Reduced Size Self-delivering Rnai Compounds
Khvorova; Anastasia ; et al.
U.S. patent application number 16/934864 was filed with the patent office on 2021-05-20 for reduced size self-delivering rnai compounds. This patent application is currently assigned to Phio Pharmaceuticals Corp.. The applicant listed for this patent is Phio Pharmaceuticals Corp.. Invention is credited to James Cardia, Joanne Kamens, Anastasia Khvorova, William Salomon, Dmitry Samarsky, Tod M. Woolf.
| Application Number | 20210147849 16/934864 |
| Document ID | / |
| Family ID | 1000005370540 |
| Filed Date | 2021-05-20 |












View All Diagrams
| United States Patent Application | 20210147849 |
| Kind Code | A1 |
| Khvorova; Anastasia ; et al. | May 20, 2021 |
REDUCED SIZE SELF-DELIVERING RNAI COMPOUNDS
Abstract
The present invention relates to RNAi constructs with minimal double-stranded regions, and their use in gene silencing. RNAi constructs associated with the invention include a double stranded region of 8-14 nucleotides and a variety of chemical modifications, and are highly effective in gene silencing.
| Inventors: | Khvorova; Anastasia; (Westborough, MA) ; Salomon; William; (Worcester, MA) ; Kamens; Joanne; (Newton, MA) ; Samarsky; Dmitry; (Westborough, MA) ; Woolf; Tod M.; (Sudbury, MA) ; Cardia; James; (Franklin, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Phio Pharmaceuticals Corp. Marlborough MA |
||||||||||
| Family ID: | 1000005370540 | ||||||||||
| Appl. No.: | 16/934864 | ||||||||||
| Filed: | July 21, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14866681 | Sep 25, 2015 | 10774330 | ||
| 16934864 | ||||
| 14278900 | May 15, 2014 | 9175289 | ||
| 14866681 | ||||
| 13120342 | Oct 7, 2011 | 8796443 | ||
| PCT/US2009/005247 | Sep 22, 2009 | |||
| 14278900 | ||||
| 61224031 | Jul 8, 2009 | |||
| 61149946 | Feb 4, 2009 | |||
| 61192954 | Sep 22, 2008 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 2310/3519 20130101; C12N 2310/321 20130101; C12N 2310/3521 20130101; C12N 2320/53 20130101; C12N 15/1136 20130101; C12N 2310/3231 20130101; C12N 2310/14 20130101; C12N 2320/51 20130101; C12N 2310/3515 20130101; C12N 15/111 20130101; C12N 15/113 20130101; C12N 15/1137 20130101; C12N 2310/3341 20130101; C12N 2310/315 20130101; C12N 2320/32 20130101; C12N 2310/322 20130101 |
| International Class: | C12N 15/113 20060101 C12N015/113; C12N 15/11 20060101 C12N015/11 |
Claims
1.-91. (canceled)
92. An isolated double stranded nucleic acid molecule comprising a guide strand and a passenger strand, wherein the isolated double stranded nucleic acid molecule includes a double stranded region and a single stranded region, wherein the single stranded region is at the 3' end of the guide strand and is 4-12 nucleotides long, wherein the single stranded region contains 3, 4, 5, 6, 7, 8, 9, 10, 11 or 12 phosphorothioate modifications, and wherein at least 40% of the nucleotides of the isolated double stranded nucleic acid molecule are modified.
93. The isolated double stranded nucleic acid molecule of claim 92, wherein the guide strand has a minimal length of 16 nucleotides.
94. The isolated double stranded nucleic acid molecule of claim 92, wherein the single stranded region is at least 6 or at least 7 nucleotides long.
95. The isolated double stranded nucleic acid molecule of claim 92, wherein each nucleotide within the single stranded region has a phosphorothioate modification.
96. The isolated double stranded nucleic acid molecule of claim 92, wherein at least one of the nucleotides of the isolated double stranded molecule that is modified comprises a 2' O-methyl or a 2'-fluoro modification and/or wherein at least one of the nucleotides of the isolated double stranded nucleic acid molecule that is modified comprises a hydrophobic modification.
97. The isolated double stranded nucleic acid molecule of claim 92, wherein a hydrophobic conjugate is attached to the double stranded nucleic acid molecule.
98. The isolated double stranded nucleic acid molecule of claim 97, wherein the hydrophobic conjugate is attached at the 3' end of the double stranded nucleic acid molecule.
99. The isolated double stranded nucleic acid molecule of claim 93, wherein the guide strand is 16-28 nucleotides long.
100. The isolated double stranded nucleic acid molecule of claim 92, wherein the passenger strand is 8-16 nucleotides long
101. An isolated asymmetric nucleic acid molecule comprising: a first polynucleotide wherein the first polynucleotide is complementary to a second polynucleotide and a target gene; and a second polynucleotide, wherein the second polynucleotide is at least 6 nucleotides shorter than the first polynucleotide, wherein the first polynucleotide includes a single stranded region of 6, 7, 8, 9, 10, 11 or 12 nucleotides, wherein the single stranded region of the first polynucleotide contains 3, 4, 5, 6, 7, 8, 9, 10, 11 or 12 phosphorothioate modifications, wherein the asymmetric nucleic acid molecule also includes a double stranded region, and wherein at least 50% of C and U nucleotides in the double stranded region are 2' O-methyl modified or 2'-fluoro modified.
102. The isolated double stranded nucleic acid molecule of claim 101, wherein the single stranded region is 6 or 7 nucleotides long and/or wherein each nucleotide within the single stranded region has a phosphorothioate modification.
103. An isolated double stranded nucleic acid molecule comprising: a guide strand of 17-21 nucleotides in length that has complementarity to a target gene, and a passenger strand of 8-16 nucleotides in length, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide strand has a 3' single stranded region of 4-12 nucleotides in length, wherein the single stranded region comprises 2-12 phosphorothioate modifications, wherein at least 40% of the nucleotides of the double stranded nucleic acid are modified, wherein if the passenger strand is 16 nucleotides long, then one end of the double-stranded nucleic acid molecule is blunt, and wherein at least one modification is a hydrophobic base modification and/or the double stranded nucleic acid molecule is linked to a hydrophobic conjugate.
104. The isolated double stranded nucleic acid molecule of claim 103, wherein the hydrophobic base modification comprises a hydrophobic modification of a pyrimidine base, optionally at position 4 or 5, optionally, wherein the hydrophobic base modification is selected from the group consisting of a phenyl, 4-pyridyl, 2-pyridyl, indolyl, isobutyl, tryptophanyl (C.sub.8H.sub.6N)CH.sub.2CH(NH.sub.2)CO), methyl, butyl, aminobenzyl, and naphthyl modification of a uridine or cytidine.
105. The isolated double stranded nucleic acid molecule of claim 103, wherein the hydrophobic conjugate is a small molecule, optionally wherein the small molecule is a sterol-type molecule, optionally wherein the sterol-type molecule is cholesterol.
106. The isolated double stranded nucleic acid molecule of claim 103, wherein the hydrophobic conjugate is attached to the double stranded nucleic acid molecule through a linker, optionally wherein the linker is a TEG linker.
107. A method for inhibiting the expression of a target gene in a mammalian cell, comprising contacting the mammalian cell with the isolated double stranded nucleic acid molecule of claim 92.
108. The method of claim 92, wherein one end of the isolated double stranded nucleic acid molecule is blunt.
109. The isolated double stranded nucleic acid molecule of claim 101, wherein the guide strand has a minimal length of 16 nucleotides.
110. The isolated double stranded nucleic acid molecule of claim 101, wherein one end of the isolated double stranded nucleic acid molecule is blunt.
Description
RELATED APPLICATIONS
[0001] This application is a continuation of U.S. application Ser. No. 14/278,900, filed on May 15, 2014, entitled "REDUCED SIZE SELF-DELIVERING RNAI COMPOUNDS," which is a continuation of U.S. application Ser. No. 13/120,342, now U.S. Pat. No. 8,796,443, which issued on Aug. 5, 2014, entitled "REDUCED SIZE SELF-DELIVERING RNAI COMPOUNDS," which is a national stage filing under 35 U.S.C. .sctn. 371 of international application PCT/US2009/005247, filed Sep. 22, 2009, which was published under PCT Article 21(2) in English, and claims the benefit under 35 U.S.C. .sctn. 119(e) of U.S. provisional application serial number U.S. 61/192,954, entitled "Chemically Modified Polynucleotides and Methods of Using the Same," filed on Sep. 22, 2008, U.S. 61/149,946, entitled "Minimum Length Triggers of RNA Interference," filed on Feb. 4, 2009, and U.S. 61/224,031, entitled "Minimum Length Triggers of RNA Interference," filed on Jul. 8, 2009, the entire disclosure of each of which is incorporated by reference herein in its entirety.
FIELD OF INVENTION
[0002] The invention pertains to the field of RNA interference (RNAi). The invention more specifically relates to nucleic acid molecules with improved in vivo delivery properties without the use of a delivering agent and their use in efficient gene silencing.
BACKGROUND OF INVENTION
[0003] Complementary oligonucleotide sequences are promising therapeutic agents and useful research tools in elucidating gene functions. However, prior art oligonucleotide molecules suffer from several problems that may impede their clinical development, and frequently make it difficult to achieve intended efficient inhibition of gene expression (including protein synthesis) using such compositions in vivo.
[0004] A major problem has been the delivery of these compounds to cells and tissues. Conventional double-stranded RNAi compounds, 19-29 bases long, form a highly negatively-charged rigid helix of approximately 1.5 by 10-15 nm in size. This rod type molecule cannot get through the cell-membrane and as a result has very limited efficacy both in vitro and in vivo. As a result, all conventional RNAi compounds require some kind of a delivery vehicle to promote their tissue distribution and cellular uptake. This is considered to be a major limitation of the RNAi technology.
[0005] There have been previous attempts to apply chemical modifications to oligonucleotides to improve their cellular uptake properties. One such modification was the attachment of a cholesterol molecule to the oligonucleotide. A first report on this approach was by Letsinger et al., in 1989. Subsequently, ISIS Pharmaceuticals, Inc. (Carlsbad, Calif.) reported on more advanced techniques in attaching the cholesterol molecule to the oligonucleotide (Manoharan, 1992).
[0006] With the discovery of siRNAs in the late nineties, similar types of modifications were attempted on these molecules to enhance their delivery profiles. Cholesterol molecules conjugated to slightly modified (Soutschek, 2004) and heavily modified (Wolfrum, 2007) siRNAs appeared in the literature. Yamada et al., 2008 also reported on the use of advanced linker chemistries which further improved cholesterol mediated uptake of siRNAs. In spite of all this effort, the uptake of these types of compounds appears to be inhibited in the presence of biological fluids resulting in highly limited efficacy in gene silencing in vivo, limiting the applicability of these compounds in a clinical setting.
[0007] Therefore, it would be of great benefit to improve upon the prior art oligonucleotides by designing oligonucleotides that have improved delivery properties in vivo and are clinically meaningful.
SUMMARY OF INVENTION
[0008] Described herein are asymmetric chemically modified nucleic acid molecules with minimal double stranded regions, and the use of such molecules in gene silencing. RNAi molecules associated with the invention contain single stranded regions and double stranded regions, and can contain a variety of chemical modifications within both the single stranded and double stranded regions of the molecule. Additionally, the RNAi molecules can be attached to a hydrophobic conjugate such as a conventional and advanced sterol-type molecule. This new class of RNAi molecules has superior efficacy both in vitro and in vivo than previously described RNAi molecules.
[0009] Aspects of the invention relate to asymmetric nucleic acid molecules including a guide strand, with a minimal length of 16 nucleotides, and a passenger strand forming a double stranded nucleic acid, having a double stranded region and a single stranded region, the double stranded region having 8-15 nucleotides in length, the single stranded region having 5-12 nucleotides in length, wherein the passenger strand is linked to a lipophilic group, wherein at least 40% of the nucleotides of the double stranded nucleic acid are modified, and wherein the single stranded region has at least 2 phosphorothioate modifications. In some embodiments position 1 of the guide strand is 5' phosphorylated. In certain embodiments, position 1 of the guide strand is 2'O-methyl modified and 5' phosphorylated.
[0010] Aspects of the invention relate to isolated double stranded nucleic acid molecules including a longer strand of 15-21 nucleotides in length that has complementarily to a miRNA sequence, a shorter strand of 8-15 nucleotides in length linked at the 3' end to a lipophilic group, wherein the longer strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the longer strand has a 3' single stranded region of 2-13 nucleotides in length, comprising at least two phosphorothioate modification, and at least 50% nucleotides are modified.
[0011] Further aspects of the invention relate to isolated double stranded nucleic acid molecules including a guide strand of 17-21 nucleotides in length that has complementarity to a target gene, a passenger strand of 8-16 nucleotides in length linked at the 3' end to a lipophilic group, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide strand has a 3' single stranded region of 2-13 nucleotides in length, each nucleotide within the single stranded region having a phosphorothioate modification, wherein the guide strand has a 5' phosphate modification and wherein at least 50% of C and U nucleotides in the double stranded region include at least one 2' O-methyl modification or 2'-fluoro modification.
[0012] In another aspect, the invention is an isolated double stranded nucleic acid molecule having a guide strand of 17-21 nucleotides in length that has complementarity to a target gene, a passenger strand of 10-16 nucleotides in length linked at the 3' end to a lipophilic group, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide strand has a 3' single stranded region of 5-11 nucleotides in length, at least two nucleotide within the single stranded region having a phosphorothioate modification, wherein the guide strand has a 5' phosphate modification and wherein at least 50% of C and U nucleotides in the double stranded region are 2' O-methyl modification or 2'-fluoro modified.
[0013] The invention in another aspect is an isolated double stranded nucleic acid molecule having a guide strand of 17-21 nucleotides in length that has complementarity to a target gene, a passenger strand of 8-16 nucleotides in length linked at the 3' end to a lipophilic group, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide strand has a 3' single stranded region of 6-8 nucleotides in length, each nucleotide within the single stranded region having a phosphorothioate modification, wherein the guide strand has a 5' phosphate modification, wherein the passenger strand includes at least two phosphorothioate modifications, wherein at least 50% of C and U nucleotides in the double stranded region include a 2' O-methyl modification or 2'-fluoro modification, and wherein the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang.
[0014] An isolated double stranded nucleic acid molecule having a guide strand of 17-21 nucleotides in length that has complementarity to a target gene, a passenger strand of 8-16 nucleotides in length linked at the 3' end to a lipophilic group, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide strand has a 3' single stranded region, each nucleotide within the single stranded region having a phosphorothioate modification, wherein the guide strand has a 5' phosphate modification, wherein every C and U nucleotide in position 11-18 of the guide strand has a 2' O-methyl modification, wherein every nucleotide of the passenger strand is 2' O-methyl modified, and wherein the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang is provided in other aspects of the invention.
[0015] In another aspect the invention is an isolated double stranded nucleic acid molecule having a guide strand of 17-21 nucleotides in length that has complementarity to a target gene, a passenger strand of 8-15 nucleotides in length linked at the 3' end to a lipophilic group, wherein the lipophilic group is selected from the group consisting of cholesterol and a sterol type molecule with C17 polycarbon chain of 5-7 or 9-18 carbons in length, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide strand has a 3' single stranded region, each nucleotide within the single stranded region having a phosphorothioate modification, wherein the guide strand has a 5' phosphate modification, wherein every C and U nucleotide in position 11-18 of the guide strand has a 2' O-methyl modification, wherein every C and U nucleotide in position 2-10 of the guide strand has a 2'F modification, wherein every nucleotide of the passenger strand is 2' O-methyl modified, and wherein the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang.
[0016] In yet another aspect the invention is an isolated nucleic acid molecule having a guide sequence that has complementarity to a target gene, a passenger sequence linked at the 3' end to a lipophilic group, wherein the guide sequence and the passenger sequence form a nucleic acid molecule having a double stranded region and a single stranded region, wherein the guide sequence has a 3' single stranded region of 2-13 nucleotides in length, each nucleotide within the single stranded region having a phosphorothioate modification, wherein the guide sequence has a 5' phosphate modification, wherein at least 50% of C and U nucleotides in the double stranded region include at least one 2' O-methyl modification or 2'-fluoro modification, and wherein the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang.
[0017] An isolated double stranded nucleic acid molecule having a guide strand and a passenger strand, wherein the region of the molecule that is double stranded is from 8-14 nucleotides long, wherein the guide strand contains a single stranded region that is 4-12 nucleotides long, and wherein the single stranded region of the guide strand contains 2-12 phosphorothioate modifications is provided in other aspects of the invention.
[0018] In some embodiments the guide strand contains 6-8 phosphorothioate modifications. In other embodiments the single stranded region of the guide strand is 6 nucleotides long.
[0019] In yet other embodiments the double stranded region is 13 nucleotides long. Optionally the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang.
[0020] In another aspect the invention is an isolated double stranded nucleic acid molecule having a guide strand, wherein the guide strand is 16-28 nucleotides long and has complementarity to a target gene, wherein the 3' terminal 10 nucleotides of the guide strand include at least two phosphate modifications, and wherein the guide strand has a 5' phosphate modification and includes at least one 2' O-methyl modification or 2'-fluoro modification, and a passenger strand, wherein the passenger strand is 8-14 nucleotides long and has complementarity to the guide strand, wherein the passenger strand is linked to a lipophilic group, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule.
[0021] In some embodiments the nucleotide in position one of the guide strand or sequence has a 2'-O-methyl modification. In other embodiments at least one C or U nucleotide in positions 2-10 of the guide strand or sequence has a 2'-fluoro modification. In yet other embodiments every C and U nucleotide in positions 2-10 of the guide strand or sequence has a 2'-fluoro modification. At least one C or U nucleotide in positions 11-18 of the guide strand or sequence may have a 2'-O-methyl modification. In some embodiments every C and U nucleotide in positions 11-18 of the guide strand or sequence has a 2'-O-methyl modification.
[0022] In yet other embodiments the 3' terminal 10 nucleotides of the guide strand include at least four phosphate modifications. Optionally the 3' terminal 10 nucleotides of the guide strand include at least eight phosphate modifications. In some embodiments the guide strand includes 4-14 phosphate modifications. In other embodiments the guide strand includes 4-10 phosphate modifications. In yet other embodiments the 3' terminal 6 nucleotides of the guide strand all include phosphate modifications. The phosphate modifications may be phosphorothioate modifications.
[0023] In some embodiments every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In other embodiments every nucleotide on the passenger strand has a 2'-O-methyl modification. In an embodiment at least one nucleotide on the passenger strand is phosphorothioate modified. At least two nucleotides on the passenger strand are phosphorothioate modified in other embodiments.
[0024] The lipophilic molecule may be a sterol, such as cholesterol.
[0025] In some embodiments the guide strand is 18-19 nucleotides long. In other embodiments the passenger strand is 11-13 nucleotides long.
[0026] The double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang in other embodiments.
[0027] In other aspects the invention is an isolated double stranded nucleic acid molecule comprising a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand, and wherein the guide stand has at least two chemical modifications. In some embodiments the at least two chemical modifications include at least two phosphorothioate modifications. In some embodiments the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang.
[0028] In some aspects the invention is an isolated double stranded nucleic acid molecule comprising a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand, and wherein the guide stand has a single stranded 3' region that is 5 nucleotides or longer and a 5' region that is 1 nucleotide or less. The single stranded region may contain at least 2 phosphorothioate modifications.
[0029] An isolated double stranded nucleic acid molecule having a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-16 nucleotides long and has complementarity to the guide strand, and wherein the guide stand has a single stranded 3' region that is 5 nucleotides or longer and a passenger strand has a sterol type molecule with C17 attached chain longer than 9 is provided in other aspects of the invention.
[0030] A duplex polynucleotide is provided in other aspects of the invention. The polynucleotide has a first polynucleotide wherein said first polynucleotide is complementary to a second polynucleotide and a target gene; and a second polynucleotide wherein said second polynucleotide is at least 6 nucleotides shorter than said first polynucleotide, wherein said first polynucleotide includes a single stranded region containing modifications selected from the group consisting of 40-90% hydrophobic base modifications, 40-90% phosphorothioates, and 40-90% modifications of the ribose moiety, or any combination thereof.
[0031] In other aspects the invention is a duplex polynucleotide having a first polynucleotide wherein said first polynucleotide is complementary to a second polynucleotide and a target gene; and a second polynucleotide wherein said second polynucleotide is at least 6 nucleotides shorter than said first polynucleotide, wherein the duplex polynucleotide includes a mismatch between nucleotides 9, 11, 12, 13 or 14 on the first polynucleotide and the opposite nucleotide on the second polynucleotide.
[0032] In other aspects the invention is a method for inhibiting the expression of a target gene in a mammalian cell, comprising contacting the mammalian cell with an isolated double stranded nucleic acid molecule described herein or a duplex polynucleotide described herein.
[0033] A method of inducing RNAi in a subject is provided in other aspects of the invention. The method involves administering to a subject an effective amount for inducing RNAi of an mRNA of a target gene, an isolated double stranded nucleic acid molecule described herein or a duplex polynucleotide described herein. In other embodiment the subject is a human. In other embodiments the target gene is PPIB, MAP4K4, or SOD1.
[0034] In other aspects an isolated hydrophobic modified polynucleotide having a polynucleotide, wherein the polynucleotide is double stranded RNA, attached to a hydrophobic molecule, wherein the hydrophobic molecule is attached to a base, a ribose or a backbone of a non-terminal nucleotide and wherein the isolated double stranded nucleic acid molecule comprises a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand is provided.
[0035] In one embodiment the hydrophobic molecule is attached to the guide strand of the double stranded RNA. In another embodiment the 3' terminal 10 nucleotides of the guide strand include at least two phosphate modifications, and wherein the guide strand has a 5' phosphate modification and includes at least one 2' O-methyl modification or 2'-fluoro modification. In yet another embodiment the hydrophobic molecule is attached to the passenger strand of the double stranded RNA.
[0036] The invention provides an isolated hydrophobic modified polynucleotide having a polynucleotide non-covalently complexed to a hydrophobic molecule, wherein the hydrophobic molecule is a polycationic molecule. In some embodiments the polycationic molecule is selected from the group consisting of protamine, arginine rich peptides, and spermine.
[0037] In other aspects the invention an isolated hydrophobic modified polynucleotide having a polynucleotide, wherein the polynucleotide is double stranded RNA, directly complexed to a hydrophobic molecule without a linker, wherein the hydrophobic molecule is not cholesterol.
[0038] A composition having a hydrophobic modified polynucleotide, wherein the polynucleotide is double stranded RNA, attached to a hydrophobic molecule, wherein the double stranded nucleic acid molecule comprises a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand, wherein position 1 of the guide strand is 5' phosphorylated or has a 2' O-methyl modification, wherein at least 40% of the nucleotides of the double stranded nucleic acid are modified, and wherein the double stranded nucleic acid molecule has one end that is blunt or includes a one-two nucleotide overhang; a neutral fatty mixture; and optionally a cargo molecule, wherein the hydrophobic modified polynucleotide and the neutral fatty mixture forms a micelle is provided in other aspects of the invention.
[0039] In some embodiments the 3' end of the passenger strand is linked to the hydrophobic molecule. In other embodiments the composition is sterile. In yet other embodiments the neutral fatty mixture comprises a DOPC (dioleoylphosphatidylcholine). In further embodiments the neutral fatty mixture comprises a DSPC (distearoylphosphatidylcholine). The neutral fatty mixture further comprises a sterol such as cholesterol in other embodiments.
[0040] In yet other embodiments the composition includes at least 20% DOPC and at least 20% cholesterol. The hydrophobic portion of the hydrophobic modified polynucleotide is a sterol in other embodiments. The sterol may be a cholesterol, a cholesteryl or modified cholesteryl residue. In other embodiments the hydrophobic portion of the hydrophobic modified polynucleotide is selected from the group consisting of bile acids, cholic acid or taurocholic acid, deoxycholate, oleyl litocholic acid, oleoyl cholenic acid, glycolipids, phospholipids, sphingolipids, isoprenoids, vitamins, saturated fatty acids, unsaturated fatty acids, fatty acid esters, triglycerides, pyrenes, porphyrines, Texaphyrine, adamantane, acridines, biotin, coumarin, fluorescein, rhodamine, Texas-Red, digoxygenin, dimethoxytrityl, t-butyldimethylsilyl, t-butyldiphenylsilyl, cyanine dyes (e.g. Cy3 or Cy5), Hoechst 33258 dye, psoralen, and ibuprofen.
[0041] In yet other embodiments the hydrophobic portion of the hydrophobic modified polynucleotide is a polycationic molecule, such as, for instance, protamine, arginine rich peptides, and/or spermine.
[0042] The composition optionally includes a cargo molecule such as a lipid, a peptide, vitamin, and/or a small molecule. In some embodiments the cargo molecule is a commercially available fat emulsions available for a variety of purposes selected from the group consisting of parenteral feeding. In some embodiments the commercially available fat emulsion is an intralipid or a nutralipid. In other embodiments the cargo molecule is a fatty acid mixture containing more then 74% of linoleic acid, a fatty acid mixture containing at least 6% of cardiolipin, or a fatty acid mixture containing at least 74% of linoleic acid and at least 6% of cardiolipin. In another embodiment the cargo molecule is a fusogenic lipid, such as for example, DOPE, and preferably is at least 10% fusogenic lipid
[0043] In some embodiments the polynucleotide includes chemical modifications. For instance it may be at least 40% modified.
[0044] A method of inducing RNAi in a subject is provided in another aspect of the invention. The method involves administering to a subject an effective amount for inducing RNAi of mRNA of a target gene, an isolated double stranded nucleic acid molecule or a duplex polynucleotide or a composition of the invention, wherein the polynucleotide has at least a region of sequence correspondence to the target gene, wherein the step of administering is systemic, intravenous, intraperitoneal, intradermal, topical, intranasal, inhalation, oral, intramucosal, local injection, subcutaneous, oral tracheal, or intraocular.
[0045] In other embodiment the subject is a human. In other embodiments the target gene is PPIB, MAP4K4, or SOD1.
[0046] In some aspects the invention is a single-stranded RNA of less than 35 nucleotides in length that forms a hairpin structure, said hairpin includes a double-stranded stem and a single-stranded loop, said double-stranded stem having a 5'-stem sequence having a 5'-end, and a 3'-stem sequence having a 3'-end; and said 5'-stem sequence and at least a portion of said loop form a guide sequence complementary to a transcript of a target gene, wherein said polynucleotide mediates sequence-dependent gene silencing of expression of said target gene, wherein each nucleotide within the single-stranded loop region has a phosphorothioate modification, and wherein at least 50% of C and U nucleotides in the double stranded region include a 2' O-methyl modification or 2'-fluoro modification. In one embodiment every C and U nucleotide in position 11-18 of the guide sequence has a 2' O-methyl modification.
[0047] A polynucleotide construct is provided in other aspects, the polynucleotide having two identical single-stranded polynucleotides, each of said single-stranded polynucleotide comprising a 5'-stem sequence having a 5'-end, a 3'-stem sequence having a 3'-end, and a linker sequence linking the 5'-stem sequence and the 3'-stem sequence, wherein: (1) the 5'-stem sequence of a first single-stranded polynucleotide hybridizes with the 3'-stem sequence of a second single-stranded polynucleotide to form a first double-stranded stem region; (2) the 5'-stem sequence of the second single-stranded polynucleotide hybridize with the 3'-stem sequence of the first single-stranded polynucleotide to form a second double-stranded stem region; and, (3) the linker sequences of the first and the second single-stranded polynucleotides form a loop or bulge connecting said first and said second double-stranded stem regions, wherein the 5'-stem sequence and at least a portion of the linker sequence form a guide sequence complementary to a transcript of a target gene, wherein said polynucleotide construct mediates sequence-dependent gene silencing of expression of said target gene, wherein each nucleotide within the single-stranded loop region has a phosphorothioate modification, and wherein at least 50% of C and U nucleotides in the double stranded regions include a 2' O-methyl modification or 2'-fluoro modification.
[0048] In one embodiment every C and U nucleotide in position 11-18 of the guide sequence has a 2' O-methyl modification.
[0049] In some embodiments, the guide strand is 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, or 29 nucleotides long. In some embodiments, the passenger strand is 8, 9, 10, 11, 12, 13 or 14 nucleotides long. In some embodiments, the nucleic acid molecule has a thermodynamic stability (.DELTA.G) of less than -20 kkal/mol.
[0050] Aspects of the invention relate to nucleic acid molecules that are chemically modified. In some embodiments, the chemical modification is selected from the group consisting of 5' Phosphate, 2'-O-methyl, 2'-O-ethyl, 2'-fluoro, ribothymidine, C-5 propynyl-dC (pdC), C-5 propynyl-dU (pdU), C-5 propynyl-C(pC), C-5 propynyl-U (pU), 5-methyl C, 5-methyl U, 5-methyl dC, 5-methyl dU methoxy, (2,6-diaminopurine), 5'-Dimethoxytrityl-N4-ethyl-2'-deoxyCytidine, C-5 propynyl-fC (pfC), C-5 propynyl-fU (pfU), 5-methyl fC, 5-methyl fU, C-5 propynyl-mC (pmC), C-5 propynyl-fU (pmU), 5-methyl mC, 5-methyl mU, LNA (locked nucleic acid), MGB (minor groove binder) and other base modifications which increase base hydrophobicity. More than one chemical modification may be present in the same molecule. In some embodiments, chemical modification increases stability and/or improves thermodynamic stability (.DELTA.G). In some embodiments, at least 90% of CU residues on a nucleic acid molecule are modified.
[0051] In some embodiments, the nucleotide in position one of the guide strand has a 2'-O-methyl modification and/or a 5' Phosphate modification. In some embodiments, at least one C or U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In certain embodiments, every C and U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In some embodiments, at least one C or U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In certain embodiments, every C and U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In some embodiments, every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In certain embodiments, every nucleotide on the passenger strand has a 2'-O-methyl modification.
[0052] In some embodiments, nucleic acid molecules associated with the invention contain a stretch of at least 4 nucleotides that are phosphorothioate modified. In certain embodiments, the stretch of nucleotides that are phosphorothioate modified is at least 12 nucleotides long. In some embodiments, the stretch of nucleotides that are phosphorothioate modified is not fully single stranded.
[0053] Nucleic acid molecules associated with the invention may be attached to a conjugate. In some embodiments, the conjugate is attached to the guide strand, while in other embodiments the conjugate is attached to the passenger strand. In some embodiments, the conjugate is hydrophobic. In some embodiments, the conjugate is a sterol such as cholesterol. In some embodiments, nucleic acid molecules associated with the invention are blunt-ended.
[0054] Aspects of the invention relate to double stranded nucleic acid molecule including a guide strand and a passenger strand, wherein the region of the molecule that is double stranded is from 8-14 nucleotides long, and wherein the molecule has a thermodynamic stability (.DELTA.G) of less than -13 kkal/mol.
[0055] In some embodiments, the region of the molecule that is double stranded is 8, 9, 10, 11, 12, 13, or 14 nucleotides long. In some embodiments, the molecule has a thermodynamic stability (.DELTA.G) of less than -20 kkal/mol. The nucleic acid molecules, in some embodiments are chemically modified. In certain embodiments, the chemical modification is selected from the group consisting of 5' Phosphate, 2'-O-methyl, 2'-O-ethyl, 2'-fluoro, ribothymidine, C-5 propynyl-dC (pdC), C-5 propynyl-dU (pdU), C-5 propynyl-C(pC), C-5 propynyl-U (pU), 5-methyl C, 5-methyl U, 5-methyl dC, 5-methyl dU methoxy, (2,6-diaminopurine), 5'-Dimethoxytrityl-N4-ethyl-2'-deoxyCytidine, C-5 propynyl-fC (pfC), C-5 propynyl-fU (pfU), 5-methyl fC, 5-methyl fU, C-5 propynyl-mC (pmC), C-5 propynyl-fU (pmU), 5-methyl mC, 5-methyl mU, LNA (locked nucleic acid), MGB (minor groove binder) and other base modifications which increase base hydrophobicity. More than one chemical modification may be present in the same molecule. In some embodiments, chemical modification increases stability and/or improves thermodynamic stability (.DELTA.G). In some embodiments, at least 90% of CU residues on a nucleic acid molecule are modified.
[0056] In some embodiments, the nucleotide in position one of the guide strand has a 2'-O-methyl modification and/or a 5' Phosphate modification. In some embodiments, at least one C or U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In certain embodiments, every C and U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In some embodiments, at least one C or U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In certain embodiments, every C and U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In some embodiments, every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In certain embodiments, every nucleotide on the passenger strand has a 2'-O-methyl modification.
[0057] The nucleic acid molecules associated with the invention may contain a stretch of at least 4 nucleotides that are phosphorothioate modified. In certain embodiments, the stretch of nucleotides that are phosphorothioate modified is at least 12 nucleotides long. In some embodiments, the stretch of nucleotides that are phosphorothioate modified is not fully single stranded. In some embodiments, the nucleic acid molecules are attached to a conjugate. In some embodiments, the conjugate is attached to the guide strand, while in other embodiments the conjugate is attached to the passenger strand. In some embodiments, the conjugate is hydrophobic. In some embodiments, the conjugate is a sterol such as cholesterol. In some embodiments, nucleic acid molecules associated with the invention are blunt-ended. In some embodiments, the nucleic acid molecules are blunt ended at the 5'end. In certain embodiments, the nucleic acid molecules are blunt ended at the 5'end where the region of complementarity between the two strands of the molecule begins.
[0058] Aspects of the invention relate to methods for inhibiting the expression of a target gene in a mammalian cell. Methods include contacting the mammalian cell with an isolated double stranded nucleic acid molecule including a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and has complementarity to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand, and wherein the double stranded nucleic acid molecule has a thermodynamic stability (.DELTA.G) of less than -13 kkal/mol.
[0059] The cell may be contacted in vivo or in vitro. In some embodiments, the guide strand is 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, or 29 nucleotides long. In some embodiments, the passenger strand is 8, 9, 10, 11, 12, 13 or 14 nucleotides long. In some embodiments, the nucleic acid molecule has a thermodynamic stability (.DELTA.G) of less than -20 kkal/mol.
[0060] The nucleic acid molecules associated with methods described herein may be chemically modified. In some embodiments, the chemical modification is selected from the group consisting of 5' Phosphate, 2'-O-methyl, 2'-O-ethyl, 2'-fluoro, ribothymidine, C-5 propynyl-dC (pdC), C-5 propynyl-dU (pdU), C-5 propynyl-C(pC), C-5 propynyl-U (pU), 5-methyl C, 5-methyl U, 5-methyl dC, 5-methyl dU methoxy, (2,6-diaminopurine), 5'-Dimethoxytrityl-N4-ethyl-2'-deoxyCytidine, C-5 propynyl-fC (pfC), C-5 propynyl-fU (pfU), 5-methyl fC, 5-methyl fU, C-5 propynyl-mC (pmC), C-5 propynyl-fU (pmU), 5-methyl mC, 5-methyl mU, LNA (locked nucleic acid), MGB (minor groove binder) and other base modifications which increase base hydrophobicity. More than one chemical modification may be present in the same molecule. In some embodiments, chemical modification increases stability and/or improves thermodynamic stability (.DELTA.G). In some embodiments, at least 90% of CU residues on a nucleic acid molecule are modified.
[0061] In some embodiments, the nucleotide in position one of the guide strand has a 2'-O-methyl modification and/or a 5' Phosphate modification. In some embodiments, at least one C or U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In certain embodiments, every C and U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In some embodiments, at least one C or U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In certain embodiments, every C and U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In some embodiments, every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In certain embodiments, every nucleotide on the passenger strand has a 2'-O-methyl modification.
[0062] In some embodiments, nucleic acid molecules associated with the invention contain a stretch of at least 4 nucleotides that are phosphorothioate modified. In certain embodiments, the stretch of nucleotides that are phosphorothioate modified is at least 12 nucleotides long. In some embodiments, the stretch of nucleotides that are phosphorothioate modified is not fully single stranded.
[0063] Nucleic acid molecules associated with the invention may be attached to a conjugate. In some embodiments, the conjugate is attached to the guide strand, while in other embodiments the conjugate is attached to the passenger strand. In some embodiments, the conjugate is hydrophobic. In some embodiments, the conjugate is a sterol such as cholesterol. In some embodiments, nucleic acid molecules associated with the invention are blunt-ended.
[0064] Methods for inhibiting the expression of a target gene in a mammalian cell described herein include contacting the mammalian cell with an isolated double stranded nucleic acid molecule including a guide strand and a passenger strand, wherein the region of the molecule that is double stranded is from 8-14 nucleotides long, and wherein the molecule has a thermodynamic stability (.DELTA.G) of less than -13 kkal/mol.
[0065] In some embodiments, the region of the molecule that is double stranded is 8, 9, 10, 11, 12, 13, or 14 nucleotides long. In some embodiments, the molecule has a thermodynamic stability (.DELTA.G) of less than -20 kkal/mol. The nucleic acid molecules, in some embodiments are chemically modified. In certain embodiments, the chemical modification is selected from the group consisting of 5' Phosphate, 2'-O-methyl, 2'-O-ethyl, 2'-fluoro, ribothymidine, C-5 propynyl-dC (pdC), C-5 propynyl-dU (pdU), C-5 propynyl-C(pC), C-5 propynyl-U (pU), 5-methyl C, 5-methyl U, 5-methyl dC, 5-methyl dU methoxy, (2,6-diaminopurine), 5'-Dimethoxytrityl-N4-ethyl-2'-deoxyCytidine, C-5 propynyl-fC (pfC), C-5 propynyl-fU (pfU), 5-methyl fC, 5-methyl fU, C-5 propynyl-mC (pmC), C-5 propynyl-fU (pmU), 5-methyl mC, 5-methyl mU, LNA (locked nucleic acid), MGB (minor groove binder) and other base modifications which increase base hydrophobicity. More than one chemical modification may be present in the same molecule. In some embodiments, chemical modification increases stability and/or improves thermodynamic stability (.DELTA.G). In some embodiments, at least 90% of CU residues on a nucleic acid molecule are modified.
[0066] In some embodiments, the nucleotide in position one of the guide strand has a 2'-O-methyl modification and/or a 5' Phosphate modification. In some embodiments, at least one C or U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In certain embodiments, every C and U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In some embodiments, at least one C or U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In certain embodiments, every C and U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In some embodiments, every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In certain embodiments, every nucleotide on the passenger strand has a 2'-O-methyl modification.
[0067] The nucleic acid molecules associated with the invention may contain a stretch of at least 4 nucleotides that are phosphorothioate modified. In certain embodiments, the stretch of nucleotides that are phosphorothioate modified is at least 12 nucleotides long. In some embodiments, the stretch of nucleotides that are phosphorothioate modified is not fully single stranded. In some embodiments, the nucleic acid molecules are attached to a conjugate. In some embodiments, the conjugate is attached to the guide strand, while in other embodiments the conjugate is attached to the passenger strand. In some embodiments, the conjugate is hydrophobic. In some embodiments, the conjugate is a sterol such as cholesterol. In some embodiments, nucleic acid molecules associated with the invention are blunt-ended.
[0068] In another embodiment, the invention provides a method for selecting an siRNA for gene silencing by (a) selecting a target gene, wherein the target gene comprises a target sequence; (b) selecting a candidate siRNA, wherein said candidate siRNA comprises a guide strand of 16-29 nucleotide base pairs and a passenger strand of 8-14 nucleotide base pairs that form a duplex comprised of an antisense region and a sense region and said antisense region of said candidate siRNA is at least 80% complementary to a region of said target sequence; (c) determining a thermodynamic stability (.DELTA.G) of the candidate siRNA; and (e) selecting said candidate siRNA as an siRNA for gene silencing, if said thermodynamic stability is less than -13 kkal/mol.
[0069] Aspects of the invention relate to isolated double stranded nucleic acid molecules including a guide strand and a passenger strand, wherein the guide strand is 18-19 nucleotides long and has complementarity to a target gene, wherein the passenger strand is 11-13 nucleotides long and has complementarity to the guide strand, and wherein the double stranded nucleic acid molecule has a thermodynamic stability (.DELTA.G) of less than -13 kkal/mol.
[0070] In some embodiments, the nucleotide in position one of the guide strand has a 2'-O-methyl modification and/or a 5' Phosphate modification. In some embodiments, at least one C or U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In certain embodiments, every C and U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In some embodiments, at least one C or U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In certain embodiments, every C and U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification.
[0071] In some embodiments, the guide strand contains a stretch of at least 4 nucleotides that are phosphorothioate modified. In certain embodiments, the guide strand contains a stretch of at least 8 nucleotides that are phosphorothioate modified. In some embodiments, every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In certain embodiments, every nucleotide on the passenger strand has a 2'-O-methyl modification. In some embodiments, at least one, or at least two nucleotides on the passenger strand is phosphorothioate modified. The nucleic acid molecule can be attached to a conjugate on either the guide or passenger strand. In some embodiments, the conjugate is a sterol such as cholesterol.
[0072] Aspects of the invention relate to isolated double stranded nucleic acid molecules including a guide strand, wherein the guide strand is 16-28 nucleotides long and has complementarity to a target gene, wherein the 3' terminal 10 nucleotides of the guide strand include at least two phosphate modifications, and wherein the guide strand includes at least one 2' O-methyl modification or 2'-fluoro modification, and a passenger strand, wherein the passenger strand is 8-28 nucleotides long and has complementarity to the guide strand, wherein the passenger strand is linked to a lipophilic group, wherein the guide strand and the passenger strand form the double stranded nucleic acid molecule.
[0073] In some embodiments, the nucleotide in position one of the guide strand has a 2'-O-methyl modification and/or a 5' Phosphate modification. In some embodiments, at least one C or U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In certain embodiments, every C and U nucleotide in positions 2-10 of the guide strand has a 2'-fluoro modification. In some embodiments, at least one C or U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification. In certain embodiments, every C and U nucleotide in positions 11-18 of the guide strand has a 2'-O-methyl modification.
[0074] In some embodiments, the 3' terminal 10 nucleotides of the guide strand include at least four, or at least eight phosphate modifications. In certain embodiments, the guide strand includes 2-14 or 4-10 phosphate modifications. In some embodiments, the 3' terminal 6 nucleotides of the guide strand all include phosphate modifications. In certain embodiments, the phosphate modifications are phosphorothioate modifications.
[0075] In some embodiments, every C and U nucleotide on the passenger strand has a 2'-O-methyl modification. In certain embodiments, every nucleotide on the passenger strand has a 2'-O-methyl modification. In some embodiments, at least one, or at least two nucleotides on the passenger strand is phosphorothioate modified. In some embodiments, the lipophilic molecule is a sterol such as cholesterol. In some embodiments, the guide strand is 18-19 nucleotides long and the passenger strand is 11-13 nucleotides long.
[0076] Aspects of the invention relate to isolated double stranded nucleic acid molecules including a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand, and wherein the guide stand has at least two chemical modifications. In some embodiments, the two chemical modifications are phosphorothioate modifications.
[0077] Further aspects of the invention relate to isolated double stranded nucleic acid molecule comprising a guide strand and a passenger strand, wherein the guide strand is from 16-29 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 8-14 nucleotides long and has complementarity to the guide strand, and wherein the guide stand has a single stranded 3' region that is 5 nucleotides or longer. In some embodiments, the single stranded region contains at least 2 phosphorothioate modifications.
[0078] Further aspects of the invention relate to isolated double stranded nucleic acid molecules including a guide strand and a passenger strand, wherein the guide strand is from 18-21 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 11-14 nucleotides long and has complementarity to the guide strand, and wherein position one of the guide stand has 2-OMe and 5' phosphate modifications, every C and U in positions 2 to 11 of the guide strand are 2-F modified, every C and U in positions 12-18 of the guide strand are 2'OMe modified, and 80% of Cs and Us on the passenger strand are 2'OMe modified
[0079] Another aspect of the invention relates to isolated double stranded nucleic acid molecules including a guide strand and a passenger strand, wherein the guide strand is from 18-21 nucleotides long and is substantially complementary to a target gene, wherein the passenger strand is from 11-14 nucleotides long and has complementarity to the guide strand, and wherein the guide stand has 2-OMe and 5' phosphate modifications at position 1, every C and U in positions 2 to 11 of the guide strand are 2-F modified, every C and U in positions 12-18 of the guide strand are 2'OMe modified, 80% of Cs and Us on the passenger strand are 2'OMe and the 3' end of the passenger strand is attached to a conjugate. In some embodiments the conjugate is selected from sterols, sterol-type molecules, hydrophobic vitamins or fatty acids.
[0080] Each of the limitations of the invention can encompass various embodiments of the invention. It is, therefore, anticipated that each of the limitations of the invention involving any one element or combinations of elements can be included in each aspect of the invention. This invention is not limited in its application to the details of construction and the arrangement of components set forth in the following description or illustrated in the drawings. The invention is capable of other embodiments and of being practiced or of being carried out in various ways.
BRIEF DESCRIPTION OF DRAWINGS
[0081] The accompanying drawings are not intended to be drawn to scale. In the drawings, each identical or nearly identical component that is illustrated in various figures is represented by a like numeral. For purposes of clarity, not every component may be labeled in every drawing. In the drawings:
[0082] FIG. 1 is a schematic depicting proposed structures of asymmetric double stranded RNA molecules (adsRNA). Bold lines represent sequences carrying modification patterns compatible with RISC loading. Striped lines represent polynucleotides carrying modifications compatible with passenger strands. Plain lines represent a single stranded polynucleotide with modification patterns optimized for cell interaction and uptake. FIG. 1A depicts adsRNA with extended guide or passenger strands; FIG. 1B depicts adsRNA with length variations of a cell penetrating polynucleotide; FIG. 1C depicts adsRNA with 3' and 5' conjugates; FIG. 1D depicts adsRNAs with mismatches.
[0083] FIG. 2 is a schematic depicting asymmetric dsRNA molecules with different chemical modification patterns. Several examples of chemical modifications that might be used to increase hydrophobicity are shown including 4-pyridyl, 2-pyridyl, isobutyl and indolyl based position 5 uridine modifications.
[0084] FIG. 3 is a schematic depicting the use of dsRNA binding domains, protamine (or other Arg rich peptides), spermidine or similar chemical structures to block duplex charge to facilitate cellular entry.
[0085] FIG. 4 is a schematic depicting positively charged chemicals that might be used for polynucleotide charge blockage.
[0086] FIG. 5 is a schematic depicting examples of structural and chemical compositions of single stranded RISC entering polynucleotides. The combination of one or more modifications including 2'd, 2'Ome, 2'F, hydrophobic and phosphorothioate modifications can be used to optimize single strand entry into the RISC.
[0087] FIG. 6 is a schematic depicting examples of structural and chemical composition of RISC substrate inhibitors. Combinations of one or more chemical modifications can be used to mediate efficient uptake and efficient binding to preloaded RISC complex.
[0088] FIG. 7 is a schematic depicting structures of polynucleotides with sterol type molecules attached, where R represent a polycarbonic tail of 9 carbons or longer. FIG. 7A depicts an adsRNA molecule; FIG. 7B depicts an siRNA molecule of approximately 17-30 bp long; FIG. 7C depicts a RISC entering strand; FIG. 7D depicts a substrate analog strand. Chemical modification patterns, as depicted in FIG. 7, can be optimized to promote desired function.
[0089] FIG. 8 is a schematic depicting examples of naturally occurring phytosterols with a polycarbon chain that is longer than 8, attached at position 17. More than 250 different types of phytosterols are known.
[0090] FIG. 9 is a schematic depicting examples of sterol-like structures, with variations in the size of the polycarbon chains attached at position 17.
[0091] FIG. 10 presents schematics and graphs demonstrating that the percentage of liver uptake and plasma clearance of lipid emulsions containing sterol type molecules is directly affected by the size of the polycarbon chain attached at position 17. This figure is adapted from Martins et al, Journal of Lipid Research (1998).
[0092] FIG. 11 is a schematic depicting micelle formation. FIG. 11A depicts a polynucleotide with a hydrophobic conjugate; FIG. 11B depicts linoleic acid; FIG. 11C depicts a micelle formed from a mixture of polynucleotides containing hydrophobic conjugates combined with fatty acids.
[0093] FIG. 12 is a schematic depicting how alteration in lipid composition can affect pharmacokinetic behavior and tissue distribution of hydrophobically modified and/or hydrophobically conjugated polynucleotides. In particular, use of lipid mixtures enriched in linoleic acid and cardiolipin results in preferential uptake by cardiomyocites.
[0094] FIG. 13 is a schematic showing examples of RNAi constructs and controls used to target MAP4K4 expression. RNAi construct 12083 corresponds to SEQ ID NOs:597 and 598. RNAi construct 12089 corresponds to SEQ ID NO:599.
[0095] FIG. 14 is a graph showing MAP4K4 expression following transfection with RNAi constructs associated with the invention. RNAi constructs tested were: 12083 (Nicked), 12085 (13nt Duplex), 12089 (No Stem Pairing) and 12134 (13nt miniRNA). Results of transfection were compared to an untransfected control sample. RNAi construct 12083 corresponds to SEQ ID NOs:597 and 598. RNAi construct 12085 corresponds to SEQ ID NOs:600 and 601. RNAi construct 12089 corresponds to SEQ ID NO:599. RNAi construct 12134 corresponds to SEQ ID NOs:602 and 603.
[0096] FIG. 15 is a graph showing expression of MAP4K4 24 hours post-transfection with RNAi constructs associated with the invention. RNAi constructs tested were: 11546 (MAP4K4 rxRNA), 12083 (MAP4K4 Nicked Construct), 12134 (12 bp soloRNA) and 12241 (14/3/14 soloRNA). Results of transfection were compared to a filler control sample. RNAi construct 11546 corresponds to SEQ ID NOs:604 and 605. RNAi construct 12083 corresponds to SEQ ID NOs:597 and 598. RNAi construct 12134 corresponds to SEQ ID NOs:602 and 603. RNAi construct 12241 corresponds to SEQ ID NOs:606 and 607.
[0097] FIG. 16 presents a graph and several tables comparing parameters associated with silencing of MAP4K4 expression following transfection with RNAi constructs associated with the invention. The rxRNA construct corresponds to SEQ ID NOs:604 and 605. The 14-3-14 soloRNA construct corresponds to SEQ ID NOs:606 and 607. The 13/19 duplex (nicked construct) corresponds to SEQ ID NOs:597 and 598. The 12-bp soloRNA construct corresponds to SEQ ID NOs:602 and 603.
[0098] FIG. 17 is a schematic showing examples of RNAi constructs and controls used to target SOD1 expression. The 12084 RNAi construct corresponds to SEQ ID NOs:612 and 613.
[0099] FIG. 18 is a graph showing SOD1 expression following transfection with RNAi constructs associated with the invention. RNAi constructs tested were: 12084 (Nicked), 12086 (13nt Duplex), 12090 (No Stem Pairing) and 12035 (13nt MiniRNA). Results of transfection were compared to an untransfected control sample. The 12084 RNAi construct corresponds to SEQ ID NOs:612 and 613. The 12086 RNAi construct corresponds to SEQ ID NOs:608 and 609. The 12035 RNAi construct corresponds to SEQ ID NOs:610 and 611.
[0100] FIG. 19 is a graph showing expression of SOD1 24 hours post-transfection with RNAi constructs associated with the invention. RNAi constructs tested were: 10015 (SOD1 rxRNA) and 12084 (SOD1 Nicked Construct). Results of transfection were compared to a filler control sample. The 10015 RNAi construct corresponds to SEQ ID NOs:614 and 615. The 12084 RNAi construct corresponds to SEQ ID NOs:612 and 613.
[0101] FIG. 20 is a schematic indicating that RNA molecules with double stranded regions that are less than 10 nucleotides are not cleaved by Dicer.
[0102] FIG. 21 is a schematic revealing a hypothetical RNAi model for RNA induced gene silencing.
[0103] FIG. 22 is a graph showing chemical optimization of asymmetric RNAi compounds. The presence of chemical modifications, in particular 2'F UC, phosphorothioate modifications on the guide strand, and complete CU 2'OMe modification of the passenger strands results in development of functional compounds. Silencing of MAP4K4 following lipid-mediated transfection is shown using RNAi molecules with specific modifications. RNAi molecules tested had sense strands that were 13 nucleotides long and contained the following modifications: unmodified; C and U 2'OMe; C and U 2'OMe and 3' Chl; rxRNA 2'OMe pattern; or full 2'OMe, except base 1. Additionally, the guide (anti-sense) strands of the RNAi molecules tested contained the following modifications: unmodified; unmodified with 5'P; C and U 2'F; C and U 2'F with 8 PS 3' end; and unmodified (17 nt length). Results for rxRNA 12/10 Duplex and negative controls are also shown.
[0104] FIG. 23 demonstrates that the chemical modifications described herein significantly increase in vitro efficacy in un-assisted delivery of RNAi molecules in HeLa cells. The structure and sequence of the compounds were not altered; only the chemical modification patterns of the molecules were modified. Compounds lacking 2' F, 2'O-me, phosphorothioate modification, or cholesterol conjugates were completely inactive in passive uptake. A combination of all 4 of these types of modifications produced the highest levels of activity (compound 12386).
