Compositions And Methods Of Phospholipase A2 Receptor Chimeric Autoantibody Receptor T Cells
Gill; Saar ; et al.
U.S. patent application number 17/051248 was filed with the patent office on 2021-04-01 for compositions and methods of phospholipase a2 receptor chimeric autoantibody receptor t cells. The applicant listed for this patent is THE TRUSTEES OF THE UNIVERSITY OF PENNSYLVANIA. Invention is credited to Saar Gill, Jonathan Hogan, Aimee S. Payne, Baomei Wang.
| Application Number | 20210095001 17/051248 |
| Document ID | / |
| Family ID | 1000005302915 |
| Filed Date | 2021-04-01 |
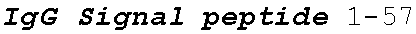




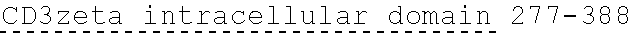
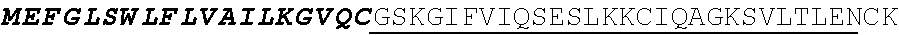





View All Diagrams
| United States Patent Application | 20210095001 |
| Kind Code | A1 |
| Gill; Saar ; et al. | April 1, 2021 |
COMPOSITIONS AND METHODS OF PHOSPHOLIPASE A2 RECEPTOR CHIMERIC AUTOANTIBODY RECEPTOR T CELLS
Abstract
The invention includes compositions comprising at least one chimeric autoantibody receptor (CAAR) specific for an anti-phospholipase A2 receptor (PLA2R) autoantibody-based B cell receptor, polynucleotides encoding the CAAR, vectors comprising a polynucleotide encoding the CAAR, and recombinant T cells comprising the CAAR. The invention also includes methods of making a genetically modified cell, e.g., a genetically modified T cell, expressing a PLA2R-CAAR wherein the expressed CAAR comprises a PLA2R extracellular domain.
| Inventors: | Gill; Saar; (Philadelphia, PA) ; Hogan; Jonathan; (Haddonfield, NJ) ; Payne; Aimee S.; (Merion Station, PA) ; Wang; Baomei; (Philadelphia, PA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005302915 | ||||||||||
| Appl. No.: | 17/051248 | ||||||||||
| Filed: | May 2, 2019 | ||||||||||
| PCT Filed: | May 2, 2019 | ||||||||||
| PCT NO: | PCT/US19/30459 | ||||||||||
| 371 Date: | October 28, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 62665863 | May 2, 2018 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 14/70578 20130101; C07K 14/70517 20130101; A61K 35/17 20130101; C07K 14/7056 20130101; C07K 2319/03 20130101; C07K 14/7051 20130101; C07K 2319/02 20130101 |
| International Class: | C07K 14/705 20060101 C07K014/705; C07K 14/725 20060101 C07K014/725; A61K 35/17 20060101 A61K035/17 |
Claims
1. A polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the CAAR comprises a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain.
2. The polynucleotide of claim 1, wherein the PLA2R autoantigen or fragment thereof is selected from the group consisting of: (a) an extracellular domain comprising a CysR or ricin B type lectin domain, a fibronectin type II domain, a C-type lectin domain 1 and a C-type lectin domain 2; and, (b) an extracellular domain comprising a CysR or ricin B type lectin domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2 and a C-type lectin domain 3.
3. The polynucleotide of claim 2, wherein the PLA2R extracellular domain (a) is encoded by a nucleic acid sequence comprising SEQ ID NO: 8, or the PLA2R extracellular domain (b) is encoded by a nucleic acid sequence comprising SEQ ID NO: 15.
4. The polynucleotide of claim 1, wherein the PLA2R autoantigen or fragment thereof is selected from the group consisting of: (a) an extracellular domain comprising a cysteine rich domain, (b) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, and a C-type lectin domain 1, (c) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, and a C-type lectin domain 3, (d) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, a C-type lectin domain 3, and a C-type lectin domain 7, (e) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, and a C-type lectin domain 7, and (f) an extracellular domain comprising a cysteine rich domain, a C-type lectin domain 1, and a C-type lectin domain 7.
5. The polynucleotide of claim 4, wherein the PLA2R extracellular domain (a) is encoded by a nucleic acid sequence comprising SEQ ID NO: 47 or SEQ ID NO: 60, the PLA2R extracellular domain (b) is encoded by a nucleic acid sequence comprising SEQ ID NO: 50 or SEQ ID NO: 62, the PLA2R extracellular domain (c) is encoded by a nucleic acid sequence comprising SEQ ID NO: 52, the PLA2R extracellular domain (d) is encoded by a nucleic acid sequence comprising SEQ ID NO: 54, the PLA2R extracellular domain (e) is encoded by a nucleic acid sequence comprising SEQ ID NO: 56, or the PLA2R extracellular domain (f) is encoded by a nucleic acid sequence comprising SEQ ID NO: 58.
6. The polynucleotide of claim 1, wherein the transmembrane domain comprises a CD8 alpha chain transmembrane domain.
7. The polynucleotide of claim 6, wherein the CD8 alpha chain transmembrane domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 20.
8. The polynucleotide of claim 6, wherein the CD8 alpha chain transmembrane domain comprises the amino acid sequence of SEQ ID NO: 19.
9. The polynucleotide of claim 1, wherein the CAAR further comprises a hinge domain.
10. The polynucleotide of claim 9, wherein the hinge domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 10 or SEQ ID NO: 42 or SEQ ID NO: 64.
11. The polynucleotide of claim 9, wherein the hinge domain comprises the amino acid sequence of SEQ ID NO: SEQ ID NO: 43 or SEQ ID NO: 44.
12. The polynucleotide of claim 1, wherein the CAAR further comprises a GS linker.
13. The polynucleotide of claim 12, wherein the GS linker is encoded by a nucleic acid sequence comprising SEQ ID NO: 68.
14. The polynucleotide of claim 12, wherein the GS linker comprises SEQ ID NO: 69.
15. The polynucleotide of claim 1, wherein the intracellular domain of a costimulatory molecule comprises 4-1BB.
16. The polynucleotide of claim 15, wherein the 4-1BB intracellular domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 22 or SEQ ID NO: 66.
17. The polynucleotide of claim 15, wherein the 4-1BB intracellular domain comprises the amino acid sequence of SEQ ID NO: 21.
18. The polynucleotide of claim 1, wherein the signaling domain comprises a CD3 zeta signaling domain.
19. The polynucleotide of claim 18, wherein the CD3 zeta signaling domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 24 or SEQ ID NO: 72.
20. The polynucleotide of claim 18, wherein the CD3 zeta signaling domain comprises an amino acid sequence of SEQ ID NO: 23 or SEQ ID NO: 45.
21. The polynucleotide of claim 1, wherein the CAAR is encoded by a nucleic acid sequence selected from the group consisting of SEQ ID NOs: 11, 17, 25, 27, 29, 31, 33, 35, 37, and 39.
22. The polynucleotide of claim 1, wherein the CAAR comprises an amino acid sequence selected from the group consisting of SEQ ID NOs: 12, 18, 26, 28, 30, 32, 34, 36, 38, and 40.
23. The polynucleotide of claim 1, wherein the CAAR comprises a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain, optionally wherein the KIR is KIRS2 or KIR2DS2.
24. A vector comprising the polynucleotide of claim 1.
25. The vector of claim 24, wherein the vector is a lentiviral vector.
26. The vector of claim 24, wherein the vector is a RNA vector.
27. (canceled)
28. A chimeric autoantibody receptor (CAAR) comprising an extracellular domain comprising a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain.
29. The CAAR of claim 28, wherein the PLA2R autoantigen or fragment thereof is selected from the group consisting of: (a) an extracellular domain comprising a ricin B type lectin domain, a fibronectin type II domain, a C-type lectin domain 1 and a C-type lectin domain 2; and, (b) an extracellular domain comprising a ricin B type lectin domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2 and a C-type lectin domain 3.
30. The CAAR of claim 29, wherein the PLA2R extracellular domain (a) is encoded by a nucleic acid sequence comprising SEQ ID NO: 8, or the PLA2R extracellular domain (b) is encoded by a nucleic acid sequence comprising SEQ ID NO: 15.
31. The CAAR of claim 28, wherein the PLA2R autoantigen or fragment thereof is selected from the group consisting of: (a) an extracellular domain comprising a cysteine rich domain, (b) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, and a C-type lectin domain 1, (c) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, a C-type lectin domain 3, and a C-type lectin domain 7, (d) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, a C-type lectin domain 3, and a C-type lectin domain 7, (e) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, and a C-type lectin domain 7, and (f) an extracellular domain comprising a cysteine rich domain, a C-type lectin domain 1, and a C-type lectin domain 7.
32. The CAAR of claim 31, wherein the PLA2R extracellular domain (a) comprises SEQ ID NO: 49 or SEQ ID NO: 61, the PLA2R extracellular domain (b) comprises the amino acid sequence of SEQ ID NO: 51 or SEQ ID NO: 63, the PLA2R extracellular domain (c) comprises the amino acid sequence of SEQ ID NO: 53, the PLA2R extracellular domain (d) comprises the amino acid sequence of SEQ ID NO: 55, or the PLA2R extracellular domain (e) comprises the amino acid sequence of SEQ ID NO: 57, or the PLA2R extracellular domain (f) comprises the amino acid sequence of SEQ ID NO: 59.
33. The CAAR of claim 28, wherein the transmembrane domain comprises a CD8 alpha chain transmembrane domain.
34. The CAAR of claim 33, wherein the CD8 alpha chain transmembrane domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 20.
35. The CAAR of claim 33, wherein the CD8 alpha chain transmembrane domain comprises the amino acid sequence of SEQ ID NO: 19.
36. The CAAR of claim 28, wherein the CAAR further comprises a hinge domain.
37. The CAAR of claim 36, wherein the hinge domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 10 or SEQ ID NO: 42 or SEQ ID NO: 64.
38. The CAAR of claim 36, wherein the hinge domain comprises the amino acid sequence of SEQ ID NO: SEQ ID NO: 43 or SEQ ID NO: 44.
39. The CAAR of claim 28, wherein the CAAR further comprises a GS linker.
40. The CAAR of claim 39, wherein the GS linker is encoded by a nucleic acid sequence comprising SEQ ID NO: 68.
41. The CAAR of claim 39, wherein the GS linker comprises SEQ ID NO: 69.
42. The CAAR of claim 28, wherein the intracellular domain of a costimulatory molecule comprises 4-1BB.
43. The CAAR of claim 42, wherein the 4-1BB intracellular domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 22 or SEQ ID NO: 66.
44. The CAAR of claim 42, wherein the 4-1BB intracellular domain comprises the amino acid sequence of SEQ ID NO: 21.
45. The CAAR of claim 28, wherein the signaling domain comprises a CD3 zeta signaling domain.
46. The CAAR of claim 45, wherein the CD3 zeta signaling domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 24 or SEQ ID NO: 72.
47. The CAAR of claim 45, wherein the CD3 zeta signaling domain comprises the amino acid sequence of SEQ ID NO: 23 or SEQ ID NO: 45.
48. The CAAR of claim 28, wherein the CAAR is encoded by a nucleic acid sequence selected from the group consisting of SEQ ID NOs: 11, 17, 25, 27, 29, 31, 33, 35, 37, and 39.
49. The CAAR of claim 28, wherein the CAAR comprises an amino acid sequence selected from the group consisting of SEQ ID NOs: 12, 18, 26, 28, 30, 32, 34, 36, 38, and 40.
50. The CAAR of claim 28, wherein the CAAR comprises an extracellular domain comprising phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain, optionally wherein the KIR is KIRS2 or KIR2DS2.
51. A genetically modified cell comprising the CAAR of claim 28.
52. The cell of claim 51, wherein the cell expresses the CAAR and has high affinity to autoantibody-based BCRs on B cells.
53. The cell of claim 51, wherein the cell expresses the CAAR and induces killing of B cells expressing autoantibodies.
54. The cell of claim 51, wherein the cell expresses the CAAR and has limited toxicity toward healthy cells.
55. The cell of claim 51, wherein the cell is selected from the group consisting of a helper T cell, a cytotoxic T cell, a memory T cell, regulatory T cell, gamma delta T cell, a natural killer cell, a cytokine induced killer cell, a cell line thereof, a T memory stem cell, a T cell derived from a pluripotent stem and other effector cell.
56. The cell of claim 51, wherein the CAAR comprises an extracellular domain comprising a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain, optionally wherein the KIR is KIRS2 or KIR2DS2; and wherein the cell further comprises DAP12.
57. A pharmaceutical composition comprising the polynucleotide of claim 1, and a pharmaceutically acceptable excipient.
58. A method for (a) preventing glomerulus damage in a subject at risk of developing an autoantibody-mediated kidney disease, (b) reducing glomerulus damage in a subject suffering from an autoantibody-mediated kidney disease, and/or (c) treating an autoantibody-mediated kidney disease in a subject, the method comprising: administering to the subject an effective amount of a genetically modified T cell comprising the polynucleotide of claim 1.
59. (canceled)
60. The method of claim 58, wherein the autoantibody mediated kidney disease is selected from the group consisting of a glomerular disease and a primary membranous nephropathy.
61. The method of claim 58, wherein the subject is a human.
62. The method of claim 58, wherein the modified T cell targets B cells.
63. The method of claim 58, wherein the CAAR comprises an extracellular domain comprising phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain, optionally wherein the KIR is KIRS2 or KIR2DS2; and wherein the cell further comprises a polynucleotide encoding DAP12.
64. (canceled)
65. (canceled)
66. (canceled)
67. (canceled)
68. The vector of claim 24, wherein the vector comprises an inducible promoter operably linked to the polynucleotide encoding the CAAR.
69. The cell of claim 51, wherein the cell comprises a polynucleotide encoding the CAAR operably linked to an inducible promoter.
70. A pharmaceutical composition comprising the CAAR of claim 28 and a pharmaceutically acceptable excipient.
71. A pharmaceutical composition comprising the cell of claim 51, and a pharmaceutically acceptable excipient.
Description
CROSS-REFERENCE TO RELATED APPLICATION
[0001] The present application is entitled to priority under 35 U.S.C. .sctn. 119(e) to U.S. Provisional Patent Application No. 62/665,863 filed May 2, 2018, which is hereby incorporated by reference in its entirety herein.
BACKGROUND OF THE INVENTION
[0002] Membranous nephropathy (MN) is among the most common primary causes of nephrotic syndrome in adults (up to one-third of cases). About 15-25% of membranous nephropathy cases are secondary membranous nephropathy, caused by drugs, infections, tumors, or immune diseases. The remaining 75-85% of membranous nephropathy cases are idiopathic, also called primary membranous nephropathy. MN is caused by immune complex formation in the glomerulus. The immune complexes are formed by binding of antibodies to antigens on the podocyte. The immune complex serves as an activator that triggers complement-mediated lysis of glomerular epithelial cells and the release of proteases and oxidants damaging capillary walls. Up to 40% of patients with primary membranous nephropathy who do not receive immunosuppressive treatment develop end-stage renal disease requiring dialysis or kidney transplantation.
[0003] The M-type phospholipase A2 receptor (PLA2R) has been described as a major autoantigen in primary membranous nephropathy. Autoantibodies to PLA2R are present in 70-80% of cases of primary membranous nephropathy, and can be used for the diagnosis and monitoring of treatment of primary membranous nephropathy both in the native kidneys and after kidney transplant.
[0004] Current guidelines suggest treatment with alkylating agents or calcineurin inhibitors as first-line therapy for the treatment of severe, primary membranous nephropathy. Rituximab, an anti-CD20 antibody, has also shown promise and is being evaluated for treating MN. However, since these treatments lack specificity for the autoreactive B cells that produce the serum anti-PLA2R antibody, they can be associated with risk of life-threatening infections. Moreover, relapse can occur in patients who achieve remission in proteinuria, and disease can recur after kidney transplantation. Importantly, relapse is usually associated with recurrence in detectable anti-PLA2R autoantibody, thus further supporting the role of this antibody in disease pathogenesis.
[0005] There is an urgent need in the art for achieving a more specific and effective treatment for primary membranous nephropathy. This invention addresses this need.
SUMMARY OF THE INVENTION
[0006] In one aspect, the invention includes a polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the CAAR comprises a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain.
[0007] In another aspect, the invention includes a vector comprising any one of the polynucleotides disclosed herein.
[0008] In yet another aspect, the invention includes a chimeric autoantibody receptor (CAAR) comprising an extracellular domain comprising a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof.
[0009] In still another aspect, the invention includes a chimeric autoantibody receptor (CAAR) comprising an extracellular domain comprising a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain.
[0010] Another aspect of the invention includes a genetically modified cell comprising any one of the CAARs disclosed herein.
[0011] Yet another aspect of the invention includes genetically modified cell comprising: (a) a chimeric autoantibody receptor (CAAR) comprising an extracellular domain comprising a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain; and (b) DAP12.
[0012] Still another aspect of the invention includes a pharmaceutical composition comprising any one of the polynucleotides disclosed herein, any one of the CAARs disclosed herein or any one of the cells disclosed herein, and a pharmaceutically acceptable excipient.
[0013] In another aspect, the invention includes a method for treating an autoantibody-mediated kidney disease in a subject. The method comprises administering to the subject an effective amount of a genetically modified T cell comprising a polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the polynucleotide encodes a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain, thereby treating the autoantibody mediated kidney disease in the subject.
[0014] In yet another aspect, the invention includes a method for preventing or reducing glomerulus damage in a subject at risk of or suffering from an autoantibody-mediated kidney disease, the method comprising: administering to the subject an effective amount of a genetically modified T cell comprising a polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the polynucleotide encodes a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain, thereby preventing or reducing glomerulus damage in the subject.
[0015] In still another aspect, the invention includes a method for treating an autoantibody-mediated kidney disease in a subject. The method comprises administering to the subject an effective amount of a genetically modified T cell comprising: (a) a polynucleotide encoding a chimeric autoantibody receptor (CAAR) comprising an extracellular domain comprising phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain; and (b) a polynucleotide encoding DAP12, thereby treating the autoantibody-mediated kidney disease in the subject.
[0016] In another aspect, the invention includes a method for preventing or reducing glomerulus damage in a subject at risk of or suffering from an autoantibody-mediated kidney disease The method comprises administering to the subject an effective amount of a genetically modified T cell comprising: (a) a polynucleotide encoding a chimeric autoantibody receptor (CAAR) comprising an extracellular domain comprising phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain; and (b) a polynucleotide encoding DAP12, thereby preventing or reducing glomerulus damage in the subject.
[0017] In various embodiments of the above aspects or any other aspect of the invention delineated herein, the PLA2R autoantigen or fragment thereof is selected from the group consisting of: (a) an extracellular domain comprising a CysR or ricin B type lectin domain, a fibronectin type II domain, a C-type lectin domain 1 and a C-type lectin domain 2; and (b) an extracellular domain comprising a CysR or ricin B type lectin domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2 and a C-type lectin domain 3.
[0018] In certain embodiments, the PLA2R autoantigen or fragment thereof is selected from the group consisting of: (a) an extracellular domain comprising a cysteine rich domain, (b) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, and a C-type lectin domain 1, (c) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, and a C-type lectin domain 3, (d) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, a C-type lectin domain 3, and a C-type lectin domain 7, (e) an extracellular domain comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, and a C-type lectin domain 7, or (f) an extracellular domain comprising a cysteine rich domain, a C-type lectin domain 1, and a C-type lectin domain 7.
[0019] In certain embodiments, the PLA2R extracellular domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 8 or SEQ ID NO: 15 or SEQ ID NO: 47 or SEQ ID NO: 60 or SEQ ID NO: 50 or SEQ ID NO: 62 or SEQ ID NO: 52 or SEQ ID NO: 54 or SEQ ID NO: 56 or SEQ ID NO: 58.
[0020] In certain embodiments, the transmembrane domain comprises a CD8 alpha chain transmembrane domain. In certain embodiments, the CD8 alpha chain transmembrane domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 20. In certain embodiments, the CD8 alpha chain transmembrane domain comprises the amino acid sequence of SEQ ID NO: 19.
[0021] In certain embodiments, the CAAR further comprises a hinge domain. In certain embodiments, the hinge domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 10 or SEQ ID NO: 42 or SEQ ID NO: 64. In certain embodiments, the hinge domain comprises the amino acid sequence of SEQ ID NO: SEQ ID NO: 43 or SEQ ID NO: 44.
[0022] In certain embodiments, the CAAR further comprises a GS linker. In certain embodiments, the GS linker is encoded by a nucleic acid sequence comprising SEQ ID NO: 68. In certain embodiments, the GS linker comprises SEQ ID NO: 69.
[0023] In certain embodiments, the intracellular domain of a costimulatory molecule comprises 4-1BB. In certain embodiments, the 4-1BB intracellular domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 22 or SEQ ID NO: 66. In certain embodiments, the 4-1BB intracellular domain comprises the amino acid sequence of SEQ ID NO: 21.
[0024] In certain embodiments, the signaling domain comprises a CD3 zeta signaling domain. In certain embodiments, the CD3 zeta signaling domain is encoded by a nucleic acid sequence comprising SEQ ID NO: 24 or SEQ ID NO: 72. In certain embodiments, the CD3 zeta signaling domain comprises an amino acid sequence of SEQ ID NO: 23 or SEQ ID NO: 45.
[0025] In certain embodiments, the CAAR is encoded by a nucleic acid sequence selected from the group consisting of SEQ ID NOs: 11, 17, 25, 27, 29, 31, 33, 35, 37, and 39. In certain embodiments, the CAAR comprises an amino acid sequence selected from the group consisting of SEQ ID NOs: 12, 18, 26, 28, 30, 32, 34, 36, 38, and 40.
[0026] In certain embodiments, the CAAR comprises a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, a killer immunoglobulin-like receptor (KIR) transmembrane domain and a KIR cytoplasmic domain.
[0027] In certain embodiments, the vector is a lentiviral vector. In certain embodiments, the vector is a RNA vector.
[0028] In certain embodiments, the cell expresses the CAAR and has high affinity to autoantibody-based BCRs on B cells. In certain embodiments, the cell expresses the CAAR and induces killing of B cells expressing autoantibodies. In certain embodiments, the cell expresses the CAAR and has limited toxicity toward healthy cells.
[0029] In certain embodiments, the cell is selected from the group consisting of a helper T cell, a cytotoxic T cell, a memory T cell, regulatory T cell, gamma delta T cell, a natural killer cell, a cytokine induced killer cell, a cell line thereof, a T memory stem cell, a T cell derived from a pluripotent stem and other effector cell.
[0030] In certain embodiments, the autoantibody mediated kidney disease is selected from the group consisting of a glomerular disease and a primary membranous nephropathy.
[0031] In certain embodiments, the subject is a human.
[0032] In certain embodiments, the modified T cell targets B cells.
[0033] In certain embodiments, the KIR is KIRS2 or KIR2DS2.
[0034] In certain embodiments, the vector comprises an inducible promoter operably linked to the polynucleotide encoding the CAAR.
[0035] In certain embodiments, the cell comprises a polynucleotide encoding the CAAR operably linked to an inducible promoter.
BRIEF DESCRIPTION OF THE DRAWINGS
[0036] The following detailed description of preferred embodiments of the invention will be better understood when read in conjunction with the appended drawings. For the purpose of illustrating the invention, there are shown in the drawings embodiments which are presently preferred. It should be understood, however, that the invention is not limited to the precise arrangements and instrumentalities of the embodiments shown in the drawings.
[0037] FIG. 1 is an illustration showing a schematic of the native PLA2R molecule. The N terminal cysteine-rich (CysR) domain (Ricin B type lectin domain) is the immunodominant epitope that is recognized by patient serum antibodies. In non-reducing conditions, serum antibody recognition has been reported to be dependent on the first 3 C-type lectin domains (CTLD). In addition to the cysteine-rich domain, serum antibody immunoreactivity to the C-type lectin domains 1 and 7 has also been reported. A commercial Novus monoclonal antibody (mAb) and polyclonal Sigma antibody bind to the CTLD2 domain.
[0038] FIGS. 2A-2G are a series of schematic diagrams illustrating the various PLA2R CAAR constructs described in this invention. Abbreviations for here and throughout: ECD: extracellular domain; TMD: transmembrane domain; BBZ: cytoplasmic tandem 41BB (CD137) and CD3-zeta signaling domain; CysR=cysteine rich domain; FNII=fibronectin type II domain; CTLD=C-type lectin domain. All constructs use a CD8 transmembrane domain and BBZ cytoplasmic domain and differ in their extracellular composition as follows: FIG. 2A depicts construct 4025.C (or construct C), whose extracellular domain consists of the PLA2R CysR domain, followed by a CD8 hinge domain. FIG. 2B depicts construct 4026.CF1 (or construct CF1), consisting of the CysR, FNII, and CTLD1 domains followed by a CD8 hinge domain. FIG. 2C depicts construct 4027.CF12, consisting of the CysR, FNII, CTLD1, and CTLD2 domains, followed by a CD8 hinge domain. FIG. 2D depicts construct 4028.CF123 (or construct CF123), consisting of the CysR, FNII, CTLD1, CTLD2, and CTLD3 domains, followed by either a CD8 hinge (4028.CF123) or GS linker (CF123). FIG. 2E depicts construct CF1237, consisting of the CysR, FNII, CTLD1, CTLD2, CTLD3, and CTLD7 domains followed by a GS linker. FIG. 2F depicts construct CF17, consisting of the CysR, FNII, CTLD1, and CTLD7 domains, followed by a GS linker. FIG. 2G depicts construct C17, consisting of the CysR, CTLD1, and CTLD7 domains, followed by a GS linker.
[0039] FIG. 3A is an illustration showing a portion of the plasmid map comprising the 4028.CF123 CAAR. FIG. 3B is a table distinguishing the compositions of the CAAR constructs described in this invention.
[0040] FIG. 4 is a series of flow cytometry plots demonstrating surface expression of a subset of PLA2R CAARs (4027.CF12 and 4028.CF123) on Jurkat NFAT-GFP cells. The expression of PLA2R CAAR by Jurkat NFAT-GFP cells transduced with different PLA2R lentiviral constructs was measured (MOI 5:1, 48 h) using a commercial Novus monoclonal antibody (mAb) specific for the PLA2R CTLD2 domain. Jurkat T cells expressing construct 4027.CF12 or 4028.CF123, but not those T cells bearing 4025.C, which does not include the CTLD2 domain and hence is not detectable by the mAb used, or untransduced Jurkat NFAT-GFP cells, exhibited detectable surface expression of the CAAR.
[0041] FIG. 5 is a series of flow cytometry plots showing the negative and positive controls for PLA2R CAAR T cell activation using an isotype control (top, negative control) or PMA+ionomycin (bottom, positive control).
[0042] FIG. 6 is a series of flow cytometry plots showing the activation of 4027.CF12 and 4028.CF123 PLA2R CAAR T cells by plate-bound monoclonal or polyclonal antibodies specific for the PLA2R CTLD2 domain, measured as GFP expression by Jurkat NFAT-GFP cells (6 h activation). No evidence of T cell activation is seen in construct 4025.C, as expected as this construct does not contain the CTLD2 domain, and untransduced cells upon exposure to the anti-PLA2R monoclonal antibody (mAb) or the polyclonal anti-PLA2R antibody (Ab). Specific T cell activation in response to cross-linking of the PLA2R CAAR by the monoclonal anti-PLA2R mAb but not the polyclonal anti-PLA2R Ab is observed.
[0043] FIGS. 7A-7B are a series of plots illustrating that PLA2R MN patient IgG detects PLA2R CAAR expression on transduced Jurkat NFAT-GFP and primary human T cells. Flow cytometry plots demonstrate robust PLA2R CAAR surface expression on Jurkat NFAT-GFP cells (FIG. 7A) and primary human T cells (FIG. 7B) for all constructs evaluated (4025.C, 4026.CF1, 4027.CF12, 4028.CF123, C, CF1, CF123, CF1237, CF17, C17). The expression of PLA2R CAAR by Jurkat NFAT-GFP and primary human T cells transduced with different PLA2R lentiviral constructs (72 h) was detected by incubation with MN patient IgG followed by APC-conjugated anti-human IgG.
[0044] FIG. 8 is a series of plots illustrating that plate-bound MN patient IgG activates PLA2R CAAR transduced Jurkat-NFAT-GFP cells. Flow cytometry plots show the activation of PLA2R CAAR T cells by MN patient IgG measured as GFP expression by Jurkat NFAT-GFP cells (6 h activation). All evaluated PLA2R CAAR T cells (4025.C, 4026.CF1, 4027.CF12, 4028.CF123, C, CF1, CF123, CF1237, CF17, C17) transduced signal after cross-linking by plate-bound anti-PLA2R IgG.
[0045] FIG. 9 is a set of graphs illustrating that plate-bound MN patient IgG stimulates IFN-.gamma. secretion from PLA2R CAAR-transduced primary human T cells. 5.times.10.sup.4 primary human T cells were cultured for 24 h in a 96-well plate coated with 10 .mu.g/ml of either MN patient IgG or control human IgG. Culture supernatants were then harvested for detection of IFN-.gamma. by ELISA. IFN.gamma. production was detectable in all culture supernatants and was elevated relative to untransduced T cells, or PLA2R CAAR T cells stimulated with control (Ctrl) human IgG.
DETAILED DESCRIPTION
[0046] The invention includes a chimeric autoantibody receptor (CAAR) specific for anti-phospholipase A2 receptor (PLA2R) B cell receptor (BCR), compositions comprising the CAAR, polynucleotides encoding the CAAR, vectors comprising a polynucleotide encoding the CAAR, and recombinant T cells comprising the CAAR. The invention also includes methods of making a genetically modified cell, e.g., a genetically modified T cell, expressing a PLA2R-CAAR wherein the expressed CAAR comprises a PLA2R extracellular domain.
[0047] The present invention also relates generally to the use of cells, e.g., T cells, engineered to express a CAAR to treat an autoantibody-mediated kidney disease associated with targeting of self-antigens (e.g. PLA2R). In one embodiment, the cells, e.g., T cells expressing the CAAR of the invention specifically bind to and kill anti-PLA2R BCR-expressing cells, but do not bind to and kill normal BCR-expressing cells.
Definitions
[0048] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which the invention pertains. Although any methods and materials similar or equivalent to those described herein can be used in the practice of and/or for the testing of the present invention, the preferred materials and methods are described herein. In describing and claiming the present invention, the following terminology will be used according to how it is defined, where a definition is provided.
[0049] It is also to be understood that the terminology used herein is for the purpose of describing particular embodiments only, and is not intended to be limiting.
[0050] The articles "a" and "an" are used herein to refer to one or to more than one (i.e., to at least one) of the grammatical object of the article. By way of example, "an element" means one element or more than one element.
[0051] "About," as used herein, when referring to a measurable value such as an amount, a temporal duration, and the like, is meant to encompass variations of .+-.20% or .+-.10%, in some instances .+-.5%, in some instances .+-.1%, and in some instance .+-.0.1% from the specified value, as such variations are appropriate to perform the disclosed methods.
[0052] The term "antibody," as used herein, refers to an immunoglobulin molecule that binds with an antigen. Antibodies can be intact immunoglobulins derived from natural sources or from recombinant sources and can be immunoreactive portions of intact immunoglobulins. Antibodies are typically tetramers of immunoglobulin molecules. The antibody in the present invention may exist in a variety of forms where the antibody is expressed as part of a contiguous polypeptide chain including, for example, a single domain antibody fragment (sdAb), a single chain antibody (scFv) and a humanized antibody (Harlow et al., 1999, In: Using Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory Press, NY; Harlow et al., 1989, In: Antibodies: A Laboratory Manual, Cold Spring Harbor, N.Y.; Houston et al., 1988, Proc. Natl. Acad. Sci. USA 85:5879-5883; Bird et al., 1988, Science 242:423-426).
[0053] The term "high affinity," as used herein, refers to high specificity in binding or interacting or attraction of a binding molecule to a target molecule. For example, in some embodiments, the binding molecule may have an affinity for the target molecule stronger than 100 nM, 50 nM, 20 nM, 15 nM, 10 nM, 9 nM, 8 nM, 7 nM, 6 nM, 5 nM, 4 nM, 3 nM, 2 nM, or 1 nM, e.g., as determined by surface plasmon resonance.
[0054] The term "antigen" or "Ag," as used herein, is defined as a molecule that provokes an immune response. This immune response may involve either antibody production, or the activation of specific immunologically competent cells, or both. The skilled artisan will understand that any macromolecule, including virtually all proteins or peptides, can serve as an antigen. Furthermore, antigens can be derived from recombinant or genomic DNA. A skilled artisan will understand that any DNA, which comprises a nucleotide sequences or a partial nucleotide sequence encoding a protein that elicits an immune response therefore encodes an "antigen" as that term is used herein. Furthermore, one skilled in the art will understand that an antigen need not be encoded solely by a full length nucleotide sequence of a gene. It is readily apparent that the present invention includes, but is not limited to, the use of partial nucleotide sequences of more than one gene and that these nucleotide sequences are arranged in various combinations to encode polypeptides that elicit the desired immune response. Moreover, a skilled artisan will understand that an antigen need not be encoded by a "gene" at all. It is readily apparent that an antigen can be generated synthesized or can be derived from a biological sample. Such a biological sample can include, but is not limited to a tissue sample, a tumor sample, a cell or a biological fluid.
[0055] By "autoantigen" is meant an endogenous antigen that stimulates production of an autoimmune response, such as production of autoantibodies. Autoantigen also includes a self-antigen or antigen from a normal tissue that is the target of a cell-mediated or an antibody-mediated immune response that may result in the development of an autoimmune disease. Examples of autoantigens include, but are not limited to, PLA2R, and fragments thereof.
[0056] The term "limited toxicity," as used herein, refers to the peptides, polynucleotides, cells and/or antibodies of the invention manifesting a lack of substantially negative biological effects, anti-tumor effects, or substantially negative physiological symptoms toward a healthy cell, non-tumor cell, non-diseased cell, non-target cell or population of such cells either in vitro or in vivo.
[0057] "Autoantibody" refers to an antibody that is specific for an autoantigen.
[0058] The term "autoimmune disease," as used herein, is defined as a disorder or condition that results from an antibody-mediated autoimmune response against autoantigens. An autoimmune disease results in the production of autoantibodies that are inappropriately produced and/or excessively produced to a self-antigen or autoantigen.
[0059] As used herein, the term "autologous" is meant to refer to any material derived from the same individual to which it is later to be re-introduced into the individual.
[0060] "Allogeneic" refers to any material derived from a different animal of the same species.
[0061] "Xenogeneic" refers to any material derived from an animal of a different species.
[0062] "Chimeric autoantibody receptor" or "CAAR" refers to an engineered receptor that is expressed on cell, e.g., a T cell or any other effector cell type, e.g., an effector cell type capable of cell-mediated cytotoxicity. The CAAR includes an antigen or fragment thereof that is specific for an autoantibody and/or BCR, e.g., a pathogenic autoantibody and/or BCR. The CAAR optionally also includes a transmembrane domain, an intracellular domain and/or a signaling domain.
[0063] As used herein, the term "conservative sequence modifications" is intended to refer to amino acid modifications that do not significantly affect or alter the binding characteristics of the antibody containing the amino acid sequence. Such conservative modifications include amino acid substitutions, additions and deletions. Modifications can be introduced into an antibody of the invention by standard techniques known in the art, such as site-directed mutagenesis and PCR-mediated mutagenesis. Conservative amino acid substitutions are ones in which the amino acid residue is replaced with an amino acid residue having a similar side chain. Families of amino acid residues having similar side chains have been defined in the art. These families include amino acids with basic side chains (e.g., lysine, arginine, histidine), acidic side chains (e.g., aspartic acid, glutamic acid), uncharged polar side chains (e.g., glycine, asparagine, glutamine, serine, threonine, tyrosine, cysteine, tryptophan), nonpolar side chains (e.g., alanine, valine, leucine, isoleucine, proline, phenylalanine, methionine), beta-branched side chains (e.g., threonine, valine, isoleucine) and aromatic side chains (e.g., tyrosine, phenylalanine, tryptophan, histidine). Thus, for example, one or more amino acid residues within the extracellular regions of the CAAR of the invention can be replaced with other amino acid residues having a similar side chain or charge and the altered CAAR can be tested for the ability to bind autoantibodies using the functional assays described herein.
[0064] "Co-stimulatory ligand," as the term is used herein, includes a molecule on an antigen presenting cell (e.g., an aAPC, dendritic cell, B cell, and the like) that specifically binds a cognate co-stimulatory molecule on a T cell, thereby providing a signal which, in addition to the primary signal provided by, for instance, binding of a TCR/CD3 complex with an MHC molecule loaded with peptide, mediates a T cell response, including, but not limited to, proliferation, activation, differentiation, and the like.
[0065] A "co-stimulatory molecule" refers to the cognate binding partner on a T cell that specifically binds with a co-stimulatory ligand, thereby mediating a co-stimulatory response by the T cell, such as, but not limited to, proliferation. Co-stimulatory molecules include, but are not limited to an MHC class I molecule, BTLA and a Toll ligand receptor.
[0066] "Encoding" refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a cDNA, or an mRNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom. Thus, a gene encodes a protein if transcription and translation of mRNA corresponding to that gene produces the protein in a cell or other biological system. Both the coding strand, the nucleotide sequence of which is identical to the mRNA sequence and is usually provided in sequence listings, and the non-coding strand, used as the template for transcription of a gene or cDNA, can be referred to as encoding the protein or other product of that gene or cDNA.
[0067] "Effective amount" or "therapeutically effective amount" are used interchangeably herein, and refer to an amount of a compound, formulation, material, or composition, as described herein effective to achieve a particular biological result. Such results may include, but are not limited to, the inhibition of virus infection as determined by any means suitable in the art.
[0068] The term "effector function" refers to a specialized function of a cell.
[0069] As used herein, "endogenous" refers to any material from or produced inside an organism, cell, tissue or system.
[0070] As used herein, the term "exogenous" refers to any material introduced from or produced outside an organism, cell, tissue or system.
[0071] The term "expression," as used herein, is defined as the transcription and/or translation of a particular nucleotide sequence driven by a promoter.
[0072] "Expression vector" refers to a vector comprising a recombinant polynucleotide comprising expression control sequences operatively linked to a nucleotide sequence to be expressed. An expression vector comprises sufficient cis-acting elements for expression; other elements for expression can be supplied by the host cell or in an in vitro expression system. Expression vectors include all those known in the art, such as cosmids, plasmids (e.g., naked or contained in liposomes), retrotransposons (e.g. piggyback, sleeping beauty), and viruses (e.g., lentiviruses, retroviruses, adenoviruses, and adeno-associated viruses) that incorporate the recombinant polynucleotide.
[0073] "Homologous," as used herein, refers to the subunit sequence identity between two polymeric molecules, e.g., between two nucleic acid molecules, such as, two DNA molecules or two RNA molecules, or between two polypeptide molecules. When a subunit position in both of the two molecules is occupied by the same monomeric subunit; e.g., if a position in each of two DNA molecules is occupied by adenine, then they are homologous at that position. The homology between two sequences is a direct function of the number of matching or homologous positions; e.g., if half (e.g., five positions in a polymer ten subunits in length) of the positions in two sequences are homologous, the two sequences are 50% homologous; if 90% of the positions (e.g., 9 of 10), are matched or homologous, the two sequences are 90% homologous.
