Peptidase And Its Uses
FRANZETTI; Bruno ; et al.
U.S. patent application number 16/771752 was filed with the patent office on 2021-03-11 for peptidase and its uses. The applicant listed for this patent is Centre national de la recherche scientifique, Commissariata I'energie atomique et aux energies alternatives, UNIVERSITE GRENOBLE ALPES. Invention is credited to Alexandre APPOLAIRE, Hind BASBOUS, Bruno FRANZETTI, Eric GIRARD.
| Application Number | 20210071162 16/771752 |
| Document ID | / |
| Family ID | 1000005272960 |
| Filed Date | 2021-03-11 |





| United States Patent Application | 20210071162 |
| Kind Code | A1 |
| FRANZETTI; Bruno ; et al. | March 11, 2021 |
PEPTIDASE AND ITS USES
Abstract
The invention relates to the uses of a new characterized TET protein showed restricted to N-terminus glycine residues exopeptidase. The invention also relates to a method comprising said use of said new characterized TET protein as a N-terminus glycine residues specific exopeptidase. The invention further relates to a support wherein it is immobilized on said new characterized TET protein as a N-terminus glycine residues specific exopeptidase.
| Inventors: | FRANZETTI; Bruno; (SASSENAGE, FR) ; GIRARD; Eric; (ROMANS-SUR-ISERE, FR) ; APPOLAIRE; Alexandre; (GRENOBLE, FR) ; BASBOUS; Hind; (MEYLAN, FR) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005272960 | ||||||||||
| Appl. No.: | 16/771752 | ||||||||||
| Filed: | December 12, 2018 | ||||||||||
| PCT Filed: | December 12, 2018 | ||||||||||
| PCT NO: | PCT/EP2018/084632 | ||||||||||
| 371 Date: | June 11, 2020 |
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A23J 3/341 20130101; A23J 3/347 20130101; C12P 21/06 20130101; A23J 3/346 20130101; A23J 3/348 20130101; C12N 9/485 20130101 |
| International Class: | C12N 9/48 20060101 C12N009/48; C12P 21/06 20060101 C12P021/06; A23J 3/34 20060101 A23J003/34 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Dec 12, 2017 | EP | 17306758.8 |
Claims
1. A method for providing a N-terminus glycine residues specific exopeptidase, wherein said N-terminus glycine residues specific exopeptidase is provided by a TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or any homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity.
2. A method for the modification of all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins harbouring a N-terminus glycine residue, wherein said modification is performed by at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said at least TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or any homologous protein derived from said at least TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity.
3. The method according to claim 1, wherein said TET protein or said derived protein originates from an extremophilic microorganism belonging to the Thermococcales order.
4. The method according to claim 3, wherein said extremophilic microorganism is Pyrococcus horikoshii.
5. The method according to claim 2, wherein peptides, polypeptides and/or proteins of said substrate are obtained from food industry or health industry or chemical industry, preferably proteins from fermented products or soya products or sea food products or cheese products.
6. Method for degrading, from the N-terminus part, a polypeptide harbouring a glycine residue at its N-terminal part, said method comprising a step of contacting said polypeptide harbouring a glycine residue at its N-terminal part with at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or at least one homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity, and possibly a step of recovering the resulting N-terminal glycine free peptides.
7. Method for modifying all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins, wherein at least one of the peptides, polypeptides and/or proteins of said substrate harbours a N-terminus glycine residue, said method comprising a step of contacting said substrate with at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or at least one homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity, and possibly a step of recovering the modified polypeptide content of said substrate.
8. The method according to claim 6, wherein said step of contacting comprises the activation of the said TET protein or derived protein using as enzyme cofactor at least one of the metal ions of the group comprising: Ni.sup.2+, Co.sup.2+ and Mn.sup.2+, preferably using Ni.sup.2+ as enzyme cofactor.
9. The method according to claim 6, wherein said step of contacting is carried out from pH 9 to pH 10, preferably at pH 9.5.
10. The method according to claim 6, wherein said step of contacting is carried out from 80.degree. C. to 100.degree. C., preferably at 85.degree. C.
11. The method according to claim 6, wherein said at least TET protein or said derived protein is immobilized on a solid support, preferably on a filter cartridge or on silica beads or on magnetic beads or on organic polymeric materials, or on inorganic polymeric materials or on membrane devices or in microcapsules.
12. A method for the modification of all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins, wherein at least one of the peptides, polypeptides and/or proteins of said substrate harbours a N-terminus glycine residue, said method comprises the contact of said polypeptide content with a solid support, and wherein is immobilized on said solid support at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or at least one homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity.
13. The method according to claim 12, wherein said solid support is a filter cartridge or silica beads or magnetic beads or on organic polymeric materials, or on inorganic polymeric materials or on membrane devices or in microcapsules.
14. The method according to claim 12, wherein peptides, polypeptides and/or proteins of said substrate are obtained from food, chemical and health industries, or from biomass.
Description
[0001] The invention is related to peptidases and their use.
[0002] Peptidases are involved in digesting polypeptide chain of peptides and proteins into shorter fragments by splitting the peptide bonds that link amino acid residues. Some detach the terminal amino acids from the protein chain and are called exopeptidases, such as aminopeptidases, carboxypeptidase A; others attack internal peptide bonds of a protein and are called endopeptidases, such as trypsin, chymotrypsin, pepsin, papain, elastase.
[0003] Aminopeptidases are enzymes that catalyse the cleavage of amino acids from the amino terminus (N-terminus) of proteins or peptides. They are widely distributed throughout the three-domain system, i.e. archaea, bacteria, and eukaryote domains, and are found in many subcellular organelles, in cytosol, and as membrane components.
[0004] Aminopeptidases which are directed to glycine residues, called glycine aminopeptidases (GAPs), are of great interest for the industrial food. Indeed, GAPs are known to better degrade peptides enriched in glycine, which can modify the taste of food preparations from fermentation, like cheeses, the tofu or the sufu. In particular, the release of glycine from the polypeptide chains is of great interest for the Japanese industry because glycine is known to be an enhancer of sweet tastes which are specific to the Japanese gastronomy. The release of the glycine is also important for the flavour of several dry cheeses (feta, parmesan, etc. . . . ).
[0005] However, glycine residues are hard to release for aminopeptidases. Nowadays, only three aminopeptidases were found to exhibit clear preference for glycine residues. One of these GAPs is a Zn-dependent metallopeptidase from M61 family secreted by the gram-negative bacteria Sphingomonas capsulata (Jamdar, S. N. (2009)). Another one is a eukaryotic S12 family serine peptidase found in the cytosol of Actinomucor oryzae (Marui, J., et al. (2012)). The last one is the cytosolic glycyl aminopeptidase of Actinomucor elegans, for which the residues implicated in enzymatic mechanism are still ambiguous (Ito K et al. (2003)).
[0006] However, these GAPs are not restricted to the glycine residues and show a significant amidolytic activity on other amino acids. Further, these GAPs shows poor yield of production, and come from mesophilic organisms which limits their scope of application.
[0007] Therefore, there is a need to provide a new aminopeptidase specific for N-terminus glycine residues, able to operate in industrial conditions.
[0008] The aim of the invention is to obviate these drawbacks.
[0009] Thus, the invention relates to the use of a TET protein as a N-terminus glycine residues specific exopeptidase, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1,
[0010] or any homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity.
[0011] The invention also relates to the use of at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, for the modification of all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins harbouring a N-terminus glycine residue, said at least TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1,
[0012] or any homologous protein derived from said at least TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity.
[0013] The invention is based on the unexpected characterization by the inventors of a new TET protein as an exopeptidase unambiguously restricted to the cleavage of glycine residues. The inventors surprisingly found that this peptidase is devoid of amidolytic activity on all other amino acid residues except glycine residues. Advantageously, the inventors discovered that this TET protein is a thermophilic protein and can be activated by heat. This is of interest in fermentation industry for example, where the processes are carried out at high temperatures. As a TET protein, this exopeptidase harbours biophysical properties favourable to its immobilization on a support. Consequently, there is advantageously no need for recovering said TET protein in the final preparation, which allows reducing the cost of production of peptides devoid of their glycine N-terminus residues.