[0105] FIG. 24 is a graph showing MAP4K4 expression in Hela cells following passive uptake transfection of: NT Accell modified siRNA, MAP4K4 Accell siRNA, Non-Chl nanoRNA (12379) and sd-nanoRNA (12386).
[0106] FIG. 25 is a graph showing expression of MAP4K4 in HeLa cells following passive uptake transfection of various concentrations of RNA molecules containing the following parameters: Nano Lead with no 3'Chl; Nano Lead; Accell MAP4K4; 21mer GS with 8 PS tail; 21mer GS with 12 PS tail; and 25mer GS with 12 PS tail.
[0107] FIG. 26 is a graph demonstrating that reduction in oligonucleotide content increases the efficacy of unassisted uptake. Similar chemical modifications were applied to assymetric compounds, traditional siRNA compounds and 25 mer RNAi compounds. The assymetric small compounds demonstrated the most significant efficacy.
[0108] FIG. 27 is a graph demonstrating the importance of phosphorothioate content for un-assisted delivery. FIG. 27A demonstrates the results of a systematic screen that revealed that the presence of at least 2-12 phosphorothioates in the guide strand significantly improves uptake; in some embodiments, 4-8 phosphorothioate modifications were found to be preferred. FIG. 27 B reveals that the presence or absence of phosphorothioate modifications in the sense strand did not alter efficacy.
[0109] FIG. 28 is a graph showing expression of MAP4K4 in primary mouse hepatocytes following passive uptake transfection of: Accell Media-Ctrl-UTC; MM APOB Alnylam; Active APOB Alnylam; nanoRNA without chl; nanoRNA MAP4K4; Mouse MAP4K4 Accell Smartpool; DY547 Accell Control; Luc Ctrl rxRNA with Dy547; MAP4K4 rxRNA with DY547; and AS Strand Alone (nano).
[0110] FIG. 29 is a graph showing expression of ApoB in mouse primary hepatocytes following passive uptake transfection of: Accell Media-Ctrl-UTC; MM APOB Alnylam; Active APOB Alnylam; nanoRNA without chl; nanoRNA MAP4K4; Mouse MAP4K4 Accell Smartpool; DY547 Accell Control; Luc Ctrl rxRNA with Dy547; MAP4K4 rxRNA with DY547; and AS Strand Alone (nano).
[0111] FIG. 30 is a graph showing expression of MAP4K4 in primary human hepatocytes following passive uptake transfection of: 11550 MAP4K4 rxRNA; 12544 MM MAP4K4 nanoRNA; 12539 Active MAP4K4 nanoRNA; Accell Media; and UTC.
[0112] FIG. 31 is a graph showing ApoB expression in primary human hepatoctyes following passive uptake transfection of: 12505 Active ApoB chol-siRNA; 12506 MM ApoB chol-siRNA; Accell Media; and UTC.
[0113] FIG. 32 is an image depicting localization of sd-rxRNA.sup.nano localization.
[0114] FIG. 33 is an image depicting localization of Chol-siRNA (Alnylam).
[0115] FIG. 34 is a schematic of 1.sup.st generation (G1) sd-rxRNA.sup.nano molecules associated with the invention indicating regions that are targeted for modification, and functions associated with different regions of the molecules.
[0116] FIG. 35 depicts modification patterns that were screened for optimization of sd-rxRNA.sup.nano (G1). The modifications that were screened included, on the guide strand, lengths of 19, 21 and 25 nucleotides, phosphorothioate modifications of 0-18 nucleotides, and replacement of 2'F modifications with 2'OMe, 5 Methyl C and/or ribo Thymidine modifications. Modifications on the sense strand that were screened included nucleotide lengths of 11, 13 and 19 nucleotides, phosphorothiote modifications of 0-4 nucleotides and 2'OMe modifications.
[0117] FIG. 36 is a schematic depicting modifications of sd-rxRNA.sup.nano that were screened for optimization.
[0118] FIG. 37 is a graph showing percent MAP4K4 expression in Hek293 cells following transfection of: Risc Free siRNA; rxRNA; Nano (unmodified); GS alone; Nano Lead (no Chl); Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 8 PS, 19 nt); Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 8 PS, 21 nt); Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 12 PS, 21 nt); and Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 12 PS, 25 nt);
[0119] FIG. 38 is a graph showing percent MAP4K4 expression in HeLa cells following passive uptake transfection of: GS alone; Nano Lead; Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 8 PS, 19 nt); Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 8 PS, 21 nt); Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 12 PS, 21 nt); Nano (GS: (3) 2'OMe at positions 1, 18, and 19, 12 PS, 25 nt).
[0120] FIG. 39 is a graph showing percent MAP4K4 expression in Hek293 cells following lipid mediated transfection of: Guide Strand alone (GS: 8PS, 19 nt); Guide Strand alone (GS: 18PS, 19 nt); Nano (GS: no PS, 19 nt); Nano (GS: 2 PS, 19 nt); Nano (GS: 4 PS, 19 nt); Nano (GS: 6 PS, 19 nt); Nano Lead (GS: 8 PS, 19 nt); Nano (GS: 10 PS, 19 nt); Nano (GS: 12 PS, 19 nt); and Nano (GS: 18 PS, 19 nt).
[0121] FIG. 40 is a graph showing percent MAP4K4 expression in Hek293 cells following lipid mediated transfection of: Guide Strand alone (GS: 8PS, 19 nt); Guide Strand alone (GS: 18PS, 19 nt); Nano (GS: no PS, 19 nt); Nano (GS: 2 PS, 19 nt); Nano (GS: 4 PS, 19 nt); Nano (GS: 6 PS, 19 nt); Nano Lead (GS: 8 PS, 19 nt); Nano (GS: 10 PS, 19 nt); Nano (GS: 12 PS, 19 nt); and Nano (GS: 18 PS, 19 nt).
[0122] FIG. 41 is a graph showing percent MAP4K4 expression in HeLa cells following passive uptake transfection of: Nano Lead (no Chl); Guide Strand alone (18 PS); Nano (GS: 0 PS, 19 nt); Nano (GS: 2 PS, 19 nt); Nano (GS: 4 PS, 19 nt); Nano (GS: 6 PS, 19 nt); Nano Lead (GS: 8 PS, 19 nt); Nano (GS: 10 PS, 19 nt); Nano (GS: 12 PS, 19 nt); and Nano (GS: 18 PS, 19 nt).
[0123] FIG. 42 is a graph showing percent MAP4K4 expression in HeLa cells following passive uptake transfection of: Nano Lead (no Chl); Guide Strand alone (18 PS); Nano (GS: 0 PS, 19 nt); Nano (GS: 2 PS, 19 nt); Nano (GS: 4 PS, 19 nt); Nano (GS: 6 PS, 19 nt); Nano Lead (GS: 8 PS, 19 nt); Nano (GS: 10 PS, 19 nt); Nano (GS: 12 PS, 19 nt); and Nano (GS: 18 PS, 19 nt).
[0124] FIG. 43 is a schematic depicting guide strand chemical modifications that were screened for optimization.
[0125] FIG. 44 is a graph showing percent MAP4K4 expression in Hek293 cells following reverse transfection of: RISC free siRNA; GS only (2'F C and Us); GS only (2'OMe C and Us); Nano Lead (2'F C and Us); nano (GS: (3) 2'OMe, positions 16-18); nano (GS: (3) 2'OMe, positions 16, 17 and 19); nano (GS: (4) 2'OMe, positions 11, 16-18); nano (GS: (10) 2'OMe, C and Us); nano (GS: (6) 2'OMe, positions 1 and 5-9); nano (GS: (3) 2'OMe, positions 1, 18 and 19); and nano (GS: (5) 2'OMe Cs).
[0126] FIG. 45 is a graph demonstrating efficacy of various chemical modification patterns. In particular, 2-OMe modification in positions 1 and 11-18 was well tolerated. 2'OMe modifications in the seed area resulted in a slight reduction of efficacy (but were still highly efficient). Ribo-modifications in the seed were well tolerated. This data enabled the generation of self delivering compounds with reduced or no 2'F modifications. This is significant because 2'F modifications may be associated with toxicity in vivo.
[0127] FIG. 46 is a schematic depicting sense strand modifications.
[0128] FIG. 47 is a graph demonstrating sense strand length optimization. A sense strand length between 10-15 bases was found to be optimal in this assay. Increasing sense strand length resulted in a reduction of passive uptake of these compounds but may be tolerated for other compounds. Sense strands containing LNA modification demonstrated similar efficacy to non-LNA containing compounds. In some embodiments, the addition of LNA or other thermodynamically stabilizing compounds can be beneficial, resulting in converting non-functional sequences into functional sequences.
[0129] FIG. 48 is a graph showing percent MAP4K4 expression in HeLa cells following passive uptake transfection of: Guide Strand Alone (2'F C and U); Nano Lead; Nano Lead (No Chl); Nano (SS: 11 nt 2'OMe C and Us, Chl); Nano (SS: lint, complete 2'OMe, Chl); Nano (SS: 19 nt, 2'OMe C and Us, Chl); Nano (SS: 19 nt, 2'OMe C and Us, no Chl).
[0130] FIG. 49 is a graph showing percent MAP4K4 expression in HeLa cells following passive uptake transfection of: Nano Lead (No Chl); Nano (SS no PS); Nano Lead (SS:2 PS); Nano (SS:4 PS).
[0131] FIG. 50 is a schematic depicting a sd-rxRNA.sup.nano second generation (GII) lead molecule.
[0132] FIG. 51 presents a graph indicating EC50 values for MAP4K4 silencing in the presence of sd-rxRNA, and images depicting localization of DY547-labeled rxRNA.sup.ori and DY547-labeled sd-rxRNA.
[0133] FIG. 52 is a graph showing percent MAP4K4 expression in HeLa cells in the presence of optimized sd-rxRNA molecules.
[0134] FIG. 53 is a graph depicting the relevance of chemistry content in optimization of sd-rxRNA efficacy.
[0135] FIG. 54 presents schematics of sterol-type molecules and a graph revealing that sd-rxRNA compounds are fully functional with a variety of linker chemistries. GII asymmetric compounds were synthesized with steroltype molecules attached through TEG and amino caproic acid linkers. Both linkers showed identical potency. This functionality independent of linker chemistry indicates a significant difference between the molecules described herein and previously described molecules, and offers significant advantages for the molecules described herein in terms of scale up and synthesis.
[0136] FIG. 55 demonstrates the stability of chemically modified sd-rxRNA compounds in human serum in comparison to non modified RNA. The oligonucleotides were incubated in 75% serum at 37.degree. C. for the number of hours indicated. The level of degradation was determined by running the samples on non-denaturing gels and staining with SYBGR.
[0137] FIG. 56 is a graph depicting optimization of cellular uptake of sd-rxRNA through minimizing oligonucleotide content.
[0138] FIG. 57 is a graph showing percent MAP4K4 expression after spontaneous cellular uptake of sd-rxRNA in mouse PEC-derived macrophages, and phase and fluorescent images showing localization of sd-rxRNA.
[0139] FIG. 58 is a graph showing percent MAP4K4 expression after spontaneous cellular uptake of sd-rxRNA (targeting) and sd-rxRNA (mismatch) in mouse primary hepatocytes, and phase and fluorescent images showing localization of sd-rxRNA.
[0140] FIG. 59 presents images depicting localization of DY547-labeled sd-rxRNA delivered to RPE cells with no formulation.
[0141] FIG. 60 is a graph showing silencing of MAP4K4 expression in RPE cells treated with sd-rxRNA.sup.nano without formulation.
[0142] FIG. 61 presents a graph and schematics of RNAi compounds showing the chemical/structural composition of highly effective sd-rxRNA compounds. Highly effective compounds were found to have the following characteristics: antisense strands of 17-21 nucleotides, sense strands of 10-15 nucleotides, single-stranded regions that contained 2-12 phosphorothioate modifications, preferentially 6-8 phosphorothioate modifications, and sense strands in which the majority of nucleotides were 2'OMe modified, with or without phosphorothioate modification. Any linker chemistry can be used to attach these molecules to hydrophobic moieties such as cholesterol at the 3' end of the sense strand. Version GIIa-b of these RNA compounds demonstrate that elimination of 2'F content has no impact on efficacy.
[0143] FIG. 62 presents a graph and schematics of RNAi compounds demonstrating the superior performance of sd-rxRNA compounds compared to compounds published by Wolfrum et. al. Nature Biotech, 2007. Both generation I and II compounds (GI and GIIa) developed herein show great efficacy. By contrast, when the chemistry described in Wolfrum et al. (all oligos contain cholesterol conjugated to the 3' end of the sense strand) was applied to the same sequence in a context of conventional siRNA (19 bp duplex with two overhang) the compound was practically inactive. These data emphasize the significance of the combination of chemical modifications and assymetrical molecules described herein, producing highly effective RNA compounds.
[0144] FIG. 63 presents images showing that sd-rxRNA accumulates inside cells while other less effective conjugate RNAs accumulate on the surface of cells.
[0145] FIG. 64 presents images showing that sd-rxRNA molecules, but not other molecules, are internalized into cells within minutes.
[0146] FIG. 65 presents images demonstrating that sd-rxRNA compounds have drastically better cellular and tissue uptake characteristics when compared to conventional cholesterol conjugated siRNAs (such as those published by Soucheck et al). FIG. 65A,B compare uptake in RPE cells, FIG. 65C,D compare uptake upon local administration to skin and FIG. 65E,F compare uptake by the liver upon systemic administration. The level of uptake is at least an order of magnitude higher for the sd-rxRNA compounds relative to the regular siRNA-cholesterol compounds.
[0147] FIG. 66 presents images depicting localization of rxRNA.sup.ori and sd-rxRNA following local delivery.
[0148] FIG. 67 presents images depicting localization of sd-rxRNA and other conjugate RNAs following local delivery.
[0149] FIG. 68 presents a graph revealing the results of a screen performed with sd-rxRNAGII chemistry to identify functional compounds targeting the SPP1 gene. Multiple effective compounds were identified, with 14131 being the most effective. The compounds were added to A-549 cells and the level of the ratio of SPP1/PPIB was determined by B-DNA after 48 hours.
[0150] FIG. 69 presents a graph and several images demonstrating efficient cellular uptake of sd-rxRNA within minutes of exposure. This is a unique characteristics of the sd-rxRNA compounds described herein, not observed with any other RNAi compounds. The Soutschek et al. compound was used as a negative control.
[0151] FIG. 70 presents a graph and several images demonstrating efficient uptake and silencing of sd-rxRNA compounds in multiple cell types with multiple sequences. In each case silencing was confirmed by looking at target gene expression using a Branched DNA assay.
[0152] FIG. 71 presents a graph revealing that sd-rxRNA is active in the presence and absence of serum. A slight reduction in efficacy (2-5 fold) was observed in the presence of serum. This minimal reduction in efficacy in the presence of serum differentiates the sd-rxRNA compounds described herein from previously described RNAi compounds, which had a greater reduction in efficacy, and thus creates a foundation for in vivo efficacy of the sd-rxRNA molecules described herein.
[0153] FIG. 72 presents images demonstrating efficient tissue penetration and cellular uptake upon single intradermal injection of sd-rxRNA compounds described herein. This represents a model for local delivery of sd-rxRNA compounds as well as an effective demonstration of delivery of sd-rxRNA compounds and silencing of genes in dermatological applications.
[0154] FIG. 73 presents images and a graph demonstrating efficient cellular uptake and in vivo silencing with sd-rxRNA following intradermal injection.
[0155] FIG. 74 presents graphs demonstrating that sd-rxRNA compounds have improved blood clearance and induce effective gene silencing in vivo in the liver upon systemic administration.
[0156] FIG. 75 presents a graph demonstrating that the presence of 5-Methyl C in an RNAi compound resulted in an increase in potency of lipid mediated transfection, demonstrating that hydrophobic modification of Cs and Us in the content of RNAi compounds can be beneficial. In some embodiments, these types of modifications can be used in the context of 2' ribose modified bases to insure optimal stability and efficacy.
[0157] FIG. 76 presents a graph showing percent MAP4K4 expression in HeLa cells following passive uptake transfection of: Guide strand alone; Nano Lead; Nano Lead (No cholesterol); Guide Strand w/5MeC and 2'F Us Alone; Nano Lead w/GS 5MeC and 2'F Us; Nano Lead w/GS riboT and 5 Methyl Cs; and Nano Lead w/Guide dT and 5 Methyl Cs.
[0158] FIG. 77 presents images comparing localization of sd-rxRNA and other RNA conjugates following systemic delivery to the liver.
[0159] FIG. 78 presents schematics demonstrating 5-uridyl modifications with improved hydrophobicity characteristics (FIG. 78A). Incorporation of such modifications into sd-rxRNA compounds can increase cellular and tissue uptake properties. FIG. 78B presents a new type of RNAi compound modification which can be applied to compounds to improve cellular uptake and pharmacokinetic behavior. This type of modification, when applied to sd-rxRNA compounds, may contribute to making such compounds orally available.
[0160] FIG. 79 presents schematics revealing the structures of synthesized modified sterol type molecules, where the length and structure of the C17 attached tail is modified. Without wishing to be bound by any theory, the length of the C17 attached tail may contribute to improving in vitro and in vivo efficacy of sd-rxRNA compounds.
[0161] FIG. 80 presents a schematic demonstrating the lithocholic acid route to long side chain cholesterols.
[0162] FIG. 81 presents a schematic demonstrating a route to 5-uridyl phosphoramidite synthesis.
[0163] FIG. 82 presents a schematic demonstrating synthesis of tri-functional hydroxyprolinol linker for 3'-cholesterol attachment.
[0164] FIG. 83 presents a schematic demonstrating synthesis of solid support for the manufacture of a shorter asymmetric RNAi compound strand.
[0165] FIG. 84 demonstrates SPPI sd-rxRNA compound selection. Sd-rxRNA compounds targeting SPP1 were added to A549 cells (using passive transfection) and the level of SPP1 expression was evaluated after 48 hours. Several novel compounds effective in SPP1 silencing were identified, the most potent of which was compound 14131.
[0166] FIG. 85 demonstrates independent validation of sd-rxRNA compounds 14116, 14121, 14131, 14134, 14139, 14149, and 14152 efficacy in SPP1 silencing.
[0167] FIG. 86 demonstrates results of sd-rxRNA compound screens to identify sd-rxRNA compounds functional in CTGF knockdown.
[0168] FIG. 87 demonstrates results of sd-rxRNA compound screens to identify sd-rxRNA functional in CTGF knockdown.
[0169] FIG. 88 demonstrates a systematic screen identifying the minimal length of the asymmetric compounds. The passenger strand of 10-19 bases was hybridized to a guide strand of 17-25 bases. In this assay, compounds with duplex regions as short as 10 bases were found to be effective in inducing.
[0170] FIG. 89 demonstrates that positioning of the sense strand relative to the guide strand is critical for RNAi Activity. In this assay, a blunt end was found to be optimal, a 3' overhang was tolerated, and a 5' overhang resulted in complete loss of functionality.
[0171] FIG. 90 demonstrates that the guide strand, which has homology to the target only at nucleotides 2-17, resulted in effective RNAi when hybridized with sense strands of different lengths. The compounds were introduced into HeLa cells via lipid mediated transfection.
[0172] FIG. 91 is a schematic depicting a panel of sterol-type molecules which can be used as a hydrophobic entity in place of cholesterol. In some instances, the use of sterol-type molecules comprising longer chains results in generation of sd-rxRNA compounds with significantly better cellular uptake and tissue distribution properties.
[0173] FIG. 92 presents schematics depicting a panel of hydrophobic molecules which might be used as a hydrophobic entity in place of cholesterol. These list just provides representative examples; any small molecule with substantial hydrophobicity can be used.
DETAILED DESCRIPTION
[0174] Aspects of the invention relate to methods and compositions involved in gene silencing. The invention is based at least in part on the surprising discovery that asymmetric nucleic acid molecules with a double stranded region of a minimal length such as 8-14 nucleotides, are effective in silencing gene expression. Molecules with such a short double stranded region have not previously been demonstrated to be effective in mediating RNA interference. It had previously been assumed that that there must be a double stranded region of 19 nucleotides or greater. The molecules described herein are optimized through chemical modification, and in some instances through attachment of hydrophobic conjugates.
[0175] The invention is based at least in part on another surprising discovery that asymmetric nucleic acid molecules with reduced double stranded regions are much more effectively taken up by cells compared to conventional siRNAs. These molecules are highly efficient in silencing of target gene expression and offer significant advantages over previously described RNAi molecules including high activity in the presence of serum, efficient self delivery, compatibility with a wide variety of linkers, and reduced presence or complete absence of chemical modifications that are associated with toxicity.
[0176] In contrast to single-stranded polynucleotides, duplex polynucleotides have been difficult to deliver to a cell as they have rigid structures and a large number of negative charges which makes membrane transfer difficult. Unexpectedly, it was found that the polynucleotides of the present invention, although partially double-stranded, are recognized in vivo as single-stranded and, as such, are capable of efficiently being delivered across cell membranes. As a result the polynucleotides of the invention are capable in many instances of self delivery. Thus, the polynucleotides of the invention may be formulated in a manner similar to conventional RNAi agents or they may be delivered to the cell or subject alone (or with non-delivery type carriers) and allowed to self deliver. In one embodiment of the present invention, self delivering asymmetric double-stranded RNA molecules are provided in which one portion of the molecule resembles a conventional RNA duplex and a second portion of the molecule is single stranded.
[0177] The polynucleotides of the invention are referred to herein as isolated double stranded or duplex nucleic acids, oligonucleotides or polynucleotides, nano molecules, nano RNA, sd-rxRNA.sup.nano, sd-rxRNA or RNA molecules of the invention.
[0178] The oligonucleotides of the invention in some aspects have a combination of asymmetric structures including a double stranded region and a single stranded region of 5 nucleotides or longer, specific chemical modification patterns and are conjugated to lipophilic or hydrophobic molecules. This new class of RNAi like compounds have superior efficacy in vitro and in vivo. Based on the data described herein it is believed that the reduction in the size of the rigid duplex region in combination with phosphorothioate modifications applied to a single stranded region are new and important for achieving the observed superior efficacy. Thus, the RNA molecules described herein are different in both structure and composition as well as in vitro and in vivo activity.
[0179] In a preferred embodiment the RNAi compounds of the invention comprise an asymmetric compound comprising a duplex region (required for efficient RISC entry of 10-15 bases long) and single stranded region of 4-12 nucleotides long; with a 13 nucleotide duplex. A 6 nucleotide single stranded region is preferred in some embodiments. The single stranded region of the new RNAi compounds also comprises 2-12 phosphorothioate internucleotide linkages (referred to as phosphorothioate modifications). 6-8 phosphorothioate internucleotide linkages are preferred in some embodiments. Additionally, the RNAi compounds of the invention also include a unique chemical modification pattern, which provides stability and is compatible with RISC entry. The combination of these elements has resulted in unexpected properties which are highly useful for delivery of RNAi reagents in vitro and in vivo.
[0180] The chemically modification pattern, which provides stability and is compatible with RISC entry includes modifications to the sense, or passenger, strand as well as the antisense, or guide, strand. For instance the passenger strand can be modified with any chemical entities which confirm stability and do not interfere with activity. Such modifications include 2' ribo modifications (O-methyl, 2' F, 2 deoxy and others) and backbone modification like phosphorothioate modifications. A preferred chemical modification pattern in the passenger strand includes Omethyl modification of C and U nucleotides within the passenger strand or alternatively the passenger strand may be completely Omethyl modified.
[0181] The guide strand, for example, may also be modified by any chemical modification which confirms stability without interfering with RISC entry. A preferred chemical modification pattern in the guide strand includes the majority of C and U nucleotides being 2' F modified and the 5' end being phosphorylated. Another preferred chemical modification pattern in the guide strand includes 2' Omethyl modification of position 1 and C/U in positions 11-18 and 5' end chemical phosphorylation. Yet another preferred chemical modification pattern in the guide strand includes 2'Omethyl modification of position 1 and C/U in positions 11-18 and 5' end chemical phosphorylation and and 2'F modification of C/U in positions 2-10.
[0182] It was surprisingly discovered according to the invention that the above-described chemical modification patterns of the oligonucleotides of the invention are well tolerated and actually improved efficacy of asymmetric RNAi compounds. See, for instance, FIG. 22.
[0183] It was also demonstrated experimentally herein that the combination of modifications to RNAi when used together in a polynucleotide results in the achievement of optimal efficacy in passive uptake of the RNAi. Elimination of any of the described components (Guide strand stabilization, phosphorothioate stretch, sense strand stabilization and hydrophobic conjugate) or increase in size results in sub-optimal efficacy and in some instances complete lost of efficacy. The combination of elements results in development of compound, which is fully active following passive delivery to cells such as HeLa cells. (FIG. 23). The degree to which the combination of elements results in efficient self delivery of RNAi molecules was completely unexpected.
[0184] The data shown in FIGS. 26, 27 and 43 demonstrated the importance of the various modifications to the RNAi in achieving stabilization and activity. For instance, FIG. 26 demonstrates that use off asymmetric configuration is important in getting efficacy in passive uptake. When the same chemical composition is applied to compounds of traditional configurations (19-21 bases duplex and 25 mer duplex) the efficacy was drastically decreased in a length dependent manner. FIG. 27 demonstrated a systematic screen of the impact of phosphorothioate chemical modifications on activity. The sequence, structure, stabilization chemical modifications, hydrophobic conjugate were kept constant and compound phosphorothioate content was varied (from 0 to 18 PS bond). Both compounds having no phosphorothioate linkages and having 18 phosphorothioate linkages were completely inactive in passive uptake. Compounds having 2-16 phosphorothioate linkages were active, with compounds having 4-10 phosphorothioate being the most active compounds.
[0185] The data in the Examples presented below demonstrates high efficacy of the oligonucleotides of the invention both in vitro in variety of cell types (supporting data) and in vivo upon local and systemic administration. For instance, the data compares the ability of several competitive RNAi molecules having different chemistries to silence a gene. Comparison of sd-rxRNA (oligonucleotides of the invention) with RNAs described in Soucheck et al. and Wolfrum at al., as applied to the same targeting region, demonstrated that only sd-rxRNA chemistry showed a significant functionality in passive uptake. The composition of the invention achieved EC50 values of 10-50 pM. This level of efficacy is un-attainable with conventional chemistries like those described in Sauthceck at al and Accell. Similar comparisons were made in other systems, such as in vitro (RPE cell line), in vivo upon local administration (wounded skin) and systemic (50 mg/kg) as well as other genes (FIGS. 65 and 68). In each case the oligonucleotides of the invention achieved better results. FIG. 64 includes data demonstrating efficient cellular uptake and resulting silencing by sd-rxRNA compounds only after 1 minute of exposure. Such an efficacy is unique to this composition and have not been seen with other types of molecules in this class. FIG. 70 demonstrates efficient uptake and silencing of sd-rxRNA compounds in multiple cell types with multiple sequences. The sd-rxRNA compounds are also active in cells in presence and absence of serum and other biological liquids. FIG. 71 demonstrates only a slight reduction in activity in the presence of serum. This ability to function in biologically aggressive environment effectively further differentiates sd-rxRNA compounds from other compounds described previously in this group, like Accell and Soucheck et al, in which uptake is drastically inhibited in a presence of serum.
[0186] Significant amounts of data also demonstrate the in vivo efficacy of the compounds of the invention. For instance FIGS. 72-74 involve multiple routes of in vivo delivery of the compounds of the invention resulting in significant activity. FIG. 72, for example, demonstrates efficient tissue penetration and cellular uptake upon single intradermal injection. This is a model for local delivery of sd-rxRNA compounds as well as an effective delivery mode for sd-rxRNA compounds and silencing genes in any dermatology applications. FIG. 73 demonstrated efficient tissue penetration, cellular uptake and silencing upon local in vivo intradermal injection of sd-rxRNA compounds. The data of FIG. 74 demonstrate that sd-rxRNA compounds result in highly effective liver uptake upon IV administration. Comparison to Souicheck at al molecule showed that the level of liver uptake at identical dose level was quite surprisingly, at least 50 fold higher with the sd-rxRNA compound than the Souicheck at al molecule.
[0187] The sd-rxRNA can be further improved in some instances by improving the hydrophobicity of compounds using of novel types of chemistries. For example one chemistry is related to use of hydrophobic base modifications. Any base in any position might be modified, as long as modification results in an increase of the partition coefficient of the base. The preferred locations for modification chemistries are positions 4 and 5 of the pyrimidines. The major advantage of these positions is (a) ease of synthesis and (b) lack of interference with base-pairing and A form helix formation, which are essential for RISC complex loading and target recognition. Examples of these chemistries is shown in FIGS. 75-83. A version of sd-rxRNA compounds where multiple deoxy Uridines are present without interfering with overall compound efficacy was used. In addition major improvement in tissue distribution and cellular uptake might be obtained by optimizing the structure of the hydrophobic conjugate. In some of the preferred embodiment the structure of sterol is modified to alter (increase/decrease) C17 attached chain. This type of modification results in significant increase in cellular uptake and improvement of tissue uptake prosperities in vivo.
[0188] This invention is not limited in its application to the details of construction and the arrangement of components set forth in the following description or illustrated in the drawings. The invention is capable of other embodiments and of being practiced or of being carried out in various ways. Also, the phraseology and terminology used herein is for the purpose of description and should not be regarded as limiting. The use of "including," "comprising," or "having," "containing," "involving," and variations thereof herein, is meant to encompass the items listed thereafter and equivalents thereof as well as additional items.
[0189] Thus, aspects of the invention relate to isolated double stranded nucleic acid molecules comprising a guide (antisense) strand and a passenger (sense) strand. As used herein, the term "double-stranded" refers to one or more nucleic acid molecules in which at least a portion of the nucleomonomers are complementary and hydrogen bond to form a double-stranded region. In some embodiments, the length of the guide strand ranges from 16-29 nucleotides long. In certain embodiments, the guide strand is 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, or 29 nucleotides long. The guide strand has complementarity to a target gene. Complementarity between the guide strand and the target gene may exist over any portion of the guide strand. Complementarity as used herein may be perfect complementarity or less than perfect complementarity as long as the guide strand is sufficiently complementary to the target that it mediates RNAi. In some embodiments complementarity refers to less than 25%, 20%, 15%, 10%, 5%, 4%, 3%, 2%, or 1% mismatch between the guide strand and the target. Perfect complementarity refers to 100% complementarity. Thus the invention has the advantage of being able to tolerate sequence variations that might be expected due to genetic mutation, strain polymorphism, or evolutionary divergence. For example, siRNA sequences with insertions, deletions, and single point mutations relative to the target sequence have also been found to be effective for inhibition. Moreover, not all positions of a siRNA contribute equally to target recognition. Mismatches in the center of the siRNA are most critical and essentially abolish target RNA cleavage. Mismatches upstream of the center or upstream of the cleavage site referencing the antisense strand are tolerated but significantly reduce target RNA cleavage. Mismatches downstream of the center or cleavage site referencing the antisense strand, preferably located near the 3' end of the antisense strand, e.g. 1, 2, 3, 4, 5 or 6 nucleotides from the 3' end of the antisense strand, are tolerated and reduce target RNA cleavage only slightly.
[0190] While not wishing to be bound by any particular theory, in some embodiments, the guide strand is at least 16 nucleotides in length and anchors the Argonaute protein in RISC. In some embodiments, when the guide strand loads into RISC it has a defined seed region and target mRNA cleavage takes place across from position 10-11 of the guide strand. In some embodiments, the 5' end of the guide strand is or is able to be phosphorylated. The nucleic acid molecules described herein may be referred to as minimum trigger RNA.
[0191] In some embodiments, the length of the passenger strand ranges from 8-14 nucleotides long. In certain embodiments, the passenger strand is 8, 9, 10, 11, 12, 13 or 14 nucleotides long. The passenger strand has complementarity to the guide strand. Complementarity between the passenger strand and the guide strand can exist over any portion of the passenger or guide strand. In some embodiments, there is 100% complementarity between the guide and passenger strands within the double stranded region of the molecule.
[0192] Aspects of the invention relate to double stranded nucleic acid molecules with minimal double stranded regions. In some embodiments the region of the molecule that is double stranded ranges from 8-14 nucleotides long. In certain embodiments, the region of the molecule that is double stranded is 8, 9, 10, 11, 12, 13 or 14 nucleotides long. In certain embodiments the double stranded region is 13 nucleotides long. There can be 100% complementarity between the guide and passenger strands, or there may be one or more mismatches between the guide and passenger strands. In some embodiments, on one end of the double stranded molecule, the molecule is either blunt-ended or has a one-nucleotide overhang. The single stranded region of the molecule is in some embodiments between 4-12 nucleotides long. For example the single stranded region can be 4, 5, 6, 7, 8, 9, 10, 11 or 12 nucleotides long. However, in certain embodiments, the single stranded region can also be less than 4 or greater than 12 nucleotides long. In certain embodiments, the single stranded region is 6 nucleotides long.
[0193] RNAi constructs associated with the invention can have a thermodynamic stability (.DELTA.G) of less than -13 kkal/mol. In some embodiments, the thermodynamic stability (.DELTA.G) is less than -20 kkal/mol. In some embodiments there is a loss of efficacy when (.DELTA.G) goes below -21 kkal/mol. In some embodiments a (.DELTA.G) value higher than -13 kkal/mol is compatible with aspects of the invention. Without wishing to be bound by any theory, in some embodiments a molecule with a relatively higher (.DELTA.G) value may become active at a relatively higher concentration, while a molecule with a relatively lower (.DELTA.G) value may become active at a relatively lower concentration. In some embodiments, the (.DELTA.G) value may be higher than -9 kkcal/mol. The gene silencing effects mediated by the RNAi constructs associated with the invention, containing minimal double stranded regions, are unexpected because molecules of almost identical design but lower thermodynamic stability have been demonstrated to be inactive (Rana et al 2004).
[0194] Without wishing to be bound by any theory, results described herein suggest that a stretch of 8-10 bp of dsRNA or dsDNA will be structurally recognized by protein components of RISC or co-factors of RISC. Additionally, there is a free energy requirement for the triggering compound that it may be either sensed by the protein components and/or stable enough to interact with such components so that it may be loaded into the Argonaute protein. If optimal thermodynamics are present and there is a double stranded portion that is preferably at least 8 nucleotides then the duplex will be recognized and loaded into the RNAi machinery.
[0195] In some embodiments, thermodynamic stability is increased through the use of LNA bases. In some embodiments, additional chemical modifications are introduced. Several non-limiting examples of chemical modifications include: 5' Phosphate, 2'-O-methyl, 2'-O-ethyl, 2'-fluoro, ribothymidine, C-5 propynyl-dC (pdC) and C-5 propynyl-dU (pdU); C-5 propynyl-C(pC) and C-5 propynyl-U (pU); 5-methyl C, 5-methyl U, 5-methyl dC, 5-methyl dU methoxy, (2,6-diaminopurine), 5'-Dimethoxytrityl-N4-ethyl-2'-deoxyCytidine and MGB (minor groove binder). It should be appreciated that more than one chemical modification can be combined within the same molecule.
[0196] Molecules associated with the invention are optimized for increased potency and/or reduced toxicity. For example, nucleotide length of the guide and/or passenger strand, and/or the number of phosphorothioate modifications in the guide and/or passenger strand, can in some aspects influence potency of the RNA molecule, while replacing 2'-fluoro (2'F) modifications with 2'-O-methyl (2'OMe) modifications can in some aspects influence toxicity of the molecule. Specifically, reduction in 2'F content of a molecule is predicted to reduce toxicity of the molecule. The Examples section presents molecules in which 2'F modifications have been eliminated, offering an advantage over previously described RNAi compounds due to a predicted reduction in toxicity. Furthermore, the number of phosphorothioate modifications in an RNA molecule can influence the uptake of the molecule into a cell, for example the efficiency of passive uptake of the molecule into a cell. Preferred embodiments of molecules described herein have no 2'F modification and yet are characterized by equal efficacy in cellular uptake and tissue penetration. Such molecules represent a significant improvement over prior art, such as molecules described by Accell and Wolfrum, which are heavily modified with extensive use of 2'F.
[0197] In some embodiments, a guide strand is approximately 18-19 nucleotides in length and has approximately 2-14 phosphate modifications. For example, a guide strand can contain 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 or more than 14 nucleotides that are phosphate-modified. The guide strand may contain one or more modifications that confer increased stability without interfering with RISC entry. The phosphate modified nucleotides, such as phosphorothioate modified nucleotides, can be at the 3' end, 5' end or spread throughout the guide strand. In some embodiments, the 3' terminal 10 nucleotides of the guide strand contains 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 phosphorothioate modified nucleotides. The guide strand can also contain 2'F and/or 2'OMe modifications, which can be located throughout the molecule. In some embodiments, the nucleotide in position one of the guide strand (the nucleotide in the most 5' position of the guide strand) is 2'OMe modified and/or phosphorylated. C and U nucleotides within the guide strand can be 2'F modified. For example, C and U nucleotides in positions 2-10 of a 19 nt guide strand (or corresponding positions in a guide strand of a different length) can be 2'F modified. C and U nucleotides within the guide strand can also be 2'OMe modified. For example, C and U nucleotides in positions 11-18 of a 19 nt guide strand (or corresponding positions in a guide strand of a different length) can be 2'OMe modified. In some embodiments, the nucleotide at the most 3' end of the guide strand is unmodified. In certain embodiments, the majority of Cs and Us within the guide strand are 2'F modified and the 5' end of the guide strand is phosphorylated. In other embodiments, position 1 and the Cs or Us in positions 11-18 are 2'OMe modified and the 5' end of the guide strand is phosphorylated. In other embodiments, position 1 and the Cs or Us in positions 11-18 are 2'OMe modified, the 5' end of the guide strand is phosphorylated, and the Cs or Us in position 2-10 are 2'F modified.
[0198] In some aspects, an optimal passenger strand is approximately 11-14 nucleotides in length. The passenger strand may contain modifications that confer increased stability. One or more nucleotides in the passenger strand can be 2'OMe modified. In some embodiments, one or more of the C and/or U nucleotides in the passenger strand is 2'OMe modified, or all of the C and U nucleotides in the passenger strand are 2'OMe modified. In certain embodiments, all of the nucleotides in the passenger strand are 2'OMe modified. One or more of the nucleotides on the passenger strand can also be phosphate-modified such as phosphorothioate modified. The passenger strand can also contain 2' ribo, 2'F and 2 deoxy modifications or any combination of the above. As demonstrated in the Examples, chemical modification patterns on both the guide and passenger strand are well tolerated and a combination of chemical modifications is shown herein to lead to increased efficacy and self-delivery of RNA molecules.
[0199] Aspects of the invention relate to RNAi constructs that have extended single-stranded regions relative to double stranded regions, as compared to molecules that have been used previously for RNAi. The single stranded region of the molecules may be modified to promote cellular uptake or gene silencing. In some embodiments, phosphorothioate modification of the single stranded region influences cellular uptake and/or gene silencing. The region of the guide strand that is phosphorothioate modified can include nucleotides within both the single stranded and double stranded regions of the molecule. In some embodiments, the single stranded region includes 2-12 phosphorothioate modifications. For example, the single stranded region can include 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, or 12 phosphorothioate modifications. In some instances, the single stranded region contains 6-8 phosphorothioate modifications.
[0200] Molecules associated with the invention are also optimized for cellular uptake. In RNA molecules described herein, the guide and/or passenger strands can be attached to a conjugate. In certain embodiments the conjugate is hydrophobic. The hydrophobic conjugate can be a small molecule with a partition coefficient that is higher than 10. The conjugate can be a sterol-type molecule such as cholesterol, or a molecule with an increased length polycarbon chain attached to C17, and the presence of a conjugate can influence the ability of an RNA molecule to be taken into a cell with or without a lipid transfection reagent. The conjugate can be attached to the passenger or guide strand through a hydrophobic linker. In some embodiments, a hydrophobic linker is 5-12C in length, and/or is hydroxypyrrolidine-based. In some embodiments, a hydrophobic conjugate is attached to the passenger strand and the CU residues of either the passenger and/or guide strand are modified. In some embodiments, at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90% or 95% of the CU residues on the passenger strand and/or the guide strand are modified. In some aspects, molecules associated with the invention are self-delivering (sd). As used herein, "self-delivery" refers to the ability of a molecule to be delivered into a cell without the need for an additional delivery vehicle such as a transfection reagent.
[0201] Aspects of the invention relate to selecting molecules for use in RNAi. Based on the data described herein, molecules that have a double stranded region of 8-14 nucleotides can be selected for use in RNAi. In some embodiments, molecules are selected based on their thermodynamic stability (.DELTA.G). In some embodiments, molecules will be selected that have a (.DELTA.G) of less than -13 kkal/mol. For example, the (.DELTA.G) value may be -13, -14, -15, -16, -17, -18, -19, -21, -22 or less than -22 kkal/mol. In other embodiments, the (.DELTA.G) value may be higher than -13 kkal/mol. For example, the (.DELTA.G) value may be -12, -11, -10, -9, -8, -7 or more than -7 kkal/mol. It should be appreciated that .DELTA.G can be calculated using any method known in the art. In some embodiments .DELTA.G is calculated using Mfold, available through the Mfold internet site (http://mfold.bioinfo.rpi.edu/cgi-bin/rna-form1.cfi). Methods for calculating .DELTA.G are described in, and are incorporated by reference from, the following references: Zuker, M. (2003) Nucleic Acids Res., 31(13):3406-15; Mathews, D. H., Sabina, J., Zuker, M. and Turner, D. H. (1999) J. Mol. Biol. 288:911-940; Mathews, D. H., Disney, M. D., Childs, J. L., Schroeder, S. J., Zuker, M., and Turner, D. H. (2004) Proc. Natl. Acad. Sci. 101:7287-7292; Duan, S., Mathews, D. H., and Turner, D. H. (2006) Biochemistry 45:9819-9832; Wuchty, S., Fontana, W., Hofacker, I. L., and Schuster, P. (1999) Biopolymers 49:145-165.
[0202] Aspects of the invention relate to using nucleic acid molecules described herein, with minimal double stranded regions and/or with a (.DELTA.G) of less than -13 kkal/mol, for gene silencing. RNAi molecules can be administered in vivo or in vitro, and gene silencing effects can be achieved in vivo or in vitro.
[0203] In certain embodiments, the polynucleotide contains 5'- and/or 3'-end overhangs. The number and/or sequence of nucleotides overhang on one end of the polynucleotide may be the same or different from the other end of the polynucleotide. In certain embodiments, one or more of the overhang nucleotides may contain chemical modification(s), such as phosphorothioate or 2'-OMe modification.
[0204] In certain embodiments, the polynucleotide is unmodified. In other embodiments, at least one nucleotide is modified. In further embodiments, the modification includes a 2'-H or 2'-modified ribose sugar at the 2nd nucleotide from the 5'-end of the guide sequence. The "2nd nucleotide" is defined as the second nucleotide from the 5'-end of the polynucleotide.
[0205] As used herein, "2'-modified ribose sugar" includes those ribose sugars that do not have a 2'-OH group. "2'-modified ribose sugar" does not include 2'-deoxyribose (found in unmodified canonical DNA nucleotides). For example, the 2'-modified ribose sugar may be 2'-O-alkyl nucleotides, 2'-deoxy-2'-fluoro nucleotides, 2'-deoxy nucleotides, or combination thereof.
[0206] In certain embodiments, the 2'-modified nucleotides are pyrimidine nucleotides (e.g., C/U). Examples of 2'-O-alkyl nucleotides include 2'-O-methyl nucleotides, or 2'-O-allyl nucleotides.
[0207] In certain embodiments, the miniRNA polynucleotide of the invention with the above-referenced 5'-end modification exhibits significantly (e.g., at least about 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90% or more) less "off-target" gene silencing when compared to similar constructs without the specified 5'-end modification, thus greatly improving the overall specificity of the RNAi reagent or therapeutics.
[0208] As used herein, "off-target" gene silencing refers to unintended gene silencing due to, for example, spurious sequence homology between the antisense (guide) sequence and the unintended target mRNA sequence.
[0209] According to this aspect of the invention, certain guide strand modifications further increase nuclease stability, and/or lower interferon induction, without significantly decreasing RNAi activity (or no decrease in RNAi activity at all).
[0210] In some embodiments, the 5'-stem sequence may comprise a 2'-modified ribose sugar, such as 2'-O-methyl modified nucleotide, at the 2.sup.nd nucleotide on the 5'-end of the polynucleotide and, in some embodiments, no other modified nucleotides. The hairpin structure having such modification may have enhanced target specificity or reduced off-target silencing compared to a similar construct without the 2'-O-methyl modification at said position.
[0211] Certain combinations of specific 5'-stem sequence and 3'-stem sequence modifications may result in further unexpected advantages, as partly manifested by enhanced ability to inhibit target gene expression, enhanced serum stability, and/or increased target specificity, etc.
[0212] In certain embodiments, the guide strand comprises a 2'-O-methyl modified nucleotide at the 2.sup.nd nucleotide on the 5'-end of the guide strand and no other modified nucleotides.
[0213] In other aspects, the miniRNA structures of the present invention mediates sequence-dependent gene silencing by a microRNA mechanism. As used herein, the term "microRNA" ("miRNA"), also referred to in the art as "small temporal RNAs" ("stRNAs"), refers to a small (10-50 nucleotide) RNA which are genetically encoded (e.g., by viral, mammalian, or plant genomes) and are capable of directing or mediating RNA silencing. An "miRNA disorder" shall refer to a disease or disorder characterized by an aberrant expression or activity of an miRNA.
[0214] microRNAs are involved in down-regulating target genes in critical pathways, such as development and cancer, in mice, worms and mammals. Gene silencing through a microRNA mechanism is achieved by specific yet imperfect base-pairing of the miRNA and its target messenger RNA (mRNA). Various mechanisms may be used in microRNA-mediated down-regulation of target mRNA expression.
[0215] miRNAs are noncoding RNAs of approximately 22 nucleotides which can regulate gene expression at the post transcriptional or translational level during plant and animal development. One common feature of miRNAs is that they are all excised from an approximately 70 nucleotide precursor RNA stem-loop termed pre-miRNA, probably by Dicer, an RNase III-type enzyme, or a homolog thereof. Naturally-occurring miRNAs are expressed by endogenous genes in vivo and are processed from a hairpin or stem-loop precursor (pre-miRNA or pri-miRNAs) by Dicer or other RNAses. miRNAs can exist transiently in vivo as a double-stranded duplex but only one strand is taken up by the RISC complex to direct gene silencing.
[0216] In some embodiments a version of sd-rxRNA compounds, which are effective in cellular uptake and inhibiting of miRNA activity are described. Essentially the compounds are similar to RISC entering version but large strand chemical modification patterns are optimized in the way to block cleavage and act as an effective inhibitor of the RISC action. For example, the compound might be completely or mostly Omethyl modified with the PS content described previously. For these types of compounds the 5' phosphorilation is not necessary. The presence of double stranded region is preferred as it is promotes cellular uptake and efficient RISC loading.
[0217] Another pathway that uses small RNAs as sequence-specific regulators is the RNA interference (RNAi) pathway, which is an evolutionarily conserved response to the presence of double-stranded RNA (dsRNA) in the cell. The dsRNAs are cleaved into .about.20-base pair (bp) duplexes of small-interfering RNAs (siRNAs) by Dicer. These small RNAs get assembled into multiprotein effector complexes called RNA-induced silencing complexes (RISCs). The siRNAs then guide the cleavage of target mRNAs with perfect complementarity.
[0218] Some aspects of biogenesis, protein complexes, and function are shared between the siRNA pathway and the miRNA pathway. The subject single-stranded polynucleotides may mimic the dsRNA in the siRNA mechanism, or the microRNA in the miRNA mechanism.
[0219] In certain embodiments, the modified RNAi constructs may have improved stability in serum and/or cerebral spinal fluid compared to an unmodified RNAi constructs having the same sequence.
[0220] In certain embodiments, the structure of the RNAi construct does not induce interferon response in primary cells, such as mammalian primary cells, including primary cells from human, mouse and other rodents, and other non-human mammals. In certain embodiments, the RNAi construct may also be used to inhibit expression of a target gene in an invertebrate organism.
[0221] To further increase the stability of the subject constructs in vivo, the 3'-end of the hairpin structure may be blocked by protective group(s). For example, protective groups such as inverted nucleotides, inverted abasic moieties, or amino-end modified nucleotides may be used. Inverted nucleotides may comprise an inverted deoxynucleotide. Inverted abasic moieties may comprise an inverted deoxyabasic moiety, such as a 3',3'-linked or 5',5'-linked deoxyabasic moiety.
[0222] The RNAi constructs of the invention are capable of inhibiting the synthesis of any target protein encoded by target gene(s). The invention includes methods to inhibit expression of a target gene either in a cell in vitro, or in vivo. As such, the RNAi constructs of the invention are useful for treating a patient with a disease characterized by the overexpression of a target gene.
[0223] The target gene can be endogenous or exogenous (e.g., introduced into a cell by a virus or using recombinant DNA technology) to a cell. Such methods may include introduction of RNA into a cell in an amount sufficient to inhibit expression of the target gene. By way of example, such an RNA molecule may have a guide strand that is complementary to the nucleotide sequence of the target gene, such that the composition inhibits expression of the target gene.
[0224] The invention also relates to vectors expressing the nucleic acids of the invention, and cells comprising such vectors or the nucleic acids. The cell may be a mammalian cell in vivo or in culture, such as a human cell.
[0225] The invention further relates to compositions comprising the subject RNAi constructs, and a pharmaceutically acceptable carrier or diluent.
[0226] Another aspect of the invention provides a method for inhibiting the expression of a target gene in a mammalian cell, comprising contacting the mammalian cell with any of the subject RNAi constructs.
[0227] The method may be carried out in vitro, ex vivo, or in vivo, in, for example, mammalian cells in culture, such as a human cell in culture.
[0228] The target cells (e.g., mammalian cell) may be contacted in the presence of a delivery reagent, such as a lipid (e.g., a cationic lipid) or a liposome.
[0229] Another aspect of the invention provides a method for inhibiting the expression of a target gene in a mammalian cell, comprising contacting the mammalian cell with a vector expressing the subject RNAi constructs.
[0230] In one aspect of the invention, a longer duplex polynucleotide is provided, including a first polynucleotide that ranges in size from about 16 to about 30 nucleotides; a second polynucleotide that ranges in size from about 26 to about 46 nucleotides, wherein the first polynucleotide (the antisense strand) is complementary to both the second polynucleotide (the sense strand) and a target gene, and wherein both polynucleotides form a duplex and wherein the first polynucleotide contains a single stranded region longer than 6 bases in length and is modified with alternative chemical modification pattern, and/or includes a conjugate moiety that facilitates cellular delivery. In this embodiment, between about 40% to about 90% of the nucleotides of the passenger strand between about 40% to about 90% of the nucleotides of the guide strand, and between about 40% to about 90% of the nucleotides of the single stranded region of the first polynucleotide are chemically modified nucleotides.
[0231] In an embodiment, the chemically modified nucleotide in the polynucleotide duplex may be any chemically modified nucleotide known in the art, such as those discussed in detail above. In a particular embodiment, the chemically modified nucleotide is selected from the group consisting of 2' F modified nucleotides, 2'-O-methyl modified and 2'deoxy nucleotides. In another particular embodiment, the chemically modified nucleotides results from "hydrophobic modifications" of the nucleotide base. In another particular embodiment, the chemically modified nucleotides are phosphorothioates. In an additional particular embodiment, chemically modified nucleotides are combination of phosphorothioates, 2'-O-methyl, 2'deoxy, hydrophobic modifications and phosphorothioates. As these groups of modifications refer to modification of the ribose ring, back bone and nucleotide, it is feasible that some modified nucleotides will carry a combination of all three modification types.
[0232] In another embodiment, the chemical modification is not the same across the various regions of the duplex. In a particular embodiment, the first polynucleotide (the passenger strand), has a large number of diverse chemical modifications in various positions. For this polynucleotide up to 90% of nucleotides might be chemically modified and/or have mismatches introduced.
[0233] In another embodiment, chemical modifications of the first or second polynucleotide include, but not limited to, 5' position modification of Uridine and Cytosine (4-pyridyl, 2-pyridyl, indolyl, phenyl (C.sub.6H.sub.5OH); tryptophanyl (C8H6N)CH2CH(NH2)CO), isobutyl, butyl, aminobenzyl; phenyl; naphthyl, etc), where the chemical modification might alter base pairing capabilities of a nucleotide. For the guide strand an important feature of this aspect of the invention is the position of the chemical modification relative to the 5' end of the antisense and sequence. For example, chemical phosphorylation of the 5' end of the guide strand is usually beneficial for efficacy. O-methyl modifications in the seed region of the sense strand (position 2-7 relative to the 5' end) are not generally well tolerated, whereas 2'F and deoxy are well tolerated. The mid part of the guide strand and the 3' end of the guide strand are more permissive in a type of chemical modifications applied. Deoxy modifications are not tolerated at the 3' end of the guide strand.
[0234] A unique feature of this aspect of the invention involves the use of hydrophobic modification on the bases. In one embodiment, the hydrophobic modifications are preferably positioned near the 5' end of the guide strand, in other embodiments, they localized in the middle of the guides strand, in other embodiment they localized at the 3' end of the guide strand and yet in another embodiment they are distributed thought the whole length of the polynucleotide. The same type of patterns is applicable to the passenger strand of the duplex.
[0235] The other part of the molecule is a single stranded region. The single stranded region is expected to range from 7 to 40 nucleotides.
[0236] In one embodiment, the single stranded region of the first polynucleotide contains modifications selected from the group consisting of between 40% and 90% hydrophobic base modifications, between 40%-90% phosphorothioates, between 40%-90% modification of the ribose moiety, and any combination of the preceding.
[0237] Efficiency of guide strand (first polynucleotide) loading into the RISC complex might be altered for heavily modified polynucleotides, so in one embodiment, the duplex polynucleotide includes a mismatch between nucleotide 9, 11, 12, 13, or 14 on the guide strand (first polynucleotide) and the opposite nucleotide on the sense strand (second polynucleotide) to promote efficient guide strand loading.