[0074] "Identity," as used herein, refers to the subunit sequence identity between two polymeric molecules particularly between two amino acid molecules, such as, between two polypeptide molecules. When two amino acid sequences have the same residues at the same positions; e.g., if a position in each of two polypeptide molecules is occupied by an Arginine, then they are identical at that position. The identity or extent to which two amino acid sequences have the same residues at the same positions in an alignment is often expressed as a percentage. The identity between two amino acid sequences is a direct function of the number of matching or identical positions; e.g., if half (e.g., five positions in a polymer ten amino acids in length) of the positions in two sequences are identical, the two sequences are 50% identical; if 90% of the positions (e.g., 9 of 10), are matched or identical, the two amino acids sequences are 90% identical.
[0075] As used herein, an "instructional material" includes a publication, a recording, a diagram, or any other medium of expression which can be used to communicate the usefulness of the compositions and methods of the invention. The instructional material of the kit of the invention may, for example, be affixed to a container which contains the nucleic acid, peptide, and/or composition of the invention or be shipped together with a container which contains the nucleic acid, peptide, and/or composition. Alternatively, the instructional material may be shipped separately from the container with the intention that the instructional material and the compound be used cooperatively by the recipient.
[0076] "Intracellular domain" refers to a portion or region of a molecule that resides inside a cell.
[0077] The term "intracellular signaling domain" is meant to include any full-length or truncated portion of the intracellular domain sufficient to transduce the effector function signal.
[0078] "Isolated" means altered or removed from the natural state. For example, a nucleic acid or a peptide naturally present in a living animal is not "isolated," but the same nucleic acid or peptide partially or completely separated from the coexisting materials of its natural state is "isolated." An isolated nucleic acid or protein can exist in substantially purified form, or can exist in a non-native environment such as, for example, a host cell.
[0079] In the context of the present invention, the following abbreviations for the commonly occurring nucleic acid bases are used. "A" refers to adenosine, "C" refers to cytosine, "G" refers to guanosine, "T" refers to thymidine, and "U" refers to uridine.
[0080] Unless otherwise specified, a "nucleotide sequence encoding an amino acid sequence" includes all nucleotide sequences that are degenerate versions of each other and that encode the same amino acid sequence. The phrase nucleotide sequence that encodes a protein or an RNA may also include introns to the extent that the nucleotide sequence encoding the protein may in some version contain an intron(s).
[0081] A "lentivirus," as used herein, refers to a genus of the Retroviridae family. Lentiviruses are unique among the retroviruses in being able to infect non-dividing cells; they can deliver a significant amount of genetic information into the DNA of the host cell, so they are one of the most efficient methods of a gene delivery vector. HIV, SIV, and FIV are all examples of lentiviruses. Vectors derived from lentiviruses offer the means to achieve significant levels of gene transfer in vivo.
[0082] The term "operably linked" refers to functional linkage between a regulatory sequence and a heterologous nucleic acid sequence resulting in expression of the latter. For example, a first nucleic acid sequence is operably linked with a second nucleic acid sequence when the first nucleic acid sequence is placed in a functional relationship with the second nucleic acid sequence. For instance, a promoter is operably linked to a coding sequence if the promoter affects the transcription or expression of the coding sequence. Generally, operably linked DNA sequences are contiguous and, where necessary to join two protein coding regions, in the same reading frame.
[0083] "Parenteral" administration of an immunogenic composition includes, e.g., subcutaneous (s.c.), intravenous (i.v.), intramuscular (i.m.), or intrasternal injection, or infusion techniques.
[0084] As used herein, "plasma cells," refer to a type of white blood cells which can produce and secrete antibodies. Plasma cells are also referred to as plasmocytes, plasmacytes, or effector B cells. In some embodiments, these cells comprise B cell precursors capable of B-cell differentiation or B cells in the early stages of B cell differentiation, expressing a .mu. chain on the cell surface as antigen receptor; B cells in which the transcription process has changed and IgM production has changed from membrane-type IgMs to secreted-type IgMs; mature B cells that have completed class-switching and secrete IgGs, IgAs, and IgMs; and B cells in the final stages of differentiation.
[0085] As used herein, the terms "phospholipase A2 receptor" or "M-type phospholipase A2 receptor" (PLA2R) are used interchangeably and refer to the major target antigen expressed in the kidney glomeruli in primary membranous nephropathy (MN) (Beck et al. 2009 N Engl J Med; 361: 11-21). The anti-PLA2R autoantibodies are predominantly of the IgG4 subclass but subclasses IgG1, IgG2, and IgG3 are also represented. Genetic variants of PLA2R were also shown to be associated with MN (Stanescu et al. 2011 N Engl J Med; 364(7):616-26).
[0086] As used herein, a PLA2R fragment refers to a shortened or truncated PLA2R protein. The polypeptide can have N-terminus or C-terminus truncations and/or also internal deletions. Examples of fragments are fragments comprising the C-type lectin domains ("CTLD") of PLA2R. In one embodiment, a PLA2R fragment includes the external domain of PLA2R, which is the amino acid residues 21-1397 of the human PLA2R (UniProtKB, Q13018) or any shorter portion of the amino acid residues 21-1397.
[0087] The term "polynucleotide," as used herein, is defined as a chain of nucleotides. Furthermore, nucleic acids are polymers of nucleotides. Thus, nucleic acids and polynucleotides, as used herein, are interchangeable. One skilled in the art has the general knowledge that nucleic acids are polynucleotides, which can be hydrolyzed into the monomeric "nucleotides." The monomeric nucleotides can be hydrolyzed into nucleosides. As used herein, polynucleotides include, but are not limited to, all nucleic acid sequences which are obtained by any means available in the art, including, without limitation, recombinant means, i.e., the cloning of nucleic acid sequences from a recombinant library or a cell genome, using ordinary cloning technology and PCR.TM., and the like, and by synthetic means. In some embodiments, a nucleic acid sequence is considered to have at least 95%, 96%, 97%, 98%, or 99% identity or homology to any nucleic acid sequence disclosed herein.
[0088] As used herein, the terms "peptide," "polypeptide," and "protein" are used interchangeably, and refer to a compound comprised of amino acid residues covalently linked by peptide bonds. A protein or peptide must contain at least two amino acids, and no limitation is placed on the maximum number of amino acids that can comprise a protein's or peptide's sequence. Polypeptides include any peptide or protein comprising two or more amino acids joined to each other by peptide bonds. As used herein, the term refers to both short chains, which also commonly are referred to in the art as peptides, oligopeptides and oligomers, for example, and to longer chains, which generally are referred to in the art as proteins, of which there are many types. "Polypeptides" include, for example, biologically active fragments, substantially homologous polypeptides, oligopeptides, homodimers, heterodimers, variants of polypeptides, modified polypeptides, derivatives, analogs, fusion proteins, among others. The polypeptides include natural peptides, recombinant peptides, synthetic peptides, or a combination thereof. In some embodiments, an amino acid sequence is considered to have at 95%, 96%, 97%, 98%, or 99% identity or homology to any amino acid sequence described herein.
[0089] The term "proinflammatory cytokine" refers to a cytokine or factor that promotes inflammation or inflammatory responses. Examples of proinflammatory cytokines include, but are not limited to, chemokines (CCL, CXCL, CX3CL, XCL), interleukins (such as, IL-1, IL-2, IL-3, IL-5, IL-6, IL-7, IL-9, IL10 and IL-15), interferons (IFN.gamma.), and tumor necrosis factors (TNF.alpha. and TNF.beta.).
[0090] The term "promoter," as used herein, is defined as a DNA sequence recognized by the synthetic machinery of the cell, or introduced synthetic machinery, required to initiate the specific transcription of a polynucleotide sequence.
[0091] As used herein, the term "promoter/regulatory sequence" means a nucleic acid sequence which is required for expression of a gene product operably linked to the promoter/regulatory sequence. In some instances, this sequence may be the core promoter sequence and in other instances, this sequence may also include an enhancer sequence and other regulatory elements which are required for expression of the gene product. The promoter/regulatory sequence may, for example, be one which expresses the gene product in a tissue specific manner.
[0092] A "constitutive" promoter is a nucleotide sequence which, when operably linked with a polynucleotide which encodes or specifies a gene product, causes the gene product to be produced in a cell under most or all physiological conditions of the cell.
[0093] An "inducible" promoter is a nucleotide sequence which, when operably linked with a polynucleotide which encodes or specifies a gene product, causes the gene product to be produced in a cell substantially only when an inducer which corresponds to the promoter is present in the cell.
[0094] A "tissue-specific" promoter is a nucleotide sequence which, when operably linked with a polynucleotide encodes or specified by a gene, causes the gene product to be produced in a cell substantially only if the cell is a cell of the tissue type corresponding to the promoter.
[0095] A "signal transduction pathway" refers to the biochemical relationship between a variety of signal transduction molecules that play a role in the transmission of a signal from one portion of a cell to another portion of a cell. The phrase "cell surface receptor" includes molecules and complexes of molecules capable of receiving a signal and transmitting signal across the membrane of a cell.
[0096] "Signaling domain" refers to the portion or region of a molecule that recruits and interacts with specific proteins in response to an activating signal.
[0097] The term "subject" is intended to include living organisms in which an immune response can be elicited (e.g., mammals).
[0098] As used herein, a "substantially purified" cell is a cell that is essentially free of other cell types. A substantially purified cell also refers to a cell which has been separated from other cell types with which it is normally associated in its naturally occurring state. In some instances, a population of substantially purified cells refers to a homogenous population of cells. In other instances, this term refers simply to cells that have been separated from the cells with which they are naturally associated in their natural state. In some embodiments, the cells are cultured in vitro. In other embodiments, the cells are not cultured in vitro.
[0099] The term "therapeutic," as used herein, means a treatment and/or prophylaxis. A therapeutic effect is obtained by suppression, remission, or eradication of a disease state.
[0100] The term "transfected" or "transformed" or "transduced," as used herein, refers to a process by which exogenous nucleic acid is transferred or introduced into the host cell. A "transfected" or "transformed" or "transduced" cell is one which has been transfected, transformed or transduced with exogenous nucleic acid. The cell includes the primary subject cell and its progeny.
[0101] "Transmembrane domain" refers to a portion or a region of a molecule that spans a lipid bilayer membrane.
[0102] The phrase "under transcriptional control" or "operatively linked," as used herein, means that the promoter is in the correct location and orientation in relation to a polynucleotide to control the initiation of transcription by RNA polymerase and expression of the polynucleotide.
[0103] A "vector" is a composition of matter which comprises an isolated nucleic acid and which can be used to deliver the isolated nucleic acid to the interior of a cell. Numerous vectors are known in the art including, but not limited to, linear polynucleotides, polynucleotides associated with ionic or amphiphilic compounds, plasmids, and viruses. Thus, the term "vector" includes an autonomously replicating plasmid or a virus. The term should also be construed to include non-plasmid and non-viral compounds which facilitate transfer of nucleic acid into cells, such as, for example, polylysine compounds, liposomes, and the like. Examples of viral vectors include, but are not limited to, adenoviral vectors, adeno-associated virus vectors, retroviral vectors, lentiviral vectors, and the like.
[0104] By the term "specifically binds," as used herein, is meant an antibody, or a ligand, which recognizes and binds with a cognate binding partner (e.g., a stimulatory and/or costimulatory molecule present on a T cell) protein present in a sample, but which antibody or ligand does not substantially recognize or bind other molecules in the sample.
[0105] By the term "stimulation," is meant a primary response induced by binding of a stimulatory molecule (e.g., a TCR/CD3 complex) with its cognate ligand thereby mediating a signal transduction event, such as, but not limited to, signal transduction via the TCR/CD3 complex. Stimulation can mediate altered expression of certain molecules, such as downregulation of TGF-.beta., and/or reorganization of cytoskeletal structures, and the like.
[0106] A "stimulatory molecule," as the term is used herein, means a molecule on a T cell that specifically binds with a cognate stimulatory ligand present on an antigen presenting cell.
[0107] A "stimulatory ligand," as used herein, means a ligand that when present on an antigen presenting cell (e.g., an aAPC, a dendritic cell, a B-cell, and the like) can specifically bind with a cognate binding partner (referred to herein as a "stimulatory molecule") on a T cell, thereby mediating a primary response by the T cell, including, but not limited to, activation, initiation of an immune response, proliferation, and the like. Stimulatory ligands are well-known in the art and encompass, inter alia, an MHC Class I molecule loaded with a peptide, an anti-CD3 antibody, a superagonist anti-CD28 antibody, and a superagonist anti-CD2 antibody.
[0108] Ranges: throughout this disclosure, various aspects of the invention can be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the invention. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range. For example, description of a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 2.7, 3, 4, 5, 5.3, and 6. This applies regardless of the breadth of the range.
Description
Chimeric Autoantibody Receptor (CAAR)
[0109] The present invention is partly based on the discovery that chimeric autoantibody receptors can be used to target B cells that express autoantibody-based B cell receptors, which after activation and autoantibody secretion, may cause an autoantibody-mediated kidney disease. The invention includes a chimeric autoantibody receptor (CAAR) specific for anti-phospholipase A2 receptor (PLA2R) B cell receptor (BCR), compositions comprising the CAAR, polynucleotides encoding the CAAR, vectors comprising a polynucleotide encoding the CAAR, and recombinant T cells comprising the CAAR. The invention also includes methods of making a genetically modified cell, e.g., a genetically modified T cell, expressing a PLA2R-CAAR wherein the expressed CAAR comprises a phospholipase PLA2R extracellular domain.
[0110] The present invention includes a technology for treating an autoantibody-mediated kidney disease. In particular, technologies that target B cells that ultimately produce the autoantibodies and display the autoantibodies on their cell surfaces, mark these B cells as disease-specific targets for therapeutic intervention. The invention therefore includes a method for efficiently targeting and killing the pathogenic B cells in autoantibody-mediated kidney diseases by targeting the disease-causing B cells using an antigen-specific (e.g., PLA2R) chimeric autoantibody receptor (or CAAR). In one embodiment of the present invention, only specific anti-PLA2R BCR-expressing B cells are killed, leaving intact the beneficial B cells and antibodies that protect from infection.
[0111] In one aspect, the invention includes a polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the polynucleotide encodes a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain.
[0112] Autoantigen Moiety
[0113] In one embodiment, the CAAR of the invention comprises an autoantibody binding domain otherwise referred to as an autoantigen or a fragment thereof. The choice of autoantigen for use in the present invention depends upon the type of autoantibody or BCR being targeted (e.g. anti-PLA2R). For example, the autoantigen may be chosen because it recognizes a BCR or autoantibody on a target cell, such as a BCR-expressing B cell, associated with a particular autoantibody mediated kidney disease state, e.g. a glomerular disease and an primary membranous nephropathy.
[0114] In some instances, it is beneficial that the autoantibody binding domain is derived from the same species in which the CAAR will ultimately be used. For example, for use in humans, it may be beneficial that the autoantibody binding domain of the CAAR comprises a human autoantigen (or fragment thereof) that binds a human BCR or autoantibody.
[0115] In one exemplary embodiment, a genetically engineered chimeric autoantibody receptor includes PLA2R or fragments thereof, which binds an anti-PLA2R BCR, e.g., anti-PLA2R BCR on a B cell in a subject.
[0116] In one embodiment, the CAAR comprises an extracellular domain of PLA2R.
[0117] In some embodiments, the extracellular domain of PLA2R comprises an N-terminal cysteine rich domain (Ricin B type lectin domain), a Fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, a C-type lectin domain 3, a C-type lectin domain 7, or a combination of any of the foregoing.
[0118] In some embodiments, the extracellular domain of PLA2R comprises the immunodominant epitope, N-terminal cysteine rich (CysR) domain (Ricin B type lectin domain), encoded by SEQ ID NO: 1, a Fibronectin type II domain encoded by SEQ ID NO: 3, a C-type lectin domain 1 encoded by SEQ ID NO: 5, and a C-type lectin domain 2 encoded by SEQ ID NO: 7. This extracellular domain of PLA2R is the extracellular domain of the construct designated as construct 4027.CF12 herein and is encoded by SEQ ID NO: 8, wherein the linker between the immunodominant epitope, N-terminal cysteine rich domain and the Fibronectin type II domain is encoded by SEQ ID NO: 2 and the linker between the Fibronectin type II domain and the C-type lectin domain 1 is encoded by SEQ ID NO: 4, and the linker between the C-type lectin domain 1 and the C-type lectin domain 2 is encoded by SEQ ID NO: 6.
[0119] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of construct 4027.CF12 and additionally a C-type lectin domain 3 encoded by SEQ ID NO: 14. This latter extracellular domain of PLA2R is the extracellular domain of the construct designated as construct 4028.CF123 herein and is encoded by SEQ ID NO: 15, wherein the linkers between the immunodominant epitope, N-terminal cysteine rich domain, the Fibronectin type II, the C-type lectin domain 1 and the C-type lectin domain 2 are the same as those listed above herein for construct 4027.CF12 (SEQ ID NOS: 2, 4 and 6) and further wherein the linker between the C-type lectin domain 2 and the C-type lectin domain 3 is encoded by SEQ ID NO: 13.
[0120] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct C, comprising a cysteine rich domain. In certain embodiments, the cysteine rich domain comprises SEQ ID NO: 49, and may be encoded by SEQ ID NO: 47. In certain embodiments, the cysteine rich domain comprises SEQ ID NO: 1.
[0121] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct CF1, comprising a cysteine rich domain, a fibronectin type II domain, and a C-type lectin domain 1. In certain embodiments, the extracellular domain comprises SEQ ID NO: 51, and may be encoded by SEQ ID NO: 50.
[0122] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct CF123, comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, and a C-type lectin domain 3. In certain embodiments, the extracellular domain comprises SEQ ID NO: 53, and may be encoded by SEQ ID NO: 52.
[0123] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct CF1237, comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, a C-type lectin domain 3, and a C-type lectin domain 7. In certain embodiments, the extracellular domain comprises SEQ ID NO: 55, and may be encoded by SEQ ID NO: 54.
[0124] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct CF17, comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, and a C-type lectin domain 7. In certain embodiments, the extracellular domain comprises SEQ ID NO: 57, and may be encoded by SEQ ID NO: 56.
[0125] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct C17, comprising a cysteine rich domain, a C-type lectin domain 1, and a C-type lectin domain 7. In certain embodiments, the extracellular domain comprises SEQ ID NO: 59, and may be encoded by SEQ ID NO: 58.
[0126] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct 4025.C, comprising a cysteine rich domain. In certain embodiments, the extracellular domain comprises SEQ ID NO: 61, and may be encoded by SEQ ID NO: 60.
[0127] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct 4026.CF1, comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1. In certain embodiments, the extracellular domain comprises SEQ ID NO: 40, and may be encoded by SEQ ID NO: 39.
[0128] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct 4027.CF12, comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, and a C-type lectin domain 2. In certain embodiments, the extracellular domain comprises SEQ ID NO: 70, and may be encoded by SEQ ID NO: 8.
[0129] In another embodiment, the extracellular domain of PLA2R comprises the extracellular domain of the construct referred to herein as construct 4028.CF123, comprising a cysteine rich domain, a fibronectin type II domain, a C-type lectin domain 1, a C-type lectin domain 2, and a C-type lectin domain 3. In certain embodiments, the extracellular domain comprises SEQ ID NO: 71, and may be encoded by SEQ ID NO: 15.
[0130] Tolerable variations of the autoantigen or a fragment thereof will be known to those of skill in the art. For example, in some embodiments the autoantigen or a fragment thereof comprises an amino acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to any of the amino acid sequences set forth in SEQ ID NO: 49, 51, 53, 55, 57, 59, 61, 63, 70 or 71. In some embodiments the autoantigen or a fragment thereof is encoded by a nucleic acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least or 99% sequence identity to the nucleic acid sequence set forth in SEQ ID NO: 8, 15, 47, 50, 52, 54, 56, 58, 60, or 62.
[0131] Transmembrane Domain
[0132] In some embodiments, the PLA2R CAAR comprises a transmembrane domain that is fused to the extracellular domain of the PLA2R CAAR. In one embodiment, the PLA2R CAAR comprises a transmembrane domain that naturally is associated with one of the domains in the PLA2R CAAR. In some instances, the transmembrane domain is selected or modified by amino acid substitution to avoid binding to the transmembrane domains of the same or different surface membrane proteins in order to minimize interactions with other members of the receptor complex.
[0133] The transmembrane domain may be derived either from a natural or from a synthetic source. When the source is natural, the domain may be derived from any membrane-bound or transmembrane protein. In one embodiment, the transmembrane domain may be synthetic, in which case it will comprise predominantly hydrophobic residues such as leucine and valine. In one aspect a triplet of phenylalanine, tryptophan and valine will be found at each end of a synthetic transmembrane domain. Optionally, a short oligo- or polypeptide linker, between 2 and 10 amino acids in length may form the linkage between the transmembrane domain and the cytoplasmic signaling domain of the PLA2R CAAR. A glycine-serine doublet provides a particularly suitable linker.
[0134] In some instances, a variety of spacer domains before the transmembrane domain can be employed as well including a hinge (e.g. a CD8 or human Ig (immunoglobulin) hinge), or a glycine-serine (GS) linker.
[0135] Examples of the hinge and/or transmembrane domain include, but are not limited to, a hinge and/or transmembrane domain of an alpha, beta or zeta chain of a T-cell receptor, CD28, CD3 epsilon, CD45, CD4, CD5, CD8, CD9, CD16, CD22, CD33, CD37, CD64, CD80, CD86, CD134, CD137, CD154, killer immunoglobulin-like receptor (KIR), OX40, CD2, CD27, LFA-1 (CD11a, CD18), ICOS (CD278), 4-1BB (CD137), GITR, CD40, BAFFR, HVEM (LIGHTR), SLAMF7, NKp80 (KLRF1), CD160, CD19, IL2R beta, IL2R gamma, IL7R .alpha., ITGA1, VLA1, CD49a, ITGA4, IA4, CD49 D, ITGA6, VLA-6, CD49f, ITGAD, CD11d, ITGAE, CD103, ITGAL, CD11a, LFA-1, ITGAM, CD11b, ITGAX, CD11c, ITGB1, CD29, ITGB2, CD18, LFA-1, ITGB7, TNFR2, DNAM1 (CD226), SLAMF4 (CD244, 2B4), CD84, CD96 (Tactile), CEACAM1, CRTAM, Ly9 (CD229), CD160 (BY55), PSGL1, CD100 (SEMA4D), SLAMF6 (NTB-A, Ly108), SLAM (SLAMF1, CD150, IPO-3), BLAME (SLAMF8), SELPLG (CD162), LTBR, PAG/Cbp, NKp44, NKp30, NKp46, NKG2D, and/or NKG2C.
[0136] In some embodiments, the PLA2R CAAR comprises a transmembrane domain, such as, but not limited to, a CD8 alpha transmembrane domain. In some embodiments, the CD8 alpha transmembrane domain comprises the amino acid sequence IYIWAPLAGTCGVLLLSLVITLYC (SEQ ID NO: 19). In some embodiments, the CD8 alpha transmembrane domain is encoded by the nucleotide sequence
TABLE-US-00001 (SEQ ID NO: 20) ATCTACATCTGGGCGCCCTTGGCCGGGACTTGTGGGGTCCTT CTCCTGTCACTGGTTATCACCCTTTACTGC.
[0137] In some embodiments the transmembrane domain comprises an amino acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 19, or is encoded by a nucleic acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 20.
[0138] In some embodiments, the PLA2R CAAR comprises a hinge domain such as, but not limited to, a CD8 alpha hinge domain. In some embodiments, the hinge domain comprises the amino acid sequence TTTPAPRPPTPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACD (SEQ ID NO: 43). In some embodiments, the hinge domain comprises the amino acid sequence FVPVFLPAKPTTTPAPRPPTPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACD (SEQ ID NO: 44). In some embodiments, the hinge domain is encoded by the nucleotide sequence of SEQ ID NO: 10. In some embodiments, the hinge domain is encoded by the nucleotide sequence ACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCGTCGC AGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGGGGGCGCAGT GCACACGAGGGGGTGGACTTCGCCTGTGAT (SEQ ID NO: 42). In some embodiments, the hinge domain is encoded by the nucleotide sequence TTCGTGCCGGTCTTCCTGCCAGCGAAGCCAACCACGACGCCAGCACCGCGAC CACCAACACCTGCGCCCACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGA GGCGTGCAGACCAGCAGCGGGGGGCAGTGCACACGAGGGGGTGGACTT CGCCTGTGAT (SEQ ID NO: 64). In some embodiments, the PLA2R CAAR comprises a transmembrane domain and a hinge domain.
[0139] In some embodiments the hinge domain comprises an amino acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO:43 or SEQ ID NO: 44, or is encoded by a nucleic acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 42 or SEQ ID NO: 64.
[0140] In some embodiments, the PLA2R CAAR comprises a glycine serine (GS) linker. In some embodiments, the PLA2R CAAR comprises a GS linker and a transmembrane domain. In some embodiments, the GS linker is encoded by the nucleotide sequence: GGTGGCGGAGGTTCTGGAGGTGGAGGTTCC (SEQ ID NO: 68). In some embodiments, the GS linker comprises the amino acid sequence: GGGGSGGGGS (SEQ ID NO: 69). Those of skill in the art would be able to select the appropriate linker sequence, when appropriate for use in a CAAR construct.
[0141] Intracellular Domain of a Costimulatory Molecule
[0142] In some embodiments, the PLA2R CAAR comprises an intracellular domain of a costimulatory molecule. The intracellular domain of a costimulatory molecule of the PLA2R CAAR of the invention is a cytoplasmic domain responsible for the activation of at least one of the normal effector functions of the immune cell in which the PLA2R CAAR has been placed in.
[0143] Effector function of a T cell, for example, may be cytolytic activity or helper activity including the secretion of cytokines. Thus the term "intracellular domain of a costimulatory molecule" refers to the portion of a protein which transduces the effector function signal and directs the cell to perform a specialized function. While the entire intracellular domain of a costimulatory molecule can be employed, in many cases it is not necessary to use the entire domain. To the extent that a truncated portion of the intracellular domain of a costimulatory molecule is used, such truncated portion may be used in place of the intact domain as long as it transduces the effector function signal.
[0144] The intracellular domain of a costimulatory molecule refers to a portion of the CAAR comprising the intracellular domain of a costimulatory molecule. A costimulatory molecule is a cell surface molecule other than an antigen receptor or its ligands that is required for an efficient response of lymphocytes to an antigen. Examples of such molecules include CD27, CD28, 4-1BB (CD137), OX40, CD30, CD40, PD-1, ICOS, lymphocyte function-associated antigen-1 (LFA-1), CD2, CD7, LIGHT, NKG2C, B7-H3, and a ligand that specifically binds with CD83, CD5, ICAM-1, GITR, BAFFR, HVEM (LIGHTR), SLAMF7, NKp80 (KLRF1), CD127, CD160, CD19, CD4, CD8 alpha, CD8 beta, IL2R beta, IL2R gamma, IL7R alpha, ITGA4, VLA1, CD49a, ITGA4, IA4, CD49D, ITGA6, VLA-6, CD49f, ITGAD, CD11d, ITGAE, CD103, ITGAL, CD11a, LFA-1, ITGAM, CD11b, ITGAX, CD11c, ITGB1, CD29, ITGB2, CD18, LFA-1, ITGB7, TNFR2, TRANCE/RANKL, DNAM1 (CD226), SLAMF4 (CD244, 2B4), CD84, CD96 (Tactile), CEACAM1, CRTAM, Ly9 (CD229), CD160 (BY55), PSGL1, CD100 (SEMA4D), CD69, SLAMF6 (NTB-A, Ly108), SLAM (SLAMF1, CD150, IPO-3), BLAME (SLAMF8), SELPLG (CD162), LTBR, LAT, GADS, SLP-76, PAG/Cbp, NKp44, NKp30, NKp46, NKG2D, other co-stimulatory molecules described herein, any derivative, variant, or fragment thereof, any synthetic sequence of a co-stimulatory molecule that has the same functional capability, and any combination thereof. Thus, while the invention is exemplified primarily with 4-1BB (CD137) as the co-stimulatory signaling domains, other costimulatory domains are within the scope of the invention.
[0145] In one embodiment, the nucleic acid sequence of the intracellular domain of a costimulatory molecule encodes an amino acid sequence comprising costimulatory molecule 4-1BB (also known and referred to as CD137) intracellular domain:
TABLE-US-00002 (SEQ ID NO: 21) KRGRKKLLYIFKQPFMRPVQTTQEEDGCSCRFPEEEEGGCEL.
In still another embodiment, the nucleic acid sequence encoding the 4-1BB intracellular domain comprises: AAACGGGGCAGAAAGAAACTCCTGTATATATTCAAACAACCATTTATGAGAC CAGTACAAACTACTCAAGAGGAAGATGGCTGTAGCTGCCGATTTCCAGAAGA AGAAGAAGGAGGATGTGAACTG (SEQ ID NO: 22). In still another embodiment, the nucleic acid sequence encoding the 4-1BB intracellular domain comprises: SEQ ID NO: 66. The human intracellular 4-1BB domain provides co-stimulatory intracellular signaling upon binding to the extracellular autoantigen, such as PLA2R, or a fragment thereof, without the need of its original ligand.
[0146] In some embodiments the intracellular domain of a costimulatory molecule comprises an amino acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 21, or is encoded by a nucleic acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 22 or SEQ ID NO: 66.
[0147] It is well recognized that signals generated through the TCR alone are insufficient for full activation of the T cell and that a secondary or co-stimulatory signal is also required. Thus, T cell activation can be said to be mediated by two distinct classes of cytoplasmic signaling sequence: those that initiate antigen-dependent primary activation through the TCR (primary cytoplasmic signaling sequences) and those that act in an antigen-independent manner to provide a secondary or co-stimulatory signal (secondary cytoplasmic signaling sequences).
[0148] Signaling Domain
[0149] In some embodiments, the PLA2R CAAR comprises a signaling domain. Primary cytoplasmic signaling sequences regulate primary activation of the TCR complex either in a stimulatory manner or in an inhibitory manner. Primary cytoplasmic signaling sequences that act in a stimulatory manner may contain signaling motifs which are known as immunoreceptor tyrosine-based activation motifs or ITAMs.
[0150] Examples of ITAM containing primary signaling sequences that are of particular use in the invention include those derived from TCR zeta, FcR gamma, FcR beta, CD3 gamma, CD3 delta, CD3 epsilon, CD5, CD22, CD79a, CD79b, and CD66d. It is particularly preferred that signaling molecule in the CAAR of the invention comprises a signaling domain derived from CD3-zeta.
[0151] In one embodiment, the signaling domain of the CAAR can be designed to comprise the CD3-zeta signaling domain by itself or combined with any other desired cytoplasmic domain(s) useful in the context of the CAAR of the invention. For example, the signaling domain of the CAAR can comprise a CD3 zeta chain portion and a costimulatory signaling domain.
[0152] In some embodiments, the PLA2R CAAR comprises a CD3-zeta signaling domain by itself or in combination with any other desired cytoplasmic domain(s) useful in the context of the PLA2R CAAR of the invention. For example, the PLA2R CAAR can comprise a CD3 zeta chain portion and an intracellular domain of a costimulatory molecule. In some embodiments, the CD3 zeta chain portion is a human T-cell surface glycoprotein CD3 zeta chain isoform 3 intracellular domain (human CD247). The human intracellular CD3 zeta domain provides stimulatory intracellular signaling upon binding to the extracellular autoantigen, such as PLA2R or a fragment thereof, without HLA restriction.
[0153] In one embodiment, the nucleic acid sequence of the signaling domain comprises a nucleic acid sequence encoding a CD3 zeta signaling domain. In another embodiment, the nucleic acid sequence of the CD3 zeta signaling domain encodes an amino acid sequence comprising RVKFSRSADAPAYKQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRK NPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPR (SEQ ID NO: 23). In another embodiment, the nucleic acid sequence of the CD3 zeta signaling domain encodes an amino acid sequence comprising RVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRK NPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPR (SEQ ID NO: 45). In another embodiment, the nucleic acid sequence encoding the CD3 zeta signaling domain comprises: AGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGGGCCAG AACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTT TGGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGA AGAACCCTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGG AGGCCTACAGTGAGATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGC ACGATGGCCTTTACCAGGGTCTCAGTACAGCCACCAAGGACACCTACGACGC CCTTCACATGCAGGCCCTGCCCCCTCGC (SEQ ID NO: 24). In another embodiment, the nucleic acid sequence encoding the CD3 zeta signaling domain comprises SEQ ID NO: 74.
[0154] In some embodiments, the signaling domain comprises an amino acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 23 or SEQ ID NO: 45, or is encoded by a nucleic acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to SEQ ID NO: 24 or SEQ ID NO: 74.
[0155] Other Domains
[0156] In some embodiments, the PLA2R CAAR and the polynucleotide encoding the PLA2R CAAR comprise a human T cell surface glycoprotein CD8 alpha chain signal peptide. The human CD8 alpha signal peptide is responsible for the translocation of the receptor to the T cell surface.
[0157] In one embodiment, the polynucleotide encoding the PLA2R CAAR comprises a nucleic acid sequence of a peptide linker. In another embodiment, the PLA2R CAAR comprises a peptide linker. In yet another embodiment, the cytoplasmic signaling sequences within the intracellular signaling domain of the PLA2R CAAR can be linked to each other in a random or specified order. Optionally, a short oligo- or polypeptide linker, for example, between 2 and 10 amino acids in length may form the linkage. A glycine-serine doublet is a particularly suitable linker.
[0158] In some embodiments, the CAAR comprises a transmembrane domain and/or a cytoplasmic (intracellular) domain from a killer immunoglobulin-like receptor (KIR) family protein. The KIR gene family has at least 15 gene loci (KIR2DL1, KIR2DL2/L3, KIR2DL4, KIR2DL5A, KIR2DL5B, KIR2DS1, KIR2DS2, KIR2DS3, KIR2DS4, KIR2DS5, KIR3DL1/S1, KIR3DL2, KIR3DL3) and two pseudogenes (KIR2DP1 and KIR3DP1) encoded within a 100-200 Kb region of the Leukocyte Receptor Complex (LRC) located on chromosome 19 (19q13.4). The LRC constitutes a large, 1 Mb, and dense cluster of rapidly evolving immune genes which contains genes encoding other cell surface molecules with distinctive Ig-like extra-cellular domains. In addition, the extended LRC contains genes encoding the transmembrane adaptor molecules DAP10 and DAP12. Thus, a cell comprising the CAAR of the invention comprising a KIR transmembrane domain and/or cytoplasmic domain may also comprise a polynucleotide encoding DAP10 or DAP12. In certain embodiments, the KIR is KIRS2 or KIR2DS2
[0159] In certain embodiments, the CAAR is encoded by a nucleic acid sequence selected from the group consisting of SEQ ID NOs: 11, 17, 25, 27, 29, 31, 33, 35, 37, and 39. In certain embodiments, the CAAR comprises an amino acid sequence selected from the group consisting of SEQ ID NOs: 12, 18, 26, 28, 30, 32, 34, 36, 38, 40, and 42.
[0160] Tolerable variations of the CAAR sequences will be known to those of skill in the art. For example, in some embodiments the CAAR comprises an amino acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to any of the amino acid sequences set forth in SEQ ID NOs: 12, and 18, 26, 28, 30, 32, 34, 36, 38, or 40. In some embodiments the CAAR is encoded by a nucleic acid sequence that has at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% sequence identity to the nucleic acid sequence set forth in SEQ ID NO: 11, 17, 25, 27, 29, 31, 33, 35, 37, or 39.
Vector Comprising the PLA2R CAAR
[0161] In some embodiments, a 3.sup.rd generation self-inactivating lentiviral vector plasmid can be used in which the expression of the CAR is regulated by the human elongation factor 1 alpha promoter. This results in stable (permanent) expression of the CAR in the host T cell. As an alternative approach, the encoding mRNA can be electroporated into the host cell, which would achieve the same therapeutic effect as the virally transduced T cells, but would not be permanent because the mRNA would dilute out with cell division.
[0162] In one aspect, the invention includes a vector comprising a polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the polynucleotide comprises an extracellular domain comprising a human PLA2R autoantigen or fragment thereof, and optionally, a transmembrane domain, and/or an intracellular signaling domain. In one embodiment, the vector comprises any of the nucleic acid sequences encoding the CAAR as described herein. In another embodiment, the vector comprises a plasmid vector, viral vector, retrotransposon (e.g. piggyback, sleeping beauty), site directed insertion vector (e.g. CRISPR, Zinc finger nucleases, TALEN), or suicide expression vector, or other known vector in the art.
[0163] All constructs disclosed herein can be used with 3rd generation lentiviral vector plasmids, other viral vectors, or RNA approved for use in human cells. In one embodiment, the vector is a viral vector, such as a lentiviral vector. In another embodiment, the vector is a RNA vector.
[0164] The expression of the PLA2R CAAR can be verified by sequencing. Expression of the full length CAAR protein may be verified using immunoblot, immunohistochemistry, flow cytometry or other technology well known and available in the art.
[0165] The present invention also provides a vector in which DNA encoding the CAAR of the present invention is inserted. Vectors, including those derived from retroviruses such as lentivirus, are suitable tools to achieve long-term gene transfer since they allow long-term, stable integration of a transgene and its propagation in daughter cells. Lentiviral vectors have the added advantage over vectors derived from onco-retroviruses, such as murine leukemia viruses, in that they can transduce non-proliferating cells, such as hepatocytes. They also have the added advantage of resulting in low immunogenicity in the subject into which they are introduced.
[0166] In brief summary, the expression of natural or synthetic polynucleotides encoding CAARs is typically achieved by operably linking a nucleic acid encoding the CAAR polypeptide or portions thereof to a promoter (e.g. EF1alpha promoter), and incorporating the construct into an expression vector. The vector is one generally capable of replication in a mammalian cell, and/or also capable of integration into the cellular genome of the mammal. Typical vectors contain transcription and translation terminators, initiation sequences, and promoters useful for regulation of the expression of the desired nucleic acid sequence.
[0167] The nucleic acid can be cloned into any number of different types of vectors. For example, the nucleic acid can be cloned into a vector including, but not limited to a plasmid, a phagemid, a phage derivative, an animal virus, and a cosmid. Vectors of particular interest include expression vectors, replication vectors, probe generation vectors, and sequencing vectors.
