Albumin Variants And Conjugates
Delahay; Karen Ann ; et al.
U.S. patent application number 16/820421 was filed with the patent office on 2021-03-11 for albumin variants and conjugates. The applicant listed for this patent is Albumedix Ltd. Invention is credited to Karen Ann Delahay, Christopher John Arthur Finnis, Karl Michael Nicholls.
| Application Number | 20210070839 16/820421 |
| Document ID | / |
| Family ID | 1000005234505 |
| Filed Date | 2021-03-11 |











View All Diagrams
| United States Patent Application | 20210070839 |
| Kind Code | A1 |
| Delahay; Karen Ann ; et al. | March 11, 2021 |
ALBUMIN VARIANTS AND CONJUGATES
Abstract
The present invention relates to conjugation-competent albumins and albumin-related polypeptides, and their conjugates with at least one moiety, and to polynucleotides encoding them.
| Inventors: | Delahay; Karen Ann; (Nottingham, GB) ; Finnis; Christopher John Arthur; (Nottingham, GB) ; Nicholls; Karl Michael; (Nottingham, GB) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005234505 | ||||||||||
| Appl. No.: | 16/820421 | ||||||||||
| Filed: | March 16, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15753947 | Feb 20, 2018 | 10633428 | ||
| PCT/EP2016/069748 | Aug 19, 2016 | |||
| 16820421 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61K 47/42 20130101; A61K 38/385 20130101; A61K 47/643 20170801; C07K 14/765 20130101; C07K 2319/31 20130101 |
| International Class: | C07K 14/765 20060101 C07K014/765; A61K 47/64 20060101 A61K047/64; A61K 38/38 20060101 A61K038/38; A61K 47/42 20060101 A61K047/42 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Aug 20, 2015 | EP | 15181822.6 |
Claims
1. (canceled)
2. A conjugation-competent polypeptide comprising an amino acid sequence which is at least 90% identical to a human albumin having a sequence as set forth in SEQ ID NO: 2, or a fragment thereof, and comprising a substitution to cysteine at a position equivalent to E294 of SEQ ID NO: 2.
3. The conjugation-competent polypeptide of claim 2, wherein at a position equivalent to position 34 of SEQ ID NO: 2 there is a conjugation-competent cysteine.
4. The conjugation-competent polypeptide of claim 2, wherein at a position equivalent to position 34 of SEQ ID NO: 2 there is not a conjugation-competent cysteine.
5. The conjugation-competent polypeptide of claim 2, in which the polypeptide has at least 98% sequence identity to SEQ ID NO: 2.
6. A method for increasing the plasma half-life of a molecule selected from the group consisting of a bioactive agent, imaging agent, diagnostic agent, contrast agent, and therapeutic compound, the method comprising conjugating, fusing or associating the molecule with a conjugation-competent polypeptide according to claim 2.
7. A fusion polypeptide comprising a conjugation-competent polypeptide of claim 2 and a fusion partner polypeptide.
8. A polynucleotide which encodes the polypeptide of claim 2.
9. A plasmid comprising the polynucleotide of claim 8.
10. A host cell comprising a polynucleotide of claim 8.
11. A conjugate comprising: (i) a bioactive agent, imaging agent, diagnostic agent, contrast agent or therapeutic compound; and (ii) the polypeptide according to claim 2, wherein the bioactive agent, imaging agent, diagnostic agent, contrast agent, or therapeutic compound is linked to the polypeptide through a conjugation-competent cysteine residue of the polypeptide.
12. The conjugate of claim 11, further comprising one or more additional molecules, wherein the additional molecule is a bioactive agent, imaging agent, diagnostic agent, contrast agent or therapeutic compound, each of the one or more additional molecules being linked to the polypeptide through a conjugation-competent cysteine residue of the polypeptide.
13. A method of producing the polynucleotide of claim 2 comprising: providing a nucleic acid molecule encoding a parent albumin or fragment thereof; and modifying the nucleic acid sequence of the nucleic acid molecule to encode a conjugation-competent polypeptide, which is at least 90% identical to human albumin, or a fragment thereof, comprising a conjugation-competent cysteine residue at a position equivalent to E294 of SEQ ID NO. 2.
14. A method of producing the polypeptide of claim 2, comprising: (a) culturing the host cell of claim 10 under conditions that allow expression of the polypeptide; and (b) recovering the polypeptide from the host cell and/or from host cell growth medium.
15. The method of claim 14, further comprising purifying the polypeptide obtained in step (b).
16. A method of producing the conjugate of claim 11, wherein the method comprises linking the polypeptide of claim 2, or the polypeptide produced by the method of claim 13, to a bioactive agent, imaging agent, diagnostic agent, contrast agent or therapeutic compound through a conjugation-competent cysteine residue of the polypeptide.
17. A particle comprising the polypeptide of claim 2, wherein the particle is a nanoparticle, a microparticle, or a liposome.
18. A composition comprising the conjugate of claim 11, and at least one pharmaceutically acceptable carriers or diluents.
19. The conjugate of claim 11, wherein the bioactive agent, imaging agent, diagnostic agent, contrast agent or therapeutic compound is selected from: (i) therapeutic compounds comprising 4-1BB ligand, 5-helix, A human C-C chemokine, A human L105 chemokine, A human L105 chemokine designated huL105_3, A monokine induced by gamma-interferon (MIG), A partial CXCR4B protein, A platelet basic protein (PBP), al-antitrypsin, ACRP-30 Homologue, Complement Component C1q C, Adenoid-expressed chemokine (ADEC), aFGF, FGF-1, AGF, AGF Protein, albumin, an etoposide, angiostatin, Anthrax vaccine, Antibodies specific for collapsin, antistasin, Anti-TGF beta family antibodies, antithrombin III, APM-1, ACRP-30, Famoxin, apo-lipoprotein species, Arylsulfatase B, b57 Protein, BCMA, Beta-thromboglobulin protein (beta-TG), bFGF, FGF2, Blood coagulation factors, BMP Processing Enzyme Furin, BMP-10, BMP-12, BMP-15, BMP-17, BMP-18, BMP-2B, BMP-4, BMP-5, BMP-6, BMP-9, Bone Morphogenic Protein-2, calcitonin, Calpain-10a, Calpain-10b, Calpain-10c, Cancer Vaccine, Carboxypeptidase, C-C chemokine, MCP2, CCR5 variant, CCR7, CCR7, CD11 a Mab, CD137, 4-1BB Receptor Protein, CD20 Mab, CD27, CD27L, CD30, CD30 ligand, CD33 immunotoxin, CD40, CD40L, CD52 Mab, Cerebus Protein, Chemokine Eotaxin, Chemokine hIL-8, Chemokine hMCP1, Chemokine hMCP1a, Chemokine hMCP1b, Chemokine hMCP2, Chemokine hMCP3, Chemokine hSDF1b, Chemokine MCP-4, chemokine TECK and TECK variant, Chemokine-like protein IL-8M1 Full-Length and Mature, Chemokine-like protein IL-8M10 Full-Length and Mature, Chemokine-like protein IL-8M3, Chemokine-like protein IL-8M8 Full-Length and Mature, Chemokine-like protein IL-8M9 Full-Length and Mature, Chemokine-like protein PF4-414 Full-Length and Mature, Chemokine-like protein PF4-426 Full-Length and Mature, Chemokine-like protein PF4-M2 Full-Length and Mature, Cholera vaccine, Chondromodulin-like protein, c-kit ligand, SCF, Mast cell growth factor, MGF, Fibrosarcoma-derived stem cell factor, CNTF and fragment thereof (such as CNTFAx15'(Axokine.TM.)), coagulation factors in both pre and active forms, collagens, Complement C5 Mab, Connective tissue activating protein-III, CTAA16.88 Mab, CTAP-III, CTLA4-Ig, CTLA-8, CXC3, CXC chemokine receptor 3, cyanovirin-N, Darbepoetin, designated exodus, designated huL105 7, DIL-40, Dnase, EDAR, EGF Receptor Mab, ENA-78, Endostatin, Eotaxin, Epithelial neutrophil activating protein-78, EPO receptor, EPOR, erythropoietin (EPO) and EPO mimics, Eutropin, Exodus protein, Factor IX, Factor VII, Factor VIII, Factor X and Factor XIII, FAS Ligand Inhibitory Protein (DcR3), FasL, FGF, FGF-12, Fibroblast growth factor homologous factor-1, FGF-15, FGF-16, FGF-18, FGF-3, INT-2, FGF-4, gelonin, HST-1, HBGF-4, FGF-5, FGF-6, Heparin binding secreted transforming factor-2, FGF-8, FGF-9, Glia activating factor, fibrinogen, flt-1, flt-3 ligand, Follicle stimulating hormone Alpha subunit, Follicle stimulating hormone Beta subunit, Follitropin, Fractalkine, fragment. myofibrillar protein Troponin I, FSH, Galactosidase, Galectin-4, G-CSF, GDF-1, Gene therapy, Glioma-derived growth factor, glucagon, glucagon-like peptides, Glucocerebrosidase, glucose oxidase, Glucosidase, Glycodelin-A, Progesterone-associated endometrial protein, GM-CSF, gonadotropin, Granulocyte chemotactic protein-2 (GCP-2), Granulocyte-macrophage colony stimulating factor, growth hormone, Growth related oncogene-alpha (GRO-alpha), Growth related oncogene-beta (GRO-beta), Growth related oncogene-gamma (GRO-gamma), hAPO-4, TROY, hCG, Hepatitus B surface Antigen, Hepatitus B Vaccine, HER2 Receptor Mab, hirudin, HIV gp120, HIV gp41, HIV Inhibitor Peptide, HIV Inhibitor Peptide, HIV Inhibitor Peptide, HIV protease inhibiting peptides, HIV-1 protease inhibitors, HPV vaccine, Human 6CKine protein, Human Act-2 protein, Human adipogenesis inhibitory factor, human B cell stimulating factor-2 receptor, Human beta-chemokine H1305 (MCP-2), Human C-C chemokine DGWCC, Human CC chemokine ELC protein, Human CC type chemokine interleukin C, Human CCC3 protein, Human CCF18 chemokine, Human CC-type chemokine protein designated SLC (secondary lymphoid chemokine), Human chemokine beta-8 short forms, Human chemokine C10, Human chemokine CC-2, Human chemokine CC-3, Human chemokine CCR-2, Human chemokine Ckbeta-7, Human chemokine ENA-78, Human chemokine eotaxin, Human chemokine GRO alpha, Human chemokine GROalpha, Human chemokine GRObeta, Human chemokine HCC-1, Human chemokine HCC-1, Human chemokine 1-309, Human chemokine IP-10, Human chemokine L105 3, Human chemokine L105 7, Human chemokine MIG, Human chemokine MIG-beta protein, Human chemokine MIP-1 alpha, Human chemokine MIP1beta, Human chemokine MIP-3alpha, Human chemokine MIP-3beta, Human chemokine PF4, Human chemokine protein 331D5, Human chemokine protein 61164, Human chemokine receptor CXCR3, Human chemokine SDF1alpha, Human chemokine SDF1beta, Human chemokine ZSIG-35, Human Chr19Kine protein, Human CKbeta-9, Human CX3C 111 amino acid chemokine, Human DNAX interleukin-40, Human DVic-1 C-C chemokine, Human EDIRF I protein sequence, Human EDIRF II protein sequence, Human eosinocyte CC type chemokine eotaxin, Human eosinophil-expressed chemokine (EEC), Human fast twitch skeletal muscle troponin C, Human fast twitch skeletal muscle troponin I, Human fast twitch skeletal muscle Troponin subunit C, Human fast twitch skeletal muscle Troponin subunit I Protein, Human fast twitch skeletal muscle Troponin subunit T, Human fast twitch skeletal muscle troponin T, Human foetal spleen expressed chemokine, FSEC, Human GM-CSF receptor, Human gro-alpha chemokine, Human gro-beta chemokine, Human gro-gamma chemokine, Human IL-16 protein, Human IL-1RD10 protein sequence, Human IL-1RD9, Human IL-5 receptor alpha chain, Human IL-6 receptor, Human IL-8 receptor protein hIL8RA, Human IL-8 receptor protein hIL8RB, Human IL-9 receptor protein, Human IL-9 receptor protein variant #3, Human IL-9 receptor protein variant fragment, Human IL-9 receptor protein variant fragment #3, Human interleukin 1 delta, Human interleukin 10, Human interleukin 18, Human interleukin 18 derivatives, Human interleukin-1 beta precursor, Human interleukin-1 beta precursor, Human interleukin-1 receptor accessory protein, Human interleukin-1 receptor antagonist beta, Human interleukin-1 type-3 receptor, Human interleukin-10 (precursor), Human interleukin-11 receptor, Human interleukin-12 40 kD subunit, Human interleukin-12 beta-1 receptor, Human interleukin-12 beta-2 receptor, Human interleukin-12 p35 protein, Human interleukin-12 p40 protein, Human interleukin-12 receptor, Human interleukin-13 alpha receptor, Human interleukin-13 beta receptor, Human interleukin-15, Human interleukin-15 receptor from clone P1, Human interleukin-17 receptor, Human interleukin-18 protein (IL-18), Human interleukin-3, human interleukin-3 receptor, Human interleukin-3 variant, Human interleukin-4 receptor, Human interleukin-5, Human interleukin-6, Human interleukin-7, Human interleukin-7, Human interleukin-8 (IL-8), Human intracellular IL-1 receptor antagonist, Human IP-10 and HIV-1 gp120 hypervariable region fusion protein, Human IP-10 and human Muc-1 core epitope (VNT) fusion protein, human liver and activation regulated chemokine (LARC), Human Lkn-1 Full-Length and Mature protein, Human mammary associated chemokine (MACK) protein Full-Length and Mature, Human mature chemokine Ckbeta-7, Human mature gro-alpha, Human mature gro-gamma polypeptide used to treat sepsis, Human MCP-3 and human Muc-1 core epitope (VNT) fusion protein, Human MI10 protein, Human MI1A protein, Human monocyte chemoattractant factor hMCP-1, Human monocyte chemoattractant factor hMCP-3, Human monocyte chemotactic proprotein (MCPP) sequence, Human neurotactin chemokine like domain, Human non-ELR CXC chemokine H174, Human non-ELR CXC chemokine IP10, Human non-ELR CXC chemokine Mig, Human PAI-1 mutants, Human protein with IL-16 activity, Human protein with IL-16 activity, Human secondary lymphoid chemokine (SLC), Human SISD protein, Human STCP-1, Human stromal cell-derived chemokine, SDF-1, Human T cell mixed lymphocyte reaction expressed chemokine (TMEC), Human thymus and activation regulated cytokine (TARC), Human thymus expressed, Human TNF-alpha, Human TNF-beta (LT-alpha), Human type CC chemokine eotaxin 3 protein sequence, Human type II interleukin-1 receptor, Human wild-type interleukin-4 (hIL-4) protein, Human ZCHEMO-8 protein, Humanized Anti-VEGF Antibodies, and fragments thereof, Humanized Anti-VEGF Antibodies, and fragments thereof, Hyaluronidase, ICE 10 kD subunit, ICE 20 kD subunit, ICE 22 kD subunit, Iduronate-2-sulfatase, Iduronidase, IL-1 alpha, IL-1 beta, IL-1 inhibitor (IL-li), IL-1 mature, IL-10 receptor, IL-11, IL-11, IL-12 p40 subunit, IL-13, IL-14, IL-15, IL-15 receptor, IL-17, IL-17 receptor, receptor, IL-19, IL-li fragments, IL1-receptor antagonist, IL-21 (TIF), IL-3 containing fusion protein, IL-3 mutant proteins, IL-3 variants, IL-4, IL-4 muteins, IL-4 mutein Y124G, IL-4 mutein Y124X, IL-5, IL-5 muteins, 11-5 receptor, IL-6, 11-6 receptor, IL-7 receptor clone, IL-8 receptor, IL-9 mature protein variant (Met117 version), immunoglobulins or immunoglobulin-based molecules or fragment of either (e.g. a Small Modular ImmunoPharmaceutical.TM. ("SMIP") or dAb, Fab' fragments, F(ab')2, scAb, scFv or scFv fragment), including but not limited to plasminogen, Influenza Vaccine, Inhibin alpha, Inhibin beta, insulin, insulin-like growth factor, Integrin Mab, inter-alpha trypsin inhibitor, inter-alpha trypsin inhibitor, Interferon gamma-inducible protein (IP-10), interferons (such as interferon alpha species and sub-species, interferon beta species and sub-species, interferon gamma species and sub-species), interleukin 6, interleukin 8 (IL-8) receptor, interleukin 8 receptor B, interleukin-1alpha, interleukin-2 receptor associated protein p43, interleukin-3, interleukin-4 muteins, interleukin-8 (IL-8) protein, interleukin-9, interleukin-9 (IL-9) mature protein (Thr117 version), interleukins (such as IL10, IL11 and IL2), Japanese encephalitis vaccine, Kalikrein Inhibitor, Keratinocyte growth factor, Kunitz domain protein (such as aprotinin, amyloid precursor protein and those described in WO 03/066824, with or without albumin fusions), LACI, lactoferrin, Latent TGF-beta binding protein II, leptin, Liver expressed chemokine-1 (LVEC-1), Liver expressed chemokine-2 (LVEC-2), LT-alpha, LT-beta, Luteinization Hormone, Lyme Vaccine, Lymphotactin, Macrophage derived chemokine analogue MDC (n+1), Macrophage derived chemokine analogue MDC-eyfy, Macrophage derived chemokine analogue MDC-yl, Macrophage-derived chemokine (MDC), Maspin, Protease Inhibitor 5, MCP-1 receptor, MCP-1a, MCP-1b, MCP-3, MCP-4 receptor, M-CSF, Melanoma inhibiting protein, Membrane-bound proteins, Met117 human interleukin 9, MIP-3 alpha, MIP-3 beta, MIP-Gamma, MIRAP, Modified Rantes, monoclonal antibody, MP52, Mutant interleukin 6 S176R, myofibrillar contractile protein Troponin I, Natriuretic Peptide, Nerve Growth Factor-beta, Nerve Growth Factor-beta2, Neuropilin-1, Neuropilin-2, Neurotactin, Neurotrophin-3, Neurotrophin-4, Neurotrophin-4a, Neurotrophin-4b, Neurotrophin-4c, Neurotrophin-4d, Neutrophil activating peptide-2 (NAP-2), NOGO-66 Receptor, NOGO-A, NOGO-B, NOGO-C, Novel beta-chemokine designated PTEC, N-terminal modified chemokine GroHEK/hSDF-1 alpha, N-terminal modified chemokine GroHEK/hSDF-1beta, N-terminal modified chemokine met-hSDF-1 alpha, N-terminal modified chemokine met-hSDF-1 beta, OPGL, Osteogenic Protein-1 (OP-1), BMP-7, Osteogenic Protein-2, OX40, ACT-4, OX40L, Oxytocin (Neurophysin I), parathyroid hormone, Patched, Patched-2, PDGF-D, Pertussis toxoid, Pituitary expressed chemokine (PGEC), Placental Growth Factor, Placental Growth Factor-2, Plasminogen Activator Inhibitor-1 (PAI-1), Plasminogen Activator Inhibitor-2 (PAI-2), Platelet derived growth factor, Platelet derived growth factor Bv-sis, Platelet derived growth factor precursor A, Platelet derived growth factor precursor B, Platelet Mab, platelet-derived endothelial cell growth factor (PD-ECGF), Platelet-Derived Growth Factor A chain, Platelet-Derived Growth Factor B chain, polypeptide used to treat sepsis, Preproapolipoprotein "milano" variant, Preproapolipoprotein "paris" variant, pre-thrombin, Primate CC chemokine "ILINCK", Primate CXC chemokine "IBICK", proinsulin, Prolactin, Prolactin2, prosaptide, Protease inhibitor peptides, Protein C, Protein S, pro-thrombin, prourokinase, RANTES, RANTES 8-68, RANTES 9-68, RANTES peptide, RANTES receptor, Recombinant interleukin-16, Resistin, restrictocin, Retroviral protease inhibitors, ricin, Rotavirus Vaccine, RSV Mab, saporin, sarcin, Secreted and Transmembrane polypeptides, serum cholinesterase, serum protein (such as a blood clotting factor), Soluble BMP Receptor Kinase Protein-3, Soluble VEGF Receptor, Stem Cell Inhibitory Factor, Straphylococcus Vaccine, Stromal Derived Factor-1 alpha, Stromal Derived Factor-1 beta, Substance P (tachykinin), T1249 peptide, T20 peptide, T4 Endonuclease, TACI, Tarc, TGF-beta 1, TGF-beta 2, Thr117 human interleukin 9, thrombin, thrombopoietin, thrombopoietin derivative 1, thrombopoietin derivative 2, thrombopoietin derivative 3, thrombopoietin derivative 4, thrombopoietin derivative 5, thrombopoietin derivative 6, thrombopoietin derivative 7, Thymus expressed chemokine (TECK), Thyroid stimulating Hormone, tick anticoagulant peptide, Tim-1 protein, TNF-alpha precursor, TNF-R, TNF-RII, TNF p75 Receptor, Death Receptor, tissue plasminogen activator (tPA), transferrin, transforming growth factor beta, Troponin peptides, Truncated monocyte chemotactic protein 2 (6-76), Truncated RANTES protein (3-68), tumour necrosis factor, Urate Oxidase, urokinase, Vasopressin (Neurophysin II), VEGF R-3, flt-4, VEGF Receptor, KDR, flk-1, VEGF-110, VEGF-121, VEGF-138, VEGF-145, VEGF-162, VEGF-165, VEGF-182, VEGF-189, VEGF-206, VEGF-D, VEGF-E, VEGF-X, von Willebrand's factor, Wild type monocyte chemotactic protein 2, or ZTGF-beta 9; (ii) chemotherapy drugs comprising 13-cis-Retinoic Acid, 2-CdA, 2-Chlorodeoxyadenosine, 5-Azacitidine, 5-Fluorouracil, 5-FU, 6-Mercaptopurine, 6-MP, 6-TG, 6-Thioguanine, Abraxane, Accutane.RTM., Actinomycin-D, Adriamycin.RTM., Adrucil.RTM., Agrylin.RTM., Ala-Cort.RTM., Aldesleukin, Alemtuzumab, ALIMTA, Alitretinoin, Alkaban-AQ.RTM., Alkeran.RTM., All-transretinoic Acid, Alpha Interferon, Altretamine, Amethopterin, Amifostine, Aminoglutethimide, Anagrelide, Anandron.RTM., Anastrozole, Arabinosylcytosine, Ara-C, Aranesp.RTM., Aredia.RTM., Arimidex.RTM., Aromasin.RTM., Arranon.RTM., Arsenic Trioxide, Asparaginase, ATRA, Avastin.RTM., Azacitidine, BCG, BCNU, Bevacizumab, Bexarotene, BEXXAR.RTM., Bicalutamide, BiCNU, Blenoxane.RTM., Bleomycin, Bortezomib, Busulfan, Busulfex.RTM., C225, Calcium Leucovorin, Campath.RTM., Camptosar.RTM., Camptothecin-11, Capecitabine, Carac.TM., Carboplatin, Carmustine, Carmustine Wafer, Casodex.RTM., CC-5013, CCNU, CDDP, CeeNU, Cerubidine.RTM., Cetuximab, Chlorambucil, Cisplatin, Citrovorum Factor, Cladribine, Cortisone, Cosmegen.RTM., CPT-11, Cyclophosphamide, Cytadren.RTM., Cytarabine, Cytarabine Liposomal, Cytosar-U.RTM., Cytoxan.RTM., Dacarbazine, Dacogen, Dactinomycin, Darbepoetin Alfa, Dasatinib, Daunomycin, Daunorubicin, Daunorubicin Hydrochloride, Daunorubicin Liposomal, DaunoXome.RTM., Decadron, Decitabine, Delta-Cortef.RTM., Deltasone.RTM., Denileukin diftitox, DepoCyt.TM., Dexamethasone, Dexamethasone acetate, Dexamethasone Sodium Phosphate, Dexasone, Dexrazoxane, DHAD, DIC, Diodex, Docetaxel, Doxil.RTM., Doxorubicin, Doxorubicin liposomal, Droxia.TM., DTIC, DTICDome.RTM., Duralone.RTM., Efudex.RTM., Eligard.TM., Ellence.TM., Eloxatin.TM., Elspar.RTM., Emcyt.RTM., Epirubicin, Epoetin alfa, Erbitux.TM., Erlotinib, Erwinia L-asparaginase, Estramustine, Ethyol, Etopophos.RTM., Etoposide, Etoposide Phosphate, Eulexin.RTM., Evista.RTM., Exemestane, Fareston.RTM., Faslodex.RTM., Femara.RTM., Filgrastim, Floxuridine, Fludara.RTM., Fludarabine, Fluoroplex.RTM., Fluorouracil, Fluoxymesterone, Flutamide, Folinic Acid, FUDR.RTM., Fulvestrant, G-CSF, Gefitinib, Gemcitabine, Gemtuzumab ozogamicin, Gemzar.RTM., Gleevec.TM., Gliadel.RTM. Wafer, GM-CSF, Goserelin, Granulocyte-Colony Stimulating Factor, Granulocyte Macrophage Colony Stimulating Factor, Halotestin.RTM., Herceptin.RTM., Hexadrol, Hexalen.RTM., Hexamethylmelamine, HMM, Hycamtin.RTM., Hydrea.RTM., Hydrocort Acetate.RTM., Hydrocortisone, Hydrocortisone Sodium Phosphate, Hydrocortisone Sodium Succinate, Hydrocortone Phosphate, Hydroxyurea, Ibritumomab, Ibritumomab Tiuxetan, Idamycin.RTM., Idarubicin, Ifex.RTM., IFN-alpha, Ifosfamide, IL-11, IL-2, Imatinib mesylate, Imidazole Carboxamide, Interferon alfa, Interferon Alfa-2b (PEG Conjugate), interleukin-2, interleukin-11, Intron A.RTM. (interferon alfa-2b), Iressa.RTM., Irinotecan, Isotretinoin, Kidrolase.RTM., Lanacort.RTM., Lapatinib, L-asparaginase, LCR, Lenalidomide, Letrozole, Leucovorin, Leukeran, Leukine.TM., Leuprolide, Leurocristine, Leustatin.TM., Liposomal Ara-C, Liquid Pred.RTM., Lomustine, L-PAM, L-Sarcolysin, Lupron.RTM., Lupron Depot.RTM., Matulane.RTM., Maxidex, Mechlorethamine, Mechlorethamine Hydrochloride, Medralone.RTM., Medrol.RTM., Megace.RTM., Megestrol, Megestrol Acetate, Melphalan, Mercaptopurine, Mesna, Mesnex.TM., Methotrexate, Methotrexate Sodium, Methylprednisolone,
Meticorten.RTM., Mitomycin, Mitomycin-C, Mitoxantrone, M-Prednisol.RTM., MTC, MTX, Mustargen.RTM., Mustine, Mutamycin.RTM., Myleran.RTM., Mylocel.TM., Mylotarg.RTM., Navelbine.RTM., Nelarabine, Neosar.RTM., Neulasta.TM., Neumega.RTM., Neupogen.RTM., Nexavar.RTM., Nilandron.RTM., Nilutamide, Nipent.RTM., Nitrogen Mustard, Novaldex.RTM., Novantrone.RTM., Octreotide, Octreotide acetate, Oncospar.RTM., Oncovin.RTM., Ontak.RTM., Onxal.TM., Oprevelkin, Orapred.RTM., Orasone.RTM., Oxaliplatin, Paclitaxel, Paclitaxel Protein-bound, Pamidronate, Panitumumab, Panretin.RTM., Paraplatin.RTM., Pediapred.RTM., PEG Interferon, Pegaspargase, Pegfilgrastim, PEG-INTRON.TM., PEG-L-asparaginase, PEMETREXED, Pentostatin, Phenylalanine Mustard, Platinol.RTM., Platinol-AQ.RTM., Prednisolone, Prednisone, Prelone.RTM., Procarbazine, PROCRIT.RTM., Proleukin.RTM., Prolifeprospan 20 with Carmustine Implant, Purinethol.RTM., Raloxifene, Revlimid.RTM., Rheumatrex.RTM., Rituxan.RTM., Rituximab, RoferonA.RTM. (Interferon Alfa-2a), Rubex.RTM., Rubidomycin hydrochloride, Sandostatin.RTM., Sandostatin LAR.RTM., Sargramostim, Solu-Cortef.RTM., Solu-Medrol Sorafenib, SPRYCEL.TM., STI-571, Streptozocin, SU11248, Sunitinib, Sutent.RTM., Tamoxifen, Tarceva.RTM., Targretin.RTM., Taxol.RTM., Taxotere.RTM., Temodar.RTM., Temozolomide, Teniposide, TESPA, Thalidomide, Thalomid.RTM., TheraCys.RTM., Thioguanine, Thioguanine Tabloid.RTM., Thiophosphoamide, Thioplex.RTM., Thiotepa, TICE.RTM., Toposar.RTM., Topotecan, Toremifene, Tositumomab, Trastuzumab, Tretinoin, Trexall.TM., Trisenox.RTM., TSPA, TYKERB.RTM., VCR, Vectibix.TM., Velban.RTM., Velcade.RTM., VePesid.RTM., Vesanoid.RTM., Viadur.TM., Vidaza.RTM., Vinblastine, Vinblastine Sulfate, Vincasar Pfs.RTM., Vincristine, Vinorelbine, Vinorelbine tartrate, VLB, VM-26, Vorinostat, VP-16, Vumon.RTM., Xeloda.RTM., Zanosar.RTM., Zevalin.TM., Zinecard.RTM., Zoladex.RTM., Zoledronic acid, Zolinza, or Zometa.RTM.; (iii) radiopharmaceuticals comprising Carbon-11, Carbon-14, Chromium-51, Cobalt-57, Cobalt-58, Erbium-169, Fluorine-18, Gallium-67, Gold-198, Indium-111, Indium-113m, Iodine-123, Iodine-125, Iodine-131, Iron-59, Krypton-81m, Nitrogen-13, Oxygen-15, Phosphorous-32, Rhenium-186, Rubidium-82, Samarium-153, Selenium-75, Strontium-89, Technetium-99m, Thallium-201, Tritium, Xenon-127, or Xenon-133, Yttrium-90, and (iv) imaging agents comprising Gadolinium, magnetite, manganese, technetium, 1125, 1131, P32, TI201, Iopamidol, or PET-FDG.
Description
REFERENCE TO SEQUENCE LISTING
[0001] This application contains a Sequence Listing in computer readable form. The computer readable form is incorporated herein by reference.
FIELD OF THE INVENTION
[0002] The present invention relates to conjugation-competent albumins and albumin-related polypeptides, and their conjugates with at least one (e.g. several) moiety, and to polynucleotides encoding them.
BACKGROUND OF THE INVENTION
[0003] Serum albumins provide valuable scaffolds to which bioactive molecules may be fused, either through genetic fusions or chemical fusions to improve the properties of the fused molecule(s) (Leger, R. et al. (2004), Bioorg Med Chem Lett 14(17): 4395-8; Thibaudeau, K., et al. (2005). Bioconjug Chem 16(4): 1000-8; Balan, V. et al. (2006), Antivir Ther 11(1): 35-45; EP 0413622; WO 90/13653; EP 1681304; WO 1997/024445). Albumin has a long plasma half-life of about 19 days and because of this property it has been suggested for use in drug delivery.
[0004] The human serum albumin (HSA) polypeptide chain has 35 cysteine residues, which form 17 disulphide bonds and one unpaired (free) cysteine at position 34 of the mature protein (SEQ ID NO. 2). Cysteine-34 has been used for conjugation of molecules to albumin (Leger et al. (2004) Bioorg Med Chem Lett 14(17): 4395-8; Thibaudeau et al. (2005), Bioconjug Chem 16(4): 1000-8), and provides a precise, well defined site for conjugation. However, conjugation at cysteine-34 provides only one site for attachment of a single moiety and thus there is no choice of conjugation site. Also, the provision of a single conjugation site means that only one moiety can be conjugated to each albumin molecule. WO 2009/126920 and WO 2010/059315 propose the substitution for cysteine of one or more (e.g. several) selected surface-exposed threonine or serine residues in albumin. However, the actual production of such variants is not disclosed. WO 2010/092135 discloses albumin variants comprising three or more (several) conjugation-competent cysteine residues: cysteine-34 and at least two further cysteine residues; or variants in which another amino acid is substituted for the cysteine-34, and there are at least three further free cysteines.
[0005] Pharmaceutical agents, or their precursors, are generally prepared as homogeneous species, to allow for quality control. In HSA, the free cysteine at position 34 is located in a hydrophobic crevice with a depth of 9.5 .ANG. (Cornell C N, Chang R, Kaplan L J. 1981. Arch. Biochem. Biophys. 209(1):1-6.), and is not thought to be involved in homodimerization of HSA. However, surface-exposed cysteine residues in polypeptides may form stable inter-molecular disulphide bridges, as occur naturally for example between the heavy and light chains of immunoglobulin. It is desirable to provide albumin variants having introduced cysteine residues which have a low propensity to form dimers or oligomers.
[0006] WO 2000/69902 discloses conjugation of pharmaceutically beneficial compounds to HSA at cysteine-34, and it was found that the conjugates maintained the long plasma half-life of albumin. The resulting plasma half-life of the conjugate was generally considerably longer than the plasma half-life of the beneficial therapeutic compound alone. Further, albumin has been genetically fused to therapeutically beneficial peptides (WO 2001/79271A and WO 2003/59934) with the typical result that the fusion has the activity of the therapeutically beneficial peptide and a considerably longer plasma half-life than the plasma half-life of the therapeutically beneficial peptide alone.