[0238] More detailed aspects of the invention are described in the sections below.
Duplex Characteristics
[0239] Double-stranded oligonucleotides of the invention may be formed by two separate complementary nucleic acid strands. Duplex formation can occur either inside or outside the cell containing the target gene.
[0240] As used herein, the term "duplex" includes the region of the double-stranded nucleic acid molecule(s) that is (are) hydrogen bonded to a complementary sequence. Double-stranded oligonucleotides of the invention may comprise a nucleotide sequence that is sense to a target gene and a complementary sequence that is antisense to the target gene. The sense and antisense nucleotide sequences correspond to the target gene sequence, e.g., are identical or are sufficiently identical to effect target gene inhibition (e.g., are about at least about 98% identical, 96% identical, 94%, 90% identical, 85% identical, or 80% identical) to the target gene sequence.
[0241] In certain embodiments, the double-stranded oligonucleotide of the invention is double-stranded over its entire length, i.e., with no overhanging single-stranded sequence at either end of the molecule, i.e., is blunt-ended. In other embodiments, the individual nucleic acid molecules can be of different lengths. In other words, a double-stranded oligonucleotide of the invention is not double-stranded over its entire length. For instance, when two separate nucleic acid molecules are used, one of the molecules, e.g., the first molecule comprising an antisense sequence, can be longer than the second molecule hybridizing thereto (leaving a portion of the molecule single-stranded). Likewise, when a single nucleic acid molecule is used a portion of the molecule at either end can remain single-stranded.
[0242] In one embodiment, a double-stranded oligonucleotide of the invention contains mismatches and/or loops or bulges, but is double-stranded over at least about 70% of the length of the oligonucleotide. In another embodiment, a double-stranded oligonucleotide of the invention is double-stranded over at least about 80% of the length of the oligonucleotide. In another embodiment, a double-stranded oligonucleotide of the invention is double-stranded over at least about 90%-95% of the length of the oligonucleotide. In another embodiment, a double-stranded oligonucleotide of the invention is double-stranded over at least about 96%-98% of the length of the oligonucleotide. In certain embodiments, the double-stranded oligonucleotide of the invention contains at least or up to 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, or 15 mismatches.
Modifications
[0243] The nucleotides of the invention may be modified at various locations, including the sugar moiety, the phosphodiester linkage, and/or the base.
[0244] Sugar moieties include natural, unmodified sugars, e.g., monosaccharide (such as pentose, e.g., ribose, deoxyribose), modified sugars and sugar analogs. In general, possible modifications of nucleomonomers, particularly of a sugar moiety, include, for example, replacement of one or more of the hydroxyl groups with a halogen, a heteroatom, an aliphatic group, or the functionalization of the hydroxyl group as an ether, an amine, a thiol, or the like.
[0245] One particularly useful group of modified nucleomonomers are 2'-O-methyl nucleotides. Such 2'-O-methyl nucleotides may be referred to as "methylated," and the corresponding nucleotides may be made from unmethylated nucleotides followed by alkylation or directly from methylated nucleotide reagents. Modified nucleomonomers may be used in combination with unmodified nucleomonomers. For example, an oligonucleotide of the invention may contain both methylated and unmethylated nucleomonomers.
[0246] Some exemplary modified nucleomonomers include sugar- or backbone-modified ribonucleotides. Modified ribonucleotides may contain a non-naturally occurring base (instead of a naturally occurring base), such as uridines or cytidines modified at the 5'-position, e.g., 5'-(2-amino)propyl uridine and 5'-bromo uridine; adenosines and guanosines modified at the 8-position, e.g., 8-bromo guanosine; deaza nucleotides, e.g., 7-deaza-adenosine; and N-alkylated nucleotides, e.g., N6-methyl adenosine. Also, sugar-modified ribonucleotides may have the 2'-OH group replaced by a H, alxoxy (or OR), R or alkyl, halogen, SH, SR, amino (such as NH2, NHR, NR.sub.2,), or CN group, wherein R is lower alkyl, alkenyl, or alkynyl.
[0247] Modified ribonucleotides may also have the phosphodiester group connecting to adjacent ribonucleotides replaced by a modified group, e.g., of phosphorothioate group. More generally, the various nucleotide modifications may be combined.
[0248] Although the antisense (guide) strand may be substantially identical to at least a portion of the target gene (or genes), at least with respect to the base pairing properties, the sequence need not be perfectly identical to be useful, e.g., to inhibit expression of a target gene's phenotype. Generally, higher homology can be used to compensate for the use of a shorter antisense gene. In some cases, the antisense strand generally will be substantially identical (although in antisense orientation) to the target gene.
[0249] The use of 2'-O-methyl modified RNA may also be beneficial in circumstances in which it is desirable to minimize cellular stress responses. RNA having 2'-O-methyl nucleomonomers may not be recognized by cellular machinery that is thought to recognize unmodified RNA. The use of 2'-O-methylated or partially 2'-O-methylated RNA may avoid the interferon response to double-stranded nucleic acids, while maintaining target RNA inhibition. This may be useful, for example, for avoiding the interferon or other cellular stress responses, both in short RNAi (e.g., siRNA) sequences that induce the interferon response, and in longer RNAi sequences that may induce the interferon response.
[0250] Overall, modified sugars may include D-ribose, 2'-O-alkyl (including 2'-O-methyl and 2'-O-ethyl), i.e., 2'-alkoxy, 2'-amino, 2'-S-alkyl, 2'-halo (including 2'-fluoro), 2'-methoxyethoxy, 2'-allyloxy (--OCH.sub.2CH.dbd.CH.sub.2), 2'-propargyl, 2'-propyl, ethynyl, ethenyl, propenyl, and cyano and the like. In one embodiment, the sugar moiety can be a hexose and incorporated into an oligonucleotide as described (Augustyns, K., et al., Nucl. Acids. Res. 18:4711 (1992)). Exemplary nucleomonomers can be found, e.g., in U.S. Pat. No. 5,849,902, incorporated by reference herein.
[0251] The term "alkyl" includes saturated aliphatic groups, including straight-chain alkyl groups (e.g., methyl, ethyl, propyl, butyl, pentyl, hexyl, heptyl, octyl, nonyl, decyl, etc.), branched-chain alkyl groups (isopropyl, tert-butyl, isobutyl, etc.), cycloalkyl (alicyclic) groups (cyclopropyl, cyclopentyl, cyclohexyl, cycloheptyl, cyclooctyl), alkyl substituted cycloalkyl groups, and cycloalkyl substituted alkyl groups. In certain embodiments, a straight chain or branched chain alkyl has 6 or fewer carbon atoms in its backbone (e.g., C.sub.1-C.sub.6 for straight chain, C.sub.3-C.sub.6 for branched chain), and more preferably 4 or fewer. Likewise, preferred cycloalkyls have from 3-8 carbon atoms in their ring structure, and more preferably have 5 or 6 carbons in the ring structure. The term C.sub.1-C.sub.6 includes alkyl groups containing 1 to 6 carbon atoms.
[0252] Moreover, unless otherwise specified, the term alkyl includes both "unsubstituted alkyls" and "substituted alkyls," the latter of which refers to alkyl moieties having independently selected substituents replacing a hydrogen on one or more carbons of the hydrocarbon backbone. Such substituents can include, for example, alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, cyano, amino (including alkyl amino, dialkylamino, arylamino, diarylamino, and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety. Cycloalkyls can be further substituted, e.g., with the substituents described above. An "alkylaryl" or an "arylalkyl" moiety is an alkyl substituted with an aryl (e.g., phenylmethyl (benzyl)). The term "alkyl" also includes the side chains of natural and unnatural amino acids. The term "n-alkyl" means a straight chain (i.e., unbranched) unsubstituted alkyl group.
[0253] The term "alkenyl" includes unsaturated aliphatic groups analogous in length and possible substitution to the alkyls described above, but that contain at least one double bond. For example, the term "alkenyl" includes straight-chain alkenyl groups (e.g., ethylenyl, propenyl, butenyl, pentenyl, hexenyl, heptenyl, octenyl, nonenyl, decenyl, etc.), branched-chain alkenyl groups, cycloalkenyl (alicyclic) groups (cyclopropenyl, cyclopentenyl, cyclohexenyl, cycloheptenyl, cyclooctenyl), alkyl or alkenyl substituted cycloalkenyl groups, and cycloalkyl or cycloalkenyl substituted alkenyl groups. In certain embodiments, a straight chain or branched chain alkenyl group has 6 or fewer carbon atoms in its backbone (e.g., C.sub.2-C.sub.6 for straight chain, C.sub.3-C.sub.6 for branched chain). Likewise, cycloalkenyl groups may have from 3-8 carbon atoms in their ring structure, and more preferably have 5 or 6 carbons in the ring structure. The term C.sub.2-C.sub.6 includes alkenyl groups containing 2 to 6 carbon atoms.
[0254] Moreover, unless otherwise specified, the term alkenyl includes both "unsubstituted alkenyls" and "substituted alkenyls," the latter of which refers to alkenyl moieties having independently selected substituents replacing a hydrogen on one or more carbons of the hydrocarbon backbone. Such substituents can include, for example, alkyl groups, alkynyl groups, halogens, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, cyano, amino (including alkyl amino, dialkylamino, arylamino, diarylamino, and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety.
[0255] The term "alkynyl" includes unsaturated aliphatic groups analogous in length and possible substitution to the alkyls described above, but which contain at least one triple bond. For example, the term "alkynyl" includes straight-chain alkynyl groups (e.g., ethynyl, propynyl, butynyl, pentynyl, hexynyl, heptynyl, octynyl, nonynyl, decynyl, etc.), branched-chain alkynyl groups, and cycloalkyl or cycloalkenyl substituted alkynyl groups. In certain embodiments, a straight chain or branched chain alkynyl group has 6 or fewer carbon atoms in its backbone (e.g., C.sub.2-C.sub.6 for straight chain, C.sub.3-C.sub.6 for branched chain). The term C.sub.2-C.sub.6 includes alkynyl groups containing 2 to 6 carbon atoms.
[0256] Moreover, unless otherwise specified, the term alkynyl includes both "unsubstituted alkynyls" and "substituted alkynyls," the latter of which refers to alkynyl moieties having independently selected substituents replacing a hydrogen on one or more carbons of the hydrocarbon backbone. Such substituents can include, for example, alkyl groups, alkynyl groups, halogens, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, cyano, amino (including alkyl amino, dialkylamino, arylamino, diarylamino, and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety.
[0257] Unless the number of carbons is otherwise specified, "lower alkyl" as used herein means an alkyl group, as defined above, but having from one to five carbon atoms in its backbone structure. "Lower alkenyl" and "lower alkynyl" have chain lengths of, for example, 2-5 carbon atoms.
[0258] The term "alkoxy" includes substituted and unsubstituted alkyl, alkenyl, and alkynyl groups covalently linked to an oxygen atom. Examples of alkoxy groups include methoxy, ethoxy, isopropyloxy, propoxy, butoxy, and pentoxy groups. Examples of substituted alkoxy groups include halogenated alkoxy groups. The alkoxy groups can be substituted with independently selected groups such as alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, cyano, amino (including alkyl amino, dialkylamino, arylamino, diarylamino, and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulffiydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfmyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moieties. Examples of halogen substituted alkoxy groups include, but are not limited to, fluoromethoxy, difluoromethoxy, trifluoromethoxy, chloromethoxy, dichloromethoxy, trichloromethoxy, etc.
[0259] The term "heteroatom" includes atoms of any element other than carbon or hydrogen. Preferred heteroatoms are nitrogen, oxygen, sulfur and phosphorus.
[0260] The term "hydroxy" or "hydroxyl" includes groups with an --OH or --O.sup.- (with an appropriate counterion).
[0261] The term "halogen" includes fluorine, bromine, chlorine, iodine, etc. The term "perhalogenated" generally refers to a moiety wherein all hydrogens are replaced by halogen atoms.
[0262] The term "substituted" includes independently selected substituents which can be placed on the moiety and which allow the molecule to perform its intended function. Examples of substituents include alkyl, alkenyl, alkynyl, aryl, (CR'R'').sub.0-3 NR'R'', (CR'R'').sub.0-3 CN, NO.sub.2, halogen, (CR'R'').sub.0-3 C(halogen).sub.3, (CR'R'').sub.0-3 CH(halogen).sub.2, (CR'R'').sub.0-3CH.sub.2(halogen), (CR'R'').sub.0-3 CONR'R'', (CR'R'').sub.0-3 S(O).sub.1-2 NR'R'', (CR'R'').sub.0-3 CHO, (CR'R'').sub.0-3O(CR'R'').sub.0-3 H, (CR'R'').sub.0-3 S(O).sub.0-2R', (CR'R'').sub.0-3O(CR'R'').sub.0-3 H, (CR'R'').sub.0-3 COR', (CR'R'').sub.0-3 CO.sub.2R', or (CR'R'').sub.0-3OR' groups; wherein each R' and R'' are each independently hydrogen, a C.sub.1-C.sub.5 alkyl, C.sub.2-C.sub.5 alkenyl, C.sub.2-C.sub.5 alkynyl, or aryl group, or R' and R'' taken together are a benzylidene group or a --(CH.sub.2).sub.2O(CH.sub.2).sub.2-- group.
[0263] The term "amine" or "amino" includes compounds or moieties in which a nitrogen atom is covalently bonded to at least one carbon or heteroatom. The term "alkyl amino" includes groups and compounds wherein the nitrogen is bound to at least one additional alkyl group. The term "dialkyl amino" includes groups wherein the nitrogen atom is bound to at least two additional alkyl groups.
[0264] The term "ether" includes compounds or moieties which contain an oxygen bonded to two different carbon atoms or heteroatoms. For example, the term includes "alkoxyalkyl," which refers to an alkyl, alkenyl, or alkynyl group covalently bonded to an oxygen atom which is covalently bonded to another alkyl group.
[0265] The term "base" includes the known purine and pyrimidine heterocyclic bases, deazapurines, and analogs (including heterocyclic substituted analogs, e.g., aminoethyoxy phenoxazine), derivatives (e.g., 1-alkyl-, 1-alkenyl-, heteroaromatic- and 1-alkynyl derivatives) and tautomers thereof. Examples of purines include adenine, guanine, inosine, diaminopurine, and xanthine and analogs (e.g., 8-oxo-N6-methyladenine or 7-diazaxanthine) and derivatives thereof. Pyrimidines include, for example, thymine, uracil, and cytosine, and their analogs (e.g., 5-methylcytosine, 5-methyluracil, 5-(1-propynyl)uracil, 5-(1-propynyl)cytosine and 4,4-ethanocytosine). Other examples of suitable bases include non-purinyl and non-pyrimidinyl bases such as 2-aminopyridine and triazines.
[0266] In a preferred embodiment, the nucleomonomers of an oligonucleotide of the invention are RNA nucleotides. In another preferred embodiment, the nucleomonomers of an oligonucleotide of the invention are modified RNA nucleotides. Thus, the oligunucleotides contain modified RNA nucleotides.
[0267] The term "nucleoside" includes bases which are covalently attached to a sugar moiety, preferably ribose or deoxyribose. Examples of preferred nucleosides include ribonucleosides and deoxyribonucleosides. Nucleosides also include bases linked to amino acids or amino acid analogs which may comprise free carboxyl groups, free amino groups, or protecting groups. Suitable protecting groups are well known in the art (see P. G. M. Wuts and T. W. Greene, "Protective Groups in Organic Synthesis", 2.sup.nd Ed., Wiley-Interscience, New York, 1999).
[0268] The term "nucleotide" includes nucleosides which further comprise a phosphate group or a phosphate analog.
[0269] As used herein, the term "linkage" includes a naturally occurring, unmodified phosphodiester moiety (--O--(PO.sup.2)--O--) that covalently couples adjacent nucleomonomers. As used herein, the term "substitute linkage" includes any analog or derivative of the native phosphodiester group that covalently couples adjacent nucleomonomers. Substitute linkages include phosphodiester analogs, e.g., phosphorothioate, phosphorodithioate, and P-ethyoxyphosphodiester, P-ethoxyphosphodiester, P-alkyloxyphosphotriester, methylphosphonate, and nonphosphorus containing linkages, e.g., acetals and amides. Such substitute linkages are known in the art (e.g., Bjergarde et al. 1991. Nucleic Acids Res. 19:5843; Caruthers et al. 1991. Nucleosides Nucleotides. 10:47). In certain embodiments, non-hydrolizable linkages are preferred, such as phosphorothiate linkages.
[0270] In certain embodiments, oligonucleotides of the invention comprise hydrophobicly modified nucleotides or "hydrophobic modifications." As used herein "hydrophobic modifications" refers to bases that are modified such that (1) overall hydrophobicity of the base is significantly increased, and/or (2) the base is still capable of forming close to regular Watson-Crick interaction. Several non-limiting examples of base modifications include 5-position uridine and cytidine modifications such as phenyl, 4-pyridyl, 2-pyridyl, indolyl, and isobutyl, phenyl (C6H5OH); tryptophanyl (C8H6N)CH2CH(NH2)CO), Isobutyl, butyl, aminobenzyl; phenyl; and naphthyl.
[0271] In certain embodiments, oligonucleotides of the invention comprise 3' and 5' termini (except for circular oligonucleotides). In one embodiment, the 3' and 5' termini of an oligonucleotide can be substantially protected from nucleases e.g., by modifying the 3' or 5' linkages (e.g., U.S. Pat. No. 5,849,902 and WO 98/13526). For example, oligonucleotides can be made resistant by the inclusion of a "blocking group." The term "blocking group" as used herein refers to substituents (e.g., other than OH groups) that can be attached to oligonucleotides or nucleomonomers, either as protecting groups or coupling groups for synthesis (e.g., FITC, propyl (CH.sub.2--CH.sub.2--CH.sub.3), glycol (--O--CH.sub.2--CH.sub.2--O--) phosphate (PO.sub.3.sup.2-), hydrogen phosphonate, or phosphoramidite). "Blocking groups" also include "end blocking groups" or "exonuclease blocking groups" which protect the 5' and 3' termini of the oligonucleotide, including modified nucleotides and non-nucleotide exonuclease resistant structures.
[0272] Exemplary end-blocking groups include cap structures (e.g., a 7-methylguanosine cap), inverted nucleomonomers, e.g., with 3'-3' or 5'-5' end inversions (see, e.g., Ortiagao et al. 1992. Antisense Res. Dev. 2:129), methylphosphonate, phosphoramidite, non-nucleotide groups (e.g., non-nucleotide linkers, amino linkers, conjugates) and the like. The 3' terminal nucleomonomer can comprise a modified sugar moiety. The 3' terminal nucleomonomer comprises a 3'-0 that can optionally be substituted by a blocking group that prevents 3'-exonuclease degradation of the oligonucleotide. For example, the 3'-hydroxyl can be esterified to a nucleotide through a 3'.fwdarw.3' internucleotide linkage. For example, the alkyloxy radical can be methoxy, ethoxy, or isopropoxy, and preferably, ethoxy. Optionally, the 3'.fwdarw.3'linked nucleotide at the 3' terminus can be linked by a substitute linkage. To reduce nuclease degradation, the 5' most 3'.fwdarw.5' linkage can be a modified linkage, e.g., a phosphorothioate or a P-alkyloxyphosphotriester linkage. Preferably, the two 5' most 3'.fwdarw.5' linkages are modified linkages. Optionally, the 5' terminal hydroxy moiety can be esterified with a phosphorus containing moiety, e.g., phosphate, phosphorothioate, or P-ethoxyphosphate.
[0273] Another type of conjugates that can be attached to the end (3' or 5' end), the loop region, or any other parts of the miniRNA might include a sterol, sterol type molecule, peptide, small molecule, protein, etc. In some embodiments, a miniRNA may contain more than one conjugates (same or different chemical nature). In some embodiments, the conjugate is cholesterol.
[0274] Another way to increase target gene specificity, or to reduce off-target silencing effect, is to introduce a 2'-modification (such as the 2'-O methyl modification) at a position corresponding to the second 5'-end nucleotide of the guide sequence. This allows the positioning of this 2'-modification in the Dicer-resistant hairpin structure, thus enabling one to design better RNAi constructs with less or no off-target silencing.
[0275] In one embodiment, a hairpin polynucleotide of the invention can comprise one nucleic acid portion which is DNA and one nucleic acid portion which is RNA. Antisense (guide) sequences of the invention can be "chimeric oligonucleotides" which comprise an RNA-like and a DNA-like region.
[0276] The language "RNase H activating region" includes a region of an oligonucleotide, e.g., a chimeric oligonucleotide, that is capable of recruiting RNase H to cleave the target RNA strand to which the oligonucleotide binds. Typically, the RNase activating region contains a minimal core (of at least about 3-5, typically between about 3-12, more typically, between about 5-12, and more preferably between about 5-10 contiguous nucleomonomers) of DNA or DNA-like nucleomonomers. (See, e.g., U.S. Pat. No. 5,849,902). Preferably, the RNase H activating region comprises about nine contiguous deoxyribose containing nucleomonomers.
[0277] The language "non-activating region" includes a region of an antisense sequence, e.g., a chimeric oligonucleotide, that does not recruit or activate RNase H. Preferably, a non-activating region does not comprise phosphorothioate DNA. The oligonucleotides of the invention comprise at least one non-activating region. In one embodiment, the non-activating region can be stabilized against nucleases or can provide specificity for the target by being complementary to the target and forming hydrogen bonds with the target nucleic acid molecule, which is to be bound by the oligonucleotide.
[0278] In one embodiment, at least a portion of the contiguous polynucleotides are linked by a substitute linkage, e.g., a phosphorothioate linkage.
[0279] In certain embodiments, most or all of the nucleotides beyond the guide sequence (2'-modified or not) are linked by phosphorothioate linkages. Such constructs tend to have improved pharmacokinetics due to their higher affinity for serum proteins. The phosphorothioate linkages in the non-guide sequence portion of the polynucleotide generally do not interfere with guide strand activity, once the latter is loaded into RISC.
[0280] Antisense (guide) sequences of the present invention may include "morpholino oligonucleotides." Morpholino oligonucleotides are non-ionic and function by an RNase H-independent mechanism. Each of the 4 genetic bases (Adenine, Cytosine, Guanine, and Thymine/Uracil) of the morpholino oligonucleotides is linked to a 6-membered morpholine ring. Morpholino oligonucleotides are made by joining the 4 different subunit types by, e.g., non-ionic phosphorodiamidate inter-subunit linkages. Morpholino oligonucleotides have many advantages including: complete resistance to nucleases (Antisense & Nucl. Acid Drug Dev. 1996. 6:267); predictable targeting (Biochemica Biophysica Acta. 1999. 1489:141); reliable activity in cells (Antisense & Nucl. Acid Drug Dev. 1997. 7:63); excellent sequence specificity (Antisense & Nucl. Acid Drug Dev. 1997. 7:151); minimal non-antisense activity (Biochemica Biophysica Acta. 1999. 1489:141); and simple osmotic or scrape delivery (Antisense & Nucl. Acid Drug Dev. 1997. 7:291). Morpholino oligonucleotides are also preferred because of their non-toxicity at high doses. A discussion of the preparation of morpholino oligonucleotides can be found in Antisense & Nucl. Acid Drug Dev. 1997. 7:187.
[0281] The chemical modifications described herein are believed, based on the data described herein, to promote single stranded polynucleotide loading into the RISC. Single stranded polynucleotides have been shown to be active in loading into RISC and inducing gene silencing. However, the level of activity for single stranded polynucleotides appears to be 2 to 4 orders of magnitude lower when compared to a duplex polynucleotide.
[0282] The present invention provides a description of the chemical modification patterns, which may (a) significantly increase stability of the single stranded polynucleotide (b) promote efficient loading of the polynucleotide into the RISC complex and (c) improve uptake of the single stranded nucleotide by the cell. FIG. 5 provides some non-limiting examples of the chemical modification patterns which may be beneficial for achieving single stranded polynucleotide efficacy inside the cell. The chemical modification patterns may include combination of ribose, backbone, hydrophobic nucleoside and conjugate type of modifications. In addition, in some of the embodiments, the 5' end of the single polynucleotide may be chemically phosphorylated.
[0283] In yet another embodiment, the present invention provides a description of the chemical modifications patterns, which improve functionality of RISC inhibiting polynucleotides. Single stranded polynucleotides have been shown to inhibit activity of a preloaded RISC complex through the substrate competition mechanism. For these types of molecules, conventionally called antagomers, the activity usually requires high concentration and in vivo delivery is not very effective. The present invention provides a description of the chemical modification patterns, which may (a) significantly increase stability of the single stranded polynucleotide (b) promote efficient recognition of the polynucleotide by the RISC as a substrate and/or (c) improve uptake of the single stranded nucleotide by the cell. FIG. 6 provides some non-limiting examples of the chemical modification patterns that may be beneficial for achieving single stranded polynucleotide efficacy inside the cell. The chemical modification patterns may include combination of ribose, backbone, hydrophobic nucleoside and conjugate type of modifications.
[0284] The modifications provided by the present invention are applicable to all polynucleotides. This includes single stranded RISC entering polynucleotides, single stranded RISC inhibiting polynucleotides, conventional duplexed polynucleotides of variable length (15-40 bp), asymmetric duplexed polynucleotides, and the like. Polynucleotides may be modified with wide variety of chemical modification patterns, including 5' end, ribose, backbone and hydrophobic nucleoside modifications.
Synthesis
[0285] Oligonucleotides of the invention can be synthesized by any method known in the art, e.g., using enzymatic synthesis and/or chemical synthesis. The oligonucleotides can be synthesized in vitro (e.g., using enzymatic synthesis and chemical synthesis) or in vivo (using recombinant DNA technology well known in the art).
[0286] In a preferred embodiment, chemical synthesis is used for modified polynucleotides. Chemical synthesis of linear oligonucleotides is well known in the art and can be achieved by solution or solid phase techniques. Preferably, synthesis is by solid phase methods. Oligonucleotides can be made by any of several different synthetic procedures including the phosphoramidite, phosphite triester, H-phosphonate, and phosphotriester methods, typically by automated synthesis methods.
[0287] Oligonucleotide synthesis protocols are well known in the art and can be found, e.g., in U.S. Pat. No. 5,830,653; WO 98/13526; Stec et al. 1984. J. Am. Chem. Soc. 106:6077; Stec et al. 1985. J. Org. Chem. 50:3908; Stec et al. J. Chromatog. 1985. 326:263; LaPlanche et al. 1986. Nucl. Acid. Res. 1986. 14:9081; Fasman G. D., 1989. Practical Handbook of Biochemistry and Molecular Biology. 1989. CRC Press, Boca Raton, Fla.; Lamone. 1993. Biochem. Soc. Trans. 21:1; U.S. Pat. Nos. 5,013,830; 5,214,135; 5,525,719; Kawasaki et al. 1993. J. Med. Chem. 36:831; WO 92/03568; U.S. Pat. Nos. 5,276,019; and 5,264,423.
[0288] The synthesis method selected can depend on the length of the desired oligonucleotide and such choice is within the skill of the ordinary artisan. For example, the phosphoramidite and phosphite triester method can produce oligonucleotides having 175 or more nucleotides, while the H-phosphonate method works well for oligonucleotides of less than 100 nucleotides. If modified bases are incorporated into the oligonucleotide, and particularly if modified phosphodiester linkages are used, then the synthetic procedures are altered as needed according to known procedures. In this regard, Uhlmann et al. (1990, Chemical Reviews 90:543-584) provide references and outline procedures for making oligonucleotides with modified bases and modified phosphodiester linkages. Other exemplary methods for making oligonucleotides are taught in Sonveaux. 1994. "Protecting Groups in Oligonucleotide Synthesis"; Agrawal. Methods in Molecular Biology 26:1. Exemplary synthesis methods are also taught in "Oligonucleotide Synthesis--A Practical Approach" (Gait, M. J. IRL Press at Oxford University Press. 1984). Moreover, linear oligonucleotides of defined sequence, including some sequences with modified nucleotides, are readily available from several commercial sources.
[0289] The oligonucleotides may be purified by polyacrylamide gel electrophoresis, or by any of a number of chromatographic methods, including gel chromatography and high pressure liquid chromatography. To confirm a nucleotide sequence, especially unmodified nucleotide sequences, oligonucleotides may be subjected to DNA sequencing by any of the known procedures, including Maxam and Gilbert sequencing, Sanger sequencing, capillary electrophoresis sequencing, the wandering spot sequencing procedure or by using selective chemical degradation of oligonucleotides bound to Hybond paper. Sequences of short oligonucleotides can also be analyzed by laser desorption mass spectroscopy or by fast atom bombardment (McNeal, et al., 1982, J. Am. Chem. Soc. 104:976; Viari, et al., 1987, Biomed. Environ. Mass Spectrom. 14:83; Grotjahn et al., 1982, Nuc. Acid Res. 10:4671). Sequencing methods are also available for RNA oligonucleotides.
[0290] The quality of oligonucleotides synthesized can be verified by testing the oligonucleotide by capillary electrophoresis and denaturing strong anion HPLC (SAX-HPLC) using, e.g., the method of Bergot and Egan. 1992. J. Chrom. 599:35.
[0291] Other exemplary synthesis techniques are well known in the art (see, e.g., Sambrook et al., Molecular Cloning: a Laboratory Manual, Second Edition (1989); DNA Cloning, Volumes I and II (D N Glover Ed. 1985); Oligonucleotide Synthesis (M J Gait Ed, 1984; Nucleic Acid Hybridisation (B D Hames and S J Higgins eds. 1984); A Practical Guide to Molecular Cloning (1984); or the series, Methods in Enzymology (Academic Press, Inc.)).
[0292] In certain embodiments, the subject RNAi constructs or at least portions thereof are transcribed from expression vectors encoding the subject constructs. Any art recognized vectors may be use for this purpose. The transcribed RNAi constructs may be isolated and purified, before desired modifications (such as replacing an unmodified sense strand with a modified one, etc.) are carried out.
Delivery/Carrier
Uptake of Oligonucleotides by Cells
[0293] Oligonucleotides and oligonucleotide compositions are contacted with (i.e., brought into contact with, also referred to herein as administered or delivered to) and taken up by one or more cells or a cell lysate. The term "cells" includes prokaryotic and eukaryotic cells, preferably vertebrate cells, and, more preferably, mammalian cells. In a preferred embodiment, the oligonucleotide compositions of the invention are contacted with human cells.
[0294] Oligonucleotide compositions of the invention can be contacted with cells in vitro, e.g., in a test tube or culture dish, (and may or may not be introduced into a subject) or in vivo, e.g., in a subject such as a mammalian subject. Oligonucleotides are taken up by cells at a slow rate by endocytosis, but endocytosed oligonucleotides are generally sequestered and not available, e.g., for hybridization to a target nucleic acid molecule. In one embodiment, cellular uptake can be facilitated by electroporation or calcium phosphate precipitation. However, these procedures are only useful for in vitro or ex vivo embodiments, are not convenient and, in some cases, are associated with cell toxicity.
[0295] In another embodiment, delivery of oligonucleotides into cells can be enhanced by suitable art recognized methods including calcium phosphate, DMSO, glycerol or dextran, electroporation, or by transfection, e.g., using cationic, anionic, or neutral lipid compositions or liposomes using methods known in the art (see e.g., WO 90/14074; WO 91/16024; WO 91/17424; U.S. Pat. No. 4,897,355; Bergan et al. 1993. Nucleic Acids Research. 21:3567). Enhanced delivery of oligonucleotides can also be mediated by the use of vectors (See e.g., Shi, Y. 2003. Trends Genet 2003 Jan. 19:9; Reichhart J M et al. Genesis. 2002. 34(1-2):1604, Yu et al. 2002. Proc. Natl. Acad Sci. USA 99:6047; Sui et al. 2002. Proc. Natl. Acad Sci. USA 99:5515) viruses, polyamine or polycation conjugates using compounds such as polylysine, protamine, or Ni, N12-bis (ethyl) spermine (see, e.g., Bartzatt, R. et al. 1989. Biotechnol. Appl. Biochem. 11:133; Wagner E. et al. 1992. Proc. Natl. Acad. Sci. 88:4255).
[0296] In certain embodiments, the miniRNA of the invention may be delivered by using various beta-glucan containing particles, such as those described in US 2005/0281781 A1, WO 2006/007372, and WO 2007/050643 (all incorporated herein by reference). In certain embodiments, the beta-glucan particle is derived from yeast. In certain embodiments, the payload trapping molecule is a polymer, such as those with a molecular weight of at least about 1000 Da, 10,000 Da, 50,000 Da, 100 kDa, 500 kDa, etc. Preferred polymers include (without limitation) cationic polymers, chitosans, or PEI (polyethylenimine), etc.
[0297] Such beta-glucan based delivery system may be formulated for oral delivery, where the orally delivered beta-glucan/miniRNA constructs may be engulfed by macrophages or other related phagocytic cells, which may in turn release the miniRNA constructs in selected in vivo sites. Alternatively or in addition, the miniRNA may changes the expression of certain macrophage target genes.
[0298] The optimal protocol for uptake of oligonucleotides will depend upon a number of factors, the most crucial being the type of cells that are being used. Other factors that are important in uptake include, but are not limited to, the nature and concentration of the oligonucleotide, the confluence of the cells, the type of culture the cells are in (e.g., a suspension culture or plated) and the type of media in which the cells are grown.
Encapsulating Agents
[0299] Encapsulating agents entrap oligonucleotides within vesicles. In another embodiment of the invention, an oligonucleotide may be associated with a carrier or vehicle, e.g., liposomes or micelles, although other carriers could be used, as would be appreciated by one skilled in the art. Liposomes are vesicles made of a lipid bilayer having a structure similar to biological membranes. Such carriers are used to facilitate the cellular uptake or targeting of the oligonucleotide, or improve the oligonucleotide's pharmacokinetic or toxicologic properties.
[0300] For example, the oligonucleotides of the present invention may also be administered encapsulated in liposomes, pharmaceutical compositions wherein the active ingredient is contained either dispersed or variously present in corpuscles consisting of aqueous concentric layers adherent to lipidic layers. The oligonucleotides, depending upon solubility, may be present both in the aqueous layer and in the lipidic layer, or in what is generally termed a liposomic suspension. The hydrophobic layer, generally but not exclusively, comprises phopholipids such as lecithin and sphingomyelin, steroids such as cholesterol, more or less ionic surfactants such as diacetylphosphate, stearylamine, or phosphatidic acid, or other materials of a hydrophobic nature. The diameters of the liposomes generally range from about 15 nm to about 5 microns.
[0301] The use of liposomes as drug delivery vehicles offers several advantages. Liposomes increase intracellular stability, increase uptake efficiency and improve biological activity. Liposomes are hollow spherical vesicles composed of lipids arranged in a similar fashion as those lipids which make up the cell membrane. They have an internal aqueous space for entrapping water soluble compounds and range in size from 0.05 to several microns in diameter. Several studies have shown that liposomes can deliver nucleic acids to cells and that the nucleic acids remain biologically active. For example, a lipid delivery vehicle originally designed as a research tool, such as Lipofectin or LIPOFECTAMINE.TM. 2000, can deliver intact nucleic acid molecules to cells.
[0302] Specific advantages of using liposomes include the following: they are non-toxic and biodegradable in composition; they display long circulation half-lives; and recognition molecules can be readily attached to their surface for targeting to tissues. Finally, cost-effective manufacture of liposome-based pharmaceuticals, either in a liquid suspension or lyophilized product, has demonstrated the viability of this technology as an acceptable drug delivery system.
[0303] In some aspects, formulations associated with the invention might be selected for a class of naturally occurring or chemically synthesized or modified saturated and unsaturated fatty acid residues. Fatty acids might exist in a form of triglycerides, diglycerides or individual fatty acids. In another embodiment, the use of well-validated mixtures of fatty acids and/or fat emulsions currently used in pharmacology for parenteral nutrition may be utilized.
[0304] Liposome based formulations are widely used for oligonucleotide delivery. However, most of commercially available lipid or liposome formulations contain at least one positively charged lipid (cationic lipids). The presence of this positively charged lipid is believed to be essential for obtaining a high degree of oligonucleotide loading and for enhancing liposome fusogenic properties. Several methods have been performed and published to identify optimal positively charged lipid chemistries. However, the commercially available liposome formulations containing cationic lipids are characterized by a high level of toxicity. In vivo limited therapeutic indexes have revealed that liposome formulations containing positive charged lipids are associated with toxicity (i.e. elevation in liver enzymes) at concentrations only slightly higher than concentration required to achieve RNA silencing.
[0305] New liposome formulations, lacking the toxicity of the prior art liposomes have been developed according to the invention. These new liposome formulations are neutral fat-based formulations for the efficient delivery of oligonucleotides, and in particular for the delivery of the RNA molecules of the invention. The compositions are referred to as neutral nanotransporters because they enable quantitative oligonucleotide incorporation into non-charged lipids mixtures. The lack of toxic levels of cationic lipids in the neutral nanotransporter compositions of the invention is an important feature.
[0306] The neutral nanotransporters compositions enable efficient loading of oligonucleotide into neutral fat formulation. The composition includes an oligonucleotide that is modified in a manner such that the hydrophobicity of the molecule is increased (for example a hydrophobic molecule is attached (covalently or no-covalently) to a hydrophobic molecule on the oligonucleotide terminus or a non-terminal nucleotide, base, sugar, or backbone), the modified oligonucleotide being mixed with a neutral fat formulation (for example containing at least 25% of cholesterol and 25% of DOPC or analogs thereof). A cargo molecule, such as another lipid can also be included in the composition. This composition, where part of the formulation is build into the oligonucleotide itself, enables efficient encapsulation of oligonucleotide in neutral lipid particles.
[0307] One of several unexpected observations associated with the invention was that the oligonucleotides of the invention could effectively be incorporated in a lipid mixture that was free of cationic lipids and that such a composition could effectively deliver the therapeutic oligonucleotide to a cell in a manner that it is functional. Another unexpected observation was the high level of activity observed when the fatty mixture is composed of a phosphatidylcholine base fatty acid and a sterol such as a cholesterol. For instance, one preferred formulation of neutral fatty mixture is composed of at least 20% of DOPC or DSPC and at least 20% of sterol such as cholesterol. Even as low as 1:5 lipid to oligonucleotide ratio was shown to be sufficient to get complete encapsulation of the oligonucleotide in a non charged formulation. The prior art demonstrated only a 1-5% oligonucleotide encapsulation with non-charged formulations, which is not sufficient to get to a desired amount of in vivo efficacy. Compared to the prior art using neutral lipids the level of oligonucleotide delivery to a cell was quite unexpected.
[0308] Stable particles ranging in size from 50 to 140 nm were formed upon complexing of hydrophobic oligonucleotides with preferred formulations. It is interesting to mention that the formulation by itself typically does not form small particles, but rather, forms agglomerates, which are transformed into stable 50-120 nm particles upon addition of the hydrophobic modified oligonucleotide.
[0309] The neutral nanotransporter compositions of the invention include a hydrophobic modified polynucleotide, a neutral fatty mixture, and optionally a cargo molecule. A "hydrophobic modified polynucleotide" as used herein is a polynucleotide of the invention (i.e. sd-rxRNA) that has at least one modification that renders the polynucleotide more hydrophobic than the polynucleotide was prior to modification. The modification may be achieved by attaching (covalently or non-covalently) a hydrophobic molecule to the polynucleotide. In some instances the hydrophobic molecule is or includes a lipophilic group.
[0310] The term "lipophilic group" means a group that has a higher affinity for lipids than its affinity for water. Examples of lipophilic groups include, but are not limited to, cholesterol, a cholesteryl or modified cholesteryl residue, adamantine, dihydrotesterone, long chain alkyl, long chain alkenyl, long chain alkynyl, olely-lithocholic, cholenic, oleoyl-cholenic, palmityl, heptadecyl, myrisityl, bile acids, cholic acid or taurocholic acid, deoxycholate, oleyl litocholic acid, oleoyl cholenic acid, glycolipids, phospholipids, sphingolipids, isoprenoids, such as steroids, vitamins, such as vitamin E, fatty acids either saturated or unsaturated, fatty acid esters, such as triglycerides, pyrenes, porphyrines, Texaphyrine, adamantane, acridines, biotin, coumarin, fluorescein, rhodamine, Texas-Red, digoxygenin, dimethoxytrityl, t-butyldimethylsilyl, t-butyldiphenylsilyl, cyanine dyes (e.g. Cy3 or Cy5), Hoechst 33258 dye, psoralen, or ibuprofen. The cholesterol moiety may be reduced (e.g. as in cholestan) or may be substituted (e.g. by halogen). A combination of different lipophilic groups in one molecule is also possible.
[0311] The hydrophobic molecule may be attached at various positions of the polynucleotide. As described above, the hydrophobic molecule may be linked to the terminal residue of the polynucleotide such as the 3' of 5'-end of the polynucleotide. Alternatively, it may be linked to an internal nucleotide or a nucleotide on a branch of the polynucleotide. The hydrophobic molecule may be attached, for instance to a 2'-position of the nucleotide. The hydrophobic molecule may also be linked to the heterocyclic base, the sugar or the backbone of a nucleotide of the polynucleotide.
[0312] The hydrophobic molecule may be connected to the polynucleotide by a linker moiety. Optionally the linker moiety is a non-nucleotidic linker moiety. Non-nucleotidic linkers are e.g. abasic residues (dSpacer), oligoethyleneglycol, such as triethyleneglycol (spacer 9) or hexaethylenegylcol (spacer 18), or alkane-diol, such as butanediol. The spacer units are preferably linked by phosphodiester or phosphorothioate bonds. The linker units may appear just once in the molecule or may be incorporated several times, e.g. via phosphodiester, phosphorothioate, methylphosphonate, or amide linkages.
[0313] Typical conjugation protocols involve the synthesis of polynucleotides bearing an aminolinker at one or more positions of the sequence, however, a linker is not required. The amino group is then reacted with the molecule being conjugated using appropriate coupling or activating reagents. The conjugation reaction may be performed either with the polynucleotide still bound to a solid support or following cleavage of the polynucleotide in solution phase. Purification of the modified polynucleotide by HPLC typically results in a pure material.
[0314] In some embodiments the hydrophobic molecule is a sterol type conjugate, a PhytoSterol conjugate, cholesterol conjugate, sterol type conjugate with altered side chain length, fatty acid conjugate, any other hydrophobic group conjugate, and/or hydrophobic modifications of the internal nucleoside, which provide sufficient hydrophobicity to be incorporated into micelles.
[0315] For purposes of the present invention, the term "sterols", refers or steroid alcohols are a subgroup of steroids with a hydroxyl group at the 3-position of the A-ring. They are amphipathic lipids synthesized from acetyl-coenzyme A via the HMG-CoA reductase pathway. The overall molecule is quite flat. The hydroxyl group on the A ring is polar. The rest of the aliphatic chain is non-polar. Usually sterols are considered to have an 8 carbon chain at position 17.
[0316] For purposes of the present invention, the term "sterol type molecules", refers to steroid alcohols, which are similar in structure to sterols. The main difference is the structure of the ring and number of carbons in a position 21 attached side chain.
[0317] For purposes of the present invention, the term "PhytoSterols" (also called plant sterols) are a group of steroid alcohols, phytochemicals naturally occurring in plants. There are more then 200 different known PhytoSterols
[0318] For purposes of the present invention, the term "Sterol side chain" refers to a chemical composition of a side chain attached at the position 17 of sterol-type molecule. In a standard definition sterols are limited to a 4 ring structure carrying a 8 carbon chain at position 17. In this invention, the sterol type molecules with side chain longer and shorter than conventional are described. The side chain may branched or contain double back bones.
[0319] Thus, sterols useful in the invention, for example, include cholesterols, as well as unique sterols in which position 17 has attached side chain of 2-7 or longer then 9 carbons. In a particular embodiment, the length of the polycarbon tail is varied between 5 and 9 carbons. FIG. 9 demonstrates that there is a correlation between plasma clearance, liver uptake and the length of the polycarbon chain. Such conjugates may have significantly better in vivo efficacy, in particular delivery to liver. These types of molecules are expected to work at concentrations 5 to 9 fold lower then oligonucleotides conjugated to conventional cholesterols.
[0320] Alternatively the polynucleotide may be bound to a protein, peptide or positively charged chemical that functions as the hydrophobic molecule. The proteins may be selected from the group consisting of protamine, dsRNA binding domain, and arginine rich peptides. Exemplary positively charged chemicals include spermine, spermidine, cadaverine, and putrescine.
[0321] In another embodiment hydrophobic molecule conjugates may demonstrate even higher efficacy when it is combined with optimal chemical modification patterns of the polynucleotide (as described herein in detail), containing but not limited to hydrophobic modifications, phosphorothioate modifications, and 2' ribo modifications.
[0322] In another embodiment the sterol type molecule may be a naturally occurring PhytoSterols such as those shown in FIG. 8. The polycarbon chain may be longer than 9 and may be linear, branched and/or contain double bonds. Some PhytoSterol containing polynucleotide conjugates may be significantly more potent and active in delivery of polynucleotides to various tissues. Some PhytoSterols may demonstrate tissue preference and thus be used as a way to delivery RNAi specifically to particular tissues.
[0323] The hydrophobic modified polynucleotide is mixed with a neutral fatty mixture to form a micelle. The neutral fatty acid mixture is a mixture of fats that has a net neutral or slightly net negative charge at or around physiological pH that can form a micelle with the hydrophobic modified polynucleotide. For purposes of the present invention, the term "micelle" refers to a small nanoparticle formed by a mixture of non charged fatty acids and phospholipids. The neutral fatty mixture may include cationic lipids as long as they are present in an amount that does not cause toxicity. In preferred embodiments the neutral fatty mixture is free of cationic lipids. A mixture that is free of cationic lipids is one that has less than 1% and preferably 0% of the total lipid being cationic lipid. The term "cationic lipid" includes lipids and synthetic lipids having a net positive charge at or around physiological pH. The term "anionic lipid" includes lipids and synthetic lipids having a net negative charge at or around physiological pH.
[0324] The neutral fats bind to the oligonucleotides of the invention by a strong but non-covalent attraction (e.g., an electrostatic, van der Waals, pi-stacking, etc. interaction).
[0325] The neutral fat mixture may include formulations selected from a class of naturally occurring or chemically synthesized or modified saturated and unsaturated fatty acid residues. Fatty acids might exist in a form of triglycerides, diglycerides or individual fatty acids. In another embodiment the use of well-validated mixtures of fatty acids and/or fat emulsions currently used in pharmacology for parenteral nutrition may be utilized.
[0326] The neutral fatty mixture is preferably a mixture of a choline based fatty acid and a sterol. Choline based fatty acids include for instance, synthetic phosphocholine derivatives such as DDPC, DLPC, DMPC, DPPC, DSPC, DOPC, POPC, and DEPC. DOPC (chemical registry number 4235-95-4) is dioleoylphosphatidylcholine (also known as dielaidoylphosphatidylcholine, dioleoyl-PC, dioleoylphosphocholine, dioleoyl-sn-glycero-3-phosphocholine, dioleylphosphatidylcholine). DSPC (chemical registry number 816-94-4) is distearoylphosphatidylcholine (also known as 1,2-Distearoyl-sn-Glycero-3-phosphocholine).
[0327] The sterol in the neutral fatty mixture may be for instance cholesterol. The neutral fatty mixture may be made up completely of a choline based fatty acid and a sterol or it may optionally include a cargo molecule. For instance, the neutral fatty mixture may have at least 20% or 25% fatty acid and 20% or 25% sterol.
[0328] For purposes of the present invention, the term "Fatty acids" relates to conventional description of fatty acid. They may exist as individual entities or in a form of two- and triglycerides. For purposes of the present invention, the term "fat emulsions" refers to safe fat formulations given intravenously to subjects who are unable to get enough fat in their diet. It is an emulsion of soy bean oil (or other naturally occurring oils) and egg phospholipids. Fat emulsions are being used for formulation of some insoluble anesthetics. In this disclosure, fat emulsions might be part of commercially available preparations like Intralipid, Liposyn, Nutrilipid, modified commercial preparations, where they are enriched with particular fatty acids or fully de novo-formulated combinations of fatty acids and phospholipids.
[0329] In one embodiment, the cells to be contacted with an oligonucleotide composition of the invention are contacted with a mixture comprising the oligonucleotide and a mixture comprising a lipid, e.g., one of the lipids or lipid compositions described supra for between about 12 hours to about 24 hours. In another embodiment, the cells to be contacted with an oligonucleotide composition are contacted with a mixture comprising the oligonucleotide and a mixture comprising a lipid, e.g., one of the lipids or lipid compositions described supra for between about 1 and about five days. In one embodiment, the cells are contacted with a mixture comprising a lipid and the oligonucleotide for between about three days to as long as about 30 days. In another embodiment, a mixture comprising a lipid is left in contact with the cells for at least about five to about 20 days. In another embodiment, a mixture comprising a lipid is left in contact with the cells for at least about seven to about 15 days. 50%-60% of the formulation can optionally be any other lipid or molecule. Such a lipid or molecule is referred to herein as a cargo lipid or cargo molecule. Cargo molecules include but are not limited to intralipid, small molecules, fusogenic peptides or lipids or other small molecules might be added to alter cellular uptake, endosomal release or tissue distribution properties. The ability to tolerate cargo molecules is important for modulation of properties of these particles, if such properties are desirable. For instance the presence of some tissue specific metabolites might drastically alter tissue distribution profiles. For example use of Intralipid type formulation enriched in shorter or longer fatty chains with various degrees of saturation affects tissue distribution profiles of these type of formulations (and their loads).
[0330] An example of a cargo lipid useful according to the invention is a fusogenic lipid. For instance, the zwiterionic lipid DOPE (chemical registry number 4004-5-1, 1,2-Dioleoyl-sn-Glycero-3-phosphoethanolamine) is a preferred cargo lipid.
[0331] Intralipid may be comprised of the following composition: 1 000 mL contain: purified soybean oil 90 g, purified egg phospholipids 12 g, glycerol anhydrous 22 g, water for injection q.s. ad 1 000 mL. pH is adjusted with sodium hydroxide to pH approximately 8. Energy content/L: 4.6 MJ (190 kcal). Osmolality (approx.): 300 mOsm/kg water. In another embodiment fat emulsion is Liposyn that contains 5% safflower oil, 5% soybean oil, up to 1.2% egg phosphatides added as an emulsifier and 2.5% glycerin in water for injection. It may also contain sodium hydroxide for pH adjustment. pH 8.0 (6.0-9.0). Liposyn has an osmolarity of 276 m Osmol/liter (actual).
[0332] Variation in the identity, amounts and ratios of cargo lipids affects the cellular uptake and tissue distribution characteristics of these compounds. For example, the length of lipid tails and level of saturability will affect differential uptake to liver, lung, fat and cardiomyocytes. Addition of special hydrophobic molecules like vitamins or different forms of sterols can favor distribution to special tissues which are involved in the metabolism of particular compounds. Complexes are formed at different oligonucleotide concentrations, with higher concentrations favoring more efficient complex formation (FIGS. 21-22).
[0333] In another embodiment, the fat emulsion is based on a mixture of lipids. Such lipids may include natural compounds, chemically synthesized compounds, purified fatty acids or any other lipids. In yet another embodiment the composition of fat emulsion is entirely artificial. In a particular embodiment, the fat emulsion is more then 70% linoleic acid. In yet another particular embodiment the fat emulsion is at least 1% of cardiolipin. Linoleic acid (LA) is an unsaturated omega-6 fatty acid. It is a colorless liquid made of a carboxylic acid with an 18-carbon chain and two cis double bonds.
[0334] In yet another embodiment of the present invention, the alteration of the composition of the fat emulsion is used as a way to alter tissue distribution of hydrophobicly modified polynucleotides. This methodology provides for the specific delivery of the polynucleotides to particular tissues (FIG. 12).
[0335] In another embodiment the fat emulsions of the cargo molecule contain more then 70% of Linoleic acid (C18H32O2) and/or cardiolipin are used for specifically delivering RNAi to heart muscle.
[0336] Fat emulsions, like intralipid have been used before as a delivery formulation for some non-water soluble drugs (such as Propofol, re-formulated as Diprivan). Unique features of the present invention include (a) the concept of combining modified polynucleotides with the hydrophobic compound(s), so it can be incorporated in the fat micelles and (b) mixing it with the fat emulsions to provide a reversible carrier. After injection into a blood stream, micelles usually bind to serum proteins, including albumin, HDL, LDL and other. This binding is reversible and eventually the fat is absorbed by cells. The polynucleotide, incorporated as a part of the micelle will then be delivered closely to the surface of the cells. After that cellular uptake might be happening though variable mechanisms, including but not limited to sterol type delivery.
Complexing Agents
[0337] Complexing agents bind to the oligonucleotides of the invention by a strong but non-covalent attraction (e.g., an electrostatic, van der Waals, pi-stacking, etc. interaction). In one embodiment, oligonucleotides of the invention can be complexed with a complexing agent to increase cellular uptake of oligonucleotides. An example of a complexing agent includes cationic lipids. Cationic lipids can be used to deliver oligonucleotides to cells. However, as discussed above, formulations free in cationic lipids are preferred in some embodiments.
[0338] The term "cationic lipid" includes lipids and synthetic lipids having both polar and non-polar domains and which are capable of being positively charged at or around physiological pH and which bind to polyanions, such as nucleic acids, and facilitate the delivery of nucleic acids into cells. In general cationic lipids include saturated and unsaturated alkyl and alicyclic ethers and esters of amines, amides, or derivatives thereof. Straight-chain and branched alkyl and alkenyl groups of cationic lipids can contain, e.g., from 1 to about 25 carbon atoms. Preferred straight chain or branched alkyl or alkene groups have six or more carbon atoms. Alicyclic groups include cholesterol and other steroid groups. Cationic lipids can be prepared with a variety of counterions (anions) including, e.g., Cl.sup.-, Br.sup.-, I.sup.-, F.sup.-, acetate, trifluoroacetate, sulfate, nitrite, and nitrate.
[0339] Examples of cationic lipids include polyethylenimine, polyamidoamine (PAMAM) starburst dendrimers, Lipofectin (a combination of DOTMA and DOPE), Lipofectase, LIPOFECTAMINE.TM. (e.g., LIPOFECTAMINE.TM. 2000), DOPE, Cytofectin (Gilead Sciences, Foster City, Calif.), and Eufectins (JBL, San Luis Obispo, Calif.). Exemplary cationic liposomes can be made from N-[1-(2,3-dioleoloxy)-propyl]-N,N,N-trimethylammonium chloride (DOTMA), N-[1-(2,3-dioleoloxy)-propyl]-N,N,N-trimethylammonium methylsulfate (DOTAP), 3.beta.-[N--(N',N'-dimethylaminoethane)carbamoyl]cholesterol (DC-Chol), 2,3,-dioleyloxy-N-[2(sperminecarboxamido)ethyl]-N,N-dimethyl-1-propanamin- ium trifluoroacetate (DOSPA), 1,2-dimyristyloxypropyl-3-dimethyl-hydroxyethyl ammonium bromide; and dimethyldioctadecylammonium bromide (DDAB). The cationic lipid N-(1-(2,3-dioleyloxy)propyl)-N,N,N-trimethylammonium chloride (DOTMA), for example, was found to increase 1000-fold the antisense effect of a phosphorothioate oligonucleotide. (Vlassov et al., 1994, Biochimica et Biophysica Acta 1197:95-108). Oligonucleotides can also be complexed with, e.g., poly (L-lysine) or avidin and lipids may, or may not, be included in this mixture, e.g., steryl-poly (L-lysine).
[0340] Cationic lipids have been used in the art to deliver oligonucleotides to cells (see, e.g., U.S. Pat. Nos. 5,855,910; 5,851,548; 5,830,430; 5,780,053; 5,767,099; Lewis et al. 1996. Proc. Natl. Acad. Sci. USA 93:3176; Hope et al. 1998. Molecular Membrane Biology 15:1). Other lipid compositions which can be used to facilitate uptake of the instant oligonucleotides can be used in connection with the claimed methods. In addition to those listed supra, other lipid compositions are also known in the art and include, e.g., those taught in U.S. Pat. Nos. 4,235,871; 4,501,728; 4,837,028; 4,737,323.