[0168] The expression vector may be provided to a cell in the form of a viral vector. Viral vector technology is well known in the art and is described, for example, in Sambrook et al., 2012, MOLECULAR CLONING: A LABORATORY MANUAL, volumes 1-4, Cold Spring Harbor Press, NY), and in other virology and molecular biology manuals. Viruses, which are useful as vectors include, but are not limited to, retroviruses, adenoviruses, adeno-associated viruses, herpes viruses, and lentiviruses. In general, a suitable vector contains an origin of replication functional in at least one organism, a promoter sequence, convenient restriction endonuclease sites, and one or more selectable markers, (e.g., WO 01/96584; WO 01/29058; and U.S. Pat. No. 6,326,193).
[0169] Additional promoter elements, e.g., enhancers, regulate the frequency of transcriptional initiation. Typically, these are located in the region 30-110 bp upstream of the start site, although a number of promoters have recently been shown to contain functional elements downstream of the start site as well. The spacing between promoter elements frequently is flexible, so that promoter function is preserved when elements are inverted or moved relative to one another. In the thymidine kinase (tk) promoter, the spacing between promoter elements can be increased to 50 bp apart before activity begins to decline. Depending on the promoter, it appears that individual elements can function either cooperatively or independently to activate transcription.
[0170] An example of a promoter is the immediate early cytomegalovirus (CMV) promoter sequence. This promoter sequence is a strong constitutive promoter sequence capable of driving high levels of expression of any polynucleotide sequence operatively linked thereto. However, other constitutive promoter sequences may also be used, including, but not limited to the simian virus 40 (SV40) early promoter, mouse mammary tumor virus (MMTV), human immunodeficiency virus (HIV) long terminal repeat (LTR) promoter, MoMuLV promoter, an avian leukemia virus promoter, an Epstein-Barr virus immediate early promoter, a Rous sarcoma virus promoter, the elongation factor-1.alpha. promoter, as well as human gene promoters such as, but not limited to, the actin promoter, the myosin promoter, the hemoglobin promoter, when appropriate for use in a CAAR construct.
[0171] Further, the invention should not be limited to the use of constitutive promoters. Inducible promoters are also contemplated as part of the invention. The use of an inducible promoter provides a molecular switch capable of turning on expression of the polynucleotide sequence, which it is operatively linked when such expression is desired, or turning off the expression when expression is not desired. Examples of inducible promoters include, but are not limited to a metallothionine promoter, a glucocorticoid promoter, a progesterone promoter, and a tetracycline promoter. In some embodiments, an inducible promoter is activated in response to a extracellular ligand. For example, in some embodiments, the inducible promoter is activated (and the expression of the CAAR is regulated) by an extracellular ligand binding to a synthetic receptor. For example, in some embodiments, a synthetic receptor, e.g., a synthetic Notch receptor (i.e., "synNotch") may be employed as a binding-triggered transcriptional switch that, when bound to its ligand, activates a promoter to which a nucleic acid sequence encoding the CAAR is operably linked. Accordingly, as a non-limiting example, such systems may require the presence of a ligand (e.g., to which the synNotch binds) for the immune cell to be responsive to a BCR or autoantibody (e.g., to which the CAAR binds). The requirement of particular combinations to generate certain signaling outputs in molecular circuits results in a logic gate. See, for example, Roybal et al., 2016 Cell 164(4):770-9.
[0172] Examples of other systems for expressing or regulating expression of a chimeric receptor include those described in Wu et al. (2015) Science 350: aab4077; Fedorov et al. (2014) Cancer Journal 20:160-165; Kloss et al. (2013) Nature Biotechnology 31: 71-75; Sakemura et al. (2016) Cancer Immunol. Res. 4:658-668; Hill et al. (2018) Nature Chemical Biology 14:112-117; Di Stasi et al. (2011) N. Engl. J. Med. 365:1673-1683; Budde et al. (2013) PLoS One 8: e82742; Wei et al. (2012) Nature 488: 384-388; Ma et al. (2016) Proc. Natl. Acad. Sci. USA 113: E450-458; Rodgers et al. (2016) Proc. Natl. Acad. Sci. USA 113: E459-468; Kudo et al. (2014) Cancer Res. 74: 93-103, and Chen et al. (2010) Proc. Natl. Acad. Sci. USA 107, 8531-8536.
[0173] In order to assess the expression of a CAAR polypeptide or portions thereof, the expression vector to be introduced into a cell can also contain either a selectable marker gene or a reporter gene or both to facilitate identification and selection of expressing cells from the population of cells sought to be transfected or infected through viral vectors. In other aspects, the selectable marker may be carried on a separate piece of DNA and used in a co-transfection procedure. Both selectable markers and reporter genes may be flanked with appropriate regulatory sequences to enable expression in the host cells. Useful selectable markers include, for example, antibiotic-resistance genes, such as neo and the like.
[0174] Reporter genes are used for identifying potentially transfected cells and for evaluating the functionality of regulatory sequences. In general, a reporter gene is a gene that is not present in or expressed by the recipient organism or tissue and that encodes a polypeptide whose expression is manifested by some easily detectable property, e.g., enzymatic activity. Expression of the reporter gene is assessed at a suitable time after the DNA has been introduced into the recipient cells. Suitable reporter genes may include genes encoding luciferase, beta-galactosidase, chloramphenicol acetyl transferase, secreted alkaline phosphatase, or the green fluorescent protein gene (e.g., Ui-Tei et al., 2000 FEBS Letters 479: 79-82). Suitable expression systems are well known and may be prepared using known techniques or obtained commercially. In general, the construct with the minimal 5' flanking region showing the highest level of expression of reporter gene is identified as the promoter. Such promoter regions may be linked to a reporter gene and used to evaluate agents for the ability to modulate promoter-driven transcription.
[0175] Methods of introducing and expressing genes into a cell are known in the art. In the context of an expression vector, the vector can be readily introduced into a host cell, e.g., mammalian, bacterial, yeast, or insect cell by any method in the art. For example, the expression vector can be transferred into a host cell by physical, chemical, or biological means.
[0176] Physical methods for introducing a polynucleotide into a host cell include calcium phosphate precipitation, lipofection, particle bombardment, microinjection, electroporation, and the like. Methods for producing cells comprising vectors and/or exogenous nucleic acids are well-known in the art. See, for example, Sambrook et al., 2012, MOLECULAR CLONING: A LABORATORY MANUAL, volumes 1-4, Cold Spring Harbor Press, NY).
[0177] Biological methods for introducing a polynucleotide of interest into a host cell include the use of DNA and RNA vectors. RNA vectors include vectors having a RNA promoter and/other relevant domains for production of a RNA transcript. Viral vectors, and especially retroviral vectors, have become the most widely used method for inserting genes into mammalian, e.g., human cells. Other viral vectors may be derived from lentivirus, poxviruses, herpes simplex virus, adenoviruses and adeno-associated viruses, and the like. See, for example, U.S. Pat. Nos. 5,350,674 and 5,585,362.
[0178] Chemical means for introducing a polynucleotide into a host cell include colloidal dispersion systems, such as macromolecule complexes, nanocapsules, microspheres, beads, and lipid-based systems including oil-in-water emulsions, micelles, mixed micelles, and liposomes. An exemplary colloidal system for use as a delivery vehicle in vitro and in vivo is a liposome (e.g., an artificial membrane vesicle).
[0179] In the case where a non-viral delivery system is utilized, an exemplary delivery vehicle is a liposome. The use of lipid formulations is contemplated for the introduction of the nucleic acids into a host cell (in vitro, ex vivo or in vivo). In another aspect, the nucleic acid may be associated with a lipid. The nucleic acid associated with a lipid may be encapsulated in the aqueous interior of a liposome, interspersed within the lipid bilayer of a liposome, attached to a liposome via a linking molecule that is associated with both the liposome and the oligonucleotide, entrapped in a liposome, complexed with a liposome, dispersed in a solution containing a lipid, mixed with a lipid, combined with a lipid, contained as a suspension in a lipid, contained or complexed with a micelle, or otherwise associated with a lipid. Lipid, lipid/DNA or lipid/expression vector associated compositions are not limited to any particular structure in solution. For example, they may be present in a bilayer structure, as micelles, or with a "collapsed" structure. They may also simply be interspersed in a solution, possibly forming aggregates that are not uniform in size or shape. Lipids are fatty substances, which may be naturally occurring or synthetic lipids. For example, lipids include the fatty droplets that naturally occur in the cytoplasm as well as the class of compounds which contain long-chain aliphatic hydrocarbons and their derivatives, such as fatty acids, alcohols, amines, amino alcohols, and aldehydes.
[0180] Lipids suitable for use can be obtained from commercial sources. For example, dimyristyl phosphatidylcholine ("DMPC") can be obtained from Sigma, St. Louis, Mo.; dicetyl phosphate ("DCP") can be obtained from K & K Laboratories (Plainview, N.Y.); cholesterol ("Choi") can be obtained from Calbiochem-Behring; dimyristyl phosphatidylglycerol ("DMPG") and other lipids may be obtained from Avanti Polar Lipids, Inc. (Birmingham, Ala.). Stock solutions of lipids in chloroform or chloroform/methanol can be stored at about -20.degree. C. Chloroform is used as the only solvent since it is more readily evaporated than methanol. "Liposome" is a generic term encompassing a variety of single and multilamellar lipid vehicles formed by the generation of enclosed lipid bilayers or aggregates. Liposomes can be characterized as having vesicular structures with a phospholipid bilayer membrane and an inner aqueous medium. Multilamellar liposomes have multiple lipid layers separated by aqueous medium. They form spontaneously when phospholipids are suspended in an excess of aqueous solution. The lipid components undergo self-rearrangement before the formation of closed structures and entrap water and dissolved solutes between the lipid bilayers (Ghosh et al., 1991 Glycobiology 5: 505-10). However, compositions that have different structures in solution than the normal vesicular structure are also encompassed. For example, the lipids may assume a micellar structure or merely exist as nonuniform aggregates of lipid molecules. Also contemplated are lipofectamine-nucleic acid complexes.
[0181] Any domains and/or fragments of the CAAR, vector, and the promoter may be synthesized gene fragments amplified by PCR or any other means known in the art.
Cells Comprising the CAAR
[0182] In another aspect, the invention includes a genetically modified cell comprising the PLA2R chimeric autoantibody receptor (CAAR) disclosed herein.
[0183] In another embodiment, the genetically modified cell expresses the PLA2R CAAR. In this embodiment, the cell has high affinity for PLA2R autoantibody-based B cell receptors (BCRs) on B cells or rarely, on B cells that have differentiated into plasma cells that have not yet downregulated their BCR. As a result, the genetically modified cell can induce direct killing of anti-PLA2R B cells or indirect killing of plasma cells expressing PLA2R autoantibodies. In yet another embodiment, the genetically modified cell has low affinity for antibodies bound to an Fc receptor.
[0184] In one embodiment, the genetically modified cell is a T cell, such as a helper T cell, a cytotoxic T cell, a memory T cell, regulatory T cell, gamma delta T cell, a natural killer cell, cytokine induced killer cell, a cell line thereof, a T memory stem cell, or other T effector cell. It is also useful for the T cell to have limited toxicity toward healthy cells and specificity to cells expressing autoantibodies. Such specificity prevents or reduces off-target toxicity that is prevalent in current therapies that are not specific for autoantibodies. In one embodiment, the T cell has limited toxicity toward healthy cells. In one embodiment the T cell is an autologous cell. In another embodiment, the T cell is an allogeneic cell.
[0185] In some embodiments, the invention includes genetically modified immune cells derived from pluripotent stem cells that were differentiated in vitro. In other embodiments, the invention includes T cells, such as primary cells, expanded T cells derived from primary T cells, T cells derived from stem cells differentiated in vitro, T cell lines such as Jurkat cells, other sources of T cells, combinations thereof, and other effector cells. For example, a transduced Jurkat cell line with a NFAT response element followed by GFP can be used to detect and isolate PLA2R specific B cells and to clone the PLA2R specific antibody repertoire in a comprehensive and unbiased fashion. The interacting B and Jurkat cells can be detected as GFP positive doublets or multimers and sorted by flow cytometry. Expression cloning of the B cell receptor encoding genes will provide further information on how autoimmunity and autoantibodies in autoantibody-mediated kidney diseases, such as glomerular disease and primary membranous nephropathy develop.
[0186] The functional ability of CAARs to specifically bind to autoantibodies and sera, for example, primary membranous nephropathy sera, can be assessed in a Jurkat reporter cell line, which depends on activation of the CAAR by binding to plate-bound autoantibody (in response to which the activated cells fluoresce green due to an NFAT-GFP reporter construct contained therein). Such methods are useful and reliable qualitative measures for functional binding ability. The proper processing of the autoantigen on the cell surface is also important and can be measured using monoclonal antibodies. Furthermore, truncations of PLA2R based on major disease epitopes are also useful and included herein. Versions using a different length hinge region or GS linker are also useful. With regard to safety, preventing or reducing possible homophilic and heterophilic interactions and activation (e.g. PLA2R-PLA2R) between the transduced cells or toward podocytes is preferred.
[0187] Further assessment of efficacy and safety of the CAAR can be performed, for example, as follows:
[0188] Constructs can be transiently transfected into human cells, such as 293T/17. The surface expression can be detected with monoclonal antibodies (either IgG or ScFv) or serum antibodies from PLA2R MN patients specific for the abovementioned extracellular domains, the linker between the domains, or other structure included in the CAAR. Binding can be verified with specific secondary antibodies and quantified by flow cytometry.
[0189] Production of membrane expressed constructs of human anti-PLA2R antibodies of any isotype can serve as target cells for testing the different PLA2R-CAARs. Additional target cell lines can be produced as needed by expression of human monoclonal antibodies on the surface of cell lines (e.g. Nalm6 or K562 cells).
Autoimmune Diseases
[0190] The present invention also provides methods for preventing, treating and/or managing a disorder or autoimmune disease associated with autoantibody-expressing cells in the context of an autoantibody-mediated kidney disease. The methods comprise administering to a subject in need thereof a genetically modified T cell comprising the CAAR of the invention that binds to the autoantibody-expressing cell. In one aspect, the subject is a human. Non-limiting examples of an autoantibody-mediated kidney disease include to glomerular disease and primary membranous nephropathy.
[0191] In the methods of treatment, T cells isolated from a subject can be modified to express the appropriate CAAR, expanded ex vivo and then reinfused into the same subject (e.g., the T cells are autologous T cells). In some embodiments, the T cells are reinfused into a different subject than the original T cells' donor (e.g., the T cells are allogeneic T cells). The modified T cells recognize target cells, such as anti-PLA2R B cells or PLA2R autoantibody producing B cells or plasma cells, and become activated, resulting in killing of the autoimmune target cells.
[0192] Relapse may also occur in patients with an autoimmune disease, for example in primary membranous nephropathy patients. In patients treated with rituximab, the relapse may be mediated by persistence of the same autoantibody B cell clones, whereas remission is associated with disappearance of these clones. By infusing PLA2R CAAR T cells, the autoimmune cells are depleted to induce long-term remission, possibly due to the longevity of the PLA2R CAAR T cells and/or autoantigen-reactive clones do not re-appear.
[0193] To monitor PLA2R CAAR-expressing cells in vitro, in situ, or in vivo, PLA2R CAAR cells can further express a detectable marker. When the PLA2R CAAR binds the target, the detectable marker is activated and expressed, which can be detected by assays known in the art, such as flow cytometry. In one embodiment, the PLA2R CAAR includes a NFAT response element and a detectable marker, such as a green fluorescent protein (GFP), to detect and quantify PLA2R CAAR expressing cells.
Sources of T Cells
[0194] Prior to expansion and genetic modification, T cells (e.g., autologous or allogeneic T cells) are obtained from a subject. Examples of subjects include humans, dogs, cats, mice, rats, and transgenic species thereof. T cells can be obtained from a number of sources, including skin, peripheral blood mononuclear cells, bone marrow, lymph node tissue, cord blood, thymus tissue, tissue from a site of infection, ascites, pleural effusion, spleen tissue, and tumors. In certain embodiments of the present invention, any number of T cell lines available in the art, may be used. In certain embodiments of the present invention, T cells can be obtained from a unit of blood collected from a subject using any number of techniques known to the skilled artisan, such as Ficoll.TM. separation. In one preferred embodiment, cells from the circulating blood of an individual are obtained by apheresis. The apheresis product typically contains lymphocytes, including T cells, monocytes, granulocytes, B cells, other nucleated white blood cells, red blood cells, and platelets. In one embodiment, the cells collected by apheresis may be washed to remove the plasma fraction and to place the cells in an appropriate buffer or media for subsequent processing steps. In one embodiment of the invention, the cells are washed with phosphate buffered saline (PBS). In an alternative embodiment, the wash solution lacks calcium and may lack magnesium or may lack many if not all divalent cations. Again, surprisingly, initial activation steps in the absence of calcium lead to magnified activation. As those of ordinary skill in the art would readily appreciate a washing step may be accomplished by methods known to those in the art, such as by using a semi-automated "flow-through" centrifuge (for example, the Cobe 2991 cell processor, the Baxter CytoMate, or the Haemonetics Cell Saver 5) according to the manufacturer's instructions. After washing, the cells may be resuspended in a variety of biocompatible buffers, such as, for example, Ca-free, Mg-free PBS, PlasmaLyte A, or other saline solution with or without buffer. Alternatively, the undesirable components of the apheresis sample may be removed and the cells directly resuspended in culture media.
[0195] In another embodiment, T cells are isolated from peripheral blood lymphocytes by lysing the red blood cells and depleting the monocytes, for example, by centrifugation through a PERCOLL.TM. gradient or by counterflow centrifugal elutriation. A specific subpopulation of T cells, such as CD3.sup.+, CD28.sup.+, CD4.sup.+, CD8.sup.+, CD45RA.sup.+, and CD45RO.sup.+T cells, can be further isolated by positive or negative selection techniques. For example, in one embodiment, T cells are isolated by incubation with anti-CD3/anti-CD28 (i.e., 3.times.28)-conjugated beads, such as DYNABEADS.RTM. M-450 CD3/CD28 T, for a time period sufficient for positive selection of the desired T cells. In one embodiment, the time period is about 30 minutes. In a further embodiment, the time period ranges from 30 minutes to 36 hours or longer and all integer values there between. In a further embodiment, the time period is at least 1, 2, 3, 4, 5, or 6 hours. In yet another preferred embodiment, the time period is 10 to 24 hours. In one preferred embodiment, the incubation time period is 24 hours. For isolation of T cells from patients with leukemia, use of longer incubation times, such as 24 hours, can increase cell yield. Longer incubation times may be used to isolate T cells in any situation where there are few T cells as compared to other cell types, such in isolating tumor infiltrating lymphocytes (TIL) from tumor tissue or from immunocompromised individuals. Further, use of longer incubation times can increase the efficiency of capture of CD8+ T cells. Thus, by simply shortening or lengthening the time T cells are allowed to bind to the CD3/CD28 beads and/or by increasing or decreasing the ratio of beads to T cells (as described further herein), subpopulations of T cells can be preferentially selected for or against at culture initiation or at other time points during the process. Additionally, by increasing or decreasing the ratio of anti-CD3 and/or anti-CD28 antibodies on the beads or other surface, subpopulations of T cells can be preferentially selected for or against at culture initiation or at other desired time points. The skilled artisan would recognize that multiple rounds of selection can also be used in the context of this invention. In certain embodiments, it may be desirable to perform the selection procedure and use the "unselected" cells in the activation and expansion process. "Unselected" cells can also be subjected to further rounds of selection.
[0196] Enrichment of a T cell population by negative selection can be accomplished with a combination of antibodies directed to surface markers unique to the negatively selected cells. One method is cell sorting and/or selection via negative magnetic immunoadherence or flow cytometry that uses a cocktail of monoclonal antibodies directed to cell surface markers present on the cells negatively selected. For example, to enrich for CD4.sup.+ cells by negative selection, a monoclonal antibody cocktail typically includes antibodies to CD14, CD20, CD11b, CD16, HLA-DR, and CD8. In certain embodiments, it may be desirable to enrich for or positively select for regulatory T cells which typically express CD4.sup.+, CD25.sup.+, CD62L.sup.+, GITR.sup.+, and FoxP3.sup.+. Alternatively, in certain embodiments, T regulatory cells are depleted by anti-CD25 conjugated beads or other similar method of selection. In other embodiments, subpopulation of T cells, such as, but not limited to, cells positive or expressing high levels of one or more surface markers e.g. CD28+, CD8+, CCR7+, CD27+, CD127+, CD45RA+, and/or CD45RO+ T cells, can be isolated by positive or negative selection techniques.
[0197] For isolation of a desired population of cells by positive or negative selection, the concentration of cells and surface (e.g., particles such as beads) can be varied. In certain embodiments, it may be desirable to significantly decrease the volume in which beads and cells are mixed together (i.e., increase the concentration of cells), to ensure maximum contact of cells and beads. For example, in one embodiment, a concentration of 2 billion cells/ml is used. In one embodiment, a concentration of 1 billion cells/ml is used. In a further embodiment, greater than 100 million cells/ml is used. In a further embodiment, a concentration of cells of 10, 15, 20, 25, 30, 35, 40, 45, or 50 million cells/ml is used. In yet another embodiment, a concentration of cells from 75, 80, 85, 90, 95, or 100 million cells/ml is used. In further embodiments, concentrations of 125 or 150 million cells/ml can be used. Using high concentrations can result in increased cell yield, cell activation, and cell expansion. Further, use of high cell concentrations allows more efficient capture of cells that may weakly express target antigens of interest, such as CD28-negative T cells, or from samples where there are many tumor cells present (i.e., leukemic blood, tumor tissue, etc.). Such populations of cells may have therapeutic value and would be desirable to obtain. For example, using high concentration of cells allows more efficient selection of CD8.sup.+ T cells that normally have weaker CD28 expression.
[0198] In a related embodiment, it may be desirable to use lower concentrations of cells. By significantly diluting the mixture of T cells and surface (e.g., particles such as beads), interactions between the particles and cells is minimized. This selects for cells that express high amounts of desired antigens to be bound to the particles. For example, CD4.sup.+ T cells express higher levels of CD28 and are more efficiently captured than CD8.sup.+ T cells in dilute concentrations. In one embodiment, the concentration of cells used is 5.times.10.sup.6/ml. In other embodiments, the concentration used can be from about 1.times.10.sup.5/ml to 1.times.10.sup.6/ml, and any integer value in between.
[0199] In other embodiments, the cells may be incubated on a rotator for varying lengths of time at varying speeds at either 2-10.degree. C. or at room temperature.
[0200] T cells for stimulation can also be frozen after a washing step. Wishing not to be bound by theory, the freeze and subsequent thaw step provides a more uniform product by removing granulocytes and to some extent monocytes in the cell population. After the washing step that removes plasma and platelets, the cells may be suspended in a freezing solution. While many freezing solutions and parameters are known in the art and will be useful in this context, one method involves using PBS containing 20% DMSO and 8% human serum albumin, or culture media containing 10% Dextran 40 and 5% Dextrose, 20% Human Serum Albumin and 7.5% DMSO, or 31.25% Plasmalyte-A, 31.25% Dextrose 5%, 0.45% NaCl, 10% Dextran 40 and 5% Dextrose, 20% Human Serum Albumin, and 7.5% DMSO or other suitable cell freezing media containing for example, Hespan and PlasmaLyte A, the cells then are frozen to -80.degree. C. at a rate of 1.degree. per minute and stored in the vapor phase of a liquid nitrogen storage tank. Other methods of controlled freezing may be used as well as uncontrolled freezing immediately at -20.degree. C. or in liquid nitrogen.
[0201] In certain embodiments, cryopreserved cells are thawed and washed as described herein and allowed to rest for one hour at room temperature prior to activation using the methods of the present invention.
[0202] Also contemplated in the context of the invention is the collection of blood samples or apheresis product from a subject at a time period prior to when the expanded cells as described herein might be needed. As such, the source of the cells to be expanded can be collected at any time point necessary, and desired cells, such as T cells, isolated and frozen for later use in T cell therapy for any number of diseases or conditions that would benefit from T cell therapy, such as those described herein. In one embodiment, a blood sample or an apheresis is taken from a generally healthy subject. In certain embodiments, a blood sample or an apheresis is taken from a generally healthy subject who is at risk of developing a disease, but who has not yet developed a disease, and the cells of interest are isolated and frozen for later use. In certain embodiments, the T cells may be expanded, frozen, and used at a later time. In certain embodiments, samples are collected from a patient shortly after diagnosis of a particular disease as described herein but prior to any treatments. In a further embodiment, the cells are isolated from a blood sample or an apheresis from a subject prior to any number of relevant treatment modalities, including but not limited to treatment with agents such as natalizumab, efalizumab, antiviral agents, chemotherapy, radiation, immunosuppressive agents, such as cyclosporin, azathioprine, methotrexate, mycophenolate, and FK506, antibodies, or other immunoablative agents such as CAMPATH, anti-CD3 antibodies, cytoxan, fludarabine, cyclosporin, FK506, rapamycin, mycophenolic acid, steroids, FR901228, and irradiation. These drugs may, for example, inhibit either the calcium dependent phosphatase calcineurin (cyclosporine and FK506) or inhibit the p70S6 kinase that is important for growth factor induced signaling (rapamycin). (Liu et al., Cell 66:807-815, 1991; Henderson et al., Immun. 73:316-321, 1991; Bierer et al., Curr. Opin. Immun. 5:763-773, 1993). In another embodiment, the cells are isolated prior to and can be frozen for later use for treatment following B-cell ablative therapy, e.g., Rituxan.
[0203] In a further embodiment of the present invention, T cells are obtained from a patient directly following treatment. In this regard, it has been observed that following certain treatments, in particular treatments with drugs that damage the immune system, shortly after treatment during the period when patients would normally be recovering from the treatment, the quality of T cells obtained may be optimal or improved for their ability to expand ex vivo. Likewise, following ex vivo manipulation using the methods described herein, these cells may be in a preferred state for enhanced engraftment and in vivo expansion. Thus, it is contemplated within the context of the present invention to collect blood cells, including T cells, dendritic cells, or other cells of the hematopoietic lineage, during this recovery phase. Further, in certain embodiments, mobilization (for example, mobilization with GM-CSF) and conditioning regimens can be used to create a condition in a subject wherein repopulation, recirculation, regeneration, and/or expansion of particular cell types is favored, especially during a defined window of time following therapy. Illustrative cell types include T cells, B cells, dendritic cells, and other cells of the immune system.
Activation and Expansion of T Cells
[0204] T cells are activated and expanded generally using methods as described, for example, in U.S. Pat. Nos. 6,352,694; 6,534,055; 6,905,680; 6,692,964; 5,858,358; 6,887,466; 6,905,681; 7,144,575; 7,067,318; 7,172,869; 7,232,566; 7,175,843; 5,883,223; 6,905,874; 6,797,514; 6,867,041; and U.S. Patent Application Publication No. 20060121005.
[0205] Generally, the T cells of the invention are expanded by contact with a surface having attached thereto an agent that stimulates a CD3/TCR complex associated signal and a ligand that stimulates a co-stimulatory molecule on the surface of the T cells. In particular, T cell populations may be stimulated as described herein, such as by contact with an anti-CD3 antibody, or antigen-binding fragment thereof, or an anti-CD2 antibody immobilized on a surface, or by contact with a protein kinase C activator (e.g., bryostatin) in conjunction with a calcium ionophore. For co-stimulation of an accessory molecule on the surface of the T cells, a ligand that binds the accessory molecule is used. For example, a population of T cells can be contacted with an anti-CD3 antibody and an anti-CD28 antibody, under conditions appropriate for stimulating proliferation of the T cells. To stimulate proliferation of either CD4.sup.+ T cells or CD8.sup.+ T cells, an anti-CD3 antibody and an anti-CD28 antibody. Examples of an anti-CD28 antibody include 9.3, B-T3, XR-CD28 (Diaclone, Besancon, France) can be used as can other methods commonly known in the art (Berg et al., Transplant Proc. 30(8):3975-3977, 1998; Haanen et al., J. Exp. Med. 190(9):13191328, 1999; Garland et al., J. Immunol Meth. 227(1-2):53-63, 1999).
[0206] In certain embodiments, the primary stimulatory signal and the co-stimulatory signal for the T cell may be provided by different protocols. For example, the agents providing each signal may be in solution or coupled to a surface. When coupled to a surface, the agents may be coupled to the same surface (i.e., in "cis" formation) or to separate surfaces (i.e., in "trans" formation). Alternatively, one agent may be coupled to a surface and the other agent in solution. In one embodiment, the agent providing the co-stimulatory signal is bound to a cell surface and the agent providing the primary activation signal is in solution or coupled to a surface. In certain embodiments, both agents can be in solution. In another embodiment, the agents may be in soluble form, and then cross-linked to a surface, such as a cell expressing Fc receptors or an antibody or other binding agent which will bind to the agents. In this regard, see for example, U.S. Patent Application Publication Nos. 20040101519 and 20060034810 for artificial antigen presenting cells (aAPCs) that are contemplated for use in activating and expanding T cells in the present invention.
[0207] In one embodiment, the two agents are immobilized on beads, either on the same bead, i.e., "cis," or to separate beads, i.e., "trans." By way of example, the agent providing the primary activation signal is an anti-CD3 antibody or an antigen-binding fragment thereof and the agent providing the co-stimulatory signal is an anti-CD28 antibody or antigen-binding fragment thereof; and both agents are co-immobilized to the same bead in equivalent molecular amounts. In one embodiment, a 1:1 ratio of each antibody bound to the beads for CD4.sup.+ T cell expansion and T cell growth is used. In certain aspects of the present invention, a ratio of anti CD3:CD28 antibodies bound to the beads is used such that an increase in T cell expansion is observed as compared to the expansion observed using a ratio of 1:1. In one particular embodiment an increase of from about 1 to about 3 fold is observed as compared to the expansion observed using a ratio of 1:1. In one embodiment, the ratio of CD3:CD28 antibody bound to the beads ranges from 100:1 to 1:100 and all integer values there between. In one aspect of the present invention, more anti-CD28 antibody is bound to the particles than anti-CD3 antibody, i.e., the ratio of CD3:CD28 is less than one. In certain embodiments of the invention, the ratio of anti CD28 antibody to anti CD3 antibody bound to the beads is greater than 2:1. In one particular embodiment, a 1:100 CD3:CD28 ratio of antibody bound to beads is used. In another embodiment, a 1:75 CD3:CD28 ratio of antibody bound to beads is used. In a further embodiment, a 1:50 CD3:CD28 ratio of antibody bound to beads is used. In another embodiment, a 1:30 CD3:CD28 ratio of antibody bound to beads is used. In one preferred embodiment, a 1:10 CD3:CD28 ratio of antibody bound to beads is used. In another embodiment, a 1:3 CD3:CD28 ratio of antibody bound to the beads is used. In yet another embodiment, a 3:1 CD3:CD28 ratio of antibody bound to the beads is used.
[0208] Ratios of particles to cells from 1:500 to 500:1 and any integer values in between may be used to stimulate T cells or other target cells. As those of ordinary skill in the art can readily appreciate, the ratio of particles to cells may depend on particle size relative to the target cell. For example, small sized beads could only bind a few cells, while larger beads could bind many. In certain embodiments the ratio of cells to particles ranges from 1:100 to 100:1 and any integer values in-between and in further embodiments the ratio comprises 1:9 to 9:1 and any integer values in between, can also be used to stimulate T cells. The ratio of anti-CD3- and anti-CD28-coupled particles to T cells that result in T cell stimulation can vary as noted above, however certain preferred values include 1:100, 1:50, 1:40, 1:30, 1:20, 1:10, 1:9, 1:8, 1:7, 1:6, 1:5, 1:4, 1:3, 1:2, 1:1, 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, 9:1, 10:1, and 15:1 with one preferred ratio being at least 1:1 particles per T cell. In one embodiment, a ratio of particles to cells of 1:1 or less is used. In one particular embodiment, a preferred particle:cell ratio is 1:5. In further embodiments, the ratio of particles to cells can be varied depending on the day of stimulation. For example, in one embodiment, the ratio of particles to cells is from 1:1 to 10:1 on the first day and additional particles are added to the cells every day or every other day thereafter for up to 10 days, at final ratios of from 1:1 to 1:10 (based on cell counts on the day of addition). In one particular embodiment, the ratio of particles to cells is 1:1 on the first day of stimulation and adjusted to 1:5 on the third and fifth days of stimulation. In another embodiment, particles are added on a daily or every other day basis to a final ratio of 1:1 on the first day, and 1:5 on the third and fifth days of stimulation. In another embodiment, the ratio of particles to cells is 2:1 on the first day of stimulation and adjusted to 1:10 on the third and fifth days of stimulation. In another embodiment, particles are added on a daily or every other day basis to a final ratio of 1:1 on the first day, and 1:10 on the third and fifth days of stimulation. One of skill in the art will appreciate that a variety of other ratios may be suitable for use in the present invention. In particular, ratios will vary depending on particle size and on cell size and type.
[0209] In further embodiments of the present invention, the cells, such as T cells, are combined with agent-coated beads, the beads and the cells are subsequently separated, and then the cells are cultured. In an alternative embodiment, prior to culture, the agent-coated beads and cells are not separated but are cultured together. In a further embodiment, the beads and cells are first concentrated by application of a force, such as a magnetic force, resulting in increased ligation of cell surface markers, thereby inducing cell stimulation.
[0210] By way of example, cell surface proteins may be ligated by allowing paramagnetic beads to which anti-CD3 and anti-CD28 are attached (3.times.28 beads) to contact the T cells. In one embodiment, the cells (for example, 10.sup.4 to 10.sup.9 T cells) and beads (for example, DYNABEADS.RTM. M-450 CD3/CD28 T paramagnetic beads at a ratio of 1:1) are combined in a buffer, for example PBS (without divalent cations such as, calcium and magnesium). Again, those of ordinary skill in the art can readily appreciate any cell concentration may be used. For example, the target cell may be very rare in the sample and comprise only 0.01% of the sample or the entire sample (i.e., 100%) may comprise the target cell of interest. Accordingly, any cell number is within the context of the present invention. In certain embodiments, it may be desirable to significantly decrease the volume in which particles and cells are mixed together (i.e., increase the concentration of cells), to ensure maximum contact of cells and particles. For example, in one embodiment, a concentration of about 2 billion cells/ml is used. In another embodiment, greater than 100 million cells/ml is used. In a further embodiment, a concentration of cells of 10, 15, 20, 25, 30, 35, 40, 45, or 50 million cells/ml is used. In yet another embodiment, a concentration of cells from 75, 80, 85, 90, 95, or 100 million cells/ml is used. In further embodiments, concentrations of 125 or 150 million cells/ml can be used. Using high concentrations can result in increased cell yield, cell activation, and cell expansion. Further, use of high cell concentrations allows more efficient capture of cells that may weakly express target antigens of interest, such as CD28-negative T cells. Such populations of cells may have therapeutic value and would be desirable to obtain in certain embodiments. For example, using high concentration of cells allows more efficient selection of CD8+ T cells that normally have weaker CD28 expression.
[0211] In one embodiment of the present invention, the mixture may be cultured for several hours (about 3 hours) to about 14 days or any hourly integer value in between. In another embodiment, the mixture may be cultured for 21 days. In one embodiment of the invention the beads and the T cells are cultured together for about eight days. In another embodiment, the beads and T cells are cultured together for 2-3 days. Several cycles of stimulation may also be desired such that culture time of T cells can be 60 days or more. Conditions appropriate for T cell culture include an appropriate media (e.g., Minimal Essential Media or RPMI Media 1640 or, X-vivo 15, (Lonza)) that may contain factors necessary for proliferation and viability, including serum (e.g., fetal bovine or human serum), interleukin-2 (IL-2), insulin, IFN-.gamma., IL-4, IL-7, GM-CSF, IL-10, IL-12, IL-15, TGF.beta., and TNF-.alpha. or any other additives for the growth of cells known to the skilled artisan. Other additives for the growth of cells include, but are not limited to, surfactant, plasmanate, and reducing agents such as N-acetyl-cysteine and 2-mercaptoethanol. Media can include RPMI 1640, AIM-V, DMEM, MEM, .alpha.-MEM, F-12, X-Vivo 15, and X-Vivo 20, Optimizer, with added amino acids, sodium pyruvate, and vitamins, either serum-free or supplemented with an appropriate amount of serum (or plasma) or a defined set of hormones, and/or an amount of cytokine(s) sufficient for the growth and expansion of T cells. Antibiotics, e.g., penicillin and streptomycin, are included only in experimental cultures, not in cultures of cells that are to be infused into a subject. The target cells are maintained under conditions necessary to support growth, for example, an appropriate temperature (e.g., 37.degree. C.) and atmosphere (e.g., air plus 5% CO.sub.2).
