Peptides Having Anti-Inflammatory Properties
Jaynes; Jesse ; et al.
U.S. patent application number 17/028662 was filed with the patent office on 2021-03-18 for peptides having anti-inflammatory properties. The applicant listed for this patent is Riptide Bioscience, Inc.. Invention is credited to Charles Garvin, Jesse Jaynes, Henry Wilfred Lopez, George R. Martin, Clayton Yates.
| Application Number | 20210077566 17/028662 |
| Document ID | / |
| Family ID | 1000005248135 |
| Filed Date | 2021-03-18 |












View All Diagrams
| United States Patent Application | 20210077566 |
| Kind Code | A1 |
| Jaynes; Jesse ; et al. | March 18, 2021 |
Peptides Having Anti-Inflammatory Properties
Abstract
Aspects of the present invention relate to peptides having anti-inflammatory activity, compositions containing one or more of the peptides, and use of the peptides to treat conditions associated with excessive inflammation in animals, particularly humans and other mammals.
| Inventors: | Jaynes; Jesse; (Auburn, AL) ; Lopez; Henry Wilfred; (Napa, CA) ; Martin; George R.; (Rockville, MD) ; Yates; Clayton; (Auburn, AL) ; Garvin; Charles; (Redwood City, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005248135 | ||||||||||
| Appl. No.: | 17/028662 | ||||||||||
| Filed: | September 22, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 16169819 | Oct 24, 2018 | |||
| 17028662 | ||||
| 15286491 | Oct 5, 2016 | 10149886 | ||
| 16169819 | ||||
| 14882293 | Oct 13, 2015 | 9492499 | ||
| 15286491 | ||||
| 62063909 | Oct 14, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 5/08 20130101; A61K 31/337 20130101; C07K 7/08 20130101; A61K 45/06 20130101; A61K 36/10 20130101; A61K 31/4745 20130101; A61K 38/04 20130101; C07K 7/06 20130101; A61K 38/08 20130101; A61K 38/16 20130101; A61K 31/282 20130101; A61K 31/513 20130101; C07K 5/10 20130101; A61K 31/519 20130101; A61K 31/7068 20130101; C07K 14/00 20130101; A61K 38/10 20130101 |
| International Class: | A61K 38/08 20060101 A61K038/08; A61K 38/16 20060101 A61K038/16; C07K 14/00 20060101 C07K014/00; C07K 7/08 20060101 C07K007/08; C07K 7/06 20060101 C07K007/06; C07K 5/10 20060101 C07K005/10; C07K 5/08 20060101 C07K005/08; A61K 38/10 20060101 A61K038/10; A61K 45/06 20060101 A61K045/06; A61K 31/7068 20060101 A61K031/7068; A61K 38/04 20060101 A61K038/04; A61K 31/282 20060101 A61K031/282; A61K 31/337 20060101 A61K031/337; A61K 31/4745 20060101 A61K031/4745; A61K 31/513 20060101 A61K031/513; A61K 31/519 20060101 A61K031/519; A61K 36/10 20060101 A61K036/10 |
Claims
1.-34. (canceled)
35. A method of modulating macrophage activity, the method comprising: contacting a macrophage with a CD206-binding agent to modulate activity of the macrophage.
36. The method according to claim 35, wherein the CD206-binding agent binds to a mannose-binding site to modulate binding of signal regulatory protein (SIRP)-mannose to CD206.
37. The method according to claim 35, wherein the CD206-binding agent binds to CD206 with a binding energy of at least -650 kcal/mol.
38. The method according to claim 35, wherein the macrophage activity that is modulated is macrophage polarization.
39. The method according to claim 35, wherein viability of the macrophage is reduced.
40. The method according to claim 35, wherein the macrophage is a M2 macrophage or a tumor associated macrophage (TAM).
41. The method according to claim 35, wherein the CD206-binding agent inhibits macrophage activity.
42. The method according to claim 35, wherein the CD206-binding agent is an anti-inflammatory peptide.
43. The method according to claim 35, wherein the macrophage is in vitro.
44. The method according to claim 35, wherein the macrophage is in vivo.
45. A method of treating a subject for a condition associated with chronic inflammation, the method comprising: administering an effective amount of a CD206-binding agent to the subject to treat the subject for the condition associated with chronic inflammation.
46. The method according to claim 45, wherein the condition associated with chronic inflammation is selected from the group consisting of scleroderma or multiple sclerosis, irritable bowel disease, ulcerative colitis, colitis, Crohn's disease, idiopathic pulmonary fibrosis, asthma, keratitis, arthritis, osteoarthritis, rheumatoid arthritis, auto-immune diseases, a feline or human immunodeficiency virus (FIV or HIV) infection, cancer, age-related inflammation and/or stem cell dysfunction, graft-versus-host disease (GVHD), keloids, obesity, diabetes, diabetic wounds, other chronic wounds, atherosclerosis, Parkinson's disease, Alzheimer's disease, macular degeneration, gout, gastric ulcers, gastritis, mucositis, toxoplasmosis, and chronic viral or microbial infections.
47. The method according to claim 45, wherein the CD206-binding agent is administered in conjunction with another drug known to be effective in treating the condition.
48. The method according to claim 45, wherein the condition is cancer.
49. The method according to claim 48, further comprising administering an effective amount of a chemotherapeutic agent or cell therapy to the subject.
50. The method according to claim 49, wherein the chemotherapeutic agent or cell therapy is selected from steroids, anthracyclines, thyroid hormone replacement drugs, thymidylate-targeted drugs, checkpoint inhibitor drugs, Chimeric Antigen Receptor/T cell therapies, and other cell therapies.
51. The method according to claim 45, wherein the condition associated with chronic inflammation is a fibrosis or scleroderma.
52. The method according to claim 45, wherein the CD206-binding agent is an immunomodulatory peptide of 18 amino acid residues or less in length.
53. The method according to claim 52, wherein the peptide comprises a sequence defined by one of the formulae: [Y.sub.1aY.sub.1b]-[X.sub.1aX.sub.1b]-[Y.sub.2aY.sub.2b]-[X.sub.2aX.sub.2- b]-[Y.sub.3a]-[X.sub.3a]; and [X.sub.3a]-[Y.sub.3a]-[X.sub.2bX.sub.2a]-[Y.sub.2bY.sub.2a]-[X.sub.1bX.su- b.1a]-[Y.sub.1bY.sub.1a]; wherein: Y.sub.1a, Y.sub.1b, Y.sub.2a, Y.sub.2b and Y.sub.3a are each phenylalanine; and X.sub.1a, X.sub.1b, X.sub.2a, X.sub.2b and X.sub.3a are each independently selected from lysine and arginine.
54. The method according to claim 46, wherein the CD206-binding agent is an immunomodulatory peptide of 18 amino acid residues or less in length, wherein the peptide comprises a sequence selected from SEQ ID NO:121-124, SEQ ID NO:148, SEQ ID NO: 106, SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 109, SEQ ID NO: 110, SEQ ID NO: 111, SEQ ID NO: 112, SEQ ID NO: 113, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 116, SEQ ID NO: 117, SEQ ID NO: 118, and SEQ ID NO: 120.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] Pursuant to 35 U.S.C. .sctn. 119 (e), this application claims the benefit of priority to U.S. Provisional Patent Application Ser. No. 62/063,909, filed Oct. 14, 2014, the disclosure of which application is hereby incorporated by reference herein in its entirety.
FIELD OF THE INVENTION
[0002] Aspects of the present invention relate to peptides having anti-inflammatory activity, compositions containing one or more of the peptides, and use of the peptides to treat conditions associated with excessive inflammation in animals, particularly humans and other mammals.
BACKGROUND OF THE INVENTION
[0003] Under normal conditions, inflammation is a process that helps an animal recover from injury. Acute inflammation is the initial response of a tissue to harmful stimuli. It involves a complex, highly regulated process that begins when cells present in the injured tissue, including macrophages, dendritic cells, histiocytes, Kupffer cells, and mastocytes, sense molecules associated with the injury and become activated. Upon activation, these cells release inflammatory mediators, such as vasodilators. The vasodilators induce increased blood flow and permeability of the blood vessels in the vicinity of the injury. This, in turn, results in the increased movement of plasma and leukocytes (including neutrophils and macrophages) from the blood into the injured tissue. Because inflammatory mediators are, in general, rapidly degraded, acute inflammation requires constant stimulation in order to be sustained. As a result, acute inflammation ends once the harmful stimulus is removed.
[0004] Various agents, including but not limited to bacteria, viruses, physical injury, chemical injury, cancer, chemotherapy, and radiation therapy, can, depending on the specific agent and the genetic makeup of the animal exposed to it, cause prolonged and excessive inflammation. Such inflammation, known as chronic inflammation, is believed to be a contributing factor to many widespread and debilitating diseases, including heart disease, cancer, respiratory disease, stroke, neurological diseases such as Alzheimer's disease, diabetes, and kidney disease. The result of chronic inflammation is the destruction of normal tissue and its replacement with collagen-rich connective tissue. Collagen-rich connective tissue, also known as scar tissue, exhibits diminished tissue function as compared to normal tissue. Persistent and prolonged formation of scar tissue, in turn, leads to fibrosis. Fibrosis is among the common symptoms of diseases affecting the lungs, skin, liver, heart, and bone marrow, and is a critical factor in diseases such as idiopathic pulmonary fibrosis, scleroderma, keloids, liver cirrhosis, myocardial fibrosis, diabetic kidney disease, myelodysplastic syndrome, and other disorders.
[0005] Studies of chronic inflammation and fibrosis have indicated that, regardless of the activating agent and the tissue affected, a common network of signaling proteins tend to function together to establish the pro-inflammatory state. This network of signaling proteins includes a number of different cytokines, cytokine receptors, transcription factors, and micro RNAs, including TGF.beta., TGF.beta.RII, and miRNA19b.
[0006] Despite growing knowledge about conditions that involve excessive inflammation, such as chronic inflammation and fibrosis, treatments for such conditions remain elusive. Many drugs and other substances have been shown to have anti-inflammatory activity, either in vitro or in vivo, but for many indications caused or potentiated by inflammation, there are still no therapies. In addition, many anti-inflammatory therapies are associated with harmful side effects. Thus, there remains a critical need to identify therapeutic agents that reduce inflammation without harmful side effects.
SUMMARY OF THE INVENTION
[0007] The present invention is based, in part, on the discovery of novel peptides that have powerful anti-inflammatory activities in vitro and in vivo. The present invention is also based, in part, on the discovery that peptides of the invention specifically bind to key functional regions on one or more signaling proteins, particularly pro-inflammatory cytokines, macrophage inhibition proteins, and histone regulation proteins. The present invention is also based, in part, on the discovery that the peptides of the invention are sufficiently stable in the circulation to allow for intravenous administration.
[0008] Accordingly, in one aspect, the invention provides a composition comprising an anti-inflammatory polypeptide. In certain embodiments, the anti-inflammatory polypeptide is 3 to 24 amino acids residues in length and includes a striapathic region consisting of alternating hydrophobic and hydrophilic modules. In certain embodiments, each hydrophilic module is made up of a sequence of one or more (e.g., 1-5, 1-4, 1-3) hydrophilic amino acid residues. In certain embodiments, each hydrophobic module is made up of a sequence of one or more (e.g., 1-5, 1-4, 1-3) hydrophobic amino acid residues.
[0009] In certain embodiments, the striapathic region of an anti-inflammatory peptide includes m hydrophilic modules and n hydrophobic modules, with m and n each being a positive integer. For example, in certain embodiments, the striapathic region includes two hydrophilic modules and two hydrophobic modules (2:2), two hydrophilic modules and three hydrophobic modules (2:3), three hydrophilic modules and two hydrophobic modules (3:2), three hydrophilic modules and three hydrophobic modules (3:3), three hydrophilic modules and four hydrophobic modules (3:4), or four hydrophilic modules and three hydrophobic modules (4:3).
[0010] In certain embodiments, the striapathic region of an anti-inflammatory polypeptide is at least 5, 6, 7, 8, 9, or 10 amino acid residues in length. In preferred embodiments, the length of the striapathic region is between 7 and 12 amino acid residues. In certain embodiments, the striapathic region makes up at least 25% of the length of the polypeptide. For example, in certain embodiments, the striapathic region comprises at least 30%, 35%, 40%, 45%, 50%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% of the length of the polypeptide.
[0011] In certain embodiments, the striapathic region of an anti-inflammatory polypeptide adopts a helical secondary structure. Examples of helical secondary structures include 3.sub.10-helices, .alpha.-helices, .pi.-helices, and poly-proline helices. In other embodiments, the striapathic region of an anti-inflammatory polypeptide adopts a beta-strand secondary structure. In preferred embodiments, the striapathic region of an anti-inflammatory polypeptides has an amphipathic conformation.
[0012] In certain embodiments, an anti-inflammatory polypeptide comprises, consists essentially of, or consists of a striapathic region having a sequence that conforms to any one of the structural formulas disclosed herein (e.g., any one of Formulas I-LIII). In certain embodiments, the anti-inflammatory polypeptide is one of the polypeptides listed in Tables 3-9. In other embodiments, the anti-inflammatory polypeptide has at least 70%, 80%, or 90% homology with any one of the polypeptides disclosed in Tables 3-9.
[0013] In certain embodiments, an anti-inflammatory polypeptide binds to at least one signaling protein. In preferred embodiments, the anti-inflammatory polypeptide binds to at least one signaling protein in vitro and/or in vivo, with sufficient affinity to modulate the activity of the signaling protein. Examples of signaling proteins that the anti-inflammatory polypeptides bind to include proteins that function as pro-inflammatory cytokines, proteins that inhibit macrophage activity, or protein that regulate histone function. In certain embodiments, the anti-inflammatory polypeptide binds to a protein target selected from the group consisting of NFkB class II proteins (e.g., Rel A, Rel B, cRel, NF-kB1, and NF-kB2), TGF.beta., Notch receptors (e.g., Notch1), Wnt receptors (e.g., Wnt8R), TRAIL, EGFR, interleukin receptors (e.g., IL6R, IL10R), cyclin dependent kinases (e.g., CDK6), CD47, SIRP-.alpha., transglutaminases (e.g., TGM2), LEGUMAIN, CD209, FAS, programmed cell death protein 1 (PD-1/CD279), mitogen-activated protein kinase kinase 7 (MKK7), ribonucleotide reductase (RNR), and histone methyl transferase. In preferred embodiments, the anti-inflammatory polypeptide binds to two, three, four, or more such signaling proteins. For example, in certain embodiments, an anti-inflammatory polypeptide binds to an NF-kB Class II protein (e.g., RelB) and at least one other signaling protein that functions as a pro-inflammatory cytokine, an inhibitor of macrophage activity, or a regulator of histone function. In preferred embodiments, the anti-inflammatory polypeptide binds to the NF-kB Class II protein and at least one other protein target, with sufficient binding affinity to each target to modulate the activity of both targets in vivo. In preferred embodiments, an anti-inflammatory polypeptide binds to the dimerization site of an NFkB Class II protein (e.g., RelB).
[0014] In certain embodiments, an anti-inflammatory polyeptides binds to a carrier protein in the blood (e.g., serum albumin).
[0015] In certain embodiments, an anti-inflammatory polypeptide is modified to include, for example, a linker, a carbohydrate, a lipid, or a polymer (e.g., PEG). In certain embodiments, a first anti-inflammatory polypeptide is linked to a second anti-inflammatory polypeptide so as to form a multimer, such as a dimer. In certain embodiments, the dimer is a homodimer. In other embodiments, the dimer is a heterodimer. In certain embodiments, the linker is a peptide linker. In preferred embodiments, the peptide linker forms a peptide bond with the C-terminus of the first anti-inflammatory polypeptide and a peptide bond with the N-terminus of the second anti-inflammatory polypeptide. In certain embodiments, the linker is a biodegradeable linker. In certain embodiments, the linker is a disulfide bond. In certain embodiments, the disulfide linkage is formed by a pair of cysteine residues (e.g., one cysteine residue from each of the polypeptides being linked).
[0016] In certain embodiments, the anti-inflammatory polypeptide is linked to a molecule other than another anti-inflammatory polypeptide. For example, the anti-inflammatory polypeptide can be linked to a label or a chemotherapeutic agent. In certain embodiments, the linker is a biodegradable linker. In certain embodiments, the linker is a di-sulfide bond (e.g., involving the sulfhydryl group of a cysteine residue located at the C-terminus or N-terminus of the anti-inflammatory polypeptide).
[0017] In another aspect, the invention provides pharmaceutical compositions that comprise an anti-inflammatory polypeptide and a pharmaceutically acceptable carrier. In certain embodiments, the pharmaceutical composition comprises a single type of anti-inflammatory polypeptide. In other embodiments, the pharmaceutical composition comprises a combination of two or more anti-inflammatory polypeptides. In preferred embodiments, the pharmaceutical composition is substantially free of blood proteins and/or metabolites found in the blood. In other embodiments, the pharmaceutical composition includes serum albumin (e.g., human serum albumin). In preferred embodiments, any serum albumin present in a pharmaceutical composition is recombinantly produced and/or substantially free of other blood proteins and/or metabolites found in the blood. In certain embodiments, the pharmaceutical composition comprises 1 mg to 1000 mg (e.g., 10 to 400 mg, 20 to 300 mg, or about 25 to 250 mg) of an anti-inflammatory polypeptide.
[0018] In another aspect, the invention provides methods of treating a subject by administering to the subject a composition (e.g., a pharmaceutical composition) comprising an anti-inflammatory polypeptide. In certain embodiments, the subject is an animal, such as a mammal (e.g., a human). In certain embodiments, the subject has elevated levels of inflammatory cytokines, is suffering from a chronic inflammatory condition, or is likely to develop a chronic inflammatory condition. In certain embodiments, the chronic inflammatory condition can be irritable bowel disease, ulcerative colitis, colitis, Crohn's disease, fibrosis, idiopathic pulmonary fibrosis, asthma, keratitis, arthritis, osteoarthritis, rheumatoid arthritis, an auto-immune disease, a feline or human immunodeficiency virus (FIV or HIV) infection, or cancer. In certain embodiments, the cancer is colon cancer, breast cancer, leukemia, lymphoma, ovarian cancer, prostate cancer, liver cancer, lung cancer, testicular cancer, cervical cancer, bladder cancer, endometrial cancer, kidney cancer, melanoma, or a cancer of the thyroid or brain. In certain embodiments, the composition is administered in combination with a chemotherapeutic agent, immunotherapeutic agent, and/or radiation therapy.
[0019] These and other features and advantages of the compositions and methods of the invention will be set forth or will become more fully apparent in the description that follows and in the appended claims. For example, suitable anti-inflammatory polypeptides may be identified by use of the Structural Algorithm described herein. Furthermore, features and advantages of the described compositions and methods may be learned by practicing the methods or will be obvious from the description.
BRIEF SUMMARY OF THE DRAWINGS
[0020] FIG. 1 depicts a structural model of human RelB, an NF-kB Class II protein.
[0021] FIG. 2 depicts a structural model of human RelB bound by RP-182.
[0022] FIG. 3 depicts a structural model of human RelB bound by RP-166.
[0023] FIG. 4 depicts a structural model of human RelB bound by RP-113.
[0024] FIG. 5 depicts a structural model of human RelB bound by RP-387.
[0025] FIG. 6 depicts a structural model of human RelB bound by RP-289.
[0026] FIG. 7 depicts a structural model of human RelB bound by NF-Contr2.
[0027] FIG. 8 depicts a structural model of human RelB bound by NF-Contr3.
[0028] FIG. 9 depicts structural models of polypeptides RP-182, RP-166, RP-113, and RP-289, with each model showing the polar and non-polar facial arc associated with the helices formed by the polypeptides.
[0029] FIG. 10 depicts structural models of polypeptides RP-387, NF-Contr2, and NF-Contr3, with each model showing the polar and non-polar amino acid residues. The facial arc associated with the helix formed by RP-387 is also shown.
[0030] FIG. 11 depicts a structural model of the binding pocket of the RelB dimerization domain.
[0031] FIG. 12 depicts a structural model of the binding pocket of the RelB dimerization domain bound by RP-183.
[0032] FIG. 13 depicts a structural model of histone methyl transferase enzyme bound by RP-182.
[0033] FIG. 14 depicts structural models of a CD47 dimer (left panel) and a CD47 dimer bound by RP-183.
[0034] FIG. 15 depicts structural models of a SIRP-.alpha. dimer (left panel) and a SIRP-.alpha. dimer bound by RP-183.
[0035] FIG. 16 depicts structural models of CD206 (left side) and CD206 bound by RP-182 (right side).
[0036] FIG. 17 depicts structural models of TGM2 (left side) and TGM2 bound by RP-182 (right side).
[0037] FIG. 18 depicts a structural model of human serum albumin bound by RP-183.
[0038] FIG. 19 shows PD-1-stained tumor cells from p53/KRAS mice treated with vehicle only (left panel) or treated with RP-182 (right panel). PD-1 expression is reduced in RP-182 treated mice.
[0039] FIG. 20 shows PD-L1-stained (left panels) and PD-L2-stained (right panels) tumor cells from p53/KRAS mice treated with vehicle only (top panel in each set) or treated with RP-182 (bottom panel in each set). PD-L1 and PD-L2 expression is reduced in RP-182 treated mice.
[0040] FIG. 21 shows MDA-MB-231 tumor volume in four cohorts of mice over time. Cohort 1: vehicle; Cohort 2: Gemcitabine treated; Cohort 3: RP-182 treated; Cohort 4: RP-182+Gemcitabine treated.
[0041] FIG. 22 shows C42B tumor volume in four cohorts of mice over time. Cohort 1: vehicle; Cohort 2: Docetaxel treated; Cohort 3: RP-182 treated; Cohort 4: RP-182+Docetaxel treated.
DETAILED DESCRIPTION OF THE INVENTION
[0042] The following description supplies specific details in order to provide a thorough understanding of the present invention. That said, to avoid obscuring aspects of the described anti-inflammatory polypeptides and related methods of treating a subject, well-known structures, materials, processes, techniques, and operations are not shown or described in detail. Additionally, the skilled artisan will understand that the described anti-inflammatory polypeptides and related methods of treating a subject can be implemented and used without employing these specific details. Indeed, the described anti-inflammatory polypeptides and methods can be placed into practice by modifying the illustrated polypeptides, compositions, and methods, and can be used in conjunction with other treatments, apparatuses, and techniques conventionally used in the industry.
[0043] As discussed above, the invention disclosed herein relates to immune-modulatory polypeptides, particularly peptides that have immunosuppressive properties, and methods of administering such immune-modulatory polypeptides to a subject, particularly a subject suffering from a medical condition associated with persistent inflammation or at risk developing such a medical condition.
[0044] The invention provides anti-inflammatory polypeptides, sometimes referred to as "RP peptides," that satisfy the requirements of the Structural Algorithm described below. The invention also provides anti-inflammatory polypeptides that share a minimum degree of homology with any of the exemplary RP peptides disclosed herein. Thus, a peptide or polypeptide of the invention is an anti-inflammatory polypeptide that satisfies the Structural Algorithm described below or shares a minimum degree of homology with any of the exemplary RP peptides disclosed herein (e.g., in Tables 3-9).
[0045] The terms "peptide" and "polypeptide" are used synonymously herein to refer to polymers constructed from amino acid residues.
[0046] The term "amino acid residue," as used herein, refers to any naturally occurring amino acid (L or D form), non-naturally occurring amino acid, or amino acid mimetic (such as a peptoid monomer).
[0047] The "length" of a polypeptide is the number of amino acid residues linked end-to-end that constitute the polypeptide, excluding any non-peptide linkers and/or modifications that the polypeptide may contain.
[0048] The term "striapathic region," as used herein, refers to an alternating sequence of hydrophobic and hydrophilic modules. A "hydrophobic module" is made up of a peptide sequence consisting of one to five hydrophobic amino acid residues. Likewise, a hydrophilic module is made up of a peptide sequence consisting of one to five hydrophilic amino acid residues.
[0049] Hydrophobic amino acid residues are characterized by a functional group ("side chain") that has predominantly non-polar chemical properties. Such hydrophobic amino acid residues can be naturally occurring (L or D form) or non-naturally occurring. Alternatively, hydrophobic amino acid residues can be amino acid mimetics characterized by a functional group ("side chain") that has predominantly non-polar chemical properties. Conversely, hydrophilic amino acid residues are characterized by a functional group ("side chain") that has predominantly polar (charged or uncharged) chemical properties. Such hydrophilic amino acid residues can be naturally occurring (L or D form) or non-naturally occurring. Alternatively, hydrophilic amino acid residues can be amino acid mimetics characterized by a functional group ("side chain") that has predominantly polar (charged or uncharged) chemical properties. Examples of hydrophilic and hydrophobic amino acid residues are shown in Table 1, below. Suitable non-naturally occurring amino acid residues and amino acid mimetics are known in the art. See, e.g., Liang et al. (2013), "An Index for Characterization of Natural and Non-Natural Amino Acids for Peptidomimetics," PLoS ONE 8(7):e67844.
[0050] Although most amino acid residues can be considered as either hydrophobic or hydrophilic, a few, depending on their context, can behave as either hydrophobic or hydrophilic. For example, due to their relatively weak non-polar characteristics, glycine, proline, and/or cysteine can sometimes function as hydrophilic amino acid residues. Conversely, due to their bulky, slightly hydrophobic side chains, histidine and arginine can sometimes function as hydrophobic amino acid residues.
TABLE-US-00001 TABLE 1 Hydrophobic and Hydrophilic Amino Acid Residues Hydrophilic Residues Hydrophobic Residues (X) (Y) Arginine Tryptophan Histidine Phenylalanine Lysine Tyrosine Aspartic Acid Isoleucine Glutamic Acid Leucine Asparagine Valine Glutamine Methionine Pyrrolysine Cysteine Threonine Serine Alanine Proline Glycine Selenocysteine N-formylmethionine Norleucine Norvaline
[0051] The term "anti-inflammatory property," as used herein, refers to any property of a polypeptide that can be evaluated in silico, in vitro, and/or in vivo, that reduces or inhibits, or would be expected to reduce or inhibit, a pro-inflammatory signal mediated by a protein target and/or reduces or inhibits inflammation in a subject.
[0052] Structural Algorithm
[0053] In its most basic form, the Structural Algorithm requires an anti-inflammatory peptide to have the following characteristics:
[0054] a length of 3 to 24 amino acid residues;
[0055] a striapathic region that comprises at least 25% of the length of the polypeptide; and
[0056] at least one anti-inflammatory property.
[0057] The anti-inflammatory peptide and/or its striapathic region can have a length that is greater than 3 amino acid residues and/or less than 24 amino acid residues. Thus, the requisite length of the polypeptide can be, for example, 3 to 20, 3 to 18, 3 to 16, 3 to 14, 3 to 12, 4 to 20, 4 to 18, 4 to 16, 4 to 14, 4 to 12, 5 to 20, 5 to 18, 5 to 16, 5 to 14, 5 to 12, 6 to 20, 6 to 18, 6 to 16, 6 to 14, 6 to 12, 7 to 20, 7 to 18, 7 to 16, 7 to 14, or in certain embodiments 7 to 12 amino acid residues. For an anti-inflammatory polypeptide that is longer than 12 amino acid residues, it can be advantageous to design a kink in the secondary structure (e.g., such as produced by a proline residue) such that the polypeptide has a striapathic region that is 12 or fewer amino acid residues in length. The striapathic region of an anti-inflammatory peptide can comprise at least 30%, 35%, 40%, 45%, 50%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% of the length of the polypeptide.
[0058] An anti-inflammatory polypeptide can have a striapathic region that includes at least two hydrophobic modules and one or more (e.g., two or three) hydrophilic modules. Alternatively, an anti-inflammatory polypeptide can have a striapathic region that includes at least three hydrophobic modules and two or more (e.g., three or four) hydrophilic modules; a striapathic region that includes at least two hydrophilic modules and one or more (e.g., two or three) hydrophilic modules; or a striapathic region that includes at least three hydrophilic modules and two or more (e.g., three or four) hydrophobic modules.
[0059] As discussed above, a striapathic region consists of alternating hydrophilic (X.sub.m) and hydrophobic (Y.sub.n) modules. In this context, the subscripts m and n are positive integers that identify different modules. Each X.sub.m module consists of a sequence according to the formula X.sub.ma-X.sub.mb-X.sub.mc-X.sub.md-X.sub.me. X.sub.ma is selected from the group consisting of a naturally occurring hydrophilic amino acid, a non-naturally occurring hydrophilic amino acid, and a hydrophilic amino acid mimetic; and X.sub.mb, X.sub.mc, X.sub.md and X.sub.me are each individually absent or selected from the group consisting of a naturally occurring hydrophilic amino acid, a non-naturally occurring hydrophilic amino acid, and a hydrophilic amino acid mimetic. Each Y.sub.n module consists of a sequence according to the formula Y.sub.na-Y.sub.nb-Y.sub.nc-Y.sub.nd-Y.sub.ne. Y.sub.na is selected from the group consisting of a naturally occurring hydrophobic amino acid, a non-naturally occurring hydrophobic amino acid, and a hydrophobic amino acid mimetic; Y.sub.nb, Y.sub.nc, Y.sub.nd, and Y.sub.ne are each individually absent or selected from the group consisting of a naturally occurring hydrophobic, a non-naturally occurring hydrophobic amino acid, and a hydrophobic amino acid mimetic.
[0060] In certain anti-inflammatory polypeptides, each X.sub.m module consists of a sequence according to the formula X.sub.ma-X.sub.mb-X.sub.mc-X.sub.md or X.sub.ma-X.sub.mb-X.sub.mc. Similarly, in certain anti-inflammatory polypeptides, each Y.sub.n module consists of a sequence according to the formula Y.sub.na-Y.sub.nb-Y.sub.nc-Y.sub.nd or Y.sub.na-Y.sub.nb-Y.sub.nc.
[0061] Anti-inflammatory peptides can include a striapathic region corresponding to a formula selected from the group consisting of:
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c (Formula I);
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.2a-- Y.sub.3a-X.sub.3a (Formula II);
X.sub.2a-Y.sub.3a-X.sub.3a-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-- Y.sub.2b-Y.sub.2c (Formula III);
X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-X.sub.2a-X.sub.2b-X.sub.2c (Formula IV);
Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-X.sub.2a-X.sub.2b-X.sub.2c-- Y.sub.3a-X.sub.3a (Formula V);
X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b (Formula VI);
Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y.sub.3a (Formula VII);
Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y.sub.3a-- Y.sub.3b-X.sub.3a (Formula VIII);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-- Y.sub.3a-Y.sub.3b (Formula IX);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-- Y.sub.3a-X.sub.3a (Formula X);
X.sub.1a-Y.sub.1a-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a-X.sub.3b-- Y.sub.3a-Y.sub.3b (Formula XI);
X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a-- X.sub.3b-Y.sub.3a (Formula XII);
Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-X.sub.2c-- Y.sub.3a-Y.sub.3b (Formula XIII);
X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-Y.sub.2a-- Y.sub.2b-Y.sub.2c (Formula XIV);
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-- X.sub.2b-X.sub.2c (Formula XV);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-Y.sub.2b-X.sub.2a-- X.sub.2b-Y.sub.3a (Formula XVI);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b (Formula XVII);
X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a (Formula XVIII);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-Y.sub.3a-- Y.sub.3b-X.sub.3a (Formula XIX);
X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-Y.sub.2a-Y.sub.2b-X.sub.3a-X.sub.3b-- Y.sub.3a-Y.sub.3b (Formula XX);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-X.sub.2a-X.sub.2b-Y.sub.3a-- Y.sub.3b (Formula XXI);
X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-X.sub.2c-Y.sub.2a-X.sub.3a-- Y.sub.3a-Y.sub.3b (Formula XXII);
Y.sub.1a-Y.sub.1b-X.sub.1a-Y.sub.2a-X.sub.2a-X.sub.2b-X.sub.2c-Y.sub.3a-- Y.sub.3b-X.sub.3a (Formula XXIII);
X.sub.1a-X.sub.1b-Y.sub.1a-X.sub.2a-Y.sub.2a-X.sub.3a-X.sub.3b (Formula XXIV);
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-Y.sub.2a-X.sub.2a-Y.sub.3a-- X.sub.3a-X.sub.3b (Formula XXV);
X.sub.1a-X.sub.1b-Y.sub.1a-X.sub.2a-Y.sub.2a-X.sub.3a-X.sub.3b-Y.sub.3a-- Y.sub.3b-Y.sub.3c (Formula XXVI);
X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.1a-Y.sub.1b-Y.sub.1c (Formula XXVII);
X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1a (Formula XXVIII);
Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-Y.sub.2a-Y.sub.2b-Y.sub.2c-- Y.sub.2a-X.sub.2a (Formula XXIX);
X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.1a-Y.sub.1b-Y.sub.1c-- Y.sub.1d-Y.sub.1e (Formula XXX);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-Y.sub.2b-Y.sub.2c-- X.sub.2a-X.sub.2b (Formula XXXI);
X.sub.1a-Y.sub.1a-X.sub.2a-Y.sub.2a-X.sub.3a-X.sub.3b-X.sub.3c-Y.sub.3a-- Y.sub.3b-Y.sub.3c (Formula XXXII);
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-X.sub.1c (Formula XXXIII);
Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1a (Formula XXXIV);
X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-X.sub.2a-X.sub.2b-X.sub.2c-- X.sub.2a-Y.sub.2a (Formula XXXV);
Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.1a-X.sub.1b-X.sub.1c-- X.sub.1d-X.sub.1e (Formula XXXVI);
X.sub.1a-X.sub.1b-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.2a-X.sub.2b-X.sub.2c-- Y.sub.2a-Y.sub.2b (Formula XXXVII);
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1a-X.sub.1c-Y.sub.2a-X.sub.2a-- Y.sub.3a-X.sub.3a (Formula XXXVIII);
Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.2a (Formula XXXIX);
Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.2a-Y.sub.2b-- Y.sub.2c-Y.sub.2a (Formula XL);
Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.2a-- Y.sub.2b-Y.sub.2c (Formula XLI);
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-- Y.sub.2a-Y.sub.2b (Formula XLII);
Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1e-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-- X.sub.1e-Y.sub.2a (Formula XLIII);
X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.2a (Formula XLIV);
X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.2a-X.sub.2b-- X.sub.2c-X.sub.2a (Formula XLV);
X.sub.1a-X.sub.1b-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.2a-- X.sub.2b-X.sub.2c (Formula XLVI);
X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-- X.sub.2a-X.sub.2b (Formula XLVII);
X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-- Y.sub.1e-X.sub.2a (Formula XLVIII);
Y.sub.1a-X.sub.1a-Y.sub.2a-X.sub.2a-Y.sub.3a-X.sub.3a (Formula XLIX);
Y.sub.1a-Y.sub.1b-X.sub.1a-Y.sub.2a-Y.sub.2b-X.sub.2a-Y.sub.3a-Y.sub.3b-- X.sub.3a-Y.sub.4a (Formula L);
X.sub.1a-X.sub.1b-Y.sub.1a-Y.sub.1b-X.sub.2a-Y.sub.2a-Y.sub.2b-Y.sub.2c-- Y.sub.2d (Formula LI);
Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1a-X.sub.1a-Y.sub.2a-Y.sub.2b-X.sub.2a-- X.sub.2b (Formula LII);
Y.sub.1a-Y.sub.1b-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.2b-Y.sub.3a-- X.sub.3a-Y.sub.4a (Formula LIII); and
Y.sub.1a-X.sub.1a-Y.sub.2a-X.sub.2a-Y.sub.3a-Y.sub.3b-Y.sub.3c-X.sub.3a-- Y.sub.4a-Y.sub.4b (Formula LIV).
[0062] Typically, the striapathic region (or a portion thereof) of an anti-inflammatory polypeptide will have an amphipathic conformation (e.g., under physiological conditions). To be considered amphipathic, the striapathic region (or portion thereof) need not be in the amphipathic conformation at all times. Rather, it is sufficient that the amphipathic conformation be present at least 50%, 60%, 70%, 80%, or more of the time, or when the anti-inflammatory polypeptide is binding to a target molecule, such as an NF-kB Class II protein (e.g., Rel B). Often, the amphipathic conformation will be associated with a particular secondary structure, such as a helical structure. Thus, the striapathic region (or a portion thereof) of the anti-inflammatory polypeptide can have an amphipathic 3.sub.10-helical conformation, an amphipathic .alpha.-helical conformation, an amphipathic n-helical conformation, or an amphipathic poly-proline helical conformation. Alternatively, the striapathic region (or a portion thereof) of the anti-inflammatory polypeptide can have an amphipathic .beta.-strand conformation.
[0063] For anti-inflammatory peptides that comprise a striapathic region that includes or has an amphipathic helical conformation (e.g., 3.sub.10-helical, .alpha.-helical, n-helical, or polyproline helical conformation), the hydrophobic surface ("side") can have a facial arc of at least 100.degree.. In certain embodiments, the facial arc of the hydrophobic surface or side is at least 125.degree., 150.degree., 175.degree., 200.degree., 225.degree., 250.degree., 275.degree., or 300.degree..
[0064] Anti-inflammatory polypeptides in certain embodiments have a striapathic region that has a relatively large hydrophobic volume. Accordingly, the striapathic region can optimally contain hydrophobic amino acid residues having a total side-chain volume of at least 600 cubic angstroms. In certain embodiments, the hydrophobic amino acid residues of the striapathic region have a hydrophobic side-chain volume of at least 650, 700, 750, 800, 850, 900, 950, 1000, or more cubic angstroms. Alternatively, or in addition, the striapathic region can be characterized by a ratio of the sum of the side-chain volume of hydrophobic amino acid residues to the sum of the side-chain volume of hydrophilic amino acid residues, wherein the ratio is at least 0.75 or higher. For example, the ratio can be at least 0.8, 0.85, 0.9, 0.95, 1.0, 1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, 1.9, 2.0, 2.1, 2.2, 2.3, 2.4, 2.5, or greater.
[0065] Because of the desirability of a striapathic region having a relatively large hydrophobic side-chain volume, it is generally preferable to include one or more (e.g., 2, 3, 4, 5, or more) large hydrophobic amino acid residues in the sequence of the striapathic region. Conversely, it is generally preferable to have two or fewer (e.g., 1 or 0) small hydrophobic amino acid residues in the sequence of the striapathic region. Examples of large hydrophobic amino acid residues include tryptophan, phenylalanine, and tyrosine. In addition, under certain circumstances, histidine or arginine can be considered a large hydrophobic amino acid residue. Examples of small hydrophobic residues include glycine, alanine, serine, cysteine, valine, threonine, and proline. Accordingly, an anti-inflammatory polypeptide can have a striapathic region that includes one or more (e.g., 2, 3, 4, 5, or more) hydrophobic residues selected from the group consisting of tryptophan, phenylalanine, and tyrosine. Alternatively, the anti-inflammatory polypeptide can have a striapathic region that includes one or more (e.g., 2, 3, 4, 5, or more) hydrophobic residues selected from (i) the group consisting of tryptophan, phenylalanine, tyrosine, and histidine, or (ii) the group consisting of tryptophan, phenylalanine, tyrosine, and arginine. In certain embodiments, the anti-inflammatory polypeptide has a striapathic region that includes two or fewer (e.g., 1 or 0) hydrophobic residues selected from the group consisting of glycine, alanine, serine, cysteine, valine, threonine, and proline. Alternatively, the anti-inflammatory polypeptide can have a striapathic region that includes no more than one hydrophobic residue selected from the group consisting of glycine, alanine, serine, cysteine, valine, threonine, and proline. In other alternatives, the anti-inflammatory polypeptide can have a striapathic region that includes no glycine residues, no alanine residues, no serine residues, no cysteine residues, no valine residues, no threonine residues, and/or no proline residues.
[0066] It is also preferable that an anti-inflammatory polypeptide have a striapathic region characterized by a moderate level of cationicity (i.e., a striapathic region that does not contain an excessive number of amino acid residues having positively charged side chains). Examples of amino acid residues having positively charged side groups (assuming physiological conditions) includes lysine, typically arginine, and sometimes histidine. Examples of amino acid residues having negatively charged side chains (assuming physiological conditions) include aspartic acid and glutamic acid. Examples of hydrophilic amino acid residues having uncharged side chains (assuming physiological conditions) include aspargine and glutamine. Accordingly, an anti-inflammatory polypeptide can have a striapathic region that includes five or fewer (e.g., 4, 3, 2) lysine residues. Alternatively, an anti-inflammatory polypeptide can have a striapathic region that includes five or fewer (e.g., 4, 3, 2) amino acid residues selected from the group consisting of lysine and arginine. In other alternatives, an anti-inflammatory polypeptide can have a striapathic region that includes five or fewer (e.g., 4, 3, 2) amino acid residues selected from the group consisting of lysine, arginine, and histidine. For anti-inflammatory polypeptides that have a striapathic region that includes one or more (e.g., two or more) positively charged amino acid residues, it can be advantageous for the striapathic region to also include some negatively charged or polar, uncharged amino acid residues. For example, the anti-inflammatory polypeptide can have a striapathic region that includes both positively and negatively charged amino acid residues, such that the net charge on the polypeptide is no more than +2 or +1 (e.g., the number of positively charged amino acid residues does not exceed the number of negatively charged amino acid residues by more than one or two). Alternatively, the anti-inflammatory polypeptide can have a striapathic region that includes both positively charged and polar, uncharged amino acid residues, such that the net charge on the polypeptide is no more than +2 or +1 (e.g., the number of positively charged amino acid residues does not exceed one or two). In other alternatives, the anti-inflammatory polypeptide can have a striapathic region that includes both positively charged, negatively charged, and hydrophilic uncharged charged amino acid residues, such that the net charge on the polypeptide is no more than +2.
[0067] To avoid certain undesired interactions between RP peptides and other molecules (whether another RP peptide, a metal ion, etc.) it can be advantageous to limit the number of certain types of amino acid residues in the polypeptide. For example, because cysteine residues form di-sulfide bonds under certain conditions (e.g., oxidative environments), it can be useful to limit the number of cysteine residues in a polypeptide of the invention to no more than one or two, or even none. Because histidine residues chelate metals under certain conditions (e.g., alkaline environments), it can be useful to limit the number of histidine residues in a polypeptide of the invention to no more than one or two, or even none. In addition, because proline residues tend to introduce kinks into secondary structure elements (e.g., .alpha.-helices and .beta.-strands), it can be useful exclude proline residues in the striapathic region of a polypeptide of the invention, or limit their number to no more than one.
[0068] Class I Polypeptides
[0069] An anti-inflammatory polypeptide of the invention can be a Class I polypeptide. Class I polypeptides comprise, consist essentially of, or consist of a striapathic region that includes a sequence selected from the group of sequences defined by Formula I:
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c (Formula I).
[0070] Each of amino acid residues Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.2a, Y.sub.2b, and Y.sub.2c in Formula I can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), His (H), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Pro (P), Thr (T), Ser (S), Ala (A), and Gly (G). In certain embodiments, at least 3, 4, 5, or 6 of amino acid residues Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.2a, Y.sub.2b, and Y.sub.2c in Formula I are selected from the group consisting of Phe (F), Trp (W), Tyr (Y), His (H), and Leu (L). In certain embodiments, at least 3, 4, 5, or 6 of amino acid residues Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.2a, Y.sub.2b, and Y.sub.2c in Formula I are selected from the group consisting of Phe (F), Trp (W), and Tyr (Y). In certain embodiments, less than two (and in certain embodiments 1 or none) of amino acid residues Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.2a, Y.sub.2b, and Y.sub.2c in Formula I are selected from the group consisting of Pro (P), Thr (T), Ser (S), Ala (A), and Gly (G).
[0071] The module Y.sub.1a-Y.sub.1b-Y.sub.1c in Formula I can have a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), Tyr-Tyr-Tyr (YYY), Leu-Leu-Leu (LLL), Cys-Cys-Cys (CCC), Met-Met-Met (MMM), Val-Val-Val (VVV), Ile-Ile-Ile (III).
[0072] Alternatively, the module Y.sub.1a-Y.sub.1b-Y.sub.1c in Formula I can have a sequence selected from the group consisting of Pro-Pro-Pro (PPP), Thr-Thr-Thr (TTT), and Ala-Ala-Ala (AAA). In certain embodiments, module Y.sub.1a-Y.sub.1b-Y.sub.1c in Formula I has a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), Tyr-Tyr-Tyr (YYY), and combinations thereof (e.g., Phe-Phe-Trp (FFW), Phe-Trp-Trp (FWW), Trp-Phe-Trp (WFW), Trp-Trp-Phe (WWF), Phe-Phe-Tyr (FFY), Phe-Tyr-Tyr (FYY), Tyr-Phe-Tyr (YFY), Tyr-Tyr-Phe (YYF), Trp-Trp-Tyr (WWY), Trp-Tyr-Tyr (WYY), Tyr-Trp-Tyr (YWY), Tyr-Tyr-Trp (YYW), Phe-Trp-Tyr (FWY), Phe-Tyr-Trp (FYW), Trp-Phe-Tyr (WFY), Trp-Tyr-Phe (WYF), Tyr-Trp-Phe (YWF), or Tyr-Phe-Trp (YFW)).
[0073] The module Y.sub.2a-Y.sub.2b-Y.sub.2c in Formula I can have a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), Tyr-Tyr-Tyr (YYY), Leu-Leu-Leu (LLL), Cys-Cys-Cys (CCC), Met-Met-Met (MMM), Val-Val-Val (VVV), and Ile-Ile-Ile (III). Alternatively, the module Y.sub.2a-Y.sub.2b-Y.sub.2c in Formula I can have a sequence selected from the group consisting of Pro-Pro-Pro (PPP), Thr-Thr-Thr (TTT), and Ala-Ala-Ala (AAA). In certain embodiments, module Y.sub.2a-Y.sub.2b-Y.sub.2c in Formula I has a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), Tyr-Tyr-Tyr (YYY), and combinations thereof (e.g., Phe-Phe-Trp (FFW), Phe-Trp-Trp (FWW), Trp-Phe-Trp (WFW), Trp-Trp-Phe (WWF), Phe-Phe-Tyr (FFY), Phe-Tyr-Tyr (FYY), Tyr-Phe-Tyr (YFY), Tyr-Tyr-Phe (YYF), Trp-Trp-Tyr (WWY), Trp-Tyr-Tyr (WYY), Tyr-Trp-Tyr (YWY), Tyr-Tyr-Trp (YYW), Phe-Trp-Tyr (FWY), Phe-Tyr-Trp (FYW), Trp-Phe-Tyr (WFY), Trp-Tyr-Phe (WYF), Tyr-Trp-Phe (YWF), or Tyr-Phe-Trp (YFW)).
[0074] Thus, a Class I anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region having a sequence selected from the group consisting of FFF-X.sub.1a-FFF (SEQ ID NO: 1), WWW-X.sub.1a-WWW (SEQ ID NO: 2), YYY-X.sub.1a-YYY (SEQ ID NO: 3), and combinations thereof. Alternatively, a Class I anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region having a sequence selected from the group consisting of LLL-X.sub.1a-LLL (SEQ ID NO: 4), CCC-X.sub.1a-CCC (SEQ ID NO: 5), MMM-X.sub.1a-MMM (SEQ ID NO: 6), VVV-X.sub.1a-VVV (SEQ ID NO: 7), and III-X.sub.1a-III (SEQ ID NO: 8). In such peptides, X.sub.1a can be selected from the group consisting of Arg (R), His (H), and Lys (K); or X.sub.1a can be selected from the group consisting of Glu (E), Gln (Q), Asn (N), and Asp (D).
[0075] A Class I anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region having a sequence selected from the group of sequences defined by Formula II or the group of sequences defined by Formula III:
Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.2a-- Y.sub.3a-X.sub.3a (Formula II);
X.sub.2a-Y.sub.3a-X.sub.3a-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-- Y.sub.2b-Y.sub.2c (Formula III).
[0076] The Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c sequences defined by Formulas II and III can be any of the sequences described above in connection with Formula I. X.sub.2a and X.sub.3a in Formulas II and III can be each individually selected from the group consisting of Arg (R), His (H), Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D). Alternatively, X.sub.2a and X.sub.3a in Formulas II and III can be each individually selected from the group consisting of Arg (R), His (H), and Lys (K). In other alternatives, X.sub.2a and X.sub.3a in Formulas II and III can be each individually selected from the group consisting of Arg (R), His (H), Lys (K), and Gln (Q). In other alternatives, X.sub.2a and X.sub.3a in Formulas II and III can be each individually selected from the group consisting Glu (E), Gln (Q), Asn (N), and Asp (D). In other alternatives, X.sub.2a in Formulas II and III can be selected from the group consisting of Arg (R), His (H), and Lys (K), and X.sub.3a in Formulas II and III can be selected from the group consisting of Glu (E), Gln (Q), Asn (N), and Asp (D). Y.sub.3a in Formulas II and III can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), and Ile (I). In certain embodiments, Y.sub.3a in Formulas II and III is selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L).
[0077] The modules X.sub.2a-Y.sub.3a-X.sub.3a in Formulas II and III can be selected from the group consisting of EFQ, EFE, EFN, EFD, NFQ, NFE, NFN, NFD, QFQ, QFE, QFN, QFD, DFQ, DFE, DFN, DFD, EWQ, EWE, EWN, EWD, NWQ, NWE, NWN, NWD, QWQ, QWE, QWN, QWD, DWQ, DWE, DWN, DWD, EYQ, EYE, EFN, EYD, NYQ, NYE, NYN, NYD, QYQ, QYE, QYN, QYD, DYQ, DYE, DYN, DYD, ELQ, ELE, ELN, ELD, NLQ, NLE, NLN, NLD, QLQ, QLE, QLN, QLD, DLQ, DLE, DLN, DLD, RFR, RFQ, RFE, RFN, RFD, RWR, RWQ, RWE, RWN, and RWD.
[0078] A Class I anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region comprising, consisting essentially of, or consisting of a sequence selected from the group of sequences listed in Table 3, e.g., RP394, RP108-RP123, RP125-131, RP133, RP135-RP141, RP143-RP146, RP148-RP150, RP152-RP165, RP179, RP395, RP211, RP230, RP232, RP258, RP267, RP268, RP271, RP273, RP280-281, and RP287. In certain embodiments, the Class I anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that comprises, consists essentially of, or consists of a sequence selected from the group of sequences consisting of RP113 (SEQ ID NO: 39), RP118 (SEQ ID NO: 44), and RP394 (SEQ ID NO: 33).
[0079] Class II Polypeptides
[0080] An anti-inflammatory polypeptide of the invention can be a Class II polypeptide. Class II anti-inflammatory polypeptides can comprise, consist essentially of, or consist of a striapathic region that includes a sequence selected from the group of sequences defined by Formula VII:
Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y.sub.3a (Formula VII).
[0081] Amino acid residue Y.sub.2a in Formula VII can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Pro (P), Thr (T), Ser (S), Ala (A), and Gly (G). In certain embodiments, amino acid residue Y.sub.2a in Formula VII is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y). Alternatively, amino acid residue Y.sub.2a in Formula VII can be selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I).
[0082] Amino acid residue Y.sub.2b in Formula VII can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Pro (P), Thr (T), Ser (S), Ala (A), and Gly (G). In certain embodiments, amino acid residue Y.sub.2b in Formula VII is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y). Alternatively, amino acid residue Y.sub.2b in Formula VII can be selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I).
[0083] Amino acid residue X.sub.1b in Formula VII can be selected from the group consisting of Arg (R), Lys (K), and His (H). Alternatively amino acid residue X.sub.1b in Formula VII can be selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0084] Amino acid residue X.sub.2a in Formula VII can be selected from the group consisting of Arg (R), Lys (K), and His (H). Alternatively, amino acid residue X.sub.2a can be selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0085] The sequence X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a in Formula VII can be selected from the group consisting of Lys-Phe-Phe-Lys (KFFK; SEQ ID NO: 386), Lys-Trp-Trp-Lys (KWWK; SEQ ID NO: 387), Lys-Tyr-Try-Lys (KYYK; SEQ ID NO: 388), Lys-Phe-Trp-Lys (KFWK; SEQ ID NO: 389), Lys-Trp-Phe-Lys (KWFK; SEQ ID NO: 390), Lys-Phe-Tyr-Lys (KFYK; SEQ ID NO: 391), Lys-Tyr-Phe-Lys (KYFK; SEQ ID NO: 392), Lys-Trp-Tyr-Lys (KWYK; SEQ ID NO: 393), and Lys-Tyr-Trp-Lys (KYWK; SEQ ID NO: 394). Alternatively, the sequence X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a in Formula VII can be selected from the group consisting of Arg-Phe-Phe-Arg (RFFR; SEQ ID NO: 395), Arg-Trp-Trp-Arg (RWWR; SEQ ID NO: 396), Arg-Tyr-Try-Arg (RYYR; SEQ ID NO: 397), Arg-Phe-Trp-Arg (RFWR; SEQ ID NO: 398), Arg-Trp-Phe-Arg (RWFR; SEQ ID NO: 399), Arg-Phe-Tyr-Arg (RFYR; SEQ ID NO: 400), Arg-Tyr-Phe-Arg (RYFR; SEQ ID NO: 401), Arg-Trp-Tyr-Arg (RWYR; SEQ ID NO: 402), and Arg-Tyr-Trp-Arg (RYWR; SEQ ID NO: 403). In other alternatives, the sequence X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a in Formula VII can be selected from the group consisting of His-Phe-Phe-His (HFFH; SEQ ID NO: 404), His-Trp-Trp-His (HWWH; SEQ ID NO: 405), His-Tyr-Try-His (HYYH; SEQ ID NO: 406), His-Phe-Trp-His (HFWH; SEQ ID NO: 407), His-Trp-Phe-His (HWFH; SEQ ID NO: 408), His-Phe-Tyr-His (HFYH; SEQ ID NO: 409), His-Tyr-Phe-His (HYFH; SEQ ID NO: 410), His-Trp-Tyr-His (HWYH; SEQ ID NO: 411), and His-Tyr-Trp-His (HYWH; SEQ ID NO:132).
[0086] Amino acid residue X.sub.1a in Formula VII can be selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E). In certain embodiments, amino acid residue X.sub.1a is selected from the group consisting of Arg (R) and Gln (Q). In certain embodiments, amino acid residue X.sub.1a in Formula VII is Arg (R). Alternatively, amino acid residue X.sub.1a in Formula VII can be selected from the group consisting of Lys (K), Gln (Q), Glu (E), and Asn (N).
[0087] Amino acid resiude X.sub.2b in Formula VII can be selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E). In certain embodiments, amino acid residue X.sub.2b is selected from the group consisting of Arg (R) and Gln (Q). In certain embodiments, amino acid residue X.sub.2b in Formula VII is Arg (R). Alternatively, amino acid residue X.sub.2b in Formula VII can be selected from the group consisting of Lys (K), Gln (Q), Glu (E), and Asn (N).
[0088] Amino acid residue Y.sub.1a in Formula VII can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G). In certain embodiments, amino acid residue Y.sub.1a in Formula VII is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y). Alternatively, amino acid residue Y.sub.1a in Formula VII can be selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I).
[0089] Amino acid residue Y.sub.3a in Formula VII can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G). In certain embodiments, amino acid residue Y.sub.3a in Formula VII is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y). Alternatively, amino acid residue Y.sub.3a in Formula VII can be selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I).
[0090] Thus, a Class II anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region having a sequence selected from the group consisting of F-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 9), F-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-W (SEQ ID NO: 10), W-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 11), F-X.sub.1a-X.sub.1b-FW-X.sub.2a-X.sub.2b-F (SEQ ID NO: 12), F-X.sub.1a-X.sub.1b-WF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 13), F-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-F (SEQ ID NO: 14), W-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-F (SEQ ID NO: 15), F-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-W (SEQ ID NO: 16), W-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-W (SEQ ID NO: 17), F-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 18), Y-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 19), F-X.sub.1a-X.sub.1b-FY-X.sub.2a-X.sub.2b-F (SEQ ID NO: 20), F-X.sub.1a-X.sub.1b-YF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 21), F-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-F (SEQ ID NO: 22), Y-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-F (SEQ ID NO: 23), F-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 24), and Y-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 25), Y-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-W (SEQ ID NO: 26), W-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 27), Y-X.sub.1a-X.sub.1b-YW-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 28), Y-X.sub.1a-X.sub.1b-WY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 29), Y-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 30), W-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 31), and Y-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-W (SEQ ID NO: 32). Amino acid residues X.sub.1a, X.sub.1b, X.sub.2a, and X.sub.2b in the foregoing sequences can be selected as discussed above.
[0091] A Class II anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes a first additional amino acid residue directly bound to amino acid residue Y.sub.1a of Formula VII. The first additional amino acid residue can be a hydrophobic amino acid residue (e.g., a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G); a residue selected from the group consisting of Phe (F), Trp (W), and Tyr (Y); a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or, a residue selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), and Ile (I)). Alternatively, the first additional amino acid residue can be a hydrophilic amino acid residue (e.g., a residue selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E); a residue selected from the group consisting of Arg (R), Lys (K), and His (H); a residue selected from the group consisting Arg (R), Lys (K), His (H), and Gln (Q); or a residue selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E)).
[0092] A Class II anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes a first additional amino acid residue directly bound to amino acid residue Y.sub.3a of Formula VII. The first additional amino acid residue can be a hydrophobic amino acid residue (e.g., a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G); a residue selected from the group consisting of Phe (F), Trp (W), and Tyr (Y); a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or, a residue selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), and Ile (I)). Alternatively, the first additional amino acid residue can be a hydrophilic amino acid residue (e.g., a residue selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E); a residue selected from the group consisting of Arg (R), Lys (K), and His (H); a residue selected from the group consisting Arg (R), Lys (K), His (H), and Gln (Q); or a residue selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E)).
