducible" or "regulatable" promoter is a promoter, which is under environmental control. Examples of environmental conditions that may affect transcription by inducible promoters include anaerobic conditions or the presence of light. Another type of promoter is a developmentally regulated promoter, for example, a promoter that drives expression during pollen development. Tissue preferred, cell type specific, developmentally regulated and inducible promoters constitute the class of "non-constitutive" promoters. A "constitutive" promoter is a promoter, which is active under most environmental conditions. Constitutive promoters include, for example, the core promoter of the Rsyn7 promoter and other constitutive promoters disclosed in WO 99/43838 and U.S. Pat. No. 6,072,050; the core CaMV 35S promoter (Odell et al. (1985) Nature 313:810-812); rice actin (McElroy et al. (1990) Plant Cell 2:163-171); ubiquitin (Christensen et al. (1989) Plant Mol. Biol. 12:619-632 and Christensen et al. (1992) Plant Mol. Biol. 18:675-689); pEMU (Last et al. (1991) Theor. Appl. Genet. 81:581-588); MAS (Velten et al. (1984) EMBO J. 3:2723-2730); ALS promoter (U.S. Pat. No. 5,659,026), and the like. Other constitutive promoters include, for example, U.S. Pat. Nos. 5,608,149; 5,608,144; 5,604,121; 5,569,597; 5,466,785; 5,399,680; 5,268,463; 5,608,142; and 6,177,611.
[0075] Also contemplated are synthetic promoters which include a combination of one or more heterologous regulatory elements.
[0076] The promoter can be any type or class of promoter known in the art, such that any one of a number of promoters can be used to express the various modified leghemoglobin sequences disclosed herein, including the native promoter of the polynucleotide sequence of interest. The promoters for use in the recombinant DNA constructs disclosed herein can be selected based on the desired outcome.
[0077] In certain embodiments, the recombinant DNA construct, described herein is expressed in a plant or seed. In certain embodiment, the plant or seed is a soybean plant or soybean seed. As used herein, the term "plant" includes plant protoplasts, plant cell tissue cultures from which plants can be regenerated, plant calli, plant clumps, and plant cells that are intact in plants or parts of plants such as embryos, pollen, ovules, seeds, leaves, flowers, branches, fruit, kernels, ears, cobs, husks, stalks, roots, root tips, anthers, and the like. Grain is intended to mean the mature seed produced by commercial growers for purposes other than growing or reproducing the species. Progeny, variants, and mutants of the regenerated plants are also included within the scope of the disclosure, provided that these parts comprise the introduced polynucleotides.
[0078] In certain embodiments, the soybean plants or soybean seeds further comprise at least one additional modification that increases the total protein in the seed as compared to a control seed (e.g., seed not comprising the at least one modification). In certain embodiments, the soybean seed comprising the at least one modification comprises at least about a 1%, 1.5%, 2%, 2.5%, 3%, 3.5%, 4%, 5%, 10%, or 15% and less than 20%, 15%, 10%, 9%, 8%, 7%, 6%, or 5% percentage point increase in total protein measured on a dry weight basis, as compared to a control seed.
[0079] In certain embodiments, the soybean plants or soybean seeds further comprise at least one additional modification decreasing the raffinose family oligosaccharides (RFO) content in the seed. In certain embodiments, the modification comprises a decrease in the expression and/or activity of a raffinose synthase. In certain embodiments, the modification comprises a decrease in the expression and/or activity of raffinose synthase 2 (RS2) and/or raffinose synthase 4 (RS4). In certain embodiments, the soybean seed comprises at least a 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 99% decrease in expression of RS2, RS4, or RS2 and RS4, as compared to a control seed. In certain embodiments, the seed comprises less than about 6%, 5.5%, 5%, 4.5%, 4%, 3.5%, 3%, 2.5%, 2%, 1.5%, 1%, or 0.5% RFO content on a dry weight basis. In certain embodiments, the introduced modification decreases RFO content by at least about a 1%, 1.5%, 2%, 2.5%, 3%, 3.5%, 4%, 5%, 10%, or 15% and less than 20%, 15%, 10%, 9%, 8%, 7%, 6%, or 5% percentage point increase in total protein measured on a dry weight basis, as compared to a control seed.
[0080] In certain embodiments, the soybean plants or soybean seeds further comprise at least one additional modification that increases the amount of oleic acid in the seed, decreases the amount of linolenic acid in the seed, increases the amount of seed protein, or a combination thereof. For example, the modification can be in a FAD2-1A, FAD2-1B, FAD3a, FAD3b gene.
[0081] In certain embodiments, the soybean plants or soybean seeds further comprise at least one additional modification that increases the amount of total protein, for example by modification of one or more of a gene encoding (i) a CCT-domain containing protein, (ii) a reticulon, (iii) a trehalose phosphate synthase, (iv) a HECT Ubiquitin Ligase (HEL or UPL3), (v) a MFT (mother of flowering) polypeptide, (vi) a raffinose synthase RS2, RS3, or RS4, such as disclosed in U.S. Pat. Nos. 5,710,365, 8,728,726, and 10,081,814 each of which are incorporated herein by reference in their entirety or (vii) any combination thereof.
[0082] For example, provided are soybean seeds which comprise leghemoglobin in an amount disclosed herein and which can be processed to produce oils and meals, and the oils produced therefrom, which soybeans and/or oils have at least or at least about 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89 or 90 percent oleic (C 18:1) acid of the total fatty acids by weight and less than or less than about 100, 99, 98, 97, 96, 95, 94, 93, 92, 91, 90, 89, 88, 87, 86, 85, 84, 83, 82, 81, 80, 79, 78, 76, 75, 74, 73, 72, 71 or 70 percent oleic acid of the total fatty acids by weight.
[0083] For example, provided are soybean seeds which comprise leghemoglobin in an amount disclosed herein in an amount disclosed herein, which can be processed to produce oils, and the oils produced therefrom, which soybeans and/or oils have at least or at least about 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, 1.9, 2.0, 2.1, 2.2, 2.3, 2.4, 2.5, 2.6, 2.7, 2.8, 2.9, or 3.0 percent linolenic (C 18:3) acid of the total fatty acids by weight and less than or less than about 6, 5.5, 5, 4.5, 4, 3.9, 3.8, 3.7, 3.6, 3.5, 3.4, 3.3, 3.2, 3.1, 3.0, 2.9, 2.8, 2.7, 2.6, 2.5, 2.4, 2.3, 2.2, 2.1 or 2.0 percent linolenic acid of the total fatty acids by weight.
[0084] For example, provided are soybean seeds which comprise leghemoglobin in an amount disclosed herein and which have a protein content of at least or at least about 35%, 36%, 37%, 38%, 39%, 40%, 41%, 42%, 43%, 44%, 45%, 46%, 47%, 48%, 49%, 50%, 51%, 52%, 53% 54% or 55% and less than or less than about 65%, 60%, 59%, 58%, 57%, 56%, 55%, 54%, 53%, 52%, 51% or 50% of the total seed weight when measured or adjusted to 13% moisture.
[0085] For example, provided are soybean seeds which comprise leghemoglobin in an amount disclosed herein and which have a stachyose content of less than or less than about 4%, 3.5%, 3%, 2.5%, 2%, 1.5%, 1%, 0.5%, 0.4%, 0.3%, 0.2%, or 0.1% and at least or at least about 0%, 0.01%, 0.05%, 0.06%, 0.07%, 0.08% or 0.09%, of the total seed weight when measured or adjusted to 13% moisture.
[0086] For example, provided are soybean seeds which comprise leghemoglobin in an amount disclosed herein and which have a raffinose content of less than or less than about 2%, 1.5%, 1.4%, 1.3%, 1.2%, 1.1%, 1%, 0.9%, 0.8%, 0.7%, 0.6%, 0.5%, 0.4%, 0.3%, 0.2%, or 0.1% and at least or at least about 0%, 0.01%, 0.05%, 0.06%, 0.07%, 0.08% or 0.09% of the total seed weight when measured or adjusted to 13% moisture.
[0087] As used herein, "soy protein composition" refers to food ingredients for humans or animals which contain soy proteins. In certain embodiments, the composition is a human food composition. In certain embodiments, the human food composition is a composition selected from the group consisting of soybean meal; soyflour; defatted soyflour; soymilk; spray-dried soymilk; soy protein concentrate; texturized soy protein concentrate; hydrolyzed soy protein; soy protein isolate; spray-dried tofu; soy meat analog; soy cheese analog; and soy coffee creamer.
[0088] In some embodiments, a soybean isolate or soy protein isolate is provided which comprises at least 0.1%, 0.2%, 0.3%, 0.4%, 0.5%, 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9% or 10% and less than 25%, 20%, 15%, 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3%, 2% or 1% leghemoglobin by weight of total protein, wherein at least about 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 80%, 85%, 90% or 95% and less than 99.9%, 99%, 98%, 97%, 96%, 95%, 90%, 85%, 80%, or 75% of the leghemoglobin is hemelated with an iron group.
[0089] In certain embodiments, plants producing seed comprising leghemoglobin as described herein comprise at least about a 10%, 15%, 20%, 25%, 30%, 35%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or 500% and less than about a 1000%, 500%, 100%, 90%, 80%, 70%, 60%, or 50% increase in the amount of one or more essential amino acids as compared to a control seed.
[0090] As used herein, "percent increase" refers to a change or difference expressed as a fraction of the control value, e.g. {[modified/transgenic/test value (%)-control value (%)]/control value (%)}.times.100%=percent change, or {[value obtained in a first location (%)-value obtained in second location (%)]/value in the second location (%)}.times.100=percent change.
[0091] In certain embodiments, the one or more essential amino acids is one or more of methionine, cystine, tryptophan, threonine, and lysine, or any combination thereof.
[0092] In certain embodiments, method, plants and seeds are provided which further comprise at least one additional modification increasing the total protein in the seed as compared to a control seed (e.g., seed not comprising the at least one modification). In certain embodiments, the introduced modification increases the protein content in the soybean seed comprising leghemoglobin to at least about a 1%, 1.5%, 2%, 2.5%, 3%, 3.5%, 4%, 5%, 10%, or 15% and less than 20%, 15%, 10%, 9%, 8%, 7%, 6%, or 5% percentage point increase in total protein measured on a dry weight basis, as compared to a control seed.
[0093] In certain embodiments, the method further comprises introducing at least one modification decreasing the raffinose family oligosaccharides (RFO) content in the seed. In certain embodiments, the modification comprises a decrease in the expression and/or activity of a raffinose synthase. In certain embodiments, the modification comprises a decrease in the expression and/or activity of raffinose synthase 2 (RS2) and/or raffinose synthase 4 (RS4). In certain embodiments, the soybean seed comprises at least a 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 99% decrease in expression of RS2, RS4, or RS2 and RS4, as compared to a control seed. In certain embodiments, the seed comprises less than about 6%, 5.5%, 5%, 4.5%, 4%, 3.5%, 3%, 2.5%, 2%, 1.5%, 1%, or 0.5% RFO content on a dry weight basis. In certain embodiments, the introduced modification decreases RFO content by at least about a 1%, 1.5%, 2%, 2.5%, 3%, 3.5%, 4%, 5%, 10% or 15% and less than 20%, 15%, 10%, 9%, 8%, 7%, 6%, or 5% percentage point increase in total protein measured on a dry weight basis, as compared to a control seed
[0094] In certain embodiments, the method comprises: (a) providing a guide RNA, at least one polynucleotide modification template, and at least one Cas endonuclease to a plant cell, wherein the at least one Cas endonuclease introduces a double stranded break at an endogenous gene to be modified in the plant cell, and wherein the polynucleotide modification template generates a modified gene that encodes any of the polypeptides described herein; (b) obtaining a plant from the plant cell; and (c) generating a progeny plant.
[0095] Methods and compositions are provided herein for modifying naturally-occurring polynucleotides or integrated transgenic sequences, including regulatory elements, coding sequences, and non-coding sequences. These methods and compositions are also useful in targeting nucleic acids to pre-engineered target recognition sequences in the genome. Modification of polynucleotides may be accomplished, for example, by introducing single- or double-strand breaks into the DNA molecule.
[0096] In certain embodiments, the method comprises: (a) providing a guide RNA, at least one polynucleotide modification template, and at least one Cas endonuclease to a plant cell, wherein the at least one Cas endonuclease introduces a double stranded break at an endogenous gene to be modified in the plant cell, and wherein the polynucleotide modification template generates a modified gene that encodes any of the polypeptides described herein; (b) obtaining a plant from the plant cell; and (c) generating a progeny plant.
[0097] Double-strand breaks induced by double-strand-break-inducing agents, such as endonucleases that cleave the phosphodiester bond within a polynucleotide chain, can result in the induction of DNA repair mechanisms, including the non-homologous end-joining pathway, and homologous recombination. Endonucleases include a range of different enzymes, including restriction endonucleases (see e.g. Roberts et al., (2003) Nucleic Acids Res 1:418-20), Roberts et al., (2003) Nucleic Acids Res 31:1805-12, and Belfort et al., (2002) in Mobile DNA II, pp. 761-783, Eds. Craigie et al., (ASM Press, Washington, D.C.)), meganucleases (see e.g., WO 2009/114321; Gao et al. (2010) Plant Journal 1:176-187), TAL effector nucleases or TALENs (see e.g., US20110145940, Christian, M., T. Cermak, et al. 2010. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186(2): 757-61 and Boch et al., (2009), Science 326(5959): 1509-12), zinc finger nucleases (see e.g. Kim, Y. G., J. Cha, et al. (1996). "Hybrid restriction enzymes: zinc finger fusions to FokI cleavage"), and CRISPR-Cas endonucleases (see e.g. WO2007/025097 application published Mar. 1, 2007).
[0098] Once a double-strand break is induced in the genome, cellular DNA repair mechanisms are activated to repair the break. There are two DNA repair pathways. One is termed nonhomologous end-joining (NHEJ) pathway (Bleuyard et al., (2006) DNA Repair 5:1-12) and the other is homology-directed repair (HDR). The structural integrity of chromosomes is typically preserved by NHEJ, but deletions, insertions, or other rearrangements (such as chromosomal translocations) are possible (Siebert and Puchta, 2002, Plant Cell 14:1121-31; Pacher et al., 2007, Genetics 175:21-9. The HDR pathway is another cellular mechanism to repair double-stranded DNA breaks and includes homologous recombination (HR) and single-strand annealing (SSA) (Lieber. 2010 Annu. Rev. Biochem. 79:181-211).
[0099] In addition to the double-strand break inducing agents, site-specific base conversions can also be achieved to engineer one or more nucleotide changes to create one or more modifications described herein into the genome. These include for example, a site-specific base edit mediated by an C.cndot.G to T.cndot.A or an A.cndot.T to G.cndot.C base editing deaminase enzymes (Gaudelli et al., Programmable base editing of A.cndot.T to G.cndot.C in genomic DNA without DNA cleavage." Nature (2017); Nishida et al. "Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems." Science 353 (6305) (2016); Komor et al. "Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage." Nature 533 (7603) (2016):420-4.
[0100] In the methods described herein, the endogenous gene may be modified by a CRISPR associated (Cas) endonuclease, a Zn-finger nuclease-mediated system, a meganuclease-mediated system, an oligonucleobase-mediated system, or any gene modification system known to one of ordinary skill in the art.
[0101] In certain embodiments the endogenous gene is modified by a CRISPR associated (Cas) endonuclease.
[0102] Class I Cas endonucleases comprise multisubunit effector complexes (Types I, III, and IV), while Class 2 systems comprise single protein effectors (Types II, V, and VI) (Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15; Zetsche et al., 2015, Cell 163, 1-13; Shmakov et al., 2015, Molecular Cell 60, 1-13; Haft et al., 2005, Computational Biology, PLoS Comput Biol 1(6): e60; and Koonin et al. 2017, Curr Opinion Microbiology 37:67-78). In Class 2 Type II systems, the Cas endonuclease acts in complex with a guide polynucleotide.
[0103] Accordingly, in certain embodiments of the methods described herein the Cas endonuclease forms a complex with a guide polynucleotide (e.g., guide polynucleotide/Cas endonuclease complex).
[0104] As used herein, the term "guide polynucleotide", relates to a polynucleotide sequence that can form a complex with a Cas endonuclease, including the Cas endonucleases described herein, and enables the Cas endonuclease to recognize, optionally bind to, and optionally cleave a DNA target site. The guide polynucleotide sequence can be a RNA sequence, a DNA sequence, or a combination thereof (a RNA-DNA combination sequence). The guide polynucleotide may further comprise a chemically-modified base, such as, but not limited, to Locked Nucleic Acid (LNA), 5-methyl dC, 2,6-Diaminopurine, 2'-Fluoro A, 2'-Fluoro U, 2'-O-Methyl RNA, Phosphorothioate bond, linkage to a cholesterol molecule, linkage to a polyethylene glycol molecule, linkage to a spacer 18 (hexaethylene glycol chain) molecule, or 5' to 3' covalent linkage resulting in circularization.
[0105] In certain embodiments, the Cas endonuclease forms a complex with a guide polynucleotide (e.g., gRNA) that directs the Cas endonuclease to cleave the DNA target to enable target recognition, binding, and cleavage by the Cas endonuclease. The guide polynucleotide (e.g., gRNA) may comprise a Cas endonuclease recognition (CER) domain that interacts with the Cas endonuclease, and a Variable Targeting (VT) domain that hybridizes to a nucleotide sequence in a target DNA. In certain embodiments, the guide polynucleotide (e.g., gRNA) comprises a CRISPR nucleotide (crNucleotide; e.g., crRNA) and a trans-activating CRISPR nucleotide (tracrNucleotide; e.g., tracrRNA) to guide the Cas endonuclease to its DNA target. The guide polynucleotide (e.g., gRNA) comprises a spacer region complementary to one strand of the double strand DNA target and a region that base pairs with the tracrNucleotide (e.g., tracrRNA), forming a nucleotide duplex (e.g. RNA duplex).
[0106] In certain embodiments, the gRNA is a "single guide RNA" (sgRNA) that comprises a synthetic fusion of crRNA and tracrRNA. In many systems, the Cas endonuclease-guide polynucleotide complex recognizes a short nucleotide sequence adjacent to the target sequence (protospacer), called a "protospacer adjacent motif" (PAM).
[0107] The terms "single guide RNA" and "sgRNA" are used interchangeably herein and relate to a synthetic fusion of two RNA molecules, a crRNA (CRISPR RNA) comprising a variable targeting domain (linked to a tracr mate sequence that hybridizes to a tracrRNA), fused to a tracrRNA (trans-activating CRISPR RNA). The single guide RNA can comprise a crRNA or crRNA fragment and a tracrRNA or tracrRNA fragment of the type II CRISPR/Cas system that can form a complex with a type II Cas endonuclease, wherein said guide RNA/Cas endonuclease complex can direct the Cas endonuclease to a DNA target site, enabling the Cas endonuclease to recognize, optionally bind to, and optionally nick or cleave (introduce a single or double-strand break) the DNA target site.
[0108] The nucleotide sequence linking the crNucleotide and the tracrNucleotide of a single guide polynucleotide can comprise a RNA sequence, a DNA sequence, or a RNA-DNA combination sequence. In one embodiment, the nucleotide sequence linking the crNucleotide and the tracrNucleotide of a single guide polynucleotide can be at least 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99 or 100 nucleotides in length. In one embodiment, the nucleotide sequence linking the crNucleotide and the tracrNucleotide of a single guide polynucleotide can comprise a tetraloop sequence, such as, but not limiting to a GAAA tetraloop sequence.
[0109] The term "variable targeting domain" or "VT domain" is used interchangeably herein and includes a nucleotide sequence that can hybridize (is complementary) to one strand (nucleotide sequence) of a double strand DNA target site. In some embodiments, the variable targeting domain comprises a contiguous stretch of 12 to 30 nucleotides. The variable targeting domain can be composed of a DNA sequence, a RNA sequence, a modified DNA sequence, a modified RNA sequence, or any combination thereof.
[0110] The term "Cas endonuclease recognition domain" or "CER domain" (of a guide polynucleotide) is used interchangeably herein and includes a nucleotide sequence that interacts with a Cas endonuclease polypeptide. A CER domain comprises a (trans-acting) tracrNucleotide mate sequence followed by a tracrNucleotide sequence. The CER domain can be composed of a DNA sequence, a RNA sequence, a modified DNA sequence, a modified RNA sequence (see for example US20150059010A1, published 26 Feb. 2015), or any combination thereof.
[0111] A "protospacer adjacent motif" (PAM) as used herein refers to a short nucleotide sequence adjacent to a target sequence (protospacer) that is recognized (targeted) by a guide polynucleotide/Cas endonuclease system described herein. In certain embodiments, the Cas endonuclease may not successfully recognize a target DNA sequence if the target DNA sequence is not adjacent to, or near, a PAM sequence. In certain embodiments, the PAM precedes the target sequence (e.g. Cas12a). In certain embodiments, the PAM follows the target sequence (e.g. S. pyogenes Cas9). The sequence and length of a PAM herein can differ depending on the Cas protein or Cas protein complex used. The PAM sequence can be of any length but is typically 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 nucleotides long.
[0112] As used herein, the terms "guide polynucleotide/Cas endonuclease complex", "guide polynucleotide/Cas endonuclease system", "guide polynucleotide/Cas complex", "guide polynucleotide/Cas system" and "guided Cas system" "polynucleotide-guided endonuclease", and "PGEN" are used interchangeably herein and refer to at least one guide polynucleotide and at least one Cas endonuclease, that are capable of forming a complex, wherein said guide polynucleotide/Cas endonuclease complex can direct the Cas endonuclease to a DNA target site, enabling the Cas endonuclease to recognize, bind to, and optionally nick or cleave (introduce a single or double-strand break) the DNA target site. A guide polynucleotide/Cas endonuclease complex herein can comprise Cas protein(s) and suitable polynucleotide component(s) of any of the known CRISPR systems (Horvath and Barrangou, 2010, Science 327:167-170; Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15; Zetsche et al., 2015, Cell 163, 1-13; Shmakov et al., 2015, Molecular Cell 60, 1-13). In certain embodiments, the guide polynucleotide/Cas endonuclease complex is provided as a ribonucleoprotein (RNP), wherein the Cas endonuclease component is provided as a protein and the guide polynucleotide component is provided as a ribonucleotide.
[0113] Examples of Cas endonucleases for use in the methods described herein include, but are not limited to, Cas9 and Cpf1. Cas9 (formerly referred to as Cas5, Csn1, or Csx12) is a Class 2 Type II Cas endonuclease (Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15). A Cas9-gRNA complex recognizes a 3' PAM sequence (NGG for the S. pyogenes Cas9) at the target site, permitting the spacer of the guide RNA to invade the double-stranded DNA target, and, if sufficient homology between the spacer and protospacer exists, generate a double-strand break cleavage. Cas9 endonucleases comprise RuvC and HNH domains that together produce double strand breaks, and separately can produce single strand breaks. For the S. pyogenes Cas9 endonuclease, the double-strand break leaves a blunt end. Cpf1 is a Clas 2 Type V Cas endonuclease, and comprises nuclease RuvC domain but lacks an HNH domain (Yamane et al., 2016, Cell 165:949-962). Cpf1 endonucleases create "sticky" overhang ends.
[0114] Some uses for Cas9-gRNA systems at a genomic target site include, but are not limited to, insertions, deletions, substitutions, or modifications of one or more nucleotides at the target site; modifying or replacing nucleotide sequences of interest (such as a regulatory elements); insertion of polynucleotides of interest; gene knock-out; gene-knock in; modification of splicing sites and/or introducing alternate splicing sites; modifications of nucleotide sequences encoding a protein of interest; amino acid and/or protein fusions; and gene silencing by expressing an inverted repeat into a gene of interest.
[0115] The terms "target site", "target sequence", "target site sequence, "target DNA", "target locus", "genomic target site", "genomic target sequence", "genomic target locus" and "protospacer", are used interchangeably herein and refer to a polynucleotide sequence such as, but not limited to, a nucleotide sequence on a chromosome, episome, a locus, or any other DNA molecule in the genome (including chromosomal, chloroplastic, mitochondrial DNA, plasmid DNA) of a cell, at which a guide polynucleotide/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave. The target site can be an endogenous site in the genome of a cell, or alternatively, the target site can be heterologous to the cell and thereby not be naturally occurring in the genome of the cell, or the target site can be found in a heterologous genomic location compared to where it occurs in nature. As used herein, terms "endogenous target sequence" and "native target sequence" are used interchangeable herein to refer to a target sequence that is endogenous or native to the genome of a cell and is at the endogenous or native position of that target sequence in the genome of the cell. An "artificial target site" or "artificial target sequence" are used interchangeably herein and refer to a target sequence that has been introduced into the genome of a cell. Such an artificial target sequence can be identical in sequence to an endogenous or native target sequence in the genome of a cell but be located in a different position (i.e., a non-endogenous or non-native position) in the genome of a cell. An "altered target site", "altered target sequence", "modified target site", "modified target sequence" are used interchangeably herein and refer to a target sequence as disclosed herein that comprises at least one alteration when compared to non-altered target sequence. Such "alterations" include, for example: (i) replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, or (iv) any combination of (i)-(iii).
[0116] A "polynucleotide modification template" is also provided that comprises at least one nucleotide modification when compared to the nucleotide sequence to be edited. For example, a modification in the endogenous gene corresponding to SEQ ID NO: 1 to induce an amino substitution in the encoded polypeptide. A nucleotide modification can be at least one nucleotide substitution, addition, deletion, or chemical alteration. Optionally, the polynucleotide modification template can further comprise homologous nucleotide sequences flanking the at least one nucleotide modification, wherein the flanking homologous nucleotide sequences provide sufficient homology to the desired nucleotide sequence to be edited.
[0117] In certain embodiments of the methods disclosed herein, a polynucleotide of interest is inserted at a target site and provided as part of a "donor DNA" molecule. As used herein, "donor DNA" is a DNA construct that comprises a polynucleotide of interest to be inserted into the target site of a Cas endonuclease. The donor DNA construct further comprises a first and a second region of homology that flank the polynucleotide of interest. The first and second regions of homology of the donor DNA share homology to a first and a second genomic region, respectively, present in or flanking the target site of the cell or organism genome. The donor DNA can be tethered to the guide polynucleotide. Tethered donor DNAs can allow for co-localizing target and donor DNA, useful in genome editing, gene insertion, and targeted genome regulation, and can also be useful in targeting post-mitotic cells where function of endogenous HR machinery is expected to be highly diminished (Mali et al., 2013, Nature Methods Vol. 10: 957-963). The amount of homology or sequence identity shared by a target and a donor polynucleotide can vary and includes total lengths and/or regions.