[0212] T cells that have been exposed to varied stimulation times may exhibit different characteristics. For example, typical blood or apheresed peripheral blood mononuclear cell products have a helper T cell population (T.sub.H, CD4.sup.+) that is greater than the cytotoxic or suppressor T cell population (T.sub.C, CD8.sup.+). Ex vivo expansion of T cells by stimulating CD3 and CD28 receptors produces a population of T cells that prior to about days 8-9 consists predominately of T.sub.H cells, while after about days 8-9, the population of T cells comprises an increasingly greater population of T.sub.C cells. Accordingly, depending on the purpose of treatment, infusing a subject with a T cell population comprising predominately of T.sub.C cells or T.sub.H cells may be advantageous. Similarly, if an antigen-specific subset of T.sub.C cells has been isolated it may be beneficial to expand this subset to a greater degree.
[0213] Further, in addition to CD4 and CD8 markers, other phenotypic markers vary significantly, but in large part, reproducibly during the course of the cell expansion process. Thus, such reproducibility enables the ability to tailor an activated T cell product for specific purposes.
Therapeutic Application
[0214] In one aspect, the invention includes a method for treating an autoantibody-mediated kidney disease in a subject. The method comprises: administering to the subject an effective amount of a genetically modified T cell comprising a polynucleotide encoding a chimeric autoantibody receptor (CAAR), wherein the polynucleotide encodes a phospholipase A2 receptor (PLA2R) autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain, thereby treating the autoantibody-mediated kidney disease in the subject.
[0215] In another aspect, the invention includes a method for preventing or reducing glomerulus damage in a subject at risk or suffering from an autoantibody-mediated kidney disease. The method comprises: administering to the subject an effective amount of a genetically modified T cell comprising a polynucleotide encoding a CAAR, wherein the polynucleotide encodes a PLA2R autoantigen or fragment thereof, and optionally, a transmembrane domain, an intracellular domain of a costimulatory molecule, and/or a signaling domain, thereby preventing or reducing glomerulus damage in the subject.
[0216] In one embodiment, the autoantibody-mediated kidney disease is selected from the group consisting of a glomerular disease and a primary membranous nephropathy, e.g., a primary membranous nephropathy mediated by or otherwise associated with a known anti-PLA2R autoantibody and/or BCR. In another embodiment, the subject is a human.
[0217] Without wishing to be bound by any particular theory, the anti-autoantibody immune response elicited by the CAAR-modified T cells may be an active or a passive immune response. In yet another embodiment, the modified T cell targets a B cell. For example, autoantibody expressing B cells may be susceptible to indirect destruction by CAAR-redirected T cells that have previously reacted against adjacent autoantibody-expressing cells.
[0218] In one embodiment, the genetically modified T cells of the invention are modified by a fully-human CAAR. In one embodiment, the fully-human CAAR-genetically modified T cells may be a type of vaccine for ex vivo immunization and/or in vivo therapy in a mammal. In one embodiment, the mammal is a human.
[0219] With respect to ex vivo immunization, at least one of the following occurs in vitro prior to administering the cell into a mammal: i) expansion of the cells, ii) introducing to the cells a polynucleotide encoding a CAAR iii) cryopreservation of the cells.
[0220] Ex vivo procedures are well known in the art and are discussed more fully below. Briefly, cells are isolated from a mammal (e.g., a human) and genetically modified (i.e., transduced or transfected in vitro) with a vector expressing a CAAR disclosed herein. The CAAR-modified cell can be administered to a mammalian recipient to provide a therapeutic benefit. The mammalian recipient may be a human and the CAAR-modified cell can be autologous with respect to the recipient. Alternatively, the cells can be allogeneic, syngeneic or xenogeneic with respect to the recipient.
[0221] The procedure for ex vivo expansion of hematopoietic stem and progenitor cells is described in U.S. Pat. No. 5,199,942, incorporated herein by reference, can be applied to the cells of the present invention. Other suitable methods are known in the art, therefore the present invention is not limited to any particular method of ex vivo expansion of the cells. Briefly, ex vivo culture and expansion of T cells comprises: (1) collecting CD34+ hematopoietic stem and progenitor cells from a mammal from peripheral blood harvest or bone marrow explants; and (2) expanding such cells ex vivo. In addition to the cellular growth factors described in U.S. Pat. No. 5,199,942, other factors such as flt3-L, IL-1, IL-3 and c-kit ligand, can be used for culturing and expansion of the cells.
[0222] In addition to using a cell-based vaccine in terms of ex vivo immunization, the present invention also includes compositions and methods for in vivo immunization to elicit an immune response directed against an antigen in a patient.
[0223] Generally, the cells activated and expanded as described herein may be utilized in the treatment and prevention of diseases that arise in individuals who are immunocompromised. In particular, the PLA2R CAAR-modified T cells of the invention are used in the treatment of diseases, disorders and conditions associated with expression of autoantibodies. In certain embodiments, the cells of the invention are used in the treatment of patients at risk for developing autoimmune kidney diseases, disorders and conditions associated with expression of autoantibodies. Thus, the present invention provides methods for the treatment or prevention of autoimmune kidney diseases, disorders and conditions associated with expression of autoantibodies (PLA2R) comprising administering to a subject in need thereof, a therapeutically effective amount of the CAAR-modified T cells of the invention.
[0224] The CAAR-modified T cells of the present invention may be administered either alone, or as a pharmaceutical composition in combination with diluents and/or with other components such as IL-2 or other cytokines or cell populations. Briefly, pharmaceutical compositions of the present invention may comprise a target cell population as described herein, in combination with one or more pharmaceutically or physiologically acceptable carriers, diluents or excipients. Such compositions may comprise buffers such as neutral buffered saline, phosphate buffered saline and the like; carbohydrates such as glucose, mannose, sucrose or dextrans, mannitol; proteins; polypeptides or amino acids such as glycine; antioxidants; chelating agents such as EDTA or glutathione; adjuvants (e.g., aluminum hydroxide); and preservatives. Compositions of the present invention are in one aspect formulated for intravenous administration.
[0225] Pharmaceutical compositions of the present invention may be administered in a manner appropriate to the disease to be treated (or prevented). The quantity and frequency of administration will be determined by such factors as the condition of the patient, and the type and severity of the patient's disease, although appropriate dosages may be determined by clinical trials.
[0226] When "an immunologically effective amount," "an anti-BCR effective amount," "an autoimmune disease-inhibiting effective amount," or "therapeutic amount" is indicated, the precise amount of the compositions of the present invention to be administered can be determined by a physician with consideration of individual differences in age, weight, tumor size, extent of infection or metastasis, and condition of the patient (subject). It can generally be stated that a pharmaceutical composition comprising the T cells described herein may be administered at a dosage of 10.sup.4 to 10.sup.9 cells/kg body weight, in some instances 10.sup.5 to 10.sup.6 cells/kg body weight, including all integer values within those ranges. T cell compositions may also be administered multiple times at these dosages. The cells can be administered by using infusion techniques that are commonly known in immunotherapy (see, e.g., Rosenberg et al., New Eng. J. of Med. 319:1676, 1988). The optimal dosage and treatment regime for a particular patient can readily be determined by one skilled in the art of medicine by monitoring the patient for signs of disease and adjusting the treatment accordingly.
[0227] In certain embodiments, activated T cells are administered to a subject. Subsequent to administration, blood is redrawn or apheresis is performed, and T cells are activated and expanded therefrom using the methods described here, and are then reinfused back into the patient. This process can be carried out multiple times every few weeks. In certain embodiments, T cells can be activated from blood draws of from 10cc to 400cc. In certain embodiments, T cells are activated from blood draws of 20cc, 30cc, 40cc, 50cc, 60cc, 70cc, 80cc, 90cc, or 100cc. Not to be bound by theory, using this multiple blood draw/multiple reinfusion protocol, may select out certain populations of T cells.
[0228] The cells of the invention to be administered may be autologous, allogeneic or xenogeneic with respect to the subject undergoing therapy.
[0229] Administration of the cells of the invention may be carried out using any convenient means, including by aerosol inhalation, injection, ingestion, transfusion, implantation or transplantation. The compositions described herein may be administered to a patient transarterially, subcutaneously, intradermally, intranodally, intramedullary, intramuscularly, by intravenous (i.v.) injection, or intraperitoneally. In one embodiment, the T cell compositions of the present invention are administered to a patient by intradermal or subcutaneous injection. In another embodiment, the T cell compositions of the present invention are administered by i.v. injection. The compositions of T cells may be injected directly into a lymph node, or other site of pathophysiologic activity.
[0230] In certain embodiments of the present invention, cells activated and expanded using the methods described herein, or other methods known in the art where T cells are expanded to therapeutic levels, are administered to a patient in conjunction with (e.g., before, simultaneously or following) any number of relevant treatment modalities, including but not limited to treatment with agents such as immunosuppressive agents, such as azathioprine, methotrexate, mycophenolate, antibodies, or other immunoablative agents such as CAMPATH, anti-CD3 antibodies or other antibody therapies, Cytoxan, fludarabine, mycophenolic acid, steroids, FR901228, cytokines, and irradiation. These drugs may, for example, inhibit either the calcium dependent phosphatase calcineurin (cyclosporine and FK506) or inhibit the p70S6 kinase that is important for growth factor induced signaling (rapamycin). (Liu et al., Cell 66:807-815, 1991; Henderson et al., Immun. 73:316-321, 1991; Bierer et al., Curr. Opin. Immun. 5:763-773, 1993). In a further embodiment, the cell compositions of the present invention are administered to a patient in conjunction with (e.g., before, simultaneously or following) T cell ablative therapy using either chemotherapy agents such as, fludarabine, external-beam radiation therapy (XRT), cyclophosphamide, or antibodies such as OKT3 or CAMPATH. In another embodiment, the cell compositions of the present invention are administered following B-cell ablative therapy such as agents that react with CD20, e.g., Rituxan. In further embodiments, the T cells of the invention may be used in combination with complement inhibitors to reduce risk of complement-mediated cytotoxicity or an antibody anti-FcRn, IVIg, or plasmapheresis in order to reduce the anti-PLA2R antibody concentration before therapy. In yet other embodiments, a mild lymphodepletion regimen (e.g. Low-dose fludarabine or Cytoxan) might precede treatment with the T cells of the invention.
[0231] The dosage of the above treatments to be administered to a patient will vary with the precise nature of the condition being treated and the recipient of the treatment. The scaling of dosages for human administration can be performed according to art-accepted practices. The dose for CAMPATH, for example, will generally be in the range 1 to about 100 mg for an adult patient, usually administered daily for a period between 1 and 30 days. The preferred daily dose is 1 to 10 mg per day although in some instances larger doses of up to 40 mg per day may be used (described in U.S. Pat. No. 6,120,766).
EXPERIMENTAL EXAMPLES
[0232] The invention is further described in detail by reference to the following experimental examples. These examples are provided for purposes of illustration only, and are not intended to be limiting unless otherwise specified. Thus, the invention should in no way be construed as being limited to the following examples, but rather, should be construed to encompass any and all variations which become evident as a result of the teaching provided herein.
[0233] Without further description, it is believed that one of ordinary skill in the art can, using the preceding description and the following illustrative examples, make and utilize the compounds of the present invention and practice the claimed methods. The following working examples therefore, specifically point out the preferred embodiments of the present invention, and are not to be construed as limiting in any way the remainder of the disclosure.
[0234] The Materials and Methods used in the performance of the experiments disclosed herein are now described.
[0235] CAAR Constructs
TABLE-US-00003 A) Domains comprised in PLA2R-CAAR Construct 4027.CF12 Cysteine-rich (CysR) domain (Ricin B type lectin domain) (SEQ ID NO: 1) AAGGGCATCTTCGTGATCCAGAGCGAGAGCCTGAAGAAGTGCATCCAGG CCGGCAAGAGCGTGCTGACCCTGGAAAATTGCAAGCAGGCCAACAAGCA CATGCTGTGGAAATGGGTGTCCAACCACGGCCTGTTCAACATCGGCGGC TCTGGATGTCTGGGCCTGAATTTCTCTGCCCCTGAGCAGCCTCTGAGCC TGTACGAGTGTGATAGCACCCTGGTGTCCCTGAGATGGCGGTGCAACCG GAAGATGATCACAGGCCCTCTGCAGTACTCTGTGCAGGTCGCCCACGAC AATACCGTGGTGGCCAGCAGAAAGTACATCCACAAGTGGATCAGCTACG GCAGCGGCGGAGGCGACATCTGTGAATAC Linker between ricin B type lectin domain and fibronectin type II domain (SEQ ID NO: 2) CTGCACAAGGATCTGCACACCATCAAGGGCAAC Fibronectin type-2 domain (SEQ ID NO: 3) ACCCACGGAATGCCCTGCATGTTCCCGTTTCAGTACAACCACCAGTGGC ACCACGAGTGCACCAGAGAAGGCAGAGAGGACGACCTGCTTTGGTGCGC CACAACCAGCAGATACGAGCGGGATGAGAAGTGGGGCTTCTGCCCTGAT Linker between fibronectin type II domain and C-type lectin domain 1 (SEQ ID NO: 4) CCTACCTCTGCCGAAGTGGGCTGCGATACCATCTGGGAGAAAGACCTG C-type lectin domain 1 (SEQ ID NO: 5) AACAGCCACATCTGCTACCAGTTCAACCTGCTGTCCAGCCTGTCTTGGA GCGAGGCCCACAGCAGCTGTCAAATGCAAGGCGGCACACTGCTGAGCAT CACCGACGAGACAGAGGAAAACTTCATCCGCGAGCACATGAGCAGCAAG ACCGTGGAAGTGTGGATGGGACTGAACCAGCTGGATGAGCATGCCGGAT GGCAGTGGAGTGATGGCACCCCTCTGAACTACCTGAACTGGTCCCCTGA AGTGAACTTCGAGCCCTTCGTGGAAGATCACTGCGGCACCTTCAGCAGC TTCATGCCCAGCGCTTGGAGAAGCAGAGACTGCGAGAGCACCCTGCCTT ACATCTGCAAG Linker between C-type lectin domain 1 and C-type lectin domain 2 (SEQ ID NO: 6) AAGTACCTGAACCACATCGACCACGAGATCGTGGAAAAGGACGCCTGGA AGTACTACGCCACACACTGCGAGCCTGGCTGGAACCCC C-type lectin domain 2 (SEQ ID NO: 7) TACAACCGGAACTGCTACAAGCTGCAGAAAGAGGAAAAGACCTGGCACG AGGCCCTGAGAAGCTGCCAGGCCGATAATAGCGCCCTGATCGACATCAC AAGCCTGGCCGAGGTGGAATTTCTGGTCACTCTGCTGGGCGACGAGAAC GCCTCTGAGACATGGATCGGCCTGTCCAGCAACAAGATCCCCGTGTCCT TCGAGTGGTCCAACGACAGCAGCGTGATCTTCACCAACTGGCACACCCT GGAACCTCACATCTTCCCCAACAGATCCCAGCTGTGTGTGTCCGCCGAG CAGTCTGAAGGCCACTGGAAAGTGAAGAACTGCGAGGAACGGCTGTTCT ACATCTGTAAA Extracellular domains of PLA2R CAAR Construct 4027.CF12 (SEQ ID NO: 8; See FIG. 2D) AAGGGCATCTTCGTGATCCAGAGCGAGAGCCTGAAGAAGTGCATCCAGG CCGGCAAGAGCGTGCTGACCCTGGAAAATTGCAAGCAGGCCAACAAGCA CATGCTGTGGAAATGGGTGTCCAACCACGGCCTGTTCAACATCGGCGGC TCTGGATGTCTGGGCCTGAATTTCTCTGCCCCTGAGCAGCCTCTGAGCC TGTACGAGTGTGATAGCACCCTGGTGTCCCTGAGATGGCGGTGCAACCG GAAGATGATCACAGGCCCTCTGCAGTACTCTGTGCAGGTCGCCCACGAC AATACCGTGGTGGCCAGCAGAAAGTACATCCACAAGTGGATCAGCTACG GCAGCGGCGGAGGCGACATCTGTGAATACCTGCACAAGGATCTGCACAC CATCAAGGGCAACACCCACGGAATGCCCTGCATGTTCCCGTTTCAGTAC AACCACCAGTGGCACCACGAGTGCACCAGAGAAGGCAGAGAGGACGACC TGCTTTGGTGCGCCACAACCAGCAGATACGAGCGGGATGAGAAGTGGGG CTTCTGCCCTGATCCTACCTCTGCCGAAGTGGGCTGCGATACCATCTGG GAGAAAGACCTGAACAGCCACATCTGCTACCAGTTCAACCTGCTGTCCA GCCTGTCTTGGAGCGAGGCCCACAGCAGCTGTCAAATGCAAGGCGGCAC ACTGCTGAGCATCACCGACGAGACAGAGGAAAACTTCATCCGCGAGCAC ATGAGCAGCAAGACCGTGGAAGTGTGGATGGGACTGAACCAGCTGGATG AGCATGCCGGATGGCAGTGGAGTGATGGCACCCCTCTGAACTACCTGAA CTGGTCCCCTGAAGTGAACTTCGAGCCCTTCGTGGAAGATCACTGCGGC ACCTTCAGCAGCTTCATGCCCAGCGCTTGGAGAAGCAGAGACTGCGAGA GCACCCTGCCTTACATCTGCAAGAAGTACCTGAACCACATCGACCACGA GATCGTGGAAAAGGACGCCTGGAAGTACTACGCCACACACTGCGAGCCT GGCTGGAACCCCTACAACCGGAACTGCTACAAGCTGCAGAAAGAGGAAA AGACCTGGCACGAGGCCCTGAGAAGCTGCCAGGCCGATAATAGCGCCCT GATCGACATCACAAGCCTGGCCGAGGTGGAATTTCTGGTCACTCTGCTG GGCGACGAGAACGCCTCTGAGACATGGATCGGCCTGTCCAGCAACAAGA TCCCCGTGTCCTTCGAGTGGTCCAACGACAGCAGCGTGATCTTCACCAA CTGGCACACCCTGGAACCTCACATCTTCCCCAACAGATCCCAGCTGTGT GTGTCCGCCGAGCAGTCTGAAGGCCACTGGAAAGTGAAGAACTGCGAGG AACGGCTGTTCTACATCTGTAAA Extracellular domains of Construct 4027.CF12 (SEQ ID NO: 70) KGIFVIQSESLKKCIQAGKSVLTLENCKQANKHMLWKWVSNHGLFNIGG SGCLGLNFSAPEQPLSLYECDSTLVSLRWRCNRKMITGPLQYSVQVAHD NTVVASRKYIHKWISYGSGGGDICEYLHKDLHTIKGNTHGMPCMFPFQY NHQWHHECTREGREDDLLWCATTSRYERDEKWGFCPDPTSAEVGCDTIW EKDLNSHICYQFNLLSSLSWSEAHSSCQMQGGTLLSITDETEENFIREH MSSKTVEVWMGLNQLDEHAGWQWSDGTPLNYLNWSPEVNFEPFVEDHCG TFSSFMPSAWRSRDCESTLPYICKKYLNHIDHEIVEKDAWKYYATHCEP GWNPYNRNCYKLQKEEKTWHEALRSCQADNSALIDITSLAEVEFLVTLL GDENASETWIGLSSNKIPVSFEWSNDSSVIFTNWHTLEPHIFPNRSQLC VSAEQSEGHWKVKNCEERLFYICKKAGHVLSDAESGCQ Link between C-type lectin domain 2 and CD8 hinge (SEQ ID NO: 9) AAGGCCGGCCACGTGCTGTCCGATGCCGAGAGTGGATGTCAATCCGGA CD8 hinge (SEQ ID NO: 10) ACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCGT CGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGGGGG CGCAGTGCACACGAGGGGGCTGGACTTCGCCTGT Nucleic acid sequence of PLA2R-CAAR Construct 4027.CF12 (SEQ ID NO: 11) ATGCTGCTGAGCCCTAGCCTGCTGCTGCTCCTGCTTCTTGGAGCCCCTA GAGGATGTGCCGGATCTGAAGGTGTTGCCGCCGCTCTGACACCCGAGAG ACTGCTGGAATGGCAGGACAAGGGCATCTTCGTGATCCAGAGCGAGAGC CTGAAGAAGTGCATCCAGGCCGGCAAGAGCGTGCTGACCCTGGAAAATT GCAAGCAGGCCAACAAGCACATGCTGTGGAAATGGGTGTCCAACCACGG CCTGTTCAACATCGGCGGCTCTGGATGTCTGGGCCTGAATTTCTCTGCC CCTGAGCAGCCTCTGAGCCTGTACGAGTGTGATAGCACCCTGGTGTCCC TGAGATGGCGGTGCAACCGGAAGATGATCACAGGCCCTCTGCAGTACTC TGTGCAGGTCGCCCACGACAATACCGTGGTGGCCAGCAGAAAGTACATC CACAAGTGGATCAGCTACGGCAGCGGCGGAGGCGACATCTGTGAATACC TGCACAAGGATCTGCACACCATCAAGGGCAACACCCACGGAATGCCCTG CATGTTCCCGTTTCAGTACAACCACCAGTGGCACCACGAGTGCACCAGA GAAGGCAGAGAGGACGACCTGCTTTGGTGCGCCACAACCAGCAGATACG AGCGGGATGAGAAGTGGGGCTTCTGCCCTGATCCTACCTCTGCCGAAGT GGGCTGCGATACCATCTGGGAGAAAGACCTGAACAGCCACATCTGCTAC CAGTTCAACCTGCTGTCCAGCCTGTCTTGGAGCGAGGCCCACAGCAGCT GTCAAATGCAAGGCGGCACACTGCTGAGCATCACCGACGAGACAGAGGA AAACTTCATCCGCGAGCACATGAGCAGCAAGACCGTGGAAGTGTGGATG GGACTGAACCAGCTGGATGAGCATGCCGGATGGCAGTGGAGTGATGGCA CCCCTCTGAACTACCTGAACTGGTCCCCTGAAGTGAACTTCGAGCCCTT CGTGGAAGATCACTGCGGCACCTTCAGCAGCTTCATGCCCAGCGCTTGG AGAAGCAGAGACTGCGAGAGCACCCTGCCTTACATCTGCAAGAAGTACC TGAACCACATCGACCACGAGATCGTGGAAAAGGACGCCTGGAAGTACTA CGCCACACACTGCGAGCCTGGCTGGAACCCCTACAACCGGAACTGCTAC AAGCTGCAGAAAGAGGAAAAGACCTGGCACGAGGCCCTGAGAAGCTGCC AGGCCGATAATAGCGCCCTGATCGACATCACAAGCCTGGCCGAGGTGGA ATTTCTGGTCACTCTGCTGGGCGACGAGAACGCCTCTGAGACATGGATC GGCCTGTCCAGCAACAAGATCCCCGTGTCCTTCGAGTGGTCCAACGACA GCAGCGTGATCTTCACCAACTGGCACACCCTGGAACCTCACATCTTCCC CAACAGATCCCAGCTGTGTGTGTCCGCCGAGCAGTCTGAAGGCCACTGG AAAGTGAAGAACTGCGAGGAACGGCTGTTCTACATCTGTAAAAAGGCCG GCCACGTGCTGTCCGATGCCGAGAGTGGATGTCAATCCGGAACCACGAC GCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCGTCGCAGCCC CTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGGGGGCGCAGTGC ACACGAGGGGGCTGGACTTCGCCTGTGATATCTACATCTGGGCGCCCTT GGCCGGGACTTGTGGGGTCCTTCTCCTGTCACTGGTTATCACCCTTTAC
TGCAAACGGGGCAGAAAGAAACTCCTGTATATATTCAAACAACCATTTA TGAGACCAGTACAAACTACTCAAGAGGAAGATGGCTGTAGCTGCCGATT TCCAGAAGAAGAAGAAGGAGGATGTGAACTGAGAGTGAAGTTCAGCAGG AGCGCAGACGCCCCCGCGTACAAGCAGGGCCAGAACCAGCTCTATAACG AGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTTGGACAAGAGACG TGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACCCTCAG GAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACA GTGAGATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGG CCTTTACCAGGGTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTT CACATGCAGGCCCTGCCCCCTCGCTAA Amino acid sequence of PLA2R-CAAR (Construct 4027.CF12; SEQ ID NO: 12) MLLSPSLLLLLLLGAPRGCAGSEGVAAALTPERLLEWQDKGIFVIQSES LKKCIQAGKSVLTLENCKQANKHMLWKWVSNHGLFNIGGSGCLGLNFSA PEQPLSLYECDSTLVSLRWRCNRKMITGPLQYSVQVAHDNTVVASRKYI HKWISYGSGGGDICEYLHKDLHTIKGNTHGMPCMFPFQYNHQWHHECTR EGREDDLLWCATTSRYERDEKWGFCPDPTSAEVGCDTIWEKDLNSHICY QFNLLSSLSWSEAHSSCQMQGGTLLSITDETEENFIREHMSSKTVEVWM GLNQLDEHAGWQWSDGTPLNYLNWSPEVNFEPFVEDHCGTFSSFMPSAW RSRDCESTLPYICKKYLNHIDHEIVEKDAWKYYATHCEPGWNPYNRNCY KLQKEEKTWHEALRSCQADNSALIDITSLAEVEFLVTLLGDENASETWI GLSSNKIPVSFEWSNDSSVIFTNWHTLEPHIFPNRSQLCVSAEQSEGHW KVKNCEERLFYICKKAGHVLSDAESGCQSGTTTPAPRPPTPAPTIASQP LSLRPEACRPAAGGAVHTRGLDFACDIYIWAPLAGTCGVLLLSLVITLY CKRGRKKLLYIFKQPFMRPVQTTQEEDGCSCRFPEEEEGGCELRVKFSR SADAPAYKQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQ EGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPR B) Domains comprised in PLA2R-CAAR Construct 4028.CF123 (SEQ ID NO: 1) Cysteine-rich (CysR) domain (Ricin B type lectin domain) (SEQ ID NO: 2) Linker between ricin B type lectin domain and fibronectin type II domain (SEQ ID NO: 3) Fibronectin type II domain (SEQ ID NO: 4) Linker between fibronectin type II domain and C-type lectin domain 1 (SEQ ID NO: 5) C-type lectin 1 domain (SEQ ID NO: 6) Linker between C-type lectin domain 1 and C-type lectin domain 2 (SEQ ID NO: 7) C-type lectin domain 2 Linker between C-type lectin domain 2 and C-type lectin domain 3 (SEQ ID NO: 13) AAGGCCGGCCACGTGCTGTCCGATGCCGAGAGTGGATGTCAAGAAGGCT GGGAGAGA C-type lectin domain 3 (SEQ ID NO: 14) CACGGCGGCTTTTGCTACAAGATCGACACCGTGCTGCGGAGCTTCGATC AGGCCAGCAGCGGCTACTATTGCCCTCCTGCTCTGGTCACCATCACCAA CAGATTCGAGCAGGCCTTCATCACCAGCCTGATCAGCAGCGTCGTGAAG ATGAAGGACAGCTACTTCTGGATCGCCCTGCAGGACCAGAACGACACCG GCGAGTACACATGGAAGCCCGTGGGACAGAAACCCGAGCCTGTGCAGTA CACCCACTGGAACACACACCAGCCTAGATACTCCGGCGGCTGCGTGGCA ATGAGAGGCAGACATCCTCTCGGCAGATGGGAAGTGAAGCACTGTCGGC ACTTCAAGGCCATGTCTCTGTGC Extracellular domains of PLA2R (SEQ ID NO: 15; See FIG. 2D, Construct 4028.CF123) AAGGGCATCTTCGTGATCCAGAGCGAGAGCCTGAAGAAGTGCATCCAGG CCGGCAAGAGCGTGCTGACCCTGGAAAATTGCAAGCAGGCCAACAAGCA CATGCTGTGGAAATGGGTGTCCAACCACGGCCTGTTCAACATCGGCGGC TCTGGATGTCTGGGCCTGAATTTCTCTGCCCCTGAGCAGCCTCTGAGCC TGTACGAGTGTGATAGCACCCTGGTGTCCCTGAGATGGCGGTGCAACCG GAAGATGATCACAGGCCCTCTGCAGTACTCTGTGCAGGTCGCCCACGAC AATACCGTGGTGGCCAGCAGAAAGTACATCCACAAGTGGATCAGCTACG GCAGCGGCGGAGGCGACATCTGTGAATACCTGCACAAGGATCTGCACAC CATCAAGGGCAACACCCACGGAATGCCCTGCATGTTCCCGTTTCAGTAC AACCACCAGTGGCACCACGAGTGCACCAGAGAAGGCAGAGAGGACGACC TGCTTTGGTGCGCCACAACCAGCAGATACGAGCGGGATGAGAAGTGGGG CTTCTGCCCTGATCCTACCTCTGCCGAAGTGGGCTGCGATACCATCTGG GAGAAAGACCTGAACAGCCACATCTGCTACCAGTTCAACCTGCTGTCCA GCCTGTCTTGGAGCGAGGCCCACAGCAGCTGTCAAATGCAAGGCGGCAC ACTGCTGAGCATCACCGACGAGACAGAGGAAAACTTCATCCGCGAGCAC ATGAGCAGCAAGACCGTGGAAGTGTGGATGGGACTGAACCAGCTGGATG AGCATGCCGGATGGCAGTGGAGTGATGGCACCCCTCTGAACTACCTGAA CTGGTCCCCTGAAGTGAACTTCGAGCCCTTCGTGGAAGATCACTGCGGC ACCTTCAGCAGCTTCATGCCCAGCGCTTGGAGAAGCAGAGACTGCGAGA GCACCCTGCCTTACATCTGCAAGAAGTACCTGAACCACATCGACCACGA GATCGTGGAAAAGGACGCCTGGAAGTACTACGCCACACACTGCGAGCCT GGCTGGAACCCCTACAACCGGAACTGCTACAAGCTGCAGAAAGAGGAAA AGACCTGGCACGAGGCCCTGAGAAGCTGCCAGGCCGATAATAGCGCCCT GATCGACATCACAAGCCTGGCCGAGGTGGAATTTCTGGTCACTCTGCTG GGCGACGAGAACGCCTCTGAGACATGGATCGGCCTGTCCAGCAACAAGA TCCCCGTGTCCTTCGAGTGGTCCAACGACAGCAGCGTGATCTTCACCAA CTGGCACACCCTGGAACCTCACATCTTCCCCAACAGATCCCAGCTGTGT GTGTCCGCCGAGCAGTCTGAAGGCCACTGGAAAGTGAAGAACTGCGAGG AACGGCTGTTCTACATCTGTAAAAAGGCCGGCCACGTGCTGTCCGATGC CGAGAGTGGATGTCAAGAAGGCTGGGAGAGACACGGCGGCTTTTGCTAC AAGATCGACACCGTGCTGCGGAGCTTCGATCAGGCCAGCAGCGGCTACT ATTGCCCTCCTGCTCTGGTCACCATCACCAACAGATTCGAGCAGGCCTT CATCACCAGCCTGATCAGCAGCGTCGTGAAGATGAAGGACAGCTACTTC TGGATCGCCCTGCAGGACCAGAACGACACCGGCGAGTACACATGGAAGC CCGTGGGACAGAAACCCGAGCCTGTGCAGTACACCCACTGGAACACACA CCAGCCTAGATACTCCGGCGGCTGCGTGGCAATGAGAGGCAGACATCCT CTCGGCAGATGGGAAGTGAAGCACTGTCGGCACTTCAAGGCCATGTCTC TGTGC Extracellular domains of Construct 4028.CF123 (SEQ ID NO: 71) KGIFVIQSESLKKCIQAGKSVLTLENCKQANKHMLWKWVSNHGLENIGG SGCLGLNFSAPEQPLSLYECDSTLVSLRWRCNRKMITGPLQYSVQVAHD NTVVASRKYIHKWISYGSGGGDICEYLHKDLHTIKGNTHGMPCMFPFQY NHQWHHECTREGREDDLLWCATTSRYERDEKWGFCPDPTSAEVGCDTIW EKDLNSHICYQFNLLSSLSWSEAHSSCQMQGGTLLSITDETEENFIREH MSSKTVEVWMGLNQLDEHAGWQWSDGTPLNYLNWSPEVNFEPFVEDHCG TFSSFMPSAWRSRDCESTLPYICKKYLNHIDHEIVEKDAWKYYATHCEP GWNPYNRNCYKLQKEEKTWHEALRSCQADNSALIDITSLAEVEFLVTLL GDENASETWIGLSSNKIPVSFEWSNDSSVIFTNWHTLEPHIFPNRSQLC VSAEQSEGHWKVKNCEERLFYICKKAGHVLSDAESGCQEGWERHGGFCY KIDTVLRSFDQASSGYYCPPALVTITNRFEQAFITSLISSVVKMKDSYF WIALQDQNDTGEYTWKPVGQKPEPVQYTHWNTHQPRYSGGCVAMRGRHP LGRWEVKHCRHFKAMSLCKQPVENQEKAEYEERWPFHPCYL Linker between C-type lectin domain 3 and CD8 hinge (SEQ ID NO: 16) AAGCAGCCCGTGGAAAATCAAGAGAAGGCCGAGTACGAGGAACGCTGGC CTTTTCACCCTTGCTACCTGTCCGGA (SEQ ID NO: 10) CD8 hinge Nucleic acid sequence of PLA2R-CAAR (Construct 4028.CF123; SEQ ID NO: 17) ATGCTGCTGAGCCCTAGCCTGCTGCTGCTCCTGCTTCTTGGAGCCCCTA GAGGATGTGCCGGATCTGAAGGTGTTGCCGCCGCTCTGACACCCGAGAG ACTGCTGGAATGGCAGGACAAGGGCATCTTCGTGATCCAGAGCGAGAGC CTGAAGAAGTGCATCCAGGCCGGCAAGAGCGTGCTGACCCTGGAAAATT GCAAGCAGGCCAACAAGCACATGCTGTGGAAATGGGTGTCCAACCACGG CCTGTTCAACATCGGCGGCTCTGGATGTCTGGGCCTGAATTTCTCTGCC CCTGAGCAGCCTCTGAGCCTGTACGAGTGTGATAGCACCCTGGTGTCCC TGAGATGGCGGTGCAACCGGAAGATGATCACAGGCCCTCTGCAGTACTC TGTGCAGGTCGCCCACGACAATACCGTGGTGGCCAGCAGAAAGTACATC CACAAGTGGATCAGCTACGGCAGCGGCGGAGGCGACATCTGTGAATACC TGCACAAGGATCTGCACACCATCAAGGGCAACACCCACGGAATGCCCTG CATGTTCCCGTTTCAGTACAACCACCAGTGGCACCACGAGTGCACCAGA GAAGGCAGAGAGGACGACCTGCTTTGGTGCGCCACAACCAGCAGATACG
AGCGGGATGAGAAGTGGGGCTTCTGCCCTGATCCTACCTCTGCCGAAGT GGGCTGCGATACCATCTGGGAGAAAGACCTGAACAGCCACATCTGCTAC CAGTTCAACCTGCTGTCCAGCCTGTCTTGGAGCGAGGCCCACAGCAGCT GTCAAATGCAAGGCGGCACACTGCTGAGCATCACCGACGAGACAGAGGA AAACTTCATCCGCGAGCACATGAGCAGCAAGACCGTGGAAGTGTGGATG GGACTGAACCAGCTGGATGAGCATGCCGGATGGCAGTGGAGTGATGGCA CCCCTCTGAACTACCTGAACTGGTCCCCTGAAGTGAACTTCGAGCCCTT CGTGGAAGATCACTGCGGCACCTTCAGCAGCTTCATGCCCAGCGCTTGG AGAAGCAGAGACTGCGAGAGCACCCTGCCTTACATCTGCAAGAAGTACC TGAACCACATCGACCACGAGATCGTGGAAAAGGACGCCTGGAAGTACTA CGCCACACACTGCGAGCCTGGCTGGAACCCCTACAACCGGAACTGCTAC AAGCTGCAGAAAGAGGAAAAGACCTGGCACGAGGCCCTGAGAAGCTGCC AGGCCGATAATAGCGCCCTGATCGACATCACAAGCCTGGCCGAGGTGGA ATTTCTGGTCACTCTGCTGGGCGACGAGAACGCCTCTGAGACATGGATC GGCCTGTCCAGCAACAAGATCCCCGTGTCCTTCGAGTGGTCCAACGACA GCAGCGTGATCTTCACCAACTGGCACACCCTGGAACCTCACATCTTCCC CAACAGATCCCAGCTGTGTGTGTCCGCCGAGCAGTCTGAAGGCCACTGG AAAGTGAAGAACTGCGAGGAACGGCTGTTCTACATCTGTAAAAAGGCCG GCCACGTGCTGTCCGATGCCGAGAGTGGATGTCAAGAAGGCTGGGAGAG ACACGGCGGCTTTTGCTACAAGATCGACACCGTGCTGCGGAGCTTCGAT CAGGCCAGCAGCGGCTACTATTGCCCTCCTGCTCTGGTCACCATCACCA ACAGATTCGAGCAGGCCTTCATCACCAGCCTGATCAGCAGCGTCGTGAA GATGAAGGACAGCTACTTCTGGATCGCCCTGCAGGACCAGAACGACACC GGCGAGTACACATGGAAGCCCGTGGGACAGAAACCCGAGCCTGTGCAGT ACACCCACTGGAACACACACCAGCCTAGATACTCCGGCGGCTGCGTGGC AATGAGAGGCAGACATCCTCTCGGCAGATGGGAAGTGAAGCACTGTCGG CACTTCAAGGCCATGTCTCTGTGCAAGCAGCCCGTGGAAAATCAAGAGA AGGCCGAGTACGAGGAACGCTGGCCTTTTCACCCTTGCTACCTGTCCGG AACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCG TCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGGGG GCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTACATCTG GGCGCCCTTGGCCGGGACTTGTGGGGTCCTTCTCCTGTCACTGGTTATC ACCCTTTACTGCAAACGGGGCAGAAAGAAACTCCTGTATATATTCAAAC AACCATTTATGAGACCAGTACAAACTACTCAAGAGGAAGATGGCTGTAG CTGCCGATTTCCAGAAGAAGAAGAAGGAGGATGTGAACTGAGAGTGAAG TTCAGCAGGAGCGCAGACGCCCCCGCGTACAAGCAGGGCCAGAACCAGC TCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTTGGA CAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAG AACCCTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGG AGGCCTACAGTGAGATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGG GCACGATGGCCTTTACCAGGGTCTCAGTACAGCCACCAAGGACACCTAC GACGCCCTTCACATGCAGGCCCTGCCCCCTCGCTAA Amino acid sequence of PLA2R-CAAR (Construct 4028.CF123; SEQ ID NO: 18) MLLSPSLLLLLLLGAPRGCAGSEGVAAALTPERLLEWQDKGIFVIQSES LKKCIQAGKSVLTLENCKQANKHMLWKWVSNHGLFNIGGSGCLGLNFSA PEQPLSLYECDSTLVSLRWRCNRKMITGPLQYSVQVAHDNTVVASRKYI HKWISYGSGGGDICEYLHKDLHTIKGNTHGMPCMFPFQYNHQWHHECTR EGREDDLLWCATTSRYERDEKWGFCPDPTSAEVGCDTIWEKDLNSHICY QFNLLSSLSWSEAHSSCQMQGGTLLSITDETEENFIREHMSSKTVEVWM GLNQLDEHAGWQWSDGTPLNYLNWSPEVNFEPFVEDHCGTFSSFMPSAW RSRDCESTLPYICKKYLNHIDHEIVEKDAWKYYATHCEPGWNPYNRNCY KLQKEEKTWHEALRSCQADNSALIDITSLAEVEFLVTLLGDENASETWI GLSSNKIPVSFEWSNDSSVIFTNWHTLEPHIFPNRSQLCVSAEQSEGHW KVKNCEERLFYICKKAGHVLSDAESGCQEGWERHGGFCYKIDTVLRSFD QASSGYYCPPALVTITNRFEQAFITSLISSVVKMKDSYFWIALQDQNDT GEYTWKPVGQKPEPVQYTHWNTHQPRYSGGCVAMRGRHPLGRWEVKHCR HFKAMSLCKQPVENQEKAEYEERWPFHPCYLSGTTTPAPRPPTPAPTIA SQPLSLRPEACRPAAGGAVHTRGLDFACDIYIWAPLAGTCGVLLLSLVI TLYCKRGRKKLLYIFKQPFMRPVQTTQEEDGCSCRFPEEEEGGCELRVK FSRSADAPAYKQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRK NPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTY DALHMQALPPR (SEQ ID NO: 25) Nucleotide sequence of pTRPE.CysR.CD8H.BBz CAAR (Construct C) (SEQ ID NO: 46) ##STR00001## (SEQ ID NO: 47) CysR 64-459 (SEQ ID NO: 64) CD8 Hinge region 466-630 (SEQ ID NO: 20) CD8 TMD 631-702 (SEQ ID NO: 66) 4-1BB intracellular domain 703-828 (SEQ ID NO: 24) ##STR00002## Stop codon 1165-1167 ##STR00003## AAAGTGCATACAGGCTGGAAAAAGCGTGCTTACCCTGGAGAACTGCAAG CAAGCTAATAAGCATATGCTTTGGAAATGGGTTAGCAACCACGGACTCT TTAATATCGGAGGCTCCGGCTGTCTGGGCCTGAACTTCAGTGCACCGGA GCAACCGCTTTCTCTGTACGAATGTGATAGCACACTTGTTAGTCTTCGG TGGCGGTGTAACCGAAAAATGATTACAGGCCCTCTGCAATATAGTGTTC AAGTGGCCCACGACAATACAGTTGTGGCGTCTAGAAAATATATTCACAA GTGGATTTCCTACGGGAGCGGCGGAGGGGATATATGTGAATATCTTCAC AAAGACTTGCATACAATCgctagcTTCGTGCCGGTCTTCCTGCCAGCGA AGCCAACCACGACGCCAGCACCGCGACCACCAACACCTGCGCCCACCAT CGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCAGACCAGCAGCG GGGGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTACA TCTGGGCGCCCTTGGCCGGGACTTGTGGGGTCCTTCTCCTGTCACTGGT TATCACCCTTTACTGCAAGCGCGGTCGCAAGAAACTGCTCTATATTTTT AAACAGCCATTCATGAGACCTGTCCAGACCACTCAAGAGGAGGACGGAT ##STR00004## (SEQ ID NO: 26) Amino acid sequence of pTRPE.CysR.CD8H.BBz CAAR (Construct C) (SEQ ID NO: 48) ##STR00005## (SEQ ID NO: 49) CysR 22-153 (SEQ ID NO: 44) CD8 Hinge region 156-210 (SEQ ID NO: 19) CD8 TMD 211-234 (SEQ ID NO: 21) 4-1BB intracellular domain 235-276 (SEQ ID NO: 45) ##STR00006## ##STR00007## QANKHMLWKWVSNHGLFNIGGSGCLGLNFSAPEQPLSLYECDSTLVSLR WRCNRKMITGPLQYSVQVAHDNTVVASRKYIHKWISYGSGGGDICEYLH KDLHTIASFVPVFLPAKPTTTPAPRPPTPAPTIASQPLSLRPEACRPAA GGAVHTRGLDFACDIYIWAPLAGTCGVLLLSLVITLYCKRGRKKLLYIF ##STR00008## (SEQ ID NO: 27) Nucleotide sequence of pTRPE.CysR-FNII- CTLD1.CD8H.BBz CAAR (construct CF1) (SEQ ID NO: 46) IgG Signal peptide 1-57 (SEQ ID NO: 50) CysR-FNII-CTLD1 64-1053 (SEQ ID NO: 64) CD8 Hinge region 1060-1224 (SEQ ID NO: 20) CD8 TMD 1225-1296 (SEQ ID NO: 66) 4-1BB intracellular domain 1297-1422 (SEQ ID NO: 24) ##STR00009## Stop codon 1759-1761 ##STR00010## AAAATGTATACAGGCTGGCAAAAGTGTACTGACTCTTGAAAATTGCAAA CAAGCCAACAAACACATGCTGTGGAAATGGGTGTCTAATCACGGTCTCT TCAATATTGGGGGAAGTGGATGCCTCGGCCTGAATTTCTCCGCTCCCGA ACAGCCCCTCTCACTTTATGAGTGTGATTCAACTCTGGTGTCCTTGAGG TGGCGATGTAACCGCAAGATGATAACCGGCCCCCTCCAGTATTCCGTCC
AAGTAGCACACGACAATACCGTGGTGGCATCTAGGAAATACATTCATAA GTGGATATCTTATGGCAGTGGTGGCGGTGACATATGCGAGTACCTGCAC AAGGACCTCCACACAATAAAGGGGAACACGCACGGGATGCCGTGTATGT TCCCGTTCCAATATAATCATCAATGGCACCATGAGTGTACGAGAGAGGG GCGAGAAGACGACCTCCTGTGGTGTGCGACCACCTCAAGATATGAACGG GATGAGAAGTGGGGCTTTTGCCCCGACCCAACCTCCGCCGAGGTTGGTT GCGACACTATTTGGGAAAAAGATTTGAACAGTCATATATGCTATCAATT TAATTTGTTGAGTTCACTCTCCTGGAGCGAAGCGCACAGCTCTTGTCAG ATGCAAGGTGGTACATTGCTTAGCATTACTGATGAAACTGAGGAGAATT TCATTAGGGAGCATATGTCCTCAAAGACAGTAGAGGTGTGGATGGGTCT GAACCAGCTCGACGAACACGCCGGTTGGCAGTGGTCAGATGGAACGCCT CTGAATTATCTCAACTGGTCCCCTGAGGTCAACTTTGAACCGTTTGTGG AAGATCATTGTGGTACTTTTTCCAGTTTTATGCCAAGCGCCTGGCGAAG CCGAGACTGCGAGTCTACGTTGCCCTATATCTGCAAGAAGTATTTGAAT CACATAGATCATGAAATTGTTGAAgctagcTTCGTGCCGGTCTTCCTGC CAGCGAAGCCAACCACGACGCCAGCACCGCGACCACCAACACCTGCGCC CACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCAGACCA GCAGCGGGGGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATA TCTACATCTGGGCGCCCTTGGCCGGGACTTGTGGGGTCCTTCTCCTGTC ACTGGTTATCACCCTTTACTGCAAGCGCGGTCGCAAGAAACTGCTCTAT ATTTTTAAACAGCCATTCATGAGACCTGTCCAGACCACTCAAGAGGAGG ACGGATGTTCCTGTAGATTTCCTGAAGAGGAAGAGGGGGGGTGCGAGCT ##STR00011## (SEQ ID NO: 28) Amino acid sequence of pTRPE.CysR-FNII- CTLD1.CD8H.BBz CAAR (construct CF1) (SEQ ID NO: 48) IgG Signal peptide 1-19 (SEQ ID NO: 51) CysR-FNII-CTLD1 22-351 (SEQ ID NO: 43) CD8 Hinge region 354-408 (SEQ ID NO: 19) CD8 TMD 409-432 (SEQ ID NO: 21) 4-1BB intracellular domain 433-474 (SEQ ID NO: 45) ##STR00012## MEFGLSWLFLVAILKGVQCGSKGIFVIQSESLKKCIQAGKSVLTLENCK QANKHMLWKWVSNHGLFNIGGSGCLGLNFSAPEQPLSLYECDSTLVSLR WRCNRKMITGPLQYSVQVAHDNTVVASRKYIHKWISYGSGGGDICEYLH KDLHTIKGNTHGMPCMFPFQYNHQWHHECTREGREDDLLWCATTSRYER DEKWGFCPDPTSAEVGCDTIWEKDLNSHICYQFNLLSSLSWSEAHSSCQ MQGGTLLSITDETEENFIREHMSSKTVEVWMGLNQLDEHAGWQWSDGTP LNYLNWSPEVNFEPFVEDHCGTFSSFMPSAWRSRDCESTLPYICKKYLN HIDHEIVEASFVPVFLPAKPTTTPAPRPPTPAPTIASQPLSLRPEACRP AAGGAVHTRGLDFACDIYIWAPLAGTCGVLLLSLVITLYCKRGRKKLLY ##STR00013## (SEQ ID NO: 29) Nucleotide sequence of pTRPE.CysR-FNII-CTLD1-3.GS linker.BBz CAAR (construct CF123) (SEQ ID NO: 46) ##STR00014## (SEQ ID NO: 52) CysR-FNII-CTLD1-3 64-1911 (SEQ ID NO: 68) GS linker 1918-1947 (SEQ ID NO: 20) CD8 TMD 1954-2025 (SEQ ID NO: 66) 4-1BB intracellular domain 2026-2151 (SEQ ID NO: 24) CD3 zeta intracellular domain 2152-2487 Stop codon 2488-2490 ##STR00015## AAAGTGTATCCAGGCAGGCAAAAGTGTTCTCACTCTGGAAAACTGCAAA CAAGCGAACAAGCACATGTTGTGGAAGTGGGTTAGTAACCATGGACTGT TCAACATCGGAGGTAGTGGATGCCTTGGTCTCAATTTCTCTGCTCCGGA ACAGCCTTTGTCACTGTACGAATGCGACTCCACTCTCGTTAGTCTTAGA TGGCGATGCAATCGCAAAATGATTACGGGACCACTTCAATATTCAGTTC AAGTGGCACATGATAACACCGTAGTGGCCTCACGGAAATACATCCATAA ATGGATTTCTTATGGTAGCGGGGGCGGCGATATATGTGAATACCTCCAT AAGGATCTCCACACCATTAAGGGTAATACTCACGGTATGCCGTGTATGT TTCCTTTTCAGTACAATCATCAGTGGCATCATGAATGCACGAGGGAAGG ACGCGAGGACGATTTGCTCTGGTGCGCAACCACCTCACGCTACGAGAGA GACGAAAAATGGGGCTTTTGCCCGGACCCCACTAGTGCTGAGGTAGGAT GTGATACGATTTGGGAAAAGGATTTGAATTCTCATATTTGCTACCAGTT TAATCTTCTTTCATCCCTGTCCTGGTCTGAGGCTCATTCTAGTTGCCAG ATGCAAGGTGGGACTTTGCTTTCAATTACTGACGAGACTGAGGAAAATT TTATCCGAGAGCATATGTCTTCTAAAACCGTAGAGGTATGGATGGGCCT GAACCAATTGGACGAACACGCGGGCTGGCAGTGGAGCGACGGGACACCT CTCAACTACCTTAATTGGAGCCCTGAGGTAAACTTTGAACCGTTTGTCG AGGATCACTGCGGAACTTTCAGCAGCTTCATGCCTAGTGCATGGCGGTC CCGAGACTGTGAGAGCACCCTTCCATACATATGTAAAAAATACCTCAAT CACATAGACCACGAGATCGTAGAGAAGGATGCATGGAAATATTATGCTA CGCACTGTGAGCCGGGATGGAATCCTTATAACCGCAACTGTTACAAGCT GCAAAAAGAAGAGAAGACATGGCATGAGGCGCTGCGCTCATGTCAAGCG GACAATTCTGCACTTATAGATATAACTAGTTTGGCGGAGGTAGAATTTT TGGTTACGCTTCTCGGCGATGAGAATGCGTCCGAGACGTGGATAGGGTT GTCAAGCAATAAAATTCCTGTAAGTTTTGAATGGTCAAATGACTCTTCT GTCATCTTCACCAATTGGCACACACTCGAACCCCATATCTTCCCAAACC GAAGCCAGTTGTGTGTCAGCGCTGAGCAATCAGAAGGACATTGGAAAGT TAAAAACTGTGAAGAAAGACTGTTCTACATCTGTAAGAAGGCAGGACAT GTGCTTTCAGATGCGGAAAGCGGCTGTCAAGAAGGTTGGGAGCGCCATG GAGGTTTCTGTTATAAAATCGACACAGTTTTGCGATCTTTCGATCAGGC TTCAAGCGGGTACTATTGTCCTCCTGCACTGGTTACAATCACGAACCGG TTTGAACAGGCTTTTATAACTTCTTTGATTTCCAGCGTGGTTAAAATGA AGGACTCTTATTTCTGGATAGCCCTGCAAGACCAAAATGATACCGGTGA GTACACATGGAAACCGGTAGGTCAAAAGCCAGAGCCAGTCCAGTACACT CATTGGAATACCCACCAGCCTAGGTACTCCGGCGGGTGTGTGGCGATGC GGGGTCGCCACCCTCTCGGACGCTGGGAGGTGAAGCATTGCCGCCACTT CAAGGCGATGAGCTTGTGTAAACAGCCCGTCGAAAATCAGGAAAAGGCA gctagcGGTGGCGGAGGTTCTGGAGGTGGAGGTTCCtccggaATCTACA TCTGGGCGCCCTTGGCCGGGACTTGTGGGGTCCTTCTCCTGTCACTGGT TATCACCCTTTACTGCAAGCGCGGTCGCAAGAAACTGCTCTATATTTTT AAACAGCCATTCATGAGACCTGTCCAGACCACTCAAGAGGAGGACGGAT ##STR00016## (SEQ ID NO: 30) Amino acid sequence of pTRPE.CysR-FNII-CTLD1-3.GS linker.BBz CAAR (construct CF123) (SEQ ID NO: 48) ##STR00017## (SEQ ID NO: 53) CysR-FNII-CTLD1-3 22-637 (SEQ ID NO: 69) GS linker 640-649 (SEQ ID NO: 19) CD8 TMD 652-675 (SEQ ID NO: 21) 4-1BB intracellular domain 676-717 (SEQ ID NO: 45) ##STR00018## ##STR00019## QANKHMLWKWVSNHGLFNIGGSGCLGLNFSAPEQPLSLYECDSTLVSLR WRCNRKMITGPLQYSVQVAHDNTVVASRKYIHKWISYGSGGGDICEYLH KDLHTIKGNTHGMPCMFPFQYNHQWHHECTREGREDDLLWCATTSRYER DEKWGFCPDPTSAEVGCDTIWEKDLNSHICYQFNLLSSLSWSEAHSSCQ MQGGTLLSITDETEENFIREHMSSKTVEVWMGLNQLDEHAGWQWSDGTP LNYLNWSPEVNFEPFVEDHCGTFSSFMPSAWRSRDCESTLPYICKKYLN HIDHEIVEKDAWKYYATHCEPGWNPYNRNCYKLQKEEKTWHEALRSCQA DNSALIDITSLAEVEFLVTLLGDENASETWIGLSSNKIPVSFEWSNDSS VIFTNWHTLEPHIFPNRSQLCVSAEQSEGHWKVKNCEERLFYICKKAGH VLSDAESGCQEGWERHGGFCYKIDTVLRSFDQASSGYYCPPALVTITNR FEQAFITSLISSVVKMKDSYFWIALQDQNDTGEYTWKPVGQKPEPVQYT HWNTHQPRYSGGCVAMRGRHPLGRWEVKHCRHFKAMSLCKQPVENQEKA ASGGGGSGGGGSSGIYIWAPLAGTCGVLLLSLVITLYCKRGRKKLLYIF ##STR00020## (SEQ ID NO: 31)
Nucleotide sequence of pTRPE.CysR-FNII-CTLD1-3 7.GS linker.BBz CAAR (construct CF1237) (SEQ ID NO: 46) ##STR00021## (SEQ ID NO: 54) CysR-FNII-CTLD1-3 7 64-2331 (SEQ ID NO: 68) GS linker 2338-2367 (SEQ ID NO: 20) CD8 TMD 2374-2445 (SEQ ID NO: 66) 4-1BB intracellular domain 2446-2571 (SEQ ID NO: 24) ##STR00022## Stop codon 2908-2910 ##STR00023## AAAGTGTATCCAGGCAGGCAAAAGTGTTCTCACTCTGGAAAACTGCAAA CAAGCGAACAAGCACATGTTGTGGAAGTGGGTTAGTAACCATGGACTGT TCAACATCGGAGGTAGTGGATGCCTTGGTCTCAATTTCTCTGCTCCGGA ACAGCCTTTGTCACTGTACGAATGCGACTCCACTCTCGTTAGTCTTAGA TGGCGATGCAATCGCAAAATGATTACGGGACCACTTCAATATTCAGTTC AAGTGGCACATGATAACACCGTAGTGGCCTCACGGAAATACATCCATAA ATGGATTTCTTATGGTAGCGGGGGCGGCGATATATGTGAATACCTCCAT AAGGATCTCCACACCATTAAGGGTAATACTCACGGTATGCCGTGTATGT TTCCTTTTCAGTACAATCATCAGTGGCATCATGAATGCACGAGGGAAGG ACGCGAGGACGATTTGCTCTGGTGCGCAACCACCTCACGCTACGAGAGA GACGAAAAATGGGGCTTTTGCCCGGACCCCACTAGTGCTGAGGTAGGAT GTGATACGATTTGGGAAAAGGATTTGAATTCTCATATTTGCTACCAGTT TAATCTTCTTTCATCCCTGTCCTGGTCTGAGGCTCATTCTAGTTGCCAG ATGCAAGGTGGGACTTTGCTTTCAATTACTGACGAGACTGAGGAAAATT TTATCCGAGAGCATATGTCTTCTAAAACCGTAGAGGTATGGATGGGCCT GAACCAATTGGACGAACACGCGGGCTGGCAGTGGAGCGACGGGACACCT CTCAACTACCTTAATTGGAGCCCTGAGGTAAACTTTGAACCGTTTGTCG AGGATCACTGCGGAACTTTCAGCAGCTTCATGCCTAGTGCATGGCGGTC CCGAGACTGTGAGAGCACCCTTCCATACATATGTAAAAAATACCTCAAT CACATAGACCACGAGATCGTAGAGAAGGATGCATGGAAATATTATGCTA CGCACTGTGAGCCGGGATGGAATCCTTATAACCGCAACTGTTACAAGCT GCAAAAAGAAGAGAAGACATGGCATGAGGCGCTGCGCTCATGTCAAGCG GACAATTCTGCACTTATAGATATAACTAGTTTGGCGGAGGTAGAATTTT TGGTTACGCTTCTCGGCGATGAGAATGCGTCCGAGACGTGGATAGGGTT GTCAAGCAATAAAATTCCTGTAAGTTTTGAATGGTCAAATGACTCTTCT GTCATCTTCACCAATTGGCACACACTCGAACCCCATATCTTCCCAAACC GAAGCCAGTTGTGTGTCAGCGCTGAGCAATCAGAAGGACATTGGAAAGT TAAAAACTGTGAAGAAAGACTGTTCTACATCTGTAAGAAGGCAGGACAT GTGCTTTCAGATGCGGAAAGCGGCTGTCAAGAAGGTTGGGAGCGCCATG GAGGTTTCTGTTATAAAATCGACACAGTTTTGCGATCTTTCGATCAGGC TTCAAGCGGGTACTATTGTCCTCCTGCACTGGTTACAATCACGAACCGG TTTGAACAGGCTTTTATAACTTCTTTGATTTCCAGCGTGGTTAAAATGA AGGACTCTTATTTCTGGATAGCCCTGCAAGACCAAAATGATACCGGTGA GTACACATGGAAACCGGTAGGTCAAAAGCCAGAGCCAGTCCAGTACACT CATTGGAATACCCACCAGCCTAGGTACTCCGGCGGGTGTGTGGCGATGC GGGGTCGCCACCCTCTCGGACGCTGGGAGGTGAAGCATTGCCGCCACTT CAAGGCGATGAGCTTGTGTAAACAGCCCGTCGAAAATCAGGAAAAGGCA GTTAACACATCTGATATGTACCCTATGCCTAACACACTCGAATATGGGA ATAGGACGTACAAGATTATAAACGCGAACATGACGTGGTATGCTGCAAT CAAGACGTGCCTCATGCACAAAGCTCAGCTTGTGTCTATTACTGACCAA TACCACCAATCATTTTTGACAGTCGTGTTGAATCGATTGGGGTACGCCC ATTGGATCGGTCTCTTCACGACGGACAATGGGCTCAATTTTGACTGGAG TGACGGTACTAAATCATCCTTTACTTTTTGGAAGGATGAAGAAAGTTCT CTGTTGGGCGATTGCGTGTTTGCTGACTCAAATGGCCGATGGCATTCCA CAGCCTGTGAAAGTTTTCTGCAGGGAGCTATTTGCCACGTGCCTCCCGA AACGCGGCAGTCCGAACACCCGGAATTGgctagcGGTGGCGGAGGTTCT GGAGGTGGAGGTTCCtccggaATCTACATCTGGGCGCCCTTGGCCGGGA CTTGTGGGGTCCTTCTCCTGTCACTGGTTATCACCCTTTACTGCAAGCG CGGTCGCAAGAAACTGCTCTATATTTTTAAACAGCCATTCATGAGACCT GTCCAGACCACTCAAGAGGAGGACGGATGTTCCTGTAGATTTCCTGAAG ##STR00024## (SEQ ID NO: 32) Amino acid sequence of pTRPE.CysR-FNII-CTLD1-3 7.GS linker.BBz CAAR (construct CF1237) (SEQ ID NO: 48) ##STR00025## (SEQ ID NO: 55) CysR-FNII-CTLD1-3 7 22-777 (SEQ ID NO: 69) GS linker 780-789 (SEQ ID NO: 19) CD8 TMD 792-815 (SEQ ID NO: 21) 4-1BB intracellular domain 816-857 (SEQ ID NO: 45) ##STR00026## ##STR00027## QANKHMLWKWVSNHGLFNIGGSGCLGLNFSAPEQPLSLYECDSTLVSLR WRCNRKMITGPLQYSVQVAHDNTVVASRKYIHKWISYGSGGGDICEYLH KDLHTIKGNTHGMPCMFPFQYNHQWHHECTREGREDDLLWCATTSRYER DEKWGFCPDPTSAEVGCDTIWEKDLNSHICYQFNLLSSLSWSEAHSSCQ MQGGTLLSITDETEENFIREHMSSKTVEVWMGLNQLDEHAGWQWSDGTP LNYLNWSPEVNFEPFVEDHCGTFSSFMPSAWRSRDCESTLPYICKKYLN HIDHEIVEKDAWKYYATHCEPGWNPYNRNCYKLQKEEKTWHEALRSCQA DNSALIDITSLAEVEFLVTLLGDENASETWIGLSSNKIPVSFEWSNDSS VIFTNWHTLEPHIFPNRSQLCVSAEQSEGHWKVKNCEERLFYICKKAGH VLSDAESGCQEGWERHGGFCYKIDTVLRSFDQASSGYYCPPALVTITNR FEQAFITSLISSVVKMKDSYFWIALQDQNDTGEYTWKPVGQKPEPVQYT HWNTHQPRYSGGCVAMRGRHPLGRWEVKHCRHFKAMSLCKQPVENQEKA VNTSDMYPMPNTLEYGNRTYKIINANMTWYAAIKTCLMHKAQLVSITDQ YHQSFLTVVLNRLGYAHWIGLFTTDNGLNFDWSDGTKSSFTFWKDEESS LLGDCVFADSNGRWHSTACESFLQGAICHVPPETRQSEHPELASGGGGS GGGGSSGIYIWAPLAGTCGVLLLSLVITLYCKRGRKKLLYIFKQPFMRP ##STR00028## (SEQ ID NO: 33) Nucleotide sequence of pTRPE.CysR-FNII-CTLD1 7.GS linker.BBz CAAR (construct CF17) (SEQ ID NO: 46) ##STR00029## (SEQ ID NO: 56) CysR-FNII-CTLD1 7 64-1473 (SEQ ID NO: 68) GS linker 1480-1509 (SEQ ID NO: 20) CD8 TMD 1516-1587 (SEQ ID NO: 66) 4-1BB intracellular domain 1588-1713 (SEQ ID NO: 24) ##STR00030## Stop codon 2050-2052 ##STR00031## AAAATGTATCCAAGCTGGCAAATCAGTACTTACCCTTGAGAACTGCAAG CAAGCCAATAAACATATGCTGTGGAAATGGGTCTCAAACCACGGCCTCT TCAATATTGGTGGGTCAGGTTGCTTGGGGTTGAATTTCTCCGCCCCAGA GCAACCACTCAGCCTTTACGAGTGTGATTCCACACTTGTCTCTTTGCGA TGGCGCTGCAATAGGAAAATGATCACAGGCCCCCTTCAGTACTCTGTGC AAGTTGCTCATGATAACACAGTCGTGGCGAGTCGGAAATATATTCACAA ATGGATTTCTTATGGGAGTGGTGGAGGAGATATATGCGAGTATTTGCAT AAGGACTTGCACACCATCAAGGGAAACACTCACGGTATGCCATGTATGT TTCCGTTCCAATATAATCATCAATGGCACCACGAATGTACCCGAGAGGG ACGCGAGGACGATCTTCTTTGGTGCGCCACAACCTCTCGATATGAACGA GATGAGAAGTGGGGGTTTTGTCCTGACCCAACCAGTGCAGAAGTAGGGT GCGATACCATCTGGGAGAAAGACTTGAACTCACACATATGCTATCAGTT TAATTTGTTGTCTTCTTTGTCATGGAGCGAAGCTCATTCATCATGCCAG ATGCAGGGCGGGACACTGCTTTCTATCACCGACGAGACTGAGGAAAATT TTATCCGCGAGCACATGTCAAGCAAGACAGTTGAGGTTTGGATGGGGCT CAATCAACTGGACGAACACGCAGGGTGGCAGTGGTCCGATGGCACTCCG CTCAACTACCTTAACTGGAGCCCAGAGGTGAACTTTGAGCCGTTTGTCG AAGATCACTGTGGTACTTTTAGCTCCTTCATGCCGTCCGCATGGAGAAG TCGCGACTGCGAGTCAACCCTCCCTTACATCTGTAAGAAATACCTCAAC CACATAGATCACGAAATCGTAGAGGTCAATACGTCCGACATGTACCCAA