[0093] A Class II anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes a first additional amino acid residue directly bound to amino acid residue Y.sub.1a of Formula VII and a second additional amino acid reside directly bound to amino acid residue Y.sub.3a of Formula VII. The first additional amino acid residue can be a hydrophobic amino acid residue and the second additional amino acid residue can be a hydrophilic amino acid residue. Alternatively, the first additional amino acid residue can be a hydrophilic amino acid residue and the second amino acid residue can be a hydrophobic amino acid residue. Regardless, the additional hydrophobic amino acid residue can be selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G); and in certain embodiments from the group consisting of Phe (F), Trp (W), and Tyr (Y); and in additional embodiments from the group consisting of Phe (F). The additional hydrophilic amino acid residue can be selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E); and in certain embodiments, a residue selected from the group consisting of Arg (R), Lys (K), and His (H); or a residue selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0094] A Class II anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region comprising, consisting essentially of, or consisting of a sequence selected from the group of sequences listed in Table 5, e.g., RP124, RP132, RP134, RP142, RP147, RP151, RP166-RP172, RP175, RP177, RP182, RP183, RP185, RP186, RP 424, RP190, RP194, RP198, RP199-RP202, RP204, RP206, RP207, RP209, RP210, RP212-RP216, RP218, RP219, RP425, RP225, RP227, RP233-RP239, RP398, RP241-RP247, RP250-RP256, RP426, RP427, RP285, and RP387. In certain embodiments, the Class II anti-inflammatory polypeptide comprises, consists essentially of, or consists of a striapathic region comprising, consisting essentially of, or consisting of a sequence selected from the group consisting of RP124 (SEQ ID NO: 106), RP166 (SEQ ID NO: 112), RP182 (SEQ ID NO: 121), and RP183 (SEQ ID NO: 122).
[0095] Class XII Polypeptides
[0096] An anti-inflammatory polypeptide of the invention can be a Class XII polypeptide. Class XII anti-inflammatory polypeptides can comprise, consist essentially of, or consist of a striapathic region that includes a sequence selected from the group of sequences defined by Formula XLIX:
Y.sub.1a-X.sub.1a-Y.sub.2a-X.sub.2a-Y.sub.3a-X.sub.3a (Formula XLIX).
[0097] Amino acid residues Y.sub.1a, Y.sub.2a, and Y.sub.3a of Formula XLIX can be each independently selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Ile (I), Cys (C), Met (M), Val (V), Pro (P), Thr (T), Ser (S), Ala (A), and Gly (G). In certain embodiments, amino acid residues Y.sub.1a, Y.sub.2a, and Y.sub.3a of Formula XLIX are each independently selected from: the group consisting of Phe (F), Trp (W), and Tyr (Y); the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Ile (I), Cys (C), Met (M), Val (V), and Ala (A).
[0098] Amino acid residues X.sub.1a, X.sub.2a, and X.sub.3a of Formula XLIX can be each independently selected from the group consisting of Arg (R), Lys (K), His (H), Gln (Q), Glu (E), Asn (N), and Asp (D). In certain embodiments, amino acid residues X.sub.1a, X.sub.2a, and X.sub.3a are each independently selected from the group consisting of Arg (R), Lys (K), and His (H). Alternatively, amino acid residues X.sub.1a, X.sub.2a, and X.sub.3a are each independently selected from the group consisting of Arg (R), Lys (K), His (H), and Gln (Q).
[0099] A Class XII anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes a first additional amino acid residue. The first additional amino acid residue can be a hydrophilic amino acid residue directly bound to amino acid residue Y.sub.1a of Formula XLIX. Thus, the first additional amino acid residue can be, for example, a residue selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E); a residue selected from the group consisting of Arg (R), Lys (K), and His (H); a residue selected from the group consisting Arg (R), Lys (K), His (H), and Gln (Q); or a residue selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E)). Alternatively, the first amino acid residue can be a hydrophobic amino acid residue directly bound to amino acid residue X.sub.3a of Formula XLIX. Thus, the first additional amino acid residue can be, for example, a residue selected from the group consisting of Phe (F), Trp (W), and Tyr (Y); a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Ile (I), Cys (C), Met (M), Val (V), and Ala (A)).
[0100] A Class XII anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes first and second additional amino acid residues. The first additional amino acid residue can be a hydrophilic amino acid residue, as discussed above, which is directly bound to amino acid residue Y.sub.1a of Formula XLIX. The second additional amino acid residue can be directly bound to the first additional amino acid residue. Thus, the second additional amino acid residue can be a hydrophobic amino acid residue, e.g., a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G); a residue selected from the group consisting of Phe (F), Trp (W), and Tyr (Y); a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or, a residue selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), and Ile (I)). Alternatively, the second additional amino acid residue can be a hydrophobic amino acid residue directly bound to amino acid residue X.sub.3a of Formula XLIX, as discussed above.
[0101] A Class XII anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes first, second, and third additional amino acid residues. The first additional amino acid residue can be a hydrophilic amino acid residue which is directly bound to amino acid residue Y.sub.1a of Formula XLIX and the second additional amino acid residue can be a hydrophobic amino acid residue which is directly bound to the first additional amino acid residue, as discussed above. The third additional amino acid residue can be a hydrophilic amino acid residue that is directly bound to the second additional amino acid residue. Thus, the third additional amino acid residue can be, for example, a residue selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E); a residue selected from the group consisting of Arg (R), Lys (K), and His (H); a residue selected from the group consisting Arg (R), Lys (K), His (H), and Gln (Q); or a residue selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E)). Alternatively, the third amino acid residue can be a hydrophobic amino acid residue directly bound to amino acid residue X.sub.3a of Formula XLIX. Thus, the third additional amino acid residue can be, for example, a residue selected from the group consisting of Phe (F), Trp (W), and Tyr (Y); a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Ile (I), Cys (C), Met (M), Val (V), and Ala (A)).
[0102] A Class XII anti-inflammatory polypeptide can comprise, consist essentially of, or consist of a striapathic region that further includes four, five, six, or more additional amino acid residues. The additional amino acid residue can be added in a manner that continues the alternating patter of a hydrophobic amino acid residue followed by a hydrophilic amino acid residue followed by a hydrophobic amino acid residue, as shown in Formula XLIX. In this manner, Class XII anti-inflammatory polypeptides can be expanded to comprise, consist essentially of, or consist of a striapathic region having 10, 11, 12, or more amino acid residues.
[0103] An anti-inflammatory polypeptide of Class XII can comprise, consist essentially of, or consist of a striapathic region comprising, consisting essentially of, or consisting of a sequence selected from the group consisting of RP393, RP391, PR392, RP390, and RP389.
[0104] Class XIV Polypeptides
[0105] An anti-inflammatory polypeptide of the invention can be a Class XIV polypeptide. Class XIV anti-inflammatory polypeptides can comprise, consist essentially of, or consist of a striapathic region that includes a sequence selected from the group of sequences defined by any one of Formulas LI through LIV:
X.sub.1a-X.sub.1b-Y.sub.1a-Y.sub.1b-X.sub.2a-Y.sub.2a-Y.sub.2b-Y.sub.2c-- Y.sub.2d (Formula LI);
Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1a-X.sub.1a-Y.sub.2a-Y.sub.2b-X.sub.2a-- X.sub.2b (Formula LII);
Y.sub.1a-Y.sub.1b-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.2b-Y.sub.3a-- X.sub.3a-Y.sub.4a (Formula LIII); and
Y.sub.1a-X.sub.1a-Y.sub.2a-X.sub.2a-Y.sub.3a-Y.sub.3b-Y.sub.3c-X.sub.3a-- Y.sub.4a-Y.sub.4b (Formula LIV).
[0106] The striapathic region of a Class XIV polypeptide can include at least 3 (e.g., 3 to 6) proline amino acid residues. For example, amino acid residues Y.sub.1a, Y.sub.2a, and Y.sub.2b in Formula LI can be proline amino acid residues. Alternatively, amino acid residues Y.sub.1c, Y.sub.1d, and Y.sub.2b in Formula LII can be proline amino acid residues. In other alternatives, amino acid residues Y.sub.1a, Y.sub.2a, Y.sub.2b, Y.sub.2c, Y.sub.3a, and Y.sub.4a in Formula LIII can be proline amino acid residues. In still other alternatives, amino acid residues Y.sub.1a, Y.sub.2b, Y.sub.3a, Y.sub.3b, Y.sub.3c, and Y.sub.4b in Formula LIV can be proline amino acid residues.
[0107] Hydrophobic amino acid residues (e.g., Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.1a, Y.sub.2a, Y.sub.2b, Y.sub.2c, Y.sub.2d, Y.sub.3a, Y.sub.3b, Y.sub.3c, Y.sub.4a, and Y.sub.4b) not designated as proline residues in Formulas LI through LIV can be each individually selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G). In certain embodiments, such hydrophobic amino acid residues are each individually selected from: the group consisting of Phe (F), Trp (W), and Tyr (Y); the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or, the group consisting of Leu (L), Cys (C), Met (M), Val (V), and Ile (I)).
[0108] Hydrophilic amino acid residues in Formulas LI through LIV (e.g., X.sub.1a, X.sub.1b, X.sub.2a, X.sub.2b, and X.sub.3a) can be each individually selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E). In certain embodiments, such hydrophilic amino acid residues are each individually selected from the group consisting of Arg (R), Lys (K), and His (H). Alternatively, such hydrophilic amino acid residues are each individually selected from: the group consisting of Arg (R), Lys (K), His (H), and Gln (Q); or the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0109] An anti-inflammatory polypeptide of Class XIV can comprise, consist essentially of, or consist of a striapathic region that comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP449, RP450, RP448, RP447, RP452, RP451, RP444, RP441, RP446, RP445, RP442, and RP443.
[0110] Other Classes of Polypeptides
[0111] An anti-inflammatory polypeptide of the invention can be from any of Classes II through XI and XIII. Such anti-inflammatory polypeptides can comprise, consist essentially of, or consist of a striapathic region that includes a sequence selected from the group of sequences defined by any one of Formulas IV through XLVIII and L.
[0112] Hydrophobic amino acid residues in Formulas IV through XLVIII and L (e.g., Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.1d, Y.sub.1e, Y.sub.2a, Y.sub.2b, Y.sub.2c, Y.sub.2d, Y.sub.2e, Y.sub.3a, Y.sub.3b, Y.sub.3c, Y.sub.4a, and Y.sub.4b) can be each individually selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G). In certain embodiments, such hydrophobic amino acid residues are each individually selected from: the group consisting of Phe (F), Trp (W), and Tyr (Y); the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or, the group consisting of Leu (L), Cys (C), Met (M), Val (V), and Ile (I)).
[0113] Hydrophilic amino acid residues in Formulas IV through XLVIII and L (e.g., X.sub.1a, X.sub.1b, X.sub.1c, X.sub.1a, X.sub.2a, X.sub.2b, X.sub.2c, X.sub.2a, X.sub.3a, X.sub.3b, X.sub.3c, X.sub.4a, and X.sub.4b) can be each individually selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E). In certain embodiments, such hydrophilic amino acid residues are each individually selected from the group consisting of Arg (R), Lys (K), and His (H). Alternatively, such hydrophilic amino acid residues are each individually selected from: the group consisting of Arg (R), Lys (K), His (H), and Gln (Q); or the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0114] An anti-inflammatory polypeptide of any one of Formulas IV through XLVIII and L can comprise, consist essentially of, or consist of a striapathic region that further includes a first additional amino acid residue directly bound to the first amino acid residue of the Formula (e.g., Y.sub.1a or X.sub.1a) or to the last amino acid residue in the formula. The first additional amino acid residue can be a hydrophilic amino acid residue (e.g., a residue selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E); a residue selected from the group consisting of Arg (R), Lys (K), and His (H); a residue selected from the group consisting Arg (R), Lys (K), His (H), and Gln (Q); or a residue selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E)). Alternatively, the first additional amino acid residue can be a hydrophobic amino acid residue (e.g., a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), Thr (T), Pro (P), Ser (S), Ala (A), and Gly (G); a residue selected from the group consisting of Phe (F), Trp (W), and Tyr (Y); a residue selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L); or, a residue selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), and Ile (I)).
[0115] An anti-inflammatory polypeptide of any one of Formulas IV through XLVIII and L can comprise, consist essentially of, or consist of a striapathic region that further includes first and second additional amino acid residues, with the first additional amino acid residue directly bound to the first amino acid residue of the Formula (e.g., Y.sub.1a or X.sub.1a) or the last amino acid residue in the formula, and the second additional amino acid residue directly bound to the first amino acid residue in the formula, the last amino acid residue in the formula, or the first additional amino acid residue. The first additional amino acid residue can be a hydrophilic or hydrophobic amino acid residue, as discussed above. The second additional amino acid residue likewise can be a hydrophilic or hydrophobic amino acid residue, as discussed above.
[0116] An anti-inflammatory polypeptide of any one of Formulas IV through XLVIII and L can comprise, consist essentially of, or consist of a striapathic region that comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP396, RP405, RP174, RP176, RP178, RP180-181, RP184, RP408, RP187, RP416, RP188, RP189, RP388, RP417, RP191-RP193, RP404, RP196, RP397, RP197, RP402, RP203, RP409, RP205, RP208, RP217, RP220-RP224, RP226, RP229, RP231, RP240, RP248, RP249, RP415, RP257, RP259-RP266, RP269, RP272, RP274, RP277-RP279, RP282, RP283, RP286, RP289, and RP414.
[0117] Variant Polypeptides
[0118] The exemplary anti-inflammatory polypeptide sequences shown in Tables 3-9 (below) are merely examples and are not the only anti-inflammatory polypeptides provided herein. Indeed, fragments and variants of the sequences of the disclosed peptides are within the scope of the invention.
[0119] A "fragment" of the invention includes at least 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, or 23 contiguous amino acid residues of a polypeptide disclosed herein (or up to one less than the number of amino acid residues in the subject polypeptide) and retains at least one anti-inflammatory property of the subject polypeptide. Thus, fragments of the invention include polypeptides that are missing one, two, three, four, or more amino acids from the N-terminus and/or the C-terminus relative to a polypeptide disclosed herein.
[0120] A "variant" of the invention is a polypeptide that is substantially similar to a polypeptide disclosed herein and retains at least one anti-inflammatory property of the subject polypeptide. Variants can include deletions (i.e., truncations) of one or more amino acid residues at the N-terminus or the C-terminus of a subject polypeptide disclosed herein; deletion and/or addition of one or more amino acid residues at one or more internal sites in the subject polypeptide disclosed herein; and/or substitution of one or more amino acid residues at one or more positions in the subject polypeptide disclosed herein. For subject polypeptides that are 12 amino acid residues in length or shorter, variant polypeptides can include three or fewer (e.g., two, one, or none) deleted amino acid residues, whether located internally, at the N-terminal end, and/or at the C-terminal end.
[0121] Accordingly, the invention further provides anti-inflammatory polypeptides that are at least 50% identical (e.g., at least 60%, 70%, 80%, 90%, or more) to any one of the anti-inflammatory polypeptides disclosed in Tables 3-9 and still retain at least one anti-inflammatory property. For example, the invention provides anti-inflammatory polypeptides that are 3 to 24 amino acids residues in length and comprise, consist essentially of, or consist of a striapathic region sharing at least 50% identity (e.g., at least 60%, 70%, 80%, 90%, or more identity) with a Class I anti-inflammatory polypeptide (e.g., any one of the sequences of Table 3). Such identity can be shared, for example, with RP-394 (SEQ ID NO: 33), RP-108 (SEQ ID NO: 34), RP-113 (SEQ ID NO: 39), RP-118 (SEQ ID NO: 44), RP-129 (SEQ ID NO: 54), or RP-179 (SEQ ID NO: 86). Alternatively, the invention provides anti-inflammatory polypeptides that are 3 to 24 amino acid residues in length and comprise, consist essentially of, or consist of a striapathic region sharing at least 50% identity (e.g., at least 60%, 70%, 80%, 90%, or more identity) with a Class II, Sub-class 1 anti-inflammatory polypeptide (e.g., any one of the sequences of Table 5). Such identity can be shared, for example, with RP-124 (SEQ ID NO: 106), RP-134 (SEQ ID NO: 108), RP-166 (SEQ ID NO: 112), RP-168 (SEQ ID NO: 114), RP-182 (SEQ ID NO: 121), or RP-183 (SEQ ID NO: 122). In other alternatives, the invention provides anti-inflammatory polypeptides that are 3 to 24 amino acid residues in length and comprise, consist essentially of, or consist of a striapathic region sharing at least 50% identity (e.g., at least 60%, 70%, 80%, 90%, or more identity) with any Class II through Class IX or Class XIII anti-inflammatory polypeptide (e.g., any one of the sequences of Table 6). In other alternatives, the invention provides anti-inflammatory polypeptides that are 3 to 24 amino acid residues in length and comprise, consist essentially of, or consist of a striapathic region sharing at least 50% identity (e.g., at least 60%, 70%, 80%, 90%, or more identity) with any Class VIII to Class XI anti-inflammatory polypeptide (e.g., any one of the sequences of Table 7). In other alternatives, the invention provides anti-inflammatory polypeptides that are 3 to 24 amino acid residues in length and comprise, consist essentially of, or consist of a striapathic region sharing at least 50% identity (e.g., at least 60%, 70%, 80%, 90%, or more identity) with a Class XII or Class XIV anti-inflammatory polypeptide (e.g., any one of the sequences of Table 8). In still other alternatives, the invention provides anti-inflammatory polypeptides that are 3 to 24 amino acid residues in length and comprise, consist essentially of, or consist of a striapathic region sharing at least 50% identity (e.g., at least 60%, 70%, 80%, 90%, or more identity) with any one of the combination sequences of Table 9.
[0122] The differences between the striapathic region of a homologous anti-inflammatory polypeptide and any one of the anti-inflammatory polypeptides of Tables 3-9 can include deletions, additions, and/or substitutions of amino acid residues, as discussed above. Substituted amino acid residues can be unrelated to the amino acid residue being replaced (e.g., unrelated in terms or hydrophobicity/hydrophilicity, size, charge, polarity, etc.), or the substituted amino acid residues can constitute similar, conservative, or highly conservative amino acid substitutions. As used herein, "similar," "conservative," and "highly conservative" amino acid substitutions are defined as shown in Table 2, below. The determination of whether an amino acid residue substitution is similar, conservative, or highly conservative is based exclusively on the side chain of the amino acid residue and not the peptide backbone, which may be modified to increase peptide stability, as discussed below.
TABLE-US-00002 TABLE 2 Classification of Amino Acid Substitutions Highly Amino Acid in Similar Conservative Conservative Subject Amino Acid Amino Acid Amino Acid Polypeptide Substitutions Substitutions Substitutions Glycine (G) A, S, N A n/a Alanine (A) S, G, T, V, C, P, Q S, G, T S Serine (S) T, A, N, G, Q T, A, N T, A Threonine (T) S, A, V, N, M S, A, V, N S Cysteine (C) A, S, T, V, I A n/a Proline (P) A, S, T, K A n/a Methionine (M) L, I, V, F L, I, V L, I Valine (V) I, L, M, T, A I, L, M I Leucine (L) M, I, V, F, T, A M, I, V, F M, I Isoleucine (I) V, L, M, F, T, C V, L, M, F V, L, M Phenylalanine (F) W, L, M, I, V W, L n/a Tyrosine (Y) F, W, H, L, I F, W F Tryptophan (W) F, L, V F n/a Asparagine (N) Q Q Q Glutamine (Q) N N N Aspartic Acid (D) E E E Glutamic Acid (E) D D D Histidine (H) R, K R, K R, K Lysine (K) R, H R, H R, H Arginine (R) KH K, H K, H
[0123] In certain embodiments, a variant polypeptide of the invention binds to two or more targets (e.g., pro-inflammatory targets). In some embodiments, a variant polypeptide binds to three, four, five, or more pro-inflammatory targets. For example, a variant polypeptide can bind to any combination of targets disclosed herein (e.g., an NF-kB Class II protein and human serum albumin (HSA)), as discussed below. Such binding can be based on in silico, in vitro, or in vivo data.
[0124] Modeling Polypeptide Binding to Target Molecules
[0125] The determination of whether a polypeptide has anti-inflammatory properties can be performed in silico. For example, the binding of a polypeptide (e.g., a polypeptide that has a length of 3 to 24 amino acid residues and includes a striapathic region comprising at least 25% of the length of the polypeptide) to a putative target molecule can be modeled in silico, using any of the numerous molecular modeling and docking platforms available in the art, to thereby assess whether the polypeptide is an anti-inflammatory polypeptide. The on-line ClusPro.TM. algorithm, version 2.0 (developed at Boston University) is particularly useful for modeling the conformation of polypeptides and their binding to target molecules, such as signaling proteins, as described in the Examples set forth below. Modeling algorithms, such as the ClusPro.TM. algorithm, that allow for docking of polypeptides on target molecules can be used, for example, to predict the binding energy associated with the polypeptide-target interaction. Such predictions provide reasonable estimates for the binding energies, but they are not necessarily equal to the binding energies that would be calculated by testing the polypeptides and protein targets in vitro. In that regard, the binding energies identified herein were all generated using the ClusPro.TM. algorithm. Accordingly, absent indication to the contrary, any numerical reference to the binding energy associated with a peptide binding to a particular target is a reference to a binding energy determined by modeling the interaction using the ClusPro.TM. algorithm.
[0126] As detailed in the Examples below, the exemplary RP peptides have been shown to interact with various signaling molecules associated with inflammation, including NF-kB Class II subunit RelB, TGF.beta., Notch1, Wnt8R, TRAIL, IL6R, IL10R, EGFR, and CDK6, as well as other membrane associated signaling molecules, including CD206, CD47 and SIRP-.alpha., translational modification protein transglutaminase 2 (TGM2), and histone modification enzyme histone methyl transferase (HMT). Upon folding of these protein targets to their normal 3-dimensional conformations, an amphipathic cleft is often generated that has high affinity for the immune-modulating peptides herein described.
[0127] For modeling interactions between potential anti-inflammatory polypeptides and NF-kB Class II subunits, any Class II subunit sequence can be used (e.g., RelA, RelB, cRel, NF-kB1, or NF-kB2). In certain embodiments, the Class II subunit sequence folds into a functional Class II subunit or a functional fragment thereof. The particular Class II subunit used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human NF-kB Class II subunit is selected if the intended subject is a human, a bovine NF-kB Class II subunit is selected if the intended subject is a cow, etc.). The NF-kB Class II subunit sequence used for modeling can be the human RelB sequence (NCBI Accession No. NP-006500), which is as follows:
TABLE-US-00003 (SEQ ID NO: 367) MLRSGPASGPSVPTGRAMPSRRVARPPAAPELGAL GSPDLSSLSLAVSRSTDELEIIDEYIKENGFGLDGGQ PGPGEGLPRLVSRGAASLSTVTLGPVAPPATPPPWG CPLGRLVSPAPGPGPQPHLVITEQPKQRGMRFRYEC EGRSAGSILGESSTEASKTLPAIELRDCGGLREVEVT ACLVWKDWPHRVHPHSLVGKDCTDGICRVRLRPHV SPRHSFNNLGIQCVRKKEIEAAIERKIQLGIDPYNAGSLK NHQEVDMNVVRICFQASYRDQQGQMRRMDPV LSEPVYDKKSTNTSELRICRINKESGPCTGGEE ##STR00001## TDGVCSEPLPFTYLPRDHDSYGVDKKRKRGMPDVLG ELNSSDPHGIESKRRKKKPAILDHFLPNHGSGPFLPPS ALLPDPDFFSGTVSLPGLEPPGGPDLLDDGFAYDPTA PTLFTMLDLLPPAPPHASAVVCSGGAGAVVGETPGP EPLTLDSYQAPGPGDGGTASLVGSNMFPNHYREAAF GGGLLSPGPEAT.
[0128] The underlined sequence in human RelB (above) has been identified as the dimerization domain. The highlighted amino acid residues (Tyr-300, Leu-302, and His-332) are believed to be particularly important in the dimerization interaction.
[0129] An anti-inflammatory polypeptide can be identified based on its ability to bind (e.g., in silico) to the dimerization pocket of the Class II subunit and/or interfere with or block the ability of the Class II subunit to dimerize. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human RelB (SEQ ID NO: 367) selected from the group consisting of Leu-281, Ile-283, Cys-284, Glu-298, Tyr-300, Leu-301, Leu-302, Cys-303, Ile-311, Ser-312, Ala-329, Asp-330, Val-331, His-332, Gln-334, and Leu-371, or the equivalent amino acid residue(s) in a different human NF-kB Class II protein or an NF-kB Class II protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human RelB (SEQ ID NO: 367) selected from the group consisting of Glu-298, Tyr-300, Leu-302, Asp-330, Gln-334, and Leu-371 or the equivalent amino acid residue(s) in a different human NF-kB Class II protein or an NF-kB Class II protein of another species.
[0130] In certain embodiments, an anti-inflammatory polypeptide binds to human RelB (SEQ ID NO: 367) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050, -1075, -1100, -1125, -1150, -1200 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0131] For modeling interactions between potential anti-inflammatory polypeptides and TGF.beta., any TGF.beta. protein sequence can be used. The TGF.beta. sequence generally folds into a functional TGF.beta. protein or a functional fragment thereof. The TGF.beta. protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human TGF.beta. is selected if the intended subject is a human, a bovine TGF.beta. is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human TGF.beta. sequence (NCBI Acc. No. NP_000651.3), which is as follows:
TABLE-US-00004 (SEQ ID NO: 368) MPPSGLRLLPLLLPLLWLLVLTPGRPAAGLSTCKTIDMELVKRKRIEAIR GQILSKLRLASPPSQGEVPPGPLPEAVLALYNSTRDRVAGESAEPEPEPE ADYYAKEVTRVLMVETHNEIYDKFKQSTHSIYMFFNTSELREAVPEPVLL SRAELRLLRLKLKVEQHVELYQKYSNNSWRYLSNRLLAPSDSPEWLSFDV TGVVRQWLSRGGEIEGFRLSAHCSCDSRDNTLQVDINGFTTGRRGDLATI HGMNRPFLLLMATPLERAQHLQSSRHRRALDTNYCFSSTEKNCCVRQLYI DFRKDLGWKWIHEPKGYHANFCLGPCPYIWSLDTQYSKVLALYNQHNPGA SAAPCCVPQALEPLPIVYYVGRKPKVEQLSNMIVRSCKCS.
[0132] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the receptor binding site on TGF.beta. and/or interfere with or block the ability of TGF.beta. to bind to its receptor. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human TGF.beta. (SEQ ID NO: 368) selected from the group consisting of Arg-25, Gly-29, Trp-30, Lys-31, Trp-32, Ile-33, His-34, Tyr-91, Val-92, Val-93, Gly-94, Arg-95, Lys-96, and Pro-97, or the equivalent amino acid residue(s) in a TGF.beta. protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human TGF.beta. (SEQ ID NO: 368) selected from the group consisting of Leu-20, Ile-22, Phe-24, Asp-27, Leu-28, Trp-30, Trp-32, Tyr-39, Phe-43, Pro-80, Leu-83, Leu-101 and Ser-112, or the equivalent amino acid residue(s) in a TGF.beta. protein of another species. In other alternatives, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human TGF.beta. (SEQ ID NO: 368) selected from the group consisting of Asp-27, Leu-28, Trp-30, and Trp-32, or the equivalent amino acid residue(s) in a TGF.beta. protein of another species.
[0133] In certain embodiments, an anti-inflammatory polypeptide can bind to human TGF.beta. (SEQ ID NO: 368) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0134] For modeling interactions between potential anti-inflammatory polypeptides and Notch1, any Notch1 protein sequence can be used. The Notch1 sequence used for modeling generally folds into a functional Notch1 protein or a calcium-binding fragment thereof. The Notch1 sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human Notch1 is selected if the intended subject is a human, a bovine Notch1 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human Notch1 sequence (GenBank Acc. No. AAG33848.1), which is as follows:
TABLE-US-00005 (SEQ ID NO: 369) MPPLLAPLLCLALLPALAARGPRCSQPGETCLNGGKCEAANGTEACVCG GAFVGPRCQDPNPCLSTPCKNAGTCHVVDRRGVADYACSCALGFSGPLC LTPLDNACLTNPCRNGGTCDLLTLTEYKCRCPPGWSGKSCQQADPCASN PCANGGQCLPFEASYICHCPPSFHGPTCRQDVNECGQKPRLCRHGGTCH NEVGSYRCVCRATHTGPNCERPYVPCSPSPCQNGGTCRPTGDVTHECAC LPGFTGQNCEENIDDCPGNNCKNGGACVDGVNTYNCPCPPEWTGQYCTE DVDECQLMPNACQNGGTCHNTHGGYNCVCVNGWTGEDCSENIDDCASAA CFHGATCHDRVASFYCECPHGRTGLLCHLNDACISNPCNEGSNCDTNPV NGKAICTCPSGYTGPACSQDVDECSLGANPCEHAGKCINTLGSFECQCL QGYTGPRCEIDVNECVSNPCQNDATCLDQIGEFQCMCMPGYEGVHCEVN TDECASSPCLHNGRCLDKINEFQCECPTGFTGHLCQYDVDECASTPCKN GAKCLDGPNTYTCVCTEGYTGTHCEVDIDECDPDPCHYGSCKDGVATFT CLCRPGYTGHHCETNINECSSQPCRLRGTCQDPDNAYLCFCLKGTTGPN CEINLDDCASSPCDSGTCLDKIDGYECACEPGYTGSMCNSNIDECAGNP CHNGGTCEDGINGFTCRCPEGYHDPTCLSEVNECNSNPCVHGACRDSLN GYKCDCDPGWSGTNCDINNNECESNPCVNGGTCKDMTSGIVCTCREGFS GPNCQTNINECASNPCLNKGTCIDDVAGYKCNCLLPYTGATCEVVLAPC APSPCRNGGECRQSEDYESFSCVCPTAGAKGQTCEVDINECVLSPCRHG ASCQNTHGXYRCHCQAGYSGRNCETDIDDCRPNPCHNGGSCTDGINTAF CDCLPGFRGTFCEEDINECASDPCRNGANCTDCVDSYTCTCPAGFSGIH CENNTPDCTESSCFNGGTCVDGINSFTCLCPPGFTGSYCQHVVNECDSR PCLLGGTCQDGRGLHRCTCPQGYTGPNCQNLVHWCDSSPCKNGGKCWQT HTQYRCECPSGWTGLYCDVPSVSCEVAAQRQGVDVARLCQHGGLCVDAG NTHHCRCQAGYTGSYCEDLVDECSPSPCQNGATCTDYLGGYSCKCVAGY HGVNCSEEIDECLSHPCQNGGTCLDLPNTYKCSCPRGTQGVHCEINVDD CNPPVDPVSRSPKCFNNGTCVDQVGGYSCTCPPGFVGERCEGDVNECLS NPCDARGTQNCVQRVNDFHCECRAGHTGRRCESVINGCKGKPCKNGGTC AVASNTARGFICKCPAGFEGATCENDARTCGSLRCLNGGTCISGPRSPT CLCLGPFTGPECQFPASSPCLGGNPCYNQGTCEPTSESPFYRCLCPAKF NGLLCHILDYSFGGGAGRDIPPPLIEEACELPECQEDAGNKVCSLQCNN HACGWDGGDCSLNFNDPWKNCTQSLQCWKYFSDGHCDSQCNSAGCLFDG FDCQRAEGQCNPLYDQYCKDHFSDGHCDQGCNSAECEWDGLDCAEHVPE RLAAGTLVVVVLMPPEQLRNSSFHFLRELSRVLHTNVVFKRDAHGQQMI FPYYGREEELRKHPIKRAAEGWAAPDALLGQVKASLLPGGSEGGRRRRE LDPMDVRGSIVYLEIDNRQCVQASSQCFQSATDVAAFLGALASLGSLNI PYKIEAVQSETVEPPPPAQLHFMYVAAAAFVLLFFVGCGVLLSRKRRRQ HGQLWFPEGFKVSEASKKKRREPLGEDSVGLKPLKNASDGALMDDNQNE WGDEDLETKKFRFEEPVVLPDLDDQTDHRQWTQQHLDAADLRMSAMAPT PPQGEVDADCMDVNVRGPDGFTPLMIASCSGGGLETGNSEEEEDAPAVI SDFIYQGASLHNQTDRTGETALHLAARYSRSDAAKRLLEASADANIQDN MGRTPLHAAVSADAQGVFQILIRNRATDLDARMHDGTTPLILAARLAVE GMLEDLINSHADVNAVDDLGKSALHWAAAVNNVDAAVVLLKNGANKDMQ NNREETPLFLAAREGSYETAKVLLDHFANRDITDHMDRLPRDIAQERMH HDIVRLLDEYNLVRSPQLHGAPLGGTPTLSPPLCSPNGYLGSLKPGVQG KKVRKPSSKGLACGSKEAKDLKARRKKSQDGKGCLLDSSGMLSPVDSLE SPHGYLSDVASPPLLPSPFQQSPSVPLNHLPGMPDTHLGIGHLNVAAKP EMAALGGGGRLAFETGPPRLSHLPVASGTSTVLGSSSGGALNFTVGGST SLNGQCEWLSRLQSGMVPNQYNPLRGSVAPGPLSTQAPSLQHGMVGPLH SSLAASALSQMMSYQGLPSTRLATQPHLVQTQQVQPQNLQMQQQNLQPA NIQQQQSLQPPPPPPQPHLGVSSAASGHLGRSFLSGEPSQADVQPLGPS SLAVHTILPQESPALPTSLPSSLVPPVTAAQFLTPPSQHSYSSPVDNTP SHQLQVPEHPFLTPSPESPDQWSSSSPHSNVSDWSEGVSSPPTSMQSQI ARIPEAFK.
[0135] An anti-inflammatory polypeptide can be identified based on its ability to bind to the calcium-binding site on Notch1 and/or interfere with or block the ability of Notch1 to bind to calcium. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human Notch1 (SEQ ID NO: 369) selected from the group consisting of Phe-1520, Gln-1523, Arg-1524, Glu-1526, Ala-1553, Glu-1556, Trp-1557, Cys-1562, His-1602, Arg-1684, Gln-1685, Cys-1686, Ser-1691, Cys-1693, Phe-1694, and Phe-1703, or the equivalent amino acid residue(s) in a Notch1 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human Notch1 (SEQ ID NO: 369) selected from the group consisting of Phe-1520, Trp-1557, Cys-1562, and Phe-1703, or the equivalent amino acid residue(s) in a Notch1 protein of another species.
[0136] In certain embodiments, a polypeptide of the invention binds to human Notch1 (SEQ ID NO: 369) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050, -1075 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0137] For modeling interactions between potential anti-inflammatory polypeptides and Wnt8R, any Wnt8R protein sequence can be used. The Wnt8R sequence used for modeling generally folds into a functional Wnt8R protein or a Wnt8-binding fragment thereof. The Wnt8R protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human Wnt8R is selected if the intended subject is a human, a bovine Wnt8R is selected if the intended subject is a cow, etc.). The sequence used for modeling can be, for example, the bovine Wnt8R sequence (NCBI Acc. No. XP_005214377.1), which is as follows:
TABLE-US-00006 (SEQ ID NO: 370) MEWGYLLEVTSLLAALALLQRSSGAAAASAKELACQEITVPLCKGIGYN YTYMPNQFNHDTQDEAGLEVHQFWPLVEIQCSPDLKFFLCSMYTPICLE DYKKPLPPCRSVCERAKAGCAPLMRQYGFAWPDRMRCDRLPEQGNPDTL CMDYNRTDLTTAASSVDGDPVAGICYVGNQSLDNLLGFVLAPLVIYLFI GTMFLLAGFVSLFRIRSVIKQQGGPTKTHKLEKLMIRLGLFTVLYTVPA AVVVACLFYEQHNRPRWEATHNCPCLRDLQPDQARRPDYAVFMLKYFMC LVVGITSGVWVWSGKTLESWRALCTRCCWASKGAGAAGAGAAGGGPGGG GPGAGGGGGPGAGGAGSLYSDVSTGLTWRSGTASSVSYPKQMPLSQV.
[0138] An anti-inflammatory polypeptide can be identified based on its ability to bind to a Wnt ligand-binding site on Wnt8R and/or interfere with or block the ability of Wnt8R to bind to a Wnt ligand (e.g., Wnt8). For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of bovine Wnt8R (SEQ ID NO: 370) selected from the group consisting of Tyr-52, Gln-56, Phe-57, Asn-58, Met-91, Tyr-100, Lys-101, Pro-103, Pro-105, Pro-106, Arg-137 and Asp-145, or the equivalent amino acid residue(s) in a Wnt8R protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of bovine Wnt8R (SEQ ID NO: 370) selected from the group consisting of Tyr-52, Phe-57, Tyr-100, and Asp-145, or the equivalent amino acid residue(s) in a Wnt8R protein of another species.
[0139] In certain embodiments, a polypeptide of the invention binds to bovine Wnt8R (SEQ ID NO: 370) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -875, -900, -925, -950, -975 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0140] For modeling interactions between potential anti-inflammatory polypeptides and TRAIL, any TRAIL protein sequence can be used. The TRAIL sequence used for modeling in certain embodiments folds into a function TRAIL protein or a functional fragment thereof. The TRAIL protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human TRAIL is selected if the intended subject is a human, a bovine TRAIL is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human TRAIL sequence (GenBank Acc. No. EAW78466.1), which is as follows:
TABLE-US-00007 (SEQ ID NO: 371) KEKQQNISPLVRERGPQRVAAHITGTRGRSNTLSSPNSKNEKALGRKINS WESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQEEIKENTKND KQMVQYIYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGGIFELKEN DRIFVSVTNEHLIDMDHEASFFGAFLVG.
[0141] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the receptor binding site on TRAIL and/or interfere with or block the ability of TRAIL to bind to its receptor. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human TRAIL (SEQ ID NO: 371) selected from the group consisting of Arg-130, Arg-158, Ser-159, Gly-160, His-161, Phe-163, Tyr-189, Arg-189, Gln-193, Glu-195, Glu-236, Tyr-237, Leu-239, Asp-267, Asp-269, His-270, and Glu-271, or the equivalent amino acid residue(s) in a TRAIL protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human TRAIL (SEQ ID NO: 371) selected from the group consisting of Ala-123, His-161, Ser-162, Phe-163, Tyr-183, Tyr-185, Tyr-243, His-270, Glu-271, Phe-274, Phe-278, Leu-279, and Val-280, or the equivalent amino acid residue(s) in a TRAIL protein of another species. In other alternatives, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human TRAIL (SEQ ID NO: 371) selected from the group consisting of Phe-163, Tyr-243, Glu-271, and Phe-278, or the equivalent amino acid residue(s) in a TRAIL protein of another species.
[0142] In certain embodiments, an anti-inflammatory polypeptide can bind to human TRAIL (SEQ ID NO: 371) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0143] For modeling interactions between potential anti-inflammatory polypeptides and IL6R, any IL6R protein sequence can be used. The IL6R sequence used for modeling generally folds into a functional IL6R protein or a IL6-binding fragment thereof. The IL6R protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human IL6R is selected if the intended subject is a human, a bovine IL6R is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human IL6R sequence (NCBI Acc. No. NP_786943.1), which is as follows:
TABLE-US-00008 (SEQ ID NO: 372) MLTLQTWLVQALFIFLTTESTGELLDPCGYISPESPVVQLHSNFTAVCVL KEKCMDYFHVNANYIVWKTNHFTIPKEQYTIINRTASSVTFTDIASLNIQ LTCNILTFGQLEQNVYGITIISGLPPEKPKNLSCIVNEGKKMRCEWDGGR ETHLETNFTLKSEWATHKFADCKAKRDTPTSCTVDYSTVYFVNIEVWVEA ENALGKVTSDHINFDPVYKVKPNPPHNLSVINSEELSSILKLTWTNPSIK SVIILKYNIQYRTKDASTWSQIPPEDTASTRSSFTVQDLKPFTEYVFRIR CMKEDGKGYWSDWSEEASGITYEDNIASF.
[0144] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the IL6-binding site on IL6R and/or interfere with or block the ability of IL6R to bind to its ligand, IL6. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human IL6R (SEQ ID NO: 372) selected from the group consisting of Glu-163, Gly-164, Phe-168, Gln-190, Phe-229, Tyr-230, Phe-279 and Gln-281, or the equivalent amino acid residue(s) in a IL6R protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human IL6R (SEQ ID NO: 372) selected from the group consisting of Leu-108, Glu-140, Pro-162, Phe-229, Tyr-230, and Phe-279, or the equivalent amino acid residue(s) in a IL6R protein of another species. In other alternatives, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human IL6R (SEQ ID NO: 372) selected from the group consisting of Glu-140, Phe-229, Tyr-230, Phe-279, or the equivalent amino acid residue(s) in a IL6R protein of another species.
[0145] In certain embodiments, an anti-inflammatory polypeptide can bind to human IL6R (SEQ ID NO: 372) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0146] For modeling interactions between potential anti-inflammatory polypeptides and IL10R, any appropriate IL10R protein sequence can be used. The IL10R sequence used for modeling generally folds into a functional IL10R protein or a IL10-binding fragment thereof. The IL10R protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human IL10R is selected if the intended subject is a human, a bovine IL10R is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human IL10R sequence (NCBI Acc. No. NP_001549.2), which is as follows:
TABLE-US-00009 (SEQ ID NO: 373) MLPCLVVLLAALLSLRLGSDAHGTELPSPPSVWFEAEFFHHILHWTPIPN QSESTCYEVALLRYGIESWNSISNCSQTLSYDLTAVTLDLYHSNGYRARV RAVDGSRHSNWTVTNTRFSVDEVTLTVGSVNLEIHNGFILGKIQLPRPKM APANDTYESIFSHFREYEIAIRKVPGNFTFTHKKVKHENFSLLTSGEVGE FCVQVKPSVASRSNKGMWSKEECISLTRQYFTVTNVIIFFAFVLLLSGAL AYCLALQLYVRRRKKLPSVLLFKKPSPFIFISQRPSPETQDTIHPLDEEA FLKVSPELKNLDLHGSTDSGFGSTKPSLQTEEPQFLLPDPHPQADRTLGN REPPVLGDSCSSGSSNSTDSGICLQEPSLSPSTGPTWEQQVGSNSRGQDD SGIDLVQNSEGRAGDTQGGSALGHHSPPEPEVPGEEDPAAVAFQGYLRQT RCAEEKATKTGCLEEESPLTDGLGPKFGRCLVDEAGLHPPALAKGYLKQD PLEMTLASSGAPTGQWNQPTEEWSLLALSSCSDLGISDWSFAHDLAPLGC VAAPGGLLGSFNSDLVTLPLISSLQSSE.
[0147] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the IL10-binding site on IL10R and/or interfere with or block the ability of IL10R to bind to its ligand, IL10. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human IL10R (SEQ ID NO: 373) selected from the group consisting of Tyr-43, Ile-45, Glu-46, Asp-61, Asn-73, Arg-76, Asn-94, Arg-96, Phe-143, Ala-189, Ser-190, and Ser-191, or the equivalent amino acid residue(s) in a IL6R protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human IL10R (SEQ ID NO: 373) selected from the group consisting of Leu-41, Arg-42, Tyr-43, Ile-45, Glu-46, Ser-47, Trp-48, Arg-76, and Arg-78, or the equivalent amino acid residue(s) in a IL10R protein of another species. In other alternatives, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human IL10R (SEQ ID NO: 373) selected from the group consisting of Tyr-43, Ile-45, Glu-46, Trp-48, or the equivalent amino acid residue(s) in a IL10R protein of another species.
[0148] In certain embodiments, an anti-inflammatory polypeptide can bind to human IL10R (SEQ ID NO: 373) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -775, -800, -825, -850, -875, -900 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0149] For modeling interactions between potential anti-inflammatory polypeptides and EGFR, any EGFR protein sequence can be used. The EGFR sequence used for modeling generally folds into a functional EGFR protein or a ligand-binding fragment thereof. The EGFR protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human EGFR is selected if the intended subject is a human, a bovine EGFR is selected if the intended subject is a cow, etc.). Alternatively, the sequence used for modeling can be the drosophila EGFR sequence (GenBank Acc. No. AAR85273.1), which is as follows:
TABLE-US-00010 (SEQ ID NO: 374) KICIGTKSRLSVPSNKEHHYRNLRDRYTNCTYVDGNLELTWLPNENLDLS FLDNIREVTGYILISHVDVKKVVFPKLQIIRGRTLFSLSVEEEKYALFVT YSKMYTLEIPDLRDVLNGQVGFHNNYNLCHMRTIQWSEIVSNGTDAYYNY DFTAPERECPKCHESCTHGCWGEGPKNCQKFSKLTCSPQCAGGRCYGPKP RECCHLFCAGGCTGPTQKDCIACKNFFDEGVCKEECPPMRKYNPTTYVLE TNPEGKYAYGATCVKECPGHLLRDNGACVRSCPQDKMDKGGECVPCNGPC PKTCPGVTVLHAGNIDSFRNCTVIDGNIRILDQTFSGFQDVYANYTMGPR YIPLDPERLEVFSTVKEITGYLNIEGTHPQFRNLSYFRNLETIHGRQLME SMFAALAIVKSSLYSLEMRNLKQISSGSVVIQHNRDLCYVSNIRWPAIQK EPEQKVWVNENLRADLCEKNGTICSDQCNEDGCWGAGTDQCLNCKNFNFN GTCIADCGYISNAYKFDNRTCKICHPECRTCNGAGADHCQECVHVRDGQH CVSECPKNKYNDRGVCRECHATCDGCTGPKDTIGIGACTTCNLAIINNDA TVKRCLLKDDKCPDGYFWEYVHPQEQGSLKPLAGRAVCRKCHPLCELCTN YGYHEQ.
[0150] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the ligand-binding site on EGFR and/or interfere with or block the ability of at least one ligand to bind to EGFR. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of drosophila EGFR (SEQ ID NO: 374) selected from the group consisting of Leu-10, Thr-40, Trp-41, Asp-48, Phe-51, Leu-63, His-66, Asp-68, Leu-88, and Tyr-101, or the equivalent amino acid residue(s) in a EGFR protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of drosophila EGFR (SEQ ID NO: 374) selected from the group consisting of Trp-41, Asp-48, Phe-51, Asp-68, and Tyr-101, or the equivalent amino acid residue(s) in a EGFR protein of another species.
[0151] In certain embodiments, an anti-inflammatory polypeptide can bind to drosophila EGFR (SEQ ID NO: 374) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0152] For modeling interactions between potential anti-inflammatory polypeptides and CDK6, any CDK6 protein sequence can be used. The CDK6 sequence used for modeling generally folds into a functional CDK6 protein or a functional fragment thereof. The CDK6 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human CDK6 is selected if the intended subject is a human, a bovine CDK6 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human CDK6 sequence (NCBI Acc. No. NP_001250.1), which is as follows:
TABLE-US-00011 (SEQ ID NO: 375) MEKDGLCRADQQYECVAEIGEGAYGKVFKARDLKNGGRFVALKRVRVQTG EEGMPLSTIREVAVLRHLETFEHPNVVRLFDVCTVSRTDRETKLTLVFEH VDQDLTTYLDKVPEPGVPTETIKDMMFQLLRGLDFLHSHRVVHRDLKPQN ILVTSSGQIKLADFGLARIYSFQMALTSVVVTLWYRAPEVLLQSSYATPV DLWSVGCIFAEMFRRKPLFRGSSDVDQLGKILDVIGLPGEEDWPRDVALP RQAFHSKSAQPIEKFVTDIDELGKDLLLKCLTFNPAKRISAYSALSHPYF QDLERCKENLDSHLPPSQNTSELNTA.
[0153] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on CDK6 and/or interfere with or block the kinase activity of CDK6 or the ability of CDK6 to phosphorylate one or more CDK6 substrates. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human CDK6 (SEQ ID NO: 375) selected from the group consisting of Val-142, Arg-144, Asp-145, Ser-171, Val-180, Val-181, Leu-183, Arg-186, Val-190, Gln-193, Tyr-196, and Val-200, or the equivalent amino acid residue(s) in a CDK6 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human CDK6 (SEQ ID NO: 375) selected from the group consisting of Asp-145, Val-180, and Tyr-196, or the equivalent amino acid residue(s) in a CDK6 protein of another species.
[0154] In certain embodiments, an anti-inflammatory polypeptide can bind to human CDK6 (SEQ ID NO: 375) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0155] For modeling interactions between potential anti-inflammatory polypeptides and histone methyl transferase (HMT), any HMT protein sequence can be used. The HMT sequence used for modeling generally folds into a functional HMT protein or a functional fragment thereof. The HMT protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human HMT is selected if the intended subject is a human, a bovine HMT is selected if the intended subject is a cow, etc.). The sequence used for modeling can be, for example, the Paramecium bursaria Chlorella virus 1 HMT sequence (NCBI Acc. No. NP_048968.1), which is as follows:
TABLE-US-00012 (SEQ ID NO: 376) MFNDRVIVKKSPLGGYGVFARKSFEKGELVEECLCIVRHNDDWGTALEDY LFSRKNMSAMALGFGAIFNHSKDPNARHELTAGLKRMRIFTIKPIAIGEE ITISYGDDYWLSRPRLTQN.
[0156] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on HMT and/or interfere with or block the methyl transferase activity of HMT or the ability of HMT to methylate histone substrates. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of Paramecium bursaria HMT (SEQ ID NO: 376) selected from the group consisting of Asn-69, His-70, Ser-71, Lys-72, Asp-73, Pro-74, and Asn-75, or the equivalent amino acid residue(s) in a HMT protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of Paramecium bursaria HMT (SEQ ID NO: 376) selected from the group consisting of Tyr-16, Glu-48, Tyr-50, Leu-51, Phe-52, and Asn-69, or the equivalent amino acid residue(s) in a HMT protein of another species.
[0157] In certain embodiments, an anti-inflammatory polypeptide can bind to Paramecium bursaria HMT (SEQ ID NO: 376) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0158] For modeling interactions between potential anti-inflammatory polypeptides and CD47, any CD47 protein sequence can be used. The CD47 sequence used for modeling generally folds into a functional CD47 protein or a SIRP-.alpha.-binding portion thereof. The CD47 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human CD47 is selected if the intended subject is a human, a bovine CD47 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human CD47 sequence (NCBI Acc. No. XP_005247966.1), which is as follows:
TABLE-US-00013 (SEQ ID NO: 377) MWPLVAALLLGSACCGSAQLLFNKTKSVEFTFCNDTVVIPCFVTNMEAQN TTEVYVKWKFKGRDIYTFDGALNKSTVPTDFSSAKIEVSQLLKGDASLKM DKSDAVSHTGNYTCEVTELTREGETIIELKYRVVSWFSPNENILIVIFPI FAILLFWGQFGIKTLKYRSGGMDEKTIALLVAGLVITVIVIVGAILFVPG EYSLKNATGLGLIVTSTGILILLHYYVFSTAIGLTSFVIAILVIQVIAYI LAVVGLSLCIAACIPMHGPLLISGLSILALAQLLGLVYMKFVE.
[0159] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the SIRP-.alpha.-binding site on HMT and/or interfere with or block the binding of CD47 to SIRP-.alpha.. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of CD47 (SEQ ID NO: 377) selected from the group consisting of Glu-29, Ala-30, Glu-35, Val-36, Tyr-37, Lys-39, Thr-49, Asp-51, Glu-97, Thr-99, Leu-101, Thr-102, Arg-103, Glu-104, and Glu-106, or the equivalent amino acid residue(s) in a CD47 protein of another species. In certain embodiments, the anti-inflammatory polypeptide can bind to at least one amino acid residue of CD47 (SEQ ID NO: 377) selected from the group consisting of Glu-29, Glu-35, Lys-39, Glu-97, Leu-101, Thr-102, Arg-103, Glu-104, and Glu-106, or the equivalent amino acid residue(s) in a CD47 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human CD47 (SEQ ID NO: 377) selected from the group consisting of Tyr-16, Glu-48, Tyr-50, Leu-51, Phe-52, and Asn-6 Tyr-37, Thr-49, Phe-50, Asp-51, Ala-53, Glu-97, Val-98, Glu-100, Leu-101, Thr-102, Glu-104, Glu-106, Gly-107, or the equivalent amino acid residue(s) in a CD47 protein of another species. In certain embodiments, the anti-inflammatory polypeptide can bind to at least one amino acid residue of CD47 (SEQ ID NO: 377) selected from the group consisting of Tyr-37, Glu-97, Glu-100, Leu-101, Glu-104, Glu-106, or the equivalent amino acid residue(s) in a CD47 protein of another species.
[0160] In certain embodiments, an anti-inflammatory polypeptide can bind to human CD47 (SEQ ID NO: 377) with an affinity of at least -550 kcal/mol, and in certain embodiments at least -600, -650, -675, -700, -725, -750, -775, -800 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0161] For modeling interactions between potential anti-inflammatory polypeptides and SIRP-.alpha., any SIRP-.alpha. protein sequence can be used. The SIRP-.alpha. sequence used for modeling generally folds into a functional SIRP-.alpha. protein or a CD47-binding fragment thereof. The SIRP-.alpha. protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human SIRP-.alpha. is selected if the intended subject is a human, a bovine SIRP-.alpha. is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human SIRP-.alpha. sequence (GenBank Acc. No. AAH26692.1), which is as follows:
TABLE-US-00014 (SEQ ID NO: 378) MEPAGPAPGRLGPLLCLLLAASCAWSGVAGEEELQVIQPDKSVSVAAGES AILHCTVTSLIPVGPIQWFRGAGPARELIYNQKEGHFPRVTTVSESTKRE NMDFSISISNITPADAGTYYCVKFRKGSPDTEFKSGAGTELSVRAKPSAP VVSGPAARATPQHTVSFTCESHGFSPRDITLKWFKNGNELSDFQTNVDPV GESVSYSIHSTAKVVLTREDVHSQVICEVAHVTLQGDPLRGTANLSETIR VPPTLEVTQQPVRAENQVNVTCQVRKFYPQRLQLTWLENGNVSRTETAST VTENKDGTYNWMSWLLVNVSAHRDDVKLTCQVEHDGQPAVSKSHDLKVSA HPKEQGSNTAAENTGSNERNIYIVVGVVCTLLVALLMAALYLVRIRQKKA QGSTSSTRLHEPEKNAREITQVQSLDTNDITYADLNLPKGKKPAPQAAEP NNHTEYASIQTSPQPASEDTLTYADLDMVHLNRTPKQPAPKPEPSFSEYA SVQVPRK.
[0162] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the HMT-binding site on SIRP-.alpha. and/or interfere with or block the binding of SIRP-.alpha. to HMT. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of SIRP-.alpha. (SEQ ID NO: 378) selected from the group consisting of Leu-30, Gln-37, Gln-52, Lys-53, Ser-66, Thr-67, Arg-69, Met-72, Phe-74, Lys-96 and Asp-100, or the equivalent amino acid residue(s) in a SIRP-.alpha. protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human SIRP-.alpha. (SEQ ID NO: 378) selected from the group consisting of Tyr-50, Gln-52, Pro-58, Ser-66, Thr-67, and Ser-77, or the equivalent amino acid residue(s) in a SIRP-.alpha. protein of another species. In certain embodiments, the anti-inflammatory polypeptide can bind to at least one amino acid residue of SIRP-.alpha. (SEQ ID NO: 378) selected from the group consisting of Tyr-50, Gln-52, Ser-66, and Thr-67, or the equivalent amino acid residue(s) in a SIRP-.alpha. protein of another species.
[0163] In certain embodiments, an anti-inflammatory polypeptide can bind to human SIRP-.alpha. (SEQ ID NO: 378) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -825, -850, -875, -900, -925, -950, -975, -1000 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0164] For modeling interactions between potential anti-inflammatory polypeptides and CD206, any CD206 protein sequence can be used. The CD206 sequence used for modeling generally folds into a functional CD206 protein or a mannose-binding fragment thereof. The CD206 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human CD206 is selected if the intended subject is a human, a bovine CD206 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human CD206 sequence (NCBI Acc. No. NP_002429.1), which is as follows:
TABLE-US-00015 (SEQ ID NO: 379) MRLPLLLVFASVIPGAVLLLDTRQFLIYNEDHKRCVDAVSPSAVQTAACN QDAESQKFRWVSESQIMSVAFKLCLGVPSKTDWVAITLYACDSKSEFQKW ECKNDTLLGIKGEDLFFNYGNRQEKNIMLYKGSGLWSRWKIYGTTDNLCS RGYEAMYTLLGNANGATCAFPFKFENKWYADCTSAGRSDGWLWCGTTTDY DTDKLFGYCPLKFEGSESLWNKDPLTSVSYQINSKSALTWHQARKSCQQQ NAELLSITEIHEQTYLTGLTSSLTSGLWIGLNSLSFNSGWQWSDRSPFRY LNWLPGSPSAEPGKSCVSLNPGKNAKWENLECVQKLGYICKKGNTTLNSF VIPSESDVPTHCPSQWWPYAGHCYKIHRDEKKIQRDALTTCRKEGGDLTS IHTIEELDFIISQLGYEPNDELWIGLNDIKIQMYFEWSDGTPVTFTKWLR GEPSHENNRQEDCVVMKGKDGYWADRGCEWPLGYICKMKSRSQGPEIVEV EKGCRKGWKKHHFYCYMIGHTLSTFAEANQTCNNENAYLTTIEDRYEQAF LTSFVGLRPEKYFWTGLSDIQTKGTFQWTIEEEVRFTHWNSDMPGRKPGC VAMRTGIAGGLWDVLKCDEKAKFVCKHWAEGVTHPPKPTTTPEPKCPEDW GASSRTSLCFKLYAKGKHEKKTWFESRDFCRALGGDLASINNKEEQQTIW RLITASGSYHKLFWLGLTYGSPSEGFTWSDGSPVSYENWAYGEPNNYQNV EYCGELKGDPTMSWNDINCEHLNNWICQIQKGQTPKPEPTPAPQDNPPVT EDGWVIYKDYQYYFSKEKETMDNARAFCKRNFGDLVSIQSESEKKFLWKY VNRNDAQSAYFIGLLISLDKKFAWMDGSKVDYVSWATGEPNFANEDENCV TMYSNSGFWNDINCGYPNAFICQRHNSSINATTVMPTMPSVPSGCKEGWN FYSNKCFKIFGFMEEERKNWQEARKACIGFGGNLVSIQNEKEQAFLTYHM KDSTFSAWTGLNDVNSEHTFLWTDGRGVHYTNWGKGYPGGRRSSLSYEDA DCVVIIGGASNEAGKWMDDTCDSKRGYICQTRSDPSLTNPPATIQTDGFV KYGKSSYSLMRQKFQWHEAETYCKLHNSLIASILDPYSNAFAWLQMETSN ERVWIALNSNLTDNQYTWTDKWRVRYTNWAADEPKLKSACVYLDLDGYWK TAHCNESFYFLCKRSDEIPATEPPQLPGRCPESDHTAWIPFHGHCYYIES SYTRNWGQASLECLRMGSSLVSIESAAESSFLSYRVEPLKSKTNFWIGLF RNVEGTWLWINNSPVSFVNWNTGDPSGERNDCVALHASSGFWSNIHCSSY KGYICKRPKIIDAKPTHELLTTKADTRKMDPSKPSSNVAGVVIIVILLIL TGAGLAAYFFYKKRRVHLPQEGAFENTLYFNSQSSPGTSDMKDLVGNIEQ NEHSVI.