[0118] The process for editing a genomic sequence at a Cas9-gRNA double-strand-break site with a modification template generally comprises: providing a host cell with a Cas9-gRNA complex that recognizes a target sequence in the genome of the host cell and is able to induce a double-strand-break in the genomic sequence, and at least one polynucleotide modification template comprising at least one nucleotide alteration when compared to the nucleotide sequence to be edited. The polynucleotide modification template can further comprise nucleotide sequences flanking the at least one nucleotide alteration, in which the flanking sequences are substantially homologous to the chromosomal region flanking the double-strand break. Genome editing using double-strand-break-inducing agents, such as Cas9-gRNA complexes, has been described, for example in US20150082478 published on 19 Mar. 2015, WO2015026886 published on 26 Feb. 2015, WO2016007347 published 14 Jan. 2016, and WO2016025131 published on 18 Feb. 2016.
[0119] To facilitate optimal expression and nuclear localization for eukaryotic cells, the gene comprising the Cas endonuclease may be optimized as described in WO2016186953 published 24 Nov. 2016, and then delivered into cells as DNA expression cassettes by methods known in the art. In certain embodiments, the Cas endonuclease is provided as a polypeptide. In certain embodiments, the Cas endonuclease is provided as a polynucleotide encoding a polypeptide. In certain embodiments, the guide RNA is provided as a DNA molecule encoding one or more RNA molecules. In certain embodiments, the guide RNA is provided as RNA or chemically-modified RNA. In certain embodiments, the Cas endonuclease protein and guide RNA are provided as a ribonucleoprotein complex (RNP).
[0120] In certain embodiments, methods for modifying the endogenous by a zinc-finger-mediated genome editing process are provided. The zinc-finger-mediated genome editing process for editing a chromosomal sequence includes for example: (a) introducing into a cell at least one nucleic acid encoding a zinc finger nuclease that recognizes a target sequence in the chromosomal sequence and is able to cleave a site in the chromosomal sequence, and, optionally, (i) at least one donor polynucleotide that includes a sequence for integration flanked by an upstream sequence and a downstream sequence that exhibit substantial sequence identity with either side of the cleavage site, or (ii) at least one exchange polynucleotide comprising a sequence that is substantially identical to a portion of the chromosomal sequence at the cleavage site and which further comprises at least one nucleotide change; and (b) culturing the cell to allow expression of the zinc finger nuclease such that the zinc finger nuclease introduces a double-stranded break into the chromosomal sequence, and wherein the double-stranded break is repaired by (i) a non-homologous end-joining repair process such that an inactivating mutation is introduced into the chromosomal sequence, or (ii) a homology-directed repair process such that the sequence in the donor polynucleotide is integrated into the chromosomal sequence or the sequence in the exchange polynucleotide is exchanged with the portion of the chromosomal sequence.
[0121] A zinc finger nuclease includes a DNA binding domain (i.e., zinc finger) and a cleavage domain (i.e., nuclease). The nucleic acid encoding a zinc finger nuclease may include DNA or RNA. Zinc finger binding domains may be engineered to recognize and bind to any nucleic acid sequence of choice. See, for example, Beerli et al. (2002) Nat. Biotechnol. 20:135-141; Pabo et al. (2001) Ann. Rev. Biochem. 70:313-340; Choo et al. (2000) Curr. Opin. Struct. Biol. 10:411-416; and Doyon et al. (2008) Nat. Biotechnol. 26:702-708; Santiago et al. (2008) Proc. Natl. Acad. Sci. USA 105:5809-5814; Urnov, et al., (2010) Nat Rev Genet. 11(9):636-46; and Shukla, et al., (2009) Nature 459 (7245):437-41. An engineered zinc finger binding domain may have a novel binding specificity compared to a naturally-occurring zinc finger protein. As an example, the algorithm of described in U.S. Pat. No. 6,453,242 may be used to design a zinc finger binding domain to target a preselected sequence. Nondegenerate recognition code tables may also be used to design a zinc finger binding domain to target a specific sequence (Sera et al. (2002) Biochemistry 41:7074-7081). Tools for identifying potential target sites in DNA sequences and designing zinc finger binding domains may be used (Mandell et al. (2006) Nuc. Acid Res. 34:W516-W523; Sander et al. (2007) Nuc. Acid Res. 35:W599-W605).
[0122] An exemplary zinc finger DNA binding domain recognizes and binds a sequence having at least about 80% sequence identity with the desired target sequence. In other embodiments, the sequence identity may be about 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100%.
[0123] A zinc finger nuclease also includes a cleavage domain. The cleavage domain portion of the zinc finger nucleases may be obtained from any endonuclease or exonuclease. Non-limiting examples of endonucleases from which a cleavage domain may be derived include, but are not limited to, restriction endonucleases and homing endonucleases. See, for example, 2010-2011 Catalog, New England Biolabs, Beverly, Mass.; and Belfort et al. (1997) Nucleic Acids Res. 25:3379-3388. Additional enzymes that cleave DNA are known (e.g., 51 Nuclease; mung bean nuclease; pancreatic DNase I; micrococcal nuclease; yeast HO endonuclease). One or more of these enzymes (or functional fragments thereof) may be used as a source of cleavage domains.
[0124] In certain embodiments of the methods described herein the endogenous gene is modified by using "custom" meganucleases produced to modify plant genomes (see e.g., WO 2009/114321; Gao et al. (2010) Plant Journal 1:176-187). The term "meganuclease" generally refers to a naturally-occurring homing endonuclease that binds double-stranded DNA at a recognition sequence that is greater than 12 base pairs and encompasses the corresponding intron insertion site. Naturally-occurring meganucleases can be monomeric (e.g., I-SceI) or dimeric (e.g., I-CreI). The term meganuclease, as used herein, can be used to refer to monomeric meganucleases, dimeric meganucleases, or to the monomers which associate to form a dimeric meganuclease.
[0125] Naturally-occurring meganucleases, for example, from the LAGLIDADG family, have been used to effectively promote site-specific genome modification in plants, yeast, Drosophila, mammalian cells and mice. Engineered meganucleases such as, for example, LIG-34 meganucleases, which recognize and cut a 22 basepair DNA sequence found in the genome of Zea mays (maize) are known (see e.g., US 20110113509).
[0126] In certain embodiments of the methods described herein the endogenous gene is modified by using TAL endonucleases (TALEN). TAL (transcription activator-like) effectors from plant pathogenic Xanthomonas are important virulence factors that act as transcriptional activators in the plant cell nucleus, where they directly bind to DNA via a central domain of tandem repeats. A transcription activator-like (TAL) effector-DNA modifying enzymes (TALE or TALEN) are also used to engineer genetic changes. See e.g., US20110145940, Boch et al., (2009), Science 326(5959): 1509-12. Fusions of TAL effectors to the FokI nuclease provide TALENs that bind and cleave DNA at specific locations. Target specificity is determined by developing customized amino acid repeats in the TAL effectors.
[0127] In certain embodiments of the methods described herein the endogenous gene is modified by using base editing, such as an oligonucleobase-mediated system. In addition to the double-strand break inducing agents, site-specific base conversions can also be achieved to engineer one or more nucleotide changes to create one or more EMEs described herein into the genome. These include for example, a site-specific base edit mediated by a C.cndot.G to T.cndot.A or an A.cndot.T to G.cndot.C base editing deaminase enzymes (Gaudelli et al., Programmable base editing of A.cndot.T to G.cndot.C in genomic DNA without DNA cleavage." Nature (2017); Nishida et al. "Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems." Science 353 (6305) (2016); Komor et al. "Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage." Nature 533 (7603) (2016):420-4. Catalytically dead dCas9 fused to a cytidine deaminase or an adenine deaminase protein becomes a specific base editor that can alter DNA bases without inducing a DNA break. Base editors convert C->T (or G->A on the opposite strand) or an adenine base editor that would convert adenine to inosine, resulting in an A->G change within an editing window specified by the gRNA.
[0128] Further provided are methods of plant breeding comprising crossing any of the soybean plants described herein with a second plant to produce a progeny seed comprising at least one modification described herein. In certain embodiments, a plant is produced from the progeny seed.
[0129] The following are examples of specific embodiments of some aspects of the invention. The examples are offered for illustrative purposes only and are not intended to limit the scope of the invention in any way.
Example 1: Expression of Soybean Leghemoglobin Protein in Soybean Seeds
[0130] A soybean leghemoglobin gene (Glyma.20g191200) was identified in the soybean genome. The gene contains 4 exons, with its CDS (SEQ ID NO: 1) encoding a leghemoglobin peptide (SEQ ID NO: 2). As shown in FIG. 1, the soybean leghemoglobin was expressed with no signal peptide in soybean seeds. In addition, several protein targeting signal sequences were used to target the leghemoglobin into protein storage vacuole in soybean seeds (Table 2). The beta-conglycinin alpha' SPP is encoded by the nucleotides from position 1 to position 195 of SEQ ID NO: 3, with its corresponding peptide sequences defined by position 1 to position 65 of SEQ ID NO: 4. The lectin SP is encoded by the nucleotides from position 1 to position 105 of SEQ ID NO: 5, with its corresponding peptide sequence defined by position 1 to position 35 of SEQ ID NO: 6. The GY1 SP is encoded by the nucleotides from position 1 to position 66 of SEQ ID NO: 7, with its corresponding peptide sequence defined by position 1 to position 22 of SEQ ID NO: 8. A strong seed specific promoter, such as a beta-conglycinin promoter (SEQ ID NO: 9) or a glycinin promoter (SEQ ID NO: 10) was used to drive the expression of the leghemoglobin. The Rubisco small subunit (Rubisco SSU) plastid targeting sequences were also used to target the leghemoglobin protein to plastids. The Rubisco SSUSP plastid targeting sequence is encoded by the nucleotide sequence from position 1 to position 165 of SEQ ID NO: 31, with the corresponding peptide targeting sequence at position 1 to position 55 of SEQ ID NO: 32. The leghemoglobin coding sequence is from position 166 to position 603 of SEQ ID NO: 31 and the corresponding peptide form position 56 to position 200 of SEQ ID NO: 32. These expression vectors were introduced into soybean plants by Ochrobactrum-mediated or Agrobacteria-mediated soybean embryonic axis transformation, the former being described in US Patent Publication No. 2018/0216123. The results are described in Example 7.
TABLE-US-00002 TABLE 2 Expression of Leghemoglobin by Protein Targeting in Soybean Seeds. Leghemoglobin (LH) with or without Signal Peptide (Vector Nucleotide Peptide name) SEQ ID NOs: SEQ ID NOs: No signal peptide::LH SEQ ID NO: 1 SEQ ID NO: 2 Beta-conglycinin alpha' SPP::LH SEQ ID NO: 3 SEQ ID NO: 4 Lectin SP::LH SEQ ID NO: 5 SEQ ID NO: 6 GY1 SP::LH::GY1-Basic subunit SEQ ID NO: 7 SEQ ID NO: 8 Rubisco SSUSP::LH SEQ ID NO: 31 SEQ ID NO: 32
Example 2: Improvement of Soybean Leghemoglobin Expression Level by Porphyrin Pathway Engineering--Glutamyl-tRNA Reductase and Ferrochelatase
[0131] To improve the soybean leghemoglobin expression level in seeds, a porphyrin pathway engineering approach was employed. There are at least nine enzymatic steps for the porphyrin pathway leading to the heme biosynthesis. Among them, as shown in FIG. 2, the glutamyl-tRNA reductase (glyma.04g089800) and ferrochelatase (glyma.04g050400) were tested for increasing heme production, to facilitate higher leghemoglobin accumulation and heme loading in soybean seeds. For this purpose, four additional soybean vectors were made, each of them contained the expression of glutamyl-tRNA reductase (SEQ ID NO: 11, 12) and ferrochelatase (SEQ ID NOs: 13, 14), in addition to the leghemoglobin expression cassettes in Example 1. The two biosynthetic genes were driven by strong seed specific promoters, such as Phaseolus vulgaris Phaseolin promoter (SEQ ID NO: 15) or Brassica napus Napin Promoter (SEQ ID NO: 16). In these four vectors, the expression cassettes of these two biosynthetic genes were stacked molecularly with the four expression cassettes of the leghemoglobin with or without different signal peptide targeting sequences. These expression vectors were introduced into soybean plants by Ochrobactrum-mediated soybean embryonic axis transformation as described in US Patent Publication No. 2018/0216123. The results are described in Example 7.
Example 3: Improvement of Soybean Leghemoglobin Expression Level by Porphyrin Enzyme Modifications or Expression
[0132] A similar technical approach to the methods described in Example 2 is used to regulate other enzymatic steps for the porphyrin pathway, such as glutamate-1-semialdehyde 2, 1-aminomutase, am inolevulinate dehydratase, hydroxymethylbilane synthase, urophorphyrinogen III synthase, urophorphyrinogen decarboxylase, coporphyrinogen III oxidase, and protoporphyrinogen oxidase. Examples of soybean genes for the porphyrin pathway that are used are listed in Table 3. Overexpressing these native metabolic enzyme genes in soybean seeds is achieved by transformation of soybean with a recombinant construct comprising a coding sequence for these polypeptides, operably linked to regulatory sequences that provide for expression in soybean seeds. Secondly, increased expression of these enzymes is achieved through gene editing. Feedback sensitive regulatory domains of these enzymes are identified and removed or inactivated by gene editing truncations, deletions, substitutions or insertions. It is expected that enhanced heme content of the leghemoglobin protein produced in soybean seeds modified to produce increased leghemoglobin protein complex is achieved. The heme biosynthetic enzymes which are modified to be feedback-insensitive or are otherwise modified or edited to enhance enzyme expression, stability or activity are expressed in soybean seeds to further increase heme production, enabling higher leghemoglobin accumulation and heme loading in soybean seeds. Specifically, Glutamyl-tRNA reductase (GTR) enzyme activity is under combinatorial, post-translational control mediated by the proteins FLUORESCENT IN BLUE LIGHT (FLU), Glutamyl-tRNA reductase-binding protein (GBP), chloroplast signal particle 43 (SRP43) (Table 4). Altered expression of a single or any combination of these three proteins achieved by gene editing, seed-preferred over-expression or RNA interference is expected to achieve higher level of heme-containing leghemoglobin by increasing heme-biosynthetic activity in developing seeds.
TABLE-US-00003 TABLE 3 The soybean genes in the porphyrin pathway Enzyme name Gene Model Name Glutamyl-tRNA reductase Glyma.02G218300 Glyma.04G089800 Glyma.06G091600 Glyma.07G184700 Glyma.08G064700 Glyma.14G185700 glutamate-1-semialdehyde 2,1-aminomutase Glyma.04G002900 Glyma.06G002900 Glyma.14G221900 aminolevulinate dehydratase (HEMB1) Glyma.04G247700 Glyma.06G115000 hydroxymethylbilane synthase (HEMC) Glyma.01G227400 Glyma.11G015400 Glyma.11G094700 Glyma.12G021100 urophorphyrinogen III synthase Glyma.04G037000 Glyma.06G037300 urophorphyrinogen decarboxylase Glyma.11G235400 Glyma.12G229700 Glyma.13G269900 Glyma.18G021500 coporphyrinogen III oxidase (HEMF, CPOX) Glyma.14G003200 protoporphyrinogen oxidase (PPOX) Glyma.10G138600 Glyma.02G007200 Glyma.19G245900 Glyma.08G173600 ferrochelatase Glyma.04G050400 Glyma.04G205600 Glyma.05G197600 Glyma.06G051100 Glyma.06G159900 Glyma.08G005000
TABLE-US-00004 TABLE 4 The soybean genes encoding proteins that regulate Glutamyl-tRNA reductase activity Enzyme name Gene Model Name Glutamyl-tRNA reductase-binding protein Glyma.08G222600 chloroplast signal particle 43 Glyma.11G097200 FLUORESCENT IN BLUE LIGHT Glyma.16G010200 Glyma.07G041700
Example 4: Genome Engineering of the Leghemoglobin Gene into the Native Soybean Glycinin Gene Loci
[0133] With the CRISPR/Cas9 system, we designed specific gRNAs (GM-GY-CR1, SEQ ID NO: 17; GM-GY-CR2, SEQ ID NO: 18; and GM-GY-CR3, SEQ ID NO: 19) to target the Glycinin 1 (GY1) gene (glyma.03g163500, SEQ ID NO: 20 for nucleotide sequences, SEQ ID NO:21 for peptide sequences). The GM-GY1-CR1 was designed to target a site near the beginning of the exon1 of the pro-glycinin 1 protein. The GM-GY1-CR2 was designed to target the junction between the acidic subunit (amino acid #1 to #310 in SEQ ID NO: 21) and basic subunit (amino acid #311 to #495 in SEQ ID NO: 21) of the pro-glycinin1. The GM-GY1-CR3 was designed to target the beginning of the 3' UTR of glycinin 1 gene. As shown in FIGS. 3 and 4, the binary vectors contained either CR1/CR2 or CR1/CR3 gRNA combinations and their corresponding donor DNA templates (SEQ ID NO: 22 and SEQ ID NO: 23). The homology recombination (HR) fragments were used to flank the leghemoglobin/GY1 sequences to facilitate the homology-mediated recombination process. The CR1 or CR2 or CR3 gRNA target sites were also used to flank the donor DNAs to enable them to be excised from the binary vectors for double strand break repair process. These sequences are defined in Table 5.
TABLE-US-00005 TABLE 5 The nucleotide sequences of HR fragments and CR Cut sites in donor DNA templates Location in SEQ ID NOs HR1 Position 34 to position 1032 of SEQ ID NO: 22 and of SEQ ID NO: 23 HR2 Position 2104 to position 3531 of SEQ ID NO: 22 HR5 Position 2108 to position 3531 of SEQ ID NO: 23 CR1 Cut Site Position 1 to position 22 of SEQ ID NO: 22 and of SEQ ID NO: 23 CR2 Cut Site Position 3543 to position 3565 of SEQ ID NO: 22 CR3 Cut site Position 3543 to position 3565 of SEQ ID NO: 23
[0134] The binary vectors were introduced into soybean plants by Agrobacterium-mediated soybean embryonic axis transformation. With site-specific integration of the donor DNA by homology-mediated double strand break DNA repair process, a genome editing variants of the glycinin 1 with the soybean leghemoglobin were created by (i) replacing the genomic sequences encoding the acidic subunit at the native Glycinin 1 gene locus or (ii) replacing the genomic sequences encoding the entire glycinin1 protein at the native Glycinin 1 gene locus. In total, 1452 T0 plants were generated for the CR1/CR3 design (FIG. 3) to replace the entire glycinin1 gene with the leghemoglobin coding sequences at the native glycinin1 gene locus. We used two PCR analyses to identify the perfect gene integration events, one is at the 5' and another at the 3' region of the glycinin1 locus. For the 1452 T0 plants, 10 potential 2.times.HDR perfect integration events were identified at T0 plants. Based on the intensity of the PCR products, we classified them into three categories: strong (4 events), medium (3 events) and weak (3 events). Of these 10 events, we performed sequencing analysis of the PCR products, two of the 10 events (1 strong and 1 weak event) had SNP variations from double strand break repair process, so we did not proceed with those two events further. T1 seeds were harvest from all remaining 8 positive events. We proceeded with leghemoglobin quantification of the T1 seeds from our top six events based on our T0 plant analyses (3 strong events: 198A, 315A, 956A and 3 medium events: 407A, 419A and 628A), the 315A events gave the highest globin accumulation (1.16% of the total protein of the seed; dry weight basis) in T1 seeds.
TABLE-US-00006 TABLE 5A Quantitative Mass Spectrometry analysis of single wildtype (yellow) and red soybeans harvested from segregating T0 plants expressing the LegHemoglobin construct targeted to the GY1 Locus Soluble LegHemoglobin LegHemoglobin protein/total protein/soluble protein/total SAMPLE ID Protein (wt %) protein (wt %) protein (wt %) GV.GM7MDH.005.198)X 70.69 0.00 0.00 (yellow) GV.GM7MDH.005.198)X 52.79 0.00 0.00 yellow GV.GM7MDH.005.198)X 67.12 0.79 0.53 GV.GM7MDH.005.198)X 46.97 0.43 0.20 GV.GM7MDH.005.198)X 38.60 1.80 0.69 GV.GM7MDH.005.198)X 57.87 0.81 0.47 GV.GM7MDH.005.198)X 60.76 0.64 0.39 GV.GM7MDH.005.198)X 37.45 1.78 0.67 GV.GMWKWK.004.315)X 48.15 0.00 0.00 (yellow) GV.GMWKWK.004.315)X 50.16 2.32 1.16 GV.GMWKWK.004.315)X 60.51 1.44 0.87 GV.GMWKWK.004.315)X 49.13 1.09 0.54 GV.GM7MDH.008.956)X 50.07 0.00 0.00 (yellow) GV.GM7MDH.008.956)X 47.90 0.00 0.00 (yellow) GV.GM7MDH.008.956)X 54.29 0.08 0.05 GV.GM7MDH.008.956)X 47.83 0.38 0.18 GV.GM7MDH.008.956)X 63.28 0.66 0.42 GV.GM7MDH.008.956)X 65.73 0.11 0.07 GV.GM7MDH.008.956)X 59.75 0.56 0.34 GV.GM7MDH.008.956)X 99.58 0.01 0.01 GV.GMWKWK.001.407)X 70.43 0.00 0.00 (yellow) GV.GMWKWK.001.407)X 57.54 0.00 0.00 (yellow) GV.GMWKWK.001.407)X 52.55 0.28 0.15 GV.GMWKWK.001.407)X 65.06 0.44 0.29 GV.GMWKWK.001.407)X 66.86 0.17 0.12 GV.GMWKWK.001.407)X 63.51 0.47 0.30 GV.GMWKWK.001.407)X 68.22 0.31 0.21 GV.GMWKWK.001.407)X 59.94 0.51 0.31 GV.GMWKWK.001.419)X 76.57 0.00 0.00 (yellow) GV.GMWKWK.001.419)X 60.10 0.00 0.00 (yellow) GV.GMWKWK.001.419)X 50.41 0.92 0.46 GV.GMWKWK.001.419)X 53.04 0.39 0.20 GV.GMWKWK.001.419)X 60.99 0.36 0.22 GV.GMWKWK.001.419)X 61.84 0.59 0.36 GV.GMWKWK.001.419)X 50.12 0.39 0.19 GV.GMWKWK.001.419)X 48.17 0.47 0.23 GV.GM7MDH.001.628)X 67.58 0.00 0.00 (yellow) GV.GM7MDH.001.628)X 61.49 0.00 0.00 (yellow) GV.GM7MDH.001.628)X 57.87 0.36 0.21 GV.GM7MDH.001.628)X 49.28 0.17 0.08 GV.GM7MDH.001.628)X 51.04 0.48 0.25 GV.GM7MDH.001.628)X 69.92 0.34 0.24 GV.GM7MDH.001.628)X 57.20 0.53 0.30 GV.GM7MDH.001.628)X 74.19 0.29 0.22
[0135] We planted T1 seeds from the same six events and performed the same PCR molecular analyses on those T1 plants. In these analyses, 2.times.HDR perfect integration T1 plants can only be consistently confirmed in three of the six events analyzed (198A, 315A, 628A), For the 956A event, only one 2.times.HDR plant can be detected from 37 T1 plants screened. For the other two events (407A and 419A), we cannot detect any 2.times.HDR PCR product, indicating that the 2.times.HDR signals from their T0 plant analyses were not transmitted to T1 plants, probably due to the chimeric nature in the transformation process. These two events were re-classified as random integrated transgenic events at T1 plants. T2 seeds will be harvested for 2.times.HDR prefect integrated events, as well as those random integrated transgenic events. The leghemoglobin level will be analyzed in all homozygous T2 seeds to compare the leghemoglobin expression level at the glycinin1 native locus to the random transgenic loci. It is expected that leghemoglobin levels will be doubled in the perfect integrated events as compared to the leghemoglobin level in T1 seeds, reaching about 2.3% or more of the total seed protein per dry weight basis.
Example 5: Genome Engineering of the Leghemoglobin Gene into Other Native Soybean Seed-Storage Protein Gene Loci
[0136] Other seed storage proteins, such as other glycinin proteins or conglycinin proteins, are shown in Tables 6 and 7. The genes encoding these storage proteins are used as the gene editing targets for soybean leghemoglobin over-expression in soybean seeds as described in this example.
TABLE-US-00007 TABLE 6 Expression profiling of glycinin 1 (bold) and other putative glycinin family members in soybean. one pod pod flow- cm shell shell seed seed seed seed seed seed seed nod- young_leaf er pod 10DAF 14DAF 10DAF 14DAF 21DAF 25DAF 28DAF 35DAF 42DAF root ule Glyma03g32020 0 0 0 0 0 2 2 12 26498 72276 218636 210908 0 0 Glyma.03g163500 0 1 0 1 0 0 0 5 34953 87714 288053 298305 0 0 Glyma.10g037100 0 0 0 0 0 0 1 3 11410 29176 166962 249297 0 0 Glyma.13g123500 0 0 0 0 0 0 0 0 5591 15500 92982 130315 0 0 Glyma.19g164800 0 0 0 0 0 0 1 10 62 48 225 186 0 0 Glyma.19g164900 1 0 0 0 0 0 0 0 883 2263 10523 12595 2 0
TABLE-US-00008 TABLE 7 Expression level of 7 .beta.-conglycinin isoforms in soybean seeds 30 or 50 days after flowering. .beta.-conglycinin Expression level measured by RNAseq Glyma.20g148200 (.beta.) 19251 (30DAF) Glyma.20g148300 (.alpha.) 67117 (30DAF) Glyma.20g148400 (.alpha.) 91647 (30DAF) Glyma.20g146200 (.beta.) 7068 (30DAF) Glyma.10g246300 (.alpha.') 86918 (30DAF) Glyma.10g246500 (.alpha.') 20492 (50DAF) Glyma.10g246400 (.alpha.) No/low expression 6 (30DAF)
[0137] Specific gRNAs are designed for these genes following the protocol in example 4. The binary vectors for each gene target are introduced into soybean plants by Agrobacterium-mediated soybean embryonic axis transformation. With site-specific integration of the donor DNA by homology-mediated double strand break DNA repair process, genome editing variants of seed-storage protein genes with the soybean leghemoglobin replacing the coding sequences are created for each seed-storage protein gene, alone or in combination with GLY1 or other seed storage protein genes. Plants are grown in the greenhouse. We expect the harvested T.sub.1 seeds to contain a leghemoglobin amount of at least 1% of total seed protein or higher.