[0014] Hereafter, the TET protein as set forth in SEQ ID NO: 1 corresponds to the protein PhTET4.
[0015] By "exopeptidase", it is meant in the invention a peptidase that catalyses the cleavage of the terminal peptide bond of an amino acid chain, starting either from the amino or carboxyl terminal of the said amino acid chain.
[0016] By "peptide", it is meant in the invention an amino acid chain comprising at least 2 amino acids. Peptides can be obtained either from protein degradation or from chemical synthesis. By "polypeptide", it is meant in the invention an amino acid chain larger than a peptide and obtained from degradation of proteins and not from chemical synthesis.
[0017] Peptides and polypeptides may harbour biological functions within the context of a protein (signal peptide, death domain, bHLH domain . . . ). By "protein", it is meant in the invention an amino acid chain containing molecule harbouring biological function and which is found naturally in an organism, said biological function being part of a natural process of the cell.
[0018] By "at least 70% of identity with the sequence as set forth in SEQ ID NO: 1", it is meant in the invention 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% and 100% of identity with the sequence as set forth in SEQ ID NO: 1. Regarding the percentage of identity, it is defined by the percentage of amino acid residues of SEQ ID NO: 1 which align with the same amino acid in the sequence of the homologous protein. The sequence alignment is carried out using dedicated algorithms and programs (such as ClustalW, for instance).
[0019] In the invention, the term "comprising" is meant to include the terms "consisting essentially of" and "consisting of".
[0020] By "modification of all or part of the polypeptide", it is meant in the invention that the modification of a peptide can result in the removal of one or more amino acid from the peptide. If the peptide contains only aromatic amino acids, the peptide can be completely degraded, i.e. can be converted into the free amino acids that constitute the peptide.
[0021] Moreover, the "modification of all or part of the polypeptide", means also in the invention that, if a composition contains two or more peptides, at least one peptide will be degraded by contacted the TET protein according to the invention. If only some peptides are degraded, the composition of peptide will be considered to be partially modified. If all the peptides are subjected to a degradation, the composition of peptide will be considered to be totally modified.
[0022] In the invention, regarding a peptide, a polypeptide, a protein or a polypeptide content, the terms "modification" and "degradation" can be used uniformly.
[0023] Advantageously, said TET protein or said derived protein originates from an extremophilic microorganism belonging to the Thermococcales order and is isolated from this extremophilic microorganism. By "extremophilic", it is meant in the invention an organism that thrives in physically or geochemically extreme conditions that are detrimental to most life on Earth. In contrast, organisms that live in more moderate environments may be termed mesophiles or neutrophiles. Among the hyperthermophilic archaea, representatives of order Thermococcales form the most numerous group to date. Members of this group are the most frequently isolated hyperthermophilic archaea. They are heterotrophic and as such regarded as the major constituents of organic matter within marine hot water ecosystems. They belong to the branch of Euryarchaeota that contains the methanogens, the genus Thermoplasma, and the extremely halophilic archaea. The Thermococcales order is actually represented by three genera: Pyrococcus, Thermococcus and the newly described Paleococcus.
[0024] Advantageously, said extremophilic microorganism is Pyrococcus horikoshii.
[0025] Advantageously, peptides, polypeptides and/or proteins of said substrate are obtained from food industry or health industry or chemical industry. Advantageously, proteins from fermented products or soya products or sea food products or cheese products.
[0026] The generation of protein-rich industrial wastes is very high (only from sunflower, about one million tons in Spain). These wastes are not used at all, or are underused in the form of low added-value products. This type of by-products constitutes a reservoir of proteins with a great economic potential.
[0027] The TET protein according to the invention, in view of its activity, can be used in various domains for instance, but without limitation: [0028] For the valorization of agriculture wastes: proteins and peptides originating from agriculture can be recycled for animal feed, or for producing feed additives. In order to be used, the vegetal proteins have to be degraded to avoid any antinutritional side effects. [0029] For the valorization of chemical wastes [0030] For the valorization of food industry waste: suitable proteins to be treated with the TET aminopeptidase according to the invention may be for instance proteins obtained from dairy products, fruit juices, beers, flours or cured products. Moreover, the products from wine industry, from cheese industry, and see food industry are particularly advantageous and can be valorized by using the TET protein according to the invention. [0031] Products from biomass: some alternative to common protein sources are now emerging in view of the need to provide more and more feed for the Earth population. For instance, algae and microorganisms, along with insects, are very rich in protein that can be used for providing new sources of amino acids or proteins that can be eaten by animal and humans. Therefore, proteins or peptides from algae or microorganisms can be relevant sources for treatment by using the TET protein according to the invention. [0032] Another kind of wastes are the wastes produced from health industry. For instance, solid, regulated medical waste can includes materials generated in the diagnosis, treatment, research, or immunization of human beings or animals. Examples of regulated medical waste includes: cultures and stocks, pathological wastes, human blood and blood products, sharps, certain animal waste and isolation wastes. The TET protein according to the invention may help to valorize such kinds of wastes.
[0033] The invention also relates to a method for degrading, from the N-terminus part, a polypeptide harbouring a glycine residue at its N-terminal part, said method comprising a step of contacting said polypeptide harbouring a glycine residue at its N-terminal part with
[0034] at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or
[0035] at least one homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity,
[0036] and possibly a step of recovering the resulting N-terminal glycine free peptides.
[0037] The invention also relates to a method for modifying all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins, wherein at least one of the peptides, polypeptides and/or proteins of said substrate harbours a N-terminus glycine residue, said method comprising a step of contacting said substrate with
[0038] at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or
[0039] at least one homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity,
[0040] and possibly a step of recovering the modified polypeptide content of said substrate.
[0041] Advantageously, in the method for modifying all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins harbouring a N-terminus glycine residue, said step of contacting comprises the activation of the said TET protein or derived protein using as enzyme cofactor at least one of the metal ions of the group consisting of Ni.sup.2+, Co.sup.2+ and Mn.sup.2+, preferably using Ni.sup.2+ as enzyme cofactor. This aspect of the invention is interesting because Ni.sup.2+ is a rare cofactor of aminopeptidase. Indeed, most of the aminopeptidases are activated by Zn.sup.2+ cofactor. Consequently, PhTET4 can be selectively activated in a pool of aminopeptidases.
[0042] In particular, said step of contacting is carried out from pH 9 to pH 10, provided that the aminopeptidase activity of said at least TET protein or said derived protein being maintained to a aminopeptidase activity of at least 80% of their maximum activity. Advantageously, said step of contacting is carried out at least at pH 9. By "at least at pH 9", it is meant in the invention pH 9, pH 9.1, pH 9.2, pH 9.3, pH 9.4, pH 9.5. Advantageously, said step of contacting is carried out at most at pH 10. By "at most at pH 10", it is meant in the invention pH 10, pH 9.9, pH 9.8, pH 9.7, pH 9.6. Advantageously said step of contacting is carried out at pH 9.5.