[0007] Albumin binds in vivo to its receptor, the neonatal Fc receptor (FcRn) "Brambell" and this interaction is known to be important for, the plasma half-life of albumin. FcRn is a membrane bound protein, expressed in many cell and tissue types. FcRn has been found to salvage albumin from intracellular degradation (Roopenian D. C. and Akilesh, S. (2007), Nat. Rev. Immunol 7, 715-725.). FcRn is a bifunctional molecule that contributes to maintaining a high level of IgGs and albumin in plasma in mammals such as humans. Data indicate that IgG and albumin bind non-cooperatively to distinct sites on FcRn (Andersen et al. (2006), Eur. J. Immunol 36, 3044-3051; Chaudhury et al. (2006), Biochemistry 45, 4983-4990). Andersen et al. (2010), Journal of Biological Chemistry 285(7): 4826-36, describes the affinity of human and mouse FcRn for each of mouse and human albumin (all possible combinations). No binding of albumin from either species was observed at physiological pH to either receptor. At acidic pH, a 100-fold difference in binding affinity was observed.
[0008] The major FcRn receptor binding site in albumin is localized within Domain III (DIII, 381-585), (Andersen et al. (2010), Clinical Biochemistry 43, 367-372). A number of key amino acid residues have been shown to be important in binding, notably histidines H464, H510 and H536 and lysine K500 of human albumin (Andersen et al. (2010), Nat. Commun. 3:610. DOI:10.1038/ncomms1607). Generally, the higher the affinity of an albumin for FcRn, the longer is its plasma half-life. WO 2011/124718 discloses a class of variant albumins having modulated binding affinity to FcRn; the variants comprise domain III of an albumin with one or more (e.g. several) other domains of albumin and optionally include one or more (e.g. several) point mutations. WO 2012/059486 discloses variants of albumin in which a C-terminal portion of Domain III is swapped with a corresponding portion of an albumin of a different animal species. WO 2013/075066, WO 2011/103076, WO 2012/112188, WO 2011/051489 and WO 2014/072481 disclose point mutations within Domain III, or combinations of such point mutations, which alter the binding affinity of albumin to FcRn.
[0009] Various amino acid residues of albumin located in Domain I or Domain II have also recently been found to affect its interaction with FcRn. WO 2013/135896 discloses albumin variants having one or more (e.g. several) alterations in Domain I and one or more (e.g. several) alterations in Domain III. WO 2015/036579 discloses albumin variants having one or more (e.g. several) alterations in Domain II.
[0010] The listing or discussion of an apparently prior-published document in this specification should not necessarily be taken as an acknowledgement that the document is part of the state of the art or is common general knowledge.
[0011] It is desirable to provide albumin variants having one or more (e.g. several) introduced cysteine residues in which an introduced free cysteine residue does not itself have a major impact on FcRn binding of albumin, or be positioned such that conjugation of a partner molecule to the free cysteine will sterically hinder FcRn binding. Such considerations could reduce the risk of unpredictable effects when introducing combinations of more than one free cysteine in a single albumin variant. Such variant polypeptides may be further modified to include alterations known to affect the binding affinity of albumin for FcRn, so as to allow the plasma half-life of the polypeptide, or conjugates thereof, to be tailored for specific applications.
SUMMARY OF THE INVENTION
[0012] Based on an analysis of the three-dimensional structure of a human serum albumin (HSA) bound to FcRn, the inventors have designed variant polypeptides (muteins) of albumin which have one or more (e.g. several) conjugation-competent cysteine residues. The term `thio-albumin` is used herein to describe an albumin variant which comprises one or more (e.g. several) unpaired cysteine residues, particularly an albumin variant in which one or more (e.g. several) of the unpaired cysteine residues does not occur in a naturally occurring variant of an albumin. Thus a thio-albumin is a `conjugation-competent albumin`. A thio-albumin may be referred to as a `cysteine variant of an albumin`. More particularly, the invention relates to a conjugation-competent polypeptide comprising an amino acid sequence which is at least 60% identical to human albumin, particularly residues 1 to 585 of the mature human albumin polypeptide sequence of SEQ ID NO. 2, or a fragment thereof; wherein at least one position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398 of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue; and wherein the conjugation-competent polypeptide preferably has a tendency to exist as a monomer in solution which is at least 70% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution.
[0013] More preferably, the polypeptide has a tendency to exist as a monomer in solution which is at least 75% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution and at least one position equivalent to a position selected from K93, E294, A226, E230, I271, E358, L24, F49, V54, D56, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, L284, K317, A322, E333, D340, E354, K359, A362, E382, and L398 comprises a conjugation-competent cysteine residue.
[0014] The invention also relates to a conjugation-competent polypeptide comprising an amino acid sequence as defined above, and at least one (e.g. several) further modification compared to SEQ ID NO. 2, such as a further modification which causes the polypeptide to have at least one (e.g. several) further conjugation-competent cysteine, or alters the binding affinity of the polypeptide for FcRn, or alters the plasma half-life of the polypeptide. The present invention also relates to isolated polynucleotides encoding the variants; nucleic acid constructs, vectors, and host cells comprising the polynucleotides; and methods of producing the variants.
[0015] The invention also relates to conjugates or associates comprising the variant albumin or fragment thereof according to the invention and a beneficial therapeutic moiety or to a fusion polypeptide comprising a variant albumin or fragment thereof of the invention and a fusion partner polypeptide.
[0016] The invention further relates to compositions comprising the variant albumin, fragment thereof, fusion polypeptide comprising variant albumin or fragment thereof or conjugates comprising the variant albumin or fragment thereof, according to the invention or associates comprising the variant albumin or fragment thereof, according to the invention. The compositions are preferably pharmaceutical compositions.
[0017] The invention further relates to a pharmaceutical composition comprising a variant albumin, fragment thereof, fusion polypeptide comprising variant albumin or fragment thereof or conjugates comprising the variant albumin or fragment thereof, or associates comprising the variant albumin or fragment thereof.
[0018] The invention also relates to the use of the variants, fragments, fusion polypeptides, conjugates, associates, nanoparticles and microparticles.
[0019] The invention also relates to a method for preparing a variant albumin, fragment thereof, fusion polypeptide comprising variant albumin or fragment thereof or conjugates comprising the variant albumin or fragment thereof, or associates comprising the variant albumin or fragment thereof.
BRIEF DESCRIPTION OF DRAWINGS
[0020] FIG. 1. Multiple alignment of amino acid sequences of (i) full length mature HSA (Hu_1_2_3), (ii) an albumin variant comprising domain I and domain III of HSA (Hu_1_3), (iii) an albumin variant comprising domain II and domain III of HSA (Hu_2_3), (iv) full-length Macaca mulatta albumin (Mac_mul), (v) full-length Rattus norvegicus albumin (Rat) and (vi) full-length Mus musculus albumin (Mouse). Positions 500, 550 and 573 (relative to full length HSA) are indicated by arrows.
[0021] FIG. 2. Multiple alignment of amino acid sequence of mature albumin from human, sheep, mouse, rabbit and goat and immature albumins from chimpanzee ("Chimp"), macaque, hamster, guinea pig, rat, cow, horse, donkey, dog, chicken, and pig. The Start and End amino acids of domains 1, 2 and 3 (as defined by Dockal et al (The Journal of Biological. Chemistry, 1999, Vol. 274(41): 29303-29310)) are indicated with respect to mature human albumin.
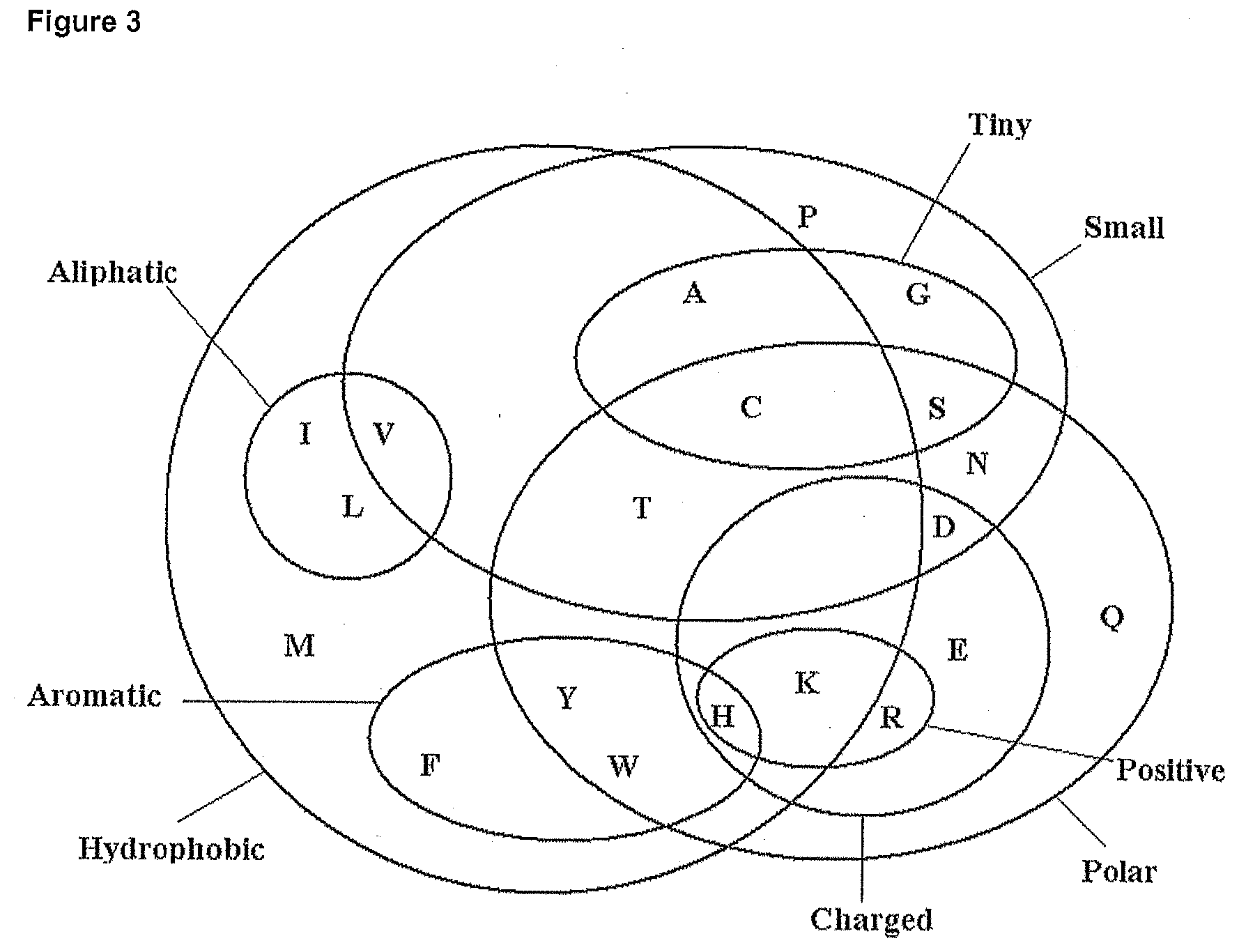
[0022] FIG. 3. Venn diagram showing the classes of and relationship between twenty amino acids.
[0023] FIG. 4. A: Reaction scheme for biotinylation of a protein comprising a free thiol group with maleimide-PEG2-biotin. B: Schematic illustrating potential retro-Michael and succinimide hydrolysis reactions of conjugates formed in scheme A.
[0024] In A, the maleimide forms an adduct with the thiol group, thus forming a succinimide moiety with a thio-ether bond.
[0025] B illustrates adduct formation. The adduct may revert back to maleimide and free thiol via a retro-Michael pathway. Alternatively, the succinimide moiety may undergo stabilizing ring opening to succinic acid, by hydrolysis at pH 9. The thio-ether bond of the conjugate is retained and the succinic acid moiety is unreactive to other thiol compounds which may be present. Free maleimide, when subjected to hydrolysis, also becomes thiol unreactive.
[0026] FIG. 5. MS spectra of purified variants (A: C34A+I271C variant; B: C34A+K93C variant) conjugated with maleimide-PEG2-biotin. A: The conjugate peak is 66924.1. The shorter peak is unconjugated protein. The relative peak heights indicate a conjugated proportion of 72%.+MS, 7.7-9.2 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). B: The conjugate peak is 66908.3, and there is no free proportion, indicating 100% conjugation. +MS, 7.6-9.4 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0027] FIG. 6. MS spectra of purified albumins (A: wild type; B: C34A+E294C variant) conjugated with maleimide-PEG2-biotin and subjected to controlled hydrolysis. In A, 53% of the albumin is present as a thiol-stable conjugate with a peak of 66978.4; and 47% is present as a free albumin following retro-Michael deconjugation. +MS, 7.0-9.6 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In B, 100% of the C34A+E294C variant is present as a thiol-stable conjugate with a peak of 66925.7. +MS, 7.6-9.5 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0028] FIG. 7. MS spectra of purified albumin variants (A: K93C+E294C; B: K93C+E294C; C: C34A+K93C+E294C) conjugated with maleimide-PEG2-biotin and subjected to controlled hydrolysis (B and C). In A, a single peak of 67967.7 for K93C+E294C indicates 100% conjugation to each of the three free thiols. +MS, 1.6-2.6 min, Baseline subtracted (0.40), Deconvoluted (MaxEnt). Smoothed (0.00,1,GA). In B, 20% of the triple conjugate of K93C+E294C is thiol stable after hydrolysis. The main peak, at 67476.2, is indicative of two thiol stable conjugate bonds, and the loss of one maleimide-PEG2-biotin through retro-Michael deconjugation. +MS, 1.8-2.9 min, Baseline subtracted (0.40), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In C, the double conjugate of C34A+K93C+E294C is the major species, at a peak of 67443.1, and the other species is the single conjugate at a peak of 66894.6. +MS, 1.7-2.8 min, Baseline subtracted (0.40), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0029] FIG. 8. MS spectra of purified albumin variant K93C+E294C+K573P (which includes native Cys34). A: indicates 100% conjugation to each of the three free thiols. +MS, 7.3-9.7 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In B, 23% of the triple conjugate of K93C+E294C+K573P (which includes native Cys34) is thiol stable after hydrolysis. The main peak, at 67447.3, is indicative of two thiol stable conjugate bonds, and the loss of one maleimide-PEG2-biotin through retro-Michael deconjugation. +MS, 7.4-9.5 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0030] FIG. 9. A: Schematic illustrating Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye. The MS spectra of purified albumin variants (B: K573P; C: K93C+E294C+K573P) conjugated with Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye are shown. In B, a single peak of 67468.5 for K573P indicates 100% conjugation to the single free thiol at Cys34. +MS, 7.6-9.7 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In C, the triple conjugate of K93C+E294C+K573P (which includes native Cys34) is the major species, at a peak of 69535.8. The shorter peak is double conjugate. The relative peak heights indicate 58% triple conjugate and 42% double conjugate respectively. +MS, 7.6-9.3 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0031] FIG. 10. A: Schematic illustrating 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye. The MS spectra of purified albumin variants (B: K573P; C: C34A+K93C+E294C+K573P) conjugated with 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye are shown. In B, a single peak of 67310.6 for K573P indicates 100% conjugation to the single free thiol at Cys34. +MS, 7.2-9.3 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In C, the double conjugate of C34A+K93C+E294C+K573P is the major species, at a peak of 68129.7. The shorter peak is single conjugated protein. The relative peak heights indicate 91% double conjugate and 9% single conjugated protein respectively. +MS, 7.3-9.3 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0032] FIG. 11. A: Schematic illustrating monobromomaleimide-paclitaxel. The MS spectra of purified albumin variants (B: K573P; C: K93C+E294C+K573P) conjugated with monobromomaleimide-paclitaxel are shown. In B, a peak of 67412.2 for K573P indicates conjugation to the single free thiol at Cys34. The shorter peak is unconjugated protein. The relative peak heights indicate 77% single conjugate and 23% unconjugated protein respectively +MS, 7.1-8.9 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In C, the double conjugate of K93C+E294C+K573P is the major species which is at a peak of 68364.2. The shorter peak is triple conjugated protein. The relative peak heights indicate 60% double conjugated and 30% triple conjugate protein respectively. +MS, 7.2-9.0 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0033] FIG. 12. A: Schematic illustrating monobromomaleimide-PEG2-exenatide peptide. The MS spectra of purified albumin variants (B: K573P; C: C34A+K93C+E294C+K573P) conjugated with monobromomaleimide-PEG2-exenatide peptide are shown. In B, a peak of 71018.7 for K573P indicates conjugation to the single free thiol at Cys34. The main peak, at 66409.2 is unconjugated protein. The relative peak heights indicate single 33% conjugate and 67% unconjugated protein respectively. +MS, 7.2-8.8 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In C, the double conjugate of C34A+K93C+E294C+K573P is 75557.3. The main peak, at 70941.7 is single conjugate. The shortest peak at 66322.4 is unconjugated protein. The relative peak heights indicate 33% double conjugate, 45% single conjugate and 22% unconjugated protein respectively. +MS, 7.2-9.2 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
[0034] FIG. 13. A: Schematic illustrating maleimide-propyl-FLAG peptide. The MS spectra of purified albumin variants (B: K573P; C: K93C+E294C+K573P) conjugated with maleimide-propyl-FLAG peptide are shown. In B, a peak of 67573.4 for K573P indicates conjugation to the single free thiol at Cys34. The main peak is unconjugated protein. The relative peak heights indicate 29% single conjugate and 71% unconjugated protein respectively. +MS, 7.3-8.7 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA). In C, the triple conjugate of K93C+E294C+K573P (which includes native Cys34) is 69850.5. The main peak, at 68685.5 is double conjugate. The peak at 67520.3 is single conjugate. The shortest peak, at 66350.2 is unconjugated protein. The relative peak heights indicate 29% triple conjugate, 50% double conjugate, 20% single conjugate and 2% unconjugated protein respectively. +MS, 7.2-8.8 min, Baseline subtracted (0.50), Deconvoluted (MaxEnt), Smoothed (0.00,1,GA).
DEFINITIONS
[0035] Variant: The term "variant" means a polypeptide derived from a parent albumin by one or more (e.g. several) alteration(s), i.e. a substitution, insertion, and/or deletion, at one or more (e.g. several) positions. A substitution means a replacement of an amino acid occupying a position with a different amino acid; a deletion means removal of an amino acid occupying a position; and an insertion means adding 1 or more (e.g. several), such as 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10, preferably 1-3 amino acids immediately adjacent an amino acid occupying a position. In relation to insertion, `immediately adjacent` may be to the N-side (`upstream`) or C-side (`downstream`) of the amino acid occupying a position (`the named amino acid`). Therefore, for an amino acid named/numbered `X`, the insertion may be at position `X+1` (`downstream`) or at position `X-1` (`upstream`).
[0036] Mutant: The term "mutant" means a polynucleotide encoding a variant.
[0037] Wild-Type Albumin: The term "wild-type" (WT) albumin means albumin having the same amino acid sequence as naturally found in an animal or in a human being.
[0038] Parent Albumin: The term "parent" or "parent albumin" means an albumin to which an alteration is made by the hand of man to produce the albumin variants of the invention. The parent may be a naturally occurring (wild-type) polypeptide or an allele thereof, or even a variant thereof.
[0039] Albumin: Albumins are proteins and constitute the most abundant protein in plasma in mammals and albumins from a long number of mammals have been characterized by biochemical methods and/or by sequence information. Several albumins, e.g. HSA, have also been characterized crystallographically and the structure determined (HSA: He X M, Carter D C (July 1992), "Atomic structure and chemistry of human serum albumin", Nature 358 (6383): 209-15; horse albumin: Ho, J. X. et al. (2001). X-ray and primary structure of horse serum albumin (Equus caballus) at 0.27-nm resolution. Eur J Biochem. 215(1):205-12). The invention relates to all albumins and their structures.
[0040] The term "albumin" means a protein having the same and/or very similar three dimensional (tertiary) structure as HSA or HSA domains and having similar properties to HSA or to the relevant domains. Similar three dimensional structures are for example the structures of the albumins from the species mentioned herein. Some of the major properties of albumin are i) its ability to regulate plasma volume (oncotic activity), ii) a long plasma half-life of around 19 days .+-.5 days, iii) binding to FcRn, iv) ligand-binding, e.g. binding of endogenous molecules such as acidic, lipophilic compounds including bilirubin, fatty acids, hemin and thyroxine (see also Table 1 of Kragh-Hansen et al., 2002, Biol. Pharm. Bull. 25, 695, hereby incorporated by reference), v) binding of small organic compounds with acidic or electronegative features e.g. drugs such as warfarin, diazepam, ibuprofen and paclitaxel (see also Table 1 of Kragh-Hansen et al., 2002, Biol. Pharm. Bull. 25, 695, hereby incorporated by reference), vi) binding to gp60, also known as albondin. Not all of these properties need to be fulfilled in order to characterize a protein or fragment as an albumin. If a fragment, for example, does not comprise a domain responsible for binding of certain ligands or organic compounds the variant of such a fragment will not be expected to have these properties either.
[0041] Albumins have generally a long plasma half-life of approximately 20 days or longer, e.g. HSA has a plasma half-life of 19 days. It is known that the long plasma half-life of HSA is mediated via interaction with its receptor FcRn, however, an understanding or knowledge of the exact mechanism behind the long half-life of HSA is not essential for the invention.
[0042] As examples of albumin proteins as starting parent "backbones" for making albumin variants according to the invention can be mentioned HSA (e.g. AAA98797 or P02768-1, SEQ ID NO. 2 (mature), SEQ ID NO. 3 (immature)), primate serum albumin, (such as chimpanzee serum albumin (e.g. predicted sequence XP_517233.2 SEQ ID NO. 4), gorilla serum albumin or macaque serum albumin (e.g. NP_001182578, SEQ ID NO. 5), rodent serum albumin (such as hamster serum albumin (e.g. A6YF56, SEQ ID NO. 6), guinea pig serum albumin (e.g. Q6WDN9-1, SEQ ID NO. 7), mouse serum albumin (e.g. AAH49971 or P07724-1 Version 3, SEQ ID NO. 8) and rat serum albumin (e.g. AAH85359 or P02770-1 Version 2, SEQ ID NO. 9), bovine serum albumin (e.g. cow serum albumin P02769-1, SEQ ID NO. 10), equine serum albumin such as horse serum albumin (e.g. P35747-1, SEQ ID NO. 11) or donkey serum albumin (e.g. Q5XLE4-1, SEQ ID NO. 12), rabbit serum albumin (e.g. P49065-1 Version 2, SEQ ID NO. 13), goat serum albumin (e.g. ACF10391, SEQ ID NO. 14), sheep serum albumin (e.g. P14639-1, SEQ ID NO. 15), dog serum albumin (e.g. P49822-1, SEQ ID NO. 16), chicken serum albumin (e.g. P19121-1 Version 2, SEQ ID NO. 17) and pig serum albumin (e.g. P08835-1 Version 2, SEQ ID NO. 18) or a polypeptide having at least 70, 75, 80, 85, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 99.2, 99.4, 99.6, or at least 99.8% amino acid identity to such an albumin. Other examples of albumin, which are also included in the scope of this application, include ovalbumin (e.g. P01012.pro: chicken ovalbumin; 073860.pro: turkey ovalbumin). A mature albumin sequence can be identified from an immature albumin sequence using techniques known to the skilled person, for example alignment with HSA (for which the mature and immature regions are known). For example, immature HSA is 609 amino acids long in which amino acids 1 to 19 are a signal sequence (also known as a leader sequence or pre sequence), amino acids 20 to 24 are a pro sequence and amino acids 25 to 609 are the mature protein. The alignment in FIG. 2 allows the skilled person to predict mature sequences for several animal albumins (see "D1 Start").
[0043] HSA as disclosed in SEQ ID NO. 2, or any naturally occurring allele thereof, is the preferred parent albumin according to the invention. HSA is a protein consisting of 585 amino acid residues and has a molecular weight of 67 kDa. In its natural form it is not glycosylated. The skilled person will appreciate that natural alleles may exist having essentially the same properties as HSA but having one or more (e.g. several) amino acid changes compared to SEQ ID NO. 2, and the inventors also contemplate the use of such natural alleles as parent albumins according to the invention.
[0044] The parent albumin, a fragment thereof, or conjugation-competent albumin variant, or albumin part of a fusion polypeptide or conjugate comprising albumin or a fragment thereof according to the invention preferably has a sequence identity to the sequence of HSA shown in SEQ ID NO. 2 of at least 60%, preferably at least 70%, preferably at least 80%, preferably at least 85%, preferably at least 86%, preferably at least 87%, preferably at least 88%, preferably at least 89%, preferably at least 90%, preferably at least 91%, preferably at least 92%, preferably at least 93%, preferably at least 94%, preferably at least 95%, more preferred at least 96%, more preferred at least 97%, more preferred at least 98% and most preferred at least 99%, at least 99.2%, at least 99.4%, at least 99.6% or at least 99.8% or 100%. It is preferred that the parent albumin maintains at least one of the major properties of albumin or a similar tertiary structure as an albumin, such as HSA. The sequence identity may be over the full-length of SEQ ID NO. 2 or over a molecule consisting or comprising of a fragment such as one or more (e.g. several) domains of SEQ ID NO. 2, such as a molecule consisting of or comprising Domain III (e.g. SEQ ID NO. 19), a molecule consisting of or comprising Domain II and Domain III (e.g. SEQ ID NO. 20), a molecule consisting of or comprising Domain I and Domain III (e.g. SEQ ID NO. 21), a molecule consisting of or comprising two copies of Domain III (e.g. SEQ ID NO. 22), a molecule consisting of or comprising three copies of Domain III (e.g. SEQ ID NO. 23) or a molecule consisting of or comprising Domain I and two copies of Domain III (e.g. SEQ ID NO. 24).
[0045] The parent albumin, a fragment thereof, or conjugation-competent albumin variant, or albumin part of a fusion polypeptide or conjugate comprising albumin or a fragment thereof according to the invention, when folded, may have several, for example at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16 and suitably all 17, of the native disulphide bonds of the polypeptide of SEQ ID NO. 2.
[0046] The parent preferably comprises or consists of the amino acid sequence of SEQ ID NO. 3 (immature sequence of HSA) or SEQ ID NO. 2 (mature sequence of HSA).
[0047] In another embodiment, the parent is an allelic variant of the mature polypeptide of SEQ ID NO. 2.
[0048] The parent albumin may be encoded by a polynucleotide that hybridizes under very low stringency conditions, low stringency conditions, medium stringency conditions, medium-high stringency conditions, high stringency conditions, or very high stringency conditions with (i) the mature polypeptide coding sequence of SEQ ID NO. 2, or (ii) the full-length complementary strand of (i) (J. Sambrook, E. F. Fritsch, and T. Maniatis, 1989, Molecular Cloning, A Laboratory Manual, 2d edition, Cold Spring Harbor, N.Y.).
[0049] The polynucleotide of SEQ ID NO. 1 or a subsequence thereof, as well as the amino acid sequence of SEQ ID NO. 2 or SEQ ID NO. 3 or a fragment thereof, may be used to design nucleic acid probes to identify and clone DNA encoding a parent from strains of different genera or species according to methods well known in the art. In particular, such probes can be used for hybridization with the genomic or cDNA of the genus or species of interest, following standard Southern blotting procedures, in order to identify and isolate the corresponding gene therein. Such probes can be considerably shorter than the entire sequence, but should be at least 14, e.g. at least 25, at least 35, or at least 70 nucleotides in length. Preferably, the nucleic acid probe is at least 100 nucleotides in length, e.g. at least 200 nucleotides, at least 300 nucleotides, at least 400 nucleotides, at least 500 nucleotides, at least 600 nucleotides, at least 700 nucleotides, at least 800 nucleotides, or at least 900 nucleotides in length. Both DNA and RNA probes can be used. The probes are typically labelled for detecting the corresponding gene (for example, with .sup.32P, .sup.3H, .sup.35S, biotin, or avidin). Such probes are encompassed by the invention.
[0050] A genomic DNA or cDNA library prepared from such other organisms may be screened for DNA that hybridizes with the probes described above and encodes a parent. Genomic or other DNA from such other organisms may be separated by agarose or polyacrylamide gel electrophoresis, or other separation techniques. DNA from the libraries or the separated DNA may be transferred to and immobilized on nitrocellulose or other suitable carrier material. In order to identify a clone or DNA that is homologous with SEQ ID NO. 1 or a subsequence thereof, the carrier material is used in a Southern blot.
[0051] For purposes of the invention, hybridization indicates that the polynucleotide hybridizes to a labelled nucleotide probe corresponding to the polynucleotide shown in SEQ ID NO. 1, its complementary strand, or a subsequence thereof, under low to very high stringency conditions. Molecules to which the probe hybridizes can be detected using, for example, X-ray film or any other detection means known in the art.
[0052] The nucleic acid probe may comprise or consist of the mature polypeptide coding sequence of SEQ ID NO. 1, i.e. nucleotides 1 to 1785 of SEQ ID NO. 1. The nucleic acid probe may comprise or consist of a polynucleotide of SEQ ID NO. 25 (nucleotide sequence encoding HSA, the nucleotide sequence has been engineered to introduce restriction enzyme sites) or a fragment thereof.
[0053] For long probes of at least 100 nucleotides in length, very low to very high stringency conditions are defined as pre-hybridization and hybridization at 42.degree. C. in 5.times.SSPE, 0.3% SDS, 200 micrograms/mL sheared and denatured salmon sperm DNA, and either 25% formamide for very low and low stringencies, 35% formamide for medium and medium-high stringencies, or 50% formamide for high and very high stringencies, following standard Southern blotting procedures for 12 to 24 hours optimally. The carrier material is finally washed three times each for 15 minutes using 2.times.SSC, 0.2% SDS at 45.degree. C. (very low stringency), 50.degree. C. (low stringency), 55.degree. C. (medium stringency), 60.degree. C. (medium-high stringency), 65.degree. C. (high stringency), or 70.degree. C. (very high stringency).
[0054] For short probes that are about 15 nucleotides to about 70 nucleotides in length, stringency conditions are defined as pre-hybridization and hybridization at about 5.degree. C. to about 10.degree. C. below the calculated T.sub.m using the calculation according to Bolton and McCarthy (1962, Proc. Natl. Acad. Sci. USA 48: 1390) in 0.9 M NaCl, 0.09 M Tris-HCl pH 7.6, 6 mM EDTA, 0.5% NP-40, 1.times.Denhardt's solution, 1 mM sodium pyrophosphate, 1 mM sodium monobasic phosphate, 0.1 mM ATP, and 0.2 mg of yeast RNA per mL following standard Southern blotting procedures for 12 to 24 hours optimally. The carrier material is finally washed once in 6.times.SCC plus 0.1% SDS for 15 minutes and twice each for 15 minutes using 6.times.SSC at 5.degree. C. to 10.degree. C. below the calculated T.sub.m.
[0055] The parent or conjugation-competent albumin may be encoded by a polynucleotide with a sequence identity to the mature polypeptide coding sequence of SEQ ID NO. 1 of at least 60%, e.g. at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100%, which encodes a polypeptide which is able to function as an albumin. In an embodiment, the parent is encoded by a polynucleotide comprising or consisting of SEQ ID NO 1.
Three Dimensional (3D) Models
[0056] The present disclosure makes reference to the crystal structure of HSA from the RCSB Protein Databank (PDB, which can be viewed at http://www.rcsb.org/pdb/) with the entry with PDB identity 1AO6 or 1ao6 (Sugio, S., A. Kashima, et al. (1999), Protein Eng 12(6): 439-46). Compared to the mature HSA sequence (SEQ ID NO. 2), the 1AO6 structure starts at residue S5 (with the first 4 amino acids absent from the structure) and finishes at A582 of SEQ ID NO. 2 (with the last 3 amino acids absent from the structure). The amino acid positions used herein to describe positions to alter to generate conjugation-competent cysteines are referring to the positions in SEQ ID NO. 2, not 1ao6. Further structures of albumin are available to the skilled person, for example the atomic coordinates for the tertiary structure of human albumin are available at the GenBank DNA database which can be viewed at www.ncbi.nlm.nih.gov. Structures may be viewed using suitable software such as RasM.1 Chime (Sayle, TIBS 20, 374, 1995). Available albumin coordinates include:
[0057] 1AO6, 1BM0 (Sugio et al. (1999), Protein Eng 12(6): 439-46), which was among the top 17 requested proteins.
[0058] 1UOR, He & Carter (1992), Nature 358(6383): 209-15.
[0059] 1 bj5 and 1bke, Curry et al. (1998), Nat Struct Biol 5(9): 827-35.
[0060] 1e7a, 1e7b, 1e7c, Bhattacharya et al. (2000), J Biol Chem 275(49): 38731-8.
[0061] 1e7e, 1e7f, 1e7g, 1e7h and 1e7i, Bhattacharya et al. (2000), J Mol Biol 303(5): 721-32.
[0062] 1GNJ, Petitpas et al. (2001), J Mol Biol 314(5): 955-60.
[0063] 1HA2 and 1H9Z Petitpas et al. (2001), J Biol Chem 276(25): 22804-9.
[0064] 4K71, Schmidt et al. (2013), Structure 21:1966-1978
[0065] 4N0F and 4N0U, Oganesyan et al. (2014), J Biol Chem 289(11):7812-24.
[0066] Albumin moiety: The albumin part of a fusion polypeptide, conjugate, associate, nanoparticle or composition comprising the albumin variant or fragment thereof according to the invention, may be referred to as an `albumin moiety` or `albumin component`. A polypeptide according to the invention may comprise or consist of an albumin moiety.
[0067] Isolated variant: The term "isolated variant" means a variant in a form or environment which does not occur in nature. Non-limiting examples of isolated variants include (1) any non-naturally occurring variant; (2) any variant that is at least partially removed from one or more (e.g. several) or all of the naturally occurring constituents with which it is associated in nature; (3) any variant modified by the hand of man relative to the polypeptide from which it is derived (e.g. the polypeptide from which it is derived as found in nature); or (4) any variant modified by increasing the amount of the variant relative to other components with which it is naturally associated (e.g. multiple copies of a gene encoding the substance; use of a stronger promoter than the promoter naturally associated with the gene encoding the substance). An isolated variant may be present in a fermentation broth sample. Isolated variants may be recombinant or synthetic.