Example 6: Improvement of Soybean Leghemoglobin Expression Level in Soybean Seeds by Protein Rebalancing
[0138] Glycinin and conglycinin are two major classes of seed storage proteins in soybean seeds. In soybean seeds, .beta.-conglycinin, the abundant 7S globulin storage protein, and glycinin consist of about 21% and 33% of total protein content, respectively (Utsumi et al., 1997). Total soybean protein content did not change after silencing a and .alpha.' subunits of .beta.-conglycinin by RNAi (Kinney et al., 2001). The resulting engineered seeds accumulated more glycinin that accounts for more than 50% of total seed protein, which compensated for the missing .beta.-conglycinin in the engineered seeds. .beta.-conglycinin consists of 3 isoforms, .alpha., .alpha.' and .beta.. If desirable, the .beta.-conglycinin gene family (6 to 7 genes for the alpha', alpha and beta subunits) can be eliminated with gene cluster dropouts or frameshift knockout mutations by CRISPR/Cas editing, then channel more protein synthesis resource to leghemoglobin production in soybean seeds.
[0139] As an example, gRNAs were designed to knockout 6 putative .beta.-conglycinin isoforms by Cas9/gRNA editing to rebalance the proteome to glycinin. Seven .beta.-conglycinin candidates including 3.alpha., 2.alpha.' and 2.beta. isoforms were identified. Except for Glyma.10g246400 (a) and Glyma.20g146200 (13), all other isoforms show relatively high expression level at 30 or 50 days after flowering (DAF) in soybean seeds (Table 7).
[0140] Four gRNAs were used to delete 6 of 7 .beta.-conglycinin isoforms. The GM-CONG-gRNA1 (SEQ ID NO: 24) and GM-CONG-gRNA2 (SEQ ID NO: 25) were used to dropout the conglycinin cluster on chromosome 20 (Gm20); the GM-CONG-gRNA3 (SEQ ID NO: 26) and GM-CONG-gRNA4 (SEQ ID NO: 27) were used to dropout the conglycinin cluster on chromosome 10 (Gm10), as illustrated in FIG. 5.
[0141] T2 homozygous seeds from the conglycinin Gm10 locus dropout experiment were generated. Seed protein analyses was conducted by SDS-PAGE Coomassie Blue gel staining analyses (FIG. 6). No alpha' subunits of conglycinin proteins can be detected in those T2 homozygous seeds from the Gm10 locus dropout variants, demonstrating complete removal of the conglycinin alpha' subunit proteins in soybean seeds, in agreement with the complete removal of their genes from soybean genome. The total protein content of these T2 seeds did not change as compared to wild type seeds, indicating other soybean proteins are compensating for the loss of conglycinin alpha' subunit proteins in these editing variants. For the second editing experiment, the T2 seeds from the Gm20 locus dropout were analyzed by protein gel analyses (FIG. 7). The results indicated that the conglycinin alpha subunit proteins had been completely removed in soybean seeds of the homozygous dropout plants. The data also indicated that the conglycinin beta subunit protein was also reduced in this dropout variant due to the elimination of Glyma.20g148200 gene. However, some of the beta subunits could still detected since the dropout design did not include the moderate expressed Glyma.20g146200 gene. These alpha' and alpha/beta dropout loci will be genetically crossed together to create complete conglycinin knockout soybean seeds.
[0142] In another editing experiment, three gRNAs (SEQ ID NOs: 28, 29, 30) were designed to do frameshift knockout of 5 highly expressed conglycinin genes (glyma.20g148200, glyma.20g148300, glyma.20g148400, glyma.10g246300 and glyma.10g246500) and one moderate-level expressed glyma.20g146200, in a multiplex frameshift knockout approach. The Homozygous T2 seeds will be analyzed for protein profile change and amino acid composition improvement.
[0143] The leghemoglobin over-expression approach and the conglycinin knockout approach are combined by either genetic crosses, or by performing gene editing in the leghemoglobin over-expression soybean lines, or by retransforming the leghemoglobin over-expression cassettes into the conglycinin knockout soybean lines. With the absence of conglycinin proteins in those soybean seeds, the contents of glycinin or other soybean proteins are expected increase to compensate the loss of conglycinin protein through protein rebalancing. It is expected that by combining the soybean leghemoglobin overexpression with the conglycinin dropout approaches, the leghemoglobin level in soybean seeds is increased.
Example 7: Characterization of Soybean Leghemoglobin Expression in Soybean Seeds
[0144] For the 8 transgenic constructs described in Example 1 and 2, T1 seeds were generated. The results were very surprising. There was almost no leghemoglobin protein accumulated in the two constructs with the GY1-SP/GY1 Basic Subunit targeting design (Experiment 4 and 8 in FIGS. 1 and 2). There is very low level of leghemoglobin accumulation (.about.0.1% of total seed protein) in the two constructs with Lectin SP targeting design (Experiment 3 and 7 in FIGS. 1 and 2) and the soybeans appeared yellow. The best expression design was from the two constructs with No Signal Peptide for the leghemoglobin (Experiment 1 and 5, shown in FIGS. 1 and 2). As shown in FIGS. 8 and 10, the "red" (i.e. pink tinged in the transverse section) colored seeds were easily identifiable by eye in these two experiments, indicating that the leghemoglobin expression level is high and the proteins were efficiently assembled as leghemoglobin protein complex containing an iron centered porphyrin (heme). Five events from the experiment 1 design (No signal peptide) and 4 events from the experiment 5 design (no signal peptide plus two heme pathway genes) were generated and all 9 events had the "red" seed phenotype. The presence of the leghemoglobin in the seeds was further verified by performing seed protein extraction, SDS PAGE gel, and Coomassie blue staining. The 16 KD leghemoglobin is easily visible by Coomassie blue staining (arrows in FIGS. 9 and 11). In FIG. 9, lanes 1, 2, 3, 5, 6, 7, 9, 10, 11 are the protein samples from the "red" pink-colored leghemoglobin positive seeds from three independent events in experiment 1 design, and lanes 4, 8, and 12 are the protein samples from the yellow null segregate seeds from the same three events. Similarly, in FIG. 11, the #14 lane is from the yellow null segregate seeds, and the #11, #21, #33, #43 are the protein samples from the "red" (pink color) leghemoglobin positive seeds from three independent events from the experiment 5 design.
[0145] Sample Preparation for Single Seed Analysis.
[0146] Single T.sub.1 red and yellow soybeans harvested from individual segregating T.sub.0 plants were placed in a Spex Certiprep 1/2.times.2'' polycarbonate vial with cap (cat #3116PC). A 3/8'' stainless steel ball bearing was added. Grinding was performed in a Spex Certiprep 2000 Geno/Grinder at 1500 strokes/min for three 30 second intervals with a 1-minute rest between each cycle.
[0147] Alternatively, soybeans were ground with a pestle, in the presence of liquid nitrogen, in a precooled mortar. The powders were then lyophilized for 48 h and kept at -20.degree. C. in a desiccator until processed.
[0148] Moisture Content Determinations were Performed According to American Oil Chemists Society (AOCS Official Method Ba 2a-38, Modified for Small Samples) as Follows:
[0149] Weigh powdered sample material (approximately 100 mg; to an accuracy of 0.1 mg) into a pre-weighed (and recorded) 13.times.100 mm glass tube VWR (53283-800) and weigh again.
[0150] Place samples into a forced air oven preheated to 130.degree. C.
[0151] Allow material to dry for 2 h.
[0152] Remove tubes into a desiccator cabinet and allow to come to room temperature before weighing again.
[0153] Cap tube and save residual dried material for subsequent combustion analysis for protein (see below).
[0154] Store in a desiccator for further analysis.
[0155] Total Protein Analysis.
[0156] Protein contents were estimated by combustion analysis of the oven dried or lyophilized powders described above. Analysis was performed on a Flash 1112EA combustion analyzer (commercially available from Thermo) running in the N-protein mode, according to the manufacturer's instructions, using aspartic acid as the standard. The powdered samples, 30-40 mg, weighed to an accuracy of 0.001 mg on a Mettler-Toledo MX5 microbalance were used for analysis. Protein contents were calculated by multiplying % N, determined by the analyzer, by 6.25. Final protein contents were assumed to be at a dry basis for the oven dried material and on an as measured basis for the lyophilized material.
[0157] Calculation of Moisture Content. The as is moisture content of the tissues was determined after oven drying using the following formula:
Moisture = ( wt . .times. tube + tissue .times. .times. as .times. .times. is - wt . .times. tube ) - ( wt . .times. tube + tissue .times. .times. dry - wt . .times. tube ) ( wt . .times. tube + tissue .times. .times. as .times. .times. is - wt . .times. tube ) .times. 100 ##EQU00001##
[0158] Quantitation of Globin Protein by LC-MS-MS.
[0159] The amino acid sequence of the globin protein (Table 1; SEQ ID 2) was assessed in-silico for potential trypsin digestion sites and the suitability of the resultant peptides for quantitative mass spectrometry. The following criteria were applied;
[0160] The peptide was between 6 and 20 amino acids in length
[0161] The amino acids within the peptide were unlikely to undergo secondary modifications.
[0162] The absence of sulfur containing amino acids
[0163] Solubility and iso-electric point.
[0164] Using these criteria, three potential peptides were identified. These were further analyzed using an online application available from Thermo Fisher Scientific at thermofisher.com/us/en/home/life-science/protein-biology/peptides-protein- s/custom-peptide-synthesis-services/peptide-analyzing-tool.html. Based on the output of this application two peptides were selected. The sequences of these peptides were subjected to a BLAST search using the NCBI Protein BLAST (protein-protein) program blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastp&PAGE_TYPE=BlastSearch&LIN- K_LOC=blasthome and were determined to be unique to the soybean globin sequence within the soybean (Glycine max) genome. The peptides were synthesized as follows:
TABLE-US-00009 Peptide 1: SEQ ID NO: 33 K.ANGTVVADAALGSIHAQK.A [position 78-95 of SEQ ID NO: 2] Peptide 2: SEQ ID NO: 34 K.AITDPQFVVVK.E [position 96-106]
[0165] Where the "." indicates the enzymatic digestion site and the parenthetical values denote the amino acid residue position relative to the N-terminal end of the mature globin protein.
[0166] Peptide stocks, at a concentration of 500 ppm, were prepared and stored as aliquots at -80.degree. C. These stocks were used to further assess the suitability of the peptides for quantitative analysis. Peptide stocks were infused into the Mass Spectrometer (SCIEX 5500 Qtrap; SCIEX LLC, Redwood City, Calif. USA) to optimize the parameters for detection. Upon analysis, the peptide AITDPQFVVVK (Peptide 2) proved to be the best candidate, with a +2 charge state of the parent ion (608.9 m/z). Following optimization of fragmentation in the collision cell, a surrogate daughter ion (816.6 m/z) with the highest abundance, was chosen to develop quantitation against. A second confirmatory ion (444.3 m/z) was also chosen.
[0167] Sample Preparation
[0168] Powder samples of .about.10-20 mg (weighed and recorded to an accuracy of 0.1 mg) were placed into 1.2 ml Micro Titer Tubes (Fisher Brand 02-681-376). Extraction buffer, 8 mM (3-[(3-Cholamidopropyl)dimethylammonio]-1-propanesulfonate hydrate, (CHAPS); 0.1% Triton X-100, pH 8.4 was added at a tissue weight to volume ratio of 25. One small steel ball was added to each vial and after capping the samples were extracted in a Geno/Grinder; 1150 oscillations per minute for 30 seconds. The contents of the homogenization tube, minus the steel ball, were quantitatively transferred to clean 1.5 ml microfuge tubes and the samples were cleared in a microcentrifuge; 10,670.times.g for 10 minutes. The supernatants were transferred to clean 1.5 ml microcentrifuge tubes and the samples were again centrifuged; 10,670.times.g, for 5 minutes. Total soluble protein concentrations of the supernatants were determined using the Bradford assay and the results were used to normalize samples to 1 mg soluble protein per ml, by dilution with trypsin digestion buffer (100 mM Ammonium Bicarbonate; 0.05% Tween-20; pH 8.3). Samples were prepared for trypsin digestion by adding 50 ul of the protein normalized extract to 100 ul of trypsin digestion buffer, 6 ul of 0.25M DTT (dithiothreitol; in digestion buffer) and incubating them at 95.degree. C. for 20 minutes. Iodoacetamide, 6 ul of 300 mM stock was added to each sample and they were incubated in the dark for one hour at room temperature. Trypsin (Pierce, MS Grade; Thermo Fisher Scientific) 10 ul of 0.1 ug/ul stock, was added to each sample and they were incubated overnight at 37.degree. C. in a static incubator. The tryptic digestions were terminated by the addition of 10 ul of 10% formic acid. Samples were then analyzed using UHPLC-MS-MS analysis.
[0169] LC/MS/MS Methods
[0170] Quantitative analysis of the tryptic digests was performed on a UHPLC (Agilent 1290) with SCIEX 5500 Qtrap detector, operating in the positive ion mode. Samples and standards (10 ul injections) were separated on a Waters Cortex C18, 2.7 um (2.1.times.100 mm) reverse phase column maintained at 40.degree. C. The solvent flow rate was 300 ul/min with starting conditions of, 90% solvent A (99.9% MS grade Water; 0.1% Formic Acid)-10 solvent B (99.9% Acetonitrile, 0.1% Formic Acid). The conditions were ramped to 60% solvent A--40% solvent B over a 7 minute period, followed by a further ramp to 10% Solvent A--90% Solvent B over 0.5 min. The solvents were then returned to the starting conditions, over a 3 minute period and the column was equilibrated under the starting conditions for a further 3 minutes before the next injection. An Electrospray Ionization (ESI) source was used to introduced samples into the MS. Source parameters were as follows: Declustering potential 135 (V), Temperature 350.degree. C., and Ion Spray voltage 350V. An MRM (Multiple Reaction Monitoring) detection technique was used to identify and quantitate the product ion (m/z: 816.6) using a collision cell energy of 35 (eV) to fragment the parent +2 molecule (m/z 608.9). Another product ion (m/z: 444.3) was used to confirm identity (based on the presence or absence). Quantitation was performed against a standard curve of the peptide that had been taken through all of the sample preparation steps described above.
[0171] Table 8 shows the Quantitative Mass Spectrometry analysis of yellow (WT) and red soybeans harvested from segregating T.sub.0 plants expressing the LegHemoglobin construct with no targeting sequences (Material from Experiment 1). Soluble protein was measured in the analyzed extracts and is expressed as a % of the total protein contents of the bean powders, as determined by combustion analysis. The LegHemoglobin protein was quantified by UHPLC-MS-MS and is expressed on a wt % of the soluble protein or total protein basis.
TABLE-US-00010 TABLE 8 Quantitative Mass Spectrometry analysis of wildtype (yellow) and red soybeans harvested from segregating T.sub.0 plants expressing the LegHemoglobin construct with no targeting sequences Soluble LegHemoglobin LegHemoglobin protein/total protein/soluble protein/total SAMPLE ID Protein (wt %) protein (wt %) protein (wt %) GMZ3A9.1.20; A 56.6 0.83 0.47 GMZ3A9.1.20; B 56.9 1.35 0.77 GMZ3A9.1.20; C 74.2 0.49 0.37 GMZ3A9.1.22; A 62.5 1.15 0.72 GMZ3A9.1.22; B 65.9 1.10 0.72 GMZ3A9.1.24; A 81.3 0.95 0.77 GMZ3A9.1.24; B 50.7 1.42 0.72 GMZ3A9.1.24; C 65.0 0.73 0.48 GMZ3A9.1.30; A NA 0.97 NA GMZ3A9.1.30; B 48.6 0.97 0.47 GMZ3A9.1.30; C 27.7 0.88 0.24 GMZ3A9.1.30 WT 59.3 0.00 0.00 GMZ3A9.1.34; A 75.8 1.06 0.80 GMZ3A9.1.34; B 35.0 1.93 0.68 GMZ3A9.1.34; C 65.7 1.24 0.81 GMZ3A9.1.20; WT 59.3 0.00 0.00 GMZ3A9.1.20; B 66.3 0.34 0.23 GMZ3A9.1.22; A 62.8 0.88 0.55 GMZ3A9.1.30; A 66.6 0.88 0.58 GMZ3A9.1.34; A 67.5 1.12 0.77 GMZ3A9.1.34; B 77.4 0.97 0.74
[0172] Expression of the soluble protein of the extracted sample as a % of the total protein content of the starting material indicated that the efficiency of extraction ranged from 27.7-81.3%, average 61.3% (Table 8). This led to substantial differences in the % globin protein contents, when expressed on a per unit soluble protein or per unit total protein basis. Yellow, wild type beans (GMZ3A9.1.30 WT and GMZ3A9.1.20; WT), had no detectable globin protein in them (Table 8). Red beans had between 0.34-1.93 wt % globin protein, when expressed on a soluble protein basis and up to 0.8 wt % when expressed on a total protein basis.
[0173] To improve the extraction efficiency and make the sample preparation more uniform the sample preparation was modified as follows; powder samples of 10+/-0.5 mg (weighed and recorded to an accuracy of 0.1 mg) were placed into 1.2 ml Micro Titer Tubes (Fisher Brand 02-681-376). Extraction buffer, 8 mM (3-[(3-Cholamidopropyl)dimethylammonio]-1-propanesulfonate hydrate, (CHAPS); 0.1% Triton X-100, pH 8.4 was added at a tissue weight to volume ratio of 50. One small steel ball was added to each vial and after capping the samples were extracted in a Geno/Grinder; 1150 oscillations per minute for 30 seconds and then on an end over end rotator for 10 minutes, the genogrinding step was then repeated. The contents of the homogenization tube, minus the steel ball, were quantitatively transferred to clean 1.5 ml microfuge tubes and the samples were cleared in a microcentrifuge; 10,670.times.g for 10 minutes. The supernatants were transferred to clean 1.5 ml microcentrifuge tubes and the samples were again centrifuged; 10,670.times.g, for 5 minutes. Total soluble protein concentrations of the supernatants were determined using the Bradford assay and the results were used to normalize samples to 1 mg soluble protein per ml, by dilution with trypsin digestion buffer (100 mM Ammonium Bicarbonate; 0.05% Tween-20; pH 8.3). Samples were prepared for trypsin digestion by adding 25 ul of the protein normalized extract to 125 ul of trypsin digestion buffer, 6 ul of 0.25M DTT (dithiothreitol; in digestion buffer) and incubating them at 95.degree. C. for 20 minutes. Iodoacetamide, 6 ul of 300 mM stock was added to each sample and they were incubated in the dark for one hour at room temperature. Trypsin (Pierce, MS Grade; Thermo Fisher Scientific) 10 ul of 0.1 ug/ul stock, was added to each sample and they were incubated overnight at 37.degree. C. in a static incubator. The tryptic digestions were terminated by the addition of 10 ul of 10% formic acid. Samples were then analyzed using UHPLC-MS-MS analysis.
[0174] The modified extraction method resulted in an average of 97% (range 95.5-100%) of the soluble protein being extracted in the first extraction. This represented an average of 71% (range 62-78%) of the total protein content of the extracted material. Using this method, yellow and red soybeans from events where only the Leg Hemoglobin was being expressed (Experiment 1) were compared to yellow and red beans from events where the Leg Hemoglobin protein was being expressed (no signal peptide) in conjunction with two heme pathway genes (Experiment 5). The results are shown in Table 9. Quantitative Mass Spectrometry analysis of yellow (WT) and red soybeans harvested from segregating T.sub.0 plants expressing only the leghemoglobin construct (no targeting sequences) Experiment 1 or the leghemoglobin construct (no targeting sequences) in conjunction with two heme pathway genes (Experiment 5). Soluble protein was measured in the analyzed extracts and is expressed as a % of the total protein contents of the bean powders, as determined by combustion analysis. The leghemoglobin protein was quantified by UHPLC-MS-MS and is expressed on a wt % of the soluble protein or total protein basis.
TABLE-US-00011 TABLE 9 Quantitative Mass Spectrometry analysis of wildtype (yellow) and red soybeans harvested from segregating T.sub.0 plants expressing the leghemoglobin construct with no targeting sequences (Experiment 1) or the leghemoglobin construct (no targeting sequences) in conjunction with two heme pathway genes (Experiment 5) Soluble Leghemoglobin Leghemoglobin protein/total protein/soluble protein/total SAMPLE ID Protein (wt %) protein (wt %) protein (wt %) EXPERIMENT 1 LegH Only GMZ3A9.1.20; WT 76.5 0 0 GMZ3A9.1.20; 29 72.8 0.31 0.21 GMZ3A9.1.22; 31 69.4 0.81 0.54 GMZ3A9.1.30; 39 78.2 0.79 0.60 GMZ3A9.1.34; 43-1 63.0 1.16 0.70 GMZ3A9.1.34; 43.2 70.9 1.13 0.80 EXPERIMENT 5 LegH plus heme pathway augmentation GM9RDV.001.5a; WT 75.1 0 0 GM9RDV.001.5a; 5.0 63.2 1.01 0.64 GM9RDV.001.5a; 5.3 75.1 1.09 0.82 GM9RDV.001.5a; 6.0 60.5 1.13 0.68 GM9RDV.001.18a; WT 59.1 0 0 GM9RDV.001.18a; 6 52.6 0.78 0.1 GM9RDV.001.18a; 7 64.7 0.64 0.42 GM9RDV.001.18a; 18.2 68.5 1.12 0.78
[0175] The data show that the quantity of the leghemoglobin protein reached similar levels in both experiments indicating that the up regulation of the heme pathway did not positively or negatively influence the level of leghemoglobin protein in these events with a maximum level, when expressed on a total protein basis reaching 0.82% for the leghemoglobin protein plus heme pathway genes (Experiment 5) and 0.80% for the leghemoglobin protein only material (Experiment 1).
[0176] Visually there were no obvious intensity differences in the red color of the beans from each experiment, which was interpreted as showing that there was no difference in the amount of leghemoglobin protein complex in the beans from the two experiments. Seed from Experiments 1 and 5 were analyzed at the next generation.
TABLE-US-00012 TABLE 9A Quantitative Mass Spectrometry analysis of wildtype (yellow) and red (pink) soybeans harvested from segregating T.sub.1 plants expressing the leghemoglobin construct with no targeting sequences (Experiment 1) or the leghemoglobin construct (no targeting sequences) in conjunction with two heme pathway genes (Experiment 5) Soluble Leghemoglobin Leghemoglobin protein/total protein/soluble protein/total SAMPLE ID Protein (wt %) protein (wt %) protein (wt %) EXPERIMENT 1 LegH Only GMZ3A9.001.24 94.34 1.45 0.66 GMZ3A9.001.20a 83.19 0.00 0.00 GMZ3A9.001.30a 66.08 1.76 0.07 GMZ3A9.001.22a 65.07 2.54 0.41 GMZ3A9.001.34a 85.76 1.14 0.93 GMZ3A9.001.24a 53.99 2.90 1.38 GMZ3A9.001.20a 88.48 1.16 0.03 GMZ3A9.001.30a 66.53 2.06 0.11 GMZ3A9.001.22a 59.80 1.96 0.41 GMZ3A9.001.20a 83.19 0.00 0.00 EXPERIMENT 5 LegH plus heme pathway augmentation 93Y21 69.60 0.00 0.00 GM9RDV.001.5a 92.14 2.26 1.84 GM9RDV.001.20a 75.51 3.02 1.07 GM9RDV.001.26a 95.73 1.76 0.34 GM9RDV.001.5a 86.66 0.00 0.00 GM9RDV.001.5a 88.09 1.77 0.05 GM9RDV.001.20a 78.21 1.47 1.02 GM9RDV.001.26a 86.02 1.68 0.58
[0177] The data in Table 9A show that homozygous seed have higher amounts of leghemoglobin than the T1 seed from the same events. For example, in T2 seed from event GMZ3A9.001.24a 1.38% of the total protein was determined to be leghemoglobin (dry basis), compared to 0.65 wt % in T1 seed, a two-fold increase. Similarly, in T2 seed from event GM9RDV.001.5a, 1.84 wt % of the total protein was determined to be leghemoglobin (dry basis) compared to a T1 seed value of 0.71 wt % (three seed average), a 2.6 fold increase. The data indicate that leghemoglobin levels double when seeds are homozygous.
Example 8: Stacking of Soybean Leghemoglobin Line with High Oleic Acid Lines
[0178] The transgenic leghemoglobin events describe above are genetically crossed with a high oleic line, such as a soybean line containing at least 50%, 70% or 75% oleic acid of the total fatty acids. Similarly, the leghemoglobin gene editing variant described above is genetically crossed with the high oleic line, such as containing at least 50%, 70% or 75% oleic acid of the total fatty acids. Alternately, leghemoglobin gene editing is performed directly in the high oleic acid line. Furthermore, both leghemoglobin gene editing and FAD2/FAD3 editing are performed together to stack the leghemoglobin trait with the high oleic trait exclusively through a gene editing approach. The resulting soybean seeds have increased nutritional value and provide an improved flavor to the soybean protein, soybean isolate or soybean concentrate.
Example 9: Extraction of Leghemoglobin Complex from Soybeans
[0179] In order for the leghemoglobin complexed protein to be used for downstream products the soybeans require processing. Typically, this involves: tempering, cracking, dehulling, solvent extraction of the oil, and toasting to remove residual solvent and to inactivate proteinaceous antinutritional factors, such as protease inhibitors. The meals or flours (typically greater than 47.5% protein) resulting from these processing steps can be further refined to concentrate the protein fraction by the removal of soluble sugars, to form soybean protein concentrates (typically greater than 65% protein). There are three processes used to create soy protein concentrates, namely alcohol washing, acid washing or hot-water extraction (Deak, N. A., Johnston, L. A., Lusas, E. W., and Rhee, K. C., 2008. Soybeans: Chemistry, Production, Processing, and utilization. Johnston, L. A., White, P. J., and Galloway, R. eds AOCS Press). All of these processes would be expected to substantially de-hemelate the leghemoglobin complexes making them unsuitable for recovery and subsequent use.