[0043] Advantageously, said step of contacting is carried out from 80.degree. C. to 100.degree. C., provided that the aminopeptidase activity of said at least TET protein or said derived protein being maintained to an aminopeptidase activity of at least 80% of their maximum activity. Advantageously, said step of contacting is carried out at least at 80.degree. C. By "at least at 80.degree. C.", it is meant in the invention, 80.degree. C., 80.1.degree. C., 80.2.degree. C., 80.3.degree. C., 80.4.degree. C., 80.5.degree. C., 80.6.degree. C., 80.7.degree. C., 80.8.degree. C., 80.9.degree. C., 81.degree. C., 81.1.degree. C., 81.2.degree. C., 81.3.degree. C., 81.4.degree. C., 81.5.degree. C., 81.6.degree. C., 81.7.degree. C., 81.8.degree. C., 81.9.degree. C., 82.degree. C., 82.1.degree. C., 82.2.degree. C., 82.3.degree. C., 82.4.degree. C., 82.5.degree. C., 82.6.degree. C., 82.7.degree. C., 82.8.degree. C., 82.9.degree. C., 83.degree. C., 83.1.degree. C., 83.2.degree. C., 83.3.degree. C., 83.4.degree. C., 83.5.degree. C., 83.6.degree. C., 83.7.degree. C., 83.8.degree. C., 83.9.degree. C., 84.degree. C., 84.1.degree. C., 84.2.degree. C., 84.3.degree. C., 84.4.degree. C., 84.5.degree. C., 84.6.degree. C., 84.7.degree. C., 84.8.degree. C., 84.9.degree. C., 85.degree. C. Advantageously, said step of contacting is carried out at most at 100.degree. C. By "at most at 100.degree. C.", it is meant in the invention 100.degree. C., 99.9.degree. C., 99.8.degree. C., 99.7.degree. C., 99.6.degree. C., 99.5.degree. C., 99.4.degree. C., 99.3.degree. C., 99.2.degree. C., 99.1.degree. C., 99.degree. C., 98.9.degree. C., 98.8.degree. C., 98.7.degree. C., 98.6.degree. C., 98.5.degree. C., 98.4.degree. C., 98.3.degree. C., 98.2.degree. C., 98.1.degree. C., 98.degree. C., 97.9.degree. C., 97.8.degree. C., 97.7.degree. C., 97.6.degree. C., 97.5.degree. C., 97.4.degree. C., 97.3.degree. C., 97.2.degree. C., 97.1.degree. C., 97.degree. C., 96.9.degree. C., 96.8.degree. C., 96.7.degree. C., 96.6.degree. C., 96.5.degree. C., 96.4.degree. C., 96.3.degree. C., 96.2.degree. C., 96.1.degree. C., 96.degree. C., 95.9.degree. C., 95.8.degree. C., 95.7.degree. C., 95.6.degree. C., 95.5.degree. C., 95.4.degree. C., 95.3.degree. C., 95.2.degree. C., 95.1.degree. C., 95.degree. C., 94.9.degree. C., 94.8.degree. C., 94.7.degree. C., 94.6.degree. C., 94.5.degree. C., 94.4.degree. C., 94.3.degree. C., 94.2.degree. C., 94.1.degree. C., 94.degree. C., 93.9.degree. C., 93.8.degree. C., 93.7.degree. C., 93.6.degree. C., 93.5.degree. C., 93.4.degree. C., 93.3.degree. C., 93.2.degree. C., 93.1.degree. C., 93.degree. C., 92.9.degree. C., 92.8.degree. C., 92.7.degree. C., 92.6.degree. C., 92.5.degree. C., 92.4.degree. C., 92.3.degree. C., 92.2.degree. C., 92.1.degree. C., 92.degree. C., 91.9.degree. C., 91.8.degree. C., 91.7.degree. C., 91.6.degree. C., 91.5.degree. C., 91.4.degree. C., 91.3.degree. C., 91.2.degree. C., 91.1.degree. C., 91.degree. C., 90.9.degree. C., 90.8.degree. C., 90.7.degree. C., 90.6.degree. C., 90.5.degree. C., 90.4.degree. C., 90.3.degree. C., 90.2.degree. C., 90.1.degree. C., 90.degree. C., 89.9.degree. C., 89.8.degree. C., 89.7.degree. C., 89.6.degree. C., 89.5.degree. C., 89.4.degree. C., 89.3.degree. C., 89.2.degree. C., 89.1.degree. C., 89.degree. C., 88.9.degree. C., 88.8.degree. C., 88.7.degree. C., 88.6.degree. C., 88.5.degree. C., 88.4.degree. C., 88.3.degree. C., 88.2.degree. C., 88.1.degree. C., 88.degree. C., 87.9.degree. C., 87.8.degree. C., 87.7.degree. C., 87.6.degree. C., 87.5.degree. C., 87.4.degree. C., 87.3.degree. C., 87.2.degree. C., 87.1.degree. C., 87.degree. C., 86.9.degree. C., 86.8.degree. C., 86.7.degree. C., 86.6.degree. C., 86.5.degree. C., 86.4.degree. C., 86.3.degree. C., 86.2.degree. C., 86.1.degree. C., 86.degree. C., 85.9.degree. C., 85.8.degree. C., 85.7.degree. C., 85.6.degree. C., 85.5.degree. C., 85.4.degree. C., 85.3.degree. C., 85.2.degree. C., 85.1.degree. C. Advantageously said step of contacting is carried out at 85.degree. C. Carried out amidolytic activity at high temperature is interesting in industry, because one use of the TET protein is about fermentation which is also carried out at high temperature. Moreover, the TET protein is heat activatable and can be therefore specifically activated during the fermentation step of an industrial process.
[0044] Compared to other TET family member peptidases, the TET protein according to the invention is active at a pressure varying from 0.1 MPa to 350 MPa.
[0045] Advantageously, said at least TET protein or said derived protein is immobilized on a solid support. As abovementioned, the biophysical properties of a TET protein allow its use on a support. Consequently, in an industrial process for a final preparation, the TET protein can be put into contact with the substrate without mixing them together so that there is no need to recover said TET protein in the final preparation. Advantageously, said at least TET protein or said derived protein is immobilized on a filter cartridge or on silica beads or on magnetic beads or on organic polymeric materials, or on inorganic polymeric materials or on membrane devices or in microcapsules. Membrane devices include hollow fibers.
[0046] In view of the robustness of the TET protein, and the stability of the structure, it is possible to immobilized the TET protein on a support. Such a support is advantageous and allows to carry out a peptide, polypeptide or peptide degradation and recover the resulting degraded peptide, polypeptides and protein, easily without additional step of separation of the enzyme and the resulting product. Current enzyme immobilization methods are described in the review of Bilal et al. (Bilal M, Iqbal H M, Guo S, Hu H, Wang W, Zhang X (2017) State-of-the-art protein engineering approaches using biological macromolecules: A review from immobilization to implementation view point. Int J Biol Macromol. November 2.).
[0047] The invention is further related to the use of a solid support for the modification of all or part of the polypeptide content of a substrate comprising peptides, polypeptides and/or proteins, wherein at least one of the peptides, polypeptides and/or proteins of said substrate harbours a N-terminus glycine residue, and wherein is immobilized on said solid support
[0048] at least a TET protein harbouring at least a N-terminus glycine residues specific exopeptidase activity, said TET protein comprising the amino acid sequence as set forth in SEQ ID NO: 1, or
[0049] at least one homologous protein derived from said TET protein as set forth in SEQ ID NO: 1 by substitution, addition or deletion of at least one amino acid, provided that the derived protein retains at least 70%, preferably at least 79% of identity with the amino acid sequence as set forth in SEQ ID NO: 1, and said derived protein retaining a N-terminus glycine residues specific exopeptidase activity.
[0050] Advantageously, said solid support is a filter cartridge or silica beads or magnetic beads or on organic polymeric materials, or on inorganic polymeric materials or on membrane devices or in microcapsules. Membrane devices include hollow fibers.
[0051] Advantageously, peptides, polypeptides and/or proteins of said substrate are obtained from food, chemical and health industries, or from biomass, as mentioned above. Advantageously, proteins from fermented products or soya products or sea food products or cheese products.
[0052] Alternatively, said at least TET protein or said derived protein is immobilized as cross-linked enzyme aggregates (CLEAs).
[0053] CLEAs are developed by precipitation of the enzyme from a solution by adding salt, such as ammonium sulphate, or water-miscible organic solvent, followed by cross-linking with a bifunctional reagent (Bilal M, Iqbal H M, Guo S, Hu H, Wang W, Zhang X (2017) State-of-the-art protein engineering approaches using biological macromolecules: A review from immobilization to implementation view point. Int J Biol Macromol. November 2.).
[0054] The invention will be better understood from the following example and the 8 following figures.
LEGEND TO THE FIGURES
[0055] FIG. 1 is a sequence alignment of PhTET1, 2, 3 and 4. Conserved residues are in a grey box and similar residues are boxed. The numbering and the secondary structure elements are those of PhTET1 (PDB ID: 2WYR). Light grey stars and heavy grey starts highlight the metal-binding and active residues, respectively.