[0068] Substantially pure variant: The term "substantially pure variant" means a preparation that contains at most 10%, at most 8%, at most 6%, at most 5%, at most 4%, at most 3%, at most 2%, at most 1%, and at most 0.5% by weight of other polypeptide material with which it is natively or recombinantly associated. Preferably, the variant is at least 92% pure, e.g. at least 94% pure, at least 95% pure, at least 96% pure, at least 97% pure, at least-98% pure, at least 99%, at least 99.5% pure, and 100% pure by weight of the total polypeptide material present in the preparation. Purity may be determined by SDS-PAGE or GP-HPLC. The variants of the invention are preferably in a substantially pure form. This can be accomplished, for example, by preparing the variant by well-known recombinant methods and by purification methods.
[0069] Mature polypeptide: The term "mature polypeptide" means a polypeptide in its final form following translation and any post-translational modifications, such as N-terminal processing, C-terminal truncation, glycosylation, phosphorylation, etc. The mature polypeptide may be amino acids 1 to 585 of SEQ ID NO. 2, e.g. with the inclusion of alterations according to the invention and/or any post-translational modifications.
[0070] Mature polypeptide coding sequence: The term "mature polypeptide coding sequence" means a polynucleotide that encodes a mature albumin polypeptide. The mature polypeptide coding sequence may be nucleotides 1 to 1758 of SEQ ID NO. 1 e.g. with the alterations required to encode a variant according to the invention.
[0071] Sequence Identity: The relatedness between two amino acid sequences or between two nucleotide sequences is described by the parameter "sequence identity".
[0072] For purposes of the present invention, the sequence identity between two amino acid sequences is determined using the Needleman-Wunsch algorithm (Needleman and Wunsch, 1970, J. Mol. Biol. 48: 443-453) as implemented in the Needle program of the EMBOSS package (EMBOSS: The European Molecular Biology Open Software Suite, Rice et al., 2000, Trends Genet. 16: 276-277), preferably version 3.0.0 or later, more preferably version 5.0.0 or later. The parameters used are gap open penalty of 10, gap extension penalty of 0.5, and the EBLOSUM62 (EMBOSS version of BLOSUM62) substitution matrix. The output of Needle labelled "longest identity" (obtained using the -nobrief option) is used as the percent identity and is calculated as follows:
(Identical Residues.times.100)/(Length of Alignment-Total Number of Gaps in Alignment)
[0073] For purposes of the present invention, the sequence identity between two deoxyribonucleotide sequences is determined using the Needleman-Wunsch algorithm (Needleman and Wunsch, 1970, supra) as implemented in the Needle program of the EMBOSS package (EMBOSS: The European Molecular Biology Open Software Suite, Rice et al., 2000, supra), preferably version 3.0.0 or later, more preferably version 5.0.0 or later. The parameters used are gap open penalty of 10, gap extension penalty of 0.5, and the EDNAFULL (EMBOSS version of NCBI NUC4.4) substitution matrix. The output of Needle labelled "longest identity" (obtained using the -nobrief option) is used as the percent identity and is calculated as follows: (Identical Deoxyribonucleotides.times.100)/(Length of Alignment-Total Number of Gaps in Alignment)
[0074] Fragment: The term "fragment" as used herein includes any fragment of full-length albumin or a variant thereof, so long as at least one (e.g. several) basic property, for example binding activity (type of and specific activity e.g. binding to bilirubin), osmolarity (oncotic pressure, colloid osmotic pressure), behaviour in a certain pH-range (pH-stability) has not significantly been changed. "Significantly" in this context means that one skilled in the art would say that the properties of the variant may still be different but would not be unobvious over the ones of the original protein. A fragment may consist of one uninterrupted sequence derived from HSA or it may comprise two or more (e.g. several) sequences derived from HSA. The fragments according to the invention have a size of more than approximately 20 amino acid residues, preferably more than 30 amino acid residues, more preferred more than 40 amino acid residues, more preferred more than 50 amino acid residues, more preferred more than 75 amino acid residues, more preferred more than 100 amino acid residues, more preferred more than 200 amino acid residues, more preferred more than 300 amino acid residues, even more preferred more than 400 amino acid residues and most preferred more than 500 amino acid residues. A fragment may comprise or consist of at least 50, 60, 70, 75, 80, 85, 90, 95, 96, 97, 98, or 99% of an albumin or of a domain of an albumin. Preferred albumin domains of the invention are domains having at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.5% or 100% identity to HSA domain I consisting of amino acid residues 1 to 194.+-.1 to 15 amino acids of SEQ ID NO. 2; at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.5% or 100% identity to HSA domain II consisting of amino acid residues 192 to 387.+-.1 to 15 amino acids of SEQ ID NO. 2 and at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.5% or 100% identity to HSA domain III consisting of amino acid residues 381 to 585.+-.1 to 15 amino acids of SEQ ID NO. 2.
[0075] Domains I, II and III may be defined with reference to HSA (SEQ ID NO. 2). For example, HSA Domain I may consist of or comprise amino acids 1 to 194 (.+-.1 to 15 amino acids) of SEQ ID NO. 2, HSA Domain II may consist of or comprise amino acids 192 (.+-.1 to 15 amino acids) to 387 (.+-.1 to 15 amino acids) of SEQ ID NO. 2 and Domain III may consist of or comprise amino acid residues 381 (.+-.1 to 15 amino acids) to 585 (.+-.1 to 15 amino acids) of SEQ ID NO. 2. ".+-.1 to 15 amino acids" means that the residue number may deviate by 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, or 15 amino acids to the C-terminus and/or to the N-terminus of the stated amino acid position. Examples of domains I, II and III are described by Dockal et al. (The Journal of Biological Chemistry, 1999, Vol. 274(41): 29303-29310) and Kjeldsen et al. (Protein Expression and Purification, 1998, Vol 13: 163-169) and are tabulated below.
TABLE-US-00001 TABLE 1 Amino acid residues of HSA domains I, II and III with reference to SEQ ID NO. 2 Dockal et al Kjeldsen et al Domain I 1 to 197 1 to 192 Domain II 189 to 385 193 to 382 Domain III 381 to 585 383 to 585
[0076] A fragment may comprise or consist of one or more (e.g. several) domains of albumin described herein such as DI+DII, DI+DIII, DII+DIII, DIII+DIII, DI+DIII DIII, DIII+DIII+DIII, or fragments of such domains or combinations of domains.
[0077] The skilled person can identify domains I, II and III in non-human albumins by amino acid sequence alignment with HSA, for example using the Needleman-Wunsch algorithm (Needleman and Wunsch, 1970, J. Mol. Biol. 48: 443-453) as implemented in the Needle program of the EMBOSS package (EMBOSS: The European Molecular Biology Open Software Suite, Rice et al., 2000, Trends Genet. 16: 276-277), preferably version 3.0.0 or later, more preferably version 5.0.0 or later. The optional parameters used are gap open penalty of 10, gap extension penalty of 0.5, and the EBLOSUM62 (EMBOSS version of BLOSUM62) substitution matrix. Other suitable software includes MUSCLE ((Multiple sequence comparison by log-expectation, Robert C. Edgar, Version 3.6, http://www.drive5.com/muscle; Edgar (2004) Nucleic Acids Research 32(5), 1792-97 and Edgar (2004) BMC Bioinformatics, 5(1):113) which may be used with the default settings as described in the User Guide (Version 3.6, September 2005). Versions of MUSCLE later than 3.6 may also be used for any aspect of the invention). Examples of suitable alignments are provided in FIGS. 1 and 2.
[0078] It is preferred that domains have at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.5% identity or 100% identity to Domain I, II or III of HSA (SEQ ID NO. 2).
[0079] Additionally, single or multiple heterologous fusions comprising any of the above; or single or multiple heterologous fusions to albumin, or a variant or fragment of any of these may be used. Such fusions include albumin N-terminal fusions, albumin C-terminal fusions and co-N-terminal and C-terminal albumin fusions as exemplified by WO 01/79271 (incorporated herein by reference).
[0080] Equivalent amino acid positions: Throughout this specification amino acid positions are defined in relation to full-length mature HSA (i.e. without leader sequence, SEQ ID NO. 2). However, the skilled person understands that the invention also relates to variants of non-human albumins (e.g. those disclosed herein) and/or fragments of a human or non-human albumin. For clarity, for albumins other than HSA (SEQ ID NO. 2), equivalent residues are favoured for mutation. Equivalent positions can be identified in fragments of HSA, in animal albumins and in fragments, fusions and other derivatives or variants thereof by comparing amino acid sequences using pairwise (e.g. ClustalW) or multiple (e.g. MUSCLE) alignments. For example, FIG. 1 shows that positions equivalent to 500, 550 and 573 in full length HSA are easily identified in fragments of HSA and in albumins of other species. Positions 500, 550 and 573 are indicated by arrows. Further details are provided in Table 2 below.
TABLE-US-00002 TABLE 2 Example of identification of equivalent positions in HSA, animal albumins and albumin fragments Organism Albumin (accession Total length Position equivalent to number of Full length Fragment of mature HSA (native amino acid): protein) or fragment details protein 500 (K) 550 (D) 573 (K) Homo Full length -- 585 500 (K) 550 (D) 573 (K) sapiens (AAA98797) Homo Fragment DI, DIII 399 314 (K) 364 (D) 387 (K) sapiens Homo Fragment DI, DIII 403 318 (K) 368 (D) 391 (K) sapiens Macaca Full length -- 584 500 (K) 550 (N) 573 (P) mulatta (NP_001182578) Rattus Full length -- 584 500 (K) 550 (D) 573 (P) norvegicus (AAH85359) Mus Full length -- 584 500 (K) 550 (D) 573 (P) musculus (AAH49971)
[0081] FIG. 1 was generated by MUSCLE using the default parameters including output in ClustalW 1.81 format. The raw output data was shaded using BoxShade 3.21 (which can be accessed at http://www.ch.embnet.org/software/BOX_form.html) using Output Format: RTF_new; Font Size: 10; Consensus Line: no consensus line; Fraction of sequences (that must agree for shading): 0.5; Input sequence format: ALN. Therefore, throughout this specification amino acid positions defined in HSA also apply to equivalent positions in fragments, derivatives or variants and fusions of HSA, albumins from other species and fragments and fusions thereof. Such equivalent positions may have (i) a different residue number in its native protein and/or (ii) a different native amino acid in its native protein. Likewise, FIG. 2 shows that equivalent positions can be identified in fragments (e.g. domains) of an albumin with reference to SEQ ID NO. 2 (HSA).
[0082] Conservative substitution: As used herein, the term "conservative" amino acid substitutions refers to substitutions made within the same group, and which typically do not substantially affect protein function. By "conservative substitutions" is intended combinations such as Gly, Ala.; Val, Ile, Leu; Asp, Glu; Asn, Gln; Ser, Thr; Lys, Arg; and Phe, Tyr. Such variants may be made by techniques well known in the art, such as by site-directed mutagenesis as disclosed in U.S. Pat. No. 4,302,386 issued 24 Nov. 1981 to Stevens, incorporated herein by reference.
[0083] In one embodiment, the Venn diagram of FIG. 3 may be used to determine conservative amino acid substitutions: Using FIG. 3, a conservation mutation score (ranging from 0 to 5) may be calculated. A score of 0 is the highest conservation, which, for cysteine, is only assigned for substitution of a cysteine residue with another cysteine residue. For changes from any other amino acid to a cysteine (or for a cysteine to any other amino acid), the score may be 1, 2, 3, 4, 5. A score of 1 is a more conservative substitution than a score of 2, 3, 4 or 5. A score of 5 is assigned to the lowest conservation between a substituted amino acid and the cysteine. The score of 0 to 5 is calculated from FIG. 3 as the number of boundaries (i.e. lines) crossed to go from cysteine to the appropriate amino acid. Thus the score for cysteine is 0 as no boundaries are crossed. Likewise, the score of aspartic acid (D) is 3, since 3 boundaries are crossed. The conservation mutation score (with respect to FIG. 3) for the 20 different amino acids are defined as (using one-letter codes for the amino acids): A=1, C=0, D=3, E=4, F=4, G=2, H=5, 1=4, K=4, L=4, M=3, N=2, P=3, Q=3, R=5, S=1, T-1, V-3, W-3, Y=3.
[0084] Alternatively, or in addition, "conservative" amino acid substitutions refers to substitutions made within the same group such as within the group of basic amino acids (such as arginine, lysine, histidine), acidic amino acids (such as glutamic acid and aspartic acid), polar amino acids (such as glutamine and asparagine), hydrophobic amino acids (such as leucine, isoleucine, valine), aromatic amino acids (such as phenylalanine, tryptophan, tyrosine) and small amino acids (such as glycine, alanine, serine, threonine, methionine).
[0085] For example, a conservative substitution of alanine-2 in SEQ ID NO. 2 can include glycine or serine. Non-conservative substitutions encompass substitutions of amino acids in one group by amino acids in another group. For example, a non-conservative substitution could include the substitution of a polar amino acid for a hydrophobic amino acid.
Conventions for Designation of Variants
[0086] For purposes of the present invention, the mature polypeptide disclosed in SEQ ID NO.
[0087] 2 is used to determine the corresponding amino acid residue in another albumin. The amino acid sequence of another albumin is aligned with the mature polypeptide disclosed in SEQ ID NO. 2, and based on the alignment, the amino acid position number corresponding to any amino acid residue in the mature polypeptide disclosed in SEQ ID NO. 2 is determined using the Needleman-Wunsch algorithm (Needleman and Wunsch, 1970, J. Mol. Biol. 48: 443-453) as implemented in the Needle program of the EMBOSS package (EMBOSS: The European Molecular Biology Open Software Suite, Rice et al., 2000, Trends Genet. 16: 276-277), preferably version 3.0.0 or later, more preferably version 5.0.0 or later. The parameters used are gap open penalty of 10, gap extension penalty of 0.5, and the EBLOSUM62 (EMBOSS version of BLOSUM62) substitution matrix.
[0088] Identification of the corresponding amino acid residue in another albumin can be determined or confirmed by an alignment of multiple polypeptide sequences using several computer programs including, but not limited to, MUSCLE (multiple sequence comparison by log-expectation; version 3.5 or later; Edgar, 2004, Nucleic Acids Research 32: 1792-1797), MAFFT (version 6.857 or later; Katoh and Kuma, 2002, Nucleic Acids Research 30: 3059-3066; Katoh et al., 2005, Nucleic Acids Research 33: 511-518; Katoh and Toh, 2007, Bioinformatics 23: 372-374; Katoh et al., 2009, Methods in Molecular Biology 537: 39-64; Katoh and Toh, 2010, Bioinformatics 26: 1899-1900), and EMBOSS EMMA employing ClustalW (1.83 or later; Thompson et al., 1994, Nucleic Acids Research 22: 4673-4680), using their respective default parameters.
[0089] When the other polypeptide (or protein) has diverged from the mature polypeptide of SEQ ID NO. 2 such that traditional sequence-based comparison fails to detect their relationship (Lindahl and Elofsson, 2000, J. Mol. Biol. 295: 613-615), other pairwise sequence comparison algorithms can be used. Greater sensitivity in sequence-based searching can be attained using search programs that utilize probabilistic representations of polypeptide families (profiles) to search databases. For example, the PSI-BLAST program generates profiles through an iterative database search process and is capable of detecting remote homologs (Altschul et al., 1997, Nucleic Acids Res. 25: 3389-3402). Even greater sensitivity can be achieved if the family or superfamily for the polypeptide has one or more (e.g. several) representatives in the protein structure databases. Programs such as GenTHREADER (Jones, 1999, J. Mol. Biol. 287: 797-815; McGuffin and Jones, 2003, Bioinformatics 19: 874-881) utilize information from a variety of sources (PSI-BLAST, secondary structure prediction, structural alignment profiles, and solvation potentials) as input to a neural network that predicts the structural fold for a query sequence. Similarly, the method of Gough et al., 2000, J. Mol. Biol. 313: 903-919, can be used to align a sequence of unknown structure with the superfamily models present in the SCOP database. These alignments can in turn be used to generate homology models for the polypeptide, and such models can be assessed for accuracy using a variety of tools developed for that purpose.
[0090] For proteins of known structure, several tools and resources are available for retrieving and generating structural alignments. For example the SCOP superfamilies of proteins have been structurally aligned, and those alignments are accessible and downloadable. Two or more (e.g. several) protein structures can be aligned using a variety of algorithms such as the distance alignment matrix (Holm and Sander, 1998, Proteins 33: 88-96) or combinatorial extension (Shindyalov and Bourne, 1998, Protein Engineering 11: 739-747), and implementation of these algorithms can additionally be utilized to query structure databases with a structure of interest in order to discover possible structural homologs (e.g. Holm and Park, 2000, Bioinformatics 16: 566-567).
[0091] In describing the albumin variants of the present invention, the nomenclature described below is adapted for ease of reference. The accepted IUPAC single letter or three letter amino acid abbreviation is employed. The term `point mutation` and/or `alteration` includes deletions, insertions and substitutions.
[0092] Substitutions. For an amino acid substitution, the following nomenclature is used: Original amino acid, position, substituted amino acid. Accordingly, the substitution of threonine at position 226 with alanine is designated as "Thr326Ala" or "T326A". Multiple mutations (or alterations) are separated by addition marks ("+"), e.g. "Gly205Arg+Ser411Phe" or "G205R S411F", representing substitutions at positions 205 and 411 of glycine (G) with arginine (R) and serine (S) with phenylalanine (F), respectively. The Figures also use ("/"), e.g. "E492T/N503D" this should be viewed as interchangeable with ("+").
[0093] Deletions. For an amino acid deletion, the following nomenclature is used: Original amino acid, position*. Accordingly, the deletion of glycine at position 195 is designated as "Gly195*" or "G195*". Multiple deletions are separated by addition marks ("+"), e.g. "Gly195*+Ser411*" or "G195*+S411*".
[0094] Insertions. As disclosed above, an insertion may be to the N-side (`upstream`, `X-1`) or C-side (`downstream`, `X+1`) of the amino acid occupying a position (`the named (or original) amino acid`, `X`).
[0095] For an amino acid insertion to the C-side (`downstream`, `X+1`) of the original amino acid (`X`), the following nomenclature is used: Original amino acid, position, original amino acid, inserted amino acid. Accordingly the insertion of lysine after glycine at position 195 is designated "Gly195GlyLys" or "G195GK". An insertion of multiple amino acids is designated [Original amino acid, position, original amino acid, inserted amino acid #1, inserted amino acid #2; etc.]. For example, the insertion of lysine and alanine after glycine at position 195 is indicated as "Gly195GlyLysAla" or "G195GKA".
[0096] In such cases the inserted amino acid residue(s) are numbered by the addition of lower case letters to the position number of the amino acid residue preceding the inserted amino acid residue(s). In the above example, the sequence would thus be:
TABLE-US-00003 Parent: Variant: 195 195 195a 195b G G - K - A
[0097] For an amino acid insertion to the N-side (`upstream`, `X-1`) of the original amino acid (X), the following nomenclature is used: Original amino acid, position, inserted amino acid, original amino acid. Accordingly the insertion of lysine (K) before glycine (G) at position 195 is designated "Gly195LysGly" or "G195KG". An insertion of multiple amino acids is designated [Original amino acid, position, inserted amino acid #1, inserted amino acid #2; etc., original amino acid]. For example, the insertion of lysine (K) and alanine (A) before glycine at position 195 is indicated as "Gly195LysAlaGly" or "G195KAG". In such cases the inserted amino acid residue(s) are numbered by the addition of lower case letters with `prime` to the position number of the amino acid residue following the inserted amino acid residue(s). In the above example, the sequence would thus be:
TABLE-US-00004 Parent: Variant: 195 195a' 195b' 195 G K - A - G
[0098] Multiple alterations. Variants comprising multiple alterations are separated by addition marks ("+"), e.g. "Arg170Tyr+Gly195Glu" or "R170Y+G195E" representing a substitution of arginine and glycine at positions 170 and 195 tyrosine and glutamic acid, respectively.
[0099] Different alterations. Where different alterations can be introduced at a position, the different alterations are separated by a comma, e.g. "Arg170Tyr,Glu" represents a substitution of arginine at position 170 with tyrosine or glutamic acid. Thus, "Tyr167Gly,Ala+Arg170Gly,Ala" designates the following variants: "Tyr167Gly+Arg170Gly", "Tyr167Gly+Arg170Ala", "Tyr167Ala+Arg170Gly", and "Tyr167Ala+Arg170Ala".
[0100] Conjugation competence: A conjugation-competent cysteine is a cysteine residue which is capable of forming an intermolecular bond with a conjugation partner, particularly a conjugation partner that is not an albumin. A conjugation-competent polypeptide, i.e. thio-albumin, is capable of forming an intermolecular bond with a conjugation partner by virtue of the conjugation-competent cysteine residue. The thio-albumin may or may not have a high level of conjugation competence, for example at least 50, 60, 70, 80, 90, 95, 96, 97, 98, 99 or 100% relative to the conjugation competence of an albumin consisting of SEQ ID NO. 2 having only one conjugation competent cysteine at Cys-34. Conjugation competence may be determined relative to any conjugatable molecule (conjugation partner) of interest, for example a bioactive molecule or a fluorescent dye. Determination may be through mass spectrometry (MS) analysis or quantification of the activity of the bioactive compound such as its fluorescence. Conjugation competence of albumin and biotin or HRP may be determined by assaying the mass of the resultant conjugate and/or the enzyme activity of the conjugated compound. Determination by fluorescent labelling and cellular uptake is described by McGraw et al., (1987), The Journal of Cell Biology, 105, 207-214; and Presley et al., (1993), The Journal of Cell Biology, 122, 1231-1241. An advantage of a thio-albumin having a high conjugation competence is that it may allow efficient conjugation of molecules to the thio-albumin. Conjugation competence may be measured with respect to time. Favoured thio-albumins may be (a) those which achieve maximal conjugation quickly or (b) slowly. The conjugation competence of a specific cysteine may be determined by methods known to those skilled in the art--for example, the protein may be digested post-conjugation and peptide mapping performed to determine the degree of conjugation at the specific cysteine.
[0101] A bioactive agent or bioactive compound is one which has the ability to interact with a living organism, system or cell. It may, for example, be a biological or chemical agent or compound.
[0102] Ligand binding: The ligand binding properties of albumin include binding to anionic and neutral ligands such as long-chain fatty acids, bilirubin and other miscellaneous ligands. The long-chain fatty acids, oleic (C18:1), palmitic (C16:0), linoleic (C18:2), stearic (C18:0), arachidonic (C20:4) and palmitoleic (C16:1) are known to bind HSA. Ligand binding studies can be performed on HSA and thio-albumins using an isothermal titration calorimetry method that had been suitably qualified for this purpose. Samples can be pre-treated by defatting (Sogami, M. and J. F. Foster (1968). Biochemistry 7(6): 2172-82, incorporated herein by reference) followed by thiol blocking (Sogami, M., H. A. Petersen, et al. (1969). Biochemistry 8(1): 49-58, incorporated herein by reference) and subsequent gel permeation chromatography. The binding curves generated for thio-albumins and HSA with octanoate, for example, may subsequently be compared, and functional similarity established. Conjugated- and/or non-conjugated thio-albumin may have at least 5%, 10%, 15%, 20%, 30%, 40% or 50%, 60%, 70%, at least 80%, 90%, 95%, 100%, 105% or more of HSA's receptor binding activity, mole for mole, to biliru bin and/or a fatty acid.
[0103] FcRn and shFcRn: The term "FcRn" means the neonatal Fc receptor (FcRn), particularly the human neonatal Fc receptor. shFcRn is a soluble recombinant form of FcRn. shFcRn is a heterodimer of SEQ ID NO. 26 (truncated heavy chain of the major histocompatibility complex class I-like Fc receptor (FCGRT)) and SEQ ID NO. 27 (beta-2-microglobulin). Together, SEQ ID NO. 26 and 27 form hFcRn.
[0104] The conjugated- and/or non-conjugated thio-albumin may or may not have an altered binding affinity to FcRn.
[0105] The thio-albumin or conjugate thereof may have a binding to FcRn that is stronger or weaker (and, preferably, is stronger) than that of the parent albumin or conjugate thereof.
[0106] The thio-albumin or conjugate thereof may have a KD to FcRn (e.g. shFcRn) that is lower than the corresponding KD for HSA or conjugate thereof to. Preferably, the KD for the thio-albumin or conjugate is less than 0.9X KD for HSA to FcRn, more preferred less than 0.5X KD for HSA to FcRn, more preferred less than 0.1X KD for HSA to FcRn, even more preferred less than 0.05X KD for HSA to FcRn, even more preferred less than 0.02X KD for HSA to FcRn, even more preferred less than 0.01X KD for HSA to FcRn and most preferred less than 0.001X KD for HSA to FcRn (where X means `multiplied by`).
[0107] For a conjugate comprising a thio-albumin, preferably the KD for the conjugate is less than 0.9X KD for the corresponding conjugate comprising HSA to FcRn, more preferred less than 0.5X KD for the corresponding conjugate to FcRn, more preferred less than 0.1X KD for the corresponding conjugate to FcRn, even more preferred less than 0.05X KD for the corresponding conjugate to FcRn, even more preferred less than 0.02X KD for the corresponding conjugate to FcRn, even more preferred less than 0.01X KD for the corresponding conjugate to FcRn and most preferred less than 0.001X KD for the corresponding conjugate to FcRn (where X means `multiplied by`). `Corresponding conjugate` means a conjugate comprising HSA (e.g. SEQ ID NO. 2) instead of the thio-albumin (i.e. albumin variant).
[0108] The thio-albumin or conjugate thereof may have a KD to FcRn that is higher than the corresponding KD for HSA or conjugate thereof to FcRn. Preferably, the KD for the thio-albumin or conjugate is more than 2X KD for HSA to FcRn, more preferred more than 5X KD for HSA to FcRn, more preferred more than 10X KD for HSA to FcRn, even more preferred more than 25X KD for HSA to FcRn, most preferred more than 50X KD for HSA to FcRn. The thio-albumin or conjugate may be a null binder to FcRn.
[0109] For a conjugate comprising a thio-albumin, preferrably the KD for the conjugate, Preferably, the KD for the corresponding conjugate comprising HSA is more than 2X KD for the corresponding conjugate to FcRn, more preferred more than 5X KD for the corresponding conjugate to FcRn, more preferred more than 10X KD for the corresponding conjugate to FcRn, even more preferred more than 25X KD for the corresponding conjugate to FcRn, most preferred more than 50X KD for the corresponding conjugate to FcRn. Corresponding conjugate' means a conjugate comprising HSA (e.g. SEQ ID NO. 2) instead of the thio-albumin (i.e. albumin variant).
[0110] When determining and/or comparing KD, one or more (e.g. several) (and preferably all) of the following parameters may be used:
[0111] Instrument: Biacore 3000 instrument (GE Healthcare)
[0112] Flow cell: CM5 sensor chip
[0113] FcRn: human FcRn, preferably soluble human FcRn, optionally coupled to a tag such as Glutathione S Transferase (GST) or Histidine (His), most preferably His such as 6 histidine residues at the C-terminus of the beta-2-microglobulin.
[0114] Quantity of FcRn: 1200-2500 RU
[0115] Coupling chemistry: amine coupling chemistry (e.g. as described in the protocol provided by the manufacturer of the instrument).
[0116] Coupling method: The coupling may be performed by injecting 20 .mu.g/mL of the protein in 10 mM sodium acetate pH 5.0 (GE Healthcare). Phosphate buffer (67 mM phosphate buffer, 0.15 M NaCl, 0.005% Tween 20) at pH 5.5 may be used as running buffer and dilution buffer. Regeneration of the surfaces may be done using injections of HBS-EP buffer (0.01 M HEPES, 0.15 M NaCl, 3 mM EDTA, 0.005% surfactant P20) at pH 7.4 (Biacore AB).
[0117] Quantity of injection of test molecule (e.g. HSA or variant) 20-0.032 .mu.M
[0118] Flow rate of injection: constant, e.g. 30 .mu.L/mL
[0119] Temperature of injection: 25.degree. C.
[0120] Data evaluation software: BIAevaluation 4.1 software (BIAcore AB).
[0121] Plasma half-life: Plasma half-life is ideally determined in vivo in suitable individuals. However, since it is time consuming and expensive and inevitably there are ethical concerns connected with doing experiments in animals or man, it is desirable to use an in vitro assay for determining whether plasma half-life is extended or reduced. It is known that the binding of albumin to its receptor (FcRn) is important for plasma half-life and the correlation between receptor binding and plasma half-life is that a higher affinity of albumin to its receptor leads to longer plasma half-life. Thus for the invention a higher affinity of albumin to FcRn is considered indicative of an increased plasma half-life and a lower affinity of albumin to its receptor is considered indicative of a reduced plasma half-life.
[0122] The binding of albumin to its receptor FcRn may be described using the term affinity and the expressions "stronger" or "weaker". Thus, it should be understood that a molecule having a higher affinity to FcRn than HSA is considered to bind more strongly to FcRn than HSA and a molecule having a lower affinity to FcRn than HSA is considered to bind more weakly to FcRn than HSA. The term `binding coefficient` can be used instead of the term `binding affinity`.
[0123] The terms "longer plasma half-life" or "shorter plasma half-life" and similar expressions are understood to be in relationship to the corresponding parent or reference or corresponding albumin molecule. Thus, a longer plasma half-life with respect to a variant albumin of the invention means that the variant has longer plasma half-life than that of the corresponding albumin having the same sequences except for the alteration(s) described herein.
[0124] Reference: a reference is an albumin, fusion, conjugate, composition, associate, nanoparticle or microparticle to which an albumin variant, fusion, conjugate, composition, associate, nanoparticle or microparticle is compared. The reference may comprise or consist of full length albumin (such as HSA or a natural allele thereof) or a fragment thereof. A reference may also be referred to as a `corresponding` albumin, fusion, conjugate, composition, associate or nanoparticle to which an albumin variant, fusion, conjugate, composition, associate or nanoparticle is compared. A reference may comprise or consist of HSA (SEQ ID NO. 2) or a fragment, fusion, conjugate, associate, nanoparticle or microparticle thereof. Preferably, the reference is identical to the polypeptide, fusion polypeptide, conjugate, composition, associate, nanoparticle or microparticle according to the invention ("being studied") with the exception of the albumin moiety. Preferably the albumin moiety of the reference comprises or consists of an albumin (e.g. HSA, SEQ ID NO. 2) or a fragment thereof. The amino acid sequence of the albumin moiety of the reference may be longer than, shorter than or, preferably, the same (.+-.1 to 15 amino acids) length as the amino sequence of the albumin moiety of the polypeptide, fusion polypeptide, conjugate, composition, associate, nanoparticle or microparticle according to the invention ("being studied").
[0125] Allelic variant: The term "allelic variant" means any of two or more (several) alternative forms of a gene occupying the same chromosomal locus. Allelic variation arises naturally through mutation, and may result in polymorphism within populations. Gene mutations can be silent (no change in the encoded polypeptide) or may encode polypeptides having altered amino acid sequences. An allelic variant of a polypeptide is a polypeptide encoded by an allelic variant of a gene. Polymorphisms known for HSA (SEQ ID NO. 2) are discussed in Minchiotti et al. (2008). Hum Mutat 29(8): 1007-16 and at http://www.uniprot.org/uniprot/P02768.
[0126] Coding sequence: The term "coding sequence" means a polynucleotide, which directly specifies the amino acid sequence of its translated polypeptide product. The boundaries of the coding sequence are generally determined by an open reading frame, which usually begins with the ATG start codon or alternative start codons such as GTG and TTG and ends with a stop codon such as TAA, TAG, and TGA. The coding sequence may be a DNA, cDNA, synthetic, or recombinant polynucleotide.
[0127] cDNA: The term "cDNA" means a DNA molecule that can be prepared by reverse transcription from a mature, spliced, mRNA molecule obtained from a eukaryotic cell. cDNA lacks intron sequences that may be present in the corresponding genomic DNA. The initial, primary RNA transcript is a precursor to mRNA that is processed through a series of steps, including splicing, before appearing as mature spliced mRNA.
[0128] Nucleic acid construct: The term "nucleic acid construct" means a nucleic acid molecule, either single- or double-stranded, which is isolated from a naturally occurring gene or is modified to contain segments of nucleic acids in a manner that would not otherwise exist in nature or which is synthetic. The term nucleic acid construct is synonymous with the term "expression cassette" when the nucleic acid construct contains the control sequences required for expression of a coding sequence of the invention.
[0129] Control sequences: The term "control sequences" means all nucleic acid sequences necessary for the expression of a polynucleotide encoding a variant of the invention. Each control sequence may be native (i.e. from the same gene) or foreign (i.e. from a different gene) to the polynucleotide encoding the variant or native or foreign to each other. Such control sequences include, but are not limited to, a leader, polyadenylation sequence, propeptide sequence, promoter, signal peptide sequence, and transcription terminator. At a minimum, the control sequences include a promoter, and transcriptional and translational stop signals. The control sequences may be provided with linkers for the purpose of introducing specific restriction sites facilitating ligation of the control sequences with the coding region of the polynucleotide encoding a variant.
[0130] Operably linked: The term "operably linked" means a configuration in which a control sequence is placed at an appropriate position relative to the coding sequence of a polynucleotide such that the control sequence directs the expression of the coding sequence.
[0131] Expression: The term "expression" includes any step involved in the production of the variant including, but not limited to, transcription, post-transcriptional modification, translation, post-translational modification, and secretion. The thio-albumin may or may not be capable of being expressed at a level of at least 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100% relative to the expression of an unmodified albumin (such as SEQ ID NO. 2) from a suitable expression system, such as yeast (e.g. Saccharomyces, e.g. S. cerevisiae) or an Aspergillus. Relative expression levels can be determined, for example, by expression of the protein followed by quantification by SDS-PAGE, GP-HPLC or Western Blotting.
[0132] Expression vector: The term "expression vector" means a linear or circular DNA molecule that comprises a polynucleotide encoding a variant and is operably linked to control sequences that provide for its expression.