[0180] Alternatively, soybeans can be processed to the form of soy protein isolates (typically greater than 90% protein). This is achieved by solubilization of the protein fraction of lightly toasted white flake (defatted meal), prior to centrifugation to remove the bulk of the insoluble carbohydrate (fiber) fraction. The protein is then precipitated, by pH adjustment, and washed to remove remaining soluble carbohydrates. The purified protein fractions are pasteurized to inactivate residual protease inhibitor activity and microbial contaminants, prior to drying to a powder. These final steps in the soy isolate production process would also be expected to substantially de-hemelate the leghemoglobin complexes making them unsuitable for recovery and subsequent use.
[0181] To investigate the solubility/extractability of the leghemoglobin protein from soybeans the following experiment was performed. Soybean powder samples from red leghemoglobin expressing soybeans (10+/-0.5 mg; weighed and recorded to an accuracy of 0.1 mg) were placed into 1.2 ml Micro Titer Tubes (Fisher Brand 02-681-376). Extraction buffer, 8 mM (3-[(3-Cholamidopropyl)dimethylammonio]-1-propanesulfonate hydrate, (CHAPS); 0.1% Triton X-100, pH 8.4 was added at a tissue weight to volume ratio of 50. One small steel ball was added to each vial and after capping the samples were extracted in a Geno/Grinder; 1150 oscillations per minute for 30 seconds and then on an end over end rotator for 10 minutes, the genogrinding step was then repeated. The contents of the homogenization tube, minus the steel ball, were quantitatively transferred to clean 1.5 ml microfuge tubes and the samples were cleared in a microcentrifuge; 10,670.times.g for 10 minutes. The supernatant was transferred to clean 1.5 ml microcentrifuge tubes. The residual pellet from this step was extracted, as described, two more times. Aliquots from the first, second and third supernatants were analyzed for soluble protein and leghemoglobin, as described in Example 7. The results are shown in Table 10.
TABLE-US-00013 TABLE 10 Efficacy of alkaline buffer extraction on whole soybean powders (average from 6 samples) on the recovery of soluble protein and leghemoglobin protein. Methods for quantitation are provided in Example 5. % Soluble Protein % Leghemoglobin Extraction # Recovery Recovery 1 97.2 100 2 2.8 0 3 0 0
[0182] The data show that all of the leghomoglobin protein was recovered in the first extraction.
Example 10: Enzymatic Soybean Processing (E-SOY) for Leghemoglobin Soybeans
[0183] A challenge for isolating proteins from soybean meals is the requirement for high water:solid ratios to yield a slurry that can be physically processed. This leads to large volumes of water in the process, which increase the processing costs associated with concentrating the extracted proteins, drying non-solubilized meal residues, and ultimately waste water disposal. A novel approach to isolating protein from defatted soybean meal was developed that uses food grade polysaccharide degrading enzymes to convert insoluble or viscous polysaccharide materials in the meal to solubilized, short chain polysaccharides, greatly reducing the volume of water required for protein extraction and isolation. This process used to isolate more traditional soybean protein concentrates and isolates, is expected to facilitate the isolation of leghemoglobin while maintaining the intact leghemoglobin complex. The generalized enzymatic soybean (E-SOY) process is outlined below.
[0184] Process Flow Chart
[0185] FIG. 12 shows an example of a generalized process flow chart for the E-SOY process. Laboratory defatted whole soybean meal (typically 40 g aliquots for the lab process) in a beaker was mixed with preheated water at a ratio 3:1 liquid:solid. After an incubation period, the beaker was placed on a regulated water bath, typically maintained at 50.degree. C. An overhead stirrer (Lightnin Mixer) equipped with a low shear impeller was used to agitate the meal dough. The viscosity of the mixture was estimated by the stirrer power (watts) required to stir the dough at 200 rpm. With stirring an aliquot of 5N HCl was added dropwise to the dough (typically 4-5 mLs) to lower the pH of the slurry to a range of pH 3.8-4.5. The exact amount of acid required to lower the pH of different feedstocks was determined separately using feedstock suspended in a larger volume of water. Aliquots of the commercial enzymes (typically 200 .mu.L of the cellulase, hemicellulase, and pectinase, 10 .mu.L for liquid pectinase). The dough was then stirred and allowed to react with the enzymes for variable lengths of time, although 3 hours was routinely used. The power reading on the overhead mixer was recorded at intervals to monitor the viscosity of the dough slurry.
[0186] When the viscosity of the solution has been reduced sufficiently to form a free flowing slurry, the mixture was quantitatively transferred to a Buchner funnel fitted with a 500 uM mesh polypropylene screen and vacuum filtered. The filter cake was washed with 60 mLs of water. The filtrate solution was passed over a second Buchner funnel fitted with a 105 um mesh polypropylene screen and rinsed with deionized water from a wash bottle. The filter cakes from the 500 um and 105 um filtrations were quantitatively transferred to a tared container and dried in a vacuum oven at 65.degree. C. for at least 48 hours. The dry weight of the combined filter cakes was determined, and the residue (Meal Residue) was ground in a coffee mill prior to further analysis.
[0187] The filtrate slurry from the 105 um filtration was adjusted to pH 6.5-7 with 2N NaOH, although for leghemoglobin isolation a pH of between 7 and 11 would be used. After stirring, the solution was transferred to centrifuge bottles and centrifuged at 7000 rpm for 10 minutes in a fixed angle rotor. The supernatant solution was carefully poured into a disposable 0.45 um-1 L filter apparatus for filtration under vacuum. The centrifuge solids (concentrated precipitated protein concentrate, or CPPC) was suspended in deionized water, quantitatively transferred to a tared flask, and shell frozen and freeze dried. The recovered dry weight of CPPC was determined after freeze drying.
[0188] The 0.45 um filtered supernatant of soluble protein solution (SPS), was transferred to an ultrafiltration device equipped with a PES membrane module (Vivaflow 200, variable MWCO depending on the experiment). The SPS was concentrated down to a volume of .about.50 mLs, then diafiltered with deionized water (100-300 mLs, depending on the experiment). The UF/DF permeate was collected in a flask for further processing. When the UF/DF step was completed, the retentate solution was quantitatively transferred to centrifuge bottles and centrifuged at 7000 rpm for 10 minutes to remove any precipitate that formed during the UF/DF process. The supernatant was decanted into a tared flask, shell frozen and lyophilized. The recovered solids (concentrated soluble protein isolate, or CSPI) was weighed and stored for further study. If any solids were recovered from the centrifugation step (soluble protein solution precipitate, or SPS-ppt), it was quantitatively transferred to another tared flask for freezing, freeze drying, and analysis.
[0189] In some variations of the UF/DF process, sequential UF/DF using a series of decreasing MWCO porosities was examined. The process was essentially the same, other than the permeate from the highest MWCO step was passed through the next lower MWCO membrane in series, and retentates were transferred to individual tared flasks to be frozen and lyophilized.
[0190] The final ultrafiltration permeate was concentrated on a rotary evaporator to dryness in a tared round bottom flask. The flask was transferred to a vacuum oven and further dried at 65.degree. C. for at least 48 hours. The solids dry weight was determined, and the material scraped from the flask wall and transferred to a container for storage.
[0191] Protein was determined as total nitrogen using a combustion analyzer (CE Elantech, Flash EA 1112 series). Total nitrogen contents were converted to total protein by multiplying % N.times.6.25.
[0192] The recovery of protein from commercial soybean meal is shown in Table 11.
TABLE-US-00014 TABLE 11 Recovery of Products from Commercial Soybean Meal. Soybean Meal Product Protein Product 50.9% Initial Yield Yield Purity Protein Protein Content (g/100 g) (g/100 g) (% Protein) (% Initial) Meal Residue (MR) 50.2 33.3 66.3 65 Protein 5.8 3.6 62.2 7 Concentrate 10 kD retentate ppt 0.4 0.3 79.8 1 10 kD retentate 4.0 4.1 102.5 8 solubles UF Permeate 25.9 2.5 9.5 5 Concentrate + Isolate 7.7 Initial Protein 15 Concentrate/Isolate 0.88
[0193] Due to the different solubilities of soy proteins, and their low solubility below pH 6, the meal residue (MR) fraction of the extracted soy meal retained 65% of the starting protein, with a protein purity of 66.3%. This actually represents an enrichment of the soy meal from the original 50.9% protein content, meeting the criteria for a protein concentrate of >65%. A much lower percentage of the initial protein was recovered in the concentrate and isolate fractions, recovering only 15% on the initial protein in these products. The concentrate fraction was actually lower in protein content than the meal residue (62.2% vs. 66.3%). The purity of the isolate fraction was excellent at 102.5%, and relatively low levels of protein was found in the UF permeate. The recovery of total protein was compromised by physical difficulties in separating the meal residue from the filter screen. Given the high protein content of the solids fraction, it might be more efficient to utilize a different physical separation of the solids from the dissolved protein of the digested meal slurry.
[0194] An unexpected finding was that the isolate protein could be recovered using a much higher MWCO ultrafiltration membrane than would have been predicted based on the molecular weight of the protein fraction. Due to an unexpected aggregated behavior in solution, much higher MWCO membranes could be used effectively. It is expected that a similar recovery is achieved for isolates produced from leghemoglobin-containing soybean. The use of higher flux membranes in the process will reduce the total surface area required, thus saving capital costs.
[0195] Steps are taken to preferentially extract the highly soluble leghemoglobin protein complex into the supernatant during the initial phases of the process and to allow its differential purification during the ultrafiltration step. This will produce a high value coproduct that could be used for downstream product formulation
[0196] The process has been tested and is generally applicable to the oilseed feedstock listed in Table 12.
TABLE-US-00015 TABLE 12 Oilseed Meals Tested with the E-SOY Process Yellow Seed Coat Canola High Oleic Canola (Nexera 845) Conventional Canola Field Mustard (Brassica rapa) Indian Mustard (Brassica juncea) Yellow Mustard (Brassica hirtu) Commercial Sunflower (defatted meal, presscake, meats) Safflower Flaxseed Sesame
[0197] Additional protein yield improvements are realized by optimizing the defatting and desolventizing processes to minimize the irreversible denaturation of protein and leghemoglobin complex in the meal matrix. Additionally, the E-Soy processing scheme can be optimized for soybeans that have been created by proteome rebalancing (such as shown Example 6) to further enhance the potential recovery of protein from the meal. Further modifications of the E-Soy process to optimize protein extractions of meals derived from high oleic oil soybeans engineered to express the leghemoglobin protein (such as shown in Example 8) are also expected to lead to improved sensory and processing properties as well. For example, see U.S. Pat. No. 9,918,485, incorporated herein by reference, for examples of suitable soybean isolates to be used as feedstock and soy processing advantages associated with high oleic soybeans.
Example 11: Soybean Seeds Expressing Leghemoglobin in Combination with Reduced Antinutritional Factors and/or Increased Protein Content
[0198] Many post-harvest processing steps which concentrate soybean protein for its subsequent incorporation into food products remove or inactivate antinutritional factors, such as protein based factors (e.g. the protease inhibitors) and carbohydrate based factors (e.g. raffino-oligosaccharides). Such steps may de-hemelate the leghemoglobin complexed protein, making it unsuitable for downstream use.
[0199] Genome editing techniques are used to reduce or knock out the expression of one or more of the Kunitz Trypsin and Bowman-Burke protease inhibitors and/or to inhibit the synthesis of one or more of raffinose and stachyose. These soybean seeds are also edited to express increased leghemoglobin or complexed leghemoglobin as described in the earlier examples. Alternatively, edited soybeans expressing increased leghemoglobin or complexed leghemoglobin are crossed with soybeans which have been edited to reduce or knock out expression of the Kunitz Trypsin protease inhibitors and Bowman-Burke protease inhibitors and/or to inhibit the synthesis of raffinose and stachyose, to create breeding stacks of the edited genes. Genome editing was used to reduce insoluble carbohydrate, such as raffinose and stachyose, by knockout of the raffinose synthase genes, such as RS2, RS3, RS4. Assays to determine the effectiveness of the engineering approaches will measure residual trypsin inhibitor activity using American Oil Chemists Society method Ba 12-75. Changes in the soluble carbohydrate profiles will be determined using the methods such as outlined in US Patent Publication No. 20190383733, which is incorporated herein by reference.
[0200] Genome editing was also used to increase total protein content in seeds by knockout or modification of key regulatory genes, such as CCT-domain containing proteins, reticulon, trehalose phosphate synthase, HECT Ubiquitin Ligase (HEL or UPL3) and/or MFT (mother of flowering time) mutated or modified plants and seeds such as disclosed in PCT/US2019/058747 incorporated herein by reference. Expression of leghemoglobin in soybeans engineered to optimize their processability as described in this example would add to the value and utility of such soybeans.
[0201] Engineering soybeans to minimize post-harvest processing steps is expected to result in greater yields of intact leghemoglobin complex. Such soybean seeds are processed without the need to use solvents to remove oil or a toasting step often employed in soybean processing. Such soybean seeds are processed using one or more of cold pressing, extrusion or Super Critical Fluid Extraction ((Friedrich J. P., List G. R., and Heakin A. J., 1982. Journal of the American Oil Chemists Society. 59 (7); 288-292).
Example 12. Transformation of the Nuclear Genomes for Seed Specific Expression of PPR10 Variants
[0202] Mature dry seed from soybean lines are surface-sterilized for 16 hours using chlorine gas, produced by mixing 3.5 mL of 12 N HCl with 100 mL of commercial bleach (5.25% sodium hypochloride), as described by Di et al. ((1996) Plant Cell Rep 15:746-750). Disinfected seeds are soaked in sterile distilled water at room temperature for 16 hours (100 seeds in a 25.times.100 mm petri dish) and imbibed on semi-solid medium containing 5 g/l sucrose and 6 g/l agar at room temperature in the dark. After overnight incubation, the seeds are soaked in distilled water for an additional 3-4 hours at room temperature in the dark. Intact embryonic axes (EA) are isolated from imbibed seeds. Agrobacterium-mediated EA transformation is carried out as described below.
[0203] A volume of 15 mL of A. tumefaciens LBA4404 or AGL-1 suspension harboring soybean nuclear transformation binary vector (FIG. 1) (OD 0.5 at 600 nm) in infection medium composed of 1/10.times. Gamborg B5 basal medium, 30 g/L sucrose, 20 mM MES, 0.25 mg/L GA3, 1.67 mg/L BAP, 200 .mu.M acetosyringone and 1 mM dithiothreitol in pH 5.4 is added to about 200-300 EAs, and they are on a 25.times.100 mm deep petri dish. The plates are sealed with parafilm (Cat No. 52858, "Parafilm M" VWR), then sonicated (Sonicator-VWR model 50T) for 30 seconds. After sonication, EAs are incubated 2 hrs at room temperature. After inoculation, excess bacterial suspension is removed and about 200-300 EAs are transferred to a single layer of autoclaved sterile filter paper (Cat No. 28320-020, VWR) in 25.times.100 mm petri dish. The plates are sealed with Micropore tape (Cat No. 1530-0, 3M, St. Paul, Minn., USA) and incubated under dim light (1-2 .mu.E/m.sup.2/s), cool white fluorescent lamps for 16 hours at 21.degree. C. for 3 days. After co-cultivation, the base of each embryonic axis is embedded in shoot induction medium (R7100, PhytoTech Labs) containing 30 g/L sucrose, 6 g/L agar and 25 mg/L spectinomycin (S742, PhytoTech Labs) as a selectable agent and 300 mg/L cefotaxime (GoldBio, ST Louis, Mo., USA) in pH5.7. Shoot induction is carried out in a Percival Biological Incubator (Percival Scientific, Perry, Iowa, USA) or growth room at 26.degree. C. with a photoperiod of 16 hours and a light intensity of 60-100 .mu.E/m.sup.2/s. After 4-6 weeks in selection medium, the spectinomycin-resistant shoots are cut and transferred to 1/2 strength MS rooting medium (M404, PhytoTech Labs) containing 15 g/L sucrose, agar 6 g/L, 10 mg/L spectinomycin and 250 mg/L cefotaxime for further shoot and root elongations. Transformation efficiency are calculated based on the number of positive transgenic soybean T0 plants divided by the total number of EAs.
[0204] For heat shock treatment of soybean to produce spectinomycin marker gene free T0 events, 2-4 cm T0 plantlets with roots in 100.times.25 mm Petri dishes or magenta boxes on spectinomycin free-rooting medium are transferred into a Percival incubator (Percival Scientific, Perry, Iowa, USA) at 45.degree. C., 70% humidity for 2 hrs in the dark. Non-heat shock treated T0 plantlets are used as a control. After the heat shock treatment, T0 plantlets are transferred to moistened Berger BM2 soil (Berger, Saint-Modeste, QC, Canada), and kept enclosed in clear plastic tray boxes in a Percival incubator at 26.degree. C. with a 16 hr photoperiod at 250-350 .mu.E/m.sup.2/s. 2-4 leaf punch samples are collected for qPCR and SbS analyses from newer growth 2 weeks after acclimatization of T0 events.
[0205] FIG. 13 is a schematic diagram showing a soybean nuclear transformation binary vector within the T-DNA. In FIG. 13, RB and LB are right border and left border of the T-DNA respectively, GM-GY1 Pro is the soybean glycinin seed specific promoter, AtUBQ10Pro is the Arabidopsis ubiquitin 10 promoter, PPR10GG is a maize or soybean RNA-binding protein PPR10 variant, UBQ10TERM is an Arabidopsis ubiquitin 10 terminator, LoxP is the lox recombination-site, Gm-HSP17.3BPro is the soybean heat shock hs6871 promoter, MoCre is the Cre recombinase, SB-GKAFTerm is the Sorghum bicolor gamma kafirin storage protein terminator, At-UBQ10 Pro is the Arabidopsis ubiquitin 10 promoter, SpcN is the soybean codon optimized spectinomycin resistant gene from Streptomyces spectabilis Genebank protein ID AAD50455, and UBQ14Term is the Arabidopsis ubiquitin 14 terminator.
Example 13. Seed Specific Leghemoglobin Expression in Transplastomic Soybean Plants: Biolistic-Mediated Soybean Chloroplast Transformation
[0206] Marker-free T1 homozygous line harboring GM-GY1 Pro:PPR10GG:SB-GAKF TERM expression cassettes are used as donor material for chloroplast transformation. Immature pods are collected from soybean marker-free T1-2 homozygous line and opened to retrieve immature seeds of about 2-8 mm in length. Immature seeds are collected and surface sterilized in a 50 mL screw cap tube containing 50 mL of a 10% bleach, 0.02% Tween-20 solution, with slight agitation for 15 minutes and are then rinsed 10 times with a total of 500 mL of sterile distilled water. Surface sterilized seeds are cut open under a microscope or under magnification. Typically, each immature seed's embryonic axis is cut off, and the two cotyledon pieces are released. Immature cotyledons are collected and transferred to flasks containing liquid S30 medium (Table 13).
TABLE-US-00016 TABLE 13 Composition of media M2 S30 M7 M8 MS salt with B5 4.44 g/L vitamins (PhytoTech M404) Gamborg B-5 basal 3.21 g/L medium (PhytoTech G398) Modified MS salt 2.68 g/L 2.68 g/L (PhytoTech M571) B5 vitamins (1000X) 1 ml 1 ml (PhytoTech G249) 2,4-D stock 10 mg/ml 4 ml 1 ml KNO.sub.3 0.93 g/L 0.93 g/L (NH.sub.4).sub.2SO.sub.4 0.463 g/L 0.463 g/L Asparagine 1 g/L 1 g/L Glutamine 4.48 g/L L-Methionine 0.149 g/L Sucrose 10 g/L 10 g/L 10 g/L Maltose 60 g/L MgCl.sub.2.cndot.6H.sub.2O 0.75 g/L Activated charcoal 5 g/L (PhytoTech C325) pH 5.8 5.8 5.7 5.7 TC agar 5 g/L 5 g/L Gelrite (Plant Media 2 g/L Cat# 714246)
[0207] Immature cotyledons are pre-cultured in S30 medium for 10 days and targeted for direct biolistic-mediated DNA transformation. After 10 days of pre-culture, twenty (20) immature cotyledons are placed on the surface of M2 solid medium supplemented with 40 mg/L of 2,4-D in the center of small petri plates (60.times.15 mm) for bombardment. The immature cotyledons are bombarded with the 0.6 .mu.m gold particle/plasmid DNA (FIG. 2) mixture at a concentration of 30 picogram/basepair/shot at 650 psi, 28 mm Hg with biolistic gene gun (PDS 1000/He, Bio-Rad). After 2 days of co-cultivation in the M2 solid medium, the bombarded immature cotyledons are transferred to liquid S30 medium containing 300 mg/L spectinomycin. Fresh S30 medium containing 300 mg/L spectinomycin is replaced bi-weekly. After 8-12 weeks of selection, spectinomycin resistant yellow green to green calli is emerged from the surface of explants. Putative transformed green callus is isolated under a microscope and plated onto petri plates with sterile filter paper overlaying M7 agar medium. The petri plates are sealed with Micropore.TM. surgical tape (3M Health Care, St. Paul, Minn., USA) and incubated at 26.degree. C. with an 18-hour photoperiod at 35-60 .mu.E/m2/s light intensity. After 3-4 weeks of maturation on M7 medium, mature somatic embryos are placed in sterile, Petri dishes and either sealed with Micropore.TM. surgical tape or placed unsealed in a plastic box for 4-7 days at room temperature for somatic embryo desiccation. After 4-7 days, desiccated embryos are plated onto M8 medium supplemented with 10 .mu.g/L spectinomycin and were allowed to germinate at 26.degree. C. with an 18-hour photoperiod at 35-60 .mu.E/m2/s light intensity. After 4-6 weeks on M8 germination medium, plantlets are transferred to 3 inchs pots containing moistened Berger BM2 soil (Berger Peat Moss, Saint-Modeste, Canada) and kept enclosed in clear plastic tray boxes until acclimatized in a culture room with a 16-hour photoperiod at 90-150 .mu.E/m2/sand 26.degree. C. day/24.degree. C. night temperatures. After acclimation, hardened plantlets are potted in 2 gallon pots containing moistened Berger MB1 (Berger Peat Moss, Saint-Modeste, Canada) and grown in a greenhouse to seed-bearing maturity.
[0208] FIG. 14 is a schematic diagram of soybean chloroplast transformation vector. GM-TRNV is the soybean plastid TRNV homology region, NT-PSBA Pro is the tobacco PSBA plastid promoter, AADA is the spectinomycin adenylyltransferase gene, NT-PSBA 3UTR is the tobacco PSBA 3UTR, DicisGG is the GG binding site into the intergenic region of a dicistronic operon (SEQ ID NO: 44), Leghemoglobin is the soybean leghemoglobin coding sequence (Glyma.20g191200), GM-RPS is the soybean plastid RPS homology region.
[0209] It is expected that the soybean seeds express leghemoglobin in the plastids of the seeds, with minimal or no expression of leghemoglobin in non-seed parts of the plant, such as roots, stems, leaves and flowers.
Example 14. Seed Specific Leghemoglobin Expression in Transplastomic Soybean Plants: Plastid Transformation Followed by Nuclear Transformation
[0210] The methodology of Example 13 is followed, except the starting donor material is a null or non-transformed soybean. The resulting soybean seeds and plants containing the transformed plastids are then transformed according to the method described in Example 12. It is expected that the soybean seeds express leghemoglobin in the plastids of the seeds, with minimal or no expression of leghemoglobin in non-seed parts of the plant, such as roots, stems, leaves and flowers.
[0211] All publications and patent applications in this specification are indicative of the level of ordinary skill in the art to which this invention pertains. All publications and patent applications are herein incorporated by reference to the same extent as if each individual publication or patent application was specifically and individually indicated by reference.
[0212] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Unless mentioned otherwise, the techniques employed or contemplated herein are standard methodologies well known to one of ordinary skill in the art. The materials, methods and examples are illustrative only and not limiting.
[0213] Many modifications and other embodiments of the inventions set forth herein will come to mind to one skilled in the art to which these inventions pertain having the benefit of the teachings presented in the foregoing descriptions and the associated drawings. Therefore, it is to be understood that the inventions are not to be limited to the specific embodiments disclosed and that modifications and other embodiments are intended to be included within the scope of the appended claims. Although specific terms are employed herein, they are used in a generic and descriptive sense only and not for purposes of limitation.
[0214] Units, prefixes and symbols may be denoted in their SI accepted form. Unless otherwise indicated, nucleic acids are written left to right in 5' to 3' orientation; amino acid sequences are written left to right in amino to carboxy orientation, respectively. Numeric ranges are inclusive of the numbers defining the range. Amino acids may be referred to herein by either their commonly known three letter symbols or by the one-letter symbols recommended by the IUPAC-IUB Biochemical Nomenclature Commission. Nucleotides, likewise, may be referred to by their commonly accepted single-letter codes.