[0056] FIGS. 2A and 2B are respectively an elution profile of PhTET4 on size-exclusion chromatography column (superdex200 10/300 GL), and a micrograph of the said eluted PhTET4 observed by negative strain electron microscopy. In FIG. 2A, X-axis represents the volume (ml) of exclusion and Y-axis represents the absorbance (mAU) at 280 nm.
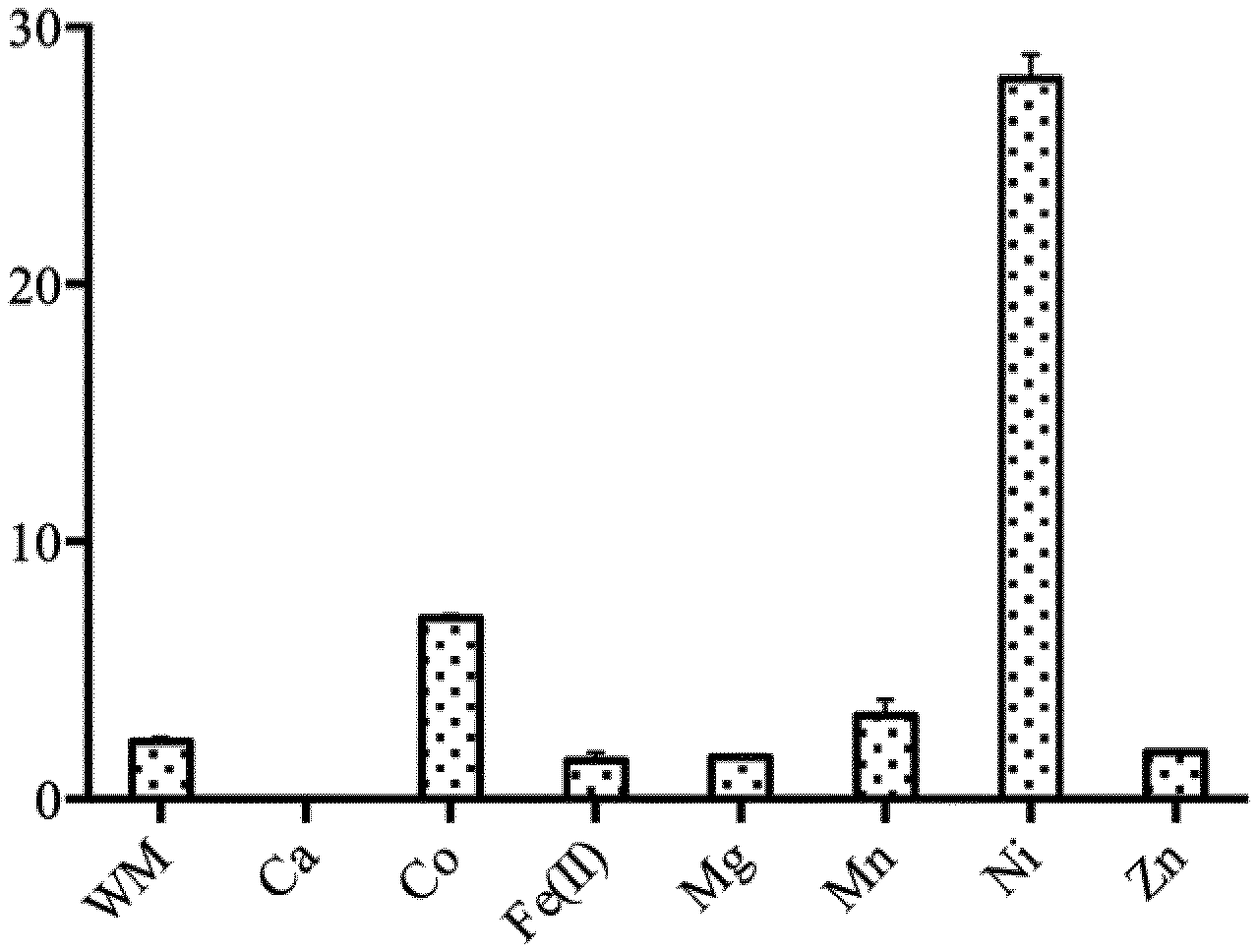
[0057] FIG. 3 is a graph representing the variation of PhTET4 specific activity in the absence (WM) of metal, or in the presence of ions Calcium (Ca), Cobalt (Co), Fer (Fell), Magnesium (Mg), Nickel (Ni), and Zinc (Zn). Y-axis represent the specific activity of PhTET4 (.mu.mol (pNA) mg-1 (PhTET4) min-1). pNA stands for p-Nitroaniline.
[0058] FIG. 4 is a graph representing the evolution of PhTET4 specific activity from pH 6 to pH 11. Y-axis represent the specific activity of PhTET4 (.mu.mol (pNA) mg-1 (PhTET4) min-1). pNA stands for p-Nitroaniline.
[0059] FIG. 5 is a graph representing the evolution of PhTET4 specific activity from 20.degree. C. to 95.degree. C. Y-axis represent the specific activity of PhTET4 (.mu.mol (pNA) mg-1 (PhTET4) min-1). pNA stands for p-Nitroaniline.
[0060] FIG. 6 represents the size exclusion chromatography of PhTET4 in absence or presence of EDTA. The continuous line represents the size exclusion chromatography of PhTET4 in absence of EDTA. The discontinuous line represents the size exclusion chromatography of PhTET4 in presence of EDTA. X-axis represent the volume (ml) of exclusion. Y-axis represents the absorbance (mAU) at 280 nm.
[0061] FIG. 7 is a graph representing the relative amidolytic activity of PhTET4 on L-Alanine (A), D-Alanine (D-A), L-Aspartate (D), L-Glutamate (E), L-Phenylalanine (F), L-Glycine (G), L-Histidine (H), L-Isoleucine (I), L-Lysine (K), L-Leucine (L), L-Methionine (M), L-Asparagine (N), L-Proline (P), L-Glutamine (Q), L-Arginine (R), L-Serine (S), L-Threonine (T), L-Valine (V), L-Tryptophan (W), and L-Tyrosine (Y). Y-axis represents the relative activity (%) of PhTET4.
[0062] FIG. 8A represents the size exclusion chromatography of GLM. FIG. 8B represents the size exclusion chromatography of GMDSLAFSGGL (SEQ ID NO: 6) peptides, respectively, after incubation with PhTET4. In both figures, the continuous line represents the size exclusion chromatography of the intact peptide, and the discontinuous line represents size exclusion chromatography of the said peptide cleaved by PhTET4. X-axis represents the elution time (sec). Y-axis represents the Absorbance (mAU) at 214 nm.
EXAMPLE
[0063] Methods
[0064] PhTET4 Expression and Purification
[0065] The gene encoding for PH0737 (SEQ ID NO: 5) was cloned in pET-41c vector GeneCust Europe, Luxembourg. Recombinant protein was overexpressed in Escherichia coli BL21 (DE3)-RIL strain during 4 hours at 37.degree. C. by induction with 0.1 mM IPTG in 1 L of lysogeny broth medium. The cell pellet was conserved at -80.degree. C. until using. The cells were re-suspended in 25 mL Tris-HCl 50 mM pH 8, NaCl 90 mM and Triton X-100 0.1%, supplemented with 6.25 mg lysozyme (Euromedex), 1.25 mg DNase I grade II (Roche), 5 mg RNase (Roche), 25 mg Pefabloc SC (Roche) and 0.25 mL MgSO4 at 2 M. Cells were disrupted by sonication at 30 watts with 5 cycles on/off of 30 sec each at 4.degree. C., and then heated for 20 min at 75.degree. C. to eliminate most mesophilic proteins of host strain. The lysate was clarified by centrifugation at 12,000 rpm, 4.degree. C. for 1 hour with JA20 rotor (Beckman), and the supernatant was loaded on ResourceQ column (GE healthcare) equilibrated with 20 mM Tris-HCl pH 7.5, 100 mM NaCl. The 30 mL of the flow-through were retained and diluted with 20 mL Tris-HCl pH 7.5 to a final concentration of 105 mM NaCl. The new supernatant volume was loaded a second time on ResourceQ column. After washing the column with 20 mM Tris-HCl pH 7.5, 100 mM NaCl, bound proteins were eluted with linear salt gradient (154-290 mM NaCl). The protein-contained fractions were pooled together and loaded on Superdex200 10/300 GL size exclusion column (GE healthcare) equilibrated with 20 mM Tris-HCl pH 7.5, 150 mM NaCl. PhTET4 protein was eluted at 11 mL of column exclusion volume. The peak fractions were combined and concentrated with Amicon Ultra 30 kDa cut-off and stored at 4.degree. C. The purity of prepared protein was checked by SDS-PAGE. 3 mg of PhTET4 were produced from 1 L of culture.