[0165] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the mannose-binding site on CD206 and/or interfere with or block the binding of SIRP-mannose to CD206. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of CD206 (SEQ ID NO: 379) selected from the group consisting of Glu-725, Tyr-729, Glu-733, Asn-747, and Asp-748, or the equivalent amino acid residue(s) in a CD206 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human CD206 (SEQ ID NO: 379) selected from the group consisting of Phe-726, Thr-727, Trp-728, Pro-733, Glu-737, Asn-738, Trp-739, Ala-740, Glu-743, Tyr-747, Glu-751, Asn-765, Asp-766, or the equivalent amino acid residue(s) in a CD206 protein of another species. In certain embodiments, the anti-inflammatory polypeptide can bind to at least one amino acid residue of CD206 (SEQ ID NO: 379) selected from the group consisting of Phe-726, Trp-728, Trp-739, Glu-743, Tyr-747, Glu-751, or the equivalent amino acid residue(s) in a CD206 protein of another species.
[0166] In certain embodiments, an anti-inflammatory polypeptide can bind to human CD206 (SEQ ID NO: 379) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0167] For modeling interactions between potential anti-inflammatory polypeptides and TGM2, any TGM2 protein sequence can be used. The TGM2 sequence used for modeling generally folds into a functional TGM2 protein or acyl-transferase catalytic fragment thereof. The TGM2 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human TGM2 is selected if the intended subject is a human, a bovine TGM2 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human TGM2 sequence (GenBank Acc. No. AAB95430.1), which is as follows:
TABLE-US-00016 (SEQ ID NO: 380) MMDASKELQVLHIDFLNQDNAVSHHTWEFQTSSPVFRRGQVFHLRLVLNQ PLQSYHQLKLEFSTGPNPSIAKHTLVVLDPRTPSDHYNWQATLQNESGKE VTVAVTSSPNAILGKYQLNVKTGNHILKSEENILYLLFNPWCKEDMVFMP DEDERKEYILNDTGCHYVGAARSIKCKPWNFGQFEKNVLDCCISLLTESS LKPTDRRDPVLVCRAMCAMMSFEKGQGVLIGNWTGDYEGGTAPYKWTGSA PILQQYYNTKQAVCFGQCWVFAGILTTVLRALGIPARSVTGFDSAHDTER NLTVDTYVNENGEKITSMTHDSVWNFHVWTDAWMKRPDLPKGYDGWQAVD ATPQERSQGVFCCGPSPLTAIRKGDIFIVYDTRFVFSEVNGDRLIWLVKM VNGQEELHVISMETTSIGKNISTKAVGQDRRRDITYEYKYPEGSSEERQV MDHAFLLLSSEREHRRPVKENFLHMSVQSDDVLLGNSVNFTVILKRKTAA LQNVNILGSFELQLYTGKKMAKLCDLNKTSQIQGQVSEVTLTLDSKTYIN SLAILDDEPVIRGFIIAEIVESKEIMASEVFTSFQYPEFSIELPNTGRIG QLLVCNCIFKNTLAIPLTDVKFSLESLGISSLQTSDHGTVQPGETIQSQI KCTPIKTGPKKFIVKLSSKQVKEINAQKIVLITK.
[0168] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on TGM2 and/or interfere with or block the acyl-transferase activity of TGM2. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of TGM2 (SEQ ID NO: 380) selected from the group consisting of Cys-277, His-335, and Asp-358, or the equivalent amino acid residue(s) in a TGM2 protein of another species.
[0169] In certain embodiments, an anti-inflammatory polypeptide can bind to human TGM2 (SEQ ID NO: 380) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0170] For modeling interactions between potential anti-inflammatory polypeptides and serum albumin, any serum albumin protein sequence can be used. The serum albumin sequence used for modeling generally folds into a functional serum albumin protein or a functional fragment thereof. The serum albumin protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human serum albumin (HSA) is selected if the intended subject is a human, a bovine serum albumin (BSA) is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human serum albumin (HSA) sequence (NCBI Acc. No. NP_000468.1), which is as follows:
TABLE-US-00017 (SEQ ID NO: 381) DAHKSEVAHRFKDLGEENFKALVLIAFAQYLQQCPFEDHVKLVNEVTEFA KTCVADESAENCDKSLHTLFGDKLCTVATLRETYGEMADCCAKQEPERNE CFLQHKDDNPNLPRLVRPEVDVMCTAFHDNEETFLKKYLYEIARRHPYFY APELLFFAKRYKAAFTECCQAADKAACLLPKLDELRDEGKASSAKQRLKC ASLQKFGERAFKAWAVARLSQRFPKAEFAEVSKLVTDLTKVHTECCHGDL LECADDRADLAKYICENQDSISSKLKECCEKPLLEKSHCIAEVENDEMPA DLPSLAADFVESKDVCKNYAEAKDVFLGMFLYEYARRHPDYSVVLLLRLA KTYETTLEKCCAAADPHECYAKVFDEFKPLVEEPQNLIKQNCELFEQLGE YKFQNALLVRYTKKVPQVSTPTLVEVSRNLGKVGSKCCKHPEAKRMPCAE DYLSVVLNQLCVLHEKTPVSDRVTKCCTESLVNRRPCFSALEVDETYVPK EFNAETFTFHADICTLSEKERQIKKQTALVELVKHKPKATKEQLKAVMDD FAAFVEKCCKADDKETCFAEEGKKLVAASQAALGL.
[0171] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to HSA under physiological conditions (e.g., in the blood stream).
[0172] In certain embodiments, an anti-inflammatory polypeptide can bind to HSA (SEQ ID NO: 381) with an affinity of at least -650 kcal/mol, and in certain embodiments at least -700, -750, -800, -850, -900, -925, -950, -975, -1000, -1025, -1050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0173] In certain embodiments, an anti-inflammatory polypeptide binds to two or more targets (e.g., pro-inflammatory targets). In some embodiments, an anti-inflammatory polypeptide binds to three, four, five, or more pro-inflammatory targets. For example, an anti-inflammatory polypeptide can bind to any combination of targets disclosed herein. Such binding can be based on in silico, in vitro, or in vivo data. Thus, an anti-inflammatory polypeptide can bind to two or more NF-kB Class II subunits (e.g., RelB and at least one other NF-kB Class II subunit, such as RelA, cRel, NF-kB1, or NF-kB2). Alternatively (or in addition), an anti-inflammatory polypeptide can bind to an NF-kB Class II subunit (e.g., RelB) and at least one other signaling molecule (e.g., at least one signaling molecule selected from the group consisting of TGF.beta., Notch1, Wnt8R, TRAIL, IL6R, IL10R, EGFR, CDK6, CD206, CD47, SIRP-.alpha., HMT, and TGM2). For example, an anti-inflammatory polypeptide can bind to an NF-kB Class II subunit (e.g., RelB) and at least one signaling molecule selected from the group consisting of TGF.beta., Notch1, Wnt8R, TRAIL, IL6R, IL10R, EGFR, and CDK6. Alternatively, an anti-inflammatory polypeptide can bind to an NF-kB Class II subunit (e.g., RelB) and at least one signaling molecule selected from the group consisting of CD206, CD47, SIRP-.alpha., and TGM2. In other alternatives, an anti-inflammatory polypeptide can bind to an NF-kB Class II subunit (e.g., RelB) and HMT. In other alternatives, an anti-inflammatory polypeptide can bind to at least one signaling molecule selected from the group consisting of TGF.beta., Notch1, Wnt8R, TRAIL, IL6R, IL10R, EGFR, and CDK6, and at least one signaling molecule selected from the group consisting of CD206, CD47, SIRP-.alpha., and TGM2. In other alternatives, an anti-inflammatory polypeptide can bind to at least one signaling molecule selected from the group consisting of TGF.beta., Notch1, Wnt8R, TRAIL, IL6R, IL10R, EGFR, and CDK6, and also bind to HMT. In still other embodiments, an anti-inflammatory polypeptide can bind to an NF-kB Class II subunit (e.g., RelB), at least one signaling molecule selected from the group consisting of TGF.beta., Notch, Wnt8R, TRAIL, IL6R, IL10R, EGFR, and CDK6, at least one signaling molecule selected from the group consisting of CD206, CD47, SIRP-.alpha., and TGM2, and also HMT. In certain embodiments, an anti-inflammatory polypeptide binds to two or more pro-inflammatory targets and also serum albumin (e.g., human serum albumin).
[0174] For modeling interactions between potential anti-inflammatory polypeptides and LEGUMAIN, any LEGUMAIN protein sequence can be used. The LEGUMAIN sequence used for modeling generally folds into a functional LEGUMAIN protein or a functional fragment thereof. The LEGUMAIN protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human LEGUMAIN is selected if the intended subject is a human, a bovine LEGUMAIN is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human LEGUMAIN sequence (GenBank Acc. No. AAH03061.1).
TABLE-US-00018 (SEQ ID NO: 137) ##STR00002## QIVVMMYDDIAYSEDNPTPGIVINRPNGTDVYQGVP KDYTGEDVTPQNFLAVLRGDAEAVKGIG ##STR00003## ##STR00004## LVKSHTNTSHVMQYGNKTISTMKVMQFQGMKRKASS PVPLPPVTHLDLTPSPDVPLTIMKRKLMNTNDLEESR QLTEEIQRHLDARHLIEKSVRKIVSLLAASEAEVEQLLS ERAPLTGHSCYPEALLHFRTHCFNWHSPTYEYALRHLY VLVNLCEKPYPLHRIKLSMDHVCLGHY.
[0175] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on LEGUMAIN and/or interfere with or block the ability of LEGUMAIN to bind to its target. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human LEGUMAIN (SEQ ID NO: 137) selected from the group consisting of Asn-44, Arg-46, His-159, Glu-189, Cys-191, Ser-217, Ser-218 and Asp-233, or the equivalent amino acid residue(s) in a LEGUMAIN protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human LEGUMAIN (SEQ ID NO: 137) selected from the group consisting of Asn-44, Glu-189 and Asp-233, or the equivalent amino acid residue(s) in a LEGUMAIN protein of another species.
[0176] In certain embodiments, an anti-inflammatory polypeptide can bind to human LEGUMAIN (SEQ ID NO: 137) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0177] For modeling interactions between potential anti-inflammatory polypeptides and CD209, any CD209 protein sequence can be used. The CD209 sequence used for modeling generally folds into a functional CD209 protein or a functional fragment thereof. The CD209 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human CD209 is selected if the intended subject is a human, a bovine CD209 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human CD209 sequence (GenBank Acc. No. NP_001138366.1).
TABLE-US-00019 (SEQ ID NO: 140) MSDSKEPRLQQLGLLVSKVPSSISQEQSRQDAIYQNL TQLKAAVGELSEKSKLQEIYQELTQLKAAVGELPEK SKLQEIYQELTRLKAAVGELPEKSKLQEIYQELTWL KAAVGELPEKSKMQEIYQELTRLKAAVGELPEKSK QQEIYQELTRLKAAVGELPEKSKQQEIYQELTRLK AAVGELPEKSKQQEIYQELTQLKAAVERLCHPCPW EWTFFQGNCYFMSNSQRNWHDSITACKEVGAQLV VIKSAEEQ ##STR00005## ##STR00006##
[0178] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on CD209 and/or interfere with or block the ability of CD209 to bind to its receptor. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human CD209 (SEQ ID NO: 140) selected from the group consisting of Phe-269, Glu-280, Glu-303, Asn-305, Asn-306, Glu-310, Asp-311, Ser-316, Gly-317, Asn-321 and Lys-324 or the equivalent amino acid residue(s) in a CD209 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human CD209 (SEQ ID NO: 140) selected from the group consisting of Phe-269, Glu-280, Glu-303, Glu-310, Asp-311, Asn-321 and Lys-324, or the equivalent amino acid residue(s) in a CD209 protein of another species.
[0179] In certain embodiments, an anti-inflammatory polypeptide can bind to human CD209 (SEQ ID NO: 140) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -1,000, -1,050 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0180] For modeling interactions between potential anti-inflammatory polypeptides and FAS, any FAS protein sequence can be used. The FAS sequence used for modeling generally folds into a functional FAS protein or a functional fragment thereof. The FAS protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human FAS is selected if the intended subject is a human, a bovine FAS is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human FAS sequence (NCBI Reference Sequence: NP_000034.1).
TABLE-US-00020 (SEQ ID NO: 152) MLGIWTLLPLVLTSVARLSSKSVNAQVTDINSKGLE LRKTVTTVETQNLEGLHHDGQFCHKPCPPGERKAR DCTVNGDEPDCVPCQEGKEYTDKAHFSSKCRRCRL CDEGHGLEVEINCTRTQNTKCRCKPNFFCNSTVCE HCDPCTKCEHGIIKECTLTSNTKCKEEGSRSNLGWL CLLLLPIPLIVWVKR ##STR00007## ##STR00008## DSENSNFRNEIQSLV.
[0181] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on FAS and/or interfere with or block the ability of FAS to bind to its ligand. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human FAS (SEQ ID NO: 152) selected from the group consisting of Lys-251, Lys-296, Lys-299, Leu-303, Leu-306, Ala-307, Glu-308, Lys-309, Gln-311, Ile-314, Leu-315, Asp-317, Ile-318 and Thr-319, or the equivalent amino acid residue(s) in a FAS protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human FAS (SEQ ID NO: 152) selected from the group consisting of Lys-296, Lys-299, Leu-306, Ala-307, Glu-308, Ile-314, Leu-315, Asp-317 and Ile-318, or the equivalent amino acid residue(s) in a FAS protein of another species.
[0182] In certain embodiments, an anti-inflammatory polypeptide can bind to human FAS (SEQ ID NO: 152) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0183] Programmed cell death protein 1, also known as PD-1 and CD279 (cluster of differentiation 279), is a protein that in humans is encoded by the PDCD1 gene. PD-1 is a cell surface receptor that belongs to the immunoglobulin superfamily and is expressed on T cells and pro-B cells. PD-1 binds two ligands, PD-L1 and PD-L2. PD-1, functioning as an immune checkpoint plays an important role in down regulating the immune system by preventing the activation of T-cells, which in turn reduces autoimmunity and promotes self-tolerance. The inhibitory effect of PD-1 is accomplished through a dual mechanism of promoting apoptosis (programmed cell death) in antigen specific T-cells in lymph nodes while simultaneously reducing apoptosis in regulatory T cells (suppressor T cells).
[0184] For modeling interactions between potential anti-inflammatory polypeptides and PD-1, any PD-1 protein sequence can be used. The PD-1 sequence used for modeling generally folds into a functional PD-1 protein or a functional fragment thereof. The PD-1 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human PD-1 is selected if the intended subject is a human, a bovine PD-1 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human PD-1 sequence (Locus: XP_006712636.1).
TABLE-US-00021 (SEQ ID NO: 159) ##STR00009## ##STR00010## APKAQIKESLRAELRVTERRAEVPTAHPSPSPRPAG QFQTLVVGVVGGLLGSLVLLVWVLAVICSRAARG.
[0185] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on PD-1 and/or interfere with or block the ability of PD-1 to bind to its receptor. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human PD-1 (SEQ ID NO: 159) selected from the group consisting of Val-64, Asn-66, Tyr-68, Met-70, Thr-76, Lys-78, Thr-120, Leu-122, Ala-125, Ser-127, or the equivalent amino acid residue(s) in a PD-1 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human PD-1 (SEQ ID NO: 159) selected from the group consisting of Tyr-68, Met-70, Lys-78 and Leu-122, or the equivalent amino acid residue(s) in a PD-1 protein of another species.
[0186] In certain embodiments, an anti-inflammatory polypeptide can bind to human PD-1 (SEQ ID NO: 159) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -1,000 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0187] Dual specificity mitogen-activated protein kinase kinase 7, also known as MAP kinase kinase 7 or MKK7, is an enzyme that in humans is encoded by the MAP2K7 gene. This protein is a member of the mitogen-activated protein kinase kinase family. The MKK7 protein exists as six different isoforms with three possible N-termini (.alpha., .beta., and .gamma. isoforms) and two possible C-termini (1 and 2 isoforms). MKK7 is involved in signal transduction mediating the cell responses to proinflammatory cytokines, and environmental stresses. This kinase specifically activates MAPK8/JNK1 and MAPK9/JNK2, and this kinase itself is phosphorylated and activated by MAP kinase kinase kinases including MAP3K1/MEKK1, MAP3K2/MEKK2, MAP3K3/MEKK5, and MAP4K2/GCK.
[0188] For modeling interactions between potential anti-inflammatory polypeptides and MKK7, any MKK7 protein sequence can be used. The MKK7 sequence used for modeling generally folds into a functional MKK7 protein or a functional fragment thereof. The MKK7 protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human MKK7 is selected if the intended subject is a human, a bovine MKK7 is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the human MKK7 sequence (NCBI Reference Sequence: NP_001284484.1).
TABLE-US-00022 (SEQ ID NO: 166) MAASSLEQKLSRLEAKLKQENREARRRIDLNLDISP QRPRPIIVITLSPAPAPSQRAALQLPLANDGGSRSPS SESSPQHPTPPARPRHMLGLPSTLFTPRSMESIEIDQ KLQEIMKQTGYLTIGGQR ##STR00011## ##STR00012## ##STR00013## VWSLGISLVELATGQFPYKNCKTDFEVLTKVLQEEPP LLPGHMGFSGDFQSFVKDCLTKDHRKRPKYNKLLEH SFIKRYETLEVDVASWFKDVMAKTESPRTSGVLSQPH LPFFR.
[0189] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on MKK7 and/or interfere with or block the ability of MKK7 to bind to its receptor. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human MKK7 (SEQ ID NO: 166) selected from the group consisting of Met-142, Val-150, Lys-152, Lys-165, Met-212, Met-215, Thr-217, Lys-221, Leu-266, Cys-276 and Asp-277, or the equivalent amino acid residue(s) in a MKK7 protein of another species. Alternatively, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human MKK7 (SEQ ID NO: 166) selected from the group consisting of Met-142, Val-150, Lys-165, Met-212, Met-215, Leu-266 and Asp-277, or the equivalent amino acid residue(s) in a MKK7 protein of another species.
[0190] In certain embodiments, an anti-inflammatory polypeptide can bind to human MKK7 (SEQ ID NO: 166) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -1,000 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0191] For modeling interactions between potential anti-inflammatory polypeptides and ribonucleotide reductase (RNR), any RNR protein sequence can be used. The RNR sequence used for modeling generally folds into a functional RNR protein or a functional fragment thereof. The RNR protein sequence used for modeling can be selected based on the type of subject that the anti-inflammatory polypeptide is intended to treat (e.g., a human RNR is selected if the intended subject is a human, a bovine RNR is selected if the intended subject is a cow, etc.). The sequence used for modeling can be the yeast RNR sequence (GenBank: AJV34160.1).
TABLE-US-00023 (SEQ ID NO: 168) MYVYKRDGRKEPVQFDKITARISRLCYGLDPKHIDA VKVTQRIISGVYEGVTTIELDNLAAETCAYMTTVHPD YATLAARIAISNLHKQTTKQFSKVVEDLYRYVNAATG KPAPMISDDVYNIVMENKDKLNSAIVYDRDFQYSYFG FKTLERSYLLRINGQVAERPQHLIMRVALGIHGRDIEA ALETYNLMSLKYYTHASPTLFNAGTPKPQMSSCFLVA MKEDSIEGIYDTLKECALISKTAGGIGLHIHNIRSTGSY IAGTNGTSNGLIPMIRVFNNTARYVDQGGNKRPGAFA LYLEPWHADIFDFIDIRKNHGKEEIRARDLFPALWIPDL FMKRVEENGTWTLFSPTSAPGLSDCYGDEFEALYTR YEKEGRGKTIK ##STR00014## ALPAFIETSEDGKTSTYNFKKLHEIAKVVTRNLNRVID RNYYPVEEARKSNMRHRPIALGVQGLADTFMLLRLP FDSEEARLLNIQIFETIYHASMEASCELAQKDGPYETF QGSPASQGILQFDMWD ##STR00015## VNPYLLRDLVDLGIWDEGMKQYLITQNGSIQGLPNVP QELKDLYKTVWEISQKTIINMAADRSVYIDQSHSLNL FLRAPTMGKLTSMHFYGWKKGLKTGMYYLRTQAAS AAIQFTIDQKIADQATENVADISNLKRPSYMPSSASYA ASDFVPAAVTANATIPSLDSSSEASREASPAPTGSHSL TKGMAELNVQESKVEVPEVPAPTKNEEKAAPIVDDEE TEFDIYNSKVIACAIDNPEACEMCSG.
[0192] An anti-inflammatory polypeptide can be identified, for example, based on its ability to bind to the active site on RNR and/or interfere with or block the ability of RNR to bind to its receptor. For example, the anti-inflammatory polypeptide can bind to at least one amino acid residue of human RNR (SEQ ID NO: 168) selected from the group consisting of Asn-426, Leu-427, Cys-428, Glu-430, Met-606, Pro-608 and Ala-610, or the equivalent amino acid residue(s) in a RNR protein of another species.
[0193] In certain embodiments, an anti-inflammatory polypeptide can bind to human RNR (SEQ ID NO: 168) with an affinity of at least -600 kcal/mol, and in certain embodiments at least -650, -700, -750, -800, -850, -900, -925, -950, -1,000 kcal/mol, or greater. The requisite binding affinity can correspond to a binding affinity that can be detected in vitro or in vivo. Alternatively, the requisite binding affinity can correspond to a binding affinity that can be detected in silico, e.g., using the ClusPro.TM. algorithm.
[0194] Excluded Polypeptides
[0195] Compositions of the invention optionally exclude polypeptides that satisfy the Structural Algorithm described herein which may have been known in the art prior to the filing of the present application. Various publications have discussed synthetic and naturally occurring anti-inflammatory polypeptides and/or polypeptides having a striapathic sequence including, for example, US Patent Application Nos. 2012/0270770 and 2003/0109452, and U.S. Pat. No. 6,559,281. Accordingly, one or more polypeptides and/or uses of such polypeptides described in such publications can be excluded from the scope of the presently disclosed composition and/or methods. For example, peptide RP-398 (SEQ ID NO: 155) is optionally excluded from compositions disclosed herein and/or methods of using such compositions. Moreover, any of the polypeptides disclosed in Tables 3-9, below, can be optionally excluded from compositions disclosed herein and/or methods of using such compounds.
[0196] Linked Anti-Inflammatory Polypeptide Combinations
[0197] The invention further includes any two anti-inflammatory polypeptides which have been linked together. The linkage can be formed by a peptide linker, such as a Gly-Gly-Gly (GGG), Gly-Gly-Gly-Arg (GGGR; SEQ ID NO: 412), Gly-Pro-Gly (GPG), or Gly-Pro-Gly-Arg (GPGR; SEQ ID NO: 413) sequence, that links the C-terminal end of a first anti-inflammatory polypeptide to the N-terminal end of a second anti-inflammatory polypeptide. Alternatively, the linkage can be a peptoid linker (e.g., a poly N-substituted version of any of the foregoing peptide linkers), a polymer containing g-amino acids (e.g., corresponding to any of the foregoing peptide linkers), or a non-peptide, chemical linker. The linked anti-inflammatory polypeptides can be any of the polypeptides disclosed herein (e.g., in Tables 3-9), and can include the same polypeptide being linked to form a homodimer or different polypeptides being linked to form a heterodimer. Techniques for linking peptides via peptide and non-peptide linkers are well known in the art, and the inventive polypeptide combinations are intended to encompass all such linkages.
[0198] Anti-inflammatory polypeptides can be linked to another molecule via a biodegradable linkage, such as a disulfide bond. The disulfide bond can be mediated by the sulfhydryl group of a cysteine residue found in the anti-inflammatory polypeptide and a sulfhydryl group in the other molecule. The cysteine residue can be, e.g., located at either the C-terminal or N-terminal end of anti-inflammatory polypeptide. Specific examples include RP-433 (FAKKFAKKFKC, SEQ ID NO: 384) and RP-434 (KFRKAFKRFFC; SEQ ID NO: 385), though any of the peptides disclosed herein could be similarly modified. Using a disulfide linkage of this sort, polypeptides of the invention can be conveniently linked to various types of useful molecules. For example, the linkage can be with another anti-inflammatory polypeptide (which optionally includes a C-terminal or N-terminal cysteine residue), a fluorescent label (e.g., Dylight 350), a chemotherapeutic agent (e.g., a taxol derivative formed by adding a sulfhydral group to an appropriate site on the taxol ring structure, followed by oxidation with a cysteine-containing peptide of the invention), or the like.
[0199] Linked anti-inflammatory polypeptides (e.g., homo- or heterodimers) can bind to a target molecule (e.g., a target protein, such as a pro-inflammatory signaling protein) with a binding energy that is greater than that of either monomer polypeptide alone. Thus, for example, the energy of binding of linked anti-inflammatory polypeptides to an NF-kB Class II protein (e.g., RelB) can be at least -700 kcal/mol, and in certain embodiments at least -750, -800, -900, -1000, -1100, -1200, -1250, -1300, -1350, -1400, -1425, -1450, -1475, -1500, -1525, -1550, -1575, -1600 kcal/mol, or greater. The energy of binding can be determined, e.g., in silico, in vitro, or in vivo, using methods well-known in the art (e.g., using the ClusPro.TM. algorithm).
[0200] Modified Polypeptides
[0201] Embodiments of the invention include the modification of any of the anti-inflammatory polypeptides of the invention, by chemical or genetic means. Examples of such modification include construction of peptides of partial or complete sequence with non-natural amino acids and/or natural amino acids in L or D forms. For example, any of the peptides disclosed herein and any variants thereof could be produced in an all-D form. Furthermore, polypeptides of the invention can be modified to contain carbohydrate or lipid moieties, such as sugars or fatty acids, covalently linked to the side chains or the N- or C-termini of the amino acids. In addition, the polypeptides of the invention can be modified to enhance solubility and/or half-life upon being administered. For example, polyethylene glycol (PEG) and related polymers have been used to enhance solubility and the half-life of protein therapeutics in the blood. Accordingly, the polypeptides of the invention can be modified by PEG polymers and the like. Polypeptides of the invention can also be modified to contain sulfur, phosphorous, halogens, metals, etc. And amino acid mimics can be used to produce polypeptides of the invention (e.g., having a structure based on the Structural Algorithm or a structure similar to any of the anti-inflammatory polypeptides disclosed herein). In certain embodiments, polypeptides of the invention that include amino acid mimics have enhanced properties, such as resistance to degradation. For example, polypeptides of the invention can include one or more (e.g., all) peptoid monomers.
[0202] Compositions
[0203] Compositions of the invention include an anti-inflammatory polypeptide that satisfies the structural algorithm described herein. For example, the anti-inflammatory polypeptide can have a striapathic region having a sequence that conforms with any one of Formulas I-LIV. In particular, the anti-inflammatory polypeptide can be any of the polypeptides listed in Table 3-9, or a fragment or variant thereof. Typically, the anti-inflammatory polypeptide included in the compositions of the invention will be a synthetic polypeptide (e.g., made by chemical synthesis and/or produced recombinantly).
[0204] The compositions of the invention can include a single anti-inflammatory polypeptide, or combinations thereof. The compositions can be substantially free of proteins and other polypeptides that do not satisfy the structural algorithm disclosed herein. As used herein, the term "substantially free of proteins and other polypeptides" means that less than 5% of the protein content of the composition is made up of proteins and other polypeptides that are not an anti-inflammatory polypeptide of the invention. A composition that is substantially free of non-anti-inflammatory polypeptides of the invention can have less than 4%, 3%, 2%, 1%, 0.5%, 0.1%, 0.05%, 0.01%, or less of proteins or other polypeptides that do not satisfy the structural algorithm disclosed herein. Thus, the compositions can be substantially free of blood proteins, such as serum albumin, globulins, fibrinogen, and clotting factors. Alternatively, the compositions can be substantially free of globulins, fibrinogen, and clotting factors, but can include purified or recombinantly produced serum albumin.
[0205] The compositions of the invention in certain embodiments contain an anti-inflammatory polypeptide that is not naturally found in a human or other mammal or animal. However, compositions of the invention can include an anti-inflammatory polypeptide that is naturally found in a human or other mammal or animal, provided that the composition is substantially free of biological molecules (such as non-anti-inflammatory polypeptides, nucleic acids, lipids, carbohydrates, and metabolites) that are associated with the anti-inflammatory polypeptide in vivo or co-purify with the anti-inflammatory polypeptide. As used herein, the term "substantially free of biological molecules" means that less than 5% of the dry weight of the composition is made up of biological molecules that are not anti-inflammatory polypeptides. A composition that is substantially free of such biological molecules can have less than 4%, 3%, 2%, 1%, 0.5%, 0.1%, 0.05%, 0.01%, or less of biological molecules that are not anti-inflammatory polypeptides. Thus, for example, the composition can be substantially free of biological molecules that are abundant in the blood, such the proteins discussed above, fatty acids, cholesterol, non-protein clotting factors, metabolites, and the like. In addition, the composition can be substantially free of cells, including red blood cells, white blood cells, and platelets, and cell fragments.
[0206] The compositions of the invention can include at least 1 mg (e.g., at least 5, 10, 20, 30, 40, 50, 75, 100, 150, 200, 250, 300, 400, 500, 600, 700, 800, 900, 1000 mg, or more) of anti-inflammatory polypeptide. Thus, for example, the compositions can include an amount of anti-inflammatory polypeptide equal to about 1 mg to about 1000 mg (e.g., about 5 mg to about 900 mg, about 5 mg to about 800 mg, about 5 mg to about 700 mg, about 5 mg to about 600 mg, about 10 mg to about 500 mg, about 10 mg to about 400 mg, about 10 mg to about 300 mg, about 10 mg to about 250 mg, about 10 mg to about 200 mg, about 10 mg to about 150 mg, about 10 mg to about 100 mg, about 50 mg to about 500 mg, about 50 mg to about 400 mg, about 50 mg to about 300 mg, about 50 mg to about 250 mg, about 50 mg to about 200 mg, about 50 mg to about 150 mg, about 50 mg to about 100 mg, about 75 mg to about 500 mg, about 75 mg to about 400 mg, about 75 mg to about 300 mg, about 75 mg to about 250 mg, about 75 mg to about 200 mg, about 75 mg to about 150 mg, about 75 mg to about 100 mg, about 100 mg to about 500 mg, about 100 mg to about 400 mg, about 100 mg to about 300 mg, about 100 mg to about 250 mg, about 100 mg to about 200 mg, or any other range containing two of the foregoing endpoints).
[0207] The compositions of the invention can include a solution that contains at least 1 mg/ml (e.g., at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, 95, 100 mg/ml or more) of an anti-inflammatory polypeptide. Thus, for example, the compositions can include a solution having an anti-inflammatory polypeptide concentration of about 1 mg/ml to about 1000 mg/ml (e.g., about 5 mg/ml to about 900 mg/ml, about 5 mg/ml to about 800 mg/ml, about 5 mg/ml to about 700 mg/ml, about 5 mg/ml to about 600 mg/ml, about 5 mg/ml to about 500 mg/ml, about 10 mg/ml to about 500 mg/ml, about 10 mg/ml to about 400 mg/ml, about 10 mg/ml to about 300 mg/ml, about 10 mg/ml to about 250 mg/ml, about 10 mg/ml to about 200 mg/ml, about 10 mg/ml to about 150 mg/ml, about 10 mg/ml to about 100 mg/ml, about 50 mg/ml to about 500 mg/ml, about 50 mg/ml to about 400 mg/ml, about 50 mg/ml to about 300 mg/ml, about 50 mg/ml to about 250 mg/ml, about 50 mg/ml to about 200 mg/ml, about 50 mg/ml to about 150 mg/ml, about 50 mg/ml to about 100 mg/ml, about 75 mg/ml to about 500 mg/ml, about 75 mg/ml to about 400 mg/ml, about 75 mg/ml to about 300 mg/ml, about 75 mg/ml to about 250 mg/ml, about 75 mg/ml to about 200 mg/ml, about 75 mg/ml to about 150 mg/ml, about 75 mg/ml to about 100 mg/ml, about 100 mg/ml to about 500 mg/ml, about 100 mg/ml to about 400 mg/ml, about 100 mg/ml to about 300 mg/ml, about 100 mg/ml to about 250 mg/ml, about 100 mg/ml to about 200 mg/ml, about 10 mg/ml to about 150 mg/ml, or any other range containing two of the foregoing endpoints).
[0208] The compositions of the invention include pharmaceutical compositions. Such pharmaceutical compositions can comprise one or more anti-inflammatory polypeptides and a pharmaceutically acceptable carrier. Pharmaceutical compositions can further include a protein other than an anti-inflammatory polypeptide of the invention and/or a chemotherapeutic agent. The other protein can be a therapeutic agent, such as a therapeutic antibody. The therapeutic protein or antibody can have anti-inflammatory properties or other properties that the anti-inflammatory polypeptides of the invention augment or are augmented by. Alternatively, the other protein can be a carrier protein, such as serum albumin (e.g., HSA). The serum albumin (e.g., HAS, BSA, etc.) can be purified or recombinantly produced. By mixing the anti-inflammatory polypeptide(s) in the pharmaceutical composition with serum album, the anti-inflammatory polypeptides can be effectively "loaded" onto the serum albumin, allowing a greater amount of anti-inflammatory polypeptide to be successfully delivered to a site of inflammation. The chemotherapeutic agent can be, for example, an anti-cancer chemotherapeutic agent. Such chemotherapeutic agents include, but are not limited to, Gemcitabine, Docetaxel, Bleomycin, Erlotinib, Gefitinib, Lapatinib, Imatinib, Dasatinib, Nilotinib, Bosutinib, Crizotinib, Ceritinib, Trametinib, Bevacizumab, Sunitinib, Sorafenib, Trastuzumab, Ado-trastuzumab emtansine, Rituximab, Ipilimumab, Rapamycin, Temsirolimus, Everolimus, Methotrexate, Doxorubicin, Abraxane, Folfirinox, Cisplatin, Carboplatin, 5-fluorouracil, Teysumo, Paclitaxel, Prednisone, Levothyroxine, and Pemetrexed.
[0209] Methods
[0210] The anti-inflammatory polypeptides of the invention provide powerful tools for reducing inflammation and/or treating conditions associated with excessive inflammation (whether acute or chronic). As used herein, the terms "treat," "treating," and similar words shall mean stabilizing, reducing the symptoms of, preventing the occurrence of, or curing a medical condition.
[0211] Accordingly, the invention provides methods of reducing the expression level and/or activity of at least one (e.g., 2, 3, 4, 5, or more) pro-inflammatory cytokine(s) at a site of inflammation in a subject. The methods include administering an anti-inflammatory polypeptide of the invention (or, for example, a pharmaceutical composition comprising an anti-inflammatory polypeptide) to the subject. The pro-inflammatory cytokine can be selected from the group consisting of NF-kB, TNF.alpha., IL-1, IL-6, IL-8, IL-12, IL-17, IL-23, MCP-1, MMP-1, and MMP-9. The reduction can be a reduction of at least 10% (e.g., 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, or more) in the expression or activity of the cytokine.
[0212] The invention also provides methods of inhibiting an increase in the expression level and/or activity of at least one (e.g., 2, 3, 4, 5, or more) pro-inflammatory cytokine(s) at a potential site of inflammation in a subject. The methods include administering an anti-inflammatory polypeptide of the invention (or, for example, a pharmaceutical composition comprising an anti-inflammatory polypeptide) to the subject. The pro-inflammatory cytokine can be selected from the group consisting of NF-kB, TNF.alpha., IL-1, IL-6, IL-8, IL-12, IL-17, IL-23, MCP-1, MMP-1, and MMP-9. The methods can inhibit increased cytokine expression and/or activity by limiting such increases to no more than 20% (e.g., 15%, 12.5%, 10%, 7.5%, 5%, 4%, 3%, 2%, 1%, or less).
[0213] The invention also provides a method of treating or preventing a condition associated with chronic inflammation. The condition associated with chronic inflammation can be irritable bowel disease, ulcerative colitis, colitis, Crohn's disease, idiopathic pulmonary fibrosis, asthma, keratitis, arthritis, osteoarthritis, rheumatoid arthritis, auto-immune diseases, a feline or human immunodeficiency virus (FIV or HIV) infection, cancer, age-related inflammation and/or stem cell dysfunction (e.g., age-related increases in Nlrp3 expression, age-related elevation of SOCS3 in muscle stem cells, etc.), graft-versus-host disease (GVHD), keloids, scleroderma, obesity, diabetes, diabetic wounds, other chronic wounds, atherosclerosis, multiple sclerosis, Parkinson's disease, Alzheimer's disease, macular degeneration, gout, gastric ulcers, gastritis, mucositis, toxoplasmosis, and chronic viral or microbial infections (e.g., such as chronic bacterial or protozoan infections). The methods includes administering an anti-inflammatory polypeptide of the invention (or, for example, a pharmaceutical composition comprising an anti-inflammatory polypeptide) to a subject suffering from or likely to develop the condition.
[0214] The invention also provides methods of treating or preventing fibrosis. The fibrosis can be, for example, pulmonary fibrosis, dermal fibrosis, hepatic fibrosis, renal fibrosis, or fibrosis caused by ionizing radiation. The methods include administering an anti-inflammatory polypeptide of the invention (or, for example, a pharmaceutical composition comprising an anti-inflammatory polypeptide) to a subject suffering from or likely to develop fibrosis.
[0215] The invention also provides methods of treating cancer. The cancer can be colon cancer, breast cancer, leukemia, lymphoma, ovarian cancer, prostate cancer, liver cancer, lung cancer, testicular cancer, cervical cancer, bladder cancer, endometrial cancer, kidney cancer, melanoma, cancers of the thyroid or brain, or ophthalmic cancer. The methods include administering an anti-inflammatory polypeptide of the invention (or, for example, a pharmaceutical composition comprising an anti-inflammatory polypeptide) to a subject suffering from cancer.
[0216] For any of the foregoing methods, the subject can be an animal, such as a domesticated animal (e.g., a horse, cow, pig, goat, sheep, rabbit, chicken, turkey, duck, etc.), a pet (e.g., a dog, cat, rabbit, hamster, gerbil, bird, fish, etc.), a lab animal (e.g., a mouse, rat, monkey, chimpanzee, owl, fish, etc.), a zoo animal (e.g., a gorilla, orangutan, chimpanzee, monkey, elephant, camel, zebra, boar, lion, tiger, giraffe, bear, bird, etc.), a wild animal (e.g., a deer, wolf, mountain lion, bird, etc.), or a human.
[0217] In conjunction with any of the foregoing methods, the anti-inflammatory polypeptide(s) can be administered at a dose and frequency that depends on the type of animal, the size of the animal, and the condition being treated. Typically, the anti-inflammatory polypeptide is administered daily (or every other day, or weekly), in an amount between about 1 mg and about 1000 mg (e.g., about 5 mg to about 900 mg, about 5 mg to about 800 mg, about 5 mg to about 700 mg, about 5 mg to about 600 mg, about 10 mg to about 500 mg, about 10 mg to about 400 mg, about 10 mg to about 300 mg, about 10 mg to about 250 mg, about 10 mg to about 200 mg, about 10 mg to about 150 mg, about 10 mg to about 100 mg, about 50 mg to about 500 mg, about 50 mg to about 400 mg, about 50 mg to about 300 mg, about 50 mg to about 250 mg, about 50 mg to about 200 mg, about 50 mg to about 150 mg, about 50 mg to about 100 mg, about 75 mg to about 500 mg, about 75 mg to about 400 mg, about 75 mg to about 300 mg, about 75 mg to about 250 mg, about 75 mg to about 200 mg, about 75 mg to about 150 mg, about 75 mg to about 100 mg, about 100 mg to about 500 mg, about 100 mg to about 400 mg, about 100 mg to about 300 mg, about 100 mg to about 250 mg, about 100 mg to about 200 mg, or any other range containing two of the foregoing endpoints). The daily dose can be administered once during the day, or broken up into smaller doses that are taken at multiple time points during the day. For a human (and other similarly-sized mammals), a dose of 5 mg/kg every other day can be administered. The anti-inflammatory polypeptide can be administered for a fixed period of time (e.g., for 2-3 weeks), at intervals (e.g., administer polypeptide for 2-3 weeks, wait 2-3 weeks, then repeat the cycle), or until such time as the pro-inflammatory cytokine levels have been reduced or stabilized, the chronic inflammatory condition or fibrosis has ameliorated, or the cancer has gone into remission.
[0218] The administration of the anti-inflammatory polypeptides (or pharmaceutical compositions comprising such polypeptides) in conjunction with any of the foregoing methods can be performed intravenously, intraperitoneally, parenteral, orthotopically, subcutaneously, topically, nasally, orally, sublingually, intraocularly, by means of an implantable depot, using nanoparticle-based delivery systems, microneedle patch, microspheres, beads, osmotic or mechanical pumps, and/or other mechanical means.
[0219] In conjunction with any of the foregoing methods, the anti-inflammatory polypeptides (or pharmaceutical compositions comprising such polypeptides) can be administered in combination with another drug designed to reduce or prevent inflammation, treat or prevent chronic inflammation or fibrosis, or treat cancer. In each case, the anti-inflammatory polypeptide can be administered prior to, at the same time as, or after the administration of the other drug. For the treatment of cancer, the anti-inflammatory polypeptide(s) can be administered in combination with a chemotherapeutic agent selected from the group consisting of steroids, anthracyclines, thyroid hormone replacement drugs, thymidylate-targeted drugs, Chimeric Antigen Receptor/T cell therapies, and other cell therapies. Specific chemotherapeutic agents include, for example, Gemcitabine, Docetaxel, Bleomycin, Erlotinib, Gefitinib, Lapatinib, Imatinib, Dasatinib, Nilotinib, Bosutinib, Crizotinib, Ceritinib, Trametinib, Bevacizumab, Sunitinib, Sorafenib, Trastuzumab, Ado-trastuzumab emtansine, Rituximab, Ipilimumab, Rapamycin, Temsirolimus, Everolimus, Methotrexate, Doxorubicin, Abraxane, Folfirinox, Cisplatin, Carboplatin, 5-fluorouracil, Teysumo, Paclitaxel, Prednisone, Levothyroxine, and Pemetrexed.
[0220] Alternatively, for the methods of treating cancer, the anti-inflammatory polypeptide(s) (or pharmaceutical compositions comprising such polypeptides) can be administered in combination with radiation therapy. Again, the anti-inflammatory polypeptide(s) can be administered prior to, or after the administration of the radiation therapy.
[0221] Any of the foregoing methods of the invention further include a step of assessing the efficacy of the therapeutic treatment. Because the anti-inflammatory polypeptides of the invention have a demonstrable ability to reduce tissue inflammation and suppress the excessive production of inflammatory mediators such as IL-1, IL-6, IL-12, and TNF.alpha., both in tissues and in serum (data not shown), the efficacy of the therapeutic treatment can be assessed by measuring the levels of such cytokines (e.g., in the serum) to determine whether the levels have responded appropriately to the treatment. Depending on the cytokine levels, the dosage of anti-inflammatory polypeptide(s) can be adjusted up or down, as needed.
EXAMPLES
Example 1: Peptide Designs
[0222] Polypeptides were designed in silico to include a striapathic region of alternating X.sub.m and Y.sub.n modules, with each X.sub.m module having one to five hydrophilic amino acid residues and each Y.sub.n module having one to five hydrophobic residues.
[0223] Initial designs focused on polypeptides consisting of a striapathic region having a total length of around 10 amino acid residues, with each X.sub.m module having one or two hydrophilic amino acid residues and each Y.sub.n module having one or two hydrophobic residues, and with the ratio of hydrophobic to hydrophilic amino acid residues being around 1:1. Such polypeptides were predicted to have an amphipathic, helical secondary structure, with a hydrophobic surface on one side of the helix and a hydrophilic surface on the opposite side of the helix.
[0224] Additional peptide designs were subsequently generated that maintained a total length of around 10 amino acid residues, but expanded the number of possible amino acid residues in a hydrophilic or hydrophobic module from two to three and varied the hydrophobic to hydrophilic ratio. For example, larger hydrophobic modules having three hydrophobic amino acid residues were coupled with shorter hydrophilic modules having one hydrophilic amino acid residue, giving rise to polypeptides predicted to have a stronger hydrophobic character. Such peptides were predicted to maintain an amphipathic, helical secondary structure, but have a larger hydrophobic surface on one side of the helix and a correspondingly smaller hydrophilic surface on the other side. Similarly, larger hydrophilic modules having three hydrophilic amino acid residues were coupled with shorter hydrophobic modules having one hydrophobic amino acid residue, giving rise to peptides having a stronger hydrophilic character. Such peptides were also predicted to maintain an amphipathic, helical secondary structure, but have a larger hydrophilic surface on one side of the helix and a correspondingly smaller hydrophobic surface on the other side.
[0225] Other peptide designs included: polypeptides having modules of four or five hydrophilic amino acid residues and/or four or five hydrophobic; polypeptides having a total length of around 10 amino acid residues but lacking hydrophobic amino acid residues; polypeptides having hydrophilic and hydrophobic modules each consisting of a single amino acid residue; and proline-rich polypeptides. Finally, larger polypeptides comprising two of the smaller peptide designs were also generated.
[0226] Exemplary polypeptides designed as described above are presented in Tables 3-9, below. To provide greater clarity into the types of polypeptides that have been developed, the peptides have been organized into Classes. Typically, the striapathic region of a specific Class of polypeptides shares a common sequence of hydrophobic and hydrophilic modules that is at least six or seven amino acid residues long. However, because the data indicates that polypeptides that have the same sequence but reversed N-terminal to C-terminal orientation have surprisingly similar anti-inflammatory activities, efforts have been made to keep such polypeptides in the same Class. Accordingly, some polypeptides have been grouped into the same Class even though the common sequence of hydrophobic and hydrophilic modules is less than six amino acid residues long. In addition, some of the polypeptides could have been grouped differently because they contain the common sequence of hydrophobic and hydrophilic modules of more than one Class. Thus, while providing a helpful framework for organizing the polypeptides around structural and functional similarities, the classification system does not capture all aspects of the relationships between different polypeptides.
[0227] Table 3 presents various Class I polypeptides, which have a striapathic region that includes a sequence corresponding to Formula I (i.e., Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c). Two different types of Class I polypeptides are presented in Table 3: peptides that have a striapathic region consisting of a sequence corresponding to Formula II (i.e., Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.2a-Y- .sub.3a-X.sub.3a); and peptide that have a striapathic region consisting of a sequence corresponding to Formula III (i.e., X.sub.2a-Y.sub.3a-X.sub.3a-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y- .sub.2b-Y.sub.2c). In addition, a peptide having a striapathic region having a sequence corresponding to Formula I, but not Formulas II or III, is presented.
TABLE-US-00024 TABLE 3 Class I Polypeptides RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 394 NFNFFFRFFF -1,286.6 III 33 108 WWWRWWWEWQ -1,278.0 II 34 109 EFNFFFRFFF -1,247.7 III 35 110 DFEFFFRFFF -1,232.0 III 36 111 QFEFFFRFFF -1,226.8 III 37 112 EFEFFFRFFF -1,216.0 III 38 113 FFFRFFFEFQ -1,208.9 II 39 114 FFFRFFFEFE -1,176.3 II 40 115 FFFRFFFEFD -1,172.3 II 41 116 FFFRFFFNFE -1,162.6 II 42 117 FFFRFFFDFE -1,147.7 II 43 118 FFFRFFFNFN -1,139.9 II 44 119 FFFHFFFEFQ -1,135.4 II 45 120 FFFHFFFNFE -1,126.4 II 46 121 FFFHFFFEFN -1,126.4 II 47 122 EFNFFFHFFF -1,125.1 III 48 123 FFFRFFFEFN -1,124.5 II 49 125 FFFHFFFEFE -1,115.4 II 50 126 QFEFFFHFFF -1,114.4 III 51 127 FFFHFFFEFD -1,114.3 II 52 128 FFFHFFFDFE -1,111.4 II 53 129 YYYRYYYEYQ -1,110.2 II 54 130 NFEFFFHFFF -1,109.1 III 55 131 FFFKFFFKFE -1,107.0 II 56 133 EFDFFFRFFF -1,103.4 III 57 135 FFFHFFFDFD -1,102.4 II 58 136 FFFHFFFNFN -1,100.4 II 59 137 FFFRFFFDFD -1,100.3 II 60 138 FFFKFFFKFN -1,098.2 II 61 139 FFFKFFFEFE -1,095.1 II 62 140 FFFEFFFKFE -1,091.8 II 63 141 FFFQFFFQFQ -1,088.8 II 64 143 FFFKFFFQFQ -1,084.4 II 65 144 FFFKFFFNFN -1,083.5 II 66 145 FFFNFFFNFN -1,083.3 II 67 146 FFFKFFFEFQ -1,082.6 II 68 148 FFFKFFFKFQ -1,080.0 II 69 149 FFFKFFFQFK -1,079.6 II 70 150 FFFKFFFKFD -1,077.4 II 71 152 FFFKFFFDFD -1,074.5 II 72 153 FFFNFFFKFN -1,074.2 II 73 154 FFFDFFFDFD -1,073.5 II 74 155 FFFKFFFEFK -1,073.3 II 75 156 FFFKFFFDFK -1,072.6 II 76 157 FFFEFFFEFE -1,070.8 II 77 158 FFFDFFFKFD -1,070.7 II 78 159 FFFKFFFKFK -1,070.7 II 79 160 FFFEFFFKFK -1,069.7 II 80 161 FFFQFFFKFK -1,069.6 II 81 162 FFFKFFFNFK -1,069.2 II 82 163 FFFNFFFKFK -1,066.7 II 83 164 FFFQFFFKFQ -1,062.5 II 84 165 FFFDFFFKFK -1,061.9 II 85 179 LLLRLLLELQ -966.7 II 86 395 FVFKFVFKFV -917.2 I 87 211 CCCRCCCECQ -818.2 II 88 230 MMMRMMMEMQ -774.6 II 89 232 VVVRVVVEVQ -771.6 II 90 258 IIIRIIIEIQ -699.2 II 91 267 GGGRGGGEGQ -640.4 II 92 268 PPPRPPPEPQ -627.1 II 93 271 TTTRTTTETQ -614.4 II 94 273 AAARAAAEAQ -609.4 II 95 280 AAAKAAAKAA -556.0 II 96 281 AAAEAAAEAE -541.6 II 97 287 SSSRSSSESQ -499.3 II 98
[0228] Table 4 presents some quasi-Class I polypeptides. These peptides include a sequence similar to the striapathic sequence of Formula II (i.e., Y.sub.1a-Y.sub.1b-Y.sub.1e-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.s- ub.2a-Y.sub.3a-X.sub.3a), but the hydrophobic amino acid residues have all been replaced with a particular hydrophilic amino acid residue.
TABLE-US-00025 TABLE 4 Quasi-Class I Polypeptides RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 173 HHHRHHHEHQ -1,002.2 II* 99 195 RRRRRRRERQ -855.2 II* 100 275 QQQRQQQEQQ -575.6 II* 101 276 EEEREEEEEQ -569.5 II* 102 284 NNNRNNNENQ -522.7 II* 103 288 DDDRDDDEDQ -463.6 II* 104 290 KKKRKKKEKQ -423.7 II* 105 *These peptides o not comply with the sequence requirements of Formula II, but instead represent an ''all hydrophilic'' variation on the sequence requirements of Formula II.
[0229] Table 5 presents various Class II, Sub-class 1 polypeptides. The presented peptides have a striapathic region consisting of a sequence corresponding to Formula X (i.e., Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y- .sub.3a-X.sub.3a), or a striapathic region consisting of a sequence corresponding to Formula XI (i.e., X.sub.1a-Y.sub.1a-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a-X.sub.3b-Y- .sub.3a-Y.sub.3b).
TABLE-US-00026 TABLE 5 Class II, Sub-class 1 Polypeptides RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 124 FFQKFFKRWR -1,121.3 X 106 132 FFRKFFKRFR -1,104.8 X 107 134 RFRKFFKRFF -1,103.3 XI 108 142 RFRKFFKQFF -1,085.5 XI 109 147 FFQKFFKRFR -1,080.3 X 110 151 RWRKFFKQFF -1,077.0 XI 111 166 FFEHFWKEFN -1,044.8 X 112 167 FFQHFWKQFN -1,024.9 X 113 168 QFNHFFKEFF -1,022.8 XI 114 169 FFDKFFHDFQ -1,014.2 X 115 170 QFDHFFKDFF -1,011.9 XI 116 171 FFEKFFHNFQ -1,009.9 X 117 172 NFEKWFHEFF -1,007.9 XI 118 175 LFRRAFKQLD -989.5 X 119 177 NFQKWFHQFF -976.3 XI 120 182 KFRKAFKRFF -944.8 XI 121 183 FFRKFAKRFK -933.2 X 122 185 FFKKFFKKFK -920.6 X 123 186 KFKKFFKKFF -919.6 XI 124 424 KARKAFKRFF -910.2 XI 125 190 WVKDAMQHLD -888.7 X 126 194 FFKKFAKKFK -859.1 X 127 198 FAEKFFKNFK -850.4 X 128 199 KFNKFFKEAF -847.1 XI 129 200 FAKQFFNKFK -846.0 X 130 201 KFNKAFKQAF -837.8 XI 131 202 KFNKAFKQAF -837.8 XI 131 204 FAQKFFKDFK -835.9 X 133 206 FAEEFAEEFE -823.1 X 134 207 KFKKFFKKAF -820.7 XI 135 209 KFKNFFQKAF -819.1 XI 136 210 KFKNFFQKAF -819.1 XI 136 212 FAKQFANKFK -817.9 X 138 213 KFKNAFQKAF -815.2 XI 139 214 KFKNAFQKAF -815.2 XI 139 215 FAKKFFKKFK -814.0 X 141 216 KFKKAFKKFF -811.2 XI 142 218 FAEKFAEKFE -807.6 X 143 219 DLHQMADKVW -807.6 XI 144 425 KARKAAKRFF -800.3 XI 145 225 FAKNFAKKFK -794.0 X 146 227 FAEKFAKNFK -786.6 X 147 233 KFKKAFKKAF -771.2 XI 148 234 FAKNFAKNFK -769.8 X 149 235 FAKEFAKEFE -768.9 X 150 236 KFDKAFKQAF -766.2 XI 151 237 KFDKAFKQAF -766.2 XI 151 238 FAEKFAKKFK -765.1 X 153 239 FAEKFAEKFK -764.2 X 154 398 FAKKFAKKFK -760.3 X 155 241 FAKNFAKNFN -758.7 X 156 242 FAQKFAKNFK -758.6 X 157 243 FANNFANNFN -755.2 X 158 244 FANNFANNFN -755.2 X 158 245 FANKFANKFN -754.0 X 160 246 FANKFAKKFK -752.2 X 161 247 FAQKFAKDFK -750.7 X 162 250 FAKEFAKEFK -745.7 X 163 251 FANKFANKFK -739.7 X 164 252 KFDKFFKQAF -739.1 XI 165 253 KFDKFFKQAF -739.1 XI 165 254 KFNKAFKEAF -738.4 XI 167 255 KFNKAFKEAF -738.4 XI 167 256 FAKEFAKKFK -702.8 X 169 426 KARKAAKRAF -634.5 XI 170 427 KARKAAKRAA -578.1 XI 171 285 AAEEAAEEAE -511.6 X 172 387 AAKKAAKKAK -301.6 X 173
[0230] Table 6 presents polypeptides that fall into a variety of different Classes, including: Class II peptides (having a striapathic region that includes a sequence corresponding to any of Formulas VI to XVI); Class II, Sub-class 2 (having a striapathic region that includes a sequence corresponding to Formulas VIII and XII); Class II, Sub-class 3 (having a striapathic region that includes a sequence corresponding to Formula IX); Class II, Sub-class 4 (having a striapathic region that includes a sequence corresponding to Formulas XIV and XV); Class II, Sub-class 5 (having a striapathic region that includes a sequence corresponding to Formulas XIII and XVI); Class III peptides (having a striapathic region that includes a sequence corresponding to any of Formulas XVII to XX); Class III, Sub-class 1 peptides (having a striapathic region that includes a sequence corresponding to Formulas XIX or XX); Class IV peptides (having a striapathic region that includes a sequence corresponding to Formulas IV and V); Class V peptides (having a striapathic region that includes a sequence corresponding to Formula XXI); Class VI peptides (having a striapathic region that includes a sequence corresponding to Formulas XXII and XXIII); Class VII peptides (having a striapathic region that includes a sequence corresponding to any of Formulas XXIV to XXVI); Class VIII peptides (having a striapathic region that includes a sequence corresponding to any of Formulas XXVII to XXXII); Class VIII, Sub-class 3 and 4 peptides (having a striapathic region that includes a sequence corresponding to Formulas XXXI and XXXII, respectively); Class IX peptides (having a striapathic region that includes a sequence corresponding to any of Formulas XXXIII to XXXVIII); Class IX, Sub-class 3 and 4 peptides (having striapathic regions that include a sequence corresponding to Formulas XXXVII and XXXVIII, respectively); and Class XIII (having a striapathic region that includes a sequence corresponding to Formula L). Because polypeptides of Class VIII, Sub-class 3 and Class IX, Sub-class 3 share the same sequence of hydrophobic and hydrophilic modules, but reversed N-terminal to C-terminal orientation, they could have been grouped into the same Class and Sub-class. Similarly, because polypeptides of Class VIII, Sub-class 4 and Class IX, Sub-class 4 share the same sequence of hydrophobic and hydrophilic modules, but reversed N-terminal to C-terminal orientation, they could have been grouped into the same Class and Sub-class.