Sequence CWU 1
1
481438DNAGlycine Max 1atgggtgctt tcactgagaa gcaagaggct ttggtgagta
gctcattcga agcattcaag 60gcaaacattc ctcaatacag cgttgtgttc tacacttcga
tactggagaa agcacccgca 120gcaaaggact tgttctcgtt tctatctaat
ggagtagatc ctagtaatcc taagctcacg 180ggccatgctg aaaagctttt
tggattggtg cgtgactcag ctggtcaact taaagcaaat 240ggaacagtag
tggctgatgc cgcacttggt tctatccatg cccaaaaagc aatcactgat
300cctcagttcg tggtggttaa agaagcactg ctgaaaacaa taaaggaggc
agttggggac 360aaatggagtg atgaattgag cagtgcttgg gaagtagcct
atgatgaatt ggcagcagct 420attaagaagg cattttag 4382145PRTGlycine Max
2Met Gly Ala Phe Thr Glu Lys Gln Glu Ala Leu Val Ser Ser Ser Phe1 5
10 15Glu Ala Phe Lys Ala Asn Ile Pro Gln Tyr Ser Val Val Phe Tyr
Thr 20 25 30Ser Ile Leu Glu Lys Ala Pro Ala Ala Lys Asp Leu Phe Ser
Phe Leu 35 40 45Ser Asn Gly Val Asp Pro Ser Asn Pro Lys Leu Thr Gly
His Ala Glu 50 55 60Lys Leu Phe Gly Leu Val Arg Asp Ser Ala Gly Gln
Leu Lys Ala Asn65 70 75 80Gly Thr Val Val Ala Asp Ala Ala Leu Gly
Ser Ile His Ala Gln Lys 85 90 95Ala Ile Thr Asp Pro Gln Phe Val Val
Val Lys Glu Ala Leu Leu Lys 100 105 110Thr Ile Lys Glu Ala Val Gly
Asp Lys Trp Ser Asp Glu Leu Ser Ser 115 120 125Ala Trp Glu Val Ala
Tyr Asp Glu Leu Ala Ala Ala Ile Lys Lys Ala 130 135
140Phe1453633DNAGlycine Max 3atgatgagag cgcggttccc attactgttg
ctgggagttg ttttcctagc atcagtttct 60gtctcatttg gcattgcgta ttgggaaaag
cagaacccca gtcacaacaa gtgcctccga 120agttgcaata gcgagaaaga
ctcctacagg aaccaagcat gccacgctcg ttgcaacctc 180cttaaggtgg
aggaaatggg tgctttcact gagaagcaag aggctttggt gagtagctca
240ttcgaagcat tcaaggcaaa cattcctcaa tacagcgttg tgttctacac
ttcgatactg 300gagaaagcac ccgcagcaaa ggacttgttc tcgtttctat
ctaatggagt agatcctagt 360aatcctaagc tcacgggcca tgctgaaaag
ctttttggat tggtgcgtga ctcagctggt 420caacttaaag caaatggaac
agtagtggct gatgccgcac ttggttctat ccatgcccaa 480aaagcaatca
ctgatcctca gttcgtggtg gttaaagaag cactgctgaa aacaataaag
540gaggcagttg gggacaaatg gagtgatgaa ttgagcagtg cttgggaagt
agcctatgat 600gaattggcag cagctattaa gaaggcattt tag
6334210PRTGlycine Max 4Met Met Arg Ala Arg Phe Pro Leu Leu Leu Leu
Gly Val Val Phe Leu1 5 10 15Ala Ser Val Ser Val Ser Phe Gly Ile Ala
Tyr Trp Glu Lys Gln Asn 20 25 30Pro Ser His Asn Lys Cys Leu Arg Ser
Cys Asn Ser Glu Lys Asp Ser 35 40 45Tyr Arg Asn Gln Ala Cys His Ala
Arg Cys Asn Leu Leu Lys Val Glu 50 55 60Glu Met Gly Ala Phe Thr Glu
Lys Gln Glu Ala Leu Val Ser Ser Ser65 70 75 80Phe Glu Ala Phe Lys
Ala Asn Ile Pro Gln Tyr Ser Val Val Phe Tyr 85 90 95Thr Ser Ile Leu
Glu Lys Ala Pro Ala Ala Lys Asp Leu Phe Ser Phe 100 105 110Leu Ser
Asn Gly Val Asp Pro Ser Asn Pro Lys Leu Thr Gly His Ala 115 120
125Glu Lys Leu Phe Gly Leu Val Arg Asp Ser Ala Gly Gln Leu Lys Ala
130 135 140Asn Gly Thr Val Val Ala Asp Ala Ala Leu Gly Ser Ile His
Ala Gln145 150 155 160Lys Ala Ile Thr Asp Pro Gln Phe Val Val Val
Lys Glu Ala Leu Leu 165 170 175Lys Thr Ile Lys Glu Ala Val Gly Asp
Lys Trp Ser Asp Glu Leu Ser 180 185 190Ser Ala Trp Glu Val Ala Tyr
Asp Glu Leu Ala Ala Ala Ile Lys Lys 195 200 205Ala Phe
2105543DNAGlycine Max 5atggctactt caaagttgaa aacccagaat gtggttgtat
ctctctccct aaccttaacc 60ttggtactgg tgctactgac cagcaaggca aactcagcgg
aaactatggg tgctttcact 120gagaagcaag aggctttggt gagtagctca
ttcgaagcat tcaaggcaaa cattcctcaa 180tacagcgttg tgttctacac
ttcgatactg gagaaagcac ccgcagcaaa ggacttgttc 240tcgtttctat
ctaatggagt agatcctagt aatcctaagc tcacgggcca tgctgaaaag
300ctttttggat tggtgcgtga ctcagctggt caacttaaag caaatggaac
agtagtggct 360gatgccgcac ttggttctat ccatgcccaa aaagcaatca
ctgatcctca gttcgtggtg 420gttaaagaag cactgctgaa aacaataaag
gaggcagttg gggacaaatg gagtgatgaa 480ttgagcagtg cttgggaagt
agcctatgat gaattggcag cagctattaa gaaggcattt 540tag
5436180PRTGlycine Max 6Met Ala Thr Ser Lys Leu Lys Thr Gln Asn Val
Val Val Ser Leu Ser1 5 10 15Leu Thr Leu Thr Leu Val Leu Val Leu Leu
Thr Ser Lys Ala Asn Ser 20 25 30Ala Glu Thr Met Gly Ala Phe Thr Glu
Lys Gln Glu Ala Leu Val Ser 35 40 45Ser Ser Phe Glu Ala Phe Lys Ala
Asn Ile Pro Gln Tyr Ser Val Val 50 55 60Phe Tyr Thr Ser Ile Leu Glu
Lys Ala Pro Ala Ala Lys Asp Leu Phe65 70 75 80Ser Phe Leu Ser Asn
Gly Val Asp Pro Ser Asn Pro Lys Leu Thr Gly 85 90 95His Ala Glu Lys
Leu Phe Gly Leu Val Arg Asp Ser Ala Gly Gln Leu 100 105 110Lys Ala
Asn Gly Thr Val Val Ala Asp Ala Ala Leu Gly Ser Ile His 115 120
125Ala Gln Lys Ala Ile Thr Asp Pro Gln Phe Val Val Val Lys Glu Ala
130 135 140Leu Leu Lys Thr Ile Lys Glu Ala Val Gly Asp Lys Trp Ser
Asp Glu145 150 155 160Leu Ser Ser Ala Trp Glu Val Ala Tyr Asp Glu
Leu Ala Ala Ala Ile 165 170 175Lys Lys Ala Phe 18071071DNAGlycine
Max 7atggccaagc tagttttttc cctttgtttt ctgcttttca gtggctgctg
cttcgctttc 60agttccatgg gtgctttcac tgagaagcaa gaggctttgg tgagtagctc
attcgaagca 120ttcaaggcaa acattcctca atacagcgtt gtgttctaca
cttcgatact ggagaaagca 180cccgcagcaa aggacttgtt ctcgtttcta
tctaatggag tagatcctag taatcctaag 240ctcacgggcc atgctgaaaa
gctttttgga ttggtgcgtg actcagctgg tcaacttaaa 300gcaaatggaa
cagtagtggc tgatgccgca cttggttcta tccatgccca aaaagcaatc
360actgatcctc agttcgtggt ggttaaagaa gcactgctga aaacaataaa
ggaggcagtt 420ggggacaaat ggagtgatga attgagcagt gcttgggaag
tagcctatga tgaattggca 480gcagctatta agaaggcatt tagcagaaga
aatggcattg acgagaccat atgcaccatg 540agacttcgcc acaacattgg
ccagacttca tcacctgaca tctacaaccc tcaagccggt 600agcgtcacaa
ccgccaccag ccttgacttc ccagccctct cgtggctcag actcagtgct
660gagtttggat ctctccgcaa gaatgcaatg ttcgtgccac actacaacct
gaacgcgaac 720agcataatat acgcattgaa tggacgggca ttgatacaag
tggtgaattg caacggtgag 780agagtgtttg atggagagct gcaagaggga
cgggtgctga tcgtgccaca aaactttgtg 840gtggctgcaa gatcacagag
tgacaacttc gagtatgtgt cattcaagac caatgataca 900cccatgatcg
gcactcttgc aggggcaaac tcattgttga acgcattacc agaggaagtg
960attcagcaca ctttcaacct aaaaagccag caggccaggc agataaagaa
caacaaccct 1020ttcaagttcc tggttccacc tcaggagtct cagaagagag
ctgtggctta g 10718356PRTGlycine Max 8Met Ala Lys Leu Val Phe Ser
Leu Cys Phe Leu Leu Phe Ser Gly Cys1 5 10 15Cys Phe Ala Phe Ser Ser
Met Gly Ala Phe Thr Glu Lys Gln Glu Ala 20 25 30Leu Val Ser Ser Ser
Phe Glu Ala Phe Lys Ala Asn Ile Pro Gln Tyr 35 40 45Ser Val Val Phe
Tyr Thr Ser Ile Leu Glu Lys Ala Pro Ala Ala Lys 50 55 60Asp Leu Phe
Ser Phe Leu Ser Asn Gly Val Asp Pro Ser Asn Pro Lys65 70 75 80Leu
Thr Gly His Ala Glu Lys Leu Phe Gly Leu Val Arg Asp Ser Ala 85 90
95Gly Gln Leu Lys Ala Asn Gly Thr Val Val Ala Asp Ala Ala Leu Gly
100 105 110Ser Ile His Ala Gln Lys Ala Ile Thr Asp Pro Gln Phe Val
Val Val 115 120 125Lys Glu Ala Leu Leu Lys Thr Ile Lys Glu Ala Val
Gly Asp Lys Trp 130 135 140Ser Asp Glu Leu Ser Ser Ala Trp Glu Val
Ala Tyr Asp Glu Leu Ala145 150 155 160Ala Ala Ile Lys Lys Ala Phe
Ser Arg Arg Asn Gly Ile Asp Glu Thr 165 170 175Ile Cys Thr Met Arg
Leu Arg His Asn Ile Gly Gln Thr Ser Ser Pro 180 185 190Asp Ile Tyr
Asn Pro Gln Ala Gly Ser Val Thr Thr Ala Thr Ser Leu 195 200 205Asp
Phe Pro Ala Leu Ser Trp Leu Arg Leu Ser Ala Glu Phe Gly Ser 210 215
220Leu Arg Lys Asn Ala Met Phe Val Pro His Tyr Asn Leu Asn Ala
Asn225 230 235 240Ser Ile Ile Tyr Ala Leu Asn Gly Arg Ala Leu Ile
Gln Val Val Asn 245 250 255Cys Asn Gly Glu Arg Val Phe Asp Gly Glu
Leu Gln Glu Gly Arg Val 260 265 270Leu Ile Val Pro Gln Asn Phe Val
Val Ala Ala Arg Ser Gln Ser Asp 275 280 285Asn Phe Glu Tyr Val Ser
Phe Lys Thr Asn Asp Thr Pro Met Ile Gly 290 295 300Thr Leu Ala Gly
Ala Asn Ser Leu Leu Asn Ala Leu Pro Glu Glu Val305 310 315 320Ile
Gln His Thr Phe Asn Leu Lys Ser Gln Gln Ala Arg Gln Ile Lys 325 330
335Asn Asn Asn Pro Phe Lys Phe Leu Val Pro Pro Gln Glu Ser Gln Lys
340 345 350Arg Ala Val Ala 3559613DNAGlycine Max 9tttgatccat
gcccttcatt tgccgcttat taattaattt ggtaacagtc cgtactaatc 60agttacttat
ccttccccca tcataattaa tcttggtagt ctcgaatgcc acaacactga
120ctagtctctt ggatcataag aaaaagccaa ggaacaaaag aagacaaaac
acaatgagag 180tatcctttgc atagcaatgt ctaagttcat aaaattcaaa
caaaaacgca atcacacaca 240gtggacatca cttatccact agctgatcag
gatcgccgcg tcaagaaaaa aaaactggac 300cccaaaagcc atgcacaaca
acacgtactc acaaaggtgt caatcgagca gcccaaaaca 360ttcaccaact
caacccatca tgagccctca catttgttgt ttctaaccca acctcaaact
420cgtattctct tccgccacct catttttgtt tatttcaaca cccgtcaaac
tgcatgccac 480cccgtggcca aatgtccatg catgttaaca agacctatga
ctataaatag ctgcaatctc 540ggcccaggtt ttcatcatca agaaccagtt
caatatccta gtacaccgta ttaaagaatt 600taagatatac tgc
613101000DNAGlycine Max 10aactcagcat ctattttttt tctctcaacc
tttttcacat cttaagtagt ctgcaccctt 60tatatatata acttatttct taccttttac
attatgtaac ttttatcacc aaaaccaaca 120actttaaatt tttattaaat
agactccaca agtaacttga cactcttaca ttcatcgaca 180ttaactttta
tctgttttat aaatattatt gtgatataat ttaatcaaaa taaccacaaa
240ctttcataaa aggttcttat taagcatggc atttaataag caaaaacaac
tcaatcactt 300tcatatagga ggtagcctaa gtacgtactc aaaatgtcaa
caaataaaaa aaaagttgct 360ttaataatgc caaaacaaat taataaaaca
cttacaacac cggatttttt ttaattaaaa 420tgtgccattt aggataaata
gttaatattt ttaataatta tttaaaaagc cgtatctact 480aaaatgattt
ttatttggtt gaaaatgtta atatgtttaa atcaacacaa tctatcaaaa
540ttaaactaaa aaaaaaataa gtgtacgtgg ttaacattag tacagtaata
taagaggaaa 600atgagaaatt aagaaattga aagcgagtct aatttataaa
ttatgaacct gcatatataa 660aaggaaagaa agaatccagg aagaaaagaa
atgaaaccat gcatggtccc ctcgtcatca 720cgagtttctg ccatttgcaa
tagaaacact gaaacacctt tctctttgtc acttaattga 780gatgccgaag
ccacctcaca ccatgaactt catgaggtgt agcacccaag gcttccatag
840ccatgcatac tgaagaatgt ctcaagctca gcaccctact tctgtgacgt
gtccctcatt 900caccttcctc tcttccctat aaataaccac gcctcaggtt
ctccgcttca caacacaaac 960attctctcca ttggtcctta aacactcatc
agtcatcacc 1000111620DNAGlycine Max 11atggccgttt caaccacttt
ctccggtgcc aaattggagg ctctattgct caaatgttct 60tcctcctctt cctcaccacc
gccttcaagg tcatcattca ccacttttcc cggccaaaac 120agaagaaccc
tcattcagag aggggttatt cgctgcgacg ctcagccctc tgatgcatca
180tctgttgctc caaataatgc caccgctctc tccgctcttg agcagctcaa
gacttctgca 240gctgatagat atacaaagga aagaagcagc attatcgcca
ttgggctcag tgtgcacact 300gcacctgtgg aaatgcgtga aaaacttgcc
attccagaag cagaatggcc tagagctatt 360gcagagctgt gtagtctgaa
tcatattgaa gaagcagctg ttctgagtac ctgcaatcga 420atggagatat
atgttcttgc cctgtcccaa catcgtggtg tcaaagaagt catggaatgg
480atgtcaaaaa caagttctgt ccctgtttca gagcttagcc agcaccggtt
tttactttac 540aacaatgatg ccacacagca tctttttgaa gtatcagcag
gtcttgactc tcttgttttg 600ggggaaggtc aaatcctttc tcaggttaag
caagttgtta aagttggaca aggagttaac 660ggctttggga gaaatatcag
tgggctattc aagcatgcaa ttactgtcgg gaaaagggtt 720agaactgaga
ctaatattgc ttctggggca gtttctgtga gctcagctgc cgttgagttg
780gcctatatga agttacctga agcctcacac gataatgcca ggatgttggt
tattggtgct 840ggcaagatgg gaaagcttgt gatcaaacat ttggtggcaa
aaggttgcaa aaagatggtg 900gttgtcaata gaactgagga gagagttgct
gcaatacgtg aagaactgaa ggatattgag 960attatctaca aacccctttc
agaaatgctc acctgtgctg gcgaagcaga tttagttttc 1020accagtactg
catcagaaaa cccattattc ttgaaagaac atgtcaagga ccttcctcct
1080gcaagtcaag aagttggagg ccgtcgcttt ttcattgata tctctgttcc
ccggaatgtg 1140ggttcatgtg tctcagacct tgagtctgtg cgagtttaca
atgttgacga ccttaaagag 1200gttgtggctg ccaataaaga ggatcgccta
agaaaagcaa tggaagcaca ggcaatcatt 1260gctgaagaat ctaagcaatt
cgaagcttgg agggactcac tggaaactgt tcctactatt 1320aagaaattga
gggcttatgc tgaaagaatc aggcttgctg agcttgagaa gtgcttaggt
1380aagatgggtg atgatatacc aaagaaaacg cggagagctg tggatgacct
tagtcggggt 1440atagtgaata agttgcttca tggtccaatg caacatttaa
ggtgtgatgg gaacgacagc 1500cggactctta gtgagacact ggagaacatg
aatgctttga ataggatgtt caaccttgag 1560acagaaatat ctgttttgga
ggagaagatt cgagcaaagg tcgaacaaaa ccagaaatga 162012539PRTGlycine Max
12Met Ala Val Ser Thr Thr Phe Ser Gly Ala Lys Leu Glu Ala Leu Leu1
5 10 15Leu Lys Cys Ser Ser Ser Ser Ser Ser Pro Pro Pro Ser Arg Ser
Ser 20 25 30Phe Thr Thr Phe Pro Gly Gln Asn Arg Arg Thr Leu Ile Gln
Arg Gly 35 40 45Val Ile Arg Cys Asp Ala Gln Pro Ser Asp Ala Ser Ser
Val Ala Pro 50 55 60Asn Asn Ala Thr Ala Leu Ser Ala Leu Glu Gln Leu
Lys Thr Ser Ala65 70 75 80Ala Asp Arg Tyr Thr Lys Glu Arg Ser Ser
Ile Ile Ala Ile Gly Leu 85 90 95Ser Val His Thr Ala Pro Val Glu Met
Arg Glu Lys Leu Ala Ile Pro 100 105 110Glu Ala Glu Trp Pro Arg Ala
Ile Ala Glu Leu Cys Ser Leu Asn His 115 120 125Ile Glu Glu Ala Ala
Val Leu Ser Thr Cys Asn Arg Met Glu Ile Tyr 130 135 140Val Leu Ala
Leu Ser Gln His Arg Gly Val Lys Glu Val Met Glu Trp145 150 155
160Met Ser Lys Thr Ser Ser Val Pro Val Ser Glu Leu Ser Gln His Arg
165 170 175Phe Leu Leu Tyr Asn Asn Asp Ala Thr Gln His Leu Phe Glu
Val Ser 180 185 190Ala Gly Leu Asp Ser Leu Val Leu Gly Glu Gly Gln
Ile Leu Ser Gln 195 200 205Val Lys Gln Val Val Lys Val Gly Gln Gly
Val Asn Gly Phe Gly Arg 210 215 220Asn Ile Ser Gly Leu Phe Lys His
Ala Ile Thr Val Gly Lys Arg Val225 230 235 240Arg Thr Glu Thr Asn
Ile Ala Ser Gly Ala Val Ser Val Ser Ser Ala 245 250 255Ala Val Glu
Leu Ala Tyr Met Lys Leu Pro Glu Ala Ser His Asp Asn 260 265 270Ala
Arg Met Leu Val Ile Gly Ala Gly Lys Met Gly Lys Leu Val Ile 275 280
285Lys His Leu Val Ala Lys Gly Cys Lys Lys Met Val Val Val Asn Arg
290 295 300Thr Glu Glu Arg Val Ala Ala Ile Arg Glu Glu Leu Lys Asp
Ile Glu305 310 315 320Ile Ile Tyr Lys Pro Leu Ser Glu Met Leu Thr
Cys Ala Gly Glu Ala 325 330 335Asp Leu Val Phe Thr Ser Thr Ala Ser
Glu Asn Pro Leu Phe Leu Lys 340 345 350Glu His Val Lys Asp Leu Pro
Pro Ala Ser Gln Glu Val Gly Gly Arg 355 360 365Arg Phe Phe Ile Asp
Ile Ser Val Pro Arg Asn Val Gly Ser Cys Val 370 375 380Ser Asp Leu
Glu Ser Val Arg Val Tyr Asn Val Asp Asp Leu Lys Glu385 390 395
400Val Val Ala Ala Asn Lys Glu Asp Arg Leu Arg Lys Ala Met Glu Ala
405 410 415Gln Ala Ile Ile Ala Glu Glu Ser Lys Gln Phe Glu Ala Trp
Arg Asp 420 425 430Ser Leu Glu Thr Val Pro Thr Ile Lys Lys Leu Arg
Ala Tyr Ala Glu 435 440 445Arg Ile Arg Leu Ala Glu Leu Glu Lys Cys
Leu Gly Lys Met Gly Asp 450 455 460Asp Ile Pro Lys Lys Thr Arg Arg
Ala Val Asp Asp Leu Ser Arg Gly465 470 475 480Ile Val Asn Lys Leu
Leu His Gly Pro Met Gln His Leu Arg Cys Asp 485 490 495Gly Asn Asp
Ser Arg Thr Leu Ser Glu Thr Leu Glu Asn Met Asn Ala 500 505 510Leu
Asn Arg Met Phe Asn Leu Glu Thr Glu Ile Ser Val Leu Glu Glu 515 520
525Lys Ile Arg Ala Lys Val Glu Gln Asn Gln Lys 530
535131446DNAGlycine Max 13atgaacgcaa cctcatactc tgctcttcct
tctacgttcc gcagtctcca tcatcggaat 60ttctcagcgt tttgttctga tatccaaaat
cctggctatg ttgattgcca ttcaaattgt 120aataagtcta catctcaagc
gtctttgttt ttgtgttccg actccaacag tagaagaaat 180ggtgtttttg
gtagaccact ttgtgtgaac ccctctggca ggagaaacct agttggtcca
240gctttttatt ctctggagac tagtgcttat gacgtggctg ctttagaatc
tccttcccgt 300gttgcagaag aaaaagttgg tgtgctgctt ctcaatctag
gaggaccaga gacattgagt 360gacgtgcaac cttttctgtt taatcttttt
gcagatcctg atatcattcg tcttccaagg 420ttgtttcggt ttctccagcg
accattggca aaattgattt ctgtacttcg ggctcctaaa 480tccaaggaag
ggtatgctgc tattggtggt ggctctcctt tacgaaaaat tacagatgac
540caggcacttg caattaaaat ggctttggaa gcaaagggca tctcttcaaa
tgtctacgtt 600gggatgcgat actggtaccc atttactgaa gaagcaattc
agcaaattaa gagggacaga 660ataacaaggc ttgtggtact acccctttat
ccccagtttt ctatatccac aactggatca 720agcatccgtg ttcttgagca
tatattcagg gaagatgcct acttgtctaa gctccctgtt 780tccattataa
actcttggta tcaacgagaa ggttatatta agtcaatggc taacttaatt
840cagaaagagc tccagagttt ttctgaacca aaagaggtaa tgatattttt
cagtgcccat 900ggtgtacctg tcagttacgt tgaggaagct ggggatccat
accgagacca aatggaggag 960tgcatcttct tgatcatgca agagttgaaa
gctagaggaa ttagtaatga gcacactctt 1020gcttatcaga gtcgagtggg
tcctgtacag tggctgaaac catatactga tgaagttctc 1080gttgagcttg
gccaaaaagg tgtgaagagt cttttagctg ttccagtgag ttttgtgagt
1140gagcatattg aaacccttga agaaattgac atggagtaca aggaattggc
tcttgaatct 1200ggcatcaaga attgggcacg tgtacctgcc cttggtgtta
ccccttcctt cattacagat 1260ttagcagatg cagtaataga agctctccca
tcagcaacag caatatatgc accgaccaga 1320acctctgaag atgttgatca
tgacccagtt agatatttta tcaagatgtt ctttggttca 1380atcttggcat
tcatcttgtt cttgtcaccc aaaatgatca cggcattcag gaatcatgtc 1440atttag
144614481PRTGlycine Max 14Met Asn Ala Thr Ser Tyr Ser Ala Leu Pro
Ser Thr Phe Arg Ser Leu1 5 10 15His His Arg Asn Phe Ser Ala Phe Cys
Ser Asp Ile Gln Asn Pro Gly 20 25 30Tyr Val Asp Cys His Ser Asn Cys
Asn Lys Ser Thr Ser Gln Ala Ser 35 40 45Leu Phe Leu Cys Ser Asp Ser
Asn Ser Arg Arg Asn Gly Val Phe Gly 50 55 60Arg Pro Leu Cys Val Asn
Pro Ser Gly Arg Arg Asn Leu Val Gly Pro65 70 75 80Ala Phe Tyr Ser
Leu Glu Thr Ser Ala Tyr Asp Val Ala Ala Leu Glu 85 90 95Ser Pro Ser
Arg Val Ala Glu Glu Lys Val Gly Val Leu Leu Leu Asn 100 105 110Leu
Gly Gly Pro Glu Thr Leu Ser Asp Val Gln Pro Phe Leu Phe Asn 115 120
125Leu Phe Ala Asp Pro Asp Ile Ile Arg Leu Pro Arg Leu Phe Arg Phe
130 135 140Leu Gln Arg Pro Leu Ala Lys Leu Ile Ser Val Leu Arg Ala
Pro Lys145 150 155 160Ser Lys Glu Gly Tyr Ala Ala Ile Gly Gly Gly
Ser Pro Leu Arg Lys 165 170 175Ile Thr Asp Asp Gln Ala Leu Ala Ile
Lys Met Ala Leu Glu Ala Lys 180 185 190Gly Ile Ser Ser Asn Val Tyr
Val Gly Met Arg Tyr Trp Tyr Pro Phe 195 200 205Thr Glu Glu Ala Ile
Gln Gln Ile Lys Arg Asp Arg Ile Thr Arg Leu 210 215 220Val Val Leu
Pro Leu Tyr Pro Gln Phe Ser Ile Ser Thr Thr Gly Ser225 230 235
240Ser Ile Arg Val Leu Glu His Ile Phe Arg Glu Asp Ala Tyr Leu Ser