[0066] PhTET4 Negative Stain Electron Microscopy
[0067] After the size exclusion chromatography step, 4 .mu.l of PhTET4 at 0.05 mg/ml were deposited onto carbon-coated 400-mesh copper grids. The samples were stained using uranyl acetate 2% and air-dried. Images were taken under low-dose conditions in a T12FEI electron microscope working at 120 kV and with a nominal magnification of 40,000 using an Orius SC1000 CCD camera.
[0068] PhTET4 Substrate Specificity
[0069] The hydrolytic activity of PhTET4 was determined by using chromogenic and fluorogenic compounds: aminoacyl-pNAs (p-Nitroaniline) and aminoacyl-AMCs (7-Amino-4-methylcoumarin), respectively. Reactions were initiated by addition of 4 .mu.g/ml (final concentration) of PhTET4 to 400 .mu.l of pre-warmed mixture containing 5 mM chromogenic or fluorogenic substrate in 50 mM PIPES, 150 mM KCl, pH 7.5 at 80.degree. C. Since all tetrahedral aminopeptidases of P. horikoshii are activated by cobalt, the inventors started by using 0.1 mM CoCl2 as metal activator for PhTET4 enzymatic activity. Then, the experimentation was later repeated using the determined optimal conditions (0.1 mM NiCl2, pH 9.5 and at 85.degree. C.). In order to avoid water evaporation, the total volume was covered by a layer of mineral oil. Catalytic activity was followed during 10 min by measuring the absorbance of released pNA at 405 nm or the AMC fluorescence using excitation and emission wavelengths of 355 and 460 nm, respectively.
[0070] PhTET4 Metallic Cofactor
[0071] In presence of cobalt low activity was identified against Gly-pNA. So, in order to enhance the enzymatic activity of PhTET4, several divalent metals were tested at 0.1 mM final concentration in a reaction volume containing 4 .mu.g/ml PhTET4, 50 mM CHES, 150 mM KCl and 5 mM Gly-pNA, pH 9.5. The reaction was followed at 80.degree. C. by measuring the absorbance of released pNA at 405 nm during 10 min.
[0072] PhTET4 Optimal pH
[0073] The effect of pH on PhTET4 enzymatic activity was studied by using different buffers: PIPES, pH 6-7.5; CHES, pH 8.2-10 and CAPS, pH 10.5-11. All buffers were used at 50 mM final concentration in presence of 4 .mu.g/ml PhTET4, 150 mM KCl, 0.1 mM NiCl2 and 5 mM Gly-pNA at 80.degree. C. The incubation was done for 10 min by measuring the absorbance of released pNA at 405 nm.
[0074] PhTET4 Optimal Temperature
[0075] The temperature impact on the enzymatic activity of PhTET4 was measured in a range from 20 to 95.degree. C. In all cases, 4 .mu.g/ml of PhTET4 were incubated with 50 mM CHES, 150 mM KCl, 0.1 mM NiCl.sub.2 and 5 mM Gly-pNA, pH 9.5 during 10 min. The reaction was assessed as described previously.
[0076] Peptide Substrates
[0077] More enzymatic studies were performed to decipher PhTET4 cleavage activity by using peptide substrates (GI; GL; GLM; GMDSLAFSGGL (SEQ ID NO: 6); LGG). 6 .mu.g/ml (final concentration) of PhTET4 were added to a pre-warmed mixture of 3 mM peptide, 50 mM CHES, 150 mM KCl, 0.1 mM NiCl.sub.2, pH 9.5. To avoid water evaporation, 20 .mu.l of mineral oil were added on the top of the total volume. The reaction incubation was done at 85.degree. C. during 6 min. Aliquot of 80 .mu.l was then removed and added to 220 .mu.l of 2% acetonitrile (ACN), 0.1% trifluoroacetic acid (TFA). Proteins were removed by centrifugation at 13,000 rpm during 15 min. 100 .mu.l of the supernatant were retrieved and injected on Nova-Pak C18 column, (4 .mu.m, 3.9.times.300 mm, (Waters)) in a HPLC purifier system (PerkinElmer), equilibrated with 2% ACN, 0.1% TFA. The elution of peptide products was achieved with a linear ACN gradient (2-33.2%) and followed by measuring the absorbance at 214 nm. Chromatographic runs were carried out at room temperature. The separated fragments were collected and submitted to N-terminal sequence analysis.
[0078] PhTET4 Inhibition
[0079] PhTET4 was incubated in presence of 5 mM EDTA, 50 mM CHES, 150 mM KCl, pH 9.5 at 85.degree. C. during 15 min. Then, the oligomeric state of inhibited protein was evaluated by using exclusion chromatography column (Superdex200) equilibrated with the same incubation buffer: 50 mM CHES, 150 mM KCl, pH 9.5.
[0080] Results
[0081] In all cell types, metallo-aminopeptidases play crucial roles in energy metabolism, protein maturation and degradation and in the regulation of biologically active peptides by removing the N-terminal amino acid from proteins and oligopeptides. Overall, metalloenzymes all have one feature in common, namely that the metal ion is bound to the protein with one labile coordination site. As with all enzymes, the shape of the active site is crucial. The metal ion is usually located in this active site or in the catalytic pocket. In most of case, metallo-aminopeptidases operate beyond the action of an endoprotease and their activity is limited to small peptides.
[0082] Several aminopeptidases assemble as large dodecameric particles, first discovered in archaea and named TET due to their peculiar tetrahedral shapes. The 13 nm hollow dodecahedrons enclose 12 active sites distributed in 4 funnel-shaped chambers located in the apices of the particles and four large access holes, formed by the junction of six subunits, situated in the facets. This organization strongly distinguishes TET from the other cytosolic compartmentalized peptidases that mostly adopt a barrel-shaped architecture.
[0083] Biochemical and structural studies of TET peptidases have accumulated over the past 10 years. They revealed that TET dodecamers represent a common scaffold for an efficient polypeptide capture and processing system. In all TET, the aminopeptidase activity is based on co-catalytic dinuclear metal active site belonging to M18 or M42 peptidase families according to MEROPS peptidase database. Bounding peptides are cleaved following common mechanism involving water molecule and glutamate residue. The nature of the metal occupying the bimetallic active site has been shown to modulate TET enzymatic activity and Co.sup.2+ ions appear to be the best activators for almost all archaeal and bacterial M42 TET.
[0084] TET machines are widespread and were found in the three life domains. Interestingly, in prokaryotes, 1 to 4 different types of TET complexes can co-exist in the cytosol depending upon the cell type. These enzymes can be categorized according to their preference for the chemical structure of the N-terminal amino acid residues present in the polypeptide chains. Three main categories have been identified so far: glutamyl/aspartyl aminopeptidases, lysine aminopeptidases and leucine aminopeptidases, these later exhibiting the broader specificities. In eukarya, M18 TET complexes displayed aspartyl aminopeptidase activity. In bacteria, M42 TET peptidases, from Clostridium thermocellum and Thermotoga maritima, were assigned as leucyl aminopeptidases. Two M42 enzymes from the pathogens Streptococcus pneumoniae and Mycoplasma hyopneumoniae displayed glutamyl-aminopeptidase activities. In archaea, the common TET structural scaffold can harbour disparate functions. So far, the unique TET complex described in the halophilic archaeon Haloarcula marismortui displayed the broader specificities with a preference for neutral and basic residues.
