Bi-specific Antibodies And Uses Thereof
ROFFLER; STEVEN R ; et al.
U.S. patent application number 16/111242 was filed with the patent office on 2021-05-27 for bi-specific antibodies and uses thereof. This patent application is currently assigned to ACADEMIA SINICA. The applicant listed for this patent is ACADEMIA SINICA, KAOHSIUNG MEDICAL UNIVERSITY. Invention is credited to BING-MAE CHEN, TIAN-LU CHENG, Kuo-Hsiang Chuang, CHIEN-HAN KAO, STEVEN R ROFFLER, YU-CHENG SU, HSIN-YI TUNG.
| Application Number | 20210154319 16/111242 |
| Document ID | / |
| Family ID | 1000005579910 |
| Filed Date | 2021-05-27 |









View All Diagrams
| United States Patent Application | 20210154319 |
| Kind Code | A9 |
| ROFFLER; STEVEN R ; et al. | May 27, 2021 |
BI-SPECIFIC ANTIBODIES AND USES THEREOF
Abstract
Disclosed herein is a bi-specific antibody that specifically directs a therapeutic agent to a cancer cell by targeting a tumor antigen of the cancer cell, and thereby suppressing the growth of the cancer or blocking the invasion or metastasis of the cancer. The bi-specific antibody of the present disclosure includes a first antigen binding site that binds to polyethylene glycol (PEG); and a second antigen binding site that binds to a target ligand, such as a tumor antigen.
| Inventors: | ROFFLER; STEVEN R; (Taipei City, TW) ; CHENG; TIAN-LU; (Kaohsiung City, TW) ; KAO; CHIEN-HAN; (Kaohsiung City, TW) ; CHEN; BING-MAE; (Taipei City, TW) ; SU; YU-CHENG; (Taipei City, TW) ; TUNG; HSIN-YI; (Taipei City, TW) ; Chuang; Kuo-Hsiang; (Taipei City, TW) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | ACADEMIA SINICA Taipei City TW KAOHSIUNG MEDICAL UNIVERSITY Kaohsiung City TW |
||||||||||
| Prior Publication: |
|
||||||||||
| Family ID: | 1000005579910 | ||||||||||
| Appl. No.: | 16/111242 | ||||||||||
| Filed: | August 24, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15123243 | Sep 1, 2016 | 10188742 | ||
| PCT/US2015/018365 | Mar 2, 2015 | |||
| 16111242 | ||||
| 61946980 | Mar 3, 2014 | |||
| 61946997 | Mar 3, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2317/35 20130101; C07K 16/2863 20130101; A61K 49/0002 20130101; C07K 16/44 20130101; C07K 16/32 20130101; C07K 2317/622 20130101; C07K 16/30 20130101; C07K 16/2803 20130101; C07K 2317/31 20130101; C07K 2317/24 20130101; C07K 2317/624 20130101; C07K 16/3061 20130101; A61K 47/6879 20170801; A61K 47/6897 20170801; C07K 16/2887 20130101; C07K 2317/73 20130101 |
| International Class: | A61K 47/68 20170101 A61K047/68; C07K 16/32 20060101 C07K016/32; A61K 49/00 20060101 A61K049/00; C07K 16/28 20060101 C07K016/28; C07K 16/30 20060101 C07K016/30; C07K 16/44 20060101 C07K016/44 |
Claims
1. A humanized bi-specific antibody against the backbone of polyethylene glycol (PEG) and a target ligand, comprising, a first antigen binding site that binds to the PEG, wherein the first antigen binding site comprises a first VL-C.kappa. domain and a first VH-CH1 domain; and a second antigen binding site that binds to the target ligand, which is EGFR, wherein, the first VL-C.kappa. domain comprises a CDR1 having the sequence at least 90% identical to SEQ ID NO: 216; a CDR2 having the sequence at least 90% identical to Trp-Ala-Ser; and a CDR3 having the sequence at least 90% identical to SEQ ID NO: 217; the first VH-CH1 domain comprises a CDR1 having the sequence at least 90% identical to SEQ ID NO: 218; a CDR2 having the sequence at least 90% identical to SEQ ID NO: 219; and a CDR3 having the sequence at least 90% identical to SEQ ID NO: 220; and the second antigen binding site comprises a single chain variable fragment (scFv) at least 90% identical to SEQ ID NO: 7.
2. The humanized bi-specific antibody of claim 1, wherein the first VL-C.kappa. domain has the sequence at least 90% identical to SEQ ID NO: 9, and the first VH-CH1 domain has the sequence at least 90% identical to SEQ ID NO: 10.
3. The humanized bi-specific antibody of claim 1, wherein the first antigen binding site further comprises a first HR-CH2-CH3 domain at least 90% identical to SEQ ID NO: 3 disposed between the first VH-CH1 domain and the scFv.
4. A pharmaceutical kit comprising the humanized bi-specific antibody of claim 1; and a PEGylated substance, wherein the substance is a protein, a peptide, or a nanoparticle, wherein the nanoparticle contains a chemotherapeutic drug or an imaging agent.
5. The pharmaceutical kit of claim 4, wherein the chemotherapeutic drug is adriamycin, amifostine, bleomycin, busulfan, cisplatin, carboplatin, oxaliplatin, camptothecin, CPT-11, cytosine arabinoside, chlorambucil, cyclophosphamide, cytarabine, daunorubicin, doxorubicin, docetaxel, dacarbazine, dactinomycin, etoposide, 5-fluorouracil (5-FU), fluoxuridine, gemcitabine, hydroxyurea, ifosfamide, idarubicin, interferon beta, irinotecan, L-asparaginase, L-aspartic acid, lomustine, mechlorethamine, mitomycin, methotrexate, mitoxantrone, megestrol, melphalan, mercaptopurine, mitotane, paclitaxel (taxol), plicamycin, pentostatin, streptozocin, topotecan, tamoxifen, teniposide, thioguanine, vinblastine, vincristine, SN38 or a combination thereof.
6. The pharmaceutical kit of claim 4, wherein the imaging agent is a quantum dot (QD), a microbubble contrast agent, a fluorescence dye, a chelated radioisotope a paramagnetic iron or a gold nanoparticle.
7. The pharmaceutical kit of claim 4, wherein the protein is a chemokine or a cytokine; and the peptide is leuprolide, goserelin, octreotide, histrelin, abarelix, cetrorelix, degarelix, cilengtide, ATN-161 or IM862.
8. A method for treating a subject suffering from a cancer comprising: mixing a first amount of the humanized bi-specific antibody of claim 1 with a second amount of a PEGylated substance to form an assembly; and administering a therapeutically effective amount of the assembly either sequentially or concurrently to the subject to inhibit the growth or metastasis of the cancer; wherein the PEGylated substance is therapeutic and is a protein, a peptide, or a nanoparticle, wherein the nanoparticle contains a chemotherapeutic drug.
9. The method of claim 8, wherein the chemotherapeutic drug is adriamycin, amifostine, bleomycin, busulfan, cisplatin, carboplatin, oxaliplatin, camptothecin, CPT-11, cytosine arabinoside, chlorambucil, cyclophosphamide, cytarabine, daunorubicin, doxorubicin, docetaxel, dacarbazine, dactinomycin, etoposide, 5-fluorouracil (5-FU), fluoxuridine, gemcitabine, hydroxyurea, ifosfamide, idarubicin, interferon beta, irinotecan, L-asparaginase, L-aspartic acid, lomustine, mechlorethamine, mitomycin, methotrexate, mitoxantrone, megestrol, melphalan, mercaptopurine, mitotane, paclitaxel (taxol), plicamycin, pentostatin, streptozocin, topotecan, tamoxifen, teniposide, thioguanine, vinblastine, vincristine, SN38 or any combination thereof.
10. The method of claim 8, wherein the protein is a chemokine or a cytokine; and the peptide is leuprolide, goserelin, octreotide, histrelin, abarelix, cetrorelix, degarelix, cilengtide, ATN-161 or IM862.
11. The method of claim 8, wherein the cancer is breast cancer, colorectal cancer, colon cancer, hepatic cancer, non-Hodgkin's lymphoma, lymphoma, pancreatic cancer, lung cancer, gastric cancer, prostate cancer, brain tumor, retinoblastoma, ovary cancer, cervical cancer, hematopoietic malignances, esophageal cancer, renal cell carcinoma, squamous cell carcinoma, glioma, or leukemia.
12. A method of imaging tissues in a subject comprising: (a) mixing a first sufficient amount of the humanized bi-specific antibody of claim 1 and a second sufficient amount of a PEGylated imaging agent to form an assembly; (b) injecting the assembly of the step (a) to the subject; and (c) imaging the tissues of the subject by fluorescence imaging, electron spin resonance (ESR) imaging, X-ray imaging, computed tomography (CT), or magnetic resonance imaging (MRI).
Description
[0001] This application contains a Sequence Listing in computer readable form. The computer readable form is incorporated herein by reference. This application is a Divisional Application of the pending U.S. patent application Ser. No. 15/123,243 filed on Sep. 1, 2016, which is the National Stage of International Application No. PCT/US2015/018365 filed on Mar. 2, 2015, which claims priority to U.S. Provisional Application No. 61/946,997, filed Mar. 3, 2014, and U.S. Provisional Appl. No. 61/946,980, filed Mar. 3, 2014. The entire contents of these documents are incorporated herein by reference in their entireties.
BACKGROUND OF THE INVENTION
1. Field of the Invention
[0002] The present disclosure relates to treatments of cancers. Specifically, the present disclosure relates to novel bi-specific antibodies and their uses for suppressing the growth or metastasis of cancers; and tracking the development of cancers.
2. Description of Related Art
[0003] Covalent attachment of poly(ethylene glycol) (PEGylation) to substances such as proteins, peptides, and nanoparticles (NPs) (e.g., liposomes, micelles, and the like) can increase drug bioavailability, enhance blood circulation half-life and hinder capture by the reticuloendothelial system (RES). These favorable attributes have led to the widespread use of PEGylation in the development of NPs including those available in clinical use, such as liposomal doxorubicin (Caelyx) for the treatment of ovarian and breast carcinomas and Kaposi's sarcoma and Genexol-PM.RTM. (Paclitaxel-loaded PEG-PLA micelles), approved for metastatic breast cancer, non-small cell lung cancer and ovarian cancer in South Korea. PEGylated nanoparticles (PEG-NPs) are highly regarded as the second generation of drug delivery systems and the mainstream of therapeutic or imaging agents.
[0004] PEGylated substances, particularly, PEG-NPs can accumulate in tumors due to the enhanced permeability and retention (EPR) effect caused by the abnormal structure of endothelial cells in tumors. PEG-NPs, however, often accumulate near tumors but do not penetrate into the tumor mass, and some drugs cannot easily diffuse from PEG-NPs to cancer cells. Therefore, several studies reported that chemical conjugation of antibodies to PEG-NPs increases specific targeting and intracellular uptake which improves therapeutic efficacy and the sensitivity of imaging. However, chemically linking antibodies to PEG-NPs is difficult to achieve. Most functional groups (e.g., amino, carboxyl, thiol groups) are abundant in ligands, which may cause loss of antibody function, or result in heterogeneous orientation of the antibody, thereby rendering it difficult to obtain a reproducible product after chemical conjugation. Further, chemical conjugation may also alter the structure of nano-carriers and encapsulated drugs. These problems limit the clinical applicability of targeted NPs.
[0005] In view of the foregoing, there exist in the related art, a need for an improved way of targeting PEGylated substances (e.g., PEG-NPs), which is reproducible and easy to use.
SUMMARY
[0006] The present disclosure provides humanized bi-specific antibodies and their uses for treating cancers or for tracking development of cancers.
[0007] Accordingly, it is the first object of the present disclosure to provide a bi-specific antibody (BsAb) that bind to two different epitopes, which are a PEG molecule (e.g., the terminal methoxy or hydroxyl group of the PEG, or the backbone of the PEG) and a target ligand (e.g., an epidermal growth factor receptor (EGFR), TAG72, CD19, or CD20). The BsAb of the present disclosure includes a first antigen binding site that binds to PEG and comprises a first light chain variable domain and a first heavy chain variable domain; a second antigen binding site that binds to a target ligand (e.g., a tumor antigen). Preferably, the BsAb of the present disclosure further includes a peptide linker between the first antigen binding site and the second antigen binding site. Optionally, the first antigen binding site may further include a first hinge domain.
[0008] The target ligand may be a protein selected from the group consisting of epidermal growth factor receptor (EGFR), human epidermal growth factor receptor (HER2), HER3, tumor-associated glycoprotein 72 (TAG-72), CD19 and, CD20.
[0009] In some embodiments, the first antigen binding site of the BsAb binds to the backbone of PEG and comprises a first VL-C.kappa. domain at least 90% identical to SEQ ID NO: 1, a first VH-CH1 domain at least 90% identical to SEQ ID NO: 2, and a first hinge domain at least 90% identical to SEQ ID NO: 3; while the second antigen binding site of the BsAb binds to any of TAG-72, EGFR, or HER2 and comprises a single chain variable fragment (scFv) at least 90% identical to SEQ ID NO: 5, 7, or 8; and the peptide linker is at least 90% identical to SEQ ID NO: 4.
[0010] In some embodiments, the first antigen binding site binds to the backbone of PEG with the first VL-C.kappa. domain at least 90% identical to SEQ ID NO: 9, and the first VH-CH1 domain at least 90% identical to SEQ ID NO: 10; while the second antigen binding site binds to EGFR or CD19 and comprises a scFv at least 90% identical to SEQ ID NO: 7 or 11; and the peptide linker is at least 90% identical to SEQ ID NO: 4.
[0011] In other embodiment, the first antigen binding site binds to the backbone of PEG and comprises a first VL-C.kappa. domain at least 90% identical to SEQ ID NO: 9, a first VH-CH1 domain at least 90% identical to SEQ ID NO: 10, and a first HR-CH2-CH3 domain at least 90% identical to SEQ ID NO: 22; while the second antigen binding site binds to CD19 or HER2 and comprises a second VL-CH1 domain at least 90% identical to SEQ ID NO: 23 or 26, a second VH-C.kappa. domain at least 90% identical to SEQ ID NO: 24 or 27, and a second HR-CH2-CH3 domain at least 90% identical to SEQ ID NO:25.
[0012] In another embodiment, the first antigen binding site binds to the terminal methoxy or hydroxyl group of PEG and comprises a first VL-C.kappa. domain at least 90% identical to SEQ ID NO: 12, a first VH-CH1 domain at least 90% identical to SEQ ID NO: 13, and a first HR-CH2-CH3 domain at least 90% identical to SEQ ID NO: 22; while the second antigen binding site binds to CD19 or HER2 and comprises a second VL-CH1 domain at least 90% identical to SEQ ID NO: 23 or 26, a second VH-C.kappa. domain at least 90% identical to SEQ ID NO: 24 or 27, and a second HR-CH2-CH3 domain at least 90% identical to SEQ ID NO: 25.
[0013] In still another embodiment, the first antigen binding site binds to the terminal methoxy or hydroxyl group of polyethylene glycol (PEG) and comprises a first VL-C.kappa. domain at least 90% identical to SEQ ID NO: 12, a first VH-CH1 domain at least 90% identical to SEQ ID NO: 13, and a first HR-CH2-CH3 domain at least 90% identical to SEQ ID NO: 22; while the second antigen binding site binds to HER2 or EGFR, and comprises a humanized single chain variable fragment (scFv) at least 90% identical to SEQ ID NO: 15 or 16.
[0014] In further embodiments, the first antigen binding site binds to the terminal methoxy or hydroxyl group of polyethylene glycol (PEG) and comprises a humanized single chain variable fragment (scFv) at least 90% identical to SEQ ID NO: 17; while the second antigen binding site binds to CD19 or CD20 and comprises a first VL-C.kappa. domain at least 90% identical to SEQ ID NO: 21, a first VH-CH1 domain at least 90% identical to SEQ ID NO: 20.
[0015] It is the second object of the present disclosure to provide a pharmaceutical kit for treating or tracking the development of cancers, including metastatic and/or drug-resistant cancers. The pharmaceutical kit includes at least, two components, which are respectively the bi-specific antibody described above; and a PEGylated substance that is either a therapeutic agent or an imaging agent. The therapeutic agent may be any of a protein, a peptide, or a nanoparticle containing therein a chemotherapeutic drug. The imaging agent may be a quantum dot (QD), a microbubble contrast agent, a fluorescence dye, an iron nanoparticle, a chelated radioisotope or a gold nanoparticle.
[0016] In practice, the bi-specific antibody and the PEGylated substance of the pharmaceutical kit are first mixed to form an assembly; and the assembly is then administered to the subject for treating cancers or for tracking cancers.
[0017] It is thus the third object of the present disclosure to provide a method of treating a subject suffering from the growth of a cancer. The method includes the steps of, administering the bi-specific antibody described above and a PEGylated substance containing a therapeutic agent, concurrently or sequentially to the subject in a dose sufficient to inhibit the growth or metastasis of the cancer of the subject. Preferably, the method comprises the steps of mixing the bi-specific antibody described above and the PEGylated substance containing a therapeutic agent to form an assembly, and administering the assembly to the subject in a dose sufficient to inhibit the growth or metastasis of the cancer of the subject. The dose administered to the subject is from about 0.1 to 50 mg/Kg body weight of the subject. In certain embodiments, the dose is administered to the subject from about 1 to 40 mg/Kg body weight of the subject, preferably from about 5 to 10 mg/Kg body weight of the subject. The dose can be administered in a single dose, or alternatively in more than one smaller doses.
[0018] Cancers, preferably those exhibit increased expression levels of EGFR, HER2, TAG72, CD19 or CD20 are treatable by the method of the present disclosure. In preferred embodiments, the method of the present disclosure is effective for treating a subject having breast cancer, head and neck cancer, colorectal cancer or ovarian cancer.
[0019] It is the fourth object of the present disclosure to provide a method of imaging tissues in a live subject. The method includes steps of, administering the bi-specific antibody described above and a PEGylated substance containing a therapeutic agent, concurrently or sequentially to the subject in an amount sufficient to imagine the tissues in the subject. Preferably, the method includes steps of: (a) mixing a first sufficient amount of any of the humanized bi-specific antibody of the present disclosure and a second sufficient amount of a PEGylated substance (e.g., a nanoparticle containing therein an imagine agent such as a quantum dot (PEG-QD) or a fluorescent dye) to form an assembly; (b) injecting the assembly of the step (a) to the subject; and (c) imaging the tissues of the subject by fluorescence imaging, electron spin resonance (ESR) imaging, gamma camera imaging, X-ray imaging, computed tomography (CT), or magnetic resonance imaging (MRI). According to some embodiments, the PEG-QD comprises a quantum dot nanocrystal selected from the group consisting of CdHgTe, CdSe, CdSe/ZnS, CdS, CdTe, CdTe/CdS, PbSe and PbS.
[0020] The details of one or more embodiments of the invention are set forth in the accompanying description below. Other features and advantages of the invention will be apparent from the detail descriptions, and from claims.
BRIEF DESCRIPTION OF THE DRAWINGS
[0021] The patent or application file contains at least one drawing executed in color. Copies of this patent or patent application publication with color drawing(s) will be provided by the Office upon request and payment of the necessary fee.
[0022] These and other features, aspects and advantages of the present invention will become better understood with reference to the following description, appended claims and the accompanying drawings, where:
[0023] FIG. 1 is a schematic diagram of IgG antibody with the domains indicated;
[0024] FIG. 2A is a schematic diagram of a dimeric BsAb structure in accordance to one embodiment of the present disclosure;
[0025] FIG. 2B is a schematic diagram of a monomeric BsAb structure in accordance to one embodiment of the present disclosure;
[0026] FIG. 2C is a schematic diagram of a monomeric BsAb structure in accordance to another embodiment of the present disclosure;
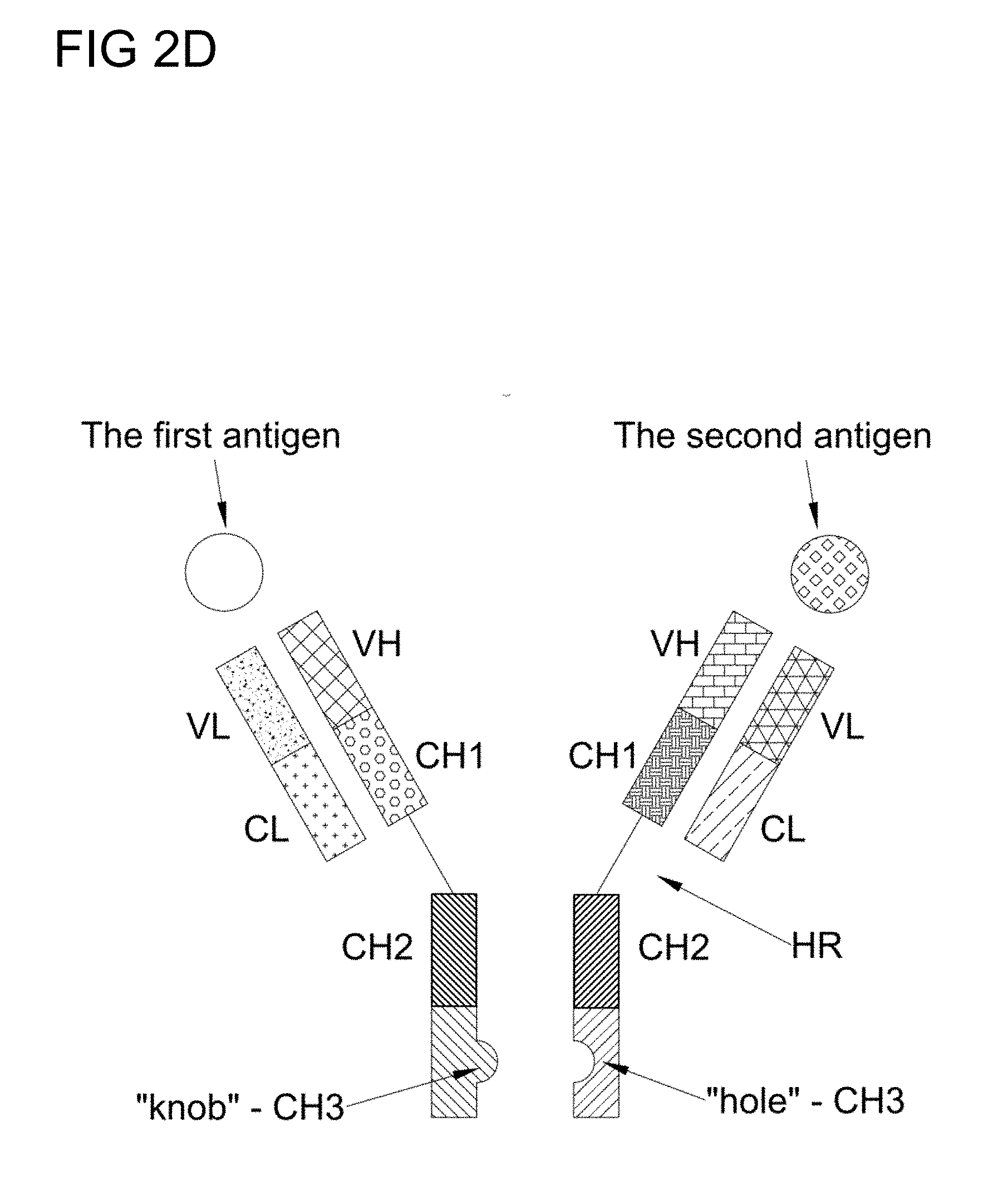
[0027] FIG. 2D is a schematic diagram of a "knob in hole" BsAb structure in accordance to one embodiment of the present disclosure;
[0028] FIG. 2E is a schematic diagram depicting the modified "knob in hole" BsAb structure having crossover heavy and light chains in accordance to one embodiment of the present disclosure;
[0029] FIG. 3 is a schematic diagram illustrating the one-step targeting and treating cancer by use of the humanized anti-mPEG BsAbs in accordance to one embodiment of the present disclosure;
[0030] FIG. 4 depicts the binding of anti-mPEG antibody secreted by hybridoma 15-2b to immobilized PEG molecules in accordance with one example of the present disclosure;
[0031] FIG. 5A is a schematic illustration of DNA constructs for humanized anti-PEG (hE11) BsAbs of example 1.2 in accordance with one embodiment of the present disclosure;
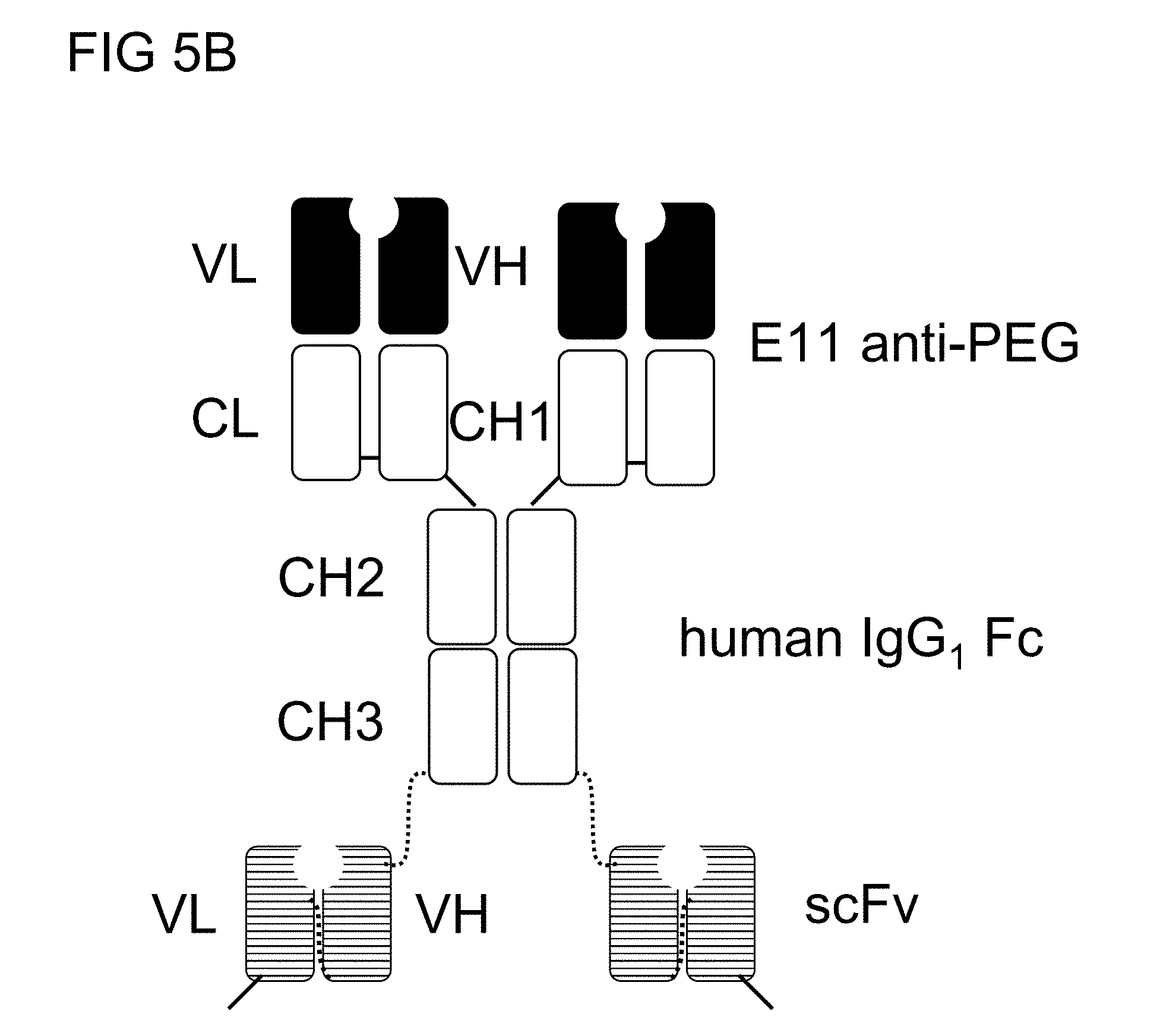
[0032] FIG. 5B is a schematic drawing of the structure of the humanized anti-PEG (hE11) BsAbs of example 1.2;
[0033] FIG. 5C illustrates the SDS-PAGE analysis of the humanized anti-PEG (hE11) BsAbs of example 1.2 in reducing or non-reducing conditions in accordance with one embodiment of the present disclosure;
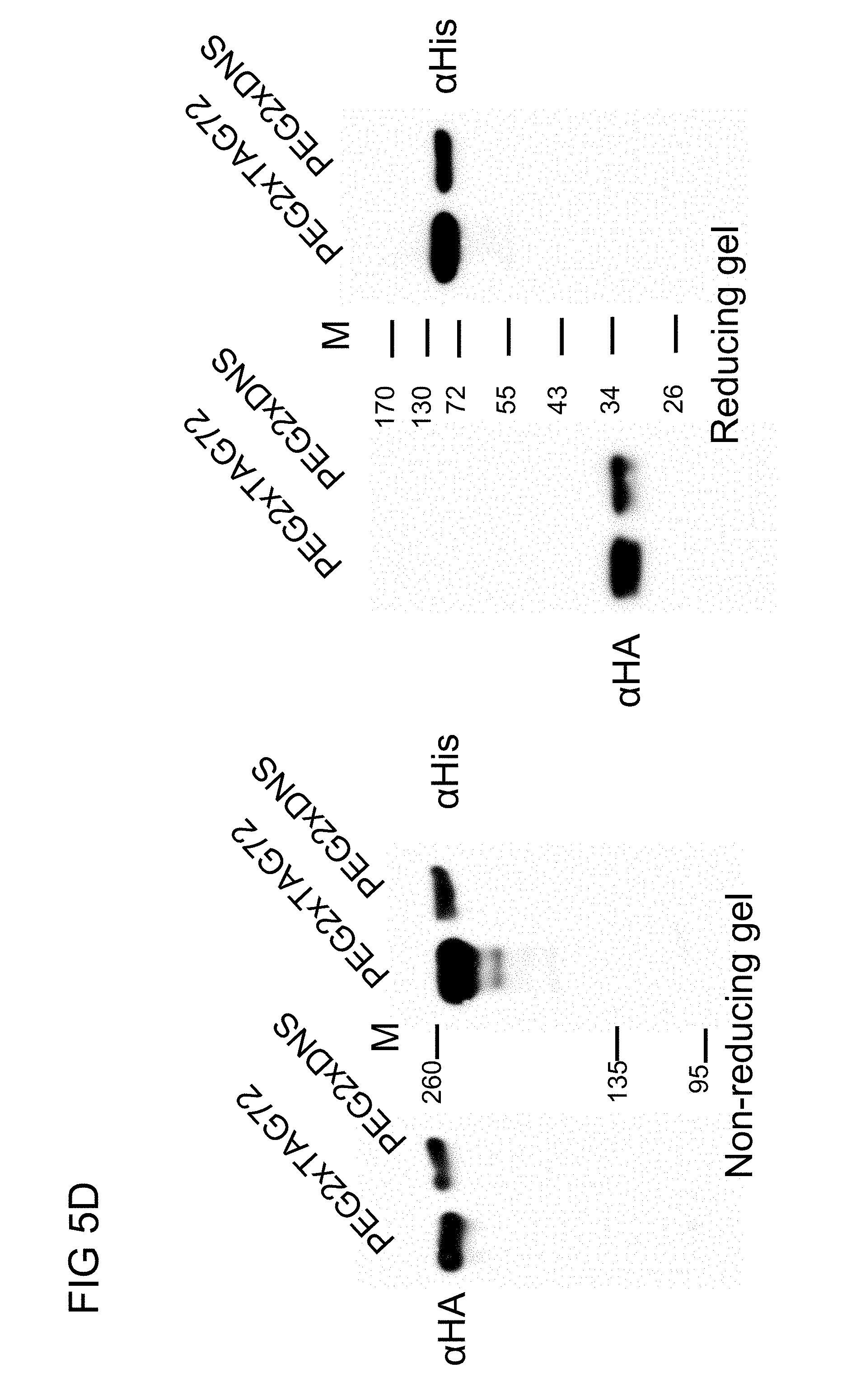
[0034] FIG. 5D illustrates the western blot analysis of the humanized anti-PEG (hE11) BsAbs of example 1.2 in accordance with one embodiment of the present disclosure;
[0035] FIGS. 6A to 6C respectively illustrate the antigen-binding activity of the humanized anti-PEG (hE11) BsAbs of example 1.2 towards (A) mucin, (B) BSA-PEG.sub.5,000 or (C) BSA in accordance with one embodiment of the present disclosure;
[0036] FIG. 7 illustrates the cancer cell selectivity of the humanized anti-PEG (hE11) BsAbs of example 1.2 in accordance with one embodiment of the present disclosure;
[0037] FIG. 8A illustrates the cancer cell selectivity of the dimeric humanized anti-PEG (hE11) BsAbs of example 1.4 in Jurkat (TAG-72+), MDA-MB-468 (EGFR+) or BT-474 (HER2+) cells in accordance with one embodiment of the present disclosure;
[0038] FIG. 8B illustrates the binding activities of the dimeric humanized anti-PEG (hE11) BsAbs of example 1.4 with the PEGylated liposomal Texas Red in Jurkat (TAG-72+), MDA-MB-468 (EGFR+) or BT-474 (HER2+) cells in accordance with one embodiment of the present disclosure;
[0039] FIG. 8C illustrates the binding activities of the dimeric humanized anti-PEG (hE11) BsAbs of example 1.4 with the PEGylated Quantum Dot (Qdot655) in Jurkat (TAG-72+), MDA-MB-468 (EGFR+) or BT-474 (HER2+) cells in accordance with one embodiment of the present disclosure;
[0040] FIG. 9 is a schematic illustration of DNA constructs for humanized monovalent anti-PEG (hE11) BsAbs of example 2.1 and the structure of the monovalent BsAb in accordance with one embodiment of the present disclosure;
[0041] FIG. 10 illustrates the cancer cell selectivity of the humanized monovalent anti-PEG (hE11) BsAbs of example 2.1 in Jurkat (TAG-72+), MDA-MB-468 (EGFR+) or BT-474 (HER2+) cells in accordance with one embodiment of the present disclosure;
[0042] FIG. 11 illustrates the binding activities of the humanized monovalent anti-PEG (hE11) BsAbs of example 2.1 with the PEGylated liposomal Texas Red in Jurkat (TAG-72+), MDA-MB-468 (EGFR+) or BT-474 (HER2+) cells in accordance with one embodiment of the present disclosure;
[0043] FIG. 12 illustrates the binding activities of the humanized monovalent anti-PEG (hE11) BsAbs of example 2.1 with the PEGylated Quantum Dot (Qdot655) in Jurkat (TAG-72+), MDA-MB-468 (EGFR+) or BT-474 (HER2+) cells in accordance with one embodiment of the present disclosure;
[0044] FIG. 13A is a schematic illustration of DNA constructs for humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 and the structure of the BsAb in accordance with one embodiment of the present disclosure;
[0045] FIG. 13B illustrates the SDS-PAGE analysis of the humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 in reducing or non-reducing conditions in accordance with one embodiment of the present disclosure;
[0046] FIGS. 14A and 14B respectively illustrate the antigen-binding activity of the humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 towards (A) NH.sub.2-PEG.sub.10,000-NH.sub.2 and (B) BSA in accordance with one embodiment of the present disclosure;
[0047] FIG. 14C illustrates the binding activities of the humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 with the PEGylated Quantum Dot (Qdot655) or PEGylated liposomal Texas Red in Raji (CD19+) or A431 (EGFR+) cells in accordance with one embodiment of the present disclosure;
[0048] FIGS. 14D and 14E respectively illustrate the binding kinetics of the humanized monovalent anti-PEG (h6.3) and antiCD19 BsAbs of example 2.2 towards CH.sub.3-PEG.sub.5,000-Alexa647 in accordance with one embodiment of the present disclosure;
[0049] FIG. 15 is a panel of real-time images illustrating the endocytic activity of the humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 with the PEGylated Quantum Dot (Qdot655) in A431 (EGFR+) cells in accordance with one embodiment of the present disclosure;
[0050] FIGS. 16A to 16C are line graphs respectively illustrate the enhanced in-vitro cytotoxity of Lipo/DOX by the humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 in (A)A431 cells (EGFR+), (B) MDA-MB-468 cells (EGFR+) and (C) Raji cells (CD19+) in accordance with one embodiment of the present disclosure;
[0051] FIG. 17 illustrates the tumor imaging enhancement of the humanized monovalent anti-PEG (h6.3) BsAbs of example 2.2 targeted PEG-NIR797 against CD19.sup.+ and EGFR.sup.+ tumor in accordance with one embodiment of the present disclosure;
[0052] FIG. 18A is a schematic illustration of DNA constructs for humanized anti-mPEG BsAbs in accordance with one embodiment of the present disclosure;
[0053] FIG. 18B illustrates the SDS-PAGE analysis of the humanized BsAbs of example 2.3 in reducing or non-reducing condition in accordance with one embodiment of the present disclosure;
[0054] FIGS. 18C and 18D illustrate the respective binding activities of the humanized BsAbs of example 2.3 with the indicated PEG-NPs in SW480 cells (EGFR.sup.+) (FIG. 2C) and SK-BR-3 cells (HER2.sup.+) (FIG. 2D) in accordance with one embodiment of the present disclosure;
[0055] FIG. 19A illustrates the cancer cell selectivity of PEG-NPs treated with the humanized BsAbs of example 2.3 in SW480 cells (EGFR.sup.+) and SW620 cells (EGFR.sup.-) (FIG. 2C) in accordance with one embodiment of the present disclosure;
[0056] FIG. 19B illustrates the cancer cell selectivity of PEG-NPs treated with the PEG.times.EGFR or PEGxHER2 of example 2.3 in SK-BR-3 cells (HER2.sup.+) and MDA-MB-468 cells (HER2.sup.-) in accordance with one embodiment of the present disclosure;
[0057] FIGS. 20A to 20D respectively illustrate the enhanced in-vitro cytotoxity of Lipo/DOX by PEG.times.EGFR of example 2.3 in (A) SW480 cells (EGFR.sup.+), (B) SW620 cells (EGFR.sup.-), (C) SK-BR-3 cells (HER2.sup.+), and (D) MDA-MB-468 cells (HER2.sup.-) in accordance with one embodiment of the present disclosure;
[0058] FIG. 21 is a panel of in vivo imaging of PEG.times.EGFR of example 2.3 targeting Lipo/IR780 in accordance with one embodiment of the present disclosure; and
[0059] FIGS. 22A and 22B illustrate the respectively size of EGFR.sup.+ and EGFR.sup.- tumors treated with PEG.times.EGFR targeted Lipo/Dox in accordance with one embodiment of the present disclosure; and
[0060] FIG. 22C is a line graph illustrating the changes in body weight of the test animals in FIGS. 22A and 22B;
[0061] FIG. 23A is a schematic drawing of DNA constructs for humanized anti-mPEG (h15-2b) anti-CD19 BsAb and anti-mPEG (h15-2b) anti-CD20 BsAb in accordance with one embodiment of the present disclosure;
[0062] FIG. 23B illustrates the cancer cell selectivity of the BsAbs of example 2.4 in Raji cells in accordance with one embodiment of the present disclosure;
[0063] FIG. 23C illustrates the mPEG binding activity of the BsAbs of example 2.4 in accordance with one embodiment of the present disclosure;
[0064] FIG. 23D illustrates the dual binding activity of the BsAbs of example 2.4 in accordance with one embodiment of the present disclosure;
[0065] FIG. 24A is a schematic illustration of DNA constructs for humanized knob in hole anti-PEG (h15-2b) BsAbs of example 3.1 and the structures of the BsAbs in accordance with one embodiment of the present disclosure;
[0066] FIG. 24B is a schematic illustration of DNA constructs for humanized knob in hole anti-PEG (h6.3) BsAbs of example 3.1 and the structures of the BsAbs in accordance with one embodiment of the present disclosure;
[0067] FIG. 24C illustrates the SDS-PAGE analysis of the humanized knob in hole anti-PEG (h15-2b or h6.3) BsAbs of example 3.1 in non-reducing condition in accordance with one embodiment of the present disclosure;
[0068] FIG. 25 illustrates the cancer cell selectivity of the humanized knob in hole anti-PEG (h15-2b) BsAbs of example 3.1 in Ramous (CD19+), Raji (CD19+) and SKBR3 (HER2.sup.+) cells in accordance with one embodiment of the present disclosure;
[0069] FIG. 26 illustrates the dual binding activities of the humanized knob in hole anti-PEG (15-2b) BsAbs of example 3.1 with the PEGylated Quantum Dot (Qdot655) in Ramos (CD19+), Raji (CD19+) and SKBR3 (HER2.sup.+) cells in accordance with one embodiment of the present disclosure;
[0070] FIG. 27 illustrates the cancer cell selectivity of the humanized knob in hole anti-PEG (h6.3) BsAbs of example 3.1 in Ramos (CD19+), Raji (CD19+) and SKBR3 (HER2.sup.+) cells in accordance with one embodiment of the present disclosure;
[0071] FIG. 28 illustrates the dual binding activities of the humanized knob in hole anti-PEG (h15-2b or h6.3) BsAbs of example 3.1 with the PEGylated Quantum Dot (Qdot655) in Raji (CD19+) and SKBR3 (HER2.sup.+) cells in accordance with one embodiment of the present disclosure;