245 250 255Lys Leu Pro Val Ser Ile Ile Asn Ser Trp Tyr Gln Arg Glu
Gly Tyr 260 265 270Ile Lys Ser Met Ala Asn Leu Ile Gln Lys Glu Leu
Gln Ser Phe Ser 275 280 285Glu Pro Lys Glu Val Met Ile Phe Phe Ser
Ala His Gly Val Pro Val 290 295 300Ser Tyr Val Glu Glu Ala Gly Asp
Pro Tyr Arg Asp Gln Met Glu Glu305 310 315 320Cys Ile Phe Leu Ile
Met Gln Glu Leu Lys Ala Arg Gly Ile Ser Asn 325 330 335Glu His Thr
Leu Ala Tyr Gln Ser Arg Val Gly Pro Val Gln Trp Leu 340 345 350Lys
Pro Tyr Thr Asp Glu Val Leu Val Glu Leu Gly Gln Lys Gly Val 355 360
365Lys Ser Leu Leu Ala Val Pro Val Ser Phe Val Ser Glu His Ile Glu
370 375 380Thr Leu Glu Glu Ile Asp Met Glu Tyr Lys Glu Leu Ala Leu
Glu Ser385 390 395 400Gly Ile Lys Asn Trp Ala Arg Val Pro Ala Leu
Gly Val Thr Pro Ser 405 410 415Phe Ile Thr Asp Leu Ala Asp Ala Val
Ile Glu Ala Leu Pro Ser Ala 420 425 430Thr Ala Ile Tyr Ala Pro Thr
Arg Thr Ser Glu Asp Val Asp His Asp 435 440 445Pro Val Arg Tyr Phe
Ile Lys Met Phe Phe Gly Ser Ile Leu Ala Phe 450 455 460Ile Leu Phe
Leu Ser Pro Lys Met Ile Thr Ala Phe Arg Asn His Val465 470 475
480Ile15505DNAPhaseolus vulgaris 15agcttggcac tctgtggtct tttggttcat
gcatggatgc ttgcgcaaga aaaagacaaa 60gaacaaagaa aaaagacaaa acagagagac
aaaacgcaat cacacaacca actcaaatta 120gtcactggct gatcaagatc
gccgcgtcca tgtatgtcta aatgccatgc aaagcaacac 180gtgcttaaca
tgcactttaa atggctcacc catctcaacc cacacacaaa cacattgcct
240ttttcttcat catcaccaca accacctgta tatattcatt ctcttccgcc
acctcaattt 300cttcacttca acacacgtca acctgcatat gcgtgtcatc
ccatgcccaa atctccatgc 360atgttccaac caccttctct cttatataat
acctataaat acctctaata tcactcactt 420ctttcatcat ccatccatcc
agagtactac tactctacta ctataatacc ccaacccaac 480tcatattcaa
tactactcta ctaac 50516993DNABrassica napus 16ctgcggatca agcagctttc
atattaagca taccaaagcg taagatggtg gatgaaactc 60aagagactct ccgcaccacc
gcctttccaa gtactcatgt caaggttggt ttctttagct 120ttgaacacag
atttggatct ttttgttttg tttccatata cataggacct gagagctttt
180ggttgaattt tttttttttc aggacaaatg ggcgaagaat ctgtacattg
catcaatatg 240ctatggcagg acagtgtgct gatgatacac acttaagcat
catgtgttgt gttagaaagc 300cgaagacaat tggagcgagc ctcagggtcg
tcataatacc aatcaaagac gtaaaaccag 360acgcagtctc tttggttgaa
tgtgatgaaa gggatgtgtc ttggtatgta tgtacgagta 420acaaaagaga
agatgcaatt gagtagtaga aagatttgag agctttttaa agcccttcaa
480gtgtgtgctt ttatcttatt gatatcatcc atttgcgttg tttaatgcgt
ctttagatat 540gtttctgttt ctttctcagt gtctgaatat ctgataagtg
caatgtgaga aagccacacc 600aaaccaaaat attcaaatct tatattttta
ataatgtcga atcactcgga gttgccacct 660tctgtgccaa ttgtgctgaa
tctatcacac taaaaaaaac atttcttcaa ggtaatgact 720tgtggactat
gttctgaatt ctcattaagt ttttattttt tgaagtttaa gtttttacct
780tcttttttga aaaatatcgt tcataagatg tcacgccagg acatgagcta
cacatcacat 840attagcatgc agatgcggac gatttgtcac tcacttcaaa
cacctaaaag agcttctctc 900tcacagcaca cacacatatg catgcaatat
ttacacgtga tcgccatgca aatctccatt 960ctcacctata aattagagcc
tcggcttcac tct 9931719DNAGlycine Max 17gtttgctgag gctgctctc
191820DNAGlycine Max 18gaaagcaaaa gcagaagaaa 201920DNAGlycine Max
19gtggggtagc acatacaaaa 20204825DNAGlycine Max 20aactcagcat
ctattttttt tctctcaacc tttttcacat cttaagtagt ctgcaccctt 60tatatatata
acttatttct taccttttac attatgtaac ttttatcacc aaaaccaaca
120actttaaatt tttattaaat agactccaca agtaacttga cactcttaca
ttcatcgaca 180ttaactttta tctgttttat aaatattatt gtgatataat
ttaatcaaaa taaccacaaa 240ctttcataaa aggttcttat taagcatggc
atttaataag caaaaacaac tcaatcactt 300tcatatagga ggtagcctaa
gtacgtactc aaaatgtcaa caaataaaaa aaaagttgct 360ttaataatgc
caaaacaaat taataaaaca cttacaacac cggatttttt ttaattaaaa
420tgtgccattt aggataaata gttaatattt ttaataatta tttaaaaagc
cgtatctact 480aaaatgattt ttatttggtt gaaaatgtta atatgtttaa
atcaacacaa tctatcaaaa 540ttaaactaaa aaaaaaataa gtgtacgtgg
ttaacattag tacagtaata taagaggaaa 600atgagaaatt aagaaattga
aagcgagtct aatttataaa ttatgaacct gcatatataa 660aaggaaagaa
agaatccagg aagaaaagaa atgaaaccat gcatggtccc ctcgtcatca
720cgagtttctg ccatttgcaa tagaaacact gaaacacctt tctctttgtc
acttaattga 780gatgccgaag ccacctcaca ccatgaactt catgaggtgt
agcacccaag gcttccatag 840ccatgcatac tgaagaatgt ctcaagctca
gcaccctact tctgtgacgt gtccctcatt 900caccttcctc tcttccctat
aaataaccac gcctcaggtt ctccgcttca caacacaaac 960attctctcca
ttggtcctta aacactcatc agtcatcacc atggccaagc tagttttttc
1020cctttgtttt ctgcttttca gtggctgctg cttcgctttc agttccagag
agcagcctca 1080gcaaaacgag tgccagatcc aaaaactcaa tgccctcaaa
ccggataacc gtatagagtc 1140agaaggtggg ttcattgaga catggaaccc
taacaacaag ccattccagt gtgccggtgt 1200tgccctctct cgctgcaccc
tcaaccgcaa cgcccttcgt agaccttcct acaccaacgg 1260tccccaggaa
atctacatcc aacaaggtcc atcttgtcca aacttcacat ataaatatat
1320aatagactta aatatgttta agggtttgat aaatgaggga attttatttt
agatttttaa 1380taatttcttt tttattttga gtttttatat attaaaattt
ttgttttgat ttcttcaata 1440tgacgtaaca taatcatatc attgataatg
ttggattcct aatttttttg tttgtttgtt 1500gttttgtaaa atgaataggt
aagggtattt ttggcatgat atacccgggt tgtcctagca 1560catttgaaga
gcctcaacaa cctcaacaaa gaggacaaag cagcagacca caagaccgtc
1620accagaagat ctataacttc agagagggtg atttgatcgc agtgcctact
ggtgttgcat 1680ggtggatgta caacaatgaa gacactcctg ttgttgccgt
ttctattatt gacaccaaca 1740gcttggagaa ccagctcgac cagatgccta
gggtgagcca catagcaata ttagatatta 1800taattcttta aaggtttaaa
tatcatttta gttcgtggag ttgcactttc taatttagta 1860cctatagatt
aaaatatgcc aattggatcc ttatagttgt gtttttttat ccaatttggt
1920tcttgtcttg aaataaatgg acaatattgt agctgataaa aaaaggaaac
tggactacat 1980tgtaacgtta agattagaat tcttaagttc taatactagc
tggttacgga ttgacaacta 2040tttgttttga caattcttgg cagagattct
atcttgctgg gaaccaagag caagagtttc 2100taaaatatca gcagcagcag
caaggaggtt cccaaagcca gaaaggaaag catcagcaag 2160aagaagaaaa
cgaaggaggc agcatattga gtggcttcac cctggaattc ttggaacatg
2220cattcagcgt ggacaagcag atagcgaaaa acctacaagg agagaacgaa
ggggaagaca 2280agggagccat tgtgacagtg aaaggaggtc tgagcgtgat
aaaaccaccc acggacgagc 2340agcaacaaag accccaggaa gaggaagaag
aagaagagga tgagaagcca cagtgcaagg 2400gtaaagacaa acactgccaa
cgcccccgag gaagccaaag caaaagcaga agaaatggca 2460ttgacgagac
catatgcacc atgagacttc gccacaacat tggccagact tcatcacctg
2520acatctacaa ccctcaagcc ggtagcgtca caaccgccac cagccttgac
ttcccagccc 2580tctcgtggct cagactcagt gctgagtttg gatctctccg
caaggtacgt acatcattca 2640tcaaagatca acatacattt atacattaaa
ctaatatttg tttccaaata tttattaatt 2700ttattgataa ttaatttttt
tagaaaattt gtttgatcac ttttaatgga gtctttcatc 2760ttaattacat
tatttatact tacactaatg atttgttgat taataataat cttagataca
2820ctataaaatg tgtgacggag ttatcttaac acttgcatgg attctatctt
ttctgtcttt 2880atatatagaa atagagagaa aaaaagaaga aaagattgat
gaaaaaagca aaacaaaaaa 2940tagtattatt ataaaaatat tggatgaatt
tgttgtgact cttgcatgca ttgatgtacg 3000atgcagaatg caatgttcgt
gccacactac aacctgaacg cgaacagcat aatatacgca 3060ttgaatggac
gggcattgat acaagtggtg aattgcaacg gtgagagagt gtttgatgga
3120gagctgcaag agggacgggt gctgatcgtg ccacaaaact ttgtggtggc
tgcaagatca 3180cagagtgaca acttcgagta tgtgtcattc aagaccaatg
atacacccat gatcggcact 3240cttgcagggg caaactcatt gttgaacgca
ttaccagagg aagtgattca gcacactttc 3300aacctaaaaa gccagcaggc
caggcagata aagaacaaca accctttcaa gttcctggtt 3360ccacctcagg
agtctcagaa gagagctgtg gcttagagcc ctttttgtat gtgctacccc
3420acttttgtct ttttggcaat agtgctagca accaataaat aataataata
ataatgaata 3480agaaaacaaa ggctttagct tgccttttgt tcactgtaaa
ataataatgt aagtactctc 3540tataatgagt cacgaaactt ttgcgggaat
aaaaggagaa attccaatga gttttctgtc 3600aaatcttctt ctgtctctct
ctctctctct ttttttttcc tttcttctga gcttcttgca 3660aaacaaaagg
caaacaataa cgattggtcc aatgatagtt agcttaatcg atgatatctt
3720taggaagtgt tggcaggaca ggacatgatg tagaagacta aaattgaaag
tattgcagac 3780ccaatagttg aagattaact ttaagaatta agacgtctgt
taggaagaag ccattgcaac 3840caataaagca tagtagatag ttagttaaat
ctgttgggtt agttaccaac aagttgttaa 3900tactgttgag ttagttatgc
atgtaaataa ttgtagtata taagagttgc agtaatgaat 3960aaaagacatg
cagaaaaact tagaatcagt ttagttgtta tggagagggt gagatagtgc
4020ctttgttcga ggcaaggctc ctcctaagaa agtctgaaac tgattttaaa
cttttctgca 4080tggcttttat tcattattgc aactcttata tactacaatt
atttacatgc ataactaact 4140caacaaacta atggtattaa caacttgttg
gtaactaacc caacagattt aactaactat 4200ctactatgct ttattggttg
caatggcttc ttcctaacaa cgtcttatca ggttcttcat 4260gacttggagc
tcaacccaac ttggaaagtt cgagagtatt tggaccattg tgctttgtgt
4320cttcaaacat aaaacatcgc tccaaattta acatgggagc taaaaaatgt
gtttttctgg 4380gattttaatt ttcaacagag tcaaggatgg tgttgcatat
gatgtcttga tgtccattgt 4440ccacactaaa tagatattgg tttcaagaaa
tattaatttc attttcatga ctttcaattc 4500ataaacctta aacgaatatt
aatttaaaat ctatcctcaa atgataaatt taaaaaaaaa 4560ttacccccaa
tcggtaattt gactcacaag ttagttagtt gatattttga agcttgaaat
4620tcgacatgga catcagacac aatatgagca cagacactct cgcatagcta
atgtgtaaaa 4680catagaatga caggacatca catatatttt tacacacaca
aaaaaagaac tctaataaaa 4740aaatatgggt agcttaacaa atatataaat
tgatggtaaa taatttactt tttaaaattc 4800atctatgttt ttttatatga taaca
482521495PRTGlycine Max 21Met Ala Lys Leu Val Phe Ser Leu Cys Phe
Leu Leu Phe Ser Gly Cys1 5 10 15Cys Phe Ala Phe Ser Ser Arg Glu Gln
Pro Gln Gln Asn Glu Cys Gln 20 25 30Ile Gln Lys Leu Asn Ala Leu Lys
Pro Asp Asn Arg Ile Glu Ser Glu 35 40 45Gly Gly Leu Ile Glu Thr Trp
Asn Pro Asn Asn Lys Pro Phe Gln Cys 50 55 60Ala Gly Val Ala Leu Ser
Arg Cys Thr Leu Asn Arg Asn Ala Leu Arg65 70 75 80Arg Pro Ser Tyr
Thr Asn Gly Pro Gln Glu Ile Tyr Ile Gln Gln Gly 85 90 95Lys Gly Ile
Phe Gly Met Ile Tyr Pro Gly Cys Pro Ser Thr Phe Glu 100 105 110Glu
Pro Gln Gln Pro Gln Gln Arg Gly Gln Ser Ser Arg Pro Gln Asp 115 120
125Arg His Gln Lys Ile Tyr Asn Phe Arg Glu Gly Asp Leu Ile Ala Val
130 135 140Pro Thr Gly Val Ala Trp Trp Met Tyr Asn Asn Glu Asp Thr
Pro Val145 150 155 160Val Ala Val Ser Ile Ile Asp Thr Asn Ser Leu
Glu Asn Gln Leu Asp 165 170 175Gln Met Pro Arg Arg Phe Tyr Leu Ala
Gly Asn Gln Glu Gln Glu Phe 180 185 190Leu Lys Tyr Gln Gln Glu Gln
Gly Gly His Gln Ser Gln Lys Gly Lys 195 200 205His Gln Gln Glu Glu
Glu Asn Glu Gly Gly Ser Ile Leu Ser Gly Phe 210 215 220Thr Leu Glu
Phe Leu Glu His Ala Phe Ser Val Asp Lys Gln Ile Ala225 230 235
240Lys Asn Leu Gln Gly Glu Asn Glu Gly Glu Asp Lys Gly Ala Ile Val
245 250 255Thr Val Lys Gly Gly Leu Ser Val Ile Lys Pro Pro Thr Asp
Glu Gln 260 265 270Gln Gln Arg Pro Gln Glu Glu Glu Glu Glu Glu Glu
Asp Glu Lys Pro 275 280 285Gln Cys Lys Gly Lys Asp Lys His Cys Gln
Arg Pro Arg Gly Ser Gln 290 295 300Ser Lys Ser Arg Arg Asn Gly Ile
Asp Glu Thr Ile Cys Thr Met Arg305 310 315 320Leu Arg His Asn Ile
Gly Gln Thr Ser Ser Pro Asp Ile Tyr Asn Pro 325 330 335Gln Ala Gly
Ser Val Thr Thr Ala Thr Ser Leu Asp Phe Pro Ala Leu 340 345 350Ser
Trp Leu Arg Leu Ser Ala Glu Phe Gly Ser Leu Arg Lys Asn Ala 355 360
365Met Phe Val Pro His Tyr Asn Leu Asn Ala Asn Ser Ile Ile Tyr Ala
370 375 380Leu Asn Gly Arg Ala Leu Ile Gln Val Val Asn Cys Asn Gly
Glu Arg385 390 395 400Val Phe Asp Gly Glu Leu Gln Glu Gly Arg Val
Leu Ile Val Pro Gln 405 410 415Asn Phe Val Val Ala Ala Arg Ser Gln
Ser Asp Asn Phe Glu Tyr Val 420 425 430Ser Phe Lys Thr Asn Asp Thr
Pro Met Ile Gly Thr Leu Ala Gly Ala 435 440 445Asn Ser Leu Leu Asn
Ala Leu Pro Glu Glu Val Ile Gln His Thr Phe 450 455 460Asn Leu Lys
Ser Gln Gln Ala Arg Gln Ile Lys Asn Asn Asn Pro Phe465 470 475
480Lys Phe Leu Val Pro Pro Gln Glu Ser Gln Lys Arg Ala Val Ala 485
490 495223565DNAGlycine Max 22ccagagagca gcctcagcaa acaatgcagg
acaactcagc atctattttt tttctctcaa 60cctttttcac atcttaagta gtctgcaccc
tttatatata taacttattt cttacctttt 120acattatgta acttttatca
ccaaaaccaa caactttaaa tttttattaa atagactcca 180caagtaactt
gacactctta cattcatcga cattaacttt tatctgtttt ataaatatta
240ttgtgatata atttaatcaa aataaccaca aactttcata aaaggttctt
attaagcatg 300gcatttaata agcaaaaaca actcaatcac tttcatatag
gaggtagcct aagtacgtac 360tcaaaatgtc aacaaataaa aaaaaagttg
ctttaataat gccaaaacaa attaataaaa 420cacttacaac accggatttt
ttttaattaa aatgtgccat ttaggataaa tagttaatat 480ttttaataat
tatttaaaaa gccgtatcta ctaaaatgat ttttatttgg ttgaaaatgt
540taatatgttt aaatcaacac aatctatcaa aattaaacta aaaaaaaaat
aagtgtacgt 600ggttaacatt agtacagtaa tataagagga aaatgagaaa
ttaagaaatt gaaagcgagt 660ctaatttata aattatgaac ctgcatatat
aaaaggaaag aaagaatcca ggaagaaaag 720aaatgaaacc atgcatggtc
ccctcgtcat cacgagtttc tgccatttgc aatagaaaca 780ctgaaacacc
tttctctttg tcacttaatt gagatgccga agccacctca caccatgaac
840ttcatgaggt gtagcaccca aggcttccat
agccatgcat actgaagaat gtctcaagct 900cagcacccta cttctgtgac
gtgtccctca ttcaccttcc tctcttccct ataaataacc 960acgcctcagg
ttctccgctt cacaacacaa acattctctc cattggtcct taaacactca
1020tcagtcatca ccatggccaa gctagttttt tccctttgtt ttctgctttt
cagtggctgc 1080tgcttcgctt tcagttccat gggtgctttc actgagaagc
aagaggcttt ggtgagtagc 1140tcattcgaag cattcaaggc aaacattcct
caatacagcg ttgtgttcta cacttcgata 1200ctggagaaag cacccgcagc
aaaggacttg ttctcgtttc tatctaatgg agtagatcct 1260agtaatccta
agctcacggg ccatgctgaa aagctttttg gattggtgcg tgactcagct
1320ggtcaactta aagcaaatgg aacagtagtg gctgatgccg cacttggttc
tatccatgcc 1380caaaaagcaa tcactgatcc tcagttcgtg gtggttaaag
aagcactgct gaaaacaata 1440aaggaggcag ttggggacaa atggagtgat
gaattgagca gtgcttggga agtagcctat 1500gatgaattgg cagcagctat
taagaaggca tttagcagaa gaaatggcat tgacgagacc 1560atatgcacca
tgagacttcg ccacaacatt ggccagactt catcacctga catctacaac
1620cctcaagccg gtagcgtcac aaccgccacc agccttgact tcccagccct
ctcgtggctc 1680agactcagtg ctgagtttgg atctctccgc aagaatgcaa
tgttcgtgcc acactacaac 1740ctgaacgcga acagcataat atacgcattg
aatggacggg cattgataca agtggtgaat 1800tgcaacggtg agagagtgtt
tgatggagag ctgcaagagg gacgggtgct gatcgtgcca 1860caaaactttg
tggtggctgc aagatcacag agtgacaact tcgagtatgt gtcattcaag
1920accaatgata cacccatgat cggcactctt gcaggggcaa actcattgtt
gaacgcatta 1980ccagaggaag tgattcagca cactttcaac ctaaaaagcc
agcaggccag gcagataaag 2040aacaacaacc ctttcaagtt cctggttcca
cctcaggagt ctcagaagag agctgtggct 2100tagagccctt tttgtatgtg
ctaccccact tttgtctttt tggcaatagt gctagcaacc 2160aataaataat
aataataata atgaataaga aaacaaaggc tttagcttgc cttttgttca
2220ctgtaaaata ataatgtaag tactctctat aatgagtcac gaaacttttg
cgggaataaa 2280aggagaaatt ccaatgagtt ttctgtcaaa tcttcttctg
tctctctctc tctctctttt 2340tttttccttt cttctgagct tcttgcaaaa
caaaaggcaa acaataacga ttggtccaat 2400gatagttagc ttaatcgatg
atatctttag gaagtgttgg caggacagga catgatgtag 2460aagactaaaa
ttgaaagtat tgcagaccca atagttgaag attaacttta agaattaaga
2520cgtctgttag gaagaagcca ttgcaaccaa taaagcatag tagatagtta
gttaaatctg 2580ttgggttagt taccaacaag ttgttaatac tgttgagtta
gttatgcatg taaataattg 2640tagtatataa gagttgcagt aatgaataaa
agacatgcag aaaaacttag aatcagttta 2700gttgttatgg agagggtgag
atagtgcctt tgttcgaggc aaggctcctc ctaagaaagt 2760ctgaaactga
ttttaaactt ttctgcatgg cttttattca ttattgcaac tcttatatac
2820tacaattatt tacatgcata actaactcaa caaactaatg gtattaacaa
cttgttggta 2880actaacccaa cagatttaac taactatcta ctatgcttta
ttggttgcaa tggcttcttc 2940ctaacaacgt cttatcaggt tcttcatgac
ttggagctca acccaacttg gaaagttcga 3000gagtatttgg accattgtgc
tttgtgtctt caaacataaa acatcgctcc aaatttaaca 3060tgggagctaa
aaaatgtgtt tttctgggat tttaattttc aacagagtca aggatggtgt
3120tgcatatgat gtcttgatgt ccattgtcca cactaaatag atattggttt
caagaaatat 3180taatttcatt ttcatgactt tcaattcata aaccttaaac
gaatattaat ttaaaatcta 3240tcctcaaatg ataaatttaa aaaaaaatta
cccccaatcg gtaatttgac tcacaagtta 3300gttagttgat attttgaagc
ttgaaattcg acatggacat cagacacaat atgagcacag 3360acactctcgc
atagctaatg tgtaaaacat agaatgacag gacatcacat atatttttac
3420acacacaaaa aaagaactct aataaaaaaa tatgggtagc ttaacaaata
tataaattga 3480tggtaaataa tttacttttt aaaattcatc tatgtttttt
tatatgataa caaggcgcgt 3540ttgaaagcaa aagcagaaga aatgg
3565232508DNAGlycine Max 23ccagagagca gcctcagcaa acaatgcagg
acaactcagc atctattttt tttctctcaa 60cctttttcac atcttaagta gtctgcaccc
tttatatata taacttattt cttacctttt 120acattatgta acttttatca
ccaaaaccaa caactttaaa tttttattaa atagactcca 180caagtaactt
gacactctta cattcatcga cattaacttt tatctgtttt ataaatatta
240ttgtgatata atttaatcaa aataaccaca aactttcata aaaggttctt
attaagcatg 300gcatttaata agcaaaaaca actcaatcac tttcatatag
gaggtagcct aagtacgtac 360tcaaaatgtc aacaaataaa aaaaaagttg
ctttaataat gccaaaacaa attaataaaa 420cacttacaac accggatttt
ttttaattaa aatgtgccat ttaggataaa tagttaatat 480ttttaataat
tatttaaaaa gccgtatcta ctaaaatgat ttttatttgg ttgaaaatgt
540taatatgttt aaatcaacac aatctatcaa aattaaacta aaaaaaaaat
aagtgtacgt 600ggttaacatt agtacagtaa tataagagga aaatgagaaa
ttaagaaatt gaaagcgagt 660ctaatttata aattatgaac ctgcatatat
aaaaggaaag aaagaatcca ggaagaaaag 720aaatgaaacc atgcatggtc
ccctcgtcat cacgagtttc tgccatttgc aatagaaaca 780ctgaaacacc
tttctctttg tcacttaatt gagatgccga agccacctca caccatgaac
840ttcatgaggt gtagcaccca aggcttccat agccatgcat actgaagaat
gtctcaagct 900cagcacccta cttctgtgac gtgtccctca ttcaccttcc
tctcttccct ataaataacc 960acgcctcagg ttctccgctt cacaacacaa
acattctctc cattggtcct taaacactca 1020tcagtcatca ccatgggtgc
tttcactgag aagcaagagg ctttggtgag tagctcattc 1080gaagcattca
aggcaaacat tcctcaatac agcgttgtgt tctacacttc gatactggag
1140aaagcacccg cagcaaagga cttgttctcg tttctatcta atggagtaga
tcctagtaat 1200cctaagctca cgggccatgc tgaaaagctt tttggattgg
tgcgtgactc agctggtcaa 1260cttaaagcaa atggaacagt agtggctgat
gccgcacttg gttctatcca tgcccaaaaa 1320gcaatcactg atcctcagtt