TGCCAAATACGTTGGAATATGGGAATAGGACATACAAGATAATTAACGC AAATATGACGTGGTATGCCGCAATCAAAACGTGCCTCATGCACAAGGCA CAGCTCGTGTCAATTACGGACCAGTACCACCAATCATTTCTCACAGTCG TTCTTAATCGATTGGGTTATGCACACTGGATAGGCTTGTTCACGACGGA CAATGGTTTGAACTTTGACTGGTCCGATGGAACTAAAAGTTCTTTCACT TTTTGGAAGGATGAGGAGTCCTCCTTGCTCGGGGACTGCGTCTTCGCAG ATTCAAACGGGCGCTGGCACTCAACGGCATGTGAGTCCTTCCTGCAGGG AGCTATATGCCATGTGCCACCAGAAACACGCCAGTCTGAGCACCCTGAG TTGgctagcGGTGGCGGAGGTTCTGGAGGTGGAGGTTCCtccggaATCT ACATCTGGGCGCCCTTGGCCGGGACTTGTGGGGTCCTTCTCCTGTCACT GGTTATCACCCTTTACTGCAAGCGCGGTCGCAAGAAACTGCTCTATATT TTTAAACAGCCATTCATGAGACCTGTCCAGACCACTCAAGAGGAGGACG ##STR00032## (SEQ ID NO: 34) Amino acid sequence of pTRPE.CysR-FNII-CTLD1 7.GS linker.BBz CAAR (construct CF17) (SEQ ID NO: 48) IgG Signal peptide 1-19 (SEQ ID NO: 57) CysR-FNII-CTLD1 7 22-491 (SEQ ID NO: 69) GS linker 494-503 (SEQ ID NO: 19) CD8 TMD 506-529 (SEQ ID NO: 21) 4-1BB intracellular domain 530-571 (SEQ ID NO: 45) ##STR00033## ##STR00034## QANKHMLWKWVSNHGLFNIGGSGCLGLNFSAPEQPLSLYECDSTLVSLR WRCNRKMITGPLQYSVQVAHDNTVVASRKYIHKWISYGSGGGDICEYLH KDLHTIKGNTHGMPCMFPFQYNHQWHHECTREGREDDLLWCATTSRYER DEKWGFCPDPTSAEVGCDTIWEKDLNSHICYQFNLLSSLSWSEAHSSCQ MQGGTLLSITDETEENFIREHMSSKTVEVWMGLNQLDEHAGWQWSDGTP LNYLNWSPEVNFEPFVEDHCGTFSSFMPSAWRSRDCESTLPYICKKYLN HIDHEIVEVNTSDMYPMPNTLEYGNRTYKIINANMTWYAAIKTCLMHKA QLVSITDQYHQSFLTVVLNRLGYAHWIGLFTTDNGLNFDWSDGTKSSFT FWKDEESSLLGDCVFADSNGRWHSTACESFLQGAICHVPPETRQSEHPE LASGGGGSGGGGSSGIYIWAPLAGTCGVLLLSLVITLYCKRGRKKLLYI ##STR00035## (SEQ ID NO: 35) Nucleotide sequence of pTRPE.CysR-CTLD1 7.GS linker.BBz CAAR (construct C17) (SEQ ID NO: 46) ##STR00036## (SEQ ID NO: 58) CysR-CTLD1 7 64-1314 (SEQ ID NO: 68) GS linker 1321-1350 (SEQ ID NO: 20) CD8 TMD 1357-1428 (SEQ ID NO: 66) 4-1BB intracellular domain 1429-1554 (SEQ ID NO: 24) CD3zeta intracellular domain 1555-1890 Stop codon 1891-1893 ##STR00037## AAAATGTATTCAGGCCGGTAAGAGCGTACTCACGCTTGAAAATTGCAAA CAGGCCAACAAACACATGCTTTGGAAATGGGTTTCAAATCACGGGTTGT TTAACATAGGGGGATCAGGATGTCTGGGCCTTAACTTTTCCGCACCTGA ACAACCTCTTAGTCTGTATGAGTGTGACTCAACGCTGGTCTCCTTGCGC TGGAGATGCAATCGGAAGATGATAACCGGGCCCCTCCAGTATTCCGTTC AGGTCGCCCACGATAATACTGTTGTTGCATCCCGAAAATATATTCATAA GTGGATCTCCTACGGGAGTGGAGGGGGCGATATTTGTGAATACCTCCAC AAGGATCTGCACACTATCACTTCTGCGGAAGTAGGCTGTGACACAATCT GGGAGAAAGATCTGAATTCACACATTTGCTATCAGTTCAATCTTCTGAG TTCTTTGAGCTGGTCCGAAGCACATTCATCCTGTCAGATGCAAGGTGGA ACACTCTTGTCAATAACAGATGAAACGGAAGAGAACTTTATTAGAGAAC ATATGTCCTCAAAGACTGTGGAGGTGTGGATGGGACTTAACCAGCTCGA TGAACATGCAGGATGGCAGTGGAGTGACGGAACGCCACTGAACTACCTG AATTGGAGCCCAGAGGTGAATTTCGAGCCTTTCGTAGAGGACCATTGCG GTACTTTTTCATCTTTTATGCCCAGCGCATGGAGATCCCGAGATTGTGA AAGCACGCTGCCCTATATTTGTAAAAAGTACCTGAACCACATAGATCAT GAGATAGTTGAGGTAAATACAAGTGATATGTACCCCATGCCGAACACAC TCGAGTACGGAAATAGAACCTACAAGATAATCAACGCTAACATGACCTG GTACGCGGCCATTAAGACCTGCCTCATGCACAAGGCTCAACTCGTCAGT ATTACTGACCAATATCACCAGTCATTTCTCACCGTCGTGTTGAATCGCC TCGGTTACGCCCACTGGATCGGTTTGTTTACAACGGACAATGGACTCAA TTTCGATTGGTCAGACGGAACCAAATCTAGTTTTACCTTCTGGAAAGAC GAGGAATCAAGCCTGCTTGGGGACTGCGTATTTGCGGACTCTAATGGCC GATGGCATAGTACAGCGTGTGAGAGCTTTTTGCAGGGGGCGATTTGTCA TGTTCCGCCGGAAACCCGCCAAAGCGAGCATCCAGAATTGgctagcGGT GGCGGAGGTTCTGGAGGTGGAGGTTCCtccggaATCTACATCTGGGCGC CCTTGGCCGGGACTTGTGGGGTCCTTCTCCTGTCACTGGTTATCACCCT TTACTGCAAGCGCGGTCGCAAGAAACTGCTCTATATTTTTAAACAGCCA TTCATGAGACCTGTCCAGACCACTCAAGAGGAGGACGGATGTTCCTGTA ##STR00038## (SEQ ID NO: 36) Amino acid sequence of pTRPE.CysR-CTLD1 7.GS linker.BBz CAAR (construct C17) (SEQ ID NO: 48) ##STR00039## (SEQ ID NO: 59) CysR-CTLD1 7 22-438 (SEQ ID NO: 69) GS linker 441-450 (SEQ ID NO: 19) CD8 TMD 453-476 (SEQ ID NO: 21) 4-1BB intracellular domain 477-518 (SEQ ID NO: 45) ##STR00040## ##STR00041## QANKHMLWKWVSNHGLFNIGGSGCLGLNFSAPEQPLSLYECDSTLVSLR WRCNRKMITGPLQYSVQVAHDNTVVASRKYIHKWISYGSGGGDICEYLH KDLHTITSAEVGCDTIWEKDLNSHICYQFNLLSSLSWSEAHSSCQMQGG TLLSITDETEENFIREHMSSKTVEVWMGLNQLDEHAGWQWSDGTPLNYL NWSPEVNFEPFVEDHCGTFSSFMPSAWRSRDCESTLPYICKKYLNHIDH EIVEVNTSDMYPMPNTLEYGNRTYKIINANMTWYAAIKTCLMHKAQLVS ITDQYHQSFLTVVLNRLGYAHWIGLFTTDNGLNFDWSDGTKSSFTFWKD EESSLLGDCVFADSNGRWHSTACESFLQGAICHVPPETRQSEHPELASG GGGSGGGGSSGIYIWAPLAGTCGVLLLSLVITLYCKRGRKKLLYIFKQP ##STR00042## (SEQ ID NO: 37) Nucleotide sequence of 4025 pTRPE.CysR.CD8H.BBz CAAR (construct 4025.C) (SEQ ID NO: 65) ##STR00043## (SEQ ID NO: 67) PLA2R pro-peptide 61-117 (SEQ ID NO: 60) ##STR00044## (SEQ ID NO: 42) CD8 Hinge region 511-645 (SEQ ID NO: 20) CD8 TMD 646-717 (SEQ ID NO: 22) 4-1BB intracellular domain 718-843 (SEQ ID NO: 72) ##STR00045## Stop codon 1180-1182 ##STR00046## ACTGCTGGAATGGCAGGACAAGGGCATCTTCGTGATCCAGAGCGAGAGC CTGAAGAAGTGCATCCAGGCCGGCAAGAGCGTGCTGACCCTGGAAAATT GCAAGCAGGCCAACAAGCACATGCTGTGGAAATGGGTGTCCAACCACGG CCTGTTCAACATCGGCGGCTCTGGATGTCTGGGCCTGAATTTCTCTGCC CCTGAGCAGCCTCTGAGCCTGTACGAGTGTGATAGCACCCTGGTGTCCC TGAGATGGCGGTGCAACCGGAAGATGATCACAGGCCCTCTGCAGTACTC TGTGCAGGTCGCCCACGACAATACCGTGGTGGCCAGCAGAAAGTACATC CACAAGTGGATCAGCTACGGCAGCGGCGGAGGCGACATCTGTGAATACC TGCACAAGGATCTGtccggaACCACGACGCCAGCGCCGCGACCACCAAC ACCGGCGCCCACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCG
TGCCGGCCAGCGGCGGGGGGCGCAGTGCACACGAGGGGGCTGGACTTCG CCTGTGATATCTACATCTGGGCGCCCTTGGCCGGGACTTGTGGGGTCCT TCTCCTGTCACTGGTTATCACCCTTTACTGCAAACGGGGCAGAAAGAAA CTCCTGTATATATTCAAACAACCATTTATGAGACCAGTACAAACTACTC AAGAGGAAGATGGCTGTAGCTGCCGATTTCCAGAAGAAGAAGAAGGAGG ##STR00047## (SEQ ID NO: 38) Amino acid sequence of 4025 pTRPE.CysR.CD8H.BBz CAAR (construct 4025.C) ##STR00048## (SEQ ID NO: 61) CysR 40-168 (SEQ ID NO: 43) CD8 Hinge region 171-215 (SEQ ID NO: 19) CD8 TMD 216-239 (SEQ ID NO: 21) 4-1BB intracellular domain 240-281 (SEQ ID NO: 23) ##STR00049## MLLSPSLLLLLLLGAPRGCAGSEGVAAALTPERLLEWQDKGIFVIQSES LKKCIQAGKSVLTLENCKQANKHMLWKWVSNHGLFNIGGSGCLGLNFSA PEQPLSLYECDSTLVSLRWRCNRKMITGPLQYSVQVAHDNTVVASRKYI HKWISYGSGGGDICEYLHKDLSGTTTPAPRPPTPAPTIASQPLSLRPEA CRPAAGGAVHTRGLDFACDIYIWAPLAGTCGVLLLSLVITLYCKRGRKK ##STR00050## (SEQ ID NO: 39) Nucleotide sequence of 4026 pTRPE. CysR-FNII-CTLD1.CD8H.BBz CAAR (construct 4026.CF1) (SEQ ID NO: 65) ##STR00051## PLA2R pro-peptide 61-117 (SEQ ID NO: 62) CysR-FNII-CTLD1 118-1143 (SEQ ID NO: 42) CD8 Hinge region 1149-1284 (SEQ ID NO: 20) CD8 TMD 1285-1356 (SEQ ID NO: 22) 4-1BB intracellular domain 1357-1482 (SEQ ID NO: 72) ##STR00052## Stop codon 1819-1821 ##STR00053## ACTGCTGGAATGGCAGGACAAGGGCATCTTCGTGATCCAGAGCGAGAGC CTGAAGAAGTGCATCCAGGCCGGCAAGAGCGTGCTGACCCTGGAAAATT GCAAGCAGGCCAACAAGCACATGCTGTGGAAATGGGTGTCCAACCACGG CCTGTTCAACATCGGCGGCTCTGGATGTCTGGGCCTGAATTTCTCTGCC CCTGAGCAGCCTCTGAGCCTGTACGAGTGTGATAGCACCCTGGTGTCCC TGAGATGGCGGTGCAACCGGAAGATGATCACAGGCCCTCTGCAGTACTC TGTGCAGGTCGCCCACGACAATACCGTGGTGGCCAGCAGAAAGTACATC CACAAGTGGATCAGCTACGGCAGCGGCGGAGGCGACATCTGTGAATACC TGCACAAGGATCTGCACACCATCAAGGGCAACACCCACGGAATGCCCTG CATGTTCCCGTTTCAGTACAACCACCAGTGGCACCACGAGTGCACCAGA GAAGGCAGAGAGGACGACCTGCTTTGGTGCGCCACAACCAGCAGATACG AGCGGGATGAGAAGTGGGGCTTCTGCCCTGATCCTACCTCTGCCGAAGT GGGCTGCGATACCATCTGGGAGAAAGACCTGAACAGCCACATCTGCTAC CAGTTCAACCTGCTGTCCAGCCTGTCTTGGAGCGAGGCCCACAGCAGCT GTCAAATGCAAGGCGGCACACTGCTGAGCATCACCGACGAGACAGAGGA AAACTTCATCCGCGAGCACATGAGCAGCAAGACCGTGGAAGTGTGGATG GGACTGAACCAGCTGGATGAGCATGCCGGATGGCAGTGGAGTGATGGCA CCCCTCTGAACTACCTGAACTGGTCCCCTGAAGTGAACTTCGAGCCCTT CGTGGAAGATCACTGCGGCACCTTCAGCAGCTTCATGCCCAGCGCTTGG AGAAGCAGAGACTGCGAGAGCACCCTGCCTTACATCTGCAAGAAGTACC TGAACCACATCGACCACGAGATCGTGGAAAAGGACGCCTGGAAGTACTA CGCCACACACTGCGAGtccggaACCACGACGCCAGCGCCGCGACCACCA ACACCGGCGCCCACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGG CGTGCCGGCCAGCGGCGGGGGGCGCAGTGCACACGAGGGGGCTGGACTT CGCCTGTGATATCTACATCTGGGCGCCCTTGGCCGGGACTTGTGGGGTC CTTCTCCTGTCACTGGTTATCACCCTTTACTGCAAACGGGGCAGAAAGA AACTCCTGTATATATTCAAACAACCATTTATGAGACCAGTACAAACTAC TCAAGAGGAAGATGGCTGTAGCTGCCGATTTCCAGAAGAAGAAGAAGGA ##STR00054## (SEQ ID NO: 40) Amino acid sequence of 4026 pTRPE. CysR-FNII-CTLD1.CD8H.BBz CAAR (construct 4026.CF1) ##STR00055## (SEQ ID NO: 63) CysR-FNII-CTLD1 40-381 (SEQ ID NO: 43) CD8 Hinge region 384-428 (SEQ ID NO: 19) CD8 TMD 429-452 (SEQ ID NO: 21) 4-1BB intracellular domain 453-494 (SEQ ID NO: 23) ##STR00056## ##STR00057## LKKCIQAGKSVLTLENCKQANKHMLWKWVSNHGLFNIGGSGCLGLNFSA PEQPLSLYECDSTLVSLRWRCNRKMITGPLQYSVQVAHDNTVVASRKYI HKWISYGSGGGDICEYLHKDLHTIKGNTHGMPCMFPFQYNHQWHHECTR EGREDDLLWCATTSRYERDEKWGFCPDPTSAEVGCDTIWEKDLNSHICY QFNLLSSLSWSEAHSSCQMQGGTLLSITDETEENFIREHMSSKTVEVWM GLNQLDEHAGWQWSDGTPLNYLNWSPEVNFEPFVEDHCGTFSSFMPSAW RSRDCESTLPYICKKYLNHIDHEIVEKDAWKYYATHCESGTTTPAPRPP TPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACDIYIWAPLAGTCGV LLLSLVITLYCKRGRKKLLYIFKQPFMRPVQTTQEEDGCSCRFPEEEEG ##STR00058## >BC144631.1 Homo sapiens phospholipase A2 receptor 1, 180kDa, mRNA (cDNA clone MGC:178179 IMAGE:9053162), complete cds (SEQ ID NO: 41) CCGGAGAGCCCAGTGGTTAGCGATGCTGCTGTCGCCGTCGCTGCTGCTG CTGCTGCTGCTGGGGGCGCCGCGGGGCTGCGCCGAGGGTGTGGCGGCGG CGCTTACCCCCGAGCGGCTCCTGGAGTGGCAGGATAAAGGAATATTTGT TATCCAAAGTGAGAGTCTCAAGAAATGCATTCAAGCAGGTAAATCGGTT CTGACCCTGGAGAACTGCAAGCAAGCAAACAAGCACATGCTGTGGAAAT GGGTTTCAAACCATGGCCTCTTTAACATAGGAGGCAGCGGTTGCCTGGG CCTGAATTTCTCCGCCCCAGAGCAGCCATTAAGCTTATATGAATGTGAC TCCACCCTCGTTTCCTTACGGTGGCGCTGTAACAGGAAGATGATCACAG GCCCGCTGCAGTACTCTGTCCAGGTGGCGCATGACAACACAGTGGTGGC CTCACGGAAGTATATTCATAAGTGGATTTCTTATGGGTCAGGTGGTGGA GACATTTGTGAATATCTACACAAAGATTTGCATACAATCAAAGGGAACA CCCACGGGATGCCGTGTATGTTTCCCTTCCAGTATAACCATCAGTGGCA TCATGAATGTACCCGTGAAGGTCGGGAAGATGACTTACTGTGGTGTGCC ACGACAAGCCGTTATGAAAGAGATGAAAAGTGGGGATTTTGCCCTGATC CCACCTCTGCAGAAGTAGGTTGTGATACTATTTGGGAGAAGGACCTCAA TTCACACATTTGCTACCAGTTCAACCTGCTTTCATCTCTCTCTTGGAGT GAGGCACATTCTTCATGCCAGATGCAAGGAGGTACGCTGTTAAGTATTA CAGATGAAACTGAAGAAAATTTCATAAGGGAGCACATGAGCAGTAAAAC AGTGGAGGTGTGGATGGGCCTCAATCAGCTGGATGAACACGCTGGCTGG CAGTGGTCTGATGGAACGCCGCTCAACTATCTGAATTGGAGCCCAGAGG TAAATTTTGAGCCATTTGTTGAAGATCACTGTGGAACATTTAGTTCATT TATGCCAAGTGCCTGGAGGAGTCGGGATTGTGAGTCCACCTTGCCATAT ATATGTAAAAAATATCTAAACCACATTGATCATGAAATAGTTGAAAAAG ATGCGTGGAAATATTATGCTACCCACTGTGAGCCTGGCTGGAATCCCTA CAATCGTAATTGCTACAAACTTCAGAAAGAAGAAAAGACCTGGCATGAG GCTCTGCGTTCTTGTCAGGCTGATAACAGTGCATTAATAGACATAACCT CATTAGCAGAGGTGGAGTTTCTTGTAACCCTCCTTGGAGATGAAAATGC ATCAGAAACATGGATTGGTTTGAGCAGCAATAAAATTCCAGTTTCCTTT GAATGGTCTAATGACTCTTCAGTCATCTTTACTAATTGGCACACACTTG AGCCCCACATTTTTCCAAATAGAAGCCAGCTGTGTGTCTCAGCAGAGCA GTCTGAGGGACACTGGAAAGTCAAAAATTGTGAAGAAAGACTTTTTTAC ATTTGTAAAAAAGCAGGCCATGTCCTCTCTGATGCTGAATCAGGATGTC AAGAGGGATGGGAGAGACATGGTGGATTCTGTTACAAAATTGACACAGT CCTTCGAAGCTTTGACCAAGCTTCCAGCGGTTATTACTGTCCTCCTGCA CTTGTAACCATTACAAACAGGTTTGAACAGGCTTTTATTACCAGTTTGA TCAGTAGTGTGGTAAAAATGAAGGACAGTTATTTTTGGATAGCTCTTCA
GGACCAAAATGATACGGGAGAATACACTTGGAAGCCAGTAGGGCAGAAA CCCGAGCCGGTGCAGTACACACACTGGAACACACACCAGCCGCGCTACA GTGGTGGCTGTGTTGCCATGCGAGGAAGGCATCCACTTGGTCGCTGGGA AGTGAAGCACTGTCGGCACTTTAAGGCAATGTCCTTGTGCAAGCAGCCA GTTGAAAATCAGGAAAAAGCAGAGTATGAAGAGAGATGGCCCTTTCACC CCTGCTATTTGGACTGGGAGTCAGAGCCTGGTCTGGCCAGTTGCTTCAA GGTATTTCATAGTGAAAAAGTTCTGATGAAAAGAACATGGAGAGAAGCT GAAGCATTTTGCGAAGAATTTGGAGCTCATCTTGCAAGCTTTGCCCATA TTGAGGAAGAGAATTTTGTGAATGAGCTCTTACATTCAAAATTTAATTG GACAGAAGAAAGGCAGTTCTGGATTGGATTTAATAAAAGAAACCCACTG AATGCCGGCTCATGGGAGTGGTCTGATAGAACTCCTGTTGTCTCTTCGT TTTTAGACAACACTTATTTTGGAGAAGATGCAAGAAACTGTGCTGTTTA TAAGGCAAACAAAACATTGCTGCCCTTACACTGTGGTTCCAAACGTGAA TGGATATGCAAAATCCCAAGAGATGTGAAACCCAAGATTCCGTTCTGGT ACCAGTACGATGTACCCTGGCTCTTTTATCAGGATGCAGAATACCTTTT TCATACCTTTGCCTCAGAATGGTTGAACTTTGAGTTTGTCTGTAGCTGG CTGCACAGTGATCTTCTCACAATTCATTCTGCACATGAGCAAGAATTCA TCCACAGCAAAATAAAAGCGCTATCAAAGTATGGTGCAAGTTGGTGGAT TGGACTTCAAGAAGAAAGAGCCAATGATGAATTTCGCTGGAGAGATGGA ACACCAGTGATATACCAGAACTGGGACACAGGAAGAGAAAGAACTGTGA ATAATCAGAGCCAGAGATGTGGCTTTATTTCTTCTATAACAGGACTCTG GGGTAGTGAAGAGTGTTCAGTTTCTATGCCTAGTATCTGTAAGCGAAAA AAGGTTTGGCTCATAGAGAAAAAGAAAGATACACCAAAACAACATGGAA CGTGTCCCAAAGGATGGCTATATTTTAACTATAAGTGCCTTCTGCTGAA TATCCCCAAAGACCCAAGCAGTTGGAAGAACTGGACGCATGCTCAACAT TTCTGTGCTGAAGAAGGGGGGACCCTGGTCGCCATTGAAAGTGAGGTGG AGCAAGCTTTCATTACTATGAATCTTTTTGGCCAGACCACCAGTGTGTG GATAGGTTTACAAAATGATGATTATGAAACATGGCTAAATGGAAAGCCT GTGGTATATTCTAACTGGTCTCCATTTGATATAATAAATATTCCAAGTC ACAATACCACTGAAGTTCAGAAACACATTCCTCTCTGTGCCTTACTCTC AAGTAATCCTAATTTTCATTTCACTGGAAAATGGTATTTTGAAGACTGT GGAAAGGAAGGCTATGGGTTTGTTTGTGAAAAAATGCAAGATACTTCTG GACACGGTGTAAATACATCTGATATGTATCCAATGCCCAATACCTTAGA ATATGGAAACAGAACTTACAAAATAATTAATGCAAATATGACTTGGTAT GCAGCAATAAAAACCTGCCTGATGCACAAAGCACAACTGGTCAGCATCA CAGACCAGTATCACCAGTCCTTCCTCACTGTTGTCCTCAACCGGCTAGG ATATGCCCACTGGATTGGACTGTTCACCACAGATAATGGTCTTAATTTT GACTGGTCTGATGGCACCAAATCTTCTTTCACTTTTTGGAAAGATGAGG AGTCCTCCCTCCTTGGTGACTGCGTTTTTGCCGACAGCAACGGACGCTG GCATAGCACAGCCTGCGAGTCATTTCTGCAAGGTGCCATTTGTCATGTG CCACCTGAAACAAGACAATCTGAACACCCAGAGTTGTGCTCAGAAACAT CTATTCCCTGGATAAAATTTAAAAGTAATTGCTACAGTTTTTCTACAGT CCTAGACAGTATGAGTTTTGAGGCTGCTCATGAATTTTGCAAAAAGGAA GGTTCTAATCTTTTAACAATCAAGGATGAGGCTGAAAATGCATTTCTCC TAGAAGAGCTGTTTGCTTTTGGTTCTTCTGTCCAGATGGTTTGGTTGAA TGCTCAATTTGATGATGAAACCATAAAGTGGTTTGATGGAACTCCCACA GACCAGTCAAACTGGGGCATTCGGAAGCCAGACACAGACTACTTCAAGC CCCATCATTGTGTTGCCTTGAGGATCCCTGAAGGATTATGGCAGCTATC CCCGTGTCAAGAAAAAAAAGGCTTTATATGTAAAATGGAGGCAGATATT CACACTGCAGAGGCGCTGCCAGAAAAAGGACCAAGTCACAGCATCATTC CTCTTGCGGTTGTACTGACACTGATAGTCATTGTGGCCATTTGCACACT TTCCTTCTGCATATACAAGCATAACGGTGGCTTCTTCAGGAGACTTGCA GGGTTTCGGAATCCTTACTATCCTGCAACCAACTTTAGTACAGTATATT TAGAAGAAAATATTCTCATTTCTGATCTTGAGAAGAGTGACCAATAATA ATGAGGTCAGAGAATGCCACAGACACCAGGG
[0236] Production of HIV-1 Based Self-Inactivating Lentivirus
[0237] 293T cells were transfected at a confluence of 90% with a mixture of the pTRPE-gene-of-interest, the packaging plasmid pGAG-Pol and pRSV-Rev and the envelope plasmid pCI VSVg in a complex with Lipofectamine 2000. Lentivirus containing supernatant was harvested after 24 and 48 hours, filtered through a 0.45 micron polyethersulfone (PES) membrane, concentrated at 12,000.times.g for 24 hours at 4.degree. C., and stored at -80.degree. C.
[0238] Stimulation, Transduction, and Expansion of Primary Human T Cells.
[0239] Primary human T cells were cultured in CTS OpTmizer T cell expansion SFM with 5% HSAB, 2.6% supplement, 1% penicillin/streptomycin and 1% glutamax. Bulk T cells (CD4.sup.+ and CD8.sup.+) were stimulated with anti-CD3 and anti-CD28 beads at a bead:cell ratio of 3:1. The culture medium was supplemented with 100 IU/ml interleukin 2. 24 hours after stimulation, 10.sup.6 T cells were transduced with CAAR or mock transduced. Expansion of the T cells were monitored for 8-12 days. Cell surface expression of the CAAR constructs was validated by flow cytometry with IgG purified from the plasma of a patient with primary membranous nephropathy and isotype control antibody.
[0240] Reporter Assay with NFAT-GFP Jurkat T Cells
[0241] Jurkat cells were cultured at 37.degree. C. with 5% CO2 in a completely humidified environment using RPMI1640, HEPES 10 mM, and FBS at 10%. To test signal transduction by CAAR-target interaction, the CAAR constructs were expressed in a Jurkat reporter cell line that has been selected (G418) for stable expression of GFP controlled under an NFAT response element, facilitating GFP expression after CAAR engagement and PLCgamma and IP3 mediated intracellular calcium release. Jurkat cells were transduced with CAAR lentivirus at a multiplicity of infection of 3 and expression of the CAAR construct was validated after >72 hours with an anti-PLA2R monoclonal antibody (Novus Biologicals.RTM.) and IgG purified from the plasma of a patient with primary membranous nephropathy by flow cytometry. For characterization of CAAR-target interaction, the CAAR Jurkat cells were incubated at 37.degree. C. for 6 hours with plate-bound anti-PLA2R monoclonal as below. GFP expression was validated by flow cytometry.
[0242] Stimulation
[0243] 1 million Jurkat-NFAT-GFP cells were transduced with each lentivirus encoding for PLA2R CAAR construct C, CF1, CF123, CF1237, CD17, C17, 4025.C, 4026.CF1, 4027.CF12, 4028.CF123 and untransduced Jurkat cells were used as negative control. Transduced and untransduced cells were plated in a 96-well plate at 2.times.10.sup.5 cells/well in 100 .mu.l cell culture medium.
[0244] The stimulation conditions were as follows: [0245] Novus monoclonal antibody (mAb) anti-PLA2R either 0.5 .mu.g/well or 1 .mu.g/well; Clone 12-6-5 [0246] Sigma polyclonal antibody anti-PLA2R 0.5 .mu.g/well; [0247] Isotype antibody for Novus mAb anti-PLA2R (0.5 .mu.g/stain) [0248] medium only (no stimulation) [0249] PMA/Ionomycin (maximum activation)
[0250] Or, transduced and untransduced cells were plated in a 96-well plate at 5.times.10.sup.4 cells/well in 100 .mu.l cell culture medium.
[0251] Cells and antibodies were incubated at 37.degree. C. for 6 hours. At the end of the 6 h, cells were examined by flow cytometry for expression of NFAT-GFP as an indicator of T cell activation.
[0252] In Vitro IFN.gamma. Cytokine Assay
[0253] PLA2R CAAR or mock transduced primary human T cells were cultured for 24 h at 5.times.10.sup.4/well in a 96-well plate coated with 10 .mu.g/ml either IgG purified from plasma of patient with primary membranous nephropathy or control human IgG. Culture supernatants were then harvested for detection of IFN-.gamma. by ELISA.
[0254] In Vivo Efficacy Testing of CAAR T Cells
[0255] To test the in vivo efficacy, anti-PLA2R BCR-expressing cells, for example hybridoma cells or human cell lines, targeting human PLA2R are generated. PLA2R BCR-expressing cells are transduced with lentiviral vectors expressing click-beetle green or click-beetle red luciferase and GFP. To test the efficacy of CAAR-T cells against the PLA2R-specific target cells, approximately 2.times.10.sup.5 anti-PLA2R cells are injected intravenously into NSG mice (age 6-8 weeks), after pre-treatment of mice daily for 2 days with 600 mg/kg intravenous immunoglobulin to minimize Fc.gamma.R-mediated toxicity against BCR-expressing cells. After 4-5 days, PLA2R CAAR T cells (approximately 10.sup.7 cells per mouse) are injected intravenously. Bioluminescence and/or serum autoantibodies are quantified every 2-3 days after injection to monitor CAAR T cell efficacy.
[0256] The results of the experiments are now described.
Example 1
[0257] Four PLA2R constructs representing different domains of the PLA2R were cloned into a CAAR expression system (4025.C, 4026.CF1, Construct 4027.CF12 and 4028.CF123). The general schematics of the extracellular domains of PLA2R of particular interest in the invention are illustrated in FIG. 1. The N terminal domain is an immunodominant epitope that is recognized by patient serum antibodies. The CysR and CTLD3 domain interact to form a ring-like structure that may better present immunodominant epitopes; hence constructs including the CysR through CTLD3 domains were generated. In addition to the cysteine-rich domain, serum antibody immunoreactivity to the C-type lectin domains 1 and 7 has also been reported.
[0258] The PLA2R CAAR constructs illustrated in FIGS. 2A-2G. All constructs use the CD8 transmembrane domain and BBZ intracellular domain and differ in their extracellular composition.
[0259] Construct 4025.C was comprised of a cysteine rich extracellular domain, a CD8 hinge region, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2A, SEQ ID NOs: 37 & 38).
[0260] Construct 4026.CF1 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, and a C-type lectin 1 domain. Construct 4026.CF1 also included a CD8 hinge region, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2B, SEQ ID NOs: 39 & 40).
[0261] Construct 4027.CF12 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, a C-type lectin 1 domain, and a C-type lectin 2 domain. Construct 4027.CF12 also included a CD8 hinge region, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2C, SEQ ID NOs: 11 & 12).
[0262] Construct 4028.CF123 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, a C-type lectin 1 domain, a C-type lectin 2 domain, and a C-type lectin 3 domain. Construct 4028.CF123 also included a CD8 hinge region, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2D, SEQ ID NOs: 17 & 18). A detailed map of PLA2R CAAR Construct 4028.CF123 is shown in FIG. 3A.
[0263] To test whether the CAAR molecules were functional, the following experiments were conducted:
[0264] 1 million Jurkat-NFAT-GFP cells were transduced with each lentivirus encoding for 4025.C, 4027.CF12, 4028.CF123 and untransduced Jurkat cells were used as negative control. Transduced and untransduced cells were plated in a 96-well plate at 2.times.10.sup.5 cells/well in 100 ul cell culture medium. FIG. 4 shows the PLA2R CAAR expression. The expression of PLA2R CAAR by Jurkat NFAT-GFP cells transduced with different PLA2R lentiviral constructs was measured by staining with soluble anti-PLA2R monoclonal antibody conjugated to PE. Jurkat T cells bearing construct 4027.CF12 or 4028.CF123, but not those T cells bearing construct 4025.C, which does not include the CTLD2 domain and hence is not detectable by the mAb used, exhibited detectable surface expression of the CAAR.
[0265] FIGS. 5 and 6 show T cell activation by GFP expression of Jurkat cells transduced with the indicated constructs. Negative and positive controls are shown in FIG. 5. Test samples are shown in FIG. 6.
[0266] Cells and antibodies were incubated at 37.degree. C. for 6 hours. At the end of the 6 h, cells were examined for expression of NFAT-GFP as an indicator of T cell activation.
[0267] FIG. 5 depicts the negative and positive controls for PLA2R CAAR T cell activation using an isotype control (top, negative control) or PMA+ionomycin (bottom, positive control).
[0268] FIG. 6 illustrates the activation of PLA2R CAAR T cells by specific antibodies measured as GFP expression by Jurkat NFAT-GFP cells (6 h activation). No evidence of T cell activation is seen in construct 4025.C and untransduced cells upon exposure to the monoclonal anti-PLA2R Mab or the polyclonal anti-PLA2R Ab. Specific T cell activation in response to cross-linking by the monoclonal anti-PLA2R Ab but not the polyclonal anti-PLA2R Ab is seen using constructs 4027.CF12 and 4028.CF123.
Example 2
[0269] Six additional PLA2R CAAR constructs were designed and generated herein. Construct C was comprised of a cysteine rich domain, a CD8 hinge region, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2A, SEQ ID NOs: 25 & 26).
[0270] Construct CF1 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, and a C-type lectin 1 domain. Construct CF1 also included a CD8 hinge region, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2B, SEQ ID NOs: 27 & 28).
[0271] Construct CF123 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, a C-type lectin 1 domain, a C-type lectin 2 domain, and a C-type lectin 3 domain. Also included in construct CF123 was a GS linker, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2C, SEQ ID NOs: 29 & 30).
[0272] Construct CF1237 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, a C-type lectin 1 domain, a C-type lectin 2 domain, a C-type lectin 3 domain and a C-type lectin 7 domain. Additionally included in construct CF1237 was a GS linker, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 2D, SEQ ID NOs: 31 & 32).
[0273] Construct CF17 was comprised of an extracellular domain comprising a cysteine rich domain, a fibronectin II domain, a C-type lectin 1 domain, and a C-type lectin 7 domain. Construct CF17 also included a GS linker, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 7E, SEQ ID NOs: 33 & 34).
[0274] Construct C17 was comprised of an extracellular domain comprising a cysteine rich domain, a C-type lectin 1 domain, and a C-type lectin 7 domain. Construct C17 also included a GS linker, a CD8 transmembrane domain, a 4-1BB intracellular domain and CD3 zeta signaling domain (FIG. 7F, SEQ ID NOs: 35 & 36).
[0275] The various PLA2R CAAR constructs were transduced into Jurkat cells and primary human T cells. Surface expression of the PLA2R CAARs was assessed by flow cytometry using MN patient IgG, followed by APC-labeled anti-human IgG secondary antibody. All of the transduced Jurkat (FIG. 7A) and primary human T cells (FIG. 7B), but not the untransduced control cells, exhibited detectable surface expression of the CAAR constructs.
[0276] To evaluate whether plate-bound anti-PLA2R IgG, which mimics the surface of an anti-PLA2R B cell, can lead to CAAR T cell activation, 5.times.10.sup.4 Jurkat NFAT-GFP reporter cells were cultured for 6 h in 96-well plate coated with 10 .mu.g/ml either MN patient IgG or control human IgG. Cells were then harvested for flow cytometry analysis. MN patient IgG, but not control human IgG activated Jurkat cells transduced with each of the 10 constructs (FIG. 8). No evidence of T cell activation was seen in untransduced Jurkat cells upon exposure to either MN patient IgG or control human IgG (FIG. 8). These studies indicate that CAAR signaling is activated upon target encounter.
[0277] To evaluate whether primary human PLA2R CAAR T cells can be activated by plate-bound anti-PLA2R IgG, PLA2R CAAR T cells were exposed to either plate-bound MN patient IgG or control human IgG for 24 h. IFN-.gamma. secretion was measured by ELISA of culture supernatants. MN patient IgG stimulated markedly higher IFN-.gamma. production compared to the control human IgG, reflecting primary human CAAR T cell activation after target engagement (FIG. 9).
[0278] The disclosures of each and every patent, patent application, and publication cited herein are hereby incorporated herein by reference in their entirety. While this invention has been disclosed with reference to specific embodiments, it is apparent that other embodiments and variations of this invention may be devised by others skilled in the art without departing from the true spirit and scope of the invention. The appended claims are intended to be construed to include all such embodiments and equivalent variations.
Sequence CWU 1
1
721372DNAArtificial SequenceCysR (Ricin B type lectin) domain
1aagggcatct tcgtgatcca gagcgagagc ctgaagaagt gcatccaggc cggcaagagc
60gtgctgaccc tggaaaattg caagcaggcc aacaagcaca tgctgtggaa atgggtgtcc
120aaccacggcc tgttcaacat cggcggctct ggatgtctgg gcctgaattt
ctctgcccct 180gagcagcctc tgagcctgta cgagtgtgat agcaccctgg
tgtccctgag atggcggtgc 240aaccggaaga tgatcacagg ccctctgcag
tactctgtgc aggtcgccca cgacaatacc 300gtggtggcca gcagaaagta
catccacaag tggatcagct acggcagcgg cggaggcgac 360atctgtgaat ac
372233DNAArtificial SequenceLinker 2ctgcacaagg atctgcacac
catcaagggc aac 333147DNAArtificial SequenceFibronectin type-2
domain 3acccacggaa tgccctgcat gttcccgttt cagtacaacc accagtggca
ccacgagtgc 60accagagaag gcagagagga cgacctgctt tggtgcgcca caaccagcag
atacgagcgg 120gatgagaagt ggggcttctg ccctgat 147448DNAArtificial
SequenceLinker 4cctacctctg ccgaagtggg ctgcgatacc atctgggaga
aagacctg 485354DNAArtificial SequenceC-type lectin domain 1
5aacagccaca tctgctacca gttcaacctg ctgtccagcc tgtcttggag cgaggcccac
60agcagctgtc aaatgcaagg cggcacactg ctgagcatca ccgacgagac agaggaaaac
120ttcatccgcg agcacatgag cagcaagacc gtggaagtgt ggatgggact
gaaccagctg 180gatgagcatg ccggatggca gtggagtgat ggcacccctc
tgaactacct gaactggtcc 240cctgaagtga acttcgagcc cttcgtggaa
gatcactgcg gcaccttcag cagcttcatg 300cccagcgctt ggagaagcag
agactgcgag agcaccctgc cttacatctg caag 354687DNAArtificial
SequenceLinker 6aagtacctga accacatcga ccacgagatc gtggaaaagg
acgcctggaa gtactacgcc 60acacactgcg agcctggctg gaacccc
877354DNAArtificial SequenceC-type lectin domain 2 7tacaaccgga
actgctacaa gctgcagaaa gaggaaaaga cctggcacga ggccctgaga 60agctgccagg
ccgataatag cgccctgatc gacatcacaa gcctggccga ggtggaattt
120ctggtcactc tgctgggcga cgagaacgcc tctgagacat ggatcggcct
gtccagcaac 180aagatccccg tgtccttcga gtggtccaac gacagcagcg
tgatcttcac caactggcac 240accctggaac ctcacatctt ccccaacaga
tcccagctgt gtgtgtccgc cgagcagtct 300gaaggccact ggaaagtgaa
gaactgcgag gaacggctgt tctacatctg taaa 35481395DNAArtificial
SequenceExtracellular domains of Construct 4027.CF12 8aagggcatct
tcgtgatcca gagcgagagc ctgaagaagt gcatccaggc cggcaagagc 60gtgctgaccc
tggaaaattg caagcaggcc aacaagcaca tgctgtggaa atgggtgtcc
120aaccacggcc tgttcaacat cggcggctct ggatgtctgg gcctgaattt
ctctgcccct 180gagcagcctc tgagcctgta cgagtgtgat agcaccctgg
tgtccctgag atggcggtgc 240aaccggaaga tgatcacagg ccctctgcag
tactctgtgc aggtcgccca cgacaatacc 300gtggtggcca gcagaaagta
catccacaag tggatcagct acggcagcgg cggaggcgac 360atctgtgaat
acctgcacaa ggatctgcac accatcaagg gcaacaccca cggaatgccc
420tgcatgttcc cgtttcagta caaccaccag tggcaccacg agtgcaccag
agaaggcaga 480gaggacgacc tgctttggtg cgccacaacc agcagatacg
agcgggatga gaagtggggc 540ttctgccctg atcctacctc tgccgaagtg
ggctgcgata ccatctggga gaaagacctg 600aacagccaca tctgctacca
gttcaacctg ctgtccagcc tgtcttggag cgaggcccac 660agcagctgtc
aaatgcaagg cggcacactg ctgagcatca ccgacgagac agaggaaaac
720ttcatccgcg agcacatgag cagcaagacc gtggaagtgt ggatgggact
gaaccagctg 780gatgagcatg ccggatggca gtggagtgat ggcacccctc
tgaactacct gaactggtcc 840cctgaagtga acttcgagcc cttcgtggaa
gatcactgcg gcaccttcag cagcttcatg 900cccagcgctt ggagaagcag
agactgcgag agcaccctgc cttacatctg caagaagtac 960ctgaaccaca
tcgaccacga gatcgtggaa aaggacgcct ggaagtacta cgccacacac
1020tgcgagcctg gctggaaccc ctacaaccgg aactgctaca agctgcagaa
agaggaaaag 1080acctggcacg aggccctgag aagctgccag gccgataata
gcgccctgat cgacatcaca 1140agcctggccg aggtggaatt tctggtcact
ctgctgggcg acgagaacgc ctctgagaca 1200tggatcggcc tgtccagcaa
caagatcccc gtgtccttcg agtggtccaa cgacagcagc 1260gtgatcttca
ccaactggca caccctggaa cctcacatct tccccaacag atcccagctg
1320tgtgtgtccg ccgagcagtc tgaaggccac tggaaagtga agaactgcga
ggaacggctg 1380ttctacatct gtaaa 1395948DNAArtificial SequenceLinker
9aaggccggcc acgtgctgtc cgatgccgag agtggatgtc aatccgga
4810132DNAArtificial SequenceCD8 hinge domain 10accacgacgc
cagcgccgcg accaccaaca ccggcgccca ccatcgcgtc gcagcccctg 60tccctgcgcc
cagaggcgtg ccggccagcg gcggggggcg cagtgcacac gagggggctg
120gacttcgcct gt 132112232DNAArtificial SequencePLA2R-CAAR
Construct 4027.CF12 11atgctgctga gccctagcct gctgctgctc ctgcttcttg
gagcccctag aggatgtgcc 60ggatctgaag gtgttgccgc cgctctgaca cccgagagac
tgctggaatg gcaggacaag 120ggcatcttcg tgatccagag cgagagcctg
aagaagtgca tccaggccgg caagagcgtg 180ctgaccctgg aaaattgcaa
gcaggccaac aagcacatgc tgtggaaatg ggtgtccaac 240cacggcctgt
tcaacatcgg cggctctgga tgtctgggcc tgaatttctc tgcccctgag
300cagcctctga gcctgtacga gtgtgatagc accctggtgt ccctgagatg
gcggtgcaac 360cggaagatga tcacaggccc tctgcagtac tctgtgcagg
tcgcccacga caataccgtg 420gtggccagca gaaagtacat ccacaagtgg
atcagctacg gcagcggcgg aggcgacatc 480tgtgaatacc tgcacaagga
tctgcacacc atcaagggca acacccacgg aatgccctgc 540atgttcccgt
ttcagtacaa ccaccagtgg caccacgagt gcaccagaga aggcagagag
600gacgacctgc tttggtgcgc cacaaccagc agatacgagc gggatgagaa
gtggggcttc 660tgccctgatc ctacctctgc cgaagtgggc tgcgatacca
tctgggagaa agacctgaac 720agccacatct gctaccagtt caacctgctg
tccagcctgt cttggagcga ggcccacagc 780agctgtcaaa tgcaaggcgg
cacactgctg agcatcaccg acgagacaga ggaaaacttc 840atccgcgagc
acatgagcag caagaccgtg gaagtgtgga tgggactgaa ccagctggat
900gagcatgccg gatggcagtg gagtgatggc acccctctga actacctgaa
ctggtcccct 960gaagtgaact tcgagccctt cgtggaagat cactgcggca
ccttcagcag cttcatgccc 1020agcgcttgga gaagcagaga ctgcgagagc
accctgcctt acatctgcaa gaagtacctg 1080aaccacatcg accacgagat
cgtggaaaag gacgcctgga agtactacgc cacacactgc 1140gagcctggct
ggaaccccta caaccggaac tgctacaagc tgcagaaaga ggaaaagacc
1200tggcacgagg ccctgagaag ctgccaggcc gataatagcg ccctgatcga
catcacaagc 1260ctggccgagg tggaatttct ggtcactctg ctgggcgacg
agaacgcctc tgagacatgg 1320atcggcctgt ccagcaacaa gatccccgtg
tccttcgagt ggtccaacga cagcagcgtg 1380atcttcacca actggcacac
cctggaacct cacatcttcc ccaacagatc ccagctgtgt 1440gtgtccgccg
agcagtctga aggccactgg aaagtgaaga actgcgagga acggctgttc
1500tacatctgta aaaaggccgg ccacgtgctg tccgatgccg agagtggatg
tcaatccgga 1560accacgacgc cagcgccgcg accaccaaca ccggcgccca
ccatcgcgtc gcagcccctg 1620tccctgcgcc cagaggcgtg ccggccagcg
gcggggggcg cagtgcacac gagggggctg 1680gacttcgcct gtgatatcta
catctgggcg cccttggccg ggacttgtgg ggtccttctc 1740ctgtcactgg
ttatcaccct ttactgcaaa cggggcagaa agaaactcct gtatatattc
1800aaacaaccat ttatgagacc agtacaaact actcaagagg aagatggctg
tagctgccga 1860tttccagaag aagaagaagg aggatgtgaa ctgagagtga
agttcagcag gagcgcagac 1920gcccccgcgt acaagcaggg ccagaaccag
ctctataacg agctcaatct aggacgaaga 1980gaggagtacg atgttttgga
caagagacgt ggccgggacc ctgagatggg gggaaagccg 2040agaaggaaga
accctcagga aggcctgtac aatgaactgc agaaagataa gatggcggag
2100gcctacagtg agattgggat gaaaggcgag cgccggaggg gcaaggggca
cgatggcctt 2160taccagggtc tcagtacagc caccaaggac acctacgacg
cccttcacat gcaggccctg 2220ccccctcgct aa 223212743PRTArtificial
SequencePLA2R-CAAR Construct 4027.CF12 12Met Leu Leu Ser Pro Ser
Leu Leu Leu Leu Leu Leu Leu Gly Ala Pro1 5 10 15Arg Gly Cys Ala Gly
Ser Glu Gly Val Ala Ala Ala Leu Thr Pro Glu 20 25 30Arg Leu Leu Glu
Trp Gln Asp Lys Gly Ile Phe Val Ile Gln Ser Glu 35 40 45Ser Leu Lys
Lys Cys Ile Gln Ala Gly Lys Ser Val Leu Thr Leu Glu 50 55 60Asn Cys
Lys Gln Ala Asn Lys His Met Leu Trp Lys Trp Val Ser Asn65 70 75
80His Gly Leu Phe Asn Ile Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe
85 90 95Ser Ala Pro Glu Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr
Leu 100 105 110Val Ser Leu Arg Trp Arg Cys Asn Arg Lys Met Ile Thr
Gly Pro Leu 115 120 125Gln Tyr Ser Val Gln Val Ala His Asp Asn Thr
Val Val Ala Ser Arg 130 135 140Lys Tyr Ile His Lys Trp Ile Ser Tyr
Gly Ser Gly Gly Gly Asp Ile145 150 155 160Cys Glu Tyr Leu His Lys
Asp Leu His Thr Ile Lys Gly Asn Thr His 165 170 175Gly Met Pro Cys
Met Phe Pro Phe Gln Tyr Asn His Gln Trp His His 180 185 190Glu Cys
Thr Arg Glu Gly Arg Glu Asp Asp Leu Leu Trp Cys Ala Thr 195 200
205Thr Ser Arg Tyr Glu Arg Asp Glu Lys Trp Gly Phe Cys Pro Asp Pro
210 215 220Thr Ser Ala Glu Val Gly Cys Asp Thr Ile Trp Glu Lys Asp
Leu Asn225 230 235 240Ser His Ile Cys Tyr Gln Phe Asn Leu Leu Ser
Ser Leu Ser Trp Ser 245 250 255Glu Ala His Ser Ser Cys Gln Met Gln
Gly Gly Thr Leu Leu Ser Ile 260 265 270Thr Asp Glu Thr Glu Glu Asn
Phe Ile Arg Glu His Met Ser Ser Lys 275 280 285Thr Val Glu Val Trp
Met Gly Leu Asn Gln Leu Asp Glu His Ala Gly 290 295 300Trp Gln Trp
Ser Asp Gly Thr Pro Leu Asn Tyr Leu Asn Trp Ser Pro305 310 315
320Glu Val Asn Phe Glu Pro Phe Val Glu Asp His Cys Gly Thr Phe Ser
325 330 335Ser Phe Met Pro Ser Ala Trp Arg Ser Arg Asp Cys Glu Ser
Thr Leu 340 345 350Pro Tyr Ile Cys Lys Lys Tyr Leu Asn His Ile Asp
His Glu Ile Val 355 360 365Glu Lys Asp Ala Trp Lys Tyr Tyr Ala Thr
His Cys Glu Pro Gly Trp 370 375 380Asn Pro Tyr Asn Arg Asn Cys Tyr
Lys Leu Gln Lys Glu Glu Lys Thr385 390 395 400Trp His Glu Ala Leu
Arg Ser Cys Gln Ala Asp Asn Ser Ala Leu Ile 405 410 415Asp Ile Thr
Ser Leu Ala Glu Val Glu Phe Leu Val Thr Leu Leu Gly 420 425 430Asp
Glu Asn Ala Ser Glu Thr Trp Ile Gly Leu Ser Ser Asn Lys Ile 435 440
445Pro Val Ser Phe Glu Trp Ser Asn Asp Ser Ser Val Ile Phe Thr Asn
450 455 460Trp His Thr Leu Glu Pro His Ile Phe Pro Asn Arg Ser Gln
Leu Cys465 470 475 480Val Ser Ala Glu Gln Ser Glu Gly His Trp Lys
Val Lys Asn Cys Glu 485 490 495Glu Arg Leu Phe Tyr Ile Cys Lys Lys
Ala Gly His Val Leu Ser Asp 500 505 510Ala Glu Ser Gly Cys Gln Ser
Gly Thr Thr Thr Pro Ala Pro Arg Pro 515 520 525Pro Thr Pro Ala Pro
Thr Ile Ala Ser Gln Pro Leu Ser Leu Arg Pro 530 535 540Glu Ala Cys
Arg Pro Ala Ala Gly Gly Ala Val His Thr Arg Gly Leu545 550 555
560Asp Phe Ala Cys Asp Ile Tyr Ile Trp Ala Pro Leu Ala Gly Thr Cys
565 570 575Gly Val Leu Leu Leu Ser Leu Val Ile Thr Leu Tyr Cys Lys
Arg Gly 580 585 590Arg Lys Lys Leu Leu Tyr Ile Phe Lys Gln Pro Phe
Met Arg Pro Val 595 600 605Gln Thr Thr Gln Glu Glu Asp Gly Cys Ser
Cys Arg Phe Pro Glu Glu 610 615 620Glu Glu Gly Gly Cys Glu Leu Arg
Val Lys Phe Ser Arg Ser Ala Asp625 630 635 640Ala Pro Ala Tyr Lys
Gln Gly Gln Asn Gln Leu Tyr Asn Glu Leu Asn 645 650 655Leu Gly Arg
Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg Arg Gly Arg 660 665 670Asp
Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro Gln Glu Gly 675 680
685Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala Tyr Ser Glu
690 695 700Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly His Asp
Gly Leu705 710 715 720Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr
Tyr Asp Ala Leu His 725 730 735Met Gln Ala Leu Pro Pro Arg
7401357DNAArtificial SequenceLinker 13aaggccggcc acgtgctgtc
cgatgccgag agtggatgtc aagaaggctg ggagaga 5714366DNAArtificial
SequenceCC-type lectin domain 3 14cacggcggct tttgctacaa gatcgacacc
gtgctgcgga gcttcgatca ggccagcagc 60ggctactatt gccctcctgc tctggtcacc
atcaccaaca gattcgagca ggccttcatc 120accagcctga tcagcagcgt
cgtgaagatg aaggacagct acttctggat cgccctgcag 180gaccagaacg
acaccggcga gtacacatgg aagcccgtgg gacagaaacc cgagcctgtg
240cagtacaccc actggaacac acaccagcct agatactccg gcggctgcgt
ggcaatgaga 300ggcagacatc ctctcggcag atgggaagtg aagcactgtc
ggcacttcaa ggccatgtct 360ctgtgc 366151818DNAArtificial
SequenceExtracellular domains of PLA2R Construct 4028.CF123
15aagggcatct tcgtgatcca gagcgagagc ctgaagaagt gcatccaggc cggcaagagc
60gtgctgaccc tggaaaattg caagcaggcc aacaagcaca tgctgtggaa atgggtgtcc
120aaccacggcc tgttcaacat cggcggctct ggatgtctgg gcctgaattt
ctctgcccct 180gagcagcctc tgagcctgta cgagtgtgat agcaccctgg
tgtccctgag atggcggtgc 240aaccggaaga tgatcacagg ccctctgcag
tactctgtgc aggtcgccca cgacaatacc 300gtggtggcca gcagaaagta
catccacaag tggatcagct acggcagcgg cggaggcgac 360atctgtgaat
acctgcacaa ggatctgcac accatcaagg gcaacaccca cggaatgccc
420tgcatgttcc cgtttcagta caaccaccag tggcaccacg agtgcaccag
agaaggcaga 480gaggacgacc tgctttggtg cgccacaacc agcagatacg
agcgggatga gaagtggggc 540ttctgccctg atcctacctc tgccgaagtg
ggctgcgata ccatctggga gaaagacctg 600aacagccaca tctgctacca
gttcaacctg ctgtccagcc tgtcttggag cgaggcccac 660agcagctgtc
aaatgcaagg cggcacactg ctgagcatca ccgacgagac agaggaaaac
720ttcatccgcg agcacatgag cagcaagacc gtggaagtgt ggatgggact
gaaccagctg 780gatgagcatg ccggatggca gtggagtgat ggcacccctc
tgaactacct gaactggtcc 840cctgaagtga acttcgagcc cttcgtggaa
gatcactgcg gcaccttcag cagcttcatg 900cccagcgctt ggagaagcag
agactgcgag agcaccctgc cttacatctg caagaagtac 960ctgaaccaca
tcgaccacga gatcgtggaa aaggacgcct ggaagtacta cgccacacac
1020tgcgagcctg gctggaaccc ctacaaccgg aactgctaca agctgcagaa
agaggaaaag 1080acctggcacg aggccctgag aagctgccag gccgataata
gcgccctgat cgacatcaca 1140agcctggccg aggtggaatt tctggtcact
ctgctgggcg acgagaacgc ctctgagaca 1200tggatcggcc tgtccagcaa
caagatcccc gtgtccttcg agtggtccaa cgacagcagc 1260gtgatcttca
ccaactggca caccctggaa cctcacatct tccccaacag atcccagctg
1320tgtgtgtccg ccgagcagtc tgaaggccac tggaaagtga agaactgcga
ggaacggctg 1380ttctacatct gtaaaaaggc cggccacgtg ctgtccgatg
ccgagagtgg atgtcaagaa 1440ggctgggaga gacacggcgg cttttgctac
aagatcgaca ccgtgctgcg gagcttcgat 1500caggccagca gcggctacta
ttgccctcct gctctggtca ccatcaccaa cagattcgag 1560caggccttca
tcaccagcct gatcagcagc gtcgtgaaga tgaaggacag ctacttctgg
1620atcgccctgc aggaccagaa cgacaccggc gagtacacat ggaagcccgt
gggacagaaa 1680cccgagcctg tgcagtacac ccactggaac acacaccagc
ctagatactc cggcggctgc 1740gtggcaatga gaggcagaca tcctctcggc
agatgggaag tgaagcactg tcggcacttc 1800aaggccatgt ctctgtgc
18181675DNAArtificial SequenceLinker 16aagcagcccg tggaaaatca
agagaaggcc gagtacgagg aacgctggcc ttttcaccct 60tgctacctgt ccgga
75172682DNAArtificial SequencePLA2R-CAAR Construct 4028.CF123
17atgctgctga gccctagcct gctgctgctc ctgcttcttg gagcccctag aggatgtgcc
60ggatctgaag gtgttgccgc cgctctgaca cccgagagac tgctggaatg gcaggacaag
120ggcatcttcg tgatccagag cgagagcctg aagaagtgca tccaggccgg
caagagcgtg 180ctgaccctgg aaaattgcaa gcaggccaac aagcacatgc
tgtggaaatg ggtgtccaac 240cacggcctgt tcaacatcgg cggctctgga
tgtctgggcc tgaatttctc tgcccctgag 300cagcctctga gcctgtacga
gtgtgatagc accctggtgt ccctgagatg gcggtgcaac 360cggaagatga
tcacaggccc tctgcagtac tctgtgcagg tcgcccacga caataccgtg
420gtggccagca gaaagtacat ccacaagtgg atcagctacg gcagcggcgg
aggcgacatc 480tgtgaatacc tgcacaagga tctgcacacc atcaagggca
acacccacgg aatgccctgc 540atgttcccgt ttcagtacaa ccaccagtgg
caccacgagt gcaccagaga aggcagagag 600gacgacctgc tttggtgcgc
cacaaccagc agatacgagc gggatgagaa gtggggcttc 660tgccctgatc
ctacctctgc cgaagtgggc tgcgatacca tctgggagaa agacctgaac
720agccacatct gctaccagtt caacctgctg tccagcctgt cttggagcga
ggcccacagc 780agctgtcaaa tgcaaggcgg cacactgctg agcatcaccg
acgagacaga ggaaaacttc 840atccgcgagc acatgagcag caagaccgtg
gaagtgtgga tgggactgaa ccagctggat 900gagcatgccg gatggcagtg
gagtgatggc acccctctga actacctgaa ctggtcccct 960gaagtgaact
tcgagccctt cgtggaagat cactgcggca ccttcagcag cttcatgccc
1020agcgcttgga gaagcagaga ctgcgagagc accctgcctt acatctgcaa
gaagtacctg 1080aaccacatcg accacgagat cgtggaaaag gacgcctgga
agtactacgc cacacactgc 1140gagcctggct ggaaccccta caaccggaac
tgctacaagc tgcagaaaga ggaaaagacc 1200tggcacgagg ccctgagaag
ctgccaggcc gataatagcg ccctgatcga catcacaagc 1260ctggccgagg
tggaatttct ggtcactctg ctgggcgacg agaacgcctc tgagacatgg
1320atcggcctgt ccagcaacaa gatccccgtg tccttcgagt ggtccaacga
cagcagcgtg 1380atcttcacca actggcacac cctggaacct cacatcttcc
ccaacagatc ccagctgtgt 1440gtgtccgccg agcagtctga aggccactgg
aaagtgaaga actgcgagga acggctgttc 1500tacatctgta aaaaggccgg
ccacgtgctg tccgatgccg agagtggatg tcaagaaggc 1560tgggagagac
acggcggctt ttgctacaag atcgacaccg tgctgcggag cttcgatcag
1620gccagcagcg gctactattg ccctcctgct ctggtcacca tcaccaacag
attcgagcag 1680gccttcatca ccagcctgat cagcagcgtc gtgaagatga
aggacagcta cttctggatc 1740gccctgcagg accagaacga
caccggcgag tacacatgga agcccgtggg acagaaaccc 1800gagcctgtgc
agtacaccca ctggaacaca caccagccta gatactccgg cggctgcgtg
1860gcaatgagag gcagacatcc tctcggcaga tgggaagtga agcactgtcg
gcacttcaag 1920gccatgtctc tgtgcaagca gcccgtggaa aatcaagaga
aggccgagta cgaggaacgc 1980tggccttttc acccttgcta cctgtccgga
accacgacgc cagcgccgcg accaccaaca 2040ccggcgccca ccatcgcgtc
gcagcccctg tccctgcgcc cagaggcgtg ccggccagcg 2100gcggggggcg
cagtgcacac gagggggctg gacttcgcct gtgatatcta catctgggcg
2160cccttggccg ggacttgtgg ggtccttctc ctgtcactgg ttatcaccct
ttactgcaaa 2220cggggcagaa agaaactcct gtatatattc aaacaaccat
ttatgagacc agtacaaact 2280actcaagagg aagatggctg tagctgccga
tttccagaag aagaagaagg aggatgtgaa 2340ctgagagtga agttcagcag
gagcgcagac gcccccgcgt acaagcaggg ccagaaccag 2400ctctataacg
agctcaatct aggacgaaga gaggagtacg atgttttgga caagagacgt
2460ggccgggacc ctgagatggg gggaaagccg agaaggaaga accctcagga
aggcctgtac 2520aatgaactgc agaaagataa gatggcggag gcctacagtg
agattgggat gaaaggcgag 2580cgccggaggg gcaaggggca cgatggcctt
taccagggtc tcagtacagc caccaaggac 2640acctacgacg cccttcacat
gcaggccctg ccccctcgct aa 268218893PRTArtificial SequencePLA2R-CAAR
Construct 4028.CF123 18Met Leu Leu Ser Pro Ser Leu Leu Leu Leu Leu
Leu Leu Gly Ala Pro1 5 10 15Arg Gly Cys Ala Gly Ser Glu Gly Val Ala
Ala Ala Leu Thr Pro Glu 20 25 30Arg Leu Leu Glu Trp Gln Asp Lys Gly
Ile Phe Val Ile Gln Ser Glu 35 40 45Ser Leu Lys Lys Cys Ile Gln Ala
Gly Lys Ser Val Leu Thr Leu Glu 50 55 60Asn Cys Lys Gln Ala Asn Lys
His Met Leu Trp Lys Trp Val Ser Asn65 70 75 80His Gly Leu Phe Asn
Ile Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe 85 90 95Ser Ala Pro Glu
Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr Leu 100 105 110Val Ser
Leu Arg Trp Arg Cys Asn Arg Lys Met Ile Thr Gly Pro Leu 115 120
125Gln Tyr Ser Val Gln Val Ala His Asp Asn Thr Val Val Ala Ser Arg
130 135 140Lys Tyr Ile His Lys Trp Ile Ser Tyr Gly Ser Gly Gly Gly
Asp Ile145 150 155 160Cys Glu Tyr Leu His Lys Asp Leu His Thr Ile
Lys Gly Asn Thr His 165 170 175Gly Met Pro Cys Met Phe Pro Phe Gln
Tyr Asn His Gln Trp His His 180 185 190Glu Cys Thr Arg Glu Gly Arg
Glu Asp Asp Leu Leu Trp Cys Ala Thr 195 200 205Thr Ser Arg Tyr Glu
Arg Asp Glu Lys Trp Gly Phe Cys Pro Asp Pro 210 215 220Thr Ser Ala
Glu Val Gly Cys Asp Thr Ile Trp Glu Lys Asp Leu Asn225 230 235
240Ser His Ile Cys Tyr Gln Phe Asn Leu Leu Ser Ser Leu Ser Trp Ser
245 250 255Glu Ala His Ser Ser Cys Gln Met Gln Gly Gly Thr Leu Leu
Ser Ile 260 265 270Thr Asp Glu Thr Glu Glu Asn Phe Ile Arg Glu His
Met Ser Ser Lys 275 280 285Thr Val Glu Val Trp Met Gly Leu Asn Gln
Leu Asp Glu His Ala Gly 290 295 300Trp Gln Trp Ser Asp Gly Thr Pro
Leu Asn Tyr Leu Asn Trp Ser Pro305 310 315 320Glu Val Asn Phe Glu
Pro Phe Val Glu Asp His Cys Gly Thr Phe Ser 325 330 335Ser Phe Met
Pro Ser Ala Trp Arg Ser Arg Asp Cys Glu Ser Thr Leu 340 345 350Pro
Tyr Ile Cys Lys Lys Tyr Leu Asn His Ile Asp His Glu Ile Val 355 360
365Glu Lys Asp Ala Trp Lys Tyr Tyr Ala Thr His Cys Glu Pro Gly Trp
370 375 380Asn Pro Tyr Asn Arg Asn Cys Tyr Lys Leu Gln Lys Glu Glu
Lys Thr385 390 395 400Trp His Glu Ala Leu Arg Ser Cys Gln Ala Asp
Asn Ser Ala Leu Ile 405 410 415Asp Ile Thr Ser Leu Ala Glu Val Glu
Phe Leu Val Thr Leu Leu Gly 420 425 430Asp Glu Asn Ala Ser Glu Thr
Trp Ile Gly Leu Ser Ser Asn Lys Ile 435 440 445Pro Val Ser Phe Glu
Trp Ser Asn Asp Ser Ser Val Ile Phe Thr Asn 450 455 460Trp His Thr
Leu Glu Pro His Ile Phe Pro Asn Arg Ser Gln Leu Cys465 470 475
480Val Ser Ala Glu Gln Ser Glu Gly His Trp Lys Val Lys Asn Cys Glu
485 490 495Glu Arg Leu Phe Tyr Ile Cys Lys Lys Ala Gly His Val Leu
Ser Asp 500 505 510Ala Glu Ser Gly Cys Gln Glu Gly Trp Glu Arg His
Gly Gly Phe Cys 515 520 525Tyr Lys Ile Asp Thr Val Leu Arg Ser Phe
Asp Gln Ala Ser Ser Gly 530 535 540Tyr Tyr Cys Pro Pro Ala Leu Val
Thr Ile Thr Asn Arg Phe Glu Gln545 550 555 560Ala Phe Ile Thr Ser