[0133] Host cell: The term "host cell" means any cell type that is susceptible to transformation, transfection, transduction, or the like with a nucleic acid construct or expression vector comprising a polynucleotide of the present invention. The term "host cell" encompasses any progeny of a parent cell that is not identical to the parent cell due to mutations that occur during replication.
DETAILED DESCRIPTION OF THE INVENTION
Conjugation-Competent Polypeptides I
[0134] A first aspect of the invention provides a conjugation-competent polypeptide comprising an amino acid sequence which is at least 60% identical to human albumin, particularly residues 1 to 585 of the mature human albumin polypeptide sequence of SEQ ID NO. 2, or a fragment thereof;
[0135] wherein at least one (e.g. several) position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398 of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue;
[0136] preferably wherein the conjugation-competent polypeptide has a tendency to exist as a monomer in solution which is at least 70% of the tendency of the parent polypeptide (such as the polypeptide of SEQ ID NO. 2) to exist as a monomer in solution, more preferably at least 75, 80, 85, 90, 95, 96, 97, 98, at least 99 or 100% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution. Preferably the parent polypeptide does not contain the conjugation-competent Cys residue or residues described herein. Preferably the parent polypeptide does not contain the additional mutation or mutations described herein. That is, preferably the parent polypeptide is identical to the conjugation-competent polypeptide with the exception of the introduced cysteine residue or residues and, if present, the introduced other mutation or mutations.
[0137] Suitably, the at least one (e.g. several) position is selected from K93, E294, A226, E230, I271, and E358, particularly from K93, E294, A226, E230, and I271.
[0138] Preferably the conjugation-competent polypeptide has at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.2, 99.4, 99.6, 99.8% sequence identity to SEQ ID NO. 2. For example, in addition to the introduced Cys residue or Cys residues, the conjugation-competent polypeptide may have at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 (e.g. several) other mutations relative to SEQ ID NO. 2. Alternatively, in addition to the introduced Cys residue or Cys residues, the conjugation-competent polypeptide may have zero other mutations relative to SEQ ID NO. 2.
[0139] Preferably, the conjugation-competent polypeptide has a tendency to exist as a monomer in solution which is at least 75% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution and at least one position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, L284, K317, A322, E333, D340, E354, K359, A362, E382, and L398 comprises a conjugation-competent cysteine residue.
[0140] Preferably the polypeptide is a recombinant polypeptide. Preferably the polypeptide is an isolated and/or purified polypeptide. Preferably the polypeptide is synthetic and/or does not naturally occur in nature.
[0141] A conjugation-competent cysteine at the position defined above may or may not be created in an albumin by insertion, for example by adding a cysteine with or without one or more (e.g. several) additional residues and without removal of an amino acid residue from the albumin sequence; or by substituting one or more (e.g. several) adjacent amino acids with a larger number of residues containing at least one (e.g. several) cysteine, thus extending the overall length of the polypeptide. For example, a cysteine residue may be introduced immediately adjacent an albumin residue identified herein. The cysteine residue may be introduced as a single cysteine residue or within a polypeptide. The polypeptide may be from 2 to 50 amino acids long, preferably from 2, 10, 20, 30, or 40 to 10, 20, 30, 40 or 50 amino acids long.
[0142] Suitably, the polypeptide comprises one or more (e.g. several) of: [0143] a) substitution of an amino acid, other than cysteine, with a cysteine at a position corresponding to a position equivalent to any of residues K93, E294, A226, E230, I271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2; and/or [0144] b) insertion of a cysteine at a position adjacent the N- or C-side of an amino acid corresponding to a position equivalent to any of residues K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2.
[0145] Substitutions are preferred, and the following disclosure of selected positions should be understood to specifically encompass substitutions, without limitation.
[0146] Suitably 2, 3, 4, 5 or more (e.g. several) positions equivalent to positions selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2 comprise a conjugation-competent cysteine residue. Suitably the 2, 3, 4, 5 or more (e.g. several) positions are selected from K93, E294, A226, E230, 1271, and E358, particularly from K93, E294, A226, E230 and 1271.
[0147] For a polypeptide comprising a Cys at a position equivalent to position E294 of SEQ ID NO. 1, preferably the polypeptide also comprises a Cys at a position equivalent to one or more of K93, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, or E382.
[0148] The inventors have found that variants of HSA in which cysteine has been substituted at a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, and E382 have the beneficial property of a tendency to exist as a monomer in solution which is at least 70% of the tendency of the HSA polypeptide of SEQ ID NO. 2 to exist as a monomer in solution. A cysteine introduced at one of the selected positions therefore has a low tendency to cause the variant to form dimers or higher order oligomers in solution. This beneficial effect is also noted in variants in which there are cysteines at more than one selected position. Without wishing to be bound by theory, the inventors ascribe the monomer tendencies of the polypeptides of the invention to the flexibility of the polypeptide chain in the region of, and surface exposure at, the site of cysteine substitution. This reflects an exercise of inventive skill, based on years of experience in protein structural biology, in the choices applied by the inventors in selecting positions within HSA for substitution with cysteine.
[0149] The tendency of albumin or variants thereof to exist as a monomer, rather than a dimer or higher order oligomer, can be determined based on measurement of monomer, dimer and higher order oligomer quantities in solutions of the albumin or variant under similar conditions.
[0150] Suitable techniques for performing such measurements include Gel Permeation High Pressure Liquid Chromatography, as described in the Examples. Results are typically expressed as "percentage monomer", which is calculated as:
amount of monomeric albumin by mass.times.100/(amount of monomeric albumin by mass+amount of dimeric albumin by mass+amount of higher order oligomer by mass).
[0151] Alternatively, the tendency to form non-monomers in solution, that is dimers and/or higher order oligomers, may be expressed. The "percentage non-monomer" is 100% minus percentage monomer.
[0152] Samples may be tested shortly after purification (for example, within 24 hours after purification) following production in shake flasks or 10 L bioreactors, or following storage at 2-8.degree. C., e.g. 5.degree. C., for time periods of up to or including 1 week, 1 month, 2 months, 3 months or 6 months. Samples are typically tested, and optionally stored, in a solution of one or more (e.g. several) salts and at a pH of about 7.0.+-.0.5. The solution may comprise a buffer comprising 50 mM ammonium acetate, 10 mM sodium octanoate, pH 7.0, preferably at a polypeptide concentration of from about 0.2 to about 2.5 mg/mL. The solution may comprise a buffer comprising 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5, preferably at a polypeptide concentration of from about 5 to about 50 mg/mL.
[0153] The percentage monomer for a given albumin may differ depending on the albumin purity and concentration. Albumin produced in shake flask culture is typically purified using a single AlbuPure.RTM. (Prometic Life Sciences Inc. or Albumedix Ltd (formerly Novozymes Biopharma UK Ltd)) chromatography step, and typically is obtained at a concentration of about 0.2 to 2.0 mg/mL, more preferably 1.+-.0.5 mg/mL and a protein purity of >95% by SDS reducing PAGE. AlbuPure.RTM. is a high-performance affinity capture adsorbent designed for albumin fusion protein purification, which comprises a synthetic triazine ligand coupled to a base matrix. Under these conditions, percentage monomer of HSA was found to be about 87%, rising to about 89% upon storage at 6 months at 2-8.degree. C. e.g. 5.degree. C. Albumin produced in 10 L bioreactor culture is typically purified by a AlbuPure.RTM. chromatography step followed by an ion exchange chromatography, is ultrafiltered, and then formulated at 50 mg/mL, and has a protein purity of >99% by SDS reducing PAGE. Under these conditions, percentage monomer of HSA was found to be about 94%, and was stable at two months of storage at 2-8.degree. C. and at 6 months storage at 2-8.degree. C. A variant having at least 70% of the tendency of HSA to exist as a monomer in solution may therefore be found to be at least 60% monomer, preferably at least 69% monomer (less than 40% non-monomer, preferably less than 31% non-monomer) when tested after typical shake flask production and purification as described above, for samples tested shortly after purification or stored for up to two or up to six months. For a variant having at least 80% of the tendency of HSA to exist as a monomer in solution, the percentage monomer should be at least 70% preferably at least 79% monomer, and the percentage non-monomer less than 30%, preferably less than 21%. A variant having at least 70% of the tendency of HSA to exist as a monomer in solution may be found to be at least 65% monomer, preferably at least 69% monomer, when tested after typical 10 L bioreactor production and purification as described above, for samples tested shortly after purification or stored for up to two months. For a variant having at least 80% of the tendency of HSA to exist as a monomer in solution, the percentage monomer should be at least 75% preferably at least 79%. The tendency is preferably measured at day 0, e.g. the day that the variant is produced, however it may also be measured later e.g. at day 1, 2, 3, 4, 5, 6, 7 or after 2, 3, 4, 5, 6, 7 weeks or after 1 or 2 months storage e.g. at 2-8.degree. C. e.g. 5.degree. C. Suitably, the percentage monomer should be stable upon storage for up to seven weeks or two months, meaning that it does not reduce by more than 10, more than 9, 8, 7, 6, 5, 4, 3, 2 or 1 percentage points between testing shortly after purification and testing after two months of storage e.g. at 2-8.degree. C. e.g. 5.degree. C. Preferably the percentage monomer should not reduce by more than 5 percentage points between testing shortly after purification and testing after 7 weeks of storage at 2-8.degree. C. e.g. 5.degree. C.
[0154] The variant may or may not have a tendency to exist as a monomer in solution which is at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or at least 100% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution. This tendency may be tested shortly after purification or after storage for up to six months e.g. at 2-8.degree. C. e.g. 5.degree. C.
[0155] The tendency of the polypeptide to exist as monomer in solution may be measured following storage for at least 7 weeks at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 8 weeks at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 3 months at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 4 months at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 6 months storage at a temperature from 2 to 8.degree. C. such as 5.degree. C., or at least 3 months storage at a tempature of about 40.degree. C. Most preferably the tendency of the polypeptide to exist as monomer in solution is measured following storage for at least 3 months at a temperature from 2 to 8.degree. C. such as 5.degree. C.
[0156] The tendency of the polypeptide to exist as a monomer in solution may be measured at a polypeptide concentration of from 0.2 to 50 mg/mL, for example at about 5 mg/mL.
[0157] The tendency of the polypeptide to exist as a monomer in solution may be measured at a pH from about 6.0, 6.1, 6.2, 6.3, 6.4, 6.5, 6.6, 6.7, 6.8, 6.9, 7.0, 7.1, 7.2, 7.3, or 7.4 to about 6.1, 6.2, 6.3, 6.4, 6.6, 6.7, 6.8, 6.9, 7.0, 7.1, 7.2, 7.3, 7.4 or 7.5, preferably about pH 7.
[0158] The tendency of the polypeptide to exist as a monomer in solution may be measured in a buffer comprising 50 mM ammonium acetate, 10 mM sodium octanoate, pH 7.0, preferably at a polypeptide concentration of from about 0.5 to about 5 mg/mL.
[0159] The tendency of the polypeptide to exist as a monomer in solution may be measured in a buffer comprising 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5, preferably at a polypeptide concentration of from about 5 to about 50 mg/mL.
[0160] The conjugation-competent polypeptide may, prior to storage, be purified for example using a triazine (such as AlbuPure.RTM.) chromatography matrix or DE-FF chromatography matrix, more preferably by triazine (such as AlbuPure.RTM.) chromatography matrix followed by DE-FF chromatography matrix. Suitable methods are disclosed in Example 10
[0161] The polypeptide sample storage may be static. The polypeptide sample storage may be vertical.
[0162] Where a variant comprises more than one conjugation-competent cysteine as provided above, the tendency to exist as a monomer may be reduced compared to the variant which differs only by virtue of having one fewer such cysteines. For example, a variant albumin having the substitutions E294C+K93C has a lower tendency to exist as a monomer than a variant albumin having either substitution alone. Suitably, the variant comprises a conjugation-competent cysteine residue at two positions selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and I271, of SEQ ID NO. 2, wherein the variant has a tendency to exist as a monomer in solution which is at least 75% of the tendency of a variant which differs only by virtue of comprising a conjugation-competent cysteine residue at only one of the two positions.
[0163] Suitably, the variant comprises a conjugation-competent cysteine residue at two positions selected from K93, E294, A226, E230, 1271, E358, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2, wherein the variant has a tendency to exist as a monomer in solution which is at least 75% of the tendency of a variant which differs only by virtue of comprising a conjugation-competent cysteine residue at only one of the two positions. Higher monomer tendencies are preferred, such as at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or at least 100%. For example, HSA comprising the substitution E294C+K93C has a tendency to exist as a monomer in solution which is at least 90% of the tendency of HSA comprising the substitution K93C, or at least 85% of the tendency of HSA comprising the substitution E294C, to exist as a monomer in solution. These results are illustrated in the Examples, with material purified from 10 L bioreactor preparations, and tested shortly after purification, or after storage for seven weeks or two months at 2-8.degree. C. e.g. 5.degree. C. The same samples were also stable following storage for 6 months. Albumin variants having more than one conjugation-competent cysteine can be prepared by introducing a further conjugation-competent cysteine residue into a variant which already has at least one (e.g. several) conjugation-competent cysteine residue. Variants comprising a further conjugation-competent cysteine residue which have at least 75% of the tendency of the reference albumin lacking the further conjugation-competent cysteine residue to exist as a monomer in solution may be preferred.
[0164] Suitable variants may comprise a conjugation-competent cysteine residue at one or two or more (e.g. several) positions selected from K93, E294, A226, E230, 1271 and E358 of SEQ ID NO. 2. Suitable combinations of positions are (i) K93+E294, A226, E230, 1271, or E358; (ii) E294+K93, A226, E230, 1271, or E358; (iii) A226+K93, E294, E230, 1271, or E358; (iv) E230+K93, E294, A226, 1271, or E358; (v) 1271+K93, E294, A226, E230, or E358; (vi) K93+E294+A226, E230, 1271, or E358 of SEQ ID NO. 2. Suitable variants may comprise a conjugation-competent cysteine residue at one or two or more (e.g. several) positions selected from L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, D237, E230, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, L398 of SEQ ID NO. 2. Suitable combinations of positions are: (1) L24+F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (2) F49+L24, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (3) V54+L24, F49, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (4) D56+L24, F49, V54, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (5) L66+L24, F49, V54, D56, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (6) A92+L24, F49, V54, D56, L66, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (7) Q94+L24, F49, V54, D56, L66, A92, K93, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (8) E97+L24, F49, V54, D56, L66, A92, K93, Q94, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (9) H128+L24, F49, V54, D56, L66, A92, K93, Q94, E97, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (10) F156+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (11) E227+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (12) D237+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E230, E227, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (13) K240+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E230, E227, D237, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (14) D259+L24, F49, V54, D56, L66, A92, K93, Q94; E97, H128, F156, A226, E230, E227, D237, K240, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (15) K262+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E230, E227, D237, K240, D259, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (16) N267+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E230, E227, D237, K240, D259, K262, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (17) 0268+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (18) L275+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (19) E277+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (20) L284+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, 0237, K240, D259, K262, N267, Q268, 1271, L275, E277, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (21) E311+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, K317, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (22) K317+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, A322, E333, D340, E354, E358, K359, A362, E382, or L398; (23) A322+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, E333, D340, E354, E358, K359, A362, E382, or L398; (24) E333+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, D340, E354, E358, K359, A362, E382, or L398; (25) D340+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, E227, D237, E230, K240, D259, K262, N267, Q268, I271, L275, E277, L284, E294, E311, K317, A322, E333, E354, E358, K359, A362, E382, or L398; (26) E354+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E358, K35,9, A362, E382, or L398; (27) K359+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, A362, E382, or L398; (28) A362+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, E382, or L398; (29) E382+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, or L398; (30) L398+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, or E382; (31) K93+L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382 or L398; (32) E294+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382 or L398; (33) A226+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382 or L398; (34) E230+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382 or L398; (35) 1271+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, E358, K359, A362, E382 or L398; and (36) E358+L24, F49, V54, D56, L66, A92, K93, Q94, E97, H128, F156, A226, E227, E230, D237, K240, D259, K262, N267, Q268, 1271, L275, E277, L284, E294, E311, K317, A322, E333, D340, E354, K359, A362, E382 or L398.
[0165] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine provided at a position equivalent to K93 in SEQ ID NO. 2.
[0166] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E294 in SEQ ID NO. 2.
[0167] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to A226 in SEQ ID NO. 2.
[0168] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E230 in SEQ ID NO. 2.
[0169] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to 1271 in SEQ ID NO. 2.
[0170] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E358 in SEQ ID NO. 2.
[0171] A particularly preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K93 in SEQ ID NO. 2 and a cysteine at a position equivalent to E294 in SEQ ID NO. 2.
[0172] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to L24 in SEQ ID NO. 2.
[0173] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to F49 in SEQ ID NO. 2.
[0174] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to V54 in SEQ ID NO. 2.
[0175] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to D56 in SEQ ID NO. 2.
[0176] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to L66 in SEQ ID NO. 2.
[0177] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to A92 in SEQ ID NO. 2.
[0178] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to Q94 in SEQ ID NO. 2.
[0179] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E97 in SEQ ID NO. 2.
[0180] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to H128 in SEQ ID NO. 2.
[0181] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to F156 in SEQ ID NO. 2.
[0182] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E227 in SEQ ID NO. 2.
[0183] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to D237 in SEQ ID NO. 2.
[0184] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to K240 in SEQ ID NO. 2.
[0185] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to D259 in SEQ ID NO. 2.
[0186] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to K262 in SEQ ID NO. 2.
[0187] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to N267 in SEQ ID NO. 2.
[0188] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to Q268 in SEQ ID NO. 2.
[0189] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to L275 in SEQ ID NO. 2.
[0190] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E277 in SEQ ID NO. 2.
[0191] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to L284 in SEQ ID NO. 2.
[0192] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E311 in SEQ ID NO. 2.
[0193] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to K317 in SEQ ID NO. 2.
[0194] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to A322 in SEQ ID NO. 2.
[0195] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E333 in SEQ ID NO. 2.
[0196] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to D340 in SEQ ID NO. 2.
[0197] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E354 in SEQ ID NO. 2.
[0198] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to K359 in SEQ ID NO. 2.
[0199] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to A362 in SEQ ID NO. 2.
[0200] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to E382 in SEQ ID NO. 2.
[0201] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 and a cysteine at a position equivalent to L398 in SEQ ID NO. 2.
[0202] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine provided at a position equivalent to K93 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0203] A particularly preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 a cysteine at a position equivalent to E294 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0204] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A226 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0205] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E230 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0206] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to 1271 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0207] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E358 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0208] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K93 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2, a cysteine at a position equivalent to E294 in SEQ ID NO. 2.
[0209] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L24 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0210] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to F49 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0211] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to V54 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0212] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D56 in SEQ ID NO. 2 and a cysteine at a position equivalent to 034 in SEQ ID NO. 2.
[0213] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L66 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0214] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A92 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0215] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to Q94 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0216] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E97 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0217] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to H128 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0218] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to F156 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0219] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E227 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0220] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D237 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0221] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K240 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0222] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D259 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0223] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K262 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0224] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to N267 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0225] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to Q268 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0226] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L275 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0227] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E277 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0228] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L284 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0229] A preferred polypeptide may have at least 90% identity to SEQ ID NO: 2, a cysteine at a position equivalent to E311 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0230] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K317 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0231] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A322 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0232] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E333 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0233] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D340 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0234] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E354 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0235] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K359 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0236] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A362 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0237] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E382 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0238] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L398 in SEQ ID NO. 2 and a cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0239] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine provided at a position equivalent to K93 and no cysteine at a position equivalent to 034 in SEQ ID NO. 2.
[0240] A particularly preferred polypeptide may have at least 90% identity to SEQ ID NO. 2 a cysteine at a position equivalent to E294 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0241] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A226 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0242] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E230 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0243] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to 1271 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0244] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E358 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0245] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K93 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2, a cysteine at a position equivalent to E294 in SEQ ID NO. 2.
[0246] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent, to L24 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0247] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to F49 SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0248] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to V54 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0249] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D56 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0250] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L66 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0251] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A92 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0252] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to Q94 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0253] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E97 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0254] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to H128 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0255] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to F156 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0256] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E227 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0257] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D237 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0258] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K240 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0259] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D259 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0260] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K262 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0261] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to N267 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0262] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to Q268 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0263] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L275 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0264] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E277 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0265] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L284 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0266] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E311 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0267] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K317 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0268] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A322 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0269] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E333 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0270] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to D340 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0271] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E354 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0272] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to K359 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0273] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to A362 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0274] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to E382 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0275] A preferred polypeptide may have at least 90% identity to SEQ ID NO. 2, a cysteine at a position equivalent to L398 in SEQ ID NO. 2 and no cysteine at a position equivalent to C34 in SEQ ID NO. 2.
[0276] The `no cysteine` at a position equivalent to C34 in SEQ ID NO. 2 may be provided, for example, by a substitution of C34 to an amino acid, such as a natural amino acid, for example, A, D, E, F, G, H, I, K, L, M, N, P, Q, R, S, T, V, W, or Y. Such a substitution may be described as C34X. The substitution C34A is preferred. The `no cysteine` at a position equivalent to C34 in SEQ ID NO. 2 may be provided, for example, by deletion of the cysteine at this position.
[0277] A thio-albumin may or may not include a polypeptide where one or more (e.g. several) naturally occurring free-thiol group(s), such as cysteine-34 in HSA (SEQ ID NO. 2), is modified to an amino acid which is not cysteine. For example, cysteine may or may not be replaced by an amino acid which has a relatively high conservation score (e.g. 1, 2 or 3 as calculated according to FIG. 3) such as alanine or serine. A thio-albumin may or may not include a polypeptide where one or more (e.g. several) naturally occurring free-thiol group(s), such as cysteine-34 in HSA (SEQ ID NO. 2) are present. Thus, the conjugation-competent polypeptide of any of the above embodiments may comprise, at a position equivalent to position 34 of SEQ ID NO. 2, a conjugation-competent cysteine. Alternatively, there may not be a conjugation-competent cysteine at a position equivalent to position 34 of SEQ ID NO. 2.
[0278] For a polypeptide comprising two or more (several) conjugation competent cysteine residues, when the polypeptide is folded, the conjugation competent cysteine residues may or may not be relatively evenly distributed over the surface of the folded protein. The term `folded` includes folding of a polypeptide/protein into its natural configuration, for example the most thermodynamically stable folded configuration. An advantage of relatively even distribution is that it allows conjugation of two or more (several) moieties to the thio-albumin with minimal steric hindrance or without steric hindrance between two or more (several) of the conjugated moieties. This has the advantage of minimising, and optionally eliminating, potential loss of activity due to issues such as steric hindrance between adjacent moieties (conjugation partners) which may be conjugated to the thio-albumin. Such moieties, for example bioactive molecules, may be relatively bulky.
[0279] Preferably the two or more (several) conjugation-competent cysteines are distributed over the surface of the thio-albumin molecule such that they are spaced as far from each other as possible, for example geometrically possible. Preferably the distance between two or more (several) conjugation-competent cysteines is at least 5, 10, 20, 30, 40, 50, 60, 70, or 80 Angstroms. Preferably each conjugation competent cysteine is at least 5, 10, 20, 30, 40, 50, 60, 70, or 80 Angstroms distant from one or several or all other conjugation-competent cysteines in the molecule. The distance between two conjugation-competent cysteines is preferably a distance which is at least 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, or 95% and most preferably 100% of the length of the longest axis of the folded albumin molecule, for example as shown in a model of an albumin. For example, the longest axis of SEQ ID NO. 2 as shown in protein structure 1A06 is approximately 85 Angstroms. Therefore, it is preferred that the two or more (several) of the cysteine residues are at least 65, 70, 75 or most preferably 80 Angstroms apart. Most preferably each conjugation-competent cysteine residue is at a distance of at least 80, 90, or 95% and most preferably 100% of the length of the longest axis of the folded albumin molecule.
[0280] Preferably the side chains of conjugation-competent cysteines are directed away from each other and/or directed so that a moiety conjugated to the cysteine will be directed away from the centre of the albumin structure. This provides the advantage of preventing interactions between the conjugated moieties and/or the albumin moiety itself.
[0281] With reference to an amino acid sequence, candidate amino acid residues may be visually inspected using software such as Yasara (Krieger and Vriend, 2014, Bioinformatics 30(20) 2981-2982; and described at http://www.yasara.org/).
[0282] Suitably, the polypeptide comprises substitution of an amino acid, other than cysteine, with a cysteine at one or both positions corresponding to a position equivalent to residues K93 or E294 of SEQ ID NO. 2. The C.alpha.-C.alpha. distance between C34 and K93 is 20.3 .ANG., between C34 and E294 is 39.9 .ANG. and K93 and E294 45.9 .ANG. in WT HSA (SEQ ID NO. 2).
[0283] Maleimide conjugation is a convenient means of conjugating a conjugation partner to an albumin. Capability to form a conjugate with maleimide-polyethylenglycol2-biotin is believed to be indicative of capability to form a conjugate with other conjugation partners containing a maleimide group. Conversely, if a conjugation-competent polypeptide has a low efficiency of conjugation with maleimide-polyethylenglycol2-biotin, or fails to conjugate, this is not indicative that it is poorly capable or not capable of conjugating with a different chemical group. Maleimide conjugates form a thio-ether bond, which may or may not be capable of stabilisation upon controlled hydrolysis. Stable conjugate formation may be preferred, such that the conjugate does not release a reactive maleimide conjugation partner during storage or use. The latter could potentially form unwanted conjugates with thiol-reactive species encountered in vivo.
[0284] As shown in the Examples, native HSA having a single free thiol at cysteine 34 forms approximately 50% stable conjugate upon maleimide conjugation and controlled hydrolysis. In contrast, polypeptides of the invention may form stable conjugates at higher efficiencies. In particular, albumins comprising a free thiol group at a position selected from those equivalent to K93, E294, and E358 of SEQ ID NO. 2 form stable maleimide conjugates at high efficiency, as shown in the Examples. Albumins comprising two or more (several) such thiols also may also form stable maleimide conjugates.
[0285] A conjugation-competent polypeptide of the invention may or may not be capable of forming a conjugate with maleimide-polyethylenglycol2-biotin (maleimide-PEG2-biotin) at a conjugation efficiency of at least 90%, preferably at least 95%, which conjugate may or may not be at least 90%, preferably at least 95% stable upon controlled hydrolysis. FIG. 4 illustrates the conjugation of maleimide-PEG2-biotin to a free thiol of a protein, and reactions which may occur to the formed conjugate.
[0286] A conjugation efficiency of a particular percentage indicates that the specified percentage of free thiol groups in the albumin form an adduct with the maleimide moiety, under suitable reaction conditions. The maleimide group reacts with thiols in the pH range 6.5-7.5 to form a thio-ether linkage with very little cross-reactivity with amines at this pH. The use of 20 mM sodium phosphate, 150 mM sodium chloride, pH 7.2 works well for this reaction. The concentration of protein should ideally be in the range of 1-10 mg/mL. Lower concentrations of protein may result in the need to increase the molar excess of reagent to obtain an acceptable level of modification (Hermanson, Greg T. (2008), Bioconjugate Techniques. Second Edition, Academic Press, San Diego, Calif.). The formation of the adduct results in an increase in mass which can be measured, for example by mass spectrometry, as in the Examples. Conveniently, the percentage conjugation efficiency is in relation to all free thiols of the albumin. Where the albumin has more than one such free thiol, a different percentage conjugation efficiency may pertain to each free thiol, and may be expressed in relation either to each individual free thiol, or collectively to all free thiols. Thus, if an albumin has two free thiols, one having 50% conjugation efficiency and the other having 100% conjugation efficiency, the overall conjugation efficiency for the albumin is the average of the two conjugation efficiencies, in this case 75%.
[0287] A stability of a particular percentage upon controlled hydrolysis indicates that the specified percentage of thiol-maleimide adduct undergoes ring-opening stabilisation, that is, the succinimide ring moiety is hydrolysed to a succinic acid moiety, and the thio-ether bond of the conjugate is maintained, as illustrated in FIG. 4. The percentage stability may be expressed in relation either to each individual free thiol or the albumin, or collectively to all free thiols. Controlled hydrolysis may be performed at alkaline pH and above ambient temperature. Suitably, adducts are incubated at pH 9.0 and 37.degree. C. for at least 18 hours, preferably 24 hours in a buffered salts solution, such as phosphate buffered saline. The hydrolysis of the succinimide moiety to a succinic acid moiety by the addition of H.sub.2O has the effect of increasing the mass of the conjugate, which can be measured, for example by mass spectrometry, as in the Examples. Where conjugation efficiency is incomplete, this must be taken into account in determining the percentage stability. For example, if 50% of an albumin having one free thiol forms a conjugate, and 40% of the albumin is conjugated following controlled hydrolysis, this represents a stability of 80%. In these circumstances, 50% of the albumin is initially unconjugated, and therefore has a mass indicative of free albumin. The mass does not change upon controlled hydrolysis. Of the 50% of the albumin that is initially conjugated, a portion, 40% of the total albumin, has an increased mass of 18 Da due to the addition of H.sub.2O. The other portion, 10% of the total albumin, does not undergo hydrolysis and therefore its mass does not change. Although this albumin is still conjugated, it may be unstable during storage or use, because it can undergo de-conjugation via the retro-Michael pathway, as illustrated in FIG. 4. In contrast, the stably hydrolysed conjugate can be expected to remain stable during storage or use (Fontaine, S. et al, Bioconjugate Chem. 2015, 26, 145-152).
[0288] Suitably conjugation efficiencies for a polypeptide of the invention may be at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or substantially 100%. Suitably conjugation efficiencies for an individual free thiol of a polypeptide of the invention may be at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95% at least 96%, at least 97%, at least 98%, at least 99%, or substantially 100%. Suitable stabilities of a polypeptide conjugate upon controlled hydrolysis may be at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or substantially 100%.
[0289] As shown in the Examples, native HSA having a single free thiol at cysteine 34 forms greater than about 90% conjugate. Albumins comprising a free thiol group at a position selected from those equivalent to K93, E294, E358, L24, V54, H128, E227, K240, K262, Q268, E277, K317, A322, K359, and A362 of SEQ ID NO. 2 form maleimide conjugates greater than about 90% efficiency, those with a free thiol group at a position selected from those equivalent to L24, V54, H128, E227, K240, K262, K359, and A362 form maleimide conjugates greater than about 95% efficiency.
[0290] Suitable stabilities of a particular thiol-ether conjugate bond of a polypeptide conjugate upon controlled hydrolysis may be at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or substantially 100%.
[0291] The polypeptide may or may not further comprise a further linker to which a conjugation partner, such as a bioactive compound, radiopharmaceutical or imaging agent, may be linked. For example a linker may comprise a primary amine such as a lysine.
[0292] It is preferred that the conjugation-competent polypeptide has an acceptable immunogenicity, particularly in humans. More preferably the conjugation-competent polypeptide has an immunogenicity that is comparable to or lower than that of a parent albumin such as WT HSA (SEQ ID NO. 2). Therefore, preferably the alteration(s) to provide a conjugation competent cysteine residue(s) do not adversely affect the immunogenicity of the polypeptide relative to the parent albumin such as WT HSA.
[0293] Preferably, the alteration(s) made to provide the conjugation competent cysteine residue(s) do not adversely affect the immunogenicity of the polypeptide in human, e.g. relative to the immunogenicity of wild-type HSA (SEQ ID NO. 2).
[0294] The immunogenicity of the polypeptide may be determined or predicted by screening for T-cell epitopes and/or for B-cell epitopes. Screening may be in silico, in vitro or ex vivo. For example, the immunogenicity of the polypeptide may be determined or predicted by an ex vivo T cell activation assay. The T cell activation assay may comprise measuring T cell responses using a proliferation assay, e.g. [3H]-thymidine uptake. Preferably, the polypeptide has less than 10% reactivity in the T cell proliferation assay, preferably less than 8, 6, 4, or 2% reactivity, most preferably 0%. `Reactivity` means that a positive response was observed. Therefore 10% reactivity means that a positive response was observed in 10% of the donor samples.
[0295] The T cell activation assay may comprise measuring T cell responses using a cytokine secretion assay, e.g. IL-2 ELISpot. Preferably the polypeptide has less than 10% reactivity in the cytokine secretion assay, preferably less than 8, 6, 4, or 2% reactivity, most preferably 0%. `Reactivity` means that a positive response was observed. Therefore 10% reactivity means that a positive response was observed in 10% of the donor samples.
[0296] More preferred, the conjugation-competent polypeptide has less than 10% reactivity in a T cell proliferation assay and in a cytokine secretion assay, e.g. an EpiScreen.TM. assay (Abzena, Cambridge, UK).
[0297] The T cell assays may comprise CD4+ T cells.
[0298] The T cell assays may use peripheral blood mononuclear cells from a cohort of 50 healthy donors representing the European and North American population (based on HLA allotypes).
[0299] Preferably, the polypeptide does not stimulate an adverse antibody response in human, such as a specific antibody response.
[0300] For a conjugate comprising the conjugation-competent polypeptide, preferably the conjugate has an immunogenicity that is comparable to or lower than that of a corresponding conjugate comprising a parent albumin such as WT HSA (SEQ ID NO. 2) instead of the conjugation-competent polypeptide. Consequently, the properties mentioned for the conjugation-competent polypeptide also apply to a conjugate comprising the conjugation-competent polypeptide, however the `control` may be a parent albumin such as WT HSA or a corresponding conjugate comprising a parent albumin such as WT HSA.
Conjugation-Competent Polypeptides II
[0301] A second aspect of the invention provides a conjugation-competent polypeptide comprising an amino acid sequence according to the first aspect of the invention, and at least one (e.g. several) further modification compared to SEQ ID NO. 2, such as a further modification which causes the polypeptide to have at least one (e.g. several) further conjugation-competent cysteine, or alters the binding affinity of the polypeptide for FcRn, or alters the plasma half-life of the polypeptide.
[0302] The second aspect of the invention allows for the favoured conjugation-competent cysteines as defined in relation to the first aspect of the invention to be combined with other modifications in an albumin background, and provides the option to further tailor the albumin for specific applications.