[0072] FIG. 29 is a schematic illustration of DNA constructs for BsAbs of example 4.1 in accordance with one embodiment of the present disclosure;
[0073] FIGS. 30A and 30B respectively illustrate the enhanced in-vitro cytotoxicity of Lipo/DOX by BsAbs of example 4.1 in (A) SKBR3 cells (HER2.sup.+) and (B) A431 cells (EGFR.sup.+) in accordance with one embodiment of the present disclosure; and
[0074] FIGS. 31A and 31B respectively illustrate the synergistic anti-cancer effects of Lipo/DOX by BsAbs of example 4.1 in (A) SKBR3 cells (HER2.sup.+) and (B) A431 cells (EGFR.sup.+) in accordance with one embodiment of the present disclosure; and
[0075] FIG. 32 illustrates the synergistic anti-cancer effects of Lipo/DOX by BsAbs of example 4.1 in SKBR3 cells (HER2.sup.+) in accordance with one embodiment of the present disclosure.
DESCRIPTION
[0076] The detailed description provided below in connection with the appended drawings is intended as a description of the present examples and is not intended to represent the only forms in which the present example may be constructed or utilized. The description sets forth the functions of the example and the sequence of steps for constructing and operating the example. However, the same or equivalent functions and sequences may be accomplished by different examples.
I. Definition
[0077] The term "antibody" is used in the broadest sense and specifically covers monoclonal antibodies, polyclonal antibodies, multispecific antibodies (e.g., bi-specific antibodies), and antibody fragments so long as they exhibit the desired biological activity, that is, to specifically bind to an antigen when it preferentially recognizes its target antigen in a complex mixture of proteins and/or other molecules.
[0078] The term "monoclonal antibody" as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, and is not to be constructed as requiring production of the antibody by any particular method. In contrast to polyclonal antibodies which typically include different antibodies directed to different epitopes, each monoclonal antibody is directed against a single determinant (i.e., epitope) on the antigen. The monoclonal antibodies of the present disclosure may be made by the hybridoma method or by recombinant DNA methods. The monoclonal antibodies herein specifically include "chimeric" or "recombinant" antibodies, in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies derived from a particular species or belonging to an antibody class or subclass, while the remainder of the chain identical with or homogolous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies, as long as they exhibit the desired biological activity.
[0079] "Humanized" forms of non-human (e.g., murine) antibodies are chimeric antibodies which contain minimal sequence derived from non-human immunoglobulin. Humanized antibodies are human immunoglobulins in which hypervarible region residues are replaced by hypervarible region residues from a non-human species such as mouse, rat, rabbit, or non-human primate having the desired specificity or affinity. In some instances, Fv framework region (FR) residues of the human immunoglobulin are replaced by corresponding non-human residues. In general, the humanized antibody will comprise substantially all of at least one, and typically two, variable domains, in which all or substantially all of the FR regions are those of a human immunoglobulin sequence. The humanized antibody may optionally comprise at least a portion of an immunoglobulin constant region (Fc), typically that of a human immunoglobulin.
[0080] An "isolated" antibody is one which has been identified and separated and/or recovered from a component of its nature environment. Containment components of its nature environment are materials which would interfere with therapeutic uses of the antibody of this invention, and may include enzymes, hormones, and other protenaceous or non-proteinaceous solutes. Isolated antibody includes the antibody in situ within recombinant cells. Ordinarily, isolated antibody will be prepared by at least one purification step.
[0081] The term "bi-specific antibody (BsAb)" refers to an antibody having specificities for at least two different antigens. For example, BsAb may have one arm having a specificity for one antigenic site, such as a tumor associated antigen, while the other arm recognizes a different target, for example, a haptan that is bound to a lethal agent (e.g., INF-.alpha. or a liposome containing an anti-cancer agent such as vinca alkaloid) or an imaging agent (e.g., a microbubble containing a contrast agent or a quantum dot or fluorescent dye). In preferred embodiments, the BsAb of the present disclosure has two antigen-binding sites, in which one is directed against a tumor antigen (e.g., TAG72, CD19, EGFR or HER2), while the other is directed against a hydrophilic polymer (e.g., polyethylene oxide (PEG)), that is bound to a nanoparticle containing a cancer therapeutic agent therein (e.g., Lipo/DOX).
[0082] The term "valent" as used herein refers to the presence of a specified number of binding sites in an antibody molecule. As such, the term "monovalent", "divalent", "trivalent" and tetravalent" refer to the presence of 1, 2, 3, and 4 binding sites, respectively in an antibody molecule. The BsAb of the present disclosure is at least "divalent", and may be multivalent, such as tetravalent.
[0083] The term "linker" and "peptide linker" are interchangeably used in the present disclosure and refers to a peptide having natural or synthetic amino acid residues for connecting two polypeptides. For example, the peptide linker may be used to connect the VH and the VL to form the single chain variable fragment (e.g., scFv); or to connect the scFv to the full length antibody to form a BsAb of the present disclosure. Preferably, the linker is a peptide having at least 5 amino acid residues in length, such as 5 to 100 amino acid residues in length, more preferably 10 to 30 amino acid residues in length. The linker within scFv is a peptide of at least 5 amino acid residues in length, preferably 15 to 20 amino acid residues in length. In one example, the linker comprises a sequence of (G.sub.nS).sub.m, with G=glycine, S=serine, n is a number between 1 to 4, and m is 1, 2 or 3. Preferably, the linker comprises a sequence of (G.sub.4S).sub.3; or a sequence of (G.sub.3S) and (G.sub.3S.sub.2).
[0084] The term "PEGylated substance" as used herein refers to a substance coated with polyethylene glycol (PEG), which includes but is not limited to, a protein (e.g., a chemokine), a peptide (e.g., leuprolide) and a nanoparticle (NP) containing therein a therapeutic agent or an imagine agent. Materials known in the state of the art that may give rise to the nanoparticle includes mesoporpous silica, as well as the material that has a hydrophilic portion and a hydrophobic portion that forms a micelle structure capable of including a therapeutic agent (e.g., anti-cancer agent) or an imaging agent (e.g., a fluorescence dye, a quantum dot, a chelated radioisotope, a paramagnetic iron, gold nanoparticle or a contrast agent) within its structure. Suitable materials for forming nanoparticles in the present disclosure include, but are not limited to, mesoporpous silica; phospholipids such as phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylserine (PS), phosphatidylglycerol (PG), phosphatidic acid (PA), phosphatidylinositol (P1), sphingomyelin (SPM), and the like, alone or in combination; biodegrable polymer such as polylactic acid (PLA), polyglycolic acid (PGA) poly(lactic-co-glycolic acid) (PLGA), polycaprolactone (PCL), polydioxanone (PDO), polyanhydrides, polyorthoesters, chitosan and the like, alone or in combination. Preferably, the PEGylated substance, such as a PEGylated NP, further contains a cancer therapeutic agent or an imagine agent within the micelle structure.
[0085] The terms "cancer" and "tumor" are used alternatively in the present disclosure and preferably refer to the physiological condition in mammals and especially in humans that is typically characterized by un-regulated cell growth. Cancers in this respect include metastases cancers, and/or drug-resistant cancers. Cancers, preferably those exhibit increased expression levels of TAG72, EGFR, HER2, CD19, and CD20. Accordingly, cancers or tumors treatable by the present disclosure are breast, lung, colon, colorectal, spleen, kidney, liver, bladder, head and neck, ovary, prostate, brain, pancreas, skin, bone, blood, thymus, uterus, testicles, cervix, and neuron. More specifically, the cancer is selected from the group consisting of breast cancer, colorectal cancer, head and neck cancer, colon cancer, hepatic cancer, non-Hodgkin's lymphoma, lymphoma, pancreatic cancer, lung cancer, gastric cancer, prostate cancer, brain tumor, retinoblastoma, ovarian cancer, cervical cancer, hematopoietic malignances, esophageal cancer, renal cell carcinoma, squamous cell carcinoma, glioma, and leukemia
[0086] The term "therapeutic agent(s)" as used herein refers to an agent utilized to treat, combat, ameliorate, prevent or improve a disease or a condition, such as a cancer, in a patient. Accordingly, therapeutic agent(s) for treating cancer preferably refers to cytotoxic agents that are known to improve the therapeutic effects of a cancer treatment; accordingly, cytotoxic agents as used in the present disclosure include, but are not limited to, radiation, chemotherapeutic agents, antibodies, and the like.
[0087] The term "drug-resistant cancer" as used herein refers to a cancer whose growth is not suppressed or retarded by the application of a well-known cytotoxic agent, which may be a chemotherapeutic agent, an antibody, a peptide or a combination thereof. In some embodiments, the drug is a chemotherapeutic agent. Examples of chemotherapeutic agent include alkylating agent such as nitrosoureas, cisplatin, or dacarbazine; antimetabolites such as folic acid, purine or pyrimidine antagonists; mitotic inhibitors such as vinca alkaloids; cytotoxic antibiotics and camptothecin derivatives. Preferred chemotherapeutic agent includes adriamycin, amifostine, bleomycin, busulfan, cisplatin, and/or other platinum compounds, preferably including carboplatin and/or oxaliplatin, camptothecin, CPT-11, cytosine arabinoside, chlorambucil, cyclophosphamide, cytarabine, daunorubicin, doxorubicin, docetaxel, dacarbazine, dactinomycin, etoposide, 5-fluorouracil (5-FU), fluoxuridine, gemcitabine, hydroxyurea, ifosfamide, idarubicin, interferon beta, irinotecan, L-asparaginase, L-aspartic acid, lomustine, mechlorethamine, mitomycin, methotrexate, mitoxantrone, megestrol, melphalan, mercaptopurine, mitotane, paclitaxel (taxol), plicamycin, pentostatin, streptozocin, topotecan, tamoxifen, teniposide, thioguanine, vinblastine, vincristine, and a combination thereof. In other embodiments, the drug is a chemokine (e.g., CC chemokine, CXC chemokine, C chemokine and CX.sub.3C chemokine) or a cytokine (e.g., interferone, interleukin, lymphokine, and tumor necrosis factor). In further embodiments, the drug is a peptide, preferably a peptide with cytotoxicity effects toward cancer cells. Preferably, the anti-cancer peptide is selected from the group consisting of leuprolide, goserelin, octreotide, histrelin, abarelix, cetrorelix, degarelix, cilengtide, ATN-161, and IM862.
[0088] The term "therapeutically effective amount" as used herein refers to an amount effective, at dosages, and for periods of time necessary, to achieve the desired therapeutically desired result with respect to the treatment of cancers, including metastatic and/or drug-resistant cancers.
[0089] The phrase "pharmaceutically acceptable" refers to molecular entities and compositions that are "generally regarded as safe", e.g., that are physiologically tolerable and do not typically produce an allergic or similar untoward reaction, such as gastric upset, dizziness and the like, when administered to a human. Preferably, as used herein, the term "pharmaceutically acceptable" means approved by a regulatory agency of the Federal or a state government or listed in the U.S. Pharmacopeia or other generally recognized pharmacopeia for use in animals, and more particularly in humans.
[0090] The term "administered", "administering" or "administration" are used interchangeably herein to refer means either directly administering a bi-specific antibody or a composition of the present disclosure.
[0091] The term "subject" or "patient" refers to an animal including the human species that is treatable with the compositions and/or methods of the present disclosure. The term "subject" or "patient" intended to refer to both the male and female gender unless one gender is specifically indicated. Accordingly, the term "subject" or "patient" comprises any mammal which may benefit from treatment of cancer. Examples of a "subject" or "patient" include, but are not limited to, a human, rat, mouse, guinea pig, monkey, pig, goat, cow, horse, dog, cat, bird and fowl. In an exemplary embodiment, the patient is a human.
[0092] The term "identical" or "percent identity" as used herein refers to two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues that are the same, when compared and aligned for maximum correspondence. To determine the percent identity, the sequences are aligned for optimal comparison purposes (e.g., gaps can be introduced in the sequence of a first amino acid sequence for optimal alignment with a second amino acid sequence). The amino acid residues at corresponding amino acid positions are then compared. When a position in the first sequence is occupied by the same amino acid residue as the corresponding position in the second sequence, then the molecules are identical at that position. The percent identity between the two sequences is a function of the number of identical positions shared by the sequences (i.e., % identity=number of identical positions/total number of positions (e.g., overlapping positions).times.100). In certain embodiments, the two sequences are the same length.
[0093] The singular forms "a", "and", and "the" are used herein to include plural referents unless the context clearly dictates otherwise.
II. Description of the Invention
[0094] Accordingly, it is the first aspect of the present disclosure to provide bi-specific antibodies (BsAbs) that convert a non-targeted PEGylated substance to tumor-targeted PEGylated substance and thereby suppress the growth of a cancer or blocking the invasion or metastasis of a cancer, including drug-resistant cancer.
[0095] 1. The Structures of BsAbs of the Present Disclosure
[0096] Antibodies belong to the immunoglobulin class of proteins that includes IgG, IgA, IgE, IgM, and IgD. The most abundant immunoglobulin found in serum is IgG, whose schematic structure is illustrated in FIG. 1. The IgG structure has four chains, two light chains and two heavy chains; each light chain has two domains and each heavy chain has four domains. The antigen-binding site is located in the fragment antigen binding (Fab) region that contains a variable light (VL) and variable heavy (VH) chain domains as well as a constant light (CL) and constant heavy (CH1) domains. The CH2 and CH3 domain region of the heavy chain is called fragment crystallizable (Fc) region. A full length antibody heavy chain is therefore a polypeptide consisting of, from N-terminus to C-terminus, a VH, a CH1, a hinge region (HR), a CH2, and a CH3; abbreviated as VH-CH1-HR-CH2-CH3. A full length antibody light chain is a polypeptide consisting in N-terminus to C-terminus direction of a VL and a CL, abbreviated as VL-CL, in which the CL can be .kappa. (kappa) or .lamda. (lambda). The IgG is regarded as a heterotetramer having two heavy chains that are held together by disulfide bonds (--S--S--) between the CL domain and the CH1 domain and between the hinge regions of the two heavy chains.
[0097] As stated above in the "definition" section, the BsAbs refer to Abs having specificities for at least two different antigens; hence, BsAbs of the present disclosure is a recombinant Ab engineered to contain sequences capable of binding to different antigens. Accordingly, various recombinant bi-specific antibody formats have been developed in the present disclosure, and the schematic structures of these BsAbs are illustrated in FIGS. 2A to 2E.
[0098] In some embodiments, the BsAb of the present disclosure is a dimeric, tetravalent bi-specific antibody, in which the two heavy chains of a full length IgG directed to the first antigens are respectively fused to single chain variable fragments (e.g., scFv) directed to the second antigens via peptide linkers (FIG. 2A). The scFv, preferably a disulfide-stabilized scFv, consists of an antibody heavy chain variable domain (VH) and an antibody light chain variable domain (VL), and a linker; abbreviated as VH-linker-VL.
[0099] Alternatively, the BsAb of the present disclosure may be a monomeric, divalent bi-specific antibody, in which a VH-CH1 domain and a light chain VL-CL domain directed to a first antigen is fused via a peptide linker to a disulfide stabilized single chain domain directed to a second antigen (FIG. 2B).
[0100] In some embodiments, the BsAb of the present disclosure is a monomeric, divalent bi-specific antibody, in which a disulfide stabilized single chain domain directed to the first antigen is connected to a monomeric antibody directed to a second antigen via a peptide linker (FIG. 2C).
[0101] In other embodiments, the BsAb of the present disclosure has a "knob into hole" structure, in which a knob in the CH3 domain of the first heavy chain is created by replacing several amino acids with alternative amino acids, and a hole in the juxtaposed position at the CH3 domain of the second heavy chain is created by replacing appropriate amino acid with alternative ones. In addition, cysteine residues are introduced to form a disulfide bond linkage between the heavy chains. A schematically presentation of the "knob into hole" BsAb structure is as depicted in FIG. 2D.
[0102] In further embodiments, the "knob in hole" BsAb as depicted in FIG. 2D is further modified, in which a monomeric antibody heavy chain is crossovered with its light chain during transcription, and thereby creating a modified antibody heavy chain hetero-polypeptide consisting in N-terminus to C-terminus direction of a VH, a CL, a hinge region (HR), a CH2, and a knob-CH3; abbreviated as VH-CL-HR-CH2-knob-CH3; and a modified antibody light chain hetero-polypeptide consisting in N-terminus to C-terminus direction of a VL and a CH1; abbreviated as VL-CH1. FIG. 2E is a schematic drawing of this modified "knob into hole" BsAb structure, in which one monomeric antibody heavy chain is crossovered with its light chain, while the other monomeric antibody structure remains unchanged.
[0103] 2. Antibody Preparation
[0104] Methods for preparing the BsAbs of the present disclosure are described in the Examples. In order to prepare a humanized BsAb, a non-human (e.g., murine) antibody is prepared and used as a starting material; relevant technology is briefly described in the following section.
[0105] 2.1 Production of Murine Anti-mPEG Antibody
[0106] To produce the desired monoclonal antibodies, animals such as mice, rats or rabbits are first immunized with mPEG-derivatized proteins (i.e., the PEG molecule has a terminal methoxy group) molecule or PEG-derivatized proteins (i.e., the PEG molecule has a terminal hydroxyl group) at a suitable dose. Generally, adjuvant and the mPEG- or PEG-derivatized protein solution are mixed together when immunizing the animals with mPEG- or PEG-derivatized proteins. Examples of adjuvants useful for this invention include Freund's complete adjuvant (FCA), Freund's incomplete adjuvant (FIA), and aluminum hydroxide adjuvant. Immunization is generally carried out mainly by intravenous, subcutaneous, intraperitoneal or intramuscular injection of the antigen. The immunization interval is not particularly limited. Immunization may be carried out at intervals of several days to several weeks, preferably 2 to 3 weeks, for 1 to 10 times, preferably 2 to 5 times. Once antibody titers in serum samples diluted by 1000 fold reaches 2 or more in the absorbance level as the result of immunization, the animals are left for about 1 month
[0107] Then, re-immunization is carried out for at least once. Several days, preferably 3 to 5 days, after the final immunization, splenic cells and regional lymph nodes are removed. Blood samples are taken regularly after immunization and subject to centrifugation to separate sera. The resultant sera are then subject to measurement of antibody titers by any suitable method, which includes, and is not limited to, enzyme linked immunosorbent assay (ELISA), enzyme immunoassay (EIA), or radio immunoassay (RIA). In one preferred example, antibody titers are measured by ELISA. Then, final immunization is given to those animals showing high antibody titers to mPEG- or PEG-derived protein isoforms.
[0108] Antibody-producing cells are prepared from splenic cells and regional lymph nodes or the like of the immunized animals. In the preparation of antibody-producing cells, it is preferably to remove tissue debris and erythrocytes as much as possible. Commercial erythrocyte remover may be used to this purpose. Alternatively, a buffer ammonium chloride and Tris may be prepared and used.
[0109] The thus prepared antibody-producing cells should be immediately fused with immortal cells such as myeloma cells to produce hybridoma cells, which semi-eternally continue to proliferate while producing antibodies. Commonly available cell strains derived from an animal such as a mouse may be used. A preferable cell strain to be used in this invention should be those that fuse efficiently, support stable high level production of antibody and are sensitive to HAT selection medium, which contains hypoxanthine, thymidine and aminopterin, and should survive there only when fused with antibody-producing cells. Examples of myeloma cells include, but are not limited to, mouse myeloma cell line (such as myeloma FO cells) and human myeloma cell line (such as Karpas 707H).
[0110] Cell fusion is usually carried out by mixing splenic cells or lymph node cells with a commercial available myeloma cells in the presence of a cell-fusion promoter, such as PEG having an average molecular weight from about 200 to 20,000 daltons or the like. Alternatively, cell fusion may be carried out in a commercial cell fusion device utilizing electric stimulation such as electro-fusion. After the fusion, the resultant cells are then diluted and cultured in HAT medium.
[0111] Hybridomas of interest are then selected from the fused cells. The fused cells surviving cultured in HAT medium would form colonies. The supernatant of each culture well is then collected and examined for the presence or absence of antibody titers to mPEG- or PEG-derivatizeded proteins. As a method of confirmation, ELISA, EIA or RIA may be used, in which CH.sub.3-PEG.sub.750-NH.sub.2 or NH.sub.2-PEG.sub.3000-NH.sub.2 is coated onto the plates and used as a screening criteria. Once antibody-positive wells are identified, cells are then cultured in a HT medium, which does not contain aminopterin. After culturing for a while, antibody titers in the culture supernatant are confirmed again. Cells that are finally selected are then subject to cloning to obtain single cells. Clones that exhibit high specificity to mPEG- or PEG-derived proteins are selected, and are proliferated to some extent to establish hybridomas.
[0112] According to preferred embodiments of the present disclosure, 3 hybridomas, E11, 15-2b and 6-3, were selected. The 15-2b hybridoma produced an anti-mPEG monoclonal antibody that specifically bound to terminal methoxy or hydroxyl group, but not the backbone, of PEG. By contrast, the E11 and 6-3 hybridomas, produced anti-PEG backbone monoclonal antibodies that bound to the backbone, instead of the end methoxy or hydroxyl group of PEG.
[0113] In some embodiments, the anti-mPEG monoclonal antibodies were selected over the anti-PEG backbone monoclonal antibodies due to space homogeneity rendered by anti-mPEG Abs once they were bound with PEGylated nanoparticles. In other embodiments, the anti-PEG backbone monoclonal antibodies were selected over the anti-mPEG monoclonal antibodies.
[0114] The thus produced anti-mPEG or anti-PEG monoclonal antibodies may be isolated or prepared by any known method. For example, antibodies may be prepared from cultured supernatant obtained by culturing hybridomas in a medium with low serum concentration. Alternatively, hybridomas may be injected into abdominal cavities of animals and the resultant abdominal dropsies are collected to prepare antibodies. Antibodies may be purified or isolated by methods that employ affinity column, gel filtration chromatography, ion exchange chromatography or the like. Any of these known methods may be appropriately selected or used in combination.
[0115] Alternatively, anti-mPEG or anti-PEG monoclonal antibodies may be produced by DNA cloning. DNA encoding anti-mPEG or anti-PEG mAbs may be easily isolated and sequenced by use of conventional procedures, such as using oliognucleotide probes that are capable of binding specifically to genes encoding the heavy and light chains of the monoclonal antibodies. The hybridoma cells (e.g., E11, 6-3 or 15-2b hybridoma) serve as a preferred source of such DNA. Once isolated, the DNA may be placed into expression vectors, which are then transfected into host cells such as E. Coli cells, simian COS cells or Chinese hamster ovary (CHO) cells or myeloma cells that do not produce immunoglobulin proteins, to synthesize the desired monoclonal antibodies in the recombinant host cells.
[0116] The monoclonal antibodies thus produced and the DNA encoding such antibodies can then be used to produce chimeric antibodies (e.g., bi-specific antibodies), humanized antibodies and/or antibody fragments derived thereof.
[0117] 2.2 Production of Humanized Anti-mPEG (15-2) or Anti-PEG (E11 or 6.3) Antibody
[0118] The major concern of a non-human origin monoclonal antibody is its immunogenicity to the recipient, in some cases, caused dangerous allergic reactions. Most monoclonal antibodies are of murine origin, and have been found to be immunogenic when injected to human. To reduce the immunogenicity of anti-mPEG or anti-PEG mAbs of this invention, humanized antibodies are produced by attaching variable domains in the heavy and light chains of murine anti-mPEG or anti-PEG Abs onto the constant regions of human antibodies.
[0119] To create humanized anti-mPEG or anti-PEG antibodies, the DNA encoding such antibodies was isolated and sequenced in accordance with methods described above in section 2.1, and then used to create humanized constructs. Detailed production method is set forth in the Examples.
[0120] According to preferred embodiments of the present disclosure, CDR (complementary determining region) grafting is employed, in which the CDR regions in the VH and VL genes of a human antibody are replaced with the appropriate CDR coding segments (such as those DNA segments in anti-mPEG or anti-PEG Abs that code amino acid segments responsible for binding PEG). The resulting antibodies therefore have variable regions in which only the CDRs are from the original mouse antibodies, while the framework regions in the VH and VL genes as well as the constant region genes (i.e., C.kappa. or CH1-H-CH2-CH3) are those of human IgG.
[0121] In preferred embodiments, the humanized anti-mPEG or anti-PEG Ab comprises a heavy chain variable domain and a light chain variable domain. Once produced, the humanized anti-mPEG or anti-PEG Abs may be purified according to standard procedures in the art, including cross-flow filtration, affinity column chromatography, gel filtration and the like. It should be understood that the humanized antibodies shall perform in a manner identical or substantially similar to that of murine anti-mPEG Abs. Preferably, the humanized anti-mPEG or anti-PEG Abs (either in the form of Fab or full length IgG) shall be more advantages to use in a human subject, as compared to the murine version. In some embodiments, the humanized anti-mPEG Abs are used in the production of bi-specific antibodies of the present disclosure. In other embodiments, the humanized anti-PEG Abs are used in the production of bi-specific antibodies of the present disclosure.
[0122] 2.3 Production of Bi-Specific Monoclonal Antibodies (BsAbs)
[0123] To produce BsAbs, the humanized anti-mPEG or anti-PEG Abs (either in the form of Fab or a full length IgG) described above in Section 2.2 are further linked with antibodies or scFv that bind tumor antigens, so as to confer cancer targeting effect. Detailed production method is set forth in the Examples.
[0124] In general, DNA sequences of the above humanized anti-mPEG or anti-PEG Abs including the heavy and light chains of humanized anti-mPEG or anti-PEG sequences are ligated with DNA sequence of a desired antibody or scFv that binds a tumor antigen via use of a linker, then the chimeric sequence is cloned into an expression vector for transfecting a host cell, and subsequently purified in accordance with similar steps described above in section 2.2. The thus produced BsAbs may then be used to treat cancers or to track the developments of cancers with an aid of an imaging system.
[0125] Accordingly, humanized monomeric and dimeric antibodies are produced, with bi-specificities to both PEGylated molecules and tumor antigens, which include, but are not limited to, TAG72, EGFR, HER2, CD19, and CD20.