[0341] In one embodiment lipid compositions can further comprise agents, e.g., viral proteins to enhance lipid-mediated transfections of oligonucleotides (Kamata, et al., 1994. Nucl. Acids. Res. 22:536). In another embodiment, oligonucleotides are contacted with cells as part of a composition comprising an oligonucleotide, a peptide, and a lipid as taught, e.g., in U.S. Pat. No. 5,736,392. Improved lipids have also been described which are serum resistant (Lewis, et al., 1996. Proc. Natl. Acad. Sci. 93:3176). Cationic lipids and other complexing agents act to increase the number of oligonucleotides carried into the cell through endocytosis.
[0342] In another embodiment N-substituted glycine oligonucleotides (peptoids) can be used to optimize uptake of oligonucleotides. Peptoids have been used to create cationic lipid-like compounds for transfection (Murphy, et al., 1998. Proc. Natl. Acad. Sci. 95:1517). Peptoids can be synthesized using standard methods (e.g., Zuckermann, R. N., et al. 1992. J. Am. Chem. Soc. 114:10646; Zuckermann, R. N., et al. 1992. Int. J. Peptide Protein Res. 40:497). Combinations of cationic lipids and peptoids, liptoids, can also be used to optimize uptake of the subject oligonucleotides (Hunag, et al., 1998. Chemistry and Biology. 5:345). Liptoids can be synthesized by elaborating peptoid oligonucleotides and coupling the amino terminal submonomer to a lipid via its amino group (Hunag, et al., 1998. Chemistry and Biology. 5:345).
[0343] It is known in the art that positively charged amino acids can be used for creating highly active cationic lipids (Lewis et al. 1996. Proc. Natl. Acad. Sci. U.S.A. 93:3176). In one embodiment, a composition for delivering oligonucleotides of the invention comprises a number of arginine, lysine, histidine or ornithine residues linked to a lipophilic moiety (see e.g., U.S. Pat. No. 5,777,153).
[0344] In another embodiment, a composition for delivering oligonucleotides of the invention comprises a peptide having from between about one to about four basic residues. These basic residues can be located, e.g., on the amino terminal, C-terminal, or internal region of the peptide. Families of amino acid residues having similar side chains have been defined in the art. These families include amino acids with basic side chains (e.g., lysine, arginine, histidine), acidic side chains (e.g., aspartic acid, glutamic acid), uncharged polar side chains (e.g., glycine (can also be considered non-polar), asparagine, glutamine, serine, threonine, tyrosine, cysteine), nonpolar side chains (e.g., alanine, valine, leucine, isoleucine, proline, phenylalanine, methionine, tryptophan), beta-branched side chains (e.g., threonine, valine, isoleucine) and aromatic side chains (e.g., tyrosine, phenylalanine, tryptophan, histidine). Apart from the basic amino acids, a majority or all of the other residues of the peptide can be selected from the non-basic amino acids, e.g., amino acids other than lysine, arginine, or histidine. Preferably a preponderance of neutral amino acids with long neutral side chains are used.
[0345] In one embodiment, a composition for delivering oligonucleotides of the invention comprises a natural or synthetic polypeptide having one or more gamma carboxyglutamic acid residues, or .gamma.-Gla residues. These gamma carboxyglutamic acid residues may enable the polypeptide to bind to each other and to membrane surfaces. In other words, a polypeptide having a series of .gamma.-Gla may be used as a general delivery modality that helps an RNAi construct to stick to whatever membrane to which it comes in contact. This may at least slow RNAi constructs from being cleared from the blood stream and enhance their chance of homing to the target.
[0346] The gamma carboxyglutamic acid residues may exist in natural proteins (for example, prothrombin has 10 .gamma.-Gla residues). Alternatively, they can be introduced into the purified, recombinantly produced, or chemically synthesized polypeptides by carboxylation using, for example, a vitamin K-dependent carboxylase. The gamma carboxyglutamic acid residues may be consecutive or non-consecutive, and the total number and location of such gamma carboxyglutamic acid residues in the polypeptide can be regulated/fine tuned to achieve different levels of "stickiness" of the polypeptide.
[0347] In one embodiment, the cells to be contacted with an oligonucleotide composition of the invention are contacted with a mixture comprising the oligonucleotide and a mixture comprising a lipid, e.g., one of the lipids or lipid compositions described supra for between about 12 hours to about 24 hours. In another embodiment, the cells to be contacted with an oligonucleotide composition are contacted with a mixture comprising the oligonucleotide and a mixture comprising a lipid, e.g., one of the lipids or lipid compositions described supra for between about 1 and about five days. In one embodiment, the cells are contacted with a mixture comprising a lipid and the oligonucleotide for between about three days to as long as about 30 days. In another embodiment, a mixture comprising a lipid is left in contact with the cells for at least about five to about 20 days. In another embodiment, a mixture comprising a lipid is left in contact with the cells for at least about seven to about 15 days.
[0348] For example, in one embodiment, an oligonucleotide composition can be contacted with cells in the presence of a lipid such as cytofectin CS or GSV (available from Glen Research; Sterling, Va.), GS3815, GS2888 for prolonged incubation periods as described herein.
[0349] In one embodiment, the incubation of the cells with the mixture comprising a lipid and an oligonucleotide composition does not reduce the viability of the cells. Preferably, after the transfection period the cells are substantially viable. In one embodiment, after transfection, the cells are between at least about 70% and at least about 100% viable. In another embodiment, the cells are between at least about 80% and at least about 95% viable. In yet another embodiment, the cells are between at least about 85% and at least about 90% viable.
[0350] In one embodiment, oligonucleotides are modified by attaching a peptide sequence that transports the oligonucleotide into a cell, referred to herein as a "transporting peptide." In one embodiment, the composition includes an oligonucleotide which is complementary to a target nucleic acid molecule encoding the protein, and a covalently attached transporting peptide.
[0351] The language "transporting peptide" includes an amino acid sequence that facilitates the transport of an oligonucleotide into a cell. Exemplary peptides which facilitate the transport of the moieties to which they are linked into cells are known in the art, and include, e.g., HIV TAT transcription factor, lactoferrin, Herpes VP22 protein, and fibroblast growth factor 2 (Pooga et al. 1998. Nature Biotechnology. 16:857; and Derossi et al. 1998. Trends in Cell Biology. 8:84; Elliott and O'Hare. 1997. Cell 88:223).
[0352] Oligonucleotides can be attached to the transporting peptide using known techniques, e.g., (Prochiantz, A. 1996. Curr. Opin. Neurobiol. 6:629; Derossi et al. 1998. Trends Cell Biol. 8:84; Troy et al. 1996. J. Neurosci. 16:253), Vives et al. 1997. J. Biol. Chem. 272:16010). For example, in one embodiment, oligonucleotides bearing an activated thiol group are linked via that thiol group to a cysteine present in a transport peptide (e.g., to the cysteine present in the .beta. turn between the second and the third helix of the antennapedia homeodomain as taught, e.g., in Derossi et al. 1998. Trends Cell Biol. 8:84; Prochiantz. 1996. Current Opinion in Neurobiol. 6:629; Allinquant et al. 1995. J Cell Biol. 128:919). In another embodiment, a Boc-Cys-(Npys)OH group can be coupled to the transport peptide as the last (N-terminal) amino acid and an oligonucleotide bearing an SH group can be coupled to the peptide (Troy et al. 1996. J. Neurosci. 16:253).
[0353] In one embodiment, a linking group can be attached to a nucleomonomer and the transporting peptide can be covalently attached to the linker. In one embodiment, a linker can function as both an attachment site for a transporting peptide and can provide stability against nucleases. Examples of suitable linkers include substituted or unsubstituted C.sub.1-C.sub.20 alkyl chains, C.sub.2-C.sub.20 alkenyl chains, C.sub.2-C.sub.20 alkynyl chains, peptides, and heteroatoms (e.g., S, O, NH, etc.). Other exemplary linkers include bifinctional crosslinking agents such as sulfosuccinimidyl-4-(maleimidophenyl)-butyrate (SMPB) (see, e.g., Smith et al. Biochem J 1991.276: 417-2).
[0354] In one embodiment, oligonucleotides of the invention are synthesized as molecular conjugates which utilize receptor-mediated endocytotic mechanisms for delivering genes into cells (see, e.g., Bunnell et al. 1992. Somatic Cell and Molecular Genetics. 18:559, and the references cited therein).
Targeting Agents
[0355] The delivery of oligonucleotides can also be improved by targeting the oligonucleotides to a cellular receptor. The targeting moieties can be conjugated to the oligonucleotides or attached to a carrier group (i.e., poly(L-lysine) or liposomes) linked to the oligonucleotides. This method is well suited to cells that display specific receptor-mediated endocytosis.
[0356] For instance, oligonucleotide conjugates to 6-phosphomannosylated proteins are internalized 20-fold more efficiently by cells expressing mannose 6-phosphate specific receptors than free oligonucleotides. The oligonucleotides may also be coupled to a ligand for a cellular receptor using a biodegradable linker. In another example, the delivery construct is mannosylated streptavidin which forms a tight complex with biotinylated oligonucleotides. Mannosylated streptavidin was found to increase 20-fold the internalization of biotinylated oligonucleotides. (Vlassov et al. 1994. Biochimica et Biophysica Acta 1197:95-108).
[0357] In addition specific ligands can be conjugated to the polylysine component of polylysine-based delivery systems. For example, transferrin-polylysine, adenovirus-polylysine, and influenza virus hemagglutinin HA-2 N-terminal fusogenic peptides-polylysine conjugates greatly enhance receptor-mediated DNA delivery in eucaryotic cells. Mannosylated glycoprotein conjugated to poly(L-lysine) in aveolar macrophages has been employed to enhance the cellular uptake of oligonucleotides. Liang et al. 1999. Pharmazie 54:559-566.
[0358] Because malignant cells have an increased need for essential nutrients such as folic acid and transferrin, these nutrients can be used to target oligonucleotides to cancerous cells. For example, when folic acid is linked to poly(L-lysine) enhanced oligonucleotide uptake is seen in promyelocytic leukaemia (HL-60) cells and human melanoma (M-14) cells. Ginobbi et al. 1997. Anticancer Res. 17:29. In another example, liposomes coated with maleylated bovine serum albumin, folic acid, or ferric protoporphyrin IX, show enhanced cellular uptake of oligonucleotides in murine macrophages, KB cells, and 2.2.15 human hepatoma cells. Liang et al. 1999. Pharmazie 54:559-566.
[0359] Liposomes naturally accumulate in the liver, spleen, and reticuloendothelial system (so-called, passive targeting). By coupling liposomes to various ligands such as antibodies are protein A, they can be actively targeted to specific cell populations. For example, protein A-bearing liposomes may be pretreated with H-2K specific antibodies which are targeted to the mouse major histocompatibility complex-encoded H-2K protein expressed on L cells. (Vlassov et al. 1994. Biochimica et Biophysica Acta 1197:95-108).
[0360] Other in vitro and/or in vivo delivery of RNAi reagents are known in the art, and can be used to deliver the subject RNAi constructs. See, for example, U.S. patent application publications 20080152661, 20080112916, 20080107694, 20080038296, 20070231392, 20060240093, 20060178327, 20060008910, 20050265957, 20050064595, 20050042227, 20050037496, 20050026286, 20040162235, 20040072785, 20040063654, 20030157030, WO 2008/036825, WO04/065601, and AU2004206255B2, just to name a few (all incorporated by reference).
Administration
[0361] The optimal course of administration or delivery of the oligonucleotides may vary depending upon the desired result and/or on the subject to be treated. As used herein "administration" refers to contacting cells with oligonucleotides and can be performed in vitro or in vivo. The dosage of oligonucleotides may be adjusted to optimally reduce expression of a protein translated from a target nucleic acid molecule, e.g., as measured by a readout of RNA stability or by a therapeutic response, without undue experimentation.
[0362] For example, expression of the protein encoded by the nucleic acid target can be measured to determine whether or not the dosage regimen needs to be adjusted accordingly. In addition, an increase or decrease in RNA or protein levels in a cell or produced by a cell can be measured using any art recognized technique. By determining whether transcription has been decreased, the effectiveness of the oligonucleotide in inducing the cleavage of a target RNA can be determined.
[0363] Any of the above-described oligonucleotide compositions can be used alone or in conjunction with a pharmaceutically acceptable carrier. As used herein, "pharmaceutically acceptable carrier" includes appropriate solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents, and the like. The use of such media and agents for pharmaceutical active substances is well known in the art. Except insofar as any conventional media or agent is incompatible with the active ingredient, it can be used in the therapeutic compositions. Supplementary active ingredients can also be incorporated into the compositions.
[0364] Oligonucleotides may be incorporated into liposomes or liposomes modified with polyethylene glycol or admixed with cationic lipids for parenteral administration. Incorporation of additional substances into the liposome, for example, antibodies reactive against membrane proteins found on specific target cells, can help target the oligonucleotides to specific cell types.
[0365] Moreover, the present invention provides for administering the subject oligonucleotides with an osmotic pump providing continuous infusion of such oligonucleotides, for example, as described in Rataiczak et al. (1992 Proc. Natl. Acad. Sci. USA 89:11823-11827). Such osmotic pumps are commercially available, e.g., from Alzet Inc. (Palo Alto, Calif.). Topical administration and parenteral administration in a cationic lipid carrier are preferred.
[0366] With respect to in vivo applications, the formulations of the present invention can be administered to a patient in a variety of forms adapted to the chosen route of administration, e.g., parenterally, orally, or intraperitoneally. Parenteral administration, which is preferred, includes administration by the following routes: intravenous; intramuscular; interstitially; intraarterially; subcutaneous; intra ocular; intrasynovial; trans epithelial, including transdermal; pulmonary via inhalation; ophthalmic; sublingual and buccal; topically, including ophthalmic; dermal; ocular; rectal; and nasal inhalation via insufflation.
[0367] Pharmaceutical preparations for parenteral administration include aqueous solutions of the active compounds in water-soluble or water-dispersible form. In addition, suspensions of the active compounds as appropriate oily injection suspensions may be administered. Suitable lipophilic solvents or vehicles include fatty oils, for example, sesame oil, or synthetic fatty acid esters, for example, ethyl oleate or triglycerides. Aqueous injection suspensions may contain substances which increase the viscosity of the suspension include, for example, sodium carboxymethyl cellulose, sorbitol, or dextran, optionally, the suspension may also contain stabilizers. The oligonucleotides of the invention can be formulated in liquid solutions, preferably in physiologically compatible buffers such as Hank's solution or Ringer's solution. In addition, the oligonucleotides may be formulated in solid form and redissolved or suspended immediately prior to use. Lyophilized forms are also included in the invention.
[0368] Pharmaceutical preparations for topical administration include transdermal patches, ointments, lotions, creams, gels, drops, sprays, suppositories, liquids and powders. In addition, conventional pharmaceutical carriers, aqueous, powder or oily bases, or thickeners may be used in pharmaceutical preparations for topical administration.
[0369] Pharmaceutical preparations for oral administration include powders or granules, suspensions or solutions in water or non-aqueous media, capsules, sachets or tablets. In addition, thickeners, flavoring agents, diluents, emulsifiers, dispersing aids, or binders may be used in pharmaceutical preparations for oral administration.
[0370] For transmucosal or transdermal administration, penetrants appropriate to the barrier to be permeated are used in the formulation. Such penetrants are known in the art, and include, for example, for transmucosal administration bile salts and fusidic acid derivatives, and detergents. Transmucosal administration may be through nasal sprays or using suppositories. For oral administration, the oligonucleotides are formulated into conventional oral administration forms such as capsules, tablets, and tonics. For topical administration, the oligonucleotides of the invention are formulated into ointments, salves, gels, or creams as known in the art.
[0371] Drug delivery vehicles can be chosen e.g., for in vitro, for systemic, or for topical administration. These vehicles can be designed to serve as a slow release reservoir or to deliver their contents directly to the target cell. An advantage of using some direct delivery drug vehicles is that multiple molecules are delivered per uptake. Such vehicles have been shown to increase the circulation half-life of drugs that would otherwise be rapidly cleared from the blood stream. Some examples of such specialized drug delivery vehicles which fall into this category are liposomes, hydrogels, cyclodextrins, biodegradable nanocapsules, and bioadhesive microspheres.
[0372] The described oligonucleotides may be administered systemically to a subject. Systemic absorption refers to the entry of drugs into the blood stream followed by distribution throughout the entire body. Administration routes which lead to systemic absorption include: intravenous, subcutaneous, intraperitoneal, and intranasal. Each of these administration routes delivers the oligonucleotide to accessible diseased cells. Following subcutaneous administration, the therapeutic agent drains into local lymph nodes and proceeds through the lymphatic network into the circulation. The rate of entry into the circulation has been shown to be a function of molecular weight or size. The use of a liposome or other drug carrier localizes the oligonucleotide at the lymph node. The oligonucleotide can be modified to diffuse into the cell, or the liposome can directly participate in the delivery of either the unmodified or modified oligonucleotide into the cell.
[0373] The chosen method of delivery will result in entry into cells. Preferred delivery methods include liposomes (10-400 nm), hydrogels, controlled-release polymers, and other pharmaceutically applicable vehicles, and microinjection or electroporation (for ex vivo treatments).
[0374] The pharmaceutical preparations of the present invention may be prepared and formulated as emulsions. Emulsions are usually heterogeneous systems of one liquid dispersed in another in the form of droplets usually exceeding 0.1 .mu.m in diameter. The emulsions of the present invention may contain excipients such as emulsifiers, stabilizers, dyes, fats, oils, waxes, fatty acids, fatty alcohols, fatty esters, humectants, hydrophilic colloids, preservatives, and anti-oxidants may also be present in emulsions as needed. These excipients may be present as a solution in either the aqueous phase, oily phase or itself as a separate phase.
[0375] Examples of naturally occurring emulsifiers that may be used in emulsion formulations of the present invention include lanolin, beeswax, phosphatides, lecithin and acacia. Finely divided solids have also been used as good emulsifiers especially in combination with surfactants and in viscous preparations. Examples of finely divided solids that may be used as emulsifiers include polar inorganic solids, such as heavy metal hydroxides, nonswelling clays such as bentonite, attapulgite, hectorite, kaolin, montrnorillonite, colloidal aluminum silicate and colloidal magnesium aluminum silicate, pigments and nonpolar solids such as carbon or glyceryl tristearate.
[0376] Examples of preservatives that may be included in the emulsion formulations include methyl paraben, propyl paraben, quaternary ammonium salts, benzalkonium chloride, esters of p-hydroxybenzoic acid, and boric acid. Examples of antioxidants that may be included in the emulsion formulations include free radical scavengers such as tocopherols, alkyl gallates, butylated hydroxyanisole, butylated hydroxytoluene, or reducing agents such as ascorbic acid and sodium metabisulfite, and antioxidant synergists such as citric acid, tartaric acid, and lecithin.
[0377] In one embodiment, the compositions of oligonucleotides are formulated as microemulsions. A microemulsion is a system of water, oil and amphiphile which is a single optically isotropic and thermodynamically stable liquid solution. Typically microemulsions are prepared by first dispersing an oil in an aqueous surfactant solution and then adding a sufficient amount of a 4th component, generally an intermediate chain-length alcohol to form a transparent system.
[0378] Surfactants that may be used in the preparation of microemulsions include, but are not limited to, ionic surfactants, non-ionic surfactants, Brij 96, polyoxyethylene oleyl ethers, polyglycerol fatty acid esters, tetraglycerol monolaurate (ML310), tetraglycerol monooleate (M0310), hexaglycerol monooleate (P0310), hexaglycerol pentaoleate (P0500), decaglycerol monocaprate (MCA750), decaglycerol monooleate (M0750), decaglycerol sequioleate (S0750), decaglycerol decaoleate (DA0750), alone or in combination with cosurfactants. The cosurfactant, usually a short-chain alcohol such as ethanol, 1-propanol, and 1-butanol, serves to increase the interfacial fluidity by penetrating into the surfactant film and consequently creating a disordered film because of the void space generated among surfactant molecules.
[0379] Microemulsions may, however, be prepared without the use of cosurfactants and alcohol-free self-emulsifying microemulsion systems are known in the art. The aqueous phase may typically be, but is not limited to, water, an aqueous solution of the drug, glycerol, PEG300, PEG400, polyglycerols, propylene glycols, and derivatives of ethylene glycol. The oil phase may include, but is not limited to, materials such as Captex 300, Captex 355, Capmul MCM, fatty acid esters, medium chain (C.sub.8-C.sub.12) mono, di, and tri-glycerides, polyoxyethylated glyceryl fatty acid esters, fatty alcohols, polyglycolized glycerides, saturated polyglycolized C.sub.8-C.sub.10 glycerides, vegetable oils and silicone oil.
[0380] Microemulsions are particularly of interest from the standpoint of drug solubilization and the enhanced absorption of drugs. Lipid based microemulsions (both oil/water and water/oil) have been proposed to enhance the oral bioavailability of drugs.
[0381] Microemulsions offer improved drug solubilization, protection of drug from enzymatic hydrolysis, possible enhancement of drug absorption due to surfactant-induced alterations in membrane fluidity and permeability, ease of preparation, ease of oral administration over solid dosage forms, improved clinical potency, and decreased toxicity (Constantinides et al., Pharmaceutical Research, 1994, 11:1385; Ho et al., J. Pharm. Sci., 1996, 85:138-143). Microemulsions have also been effective in the transdermal delivery of active components in both cosmetic and pharmaceutical applications. It is expected that the microemulsion compositions and formulations of the present invention will facilitate the increased systemic absorption of oligonucleotides from the gastrointestinal tract, as well as improve the local cellular uptake of oligonucleotides within the gastrointestinal tract, vagina, buccal cavity and other areas of administration.
[0382] In an embodiment, the present invention employs various penetration enhancers to affect the efficient delivery of nucleic acids, particularly oligonucleotides, to the skin of animals. Even non-lipophilic drugs may cross cell membranes if the membrane to be crossed is treated with a penetration enhancer. In addition to increasing the diffusion of non-lipophilic drugs across cell membranes, penetration enhancers also act to enhance the permeability of lipophilic drugs.
[0383] Five categories of penetration enhancers that may be used in the present invention include: surfactants, fatty acids, bile salts, chelating agents, and non-chelating non-surfactants. Other agents may be utilized to enhance the penetration of the administered oligonucleotides include: glycols such as ethylene glycol and propylene glycol, pyrrols such as 2-15 pyrrol, azones, and terpenes such as limonene, and menthone.
[0384] The oligonucleotides, especially in lipid formulations, can also be administered by coating a medical device, for example, a catheter, such as an angioplasty balloon catheter, with a cationic lipid formulation. Coating may be achieved, for example, by dipping the medical device into a lipid formulation or a mixture of a lipid formulation and a suitable solvent, for example, an aqueous-based buffer, an aqueous solvent, ethanol, methylene chloride, chloroform and the like. An amount of the formulation will naturally adhere to the surface of the device which is subsequently administered to a patient, as appropriate. Alternatively, a lyophilized mixture of a lipid formulation may be specifically bound to the surface of the device. Such binding techniques are described, for example, in K. Ishihara et al., Journal of Biomedical Materials Research, Vol. 27, pp. 1309-1314 (1993), the disclosures of which are incorporated herein by reference in their entirety.
[0385] The useful dosage to be administered and the particular mode of administration will vary depending upon such factors as the cell type, or for in vivo use, the age, weight and the particular animal and region thereof to be treated, the particular oligonucleotide and delivery method used, the therapeutic or diagnostic use contemplated, and the form of the formulation, for example, suspension, emulsion, micelle or liposome, as will be readily apparent to those skilled in the art. Typically, dosage is administered at lower levels and increased until the desired effect is achieved. When lipids are used to deliver the oligonucleotides, the amount of lipid compound that is administered can vary and generally depends upon the amount of oligonucleotide agent being administered. For example, the weight ratio of lipid compound to oligonucleotide agent is preferably from about 1:1 to about 15:1, with a weight ratio of about 5:1 to about 10:1 being more preferred. Generally, the amount of cationic lipid compound which is administered will vary from between about 0.1 milligram (mg) to about 1 gram (g). By way of general guidance, typically between about 0.1 mg and about 10 mg of the particular oligonucleotide agent, and about 1 mg to about 100 mg of the lipid compositions, each per kilogram of patient body weight, is administered, although higher and lower amounts can be used.
[0386] The agents of the invention are administered to subjects or contacted with cells in a biologically compatible form suitable for pharmaceutical administration. By "biologically compatible form suitable for administration" is meant that the oligonucleotide is administered in a form in which any toxic effects are outweighed by the therapeutic effects of the oligonucleotide. In one embodiment, oligonucleotides can be administered to subjects. Examples of subjects include mammals, e.g., humans and other primates; cows, pigs, horses, and farming (agricultural) animals; dogs, cats, and other domesticated pets; mice, rats, and transgenic non-human animals.
[0387] Administration of an active amount of an oligonucleotide of the present invention is defined as an amount effective, at dosages and for periods of time necessary to achieve the desired result. For example, an active amount of an oligonucleotide may vary according to factors such as the type of cell, the oligonucleotide used, and for in vivo uses the disease state, age, sex, and weight of the individual, and the ability of the oligonucleotide to elicit a desired response in the individual. Establishment of therapeutic levels of oligonucleotides within the cell is dependent upon the rates of uptake and efflux or degradation. Decreasing the degree of degradation prolongs the intracellular half-life of the oligonucleotide. Thus, chemically-modified oligonucleotides, e.g., with modification of the phosphate backbone, may require different dosing.
[0388] The exact dosage of an oligonucleotide and number of doses administered will depend upon the data generated experimentally and in clinical trials. Several factors such as the desired effect, the delivery vehicle, disease indication, and the route of administration, will affect the dosage. Dosages can be readily determined by one of ordinary skill in the art and formulated into the subject pharmaceutical compositions. Preferably, the duration of treatment will extend at least through the course of the disease symptoms.
[0389] Dosage region may be adjusted to provide the optimum therapeutic response. For example, the oligonucleotide may be repeatedly administered, e.g., several doses may be administered daily or the dose may be proportionally reduced as indicated by the exigencies of the therapeutic situation. One of ordinary skill in the art will readily be able to determine appropriate doses and schedules of administration of the subject oligonucleotides, whether the oligonucleotides are to be administered to cells or to subjects.
[0390] Physical methods of introducing nucleic acids include injection of a solution containing the nucleic acid, bombardment by particles covered by the nucleic acid, soaking the cell or organism in a solution of the nucleic acid, or electroporation of cell membranes in the presence of the nucleic acid. A viral construct packaged into a viral particle would accomplish both efficient introduction of an expression construct into the cell and transcription of nucleic acid encoded by the expression construct. Other methods known in the art for introducing nucleic acids to cells may be used, such as lipid-mediated carrier transport, chemical-mediated transport, such as calcium phosphate, and the like. Thus the nucleic acid may be introduced along with components that perform one or more of the following activities: enhance nucleic acid uptake by the cell, inhibit annealing of single strands, stabilize the single strands, or other-wise increase inhibition of the target gene.
[0391] Nucleic acid may be directly introduced into the cell (i.e., intracellularly); or introduced extracellularly into a cavity, interstitial space, into the circulation of an organism, introduced orally or by inhalation, or may be introduced by bathing a cell or organism in a solution containing the nucleic acid. Vascular or extravascular circulation, the blood or lymph system, and the cerebrospinal fluid are sites where the nucleic acid may be introduced.
[0392] The cell with the target gene may be derived from or contained in any organism. The organism may a plant, animal, protozoan, bacterium, virus, or fungus. The plant may be a monocot, dicot or gymnosperm; the animal may be a vertebrate or invertebrate. Preferred microbes are those used in agriculture or by industry, and those that are pathogenic for plants or animals.
[0393] Alternatively, vectors, e.g., transgenes encoding a siRNA of the invention can be engineered into a host cell or transgenic animal using art recognized techniques.
[0394] Another use for the nucleic acids of the present invention (or vectors or transgenes encoding same) is a functional analysis to be carried out in eukaryotic cells, or eukaryotic non-human organisms, preferably mammalian cells or organisms and most preferably human cells, e.g. cell lines such as HeLa or 293 or rodents, e.g. rats and mice. By administering a suitable nucleic acid of the invention which is sufficiently complementary to a target mRNA sequence to direct target-specific RNA interference, a specific knockout or knockdown phenotype can be obtained in a target cell, e.g. in cell culture or in a target organism.
[0395] Thus, a further subject matter of the invention is a eukaryotic cell or a eukaryotic non-human organism exhibiting a target gene-specific knockout or knockdown phenotype comprising a fully or at least partially deficient expression of at least one endogenous target gene wherein said cell or organism is transfected with at least one vector comprising DNA encoding an RNAi agent capable of inhibiting the expression of the target gene. It should be noted that the present invention allows a target-specific knockout or knockdown of several different endogenous genes due to the specificity of the RNAi agent.
[0396] Gene-specific knockout or knockdown phenotypes of cells or non-human organisms, particularly of human cells or non-human mammals may be used in analytic to procedures, e.g. in the functional and/or phenotypical analysis of complex physiological processes such as analysis of gene expression profiles and/or proteomes. Preferably the analysis is carried out by high throughput methods using oligonucleotide based chips.
Assays of Oligonucleotide Stability
[0397] In some embodiments, the oligonucleotides of the invention are stabilized, i.e., substantially resistant to endonuclease and exonuclease degradation. An oligonucleotide is defined as being substantially resistant to nucleases when it is at least about 3-fold more resistant to attack by an endogenous cellular nuclease, and is highly nuclease resistant when it is at least about 6-fold more resistant than a corresponding oligonucleotide. This can be demonstrated by showing that the oligonucleotides of the invention are substantially resistant to nucleases using techniques which are known in the art.
[0398] One way in which substantial stability can be demonstrated is by showing that the oligonucleotides of the invention function when delivered to a cell, e.g., that they reduce transcription or translation of target nucleic acid molecules, e.g., by measuring protein levels or by measuring cleavage of mRNA. Assays which measure the stability of target RNA can be performed at about 24 hours post-transfection (e.g., using Northern blot techniques, RNase Protection Assays, or QC-PCR assays as known in the art). Alternatively, levels of the target protein can be measured. Preferably, in addition to testing the RNA or protein levels of interest, the RNA or protein levels of a control, non-targeted gene will be measured (e.g., actin, or preferably a control with sequence similarity to the target) as a specificity control. RNA or protein measurements can be made using any art-recognized technique. Preferably, measurements will be made beginning at about 16-24 hours post transfection. (M. Y. Chiang, et al. 1991. J Biol Chem. 266:18162-71; T. Fisher, et al. 1993. Nucleic Acids Research. 21 3857).
[0399] The ability of an oligonucleotide composition of the invention to inhibit protein synthesis can be measured using techniques which are known in the art, for example, by detecting an inhibition in gene transcription or protein synthesis. For example, Nuclease S1 mapping can be performed. In another example, Northern blot analysis can be used to measure the presence of RNA encoding a particular protein. For example, total RNA can be prepared over a cesium chloride cushion (see, e.g., Ausebel et al., 1987. Current Protocols in Molecular Biology (Greene & Wiley, New York)). Northern blots can then be made using the RNA and probed (see, e.g., Id.). In another example, the level of the specific mRNA produced by the target protein can be measured, e.g., using PCR. In yet another example, Western blots can be used to measure the amount of target protein present. In still another embodiment, a phenotype influenced by the amount of the protein can be detected. Techniques for performing Western blots are well known in the art, see, e.g., Chen et al. J. Biol. Chem. 271:28259.
[0400] In another example, the promoter sequence of a target gene can be linked to a reporter gene and reporter gene transcription (e.g., as described in more detail below) can be monitored. Alternatively, oligonucleotide compositions that do not target a promoter can be identified by fusing a portion of the target nucleic acid molecule with a reporter gene so that the reporter gene is transcribed. By monitoring a change in the expression of the reporter gene in the presence of the oligonucleotide composition, it is possible to determine the effectiveness of the oligonucleotide composition in inhibiting the expression of the reporter gene. For example, in one embodiment, an effective oligonucleotide composition will reduce the expression of the reporter gene.
[0401] A "reporter gene" is a nucleic acid that expresses a detectable gene product, which may be RNA or protein. Detection of mRNA expression may be accomplished by Northern blotting and detection of protein may be accomplished by staining with antibodies specific to the protein. Preferred reporter genes produce a readily detectable product. A reporter gene may be operably linked with a regulatory DNA sequence such that detection of the reporter gene product provides a measure of the transcriptional activity of the regulatory sequence. In preferred embodiments, the gene product of the reporter gene is detected by an intrinsic activity associated with that product. For instance, the reporter gene may encode a gene product that, by enzymatic activity, gives rise to a detectable signal based on color, fluorescence, or luminescence. Examples of reporter genes include, but are not limited to, those coding for chloramphenicol acetyl transferase (CAT), luciferase, beta-galactosidase, and alkaline phosphatase.
[0402] One skilled in the art would readily recognize numerous reporter genes suitable for use in the present invention. These include, but are not limited to, chloramphenicol acetyltransferase (CAT), luciferase, human growth hormone (hGH), and beta-galactosidase. Examples of such reporter genes can be found in F. A. Ausubel et al., Eds., Current Protocols in Molecular Biology, John Wiley & Sons, New York, (1989). Any gene that encodes a detectable product, e.g., any product having detectable enzymatic activity or against which a specific antibody can be raised, can be used as a reporter gene in the present methods.
[0403] One reporter gene system is the firefly luciferase reporter system. (Gould, S. J., and Subramani, S. 1988. Anal. Biochem., 7:404-408 incorporated herein by reference). The luciferase assay is fast and sensitive. In this assay, a lysate of the test cell is prepared and combined with ATP and the substrate luciferin. The encoded enzyme luciferase catalyzes a rapid, ATP dependent oxidation of the substrate to generate a light-emitting product. The total light output is measured and is proportional to the amount of luciferase present over a wide range of enzyme concentrations.
[0404] CAT is another frequently used reporter gene system; a major advantage of this system is that it has been an extensively validated and is widely accepted as a measure of promoter activity. (Gorman C. M., Moffat, L. F., and Howard, B. H. 1982. Mol. Cell. Biol., 2:1044-1051). In this system, test cells are transfected with CAT expression vectors and incubated with the candidate substance within 2-3 days of the initial transfection. Thereafter, cell extracts are prepared. The extracts are incubated with acetyl CoA and radioactive chloramphenicol. Following the incubation, acetylated chloramphenicol is separated from nonacetylated form by thin layer chromatography. In this assay, the degree of acetylation reflects the CAT gene activity with the particular promoter.
[0405] Another suitable reporter gene system is based on immunologic detection of hGH. This system is also quick and easy to use. (Selden, R., Burke-Howie, K. Rowe, M. E., Goodman, H. M., and Moore, D. D. (1986), Mol. Cell, Biol., 6:3173-3179 incorporated herein by reference). The hGH system is advantageous in that the expressed hGH polypeptide is assayed in the media, rather than in a cell extract. Thus, this system does not require the destruction of the test cells. It will be appreciated that the principle of this reporter gene system is not limited to hGH but rather adapted for use with any polypeptide for which an antibody of acceptable specificity is available or can be prepared.
[0406] In one embodiment, nuclease stability of a double-stranded oligonucleotide of the invention is measured and compared to a control, e.g., an RNAi molecule typically used in the art (e.g., a duplex oligonucleotide of less than 25 nucleotides in length and comprising 2 nucleotide base overhangs) or an unmodified RNA duplex with blunt ends.
[0407] The target RNA cleavage reaction achieved using the siRNAs of the invention is highly sequence specific. Sequence identity may determined by sequence comparison and alignment algorithms known in the art. To determine the percent identity of two nucleic acid sequences (or of two amino acid sequences), the sequences are aligned for optimal comparison purposes (e.g., gaps can be introduced in the first sequence or second sequence for optimal alignment). A preferred, non-limiting example of a local alignment algorithm utilized for the comparison of sequences is the algorithm of Karlin and Altschul (1990) Proc. Natl. Acad. Sci. USA 87:2264-68, modified as in Karlin and Altschul (1993) Proc. Natl. Acad. Sci. USA 90:5873-77. Such an algorithm is incorporated into the BLAST programs (version 2.0) of Altschul, et al. (1990) J. Mol. Biol. 215:403-10. Additionally, numerous commercial entities, such as Dharmacon, and Invitrogen provide access to algorithms on their website. The Whitehead Institute also offers a free siRNA Selection Program. Greater than 90% sequence identity, e.g., 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or even 100% sequence identity, between the siRNA and the portion of the target gene is preferred. Alternatively, the siRNA may be defined functionally as a nucleotide sequence (or oligonucleotide sequence) that is capable of hybridizing with a portion of the target gene transcript. Examples of stringency conditions for polynucleotide hybridization are provided in Sambrook, J., E. F. Fritsch, and T. Maniatis, 1989, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., chapters 9 and 11, and Current Protocols in Molecular Biology, 1995, F. M. Ausubel et al., eds., John Wiley & Sons, Inc., sections 2.10 and 6.3-6.4, incorporated herein by reference.
Therapeutic Use
[0408] By inhibiting the expression of a gene, the oligonucleotide compositions of the present invention can be used to treat any disease involving the expression of a protein. Examples of diseases that can be treated by oligonucleotide compositions, just to illustrate, include: cancer, retinopathies, autoimmune diseases, inflammatory diseases (i.e., ICAM-1 related disorders, Psoriasis, Ulcerative Colitus, Crohn's disease), viral diseases (i.e., HIV, Hepatitis C), miRNA disorders, and cardiovascular diseases.
[0409] In one embodiment, in vitro treatment of cells with oligonucleotides can be used for ex vivo therapy of cells removed from a subject (e.g., for treatment of leukemia or viral infection) or for treatment of cells which did not originate in the subject, but are to be administered to the subject (e.g., to eliminate transplantation antigen expression on cells to be transplanted into a subject). In addition, in vitro treatment of cells can be used in non-therapeutic settings, e.g., to evaluate gene function, to study gene regulation and protein synthesis or to evaluate improvements made to oligonucleotides designed to modulate gene expression or protein synthesis. In vivo treatment of cells can be useful in certain clinical settings where it is desirable to inhibit the expression of a protein. There are numerous medical conditions for which antisense therapy is reported to be suitable (see, e.g., U.S. Pat. No. 5,830,653) as well as respiratory syncytial virus infection (WO 95/22,553) influenza virus (WO 94/23,028), and malignancies (WO 94/08,003). Other examples of clinical uses of antisense sequences are reviewed, e.g., in Glaser. 1996. Genetic Engineering News 16:1. Exemplary targets for cleavage by oligonucleotides include, e.g., protein kinase Ca, ICAM-1, c-raf kinase, p53, c-myb, and the bcr/abl fusion gene found in chronic myelogenous leukemia.
[0410] The subject nucleic acids can be used in RNAi-based therapy in any animal having RNAi pathway, such as human, non-human primate, non-human mammal, non-human vertebrates, rodents (mice, rats, hamsters, rabbits, etc.), domestic livestock animals, pets (cats, dogs, etc.), Xenopus, fish, insects (Drosophila, etc.), and worms (C. elegans), etc.
[0411] The invention provides methods for inhibiting or preventing in a subject, a disease or condition associated with an aberrant or unwanted target gene expression or activity, by administering to the subject a nucleic acid of the invention. If appropriate, subjects are first treated with a priming agent so as to be more responsive to the subsequent RNAi therapy. Subjects at risk for a disease which is caused or contributed to by aberrant or unwanted target gene expression or activity can be identified by, for example, any or a combination of diagnostic or prognostic assays known in the art. Administration of a prophylactic agent can occur prior to the manifestation of symptoms characteristic of the target gene aberrancy, such that a disease or disorder is prevented or, alternatively, delayed in its progression. Depending on the type of target gene aberrancy, for example, a target gene, target gene agonist or target gene antagonist agent can be used for treating the subject.
[0412] In another aspect, the invention pertains to methods of modulating target gene expression, protein expression or activity for therapeutic purposes. Accordingly, in an exemplary embodiment, the methods of the invention involve contacting a cell capable of expressing target gene with a nucleic acid of the invention that is specific for the target gene or protein (e.g., is specific for the mRNA encoded by said gene or specifying the amino acid sequence of said protein) such that expression or one or more of the activities of target protein is modulated. These methods can be performed in vitro (e.g., by culturing the cell with the agent), in vivo (e.g., by administering the agent to a subject), or ex vivo. The subjects may be first treated with a priming agent so as to be more responsive to the subsequent RNAi therapy if desired. As such, the present invention provides methods of treating a subject afflicted with a disease or disorder characterized by aberrant or unwanted expression or activity of a target gene polypeptide or nucleic acid molecule. Inhibition of target gene activity is desirable in situations in which target gene is abnormally unregulated and/or in which decreased target gene activity is likely to have a beneficial effect.
[0413] Thus the therapeutic agents of the invention can be administered to subjects to treat (prophylactically or therapeutically) disorders associated with aberrant or unwanted target gene activity. In conjunction with such treatment, pharmacogenomics (i.e., the study of the relationship between an individual's genotype and that individual's response to a foreign compound or drug) may be considered. Differences in metabolism of therapeutics can lead to severe toxicity or therapeutic failure by altering the relation between dose and blood concentration of the pharmacologically active drug. Thus, a physician or clinician may consider applying knowledge obtained in relevant pharmacogenomics studies in determining whether to administer a therapeutic agent as well as tailoring the dosage and/or therapeutic regimen of treatment with a therapeutic agent. Pharmacogenomics deals with clinically significant hereditary variations in the response to drugs due to altered drug disposition and abnormal action in affected persons.
[0414] For the purposes of the invention, ranges may be expressed herein as from "about" one particular value, and/or to "about" another particular value. When such a range is expressed, another embodiment includes from the one particular value and/or to the other particular value. Similarly, when values are expressed as approximations, by use of the antecedent "about," it will be understood that the particular value forms another embodiment. It will be further understood that the endpoints of each of the ranges are significant both in relation to the other endpoint, and independently of the other endpoint.
[0415] Moreover, for the purposes of the present invention, the term "a" or "an" entity refers to one or more of that entity; for example, "a protein" or "a nucleic acid molecule" refers to one or more of those compounds or at least one compound. As such, the terms "a" (or "an"), "one or more" and "at least one" can be used interchangeably herein. It is also to be noted that the terms "comprising", "including", and "having" can be used interchangeably. Furthermore, a compound "selected from the group consisting of" refers to one or more of the compounds in the list that follows, including mixtures (i.e., combinations) of two or more of the compounds. According to the present invention, an isolated, or biologically pure, protein or nucleic acid molecule is a compound that has been removed from its natural milieu. As such, "isolated" and "biologically pure" do not necessarily reflect the extent to which the compound has been purified. An isolated compound of the present invention can be obtained from its natural source, can be produced using molecular biology techniques or can be produced by chemical synthesis.
[0416] The present invention is further illustrated by the following Examples, which in no way should be construed as further limiting. The entire contents of all of the references (including literature references, issued patents, published patent applications, and co-pending patent applications) cited throughout this application are hereby expressly incorporated by reference.
EXAMPLES
Example 1: Inhibition of Gene Expression Using Minimum Length Trigger RNAs
[0417] Transfection of Minimum Length Trigger (mlt) RNA
[0418] mltRNA constructs were chemically synthesized (Integrated DNA Technologies, Coralville, Iowa) and transfected into HEK293 cells (ATCC, Manassas, Va.) using the Lipofectamine RNAiMAX (Invitrogen, Carlsbad, Calif.) reagent according to manufacturer's instructions. In brief, RNA was diluted to a 12.times. concentration and then combined with a 12.times. concentration of Lipofectamine RNAiMAX to complex. The RNA and transfection reagent were allowed to complex at room temperature for 20 minutes and make a 6.times. concentration. While complexing, HEK293 cells were washed, trypsinized and counted. The cells were diluted to a concentration recommended by the manufacturer and previously described conditions which was at 1.times.10.sup.5 cells/ml. When RNA had completed complexing with the RNAiMAX transfection reagent, 20 ul of the complexes were added to the appropriate well of the 96-well plate in triplicate. Cells were added to each well (100 ul volume) to make the final cell count per well at 1.times.10.sup.4 cells/well. The volume of cells diluted the 6.times. concentration of complex to 1.times. which was equal to a concentration noted (between 10-0.05 nM). Cells were incubated for 24 or 48 hours under normal growth conditions.
[0419] After 24 or 48 hour incubation cells were lysed and gene silencing activity was measured using the QuantiGene assay (Panomics, Freemont, Calif.) which employs bDNA hybridization technology. The assay was carried out according to manufacturer's instructions.
.DELTA.G Calculation
[0420] .DELTA.G was calculated using Mfold, available through the Mfold internet site (http://mfold.bioinfo.rpi.edu/cgi-bin/rna-form1.cfi). Methods for calculating .DELTA.G are described in, and are incorporated by reference from, the following references: Zuker, M. (2003) Nucleic Acids Res., 31(13):3406-15; Mathews, D. H., Sabina, J., Zuker, M. and Turner, D. H. (1999) J. Mol. Biol. 288:911-940; Mathews, D. H., Disney, M. D., Childs, J. L., Schroeder, S. J., Zuker, M., and Turner, D. H. (2004) Proc. Natl. Acad. Sci. 101:7287-7292; Duan, S., Mathews, D. H., and Turner, D. H. (2006) Biochemistry 45:9819-9832; Wuchty, S., Fontana, W., Hofacker, I. L., and Schuster, P. (1999) Biopolymers 49:145-165.
Example 2: Optimization of sd-rxRNA.sup.nano Molecules for Gene Silencing
[0421] Asymmetric double stranded RNAi molecules, with minimal double stranded regions, were developed herein and are highly effective at gene silencing. These molecules can contain a variety of chemical modifications on the sense and/or anti-sense strands, and can be conjugated to sterol-like compounds such as cholesterol.
[0422] FIGS. 1-3 present schematics of RNAi molecules associated with the invention. In the asymmetric molecules, which contain a sense and anti-sense strand, either of the strands can be the longer strand. Either strand can also contain a single-stranded region. There can also be mismatches between the sense and anti-sense strand, as indicated in FIG. 1D. Preferably, one end of the double-stranded molecule is either blunt-ended or contains a short overhang such as an overhang of one nucleotide. FIG. 2 indicates types of chemical modifications applied to the sense and anti-sense strands including 2'F, 2'OMe, hydrophobic modifications and phosphorothioate modifications. Preferably, the single stranded region of the molecule contains multiple phosphorothioate modifications. Hydrophobicity of molecules can be increased using such compounds as 4-pyridyl at 5-U, 2-pyridyl at 5-U, isobutyl at 5-U and indolyl at 5-U (FIG. 2). Proteins or peptides such as protamine (or other Arg rich peptides), spermidine or other similar chemical structures can also be used to block duplex charge and facilitate cellular entry (FIG. 3). Increased hydrophobicity can be achieved through either covalent or non-covalent modifications. Several positively charged chemicals, which might be used for polynucleotide charge blockage are depicted in FIG. 4.
[0423] Chemical modifications of polynucleotides, such as the guide strand in a duplex molecule, can facilitate RISC entry. FIG. 5 depicts single stranded polynucleotides, representing a guide strand in a duplex molecule, with a variety of chemical modifications including 2'd, 2'OMe, 2'F, hydrophobic modifications, phosphorothioate modifications, and attachment of conjugates such as "X" in FIG. 5, where X can be a small molecule with high affinity to a PAZ domain, or sterol-type entity. Similarly, FIG. 6 depicts single stranded polynucleotides, representing a passenger strand in a duplex molecule, with proposed structural and chemical compositions of RISC substrate inhibitors. Combinations of chemical modifications can ensure efficient uptake and efficient binding to preloaded RISC complexes.
[0424] FIG. 7 depicts structures of polynucleotides with sterol-type molecules attached, where R represents a polycarbonic tail of 9 carbons or longer. FIG. 8 presents examples of naturally occurring phytosterols with a polycarbon chain longer than 8 attached at position 17. More than 250 different types of phytosterols are known. FIG. 9 presents examples of sterol-like structures with variations in the sizes of the polycarbon chains attached at position 17. FIG. 91 presents further examples of sterol-type molecules that can be used as a hydrophobic entity in place of cholesterol. FIG. 92 presents further examples of hydrophobic molecules that might be used as hydrophobic entities in place of cholestesterol. Optimization of such characteristics can improve uptake properties of the RNAi molecules. FIG. 10 presents data adapted from Martins et al. (J Lipid Research), showing that the percentage of liver uptake and plasma clearance of lipid emulsions containing sterol-type molecules is directly affected by the size of the attached polycarbon chain at position 17. FIG. 11 depicts a micelle formed from a mixture of polynucleotides attached to hydrophobic conjugates and fatty acids. FIG. 12 describes how alteration in lipid composition can affect pharmacokinetic behavior and tissue distribution of hydrophobically modified and/or hydrophobically conjugated polynucleotides. In particular, the use of lipid mixtures that are enriched in linoleic acid and cardiolipin results in preferential uptake by cardiomyocites.
[0425] FIG. 13 depicts examples of RNAi constructs and controls designed to target MAP4K4 expression. FIGS. 14 and 15 reveal that RNAi constructs with minimal duplex regions (such as duplex regions of approximately 13 nucleotides) are effective in mediating RNA silencing in cell culture. Parameters associated with these RNA molecules are shown in FIG. 16. FIG. 17 depicts examples of RNAi constructs and controls designed to target SOD1 expression. FIGS. 18 and 19 reveal the results of gene silencing experiments using these RNAi molecules to target SOD1 in cells. FIG. 20 presents a schematic indicating that RNA molecules with double stranded regions that are less than 10 nucleotides are not cleaved by Dicer, and FIG. 21 presents a schematic of a hypothetical RNAi model for RNA induced gene silencing.
[0426] The RNA molecules described herein were subject to a variety of chemical modifications on the sense and antisense strands, and the effects of such modifications were observed. RNAi molecules were synthesized and optimized through testing of a variety of modifications. In first generation optimization, the sense (passenger) and anti-sense (guide) strands of the sd-rxRNA' molecules were modified for example through incorporation of C and U 2'OMe modifications, 2'F modifications, phosphorothioate modifications, phosphorylation, and conjugation of cholesterol. Molecules were tested for inhibition of MAP4K4 expression in cells including HeLa, primary mouse hepatocytes and primary human hepatocytes through both lipid-mediated and passive uptake transfection.
[0427] FIG. 22 reveals that chemical modifications can enhance gene silencing. In particular, modifying the guide strand with 2'F UC modifications, and with a stretch of phosphorothioate modifications, combined with complete CU O'Me modification of the passenger strands, resulted in molecules that were highly effective in gene silencing. The effect of chemical modification on in vitro efficacy in un-assisted delivery in HeLa cells was also examined. FIG. 23 reveals that compounds lacking any of 2'F, 2'OMe, a stretch of phosphorothioate modifications, or cholesterol conjugates, were completely inactive in passive uptake. A combination of all 4 types of chemical modifications, for example in compound 12386, was found to be highly effective in gene silencing. FIG. 24 also shows the effectiveness of compound 12386 in gene silencing.
[0428] Optimization of the length of the oligonucleotide was also investigated. FIGS. 25 and 26 reveal that oligonucleotides with a length of 21 nucleotides were more effective than oligonucletides with a length of 25 nucleotides, indicating that reduction in the size of an RNA molecule can improve efficiency, potentially by assisting in its uptake. Screening was also conducted to optimize the size of the duplex region of double stranded RNA molecules. FIG. 88 reveals that compounds with duplexes of 10 nucleotides were effective in inducing gene silencing. Positioning of the sense strand relative to the guide strand can also be critical for silencing gene expression (FIG. 89). In this assay, a blunt end was found to be most effective. 3' overhangs were tolerated, but 5' overhangs resulted in a complete loss of functionality. The guide strand can be effective in gene silencing when hybiridized to a sense strand of varying lengths (FIG. 90). In this assay presented in FIG. 90, the compounds were introduced into HeLa cells via lipid mediated transfection.