Leu Ile Ser Ser Val Val Lys Met Lys Asp Ser 565 570 575Tyr Phe Trp
Ile Ala Leu Gln Asp Gln Asn Asp Thr Gly Glu Tyr Thr 580 585 590Trp
Lys Pro Val Gly Gln Lys Pro Glu Pro Val Gln Tyr Thr His Trp 595 600
605Asn Thr His Gln Pro Arg Tyr Ser Gly Gly Cys Val Ala Met Arg Gly
610 615 620Arg His Pro Leu Gly Arg Trp Glu Val Lys His Cys Arg His
Phe Lys625 630 635 640Ala Met Ser Leu Cys Lys Gln Pro Val Glu Asn
Gln Glu Lys Ala Glu 645 650 655Tyr Glu Glu Arg Trp Pro Phe His Pro
Cys Tyr Leu Ser Gly Thr Thr 660 665 670Thr Pro Ala Pro Arg Pro Pro
Thr Pro Ala Pro Thr Ile Ala Ser Gln 675 680 685Pro Leu Ser Leu Arg
Pro Glu Ala Cys Arg Pro Ala Ala Gly Gly Ala 690 695 700Val His Thr
Arg Gly Leu Asp Phe Ala Cys Asp Ile Tyr Ile Trp Ala705 710 715
720Pro Leu Ala Gly Thr Cys Gly Val Leu Leu Leu Ser Leu Val Ile Thr
725 730 735Leu Tyr Cys Lys Arg Gly Arg Lys Lys Leu Leu Tyr Ile Phe
Lys Gln 740 745 750Pro Phe Met Arg Pro Val Gln Thr Thr Gln Glu Glu
Asp Gly Cys Ser 755 760 765Cys Arg Phe Pro Glu Glu Glu Glu Gly Gly
Cys Glu Leu Arg Val Lys 770 775 780Phe Ser Arg Ser Ala Asp Ala Pro
Ala Tyr Lys Gln Gly Gln Asn Gln785 790 795 800Leu Tyr Asn Glu Leu
Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu 805 810 815Asp Lys Arg
Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg 820 825 830Lys
Asn Pro Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met 835 840
845Ala Glu Ala Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly
850 855 860Lys Gly His Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr
Lys Asp865 870 875 880Thr Tyr Asp Ala Leu His Met Gln Ala Leu Pro
Pro Arg 885 8901924PRTArtificial SequenceCD8 alpha transmembrane
domain 19Ile Tyr Ile Trp Ala Pro Leu Ala Gly Thr Cys Gly Val Leu
Leu Leu1 5 10 15Ser Leu Val Ile Thr Leu Tyr Cys 202072DNAArtificial
SequenceCD8 alpha transmembrane domain 20atctacatct gggcgccctt
ggccgggact tgtggggtcc ttctcctgtc actggttatc 60accctttact gc
722142PRTArtificial Sequence4-1BB 21Lys Arg Gly Arg Lys Lys Leu Leu
Tyr Ile Phe Lys Gln Pro Phe Met1 5 10 15Arg Pro Val Gln Thr Thr Gln
Glu Glu Asp Gly Cys Ser Cys Arg Phe 20 25 30Pro Glu Glu Glu Glu Gly
Gly Cys Glu Leu 35 4022126DNAArtificial Sequence4-1BB 22aaacggggca
gaaagaaact cctgtatata ttcaaacaac catttatgag accagtacaa 60actactcaag
aggaagatgg ctgtagctgc cgatttccag aagaagaaga aggaggatgt 120gaactg
12623112PRTArtificial SequenceCD3 zeta signaling domain 23Arg Val
Lys Phe Ser Arg Ser Ala Asp Ala Pro Ala Tyr Lys Gln Gly1 5 10 15Gln
Asn Gln Leu Tyr Asn Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr 20 25
30Asp Val Leu Asp Lys Arg Arg Gly Arg Asp Pro Glu Met Gly Gly Lys
35 40 45Pro Arg Arg Lys Asn Pro Gln Glu Gly Leu Tyr Asn Glu Leu Gln
Lys 50 55 60Asp Lys Met Ala Glu Ala Tyr Ser Glu Ile Gly Met Lys Gly
Glu Arg65 70 75 80Arg Arg Gly Lys Gly His Asp Gly Leu Tyr Gln Gly
Leu Ser Thr Ala 85 90 95Thr Lys Asp Thr Tyr Asp Ala Leu His Met Gln
Ala Leu Pro Pro Arg 100 105 11024336DNAArtificial SequenceCD3 zeta
signaling domain 24agagtgaagt tcagcaggag cgcagacgcc cccgcgtacc
agcagggcca gaaccagctc 60tataacgagc tcaatctagg acgaagagag gagtacgatg
ttttggacaa gagacgtggc 120cgggaccctg agatgggggg aaagccgaga
aggaagaacc ctcaggaagg cctgtacaat 180gaactgcaga aagataagat
ggcggaggcc tacagtgaga ttgggatgaa aggcgagcgc 240cggaggggca
aggggcacga tggcctttac cagggtctca gtacagccac caaggacacc
300tacgacgccc ttcacatgca ggccctgccc cctcgc 336251167DNAArtificial
SequencePLA2R-CAAR Construct C 25atggagtttg ggctgagctg gctttttctt
gtggctattt taaaaggtgt ccagtgcgga 60tccaaaggaa tatttgtcat tcagtcagag
tctttgaaaa agtgcataca ggctggaaaa 120agcgtgctta ccctggagaa
ctgcaagcaa gctaataagc atatgctttg gaaatgggtt 180agcaaccacg
gactctttaa tatcggaggc tccggctgtc tgggcctgaa cttcagtgca
240ccggagcaac cgctttctct gtacgaatgt gatagcacac ttgttagtct
tcggtggcgg 300tgtaaccgaa aaatgattac aggccctctg caatatagtg
ttcaagtggc ccacgacaat 360acagttgtgg cgtctagaaa atatattcac
aagtggattt cctacgggag cggcggaggg 420gatatatgtg aatatcttca
caaagacttg catacaatcg ctagcttcgt gccggtcttc 480ctgccagcga
agccaaccac gacgccagca ccgcgaccac caacacctgc gcccaccatc
540gcgtcgcagc ccctgtccct gcgcccagag gcgtgcagac cagcagcggg
gggcgcagtg 600cacacgaggg ggctggactt cgcctgtgat atctacatct
gggcgccctt ggccgggact 660tgtggggtcc ttctcctgtc actggttatc
accctttact gcaagcgcgg tcgcaagaaa 720ctgctctata tttttaaaca
gccattcatg agacctgtcc agaccactca agaggaggac 780ggatgttcct
gtagatttcc tgaagaggaa gagggggggt gcgagctgag agtgaagttc
840agcaggagcg cagacgcccc cgcgtaccag cagggccaga accagctcta
taacgagctc 900aatctaggac gaagagagga gtacgatgtt ttggacaaga
gacgtggccg ggaccctgag 960atggggggaa agccgagaag gaagaaccct
caggaaggcc tgtacaatga actgcagaaa 1020gataagatgg cggaggccta
cagtgagatt gggatgaaag gcgagcgccg gaggggcaag 1080gggcacgatg
gcctttacca gggtctcagt acagccacca aggacaccta cgacgccctt
1140cacatgcagg ccctgccccc tcgctaa 116726388PRTArtificial
SequencePLA2R-CAAR Construct C 26Met Glu Phe Gly Leu Ser Trp Leu
Phe Leu Val Ala Ile Leu Lys Gly1 5 10 15Val Gln Cys Gly Ser Lys Gly
Ile Phe Val Ile Gln Ser Glu Ser Leu 20 25 30Lys Lys Cys Ile Gln Ala
Gly Lys Ser Val Leu Thr Leu Glu Asn Cys 35 40 45Lys Gln Ala Asn Lys
His Met Leu Trp Lys Trp Val Ser Asn His Gly 50 55 60Leu Phe Asn Ile
Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala65 70 75 80Pro Glu
Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser 85 90 95Leu
Arg Trp Arg Cys Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr 100 105
110Ser Val Gln Val Ala His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr
115 120 125Ile His Lys Trp Ile Ser Tyr Gly Ser Gly Gly Gly Asp Ile
Cys Glu 130 135 140Tyr Leu His Lys Asp Leu His Thr Ile Ala Ser Phe
Val Pro Val Phe145 150 155 160Leu Pro Ala Lys Pro Thr Thr Thr Pro
Ala Pro Arg Pro Pro Thr Pro 165 170 175Ala Pro Thr Ile Ala Ser Gln
Pro Leu Ser Leu Arg Pro Glu Ala Cys 180 185 190Arg Pro Ala Ala Gly
Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala 195 200 205Cys Asp Ile
Tyr Ile Trp Ala Pro Leu Ala Gly Thr Cys Gly Val Leu 210 215 220Leu
Leu Ser Leu Val Ile Thr Leu Tyr Cys Lys Arg Gly Arg Lys Lys225 230
235 240Leu Leu Tyr Ile Phe Lys Gln Pro Phe Met Arg Pro Val Gln Thr
Thr 245 250 255Gln Glu Glu Asp Gly Cys Ser Cys Arg Phe Pro Glu Glu
Glu Glu Gly 260 265 270Gly Cys Glu Leu Arg Val Lys Phe Ser Arg Ser
Ala Asp Ala Pro Ala 275 280 285Tyr Gln Gln Gly Gln Asn Gln Leu Tyr
Asn Glu Leu Asn Leu Gly Arg 290 295 300Arg Glu Glu Tyr Asp Val Leu
Asp Lys Arg Arg Gly Arg Asp Pro Glu305 310 315 320Met Gly Gly Lys
Pro Arg Arg Lys Asn Pro Gln Glu Gly Leu Tyr Asn 325 330 335Glu Leu
Gln Lys Asp Lys Met Ala Glu Ala Tyr Ser Glu Ile Gly Met 340 345
350Lys Gly Glu Arg Arg Arg Gly Lys Gly His Asp Gly Leu Tyr Gln Gly
355 360 365Leu Ser Thr Ala Thr Lys Asp Thr Tyr Asp Ala Leu His Met
Gln Ala 370 375 380Leu Pro Pro Arg385271761DNAArtificial
SequencePLA2R-CAAR construct CF1 27atggagtttg ggctgagctg gctttttctt
gtggctattt taaaaggtgt ccagtgcgga 60tccaaaggga tctttgttat acaaagtgag
agcttgaaaa aatgtataca ggctggcaaa 120agtgtactga ctcttgaaaa
ttgcaaacaa gccaacaaac acatgctgtg gaaatgggtg 180tctaatcacg
gtctcttcaa tattggggga agtggatgcc tcggcctgaa tttctccgct
240cccgaacagc ccctctcact ttatgagtgt gattcaactc tggtgtcctt
gaggtggcga 300tgtaaccgca agatgataac cggccccctc cagtattccg
tccaagtagc acacgacaat 360accgtggtgg catctaggaa atacattcat
aagtggatat cttatggcag tggtggcggt 420gacatatgcg agtacctgca
caaggacctc cacacaataa aggggaacac gcacgggatg 480ccgtgtatgt
tcccgttcca atataatcat caatggcacc atgagtgtac gagagagggg
540cgagaagacg acctcctgtg gtgtgcgacc acctcaagat atgaacggga
tgagaagtgg 600ggcttttgcc ccgacccaac ctccgccgag gttggttgcg
acactatttg ggaaaaagat 660ttgaacagtc atatatgcta tcaatttaat
ttgttgagtt cactctcctg gagcgaagcg 720cacagctctt gtcagatgca
aggtggtaca ttgcttagca ttactgatga aactgaggag 780aatttcatta
gggagcatat gtcctcaaag acagtagagg tgtggatggg tctgaaccag
840ctcgacgaac acgccggttg gcagtggtca gatggaacgc ctctgaatta
tctcaactgg 900tcccctgagg tcaactttga accgtttgtg gaagatcatt
gtggtacttt ttccagtttt 960atgccaagcg cctggcgaag ccgagactgc
gagtctacgt tgccctatat ctgcaagaag 1020tatttgaatc acatagatca
tgaaattgtt gaagctagct tcgtgccggt cttcctgcca 1080gcgaagccaa
ccacgacgcc agcaccgcga ccaccaacac ctgcgcccac catcgcgtcg
1140cagcccctgt ccctgcgccc agaggcgtgc agaccagcag cggggggcgc
agtgcacacg 1200agggggctgg acttcgcctg tgatatctac atctgggcgc
ccttggccgg gacttgtggg 1260gtccttctcc tgtcactggt tatcaccctt
tactgcaagc gcggtcgcaa gaaactgctc 1320tatattttta aacagccatt
catgagacct gtccagacca ctcaagagga ggacggatgt 1380tcctgtagat
ttcctgaaga ggaagagggg gggtgcgagc tgagagtgaa gttcagcagg
1440agcgcagacg cccccgcgta ccagcagggc cagaaccagc tctataacga
gctcaatcta 1500ggacgaagag aggagtacga tgttttggac aagagacgtg
gccgggaccc tgagatgggg 1560ggaaagccga gaaggaagaa ccctcaggaa
ggcctgtaca atgaactgca gaaagataag 1620atggcggagg cctacagtga
gattgggatg aaaggcgagc gccggagggg caaggggcac 1680gatggccttt
accagggtct cagtacagcc accaaggaca cctacgacgc ccttcacatg
1740caggccctgc cccctcgcta a 176128586PRTArtificial
SequencePLA2R-CAAR construct CF1 28Met Glu Phe Gly Leu Ser Trp Leu
Phe Leu Val Ala Ile Leu Lys Gly1 5 10 15Val Gln Cys Gly Ser Lys Gly
Ile Phe Val Ile Gln Ser Glu Ser Leu 20 25 30Lys Lys Cys Ile Gln Ala
Gly Lys Ser Val Leu Thr Leu Glu Asn Cys 35 40 45Lys Gln Ala Asn Lys
His Met Leu Trp Lys Trp Val Ser Asn His Gly 50 55 60Leu Phe Asn Ile
Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala65 70 75 80Pro Glu
Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser 85 90 95Leu
Arg Trp Arg Cys Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr 100 105
110Ser Val Gln Val Ala His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr
115 120 125Ile His Lys Trp Ile Ser Tyr Gly Ser Gly Gly Gly Asp Ile
Cys Glu 130 135 140Tyr Leu His Lys Asp Leu His Thr Ile Lys Gly Asn
Thr His Gly Met145 150 155 160Pro Cys Met Phe Pro Phe Gln Tyr Asn
His Gln Trp His His Glu Cys 165 170 175Thr Arg Glu Gly Arg Glu Asp
Asp Leu Leu Trp Cys
Ala Thr Thr Ser 180 185 190Arg Tyr Glu Arg Asp Glu Lys Trp Gly Phe
Cys Pro Asp Pro Thr Ser 195 200 205Ala Glu Val Gly Cys Asp Thr Ile
Trp Glu Lys Asp Leu Asn Ser His 210 215 220Ile Cys Tyr Gln Phe Asn
Leu Leu Ser Ser Leu Ser Trp Ser Glu Ala225 230 235 240His Ser Ser
Cys Gln Met Gln Gly Gly Thr Leu Leu Ser Ile Thr Asp 245 250 255Glu
Thr Glu Glu Asn Phe Ile Arg Glu His Met Ser Ser Lys Thr Val 260 265
270Glu Val Trp Met Gly Leu Asn Gln Leu Asp Glu His Ala Gly Trp Gln
275 280 285Trp Ser Asp Gly Thr Pro Leu Asn Tyr Leu Asn Trp Ser Pro
Glu Val 290 295 300Asn Phe Glu Pro Phe Val Glu Asp His Cys Gly Thr
Phe Ser Ser Phe305 310 315 320Met Pro Ser Ala Trp Arg Ser Arg Asp
Cys Glu Ser Thr Leu Pro Tyr 325 330 335Ile Cys Lys Lys Tyr Leu Asn
His Ile Asp His Glu Ile Val Glu Ala 340 345 350Ser Phe Val Pro Val
Phe Leu Pro Ala Lys Pro Thr Thr Thr Pro Ala 355 360 365Pro Arg Pro
Pro Thr Pro Ala Pro Thr Ile Ala Ser Gln Pro Leu Ser 370 375 380Leu
Arg Pro Glu Ala Cys Arg Pro Ala Ala Gly Gly Ala Val His Thr385 390
395 400Arg Gly Leu Asp Phe Ala Cys Asp Ile Tyr Ile Trp Ala Pro Leu
Ala 405 410 415Gly Thr Cys Gly Val Leu Leu Leu Ser Leu Val Ile Thr
Leu Tyr Cys 420 425 430Lys Arg Gly Arg Lys Lys Leu Leu Tyr Ile Phe
Lys Gln Pro Phe Met 435 440 445Arg Pro Val Gln Thr Thr Gln Glu Glu
Asp Gly Cys Ser Cys Arg Phe 450 455 460Pro Glu Glu Glu Glu Gly Gly
Cys Glu Leu Arg Val Lys Phe Ser Arg465 470 475 480Ser Ala Asp Ala
Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 485 490 495Glu Leu
Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 500 505
510Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro
515 520 525Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala
Glu Ala 530 535 540Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg
Gly Lys Gly His545 550 555 560Asp Gly Leu Tyr Gln Gly Leu Ser Thr
Ala Thr Lys Asp Thr Tyr Asp 565 570 575Ala Leu His Met Gln Ala Leu
Pro Pro Arg 580 585292490DNAArtificial SequencePLA2R-CAAR Construct
CF123 29atggagtttg ggctgagctg gctttttctt gtggctattt taaaaggtgt
ccagtgcgga 60tccaaaggga ttttcgtgat acagtccgag agtctcaaaa agtgtatcca
ggcaggcaaa 120agtgttctca ctctggaaaa ctgcaaacaa gcgaacaagc
acatgttgtg gaagtgggtt 180agtaaccatg gactgttcaa catcggaggt
agtggatgcc ttggtctcaa tttctctgct 240ccggaacagc ctttgtcact
gtacgaatgc gactccactc tcgttagtct tagatggcga 300tgcaatcgca
aaatgattac gggaccactt caatattcag ttcaagtggc acatgataac
360accgtagtgg cctcacggaa atacatccat aaatggattt cttatggtag
cgggggcggc 420gatatatgtg aatacctcca taaggatctc cacaccatta
agggtaatac tcacggtatg 480ccgtgtatgt ttccttttca gtacaatcat
cagtggcatc atgaatgcac gagggaagga 540cgcgaggacg atttgctctg
gtgcgcaacc acctcacgct acgagagaga cgaaaaatgg 600ggcttttgcc
cggaccccac tagtgctgag gtaggatgtg atacgatttg ggaaaaggat
660ttgaattctc atatttgcta ccagtttaat cttctttcat ccctgtcctg
gtctgaggct 720cattctagtt gccagatgca aggtgggact ttgctttcaa
ttactgacga gactgaggaa 780aattttatcc gagagcatat gtcttctaaa
accgtagagg tatggatggg cctgaaccaa 840ttggacgaac acgcgggctg
gcagtggagc gacgggacac ctctcaacta ccttaattgg 900agccctgagg
taaactttga accgtttgtc gaggatcact gcggaacttt cagcagcttc
960atgcctagtg catggcggtc ccgagactgt gagagcaccc ttccatacat
atgtaaaaaa 1020tacctcaatc acatagacca cgagatcgta gagaaggatg
catggaaata ttatgctacg 1080cactgtgagc cgggatggaa tccttataac
cgcaactgtt acaagctgca aaaagaagag 1140aagacatggc atgaggcgct
gcgctcatgt caagcggaca attctgcact tatagatata 1200actagtttgg
cggaggtaga atttttggtt acgcttctcg gcgatgagaa tgcgtccgag
1260acgtggatag ggttgtcaag caataaaatt cctgtaagtt ttgaatggtc
aaatgactct 1320tctgtcatct tcaccaattg gcacacactc gaaccccata
tcttcccaaa ccgaagccag 1380ttgtgtgtca gcgctgagca atcagaagga
cattggaaag ttaaaaactg tgaagaaaga 1440ctgttctaca tctgtaagaa
ggcaggacat gtgctttcag atgcggaaag cggctgtcaa 1500gaaggttggg
agcgccatgg aggtttctgt tataaaatcg acacagtttt gcgatctttc
1560gatcaggctt caagcgggta ctattgtcct cctgcactgg ttacaatcac
gaaccggttt 1620gaacaggctt ttataacttc tttgatttcc agcgtggtta
aaatgaagga ctcttatttc 1680tggatagccc tgcaagacca aaatgatacc
ggtgagtaca catggaaacc ggtaggtcaa 1740aagccagagc cagtccagta
cactcattgg aatacccacc agcctaggta ctccggcggg 1800tgtgtggcga
tgcggggtcg ccaccctctc ggacgctggg aggtgaagca ttgccgccac
1860ttcaaggcga tgagcttgtg taaacagccc gtcgaaaatc aggaaaaggc
agctagcggt 1920ggcggaggtt ctggaggtgg aggttcctcc ggaatctaca
tctgggcgcc cttggccggg 1980acttgtgggg tccttctcct gtcactggtt
atcacccttt actgcaagcg cggtcgcaag 2040aaactgctct atatttttaa
acagccattc atgagacctg tccagaccac tcaagaggag 2100gacggatgtt
cctgtagatt tcctgaagag gaagaggggg ggtgcgagct gagagtgaag
2160ttcagcagga gcgcagacgc ccccgcgtac cagcagggcc agaaccagct
ctataacgag 2220ctcaatctag gacgaagaga ggagtacgat gttttggaca
agagacgtgg ccgggaccct 2280gagatggggg gaaagccgag aaggaagaac
cctcaggaag gcctgtacaa tgaactgcag 2340aaagataaga tggcggaggc
ctacagtgag attgggatga aaggcgagcg ccggaggggc 2400aaggggcacg
atggccttta ccagggtctc agtacagcca ccaaggacac ctacgacgcc
2460cttcacatgc aggccctgcc ccctcgctaa 249030829PRTArtificial
SequencePLA2R-CAAR Construct CF123 30Met Glu Phe Gly Leu Ser Trp
Leu Phe Leu Val Ala Ile Leu Lys Gly1 5 10 15Val Gln Cys Gly Ser Lys
Gly Ile Phe Val Ile Gln Ser Glu Ser Leu 20 25 30Lys Lys Cys Ile Gln
Ala Gly Lys Ser Val Leu Thr Leu Glu Asn Cys 35 40 45Lys Gln Ala Asn
Lys His Met Leu Trp Lys Trp Val Ser Asn His Gly 50 55 60Leu Phe Asn
Ile Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala65 70 75 80Pro
Glu Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser 85 90
95Leu Arg Trp Arg Cys Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr
100 105 110Ser Val Gln Val Ala His Asp Asn Thr Val Val Ala Ser Arg
Lys Tyr 115 120 125Ile His Lys Trp Ile Ser Tyr Gly Ser Gly Gly Gly
Asp Ile Cys Glu 130 135 140Tyr Leu His Lys Asp Leu His Thr Ile Lys
Gly Asn Thr His Gly Met145 150 155 160Pro Cys Met Phe Pro Phe Gln
Tyr Asn His Gln Trp His His Glu Cys 165 170 175Thr Arg Glu Gly Arg
Glu Asp Asp Leu Leu Trp Cys Ala Thr Thr Ser 180 185 190Arg Tyr Glu
Arg Asp Glu Lys Trp Gly Phe Cys Pro Asp Pro Thr Ser 195 200 205Ala
Glu Val Gly Cys Asp Thr Ile Trp Glu Lys Asp Leu Asn Ser His 210 215
220Ile Cys Tyr Gln Phe Asn Leu Leu Ser Ser Leu Ser Trp Ser Glu
Ala225 230 235 240His Ser Ser Cys Gln Met Gln Gly Gly Thr Leu Leu
Ser Ile Thr Asp 245 250 255Glu Thr Glu Glu Asn Phe Ile Arg Glu His
Met Ser Ser Lys Thr Val 260 265 270Glu Val Trp Met Gly Leu Asn Gln
Leu Asp Glu His Ala Gly Trp Gln 275 280 285Trp Ser Asp Gly Thr Pro
Leu Asn Tyr Leu Asn Trp Ser Pro Glu Val 290 295 300Asn Phe Glu Pro
Phe Val Glu Asp His Cys Gly Thr Phe Ser Ser Phe305 310 315 320Met
Pro Ser Ala Trp Arg Ser Arg Asp Cys Glu Ser Thr Leu Pro Tyr 325 330
335Ile Cys Lys Lys Tyr Leu Asn His Ile Asp His Glu Ile Val Glu Lys
340 345 350Asp Ala Trp Lys Tyr Tyr Ala Thr His Cys Glu Pro Gly Trp
Asn Pro 355 360 365Tyr Asn Arg Asn Cys Tyr Lys Leu Gln Lys Glu Glu
Lys Thr Trp His 370 375 380Glu Ala Leu Arg Ser Cys Gln Ala Asp Asn
Ser Ala Leu Ile Asp Ile385 390 395 400Thr Ser Leu Ala Glu Val Glu
Phe Leu Val Thr Leu Leu Gly Asp Glu 405 410 415Asn Ala Ser Glu Thr
Trp Ile Gly Leu Ser Ser Asn Lys Ile Pro Val 420 425 430Ser Phe Glu
Trp Ser Asn Asp Ser Ser Val Ile Phe Thr Asn Trp His 435 440 445Thr
Leu Glu Pro His Ile Phe Pro Asn Arg Ser Gln Leu Cys Val Ser 450 455
460Ala Glu Gln Ser Glu Gly His Trp Lys Val Lys Asn Cys Glu Glu
Arg465 470 475 480Leu Phe Tyr Ile Cys Lys Lys Ala Gly His Val Leu
Ser Asp Ala Glu 485 490 495Ser Gly Cys Gln Glu Gly Trp Glu Arg His
Gly Gly Phe Cys Tyr Lys 500 505 510Ile Asp Thr Val Leu Arg Ser Phe
Asp Gln Ala Ser Ser Gly Tyr Tyr 515 520 525Cys Pro Pro Ala Leu Val
Thr Ile Thr Asn Arg Phe Glu Gln Ala Phe 530 535 540Ile Thr Ser Leu
Ile Ser Ser Val Val Lys Met Lys Asp Ser Tyr Phe545 550 555 560Trp
Ile Ala Leu Gln Asp Gln Asn Asp Thr Gly Glu Tyr Thr Trp Lys 565 570
575Pro Val Gly Gln Lys Pro Glu Pro Val Gln Tyr Thr His Trp Asn Thr
580 585 590His Gln Pro Arg Tyr Ser Gly Gly Cys Val Ala Met Arg Gly
Arg His 595 600 605Pro Leu Gly Arg Trp Glu Val Lys His Cys Arg His
Phe Lys Ala Met 610 615 620Ser Leu Cys Lys Gln Pro Val Glu Asn Gln
Glu Lys Ala Ala Ser Gly625 630 635 640Gly Gly Gly Ser Gly Gly Gly
Gly Ser Ser Gly Ile Tyr Ile Trp Ala 645 650 655Pro Leu Ala Gly Thr
Cys Gly Val Leu Leu Leu Ser Leu Val Ile Thr 660 665 670Leu Tyr Cys
Lys Arg Gly Arg Lys Lys Leu Leu Tyr Ile Phe Lys Gln 675 680 685Pro
Phe Met Arg Pro Val Gln Thr Thr Gln Glu Glu Asp Gly Cys Ser 690 695
700Cys Arg Phe Pro Glu Glu Glu Glu Gly Gly Cys Glu Leu Arg Val
Lys705 710 715 720Phe Ser Arg Ser Ala Asp Ala Pro Ala Tyr Gln Gln
Gly Gln Asn Gln 725 730 735Leu Tyr Asn Glu Leu Asn Leu Gly Arg Arg
Glu Glu Tyr Asp Val Leu 740 745 750Asp Lys Arg Arg Gly Arg Asp Pro
Glu Met Gly Gly Lys Pro Arg Arg 755 760 765Lys Asn Pro Gln Glu Gly
Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met 770 775 780Ala Glu Ala Tyr
Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly785 790 795 800Lys
Gly His Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp 805 810
815Thr Tyr Asp Ala Leu His Met Gln Ala Leu Pro Pro Arg 820
825312910DNAArtificial SequencePLA2R-CAAR Construct CF1237
31atggagtttg ggctgagctg gctttttctt gtggctattt taaaaggtgt ccagtgcgga
60tccaaaggga ttttcgtgat acagtccgag agtctcaaaa agtgtatcca ggcaggcaaa
120agtgttctca ctctggaaaa ctgcaaacaa gcgaacaagc acatgttgtg
gaagtgggtt 180agtaaccatg gactgttcaa catcggaggt agtggatgcc
ttggtctcaa tttctctgct 240ccggaacagc ctttgtcact gtacgaatgc
gactccactc tcgttagtct tagatggcga 300tgcaatcgca aaatgattac
gggaccactt caatattcag ttcaagtggc acatgataac 360accgtagtgg
cctcacggaa atacatccat aaatggattt cttatggtag cgggggcggc
420gatatatgtg aatacctcca taaggatctc cacaccatta agggtaatac
tcacggtatg 480ccgtgtatgt ttccttttca gtacaatcat cagtggcatc
atgaatgcac gagggaagga 540cgcgaggacg atttgctctg gtgcgcaacc
acctcacgct acgagagaga cgaaaaatgg 600ggcttttgcc cggaccccac
tagtgctgag gtaggatgtg atacgatttg ggaaaaggat 660ttgaattctc
atatttgcta ccagtttaat cttctttcat ccctgtcctg gtctgaggct
720cattctagtt gccagatgca aggtgggact ttgctttcaa ttactgacga
gactgaggaa 780aattttatcc gagagcatat gtcttctaaa accgtagagg
tatggatggg cctgaaccaa 840ttggacgaac acgcgggctg gcagtggagc
gacgggacac ctctcaacta ccttaattgg 900agccctgagg taaactttga
accgtttgtc gaggatcact gcggaacttt cagcagcttc 960atgcctagtg
catggcggtc ccgagactgt gagagcaccc ttccatacat atgtaaaaaa
1020tacctcaatc acatagacca cgagatcgta gagaaggatg catggaaata
ttatgctacg 1080cactgtgagc cgggatggaa tccttataac cgcaactgtt
acaagctgca aaaagaagag 1140aagacatggc atgaggcgct gcgctcatgt
caagcggaca attctgcact tatagatata 1200actagtttgg cggaggtaga
atttttggtt acgcttctcg gcgatgagaa tgcgtccgag 1260acgtggatag
ggttgtcaag caataaaatt cctgtaagtt ttgaatggtc aaatgactct
1320tctgtcatct tcaccaattg gcacacactc gaaccccata tcttcccaaa
ccgaagccag 1380ttgtgtgtca gcgctgagca atcagaagga cattggaaag
ttaaaaactg tgaagaaaga 1440ctgttctaca tctgtaagaa ggcaggacat
gtgctttcag atgcggaaag cggctgtcaa 1500gaaggttggg agcgccatgg
aggtttctgt tataaaatcg acacagtttt gcgatctttc 1560gatcaggctt
caagcgggta ctattgtcct cctgcactgg ttacaatcac gaaccggttt
1620gaacaggctt ttataacttc tttgatttcc agcgtggtta aaatgaagga
ctcttatttc 1680tggatagccc tgcaagacca aaatgatacc ggtgagtaca
catggaaacc ggtaggtcaa 1740aagccagagc cagtccagta cactcattgg
aatacccacc agcctaggta ctccggcggg 1800tgtgtggcga tgcggggtcg
ccaccctctc ggacgctggg aggtgaagca ttgccgccac 1860ttcaaggcga
tgagcttgtg taaacagccc gtcgaaaatc aggaaaaggc agttaacaca
1920tctgatatgt accctatgcc taacacactc gaatatggga ataggacgta
caagattata 1980aacgcgaaca tgacgtggta tgctgcaatc aagacgtgcc
tcatgcacaa agctcagctt 2040gtgtctatta ctgaccaata ccaccaatca
tttttgacag tcgtgttgaa tcgattgggg 2100tacgcccatt ggatcggtct
cttcacgacg gacaatgggc tcaattttga ctggagtgac 2160ggtactaaat
catcctttac tttttggaag gatgaagaaa gttctctgtt gggcgattgc
2220gtgtttgctg actcaaatgg ccgatggcat tccacagcct gtgaaagttt
tctgcaggga 2280gctatttgcc acgtgcctcc cgaaacgcgg cagtccgaac
acccggaatt ggctagcggt 2340ggcggaggtt ctggaggtgg aggttcctcc
ggaatctaca tctgggcgcc cttggccggg 2400acttgtgggg tccttctcct
gtcactggtt atcacccttt actgcaagcg cggtcgcaag 2460aaactgctct
atatttttaa acagccattc atgagacctg tccagaccac tcaagaggag
2520gacggatgtt cctgtagatt tcctgaagag gaagaggggg ggtgcgagct
gagagtgaag 2580ttcagcagga gcgcagacgc ccccgcgtac cagcagggcc
agaaccagct ctataacgag 2640ctcaatctag gacgaagaga ggagtacgat
gttttggaca agagacgtgg ccgggaccct 2700gagatggggg gaaagccgag
aaggaagaac cctcaggaag gcctgtacaa tgaactgcag 2760aaagataaga
tggcggaggc ctacagtgag attgggatga aaggcgagcg ccggaggggc
2820aaggggcacg atggccttta ccagggtctc agtacagcca ccaaggacac
ctacgacgcc 2880cttcacatgc aggccctgcc ccctcgctaa
291032969PRTArtificial SequencePLA2R-CAAR Construct CF1237 32Met
Glu Phe Gly Leu Ser Trp Leu Phe Leu Val Ala Ile Leu Lys Gly1 5 10
15Val Gln Cys Gly Ser Lys Gly Ile Phe Val Ile Gln Ser Glu Ser Leu
20 25 30Lys Lys Cys Ile Gln Ala Gly Lys Ser Val Leu Thr Leu Glu Asn
Cys 35 40 45Lys Gln Ala Asn Lys His Met Leu Trp Lys Trp Val Ser Asn
His Gly 50 55 60Leu Phe Asn Ile Gly Gly Ser Gly Cys Leu Gly Leu Asn
Phe Ser Ala65 70 75 80Pro Glu Gln Pro Leu Ser Leu Tyr Glu Cys Asp
Ser Thr Leu Val Ser 85 90 95Leu Arg Trp Arg Cys Asn Arg Lys Met Ile
Thr Gly Pro Leu Gln Tyr 100 105 110Ser Val Gln Val Ala His Asp Asn
Thr Val Val Ala Ser Arg Lys Tyr 115 120 125Ile His Lys Trp Ile Ser
Tyr Gly Ser Gly Gly Gly Asp Ile Cys Glu 130 135 140Tyr Leu His Lys
Asp Leu His Thr Ile Lys Gly Asn Thr His Gly Met145 150 155 160Pro
Cys Met Phe Pro Phe Gln Tyr Asn His Gln Trp His His Glu Cys 165 170
175Thr Arg Glu Gly Arg Glu Asp Asp Leu Leu Trp Cys Ala Thr Thr Ser
180 185 190Arg Tyr Glu Arg Asp Glu Lys Trp Gly Phe Cys Pro Asp Pro
Thr Ser 195 200 205Ala Glu Val Gly Cys Asp Thr Ile Trp Glu Lys Asp
Leu Asn Ser His 210 215 220Ile Cys Tyr Gln Phe Asn Leu Leu Ser Ser
Leu Ser Trp Ser Glu Ala225 230 235 240His Ser Ser Cys Gln Met Gln
Gly Gly Thr Leu Leu Ser Ile Thr Asp 245 250 255Glu Thr Glu Glu Asn
Phe Ile Arg Glu His Met Ser Ser Lys Thr Val 260 265 270Glu Val Trp
Met Gly Leu Asn Gln Leu Asp Glu His Ala Gly Trp Gln 275 280 285Trp
Ser Asp Gly Thr Pro Leu Asn Tyr Leu Asn Trp Ser Pro Glu Val 290 295
300Asn Phe Glu Pro Phe Val Glu Asp His Cys Gly
Thr Phe Ser Ser Phe305 310 315 320Met Pro Ser Ala Trp Arg Ser Arg
Asp Cys Glu Ser Thr Leu Pro Tyr 325 330 335Ile Cys Lys Lys Tyr Leu
Asn His Ile Asp His Glu Ile Val Glu Lys 340 345 350Asp Ala Trp Lys
Tyr Tyr Ala Thr His Cys Glu Pro Gly Trp Asn Pro 355 360 365Tyr Asn
Arg Asn Cys Tyr Lys Leu Gln Lys Glu Glu Lys Thr Trp His 370 375
380Glu Ala Leu Arg Ser Cys Gln Ala Asp Asn Ser Ala Leu Ile Asp
Ile385 390 395 400Thr Ser Leu Ala Glu Val Glu Phe Leu Val Thr Leu
Leu Gly Asp Glu 405 410 415Asn Ala Ser Glu Thr Trp Ile Gly Leu Ser
Ser Asn Lys Ile Pro Val 420 425 430Ser Phe Glu Trp Ser Asn Asp Ser
Ser Val Ile Phe Thr Asn Trp His 435 440 445Thr Leu Glu Pro His Ile
Phe Pro Asn Arg Ser Gln Leu Cys Val Ser 450 455 460Ala Glu Gln Ser
Glu Gly His Trp Lys Val Lys Asn Cys Glu Glu Arg465 470 475 480Leu
Phe Tyr Ile Cys Lys Lys Ala Gly His Val Leu Ser Asp Ala Glu 485 490
495Ser Gly Cys Gln Glu Gly Trp Glu Arg His Gly Gly Phe Cys Tyr Lys
500 505 510Ile Asp Thr Val Leu Arg Ser Phe Asp Gln Ala Ser Ser Gly
Tyr Tyr 515 520 525Cys Pro Pro Ala Leu Val Thr Ile Thr Asn Arg Phe
Glu Gln Ala Phe 530 535 540Ile Thr Ser Leu Ile Ser Ser Val Val Lys
Met Lys Asp Ser Tyr Phe545 550 555 560Trp Ile Ala Leu Gln Asp Gln
Asn Asp Thr Gly Glu Tyr Thr Trp Lys 565 570 575Pro Val Gly Gln Lys
Pro Glu Pro Val Gln Tyr Thr His Trp Asn Thr 580 585 590His Gln Pro
Arg Tyr Ser Gly Gly Cys Val Ala Met Arg Gly Arg His 595 600 605Pro
Leu Gly Arg Trp Glu Val Lys His Cys Arg His Phe Lys Ala Met 610 615
620Ser Leu Cys Lys Gln Pro Val Glu Asn Gln Glu Lys Ala Val Asn
Thr625 630 635 640Ser Asp Met Tyr Pro Met Pro Asn Thr Leu Glu Tyr
Gly Asn Arg Thr 645 650 655Tyr Lys Ile Ile Asn Ala Asn Met Thr Trp
Tyr Ala Ala Ile Lys Thr 660 665 670Cys Leu Met His Lys Ala Gln Leu
Val Ser Ile Thr Asp Gln Tyr His 675 680 685Gln Ser Phe Leu Thr Val
Val Leu Asn Arg Leu Gly Tyr Ala His Trp 690 695 700Ile Gly Leu Phe
Thr Thr Asp Asn Gly Leu Asn Phe Asp Trp Ser Asp705 710 715 720Gly
Thr Lys Ser Ser Phe Thr Phe Trp Lys Asp Glu Glu Ser Ser Leu 725 730
735Leu Gly Asp Cys Val Phe Ala Asp Ser Asn Gly Arg Trp His Ser Thr
740 745 750Ala Cys Glu Ser Phe Leu Gln Gly Ala Ile Cys His Val Pro
Pro Glu 755 760 765Thr Arg Gln Ser Glu His Pro Glu Leu Ala Ser Gly
Gly Gly Gly Ser 770 775 780Gly Gly Gly Gly Ser Ser Gly Ile Tyr Ile
Trp Ala Pro Leu Ala Gly785 790 795 800Thr Cys Gly Val Leu Leu Leu
Ser Leu Val Ile Thr Leu Tyr Cys Lys 805 810 815Arg Gly Arg Lys Lys
Leu Leu Tyr Ile Phe Lys Gln Pro Phe Met Arg 820 825 830Pro Val Gln
Thr Thr Gln Glu Glu Asp Gly Cys Ser Cys Arg Phe Pro 835 840 845Glu
Glu Glu Glu Gly Gly Cys Glu Leu Arg Val Lys Phe Ser Arg Ser 850 855
860Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn
Glu865 870 875 880Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu
Asp Lys Arg Arg 885 890 895Gly Arg Asp Pro Glu Met Gly Gly Lys Pro
Arg Arg Lys Asn Pro Gln 900 905 910Glu Gly Leu Tyr Asn Glu Leu Gln
Lys Asp Lys Met Ala Glu Ala Tyr 915 920 925Ser Glu Ile Gly Met Lys
Gly Glu Arg Arg Arg Gly Lys Gly His Asp 930 935 940Gly Leu Tyr Gln
Gly Leu Ser Thr Ala Thr Lys Asp Thr Tyr Asp Ala945 950 955 960Leu
His Met Gln Ala Leu Pro Pro Arg 965332052DNAArtificial
SequencePLA2R-CAAR Construct CF17 33atggagtttg ggctgagctg
gctttttctt gtggctattt taaaaggtgt ccagtgcgga 60tccaaaggca tcttcgtaat
ccagtcagaa agtttgaaaa aatgtatcca agctggcaaa 120tcagtactta
cccttgagaa ctgcaagcaa gccaataaac atatgctgtg gaaatgggtc
180tcaaaccacg gcctcttcaa tattggtggg tcaggttgct tggggttgaa
tttctccgcc 240ccagagcaac cactcagcct ttacgagtgt gattccacac
ttgtctcttt gcgatggcgc 300tgcaatagga aaatgatcac aggccccctt
cagtactctg tgcaagttgc tcatgataac 360acagtcgtgg cgagtcggaa
atatattcac aaatggattt cttatgggag tggtggagga 420gatatatgcg
agtatttgca taaggacttg cacaccatca agggaaacac tcacggtatg
480ccatgtatgt ttccgttcca atataatcat caatggcacc acgaatgtac
ccgagaggga 540cgcgaggacg atcttctttg gtgcgccaca acctctcgat
atgaacgaga tgagaagtgg 600gggttttgtc ctgacccaac cagtgcagaa
gtagggtgcg ataccatctg ggagaaagac 660ttgaactcac acatatgcta
tcagtttaat ttgttgtctt ctttgtcatg gagcgaagct 720cattcatcat
gccagatgca gggcgggaca ctgctttcta tcaccgacga gactgaggaa
780aattttatcc gcgagcacat gtcaagcaag acagttgagg tttggatggg
gctcaatcaa 840ctggacgaac acgcagggtg gcagtggtcc gatggcactc
cgctcaacta ccttaactgg 900agcccagagg tgaactttga gccgtttgtc
gaagatcact gtggtacttt tagctccttc 960atgccgtccg catggagaag
tcgcgactgc gagtcaaccc tcccttacat ctgtaagaaa 1020tacctcaacc
acatagatca cgaaatcgta gaggtcaata cgtccgacat gtacccaatg
1080ccaaatacgt tggaatatgg gaataggaca tacaagataa ttaacgcaaa
tatgacgtgg 1140tatgccgcaa tcaaaacgtg cctcatgcac aaggcacagc
tcgtgtcaat tacggaccag 1200taccaccaat catttctcac agtcgttctt
aatcgattgg gttatgcaca ctggataggc 1260ttgttcacga cggacaatgg
tttgaacttt gactggtccg atggaactaa aagttctttc 1320actttttgga
aggatgagga gtcctccttg ctcggggact gcgtcttcgc agattcaaac
1380gggcgctggc actcaacggc atgtgagtcc ttcctgcagg gagctatatg
ccatgtgcca 1440ccagaaacac gccagtctga gcaccctgag ttggctagcg
gtggcggagg ttctggaggt 1500ggaggttcct ccggaatcta catctgggcg
cccttggccg ggacttgtgg ggtccttctc 1560ctgtcactgg ttatcaccct
ttactgcaag cgcggtcgca agaaactgct ctatattttt 1620aaacagccat
tcatgagacc tgtccagacc actcaagagg aggacggatg ttcctgtaga
1680tttcctgaag aggaagaggg ggggtgcgag ctgagagtga agttcagcag
gagcgcagac 1740gcccccgcgt accagcaggg ccagaaccag ctctataacg
agctcaatct aggacgaaga 1800gaggagtacg atgttttgga caagagacgt
ggccgggacc ctgagatggg gggaaagccg 1860agaaggaaga accctcagga
aggcctgtac aatgaactgc agaaagataa gatggcggag 1920gcctacagtg
agattgggat gaaaggcgag cgccggaggg gcaaggggca cgatggcctt
1980taccagggtc tcagtacagc caccaaggac acctacgacg cccttcacat
gcaggccctg 2040ccccctcgct aa 205234683PRTArtificial
SequencePLA2R-CAAR Construct CF17 34Met Glu Phe Gly Leu Ser Trp Leu
Phe Leu Val Ala Ile Leu Lys Gly1 5 10 15Val Gln Cys Gly Ser Lys Gly
Ile Phe Val Ile Gln Ser Glu Ser Leu 20 25 30Lys Lys Cys Ile Gln Ala
Gly Lys Ser Val Leu Thr Leu Glu Asn Cys 35 40 45Lys Gln Ala Asn Lys
His Met Leu Trp Lys Trp Val Ser Asn His Gly 50 55 60Leu Phe Asn Ile
Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala65 70 75 80Pro Glu
Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser 85 90 95Leu
Arg Trp Arg Cys Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr 100 105
110Ser Val Gln Val Ala His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr
115 120 125Ile His Lys Trp Ile Ser Tyr Gly Ser Gly Gly Gly Asp Ile
Cys Glu 130 135 140Tyr Leu His Lys Asp Leu His Thr Ile Lys Gly Asn
Thr His Gly Met145 150 155 160Pro Cys Met Phe Pro Phe Gln Tyr Asn
His Gln Trp His His Glu Cys 165 170 175Thr Arg Glu Gly Arg Glu Asp
Asp Leu Leu Trp Cys Ala Thr Thr Ser 180 185 190Arg Tyr Glu Arg Asp
Glu Lys Trp Gly Phe Cys Pro Asp Pro Thr Ser 195 200 205Ala Glu Val
Gly Cys Asp Thr Ile Trp Glu Lys Asp Leu Asn Ser His 210 215 220Ile
Cys Tyr Gln Phe Asn Leu Leu Ser Ser Leu Ser Trp Ser Glu Ala225 230
235 240His Ser Ser Cys Gln Met Gln Gly Gly Thr Leu Leu Ser Ile Thr
Asp 245 250 255Glu Thr Glu Glu Asn Phe Ile Arg Glu His Met Ser Ser
Lys Thr Val 260 265 270Glu Val Trp Met Gly Leu Asn Gln Leu Asp Glu
His Ala Gly Trp Gln 275 280 285Trp Ser Asp Gly Thr Pro Leu Asn Tyr
Leu Asn Trp Ser Pro Glu Val 290 295 300Asn Phe Glu Pro Phe Val Glu
Asp His Cys Gly Thr Phe Ser Ser Phe305 310 315 320Met Pro Ser Ala
Trp Arg Ser Arg Asp Cys Glu Ser Thr Leu Pro Tyr 325 330 335Ile Cys
Lys Lys Tyr Leu Asn His Ile Asp His Glu Ile Val Glu Val 340 345
350Asn Thr Ser Asp Met Tyr Pro Met Pro Asn Thr Leu Glu Tyr Gly Asn
355 360 365Arg Thr Tyr Lys Ile Ile Asn Ala Asn Met Thr Trp Tyr Ala
Ala Ile 370 375 380Lys Thr Cys Leu Met His Lys Ala Gln Leu Val Ser
Ile Thr Asp Gln385 390 395 400Tyr His Gln Ser Phe Leu Thr Val Val
Leu Asn Arg Leu Gly Tyr Ala 405 410 415His Trp Ile Gly Leu Phe Thr
Thr Asp Asn Gly Leu Asn Phe Asp Trp 420 425 430Ser Asp Gly Thr Lys
Ser Ser Phe Thr Phe Trp Lys Asp Glu Glu Ser 435 440 445Ser Leu Leu
Gly Asp Cys Val Phe Ala Asp Ser Asn Gly Arg Trp His 450 455 460Ser
Thr Ala Cys Glu Ser Phe Leu Gln Gly Ala Ile Cys His Val Pro465 470
475 480Pro Glu Thr Arg Gln Ser Glu His Pro Glu Leu Ala Ser Gly Gly
Gly 485 490 495Gly Ser Gly Gly Gly Gly Ser Ser Gly Ile Tyr Ile Trp
Ala Pro Leu 500 505 510Ala Gly Thr Cys Gly Val Leu Leu Leu Ser Leu
Val Ile Thr Leu Tyr 515 520 525Cys Lys Arg Gly Arg Lys Lys Leu Leu
Tyr Ile Phe Lys Gln Pro Phe 530 535 540Met Arg Pro Val Gln Thr Thr
Gln Glu Glu Asp Gly Cys Ser Cys Arg545 550 555 560Phe Pro Glu Glu
Glu Glu Gly Gly Cys Glu Leu Arg Val Lys Phe Ser 565 570 575Arg Ser
Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr 580 585
590Asn Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys
595 600 605Arg Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg
Lys Asn 610 615 620Pro Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp
Lys Met Ala Glu625 630 635 640Ala Tyr Ser Glu Ile Gly Met Lys Gly
Glu Arg Arg Arg Gly Lys Gly 645 650 655His Asp Gly Leu Tyr Gln Gly
Leu Ser Thr Ala Thr Lys Asp Thr Tyr 660 665 670Asp Ala Leu His Met
Gln Ala Leu Pro Pro Arg 675 680351893DNAArtificial
SequencePLA2R-CAAR Construct C17 35atggagtttg ggctgagctg gctttttctt
gtggctattt taaaaggtgt ccagtgcgga 60tccaaaggaa tcttcgtaat tcaatctgag
agtctgaaaa aatgtattca ggccggtaag 120agcgtactca cgcttgaaaa
ttgcaaacag gccaacaaac acatgctttg gaaatgggtt 180tcaaatcacg
ggttgtttaa cataggggga tcaggatgtc tgggccttaa cttttccgca
240cctgaacaac ctcttagtct gtatgagtgt gactcaacgc tggtctcctt
gcgctggaga 300tgcaatcgga agatgataac cgggcccctc cagtattccg
ttcaggtcgc ccacgataat 360actgttgttg catcccgaaa atatattcat
aagtggatct cctacgggag tggagggggc 420gatatttgtg aatacctcca
caaggatctg cacactatca cttctgcgga agtaggctgt 480gacacaatct
gggagaaaga tctgaattca cacatttgct atcagttcaa tcttctgagt
540tctttgagct ggtccgaagc acattcatcc tgtcagatgc aaggtggaac
actcttgtca 600ataacagatg aaacggaaga gaactttatt agagaacata
tgtcctcaaa gactgtggag 660gtgtggatgg gacttaacca gctcgatgaa
catgcaggat ggcagtggag tgacggaacg 720ccactgaact acctgaattg
gagcccagag gtgaatttcg agcctttcgt agaggaccat 780tgcggtactt
tttcatcttt tatgcccagc gcatggagat cccgagattg tgaaagcacg
840ctgccctata tttgtaaaaa gtacctgaac cacatagatc atgagatagt
tgaggtaaat 900acaagtgata tgtaccccat gccgaacaca ctcgagtacg
gaaatagaac ctacaagata 960atcaacgcta acatgacctg gtacgcggcc
attaagacct gcctcatgca caaggctcaa 1020ctcgtcagta ttactgacca
atatcaccag tcatttctca ccgtcgtgtt gaatcgcctc 1080ggttacgccc
actggatcgg tttgtttaca acggacaatg gactcaattt cgattggtca
1140gacggaacca aatctagttt taccttctgg aaagacgagg aatcaagcct
gcttggggac 1200tgcgtatttg cggactctaa tggccgatgg catagtacag
cgtgtgagag ctttttgcag 1260ggggcgattt gtcatgttcc gccggaaacc
cgccaaagcg agcatccaga attggctagc 1320ggtggcggag gttctggagg
tggaggttcc tccggaatct acatctgggc gcccttggcc 1380gggacttgtg
gggtccttct cctgtcactg gttatcaccc tttactgcaa gcgcggtcgc
1440aagaaactgc tctatatttt taaacagcca ttcatgagac ctgtccagac
cactcaagag 1500gaggacggat gttcctgtag atttcctgaa gaggaagagg
gggggtgcga gctgagagtg 1560aagttcagca ggagcgcaga cgcccccgcg
taccagcagg gccagaacca gctctataac 1620gagctcaatc taggacgaag
agaggagtac gatgttttgg acaagagacg tggccgggac 1680cctgagatgg
ggggaaagcc gagaaggaag aaccctcagg aaggcctgta caatgaactg
1740cagaaagata agatggcgga ggcctacagt gagattggga tgaaaggcga
gcgccggagg 1800ggcaaggggc acgatggcct ttaccagggt ctcagtacag
ccaccaagga cacctacgac 1860gcccttcaca tgcaggccct gccccctcgc taa
189336630PRTArtificial SequencePLA2R-CAAR Construct C17 36Met Glu
Phe Gly Leu Ser Trp Leu Phe Leu Val Ala Ile Leu Lys Gly1 5 10 15Val
Gln Cys Gly Ser Lys Gly Ile Phe Val Ile Gln Ser Glu Ser Leu 20 25
30Lys Lys Cys Ile Gln Ala Gly Lys Ser Val Leu Thr Leu Glu Asn Cys
35 40 45Lys Gln Ala Asn Lys His Met Leu Trp Lys Trp Val Ser Asn His
Gly 50 55 60Leu Phe Asn Ile Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe
Ser Ala65 70 75 80Pro Glu Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser
Thr Leu Val Ser 85 90 95Leu Arg Trp Arg Cys Asn Arg Lys Met Ile Thr
Gly Pro Leu Gln Tyr 100 105 110Ser Val Gln Val Ala His Asp Asn Thr
Val Val Ala Ser Arg Lys Tyr 115 120 125Ile His Lys Trp Ile Ser Tyr
Gly Ser Gly Gly Gly Asp Ile Cys Glu 130 135 140Tyr Leu His Lys Asp
Leu His Thr Ile Thr Ser Ala Glu Val Gly Cys145 150 155 160Asp Thr
Ile Trp Glu Lys Asp Leu Asn Ser His Ile Cys Tyr Gln Phe 165 170
175Asn Leu Leu Ser Ser Leu Ser Trp Ser Glu Ala His Ser Ser Cys Gln
180 185 190Met Gln Gly Gly Thr Leu Leu Ser Ile Thr Asp Glu Thr Glu
Glu Asn 195 200 205Phe Ile Arg Glu His Met Ser Ser Lys Thr Val Glu
Val Trp Met Gly 210 215 220Leu Asn Gln Leu Asp Glu His Ala Gly Trp
Gln Trp Ser Asp Gly Thr225 230 235 240Pro Leu Asn Tyr Leu Asn Trp
Ser Pro Glu Val Asn Phe Glu Pro Phe 245 250 255Val Glu Asp His Cys
Gly Thr Phe Ser Ser Phe Met Pro Ser Ala Trp 260 265 270Arg Ser Arg
Asp Cys Glu Ser Thr Leu Pro Tyr Ile Cys Lys Lys Tyr 275 280 285Leu
Asn His Ile Asp His Glu Ile Val Glu Val Asn Thr Ser Asp Met 290 295
300Tyr Pro Met Pro Asn Thr Leu Glu Tyr Gly Asn Arg Thr Tyr Lys
Ile305 310 315 320Ile Asn Ala Asn Met Thr Trp Tyr Ala Ala Ile Lys
Thr Cys Leu Met 325 330 335His Lys Ala Gln Leu Val Ser Ile Thr Asp
Gln Tyr His Gln Ser Phe 340 345 350Leu Thr Val Val Leu Asn Arg Leu
Gly Tyr Ala His Trp Ile Gly Leu 355 360 365Phe Thr Thr Asp Asn Gly
Leu Asn Phe Asp Trp Ser Asp Gly Thr Lys 370 375 380Ser Ser Phe Thr
Phe Trp Lys Asp Glu Glu Ser Ser Leu Leu Gly Asp385 390 395 400Cys
Val Phe Ala Asp Ser Asn Gly Arg Trp His Ser Thr Ala Cys Glu 405 410
415Ser Phe Leu Gln Gly Ala Ile Cys His Val Pro Pro Glu Thr Arg Gln
420 425 430Ser Glu His Pro Glu Leu Ala Ser Gly Gly
Gly Gly Ser Gly Gly Gly 435 440 445Gly Ser Ser Gly Ile Tyr Ile Trp
Ala Pro Leu Ala Gly Thr Cys Gly 450 455 460Val Leu Leu Leu Ser Leu
Val Ile Thr Leu Tyr Cys Lys Arg Gly Arg465 470 475 480Lys Lys Leu
Leu Tyr Ile Phe Lys Gln Pro Phe Met Arg Pro Val Gln 485 490 495Thr
Thr Gln Glu Glu Asp Gly Cys Ser Cys Arg Phe Pro Glu Glu Glu 500 505
510Glu Gly Gly Cys Glu Leu Arg Val Lys Phe Ser Arg Ser Ala Asp Ala
515 520 525Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn Glu Leu
Asn Leu 530 535 540Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg
Arg Gly Arg Asp545 550 555 560Pro Glu Met Gly Gly Lys Pro Arg Arg
Lys Asn Pro Gln Glu Gly Leu 565 570 575Tyr Asn Glu Leu Gln Lys Asp
Lys Met Ala Glu Ala Tyr Ser Glu Ile 580 585 590Gly Met Lys Gly Glu
Arg Arg Arg Gly Lys Gly His Asp Gly Leu Tyr 595 600 605Gln Gly Leu
Ser Thr Ala Thr Lys Asp Thr Tyr Asp Ala Leu His Met 610 615 620Gln
Ala Leu Pro Pro Arg625 630371182DNAArtificial SequencePLA2R-CAAR
Construct 4025.C 37atgctgctga gccctagcct gctgctgctc ctgcttcttg
gagcccctag aggatgtgcc 60ggatctgaag gtgttgccgc cgctctgaca cccgagagac
tgctggaatg gcaggacaag 120ggcatcttcg tgatccagag cgagagcctg
aagaagtgca tccaggccgg caagagcgtg 180ctgaccctgg aaaattgcaa
gcaggccaac aagcacatgc tgtggaaatg ggtgtccaac 240cacggcctgt
tcaacatcgg cggctctgga tgtctgggcc tgaatttctc tgcccctgag
300cagcctctga gcctgtacga gtgtgatagc accctggtgt ccctgagatg
gcggtgcaac 360cggaagatga tcacaggccc tctgcagtac tctgtgcagg
tcgcccacga caataccgtg 420gtggccagca gaaagtacat ccacaagtgg
atcagctacg gcagcggcgg aggcgacatc 480tgtgaatacc tgcacaagga
tctgtccgga accacgacgc cagcgccgcg accaccaaca 540ccggcgccca
ccatcgcgtc gcagcccctg tccctgcgcc cagaggcgtg ccggccagcg
600gcggggggcg cagtgcacac gagggggctg gacttcgcct gtgatatcta
catctgggcg 660cccttggccg ggacttgtgg ggtccttctc ctgtcactgg
ttatcaccct ttactgcaaa 720cggggcagaa agaaactcct gtatatattc
aaacaaccat ttatgagacc agtacaaact 780actcaagagg aagatggctg
tagctgccga tttccagaag aagaagaagg aggatgtgaa 840ctgagagtga
agttcagcag gagcgcagac gcccccgcgt acaagcaggg ccagaaccag
900ctctataacg agctcaatct aggacgaaga gaggagtacg atgttttgga
caagagacgt 960ggccgggacc ctgagatggg gggaaagccg agaaggaaga
accctcagga aggcctgtac 1020aatgaactgc agaaagataa gatggcggag
gcctacagtg agattgggat gaaaggcgag 1080cgccggaggg gcaaggggca
cgatggcctt taccagggtc tcagtacagc caccaaggac 1140acctacgacg
cccttcacat gcaggccctg ccccctcgct aa 118238393PRTArtificial
SequencePLA2R-CAAR Construct 4025.C 38Met Leu Leu Ser Pro Ser Leu
Leu Leu Leu Leu Leu Leu Gly Ala Pro1 5 10 15Arg Gly Cys Ala Gly Ser
Glu Gly Val Ala Ala Ala Leu Thr Pro Glu 20 25 30Arg Leu Leu Glu Trp
Gln Asp Lys Gly Ile Phe Val Ile Gln Ser Glu 35 40 45Ser Leu Lys Lys
Cys Ile Gln Ala Gly Lys Ser Val Leu Thr Leu Glu 50 55 60Asn Cys Lys
Gln Ala Asn Lys His Met Leu Trp Lys Trp Val Ser Asn65 70 75 80His
Gly Leu Phe Asn Ile Gly Gly Ser Gly Cys Leu Gly Leu Asn Phe 85 90
95Ser Ala Pro Glu Gln Pro Leu Ser Leu Tyr Glu Cys Asp Ser Thr Leu
100 105 110Val Ser Leu Arg Trp Arg Cys Asn Arg Lys Met Ile Thr Gly
Pro Leu 115 120 125Gln Tyr Ser Val Gln Val Ala His Asp Asn Thr Val
Val Ala Ser Arg 130 135 140Lys Tyr Ile His Lys Trp Ile Ser Tyr Gly
Ser Gly Gly Gly Asp Ile145 150 155 160Cys Glu Tyr Leu His Lys Asp
Leu Ser Gly Thr Thr Thr Pro Ala Pro 165 170 175Arg Pro Pro Thr Pro
Ala Pro Thr Ile Ala Ser Gln Pro Leu Ser Leu 180 185 190Arg Pro Glu
Ala Cys Arg Pro Ala Ala Gly Gly Ala Val His Thr Arg 195 200 205Gly
Leu Asp Phe Ala Cys Asp Ile Tyr Ile Trp Ala Pro Leu Ala Gly 210 215
220Thr Cys Gly Val Leu Leu Leu Ser Leu Val Ile Thr Leu Tyr Cys
Lys225 230 235 240Arg Gly Arg Lys Lys Leu Leu Tyr Ile Phe Lys Gln
Pro Phe Met Arg 245 250 255Pro Val Gln Thr Thr Gln Glu Glu Asp Gly
Cys Ser Cys Arg Phe Pro 260 265 270Glu Glu Glu Glu Gly Gly Cys Glu
Leu Arg Val Lys Phe Ser Arg Ser 275 280 285Ala Asp Ala Pro Ala Tyr
Lys Gln Gly Gln Asn Gln Leu Tyr Asn Glu 290 295 300Leu Asn Leu Gly
Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg Arg305 310 315 320Gly
Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro Gln 325 330
335Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala Tyr
340 345 350Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly
His Asp 355 360 365Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp
Thr Tyr Asp Ala 370 375 380Leu His Met Gln Ala Leu Pro Pro Arg385
390391821DNAArtificial SequencePLA2R-CAAR Construct 4026.CF1
39atgctgctga gccctagcct gctgctgctc ctgcttcttg gagcccctag aggatgtgcc
60ggatctgaag gtgttgccgc cgctctgaca cccgagagac tgctggaatg gcaggacaag
120ggcatcttcg tgatccagag cgagagcctg aagaagtgca tccaggccgg
caagagcgtg 180ctgaccctgg aaaattgcaa gcaggccaac aagcacatgc
tgtggaaatg ggtgtccaac 240cacggcctgt tcaacatcgg cggctctgga
tgtctgggcc tgaatttctc tgcccctgag 300cagcctctga gcctgtacga
gtgtgatagc accctggtgt ccctgagatg gcggtgcaac 360cggaagatga
tcacaggccc tctgcagtac tctgtgcagg tcgcccacga caataccgtg
420gtggccagca gaaagtacat ccacaagtgg atcagctacg gcagcggcgg
aggcgacatc 480tgtgaatacc tgcacaagga tctgcacacc atcaagggca
acacccacgg aatgccctgc 540atgttcccgt ttcagtacaa ccaccagtgg
caccacgagt gcaccagaga aggcagagag 600gacgacctgc tttggtgcgc
cacaaccagc agatacgagc gggatgagaa gtggggcttc 660tgccctgatc
ctacctctgc cgaagtgggc tgcgatacca tctgggagaa agacctgaac
720agccacatct gctaccagtt caacctgctg tccagcctgt cttggagcga
ggcccacagc 780agctgtcaaa tgcaaggcgg cacactgctg agcatcaccg
acgagacaga ggaaaacttc 840atccgcgagc acatgagcag caagaccgtg
gaagtgtgga tgggactgaa ccagctggat 900gagcatgccg gatggcagtg
gagtgatggc acccctctga actacctgaa ctggtcccct 960gaagtgaact
tcgagccctt cgtggaagat cactgcggca ccttcagcag cttcatgccc
1020agcgcttgga gaagcagaga ctgcgagagc accctgcctt acatctgcaa
gaagtacctg 1080aaccacatcg accacgagat cgtggaaaag gacgcctgga
agtactacgc cacacactgc 1140gagtccggaa ccacgacgcc agcgccgcga
ccaccaacac cggcgcccac catcgcgtcg 1200cagcccctgt ccctgcgccc
agaggcgtgc cggccagcgg cggggggcgc agtgcacacg 1260agggggctgg
acttcgcctg tgatatctac atctgggcgc ccttggccgg gacttgtggg
1320gtccttctcc tgtcactggt tatcaccctt tactgcaaac ggggcagaaa
gaaactcctg 1380tatatattca aacaaccatt tatgagacca gtacaaacta
ctcaagagga agatggctgt 1440agctgccgat ttccagaaga agaagaagga
ggatgtgaac tgagagtgaa gttcagcagg 1500agcgcagacg cccccgcgta
caagcagggc cagaaccagc tctataacga gctcaatcta 1560ggacgaagag
aggagtacga tgttttggac aagagacgtg gccgggaccc tgagatgggg
1620ggaaagccga gaaggaagaa ccctcaggaa ggcctgtaca atgaactgca
gaaagataag 1680atggcggagg cctacagtga gattgggatg aaaggcgagc
gccggagggg caaggggcac 1740gatggccttt accagggtct cagtacagcc
accaaggaca cctacgacgc ccttcacatg 1800caggccctgc cccctcgcta a
182140606PRTArtificial SequencePLA2R-CAAR Construct 4026.CF1 40Met
Leu Leu Ser Pro Ser Leu Leu Leu Leu Leu Leu Leu Gly Ala Pro1 5 10