[0126] In some embodiments, monomeric BsAbs including PEG.times.EGFR (anti-PEG anti-EGFR), PEGxTAG72 (anti-PEG anti-TAG72), and PEGxHER2 (anti-PEG anti-HER2) are produced, with the anti-PEG portion derived from the hE11 Fab fragment. In another embodiment, monomeric h6.3 Fab.times.EGFR (anti-PEG anti-EGFR) and h6.3 Fab.times.CD19 (anti-PEG anti-CD19) are produced, in which h6.3 Fab, instead of hE11 Fab, is fused with scFv against EGFR or CD19. In a further embodiment, monomeric h15-2b Fab.times.EGFR scFv (anti-PEG anti-EGFR), h15-2b Fab.times.HER2 scFv (anti-PEG anti-HER2), are produced, in which h15-2b Fab is fused with scFv against EGFR or HER2. In still further embodiments, monomeric h15-2b scFv xCD19 Fab (anti-PEG anti-CD19) and h15-2b scFv xCD20 Fab (anti-PEG anti-CD20) are produced, in which h15-2b scFv is fused with Fab against CD19 or CD20.
[0127] In other embodiments, dimeric BsAbs, including PEG2.times.EGFR (anti-PEG anti-EGFR), PEG2.times.TAG72 (anti-PEG anti-TAG72), and PEG2.times.HER2 (anti-PEG anti-HER2) are produced. Unlike the monomeric BsAb, each dimeric BsAb includes a full length IgG, with each heavy chain being linked to the scFv that binds a tumor antigen (e.g., TAG 72, EGFR or HER2). Further, monomeric BsAbs of PEG.times.EGFR, PEG.times.HER2, and PEG.times.TAG72 of the present disclosure differ from their counterparts in the dimeric forms (i.e., PEG2.times.EGFR, PEG2.times.HER2, and PEG2.times.TAG72) in that they do not possess HR-CH2-CH3 domains in their respective structures.
[0128] In still some other embodiments, "knob in hole" BsAbs are created, in which DNA sequences encoding antibody heavy chains, particularly the CH3 domains of the two heavy chains, are designed to introduce specific and complementary interactions at the interface of the respective CH3 domains of the two heavy chains. For example, several amino acids are substituted with alternative amino acids in the first heavy chain CH3 domain to create a "knob" structure, and several amino acids in the second heavy chain CH3 domain are altered to create a "hole" such that antibody heavy chains expressed from these DNA sequences are unlikely to form a combination of just the first pairs or just the second pairs, but rather the "knob in hole" heavy chain pairs. The knob-in-hole technique is well known to those skilled in the art, and can be readily applied in forming the BsAbs of the present disclosure. Additionally, the "knob in hole" BsAbs may be further modified by crossing over the antibody heavy chain and the antibody light chain, and thereby creating an antibody heavy chain hetero-polypeptide consisting in N-terminus to C-terminus direction of a VH, a CL, a hinge region (HR), a CH2, and a knob-CH3; abbreviated as VH-CL-HR-CH2-knob-CH3; and an antibody light chain hetero-polypeptide consisting in N-terminus to C-terminus direction of a VL and a CH1; abbreviated as VL-CH1.
[0129] Accordingly, in one specific embodiment, a "knob in hole" anti-mPEG, anti-CD19 BsAb is produced. Specifically, two point mutations, S354C and T366W are introduced into the CH3 region of one h15-2b (anti-mPEG) heavy chain to create a knob structure; whereas additional four point mutations at S349C, T366S, L368A, and Y407V are introduced into the CH3 region of one BU12 (anti-CD19) heavy chain to generate a hole structure. In addition to creating the knob and hole structures on respective heavy chains, the Bu12-hole heavy chain may be further modified by crossing over with its light chain to generate a hetero heavy chain polypeptide and a hetero light chain polypeptide as described above. Therefore, each arms of the Y-shape h15-2b knob/Bu12-hole BsAb respectively recognize different antigens, that is, a PEGylated molecule and CD19. In one specific embodiment, h15-2b knob/HER2-hole BsAb is provided, in which the two arms of the Y-shape h15-2b knob/HER2-hole BsAb respectively recognize a PEGylated molecule and HER2.
[0130] The components and their respective amino acid sequences of BsAbs of the present disclosure are summarized in Tables 1 to 13.
TABLE-US-00001 TABLE 1 Amino Acid Sequence of PEG2 .times. TAG72 Name Amino Acid Sequence SEQ ID NO Humanized DVVMTQSPLSLPVTLGQPASISCRSSKSIVHSNGNTYLEWFQQR 1 E11 VL-C.kappa. PGQSPRRLIYKVSKRMSGVPDRFSGSGSGTDFTLKISRVEAEDV GVYYCSQGSHVPPTFGGGTKVEIKRTVAAPSVFIFPPSDEQLKS GTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGAEVKKPGASVKVSCKASGYTFTTYTMNWVRQAP 2 E11 GQGLEWMGYIIPSSGYVDYNQKFKGRVTMTRDTSTSTVYMEL VH-CH1 SSLRSEDTAVYYCVRSLDGYFWFAYWGQGTLVTVSSASTKGPS VFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGV HTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKV DKRV Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 3 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK Peptide VDLVTVSSASTGGGSGQLGGGGS 4 Linker Hcc49 dsFv QVQLVQSGAEVKKPGASVKVSCKASGYTFTDHAIHWVRQAPG 5 QCLEWMGYFSPGNDDFKYSQKFQGRVTITADKSASTAYMELSS LRSEDTAVYYCARSWIMQYWGQGTLVTVSSGGGGSGGGGSG GGGSDIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLA WYQQKPGQPPKLLIYWASTRESGVPDRFSGSGSGTDFTLTISSL QAEDVAVYYCQQYYSYPLTFGCGTKVEIK 6xHis Tag TRHHHHHH 6
TABLE-US-00002 TABLE 2 Amino Acid Sequence of PEG2 .times. EGFR Name Amino Acid Sequence SEQ ID NO Humanized DVVMTQSPLSLPVTLGQPASISCRSSKSIVHSNGNTYLEWFQQR 1 E11 VL-C.kappa. PGQSPRRLIYKVSKRMSGVPDRFSGSGSGTDFTLKISRVEAEDV GVYYCSQGSHVPPTFGGGTKVEIKRTVAAPSVFIFPPSDEQLKS GTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGAEVKKPGASVKVSCKASGYTFTTYTMNWVRQAP 2 E11 GQGLEWMGYIIPSSGYVDYNQKFKGRVTMTRDTSTSTVYMEL VH-CH1 SSLRSEDTAVYYCVRSLDGYFWFAYWGQGTLVTVSSASTKGPS VFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGV HTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKV DKRV Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 3 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK Peptide VDLVTVSSASTGGGSGQLGGGGS 4 Linker 11F8 QVQLQESGPGLVKPSQTLSLTCTVSGGSISSGDYYWSWIRQPPG 7 anti-EGFR KCLEWIGYIYYSGSTDYNPSLKSRVTMSVDTSKNQFSLKVNSV dsFv TAADTAVYYCARVSIFGVGTFDYWGQGTLVTVSSGGGGSGGG GSGGGGSEIVMTQSPATLSLSPGERATLSCRASQSVSSYLAWYQ QKPGQAPRLLIYDASNRATGIPARFSGSGSGTDFTLTISSLEPEDF AVYYCHQYGSTPLTFGCGTKAEIK 6xHis Tag TRHHHHHH 6
TABLE-US-00003 TABLE 3 Amino Acid Sequence of PEG2 .times. HER2 Name Amino Acid Sequence SEQ ID NO Humanized DVVMTQSPLSLPVTLGQPASISCRSSKSIVHSNGNTYLEWFQQR 1 E11 VL-C.kappa. PGQSPRRLIYKVSKRMSGVPDRFSGSGSGTDFTLKISRVEAEDV GVYYCSQGSHVPPTFGGGTKVEIKRTVAAPSVFIFPPSDEQLKS GTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGAEVKKPGASVKVSCKASGYTFTTYTMNWVRQAP 2 E11 GQGLEWMGYIIPSSGYVDYNQKFKGRVTMTRDTSTSTVYMEL VH-CH1 SSLRSEDTAVYYCVRSLDGYFWFAYWGQGTLVTVSSASTKGPS VFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGV HTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKV DKRV Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 3 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK Peptide VDLVTVSSASTGGGSGQLGGGGS 4 Linker C6ML3-9 QVQLLQSGAEVKKPGESLKISCKGSGYSFTSYWIAVVVRQMPG 8 anti-HER2 KGLEYMGLIYPGDSDTKYSPSFQGQVTISVDKSVSTAYLQWS dsFv SLKPSDSAVYFCARHDVGYCSSSNCAKWPEYFQHWGQGTLV TVSSGGGGSGGGGSGGGGSQSVLTQPPSVSAAPGQKVTISC SGSSSNIGNNYVSVVYQQLPGTAPKLLIYDHTNRPAGVPDRFS GSKSGTSASLAISGFRSEDEADYYCASWDYTLSGVVVFGGGT KLTVLG 6xHis Tag TRHHHHHH 6
TABLE-US-00004 TABLE 4 Amino Acid Sequence of h6.3 Fab .times. EGFR Name Amino Acid Sequence SEQ ID NO Humanized DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNQMNYLAWYQ 9 6.3 VL-C.kappa. QKPGQPPKLLIYWASTRESGVPDRFSGSGSGTDFTLTISSLQAED VAVYYCLQYLSSWTFGGGTKLEIKTYSLSSTLTLSKADYEKHK LYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGSELKKPGASVKVSCKASGYTFKNYGMNWVRQAP 10 6.3 GQGLEWMGWINTYTGQPIYANDFKGRFVFSLDTSVSTAYLQIS VH-CH1 SLKAEDTAVYYCARDWGPYWGQGTLVTVSSASTKGPSVFPLA PSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPA VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVE PKSCDK Peptide VDLVTVSSASTGGGSGQLGGGGS 4 Linker 11F8 QVQLQESGPGLVKPSQTLSLTCTVSGGSISSGDYYWSWIRQPPG 7 anti-EGFR KCLEWIGYIYYSGSTDYNPSLKSRVTMSVDTSKNQFSLKVNSV dsFv TAADTAVYYCARVSIFGVGTFDYWGQGTLVTVSSGGGGSGGG GSGGGGSEIVMTQSPATLSLSPGERATLSCRASQSVSSYLAWYQ QKPGQAPRLLIYDASNRATGIPARFSGSGSGTDFTLTISSLEPEDF AVYYCHQYGSTPLTFGCGTKAEIK 6xHis Tag TRHHHHHH 6
TABLE-US-00005 TABLE 5 Amino Acid Sequence of h6.3 Fab .times. CD19 Name Amino Acid Sequence SEQ ID NO Humanized DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNQMNYLAWYQ 9 6.3 VL-C.kappa. QKPGQPPKLLIYWASTRESGVPDRFSGSGSGTDFTLTISSLQAED VAVYYCLQYLSSWTFGGGTKLEIKTYSLSSTLTLSKADYEKHK LYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGSELKKPGASVKVSCKASGYTFKNYGMNWVRQAP 10 6.3 GQGLEWMGWINTYTGQPIYANDFKGRFVFSLDTSVSTAYLQIS VH-CH1 SLKAEDTAVYYCARDWGPYWGQGTLVTVSSASTKGPSVFPLA PSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPA VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVE PKSCDK Peptide VDLVTVSSASTGGGSGQLGGGGS 4 Linker hBU12 QVQLQESGPGLVKPSQTLSLTCTVSGGSISTSGMGVGWIRQHPG 11 dsPv KCLEWIGHIWWDDDKRYNPALKSRVTISVDTSKNQFSLKLSSV TAADTAVYYCARMELWSYYFDYWGQGTLVTVSSGGGGSGGG GSGGGGSEIVLTQSPATLSLSPGERATLSCSASSSVSYMHWYQQ KPGQAPRLLIYDTSKLASGIPARFSGSGSGTDFTLTISSLEPEDVA VYYCFQGSVYPFTFGCGTKLEIKR 6xHisTag TRHHHHHH 6
TABLE-US-00006 TABLE 6 Amino Acid Sequence of h15-2b Fab .times. HER2 scFv Name Amino Acid Sequence SEQ ID NO Humanized DIQMTQSPSSLSASVGDRVTITCKASQDVNTSVAVVYQQKPGK 12 15-2b APKLLIYWASTRHTGVPSRFSGSGSGTDFTFTISSLQPEDIATY VL-C.kappa. YCLQYINYPYTFGQGTKLEIKRTVAAPSVFIFPPSDEQLKSGTA SVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDS TYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized EVQLVESGGGLVQPGGSLKLSCAASGFTFSNYWMNVVVRQAS 13 15-2b GKGLEVVVGEIRSKSNNYATHYAESVKGRFTISRDDSKNTAYL VH-CH1 QMNSLKTEDTAVYYCSNRYYWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTF PAVLQSSGLYSLSSVVTVPSSNFGTQTYTCNVDHKPSNTKVD KTVERK G-MYC-(G GEQKLISEEDLGGGGSGGGGSGGGGSQL 14 4S)3 Linker C6ML3-9 QVQLLQSGAEVKKPGESLKISCKGSGYSFTSYWIAVVVRQMPG 15 (Anti-HER2) KGLEYMGLIYPGDSDTKYSPSFQGQVTISVDKSVSTAYLQWS scFv SLKPSDSAVYFCARHDVGYCSSSNCAKWPEYFQHWGQGTLV TVSSGGGGSGGGGSGGGGSQSVLTQPPSVSAAPGQKVTISC SGSSSNIGNNYVSVVYQQLPGTAPKLLIYDHTNRPAGVPDRFS GSKSGTSASLAISGFRSEDEADYYCASWDYTLSGVVVFGGGT KLTVLG
TABLE-US-00007 TABLE 7 Amino Acid Sequence of h15-2b Fab .times. EGFR scFv Name Amino Acid Sequence SEQ ID NO Humanized DIQMTQSPSSLSASVGDRVTITCKASQDVNTSVAVVYQQKPGK 12 15-2b APKLLIYWASTRHTGVPSRFSGSGSGTDFTFTISSLQPEDIATY VL-C.kappa. YCLQYINYPYTFGQGTKLEIKRTVAAPSVFIFPPSDEQLKSGTA SVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDS TYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized EVQLVESGGGLVQPGGSLKLSCAASGFTFSNYWMNVVVRQAS 13 15-2b GKGLEVVVGEIRSKSNNYATHYAESVKGRFTISRDDSKNTAYL VH-CH1 QMNSLKTEDTAVYYCSNRYYWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTF PAVLQSSGLYSLSSVVTVPSSNFGTQTYTCNVDHKPSNTKVD KTVERK G-MYC-(G GEQKLISEEDLGGGGSGGGGSGGGGSQL 14 4S)3 Linker h528 DIVMTQSPLSLPVTPGEPASISCRSSQNIVHNNGITYLEVVYLQK 16 (Anti-EGFR) PGQSPQLLIYKVSDRFSGVPDRFSGSGSGTDFTLKISRVEAED scFv VGVYYCFQGSHIPPTFGQGTKVEIKRAGGGGSGGGGSGGGG SQVQLVQSGAEVKKPGASVKVSCKASGYTFTSYWMHVVVRQ APGQGLEWMGNIYPGSGGTNYAEKFKNRVTMTRDTSISTAYM ELSRLRSDDTAVYYCARSGGPYFFDYWGQGTLVTVSS
TABLE-US-00008 TABLE 8 Amino Acid Sequence of h15-2b scFv .times. CD19 Fab Name Amino Acid Sequence SEQ ID NO Humanized DIQMTQSPSSLSASVGDRVTITCKASQDVNTSVAVVYQQKPGK 17 15-2b scFv APKLLIYWASTRHTGVPSRFSGSGSGTDFTFTISSLQPEDIATY YCLQYINYPYTFGQGTKLEIKRGGGGSEVQLVESGGGLVQPG GSLKLSCAASGFTFSNYWMNVVVRQASGKGLEVVVGEIRSKSN NYATHYAESVKGRFTISRDDSKNTAYLQMNSLKTEDTAVYYCT NRYYWGQGTLVTVSS G-MYC-(G GEQKLISEEDLGGGGSGGGGSGGGGSQL 14 4S)3 Linker hHB12b EVQLVESGGGLVQPGGSLRLSCAASGFTFSSSWMNVVVRQAP 18 (Anti-CD19) GKGLEVVVGRIYPGDGDTNYNGKFKGRFTISRDDSKNSLYLQM VH-CH1 NSLKTEDTAVYYCARSGFITTVLDFDYWGQGTLVTVSSASTKG PSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALT SGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPS NTKVDKRV hHB12b EIVLTQSPDFQSVTPKEKVTITCRASESVDTFGISFMNWFQQK 19 (Anti-CD19) PDQSPKLLIHAASNQGSGVPSRFSGSGSGTDFTLTINSLEAED VL-C.kappa. AATYYCQQSKEVPFTFGGGTKVEIKTVAAPSVFIFPPSDEQLK SGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQD SKDSTYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFN RGEC
TABLE-US-00009 TABLE 9 Amino Acid Sequence of h15-2b scFv .times. CD20 Fab Name Amino Acid Sequence SEQ ID NO Humanized DIQMTQSPSSLSASVGDRVTITCKASQDVNTSVAVVYQQKPGK 17 15-2b scFv APKLLIYWASTRHTGVPSRFSGSGSGTDFTFTISSLQPEDIATY YCLQYINYPYTFGQGTKLEIKRGGGGSEVQLVESGGGLVQPG GSLKLSCAASGFTFSNYWMNVVVRQASGKGLEVVVGEIRSKSN NYATHYAESVKGRFTISRDDSKNTAYLQMNSLKTEDTAVYYCT NRYYWGQGTLVTVSS G-MYC-(G GEQKLISEEDLGGGGSGGGGSGGGGSQL 14 4S)3 Linker 2F2 MELGLSWIFLLAILKGVQCEVQLVESGGGLVQPGRSLRLSCAA 20 (Anti-CD20) SGFTFNDYAMHVVVRQAPGKGLEVVVSTISWNSGSIGYADSVK VH-CH1 GRFTISRDNAKKSLYLQMNSLRAEDTALYYCAKDIQYGNYYYG MDVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCL VKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVT VPSSSLGTQTYICNVNHKPSNTKVDKRV 2F2 MEAPAQLLFLLLLWLPDTTGEIVLTQSPATLSLSPGERATLSCR 21 (Anti-CD20) ASQSVSSYLAVVYQQKPGQAPRLLIYDASNRATGIPARFSGSG VL-C.kappa. SGTDFTLTISSLEPEDFAVYYCQQRSNWPITFGQGTRLEIKTVA APSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNAL QSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKLYACEV THQGLSSPVTKSFNRGEC
TABLE-US-00010 TABLE 10 Amino Acid Sequence of 15-2b knob/Bu12 hole Name Amino Acid Sequence SEQ ID NO 15-2b knob heavy chain Humanized DIQMTQSPSSLSASVGDRVTITCKASQDVNTSVAWYQQKPGKA 12 15-2b PKLLIYWASTRHTGVPSRFSGSGSGTDFTFTISSLQPEDIATYYC VL-C.kappa. LQYINYPYTFGQGTKLEIKRTVAAPSVFIFPPSDEQLKSGTASVV CLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLS STLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized EVQLVESGGGLVQPGGSLKLSCAASGFTFSNYWMNWVRQASG 13 15-2b KGLEWVGEIRSKSNNYATHYAESVKGRFTISRDDSKNTAYLQM VH-CH1 NSLKTEDTAVYYCTNRYYWGQGTLVTVSSASTKGPSVFPLAPC SRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQ SSGLYSLSSVVTVPSSNFGTQTYTCNVDHKPSNTK Knob Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 22 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPCRDELTKNQVSLWCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK hBU12 hole hBU12 EIVLTQSPATLSLSPGERATLSCSASSSVSYMHWYQQKPGQAPR 23 VL-crossover LLIYDTSKLASGIPARFSGSGSGTDFTLTISSLEPEDVAVYYCFQ CH1 GSVYPFTFGQGTKLEIKRSSASTKGPSVFPLAPSSKSTSGGTAAL GCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSV VTVPSSSLGTQTYICNVNHKPSNTKVDKKV hBU12 QVQLQESGPGLVKPSQTLSLTCTVSGGSISTSGMGVGWIRQHPG 24 VH-crossover KGLEWIGHIWWDDDKRYNPALKSRVTISVDTSKNQFSLKLSSV C.kappa. TAADTAVYYCARMELWSYYFDYWGQGTLVTVSSASVAAPSV FIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGN SQESVTEQDSKDSTYSLSSTLTLSKADYEKHKLYACEVTHQGLS SPVTKSFNRGEC hole DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVD 25 hinge-CH2- VSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLT CH3 VLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVCT LPPSRDELTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKT TPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNH YTQKSLSLSPGK
TABLE-US-00011 TABLE 11 Amino Acid Sequence of 15-2b knob/anti-HER2 hole Name Amino Acid Sequence SEQ ID NO 15-2b knob heavy chain Humanized DIQMTQSPSSLSASVGDRVTITCKASQDVNTSVAWYQQKPGKA 12 15-2b PKLLIYWASTRHTGVPSRFSGSGSGTDFTFTISSLQPEDIATYYC VL-C.kappa. LQYINYPYTFGQGTKLEIKRTVAAPSVFIFPPSDEQLKSGTASVV CLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLS STLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC Humanized EVQLVESGGGLVQPGGSLKLSCAASGFTFSNYWMNVVVRQAS 13 15-2b GKGLEVVVGEIRSKSNNYATHYAESVKGRFTISRDDSKNTAYL VH-CH1 QMNSLKTEDTAVYYCSNRYYWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTF PAVLQSSGLYSLSSVVTVPSSNFGTQTYTCNVDHKPSNTKVD KTVERK Knob Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 22 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPCRDELTKNQVSLWCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK Anti-HER2 hole C6ML3-9V QSVLTQPPSVSAAPGQKVTISCSGSSSNIGNNYVSWYQQLPGTA 26 L-crossover PKLLIYDHTNRPAGVPDRFSGSKSGTSASLAISGFRSEDEADYY CH1 CASWDYTLSGWVFGGGTKLTVLGSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGL YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKV C6ML3-9 QVQLLQSGAEVKKPGESLKISCKGSGYSFTSYWIAWVRQMPGK 27 VH-crossover GLEYMGLIYPGDSDTKYSPSFQGQVTISVDKSVSTAYLQWSSLK C.kappa. PSDSAVYFCARHDVGYCSSSNCAKWPEYFQHWGQGTLVTVSS ASVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKV DNALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKLYA CEVTHQGLSSPVTKSFNRGEC hole DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVD 25 hinge-CH2- VSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLT CH3 VLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVCT LPPSRDELTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKT TPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNH YTQKSLSLSPGK
TABLE-US-00012 TABLE 12 Amino Acid Sequence of h6.3 knob/BU12 hole Name Amino Acid Sequence SEQ ID NO h6.3 knob heavy chain Humanized DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNQMNYLAWYQ 9 6.3 VL-C.kappa. QKPGQPPKLLIYWASTRESGVPDRFSGSGSGTDFTLTISSLQAED VAVYYCLQYLSSWTFGGGTKLEIKTYSLSSTLTLSKADYEKHK LYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGSELKKPGASVKVSCKASGYTFKNYGMNWVRQAP 10 6.3 GQGLEWMGWINTYTGQPIYANDFKGRFVFSLDTSVSTAYLQIS VH-CH1 SLKAEDTAVYYCARDWGPYWGQGTLVTVSSASTKGPSVFPLA PSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPA VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVE PKSCDK Knob Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 22 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPCRDELTKNQVSLWCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK BU12 hole hBU12 EIVLTQSPATLSLSPGERATLSCSASSSVSYMHWYQQKPGQAPR 23 VL-crossover LLIYDTSKLASGIPARFSGSGSGTDFTLTISSLEPEDVAVYYCFQ CH1 GSVYPFTFGQGTKLEIKRSSASTKGPSVFPLAPSSKSTSGGTAAL GCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSV VTVPSSSLGTQTYICNVNHKPSNTKVDKKV hBU12 QVQLQESGPGLVKPSQTLSLTCTVSGGSISTSGMGVGWIRQHPG 24 VH-crossover KGLEWIGHIWWDDDKRYNPALKSRVTISVDTSKNQFSLKLSSV C.kappa. TAADTAVYYCARMELWSYYFDYWGQGTLVTVSSASVAAPSV FIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGN SQESVTEQDSKDSTYSLSSTLTLSKADYEKHKLYACEVTHQGLS SPVTKSFNRGEC hole DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVD 25 hinge-CH2- VSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLT CH3 VLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVCT LPPSRDELTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKT TPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNH YTQKSLSLSPGK
TABLE-US-00013 TABLE 13 Amino Acid Sequence of h6.3 knob/anti-HER2 hole Name Amino Acid Sequence SEQ ID NO h6.3 knob heavy chain Humanized DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNQMNYLAWYQ 9 6.3 VL-C.kappa. QKPGQPPKLLIYWASTRESGVPDRFSGSGSGTDFTLTISSLQAED VAVYYCLQYLSSWTFGGGTKLEIKTYSLSSTLTLSKADYEKHK LYACEVTHQGLSSPVTKSFNRGEC Humanized QVQLVQSGSELKKPGASVKVSCKASGYTFKNYGMNWVRQAP 10 6.3 GQGLEWMGWINTYTGQPIYANDFKGRFVFSLDTSVSTAYLQIS VH-CH1 SLKAEDTAVYYCARDWGPYWGQGTLVTVSSASTKGPSVFPLA PSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPA VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVE PKSCDK Knob Hinge EPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT 22 CH2-CH3 CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPCRDELTKNQVSLWCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHE ALHNHYTQKSLSLSPGK Anti-HER2 hole C6ML3-9V QSVLTQPPSVSAAPGQKVTISCSGSSSNIGNNYVSWYQQLPGTA 26 L-crossover PKLLIYDHTNRPAGVPDRFSGSKSGTSASLAISGFRSEDEADYY CH1 CASWDYTLSGWVFGGGTKLTVLGSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGL YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKV C6ML3-9 QVQLLQSGAEVKKPGESLKISCKGSGYSFTSYWIAWVRQMPGK 27 VH-crossover GLEYMGLIYPGDSDTKYSPSFQGQVTISVDKSVSTAYLQWSSLK C.kappa. PSDSAVYFCARHDVGYCSSSNCAKWPEYFQHWGQGTLVTVSS ASVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKV DNALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKLYA CEVTHQGLSSPVTKSFNRGEC hole DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVD 25 hinge-CH2- VSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLT CH3 VLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVCT LPPSRDELTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKT TPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNH YTQKSLSLSPGK
[0131] 3. Pharmaceutical Kit
[0132] It is the second aspect of the present disclosure to provide a pharmaceutical kit for treating or imaging cancers, including metastatic and/or drug-resistant cancers. The pharmaceutical kit includes at least two components, the first component being the BsAbs of the present disclosure; and the second component being a PEGylated substance, which includes a cancer therapeutic agent (e.g., vinca alkaloid) or an imaging agent (e.g., a microbubble containing therein a contrast agent, or a quantum dot) inside the PEGylated substance. Typically, each component is contained in respective separate container. Preferably, the first and second components can be respectively present in the container in a dry solid form or as a suspension in a physiologically acceptable aqueous carrier. The kit may optionally include a physiologically acceptable aqueous carrier such as a saline for reconstitution of the dry components before injection. The two components will be reconstituted separately with the respective carriers, then mixed to form an assembly, which is administered to the subject (e.g., by injection).
[0133] 4. One-Step Method of Targeting and Treating Cancer
[0134] Accordingly, it is the third aspect of the present disclosure to provide a one-step method of targeting and treating cancers, including metastatic and/or drug-resistant cancers. The method takes advantages of the pharmaceutical kit described in Section 3, in which the isolated humanized anti-PEG bi-specific antibody (BsAbs) as described in Section 2 is mixed with a PEGylated substance to form an assembly before being administered to the subject. Alternatively, the humanized BsAbs as described in Section 2 is injected to the test subject (e.g., human) first, then followed by the injection of a PEGylated substance (data not shown).
[0135] FIG. 3 is a schematic drawing illustrating the one-step targeting and treating cancer by use of the pharmaceutical kit 300 of the present disclosure. The pharmaceutical kit 300 includes a humanized anti-PEG BsAb 310 and a PEGylated substance 320. The humanized anti-PEG BsAb 310 is composed of a first antigen binding site 311 that selectively binds to PEGylated substance 320, which contains a cancer therapeutic agent within its structure; and a second antigen binding site 312 that selectively binds to a target protein, such as a tumor antigen 340. In this embodiment, the PEGylated substance 320 is depicted as a liposome or micelle, and is characterized in having a cancer therapeutic agent 321 within the liposome or micelle structure and a plurality of PEG molecules 322 extended from the surface of the liposome or micelle. Upon binding to the PEGylated substance 320, the first antigen binding site 311 of the BsAb 310 allows the BsAb 310 to orient the second antigen binding site 312 outward from the surface of the PEGylated substance 320, thereby converting the non-targeted PEGylated substance 320 to tumor cell-targeted PEGylated substance.
[0136] In practice, to achieve one-step targeting and treating purpose, the humanized anti-PEG BsAb 310 is mixed with the PEGylated substance 320 to form an assembly 330, the assembly 330 is then immediately administered (e.g., injection) to a subject (e.g., a human 360 as depicted in FIG. 3).
[0137] Accordingly, it is the third aspect of the present disclosure to provide a method of treating cancers. The method includes the step of, administering to the subject, the BsAb of the present disclosure and a PEGylated substance containing a cancer therapeutic agent therein, in a dose sufficient to inhibit the growth or metastasis of the cancer of the subject. The dose administered to the subject is from about 0.1 to 50 mg/Kg body weight of the subject. In certain embodiments, the dose is administered to the subject from about 1 to 40 mg/Kg body weight of the subject, such as 10 to 30 mg/Kg body weight of the subject. The dose can be administered in a single dose, or alternatively in more than one smaller doses.
[0138] The BsAb of the present disclosure and the PEGylated substance may be administered to a mammal, preferably human, by any route that may effectively transports the cancer therapeutic agent contained in the PEGylated substance to the appropriate or desired site of action, such as oral, nasal, pulmonary, transdermal, such as passive or iontophoretic delivery, or parenteral, e.g., rectal, depot, subcutaneous, intravenous, intramuscular, intranasal, intra-cerebella, ophthalmic solution or an ointment. It will be appreciated that the dosage of the present disclosure will vary from patient to patient not only for the cancer therapeutic agent selected, the route of administration, and the ability of the BsAb in combination with a PEGylated substance, to elicit a desired response in the patient, but also factors such as disease state or severity of the condition to be alleviated, age, sex, weight of the patient, the state of being of the patient, and the severity of the pathological condition being treated, concurrent medication or special diets then being followed by the patient, and other factors which those skilled in the art will recognize, with the appropriate dosage ultimately being at the discretion of the attendant physician. Dosage regimens may be adjusted to provide the improved therapeutic response. A therapeutically effective amount is also one in which any toxic or detrimental effects of the cancer therapeutic agent are outweighed by the therapeutically beneficial effects. Preferably, the BsAb and the PEGylated substance of the present disclosure are administered at a dosage and for a time such that the number and/or severity of the symptoms are decreased.
[0139] 5. Method of Imaging a Targeted Tissues
[0140] The fourth aspect of the present disclosure is to provide a method of imaging tissues, particularly the cancerous tissues, of a live subject. The method also takes advantages of the pharmaceutical kit described in Section 3, in which the isolated humanized anti-PEG bi-specific antibody (BsAbs) as described in Section 2 is mixed with a PEGylated substance to form an assembly before being administered to the subject.
[0141] The method includes the steps of, (a) mixing a first sufficient amount of the humanized BsAb of the present disclosure and a second sufficient amount of a PEGylated quantum dot (PEG-QD) or a PEGylated liposome containing a fluorescent dye, to form an assembly; (b) injecting the assembly of the step (a) to a body portion of the subject; and (c) imaging the body portion of the subject by fluorescence imaging, electron spin resonance (ESR) imaging, X-ray imaging, computed tomography (CT), or magnetic resonance imaging (MRI). The PEG-QD includes a quantum dot nanocrystal selected from the group consisting of CdHgTe, CdSe, CdSe/ZnS, CdS, CdTe, CdTe/CdS, PbSe and PbS.
[0142] The 6.3 antibody comprises the sequence of VL-C.kappa. domain (SEQ ID NO: 9) and the sequence of VH-CH1 domain (SEQ ID NO: 10), wherein the sequence of VL-C.kappa. domain comprises a CDR1 having the sequence of SEQ ID NO: 216; a CDR2 having the sequence of Trp-Ala-Ser; and a CDR3 having the sequence of SEQ ID NO: 217, wherein the sequence of VH-CH1 domain comprises a CDR1 having the sequence of SEQ ID NO: 218; a CDR2 having the sequence of SEQ ID NO: 219; and a CDR3 having the sequence of SEQ ID NO: 220.
[0143] The h15-2b antibody comprises the sequence of VL-C.kappa. domain (SEQ ID NO: 12) and the sequence of VH-CH1 domain (SEQ ID NO: 13), wherein the sequence of VL-C.kappa. domain comprises a CDR1 having the sequence of SEQ ID NO: 221; a CDR2 having the sequence of SEQ ID NO: 222; and a CDR3 having the sequence of SEQ ID NO: 223, wherein the sequence of VH-CH1 domain comprises a CDR1 having the sequence of SEQ ID NO: 224; a CDR2 having the sequence of SEQ ID NO: 225; and a CDR3 having the sequence of SEQ ID NO: 226.
[0144] The present invention further provides a humanized bi-specific antibody against the backbone of polyethylene glycol (PEG) and a target ligand, comprising, a first antigen binding site that binds to the PEG, wherein the first antigen binding site comprises a first VL-C.kappa. domain and a first VH-CH1 domain; and a second antigen binding site that binds to the target ligand, which is EGFR, wherein the first VL-C.kappa. domain comprises a CDR1 having the sequence at least 90% identical to SEQ ID NO: 216; a CDR2 having the sequence at least 90% identical to Trp-Ala-Ser; and a CDR3 having the sequence at least 90% identical to SEQ ID NO: 217; the first VH-CH1 domain comprises a CDR1 having the sequence at least 90% identical to SEQ ID NO: 218; a CDR2 having the sequence at least 90% identical to SEQ ID NO: 219; and a CDR3 having the sequence at least 90% identical to SEQ ID NO: 220; and the second antigen binding site comprises a single chain variable fragment (scFv) at least 90% identical to SEQ ID NO:7.
[0145] In another embodiment, the first VL-C.kappa. domain has the sequence at least 90% identical to SEQ ID NO: 9, and the first VH-CH1 domain has the sequence at least 90% identical to SEQ ID NO: 10.
[0146] In one embodiment, the first antigen binding site further comprises a first HR-CH2-CH3 domain at least 90% identical to SEQ ID NO: 3 disposed between the first VH-CH1 domain and the scFv.
[0147] The present invention will now be described more specifically with reference to the following embodiments, which are provided for the purpose of demonstration rather than limitation.
EXAMPLES
[0148] Materials and Methods.
[0149] Cells and Animals
[0150] Breast cancer cell line MCF-7, human T lymphocyte cell line Jurkat, ovarian cancer cell line OVCAR-3, epidermoid carcinoma cell line A431 (EGFR.sup.+) (ATCC CRL1555), B lymphocyte cell line Raji (CD19+) (ATCC CCL86), malignant melanoma cell line A-375, 293FT cells, B-lymphoblastoid cell line Ramos (CD19.sup.+) (ATCC CRL-1596), SW480 (EGFR.sup.+), SW620 human colon carcinoma cells, human breast adenocarcinoma cell line SKBR3 (HER2.sup.+), human breast adenocarcinoma cell line MDA-MB-468, BALB 3T3 cells, and GP2-293 retrovirus packaging cells were used in the present disclosure. In general, cells were cultured in Dulbecco's modified Eagle's medium (Sigma, St Louis, Mo., USA) supplemented with 10% fetal calf serum (HyClone, Logan, Utah), 100 U/mL penicillin and 100 .mu.g/mL streptomycin at 37.degree. C. in an atmosphere of 5% CO.sub.2 in air. A431, Raji and Ramos cells were grown in RPMI-1640 containing the same supplements but with 10% bovine serum source.