[0429] The importance of phosphorothioate content of the RNA molecule for unassisted delivery was also investigated. FIG. 27 presents the results of a systematic screen that identified that the presence of at least 2-12 phosphorothioates in the guide strand as being highly advantageous for achieving uptake, with 4-8 being the preferred number. FIG. 27 also shows that presence or absence of phosphorothioate modifications in the sense strand did not alter efficacy.
[0430] FIGS. 28-29 reveal the effects of passive uptake of RNA compounds on gene silencing in primary mouse hepatocytes. nanoRNA molecules were found to be highly effective, especially at a concentration of 1 .mu.M (FIG. 28). FIGS. 30 and 31 reveal that the RNA compounds associated with the invention were also effective in gene silencing following passive uptake in primary human hepatocytes. The cellular localization of the RNA molecules associated with the invention was examined and compared to the localization of Chol-siRNA (Alnylam) molecules, as shown in FIGS. 32 and 33.
[0431] A summary of 1.sup.st generation sd-rxRNA molecules is presented in FIG. 21. Chemical modifications were introduced into the RNA molecules, at least in part, to increase potency, such as through optimization of nucleotide length and phosphorothioate content, to reduce toxicity, such as through replacing 2'F modifications on the guide strand with other modifications, to improve delivery such as by adding or conjugating the RNA molecules to linker and sterol modalities, and to improve the ease of manufacturing the RNA molecules. FIG. 35 presents schematic depictions of some of the chemical modifications that were screened in 1.sup.st generation molecules. Parameters that were optimized for the guide strand included nucleotide length (e.g., 19, 21 and 25 nucleotides), phosphorothioate content (e.g., 0-18 phosphorothioate linkages) and replacement of 2'F groups with 2'OMe and 5 Me C or riboThymidine. Parameters that were optimized for the sense strand included nucleotide length (e.g., 11, 13 and 19 nucleotides), phosphorothioate content (e.g., 0-4 phosphorothioate linkages), and 2'OMe modifications. FIG. 36 summarizes parameters that were screened. For example, the nucleotide length and the phosphorothioate tail length were modified and screened for optimization, as were the additions of 2'OMe C and U modifications. Guide strand length and the length of the phosphorothioate modified stretch of nucleotides were found to influence efficacy (FIGS. 37-38). Phosphorothioate modifications were tolerated in the guide strand and were found to influence passive uptake (FIGS. 39-42).
[0432] FIG. 43 presents a schematic revealing guide strand chemical modifications that were screened. FIGS. 44 and 45 reveal that 2'OMe modifications were tolerated in the 3' end of the guide strand. In particular, 2'OMe modifications in positions 1 and 11-18 were well tolerated. The 2'OMe modifications in the seed area were tolerated but resulted in slight reduction of efficacy. Ribo-modifications in the seed were also well tolerated. These data indicate that the molecules associated with the invention offer the significant advantage of having reduced or no 2'F modification content. This is advantageous because 2'F modifications are thought to generate toxicity in vivo. In some instances, a complete substitution of 2'F modifications with 2'OMe was found to lead to some reduction in potency. However, the 2' OMe substituted molecules were still very active. A molecule with 50% reduction in 2'F content (including at positions 11, 16-18 which were changed to 2'OMe modifications), was found to have comparable efficacy to a compound with complete 2'F C and U modification. 2'OMe modification in position was found in some instances to reduce efficacy, although this can be at least partially compensated by 2'OMe modification in position 1 (with chemical phosphate). In some instances, 5 Me C and/or ribothymidine substitution for 2'F modifications led to a reduction in passive uptake efficacy, but increased potency in lipid mediated transfections compared to 2'F modifications. Optimization results for lipid mediated transfection were not necessarily the same as for passive uptake.
[0433] Modifications to the sense strand were also developed and tested, as depicted in FIG. 46. FIG. 47 reveals that in some instances, a sense strand length between 10-15 bases was found to be optimal. For the molecules tested in FIG. 47, an increase in the sense strand length resulted in reduction of passive uptake, however an increase in sense strand length may be tolerated for some compounds. FIG. 47 also reveals that LNA modification of the sense strand demonstrated similar efficacy to non-LNA containing compounds. In general, the addition of LNA or other thermodynamically stabilizing compounds has been found to be beneficial, in some instances resulting in converting non-functional sequences to functional sequences. FIG. 48 also presents data on sense strand length optimization, while FIG. 49 shows that phosphorothioate modification of the sense strand is not required for passive uptake.
[0434] Based on the above-described optimization experiments, 2.sup.nd generation RNA molecules were developed. As shown in FIG. 50, these molecules contained reduced phosphorothioate modification content and reduced 2'F modification content, relative to 1.sup.st generation RNA molecules. Significantly, these RNA molecules exhibit spontaneous cellular uptake and efficacy without a delivery vehicle (FIG. 51). These molecules can achieve self-delivery (i.e., with no transfection reagent) and following self-delivery can exhibit nanomolar activity in cell culture. These molecules can also be delivered using lipid-mediated transfection, and exhibit picomolar activity levels following transfection. Significantly, these molecules exhibit highly efficient uptake, 95% by most cells in cell culture, and are stable for more than three days in the presence of 100% human serum. These molecules are also highly specific and exhibit little or no immune induction. FIGS. 52 and 53 reveal the significance of chemical modifications and the configurations of such modifications in influencing the properties of the RNA molecules associated with the invention.
[0435] Linker chemistry was also tested in conjunction with the RNA molecules associated with the invention. As depicted in FIG. 54, 2.sup.nd generation RNA molecules were synthesized with sterol-type molecules attached through TEG and amino caproic acid linkers. Both linkers showed identical potency. This functionality of the RNA molecules, independent of linker chemistry offers additional advantages in terms of scale up and synthesis and demonstrates that the mechanism of function of these RNA molecules is very different from other previously described RNA molecules.
[0436] Stability of the chemically modified sd-rxRNA molecules described herein in human serum is shown in FIG. 55 in comparison to unmodified RNA. The duplex molecules were incubated in 75% serum at 37.degree. C. for the indicated periods of time. The level of degradation was determined by running the samples on non-denaturing gels and staining with SYBGR.
[0437] FIGS. 56 and 57 present data on cellular uptake of the sd-rxRNA molecules. FIG. 56 shows that minimizing the length of the RNA molecule is importance for cellular uptake, while FIG. 57 presents data showing target gene silencing after spontaneous cellular uptake in mouse PEC-derived macrophages. FIG. 58 demonstrates spontaneous uptake and target gene silencing in primary cells. FIG. 59 shows the results of delivery of sd-rxRNA molecules associated with the invention to RPE cells with no formulation. Imaging with Hoechst and DY547 reveals the clear presence of a signal representing the RNA molecule in the sd-rxRNA sample, while no signal is detectable in the other samples including the samples competing a competing conjugate, an rxRNA, and an untransfected control. FIG. 60 reveals silencing of target gene expression in RPE cells treated with sd-rxRNA molecules associated with the invention following 24-48 hours without any transfection formulation.
[0438] FIG. 61 shows further optimization of the chemical/structural composition of sd-rxRNA compounds. In some instances, preferred properties included an antisense strand that was 17-21 nucleotides long, a sense strand that was 10-15 nucleotides long, phosphorothioate modification of 2-12 nucleotides within the single stranded region of the molecule, preferentially phosphorothioate modification of 6-8 nucleotides within the single stranded region, and 2'OMe modification at the majority of positions within the sense strand, with or without phosphorothioate modification. Any linker chemistry can be used to attach the hydrophobic moiety, such as cholesterol, to the 3' end of the sense strand. Version GM molecules, as shown in FIG. 61, have no 2'F modifications. Significantly, there is was no impact on efficacy in these molecules.
[0439] FIG. 62 demonstrates the superior performance of sd-rxRNA compounds compared to compounds published by Wolfrum et. al. Nature Biotech, 2007. Both generation I and II compounds (GI and GIIa) developed herein show great efficacy in reducing target gene expression. By contrast, when the chemistry described in Wolfrum et al. (all oligos contain cholesterol conjugated to the 3' end of the sense strand) was applied to the same sequence in a context of conventional siRNA (19 bp duplex with two overhang) the compound was practically inactive. These data emphasize the significance of the combination of chemical modifications and assymetrical molecules described herein, producing highly effective RNA compounds.
[0440] FIG. 63 shows localization of sd-rxRNA molecules developed herein compared to localization of other RNA molecules such as those described in Soutschek et al. (2004) Nature, 432:173. sd-rxRNA molecules accumulate inside the cells whereas competing conjugate RNAs accumulate on the surface of cells. Significantly, FIG. 64 shows that sd-rxRNA molecules, but not competitor molecules such as those described in Soutschek et al. are internalized within minutes. FIG. 65 compares localization of sd-rxRNA molecules compared to regular siRNA-cholesterol, as described in Soutschek et al. A signal representing the RNA molecule is clearly detected for the sd-rxRNA molecule in tissue culture RPE cells, following local delivery to compromised skin, and following systemic delivery where uptake to the liver is seen. In each case, no signal is detected for the regular siRNA-cholesterol molecule. The sd-rxRNA molecule thus has drastically better cellular and tissue uptake characteristics when compared to conventional cholesterol conjugated siRNAs such as those described in Soutschek et al. The level of uptake is at least order of magnitude higher and is due at least in part to the unique combination of chemistries and conjugated structure. Superior delivery of sd-rxRNA relative to previously described RNA molecules is also demonstrated in FIGS. 66 and 67.
[0441] Based on the analysis of 2.sup.nd generation RNA molecules associated with the invention, a screen was performed to identify functional molecules for targeting the SPP1/PPIB gene. As revealed in FIG. 68, several effective molecules were identified, with 14131 being the most effective. The compounds were added to A-549 cells and then the level of SPP1/PPIB ratio was determined by B-DNA after 48 hours.
[0442] FIG. 69 reveals efficient cellular uptake of sd-rxRNA within minutes of exposure. This is a unique characteristics of these molecules, not observed with any other RNAi compounds. Compounds described in Soutschek et al. were used as negative controls. FIG. 70 reveals that the uptake and gene silencing of the sd-rxRNA is effective in multiple different cell types including SH-SY5Y neuroblastoma derived cells, ARPE-19 (retinal pigment epithelium) cells, primary hepatocytes, and primary macrophages. In each case silencing was confirmed by looking at target gene expression by a Branched DNA assay.
[0443] FIG. 70 reveals that sd-rxRNA is active in the presence or absence of serum. While a slight reduction in efficacy (2-5 fold) was observed in the presence of serum, this small reduction in efficacy in the presence of serum differentiate the sd-rxRNA molecules from previously described molecules which exhibited a larger reduction in efficacy in the presence of serum. This demonstrated level of efficacy in the presence of serum creates a foundation for in vivo efficacy.
[0444] FIG. 72 reveals efficient tissue penetration and cellular uptake upon single intradermal injection. This data indicates the potential of the sd-rxRNA compounds described herein for silencing genes in any dermatology applications, and also represents a model for local delivery of sd-rxRNA compounds. FIG. 73 also demonstrates efficient cellular uptake and in vivo silencing with sd-rxRNA following intradermal injection. Silencing is determined as the level of MAP4K4 knockdown in several individual biopsies taken from the site of injection as compared to biopsies taken from a site injected with a negative control. FIG. 74 reveals that sd-rxRNA compounds has improved blood clearance and induced effective gene silencing in vivo in the liver upon systemic administration. In comparison to the RNA molecules described by Soutschek et al., the level of liver uptake at identical dose level is at least 50 fold higher with the sd-rxRNA molecules. The uptake results in productive silencing. sd-rxRNA compounds are also characterized by improved blood clearance kinetics.
[0445] The effect of 5-Methly C modifications was also examined. FIG. 75 demonstrates that the presence of 5-Methyl C in an RNAi molecule resulted in increased potency in lipid mediated transfection. This suggests that hydrophobic modification of Cs and Us in an RNAi molecule can be beneficial. These types of modifications can also be used in the context 2' ribose modified bases to ensure optimal stability and efficacy. FIG. 76 presents data showing that incorporation of 5-Methyl C and/or ribothymidine in the guide strand can in some instances reduce efficacy.
[0446] FIG. 77 reveals that sd-rxRNA molecules are more effective than competitor molecules such as molecules described in Soutschek et al., in systemic delivery to the liver. A signal representing the RNA molecule is clearly visible in the sample containing sd-rxRNA, while no signal representing the RNA molecule is visible in the sample containing the competitor RNA molecule.
[0447] The addition of hydrophobic conjugates to the sd-rxRNA molecules was also explored (FIGS. 78-83). FIG. 78 presents schematics demonstrating 5-uridyl modifications with improved hydrophobicity characteristics. Incorporation of such modifications into sd-rxRNA compounds can increase cellular and tissue uptake properties. FIG. 78B presents a new type of RNAi compound modification which can be applied to compounds to improve cellular uptake and pharmacokinetic behavior. Significantly, this type of modification, when applied to sd-rxRNA compounds, may contribute to making such compounds orally available. FIG. 79 presents schematics revealing the structures of synthesized modified sterol-type molecules, where the length and structure of the C17 attached tail is modified. Without wishing to be bound by any theory, the length of the C17 attached tail may contribute to improving in vitro and in vivo efficacy of sd-rxRNA compounds.
[0448] FIG. 80 presents a schematic demonstrating the lithocholic acid route to long side chain cholesterols. FIG. 81 presents a schematic demonstrating a route to 5-uridyl phosphoramidite synthesis. FIG. 82 presents a schematic demonstrating synthesis of tri-functional hydroxyprolinol linker for 3'-cholesterol attachment. FIG. 83 presents a schematic demonstrating synthesis of solid support for the manufacture of a shorter asymmetric RNAi compound strand.
[0449] A screen was conducted to identify compounds that could effectively silence expression of SPP1 (Osteopontin). Compounds targeting SPP1 were added to A549 cells (using passive transfection), and the level of SPP1 expression was evaluated at 48 hours. Several novel compounds effective in SPP1 silencing were identified. Compounds that were effective in silencing of SPP1 included 14116, 14121, 14131, 14134, 14139, 14149, and 14152 (FIGS. 84-86). The most potent compound in this assay was 14131 (FIG. 84). The efficacy of these sd-rxRNA compounds in silencing SPP1 expression was independently validated (FIG. 85).
[0450] A similar screen was conducted to identify compounds that could effectively silence expression of CTGF (FIGS. 86-87). Compounds that were effective in silencing of CTGF included 14017, 14013, 14016, 14022, 14025, 14027.
Methods
Transfection of Sd-rxRNA.sup.nano
Lipid Mediated Transfection
[0451] sd-rxRNA.sup.nano constructs were chemically synthesized (Dharmacon, Lafayette, Colo.) and transfected into HEK293 cells (ATCC, Manassas, Va.) using Lipofectamine RNAiMAX (Invitrogen, Carlsbad, Calif.) according to the manufacturer's instructions. In brief, RNA was diluted to a 12.times. concentration in Opti-MEM.RTM. Reduced Serum Media (Invitrogen, Carlsbad, Calif.) and then combined with a 12.times. concentration of Lipofectamine RNAiMAX. The RNA and transfection reagent were allowed to complex at room temperature for 20 minutes and make a 6.times. concentration. While complexing, HEK293 cells were washed, trypsinized and counted. The cells were diluted to a concentration recommended by the manufacturer and previously described of 1.times.10.sup.5 cells/ml. When RNA had completed complexing with the RNAiMAX transfection reagent, 20 ul of the complexes were added to the appropriate well of the 96-well plate in triplicate. Cells were added to each well (100 ul volume) to make the final cell count per well 1.times.10.sup.4 cells/well. The volume of cells diluted the 6.times. concentration of complex to 1.times. (between 10-0.05 nM). Cells were incubated for 24 or 48 hours under normal growth conditions. After 24 or 48 hour incubation, cells were lysed and gene silencing activity was measured using the QuantiGene assay (Panomics, Freemont, Calif.) which employs bDNA hybridization technology. The assay was carried out according to manufacturer's instructions.
Passive Uptake Transfection
[0452] sd-rxRNA.sup.nano constructs were chemically synthesized (Dharmacon, Lafayette, Colo.). 24 hours prior to transfection, HeLa cells (ATCC, Manassas, Va.) were plated at 1.times.10.sup.4 cells/well in a 96 well plate under normal growth conditions (DMEM, 10% FBS and 1% Penicillin and Streptomycin). Prior to transfection of HeLa cells, sd-rxRNA.sup.nano were diluted to a final concentration of 0.01 uM to 1 uM in Accell siRNA Delivery Media (Dharmacon, Lafayette, Colo.). Normal growth media was aspirated off cells and 100 uL of Accell Delivery media containing the appropriate concentration of sd-rxRNAnano was applied to the cells. 48 hours post transfection, delivery media was aspirated off the cells and normal growth media was applied to cells for an additional 24 hours.
[0453] After 48 or 72 hour incubation, cells were lysed and gene silencing activity was measured using the QuantiGene assay (Panomics, Freemont, Calif.) according to manufacturer's instructions.
TABLE-US-00001 TABLE 1 Oligo Accession Gene ID Number Number number Gene Name Symbol APOB-10167-20-12138 12138 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-10167-20-12139 12139 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4-2931-13-12266 12266 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12293 12293 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12383 12383 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12384 12384 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12385 12385 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12386 12386 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12387 12387 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-15-12388 12388 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-13-12432 12432 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-13-12266.2 12266.2 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 APOB--21-12434 12434 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--21-12435 12435 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4-2931-16-12451 12451 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12452 12452 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-16-12453 12453 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-17-12454 12454 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-17-12455 12455 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-19-12456 12456 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 --27-12480 12480 --27-12481 12481 APOB-10167-21-12505 12505 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-10167-21-12506 12506 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4-2931-16-12539 12539 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 APOB-10167-21-12505.2 12505.2 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-10167-21-12506.2 12506.2 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4--13-12565 12565 MAP4K4 MAP4K4-2931-16-12386.2 12386.2 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-13-12815 12815 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 APOB--13-12957 12957 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4--16-12983 12983 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12984 12984 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12985 12985 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12986 12986 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12987 12987 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12988 12988 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12989 12989 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12990 12990 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12991 12991 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12992 12992 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12993 12993 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12994 12994 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4--16-12995 12995 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-19-13012 13012 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 MAP4K4-2931-19-13016 13016 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 PPIB--13-13021 13021 NM_000942 Peptidylprolyl Isomerase B PPIB (cyclophilin B) pGL3-1172-13-13038 13038 U47296 Cloning vector pGL3-Control pGL3 pGL3-1172-13-13040 13040 U47296 Cloning vector pGL3-Control pGL3 --16-13047 13047 SOD1-530-13-13090 13090 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-523-13-13091 13091 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-535-13-13092 13092 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-536-13-13093 13093 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-396-13-13094 13094 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-385-13-13095 13095 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-195-13-13096 13096 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) APOB-4314-13-13115 13115 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-3384-13-13116 13116 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-3547-13-13117 13117 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-4318-13-13118 13118 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-3741-13-13119 13119 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) PPIB--16-13136 13136 NM_000942 Peptidylprolyl Isomerase B PPIB (cyclophilin B) APOB-4314-15-13154 13154 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-3547-15-13155 13155 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-4318-15-13157 13157 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-3741-15-13158 13158 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--13-13159 13159 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--15-13160 13160 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) SOD1-530-16-13163 13163 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-523-16-13164 13164 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-535-16-13165 13165 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-536-16-13166 13166 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-396-16-13167 13167 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-385-16-13168 13168 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) SOD1-195-16-13169 13169 NM_000454 Superoxide Dismutase 1, soluble SOD1 (amyotrophic lateral sclerosis 1 (adult)) pGL3-1172-16-13170 13170 U47296 Cloning vector pGL3-Control pGL3 pGL3-1172-16-13171 13171 U47296 Cloning vector pGL3-Control pGL3 MAP4k4-2931-19-13189 13189 NM_004834 Mitogen-Activated Protein Kinase MAP4k4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 CTGF-1222-13-13190 13190 NM_001901.2 connective tissue growth factor CTGF CTGF-813-13-13192 13192 NM_001901.2 connective tissue growth factor CTGF CTGF-747-13-13194 13194 NM_001901.2 connective tissue growth factor CTGF CTGF-817-13-13196 13196 NM_001901.2 connective tissue growth factor CTGF CTGF-1174-13-13198 13198 NM_001901.2 connective tissue growth factor CTGF CTGF-1005-13-13200 13200 NM_001901.2 connective tissue growth factor CTGF CTGF-814-13-13202 13202 NM_001901.2 connective tissue growth factor CTGF CTGF-816-13-13204 13204 NM_001901.2 connective tissue growth factor CTGF CTGF-1001-13-13206 13206 NM_001901.2 connective tissue growth factor CTGF CTGF-1173-13-13208 13208 NM_001901.2 connective tissue growth factor CTGF CTGF-749-13-13210 13210 NM_001901.2 connective tissue growth factor CTGF CTGF-792-13-13212 13212 NM_001901.2 connective tissue growth factor CTGF CTGF-1162-13-13214 13214 NM_001901.2 connective tissue growth factor CTGF CTGF-811-13-13216 13216 NM_001901.2 connective tissue growth factor CTGF CTGF-797-13-13218 13218 NM_001901.2 connective tissue growth factor CTGF CTGF-1175-13-13220 13220 NM_001901.2 connective tissue growth factor CTGF CTGF-1172-13-13222 13222 NM_001901.2 connective tissue growth factor CTGF CTGF-1177-13-13224 13224 NM_001901.2 connective tissue growth factor CTGF CTGF-1176-13-13226 13226 NM_001901.2 connective tissue growth factor CTGF CTGF-812-13-13228 13228 NM_001901.2 connective tissue growth factor CTGF CTGF-745-13-13230 13230 NM_001901.2 connective tissue growth factor CTGF CTGF-1230-13-13232 13232 NM_001901.2 connective tissue growth factor CTGF CTGF-920-13-13234 13234 NM_001901.2 connective tissue growth factor CTGF
CTGF-679-13-13236 13236 NM_001901.2 connective tissue growth factor CTGF CTGF-992-13-13238 13238 NM_001901.2 connective tissue growth factor CTGF CTGF-1045-13-13240 13240 NM_001901.2 connective tissue growth factor CTGF CTGF-1231-13-13242 13242 NM_001901.2 connective tissue growth factor CTGF CTGF-991-13-13244 13244 NM_001901.2 connective tissue growth factor CTGF CTGF-998-13-13246 13246 NM_001901.2 connective tissue growth factor CTGF CTGF-1049-13-13248 13248 NM_001901.2 connective tissue growth factor CTGF CTGF-1044-13-13250 13250 NM_001901.2 connective tissue growth factor CTGF CTGF-1327-13-13252 13252 NM_001901.2 connective tissue growth factor CTGF CTGF-1196-13-13254 13254 NM_001901.2 connective tissue growth factor CTGF CTGF-562-13-13256 13256 NM_001901.2 connective tissue growth factor CTGF CTGF-752-13-13258 13258 NM_001901.2 connective tissue growth factor CTGF CTGF-994-13-13260 13260 NM_001901.2 connective tissue growth factor CTGF CTGF-1040-13-13262 13262 NM_001901.2 connective tissue growth factor CTGF CTGF-1984-13-13264 13264 NM_001901.2 connective tissue growth factor CTGF CTGF-2195-13-13266 13266 NM_001901.2 connective tissue growth factor CTGF CTGF-2043-13-13268 13268 NM_001901.2 connective tissue growth factor CTGF CTGF-1892-13-13270 13270 NM_001901.2 connective tissue growth factor CTGF CTGF-1567-13-13272 13272 NM_001901.2 connective tissue growth factor CTGF CTGF-1780-13-13274 13274 NM_001901.2 connective tissue growth factor CTGF CTGF-2162-13-13276 13276 NM_001901.2 connective tissue growth factor CTGF CTGF-1034-13-13278 13278 NM_001901.2 connective tissue growth factor CTGF CTGF-2264-13-13280 13280 NM_001901.2 connective tissue growth factor CTGF CTGF-1032-13-13282 13282 NM_001901.2 connective tissue growth factor CTGF CTGF-1535-13-13284 13284 NM_001901.2 connective tissue growth factor CTGF CTGF-1694-13-13286 13286 NM_001901.2 connective tissue growth factor CTGF CTGF-1588-13-13288 13288 NM_001901.2 connective tissue growth factor CTGF CTGF-928-13-13290 13290 NM_001901.2 connective tissue growth factor CTGF CTGF-1133-13-13292 13292 NM_001901.2 connective tissue growth factor CTGF CTGF-912-13-13294 13294 NM_001901.2 connective tissue growth factor CTGF CTGF-753-13-13296 13296 NM_001901.2 connective tissue growth factor CTGF CTGF-918-13-13298 13298 NM_001901.2 connective tissue growth factor CTGF CTGF-744-13-13300 13300 NM_001901.2 connective tissue growth factor CTGF CTGF-466-13-13302 13302 NM_001901.2 connective tissue growth factor CTGF CTGF-917-13-13304 13304 NM_001901.2 connective tissue growth factor CTGF CTGF-1038-13-13306 13306 NM_001901.2 connective tissue growth factor CTGF CTGF-1048-13-13308 13308 NM_001901.2 connective tissue growth factor CTGF CTGF-1235-13-13310 13310 NM_001901.2 connective tissue growth factor CTGF CTGF-868-13-13312 13312 NM_001901.2 connective tissue growth factor CTGF CTGF-1131-13-13314 13314 NM_001901.2 connective tissue growth factor CTGF CTGF-1043-13-13316 13316 NM_001901.2 connective tissue growth factor CTGF CTGF-751-13-13318 13318 NM_001901.2 connective tissue growth factor CTGF CTGF-1227-13-13320 13320 NM_001901.2 connective tissue growth factor CTGF CTGF-867-13-13322 13322 NM_001901.2 connective tissue growth factor CTGF CTGF-1128-13-13324 13324 NM_001901.2 connective tissue growth factor CTGF CTGF-756-13-13326 13326 NM_001901.2 connective tissue growth factor CTGF CTGF-1234-13-13328 13328 NM_001901.2 connective tissue growth factor CTGF CTGF-916-13-13330 13330 NM_001901.2 connective tissue growth factor CTGF CTGF-925-13-13332 13332 NM_001901.2 connective tissue growth factor CTGF CTGF-1225-13-13334 13334 NM_001901.2 connective tissue growth factor CTGF CTGF-445-13-13336 13336 NM_001901.2 connective tissue growth factor CTGF CTGF-446-13-13338 13338 NM_001901.2 connective tissue growth factor CTGF CTGF-913-13-13340 13340 NM_001901.2 connective tissue growth factor CTGF CTGF-997-13-13342 13342 NM_001901.2 connective tissue growth factor CTGF CTGF-277-13-13344 13344 NM_001901.2 connective tissue growth factor CTGF CTGF-1052-13-13346 13346 NM_001901.2 connective tissue growth factor CTGF CTGF-887-13-13348 13348 NM_001901.2 connective tissue growth factor CTGF CTGF-914-13-13350 13350 NM_001901.2 connective tissue growth factor CTGF CTGF-1039-13-13352 13352 NM_001901.2 connective tissue growth factor CTGF CTGF-754-13-13354 13354 NM_001901.2 connective tissue growth factor CTGF CTGF-1130-13-13356 13356 NM_001901.2 connective tissue growth factor CTGF CTGF-919-13-13358 13358 NM_001901.2 connective tissue growth factor CTGF CTGF-922-13-13360 13360 NM_001901.2 connective tissue growth factor CTGF CTGF-746-13-13362 13362 NM_001901.2 connective tissue growth factor CTGF CTGF-993-13-13364 13364 NM_001901.2 connective tissue growth factor CTGF CTGF-825-13-13366 13366 NM_001901.2 connective tissue growth factor CTGF CTGF-926-13-13368 13368 NM_001901.2 connective tissue growth factor CTGF CTGF-923-13-13370 13370 NM_001901.2 connective tissue growth factor CTGF CTGF-866-13-13372 13372 NM_001901.2 connective tissue growth factor CTGF CTGF-563-13-13374 13374 NM_001901.2 connective tissue growth factor CTGF CTGF-823-13-13376 13376 NM_001901.2 connective tissue growth factor CTGF CTGF-1233-13-13378 13378 NM_001901.2 connective tissue growth factor CTGF CTGF-924-13-13380 13380 NM_001901.2 connective tissue growth factor CTGF CTGF-921-13-13382 13382 NM_001901.2 connective tissue growth factor CTGF CTGF-443-13-13384 13384 NM_001901.2 connective tissue growth factor CTGF CTGF-1041-13-13386 13386 NM_001901.2 connective tissue growth factor CTGF CTGF-1042-13-13388 13388 NM_001901.2 connective tissue growth factor CTGF CTGF-755-13-13390 13390 NM_001901.2 connective tissue growth factor CTGF CTGF-467-13-13392 13392 NM_001901.2 connective tissue growth factor CTGF CTGF-995-13-13394 13394 NM_001901.2 connective tissue growth factor CTGF CTGF-927-13-13396 13396 NM_001901.2 connective tissue growth factor CTGF SPP1-1025-13-13398 13398 NM_000582.2 Osteopontin SPP1 SPP1-1049-13-13400 13400 NM_000582.2 Osteopontin SPP1 SPP1-1051-13-13402 13402 NM_000582.2 Osteopontin SPP1 SPP1-1048-13-13404 13404 NM_000582.2 Osteopontin SPP1 SPP1-1050-13-13406 13406 NM_000582.2 Osteopontin SPP1 SPP1-1047-13-13408 13408 NM_000582.2 Osteopontin SPP1 SPP1-800-13-13410 13410 NM_000582.2 Osteopontin SPP1 SPP1-492-13-13412 13412 NM_000582.2 Osteopontin SPP1 SPP1-612-13-13414 13414 NM_000582.2 Osteopontin SPP1 SPP1-481-13-13416 13416 NM_000582.2 Osteopontin SPP1 SPP1-614-13-13418 13418 NM_000582.2 Osteopontin SPP1 SPP1-951-13-13420 13420 NM_000582.2 Osteopontin SPP1 SPP1-482-13-13422 13422 NM_000582.2 Osteopontin SPP1 SPP1-856-13-13424 13424 NM_000582.2 Osteopontin SPP1 SPP1-857-13-13426 13426 NM_000582.2 Osteopontin SPP1 SPP1-365-13-13428 13428 NM_000582.2 Osteopontin SPP1 SPP1-359-13-13430 13430 NM_000582.2 Osteopontin SPP1 SPP1-357-13-13432 13432 NM_000582.2 Osteopontin SPP1 SPP1-858-13-13434 13434 NM_000582.2 Osteopontin SPP1 SPP1-1012-13-13436 13436 NM_000582.2 Osteopontin SPP1 SPP1-1014-13-13438 13438 NM_000582.2 Osteopontin SPP1 SPP1-356-13-13440 13440 NM_000582.2 Osteopontin SPP1 SPP1-368-13-13442 13442 NM_000582.2 Osteopontin SPP1 SPP1-1011-13-13444 13444 NM_000582.2 Osteopontin SPP1 SPP1-754-13-13446 13446 NM_000582.2 Osteopontin SPP1 SPP1-1021-13-13448 13448 NM_000582.2 Osteopontin SPP1 SPP1-1330-13-13450 13450 NM_000582.2 Osteopontin SPP1 SPP1-346-13-13452 13452 NM_000582.2 Osteopontin SPP1 SPP1-869-13-13454 13454 NM_000582.2 Osteopontin SPP1 SPP1-701-13-13456 13456 NM_000582.2 Osteopontin SPP1 SPP1-896-13-13458 13458 NM_000582.2 Osteopontin SPP1 SPP1-1035-13-13460 13460 NM_000582.2 Osteopontin SPP1 SPP1-1170-13-13462 13462 NM_000582.2 Osteopontin SPP1 SPP1-1282-13-13464 13464 NM_000582.2 Osteopontin SPP1 SPP1-1537-13-13466 13466 NM_000582.2 Osteopontin SPP1 SPP1-692-13-13468 13468 NM_000582.2 Osteopontin SPP1 SPP1-840-13-13470 13470 NM_000582.2 Osteopontin SPP1 SPP1-1163-13-13472 13472 NM_000582.2 Osteopontin SPP1 SPP1-789-13-13474 13474 NM_000582.2 Osteopontin SPP1 SPP1-841-13-13476 13476 NM_000582.2 Osteopontin SPP1 SPP1-852-13-13478 13478 NM_000582.2 Osteopontin SPP1 SPP1-209-13-13480 13480 NM_000582.2 Osteopontin SPP1 SPP1-1276-13-13482 13482 NM_000582.2 Osteopontin SPP1 SPP1-137-13-13484 13484 NM_000582.2 Osteopontin SPP1 SPP1-711-13-13486 13486 NM_000582.2 Osteopontin SPP1 SPP1-582-13-13488 13488 NM_000582.2 Osteopontin SPP1 SPP1-839-13-13490 13490 NM_000582.2 Osteopontin SPP1 SPP1-1091-13-13492 13492 NM_000582.2 Osteopontin SPP1 SPP1-884-13-13494 13494 NM_000582.2 Osteopontin SPP1 SPP1-903-13-13496 13496 NM_000582.2 Osteopontin SPP1 SPP1-1090-13-13498 13498 NM_000582.2 Osteopontin SPP1 SPP1-474-13-13500 13500 NM_000582.2 Osteopontin SPP1 SPP1-575-13-13502 13502 NM_000582.2 Osteopontin SPP1 SPP1-671-13-13504 13504 NM_000582.2 Osteopontin SPP1 SPP1-924-13-13506 13506 NM_000582.2 Osteopontin SPP1 SPP1-1185-13-13508 13508 NM_000582.2 Osteopontin SPP1 SPP1-1221-13-13510 13510 NM_000582.2 Osteopontin SPP1 SPP1-347-13-13512 13512 NM_000582.2 Osteopontin SPP1 SPP1-634-13-13514 13514 NM_000582.2 Osteopontin SPP1 SPP1-877-13-13516 13516 NM_000582.2 Osteopontin SPP1 SPP1-1033-13-13518 13518 NM_000582.2 Osteopontin SPP1 SPP1-714-13-13520 13520 NM_000582.2 Osteopontin SPP1 SPP1-791-13-13522 13522 NM_000582.2 Osteopontin SPP1 SPP1-813-13-13524 13524 NM_000582.2 Osteopontin SPP1 SPP1-939-13-13526 13526 NM_000582.2 Osteopontin SPP1 SPP1-1161-13-13528 13528 NM_000582.2 Osteopontin SPP1 SPP1-1164-13-13530 13530 NM_000582.2 Osteopontin SPP1 SPP1-1190-13-13532 13532 NM_000582.2 Osteopontin SPP1 SPP1-1333-13-13534 13534 NM_000582.2 Osteopontin SPP1 SPP1-537-13-13536 13536 NM_000582.2 Osteopontin SPP1 SPP1-684-13-13538 13538 NM_000582.2 Osteopontin SPP1 SPP1-707-13-13540 13540 NM_000582.2 Osteopontin SPP1 SPP1-799-13-13542 13542 NM_000582.2 Osteopontin SPP1 SPP1-853-13-13544 13544 NM_000582.2 Osteopontin SPP1 SPP1-888-13-13546 13546 NM_000582.2 Osteopontin SPP1 SPP1-1194-13-13548 13548 NM_000582.2 Osteopontin SPP1 SPP1-1279-13-13550 13550 NM_000582.2 Osteopontin SPP1 SPP1-1300-13-13552 13552 NM_000582.2 Osteopontin SPP1 SPP1-1510-13-13554 13554 NM_000582.2 Osteopontin SPP1 SPP1-1543-13-13556 13556 NM_000582.2 Osteopontin SPP1 SPP1-434-13-13558 13558 NM_000582.2 Osteopontin SPP1 SPP1-600-13-13560 13560 NM_000582.2 Osteopontin SPP1 SPP1-863-13-13562 13562 NM_000582.2 Osteopontin SPP1 SPP1-902-13-13564 13564 NM_000582.2 Osteopontin SPP1 SPP1-921-13-13566 13566 NM_000582.2 Osteopontin SPP1 SPP1-154-13-13568 13568 NM_000582.2 Osteopontin SPP1 SPP1-217-13-13570 13570 NM_000582.2 Osteopontin SPP1 SPP1-816-13-13572 13572 NM_000582.2 Osteopontin SPP1 SPP1-882-13-13574 13574 NM_000582.2 Osteopontin SPP1 SPP1-932-13-13576 13576 NM_000582.2 Osteopontin SPP1 SPP1-1509-13-13578 13578 NM_000582.2 Osteopontin SPP1 SPP1-157-13-13580 13580 NM_000582.2 Osteopontin SPP1 SPP1-350-13-13582 13582 NM_000582.2 Osteopontin SPP1 SPP1-511-13-13584 13584 NM_000582.2 Osteopontin SPP1 SPP1-605-13-13586 13586 NM_000582.2 Osteopontin SPP1 SPP1-811-13-13588 13588 NM_000582.2 Osteopontin SPP1 SPP1-892-13-13590 13590 NM_000582.2 Osteopontin SPP1 SPP1-922-13-13592 13592 NM_000582.2 Osteopontin SPP1 SPP1-1169-13-13594 13594 NM_000582.2 Osteopontin SPP1 SPP1-1182-13-13596 13596 NM_000582.2 Osteopontin SPP1 SPP1-1539-13-13598 13598 NM_000582.2 Osteopontin SPP1 SPP1-1541-13-13600 13600 NM_000582.2 Osteopontin SPP1 SPP1-427-13-13602 13602 NM_000582.2 Osteopontin SPP1 SPP1-533-13-13604 13604 NM_000582.2 Osteopontin SPP1 APOB--13-13763 13763 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--13-13764 13764 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4--16-13766 13766 MAP4K4 PPIB--13-13767 13767 NM_000942 peptidylprolyl isomerase B PPIB (cyclophilin B) PPIB--15-13768 13768 NM_000942 peptidylprolyl isomerase B PPIB (cyclophilin B) PPIB--17-13769 13769 NM_000942 peptidylprolyl isomerase B PPIB (cyclophilin B) MAP4K4--16-13939 13939 MAP4K4 APOB-4314-16-13940 13940 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-4314-17-13941 13941 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--16-13942 13942 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--18-13943 13943 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--17-13944 13944 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--19-13945 13945 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-4314-16-13946 13946 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB-4314-17-13947 13947 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--16-13948 13948 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--17-13949 13949 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--16-13950 13950 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--18-13951 13951 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--17-13952 13952 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) APOB--19-13953 13953 NM_000384 Apolipoprotein B (including APOB Ag(x) antigen) MAP4K4--16-13766.2 13766.2 MAP4K4 CTGF-1222-16-13980 13980 NM_001901.2 connective tissue growth factor CTGF CTGF-813-16-13981 13981 NM_001901.2 connective tissue growth factor CTGF CTGF-747-16-13982 13982 NM_001901.2 connective tissue growth factor CTGF CTGF-817-16-13983 13983 NM_001901.2 connective tissue growth factor CTGF CTGF-1174-16-13984 13984 NM_001901.2 connective tissue growth factor CTGF CTGF-1005-16-13985 13985 NM_001901.2 connective tissue growth factor CTGF CTGF-814-16-13986 13986 NM_001901.2 connective tissue growth factor CTGF CTGF-816-16-13987 13987 NM_001901.2 connective tissue growth factor CTGF CTGF-1001-16-13988 13988 NM_001901.2 connective tissue growth factor CTGF CTGF-1173-16-13989 13989 NM_001901.2 connective tissue growth factor CTGF CTGF-749-16-13990 13990 NM_001901.2 connective tissue growth factor CTGF CTGF-792-16-13991 13991 NM_001901.2 connective tissue growth factor CTGF CTGF-1162-16-13992 13992 NM_001901.2 connective tissue growth factor CTGF CTGF-811-16-13993 13993 NM_001901.2 connective tissue growth factor CTGF CTGF-797-16-13994 13994 NM_001901.2 connective tissue growth factor CTGF CTGF-1175-16-13995 13995 NM_001901.2 connective tissue growth factor CTGF CTGF-1172-16-13996 13996 NM_001901.2 connective tissue growth factor CTGF CTGF-1177-16-13997 13997 NM_001901.2 connective tissue growth factor CTGF CTGF-1176-16-13998 13998 NM_001901.2 connective tissue growth factor CTGF CTGF-812-16-13999 13999 NM_001901.2 connective tissue growth factor CTGF CTGF-745-16-14000 14000 NM_001901.2 connective tissue growth factor CTGF CTGF-1230-16-14001 14001 NM_001901.2 connective tissue growth factor CTGF CTGF-920-16-14002 14002 NM_001901.2 connective tissue growth factor CTGF CTGF-679-16-14003 14003 NM_001901.2 connective tissue growth factor CTGF CTGF-992-16-14004 14004 NM_001901.2 connective tissue growth factor CTGF
CTGF-1045-16-14005 14005 NM_001901.2 connective tissue growth factor CTGF CTGF-1231-16-14006 14006 NM_001901.2 connective tissue growth factor CTGF CTGF-991-16-14007 14007 NM_001901.2 connective tissue growth factor CTGF CTGF-998-16-14008 14008 NM_001901.2 connective tissue growth factor CTGF CTGF-1049-16-14009 14009 NM_001901.2 connective tissue growth factor CTGF CTGF-1044-16-14010 14010 NM_001901.2 connective tissue growth factor CTGF CTGF-1327-16-14011 14011 NM_001901.2 connective tissue growth factor CTGF CTGF-1196-16-14012 14012 NM_001901.2 connective tissue growth factor CTGF CTGF-562-16-14013 14013 NM_001901.2 connective tissue growth factor CTGF CTGF-752-16-14014 14014 NM_001901.2 connective tissue growth factor CTGF CTGF-994-16-14015 14015 NM_001901.2 connective tissue growth factor CTGF CTGF-1040-16-14016 14016 NM_001901.2 connective tissue growth factor CTGF CTGF-1984-16-14017 14017 NM_001901.2 connective tissue growth factor CTGF CTGF-2195-16-14018 14018 NM_001901.2 connective tissue growth factor CTGF CTGF-2043-16-14019 14019 NM_001901.2 connective tissue growth factor CTGF CTGF-1892-16-14020 14020 NM_001901.2 connective tissue growth factor CTGF CTGF-1567-16-14021 14021 NM_001901.2 connective tissue growth factor CTGF CTGF-1780-16-14022 14022 NM_001901.2 connective tissue growth factor CTGF CTGF-2162-16-14023 14023 NM_001901.2 connective tissue growth factor CTGF CTGF-1034-16-14024 14024 NM_001901.2 connective tissue growth factor CTGF CTGF-2264-16-14025 14025 NM_001901.2 connective tissue growth factor CTGF CTGF-1032-16-14026 14026 NM_001901.2 connective tissue growth factor CTGF CTGF-1535-16-14027 14027 NM_001901.2 connective tissue growth factor CTGF CTGF-1694-16-14028 14028 NM_001901.2 connective tissue growth factor CTGF CTGF-1588-16-14029 14029 NM_001901.2 connective tissue growth factor CTGF CTGF-928-16-14030 14030 NM_001901.2 connective tissue growth factor CTGF CTGF-1133-16-14031 14031 NM_001901.2 connective tissue growth factor CTGF CTGF-912-16-14032 14032 NM_001901.2 connective tissue growth factor CTGF CTGF-753-16-14033 14033 NM_001901.2 connective tissue growth factor CTGF CTGF-918-16-14034 14034 NM_001901.2 connective tissue growth factor CTGF CTGF-744-16-14035 14035 NM_001901.2 connective tissue growth factor CTGF CTGF-466-16-14036 14036 NM_001901.2 connective tissue growth factor CTGF CTGF-917-16-14037 14037 NM_001901.2 connective tissue growth factor CTGF CTGF-1038-16-14038 14038 NM_001901.2 connective tissue growth factor CTGF CTGF-1048-16-14039 14039 NM_001901.2 connective tissue growth factor CTGF CTGF-1235-16-14040 14040 NM_001901.2 connective tissue growth factor CTGF CTGF-868-16-14041 14041 NM_001901.2 connective tissue growth factor CTGF CTGF-1131-16-14042 14042 NM_001901.2 connective tissue growth factor CTGF CTGF-1043-16-14043 14043 NM_001901.2 connective tissue growth factor CTGF CTGF-751-16-14044 14044 NM_001901.2 connective tissue growth factor CTGF CTGF-1227-16-14045 14045 NM_001901.2 connective tissue growth factor CTGF CTGF-867-16-14046 14046 NM_001901.2 connective tissue growth factor CTGF CTGF-1128-16-14047 14047 NM_001901.2 connective tissue growth factor CTGF CTGF-756-16-14048 14048 NM_001901.2 connective tissue growth factor CTGF CTGF-1234-16-14049 14049 NM_001901.2 connective tissue growth factor CTGF CTGF-916-16-14050 14050 NM_001901.2 connective tissue growth factor CTGF CTGF-925-16-14051 14051 NM_001901.2 connective tissue growth factor CTGF CTGF-1225-16-14052 14052 NM_001901.2 connective tissue growth factor CTGF CTGF-445-16-14053 14053 NM_001901.2 connective tissue growth factor CTGF CTGF-446-16-14054 14054 NM_001901.2 connective tissue growth factor CTGF CTGF-913-16-14055 14055 NM_001901.2 connective tissue growth factor CTGF CTGF-997-16-14056 14056 NM_001901.2 connective tissue growth factor CTGF CTGF-277-16-14057 14057 NM_001901.2 connective tissue growth factor CTGF CTGF-1052-16-14058 14058 NM_001901.2 connective tissue growth factor CTGF CTGF-887-16-14059 14059 NM_001901.2 connective tissue growth factor CTGF CTGF-914-16-14060 14060 NM_001901.2 connective tissue growth factor CTGF CTGF-1039-16-14061 14061 NM_001901.2 connective tissue growth factor CTGF CTGF-754-16-14062 14062 NM_001901.2 connective tissue growth factor CTGF CTGF-1130-16-14063 14063 NM_001901.2 connective tissue growth factor CTGF CTGF-919-16-14064 14064 NM_001901.2 connective tissue growth factor CTGF CTGF-922-16-14065 14065 NM_001901.2 connective tissue growth factor CTGF CTGF-746-16-14066 14066 NM_001901.2 connective tissue growth factor CTGF CTGF-993-16-14067 14067 NM_001901.2 connective tissue growth factor CTGF CTGF-825-16-14068 14068 NM_001901.2 connective tissue growth factor CTGF CTGF-926-16-14069 14069 NM_001901.2 connective tissue growth factor CTGF CTGF-923-16-14070 14070 NM_001901.2 connective tissue growth factor CTGF CTGF-866-16-14071 14071 NM_001901.2 connective tissue growth factor CTGF CTGF-563-16-14072 14072 NM_001901.2 connective tissue growth factor CTGF CTGF-823-16-14073 14073 NM_001901.2 connective tissue growth factor CTGF CTGF-1233-16-14074 14074 NM_001901.2 connective tissue growth factor CTGF CTGF-924-16-14075 14075 NM_001901.2 connective tissue growth factor CTGF CTGF-921-16-14076 14076 NM_001901.2 connective tissue growth factor CTGF CTGF-443-16-14077 14077 NM_001901.2 connective tissue growth factor CTGF CTGF-1041-16-14078 14078 NM_001901.2 connective tissue growth factor CTGF CTGF-1042-16-14079 14079 NM_001901.2 connective tissue growth factor CTGF CTGF-755-16-14080 14080 NM_001901.2 connective tissue growth factor CTGF CTGF-467-16-14081 14081 NM_001901.2 connective tissue growth factor CTGF CTGF-995-16-14082 14082 NM_001901.2 connective tissue growth factor CTGF CTGF-927-16-14083 14083 NM_001901.2 connective tissue growth factor CTGF SPP1-1091-16-14131 14131 NM_000582.2 Osteopontin SPP1 PPIB--16-14188 14188 NM_000942 peptidylprolyl isomerase B PPIB (cyclophilin B) PPIB--17-14189 14189 NM_000942 peptidylprolyl isomerase B PPIB (cyclophilin B) PPIB--18-14190 14190 NM_000942 peptidylprolyl isomerase B PPIB (cyclophilin B) pGL3-1172-16-14386 14386 U47296 Cloning vector pGL3-Control pGL3 pGL3-1172-16-14387 14387 U47296 Cloning vector pGL3-Control pGL3 MAP4K4-2931-25-14390 14390 NM_004834 Mitogen-Activated Protein Kinase MAP4K4 Kinase Kinase Kinase 4 (MAP4K4), transcript variant 1 miR-122--23-14391 14391 miR-122 14084 NM_000582.2 Osteopontin SPP1 14085 NM_000582.2 Osteopontin SPP1 14086 NM_000582.2 Osteopontin SPP1 14087 NM_000582.2 Osteopontin SPP1 14088 NM_000582.2 Osteopontin SPP1 14089 NM_000582.2 Osteopontin SPP1 14090 NM_000582.2 Osteopontin SPP1 14091 NM_000582.2 Osteopontin SPP1 14092 NM_000582.2 Osteopontin SPP1 14093 NM_000582.2 Osteopontin SPP1 14094 NM_000582.2 Osteopontin SPP1 14095 NM_000582.2 Osteopontin SPP1 14096 NM_000582.2 Osteopontin SPP1 14097 NM_000582.2 Osteopontin SPP1 14098 NM_000582.2 Osteopontin SPP1 14099 NM_000582.2 Osteopontin SPP1 14100 NM_000582.2 Osteopontin SPP1 14101 NM_000582.2 Osteopontin SPP1 14102 NM_000582.2 Osteopontin SPP1 14103 NM_000582.2 Osteopontin SPP1 14104 NM_000582.2 Osteopontin SPP1 14105 NM_000582.2 Osteopontin SPP1 14106 NM_000582.2 Osteopontin SPP1 14107 NM_000582.2 Osteopontin SPP1 14108 NM_000582.2 Osteopontin SPP1 14109 NM_000582.2 Osteopontin SPP1 14110 NM_000582.2 Osteopontin SPP1 14111 NM_000582.2 Osteopontin SPP1 14112 NM_000582.2 Osteopontin SPP1 14113 NM_000582.2 Osteopontin SPP1 14114 NM_000582.2 Osteopontin SPP1 14115 NM_000582.2 Osteopontin SPP1 14116 NM_000582.2 Osteopontin SPP1 14117 NM_000582.2 Osteopontin SPP1 14118 NM_000582.2 Osteopontin SPP1 14119 NM_000582.2 Osteopontin SPP1 14120 NM_000582.2 Osteopontin SPP1 14121 NM_000582.2 Osteopontin SPP1 14122 NM_000582.2 Osteopontin SPP1 14123 NM_000582.2 Osteopontin SPP1 14124 NM_000582.2 Osteopontin SPP1 14125 NM_000582.2 Osteopontin SPP1 14126 NM_000582.2 Osteopontin SPP1 14127 NM_000582.2 Osteopontin SPP1 14128 NM_000582.2 Osteopontin SPP1 14129 NM_000582.2 Osteopontin SPP1 14130 NM_000582.2 Osteopontin SPP1 14132 NM_000582.2 Osteopontin SPP1 14133 NM_000582.2 Osteopontin SPP1 14134 NM_000582.2 Osteopontin SPP1 14135 NM_000582.2 Osteopontin SPP1 14136 NM_000582.2 Osteopontin SPP1 14137 NM_000582.2 Osteopontin SPP1 14138 NM_000582.2 Osteopontin SPP1 14139 NM_000582.2 Osteopontin SPP1 14140 NM_000582.2 Osteopontin SPP1 14141 NM_000582.2 Osteopontin SPP1 14142 NM_000582.2 Osteopontin SPP1 14143 NM_000582.2 Osteopontin SPP1 14144 NM_000582.2 Osteopontin SPP1 14145 NM_000582.2 Osteopontin SPP1 14146 NM_000582.2 Osteopontin SPP1 14147 NM_000582.2 Osteopontin SPP1 14148 NM_000582.2 Osteopontin SPP1 14149 NM_000582.2 Osteopontin SPP1 14150 NM_000582.2 Osteopontin SPP1 14151 NM_000582.2 Osteopontin SPP1 14152 NM_000582.2 Osteopontin SPP1 14153 NM_000582.2 Osteopontin SPP1 14154 NM_000582.2 Osteopontin SPP1 14155 NM_000582.2 Osteopontin SPP1 14156 NM_000582.2 Osteopontin SPP1 14157 NM_000582.2 Osteopontin SPP1 14158 NM_000582.2 Osteopontin SPP1 14159 NM_000582.2 Osteopontin SPP1 14160 NM_000582.2 Osteopontin SPP1 14161 NM_000582.2 Osteopontin SPP1 14162 NM_000582.2 Osteopontin SPP1 14163 NM_000582.2 Osteopontin SPP1 14164 NM_000582.2 Osteopontin SPP1 14165 NM_000582.2 Osteopontin SPP1 14166 NM_000582.2 Osteopontin SPP1 14167 NM_000582.2 Osteopontin SPP1 14168 NM_000582.2 Osteopontin SPP1 14169 NM_000582.2 Osteopontin SPP1 14170 NM_000582.2 Osteopontin SPP1 14171 NM_000582.2 Osteopontin SPP1 14172 NM_000582.2 Osteopontin SPP1 14173 NM_000582.2 Osteopontin SPP1 14174 NM_000582.2 Osteopontin SPP1 14175 NM_000582.2 Osteopontin SPP1 14176 NM_000582.2 Osteopontin SPP1 14177 NM_000582.2 Osteopontin SPP1 14178 NM_000582.2 Osteopontin SPP1 14179 NM_000582.2 Osteopontin SPP1 14180 NM_000582.2 Osteopontin SPP1 14181 NM_000582.2 Osteopontin SPP1 14182 NM_000582.2 Osteopontin SPP1 14183 NM_000582.2 Osteopontin SPP1 14184 NM_000582.2 Osteopontin SPP1 14185 NM_000582.2 Osteopontin SPP1 14186 NM_000582.2 Osteopontin SPP1 14187 NM_000582.2 Osteopontin SPP1
TABLE-US-00002 TABLE 2 Antisense backbone, chemistry, and sequence information. ID Oligo AntiSense AntiSense AntiSense SEQ ID Number Number Backbone Chemistry Sequence NO: APOB-10167-20-12138 12138 ooooooooooooo 00000000000000 AUUGGUAUUCAGUGUGA 1 oooooo 000000m UG APOB-10167-20-12139 12139 ooooooooooooo 00000000000000 AUUCGUAUUGAGUCUGA 2 oooooo 000000m UC MAP4K4-2931-13-12266 12266 MAP4K4-2931-16-12293 12293 ooooooooooooo Pf000fffff0f00 UAGACUUCCACAGAACU 3 oooooo 00fff0 CU MAP4K4-2931-16-12383 12383 ooooooooooooo 00000000000000 UAGACUUCCACAGAACU 4 oooooo 00000 CU MAP4K4-2931-16-12384 12384 ooooooooooooo P0000000000000 UAGACUUCCACAGAACU 5 oooooo 000000 CU MAP4K4-2931-16-12385 12385 ooooooooooooo Pf000fffff0f00 UAGACUUCCACAGAACU 6 oooooo 00fff0 CU MAP4K4-2931-16-12386 12386 oooooooooosss Pf000fffff0f00 UAGACUUCCACAGAACU 7 ssssso 00fff0 CU MAP4K4-2931-16-12387 12387 oooooooooosss P0000000000000 UAGACUUCCACAGAACU 8 ssssso 000000 CU MAP4K4-2931-15-12388 12388 ooooooooooooo 00000000000000 UAGACUUCCACAGAACU 9 oooo 000 MAP4K4-2931-13-12432 12432 MAP4K4-2931-13-12266.2 12266.2 APOB--21-12434 12434 ooooooooooooo 00000000000000 AUUGGUAUUCAGUGUGA 10 oooooooo 000000m UGAC APOB--21-12435 12435 ooooooooooooo 00000000000000 AUUCGUAUUGAGUCUGA 11 oooooooo 000000m UCAC MAP4K4-2931-16-12451 12451 oooooooooosss Pf000fffff0f00 UAGACUUCCACAGAACU 12 ssssso 00ffmm CU MAP4K4-2931-16-12452 12452 oooooooooosss Pm000fffff0f00 UAGACUUCCACAGAACU 13 ssssso 00ffmm CU MAP4K4-2931-16-12453 12453 oooooosssssss Pm000fffff0f00 UAGACUUCCACAGAACU 14 ssssso 00ffmm CU MAP4K4-2931-17-12454 12454 oooooooooooos Pm000fffff0f00 UAGACUUCCACAGAACU 15 ssssssso 00ffffmm CUUC MAP4K4-2931-17-12455 12455 oooooooosssss Pm000fffff0f00 UAGACUUCCACAGAACU 16 ssssssso 00ffffmm CUUC MAP4K4-2931-19-12456 12456 oooooooooooos Pm000fffff0f00 UAGACUUCCACAGAACU 17 ssssssssssso 00ffffff00mm CUUCAAAG --27-12480 12480 --27-12481 12481 APOB-10167-21-12505 12505 ooooooooooooo 00000000000000 AUUGGUAUUCAGUGUGA 18 ooooooos 000000m UGAC APOB-10167-21-12506 12506 ooooooooooooo 00000000000000 AUUCGUAUUGAGUCUGA 19 ooooooos 000000m UCAC MAP4K4-2931-16-12539 12539 oooooooooooss Pf000fffff0f00 UAGACUUCCACAGAACU 20 ssssss 00fff0 CU APOB-10167-21-12505.2 12505.2 ooooooooooooo 00000000000000 AUUGGUAUUCAGUGUGA 21 oooooooo 000000m UGAC APOB-10167-21-12506.2 12506.2 ooooooooooooo 00000000000000 AUUCGUAUUGAGUCUGA 22 oooooooo 000000m UCAC MAP4K4--13-12565 12565 MAP4K4-2931-16-12386.2 12386.2 oooooooooosss Pf000fffff0f00 UAGACUUCCACAGAACU 23 ssssso 00fff0 CU MAP4K4-2931-13-12815 12815 APOB--13-12957 12957 MAP4K4--16-12983 12983 oooooooooooos Pm000fffff0m00 uagacuuccacagaacu 24 ssssso 00mmm0 cu MAP4K4--16-12984 12984 oooooooooooos Pm000fffff0m00 uagacuuccacagaacu 25 sssss 00mmm0 cu MAP4K4--16-12985 12985 oooooooooooos Pm000fffff0m00 uagacuuccacagaacu 26 ssssso 00mmm0 cu MAP4K4--16-12986 12986 oooooooooosss Pf000fffff0f00 UAGACUUCCACAGAACU 27 ssssso 00fff0 CU MAP4K4--16-12987 12987 ooooooooooooo P0000f00ff0m00 UagacUUccacagaacU 28 ssssss 00m0m0 cU MAP4K4--16-12988 12988 ooooooooooooo P0000f00ff0m00 UagacUUccacagaacU 29 ssssss 00m0m0 cu MAP4K4--16-12989 12989 ooooooooooooo P0000ff0ff0m00 UagacuUccacagaacU 30 ssssss 00m0m0 cu MAP4K4--16-12990 12990 ooooooooooooo Pf0000ff000000 uagaCuuCCaCagaaCu 31 ssssss 000m00 Cu MAP4K4--16-12991 12991 ooooooooooooo Pf0000fff00m00 uagaCuucCacagaaCu 32 ssssss 000mm0 cu MAP4K4--16-12992 12992 ooooooooooooo Pf000fffff0000 uagacuuccaCagaaCu 33 ssssss 000m00 Cu MAP4K4--16-12993 12993 ooooooooooooo P0000000000000 UagaCUUCCaCagaaCU 34 ssssss 000000 CU MAP4K4--16-12994 12994 ooooooooooooo P0000f0f0f0000 UagacUuCcaCagaaCu 35 ssssss 000m00 Cu MAP4K4--16-12995 12995 oooooooooooos Pf000fffff0000 uagacuuccaCagaaCu 36 ssssso 000000 CU MAP4K4-2931-19-13012 13012 MAP4K4-2931-19-13016 13016 PPIB--13-13021 13021 pGL3-1172-13-13038 13038 pGL3-1172-13-13040 13040 --16-13047 13047 oooooooooooos Pm000000000m00 UAGACUUCCACAGAACU 37 sssss 00mmm0 CU SOD1-530-13-13090 13090 SOD1-523-13-13091 13091 SOD1-535-13-13092 13092 SOD1-536-13-13093 13093 SOD1-396-13-13094 13094 SOD1-385-13-13095 13095 SOD1-195-13-13096 13096 APOB-4314-13-13115 13115 APOB-3384-13-13116 13116 APOB-3547-13-13117 13117 APOB-4318-13-13118 13118 APOB-3741-13-13119 13119 PPIB--16-13136 13136 oooooooooooos Pm0fffff0f00mm UGUUUUUGUAGCCAAAU 38 sssss 000mm0 CC APOB-4314-15-13154 13154 APOB-3547-15-13155 13155 APOB-4318-15-13157 13157 APOB-3741-15-13158 13158 APOB--13-13159 13159 APOB--15-13160 13160 SOD1-530-16-13163 13163 oooooooooooos Pm0ffffffff0mm UACUUUCUUCAUUUCCA 39 ssssso mmm0m0 CC SOD1-523-16-13164 13164 oooooooooooos Pmff0fffff0fmm UUCAUUUCCACCUUUGC 40 ssssso mm0mm0 CC SOD1-535-16-13165 13165 oooooooooooos Pmfff0f0ffffmm CUUUGUACUUUCUUCAU 41 ssssso mm0mm0 UU SOD1-536-16-13166 13166 oooooooooooos Pmffff0f0fffmm UCUUUGUACUUUCUUCA 42 ssssso mmm0m0 UU SOD1-396-16-13167 13167 oooooooooooos Pmf00f00ff0f0m UCAGCAGUCACAUUGCC 43 ssssso m0mmm0 CA SOD1-385-16-13168 13168 oooooooooooos Pmff0fff000fmm AUUGCCCAAGUCUCCAA 44 ssssso mm00m0 CA SOD1-195-16-13169 13169 oooooooooooos Pmfff0fff0000m UUCUGCUCGAAAUUGAU 45 ssssso m00m00 GA pGL3-1172-16-13170 13170 oooooooooooos Pm00ff0f0ffm0f AAAUCGUAUUUGUCAAU 46 ssssso f00mm0 CA pGL3-l172-16-13171 13171 ooooooooooooo Pm00ff0f0ffm0f AAAUCGUAUUUGUCAAU 47 ssssss f00mm0 CA MAP4k4-2931-19-13189 13189 ooooooooooooo 00000000000000 UAGACUUCCACAGAACU 48 oooooo 00000 CU CTGF-1222-13-13190 13190 CTGF-813-13-13192 13192 CTGF-747-13-13194 13194 CTGF-817-13-13196 13196
CTGF-1174-13-13198 13198 CTGF-1005-13-13200 13200 CTGF-814-13-13202 13202 CTGF-816-13-13204 13204 CTGF-1001-13-13206 13206 CTGF-1173-13-13208 13208 CTGF-749-13-13210 13210 CTGF-792-13-13212 13212 CTGF-1162-13-13214 13214 CTGF-811-13-13216 13216 CTGF-797-13-13218 13218 CTGF-1175-13-13220 13220 CTGF-1172-13-13222 13222 CTGF-1177-13-13224 13224 CTGF-1176-13-13226 13226 CTGF-812-13-13228 13228 CTGF-745-13-13230 13230 CTGF-1230-13-13232 13232 CTGF-920-13-13234 13234 CTGF-679-13-13236 13236 CTGF-992-13-13238 13238 CTGF-1045-13-13240 13240 CTGF-1231-13-13242 13242 CTGF-991-13-13244 13244 CTGF-998-13-13246 13246 CTGF-1049-13-13248 13248 CTGF-1044-13-13250 13250 CTGF-1327-13-13252 13252 CTGF-1196-13-13254 13254 CTGF-562-13-13256 13256 CTGF-752-13-13258 13258 CTGF-994-13-13260 13260 CTGF-1040-13-13262 13262 CTGF-1984-13-13264 13264 CTGF-2195-13-13266 13266 CTGF-2043-13-13268 13268 CTGF-1892-13-13270 13270 CTGF-1567-13-13272 13272 CTGF-1780-13-13274 13274 CTGF-2162-13-13276 13276 CTGF-1034-13-13278 13278 CTGF-2264-13-13280 13280 CTGF-1032-13-13282 13282 CTGF-1535-13-13284 13284 CTGF-1694-13-13286 13286 CTGF-1588-13-13288 13288 CTGF-928-13-13290 13290 CTGF-1133-13-13292 13292 CTGF-912-13-13294 13294 CTGF-753-13-13296 13296 CTGF-918-13-13298 13298 CTGF-744-13-13300 13300 CTGF-466-13-13302 13302 CTGF-917-13-13304 13304 CTGF-1038-13-13306 13306 CTGF-1048-13-13308 13308 CTGF-1235-13-13310 13310 CTGF-868-13-13312 13312 CTGF-1131-13-13314 13314 CTGF-1043-13-13316 13316 CTGF-751-13-13318 13318 CTGF-1227-13-13320 13320 CTGF-867-13-13322 13322 CTGF-1128-13-13324 13324 CTGF-756-13-13326 13326 CTGF-1234-13-13328 13328 CTGF-916-13-13330 13330 CTGF-925-13-13332 13332 CTGF-1225-13-13334 13334 CTGF-445-13-13336 13336 CTGF-446-13-13338 13338 CTGF-913-13-13340 13340 CTGF-997-13-13342 13342 CTGF-277-13-13344 13344 CTGF-1052-13-13346 13346 CTGF-887-13-13348 13348 CTGF-914-13-13350 13350 CTGF-1039-13-13352 13352 CTGF-754-13-13354 13354 CTGF-1130-13-13356 13356 CTGF-919-13-13358 13358 CTGF-922-13-13360 13360 CTGF-746-13-13362 13362 CTGF-993-13-13364 13364 CTGF-825-13-13366 13366 CTGF-926-13-13368 13368 CTGF-923-13-13370 13370 CTGF-866-13-13372 13372 CTGF-563-13-13374 13374 CTGF-823-13-13376 13376 CTGF-1233-13-13378 13378 CTGF-924-13-13380 13380 CTGF-921-13-13382 13382 CTGF-443-13-13384 13384 CTGF-1041 13-13386 13386 CTGF-1042-13-13388 13388 CTGF-755-13-13390 13390 CTGF-467-13-13392 13392 CTGF-995-13-13394 13394 CTGF-927-13-13396 13396 SPP1-1025-13-13398 13398 SPP1-1049-13-13400 13400 SPP1-1051-13-13402 13402 SPP1-1048-13-13404 13404 SPP1-1050-13-13406 13406 SPP1-1047-13-13408 13408 SPP1-800-13-13410 13410 SPP1-492-13-13412 13412 SPP1-612-13-13414 13414 SPP1-481-13-13416 13416 SPP1-614-13-13418 13418 SPP1-951-13-13420 13420 SPP1-482-13-13422 13422 SPP1-856-13-13424 13424 SPP1-857-13-13426 13426 SPP1-365-13-13428 13428 SPP1-359-13-13430 13430 SPP1-357-13-13432 13432 SPP1-858-13-13434 13434 SPP1-1012-13-13436 13436 SPP1-1014-13-13438 13438 SPP1-356-13-13440 13440 SPP1-368-13-13442 13442 SPP1-1011-13-13444 13444 SPP1-754-13-13446 13446