TABLE-US-00027 TABLE 6 Class II to Class IX and Class XIII Polypeptides RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 396 FVKFVKFVKF -1,039.7 L 174 405 KRKAFRKFFF -1,026.6 XIV 175 174 LHKMYNQVW -1,000.2 VII 176 176 WVQNYMKHL -979.3 VII 177 178 RLVEMMRQIW -972.2 XX 178 180 FLKRLLQEI -955.9 VII 179 181 LRLLHRLL -950.2 XVII 180 184 WVRDSMKHL -925.6 VII 181 408 KFFRKKFRFA -917.4 XXII 182 187 WVQRVVEKFL -906.4 IX 183 416 AFFRRFKFKK -904.1 XXV 184 188 LFKEVVRQVW -902.9 IX 185 189 MDKIYDQVWK -893.3 VIII 186 388 FVKKFVKKFV -891.9 X 187 417 KKFKFRRFFA -888.8 XXVI 188 191 WVRDVVRSMD -874.1 XIX 189 192 ELSNIYERVW -872.4 XX 190 193 WIQRMMEVLR -866.9 XIX 191 404 FFFKRFAKRK -856.7 XV 192 196 LHKMSDRVW -852.4 VII 193 197 WVREYINSLE -851.2 XIX 195 402 FFKKRFAFRK -851.0 XXXI 196 203 KWVQDYIKDM -837.0 XII 197 409 AFRFKKRFFK -832.7 XXIII 198 205 LLRHLLRL -830.0 XVII 199 208 WIKKLLESSQ -819.7 XIX 200 217 DMSRVVDRVW -810.4 XX 201 220 FEEEFEEEFE -804.8 V 202 221 WVKNSINHL -803.7 VII 203 222 LTKKGRRFC -799.7 XXI 204 223 IEQLLRKLF -796.8 VII 205 224 LHNISNKVW -794.5 VII 206 226 CFRRGKKTL -786.7 XXI 207 229 IVRRADRAAV -781.5 XXI 208 231 TVERFKNLS -771.8 XXI 209 240 QSSELLKKIW -761.9 XX 210 248 SLNKFREVT -750.5 XXI 211 249 LIKQIVKKLF -750.5 IX 212 397 FAKKFAKKF -739.3 VII 194 415 KKKFFF -706.8 XXVII 213 257 LYKKIIKKLL -699.8 IX 214 259 FKKKFKKKFK -686.5 V 215 260 VAARDARRVI -684.6 XXI 216 261 FLKKVIQKIL -679.4 IX 217 262 LIKEIIKQVM -668.4 IX 218 263 LLKKIIKKYL -666.7 IX 219 264 AFFEEEAEFE -652.2 XXXVIII 220 265 KKWVQDSMK -650.1 XVIII 221 266 NFANKVQEVA -644.1 XXI 222 269 AVEQVKNAFN -621.1 XXI 223 272 MVQKIIEKIL -613.1 IX 224 274 KMSDQVWKK -595.9 XVIII 225 277 MVKKIIEKM -569.2 VII 226 278 ALKKQVIKKI -559.1 XVI 227 279 IKKIVQKKLA -556.7 XIII 228 282 AFFKKKAKFK -537.6 XXXVIII 229 283 MKEIIKVM -533.1 VII 230 286 AEEEAEEEAE -504.4 V 231 289 AKKKAKKKAK -431.6 V 232 414 KKKAAA 0.0 XXVII 233
[0231] Table 7 presents polypeptide of Classes VIII through XI. All of the peptides presented in Table 7 have a striapathic region that includes a hydrophilic module having four or five hydrophilic amino acid residues and/or a hydrophobic module having four or five hydrophobic amino acid residues. Class VIII, Sub-class 1 peptides have a striapathic region that includes a sequence corresponding to Formulas XXVIII or XXIX; Class VIII, Sub-class 2 peptides have a striapathic region that includes a sequence corresponding to Formula XXX; Class IX, Sub-class 1 peptides have a striapathic region that includes a sequence corresponding to Formulas XXXIV or XXXV; Class IX, Sub-class 2 peptides have a striapathic region that includes a sequence corresponding to Formula XXXVI; Class X peptides have a striapathic region that includes a sequence corresponding to any of Formulas XXXIX to XLIII; and Class XI peptides have a striapathic region that includes a sequence corresponding to any of Formulas XLIV to XLVIII. Because polypeptides of Class VIII, Sub-class 1 and Class IX, Sub-class 1 share the same sequence of hydrophobic and hydrophilic modules, but reversed N-terminal to C-terminal orientation, they could have been grouped into the same Class and Sub-class. Similarly, because polypeptides of Class VIII, Sub-class 2 and Class IX, Sub-class 2 share the same sequence of hydrophobic and hydrophilic modules, but reversed N-terminal to C-terminal orientation, they could have been grouped into the same Class and Sub-class.
TABLE-US-00028 TABLE 7 Class VIII to XI Polypeptides RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 406 KRKKRFAFFF -993.5 XXX 234 422 RKRKFFAFFK -948.2 XLVIII 235 407 FFFAFRKKRK -914.7 XXXVI 236 400 FRKKRFAFFK -900.5 XXIX 237 419 FFFRRKKKFA -881.9 XLII 238 401 KFFAFRKKRF -880.1 XXXV 239 423 KFFAFFKRKR -877.1 XLV 240 411 KKKKKFFFFF -863.7 XXX 241 418 AFKKKRRFFF -854.1 XLI 242 428 KRKKRAAFFF -842.0 XXX 243 420 KKFFAFFRKR -840.2 XLVI 244 421 RKRFFAFFKK -835.5 XLVII 245 429 KRKKRAAAFF -758.1 XXX 246 413 KKKKFFFF -715.8 XXVIII 247 430 KRKKRAAAAF -676.7 XXX 248 270 KKKAFFFAKK -614.4 XLVII 249 431 KRKKRAAAAA -544.9 XXX 250 410 KKKKKAAAAA -385.3 XXX 251 412 KKKKAAAA -382.8 XXVIII 252
[0232] Table 8 presents polypeptides of Class XII and Class XIV. Class XII peptides have astriapathic region that includes a sequence corresponding to Formula XLIX (i.e., Y.sub.1a-X.sub.1a-Y.sub.2a-X.sub.2a-Y.sub.3a-X.sub.3a). Class XII peptides are predicted to adopt a beta-strand secondary structure. Class XIV peptides are proline-rich peptides that have astriapathic region that includes a sequence corresponding to one of Formulas LI-LIV.
TABLE-US-00029 TABLE 8 Beta-Strand and Proline-Rich Polypeptides RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 393 FKFKFKFKF -1,193.2 XLIX 253 391 FRFKFKFR -1,190.8 XLIX 254 392 RFQFKFRF -1,170.3 XLIX 255 390 FRFKFKF -1,083.3 XLIX 256 389 FRFKFA -1,009.8 XLIX 257 449 RRFPRPPFF -1,116.8 LI 258 450 FFPPRPFRR -1,100.0 LII 259 448 LYPPRPFRR -1,059.3 LII 260 447 RRIPRPPYL -1.050.5 LI 261 452 PFRPPPRPRF -1,012.2 LIII 262 451 PRPRPPPRFF -1,002.1 LIV 263 444 FFPPKPFKK -954.8 LII 264 441 KKIPKPPYL -922.1 LI 265 446 PFKPPPKPKP -882.3 LIII 266 445 PKPKPPPKFP -866.3 LIV 267 442 LYPPKPIKK -846.6 LII 268 443 KKFPKPPFF -802.8 LI 269
[0233] Table 9 presents fusion peptides, which include combinations of Class I, Class II, and/or Class III peptides linked together by a peptide bond and, optionally, a short peptide linker (e.g., a tri-glycine (GGG) linker).
TABLE-US-00030 TABLE 9 Peptide Combinations RelB RP Binding E SEQ ID # Sequence (kCal/mol) Formula NO: 292 EFEFFFRFFFGGGEFEFFFRFFF -1,606.1 III + III 270 293 QFEFFFRFFFGGGQFEFFFRFFF -1,602.0 III + III 271 294 DFEFFFRFFFGGGDFEFFFRFFF -1,591.8 III + III 272 295 EFNFFFRFFFGGGEFNFFFRFFF -1,591.8 III + III 273 296 FFFRFFFEFQFFFRFFFEFQ -1,511.6 II + II 274 297 FFFRFFFEFQGGGFFFRFFFEFQ -1,511.5 II + II 275 298 RWRKFFKRFFQFEFFFRFFF -1,505.2 XI + III 276 299 RWRKFFKRFFGGGFFFRFFFNFN -1,501.3 XI + II 277 300 RFRKFFKRFFQFEFFFRFFF -1,486.0 XI + III 278 301 RFRKFFKRFFGGGFFFRFFFNFN -1,485.0 XI + II 279 302 RWRKFFKRFFGGGFFFRFFFEFQ -1,479.6 XI + II 280 303 RFRKFFKRFFGGGFFFRFFFEFQ -1,476.8 XI + II 281 304 EFEFFFRFFFEFEFFFRFFF -1,476.0 III + III 282 305 RWRKFFKRFFNFNFFFRFFF -1,474.2 XI + III 283 306 QFEFFFRFFFQFEFFFRFFF -1,467.0 III + III 284 307 RWRKFFKRFFGGGNFNFFFRFFF -1,464.2 XI + III 285 308 EFNFFFRFFFEFNFFFRFFF -1,460.5 III + III 286 309 RFRKFFKRFFNFNFFFRFFF -1,458.4 XI + III 287 310 FFRKFFKRFRGGGNFNFFFRFFF -1,447.1 X + III 288 311 RFRKFFKRFFGGGNFNFFFRFFF -1,432.1 XI + III 289 312 DFEFFFRFFFDFEFFFRFFF -1,430.0 III + III 290 313 RWRKFFKRFFFFFRFFFEFQ -1,427.4 XI + II 291 314 RFRKFFKRFFFFFRFFFEFQ -1,425.6 XI + II 292 315 FFRKFFKRFRGGGFFFRFFFNFN -1,420.6 X + II 293 316 FFRKFFKRWRGGGFFFRFFFNFN -1,417.5 X + II 294 317 RFRKFFKRFFFFFRFFFNFN -1,406.6 XI + II 295 318 FFRKFFKRFRFFFRFFFEFQR -1,402.0 X + II 296 291 FFEHFWKEFNGGGNFQKWFHQFF -1,401.6 X + XI 297 319 FFRKFFKRWRQFEFFFRFFF -1,400.7 X + III 298 320 RWRKFFKRFFFFFRFFFNFN -1,397.9 XI + II 299 321 NFQKWFHQFFGGGFFEHFWKEFN -1,396.0 XI + X 300 322 FFRKFFKRWRGGGNFNFFFRFFF -1,394.4 X + III 301 323 FFRKFFKRWRFFFRFFFEFQR -1,394.3 X + II 302 324 FFRKFFKRWRNFNFFFRFFF -1,393.7 X + III 303 325 FFRKFFKRFRGGGFFFRFFFEFQR -1,386.8 X + II 304 326 FFRKFFKRFRQFEFFFRFFF -1,382.8 X + III 305 327 FFRKFFKRFRNFNFFFRFFF -1,378.2 X + III 306 328 RFRKFFKRFFGGGQFEFFFRFFF -1,368.5 XI + III 307 329 FFRKFFKRWRGGGFFFRFFFEFQR -1,354.5 X + II 308 330 FFRKFFKRFRGGGQFEFFFRFFF -1,352.8 X + III 309 331 FFRKFFKRWRGGGQFEFFFRFFF -1,352.2 X + III 310 332 RWRKFFKRFFGGGQFEFFFRFFF -1,349.8 XI + III 311 333 QFNHFFKEFGGGQFNHFFKEFF -1,340.0 VII + XI 312 334 FFRKFFKRFRFFFRFFFNFN -1,337.5 X + II 313 335 FFRKFFKRWRFFFRFFFNFN -1,337.0 X + II 314 336 FFEHFWKEFNGGGFFEHFWKEFN -1,325.5 X + X 315 337 FFEHFWKEFGGGNFQKWFHQFF -1,324.8 VII + XI 316 338 NFQKWFHQFGGGFFEHFWKEFN -1,317.9 VII + X 317 339 FFEHFWKEFNGGGLHKMYNQVW -1,315.4 X + VII 318 340 NFQKWFHQFFGGGNFQKWFHQFF -1,309.9 XI + XI 319 341 FAKKFAKKFKGGGNFQKWFHQFF -1,308.3 X + XI 320 342 FFEKFFHNFQGGGFFEKFFHNFQ -1,304.6 X + X 321 343 FFQHFWKQFNGGGFFQHFWKQFN -1,300.2 X + X 322 344 NFQKWFHQFFNFQKWFHQFF -1,293.5 XI + XI 323 345 FAKKFAQKFKGGGNFQKWFHQFF -1,291.9 X + XI 324 346 FAKKFAKKFKGGGQFEFFFRFFF -1,290.9 X + III 325 347 QFNHFFKEFQFNHFFKEFF -1,279.8 VII + XI 326 348 FAKKFAKKFKGGGDFEFFFRFFF -1,278.4 X + III 327 349 FFEHFWKEFNGGGWVQNYMKHL -1,268.8 X + VII 328 350 FAKKFAKKFKQFEFFFRFFF -1,268.5 X + III 329 351 FFQHFWKQFNFFQHFWKQFN -1,263.2 X + X 330 352 FFEHFWKEFNFFEHFWKEFN -1,251.5 X + X 331 353 NFEKWFHEFFNFEKWFHEFF -1,247.0 XI + XI 332 354 FAKKFAKKFKGGGQFNHFFKEFF -1,244.6 X + XI 333 355 NFEKWFHEFFGGGNFEKWFHEFF -1,241.4 XI + XI 334 356 FAKKFAKKFKGGGFFFRFFFEFQ -1,237.9 X + II 335 357 FAKKFAKKFKDFEFFFRFFF -1,235.3 X + III 336 358 QFNHFFKEFFGGGQFNHFFKEFF -1,230.0 XI + XI 337 359 FAKKFAKKFKGGGEFEFFFRFFF -1,221.7 X + III 338 360 FAKKFAKKFKGGGEFNFFFRFFF -1,221.0 X + III 339 361 FAKKFAKKFKGGGNFEKWFHEFF -1,212.3 X + XI 340 362 FAKKFAKKFKGGGFFEKFFHNFQ -1,210.8 X + X 341 363 QFNHFFKEFFQFNHFFKEFF -1,208.6 XI + XI 342 364 FFEKFFHNFQFFEKFFHNFQ -1,207.5 X + X 343 365 FAKKFAKKFKEFEFFFRFFF -1,204.2 X + III 344 366 FAKKFAKKFKEFNFFFRFFF -1,187.6 X + III 345 367 FAKKFAKKFKFFEHFWKEFN -1,168.1 X + X 346 368 FAKKFAKKFKFFFRFFFEFQ -1,166.4 X + II 347 369 FAKKFAKKFKLHKMYNQVW -1,159.5 X + VII 348 370 FAKKFAKKFKGGGFFEHFWKEFN -1,140.4 X + X 349 371 FAKKFAKKFKGGGWVQNYMKHL -1,130.4 X + VII 350 372 FAKKFAKKFKNFQKWFHQFF -1,126.1 X + XI 351 373 FAKKFAKKFKFFQHFWKQFN -1,119.8 X + X 352 374 FAKKFAKKFKGGGFFQHFWKQFN -1,119.6 X + X 353 375 FAKKFAKKFKWVQNYMKHL -1,119.2 X + VII 354 376 FAKKFAKKFKQFNHFFKEFF -1,108.3 X + XI 355 377 FAKKFAKKFKGGGLHKMYNQVW -1,100.3 X + VII 356 378 FAKKFAKKFKNFEKWFHEFF -1,081.4 X + XI 357 379 FAKKFAKKFKFFEKFFHNFQ -1,046.8 X + X 358 380 FAKKFAKKFKGGGAFFKKKAKFK -950.9 X + 359 XXXVIII 381 AFFKKKAKFKGGGAFFKKKAKFK -935.5 XXXVIII + 360 XXXVIII 382 KFKKAFKKAFKFKKAFKKAF -925.2 XI + XI 361 383 KFKKAFKKAFGGGKFKKAFKKAF -923.8 XI + XI 362 384 FAKKFAKKFKGGGFAKKFAKKFK -909.2 X + X 363 385 FAKKFAKKFKAFFKKKAKFK -839.9 X + 364 XXXVIII 228 PSRKSMEKSVAKLLNKIAKSEP -782.4 IX + 365 XVIII 386 AFFKKKAKFKAFFKKKAKFK -716.0 XXXVIII + 366 XXXVIII
[0234] In each of Tables 3-9, the RP# is a randomly assigned designation used to identify specific peptide sequences. The "Binding E" (see column 3 in each of the Tables) corresponds to a predicted measure of the energy released when individual peptides bind to the protein dimerization domain of RelB, an NFkB Class II protein (see Example 2, below).
Example 2: Predicted Binding of Peptides to Rel B
[0235] To identify peptides having anti-inflammatory activity, the NF-kB complex was selected as a target, since it is known to be a key component in the signaling pathways that regulate inflammation. Dimerization of NF-kB (either homologous or heterologous), which is mediated by the dimerization domains found in NF-kB Class II proteins (e.g., RelA, RelB, cRel, NF-kB1, and NF-kB2), is essential for activation of the NF-kB complex and its generation of pro-inflammatory signals. Accordingly, peptide designs were selected for their ability to specifically bind to the dimerization domain of RelB (NCBI Acc. No. NP_033072.2), with the goal that such binding would inhibit NF-kB dimerization and activation.
[0236] Peptide binding to the dimerization domain of Rel B was evaluated in silico, using the web-based ClusPro.TM. algorithm developed at Boston University. The ClusPro.TM. algorithm filters docked conformations between a protein target and a putative ligand and determines surface complementarity, ranking the conformations based on their clustering properties. The free energy filters select complexes with the lowest desolvation and electrostatic energies. Clustering is then used to smooth the local minima and to select the ones with the broadest energy wells, a property associated with the free energy at the binding site. Using this method, it is possible to evaluate the affinity a ligand possesses for a particular target, whereupon the ligands can be ranked and then tested for biological activity in vitro or in vivo.
[0237] The binding energies calculated by the ClusPro.TM. algorithm are shown for each of the peptides in Tables 3-9, in the third columns of the tables. In each of Tables 3-9, the peptides are ranked according to the calculated RelB binding energy, from highest to lowest binding energy. The RelB binding energies were used to explore the structure-function relationship of the peptides, particularly with regard to (i) increasing or decreasing hydrophobicity, (ii) positive/negative charge density, and (iii) altering the arc of the hydrophobic and hydrophilic faces of the peptides. The peptides shown in Table 10 (below) will be used to illustrate the results of the study.
TABLE-US-00031 TABLE 10 Predicted Binding of Select Peptides to RelB RelB RP Binding E SEQ ID # Sequence (kCal/mol)* Formula NO: RP-182 KFRKAFKRFF -944.8 XI 121 RP-166 FFEHFWKEFN -1,044.8 X 112 RP-113 FFFRFFFEFQ -1,208.9 II 39 RP-289 AKKKAKKKAK -431.6 V 232 RP-387 AAKKAAKKAK -338.3 X 173 NF-CONTR2 IESKRRKKKP -476.6 N/A 382 NF-CONTR3 APGPGDGGTA -621.1 N/A 383 *The lower the energy value, the greater affinity the ligand possesses for the binding site on the target protein.
[0238] A structural model of the RelB subunit of NF-kB is shown in FIG. 1. Amino acids with the dimerization site are shaded dark gray to indicate their hydrophobic or hydrophilic character. In particular, the amino acid residues circled are hydrophilic, while the remaining dark gray amino acid residues are hydrophobic. Given the distinct locations of the hydrophilic and hydrophobic amino acid residues within the binding pocket of the dimerization domain, it is evident that striapathic peptides having an amphipathic secondary structure have the potential to bind site-specifically to the dimerization domain binding pocket.
[0239] The secondary structure of RP-182 (SEQ ID NO: 121) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 2. As can be seen in the panels on the right, RP-182's predicted secondary structure has distinct hydrophobic and hydrophilic sides that comprise approximately equal facial arcs (see also FIG. 9) and are of high volume. Overall, the structure of RP-182 possesses high hydrophobicity and high cationicity (with a total of five cationic amino acid residues). These characteristics of RP-182 are summarized in Table 11, below. Based on the structural modeling, RP-182 binds with high affinity to the RelB dimerization domain binding pocket, with an estimated binding energy of -944.8 kcal/mol.
[0240] The secondary structure of RP-166 (SEQ ID NO: 112) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 3. As can be seen in the panels on the right, RP-166's predicted secondary structure also has distinct hydrophobic and hydrophilic sides that comprise approximately equal facial arcs (see also FIG. 9). These characteristics are not surprising, as the striapathic region of RP-166 has a modular structure that is identical (albeit reversed) to that of RP-182's (compare Formulas X and XI). As with RP-182, the hydrophobic and hydrophilic surfaces of RP-166 are of high volume, but RP-166 has a greater ratio of hydrophobic volume to hydrophilic volume as compared to RP-182. In addition, the cationicity of RP-166 is significantly reduced relative to that of RP-182, since RP-166 has an equal number of cationic amino acid residues and anionic amino acid residues. These characteristics of RP-166 are summarized in Table 11, below. Based on the structural modeling, RP-166 binds to the RelB dimerization domain binding pocket with even higher affinity than RP-182, having an estimated binding energy of -1,044.8 kcal/mol.
[0241] The secondary structure of RP-113 (SEQ ID NO: 39) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 4. As can be seen in the panels on the right, RP-113's predicted secondary structure also has distinct hydrophobic and hydrophilic sides, but the hydrophobic side comprises a much larger facial arc than the hydrophilic side. As shown in FIG. 9, the facial arc of the polar side of RP-113 is only 60.degree., while the facial arc of the non-polar side is 300.degree.. Consistent with this shift toward a larger hydrophobic surface, RP-113 has a larger hydrophobic volume than either RP-182 or RP-166, as well as a significantly larger ratio of hydrophobic to hydrophilic volume. See Table 11, below. Like RP-166, the cationicity of RP-113 is significantly reduced relative to that of RP-182, since RP-113 has an equal number of cationic amino acid residues and anionic amino acid residues. Based on the structural modeling, RP-113 binds to the RelB dimerization domain binding pocket with one of the highest affinities predicted for the peptides of the invention, having an estimated binding energy of -1,208.9 kcal/mol.
[0242] The secondary structure of RP-387 (SEQ ID NO: 173) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 5. As can be seen in the panels on the right, RP-387's predicted secondary structure has distinct hydrophobic and hydrophilic sides. However, in contrast to RP-113, the hydrophilic side of RP-387 comprises a much larger facial arc than the hydrophobic side. As shown in FIG. 10, the facial arc of the polar side of RP-387 is 245.degree., while the facial arc of the non-polar side is 115.degree.. Consistent with this shift toward a larger hydrophilic surface, RP-387 has a smaller hydrophobic volume than any of RP-182, RP-166, and RP-113, as well as a significantly smaller ratio of hydrophobic to hydrophilic volume. See Table 11, below. With regard to cationicity, RP-387 is similar to RP-182, having a total of five cationic amino acid residues. Based on the structural modeling, RP-387 binds to the RelB dimerization domain binding pocket, but is does so relatively poorly, having an estimated binding energy of only -338.3 kcal/mol.
[0243] The secondary structure of RP-289 (SEQ ID NO: 232) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 6. As can be seen in the panels on the right, RP-289's predicted secondary structure has distinct hydrophobic and hydrophilic sides. However, RP-289's hydrophobic side is one of the smallest of the peptides screened. As shown in FIG. 9, the facial arc of the polar side of RP-289 is 290.degree., while the facial arc of the non-polar side is only 70.degree.. Of the peptides listed in Table 11, RP-289 has the smallest hydrophobic volume and the smallest ratio of hydrophobic to hydrophilic volume. RP-289 also has the highest cationicity of the peptides listed in Table 11, having a total of seven cationic amino acid residues. Based on the structural modeling, RP-289 binds to the RelB dimerization domain, though comparatively much more weakly than RP-182, RP-166, and RP-113, having an estimated binding energy of only -431.6 kcal/mol.
[0244] Tables 10 and 11 also identify two control peptides, NF-CONTR2 and NF-CONTR3, which are fragments of the RelB subunit of NF-kB. The sequences of NF-CONTR2 and NF-CONTR3 do not conform to any of structural Formulas I-LIII. The secondary structure of NF-CONTR2 (SEQ ID NO: 382) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 7. The secondary structure of NF-CONTR3 (SEQ ID NO: 383) and its binding to RelB (SEQ ID NO: 367) is modeled in FIG. 8. Neither peptide is predicted to adopt a clearly amphipathic secondary structure throughout the length of the peptide. Moreover, although the ClusPro.TM. algorithm identifies a binding interaction between each of NF-CONTR2 and NF-CONTR3 and RelB, the binding energies are not very strong and neither peptide displays a preference for binding to the RelB dimerization domain binding pocket.
TABLE-US-00032 TABLE 11 Physical Characteristics of Select Peptides HPB SEQ RelB Vol/ RP ID Binding HPL HPB HPL tH tH th HPB/ # Sequence NO: Energy (+) (-) Vol Vol Vol HPL HPB th HPL 182 KFRKAFKRFF 121 -944.8 6 1 696.9 659.8 0.95 -50.8 16.4 -0.32 166 FFEHFWKEFN 112 -1,044.8 3 3 637.7 775.0 1.22 -33.0 16.7 -0.51 113 FFFRFFFEFQ 39 -1,208.9 2 2 414.5 1030.4 2.49 -23.5 25.9 -1.10 289 AKKKAKKKAK 232 -431.6 8 1 896.8 213.3 0.24 -61.6 4.8 -0.08 387 AAKKAAKKAK 173 -338.3 6 1 640.5 355.5 0.55 -44.0 8.0 -0.18 NF-C2 IESKRRKKKP 382 -476.6 7 2 954.9 297.4 0.31 -66.8 3.5 -0.05 NF-C3 APGPGDGGTA 383 -621.1 1 1 115.1 665.7 5.78 -9.2 8.0 -0.87 *Binding energies are in kcal/mol. Volumes are in cubic angstroms. HPL means hydrophilic; HPB means hydrophobic. ''tH'' is the total hydrophobicity (in kcal/mol), as defined by Engleman et al. (1986), ''Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins,'' Annu. Rev.
Biophys. Bioeng. 15: 321-53.
[0245] FIGS. 1 through 10 and Table 11 reveal some important aspects of the structure-function relationship for the peptides of the invention. First, all of the peptides that are predicted to bind the RelB dimerization domain binding pocket have an amphipathic secondary structure. Second, greater hydrophobic volume, a greater ratio of hydrophobic to hydrophilic volume, and a greater hydrophobic arc are all associated with increased affinity for the binding pocket of the RelB dimerization domain. Third, increased cationicity is associated with decreased binding affinity for the binding pocket of the RelB dimerization domain.
[0246] Table 4, which lists some "all hydrophilic" variants of the Class I peptides, appears to potentially refute the conclusion that increased cationicity is associated with decreased binding affinity for the binding pocket of the RelB dimerization domain. In each of the peptides in Table 4, the hydrophobic residues of a Class I, Formula II peptide have been replaced with a single type of hydrophilic residue. Significantly, RP-173 (HHHRHHHEHQ; SEQ ID NO: 99) and RP-195 (RRRRRRRERQ; SEQ ID NO: 100) both have a high affinity for the binding pocket of the RelB dimerization domain (-1,002.2 and -855.2 kcal/mol, respectively), despite have eight amino acid residues that generally have a cationic charge in solution. Because both histidine and arginine have large side chains, a potential explanation for their high RelB binding affinities is that the uncharged hydrocarbon groups in the side chains provide some hydrophobicity that would otherwise have been lost by switching from a hydrophobic residue to a hydrophilic residue. In addition, when bound to RelB, some of the histidine and arginine residues may adopt an uncharged state. Table 4 therefore sheds further light on the structure-function relationship of the peptides of the invention by indicating that histidine and arginine can function in a quasi-hydrophobic capacity, at least with regard to the binding affinities of peptides for the RelB dimerization domain binding pocket. Accordingly, in some peptides of the invention, it can be energetically advantageous to place a histidine or arginine adjacent to a hydrophobic module that is made up of one or two hydrophobic amino acid residues.
Example 3: RelB Amino Acid Residues Involved in Peptide Binding
[0247] A model of the amino acid residues that line the binding pocket of the RelB dimerization domain is shown in FIG. 11. The model shows that Glu-298, Asp-330, and His-332 are key hydrophilic amino acid residues that line the binding pocket, while Tyr-300, Leu-301, Leu-302, and Leu-371 are important hydrophobic residues. The same model, with the addition of a stick diagram of the RP-182 peptide (SEQ ID NO: 121) is shown in FIG. 12. The dotted lines in FIG. 12 show ionic bonds between (1) Lys-7 of RP-183 and Asp-330 of RelB, and (2) Lys-4 of RP-183 and Glu-298 of RelB. Further stabilizing the interaction is an intra-ionic bond formed between Arg-8 of RP-183 and the carboxy terminal Lys-10 of RP-183. In addition to the ionic binds, there are numerous Van der Waals interactions. For the sake of clarity, only that of Phe-9 of RP-182 with Leu-371 of Rel-B is shown. However, the other hydrophobic amino acid residues on the hydrophobic face of RP-183 interact with the hydrophobic "floor" of the cleft of dimerization site of Rel-B.
[0248] An analysis of the ionic and Van der Waals interactions involved with the binding of different peptides of the invention has revealed that the peptides bind to a subset of the RelB amino acid residues selected from the group consisting of Leu-281, Ile-283, Cys-284, Glu-298, Tyr-300, Leu-301, Leu-302, Cys-303, Ile-311, Ser-312, Ala-329, Asp-330, Val-331, His-332, Gln-334, and Leu-371. See Table 13, below. Tyr-300, Leu-302, and His-332 are designated in the literature as being critical for dimerization. The amino acids most critical to binding by peptides of the invention include Glu-298, Tyr-300, Leu-302, Asp-330, Gln-334, and Leu-371.
Example 4: Binding of Peptides to Protein Targets Other Than RelB
[0249] A subset of the peptides shown in Tables 3-9 were further evaluated in silico to determine whether they bind to signaling proteins involved in the inflammatory response other than RelB. In doing so, it was discovered the dimerization cleft of the RelB subunit of NF-kB has structural parallels in a number of other signaling molecules. Consistent with these structural parallels, the peptides of the invention are predicted (by the ClusPro.TM. algorithm) to bind with high affinity to important signaling molecules in the inflammatory cascade, including: TGF.beta. (NCBI Acc. No. NP_000651.3; SEQ ID NO: 368); Notch1 (GenBank Acc. No. AAG33848.1; SEQ ID NO: 369); Wnt8R (NCBI Acc. No. XP_005214377.1; SEQ ID NO: 370); TRAIL (GenBank Acc. No. EAW78466.1; SEQ ID NO: 371); IL6R (NCBI Acc. No. NP_786943.1; SEQ ID NO: 372); IL10R (NCBI Acc. No. NP_001549.2; SEQ ID NO: 373); EGFR (GenBank Acc. No. AAR85273.1; SEQ ID NO: 374); and CDK6 (NCBI Acc. No. NP_001250.1; SEQ ID NO: 375). Representative peptides of the invention and the predicted binding energies between the peptides and each of these signaling molecules is shown in Tables 12A and 12B, below.
TABLE-US-00033 TABLE 12A Predicted Binding of Select Peptides to Different Inflammatory Targets RP SEQ ID # Sequence NO: RelB TGF.beta. NOTCH1 WNT8R TRAIL 185 FFKKFFKKFK 123 -920.6 -880.1 -817.7 -747.2 -904.5 186 KFKKFFKKFF 124 -919.6 -846.0 -887.7 -739.1 -884.3 183 FFRKFAKRFK 122 -933.2 -878.9 -890.8 -763.1 -938.8 182 KFRKAFKRFF 121 -944.8 -851.8 -1,096.3 -733.7 -938.8 118 FFFRFFFNFN 44 -1,139.9 -1,074.7 -1,032.4 -990.9 -995.4 394 NFNFFFRFFF 33 -1,286.6 -1,002.6 -1,059.6 -971.2 -943.8 389 FRFKFA 257 -1,009.8 -878.4 -846.4 -804.5 -916.8 390 FRFKFKF 256 -1,083.3 -933.2 -1,005.3 -871.0 -1,014.4 391 FRFKFKFR 254 -1,190.8 -987.5 -1,005.4 -897.9 -1,049.2 392 RFQFKFRF 255 -1,170.3 -943.2 -923.1 -853.8 -1,039.6 387 AAKKAAKKAK 173 -301.6 -397.7 -385.5 -394.9 -397.7 *All binding affinities are in kcal/mol.
TABLE-US-00034 TABLE 12A Predicted Binding of Select Peptides to Different Inflammatory Targets RP SEQ ID # Sequence NO: RelB EGFR IL6R IL10R CDK6 185 FFKKFFKKFK 123 -920.6 -785.4 -747.5 -756.3 -753.9 186 KFKKFFKKFF 124 -919.6 -866.3 -755.0 -742.0 -718.1 183 FFRKFAKRFK 122 -933.2 -795.6 -696.7 -738.6 -783.0 182 KFRKAFKRFF 121 -944.8 -853.8 -784.5 -785.9 -781.5 118 FFFRFFFNFN 44 -1,139.9 -1,039.4 -1,074.8 -881.4 -1,020.8 394 NFNFFFRFFF 33 -1,286.6 -1,061.4 -1,069.9 -850.8 -1,075.3 389 FRFKFA 257 -1,009.8 -896.0 -812.3 -779.2 -900.5 390 FRFKFKF 256 -1,083.3 -1,036.3 -952.2 -876.2 -861.1 391 FRFKFKFR 254 -1,190.8 -1,024.9 -957.6 -882.3 -899.9 392 RFQFKFRF 255 -1,170.3 -1,010.4 -1,052.3 -901.7 -870.0 387 AAKKAAKKAK 173 -301.6 -395.9 -342.0 -338.1 -351.4
[0250] The data reveals that the strength of binding to RelB is highly correlated with the strength of binding to the various inflammatory targets. In other words, peptides that are predicted to bind with high affinity to RelB are likewise predicted to bind with high affinity to TGF.beta., Notch1, Wnt8R, TRAIL, EGFR, IL6R, and IL10R.
[0251] A closer evaluation of the predicted binding interactions between the peptides of the invention and each of TGF.beta., Notch1, Wnt8R, TRAIL, EGFR, IL6R, and IL10R reveals that the peptides not only bind with high affinity, but also bind to functionally critical sites on the targets. For example, peptides of the invention are predicted to bind to the receptor-binding site on TGF.beta., the calcium-binding site on Notch1, the Wnt8-binding site on Wnt8R, the receptor-binding site on TRAIL, the IL6-binding site on IL6R, the IL10-binding site on IL10R, and the general ligand-binding site on EGFR. A non-exhaustive list of amino acid residues in each of these targets that are bound by the peptides of the invention is shown in Table 13.
TABLE-US-00035 TABLE 13 Amino Acid Residues in Target Proteins Contacted by Peptides of the Invention SEQ ID Most Target NO: AA Residue Contacts Critical AAs RelB 367 Leu-281, Ile-283, Cys-284, Glu-298, Glu-298, Tyr-300, Tyr-300, Leu- Leu-301, Leu-302, Cys-303, 302, Asp-330, Ile-311, Ser-312, Gln-334, Ala-329, Asp-330, Val-331, Leu-371 His-332, Gln-334, Leu-371 TGF.beta. 368 Leu-20, Ile-22, Phe-24, Asp-27, Leu-28, Asp-27, Leu-28, Trp-30, Trp-30, Trp-32, Tyr-39, Phe-43, Trp-32 Pro-80, Leu-83, Leu-101, Ser-112 Notch1 369 Phe-1520, Gln-1523, Phe-1520, Arg-1524, Glu-1526, Ala- Trp-1557, 1553, Glu-1556, Trp-1557, Cys-1562, Cys-1562, His-1602, Phe-1703 Arg-1684, Gln-1685, Cys-1686, Ser-1691, Cys- 1693, Phe-1694, Phe-1703 Wnt8R 370 Tyr-52, Gln-56, Phe-57, Tyr-52, Phe-57, Asn-58, Met-91, Tyr- Tyr-100, 100, Lys-101, Pro-103, Asp-145 Pro-105, Pro-106, Arg- 137, Asp-145 TRAIL 371 Ala-123, His-161, Ser-162, Phe-163, Phe-163, Tyr-183, Tyr-243, Glu- Tyr-185, Tyr-243, His-270, 271, Phe-278 Glu-271, Phe-274, Phe-278, Leu-279, Val-280 IL6R 372 Leu-108, Glu-140, Pro-162, Glu-140, Phe-229, Tyr-230, Phe-229, Tyr- Phe-279 230, Phe-279 IL10R 373 Leu-41, Arg-42, Tyr-43, Tyr-43, Ile-45, Glu-46, Ser-47, Ile-45, Glu-46, Trp-48, Arg-76, Arg-78 Trp-48 EGFR 374 Leu-10, Thr-40, Trp-41, Trp-41, Asp-48, Asp-48, Phe-51, Leu-63, Phe-51, His-66, Asp-68, Leu-88, Asp-68, Tyr-101, Tyr-101 CDK6 375 Val-142, Arg-144, Asp-145, Asp-145, Ser-171, Val-180, Val-180, Tyr- Val-181, Leu-183, Arg-186, 196 Val-190, Gln-193, Tyr-196, Val-200 HMT 376 Tyr-16, Glu-48, Tyr-50, Tyr-16, Glu-48, Leu-51, Phe-52, Asn-69 Tyr-50, Leu-51, Phe-52, Asn-69 CD47 377 Tyr-37, Thr-49, Phe-50, Tyr-37, Glu-97, Asp-51, Ala-53, Glu-97, Glu-100, Val-98, Glu-100, Leu-101, Leu-101, Thr-102, Glu-104, Glu-104, Glu- Glu-106, Gly-107 106 SIRP-.alpha. 378 Tyr-50, Gln-52, Pro-58, Tyr-50, Gln-52, Ser-66, Thr-67, Ser-77 Ser-66, Thr-67 CD206 379 Phe-726, Thr-727, Trp-728, Phe-726, Trp- Pro-733, Glu-737, 728, Trp- Asn-738, Trp-739, Ala-740, 739, Glu-743, Glu-743, Tyr-747, Tyr-747, Glu-751, Asn-765, Asp-766 Glu-751 TGM2 380 Cys-277, His-335, Asp-358 Cys-277, His-335, Asp-358
[0252] Given the large number of immunologically important signaling proteins that are targeted by the peptides of the invention, the data suggests that the peptides act in a pleiotropic manner, making many different interactions that sum together to create a broad anti-inflammatory response. This may make possible a reduction in disease conditions without the toxicity observed in the use of more highly-targeted chemotherapeutic agents.
Example 5: Binding of Peptides to Histone Modifying Enzymes and Ribonuclease Reductase
[0253] A number of the peptides of the invention were observed to share structural characteristics of the N-terminal regions of histones. Accordingly, representative peptides were evaluated in silico for their ability to bind to histone modification enzymes. In this manner, it was discovered that the peptides of the invention have high binding affinity for histone methyl transferase (HMT)(NCBI Acc. No. NP_048968.1; SEQ ID NO: 376), binding close to the active site of the enzyme. Predicted binding energies of select peptides of the invention for HMT, calculated using the ClusPro.TM. algorithm, are shown in Table 14. Again, the predicted binding energies correlate well with the predicted energies for binding RelB.
TABLE-US-00036 TABLE 14 Binding Affinities of Select Peptides to HMT, MKK7, and RNR RP SEQ ID # Sequence NO: RelB HMT 185 FFKKFFKKFK 123 -920.6 -846.4 186 KFKKFFKKFF 124 -919.6 -795.7 183 FFRKFAKRFK 122 -933.2 -839.4 182 KFRKAFKRFF 121 -944.8 -826.6 118 FFFRFFFNFN 44 -1,139.9 -1,000.2 394 NFNFFFRFFF 33 -1,286.6 -998.4 389 FRFKFA 257 -1,009.8 -836.8 390 FRFKFKF 256 -1,083.3 -906.6 391 FRFKFKFR 254 -1,190.8 -949.2 392 RFQFKFRF 255 -1,170.3 -962.2 387 AAKKAAKKAK 173 -301.6 -334.5 *All binding affinities are in kcal/mol.
[0254] A model of Histone Methyl Transferase (HMT) bound by RP-182 is shown in FIG. 13. The circled amino acids are the active site of the histone methyl transferase enzyme. Inhibition of methyl transferase activity by RP-182 is expected since RP-182 binds to at least one residue of the active site, in a manner that appears to obstruct access to the active site. A non-exhaustive list of amino acid residues in HMT that are bound by the peptides of the invention is shown in Table 13, above.
[0255] Peptides of the invention are also observed to display strong predicted affinities to MAP kinase kinase 7 (MKK7; SEQ ID NO: 166), a member of the mitogen-activated protein kinase kinase family involved in signal transduction mediating cell responses to proinflammatory cytokines, and therefore likely involved in peptides' anti-inflammatory activity. The predicted affinity of e.g. RP-182 for MKK7 is -738.2 kcals/mol.
[0256] In addition, peptides of the invention were observed to display substantial predicted affinities to ribonuclease reductase (RNR; SEQ ID NO: 168) also known as ribonucleoside diphosphate reductase. This is an enzyme that catalyzes the formation of deoxyribonucleotides from ribonucleotides. Deoxyribonucleotides in turn are used in the synthesis of DNA. The reaction catalyzed by RNR is strictly conserved in all living organisms. Furthermore, RNR plays a critical role in regulating the total rate of DNA synthesis, so that DNA to cell mass is maintained at a constant ratio during cell division and DNA repair. A somewhat unusual feature of the RNR enzyme is that it catalyzes a reaction that proceeds via a free radical mechanism of action. The substrates for RNR are ADP, GDP, CDP and UDP. dTDP (deoxythymidine diphosphate) is synthesized by another enzyme (thymidylate kinase) from dTMP (deoxythymidine monophosphate). The predicted affinity of e.g. RP-182 for RNR is -814.0 kcals/mol.
Example 6: Binding of Peptides to Targets Associated with Macrophage Activation
[0257] Peptides of the invention are also predicted to interact with several proteins relevant to macrophage activity and apoptosis, properties associated with inflammation and with tumor genesis and metastasis. Targets identified to date include CD47, SIRP-.alpha., CD206, TGM2, LEGUMAIN, DC-SIGN, CSF1, CSF1R, and IL34.
[0258] CD47 (or "Cluster of Differentiation 47"), also known as integrin associated protein (IAP), is a transmembrane protein that belongs to the immunoglobulin superfamily. CD47 protein partners with membrane-bound cellular adhesion receptors known as integrins and also binds the ligands thrombospondin-1 (TSP-1) and signal-regulatory protein alpha (SIRP-.alpha.). CD47 is involved in a range of cellular processes, including apoptosis, proliferation, adhesion, and migration. Furthermore, it plays a key role in immune and angiogenic responses. CD47 is expressed in many types of human cells and has been found to be overexpressed in many different types of tumors. The overexpression of CD47 has received considerable attention as a possible protective agent for human cancers. By binding to SIRP-.alpha. on the surface of macrophages, CD47 is believed to send a "don't eat me" signal that disables the macrophages from attacking the cancer cell.
[0259] CD206 and TGM2 have likewise been identified as potentially important regulators of macrophage activity. CD206 is a C-type lectin, primarily present on the surface of macrophages and dendritic cells. It is the first member of a family of endocytic receptors that includes Endo180 (CD280), M-type PLA2R, and DEC-205 (CD205). The receptor recognizes terminal mannose, N-acetylglucosamine, and fucose residues that make up glycans, which are attached to proteins found on the surface of some microorganisms. Accordingly, the CD206 receptor appears to play a role in both the innate and adaptive immune systems. In addition, tumor-associated macrophages may use CD206 to ingest collagen, yielding degradation products capable of nourishing both themselves and tumor cells, and weakening collagen binding of tumor cells so as to encourage metastasis.
[0260] TGM2 belongs to a family of enzymes that catalyze the calcium-dependent translational modification of proteins. The family members are found both intracellularly and extracellularly. TGM2 is unique in the family because of its multi-functionality and specialized structure, which includes four distinct domains: an N-terminal .beta.-sandwich that contains fibronectin and integrin binding sites; a catalytic core that contains the catalytic triad for acyl-transfer reactions (Cys-277, His-335, and Asp-358); and two C-terminal .beta.-barrel domains, with the second having a phospholipase-binding sequence. TGM2 has been implicated as a regulator of extracellular matrix functions, including cell adhesion and migration, cellular growth and differentiation, apoptosis, tumor growth, and wound healing. Although TGM2 is ubiquitously expressed, it is most highly expressed in M2 macrophages. Furthermore, increased TGM2 levels are associated with scleroderma, lung and kidney fibrosis, worsening symptoms for diabetes, arthritis, and EAE, and poor outcomes in a number of different cancers, all of which can be linked to M2 macrophages.
[0261] Predicted binding energies of select peptides of the invention for CD47 (NCBI Acc. No. XP_005247966.1; SEQ ID NO: 377), SIRP-.alpha. (GenBank Acc. No. AAH26692.1; SEQ ID NO: 378), CD206 (NCBI Acc. No. NP_002429.1; SEQ ID NO: 379), and TGM2 (GenBank Acc. No. AAB95430.1; SEQ ID NO: 380) calculated using the ClusPro.TM. algorithm, are shown in Table 15. As with the other targets discussed above, the predicted binding energies correlate well with the predicted energies for binding RelB.
[0262] LEGUMAIN is a protein that in humans is encoded by the LGMN gene. This gene encodes a cysteine protease, legumain that has a strict specificity for hydrolysis of asparaginyl bonds. This enzyme may be involved in the processing of bacterial peptides and endogenous proteins for MHC class II presentation in the lysosomal/endosomal systems. Enzyme activation is triggered by acidic pH and appears to be autocatalytic. Protein expression occurs after monocytes differentiate into dendritic cells. A fully mature, active enzyme is produced following lipopolysaccharide expression in mature dendritic cells. Overexpression of this gene may be associated with the majority of solid tumor types. LEGUMAIN is also overexpressed in M2 macrophages, and inhibition of its activity by the disclosed peptides is expected to downregulate M2-activated macrophages.
[0263] DC-SIGN (Dendritic Cell-Specific Intercellular adhesion molecule-3-Grabbing Non-integrin) also known as CD209 (Cluster of Differentiation 209) is a protein that in humans is encoded by the CD209 gene. DC-SIGN is a C-type lectin receptor present on the surface of both macrophages and dendritic cells. DC-SIGN on macrophages recognizes and binds to mannose type carbohydrates, a class of pathogen associated molecular patterns PAMPs commonly found on viruses, bacteria and fungi. This binding interaction activates phagocytosis. On myeloid and pre-plasmacytoid dendritic cells DC-SIGN mediates dendritic cell rolling interactions with blood endothelium and activation of CD4+ T cells, as well as recognition of pathogen haptens. DC-SIGN is significantly overexpressed in M2 macrophages, and inhibition of its activity by the disclosed peptides is expected to downregulate M2-activated macrophages.
TABLE-US-00037 TABLE 15 Binding Affinities of Select Peptides to CD47, SIRP-.alpha., CD206, and TGM2 RP SEQ ID # Sequence NO: RelB SIRP-.alpha. CD47 CD206 TGM2 185 FFKKFFKKFK 123 -920.6 -799.2 -639.3 -807.1 -827.2 186 KFKKFFKKFF 124 -919.6 -711.8 -637.4 -881.3 -885.3 183 FFRKFAKRFK 122 -933.2 -834.2 -658.1 -786.7 -860.7 182 KFRKAFKRFF 121 -944.8 -733.1 -723.1 -844.5 -869.1 118 FFFRFFFNFN 44 -1,139.9 -805.2 -751.5 -1,048.7 n/a 394 NFNFFFRFFF 33 -1,286.6 -854.2 -751.5 -986.6 n/a 389 FRFKFA 257 -1,009.8 -934.6 -688.3 -861.9 n/a 390 FRFKFKF 256 -1,083.3 -887.2 -783.5 -978.1 n/a 391 FRFKFKFR 254 -1,190.8 -932.1 -790.1 -941.3 n/a 392 RFQFKFRF 255 -1,170.3 -982.5 -792.1 -981.6 n/a 387 AAKKAAKKAK 173 -301.6 -392.3 -308.7 -416.6 n/a *All binding affinities are in kcal/mol.
[0264] FIG. 14 (left panel) shows a model of the ecto-domain of a CD47 dimer (top view) (SEQ ID NO: 377), with dark gray shaded surfaces representing the polar and non-polar amino acids that are involved in the binding of CD47 to the SIRP-.alpha. receptor, wherein the non-polar amino acids are circled. FIG. 14 (right panel) is a model of the ecto-domain of the CD47 dimer when bound by RP-183 (SEQ ID NO: 121). Based on this predicted interaction between RP-183 and CD47, peptides of the invention are expected to block the interaction between CD47 and SIRP-.alpha..
[0265] FIG. 15 shows a model of a SIRP-.alpha. dimer (SEQ ID NO: 378), with dark gray shaded surfaces representing the polar and non-polar amino acids involved in its binding to CD47 (see left-most dimer). In a slightly-skewed view of the same SIRP-.alpha. dimer bound by RP-183 (SEQ ID NO: 122) (see right-most dimer), it can be seen that RP-183 binds tightly to the amino acids involved in binding to the CD47 receptor. It therefore appears that RP-183 (and other peptides of the invention) block the interaction between CD47 and SIRP-.alpha. by two distinct mechanisms, binding to the corresponding binding sites in both CD47 and SIRP-.alpha.. Thus, predicted activities associated with the peptides of the invention include thwarting of an important defense mechanism for cancer cells.
[0266] Peptides of the invention are also predicted to block key sites on the CD206 receptor subunit. FIG. 16 shows a model of CD206 (SEQ ID NO: 379) bound by RP-182 (SEQ ID NO: 121). The dark gray shaded tyrosine residue on the bend region of CD206 (left-most molecule) forms a planar, hydrophobic stacking interactions with the mannose ligands on the surface of target cells. The remaining dark gray shaded amino acids are acidic residues that help chelate the required calcium ion necessary for stable interactions with the mannose receptor. The RP-182 peptide (seen in mesh on the right-most molecule) blocks activity by interacting with both of these key sites on the receptor subunit. Peptides of the invention are therefore expected to reduce the viability of M2 macrophages, which has been experimentally confirmed (as set forth below).
[0267] Furthermore, peptides of the invention are predicted to block the active site of TGM2. FIG. 17 (left panel) shows a model of TGM2 (SEQ ID NO: 380) with the active site residues in the center. FIG. 17 (right panel) shows the same model of TGM2 bound by RP-182 (SEQ ID NO: 121), which is shaded dark gray. As can be seen, RP-182 is predicted to bind to TGM2 in a manner that completely covers the active site, thereby obstructing substrate access and inhibiting TGM2 function. Significantly, decreased levels of TGM2 is associated with reduced NF-kB activation, so the interaction of the polypeptides of the invention with TGM2 would appear to reinforce and/or augment their suppression of NF-kB activity.
[0268] Non-exhaustive lists of specific amino acid residues in CD47, SIRP-.alpha., CD206, and TGM2 that are bound by the peptides of the invention are shown in Table 13, above.
Example 7: Binding of Peptides to Checkpoint Inhibitors and Related Targets
[0269] It has also been observed that peptides of the present invention display substantial affinity to checkpoint inhibitor proteins and their ligands. Such proteins, including cytotoxic T-lymphocyte antigen 4 (CTLA-4), PD-1, and other inhibitory coreceptors, expressed on the surface of effector immune cells, when activated appear to exhaust the activity of the immune cells, serving as immune checkpoints in order to prevent uncontrolled immune reactions. Tumor cells often express ligands to the checkpoint inhibitors, e.g. PD-L1 and PD-L2, attenuating the capacity of the immune system to attack the tumor.
[0270] In particular, programmed cell death protein 1, also known as PD-1 and CD279 (cluster of differentiation 279), is a protein that in humans is encoded by the PDCD1 gene. PD-1 is a cell surface receptor that belongs to the immunoglobulin superfamily and is expressed on T cells and pro-B cells. PD-1 binds two ligands, PD-L1 and PD-L2. PD-1, functioning as an immune checkpoint plays an important role in downregulating the immune system by preventing the activation of T-cells, which in turn reduces autoimmunity and promotes self-tolerance. The inhibitory effect of PD-1 is accomplished through a dual mechanism of promoting apoptosis (programmed cell death) in antigen specific T-cells in lymph nodes while simultaneously reducing apoptosis in regulatory T cells (suppressor T cells).
[0271] Programmed death-ligand 1 (PD-L1) also known as cluster of differentiation 274 (CD274) or B7 homolog 1 (B7-H1) is a protein that in humans is encoded by the CD274 gene. Programmed death-ligand 1 (PD-L1) is a 40 kDa type 1 transmembrane protein that has been speculated to play a major role in suppressing the immune system during particular events such as pregnancy, tissue allografts, autoimmune disease and other disease states such as hepatitis. Normally the immune system reacts to foreign antigens where there is some accumulation in the lymph nodes or spleen that triggers a proliferation of antigen-specific CD8+ T cell. The formation of PD-1 receptor/PD-L1 or B7.1 receptor/PD-L1 ligand complex transmits an inhibitory signal which reduces the proliferation of these CD8+ T cells at the lymph nodes and supplementary to that PD-1 is also able to control the accumulation of foreign antigen specific T cells in the lymph nodes through apoptosis which is further mediated by a lower regulation of the gene Bcl-2.
[0272] As illustrations of the binding of peptides of the present invention with checkpoint inhibitors and their ligands, the predicted affinity of RP-182 to PD-1 is -742.9, and that of RP-621 is -1,008.8. The affinity of RP-182 to PD-L1 is -677.4, and that of RP-621 to PD-L1 is -1,010.6. As with inflammatory targets, there is a striking correlation among predicted affinities to several other checkpoint inhibitors and their ligands, as well as other proteins known to play a role in modulating the immune apparatus. These include: TIM-1 (believed to play a role in T-helper cell development: predicted affinity to RP-182, -850.1); CTLA-4 (checkpoint inhibitor: predicted affinity to RP-182, -663.2); ADORA2a (modulates activity of neutrophils and mast cells: predicted affinity to RP-182, -938.7); OX40 (secondary co-stimulatory immune checkpoint: predicted affinity to RP-182, -759.9); IDO (immune checkpoint: predicted affinity to RP-182, -934.0); LAG-3 (immune checkpoint receptor: predicted affinity to RP-182, -873.1); CD73 (enzyme limiting T cell activity through adenosine receptor signaling: predicted affinity of CD73-I to RP-182, -808.7; predicted affinity of CD73-II to RP-182, -949.1); Arginase-1 (blocks activity of cytotoxic T lymphocytes: predicted affinity to RP-182, -984.2); Colony Stimulating Factor 1 (blockade shown to upregulate checkpoint molecules, as well as reprogramming macrophage responses; predicted affinity of CSF1 to RP-182, -854.7; predicted affinity of CSF1D to RP-182, -847.1; predicted affinity of CSF1R to RP-182, -774.1); and IL34 (also activates CSF1R; predicted affinity to RP-182, -828.5).
Example 8: Binding of Peptides to MKK7
[0273] Dual specificity mitogen-activated protein kinase kinase 7, also known as MAP kinase kinase 7 or MKK7, is an enzyme that in humans is encoded by the MAP2K7 gene. This protein is a member of the mitogen-activated protein kinase kinase family. The MKK7 protein exists as six different isoforms with three possible N-termini (.alpha., .beta., and .gamma. isoforms) and two possible C-termini (1 and 2 isoforms). MKK7 is involved in signal transduction mediating the cell responses to proinflammatory cytokines, and environmental stresses. This kinase specifically activates MAPK8/JNK1 and MAPK9/JNK2, and this kinase itself is phosphorylated and activated by MAP kinase kinase kinases including MAP3K1/MEKK1, MAP3K2/MEKK2, MAP3K3/MEKK5, and MAP4K2/GCK.
Example 9: Binding of Peptides to Serum Albumin
[0274] It is well-known that the most abundant protein in the circulation is serum albumin. It is also known that solid tumors will take up serum albumin into their cells (through the process of pinocytosis) and use it as an energy source. Therefore, peptides of the invention were evaluated in silico for their ability to bind to human serum albumin (HSA)(NCBI Acc. No. NP_000468.1; SEQ ID NO: 381). It was discovered that peptides of the invention have the capacity to bind to HSA with high affinity. Predicted binding energies of select peptides of the invention for binding to HSA are shown in Table 16, below.
TABLE-US-00038 TABLE 16 Binding Affinities of Select Peptides to Human Serum Albumin (HSA) RP SEQ ID # Sequence NO: RelB HSA 185 FFKKFFKKFK 123 -920.6 -880.2 186 KFKKFFKKFF 124 -919.6 -850.5 183 FFRKFAKRFK 122 -933.2 -860.1 182 KFRKAFKRFF 121 -944.8 -789.0 118 FFFRFFFNFN 44 -1,139.9 -1,064.7 394 NFNFFFRFFF 33 -1,286.6 -1,016.5 389 FRFKFA 257 -1,009.8 -904.8 390 FRFKFKF 256 -1,083.3 -1,046.0 391 FRFKFKFR 254 -1,190.8 -1,021.9 392 RFQFKFRF 255 -1,170.3 -1,037.4 387 AAKKAAKKAK 173 -301.6 -410.7 *All binding affinities are in kcal/mol.