[0151] Female BALB/c nude mice (6-8 weeks old) were obtained from the National Laboratory Animal Center, Taipei, Taiwan. All animal experiments were performed in accordance with institutional guidelines and approved by the Laboratory Animal Facility and Pathology Core Committee of IBMS, Academia Sinica.
[0152] Generating Murine Anti-PEG Antibodies (Abs)
[0153] Hybridoma cells secreting anti-PEG Ab were generated by immunizing female BALB/c mice with mPEG-derived proteins or PEG-derived proteins as described previously (Su et al., Bioconjugate Chemistry (2010) 21(7), 1264-1270.). The hybridomas were then screened by ELISA. Specifically, 96-well plates were coated with 1 .mu.g/well CH3-PEG.sub.750-NH2, NH2-PEG.sub.3000-NH2 (Sigma-Aldrich), or CH3-PEG.sub.5000-NH2 in 5 .mu.L/well 0.1M NaHCO.sub.3/Na.sub.2CO.sub.3 for 3 hr at 37.degree. C. and then blocked with 200 .mu.L/well dilution buffer (2% skim mile in PBS) at 4.degree. C. overnight. Graded concentrations of antibodies in 50 .mu.L 2% skim milk were added to plates at room temperature for 1 hr. The plates were washed with PBS-T (PBS containing 0.05% Tween-20) 3 times and with PBS 2 times. HRP-conjugated goat anti-mouse IgMpchain (2 .mu.g/mL) or HRP-conjugated donkey antimouse IgG Fc (2 .mu.g/mL) in 50 .mu.L dilution buffer were added for 1 hr. The plates were washed and peroxidase activity was measured by adding 100 .mu.L/well TMB substrate solution (BioLegend, San Diego, Calif.) for 30 min at room temperature. After adding stop buffer (2N H2504, 50 .mu.L/well), the absorbance (405 nm) were read. Selected hybridomas were cloned three times by limiting dilution in 96-well plates containing thymocyte feeder cells in HT medium (Sigma-Aldrich) supplemented with 15% fetal calf serum (Hyclone), and then three hybridoma cells, E11, 6-3, and 15-2b were produced, in which E11 and 6-3 secreted anti-PEG backbone Abs, whereas 15-2b secreted anti-mPEG Abs.
[0154] Construction of DNA plasmids for PEG2.times.EGFR, PEG2.times.TAG72, PEG2.times.HER2, h6.3Fab.times.EGFR, and h6.3Fab.times.CD19
[0155] To generate the anti-PEG Fab or IgG based BsAbs, the mouse VL and VH domains of the anti-PEG antibodies were cloned from cDNA respectively prepared from the E11, 6-3, and 15-2b hybridoma cells. The humanized VL and VH domains of the anti-PEG (hE11, h6-3) and anti-mPEG (h15-2b) antibodies, generated by grafting the DNA sequences coding the complementarity-determining regions (CDRs) of the light and heavy chain variable region genes of E11, h6-3 and 15-25 into the framework regions of human IgGVL and VH genes, then were fused with the DNA sequence encoding the remaining human IgG1 constant region genes.
[0156] The C.kappa., CH1, and partial CH.sub.1-hinge-CH.sub.2-CH.sub.3 (Fc), constant domains of human IgG.sub.1 were cloned from extracted human PBMC cDNA by using the primers in Table 14.
TABLE-US-00014 TABLE 14 Primers For Cloning Human CH.sub.1, C.kappa. and F.sub.c Fragments SEQ Name Sequence (5'-3') ID NO CH.sub.1 ctggtcaccgtctcctcagcctccaccaagggaccatcg 28 (forward) gtcgactttgtcacaagatttgggc 29 (reverse) C.kappa. accaaggtggagatcaaacggactgtggctgcaccatct 30 (forward) ctcgaggcactctcccctgttgaagc 31 (reverse) F.sub.c ggtggacaagagagttgagcccaaatcttgtgac 32 (forward) caattgtccactgccacccccgcttga 33 (reverse)
[0157] The humanized variable domains (V.sub.L or V.sub.H) and human antibody constant domains (C.kappa., CH.sub.1, or Hinge-CH.sub.2--CH.sub.3) were joined by overlap PCR. To this aim, all the humanized V.sub.L fragments were amplified by PCR to introduce a partial C.kappa. fragment at the C-terminus using primers in Table 15.
TABLE-US-00015 TABLE 15 Primers For Cloning the V.sub.L Domains of humanized E11, 6-3, 15-2b anti-PEG fragments SEQ Name Sequence (5'-3') ID NO hE11V.sub.L ggcccagccggccgatgttgtgatg 34 (forward) actcagtc hE11V.sub.L-partial gtgcagccacagtccgtttgatctc 35 C.kappa. caccttggtc (reverse) h6-3V.sub.L(forward) ggcccagccggcc gacatcgtgat 36 gacccag h6-3V.sub.L-partial C.kappa. gtgcagccacagtccgtttgatttc 37 (reverse) caccttggtc h15-2bV.sub.L ggcccagccggccgacatccagatg 38 (forward) acccag h15-2bV.sub.L-partial gtgcagccacagtccgtttgatctc 39 C.kappa. cagcttggtc (reverse)
[0158] The VL-partial C.kappa. and C.kappa. fragments were again joined by overlap PCR to generate V.sub.L-C.kappa. fragments using the forward primers of V.sub.L domains and the C.kappa. reverse primer as shown above in Tables 14 and 15.
[0159] Likewise, all the humanized VH fragments were amplified by PCR to introducing partial CH1 fragment at the C-terminus using primers shown in Table 16.
TABLE-US-00016 TABLE 16 Primers For Cloning the V.sub.H Domains of humanized E11, 6-3, 15-2b anti-PEG Fragments SEQ Name Sequence (5'-3') ID NO hE11V.sub.H agatctcaggtgcagctggtgcag 40 (forward) hE11V.sub.H-partial tcccttggtggaggctgaggagacg 41 CH.sub.1 gtgaccaggg (reverse) h6-3V.sub.LH agatctcaggtgcagctggtgcaatc 42 (forward) h6-3V.sub.H-partial CH.sub.1 gtcccttggtggaggctgaggagac 43 (reverse) ggtgaccag h15-2bV.sub.H agatctgaggtgcagctggtggag 44 (forward) h15-2bV.sub.H-partial gcccttggtggaggctgaggagacg 45 CH.sub.1 gtgaccaggg (reverse)
[0160] The V.sub.H-partial CH.sub.1, CH.sub.1 and Fc fragments were joined by overlap PCR to generate V.sub.H--CH.sub.1 or V.sub.H--CH.sub.1-hinge-CH.sub.2--CH.sub.3 fragments using the forward primers of VH domains, and the CH.sub.1 and Fc reverse primers as indicated above in Tables 14 and 16.
[0161] The hBU12 dsFv DNA fragment was synthesized by assembly PCR based on the VH and V.sub.L sequences of hBU12 described in U.S. Pat. No. 7,968,68762, the entirety of which is incorporated herein by reference. PCR was carried out as follows: 95.degree. C. for 3 min; 10 cycles at 95.degree. C. for 30 s, 63 to 53.degree. C. touchdown for 1 min (decrease 1.degree. C. every cycles), 72.degree. C. for 1 min; 25 cycles at 95.degree. C. for 30 s, 53.degree. C. for 1 min, 72.degree. C. for 1 min; 72.degree. C. for 10 min. The VH and V.sub.L fragments were joined and amplified using P1 and P22 primers described in Table 17. The 11F8 dsFv DNA fragment was synthesized by assembly PCR based on the V.sub.H and V.sub.L sequences of 11F8 described in EP2332990 A1, the entirety of which is incorporated herein by reference. PCR was carried out as described above. The VH and VL fragments were assembled by assembly PCR using primers P23 to P34 and primers P35 to P44, respectively described in Table 18. The hCC49 scFv and DNS scFv DNA fragments were amplified by PCR using the primers as described in our previous studies (K C Chen et al., Bioconjugate Chemistry 22: 938-948, 2011). The production of C6ML3-9 dsFv plasmid was described in EP2258726A1. We next generated SalI-linker-MfeI-hBU12 scdsFv-MluI-6.times.His-ClaI by PCR using primers set forth in Table 19; whereas MfeI-linker-11F8 dsFv-MluI, MfeI-linker-DNS dsFv-MluI and MfeI-linker-hCC49 dsFv-MluI are generated by PCR using primers as described in Table 20.
TABLE-US-00017 TABLE 17 Primers For Cloning hBU12 dsFv Primer SEQ ID name Primer Sequence (5'-xxxxxxx-3') NO P1 CAATTGCAGGTTCAGCTGCAAGAGTCTGGCCCTGGGTTGGTTAAGCCC 46 P2 CAGTACAAGTCAGACTGAGGGTCTGGGAGGGCTTAACCAACCCAGGGCC 47 P3 CAGTCTGACTTGTACTGTGTCTGGGGGTTCAATCAGCACTTCTGGTATG 48 P4 CTGGGTGCTGCCTAATCCAGCCTACACCCATACCAGAAGTGCTGATTG 49 P5 GGATTAGGCAGCACCCAGGGAAGTGTCTGGAGTGGATTGGACACATTTGG 50 P6 AACAGTAATAGACAGCAACATCCTCTGGCTCCAGGCTGCTGATTGTG 51 P7 CAAGAGATATAACCCAGCCCTGAAGAGCAGAGTGACAATCTCTGTGGATAC 52 P8 GACAGCTTGAGGCTAAACTGGTTCTTGGAGGTATCCACAGAGATTGTCAC 53 P9 GTTTAGCCTCAAGCTGTCCAGTGTGACAGCTGCAGATACTGCTGTCTAC 54 P10 AAACAGTAATAGACAGCAACATCCTCTGGCTCCAGGCTGCTGATTGTG 55 P11 GGAACTTTGGTCCTACTATTTTGACTACTGGGGCCAAGGCACCCTTG 56 P12 GCCCCCTGACCCGCCACCTCCTGAGGAGACTGTGACAAGGGTGCCTTGGCCCC 57 P13 GGTGGATCGGGGGGTGGCGGATCTGAAATTGTTCTCACCCAGTCTCCAGCAAC 58 P14 CAGGGTAGCCCTTTCCCCTGGAGAGAGAGACAGGGTTGCTGGAGACTGGGTG 59 P15 GGGGAAAGGGCTACCCTGAGCTGCAGTGCCAGCTCAAGTGTAAGTTACATGC 60 P16 CTGGGAGCCTGCCCTGGCTTCTGCTGGTACCAGTGCATGTAACTTACACTTG 61 P17 GCCAGGGCAGGCTCCCAGACTCCTGATTTATGACACATCCAAACTGGCTTC 62 P18 CCAGACCCACTGCCACTGAACCTTGCTGGAATACCAGAAGCCAGTTTGGATG 63 P19 CAGTGGCAGTGGGTCTGGAACAGATTTTACACTCACAATCAGCAGCCTGG 64 P20 GAAAACAGTAATAGACAGCAACATCCTCTGGCTCCAGGCTGCTGATTGTG 65 P21 GCTGTCTATTACTGTTTTCAGGGGAGTGTATACCCATTCACTTTTGGC 66 P22 ACGCGTTCTTTTGATTTCCAACTTTGTCCCGCAGCCAAAAGTGAATGGG 67
TABLE-US-00018 TABLE 18 Primers For Cloning 11F8 dsFv SEQ ID Primer name primer sequence (5'-xxxxxxx-3') NO P23 GTCGACCAATTGGGAGGTGGCGGATCCCAGGTGCAGCTGCAGGAGTCGGG 68 P24 ACAGGGTCTGTGAAGGCTTCACCAGTCCTGGGCCCGACTCCTGCAGCTGC 69 P25 AGCCTTCACAGACCCTGTCCCTCACCTGCACTGTCTCTGGTGGCTCCATCAG 70 P26 GGCGGATCCAACTCCAGTAGTAATCACCACTGCTGATGGAGCCACCAGAGAC 71 P27 ACTGGAGTTGGATCCGCCAGCCCCCAGGGAAGTGCCTGGAGTGGATTGGG 72 P28 GGTTGTAGTCGGTGCTCCCACTGTAATAGATGTACCCAATCCACTCCAGGCA 73 P29 TGGGAGCACCGACTACAACCCGTCCCTCAAGAGTCGAGTCACCATGTCCGTA 74 P30 ACCTTCAGGGAAAACTGATTCTTGGACGTGTCTACGGACATGGTGACTCG 75 P31 TCAGTTTTCCCTGAAGGTCAACTCTGTGACCGCCGCAGACACGGCTGTGT 76 P32 CCCCACTCCAAAAATCGACACTCTCGCACAGTAATACACAGCCGTGTCTGCGG 77 P33 TCGATTTTTGGAGTGGGGACATTTGACTACTGGGGCCAGGGCACCCTGGT 78 P34 ACCGCCCCCTGACCCGCCACCTCCGCTTGAGACGGTGACCAGGGTGCCCTGGCC 79 P35 GGATCGGGGGGTGGCGGATCTGAAATTGTGATGACACAGTCTCCAGCCACCCTGTC 80 P36 GCAGGAGAGGGTGGCTCTTTCCCCTGGAGACAAAGACAGGGTGGCTGGAGAC 81 P37 AGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCTACTTA 82 P38 AGCCTGGCCAGGTTTCTGTTGGTACCAGGCTAAGTAGCTGCTAACACT 83 P39 CAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCATCCAACAG 84 P40 ACTGCCACTGAACCTGGCTGGGATGCCAGTGGCCCTGTTGGATGCATCATAG 85 P41 GCCAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACTCTCACCATCAG 86 P42 AATACACTGCAAAATCTTCAGGCTCTAGGCTGCTGATGGTGAGAGTGAAG 87 P43 GAAGATTTTGCAGTGTATTACTGTCACCAGTATGGTAGCACACCTCTCACTT 88 P44 ACGCGTTTTGATCTCCGCCTTGGTCCCGCAGCCGAAAGTGAGAGGTGTGCTA 89
TABLE-US-00019 TABLE 19 Primers For Cloning Sall-linker-Mfel-hBU12 scFv-Mlul-6xHis-Clal SEQ ID Name Sequence (5'-3') NO linker-Mfel-hBU12 gtggtggttcaggacaattgggagg 90 V.sub.H (forward) tggcggatcccaggttcagctgcaa gag Sall-linker- gtcgacctggtcaccgtctcctcag 91 Mfel-hBU12 cctccaccggtggtggttcaggaca V.sub.H (forward) at hBU12V.sub.L-Mlul- atcgatttaatgatgatgatgatga 92 6xHis-Clal tgacgcgttatttgatttccaactt (reverse) tg
TABLE-US-00020 TABLE 20 Primers For Cloning Mfel-linker-11F8 dsFv-Mlul, Mfel-linker-DNS dsFv-Mlul and Mfel-linker-hCC49 dsFv-Mlul SEQ ID Name Sequence (5'-3') NO Mfel-hl1F8 V.sub.H caattgggaggtggcggatcccaggtgcagc 93 (forward) tgcaggag Mlul-11F8V.sub.L acgcgttttgatctccgccttggtc 94 (reverse) Mfel-DNSV.sub.H caattgggaggtggcggatccagtgaagtga 95 (forward) agcttgag Mlul-DNSV.sub.L acgcgtccgttttatttccaactt 96 (reverse) Mfel-hCC49V.sub.H caattgggaggtggcggatcccaggtgcagc 97 (forward) tggtgcag Mlul-hCC49V.sub.L acgcgttttgatctccaccttggtc 98 (reverse)
[0162] The pLNCX-SfiI-mAGP3 V.sub.L-C.kappa.-XhoI-F2A-BglII-mAGP3 VH-CH.sub.1-SalI-eB7-ClaI plasmid was used as template for further sub-cloning. These SfiI-V.sub.L-C.kappa.-XhoI or BglII-VH-CH.sub.1-SalI or BglII-VH-CH.sub.1-hinge-CH2-CH3-SalI fragments were sub-cloned into the template DNA plasmid described above by digesting with proper restriction enzyme to generate pLNCX-SfiI-anti-PEG V.sub.L-C.kappa.-XhoI-F2A-BglII-anti-PEG V.sub.H-CH.sub.1-SalI-eB7-ClaI orpLNCX-SfiI-anti-PEGVL-C.kappa.-XhoI-F2A-BglII-anti-PEGVH-CH.sub.1-hinge- -CH.sub.2-CH.sub.3-SalI-eB7-ClaI plasmids. Furthermore, the single chain variable fragments (scFv or scdsFv) were sub-cloned into these plasmids by using SalI and ClaI or followed by MfeI and ClaI enzyme digestion.
[0163] Construction of DNA plasmids for 15-2b knob/Bu12 hole, 15-2bknob/anti-HER2 hole, h6.3 knob/Bu12 hole, and h6.3 knob/anti-HER2 hole
[0164] To construct the knob into hole BsAbs, V.sub.H--CH.sub.1 of h15-2b or h6-3 was fused with the modified human IgG.sub.1 CH.sub.2--CH.sub.3 domain to form the heavy chain of h15-2b-knob or h6-3-knob by assembly PCR. The heavy chain sequence, which is followed by a sequence derived from furin-2A (F2A) and the light chain sequence of h15-2b or h6-3, was cloned into lentival vector pLKOAS3W-hyg (RNAi core, Academia Sinica, Taipei, Taiwan) by use of NheI and PmeI restriction sites. We adopted the immunoglobulin domain crossover approach, alone with modifications of the locations of the C.sub.H1 and hinge regions, to generate BU12-hole and .alpha.-Her2-hole. In brief, V.sub.H-partialCH.sub.1-C.kappa.-partial hinge, with a human influenza virus hemagglutinin (HA) tag protein fused at the N terminal of the heavy chain was fused with human IgG1 CH.sub.2--CH.sub.3 domain to form the new heavy chain. V.sub.L-CH.sub.1-partial hinge sequences of .alpha.-Her2 Ab or BU12 were connected with the new heavy chain sequences by a F2A sequence and cloned into lentiviral vector pLKOAS3W-pur (RNAi core, Academia Sinica, Taipei, Taiwan) by use of SfiI and PmeI restriction sites. Primers used for cloning were as described in Table 21.
TABLE-US-00021 TABLE 21 Primers For Cloning knob into hole BsAbs SEQ ID Name Sequence (5'-3') NO 15-2b-knob: V.sub.L-C.sub..kappa. agatctgacatccagatgacccag 99 (forward) tatcgatgtttaaacctagcactctcccctgttgaa 100 (reverse) V.sub.H-CH.sub.1 ggcccagccggccgaggtgcagctggtggag 101 (forward) aagttttttgtcgaccgtgg 102 (reverse) hulgG1-upper hinge gacaaaactcacacatgcccaccgtgc 103 (forward) CH.sub.1-partical hinge gcatgtgtgagttttgtcacaagatttgggctcaac 104 (reverse) CH.sub.3 ctcgagtttacccggagacaggga 105 (reverse) h6-3-knob: 104 V.sub.L-C.sub..kappa. agatctgacatcgtgatgacccagtctc 106 (forward) tatcgatgtttaaacctagcactctcccctgttgaa 107 (reverse) V.sub.H-CH.sub.1 caggtgcagctggtgcaat 108 (forward) aactctcttgtccaccttgg 109 (reverse) BU12-hole: V.sub.H-C.sub..kappa.-partial hinge agccggcccaggttcagctgcaagagtctggc 110 (forward) gcatgtgtgagttttgtcacactctcccctgttgaagct 111 (reverse) V.sub.L -partial gaaattgttctcacccagtctcc 112 V.sub.H-CH.sub.1-upperHinge (forward) ttaacaagatttgggctcaac 113 (reverse) partial V.sub.L-hCH.sub.1 gttggaaatcaaaagatcctcagcctccaccaagggcccatcg 114 (forward) .alpha.-Her2-hole: V.sub.H ggcccagccggcccaggtgcagctgttgcagtctggg 115 (forward) V.sub.H-C.sub..kappa.-partial hinge aggtgcagctgttgcagtctggg 116 (forward) gcatgtgtgagttttgtcacactctcccctgttgaagct 117 (reverse) V.sub.L-partial V.sub.H ctgaccgtcctaggttcctcagcctccaccaagggcccatcg 118 (forward) acctaggacggtcagcttggtcccgccgccgaacacccagcccga 119 (reverse) V.sub.L ctgccagatctcagtctgtgttgacgcag 120 (forward) acctaggacggtcagcttggtcccgccgccgaacacccagcccga 121 (reverse) CH.sub.1-upper hinge ttaacaagatttgggctcaac 122 (reverse)
[0165] Constructing DNA Plasmids for 15-2b Fab.times.HER2 scFv, 15-2b Fab.times.EGFR scFv, 15-2b scFvxCD19 Fab, and 15-2b scFvxCD20 Fab
[0166] The V.sub.L-C.kappa. and V.sub.H--C.sub.H1 domains of the anti-mPEG antibody were cloned from the cDNA of the 15-2b hybridoma and humanized as described previously (Chuang K-H et al., J. Nucl. Med. 2010 (51): 933-941). The humanized anti-mPEG VL and VH segments were synthesized by assembly polymerase chain reaction (PCR) and were subcloned into retroviral vector pLNCX-anti-PEG-eB7) in the unique BgIII, SalI, SfiI, and XhoI restriction enzyme sites, respectively. Primers for cloning 15-2b Fab sequence are given in Table 22. The human anti-EGFR scFv was cloned based on the h528Fv DNA sequence (Makabe et al., (2008) J. Biol. Chem, 283, 1156-1166.). Therefore, h528Fv DNA sequence was generated by assembly PCR. Primers used in the cloning of anti-EGFR V.sub.H and V.sub.L are given in Table 23. Then, using Mfe-ahEGFR V.sub.L primer, ahEGFR V.sub.L-(G4S)2 primer, (G4S)2-ahEGFR V.sub.H primer and ahEGFR V.sub.H-stop-Cal primer to create human anti-EGFR scFv, which contained a myc tag and fifteen amino acid (GGGGS)3 flexible linker in front of the sequence.
[0167] A human anti-HER2 scFv and anti-DNS scFv were cloned from the pBub-YCMC plasmid (Shahied et al., (2004) J. Biol. Chem. 279, 53907-53914) and pLNCX-DNS-B7 (Chuang et al., (2006) Bioconjugate Chemistry 17, 707-714), respectively. Primers for cloning human anti-HER2 scFv and anti-DNS scFv are given in Tables 24 and 25, respectively. A myc tag and fifteen amino acid (GGGGS)3 flexible linker was placed between the anti-mPEG Fab and scFv genes to generate pLNCX-PEG.times.EGFR, pLNCX-PEGxHER2 and pLNCX-PEGxDNS plasmids by using SalI and Call restriction enzyme sites.
TABLE-US-00022 TABLE 22 Primers For Cloning h15-2b Fab Sequence SEQ ID Name of Primer Sequence NO h15-2b VH-CH1 h15-2b Bgl-VH-1 5'-gaagatctgaggtgcagctggtggagtctgggggaggcttggtccag-3' 123 (forward) h15-2b VH-2 5'-agaggctgcacaggagagtttcagggaccccccaggctggaccaagcctcc-3' 124 (reverse) h15-2b VH-3 5'-tcctgtgcagcctctgggttcaccttcagtaactactggatgaactgggtc-3' 125 (forward) h15-2b VH-4 5'-gccaacccactccagccctttcccggaagcctggcggacccagttcatcca-3' 126 (reverse) h15-2b VH-5 5'-ctggagtgggttggcgaaattagatcgaaatctaataattatgcgacacat-3' 127 (forward) h15-2b VH-6 5'-ggagatggtgaacctccctttcacagactccgcataatgtgtcgcataatt-3' 128 (reverse) h15-2b VH-7 5'-aggttcaccatctccagagatgattcaaagaacacggcgtatctgcaaatg-3' 129 (forward) h15-2b VH-8 5'-gtaatacacggccgtgtcctcggttttcaggctgttcatttgcagatacgc-3' 130 (reverse) h15-2b VH-9 5'-acggccgtgtattactgttccaacagatactactggggccaaggaaccctg-3' 131 (T93S) (forward) h15-2b VH-10 5'-acctttggtggaggctgaggagacggtgaccagggttccttggcc-3' 132 (reverse) homo 3' IgG2 5'-acgcgtcgactttgcgctcaactgtctt-3' 133 CH1-Sall (reverse) h15-2b VL-C.kappa. h15-2b sfi-VL-1 5'-tgctggggcccagccggccgacatccagatgacccagtctcca-3' 134 (forward) h15-2b VL-2 5'-ggtgactctgtctcctacagatgcagacagggaggatggagactgggtcat-3' 135 (reverse) h15-2b VL-3 5'-ggagacagagtcaccatcacttgcaaggccagtcaggatgtaaatacttct-3' 136 (forward) h15-2b VL-4 5'-aggggctttccctggtttctgctgataccaggctacagaagtatttacatc-3' 137 (reverse) h15-2b VL-5 5'-ccagggaaagcccctaagctcctgatctactgggcatccacccggcacact-3' 138 (forward) h15-2b VL-6 5'-cccagatccacttccactgaaccttgatgggaccccagtgtgccgggtgga-3' 139 (reverse) h15-2b VL-7 5'-ggaagtggatctgggacagattttactttcaccatcagcagcctgcagcct-3' 140 (forward) h15-2b VL-8 5'-gatatattgcagacagtaatatgttgcaatatcttcaggctgcaggctgct-3' 141 (reverse) h15-2b VL-9 5'-tgtctgcaatatatcaactatccgtacacgtttggccaggggaccaagctg-3' 142 (forward) h15-2b VL-10 5'-tggtgcagccacagtccgtttgatctccagcttggtcccctg-3' 143 (reverse) homo 3' 5'-ccgctcgaggcactctcccctgttgaagctctttgtgacgggcgagctcaggccctg-3' 144 C.kappa.cys-Xhol (reverse)
TABLE-US-00023 TABLE 23 Primers for the cloning of h528 (Anti-EGFR) scFv SEQ ID Name of Primer Sequence NO h528VH01 5'-caggtgcaactggttcagagcggcgcggaagtgaaaaagccgggcgcgtcggtt-3' 145 (forward) h528VH02 5'-aaaggtatagcctgaggctttgcagctcactttaaccgacgcgcccgg-3' 146 (reverse) h528VH03 5'-tcaggctatacctttacgagctactggatgcattgggtgcgccaggcc-3' 147 (forward) h528VH04 5'-aatgttacccatccattccaggccctgacccggggcctggcgcaccca-3' 148 (reverse) h528VH05 5'-tggatgggtaacatttatccgggcagcggtggcaccaactatgcggaa-3' 149 (forward) h528VH06 5'-atcacgcgtcatggtcacgcggttcttaaatttttccgcatagttggt-3' 150 (reverse) h528VH07 5'-accatgacgcgtgataccagcatttcgacggcctatatggaactgagc-3' 151 (forward) h528VH08 5'-gtaatacacggcggtgtcatcgctacgcaggcggctcagttccatata-3' 152 (reverse) h528VH09 5'-accgccgtgtattactgcgcgcgcagtggcggtccgtattttttcgat-3' 153 (forward) h528VH10 5'-cgagctcacggtaaccagcgtaccctggccccagtaatcgaaaaaatacgg-3' 154 (reverse) (G4S)2-ahEGFR 5'-ggcggtggtgggtcgggtggcggcggatctcaggtgcaactggtt-3' 155 VH (forward) ahEGFR 5'-ccatcgatttacgagctcacggtaac-3' 156 VH-stop-Cal (reverse) h528VL01 5'-gatattgtgatgacccagagcccgctgagcctgccggtgaccccaggc-3' 157 (forward) h528VL02 5'-ctgcgagctgcggcagctaatcgacgccggttcgcctggggtcaccgg-3' 158 (reverse) h528VL03 5'-tgccgcagctcgcagaacatcgtgcataataacggcattacctatctg-3' 159 (forward) h528VL04 5'-cgggctttggcccggtttctgcagataccattccagataggtaatgcc-3' 160 (reverse) h528VL05 5'-ccgggccaaagcccgcagctgttaatttataaagtgagcgatcgcttt-3' 161 (forward) h528VL06 5'-accgctgcccgaaaagcgatccggcacgccgctaaagcgatcgctcac-3' 162 (reverse) h528VL07 5'-ttttcgggcagcggtagtggcaccgattttacgctgaaaattagccgc-3' 163 (forward) h528VL08 5'-gcagtaatacacgccaacatcctccgcttccacgcggctaattttcag-3' 164 (reverse) h528VL09 5'-ggcgtgtattactgctttcagggcagccatatcccgccaacctttggc-3' 165 (forward) h528VL10 5'-cgcgcgtttaatttccactttggtgccttggccaaaggttggcgg-3' 166 (reverse) Mfe-ahEGFR VL 5'-caattggatattgtgatgacccag-3' 167 (forward) ahEGFR 5'-cgacccaccaccgcccgagccaccgccacccgcgcgtttaatttc-3' 168 VL-(G4S)2 (reverse)
TABLE-US-00024 TABLE 24 Primers for the cloning of C6ML3-9 (Anti-HER2) scFv SEQ ID Name of Primer Sequence NO Sal-G-myc-G4S 5'-acgcgtcgacggggaacaaaaactca 169 (forward) tctcagaagaggatctgggaggcggt ggcagt-3' G2S-G4SX2-Mfel 5'-ggtggcagtggtggtggtggatcagg 170 (forward) aggtggcggatcccaattgcaggtgc agctg-3' Her2 5'-atcgattcaacctaggacggtcagct 171 scFv-stop-Clal t-3' (reverse)
TABLE-US-00025 TABLE 25 Primers for the cloning of h15-2b scFv SEQ ID Name of Primer Sequence NO mfe1-h15-2bVL 5'-caattggacatccagatga 172 (forward) cccagtctcca-3' h15-2bscFv-Clal-Sbfl 5'-cccctgcaggcatcgattt 173 (reverse) atgaggagacggtgac-3'
[0168] Primers for cloning Human anti-CD19 VH and VL are given in Tables 26. The cloned human anti-CD19 VH and VL sequences (Table 13) were then fused with DNA sequence of h15-2b scFv to generate DNA construct for expressing BsAbs of PEGxCD19. Similarly, human anti-CD20 VH and VL were cloned by primers given in Table 27 and the cloned human anti-CD20 and anti-CD22 sequences were then fused with DNA sequence of h15-2b scFv to produce constructs for expressing BsAbs of PEGxCD20 and PEGxCD22, respectively.
TABLE-US-00026 TABLE 26 Primers for the cloning of hHB12b (Anti-CD19) VL and VH SEQ ID Name of Primer Sequence NO hHB12b VL Nael-X + hHB12bVL-1 5'-gccggccgagatcgtgctgacccagagccccgacttccagagc-3' 174 (forward) hHB12bVL-2 5'-ctctgcaggtgatggtcaccttctccttgggggtcacgctctggaagtcgggg-3' 175 (reverse) hHB12bVL-3 5'-gtgaccatcacctgcagagccagcgagagcgtggacaccttcggcatcagcttc-3' 176 (forward) hHB12bVL-4 5'-gctctggtcgggcttctgctggaaccagttcatgaagctgatgccgaaggtg-3' 177 (reverse) hHB12bVL-5 5'-gaagcccgaccagagccccaagctgctgatccacgccgccagcaaccaggg-3' 178 (forward) hHB12bVL-6 5'-cttccgctgccgctgaatctgctgggcacgccgctgccctggttgctggcg-3' 179 (reverse) hHB12bVL-7 5'-ttcagcggcagcggaagcggcaccgacttcaccctgaccatcaacagcctgg-3' 180 (forward) hHB12bVL-8 5'-ctctgctggcagtagtaggttgctgcgtcctcggcctccaggctgttgatggtc -3' 181 (reverse) hHB12bVL-9 5'-aacctactactgccagcagagcaaggaggtgcccttcaccttcggcggcggc-3' 182 (forward) DraIII + hHB12bVL- 5'-gacactcggtgcagccacagtcttgatctccaccttggtgccgccgccgaag-3' 183 10 (reverse) hHB12b VH Hpal + hHB12bVH-1 5'-gttaacgaggtgcagctggtggagagcggcggcggcctggtgca-3' 184 (forward) hHB12bVH-2 5'-cgctggcggcgcagctcagtctcaggctgccgccgggctgcaccaggccgccgc- 185 (reverse) 3' hHB12bVH-3 5'-gctgcgccgccagcggcttcaccttcagcagcagctggatgaactgggtgagac- 186 (forward) 3' hHB12bVH-4 5'-gattctgcccacccactccaggcccttgccgggggcctgtctcacccagttcatcc-3' 187 (reverse) hHB12bVH-5 5'-gagtgggtgggcagaatctaccccggcgacggcgacaccaactacaacggcaa 188 (forward) gttc-3' hHB12bVH-6 5'-tcttgctgtcgtctctgctgatggtgaatctgcccttgaacttgccgttgtagttg-3' 189 (reverse) hHB12bVH-7 5'-ttcagcggcagcggaagcggcaccgacttcaccctgaccatcaacagcctgg-3' 190 (forward) hHB12bVH-8 5'-atgaagccgcttctggcgcagtagtacacggcggtgtcctcggtcttcaggctgtt- 191 (reverse) 3' hHB12bVH-9 5'-cgccagaagcggcttcatcaccaccgtgctggacttcgactactggggccagggc- 192 (forward) 3' Apal + hHB12bVH-10 5'-gggccctttggtggaggcgctgctcacggtcaccagggtgccctggccccagtag- 193 (reverse) 3'
TABLE-US-00027 TABLE 27 Primers for the cloning of F2F (Anti-CD20) VL and VH SEQ ID Name of Primer Sequence NO F2F VL Nael-X-aCD20VL-1 5'-gccggccatggaagccccagctcagcttctcttcctcctgctactctggc-3' 194 (forward) aCD20VL-2 5'-ctggagactgtgtcaacacaatttctccggtggtatctgggagccagagtagcaggagg 195 (reverse) aag-3' aCD20VL-3 5'-aattgtgttgacacagtctccagccaccctgtattgtctccaggggaaagagccaccc-3' 196 (forward) aCD20VL-4 5'-caggctaagtagctgctaacactctgactggccctgcaggagagggtggctctttcccc- 197 (reverse) 3' aCD20VL-5 5'-tgttagcagctacttagcctggtaccaacagaaacctggccaggctcccaggctcctc-3' 198 (forward) aCD20VL-6 5'-ctggctgggatgccagtggccctgttggatgcatcatagatgaggagcctgggagcc-3' 199 (reverse) aCD20VL-7 5'-actggcatcccagccaggttcagtggcagtgggtctgggacagacttcactctcaccat- 200 (forward) 3' aCD20VL-8 5'-ctgacagtaataaactgcaaaatcttcaggctctaggctgctgatggtgagagtgaagtct 201 (reverse) gtcc-3' aCD20VL-9 5'-gaagattttgcagtttattactgtcagcagcgtagcaactggccgatcaccttcggccaag 202 (forward) g-3' DraIII - aCD20VL- 5'-gacactcggtgcagccacagttttaatctccagtcgtgtcccttggccgaaggtgatc-3' 203 10 (reverse) F2F VH Hpal + aCD20VH-1 5'-gttaacatggagttgggactgagctggattttccttttggctatttta-3' 204 (forward) aCD20VH-2 5'-ctccaccagctgcacttcacactggacaccttttaaaatagccaaaaggaaaatccagc- 205 (reverse) -3' aCD20VhH-3 5'-gaagtgcagctggtggagtctgggggaggcttggtacagcctggcaggtccctg-3' 206 (forward) aCD20VH-4 5'-cataatcattaaaggtgaatccagaggctgcacaggagagtctcagggacctgccagg- 207 (reverse) 3' aCD20VH-5 5'-gcctctggattcacctttaatgattatgccatgcactgggtccggcaagctccagggaag- 208 (forward) 3' aCD20VH-6 5'-ggaaccactattccaactaatagttgagacccactccaggcccttccctggagcttgcc-3- ' 209 (reverse) aCD20VH-7 5'-tcaactattagttggaatagtggttccataggctatgcggactctgtgaagggccgattc-- 3' 210 (forward) aCD20VH-8 5'-gatacagggacttcttggcgttgtctctggagatggtgaatcggcccttcacagag -3' 211 (reverse) aCD20VH-9 5'-cgccaagaagtccctgtatctgcaaatgaacagtctgagagctgaggacacggcc-3' 212 (forward) aCD20VH-10 5'-gtagtagttgccgtactgtatatcttttgcacagtaatacaaggccgtgtcctcagc-3' 213 (reverse) aCD20VH-11 5'-agatatacagtacggcaactactactacggtatggacgtaggggccaagggaccac- 214 (forward) 3' Apal - aCD20VH-12 5'-gggccattggtggaggctgaggagacggtgaccgtggtcccttggccc-3' 215 (reverse)
[0169] Production of recombinant PEG2.times.TAG72, PEG2.times.EGFR, PEG2-HER2, PEGxTAG72, PEG.times.EGFR and PEGxHER BsAbs
[0170] CHO-K1/PEG2.times.TAG72 and CHO-K1/PEG2.times.DNS cells that stably secrete PEG2.times.TAG72 and PEG2.times.DNS BsAbs were generated by retroviral transduction of CHO-K1 Chinese hamster ovary cells. PEG2.times.TAG72, PEG2.times.EGFR, PEG2-HER2, PEGxTAG72, PEG.times.EGFR and PEGxHER BsAbs were produced by transient transfection of 293FT cells with corresponding plasmids. 293FT/h6.3Fab.times.CD19 and 293FT/h6.3Fab.times.EGFR cells that stably secreted h6.3Fab.times.CD19 and h6.3Fab.times.EGFR BsAbs were generated by lentiviral transduction. Recombinant lentiviral particles were packaged by co-transfection of pAS3w. Ppuro-pAS3w. Ppuro-h6.3Fab.times.CD19 and pAS3w. Ppuro-h6.3Fab.times.EGFR (7.5 .mu.g) with pCMV.DELTA.R8.91 (6.75 .mu.g) and pMD.G (0.75 .mu.g) using TranslT-LT1 transfection reagent (Mirus Bio, Madison, Wis.) (45 .mu.L) in 293FT cells grown in a 10 cm culture dish (90% confluency). After 48 hr, lentiviral particles were harvested and concentrated by ultracentrifugation (Beckman SW 41 Ti Ultracentrifuge Swing Bucket Rotor, 50,000 g, 1.5 hr, 4.degree. C.). Lentiviral particles were suspended in culture medium containing 5 .mu.g/mL polybrene and filtered through a 0.45 .mu.m filter. 293FT cells were seeded in 6-well plates (1.times.10.sup.5 cells/well) one day before viral infection. Lentivirus containing medium was added to cells and then centrifuged for 1.5 hr (500 g, 32.degree. C.). The cells were selected in puromycin (5 .mu.g/mL) to generate stable cell lines. These anti-PEG BsAbs were purified by affinity chromatography on a TALON column. Briefly, the medium was harvested from CELLine adhere 1000 bioreactors (INTEGRA Biosciences AG, Switzerland) every 7-10 days. Poly-histidine-tagged BsAbs were purified on a Co.sup.2+-TALON column (GE Healthcare Life Sciences, Piscataway, N.J.). The columns were washed by 5-fold bed volumes of binding buffer (0.3 M NaCl/20 mM phosphate/HCl, pH 7.4) and followed by 10-fold bed volumes of washing buffer (0.3 M NaCl/20 mM phosphate/5 mM imidazole/HCl, pH 7.4). These poylhistidine-tagged BsAbs were eluted by elution buffer (0.3 M NaCl/20 mM phosphate/150 mM imidazole/HCl, pH 7.4). The eluted proteins were desalted on Sephadex G-25, equilibrated with PBS and concentrated by ultrafiltration. Protein concentrations were determined by the bicinchoninic acid protein assay (Pierce, Rockford, Ill., U.S.A.).
[0171] Production of PEG.times.EGFR, PEGxHER2 and PEGxDNS BsAbs
[0172] To produce desired BsAbs, the BALB 3T3 producer cells were transfected with plasmids of this invention as described above, and were subsequently sorted by FACS on a MoFIo.TM. XDP (Beckman coulter, Inc., Brea, Calif.) at 4.degree. C. and then incubated into CELLine (INTEGRA Biosciences AG, Zizers, Switzerland) with 1% CCS DMEM. After collecting the culture medium, BsAbs were purified by mPEG affinity chromatography, which was made by coupling 36 mg of o-(2-aminoethyl)-o'-methylpolyethylene glycol 750 (Fluka-Sigma-Aldrich, St. Louis, Mo.) on 1 g of CNBr-activated Sepharose.TM. 4B (GE Healthcare, Little Chalfont, UK). This procedure was performed by following the instruction manual of CNBr-activated Sepharose.TM. 4B (GE Healthcare).
[0173] Purification of Knob in Hole BsAbs
[0174] 293FT cells stably expressing BsAbs were cultured in CellLine adhere 1000 (Integra Biosciences AG, Zizers, Switzerland) in DMEM with 10% low bovine IgG medium (serum was pre-absorbed with protein A resin) at a starting cell number of 5.times.10.sup.7. Culture supernatant was harvested every week. The pooled supernatant was centrifuged at 800 g for 10 min at 4.degree. C. to remove cells and subsequently centrifuged at 15000 rpm for 25 min at 4.degree. C. to remove cell debris. Later on, the supernatant was passed through a 0.45 .mu.M filter and G25 column in phosphate saline buffer (PBS), and finally the bsAb was affinity purified using protein A sepharose. After protein A purification, the purified products were further purified by affinity chromatography by CNBr-activated Sepharose.TM. 4B (Sigma-Aldrich Chemical Co, St. Louis, Mo., USA) conjugated with 35 mg of methyl-PEG.sub.1000-NH.sub.2 per gram of CNBr activated Sepharose.TM. 4B. Subsequently, the purified bsAb were further purified by affinity chromatography using Pierce anti-HA agarose (Thermo Scientific, MA, USA). Purified bsAb fractions were dialyzed against 1000 volumes of PBS three times and concentrated using Amicon Ultra (30 kD cutoff) (Millipore).