SPP1-1021-13-13448 13448 SPP1-1330-13-13450 13450 SPP1-346-13-13452 13452 SPP1-869-13-13454 13454 SPP1-701-13-13456 13456 SPP1-896-13-13458 13458 SPP1-1035-13-13460 13460 SPP1-1170-13-13462 13462 SPP1-1282-13-13464 13464 SPP1-1537-13-13466 13466 SPP1-692-13-13468 13468 SPP1-840-13-13470 13470 SPP1-1163-13-13472 13472 SPP1-789-13-13474 13474 SPP1-841-13-13476 13476 SPP1-852-13-13478 13478 SPP1-209-13-13480 13480 SPP1-1276-13-13482 13482 SPP1-137-13-13484 13484 SPP1-711-13-13486 13486 SPP1-582-13-13488 13488 SPP1-839-13-13490 13490 SPP1-1091-13-13492 13492 SPP1-884-13-13494 13494 SPP1-903-13-13496 13496 SPP1-1090-13-13498 13498 SPP1-474-13-13500 13500 SPP1-575-13-13502 13502 SPP1-671-13-13504 13504 SPP1-924-13-13506 13506 SPP1-1185-13-13508 13508 SPP1-1221-13-13510 13510 SPP1-347-13-13512 13512 SPP1-634-13-13514 13514 SPP1-877-13-13516 13516 SPP1-1033-13-13518 13518 SPP1-714-13-13520 13520 SPP1-791-13-13522 13522 SPP1-813-13-13524 13524 SPP1-939-13-13526 13526 SPP1-1161-13-13528 13528 SPP1-1164-13-13530 13530 SPP1-1190-13-13532 13532 SPP1-1333-13-13534 13534 SPP1-537-13-13536 13536 SPP1-684-13-13538 13538 SPP1-707-13-13540 13540 SPP1-799-13-13542 13542 SPP1-853-13-13544 13544 SPP1-888-13-13546 13546 SPP1-1194-13-13548 13548 SPP1-1279-13-13550 13550 SPP1-1300-13-13552 13552 SPP1-1510-13-13554 13554 SPP1-1543-13-13556 13556 SPP1-434-13-13558 13558 SPP1-600-13-13560 13560 SPP1-863-13-13562 13562 SPP1-902-13-13564 13564 SPP1-921-13-13566 13566 SPP1-154-13-13568 13568 SPP1-217-13-13570 13570 SPP1-816-13-13572 13572 SPP1-882-13-13574 13574 SPP1-932-13-13576 13576 SPP1-1509-13-13578 13578 SPP1-157-13-13580 13580 SPP1-350-13-13582 13582 SPP1-511-13-13584 13584 SPP1-605-13-13586 13586 SPP1-811-13-13588 13588 SPP1-892-13-13590 13590 SPP1-922-13-13592 13592 SPP1-1169-13-13594 13594 SPP1-1182-13-13596 13596 SPP1-1539-13-13598 13598 SPP1-1541-13-13600 13600 SPP1-427-13-13602 13602 SPP1-533-13-13604 13604 APOB--13-13763 13763 APOB--13-13764 13764 MAP4K4--16-13766 13766 oooooooooooos Pm000fffff0m00 UAGACUUCCACAGAACU 49 ssssso 00mmm0 CU PPIB--13-13767 13767 PPIB--15-13768 13768 PPIB--17-13769 13769 MAP4K4--16-13939 13939 oooooooooooos m000f0ffff0m0m UAGACAUCCUACACAGC 50 ssssso 00m0m AC APOB-4314-16-13940 13940 oooooooooooos Pm0fffffff000m UGUUUCUCCAGAUCCUU 51 ssssso mmmm00 GC APOB-4314-17-13941 13941 oooooooooooos Pm0fffffff000m UGUUUCUCCAGAUCCUU 52 ssssso mmmm00 GC APOB--16-13942 13942 oooooooooooos Pm00f000f000mm UAGCAGAUGAGUCCAUU 53 ssssso m0mmm0 UG APOB--18-13943 13943 ooooooooooooo Pm00f000f000mm UAGCAGAUGAGUCCAUU 54 ooosssssso m0mmm00000 UGGAGA APOB--17-13944 13944 oooooooooooos Pm00f000f000mm UAGCAGAUGAGUCCAUU 55 ssssso m0mmm0 UG APOB--19-13945 13945 ooooooooooooo Pm00f000f000mm UAGCAGAUGAGUCCAUU 56 ooosssssso m0mmm00000 UGGAGA APOB-4314-16-13946 13946 oooooooooooos Pmf0ff0ffffmmm AUGUUGUUUCUCCAGAU 57 ssssso 000mm0 CC APOB-4314-17-13947 13947 oooooooooooos Pmf0ff0ffffmmm AUGUUGUUUCUCCAGAU 58 ssssso 000mm0 CC APOB--16-13948 13948 oooooooooooos Pm0fff000000mm UGUUUGAGGGACUCUGU 59 ssssso mm0m00 GA APOB--17- 13949 oooooooooooos Pm0fff000000mm UGUUUGAGGGACUCUGU 60 13949 ssssso mm0m00 GA APOB--16- 13950 oooooooooooos Pmff00f0fff00m AUUGGUAUUCAGUGUGA 61 13950 ssssso 0m00m0 UG APOB--18- 13951 ooooooooooooo Pmff00f0fff00m AUUGGUAUUCAGUGUGA 62 13951 ooosssssso 0m00m00m00 UGACAC APOB--17- 13952 oooooooooooos Pmff00f0fff00m AUUGGUAUUCAGUGUGA 63 13952 ssssso 0m00m0 UG APOB--19- 13953 ooooooooooooo Pmff00f0fff00m AUUGGUAUUCAGUGUGA 64 13953 ooosssssso 0m00m00m00 UGACAC MAP4K4-- 13766.2 oooooooooooos Pm000fffff0m00 UAGACUUCCACAGAACU 65 16-13766.2 ssssso 00mmm0 CU CTGF-1222- 13980 oooooooooooos Pm0f0ffffffm0m UACAUCUUCCUGUAGUA 66 16-13980 ssssso 00m0m0 CA CTGF-813- 13981 oooooooooooos Pm0f0ffff0mmmm AGGCGCUCCACUCUGUG 67 16-13981 ssssso 0m000 GU CTGF-747- 13982 oooooooooooos Pm0ffffff00mm0 UGUCUUCCAGUCGGUAA 68 16-13982 ssssso m0000 GC CTGF-817- 13983 oooooooooooos Pm00f000f0fmmm GAACAGGCGCUCCACUC 69 16-13983 ssssso 0mmmm0 UG CTGF-1174- 13984 oooooooooooos Pm00ff0f00f00m CAGUUGUAAUGGCAGGC 70 16-13984 ssssso 000m00 AC CTGF-1005- 13985 oooooooooooos Pmff000000mmm0 AGCCAGAAAGCUCAAAC 71 16-13985 ssssso 00mm0 UU CTGF-814- 13986 oooooooooooos Pm000f0ffff0mm CAGGCGCUCCACUCUGU 72 16-13986 ssssso mm0m00 GG CTGF-816- 13987 oooooooooooos Pm0f000f0ffmm0 AACAGGCGCUCCACUCU 73 16-13987 ssssso mmmm00 GU CTGF-1001- 13988 oooooooooooos Pm0000fff000mm AGAAAGCUCAAACUUGA 74 16-13988 ssssso m00m0 UA CTGF-1173- 13989 oooooooooooos Pmff0f00f00m00 AGUUGUAAUGGCAGGCA 75
16-13989 ssssso 0m0m0 CA CTGF-749- 13990 oooooooooooos Pmf0ffffff00mm CGUGUCUUCCAGUCGGU 76 16-13990 ssssso 00m00 AA CTGF-792- 13991 oooooooooooos Pm00ff000f00mm GGACCAGGCAGUUGGCU 77 16-13991 ssssso 00mmm0 CU CTGF-1162- 13992 oooooooooooos Pm000f0f000mmm CAGGCACAGGUCUUGAU 78 16-13992 ssssso m00m00 GA CTGF-811- 13993 oooooooooooos Pmf0ffff0ffmm0 GCGCUCCACUCUGUGGU 79 16-13993 ssssso m00mm0 CU CTGF-797- 13994 oooooooooooos Pm0fff000ff000 GGUCUGGACCAGGCAGU 80 16-13994 ssssso m00mm0 UG CTGF-1175- 13995 oooooooooooos Pmf00ff0f00m00 ACAGUUGUAAUGGCAGG 81 16-13995 ssssso m000m0 CA CTGF-1172- 13996 oooooooooooos Pmff0f00f00m00 GUUGUAAUGGCAGGCAC 82 16-13996 ssssso 0m0m00 AG CTGF-1177- 13997 oooooooooooos Pm00f00ff0f00m GGACAGUUGUAAUGGCA 83 16-13997 ssssso 00m000 GG CTGF-1176- 13998 oooooooooooos Pm0f00ff0f00m0 GACAGUUGUAAUGGCAG 84 16-13998 ssssso 0m0000 GC CTGF-812- 13999 oooooooooooos Pm0f0ffff0fmmm GGCGCUCCACUCUGUGG 85 16-13999 ssssso 0m00m0 UC CTGF-745- 14000 oooooooooooos Pmfffff00ff00m UCUUCCAGUCGGUAAGC 86 16-14000 ssssso 000mm0 CG CTGF-1230- 14001 oooooooooooos Pm0fffff0f0m0m UGUCUCCGUACAUCUUC 87 16-14001 ssssso mmmmm0 CU CTGF-920- 14002 oooooooooooos Pmffff0f0000mm AGCUUCGCAAGGCCUGA 88 16-14002 ssssso m00m0 CC CTGF-679- 14003 oooooooooooos Pm0ffffff0f00m CACUCCUCGCAGCAUUU 89 16-14003 ssssso 0mmmm0 CC CTGF-992- 14004 oooooooooooos Pm00fff00f000m AAACUUGAUAGGCUUGG 90 16-14004 ssssso mm0000 AG CTGF-1045- 14005 oooooooooooos Pmffff0f0000mm ACUCCACAGAAUUUAGC 91 16-14005 ssssso m00mm0 UC CTGF-1231- 14006 oooooooooooos Pmf0fffff0f0m0 AUGUCUCCGUACAUCUU 92 16-14006 ssssso mmmmm0 CC CTGF-991- 14007 oooooooooooos Pm0fff00f000mm AACUUGAUAGGCUUGGA 93 16-14007 ssssso m00000 GA CTGF-998- 14008 oooooooooooos Pm00fff000fmm0 AAGCUCAAACUUGAUAG 94 16-14008 ssssso 0m0000 GC CTGF-1049- 14009 oooooooooooos Pmf0f0ffff0m00 ACAUACUCCACAGAAUU 95 16-14009 ssssso 00mmm0 UA CTGF-1044- 14010 oooooooooooos Pmfff0f0000mmm CUCCACAGAAUUUAGCU 96 16-14010 ssssso 00mmm0 CG CTGF-1327- 14011 oooooooooooos Pm0f0ff0ff0000 UGUGCUACUGAAAUCAU 97 16-14011 ssssso mm0mm0 UU CTGF-1196- 14012 oooooooooooos Pm0000f0ff0mm0 AAAGAUGUCAUUGUCUC 98 16-14012 ssssso mmmmm0 CG CTGF-562- 14013 oooooooooooos Pmf0f0ff00f0mm GUGCACUGGUACUUGCA 99 16-14013 ssssso m0m000 GC CTGF-752- 14014 oooooooooooos Pm00f0f0fffmmm AAACGUGUCUUCCAGUC 100 16-14014 ssssso 00mm00 GG CTGF-994- 14015 oooooooooooos Pmf000fff00m00 UCAAACUUGAUAGGCUU 101 16-14015 ssssso 0mmm00 GG CTGF-1040- 14016 oooooooooooos Pmf0000fff00mm ACAGAAUUUAGCUCGGU 102 16-14016 ssssso m00m00 AU CTGF-1984- 14017 oooooooooooos Pmf0f0ffff0mmm UUACAUUCUACCUAUGG 103 16-14017 ssssso 0m00m0 UG CTGF-2195- 14018 oooooooooooos Pm00ff00ff00mm AAACUGAUCAGCUAUAU 104 16-14018 ssssso 0m0m00 AG CTGF-2043- 14019 oooooooooooos Pm0fff000f0000 UAUCUGAGCAGAAUUUC 105 16-14019 ssssso mmmmm0 CA CTGF-1892- 14020 oooooooooooos Pmf00fff000m00 UUAACUUAGAUAACUGU 106 16-14020 ssssso mm0m00 AC CTGF-1567- 14021 oooooooooooos Pm0ff0fff0f0m0 UAUUACUCGUAUAAGAU 107 16-14021 ssssso 000m00 GC CTGF-1780- 14022 oooooooooooos Pm00ff0fff00mm AAGCUGUCCAGUCUAAU 108 16-14022 ssssso m00mm0 CG CTGF-2162- 14023 oooooooooooos Pm00f00000fm0m UAAUAAAGGCCAUUUGU 109 16-14023 ssssso mm0mm0 UC CTGF-1034- 14024 oooooooooooos Pmff00fff00m0m UUUAGCUCGGUAUGUCU 110 16-14024 ssssso 0mmmm0 UC CTGF-2264- 14025 oooooooooooos Pmf0fffff00m00 ACACUCUCAACAAAUAA 111 16-14025 ssssso 0m0000 AC CTGF-1032- 14026 oooooooooooos Pm00fff00f0m0m UAGCUCGGUAUGUCUUC 112 16-14026 ssssso mmmm00 AU CTGF-1535- 14027 oooooooooooos Pm00fffffff0mm UAACCUUUCUGCUGGUA 113 16-14027 ssssso 00m0m0 CC CTGF-1694- 14028 oooooooooooos Pmf000000f00mm UUAAGGAACAACUUGAC 114 16-14028 ssssso m00mm0 UC CTGF-1588- 14029 oooooooooooos Pmf0f0ffff000m UUACACUUCAAAUAGCA 115 16-14029 ssssso 00m000 GG CTGF-928- 14030 oooooooooooos Pmff000ff00mmm UCCAGGUCAGCUUCGCA 116 16-14030 ssssso m0m000 AG CTGF-1133- 14031 oooooooooooos Pmffffff0f00mm CUUCUUCAUGACCUCGC 117 16-14031 ssssso mm0mm0 CG CTGF-912- 14032 oooooooooooos Pm000fff00fm0m AAGGCCUGACCAUGCAC 118 16-14032 ssssso 0m0m00 AG CTGF-753- 14033 oooooooooooos Pm000f0f0ffmmm CAAACGUGUCUUCCAGU 119 16-14033 ssssso m00mm0 CG CTGF-918- 14034 oooooooooooos Pmfff0f0000mmm CUUCGCAAGGCCUGACC 120 16-14034 ssssso 00mm00 AU CTGF-744- 14035 oooooooooooos Pmffff00ff00m0 CUUCCAGUCGGUAAGCC 121 16-14035 ssssso 00mm00 GC CTGF-466- 14036 oooooooooooos Pmf00ffff0f00m CCGAUCUUGCGGUUGGC 122 16-14036 ssssso m00mm0 CG CTGF-917- 14037 oooooooooooos Pmff0f0000fmm0 UUCGCAAGGCCUGACCA 123 16-14037 ssssso 0mm0m0 UG CTGF-1038- 14038 oooooooooooos Pm00fff00fmm0m AGAAUUUAGCUCGGUAU 124 16-14038 ssssso 0m00 GU CTGF-1048- 14039 oooooooooooos Pm0f0ffff0f000 CAUACUCCACAGAAUUU 125 16-14039 ssssso 0mmm00 AG CTGF-1235- 14040 oooooooooooos Pm0ff0f0fffmmm UGCCAUGUCUCCGUACA 126 16-14040 ssssso 0m0m0 UC CTGF-868- 14041 oooooooooooos Pm000f0ff0fm0m GAGGCGUUGUCAUUGGU 127 16-14041 ssssso m00m00 AA CTGF-1131- 14042 oooooooooooos Pmffff0f00fmmm UCUUCAUGACCUCGCCG 128 16-14042 ssssso 0mm0m0 UC CTGF-1043- 14043 oooooooooooos Pmff0f0000fmm0 UCCACAGAAUUUAGCUC 129 16-14043 ssssso 0mmm00 GG CTGF-751- 14044 oooooooooooos Pm0f0f0ffffmm0 AACGUGUCUUCCAGUCG 130 16-14044 ssssso 0mm000 GU CTGF-1227- 14045 oooooooooooos Pmfff0f0f0fmmm CUCCGUACAUCUUCCUG 131 16-14045 ssssso mmm0m0 UA CTGF-867- 14046 oooooooooooos Pm0f0ff0ff0mm0 AGGCGUUGUCAUUGGUA 132 16-14046 ssssso 0m000 AC CTGF-1128- 14047 oooooooooooos PmfCf00ffff0mm UCAUGACCUCGCCGUCA 133 16-14047 ssssso 0mm000 GG CTGF-756- 14048 oooooooooooos Pm0ff000f0f0mm GGCCAAACGUGUCUUCC 134 16-14048 ssssso mmmm00 AG CTGF-1234- 14049 oooooooooooos Pmff0f0ffffmm0 GCCAUGUCUCCGUACAU 135 16-14049 ssssso m0mm0 CU CTGF-916- 14050 oooooooooooos Pmf0f0000ffm00 UCGCAAGGCCUGACCAU 136 16-14050 ssssso mm0m00 GC CTGF-925- 14051 oooooooooooos Pm0ff00fffmm00 AGGUCAGCUUCGCAAGG 137 16-14051 ssssso 00m0 CC CTGF-1225- 14052 oooooooooooos Pmf0f0f0fffmmm CCGUACAUCUUCCUGUA 138 16-14052 ssssso m0m000 GU CTGF-445- 14053 oooooooooooos Pm00ff0000fm0m GAGCCGAAGUCACAGAA 139 16-14053 ssssso 000000 GA CTGF-446- 14054 oooooooooooos Pm000ff0000mm0 GGAGCCGAAGUCACAGA 140 16-14054 ssssso m00000 AG CTGF-913- 14055 oooooooooooos Pm0000fff00mm0 CAAGGCCUGACCAUGCA 141 16-14055 ssssso m0m0m0 CA CTGF-997- 14056 oooooooooooos Pmfff000ffm00m AGCUCAAACUUGAUAGG 142 16-14056 ssssso 000m0 CU CTGF-277- 14057 oooooooooooos Pmf0f00ffff00m CUGCAGUUCUGGCCGAC 143 16-14057 ssssso m00m00 GG CTGF-1052- 14058 oooooooooooos Pm0f0f0f0ffmm0 GGUACAUACUCCACAGA 144 16-14058 ssssso m00000 AU CTGF-887- 14059 oooooooooooos Pmf0fffffff00m CUGCUUCUCUAGCCUGC 145 16-14059 ssssso mm0m00 AG CTGF-914- 14060 oooooooooooos Pmf0000fff00mm GCAAGGCCUGACCAUGC 146 16-14060 ssssso 0m0m00 AC CTGF-1039- 14061 oooooooooooos Pm0000fff00mmm CAGAAUUUAGCUCGGUA 147 16-14061 ssssso 00m0m0 UG CTGF-754- 14062 oooooooooooos Pmf000f0f0fmmm CCAAACGUGUCUUCCAG 148 16-14062 ssssso mm00m0 UC CTGF-1130- 14063 oooooooooooos Pmfff0f00ffmmm CUUCAUGACCUCGCCGU 149 16-14063 ssssso m0mm0 CA CTGF-919- 14064 oooooooooooos Pmffff0f0000mm GCUUCGCAAGGCCUGAC 150 16-14064 ssssso m00mm0 CA CTGF-922- 14065 oooooooooooos Pmf00ffff0f000 UCAGCUUCGCAAGGCCU 151 16-14065 ssssso 0mmm00 GA CTGF-746- 14066 oooooooooooos Pmffffff00fm0m GUCUUCCAGUCGGUAAG 152 16-14066 ssssso 000m0 CC CTGF-993- 14067 oooooooooooos Pm000fff00f000 CAAACUUGAUAGGCUUG 153 16-14067 ssssso mmm000 GA CTGF-825- 14068 oooooooooooos Pm0ffff0000m00 AGGUCUUGGAACAGGCG 154 16-14068 ssssso 0m0m0 CU CTGF-926- 14069 oooooooooooos Pm000ff00ffmmm CAGGUCAGCUUCGCAAG 155 16-14069 ssssso 00000 GC CTGF-923- 14070 oooooooooooos Pmff00ffff0m00 GUCAGCUUCGCAAGGCC 156 16-14070 ssssso 00mmm0 UG CTGF-866- 14071 oooooooooooos Pm0f0ff0ff0mm0 GGCGUUGUCAUUGGUAA 157 16-14071 ssssso 0m00m0 CC CTGF-563- 14072 oooooooooooos Pmf0f0ff00m0mm CGUGCACUGGUACUUGC 158 16-14072 ssssso m0m00 AG
CTGF-823- 14073 oooooooooooos Pmffff0000f000 GUCUUGGAACAGGCGCU 159 16-14073 ssssso m0mmm0 CC CTGF-1233- 14074 oooooooooooos Pmf0f0fffff0m0 CCAUGUCUCCGUACAUC 160 16-14074 ssssso m0mmm0 UU CTGF-924- 14075 oooooooooooos Pm0ff00ffff0m0 GGUCAGCUUCGCAAGGC 161 16-14075 ssssso 000mm0 CU CTGF-921- 14076 oooooooooooos Pm00ffff0f0000 CAGCUUCGCAAGGCCUG 162 16-14076 ssssso mmm000 AC CTGF-443- 14077 oooooooooooos Pmff0000ff0m00 GCCGAAGUCACAGAAGA 163 16-14077 ssssso 000000 GG CTGF-1041- 14078 oooooooooooos Pm0f0000fff00m CACAGAAUUUAGCUCGG 164 16-14078 ssssso mm00m0 UA CTGF-1042- 14079 oooooooooooos Pmf0f0000ffm00 CCACAGAAUUUAGCUCG 165 16-14079 ssssso mmm000 GU CTGF-755- 14080 oooooooooooos Pmff000f0f0mmm GCCAAACGUGUCUUCCA 166 16-14080 ssssso mmm000 GU CTGF-467- 14081 oooooooooooos Pmf0f00ffff0m0 GCCGAUCUUGCGGUUGG 167 16-14081 ssssso mm00m0 CC CTGF-995- 14082 oooooooooooos Pmff000fff00m0 CUCAAACUUGAUAGGCU 168 16-14082 ssssso 00mmm0 UG CTGF-927- 14083 oooooooooooos Pmf000ff00fmmm CCAGGUCAGCUUCGCAA 169 16-14083 ssssso 0m0000 GG SPP1-1091- 14131 oooooooooooos Pmff00ff000m0m UUUGACUAAAUGCAAAG 170 16-14131 ssssso 0000m0 UG PPIB--16-14188 14188 ooooooooooooo Pm0fffff0f00mm UGUUUUUGUAGCCAAAU 171 ssssss 000mm0 CC PPIB--17-14189 14189 ooooooooooooo Pm0fffff0f00mm UGUUUUUGUAGCCAAAU 172 ssssss 000mm0 CC PPIB--18-14190 14190 ooooooooooooo Pm0fffff0f00mm UGUUUUUGUAGCCAAAU 173 ssssss 000mm0 CC pGL3-l172- 14386 oooooooooooos Pm00ff0f0ffm0m AAAUCGUAUUUGUCAAU 174 16-14386 ssssso m00mm0 CA pGL3-l172- 14387 oooooooooooos Pm00ff0f0ffm0m AAAUCGUAUUUGUCAAU 175 16-14387 ssssso m00mm0 CA MAP4K4-2931- 14390 25-14390 miR-122--23-14391 14391 14084 oooooooooooos Pmff00fff0f000 UCUAAUUCAUGAGAAAU 616 ssssso 000m00 AC 14085 oooooooooooos Pm00ff00fffm00 UAAUUGACCUCAGAAGA 617 ssssso 0000m0 UG 14086 oooooooooooos Pmff00ff00fmmm UUUAAUUGACCUCAGAA 618 ssssso 000000 GA 14087 oooooooooooos Pm0ff00ffff000 AAUUGACCUCAGAAGAU 619 ssssso 000m00 GC 14088 oooooooooooos Pmf00ff00ffmm0 UUAAUUGACCUCAGAAG 620 ssssso 000000 AU 14089 oooooooooooos Pmff00ffff0000 AUUGACCUCAGAAGAUG 621 ssssso 00m0m0 CA 14090 oooooooooooos Pmf0fff00ff00m UCAUCCAGCUGACUCGU 622 ssssso mm0mm0 UU 14091 oooooooooooos Pm0fff0ff0000m AGAUUCAUCAGAAUGGU 623 ssssso 00m00 GA 14092 oooooooooooos Pm0Cffff00fmm0 UGACCUCAGUCCAUAAA 624 ssssso m000m0 CC 14093 oooooooooooos Pm0f00f0000mmm AAUGGUGAGACUCAUCA 625 ssssso 0mm000 GA 14094 oooooooooooos Pmff00ffff00mm UUUGACCUCAGUCCAUA 626 ssssso m0m000 AA 14095 oooooooooooos Pmff0f00ff0m00 UUCAUGGCUGUGAAAUU 627 ssssso 00mmm0 CA 14096 oooooooooooos Pm00f00f0000mm GAAUGGUGAGACUCAUC 628 ssssso m0mm00 AG 14097 oooooooooooos Pm0Cffffff0mmm UGGCUUUCCGCUUAUAU 629 ssssso 0m0m00 AA 14098 oooooooooooos Pmf00ffffff0mm UUGGCUUUCCGCUUAUA 630 ssssso m0m0m0 UA 14099 oooooooooooos Pmf0fff0f0f00m UCAUCCAUGUGGUCAUG 631 ssssso m0m000 GC 14100 oooooooooooos Pmf0f00ff0f00m AUGUGGUCAUGGCUUUC 632 ssssso mmmm00 GU 14101 oooooooooooos Pmf00ff0f00mmm GUGGUCAUGGCUUUCGU 633 ssssso mm0mm0 UG 14102 oooooooooooos Pmff00fffffmmm AUUGGCUUUCCGCUUAU 634 ssssso m0m00 AU 14103 oooooooooooos Pm00f0f0000mmm AAAUACGAAAUUUCAGG 635 ssssso m000m0 UG 14104 oooooooooooos Pm000f0f0000mm AGAAAUACGAAAUUUCA 636 ssssso mm000 GG 14105 oooooooooooos Pm00ff0f00fmmm UGGUCAUGGCUUUCGUU 637 ssssso m0mm00 GG 14106 oooooooooooos PmfCff0fff0m0m AUAUCAUCCAUGUGGUC 638 ssssso 00mm00 AU 14107 oooooooooooos Pm0f0f0000fmmm AAUACGAAAUUUCAGGU 639 ssssso 000m00 GU 14108 oooooooooooos Pm0ff000000mm0 AAUCAGAAGGCGCGUUC 640 ssssso mmm00 AG 14109 oooooooooooos Pmfff0f000000m AUUCAUGAGAAAUACGA 641 ssssso 0m0000 AA 14110 oooooooooooos Pmf0fff0f00000 CUAUUCAUGAGAGAAUA 642 ssssso 00m000 AC 14111 oooooooooooos Pmfff0ff000mmm UUUCGUUGGACUUACUU 643 ssssso 0mmm00 GG 14112 oooooooooooos Pmf0fffff0fm0m UUGCUCUCAUCAUUGGC 644 ssssso m00mm0 UU 14113 oooooooooooos Pmff00fffffmmm UUCAACUCCUCGCUUUC 645 ssssso mmmm0 CA 14114 oooooooooooos Pm00ff0ff00mm0 UGACUAUCAAUCACAUC 646 ssssso m0mm00 GG 14115 oooooooooooos Pm0f0f0ff0mmm0 AGAUGCACUAUCUAAUU 647 ssssso 0mmm0 CA 14116 oooooooooooos Pm0f000f0f0m0m AAUAGAUACACAUUCAA 648 ssssso mm00m0 CC 14117 oooooooooooos Pmffffff0f0000 UUCUUCUAUAGAAUGAA 649 ssssso m000m0 CA 14118 oooooooooooos Pm0ff0ff000m00 AAUUGCUGGACAACCGU 650 ssssso mm0m00 GG 14119 oooooooooooos Pmf0ffffff0m0m UCGCUUUCCAUGUGUGA 651 ssssso 0m0000 GG 14120 oooooooooooos Pm0Cfff000fm0m UAAUCUGGACUGCUUGU 652 ssssso mm0m00 GG 14121 oooooooooooos Pmf0f0fff00mm0 ACACAUUCAACCAAUAA 653 ssssso 0m0000 AC 14122 oooooooooooos Pmfff0ffff0m00 ACUCGUUUCAUAACUGU 654 ssssso mm0mm0 CC 14123 oooooooooooos Pmf00fff000mm0 AUAAUCUGGACUGCUUG 655 ssssso mmm0m0 UG 14124 oooooooooooos Pmffff0fff0m0m UUUCCGCUUAUAUAAUC 656 ssssso 00mmm0 UG 14125 oooooooooooos Pm0fff00ff00m0 UGUUUAACUGGUAUGGC 657 ssssso m00m00 AC 14126 oooooooooooos Pm0f0000f000m0 UAUAGAAUGAACAUAGA 658 ssssso m000m0 CA 14127 oooooooooooos Pmffffff00fm0m UUUCCUUGGUCGGCGUU 659 ssssso 0mmm0 UG 14128 oooooooooooos Pmf0f0f0ff0mmm GUAUGCACCAUUCAACU 660 ssssso 00mmm0 CC 14129 oooooooooooos Pmf00ff0ff0m0m UCGGCCAUCAUAUGUGU 661 ssssso 0m0mm0 CU 14130 oooooooooooos Pm0fff000ff0mm AAUCUGGACUGCUUGUG 662 ssssso m0m000 GC 14132 oooooooooooos Pmf0ff0000f0mm ACAUCGGAAUGCUCAUU 663 ssssso m0mm00 GC 14133 oooooooooooos Pm00fffff00mm0 AAGUUCCUGACUAUCAA 664 ssssso mm00m0 UC 14134 oooooooooooos Pmf00ff000f0m0 UUGACUAAAUGCAAAGU 665 ssssso 000m00 GA 14135 oooooooooooos Pm0fff0ff000mm AGACUCAUCAGACUGGU 666 ssssso 00m00 GA 14136 oooooooooooos Pmf0f0f0f0fmm0 UCAUAUGUGUCUACUGU 667 ssssso mm0mC0 GG 14137 oooooooooooos Pmf0fffff0fmm0 AUGUCCUCGUCUGUAGC 668 ssssso m00m00 AU 14138 oooooooooooos Pm00fff0f00mm0 GAAUUCACGGCUGACUU 669 ssssso 0mmmm0 UG 14139 oooooooooooos Pmf0fffff000mm UUAUUUCCAGACUCAAA 670 ssssso m000m0 UA 14140 oooooooooooos Pm000ff0f000mm GAAGCCACAAACUAAAC 671 ssssso 000mm0 UA 14141 oooooooooooos Pmffff0ff000mm CUUUCGUUGGACUUACU 672 ssssso m0mmm0 UG 14142 oooooooooooos Pmfff0f0000mmm GUCUGCGAAACUUCUUA 673 ssssso mmm000 GA 14143 oooooooooooos Pm0f0fff0ff0mm AAUGCUCAUUGCUCUCA 674 ssssso mmm0m0 UC 14144 oooooooooooos Pmf0f0ff0ffm00 AUGCACUAUCUAAUUCA 675 ssssso mmm0m0 UG 14145 oooooooooooos Pmff0f0f0f0mm0 CUUGUAUGCACCAUUCA 676 ssssso mmm000 AC 14146 oooooooooooos Pm00fff0fffm0m UGACUCGUUUCAUAACU 677 ssssso 00mm00 GU 14147 oooooooooooos Pmff00f0fffm00 UUCAGCACUCUGGUCAU 678 ssssso mm0mm0 CC 14148 oooooooooooos Pm00fff0f00mm0 AAAUUCAUGGCUGUGGA 679 ssssso m00000 AU 14149 oooooooooooos Pmf0fff00ff00m ACAUUCAACCAAUAAAC 680 ssssso 000mm0 UG
14150 oooooooooooos Pm0f0f0fff00mm UACACAUUCAACCAAUA 681 ssssso 00m000 AA 14151 oooooooooooos Pmff00ff0ffmmm AUUAGUUAUUUCCAGAC 682 ssssso 000mm0 UC 14152 oooooooooooos Pmffff0fff0m00 UUUCUAUUCAUGAGAGA 683 ssssso 000000 AU 14153 oooooooooooos Pmff00ff0ff00m UUCGGUUGCUGGCAGGU 684 ssssso 000mm0 CC 14154 oooooooooooos Pm0f0f0f0000m0 CAUGUGUGAGGUGAUGU 685 ssssso 0m0mm0 CC 14155 oooooooooooos Pmf0ff0fff00mm GCACCAUUCAACUCCUC 686 ssssso mmmm00 GC 14156 oooooooooooos Pm0fff00ff00mm CAUCCAGCUGACUCGUU 687 ssssso m0mmm0 UC 14157 oooooooooooos Pmfffff0fff0m0 CUUUCCGCUUAUAUAAU 688 ssssso m00mm0 CU 14158 oooooooooooos Pm0ff0f0ff0000 AAUCACAUCGGAAUGCU 689 ssssso m0mmm0 CA 14159 oooooooooooos Pmf0f0ff00fm0m ACACAUUAGUUAUUUCC 690 ssssso mmmm00 AG 14160 oooooooooooos Pmfff0f0000m00 UUCUAUAGAAUGAACAU 691 ssssso 0m0m00 AG 14161 oooooooooooos Pm0f00f00f00mm UACAGUGAUAGUUUGCA 692 ssssso m0m0m0 UU 14162 oooooooooooos Pmf000f00ff00m AUAAGCAAUUGACACCA 693 ssssso 0mm0m0 CC 14163 oooooooooooos Pmff0ff00ff0mm UUUAUUAAUUGCUGGAC 694 ssssso 000m00 AA 14164 oooooooooooos Pmf0ff0000fmmm UCAUCAGAGUCGUUCGA 695 ssssso m0000 GU 14165 oooooooooooos Pmf000ff0f0mm0 AUAAACCACACUAUCAC 696 ssssso mm0mm0 CU 14166 oooooooooooos Pmf0ff0ff00mmm UCAUCAUUGGCUUUCCG 697 ssssso mmm0m0 CU 14167 oooooooooooos Pmfffff00fm0mm AGUUCCUGACUAUCAAU 698 ssssso 00mm0 CA 14168 oooooooooooos Pmff0f00ff00mm UUCACGGCUGACUUUGG 699 ssssso mm0000 AA 14169 oooooooooooos Pmffff0f00f00m UUCUCAUGGUAGUGAGU 700 ssssso 000mm0 UU 14170 oooooooooooos Pm0ff00fff0mmm AAUCAGCCUGUUUAACU 701 ssssso 00mm00 GG 14171 oooooooooooos Pm0ffff00f0mmm GGUUUCAGCACUCUGGU 702 ssssso m00mm0 CA 14172 oooooooooooos Pmff0000f0fmm0 AUCGGAAUGCUCAUUGC 703 ssssso mm0mm0 UC 14173 oooooooooooos Pm00ff0f0000mm UGGCUGUGGAAUUCACG 704 ssssso m0m000 GC 14174 oooooooooooos Pm000f00ff00m0 UAAGCAAUUGACACCAC 705 ssssso mm0mm0 CA 14175 oooooooooooos Pm00fffff0f00m CAAUUCUCAUGGUAGUG 706 ssssso 00m000 AG 14176 oooooooooooos Pm00fffff0fm00 UGGCUUUCGUUGGACUU 707 ssssso 0mmm00 AC 14177 oooooooooooos Pm0ff00f00fm00 AAUCAGUGACCAGUUCA 708 ssssso mmm0m0 UC 14178 oooooooooooos Pmfff0f000mm0m AGUCCAUAAACCACACU 709 ssssso 0mm0C AU 14179 oooooooooooos Pm00f0ffff00mm CAGCACUCUGGUCAUCC 710 ssssso 0mmm00 AG 14180 oooooooooooos Pm0ff00ff0f0mm UAUCAAUCACAUCGGAA 711 ssssso 0000m0 UG 14181 oooooooooooos Pmfff0f00ff00m AUUCACGGCUGACUUUG 712 ssssso mmm000 GA 14182 oooooooooooos Pmf000f0f0f0mm AUAGAUACACAUUCAAC 713 ssssso m00mm0 CA 14183 oooooooooooos Pmffff000ffm00 UUUCCAGACUCAAAUAG 714 ssssso 0m0000 AU 14184 oooooooooooos Pmf00ff0ff000m UUAAUUGCUGGACAACC 715 ssssso 00mm00 GU 14185 oooooooooooos Pm0ff00ff0fm00 UAUUAAUUGCUGGACAA 716 ssssso 0m00m0 CC 14186 oooooooooooos Pmff0fff000mm0 AGUCGUUCGAGUCAAUG 717 ssssso 0m000 GA 14187 oooooooooooos Pmff0ff00f000m GUUGCUGGCAGGUCCGU 718 ssssso mm0m00 GG o: phosphodiester; s: phosphorothioate; P: 5' phosphorylation; 0: 2'-OH; F: 2'-fluoro; m: 2' O-methyl; +: LNA modification. Capital letters in the sequence signify riobonucleotides, lower case letters signify deoxyribonucleotides.
TABLE-US-00003 TABLE 3 Sense backbone, chemistry, and sequence information. OHang SEQ Oligo Sense Sense Sense Sense ID ID Number Number Chem. Backbone Chemistry Sequence NO: APOB-10167- 12138 chl oooooooooooo 0000000000000 GUCAUCACACUGA 176 20-12138 oooooooso 0000000 AUACCAAU APOB-10167- 12139 chl oooooooooooo 0000000000000 GUGAUCAGACUCA 177 20-12139 oooooooso 0000000 AUACGAAU MAP4K4- 12266 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 178 2931-13- o 12266 MAP4K4- 12293 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 179 2931-16- o 12293 MAP4K4- 12383 chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 180 2931-16- o 12383 MAP4K4- 12384 chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 181 2931-16- o 12384 MAP4K4- 12385 chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 182 2931-16- o 12385 MAP4K4- 12386 chl ooooooooooss 0mm0m00000mmm CUGUGGAAGUCUA 183 2931-16- o 0 12386 MAP4K4- 12387 chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 184 2931-16- o 12387 MAP4K4- 12388 chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 185 2931-15- o 12388 MAP4K4- 12432 chl oooooooooooo DY547mm0m0000 CUGUGGAAGUCUA 186 2931-13- o 0mmm0 12432 MAP4K4- 12266.2 chl ooooooooooos mm0m00000mmm0 CUGUGGAAGUCUA 187 2931-13- s 12266.2 APOB--21- 12434 chl oooooooooooo 0000000000000 GUCAUCACACUGA 188 12434 oooooooso 0000000 AUACCAAU APOB--21- 12435 chl oooooooooooo DY54700000000 GUGAUCAGACUCA 189 12435 oooooooso 000000000000 AUACGAAU MAP4K4- 12451 chl ooooooooooos 0mm0m00000mmm CUGUGGAAGUCUA 190 2931-16- s 0 12451 MAP4K4- 12452 chl ooooooooooos mm0m00000mmm0 CUGUGGAAGUCUA 191 2931-16- s 12452 MAP4K4- 12453 chl ooooooooooos mm0m00000mmm0 CUGUGGAAGUCUA 192 2931-16- s 12453 MAP4K4- 12454 chl ooooooooooos 0mm0m00000mmm CUGUGGAAGUCUA 193 2931-17- s 0 12454 MAP4K4- 12455 chl ooooooooooos mm0m00000mmm0 CUGUGGAAGUCUA 194 2931-17- s 12455 MAP4K4- 12456 chl ooooooooooos mm0m00000mmm0 CUGUGGAAGUCUA 195 2931-19- s 12456 --27-12480 12480 chl oooooooooooo DY547mm0f000f UCAUAGGUAACCU 196 oooooooooooo 0055f5f00mm00 CUGGUUGAAAGUG sso 000m000 A --27-12481 12481 chl oooooooooooo DY547mm05f050 CGGCUACAGGUGC 197 oooooooooooo 00f05ff0m0000 UUAUGAAGAAAGU sso 0000m00 A APOB-10167- 12505 chl oooooooooooo 0000000000000 GUCAUCACACUGA 198 21-12505 oooooooos 00000000 AUACCAAU APOB-10167- 12506 chl oooooooooooo 0000000000000 GUGAUCAGACUCA 199 21-12506 oooooooos 00000000 AUACGAAU MAP4K4- 12539 chl ooooooooooos DY547mm0m0000 CUGUGGAAGUCUA 200 2931-16- s 0mmm0 12539 APOB-10167- 12505.2 chl oooooooooooo 0000000000000 GUCAUCACACUGA 201 21-12505.2 oooooooso 0000000 AUACCAAU APOB-10167- 12506.2 chl oooooooooooo 0000000000000 GUGAUCAGACUCA 202 21-12506.2 oooooooso 0000000 AUACGAAU MAP4K4--13- 12565 Chl oooooooooooo m0m0000m0mmm0 UGUAGGAUGUCUA 203 12565 o MAP4K4- 12386.2 chl oooooooooooo 0mm0m00000mmm CUGUGGAAGUCUA 204 2931-16- o 0 12386.2 MAP4K4- 12815 chl oooooooooooo m0m0m0m0m0m0m CUGUGGAAGUCUA 205 2931-13- o 0m0m0m0m0m0m0 12815 APOB--13- 12957 Chl ooooooooooos 0mmmmmmmmmmmm ACUGAAUACCAAU 206 12957 TEG s m MAP4K4--16- 12983 chl ooooooooooos mm0m00000mmm0 CUGUGGAAGUCUA 207 12983 s MAP4K4--16- 12984 Chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 208 12984 oo MAP4K4--16- 12985 chl ooooooooooss mmmmmmmmmmmmm CUGUGGAAGUCUA 209 12985 o MAP4K4--16- 12986 chl ooooooooooss mmmmmmmmmmmmm CUGUGGAAGUCUA 210 12986 o MAP4K4--16- 12987 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 211 12987 o MAP4K4--16- 12988 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 212 12988 o MAP4K4--16- 12989 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 213 12989 o MAP4K4--16- 12990 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 214 12990 o MAP4K4--16- 12991 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 215 12991 o MAP4K4--16- 12992 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 216 12992 o MAP4K4--16- 12993 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 217 12993 o MAP4K4--16- 12994 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 218 12994 o MAP4K4--16- 12995 chl ooooooooooss mm0m00000mmm0 CUGUGGAAGUCUA 219 12995 o MAP4K4- 13012 chl oooooooooooo 0000000000000 AGAGUUCUGUGGA 220 2931-19- ooooooo 00000000 AGUCUA 13012 MAP4K4- 13016 chl oooooooooooo DY54700000000 AGAGUUCUGUGGA 221 2931-19- ooooooo 0000000000000 AGUCUA 13016 PPIB--13- 13021 Chl oooooooooooo 0mmm00mm0m000 AUUUGGCUACAAA 222 13021 o pGL3-1172- 13038 chl oooooooooooo 00m000m0m00mm ACAAAUACGAUUU 223 13-13038 o m pGL3-1172- 13040 chl oooooooooooo DY5470m000m0m ACAAAUACGAUUU 224 13-13040 00mm --16-13047 13047 Chl oooooooooooo mm0m00000mmm0 CUGUGGAAGUCUA 225 oo SOD1-530- 13090 chl oooooooooooo 00m00000000m0 AAUGAAGAAAGUA 226 13-13090 o SOD1-523- 13091 chl oooooooooooo 000m00000m000 AGGUGGAAAUGAA 227 13-13091 o SOD1-535- 13092 chl oooooooooooo 000000m0m0000 AGAAAGUACAAAG 228 13-13092 o SOD1-536- 13093 chl oooooooooooo 00000m0m00000 GAAAGUACAAAGA 229 13-13093 o SOD1-396- 13094 chl oooooooooooo 0m0m00mm0mm00 AUGUGACUGCUGA 230 13-13094 o SOD1-385- 13095 chl oooooooooooo 000mmm000m00m AGACUUGGGCAAU 231 13-13095 o SOD1-195- 13096 chl oooooooooooo 0mmmm000m0000 AUUUCGAGCAGAA 232 13-13096 o APOB-4314- 13115 Chl oooooooooooo 0mmm0000000m0 AUCUGGAGAAACA 233 13-13115 0 APOB-3384- 13116 Chl oooooooooooo mm0000m000000 UCAGAACAAGAAA 234 13-13116 o APOB-3547- 13117 Chl oooooooooooo 00mmm0mmm0mm0 GACUCAUCUGCUA 235 13-13117 o APOB-4318- 13118 Chl oooooooooooo 0000000m00m0m GGAGAAACAACAU 236 13-13118 o APOB-3741- 13119 Chl oooooooooooo 00mmmmmm000m0 AGUCCCUCAAACA 237 13-13119 o PPIB--16- 13136 Chl oooooooooooo 00mm0m00000m0 GGCUACAAAAACA 238 13136 oo APOB-4314- 13154 chl oooooooooooo 000mmm0000000 AGAUCUGGAGAAA 239 15-13154 oo m0 CA APOB-3547- 13155 chl oooooooooooo m000mmm0mmm0m UGGACUCAUCUGC 240 15-13155 oo m0 UA APOB-4318- 13157 chl oooooooooooo mm0000000m00m CUGGAGAAACAAC 241 15-13157 oo 0m AU APOB-3741- 13158 chl oooooooooooo 0000mmmmmm000 AGAGUCCCUCAAA 242 15-13158 oo m0 CA APOB--13- 13159 chl oooooooooooo 0mm000m0mm00m ACUGAAUACCAAU 243 13159 APOB--15- 13160 chl oooooooooooo 0m0mm000m0mm0 ACACUGAAUACCA 244 13160 oo 0m AU SOD1-530- 13163 chl oooooooooooo 00m00000000m0 AAUGAAGAAAGUA 245 16-13163 o SOD1-523- 13164 chl oooooooooooo 000m00000m000 AGGUGGAAAUGAA 246 16-13164 o SOD1-535- 13165 chl oooooooooooo 000000m0m0000 AGAAAGUACAAAG 247 16-13165 o SOD1-536- 13166 chl oooooooooooo 00000m0m00000 GAAAGUACAAAGA 248 16-13166 o SOD1-396- 13167 chl oooooooooooo 0m0m00mm0mm00 AUGUGACUGCUGA 249
16-13167 o SOD1-385- 13168 chl oooooooooooo 000mmm000m00m AGACUUGGGCAAU 250 16-13168 o SOD1-195- 13169 chl oooooooooooo 0mmmm000m0000 AUUUCGAGCAGAA 251 16-13169 o pGL3-1172- 13170 chl oooooooooooo 0m000m0m00mmm ACAAAUACGAUUU 252 16-13170 o pGL3-1172- 13171 chl oooooooooooo DY5470m000m0m ACAAAUACGAUUU 253 16-13171 o 00mmm MAP4k4- 13189 chl oooooooooooo 0000000000000 AGAGUUCUGUGGA 254 2931-19- ooooooo 00000000 AGUCUA 13189 CTGF-1222- 13190 Chl oooooooooooo 0m0000000m0m0 ACAGGAAGAUGUA 255 13-13190 o CTGF-813- 13192 Chl oooooooooooo 000m0000m0mmm GAGUGGAGCGCCU 256 13-13192 o CTGF-747- 13194 Chl oooooooooooo m00mm000000m0 CGACUGGAAGACA 257 13-13194 o CTGF-817- 13196 Chl oooooooooooo 0000m0mmm0mmm GGAGCGCCUGUUC 258 13-13196 o CTGF-1174- 13198 Chl oooooooooooo 0mm0mm0m00mm0 GCCAUUACAACUG 259 13-13198 o CTGF-1005- 13200 Chl oooooooooooo 000mmmmmm00mm GAGCUUUCUGGCU 260 13-13200 o CTGF-814- 13202 Chl oooooooooooo 00m0000m0mmm0 AGUGGAGCGCCUG 261 13-13202 o CTGF-816- 13204 Chl oooooooooooo m0000m0mmm0mm UGGAGCGCCUGUU 262 13-13204 o CTGF-1001- 13206 Chl oooooooooooo 0mmm000mmmmmm GUUUGAGCUUUCU 263 13-13206 o CTGF-1173- 13208 Chl oooooooooooo m0mm0mm0m00mm UGCCAUUACAACU 264 13-13208 o CTGF-749- 13210 Chl oooooooooooo 0mm000000m0m0 ACUGGAAGACACG 265 13-13210 o CTGF-792- 13212 Chl oooooooooooo 00mm0mmm00mmm AACUGCCUGGUCC 266 13-13212 o CTGF-1162- 13214 Chl oooooooooooo 000mmm0m0mmm0 AGACCUGUGCCUG 267 13-13214 o CTGF-811- 13216 Chl oooooooooooo m0000m0000m0m CAGAGUGGAGCGC 268 13-13216 o CTGF-797- 13218 Chl oooooooooooo mmm00mmm000mm CCUGGUCCAGACC 269 13-13218 o CTGF-1175- 13220 Chl oooooooooooo mm0mm0m00mm0m CCAUUACAACUGU 270 13-13220 o CTGF-1172- 13222 Chl oooooooooooo mm0mm0mm0m00m CUGCCAUUACAAC 271 13-13222 o CTGF-1177- 13224 Chl oooooooooooo 0mm0m00mm0mmm AUUACAACUGUCC 272 13-13224 o CTGF-1176- 13226 Chl oooooooooooo m0mm0m00mm0mm CAUUACAACUGUC 273 13-13226 o CTGF-812- 13228 Chl oooooooooooo 0000m0000m0mm AGAGUGGAGCGCC 274 13-13228 o CTGF-745- 13230 Chl oooooooooooo 0mm00mm000000 ACCGACUGGAAGA 275 13-13230 o CTGF-1230- 13232 Chl oooooooooooo 0m0m0m00000m0 AUGUACGGAGACA 276 13-13232 o CTGF-920- 13234 Chl oooooooooooo 0mmmm0m0000mm GCCUUGCGAAGCU 277 13-13234 o CTGF-679- 13236 Chl oooooooooooo 0mm0m000000m0 GCUGCGAGGAGUG 278 13-13236 o CTGF-992- 13238 Chl oooooooooooo 0mmm0mm000mmm GCCUAUCAAGUUU 279 13-13238 o CTGF-1045- 13240 Chl oooooooooooo 00mmmm0m0000m AAUUCUGUGGAGU 280 13-13240 o CTGF-1231- 13242 Chl oooooooooooo m0m0m00000m0m UGUACGGAGACAU 281 13-13242 o CTGF-991- 13244 Chl oooooooooooo 00mmm0mm000mm AGCCUAUCAAGUU 282 13-13244 o CTGF-998- 13246 Chl oooooooooooo m000mmm000mmm CAAGUUUGAGCUU 283 13-13246 o CTGF-1049- 13248 Chl oooooooooooo mm0m0000m0m0m CUGUGGAGUAUGU 284 13-13248 o CTGF-1044- 13250 Chl oooooooooooo 000mmmm0m0000 AAAUUCUGUGGAG 285 13-13250 o CTGF-1327- 13252 Chl oooooooooooo mmmm00m00m0m0 UUUCAGUAGCACA 286 13-13252 o CTGF-1196- 13254 Chl oooooooooooo m00m00m0mmmmm CAAUGACAUCUUU 287 13-13254 o CTGF-562- 13256 Chl oooooooooooo 00m0mm00m0m0m AGUACCAGUGCAC 288 13-13256 o CTGF-752- 13258 Chl oooooooooooo 000000m0m0mmm GGAAGACACGUUU 289 13-13258 o CTGF-994- 13260 Chl oooooooooooo mm0mm000mmm00 CUAUCAAGUUUGA 290 13-13260 o CTGF-1040- 13262 Chl oooooooooooo 00mm000mmmm0m AGCUAAAUUCUGU 291 13-13262 o CTGF-1984- 13264 Chl oooooooooooo 000m0000m0m00 AGGUAGAAUGUAA 292 13-13264 o CTGF-2195- 13266 Chl oooooooooooo 00mm00mm00mmm AGCUGAUCAGUUU 293 13-13266 o CTGF-2043- 13268 Chl oooooooooooo mmmm0mmm000m0 UUCUGCUCAGAUA 294 13-13268 o CTGF-1892- 13270 Chl oooooooooooo mm0mmm000mm00 UUAUCUAAGUUAA 295 13-13270 o CTGF-1567- 13272 Chl oooooooooooo m0m0m000m00m0 UAUACGAGUAAUA 296 13-13272 o CTGF-1780- 13274 Chl oooooooooooo 00mm000m00mmm GACUGGACAGCUU 297 13-13274 o CTGF-2162- 13276 Chl oooooooooooo 0m00mmmmm0mm0 AUGGCCUUUAUUA 298 13-13276 o CTGF-1034- 13278 Chl oooooooooooo 0m0mm000mm000 AUACCGAGCUAAA 299 13-13278 o CTGF-2264- 13280 Chl oooooooooooo mm0mm00000m0m UUGUUGAGAGUGU 300 13-13280 o CTGF-1032- 13282 Chl oooooooooooo 0m0m0mm000mm0 ACAUACCGAGCUA 301 13-13282 o CTGF-1535- 13284 Chl oooooooooooo 00m0000000mm0 AGCAGAAAGGUUA 302 13-13284 o CTGF-1694- 13286 Chl oooooooooooo 00mm0mmmmmm00 AGUUGUUCCUUAA 303 13-13286 o CTGF-1588- 13288 Chl oooooooooooo 0mmm0000m0m00 AUUUGAAGUGUAA 304 13-13288 o CTGF-928- 13290 Chl oooooooooooo 000mm00mmm000 AAGCUGACCUGGA 305 13-13290 o CTGF-1133- 13292 Chl oooooooooooo 00mm0m0000000 GGUCAUGAAGAAG 306 13-13292 o CTGF-912- 13294 Chl oooooooooooo 0m00mm000mmmm AUGGUCAGGCCUU 307 13-13294 o CTGF-753- 13296 Chl oooooooooooo 00000m0m0mmm0 GAAGACACGUUUG 308 13-13296 o CTGF-918- 13298 Chl oooooooooooo 000mmmm0m0000 AGGCCUUGCGAAG 309 13-13298 o CTGF-744- 13300 Chl oooooooooooo m0mm0mm00000 UACCGACUGGAAG 310 13-13300 o CTGF-466- 13302 Chl oooooooooooo 0mm0m0000mm0 ACCGCAAGAUCGG 311 13-13302 o CTGF-917- 13304 Chl oooooooooooo m000mmmm0m000 CAGGCCUUGCGAA 312 13-13304 o CTGF-1038- 13306 Chl oooooooooooo m000mm000mmmm CGAGCUAAAUUCU 313 13-13306 o CTGF-1048- 13308 Chl oooooooooooo mmm0m0000m0m0 UCUGUGGAGUAUG 314 13-13308 o CTGF-1235- 13310 Chl oooooooooooo m00000m0m00m0 CGGAGACAUGGCA 315 13-13310 o CTGF-868- 13312 Chl oooooooooooo 0m00m00m0mmmm AUGACAACGCCUC 316 13-13312 o CTGF-1131- 13314 Chl oooooooooooo 0000mm0m00000 GAGGUCAUGAAGA 317 13-13314 o CTGF-1043- 13316 Chl oooooooooooo m000mmmm0m000 UAAAUUCUGUGGA 318 13-13316 o CTGF-751- 13318 Chl oooooooooooo m000000m0m0mm UGGAAGACACGUU 319 13-13318 o CTGF-1227- 13320 Chl oooooooooooo 0000m0m0m0000 AAGAUGUACGGAG 320 13-13320 0 CTGF-867- 13322 Chl oooooooooooo 00m00m00m0mmm AAUGACAACGCCU 321 13-13322 o CTGF-1128- 13324 Chl oooooooooooo 00m0000mm0m00 GGCGAGGUCAUGA 322 13-13324 o CTGF-756- 13326 Chl oooooooooooo 00m0m0mmm00mm GACACGUUUGGCC 323 13-13326 o CTGF-1234- 13328 Chl oooooooooooo 0m00000m0m00m ACGGAGACAUGGC 324 13-13328 o CTGF-916- 13330 Chl oooooooooooo mm000mmmm0m00 UCAGGCCUUGCGA 325 13-13330 o CTGF-925- 13332 Chl oooooooooooo 0m0000mm00mmm GCGAAGCUGACCU 326 13-13332 o CTGF-1225- 13334 Chl oooooooooooo 000000m0m0m00 GGAAGAUGUACGG 327 13-13334 o CTGF-445- 13336 Chl oooooooooooo 0m00mmmm00mmm GUGACUUCGGCUC 328 13-13336 o CTGF-446- 13338 Chl oooooooooooo m00mmmm00mmmm UGACUUCGGCUCC 329 13-13338 o CTGF-913- 13340 Chl oooooooooooo m00mm000mmmm0 UGGUCAGGCCUUG 330 13-13340 o CTGF-997- 13342 Chl oooooooooooo mm000mmm000mm UCAAGUUUGAGCU 331 13-13342 o CTGF-277- 13344 Chl oooooooooooo 0mm0000mm0m00 GCCAGAACUGCAG 332 13-13344 o
CTGF-1052- 13346 Chl oooooooooooo m0000m0m0m0mm UGGAGUAUGUACC 333 13-13346 o CTGF-887- 13348 Chl oooooooooooo 0mm0000000m00 GCUAGAGAAGCAG 334 13-13348 o CTGF-914- 13350 Chl oooooooooooo 00mm000mmmm0m GGUCAGGCCUUGC 335 13-13350 o CTGF-1039- 13352 Chl oooooooooooo 000mm000mmmm0 GAGCUAAAUUCUG 336 13-13352 o CTGF-754- 13354 Chl oooooooooooo 0000m0m0mmm00 AAGACACGUUUGG 337 13-13354 o CTGF-1130- 13356 Chl oooooooooooo m0000mm0m0000 CGAGGUCAUGAAG 338 13-13356 o CTGF-919- 13358 Chl oooooooooooo 00mmmm0m0000m GGCCUUGCGAAGC 339 13-13358 o CTGF-922- 13360 Chl oooooooooooo mmm0m0000mm00 CUUGCGAAGCUGA 340 13-13360 o CTGF-746- 13362 Chl oooooooooooo mm00mm000000m CCGACUGGAAGAC 341 13-13362 o CTGF-993- 13364 Chl oooooooooooo mmm0mm000mmm0 CCUAUCAAGUUUG 342 13-13364 o CTGF-825- 13366 Chl oooooooooooo m0mmmm0000mmm UGUUCCAAGACCU 343 13-13366 o CTGF-926- 13368 Chl oooooooooooo m0000mm00mmm0 CGAAGCUGACCUG 344 13-13368 o CTGF-923- 13370 Chl oooooooooooo mm0m0000mm00m UUGCGAAGCUGAC 345 13-13370 o CTGF-866- 13372 Chl oooooooooooo m00m00m00m0mm CAAUGACAACGCC 346 13-13372 o CTGF-563- 13374 Chl oooooooooooo 0m0mm00m0m0m0 GUACCAGUGCACG 347 13-13374 o CTGF-823- 13376 Chl oooooooooooo mmm0mmmm0000m CCUGUUCCAAGAC 348 13-13376 o CTGF-1233- 13378 Chl oooooooooooo m0m00000m0m00 UACGGAGACAUGG 349 13-13378 o CTGF-924- 13380 Chl oooooooooooo m0m0000mm00mm UGCGAAGCUGACC 350 13-13380 o CTGF-921- 13382 Chl oooooooooooo mmmm0m0000mm0 CCUUGCGAAGCUG 351 13-13382 o CTGF-443- 13384 Chl oooooooooooo mm0m00mmmm00m CUGUGACUUCGGC 352 13-13384 o CTGF-1041- 13386 Chl oooooooooooo 0mm000mmmm0m0 GCUAAAUUCUGUG 353 13-13386 o CTGF-1042- 13388 Chl oooooooooooo mm000mmmm0m00 CUAAAUUCUGUGG 354 13-13388 o CTGF-755- 13390 Chl oooooooooooo 000m0m0mmm00m AGACACGUUUGGC 355 13-13390 o CTGF-467- 13392 Chl oooooooooooo mm0m0000mm00m CCGCAAGAUCGGC 356 13-13392 o CTGF-995- 13394 Chl oooooooooooo m0mm000mmm000 UAUCAAGUUUGAG 357 13-13394 o CTGF-927- 13396 Chl oooooooooooo 0000mm00mmm00 GAAGCUGACCUGG 358 13-13396 o SPP1-1025- 13398 Chl oooooooooooo mmm0m000mm000 CUCAUGAAUUAGA 359 13-13398 o SPP1-1049- 13400 Chl oooooooooooo mm0000mm00mm0 CUGAGGUCAAUUA 360 13-13400 o SPP1-1051- 13402 Chl oooooooooooo 0000mm00mm000 GAGGUCAAUUAAA 361 13-13402 o SPP1-1048- 13404 Chl oooooooooooo mmm0000mm00mm UCUGAGGUCAAUU 362 13-13404 o SPP1-1050- 13406 Chl oooooooooooo m0000mm00mm00 UGAGGUCAAUUAA 363 13-13406 o SPP1-1047- 13408 Chl oooooooooooo mmmm0000mm00m UUCUGAGGUCAAU 364 13-13408 o SPP1-800- 13410 Chl oooooooooooo 0mm00mm000m00 GUCAGCUGGAUGA 365 13-13410 o SPP1-492- 13412 Chl oooooooooooo mmmm00m000mmm UUCUGAUGAAUCU 366 13-13412 o SPP1-612- 13414 Chl oooooooooooo m000mm0000mm0 UGGACUGAGGUCA 367 13-13414 o SPP1-481- 13416 Chl oooooooooooo 000mmmm0mm0mm GAGUCUCACCAUU 368 13-13416 o SPP1-614- 13418 Chl oooooooooooo 00mm0000mm000 GACUGAGGUCAAA 369 13-13418 o SPP1-951- 13420 Chl oooooooooooo mm0m00mm0m000 UCACAGCCAUGAA 370 13-13420 o SPP1-482- 13422 Chl oooooooooooo 00mmmm0mm0mmm AGUCUCACCAUUC 371 13-13422 o SPP1-856- 13424 Chl oooooooooooo 000m000000mm0 AAGCGGAAAGCCA 372 13-13424 o SPP1-857- 13426 Chl oooooooooooo 00m000000mm00 AGCGGAAAGCCAA 373 13-13426 o SPP1-365- 13428 Chl oooooooooooo 0mm0m0m000m00 ACCACAUGGAUGA 374 13-13428 o SPP1-359- 13430 Chl oooooooooooo 0mm0m00mm0m0m GCCAUGACCACAU 375 13-13430 o SPP1-357- 13432 Chl oooooooooooo 000mm0m00mm0m AAGCCAUGACCAC 376 13-13432 o SPP1-858- 13434 Chl oooooooooooo 0m000000mm00m GCGGAAAGCCAAU 377 13-13434 o SPP1-1012- 13436 Chl oooooooooooo 000mmmm0m0mmm AAAUUUCGUAUUU 378 13-13436 o SPP1-1014- 13438 Chl oooooooooooo 0mmmm0m0mmmmm AUUUCGUAUUUCU 379 13-13438 o SPP1-356- 13440 Chl oooooooooooo 0000mm0m00mm0 AAAGCCAUGACCA 380 13-13440 o SPP1-368- 13442 Chl oooooooooooo 0m0m000m00m0m ACAUGGAUGAUAU 381 13-13442 o SPP1-1011- 13444 Chl oooooooooooo 0000mmmm0m0mm GAAAUUUCGUAUU 382 13-13444 o SPP1-754- 13446 Chl oooooooooooo 0m0mmmmmm00mm GCGCCUUCUGAUU 383 13-13446 o SPP1-1021- 13448 Chl oooooooooooo 0mmmmmm0m000m AUUUCUCAUGAAU 384 13-13448 o SPP1-1330- 13450 Chl oooooooooooo mmmmm0m000m00 CUCUCAUGAAUAG 385 13-13450 o SPP1-346- 13452 Chl oooooooooooo 000mmm00m0000 AAGUCCAACGAAA 386 13-13452 o SPP1-869- 13454 Chl oooooooooooo 0m00m00000m00 AUGAUGAGAGCAA 387 13-13454 o SPP1-701- 13456 Chl oooooooooooo 0m000000mm000 GCGAGGAGUUGAA 388 13-13456 o SPP1-896- 13458 Chl oooooooooooo m00mm00m00mm0 UGAUUGAUAGUCA 389 13-13458 o SPP1-1035- 13460 Chl oooooooooooo 000m00m0m0mmm AGAUAGUGCAUCU 390 13-13460 o SPP1-1170- 13462 Chl oooooooooooo 0m0m0m0mmm0mm AUGUGUAUCUAUU 391 13-13462 o SPP1-1282- 13464 Chl oooooooooooo mmmm0m0000000 UUCUAUAGAAGAA 392 13-13464 o SPP1-1537- 13466 Chl oooooooooooo mm0mmm00m00mm UUGUCCAGCAAUU 393 13-13466 o SPP1-692- 13468 Chl oooooooooooo 0m0m000000m00 ACAUGGAAAGCGA 394 13-13468 o SPP1-840- 13470 Chl oooooooooooo 0m00mmm000mm0 GCAGUCCAGAUUA 395 13-13470 o SPP1-1163- 13472 Chl oooooooooooo m00mm000m0m0m UGGUUGAAUGUGU 396 13-13472 o SPP1-789- 13474 Chl oooooooooooo mm0m0000m000m UUAUGAAACGAGU 397 13-13474 o SPP1-841- 13476 Chl oooooooooooo m00mmm000mm0m CAGUCCAGAUUAU 398 13-13476 o SPP1-852- 13478 Chl oooooooooooo 0m0m000m00000 AUAUAAGCGGAAA 399 13-13478 o SPP1-209- 13480 Chl oooooooooooo m0mm00mm000m0 UACCAGUUAAACA 400 13-13480 o SPP1-1276- 13482 Chl oooooooooooo m0mmm0mmmm0m0 UGUUCAUUCUAUA 401 13-13482 o SPP1-137- 13484 Chl oooooooooooo mm00mm0000000 CCGACCAAGGAAA 402 13-13484 o SPP1-711- 13486 Chl oooooooooooo 000m00m0m0m0m GAAUGGUGCAUAC 403 13-13486 o SPP1-582- 13488 Chl oooooooooooo 0m0m00m00mm00 AUAUGAUGGCCGA 404 13-13488 o SPP1-839- 13490 Chl oooooooooooo 00m00mmm000mm AGCAGUCCAGAUU 405 13-13490 o SPP1-1091- 13492 Chl oooooooooooo 0m0mmm00mm000 GCAUUUAGUCAAA 406 13-13492 o SPP1-884- 13494 Chl oooooooooooo 00m0mmmm00m0m AGCAUUCCGAUGU 407 13-13494 o SPP1-903- 13496 Chl oooooooooooo m00mm00000mmm UAGUCAGGAACUU 408 13-13496 o SPP1-1090- 13498 Chl oooooooooooo m0m0mmm00mm00 UGCAUUUAGUCAA 409 13-13498 o SPP1-474- 13500 Chl oooooooooooo 0mmm00m000mmm GUCUGAUGAGUCU 410 13-13500 o SPP1-575- 13502 Chl oooooooooooo m000m0m0m0m00 UAGACACAUAUGA 411 13-13502 o SPP1-671- 13504 Chl oooooooooooo m000m00000m0m CAGACGAGGACAU 412 13-13504 o SPP1-924- 13506 Chl oooooooooooo m00mm0m000mmm CAGCCGUGAAUUC 413 13-13506 o SPP1-1185- 13508 Chl oooooooooooo 00mmm00000m00 AGUCUGGAAAUAA 414 13-13508 o SPP1-1221- 13510 Chl oooooooooooo 00mmm0m00mmmm AGUUUGUGGCUUC 415 13-13510 o SPP1-347- 13512 Chl oooooooooooo 00mmm00m00000 AGUCCAACGAAAG 416
13-13512 o SPP1-634- 13514 Chl oooooooooooo 000mmmm0m000m AAGUUUCGCAGAC 417 13-13514 o SPP1-877- 13516 Chl oooooooooooo 00m00m000m0mm AGCAAUGAGCAUU 418 13-13516 o SPP1-1033- 13518 Chl oooooooooooo mm000m00m0m0m UUAGAUAGUGCAU 419 13-13518 o SPP1-714- 13520 Chl oooooooooooo m00m0m0m0m000 UGGUGCAUACAAG 420 13-13520 o SPP1-791- 13522 Chl oooooooooooo 0m0000m000mm0 AUGAAACGAGUCA 421 13-13522 o SPP1-813- 13524 Chl oooooooooooo mm0000m0mm000 CCAGAGUGCUGAA 422 13-13524 o SPP1-939- 13526 Chl oooooooooooo m00mm0m000mmm CAGCCAUGAAUUU 423 13-13526 o SPP1-1161- 13528 Chl oooooooooooo 0mm00mm000m0m AUUGGUUGAAUGU 424 13-13528 o SPP1-1164- 13530 Chl oooooooooooo 00mm000m0m0m0 GGUUGAAUGUGUA 425 13-13530 o SPP1-1190- 13532 Chl oooooooooooo 00000m00mm00m GGAAAUAACUAAU 426 13-13532 o SPP1-1333- 13534 Chl oooooooooooo mm0m000m00000 UCAUGAAUAGAAA 427 13-13534 o SPP1-537- 13536 Chl oooooooooooo 0mm00m00mm000 GCCAGCAACCGAA 428 13-13536 o SPP1-684- 13538 Chl oooooooooooo m0mmmm0m0m0m0 CACCUCACACAUG 429 13-13538 o SPP1-707- 13540 Chl oooooooooooo 00mm000m00m0m AGUUGAAUGGUGC 430 13-13540 o SPP1-799- 13542 Chl oooooooooooo 00mm00mm000m0 AGUCAGCUGGAUG 431 13-13542 o SPP1-853- 13544 Chl oooooooooooo m0m000m000000 UAUAAGCGGAAAG 432 13-13544 o SPP1-888- 13546 Chl oooooooooooo mmmm00m0m00mm UUCCGAUGUGAUU 433 13-13546 o SPP1-1194- 13548 Chl oooooooooooo 0m00mm00m0m0m AUAACUAAUGUGU 434 13-13548 o SPP1-1279- 13550 Chl oooooooooooo mm0mmmm0m0000 UCAUUCUAUAGAA 435 13-13550 o SPP1-1300- 13552 Chl oooooooooooo 00mm0mm0mm0m0 AACUAUCACUGUA 436 13-13552 o SPP1-1510- 13554 Chl oooooooooooo 0mm00mm0mmm0m GUCAAUUGCUUAU 437 13-13554 o SPP1-1543- 13556 Chl oooooooooooo 00m00mm00m000 AGCAAUUAAUAAA 438 13-13556 o SPP1-434- 13558 Chl oooooooooooo 0m00mmmm00m00 ACGACUCUGAUGA 439 13-13558 o SPP1-600- 13560 Chl oooooooooooo m00m0m00mmm0m UAGUGUGGUUUAU 440 13-13560 o SPP1-863- 13562 Chl oooooooooooo 000mm00m00m00 AAGCCAAUGAUGA 441 13-13562 o SPP1-902- 13564 Chl oooooooooooo 0m00mm00000mm AUAGUCAGGAACU 442 13-13564 o SPP1-921- 13566 Chl oooooooooooo 00mm00mm0m000 AGUCAGCCGUGAA 443 13-13566 o SPP1-154- 13568 Chl oooooooooooo 0mm0mm0m00000 ACUACCAUGAGAA 444 13-13568 o SPP1-217- 13570 Chl oooooooooooo 000m000mm00mm AAACAGGCUGAUU 445 13-13570 o SPP1-816- 13572 Chl oooooooooooo 000m0mm0000mm GAGUGCUGAAACC 446 13-13572 o SPP1-882- 13574 Chl oooooooooooo m000m0mmmm00m UGAGCAUUCCGAU 447 13-13574 o SPP1-932- 13576 Chl oooooooooooo 00mmmm0m00mm0 AAUUCCACAGCCA 448 13-13576 o SPP1-1509- 13578 Chl oooooooooooo m0mm00mm0mmm0 UGUCAAUUGCUUA 449 13-13578 o SPP1-157- 13580 Chl oooooooooooo 0mm0m00000mm0 ACCAUGAGAAUUG 450 13-13580 o SPP1-350- 13582 Chl oooooooooooo mm00m00000mm0 CCAACGAAAGCCA 451 13-13582 o SPP1-511- 13584 Chl oooooooooooo mm00mm0mm00mm CUGGUCACUGAUU 452 13-13584 o SPP1-605- 13586 Chl oooooooooooo m00mmm0m000mm UGGUUUAUGGACU 453 13-13586 o SPP1-811- 13588 Chl oooooooooooo 00mm0000m0mm0 GACCAGAGUGCUG 454 13-13588 o SPP1-892- 13590 Chl oooooooooooo 00m0m00mm00m0 GAUGUGAUUGAUA 455 13-13590 o SPP1-922- 13592 Chl oooooooooooo 0mm00mm0m000m GUCAGCCGUGAAU 456 13-13592 o SPP1-1169- 13594 Chl oooooooooooo 00m0m0m0mmm0m AAUGUGUAUCUAU 457 13-13594 o SPP1-1182- 13596 Chl oooooooooooo mm000mmm00000 UUGAGUCUGGAAA 458 13-13596 o SPP1-1539- 13598 Chl oooooooooooo 0mmm00m00mm00 GUCCAGCAAUUAA 459 13-13598 o SPP1-1541- 13600 Chl oooooooooooo mm00m00mm00m0 CCAGCAAUUAAUA 460 13-13600 o SPP1-427- 13602 Chl oooooooooooo 00mmm000m00mm GACUCGAACGACU 461 13-13602 o SPP1-533- 13604 Chl oooooooooooo 0mmm0mm00m00m ACCUGCCAGCAAC 462 13-13604 o APOB--13- 13763 Chl oooooooooooo 0m + 00 + m0 + m0 + m ACtGAaUAcCAaU 463 13763 TEG o APOB--13- 13764 Chl oooooooooooo 0mm000m0mm00m ACUGAAUACCAAU 464 13764 TEG o MAP4K4--16- 13766 Chl oooooooooooo DY547mm0m0000 CUGUGGAAGUCUA 465 13766 o 0mmm0 PPIB--13- 13767 Chl oooooooooooo mmmmmmmmmmmmm GGCUACAAAAACA 466 13767 o PPIB--15- 13768 Chl oooooooooooo mm00mm0m00000 UUGGCUACAAAAA 467 13768 ooo m0 CA PPIB--17- 13769 Chl oooooooooooo 0mmm00mm0m000 AUUUGGCUACAAA 468 13769 ooooo 00m0 AACA MAP4K4--16- 13939 Chl oooooooooooo m0m0000m0mmm0 UGUAGGAUGUCUA 469 13939 o APOB-4314- 13940 Chl oooooooooooo 0mmm0000000m0 AUCUGGAGAAACA 470 16-13940 o APOB-4314- 13941 Chl oooooooooooo 000mmm0000000 AGAUCUGGAGAAA 471 17-13941 ooo m0 CA APOB--16- 13942 Chl oooooooooooo 00mmm0mmm0mm0 GACUCAUCUGCUA 472 13942 o APOB--18- 13943 Chl oooooooooooo 00mmm0mmm0mm0 GACUCAUCUGCUA 473 13943 o APOB--17- 13944 Chl oooooooooooo m000mmm0mmm0m UGGACUCAUCUGC 474 13944 ooo m0 UA APOB--19- 13945 Chl oooooooooooo m000mmm0mmm0m UGGACUCAUCUGC 475 13945 ooo m0 UA APOB-4314- 13946 Chl oooooooooooo 0000000m00m0m GGAGAAACAACAU 476 16-13946 o APOB-4314- 13947 Chl oooooooooooo mm0000000m00m CUGGAGAAACAAC 477 17-13947 ooo 0m AU APOB--16- 13948 Chl oooooooooooo 00mmmmmm000m0 AGUCCCUCAAACA 478 13948 o APOB--17- 13949 Chl oooooooooooo 0000mmmmmm000 AGAGUCCCUCAAA 479 13949 ooo m0 CA APOB--16- 13950 Chl oooooooooooo 0mm000m0mm00m ACUGAAUACCAAU 480 13950 o APOB--18- 13951 Chl oooooooooooo 0mm000m0mm00m ACUGAAUACCAAU 481 13951 o APOB--17- 13952 Chl oooooooooooo 0m0mm000m0mm0 ACACUGAAUACCA 482 13952 ooo 0m AU APOB--19- 13953 Chl oooooooooooo 0m0mm000m0mm0 ACACUGAAUACCA 483 13953 ooo 0m AU MAP4K4--16- 13766.2 Chl oooooooooooo DY547mm0m0000 CUGUGGAAGUCUA 484 13766.2 o 0mmm0 CTGF-1222- 13980 Chl oooooooooooo 0m0000000m0m0 ACAGGAAGAUGUA 485 16-13980 o CTGF-813- 13981 Chl oooooooooooo 000m0000mmmm GAGUGGAGCGCCU 486 16-13981 o CTGF-747- 13982 Chl oooooooooooo m0mm000000m0 CGACUGGAAGACA 487 16-13982 o CTGF-817- 13983 Chl oooooooooooo 0000mmmm0mmm GGAGCGCCUGUUC 488 16-13983 o CTGF-1174- 13984 Chl oooooooooooo 0mm0mm0m00mm0 GCCAUUACAACUG 489 16-13984 o CTGF-1005- 13985 Chl oooooooooooo 000mmmmmm00mm GAGCUUUCUGGCU 490 16-13985 o CTGF-814- 13986 Chl oooooooooooo 00m0000mmmm0 AGUGGAGCGCCUG 491 16-13986 o CTGF-816- 13987 Chl oooooooooooo m0000mmmm0mm UGGAGCGCCUGUU 492 16-13987 o CTGF-1001- 13988 Chl oooooooooooo 0mmm000mmmmmm GUUUGAGCUUUCU 493 16-13988 o CTGF-1173- 13989 Chl oooooooooooo m0mm0mm0m00mm UGCCAUUACAACU 494 16-13989 o CTGF-749- 13990 Chl oooooooooooo 0mm000000m0m ACUGGAAGACACG 495 16-13990 o CTGF-792- 13991 Chl oooooooooooo 00mm0mmm00mmm AACUGCCUGGUCC 496 16-13991 o CTGF-1162- 13992 Chl oooooooooooo 000mmm0m0mmm0 AGACCUGUGCCUG 497 16-13992 o CTGF-811- 13993 Chl oooooooooooo m0000m0000mm CAGAGUGGAGCGC 498 16-13993 o CTGF-797- 13994 Chl oooooooooooo mmm00mmm000mm CCUGGUCCAGACC 499 16-13994 o
CTGF-1175- 13995 Chl oooooooooooo mm0mm0m00mm0m CCAUUACAACUGU 500 16-13995 o CTGF-1172- 13996 Chl oooooooooooo mm0mm0mm0m00m CUGCCAUUACAAC 501 16-13996 o CTGF-1177- 13997 Chl oooooooooooo 0mm0m00mm0mmm AUUACAACUGUCC 502 16-13997 o CTGF-1176- 13998 Chl oooooooooooo m0mm0m00mm0mm CAUUACAACUGUC 503 16-13998 o CTGF-812- 13999 Chl oooooooooooo 0000m0000mmm AGAGUGGAGCGCC 504 16-13999 o CTGF-745- 14000 Chl oooooooooooo 0mm0mm000000 ACCGACUGGAAGA 505 16-14000 o CTGF-1230- 14001 Chl oooooooooooo 0m0m0m0000m0 AUGUACGGAGACA 506 16-14001 o CTGF-920- 14002 Chl oooooooooooo 0mmmm0m000mm GCCUUGCGAAGCU 507 16-14002 o CTGF-679- 14003 Chl oooooooooooo 0mm0m00000m0 GCUGCGAGGAGUG 508 16-14003 o CTGF-992- 14004 Chl oooooooooooo 0mmm0mm000mmm GCCUAUCAAGUUU 509 16-14004 o CTGF-1045- 14005 Chl oooooooooooo 00mmmm0m0000m AAUUCUGUGGAGU 510 16-14005 o CTGF-1231- 14006 Chl oooooooooooo m0m0m0000m0m UGUACGGAGACAU 511 16-14006 o CTGF-991- 14007 Chl oooooooooooo 00mmm0mm000mm AGCCUAUCAAGUU 512 16-14007 o CTGF-998- 14008 Chl oooooooooooo m000mmm000mmm CAAGUUUGAGCUU 513 16-14008 o CTGF-1049- 14009 Chl oooooooooooo mm0m0000m0m0m CUGUGGAGUAUGU 514 16-14009 o CTGF-1044- 14010 Chl oooooooooooo 000mmmm0m0000 AAAUUCUGUGGAG 515 16-14010 o CTGF-1327- 14011 Chl oooooooooooo mmmm00m00m0m0 UUUCAGUAGCACA 516 16-14011 o CTGF-1196- 14012 Chl oooooooooooo m00m00m0mmmmm CAAUGACAUCUUU 517 16-14012 o CTGF-562- 14013 Chl oooooooooooo 00m0mm00m0m0m AGUACCAGUGCAC 518 16-14013 o CTGF-752- 14014 Chl oooooooooooo 000000m0mmmm GGAAGACACGUUU 519 16-14014 o CTGF-994- 14015 Chl oooooooooooo mm0mm000mmm00 CUAUCAAGUUUGA 520 16-14015 o CTGF-1040- 14016 Chl oooooooooooo 00mm000mmmm0m AGCUAAAUUCUGU 521 16-14016 o CTGF-1984- 14017 Chl oooooooooooo 000m0000m0m00 AGGUAGAAUGUAA 522 16-14017 o CTGF-2195- 14018 Chl oooooooooooo 00mm00mm00mmm AGCUGAUCAGUUU 523 16-14018 o CTGF-2043- 14019 Chl oooooooooooo mmmm0mmm000m0 UUCUGCUCAGAUA 524 16-14019 o CTGF-1892- 14020 Chl oooooooooooo mm0mmm000mm00 UUAUCUAAGUUAA 525 16-14020 o CTGF-1567- 14021 Chl oooooooooooo m0m0m00m00m0 UAUACGAGUAAUA 526 16-14021 o CTGF-1780- 14022 Chl oooooooooooo 00mm000m00mmm GACUGGACAGCUU 527 16-14022 o CTGF-2162- 14023 Chl oooooooooooo 0m00mmmmm0mm0 AUGGCCUUUAUUA 528 16-14023 o CTGF-1034- 14024 Chl oooooooooooo 0m0mm00mm000 AUACCGAGCUAAA 529 16-14024 o CTGF-2264- 14025 Chl oooooooooooo mm0mm00000m0m UUGUUGAGAGUGU 530 16-14025 o CTGF-1032- 14026 Chl oooooooooooo 0m0m0mm00mm0 ACAUACCGAGCUA 531 16-14026 o CTGF-1535- 14027 Chl oooooooooooo 00m0000000mm0 AGCAGAAAGGUUA 532 16-14027 o CTGF-1694- 14028 Chl oooooooooooo 00mm0mmmmmm00 AGUUGUUCCUUAA 533 16-14028 o CTGF-1588- 14029 Chl oooooooooooo 0mmm0000m0m00 AUUUGAAGUGUAA 534 16-14029 o CTGF-928- 14030 Chl oooooooooooo 000mm00mmm000 AAGCUGACCUGGA 535 16-14030 o CTGF-1133- 14031 Chl oooooooooooo 00mm0m0000000 GGUCAUGAAGAAG 536 16-14031 o CTGF-912- 14032 Chl oooooooooooo 0m00mm000mmmm AUGGUCAGGCCUU 537 16-14032 o CTGF-753- 14033 Chl oooooooooooo 00000m0mmmm0 GAAGACACGUUUG 538 16-14033 o CTGF-918- 14034 Chl oooooooooooo 000mmmm0m000 AGGCCUUGCGAAG 539 16-14034 o CTGF-744- 14035 Chl oooooooooooo m0mm0mm00000 UACCGACUGGAAG 540 16-14035 o CTGF-466- 14036 Chl oooooooooooo 0mmm0000mm0 ACCGCAAGAUCGG 541 16-14036 o CTGF-917- 14037 Chl oooooooooooo m000mmmm0m00 CAGGCCUUGCGAA 542 16-14037 o CTGF-1038- 14038 Chl oooooooooooo m00mm000mmmm CGAGCUAAAUUCU 543 16-14038 o CTGF-1048- 14039 Chl oooooooooooo mmm0m0000m0m0 UCUGUGGAGUAUG 544 16-14039 o CTGF-1235- 14040 Chl oooooooooooo m0000m0m00m0 CGGAGACAUGGCA 545 16-14040 o CTGF-868- 14041 Chl oooooooooooo 0m00m00mmmmm AUGACAACGCCUC 546 16-14041 o CTGF-1131- 14042 Chl oooooooooooo 0000mm0m00000 GAGGUCAUGAAGA 547 16-14042 o CTGF-1043- 14043 Chl oooooooooooo m000mmmm0m000 UAAAUUCUGUGGA 548 16-14043 o CTGF-751- 14044 Chl oooooooooooo m000000m0mmm UGGAAGACACGUU 549 16-14044 o CTGF-1227- 14045 Chl oooooooooooo 0000m0m0m000 AAGAUGUACGGAG 550 16-14045 o CTGF-867- 14046 Chl oooooooooooo 00m00m00mmmm AAUGACAACGCCU 551 16-14046 o CTGF-1128- 14047 Chl oooooooooooo 00m000mm0m00 GGCGAGGUCAUGA 552 16-14047 o CTGF-756- 14048 Chl oooooooooooo 00m0m0mmm00mm GACACGUUUGGCC 553 16-14048 o CTGF-1234- 14049 Chl oooooooooooo 0m00000m0m00m ACGGAGACAUGGC 554 16-14049 o CTGF-916- 14050 Chl oooooooooooo mm000mmmm0m00 UCAGGCCUUGCGA 555 16-14050 o CTGF-925- 14051 Chl oooooooooooo 0m0000mm00mmm GCGAAGCUGACCU 556 16-14051 o CTGF-1225- 14052 Chl oooooooooooo 000000m0m0m00 GGAAGAUGUACGG 557 16-14052 o CTGF-445- 14053 Chl oooooooooooo 0m00mmmm00mmm GUGACUUCGGCUC 558 16-14053 o CTGF-446- 14054 Chl oooooooooooo m00mmmm00mmmm UGACUUCGGCUCC 559 16-14054 o CTGF-913- 14055 Chl oooooooooooo m00mm000mmmm0 UGGUCAGGCCUUG 560 16-14055 o CTGF-997- 14056 Chl oooooooooooo mm000mmm000mm UCAAGUUUGAGCU 561 16-14056 o CTGF-277- 14057 Chl oooooooooooo 0mm0000mm0m00 GCCAGAACUGCAG 562 16-14057 o CTGF-1052- 14058 Chl oooooooooooo m0000m0m0m0mm UGGAGUAUGUACC 563 16-14058 o CTGF-887- 14059 Chl oooooooooooo 0mm0000000m00 GCUAGAGAAGCAG 564 16-14059 o CTGF-914- 14060 Chl oooooooooooo 00mm000mmmm0m GGUCAGGCCUUGC 565 16-14060 O CTGF-1039- 14061 Chl oooooooooooo 000mm000mmmm0 GAGCUAAAUUCUG 566 16-14061 o CTGF-754- 14062 Chl oooooooooooo 0000m0m0mmm00 AAGACACGUUUGG 567 16-14062 o CTGF-1130- 14063 Chl oooooooooooo m0000mm0m0000 CGAGGUCAUGAAG 568 16-14063 o CTGF-919- 14064 Chl oooooooooooo 00mmmm0m0000m GGCCUUGCGAAGC 569 16-14064 o CTGF-922- 14065 Chl oooooooooooo mmm0m0000mm00 CUUGCGAAGCUGA 570 16-14065 o CTGF-746- 14066 Chl oooooooooooo mm00mm000000m CCGACUGGAAGAC 571 16-14066 o CTGF-993- 14067 Chl oooooooooooo mmm0mm000mmm0 CCUAUCAAGUUUG 572 16-14067 o CTGF-825- 14068 Chl oooooooooooo m0mmmm0000mmm UGUUCCAAGACCU 573 16-14068 o CTGF-926- 14069 Chl oooooooooooo m0000mm00mmm0 CGAAGCUGACCUG 574 16-14069 o CTGF-923- 14070 Chl oooooooooooo mm0m0000mm00m UUGCGAAGCUGAC 575 16-14070 o CTGF-866- 14071 Chl oooooooooooo m00m00m00m0mm CAAUGACAACGCC 576 16-14071 o CTGF-563- 14072 Chl oooooooooooo 0m0mm00m0m0m0 GUACCAGUGCACG 577 16-14072 o CTGF-823- 14073 Chl oooooooooooo mmm0mmmm0000m CCUGUUCCAAGAC 578 16-14073 o CTGF-1233- 14074 Chl oooooooooooo m0m00000m0m00 UACGGAGACAUGG 579 16-14074 o CTGF-924- 14075 Chl oooooooooooo m0m0000mm00mm UGCGAAGCUGACC 580 16-14075 o CTGF-921- 14076 Chl oooooooooooo mmmm0m0000mm0 CCUUGCGAAGCUG 581 16-14076 o CTGF-443- 14077 Chl oooooooooooo mm0m00mmmm00m CUGUGACUUCGGC 582 16-14077 o CTGF-1041- 14078 Chl oooooooooooo 0mm000mmmm0m0 GCUAAAUUCUGUG 583 16-14078 o
CTGF-1042- 14079 Chl oooooooooooo mm000mmmm0m00 CUAAAUUCUGUGG 584 16-14079 o CTGF-755- 14080 Chl oooooooooooo 000m0m0mmm00m AGACACGUUUGGC 585 16-14080 o CTGF-467- 14081 Chl oooooooooooo mm0m0000mm00m CCGCAAGAUCGGC 586 16-14081 o CTGF-995- 14082 Chl oooooooooooo m0mm000mmm000 UAUCAAGUUUGAG 587 16-14082 o CTGF-927- 14083 Chl oooooooooooo 0000mm00mmm00 GAAGCUGACCUGG 588 16-14083 o SPP1-1091- 14131 Chl oooooooooooo 0m0mmm00mm000 GCAUUUAGUCAAA 589 16-14131 o PPIB--16- 14188 Chl oooooooooooo mmmmmmmmmmmmm GGCUACAAAAACA 590 14188 o PPIB--17- 14189 Chl oooooooooooo mm00mm0m00000 UUGGCUACAAAAA 591 14189 ooo m0 CA PPIB--18- 14190 Chl oooooooooooo 0mmm00mm0m000 AUUUGGCUACAAA 592 14190 ooooo 00m0 AACA pGL3-1172- 14386 chl oooooooooooo 0m000m0m00mmm ACAAAUACGAUUU 593 16-14386 o pGL3-1172- 14387 chl oooooooooooo DY5470m000m0m ACAAAUACGAUUU 594 16-14387 o 00mmm MAP4K4- 14390 Chl oooooooooooo Pmmmmmmmmmmmm CUUUGAAGAGUUC 595 2931-25- oooooooooooo 000mmmmmmmmmm UGUGGAAGUCUA 14390 o miR-122-- 14391 Chl ssoooooooooo mmmmmmmmmmmmm ACAAACACCAUUG 596 23-14391 ooooooossss mmmmmmmmmm UCACACUCCA 14084 Chl oooooooooooo mmm0m000mm000 CUCAUGAAUUAGA 719 o 14085 Chl oooooooooooo mm0000mm00mm0 CUGAGGUCAAUUA 720 o 14086 Chl oooooooooooo 0000mm00mm000 GAGGUCAAUUAAA 721 o 14087 Chl oooooooooooo mmm0000mm00mm UCUGAGGUCAAUU 722 o 14088 Chl oooooooooooo m0000mm00mm00 UGAGGUCAAUUAA 723 o 14089 Chl oooooooooooo mmmm0000mm00m UUCUGAGGUCAAU 724 o 14090 Chl oooooooooooo 0mm00mm000m00 GUCAGCUGGAUGA 725 o 14091 Chl oooooooooooo mmmm00m000mmm UUCUGAUGAAUCU 726 o 14092 Chl oooooooooooo m000mm0000mm0 UGGACUGAGGUCA 727 o 14093 Chl oooooooooooo 000mmmm0mm0mm GAGUCUCACCAUU 728 o 14094 Chl oooooooooooo 00mm0000mm000 GACUGAGGUCAAA 729 o 14095 Chl oooooooooooo mm0m00mm0m000 UCACAGCCAUGAA 730 o 14096 Chl oooooooooooo 00mmmm0mm0mmm AGUCUCACCAUUC 731 o 14097 Chl oooooooooooo 000m00000mm0 AAGCGGAAAGCCA 732 o 14098 Chl oooooooooooo 00m00000mm00 AGCGGAAAGCCAA 733 o 14099 Chl oooooooooooo 0mm0m0m000m00 ACCACAUGGAUGA 734 o 14100 Chl oooooooooooo 0mm0m00mm0m0m GCCAUGACCACAU 735 o 14101 Chl oooooooooooo 000mm0m00mm0m AAGCCAUGACCAC 736 o 14102 Chl oooooooooooo 0m00000mm00m GCGGAAAGCCAAU 737 o 14103 Chl oooooooooooo 000mmmmm0mmm AAAUUUCGUAUUU 738 o 14104 Chl oooooooooooo 0mmmmm0mmmmm AUUUCGUAUUUCU 739 o 14105 Chl oooooooooooo 0000mm0m00mm0 AAAGCCAUGACCA 740 o 14106 Chl oooooooooooo 0m0m000m00m0m ACAUGGAUGAUAU 741 o 14107 Chl oooooooooooo 0000mmmmm0mm GAAAUUUCGUAUU 742 o 14108 Chl oooooooooooo 0mmmmmmm00mm GCGCCUUCUGAUU 743 o 14109 Chl oooooooooooo 0mmmmmm0m000m AUUUCUCAUGAAU 744 o 14110 Chl oooooooooooo mmmmm0m000m00 CUCUCAUGAAUAG 745 o 14111 Chl oooooooooooo 000mmm00m000 AAGUCCAACGAAA 746 o 14112 Chl oooooooooooo 0m00m00000m00 AUGAUGAGAGCAA 747 o 14113 Chl oooooooooooo 0m00000mm000 GCGAGGAGUUGAA 748 o 14114 Chl oooooooooooo m00mm00m00mm0 UGAUUGAUAGUCA 749 o 14115 Chl oooooooooooo 000m00m0m0mmm AGAUAGUGCAUCU 750 o 14116 Chl oooooooooooo 0m0m0m0mmm0mm AUGUGUAUCUAUU 751 o 14117 Chl oooooooooooo mmmm0m0000000 UUCUAUAGAAGAA 752 o 14118 Chl oooooooooooo mm0mmm00m00mm UUGUCCAGCAAUU 753 o 14119 Chl oooooooooooo 0m0m000000m0 ACAUGGAAAGCGA 754 o 14120 Chl oooooooooooo 0m00mmm000mm0 GCAGUCCAGAUUA 755 o 14121 Chl oooooooooooo m00mm000m0m0m UGGUUGAAUGUGU 756 o 14122 Chl oooooooooooo mm0m0000m00m UUAUGAAACGAGU 757 o 14123 Chl oooooooooooo m00mmm000mm0m CAGUCCAGAUUAU 758 o 14124 Chl oooooooooooo 0m0m000m0000 AUAUAAGCGGAAA 759 o 14125 Chl oooooooooooo m0mm00mm000m0 UACCAGUUAAACA 760 o 14126 Chl oooooooooooo m0mmm0mmmm0m0 UGUUCAUUCUAUA 761 o 14127 Chl oooooooooooo mm0mm0000000 CCGACCAAGGAAA 762 o 14128 Chl oooooooooooo 000m00m0m0m0m GAAUGGUGCAUAC 763 o 14129 Chl oooooooooooo 0m0m00m00mm0 AUAUGAUGGCCGA 764 o 14130 Chl oooooooooooo 00m00mmm000mm AGCAGUCCAGAUU 765 o 14132 Chl oooooooooooo 00m0mmmm0m0m AGCAUUCCGAUGU 766 o 14133 Chl oooooooooooo m00mm00000mmm UAGUCAGGAACUU 767 o 14134 Chl oooooooooooo m0m0mmm00mm00 UGCAUUUAGUCAA 768 o 14135 Chl oooooooooooo 0mmm00m000mmm GUCUGAUGAGUCU 769 o 14136 Chl oooooooooooo m000m0m0m0m00 UAGACACAUAUGA 770 o 14137 Chl oooooooooooo m000m0000m0m CAGACGAGGACAU 771 o 14138 Chl oooooooooooo m00mmm000mmm CAGCCGUGAAUUC 772 o 14139 Chl oooooooooooo 00mmm00000m00 AGUCUGGAAAUAA 773 o 14140 Chl oooooooooooo 00mmm0m00mmmm AGUUUGUGGCUUC 774 o 14141 Chl oooooooooooo 00mmm00m0000 AGUCCAACGAAAG 775 o 14142 Chl oooooooooooo 000mmmmm000m AAGUUUCGCAGAC 776 o 14143 Chl oooooooooooo 00m00m000m0mm AGCAAUGAGCAUU 777 o 14144 Chl oooooooooooo mm000m00m0m0m UUAGAUAGUGCAU 778 o 14145 Chl oooooooooooo m00m0m0m0m000 UGGUGCAUACAAG 779 o 14146 Chl oooooooooooo 0m0000m00mm0 AUGAAACGAGUCA 780 o 14147 Chl oooooooooooo mm0000m0mm000 CCAGAGUGCUGAA 781 o 14148 Chl oooooooooooo m00mm0m000mmm CAGCCAUGAAUUU 782 o 14149 Chl oooooooooooo 0mm00mm000m0m AUUGGUUGAAUGU 783 o 14150 Chl oooooooooooo 00mm000m0m0m0 GGUUGAAUGUGUA 784 o 14151 Chl oooooooooooo 00000m00mm00m GGAAAUAACUAAU 785 o 14152 Chl oooooooooooo mm0m000m00000 UCAUGAAUAGAAA 786 o 14153 Chl oooooooooooo 0mm00m00mm00 GCCAGCAACCGAA 787 o 14154 Chl oooooooooooo m0mmmm0m0m0m0 CACCUCACACAUG 788 o
14155 Chl oooooooooooo 00mm000m00m0m AGUUGAAUGGUGC 789 o 14156 Chl oooooooooooo 00mm00mm000m0 AGUCAGCUGGAUG 790 o 14157 Chl oooooooooooo m0m000m00000 UAUAAGCGGAAAG 791 o 14158 Chl oooooooooooo mmmm0m0m00mm UUCCGAUGUGAUU 792 o 14159 Chl oooooooooooo 0m00mm00m0m0m AUAACUAAUGUGU 793 o 14160 Chl oooooooooooo mm0mmmm0m0000 UCAUUCUAUAGAA 794 o 14161 Chl oooooooooooo 00mm0mm0mm0m0 AACUAUCACUGUA 795 o 14162 Chl oooooooooooo 0mm00mm0mmm0m GUCAAUUGCUUAU 796 o 14163 Chl oooooooooooo 00m00mm00m000 AGCAAUUAAUAAA 797 o 14164 Chl oooooooooooo 0m0mmmm00m00 ACGACUCUGAUGA 798 o 14165 Chl oooooooooooo m00m0m00mmm0m UAGUGUGGUUUAU 799 o 14166 Chl oooooooooooo 000mm00m00m00 AAGCCAAUGAUGA 800 o 14167 Chl oooooooooooo 0m00mm00000mm AUAGUCAGGAACU 801 o 14168 Chl oooooooooooo 00mm00mmm000 AGUCAGCCGUGAA 802 o 14169 Chl oooooooooooo 0mm0mm0m00000 ACUACCAUGAGAA 803 o 14170 Chl oooooooooooo 000m000mm00mm AAACAGGCUGAUU 804 o 14171 Chl oooooooooooo 000m0mm0000mm GAGUGCUGAAACC 805 o 14172 Chl oooooooooooo m000m0mmmm0m UGAGCAUUCCGAU 806 o 14173 Chl oooooooooooo 00mmmm0m00mm0 AAUUCCACAGCCA 807 o 14174 Chl oooooooooooo m0mm00mm0mmm0 UGUCAAUUGCUUA 808 o 14175 Chl oooooooooooo 0mm0m00000mm0 ACCAUGAGAAUUG 809 o 14176 Chl oooooooooooo mm00m0000mm0 CCAACGAAAGCCA 810 o 14177 Chl oooooooooooo mm00mm0mm00mm CUGGUCACUGAUU 811 o 14178 Chl oooooooooooo m00mmm0m000mm UGGUUUAUGGACU 812 o 14179 Chl oooooooooooo 00mm0000m0mm0 GACCAGAGUGCUG 813 o 14180 Chl oooooooooooo 00m0m00mm00m0 GAUGUGAUUGAUA 814 o 14181 Chl oooooooooooo 0mm00mmm000m GUCAGCCGUGAAU 815 o 14182 Chl oooooooooooo 00m0m0m0mmm0m AAUGUGUAUCUAU 816 o 14183 Chl oooooooooooo mm000mmm00000 UUGAGUCUGGAAA 817 o 14184 Chl oooooooooooo 0mmm00m00mm00 GUCCAGCAAUUAA 818 o 14185 Chl oooooooooooo mm00m00mm00m0 CCAGCAAUUAAUA 819 o 14186 Chl oooooooooooo 00mmm00m0mm GACUCGAACGACU 820 o 14187 Chl oooooooooooo 0mmm0mm00m00m ACCUGCCAGCAAC 821 o o: phosphodiester; s: phosphorothioate; P: 5' phosphorylation; 0: 2'-OH; F: 2'-fluoro; m: 2' O-methyl; +: LNA modification. Capital letters in the sequence signify ribonucleotides, lower case letters signify deoxyribonucleotides.
[0454] Having thus described several aspects of at least one embodiment of this invention, it is to be appreciated various alterations, modifications, and improvements will readily occur to those skilled in the art. Such alterations, modifications, and improvements are intended to be part of this disclosure, and are intended to be within the spirit and scope of the invention. Accordingly, the foregoing description and drawings are by way of example only.
EQUIVALENTS
[0455] Those skilled in the art will recognize, or be able to ascertain using no more than routine experimentation, many equivalents to the specific embodiments of the invention described herein. Such equivalents are intended to be encompassed by the following claims.
[0456] All references, including patent documents, disclosed herein are incorporated by reference in their entirety. This application incorporates by reference the entire contents, including all the drawings and all parts of the specification (including sequence listing or amino acid/polynucleotide sequences) of the co-pending U.S. Provisional Application No. 61/135,855, filed on Jul. 24, 2008, entitled "SHORT HAIRPIN RNAI CONSTRUCTS AND USES THEREOF," and U.S. Provisional Application No. 61/197,768, filed on Oct. 30, 2008, entitled "MINIRNA CONSTRUCTS AND USES THEREOF."
Sequence CWU 1
1
821119RNAArtificial Sequencesynthetic oligonucleotide 1auugguauuc
agugugaug 19219RNAArtificial Sequencesynthetic oligonucleotide
2auucguauug agucugauc 19319RNAArtificial Sequencesynthetic
oligonucleotide 3uagacuucca cagaacucu 19419RNAArtificial
Sequencesynthetic oligonucleotide 4uagacuucca cagaacucu
19519RNAArtificial Sequencesynthetic oligonucleotide 5uagacuucca
cagaacucu 19619RNAArtificial Sequencesynthetic oligonucleotide
6uagacuucca cagaacucu 19719RNAArtificial Sequencesynthetic