Further Conjugation-Competent Cysteines
[0303] The at least one (e.g. several) further modification may or may not cause the polypeptide to have at least one (e.g. several) further conjugation-competent cysteine. The polypeptide may or may not comprise a total of 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 conjugation competent cysteine residues. The polypeptide may or may not comprise at least one (e.g. several) further conjugation-competent cysteine as defined in relation to the first aspect of the invention.
[0304] The polypeptide may or may not comprise at least one (e.g. several) further conjugation-competent cysteine, other than at a position corresponding to least one position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2. Suitable conjugation-competent cysteines are disclosed in WO 2010/092135 (incorporated by reference, particularly FIGS. 5 and 6). Suitably, at least one (e.g. several) position equivalent to a position selected from D1, A2, H3, S5, A55, S58, C75, T76, T79, E82, T83, E86, C91, D121, V122, C124, T125, D129, C169, C177, A229, T236, E266, D269, S270, 5273, S304, K313, D314, C316, N318, A320, C361, A364, C369, A371, N386, Q390, Q397, S435, T478, T496, A504, E505, T506, T508, D549, C558, D562, C567, A581, L585 and A578 of SEQ ID NO. 2 may comprise a conjugation-competent cysteine. Suitably, the polypeptide may comprise one or more (e.g. several) of: (a) substitution of an amino acid, other than cysteine, with a cysteine at a position corresponding to a position equivalent to any of residues D1, A2, H3, S5, A55, S58, C75, T76, T79, E82, T83, E86, C91, D121, V122, C124, T125, D129, C169, C177, A229, T236, E266, D269, 8270, S273, S304, K313, D314, C316, N318, A320, C361, A364, 0369, A371, N386, Q390, Q397, S435, T478, T496, A504, E505, T506, T508, D549, C558, D562, C567, A581, L585 and A578 of SEQ ID NO. 2; (b) insertion of a cysteine at a position adjacent the N- or C-side of an amino acid corresponding to a position equivalent to any of residues D1, A2, H3, S5, A55, S58, C75, T76, T79, E82, T83, E86, C91, D121, V122, C124, T125, D129, C169, C177, A229, T236, E266, D269, S270, S273, S304, K313, D314, 0316, N318, A320, C361, A364, C369, A371, N386, Q390, Q397, S435, T478, T496, A504, E505, T506, T508, D549, C558, D562, C567, A581, L585 and A578 of SEQ ID NO. 2 so as to generate a conjugation competent cysteine at any of C369, C361, C91, C177, C567, C316, C75, C169, C124 and C558; and (c) addition of a cysteine to the N-side of the N-terminal residue of an albumin sequence or to the C-side of the C-terminal residue of an albumin sequence. Exemplary combinations include conjugation-competent cysteines located at: (a) A2+L585, (b) A2+A364+D562+L585C, (c) A2 and adjacent the C-side of the C-terminus of the albumin (d) T79+A364; (e) A364+D1; (f) T79+D562+A364; (g) D562+A364+D1; (h) T79+D562+A364+A504; (i) T79+D562+A364+L585; (j) T79+D562+A364+D1; (k) T79+D562+A364+L585+D1; (I) E86+D562+A364+A504+A2; (m) S270+A581; (n) 5270+D129; (o) S270+A581+E82; (p) S270+A581+D129; (q) S270+A581+E82+D129; (r) 5270+A581+E82+D129+Q397; (s) C369+C177; (t) A364+A581; (u) T79+A364+A581; (v) A364+A581+D129; (w) A364+0177; (x) D562+C369; (y) D129+C369; (z) A581+C369; or (aa) D562+D129+C369.
[0305] Further suitable cysteine residues may be introduced as disclosed in WO 2009/126920 or WO 2010/059315 (incorporated herein by reference). Specifically, one or more (e.g. several) surface-exposed amino acid residues may be substituted for a cysteine residue, corresponding to one or more (e.g. several) positions corresponding S58, T76, T79, T83, T125, T236, S270, S273, 5304, S435, T478, T496, T506 and T508 of SEQ ID NO. 2.
[0306] As noted in relation to the first aspect of the invention, increasing the number of conjugation-competent cysteine residues in an albumin variant may reduce its tendency to exist as a monomer in solution. It is preferred that the conjugation-competent polypeptide of the second aspect of the invention has a tendency to exist as a monomer in solution which is at least 70% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution, and optionally at least 75%, at least 80%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or at least 100%. This preference applies whether or not the polypeptide comprises a further conjugation-competent cysteine as defined in relation to the second aspect. Nevertheless, useful conjugation-competent polypeptides may still be provided which have a lower tendency to exist as a monomer in solution. Because the conjugation-competent cysteine residues defined in relation to the first aspect of the invention themselves contribute relatively minimally to non-monomer formation, combining one or more (e.g. several) of them with one or more (e.g. several) other conjugation-competent cysteine residues can be expected to result in a variant having increased monomer percentage compared to a variant having the same number of conjugation-competent cysteine residues selected from the prior art.
Albumin Variants with Altered Binding to FcRn and/or Altered Plasma Half-Life
[0307] The at least one (e.g. several) further modification may or may not alter the binding affinity of the albumin variant to FcRn and/or alter the plasma half-life. Preferably the albumin variant may have at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.2, 99.4, 99.6, 99.8% sequence identity to SEQ ID NO. 2. For example, in addition to the introduced Cys residue or Cys residues, the albumin variant may have at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 (e.g. several) other mutations relative to SEQ ID NO. 2. Alternatively, in addition to the introduced Cys residue or Cys residues, the albumin variant may have zero other mutations relative to SEQ ID NO. 2.
[0308] The thio-albumin or conjugate may have a plasma half-life that is either longer or shorter, preferably longer, than that of the parent albumin or conjugate thereof, or a binding to FcRn that is stronger or weaker, preferably stronger. Preferably the thio-albumin or conjugate has a plasma half-life that is longer than that of HSA or the corresponding conjugate thereof.
[0309] Alternatively, this may be expressed as the thio-albumin or conjugate having a KD to FcRn (e.g. shFcRn) that is lower than the corresponding KD for HSA or conjugate thereof to. Preferably, the KD for the thio-albumin or conjugate is less than 0.9X KD for HSA to FcRn, more preferred less than 0.5X KD for HSA to FcRn, more preferred less than 0.1X KD for HSA to FcRn, even more preferred less than 0.05X KD for HSA to FcRn, even more preferred less than 0.02X KD for HSA to FcRn, even more preferred less than 0.01X KD for HSA to FcRn and most preferred less than 0.001X KD for HSA to FcRn (where X means `multiplied by`).
[0310] For a conjugate comprising a thio-albumin, preferrably the KD for the conjugate is less than 0.9X KD for the corresponding conjugate comprising HSA to FcRn, more preferred less than 0.5X KD for the corresponding conjugate to FcRn, more preferred less than 0.1X KD for the corresponding conjugate to FcRn, even more preferred less than 0.05X KD for the corresponding conjugate to FcRn, even more preferred less than 0.02X KD for the corresponding conjugate to FcRn, even more preferred less than 0.01X KD for the corresponding conjugate to FcRn and most preferred less than 0.001X KD for the corresponding conjugate to FcRn (where X means `multiplied by`). `Corresponding conjugate` means a conjugate comprising HSA (e.g. SEQ ID NO. 2) instead of the thio-albumin albumin variant).
[0311] Alternatively, the thio-albumin or conjugate may have a plasma half-life that is shorter than that of HSA or the conjugate thereof.
[0312] This may be expressed as the thio-albumin or conjugate having a KD to FcRn that is higher than the corresponding KD for HSA or conjugate thereof to FcRn. Preferably, the KD for the thio-albumin or conjugate is more than 2X KD for HSA to FcRn, more preferred more than 5X KD for HSA to FcRn, more preferred more than 10X KD for HSA to FcRn, even more preferred more than 25X KD for HSA to FcRn, most preferred more than 50X KD for HSA to FcRn. The thio-albumin or conjugate may be a null binder to FcRn.
[0313] For a conjugate comprising a thio-albumin, preferrably the KD for the conjugate, Preferably, the KD for the corresponding conjugate comprising HSA is more than 2X KD for the corresponding conjugate to FcRn, more preferred more than 5X KD for the corresponding conjugate to FcRn, more preferred more than 10X KD for the corresponding conjugate to FcRn, even more preferred more than 25X KD for the corresponding conjugate to FcRn, most preferred more than 50X KD for the corresponding conjugate to FcRn. Corresponding conjugate' means a conjugate comprising HSA (e.g. SEQ ID NO. 2) instead of the thio-albumin (i.e. albumin variant).
[0314] The half-life of the thio-albumin or conjugate or product made from associate, nanoparticle, microparticle or liposome may be tailored in order to achieve a binding affinity or half-life which meets the needs of the user.
[0315] When determining and/or comparing KD, one or more (e.g. several) (and preferably all) of the following parameters may be used:
[0316] Instrument: Biacore 3000 instrument (GE Healthcare)
[0317] Flow cell: CM5 sensor chip
[0318] FcRn: human FcRn, preferably soluble human FcRn, optionally coupled to a tag such as
[0319] Glutathione S Transferase (GST) or Histidine (His), most preferably His such as 6 histidine residues at the C-terminus of the beta-2-microglobulin.
[0320] Quantity of FcRn: 1200-2500 RU
[0321] Coupling chemistry: amine coupling chemistry (e.g. as described in the protocol provided by the manufacturer of the instrument).
[0322] Coupling method: The coupling may be performed by injecting 20 .mu.g/mL of the protein in 10 mM sodium acetate pH 5.0 (GE Healthcare). Phosphate buffer (67 mM phosphate buffer, 0.15 M NaCl, 0.005% Tween 20) at pH 5.5 may be used as running buffer and dilution buffer. Regeneration of the surfaces may be done using injections of HBS-EP buffer (0.01 M HEPES, 0.15 M NaCl, 3 mM EDTA, 0.005% surfactant P20) at pH 7.4 (Biacore AB).
[0323] Quantity of injection of test molecule (e.g. HSA or variant) 20-0.032 .mu.M
[0324] Flow rate of injection: constant, e.g. 30 .mu.L/mL
[0325] Temperature of injection: 25.degree. C.
[0326] Data evaluation software: BIAevaluation 4.1 software (BIAcore AB).
[0327] Domain III of albumin is primarily responsible for binding FcRn. The conjugation-competent polypeptide may or may not comprise or consist of albumin domain III or a variant thereof and at least one (e.g. several) additional albumin domain or fragment thereof, such as a second albumin domain III or a variant thereof, as disclosed in WO 2011/124718 (incorporated herein by reference). Suitably, the polypeptide comprises or consists of at least one (e.g. several) albumin domain III or variant or fragment thereof, wherein at least one (e.g. several) albumin domain III comprises one or more (e.g. several) substitutions in positions corresponding to the positions in SEQ ID NO. 2 selected among: 573, 500, 550, 417, 440, 464, 490, 492, 493, 494, 495, 496, 499, 501, 503, 504, 505, 506, 510, 535, 536, 537, 538, 540, 541, 542, 574, 575, 577, 578, 579, 580, 581, 582 and 584, as disclosed in WO 2011/051489 (incorporated herein by reference). Suitable substitutions include one or more (e.g. several) substitutions in positions corresponding to the positions in SEQ ID NO. 2 selected among: K573Y, W, P, H, F, V, I, T, N, S, G, M, C, A, E, Q, R, L, D, K500E, G, D, A, 5, C, P, H, F, N, W, T, M, Y, V, Q, L, I, R, Q417A, H440A, H464Q, E492G, D494N,Q,A, E495Q,A, T496A, D494E+Q417H, D494N+T496A, E492G+V493P, P499A, E501A,Q, N503H,K, H510Q, H535Q, K536A, P537A, K538A, K541G,D, D550E,N, E492G+K573P,A, or E492G/N503H/K573P.
[0328] In an alternative embodiment, the polypeptide may comprise alterations at two or more (several) positions selected from positions corresponding to positions (a) 492 and 580; (b) 492 and 574; (c) 492 and 550; (d) 550 and 573; (e) 550 and 574; (f) 550 and 580 in SEQ ID NO. 2, as disclosed in WO 2014/072481 (incorporated herein by reference).
[0329] In an alternative embodiment, the conjugation-competent polypeptide may comprise: (i) an N-terminal region comprising a first albumin which is a human albumin variant, in which the N-terminal of the first albumin comprises all amino acids of the human albumin variant except the C-terminal 2 to 30 amino acids; and (ii) a C-terminal region of a second albumin, which is selected from macaque albumin, mouse albumin, rabbit albumin, sheep albumin, human albumin, goat albumin, chimpanzee albumin, hamster albumin, guinea pig albumin, rat albumin, cow albumin, horse albumin, donkey albumin, dog albumin, chicken albumin, or pig albumin, or a variant thereof, in which the C-terminal of the second albumin or albumin variant comprises the C-terminal 2 to 30 amino acids of the second albumin or albumin variant; wherein the polypeptide has (i) an altered plasma half-life compared with the human albumin variant and/or (ii) an altered binding affinity to FcRn compared with the human albumin variant, as disclosed in WO 2012/059486 (incorporated herein by reference).
[0330] In an alternative embodiment, the polypeptide may comprise one or more (e.g. several) alterations in Domain I of the mature human albumin polypeptide sequence of SEQ ID NO. 2; and one or more (e.g. several) alterations in Domain III of the mature human albumin polypeptide sequence of SEQ ID NO. 2, wherein the one or more (e.g. several) alterations cause the polypeptide to have an altered binding affinity to FcRn, as disclosed in WO 2013/135896 (incorporated herein by reference). Suitably, the alteration(s) in Domain I are selected from positions corresponding to any of positions 78 to 120 of SEQ ID NO. 2, such as any of positions 78 to 88 and/or from any of 105 to 120; and the alteration(s) in Domain III are selected from positions corresponding to any of positions 425, 505, 510, 512, 524, 527, 531, 534, 569, 573, 575 of SEQ ID NO. 2. Suitably, the alteration at the position corresponding to positions 78 to 120 or 425, 505, 510, 512, 524, 527, 531, 534, 569, 573, and/or 575 of SEQ ID NO. 2 is a substitution; and the alteration is optionally a substitution selected from (i) 83N, K or S; (ii) 111D, G, H, R, Q or E; or (iii) 573P, Y, W, H, F, T, I or V.
[0331] In an alternative embodiment, the polypeptide may comprise one or more (e.g. several) alterations in Domain II of the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions 349, 342, 381, 345, 384, 198, 206, 340, 341, 343, 344, 352, 382, 348, and/or 383 in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn, as disclosed in WO 2015/036579 (incorporated herein by reference). Suitably, the alteration at the position corresponding to position 349, 342, 381, 345, 384, 198, 206, 340, 341, 343, 344, 352, 382, 348, and/or 383 is a substitution; and the alteration is optionally a substitution selected from (i) 349F, W, Y, H, P, K or Q, preferably F; (ii) 342Y, W, F, H, T, N, Q, A, C, I, L, P, V, preferably Y; (iii) 381G or A, preferably G; or (iv) 345E, H, I or Q.
[0332] In an alternative embodiment, the polypeptide may comprise a variant Domain III of an albumin, or fragment thereof, comprising a mutation, such as a substitution, corresponding to one or more (e.g. several) positions corresponding to V418, T420, V424, E505 and V547 of SEQ ID NO. 2. These mutations are disclosed in WO 2013/075066 (incorporated herein by reference). Substitutions may be at one, two or more (several, e.g. at two, three, four, or five) of the positions corresponding to V418, T420, V424, E505 and V547; for example, there may be one or more (e.g. several) substitutions selected from V418M, T420A, V4241, E505(R/K/G) and V547A. In a particular embodiment, the albumin comprises the substitutions V418M, T420A and E505R; or V418M, T420A, E505G and V547A. The albumin may comprise one or more (e.g. several) additional substitutions at positions selected from N429, M446, A449, T467, and A552; such as selected from N429D, M446V, A449V, T467M, and A552T.
[0333] In an alternative embodiment, the variant may comprise a variant Domain III of an albumin, or fragment thereof, comprising one to eighteen amino acid substitutions to increase one or both of affinity for FcRn and serum half-life of the polypeptide, as disclosed in WO 2011/103076 (incorporated herein by reference). Substitutions may be at any one or more (e.g. several) of positions corresponding to positions 381, 383, 391, 401, 402, 407, 411, 413, 414, 415, 416, 424, 426, 434, 442, 445, 447, 450, 454, 455, 456, 457, 459, 463, 495, 506, 508, 509, 511, 512, 515, 516, 517, 519, 521, 523, 524, 525, 526, 527, 531, 535, 538, 539, 541, 557, 561, 566 or 569 of SEQ ID NO. 2. Suitable substitutions may be selected from V381N, V381Q, E383A, E383G, E3831, E383L, E383V, N391A, N391G, N391I, N391L, N391V, Y401D, Y401E, K402A, K402G, K4021, K402L, K402V, L407F, L407N, L407Q, L407W, L407Y, Y411Q, Y411N, K413C, K413S, K413T, K414S, K414T, V415C, V415S, V415T, Q416H, Q416P, V424A, V424G, V424I, V424L, V424N, V424Q, V426D, V426E, V426H, V426P, G434C, G4345, G434T, E442K, E442R, R445F, R445W, R445Y, P447S, P447T, E450D, E450E, S454C, S454M, 5454T, V455N, V455Q, V456N, V456Q, L457F, L457W, L457Y, Q459K, Q459R, L463N, L463Q, E495D, 1506F, T506W, 1506Y, T508K, T508R, T508S, F509C, F5091, F509L, F509M, F509V, F509W, F509Y, A511F, A511W, A511Y, D512F, D512W, D512Y, T515C, T515H, T515N, T515P, T515Q, T515S, L516F, L516S, L516T, L516W, L516Y, S5170, 5517F, S517M, 5517T, S517W, S517Y, K519A, K519G, K5191, K519L, K519V, R521F, R521W, R521Y, I523A, I523D, 1523E, I523F, I523G, I523K, I523L, I523N, 1523Q, I523R, I523V, 1523W, I523Y, K524A, K524G, K5241, K524L, K524V, K525A, K525G, K5251, K525L, K525V, Q526C, Q526M, Q526S, Q526T, Q526Y, T527F, T527W, T527Y, E531A, E531G, E5311, E531L, E531V, H535D, H535E, H535P, K538F, K538W, K538Y, A539I, A539L, A539V, K541F, K541W, K541Y, K557A, K557G, K557I, K557L, K557V, A561F, A561W, A561Y, T566F, T566W, T566Y, A569H, and A569P; such as selected from L407N, L407Y, V415T, V4241, V424Q, V426E, V426H, P447S, V455N, V456N, L463N, E495D, T506Y, 1508R, F509M, F509W, A511F, D512Y, T515Q, L516T, L516W, S517W, R521W, I523D, 1523E, I523G, 1523K, I523R, K524L, Q526M, T527Y, H535P and K557G.
[0334] The variant may comprise a variant Domain III of an albumin, or fragment thereof, comprising amino acid substitutions at positions corresponding to the following positions of SEQ ID NO. 2: (a) residues 383 and 413; (b) residues 401 and 523; (c) residues 407 and 447; (d) residues 407 and 447 and 539; (e) residues 407 and 509; (f) residues 407 and 526; (g) residues 411 and 535; (h) residues 414 and 456; (i) residues 415 and 569; (j) residues 426 and 526; (k) residues 442 and 450 and 459; (I) residues 463 and 508; (m) residues 508 and 519 and 525; (n) residues 509 and 527; (o) residues 523 and 538; (p) residues 526 and 557; (q) residues 541 and 561; (r) residues 463 and 523; (s) residues 508 and 523; (t) residues 508 and 524; (u) residues 463, 508 and 523; (v) residues 463, 508 and 524; (w) residue 508, 523 and 524; (x) residue 463, 508, 523 and 524; (y) residues 463 and 524; (z) residues 523 and 524; and (aa) residues 463, 523, and 524, wherein the substitutions increase one or both of affinity for FcRn and serum half-life of the polypeptide, as disclosed in WO 2012/112188 (incorporated herein by reference). Suitable substitutions may be selected from (a) L463C, F, G, H, I, N, S or Q; (b) T5080, E, I, K, R or S; (c) I523A, C, D, E, F, G, H, I, K, L, M, N, P, Q, R, S, T, V, W or Y; (d) K524A, F, G, H, I, L, M, Q, T or V; (e) L463F or N; (f) T508R or S; (g) I523D, E, F, G, K or R; and (h) K524L.
[0335] The variant albumin may comprise one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions V418, T420, V424, E505, V547, K573 in SEQ ID NO. 2; wherein the one or more (several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn.
[0336] The variant albumin may comprise one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions V381, preferably V381N or Q; E383, preferably E383A, G, I, L, or V; N391, preferably N391A, G, I, L or V; Y401 preferably Y401D or E; K402, preferably K402A, G, I, L, or V; L407, preferably L407F, N, Q, W, or Y; Y411, preferably Y411Q, or N; K413, preferably K413C, S, or T; K414, preferably K414S or T; V415C, preferably V415C, S, or T; Q416, preferably Q416H or P; V424, preferably V424A, G, I, L, N, or Q; V4260, preferably V426D, E, H, or P; G434, preferably G434C, S, or T; E442, preferably E442K or R; R445, preferably R445F, W or Y; P447, preferably P447S or T; E450, preferably E450D or E; S454, preferably S454C, M or T; V455, preferably V455N or Q; V456, preferably V456N or Q; L457, preferably L457F, W or Y; Q459, preferably Q459K or R; L463, preferably L463N or Q; E495, preferably E495D; T506, preferably T506F, W or Y; T508, preferably T508K, R, or S; F509, preferably F5090, I, L, M, V, W or Y; A511, preferably A511F, W, or Y; D512, preferably D512F, W or Y; T515, preferably T515C, H, N, P, Q or S; L516, preferably L516F, S, T, W or Y; S517, preferably S517C, F, M, T, W or Y; K519, preferably K519A, G, I, L, or V; R521, preferably R521F, W or Y; 1523, preferably I523A, D, E, F, G, K, L, N, Q, R, V, W or Y; K524, preferably K524A, G, I, L or V; K525, preferably K525A, G, I, L or V; Q526, preferably Q526C, M, S, T or Y; T527, preferably T527F, W or Y; E531, preferably E531A, G, I, L or V; H535, preferably H535D, E or P; K538, preferably K538F, W or Y; A539, preferably A5391, L or V; K541, preferably, K541F, W or Y; K557, preferably K557A, G, I, L or V; A561, preferably A561F, W or Y; T566, preferably T566F, W or Y; A569, preferably A569H or P in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn.
[0337] The variant albumin may comprise one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions V547, preferably V457A; K573, preferably K573P or Y; I523, preferably I523A or G, T527, preferably T527M, K500, preferably K500A; or E505, preferably E505Q in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn.
[0338] The variant albumin may comprise one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions 573, 523, 527 or 505 of SEQ ID NO. 2, preferably K573Y; I523G; I523A; T527M; E5050; or K573P, for example K573Y and 1523G; K573Y, I523G and T527M; K573Y, E5050 and T527M; K573Y and T527M; K573P and I523G; K573P, 1523G and T527M; K573P, E505Q and T527M; K573P and T527M; V547A; V547A and K573P; V547A, E5050, K573P and T527M; or K500A and H510Q of SEQ ID NO. 2.
Other Modifications
[0339] The second aspect of the invention encompasses other modifications. For example, the polypeptide may or may not comprise at least one (e.g. several) mutation that reduces glycosylation.
Fusion Polypeptide
[0340] A third aspect of the invention provides a fusion polypeptide comprising a conjugation-competent polypeptide of either the first or the second aspect of the invention.
[0341] Polypeptides of the invention may be fused with a non-albumin polypeptide fusion partner. The fusion partner may in principle be any polypeptide but generally it is preferred that the fusion partner is a polypeptide having bioactive, therapeutic, prophylactic (including vaccine), diagnostic, imaging or other beneficial properties. Such properties may be referred to as `pharmaceutically beneficial properties`. Fusion polypeptides comprising albumin or fragments thereof are known in the art. It has been found that such fusion polypeptides comprising albumin or a fragment thereof and a fusion partner polypeptide have a longer plasma half-life compared to the unfused fusion partner polypeptide alone.
[0342] One or more (e.g. several) bioactive, therapeutic, prophylactic (including vaccine), diagnostic, imaging or other beneficial polypeptides may be fused to the N-terminus, the C-terminus of albumin, inserted into a loop in the albumin structure or any combination thereof. It may or it may not comprise linker sequences separating the various components of the fusion polypeptide. By way of non-limiting examples, a fusion may comprise N'-partner-albumin-C', N'-albumin-partner-C', N'-albumin-partner-albumin-C', N'-partner-albumin-partner-C' where `partner` is the fusion partner.
[0343] Teachings relating to fusions of albumin or a fragment thereof are known in the art and the skilled person will appreciate that such teachings can also be applied to the invention. WO 2001/79271A (particularly page 9 and/or Table 1), WO 2003/59934 (particularly Table 1), WO 03/060071 (particularly Table 1) and WO 01/079480 (particularly Table 1) (each incorporated herein by reference in their entirety) also contain examples of bioactive, therapeutic, prophylactic (including vaccine), diagnostic, imaging or other beneficial polypeptides that may be fused to albumin or fragments thereof, and these examples apply also to the invention.
[0344] An advantage of using a genetically or chemically fused albumin is that either or all of the molecules which contribute to the fusion may have improved properties relative to the unfused molecule(s) (Balan et al. (2006), Antivir Ther 11(1): 35-45). Albumins and albumin particles are also important for carrying and delivering drugs and prodrugs to their sites of action (Kratz, F. (2008), Journal of Controlled Release, 132 (3), p. 171-183). Fusion and particle technologies offer improved dosing regimens due to improved pharmacokinetic properties, such as half-life extension, and may improve bioavailability and protect the fused conjugation partner, for example bioactive molecule, radiopharmaceutical or imaging agent, from inactivation.
[0345] The polypeptide may also be fused to one or more (e.g. several) purification tags such as (Ala-Trp-Trp-Pro).sub.n, avidin/streptavidin/Strep-tag, FLAG.TM. peptide (DYKDDDDK), His-tag.
[0346] Further preferences for the third aspect of the invention include those of the first and second aspects of the invention. The skilled person understands that any aspect of the invention may be combined with another aspect or aspects of the invention and/or with one or more (e.g. several) of the preferences for the aspects of the invention and/or other disclosures made herein.
Polynucleotides
[0347] A. fourth aspect of the invention provides a polynucleotide which encodes the polypeptide according to the first, second or third aspects of the invention.
[0348] The polynucleotide may be an isolated polynucleotide. The polynucleotide may be comprised in a vector (such as a plasmid) and/or in a host cell.
[0349] The polynucleotide may or may not be codon-optimised relative to the host from which it is to be expressed. SEQ ID NO. 1 provides the usual coding sequence of HSA (SEQ ID NO. 2). SEQ ID NO. 28 provides a coding sequence of HSA (SEQ ID NO. 1) which is codon-optimised for expression from S. cerevisiae. SEQ ID NO. 1 or SEQ ID NO. 28 may be mutated in order to provide a polynucleotide which encodes a polypeptide according to the invention. Preferably the polynucleotide is synthetic and/or recombinant. Preferably the polynucleotide is an isolated polynucleotide. The polynucleotide may encode an HSA with or without a leader sequence. For example, the polynucleotide may encode an HSA with the natural leader sequence of HSA (amino acids 1 to 24 of SEQ ID NO. 3) or an HSA with a fusion leader sequence (amino acids 1 to 24 of SEQ ID NO. 29).
[0350] The polypeptide may be provided as a nucleic acid construct comprising a polynucleotide encoding a polypeptide of the invention operably linked to one or more (e.g. several) control sequences that direct the expression of the coding sequence in a suitable host cell under conditions compatible with the control sequences.
[0351] A polynucleotide may be manipulated in a variety of ways to provide for expression of a variant. Manipulation of the polynucleotide prior to its insertion into a vector may be desirable or necessary depending on the expression vector. The techniques for modifying polynucleotides utilizing recombinant DNA methods are well known in the art.
[0352] The control sequence may be a promoter sequence, which is recognized by a host cell for expression of the polynucleotide. The promoter sequence contains transcriptional control sequences that mediate the expression of the variant. The promoter may be any nucleic acid sequence that shows transcriptional activity in the host cell including mutant, truncated, and hybrid promoters, and may be obtained from genes encoding extracellular or intracellular polypeptides either homologous or heterologous to the host cell.
[0353] In a yeast host, useful promoters are obtained from the genes for Saccharomyces cerevisiae enolase (ENO-1), Saccharomyces cerevisiae protease A (PRA1), Saccharomyces cerevisiae protease B (PRB1), Saccharomyces cerevisiae translation elongation factor (TEF1), Saccharomyces cerevisiae translation elongation factor (TEF2), Saccharomyces cerevisiae galactokinase (GAL1), Saccharomyces cerevisiae alcohol dehydrogenase/glyceraldehyde-3-phosphate dehydrogenase (ADH1, ADH2/GAP), Saccharomyces cerevisiae triose phosphate isomerase (TPI), Saccharomyces cerevisiae metallothionein (CUP1), and Saccharomyces cerevisiae 3-phosphoglycerate kinase. Other useful promoters for yeast host cells are described by Romanos et al., 1992, Yeast 8: 423-488.
[0354] The skilled person knows useful promoters for use in rice and mammalian cells, such as CHO or HEK. In a rice host, useful promoters are obtained from cauliflower mosaic virus 35S RNA gene (CaMV35S), maize alcohol dehydrogenase (Adh1) and alpha Amy3.
[0355] In a mammalian host cell, such as CHO or HEK, useful promoters are obtained from Cytomegalovirus (CMV) and CAG hybrid promoter (hybrid of CMV early enhancer element and chicken beta-actin promoter), Simian vacuolating virus 40 (SV40).
[0356] The control sequence may also be a suitable transcription terminator sequence, which is recognized by a host cell to terminate transcription. The terminator sequence is operably linked to the 3'-terminus of the polynucleotide encoding the variant. Any terminator that is functional in the host cell may be used.
[0357] Preferred terminators for yeast host cells are obtained from the genes for Saccharomyces cerevisiae enolase, Saccharomyces cerevisiae cytochrome C (CYC1), Saccharomyces cerevisiae alcohol dehydrogenase (ADH1) and Saccharomyces cerevisiae glyceraldehyde-3-phosphate dehydrogenase. Other useful terminators for yeast host cells are described by Romanos et al., 1992, supra. The skilled person knows useful terminators for use in rice and mammalian cells, such as CHO or HEK. For example, in a rice host, preferred terminators are obtained from Agrobacterium tumefaciens nopaline synthase (Nos) and cauliflower mosaic virus 35S RNA gene (CaMV35S).
[0358] The control sequence may also be a suitable leader sequence, a nontranslated region of an mRNA that is important for translation by the host cell. The leader sequence is operably linked to the 5'-terminus of the polynucleotide encoding the variant. Any leader sequence that is functional in the host cell may be used.
[0359] Suitable leaders for yeast host cells are obtained from the genes for Saccharomyces cerevisiae enolase (ENO-1), Saccharomyces cerevisiae 3-phosphoglycerate kinase, Saccharomyces cerevisiae alpha-factor, and Saccharomyces cerevisiae alcohol dehydrogenase/glyceraldehyde-3-phosphate dehydrogenase (ADH2/GAP).
[0360] The control sequence may also be a polyadenylation sequence, a sequence operably linked to the 3'-terminus of the variant-encoding sequence and, when transcribed, is recognized by the host cell as a signal to add polyadenosine residues to transcribed mRNA. Any polyadenylation sequence that is functional in the host cell may be used.
[0361] Useful polyadenylation sequences for yeast host cells are described by Guo and Sherman, 1995, Mol. Cellular Biol. 15: 5983-5990.
[0362] The control sequence may also be a signal peptide coding region that encodes a signal peptide linked to the N-terminus of a variant and directs the variant into the cell's secretory pathway. The 5'-end of the coding sequence of the polynucleotide may inherently contain a signal peptide coding region naturally linked in translation reading frame with the segment of the coding region that encodes the variant. Alternatively, the 5'-end of the coding sequence may contain a signal peptide coding region that is foreign to the coding sequence. The foreign signal peptide coding region may be required where the coding sequence does not naturally contain a signal peptide coding region. Alternatively, the foreign signal peptide coding region may simply replace the natural signal peptide coding region in order to enhance secretion of the variant. However, any signal peptide coding region that directs the expressed variant into the secretory pathway of a host cell may be used.
[0363] Useful signal peptides for yeast host cells are obtained from the genes for Saccharomyces cerevisiae alpha-factor and Saccharomyces cerevisiae invertase. Other useful signal peptide coding sequences are described by Romanos et al., 1992, supra. The skilled person knows useful signal peptides for use In rice and mammalian cells, such as CHO or HEK.
[0364] Where both signal peptide and propeptide regions are present at the N-terminus of a variant, the propeptide region is positioned next to the N-terminus of the variant and the signal peptide region is positioned next to the N-terminus of the propeptide region.
Plasmids
[0365] A fifth aspect of the invention provides a plasmid comprising the polynucleotide of the fourth aspect of the invention. The plasmid may be a 2 micron based plasmid such as those described in WO 2005/061719, WO 2005/061718 and WO 2006/067511 (all incorporated herein by reference). The plasmid may exhibit enhanced chaperone activity, for example through over expression of a chaperone, particularly PDI. Preferred helper proteins include PDI1, AHA1, ATP11, CCT2, CCT3, CCT4, CCT5, CCT6, CCT7, CCT8, CNS1, CPR3, CPR6, DER1, DER3, DOA4, ERO1, EUG1, ERV2, EPS1, FKB2, FMO1, HCH1, HRD3, HSP10, HSP12, HSP104, HSP26, HSP30, HSP42, HSP60, HSP78, HSP82, KAR2, JEM1, MDJ1, MDJ2, MPD1, MPD2, PDI1, PFD1, ABC1, APJ1, ATP11, ATP12, BTT1, CDC37, CPR7, HSC82, KAR2, LHS1, MGE1, MRS11, NOB1, ECM10, SCJ1, SSA1, SSA2, SSA3, SSA4, SSB1, SSB2, SSC1, SSE2, SIL1, SLS1, ORM1, ORM2, PER1, PTC2, PSE1, UBC7, UBI4 and HAC1 or a truncated intronless HAC1 (Valkonen et al. 2003, Applied Environ. Micro., 69, 2065). Such helper proteins are disclosed in WO 2005/061718, WO 2006/067511 and WO 2006/136831 (all incorporated herein by reference).