15Arg Gly Cys Ala Gly Ser Glu Gly Val Ala Ala Ala Leu Thr Pro Glu
20 25 30Arg Leu Leu Glu Trp Gln Asp Lys Gly Ile Phe Val Ile Gln Ser
Glu 35 40 45Ser Leu Lys Lys Cys Ile Gln Ala Gly Lys Ser Val Leu Thr
Leu Glu 50 55 60Asn Cys Lys Gln Ala Asn Lys His Met Leu Trp Lys Trp
Val Ser Asn65 70 75 80His Gly Leu Phe Asn Ile Gly Gly Ser Gly Cys
Leu Gly Leu Asn Phe 85 90 95Ser Ala Pro Glu Gln Pro Leu Ser Leu Tyr
Glu Cys Asp Ser Thr Leu 100 105 110Val Ser Leu Arg Trp Arg Cys Asn
Arg Lys Met Ile Thr Gly Pro Leu 115 120 125Gln Tyr Ser Val Gln Val
Ala His Asp Asn Thr Val Val Ala Ser Arg 130 135 140Lys Tyr Ile His
Lys Trp Ile Ser Tyr Gly Ser Gly Gly Gly Asp Ile145 150 155 160Cys
Glu Tyr Leu His Lys Asp Leu His Thr Ile Lys Gly Asn Thr His 165 170
175Gly Met Pro Cys Met Phe Pro Phe Gln Tyr Asn His Gln Trp His His
180 185 190Glu Cys Thr Arg Glu Gly Arg Glu Asp Asp Leu Leu Trp Cys
Ala Thr 195 200 205Thr Ser Arg Tyr Glu Arg Asp Glu Lys Trp Gly Phe
Cys Pro Asp Pro 210 215 220Thr Ser Ala Glu Val Gly Cys Asp Thr Ile
Trp Glu Lys Asp Leu Asn225 230 235 240Ser His Ile Cys Tyr Gln Phe
Asn Leu Leu Ser Ser Leu Ser Trp Ser 245 250 255Glu Ala His Ser Ser
Cys Gln Met Gln Gly Gly Thr Leu Leu Ser Ile 260 265 270Thr Asp Glu
Thr Glu Glu Asn Phe Ile Arg Glu His Met Ser Ser Lys 275 280 285Thr
Val Glu Val Trp Met Gly Leu Asn Gln Leu Asp Glu His Ala Gly 290 295
300Trp Gln Trp Ser Asp Gly Thr Pro Leu Asn Tyr Leu Asn Trp Ser
Pro305 310 315 320Glu Val Asn Phe Glu Pro Phe Val Glu Asp His Cys
Gly Thr Phe Ser 325 330 335Ser Phe Met Pro Ser Ala Trp Arg Ser Arg
Asp Cys Glu Ser Thr Leu 340 345 350Pro Tyr Ile Cys Lys Lys Tyr Leu
Asn His Ile Asp His Glu Ile Val 355 360 365Glu Lys Asp Ala Trp Lys
Tyr Tyr Ala Thr His Cys Glu Ser Gly Thr 370 375 380Thr Thr Pro Ala
Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala Ser385 390 395 400Gln
Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala Ala Gly Gly 405 410
415Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys Asp Ile Tyr Ile Trp
420 425 430Ala Pro Leu Ala Gly Thr Cys Gly Val Leu Leu Leu Ser Leu
Val Ile 435 440 445Thr Leu Tyr Cys Lys Arg Gly Arg Lys Lys Leu Leu
Tyr Ile Phe Lys 450 455 460Gln Pro Phe Met Arg Pro Val Gln Thr Thr
Gln Glu Glu Asp Gly Cys465 470 475 480Ser Cys Arg Phe Pro Glu Glu
Glu Glu Gly Gly Cys Glu Leu Arg Val 485 490 495Lys Phe Ser Arg Ser
Ala Asp Ala Pro Ala Tyr Lys Gln Gly Gln Asn 500 505 510Gln Leu Tyr
Asn Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val 515 520 525Leu
Asp Lys Arg Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg 530 535
540Arg Lys Asn Pro Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp
Lys545 550 555 560Met Ala Glu Ala Tyr Ser Glu Ile Gly Met Lys Gly
Glu Arg Arg Arg 565 570 575Gly Lys Gly His Asp Gly Leu Tyr Gln Gly
Leu Ser Thr Ala Thr Lys 580 585 590Asp Thr Tyr Asp Ala Leu His Met
Gln Ala Leu Pro Pro Arg 595 600 605414441DNAHomo sapiens
41ccggagagcc cagtggttag cgatgctgct gtcgccgtcg ctgctgctgc tgctgctgct
60gggggcgccg cggggctgcg ccgagggtgt ggcggcggcg cttacccccg agcggctcct
120ggagtggcag gataaaggaa tatttgttat ccaaagtgag agtctcaaga
aatgcattca 180agcaggtaaa tcggttctga ccctggagaa ctgcaagcaa
gcaaacaagc acatgctgtg 240gaaatgggtt tcaaaccatg gcctctttaa
cataggaggc agcggttgcc tgggcctgaa 300tttctccgcc ccagagcagc
cattaagctt atatgaatgt gactccaccc tcgtttcctt 360acggtggcgc
tgtaacagga agatgatcac aggcccgctg cagtactctg tccaggtggc
420gcatgacaac acagtggtgg cctcacggaa gtatattcat aagtggattt
cttatgggtc 480aggtggtgga gacatttgtg aatatctaca caaagatttg
catacaatca aagggaacac 540ccacgggatg ccgtgtatgt ttcccttcca
gtataaccat cagtggcatc atgaatgtac 600ccgtgaaggt cgggaagatg
acttactgtg gtgtgccacg acaagccgtt atgaaagaga 660tgaaaagtgg
ggattttgcc ctgatcccac ctctgcagaa gtaggttgtg atactatttg
720ggagaaggac ctcaattcac acatttgcta ccagttcaac ctgctttcat
ctctctcttg 780gagtgaggca cattcttcat gccagatgca aggaggtacg
ctgttaagta ttacagatga 840aactgaagaa aatttcataa gggagcacat
gagcagtaaa acagtggagg tgtggatggg 900cctcaatcag ctggatgaac
acgctggctg gcagtggtct gatggaacgc cgctcaacta 960tctgaattgg
agcccagagg taaattttga gccatttgtt gaagatcact gtggaacatt
1020tagttcattt atgccaagtg cctggaggag tcgggattgt gagtccacct
tgccatatat 1080atgtaaaaaa tatctaaacc acattgatca tgaaatagtt
gaaaaagatg cgtggaaata 1140ttatgctacc cactgtgagc ctggctggaa
tccctacaat cgtaattgct acaaacttca 1200gaaagaagaa aagacctggc
atgaggctct gcgttcttgt caggctgata acagtgcatt 1260aatagacata
acctcattag cagaggtgga gtttcttgta accctccttg gagatgaaaa
1320tgcatcagaa acatggattg gtttgagcag caataaaatt ccagtttcct
ttgaatggtc 1380taatgactct tcagtcatct ttactaattg gcacacactt
gagccccaca tttttccaaa 1440tagaagccag ctgtgtgtct cagcagagca
gtctgaggga cactggaaag tcaaaaattg 1500tgaagaaaga cttttttaca
tttgtaaaaa agcaggccat gtcctctctg atgctgaatc 1560aggatgtcaa
gagggatggg agagacatgg tggattctgt tacaaaattg acacagtcct
1620tcgaagcttt gaccaagctt ccagcggtta ttactgtcct cctgcacttg
taaccattac 1680aaacaggttt gaacaggctt ttattaccag tttgatcagt
agtgtggtaa aaatgaagga 1740cagttatttt tggatagctc ttcaggacca
aaatgatacg ggagaataca cttggaagcc 1800agtagggcag aaacccgagc
cggtgcagta cacacactgg aacacacacc agccgcgcta 1860cagtggtggc
tgtgttgcca tgcgaggaag gcatccactt ggtcgctggg aagtgaagca
1920ctgtcggcac tttaaggcaa tgtccttgtg caagcagcca gttgaaaatc
aggaaaaagc 1980agagtatgaa gagagatggc cctttcaccc ctgctatttg
gactgggagt cagagcctgg 2040tctggccagt tgcttcaagg tatttcatag
tgaaaaagtt ctgatgaaaa gaacatggag 2100agaagctgaa gcattttgcg
aagaatttgg agctcatctt gcaagctttg cccatattga 2160ggaagagaat
tttgtgaatg agctcttaca ttcaaaattt aattggacag aagaaaggca
2220gttctggatt ggatttaata aaagaaaccc actgaatgcc ggctcatggg
agtggtctga 2280tagaactcct gttgtctctt cgtttttaga caacacttat
tttggagaag atgcaagaaa 2340ctgtgctgtt tataaggcaa acaaaacatt
gctgccctta cactgtggtt ccaaacgtga 2400atggatatgc aaaatcccaa
gagatgtgaa acccaagatt ccgttctggt accagtacga 2460tgtaccctgg
ctcttttatc aggatgcaga ataccttttt catacctttg cctcagaatg
2520gttgaacttt gagtttgtct gtagctggct gcacagtgat cttctcacaa
ttcattctgc 2580acatgagcaa gaattcatcc acagcaaaat aaaagcgcta
tcaaagtatg gtgcaagttg 2640gtggattgga cttcaagaag aaagagccaa
tgatgaattt cgctggagag atggaacacc 2700agtgatatac cagaactggg
acacaggaag agaaagaact gtgaataatc agagccagag 2760atgtggcttt
atttcttcta taacaggact ctggggtagt gaagagtgtt cagtttctat
2820gcctagtatc tgtaagcgaa aaaaggtttg gctcatagag aaaaagaaag
atacaccaaa 2880acaacatgga acgtgtccca aaggatggct atattttaac
tataagtgcc ttctgctgaa 2940tatccccaaa gacccaagca gttggaagaa
ctggacgcat gctcaacatt tctgtgctga 3000agaagggggg accctggtcg
ccattgaaag tgaggtggag caagctttca ttactatgaa 3060tctttttggc
cagaccacca gtgtgtggat aggtttacaa aatgatgatt atgaaacatg
3120gctaaatgga aagcctgtgg tatattctaa ctggtctcca tttgatataa
taaatattcc 3180aagtcacaat accactgaag ttcagaaaca cattcctctc
tgtgccttac tctcaagtaa 3240tcctaatttt catttcactg gaaaatggta
ttttgaagac tgtggaaagg aaggctatgg 3300gtttgtttgt gaaaaaatgc
aagatacttc tggacacggt gtaaatacat ctgatatgta 3360tccaatgccc
aataccttag aatatggaaa cagaacttac aaaataatta atgcaaatat
3420gacttggtat gcagcaataa aaacctgcct gatgcacaaa gcacaactgg
tcagcatcac 3480agaccagtat caccagtcct tcctcactgt tgtcctcaac
cggctaggat atgcccactg 3540gattggactg ttcaccacag ataatggtct
taattttgac tggtctgatg gcaccaaatc 3600ttctttcact ttttggaaag
atgaggagtc ctccctcctt ggtgactgcg tttttgccga 3660cagcaacgga
cgctggcata gcacagcctg cgagtcattt ctgcaaggtg ccatttgtca
3720tgtgccacct gaaacaagac aatctgaaca cccagagttg tgctcagaaa
catctattcc 3780ctggataaaa tttaaaagta attgctacag tttttctaca
gtcctagaca gtatgagttt 3840tgaggctgct catgaatttt gcaaaaagga
aggttctaat cttttaacaa tcaaggatga 3900ggctgaaaat gcatttctcc
tagaagagct gtttgctttt ggttcttctg tccagatggt 3960ttggttgaat
gctcaatttg atgatgaaac cataaagtgg tttgatggaa ctcccacaga
4020ccagtcaaac tggggcattc ggaagccaga cacagactac ttcaagcccc
atcattgtgt 4080tgccttgagg atccctgaag gattatggca gctatccccg
tgtcaagaaa aaaaaggctt 4140tatatgtaaa atggaggcag atattcacac
tgcagaggcg ctgccagaaa aaggaccaag 4200tcacagcatc attcctcttg
cggttgtact gacactgata gtcattgtgg ccatttgcac 4260actttccttc
tgcatataca agcataacgg tggcttcttc aggagacttg cagggtttcg
4320gaatccttac tatcctgcaa ccaactttag tacagtatat ttagaagaaa
atattctcat 4380ttctgatctt gagaagagtg accaataata atgaggtcag
agaatgccac agacaccagg 4440g 444142135DNAArtificial SequenceCD8
alpha hinge
domain 42accacgacgc cagcgccgcg accaccaaca ccggcgccca ccatcgcgtc
gcagcccctg 60tccctgcgcc cagaggcgtg ccggccagcg gcggggggcg cagtgcacac
gagggggctg 120gacttcgcct gtgat 1354345PRTArtificial SequenceCD8
alpha hinge domain 43Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro
Ala Pro Thr Ile Ala1 5 10 15Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala
Cys Arg Pro Ala Ala Gly 20 25 30Gly Ala Val His Thr Arg Gly Leu Asp
Phe Ala Cys Asp 35 40 454455PRTArtificial SequenceCD8 alpha hinge
domain 44Phe Val Pro Val Phe Leu Pro Ala Lys Pro Thr Thr Thr Pro
Ala Pro1 5 10 15Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala Ser Gln Pro
Leu Ser Leu 20 25 30Arg Pro Glu Ala Cys Arg Pro Ala Ala Gly Gly Ala
Val His Thr Arg 35 40 45Gly Leu Asp Phe Ala Cys Asp 50
5545112PRTArtificial SequenceCD3 zeta signaling domain 45Arg Val
Lys Phe Ser Arg Ser Ala Asp Ala Pro Ala Tyr Gln Gln Gly1 5 10 15Gln
Asn Gln Leu Tyr Asn Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr 20 25
30Asp Val Leu Asp Lys Arg Arg Gly Arg Asp Pro Glu Met Gly Gly Lys
35 40 45Pro Arg Arg Lys Asn Pro Gln Glu Gly Leu Tyr Asn Glu Leu Gln
Lys 50 55 60Asp Lys Met Ala Glu Ala Tyr Ser Glu Ile Gly Met Lys Gly
Glu Arg65 70 75 80Arg Arg Gly Lys Gly His Asp Gly Leu Tyr Gln Gly
Leu Ser Thr Ala 85 90 95Thr Lys Asp Thr Tyr Asp Ala Leu His Met Gln
Ala Leu Pro Pro Arg 100 105 1104657DNAArtificial SequenceIgG Signal
peptide 46atggagtttg ggctgagctg gctttttctt gtggctattt taaaaggtgt
ccagtgc 5747396DNAArtificial SequenceCysR 47aaaggaatat ttgtcattca
gtcagagtct ttgaaaaagt gcatacaggc tggaaaaagc 60gtgcttaccc tggagaactg
caagcaagct aataagcata tgctttggaa atgggttagc 120aaccacggac
tctttaatat cggaggctcc ggctgtctgg gcctgaactt cagtgcaccg
180gagcaaccgc tttctctgta cgaatgtgat agcacacttg ttagtcttcg
gtggcggtgt 240aaccgaaaaa tgattacagg ccctctgcaa tatagtgttc
aagtggccca cgacaataca 300gttgtggcgt ctagaaaata tattcacaag
tggatttcct acgggagcgg cggaggggat 360atatgtgaat atcttcacaa
agacttgcat acaatc 3964819PRTArtificial SequenceIgG Signal peptide
48Met Glu Phe Gly Leu Ser Trp Leu Phe Leu Val Ala Ile Leu Lys Gly1
5 10 15Val Gln Cys49132PRTArtificial SequenceCysR 49Lys Gly Ile Phe
Val Ile Gln Ser Glu Ser Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys
Ser Val Leu Thr Leu Glu Asn Cys Lys Gln Ala Asn Lys 20 25 30His Met
Leu Trp Lys Trp Val Ser Asn His Gly Leu Phe Asn Ile Gly 35 40 45Gly
Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala Pro Glu Gln Pro Leu 50 55
60Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser Leu Arg Trp Arg Cys65
70 75 80Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr Ser Val Gln Val
Ala 85 90 95His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr Ile His Lys
Trp Ile 100 105 110Ser Tyr Gly Ser Gly Gly Gly Asp Ile Cys Glu Tyr
Leu His Lys Asp 115 120 125Leu His Thr Ile 13050990DNAArtificial
SequenceCysR-FNII-CTLD1 50aaagggatct ttgttataca aagtgagagc
ttgaaaaaat gtatacaggc tggcaaaagt 60gtactgactc ttgaaaattg caaacaagcc
aacaaacaca tgctgtggaa atgggtgtct 120aatcacggtc tcttcaatat
tgggggaagt ggatgcctcg gcctgaattt ctccgctccc 180gaacagcccc
tctcacttta tgagtgtgat tcaactctgg tgtccttgag gtggcgatgt
240aaccgcaaga tgataaccgg ccccctccag tattccgtcc aagtagcaca
cgacaatacc 300gtggtggcat ctaggaaata cattcataag tggatatctt
atggcagtgg tggcggtgac 360atatgcgagt acctgcacaa ggacctccac
acaataaagg ggaacacgca cgggatgccg 420tgtatgttcc cgttccaata
taatcatcaa tggcaccatg agtgtacgag agaggggcga 480gaagacgacc
tcctgtggtg tgcgaccacc tcaagatatg aacgggatga gaagtggggc
540ttttgccccg acccaacctc cgccgaggtt ggttgcgaca ctatttggga
aaaagatttg 600aacagtcata tatgctatca atttaatttg ttgagttcac
tctcctggag cgaagcgcac 660agctcttgtc agatgcaagg tggtacattg
cttagcatta ctgatgaaac tgaggagaat 720ttcattaggg agcatatgtc
ctcaaagaca gtagaggtgt ggatgggtct gaaccagctc 780gacgaacacg
ccggttggca gtggtcagat ggaacgcctc tgaattatct caactggtcc
840cctgaggtca actttgaacc gtttgtggaa gatcattgtg gtactttttc
cagttttatg 900ccaagcgcct ggcgaagccg agactgcgag tctacgttgc
cctatatctg caagaagtat 960ttgaatcaca tagatcatga aattgttgaa
99051330PRTArtificial SequenceCysR-FNII-CTLD1 51Lys Gly Ile Phe Val
Ile Gln Ser Glu Ser Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys Ser
Val Leu Thr Leu Glu Asn Cys Lys Gln Ala Asn Lys 20 25 30His Met Leu
Trp Lys Trp Val Ser Asn His Gly Leu Phe Asn Ile Gly 35 40 45Gly Ser
Gly Cys Leu Gly Leu Asn Phe Ser Ala Pro Glu Gln Pro Leu 50 55 60Ser
Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser Leu Arg Trp Arg Cys65 70 75
80Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr Ser Val Gln Val Ala
85 90 95His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr Ile His Lys Trp
Ile 100 105 110Ser Tyr Gly Ser Gly Gly Gly Asp Ile Cys Glu Tyr Leu
His Lys Asp 115 120 125Leu His Thr Ile Lys Gly Asn Thr His Gly Met
Pro Cys Met Phe Pro 130 135 140Phe Gln Tyr Asn His Gln Trp His His
Glu Cys Thr Arg Glu Gly Arg145 150 155 160Glu Asp Asp Leu Leu Trp
Cys Ala Thr Thr Ser Arg Tyr Glu Arg Asp 165 170 175Glu Lys Trp Gly
Phe Cys Pro Asp Pro Thr Ser Ala Glu Val Gly Cys 180 185 190Asp Thr
Ile Trp Glu Lys Asp Leu Asn Ser His Ile Cys Tyr Gln Phe 195 200
205Asn Leu Leu Ser Ser Leu Ser Trp Ser Glu Ala His Ser Ser Cys Gln
210 215 220Met Gln Gly Gly Thr Leu Leu Ser Ile Thr Asp Glu Thr Glu
Glu Asn225 230 235 240Phe Ile Arg Glu His Met Ser Ser Lys Thr Val
Glu Val Trp Met Gly 245 250 255Leu Asn Gln Leu Asp Glu His Ala Gly
Trp Gln Trp Ser Asp Gly Thr 260 265 270Pro Leu Asn Tyr Leu Asn Trp
Ser Pro Glu Val Asn Phe Glu Pro Phe 275 280 285Val Glu Asp His Cys
Gly Thr Phe Ser Ser Phe Met Pro Ser Ala Trp 290 295 300Arg Ser Arg
Asp Cys Glu Ser Thr Leu Pro Tyr Ile Cys Lys Lys Tyr305 310 315
320Leu Asn His Ile Asp His Glu Ile Val Glu 325
330521848DNAArtificial SequenceCysR-FNII-CTLD1-3 52aaagggattt
tcgtgataca gtccgagagt ctcaaaaagt gtatccaggc aggcaaaagt 60gttctcactc
tggaaaactg caaacaagcg aacaagcaca tgttgtggaa gtgggttagt
120aaccatggac tgttcaacat cggaggtagt ggatgccttg gtctcaattt
ctctgctccg 180gaacagcctt tgtcactgta cgaatgcgac tccactctcg
ttagtcttag atggcgatgc 240aatcgcaaaa tgattacggg accacttcaa
tattcagttc aagtggcaca tgataacacc 300gtagtggcct cacggaaata
catccataaa tggatttctt atggtagcgg gggcggcgat 360atatgtgaat
acctccataa ggatctccac accattaagg gtaatactca cggtatgccg
420tgtatgtttc cttttcagta caatcatcag tggcatcatg aatgcacgag
ggaaggacgc 480gaggacgatt tgctctggtg cgcaaccacc tcacgctacg
agagagacga aaaatggggc 540ttttgcccgg accccactag tgctgaggta
ggatgtgata cgatttggga aaaggatttg 600aattctcata tttgctacca
gtttaatctt ctttcatccc tgtcctggtc tgaggctcat 660tctagttgcc
agatgcaagg tgggactttg ctttcaatta ctgacgagac tgaggaaaat
720tttatccgag agcatatgtc ttctaaaacc gtagaggtat ggatgggcct
gaaccaattg 780gacgaacacg cgggctggca gtggagcgac gggacacctc
tcaactacct taattggagc 840cctgaggtaa actttgaacc gtttgtcgag
gatcactgcg gaactttcag cagcttcatg 900cctagtgcat ggcggtcccg
agactgtgag agcacccttc catacatatg taaaaaatac 960ctcaatcaca
tagaccacga gatcgtagag aaggatgcat ggaaatatta tgctacgcac
1020tgtgagccgg gatggaatcc ttataaccgc aactgttaca agctgcaaaa
agaagagaag 1080acatggcatg aggcgctgcg ctcatgtcaa gcggacaatt
ctgcacttat agatataact 1140agtttggcgg aggtagaatt tttggttacg
cttctcggcg atgagaatgc gtccgagacg 1200tggatagggt tgtcaagcaa
taaaattcct gtaagttttg aatggtcaaa tgactcttct 1260gtcatcttca
ccaattggca cacactcgaa ccccatatct tcccaaaccg aagccagttg
1320tgtgtcagcg ctgagcaatc agaaggacat tggaaagtta aaaactgtga
agaaagactg 1380ttctacatct gtaagaaggc aggacatgtg ctttcagatg
cggaaagcgg ctgtcaagaa 1440ggttgggagc gccatggagg tttctgttat
aaaatcgaca cagttttgcg atctttcgat 1500caggcttcaa gcgggtacta
ttgtcctcct gcactggtta caatcacgaa ccggtttgaa 1560caggctttta
taacttcttt gatttccagc gtggttaaaa tgaaggactc ttatttctgg
1620atagccctgc aagaccaaaa tgataccggt gagtacacat ggaaaccggt
aggtcaaaag 1680ccagagccag tccagtacac tcattggaat acccaccagc
ctaggtactc cggcgggtgt 1740gtggcgatgc ggggtcgcca ccctctcgga
cgctgggagg tgaagcattg ccgccacttc 1800aaggcgatga gcttgtgtaa
acagcccgtc gaaaatcagg aaaaggca 184853616PRTArtificial
SequenceCysR-FNII-CTLD1-3 53Lys Gly Ile Phe Val Ile Gln Ser Glu Ser
Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys Ser Val Leu Thr Leu Glu
Asn Cys Lys Gln Ala Asn Lys 20 25 30His Met Leu Trp Lys Trp Val Ser
Asn His Gly Leu Phe Asn Ile Gly 35 40 45Gly Ser Gly Cys Leu Gly Leu
Asn Phe Ser Ala Pro Glu Gln Pro Leu 50 55 60Ser Leu Tyr Glu Cys Asp
Ser Thr Leu Val Ser Leu Arg Trp Arg Cys65 70 75 80Asn Arg Lys Met
Ile Thr Gly Pro Leu Gln Tyr Ser Val Gln Val Ala 85 90 95His Asp Asn
Thr Val Val Ala Ser Arg Lys Tyr Ile His Lys Trp Ile 100 105 110Ser
Tyr Gly Ser Gly Gly Gly Asp Ile Cys Glu Tyr Leu His Lys Asp 115 120
125Leu His Thr Ile Lys Gly Asn Thr His Gly Met Pro Cys Met Phe Pro
130 135 140Phe Gln Tyr Asn His Gln Trp His His Glu Cys Thr Arg Glu
Gly Arg145 150 155 160Glu Asp Asp Leu Leu Trp Cys Ala Thr Thr Ser
Arg Tyr Glu Arg Asp 165 170 175Glu Lys Trp Gly Phe Cys Pro Asp Pro
Thr Ser Ala Glu Val Gly Cys 180 185 190Asp Thr Ile Trp Glu Lys Asp
Leu Asn Ser His Ile Cys Tyr Gln Phe 195 200 205Asn Leu Leu Ser Ser
Leu Ser Trp Ser Glu Ala His Ser Ser Cys Gln 210 215 220Met Gln Gly
Gly Thr Leu Leu Ser Ile Thr Asp Glu Thr Glu Glu Asn225 230 235
240Phe Ile Arg Glu His Met Ser Ser Lys Thr Val Glu Val Trp Met Gly
245 250 255Leu Asn Gln Leu Asp Glu His Ala Gly Trp Gln Trp Ser Asp
Gly Thr 260 265 270Pro Leu Asn Tyr Leu Asn Trp Ser Pro Glu Val Asn
Phe Glu Pro Phe 275 280 285Val Glu Asp His Cys Gly Thr Phe Ser Ser
Phe Met Pro Ser Ala Trp 290 295 300Arg Ser Arg Asp Cys Glu Ser Thr
Leu Pro Tyr Ile Cys Lys Lys Tyr305 310 315 320Leu Asn His Ile Asp
His Glu Ile Val Glu Lys Asp Ala Trp Lys Tyr 325 330 335Tyr Ala Thr
His Cys Glu Pro Gly Trp Asn Pro Tyr Asn Arg Asn Cys 340 345 350Tyr
Lys Leu Gln Lys Glu Glu Lys Thr Trp His Glu Ala Leu Arg Ser 355 360
365Cys Gln Ala Asp Asn Ser Ala Leu Ile Asp Ile Thr Ser Leu Ala Glu
370 375 380Val Glu Phe Leu Val Thr Leu Leu Gly Asp Glu Asn Ala Ser
Glu Thr385 390 395 400Trp Ile Gly Leu Ser Ser Asn Lys Ile Pro Val
Ser Phe Glu Trp Ser 405 410 415Asn Asp Ser Ser Val Ile Phe Thr Asn
Trp His Thr Leu Glu Pro His 420 425 430Ile Phe Pro Asn Arg Ser Gln
Leu Cys Val Ser Ala Glu Gln Ser Glu 435 440 445Gly His Trp Lys Val
Lys Asn Cys Glu Glu Arg Leu Phe Tyr Ile Cys 450 455 460Lys Lys Ala
Gly His Val Leu Ser Asp Ala Glu Ser Gly Cys Gln Glu465 470 475
480Gly Trp Glu Arg His Gly Gly Phe Cys Tyr Lys Ile Asp Thr Val Leu
485 490 495Arg Ser Phe Asp Gln Ala Ser Ser Gly Tyr Tyr Cys Pro Pro
Ala Leu 500 505 510Val Thr Ile Thr Asn Arg Phe Glu Gln Ala Phe Ile
Thr Ser Leu Ile 515 520 525Ser Ser Val Val Lys Met Lys Asp Ser Tyr
Phe Trp Ile Ala Leu Gln 530 535 540Asp Gln Asn Asp Thr Gly Glu Tyr
Thr Trp Lys Pro Val Gly Gln Lys545 550 555 560Pro Glu Pro Val Gln
Tyr Thr His Trp Asn Thr His Gln Pro Arg Tyr 565 570 575Ser Gly Gly
Cys Val Ala Met Arg Gly Arg His Pro Leu Gly Arg Trp 580 585 590Glu
Val Lys His Cys Arg His Phe Lys Ala Met Ser Leu Cys Lys Gln 595 600
605Pro Val Glu Asn Gln Glu Lys Ala 610 615542268DNAArtificial
SequenceCysR-FNII-CTLD1-3 7 54aaagggattt tcgtgataca gtccgagagt
ctcaaaaagt gtatccaggc aggcaaaagt 60gttctcactc tggaaaactg caaacaagcg
aacaagcaca tgttgtggaa gtgggttagt 120aaccatggac tgttcaacat
cggaggtagt ggatgccttg gtctcaattt ctctgctccg 180gaacagcctt
tgtcactgta cgaatgcgac tccactctcg ttagtcttag atggcgatgc
240aatcgcaaaa tgattacggg accacttcaa tattcagttc aagtggcaca
tgataacacc 300gtagtggcct cacggaaata catccataaa tggatttctt
atggtagcgg gggcggcgat 360atatgtgaat acctccataa ggatctccac
accattaagg gtaatactca cggtatgccg 420tgtatgtttc cttttcagta
caatcatcag tggcatcatg aatgcacgag ggaaggacgc 480gaggacgatt
tgctctggtg cgcaaccacc tcacgctacg agagagacga aaaatggggc
540ttttgcccgg accccactag tgctgaggta ggatgtgata cgatttggga
aaaggatttg 600aattctcata tttgctacca gtttaatctt ctttcatccc
tgtcctggtc tgaggctcat 660tctagttgcc agatgcaagg tgggactttg
ctttcaatta ctgacgagac tgaggaaaat 720tttatccgag agcatatgtc
ttctaaaacc gtagaggtat ggatgggcct gaaccaattg 780gacgaacacg
cgggctggca gtggagcgac gggacacctc tcaactacct taattggagc
840cctgaggtaa actttgaacc gtttgtcgag gatcactgcg gaactttcag
cagcttcatg 900cctagtgcat ggcggtcccg agactgtgag agcacccttc
catacatatg taaaaaatac 960ctcaatcaca tagaccacga gatcgtagag
aaggatgcat ggaaatatta tgctacgcac 1020tgtgagccgg gatggaatcc
ttataaccgc aactgttaca agctgcaaaa agaagagaag 1080acatggcatg
aggcgctgcg ctcatgtcaa gcggacaatt ctgcacttat agatataact
1140agtttggcgg aggtagaatt tttggttacg cttctcggcg atgagaatgc
gtccgagacg 1200tggatagggt tgtcaagcaa taaaattcct gtaagttttg
aatggtcaaa tgactcttct 1260gtcatcttca ccaattggca cacactcgaa
ccccatatct tcccaaaccg aagccagttg 1320tgtgtcagcg ctgagcaatc
agaaggacat tggaaagtta aaaactgtga agaaagactg 1380ttctacatct
gtaagaaggc aggacatgtg ctttcagatg cggaaagcgg ctgtcaagaa
1440ggttgggagc gccatggagg tttctgttat aaaatcgaca cagttttgcg
atctttcgat 1500caggcttcaa gcgggtacta ttgtcctcct gcactggtta
caatcacgaa ccggtttgaa 1560caggctttta taacttcttt gatttccagc
gtggttaaaa tgaaggactc ttatttctgg 1620atagccctgc aagaccaaaa
tgataccggt gagtacacat ggaaaccggt aggtcaaaag 1680ccagagccag
tccagtacac tcattggaat acccaccagc ctaggtactc cggcgggtgt
1740gtggcgatgc ggggtcgcca ccctctcgga cgctgggagg tgaagcattg
ccgccacttc 1800aaggcgatga gcttgtgtaa acagcccgtc gaaaatcagg
aaaaggcagt taacacatct 1860gatatgtacc ctatgcctaa cacactcgaa
tatgggaata ggacgtacaa gattataaac 1920gcgaacatga cgtggtatgc
tgcaatcaag acgtgcctca tgcacaaagc tcagcttgtg 1980tctattactg
accaatacca ccaatcattt ttgacagtcg tgttgaatcg attggggtac
2040gcccattgga tcggtctctt cacgacggac aatgggctca attttgactg
gagtgacggt 2100actaaatcat cctttacttt ttggaaggat gaagaaagtt
ctctgttggg cgattgcgtg 2160tttgctgact caaatggccg atggcattcc
acagcctgtg aaagttttct gcagggagct 2220atttgccacg tgcctcccga
aacgcggcag tccgaacacc cggaattg 226855756PRTArtificial
SequenceCysR-FNII-CTLD1-3 7 55Lys Gly Ile Phe Val Ile Gln Ser Glu
Ser Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys Ser Val Leu Thr Leu
Glu Asn Cys Lys Gln Ala Asn Lys 20 25 30His Met Leu Trp Lys Trp Val
Ser Asn His Gly Leu Phe Asn Ile Gly 35 40 45Gly Ser Gly Cys Leu Gly
Leu Asn Phe Ser Ala Pro Glu Gln Pro Leu 50 55 60Ser Leu Tyr Glu Cys
Asp Ser Thr Leu Val Ser Leu Arg Trp Arg Cys65 70 75 80Asn Arg Lys
Met Ile Thr Gly Pro Leu Gln Tyr Ser Val Gln Val Ala 85 90 95His Asp
Asn Thr Val Val Ala Ser Arg Lys Tyr Ile His Lys Trp Ile 100 105
110Ser Tyr Gly Ser Gly Gly Gly Asp Ile Cys Glu Tyr Leu His Lys
Asp
115 120 125Leu His Thr Ile Lys Gly Asn Thr His Gly Met Pro Cys Met
Phe Pro 130 135 140Phe Gln Tyr Asn His Gln Trp His His Glu Cys Thr
Arg Glu Gly Arg145 150 155 160Glu Asp Asp Leu Leu Trp Cys Ala Thr
Thr Ser Arg Tyr Glu Arg Asp 165 170 175Glu Lys Trp Gly Phe Cys Pro
Asp Pro Thr Ser Ala Glu Val Gly Cys 180 185 190Asp Thr Ile Trp Glu
Lys Asp Leu Asn Ser His Ile Cys Tyr Gln Phe 195 200 205Asn Leu Leu
Ser Ser Leu Ser Trp Ser Glu Ala His Ser Ser Cys Gln 210 215 220Met
Gln Gly Gly Thr Leu Leu Ser Ile Thr Asp Glu Thr Glu Glu Asn225 230
235 240Phe Ile Arg Glu His Met Ser Ser Lys Thr Val Glu Val Trp Met
Gly 245 250 255Leu Asn Gln Leu Asp Glu His Ala Gly Trp Gln Trp Ser
Asp Gly Thr 260 265 270Pro Leu Asn Tyr Leu Asn Trp Ser Pro Glu Val
Asn Phe Glu Pro Phe 275 280 285Val Glu Asp His Cys Gly Thr Phe Ser
Ser Phe Met Pro Ser Ala Trp 290 295 300Arg Ser Arg Asp Cys Glu Ser
Thr Leu Pro Tyr Ile Cys Lys Lys Tyr305 310 315 320Leu Asn His Ile
Asp His Glu Ile Val Glu Lys Asp Ala Trp Lys Tyr 325 330 335Tyr Ala
Thr His Cys Glu Pro Gly Trp Asn Pro Tyr Asn Arg Asn Cys 340 345
350Tyr Lys Leu Gln Lys Glu Glu Lys Thr Trp His Glu Ala Leu Arg Ser
355 360 365Cys Gln Ala Asp Asn Ser Ala Leu Ile Asp Ile Thr Ser Leu
Ala Glu 370 375 380Val Glu Phe Leu Val Thr Leu Leu Gly Asp Glu Asn
Ala Ser Glu Thr385 390 395 400Trp Ile Gly Leu Ser Ser Asn Lys Ile
Pro Val Ser Phe Glu Trp Ser 405 410 415Asn Asp Ser Ser Val Ile Phe
Thr Asn Trp His Thr Leu Glu Pro His 420 425 430Ile Phe Pro Asn Arg
Ser Gln Leu Cys Val Ser Ala Glu Gln Ser Glu 435 440 445Gly His Trp
Lys Val Lys Asn Cys Glu Glu Arg Leu Phe Tyr Ile Cys 450 455 460Lys
Lys Ala Gly His Val Leu Ser Asp Ala Glu Ser Gly Cys Gln Glu465 470
475 480Gly Trp Glu Arg His Gly Gly Phe Cys Tyr Lys Ile Asp Thr Val
Leu 485 490 495Arg Ser Phe Asp Gln Ala Ser Ser Gly Tyr Tyr Cys Pro
Pro Ala Leu 500 505 510Val Thr Ile Thr Asn Arg Phe Glu Gln Ala Phe
Ile Thr Ser Leu Ile 515 520 525Ser Ser Val Val Lys Met Lys Asp Ser
Tyr Phe Trp Ile Ala Leu Gln 530 535 540Asp Gln Asn Asp Thr Gly Glu
Tyr Thr Trp Lys Pro Val Gly Gln Lys545 550 555 560Pro Glu Pro Val
Gln Tyr Thr His Trp Asn Thr His Gln Pro Arg Tyr 565 570 575Ser Gly
Gly Cys Val Ala Met Arg Gly Arg His Pro Leu Gly Arg Trp 580 585
590Glu Val Lys His Cys Arg His Phe Lys Ala Met Ser Leu Cys Lys Gln
595 600 605Pro Val Glu Asn Gln Glu Lys Ala Val Asn Thr Ser Asp Met
Tyr Pro 610 615 620Met Pro Asn Thr Leu Glu Tyr Gly Asn Arg Thr Tyr
Lys Ile Ile Asn625 630 635 640Ala Asn Met Thr Trp Tyr Ala Ala Ile
Lys Thr Cys Leu Met His Lys 645 650 655Ala Gln Leu Val Ser Ile Thr
Asp Gln Tyr His Gln Ser Phe Leu Thr 660 665 670Val Val Leu Asn Arg
Leu Gly Tyr Ala His Trp Ile Gly Leu Phe Thr 675 680 685Thr Asp Asn
Gly Leu Asn Phe Asp Trp Ser Asp Gly Thr Lys Ser Ser 690 695 700Phe
Thr Phe Trp Lys Asp Glu Glu Ser Ser Leu Leu Gly Asp Cys Val705 710
715 720Phe Ala Asp Ser Asn Gly Arg Trp His Ser Thr Ala Cys Glu Ser
Phe 725 730 735Leu Gln Gly Ala Ile Cys His Val Pro Pro Glu Thr Arg
Gln Ser Glu 740 745 750His Pro Glu Leu 755561410DNAArtificial
SequenceCysR-FNII-CTLD1 7 56aaaggcatct tcgtaatcca gtcagaaagt
ttgaaaaaat gtatccaagc tggcaaatca 60gtacttaccc ttgagaactg caagcaagcc
aataaacata tgctgtggaa atgggtctca 120aaccacggcc tcttcaatat
tggtgggtca ggttgcttgg ggttgaattt ctccgcccca 180gagcaaccac
tcagccttta cgagtgtgat tccacacttg tctctttgcg atggcgctgc
240aataggaaaa tgatcacagg cccccttcag tactctgtgc aagttgctca
tgataacaca 300gtcgtggcga gtcggaaata tattcacaaa tggatttctt
atgggagtgg tggaggagat 360atatgcgagt atttgcataa ggacttgcac
accatcaagg gaaacactca cggtatgcca 420tgtatgtttc cgttccaata
taatcatcaa tggcaccacg aatgtacccg agagggacgc 480gaggacgatc
ttctttggtg cgccacaacc tctcgatatg aacgagatga gaagtggggg
540ttttgtcctg acccaaccag tgcagaagta gggtgcgata ccatctggga
gaaagacttg 600aactcacaca tatgctatca gtttaatttg ttgtcttctt
tgtcatggag cgaagctcat 660tcatcatgcc agatgcaggg cgggacactg
ctttctatca ccgacgagac tgaggaaaat 720tttatccgcg agcacatgtc
aagcaagaca gttgaggttt ggatggggct caatcaactg 780gacgaacacg
cagggtggca gtggtccgat ggcactccgc tcaactacct taactggagc
840ccagaggtga actttgagcc gtttgtcgaa gatcactgtg gtacttttag
ctccttcatg 900ccgtccgcat ggagaagtcg cgactgcgag tcaaccctcc
cttacatctg taagaaatac 960ctcaaccaca tagatcacga aatcgtagag
gtcaatacgt ccgacatgta cccaatgcca 1020aatacgttgg aatatgggaa
taggacatac aagataatta acgcaaatat gacgtggtat 1080gccgcaatca
aaacgtgcct catgcacaag gcacagctcg tgtcaattac ggaccagtac
1140caccaatcat ttctcacagt cgttcttaat cgattgggtt atgcacactg
gataggcttg 1200ttcacgacgg acaatggttt gaactttgac tggtccgatg
gaactaaaag ttctttcact 1260ttttggaagg atgaggagtc ctccttgctc
ggggactgcg tcttcgcaga ttcaaacggg 1320cgctggcact caacggcatg
tgagtccttc ctgcagggag ctatatgcca tgtgccacca 1380gaaacacgcc
agtctgagca ccctgagttg 141057470PRTArtificial
SequenceCysR-FNII-CTLD1 7 57Lys Gly Ile Phe Val Ile Gln Ser Glu Ser
Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys Ser Val Leu Thr Leu Glu
Asn Cys Lys Gln Ala Asn Lys 20 25 30His Met Leu Trp Lys Trp Val Ser
Asn His Gly Leu Phe Asn Ile Gly 35 40 45Gly Ser Gly Cys Leu Gly Leu
Asn Phe Ser Ala Pro Glu Gln Pro Leu 50 55 60Ser Leu Tyr Glu Cys Asp
Ser Thr Leu Val Ser Leu Arg Trp Arg Cys65 70 75 80Asn Arg Lys Met
Ile Thr Gly Pro Leu Gln Tyr Ser Val Gln Val Ala 85 90 95His Asp Asn
Thr Val Val Ala Ser Arg Lys Tyr Ile His Lys Trp Ile 100 105 110Ser
Tyr Gly Ser Gly Gly Gly Asp Ile Cys Glu Tyr Leu His Lys Asp 115 120
125Leu His Thr Ile Lys Gly Asn Thr His Gly Met Pro Cys Met Phe Pro
130 135 140Phe Gln Tyr Asn His Gln Trp His His Glu Cys Thr Arg Glu
Gly Arg145 150 155 160Glu Asp Asp Leu Leu Trp Cys Ala Thr Thr Ser
Arg Tyr Glu Arg Asp 165 170 175Glu Lys Trp Gly Phe Cys Pro Asp Pro
Thr Ser Ala Glu Val Gly Cys 180 185 190Asp Thr Ile Trp Glu Lys Asp
Leu Asn Ser His Ile Cys Tyr Gln Phe 195 200 205Asn Leu Leu Ser Ser
Leu Ser Trp Ser Glu Ala His Ser Ser Cys Gln 210 215 220Met Gln Gly
Gly Thr Leu Leu Ser Ile Thr Asp Glu Thr Glu Glu Asn225 230 235
240Phe Ile Arg Glu His Met Ser Ser Lys Thr Val Glu Val Trp Met Gly
245 250 255Leu Asn Gln Leu Asp Glu His Ala Gly Trp Gln Trp Ser Asp
Gly Thr 260 265 270Pro Leu Asn Tyr Leu Asn Trp Ser Pro Glu Val Asn
Phe Glu Pro Phe 275 280 285Val Glu Asp His Cys Gly Thr Phe Ser Ser
Phe Met Pro Ser Ala Trp 290 295 300Arg Ser Arg Asp Cys Glu Ser Thr
Leu Pro Tyr Ile Cys Lys Lys Tyr305 310 315 320Leu Asn His Ile Asp
His Glu Ile Val Glu Val Asn Thr Ser Asp Met 325 330 335Tyr Pro Met
Pro Asn Thr Leu Glu Tyr Gly Asn Arg Thr Tyr Lys Ile 340 345 350Ile
Asn Ala Asn Met Thr Trp Tyr Ala Ala Ile Lys Thr Cys Leu Met 355 360
365His Lys Ala Gln Leu Val Ser Ile Thr Asp Gln Tyr His Gln Ser Phe
370 375 380Leu Thr Val Val Leu Asn Arg Leu Gly Tyr Ala His Trp Ile
Gly Leu385 390 395 400Phe Thr Thr Asp Asn Gly Leu Asn Phe Asp Trp
Ser Asp Gly Thr Lys 405 410 415Ser Ser Phe Thr Phe Trp Lys Asp Glu
Glu Ser Ser Leu Leu Gly Asp 420 425 430Cys Val Phe Ala Asp Ser Asn
Gly Arg Trp His Ser Thr Ala Cys Glu 435 440 445Ser Phe Leu Gln Gly
Ala Ile Cys His Val Pro Pro Glu Thr Arg Gln 450 455 460Ser Glu His
Pro Glu Leu465 470581251DNAArtificial SequenceCysR-CTLD1 7
58aaaggaatct tcgtaattca atctgagagt ctgaaaaaat gtattcaggc cggtaagagc
60gtactcacgc ttgaaaattg caaacaggcc aacaaacaca tgctttggaa atgggtttca
120aatcacgggt tgtttaacat agggggatca ggatgtctgg gccttaactt
ttccgcacct 180gaacaacctc ttagtctgta tgagtgtgac tcaacgctgg
tctccttgcg ctggagatgc 240aatcggaaga tgataaccgg gcccctccag
tattccgttc aggtcgccca cgataatact 300gttgttgcat cccgaaaata
tattcataag tggatctcct acgggagtgg agggggcgat 360atttgtgaat
acctccacaa ggatctgcac actatcactt ctgcggaagt aggctgtgac
420acaatctggg agaaagatct gaattcacac atttgctatc agttcaatct
tctgagttct 480ttgagctggt ccgaagcaca ttcatcctgt cagatgcaag
gtggaacact cttgtcaata 540acagatgaaa cggaagagaa ctttattaga
gaacatatgt cctcaaagac tgtggaggtg 600tggatgggac ttaaccagct
cgatgaacat gcaggatggc agtggagtga cggaacgcca 660ctgaactacc
tgaattggag cccagaggtg aatttcgagc ctttcgtaga ggaccattgc
720ggtacttttt catcttttat gcccagcgca tggagatccc gagattgtga
aagcacgctg 780ccctatattt gtaaaaagta cctgaaccac atagatcatg
agatagttga ggtaaataca 840agtgatatgt accccatgcc gaacacactc
gagtacggaa atagaaccta caagataatc 900aacgctaaca tgacctggta
cgcggccatt aagacctgcc tcatgcacaa ggctcaactc 960gtcagtatta
ctgaccaata tcaccagtca tttctcaccg tcgtgttgaa tcgcctcggt
1020tacgcccact ggatcggttt gtttacaacg gacaatggac tcaatttcga
ttggtcagac 1080ggaaccaaat ctagttttac cttctggaaa gacgaggaat
caagcctgct tggggactgc 1140gtatttgcgg actctaatgg ccgatggcat
agtacagcgt gtgagagctt tttgcagggg 1200gcgatttgtc atgttccgcc
ggaaacccgc caaagcgagc atccagaatt g 125159417PRTArtificial
SequenceCysR-CTLD1 7 59Lys Gly Ile Phe Val Ile Gln Ser Glu Ser Leu
Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys Ser Val Leu Thr Leu Glu Asn
Cys Lys Gln Ala Asn Lys 20 25 30His Met Leu Trp Lys Trp Val Ser Asn
His Gly Leu Phe Asn Ile Gly 35 40 45Gly Ser Gly Cys Leu Gly Leu Asn
Phe Ser Ala Pro Glu Gln Pro Leu 50 55 60Ser Leu Tyr Glu Cys Asp Ser
Thr Leu Val Ser Leu Arg Trp Arg Cys65 70 75 80Asn Arg Lys Met Ile
Thr Gly Pro Leu Gln Tyr Ser Val Gln Val Ala 85 90 95His Asp Asn Thr
Val Val Ala Ser Arg Lys Tyr Ile His Lys Trp Ile 100 105 110Ser Tyr
Gly Ser Gly Gly Gly Asp Ile Cys Glu Tyr Leu His Lys Asp 115 120
125Leu His Thr Ile Thr Ser Ala Glu Val Gly Cys Asp Thr Ile Trp Glu
130 135 140Lys Asp Leu Asn Ser His Ile Cys Tyr Gln Phe Asn Leu Leu
Ser Ser145 150 155 160Leu Ser Trp Ser Glu Ala His Ser Ser Cys Gln
Met Gln Gly Gly Thr 165 170 175Leu Leu Ser Ile Thr Asp Glu Thr Glu
Glu Asn Phe Ile Arg Glu His 180 185 190Met Ser Ser Lys Thr Val Glu
Val Trp Met Gly Leu Asn Gln Leu Asp 195 200 205Glu His Ala Gly Trp
Gln Trp Ser Asp Gly Thr Pro Leu Asn Tyr Leu 210 215 220Asn Trp Ser
Pro Glu Val Asn Phe Glu Pro Phe Val Glu Asp His Cys225 230 235
240Gly Thr Phe Ser Ser Phe Met Pro Ser Ala Trp Arg Ser Arg Asp Cys
245 250 255Glu Ser Thr Leu Pro Tyr Ile Cys Lys Lys Tyr Leu Asn His
Ile Asp 260 265 270His Glu Ile Val Glu Val Asn Thr Ser Asp Met Tyr
Pro Met Pro Asn 275 280 285Thr Leu Glu Tyr Gly Asn Arg Thr Tyr Lys
Ile Ile Asn Ala Asn Met 290 295 300Thr Trp Tyr Ala Ala Ile Lys Thr
Cys Leu Met His Lys Ala Gln Leu305 310 315 320Val Ser Ile Thr Asp
Gln Tyr His Gln Ser Phe Leu Thr Val Val Leu 325 330 335Asn Arg Leu
Gly Tyr Ala His Trp Ile Gly Leu Phe Thr Thr Asp Asn 340 345 350Gly
Leu Asn Phe Asp Trp Ser Asp Gly Thr Lys Ser Ser Phe Thr Phe 355 360
365Trp Lys Asp Glu Glu Ser Ser Leu Leu Gly Asp Cys Val Phe Ala Asp
370 375 380Ser Asn Gly Arg Trp His Ser Thr Ala Cys Glu Ser Phe Leu
Gln Gly385 390 395 400Ala Ile Cys His Val Pro Pro Glu Thr Arg Gln
Ser Glu His Pro Glu 405 410 415Leu60387DNAArtificial SequenceCysR
60aagggcatct tcgtgatcca gagcgagagc ctgaagaagt gcatccaggc cggcaagagc
60gtgctgaccc tggaaaattg caagcaggcc aacaagcaca tgctgtggaa atgggtgtcc
120aaccacggcc tgttcaacat cggcggctct ggatgtctgg gcctgaattt
ctctgcccct 180gagcagcctc tgagcctgta cgagtgtgat agcaccctgg
tgtccctgag atggcggtgc 240aaccggaaga tgatcacagg ccctctgcag
tactctgtgc aggtcgccca cgacaatacc 300gtggtggcca gcagaaagta
catccacaag tggatcagct acggcagcgg cggaggcgac 360atctgtgaat
acctgcacaa ggatctg 38761129PRTArtificial SequenceCysR 61Lys Gly Ile
Phe Val Ile Gln Ser Glu Ser Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly
Lys Ser Val Leu Thr Leu Glu Asn Cys Lys Gln Ala Asn Lys 20 25 30His
Met Leu Trp Lys Trp Val Ser Asn His Gly Leu Phe Asn Ile Gly 35 40
45Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala Pro Glu Gln Pro Leu
50 55 60Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser Leu Arg Trp Arg
Cys65 70 75 80Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr Ser Val
Gln Val Ala 85 90 95His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr Ile
His Lys Trp Ile 100 105 110Ser Tyr Gly Ser Gly Gly Gly Asp Ile Cys
Glu Tyr Leu His Lys Asp 115 120 125Leu621026DNAArtificial
SequenceCysR-FNII-CTLD1 62aagggcatct tcgtgatcca gagcgagagc
ctgaagaagt gcatccaggc cggcaagagc 60gtgctgaccc tggaaaattg caagcaggcc
aacaagcaca tgctgtggaa atgggtgtcc 120aaccacggcc tgttcaacat
cggcggctct ggatgtctgg gcctgaattt ctctgcccct 180gagcagcctc
tgagcctgta cgagtgtgat agcaccctgg tgtccctgag atggcggtgc
240aaccggaaga tgatcacagg ccctctgcag tactctgtgc aggtcgccca
cgacaatacc 300gtggtggcca gcagaaagta catccacaag tggatcagct
acggcagcgg cggaggcgac 360atctgtgaat acctgcacaa ggatctgcac
accatcaagg gcaacaccca cggaatgccc 420tgcatgttcc cgtttcagta
caaccaccag tggcaccacg agtgcaccag agaaggcaga 480gaggacgacc
tgctttggtg cgccacaacc agcagatacg agcgggatga gaagtggggc
540ttctgccctg atcctacctc tgccgaagtg ggctgcgata ccatctggga
gaaagacctg 600aacagccaca tctgctacca gttcaacctg ctgtccagcc
tgtcttggag cgaggcccac 660agcagctgtc aaatgcaagg cggcacactg
ctgagcatca ccgacgagac agaggaaaac 720ttcatccgcg agcacatgag
cagcaagacc gtggaagtgt ggatgggact gaaccagctg 780gatgagcatg
ccggatggca gtggagtgat ggcacccctc tgaactacct gaactggtcc
840cctgaagtga acttcgagcc cttcgtggaa gatcactgcg gcaccttcag
cagcttcatg 900cccagcgctt ggagaagcag agactgcgag agcaccctgc
cttacatctg caagaagtac 960ctgaaccaca tcgaccacga gatcgtggaa
aaggacgcct ggaagtacta cgccacacac 1020tgcgag 102663342PRTArtificial
SequenceCysR-FNII-CTLD1 63Lys Gly Ile Phe Val Ile Gln Ser Glu Ser
Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly Lys Ser Val Leu Thr Leu Glu
Asn Cys Lys Gln Ala Asn Lys 20 25 30His Met Leu Trp Lys Trp Val Ser
Asn His Gly Leu Phe Asn Ile Gly 35 40 45Gly Ser Gly Cys Leu Gly Leu
Asn Phe Ser Ala Pro Glu Gln Pro Leu 50 55 60Ser Leu Tyr Glu Cys Asp
Ser Thr Leu Val Ser Leu Arg Trp Arg Cys65 70 75 80Asn Arg Lys Met
Ile Thr Gly Pro Leu Gln Tyr
Ser Val Gln Val Ala 85 90 95His Asp Asn Thr Val Val Ala Ser Arg Lys
Tyr Ile His Lys Trp Ile 100 105 110Ser Tyr Gly Ser Gly Gly Gly Asp
Ile Cys Glu Tyr Leu His Lys Asp 115 120 125Leu His Thr Ile Lys Gly
Asn Thr His Gly Met Pro Cys Met Phe Pro 130 135 140Phe Gln Tyr Asn
His Gln Trp His His Glu Cys Thr Arg Glu Gly Arg145 150 155 160Glu
Asp Asp Leu Leu Trp Cys Ala Thr Thr Ser Arg Tyr Glu Arg Asp 165 170
175Glu Lys Trp Gly Phe Cys Pro Asp Pro Thr Ser Ala Glu Val Gly Cys
180 185 190Asp Thr Ile Trp Glu Lys Asp Leu Asn Ser His Ile Cys Tyr
Gln Phe 195 200 205Asn Leu Leu Ser Ser Leu Ser Trp Ser Glu Ala His
Ser Ser Cys Gln 210 215 220Met Gln Gly Gly Thr Leu Leu Ser Ile Thr
Asp Glu Thr Glu Glu Asn225 230 235 240Phe Ile Arg Glu His Met Ser
Ser Lys Thr Val Glu Val Trp Met Gly 245 250 255Leu Asn Gln Leu Asp
Glu His Ala Gly Trp Gln Trp Ser Asp Gly Thr 260 265 270Pro Leu Asn
Tyr Leu Asn Trp Ser Pro Glu Val Asn Phe Glu Pro Phe 275 280 285Val
Glu Asp His Cys Gly Thr Phe Ser Ser Phe Met Pro Ser Ala Trp 290 295
300Arg Ser Arg Asp Cys Glu Ser Thr Leu Pro Tyr Ile Cys Lys Lys
Tyr305 310 315 320Leu Asn His Ile Asp His Glu Ile Val Glu Lys Asp
Ala Trp Lys Tyr 325 330 335Tyr Ala Thr His Cys Glu
34064165DNAArtificial SequenceCD8 alpha hinge domain 64ttcgtgccgg
tcttcctgcc agcgaagcca accacgacgc cagcaccgcg accaccaaca 60cctgcgccca
ccatcgcgtc gcagcccctg tccctgcgcc cagaggcgtg cagaccagca
120gcggggggcg cagtgcacac gagggggctg gacttcgcct gtgat
1656560DNAArtificial SequencePLA2R Signal peptide 65atgctgctga
gccctagcct gctgctgctc ctgcttcttg gagcccctag aggatgtgcc
6066126DNAArtificial Sequence4-1BB intracellular domain
66aagcgcggtc gcaagaaact gctctatatt tttaaacagc cattcatgag acctgtccag
60accactcaag aggaggacgg atgttcctgt agatttcctg aagaggaaga gggggggtgc
120gagctg 1266757DNAArtificial SequencePLA2R pro-peptide
67ggatctgaag gtgttgccgc cgctctgaca cccgagagac tgctggaatg gcaggac
576830DNAArtificial SequenceGS linker 68ggtggcggag gttctggagg
tggaggttcc 306910DNAArtificial SequenceGS linker 69ggggsggggs
1070479PRTArtificial SequenceExtracellular domains of Construct
4027.CF12 70Lys Gly Ile Phe Val Ile Gln Ser Glu Ser Leu Lys Lys Cys
Ile Gln1 5 10 15Ala Gly Lys Ser Val Leu Thr Leu Glu Asn Cys Lys Gln
Ala Asn Lys 20 25 30His Met Leu Trp Lys Trp Val Ser Asn His Gly Leu
Phe Asn Ile Gly 35 40 45Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala
Pro Glu Gln Pro Leu 50 55 60Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val
Ser Leu Arg Trp Arg Cys65 70 75 80Asn Arg Lys Met Ile Thr Gly Pro
Leu Gln Tyr Ser Val Gln Val Ala 85 90 95His Asp Asn Thr Val Val Ala
Ser Arg Lys Tyr Ile His Lys Trp Ile 100 105 110Ser Tyr Gly Ser Gly
Gly Gly Asp Ile Cys Glu Tyr Leu His Lys Asp 115 120 125Leu His Thr
Ile Lys Gly Asn Thr His Gly Met Pro Cys Met Phe Pro 130 135 140Phe
Gln Tyr Asn His Gln Trp His His Glu Cys Thr Arg Glu Gly Arg145 150
155 160Glu Asp Asp Leu Leu Trp Cys Ala Thr Thr Ser Arg Tyr Glu Arg
Asp 165 170 175Glu Lys Trp Gly Phe Cys Pro Asp Pro Thr Ser Ala Glu
Val Gly Cys 180 185 190Asp Thr Ile Trp Glu Lys Asp Leu Asn Ser His
Ile Cys Tyr Gln Phe 195 200 205Asn Leu Leu Ser Ser Leu Ser Trp Ser
Glu Ala His Ser Ser Cys Gln 210 215 220Met Gln Gly Gly Thr Leu Leu
Ser Ile Thr Asp Glu Thr Glu Glu Asn225 230 235 240Phe Ile Arg Glu
His Met Ser Ser Lys Thr Val Glu Val Trp Met Gly 245 250 255Leu Asn
Gln Leu Asp Glu His Ala Gly Trp Gln Trp Ser Asp Gly Thr 260 265
270Pro Leu Asn Tyr Leu Asn Trp Ser Pro Glu Val Asn Phe Glu Pro Phe
275 280 285Val Glu Asp His Cys Gly Thr Phe Ser Ser Phe Met Pro Ser
Ala Trp 290 295 300Arg Ser Arg Asp Cys Glu Ser Thr Leu Pro Tyr Ile
Cys Lys Lys Tyr305 310 315 320Leu Asn His Ile Asp His Glu Ile Val
Glu Lys Asp Ala Trp Lys Tyr 325 330 335Tyr Ala Thr His Cys Glu Pro
Gly Trp Asn Pro Tyr Asn Arg Asn Cys 340 345 350Tyr Lys Leu Gln Lys
Glu Glu Lys Thr Trp His Glu Ala Leu Arg Ser 355 360 365Cys Gln Ala
Asp Asn Ser Ala Leu Ile Asp Ile Thr Ser Leu Ala Glu 370 375 380Val
Glu Phe Leu Val Thr Leu Leu Gly Asp Glu Asn Ala Ser Glu Thr385 390
395 400Trp Ile Gly Leu Ser Ser Asn Lys Ile Pro Val Ser Phe Glu Trp
Ser 405 410 415Asn Asp Ser Ser Val Ile Phe Thr Asn Trp His Thr Leu
Glu Pro His 420 425 430Ile Phe Pro Asn Arg Ser Gln Leu Cys Val Ser
Ala Glu Gln Ser Glu 435 440 445Gly His Trp Lys Val Lys Asn Cys Glu
Glu Arg Leu Phe Tyr Ile Cys 450 455 460Lys Lys Ala Gly His Val Leu
Ser Asp Ala Glu Ser Gly Cys Gln465 470 47571629PRTArtificial
SequenceExtracellular domains of Construct 4028.CF123 71Lys Gly Ile
Phe Val Ile Gln Ser Glu Ser Leu Lys Lys Cys Ile Gln1 5 10 15Ala Gly
Lys Ser Val Leu Thr Leu Glu Asn Cys Lys Gln Ala Asn Lys 20 25 30His
Met Leu Trp Lys Trp Val Ser Asn His Gly Leu Phe Asn Ile Gly 35 40
45Gly Ser Gly Cys Leu Gly Leu Asn Phe Ser Ala Pro Glu Gln Pro Leu
50 55 60Ser Leu Tyr Glu Cys Asp Ser Thr Leu Val Ser Leu Arg Trp Arg
Cys65 70 75 80Asn Arg Lys Met Ile Thr Gly Pro Leu Gln Tyr Ser Val
Gln Val Ala 85 90 95His Asp Asn Thr Val Val Ala Ser Arg Lys Tyr Ile
His Lys Trp Ile 100 105 110Ser Tyr Gly Ser Gly Gly Gly Asp Ile Cys
Glu Tyr Leu His Lys Asp 115 120 125Leu His Thr Ile Lys Gly Asn Thr
His Gly Met Pro Cys Met Phe Pro 130 135 140Phe Gln Tyr Asn His Gln
Trp His His Glu Cys Thr Arg Glu Gly Arg145 150 155 160Glu Asp Asp
Leu Leu Trp Cys Ala Thr Thr Ser Arg Tyr Glu Arg Asp 165 170 175Glu
Lys Trp Gly Phe Cys Pro Asp Pro Thr Ser Ala Glu Val Gly Cys 180 185
190Asp Thr Ile Trp Glu Lys Asp Leu Asn Ser His Ile Cys Tyr Gln Phe
195 200 205Asn Leu Leu Ser Ser Leu Ser Trp Ser Glu Ala His Ser Ser
Cys Gln 210 215 220Met Gln Gly Gly Thr Leu Leu Ser Ile Thr Asp Glu
Thr Glu Glu Asn225 230 235 240Phe Ile Arg Glu His Met Ser Ser Lys
Thr Val Glu Val Trp Met Gly 245 250 255Leu Asn Gln Leu Asp Glu His
Ala Gly Trp Gln Trp Ser Asp Gly Thr 260 265 270Pro Leu Asn Tyr Leu
Asn Trp Ser Pro Glu Val Asn Phe Glu Pro Phe 275 280 285Val Glu Asp
His Cys Gly Thr Phe Ser Ser Phe Met Pro Ser Ala Trp 290 295 300Arg
Ser Arg Asp Cys Glu Ser Thr Leu Pro Tyr Ile Cys Lys Lys Tyr305 310
315 320Leu Asn His Ile Asp His Glu Ile Val Glu Lys Asp Ala Trp Lys
Tyr 325 330 335Tyr Ala Thr His Cys Glu Pro Gly Trp Asn Pro Tyr Asn
Arg Asn Cys 340 345 350Tyr Lys Leu Gln Lys Glu Glu Lys Thr Trp His
Glu Ala Leu Arg Ser 355 360 365Cys Gln Ala Asp Asn Ser Ala Leu Ile
Asp Ile Thr Ser Leu Ala Glu 370 375 380Val Glu Phe Leu Val Thr Leu
Leu Gly Asp Glu Asn Ala Ser Glu Thr385 390 395 400Trp Ile Gly Leu
Ser Ser Asn Lys Ile Pro Val Ser Phe Glu Trp Ser 405 410 415Asn Asp
Ser Ser Val Ile Phe Thr Asn Trp His Thr Leu Glu Pro His 420 425
430Ile Phe Pro Asn Arg Ser Gln Leu Cys Val Ser Ala Glu Gln Ser Glu
435 440 445Gly His Trp Lys Val Lys Asn Cys Glu Glu Arg Leu Phe Tyr
Ile Cys 450 455 460Lys Lys Ala Gly His Val Leu Ser Asp Ala Glu Ser
Gly Cys Gln Glu465 470 475 480Gly Trp Glu Arg His Gly Gly Phe Cys
Tyr Lys Ile Asp Thr Val Leu 485 490 495Arg Ser Phe Asp Gln Ala Ser
Ser Gly Tyr Tyr Cys Pro Pro Ala Leu 500 505 510Val Thr Ile Thr Asn
Arg Phe Glu Gln Ala Phe Ile Thr Ser Leu Ile 515 520 525Ser Ser Val
Val Lys Met Lys Asp Ser Tyr Phe Trp Ile Ala Leu Gln 530 535 540Asp
Gln Asn Asp Thr Gly Glu Tyr Thr Trp Lys Pro Val Gly Gln Lys545 550
555 560Pro Glu Pro Val Gln Tyr Thr His Trp Asn Thr His Gln Pro Arg
Tyr 565 570 575Ser Gly Gly Cys Val Ala Met Arg Gly Arg His Pro Leu
Gly Arg Trp 580 585 590Glu Val Lys His Cys Arg His Phe Lys Ala Met
Ser Leu Cys Lys Gln 595 600 605Pro Val Glu Asn Gln Glu Lys Ala Glu
Tyr Glu Glu Arg Trp Pro Phe 610 615 620His Pro Cys Tyr
Leu62572336DNAArtificial SequenceCD3zeta intracellular domain
72agagtgaagt tcagcaggag cgcagacgcc cccgcgtaca agcagggcca gaaccagctc
60tataacgagc tcaatctagg acgaagagag gagtacgatg ttttggacaa gagacgtggc
120cgggaccctg agatgggggg aaagccgaga aggaagaacc ctcaggaagg
cctgtacaat 180gaactgcaga aagataagat ggcggaggcc tacagtgaga
ttgggatgaa aggcgagcgc 240cggaggggca aggggcacga tggcctttac
cagggtctca gtacagccac caaggacacc 300tacgacgccc ttcacatgca
ggccctgccc cctcgc 336
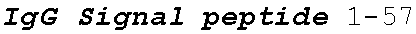