[0275] FIG. 18 is a model of HSA (shown in light gray) bound by RP-183 (dark gray). The computational modeling has identified a number of possible peptide binding sites on HSA. Therefore, it is believed that a single HSA molecule is able to bind to multiple peptides of the invention. The binding interaction between peptides of the invention and HSA suggest that HSA could be used as an in vivo carrier of the peptides. In this manner, HSA could protect the peptides from degradation in the blood and carry the peptides to sites of action, such as sites of inflammation and/or cancer cells, thereby increasing the efficacy of the peptides.
Example 10: In Vitro Modulation of NF-kB Activity
[0276] NF-kB activity was monitored using the a 3T3-L1 preadipocyte cell line stably transformed with a Nfkb-RE/GFP construct, as described in Shen et al. (2013), "Adipocyte reporter assays: Application for identification of anti-inflammatory and antioxidant properties of mangosteen xanthones," Mol. Nutr. Food Res. 00:1-9, the entire contents of which are incorporated herein by reference. NF-kB expressing adipocyte reporter cells were plated in DMEM in wells of a 24-well plate, at a seeding density of 5.times.10.sup.4. On the second and third days post-plating, test peptides were individually added to the wells to a final concentration of 0.01 .mu.M. The test peptides included RP-398 (SEQ ID NO: 155), and RP-185 (SEQ ID NO: 123). On day 4 post-plating, lipopolysaccharide was added to the medium to a final concentration of 20 ng/ml. Finally, on day 5 post-plating, the cells were harvested and a fluorescence assay performed to detect GFP expression levels.
[0277] In this experiment, NF-kB expression was reduced approximately 58% relative to control cells that were not exposed to RP-398 or RP-185 peptide.
Example 11: In Vitro Modulation of Macrophage Activity
[0278] A frequently observed phenotype associated with tumor genesis and metastasis is the polarization of macrophage cells into the "M2" transition state, in which they are in an inflammatory state. Such macrophages are among those designated as "tumor-associated macrophages" (TAMs). To determine whether the peptides of the invention could influence macrophage polarization, the following experiment was performed.
[0279] Primary bone marrow cells were collected from male C57BL/6J (The Jackson Laboratory, Bar Harbor, Me.). Mouse bone marrow macrophages were differentiated in vitro from the primary bone marrow cells by culturing in Dulbecco's minimal essential medium (DMEM) with 10% FBS and 30 ng/ml murine M-CSF (colony stimulating factor) for 6 days. At day 7, macrophages were plated into 12-well plates and cultured in DMEM (10% FBS) with (i) IL-4 peptide (20 ng/mL), (ii) INF-.gamma. (10 ng/mL), or (iii) neither IL-4 nor INF-7 for 24 hours. After 24 hours, the media was replaced with pure DMEM and the cells were cultured for an additional 48 hours. The resulting macrophages were (i) M2-polarized, (ii) M1-polarized, or (iii) undifferentiated, respectively.
[0280] A macrophage sample containing approximately 70,000 undifferentiated macrophages per ml was incubated for 72 hours with 100 nM RP-182 (SEQ ID NO: 121). Following the incubation, a count of viable cells revealed that there were 68,000 cells per ml. Similarly, incubating M1-polarized macrophages for 72 hours with 100 nM RP-182 resulted in a viable cell count of 68,000 cells per ml. Thus, the RP-182 had little effect on the viability of M1 macrophages. In contrast, incubating M2-polarized macrophages for 72 hours with 100 nM RP-182 resulted in a viable cell count of only 20,000 cells per ml. The results indicate that RP-182 reduces the viability of M2 macrophages.
Example 12: Downregulation of Checkpoint Inhibitors and Ligands
[0281] Based on their predicted affinity to checkpoint inhibitors (e.g. PD-1) and their ligands (e.g. PD-L1 and PD-L2), the polypeptides of the invention were also evaluated to determine whether the concentration of these proteins in treated tissue would be downregulated in vivo. In one experiment, tumors in transgenic p53/KRAS mice were allowed to grow to approximately 100 m.sup.3 in volume, and the animals were then treated daily subcu for one week with either vehicle only, or 10 mg/kg RP-182, following which the animals were sacrificed and the tumors resected, formalin-fixed, and stained with antibodies to PD-1 (FIG. 19), PD-L1 and PD-L2 (FIG. 20). It is clear from the figures that both the checkpoint inhibitor PD-1 and each of its ligands PD-L1 and PD-L2 are significantly downregulated in vivo in tissue treated with peptides of the present disclosure.
Example 13: Suppression of Tumor Growth
[0282] The polypeptides of the invention were also tested for their effect on tumor growth in a mouse model of non-metastatic breast cancer. MCF-7 human non-metastatic breast cancer cells were cultured at 37.degree. C., 5% CO2 in normal growth media. Cells were harvested at 80% to 90% confluence. Immune compromised athymic nude mice (J:NU) were divided into 2 groups (9 animals per group). All mice were injected with .about.4.5.times.10.sup.6 MCF-7 cells which had been stained with VIVO Tracker 680 and suspended in 200 .mu.l of PBS/Matrigel mixture. Cells were injected subcutaneously on the dorsal surface of treated animals using a 22 gauge needle fitted with a 500 .mu.l syringe.
[0283] Animals were designated vehicle and peptide treated. The peptide treated animals were treated with the RP-397 polypeptide (SEQ ID NO: 194). Freshly prepared RP-397 peptide was dissolved in sterile saline at a concentration of 100 .mu.M and was used to treat the animals in the peptide group. Vehicle treated animals were injected with saline buffer alone. All treatments were injected into the tumor mass two times weekly for 5 weeks using a 271/2 gauge needle fitted with a 1 ml syringe. Animal weights and tumor volumes were measured 3 times weekly and the fluorescence labeling was followed by VIVO Tracker 680 and IVIS Imaging. The results are shown in Table 17, below.
TABLE-US-00039 TABLE 17 Suppression of Tumor Growth Using RP-397 Avg. Rate of Body Body Tumor Tumor Weight Weight Weight Growth Before After Vehicle 1.5 g 63 25.2 30.2 RP-397 0.75 g* 20* 25.1 30.1 The rate of tumor growth was measured in mm.sup.3/day. The "*" denotes a statistically significant difference from the vehicle control.
[0284] The data shows that polypeptides of the invention can suppress tumor growth in vivo.
Example 14: Administering Peptides in Combination with Chemotherapy
[0285] Given the significant role of inflammation in tumor genesis and metastasis, as well as the known association of M2 macrophage activity with tumor development, it was anticipated that the administration of peptides of the invention could positively influence the outcomes of cancer treatment.
[0286] To test this theory, cohorts of immunocompromised ("nude") mice were injected with .about.5.times.10.sup.6 human triple-negative breast cancer cells (MDA-MB-231) under the upper left teat. Following this administration, one cohort received only vehicle; two of the cohorts received the chemotherapeutic agent Gemcitabine, at a q4d dose of 40 mg/kg of body weight. One of these cohorts also received RP-182 (SEQ ID NO: 121) at a daily dose of 5 mg/kg body weight; and a fourth cohort received only RP-182 at a daily dose of 5 mg/kg body weight. Beginning on day 32 of the study, in the Gemcitabine+RP-182 cohort, concentrations of RP-182 were increased to 20 mg/kg body weight. Tumor volume was assessed at various time points following initial cell administration (FIG. 21). After 50 days, the mice were sacrificed.
[0287] The data demonstrates that, as compared to treatment with Gemcitabine alone, combined treatment with Gemcitabine and polypeptides of the invention resulted in reductions in mean tumor volume. When RP-182 concentration was increased to 20 mg/kg body weight, the increase in tumor volume was essentially halted.
[0288] In a second experiment, xenografts of C42B prostate cancer cells were introduced into four cohorts of mice, and the tumors allowed to grow to approximately 100 m.sup.3 before treatment. One cohort was treated only with vehicle; a second with Docetaxel at 2.5 mg/kg body weight administered weekly; a third with RP-182 administered daily subcu at 10 mg/kg body weight; and a fourth with both Docetaxel at 2.5 mg/kg weekly and RP-182 at 10 mg/kg daily. Tumor volume was assessed at various time points following initial cell administration (FIG. 22); after 27 days, the mice were sacrificed. Similarly, the administration of RP-182 plus Docetaxel resulted in decreases in mean tumor volume compared to Docetaxel alone.
[0289] It is anticipated that the peptides of the invention will produce synergistic effects when administered with chemotherapeutic agents other than Gemcitabine and Docetaxel, as well as checkpoint inhibitor therapies and other immunotherapies. In particular, the peptides of the invention may be particularly useful when used in conjunction with recently-developed CAR-T (chimeric antigen receptor/T cell) therapies. Such therapies, while destroying tumor cells, create a very high systemic burden of dead cell material, overstimulating the immune system and creating a "cytokine storm" which can be fatal to the patient.
Embodiments
[0290] The following embodiments are provided to illustrate aspects of the present invention.
[0291] 1. An anti-inflammatory composition comprising a peptide, wherein the peptide is 3 to 24 amino acid residues in length and comprises a striapathic region consisting of alternating X.sub.m and Y.sub.n modules, wherein m and n are positive integers that identify different modules, wherein each X.sub.m module consists of a sequence according to the formula X.sub.ma-X.sub.mb-X.sub.m-X.sub.ma-X.sub.me, wherein X.sub.ma is selected from the group consisting of a naturally occurring hydrophilic amino acid, a non-naturally occurring hydrophilic amino acid, and a hydrophilic amino acid mimetic, and wherein X.sub.mb, X.sub.mc, X.sub.md and X.sub.me are each individually absent or selected from the group consisting of a naturally occurring hydrophilic amino acid, a non-naturally occurring hydrophilic amino acid, and a hydrophilic amino acid mimetic, wherein each Y.sub.b module consists of a sequence according to the formula Y.sub.na-Y.sub.nb-Y.sub.nc-Y.sub.na-Y.sub.ne, wherein Y.sub.na is selected from the group consisting of a naturally occurring hydrophobic amino acid, a non-naturally occurring hydrophobic amino acid, and a hydrophobic amino acid mimetic, and wherein Y.sub.nb, Y.sub.nc, Y.sub.nd, and Y.sub.ne are each individually absent or selected from the group consisting of a naturally occurring hydrophobic, a non-naturally occurring hydrophobic amino acid, and a hydrophobic amino acid mimetic, and wherein the peptide binds to the dimerization site on a NFkB Class II protein.
[0292] 2. The anti-inflammatory composition of embodiment 1, wherein each X.sub.m module consists of a sequence according to the formula X.sub.ma-X.sub.mb-X.sub.mc-X.sub.ma, and each Y.sub.n module consists of a sequence according to the formula Y.sub.na-Y.sub.nb-Y.sub.nc-Y.sub.na.
[0293] 3. The anti-inflammatory composition of embodiment 1, wherein each X.sub.m module consists of a sequence according to the formula X.sub.ma-X.sub.mb-X.sub.mc, and each Y.sub.n module consists of a sequence according to the formula Y.sub.na-Y.sub.nb-Y.sub.nc.
[0294] 4. The anti-inflammatory composition of any one of embodiments 1 to 3, wherein the peptide also binds to human serum albumin.
[0295] 5. The anti-inflammatory composition of any one of embodiments 1 to 4, wherein the striapathic region of the peptide contains at least two X.sub.m modules (X.sub.1, X.sub.2, and X.sub.3) and at least two Y.sub.n modules (Y.sub.1, Y.sub.2, and Y.sub.3).
[0296] 6. The anti-inflammatory composition of any one of embodiments 1 to 5, wherein the striapathic region of the peptide contains at least seven amino acid residues.
[0297] 7. The anti-inflammatory composition of any one of embodiments 1 to 6, wherein the striapathic region of the peptide has a length of 7 to 12 amino acid residues.
[0298] 8. The anti-inflammatory composition of any one of embodiments 1 to 7, wherein the striapathic region of the peptide constitutes at least 25% of the length of the peptide.
[0299] 9. The anti-inflammatory composition of any one of embodiments 1 to 8, wherein the striapathic region of the peptide has an amphipathic conformation under physiological conditions.
[0300] 10. The anti-inflammatory composition of embodiment 9, wherein the striapathic region of the peptide has an amphipathic 3.sub.10-helical conformation, an amphipathic .alpha.-helical conformation, or an amphipathic n-helical conformation when bound to the NFkB Class II protein.
[0301] 11. The anti-inflammatory composition of embodiment 10, wherein the amphipathic 3.sub.10-helical, .alpha.-helical, or n-helical conformation includes a hydrophobic portion having a facial arc of at least 100.degree..
[0302] 12. The anti-inflammatory composition of any one of embodiments 1 to 11, wherein the striapathic region contains hydrophobic amino acid residues having a total volume of at least 650 cubic angstroms.
[0303] 13. The anti-inflammatory composition of any one of embodiments 1 to 12, wherein the striapathic region is characterized by a ratio of the sum of the volume of hydrophobic amino acid residues to the sum of the volume of hydrophilic amino acid residues, wherein the ratio is at least 0.75 or higher.
[0304] 14. The anti-inflammatory composition of embodiment 9, wherein the striapathic region of the peptide comprises at least one proline residue and adopts an amphipathic conformation that includes a proline-rich helix.
[0305] 15. The anti-inflammatory composition of embodiment 9, wherein the striapathic region of the peptide adopts an amphipathic beta-strand conformation.
[0306] 16. The anti-inflammatory composition of any one of embodiments 1 to 13, wherein the striapathic region includes a sequence selected from the group of sequences defined by Formula I: Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c (Formula I).
[0307] 17. The anti-inflammatory composition of embodiment 16, wherein the module Y.sub.1a-Y.sub.1b-Y.sub.1c has a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), Tyr-Tyr-Tyr (YYY), Leu-Leu-Leu (LLL), Cys-Cys-Cys (CCC), Met-Met-Met (MMM), Val-Val-Val (VVV), and Ile-Ile-Ile (III).
[0308] 18. The anti-inflammatory composition of embodiment 16, wherein the module Y.sub.1a-Y.sub.1b-Y.sub.1c has a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), and Tyr-Tyr-Tyr (YYY).
[0309] 19. The anti-inflammatory composition of any one of embodiments 16 to 18, wherein the module Y.sub.2a-Y.sub.2b-Y.sub.2c has a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), Tyr-Tyr-Tyr (YYY), Leu-Leu-Leu (LLL), Cys-Cys-Cys (CCC), Met-Met-Met (MMM), Val-Val-Val (VVV), and Ile-Ile-Ile (III).
[0310] 20. The anti-inflammatory composition of any one of embodiments 16 to 18, wherein the module Y.sub.2a-Y.sub.2b-Y.sub.2c has a sequence selected from the group consisting of Phe-Phe-Phe (FFF), Trp-Trp-Trp (WWW), and Tyr-Tyr-Tyr (YYY).
[0311] 21. The anti-inflammatory composition of embodiment 16, wherein the striapathic region includes a sequence selected from the group consisting of FFF-X.sub.1a-FFF (SEQ ID NO: 1), WWW-X.sub.1a-WWW (SEQ ID NO: 2), and YYY-X.sub.1a-YYY (SEQ ID NO: 3).
[0312] 22. The anti-inflammatory composition of embodiment 16, wherein the sequence of the three modules is selected from the group consisting of LLL-X.sub.1a-LLL (SEQ ID NO: 4), CCC-X.sub.1a-CCC (SEQ ID NO: 5), MMM-X.sub.1a-MMM (SEQ ID NO: 6), VVV-X.sub.1a-VVV (SEQ ID NO: 7), and III-X.sub.1a-III (SEQ ID NO: 8).
[0313] 23. The anti-inflammatory composition of any one of embodiments 16 to 22, wherein X.sub.1a is selected from the group consisting of Arg (R), His (H), and Lys (K).
[0314] 24. The anti-inflammatory composition of any one of embodiments 16 to 22, wherein X.sub.1a is selected from the group consisting of Glu (E), Gln (Q), Asn (N), and Asp (D).
[0315] 25. The anti-inflammatory composition of any one of embodiments 16 to 24, wherein the striapathic region includes a sequence selected from the group of sequences defined by Formula II or the group of sequences defined by Formula III: Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.2a-Y- .sub.3a-X.sub.3a (Formula II); X.sub.2a-Y.sub.3a-X.sub.3a-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y- .sub.2b-Y.sub.2c (Formula III).
[0316] 26. The anti-inflammatory composition of embodiment 25, wherein X.sub.2a and X.sub.3a are each individually selected from the group consisting of Arg (R), His (H), Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D).
[0317] 27. The anti-inflammatory composition of embodiment 25, wherein X.sub.2a and X.sub.3a are each individually selected from the group consisting of Glu (E), Gln (Q), Asn (N), and Asp (D).
[0318] 28. The anti-inflammatory composition of any one of embodiments 25 to 27, wherein Y.sub.3a is selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), and Ile (I).
[0319] 29. The anti-inflammatory composition of any one of embodiments 25 to 27, wherein Y.sub.3a is selected from the group consisting of Phe (F), Trp (W), Tyr (Y), and Leu (L).
[0320] 30. The anti-inflammatory composition of embodiment 25, wherein the sequence of X.sub.2a-Y.sub.3a-X.sub.3a is selected from the group consisting of EFQ, EFE, EFN, EFD, NFQ, NFE, NFN, NFD, QFQ, QFE, QFN, QFD, DFQ, DFE, DFN, DFD, EWQ, EWE, EWN, EWD, NWQ, NWE, NWN, NWD, QWQ, QWE, QWN, QWD, DWQ, DWE, DWN, DWD, EYQ, EYE, EFN, EYD, NYQ, NYE, NYN, NYD, QYQ, QYE, QYN, QYD, DYQ, DYE, DYN, DYD, ELQ, ELE, ELN, ELD, NLQ, NLE, NLN, NLD, QLQ, QLE, QLN, QLD, DLQ, DLE, DLN, DLD, RFR, RFQ, RFE, RFN, RFD, RWR, RWQ, RWE, RWN, and RWD.
[0321] 31. The anti-inflammatory composition of embodiment 25, wherein the striapathic region comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP394, RP108-RP123, RP125-131, RP133, RP135-RP141, RP143-RP146, RP148-RP150, RP152-RP165, RP179, RP395, RP211, RP230, RP232, RP258, RP267, RP268, RP271, and RP273.
[0322] 32. The anti-inflammatory composition of embodiment 25, wherein the striapathic region comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP113 (SEQ ID NO: 39), RP118 (SEQ ID NO: 44), and RP394 (SEQ ID NO: 33).
[0323] 33. The anti-inflammatory composition of any one of embodiments 1 to 13, wherein the striapathic region includes a sequence selected from the group of sequences defined by Formula VII: Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y.sub.3a (Formula VII).
[0324] 34. The anti-inflammatory composition of embodiment 33, wherein Y.sub.2a is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0325] 35. The anti-inflammatory composition of embodiment 33, wherein Y.sub.2a is selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I), and Ala (A).
[0326] 36. The anti-inflammatory composition of any one of embodiments 33 to 35, wherein Y.sub.2b is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0327] 37. The anti-inflammatory composition of any one of embodiments 33 to 35, wherein Y.sub.2b is selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I), and Ala (A).
[0328] 38. The anti-inflammatory composition of any one of embodiments 33 to 37, wherein X.sub.1b is selected from the group consisting of Arg (R), Lys (K), and His (H).
[0329] 39. The anti-inflammatory composition of any one of embodiments 33 to 37, wherein X.sub.1b is selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0330] 40. The anti-inflammatory composition of any one of embodiments 33 to 39, wherein X.sub.2a is selected from the group consisting of Arg (R), Lys (K), and His (H).
[0331] 41. The anti-inflammatory composition of any one of embodiments 33 to 39, wherein X.sub.2a is selected from the group consisting of Asn (N), Gln (Q), Asp (D), and Glu (E).
[0332] 42. The anti-inflammatory composition of embodiment 33, wherein the sequence X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a is selected from the group consisting of Lys-Phe-Phe-Lys (KFFK; SEQ ID NO: 386), Lys-Trp-Trp-Lys (KWWK; SEQ ID NO: 387), Lys-Tyr-Try-Lys (KYYK; SEQ ID NO: 388), Lys-Phe-Trp-Lys (KFWK; SEQ ID NO: 389), Lys-Trp-Phe-Lys (KWFK; SEQ ID NO: 390), Lys-Phe-Tyr-Lys (KFYK; SEQ ID NO: 391), Lys-Tyr-Phe-Lys (KYFK; SEQ ID NO: 392), Lys-Trp-Tyr-Lys (KWYK; SEQ ID NO: 393), and Lys-Tyr-Trp-Lys (KYWK; SEQ ID NO: 394).
[0333] 43. The anti-inflammatory composition of embodiment 33, wherein the sequence X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a is selected from the group consisting of Arg-Phe-Phe-Arg (RFFR; SEQ ID NO: 395), Arg-Trp-Trp-Arg (RWWR; SEQ ID NO: 396), Arg-Tyr-Try-Arg (RYYR; SEQ ID NO: 397), Arg-Phe-Trp-Arg (RFWR; SEQ ID NO: 398), Arg-Trp-Phe-Arg (RWFR; SEQ ID NO: 399), Arg-Phe-Tyr-Arg (RFYR; SEQ ID NO: 400), Arg-Tyr-Phe-Arg (RYFR; SEQ ID NO: 401), Arg-Trp-Tyr-Arg (RWYR; SEQ ID NO: 402), and Arg-Tyr-Trp-Arg (RYWR; SEQ ID NO: 403).
[0334] 44. The anti-inflammatory composition of embodiment 33, wherein the sequence X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a is selected from the group consisting of His-Phe-Phe-His (HFFH; SEQ ID NO: 404), His-Trp-Trp-His (HWWH; SEQ ID NO: 405), His-Tyr-Try-His (HYYH; SEQ ID NO: 406), His-Phe-Trp-His (HFWH; SEQ ID NO: 407), His-Trp-Phe-His (HWFH; SEQ ID NO: 408), His-Phe-Tyr-His (HFYH; SEQ ID NO: 409), His-Tyr-Phe-His (HYFH; SEQ ID NO: 410), His-Trp-Tyr-His (HWYH; SEQ ID NO: 411), and His-Tyr-Trp-His (HYWH; SEQ ID NO: 132).
[0335] 45. The anti-inflammatory composition of any one of embodiments 33 to 44, wherein X.sub.1a is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0336] 46. The anti-inflammatory composition of any one of embodiments 33 to 44, wherein X.sub.1a is selected from the group consisting of Arg (R) and Gln (Q).
[0337] 47. The anti-inflammatory composition of any one of embodiments 33 to 46, wherein X.sub.2b is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0338] 48. The anti-inflammatory composition of any one of embodiments 33 to 46, wherein X.sub.2b is selected from the group consisting of Arg (R) and Gln (Q).
[0339] 49. The anti-inflammatory composition of any one of embodiments 33 to 48, wherein Y.sub.1a is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0340] 50. The anti-inflammatory composition of any one of embodiments 33 to 48, wherein Y.sub.1a is selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I), and Ala (A).
[0341] 51. The anti-inflammatory composition of any one of embodiments 33 to 50, wherein Y.sub.3a is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0342] 52. The anti-inflammatory composition of any one of embodiments 33 to 50, wherein Y.sub.3a is selected from the group consisting of Leu (L), Cys (C), Met (M), Val (V), Ile (I), and Ala (A).
[0343] 53. The anti-inflammatory composition of embodiment 33, wherein the striapathic region includes a sequence selected from the group consisting of F-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 9), F-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-W (SEQ ID NO: 10), W-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 11), F-X.sub.1a-X.sub.1b-FW-X.sub.2a-X.sub.2b-F (SEQ ID NO: 12), F-X.sub.1a-X.sub.1b-WF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 13), F-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-F (SEQ ID NO: 14), W-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-F (SEQ ID NO: 15), F-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-W (SEQ ID NO: 16), W-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-W (SEQ ID NO: 17), F-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 18), Y-X.sub.1a-X.sub.1b-FF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 19), F-X.sub.1a-X.sub.1b-FY-X.sub.2a-X.sub.2b-F (SEQ ID NO: 20), F-X.sub.1a-X.sub.1b-YF-X.sub.2a-X.sub.2b-F (SEQ ID NO: 21), F-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-F (SEQ ID NO: 22), Y-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-F (SEQ ID NO: 23), F-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 24), and Y-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 25), Y-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-W (SEQ ID NO: 26), W-X.sub.1a-X.sub.1b-YY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 27), Y-X.sub.1a-X.sub.1b-YW-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 28), Y-X.sub.1a-X.sub.1b-WY-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 29), Y-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 30), W-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-Y (SEQ ID NO: 31), and Y-X.sub.1a-X.sub.1b-WW-X.sub.2a-X.sub.2b-W (SEQ ID NO: 32).
[0344] 54. The anti-inflammatory composition of embodiment 53, wherein X.sub.1a, X.sub.1b, X.sub.2a, and X.sub.2b are each independently selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0345] 55. The anti-inflammatory composition of embodiment 53 or 54, wherein X.sub.1b and X.sub.2a are each independently selected from the group consisting of Arg (R), Lys (K), and His (H).
[0346] 56. The anti-inflammatory composition of any one of embodiments 33 to 55, wherein the striapathic region includes a first additional amino acid residue directly bound to Y.sub.1a of Formula VII, wherein the first additional amino acid residue is a hydrophobic amino acid residue.
[0347] 57. The anti-inflammatory composition of embodiment 56, wherein the first additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0348] 58. The anti-inflammatory composition of any one of embodiments 33 to 55, wherein the striapathic region includes a first additional amino acid residue directly bound to Y.sub.3a of Formula VII, wherein the first additional amino acid residue is a hydrophobic amino acid residue.
[0349] 59. The anti-inflammatory composition of embodiment 58, wherein the first additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0350] 60. The anti-inflammatory composition of any one of embodiments 33 to 55, wherein the striapathic region includes a first additional amino acid residue directly bound to Y.sub.1a of Formula VII, wherein the first additional amino acid residue is a hydrophilic amino acid residue.
[0351] 61. The anti-inflammatory composition of embodiment 60, wherein the first additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0352] 62. The anti-inflammatory composition of any one of embodiments 33 to 55, wherein the striapathic region includes a first additional amino acid residue directly bound to Y.sub.3a of Formula VII, wherein the first additional amino acid residue is a hydrophilic amino acid residue.
[0353] 63. The anti-inflammatory composition of embodiment 62, wherein the first additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0354] 64. The anti-inflammatory composition of 56, 57, 60, or 61, wherein the striapathic region includes a second additional amino acid residue directly bound to Y.sub.3a of Formula VII, wherein the second additional amino acid residue is a hydrophobic amino acid residue.
[0355] 65. The anti-inflammatory composition of embodiment 64, wherein the second additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0356] 66. The anti-inflammatory composition of 58, 59, 62, or 63, wherein the striapathic region includes a second additional amino acid residue directly bound to Y.sub.1a of Formula VII, wherein the second additional amino acid residue is a hydrophilic amino acid residue.
[0357] 67. The anti-inflammatory composition of embodiment 66, wherein the second additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0358] 68. The anti-inflammatory composition of embodiment 33, wherein the striapathic region comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP124, RP132, RP134, RP142, RP147, RP151, RP166-RP172, RP175, RP177, RP182, RP183, RP185, RP186, RP 424, RP190, RP194, RP198, RP199-RP202, RP204, RP206, RP207, RP209, RP210, RP212-RP216, RP218, RP219, RP425, RP225, RP227, RP233-RP239, RP398, RP241-RP247, RP250-RP256, and RP426.
[0359] 69. The anti-inflammatory composition of embodiment 33, wherein the striapathic region comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP124 (SEQ ID NO: 106), RP166 (SEQ ID NO: 112), RP182 (SEQ ID NO: 121), and RP183 (SEQ ID NO: 122).
[0360] 70. The anti-inflammatory composition of any one of embodiments 1 to 15, wherein the striapathic region includes a sequence selected from the group of sequences defined by any one of Formulas I-XLVIII and L: Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c (Formula I); Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y.sub.2b-Y.sub.2c-X.sub.- 2a-Y.sub.3a-X.sub.3a (Formula II); X.sub.2a-Y.sub.3a-X.sub.3a-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-Y.sub.2a-Y- .sub.2b-Y.sub.2c (Formula III); X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-X.sub.2a-X.sub.2b-X.sub.2c (Formula IV); Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-X.sub.2a-X.sub.2b-X.sub- .2c-Y.sub.3a-X.sub.3a (Formula V); X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b (Formula VI); Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y.sub.3a (Formula VII); Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y.sub.3a-Y- .sub.3b-X.sub.3a (Formula VIII); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y- .sub.3a-Y.sub.3b (Formula IX); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-Y- .sub.3a-X.sub.3a (Formula X); X.sub.1a-Y.sub.1a-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a-X.sub.3b-Y- .sub.3a-Y.sub.3b (Formula XI); X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a-X- .sub.3b-Y.sub.3a (Formula XII); Y.sub.1a-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X.sub.2b-X.sub.2c-Y- .sub.3a-Y.sub.3b (Formula XIII); X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-Y.sub.2a-Y- .sub.2b-Y.sub.2c (Formula XIV); Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-X- .sub.2b-X.sub.2c (Formula XV); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-Y.sub.2b-X.sub.2a-X- .sub.2b-Y.sub.3a (Formula XVI); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b (Formula XVII); X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-Y.sub.2a-Y.sub.2b-X.sub.3a (Formula XVIII); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-Y.sub.2b-X.sub.2a-Y.sub.3a-Y- .sub.3b-X.sub.3a (Formula XIX); X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-Y.sub.2a-Y.sub.2b-X.sub.3a-X.sub.3b-Y- .sub.3a-Y.sub.3b (Formula XX); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-Y.sub.2a-X.sub.2a-X.sub.2b-Y.sub.3a-Y- .sub.3b (Formula XXI); X.sub.1a-Y.sub.1a-Y.sub.1b-X.sub.2a-X.sub.2b-X.sub.2c-Y.sub.2a-X.sub.3a-Y- .sub.3a-Y.sub.3b (Formula XXII); Y.sub.1a-Y.sub.1b-X.sub.1a-Y.sub.2a-X.sub.2a-X.sub.2b-X.sub.2c-Y.sub.3a-Y- .sub.3b-X.sub.3a (Formula XXIII); X.sub.1a-X.sub.1b-Y.sub.1a-X.sub.2a-Y.sub.2a-X.sub.3a-X.sub.3b (Formula XXIV); Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-Y.sub.2a-X.sub.2a-Y.s- ub.3a-X.sub.3a-X.sub.3b (Formula XXV); X.sub.1a-X.sub.1b-Y.sub.1a-X.sub.2a-Y.sub.2a-X.sub.3a-X.sub.3b-Y.sub.3a-Y- .sub.3b-Y.sub.3c (Formula XXVI); X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.1a-Y.sub.1b-Y.sub.1c (Formula XXVII); X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1a (Formula XXVIII); Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-Y.sub.2a-Y.sub.2b-Y.sub.2c-Y- .sub.2a-X.sub.2a (Formula XXIX); X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y- .sub.1d-Y.sub.1e (Formula XXX); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.2a-Y.sub.2b-Y.sub.2c-X- .sub.2a-X.sub.2b (Formula XXXI); X.sub.1a-Y.sub.1a-X.sub.2a-Y.sub.2a-X.sub.3a-X.sub.3b-X.sub.3c-Y.sub.3a-Y- .sub.3b-Y.sub.3c (Formula XXXII); Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-X.sub.1c (Formula XXXIII); Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d (Formula XXXIV); X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-X.sub.2a-X.sub.2b-X.sub.2c-X- .sub.2a-Y.sub.2a (Formula XXXV); Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.1a-X.sub.1b-X.sub.1c-X- .sub.1a-X.sub.1e (Formula XXXVI); X.sub.1a-X.sub.1b-Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.2a-X.sub.2b-X.sub.2c-Y- .sub.2a-Y.sub.2b (Formula XXXVII); Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1a-X.sub.1c-Y.sub.2a-X.sub.2a-Y- .sub.3a-X.sub.3a (Formula XXXVIII); Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1a-X.sub.1e-Y.sub.2a (Formula XXXIX); Y.sub.1a-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.2a-Y.- sub.2b-Y.sub.2c-Y.sub.2a (Formula XL); Y.sub.1a-Y.sub.1b-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y.sub.2a-Y- .sub.2b-Y.sub.2c (Formula XLI); Y.sub.1a-Y.sub.1b-Y.sub.1c-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X.sub.1e-Y- .sub.2a-Y.sub.2b (Formula XLII); Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1e-X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1d-X- .sub.1e-Y.sub.2a (Formula XLIII); X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.2a (Formula XLIV); X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.2a-X.s- ub.2b-X.sub.2c-X.sub.2a (Formula XLV); X.sub.1a-X.sub.1b-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X.sub.2a-X- .sub.2b-X.sub.2c (Formula XLVI); X.sub.1a-X.sub.1b-X.sub.1c-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y.sub.1e-X- .sub.2a-X.sub.2b (Formula XLVII); X.sub.1a-X.sub.1b-X.sub.1c-X.sub.1a-Y.sub.1a-Y.sub.1b-Y.sub.1c-Y.sub.1d-Y- .sub.1e-X.sub.2a (Formula XLVIII); and Y.sub.1a-Y.sub.1b-X.sub.1a-Y.sub.2a-Y.sub.2b-X.sub.2a-Y.sub.3a-Y.sub.3b-X- .sub.3a-Y.sub.4a (Formula L).
[0361] 71. The anti-inflammatory composition of embodiment 70, wherein Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.2a, Y.sub.2b, Y.sub.2c, Y.sub.3a, Y.sub.3b, and Y.sub.3c are each individually selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Cys (C), Met (M), Val (V), Ile (I), and Ala (A).
[0362] 72. The anti-inflammatory composition of embodiment 70, wherein Y.sub.1a, Y.sub.1b, Y.sub.1c, Y.sub.2a, Y.sub.2b, Y.sub.2c, Y.sub.3a, Y.sub.3b, and Y.sub.3c are each individually selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0363] 73. The anti-inflammatory composition of any one of embodiments 70 to 72, wherein X.sub.1a, X.sub.1b, X.sub.1c, X.sub.2a, X.sub.2b, X.sub.2c, X.sub.3a, and X.sub.3b are each individually selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0364] 74. The anti-inflammatory composition of any one of embodiments 70 to 73, wherein X.sub.1a, X.sub.1b, X.sub.1c, X.sub.2a, X.sub.2b, X.sub.2c, X.sub.3a, and X.sub.3b are each individually selected from the group consisting of Arg (R), Lys (K), His (H), and Gln (Q).
[0365] 75. The anti-inflammatory composition of any one of embodiments 70 to 74, wherein the striapathic region includes a first additional amino acid residue directly bound to the N-terminal end of any one of Formulas I-XLVIII and L, wherein the first additional amino acid residue is a hydrophobic amino acid residue.
[0366] 76. The anti-inflammatory composition of embodiment 70, wherein the first additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0367] 77. The anti-inflammatory composition of any one of embodiments 70 to 74, wherein the striapathic region includes a first additional amino acid residue directly bound to the C-terminal end of any one of Formulas I-XLVIII and L, wherein the first additional amino acid residue is a hydrophobic amino acid residue.
[0368] 78. The anti-inflammatory composition of embodiment 77, wherein the first additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0369] 79. The anti-inflammatory composition of any one of embodiments 70 to 74, wherein the striapathic region includes a first additional amino acid residue directly bound to the N-terminal end of any one of Formulas I-XLVIII and L, wherein the first additional amino acid residue is a hydrophilic amino acid residue.
[0370] 80. The anti-inflammatory composition of embodiment 79, wherein the first additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0371] 81. The anti-inflammatory composition of any one of embodiments 70 to 74, wherein the striapathic region includes a first additional amino acid residue directly bound to the C-terminal end of any one of Formulas I-XLVIII and L, wherein the first additional amino acid residue is a hydrophilic amino acid residue.
[0372] 82. The anti-inflammatory composition of embodiment 81, wherein the first additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0373] 83. The anti-inflammatory composition of any one of embodiments 75, 76, 79, or 80, wherein the striapathic region includes a second additional amino acid residue directly bound to the C-terminal end of any one of Formulas I-XLVIII and L, wherein the second additional amino acid residue is a hydrophobic amino acid residue.
[0374] 84. The anti-inflammatory composition of embodiment 83, wherein the second additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0375] 85. The anti-inflammatory composition of any one of embodiments 77, 78, 81, or 82, wherein the striapathic region includes a second additional amino acid residue directly bound to the N-terminal end of any one of Formulas I-XLVIII and L, wherein the second additional amino acid residue is a hydrophilic amino acid residue.
[0376] 86. The anti-inflammatory composition of embodiment 81, wherein the second additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0377] 87. The anti-inflammatory composition of embodiment 70, wherein the striapathic region comprises, consists essentially of, or consists of a sequence selected from the group consisting of RP396, RP405, RP174, RP176, RP178, RP180-181, RP184, RP408, RP187, RP416, RP188, RP189, RP388, RP417, RP191-RP193, RP404, RP196, RP397, RP197, RP402, RP203, RP409, RP205, RP208, RP217, RP220-RP224, RP226, RP229, RP231, RP240, RP248, RP249, RP415, RP257, RP259-RP266, RP269, RP272, RP406, RP422, RP407, RP400, RP419, RP401, RP423, RP411, RP418, RP428, RP420, RP421, RP429, RP413, RP430, RP270.
[0378] 88. The anti-inflammatory composition of any one of embodiments 1 to 9 or 15, wherein the striapathic region includes a sequence selected from the group of sequences defined by Formula XLIX:
Y.sub.1a-X.sub.1a-Y.sub.2a-X.sub.2a-Y.sub.3a-X.sub.3a (Formula XLIX).
[0379] 89. The anti-inflammatory composition of embodiment 88, wherein Y.sub.1a, Y.sub.2a, and Y.sub.3a are each independently selected from the group consisting of Phe (F), Trp (W), Tyr (Y), Leu (L), Ile (I), Cys (C), and Met (M).
[0380] 90. The anti-inflammatory composition of embodiment 88, wherein Y.sub.1a, Y.sub.2a, and Y.sub.3a are each independently selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0381] 91. The anti-inflammatory composition of any one of embodiments 88 to 90, wherein X.sub.1a, X.sub.2a, and X.sub.3a are each independently selected from the group consisting of Arg (R), Lys (K), His (H), Gln (Q), Glu (E), Asn (N), and Asp (D).
[0382] 92. The anti-inflammatory composition of any one of embodiments 88 to 90, wherein X.sub.1a, X.sub.2a, and X.sub.3a are each independently selected from the group consisting of Arg (R), Lys (K), and His (H).
[0383] 93. The anti-inflammatory composition of any one of embodiments 88 to 92, wherein the striapathic region includes a first additional amino acid residue directly bound to Y.sub.1a of Formula XLIX, wherein the first additional amino acid residue is a hydrophilic amino acid residue.
[0384] 94. The anti-inflammatory composition of embodiment 93, wherein the first additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), His (H), Asn (N), Gln (Q), Asp (D), and Glu (E).
[0385] 95. The anti-inflammatory composition of embodiment 93, wherein the first additional amino acid residue is selected from the group consisting of Arg (R), Lys (K), and His (H).
[0386] 96. The anti-inflammatory composition of any one of embodiments 88 to 92, wherein the striapathic region includes a first additional amino acid residue directly bound to X.sub.3a of Formula XLIX, wherein the first additional amino acid residue is a hydrophobic amino acid residue.
[0387] 97. The anti-inflammatory composition of embodiment 96, wherein the first additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), (Tyr), Leu (L), Ile (I), Cys (C), and Met (M).
[0388] 98. The anti-inflammatory composition of embodiment 96, wherein the first additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and (Tyr).
[0389] 99. The anti-inflammatory composition of any one of embodiments 93 to 95, wherein the striapathic region includes a second additional amino acid residue directly bound to X.sub.3a of Formula XLIX, wherein the second additional amino acid residue is a hydrophobic amino acid residue.
[0390] 100. The anti-inflammatory composition of embodiment 99, wherein the second additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), (Tyr), Leu (L), Ile (I), Cys (C), and Met (M).
[0391] 101. The anti-inflammatory composition of embodiment 99, wherein the second additional amino acid residue is selected from the group consisting of Phe (F), Trp (W), and Tyr (Y).
[0392] 102. An anti-inflammatory composition comprising a peptide, wherein the peptide is 3 to 24 amino acids residues in length and comprises a striapathic region having at least 70% identity with the sequence NFNFFFRFFF (RP394, SEQ ID NO: 33), wherein the peptide binds to the dimerization site on a NFkB Class II protein.
[0393] 103. The anti-inflammatory composition of embodiment 102, wherein the peptide also binds to human serum albumin.
[0394] 104. The anti-inflammatory composition of embodiment 102 or 103, wherein the differences between the striapathic region of the peptide and the sequence NFNFFFRFFF (SEQ ID NO: 33) are limited to conservative or highly conservative amino acid substitutions.
[0395] 105. The anti-inflammatory composition of embodiment 102 or 103, wherein the striapathic region of the peptide differs from the sequence NFNFFFRFFF (SEQ ID NO: 33) by substitution of one or more of the phenylalanine (F) residues with an amino acid residue selected from the group consisting of Trp (W), Tyr (Y), His (H), and Leu (L).
[0396] 106. The anti-inflammatory composition of embodiment 102 or 103, wherein the striapathic region of the peptide differs from the sequence NFNFFFRFFF (SEQ ID NO: 33) by the deletion of one, two, or three amino acids.
[0397] 107. The anti-inflammatory composition of embodiment 106, wherein the deleted amino acids are located at the N-terminal end, the C-terminal end, or both ends of the sequence NFNFFFRFFF (SEQ ID NO: 33).
[0398] 108. An anti-inflammatory composition comprising a peptide, wherein the peptide is 3 to 24 amino acids residues in length and comprises a striapathic region having at least 70% identity with the sequence FFFRFFFNFN (RP118, SEQ ID NO: 44), wherein the peptide binds to the dimerization site on a NFkB Class II protein.
[0399] 109. The anti-inflammatory composition of embodiment 108, wherein the peptide also binds to human serum albumin.
[0400] 110. The anti-inflammatory composition of embodiment 108 or 109, wherein the differences between the striapathic region of the peptide and the sequence FFFRFFFNFN (SEQ ID NO: 44) are limited to conservative or highly conservative amino acid substitutions.
[0401] 111. The anti-inflammatory composition of embodiment 108 or 109, wherein the striapathic region of the peptide differs from the sequence FFFRFFFNFN (SEQ ID NO: 44) by substitution of one or more of the phenylalanine (F) residues with an amino acid residue selected from the group consisting of Trp (W), Tyr (Y), His (H), and Leu (L).
[0402] 112. The anti-inflammatory composition of embodiment 108 or 109, wherein the striapathic region of the peptide differs from the sequence FFFRFFFNFN (SEQ ID NO: 44) by the deletion of one, two, or three amino acids.
[0403] 113. The anti-inflammatory composition of embodiment 112, wherein the deleted amino acids are located at the N-terminal end, the C-terminal end, or both ends of the sequence FFFRFFFNFN (SEQ ID NO: 44).
[0404] 114. An anti-inflammatory composition comprising a peptide, wherein the peptide is 3 to 24 amino acids residues in length and comprises a striapathic region having at least 70% identity with the sequence FFRKFAKRFK (RP183, SEQ ID NO: 122), wherein the peptide binds to the dimerization site on a NFkB Class II protein.
[0405] 115. The anti-inflammatory composition of embodiment 114, wherein the peptide also binds to human serum albumin.
[0406] 116. The anti-inflammatory composition of embodiment 114 or 115, wherein the differences between the striapathic region of the peptide and the sequence FFRKFAKRFK (SEQ ID NO: 122) are limited to conservative or highly conservative amino acid substitutions.
[0407] 117. The anti-inflammatory composition of embodiment 114 or 115, wherein the striapathic region of the peptide differs from the sequence FFRKFAKRFK (SEQ ID NO: 122) by substitution of one or more of the phenylalanine (F) residues with an amino acid residue selected from the group consisting of Trp (W), Tyr (Y), and Leu (L).
[0408] 118. The anti-inflammatory composition of embodiment 114 or 115, wherein the striapathic region of the peptide differs from the sequence FFRKFAKRFK (SEQ ID NO: 122) by the deletion of one, two, or three amino acids.
[0409] 119. The anti-inflammatory composition of embodiment 118, wherein the deleted amino acids are located at the N-terminal end, the C-terminal end, or both ends of the sequence FFRKFAKRFK (SEQ ID NO: 122).
[0410] 120. An anti-inflammatory composition comprising a peptide, wherein the peptide is 3 to 24 amino acids residues in length and comprises a striapathic region having at least 70% identity with the sequence KFRKAFKRFF (RP182, SEQ ID NO: 121), wherein the peptide binds to the dimerization site on a NFkB Class II protein.
[0411] 121. The anti-inflammatory composition of embodiment 120, wherein the peptide also binds to human serum albumin.
[0412] 122. The anti-inflammatory composition of embodiment 120 or 121, wherein the differences between the striapathic region of the peptide and the sequence KFRKAFKRFF (SEQ ID NO: 121) are limited to conservative or highly conservative amino acid substitutions.
[0413] 123. The anti-inflammatory composition of embodiment 120 or 121, wherein the striapathic region of the peptide differs from the sequence KFRKAFKRFF (SEQ ID NO: 121) by substitution of one or more of the phenylalanine (F) residues with an amino acid residue selected from the group consisting of Trp (W), Tyr (Y), and Leu (L).
[0414] 124. The anti-inflammatory composition of embodiment 120 or 121, wherein the striapathic region of the peptide differs from the sequence KFRKAFKRFF (SEQ ID NO: 121) by the deletion of one, two, or three amino acids.
[0415] 125. The anti-inflammatory composition of embodiment 124, wherein the deleted amino acids are located at the N-terminal end, the C-terminal end, or both ends of the sequence KFRKAFKRFF (SEQ ID NO: 121).
[0416] 126. The anti-inflammatory composition of any one of embodiments 1 to 125, wherein the peptide binds to the dimerization site on Rel B (SEQ ID NO: 367) with a binding energy of at least -650 kcal/mol.
[0417] 127. The anti-inflammatory composition of any one of embodiments 1 to 126, wherein the peptide binds to the dimerization site on Rel B (SEQ ID NO: 367) and directly contacts at least one amino acid residue of Rel B selected from the group consisting of Glu 298, Tyr-300, Leu-301, Leu-302, Asp-330, His-332, and Leu-371.
[0418] 128. The anti-inflammatory composition of embodiment 127, wherein the peptide, when bound to the dimerization site on Rel B, forms an ionic bond with Asp-330, forms an ionic bond with His-332, and/or makes a hydrophobic contact with Leu-371.
[0419] 129. The anti-inflammatory composition of any one of embodiments 1 to 128, wherein the peptide binds to at least one signaling molecule selected from the group consisting of TGF.beta. (SEQ ID NO: 368), Notch1 (SEQ ID NO: 369), Wnt8R (SEQ ID NO: 370), TRAIL (SEQ ID NO: 371), IL6R (SEQ ID NO: 372), IL10R (SEQ ID NO: 373), EGFR (SEQ ID NO: 374), CDK6 (SEQ ID NO: 375), Histone Methyl Transferase (HMT) (SEQ ID NO: 376), CD47 (SEQ ID NO: 377), SIRP-.alpha. (SEQ ID NO: 378), CD206 (SEQ ID NO: 379), TGM2 (SEQ ID NO: 380); LEGUMAIN (SEQ ID NO: 137), CD209 (SEQ ID NO: 140), FAS (SEQ ID NO: 152), PD-1 (SEQ ID NO: 159), MKK7 (SEQ ID NO: 166), and RNR (SEQ ID NO: 168).
[0420] 130. The anti-inflammatory composition of embodiment 129, wherein the peptide binds to TGF.beta. (SEQ ID NO: 368) with a binding energy of at least -650 kcal/mol.
[0421] 131. The anti-inflammatory composition of embodiment 129 or 130, wherein the peptide binds to TGF.beta. (SEQ ID NO: 368) and directly contacts at least one amino acid residue of TGF.beta. selected from the group consisting of Leu-20, Ile-22, Phe-24, Asp-27, Leu-28, Trp-30, Trp-32, Tyr-39, Phe-43, Pro-80, Leu-83, Leu-101, and Ser-112.
[0422] 132. The anti-inflammatory composition of any one of embodiments 129 to 131, wherein the peptide binds to Notch1 (SEQ ID NO: 369) with a binding energy of at least -650 kcal/mol.
[0423] 133. The anti-inflammatory composition of any one of embodiments 120 to 123, wherein the peptide binds to Notch (SEQ ID NO: 369) and directly contacts at least one amino acid residue of Notch selected from the group consisting of Phe-1520, Gln-1523, Arg-1524, Glu-1526, Ala-1553, Glu-1556, Trp-1557, Cys-1562, His-1602, Arg-1684, Gln-1685, Cys-1686, Ser-1691, Cys-1693, Phe-1694, and Phe-1703.
[0424] 134. The anti-inflammatory composition of any one of embodiments 129 to 133, wherein the peptide binds to Wnt8R (SEQ ID NO: 370) with a binding energy of at least -600 kcal/mol.
[0425] 135. The anti-inflammatory composition of any one of embodiments 129 to 134, wherein the peptide binds to Wnt8R (SEQ ID NO: 370) and directly contacts at least one amino acid residue of Wnt8R selected from the group consisting of Tyr-52, Gln-56, Phe-57, Asn-58, Met-91, Tyr-100, Lys-101, Pro-103, Pro-105, Pro-106, Arg-137, and Asp-145.
[0426] 136. The anti-inflammatory composition of any one of embodiments 129 to 135, wherein the peptide binds to TRAIL (SEQ ID NO: 371) with a binding energy of at least -650 kcal/mol.
[0427] 137. The anti-inflammatory composition of any one of embodiments 120 to 127, wherein the peptide binds to TRAIL (SEQ ID NO: 371) and directly contacts at least one amino acid residue of TRAIL selected from the group consisting of Arg-130, Arg-158, Ser-159, Gly-160, His-161, Phe-163, Tyr-189, Arg-189, Gln-193, Glu-195, Glu-236, Tyr-237, Leu-239, Asp-267, Asp-269, His-270, and Glu-271.
[0428] 138. The anti-inflammatory composition of any one of embodiments 129 to 137, wherein the peptide binds to IL6R (SEQ ID NO: 372) with a binding energy of at least -600 kcal/mol.
[0429] 139. The anti-inflammatory composition of any one of embodiments 129 to 138, wherein the peptide binds to IL6R (SEQ ID NO: 372) and directly contacts at least one amino acid residue of IL6R selected from the group consisting of Glu-163, Gly-164, Phe-168, Gln-190, Phe-229, Tyr-230, Phe-279, and Gln-281.
[0430] 140. The anti-inflammatory composition of any one of embodiments 129 to 139, wherein the peptide binds to IL10R (SEQ ID NO: 373) with a binding energy of at least -600 kcal/mol.
[0431] 141. The anti-inflammatory composition of any one of embodiments 129 to 140, wherein the peptide binds to IL10R (SEQ ID NO: 373) and directly contacts at least one amino acid residue of IL10R selected from the group consisting of Tyr-43, Ile-45, Glu-46, Asp-61, Asn-73, Arg-76, Asn-94, Arg-96, Phe-143, Ala-189, Ser-190, and Ser-191.
[0432] 142. The anti-inflammatory composition of any one of embodiments 129 to 141, wherein the peptide binds to EGFR (SEQ ID NO: 374) with a binding energy of at least -650 kcal/mol.
[0433] 143. The anti-inflammatory composition of any one of embodiments 129 to 142, wherein the peptide binds to EGFR (SEQ ID NO: 374) and directly contacts at least one amino acid residue of EGFR selected from the group consisting of Leu-10, Thr-40, Trp-41, Leu-63, His-66, Asp-68, Leu-88, Tyr-101, Asp-48, and Phe-51.
[0434] 144. The anti-inflammatory composition of any one of embodiments 129 to 143, wherein the peptide binds to CDK6 (SEQ ID NO: 375) with a binding energy of at least -600 kcal/mol.
[0435] 145. The anti-inflammatory composition of any one of embodiments 129 to 144, wherein the peptide binds to CDK6 (SEQ ID NO: 375) and directly contacts at least one amino acid residue of CDK6 selected from the group consisting of Val-142, Arg-144, Asp-145, Ser-171, Val-180, Val-181, Leu-183, Arg-186, Val-190, Gln-193, Tyr-196, and Val-200.
[0436] 146. The anti-inflammatory composition of any one of embodiments 129 to 145, wherein the peptide binds to histone methyl transferase (HMT) (SEQ ID NO: 376) with a binding energy of at least -600 kcal/mol.
[0437] 147. The anti-inflammatory composition of any one of embodiments 129 to 146, wherein the peptide binds to HMT (SEQ ID NO: 376) and directly contacts at least one amino acid residue of HMT selected from the group consisting of Asn-69, His-70, Ser-71, Lys-72, Asp-73, Pro-74, and Asn-75.
[0438] 148. The anti-inflammatory composition of any one of embodiments 129 to 147, wherein the peptide binds to the SIRP-.alpha. binding site on CD47 (SEQ ID NO: 377) with a binding energy of at least -550 kcal/mol.
[0439] 149. The anti-inflammatory composition of any one of embodiments 129 to 148, wherein the peptide binds to CD47 (SEQ ID NO: 377) and directly contacts at least one amino acid residue of CD47 selected from the group consisting of Glu-29, Ala-30, Glu-35, Val-36, Tyr-37, Lys-39, Thr-49, Asp-51, Glu-97, Thr-99, Leu-101, Thr-102, Arp-103, Glu-104, and Glu-106.
[0440] 150. The anti-inflammatory composition of any one of embodiments 129 to 149, wherein the peptide binds to the CD47 binding site on SIRP-.alpha. (SEQ ID NO: 378) with a binding energy of at least -600 kcal/mol.
[0441] 151. The anti-inflammatory composition of any one of embodiments 129 to 150, wherein the peptide binds to SIRP-.alpha. (SEQ ID NO: 378) and directly contacts at least one amino acid residue of SIRP-.alpha. selected from the group consisting of Leu-30, Gln-37, Gln-52, Lys-53, Ser-66, Thr-67, Arg-69, Met-72, Phe-74, Lys-96, and Asp-100.
[0442] 152. The anti-inflammatory composition of any one of embodiments 129 to 151, wherein the peptide binds to CD206 (SEQ ID NO: 379) with a binding energy of at least -650 kcal/mol.
[0443] 153. The anti-inflammatory composition of any one of embodiments 129 to 152, wherein the peptide binds to CD206 (SEQ ID NO: 379) and directly contacts at least one amino acid residue of CD206 selected from the group consisting of Glu-725, Tyr-729, Glu-733, Asn-747, and Asp-748.
[0444] 154. The anti-inflammatory composition of any one of embodiments 129 to 153, wherein the peptide binds to TGM2 (SEQ ID NO: 380) with a binding energy of at least -650 kcal/mol.
[0445] 155. The anti-inflammatory composition of any one of embodiments 129 to 154, wherein the peptide binds to TGM2 (SEQ ID NO: 380) and directly contacts at least one amino acid residue of TGM2 selected from the group consisting of Cys-277, His-335, and Asp-358.
[0446] 156. The anti-inflammatory composition of any one of embodiments 129 to 155, wherein the peptide binds to LEGUMAIN (SEQ ID NO: 137) with a binding energy of at least -600 kcal/mol.
[0447] 157. The anti-inflammatory composition of any one of embodiments 129 to 156, wherein the peptide binds to LEGUMAIN (SEQ ID NO: 137) and directly contacts at least one amino acid residue of LEGUMAIN selected from the group consisting of Asn-44, Arg-46, His-159, Glu-189, Cys-191, Ser-217, Ser-218 and Asp-233.
[0448] 158. The anti-inflammatory composition of any one of embodiments 129 to 157, wherein the peptide binds to CD209 (SEQ ID NO: 140) with a binding energy of at least -600 kcal/mol.
[0449] 159. The anti-inflammatory composition of any one of embodiments 129 to 158, wherein the peptide binds to CD209 (SEQ ID NO: 140) and directly contacts at least one amino acid residue of CD209 selected from the group consisting of Phe-269, Glu-280, Glu-303, Asn-305, Asn-306, Glu-310, Asp-311, Ser-316, Gly-317, Asn-321 and Lys-324.
[0450] 160. The anti-inflammatory composition of any one of embodiments 129 to 159, wherein the peptide binds to FAS (SEQ ID NO: 152) with a binding energy of at least -600 kcal/mol.
[0451] 161. The anti-inflammatory composition of any one of embodiments 129 to 160, wherein the peptide binds to FAS (; SEQ ID NO: 152) and directly contacts at least one amino acid residue of FAS selected from the group consisting of Lys-251, Lys-296, Lys-299, Leu-303, Leu-306, Ala-307, Glu-308, Lys-309, Gln-311, Ile-314, Leu-315, Asp-317, Ile-318 and Thr-319.
[0452] 162. The anti-inflammatory composition of any one of embodiments 129 to 161, wherein the peptide binds to PD-1 (SEQ ID NO: 159) with a binding energy of at least -600 kcal/mol.
[0453] 163. The anti-inflammatory composition of any one of embodiments 129 to 162, wherein the peptide binds to PD-1 (SEQ ID NO: 159) and directly contacts at least one amino acid residue of PD-1 selected from the group consisting of Val-64, Asn-66, Tyr-68, Met-70, Thr-76, Lys-78, Thr-120, Leu-122, Ala-125, and Ser-127.
[0454] 164. The anti-inflammatory composition of any one of embodiments 129 to 163, wherein the peptide binds to MKK7 (SEQ ID NO: 166) with a binding energy of at least -600 kcal/mol.