Host Cells
[0366] A sixth aspect of the invention provides an expression system such as a host cell comprising a polynucleotide according to the fourth aspect of the invention and/or a plasmid of the fifth aspect of the invention. Preferably the host cell is a mammalian cell such as a human or bovine cell, or a fungal cell such as a yeast cell. Alternatively, the host cell may be a bacterial cell such as a Bacillus or Escherichia coli or a viral cell such as Baculovirus or a plant cell such as a rice e.g. Oryza sativa. Most preferably, the cell is a yeast cell such as a Saccharomyces (e.g. S. cerevisiae), a Pichia or an Aspergillus cell.
Conjugates
[0367] A seventh aspect of the invention provides a conjugate which comprises a conjugation partner, such as a bioactive compound, radiopharmaceutical or imaging agent, and a polypeptide according to the first, second or third aspect of the invention, wherein the conjugation partner is linked to the polypeptide through a conjugation-competent cysteine residue of the polypeptide. The conjugation partner may be a bioactive, therapeutic, diagnostic or imaging compound such as those mentioned herein. The conjugate may comprise 2 or more, (several, for example 2, 3, 4, 5, 6, 7,8, 9 or 10), conjugation partners which may each be different and/or may be multiple copies of the same compound. Preferably, each conjugation partner is linked to the polypeptide through a conjugation-competent cysteine residue of the polypeptide, however conjugation partners may be linked by other means for example by a genetic fusion or covalent bonds to non-cysteine amino acids such as lysine.
[0368] A related aspect provides a use of a polypeptide according to the invention for the production of a thio-albumin-conjugate.
Conjugation Partner
[0369] The term `conjugation partner` includes bioactive agents, imaging agents, diagnostic agents, contrast agents, radiopharmaceuticals and therapeutic compounds such as chemotherapeutic drugs and radiopharmaceuticals. A thio-albumin of the invention may be conjugated to one or more (e.g. several) conjugation partners.
Imaging Agents, Diagnostic Compounds, Contrast Agents and Therapeutic Compounds
[0370] The use of diagnostic agents, imaging agents and biological "contrast" agents are well known to the art. A diagnostic agent is any pharmaceutical product used as part of a diagnostic test (i.e. together with the equipment and procedures that are needed to assess the test result). The diagnostic agent may be used in vivo, ex vivo or in vitro.
[0371] The ability of albumin to accumulate in damaged muscle fibres of dystrophic muscle has been well described. For example, a Gadolinium-DTPA-albumin conjugate may be used as a combined diagnostic and therapeutic tool to visualize and monitor, for example, dystrophic muscle by magnetic resonance imaging (MRI) and for the delivery of putative therapeutics bound to albumin for effective targeting to dystrophic muscle (Amthor et al. (2004), Neuromuscular Disorders 14912: 791-796). Malignant tumours often show an increased uptake and metabolism of albumin. The use of gadolinium-albumin conjugate has also been described for improved imaging of malignant tumours and to determine by MRI tumours sensitive to a therapy with drug-conjugated albumin (Kiessling et al. (2002), Investigative Radiology 37(4): 93-198).
[0372] Current imaging agents often degrade quickly whilst longer-lasting agents are often toxic. The use of albumin conjugates may be especially useful to increase the half-life of imaging agents and would therefore permit imaging over an extended period of time. WO 2005/082423 (incorporated herein by reference) describes the use of serum albumin conjugated to fluorescent substances for imaging.
[0373] A thio-albumin of this invention may be conjugated to two or more (several) molecules selected from bioactive, imaging agents, diagnostic agents, therapeutic compounds and contrast agents.
[0374] Tumours (and muscle degeneration) show enhanced uptake of albumin (EPR: Enhanced Permeation and Retention). Albumin conjugates may be used for enhanced imaging, and also to assess whether tumours (or other tissues and organs) would be suitable for albumin conjugated drugs.
Bioactive Compounds
[0375] The bioactive compound may be a therapeutic or diagnostic compound. The therapeutic compound may be a chemotherapy drug for use in cancer chemotherapy. It may be cytostatic or cytotoxic; it may be a tumor-inhibiting agent.
[0376] The bioactive compound may already contain a free thiol group, e.g. a polypeptide containing a Cysteine residue with a free thiol group. Alternatively, the bioactive compound may be modified so as to contain a free thiol group. Thus, the amino acid sequence of a polypeptide may be altered so as to include a Cysteine residue with a free thiol group, or the bioactive compound may be chemically derivatized to include a free thiol group.
[0377] The bioactive compound may be a polypeptide (protein), particularly a recombinant protein pharmaceutical. It may be a chemotherapy or radiotherapy drug used to treat cancers and other related diseases.
[0378] The free thiol containing albumin mutein of the invention (thio-albumin) can be conjugated via the free thiol group, or groups if the albumin mutein of the invention contains more than one free thiol, to at least one (e.g. several) bioactive compound by methods know to the art. The bioactive compound includes but is not limited to, peptides, polypeptides or proteins (either natural, recombinant, or synthetic) (Debinski, (2002) Cancer Investigation 20, 801-809, O'Keefe and Draper et al., (1985) JBC 260, 932-937, Xia et al., (2000) J. Pharmacology Experimental Therapeutics 295, 594-600, Kavimandan et al., (2006), Bioconjugate Chem. 17, 1376-1384, Humphries, et al., (1994) J. Tissue Culture Methods 16, 239-242, Wenning et al., (1998) Biotech. Bioeng. 57, 484-496, Yazdi and Murphy, (1994), Cancer Research 54, 6387-6394, Weaver and Laske (2003) J. Neuro-Oncology 65, 3-13, Widera et al., (2003) Pharmaceutical Research 20, 1231-1238, Daniels, T. R. et al. (2006) Clinical Immunology 121, 159-176 and the references included therein); therapeutic and diagnostic drugs or compounds (Mishra et al., (2006) J. Drug Targeting 14, 45-53, Lim and Shen, (2004) Pharmaceutical Research 21, 1985-1992, Fritzer et al., (1996) Biochemical Pharmacology 51, 489-493, Lubgan and Jozwiak (2002) Cell. Mol. Biol. Lett. 7, 98, Daniels, T. R. et al. (2006) Clinical Immunology 121, 159-176 and the references included therein); high molecular weight complexes including but not limited to liposomes, viruses and nanoparticles (Mishra et al., (2006) J. Drug Targeting 14, 45-53, Daniels, T. R. et al. (2006) Clinical Immunology 121, 159-176 and the references included therein); nucleic acids and radionuclides, including DNA, RNA (including siRNA) and their analogs (Lee et al., (2005), Arch. Pharm. Res. 28, 722-729, Huang et al., (2007) FASEB J. 21, 1117-1125, Daniels, T. R. et al. (2006) Clinical Immunology 121, 159-176 and the references included therein) and devices (Humphries, et al, (1994) J. Tissue Culture Methods 16, 239-242 and the references included therein). Additionally the entity can itself be modified by methods known to the art.
Therapeutic Compounds
[0379] Examples of therapeutic compounds include: 4-1BB ligand, 5-helix, A human C-C chemokine, A human L105 chemokine, A human L105 chemokine designated huL105_3, A monokine induced by gamma-interferon (MIG), A partial CXCR4B protein, A platelet basic protein (PBP), al-antitrypsin, ACRP-30 Homologue, Complement Component C1q C, Adenoid-expressed chemokine (ADEC), aFGF, FGF-1, AGF, AGF Protein, albumin, an etoposide, angiostatin, Anthrax vaccine, Antibodies specific for collapsin, antistasin, Anti-TGF beta family antibodies, antithrombin III, APM-1, ACRP-30, Famoxin, apo-lipoprotein species, Arylsulfatase B, b57 Protein, BCMA, Beta-thromboglobulin protein (beta-TG), bFGF, FGF2, Blood coagulation factors, BMP Processing Enzyme Furin, BMP-10, BMP-12, BMP-15, BMP-17, BMP-18, BMP-2B, BMP-4, BMP-5, BMP-6, BMP-9, Bone Morphogenic Protein-2, calcitonin, Calpain-10a, Calpain-10b, Calpain-10c, Cancer Vaccine, Carboxypeptidase, C-C chemokine, MCP2, CCR5 variant, CCR7, CCR7, CD11a Mab, CD137, 4-1BB Receptor Protein, CD20 Mab, CD27, CD27L, CD30, CD30 ligand, CD33 immunotoxin, CD40, CD40L, CD52 Mab, Cerebus Protein, Chemokine Eotaxin, Chemokine hIL-8, Chemokine hMCP1, Chemokine hMCP1a, Chemokine hMCP1b, Chemokine hMCP2, Chemokine hMCP3, Chemokine hSDF1b, Chemokine MCP-4, chemokine TECK and TECK variant, Chemokine-like protein IL-8M1 Full-Length and Mature, Chemokine-like protein IL-8M10 Full-Length and Mature, Chemokine-like protein IL-8M3, Chemokine-like protein IL-8M8 Full-Length and Mature, Chemokine-like protein IL-8M9 Full-Length and Mature, Chemokine-like protein PF4-414 Full-Length and Mature, Chemokine-like protein PF4-426 Full-Length and Mature, Chemokine-like protein PF4-M2 Full-Length and Mature, Cholera vaccine, Chondromodulin-like protein, c-kit ligand, SCF, Mast cell growth factor, MGF, Fibrosarcoma-derived stem cell factor, CNTF and fragment thereof (such as CNTFA.times.15'(Axokinen.TM.)), coagulation factors in both pre and active forms, collagens, Complement C5 Mab, Connective tissue activating protein-III, CTAA16.88 Mab, CTAP-III, CTLA4-Ig, CTLA-8, CXCR3, CXC chemokine receptor 3, cyanovirin-N, Darbepoetin, designated exodus, designated huL105_7, DIL-40, Dnase, EDAR, EGF Receptor Mab, ENA-78, Endostatin, Eotaxin, Epithelial neutrophil activating protein-78, EPO receptor, EPOR, erythropoietin (EPO) and EPO mimics, Eutropin, Exodus protein, Factor IX, Factor VII, Factor VIII, Factor X and Factor XIII, FAS Ligand Inhibitory Protein (DcR3), FasL, FGF, FGF-12, Fibroblast growth factor homologous factor-1, FGF-15, FGF-16, FGF-18, FGF-3, INT-2, FGF-4, gelonin, HST-1, HBGF-4, FGF-5, FGF-6, Heparin binding secreted transforming factor-2, FGF-8, FGF-9, Glia activating factor, fibrinogen, flt-1, flt-3 ligand, Follicle stimulating hormone Alpha subunit, Follicle stimulating hormone Beta subunit, Follitropin, Fractalkine, fragment. myofibrillar protein Troponin I, FSH, Galactosidase, Galectin-4, G-CSF, GDF-1, Gene therapy, Glioma-derived growth factor, glucagon, glucagon-like peptides, Glucocerebrosidase, glucose oxidase, Glucosidase, Glycodelin-A, Progesterone-associated endometrial protein, GM-CSF, gonadotropin, Granulocyte chemotactic protein-2 (GCP-2), Granulocyte-macrophage colony stimulating factor, growth hormone, Growth related oncogene-alpha (GRO-alpha), Growth related oncogene-beta (GRO-beta), Growth related oncogene-gamma (GRO-gamma), hAPO-4, TROY, hCG, Hepatitus B surface Antigen, Hepatitus B Vaccine, HER2 Receptor Mab, hirudin, HIV gp120, HIV gp41, HIV Inhibitor Peptide, HIV Inhibitor Peptide, HIV Inhibitor Peptide, HIV protease inhibiting peptides, HIV-1 protease inhibitors, HPV vaccine, Human 6CKine protein, Human Act-2 protein, Human adipogenesis inhibitory factor, human B cell stimulating factor-2 receptor, Human beta-chemokine H1305 (MCP-2), Human C-C chemokine DGWCC, Human CC chemokine ELC protein, Human CC type chemokine interleukin C, Human CCC3 protein, Human CCF18 chemokine, Human CC-type chemokine protein designated SLC (secondary lymphoid chemokine), Human chemokine beta-8 short forms, Human chemokine C10, Human chemokine CC-2, Human chemokine CC-3, Human chemokine CCR-2, Human chemokine Ckbeta-7, Human chemokine ENA-78, Human chemokine eotaxin, Human chemokine GRO alpha, Human chemokine GROalpha, Human chemokine GRObeta, Human chemokine HCC-1, Human chemokine HCC-1, Human chemokine 1-309, Human chemokine IP-10, Human chemokine L105_3, Human chemokine L105_7, Human chemokine MIG, Human chemokine MIG-beta protein, Human chemokine MIP-1alpha, Human chemokine MIP1beta, Human chemokine MIP-3alpha, Human chemokine MIP-3beta, Human chemokine PF4, Human chemokine protein 331D5, Human chemokine protein 61164, Human chemokine receptor CXCR3, Human chemokine SDF1alpha, Human chemokine SDF1beta, Human chemokine ZSIG-35, Human Chr19Kine protein, Human CKbeta-9, Human CX3C 111 amino acid chemokine, Human DNAX interleukin-40, Human DVic-1 C-C chemokine, Human EDIRF I protein sequence, Human EDIRF II protein sequence, Human eosinocyte CC type chemokine eotaxin, Human eosinophil-expressed chemokine (EEC), Human fast twitch skeletal muscle troponin C, Human fast twitch skeletal muscle troponin I, Human fast twitch skeletal muscle Troponin subunit C, Human fast twitch skeletal muscle Troponin subunit I Protein, Human fast twitch skeletal muscle Troponin subunit T, Human fast twitch skeletal muscle troponin T, Human foetal spleen expressed chemokine, FSEC, Human GM-CSF receptor, Human gro-alpha chemokine, Human gro-beta chemokine, Human gro-gamma chemokine, Human IL-16 protein, Human IL-1RD10 protein sequence, Human IL-1RD9, Human IL-5 receptor alpha chain, Human IL-6 receptor, Human IL-8 receptor protein hIL8RA, Human IL-8 receptor protein hIL8RB, Human IL-9 receptor protein, Human IL-9 receptor protein variant #3, Human IL-9 receptor protein variant fragment, Human IL-9 receptor protein variant fragment #3, Human interleukin 1 delta, Human interleukin 10, Human interleukin 18, Human interleukin 18 derivatives, Human interleukin-1 beta precursor, Human interleukin-1 beta precursor, Human interleukin-1 receptor accessory protein, Human interleukin-1 receptor antagonist beta, Human interleukin-1 type-3 receptor, Human interleukin-10 (precursor), Human interleukin-11 receptor, Human interleukin-12 40 kD subunit, Human interleukin-12 beta-1 receptor, Human interleukin-12 beta-2 receptor, Human interleukin-12 p35 protein, Human interleukin-12 p40 protein, Human interleukin-12 receptor, Human interleukin-13 alpha receptor, Human interleukin-13 beta receptor, Human interleukin-15, Human interleukin-15 receptor from clone P1, Human interleukin-17 receptor, Human interleukin-18 protein (IL-18), Human interleukin-3, human interleukin-3 receptor, Human interleukin-3 variant, Human interleukin-4 receptor, Human interleukin-5, Human interleukin-6, Human interleukin-7, Human interleukin-7, Human interleukin-8 (IL-8), Human intracellular IL-1 receptor antagonist, Human IP-10 and HIV-1 gp120 hypervariable region fusion protein, Human IP-10 and human Muc-1 core epitope (VNT) fusion protein, human liver and activation regulated chemokine (LARC), Human Lkn-1 Full-Length and Mature protein, Human mammary associated chemokine (MACK) protein Full-Length and Mature, Human mature chemokine Ckbeta-7, Human mature gro-alpha, Human mature gro-gamma polypeptide used to treat sepsis, Human MCP-3 and human Muc-1 core epitope (VNT) fusion protein, Human MI10 protein, Human MI1A protein, Human monocyte chemoattractant factor hMCP-1, Human monocyte chemoattractant factor hMCP-3, Human monocyte chemotactic proprotein (MCPP) sequence, Human neurotactin chemokine like domain, Human non-ELR CXC chemokine H174, Human non-ELR CXC chemokine IP10, Human non-ELR CXC chemokine Mig, Human PAI-1 mutants, Human protein with IL-16 activity, Human protein with IL-16 activity, Human secondary lymphoid chemokine (SLC), Human SISD protein, Human STCP-1, Human stromal cell-derived chemokine, SDF-1, Human T cell mixed lymphocyte reaction expressed chemokine (TMEC), Human thymus and activation regulated cytokine (TARC), Human thymus expressed, Human TNF-alpha, Human TNF-beta (LT-alpha), Human type CC chemokine eotaxin 3 protein sequence, Human type II interleukin-1 receptor, Human wild-type interleukin-4 (hIL-4) protein, Human ZCHEMO-8 protein, Humanized Anti-VEGF Antibodies, and fragments thereof, Humanized Anti-VEGF Antibodies, and fragments thereof, Hyaluronidase, ICE 10 kD subunit, ICE 20 kD subunit, ICE 22 kD subunit, Iduronate-2-sulfatase, Iduronidase, IL-1 alpha, IL-1 beta, IL-1 inhibitor (IL-1i), IL-1 mature, IL-10 receptor, IL-11, IL-11, IL-12 p40 subunit, IL-13, IL-14, IL-15, IL-15 receptor, IL-17, IL-17 receptor, IL-19, IL-1i fragments, IL1-receptor antagonist, IL-21 (TIF), IL-3 containing fusion protein, IL-3 mutant proteins, IL-3 variants, IL-4, IL-4 muteins, IL-4 mutein Y124G, IL-4 mutein Y124X, IL-5, IL-5 muteins, 11-5 receptor, IL-6, 11-6 receptor, IL-7 receptor clone, IL-8 receptor, IL-9 mature protein variant (Met117 version), immunoglobulins or immunoglobulin-based molecules or fragment of either (e.g. a Small Modular ImmunoPharmaceutical.TM. ("SMIP") or dAb, Fab' fragments, F(ab')2, scAb, scFv or scFv fragment), including but not limited to plasminogen, Influenza Vaccine, Inhibin alpha, Inhibin beta, insulin, insulin-like growth factor, Integrin Mab, inter-alpha trypsin inhibitor, inter-alpha trypsin inhibitor, Interferon gamma-inducible protein (IP-10), interferons (such as interferon alpha species and sub-species, interferon beta species and sub-species, interferon gamma species and sub-species), interleukin 6, interleukin 8 (IL-8) receptor, interleukin 8 receptor B, interleukin-1alpha, interleukin-2 receptor associated protein p43, interleukin-3, interleukin-4 muteins, interleukin-8 (IL-8) protein, interleukin-9, interleukin-9 (IL-9) mature protein (Thr117 version), interleukins (such as IL10, IL11 and IL2), Japanese encephalitis vaccine, Kalikrein Inhibitor, Keratinocyte growth factor, Kunitz domain protein (such as aprotinin, amyloid precursor protein and those described in WO 03/066824, with or without albumin fusions), LACI, lactoferrin, Latent TGF-beta binding protein II, leptin, Liver expressed chemokine-1 (LVEC-1), Liver expressed chemokine-2 (LVEC-2), LT-alpha, LT-beta, Luteinization Hormone, Lyme Vaccine, Lymphotactin, Macrophage derived chemokine analogue MDC (n+1), Macrophage derived chemokine analogue MDC-eyfy, Macrophage derived chemokine analogue MDC-yl, Macrophage-derived chemokine (MDC), Maspin, Protease Inhibitor 5, MCP-1 receptor, MCP-1a, MCP-1b, MCP-3, MCP-4 receptor, M-CSF, Melanoma inhibiting protein, Membrane-bound proteins, Met117 human interleukin 9, MIP-3 alpha, MIP-3 beta, MIP-Gamma, MIRAP, Modified Rantes, monoclonal antibody, MP52, Mutant interleukin 6 S176R, myofibrillar contractile protein Troponin I, Natriuretic Peptide, Nerve Growth Factor-beta, Nerve Growth Factor-beta2, Neuropilin-1, Neuropilin-2, Neurotactin, Neurotrophin-3, Neurotrophin-4, Neurotrophin-4a, Neurotrophin-4b, Neurotrophin-4c, Neurotrophin-4d, Neutrophil activating peptide-2 (NAP-2), NOGO-66 Receptor, NOGO-A, NOGO-B, NOGO-C, Novel beta-chemokine designated PTEC, N-terminal modified chemokine GroHEK/hSDF-1alpha, N-terminal modified chemokine GroHEK/hSDF-1beta, N-terminal modified chemokine met-hSDF-1 alpha, N-terminal modified chemokine met-hSDF-1 beta, OPGL, Osteogenic Protein-1 (OP-1), BMP-7, Osteogenic Protein-2, OX40, ACT-4, OX40L, Oxytocin (Neurophysin I), parathyroid hormone, Patched, Patched-2, PDGF-D, Pertussis toxoid, Pituitary expressed chemokine (PGEC), Placental Growth Factor, Placental Growth Factor-2, Plasminogen Activator Inhibitor-1 (PAI-1), Plasminogen Activator Inhibitor-2 (PAI-2), Platelet derived growth factor, Platelet derived growth factor Bv-sis, Platelet derived growth factor precursor A, Platelet derived growth factor precursor B, Platelet Mab, platelet-derived endothelial cell growth factor (PD-ECGF), Platelet-Derived Growth Factor A chain, Platelet-Derived Growth Factor B chain, polypeptide used to treat sepsis, Preproapolipoprotein "milano" variant, Preproapolipoprotein "paris" variant, pre-thrombin, Primate CC chemokine "ILINCK", Primate CXC chemokine "IBICK", proinsulin, Prolactin, Prolactin2, prosaptide, Protease inhibitor peptides, Protein C, Protein S, pro-thrombin, prourokinase, RANTES, RANTES 8-68, RANTES 9-68, RANTES peptide, RANTES receptor, Recombinant interleukin-16, Resistin, restrictocin, Retroviral protease inhibitors, ricin, Rotavirus Vaccine, RSV Mab, saporin, sarcin, Secreted and Transmembrane polypeptides, serum cholinesterase, serum protein (such as a blood clotting factor), Soluble BMP Receptor Kinase Protein-3, Soluble VEGF Receptor, Stem Cell Inhibitory Factor, Straphylococcus Vaccine, Stromal Derived Factor-1 alpha, Stromal Derived Factor-1 beta, Substance P (tachykinin), T1249 peptide, T20 peptide, T4 Endonuclease, TACI, Tarc, TGF-beta 1, TGF-beta 2, Thr117 human interleukin 9, thrombin, thrombopoietin, thrombopoietin derivative 1, thrombopoietin derivative 2, thrombopoietin derivative 3, thrombopoietin derivative 4, thrombopoietin derivative 5, thrombopoietin derivative 6, thrombopoietin derivative 7, Thymus expressed chemokine (TECK), Thyroid stimulating Hormone, tick anticoagulant peptide, Tim-1 protein, TNF-alpha precursor, TNF-R, TNF-RII, TNF p75 Receptor, Death Receptor, tissue plasminogen activator (tPA), transferrin, transforming growth factor beta, Troponin peptides, Truncated monocyte chemotactic protein 2 (6-76), Truncated RANTES protein (3-68), tumour necrosis factor, Urate Oxidase, urokinase, Vasopressin (Neurophysin II), VEGF R-3, flt-4, VEGF Receptor, KDR, flk-1, VEGF-110, VEGF-121, VEGF-138, VEGF-145, VEGF-162, VEGF-165, VEGF-182, VEGF-189, VEGF-206, VEGF-D, VEGF-E, VEGF-X, von Willebrand's factor, Wild type monocyte chemotactic protein 2, ZTGF-beta 9.
Chemotherapy Drugs
[0380] Examples of chemotherapy drugs include: 13-cis-Retinoic Acid, 2-CdA, 2-Chlorodeoxyadenosine, 5-Azacitidine, 5-Fluorouracil, 5-FU, 6-Mercaptopurine, 6-MP, 6-TG, 6-Thioguanine, Abraxane, Accutane.RTM., Actinomycin-D, Adriamycin.RTM., Adrucil.RTM., Agrylin.RTM., Ala-Cort.RTM., Aldesleukin, Alemtuzumab, ALIMTA, Alitretinoin, Alkaban-AQ.RTM., Alkeran.RTM., All-transretinoic Acid, Alpha Interferon, Altretamine, Amethopterin, Amifostine, Aminoglutethimide, Anagrelide, Anandron.RTM., Anastrozole, Arabinosylcytosine, Ara-C, Aranesp.RTM., Aredia.RTM., Arimidex.RTM., Aromasin.RTM., Arranon.RTM., Arsenic Trioxide, Asparaginase, ATRA, Avastin.RTM., Azacitidine, BCG, BCNU, Bevacizumab, Bexarotene, BEXXAR.RTM., Bicalutamide, BiCNU, Blenoxane.RTM., Bleomycin, Bortezomib, Busulfan, Busulfex.RTM., C225, Calcium Leucovorin, Campath.RTM., Camptosar.RTM., Camptothecin-11, Capecitabine, Carac.TM., Carboplatin, Carmustine, Carmustine Wafer, Casodex.RTM., CC-5013, CCNU, CDDP, CeeNU, Cerubidine.RTM., Cetuximab, Chlorambucil, Cisplatin, Citrovorum Factor, Cladribine, Cortisone, Cosmegen.RTM., CPT-11, Cyclophosphamide, Cytadren.RTM., Cytarabine, Cytarabine Liposomal, Cytosar-U.RTM., Cytoxan.RTM., Dacarbazine, Dacogen, Dactinomycin, Darbepoetin Alfa, Dasatinib, Daunomycin, Daunorubicin, Daunorubicin Hydrochloride, Daunorubicin Liposomal, DaunoXome.RTM., Decadron, Decitabine, Delta-Cortef.RTM., Deltasone.RTM., Denileukin diftitox, DepoCyt.TM., Dexamethasone, Dexamethasone acetate, Dexamethasone Sodium Phosphate, Dexasone, Dexrazoxane, DHAD, DIC, Diodex, Docetaxel, Doxil.RTM., Doxorubicin, Doxorubicin liposomal, Droxia, DTIC, DTIC-Dome.RTM., Duralone.RTM., Efudex.RTM., Eligard.TM., Ellence.TM., Eloxatin.TM., Elspar.RTM., Emcyt.RTM., Epirubicin, Epoetin alfa, Erbitux.TM., Erlotinib, Erwinia L-asparaginase, Estramustine, Ethyol, Etopophos.RTM., Etoposide, Etoposide Phosphate, Eulexin.RTM., Evista.RTM., Exemestane, Fareston.RTM., Faslodex.RTM., Femara.RTM., Filgrastim, Floxuridine, Fludara.RTM., Fludarabine, Fluoroplex.RTM., Fluorouracil, Fluoxymesterone, Flutamide, Folinic Acid, FUDR.RTM., Fulvestrant, G-CSF, Gefitinib, Gemcitabine, Gemtuzumab ozogamicin, Gemzar.RTM., Gleevec.TM., Gliadel.RTM. Wafer, GM-CSF, Goserelin, Granulocyte-Colony Stimulating Factor, Granulocyte Macrophage Colony Stimulating Factor, Halotestin.RTM., Herceptin.RTM., Hexadrol, Hexalen.RTM., Hexamethylmelamine, HMM, Hycamtin.RTM., Hydrea.RTM., Hydrocort Acetate.RTM., Hydrocortisone, Hydrocortisone Sodium Phosphate, Hydrocortisone Sodium Succinate, Hydrocortone Phosphate, Hydroxyurea, Ibritumomab, Ibritumomab Tiuxetan, Idamycin.RTM., Idarubicin, Ifex.RTM., IFN-alpha, Ifosfamide, IL-11, IL-2, Imatinib mesylate, Imidazole Carboxamide, Interferon alfa, Interferon Alfa-2b (PEG Conjugate), interleukin-2, interleukin-11, Intron A.degree. (interferon alfa-2b), Iressa.RTM., Irinotecan, Isotretinoin, Kidrolase.RTM., Lanacort.RTM., Lapatinib, L-asparaginase, LCR, Lenalidomide, Letrozole, Leucovorin, Leukeran, Leukine.TM., Leuprolide, Leurocristine, Leustatin.TM., Liposomal Ara-C, Liquid Pred.RTM., Lomustine, L-PAM, L-Sarcolysin, Lupron.RTM., Lupron Depot.RTM., Matulane.RTM., Maxidex, Mechlorethamine, Mechlorethamine Hydrochloride, Medralone.RTM., Medrol.RTM., Megace.RTM., Megestrol, Megestrol Acetate, Melphalan, Mercaptopurine, Mesna, Mesnex.TM., Methotrexate, Methotrexate Sodium, Methylprednisolone, Meticorten.RTM., Mitomycin, Mitomycin-C, Mitoxantrone, M-Prednisol.RTM., MTC, MTX, Mustargen.RTM., Mustine, Mutamycin.RTM., Myleran.RTM., Mylocel.TM., Mylotarg.RTM., Navelbine.RTM., Nelarabine, Neosar.RTM., Neulasta.TM., Neumega.RTM., Neupogen.RTM., Nexavar.RTM., Nilandron.RTM., Nilutamide, Nipent.RTM., Nitrogen Mustard, Novaldex.RTM., Novantrone.RTM., Octreotide, Octreotide acetate, Oncospar.RTM., Oncovin.RTM., Ontak.RTM., Onxal.TM., Oprevelkin, Orapred.RTM., Orasone.RTM., Oxaliplatin, Paclitaxel, Paclitaxel Protein-bound, Pamidronate, Panitumumab, Panretin.RTM., Paraplatin.RTM., Pediapred.RTM., PEG Interferon, Pegaspargase, Pegfilgrastim, PEG-INTRON.TM., PEG-L-asparaginase, PEMETREXED, Pentostatin, Phenylalanine Mustard, Platinol.RTM., Platinol-AQ.RTM., Prednisolone, Prednisone, Prelone.RTM., Procarbazine, PROCRIT.RTM., Proleukin.RTM., Prolifeprospan 20 with Carmustine Implant, Purinethol.RTM., Raloxifene, Revlimid.RTM., Rheumatrex.RTM., Rituxan.RTM., Rituximab, Roferon-A.RTM. (Interferon Alfa-2a), Rubex.RTM., Rubidomycin hydrochloride, Sandostatin.RTM., Sandostatin LAR.RTM., Sargramostim, Solu-Cortef.RTM., Solu-Medrol.RTM., Sorafenib, SPRYCEL.TM., STI-571, Streptozocin, SU11248, Sunitinib, Sutent.RTM., Tamoxifen, Tarceva.RTM., Targretin.RTM., Taxol.RTM., Taxotere.RTM., Temodar.RTM., Temozolomide, Teniposide, TESPA, Thalidomide, Thalomid.RTM., TheraCys.RTM., Thioguanine, Thioguanine Tabloid.RTM., Thiophosphoamide, Thioplex.RTM., Thiotepa, TICE.RTM., Toposar.RTM., Topotecan, Toremifene, Tositumomab, Trastuzumab, Tretinoin, Trexall.TM., Trisenox.RTM., TSPA, TYKERB.RTM., VCR, Vectibix.TM., Velban.RTM., Velcade.RTM., VePesid.RTM., Vesanoid.RTM., Viadur.TM., Vidaza.RTM., Vinblastine, Vinblastine Sulfate, Vincasar Pfs.RTM., Vincristine, Vinorelbine, Vinorelbine tartrate, VLB, VM-26, Vorinostat, VP-16, Vumon.RTM., Xeloda.RTM., Zanosar.RTM., Zevalin.TM., Zinecard.RTM., Zoladex.RTM., Zoledronic acid, Zolinza, Zometa.RTM..
Radiopharmaceuticals
[0381] Examples of radiopharmaceuticals include: Carbon-11, Carbon-14, Chromium-51, Cobalt-57, Cobalt-58, Erbium-169, Fluorine-18, Gallium-67, Gold-198, Indium-111, Indium-113m, Iodine-123, Iodine-125, Iodine-131, Iron-59, Krypton-81m, Nitrogen-13, Oxygen-15, Phosphorous-32, Rhenium-186, Rubidium-82, Samarium-153, Selenium-75, Strontium-89, Technetium-99m, Thallium-201, Tritium, Xenon-127, Xenon-133, Yttrium-90.
Imaging Agents
[0382] Examples of imaging agents include: Gadolinium, magnetite, manganese, technetium, 1125,1131, P32, TI201, Iopamidol, PET-FDG.
Preparation of a Polynucleotide
[0383] An eighth aspect of the invention provides a method of producing a polynucleotide comprising:
[0384] (a) providing a nucleic acid molecule encoding a parent albumin or fragment thereof; and
[0385] (b) modifying the nucleic acid sequence of the nucleic acid molecule to encode a conjugation-competent polypeptide which is at least 60% identical to human albumin, particularly residues 1 to 585 of the mature human albumin polypeptide sequence of SEQ ID NO. 2, or a fragment thereof, wherein at least one (e.g. several) position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue.
[0386] Suitably, modifying the nucleic acid sequence comprises introducing an alteration such that at least one (e.g. several) conjugation-competent cysteine as provided for in step (b) is introduced into the encoded polypeptide. Preferred alterations are as described in relation to the first and second aspects of the invention.
[0387] It is preferred that the parent albumin comprises or consists of:
[0388] (a) a polypeptide having at least 70% sequence identity to the mature polypeptide of SEQ ID NO. 2;
[0389] (b) a polypeptide encoded by a polynucleotide that hybridizes under low stringency conditions with (i) the mature polypeptide coding sequence of SEQ ID NO. 2, or (ii) the full-length complement of (i);
[0390] (c) a polypeptide encoded by a polynucleotide having at least 60% identity to the mature polypeptide coding sequence of SEQ ID NO. 2; and/or
[0391] (d) a fragment of the mature polypeptide of SEQ ID NO. 2.