[0175] Analysis of the Purified BsAbs
[0176] Five microgram of BsAbs (such as PEG2.times.TAG72, PEG2.times.DNS, mPEG.times.EGFR, mPEGxHER2, mPEGxDNS, 15-2b/BU12, h6.3/BU12, h6.3Fab.times.EGFR, and h6.3Fab.times.CD19 BsAbs) were electrophoresed in 10% SDS-PAGE gels under reducing or non-reducing conditions and then stained by Coomassie Blue.
[0177] ELISA
[0178] The anti-PEG binding specificity of BsAbs was measured by adding graded concentrations of PEG2.times.TAG72,PEG2.times.DNS, hCC49 scFv, h6.3Fab.times.EGFR or h6.3Fab.times.CD19 in 50 .mu.L 2% skim milk to the mucin, BSA-PEG5000, NH.sub.2-PEG.sub.3k-NH.sub.2 or BSA coated plates at RT for 1 h. The plates were washed with PBS-T (PBS containing 0.05% Tween-20) three times. Rabbit anti-6.times.His (2 .mu.g/mL) supplemented with HRP-conjugated goat anti-rabbit (2 .mu.g/mL) or HRP-conjugated goat Ig anti human IgG Fab (2 .mu.g/mL) (Jackson ImmunoResearch Laboratories, West Grove, Pa.) in 50 .mu.L dilution buffer were added for 1 h at room temperature. The plates were washed with PBS-T (PBS containing 0.05% Tween-20) three times and with PBS two times. Bound peroxidase activity was measured by adding 150 .mu.L/well ABTS substrate solution (BioLegend, San Diego, Calif.) for 30 min at room temperature. The absorbance (405 nm) of wells was measured in a microplate reader (Molecular Device, Menlo Park, Calif.).
[0179] Flow Cytometer Analysis
[0180] PEG2.times.TAG72 or control PEG2.times.DNS BsAbs (10 .mu.g/mL) were incubated with A-375(TAG72-), MCF-7(TAG72+), Jurkat (TAG72+) or OVCAR-3(TAG72+) cells at 4.degree. C. for 30 min followed by FITC-labeled goat anti-human immunoglobulin second antibody (2 .mu.g/m L) (Jackson ImmunoResearch Laboratories, West Grove, Pa.) or FITC-labeled 4arm-PEG (2 .mu.g/mL). Tumor-specific targeting of PEGylated compounds was also examined by staining A431 (EGFR.sup.high), and Raji (CD19.sup.high) cell lines with 10 .mu.g/mL of h6.3Fab.times.EGFR or h6.3Fab.times.CD19 BsAbs in PBS containing 0.05% BSA (staining buffer) at 4.degree. C. for 30 min. The cells were washed with cold PBS for 3 times. PEG-Qdot655 (8 nM) (Invitrogen, Grand Island, N.Y.) or PEG-liposomal Texas-Red (100 nm size, 50 .mu.M, lipid conc.) (FormuMax Scientific, Palo Alto, Calif.) in staining buffer was added to cells for 30 min at 4.degree. C. Raji cells (CD19+) or SKBR3 cells (HER2+) were incubated with 10 .mu.g/ml h15-2b-knob/BU12-hole, h15-2b-knob/anti-Her2-hole, h6-3-knob/BU12-hole or h6-3-knob/anti-Her2-hole BsAbs, washed and incubated with 0.25 .mu.g/ml FITC-labeled goat anti-human IgG or 10 nM methoxy-PEG Qdot 655 at 4.degree. C. for 30 min. After washing with cold PBS, the surface fluorescence of 10.sup.4 viable cells was measured by FACScaliber flow cytometer (Becton Dickinson, Mountain View, Calif., USA) then analyzed with Flowjo (Tree Star Inc., San Carlos, Calif., USA).
[0181] Confocal Microscopy of BsAb-Targeted Nano-Particles
[0182] The coverslips (30 mm) in POC chambers were coated with 10 .mu.g/mL poly-L-lysine in PBS for 30 min at room temperature. The coverslips were washed twice with PBS and then 5.times.10.sup.4 cells/chamber of A431 (EGFR.sup.+) tumor cells were seeded on the coverslips. A431 cells were incubated with 10 .mu.g/mL of h6.3Fab.times.EGFR or h6.3Fab.times.CD19 BsAbs at 37.degree. C. for 30 min containing 1 .mu.g/mL of Hoechst 33342 and 100 nM of LysoTracker.RTM. Red DND-99 (Invitrogen Life Technologies Corporation, NY, USA). The cells were washed with culture medium for 2 times. Cell imaging was recorded with an Axiovert 200M Confocal Microscope (Carl Ziess Inc., Thornwood, N.Y.) after adding 16 nM of PEG-Qdot655 solution (Invitrogen Life Technologies Corporation, NY, USA).
[0183] Cytotoxicity Assay
[0184] A431 (EGFR.sup.high) and Raji (CD19.sup.high) cells (5000 cells/well) were seeded in 96-well plates overnight. Fifteen microgram per mL of h6.3Fab.times.EGFR or h6.3Fab.times.CD19 BsAbs were added to the cells for 30 min at 37.degree. C. and followed by graded concentrations of free doxorubicin or PEGylated liposomal doxorubicin (Doxisome.RTM., Taiwan Liposome Company Ltd., Taipei, Taiwan) was added to the cells in triplicate at 37.degree. C. for 4 h. The cells were subsequently washed once and incubated for an additional 48 h in fresh culture medium and then pulsed for 16 h with .sup.3H-thymidine (1 .mu.Ci/well). Results are expressed as percent inhibition of .sup.3H-thymidine incorporation into cellular DNA in comparison to untreated cells.
[0185] In Vivo Optical Imaging of PEG-N1R797 Probes
[0186] BALB/c nude mice bearing Ramos (CD19+) and A431 (EGFR.sup.+) tumor (.about.250 mm.sup.3) in their hind leg regions were intravenously injected with h6.3Fab.times.EGFR (50 .mu.g) and PEG-NIR791 (50 .mu.g). Pentobarbital anesthetized mice were sequentially imaged with an IVIS spectrum optical imaging system (excitation, 745 nm; emission, 840 nm; Perkin-Elmer, Inc., MA, USA) at 45 min, 24 and 48 hr after injection.
[0187] Detecting the Expressed Level of Tumor Markers on Colon and Breast Cancer Cells
[0188] EGFR expression was measured by staining SW480 or SW620 cells with 1 mg/ml Erbitux followed by 1 mg/ml FITC conjugated goat anti-human IgG Fc (Jackson Immuno-Research Laboratories, Westgrove, Pa.) at 4.degree. C. The same procedure were used to measure HER2 expression of SK-BR-3 or MDA-MB-468 cells, which were stained by 1 mg/ml Herceptin followed by 1 mg/ml FITC conjugated goat anti-human IgG Fc. After extensive washing with ice cold PBS, the surface immunofluorescence of viable cells was measured with a FACScan flow cytometer (BD Biosciences, San Diego, Calif.) and fluorescence intensities were analyzed with Cellquest pro software (BD Biosciences).
[0189] Bi-Functional Assay of PEG.times.EGFR and PEGxHER2
[0190] Ninety-six well plates were coated with 2 .mu.g/well of poly-L-lysine (40 .mu.g/ml) in PBS for 5 min at room temperature, washed twice with PBS and then coated with 2.times.10.sup.5 cells/well of SW480 (EGFR.sup.+) or SK-BR-3 (HER2.sup.+) tumor cells. PEG.times.EGFR, PEGxHER2 and PEGxDNS (10 .mu.g/ml) were added to the wells at room temperature for 1 h. The wells were then washed three times with DMEM and 200 ng/ml of Lipo/DOX, 66.7 ng/ml of Lipo/IR780, 100 ng/ml of SN38/PM, 600 ng/ml of FeOdots, 0.5 nM of AuNP and 0.5 nM of Qdot565 nm were added to the wells for 20 mins. After extensive washing with DMEM, the concentrations of PEG-NPs were determined by adding 5 .mu.g/ml of anti-PEG backbone Ab (6-3 Ab from 6-3 hybridoma) for 1 hr, and then DMEM washing three times. In order to amplifying the signals, 0.4 .mu.g/ml of goat anti-mouse IgG Fc-HRP (Jackson ImmunoResearch Laboratories, Inc., PA, USA) was added to the wells. Washing wells fourth times with DMEM, followed by ABTS substrate before absorbance values at 405 nm were measured in a microplate reader (Molecular Device, Menlo Park, Calif., USA).
[0191] Non-Covalent Modification of PEG-NPs with PEG.times.EGFR and PEGxHER2
[0192] BsAbs were added to the PEG-NPs in BSA/PBS buffer (0.05% BSA in 1.times.PBS buffer) at 4.degree. C. for 1 h at protein/PEG-NP ratios of 380-570 .mu.g BsAb/pmol doxorubicin (for Lipo/DOX), 550 .mu.g BsAbs/pmol FeOdot and 140 ng BsAbs/pmol Qdots. After PEG.times.EGFR or PEGxHER2 modification, PEG-NPs became .alpha.EGFR-NPs or .alpha.HER2-NPs.
[0193] Confirm the Conversion of Non-Targeted NPs to Targeted NPs
[0194] Ninety-six well plates were coated with 2 .mu.g/well of poly-L-lysine (40 .mu.g/ml) in PBS for 5 min at room temperature, washed twice with PBS and then coated with 2.times.10.sup.5 cells/well of SW480 (EGFR.sup.+), SW620 (EGFR.sup.-), SK-BR-3 (HER2.sup.+) or MDA-MB-468 (HER2.sup.-) tumor cells.SW480 (EGFR.sup.+) and SW620 (EGFR.sup.-) cells were incubated with 4 .mu.g/ml of .alpha.EGFR-Lipo/DOX, 1 .mu.g/ml of .alpha.EGFR-Lipo/IR780 and 4 .mu.g/ml of FeOdots for 20 mins. After extensive washing with DMEM, the concentrations of PEGylated NPs were determined by adding 5 .mu.g/ml of anti-PEG backbone Ab (6-3 Ab) for 1 hr, and then DMEM washing three times. In order to amplifying the signals, 0.4 .mu.g/ml of goat anti-mouse IgG Fc-HRP (Jacksonlmmuno Research Laboratories, Inc., PA, USA) was added to the wells. Washing wells with DMEM, followed by adding ABTS substrate before absorbance values at 405 nm were measured in a microplate reader (Molecular Device, Menlo Park, Calif., USA). The same procedure was used to examine SK-BR-3 (HER2.sup.+) and MDA-MB-468 (HER2.sup.-) cells that were stained with 4 .mu.g/ml of .alpha.HER2-Lipo/DOX, 4 .mu.g/ml of FeOdots and 2 nM of .alpha.EGFR-Qdot565 nm for 20 mins.
[0195] Confocal Microscopy of BsAb-Targeted NPs
[0196] Circular coverslips (18 mm) in 12 wells plate were coated with 20 .mu.g/well of poly-L-lysine (40 .mu.g/ml) in PBS for 5 min at room temperature. The coverslips were washed twice with PBS and then 4.times.10.sup.4 cells/well of SW480 (EGFR.sup.+), SW620 (EGFR.sup.-), SK-BR-3 (HER2.sup.+) or MDA-MB-468 (HER2.sup.-) tumor cells were coated on the coverslips. SW480 (EGFR.sup.+) and SW620 cells (EGFR.sup.-) were incubated with 300 ng/ml of .alpha.EGFR-Lipo/Rho and .alpha.DNS-Lipo/Rho at 37.degree. C. for 1 h. The cells were fixed with 2% paraformaldehyde in PBS for 30 min at 4.degree. C. and were stained with DAPI for 45 min at 4.degree. C. Then, the coverslips were washed 4 times with PBS, and then mounted with fluorescent mounting medium (Dako, Glostrup, Denmark) on glass microscope slide. The fluorescent signals of .alpha.EGFR-Lipo/Rho and .alpha.DNS-Lipo/Rhowere recorded with an Olympus FluoView.TM. FV1000 Confocal Microscope (Olympus Imaging America Inc., Center Valley, Pa.). The same procedure was used to image SK-BR-3 (HER2.sup.+) and MDA-MB-468 (HER2.sup.-) cells which were stained with 4 nM of .alpha.HER2-Qdot.sub.565 nm and .alpha.DNS-Qdot.sub.565 nm, respectively.
[0197] Targeting of BsAb-Targeted FeOdots by MR Imaging
[0198] MR imaging was performed with a clinical 3.0 T MR imager (Signa; GE Healthcare, Little Chalfont, UK). 1.times.10.sup.7 SW480 (EGFR.sup.+) or SW620 (EGFR.sup.-) cells were incubated with different concentrations of .alpha.EGFR-FeOdots or to .alpha.DNS-FeOdots (7 .mu.M, 14 .mu.M and 28 .mu.M) at 4.degree. C. for 30 min. The cells were washed with PBS 3 times and then the accumulation of BsAbs-FeOdots were scanned by T2-weighted fast spin-echo sequence (TR/TE=2500 ms/60 ms). The same protocol was used to examine localization of .alpha.HER2-FeOdots and .alpha.DNS-FeOdots at SK-BR-3 (HER2.sup.+) and MDA-MB-468 (HER2.sup.-) cells.
[0199] In Vitro Cytotoxicity of BsAb-Targeted Lipo/DOX
[0200] SW480 (EGFR.sup.+) and SW620 (EGFR.sup.-) cells (3.times.10.sup.3/well) were seeded in 96-wells plates. 2 .mu.g/ml or 4 .mu.g/ml of .alpha.EGFR-Lipo/DOX, .alpha.DNS-Lipo/DOX, and Lipo/DOX were added to each well and incubated at 37.degree. C. for 1 h. The medium was replenished and the cells were incubated for 72 h before cell viability was measured with the ATPlite.TM. Luminescence Assay System (Perkin-Elmer, Inc., Waltham, Mass.). Cell viability for SK-BR-3 (HER2.sup.+) or MDA-MB-468 (HER2.sup.-) cells incubated with .alpha.HER2-Lipo/DOX, .alpha.DNS-Lipo/DOX, or Lipo/DOX at 37.degree. C. for 3 h were measured in accordance with the same procedures. Results were expressed as percent inhibition of luminescence as compared with untreated cells by the following formula: % inhibition=100.times.(treated luminescence/untreated luminescence). The standard deviation for each data point was averaged over three samples (n=3).
[0201] In Vivo Optical Imaging of BsAb-Lipo/IR780 and Lipo/IR780
[0202] BALB/c nude mice bearing SW480 (EGFR.sup.+) and SW620 (EGFR.sup.-) tumor (approximately 100 mm.sup.3) in their hind leg regions, were intravenously injected with .alpha.EGFR-Lipo/IR780, .alpha.DNS-Lipo/IR780, and Lipo/IR780 (100 .mu.g per mouse), respectively. Pentobarbital anesthetized mice were sequentially imaged with an IVIS spectrum optical imaging system (excitation, 745 nm; emission, 840 nm; Perkin-Elmer, Inc., Waltham, Mass.) at 24, 48 and 72 h after injection.
[0203] Treatment of EGFR.sup.+ and EGFR.sup.- Tumors with BsAb-Lipo/DOX and Lipo/DOX
[0204] BALB/c nude mice (n=6) were inoculated s.c. with 4.times.10.sup.6 SW480 (EGFR.sup.+) cells and 1.times.10.sup.6 SW620 (EGFR.sup.-) cells in their hind leg regions. After tumor sizes reached to about 20 mm.sup.3, Lipo/DOX, .alpha.DNS-Lipo/DOX and .alpha.EGFR-Lipo/DOX were i.v. administered at 5 mg DOX/kg once weekly for 3 weeks, for a total dose of 15 mg DOX/kg. Other treatment groups included saline. Tumor measurements were performed 3 times a week using a caliper, and tumor sizes were calculated using the equation: (length.times.width.times.high)/2. Mice were weighted once a week to examine the toxicity.
[0205] Statistic Analysis.
[0206] Statistical significance of differences between mean values was estimated with JMP 9.0 software (SAS Institute, Inc., Cary, N.C.) using the nonparametric Mann-Whitney test. P-values in the cytotoxicity assay and in vivo toxicity<0.05 and the P-values in the in vivo treatment<0.01 were considered to be statistically significant.
Example 1 Production and Characterization of Dimerichumanized Bi-Specific Antibodies (BsAbs)
[0207] 1.1 Production of Murine Anti-mPEG or Anti-PEG Abs
[0208] In order to produce humanized BsAbs, three hybridoma cells, E11, 15-2b and 6-3, were identified, and their respective monoclonal Abs were collected by affinity chromatography. The binding specificity of the collected antibodies toward immobilized PEG was then determined. An exemplary binding specificity between monoclonal antibody produced by hybridoma 15-2b and PEG is illustrated in FIG. 4.
[0209] As evident from FIG. 4, monoclonal antibody produced by hybridoma 15-2b bound with CH.sub.3-PEG.sub.750-NH2, instead of NH.sub.2-PEG.sub.3000-NH2; which indicates that such monoclonal antibody specifically recognized the terminal methoxy group of the CH.sub.3-PEG molecules or the terminal hydroxyl group of PEG molecules (FIG. 4), and is thus termed anti-mPEG Ab; whereas the antibody produced by hybridoma 6-3 or E11 specifically recognized the backbone portion and not the terminal methoxy or hydroxyl group or the PEG molecules, and is thus termed anti-PEG Ab.
[0210] DNA encoding the anti-mPEG or anti-PEG Abs was then isolated and sequenced using conventional procedures (i.e., by using oligonucleotide probes that are capable of binding specifically to genes encoding the heavy and light chains of the monoclonal Abs).
[0211] 1.2 Production of Dimerichumanized Anti-PEG (hE11) Anti-TAG72 BsAb
[0212] To produce humanized Abs with bispecificity, the DNA sequence of murine anti-mPEG mAb of example 1.1 was humanized and fused with a humanized single-chain antibody fragment gene against tumor-associated glycoprotein 72 (TAG-72) antigen (hcc49scFv) ora dansyl (DNS) hapten in accordance with procedures described in the Materials and Methods section.
[0213] FIG. 5A is a schematic illustration of the DNA constructs of the humanized bi-specific Abs prepared in this example. In general, each construct encoded in sequence, HA epitope tag (HA), the hE11 anti-PEG light chain, a F2A bicistronic element, the hE11 anti-PEG heavy chain, a hinge-CH.sub.2-CH.sub.3domain, a linker peptide (L), an anti-tumor scFv sequence (e.g., hcc49 scFv for PEG2.times.TAG72 plasmid, and anti-dansyl scFv for the control PEG2.times.DNS plasmid), and a histidine tag. FIG. 5B is a schematic illustration of the dimeric humanized anti-PEG BsAb of this example. Accordingly, BsAbs including PEG2.times.TAG72 and PEG2.times.DNS were produced. SDS-PAGE analysis indicated that BsAbs were composed by a VH-CH1-H-CH2-CH3-scFv fragment (72 kDa) and light chain (35 kDa) under reducing condition (FIG. 5C, right panel); by contrast, a 230 kDa disulfide-linked BsAbs was observed under non-reducing condition (FIG. 5C, left panel). The result was further confirmed by a western blot analysis, in which the HA epitope tag on the N-terminus of the hE11 anti-PEG light chain and the His epitope tag present on the C-terminus of the scFv attached to the hE11 anti-PEG heavy chain were detected, demonstrating that the bispecific antibody was present in the expected conformation (FIG. 5D).
[0214] 1.3 Characterizing the Function of BsAbs of Example 1.2
[0215] Bi-functional activity of the humanized hE11 BsAbs of example 1.2 was examined in this example.
[0216] Binding of the BsAbs was detected by ELISA. Microtiter plates were first coated with antigens and then PEG2.times.TAG72, PEG2.times.DNS BsAbs or hCC49 (anti-TAG72) single chain antibody was added. After the plates were extensively washed to remove unbound antibodies, the remaining bound antibodies in each well were detected with HRP-conjugated secondary antibody. The PEG2.times.TAG72 BsAb was able to bind to both mucin (TAG-72 tumor antigen) (FIG. 6A) as well as BSA-PEG (FIG. 6B) but not to BSA (FIG. 6C), demonstrating that PEG2.times.TAG72 displayed dual antigen specificity to both PEG and mucin. By contrast, the control PEG2.times.DNS BsAb bound to BSA-PEG but not to mucin. Likewise, the hcc49 scFv bound to mucin but not to BSA-PEG. In sum, PEG2.times.TAG72 anti-PEG (hE11) BsAb of example 1.2 can bind both PEG and tumor antigens.
[0217] To determine if the hE11 BsAbs of example 1.2 could bind target cells, MCF-7 breast cancer, Jurkat T cells and OVCAR-3 ovarian cancer cells, which express the TAG-72 antigen recognized by the hCC49 antibody, were incubated with PEG2.times.TAG72 and PEG2.times.DNS BsAbs. A-375 malignant melanoma cells, which do not express the TAG-72 antigen, were used as a negative control cell line. After washing unbound BsAbs from the cells, the BsAbs that remained bound to cells were detected with FITC-conjugated anti-human immunoglobulin antibody. Detection of surface immunofluorescence in a flow cytometer demonstrated that TAG-72 positive cells bound PEG2.times.TAG72 BsAb but not the control PEG2.times.DNS BsAb (FIG. 7, left panels). To test if PEG2.times.TAG72 could simultaneously bind to cancer cells and PEGylated molecules, the cells were first incubated with BsAbs, washed and then incubated with FITC-labeled PEG molecules. TAG-72 positive cells incubated with PEG2.times.TAG72 BsAb but not the control PEG2.times.DNS BsAb could bind PEG-FITC, demonstrating that PEG2.times.TAG72 acted as a true BsAb which could simultaneously bind tumor antigens and PEG molecules (FIG. 7, right panels).
[0218] 1.4 Production and Characterization of Dimerichumanized Anti-PEG (hE11) Anti-EGFR or Anti-HER BsAbs
[0219] To assess whether other cellular targets could be targeted by anti-PEG BsAbs, single-chain antibody fragment genes against epidermal growth factor receptor (EGFR) and the HER2 antigen was fused to the C-terminus of the heavy chain CH3 region gene of the humanized E11 antibody to generate PEG2.times.EGFR and PEG2.times.HER2, respectively in accordance with similar procedures of example 1.2.
[0220] Assessment of the ability of these BsAbs to bind cancer cells indicated that PEG2.times.TAG72 bound with Jurkat T cells (Tag-72 positive) but not MDA-MB-468 cells or BT-474 cells. By contrast, PEG2.times.EGFR BsAb bound to MDA-MB-468 cells (EGFR positive) but not the other two cells, whereas PEG2.times.HER2 BsAbs bound specifically with BT-474 cells (HER2 positive) (FIG. 8A). Thus, these BsAbs bound to respective target cells in an antigen-dependent manner.
[0221] The ability of the bispecific antibodies to simultaneously bind cancer cells and PEGylated compounds was further investigated by first incubating cells with BsAbs, washing unbound antibody from the cells and then adding
[0222] PEG-liposomal Texas Red or PEG-Qdot655 (PEGylated quantum dots). Each BsAb selectively accumulated PEGylated liposomes (FIG. 8B) or PEGylated nanoparticles (FIG. 8C) at cells that expressed the corresponding target antigen on their surface.
[0223] In sum, the anti-PEG BsAbs of this example can simultaneously bind to target antigens and PEGylated substances to selectively accumulate PEGylated compounds and nanoparticles on their respective target cells.
Example 2 Production and Characterization of Monovalent Humanized BsAbs
[0224] 2.1 Production and Characterization of Monovalent Anti-PEG (E11) BsAbs
[0225] Monovalent anti-PEG BsAbs were generated by fusing the Fab fragment of a humanized antibody derived from the anti-PEG antibody E11 to single chain antibodies with specificity for tumor-associated antigens.
[0226] Specifically, the hE11 Fab fragment was fused a single-chain antibody fragment (scFv) derived from anti-TAG72, anti-EGFR (epidermal growth factor receptor) or anti-HER2/Neu antibodies (FIG. 9). CHO cells that stably expressed the monovalent BsAbs were generated and culture medium from each expression cell line was collected.
[0227] The binding specificity of monovalent BsAbs with their target proteins was measured by collecting the culture medium of BsAbs producing cells, adding the collected medium to cells that expressed the target protein, then determining the binding by ELSA. After washing the cells, the bound BsAbs were detected by FITC-labeled goat anti-human immunoglobulin second antibody. It was found that PEGxTAG72 bound to Jurkat T cells (Tag-72 positive), but not MDA-MB-468 cells or BT-474 cells. By contrast, PEG.times.EGFR BsAb bound to MDA-MB-468 cells (EGFR positive) but not the other two cells; whereas PEGxHER2 BsAb was found to bind with BT-474 cells (HER2 positive) specifically (FIG. 10). Thus, monovalent anti-PEG (hE11) BsAbs bound to target cells in an antigen-dependent and selective manner.
[0228] The ability of the monovalent BsAb to simultaneously bind with cancer cells and PEGylated compounds was examined by incubating target cells with BsAbs and PEGylated substances. After washing out the unbound antibody from the cells, PEG-liposomal Texas Red or PEG-Qdot655 (PEGylated quantum dots) were then added. Each BsAbs selectively accumulated PEGylated liposomes (FIG. 11) or PEGylated nanoparticles (FIG. 12) at cells that expressed the corresponding target antigen on their surface. Thus, monovalent anti-PEG (hE11) BsAbs can simultaneously bind to target antigens and PEGylated substances to selectively accumulate PEGylated compounds and nanoparticles on the surface of the target cells.
[0229] 2.2 Production and Characterization of Monovalent Anti-PEG (h6.3) BsAbs
[0230] The humanized anti-PEG (h6.3) Fab was constructed as a single open reading frame by fusing V.sub.L-C.sub..kappa. and V.sub.H-CH.sub.1 with a F2A (furin-2A) peptide linker, allowing the expression of light chain and heavy chain separately in accordance with procedures described in the "Materials and Methods" section. The single chain disulfide-stabilized variable fragments (dsFv) were linked to the C-termius of the CH1 domain in the 6.3Fab via a peptide linker to generate h6.3-11F8 (h6-3Fab.times.EGFR) and h6.3-hBU12 (h6.3Fab.times.CD19) BsAbs (FIG. 13A). These two BsAbs were then inserted into a lentiviral expression vector to generate stable 293FT producer cell lines. BsAbs (including h6-3Fab.times.EGFR and h6.3Fab.times.CD19) that were purified from the culture medium displayed the expected molecular sizes on a 10% SDS-PAGE (FIG. 13B).
[0231] Further, both h6-3Fab.times.EGFR and h6.3Fab.times.CD19 BsAbs bound to the NH.sub.2-PEG.sub.10,000-NH2, but not to the control BSA protein, indicating their binding specificity toward PEG molecule (FIGS. 14A and 14B). As to their binding specificity toward the target antigen, flow cytometer analysis revealed that h6.3Fab.times.EGFR, but not h6.3Fab.times.CD19, was capable of directing the PEGylated substance (including PEG-liposome Texas Red and PEG-Qdot655) to A431 cells (EGFR+); whereas h6.3Fab.times.CD19, but not h6.3Fab.times.EGFR, was capable of directing PEGylated substance to Raji cells (CD19+) (FIG. 14C). The PEG-binding kinetics of h6-3Fab.times.EGFR and h6.3Fab.times.CD19 BsAbs were verified by Microscale Thermophoresis (MST) (FIGS. 14D and 14E).
[0232] To verify whether BsAb-targeted substance could be internalized by antigen-positive cancer cells, live cell imaging was performed by staining cells with lysosome tracker and BsAbs, followed by the addition and incubation of PEG-Qdot655. It was found that h6.3Fab.times.EGFR treated A431 cells displayed red fluorescence within endocytic vesicles, whereas h6.3Fab.times.CD19 treated A431 cells failed to produce PEG-Qdot655 signals (FIG. 15). This observation indicates that h6.3Fab.times.EGFR mediated EGFR endocytosis allows the uptake of PEG-substance into A431 cells.
[0233] Next, the ability of whether h6.3Fab.times.CD19 and h6.3Fab.times.EGFR BsAbs could increase the cytotoxicity of the drug-loaded nanoparticles (NPs) to antigen-positive cancer cells was investigated. Raji (CD19.sup.+) and A431 (EGFR.sup.+) cells were incubated with h6.3Fab.times.CD19 and h6.3Fab.times.EGFR, followed by the addition of graded concentrations of Doxisome (i.e., PEGylated liposomal doxorubicin). When compared with Doxisome alone, both h6.3Fab.times.CD19 and h6.3Fab.times.EGFR enhanced cytotoxicity of Doxisome to Raji, MDA-MB-468 and A431 cells, respectively (FIGS. 16A to 16C). The results indicate that anti-PEG (h6.3) BsAbs may confer tumor selectivity and increase the cytotoxicity of a PEGylated NPs (e.g., doxisome) to antigen-positive cancer cells.
[0234] To determine tumor targeting of anti-PEG (h6.3) BsAbs-NPs in vivo, BALB/c nude mice bearing Ramos (CD19, left side) and SW480 (EGFR, right side) tumors in their hind leg regions were intravenously injected with h6.3 Fab.times.EGFR and PEG-NIR797 probes. The mice were then imaged at 45 min, 24 and 48 hours after injection using an IVIS spectrum optical imaging system. Enhanced signals of PEG-NIR797 were observed in A431 tumors but not in Ramos tumors (FIG. 17), demonstrating that h6.3Fab.times.EGFR BsAbs preferentially deliver PEG probes to EGFR on antigen-positive tumors in vivo.
[0235] 2.3 Production and Characterization of Monovalent Anti-mPEG (h15.2b) BsAbs
[0236] In this example, the DNA sequence of murine anti-mPEG mAb of example 1.1 was humanized and combined with another nucleic acid encoding a single chain variable fragment (scFv) of EGFR or HER2 in accordance with procedures described in the Materials and Methods section.
[0237] FIG. 18A is a schematic illustration of the DNA constructs of the humanized bi-specific Abs prepared in this example, and 3 BsAbs including PEG.times.EGFR, PEGxHER2 and PEGxDNS were produced. In general, each construct encoded in sequence, a signal peptide (SP), HA epitope tag (HA), the anti-mPEG light chain, a F2A bicistronic element, the anti-mPEG heavy chain Fd fragment, a myc epitope tag, a 15 amino acid flexible linker peptide (L) and scFv against an anti-tumor marker sequence, such as anti-EGFR scFv for PEG.times.EGFR plasmid, and anti-HER2 scFv for PEGxHER2 plasmid, and anti-dansyl scFv for the control PEGxDNS plasmid. Accordingly, 3 BsAbs including PEG.times.EGFR, PEGxHER2 and PEGxDNS were produced. SDS-PAGE analysis indicated that BsAbs were composed by a Fd-scFv fragment (56 kDa) and light chain (35 kDa) under reducing condition; by contrast, a 91 kDa disulfide-linked BsAbs was observed under non-reducing condition (FIG. 18B).
[0238] The thus produced humanized BsAbs were then subjected to bi-functional activity assay in antigen-positive or antigen-deficient cancer cells. Briefly, cells with over expressed EGFR (i.e., SW480, EGFR.sup.+) or HER2 (i.e., SK-BR-3, HER2.sup.+) were first incubated with humanized BsAbs of this example, and the unbound BsAbs were washed out with a buffer solution, various PEG-NPs (i.e., Lipo/DOX, Lipo/IR780, SN38/PM, FeOdot, AuNP, and Qdot565 nm) were then added, and the respective binding activities of the BsAbs were detected by ELISA with an anti-mPEG antibody.
[0239] It is noted that PEG.times.EGFR, instead of the negative control PEGxDNS, mediated binding of all the tested PEG-NPs to EGFR.sup.+ SW480 cancer cells (FIG. 18C). Likewise, PEGxHER2, but not PEGxDNS, mediated the binding of PEG-NPs to HER2.sup.+ SK-BR-3 cancer cells (FIG. 18D). In sum, both PEG.times.EGFR and PEGxHER2 display bi-functional binding activity, and are capable of mediating the cross-linking of PEG-NPs to cells that express the EGFR or HER2 tumor markers.
[0240] To evaluate whether the BsAb of this example may confer cancer cell specificity to the PEG-NP, the BsAbs of this example were added to various PEG-NPs (e.g., Lipo/DOX, Lipo/IR780 and FeOdot) to generate targeted PEG-NPs. Binding specificity was then measured by ELISA. Results are depicted in FIGS. 19A to 19B. It is found that PEG-NPs targeting depends on the anti-EGFR portion of the BsAb, for the control .alpha.DNS-NPs failed to bind to either SW480 (EGFR.sup.+) or SW620 (EGFR.sup.-) tumor cells (FIG. 19A). Likewise, incubating PEG-NPs with PEGxHER2, allowed the NPs to bind with SK-BR-3 (HER2.sup.+) tumor cells, but not MDA-MB-468 (HER2.sup.-) tumor cells (FIG. 19B). These results demonstrate that mixing PEG.times.EGFR or PEGxHER2 with PEG-NPs can endow the NPs with specificity to EGFR or HER2 on cancer cells.
[0241] The ability of the targeted PEG-NP in killing antigen-positive cancer cells were further investigated, and results are provided in FIGS. 20A to 20D. As depicted in FIG. 20A, .alpha.EGFR-Lipo/DOX exhibited higher cytotoxicity to SW480 (EGFR.sup.+) cancer cells, as compared with that of Lipo/DOX or .alpha.DNS-Lipo/DOX (FIG. 20A). By contrast, .alpha.EGFR-Lipo/DOX displayed similar cytotoxicity as to that of Lipo/DOX or .alpha.DNS-Lipo/DOX to SW620 (EGFR.sup.-) tumor cells (FIG. 20B). Likewise, .alpha.HER2-Lipo/DOX was significantly more cytotoxic to SK-BR-3 (HER2.sup.+) cancer cells than that caused by Lipo/DOX or .alpha.DNS-Lipo/DOX (FIG. 20C). However, .alpha.HER2-Lipo/DOX displayed similar cytotoxicity to MDA-MB-468 (HER2.sup.-) cancer cells as compared with that of Lipo/DOX or .alpha.DNS-Lipo/DOX (FIG. 20D). Accordingly, it is reasonable to conclude that anti-mPEG BsAbs may confer tumor selectivity and increase the cytotoxicity of a PEG-NP (Lipo/DOX) to antigen-positive cancer cells.