[0455] 165. The anti-inflammatory composition of any one of embodiments 129 to 164, wherein the peptide binds to MKK7 (SEQ ID NO: 166) and directly contacts at least one amino acid residue of MKK7 selected from the group consisting of Met-142, Val-150, Lys-152, Lys-165, Met-212, Met-215, Thr-217, Lys-221, Leu-266, Cys-276 and Asp-277.
[0456] 166. The anti-inflammatory composition of any one of embodiments 129 to 165, wherein the peptide binds to RNR (SEQ ID NO: 168) with a binding energy of at least -600 kcal/mol.
[0457] 167. The anti-inflammatory composition of any one of embodiments 129 to 166, wherein the peptide binds to RNR (SEQ ID NO: 168) and directly contacts at least one amino acid residue of RNR selected from the group consisting of Asn-426, Leu-427, Cys-428, Glu-430, Met-606, Pro-608 and Ala-610.
[0458] 168. The anti-inflammatory composition of any one of embodiments 1 to 167, wherein the peptide binds to human serum albumin (HSA) (SEQ ID NO: 381) with a binding energy of at least -650 kcal/mol.
[0459] 169. The anti-inflammatory composition of any one of embodiments 1 to 168, wherein the peptide comprises a striapathic region that is composed exclusively of D-form amino acid residues.
[0460] 170. The anti-inflammatory composition of any one of embodiments 1 to 169, wherein the peptide is in solution at a concentration of about 0.1 mg/ml to about 100 mg/ml.
[0461] 171. The anti-inflammatory composition of any one of embodiments 1 to 170, wherein the composition contains about 1 mg to about 500 mg of the peptide.
[0462] 172. The anti-inflammatory composition of embodiment 158 or 171, wherein the composition is substantially free of protein other than the peptide.
[0463] 173. An anti-inflammatory composition comprising a first peptide as defined in any one of embodiments 1 to 171 in combination with a second peptide as defined in any one of embodiments 1 to 171, wherein the first and second peptides can have the same sequence or different sequences.
[0464] 174. The anti-inflammatory composition of embodiment 173, wherein the first and second peptides are linked together by a peptide bond, a peptide linker, or a non-peptide linker.
[0465] 175. The anti-inflammatory composition of embodiment 173, wherein the first and second peptides are linked together by a peptide linker, wherein the peptide linker has a sequence selected from the group consisting of Gly-Gly-Gly (GGG), Gly-Gly-Gly-Arg (GGGR; SEQ ID NO: 412), Gly-Pro-Gly (GPG), and Gly-Pro-Gly-Arg (GPGR; SEQ ID NO: 413).
[0466] 176. The anti-inflammatory composition of embodiment 174 or 175, wherein the linked first and second peptides bind to the dimerization site on Rel B (SEQ ID NO: 367) with a binding energy of at least -700 kcal/mol.
[0467] 177. The anti-inflammatory composition of any one of embodiments 1 to 171 and embodiments 173 to 176, further comprising serum albumin.
[0468] 178. The anti-inflammatory composition of embodiment 177, wherein the composition is substantially free of blood proteins other than serum albumin.
[0469] 179. A pharmaceutical composition comprising the anti-inflammatory composition of any one of embodiments 1 to 178, and a pharmaceutically acceptable carrier.
[0470] 180. The pharmaceutical composition of embodiment 179, wherein the composition comprises a chemotherapeutic agent.
[0471] 181. A method of treating a condition associated with chronic inflammation, the method comprising administering a composition according to any one of embodiments 1 to 180 to a subject suffering from the condition.
[0472] 182. The method of embodiment 181, wherein the condition is selected from the group consisting of irritable bowel disease, ulcerative colitis, colitis, Crohn's disease, idiopathic pulmonary fibrosis, asthma, keratitis, arthritis, osteoarthritis, rheumatoid arthritis, auto-immune diseases, a feline or human immunodeficiency virus (FIV or HIV) infection, and cancer.
[0473] 183. The method of embodiment 181 or 182, wherein the subject is a mammal.
[0474] 184. The method of any one of embodiments 181 to 183, wherein the subject is a human.
[0475] 185. The method of any one of embodiments 181 to 184, wherein the anti-inflammatory composition is administered in a dosage that includes between about 1 mg and about 500 mg of peptide.
[0476] 186. The method of any one of embodiments 181 to 185, wherein the anti-inflammatory composition is administered intravenously, intraperitoneally, parenteral, orthotopically, subcutaneously, topically, nasally, by means of an implantable depot, using nanoparticle-based delivery systems, microneedle patch, microspheres, beads, osmotic or mechanical pumps, and/or other mechanical means.
[0477] 187. The method of any one of embodiments 181 to 186, wherein the anti-inflammatory composition is administered in conjunction with another drug known to be effective in treating the condition.
[0478] 188. The method of embodiment 187, wherein the anti-inflammatory composition is administered prior to, at the same time as, or after the administration of the other drug.
[0479] 189. A method of treating fibrosis in a subject, the method comprising administering a composition according to any one of embodiments 1 to 180 to the subject.
[0480] 190. The method of embodiment 189, wherein the fibrosis is selected from the group consisting of pulmonary fibrosis, dermal fibrosis, hepatic fibrosis, renal fibrosis, and fibrosis caused by ionizing radiation.
[0481] 191. The method of embodiment 189 or 190, wherein the subject is a mammal.
[0482] 192. The method of any one of embodiments 189 to 191, wherein the subject is a human.
[0483] 193. The method of any one of embodiments 189 to 192, wherein the anti-inflammatory composition is administered in a dosage that includes between about 1 mg and about 500 mg of peptide.
[0484] 194. The method of any one of embodiments 189 to 193, wherein the anti-inflammatory composition is administered intravenously, intraperitoneally, parenteral, orthotopically, subcutaneously, topically, nasally, by means of an implantable depot, using nanoparticle-based delivery systems, microneedle patch, microspheres, beads, osmotic or mechanical pumps, and/or other mechanical means.
[0485] 195. The method of any one of embodiments 189 to 194, wherein the anti-inflammatory composition is administered in conjunction with another drug known to be effective in treating fibrosis.
[0486] 196. The method of embodiment 195, wherein the anti-inflammatory composition is administered prior to, at the same time as, or after the administration of the other drug.
[0487] 197. A method of reducing pro-inflammatory cytokine levels in a subject suffering from a chronic inflammatory condition, the method comprising administering a composition according to any one of embodiments 1 to 180 to the subject.
[0488] 198. The method of embodiment 197, wherein the chronic inflammatory condition is selected from the group consisting of irritable bowel disease, ulcerative colitis, colitis, Crohn's disease, idiopathic pulmonary fibrosis, asthma, keratitis, arthritis, osteoarthritis, rheumatoid arthritis, auto-immune diseases, a feline or human immunodeficiency virus (FIV or HIV) infection, and cancer.
[0489] 199. The method of embodiment 197 or 198, wherein the method reduces the level of at least one cytokine selected from group consisting of NF-kB, TNF.alpha., IL1, IL6, IL12, MMP-1, MMP-9, MCP-1, IL8, IL17, and IL23.
[0490] 200. The method of embodiment 199, wherein the level of the at least one cytokine is reduced by at least 10%.
[0491] 201. The method of any one of embodiments 197 to 200, wherein the subject is a mammal.
[0492] 202. The method of any one of embodiments 197 to 201, wherein the subject is a human.
[0493] 203. The method of any one of embodiments 197 to 202, wherein the anti-inflammatory composition is administered in a dosage that includes between about 1 mg and about 500 mg of peptide.
[0494] 204. The method of any one of embodiments 197 to 203, wherein the anti-inflammatory composition is administered intravenously, intraperitoneally, parenteral, orthotopically, subcutaneously, topically, nasally, by means of an implantable depot, using nanoparticle-based delivery systems, microneedle patch, microspheres, beads, osmotic or mechanical pumps, and/or other mechanical means.
[0495] 205. The method of any one of embodiments 197 to 204, wherein the anti-inflammatory composition is administered in conjunction with another drug known to be effective in treating the chronic inflammatory condition that the subject is suffering from.
[0496] 206. The method of embodiment 205, wherein the anti-inflammatory composition is administered prior to, at the same time as, or after the administration of the other drug.
[0497] 207. A method of treating cancer in a subject, the method comprising administering an anti-inflammatory composition according to any one of embodiments 1 to 180 to the subject.
[0498] 208. The method of embodiment 207, wherein the cancer is selected from the group consisting of colon cancer, and breast cancer.
[0499] 209. The method of embodiment 207 or 208, wherein the anti-inflammatory composition is administered in conjunction with a chemotherapeutic agent or cell therapy.
[0500] 210. The method of embodiment 209, wherein the chemotherapeutic agent or cell therapy is selected from the group consisting of steroids, anthracyclines, thyroid hormone replacement drugs, thymidylate-targeted drugs, checkpoint inhibitor drugs, Chimeric Antigen Receptor/T cell therapies, and other cell therapies.
[0501] 211. The method of embodiment 209, wherein the chemotherapeutic agent is selected from the group consisting of Gemcitabine, Docetaxel, Bleomycin, Erlotinib, Gefitinib, Lapatinib, Imatinib, Dasatinib, Nilotinib, Bosutinib, Crizotinib, Ceritinib, Trametinib, Bevacizumab, Sunitinib, Sorafenib, Trastuzumab, Ado-trastuzumab emtansine, Rituximab, Ipilimumab, Rapamycin, Temsirolimus, Everolimus, Methotrexate, Doxorubicin, Abraxane, Folfirinox, Cisplatin, Carboplatin, 5-fluorouracil, Teysumo, Paclitaxel, Prednisone, Levothyroxine, and Pemetrexed.
[0502] 212. The method of any one of embodiments 209 to 211, wherein the anti-inflammatory composition is administered prior to, at the same time as, or after the administration of the chemotherapeutic agent or cell therapy.
[0503] 213. The method of embodiment 207 or 208, wherein the anti-inflammatory composition is administered in conjunction with radiation therapy.
[0504] 214. The method of embodiment 213, wherein the anti-inflammatory composition is administered prior to, or after the administration of the radiation therapy.
[0505] 215. The method of any one of embodiments 207 to 214, wherein the subject is a mammal.
[0506] 216. The method of any one of embodiments 207 to 215, wherein the subject is a human.
[0507] 217. The method of any one of embodiments 207 to 216, wherein the anti-inflammatory composition is administered in a dosage that includes between about 1 mg and about 500 mg of peptide.
[0508] 218. The method of any one of embodiments 207 to 217, wherein the anti-inflammatory composition is administered intravenously, intraperitoneally, parenteral, orthotopically, subcutaneously, nasally, by means of an implantable depot, using nanoparticle-based delivery systems, microneedle patch, microspheres, beads, osmotic or mechanical pumps, and/or other mechanical means.
Sequence CWU 1
1
41317PRTArtificial SequenceSynthetic peptideMISC_FEATURE(4)..(4)Xaa
is selected from Arg (R), His (H), Lys (K), Glu (E), Gln (Q), Asn
(N), and Asp (D) 1Phe Phe Phe Xaa Phe Phe Phe1 527PRTArtificial
SequenceSynthetic peptideMISC_FEATURE(4)..(4)Xaa is selected from
Arg (R), His (H), Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D)
2Trp Trp Trp Xaa Trp Trp Trp1 537PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(4)..(4)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D) 3Tyr Tyr Tyr Xaa
Tyr Tyr Tyr1 547PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(4)..(4)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D) 4Leu Leu Leu Xaa
Leu Leu Leu1 557PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(4)..(4)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D) 5Cys Cys Cys Xaa
Cys Cys Cys1 567PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(4)..(4)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D) 6Met Met Met Xaa
Met Met Met1 577PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(4)..(4)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D) 7Val Val Val Xaa
Val Val Val1 587PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(4)..(4)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp (D) 8Ile Ile Ile Xaa
Ile Ile Ile1 598PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 9Phe Xaa Xaa Phe Phe
Xaa Xaa Phe1 5108PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 10Phe Xaa Xaa Phe Phe
Xaa Xaa Trp1 5118PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 11Trp Xaa Xaa Phe Phe
Xaa Xaa Phe1 5128PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 12Phe Xaa Xaa Phe Trp
Xaa Xaa Phe1 5138PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 13Phe Xaa Xaa Trp Phe
Xaa Xaa Phe1 5148PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 14Phe Xaa Xaa Trp Trp
Xaa Xaa Phe1 5158PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 15Trp Xaa Xaa Trp Trp
Xaa Xaa Phe1 5168PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 16Phe Xaa Xaa Trp Trp
Xaa Xaa Trp1 5178PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 17Trp Xaa Xaa Trp Trp
Xaa Xaa Trp1 5188PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 18Phe Xaa Xaa Phe Phe
Xaa Xaa Tyr1 5198PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 19Tyr Xaa Xaa Phe Phe
Xaa Xaa Phe1 5208PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 20Phe Xaa Xaa Phe Tyr
Xaa Xaa Phe1 5218PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 21Phe Xaa Xaa Tyr Phe
Xaa Xaa Phe1 5228PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 22Phe Xaa Xaa Tyr Tyr
Xaa Xaa Phe1 5238PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 23Tyr Xaa Xaa Tyr Tyr
Xaa Xaa Phe1 5248PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 24Phe Xaa Xaa Tyr Tyr
Xaa Xaa Tyr1 5258PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 25Tyr Xaa Xaa Tyr Tyr
Xaa Xaa Tyr1 5268PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 26Tyr Xaa Xaa Tyr Tyr
Xaa Xaa Trp1 5278PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 27Trp Xaa Xaa Tyr Tyr
Xaa Xaa Tyr1 5288PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 28Tyr Xaa Xaa Tyr Trp
Xaa Xaa Tyr1 5298PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 29Tyr Xaa Xaa Trp Tyr
Xaa Xaa Tyr1 5308PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 30Tyr Xaa Xaa Trp Trp
Xaa Xaa Tyr1 5318PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 31Trp Xaa Xaa Trp Trp
Xaa Xaa Tyr1 5328PRTArtificial SequenceSynthetic
peptideMISC_FEATURE(2)..(3)Xaa is selected from Arg (R), His (H),
Lys (K), Glu (E), Gln (Q), Asn (N), and Asp
(D)MISC_FEATURE(6)..(7)Xaa is selected from Arg (R), His (H), Lys
(K), Glu (E), Gln (Q), Asn (N), and Asp (D) 32Tyr Xaa Xaa Trp Trp
Xaa Xaa Trp1 53310PRTArtificial SequenceSynthetic peptide 33Asn Phe
Asn Phe Phe Phe Arg Phe Phe Phe1 5 103410PRTArtificial
SequenceSynthetic peptide 34Trp Trp Trp Arg Trp Trp Trp Glu Trp
Gln1 5 103510PRTArtificial SequenceSynthetic peptide 35Glu Phe Asn
Phe Phe Phe Arg Phe Phe Phe1 5 103610PRTArtificial
SequenceSynthetic peptide 36Asp Phe Glu Phe Phe Phe Arg Phe Phe
Phe1 5 103710PRTArtificial SequenceSynthetic peptide 37Gln Phe Glu
Phe Phe Phe Arg Phe Phe Phe1 5 103810PRTArtificial
SequenceSynthetic peptide 38Glu Phe Glu Phe Phe Phe Arg Phe Phe
Phe1 5 103910PRTArtificial SequenceSynthetic peptide 39Phe Phe Phe
Arg Phe Phe Phe Glu Phe Gln1 5 104010PRTArtificial
SequenceSynthetic peptide 40Phe Phe Phe Arg Phe Phe Phe Glu Phe
Glu1 5 104110PRTArtificial SequenceSynthetic peptide 41Phe Phe Phe
Arg Phe Phe Phe Glu Phe Asp1 5 104210PRTArtificial
SequenceSynthetic peptide 42Phe Phe Phe Arg Phe Phe Phe Asn Phe
Glu1 5 104310PRTArtificial SequenceSynthetic peptide 43Phe Phe Phe
Arg Phe Phe Phe Asp Phe Glu1 5 104410PRTArtificial
SequenceSynthetic peptide 44Phe Phe Phe Arg Phe Phe Phe Asn Phe
Asn1 5 104510PRTArtificial SequenceSynthetic peptide 45Phe Phe Phe
His Phe Phe Phe Glu Phe Gln1 5 104610PRTArtificial
SequenceSynthetic peptide 46Phe Phe Phe His Phe Phe Phe Asn Phe
Glu1 5 104710PRTArtificial SequenceSynthetic peptide 47Phe Phe Phe
His Phe Phe Phe Glu Phe Asn1 5 104810PRTArtificial
SequenceSynthetic peptide 48Glu Phe Asn Phe Phe Phe His Phe Phe
Phe1 5 104910PRTArtificial SequenceSynthetic peptide 49Phe Phe Phe
Arg Phe Phe Phe Glu Phe Asn1 5 105010PRTArtificial
SequenceSynthetic peptide 50Phe Phe Phe His Phe Phe Phe Glu Phe
Glu1 5 105110PRTArtificial SequenceSynthetic peptide 51Gln Phe Glu
Phe Phe Phe His Phe Phe Phe1 5 105210PRTArtificial
SequenceSynthetic peptide 52Phe Phe Phe His Phe Phe Phe Glu Phe
Asp1 5 105310PRTArtificial SequenceSynthetic peptide 53Phe Phe Phe
His Phe Phe Phe Asp Phe Glu1 5 105410PRTArtificial
SequenceSynthetic peptide 54Tyr Tyr Tyr Arg Tyr Tyr Tyr Glu Tyr
Gln1 5 105510PRTArtificial SequenceSynthetic peptide 55Asn Phe Glu
Phe Phe Phe His Phe Phe Phe1 5 105610PRTArtificial
SequenceSynthetic peptide 56Phe Phe Phe Lys Phe Phe Phe Lys Phe
Glu1 5 105710PRTArtificial SequenceSynthetic peptide 57Glu Phe Asp
Phe Phe Phe Arg Phe Phe Phe1 5 105810PRTArtificial
SequenceSynthetic peptide 58Phe Phe Phe His Phe Phe Phe Asp Phe
Asp1 5 105910PRTArtificial SequenceSynthetic peptide 59Phe Phe Phe
His Phe Phe Phe Asn Phe Asn1 5 106010PRTArtificial
SequenceSynthetic peptide 60Phe Phe Phe Arg Phe Phe Phe Asp Phe
Asp1 5 106110PRTArtificial SequenceSynthetic peptide 61Phe Phe Phe
Lys Phe Phe Phe Lys Phe Asn1 5 106210PRTArtificial
SequenceSynthetic peptide 62Phe Phe Phe Lys Phe Phe Phe Glu Phe
Glu1 5 106310PRTArtificial SequenceSynthetic peptide 63Phe Phe Phe
Glu Phe Phe Phe Lys Phe Glu1 5 106410PRTArtificial
SequenceSynthetic peptide 64Phe Phe Phe Gln Phe Phe Phe Gln Phe
Gln1 5 106510PRTArtificial SequenceSynthetic peptide 65Phe Phe Phe
Lys Phe Phe Phe Gln Phe Gln1 5 106610PRTArtificial
SequenceSynthetic peptide 66Phe Phe Phe Lys Phe Phe Phe Asn Phe
Asn1 5 106710PRTArtificial SequenceSynthetic peptide 67Phe Phe Phe
Asn Phe Phe Phe Asn Phe Asn1 5 106810PRTArtificial
SequenceSynthetic peptide 68Phe Phe Phe Lys Phe Phe Phe Glu Phe
Gln1 5 106910PRTArtificial SequenceSynthetic peptide 69Phe Phe Phe
Lys Phe Phe Phe Lys Phe Gln1 5 107010PRTArtificial
SequenceSynthetic peptide 70Phe Phe Phe Lys Phe Phe Phe Gln Phe
Lys1 5 107110PRTArtificial SequenceSynthetic peptide 71Phe Phe Phe
Lys Phe Phe Phe Lys Phe Asp1 5 107210PRTArtificial
SequenceSynthetic peptide 72Phe Phe Phe Lys Phe Phe Phe Asp Phe
Asp1 5 107310PRTArtificial SequenceSynthetic peptide 73Phe Phe Phe
Asn Phe Phe Phe Lys Phe Asn1 5 107410PRTArtificial
SequenceSynthetic peptide 74Phe Phe Phe Asp Phe Phe Phe Asp Phe
Asp1 5 107510PRTArtificial SequenceSynthetic peptide 75Phe Phe Phe
Lys Phe Phe Phe Glu Phe Lys1 5 107610PRTArtificial
SequenceSynthetic peptide 76Phe Phe Phe Lys Phe Phe Phe Asp Phe
Lys1 5 107710PRTArtificial SequenceSynthetic peptide 77Phe Phe Phe
Glu Phe Phe Phe Glu Phe Glu1 5 107810PRTArtificial
SequenceSynthetic peptide 78Phe Phe Phe Asp Phe Phe Phe Lys Phe
Asp1 5 107910PRTArtificial SequenceSynthetic peptide 79Phe Phe Phe
Lys Phe Phe Phe Lys Phe Lys1 5 108010PRTArtificial
SequenceSynthetic peptide 80Phe Phe Phe Glu Phe Phe Phe Lys Phe
Lys1 5 108110PRTArtificial SequenceSynthetic peptide 81Phe Phe Phe
Gln Phe Phe Phe Lys Phe Lys1 5 108210PRTArtificial
SequenceSynthetic peptide 82Phe Phe Phe Lys Phe Phe Phe Asn Phe
Lys1 5 108310PRTArtificial SequenceSynthetic peptide 83Phe Phe Phe
Asn Phe Phe Phe Lys Phe Lys1 5 108410PRTArtificial
SequenceSynthetic peptide 84Phe Phe Phe Gln Phe Phe Phe Lys Phe
Gln1 5 108510PRTArtificial SequenceSynthetic peptide 85Phe Phe Phe
Asp Phe Phe Phe Lys Phe Lys1 5 108610PRTArtificial
SequenceSynthetic peptide 86Leu Leu Leu Arg Leu Leu Leu Glu Leu
Gln1 5 108710PRTArtificial SequenceSynthetic peptide 87Phe Val Phe
Lys Phe Val Phe Lys Phe Val1 5 108810PRTArtificial
SequenceSynthetic peptide 88Cys Cys Cys Arg Cys Cys Cys Glu Cys
Gln1 5 108910PRTArtificial SequenceSynthetic peptide 89Met Met Met
Arg Met Met Met Glu Met Gln1 5 109010PRTArtificial
SequenceSynthetic peptide 90Val Val Val Arg Val Val Val Glu Val
Gln1 5 109110PRTArtificial SequenceSynthetic peptide 91Ile Ile Ile
Arg Ile Ile Ile Glu Ile Gln1 5 109210PRTArtificial
SequenceSynthetic peptide 92Gly Gly Gly Arg Gly Gly Gly Glu Gly
Gln1 5 109310PRTArtificial SequenceSynthetic peptide 93Pro Pro Pro
Arg Pro Pro Pro Glu Pro Gln1 5 109410PRTArtificial
SequenceSynthetic peptide 94Thr Thr Thr Arg Thr Thr Thr Glu Thr
Gln1 5 109510PRTArtificial SequenceSynthetic peptide 95Ala Ala Ala
Arg Ala Ala Ala Glu Ala Gln1 5 109610PRTArtificial
SequenceSynthetic peptide 96Ala Ala Ala Lys Ala Ala Ala Lys Ala
Ala1 5 109710PRTArtificial SequenceSynthetic peptide 97Ala Ala Ala
Glu Ala Ala Ala Glu Ala Glu1 5 109810PRTArtificial
SequenceSynthetic peptide 98Ser Ser Ser Arg Ser Ser Ser Glu Ser
Gln1 5 109910PRTArtificial SequenceSynthetic peptide 99His His His
Arg His His His Glu His Gln1 5 1010010PRTArtificial
SequenceSynthetic peptide 100Arg Arg Arg Arg Arg Arg Arg Glu Arg
Gln1 5 1010110PRTArtificial SequenceSynthetic
peptide 101Gln Gln Gln Arg Gln Gln Gln Glu Gln Gln1 5
1010210PRTArtificial SequenceSynthetic peptide 102Glu Glu Glu Arg
Glu Glu Glu Glu Glu Gln1 5 1010310PRTArtificial SequenceSynthetic
peptide 103Asn Asn Asn Arg Asn Asn Asn Glu Asn Gln1 5
1010410PRTArtificial SequenceSynthetic peptide 104Asp Asp Asp Arg
Asp Asp Asp Glu Asp Gln1 5 1010510PRTArtificial SequenceSynthetic
peptide 105Lys Lys Lys Arg Lys Lys Lys Glu Lys Gln1 5
1010610PRTArtificial SequenceSynthetic peptide 106Phe Phe Gln Lys
Phe Phe Lys Arg Trp Arg1 5 1010710PRTArtificial SequenceSynthetic
peptide 107Phe Phe Arg Lys Phe Phe Lys Arg Phe Arg1 5
1010810PRTArtificial SequenceSynthetic peptide 108Arg Phe Arg Lys
Phe Phe Lys Arg Phe Phe1 5 1010910PRTArtificial SequenceSynthetic
peptide 109Arg Phe Arg Lys Phe Phe Lys Gln Phe Phe1 5
1011010PRTArtificial SequenceSynthetic peptide 110Phe Phe Gln Lys
Phe Phe Lys Arg Phe Arg1 5 1011110PRTArtificial SequenceSynthetic
peptide 111Arg Trp Arg Lys Phe Phe Lys Gln Phe Phe1 5
1011210PRTArtificial SequenceSynthetic peptide 112Phe Phe Glu His
Phe Trp Lys Glu Phe Asn1 5 1011310PRTArtificial SequenceSynthetic
peptide 113Phe Phe Gln His Phe Trp Lys Gln Phe Asn1 5
1011410PRTArtificial SequenceSynthetic peptide 114Gln Phe Asn His
Phe Phe Lys Glu Phe Phe1 5 1011510PRTArtificial SequenceSynthetic
peptide 115Phe Phe Asp Lys Phe Phe His Asp Phe Gln1 5
1011610PRTArtificial SequenceSynthetic peptide 116Gln Phe Asp His
Phe Phe Lys Asp Phe Phe1 5 1011710PRTArtificial SequenceSynthetic
peptide 117Phe Phe Glu Lys Phe Phe His Asn Phe Gln1 5
1011810PRTArtificial SequenceSynthetic peptide 118Asn Phe Glu Lys
Trp Phe His Glu Phe Phe1 5 1011910PRTArtificial SequenceSynthetic
peptide 119Leu Phe Arg Arg Ala Phe Lys Gln Leu Asp1 5
1012010PRTArtificial SequenceSynthetic peptide 120Asn Phe Gln Lys
Trp Phe His Gln Phe Phe1 5 1012110PRTArtificial SequenceSynthetic
peptide 121Lys Phe Arg Lys Ala Phe Lys Arg Phe Phe1 5
1012210PRTArtificial SequenceSynthetic peptide 122Phe Phe Arg Lys
Phe Ala Lys Arg Phe Lys1 5 1012310PRTArtificial SequenceSynthetic
peptide 123Phe Phe Lys Lys Phe Phe Lys Lys Phe Lys1 5
1012410PRTArtificial SequenceSynthetic peptide 124Lys Phe Lys Lys
Phe Phe Lys Lys Phe Phe1 5 1012510PRTArtificial SequenceSynthetic
peptide 125Lys Ala Arg Lys Ala Phe Lys Arg Phe Phe1 5
1012610PRTArtificial SequenceSynthetic peptide 126Trp Val Lys Asp
Ala Met Gln His Leu Asp1 5 1012710PRTArtificial SequenceSynthetic
peptide 127Phe Phe Lys Lys Phe Ala Lys Lys Phe Lys1 5
1012810PRTArtificial SequenceSynthetic peptide 128Phe Ala Glu Lys
Phe Phe Lys Asn Phe Lys1 5 1012910PRTArtificial SequenceSynthetic
peptide 129Lys Phe Asn Lys Phe Phe Lys Glu Ala Phe1 5
1013010PRTArtificial SequenceSynthetic peptide 130Phe Ala Lys Gln
Phe Phe Asn Lys Phe Lys1 5 1013110PRTArtificial SequenceSynthetic
peptide 131Lys Phe Asn Lys Ala Phe Lys Gln Ala Phe1 5
101324PRTArtificial SequenceSynthetic peptide 132His Tyr Trp
His113310PRTArtificial SequenceSynthetic peptide 133Phe Ala Gln Lys
Phe Phe Lys Asp Phe Lys1 5 1013410PRTArtificial SequenceSynthetic
peptide 134Phe Ala Glu Glu Phe Ala Glu Glu Phe Glu1 5
1013510PRTArtificial SequenceSynthetic peptide 135Lys Phe Lys Lys
Phe Phe Lys Lys Ala Phe1 5 1013610PRTArtificial SequenceSynthetic
peptide 136Lys Phe Lys Asn Phe Phe Gln Lys Ala Phe1 5
10137433PRTHomo sapiens 137Met Val Trp Lys Val Ala Val Phe Leu Ser
Val Ala Leu Gly Ile Gly1 5 10 15Ala Ile Pro Ile Asp Asp Pro Glu Asp
Gly Gly Lys His Trp Val Val 20 25 30Ile Val Ala Gly Ser Asn Gly Trp
Tyr Asn Tyr Arg His Gln Ala Asp 35 40 45Ala Cys His Ala Tyr Gln Ile
Ile His Arg Asn Gly Ile Pro Asp Glu 50 55 60Gln Ile Val Val Met Met
Tyr Asp Asp Ile Ala Tyr Ser Glu Asp Asn65 70 75 80Pro Thr Pro Gly
Ile Val Ile Asn Arg Pro Asn Gly Thr Asp Val Tyr 85 90 95Gln Gly Val
Pro Lys Asp Tyr Thr Gly Glu Asp Val Thr Pro Gln Asn 100 105 110Phe
Leu Ala Val Leu Arg Gly Asp Ala Glu Ala Val Lys Gly Ile Gly 115 120
125Ser Gly Lys Val Leu Lys Ser Gly Pro Gln Asp His Val Phe Ile Tyr
130 135 140Phe Thr Asp His Gly Ser Thr Gly Ile Leu Val Phe Pro Asn
Glu Asp145 150 155 160Leu His Val Lys Asp Leu Asn Glu Thr Ile His
Tyr Met Tyr Lys His 165 170 175Lys Met Tyr Arg Lys Met Val Phe Tyr
Ile Glu Ala Cys Glu Ser Gly 180 185 190Ser Met Met Asn His Leu Pro
Asp Asn Ile Asn Val Tyr Ala Thr Thr 195 200 205Ala Ala Asn Pro Arg
Glu Ser Ser Tyr Ala Cys Tyr Tyr Asp Glu Lys 210 215 220Arg Ser Thr
Tyr Leu Gly Asp Trp Tyr Ser Val Asn Trp Met Glu Asp225 230 235
240Ser Asp Val Glu Asp Leu Thr Lys Glu Thr Leu His Lys Gln Tyr His
245 250 255Leu Val Lys Ser His Thr Asn Thr Ser His Val Met Gln Tyr
Gly Asn 260 265 270Lys Thr Ile Ser Thr Met Lys Val Met Gln Phe Gln
Gly Met Lys Arg 275 280 285Lys Ala Ser Ser Pro Val Pro Leu Pro Pro
Val Thr His Leu Asp Leu 290 295 300Thr Pro Ser Pro Asp Val Pro Leu
Thr Ile Met Lys Arg Lys Leu Met305 310 315 320Asn Thr Asn Asp Leu
Glu Glu Ser Arg Gln Leu Thr Glu Glu Ile Gln 325 330 335Arg His Leu
Asp Ala Arg His Leu Ile Glu Lys Ser Val Arg Lys Ile 340 345 350Val
Ser Leu Leu Ala Ala Ser Glu Ala Glu Val Glu Gln Leu Leu Ser 355 360
365Glu Arg Ala Pro Leu Thr Gly His Ser Cys Tyr Pro Glu Ala Leu Leu
370 375 380His Phe Arg Thr His Cys Phe Asn Trp His Ser Pro Thr Tyr
Glu Tyr385 390 395 400Ala Leu Arg His Leu Tyr Val Leu Val Asn Leu
Cys Glu Lys Pro Tyr 405 410 415Pro Leu His Arg Ile Lys Leu Ser Met
Asp His Val Cys Leu Gly His 420 425 430Tyr13810PRTArtificial
SequenceSynthetic peptide 138Phe Ala Lys Gln Phe Ala Asn Lys Phe
Lys1 5 1013910PRTArtificial SequenceSynthetic peptide 139Lys Phe
Lys Asn Ala Phe Gln Lys Ala Phe1 5 10140360PRTHomo sapiens 140Met
Ser Asp Ser Lys Glu Pro Arg Leu Gln Gln Leu Gly Leu Leu Val1 5 10
15Ser Lys Val Pro Ser Ser Ile Ser Gln Glu Gln Ser Arg Gln Asp Ala
20 25 30Ile Tyr Gln Asn Leu Thr Gln Leu Lys Ala Ala Val Gly Glu Leu
Ser 35 40 45Glu Lys Ser Lys Leu Gln Glu Ile Tyr Gln Glu Leu Thr Gln
Leu Lys 50 55 60Ala Ala Val Gly Glu Leu Pro Glu Lys Ser Lys Leu Gln
Glu Ile Tyr65 70 75 80Gln Glu Leu Thr Arg Leu Lys Ala Ala Val Gly
Glu Leu Pro Glu Lys 85 90 95Ser Lys Leu Gln Glu Ile Tyr Gln Glu Leu
Thr Trp Leu Lys Ala Ala 100 105 110Val Gly Glu Leu Pro Glu Lys Ser
Lys Met Gln Glu Ile Tyr Gln Glu 115 120 125Leu Thr Arg Leu Lys Ala
Ala Val Gly Glu Leu Pro Glu Lys Ser Lys 130 135 140Gln Gln Glu Ile
Tyr Gln Glu Leu Thr Arg Leu Lys Ala Ala Val Gly145 150 155 160Glu
Leu Pro Glu Lys Ser Lys Gln Gln Glu Ile Tyr Gln Glu Leu Thr 165 170
175Arg Leu Lys Ala Ala Val Gly Glu Leu Pro Glu Lys Ser Lys Gln Gln
180 185 190Glu Ile Tyr Gln Glu Leu Thr Gln Leu Lys Ala Ala Val Glu
Arg Leu 195 200 205Cys His Pro Cys Pro Trp Glu Trp Thr Phe Phe Gln
Gly Asn Cys Tyr 210 215 220Phe Met Ser Asn Ser Gln Arg Asn Trp His
Asp Ser Ile Thr Ala Cys225 230 235 240Lys Glu Val Gly Ala Gln Leu
Val Val Ile Lys Ser Ala Glu Glu Gln 245 250 255Asn Phe Leu Gln Leu
Gln Ser Ser Arg Ser Asn Arg Phe Thr Trp Met 260 265 270Gly Leu Ser
Asp Leu Asn Gln Glu Gly Thr Trp Gln Trp Val Asp Gly 275 280 285Ser
Pro Leu Leu Pro Ser Phe Lys Gln Tyr Trp Asn Arg Gly Glu Pro 290 295
300Asn Asn Val Gly Glu Glu Asp Cys Ala Glu Phe Ser Gly Asn Gly
Trp305 310 315 320Asn Asp Asp Lys Cys Asn Leu Ala Lys Phe Trp Ile
Cys Lys Lys Ser 325 330 335Ala Ala Ser Cys Ser Arg Asp Glu Glu Gln
Phe Leu Ser Pro Ala Pro 340 345 350Ala Thr Pro Asn Pro Pro Pro Ala
355 36014110PRTArtificial SequenceSynthetic peptide 141Phe Ala Lys
Lys Phe Phe Lys Lys Phe Lys1 5 1014210PRTArtificial
SequenceSynthetic peptide 142Lys Phe Lys Lys Ala Phe Lys Lys Phe
Phe1 5 1014310PRTArtificial SequenceSynthetic peptide 143Phe Ala
Glu Lys Phe Ala Glu Lys Phe Glu1 5 1014410PRTArtificial
SequenceSynthetic peptide 144Asp Leu His Gln Met Ala Asp Lys Val
Trp1 5 1014510PRTArtificial SequenceSynthetic peptide 145Lys Ala
Arg Lys Ala Ala Lys Arg Phe Phe1 5 1014610PRTArtificial
SequenceSynthetic peptide 146Phe Ala Lys Asn Phe Ala Lys Lys Phe
Lys1 5 1014710PRTArtificial SequenceSynthetic peptide 147Phe Ala
Glu Lys Phe Ala Lys Asn Phe Lys1 5 1014810PRTArtificial
SequenceSynthetic peptide 148Lys Phe Lys Lys Ala Phe Lys Lys Ala
Phe1 5 1014910PRTArtificial SequenceSynthetic peptide 149Phe Ala
Lys Asn Phe Ala Lys Asn Phe Lys1 5 1015010PRTArtificial
SequenceSynthetic peptide 150Phe Ala Lys Glu Phe Ala Lys Glu Phe
Glu1 5 1015110PRTArtificial SequenceSynthetic peptide 151Lys Phe
Asp Lys Ala Phe Lys Gln Ala Phe1 5 10152335PRTHomo sapiens 152Met
Leu Gly Ile Trp Thr Leu Leu Pro Leu Val Leu Thr Ser Val Ala1 5 10
15Arg Leu Ser Ser Lys Ser Val Asn Ala Gln Val Thr Asp Ile Asn Ser
20 25 30Lys Gly Leu Glu Leu Arg Lys Thr Val Thr Thr Val Glu Thr Gln
Asn 35 40 45Leu Glu Gly Leu His His Asp Gly Gln Phe Cys His Lys Pro
Cys Pro 50 55 60Pro Gly Glu Arg Lys Ala Arg Asp Cys Thr Val Asn Gly
Asp Glu Pro65 70 75 80Asp Cys Val Pro Cys Gln Glu Gly Lys Glu Tyr
Thr Asp Lys Ala His 85 90 95Phe Ser Ser Lys Cys Arg Arg Cys Arg Leu
Cys Asp Glu Gly His Gly 100 105 110Leu Glu Val Glu Ile Asn Cys Thr
Arg Thr Gln Asn Thr Lys Cys Arg 115 120 125Cys Lys Pro Asn Phe Phe
Cys Asn Ser Thr Val Cys Glu His Cys Asp 130 135 140Pro Cys Thr Lys
Cys Glu His Gly Ile Ile Lys Glu Cys Thr Leu Thr145 150 155 160Ser
Asn Thr Lys Cys Lys Glu Glu Gly Ser Arg Ser Asn Leu Gly Trp 165 170
175Leu Cys Leu Leu Leu Leu Pro Ile Pro Leu Ile Val Trp Val Lys Arg
180 185 190Lys Glu Val Gln Lys Thr Cys Arg Lys His Arg Lys Glu Asn
Gln Gly 195 200 205Ser His Glu Ser Pro Thr Leu Asn Pro Glu Thr Val
Ala Ile Asn Leu 210 215 220Ser Asp Val Asp Leu Ser Lys Tyr Ile Thr
Thr Ile Ala Gly Val Met225 230 235 240Thr Leu Ser Gln Val Lys Gly
Phe Val Arg Lys Asn Gly Val Asn Glu 245 250 255Ala Lys Ile Asp Glu
Ile Lys Asn Asp Asn Val Gln Asp Thr Ala Glu 260 265 270Gln Lys Val
Gln Leu Leu Arg Asn Trp His Gln Leu His Gly Lys Lys 275 280 285Glu
Ala Tyr Asp Thr Leu Ile Lys Asp Leu Lys Lys Ala Asn Leu Cys 290 295
300Thr Leu Ala Glu Lys Ile Gln Thr Ile Ile Leu Lys Asp Ile Thr
Ser305 310 315 320Asp Ser Glu Asn Ser Asn Phe Arg Asn Glu Ile Gln
Ser Leu Val 325 330 33515310PRTArtificial SequenceSynthetic peptide
153Phe Ala Glu Lys Phe Ala Lys Lys Phe Lys1 5 1015410PRTArtificial
SequenceSynthetic peptide 154Phe Ala Glu Lys Phe Ala Glu Lys Phe
Lys1 5 1015510PRTArtificial SequenceSynthetic peptide 155Phe Ala
Lys Lys Phe Ala Lys Lys Phe Lys1 5 1015610PRTArtificial
SequenceSynthetic peptide 156Phe Ala Lys Asn Phe Ala Lys Asn Phe
Asn1 5 1015710PRTArtificial SequenceSynthetic peptide 157Phe Ala
Gln Lys Phe Ala Lys Asn Phe Lys1 5 1015810PRTArtificial
SequenceSynthetic peptide 158Phe Ala Asn Asn Phe Ala Asn Asn Phe
Asn1 5 10159198PRTHomo sapiens 159Met Gln Ile Pro Gln Ala Pro Trp
Pro Val Val Trp Ala Val Leu Gln1 5 10 15Leu Gly Trp Arg Pro Gly Trp
Phe Leu Asp Ser Pro Asp Arg Pro Trp 20 25 30Asn Pro Pro Thr Phe Ser
Pro Ala Leu Leu Val Val Thr Glu Gly Asp 35 40 45Asn Ala Thr Phe Thr
Cys Ser Phe Ser Asn Thr Ser Glu Ser Phe Val 50 55 60Leu Asn Trp Tyr
Arg Met Ser Pro Ser Asn Gln Thr Asp Lys Leu Ala65 70 75 80Ala Phe
Pro Glu Asp Arg Ser Gln Pro Gly Gln Asp Cys Arg Phe Arg 85 90 95Val
Thr Gln Leu Pro Asn Gly Arg Asp Phe His Met Ser Val Val Arg 100 105
110Ala Arg Arg Asn Asp Ser Gly Thr Tyr Leu Cys Gly Ala Ile Ser Leu
115 120 125Ala Pro Lys Ala Gln Ile Lys Glu Ser Leu Arg Ala Glu Leu
Arg Val 130 135 140Thr Glu Arg Arg Ala Glu Val Pro Thr Ala His Pro
Ser Pro Ser Pro145 150 155 160Arg Pro Ala Gly Gln Phe Gln Thr Leu
Val Val Gly Val Val Gly Gly 165 170 175Leu Leu Gly Ser Leu Val Leu
Leu Val Trp Val Leu Ala Val Ile Cys 180 185 190Ser Arg Ala Ala Arg
Gly 19516010PRTArtificial SequenceSynthetic peptide 160Phe Ala Asn
Lys Phe Ala Asn Lys Phe Asn1 5 1016110PRTArtificial
SequenceSynthetic peptide 161Phe Ala Asn Lys Phe Ala Lys Lys Phe
Lys1 5 1016210PRTArtificial SequenceSynthetic peptide 162Phe Ala
Gln Lys Phe Ala Lys Asp Phe Lys1 5 1016310PRTArtificial
SequenceSynthetic peptide 163Phe Ala Lys Glu Phe Ala Lys Glu Phe
Lys1 5 1016410PRTArtificial SequenceSynthetic peptide 164Phe Ala
Asn Lys Phe Ala Asn Lys Phe Lys1 5 1016510PRTArtificial
SequenceSynthetic peptide 165Lys Phe Asp Lys Phe Phe Lys Gln Ala
Phe1 5 10166435PRTHomo sapiens 166Met Ala Ala Ser Ser Leu Glu Gln
Lys Leu Ser Arg Leu Glu Ala Lys1 5 10 15Leu Lys Gln Glu Asn Arg Glu
Ala Arg Arg Arg Ile Asp Leu Asn Leu 20 25 30Asp Ile Ser Pro Gln Arg
Pro Arg Pro Ile Ile Val Ile Thr Leu Ser 35 40 45Pro Ala Pro Ala Pro
Ser Gln Arg Ala Ala Leu Gln Leu Pro Leu Ala 50 55 60Asn Asp Gly Gly
Ser Arg Ser Pro Ser Ser Glu Ser Ser Pro Gln His65 70 75 80Pro Thr
Pro Pro Ala Arg Pro Arg His Met Leu
Gly Leu Pro Ser Thr 85 90 95Leu Phe Thr Pro Arg Ser Met Glu Ser Ile
Glu Ile Asp Gln Lys Leu 100 105 110Gln Glu Ile Met Lys Gln Thr Gly
Tyr Leu Thr Ile Gly Gly Gln Arg 115 120 125Tyr Gln Ala Glu Ile Asn
Asp Leu Glu Asn Leu Gly Glu Met Gly Ser 130 135 140Gly Thr Cys Gly
Gln Val Trp Lys Met Arg Phe Arg Lys Thr Gly His145 150 155 160Val
Ile Ala Val Lys Gln Met Arg Arg Ser Gly Asn Lys Glu Glu Asn 165 170
175Lys Arg Ile Leu Met Asp Leu Asp Val Val Leu Lys Ser His Asp Cys
180 185 190Pro Tyr Ile Val Gln Cys Phe Gly Thr Phe Ile Thr Asn Thr
Asp Val 195 200 205Phe Ile Ala Met Glu Leu Met Gly Thr Cys Ala Glu
Lys Leu Lys Lys 210 215 220Arg Met Gln Gly Pro Ile Pro Glu Arg Ile
Leu Gly Lys Met Thr Val225 230 235 240Ala Ile Val Lys Ala Leu Tyr
Tyr Leu Lys Glu Lys His Gly Val Ile 245 250 255His Arg Asp Val Lys
Pro Ser Asn Ile Leu Leu Asp Glu Arg Gly Gln 260 265 270Ile Lys Leu
Cys Asp Phe Gly Ile Ser Gly Arg Leu Val Asp Ser Lys 275 280 285Ala
Lys Thr Arg Ser Ala Gly Cys Ala Ala Tyr Met Ala Pro Glu Arg 290 295
300Ile Asp Pro Pro Asp Pro Thr Lys Pro Asp Tyr Asp Ile Arg Ala
Asp305 310 315 320Val Trp Ser Leu Gly Ile Ser Leu Val Glu Leu Ala
Thr Gly Gln Phe 325 330 335Pro Tyr Lys Asn Cys Lys Thr Asp Phe Glu
Val Leu Thr Lys Val Leu 340 345 350Gln Glu Glu Pro Pro Leu Leu Pro
Gly His Met Gly Phe Ser Gly Asp 355 360 365Phe Gln Ser Phe Val Lys
Asp Cys Leu Thr Lys Asp His Arg Lys Arg 370 375 380Pro Lys Tyr Asn
Lys Leu Leu Glu His Ser Phe Ile Lys Arg Tyr Glu385 390 395 400Thr
Leu Glu Val Asp Val Ala Ser Trp Phe Lys Asp Val Met Ala Lys 405 410
415Thr Glu Ser Pro Arg Thr Ser Gly Val Leu Ser Gln Pro His Leu Pro
420 425 430Phe Phe Arg 43516710PRTArtificial SequenceSynthetic
peptide 167Lys Phe Asn Lys Ala Phe Lys Glu Ala Phe1 5
10168888PRTSaccharomyces cerevisiae 168Met Tyr Val Tyr Lys Arg Asp
Gly Arg Lys Glu Pro Val Gln Phe Asp1 5 10 15Lys Ile Thr Ala Arg Ile
Ser Arg Leu Cys Tyr Gly Leu Asp Pro Lys 20 25 30His Ile Asp Ala Val
Lys Val Thr Gln Arg Ile Ile Ser Gly Val Tyr 35 40 45Glu Gly Val Thr
Thr Ile Glu Leu Asp Asn Leu Ala Ala Glu Thr Cys 50 55 60Ala Tyr Met
Thr Thr Val His Pro Asp Tyr Ala Thr Leu Ala Ala Arg65 70 75 80Ile
Ala Ile Ser Asn Leu His Lys Gln Thr Thr Lys Gln Phe Ser Lys 85 90
95Val Val Glu Asp Leu Tyr Arg Tyr Val Asn Ala Ala Thr Gly Lys Pro
100 105 110Ala Pro Met Ile Ser Asp Asp Val Tyr Asn Ile Val Met Glu
Asn Lys 115 120 125Asp Lys Leu Asn Ser Ala Ile Val Tyr Asp Arg Asp
Phe Gln Tyr Ser 130 135 140Tyr Phe Gly Phe Lys Thr Leu Glu Arg Ser
Tyr Leu Leu Arg Ile Asn145 150 155 160Gly Gln Val Ala Glu Arg Pro
Gln His Leu Ile Met Arg Val Ala Leu 165 170 175Gly Ile His Gly Arg
Asp Ile Glu Ala Ala Leu Glu Thr Tyr Asn Leu 180 185 190Met Ser Leu
Lys Tyr Tyr Thr His Ala Ser Pro Thr Leu Phe Asn Ala 195 200 205Gly
Thr Pro Lys Pro Gln Met Ser Ser Cys Phe Leu Val Ala Met Lys 210 215
220Glu Asp Ser Ile Glu Gly Ile Tyr Asp Thr Leu Lys Glu Cys Ala
Leu225 230 235 240Ile Ser Lys Thr Ala Gly Gly Ile Gly Leu His Ile
His Asn Ile Arg 245 250 255Ser Thr Gly Ser Tyr Ile Ala Gly Thr Asn
Gly Thr Ser Asn Gly Leu 260 265 270Ile Pro Met Ile Arg Val Phe Asn
Asn Thr Ala Arg Tyr Val Asp Gln 275 280 285Gly Gly Asn Lys Arg Pro
Gly Ala Phe Ala Leu Tyr Leu Glu Pro Trp 290 295 300His Ala Asp Ile
Phe Asp Phe Ile Asp Ile Arg Lys Asn His Gly Lys305 310 315 320Glu
Glu Ile Arg Ala Arg Asp Leu Phe Pro Ala Leu Trp Ile Pro Asp 325 330