[0085] 1. Characterisation of a New TET Protein, PhTET4
[0086] Pyrococcus horikoshii is a deep-sea hyperthermophilic archaeon belonging to the Thermococcales order that was isolated at 1395 m depth from a hydrothermal vent. P. horikoshii, and other related hyperthermophilic Thermococcales are distinguished by the fact that they possess three different TET complexes: PhTET1 (SEQ ID NO: 2), a glutamyl/aspartyl aminopeptidase, PhTET2 (SEQ ID NO: 3), a leucyl aminopeptidase with a broad activity against neutral amino-acids and PhTET3 (SEQ ID NO: 4), a lysyl aminopeptidase with a clear preference for positively charged residues. The analysis of their activities on synthetic peptides of different sizes and compositions using reverse phase HPLC indicated that the TET peptidases degrade oligopeptides in a sequential manner and displayed strict aminopeptidase behaviour.
[0087] An analysis of the genomes of P. horikoshii revealed the existence of a conserved gene PH0737 encoding for an unassigned peptidase of MH clan in MEROPS peptidase database. The protein encoded by PH0737 gene shares 20.6%, 22.5% and 22% of sequence identity with the three characterized aminopeptidases PhTET1, PhTET2 and PhTET3 respectively (FIG. 1). The residues involved in the coordination of metal ions in M42 peptidase family are well conserved between PhTET1, 2, 3 and the said unassigned peptidase. Moreover, the unassigned peptidase comprises two regions (the catalytic domain and the little .beta. sheet domain localized on the top of the former) that conferring the ability of M42 peptidases to form large multimers. Most likely, a shift was occurred for the two latest putative ligands of the unassigned peptidase, Asp231 and His311, which are highlighted by light grey triangle in comparison with the conserved position of PhTET1, 2 and 3 ligands. Hence, said unassigned peptidase was named PhTET4. PhTET4 refers to SEQ ID NO: 1.
[0088] 2. Determination of the Three-Dimensional Structure of PhTET4
[0089] The three-dimensional structure of PhTET4 was assessed by producing the recombinant PhTET4 in E. coll. The cellular extract was clarified by heat shock precipitation and the recombinant protein was purified by ion-exchange and size-exclusion chromatographies. At the final step, PhTET4 was eluted at the same exclusion volume (11 mL) than PhTET1 to 3, suggesting that it forms large molecular weight assembly similar to the .about.500 kDa TET dodecameric complexes (FIG. 2A). Negative stain electron microscopy micrographs realized on PhTET4 peak fractions, showed that the protein self-assembled in a hollow tetrahedral-shaped complex of homogenous size (FIG. 2B). The dimensions and the shape of PhTET4 are analogous to the three P. horikoshii tetrahedral TET edifices: PhTET1, PhTET2 and PhTET3.
[0090] 3. Characterisation of PhTET4 Amidolytic Activity
[0091] To determine PhTET4 functional identity the inventors tested its amidolytic activity toward the 20 amino acids by using a broad array of chromogenic p-nitroaniline (pNA) or fluorogenic 7-amino-4-methylcoumarin (AMC) conjugated aminoacyl compounds. Because of the homologies between phTET4 with PhTET1, 2 and 3, the inventors first test the amidolytic activity of PhTET4 using the known operative conditions of PhTET1, 2 and 3. Co.sup.2+ being the main activating metal of the 3 TET enzymes from P. horikoshii, the inventors first assayed PhTET4 activity in the presence of 0.1 mM CoCl.sub.2 as enzyme cofactor, at 80.degree. C. and in identical buffer conditions (5 mM substrate, 50 mM PIPES, 150 mM KCl, pH 7.5). Surprisingly, the results showed that no hydrolysis was observed against all tested substrates with the notable exception of Gly-pNA toward which PhTET4 exhibited a weak catalytic activity.
[0092] 4. Characterisation of PhTET4 Operatives Conditions
[0093] a) Metal Cofactors
[0094] Since metal cofactors have been shown to be essential to control the activity and the oligomeric state of the various TET edifices characterized so far, the inventors tested the influence of various metal ions on PhTET4 glycyl aminopeptidase activity. The results are shown on FIG. 3. Surprisingly, the inventors revealed that Ni.sup.2+ shows the most important stimulating effect on PhTET4 cleavage activity, with 12 times greater activity compared with control assay where no metallic ion (WM) was added to the reaction volume. Co.sup.2+ and Mn.sup.2+ also stimulated PhTET4 activity but less efficiently than Ni.sup.2+ (3 and 1.4 fold activation, respectively). Interestingly, Zn.sup.2+, Ca.sup.2+, Fe.sup.2+ and Mg.sup.2+ were found to inhibit PhTET4 hydrolytic activity, and total inhibition was observed in presence of Ca.sup.2+ ions. This is the first time that Ni.sup.2+ ions have been described as an essential activating cofactor of an aminopeptidase from the M42 family. This is an advantage and allow user to selectively activate PhTET4 in a pull of peptidases.
[0095] b) pH Conditions
[0096] In order to determine the influence of pH on PhTET4 enzymatic behaviour, the amidolytic activity was measured between pH 6 and pH 11, against Gly-pNA at 80.degree. C. The results are shown on FIG. 4. The optimal activity was found at pH 9.5 and a significant percentage of activity (beyond 80% of the maximal activity) was observed from pH 9 to pH 10. Consequently, a significant percentage of activity is maintained at elevated pH. These experiments revealed that, compared to the other 3 PhTETs enzymes, PhTET4 can be defined as an alkaline peptidase.
[0097] c) Temperature Conditions
[0098] To assess the hyperthermophilic properties of the enzyme, the temperature dependence of PhTET4 activity was studied at different temperatures varying from 20 to 95.degree. C. The results are shown on FIG. 5. PhTET4 enzymatic activity increases in parallel with the augmentation of heating temperature, with a maximal activity observed at 95.degree. C. in the measurable temperature range. A significant percentage of activity (beyond 80% of the maximal activity) was observed from 80.degree. C. to 95.degree. C. Thus PhTET4 displays a high hyperthermophilic behaviour comparable to the ones reported for the 3 other PhTETs aminopeptidases present in Pyrococcus horikoshii cells. Therefore, PhTET4 can advantageously work at high temperature required by industrial fermentation process.
[0099] d) Inhibition of PhTET4 Amidolytic Activity Using EDTA
[0100] In case of PhTET1, 2 and 3 aminopeptidases, it has been shown that the two metal ions present in the catalytic site are essential both for catalysis and for assembling the dodecameric edifice. For this reason, treatment with the divalent ions chelating agent EDTA leads to the dissociation of the TET quaternary structures and to enzymes inactivation. Indeed, in the optimal conditions for PhTET4 activity, the addition of 5 mM EDTA completely inhibited the glycine aminopeptidase activity. However, analysis of the oligomeric state of EDTA-treated PhTET4 samples by size-exclusion chromatography revealed that the PhTET4 dodecameric edifice remained unaffected by the EDTA treatment unlike what was reported for all the other TET enzymes (FIG. 6). This suggests that the contribution of the metals ions situated in position M1 and/or M2 for PhTET4 oligomerization is not as important as for the other TET peptidases.
[0101] e) Amidolytic Activity of PhTET4 Using Optimal Operation Conditions
[0102] The initial characterization of PhTET4 aminopeptidase activity indicated that the enzyme displayed narrow substrate specificity with a strong preference for glycine residues. In order to consolidate this finding, the experiments were repeated in the presence of nickel and in the optimal temperature and pH conditions defined above (0.1 mM NiCl.sub.2, pH 9.5 and at 85.degree. C.), and results are represented in FIG. 7. These results show unambiguously that the enzyme acted only on Gly-pNA. No hydrolytic activity could be detected toward all other amino acids, even with long incubation times. This confirms the previous results.
[0103] The inventors also tried to investigate if PhTET4 exhibits high D-stereospecificity against D-Alanine as shown for Aspergillus oryzae glycine aminopeptidase (Marui, J., et al. (2012)). For this, D-Ala-pNA was used as chromogenic substrate in optimal activity conditions (0.1 mM NiCl.sub.2, pH 9.5 and 85.degree. C.) and the results are reported in FIG. 7. The experiment showed that PhTET4 is unable to cleave residue alanine in D-conformation, thus demonstrating that PhTET4 is devoid from D-stereospecificity.