[0242] To investigate tumor targeting of BsAbs-NPs in vivo, BALB/c nude mice bearing EGFR.sup.- SW620 (left side) and EGFR.sup.+ SW480 (right side) tumors in their hind leg regions were intravenously injected with .alpha.EGFR-Lipo/IR780, .alpha.DNS-Lipo/IR780 or Lipo/IR780. The mice were imaged at 24, 48 and 72 hrs after injection with an IVIS spectrum optical imaging system. The fluorescent signal of .alpha.EGFR-Lipo/IR780 was enhanced in SW480 (EGFR.sup.+) tumors as compared to SW620 (EGFR.sup.-) tumors from 24 to 72 hrs after probe injection (FIG. 21, bottom row). The fluorescent intensity of .alpha.EGFR-Lipo/IR780 in SW480 (EGFR.sup.+) tumor were 2.037, 2.318 and 2.328-fold greater at 24, 48 and 72 hrs than SW620 (EGFR.sup.-) tumor, respectively (Table 28). By contrast, Lipo/IR780 and .alpha.DNS-Lipo/IR780 localized more strongly in SW620 tumors, presumably by the EPR effect. These data indicate that .alpha.EGFR-Lipo/IR780 possessed selectivity for EGFR.sup.+ cancer cells, thereby facilitating enhanced accumulation in EGFR.sup.+ tumors.
TABLE-US-00028 TABLE 28 The region of interest (ROI) ratio of SW480 (EGFR.sup.+) to SW620 (EGFR.sup.-) tumors was determined at the indicated times Time i.v. injection ROI ratio 24 h 48 h 72 h Lipo/IR780 EGFR.sup.+ 0.93 0.93 0.92 EGFR.sup.- .alpha.DNS-Lipo/IR780 EGFR.sup.+ 1.09 0.97 1.01 EGFR.sup.- .alpha.EGFR-Lipo/IR780 EGFR.sup.+ 2.04 2.32 2.33 EGFR.sup.-
[0243] To examine whether PEG.times.EGFR of this example may increase the therapeutic efficacy of Lipo/DOX to EGFR.sup.+ tumors in vivo, BALB/c nude mice bearing SW480 (EGFR.sup.+) and SW620 (EGFR.sup.-) tumor in their hind leg regions were treated with Lipo/DOX .alpha.EGFR-Lipo/DOX, .alpha.DNS-Lipo/DOX or saline. It was found that .alpha.EGFR-Lipo/DOX suppressed the growth of SW480 (EGFR.sup.+) tumors significantly more than that treated by Lipo/DOX (P<0.01 on day 8 to 45) (FIG. 22A) without any apparent toxicity, as determined by mouse body weight (FIG. 22C). In the SW620 (EGFR.sup.-) tumor model, there were no significant differences between tumor sizes in mice treated with .alpha.EGFR-Lipo/DOX, .alpha.DNS-Lipo/DOX or Lipo/DOX (FIG. 22B). Accordingly, it is reasonable to conclude that PEG.times.EGFR of this example may indeed enhance the anti-tumor efficacy of Lipo/DOX to EGFR.sup.+ tumors in vivo.
[0244] 2.4 Production and Characterization of Monovalent Anti-mPEG (h15.2b) Anti-CD19 or Anti-CD20 BsAbs
[0245] In this example, the humanized single chain variable fragment (scFv) of murine anti-mPEG mAb was combined with another nucleic acid encoding the monomeric IgG of CD19 or CD20 in accordance with procedures described in the Materials and Methods section.
[0246] FIG. 23A is a schematic illustration of the DNA constructs of the humanized bi-specific Abs prepared in this example. In general, each construct encoded in sequence, a signal peptide (SP), an anti-CD19 or anti-CD20 heavy chain sequence (VH-CH1), an anti-CD19 or anti-CD 20 light chain sequence (VL-C.kappa.), an amino acid flexible linker peptide (L), and the anti-mPEG scFv. Accordingly, 2anti-mPEG BsAbs respectively directed to CD19 and CD20 were produced. Binding results confirmed that the anti-mPEG BsAbs of the present example specifically bound to cells that positively expressed CD19 and CD20 (e.g., Raji cells) (FIG. 23B), as well as the terminal methoxy or hydroxyl group of PEG molecules (FIG. 23C). Further, once the anti-mPEG BsAbs of the present example was mixed with therapeutic nanoparticles (e.g., Lipo/DOX), they were able to target deliver the therapeutic nanoparticles to CD19 or CD-20 positive cancer cells (FIG. 23D).
Example 3 Construction and Characterization of Dimeric Humanized Knob in Hole BsAbs
[0247] 3.1 Production of Knob in Hole Anti-PEG (h6.3 or h15.2b) Anti-HER2 or Anti-CD19 BsAbs
[0248] Recombinant DNA technology was utilized to create BsAbs derived from the cDNA coding regions of V.sub.H and V.sub.L of either an anti-HER2 antibody (C6) or an anti-CD19 antibodies (BU12), and the humanized anti-methoxy-PEG monoclonal h15-2b or the humanized anti-PEG antibody h6.3, with the employment of the "knobs-into-holes" strategy and immunoglobulin domain crossover approach, for heterodimer formation and correct antibody heavy chain and light assembly.
[0249] For generating correctly assembled antibody, the light chain C.kappa. domain and heavy chain CH1 domain within the antigen binding fragment (Fab) of either BU12 or C6 antibody were exchanged, while the anti-PEG antibodies were kept unmodified. In particular, C.sub..kappa. of the tumor antigen antibodies BU12 and C6 was replaced with the partial C.sub.H1 fragment and partial hinge of that antibody heavy chain, and the original C.sub.H1 fragment site within antibody heavy chain was replaced with the C.kappa. sequence of the antibody light chain. This allows the light chain to pair with its cognate heavy chain, instead of pairing with the heavy chain of the anti-PEG-knob antibody. Also, for heavy-chain heterodimer formation, a knob structure (T366W and S354C) was introduced into the h15-2b and h6.3 CH3 region and a hole structure (T366S, L368A, Y407V and Y349C) was introduced into CH3 of C6 and BU12, respectively. The construction maps of BsAbs constructed from the anti-mPEG (h15-2b)-knob antibody and BU12-hole or anti-HER2-hole antibodies are depicted in FIG. 24A; whereas the DNA maps of BsAbs constructed from the anti-PEG antibody (h6.3)-knob and BU12-hole or anti-HER2-hole antibodies are illustrated in FIG. 24B.
[0250] The DNA constructs of BsAbs were then inserted into a lentiviral expression vector to generate stable 293FT producer cell lines. BsAbs (including h15-2b knob+BU12-hole, h6.3+BU12-hole) that were purified from the culture medium displayed the expected molecular sizes on a 10% SDS-PAGE (FIG. 24C).
[0251] 3.2 Characterization of Knob in Hole BsAbs of Example 3.1
[0252] In this example, the bi-specificity of the purified knob in hole BsAbs of example 3.1 was investigated. Briefly, Ramos cells (CD19.sup.+) and SKBR3 cells (HER2.sup.+) were used to verify whether purified BsAbs can bind to both PEG compounds and cancer cells that express the CD19 or Her2/neu tumor antigens. Briefly, Ramos cells (CD19.sup.+), Raji cells (CD19.sup.+) or SKBR3 cells (HER2.sup.+) were incubated with 10 .mu.g/ml h15-2b-knob/BU12-hole or h15-2b-knob/anti-Her2-hole BsAbs, washed and incubated with 0.25 .mu.g/ml FITC-labeled goat anti-human IgGor 10 nM methoxy-PEG Qdot 655 at 4.degree. C. for 30 min. The surface fluorescence of viable cells was measured on a FACSCalibur. Results are illustrated in FIGS. 25 to 28.
[0253] It was found that h15-2b-knob/BU12-hole BsAbs bound to CD19-positive Ramos cells, but not to SKBR3 cells; whereas h15-2b-knob/anti-HER2-hole BsAbs bound to SKBR3 (Her2-positive) (FIG. 25). This indicates that h15-2b BsAbs retained the ability to specifically bind to cancer cells in an antigen-dependent fashion. More importantly, h15-2b-knob/BU12-hole could stably retain PEGylated Qdots at Ramos and Raji cells; and h15-2b-knob/anti-HER2-hole could retain the Qdots at SKBR3 cell (FIG. 26). Thus, these reagents acted as true bispecific molecules. Similar results were observed for h6.3-knob/BU12-hole BsAbs, which could bind to cells expressing CD19 (Ramos and Raji) as measured by FACs (FIG. 27). h6.3-knob/BU12-hole also effectively bound PEG-modified Qdots to Raji cells (FIG. 28), demonstrating that this BsAb could simultaneously bind to CD19 on cancer cells and the PEG molecule of a PEGylated nanoparticle.
Example 4 Construction and Characterization of Recombinant Intact Anti-Cancer BsAbs
[0254] 4.1 Production of Herceptin/h-.alpha.-PEG and Erbitux/h-.alpha.PEG Abs
[0255] To create reagents that may synergistically attack cancer cells, a functional and humanized anti-PEG single-chain Ab (h-.alpha.PEG scFv) was fused to the C-terminal of commercial available targeted antibodies (including Herceptin and Erbitux) to form bi-functional Herceptin/h-.alpha.PEG, and Erbitux/h-.alpha.PEG Abs (FIG. 29). Accordingly, not only have the original anticancer effects of the herceptin and/or Erbitux antibodies been retained, but the newly produced BsAbs can also actively bind to PEGylated drugs at tumor sites, to produce synergistic anticancer effects (i.e., double-attack strategy).
[0256] 4.2 Characterization of the Function of BsAbs of Example 4.1
[0257] In this example, the bi-functional activity of BsAbs of example 4.1 was investigated. Briefly, SKBR-3 human breast adenocarcinoma cells, which overexpress the HER2/c-erb-2 gene product, were coated in 96-well microtiter plates; then Herceptin/h-.alpha.PEG antibodies and control Herceptin antibodies were added to the microtiter plates. After the unbound bi-functional antibodies were washed out, PEGylated liposomes containing doxorubicin therein (herein Lipo-DOX) were added to the wells. Binding of the PEGylated compounds was determined by ELISA.
[0258] As expected, Herceptin/h-.alpha.PEG, but not control Herceptin antibodies, selectively bound Lipo-DOX to SKBR-3 cells (FIG. 30A). Similar results were also observed for A431 Human epithelial carcinoma cells, which exhibited anover expressed level of EGFR. Erbitux/h-.alpha.PEG, but not the control Erbitux antibodies, directed PEGylated compounds to be accumulated on the surface of A431 cells (EGFR.sup.+) (FIG. 30B).
[0259] The anticancer effects of using BsAbs of example 4.1 to target Lipo-Dox to cancer cells was further examined in vitro. Results are depicted in FIGS. 31A and 31B.
[0260] It was confirmed that Herceptin/h-.alpha.PEG (FIG. 31A) and Erbitux/h-.alpha.PEG (FIG. 31B), but not control Abs, when respectively combined with Lipo-Dox exhibited synergistic anti-cancer effects, indicating that the double-attack strategy help attain a higher level of tumor-killing effect.
[0261] In a similar experiment, HER-2 positive SKBR-3 cells were pre-incubated with 5 .mu.g/mL Herceptin/h-.alpha.PEG or Herceptin at 37.degree. C. for 1 h. After washing to remove unbound Abs, cells were then treated with graded concentrations of Lipo-Dox (9, 3, and 0.33 .mu.g/mL) in triplicate for 6 h. Drug-containing medium was then replaced with fresh medium and allowed the cells to continue incubation for an additional 72 hr. Cellular ATP synthesis in the drug-treated cells was then compared with that of the untreated cells.
[0262] It was found that cytotoxicity level was much higher in cells pre-treated with bi-specific Herceptin antibody and Lipo-Dox, than that of the cells treated with either Herceptin alone or Lipo-Dox alone (FIG. 32). The finding is in line with that of FIGS. 31A and 31B, in which a synergistic killing effect caused by Lipo-Dox and the anti-PEG BsAb was observed.
[0263] It will be understood that the above description of embodiments is given by way of example only and that various modifications may be made by those with ordinary skill in the art. The above specification, examples and data provide a complete description of the structure and use of exemplary embodiments of the invention. Although various embodiments of the invention have been described above with a certain degree of particularity, or with reference to one or more individual embodiments, those with ordinary skill in the art could make numerous alterations to the disclosed embodiments without departing from the spirit or scope of this invention.
Sequence CWU 1
1
2261219PRTArtificial SequenceHumanized E11 VL-Ck 1Asp Val Val Met
Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Leu Gly 1 5 10 15 Gln Pro
Ala Ser Ile Ser Cys Arg Ser Ser Lys Ser Ile Val His Ser 20 25 30
Asn Gly Asn Thr Tyr Leu Glu Trp Phe Gln Gln Arg Pro Gly Gln Ser 35
40 45 Pro Arg Arg Leu Ile Tyr Lys Val Ser Lys Arg Met Ser Gly Val
Pro 50 55 60 Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr
Leu Lys Ile 65 70 75 80 Ser Arg Val Glu Ala Glu Asp Val Gly Val Tyr
Tyr Cys Ser Gln Gly 85 90 95 Ser His Val Pro Pro Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 110 Arg Thr Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu 115 120 125 Gln Leu Lys Ser Gly
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe 130 135 140 Tyr Pro Arg
Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln 145 150 155 160
Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 165
170 175 Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu 180 185 190 Lys His Lys Leu Tyr Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser 195 200 205 Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys
210 215 2217PRTArtificial SequenceHumanized E11 VH-CH1 2Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Thr Tyr 20 25
30 Thr Met Asn Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Tyr Ile Ile Pro Ser Ser Gly Tyr Val Asp Tyr Asn Gln
Lys Phe 50 55 60 Lys Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr
Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Val Arg Ser Leu Asp Gly Tyr Phe
Trp Phe Ala Tyr Trp Gly Gln Gly 100 105 110 Thr Leu Val Thr Val Ser
Ser Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125 Pro Leu Ala Pro
Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu 130 135 140 Gly Cys
Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp 145 150 155
160 Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
165 170 175 Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser 180 185 190 Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val
Asn His Lys Pro 195 200 205 Ser Asn Thr Lys Val Asp Lys Arg Val 210
215 3232PRTArtificial SequenceHinge CH2-CH3 3Glu Pro Lys Ser Cys
Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala 1 5 10 15 Pro Glu Leu
Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro 20 25 30 Lys
Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val 35 40
45 Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val
50 55 60 Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
Glu Gln 65 70 75 80 Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr
Val Leu His Gln 85 90 95 Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn Lys Ala 100 105 110 Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Ala Lys Gly Gln Pro 115 120 125 Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr 130 135 140 Lys Asn Gln Val
Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser 145 150 155 160 Asp
Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr 165 170
175 Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr
180 185 190 Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
Val Phe 195 200 205 Ser Cys Ser Val Met His Glu Ala Leu His Asn His
Tyr Thr Gln Lys 210 215 220 Ser Leu Ser Leu Ser Pro Gly Lys 225 230
423PRTArtificial SequencePeptide Linker 4Val Asp Leu Val Thr Val
Ser Ser Ala Ser Thr Gly Gly Gly Ser Gly 1 5 10 15 Gln Leu Gly Gly
Gly Gly Ser 20 5243PRTArtificial SequenceHcc49 dsFv 5Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp His 20 25
30 Ala Ile His Trp Val Arg Gln Ala Pro Gly Gln Cys Leu Glu Trp Met
35 40 45 Gly Tyr Phe Ser Pro Gly Asn Asp Asp Phe Lys Tyr Ser Gln
Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Lys Ser Ala
Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Ser Trp Ile Met Gln Tyr
Trp Gly Gln Gly Thr Leu Val Thr 100 105 110 Val Ser Ser Gly Gly Gly
Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly 115 120 125 Gly Ser Asp Ile
Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser 130 135 140 Leu Gly
Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val Leu 145 150 155
160 Tyr Ser Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro
165 170 175 Gly Gln Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg
Glu Ser 180 185 190 Gly Val Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr 195 200 205 Leu Thr Ile Ser Ser Leu Gln Ala Glu Asp
Val Ala Val Tyr Tyr Cys 210 215 220 Gln Gln Tyr Tyr Ser Tyr Pro Leu
Thr Phe Gly Cys Gly Thr Lys Val 225 230 235 240 Glu Ile Lys
68PRTArtificial Sequence6xHis Tag 6Thr Arg His His His His His His
1 5 7243PRTArtificial Sequence11F8 anti-EGFR dsFv 7Gln Val Gln Leu
Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Gln 1 5 10 15 Thr Leu
Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Gly 20 25 30
Asp Tyr Tyr Trp Ser Trp Ile Arg Gln Pro Pro Gly Lys Cys Leu Glu 35
40 45 Trp Ile Gly Tyr Ile Tyr Tyr Ser Gly Ser Thr Asp Tyr Asn Pro
Ser 50 55 60 Leu Lys Ser Arg Val Thr Met Ser Val Asp Thr Ser Lys
Asn Gln Phe 65 70 75 80 Ser Leu Lys Val Asn Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr 85 90 95 Cys Ala Arg Val Ser Ile Phe Gly Val
Gly Thr Phe Asp Tyr Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser Gly Gly Gly Gly Ser Gly Gly 115 120 125 Gly Gly Ser Gly Gly
Gly Gly Ser Glu Ile Val Met Thr Gln Ser Pro 130 135 140 Ala Thr Leu
Ser Leu Ser Pro Gly Glu Arg Ala Thr Leu Ser Cys Arg 145 150 155 160
Ala Ser Gln Ser Val Ser Ser Tyr Leu Ala Trp Tyr Gln Gln Lys Pro 165
170 175 Gly Gln Ala Pro Arg Leu Leu Ile Tyr Asp Ala Ser Asn Arg Ala
Thr 180 185 190 Gly Ile Pro Ala Arg Phe Ser Gly Ser Gly Ser Gly Thr
Asp Phe Thr 195 200 205 Leu Thr Ile Ser Ser Leu Glu Pro Glu Asp Phe
Ala Val Tyr Tyr Cys 210 215 220 His Gln Tyr Gly Ser Thr Pro Leu Thr
Phe Gly Cys Gly Thr Lys Ala 225 230 235 240 Glu Ile Lys
8255PRTArtificial SequenceC6ML3-9 anti-HER2 dsFv 8Gln Val Gln Leu
Leu Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Glu 1 5 10 15 Ser Leu
Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr 20 25 30
Trp Ile Ala Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu Tyr Met 35
40 45 Gly Leu Ile Tyr Pro Gly Asp Ser Asp Thr Lys Tyr Ser Pro Ser
Phe 50 55 60 Gln Gly Gln Val Thr Ile Ser Val Asp Lys Ser Val Ser
Thr Ala Tyr 65 70 75 80 Leu Gln Trp Ser Ser Leu Lys Pro Ser Asp Ser
Ala Val Tyr Phe Cys 85 90 95 Ala Arg His Asp Val Gly Tyr Cys Ser
Ser Ser Asn Cys Ala Lys Trp 100 105 110 Pro Glu Tyr Phe Gln His Trp
Gly Gln Gly Thr Leu Val Thr Val Ser 115 120 125 Ser Gly Gly Gly Gly
Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser 130 135 140 Gln Ser Val
Leu Thr Gln Pro Pro Ser Val Ser Ala Ala Pro Gly Gln 145 150 155 160
Lys Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Asn Asn 165
170 175 Tyr Val Ser Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu
Leu 180 185 190 Ile Tyr Asp His Thr Asn Arg Pro Ala Gly Val Pro Asp
Arg Phe Ser 195 200 205 Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala
Ile Ser Gly Phe Arg 210 215 220 Ser Glu Asp Glu Ala Asp Tyr Tyr Cys
Ala Ser Trp Asp Tyr Thr Leu 225 230 235 240 Ser Gly Trp Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 245 250 255 9155PRTArtificial
SequenceHumanized 6.3 VL-Ck 9Asp Ile Val Met Thr Gln Ser Pro Asp
Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Val Leu Tyr Ser 20 25 30 Ser Asn Gln Met Asn
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45 Pro Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60 Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 65 70
75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Leu
Gln 85 90 95 Tyr Leu Ser Ser Trp Thr Phe Gly Gly Gly Thr Lys Leu
Glu Ile Lys 100 105 110 Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser
Lys Ala Asp Tyr Glu 115 120 125 Lys His Lys Leu Tyr Ala Cys Glu Val
Thr His Gln Gly Leu Ser Ser 130 135 140 Pro Val Thr Lys Ser Phe Asn
Arg Gly Glu Cys 145 150 155 10219PRTArtificial SequenceHumanized
6.3 VH-CH1 10Gln Val Gln Leu Val Gln Ser Gly Ser Glu Leu Lys Lys
Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr
Thr Phe Lys Asn Tyr 20 25 30 Gly Met Asn Trp Val Arg Gln Ala Pro
Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Trp Ile Asn Thr Tyr Thr
Gly Gln Pro Ile Tyr Ala Asn Asp Phe 50 55 60 Lys Gly Arg Phe Val
Phe Ser Leu Asp Thr Ser Val Ser Thr Ala Tyr 65 70 75 80 Leu Gln Ile
Ser Ser Leu Lys Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala
Arg Asp Trp Gly Pro Tyr Trp Gly Gln Gly Thr Leu Val Thr Val 100 105
110 Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser
115 120 125 Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu
Val Lys 130 135 140 Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp Asn
Ser Gly Ala Leu 145 150 155 160 Thr Ser Gly Val His Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu 165 170 175 Tyr Ser Leu Ser Ser Val Val
Thr Val Pro Ser Ser Ser Leu Gly Thr 180 185 190 Gln Thr Tyr Ile Cys
Asn Val Asn His Lys Pro Ser Asn Thr Lys Val 195 200 205 Asp Lys Arg
Val Glu Pro Lys Ser Cys Asp Lys 210 215 11242PRTArtificial
SequencehBU12 dsFv 11Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu
Val Lys Pro Ser Gln 1 5 10 15 Thr Leu Ser Leu Thr Cys Thr Val Ser
Gly Gly Ser Ile Ser Thr Ser 20 25 30 Gly Met Gly Val Gly Trp Ile
Arg Gln His Pro Gly Lys Cys Leu Glu 35 40 45 Trp Ile Gly His Ile
Trp Trp Asp Asp Asp Lys Arg Tyr Asn Pro Ala 50 55 60 Leu Lys Ser
Arg Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln Phe 65 70 75 80 Ser
Leu Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr 85 90
95 Cys Ala Arg Met Glu Leu Trp Ser Tyr Tyr Phe Asp Tyr Trp Gly Gln
100 105 110 Gly Thr Leu Val Thr Val Ser Ser Gly Gly Gly Gly Ser Gly
Gly Gly 115 120 125 Gly Ser Gly Gly Gly Gly Ser Glu Ile Val Leu Thr
Gln Ser Pro Ala 130 135 140 Thr Leu Ser Leu Ser Pro Gly Glu Arg Ala
Thr Leu Ser Cys Ser Ala 145 150 155 160 Ser Ser Ser Val Ser Tyr Met
His Trp Tyr Gln Gln Lys Pro Gly Gln 165 170 175 Ala Pro Arg Leu Leu
Ile Tyr Asp Thr Ser Lys Leu Ala Ser Gly Ile 180 185 190 Pro Ala Arg
Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 195 200 205 Ile
Ser Ser Leu Glu Pro Glu Asp Val Ala Val Tyr Tyr Cys Phe Gln 210 215
220 Gly Ser Val Tyr Pro Phe Thr Phe Gly Cys Gly Thr Lys Leu Glu Ile
225 230 235 240 Lys Arg 12214PRTArtificial SequenceHumanized 15-2b
VL-Ck 12Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val
Gly 1 5 10 15 Asp Arg Val Thr Ile Thr Cys Lys Ala Ser Gln Asp Val
Asn Thr Ser 20 25 30 Val Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala
Pro Lys Leu Leu Ile 35 40 45 Tyr Trp Ala Ser Thr Arg His Thr Gly
Val Pro Ser Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe
Thr Phe Thr Ile Ser Ser Leu Gln Pro 65 70 75 80 Glu Asp Ile Ala Thr
Tyr Tyr Cys Leu Gln Tyr Ile Asn Tyr Pro Tyr 85 90 95 Thr Phe Gly
Gln Gly Thr Lys Leu Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro
Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120
125 Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala
130 135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn
Ser Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser
Thr Tyr Ser Leu Ser 165 170 175 Ser Thr Leu
Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Leu Tyr 180 185 190 Ala
Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205 Phe Asn Arg Gly Glu Cys 210 13215PRTArtificial
SequenceHumanized 15-2b VH-CH1 13Glu Val Gln Leu Val Glu Ser Gly
Gly Gly Leu Val Gln Pro Gly Gly 1 5 10 15 Ser Leu Lys Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr 20 25 30 Trp Met Asn Trp
Val Arg Gln Ala Ser Gly Lys Gly Leu Glu Trp Val 35 40 45 Gly Glu
Ile Arg Ser Lys Ser Asn Asn Tyr Ala Thr His Tyr Ala Glu 50 55 60
Ser Val Lys Gly Arg Phe Thr Ile Ser Arg Asp Asp Ser Lys Asn Thr 65
70 75 80 Ala Tyr Leu Gln Met Asn Ser Leu Lys Thr Glu Asp Thr Ala
Val Tyr 85 90 95 Tyr Cys Ser Asn Arg Tyr Tyr Trp Gly Gln Gly Thr
Leu Val Thr Val 100 105 110 Ser Ser Ala Ser Thr Lys Gly Pro Ser Val
Phe Pro Leu Ala Pro Cys 115 120 125 Ser Arg Ser Thr Ser Glu Ser Thr
Ala Ala Leu Gly Cys Leu Val Lys 130 135 140 Asp Tyr Phe Pro Glu Pro
Val Thr Val Ser Trp Asn Ser Gly Ala Leu 145 150 155 160 Thr Ser Gly
Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu 165 170 175 Tyr
Ser Leu Ser Ser Val Val Thr Val Pro Ser Ser Asn Phe Gly Thr 180 185
190 Gln Thr Tyr Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val
195 200 205 Asp Lys Thr Val Glu Arg Lys 210 215 1428PRTArtificial
SequenceG-MYC-(G4S)3 Linker 14Gly Glu Gln Lys Leu Ile Ser Glu Glu
Asp Leu Gly Gly Gly Gly Ser 1 5 10 15 Gly Gly Gly Gly Ser Gly Gly
Gly Gly Ser Gln Leu 20 25 15255PRTArtificial SequenceC6ML3-9
(Anti-HER2) scFv 15Gln Val Gln Leu Leu Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Glu 1 5 10 15 Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly
Tyr Ser Phe Thr Ser Tyr 20 25 30 Trp Ile Ala Trp Val Arg Gln Met
Pro Gly Lys Gly Leu Glu Tyr Met 35 40 45 Gly Leu Ile Tyr Pro Gly
Asp Ser Asp Thr Lys Tyr Ser Pro Ser Phe 50 55 60 Gln Gly Gln Val
Thr Ile Ser Val Asp Lys Ser Val Ser Thr Ala Tyr 65 70 75 80 Leu Gln
Trp Ser Ser Leu Lys Pro Ser Asp Ser Ala Val Tyr Phe Cys 85 90 95
Ala Arg His Asp Val Gly Tyr Cys Ser Ser Ser Asn Cys Ala Lys Trp 100
105 110 Pro Glu Tyr Phe Gln His Trp Gly Gln Gly Thr Leu Val Thr Val
Ser 115 120 125 Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly
Gly Gly Ser 130 135 140 Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser
Ala Ala Pro Gly Gln 145 150 155 160 Lys Val Thr Ile Ser Cys Ser Gly
Ser Ser Ser Asn Ile Gly Asn Asn 165 170 175 Tyr Val Ser Trp Tyr Gln
Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu 180 185 190 Ile Tyr Asp His
Thr Asn Arg Pro Ala Gly Val Pro Asp Arg Phe Ser 195 200 205 Gly Ser
Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly Phe Arg 210 215 220
Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Ser Trp Asp Tyr Thr Leu 225
230 235 240 Ser Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu
Gly 245 250 255 16247PRTArtificial Sequenceh528 (Anti-EGFR) scFv
16Asp Ile Val Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly 1
5 10 15 Glu Pro Ala Ser Ile Ser Cys Arg Ser Ser Gln Asn Ile Val His
Asn 20 25 30 Asn Gly Ile Thr Tyr Leu Glu Trp Tyr Leu Gln Lys Pro
Gly Gln Ser 35 40 45 Pro Gln Leu Leu Ile Tyr Lys Val Ser Asp Arg
Phe Ser Gly Val Pro 50 55 60 Asp Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Lys Ile 65 70 75 80 Ser Arg Val Glu Ala Glu Asp
Val Gly Val Tyr Tyr Cys Phe Gln Gly 85 90 95 Ser His Ile Pro Pro
Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 110 Arg Ala Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 115 120 125 Ser
Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly 130 135
140 Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
145 150 155 160 Tyr Trp Met His Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp 165 170 175 Met Gly Asn Ile Tyr Pro Gly Ser Gly Gly Thr
Asn Tyr Ala Glu Lys 180 185 190 Phe Lys Asn Arg Val Thr Met Thr Arg
Asp Thr Ser Ile Ser Thr Ala 195 200 205 Tyr Met Glu Leu Ser Arg Leu
Arg Ser Asp Asp Thr Ala Val Tyr Tyr 210 215 220 Cys Ala Arg Ser Gly
Gly Pro Tyr Phe Phe Asp Tyr Trp Gly Gln Gly 225 230 235 240 Thr Leu
Val Thr Val Ser Ser 245 17227PRTArtificial SequenceHumanized 15-2b
scFv 17Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val
Gly 1 5 10 15 Asp Arg Val Thr Ile Thr Cys Lys Ala Ser Gln Asp Val
Asn Thr Ser 20 25 30 Val Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala
Pro Lys Leu Leu Ile 35 40 45 Tyr Trp Ala Ser Thr Arg His Thr Gly
Val Pro Ser Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe
Thr Phe Thr Ile Ser Ser Leu Gln Pro 65 70 75 80 Glu Asp Ile Ala Thr
Tyr Tyr Cys Leu Gln Tyr Ile Asn Tyr Pro Tyr 85 90 95 Thr Phe Gly
Gln Gly Thr Lys Leu Glu Ile Lys Arg Gly Gly Gly Gly 100 105 110 Ser
Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly 115 120
125 Gly Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn
130 135 140 Tyr Trp Met Asn Trp Val Arg Gln Ala Ser Gly Lys Gly Leu
Glu Trp 145 150 155 160 Val Gly Glu Ile Arg Ser Lys Ser Asn Asn Tyr
Ala Thr His Tyr Ala 165 170 175 Glu Ser Val Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asp Ser Lys Asn 180 185 190 Thr Ala Tyr Leu Gln Met Asn
Ser Leu Lys Thr Glu Asp Thr Ala Val 195 200 205 Tyr Tyr Cys Thr Asn
Arg Tyr Tyr Trp Gly Gln Gly Thr Leu Val Thr 210 215 220 Val Ser Ser
225 18219PRTArtificial SequencehHB12b (Anti-CD19) VH-CH1 18Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly 1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Ser 20
25 30 Trp Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp
Val 35 40 45 Gly Arg Ile Tyr Pro Gly Asp Gly Asp Thr Asn Tyr Asn
Gly Lys Phe 50 55 60 Lys Gly Arg Phe Thr Ile Ser Arg Asp Asp Ser
Lys Asn Ser Leu Tyr 65 70 75 80 Leu Gln Met Asn Ser Leu Lys Thr Glu
Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Ser Gly Phe Ile Thr
Thr Val Leu Asp Phe Asp Tyr Trp Gly 100 105 110 Gln Gly Thr Leu Val
Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro
Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala 130 135 140 Ala
Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150
155 160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro
Ala 165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val
Val Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile
Cys Asn Val Asn His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys
Arg Val 210 215 19217PRTArtificial SequencehHB12b (Anti-CD19) VL-Ck
19Glu Ile Val Leu Thr Gln Ser Pro Asp Phe Gln Ser Val Thr Pro Lys 1
5 10 15 Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Glu Ser Val Asp Thr
Phe 20 25 30 Gly Ile Ser Phe Met Asn Trp Phe Gln Gln Lys Pro Asp
Gln Ser Pro 35 40 45 Lys Leu Leu Ile His Ala Ala Ser Asn Gln Gly
Ser Gly Val Pro Ser 50 55 60 Arg Phe Ser Gly Ser Gly Ser Gly Thr
Asp Phe Thr Leu Thr Ile Asn 65 70 75 80 Ser Leu Glu Ala Glu Asp Ala
Ala Thr Tyr Tyr Cys Gln Gln Ser Lys 85 90 95 Glu Val Pro Phe Thr
Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Thr 100 105 110 Val Ala Ala
Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu 115 120 125 Lys
Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro 130 135
140 Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly
145 150 155 160 Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp
Ser Thr Tyr 165 170 175 Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala
Asp Tyr Glu Lys His 180 185 190 Lys Leu Tyr Ala Cys Glu Val Thr His