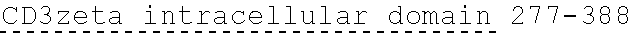
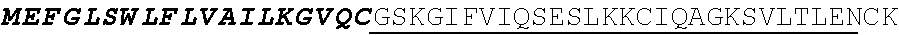






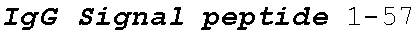


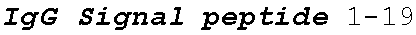
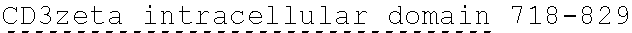
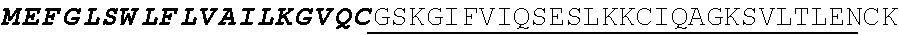

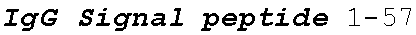
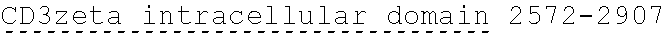


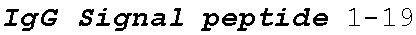




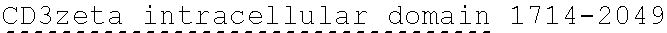


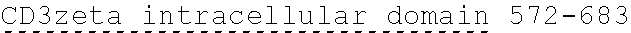
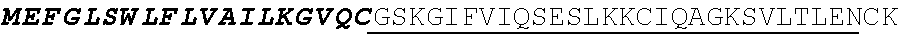




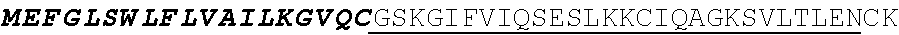

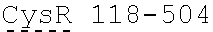
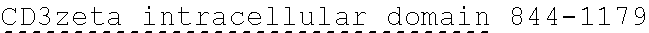





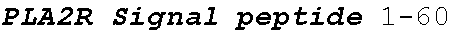
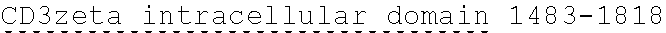


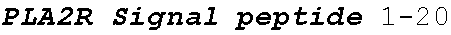
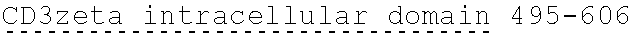
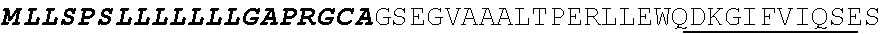

D00000
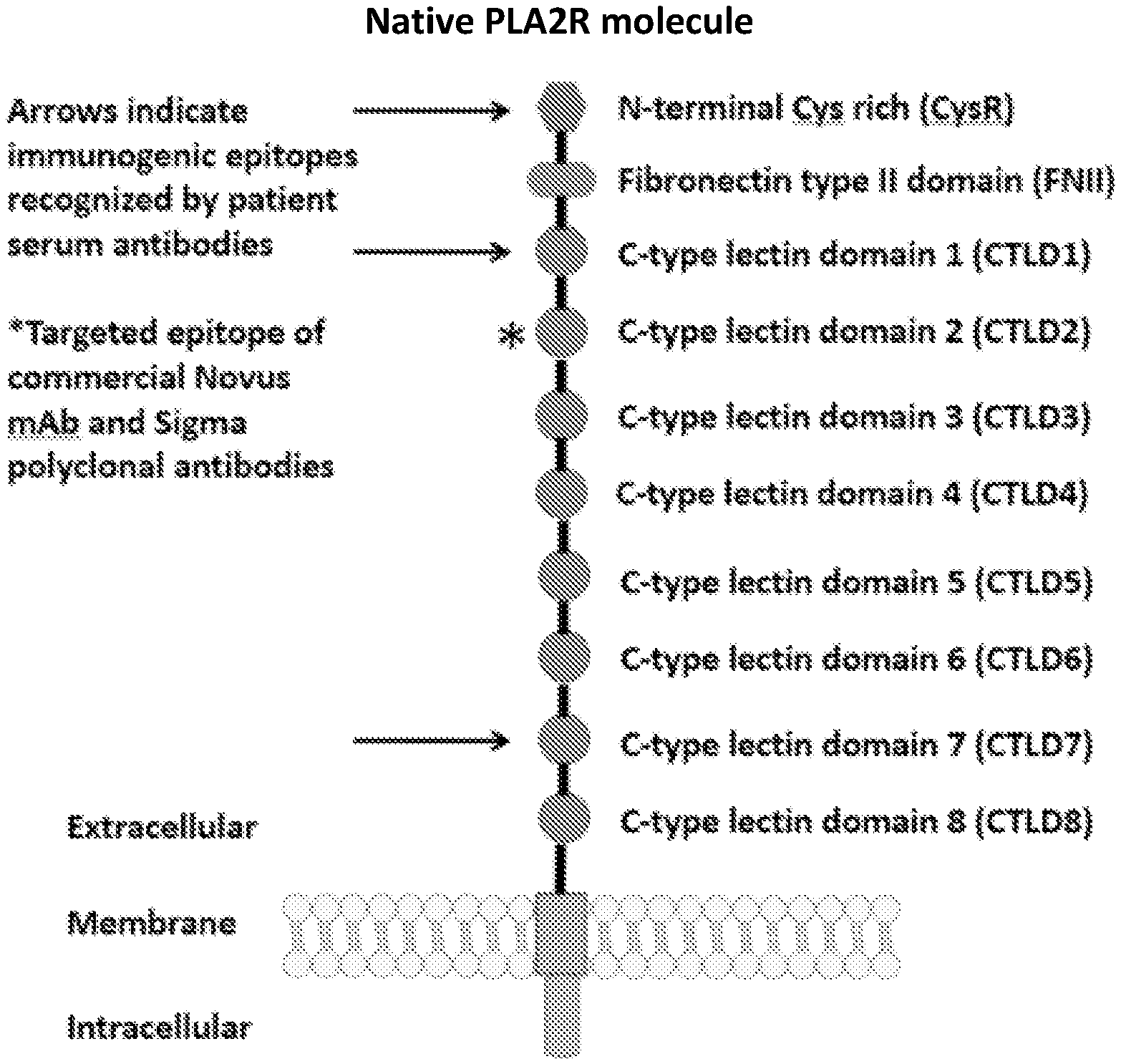
D00001
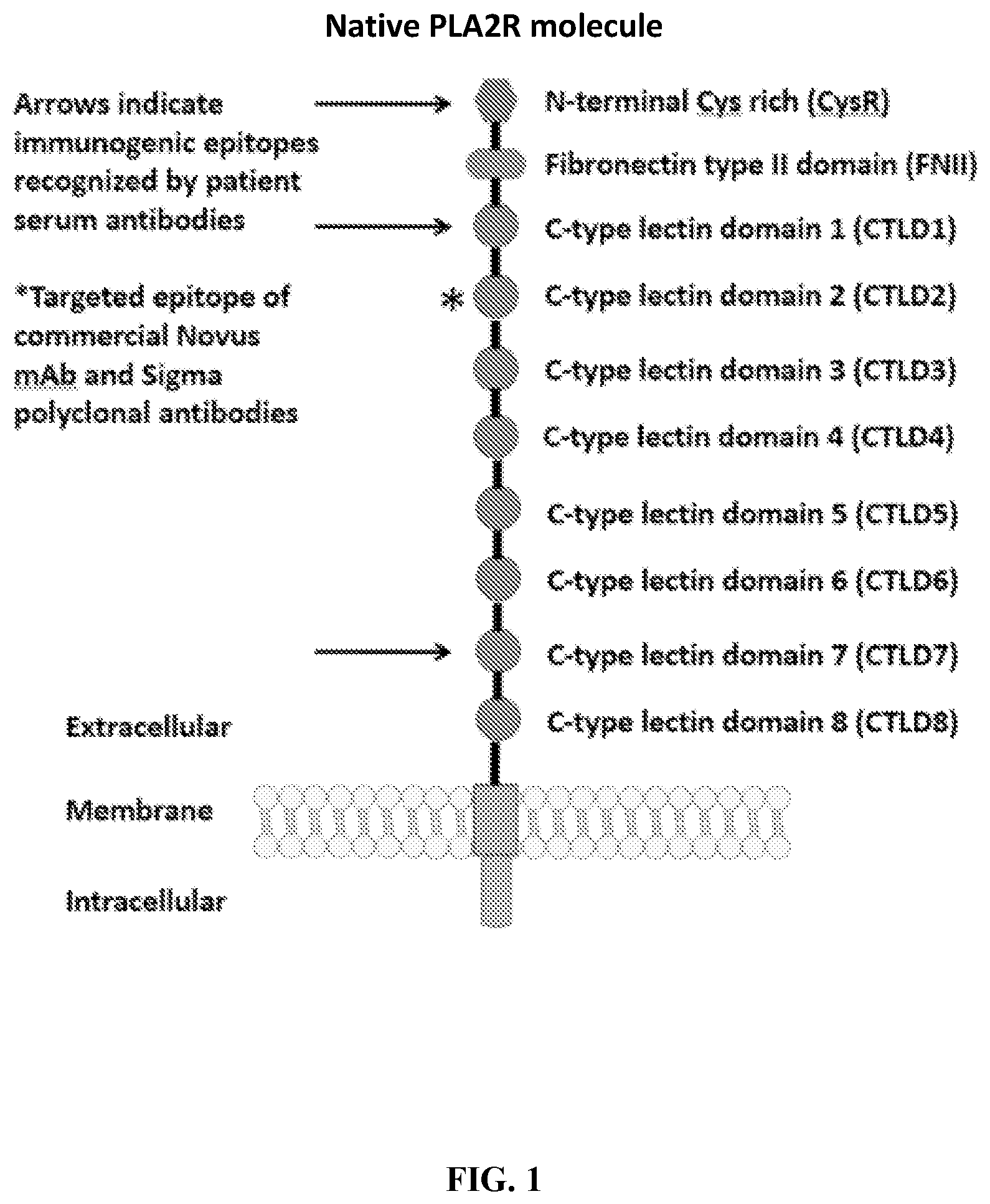
D00002

D00003
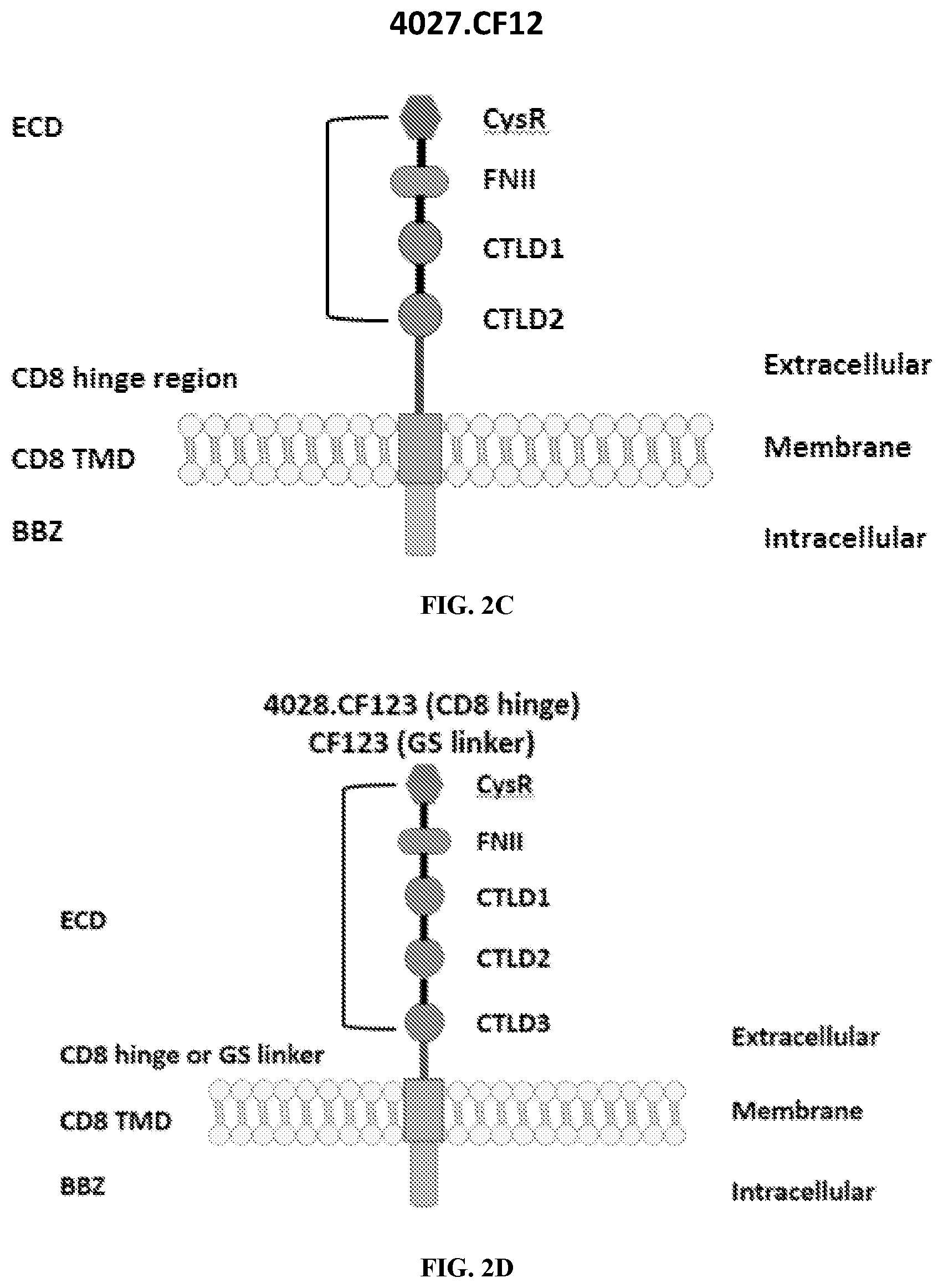
D00004
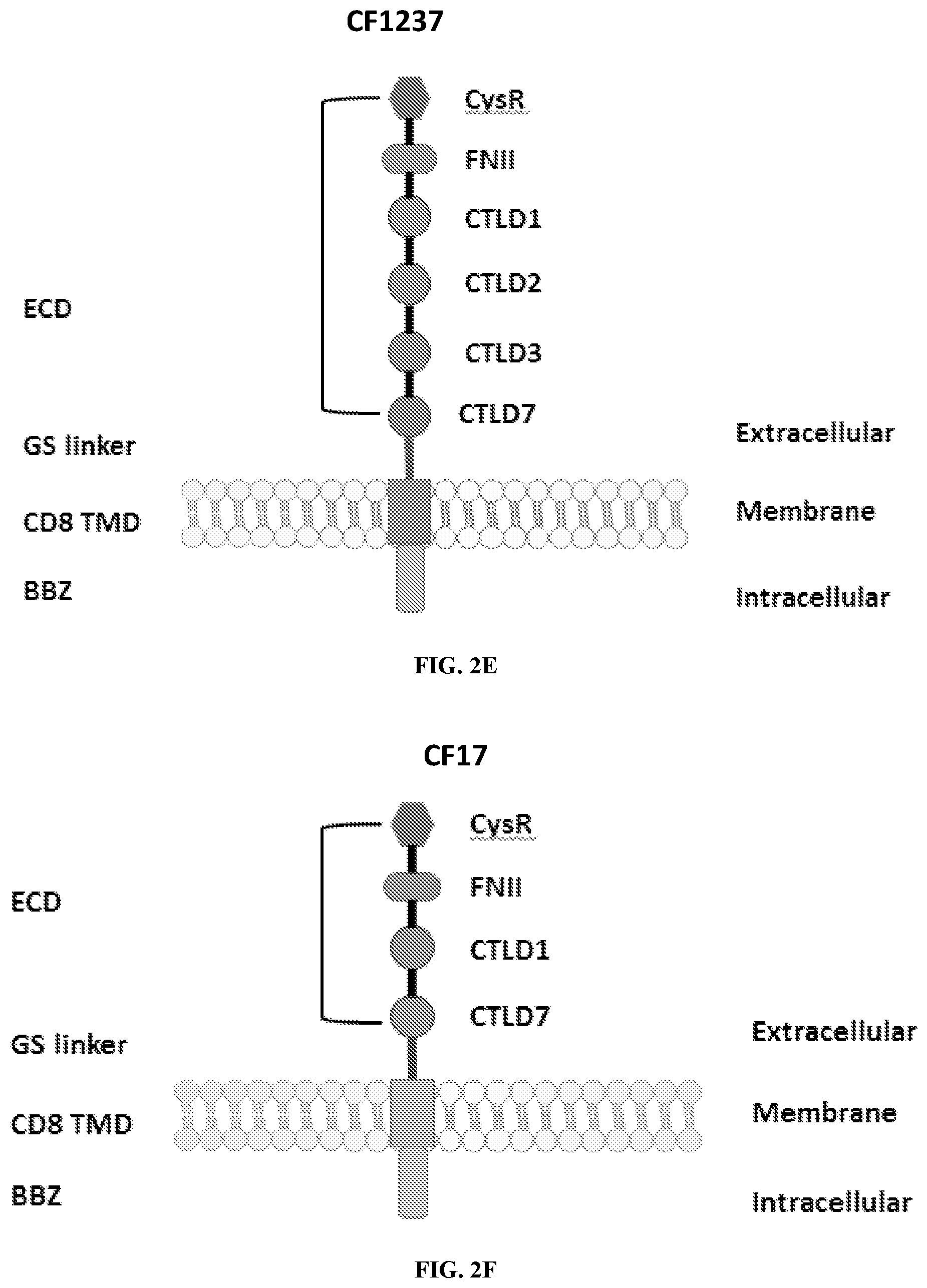
D00005
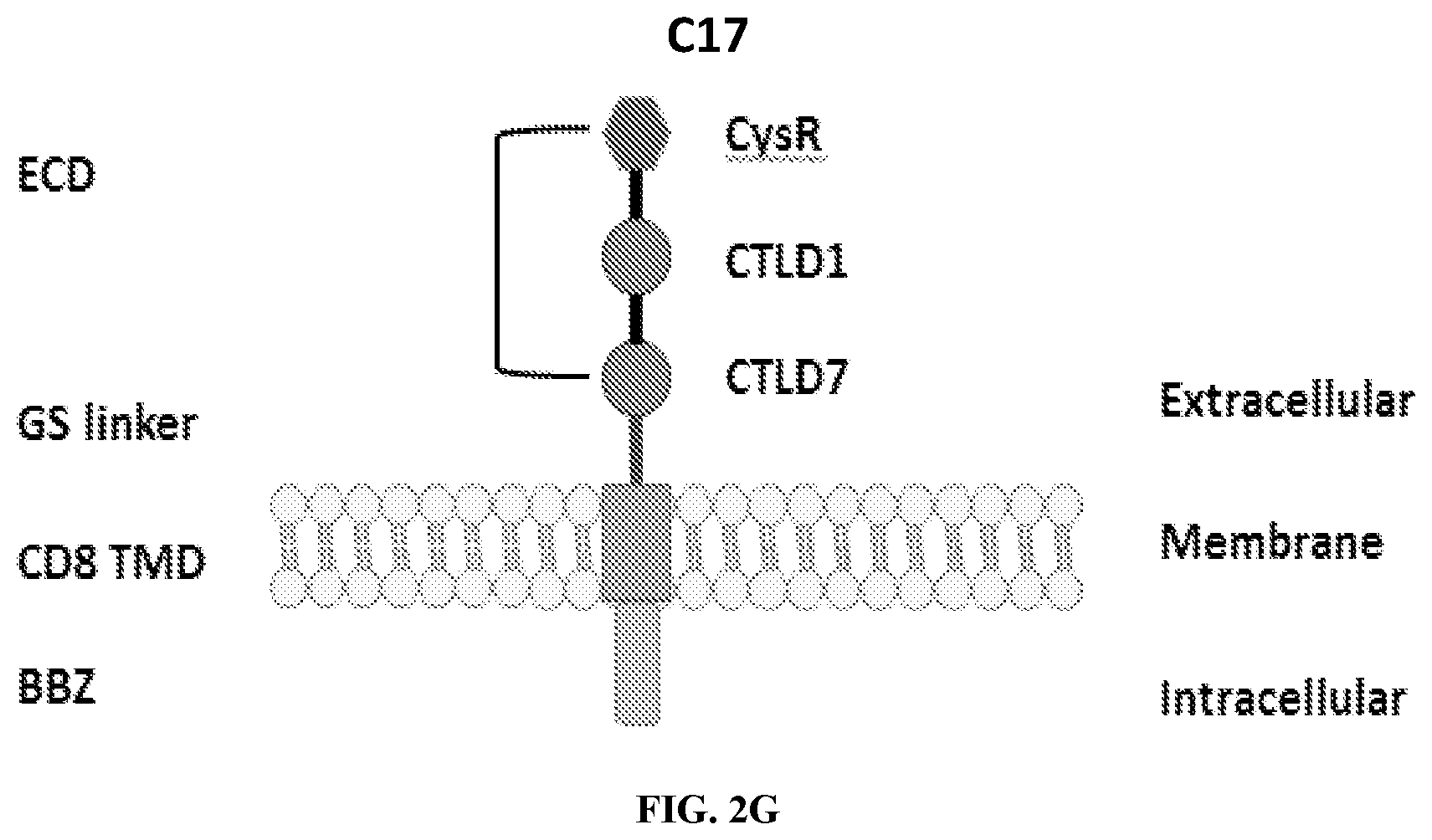
D00006
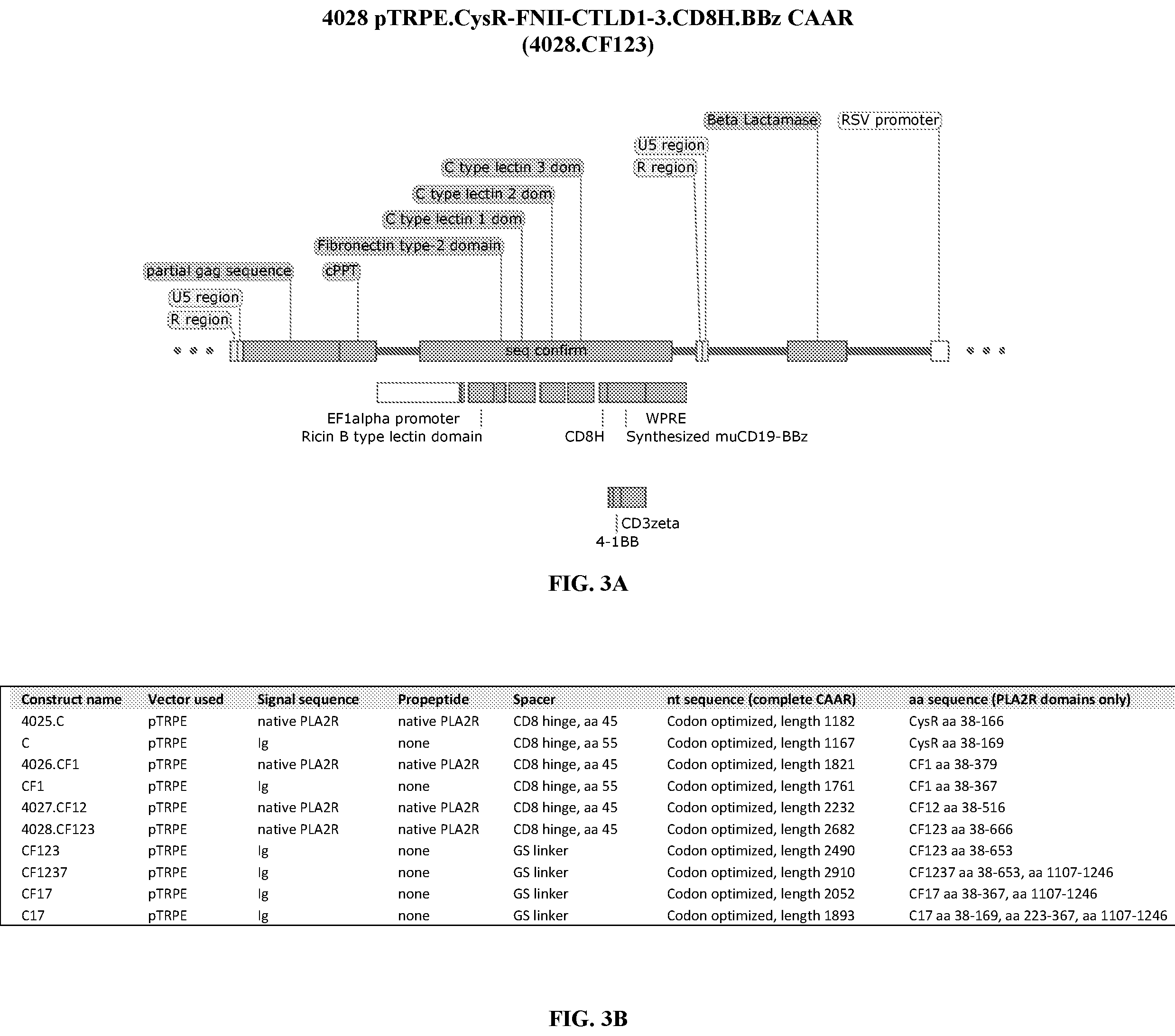
D00007
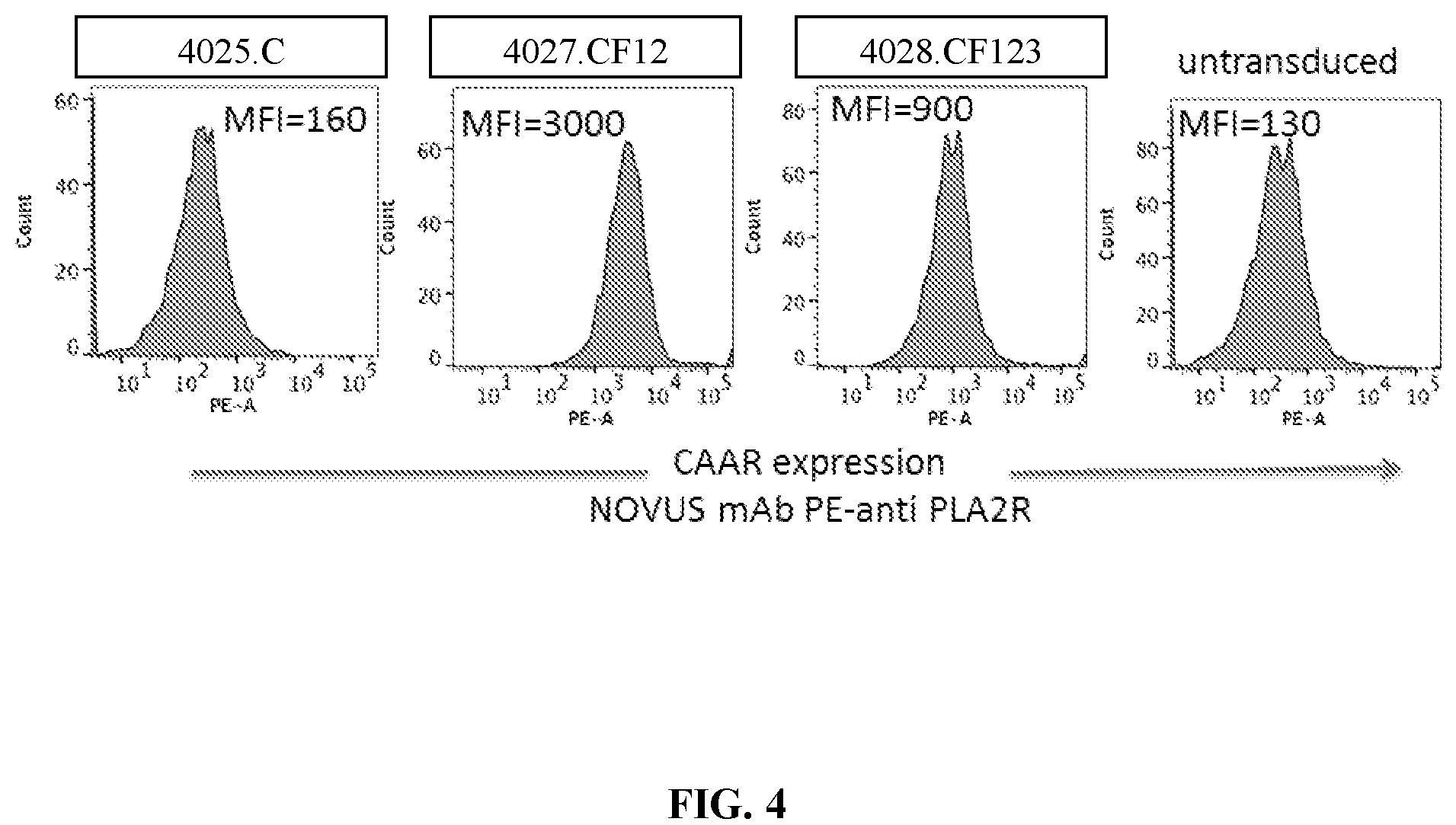
D00008
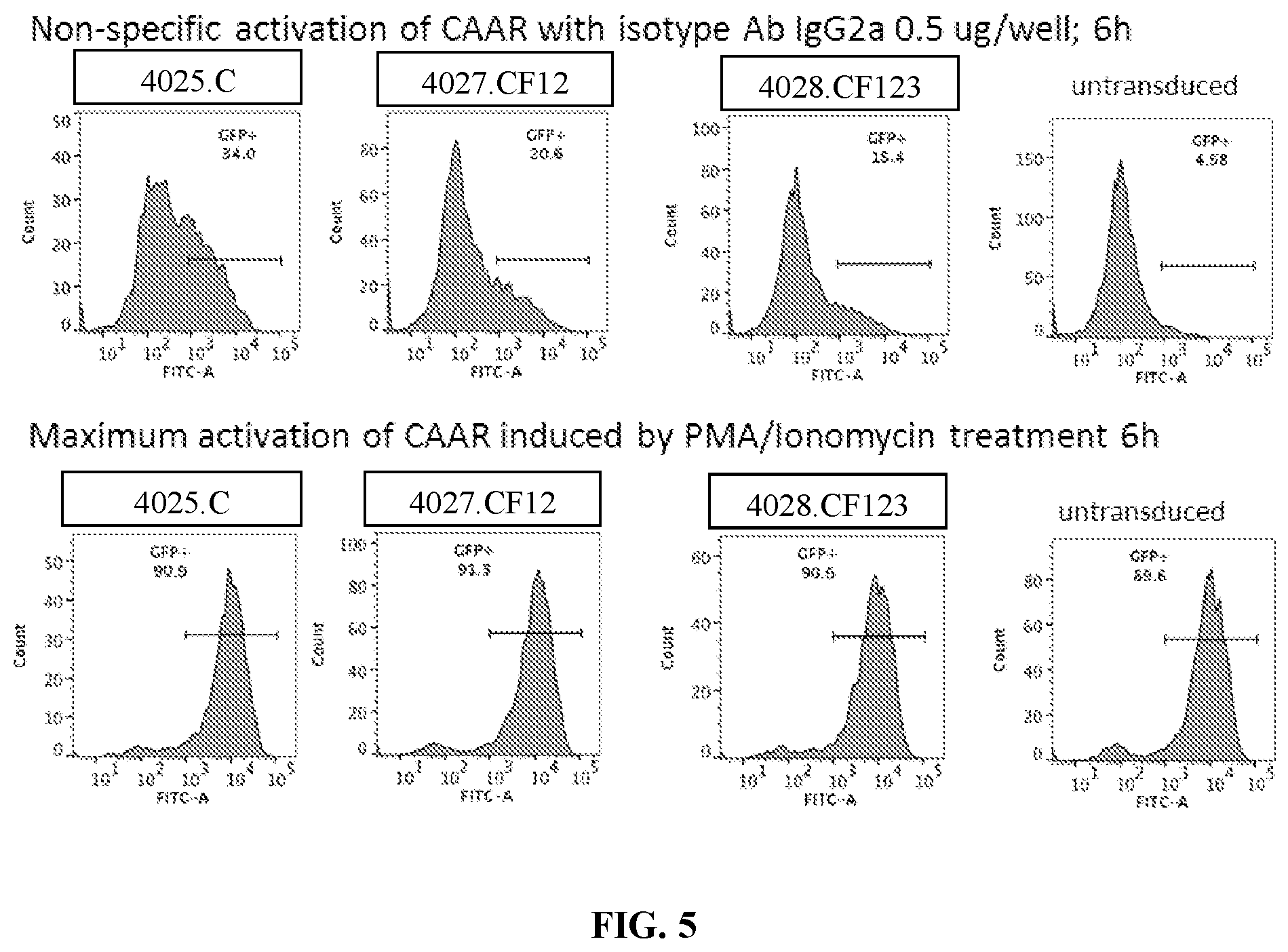
D00009
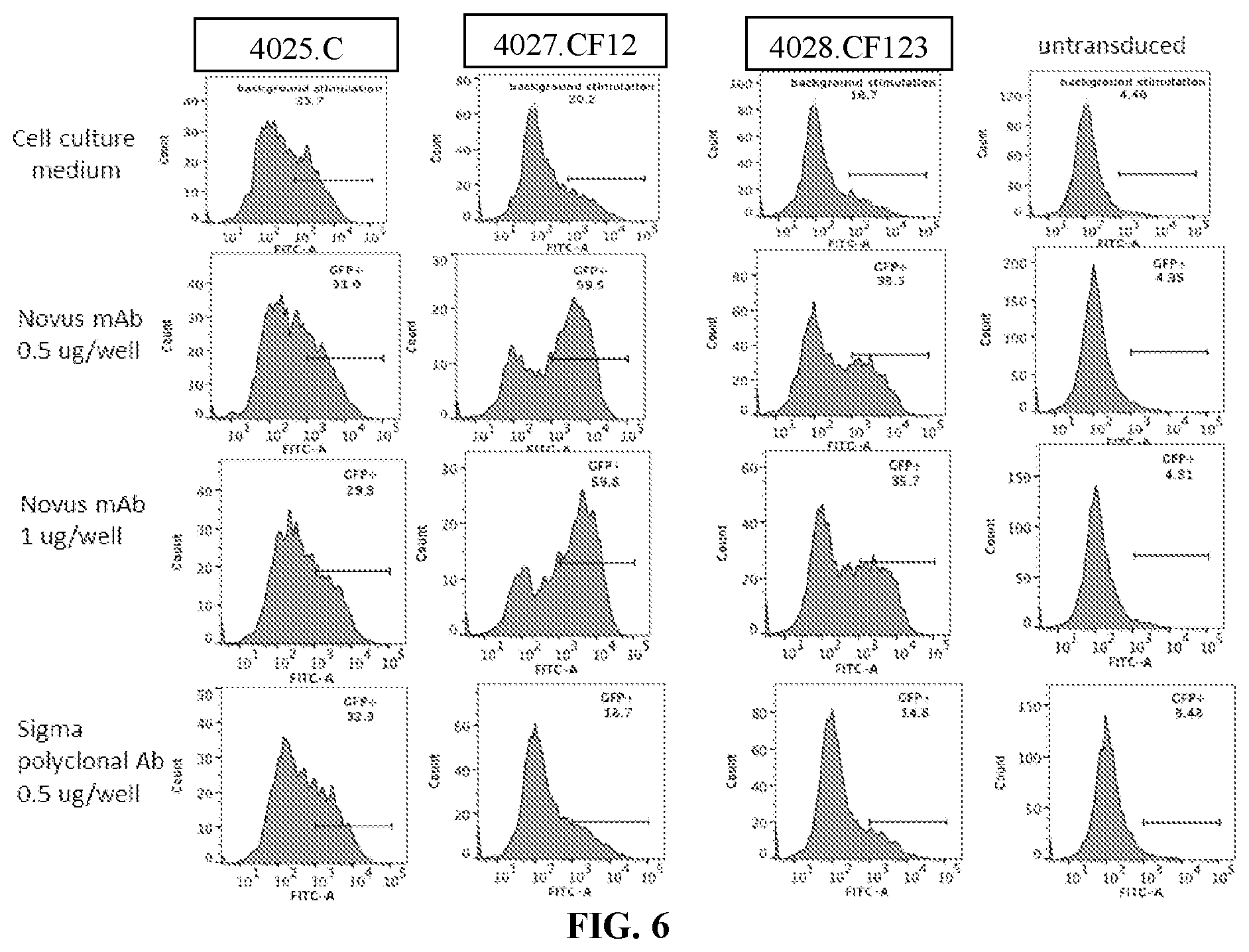
D00010

D00011
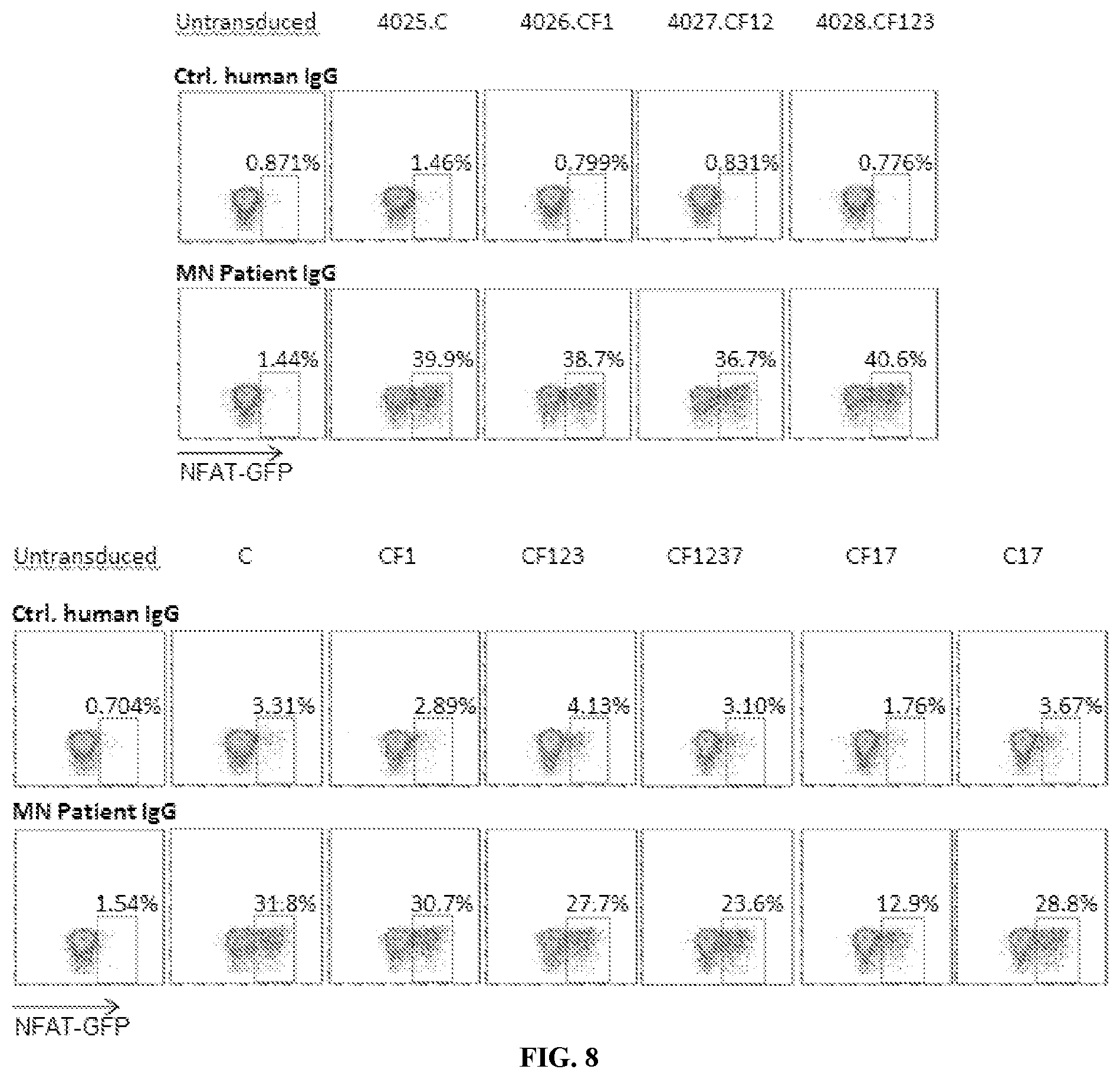
D00012
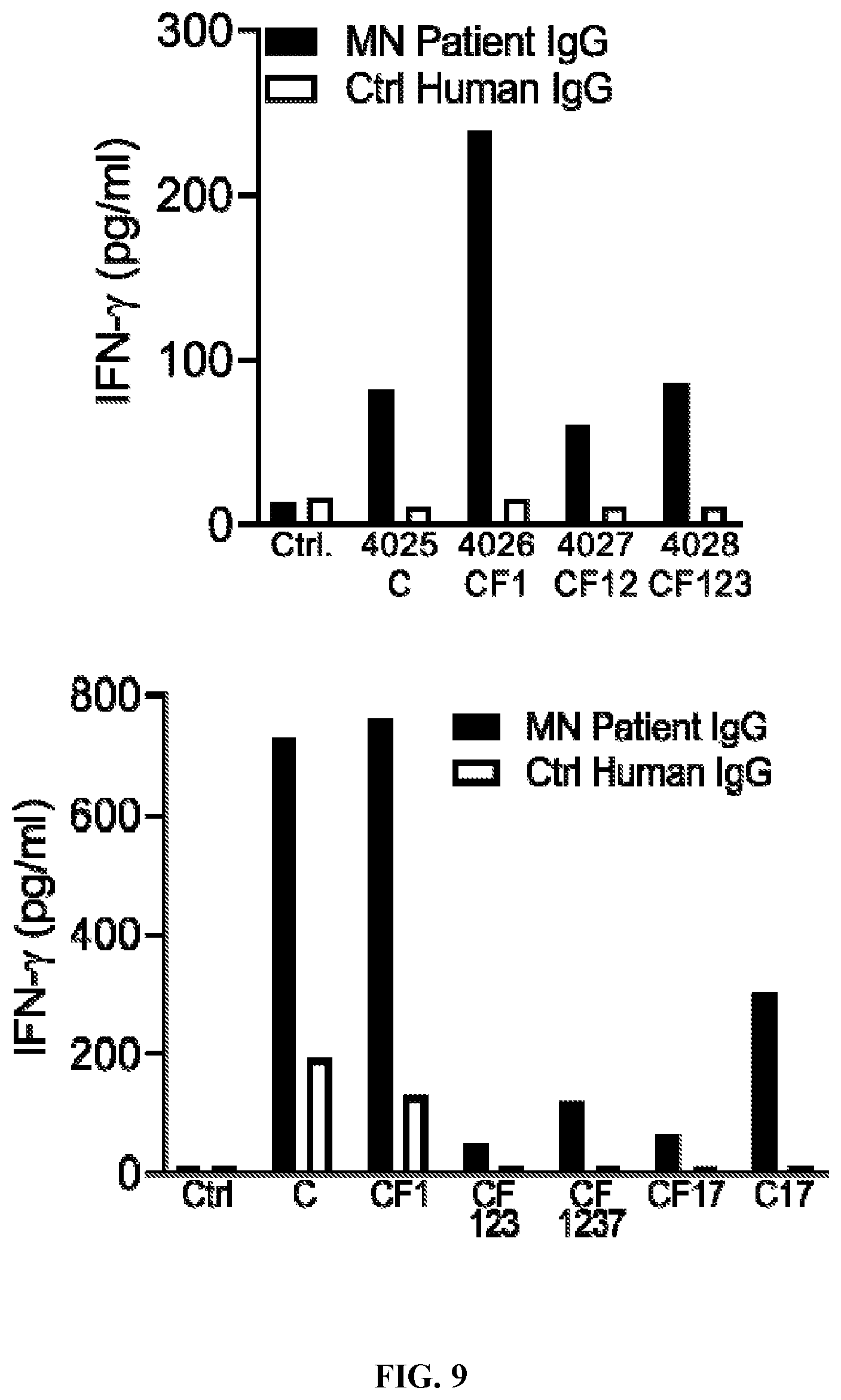
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.