[0104] 5. Endopeptidase and Exopeptidase Activities of PhTET4
[0105] In aminopeptidases, it is known that the catalytic activities and specificities can be affected by the length and N-terminal amino acid composition of the peptide substrate. Consequently, the inventors tested if PhTET4 maintains its narrow specificity toward glycine residues in a peptide context. For this, the inventors measured PhTET4 capacity to cleave N-terminal residue of the following peptides: GI; GL; GLM; GMDSLAFSGGL and LGG. After incubation of PhTET4 with the peptide substrates in optimal activity conditions (0.1 mM NiCl.sub.2, pH 9.5 and 85.degree. C.), the reaction products were separated by reverse phase HPLC and identified by N-terminal sequencing.
[0106] The HPLC chromatographic profiles of the degradation of GLM and GMDSLAFSGGL peptides are shown on FIGS. 8A and 8B, respectively. The sequences of the detected accumulating peptides were determined and the results clearly demonstrated that PhTET4 does not exert amidolytic activity beyond the N-ter glycine in a peptide context. Therefore, PhTET4 is exclusively an exopeptidase. Overall, the results of these experiments showed that no enzymatic activity was detected against peptide that do not start by glycine residue even if a glycine residue is present at P1' position as demonstrated with the LGG tripeptide.
[0107] To assess if PhTET4 can process consequently several glycine residues in a peptide sequence, the inventors tested the enzymatic activity against the chromogenic peptide Gly-Gly-pNA in the same conditions as described previously. In the results, PhTET4 displayed significant activity against this substrate, corresponding to 10% of the total activity exhibited in presence of monoacyl compound Gly-pNA. Taken together, these experiments clearly mark PhTET4 as an aminopeptidase strictly specialized in the hydrolysis of N-terminal glycine residues.
BIBLIOGRAPHY
[0108] Jamdar, S. N. (2009) A novel aminopeptidase from Burkholderia cepacia specific for acidic amino acids. FEMS Microbiol Lett 295, 230-237 [0109] Marui, J., Matsushita-Morita, M., Tada, S., Hattori, R., Suzuki, S., Amano, H., Ishida, H., Yamagata, Y., Takeuchi, M., and Kusumoto, K. (2012) Enzymatic properties of the glycine D-alanine [corrected] aminopeptidase of Aspergillus oryzae and its activity profiles in liquid-cultured mycelia and solid-state rice culture (rice koji). Applied microbiology and biotechnology 93, 655-669 [0110] Ito K, Ma X, Azmi N, Huang H S, Fujii M, Yoshimoto T. (2003) Novel aminopeptidase specific for glycine from Actinomucor elegans. Biosci Biotechnol Biochem. January; 67(1):83-8
Sequence CWU 1
1
61336PRTPYROCOCCUS HORIKOSHII 1Met Glu Arg Ile Val Lys Ile Leu Arg
Glu Ile Leu Glu Ile Pro Ser1 5 10 15Pro Thr Gly Tyr Thr Lys Glu Val
Met Ser Tyr Leu Glu Lys Phe Leu 20 25 30Lys Glu Asn Glu Val Asn Phe
Tyr Tyr Thr Asn Lys Gly Ala Leu Ile 35 40 45Ala Gly Asn His Pro Lys
Pro Glu Leu Val Val Ile Ala His Val Asp 50 55 60Thr Leu Gly Ala Met
Val Lys Glu Ile Leu Pro Asp Gly His Leu Ala65 70 75 80Phe Ser Arg
Ile Gly Gly Leu Val Leu Pro Thr Phe Glu Gly Glu Tyr 85 90 95Cys Thr
Ile Ile Thr Arg Lys Gly Lys Lys Phe Arg Gly Thr Leu Leu 100 105
110Leu Arg Asn Pro Ser Ala His Val Asn Arg Glu Val Gly Lys Lys Glu
115 120 125Arg Lys Glu Glu Asn Met Tyr Ile Arg Leu Asp Glu Leu Val
Glu Lys 130 135 140Arg Glu Asp Thr Glu Lys Leu Gly Ile Arg Pro Gly
Asp Phe Ile Ala145 150 155 160Phe Asp Pro Lys Phe Glu Tyr Val Asn
Gly Phe Val Lys Ser His Phe 165 170 175Leu Asp Asp Lys Ala Ser Val
Ala Ala Ile Leu Asp Leu Ile Ile Asp 180 185 190Met Lys Asp Glu Leu
Glu Lys Tyr Pro Val Ala Phe Phe Phe Ser Pro 195 200 205Tyr Glu Glu
Val Gly His Gly Gly Ser Ala Gly Tyr Pro Pro Thr Thr 210 215 220Lys
Glu Leu Leu Val Val Asp Met Gly Val Val Gly Glu Gly Val Ser225 230
235 240Gly Lys Glu Thr Ala Val Ser Ile Ala Ala Lys Asp Thr Thr Gly
Pro 245 250 255Tyr Asp Tyr Asp Met Thr Asn Arg Leu Ile Glu Leu Ala
Glu Glu Asn 260 265 270Asn Ile Pro Tyr Val Val Asp Val Phe Pro Tyr
Tyr Gly Ser Asp Gly 275 280 285Ser Ala Ala Leu Arg Ala Gly Trp Asp
Phe Arg Val Ala Leu Ile Gly 290 295 300Pro Gly Val His Ala Ser His
Gly Met Glu Arg Thr His Val Lys Gly305 310 315 320Leu Leu Ala Thr
Lys Glu Leu Ile Arg Ala Tyr Ile Lys Trp Lys Gly 325 330
3352332PRTPYROCOCCUS HORIKOSHII 2Met Met Ser Met Ile Glu Lys Leu
Lys Lys Phe Thr Gln Ile Pro Gly1 5 10 15Ile Ser Gly Tyr Glu Glu Arg
Ile Arg Glu Glu Ile Ile Arg Glu Ile 20 25 30Lys Asp Phe Ala Asp Tyr
Lys Val Asp Ala Ile Gly Asn Leu Ile Val 35 40 45Glu Leu Gly Glu Gly
Glu Glu Arg Ile Leu Phe Met Ala His Met Asp 50 55 60Glu Ile Gly Leu
Leu Ile Thr Gly Ile Thr Asp Glu Gly Lys Leu Arg65 70 75 80Phe Arg
Lys Val Gly Gly Ile Asp Asp Arg Leu Leu Tyr Gly Arg His 85 90 95Val
Asn Val Val Thr Glu Lys Gly Ile Leu Asp Gly Val Ile Gly Ala 100 105
110Thr Pro Pro His Leu Ser Leu Glu Arg Asp Lys Ser Val Ile Pro Trp
115 120 125Tyr Asp Leu Val Ile Asp Ile Gly Ala Glu Ser Lys Glu Glu
Ala Leu 130 135 140Glu Leu Val Lys Pro Leu Asp Phe Ala Val Phe Lys
Lys His Phe Ser145 150 155 160Val Leu Asn Gly Lys Tyr Val Ser Thr
Arg Gly Leu Asp Asp Arg Phe 165 170 175Gly Val Val Ala Leu Ile Glu
Ala Ile Lys Asp Leu Val Asp His Glu 180 185 190Leu Glu Gly Lys Val
Ile Phe Ala Phe Thr Val Gln Glu Glu Val Gly 195 200 205Leu Lys Gly
Ala Lys Phe Leu Ala Asn His Tyr Tyr Pro Gln Tyr Ala 210 215 220Phe