Gln Gly Leu Ser Ser Pro Val 195 200 205 Thr Lys Ser Phe Asn Arg Gly
Glu Cys 210 215 20239PRTArtificial Sequence2F2 (Anti-CD20) VH-CH1
20Met Glu Leu Gly Leu Ser Trp Ile Phe Leu Leu Ala Ile Leu Lys Gly 1
5 10 15 Val Gln Cys Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val
Gln 20 25 30 Pro Gly Arg Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
Phe Thr Phe 35 40 45 Asn Asp Tyr Ala Met His Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu 50 55 60 Glu Trp Val Ser Thr Ile Ser Trp Asn
Ser Gly Ser Ile Gly Tyr Ala 65 70 75 80 Asp Ser Val Lys Gly Arg Phe
Thr Ile Ser Arg Asp Asn Ala Lys Lys 85 90 95 Ser Leu Tyr Leu Gln
Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Leu 100 105 110 Tyr Tyr Cys
Ala Lys Asp Ile Gln Tyr Gly Asn Tyr Tyr Tyr Gly Met 115 120 125 Asp
Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Ala Ser Thr 130 135
140 Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser
145 150 155 160 Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr
Phe Pro Glu 165 170 175 Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu
Thr Ser Gly Val His 180 185 190 Thr Phe Pro Ala Val Leu Gln Ser Ser
Gly Leu Tyr Ser Leu Ser Ser 195 200 205 Val Val Thr Val Pro Ser Ser
Ser Leu Gly Thr Gln Thr Tyr Ile Cys 210 215 220 Asn Val Asn His Lys
Pro Ser Asn Thr Lys Val Asp Lys Arg Val 225 230 235
21233PRTArtificial Sequence2F2 (Anti-CD20) VL-Ck 21Met Glu Ala Pro
Ala Gln Leu Leu Phe Leu Leu Leu Leu Trp Leu Pro 1 5 10 15 Asp Thr
Thr Gly Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser 20 25 30
Leu Ser Pro Gly Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser 35
40 45 Val Ser Ser Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro 50 55 60 Arg Leu Leu Ile Tyr Asp Ala Ser Asn Arg Ala Thr Gly
Ile Pro Ala 65 70 75 80 Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser 85 90 95 Ser Leu Glu Pro Glu Asp Phe Ala Val
Tyr Tyr Cys Gln Gln Arg Ser 100 105 110 Asn Trp Pro Ile Thr Phe Gly
Gln Gly Thr Arg Leu Glu Ile Lys Thr 115 120 125 Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu 130 135 140 Lys Ser Gly
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro 145 150 155 160
Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly 165
170 175 Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr 180 185 190 Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His 195 200 205 Lys Leu Tyr Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val 210 215 220 Thr Lys Ser Phe Asn Arg Gly Glu Cys
225 230 22232PRTArtificial SequenceKnob Hinge CH2-CH3 22Glu Pro Lys
Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala 1 5 10 15 Pro
Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro 20 25
30 Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val
35 40 45 Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp
Tyr Val 50 55 60 Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu Glu Gln 65 70 75 80 Tyr Asn Ser Thr Tyr Arg Val Val Ser Val
Leu Thr Val Leu His Gln 85 90 95 Asp Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser Asn Lys Ala 100 105 110 Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro 115 120 125 Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Cys Arg Asp Glu Leu Thr 130 135 140 Lys Asn
Gln Val Ser Leu Trp Cys Leu Val Lys Gly Phe Tyr Pro Ser 145 150 155
160 Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr
165 170 175 Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe
Leu Tyr 180 185 190 Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly Asn Val Phe 195 200 205 Ser Cys Ser Val Met His Glu Ala Leu His
Asn His Tyr Thr Gln Lys 210 215 220 Ser Leu Ser Leu Ser Pro Gly Lys
225 230 23207PRTArtificial SequencehBU12 VL-crossover CH1 23Glu Ile
Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Ser Ala Ser Ser Ser Val Ser Tyr Met 20
25 30 His Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
Tyr 35 40 45 Asp Thr Ser Lys Leu Ala Ser Gly Ile Pro Ala Arg Phe
Ser Gly Ser 50 55 60 Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Glu Pro Glu 65 70 75 80 Asp Val Ala Val Tyr Tyr Cys Phe Gln
Gly Ser
Val Tyr Pro Phe Thr 85 90 95 Phe Gly Gln Gly Thr Lys Leu Glu Ile
Lys Arg Ser Ser Ala Ser Thr 100 105 110 Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Ser Ser Lys Ser Thr Ser 115 120 125 Gly Gly Thr Ala Ala
Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu 130 135 140 Pro Val Thr
Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His 145 150 155 160
Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser 165
170 175 Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile
Cys 180 185 190 Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys
Lys Val 195 200 205 24227PRTArtificial SequencehBU12 VH-crossover
Ck 24Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser
Gln 1 5 10 15 Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile
Ser Thr Ser 20 25 30 Gly Met Gly Val Gly Trp Ile Arg Gln His Pro
Gly Lys Gly Leu Glu 35 40 45 Trp Ile Gly His Ile Trp Trp Asp Asp
Asp Lys Arg Tyr Asn Pro Ala 50 55 60 Leu Lys Ser Arg Val Thr Ile
Ser Val Asp Thr Ser Lys Asn Gln Phe 65 70 75 80 Ser Leu Lys Leu Ser
Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr 85 90 95 Cys Ala Arg
Met Glu Leu Trp Ser Tyr Tyr Phe Asp Tyr Trp Gly Gln 100 105 110 Gly
Thr Leu Val Thr Val Ser Ser Ala Ser Val Ala Ala Pro Ser Val 115 120
125 Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly Thr Ala Ser
130 135 140 Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala Lys
Val Gln 145 150 155 160 Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn
Ser Gln Glu Ser Val 165 170 175 Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser Ser Thr Leu 180 185 190 Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Leu Tyr Ala Cys Glu 195 200 205 Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser Phe Asn Arg 210 215 220 Gly Glu Cys
225 25227PRTArtificial Sequencehole hinge-CH2-CH3 25Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly 1 5 10 15 Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met 20 25 30
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser His 35
40 45 Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu
Val 50 55 60 His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn
Ser Thr Tyr 65 70 75 80 Arg Val Val Ser Val Leu Thr Val Leu His Gln
Asp Trp Leu Asn Gly 85 90 95 Lys Glu Tyr Lys Cys Lys Val Ser Asn
Lys Ala Leu Pro Ala Pro Ile 100 105 110 Glu Lys Thr Ile Ser Lys Ala
Lys Gly Gln Pro Arg Glu Pro Gln Val 115 120 125 Cys Thr Leu Pro Pro
Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser 130 135 140 Leu Ser Cys
Ala Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu 145 150 155 160
Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro 165
170 175 Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Val Ser Lys Leu Thr
Val 180 185 190 Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys
Ser Val Met 195 200 205 His Glu Ala Leu His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu Ser 210 215 220 Pro Gly Lys 225 26211PRTArtificial
SequenceC6ML3-9 VL-crossover CH1 26Gln Ser Val Leu Thr Gln Pro Pro
Ser Val Ser Ala Ala Pro Gly Gln 1 5 10 15 Lys Val Thr Ile Ser Cys
Ser Gly Ser Ser Ser Asn Ile Gly Asn Asn 20 25 30 Tyr Val Ser Trp
Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu 35 40 45 Ile Tyr
Asp His Thr Asn Arg Pro Ala Gly Val Pro Asp Arg Phe Ser 50 55 60
Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly Phe Arg 65
70 75 80 Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Ser Trp Asp Tyr
Thr Leu 85 90 95 Ser Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr
Val Leu Gly Ser 100 105 110 Ser Ala Ser Thr Lys Gly Pro Ser Val Phe
Pro Leu Ala Pro Ser Ser 115 120 125 Lys Ser Thr Ser Gly Gly Thr Ala
Ala Leu Gly Cys Leu Val Lys Asp 130 135 140 Tyr Phe Pro Glu Pro Val
Thr Val Ser Trp Asn Ser Gly Ala Leu Thr 145 150 155 160 Ser Gly Val
His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr 165 170 175 Ser
Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln 180 185
190 Thr Tyr Ile Cys Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp
195 200 205 Lys Lys Val 210 27236PRTArtificial SequenceC6ML3-9
VH-crossover Ck 27Gln Val Gln Leu Leu Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Glu 1 5 10 15 Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly
Tyr Ser Phe Thr Ser Tyr 20 25 30 Trp Ile Ala Trp Val Arg Gln Met
Pro Gly Lys Gly Leu Glu Tyr Met 35 40 45 Gly Leu Ile Tyr Pro Gly
Asp Ser Asp Thr Lys Tyr Ser Pro Ser Phe 50 55 60 Gln Gly Gln Val
Thr Ile Ser Val Asp Lys Ser Val Ser Thr Ala Tyr 65 70 75 80 Leu Gln
Trp Ser Ser Leu Lys Pro Ser Asp Ser Ala Val Tyr Phe Cys 85 90 95
Ala Arg His Asp Val Gly Tyr Cys Ser Ser Ser Asn Cys Ala Lys Trp 100
105 110 Pro Glu Tyr Phe Gln His Trp Gly Gln Gly Thr Leu Val Thr Val
Ser 115 120 125 Ser Ala Ser Val Ala Ala Pro Ser Val Phe Ile Phe Pro
Pro Ser Asp 130 135 140 Glu Gln Leu Lys Ser Gly Thr Ala Ser Val Val
Cys Leu Leu Asn Asn 145 150 155 160 Phe Tyr Pro Arg Glu Ala Lys Val
Gln Trp Lys Val Asp Asn Ala Leu 165 170 175 Gln Ser Gly Asn Ser Gln
Glu Ser Val Thr Glu Gln Asp Ser Lys Asp 180 185 190 Ser Thr Tyr Ser
Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr 195 200 205 Glu Lys
His Lys Leu Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser 210 215 220
Ser Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 225 230 235
2839DNAArtificial SequenceCH1 primer F 28ctggtcaccg tctcctcagc
ctccaccaag ggaccatcg 392925DNAArtificial SequenceCH1 primer R
29gtcgactttg tcacaagatt tgggc 253039DNAArtificial SequenceCk primer
F 30accaaggtgg agatcaaacg gactgtggct gcaccatct 393126DNAArtificial
SequenceCk primer R 31ctcgaggcac tctcccctgt tgaagc
263234DNAArtificial SequenceFc primer F 32ggtggacaag agagttgagc
ccaaatcttg tgac 343327DNAArtificial SequenceFc primer R
33caattgtcca ctgccacccc cgcttga 273433DNAArtificial SequencehE11VL
primer F 34ggcccagccg gccgatgttg tgatgactca gtc 333535DNAArtificial
SequencehE11VL- partial Ck primer R 35gtgcagccac agtccgtttg
atctccacct tggtc 353631DNAArtificial Sequenceh6-3VL primer F
36ggcccagccg gccgacatcg tgatgaccca g 313735DNAArtificial
Sequenceh6-3VL-partial Ck primer R 37gtgcagccac agtccgtttg
atttccacct tggtc 353831DNAArtificial Sequenceh15-2bVL primer F
38ggcccagccg gccgacatcc agatgaccca g 313935DNAArtificial
Sequenceh15-2bVL- partial Ck primer R 39gtgcagccac agtccgtttg
atctccagct tggtc 354024DNAArtificial SequencehE11VH primer F
40agatctcagg tgcagctggt gcag 244135DNAArtificial SequencehE11VH-
partial CH1 primer R 41tcccttggtg gaggctgagg agacggtgac caggg
354226DNAArtificial Sequenceh6-3VLH primer F 42agatctcagg
tgcagctggt gcaatc 264334DNAArtificial Sequenceh6-3VH-partial CH1
primer R 43gtcccttggt ggaggctgag gagacggtga ccag
344424DNAArtificial Sequenceh15-2bVH primer F 44agatctgagg
tgcagctggt ggag 244535DNAArtificial Sequenceh15-2bVH- partial CH1
primer R 45gcccttggtg gaggctgagg agacggtgac caggg
354648DNAArtificial SequencehBU12 dsFv primer P1 46caattgcagg
ttcagctgca agagtctggc cctgggttgg ttaagccc 484749DNAArtificial
SequencehBU12 dsFv primer P2 47cagtacaagt cagactgagg gtctgggagg
gcttaaccaa cccagggcc 494849DNAArtificial SequencehBU12 dsFv primer
P3 48cagtctgact tgtactgtgt ctgggggttc aatcagcact tctggtatg
494948DNAArtificial SequencehBU12 dsFv primer P4 49ctgggtgctg
cctaatccag cctacaccca taccagaagt gctgattg 485050DNAArtificial
SequencehBU12 dsFv primer P5 50ggattaggca gcacccaggg aagtgtctgg
agtggattgg acacatttgg 505147DNAArtificial SequencehBU12 dsFv primer
P6 51aacagtaata gacagcaaca tcctctggct ccaggctgct gattgtg
475251DNAArtificial SequencehBU12 dsFv primer P7 52caagagatat
aacccagccc tgaagagcag agtgacaatc tctgtggata c 515350DNAArtificial
SequencehBU12 dsFv primer P8 53gacagcttga ggctaaactg gttcttggag
gtatccacag agattgtcac 505449DNAArtificial SequencehBU12 dsFv primer
P9 54gtttagcctc aagctgtcca gtgtgacagc tgcagatact gctgtctac
495548DNAArtificial SequencehBU12 dsFv primer P10 55aaacagtaat
agacagcaac atcctctggc tccaggctgc tgattgtg 485647DNAArtificial
SequencehBU12 dsFv primer P11 56ggaactttgg tcctactatt ttgactactg
gggccaaggc acccttg 475753DNAArtificial SequencehBU12 dsFv primer
P12 57gccccctgac ccgccacctc ctgaggagac tgtgacaagg gtgccttggc ccc
535853DNAArtificial SequencehBU12 dsFv primer P13 58ggtggatcgg
ggggtggcgg atctgaaatt gttctcaccc agtctccagc aac 535952DNAArtificial
SequencehBU12 dsFv primer P14 59cagggtagcc ctttcccctg gagagagaga
cagggttgct ggagactggg tg 526052DNAArtificial SequencehBU12 dsFv
primer P15 60ggggaaaggg ctaccctgag ctgcagtgcc agctcaagtg taagttacat
gc 526152DNAArtificial SequencehBU12 dsFv primer P16 61ctgggagcct
gccctggctt ctgctggtac cagtgcatgt aacttacact tg 526251DNAArtificial
SequencehBU12 dsFv primer P17 62gccagggcag gctcccagac tcctgattta
tgacacatcc aaactggctt c 516352DNAArtificial SequencehBU12 dsFv
primer P18 63ccagacccac tgccactgaa ccttgctgga ataccagaag ccagtttgga
tg 526450DNAArtificial SequencehBU12 dsFv primer P19 64cagtggcagt
gggtctggaa cagattttac actcacaatc agcagcctgg 506550DNAArtificial
SequencehBU12 dsFv primer P20 65gaaaacagta atagacagca acatcctctg
gctccaggct gctgattgtg 506648DNAArtificial SequencehBU12 dsFv primer
P21 66gctgtctatt actgttttca ggggagtgta tacccattca cttttggc
486749DNAArtificial SequencehBU12 dsFv primer P22 67acgcgttctt
ttgatttcca actttgtccc gcagccaaaa gtgaatggg 496850DNAArtificial
Sequence11F8 dsFv primer P23 68gtcgaccaat tgggaggtgg cggatcccag
gtgcagctgc aggagtcggg 506950DNAArtificial Sequence11F8 dsFv primer
P24 69acagggtctg tgaaggcttc accagtcctg ggcccgactc ctgcagctgc
507052DNAArtificial Sequence11F8 dsFv primer P25 70agccttcaca
gaccctgtcc ctcacctgca ctgtctctgg tggctccatc ag 527152DNAArtificial
Sequence11F8 dsFv primer P26 71ggcggatcca actccagtag taatcaccac
tgctgatgga gccaccagag ac 527250DNAArtificial Sequence11F8 dsFv
primer P27 72actggagttg gatccgccag cccccaggga agtgcctgga gtggattggg
507352DNAArtificial Sequence11F8 dsFv primer P28 73ggttgtagtc
ggtgctccca ctgtaataga tgtacccaat ccactccagg ca 527452DNAArtificial
Sequence11F8 dsFv primer P29 74tgggagcacc gactacaacc cgtccctcaa
gagtcgagtc accatgtccg ta 527550DNAArtificial Sequence11F8 dsFv
primer P30 75accttcaggg aaaactgatt cttggacgtg tctacggaca tggtgactcg
507650DNAArtificial Sequence11F8 dsFv primer P31 76tcagttttcc
ctgaaggtca actctgtgac cgccgcagac acggctgtgt 507753DNAArtificial
Sequence11F8 dsFv primer P32 77ccccactcca aaaatcgaca ctctcgcaca
gtaatacaca gccgtgtctg cgg 537850DNAArtificial Sequence11F8 dsFv
primer P33 78tcgatttttg gagtggggac atttgactac tggggccagg gcaccctggt
507954DNAArtificial Sequence11F8 dsFv primer P34 79accgccccct
gacccgccac ctccgcttga gacggtgacc agggtgccct ggcc
548056DNAArtificial Sequence11F8 dsFv primer P35 80ggatcggggg
gtggcggatc tgaaattgtg atgacacagt ctccagccac cctgtc
568152DNAArtificial Sequence11F8 dsFv primer P36 81gcaggagagg
gtggctcttt cccctggaga caaagacagg gtggctggag ac 528248DNAArtificial
Sequence11F8 dsFv primer P37 82agagccaccc tctcctgcag ggccagtcag
agtgttagca gctactta 488348DNAArtificial Sequence11F8 dsFv primer
P38 83agcctggcca ggtttctgtt ggtaccaggc taagtagctg ctaacact
488450DNAArtificial Sequence11F8 dsFv primer P39 84cagaaacctg
gccaggctcc caggctcctc atctatgatg catccaacag 508552DNAArtificial
Sequence11F8 dsFv primer P40 85actgccactg aacctggctg ggatgccagt
ggccctgttg gatgcatcat ag 528650DNAArtificial Sequence11F8 dsFv
primer P41 86gccaggttca gtggcagtgg gtctgggaca gacttcactc tcaccatcag
508750DNAArtificial Sequence11F8 dsFv primer P42 87aatacactgc
aaaatcttca ggctctaggc tgctgatggt gagagtgaag 508852DNAArtificial
Sequence11F8 dsFv primer P43 88gaagattttg cagtgtatta ctgtcaccag
tatggtagca cacctctcac tt 528952DNAArtificial Sequence11F8 dsFv
primer P44 89acgcgttttg atctccgcct tggtcccgca gccgaaagtg agaggtgtgc
ta 529053DNAArtificial Sequencelinker-Mfel-hBU12 VH primer F
90gtggtggttc aggacaattg ggaggtggcg gatcccaggt tcagctgcaa gag
539152DNAArtificial SequenceSall-linker-Mfel-hBU12 VH primer R
91gtcgacctgg tcaccgtctc ctcagcctcc accggtggtg gttcaggaca at
529253DNAArtificial SequencehBU12VL-Mlu l-6xHis-Clal primer R
92atcgatttaa tgatgatgat gatgatgacg cgttcttttg atttccaact ttg
539339DNAArtificial SequenceMfeI-h11F8 VH primer F 93caattgggag
gtggcggatc ccaggtgcag ctgcaggag 399425DNAArtificial
SequenceMluI-11F8VL primer R 94acgcgttttg atctccgcct tggtc
259539DNAArtificial SequenceMfeI-DNSVH primer F 95caattgggag
gtggcggatc cagtgaagtg aagcttgag 399624DNAArtificial
SequenceMluI-DNSVL primer R 96acgcgtccgt tttatttcca actt
249739DNAArtificial SequenceMfeI-hCC49VH primer F 97caattgggag
gtggcggatc ccaggtgcag ctggtgcag 399825DNAArtificial
SequenceMluI-hCC49VL primer R 98acgcgttttg atctccacct tggtc
259924DNAArtificial Sequence15-2b- knob VL-Ck primer F 99agatctgaca
tccagatgac ccag 2410036DNAArtificial Sequence15-2b- knob VL-Ck
primer R 100tatcgatgtt taaacctagc actctcccct gttgaa
3610131DNAArtificial Sequence15-2b- knob VH-CH1 primer F
101ggcccagccg gccgaggtgc agctggtgga g 3110220DNAArtificial
Sequence15-2b- knob VH-CH1 primer R 102aagttttttg tcgaccgtgg
2010327DNAArtificial Sequence15-2b- knob huIgG1-upper hinge primer
F 103gacaaaactc acacatgccc accgtgc 2710436DNAArtificial
Sequence15-2b- knob CH1-partical hinge primer R 104gcatgtgtga
gttttgtcac aagatttggg ctcaac 3610524DNAArtificial Sequence15-2b-
knob CH3 primer R 105ctcgagttta
cccggagaca ggga 2410628DNAArtificial Sequenceh6-3- knob104 VL-Ck
primer F 106agatctgaca tcgtgatgac ccagtctc 2810736DNAArtificial
Sequenceh6-3- knob104 VL-Ck primer R 107tatcgatgtt taaacctagc
actctcccct gttgaa 3610819DNAArtificial Sequenceh6-3- knob104 VH-CH1
primer F 108caggtgcagc tggtgcaat 1910920DNAArtificial Sequenceh6-3-
knob104 VH-CH1 primer R 109aactctcttg tccaccttgg
2011032DNAArtificial SequenceBU12-hole VH-Ck-partial hinge primer F
110agccggccca ggttcagctg caagagtctg gc 3211139DNAArtificial
SequenceBU12-hole VH-Ck-partial hinge primer R 111gcatgtgtga
gttttgtcac actctcccct gttgaagct 3911223DNAArtificial
SequenceBU12-hole VL-partial VH-CH1-upper Hinge primer F
112gaaattgttc tcacccagtc tcc 2311321DNAArtificial SequenceBU12-hole
VL-partial VH-CH1-upper Hinge primer R 113ttaacaagat ttgggctcaa c
2111443DNAArtificial SequenceBU12-hole partial VL-hCH1 primer F
114gttggaaatc aaaagatcct cagcctccac caagggccca tcg
4311537DNAArtificial Sequencea-Her2-hole VH primer 115ggcccagccg
gcccaggtgc agctgttgca gtctggg 3711623DNAArtificial
Sequencea-Her2-hole VH-Ck-partial hinge primer F 116aggtgcagct
gttgcagtct ggg 2311739DNAArtificial Sequencea-Her2-hole
VH-Ck-partial hinge primer R 117gcatgtgtga gttttgtcac actctcccct
gttgaagct 3911842DNAArtificial Sequencea-Her2-hole VL-partial VH
primer F 118ctgaccgtcc taggttcctc agcctccacc aagggcccat cg
4211945DNAArtificial Sequencea-Her2-hole VL-partial VH primer R
119acctaggacg gtcagcttgg tcccgccgcc gaacacccag cccga
4512029DNAArtificial Sequencea-Her2-hole VL primer F 120ctgccagatc
tcagtctgtg ttgacgcag 2912145DNAArtificial Sequencea-Her2-hole VL
primer R 121acctaggacg gtcagcttgg tcccgccgcc gaacacccag cccga
4512221DNAArtificial Sequencea-Her2-hole CH1-upper hinge primer R
122ttaacaagat ttgggctcaa c 2112347DNAArtificial Sequenceh15-2b
Bgl-VH-1 primer F 123gaagatctga ggtgcagctg gtggagtctg ggggaggctt
ggtccag 4712451DNAArtificial Sequenceh15-2b VH-2 primer R
124agaggctgca caggagagtt tcagggaccc cccaggctgg accaagcctc c
5112551DNAArtificial Sequenceh15-2b VH-3 primer F 125tcctgtgcag
cctctgggtt caccttcagt aactactgga tgaactgggt c 5112651DNAArtificial
Sequenceh15-2b VH-4 primer R 126gccaacccac tccagccctt tcccggaagc
ctggcggacc cagttcatcc a 5112751DNAArtificial Sequenceh15-2b VH-5
primer F 127ctggagtggg ttggcgaaat tagatcgaaa tctaataatt atgcgacaca
t 5112851DNAArtificial Sequenceh15-2b VH-6 primer R 128ggagatggtg
aacctccctt tcacagactc cgcataatgt gtcgcataat t 5112951DNAArtificial
Sequenceh15-2b VH-7 primer F 129aggttcacca tctccagaga tgattcaaag
aacacggcgt atctgcaaat g 5113051DNAArtificial Sequenceh15-2b VH-8
130gtaatacacg gccgtgtcct cggttttcag gctgttcatt tgcagatacg c
5113151DNAArtificial Sequenceh15-2b VH-9 (T93S) primer F
131acggccgtgt attactgttc caacagatac tactggggcc aaggaaccct g
5113245DNAArtificial Sequenceh15-2b VH-10 primer R 132acctttggtg
gaggctgagg agacggtgac cagggttcct tggcc 4513328DNAArtificial
Sequencehomo 3' IgG2 CH1-SalI primer R 133acgcgtcgac tttgcgctca
actgtctt 2813443DNAArtificial Sequenceh15-2b sfi-VL-1 primer F
134tgctggggcc cagccggccg acatccagat gacccagtct cca
4313551DNAArtificial Sequenceh15-2b VL-2 primer R 135ggtgactctg
tctcctacag atgcagacag ggaggatgga gactgggtca t 5113651DNAArtificial
Sequenceh15-2b VL-3 primer F 136ggagacagag tcaccatcac ttgcaaggcc
agtcaggatg taaatacttc t 5113751DNAArtificial Sequenceh15-2b VL-4
primer R 137aggggctttc cctggtttct gctgatacca ggctacagaa gtatttacat
c 5113851DNAArtificial Sequenceh15-2b VL-5 primer F 138ccagggaaag
cccctaagct cctgatctac tgggcatcca cccggcacac t 5113951DNAArtificial
Sequenceh15-2b VL-6 primer R 139cccagatcca cttccactga accttgatgg
gaccccagtg tgccgggtgg a 5114051DNAArtificial Sequenceh15-2b VL-7
primer F 140ggaagtggat ctgggacaga ttttactttc accatcagca gcctgcagcc
t 5114151DNAArtificial Sequenceh15-2b VL-8 primer R 141gatatattgc
agacagtaat atgttgcaat atcttcaggc tgcaggctgc t 5114251DNAArtificial
Sequenceh15-2b VL-9 primer F 142tgtctgcaat atatcaacta tccgtacacg
tttggccagg ggaccaagct g 5114342DNAArtificial Sequenceh15-2b VL-10
primer R 143tggtgcagcc acagtccgtt tgatctccag cttggtcccc tg
4214457DNAArtificial Sequencehomo 3' Ck cys-XhoI primer R
144ccgctcgagg cactctcccc tgttgaagct ctttgtgacg ggcgagctca ggccctg
5714554DNAArtificial Sequenceh528VH01 primer F 145caggtgcaac
tggttcagag cggcgcggaa gtgaaaaagc cgggcgcgtc ggtt
5414648DNAArtificial Sequenceh528VH02 primer R 146aaaggtatag
cctgaggctt tgcagctcac tttaaccgac gcgcccgg 4814748DNAArtificial
Sequenceh528VH03 primer F 147tcaggctata cctttacgag ctactggatg
cattgggtgc gccaggcc 4814848DNAArtificial Sequenceh528VH04 primer R
148aatgttaccc atccattcca ggccctgacc cggggcctgg cgcaccca
4814948DNAArtificial Sequenceh528VH05 primer F 149tggatgggta
acatttatcc gggcagcggt ggcaccaact atgcggaa 4815048DNAArtificial
Sequenceh528VH06 primer R 150atcacgcgtc atggtcacgc ggttcttaaa
tttttccgca tagttggt 4815148DNAArtificial Sequenceh528VH07 primer F
151accatgacgc gtgataccag catttcgacg gcctatatgg aactgagc
4815248DNAArtificial Sequenceh528VH08 primer R 152gtaatacacg
gcggtgtcat cgctacgcag gcggctcagt tccatata 4815348DNAArtificial
Sequenceh528VH09 primer F 153accgccgtgt attactgcgc gcgcagtggc
ggtccgtatt ttttcgat 4815451DNAArtificial Sequenceh528VH10 primer R
154cgagctcacg gtaaccagcg taccctggcc ccagtaatcg aaaaaatacg g
5115545DNAArtificial Sequence(G4S)2-ahEGFR VH primer F
155ggcggtggtg ggtcgggtgg cggcggatct caggtgcaac tggtt
4515626DNAArtificial SequenceahEGFR VH-stop-Cal primer R
156ccatcgattt acgagctcac ggtaac 2615748DNAArtificial
Sequenceh528VL01 primer F 157gatattgtga tgacccagag cccgctgagc
ctgccggtga ccccaggc 4815848DNAArtificial Sequenceh528VL02 primer R
158ctgcgagctg cggcagctaa tcgacgccgg ttcgcctggg gtcaccgg
4815948DNAArtificial Sequenceh528VL03 primer F 159tgccgcagct
cgcagaacat cgtgcataat aacggcatta cctatctg 4816048DNAArtificial
Sequenceh528VL04 primer R 160cgggctttgg cccggtttct gcagatacca
ttccagatag gtaatgcc 4816148DNAArtificial Sequenceh528VL05 primer F
161ccgggccaaa gcccgcagct gttaatttat aaagtgagcg atcgcttt
4816248DNAArtificial Sequenceh528VL06 primer R 162accgctgccc
gaaaagcgat ccggcacgcc gctaaagcga tcgctcac 4816348DNAArtificial
Sequenceh528VL07 primer F 163ttttcgggca gcggtagtgg caccgatttt
acgctgaaaa ttagccgc 4816448DNAArtificial Sequenceh528VL08 primer R
164gcagtaatac acgccaacat cctccgcttc cacgcggcta attttcag
4816548DNAArtificial Sequenceh528VL09 primer F 165ggcgtgtatt
actgctttca gggcagccat atcccgccaa cctttggc 4816645DNAArtificial
Sequenceh528VL10 primer R 166cgcgcgttta atttccactt tggtgccttg
gccaaaggtt ggcgg 4516724DNAArtificial SequenceMfe-ahEGFR VL primer
F 167caattggata ttgtgatgac ccag 2416845DNAArtificial SequenceahEGFR
VL-(G4S)2 primer R 168cgacccacca ccgcccgagc caccgccacc cgcgcgttta
atttc 4516958DNAArtificial SequenceSal-G-myc-G4S primer F
169acgcgtcgac ggggaacaaa aactcatctc agaagaggat ctgggaggcg gtggcagt
5817057DNAArtificial SequenceG2S-G4SX2-MfeI primer F 170ggtggcagtg
gtggtggtgg atcaggaggt ggcggatccc aattgcaggt gcagctg
5717127DNAArtificial SequenceHer2 scFv-stop-ClaI primer R
171atcgattcaa cctaggacgg tcagctt 2717230DNAArtificial
Sequencemfe1-h15-2bVL primer F 172caattggaca tccagatgac ccagtctcca
3017335DNAArtificial Sequenceh15-2bscFv-ClaI-Sbfl primer R
173cccctgcagg catcgattta tgaggagacg gtgac 3517443DNAArtificial
SequenceNaeI-X+hHB12bVL-1 primer F 174gccggccgag atcgtgctga
cccagagccc cgacttccag agc 4317553DNAArtificial SequencehHB12bVL-2
primer R 175ctctgcaggt gatggtcacc ttctccttgg gggtcacgct ctggaagtcg
ggg 5317654DNAArtificial SequencehHB12bVL-3 primer F 176gtgaccatca
cctgcagagc cagcgagagc gtggacacct tcggcatcag cttc
5417752DNAArtificial SequencehHB12bVL-4 priemr R 177gctctggtcg
ggcttctgct ggaaccagtt catgaagctg atgccgaagg tg 5217851DNAArtificial
SequencehHB12bVL-5 primer F 178gaagcccgac cagagcccca agctgctgat
ccacgccgcc agcaaccagg g 5117951DNAArtificial SequencehHB12bVL-6
primer R 179cttccgctgc cgctgaatct gctgggcacg ccgctgccct ggttgctggc
g 5118052DNAArtificial SequencehHB12bVL-7 primer F 180ttcagcggca
gcggaagcgg caccgacttc accctgacca tcaacagcct gg 5218154DNAArtificial
SequencehHB12bVL-8 primer R 181ctctgctggc agtagtaggt tgctgcgtcc
tcggcctcca ggctgttgat ggtc 5418252DNAArtificial SequencehHB12bVL-9
primer F 182aacctactac tgccagcaga gcaaggaggt gcccttcacc ttcggcggcg
gc 5218352DNAArtificial SequenceDraIII+hHB12bVL-10 primer R
183gacactcggt gcagccacag tcttgatctc caccttggtg ccgccgccga ag
5218444DNAArtificial SequenceHpaI+hHB12bVH-1 primer F 184gttaacgagg
tgcagctggt ggagagcggc ggcggcctgg tgca 4418554DNAArtificial
SequencehHB12bVH-2 primer R 185cgctggcggc gcagctcagt ctcaggctgc
cgccgggctg caccaggccg ccgc 5418654DNAArtificial SequencehHB12bVH-3
primer F 186gctgcgccgc cagcggcttc accttcagca gcagctggat gaactgggtg
agac 5418756DNAArtificial SequencehHB12bVH-4 primer R 187gattctgccc
acccactcca ggcccttgcc gggggcctgt ctcacccagt tcatcc
5618857DNAArtificial SequencehHB12bVH-5 primrer F 188gagtgggtgg
gcagaatcta ccccggcgac ggcgacacca actacaacgg caagttc
5718956DNAArtificial SequencehHB12bVH-6 primer R 189tcttgctgtc
gtctctgctg atggtgaatc tgcccttgaa cttgccgttg tagttg
5619052DNAArtificial SequencehHB12bVH-7 primer F 190ttcagcggca
gcggaagcgg caccgacttc accctgacca tcaacagcct gg 5219156DNAArtificial
SequencehHB12bVH-8 primer R 191atgaagccgc ttctggcgca gtagtacacg
gcggtgtcct cggtcttcag gctgtt 5619255DNAArtificial
SequencehHB12bVH-9 primer F 192cgccagaagc ggcttcatca ccaccgtgct
ggacttcgac tactggggcc agggc 5519355DNAArtificial
SequenceApaI+hHB12bVH-10 primer R 193gggccctttg gtggaggcgc
tgctcacggt caccagggtg ccctggcccc agtag 5519450DNAArtificial
SequenceNaeI-X-aCD20VL-1 primer F 194gccggccatg gaagccccag
ctcagcttct cttcctcctg ctactctggc 5019562DNAArtificial
SequenceaCD20VL-2 primer R 195ctggagactg tgtcaacaca atttctccgg
tggtatctgg gagccagagt agcaggagga 60ag 6219659DNAArtificial
SequenceaCD20VL-3 primer F 196aattgtgttg acacagtctc cagccaccct
gtctttgtct ccaggggaaa gagccaccc 5919759DNAArtificial
SequenceaCD20VL-4 primer R 197caggctaagt agctgctaac actctgactg
gccctgcagg agagggtggc tctttcccc 5919858DNAArtificial
SequenceaCD20VL-5 primer F 198tgttagcagc tacttagcct ggtaccaaca
gaaacctggc caggctccca ggctcctc 5819957DNAArtificial
SequenceaCD20VL-6 primer R 199ctggctggga tgccagtggc cctgttggat
gcatcataga tgaggagcct gggagcc 5720059DNAArtificial
SequenceaCD20VL-7 primer F 200actggcatcc cagccaggtt cagtggcagt
gggtctggga cagacttcac tctcaccat 5920165DNAArtificial
SequenceaCD20VL-8 primer R 201ctgacagtaa taaactgcaa aatcttcagg
ctctaggctg ctgatggtga gagtgaagtc 60tgtcc 6520262DNAArtificial
SequenceaCD20VL-9 primer F 202gaagattttg cagtttatta ctgtcagcag
cgtagcaact ggccgatcac cttcggccaa 60gg 6220358DNAArtificial
SequenceDraIII-aCD20VL-10 primer R 203gacactcggt gcagccacag
ttttaatctc cagtcgtgtc ccttggccga aggtgatc 5820448DNAArtificial
SequenceHpaI+aCD20VH-1 primer F 204gttaacatgg agttgggact gagctggatt
ttccttttgg ctatttta 4820559DNAArtificial SequenceaCD20VH-2 primer R
205ctccaccagc tgcacttcac actggacacc ttttaaaata gccaaaagga aaatccagc
5920654DNAArtificial SequenceaCD20VhH-3 primer F 206gaagtgcagc
tggtggagtc tgggggaggc ttggtacagc ctggcaggtc cctg
5420758DNAArtificial SequenceaCD20VH-4 primer R 207cataatcatt
aaaggtgaat ccagaggctg cacaggagag tctcagggac ctgccagg
5820860DNAArtificial SequenceaCD20VH-5 primer F 208gcctctggat
tcacctttaa tgattatgcc atgcactggg tccggcaagc tccagggaag
6020959DNAArtificial SequenceaCD20VH-6 primer R 209ggaaccacta
ttccaactaa tagttgagac ccactccagg cccttccctg gagcttgcc
5921060DNAArtificial SequenceaCD20VH-7 primer F 210tcaactatta
gttggaatag tggttccata ggctatgcgg actctgtgaa gggccgattc
6021156DNAArtificial SequenceaCD20VH-8 primer R 211gatacaggga
cttcttggcg ttgtctctgg agatggtgaa tcggcccttc acagag
5621255DNAArtificial SequenceaCD20VH-9 primer F 212cgccaagaag
tccctgtatc tgcaaatgaa cagtctgaga gctgaggaca cggcc
5521357DNAArtificial SequenceaCD20VH-10 primer R 213gtagtagttg
ccgtactgta tatcttttgc acagtaatac aaggccgtgt cctcagc
5721457DNAArtificial SequenceaCD20VH-11 primer F 214agatatacag
tacggcaact actactacgg tatggacgtc tggggccaag ggaccac
5721549DNAArtificial SequenceApaI-aCD20VH-12 primer R 215gggccctttg
gtggaggctg aggagacggt gaccgtggtc ccttggccc 4921612PRTArtificial
SequenceHumanized 6.3 VL 216Gln Ser Val Leu Tyr Ser Ser Asn Gln Met
Asn Tyr 1 5 10 2178PRTArtificial SequenceHumanized 6.3 VL 217Leu
Gln Tyr Leu Ser Ser Trp Thr 1 5 2188PRTArtificial SequenceHumanized
6.3 VH 218Gly Tyr Thr Phe Lys Asn Tyr Gly 1 5 2198PRTArtificial
SequenceHumanized 6.3 VH 219Ile Asn Thr Tyr Thr Gly Gln Pro 1 5
2207PRTArtificial SequenceHumanized 6.3 VH 220Ala Arg Asp Trp Gly
Pro Tyr 1 5 22111PRTArtificial SequenceHumanized 15-2b VL 221Lys
Ala Ser Gln Asp Val Asn Thr Ser Val Ala 1 5 10 2227PRTArtificial
SequenceHumanized 15-2b VL 222Trp Ala Ser Thr Arg His Thr 1 5
2239PRTArtificial SequenceHumanized 15-2b VL 223Leu Gln Tyr Ile Asn
Tyr Pro Tyr Thr 1 5 22410PRTArtificial SequenceHumanized 15-2b VH
224Gly Phe Thr Phe Ser Asn Tyr Trp Met Asn 1 5 10
22519PRTArtificial SequenceHumanized 15-2b VH 225Glu Ile Arg Ser
Lys Ser Asn Asn Tyr Ala Thr His Tyr Ala Glu Ser 1 5 10 15 Val Lys
Gly 2266PRTArtificial SequenceHumanized 15-2b VH 226Ser Asn Arg Tyr
Tyr Trp 1 5
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021
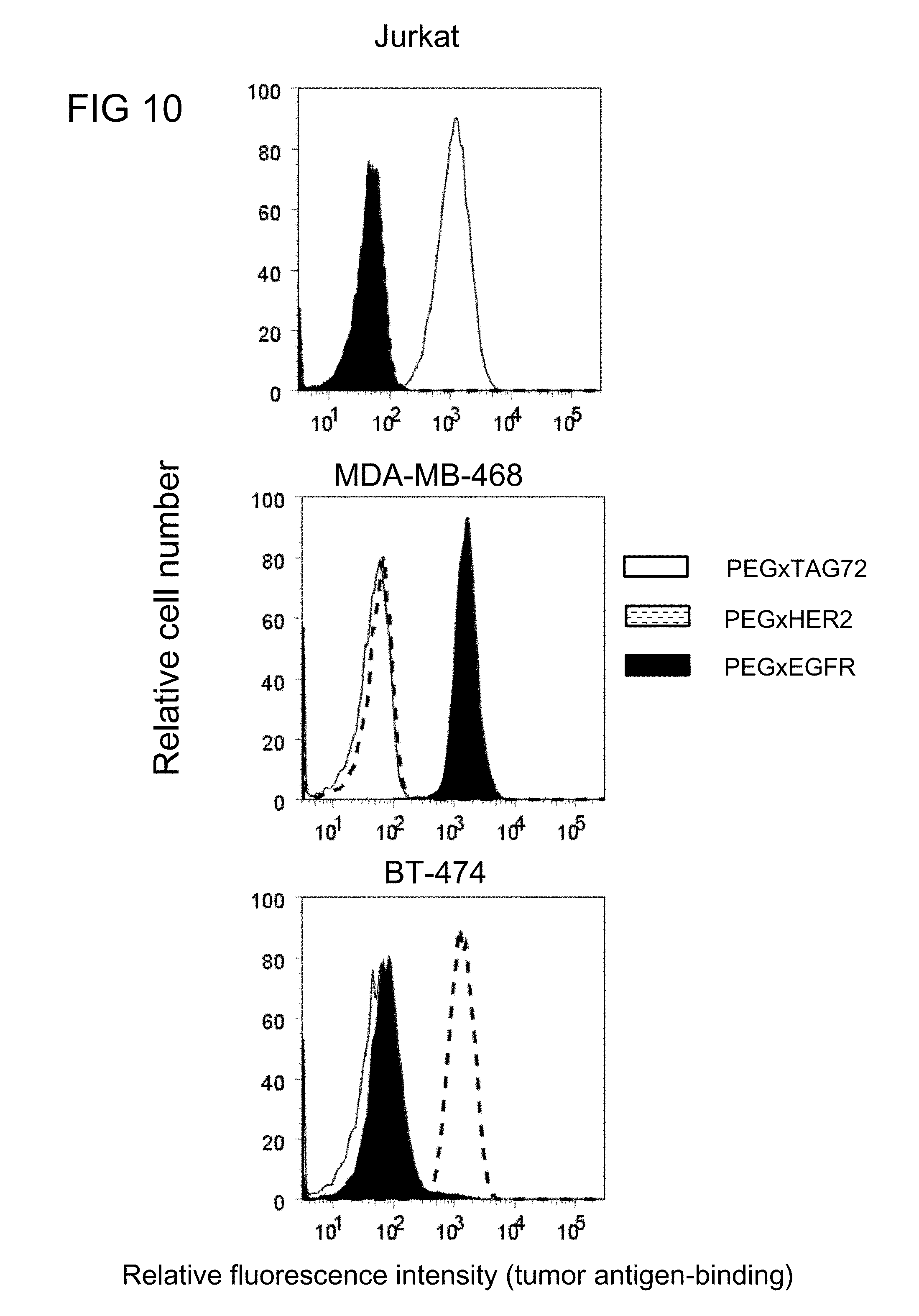
D00022