335Leu Phe Met Lys Arg Val Glu Glu Asn Gly Thr Trp Thr Leu Phe Ser
340 345 350Pro Thr Ser Ala Pro Gly Leu Ser Asp Cys Tyr Gly Asp Glu
Phe Glu 355 360 365Ala Leu Tyr Thr Arg Tyr Glu Lys Glu Gly Arg Gly
Lys Thr Ile Lys 370 375 380Ala Gln Lys Leu Trp Tyr Ser Ile Leu Glu
Ala Gln Thr Glu Thr Gly385 390 395 400Thr Pro Phe Val Val Tyr Lys
Asp Ala Cys Asn Arg Lys Ser Asn Gln 405 410 415Lys Asn Leu Gly Val
Ile Lys Ser Ser Asn Leu Cys Cys Glu Ile Val 420 425 430Glu Tyr Ser
Ala Pro Asp Glu Thr Ala Val Cys Asn Leu Ala Ser Val 435 440 445Ala
Leu Pro Ala Phe Ile Glu Thr Ser Glu Asp Gly Lys Thr Ser Thr 450 455
460Tyr Asn Phe Lys Lys Leu His Glu Ile Ala Lys Val Val Thr Arg
Asn465 470 475 480Leu Asn Arg Val Ile Asp Arg Asn Tyr Tyr Pro Val
Glu Glu Ala Arg 485 490 495Lys Ser Asn Met Arg His Arg Pro Ile Ala
Leu Gly Val Gln Gly Leu 500 505 510Ala Asp Thr Phe Met Leu Leu Arg
Leu Pro Phe Asp Ser Glu Glu Ala 515 520 525Arg Leu Leu Asn Ile Gln
Ile Phe Glu Thr Ile Tyr His Ala Ser Met 530 535 540Glu Ala Ser Cys
Glu Leu Ala Gln Lys Asp Gly Pro Tyr Glu Thr Phe545 550 555 560Gln
Gly Ser Pro Ala Ser Gln Gly Ile Leu Gln Phe Asp Met Trp Asp 565 570
575Gln Lys Pro Tyr Gly Met Trp Asp Trp Asp Thr Leu Arg Lys Asp Ile
580 585 590Met Lys His Gly Ile Arg Asn Ser Leu Thr Met Ala Pro Met
Pro Thr 595 600 605Ala Ser Thr Ser Gln Ile Leu Gly Tyr Asn Glu Cys
Phe Glu Pro Val 610 615 620Thr Ser Asn Met Tyr Ser Arg Arg Val Leu
Ser Gly Glu Phe Gln Val625 630 635 640Val Asn Pro Tyr Leu Leu Arg
Asp Leu Val Asp Leu Gly Ile Trp Asp 645 650 655Glu Gly Met Lys Gln
Tyr Leu Ile Thr Gln Asn Gly Ser Ile Gln Gly 660 665 670Leu Pro Asn
Val Pro Gln Glu Leu Lys Asp Leu Tyr Lys Thr Val Trp 675 680 685Glu
Ile Ser Gln Lys Thr Ile Ile Asn Met Ala Ala Asp Arg Ser Val 690 695
700Tyr Ile Asp Gln Ser His Ser Leu Asn Leu Phe Leu Arg Ala Pro
Thr705 710 715 720Met Gly Lys Leu Thr Ser Met His Phe Tyr Gly Trp
Lys Lys Gly Leu 725 730 735Lys Thr Gly Met Tyr Tyr Leu Arg Thr Gln
Ala Ala Ser Ala Ala Ile 740 745 750Gln Phe Thr Ile Asp Gln Lys Ile
Ala Asp Gln Ala Thr Glu Asn Val 755 760 765Ala Asp Ile Ser Asn Leu
Lys Arg Pro Ser Tyr Met Pro Ser Ser Ala 770 775 780Ser Tyr Ala Ala
Ser Asp Phe Val Pro Ala Ala Val Thr Ala Asn Ala785 790 795 800Thr
Ile Pro Ser Leu Asp Ser Ser Ser Glu Ala Ser Arg Glu Ala Ser 805 810
815Pro Ala Pro Thr Gly Ser His Ser Leu Thr Lys Gly Met Ala Glu Leu
820 825 830Asn Val Gln Glu Ser Lys Val Glu Val Pro Glu Val Pro Ala
Pro Thr 835 840 845Lys Asn Glu Glu Lys Ala Ala Pro Ile Val Asp Asp
Glu Glu Thr Glu 850 855 860Phe Asp Ile Tyr Asn Ser Lys Val Ile Ala
Cys Ala Ile Asp Asn Pro865 870 875 880Glu Ala Cys Glu Met Cys Ser
Gly 88516910PRTArtificial SequenceSynthetic peptide 169Phe Ala Lys
Glu Phe Ala Lys Lys Phe Lys1 5 1017010PRTArtificial
SequenceSynthetic peptide 170Lys Ala Arg Lys Ala Ala Lys Arg Ala
Phe1 5 1017110PRTArtificial SequenceSynthetic peptide 171Lys Ala
Arg Lys Ala Ala Lys Arg Ala Ala1 5 1017210PRTArtificial
SequenceSynthetic peptide 172Ala Ala Glu Glu Ala Ala Glu Glu Ala
Glu1 5 1017310PRTArtificial SequenceSynthetic peptide 173Ala Ala
Lys Lys Ala Ala Lys Lys Ala Lys1 5 1017410PRTArtificial
SequenceSynthetic peptide 174Phe Val Lys Phe Val Lys Phe Val Lys
Phe1 5 1017510PRTArtificial SequenceSynthetic peptide 175Lys Arg
Lys Ala Phe Arg Lys Phe Phe Phe1 5 101769PRTArtificial
SequenceSynthetic peptide 176Leu His Lys Met Tyr Asn Gln Val Trp1
51779PRTArtificial SequenceSynthetic peptide 177Trp Val Gln Asn Tyr
Met Lys His Leu1 517810PRTArtificial SequenceSynthetic peptide
178Arg Leu Val Glu Met Met Arg Gln Ile Trp1 5 101799PRTArtificial
SequenceSynthetic peptide 179Phe Leu Lys Arg Leu Leu Gln Glu Ile1
51808PRTArtificial SequenceSynthetic peptide 180Leu Arg Leu Leu His
Arg Leu Leu1 51819PRTArtificial SequenceSynthetic peptide 181Trp
Val Arg Asp Ser Met Lys His Leu1 518210PRTArtificial
SequenceSynthetic peptide 182Lys Phe Phe Arg Lys Lys Phe Arg Phe
Ala1 5 1018310PRTArtificial SequenceSynthetic peptide 183Trp Val
Gln Arg Val Val Glu Lys Phe Leu1 5 1018410PRTArtificial
SequenceSynthetic peptide 184Ala Phe Phe Arg Arg Phe Lys Phe Lys
Lys1 5 1018510PRTArtificial SequenceSynthetic peptide 185Leu Phe
Lys Glu Val Val Arg Gln Val Trp1 5 1018610PRTArtificial
SequenceSynthetic peptide 186Met Asp Lys Ile Tyr Asp Gln Val Trp
Lys1 5 1018710PRTArtificial SequenceSynthetic peptide 187Phe Val
Lys Lys Phe Val Lys Lys Phe Val1 5 1018810PRTArtificial
SequenceSynthetic peptide 188Lys Lys Phe Lys Phe Arg Arg Phe Phe
Ala1 5 1018910PRTArtificial SequenceSynthetic peptide 189Trp Val
Arg Asp Val Val Arg Ser Met Asp1 5 1019010PRTArtificial
SequenceSynthetic peptide 190Glu Leu Ser Asn Ile Tyr Glu Arg Val
Trp1 5 1019110PRTArtificial SequenceSynthetic peptide 191Trp Ile
Gln Arg Met Met Glu Val Leu Arg1 5 1019210PRTArtificial
SequenceSynthetic peptide 192Phe Phe Phe Lys Arg Phe Ala Lys Arg
Lys1 5 101939PRTArtificial SequenceSynthetic peptide 193Leu His Lys
Met Ser Asp Arg Val Trp1 51949PRTArtificial SequenceSynthetic
peptide 194Phe Ala Lys Lys Phe Ala Lys Lys Phe1 519510PRTArtificial
SequenceSynthetic peptide 195Trp Val Arg Glu Tyr Ile Asn Ser Leu
Glu1 5 1019610PRTArtificial SequenceSynthetic peptide 196Phe Phe
Lys Lys Arg Phe Ala Phe Arg Lys1 5 1019710PRTArtificial
SequenceSynthetic peptide 197Lys Trp Val Gln Asp Tyr Ile Lys Asp
Met1 5 1019810PRTArtificial SequenceSynthetic peptide 198Ala Phe
Arg Phe Lys Lys Arg Phe Phe Lys1 5 101998PRTArtificial
SequenceSynthetic peptide 199Leu Leu Arg His Leu Leu Arg Leu1
520010PRTArtificial SequenceSynthetic peptide 200Trp Ile Lys Lys
Leu Leu Glu Ser Ser Gln1 5 1020110PRTArtificial SequenceSynthetic
peptide 201Asp Met Ser Arg Val Val Asp Arg Val Trp1 5
1020210PRTArtificial SequenceSynthetic peptide 202Phe Glu Glu Glu
Phe Glu Glu Glu Phe Glu1 5 102039PRTArtificial SequenceSynthetic
peptide 203Trp Val Lys Asn Ser Ile Asn His Leu1 52049PRTArtificial
SequenceSynthetic peptide 204Leu Thr Lys Lys Gly Arg Arg Phe Cys1
52059PRTArtificial SequenceSynthetic peptide 205Ile Glu Gln Leu Leu
Arg Lys Leu Phe1 52069PRTArtificial SequenceSynthetic peptide
206Leu His Asn Ile Ser Asn Lys Val Trp1 52079PRTArtificial
SequenceSynthetic peptide 207Cys Phe Arg Arg Gly Lys Lys Thr Leu1
520810PRTArtificial SequenceSynthetic peptide 208Ile Val Arg Arg
Ala Asp Arg Ala Ala Val1 5 102099PRTArtificial SequenceSynthetic
peptide 209Thr Val Glu Arg Phe Lys Asn Leu Ser1 521010PRTArtificial
SequenceSynthetic peptide 210Gln Ser Ser Glu Leu Leu Lys Lys Ile
Trp1 5 102119PRTArtificial SequenceSynthetic peptide 211Ser Leu Asn
Lys Phe Arg Glu Val Thr1 521210PRTArtificial SequenceSynthetic
peptide 212Leu Ile Lys Gln Ile Val Lys Lys Leu Phe1 5
102136PRTArtificial SequenceSynthetic peptide 213Lys Lys Lys Phe
Phe Phe1 521410PRTArtificial SequenceSynthetic peptide 214Leu Tyr
Lys Lys Ile Ile Lys Lys Leu Leu1 5 1021510PRTArtificial
SequenceSynthetic peptide 215Phe Lys Lys Lys Phe Lys Lys Lys Phe
Lys1 5 1021610PRTArtificial SequenceSynthetic peptide 216Val Ala
Ala Arg Asp Ala Arg Arg Val Ile1 5 1021710PRTArtificial
SequenceSynthetic peptide 217Phe Leu Lys Lys Val Ile Gln Lys Ile
Leu1 5 1021810PRTArtificial SequenceSynthetic peptide 218Leu Ile
Lys Glu Ile Ile Lys Gln Val Met1 5 1021910PRTArtificial
SequenceSynthetic peptide 219Leu Leu Lys Lys Ile Ile Lys Lys Tyr
Leu1 5 1022010PRTArtificial SequenceSynthetic peptide 220Ala Phe
Phe Glu Glu Glu Ala Glu Phe Glu1 5 102219PRTArtificial
SequenceSynthetic peptide 221Lys Lys Trp Val Gln Asp Ser Met Lys1
522210PRTArtificial SequenceSynthetic peptide 222Asn Phe Ala Asn
Lys Val Gln Glu Val Ala1 5 1022310PRTArtificial SequenceSynthetic
peptide 223Ala Val Glu Gln Val Lys Asn Ala Phe Asn1 5
1022410PRTArtificial SequenceSynthetic peptide 224Met Val Gln Lys
Ile Ile Glu Lys Ile Leu1 5 102259PRTArtificial SequenceSynthetic
peptide 225Lys Met Ser Asp Gln Val Trp Lys Lys1 52269PRTArtificial
SequenceSynthetic peptide 226Met Val Lys Lys Ile Ile Glu Lys Met1
522710PRTArtificial SequenceSynthetic peptide 227Ala Leu Lys Lys
Gln Val Ile Lys Lys Ile1 5 1022810PRTArtificial SequenceSynthetic
peptide 228Ile Lys Lys Ile Val Gln Lys Lys Leu Ala1 5
1022910PRTArtificial SequenceSynthetic peptide 229Ala Phe Phe Lys
Lys Lys Ala Lys Phe Lys1 5 102308PRTArtificial SequenceSynthetic
peptide 230Met Lys Glu Ile Ile Lys Val Met1 523110PRTArtificial
SequenceSynthetic peptide 231Ala Glu Glu Glu Ala Glu Glu Glu Ala
Glu1 5 1023210PRTArtificial SequenceSynthetic peptide 232Ala Lys
Lys Lys Ala Lys Lys Lys Ala Lys1 5 102336PRTArtificial
SequenceSynthetic peptide 233Lys Lys Lys Ala Ala Ala1
523410PRTArtificial SequenceSynthetic peptide 234Lys Arg Lys Lys
Arg Phe Ala Phe Phe Phe1 5 1023510PRTArtificial SequenceSynthetic
peptide 235Arg Lys Arg Lys Phe Phe Ala Phe Phe Lys1 5
1023610PRTArtificial SequenceSynthetic peptide 236Phe Phe Phe Ala
Phe Arg Lys Lys Arg Lys1 5 1023710PRTArtificial SequenceSynthetic
peptide 237Phe Arg Lys Lys Arg Phe Ala Phe Phe Lys1 5
1023810PRTArtificial SequenceSynthetic peptide 238Phe Phe Phe Arg
Arg Lys Lys Lys Phe Ala1 5 1023910PRTArtificial SequenceSynthetic
peptide 239Lys Phe Phe Ala Phe Arg Lys Lys Arg Phe1 5
1024010PRTArtificial SequenceSynthetic peptide 240Lys Phe Phe Ala
Phe Phe Lys Arg Lys Arg1 5 1024110PRTArtificial SequenceSynthetic
peptide 241Lys Lys Lys Lys Lys Phe Phe Phe Phe Phe1 5
1024210PRTArtificial SequenceSynthetic peptide 242Ala Phe Lys Lys
Lys Arg Arg Phe Phe Phe1 5 1024310PRTArtificial
SequenceSynthetic peptide 243Lys Arg Lys Lys Arg Ala Ala Phe Phe
Phe1 5 1024410PRTArtificial SequenceSynthetic peptide 244Lys Lys
Phe Phe Ala Phe Phe Arg Lys Arg1 5 1024510PRTArtificial
SequenceSynthetic peptide 245Arg Lys Arg Phe Phe Ala Phe Phe Lys
Lys1 5 1024610PRTArtificial SequenceSynthetic peptide 246Lys Arg
Lys Lys Arg Ala Ala Ala Phe Phe1 5 102478PRTArtificial
SequenceSynthetic peptide 247Lys Lys Lys Lys Phe Phe Phe Phe1
524810PRTArtificial SequenceSynthetic peptide 248Lys Arg Lys Lys
Arg Ala Ala Ala Ala Phe1 5 1024910PRTArtificial SequenceSynthetic
peptide 249Lys Lys Lys Ala Phe Phe Phe Ala Lys Lys1 5
1025010PRTArtificial SequenceSynthetic peptide 250Lys Arg Lys Lys
Arg Ala Ala Ala Ala Ala1 5 1025110PRTArtificial SequenceSynthetic
peptide 251Lys Lys Lys Lys Lys Ala Ala Ala Ala Ala1 5
102528PRTArtificial SequenceSynthetic peptide 252Lys Lys Lys Lys
Ala Ala Ala Ala1 52539PRTArtificial SequenceSynthetic peptide
253Phe Lys Phe Lys Phe Lys Phe Lys Phe1 52548PRTArtificial
SequenceSynthetic peptide 254Phe Arg Phe Lys Phe Lys Phe Arg1
52558PRTArtificial SequenceSynthetic peptide 255Arg Phe Gln Phe Lys
Phe Arg Phe1 52567PRTArtificial SequenceSynthetic peptide 256Phe
Arg Phe Lys Phe Lys Phe1 52576PRTArtificial SequenceSynthetic
peptide 257Phe Arg Phe Lys Phe Ala1 52589PRTArtificial
SequenceSynthetic peptide 258Arg Arg Phe Pro Arg Pro Pro Phe Phe1
52599PRTArtificial SequenceSynthetic peptide 259Phe Phe Pro Pro Arg
Pro Phe Arg Arg1 52609PRTArtificial SequenceSynthetic peptide
260Leu Tyr Pro Pro Arg Pro Phe Arg Arg1 52619PRTArtificial
SequenceSynthetic peptide 261Arg Arg Ile Pro Arg Pro Pro Tyr Leu1
526210PRTArtificial SequenceSynthetic peptide 262Pro Phe Arg Pro
Pro Pro Arg Pro Arg Phe1 5 1026310PRTArtificial SequenceSynthetic
peptide 263Pro Arg Pro Arg Pro Pro Pro Arg Phe Phe1 5
102649PRTArtificial SequenceSynthetic peptide 264Phe Phe Pro Pro
Lys Pro Phe Lys Lys1 52659PRTArtificial SequenceSynthetic peptide
265Lys Lys Ile Pro Lys Pro Pro Tyr Leu1 526610PRTArtificial
SequenceSynthetic peptide 266Pro Phe Lys Pro Pro Pro Lys Pro Lys
Pro1 5 1026710PRTArtificial SequenceSynthetic peptide 267Pro Lys
Pro Lys Pro Pro Pro Lys Phe Pro1 5 102689PRTArtificial
SequenceSynthetic peptide 268Leu Tyr Pro Pro Lys Pro Ile Lys Lys1
52699PRTArtificial SequenceSynthetic peptide 269Lys Lys Phe Pro Lys
Pro Pro Phe Phe1 527023PRTArtificial SequenceSynthetic peptide
270Glu Phe Glu Phe Phe Phe Arg Phe Phe Phe Gly Gly Gly Glu Phe Glu1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2027123PRTArtificial
SequenceSynthetic peptide 271Gln Phe Glu Phe Phe Phe Arg Phe Phe
Phe Gly Gly Gly Gln Phe Glu1 5 10 15Phe Phe Phe Arg Phe Phe Phe
2027223PRTArtificial SequenceSynthetic peptide 272Asp Phe Glu Phe
Phe Phe Arg Phe Phe Phe Gly Gly Gly Asp Phe Glu1 5 10 15Phe Phe Phe
Arg Phe Phe Phe 2027323PRTArtificial SequenceSynthetic peptide
273Glu Phe Asn Phe Phe Phe Arg Phe Phe Phe Gly Gly Gly Glu Phe Asn1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2027420PRTArtificial
SequenceSynthetic peptide 274Phe Phe Phe Arg Phe Phe Phe Glu Phe
Gln Phe Phe Phe Arg Phe Phe1 5 10 15Phe Glu Phe Gln
2027523PRTArtificial SequenceSynthetic peptide 275Phe Phe Phe Arg
Phe Phe Phe Glu Phe Gln Gly Gly Gly Phe Phe Phe1 5 10 15Arg Phe Phe
Phe Glu Phe Gln 2027620PRTArtificial SequenceSynthetic peptide
276Arg Trp Arg Lys Phe Phe Lys Arg Phe Phe Gln Phe Glu Phe Phe Phe1
5 10 15Arg Phe Phe Phe 2027723PRTArtificial SequenceSynthetic
peptide 277Arg Trp Arg Lys Phe Phe Lys Arg Phe Phe Gly Gly Gly Phe
Phe Phe1 5 10 15Arg Phe Phe Phe Asn Phe Asn 2027820PRTArtificial
SequenceSynthetic peptide 278Arg Phe Arg Lys Phe Phe Lys Arg Phe
Phe Gln Phe Glu Phe Phe Phe1 5 10 15Arg Phe Phe Phe
2027923PRTArtificial SequenceSynthetic peptide 279Arg Phe Arg Lys
Phe Phe Lys Arg Phe Phe Gly Gly Gly Phe Phe Phe1 5 10 15Arg Phe Phe
Phe Asn Phe Asn 2028023PRTArtificial SequenceSynthetic peptide
280Arg Trp Arg Lys Phe Phe Lys Arg Phe Phe Gly Gly Gly Phe Phe Phe1
5 10 15Arg Phe Phe Phe Glu Phe Gln 2028123PRTArtificial
SequenceSynthetic peptide 281Arg Phe Arg Lys Phe Phe Lys Arg Phe
Phe Gly Gly Gly Phe Phe Phe1 5 10 15Arg Phe Phe Phe Glu Phe Gln
2028220PRTArtificial SequenceSynthetic peptide 282Glu Phe Glu Phe
Phe Phe Arg Phe Phe Phe Glu Phe Glu Phe Phe Phe1 5 10 15Arg Phe Phe
Phe 2028320PRTArtificial SequenceSynthetic peptide 283Arg Trp Arg
Lys Phe Phe Lys Arg Phe Phe Asn Phe Asn Phe Phe Phe1 5 10 15Arg Phe
Phe Phe 2028420PRTArtificial SequenceSynthetic peptide 284Gln Phe
Glu Phe Phe Phe Arg Phe Phe Phe Gln Phe Glu Phe Phe Phe1 5 10 15Arg
Phe Phe Phe 2028523PRTArtificial SequenceSynthetic peptide 285Arg
Trp Arg Lys Phe Phe Lys Arg Phe Phe Gly Gly Gly Asn Phe Asn1 5 10
15Phe Phe Phe Arg Phe Phe Phe 2028620PRTArtificial
SequenceSynthetic peptide 286Glu Phe Asn Phe Phe Phe Arg Phe Phe
Phe Glu Phe Asn Phe Phe Phe1 5 10 15Arg Phe Phe Phe
2028720PRTArtificial SequenceSynthetic peptide 287Arg Phe Arg Lys
Phe Phe Lys Arg Phe Phe Asn Phe Asn Phe Phe Phe1 5 10 15Arg Phe Phe
Phe 2028823PRTArtificial SequenceSynthetic peptide 288Phe Phe Arg
Lys Phe Phe Lys Arg Phe Arg Gly Gly Gly Asn Phe Asn1 5 10 15Phe Phe
Phe Arg Phe Phe Phe 2028923PRTArtificial SequenceSynthetic peptide
289Arg Phe Arg Lys Phe Phe Lys Arg Phe Phe Gly Gly Gly Asn Phe Asn1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2029020PRTArtificial
SequenceSynthetic peptide 290Asp Phe Glu Phe Phe Phe Arg Phe Phe
Phe Asp Phe Glu Phe Phe Phe1 5 10 15Arg Phe Phe Phe
2029120PRTArtificial SequenceSynthetic peptide 291Arg Trp Arg Lys
Phe Phe Lys Arg Phe Phe Phe Phe Phe Arg Phe Phe1 5 10 15Phe Glu Phe
Gln 2029220PRTArtificial SequenceSynthetic peptide 292Arg Phe Arg
Lys Phe Phe Lys Arg Phe Phe Phe Phe Phe Arg Phe Phe1 5 10 15Phe Glu
Phe Gln 2029323PRTArtificial SequenceSynthetic peptide 293Phe Phe
Arg Lys Phe Phe Lys Arg Phe Arg Gly Gly Gly Phe Phe Phe1 5 10 15Arg
Phe Phe Phe Asn Phe Asn 2029423PRTArtificial SequenceSynthetic
peptide 294Phe Phe Arg Lys Phe Phe Lys Arg Trp Arg Gly Gly Gly Phe
Phe Phe1 5 10 15Arg Phe Phe Phe Asn Phe Asn 2029520PRTArtificial
SequenceSynthetic peptide 295Arg Phe Arg Lys Phe Phe Lys Arg Phe
Phe Phe Phe Phe Arg Phe Phe1 5 10 15Phe Asn Phe Asn
2029621PRTArtificial SequenceSynthetic peptide 296Phe Phe Arg Lys
Phe Phe Lys Arg Phe Arg Phe Phe Phe Arg Phe Phe1 5 10 15Phe Glu Phe
Gln Arg 2029723PRTArtificial SequenceSynthetic peptide 297Phe Phe
Glu His Phe Trp Lys Glu Phe Asn Gly Gly Gly Asn Phe Gln1 5 10 15Lys
Trp Phe His Gln Phe Phe 2029820PRTArtificial SequenceSynthetic
peptide 298Phe Phe Arg Lys Phe Phe Lys Arg Trp Arg Gln Phe Glu Phe
Phe Phe1 5 10 15Arg Phe Phe Phe 2029920PRTArtificial
SequenceSynthetic peptide 299Arg Trp Arg Lys Phe Phe Lys Arg Phe
Phe Phe Phe Phe Arg Phe Phe1 5 10 15Phe Asn Phe Asn
2030023PRTArtificial SequenceSynthetic peptide 300Asn Phe Gln Lys
Trp Phe His Gln Phe Phe Gly Gly Gly Phe Phe Glu1 5 10 15His Phe Trp
Lys Glu Phe Asn 2030123PRTArtificial SequenceSynthetic peptide
301Phe Phe Arg Lys Phe Phe Lys Arg Trp Arg Gly Gly Gly Asn Phe Asn1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2030221PRTArtificial
SequenceSynthetic peptide 302Phe Phe Arg Lys Phe Phe Lys Arg Trp
Arg Phe Phe Phe Arg Phe Phe1 5 10 15Phe Glu Phe Gln Arg
2030320PRTArtificial SequenceSynthetic peptide 303Phe Phe Arg Lys
Phe Phe Lys Arg Trp Arg Asn Phe Asn Phe Phe Phe1 5 10 15Arg Phe Phe
Phe 2030424PRTArtificial SequenceSynthetic peptide 304Phe Phe Arg
Lys Phe Phe Lys Arg Phe Arg Gly Gly Gly Phe Phe Phe1 5 10 15Arg Phe
Phe Phe Glu Phe Gln Arg 2030520PRTArtificial SequenceSynthetic
peptide 305Phe Phe Arg Lys Phe Phe Lys Arg Phe Arg Gln Phe Glu Phe
Phe Phe1 5 10 15Arg Phe Phe Phe 2030620PRTArtificial
SequenceSynthetic peptide 306Phe Phe Arg Lys Phe Phe Lys Arg Phe
Arg Asn Phe Asn Phe Phe Phe1 5 10 15Arg Phe Phe Phe
2030723PRTArtificial SequenceSynthetic peptide 307Arg Phe Arg Lys
Phe Phe Lys Arg Phe Phe Gly Gly Gly Gln Phe Glu1 5 10 15Phe Phe Phe
Arg Phe Phe Phe 2030824PRTArtificial SequenceSynthetic peptide
308Phe Phe Arg Lys Phe Phe Lys Arg Trp Arg Gly Gly Gly Phe Phe Phe1
5 10 15Arg Phe Phe Phe Glu Phe Gln Arg 2030923PRTArtificial
SequenceSynthetic peptide 309Phe Phe Arg Lys Phe Phe Lys Arg Phe
Arg Gly Gly Gly Gln Phe Glu1 5 10 15Phe Phe Phe Arg Phe Phe Phe
2031023PRTArtificial SequenceSynthetic peptide 310Phe Phe Arg Lys
Phe Phe Lys Arg Trp Arg Gly Gly Gly Gln Phe Glu1 5 10 15Phe Phe Phe
Arg Phe Phe Phe 2031123PRTArtificial SequenceSynthetic peptide
311Arg Trp Arg Lys Phe Phe Lys Arg Phe Phe Gly Gly Gly Gln Phe Glu1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2031222PRTArtificial
SequenceSynthetic peptide 312Gln Phe Asn His Phe Phe Lys Glu Phe
Gly Gly Gly Gln Phe Asn His1 5 10 15Phe Phe Lys Glu Phe Phe
2031320PRTArtificial SequenceSynthetic peptide 313Phe Phe Arg Lys
Phe Phe Lys Arg Phe Arg Phe Phe Phe Arg Phe Phe1 5 10 15Phe Asn Phe
Asn 2031420PRTArtificial SequenceSynthetic peptide 314Phe Phe Arg
Lys Phe Phe Lys Arg Trp Arg Phe Phe Phe Arg Phe Phe1 5 10 15Phe Asn
Phe Asn 2031523PRTArtificial SequenceSynthetic peptide 315Phe Phe
Glu His Phe Trp Lys Glu Phe Asn Gly Gly Gly Phe Phe Glu1 5 10 15His
Phe Trp Lys Glu Phe Asn 2031622PRTArtificial SequenceSynthetic
peptide 316Phe Phe Glu His Phe Trp Lys Glu Phe Gly Gly Gly Asn Phe
Gln Lys1 5 10 15Trp Phe His Gln Phe Phe 2031722PRTArtificial
SequenceSynthetic peptide 317Asn Phe Gln Lys Trp Phe His Gln Phe
Gly Gly Gly Phe Phe Glu His1 5 10 15Phe Trp Lys Glu Phe Asn
2031822PRTArtificial SequenceSynthetic peptide 318Phe Phe Glu His
Phe Trp Lys Glu Phe Asn Gly Gly Gly Leu His Lys1 5 10 15Met Tyr Asn
Gln Val Trp 2031923PRTArtificial SequenceSynthetic peptide 319Asn
Phe Gln Lys Trp Phe His Gln Phe Phe Gly Gly Gly Asn Phe Gln1 5 10
15Lys Trp Phe His Gln Phe Phe 2032023PRTArtificial
SequenceSynthetic peptide 320Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Gly Gly Gly Asn Phe Gln1 5 10 15Lys Trp Phe His Gln Phe Phe
2032123PRTArtificial SequenceSynthetic peptide 321Phe Phe Glu Lys
Phe Phe His Asn Phe Gln Gly Gly Gly Phe Phe Glu1 5 10 15Lys Phe Phe
His Asn Phe Gln 2032223PRTArtificial SequenceSynthetic peptide
322Phe Phe Gln His Phe Trp Lys Gln Phe Asn Gly Gly Gly Phe Phe Gln1
5 10 15His Phe Trp Lys Gln Phe Asn 2032320PRTArtificial
SequenceSynthetic peptide 323Asn Phe Gln Lys Trp Phe His Gln Phe
Phe Asn Phe Gln Lys Trp Phe1 5 10 15His Gln Phe Phe
2032423PRTArtificial SequenceSynthetic peptide 324Phe Ala Lys Lys
Phe Ala Gln Lys Phe Lys Gly Gly Gly Asn Phe Gln1 5 10 15Lys Trp Phe
His Gln Phe Phe 2032523PRTArtificial SequenceSynthetic peptide
325Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Gln Phe Glu1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2032619PRTArtificial
SequenceSynthetic peptide 326Gln Phe Asn His Phe Phe Lys Glu Phe
Gln Phe Asn His Phe Phe Lys1 5 10 15Glu Phe Phe32723PRTArtificial
SequenceSynthetic peptide 327Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Gly Gly Gly Asp Phe Glu1 5 10 15Phe Phe Phe Arg Phe Phe Phe
2032822PRTArtificial SequenceSynthetic peptide 328Phe Phe Glu His
Phe Trp Lys Glu Phe Asn Gly Gly Gly Trp Val Gln1 5 10 15Asn Tyr Met
Lys His Leu 2032920PRTArtificial SequenceSynthetic peptide 329Phe
Ala Lys Lys Phe Ala Lys Lys Phe Lys Gln Phe Glu Phe Phe Phe1 5 10
15Arg Phe Phe Phe 2033020PRTArtificial SequenceSynthetic peptide
330Phe Phe Gln His Phe Trp Lys Gln Phe Asn Phe Phe Gln His Phe Trp1
5 10 15Lys Gln Phe Asn 2033120PRTArtificial SequenceSynthetic
peptide 331Phe Phe Glu His Phe Trp Lys Glu Phe Asn Phe Phe Glu His
Phe Trp1 5 10 15Lys Glu Phe Asn 2033220PRTArtificial
SequenceSynthetic peptide 332Asn Phe Glu Lys Trp Phe His Glu Phe
Phe Asn Phe Glu Lys Trp Phe1 5 10 15His Glu Phe Phe
2033323PRTArtificial SequenceSynthetic peptide 333Phe Ala Lys Lys
Phe Ala Lys Lys Phe Lys Gly Gly Gly Gln Phe Asn1 5 10 15His Phe Phe
Lys Glu Phe Phe 2033423PRTArtificial SequenceSynthetic peptide
334Asn Phe Glu Lys Trp Phe His Glu Phe Phe Gly Gly Gly Asn Phe Glu1
5 10 15Lys Trp Phe His Glu Phe Phe 2033523PRTArtificial
SequenceSynthetic peptide 335Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Gly Gly Gly Phe Phe Phe1 5 10 15Arg Phe Phe Phe Glu Phe Gln
2033620PRTArtificial SequenceSynthetic peptide 336Phe Ala Lys Lys
Phe Ala Lys Lys Phe Lys Asp Phe Glu Phe Phe Phe1 5 10 15Arg Phe Phe
Phe 2033723PRTArtificial SequenceSynthetic peptide 337Gln Phe Asn
His Phe Phe Lys Glu Phe Phe Gly Gly Gly Gln Phe Asn1 5 10 15His Phe
Phe Lys Glu Phe Phe 2033823PRTArtificial SequenceSynthetic peptide
338Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Glu Phe Glu1
5 10 15Phe Phe Phe Arg Phe Phe Phe 2033923PRTArtificial
SequenceSynthetic peptide 339Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Gly Gly Gly Glu Phe Asn1 5 10 15Phe Phe Phe Arg Phe Phe Phe
2034023PRTArtificial SequenceSynthetic peptide 340Phe Ala Lys Lys
Phe Ala Lys Lys Phe Lys Gly Gly Gly Asn Phe Glu1 5 10 15Lys Trp Phe
His Glu Phe Phe 2034123PRTArtificial SequenceSynthetic peptide
341Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Phe Phe Glu1
5 10 15Lys Phe Phe His Asn Phe Gln 2034220PRTArtificial
SequenceSynthetic peptide 342Gln Phe Asn His Phe Phe Lys Glu Phe
Phe Gln Phe Asn His Phe Phe1 5 10
15Lys Glu Phe Phe 2034320PRTArtificial SequenceSynthetic peptide
343Phe Phe Glu Lys Phe Phe His Asn Phe Gln Phe Phe Glu Lys Phe Phe1
5 10 15His Asn Phe Gln 2034420PRTArtificial SequenceSynthetic
peptide 344Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Glu Phe Glu Phe
Phe Phe1 5 10 15Arg Phe Phe Phe 2034520PRTArtificial
SequenceSynthetic peptide 345Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Glu Phe Asn Phe Phe Phe1 5 10 15Arg Phe Phe Phe
2034620PRTArtificial SequenceSynthetic peptide 346Phe Ala Lys Lys
Phe Ala Lys Lys Phe Lys Phe Phe Glu His Phe Trp1 5 10 15Lys Glu Phe
Asn 2034720PRTArtificial SequenceSynthetic peptide 347Phe Ala Lys
Lys Phe Ala Lys Lys Phe Lys Phe Phe Phe Arg Phe Phe1 5 10 15Phe Glu
Phe Gln 2034819PRTArtificial SequenceSynthetic peptide 348Phe Ala
Lys Lys Phe Ala Lys Lys Phe Lys Leu His Lys Met Tyr Asn1 5 10 15Gln
Val Trp34923PRTArtificial SequenceSynthetic peptide 349Phe Ala Lys
Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Phe Phe Glu1 5 10 15His Phe
Trp Lys Glu Phe Asn 2035022PRTArtificial SequenceSynthetic peptide
350Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Trp Val Gln1
5 10 15Asn Tyr Met Lys His Leu 2035120PRTArtificial
SequenceSynthetic peptide 351Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Asn Phe Gln Lys Trp Phe1 5 10 15His Gln Phe Phe
2035220PRTArtificial SequenceSynthetic peptide 352Phe Ala Lys Lys
Phe Ala Lys Lys Phe Lys Phe Phe Gln His Phe Trp1 5 10 15Lys Gln Phe
Asn 2035323PRTArtificial SequenceSynthetic peptide 353Phe Ala Lys
Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Phe Phe Gln1 5 10 15His Phe
Trp Lys Gln Phe Asn 2035419PRTArtificial SequenceSynthetic peptide
354Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Trp Val Gln Asn Tyr Met1
5 10 15Lys His Leu35520PRTArtificial SequenceSynthetic peptide
355Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gln Phe Asn His Phe Phe1
5 10 15Lys Glu Phe Phe 2035622PRTArtificial SequenceSynthetic
peptide 356Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Leu
His Lys1 5 10 15Met Tyr Asn Gln Val Trp 2035720PRTArtificial
SequenceSynthetic peptide 357Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Asn Phe Glu Lys Trp Phe1 5 10 15His Glu Phe Phe
2035820PRTArtificial SequenceSynthetic peptide 358Phe Ala Lys Lys
Phe Ala Lys Lys Phe Lys Phe Phe Glu Lys Phe Phe1 5 10 15His Asn Phe
Gln 2035923PRTArtificial SequenceSynthetic peptide 359Phe Ala Lys
Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Ala Phe Phe1 5 10 15Lys Lys
Lys Ala Lys Phe Lys 2036023PRTArtificial SequenceSynthetic peptide
360Ala Phe Phe Lys Lys Lys Ala Lys Phe Lys Gly Gly Gly Ala Phe Phe1
5 10 15Lys Lys Lys Ala Lys Phe Lys 2036120PRTArtificial
SequenceSynthetic peptide 361Lys Phe Lys Lys Ala Phe Lys Lys Ala
Phe Lys Phe Lys Lys Ala Phe1 5 10 15Lys Lys Ala Phe
2036223PRTArtificial SequenceSynthetic peptide 362Lys Phe Lys Lys
Ala Phe Lys Lys Ala Phe Gly Gly Gly Lys Phe Lys1 5 10 15Lys Ala Phe
Lys Lys Ala Phe 2036323PRTArtificial SequenceSynthetic peptide
363Phe Ala Lys Lys Phe Ala Lys Lys Phe Lys Gly Gly Gly Phe Ala Lys1
5 10 15Lys Phe Ala Lys Lys Phe Lys 2036420PRTArtificial
SequenceSynthetic peptide 364Phe Ala Lys Lys Phe Ala Lys Lys Phe
Lys Ala Phe Phe Lys Lys Lys1 5 10 15Ala Lys Phe Lys
2036522PRTArtificial SequenceSynthetic peptide 365Pro Ser Arg Lys
Ser Met Glu Lys Ser Val Ala Lys Leu Leu Asn Lys1 5 10 15Ile Ala Lys
Ser Glu Pro 2036620PRTArtificial SequenceSynthetic peptide 366Ala
Phe Phe Lys Lys Lys Ala Lys Phe Lys Ala Phe Phe Lys Lys Lys1 5 10
15Ala Lys Phe Lys 20367579PRTHomo sapiens 367Met Leu Arg Ser Gly
Pro Ala Ser Gly Pro Ser Val Pro Thr Gly Arg1 5 10 15Ala Met Pro Ser
Arg Arg Val Ala Arg Pro Pro Ala Ala Pro Glu Leu 20 25 30Gly Ala Leu
Gly Ser Pro Asp Leu Ser Ser Leu Ser Leu Ala Val Ser 35 40 45Arg Ser
Thr Asp Glu Leu Glu Ile Ile Asp Glu Tyr Ile Lys Glu Asn 50 55 60Gly
Phe Gly Leu Asp Gly Gly Gln Pro Gly Pro Gly Glu Gly Leu Pro65 70 75
80Arg Leu Val Ser Arg Gly Ala Ala Ser Leu Ser Thr Val Thr Leu Gly
85 90 95Pro Val Ala Pro Pro Ala Thr Pro Pro Pro Trp Gly Cys Pro Leu
Gly 100 105 110Arg Leu Val Ser Pro Ala Pro Gly Pro Gly Pro Gln Pro
His Leu Val 115 120 125Ile Thr Glu Gln Pro Lys Gln Arg Gly Met Arg
Phe Arg Tyr Glu Cys 130 135 140Glu Gly Arg Ser Ala Gly Ser Ile Leu
Gly Glu Ser Ser Thr Glu Ala145 150 155 160Ser Lys Thr Leu Pro Ala
Ile Glu Leu Arg Asp Cys Gly Gly Leu Arg 165 170 175Glu Val Glu Val
Thr Ala Cys Leu Val Trp Lys Asp Trp Pro His Arg 180 185 190Val His
Pro His Ser Leu Val Gly Lys Asp Cys Thr Asp Gly Ile Cys 195 200
205Arg Val Arg Leu Arg Pro His Val Ser Pro Arg His Ser Phe Asn Asn
210 215 220Leu Gly Ile Gln Cys Val Arg Lys Lys Glu Ile Glu Ala Ala
Ile Glu225 230 235 240Arg Lys Ile Gln Leu Gly Ile Asp Pro Tyr Asn
Ala Gly Ser Leu Lys 245 250 255Asn His Gln Glu Val Asp Met Asn Val
Val Arg Ile Cys Phe Gln Ala 260 265 270Ser Tyr Arg Asp Gln Gln Gly
Gln Met Arg Arg Met Asp Pro Val Leu 275 280 285Ser Glu Pro Val Tyr
Asp Lys Lys Ser Thr Asn Thr Ser Glu Leu Arg 290 295 300Ile Cys Arg
Ile Asn Lys Glu Ser Gly Pro Cys Thr Gly Gly Glu Glu305 310 315
320Leu Tyr Leu Leu Cys Asp Lys Val Gln Lys Glu Asp Ile Ser Val Val
325 330 335Phe Ser Arg Ala Ser Trp Glu Gly Arg Ala Asp Phe Ser Gln
Ala Asp 340 345 350Val His Arg Gln Ile Ala Ile Val Phe Lys Thr Pro
Pro Tyr Glu Asp 355 360 365Leu Glu Ile Val Glu Pro Val Thr Val Asn
Val Phe Leu Gln Arg Leu 370 375 380Thr Asp Gly Val Cys Ser Glu Pro
Leu Pro Phe Thr Tyr Leu Pro Arg385 390 395 400Asp His Asp Ser Tyr
Gly Val Asp Lys Lys Arg Lys Arg Gly Met Pro 405 410 415Asp Val Leu
Gly Glu Leu Asn Ser Ser Asp Pro His Gly Ile Glu Ser 420 425 430Lys
Arg Arg Lys Lys Lys Pro Ala Ile Leu Asp His Phe Leu Pro Asn 435 440
445His Gly Ser Gly Pro Phe Leu Pro Pro Ser Ala Leu Leu Pro Asp Pro
450 455 460Asp Phe Phe Ser Gly Thr Val Ser Leu Pro Gly Leu Glu Pro
Pro Gly465 470 475 480Gly Pro Asp Leu Leu Asp Asp Gly Phe Ala Tyr
Asp Pro Thr Ala Pro 485 490 495Thr Leu Phe Thr Met Leu Asp Leu Leu
Pro Pro Ala Pro Pro His Ala 500 505 510Ser Ala Val Val Cys Ser Gly
Gly Ala Gly Ala Val Val Gly Glu Thr 515 520 525Pro Gly Pro Glu Pro
Leu Thr Leu Asp Ser Tyr Gln Ala Pro Gly Pro 530 535 540Gly Asp Gly
Gly Thr Ala Ser Leu Val Gly Ser Asn Met Phe Pro Asn545 550 555
560His Tyr Arg Glu Ala Ala Phe Gly Gly Gly Leu Leu Ser Pro Gly Pro
565 570 575Glu Ala Thr368390PRTHomo sapiens 368Met Pro Pro Ser Gly
Leu Arg Leu Leu Pro Leu Leu Leu Pro Leu Leu1 5 10 15Trp Leu Leu Val
Leu Thr Pro Gly Arg Pro Ala Ala Gly Leu Ser Thr 20 25 30Cys Lys Thr
Ile Asp Met Glu Leu Val Lys Arg Lys Arg Ile Glu Ala 35 40 45Ile Arg
Gly Gln Ile Leu Ser Lys Leu Arg Leu Ala Ser Pro Pro Ser 50 55 60Gln
Gly Glu Val Pro Pro Gly Pro Leu Pro Glu Ala Val Leu Ala Leu65 70 75
80Tyr Asn Ser Thr Arg Asp Arg Val Ala Gly Glu Ser Ala Glu Pro Glu
85 90 95Pro Glu Pro Glu Ala Asp Tyr Tyr Ala Lys Glu Val Thr Arg Val
Leu 100 105 110Met Val Glu Thr His Asn Glu Ile Tyr Asp Lys Phe Lys
Gln Ser Thr 115 120 125His Ser Ile Tyr Met Phe Phe Asn Thr Ser Glu
Leu Arg Glu Ala Val 130 135 140Pro Glu Pro Val Leu Leu Ser Arg Ala
Glu Leu Arg Leu Leu Arg Leu145 150 155 160Lys Leu Lys Val Glu Gln
His Val Glu Leu Tyr Gln Lys Tyr Ser Asn 165 170 175Asn Ser Trp Arg
Tyr Leu Ser Asn Arg Leu Leu Ala Pro Ser Asp Ser 180 185 190Pro Glu
Trp Leu Ser Phe Asp Val Thr Gly Val Val Arg Gln Trp Leu 195 200
205Ser Arg Gly Gly Glu Ile Glu Gly Phe Arg Leu Ser Ala His Cys Ser
210 215 220Cys Asp Ser Arg Asp Asn Thr Leu Gln Val Asp Ile Asn Gly
Phe Thr225 230 235 240Thr Gly Arg Arg Gly Asp Leu Ala Thr Ile His
Gly Met Asn Arg Pro 245 250 255Phe Leu Leu Leu Met Ala Thr Pro Leu
Glu Arg Ala Gln His Leu Gln 260 265 270Ser Ser Arg His Arg Arg Ala
Leu Asp Thr Asn Tyr Cys Phe Ser Ser 275 280 285Thr Glu Lys Asn Cys
Cys Val Arg Gln Leu Tyr Ile Asp Phe Arg Lys 290 295 300Asp Leu Gly
Trp Lys Trp Ile His Glu Pro Lys Gly Tyr His Ala Asn305 310 315
320Phe Cys Leu Gly Pro Cys Pro Tyr Ile Trp Ser Leu Asp Thr Gln Tyr
325 330 335Ser Lys Val Leu Ala Leu Tyr Asn Gln His Asn Pro Gly Ala
Ser Ala 340 345 350Ala Pro Cys Cys Val Pro Gln Ala Leu Glu Pro Leu
Pro Ile Val Tyr 355 360 365Tyr Val Gly Arg Lys Pro Lys Val Glu Gln
Leu Ser Asn Met Ile Val 370 375 380Arg Ser Cys Lys Cys Ser385
3903692556PRTHomo sapiensMISC_FEATURE(891)..(891)Xaa is denoted "X"
in GenBank Sequence for Homo sapiens Notch1 (GenBank Acc. No.
AAG33848.1) 369Met Pro Pro Leu Leu Ala Pro Leu Leu Cys Leu Ala Leu
Leu Pro Ala1 5 10 15Leu Ala Ala Arg Gly Pro Arg Cys Ser Gln Pro Gly
Glu Thr Cys Leu 20 25 30Asn Gly Gly Lys Cys Glu Ala Ala Asn Gly Thr
Glu Ala Cys Val Cys 35 40 45Gly Gly Ala Phe Val Gly Pro Arg Cys Gln
Asp Pro Asn Pro Cys Leu 50 55 60Ser Thr Pro Cys Lys Asn Ala Gly Thr
Cys His Val Val Asp Arg Arg65 70 75 80Gly Val Ala Asp Tyr Ala Cys
Ser Cys Ala Leu Gly Phe Ser Gly Pro 85 90 95Leu Cys Leu Thr Pro Leu
Asp Asn Ala Cys Leu Thr Asn Pro Cys Arg 100 105 110Asn Gly Gly Thr
Cys Asp Leu Leu Thr Leu Thr Glu Tyr Lys Cys Arg 115 120 125Cys Pro
Pro Gly Trp Ser Gly Lys Ser Cys Gln Gln Ala Asp Pro Cys 130 135
140Ala Ser Asn Pro Cys Ala Asn Gly Gly Gln Cys Leu Pro Phe Glu
Ala145 150 155 160Ser Tyr Ile Cys His Cys Pro Pro Ser Phe His Gly
Pro Thr Cys Arg 165 170 175Gln Asp Val Asn Glu Cys Gly Gln Lys Pro
Arg Leu Cys Arg His Gly 180 185 190Gly Thr Cys His Asn Glu Val Gly
Ser Tyr Arg Cys Val Cys Arg Ala 195 200 205Thr His Thr Gly Pro Asn
Cys Glu Arg Pro Tyr Val Pro Cys Ser Pro 210 215 220Ser Pro Cys Gln
Asn Gly Gly Thr Cys Arg Pro Thr Gly Asp Val Thr225 230 235 240His
Glu Cys Ala Cys Leu Pro Gly Phe Thr Gly Gln Asn Cys Glu Glu 245 250
255Asn Ile Asp Asp Cys Pro Gly Asn Asn Cys Lys Asn Gly Gly Ala Cys
260 265 270Val Asp Gly Val Asn Thr Tyr Asn Cys Pro Cys Pro Pro Glu
Trp Thr 275 280 285Gly Gln Tyr Cys Thr Glu Asp Val Asp Glu Cys Gln
Leu Met Pro Asn 290 295 300Ala Cys Gln Asn Gly Gly Thr Cys His Asn
Thr His Gly Gly Tyr Asn305 310 315 320Cys Val Cys Val Asn Gly Trp
Thr Gly Glu Asp Cys Ser Glu Asn Ile 325 330 335Asp Asp Cys Ala Ser
Ala Ala Cys Phe His Gly Ala Thr Cys His Asp 340 345 350Arg Val Ala
Ser Phe Tyr Cys Glu Cys Pro His Gly Arg Thr Gly Leu 355 360 365Leu
Cys His Leu Asn Asp Ala Cys Ile Ser Asn Pro Cys Asn Glu Gly 370 375
380Ser Asn Cys Asp Thr Asn Pro Val Asn Gly Lys Ala Ile Cys Thr
Cys385 390 395 400Pro Ser Gly Tyr Thr Gly Pro Ala Cys Ser Gln Asp
Val Asp Glu Cys 405 410 415Ser Leu Gly Ala Asn Pro Cys Glu His Ala
Gly Lys Cys Ile Asn Thr 420 425 430Leu Gly Ser Phe Glu Cys Gln Cys
Leu Gln Gly Tyr Thr Gly Pro Arg 435 440 445Cys Glu Ile Asp Val Asn
Glu Cys Val Ser Asn Pro Cys Gln Asn Asp 450 455 460Ala Thr Cys Leu
Asp Gln Ile Gly Glu Phe Gln Cys Met Cys Met Pro465 470 475 480Gly
Tyr Glu Gly Val His Cys Glu Val Asn Thr Asp Glu Cys Ala Ser 485 490
495Ser Pro Cys Leu His Asn Gly Arg Cys Leu Asp Lys Ile Asn Glu Phe
500 505 510Gln Cys Glu Cys Pro Thr Gly Phe Thr Gly His Leu Cys Gln
Tyr Asp 515 520 525Val Asp Glu Cys Ala Ser Thr Pro Cys Lys Asn Gly
Ala Lys Cys Leu 530 535 540Asp Gly Pro Asn Thr Tyr Thr Cys Val Cys
Thr Glu Gly Tyr Thr Gly545 550 555 560Thr His Cys Glu Val Asp Ile
Asp Glu Cys Asp Pro Asp Pro Cys His 565 570 575Tyr Gly Ser Cys Lys
Asp Gly Val Ala Thr Phe Thr Cys Leu Cys Arg 580 585 590Pro Gly Tyr
Thr Gly His His Cys Glu Thr Asn Ile Asn Glu Cys Ser 595 600 605Ser
Gln Pro Cys Arg Leu Arg Gly Thr Cys Gln Asp Pro Asp Asn Ala 610 615
620Tyr Leu Cys Phe Cys Leu Lys Gly Thr Thr Gly Pro Asn Cys Glu
Ile625 630 635 640Asn Leu Asp Asp Cys Ala Ser Ser Pro Cys Asp Ser
Gly Thr Cys Leu 645 650 655Asp Lys Ile Asp Gly Tyr Glu Cys Ala Cys
Glu Pro Gly Tyr Thr Gly 660 665 670Ser Met Cys Asn Ser Asn Ile Asp
Glu Cys Ala Gly Asn Pro Cys His 675 680 685Asn Gly Gly Thr Cys Glu
Asp Gly Ile Asn Gly Phe Thr Cys Arg Cys 690 695 700Pro Glu Gly Tyr
His Asp Pro Thr Cys Leu Ser Glu Val Asn Glu Cys705 710 715 720Asn
Ser Asn Pro Cys Val His Gly Ala Cys Arg Asp Ser Leu Asn Gly 725 730
735Tyr Lys Cys Asp Cys Asp Pro Gly Trp Ser Gly Thr Asn Cys Asp Ile
740 745 750Asn Asn Asn Glu Cys Glu Ser Asn Pro Cys Val Asn Gly Gly
Thr Cys 755 760 765Lys Asp Met Thr Ser Gly Ile Val Cys Thr Cys Arg
Glu Gly Phe Ser 770 775 780Gly Pro Asn Cys Gln Thr Asn Ile Asn Glu
Cys Ala Ser Asn Pro Cys785 790 795 800Leu Asn Lys Gly Thr Cys Ile
Asp Asp Val Ala Gly Tyr Lys Cys Asn
805 810 815Cys Leu Leu Pro Tyr Thr Gly Ala Thr Cys Glu Val Val Leu
Ala Pro 820 825 830Cys Ala Pro Ser Pro Cys Arg Asn Gly Gly Glu Cys
Arg Gln Ser Glu 835 840 845Asp Tyr Glu Ser Phe Ser Cys Val Cys Pro
Thr Ala Gly Ala Lys Gly 850 855 860Gln Thr Cys Glu Val Asp Ile Asn
Glu Cys Val Leu Ser Pro Cys Arg865 870 875 880His Gly Ala Ser Cys
Gln Asn Thr His Gly Xaa Tyr Arg Cys His Cys 885 890 895Gln Ala Gly
Tyr Ser Gly Arg Asn Cys Glu Thr Asp Ile Asp Asp Cys 900 905 910Arg
Pro Asn Pro Cys His Asn Gly Gly Ser Cys Thr Asp Gly Ile Asn 915 920
925Thr Ala Phe Cys Asp Cys Leu Pro Gly Phe Arg Gly Thr Phe Cys Glu
930 935 940Glu Asp Ile Asn Glu Cys Ala Ser Asp Pro Cys Arg Asn Gly
Ala Asn945 950 955 960Cys Thr Asp Cys Val Asp Ser Tyr Thr Cys Thr
Cys Pro Ala Gly Phe 965 970 975Ser Gly Ile His Cys Glu Asn Asn Thr
Pro Asp Cys Thr Glu Ser Ser 980 985 990Cys Phe Asn Gly Gly Thr Cys
Val Asp Gly Ile Asn Ser Phe Thr Cys 995 1000 1005Leu Cys Pro Pro
Gly Phe Thr Gly Ser Tyr Cys Gln His Val Val 1010 1015 1020Asn Glu
Cys Asp Ser Arg Pro Cys Leu Leu Gly Gly Thr Cys Gln 1025 1030
1035Asp Gly Arg Gly Leu His Arg Cys Thr Cys Pro Gln Gly Tyr Thr
1040 1045 1050Gly Pro Asn Cys Gln Asn Leu Val His Trp Cys Asp Ser
Ser Pro 1055 1060 1065Cys Lys Asn Gly Gly Lys Cys Trp Gln Thr His
Thr Gln Tyr Arg 1070 1075 1080Cys Glu Cys Pro Ser Gly Trp Thr Gly
Leu Tyr Cys Asp Val Pro 1085 1090 1095Ser Val Ser Cys Glu Val Ala
Ala Gln Arg Gln Gly Val Asp Val 1100 1105 1110Ala Arg Leu Cys Gln
His Gly Gly Leu Cys Val Asp Ala Gly Asn 1115 1120 1125Thr His His
Cys Arg Cys Gln Ala Gly Tyr Thr Gly Ser Tyr Cys 1130 1135 1140Glu
Asp Leu Val Asp Glu Cys Ser Pro Ser Pro Cys Gln Asn Gly 1145 1150
1155Ala Thr Cys Thr Asp Tyr Leu Gly Gly Tyr Ser Cys Lys Cys Val
1160 1165 1170Ala Gly Tyr His Gly Val Asn Cys Ser Glu Glu Ile Asp
Glu Cys 1175 1180 1185Leu Ser His Pro Cys Gln Asn Gly Gly Thr Cys
Leu Asp Leu Pro 1190 1195 1200Asn Thr Tyr Lys Cys Ser Cys Pro Arg
Gly Thr Gln Gly Val His 1205 1210 1215Cys Glu Ile Asn Val Asp Asp
Cys Asn Pro Pro Val Asp Pro Val 1220 1225 1230Ser Arg Ser Pro Lys
Cys Phe Asn Asn Gly Thr Cys Val Asp Gln 1235 1240 1245Val Gly Gly
Tyr Ser Cys Thr Cys Pro Pro Gly Phe Val Gly Glu 1250 1255 1260Arg
Cys Glu Gly Asp Val Asn Glu Cys Leu Ser Asn Pro Cys Asp 1265 1270
1275Ala Arg Gly Thr Gln Asn Cys Val Gln Arg Val Asn Asp Phe His
1280 1285 1290Cys Glu Cys Arg Ala Gly His Thr Gly Arg Arg Cys Glu
Ser Val 1295 1300 1305Ile Asn Gly Cys Lys Gly Lys Pro Cys Lys Asn
Gly Gly Thr Cys 1310 1315 1320Ala Val Ala Ser Asn Thr Ala Arg Gly
Phe Ile Cys Lys Cys Pro 1325 1330 1335Ala Gly Phe Glu Gly Ala Thr