Ala Ile Asp Ser Phe Ala Cys Cys Ser Pro Leu Thr Gly Asp Val225 230
235 240Lys Leu Gly Lys Gly Pro Val Ile Arg Ala Val Asp Asn Ser Ala
Ile 245 250 255Tyr Ser Arg Asp Leu Ala Arg Lys Val Trp Ser Ile Ala
Glu Lys Asn 260 265 270Gly Ile Glu Ile Gln Ile Gly Val Thr Gly Gly
Gly Thr Asp Ala Ser 275 280 285Ala Phe Gln Asp Arg Ser Lys Thr Leu
Ala Leu Ser Val Pro Ile Lys 290 295 300Tyr Leu His Ser Glu Val Glu
Thr Leu His Leu Asn Asp Leu Glu Lys305 310 315 320Leu Val Lys Leu
Ile Glu Ala Leu Ala Phe Glu Leu 325 3303353PRTPYROCOCCUS HORIKOSHII
3Met Glu Val Arg Asn Met Val Asp Tyr Glu Leu Leu Lys Lys Val Val1 5
10 15Glu Ala Pro Gly Val Ser Gly Tyr Glu Phe Leu Gly Ile Arg Asp
Val 20 25 30Val Ile Glu Glu Ile Lys Asp Tyr Val Asp Glu Val Lys Val
Asp Lys 35 40 45Leu Gly Asn Val Ile Ala His Lys Lys Gly Glu Gly Pro
Lys Val Met 50 55 60Ile Ala Ala His Met Asp Gln Ile Gly Leu Met Val
Thr His Ile Glu65 70 75 80Lys Asn Gly Phe Leu Arg Val Ala Pro Ile
Gly Gly Val Asp Pro Lys 85 90 95Thr Leu Ile Ala Gln Arg Phe Lys Val
Trp Ile Asp Lys Gly Lys Phe 100 105 110Ile Tyr Gly Val Gly Ala Ser
Val Pro Pro His Ile Gln Lys Pro Glu 115 120 125Asp Arg Lys Lys Ala
Pro Asp Trp Asp Gln Ile Phe Ile Asp Ile Gly 130 135 140Ala Glu Ser
Lys Glu Glu Ala Glu Asp Met Gly Val Lys Ile Gly Thr145 150 155
160Val Ile Thr Trp Asp Gly Arg Leu Glu Arg Leu Gly Lys His Arg Phe
165 170 175Val Ser Ile Ala Phe Asp Asp Arg Ile Ala Val Tyr Thr Ile
Leu Glu 180 185 190Val Ala Lys Gln Leu Lys Asp Ala Lys Ala Asp Val
Tyr Phe Val Ala 195 200 205Thr Val Gln Glu Glu Val Gly Leu Arg Gly
Ala Arg Thr Ser Ala Phe 210 215 220Gly Ile Glu Pro Asp Tyr Gly Phe
Ala Ile Asp Val Thr Ile Ala Ala225 230 235 240Asp Ile Pro Gly Thr
Pro Glu His Lys Gln Val Thr His Leu Gly Lys 245 250 255Gly Thr Ala
Ile Lys Ile Met Asp Arg Ser Val Ile Cys His Pro Thr 260 265 270Ile
Val Arg Trp Leu Glu Glu Leu Ala Lys Lys His Glu Ile Pro Tyr 275 280
285Gln Leu Glu Ile Leu Leu Gly Gly Gly Thr Asp Ala Gly Ala Ile His
290 295 300Leu Thr Lys Ala Gly Val Pro Thr Gly Ala Leu Ser Val Pro
Ala Arg305 310 315 320Tyr Ile His Ser Asn Thr Glu Val Val Asp Glu
Arg Asp Val Asp Ala 325 330 335Thr Val Glu Leu Met Thr Lys Ala Leu
Glu Asn Ile His Glu Leu Lys 340 345 350Ile4354PRTPYROCOCCUS
HORIKOSHII 4Met Asp Leu Lys Gly Gly Glu Ser Met Val Asp Trp Lys Leu
Met Gln1 5 10 15Glu Ile Ile Glu Ala Pro Gly Val Ser Gly Tyr Glu His
Leu Gly Ile 20 25 30Arg Asp Ile Val Val Asp Val Leu Lys Glu Val Ala
Asp Glu Val Lys 35 40 45Val Asp Lys Leu Gly Asn Val Ile Ala His Phe
Lys Gly Ser Ser Pro 50 55 60Arg Ile Met Val Ala Ala His Met Asp Lys
Ile Gly Val Met Val Asn65 70 75 80His Ile Asp Lys Asp Gly Tyr Leu
His Ile Val Pro Ile Gly Gly Val 85 90 95Leu Pro Glu Thr Leu Val Ala
Gln Arg Ile Arg Phe Phe Thr Glu Lys 100 105 110Gly Glu Arg Tyr Gly
Val Val Gly Val Leu Pro Pro His Leu Arg Arg 115 120 125Gly Gln Glu
Asp Lys Gly Ser Lys Ile Asp Trp Asp Gln Ile Val Val 130 135 140Asp
Val Gly Ala Ser Ser Lys Glu Glu Ala Glu Glu Met Gly Phe Arg145 150
155 160Val Gly Thr Val Gly Glu Phe Ala Pro Asn Phe Thr Arg Leu Asn
Glu 165 170 175His Arg Phe Ala Thr Pro Tyr Leu Asp Asp Arg Ile Cys
Leu Tyr Ala 180 185 190Met Ile Glu Ala Ala Arg Gln Leu Gly Asp His
Glu Ala Asp Ile Tyr 195 200 205Ile Val Gly Ser Val Gln Glu Glu Val
Gly Leu Arg Gly Ala Arg Val 210 215 220Ala Ser Tyr Ala Ile Asn Pro
Glu Val Gly Ile Ala Met Asp Val Thr225 230 235 240Phe Ala Lys Gln
Pro His Asp Lys Gly Lys Ile Val Pro Glu Leu Gly 245 250 255Lys Gly
Pro Val Met Asp Val Gly Pro Asn Ile Asn Pro Lys Leu Arg 260 265
270Ala Phe Ala Asp Glu Val Ala Lys Lys Tyr Glu Ile Pro Leu Gln Val
275 280 285Glu Pro Ser Pro Arg Pro Thr Gly Thr Asp Ala Asn Val Met
Gln Ile 290 295 300Asn Lys Glu Gly Val Ala Thr Ala Val Leu Ser Ile
Pro Ile Arg Tyr305 310 315 320Met His Ser Gln Val Glu Leu Ala Asp
Ala Arg Asp Val Asp Asn Thr 325 330 335Ile Lys Leu Ala Lys Ala Leu
Leu Glu Glu Leu Lys Pro Met Asp Phe 340 345 350Thr
Pro51011DNAPYROCOCCUS HORIKOSHII 5atggaaagga tagtcaagat cttaagggaa
atcttagaga taccttctcc aacgggctac 60acgaaggagg taatgagtta cctagaaaaa
tttctaaagg aaaatgaagt aaacttttac 120tatacgaaca agggggccct
aatagccggt aatcatccaa agcctgagct cgttgttata 180gcccacgtag
acacgcttgg ggcaatggtt aaggagatac taccagacgg acacttagca
240ttttcaagga taggagggct cgttctacct acgtttgaag gcgaatactg
tactataata 300acgagaaaag gaaagaagtt tagaggaacg ctcctcctta
gaaatccgag cgctcatgta 360aatagggaag taggtaaaaa ggagagaaaa
gaggagaata tgtatataag attggacgag 420ctcgtggaga agagagagga
tacagaaaag ctggggataa ggccaggaga cttcatagct 480tttgatccca
aatttgaata cgtaaacggc tttgttaaat cacacttcct agatgacaag
540gctagcgtag ctgcaatact cgatctaata atagatatga aggatgaact
cgagaagtat 600ccagttgcat tcttcttctc accgtatgag gaagttggcc
acggaggctc agctggctac 660ccaccaacga ctaaggaact gctcgtggtt
gatatgggag tagtgggtga aggtgtttca 720ggaaaagaaa ccgccgtatc
tatagcggcc aaggatacaa ctgggcctta tgactatgac 780atgacgaaca
ggttaataga gcttgctgaa gagaacaata tcccatatgt agttgacgtg
840ttcccctact atggttccga tggttcagct gcactaagag ctggatggga
tttcagggtt 900gccctaattg ggccaggtgt gcacgcaagc cacggaatgg
agagaaccca cgttaaggga 960ttgttggcaa ctaaagagct tataagggct
tacataaaat ggaaggggta a 1011611PRTArtificial SequenceGenerated
peptide for amidolytic activity assay 6Gly Met Asp Ser Leu Ala Phe
Ser Gly Gly Leu1 5 10
D00000

D00001

D00002

D00003

D00004

D00005

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.