D00023

D00024

D00025
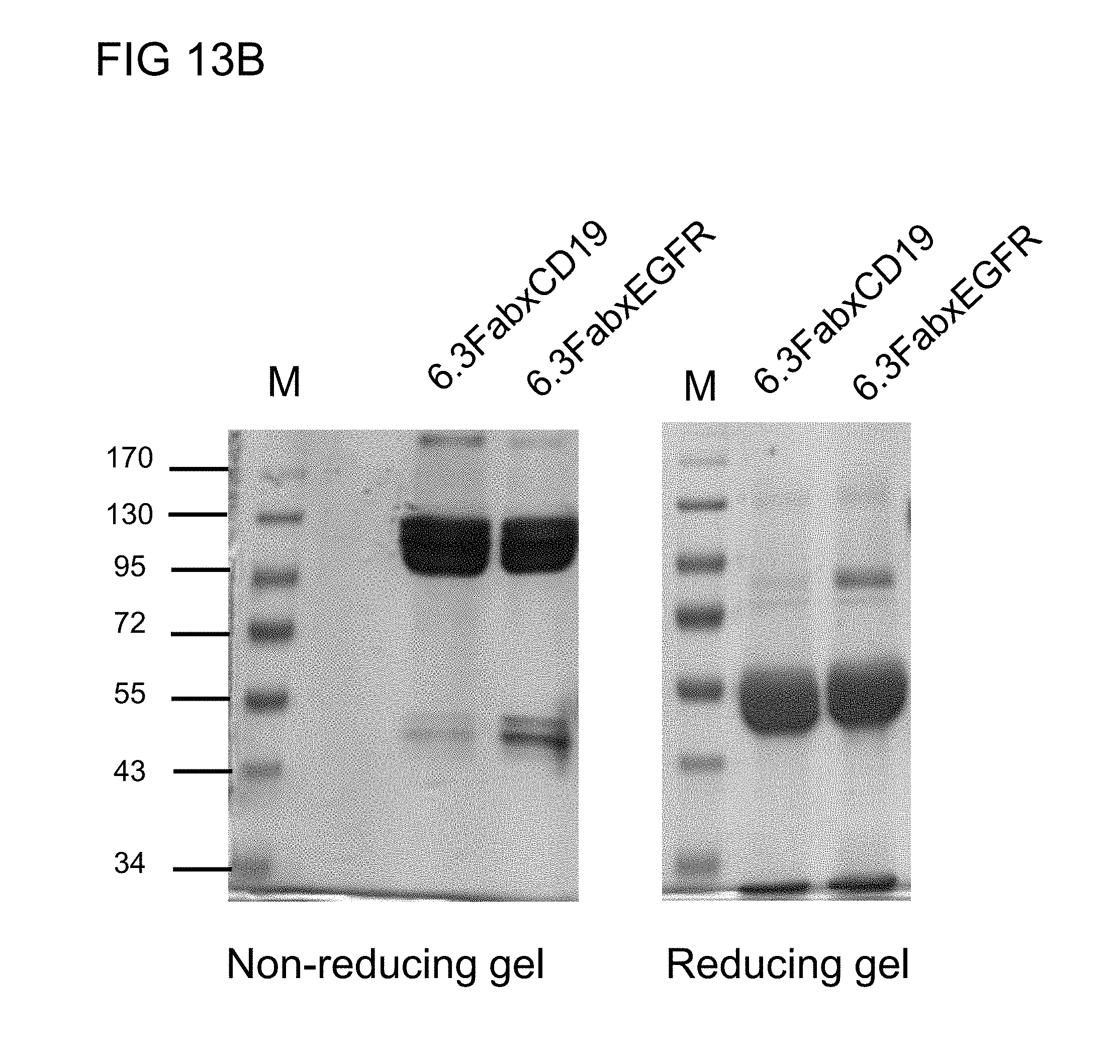
D00026

D00027

D00028

D00029
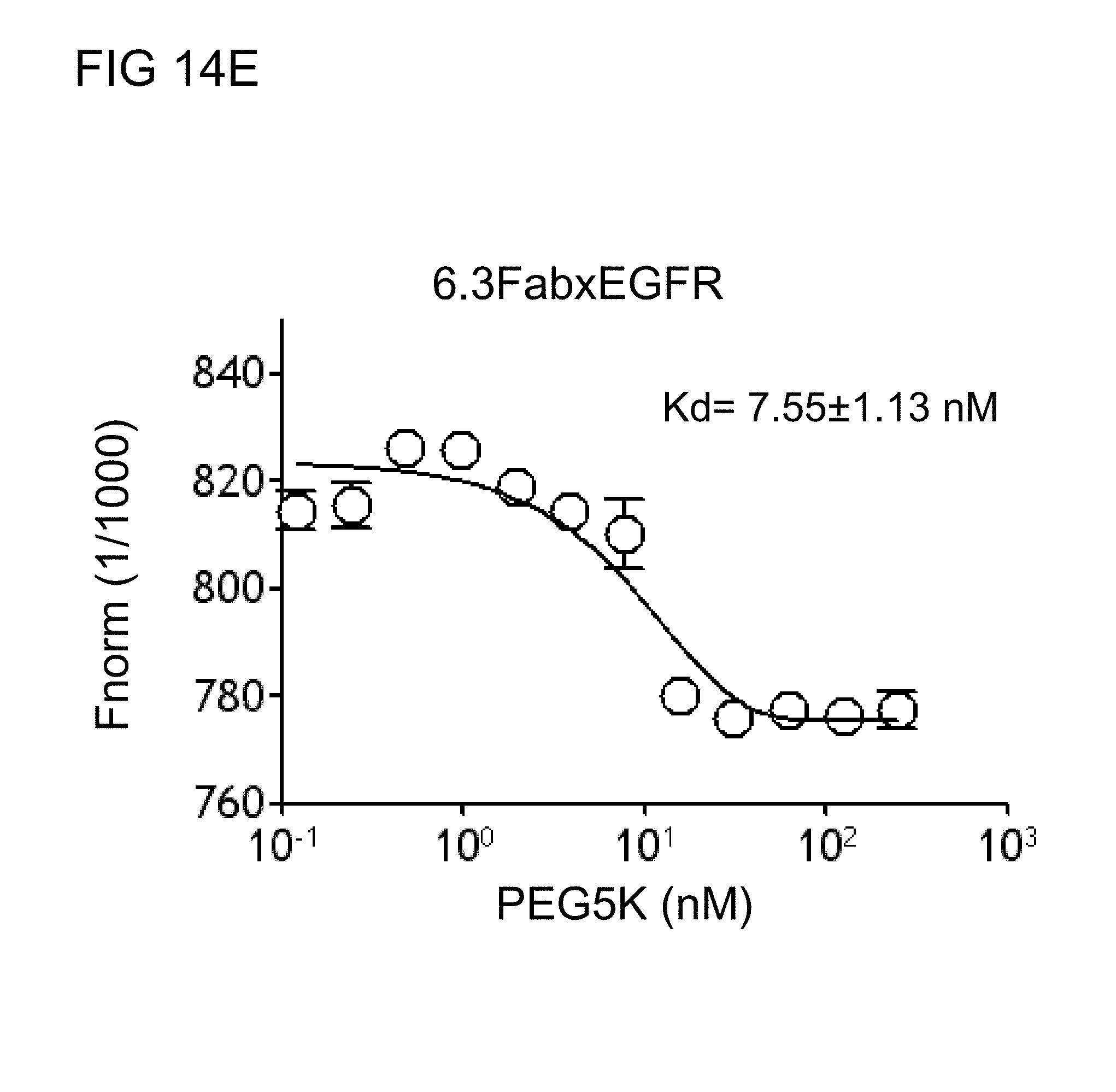
D00030

D00031

D00032
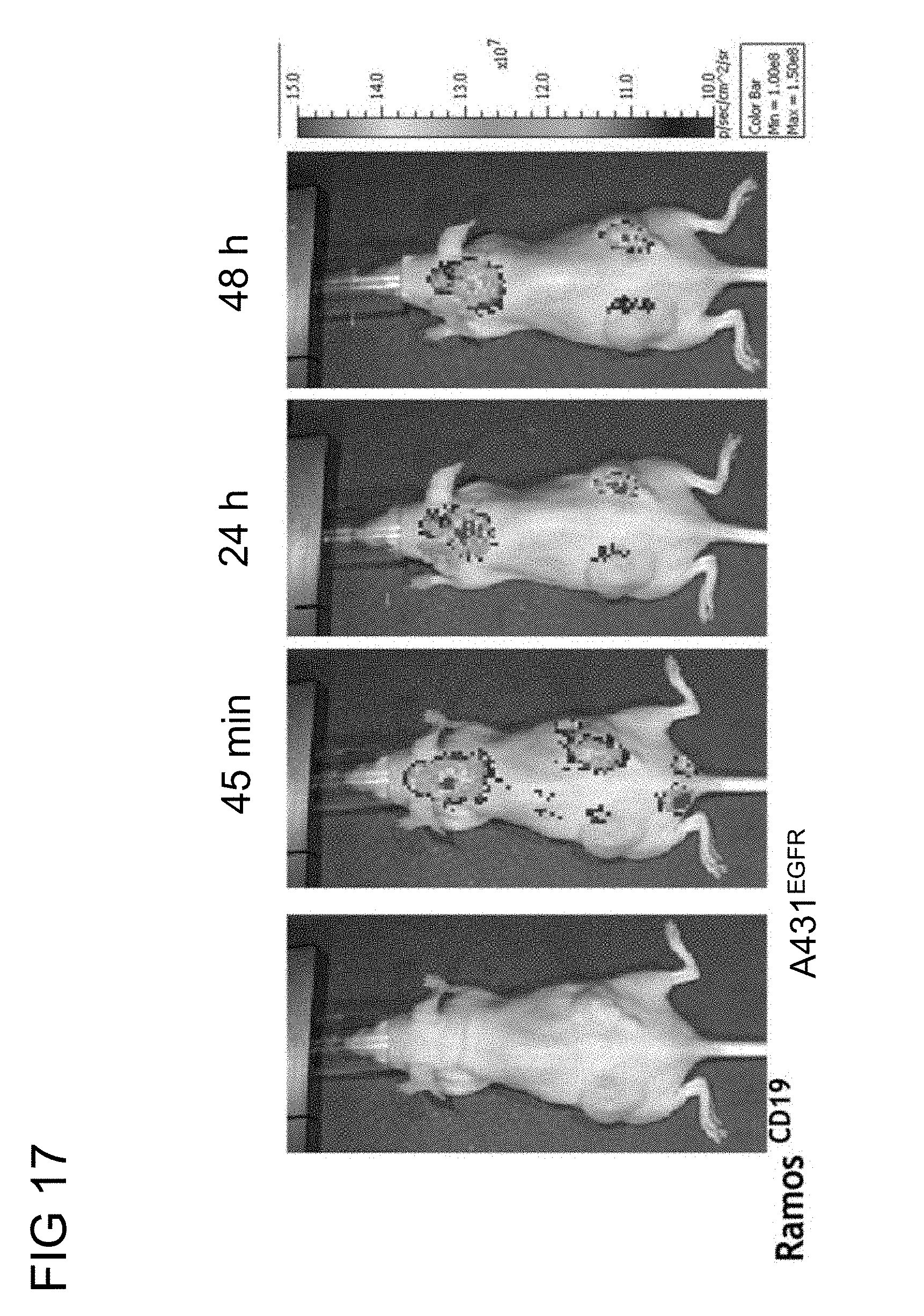
D00033

D00034
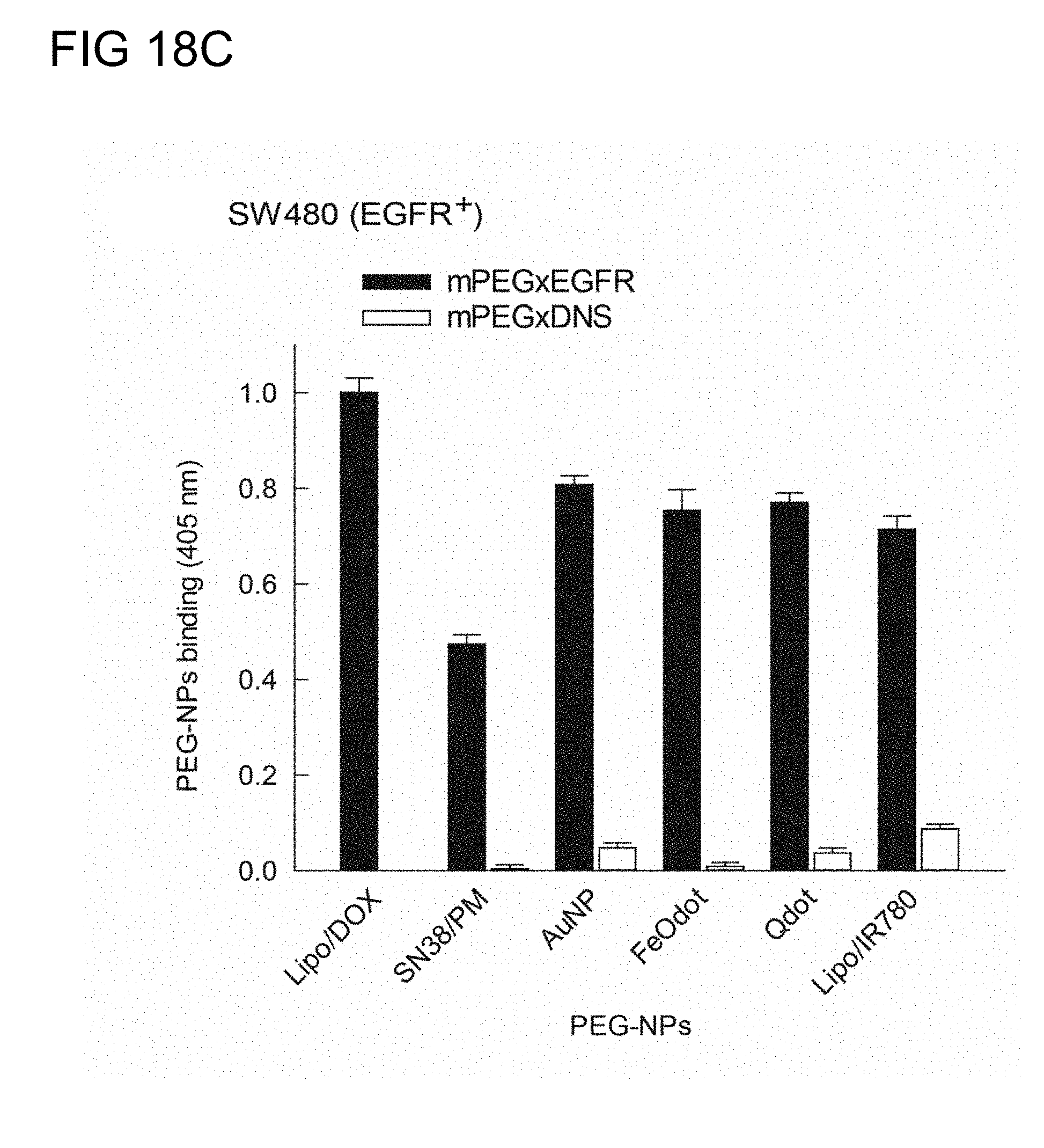
D00035

D00036

D00037

D00038
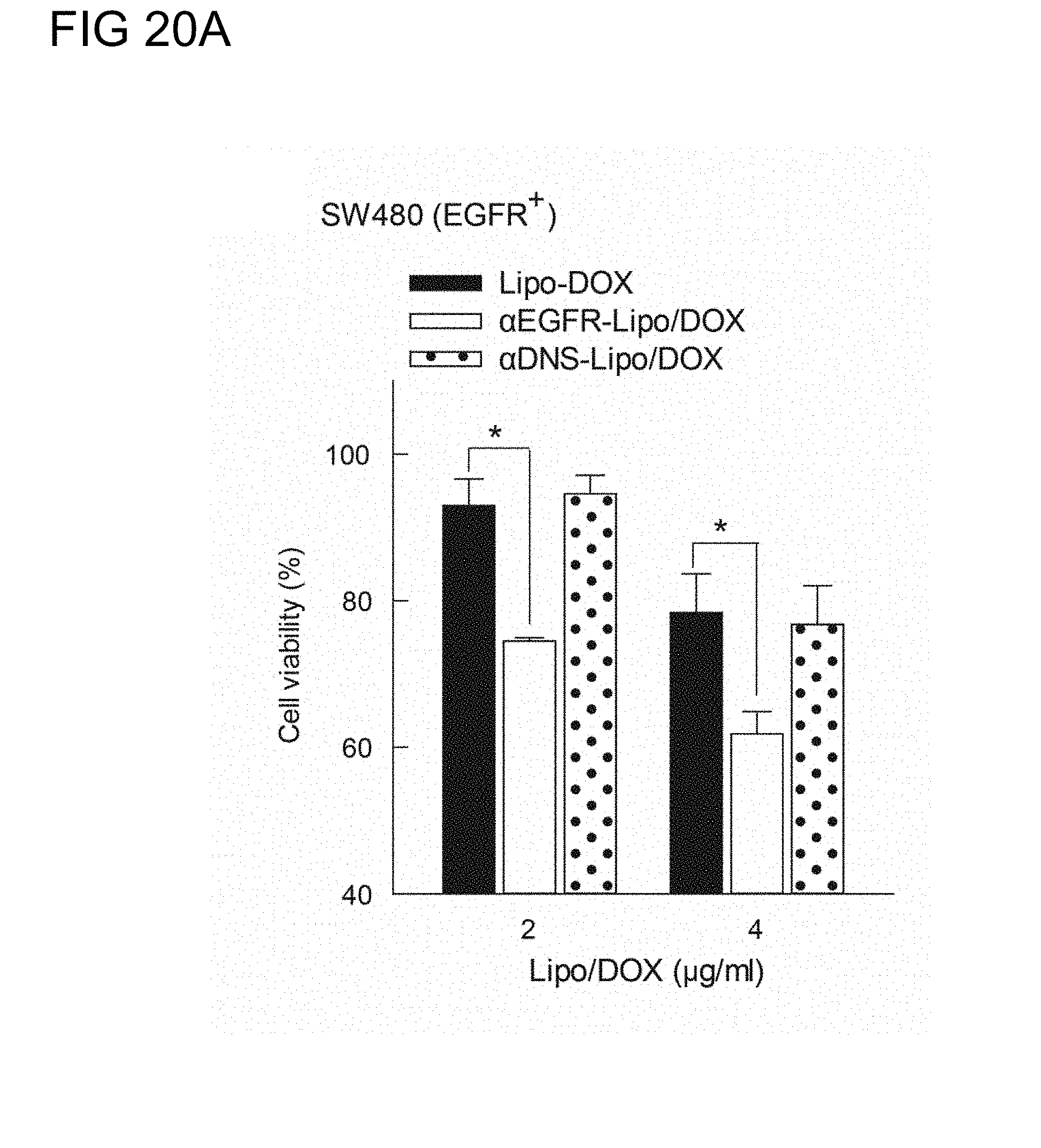
D00039

D00040

D00041

D00042

D00043

D00044

D00045

D00046
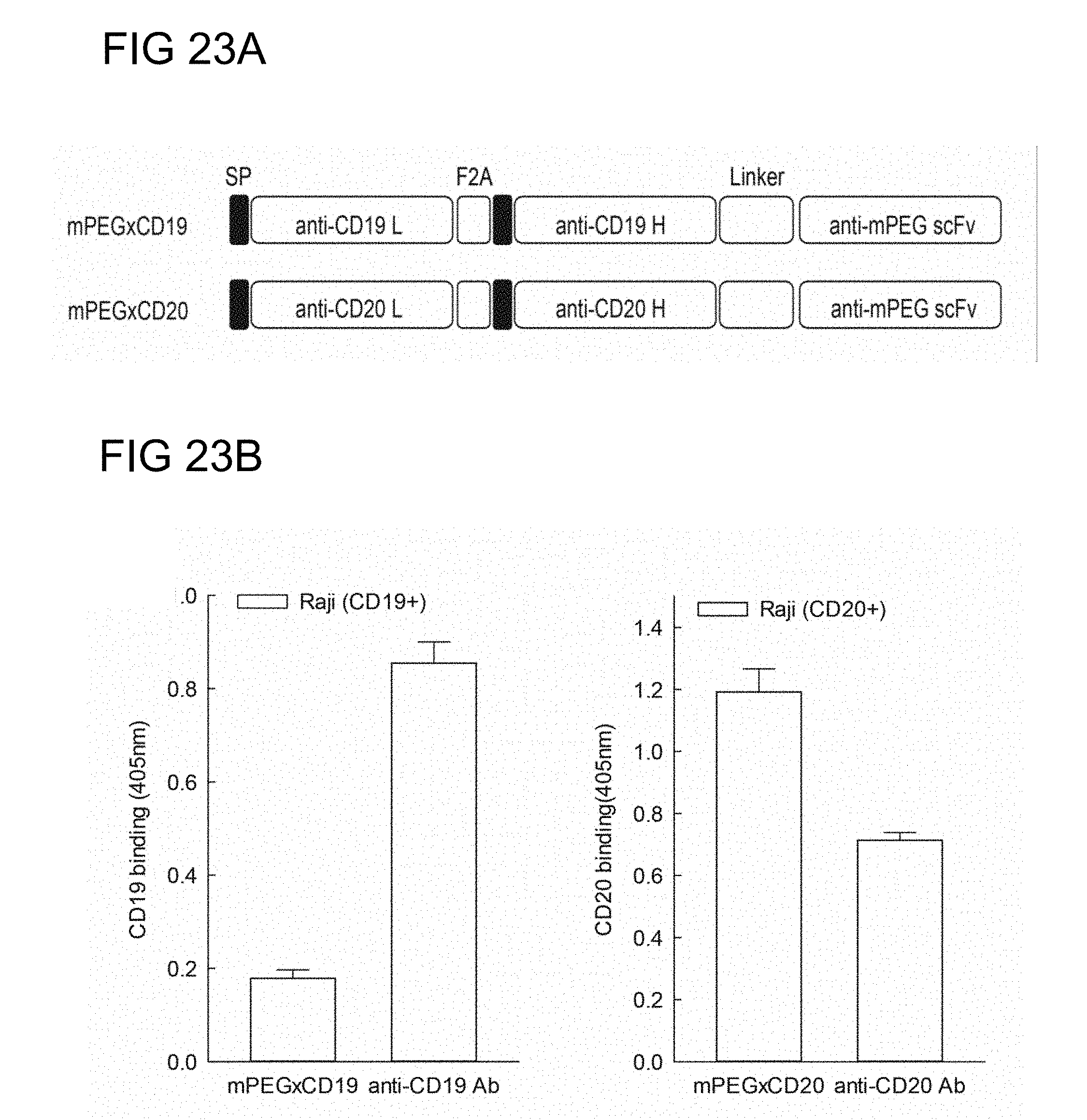
D00047

D00048

D00049

D00050

D00051

D00052
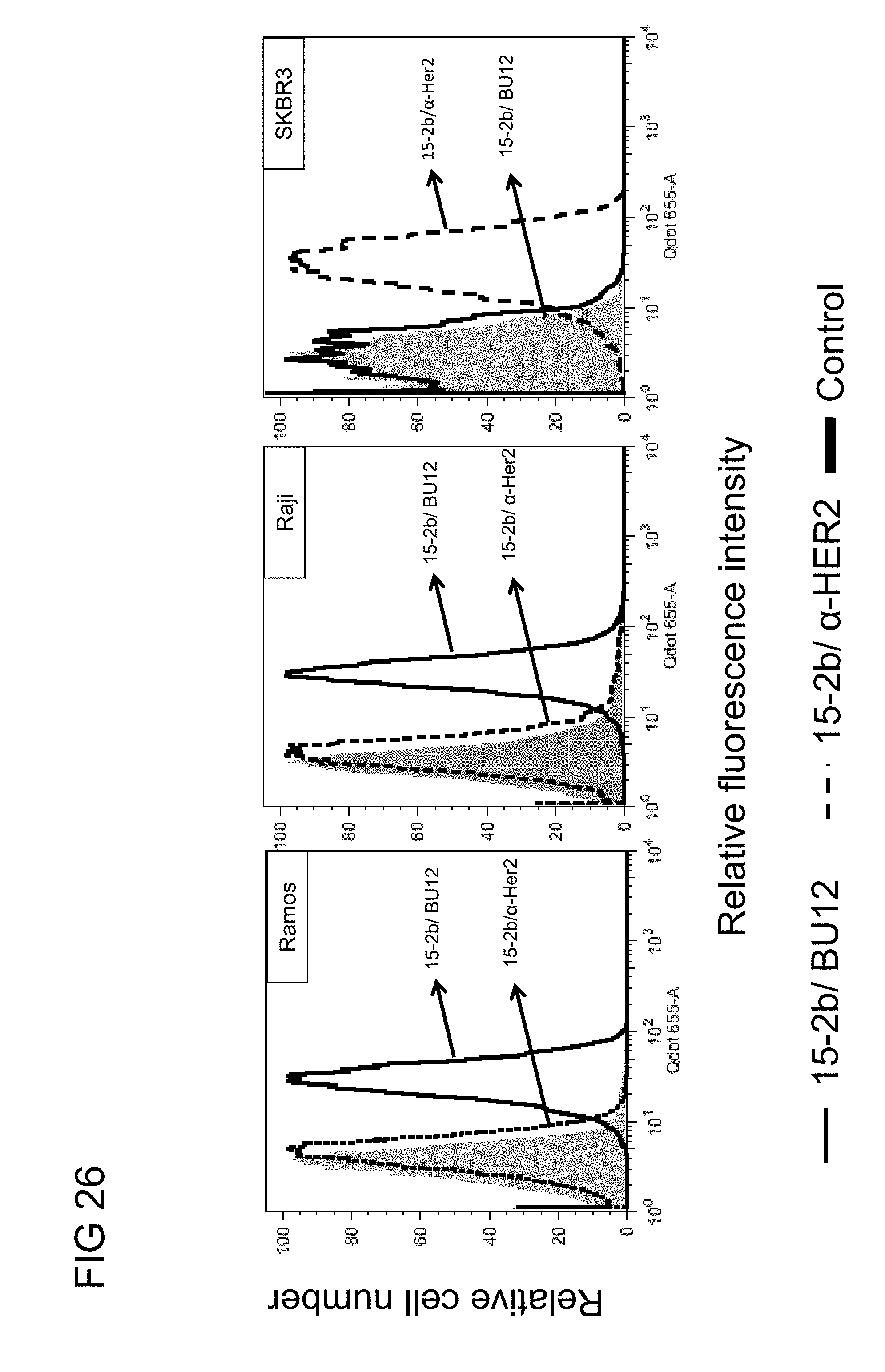
D00053
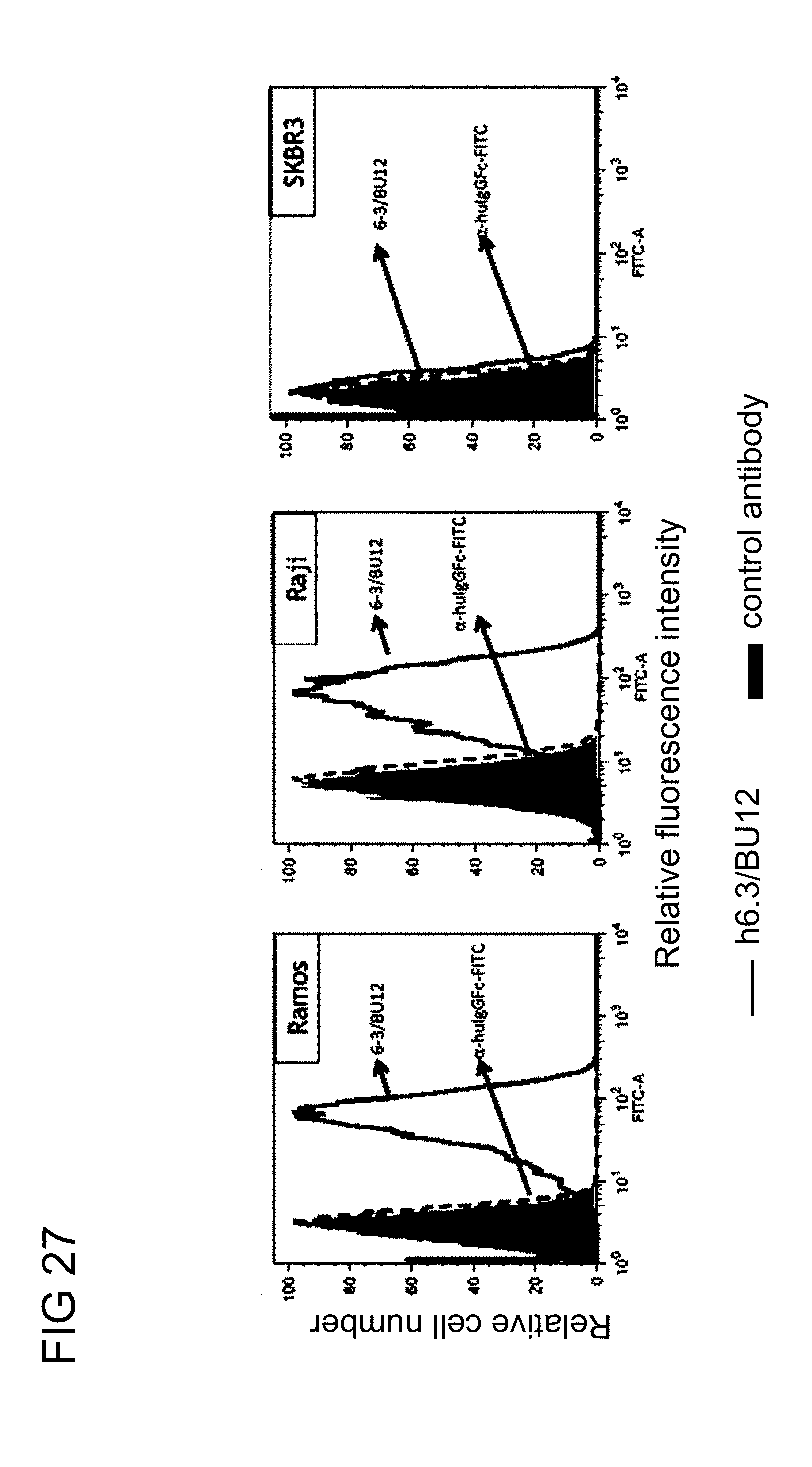
D00054

D00055

D00056

D00057

D00058

D00059

D00060

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.