[0392] Suitably, the parent albumin comprises or consists of the HSA polypeptide sequence of SEQ ID NO. 2 or a variant or fragment thereof.
[0393] The variant polynucleotides can be prepared by those skilled persons using any mutagenesis procedure known in the art, such as site-directed mutagenesis, synthetic gene construction, semi-synthetic gene construction, random mutagenesis, shuffling, etc.
[0394] Site-directed mutagenesis is a technique in which one or more (e.g. several) mutations (alterations) are created at one or more (e.g. several) defined sites in a polynucleotide encoding the parent.
[0395] Site-directed mutagenesis can be accomplished in vitro by PCR involving the use of oligonucleotide primers containing the desired mutation. Site-directed mutagenesis can also be performed in vitro by cassette mutagenesis involving the cleavage by a restriction enzyme at a site in the plasmid comprising a polynucleotide encoding the parent and subsequent ligation of an oligonucleotide containing the mutation in the polynucleotide. Usually the restriction enzyme that digests at the plasmid and the oligonucleotide is the same, permitting ligation of the plasmid and insert to one another. See, e.g. Scherer and Davis, 1979, Proc. Natl. Acad. Sci. USA 76: 4949-4955; and Barton et al., 1990, Nucleic Acids Res. 18: 7349-4966.
[0396] Site-directed mutagenesis can also be accomplished in vivo by methods known in the art, see, e.g. U.S. Patent Application Publication: 2004/0171154; Storici et al., 2001, Nature Biotechnol. 19: 773-776; Kren et al., 1998, Nat. Med. 4: 285-290; and Calissano and Macino, 1996, Fungal Genet. Newslett. 43: 15-16.
[0397] Any site-directed mutagenesis procedure can be used in the invention. There are many commercial kits available that can be used to prepare variants.
[0398] Synthetic gene construction entails in vitro synthesis of a designed polynucleotide molecule to encode a polypeptide of interest. Gene synthesis can be performed utilizing a number of techniques, such as the multiplex microchip-based technology described by Tian et al. (2004, Nature 432: 1050-1054) and similar technologies wherein oligonucleotides are synthesized and assembled upon photo-programmable microfluidic chips.
[0399] Single or multiple amino acid substitutions, deletions, and/or insertions can be made and tested using known methods of mutagenesis, recombination, and/or shuffling, followed by a relevant screening procedure, such as those disclosed by Reidhaar-Olson and Sauer, 1988, Science 241: 53-57; Bowie and Sauer, 1989, Proc. Natl. Acad. Sci. USA 86: 2152-2156; WO 95/17413; or WO 95/22625. Other methods that can be used include error-prone PCR, phage display (e.g. Lowman et al., 1991, Biochemistry 30: 10832-10837; U.S. Pat. No. 5,223,409; WO 92/06204) and region-directed mutagenesis (Derbyshire et al., 1986, Gene 46: 145; Ner et al., 1988, DNA 7: 127).
[0400] Mutagenesis/shuffling methods can be combined with high-throughput, automated screening methods to detect activity of cloned, mutagenized polypeptides expressed by host cells (Ness et al., 1999, Nature Biotechnology 17: 893-896). Mutagenized DNA molecules that encode active polypeptides can be recovered from the host cells and rapidly sequenced using standard methods in the art. These methods allow the rapid determination of the importance of individual amino acid residues in a polypeptide.
[0401] Semi-synthetic gene construction is accomplished by combining aspects of synthetic gene construction, and/or site-directed mutagenesis, and/or random mutagenesis, and/or shuffling. Semi-synthetic construction is typified by a process utilizing polynucleotide fragments that are synthesized, in combination with PCR techniques. Defined regions of genes may thus be synthesized de novo, while other regions may be amplified using site-specific mutagenic primers, while yet other regions may be subjected to error-prone PCR or non-error prone PCR amplification. Polynucleotide sub sequences may then be shuffled.
Method of Producing a Polypeptide
[0402] A ninth aspect of the invention provides a method of producing a polypeptide of the invention comprising:
[0403] (a) culturing a host cell according to the invention under conditions that allow expression of the polypeptide; and
[0404] (b) recovering the polypeptide from the host cell and/or from host cell growth medium.
[0405] The method may or may not further comprise determining the receptor binding capacity and/or the conjugation competence of the polypeptide and/or the tendency to exist as a monomer in solution, and optionally selecting a polypeptide which does or does not have a receptor binding capacity and/or conjugation competence and/or selected range of percentage monomer tendency.
[0406] The variants of the invention can be prepared using techniques well known to the skilled person. One convenient way is by cloning a nucleic acid molecule encoding a parent albumin or a fragment thereof and modifying the sequence of the nucleic acid molecule according to the method of the eighth aspect of the invention, preparing a suitable genetic construct where the modified nucleic acid molecule is placed in operative connection with suitable regulatory genetic elements, such as promoter, terminator, activation sites, ribosome binding sites etc., introducing the genetic construct into a suitable host organism, culturing the transformed host organism under conditions leading to expression of the variant and recovering the variant. All these techniques are known in the art and it is within the skills of the average practitioner to design a suitable method for preparing a particular variant according to the invention.
[0407] The variant polypeptide of the invention may also be connected to a signal sequence in order to have the variant polypeptide secreted into the growth medium during culturing of the transformed host organism. It is generally advantageous to have the variant polypeptide secreted into the growth medium in order to ease recovery and purification. The polypeptide may be prepared as a fusion polypeptide as described in relation to the third aspect of the invention. Techniques for preparing variant polypeptides have been disclosed in WO 2009/019314 (included by reference) and these techniques may also be applied to the invention.
[0408] Albumins have been successfully expressed as recombinant proteins in a range of hosts including fungi (including but not limited to Aspergillus (WO 06066595), Kluyveromyces (Fleer 1991, Bio/technology 9, 968-975), Pichia (Kobayashi 1998 Therapeutic Apheresis 2, 257-262) and Saccharomyces (Sleep 1990, Bio/technology 8, 42-46)), bacteria (Pandjaitab 2000, J. Allergy Clin. Immunol. 105, 279-285)), animals (Barash 1993, Transgenic Research 2, 266-276) and plants (including but not limited to potato and tobacco (Sijmons 1990, Bio/technology 8, 217 and Farran 2002, Transgenic Research 11, 337-346) and rice e.g. Oryza sativa) and mammalian cells such as CHO and HEK. The variant polypeptide of the invention is preferably produced recombinantly in a suitable host cell. In principle any host cell capable of producing a polypeptide in suitable amounts may be used and it is within the skills of the average practitioner to select a suitable host cell according to the invention. A preferred host organism is yeast, preferably selected among Saccharomycacae, more preferred Saccharomyces cerevisiae.
[0409] The variant polypeptides of the invention may be recovered and purified from the growth medium using a combination of known separation techniques such as filtration, centrifugation, chromatography, and affinity separation techniques etc. It is within the skills of the average practitioner to purify the variants of the invention using a particular combination of such known separation steps. As an example of purification techniques that may be applied to the variants of the invention can be mentioned the teaching of WO 00/44772.
[0410] In the method of the invention, the host cell may or may not exhibit enhanced chaperone activity. Accordingly, the present invention also provides a method for producing a polypeptide (or protein) of the invention, the method comprising: (a) providing a host cell of the invention comprising a polynucleotide encoding protein product of choice as defined above; and (b) growing the host cell (for example, culturing the host cell in a culture medium); thereby to produce a cell culture or recombinant organism comprising an increased level of the protein product of choice compared to the level of production of the protein product of choice achieved by growing (for example, culturing), under the same conditions, the same host cell that has not been genetically modified to cause over-expression of one or more (e.g. several) helper proteins.
[0411] The step of growing the host cell may or may not involve allowing a host cell derived from a multicellular organism to be regrown into a multicellular recombinant organism (such as a plant or animal) and, optionally, producing one or more (e.g. several) generations of progeny therefrom.
[0412] The thio-albumin may or may not be capable of being expressed at a level of at least 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100% relative to the expression of an unmodified albumin (such as SEQ ID NO. 2) from a suitable expression system, such as yeast (e.g. Saccharomyces, e.g. S. cerevisiae) or an Aspergillus. Relative expression levels can be determined, for example, by expression of the protein followed by quantification by SDS-PAGE, HPLC or Western Blotting. Relative expression levels may be determined in at least 10 liter scale.
[0413] The method may or may not further comprise the step of purifying the thus expressed protein product of choice from the cultured host cell, recombinant organism or culture medium.
[0414] The production method may comprise linking a conjugation partner to the polypeptide of the invention through a conjugation competent cysteine residue of the polypeptide. Suitable conjugation methods and conjugation partners are described herein.
[0415] The thio-albumin or fusions of thio-albumin and another protein or proteins can be expressed as variants with reduced N-linked glycosylation. Accordingly, in case of HSA, it may be particularly advantageous to use a yeast deficient in one or more (e.g. several) protein mannosyl transferases involved in O-glycosylation of proteins, for instance by disruption of the gene coding sequence. Recombinantly expressed proteins can be subject to undesirable post-. translational modifications by the producing host cell. The mannosylated albumin would be able to bind to the lectin Concanavalin A. The amount of mannosylated albumin produced by the yeast can be reduced by using a yeast strain deficient in one or more (e.g. several) of the PMT genes (WO 94/04687). The most convenient way of achieving this is to create a yeast which has a defect in its genome such that a reduced level of one of the Pmt proteins is produced. For example, there may or may not be a deletion, insertion or transposition in the coding sequence or the regulatory regions (or in another gene regulating the expression of one of the PMT genes) such that little or no Pmt protein is produced. Alternatively, the yeast could be transformed to produce an anti-Pmt agent, such as an anti-Pmt antibody. Alternatively, the yeast could be cultured in the presence of a compound that inhibits the activity of one of the PMT genes (Duffy et al, "Inhibition of protein mannosyltransferase 1 (PMT1) activity in the pathogenic yeast Candida albicans", International Conference on Molecular Mechanisms of Fungal Cell Wall Biogenesis, 26-31 Aug. 2001, Monte Verita, Switzerland, Poster Abstract P38). If a yeast other than S. cerevisiae is used, disruption of one or more (e.g. several) of the genes equivalent to the PMT genes of S. cerevisiae is also beneficial, e.g. in Pichia pastoris or Kluyveromyces lactis. The sequence of PMT1 (or any other PMT gene) isolated from S. cerevisiae may be used for the identification or disruption of genes encoding similar enzymatic activities in other fungal species. The cloning of the PMT1 homologue of Kluyveromyces lactis is described in WO 94/04687.
[0416] The variant polypeptides of the invention may be used for delivering a therapeutically beneficial compound (including prophylactically beneficial compound such as a vaccine) to an animal or a human individual in need thereof. Such therapeutically beneficial compounds include, but are not limited to, labels and readily detectable compounds for use in diagnostics, such as various imaging techniques; pharmaceutical active compounds such as drugs, or specifically binding moieties such as antibodies. The variants of the invention may even be connected to two or more (several) different therapeutically beneficial compounds, e.g. an antibody and a drug, which gives the combined molecule the ability to bind specifically to a desired target and thereby provide a high concentration of the connected drug at that particular target.
[0417] The method may further comprise the step of purifying the polypeptide recovered from the host cell and/or from the host cell growth medium. The purification step optionally comprises cell immobilisation, cell separation and/or cell breakage, but always comprises at least one (e.g. several) other purification step different from the step or steps of cell immobilisation, separation and/or breakage.
[0418] Thio-albumin of the invention may be purified from the culture medium by any technique that has been found to be useful for purifying such proteins. Similarly, cell separation techniques, such as centrifugation, filtration (e.g. cross-flow filtration, expanded bed chromatography and the like) are well known in the art. Likewise, methods of cell breakage, including beadmilling, sonication, enzymatic exposure and the like are well known in the art.
[0419] The "at least one (e.g. several) other purification step" may be any other step suitable for protein purification known in the art. For example purification techniques for the recovery of recombinantly expressed albumin have been disclosed in: WO 92/04367, removal of matrix-derived dye; EP 464590, removal of yeast-derived colorants; EP 319067, alkaline precipitation and subsequent application of the albumin to a lipophilic phase; and WO 96/37515, U.S. Pat. No. 5,728,553 and WO 00/44772, which describe complete purification processes; all of which are incorporated herein by reference. Suitable methods include ammonium sulphate or ethanol precipitation, acid or solvent extraction, anion or cation exchange chromatography, phosphocellulose chromatography, hydrophobic interaction chromatography, affinity chromatography, hydroxyapatite chromatography, lectin chromatography, concentration, dilution, pH adjustment, diafiltration, ultrafiltration, high performance liquid chromatography ("HPLC"), reverse phase HPLC, conductivity adjustment and the like.
[0420] The polypeptide may be purified to a commercially or industrially acceptable level of purity. By commercially or industrially acceptable level of purity, we include the provision of the thio-albumin and/or thio-albumin-conjugate in which other material (for example, one or more (e.g. several) contaminants) are present at a level of less than 50%, 40%, 30%, 20%, 10%, 5%, 4%, 3%, 2%, 1%, 0.5%, 0.1%, 0.01%, 0.001%, 0.0001%, 0.00001%, or 0.000001% and, most preferably at a level of 0%.
[0421] A commercially or industrially acceptable level of purity may be obtained by a relatively crude purification method by which the protein product of choice is put into a form suitable for its intended purpose. A protein preparation that has been purified to a commercially or industrially acceptable level of purity may, in addition to the protein product of choice, also comprise, for example, cell culture components such as host cells or debris derived therefrom. Alternatively, high molecular weight components (such as host cells or debris derived therefrom) may or may not be removed (such as by filtration or centrifugation) to obtain a composition comprising the protein product of choice and, optionally, a functionally acceptable level of low molecular weight contaminants derived from the cell culture process.
[0422] The protein may or may not be purified to achieve a pharmaceutically acceptable level of purity. A protein has a pharmaceutically acceptable level of purity if it is essentially pyrogen free and can be used for its intended purpose and hence be administered in a pharmaceutically efficacious amount without causing medical effects not associated with the activity of the protein.
[0423] The thio-albumin and/or thio-albumin-conjugate may be provided at a concentration of at least 10.sup.-4 gL.sup.-1, 10.sup.-3 gL.sup.-1, 0.01 gL.sup.-1, 0.02 gL.sup.-1, 0.03 gL.sup.-1, 0.04 gL.sup.-1, 0.05 gL.sup.-1, 0.06 gL.sup.-1, 0.07 gL.sup.-1, 0.08 gL.sup.-1, 0.09 gL.sup.-1, 0.1 gL.sup.-1, 0.2 gL.sup.-1, 0.3 gL.sup.-1, 0.4 gL.sup.-1, 0.5 gL.sup.-1, 0.6 gL.sup.-1, 0.7 gL.sup.-1, 0.8 gL.sup.-1, 0.9 gL.sup.-1, 1 gL.sup.-1, 2 gL.sup.-1, 3 gL.sup.-1, 4 gL.sup.-1, 5 gL.sup.-1, 6 gL.sup.-1, 7 gL.sup.-1, 8 gL.sup.-1, 9 gL.sup.-1, 10 gL.sup.-1, 15 gL.sup.-1, 20 gL.sup.-1, 25 gL.sup.-1, 30 gL.sup.-1, 40 gL.sup.-1, 50 gL.sup.-1, 60 gL.sup.-1, 70 gL.sup.-1, 80 gL.sup.-1, 90 gL.sup.-1, 100 gL.sup.-1, 150 gL.sup.-1, 200 gL.sup.-1, 250 gL.sup.-1, 300 gL.sup.-1, 350 gL.sup.-1, 400 gL.sup.-1, 500 gL.sup.-1, 600 gL.sup.-1, 700 gL.sup.-1, 800 gL.sup.-1, 900 gL.sup.-1, 1000 gL.sup.-1.
[0424] A method of the present invention may or may not further comprise the step of formulating the purified protein product of choice with a carrier or diluent and optionally presenting the thus formulated protein in a unit dosage form.
[0425] Although it is possible for a therapeutically useful protein obtained by a process of the invention to be administered alone, it is preferable to present it as a pharmaceutical formulation, together with one or more (e.g. several) acceptable carriers or diluents. The carrier(s) or diluent(s) must be "acceptable" in the sense of being compatible with the desired protein. Typically, the carriers or diluents will be water or saline which will be sterile and pyrogen free. Alternatively, a method of the present invention may or may not further comprise the step of lyophilising the thus purified protein product of choice.
[0426] The thio-albumin may be formulated by strategies given in "Protein Formulation and Delivery", E. J. McNally (Ed.), published by Marcel Dekker Inc. New York 2000 and "Rational Design of Stable Protein Formulations--Theory and Practice"; J. F. Carpenter and M. C. Manning (Ed.) Pharmaceutical Biotechnology Vol 13. Kluwer Academic/Plenum Publishers, New York 2002, Yazdi and Murphy, (1994), Cancer Research 54, 6387-6394, Widera et al., (2003) Pharmaceutical Research 20, 1231-1238; Lee et al., (2005), Arch. Pharm. Res. 28, 722-729. Examples of formulation methods are as follows:
[0427] Method #1: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be stored at 4.degree. C., -20.degree. C. or -80.degree. C. in 0.01 M-0.1 M phosphate buffered saline (pH 7.0-8.0) containing 0.01 M-0.25 M NaCl.
[0428] Method #2: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be stored at 4.degree. C., -20.degree. C. or -80.degree. C. in 0.01 M-0.1 M phosphate buffered saline (pH 7.0-8.0) containing 0.01 M-0.25 M NaCl and containing 10-20 mg/L Polysorbate 80.
[0429] Method #3: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be stored at 4.degree. C., -20.degree. C. or -80.degree. C. in 0.01 M-0.25 M NaCl (pH 7.0-8.0).
[0430] Method #4: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be stored at 4.degree. C., -20.degree. C. or -80.degree. C. in 0.01 M-0.25 M NaCl (pH 7.0-8.0) containing 10-20 mg/L Polysorbate 80.
[0431] Freeze-Dried Formulations
[0432] Method #5: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be dialysed against water, freeze dried and stored at 4.degree. C., -20.degree. C. or -80.degree. C.
[0433] Method #6: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be dialysed against 0.01 M-0.25 M NaCl (pH 7.0-8.0), freeze dried and stored at 4.degree. C., -20.degree. C. or -80.degree. C.
Conjugation Methods
[0434] A tenth aspect of the invention provides a method of producing the conjugate of the seventh aspect of the invention, the method comprising linking a polypeptide of the first, second or third aspect of the invention, or produced by the method of the ninth aspect of the invention, to a bioactive compound through a conjugation-competent cysteine residue of, the polypeptide. The linking may be carried out using a linker.
[0435] The albumin mutein (thio-albumin) of the invention can be covalently linked to one or more (e.g. several) conjugation partners such as bioactive compounds by methods known in the art (for example those provided by Pierce, Thermo Fisher Scientific, Rockford, Ill., USA; https://tools.lifetechnologies.com/content/sfs/brochures/1602163-Crosslin- king-Reagents-Handbook.pdf). These include, but are not limited to incorporating or engineering a thiol reactive group into or onto the conjugation partner, for example by incorporating or engineering another free thiol present on the conjugation partner; or by incorporating or engineering a pyridyl disulphide group on the conjugation partner; or by incorporating or engineering an haloacetyl group on the bioactive compound or by incorporating or engineering a maleimide group on the conjugation partner, or by incorporating or engineering a thiosulfonate group on the conjugation partner, or by incorporating or engineering vinylsulfone group on the conjugation partner. For example, but not limited to, N-ethylmaleimide (NEM, Pierce), 2-amino-2'-aminoethanethiolsulfonate (Pierce), N-beta-maleimidoprpionic acid (BMPA Pierce), methyl methane thiosulfonate (MMTS, Pierce), fluorescein-5-maleimide (Pierce), 5-iodoacetamido-fluorescein (5-IAF, Pierce) or N-[6-7-amino-4-methylcoumarin-3-acetamido) hexyl]-3'-[2'-pyridyldithio] propionamide (AMCA-HPDP, Pierce).
[0436] If the conjugation partner contains at least one (e.g. several) thiol group, then the conjugation partner may be cross-linked to the albumin mutein of the invention by methods known to the art such as, but not limited to, oxidation or by the use of cross-linking reagents such as, but not limited to, 1,4-Bis-maleimidibutane (BMB, Pierce); 1,4-Bis-maleimidyl-2,3-dihydroxybutane (BMDB, Pierce); Bis-maleimidohexane (BMH, Pierce), Bis-maleimidoethane (BMOE, Pierce); 1,8-Bis-Maleimidotriethyleneglycol (BM[PEO]3 Pierce); 1,11-Bis-Maleimidotetraethyleneglycol (BM[PEO]4 Pierce); 1,4-DI-[3'-(2'-pyridyldithio)-propionamido]butane (DPDPB, Pierce); dithio-bis-maleimidoethane (DTME Pierce); 1,6-Hexane-bis-vinylsulfone (HBVS, Pierce) and Tris[2-maleimimidoethyl]amine (TMEA, Pierce).
[0437] If the conjugation partner does not contain a thiol reactive group then it may be modified to incorporate one or more (e.g. several) such groups by either chemical modification or genetic engineering by methods know to the art (Chapman, A. P. (2002) Adv. Drug Deliv. Rev., 54 531-545: Humphreys, D. P. et al. Protein Engineering, Design & Selection vol. 20 no. 5 pp. 227-234, 2007). While these two references describe methodologies to cross-link PEG to an engineered free thiol within an antibody or antibody fragment, the techniques may be used to cross-link a conjugation partner to an engineered free thiol within the albumin mutein of the invention. Alternatively the Drug Affinity Complex (DAC.TM.) technology developed by ConjuChem Inc. (Montreal, Quebec, Canada, H2X 3Y8) may be used, e.g. as described in WO 200069902. There are three parts of each DAC.TM. construct: 1) the drug component (the portion responsible for biologic activity); 2) a linker attached to the drug component, and 3) a reactive chemistry group at the opposite end of the linker, usually a soft electrophile selective for thiols; a maleimide is the most useful embodiment. Other applicable conjugation methods are described in WO 2007/071068 incorporated herein by reference.
[0438] If the conjugation partner does not contain a thiol reactive group but does contain one or more (e.g. several) amino groups then it may be modified to incorporate one or more (e.g. several) thiol reactive groups by chemical modification by methods known to the art such as the use of cross-linking reagents such as, but not limited to, N-5-azido-2-nitrobenzoyloxysuccinimide (AMAS, Pierce), N-[beta-maleimidopropyloxy] succinimide ester (BMPS, Pierce), N-eta-maleimidocaproic acid (EMCA, Pierce), N-[eta-maleimidocaproyloxy]succinimide ester (EMCS, Pierce), N-[eta-maleimidocaproyloxy]sulfosuccinimide ester (sulfa-EMCS, Pierce), N-[gamma-maleimidobutyryloxy]succinimide ester (GMBS, Pierce), N-[gamma-maleimidobutyryloxy]sulfosuccinimide ester (sulfo-GMBS, Pierce), N-kappa-maleimidoundecanoic acid (KMUA, Pierce), N-[kappa maleimidoundecanoyloxy]sulfosuccinimide ester (sulfo-KMUS, Pierce), m-maleimidobenzoyl-N-hydroxysuccinimide (MBS, Pierce), m-maleimidobenzoyl-N-hydroxysulfosuccinimide ester (sulfo-MBS, Pierce), N-succinimidyl S-acetylthio-acetate (SATA, Pierce), N-succinimidyl S-acetylthiopropionate (SATP, Pierce), succinimidyl 3-[bromoacetamido]propionate (SBAP, Pierce), N-succinimidyl iodoacetate (SIA, Pierce), N-succinimidyl[4-iodoacetyl]aminobenzoate (SIAB, Pierce), sulfosuccinimidyl[4-iodoacetyl]aminobenzoate (sulfo-SIAB, Pierce), succinimidyl [4[N-maleimidomethyl]cyclohexane-1-carboxylate (SMCC, Pierce), sulfosuccinimidyl [4-[N-maleimidomethyl]cyclohexane-1-carboxylate (sulfo-SMCC, Pierce), succinimidyl-[4-[N-maleimidomethyl]cyclohexane-1-carboxy-[6-amidocaproate (LC-SM CC, Pierce), 4-succinimidyloxycarbonyl-methyl-alpha[2-pyridyld ithio]toluene (SMPT, Pierce), sulfosuccinimidyl6-[alpha-methyl-alpha-(2-pyridyldithio)toluamido]hexanoa- te (sulfo-LC-SMPT, Pierce), succinimidyl 4-[p-maleimidophenyl]-butyrate (SMPB, Pierce), sulfosuccinimidyl 4-[p-maleimidophenyl]-butyrate (sulfo-SMPB, Pierce), succinimidyl-6-[(beta-maleimidopropionamido)hexanoate] (SMPH, Pierce), N-succinimidyl 3-[2-pyridyld ithio]propionate (SP DP, Pierce), succinimidyl [3-(2-pyridyldithio)propionamido]hexanoate (LC-SPDP, Pierce), sulfosuccinimidyl [3'-(2-pyridyldithio)propionamido]hexanoate (sulfo-LC-SPDP, Pierce) and N-succinimidyl-[4-vinylsulfonyl]benzoate (SVSB Pierce). It may be advantageous to block certain amine residue as described by Kavimandan et al., (2006), Bioconjugate Chem. 17, 1376-1384.
[0439] Suitable linkers include bromomaleimide linkers such as monobromomaleimide linkers. Monobromomaleimides are next generation maleimides for the construction of stable conjugates, as described in Smith et al Organic & Biomolecular Chemistry, (2015), 13, pages 7946-7949. Preferred monobromomaleimide linkers include those described in WO 2011/018611 (incorporated herein by reference).
[0440] If the conjugation partner does not contain a thiol reactive group but does contain one or more (e.g. several) carbonyl (oxidised carbohydrate) groups then it can be modified to incorporate one or more (e.g. several) thiol reactive groups by chemical modification by methods known to the art such as the use of cross-linking reagents such as, but not limited to, (N-.beta.-maleimidopropionic acid hydrazide (BMPH, Pierce) N-[eta-maleimidocaproic acid]hydrazide (EMCH, Pierce), 4-[N-maleimidomethyl]cyclohexane-1carboxylhydrazide.HCl.1/2 dioxane (MMCCH, Pierce), 3-maleimidophenyl boronic acid (MPBH, Pierce), N-[kappa-maleimidoundecanoic acid]hydrazide (KMUH, Pierce) and 3-[2-pyridyldithio]propionyl hydrazide (PDPH, Pierce).
[0441] If the conjugation partner does not contain a thiol reactive group but does contain one or more (e.g. several) hydroxyl groups then it may be modified to incorporate one or more (e.g. several) thiol reactive groups by chemical modification by methods known to the art such as the use of cross-linking reagents such as, but not limited to, N-[p-maleimidophenyl]isocyanate (PMPI, Pierce).
Associates
[0442] An eleventh aspect of the invention provides an associate comprising the conjugate of the seventh aspect of the invention and a bioactive, therapeutic, prophylactic, diagnostic, imaging or other beneficial moiety.
[0443] The conjugates may further be used in the form of "associates". In this connection the term "associate" is intended to mean a compound comprising a conjugate of a variant of albumin or a fragment thereof and another compound bound or associated to the conjugate by non-covalent binding. As an example of such an associate can be mentioned an associate consisting of a variant albumin conjugate and a lipid associated to albumin by a hydrophobic interaction. Such associates are known in the art and they may be prepared using well known techniques. As an example of a preferred associate according to the invention can be mentioned, an associate comprising a variant albumin conjugate and a taxane, a taxol or taxol derivative (e.g. paclitaxel). Further examples of associates comprise a bioactive, therapeutic, prophylactic (including vaccine), diagnostic, imaging or other beneficial moiety.
[0444] Methods for the preparation of associates are well-known to the skilled person, for example, formulation (by association) of HSA with lipo-compounds is described in Hussain, R. and Siligardi, G. (2006), International Journal of Peptide Research and Therapeutics, Vol. 12, NO: 3, pp. 311-315.
Nanoparticle, Microparticle or Liposome
[0445] A twelfth aspect of the invention provides a nanoparticle, a microparticle or a liposome comprising the polypeptide or the first, second or third aspect of the invention, the conjugate of the seventh aspect of the invention or the associate of the eleventh aspect of the invention.
[0446] Albumins and albumin particles are important for carrying and delivering drugs and prodrugs to their sites of action (Kratz (2008), Journal of Controlled Release, 132 (3), p. 171-183). Fusion and particle technologies offer improved dosing regimens due to improved pharmacokinetic properties, such as plasma half-life extension, and may improve bioavailability and protect the fused bioactive molecule from inactivation.
[0447] Techniques for incorporation of a molecule into nano- or microparticles are known in the art. Preferred methods for preparing nano- or microparticles that may be applied to the variant albumin conjugate or associate thereof according to the invention are disclosed in WO 2004/071536 or WO 2008/007146 or Oner & Groves (Pharmaceutical Research, Vol 10(9), 1993, pages 1387 to 1388) which are incorporated herein by reference. Preferably the average diameter of a nano-particle is from 5 to 1000 nm, more preferably from 5, 10, 20, 30, 40, 50, 80, 100, 130, 150, 200, 300, 400, 500, 600, 700, 800, 900, or 999 to 5, 10, 20, 30, 40, 50, 80, 100, 130, 150, 200, 300, 400, 500, 600, 700, 800, 900, or 1000 nm. An advantage of a microparticle less than 200 nm diameter, and more particularly less than 130 nm, is that is amenable to sterilization by filtration through a 0.2 .mu.m (micron) filter. Preferably, the average diameter of a microparticle is from 1000 nm (1 .mu.m (micron)) to 100 .mu.m (micron), more preferably from 1, 2, 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100 to 1, 2, 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100 .mu.m (micron).
[0448] The thio-albumin of the invention (and/or its conjugated form) may be used to produce nanoparticles and/or be entrapped within a nanoparticle or liposome.
[0449] The thio-albumin of the invention may be used with and/or in and/or as a nanoparticle and/or liposome. A problem of current conjugation strategies is maintaining both the pharmacological and immunological activity of the conjugation partner, such as a bioactive-targeting ligand conjugate. There is likely to be a maximum number of protein targeting ligand or bioactive moieties (conjugation partners) possible for conjugation to a protein and if this number is exceeded the targeting ligand does not retain its biological activity. Preferably the biological activity of the conjugation partner is not reduced by conjugation to an albumin of the invention.
[0450] Liposomes and nanoparticles may be used to entrap bioactive compounds. They provide a mechanism for enhanced delivery of drugs such as bioactive compounds, or uptake by target cells and/or a reduction in the toxicity of the free bioactive to non-target organs which may result in an increased therapeutic index and/or reduced side effects. In addition, many solvent-based formulations required for the delivery of some bioactive compounds (e.g. taxanes) are associated with toxicity which limits the maximum dose which can be given to a patient. Liposome and nanoparticle delivery may also be advantageous for such bioactive compounds, since they would allow larger amounts of the bioactive compound to be delivered whilst avoiding some of the toxicities of solvent-based formulations (Hawkins et al (2008), Advanced Drug Delivery Reviews, 60, 8, p876-885).
[0451] Methods for attaching targeting ligands to liposomes and nanoparticles are known in the art (reviewed in Nobs et al (2004), Journal of Pharmaceutical Sciences Vol 93 p1980-1992) and may be used in accordance with the invention. Attachment methods may be non-covalent or covalent. Covalent reactions appear to be favourable, because covalent linkage is more stable than noncovalent methods. Lipids for the covalent or non-covalent attachment of proteins, peptides, or drugs to the liposome surface are available commercially (for example Avanti Polar Lipids Inc Alabaster, Ala., USA). There are 3 major classes of functionality: conjugation through disulphide or thioether formation, amide bond formation, or biotin/streptavidin binding, any of these may be used in the invention.
[0452] A number of methods relying on covalent coupling ligands to the surface of liposomes via thioether bonds have been described, most commonly utilizing the highly efficient reaction of maleimide with thiol groups. Functionalized lipid anchors commonly added to liposomes, and which may be used in or with the invention, include, but are not limited to those containing maleimide such as N-[4-(p-maleimidophenyl) butyramide]-PE (N-MPB]-PE) or N-[4-(p-maleimidomethyl) cyclohexane-carboxamide) (MCC-PE) which allow convenient covalent coupling of the targeting moiety via a stable thioether bond (Martin & Papahadjopoulos (1982), J. Biol. Chem. 257, 286-288).
[0453] Method #7: Following purification the free thiol containing albumin mutein of the invention or the conjugate can be formulated into nanoparticles prepared according to known procedures for preparing nanoparticles, such as procedures disclosed in WO 2004/071536 A1 and WO 2008/007146 A1, both incorporated herein by reference.
[0454] Similarly materials for the formation of nanoparticles, including but are not limited to poly(lactic acid) (PLA), poly(lactic-co-glycolic acid) (PLGA), and COOH-PLA are commercially available and may be functionalized with maleimide or other known chemistries according to known literature for nanoparticle formation. Any of these may be used in or with the invention.
[0455] Another convenient way for covalent coupling of ligands to liposomes involves conjugation of two thiols to form a disulphide; however under the reductive conditions in serum more stable conjugation chemistries involving one free thiol group may be preferred. Chemistries such as (PDP-PE) allow covalent coupling via a disulphide bond. Modification of the ligand to introduce a free thiol group or a functionalized linker may be used. An advantage of the thio-albumin of the invention is that no ligand modification is required. However, ligand modification may optionally be used in addition to the invention.
[0456] Frequently thiol groups are not present in proteins, or are not present in sufficient amounts or at the desired location. Thus, most cases of covalent coupling of one of more ligands to a liposome via thioether or disulphide bonds requires the use of heterobifunctional cross linking agents (described herein with reference to conjugation). Some heterobifunctional cross linking agents (such as SPDP and SATA) require a de-protection step. The thio-albumin of the invention overcomes the requirement for this additional processing.