oligonucleotide 7uagacuucca cagaacucu 19819RNAArtificial
Sequencesynthetic oligonucleotide 8uagacuucca cagaacucu
19917RNAArtificial Sequencesynthetic oligonucleotide 9uagacuucca
cagaacu 171021RNAArtificial Sequencesynthetic oligonucleotide
10auugguauuc agugugauga c 211121RNAArtificial Sequencesynthetic
oligonucleotide 11auucguauug agucugauca c 211219RNAArtificial
Sequencesynthetic oligonucleotide 12uagacuucca cagaacucu
191319RNAArtificial Sequencesynthetic oligonucleotide 13uagacuucca
cagaacucu 191419RNAArtificial Sequencesynthetic oligonucleotide
14uagacuucca cagaacucu 191521RNAArtificial Sequencesynthetic
oligonucleotide 15uagacuucca cagaacucuu c 211621RNAArtificial
Sequencesynthetic oligonucleotide 16uagacuucca cagaacucuu c
211725RNAArtificial Sequencesynthetic oligonucleotide 17uagacuucca
cagaacucuu caaag 251821RNAArtificial Sequencesynthetic
oligonucleotide 18auugguauuc agugugauga c 211921RNAArtificial
Sequencesynthetic oligonucleotide 19auucguauug agucugauca c
212019RNAArtificial Sequencesynthetic oligonucleotide 20uagacuucca
cagaacucu 192121RNAArtificial Sequencesynthetic oligonucleotide
21auugguauuc agugugauga c 212221RNAArtificial Sequencesynthetic
oligonucleotide 22auucguauug agucugauca c 212319RNAArtificial
Sequencesynthetic oligonucleotide 23uagacuucca cagaacucu
192419RNAArtificial Sequencesynthetic oligonucleotide 24uagacuucca
cagaacucu 192519RNAArtificial Sequencesynthetic oligonucleotide
25uagacuucca cagaacucu 192619RNAArtificial Sequencesynthetic
oligonucleotide 26uagacuucca cagaacucu 192719RNAArtificial
Sequencesynthetic oligonucleotide 27uagacuucca cagaacucu
192819RNAArtificial Sequencesynthetic oligonucleotide 28uagacuucca
cagaacucu 192919RNAArtificial Sequencesynthetic oligonucleotide
29uagacuucca cagaacucu 193019RNAArtificial Sequencesynthetic
oligonucleotide 30uagacuucca cagaacucu 193119RNAArtificial
Sequencesynthetic oligonucleotide 31uagacuucca cagaacucu
193219RNAArtificial Sequencesynthetic oligonucleotide 32uagacuucca
cagaacucu 193319RNAArtificial Sequencesynthetic oligonucleotide
33uagacuucca cagaacucu 193419RNAArtificial Sequencesynthetic
oligonucleotide 34uagacuucca cagaacucu 193519RNAArtificial
Sequencesynthetic oligonucleotide 35uagacuucca cagaacucu
193619RNAArtificial Sequencesynthetic oligonucleotide 36uagacuucca
cagaacucu 193719RNAArtificial Sequencesynthetic oligonucleotide
37uagacuucca cagaacucu 193819RNAArtificial Sequencesynthetic
oligonucleotide 38uguuuuugua gccaaaucc 193919RNAArtificial
Sequencesynthetic oligonucleotide 39uacuuucuuc auuuccacc
194019RNAArtificial Sequencesynthetic oligonucleotide 40uucauuucca
ccuuugccc 194119RNAArtificial Sequencesynthetic oligonucleotide
41cuuuguacuu ucuucauuu 194219RNAArtificial Sequencesynthetic
oligonucleotide 42ucuuuguacu uucuucauu 194319RNAArtificial
Sequencesynthetic oligonucleotide 43ucagcaguca cauugccca
194419RNAArtificial Sequencesynthetic oligonucleotide 44auugcccaag
ucuccaaca 194519RNAArtificial Sequencesynthetic oligonucleotide
45uucugcucga aauugauga 194619RNAArtificial Sequencesynthetic
oligonucleotide 46aaaucguauu ugucaauca 194719RNAArtificial
Sequencesynthetic oligonucleotide 47aaaucguauu ugucaauca
194819RNAArtificial Sequencesynthetic oligonucleotide 48uagacuucca
cagaacucu 194919RNAArtificial Sequencesynthetic oligonucleotide
49uagacuucca cagaacucu 195019RNAArtificial Sequencesynthetic
oligonucleotide 50uagacauccu acacagcac 195119RNAArtificial
Sequencesynthetic oligonucleotide 51uguuucucca gauccuugc
195219RNAArtificial Sequencesynthetic oligonucleotide 52uguuucucca
gauccuugc 195319RNAArtificial Sequencesynthetic oligonucleotide
53uagcagauga guccauuug 195423RNAArtificial Sequencesynthetic
oligonucleotide 54uagcagauga guccauuugg aga 235519RNAArtificial
Sequencesynthetic oligonucleotide 55uagcagauga guccauuug
195623RNAArtificial Sequencesynthetic oligonucleotide 56uagcagauga
guccauuugg aga 235719RNAArtificial Sequencesynthetic
oligonucleotide 57auguuguuuc uccagaucc 195819RNAArtificial
Sequencesynthetic oligonucleotide 58auguuguuuc uccagaucc
195919RNAArtificial Sequencesynthetic oligonucleotide 59uguuugaggg
acucuguga 196019RNAArtificial Sequencesynthetic oligonucleotide
60uguuugaggg acucuguga 196119RNAArtificial Sequencesynthetic
oligonucleotide 61auugguauuc agugugaug 196223RNAArtificial
Sequencesynthetic oligonucleotide 62auugguauuc agugugauga cac
236319RNAArtificial Sequencesynthetic oligonucleotide 63auugguauuc
agugugaug 196423RNAArtificial Sequencesynthetic oligonucleotide
64auugguauuc agugugauga cac 236519RNAArtificial Sequencesynthetic
oligonucleotide 65uagacuucca cagaacucu 196619RNAArtificial
Sequencesynthetic oligonucleotide 66uacaucuucc uguaguaca
196719RNAArtificial Sequencesynthetic oligonucleotide 67aggcgcucca
cucuguggu 196819RNAArtificial Sequencesynthetic oligonucleotide
68ugucuuccag ucgguaagc 196919RNAArtificial Sequencesynthetic
oligonucleotide 69gaacaggcgc uccacucug 197019RNAArtificial
Sequencesynthetic oligonucleotide 70caguuguaau ggcaggcac
197119RNAArtificial Sequencesynthetic oligonucleotide 71agccagaaag
cucaaacuu 197219RNAArtificial Sequencesynthetic oligonucleotide
72caggcgcucc acucugugg 197319RNAArtificial Sequencesynthetic
oligonucleotide 73aacaggcgcu ccacucugu 197419RNAArtificial
Sequencesynthetic oligonucleotide 74agaaagcuca aacuugaua
197519RNAArtificial Sequencesynthetic oligonucleotide 75aguuguaaug
gcaggcaca 197619RNAArtificial Sequencesynthetic oligonucleotide
76cgugucuucc agucgguaa 197719RNAArtificial Sequencesynthetic
oligonucleotide 77ggaccaggca guuggcucu 197819RNAArtificial
Sequencesynthetic oligonucleotide 78caggcacagg ucuugauga
197919RNAArtificial Sequencesynthetic oligonucleotide 79gcgcuccacu
cuguggucu 198019RNAArtificial Sequencesynthetic oligonucleotide
80ggucuggacc aggcaguug 198119RNAArtificial Sequencesynthetic
oligonucleotide 81acaguuguaa uggcaggca 198219RNAArtificial
Sequencesynthetic oligonucleotide 82guuguaaugg caggcacag
198319RNAArtificial Sequencesynthetic oligonucleotide 83ggacaguugu
aauggcagg 198419RNAArtificial Sequencesynthetic oligonucleotide
84gacaguugua auggcaggc 198519RNAArtificial Sequencesynthetic
oligonucleotide 85ggcgcuccac ucugugguc 198619RNAArtificial
Sequencesynthetic oligonucleotide 86ucuuccaguc gguaagccg
198719RNAArtificial Sequencesynthetic oligonucleotide 87ugucuccgua
caucuuccu 198819RNAArtificial Sequencesynthetic oligonucleotide
88agcuucgcaa ggccugacc 198919RNAArtificial Sequencesynthetic
oligonucleotide 89cacuccucgc agcauuucc 199019RNAArtificial
Sequencesynthetic oligonucleotide 90aaacuugaua ggcuuggag
199119RNAArtificial Sequencesynthetic oligonucleotide 91acuccacaga
auuuagcuc 199219RNAArtificial Sequencesynthetic oligonucleotide
92augucuccgu acaucuucc 199319RNAArtificial Sequencesynthetic
oligonucleotide 93aacuugauag gcuuggaga 199419RNAArtificial
Sequencesynthetic oligonucleotide 94aagcucaaac uugauaggc
199519RNAArtificial Sequencesynthetic oligonucleotide 95acauacucca
cagaauuua 199619RNAArtificial Sequencesynthetic oligonucleotide
96cuccacagaa uuuagcucg 199719RNAArtificial Sequencesynthetic
oligonucleotide 97ugugcuacug aaaucauuu 199819RNAArtificial
Sequencesynthetic oligonucleotide 98aaagauguca uugucuccg
199919RNAArtificial Sequencesynthetic oligonucleotide 99gugcacuggu
acuugcagc 1910019RNAArtificial Sequencesynthetic oligonucleotide
100aaacgugucu uccagucgg 1910119RNAArtificial Sequencesynthetic
oligonucleotide 101ucaaacuuga uaggcuugg 1910219RNAArtificial
Sequencesynthetic oligonucleotide 102acagaauuua gcucgguau
1910319RNAArtificial Sequencesynthetic oligonucleotide
103uuacauucua ccuauggug 1910419RNAArtificial Sequencesynthetic
oligonucleotide 104aaacugauca gcuauauag 1910519RNAArtificial
Sequencesynthetic oligonucleotide 105uaucugagca gaauuucca
1910619RNAArtificial Sequencesynthetic oligonucleotide
106uuaacuuaga uaacuguac 1910719RNAArtificial Sequencesynthetic
oligonucleotide 107uauuacucgu auaagaugc 1910819RNAArtificial
Sequencesynthetic oligonucleotide 108aagcugucca gucuaaucg
1910919RNAArtificial Sequencesynthetic oligonucleotide
109uaauaaaggc cauuuguuc 1911019RNAArtificial Sequencesynthetic
oligonucleotide 110uuuagcucgg uaugucuuc 1911119RNAArtificial
Sequencesynthetic oligonucleotide 111acacucucaa caaauaaac
1911219RNAArtificial Sequencesynthetic oligonucleotide
112uagcucggua ugucuucau 1911319RNAArtificial Sequencesynthetic
oligonucleotide 113uaaccuuucu gcugguacc 1911419RNAArtificial
Sequencesynthetic oligonucleotide 114uuaaggaaca acuugacuc
1911519RNAArtificial Sequencesynthetic oligonucleotide
115uuacacuuca aauagcagg 1911619RNAArtificial Sequencesynthetic
oligonucleotide 116uccaggucag cuucgcaag 1911719RNAArtificial
Sequencesynthetic oligonucleotide 117cuucuucaug accucgccg
1911819RNAArtificial Sequencesynthetic oligonucleotide
118aaggccugac caugcacag 1911919RNAArtificial Sequencesynthetic
oligonucleotide 119caaacguguc uuccagucg 1912019RNAArtificial
Sequencesynthetic oligonucleotide 120cuucgcaagg ccugaccau
1912119RNAArtificial Sequencesynthetic oligonucleotide
121cuuccagucg guaagccgc 1912219RNAArtificial Sequencesynthetic
oligonucleotide 122ccgaucuugc gguuggccg 1912319RNAArtificial
Sequencesynthetic oligonucleotide 123uucgcaaggc cugaccaug
1912419RNAArtificial Sequencesynthetic oligonucleotide
124agaauuuagc ucgguaugu 1912519RNAArtificial Sequencesynthetic
oligonucleotide 125cauacuccac agaauuuag 1912619RNAArtificial
Sequencesynthetic oligonucleotide 126ugccaugucu ccguacauc
1912719RNAArtificial Sequencesynthetic oligonucleotide
127gaggcguugu cauugguaa 1912819RNAArtificial Sequencesynthetic
oligonucleotide 128ucuucaugac cucgccguc 1912919RNAArtificial
Sequencesynthetic oligonucleotide 129uccacagaau uuagcucgg
1913019RNAArtificial Sequencesynthetic oligonucleotide
130aacgugucuu ccagucggu 1913119RNAArtificial Sequencesynthetic
oligonucleotide 131cuccguacau cuuccugua 1913219RNAArtificial
Sequencesynthetic oligonucleotide 132aggcguuguc auugguaac
1913319RNAArtificial Sequencesynthetic oligonucleotide
133ucaugaccuc gccgucagg 1913419RNAArtificial Sequencesynthetic
oligonucleotide 134ggccaaacgu gucuuccag 1913519RNAArtificial
Sequencesynthetic oligonucleotide 135gccaugucuc cguacaucu
1913619RNAArtificial Sequencesynthetic oligonucleotide
136ucgcaaggcc ugaccaugc 1913719RNAArtificial Sequencesynthetic
oligonucleotide 137aggucagcuu cgcaaggcc 1913819RNAArtificial
Sequencesynthetic oligonucleotide 138ccguacaucu uccuguagu
1913919RNAArtificial Sequencesynthetic oligonucleotide
139gagccgaagu cacagaaga 1914019RNAArtificial Sequencesynthetic
oligonucleotide 140ggagccgaag ucacagaag 1914119RNAArtificial
Sequencesynthetic oligonucleotide 141caaggccuga ccaugcaca
1914219RNAArtificial Sequencesynthetic oligonucleotide
142agcucaaacu ugauaggcu 1914319RNAArtificial Sequencesynthetic
oligonucleotide 143cugcaguucu ggccgacgg 1914419RNAArtificial
Sequencesynthetic oligonucleotide 144gguacauacu ccacagaau
1914519RNAArtificial Sequencesynthetic oligonucleotide
145cugcuucucu agccugcag 1914619RNAArtificial Sequencesynthetic
oligonucleotide 146gcaaggccug accaugcac 1914719RNAArtificial
Sequencesynthetic oligonucleotide 147cagaauuuag cucgguaug
1914819RNAArtificial Sequencesynthetic oligonucleotide
148ccaaacgugu cuuccaguc 1914919RNAArtificial Sequencesynthetic
oligonucleotide 149cuucaugacc ucgccguca 1915019RNAArtificial
Sequencesynthetic oligonucleotide 150gcuucgcaag gccugacca
1915119RNAArtificial Sequencesynthetic oligonucleotide
151ucagcuucgc aaggccuga 1915219RNAArtificial Sequencesynthetic
oligonucleotide 152gucuuccagu cgguaagcc 1915319RNAArtificial
Sequencesynthetic oligonucleotide 153caaacuugau aggcuugga
1915419RNAArtificial Sequencesynthetic oligonucleotide
154aggucuugga acaggcgcu 1915519RNAArtificial Sequencesynthetic
oligonucleotide 155caggucagcu ucgcaaggc 1915619RNAArtificial
Sequencesynthetic oligonucleotide 156gucagcuucg caaggccug
1915719RNAArtificial Sequencesynthetic oligonucleotide
157ggcguuguca uugguaacc 1915819RNAArtificial Sequencesynthetic
oligonucleotide 158cgugcacugg uacuugcag 1915919RNAArtificial
Sequencesynthetic oligonucleotide 159gucuuggaac aggcgcucc
1916019RNAArtificial Sequencesynthetic oligonucleotide
160ccaugucucc guacaucuu 1916119RNAArtificial Sequencesynthetic
oligonucleotide 161ggucagcuuc gcaaggccu 1916219RNAArtificial
Sequencesynthetic oligonucleotide 162cagcuucgca aggccugac
1916319RNAArtificial Sequencesynthetic oligonucleotide
163gccgaaguca cagaagagg 1916419RNAArtificial Sequencesynthetic
oligonucleotide 164cacagaauuu agcucggua 1916519RNAArtificial
Sequencesynthetic oligonucleotide 165ccacagaauu uagcucggu
1916619RNAArtificial Sequencesynthetic oligonucleotide
166gccaaacgug ucuuccagu 1916719RNAArtificial Sequencesynthetic
oligonucleotide 167gccgaucuug cgguuggcc 1916819RNAArtificial
Sequencesynthetic oligonucleotide 168cucaaacuug auaggcuug
1916919RNAArtificial Sequencesynthetic oligonucleotide
169ccaggucagc uucgcaagg 1917019RNAArtificial Sequencesynthetic
oligonucleotide 170uuugacuaaa ugcaaagug 1917119RNAArtificial
Sequencesynthetic oligonucleotide 171uguuuuugua gccaaaucc
1917219RNAArtificial Sequencesynthetic oligonucleotide
172uguuuuugua gccaaaucc 1917319RNAArtificial Sequencesynthetic
oligonucleotide 173uguuuuugua gccaaaucc 1917419RNAArtificial
Sequencesynthetic oligonucleotide 174aaaucguauu ugucaauca
1917519RNAArtificial Sequencesynthetic oligonucleotide
175aaaucguauu ugucaauca 1917621RNAArtificial Sequencesynthetic
oligonucleotide 176gucaucacac ugaauaccaa u 2117721RNAArtificial
Sequencesynthetic oligonucleotide 177gugaucagac ucaauacgaa u
2117813RNAArtificial Sequencesynthetic oligonucleotide
178cuguggaagu cua 1317913RNAArtificial Sequencesynthetic
oligonucleotide 179cuguggaagu cua 1318013RNAArtificial
Sequencesynthetic oligonucleotide 180cuguggaagu cua
1318113RNAArtificial Sequencesynthetic oligonucleotide
181cuguggaagu cua 1318213RNAArtificial Sequencesynthetic
oligonucleotide 182cuguggaagu cua 1318313RNAArtificial
Sequencesynthetic oligonucleotide 183cuguggaagu cua
1318413RNAArtificial Sequencesynthetic oligonucleotide
184cuguggaagu cua 1318513RNAArtificial Sequencesynthetic
oligonucleotide 185cuguggaagu cua 1318613RNAArtificial
Sequencesynthetic oligonucleotide 186cuguggaagu cua
1318713RNAArtificial Sequencesynthetic oligonucleotide
187cuguggaagu cua 1318821RNAArtificial Sequencesynthetic
oligonucleotide 188gucaucacac ugaauaccaa u 2118921RNAArtificial
Sequencesynthetic oligonucleotide 189gugaucagac ucaauacgaa u
2119013RNAArtificial Sequencesynthetic oligonucleotide
190cuguggaagu cua 1319113RNAArtificial Sequencesynthetic
oligonucleotide 191cuguggaagu cua 1319213RNAArtificial
Sequencesynthetic oligonucleotide 192cuguggaagu cua
1319313RNAArtificial Sequencesynthetic oligonucleotide
193cuguggaagu cua 1319413RNAArtificial Sequencesynthetic
oligonucleotide 194cuguggaagu cua 1319513RNAArtificial
Sequencesynthetic oligonucleotide 195cuguggaagu cua
1319627RNAArtificial Sequencesynthetic oligonucleotide
196ucauagguaa ccucugguug aaaguga 2719727RNAArtificial
Sequencesynthetic oligonucleotide 197cggcuacagg ugcuuaugaa gaaagua
2719821RNAArtificial Sequencesynthetic oligonucleotide
198gucaucacac ugaauaccaa u 2119921RNAArtificial Sequencesynthetic
oligonucleotide 199gugaucagac ucaauacgaa u 2120013RNAArtificial
Sequencesynthetic oligonucleotide 200cuguggaagu cua
1320121RNAArtificial Sequencesynthetic oligonucleotide
201gucaucacac ugaauaccaa u 2120221RNAArtificial Sequencesynthetic
oligonucleotide 202gugaucagac ucaauacgaa u 2120313RNAArtificial
Sequencesynthetic oligonucleotide 203uguaggaugu cua
1320413RNAArtificial Sequencesynthetic oligonucleotide
204cuguggaagu cua 1320513RNAArtificial Sequencesynthetic
oligonucleotide 205cuguggaagu cua 1320613RNAArtificial
Sequencesynthetic oligonucleotide 206acugaauacc aau
1320713RNAArtificial Sequencesynthetic oligonucleotide
207cuguggaagu cua 1320813RNAArtificial Sequencesynthetic
oligonucleotide 208cuguggaagu cua 1320913RNAArtificial
Sequencesynthetic oligonucleotide 209cuguggaagu cua
1321013RNAArtificial Sequencesynthetic oligonucleotide
210cuguggaagu cua 1321113RNAArtificial Sequencesynthetic
oligonucleotide 211cuguggaagu cua 1321213RNAArtificial
Sequencesynthetic oligonucleotide 212cuguggaagu cua
1321313RNAArtificial Sequencesynthetic oligonucleotide
213cuguggaagu cua 1321413RNAArtificial Sequencesynthetic
oligonucleotide 214cuguggaagu cua 1321513RNAArtificial
Sequencesynthetic oligonucleotide 215cuguggaagu cua
1321613RNAArtificial Sequencesynthetic oligonucleotide
216cuguggaagu cua 1321713RNAArtificial Sequencesynthetic
oligonucleotide 217cuguggaagu cua 1321813RNAArtificial
Sequencesynthetic oligonucleotide 218cuguggaagu cua
1321913RNAArtificial Sequencesynthetic oligonucleotide
219cuguggaagu cua 1322019RNAArtificial Sequencesynthetic
oligonucleotide 220agaguucugu ggaagucua 1922119RNAArtificial
Sequencesynthetic oligonucleotide 221agaguucugu ggaagucua
1922213RNAArtificial Sequencesynthetic oligonucleotide
222auuuggcuac aaa 1322313RNAArtificial Sequencesynthetic
oligonucleotide 223acaaauacga uuu 1322413RNAArtificial
Sequencesynthetic oligonucleotide 224acaaauacga uuu
1322513RNAArtificial Sequencesynthetic oligonucleotide
225cuguggaagu cua 1322613RNAArtificial Sequencesynthetic
oligonucleotide 226aaugaagaaa gua 1322713RNAArtificial
Sequencesynthetic oligonucleotide 227agguggaaau gaa
1322813RNAArtificial Sequencesynthetic oligonucleotide
228agaaaguaca aag 1322913RNAArtificial Sequencesynthetic
oligonucleotide 229gaaaguacaa aga 1323013RNAArtificial
Sequencesynthetic oligonucleotide 230augugacugc uga
1323113RNAArtificial Sequencesynthetic oligonucleotide
231agacuugggc aau 1323213RNAArtificial Sequencesynthetic
oligonucleotide 232auuucgagca gaa 1323313RNAArtificial
Sequencesynthetic oligonucleotide 233aucuggagaa aca
1323413RNAArtificial Sequencesynthetic oligonucleotide
234ucagaacaag aaa 1323513RNAArtificial Sequencesynthetic
oligonucleotide 235gacucaucug cua 1323613RNAArtificial
Sequencesynthetic oligonucleotide 236ggagaaacaa cau
1323713RNAArtificial Sequencesynthetic oligonucleotide
237agucccucaa aca 1323813RNAArtificial Sequencesynthetic
oligonucleotide 238ggcuacaaaa aca 1323915RNAArtificial
Sequencesynthetic oligonucleotide 239agaucuggag aaaca
1524015RNAArtificial Sequencesynthetic oligonucleotide
240uggacucauc ugcua 1524115RNAArtificial Sequencesynthetic
oligonucleotide 241cuggagaaac aacau 1524215RNAArtificial
Sequencesynthetic oligonucleotide 242agagucccuc aaaca
1524313RNAArtificial Sequencesynthetic oligonucleotide
243acugaauacc aau 1324415RNAArtificial Sequencesynthetic
oligonucleotide 244acacugaaua ccaau 1524513RNAArtificial
Sequencesynthetic oligonucleotide 245aaugaagaaa gua
1324613RNAArtificial Sequencesynthetic oligonucleotide
246agguggaaau gaa 1324713RNAArtificial Sequencesynthetic
oligonucleotide 247agaaaguaca aag 1324813RNAArtificial
Sequencesynthetic oligonucleotide 248gaaaguacaa aga
1324913RNAArtificial Sequencesynthetic oligonucleotide
249augugacugc uga 1325013RNAArtificial Sequencesynthetic
oligonucleotide 250agacuugggc aau 1325113RNAArtificial
Sequencesynthetic oligonucleotide 251auuucgagca gaa
1325213RNAArtificial Sequencesynthetic oligonucleotide
252acaaauacga uuu
1325313RNAArtificial Sequencesynthetic oligonucleotide
253acaaauacga uuu 1325419RNAArtificial Sequencesynthetic
oligonucleotide 254agaguucugu ggaagucua 1925513RNAArtificial
Sequencesynthetic oligonucleotide 255acaggaagau gua
1325613RNAArtificial Sequencesynthetic oligonucleotide
256gaguggagcg ccu 1325713RNAArtificial Sequencesynthetic
oligonucleotide 257cgacuggaag aca 1325813RNAArtificial
Sequencesynthetic oligonucleotide 258ggagcgccug uuc
1325913RNAArtificial Sequencesynthetic oligonucleotide
259gccauuacaa cug 1326013RNAArtificial Sequencesynthetic
oligonucleotide 260gagcuuucug gcu 1326113RNAArtificial
Sequencesynthetic oligonucleotide 261aguggagcgc cug
1326213RNAArtificial Sequencesynthetic oligonucleotide
262uggagcgccu guu 1326313RNAArtificial Sequencesynthetic
oligonucleotide 263guuugagcuu ucu 1326413RNAArtificial
Sequencesynthetic oligonucleotide 264ugccauuaca acu
1326513RNAArtificial Sequencesynthetic oligonucleotide
265acuggaagac acg 1326613RNAArtificial Sequencesynthetic
oligonucleotide 266aacugccugg ucc 1326713RNAArtificial
Sequencesynthetic oligonucleotide 267agaccugugc cug
1326813RNAArtificial Sequencesynthetic oligonucleotide
268cagaguggag cgc 1326913RNAArtificial Sequencesynthetic
oligonucleotide 269ccugguccag acc 1327013RNAArtificial
Sequencesynthetic oligonucleotide 270ccauuacaac ugu
1327113RNAArtificial Sequencesynthetic oligonucleotide
271cugccauuac aac 1327213RNAArtificial Sequencesynthetic
oligonucleotide 272auuacaacug ucc 1327313RNAArtificial
Sequencesynthetic oligonucleotide 273cauuacaacu guc
1327413RNAArtificial Sequencesynthetic oligonucleotide
274agaguggagc gcc 1327513RNAArtificial Sequencesynthetic
oligonucleotide 275accgacugga aga 1327613RNAArtificial
Sequencesynthetic oligonucleotide 276auguacggag aca
1327713RNAArtificial Sequencesynthetic oligonucleotide
277gccuugcgaa gcu 1327813RNAArtificial Sequencesynthetic
oligonucleotide 278gcugcgagga gug 1327913RNAArtificial
Sequencesynthetic oligonucleotide 279gccuaucaag uuu
1328013RNAArtificial Sequencesynthetic oligonucleotide
280aauucugugg agu 1328113RNAArtificial Sequencesynthetic
oligonucleotide 281uguacggaga cau 1328213RNAArtificial
Sequencesynthetic oligonucleotide 282agccuaucaa guu
1328313RNAArtificial Sequencesynthetic oligonucleotide
283caaguuugag cuu 1328413RNAArtificial Sequencesynthetic
oligonucleotide 284cuguggagua ugu 1328513RNAArtificial
Sequencesynthetic oligonucleotide 285aaauucugug gag
1328613RNAArtificial Sequencesynthetic oligonucleotide
286uuucaguagc aca 1328713RNAArtificial Sequencesynthetic
oligonucleotide 287caaugacauc uuu 1328813RNAArtificial
Sequencesynthetic oligonucleotide 288aguaccagug cac
1328913RNAArtificial Sequencesynthetic oligonucleotide
289ggaagacacg uuu 1329013RNAArtificial Sequencesynthetic
oligonucleotide 290cuaucaaguu uga 1329113RNAArtificial
Sequencesynthetic oligonucleotide 291agcuaaauuc ugu
1329213RNAArtificial Sequencesynthetic oligonucleotide
292agguagaaug uaa 1329313RNAArtificial Sequencesynthetic
oligonucleotide 293agcugaucag uuu 1329413RNAArtificial
Sequencesynthetic oligonucleotide 294uucugcucag aua
1329513RNAArtificial Sequencesynthetic oligonucleotide
295uuaucuaagu uaa 1329613RNAArtificial Sequencesynthetic
oligonucleotide 296uauacgagua aua 1329713RNAArtificial
Sequencesynthetic oligonucleotide 297gacuggacag cuu
1329813RNAArtificial Sequencesynthetic oligonucleotide
298auggccuuua uua 1329913RNAArtificial Sequencesynthetic
oligonucleotide 299auaccgagcu aaa 1330013RNAArtificial
Sequencesynthetic oligonucleotide 300uuguugagag ugu
1330113RNAArtificial Sequencesynthetic oligonucleotide
301acauaccgag cua 1330213RNAArtificial Sequencesynthetic
oligonucleotide 302agcagaaagg uua 1330313RNAArtificial
Sequencesynthetic oligonucleotide 303aguuguuccu uaa
1330413RNAArtificial Sequencesynthetic oligonucleotide
304auuugaagug uaa 1330513RNAArtificial Sequencesynthetic
oligonucleotide 305aagcugaccu gga 1330613RNAArtificial
Sequencesynthetic oligonucleotide 306ggucaugaag aag
1330713RNAArtificial Sequencesynthetic oligonucleotide
307auggucaggc cuu 1330813RNAArtificial Sequencesynthetic
oligonucleotide 308gaagacacgu uug 1330913RNAArtificial
Sequencesynthetic oligonucleotide 309aggccuugcg aag
1331013RNAArtificial Sequencesynthetic oligonucleotide
310uaccgacugg aag 1331113RNAArtificial Sequencesynthetic
oligonucleotide 311accgcaagau cgg 1331213RNAArtificial
Sequencesynthetic oligonucleotide 312caggccuugc gaa
1331313RNAArtificial Sequencesynthetic oligonucleotide
313cgagcuaaau ucu 1331413RNAArtificial Sequencesynthetic
oligonucleotide 314ucuguggagu aug 1331513RNAArtificial
Sequencesynthetic oligonucleotide 315cggagacaug gca
1331613RNAArtificial Sequencesynthetic oligonucleotide
316augacaacgc cuc 1331713RNAArtificial Sequencesynthetic
oligonucleotide 317gaggucauga aga 1331813RNAArtificial
Sequencesynthetic oligonucleotide 318uaaauucugu gga
1331913RNAArtificial Sequencesynthetic oligonucleotide
319uggaagacac guu 1332013RNAArtificial Sequencesynthetic
oligonucleotide 320aagauguacg gag 1332113RNAArtificial
Sequencesynthetic oligonucleotide 321aaugacaacg ccu
1332213RNAArtificial Sequencesynthetic oligonucleotide
322ggcgagguca uga 1332313RNAArtificial Sequencesynthetic
oligonucleotide 323gacacguuug gcc 1332413RNAArtificial
Sequencesynthetic oligonucleotide 324acggagacau ggc
1332513RNAArtificial Sequencesynthetic oligonucleotide
325ucaggccuug cga 1332613RNAArtificial Sequencesynthetic
oligonucleotide 326gcgaagcuga ccu 1332713RNAArtificial
Sequencesynthetic oligonucleotide 327ggaagaugua cgg
1332813RNAArtificial Sequencesynthetic oligonucleotide
328gugacuucgg cuc 1332913RNAArtificial Sequencesynthetic
oligonucleotide 329ugacuucggc ucc 1333013RNAArtificial
Sequencesynthetic oligonucleotide 330uggucaggcc uug
1333113RNAArtificial Sequencesynthetic oligonucleotide
331ucaaguuuga gcu 1333213RNAArtificial Sequencesynthetic
oligonucleotide 332gccagaacug cag 1333313RNAArtificial
Sequencesynthetic oligonucleotide 333uggaguaugu acc
1333413RNAArtificial Sequencesynthetic oligonucleotide
334gcuagagaag cag 1333513RNAArtificial Sequencesynthetic
oligonucleotide 335ggucaggccu ugc 1333613RNAArtificial
Sequencesynthetic oligonucleotide 336gagcuaaauu cug
1333713RNAArtificial Sequencesynthetic oligonucleotide
337aagacacguu ugg 1333813RNAArtificial Sequencesynthetic
oligonucleotide 338cgaggucaug aag 1333913RNAArtificial
Sequencesynthetic oligonucleotide 339ggccuugcga agc
1334013RNAArtificial Sequencesynthetic oligonucleotide
340cuugcgaagc uga 1334113RNAArtificial Sequencesynthetic
oligonucleotide 341ccgacuggaa gac 1334213RNAArtificial
Sequencesynthetic oligonucleotide 342ccuaucaagu uug
1334313RNAArtificial Sequencesynthetic oligonucleotide
343uguuccaaga ccu 1334413RNAArtificial Sequencesynthetic
oligonucleotide 344cgaagcugac cug 1334513RNAArtificial
Sequencesynthetic oligonucleotide 345uugcgaagcu gac
1334613RNAArtificial Sequencesynthetic oligonucleotide
346caaugacaac gcc 1334713RNAArtificial Sequencesynthetic
oligonucleotide 347guaccagugc acg 1334813RNAArtificial
Sequencesynthetic oligonucleotide 348ccuguuccaa gac
1334913RNAArtificial Sequencesynthetic oligonucleotide
349uacggagaca ugg 1335013RNAArtificial Sequencesynthetic
oligonucleotide 350ugcgaagcug acc 1335113RNAArtificial
Sequencesynthetic oligonucleotide 351ccuugcgaag cug
1335213RNAArtificial Sequencesynthetic oligonucleotide
352cugugacuuc ggc 1335313RNAArtificial Sequencesynthetic
oligonucleotide 353gcuaaauucu gug 1335413RNAArtificial
Sequencesynthetic oligonucleotide 354cuaaauucug ugg
1335513RNAArtificial Sequencesynthetic oligonucleotide
355agacacguuu ggc 1335613RNAArtificial Sequencesynthetic
oligonucleotide 356ccgcaagauc ggc 1335713RNAArtificial
Sequencesynthetic oligonucleotide 357uaucaaguuu gag
1335813RNAArtificial Sequencesynthetic oligonucleotide
358gaagcugacc ugg 1335913RNAArtificial Sequencesynthetic
oligonucleotide 359cucaugaauu aga 1336013RNAArtificial
Sequencesynthetic oligonucleotide 360cugaggucaa uua
1336113RNAArtificial Sequencesynthetic oligonucleotide
361gaggucaauu aaa 1336213RNAArtificial Sequencesynthetic
oligonucleotide 362ucugagguca auu 1336313RNAArtificial
Sequencesynthetic oligonucleotide 363ugaggucaau uaa
1336413RNAArtificial Sequencesynthetic oligonucleotide
364uucugagguc aau 1336513RNAArtificial Sequencesynthetic
oligonucleotide 365gucagcugga uga 1336613RNAArtificial
Sequencesynthetic oligonucleotide 366uucugaugaa ucu
1336713RNAArtificial Sequencesynthetic oligonucleotide
367uggacugagg uca 1336813RNAArtificial Sequencesynthetic
oligonucleotide 368gagucucacc auu 1336913RNAArtificial
Sequencesynthetic oligonucleotide 369gacugagguc aaa
1337013RNAArtificial Sequencesynthetic oligonucleotide
370ucacagccau gaa 1337113RNAArtificial Sequencesynthetic
oligonucleotide 371agucucacca uuc 1337213RNAArtificial
Sequencesynthetic oligonucleotide 372aagcggaaag cca
1337313RNAArtificial Sequencesynthetic oligonucleotide
373agcggaaagc caa 1337413RNAArtificial Sequencesynthetic
oligonucleotide 374accacaugga uga 1337513RNAArtificial
Sequencesynthetic oligonucleotide 375gccaugacca cau
1337613RNAArtificial Sequencesynthetic oligonucleotide
376aagccaugac cac 1337713RNAArtificial Sequencesynthetic
oligonucleotide 377gcggaaagcc aau
1337813RNAArtificial Sequencesynthetic oligonucleotide
378aaauuucgua uuu 1337913RNAArtificial Sequencesynthetic
oligonucleotide 379auuucguauu ucu 1338013RNAArtificial
Sequencesynthetic oligonucleotide 380aaagccauga cca
1338113RNAArtificial Sequencesynthetic oligonucleotide
381acauggauga uau 1338213RNAArtificial Sequencesynthetic
oligonucleotide 382gaaauuucgu auu 1338313RNAArtificial
Sequencesynthetic oligonucleotide 383gcgccuucug auu
1338413RNAArtificial Sequencesynthetic oligonucleotide
384auuucucaug aau 1338513RNAArtificial Sequencesynthetic
oligonucleotide 385cucucaugaa uag 1338613RNAArtificial
Sequencesynthetic oligonucleotide 386aaguccaacg aaa
1338713RNAArtificial Sequencesynthetic oligonucleotide
387augaugagag caa 1338813RNAArtificial Sequencesynthetic
oligonucleotide 388gcgaggaguu gaa 1338913RNAArtificial
Sequencesynthetic oligonucleotide 389ugauugauag uca
1339013RNAArtificial Sequencesynthetic oligonucleotide
390agauagugca ucu 1339113RNAArtificial Sequencesynthetic
oligonucleotide 391auguguaucu auu 1339213RNAArtificial
Sequencesynthetic oligonucleotide 392uucuauagaa gaa
1339313RNAArtificial Sequencesynthetic oligonucleotide
393uuguccagca auu 1339413RNAArtificial Sequencesynthetic
oligonucleotide 394acauggaaag cga 1339513RNAArtificial
Sequencesynthetic oligonucleotide 395gcaguccaga uua
1339613RNAArtificial Sequencesynthetic oligonucleotide
396ugguugaaug ugu 1339713RNAArtificial Sequencesynthetic
oligonucleotide 397uuaugaaacg agu 1339813RNAArtificial
Sequencesynthetic oligonucleotide 398caguccagau uau
1339913RNAArtificial Sequencesynthetic oligonucleotide
399auauaagcgg aaa 1340013RNAArtificial Sequencesynthetic
oligonucleotide 400uaccaguuaa aca 1340113RNAArtificial
Sequencesynthetic oligonucleotide 401uguucauucu aua
1340213RNAArtificial Sequencesynthetic oligonucleotide
402ccgaccaagg aaa 1340313RNAArtificial Sequencesynthetic
oligonucleotide 403gaauggugca uac 1340413RNAArtificial
Sequencesynthetic oligonucleotide 404auaugauggc cga
1340513RNAArtificial Sequencesynthetic oligonucleotide
405agcaguccag auu 1340613RNAArtificial Sequencesynthetic
oligonucleotide 406gcauuuaguc aaa 1340713RNAArtificial
Sequencesynthetic oligonucleotide 407agcauuccga ugu
1340813RNAArtificial Sequencesynthetic oligonucleotide
408uagucaggaa cuu 1340913RNAArtificial Sequencesynthetic
oligonucleotide 409ugcauuuagu caa 1341013RNAArtificial
Sequencesynthetic oligonucleotide 410gucugaugag ucu
1341113RNAArtificial Sequencesynthetic oligonucleotide
411uagacacaua uga 1341213RNAArtificial Sequencesynthetic
oligonucleotide 412cagacgagga cau 1341313RNAArtificial
Sequencesynthetic oligonucleotide 413cagccgugaa uuc
1341413RNAArtificial Sequencesynthetic oligonucleotide
414agucuggaaa uaa 1341513RNAArtificial Sequencesynthetic
oligonucleotide 415aguuuguggc uuc 1341613RNAArtificial
Sequencesynthetic oligonucleotide 416aguccaacga aag
1341713RNAArtificial Sequencesynthetic oligonucleotide
417aaguuucgca gac 1341813RNAArtificial Sequencesynthetic
oligonucleotide 418agcaaugagc auu 1341913RNAArtificial
Sequencesynthetic oligonucleotide 419uuagauagug cau
1342013RNAArtificial Sequencesynthetic oligonucleotide
420uggugcauac aag 1342113RNAArtificial Sequencesynthetic
oligonucleotide 421augaaacgag uca 1342213RNAArtificial
Sequencesynthetic oligonucleotide 422ccagagugcu gaa
1342313RNAArtificial Sequencesynthetic oligonucleotide
423cagccaugaa uuu 1342413RNAArtificial Sequencesynthetic
oligonucleotide 424auugguugaa ugu 1342513RNAArtificial
Sequencesynthetic oligonucleotide 425gguugaaugu gua
1342613RNAArtificial Sequencesynthetic oligonucleotide
426ggaaauaacu aau 1342713RNAArtificial Sequencesynthetic
oligonucleotide 427ucaugaauag aaa 1342813RNAArtificial
Sequencesynthetic oligonucleotide 428gccagcaacc gaa
1342913RNAArtificial Sequencesynthetic oligonucleotide
429caccucacac aug 1343013RNAArtificial Sequencesynthetic
oligonucleotide 430aguugaaugg ugc 1343113RNAArtificial
Sequencesynthetic oligonucleotide 431agucagcugg aug
1343213RNAArtificial Sequencesynthetic oligonucleotide
432uauaagcgga aag 1343313RNAArtificial Sequencesynthetic
oligonucleotide 433uuccgaugug auu 1343413RNAArtificial
Sequencesynthetic oligonucleotide 434auaacuaaug ugu
1343513RNAArtificial Sequencesynthetic oligonucleotide
435ucauucuaua gaa 1343613RNAArtificial Sequencesynthetic
oligonucleotide 436aacuaucacu gua 1343713RNAArtificial
Sequencesynthetic oligonucleotide 437gucaauugcu uau
1343813RNAArtificial Sequencesynthetic oligonucleotide
438agcaauuaau aaa 1343913RNAArtificial Sequencesynthetic
oligonucleotide 439acgacucuga uga 1344013RNAArtificial
Sequencesynthetic oligonucleotide 440uagugugguu uau
1344113RNAArtificial Sequencesynthetic oligonucleotide
441aagccaauga uga 1344213RNAArtificial Sequencesynthetic
oligonucleotide 442auagucagga acu 1344313RNAArtificial
Sequencesynthetic oligonucleotide 443agucagccgu gaa
1344413RNAArtificial Sequencesynthetic oligonucleotide
444acuaccauga gaa 1344513RNAArtificial Sequencesynthetic
oligonucleotide 445aaacaggcug auu 1344613RNAArtificial
Sequencesynthetic oligonucleotide 446gagugcugaa acc
1344713RNAArtificial Sequencesynthetic oligonucleotide
447ugagcauucc gau 1344813RNAArtificial Sequencesynthetic
oligonucleotide 448aauuccacag cca 1344913RNAArtificial
Sequencesynthetic oligonucleotide 449ugucaauugc uua
1345013RNAArtificial Sequencesynthetic oligonucleotide
450accaugagaa uug 1345113RNAArtificial Sequencesynthetic
oligonucleotide 451ccaacgaaag cca 1345213RNAArtificial
Sequencesynthetic oligonucleotide 452cuggucacug auu
1345313RNAArtificial Sequencesynthetic oligonucleotide
453ugguuuaugg acu 1345413RNAArtificial Sequencesynthetic
oligonucleotide 454gaccagagug cug 1345513RNAArtificial
Sequencesynthetic oligonucleotide 455gaugugauug aua
1345613RNAArtificial Sequencesynthetic oligonucleotide
456gucagccgug aau 1345713RNAArtificial Sequencesynthetic
oligonucleotide 457aauguguauc uau 1345813RNAArtificial
Sequencesynthetic oligonucleotide 458uugagucugg aaa
1345913RNAArtificial Sequencesynthetic oligonucleotide
459guccagcaau uaa 1346013RNAArtificial Sequencesynthetic
oligonucleotide 460ccagcaauua aua 1346113RNAArtificial
Sequencesynthetic oligonucleotide 461gacucgaacg acu
1346213RNAArtificial Sequencesynthetic oligonucleotide
462accugccagc aac 1346313DNAArtificial Sequencesynthetic
oligonucleotide 463actgaauacc aau 1346413RNAArtificial
Sequencesynthetic oligonucleotide 464acugaauacc aau
1346513RNAArtificial Sequencesynthetic oligonucleotide
465cuguggaagu cua 1346613RNAArtificial Sequencesynthetic
oligonucleotide 466ggcuacaaaa aca 1346715RNAArtificial
Sequencesynthetic oligonucleotide 467uuggcuacaa aaaca
1546817RNAArtificial Sequencesynthetic oligonucleotide
468auuuggcuac aaaaaca 1746913RNAArtificial Sequencesynthetic
oligonucleotide 469uguaggaugu cua 1347013RNAArtificial
Sequencesynthetic oligonucleotide 470aucuggagaa aca
1347115RNAArtificial Sequencesynthetic oligonucleotide
471agaucuggag aaaca 1547213RNAArtificial Sequencesynthetic
oligonucleotide 472gacucaucug cua 1347313RNAArtificial
Sequencesynthetic oligonucleotide 473gacucaucug cua
1347415RNAArtificial Sequencesynthetic oligonucleotide
474uggacucauc ugcua 1547515RNAArtificial Sequencesynthetic
oligonucleotide 475uggacucauc ugcua 1547613RNAArtificial
Sequencesynthetic oligonucleotide 476ggagaaacaa cau
1347715RNAArtificial Sequencesynthetic oligonucleotide
477cuggagaaac aacau 1547813RNAArtificial Sequencesynthetic
oligonucleotide 478agucccucaa aca 1347915RNAArtificial
Sequencesynthetic oligonucleotide 479agagucccuc aaaca
1548013RNAArtificial Sequencesynthetic oligonucleotide
480acugaauacc aau 1348113RNAArtificial Sequencesynthetic
oligonucleotide 481acugaauacc aau 1348215RNAArtificial
Sequencesynthetic oligonucleotide 482acacugaaua ccaau
1548315RNAArtificial Sequencesynthetic oligonucleotide
483acacugaaua ccaau 1548413RNAArtificial Sequencesynthetic
oligonucleotide 484cuguggaagu cua 1348513RNAArtificial
Sequencesynthetic oligonucleotide 485acaggaagau gua
1348613RNAArtificial Sequencesynthetic oligonucleotide
486gaguggagcg ccu 1348713RNAArtificial Sequencesynthetic
oligonucleotide 487cgacuggaag aca 1348813RNAArtificial
Sequencesynthetic oligonucleotide 488ggagcgccug uuc
1348913RNAArtificial Sequencesynthetic oligonucleotide
489gccauuacaa cug 1349013RNAArtificial Sequencesynthetic
oligonucleotide 490gagcuuucug gcu 1349113RNAArtificial
Sequencesynthetic oligonucleotide 491aguggagcgc cug
1349213RNAArtificial Sequencesynthetic oligonucleotide
492uggagcgccu guu 1349313RNAArtificial Sequencesynthetic
oligonucleotide 493guuugagcuu ucu 1349413RNAArtificial
Sequencesynthetic oligonucleotide 494ugccauuaca acu
1349513RNAArtificial Sequencesynthetic oligonucleotide
495acuggaagac acg 1349613RNAArtificial Sequencesynthetic
oligonucleotide 496aacugccugg ucc 1349713RNAArtificial
Sequencesynthetic oligonucleotide 497agaccugugc cug
1349813RNAArtificial Sequencesynthetic oligonucleotide
498cagaguggag cgc 1349913RNAArtificial Sequencesynthetic
oligonucleotide 499ccugguccag acc 1350013RNAArtificial
Sequencesynthetic oligonucleotide 500ccauuacaac ugu
1350113RNAArtificial Sequencesynthetic oligonucleotide
501cugccauuac aac 1350213RNAArtificial Sequencesynthetic
oligonucleotide 502auuacaacug ucc 1350313RNAArtificial
Sequencesynthetic oligonucleotide 503cauuacaacu guc
1350413RNAArtificial Sequencesynthetic oligonucleotide
504agaguggagc gcc 1350513RNAArtificial Sequencesynthetic
oligonucleotide 505accgacugga aga 1350613RNAArtificial
Sequencesynthetic oligonucleotide 506auguacggag aca
1350713RNAArtificial Sequencesynthetic oligonucleotide
507gccuugcgaa gcu 1350813RNAArtificial Sequencesynthetic
oligonucleotide 508gcugcgagga gug 1350913RNAArtificial
Sequencesynthetic oligonucleotide 509gccuaucaag uuu
1351013RNAArtificial Sequencesynthetic oligonucleotide
510aauucugugg agu 1351113RNAArtificial Sequencesynthetic
oligonucleotide 511uguacggaga cau 1351213RNAArtificial
Sequencesynthetic oligonucleotide 512agccuaucaa guu
1351313RNAArtificial Sequencesynthetic oligonucleotide
513caaguuugag cuu 1351413RNAArtificial Sequencesynthetic
oligonucleotide 514cuguggagua ugu 1351513RNAArtificial
Sequencesynthetic oligonucleotide 515aaauucugug gag
1351613RNAArtificial Sequencesynthetic oligonucleotide
516uuucaguagc aca 1351713RNAArtificial Sequencesynthetic
oligonucleotide 517caaugacauc uuu 1351813RNAArtificial
Sequencesynthetic oligonucleotide 518aguaccagug cac
1351913RNAArtificial Sequencesynthetic oligonucleotide
519ggaagacacg uuu 1352013RNAArtificial Sequencesynthetic
oligonucleotide 520cuaucaaguu uga 1352113RNAArtificial
Sequencesynthetic oligonucleotide 521agcuaaauuc ugu
1352213RNAArtificial Sequencesynthetic oligonucleotide
522agguagaaug uaa 1352313RNAArtificial Sequencesynthetic
oligonucleotide 523agcugaucag uuu 1352413RNAArtificial
Sequencesynthetic oligonucleotide 524uucugcucag aua
1352513RNAArtificial Sequencesynthetic oligonucleotide
525uuaucuaagu uaa 1352613RNAArtificial Sequencesynthetic
oligonucleotide 526uauacgagua aua 1352713RNAArtificial
Sequencesynthetic oligonucleotide 527gacuggacag cuu
1352813RNAArtificial Sequencesynthetic oligonucleotide
528auggccuuua uua 1352913RNAArtificial Sequencesynthetic
oligonucleotide 529auaccgagcu aaa 1353013RNAArtificial
Sequencesynthetic oligonucleotide 530uuguugagag ugu
1353113RNAArtificial Sequencesynthetic oligonucleotide
531acauaccgag cua 1353213RNAArtificial Sequencesynthetic
oligonucleotide 532agcagaaagg uua 1353313RNAArtificial
Sequencesynthetic oligonucleotide 533aguuguuccu uaa
1353413RNAArtificial Sequencesynthetic oligonucleotide
534auuugaagug uaa 1353513RNAArtificial Sequencesynthetic
oligonucleotide 535aagcugaccu gga 1353613RNAArtificial
Sequencesynthetic oligonucleotide 536ggucaugaag aag
1353713RNAArtificial Sequencesynthetic oligonucleotide
537auggucaggc cuu 1353813RNAArtificial Sequencesynthetic
oligonucleotide 538gaagacacgu uug 1353913RNAArtificial
Sequencesynthetic oligonucleotide 539aggccuugcg aag
1354013RNAArtificial Sequencesynthetic oligonucleotide
540uaccgacugg aag 1354113RNAArtificial Sequencesynthetic
oligonucleotide 541accgcaagau cgg 1354213RNAArtificial
Sequencesynthetic oligonucleotide 542caggccuugc gaa
1354313RNAArtificial Sequencesynthetic oligonucleotide
543cgagcuaaau ucu 1354413RNAArtificial Sequencesynthetic
oligonucleotide 544ucuguggagu aug 1354513RNAArtificial
Sequencesynthetic oligonucleotide 545cggagacaug gca
1354613RNAArtificial Sequencesynthetic oligonucleotide
546augacaacgc cuc 1354713RNAArtificial Sequencesynthetic
oligonucleotide 547gaggucauga aga 1354813RNAArtificial
Sequencesynthetic oligonucleotide 548uaaauucugu gga
1354913RNAArtificial Sequencesynthetic oligonucleotide
549uggaagacac guu 1355013RNAArtificial Sequencesynthetic
oligonucleotide 550aagauguacg gag 1355113RNAArtificial
Sequencesynthetic oligonucleotide 551aaugacaacg ccu
1355213RNAArtificial Sequencesynthetic oligonucleotide
552ggcgagguca uga 1355313RNAArtificial Sequencesynthetic
oligonucleotide 553gacacguuug gcc 1355413RNAArtificial
Sequencesynthetic oligonucleotide 554acggagacau ggc
1355513RNAArtificial Sequencesynthetic oligonucleotide
555ucaggccuug cga 1355613RNAArtificial Sequencesynthetic
oligonucleotide 556gcgaagcuga ccu 1355713RNAArtificial
Sequencesynthetic oligonucleotide 557ggaagaugua cgg
1355813RNAArtificial Sequencesynthetic oligonucleotide
558gugacuucgg cuc 1355913RNAArtificial Sequencesynthetic
oligonucleotide 559ugacuucggc ucc 1356013RNAArtificial
Sequencesynthetic oligonucleotide 560uggucaggcc uug
1356113RNAArtificial Sequencesynthetic oligonucleotide
561ucaaguuuga gcu 1356213RNAArtificial Sequencesynthetic
oligonucleotide 562gccagaacug cag 1356313RNAArtificial
Sequencesynthetic oligonucleotide 563uggaguaugu acc
1356413RNAArtificial Sequencesynthetic oligonucleotide
564gcuagagaag cag 1356513RNAArtificial Sequencesynthetic
oligonucleotide 565ggucaggccu ugc 1356613RNAArtificial
Sequencesynthetic oligonucleotide 566gagcuaaauu cug
1356713RNAArtificial Sequencesynthetic oligonucleotide
567aagacacguu ugg 1356813RNAArtificial Sequencesynthetic
oligonucleotide 568cgaggucaug aag 1356913RNAArtificial
Sequencesynthetic oligonucleotide 569ggccuugcga agc
1357013RNAArtificial Sequencesynthetic oligonucleotide
570cuugcgaagc uga 1357113RNAArtificial Sequencesynthetic
oligonucleotide 571ccgacuggaa gac 1357213RNAArtificial
Sequencesynthetic oligonucleotide 572ccuaucaagu uug
1357313RNAArtificial Sequencesynthetic oligonucleotide
573uguuccaaga ccu 1357413RNAArtificial Sequencesynthetic
oligonucleotide 574cgaagcugac cug 1357513RNAArtificial
Sequencesynthetic oligonucleotide 575uugcgaagcu gac
1357613RNAArtificial Sequencesynthetic oligonucleotide
576caaugacaac gcc 1357713RNAArtificial Sequencesynthetic
oligonucleotide 577guaccagugc acg 1357813RNAArtificial
Sequencesynthetic oligonucleotide 578ccuguuccaa gac
1357913RNAArtificial Sequencesynthetic oligonucleotide
579uacggagaca ugg 1358013RNAArtificial Sequencesynthetic
oligonucleotide 580ugcgaagcug acc 1358113RNAArtificial
Sequencesynthetic oligonucleotide 581ccuugcgaag cug
1358213RNAArtificial Sequencesynthetic oligonucleotide
582cugugacuuc ggc 1358313RNAArtificial Sequencesynthetic
oligonucleotide 583gcuaaauucu gug 1358413RNAArtificial
Sequencesynthetic oligonucleotide 584cuaaauucug ugg
1358513RNAArtificial Sequencesynthetic oligonucleotide
585agacacguuu ggc 1358613RNAArtificial Sequencesynthetic
oligonucleotide 586ccgcaagauc ggc 1358713RNAArtificial
Sequencesynthetic oligonucleotide 587uaucaaguuu gag
1358813RNAArtificial Sequencesynthetic oligonucleotide
588gaagcugacc ugg 1358913RNAArtificial Sequencesynthetic
oligonucleotide 589gcauuuaguc aaa 1359013RNAArtificial
Sequencesynthetic oligonucleotide 590ggcuacaaaa aca
1359115RNAArtificial Sequencesynthetic oligonucleotide
591uuggcuacaa aaaca 1559217RNAArtificial Sequencesynthetic
oligonucleotide 592auuuggcuac aaaaaca 1759313RNAArtificial
Sequencesynthetic oligonucleotide 593acaaauacga uuu
1359413RNAArtificial Sequencesynthetic oligonucleotide
594acaaauacga uuu 1359525RNAArtificial Sequencesynthetic
oligonucleotide 595cuuugaagag uucuguggaa gucua 2559623RNAArtificial
Sequencesynthetic oligonucleotide 596acaaacacca uugucacacu cca
2359719RNAArtificial sequencesynthetic oligonucleotide
597uagacuucca cagaacucu 1959813RNAArtificial sequencesynthetic
oligonucleotide 598cuguggaagu cua 1359932RNAArtificial
sequencesynthetic oligonuclotide 599uagacuucca cagaacucug
acaccuucag au 3260013RNAArtificial sequencesynthetic
oligonucleotide 600uagacuucca cag 1360113RNAArtificial
sequencesynthetic oligonucleotide 601cuguggaagu cua
1360231RNAArtificial sequencesynthetic oligonucleotide
602uagacuucca cagaacucuu guggaagucu a 3160331RNAArtificial
sequencesynthetic oligonucleotide 603uagacuucca cagaacucuu
guggaagucu a 3160425RNAArtificial sequencesynthetic oligonucleotide
604cuuugaagag uucuguggaa gucua 2560525RNAArtificial
sequencesynthetic oligonucleotide 605uagacuucca cagaacucuu caaag
2560631RNAArtificial sequencesynthetic oligonucleotide
606uagacuucca cagaacuucu guggaagucu a 3160731RNAArtificial
sequencesynthetic oligonucleotide 607uagacuucca cagaacuucu
guggaagucu a 3160813RNAArtificial sequencesynthetoc oligonucleotide
608uacuuucuuc auu 1360913RNAArtificial sequencesynthetic
oligonucleotide 609aaugaagaaa gua 1361032RNAArtificial
sequencesynthetic oligonucleotide 610uacuuucuuc auuuccacca
augaagaaag ua 3261132RNAArtificial sequencesynthetic
oligonucleotide 611uacuuucuuc auuuccacca augaagaaag ua
3261219RNAArtificial sequencesynthetic oligonucleotide
612uacuuucuuc auuuccacc 1961313RNAArtificial sequencesynthetic
oligonucleotide 613aaugaagaaa gua 1361425RNAArtificial
sequencesynthetic oligonucleotide 614uacuuucuuc auuuccaccu uugcc
2561525RNAArtificial sequencesynthetic oligonucleotide
615ggcaaaggug gaaaugaaga aagua 2561619RNAArtificial
Sequencesynthetic oligonucleotide 616ucuaauucau gagaaauac
1961719RNAArtificial Sequencesynthetic oligonucleotide
617uaauugaccu cagaagaug 1961819RNAArtificial Sequencesynthetic
oligonucleotide 618uuuaauugac cucagaaga 1961919RNAArtificial
Sequencesynthetic oligonucleotide 619aauugaccuc agaagaugc
1962019RNAArtificial Sequencesynthetic oligonucleotide
620uuaauugacc ucagaagau 1962119RNAArtificial Sequencesynthetic
oligonucleotide 621auugaccuca gaagaugca 1962219RNAArtificial
Sequencesynthetic oligonucleotide 622ucauccagcu gacucguuu
1962319RNAArtificial Sequencesynthetic oligonucleotide
623agauucauca gaaugguga 1962419RNAArtificial Sequencesynthetic
oligonucleotide 624ugaccucagu ccauaaacc 1962519RNAArtificial
Sequencesynthetic oligonucleotide 625aauggugaga cucaucaga
1962619RNAArtificial Sequencesynthetic oligonucleotide
626uuugaccuca guccauaaa 1962719RNAArtificial Sequencesynthetic
oligonucleotide 627uucauggcug ugaaauuca 1962819RNAArtificial
Sequencesynthetic oligonucleotide 628gaauggugag acucaucag
1962919RNAArtificial Sequencesynthetic oligonucleotide
629uggcuuuccg cuuauauaa 1963019RNAArtificial Sequencesynthetic
oligonucleotide 630uuggcuuucc gcuuauaua 1963119RNAArtificial
Sequencesynthetic oligonucleotide 631ucauccaugu ggucauggc
1963219RNAArtificial Sequencesynthetic oligonucleotide
632auguggucau ggcuuucgu 1963319RNAArtificial Sequencesynthetic
oligonucleotide 633guggucaugg cuuucguug 1963419RNAArtificial
Sequencesynthetic oligonucleotide 634auuggcuuuc cgcuuauau
1963519RNAArtificial Sequencesynthetic oligonucleotide
635aaauacgaaa uuucaggug 1963619RNAArtificial Sequencesynthetic
oligonucleotide 636agaaauacga aauuucagg 1963719RNAArtificial
Sequencesynthetic oligonucleotide 637uggucauggc uuucguugg
1963819RNAArtificial Sequencesynthetic oligonucleotide
638auaucaucca uguggucau 1963919RNAArtificial Sequencesynthetic
oligonucleotide 639aauacgaaau uucaggugu 1964019RNAArtificial
Sequencesynthetic oligonucleotide 640aaucagaagg cgcguucag
1964119RNAArtificial Sequencesynthetic oligonucleotide
641auucaugaga aauacgaaa 1964219RNAArtificial Sequencesynthetic
oligonucleotide 642cuauucauga gagaauaac 1964319RNAArtificial
Sequencesynthetic oligonucleotide 643uuucguugga cuuacuugg
1964419RNAArtificial Sequencesynthetic oligonucleotide
644uugcucucau cauuggcuu 1964519RNAArtificial Sequencesynthetic
oligonucleotide 645uucaacuccu cgcuuucca 1964619RNAArtificial
Sequencesynthetic oligonucleotide 646ugacuaucaa ucacaucgg
1964719RNAArtificial Sequencesynthetic oligonucleotide
647agaugcacua ucuaauuca 1964819RNAArtificial Sequencesynthetic
oligonucleotide 648aauagauaca cauucaacc 1964919RNAArtificial
Sequencesynthetic oligonucleotide 649uucuucuaua gaaugaaca
1965019RNAArtificial Sequencesynthetic oligonucleotide
650aauugcugga caaccgugg 1965119RNAArtificial Sequencesynthetic
oligonucleotide 651ucgcuuucca ugugugagg 1965219RNAArtificial
Sequencesynthetic oligonucleotide 652uaaucuggac ugcuugugg
1965319RNAArtificial Sequencesynthetic oligonucleotide
653acacauucaa ccaauaaac 1965419RNAArtificial Sequencesynthetic
oligonucleotide 654acucguuuca uaacugucc 1965519RNAArtificial
Sequencesynthetic oligonucleotide 655auaaucugga cugcuugug
1965619RNAArtificial Sequencesynthetic oligonucleotide
656uuuccgcuua uauaaucug 1965719RNAArtificial Sequencesynthetic
oligonucleotide 657uguuuaacug guauggcac 1965819RNAArtificial
Sequencesynthetic oligonucleotide 658uauagaauga acauagaca
1965919RNAArtificial Sequencesynthetic oligonucleotide
659uuuccuuggu cggcguuug 1966019RNAArtificial Sequencesynthetic
oligonucleotide 660guaugcacca uucaacucc 1966119RNAArtificial
Sequencesynthetic oligonucleotide 661ucggccauca uaugugucu
1966219RNAArtificial Sequencesynthetic oligonucleotide
662aaucuggacu gcuuguggc 1966319RNAArtificial Sequencesynthetic
oligonucleotide 663acaucggaau gcucauugc 1966419RNAArtificial
Sequencesynthetic oligonucleotide 664aaguuccuga cuaucaauc
1966519RNAArtificial Sequencesynthetic oligonucleotide
665uugacuaaau gcaaaguga 1966619RNAArtificial Sequencesynthetic
oligonucleotide 666agacucauca gacugguga 1966719RNAArtificial
Sequencesynthetic oligonucleotide 667ucauaugugu cuacugugg
1966819RNAArtificial Sequencesynthetic oligonucleotide
668auguccucgu cuguagcau 1966919RNAArtificial Sequencesynthetic
oligonucleotide 669gaauucacgg cugacuuug 1967019RNAArtificial
Sequencesynthetic oligonucleotide 670uuauuuccag acucaaaua
1967119RNAArtificial Sequencesynthetic oligonucleotide
671gaagccacaa acuaaacua 1967219RNAArtificial Sequencesynthetic
oligonucleotide 672cuuucguugg acuuacuug 1967319RNAArtificial
Sequencesynthetic oligonucleotide 673gucugcgaaa cuucuuaga
1967419RNAArtificial Sequencesynthetic oligonucleotide
674aaugcucauu gcucucauc 1967519RNAArtificial Sequencesynthetic
oligonucleotide 675augcacuauc uaauucaug 1967619RNAArtificial
Sequencesynthetic oligonucleotide 676cuuguaugca ccauucaac
1967719RNAArtificial Sequencesynthetic oligonucleotide
677ugacucguuu cauaacugu 1967819RNAArtificial Sequencesynthetic
oligonucleotide 678uucagcacuc uggucaucc 1967919RNAArtificial
Sequencesynthetic oligonucleotide 679aaauucaugg cuguggaau
1968019RNAArtificial Sequencesynthetic oligonucleotide
680acauucaacc aauaaacug 1968119RNAArtificial Sequencesynthetic
oligonucleotide 681uacacauuca accaauaaa 1968219RNAArtificial
Sequencesynthetic oligonucleotide 682auuaguuauu uccagacuc
1968319RNAArtificial Sequencesynthetic oligonucleotide
683uuucuauuca ugagagaau 1968419RNAArtificial Sequencesynthetic
oligonucleotide 684uucgguugcu ggcaggucc 1968519RNAArtificial
Sequencesynthetic oligonucleotide 685caugugugag gugaugucc
1968619RNAArtificial Sequencesynthetic oligonucleotide
686gcaccauuca acuccucgc 1968719RNAArtificial Sequencesynthetic
oligonucleotide 687cauccagcug acucguuuc 1968819RNAArtificial
Sequencesynthetic oligonucleotide 688cuuuccgcuu auauaaucu
1968919RNAArtificial Sequencesynthetic oligonucleotide
689aaucacaucg gaaugcuca 1969019RNAArtificial Sequencesynthetic
oligonucleotide 690acacauuagu uauuuccag 1969119RNAArtificial
Sequencesynthetic oligonucleotide 691uucuauagaa ugaacauag
1969219RNAArtificial Sequencesynthetic oligonucleotide
692uacagugaua guuugcauu 1969319RNAArtificial Sequencesynthetic
oligonucleotide 693auaagcaauu gacaccacc 1969419RNAArtificial
Sequencesynthetic oligonucleotide 694uuuauuaauu gcuggacaa
1969519RNAArtificial Sequencesynthetic oligonucleotide
695ucaucagagu cguucgagu 1969619RNAArtificial Sequencesynthetic
oligonucleotide 696auaaaccaca cuaucaccu 1969719RNAArtificial
Sequencesynthetic oligonucleotide 697ucaucauugg cuuuccgcu
1969819RNAArtificial Sequencesynthetic oligonucleotide
698aguuccugac uaucaauca 1969919RNAArtificial Sequencesynthetic
oligonucleotide 699uucacggcug acuuuggaa 1970019RNAArtificial
Sequencesynthetic oligonucleotide 700uucucauggu agugaguuu
1970119RNAArtificial Sequencesynthetic oligonucleotide
701aaucagccug uuuaacugg 1970219RNAArtificial Sequencesynthetic
oligonucleotide 702gguuucagca cucugguca 1970319RNAArtificial
Sequencesynthetic oligonucleotide 703aucggaaugc ucauugcuc
1970419RNAArtificial Sequencesynthetic oligonucleotide
704uggcugugga auucacggc 1970519RNAArtificial Sequencesynthetic
oligonucleotide 705uaagcaauug acaccacca 1970619RNAArtificial
Sequencesynthetic oligonucleotide 706caauucucau gguagugag
1970719RNAArtificial Sequencesynthetic oligonucleotide
707uggcuuucgu uggacuuac 1970819RNAArtificial Sequencesynthetic
oligonucleotide 708aaucagugac caguucauc 1970919RNAArtificial
Sequencesynthetic oligonucleotide 709aguccauaaa ccacacuau
1971019RNAArtificial Sequencesynthetic oligonucleotide
710cagcacucug gucauccag 1971119RNAArtificial Sequencesynthetic
oligonucleotide 711uaucaaucac aucggaaug 1971219RNAArtificial
Sequencesynthetic oligonucleotide 712auucacggcu gacuuugga
1971319RNAArtificial Sequencesynthetic oligonucleotide
713auagauacac auucaacca 1971419RNAArtificial Sequencesynthetic
oligonucleotide 714uuuccagacu caaauagau 1971519RNAArtificial
Sequencesynthetic oligonucleotide 715uuaauugcug gacaaccgu
1971619RNAArtificial Sequencesynthetic oligonucleotide
716uauuaauugc uggacaacc 1971719RNAArtificial Sequencesynthetic
oligonucleotide 717agucguucga gucaaugga 1971819RNAArtificial
Sequencesynthetic oligonucleotide 718guugcuggca gguccgugg
1971913RNAArtificial Sequencesynthetic oligonucleotide
719cucaugaauu aga 1372013RNAArtificial Sequencesynthetic
oligonucleotide 720cugaggucaa uua 1372113RNAArtificial
Sequencesynthetic oligonucleotide 721gaggucaauu aaa
1372213RNAArtificial Sequencesynthetic oligonucleotide
722ucugagguca auu 1372313RNAArtificial Sequencesynthetic
oligonucleotide 723ugaggucaau uaa 1372413RNAArtificial
Sequencesynthetic oligonucleotide 724uucugagguc aau
1372513RNAArtificial Sequencesynthetic oligonucleotide
725gucagcugga uga 1372613RNAArtificial Sequencesynthetic
oligonucleotide 726uucugaugaa ucu 1372713RNAArtificial
Sequencesynthetic oligonucleotide 727uggacugagg uca
1372813RNAArtificial Sequencesynthetic oligonucleotide
728gagucucacc auu 1372913RNAArtificial Sequencesynthetic
oligonucleotide 729gacugagguc aaa 1373013RNAArtificial
Sequencesynthetic oligonucleotide 730ucacagccau gaa
1373113RNAArtificial Sequencesynthetic oligonucleotide
731agucucacca uuc 1373213RNAArtificial Sequencesynthetic
oligonucleotide 732aagcggaaag cca 1373313RNAArtificial
Sequencesynthetic oligonucleotide 733agcggaaagc caa
1373413RNAArtificial Sequencesynthetic oligonucleotide
734accacaugga uga 1373513RNAArtificial Sequencesynthetic
oligonucleotide 735gccaugacca cau 1373613RNAArtificial
Sequencesynthetic oligonucleotide 736aagccaugac cac
1373713RNAArtificial Sequencesynthetic oligonucleotide
737gcggaaagcc aau 1373813RNAArtificial Sequencesynthetic
oligonucleotide 738aaauuucgua uuu 1373913RNAArtificial
Sequencesynthetic oligonucleotide 739auuucguauu ucu
1374013RNAArtificial Sequencesynthetic oligonucleotide
740aaagccauga cca 1374113RNAArtificial Sequencesynthetic
oligonucleotide 741acauggauga uau 1374213RNAArtificial
Sequencesynthetic oligonucleotide 742gaaauuucgu auu
1374313RNAArtificial Sequencesynthetic oligonucleotide
743gcgccuucug auu 1374413RNAArtificial Sequencesynthetic
oligonucleotide 744auuucucaug aau 1374513RNAArtificial
Sequencesynthetic oligonucleotide 745cucucaugaa uag
1374613RNAArtificial Sequencesynthetic oligonucleotide
746aaguccaacg aaa 1374713RNAArtificial Sequencesynthetic
oligonucleotide 747augaugagag caa 1374813RNAArtificial
Sequencesynthetic oligonucleotide 748gcgaggaguu gaa
1374913RNAArtificial Sequencesynthetic oligonucleotide
749ugauugauag uca 1375013RNAArtificial Sequencesynthetic
oligonucleotide 750agauagugca ucu 1375113RNAArtificial
Sequencesynthetic oligonucleotide 751auguguaucu auu
1375213RNAArtificial Sequencesynthetic oligonucleotide
752uucuauagaa gaa 1375313RNAArtificial Sequencesynthetic
oligonucleotide 753uuguccagca auu 1375413RNAArtificial
Sequencesynthetic oligonucleotide 754acauggaaag cga
1375513RNAArtificial Sequencesynthetic oligonucleotide
755gcaguccaga uua 1375613RNAArtificial Sequencesynthetic
oligonucleotide 756ugguugaaug ugu 1375713RNAArtificial
Sequencesynthetic oligonucleotide 757uuaugaaacg agu
1375813RNAArtificial Sequencesynthetic oligonucleotide
758caguccagau uau 1375913RNAArtificial Sequencesynthetic
oligonucleotide 759auauaagcgg aaa 1376013RNAArtificial
Sequencesynthetic oligonucleotide 760uaccaguuaa aca
1376113RNAArtificial Sequencesynthetic oligonucleotide
761uguucauucu aua 1376213RNAArtificial Sequencesynthetic
oligonucleotide 762ccgaccaagg aaa 1376313RNAArtificial
Sequencesynthetic oligonucleotide 763gaauggugca uac
1376413RNAArtificial Sequencesynthetic oligonucleotide
764auaugauggc cga 1376513RNAArtificial Sequencesynthetic
oligonucleotide 765agcaguccag auu 1376613RNAArtificial
Sequencesynthetic oligonucleotide 766agcauuccga ugu
1376713RNAArtificial Sequencesynthetic oligonucleotide
767uagucaggaa cuu 1376813RNAArtificial Sequencesynthetic
oligonucleotide 768ugcauuuagu caa 1376913RNAArtificial
Sequencesynthetic oligonucleotide 769gucugaugag ucu
1377013RNAArtificial Sequencesynthetic oligonucleotide
770uagacacaua uga 1377113RNAArtificial Sequencesynthetic
oligonucleotide 771cagacgagga cau 1377213RNAArtificial
Sequencesynthetic oligonucleotide 772cagccgugaa uuc
1377313RNAArtificial Sequencesynthetic oligonucleotide
773agucuggaaa uaa 1377413RNAArtificial Sequencesynthetic
oligonucleotide 774aguuuguggc uuc 1377513RNAArtificial
Sequencesynthetic oligonucleotide 775aguccaacga aag
1377613RNAArtificial Sequencesynthetic oligonucleotide
776aaguuucgca gac 1377713RNAArtificial Sequencesynthetic
oligonucleotide 777agcaaugagc auu 1377813RNAArtificial
Sequencesynthetic oligonucleotide 778uuagauagug cau
1377913RNAArtificial Sequencesynthetic oligonucleotide
779uggugcauac aag 1378013RNAArtificial Sequencesynthetic
oligonucleotide 780augaaacgag uca 1378113RNAArtificial
Sequencesynthetic oligonucleotide 781ccagagugcu gaa
1378213RNAArtificial Sequencesynthetic oligonucleotide
782cagccaugaa uuu 1378313RNAArtificial Sequencesynthetic
oligonucleotide 783auugguugaa ugu 1378413RNAArtificial
Sequencesynthetic oligonucleotide 784gguugaaugu gua
1378513RNAArtificial Sequencesynthetic oligonucleotide
785ggaaauaacu aau 1378613RNAArtificial Sequencesynthetic
oligonucleotide 786ucaugaauag aaa 1378713RNAArtificial
Sequencesynthetic oligonucleotide 787gccagcaacc gaa
1378813RNAArtificial Sequencesynthetic oligonucleotide
788caccucacac aug 1378913RNAArtificial Sequencesynthetic
oligonucleotide 789aguugaaugg ugc 1379013RNAArtificial
Sequencesynthetic oligonucleotide 790agucagcugg aug
1379113RNAArtificial Sequencesynthetic oligonucleotide
791uauaagcgga aag 1379213RNAArtificial Sequencesynthetic
oligonucleotide 792uuccgaugug auu 1379313RNAArtificial
Sequencesynthetic oligonucleotide 793auaacuaaug ugu
1379413RNAArtificial Sequencesynthetic oligonucleotide
794ucauucuaua gaa 1379513RNAArtificial Sequencesynthetic
oligonucleotide 795aacuaucacu gua 1379613RNAArtificial
Sequencesynthetic oligonucleotide 796gucaauugcu uau
1379713RNAArtificial Sequencesynthetic oligonucleotide
797agcaauuaau aaa 1379813RNAArtificial Sequencesynthetic
oligonucleotide 798acgacucuga uga 1379913RNAArtificial
Sequencesynthetic oligonucleotide 799uagugugguu uau
1380013RNAArtificial Sequencesynthetic oligonucleotide
800aagccaauga uga 1380113RNAArtificial Sequencesynthetic
oligonucleotide 801auagucagga acu 1380213RNAArtificial
Sequencesynthetic oligonucleotide 802agucagccgu gaa
1380313RNAArtificial Sequencesynthetic oligonucleotide
803acuaccauga gaa 1380413RNAArtificial Sequencesynthetic
oligonucleotide 804aaacaggcug auu 1380513RNAArtificial
Sequencesynthetic oligonucleotide 805gagugcugaa acc
1380613RNAArtificial Sequencesynthetic oligonucleotide
806ugagcauucc gau 1380713RNAArtificial Sequencesynthetic
oligonucleotide 807aauuccacag cca 1380813RNAArtificial
Sequencesynthetic oligonucleotide 808ugucaauugc uua
1380913RNAArtificial Sequencesynthetic oligonucleotide
809accaugagaa uug 1381013RNAArtificial Sequencesynthetic
oligonucleotide 810ccaacgaaag cca 1381113RNAArtificial
Sequencesynthetic oligonucleotide 811cuggucacug auu
1381213RNAArtificial Sequencesynthetic oligonucleotide
812ugguuuaugg acu 1381313RNAArtificial Sequencesynthetic
oligonucleotide 813gaccagagug cug 1381413RNAArtificial
Sequencesynthetic oligonucleotide 814gaugugauug aua
1381513RNAArtificial Sequencesynthetic oligonucleotide
815gucagccgug aau 1381613RNAArtificial Sequencesynthetic
oligonucleotide 816aauguguauc uau 1381713RNAArtificial
Sequencesynthetic oligonucleotide 817uugagucugg aaa
1381813RNAArtificial Sequencesynthetic oligonucleotide
818guccagcaau uaa 1381913RNAArtificial Sequencesynthetic
oligonucleotide 819ccagcaauua aua 1382013RNAArtificial
Sequencesynthetic oligonucleotide 820gacucgaacg acu
1382113RNAArtificial Sequencesynthetic oligonucleotide
821accugccagc aac 13
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022
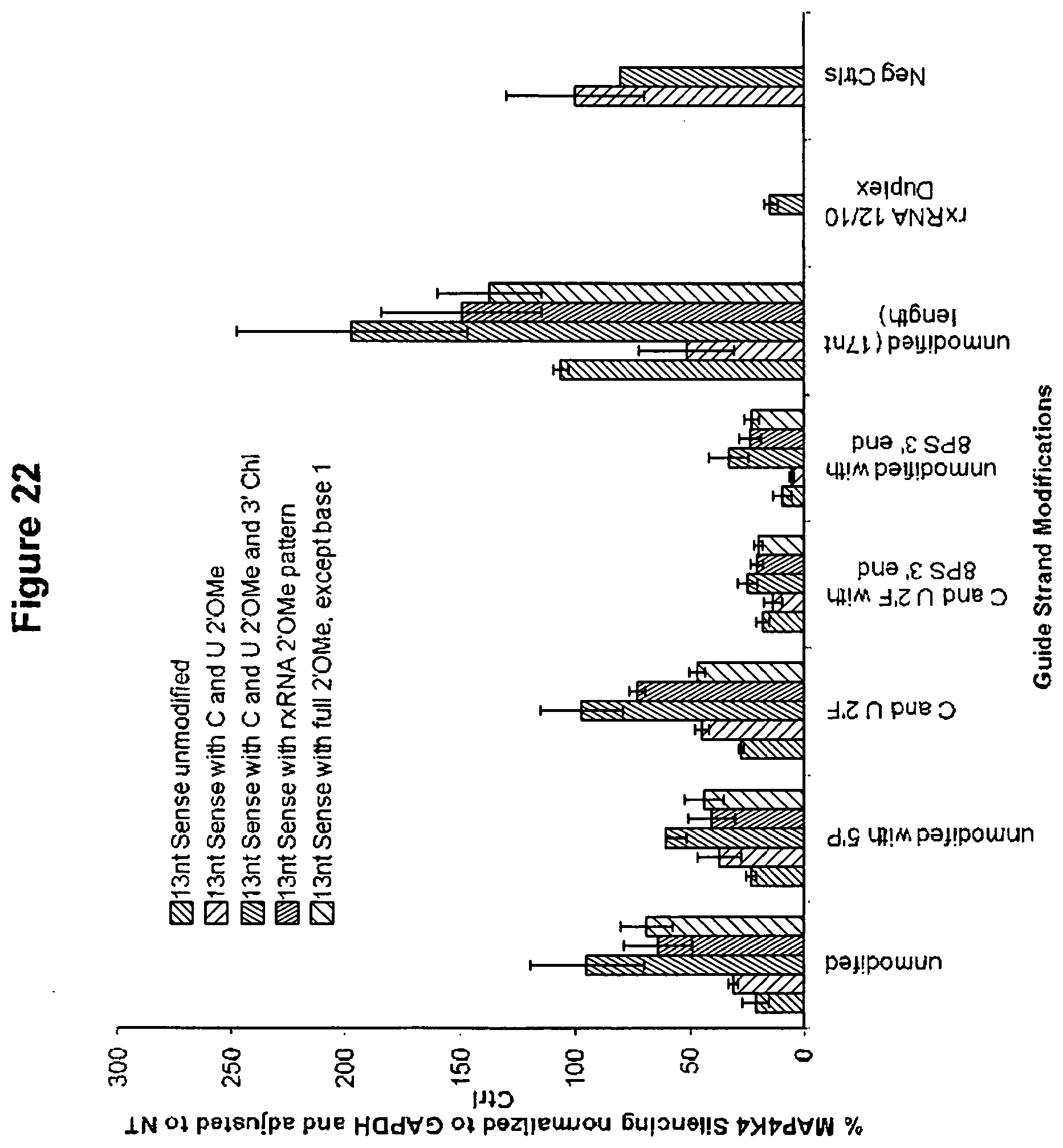
D00023