Cys Glu Asn Asp Ala Arg Thr Cys 1340 1345 1350Gly Ser Leu Arg Cys
Leu Asn Gly Gly Thr Cys Ile Ser Gly Pro 1355 1360 1365Arg Ser Pro
Thr Cys Leu Cys Leu Gly Pro Phe Thr Gly Pro Glu 1370 1375 1380Cys
Gln Phe Pro Ala Ser Ser Pro Cys Leu Gly Gly Asn Pro Cys 1385 1390
1395Tyr Asn Gln Gly Thr Cys Glu Pro Thr Ser Glu Ser Pro Phe Tyr
1400 1405 1410Arg Cys Leu Cys Pro Ala Lys Phe Asn Gly Leu Leu Cys
His Ile 1415 1420 1425Leu Asp Tyr Ser Phe Gly Gly Gly Ala Gly Arg
Asp Ile Pro Pro 1430 1435 1440Pro Leu Ile Glu Glu Ala Cys Glu Leu
Pro Glu Cys Gln Glu Asp 1445 1450 1455Ala Gly Asn Lys Val Cys Ser
Leu Gln Cys Asn Asn His Ala Cys 1460 1465 1470Gly Trp Asp Gly Gly
Asp Cys Ser Leu Asn Phe Asn Asp Pro Trp 1475 1480 1485Lys Asn Cys
Thr Gln Ser Leu Gln Cys Trp Lys Tyr Phe Ser Asp 1490 1495 1500Gly
His Cys Asp Ser Gln Cys Asn Ser Ala Gly Cys Leu Phe Asp 1505 1510
1515Gly Phe Asp Cys Gln Arg Ala Glu Gly Gln Cys Asn Pro Leu Tyr
1520 1525 1530Asp Gln Tyr Cys Lys Asp His Phe Ser Asp Gly His Cys
Asp Gln 1535 1540 1545Gly Cys Asn Ser Ala Glu Cys Glu Trp Asp Gly
Leu Asp Cys Ala 1550 1555 1560Glu His Val Pro Glu Arg Leu Ala Ala
Gly Thr Leu Val Val Val 1565 1570 1575Val Leu Met Pro Pro Glu Gln
Leu Arg Asn Ser Ser Phe His Phe 1580 1585 1590Leu Arg Glu Leu Ser
Arg Val Leu His Thr Asn Val Val Phe Lys 1595 1600 1605Arg Asp Ala
His Gly Gln Gln Met Ile Phe Pro Tyr Tyr Gly Arg 1610 1615 1620Glu
Glu Glu Leu Arg Lys His Pro Ile Lys Arg Ala Ala Glu Gly 1625 1630
1635Trp Ala Ala Pro Asp Ala Leu Leu Gly Gln Val Lys Ala Ser Leu
1640 1645 1650Leu Pro Gly Gly Ser Glu Gly Gly Arg Arg Arg Arg Glu
Leu Asp 1655 1660 1665Pro Met Asp Val Arg Gly Ser Ile Val Tyr Leu
Glu Ile Asp Asn 1670 1675 1680Arg Gln Cys Val Gln Ala Ser Ser Gln
Cys Phe Gln Ser Ala Thr 1685 1690 1695Asp Val Ala Ala Phe Leu Gly
Ala Leu Ala Ser Leu Gly Ser Leu 1700 1705 1710Asn Ile Pro Tyr Lys
Ile Glu Ala Val Gln Ser Glu Thr Val Glu 1715 1720 1725Pro Pro Pro
Pro Ala Gln Leu His Phe Met Tyr Val Ala Ala Ala 1730 1735 1740Ala
Phe Val Leu Leu Phe Phe Val Gly Cys Gly Val Leu Leu Ser 1745 1750
1755Arg Lys Arg Arg Arg Gln His Gly Gln Leu Trp Phe Pro Glu Gly
1760 1765 1770Phe Lys Val Ser Glu Ala Ser Lys Lys Lys Arg Arg Glu
Pro Leu 1775 1780 1785Gly Glu Asp Ser Val Gly Leu Lys Pro Leu Lys
Asn Ala Ser Asp 1790 1795 1800Gly Ala Leu Met Asp Asp Asn Gln Asn
Glu Trp Gly Asp Glu Asp 1805 1810 1815Leu Glu Thr Lys Lys Phe Arg
Phe Glu Glu Pro Val Val Leu Pro 1820 1825 1830Asp Leu Asp Asp Gln
Thr Asp His Arg Gln Trp Thr Gln Gln His 1835 1840 1845Leu Asp Ala
Ala Asp Leu Arg Met Ser Ala Met Ala Pro Thr Pro 1850 1855 1860Pro
Gln Gly Glu Val Asp Ala Asp Cys Met Asp Val Asn Val Arg 1865 1870
1875Gly Pro Asp Gly Phe Thr Pro Leu Met Ile Ala Ser Cys Ser Gly
1880 1885 1890Gly Gly Leu Glu Thr Gly Asn Ser Glu Glu Glu Glu Asp
Ala Pro 1895 1900 1905Ala Val Ile Ser Asp Phe Ile Tyr Gln Gly Ala
Ser Leu His Asn 1910 1915 1920Gln Thr Asp Arg Thr Gly Glu Thr Ala
Leu His Leu Ala Ala Arg 1925 1930 1935Tyr Ser Arg Ser Asp Ala Ala
Lys Arg Leu Leu Glu Ala Ser Ala 1940 1945 1950Asp Ala Asn Ile Gln
Asp Asn Met Gly Arg Thr Pro Leu His Ala 1955 1960 1965Ala Val Ser
Ala Asp Ala Gln Gly Val Phe Gln Ile Leu Ile Arg 1970 1975 1980Asn
Arg Ala Thr Asp Leu Asp Ala Arg Met His Asp Gly Thr Thr 1985 1990
1995Pro Leu Ile Leu Ala Ala Arg Leu Ala Val Glu Gly Met Leu Glu
2000 2005 2010Asp Leu Ile Asn Ser His Ala Asp Val Asn Ala Val Asp
Asp Leu 2015 2020 2025Gly Lys Ser Ala Leu His Trp Ala Ala Ala Val
Asn Asn Val Asp 2030 2035 2040Ala Ala Val Val Leu Leu Lys Asn Gly
Ala Asn Lys Asp Met Gln 2045 2050 2055Asn Asn Arg Glu Glu Thr Pro
Leu Phe Leu Ala Ala Arg Glu Gly 2060 2065 2070Ser Tyr Glu Thr Ala
Lys Val Leu Leu Asp His Phe Ala Asn Arg 2075 2080 2085Asp Ile Thr
Asp His Met Asp Arg Leu Pro Arg Asp Ile Ala Gln 2090 2095 2100Glu
Arg Met His His Asp Ile Val Arg Leu Leu Asp Glu Tyr Asn 2105 2110
2115Leu Val Arg Ser Pro Gln Leu His Gly Ala Pro Leu Gly Gly Thr
2120 2125 2130Pro Thr Leu Ser Pro Pro Leu Cys Ser Pro Asn Gly Tyr
Leu Gly 2135 2140 2145Ser Leu Lys Pro Gly Val Gln Gly Lys Lys Val
Arg Lys Pro Ser 2150 2155 2160Ser Lys Gly Leu Ala Cys Gly Ser Lys
Glu Ala Lys Asp Leu Lys 2165 2170 2175Ala Arg Arg Lys Lys Ser Gln
Asp Gly Lys Gly Cys Leu Leu Asp 2180 2185 2190Ser Ser Gly Met Leu
Ser Pro Val Asp Ser Leu Glu Ser Pro His 2195 2200 2205Gly Tyr Leu
Ser Asp Val Ala Ser Pro Pro Leu Leu Pro Ser Pro 2210 2215 2220Phe
Gln Gln Ser Pro Ser Val Pro Leu Asn His Leu Pro Gly Met 2225 2230
2235Pro Asp Thr His Leu Gly Ile Gly His Leu Asn Val Ala Ala Lys
2240 2245 2250Pro Glu Met Ala Ala Leu Gly Gly Gly Gly Arg Leu Ala
Phe Glu 2255 2260 2265Thr Gly Pro Pro Arg Leu Ser His Leu Pro Val
Ala Ser Gly Thr 2270 2275 2280Ser Thr Val Leu Gly Ser Ser Ser Gly
Gly Ala Leu Asn Phe Thr 2285 2290 2295Val Gly Gly Ser Thr Ser Leu
Asn Gly Gln Cys Glu Trp Leu Ser 2300 2305 2310Arg Leu Gln Ser Gly
Met Val Pro Asn Gln Tyr Asn Pro Leu Arg 2315 2320 2325Gly Ser Val
Ala Pro Gly Pro Leu Ser Thr Gln Ala Pro Ser Leu 2330 2335 2340Gln
His Gly Met Val Gly Pro Leu His Ser Ser Leu Ala Ala Ser 2345 2350
2355Ala Leu Ser Gln Met Met Ser Tyr Gln Gly Leu Pro Ser Thr Arg
2360 2365 2370Leu Ala Thr Gln Pro His Leu Val Gln Thr Gln Gln Val
Gln Pro 2375 2380 2385Gln Asn Leu Gln Met Gln Gln Gln Asn Leu Gln
Pro Ala Asn Ile 2390 2395 2400Gln Gln Gln Gln Ser Leu Gln Pro Pro
Pro Pro Pro Pro Gln Pro 2405 2410 2415His Leu Gly Val Ser Ser Ala
Ala Ser Gly His Leu Gly Arg Ser 2420 2425 2430Phe Leu Ser Gly Glu
Pro Ser Gln Ala Asp Val Gln Pro Leu Gly 2435 2440 2445Pro Ser Ser
Leu Ala Val His Thr Ile Leu Pro Gln Glu Ser Pro 2450 2455 2460Ala
Leu Pro Thr Ser Leu Pro Ser Ser Leu Val Pro Pro Val Thr 2465 2470
2475Ala Ala Gln Phe Leu Thr Pro Pro Ser Gln His Ser Tyr Ser Ser
2480 2485 2490Pro Val Asp Asn Thr Pro Ser His Gln Leu Gln Val Pro
Glu His 2495 2500 2505Pro Phe Leu Thr Pro Ser Pro Glu Ser Pro Asp
Gln Trp Ser Ser 2510 2515 2520Ser Ser Pro His Ser Asn Val Ser Asp
Trp Ser Glu Gly Val Ser 2525 2530 2535Ser Pro Pro Thr Ser Met Gln
Ser Gln Ile Ala Arg Ile Pro Glu 2540 2545 2550Ala Phe Lys
2555370390PRTBos taurus 370Met Glu Trp Gly Tyr Leu Leu Glu Val Thr
Ser Leu Leu Ala Ala Leu1 5 10 15Ala Leu Leu Gln Arg Ser Ser Gly Ala
Ala Ala Ala Ser Ala Lys Glu 20 25 30Leu Ala Cys Gln Glu Ile Thr Val
Pro Leu Cys Lys Gly Ile Gly Tyr 35 40 45Asn Tyr Thr Tyr Met Pro Asn
Gln Phe Asn His Asp Thr Gln Asp Glu 50 55 60Ala Gly Leu Glu Val His
Gln Phe Trp Pro Leu Val Glu Ile Gln Cys65 70 75 80Ser Pro Asp Leu
Lys Phe Phe Leu Cys Ser Met Tyr Thr Pro Ile Cys 85 90 95Leu Glu Asp
Tyr Lys Lys Pro Leu Pro Pro Cys Arg Ser Val Cys Glu 100 105 110Arg
Ala Lys Ala Gly Cys Ala Pro Leu Met Arg Gln Tyr Gly Phe Ala 115 120
125Trp Pro Asp Arg Met Arg Cys Asp Arg Leu Pro Glu Gln Gly Asn Pro
130 135 140Asp Thr Leu Cys Met Asp Tyr Asn Arg Thr Asp Leu Thr Thr
Ala Ala145 150 155 160Ser Ser Val Asp Gly Asp Pro Val Ala Gly Ile
Cys Tyr Val Gly Asn 165 170 175Gln Ser Leu Asp Asn Leu Leu Gly Phe
Val Leu Ala Pro Leu Val Ile 180 185 190Tyr Leu Phe Ile Gly Thr Met
Phe Leu Leu Ala Gly Phe Val Ser Leu 195 200 205Phe Arg Ile Arg Ser
Val Ile Lys Gln Gln Gly Gly Pro Thr Lys Thr 210 215 220His Lys Leu
Glu Lys Leu Met Ile Arg Leu Gly Leu Phe Thr Val Leu225 230 235
240Tyr Thr Val Pro Ala Ala Val Val Val Ala Cys Leu Phe Tyr Glu Gln
245 250 255His Asn Arg Pro Arg Trp Glu Ala Thr His Asn Cys Pro Cys
Leu Arg 260 265 270Asp Leu Gln Pro Asp Gln Ala Arg Arg Pro Asp Tyr
Ala Val Phe Met 275 280 285Leu Lys Tyr Phe Met Cys Leu Val Val Gly
Ile Thr Ser Gly Val Trp 290 295 300Val Trp Ser Gly Lys Thr Leu Glu
Ser Trp Arg Ala Leu Cys Thr Arg305 310 315 320Cys Cys Trp Ala Ser
Lys Gly Ala Gly Ala Ala Gly Ala Gly Ala Ala 325 330 335Gly Gly Gly
Pro Gly Gly Gly Gly Pro Gly Ala Gly Gly Gly Gly Gly 340 345 350Pro
Gly Ala Gly Gly Ala Gly Ser Leu Tyr Ser Asp Val Ser Thr Gly 355 360
365Leu Thr Trp Arg Ser Gly Thr Ala Ser Ser Val Ser Tyr Pro Lys Gln
370 375 380Met Pro Leu Ser Gln Val385 390371178PRTHomo sapiens
371Lys Glu Lys Gln Gln Asn Ile Ser Pro Leu Val Arg Glu Arg Gly Pro1
5 10 15Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser Asn
Thr 20 25 30Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg
Lys Ile 35 40 45Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser Phe Leu
Ser Asn Leu 50 55 60His Leu Arg Asn Gly Glu Leu Val Ile His Glu Lys
Gly Phe Tyr Tyr65 70 75 80Ile Tyr Ser Gln Thr Tyr Phe Arg Phe Gln
Glu Glu Ile Lys Glu Asn 85 90 95Thr Lys Asn Asp Lys Gln Met Val Gln
Tyr Ile Tyr Lys Tyr Thr Ser 100 105 110Tyr Pro Asp Pro Ile Leu Leu
Met Lys Ser Ala Arg Asn Ser Cys Trp 115 120 125Ser Lys Asp Ala Glu
Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile 130 135 140Phe Glu Leu
Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu145 150 155
160His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe Leu
165 170 175Val Gly372329PRTHomo sapiens 372Met Leu Thr Leu Gln Thr
Trp Leu Val Gln Ala Leu Phe Ile Phe Leu1 5 10 15Thr Thr Glu Ser Thr
Gly Glu Leu Leu Asp Pro Cys Gly Tyr Ile Ser 20 25 30Pro Glu Ser Pro
Val Val Gln Leu His Ser Asn Phe Thr Ala Val Cys 35 40 45Val Leu Lys
Glu Lys Cys Met Asp Tyr Phe His Val Asn Ala Asn Tyr 50 55 60Ile Val
Trp Lys Thr Asn His Phe Thr Ile Pro Lys Glu Gln Tyr Thr65 70 75
80Ile Ile Asn Arg Thr Ala Ser Ser Val Thr Phe Thr Asp Ile Ala Ser
85 90 95Leu Asn Ile Gln Leu Thr Cys Asn Ile Leu Thr Phe Gly Gln Leu
Glu 100 105 110Gln Asn Val Tyr Gly Ile Thr Ile Ile Ser Gly Leu Pro
Pro Glu Lys 115 120 125Pro Lys Asn Leu Ser Cys Ile Val Asn Glu
Gly Lys Lys Met Arg Cys 130 135 140Glu Trp Asp Gly Gly Arg Glu Thr
His Leu Glu Thr Asn Phe Thr Leu145 150 155 160Lys Ser Glu Trp Ala
Thr His Lys Phe Ala Asp Cys Lys Ala Lys Arg 165 170 175Asp Thr Pro
Thr Ser Cys Thr Val Asp Tyr Ser Thr Val Tyr Phe Val 180 185 190Asn
Ile Glu Val Trp Val Glu Ala Glu Asn Ala Leu Gly Lys Val Thr 195 200
205Ser Asp His Ile Asn Phe Asp Pro Val Tyr Lys Val Lys Pro Asn Pro
210 215 220Pro His Asn Leu Ser Val Ile Asn Ser Glu Glu Leu Ser Ser
Ile Leu225 230 235 240Lys Leu Thr Trp Thr Asn Pro Ser Ile Lys Ser
Val Ile Ile Leu Lys 245 250 255Tyr Asn Ile Gln Tyr Arg Thr Lys Asp
Ala Ser Thr Trp Ser Gln Ile 260 265 270Pro Pro Glu Asp Thr Ala Ser
Thr Arg Ser Ser Phe Thr Val Gln Asp 275 280 285Leu Lys Pro Phe Thr
Glu Tyr Val Phe Arg Ile Arg Cys Met Lys Glu 290 295 300Asp Gly Lys
Gly Tyr Trp Ser Asp Trp Ser Glu Glu Ala Ser Gly Ile305 310 315
320Thr Tyr Glu Asp Asn Ile Ala Ser Phe 325373577PRTHomo sapiens
373Met Leu Pro Cys Leu Val Val Leu Leu Ala Ala Leu Leu Ser Leu Arg1
5 10 15Leu Gly Ser Asp Ala His Gly Thr Glu Leu Pro Ser Pro Pro Ser
Val 20 25 30Trp Phe Glu Ala Glu Phe Phe His His Ile Leu His Trp Thr
Pro Ile 35 40 45Pro Asn Gln Ser Glu Ser Thr Cys Tyr Glu Val Ala Leu
Leu Arg Tyr 50 55 60Gly Ile Glu Ser Trp Asn Ser Ile Ser Asn Cys Ser
Gln Thr Leu Ser65 70 75 80Tyr Asp Leu Thr Ala Val Thr Leu Asp Leu
Tyr His Ser Asn Gly Tyr 85 90 95Arg Ala Arg Val Arg Ala Val Asp Gly
Ser Arg His Ser Asn Trp Thr 100 105 110Val Thr Asn Thr Arg Phe Ser
Val Asp Glu Val Thr Leu Thr Val Gly 115 120 125Ser Val Asn Leu Glu
Ile His Asn Gly Phe Ile Leu Gly Lys Ile Gln 130 135 140Leu Pro Arg
Pro Lys Met Ala Pro Ala Asn Asp Thr Tyr Glu Ser Ile145 150 155
160Phe Ser His Phe Arg Glu Tyr Glu Ile Ala Ile Arg Lys Val Pro Gly
165 170 175Asn Phe Thr Phe Thr His Lys Lys Val Lys His Glu Asn Phe
Ser Leu 180 185 190Leu Thr Ser Gly Glu Val Gly Glu Phe Cys Val Gln
Val Lys Pro Ser 195 200 205Val Ala Ser Arg Ser Asn Lys Gly Met Trp
Ser Lys Glu Glu Cys Ile 210 215 220Ser Leu Thr Arg Gln Tyr Phe Thr
Val Thr Asn Val Ile Ile Phe Phe225 230 235 240Ala Phe Val Leu Leu
Leu Ser Gly Ala Leu Ala Tyr Cys Leu Ala Leu 245 250 255Gln Leu Tyr
Val Arg Arg Arg Lys Lys Leu Pro Ser Val Leu Leu Phe 260 265 270Lys
Lys Pro Ser Pro Phe Ile Phe Ile Ser Gln Arg Pro Ser Pro Glu 275 280
285Thr Gln Asp Thr Ile His Pro Leu Asp Glu Glu Ala Phe Leu Lys Val
290 295 300Ser Pro Glu Leu Lys Asn Leu Asp Leu His Gly Ser Thr Asp
Ser Gly305 310 315 320Phe Gly Ser Thr Lys Pro Ser Leu Gln Thr Glu
Glu Pro Gln Phe Leu 325 330 335Leu Pro Asp Pro His Pro Gln Ala Asp
Arg Thr Leu Gly Asn Arg Glu 340 345 350Pro Pro Val Leu Gly Asp Ser
Cys Ser Ser Gly Ser Ser Asn Ser Thr 355 360 365Asp Ser Gly Ile Cys
Leu Gln Glu Pro Ser Leu Ser Pro Ser Thr Gly 370 375 380Pro Thr Trp
Glu Gln Gln Val Gly Ser Asn Ser Arg Gly Gln Asp Asp385 390 395
400Ser Gly Ile Asp Leu Val Gln Asn Ser Glu Gly Arg Ala Gly Asp Thr
405 410 415Gln Gly Gly Ser Ala Leu Gly His His Ser Pro Pro Glu Pro
Glu Val 420 425 430Pro Gly Glu Glu Asp Pro Ala Ala Val Ala Phe Gln
Gly Tyr Leu Arg 435 440 445Gln Thr Arg Cys Ala Glu Glu Lys Ala Thr
Lys Thr Gly Cys Leu Glu 450 455 460Glu Glu Ser Pro Leu Thr Asp Gly
Leu Gly Pro Lys Phe Gly Arg Cys465 470 475 480Leu Val Asp Glu Ala
Gly Leu His Pro Pro Ala Leu Ala Lys Gly Tyr 485 490 495Leu Lys Gln
Asp Pro Leu Glu Met Thr Leu Ala Ser Ser Gly Ala Pro 500 505 510Thr
Gly Gln Trp Asn Gln Pro Thr Glu Glu Trp Ser Leu Leu Ala Leu 515 520
525Ser Ser Cys Ser Asp Leu Gly Ile Ser Asp Trp Ser Phe Ala His Asp
530 535 540Leu Ala Pro Leu Gly Cys Val Ala Ala Pro Gly Gly Leu Leu
Gly Ser545 550 555 560Phe Asn Ser Asp Leu Val Thr Leu Pro Leu Ile
Ser Ser Leu Gln Ser 565 570 575Ser374656PRTDrosophila melanogaster
374Lys Ile Cys Ile Gly Thr Lys Ser Arg Leu Ser Val Pro Ser Asn Lys1
5 10 15Glu His His Tyr Arg Asn Leu Arg Asp Arg Tyr Thr Asn Cys Thr
Tyr 20 25 30Val Asp Gly Asn Leu Glu Leu Thr Trp Leu Pro Asn Glu Asn
Leu Asp 35 40 45Leu Ser Phe Leu Asp Asn Ile Arg Glu Val Thr Gly Tyr
Ile Leu Ile 50 55 60Ser His Val Asp Val Lys Lys Val Val Phe Pro Lys
Leu Gln Ile Ile65 70 75 80Arg Gly Arg Thr Leu Phe Ser Leu Ser Val
Glu Glu Glu Lys Tyr Ala 85 90 95Leu Phe Val Thr Tyr Ser Lys Met Tyr
Thr Leu Glu Ile Pro Asp Leu 100 105 110Arg Asp Val Leu Asn Gly Gln
Val Gly Phe His Asn Asn Tyr Asn Leu 115 120 125Cys His Met Arg Thr
Ile Gln Trp Ser Glu Ile Val Ser Asn Gly Thr 130 135 140Asp Ala Tyr
Tyr Asn Tyr Asp Phe Thr Ala Pro Glu Arg Glu Cys Pro145 150 155
160Lys Cys His Glu Ser Cys Thr His Gly Cys Trp Gly Glu Gly Pro Lys
165 170 175Asn Cys Gln Lys Phe Ser Lys Leu Thr Cys Ser Pro Gln Cys
Ala Gly 180 185 190Gly Arg Cys Tyr Gly Pro Lys Pro Arg Glu Cys Cys
His Leu Phe Cys 195 200 205Ala Gly Gly Cys Thr Gly Pro Thr Gln Lys
Asp Cys Ile Ala Cys Lys 210 215 220Asn Phe Phe Asp Glu Gly Val Cys
Lys Glu Glu Cys Pro Pro Met Arg225 230 235 240Lys Tyr Asn Pro Thr
Thr Tyr Val Leu Glu Thr Asn Pro Glu Gly Lys 245 250 255Tyr Ala Tyr
Gly Ala Thr Cys Val Lys Glu Cys Pro Gly His Leu Leu 260 265 270Arg
Asp Asn Gly Ala Cys Val Arg Ser Cys Pro Gln Asp Lys Met Asp 275 280
285Lys Gly Gly Glu Cys Val Pro Cys Asn Gly Pro Cys Pro Lys Thr Cys
290 295 300Pro Gly Val Thr Val Leu His Ala Gly Asn Ile Asp Ser Phe
Arg Asn305 310 315 320Cys Thr Val Ile Asp Gly Asn Ile Arg Ile Leu
Asp Gln Thr Phe Ser 325 330 335Gly Phe Gln Asp Val Tyr Ala Asn Tyr
Thr Met Gly Pro Arg Tyr Ile 340 345 350Pro Leu Asp Pro Glu Arg Leu
Glu Val Phe Ser Thr Val Lys Glu Ile 355 360 365Thr Gly Tyr Leu Asn
Ile Glu Gly Thr His Pro Gln Phe Arg Asn Leu 370 375 380Ser Tyr Phe
Arg Asn Leu Glu Thr Ile His Gly Arg Gln Leu Met Glu385 390 395
400Ser Met Phe Ala Ala Leu Ala Ile Val Lys Ser Ser Leu Tyr Ser Leu
405 410 415Glu Met Arg Asn Leu Lys Gln Ile Ser Ser Gly Ser Val Val
Ile Gln 420 425 430His Asn Arg Asp Leu Cys Tyr Val Ser Asn Ile Arg
Trp Pro Ala Ile 435 440 445Gln Lys Glu Pro Glu Gln Lys Val Trp Val
Asn Glu Asn Leu Arg Ala 450 455 460Asp Leu Cys Glu Lys Asn Gly Thr
Ile Cys Ser Asp Gln Cys Asn Glu465 470 475 480Asp Gly Cys Trp Gly
Ala Gly Thr Asp Gln Cys Leu Asn Cys Lys Asn 485 490 495Phe Asn Phe
Asn Gly Thr Cys Ile Ala Asp Cys Gly Tyr Ile Ser Asn 500 505 510Ala
Tyr Lys Phe Asp Asn Arg Thr Cys Lys Ile Cys His Pro Glu Cys 515 520
525Arg Thr Cys Asn Gly Ala Gly Ala Asp His Cys Gln Glu Cys Val His
530 535 540Val Arg Asp Gly Gln His Cys Val Ser Glu Cys Pro Lys Asn
Lys Tyr545 550 555 560Asn Asp Arg Gly Val Cys Arg Glu Cys His Ala
Thr Cys Asp Gly Cys 565 570 575Thr Gly Pro Lys Asp Thr Ile Gly Ile
Gly Ala Cys Thr Thr Cys Asn 580 585 590Leu Ala Ile Ile Asn Asn Asp
Ala Thr Val Lys Arg Cys Leu Leu Lys 595 600 605Asp Asp Lys Cys Pro
Asp Gly Tyr Phe Trp Glu Tyr Val His Pro Gln 610 615 620Glu Gln Gly
Ser Leu Lys Pro Leu Ala Gly Arg Ala Val Cys Arg Lys625 630 635
640Cys His Pro Leu Cys Glu Leu Cys Thr Asn Tyr Gly Tyr His Glu Gln
645 650 655375326PRTHomo sapiens 375Met Glu Lys Asp Gly Leu Cys Arg
Ala Asp Gln Gln Tyr Glu Cys Val1 5 10 15Ala Glu Ile Gly Glu Gly Ala
Tyr Gly Lys Val Phe Lys Ala Arg Asp 20 25 30Leu Lys Asn Gly Gly Arg
Phe Val Ala Leu Lys Arg Val Arg Val Gln 35 40 45Thr Gly Glu Glu Gly
Met Pro Leu Ser Thr Ile Arg Glu Val Ala Val 50 55 60Leu Arg His Leu
Glu Thr Phe Glu His Pro Asn Val Val Arg Leu Phe65 70 75 80Asp Val
Cys Thr Val Ser Arg Thr Asp Arg Glu Thr Lys Leu Thr Leu 85 90 95Val
Phe Glu His Val Asp Gln Asp Leu Thr Thr Tyr Leu Asp Lys Val 100 105
110Pro Glu Pro Gly Val Pro Thr Glu Thr Ile Lys Asp Met Met Phe Gln
115 120 125Leu Leu Arg Gly Leu Asp Phe Leu His Ser His Arg Val Val
His Arg 130 135 140Asp Leu Lys Pro Gln Asn Ile Leu Val Thr Ser Ser
Gly Gln Ile Lys145 150 155 160Leu Ala Asp Phe Gly Leu Ala Arg Ile
Tyr Ser Phe Gln Met Ala Leu 165 170 175Thr Ser Val Val Val Thr Leu
Trp Tyr Arg Ala Pro Glu Val Leu Leu 180 185 190Gln Ser Ser Tyr Ala
Thr Pro Val Asp Leu Trp Ser Val Gly Cys Ile 195 200 205Phe Ala Glu
Met Phe Arg Arg Lys Pro Leu Phe Arg Gly Ser Ser Asp 210 215 220Val
Asp Gln Leu Gly Lys Ile Leu Asp Val Ile Gly Leu Pro Gly Glu225 230
235 240Glu Asp Trp Pro Arg Asp Val Ala Leu Pro Arg Gln Ala Phe His
Ser 245 250 255Lys Ser Ala Gln Pro Ile Glu Lys Phe Val Thr Asp Ile
Asp Glu Leu 260 265 270Gly Lys Asp Leu Leu Leu Lys Cys Leu Thr Phe
Asn Pro Ala Lys Arg 275 280 285Ile Ser Ala Tyr Ser Ala Leu Ser His
Pro Tyr Phe Gln Asp Leu Glu 290 295 300Arg Cys Lys Glu Asn Leu Asp
Ser His Leu Pro Pro Ser Gln Asn Thr305 310 315 320Ser Glu Leu Asn
Thr Ala 325376119PRTParamecium bursaria Chlorella virus 1 376Met
Phe Asn Asp Arg Val Ile Val Lys Lys Ser Pro Leu Gly Gly Tyr1 5 10
15Gly Val Phe Ala Arg Lys Ser Phe Glu Lys Gly Glu Leu Val Glu Glu
20 25 30Cys Leu Cys Ile Val Arg His Asn Asp Asp Trp Gly Thr Ala Leu
Glu 35 40 45Asp Tyr Leu Phe Ser Arg Lys Asn Met Ser Ala Met Ala Leu
Gly Phe 50 55 60Gly Ala Ile Phe Asn His Ser Lys Asp Pro Asn Ala Arg
His Glu Leu65 70 75 80Thr Ala Gly Leu Lys Arg Met Arg Ile Phe Thr
Ile Lys Pro Ile Ala 85 90 95Ile Gly Glu Glu Ile Thr Ile Ser Tyr Gly
Asp Asp Tyr Trp Leu Ser 100 105 110Arg Pro Arg Leu Thr Gln Asn
115377293PRTHomo sapiens 377Met Trp Pro Leu Val Ala Ala Leu Leu Leu
Gly Ser Ala Cys Cys Gly1 5 10 15Ser Ala Gln Leu Leu Phe Asn Lys Thr
Lys Ser Val Glu Phe Thr Phe 20 25 30Cys Asn Asp Thr Val Val Ile Pro
Cys Phe Val Thr Asn Met Glu Ala 35 40 45Gln Asn Thr Thr Glu Val Tyr
Val Lys Trp Lys Phe Lys Gly Arg Asp 50 55 60Ile Tyr Thr Phe Asp Gly
Ala Leu Asn Lys Ser Thr Val Pro Thr Asp65 70 75 80Phe Ser Ser Ala
Lys Ile Glu Val Ser Gln Leu Leu Lys Gly Asp Ala 85 90 95Ser Leu Lys
Met Asp Lys Ser Asp Ala Val Ser His Thr Gly Asn Tyr 100 105 110Thr
Cys Glu Val Thr Glu Leu Thr Arg Glu Gly Glu Thr Ile Ile Glu 115 120
125Leu Lys Tyr Arg Val Val Ser Trp Phe Ser Pro Asn Glu Asn Ile Leu
130 135 140Ile Val Ile Phe Pro Ile Phe Ala Ile Leu Leu Phe Trp Gly
Gln Phe145 150 155 160Gly Ile Lys Thr Leu Lys Tyr Arg Ser Gly Gly
Met Asp Glu Lys Thr 165 170 175Ile Ala Leu Leu Val Ala Gly Leu Val
Ile Thr Val Ile Val Ile Val 180 185 190Gly Ala Ile Leu Phe Val Pro
Gly Glu Tyr Ser Leu Lys Asn Ala Thr 195 200 205Gly Leu Gly Leu Ile
Val Thr Ser Thr Gly Ile Leu Ile Leu Leu His 210 215 220Tyr Tyr Val
Phe Ser Thr Ala Ile Gly Leu Thr Ser Phe Val Ile Ala225 230 235
240Ile Leu Val Ile Gln Val Ile Ala Tyr Ile Leu Ala Val Val Gly Leu
245 250 255Ser Leu Cys Ile Ala Ala Cys Ile Pro Met His Gly Pro Leu
Leu Ile 260 265 270Ser Gly Leu Ser Ile Leu Ala Leu Ala Gln Leu Leu
Gly Leu Val Tyr 275 280 285Met Lys Phe Val Glu 290378507PRTHomo
sapiens 378Met Glu Pro Ala Gly Pro Ala Pro Gly Arg Leu Gly Pro Leu
Leu Cys1 5 10 15Leu Leu Leu Ala Ala Ser Cys Ala Trp Ser Gly Val Ala
Gly Glu Glu 20 25 30Glu Leu Gln Val Ile Gln Pro Asp Lys Ser Val Ser
Val Ala Ala Gly 35 40 45Glu Ser Ala Ile Leu His Cys Thr Val Thr Ser
Leu Ile Pro Val Gly 50 55 60Pro Ile Gln Trp Phe Arg Gly Ala Gly Pro
Ala Arg Glu Leu Ile Tyr65 70 75 80Asn Gln Lys Glu Gly His Phe Pro
Arg Val Thr Thr Val Ser Glu Ser 85 90 95Thr Lys Arg Glu Asn Met Asp
Phe Ser Ile Ser Ile Ser Asn Ile Thr 100 105 110Pro Ala Asp Ala Gly
Thr Tyr Tyr Cys Val Lys Phe Arg Lys Gly Ser 115 120 125Pro Asp Thr
Glu Phe Lys Ser Gly Ala Gly Thr Glu Leu Ser Val Arg 130 135 140Ala
Lys Pro Ser Ala Pro Val Val Ser Gly Pro Ala Ala Arg Ala Thr145 150
155 160Pro Gln His Thr Val Ser Phe Thr Cys Glu Ser His Gly Phe Ser
Pro 165 170 175Arg Asp Ile Thr Leu Lys Trp Phe Lys Asn Gly Asn Glu
Leu Ser Asp 180 185 190Phe Gln Thr Asn Val Asp Pro Val Gly Glu Ser
Val Ser Tyr Ser Ile 195 200 205His Ser Thr Ala Lys Val Val Leu Thr
Arg Glu Asp Val His Ser Gln 210 215 220Val Ile Cys Glu Val Ala His
Val Thr Leu Gln Gly Asp Pro Leu Arg225 230 235 240Gly Thr Ala Asn
Leu Ser Glu Thr Ile Arg Val Pro Pro Thr Leu Glu 245 250 255Val Thr
Gln Gln Pro Val Arg Ala Glu Asn Gln Val Asn Val Thr Cys 260 265
270Gln Val Arg Lys Phe Tyr Pro Gln Arg Leu Gln Leu Thr Trp Leu Glu
275 280 285Asn Gly
Asn Val Ser Arg Thr Glu Thr Ala Ser Thr Val Thr Glu Asn 290 295
300Lys Asp Gly Thr Tyr Asn Trp Met Ser Trp Leu Leu Val Asn Val
Ser305 310 315 320Ala His Arg Asp Asp Val Lys Leu Thr Cys Gln Val
Glu His Asp Gly 325 330 335Gln Pro Ala Val Ser Lys Ser His Asp Leu
Lys Val Ser Ala His Pro 340 345 350Lys Glu Gln Gly Ser Asn Thr Ala
Ala Glu Asn Thr Gly Ser Asn Glu 355 360 365Arg Asn Ile Tyr Ile Val
Val Gly Val Val Cys Thr Leu Leu Val Ala 370 375 380Leu Leu Met Ala
Ala Leu Tyr Leu Val Arg Ile Arg Gln Lys Lys Ala385 390 395 400Gln
Gly Ser Thr Ser Ser Thr Arg Leu His Glu Pro Glu Lys Asn Ala 405 410
415Arg Glu Ile Thr Gln Val Gln Ser Leu Asp Thr Asn Asp Ile Thr Tyr
420 425 430Ala Asp Leu Asn Leu Pro Lys Gly Lys Lys Pro Ala Pro Gln
Ala Ala 435 440 445Glu Pro Asn Asn His Thr Glu Tyr Ala Ser Ile Gln
Thr Ser Pro Gln 450 455 460Pro Ala Ser Glu Asp Thr Leu Thr Tyr Ala
Asp Leu Asp Met Val His465 470 475 480Leu Asn Arg Thr Pro Lys Gln
Pro Ala Pro Lys Pro Glu Pro Ser Phe 485 490 495Ser Glu Tyr Ala Ser
Val Gln Val Pro Arg Lys 500 5053791456PRTHomo sapiens 379Met Arg
Leu Pro Leu Leu Leu Val Phe Ala Ser Val Ile Pro Gly Ala1 5 10 15Val
Leu Leu Leu Asp Thr Arg Gln Phe Leu Ile Tyr Asn Glu Asp His 20 25
30Lys Arg Cys Val Asp Ala Val Ser Pro Ser Ala Val Gln Thr Ala Ala
35 40 45Cys Asn Gln Asp Ala Glu Ser Gln Lys Phe Arg Trp Val Ser Glu
Ser 50 55 60Gln Ile Met Ser Val Ala Phe Lys Leu Cys Leu Gly Val Pro
Ser Lys65 70 75 80Thr Asp Trp Val Ala Ile Thr Leu Tyr Ala Cys Asp
Ser Lys Ser Glu 85 90 95Phe Gln Lys Trp Glu Cys Lys Asn Asp Thr Leu
Leu Gly Ile Lys Gly 100 105 110Glu Asp Leu Phe Phe Asn Tyr Gly Asn
Arg Gln Glu Lys Asn Ile Met 115 120 125Leu Tyr Lys Gly Ser Gly Leu
Trp Ser Arg Trp Lys Ile Tyr Gly Thr 130 135 140Thr Asp Asn Leu Cys
Ser Arg Gly Tyr Glu Ala Met Tyr Thr Leu Leu145 150 155 160Gly Asn
Ala Asn Gly Ala Thr Cys Ala Phe Pro Phe Lys Phe Glu Asn 165 170
175Lys Trp Tyr Ala Asp Cys Thr Ser Ala Gly Arg Ser Asp Gly Trp Leu
180 185 190Trp Cys Gly Thr Thr Thr Asp Tyr Asp Thr Asp Lys Leu Phe
Gly Tyr 195 200 205Cys Pro Leu Lys Phe Glu Gly Ser Glu Ser Leu Trp
Asn Lys Asp Pro 210 215 220Leu Thr Ser Val Ser Tyr Gln Ile Asn Ser
Lys Ser Ala Leu Thr Trp225 230 235 240His Gln Ala Arg Lys Ser Cys
Gln Gln Gln Asn Ala Glu Leu Leu Ser 245 250 255Ile Thr Glu Ile His
Glu Gln Thr Tyr Leu Thr Gly Leu Thr Ser Ser 260 265 270Leu Thr Ser
Gly Leu Trp Ile Gly Leu Asn Ser Leu Ser Phe Asn Ser 275 280 285Gly
Trp Gln Trp Ser Asp Arg Ser Pro Phe Arg Tyr Leu Asn Trp Leu 290 295
300Pro Gly Ser Pro Ser Ala Glu Pro Gly Lys Ser Cys Val Ser Leu
Asn305 310 315 320Pro Gly Lys Asn Ala Lys Trp Glu Asn Leu Glu Cys
Val Gln Lys Leu 325 330 335Gly Tyr Ile Cys Lys Lys Gly Asn Thr Thr
Leu Asn Ser Phe Val Ile 340 345 350Pro Ser Glu Ser Asp Val Pro Thr
His Cys Pro Ser Gln Trp Trp Pro 355 360 365Tyr Ala Gly His Cys Tyr
Lys Ile His Arg Asp Glu Lys Lys Ile Gln 370 375 380Arg Asp Ala Leu
Thr Thr Cys Arg Lys Glu Gly Gly Asp Leu Thr Ser385 390 395 400Ile
His Thr Ile Glu Glu Leu Asp Phe Ile Ile Ser Gln Leu Gly Tyr 405 410
415Glu Pro Asn Asp Glu Leu Trp Ile Gly Leu Asn Asp Ile Lys Ile Gln
420 425 430Met Tyr Phe Glu Trp Ser Asp Gly Thr Pro Val Thr Phe Thr
Lys Trp 435 440 445Leu Arg Gly Glu Pro Ser His Glu Asn Asn Arg Gln
Glu Asp Cys Val 450 455 460Val Met Lys Gly Lys Asp Gly Tyr Trp Ala
Asp Arg Gly Cys Glu Trp465 470 475 480Pro Leu Gly Tyr Ile Cys Lys
Met Lys Ser Arg Ser Gln Gly Pro Glu 485 490 495Ile Val Glu Val Glu
Lys Gly Cys Arg Lys Gly Trp Lys Lys His His 500 505 510Phe Tyr Cys
Tyr Met Ile Gly His Thr Leu Ser Thr Phe Ala Glu Ala 515 520 525Asn
Gln Thr Cys Asn Asn Glu Asn Ala Tyr Leu Thr Thr Ile Glu Asp 530 535
540Arg Tyr Glu Gln Ala Phe Leu Thr Ser Phe Val Gly Leu Arg Pro
Glu545 550 555 560Lys Tyr Phe Trp Thr Gly Leu Ser Asp Ile Gln Thr
Lys Gly Thr Phe 565 570 575Gln Trp Thr Ile Glu Glu Glu Val Arg Phe
Thr His Trp Asn Ser Asp 580 585 590Met Pro Gly Arg Lys Pro Gly Cys
Val Ala Met Arg Thr Gly Ile Ala 595 600 605Gly Gly Leu Trp Asp Val
Leu Lys Cys Asp Glu Lys Ala Lys Phe Val 610 615 620Cys Lys His Trp
Ala Glu Gly Val Thr His Pro Pro Lys Pro Thr Thr625 630 635 640Thr
Pro Glu Pro Lys Cys Pro Glu Asp Trp Gly Ala Ser Ser Arg Thr 645 650
655Ser Leu Cys Phe Lys Leu Tyr Ala Lys Gly Lys His Glu Lys Lys Thr
660 665 670Trp Phe Glu Ser Arg Asp Phe Cys Arg Ala Leu Gly Gly Asp
Leu Ala 675 680 685Ser Ile Asn Asn Lys Glu Glu Gln Gln Thr Ile Trp
Arg Leu Ile Thr 690 695 700Ala Ser Gly Ser Tyr His Lys Leu Phe Trp
Leu Gly Leu Thr Tyr Gly705 710 715 720Ser Pro Ser Glu Gly Phe Thr
Trp Ser Asp Gly Ser Pro Val Ser Tyr 725 730 735Glu Asn Trp Ala Tyr
Gly Glu Pro Asn Asn Tyr Gln Asn Val Glu Tyr 740 745 750Cys Gly Glu
Leu Lys Gly Asp Pro Thr Met Ser Trp Asn Asp Ile Asn 755 760 765Cys
Glu His Leu Asn Asn Trp Ile Cys Gln Ile Gln Lys Gly Gln Thr 770 775
780Pro Lys Pro Glu Pro Thr Pro Ala Pro Gln Asp Asn Pro Pro Val
Thr785 790 795 800Glu Asp Gly Trp Val Ile Tyr Lys Asp Tyr Gln Tyr
Tyr Phe Ser Lys 805 810 815Glu Lys Glu Thr Met Asp Asn Ala Arg Ala
Phe Cys Lys Arg Asn Phe 820 825 830Gly Asp Leu Val Ser Ile Gln Ser
Glu Ser Glu Lys Lys Phe Leu Trp 835 840 845Lys Tyr Val Asn Arg Asn
Asp Ala Gln Ser Ala Tyr Phe Ile Gly Leu 850 855 860Leu Ile Ser Leu
Asp Lys Lys Phe Ala Trp Met Asp Gly Ser Lys Val865 870 875 880Asp
Tyr Val Ser Trp Ala Thr Gly Glu Pro Asn Phe Ala Asn Glu Asp 885 890
895Glu Asn Cys Val Thr Met Tyr Ser Asn Ser Gly Phe Trp Asn Asp Ile
900 905 910Asn Cys Gly Tyr Pro Asn Ala Phe Ile Cys Gln Arg His Asn
Ser Ser 915 920 925Ile Asn Ala Thr Thr Val Met Pro Thr Met Pro Ser
Val Pro Ser Gly 930 935 940Cys Lys Glu Gly Trp Asn Phe Tyr Ser Asn
Lys Cys Phe Lys Ile Phe945 950 955 960Gly Phe Met Glu Glu Glu Arg
Lys Asn Trp Gln Glu Ala Arg Lys Ala 965 970 975Cys Ile Gly Phe Gly
Gly Asn Leu Val Ser Ile Gln Asn Glu Lys Glu 980 985 990Gln Ala Phe
Leu Thr Tyr His Met Lys Asp Ser Thr Phe Ser Ala Trp 995 1000
1005Thr Gly Leu Asn Asp Val Asn Ser Glu His Thr Phe Leu Trp Thr
1010 1015 1020Asp Gly Arg Gly Val His Tyr Thr Asn Trp Gly Lys Gly
Tyr Pro 1025 1030 1035Gly Gly Arg Arg Ser Ser Leu Ser Tyr Glu Asp
Ala Asp Cys Val 1040 1045 1050Val Ile Ile Gly Gly Ala Ser Asn Glu
Ala Gly Lys Trp Met Asp 1055 1060 1065Asp Thr Cys Asp Ser Lys Arg
Gly Tyr Ile Cys Gln Thr Arg Ser 1070 1075 1080Asp Pro Ser Leu Thr
Asn Pro Pro Ala Thr Ile Gln Thr Asp Gly 1085 1090 1095Phe Val Lys
Tyr Gly Lys Ser Ser Tyr Ser Leu Met Arg Gln Lys 1100 1105 1110Phe
Gln Trp His Glu Ala Glu Thr Tyr Cys Lys Leu His Asn Ser 1115 1120
1125Leu Ile Ala Ser Ile Leu Asp Pro Tyr Ser Asn Ala Phe Ala Trp
1130 1135 1140Leu Gln Met Glu Thr Ser Asn Glu Arg Val Trp Ile Ala
Leu Asn 1145 1150 1155Ser Asn Leu Thr Asp Asn Gln Tyr Thr Trp Thr
Asp Lys Trp Arg 1160 1165 1170Val Arg Tyr Thr Asn Trp Ala Ala Asp
Glu Pro Lys Leu Lys Ser 1175 1180 1185Ala Cys Val Tyr Leu Asp Leu
Asp Gly Tyr Trp Lys Thr Ala His 1190 1195 1200Cys Asn Glu Ser Phe
Tyr Phe Leu Cys Lys Arg Ser Asp Glu Ile 1205 1210 1215Pro Ala Thr
Glu Pro Pro Gln Leu Pro Gly Arg Cys Pro Glu Ser 1220 1225 1230Asp
His Thr Ala Trp Ile Pro Phe His Gly His Cys Tyr Tyr Ile 1235 1240
1245Glu Ser Ser Tyr Thr Arg Asn Trp Gly Gln Ala Ser Leu Glu Cys
1250 1255 1260Leu Arg Met Gly Ser Ser Leu Val Ser Ile Glu Ser Ala
Ala Glu 1265 1270 1275Ser Ser Phe Leu Ser Tyr Arg Val Glu Pro Leu
Lys Ser Lys Thr 1280 1285 1290Asn Phe Trp Ile Gly Leu Phe Arg Asn
Val Glu Gly Thr Trp Leu 1295 1300 1305Trp Ile Asn Asn Ser Pro Val
Ser Phe Val Asn Trp Asn Thr Gly 1310 1315 1320Asp Pro Ser Gly Glu
Arg Asn Asp Cys Val Ala Leu His Ala Ser 1325 1330 1335Ser Gly Phe
Trp Ser Asn Ile His Cys Ser Ser Tyr Lys Gly Tyr 1340 1345 1350Ile
Cys Lys Arg Pro Lys Ile Ile Asp Ala Lys Pro Thr His Glu 1355 1360
1365Leu Leu Thr Thr Lys Ala Asp Thr Arg Lys Met Asp Pro Ser Lys
1370 1375 1380Pro Ser Ser Asn Val Ala Gly Val Val Ile Ile Val Ile
Leu Leu 1385 1390 1395Ile Leu Thr Gly Ala Gly Leu Ala Ala Tyr Phe
Phe Tyr Lys Lys 1400 1405 1410Arg Arg Val His Leu Pro Gln Glu Gly
Ala Phe Glu Asn Thr Leu 1415 1420 1425Tyr Phe Asn Ser Gln Ser Ser
Pro Gly Thr Ser Asp Met Lys Asp 1430 1435 1440Leu Val Gly Asn Ile
Glu Gln Asn Glu His Ser Val Ile 1445 1450 1455380684PRTHomo sapiens
380Met Met Asp Ala Ser Lys Glu Leu Gln Val Leu His Ile Asp Phe Leu1
5 10 15Asn Gln Asp Asn Ala Val Ser His His Thr Trp Glu Phe Gln Thr
Ser 20 25 30Ser Pro Val Phe Arg Arg Gly Gln Val Phe His Leu Arg Leu
Val Leu 35 40 45Asn Gln Pro Leu Gln Ser Tyr His Gln Leu Lys Leu Glu
Phe Ser Thr 50 55 60Gly Pro Asn Pro Ser Ile Ala Lys His Thr Leu Val
Val Leu Asp Pro65 70 75 80Arg Thr Pro Ser Asp His Tyr Asn Trp Gln
Ala Thr Leu Gln Asn Glu 85 90 95Ser Gly Lys Glu Val Thr Val Ala Val
Thr Ser Ser Pro Asn Ala Ile 100 105 110Leu Gly Lys Tyr Gln Leu Asn
Val Lys Thr Gly Asn His Ile Leu Lys 115 120 125Ser Glu Glu Asn Ile
Leu Tyr Leu Leu Phe Asn Pro Trp Cys Lys Glu 130 135 140Asp Met Val
Phe Met Pro Asp Glu Asp Glu Arg Lys Glu Tyr Ile Leu145 150 155
160Asn Asp Thr Gly Cys His Tyr Val Gly Ala Ala Arg Ser Ile Lys Cys
165 170 175Lys Pro Trp Asn Phe Gly Gln Phe Glu Lys Asn Val Leu Asp
Cys Cys 180 185 190Ile Ser Leu Leu Thr Glu Ser Ser Leu Lys Pro Thr
Asp Arg Arg Asp 195 200 205Pro Val Leu Val Cys Arg Ala Met Cys Ala
Met Met Ser Phe Glu Lys 210 215 220Gly Gln Gly Val Leu Ile Gly Asn
Trp Thr Gly Asp Tyr Glu Gly Gly225 230 235 240Thr Ala Pro Tyr Lys
Trp Thr Gly Ser Ala Pro Ile Leu Gln Gln Tyr 245 250 255Tyr Asn Thr
Lys Gln Ala Val Cys Phe Gly Gln Cys Trp Val Phe Ala 260 265 270Gly
Ile Leu Thr Thr Val Leu Arg Ala Leu Gly Ile Pro Ala Arg Ser 275 280
285Val Thr Gly Phe Asp Ser Ala His Asp Thr Glu Arg Asn Leu Thr Val
290 295 300Asp Thr Tyr Val Asn Glu Asn Gly Glu Lys Ile Thr Ser Met
Thr His305 310 315 320Asp Ser Val Trp Asn Phe His Val Trp Thr Asp
Ala Trp Met Lys Arg 325 330 335Pro Asp Leu Pro Lys Gly Tyr Asp Gly
Trp Gln Ala Val Asp Ala Thr 340 345 350Pro Gln Glu Arg Ser Gln Gly
Val Phe Cys Cys Gly Pro Ser Pro Leu 355 360 365Thr Ala Ile Arg Lys
Gly Asp Ile Phe Ile Val Tyr Asp Thr Arg Phe 370 375 380Val Phe Ser
Glu Val Asn Gly Asp Arg Leu Ile Trp Leu Val Lys Met385 390 395
400Val Asn Gly Gln Glu Glu Leu His Val Ile Ser Met Glu Thr Thr Ser
405 410 415Ile Gly Lys Asn Ile Ser Thr Lys Ala Val Gly Gln Asp Arg
Arg Arg 420 425 430Asp Ile Thr Tyr Glu Tyr Lys Tyr Pro Glu Gly Ser
Ser Glu Glu Arg 435 440 445Gln Val Met Asp His Ala Phe Leu Leu Leu
Ser Ser Glu Arg Glu His 450 455 460Arg Arg Pro Val Lys Glu Asn Phe
Leu His Met Ser Val Gln Ser Asp465 470 475 480Asp Val Leu Leu Gly
Asn Ser Val Asn Phe Thr Val Ile Leu Lys Arg 485 490 495Lys Thr Ala
Ala Leu Gln Asn Val Asn Ile Leu Gly Ser Phe Glu Leu 500 505 510Gln
Leu Tyr Thr Gly Lys Lys Met Ala Lys Leu Cys Asp Leu Asn Lys 515 520
525Thr Ser Gln Ile Gln Gly Gln Val Ser Glu Val Thr Leu Thr Leu Asp
530 535 540Ser Lys Thr Tyr Ile Asn Ser Leu Ala Ile Leu Asp Asp Glu
Pro Val545 550 555 560Ile Arg Gly Phe Ile Ile Ala Glu Ile Val Glu
Ser Lys Glu Ile Met 565 570 575Ala Ser Glu Val Phe Thr Ser Phe Gln
Tyr Pro Glu Phe Ser Ile Glu 580 585 590Leu Pro Asn Thr Gly Arg Ile
Gly Gln Leu Leu Val Cys Asn Cys Ile 595 600 605Phe Lys Asn Thr Leu
Ala Ile Pro Leu Thr Asp Val Lys Phe Ser Leu 610 615 620Glu Ser Leu
Gly Ile Ser Ser Leu Gln Thr Ser Asp His Gly Thr Val625 630 635
640Gln Pro Gly Glu Thr Ile Gln Ser Gln Ile Lys Cys Thr Pro Ile Lys
645 650 655Thr Gly Pro Lys Lys Phe Ile Val Lys Leu Ser Ser Lys Gln
Val Lys 660 665 670Glu Ile Asn Ala Gln Lys Ile Val Leu Ile Thr Lys
675 680381585PRTHomo sapiens 381Asp Ala His Lys Ser Glu Val Ala His
Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu
Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Cys Pro Phe Glu Asp His
Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val
Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu
Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr
Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg
Asn Glu Cys Phe Leu
Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg
Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu
Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg
His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150
155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala
Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly
Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu
Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala
Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val
Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu
Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala
Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265
270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His
275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu
Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys
Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met
Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val
Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu
Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala
Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln
Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390
395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val
Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn
Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala
Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu
Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp
Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg
Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val
Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505
510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala
515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu
Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu
Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala
Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu
Gly Leu 580 58538210PRTArtificial SequenceSynthetic peptide 382Ile
Glu Ser Lys Arg Arg Lys Lys Lys Pro1 5 1038310PRTArtificial
SequenceSynthetic peptide 383Ala Pro Gly Pro Gly Asp Gly Gly Thr
Ala1 5 1038411PRTArtificial SequenceSynthetic peptide 384Phe Ala
Lys Lys Phe Ala Lys Lys Phe Lys Cys1 5 1038511PRTArtificial
SequenceSynthetic peptide 385Lys Phe Arg Lys Ala Phe Lys Arg Phe
Phe Cys1 5 103864PRTArtificial SequenceSynthetic peptide 386Lys Phe
Phe Lys13874PRTArtificial SequenceSynthetic peptide 387Lys Trp Trp
Lys13884PRTArtificial SequenceSynthetic peptide 388Lys Tyr Tyr
Lys13894PRTArtificial SequenceSynthetic peptide 389Lys Phe Trp
Lys13904PRTArtificial SequenceSynthetic peptide 390Lys Trp Phe
Lys13914PRTArtificial SequenceSynthetic peptide 391Lys Phe Tyr
Lys13924PRTArtificial SequenceSynthetic peptide 392Lys Tyr Phe
Lys13934PRTArtificial SequenceSynthetic peptide 393Lys Trp Tyr
Lys13944PRTArtificial SequenceSynthetic peptide 394Lys Tyr Trp
Lys13954PRTArtificial SequenceSynthetic peptide 395Arg Phe Phe
Arg13964PRTArtificial SequenceSynthetic peptide 396Arg Trp Trp
Arg13974PRTArtificial SequenceSynthetic peptide 397Arg Tyr Tyr
Arg13984PRTArtificial SequenceSynthetic peptide 398Arg Phe Trp
Arg13994PRTArtificial SequenceSynthetic peptide 399Arg Trp Phe
Arg14004PRTArtificial SequenceSynthetic peptide 400Arg Phe Tyr
Arg14014PRTArtificial SequenceSynthetic peptide 401Arg Tyr Phe
Arg14024PRTArtificial SequenceSynthetic peptide 402Arg Trp Tyr
Arg14034PRTArtificial SequenceSynthetic peptide 403Arg Tyr Trp
Arg14044PRTArtificial SequenceSynthetic peptide 404His Phe Phe
His14054PRTArtificial SequenceSynthetic peptide 405His Trp Trp
His14064PRTArtificial SequenceSynthetic peptide 406His Tyr Tyr
His14074PRTArtificial SequenceSynthetic peptide 407His Phe Trp
His14084PRTArtificial SequenceSynthetic peptide 408His Trp Phe
His14094PRTArtificial SequenceSynthetic peptide 409His Phe Tyr
His14104PRTArtificial SequenceSynthetic peptide 410His Tyr Phe
His14114PRTArtificial SequenceSynthetic peptide 411His Trp Tyr
His14124PRTArtificial SequenceSynthetic peptide 412Gly Gly Gly
Arg14134PRTArtificial SequenceSynthetic peptide 413Gly Pro Gly
Arg1















D00001

D00002

D00003

D00004

D00005
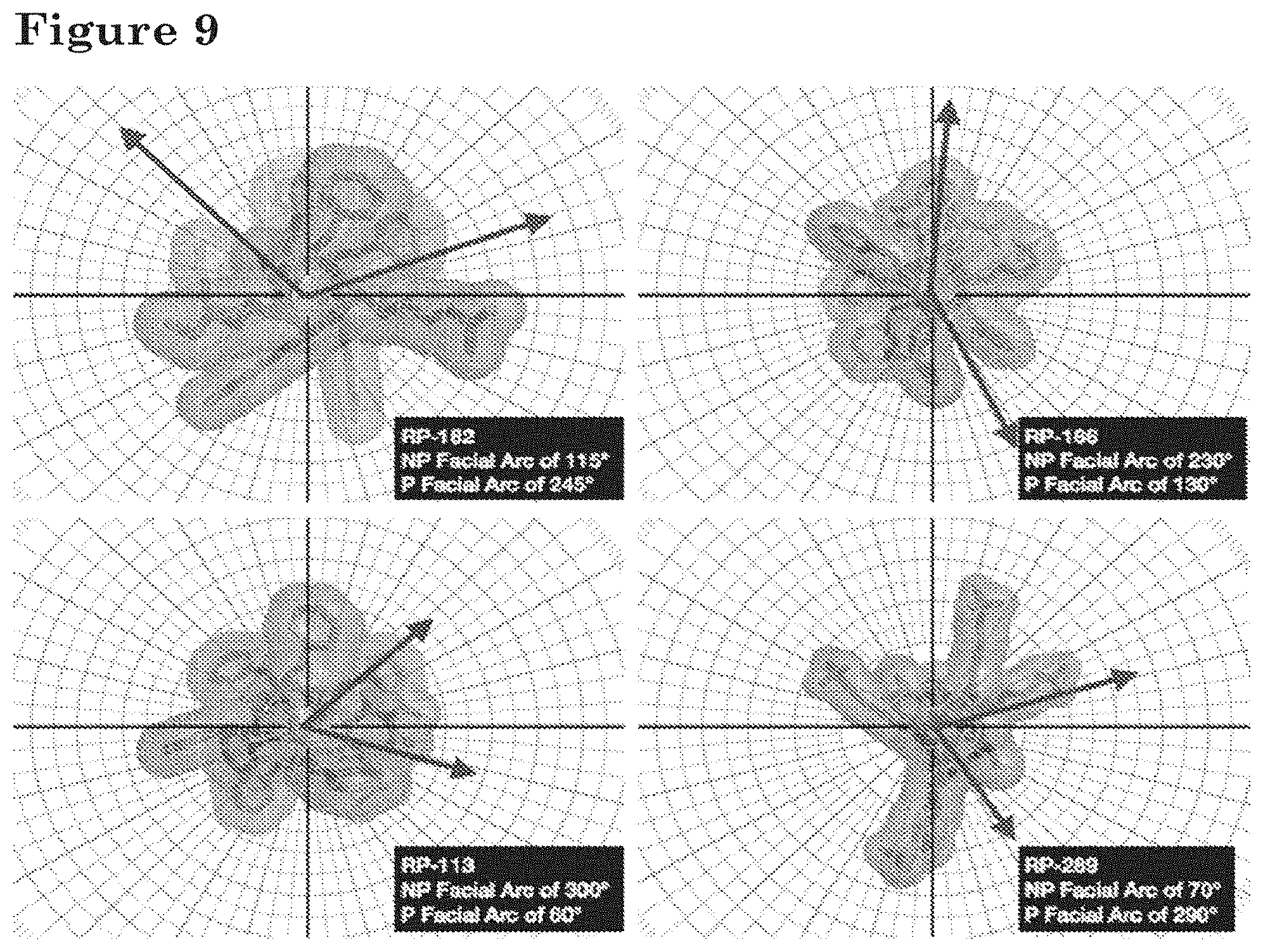
D00006

D00007
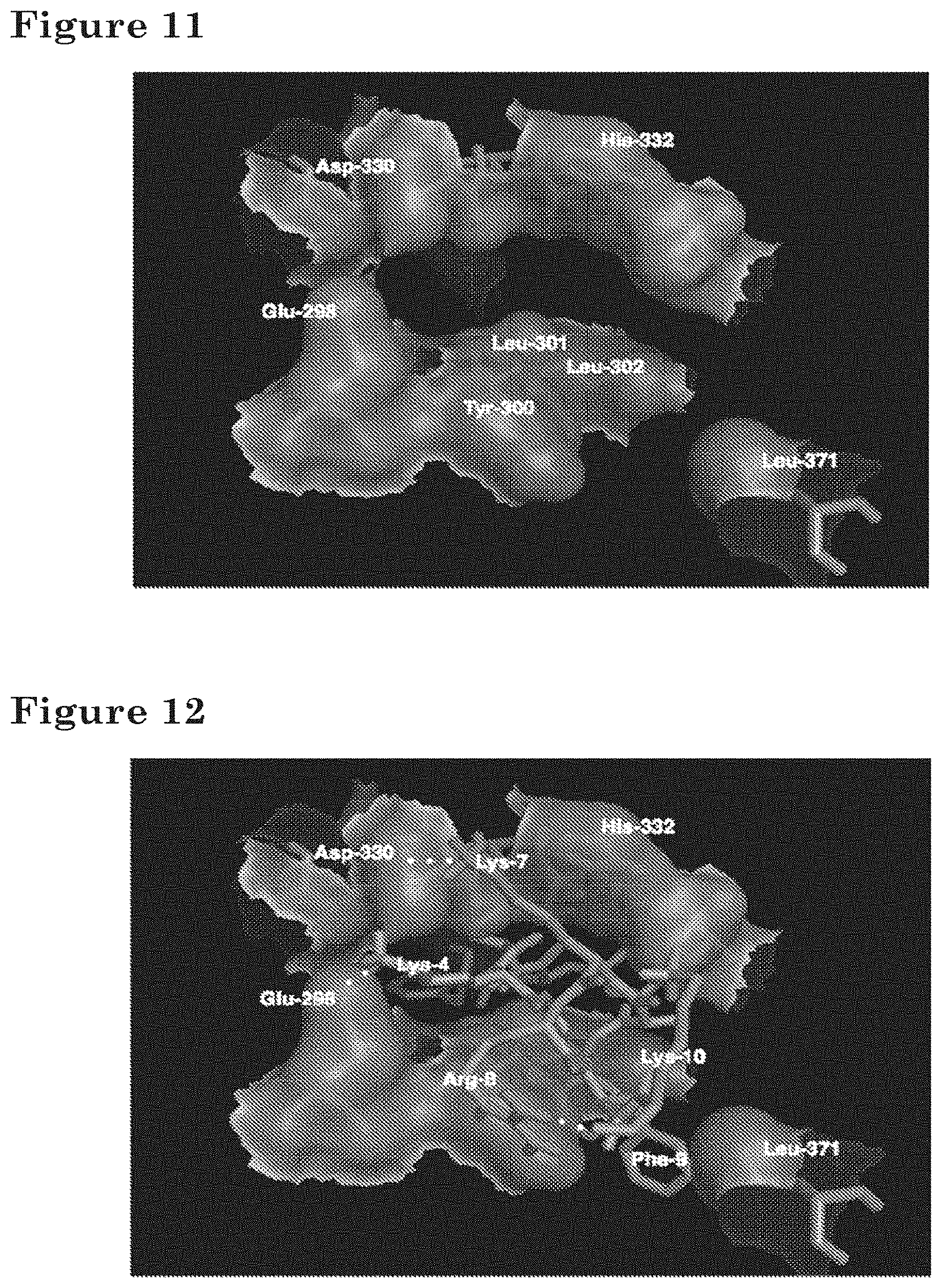
D00008

D00009

D00010

D00011

D00012

D00013

D00014

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.