cgtggtggtt aaagaagcac tgctgaaaac aataaaggag 1380gcagttgggg
acaaatggag tgatgaattg agcagtgctt gggaagtagc ctatgatgaa
1440ttggcagcag ctattaagaa ggcattttag agcgcttttt gtatgtgcta
ccccactttt 1500gtctttttgg caatagtgct agcaaccaat aaataataat
aataataatg aataagaaaa 1560caaaggcttt agcttgcctt ttgttcactg
taaaataata atgtaagtac tctctataat 1620gagtcacgaa acttttgcgg
gaataaaagg agaaattcca atgagttttc tgtcaaatct 1680tcttctgtct
ctctctctct ctcttttttt ttcctttctt ctgagcttct tgcaaaacaa
1740aaggcaaaca ataacgattg gtccaatgat agttagctta atcgatgata
tctttaggaa 1800gtgttggcag gacaggacat gatgtagaag actaaaattg
aaagtattgc agacccaata 1860gttgaagatt aactttaaga attaagacgt
ctgttaggaa gaagccattg caaccaataa 1920agcatagtag atagttagtt
aaatctgttg ggttagttac caacaagttg ttaatactgt 1980tgagttagtt
atgcatgtaa ataattgtag tatataagag ttgcagtaat gaataaaaga
2040catgcagaaa aacttagaat cagtttagtt gttatggaga gggtgagata
gtgcctttgt 2100tcgaggcaag gctcctccta agaaagtctg aaactgattt
taaacttttc tgcatggctt 2160ttattcatta ttgcaactct tatatactac
aattatttac atgcataact aactcaacaa 2220actaatggta ttaacaactt
gttggtaact aacccaacag atttaactaa ctatctacta 2280tgctttattg
gttgcaatgg cttcttccta acaacgtctt atcaggttct tcatgacttg
2340gagctcaacc caacttggaa agttcgagag tatttggacc attgtgcttt
gtgtcttcaa 2400acataaaaca tcgctccaaa tttaacatgg gagctaaaaa
atgtgttttt ctgggatttt 2460aattttcaac agagaaggcg cgtttccttt
ttgtatgtgc taccccac 25082420DNAGlycine max 24gatacgtgga tatattattg
202520DNAGlycine max 25gaattaagcg atgaagatca 202620DNAGlycine max
26gttgtctcta tgttattgtt 202720DNAGlycine max 27gttgttgtgt
agcaatccga 202820DNAGlycine max 28ggcaacgagc gtggcatgct
202920DNAGlycine max 29gtcagtttgt gtctcattaa 203020DNAGlycine max
30gtcagtttct gtctcattac 2031603DNAGlycine Max 31atggcttcct
caatgatctc ctccccagct gttaccaccg tcaaccgtgc cggtgccggc 60atggttgctc
cattcaccgg cctcaaatcc atggctggct tccccacgag gaagaccaac
120aatgacatta cctccattgc tagcaacggt ggaagagtac aatgcatggg
tgctttcact 180gagaagcaag aggctttggt gagtagctca ttcgaagcat
tcaaggcaaa cattcctcaa 240tacagcgttg tgttctacac ttcgatactg
gagaaagcac ccgcagcaaa ggacttgttc 300tcgtttctat ctaatggagt
agatcctagt aatcctaagc tcacgggcca tgctgaaaag 360ctttttggat
tggtgcgtga ctcagctggt caacttaaag caaatggaac agtagtggct
420gatgccgcac ttggttctat ccatgcccaa aaagcaatca ctgatcctca
gttcgtggtg 480gttaaagaag cactgctgaa aacaataaag gaggcagttg
gggacaaatg gagtgatgaa 540ttgagcagtg cttgggaagt agcctatgat
gaattggcag cagctattaa gaaggcattt 600tag 60332200PRTGlycine Max
32Met Ala Ser Ser Met Ile Ser Ser Pro Ala Val Thr Thr Val Asn Arg1
5 10 15Ala Gly Ala Gly Met Val Ala Pro Phe Thr Gly Leu Lys Ser Met
Ala 20 25 30Gly Phe Pro Thr Arg Lys Thr Asn Asn Asp Ile Thr Ser Ile
Ala Ser 35 40 45Asn Gly Gly Arg Val Gln Cys Met Gly Ala Phe Thr Glu
Lys Gln Glu 50 55 60Ala Leu Val Ser Ser Ser Phe Glu Ala Phe Lys Ala
Asn Ile Pro Gln65 70 75 80Tyr Ser Val Val Phe Tyr Thr Ser Ile Leu
Glu Lys Ala Pro Ala Ala 85 90 95Lys Asp Leu Phe Ser Phe Leu Ser Asn
Gly Val Asp Pro Ser Asn Pro 100 105 110Lys Leu Thr Gly His Ala Glu
Lys Leu Phe Gly Leu Val Arg Asp Ser 115 120 125Ala Gly Gln Leu Lys
Ala Asn Gly Thr Val Val Ala Asp Ala Ala Leu 130 135 140Gly Ser Ile
His Ala Gln Lys Ala Ile Thr Asp Pro Gln Phe Val Val145 150 155
160Val Lys Glu Ala Leu Leu Lys Thr Ile Lys Glu Ala Val Gly Asp Lys
165 170 175Trp Ser Asp Glu Leu Ser Ser Ala Trp Glu Val Ala Tyr Asp
Glu Leu 180 185 190Ala Ala Ala Ile Lys Lys Ala Phe 195
2003320PRTGlycine max 33Lys Ala Asn Gly Thr Val Val Ala Asp Ala Ala
Leu Gly Ser Ile His1 5 10 15Ala Gln Lys Ala 203413PRTGlycine max
34Lys Ala Ile Thr Asp Pro Gln Phe Val Val Val Lys Glu1 5
1035100DNAGlycine max 35atgccccatt tttctctctc ccctctctcc tgtgcacctt
ctcctctccc acctcgccgc 60cgaatctttc tcgctcctcc ccgaaccgcc gcatcttccg
1003683PRTGlycine max 36Met Pro His Phe Ser Leu Ser Pro Leu Ser Cys
Ala Pro Ser Pro Leu1 5 10 15Pro Pro Arg Arg Arg Ile Phe Leu Ala Pro
Pro Arg Thr Ala Ala Ser 20 25 30Ser Ala Thr Asp Ala Ala Ser Ser Ser
Thr Ser Ser Ser Ala Ser Asn 35 40 45Phe Ala Pro Lys Val Val Val Thr
Arg Glu Arg Gly Lys Asn Ala Lys 50 55 60Leu Ile Ala Ala Leu Ala Lys
His Glu Ile Asn Cys Leu Glu Leu Pro65 70 75 80Leu Ile
Glu371413DNAGlycine max 37atggctgttt cggctatcac aggagcgagg
ctaaccctag ggatagggtt ggcgatacct 60ctttcctctc ccacgcgctc tcgaaccgtc
gcaatggccg tatccgtcga ccccaagacc 120gacaacaaac tcactcttac
caagtccgag gaagctttcg ctgctgccaa ggagctgatg 180cctggaggtg
tcaactcccc agttcgtgcc ttcaaatccg tgggtggtca accaattgtg
240attgattcag tcaaagggtc tcgtatgtgg gacatcgacg gcaatgagta
cattgactac 300gtcggttctt ggggtcccgc aatcattggt cacgctgatg
atcaagtgct ttcagctctg 360gttgaaacca tgaagaaagg aactagcttt
ggtgcaccct gtctgctgga aaacactttg 420gcagagctgg ttatcaatgc
ggtccccagc attgaaatgg ttcgctttgt caattcaggc 480accgaagctt
gcatgggtgc actacgtctc gcccgagctt ataccggaag agagaagatc
540atcaagtttg agggctgtta ccatggccat gctgatcctt ttcttgttaa
ggcaggtagt 600ggagttgcca ccttgggact tcctgattct cccggtgtcc
ccaaagctgc cacttttgaa 660acccttacag ccccctacaa tgacactgcc
gccgttgaga agctctttga ggctaacaaa 720ggagaaatcg ctgctgtttt
cctcgaacct gttgttggaa acgctggttt cattgttcct 780aaacccgatt
ttcataattt cttgcgcaag atcaccaagg agaacaatac ccttcttgtg
840tttgatgaag ttatgactgg gtttcgtttg tcatacggag gtgctcaaga
gtattttggc 900ataactcctg atataacaac tctaggaaag atcattggtg
gaggtctgcc ggtgggggct 960tatggaggga ggagggatat tatggagaag
gtggcaccag ctggcccaat gtatcaggct 1020gggaccttga gtgggaaccc
tttggccatg actgcaggaa tacagaccct gcagcgtatt 1080aaggagccag
gaacttatga gtacttggac aaaatcaccg gtgagcttgt tcagggcatt
1140attgaagctg ggaagagggc aggccatgca atatgtggtg gtcatataag
ggggatgttt 1200gggtttttct tcacagaagg accagtgtat aattttgcag
atgccaaaaa gagtgatacg 1260gacaagtttg ctaggttctt ttggggaatg
ttggcggaag gtgtctattt ggcaccttcc 1320cagtttgagg ctggctttac
cagcttggca catacctctg atgacataaa aaagacaata 1380gccgctgctg
agaaggtttt cagggagatc tga 141338470PRTGlycine max 38Met Ala Val Ser
Ala Ile Thr Gly Ala Arg Leu Thr Leu Gly Ile Gly1 5 10 15Leu Ala Ile
Pro Leu Ser Ser Pro Thr Arg Ser Arg Thr Val Ala Met 20 25 30Ala Val
Ser Val Asp Pro Lys Thr Asp Asn Lys Leu Thr Leu Thr Lys 35 40 45Ser
Glu Glu Ala Phe Ala Ala Ala Lys Glu Leu Met Pro Gly Gly Val 50 55
60Asn Ser Pro Val Arg Ala Phe Lys Ser Val Gly Gly Gln Pro Ile Val65
70 75 80Ile Asp Ser Val Lys Gly Ser Arg Met Trp Asp Ile Asp Gly Asn
Glu 85 90 95Tyr Ile Asp Tyr Val Gly Ser Trp Gly Pro Ala Ile Ile Gly
His Ala 100 105 110Asp Asp Gln Val Leu Ser Ala Leu Val Glu Thr Met
Lys Lys Gly Thr 115 120 125Ser Phe Gly Ala Pro Cys Leu Leu Glu Asn
Thr Leu Ala Glu Leu Val 130 135 140Ile Asn Ala Val Pro Ser Ile Glu
Met Val Arg Phe Val Asn Ser Gly145 150 155 160Thr Glu Ala Cys Met
Gly Ala Leu Arg Leu Ala Arg Ala Tyr Thr Gly 165 170 175Arg Glu Lys
Ile Ile Lys Phe Glu Gly Cys Tyr His Gly His Ala Asp 180 185 190Pro
Phe Leu Val Lys Ala Gly Ser Gly Val Ala Thr Leu Gly Leu Pro 195 200
205Asp Ser Pro Gly Val Pro Lys Ala Ala Thr Phe Glu Thr Leu Thr Ala
210 215 220Pro Tyr Asn Asp Thr Ala Ala Val Glu Lys Leu Phe Glu Ala
Asn Lys225 230 235 240Gly Glu Ile Ala Ala Val Phe Leu Glu Pro Val
Val Gly Asn Ala Gly 245 250 255Phe Ile Val Pro Lys Pro Asp Phe His
Asn Phe Leu Arg Lys Ile Thr 260 265 270Lys Glu Asn Asn Thr Leu Leu
Val Phe Asp Glu Val Met Thr Gly Phe 275 280 285Arg Leu Ser Tyr Gly
Gly Ala Gln Glu Tyr Phe Gly Ile Thr Pro Asp 290 295 300Ile Thr Thr
Leu Gly Lys Ile Ile Gly Gly Gly Leu Pro Val Gly Ala305 310 315
320Tyr Gly Gly Arg Arg Asp Ile Met Glu Lys Val Ala Pro Ala Gly Pro
325 330 335Met Tyr Gln Ala Gly Thr Leu Ser Gly Asn Pro Leu Ala Met
Thr Ala 340 345 350Gly Ile Gln Thr Leu Gln Arg Ile Lys Glu Pro Gly
Thr Tyr Glu Tyr 355 360 365Leu Asp Lys Ile Thr Gly Glu Leu Val Gln
Gly Ile Ile Glu Ala Gly 370 375 380Lys Arg Ala Gly His Ala Ile Cys
Gly Gly His Ile Arg Gly Met Phe385 390 395 400Gly Phe Phe Phe Thr
Glu Gly Pro Val Tyr Asn Phe Ala Asp Ala Lys 405 410 415Lys Ser Asp
Thr Asp Lys Phe Ala Arg Phe Phe Trp Gly Met Leu Ala 420 425 430Glu
Gly Val Tyr Leu Ala Pro Ser Gln Phe Glu Ala Gly Phe Thr Ser 435 440
445Leu Ala His Thr Ser Asp Asp Ile Lys Lys Thr Ile Ala Ala Ala Glu
450 455 460Lys Val Phe Arg Glu Ile465 47039918DNAGlycine max
39atgcttcttc aaacgaggca atttggcaca gcgtgtccat gggtcagact cacacccgca
60agcaaaagaa atagctcagt ctcattgaga aggagcaatt gtggcccaaa atgttcggtg
120tcccaattgg aattgaagag tgagagtgag agcaaaacga agccgtatcc
agcagaggca 180tcgaggacca taatggagtt ggctaaggtg ggcacgctgt
gtacgttgac ccaagagggt 240ttgccccttg gtactggggt tcgattcgcg
gttgaccctg aacatggcac tcccttgttc 300tgcttcaatt ccactgacaa
cactaacatc ccctcttctc ttcatgttca gtttgaacaa 360tttggattgc
gcactcctca gtgcactatt caaggaaccc ttaccaaacc acaggatcca
420aagcgtcttg tttctgtgtg gaggaaaagg tttggagaag aagttgatcg
agattttatt 480tacattattg ccgtggatcg ggtactccaa ttggaagaca
ttcaggagga tggcatatgg 540gtcacctctt cagattacaa aaatgctcaa
cccgatcctc ttcgagactc tgcacacaac 600ttagtcactg aaatcaacac
caacaatatg gaagacatta ctcgcttttg caatgtctac 660gttgatttgg
atttcctggt ctccgaggca aagatgctat gggttgatcg cttgggcttt
720gacatgcgtt tgtcttcccc tcacaaaggc gtgtttgacg tccgcattcc
tttccccaga 780gaagtcaccg atgagaaagg tgccaagtcg acgtttaatt
gtatgtcaca actcgcttgg 840gaggtagaaa gaaacttcca acatccagac
ttttcaaagg ttaaagagtt gaagccagtc 900aagtcccctt ttctgtaa
91840305PRTGlycine max 40Met Leu Leu Gln Thr Arg Gln Phe Gly Thr
Ala Cys Pro Trp Val Arg1 5 10 15Leu Thr Pro Ala Ser Lys Arg Asn Ser
Ser Val Ser Leu Arg Arg Ser 20 25 30Asn Cys Gly Pro Lys Cys Ser Val
Ser Gln Leu Glu Leu Lys Ser Glu 35 40 45Ser Glu Ser Lys Thr Lys Pro
Tyr Pro Ala Glu Ala Ser Arg Thr Ile 50 55 60Met Glu Leu Ala Lys Val
Gly Thr Leu Cys Thr Leu Thr Gln Glu Gly65 70 75 80Leu Pro Leu Gly
Thr Gly Val Arg Phe Ala Val Asp Pro Glu His Gly 85 90 95Thr Pro Leu
Phe Cys Phe Asn Ser Thr Asp Asn Thr Asn Ile Pro Ser 100 105 110Ser
Leu His Val Gln Phe Glu Gln Phe Gly Leu Arg Thr Pro Gln Cys 115 120
125Thr Ile Gln Gly Thr Leu Thr Lys Pro Gln Asp Pro Lys Arg Leu Val
130 135 140Ser Val Trp Arg Lys Arg Phe Gly Glu Glu Val Asp Arg Asp
Phe Ile145 150 155 160Tyr Ile Ile Ala Val Asp Arg Val Leu Gln Leu
Glu Asp Ile Gln Glu 165 170 175Asp Gly Ile Trp Val Thr Ser
Ser Asp Tyr Lys Asn Ala Gln Pro Asp 180 185 190Pro Leu Arg Asp Ser
Ala His Asn Leu Val Thr Glu Ile Asn Thr Asn 195 200 205Asn Met Glu
Asp Ile Thr Arg Phe Cys Asn Val Tyr Val Asp Leu Asp 210 215 220Phe
Leu Val Ser Glu Ala Lys Met Leu Trp Val Asp Arg Leu Gly Phe225 230
235 240Asp Met Arg Leu Ser Ser Pro His Lys Gly Val Phe Asp Val Arg
Ile 245 250 255Pro Phe Pro Arg Glu Val Thr Asp Glu Lys Gly Ala Lys
Ser Thr Phe 260 265 270Asn Cys Met Ser Gln Leu Ala Trp Glu Val Glu
Arg Asn Phe Gln His 275 280 285Pro Asp Phe Ser Lys Val Lys Glu Leu
Lys Pro Val Lys Ser Pro Phe 290 295 300Leu305411149DNAGlycine max
41atggaagcca tattcgccac caaacccgct tcccattctc tcctcttaac taaactctct
60ccgaatccca aacacttgtt ccctccacac caacaatcct ttcacaacat ccgccacaaa
120cccacgcgct tccgccccgt caccgctgtt ttccaaaacc aacatcaaca
agatgcagct 180gcagcttcca accacaccga agatgagtcc tacggcgaag
tcaaaggcat cattggaagc 240agagccttgg aagccgccac cggaatggag
tacctcatcg agtggaacga cggccacgcg 300ccgtcctggg ttcccgccga
cttcatagcc aaagacgtcg tcgacgagta cgaaactccc 360tggtggactg
ccgccaagaa agccgacgag tccgcgttga aaaacttaac caaatccgac
420gacggccgcg acgtcgacgc cgtggacgcc gacggccgca ctgcgctcct
cttcgtcgcc 480ggactcggct cggagtcctg cgtgaagctg ctagcggagg
ccggcgcgaa tctggaccac 540cgcgaccgga gcggcggcct cgcggctctg
cacatggcgg cggggtacgt caggcccggc 600gtggcgaagg ttctcttgga
tctcggcgcg gatcccgagg tggcggacga ccgcgggaga 660acggcgttgg
atctggcgag ggagattctg aaggtgacgc cgaaggggaa tccgatgcag
720ttcggacgca ggattggact ggaaggtgtg attagggttt tggaaggggc
agtgttcgag 780tacgcggagg tgcaggagat tctggaacgg agaggaaagg
gtgagaattt ggagtatctt 840gtgcggtgga aggacggtgg tgccaacgag
tgggtgaagg cgaagtttgt ggcggaggat 900ttggtgaaag actacgaggc
tggcctcgag tacgccgtcg ctgaggcggt gctcgcgaaa 960agggtagcgg
atgaagggac gccggagttt ttggttaaat gggccgattt ggaggagccc
1020acatgggagc ccgaggagaa tgtggaccca gagcttgtca aagctttcga
gggaagtaac 1080aaccaggccc agcccagtag taatgggccc gctgtggtct
tttccaatca ggatagccct 1140agcctgtga 114942382PRTGlycine max 42Met
Glu Ala Ile Phe Ala Thr Lys Pro Ala Ser His Ser Leu Leu Leu1 5 10
15Thr Lys Leu Ser Pro Asn Pro Lys His Leu Phe Pro Pro His Gln Gln
20 25 30Ser Phe His Asn Ile Arg His Lys Pro Thr Arg Phe Arg Pro Val
Thr 35 40 45Ala Val Phe Gln Asn Gln His Gln Gln Asp Ala Ala Ala Ala
Ser Asn 50 55 60His Thr Glu Asp Glu Ser Tyr Gly Glu Val Lys Gly Ile
Ile Gly Ser65 70 75 80Arg Ala Leu Glu Ala Ala Thr Gly Met Glu Tyr
Leu Ile Glu Trp Asn 85 90 95Asp Gly His Ala Pro Ser Trp Val Pro Ala
Asp Phe Ile Ala Lys Asp 100 105 110Val Val Asp Glu Tyr Glu Thr Pro
Trp Trp Thr Ala Ala Lys Lys Ala 115 120 125Asp Glu Ser Ala Leu Lys
Asn Leu Thr Lys Ser Asp Asp Gly Arg Asp 130 135 140Val Asp Ala Val
Asp Ala Asp Gly Arg Thr Ala Leu Leu Phe Val Ala145 150 155 160Gly
Leu Gly Ser Glu Ser Cys Val Lys Leu Leu Ala Glu Ala Gly Ala 165 170
175Asn Leu Asp His Arg Asp Arg Ser Gly Gly Leu Ala Ala Leu His Met
180 185 190Ala Ala Gly Tyr Val Arg Pro Gly Val Ala Lys Val Leu Leu
Asp Leu 195 200 205Gly Ala Asp Pro Glu Val Ala Asp Asp Arg Gly Arg
Thr Ala Leu Asp 210 215 220Leu Ala Arg Glu Ile Leu Lys Val Thr Pro
Lys Gly Asn Pro Met Gln225 230 235 240Phe Gly Arg Arg Ile Gly Leu
Glu Gly Val Ile Arg Val Leu Glu Gly 245 250 255Ala Val Phe Glu Tyr
Ala Glu Val Gln Glu Ile Leu Glu Arg Arg Gly 260 265 270Lys Gly Glu
Asn Leu Glu Tyr Leu Val Arg Trp Lys Asp Gly Gly Ala 275 280 285Asn
Glu Trp Val Lys Ala Lys Phe Val Ala Glu Asp Leu Val Lys Asp 290 295
300Tyr Glu Ala Gly Leu Glu Tyr Ala Val Ala Glu Ala Val Leu Ala
Lys305 310 315 320Arg Val Ala Asp Glu Gly Thr Pro Glu Phe Leu Val
Lys Trp Ala Asp 325 330 335Leu Glu Glu Pro Thr Trp Glu Pro Glu Glu
Asn Val Asp Pro Glu Leu 340 345 350Val Lys Ala Phe Glu Gly Ser Asn
Asn Gln Ala Gln Pro Ser Ser Asn 355 360 365Gly Pro Ala Val Val Phe
Ser Asn Gln Asp Ser Pro Ser Leu 370 375 380435214DNAGlycine max
43tctaactaag taagaaattg aagtatttta taaaaaaaaa ttgactcatc gaatttataa
60agtaatttat gtaacaaaaa ttaagagtaa tttaaaagca atacatagat gaataggagt
120gtttttatta ggaagaataa gagtgttaat aatatcacac gtgaacctct
taaatcaagt 180aagctaaatc tctttggtaa aattaagatt cacctacatt
gttttttttt ttttatcaat 240acctacatca atataaaaac aataattttt
tatcaatacc tacattaata taaaaaaaat 300aaatgataca gttacattaa
atattcagag taaaagcttt gacactcatt tatttttatc 360taactcaaat
ataattttct ctattaaata atatttatgg tctaaatttt ttttagctat
420attgcagctc gagtcttgat atcatgacat cttagccaac taaagctcaa
ctggctcaaa 480ggcaggaata tttcaaaaaa aattatttca tatatattaa
tatattgatt aagaaaataa 540aaaaattata ctaaaagaac catgacttta
cccagcacca aggtgttcaa ttttcaaaag 600ccacaccact ggaagaatga
gaagccaaca tagtctttca tggactaaca catccacaaa 660cgtcacggtt
gtctttcatg caccacacga aattgccttt ttctttattc ttcaagaaat
720tatggcatca attagtctta atagtcaaat tcgtaaaaac taacaaattg
acttttaaaa 780aattaaaaaa ttcaaattaa tcacgggata tgtataaatt
atgatagact aatcattgta 840aataagttaa tgataaattc tcaaaaagat
aatttatttt taaattaatt attaaacatt 900ataactaaat aataaaatga
tctcacaaat taaatgacat gactaatttg ttatattttt 960tcatatttga
aaactacaat tttatgtcat ttttatggat taatttatat acttttaaga
1020ataaatttaa ctatttactc cataaattat cgattattga tgtttactaa
ttttgaagtt 1080tttgctaaca agtgtcgtcc taaagtaatt taattttcac
tcattaacaa aaaaataatt 1140taagaattat aaaattagtt taataattga
aacgagaaga aaggaagaac agattgtgag 1200ttacagtctc tttgtcccct
aacaattaat atttaatgat aaaaaatagt aatttgactg 1260ttaaaaatta
taagaaagaa aataatatat tttataaatt ctttagtatc cgaattgtat
1320atatatttat tcaatactga ttgatacaat tcatattaat ataatcgagt
ttttactgaa 1380catacattta ttaaaaaaaa tctctctagt gtccatttat
tcggcgagaa gccttctcgt 1440gctttacaca ctttaatatt attatatccc
cacccccacc aaaaaaaaaa aaactgttat 1500atctttccag tacatttatt
tcttttattt ttacaaagga aacttcacga aagtaattac 1560aaaaaagata
gtgaacatca tttttttagt taagatgaat tttaaaatca cactttttta
1620tatttttttg ttaccctttt cattattggg tgaaatctca tagtgaaact
attaaatagt 1680ttgggctcaa gttttattag taaagtctgc atgaaattta
acttaataat agagagagtt 1740ttggaaaggt aacgaatgtt agaaagtgtg
atattattat agttttattt agattaataa 1800ttatgtttac atgaaaattg
acaatttatt tttaaaattc agagtaatac ttaaattact 1860tatttacttt
aagattttga aaagatcatt tggctcttca tcatgccgat tgacaccctc
1920cacaagccaa gagaaactta agttgtaatt tttctaactc caagccttct
atataaacac 1980gtattggatg tgaagttgtt gcataacttg cattgaacaa
tagaaataac aacaaagaaa 2040ataagtgaaa aaagaaatat gggtgctttc
actgagaagc aagaggcttt ggtgagtagc 2100tcattcgaag cattcaaggc
aaacattcct caatacagcg ttgtgttcta cacttcgtaa 2160gttttctctt
aaagcatgta tctttcattc tctgtttttc ctttcgacat tttttgtgtt
2220tgaaaagaga tagtgtcaat gtgagtgggt attttttttt attaaaaatt
aacaggatac 2280tggagaaagc acccgcagca aaggacttgt tctcgtttct
atctaatgga gtagatccta 2340gtaatcctaa gctcacgggc catgctgaaa
agctttttgg attggtaagt atcatccaac 2400taaaattata gctattttat
gtgattaatt ttaagattaa acatgtattt aacactctta 2460aacatgtatt
taacactctt aagattaaac atgtatttaa ctaaaacatg tatttgctga
2520ttattttttt tttataatta tcttgtcaca tattatatat tttttgaatt