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

D00032

D00033

D00034

D00035

D00036

D00037

D00038

D00039

D00040

D00041

D00042

D00043

D00044

D00045
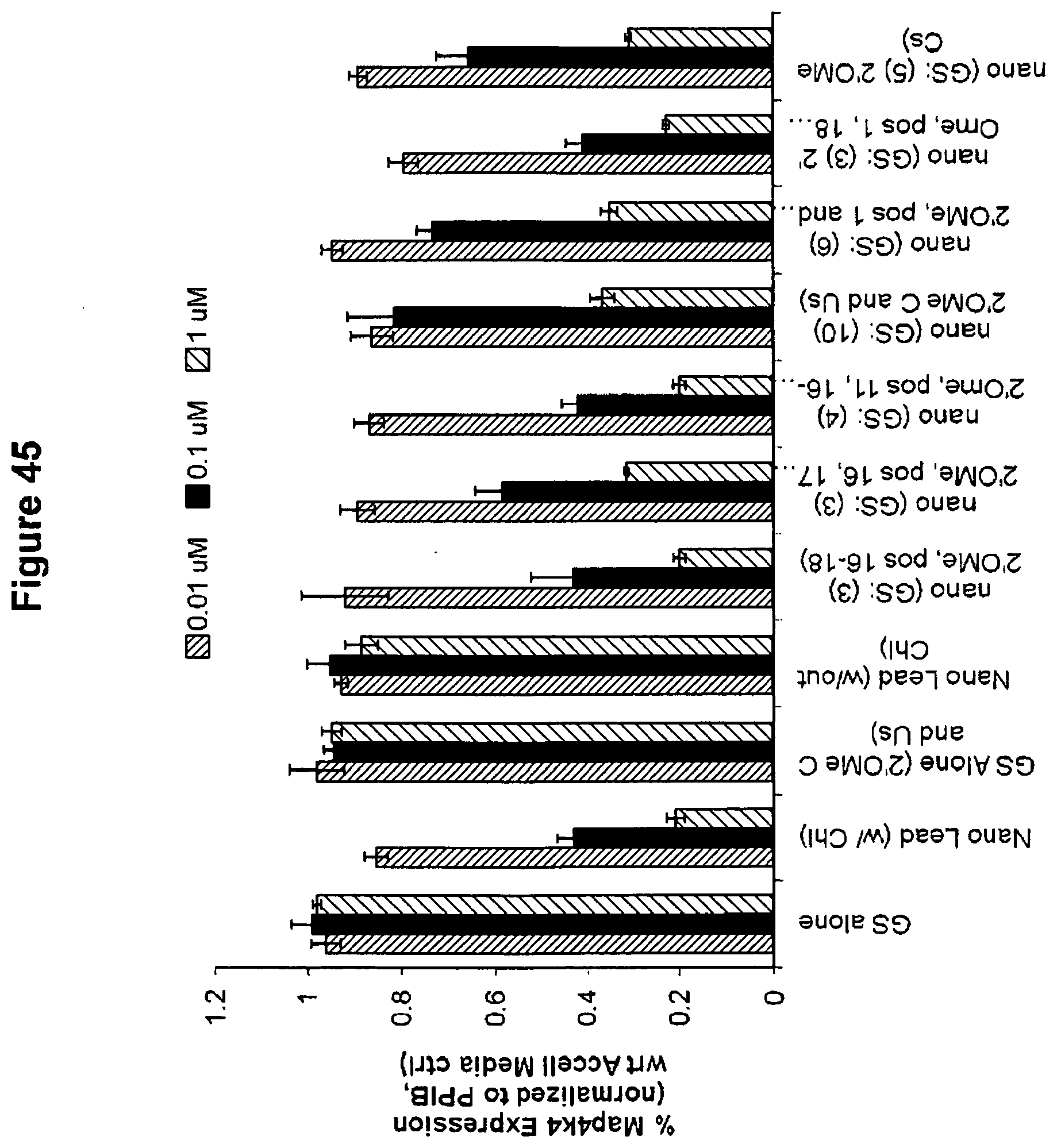
D00046

D00047

D00048

D00049

D00050

D00051
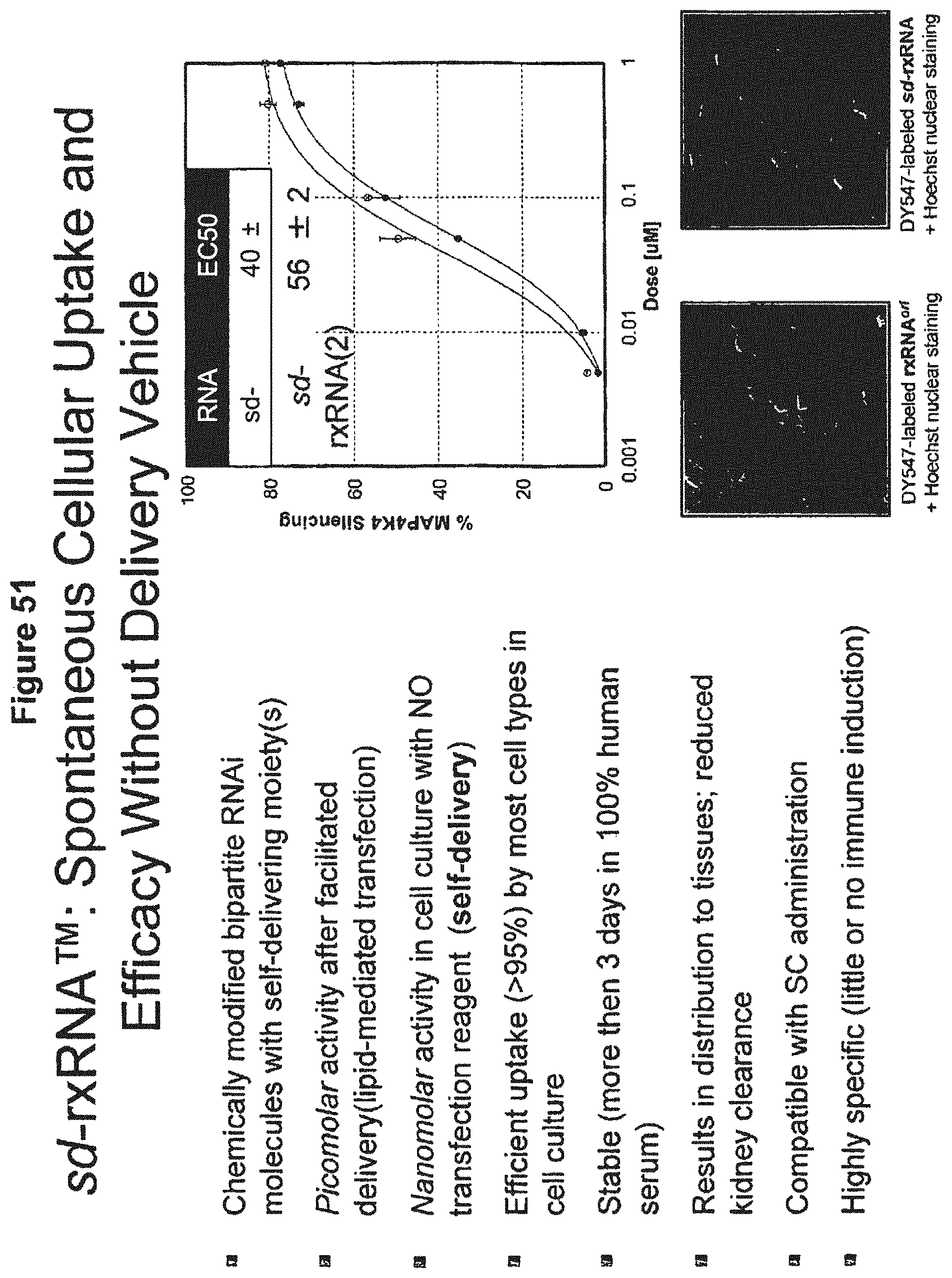
D00052

D00053

D00054

D00055

D00056

D00057

D00058

D00059

D00060

D00061

D00062

D00063

D00064

D00065

D00066

D00067

D00068

D00069

D00070

D00071

D00072

D00073

D00074

D00075

D00076

D00077

D00078

D00079

D00080

D00081

D00082

D00083

D00084

D00085

D00086

D00087

D00088

D00089

D00090

D00091

D00092

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.