gtaggtgcgt 2580gactcagctg gtcaacttaa agcaaatgga acagtagtgg
ctgatgccgc acttggttct 2640atccatgccc aaaaagcaat cactgatcct
cagttcgtgg tatgataaat aataaaatgt 2700tacaataaat gcacatatac
ttaaatttta catggtgcag tgttatgatc atcatttttg 2760tttagtaatg
aatttactta aaatcttaaa ttatgtactt tttgaaagtt ttatatggaa
2820ttttaattat agggaaaaat gtaagagcta atccattagt gatgttttgt
ctgtaggtgg 2880ttaaagaagc actgctgaaa acaataaagg aggcagttgg
ggacaaatgg agtgatgaat 2940tgagcagtgc ttgggaagta gcctatgatg
aattggcagc agctattaag aaggcatttt 3000aggatctact attgccgtca
agtgtaataa ataaattttg tttcactaaa acttgttatt 3060aaacaagtcc
ccgatatata aatgttggtt aaaataagta aattatacgg tattgataaa
3120caatcttaag ttttatatat agttccatat actaaagttt gtgaatcata
atcgatgctt 3180tatataaata agttcataag aatttatatg ttttaaaatt
attttacccc tataaattca 3240aataaactta actaaatggg atagatgttt
tcctacgaaa aaaatgaaaa taagtaacag 3300taaaatgcgt tagaaatagt
tattgtggta gataaattca aatacattga aaatattttt 3360atcaattaga
ttaatatgcc aatgtttctg attttattac tttaaagaat agcaggggta
3420agagggggat tagcactcaa aaactgacat gaggccagaa tgttaccatg
gtagcgcaat 3480cgatcaacta ttaacatata cagtaaatct aaaagattaa
gggttcgttt cgtagaatta 3540aaaaaaataa ttcaagtttt gaattaaaat
atgatttaaa atgttaattc tatatttttt 3600ttctttttca ctttctttct
ctacaaatga gaaattagaa gctcaattcc attgcacggt 3660atactcaaac
cataattaca atttatcacg cgtaaatagc agaacgtacg ggttgctgag
3720agactaacaa agatattcaa ggtccaattt gttatacatt tccgccactg
atgaaagaac 3780attactatgc aaattcctta tagaagaaaa catttaccct
tctcaattct catttgcagc 3840aaaatcacac tctttaagct aattgattga
ttaaactggt gtagaattat aaataaatca 3900ataaaatgag atcattttga
acaaagtgaa gttgatttct cacttcacaa agttcgccaa 3960tactgatggt
aacttttagg tgtgaaataa gatgggctga ataaagtttt gaataaattg
4020gtttcatgag cctatttgaa aaatggtttg ttttgtgtga gcgttttatt
taggagaatg 4080acgaaacaca tatttcgcca agagaaataa tgaaataggt
gcatatttcg tcaataggaa 4140caacgaaata ggtgtatgtt aggtcaggtg
cagtgtgcag agtggatttt gtcatcctta 4200ttggcgaaat aggtggtgct
gttgctttgt caatttatct aatagtgagt tgaaaataat 4260ttatctaata
atgagtcaga aataatttat ctaataatga gtcaaaaccg ccaaaaaatc
4320cacaaagcag caccacttat ttcgccaatg aggatggcaa aatccactct
gcatgctgca 4380cttatttcgc caatgtggat ggcgaaatcc actctgcaca
ctgcacccct acgcgctgca 4440tactgctgca cctgacctga cctacaccta
tttcgctatt ccggttggcg aaatatgcac 4500ctatttcact gttcctcttg
gcgaaatatg tgtttcgcca ttcctttcag caaaacgctc 4560acacaaaaca
gaccacatta gcaaaatgtt tcaaaacaga ttgggaatta gtttctaaaa
4620gaaacccccc gaagtcaatt tgctataaaa tttaaactta tttaaataat
tgagtgcaac 4680tgtaaactta aatgtatttt tttttgttta aacatataag
tacacaaata atgaatccaa 4740atataataaa ttcactaata actcaactcg
gtcatctttg aaatcatgaa tatatttatt 4800atatctatac ataatccaac
aattatataa tgttttattt aacaaacagt gattaatgta 4860ttgaaattaa
aattaaatca ttcaggtaat accaaaattc aatcattagc agaaataatt
4920tttgatcaga ttttatttac ttgttcgcta aattttaaat tagttagagt
ttcttttctt 4980taatgaaccg gagggtcaag ataaaaaaat acattttttt
tttaataatt atcatattat 5040aacaggtagt tatttcaaag caattcaatg
cttcatttca accaccaaca tttaaaagaa 5100ataaaataca aaatagtact
cgtggttttg tgtctggaaa gctccaacca taatttaatc 5160caaaagttgt
ctattgtaac tttagcttaa ataaaccatg gtttaagact ttca 521444100DNAZea
mays 44ttacttctac ccgatagagc ttagaagttg gaagtaataa tttcttggtt
gattgtaggc 60ttaaccattt cttttttttt gacacgagga actcatcatg
100452361DNAZea mays 45atggaggcca ccggcagggg gctgttcccg aacaagccca
ccctcccggc ggggccgagg 60aaacggggcc cgctcctccc ggccgcgccc ccgccaccgt
ccccctcctc gctcccgctc 120gactcgctcc tgctccacct caccgcgccc
gcccccgcgc cggcccccgc gccgcggcgg 180tcgcaccaga cgccgacgcc
gccgcactcc ttcctctccc ccgacgcgca ggtgctggtg 240ctcgccatct
cctcgcaccc gctccccacg ctggcggcct tcctggcctc ccgccgcgac
300gagctcctcc gcgcggacat cacgtccctg ctcaaggcgc tggagctctc
ggggcactgg 360gagtgggcgc tcgcgctcct ccggtgggca ggcaaggagg
gtgccgccga cgcgtcggcg 420ctcgagatgg tcgtccgcgc gctgggccgc
gagggccagc acgacgccgt ctgcgcgctg 480ctcgacgaaa cgccgctccc
gccgggctcc cgcctcgacg tccgcgccta caccaccgtg 540ctgcacgcgc
tctcccgcgc gggccggtac gagcgcgcgc tcgagctctt cgccgagctc
600cggcgccagg gggtggcgcc cacgctcgtc acctacaacg tcgtgctgga
cgtgtacggg 660cggatgggcc ggtcgtggcc gcggatcgtc gccctcctcg
atgagatgcg cgccgccggg 720gtcgagcccg acggcttcac cgccagcacg
gtgatcgccg cgtgctgccg cgacgggctg 780gttgacgagg cggtggcgtt
cttcgaggac ctcaaggccc gcggccacgc cccgtgcgtc 840gtcacgtaca
cagcgttgct ccaggtgttc ggcaaggccg ggaactacac ggaggcgctg
900cgcgtgctcg gggagatgga gcagaacggc tgccagccag atgctgtgac
gtacaccgag 960ctcgccggaa cgtacgcccg ggctgggttc ttcgaggagg
ctgccaggtg cctggacaca 1020atggcatcca agggtctgtt gccaaacgca
ttcacgtaca acaccgtgat gacagcctat 1080gggaatgttg ggaaggtgga
tgaggcgctc gctctgtttg accagatgaa gaagaccggg 1140ttcgtgccga
acgtgaacac gtacaatctt gtccttggca tgcttggcaa gaagtcaagg
1200ttcacggtga tgctagagat gcttggagag atgtcgagga gcggatgcac
accgaaccgg 1260gtaacatgga acacaatgct tgcagtctgt gggaagcgtg
gcatggagga ctacgtcacc 1320cgggttctgg aggggatgag gtcttgcggg
gttgaactga gccgagacac ctacaacacc 1380ctgatagctg cgtacggccg
gtgtggctcg aggactaatg ccttcaagat gtacaacgag 1440atgaccagcg
ctggattcac cccctgcatc accacgtaca acgcgttgct gaacgtgctg
1500tcgcggcagg gcgactggtc caccgcccag tcgatcgtaa gcaaaatgag
gaccaagggg 1560ttcaagccga acgagcagtc gtattcgctg ctgctccagt
gctacgcgaa ggggggcaac 1620gtggcaggga tagccgcgat cgagaacgag
gtgtacggat caggtgccgt tttcccaagc 1680tgggtgatcc tgaggaccct
tgtcatcgcc aatttcaagt gccggcgact ggatggcatg 1740gagacggcgt
ttcaagaggt gaaggccaga ggctacaacc cggacctcgt gatattcaac
1800tccatgctgt ccatctacgc gaagaacggg atgtacagca aggccaccga
ggtcttcgac 1860tccatcaagc ggagcgggct gagccccgac ctcatcacct
acaacagcct gatggacatg 1920tacgccaagt gcagcgagtc gtgggaggcc
gagaagatac tgaaccagct caagtgctcc 1980cagacgatga agcccgacgt
ggtgtcctac aacacggtca taaacgggtt ctgcaagcag 2040gggctggtga
aagaggccca gagggtcctc tcggagatgg tcgccgacgg catggccccc
2100tgcgccgtga cctaccacac gctcgtcggg ggttactcca gcctggagat
gttcagcgag 2160gccagggagg tcatcggcta catggtccag cacggcctca
agcctatgga gctgacctac 2220aggagagtcg tcgagagcta ctgcagagcg
aagcggttcg aggaggctcg cggcttcctg 2280tccgaggtct cggagaccga
cctggatttt gacaagaagg cgctcgaagc ctatatagag 2340gatgcgcagt
ttggaaggta g 236146786PRTZea mays 46Met Glu Ala Thr Gly Arg Gly Leu
Phe Pro Asn Lys Pro Thr Leu Pro1 5 10 15Ala Gly Pro Arg Lys Arg Gly
Pro Leu Leu Pro Ala Ala Pro Pro Pro 20 25 30Pro Ser Pro Ser Ser Leu
Pro Leu Asp Ser Leu Leu Leu His Leu Thr 35 40 45Ala Pro Ala Pro Ala
Pro Ala Pro Ala Pro Arg Arg Ser His Gln Thr 50 55 60Pro Thr Pro Pro
His Ser Phe Leu Ser Pro Asp Ala Gln Val Leu Val65 70 75 80Leu Ala
Ile Ser Ser His Pro Leu Pro Thr Leu Ala Ala Phe Leu Ala 85 90 95Ser
Arg Arg Asp Glu Leu Leu Arg Ala Asp Ile Thr Ser Leu Leu Lys 100 105
110Ala Leu Glu Leu Ser Gly His Trp Glu Trp Ala Leu Ala Leu Leu Arg
115 120 125Trp Ala Gly Lys Glu Gly Ala Ala Asp Ala Ser Ala Leu Glu
Met Val 130 135 140Val Arg Ala Leu Gly Arg Glu Gly Gln His Asp Ala
Val Cys Ala Leu145 150 155 160Leu Asp Glu Thr Pro Leu Pro Pro Gly
Ser Arg Leu Asp Val Arg Ala 165 170 175Tyr Thr Thr Val Leu His Ala
Leu Ser Arg Ala Gly Arg Tyr Glu Arg 180 185 190Ala Leu Glu Leu Phe
Ala Glu Leu Arg Arg Gln Gly Val Ala Pro Thr 195 200 205Leu Val Thr
Tyr Asn Val Val Leu Asp Val Tyr Gly Arg Met Gly Arg 210 215 220Ser
Trp Pro Arg Ile Val Ala Leu Leu Asp Glu Met Arg Ala Ala Gly225 230
235 240Val Glu Pro Asp Gly Phe Thr Ala Ser Thr Val Ile Ala Ala Cys
Cys 245 250 255Arg Asp Gly Leu Val Asp Glu Ala Val Ala Phe Phe Glu
Asp Leu Lys 260 265 270Ala Arg Gly His Ala Pro Cys Val Val Thr Tyr
Thr Ala Leu Leu Gln 275 280 285Val Phe Gly Lys Ala Gly Asn Tyr Thr
Glu Ala Leu Arg Val Leu Gly 290 295 300Glu Met Glu Gln Asn Gly Cys
Gln Pro Asp Ala Val Thr Tyr Thr Glu305 310 315 320Leu Ala Gly Thr
Tyr Ala Arg Ala Gly Phe Phe Glu Glu Ala Ala Arg 325 330 335Cys Leu
Asp Thr Met Ala Ser Lys Gly Leu Leu Pro Asn Ala Phe Thr 340 345
350Tyr Asn Thr Val Met Thr Ala Tyr Gly Asn Val Gly Lys Val Asp Glu
355 360 365Ala Leu Ala Leu Phe Asp Gln Met Lys Lys Thr Gly Phe Val
Pro Asn 370 375 380Val Asn Thr Tyr Asn Leu Val Leu Gly Met Leu Gly
Lys Lys Ser Arg385 390 395 400Phe Thr Val Met Leu Glu Met Leu Gly
Glu Met Ser Arg Ser Gly Cys 405 410 415Thr Pro Asn Arg Val Thr Trp
Asn Thr Met Leu Ala Val Cys Gly Lys 420 425 430Arg Gly Met Glu Asp
Tyr Val Thr Arg Val Leu Glu Gly Met Arg Ser 435 440 445Cys Gly Val
Glu Leu Ser Arg Asp Thr Tyr Asn Thr Leu Ile Ala Ala 450 455 460Tyr
Gly Arg Cys Gly Ser Arg Thr Asn Ala Phe Lys Met Tyr Asn Glu465 470
475
480Met Thr Ser Ala Gly Phe Thr Pro Cys Ile Thr Thr Tyr Asn Ala Leu
485 490 495Leu Asn Val Leu Ser Arg Gln Gly Asp Trp Ser Thr Ala Gln
Ser Ile 500 505 510Val Ser Lys Met Arg Thr Lys Gly Phe Lys Pro Asn
Glu Gln Ser Tyr 515 520 525Ser Leu Leu Leu Gln Cys Tyr Ala Lys Gly
Gly Asn Val Ala Gly Ile 530 535 540Ala Ala Ile Glu Asn Glu Val Tyr
Gly Ser Gly Ala Val Phe Pro Ser545 550 555 560Trp Val Ile Leu Arg
Thr Leu Val Ile Ala Asn Phe Lys Cys Arg Arg 565 570 575Leu Asp Gly
Met Glu Thr Ala Phe Gln Glu Val Lys Ala Arg Gly Tyr 580 585 590Asn
Pro Asp Leu Val Ile Phe Asn Ser Met Leu Ser Ile Tyr Ala Lys 595 600
605Asn Gly Met Tyr Ser Lys Ala Thr Glu Val Phe Asp Ser Ile Lys Arg
610 615 620Ser Gly Leu Ser Pro Asp Leu Ile Thr Tyr Asn Ser Leu Met
Asp Met625 630 635 640Tyr Ala Lys Cys Ser Glu Ser Trp Glu Ala Glu
Lys Ile Leu Asn Gln 645 650 655Leu Lys Cys Ser Gln Thr Met Lys Pro
Asp Val Val Ser Tyr Asn Thr 660 665 670Val Ile Asn Gly Phe Cys Lys
Gln Gly Leu Val Lys Glu Ala Gln Arg 675 680 685Val Leu Ser Glu Met
Val Ala Asp Gly Met Ala Pro Cys Ala Val Thr 690 695 700Tyr His Thr
Leu Val Gly Gly Tyr Ser Ser Leu Glu Met Phe Ser Glu705 710 715
720Ala Arg Glu Val Ile Gly Tyr Met Val Gln His Gly Leu Lys Pro Met
725 730 735Glu Leu Thr Tyr Arg Arg Val Val Glu Ser Tyr Cys Arg Ala
Lys Arg 740 745 750Phe Glu Glu Ala Arg Gly Phe Leu Ser Glu Val Ser
Glu Thr Asp Leu 755 760 765Asp Phe Asp Lys Lys Ala Leu Glu Ala Tyr
Ile Glu Asp Ala Gln Phe 770 775 780Gly Arg785472490DNAGlycine max
47atggagggta cccttttccc caataggcca gttttgcctg ctccttcaca caaaccaaca
60caacaacctt tgaaattcaa gccaactttt ttgcctccac agtctccacc accaccacct
120ccttcttttc agttagattc ccttctccaa caccttcagc atctctcttc
agttcccatc 180accactcaca ctctcacact tgtgcctcct tctcatgata
acaccaaaga ttttaataat 240tcagttcatt caaagcaccc cactttaggt
tctggctcca taattgatga ggacaagttt 300gatgatgcaa agtttggatt
tttatcagac aagggtaagt tgctgttcag ttcaattgtt 360gggtcacctt
tgcatgaatt gaatgacttt ttcaactctg ttaagtttga gttgcttgag
420gctgattttc ccagcttgtt gaaggctttg gacctttctg ggaactggga
aagggcactc 480ttgctgtttg aatggggttg gttgcatttt gggagtgatc
agaatttgag gttggacaac 540caggttgttg aattgatggt taggatattg
gggagggagt cacagcattc aattgcatcc 600aagttgtttg atttaattcc
tgtggaaaaa tactcgcttg atgtccgggc ttacaccacc 660attcttcatt
cctatgctcg cactggcaag tacaaacggg ctattgactt gtttgggaag
720atgaaggaga ttggtcttga tccaactttg gtcacttaca atgttatgct
tgatgtttac 780ggcaagatgg gtcgttcttg ggatagaatc ttggagttgt
tggatgagat gaggagtaaa 840gggcttgagt tagatgagtt tacctgcagc
actgtgattt ctgcttgtgg gagagagggt 900atgctggatg aagcgaggaa
gtttttggct gaattgaaat ttaatggcta taaaccggga 960actgttacgt
ataattctat gttgcaggtt tttggaaagg caggaattta cactgaggcc
1020ttgagcatat tgaaagaaat ggaggataat aattgccctc ctgattctgt
tacttacaat 1080gagcttgcgg caacatatgt aagagctggt tttctggacg
aagggatggc tgtcatagat 1140acaatgacaa gcaaaggggt aatgccaaat
gctattacct ataccactgt aatagatgcc 1200tatggtaagg cgggaaggga
ggatgatgca ttaaggttgt tcagcctgat gaaggacttg 1260ggttgtgctc
ccaatgtgta cacatataac tctgttcttg ccatgctagg caagaaatca
1320agaacagaag atgttattaa ggttctctgt gagatgaaat tgaatggatg
tgctcctaat 1380cgtgctacat ggaacaccat gcttgctgta tgtagcgagg
agggtaagca caattatgtt 1440aacaaggtct tgagggaaat gaaaaactgt
ggatttgagc ctgataaaga cacattcaat 1500acattgatta gtgcatatgc
tcgttgcgga tctgaagttg attctgcaaa aatgtatggg 1560gaaatggtta
aatcaggctt tactccatgt gtaacaactt ataatgctct tctaaatgcc
1620ctggctcggc gaggtgattg gaaagcggca gaatctgtca ttcaggacat
gcgaaccaag 1680ggctttaagc ctaatgaaaa ttcatactca ctgttgctcc
attgttattc caaggctggg 1740aatgtcaagg ggatagagaa ggtcgagaaa
gaaatttatg atggtcatgt ctttcctagc 1800tggatacttt tgagaaccct
tgtccttaca aaccacaagt gcagacacct taggggaatg 1860gaaagggcat
ttgatcaact gcaaaagtat ggatacaaac ctgatttggt tgtcatcaac
1920tccatgcttt cgatgtttgc ccgaaacaag atgttttcga aggcccgtga
aatgctgcat 1980ttcattcatg aatgtggatt gcagccaaat cttttcacct
acaattgctt gatggatttg 2040tatgtccgag agggcgagtg ttggaaagca
gaagaagtgc tcaagggaat tcaaaactct 2100ggtccagagc cagatgttgt
gtcttacaat actgttatca agggattttg cagaaaaggg 2160ctcatgcagg
aggctattgg agttctctca gaaatgacca ctaaggggat tcaaccaact
2220atagttacat acaatacttt cttgtcaggc tatgcaggga tggagttgtt
tgatgaagca 2280aatgaagtca ttagatttat gattgagcac aattgcaggc
caagtgaact aacttacaag 2340attctagttg atggttactg taaagctggg
aagtatgaag aagccatgga ctttgtgtct 2400aagattaagg agcttgatat
ctcctttgat gatcaatctg tgaaaagact tggttcttgt 2460attagggaga
gagtggggtc tactttgtga 249048829PRTGlycine max 48Met Glu Gly Thr Leu
Phe Pro Asn Arg Pro Val Leu Pro Ala Pro Ser1 5 10 15His Lys Pro Thr
Gln Gln Pro Leu Lys Phe Lys Pro Thr Phe Leu Pro 20 25 30Pro Gln Ser
Pro Pro Pro Pro Pro Pro Ser Phe Gln Leu Asp Ser Leu 35 40 45Leu Gln
His Leu Gln His Leu Ser Ser Val Pro Ile Thr Thr His Thr 50 55 60Leu
Thr Leu Val Pro Pro Ser His Asp Asn Thr Lys Asp Phe Asn Asn65 70 75
80Ser Val His Ser Lys His Pro Thr Leu Gly Ser Gly Ser Ile Ile Asp
85 90 95Glu Asp Lys Phe Asp Asp Ala Lys Phe Gly Phe Leu Ser Asp Lys
Gly 100 105 110Lys Leu Leu Phe Ser Ser Ile Val Gly Ser Pro Leu His
Glu Leu Asn 115 120 125Asp Phe Phe Asn Ser Val Lys Phe Glu Leu Leu
Glu Ala Asp Phe Pro 130 135 140Ser Leu Leu Lys Ala Leu Asp Leu Ser
Gly Asn Trp Glu Arg Ala Leu145 150 155 160Leu Leu Phe Glu Trp Gly
Trp Leu His Phe Gly Ser Asp Gln Asn Leu 165 170 175Arg Leu Asp Asn
Gln Val Val Glu Leu Met Val Arg Ile Leu Gly Arg 180 185 190Glu Ser
Gln His Ser Ile Ala Ser Lys Leu Phe Asp Leu Ile Pro Val 195 200
205Glu Lys Tyr Ser Leu Asp Val Arg Ala Tyr Thr Thr Ile Leu His Ser
210 215 220Tyr Ala Arg Thr Gly Lys Tyr Lys Arg Ala Ile Asp Leu Phe
Gly Lys225 230 235 240Met Lys Glu Ile Gly Leu Asp Pro Thr Leu Val
Thr Tyr Asn Val Met 245 250 255Leu Asp Val Tyr Gly Lys Met Gly Arg
Ser Trp Asp Arg Ile Leu Glu 260 265 270Leu Leu Asp Glu Met Arg Ser
Lys Gly Leu Glu Leu Asp Glu Phe Thr 275 280 285Cys Ser Thr Val Ile
Ser Ala Cys Gly Arg Glu Gly Met Leu Asp Glu 290 295 300Ala Arg Lys
Phe Leu Ala Glu Leu Lys Phe Asn Gly Tyr Lys Pro Gly305 310 315
320Thr Val Thr Tyr Asn Ser Met Leu Gln Val Phe Gly Lys Ala Gly Ile
325 330 335Tyr Thr Glu Ala Leu Ser Ile Leu Lys Glu Met Glu Asp Asn
Asn Cys 340 345 350Pro Pro Asp Ser Val Thr Tyr Asn Glu Leu Ala Ala
Thr Tyr Val Arg 355 360 365Ala Gly Phe Leu Asp Glu Gly Met Ala Val
Ile Asp Thr Met Thr Ser 370 375 380Lys Gly Val Met Pro Asn Ala Ile
Thr Tyr Thr Thr Val Ile Asp Ala385 390 395 400Tyr Gly Lys Ala Gly
Arg Glu Asp Asp Ala Leu Arg Leu Phe Ser Leu 405 410 415Met Lys Asp
Leu Gly Cys Ala Pro Asn Val Tyr Thr Tyr Asn Ser Val 420 425 430Leu
Ala Met Leu Gly Lys Lys Ser Arg Thr Glu Asp Val Ile Lys Val 435 440
445Leu Cys Glu Met Lys Leu Asn Gly Cys Ala Pro Asn Arg Ala Thr Trp
450 455 460Asn Thr Met Leu Ala Val Cys Ser Glu Glu Gly Lys His Asn
Tyr Val465 470 475 480Asn Lys Val Leu Arg Glu Met Lys Asn Cys Gly
Phe Glu Pro Asp Lys 485 490 495Asp Thr Phe Asn Thr Leu Ile Ser Ala
Tyr Ala Arg Cys Gly Ser Glu 500 505 510Val Asp Ser Ala Lys Met Tyr
Gly Glu Met Val Lys Ser Gly Phe Thr 515 520 525Pro Cys Val Thr Thr
Tyr Asn Ala Leu Leu Asn Ala Leu Ala Arg Arg 530 535 540Gly Asp Trp
Lys Ala Ala Glu Ser Val Ile Gln Asp Met Arg Thr Lys545 550 555
560Gly Phe Lys Pro Asn Glu Asn Ser Tyr Ser Leu Leu Leu His Cys Tyr
565 570 575Ser Lys Ala Gly Asn Val Lys Gly Ile Glu Lys Val Glu Lys
Glu Ile 580 585 590Tyr Asp Gly His Val Phe Pro Ser Trp Ile Leu Leu
Arg Thr Leu Val 595 600 605Leu Thr Asn His Lys Cys Arg His Leu Arg
Gly Met Glu Arg Ala Phe 610 615 620Asp Gln Leu Gln Lys Tyr Gly Tyr
Lys Pro Asp Leu Val Val Ile Asn625 630 635 640Ser Met Leu Ser Met
Phe Ala Arg Asn Lys Met Phe Ser Lys Ala Arg 645 650 655Glu Met Leu
His Phe Ile His Glu Cys Gly Leu Gln Pro Asn Leu Phe 660 665 670Thr
Tyr Asn Cys Leu Met Asp Leu Tyr Val Arg Glu Gly Glu Cys Trp 675 680
685Lys Ala Glu Glu Val Leu Lys Gly Ile Gln Asn Ser Gly Pro Glu Pro
690 695 700Asp Val Val Ser Tyr Asn Thr Val Ile Lys Gly Phe Cys Arg
Lys Gly705 710 715 720Leu Met Gln Glu Ala Ile Gly Val Leu Ser Glu
Met Thr Thr Lys Gly 725 730 735Ile Gln Pro Thr Ile Val Thr Tyr Asn
Thr Phe Leu Ser Gly Tyr Ala 740 745 750Gly Met Glu Leu Phe Asp Glu
Ala Asn Glu Val Ile Arg Phe Met Ile 755 760 765Glu His Asn Cys Arg
Pro Ser Glu Leu Thr Tyr Lys Ile Leu Val Asp 770 775 780Gly Tyr Cys
Lys Ala Gly Lys Tyr Glu Glu Ala Met Asp Phe Val Ser785 790 795
800Lys Ile Lys Glu Leu Asp Ile Ser Phe Asp Asp Gln Ser Val Lys Arg
805 810 815Leu Gly Ser Cys Ile Arg Glu Arg Val Gly Ser Thr Leu 820
825
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

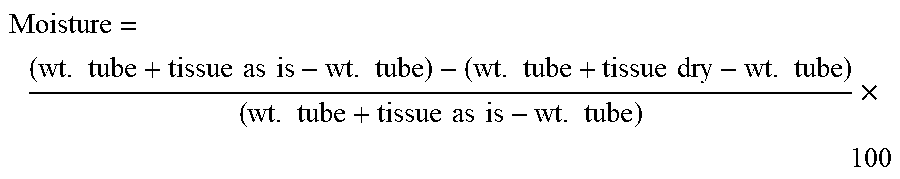
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.