[0457] Alternatively thio-albumin could be conjugated to liposomes or nanoparticles by other chemistries, known to the art. For example, thio-albumin could be attached by an amide bond using a functionalised lipid anchor with either amine or carboxyl functional groups (examples include DSPE-PEG-COOH) which reacts with the primary amine of the ligand. Direct cross linking between primary amines and the surface of liposomes may also be used. The one or more (e.g. several) free thiol groups of thio-albumin would then be available for conjugation to another conjugation partner.
[0458] Following conjugation, a conjugation partner (e.g. bioactive molecule) may show a reduction in its activity (e.g. bioactivity). Thio-albumin described in this invention may overcome this problem by providing a conjugate, nanoparticle and/or liposome in which the conjugation partner is located and/or orientated with respect to a thio-albumin such that the conjugation partner retains at least 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100% of its unconjugated activity.
[0459] Nanoparticles may be used, for example, in angiogenic applications, anti-angiogenic applications and to coat a medical device such as a stent. Nanoparticles are effective at targeting, for example to non tight-junctions, and therefore can be useful for targeting tumours such as cancerous tumours. Nanoparticles can also be useful to target antigen in order to provoke an immune response since nanoparticles are particularly susceptible to engulfment and presentation by phagocytes. The invention provides nanoparticles consisting only of thio-albumin according to the invention which may or may not be conjugated to a moiety (conjugation partner). The invention also provides nanoparticles comprising thio-albumin according to the invention, which may or may not be conjugated to a moiety, and one or more (e.g. several) other constituents of a nanoparticle which may or may not be albumin related. In a preferred embodiment, a thio-albumin according to the invention comprises at least two conjugation competent cysteine residues located on the surface of the polypeptide. Such a thio-albumin may be used for the preparation of nanoparticles in which one or more (e.g. several) conjugation competent cysteine residues may be used in the formation of a nanoparticle and one or more (e.g. several) conjugation competent residues is used for conjugation to a conjugation partner, for example to a bioactive molecule.
Compositions
[0460] A thirteenth aspect of the invention provides a composition comprising a polypeptide, fusion polypeptide, conjugate, associate, nanoparticle, microparticle or liposome according to the invention and at least one (e.g. several) pharmaceutically acceptable carrier and/or diluent.
[0461] Various formulations are described herein in relation to the corresponding products.
[0462] A related aspect of the invention provides a method for making a pharmaceutical ingredient and/or a pharmaceutical product comprising making a thio-albumin according to the present invention, optionally conjugating a further molecule to the thio-albumin, optionally formulating the resultant conjugate with a pharmaceutically acceptable diluent and/or carrier and optionally preparing the product in unit dosage form.
Medical Uses
[0463] A fourteenth aspect of the invention provides use of a polypeptide, fusion polypeptide, conjugate according to the invention and/or produced by a method according to the invention, or an associate, nanoparticle, microparticle or liposome for treatment of disease, treatment of illness and/or diagnosis.
[0464] Various medical uses are described herein in relation to the corresponding products.
[0465] In addition, in some embodiments, the thio-albumin or conjugate has a binding affinity to FcRn and/or plasma half-life that is altered compared to the parent or reference albumin or conjugate. This has the advantage that the binding affinity to FcRn and/or plasma half-life of conjugates, associates, nanoparticle, microparticle or liposome according to the invention can be selected in accordance with the particular therapeutic purpose. An increased half-life could have the benefit that the administration would be needed less frequently or at a reduced dose (and consequently with fewer side effects) compared to the situation where the reference molecule or composition was used. Alternatively, a shorter plasma half-life than the reference molecule or composition would have the benefit that the administration can be carried out at a higher dose compared to the situation where the reference molecule or composition was used with the benefit that the administered compound clears from the recipient more quickly than if the reference molecule or composition was used.
[0466] For example for a conjugate, associate or fusion polypeptide used for imaging purposes in animals or humans, where the imaging moiety has a very short half-life and a conjugate or a fusion polypeptide comprising HSA has a plasma half-life that is far longer than needed for the imaging purposes it would be advantageous to use a variant albumin or fragment thereof of the invention having a shorter plasma half-life than the parent or reference albumin or fragment thereof, to provide conjugates or fusion polypeptides having a plasma half-life that is sufficiently long for the imaging purpose but sufficiently short to be cleared form the body of the particular patient on which it is applied.
[0467] In another example for a conjugate, an associate or fusion polypeptide comprising a therapeutic compound effective to treat or alleviate a particular condition in a patient in need for such a treatment it would be advantageous to use the variant albumin or fragment thereof having a longer plasma half-life than the parent or reference albumin or fragment thereof, to provide associates or conjugates or fusion polypeptides having longer plasma half-lives which would have the benefit that the administration of the associate or conjugate or fusion polypeptide of the invention would be needed less frequently or at reduced dose with less side effects compared to the situation where the parent or reference albumin or associates thereof or fragment thereof was used. For example, the invention provides a method of treating a proliferative disease in an individual, comprising administering the individual an effective amount of an associate according to the invention in which the associate comprises a taxane, a taxol or taxol derivative (e.g. paclitaxel).
Use to Increase Half-Life
[0468] A fifteenth aspect of the invention provides for use of a polypeptide as defined in any previous aspect of the invention to increase the half-life of a molecule such as a bioactive agent, an imaging agent, a diagnostic agent, a contrast agent or a therapeutic compound such as a chemotherapeutic drug or radiopharmaceutical. Preferably, the half-life is increased by at least 10, 20, 30, 40, 50, 60, 70, 80, 90 or at least 100% relative to the half-life of the molecule alone. Preferably, the half-life is increased by at least 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22 hours or by at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13 or at least 14 days relative to the half-life of the molecule alone.
[0469] For example, the half-life of a molecule may be increased by conjugating it to the polypeptide as defined in any previous aspect of the invention for example via a conjugatable cysteine residue; by genetically fusing the molecule to the polypeptide, by associating the molecule with the polypeptide and/or by incorporating it into a particle according to any previous aspect of the invention.
Embodiments of the Invention
[0470] The invention is further described with reference to the following numbered paragraphs: 1. A conjugation-competent polypeptide comprising an amino acid sequence which is at least 70% identical to human albumin, particularly residues 1 to 585 of the mature human albumin polypeptide sequence of SEQ ID NO. 2, or a fragment thereof;
[0471] wherein at least one (e.g. several) position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, 0268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue; and
[0472] preferably wherein the conjugation-competent polypeptide has a tendency to exist as a monomer in solution which is at least 70% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution.
2. The conjugation-competent polypeptide of Paragraph 1, wherein the polypeptide comprises one or more (e.g. several) of:
[0473] substitution of an amino acid, other than cysteine, with a cysteine at a position corresponding to a position equivalent to any of residues K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2; and/or
[0474] insertion of a cysteine at a position adjacent the N- or C-side of an amino acid corresponding to a position equivalent to any of residues K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2.
3. The conjugation-competent polypeptide of Paragraph 1 or 2 wherein two, three, four, five or more (e.g. several) positions equivalent to positions selected from K93, E294, A226, E230, I271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, 0340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2 comprise a conjugation-competent cysteine residue. 4. The conjugation-competent polypeptide of any preceding Paragraph, wherein the polypeptide has a tendency to exist as a monomer in solution which is at least 75%, at least 80%, at least 85%, at least 90%, at least 95% or at least 100% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution. 5. The conjugation-competent polypeptide of any preceding Paragraph wherein the tendency of the polypeptide to exist as monomer in solution is measured following storage for at least 7 weeks at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 8 weeks at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 3 months at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 4 months at a temperature from 2 to 8.degree. C. such as 5.degree. C., at least 6 months storage at a temperature from 2 to 8.degree. C. such as 5.degree. C., or at least 3 months storage at a temperature of about 40.degree. C. 6. The conjugation-competent polypeptide of Paragraph 5 wherein the tendency of the polypeptide to exist as monomer in solution is measured following storage for at least 3 months at a temperature from 2 to 8.degree. C., such as 5.degree. C. 7. The conjugation-competent polypeptide of paragraph 5 or 6, prior to storage, wherein the polypeptide is purified using triazine (such as AlbuPure.RTM.) chromatography matrix or DE-FF chromatography matrix prior to storage. 8. The conjugation-competent polypeptide of any of paragraphs 5, 6, or 7 wherein, prior to storage, the polypeptide is purified using triazine (such as AlbuPure.RTM.) chromatography matrix followed by DE-FF chromatography matrix. 9. The conjugation-competent polypeptide of any of paragraphs 5 to 8 wherein, prior to storage, the polypeptide is purified using triazine (such as AlbuPure.RTM.) chromatography matrix followed by DE-FF chromatography matrix followed by size exclusion (e.g size exclusion limit (Mr) of about 5.times.10.sup.3 to 2.5.times.10.sup.5 such as Sephacryl 5-200 HR) chromatography. 10. The conjugation-competent polypeptide of any of Paragraphs 5 to 9 wherein the storage uses a polypeptide concentration of from 0.5 to 50 mg/mL. 11. The conjugation-competent polypeptide of any of Paragraphs 5 to 10 wherein the storage uses a polypeptide concentration of about 5 mg/mL. 12. The conjugation-competent polypeptide of any of Paragraphs 5 to 11 wherein the storage is at a pH between about 6.0 and about 7.5. 13. The conjugation-competent polypeptide of any of Paragraphs 5 to 12 wherein the storage is at a pH about 7. 14. The conjugation-competent polypeptide of any of Paragraphs 5 to 13 wherein the storage uses a buffer comprising 50 mM ammonium acetate, 10 mM sodium octanoate, pH 7.0, preferably at a polypeptide concentration of from about 0.2 to about 2.5 mg/mL. 15. The conjugation-competent polypeptide of any of Paragraphs 5 to 14 wherein the storage, uses a buffer comprising 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5, preferably at a polypeptide concentration of from about 5 to about 50 mg/mL. 16. The conjugation-competent polypeptide of any preceding Paragraph, wherein at least one (e.g. several) position equivalent to a position selected from K93, E294, A226, E230, I271, E358, L24, F49, V54, D56, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, L284, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue; and wherein the tendency to exist as monomer in solution is at least 75% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution. 17. The conjugation-competent polypeptide of any preceding Paragraph wherein the amino acid sequence is at least 95% identical to human albumin, particularly residues 1 to 585 of the mature human albumin polypeptide sequence of SEQ ID NO. 2, or a fragment thereof and the conjugation-competent polypeptide has a tendency to exist as a monomer in solution which is at least 80% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution. 18. The conjugation-competent polypeptide of any preceding Paragraph, wherein at a position equivalent to position 34 of SEQ ID NO. 2 there is a conjugation-competent cysteine. 19. The conjugation-competent polypeptide of any of Paragraphs 1 to 18, wherein at a position equivalent to position 34 of SEQ ID NO. 2 there is not a conjugation-competent cysteine. 20. The conjugation-competent polypeptide of any preceding Paragraph in which the polypeptide comprises two or more (several) conjugation-competent cysteine residues wherein, when the polypeptide is folded, there is a distance of at least 5 .ANG. between at least one pair of the conjugation-competent cysteine residues. 21. The conjugation-competent polypeptide of any preceding Paragraph, wherein the polypeptide comprises substitution of an amino acid, other than cysteine, with a cysteine at one or both positions corresponding to a position equivalent to residues K93 or E294 of SEQ ID NO. 2. 22. The conjugation-competent polypeptide of any preceding Paragraph which is capable of forming a conjugate with maleimide-polyethylenglycol2-biotin, at a conjugation efficiency of at least 90%, preferably at least 95%, suitably wherein the conjugate is at 90%, preferably at least 95% stable upon controlled hydrolysis. 23. The conjugation-competent polypeptide of Paragraph 22 wherein the capability of forming a conjugate with maleimide-polyethylenglycol2-biotin is determined by incubating at ambient temperature overnight in phosphate buffered saline buffer pH 7.4. 24. The conjugation-competent polypeptide of Paragraph 22 or 23 wherein stability is determined by incubating at pH 9.0 and 37.degree. C. for at least 18 hours, preferably 24 hours, in a buffered salts solution, such as phosphate buffered saline. 25. A conjugation-competent polypeptide comprising an amino acid sequence which is at least 70% identical to human albumin (SEQ ID NO. 2), or a fragment thereof;
[0475] wherein at least one (e.g. several) position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and 1271, of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue; and
[0476] comprising at least one (e.g. several) further conjugation-competent cysteine, or at least one (e.g. several) modification that alters the binding affinity of the polypeptide for FcRn, or alters the plasma half-life of the polypeptide.
26. The conjugation-competent polypeptide of Paragraph 25 wherein the at least one (e.g. several) further modification comprises at least one (e.g. several) further conjugation-competent cysteine as defined in any one of Paragraphs 1, 2, 3 or 21. 27. The conjugation-competent polypeptide of any preceding Paragraph wherein at least one (e.g. several) position equivalent to a position selected from D1, A2, H3, S5, A55, S58, C75, T76, T79, E82, T83, E86, C91, D121, V122, C124, T125, D129, C169, C177, A229, T236, E266, D269, S270, S273, S304, K313, D314, 0316, N318, A320, C361, A364, C369, A371, N386, Q390, Q397, S435, T478, T496, A504, E505, T506, T508, D549, C558, D562, C567, A581, L585 and A578 of SEQ ID NO. 2 comprises a conjugation-competent cysteine. 28. The conjugation-competent polypeptide of any preceding Paragraph in which the polypeptide comprises one or more (e.g. several) of:
[0477] substitution of an amino acid, other than cysteine, with a cysteine at a position corresponding to a position equivalent to any of residues D1, A2, H3, S5, A55, S58, C75, T76, T79, E82, T83, E86, C91, D121, V122, C124, T125, D129, C169, C177, A229, T236, E266, D269, S270, S273, S304, K313, D314, 0316, N318, A320, C361, A364, C369, A371, N386, Q390, Q397, 5435, T478, T496, A504, E505, T506, T508, D549, C558, D562, C567, A581, L585 and A578 of SEQ ID NO. 2; and/or
[0478] insertion of a cysteine at a position adjacent the N- or C-side of an amino acid corresponding to a position equivalent to any of residues D1, A2, H3, S5, A55, S58, C75, T76, T79, E82, T83, E86, C91, D121, V122, C124, T125, D129, C169, C177, A229, T236, E266, D269, S270, S273, S304, K313, D314, C316, N318, A320, C361, A364, C369, A371, N386, Q390, Q397, 5435, T478, T496, A504, E505, T506, T508, D549, C558, D562, C567, A581, L585 and A578 of SEQ ID NO. 2; and/or
[0479] deletion or substitution of a cysteine at a position corresponding to any of C360, C316, C75, C168, C558, C361, C91, C124, C169 and 0567 of SEQ ID NO. 2 so as to generate a conjugation competent cysteine at any of C369, 0361, C91, C177, C567, C316, C75, C169, C124 and C558; and/or
[0480] addition of a cysteine to the N-side of the N-terminal residue of an albumin sequence or to the C-side of the C-terminal residue of an albumin sequence.
29. The conjugation-competent polypeptide of any preceding Paragraph in which the polypeptide comprises conjugation-competent cysteines located at: (a) A2+L585, (b) A2+A364+D562+L585C, (c) A2 and adjacent the C-side of the C-terminus of the albumin (d) T79+A364; (e) A364+D1; (f) T79+D562+A364; (g) D562+A364+D1; (h) T79+D562+A364+A504; (i) T79+D562+A364+L585; (j) T79+D562+A364+D1; (k) T79+D562+A364+L585+D1; (I) E86+D562+A364+A504+A2; (m) S270+A581; (n) 5270+D129; (o) 5270+A581+E82; (p) S270+A581+D129; (q) 5270+A581+E82+D129; (r) S270+A581+E82+D129+Q397; (s) C369+C177; (t) A364+A581; (u) T79+A364+A581; (v) A364+A581+D129; (w) A364+C177; (x) D562+C369; (y) D129+C369; (z) A581+C369; or (aa) D562+D129+C369. 30. The conjugation-competent polypeptide of any preceding Paragraph which comprises or consists of albumin domain III or a variant thereof and at least one (e.g. several) additional albumin domain or fragment thereof, such as a second albumin domain III or a variant thereof. 31. The conjugation-competent polypeptide of any preceding Paragraph which comprises or consists of at least one (e.g. several) albumin domain III or variant or fragment thereof wherein at least one (e.g. several) albumin domain III comprises one or more (e.g. several) substitutions in positions corresponding to the positions in SEQ ID NO. 2 selected among: 573, 500, 550, 417, 440, 464, 490, 492, 493, 494, 495, 496, 499, 501, 503, 504, 505, 506, 510, 535, 536, 537, 538, 540, 541, 542, 574, 575, 577, 578, 579, 580, 581, 582 and 584. 32. The conjugation-competent polypeptide of Paragraph 31, wherein the one or more (e.g. several) substitutions in positions corresponding to the positions in SEQ ID NO. 2 is selected among: K573Y, W, P, H, F, V, I, T, N, 5, G, M, C, A, E, Q, R, L, D, K500E, G, D, A, S, C, P, H, F, N, W, T, M, Y, V, Q, L, I, R, Q417A, H440A, H464Q, E492G, D494N,Q,A, E495Q,A, T496A, D494E+Q417H, D494N+T496A, E492G+V493P, P499A, E501A,Q, N503H,K, H510Q, H535Q, K536A, P537A, K538A, K541G,D, D550E,N, E492G+K573P,A, or E492G/N503H/K573P. 33. The conjugation-competent polypeptide of any preceding Paragraph wherein the polypeptide comprises alterations at two or more (e.g. several) positions selected from positions corresponding to positions (a) 492 and 580; (b) 492 and 574; (c) 492 and 550; (d) 550 and 573; (e) 550 and 574; (f) 550 and 580 in SEQ ID NO. 2. 34. The conjugation-competent polypeptide of any preceding Paragraph comprising: (i) an N-terminal region comprising a first albumin which is a human albumin variant, in which the N-terminal of the first albumin comprises all amino acids of the human albumin variant except the C-terminal 2 to 30 amino acids; and (ii) a C-terminal region of a second albumin, which is selected from macaque albumin, mouse albumin, rabbit albumin, sheep albumin, human albumin, goat albumin, chimpanzee albumin, hamster albumin, guinea pig albumin, rat albumin, cow albumin, horse albumin, donkey albumin, dog albumin, chicken albumin, or pig albumin, or a variant thereof, in which the C-terminal of the second albumin or albumin variant comprises the C-terminal 2 to 30 amino acids of the second albumin or albumin variant; wherein the polypeptide has (i) an altered plasma half-life compared with the human albumin variant and/or (ii) an altered binding affinity to FcRn compared with the human albumin variant. 35. The conjugation-competent polypeptide of any preceding Paragraph comprising one or more (e.g. several) alterations in Domain I of the mature human albumin polypeptide sequence of SEQ ID NO. 2; and one or more (e.g. several) alterations in Domain III of the mature human albumin polypeptide sequence of SEQ ID NO. 2, wherein the one or more (e.g. several) alterations cause the polypeptide to have an altered binding affinity to FcRn. 36. The conjugation-competent polypeptide of Paragraph 35 wherein the alteration(s) in Domain I are selected from positions corresponding to any of positions 78 to 120 of SEQ ID NO. 2, such as any of positions 78 to 88 and/or from any of 105 to 120; and the alteration(s) in Domain III are selected from positions corresponding to any of positions 425, 505, 510, 512, 524, 527, 531, 534, 569, 573, or 575 of SEQ ID NO. 2. 37. The conjugation-competent polypeptide of Paragraph 36 wherein the alteration at the position corresponding to positions is selected among 78 to 120 or 425, 505, 510, 512, 524, 527, 531, 534, 569, 573, and/or 575 of SEQ ID NO. 2 is a substitution; and the alteration is optionally a substitution selected from (i) 83N, K or S; (ii) 111D, G, H, R, Q or E; or (iii) 573P, Y, W, H, F, T, I or V. 38. The conjugation-competent polypeptide of any preceding Paragraph comprising one or more (e.g. several) alterations in Domain II of the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions 349, 342, 381, 345, 384, 198, 206, 340, 341, 343, 344, 352, 382, 348, and/or 383 in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn. 39. The conjugation-competent polypeptide of Paragraph 38 wherein the alteration at the position corresponding to position 349, 342, 381, 345, 384, 198, 206, 340, 341, 343, 344, 352, 382, 348, and/or 383 is a substitution; and the alteration is optionally a substitution selected from (i) 349F, W, Y, H, P, K or Q, preferably F; (ii) 342Y, W, F, H, T, N, Q, A, C, I, L, P, V, preferably Y; (iii) 381G or A, preferably G; or (iv) 345E, H, I or Q. 40. The conjugation-competent polypeptide of any preceding Paragraph comprising one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions V418, T420, V424, E505, V547, K573 in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn. 41. The conjugation-competent polypeptide of any preceding Paragraph comprising one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions V381, preferably V381N or Q; E383, preferably E383A, G, I, L, or V; N391, preferably N391A, G, I, L or V; Y401 preferably Y401D or E; K402, preferably K402A, G, I, L, or V; L407, preferably L407F, N, Q, W, or Y; Y411, preferably Y411Q, or N; K413, preferably K4130, S, or T; K414, preferably K414S or T; V415C, preferably V415C, S, or T; Q416, preferably Q416H or P; V424, preferably V424A, G, I, L, N, or Q; V426D, preferably V426D, E, H, or P; G434, preferably G434C, S, or T; E442, preferably E442K or R; R445, preferably R445F, W or Y; P447, preferably P447S or T; E450, preferably E450D or E; S454, preferably S454C, M or T; V455, preferably V455N or Q; V456, preferably V456N or Q; L457, preferably L457F, W or Y; Q459, preferably Q459K or R; L463, preferably L463N or Q; E495, preferably E495D; T506, preferably T506F, W or Y; T508, preferably T508K, R, or S; F509, preferably F5090, I, L, M, V, W or Y; A511, preferably A511F, W, or Y; D512, preferably D512F, W or Y; T515, preferably T515C, H, N, P, Q or S; L516, preferably L516F, S, T, W or Y; S517, preferably S517C, F, M, T, W or Y; K519, preferably K519A, G, I, L, or V; R521, preferably R521F, W or Y; 1523, preferably I523A, D, E, F, G, K, L, N, Q, R, V, W or Y; K524, preferably K524A, G, I, L or V; K525, preferably K525A, G, 1, L or V; Q526, preferably Q526C, M, S, T or Y; T527, preferably T527F, W or Y; E531, preferably E531A, G, I, L or V; H535, preferably H535D, E or P; K538, preferably K538F, W or Y; A539, preferably A5391, L or V; K541, preferably, K541F, W or Y; K557, preferably K557A, G, 1, L or V; A561, preferably A561F, W or Y; T566, preferably T566F, W or Y; A569, preferably A569H or P in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn. 42. The conjugation-competent polypeptide of any preceding Paragraph comprising one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions V547, preferably V457A; K573, preferably K573P or Y; 1523, preferably I523A or G, T527, preferably T527M, K500, preferably K500A; or E505, preferably E505Q in SEQ ID NO. 2; wherein the one or more (e.g. several) alterations causes the conjugation-competent polypeptides to have (i) an altered plasma half-life and/or (ii) an altered binding affinity to FcRn. 43. The conjugation-competent polypeptide of any preceding Paragraph comprising one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions 573,523,527 or 505 of SEQ ID NO. 2, preferably K573Y; I523G; 1523A; T527M; E505Q; or K573P. 44. The conjugation-competent polypeptide of Paragraph 43 comprising one or more (e.g. several) alterations in the mature human albumin polypeptide sequence of SEQ ID NO. 2 selected from the group consisting of positions corresponding to positions K573Y and I523G; K573Y, I523G and T527M; K573Y, E505Q and T527M; K573Y and T527M; K573P and I523G; K573P, I523G and T527M; K573P, E505Q and T527M; K573P and T527M; V547A; V547A and K573P; V547A, E505Q, K573P and T527M; or K500A and H510Q. 45. The conjugation-competent polypeptide of any of Paragraphs 25 to 44 wherein the conjugation-competent polypeptide has a tendency to exist as a monomer in solution which is at least 70% of the tendency of the polypeptide of SEQ ID NO. 2 to exist as a monomer in solution, and optionally at least 75%, at least 80%, at least 90%, at least 95% or at least 100%. 46. The conjugation-competent polypeptide of any preceding Paragraph, in which the polypeptide has at least 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 99.2, 99.4, 99.6, 99.8% sequence identity to SEQ ID NO. 2. 47. The conjugation-competent polypeptide of any preceding Paragraph wherein, when the polypeptide is folded, there are at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, and preferably all 17 of the native disulphide bonds of the polypeptide of SEQ ID NO. 2. 48. The conjugation-competent polypeptide of any preceding Paragraph in which the polypeptide further comprises a further linker to which a bioactive compound, radiopharmaceutical or imaging agent may be linked. 49. The conjugation-competent polypeptide of any preceding Paragraph wherein the alteration(s) to provide a conjugation competent cysteine residue(s) result in a polypeptide with acceptable immunogenicity in human, preferably an immunogenicity which is comparable to or lower than that of wild-type HSA (SEQ ID NO. 2). 50. The conjugation-competent polypeptide of any preceding Paragraph wherein the alteration(s) to provide a conjugation competent cysteine residue(s) does not adversely affect the immunogenicity of the polypeptide in human, e.g. relative to the immunogenicity of wild-type HSA (SEQ ID NO. 2). 51. The conjugation-competent polypeptide of Paragraph 49 or 50 wherein the immunogenicity of the polypeptide is determined or predicted by screening for T-cell epitopes and/or for B-cell epitopes. 52. The conjugation-competent polypeptide of any of Paragraphs 50 to 51 wherein the immunogenicity of the polypeptide is determined or predicted by an ex vivo T cell activation assay. 53. The conjugation-competent polypeptide of Paragraph 52 wherein the T cell activation assay comprises measuring T cell responses using a proliferation assay, e.g. [3H]-thymidine uptake. 54. The conjugation-competent polypeptide of Paragraph 52 or 53 wherein the polypeptide has less than 10% reactivity in the T cell proliferation assay, preferably less than 8, 6, 4, or 2% reactivity, most preferably 0%. 55. The conjugation-competent polypeptide of any of Paragraphs 52 to 54 wherein the T cell activation assay comprises measuring T cell responses using a cytokine secretion assay, e.g. IL-2 ELISpot. 56. The conjugation-competent polypeptide of Paragraph 55 wherein the polypeptide has less than 10% reactivity in the cytokine secretion assay, preferably less than 8, 6, 4, or 2% reactivity, most preferably 0%. 57. The conjugation-competent polypeptide of any of Paragraphs 49 to 56 wherein the polypeptide has less than 10% reactivity in a T cell proliferation assay and in a cytokine secretion assay. 58. The conjugation-competent polypeptide of any preceding Paragraph wherein the polypeptide does not stimulate an adverse antibody response in human. 59. A fusion polypeptide comprising a conjugation-competent polypeptide of any preceding Paragraph and a fusion partner polypeptide. 60. A polynucleotide which encodes the polypeptide of any of Paragraphs 1 to 59. 61. A plasmid comprising the polynucleotide of Paragraph 60. 62. A host cell comprising a polynucleotide of Paragraph 60 and/or a plasmid of Paragraph 61. 63. The host cell of Paragraph 62, which is a yeast cell, particularly a Saccharomyces cerevisiae cell. 64. A conjugate which comprises a bioactive compound, radiopharmaceutical or imaging agent, and a polypeptide according to any of Paragraphs 1 to 59, wherein the bioactive compound is radiopharmaceutical or imaging agent, linked to the polypeptide through a conjugation-competent cysteine residue of the polypeptide. 65. The conjugate of Paragraph 64 further comprising one or more (e.g. several) further bioactive compounds radiopharmaceuticals or imaging agents, each bioactive compound, radiopharmaceutical or imaging agent, being linked to the polypeptide through a conjugation-competent cysteine residue of the polypeptide. 66. A method of producing the polynucleotide of Paragraph 60 comprising: [0481] (a) providing a nucleic acid molecule encoding a parent albumin or fragment thereof; and [0482] (b) modifying the nucleic acid sequence of the nucleic acid molecule to encode a conjugation-competent polypeptide which is at least 70% identical to human albumin, particularly residues 1 to 585 of the mature human albumin polypeptide sequence of SEQ ID NO. 2, or a fragment thereof, wherein at least one position equivalent to a position selected from K93, E294, A226, E230, 1271, E358, L24, F49, V54, D56, L66, A92, Q94, E97, H128, F156, E227, D237, K240, D259, K262, N267, Q268, L275, E277, L284, E311, K317, A322, E333, D340, E354, K359, A362, E382, and L398, particularly from K93, E294, A226, E230, and I271, of SEQ ID NO. 2 comprises a conjugation-competent cysteine residue. 67. A method of producing the polypeptide of any of Paragraphs 1 to 59, comprising: [0483] (a) culturing the host cell of Paragraph 62 or 63 under conditions that allow expression of the polypeptide; and
[0484] (b) recovering the polypeptide from the host cell and/or from host cell growth medium. 68. The method of paragraph 67 in which the host cell exhibits enhanced chaperone activity. 69. The method of Paragraph 67 or 68 further comprising purifying the polypeptide obtained in step (b). 70. A method of producing the conjugate of Paragraph 64 or 65 which comprises linking a polypeptide of any one of Paragraphs 1 to 59, or produced by the method of any one of Paragraphs 67 to 69, to a bioactive compound, radiopharmaceutical or imaging agent, through a conjugation-competent cysteine residue of the polypeptide. 71. An associate comprising the conjugate of Paragraph 64 or 65 and a bioactive, therapeutic, prophylactic, diagnostic, imaging or other beneficial moiety. 72. A nanoparticle or a microparticle or a liposome comprising the polypeptide of any one of Paragraphs 1 to 59, the conjugate of Paragraph 64 or 65 or the associate of Paragraph 71. 73. A composition comprising the conjugate of Paragraph 64 or 65, the associate of Paragraph 71 or the nanoparticle or microparticle or liposome of Paragraph 72 and at least one (e.g. several) pharmaceutically acceptable carrier or diluent. 74. The conjugate of Paragraph 64 or 65, the associate of Paragraph 71, the nanoparticle or microparticle or liposome of Paragraph 72, or the composition of Paragraph 73, wherein the bioactive molecule, radiopharmaceutical or imaging agent, is selected from those described herein. 75. The conjugate of Paragraph 64, 65 or 74, or the associate of Paragraph 71, the nanoparticle or microparticle or liposome of Paragraph 72 for treatment of disease, treatment of illness and/or for diagnosis. 76. Use of a polypeptide as defined in any of Paragraphs 1 to 59 to increase half-life of a bioactive molecule, radiopharmaceutical or imaging agent. The invention is further described by the following examples that should not be construed as limiting the scope of the invention.
Examples
Example 1: Preparation of Variants
Preparation of Specific HSA Variant Expression Plasmids.
[0485] Methods for the expression of HSA variants were performed using several techniques, employing standard molecular biology techniques throughout, such as described in Sambrook, J. and D. W. Russell, 2001 (Molecular Cloning: a laboratory manual, 3.sup.rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y).
Method 1.
[0486] Single amino acid mutations (K93C, A226C, E230C, I271C, E294C, and E358C) were introduced into the pDB5155 plasmid (encoding mutated C34A HSA, SEQ ID NO. 30) using a mutagenic forward primer and non-mutagenic reverse primer (Table 3). pDB5155, encoding a C34A mutant, based on the plasmid pDB5102 was made using a mutagenic forward primer and a non-mutagenic reverse primer (Table 3). pDB5102 is described in WO 2015/036579. Methylated template DNA was prepared by mixing about 1.7 .mu.g of plasmid DNA with 5 .mu.L 10.times. buffer (50 mM Tris-HCl mM .beta.-mercaptoethanol, 10 mM EDTA pH 7.5 at 25.degree. C.--New England Biolabs), 1 .mu.L dam methyltransferase (New England Biolabs), 12.5 .mu.L s-adenosylmethionine (New England Biolabs 80 .mu.M final concentration) and water to 50 .mu.l final volume and incubating at 37.degree. C. for 1.5 hours. Reaction mixtures were then purified using a QIAquick PCR purification kit (Qiagen) according to the manufacturer's instructions. The relevant primers were employed in the PCR reaction (described in Tables 4 and 5) using dam-methylated pDB5102 as template and Q5 DNA polymerase (New England Biolabs). Amplification of the plasmid was confirmed by analysis of 5 .mu.l of PCR product on a 1% TBE agarose gel. The remaining PCR product was supplemented with 5 .mu.l buffer 4 (50 mM potassium acetate, 20 mM Tris-acetate, 10 mM magnesium acetate, 1 mM DTT, pH 7.9 at 25.degree. C.--New England Biolabs) and 1 .mu.l DpnI enzyme, followed by incubation at 37.degree. C. for two hours. The reaction mixtures were then purified using a QIAquick PCR purification kit (Qiagen) according to the manufacturer's instructions. 1 .mu.l of purified plasmid was transformed into E. co/i 10-beta cells (New England Biolabs) and plated onto LB plates (5 g/L yeast extract, 10 g/L peptone from casein, 10 g/L NaCl, 12 g/L agar agar (Millers LB agar, Merck Millipore)) supplemented with 50 .mu.g/mL ampicillin. Plasmids were isolated using a Qiagen Plasmid Plus Kit (Qiagen--according to manufacturer's instructions) and sequenced to confirm the presence of the desired mutation within the HSA sequence and the plasmid named pDB5155.
[0487] Methylated pDB5155 template DNA was prepared by mixing about 3.0 .mu.g of plasmid DNA with 5 .mu.L 10x buffer (50 mM Tris-HCl mM .beta.-mercaptoethanol, 10 mM EDTA pH 7.5 at 25.degree. C. --New England Biolabs), 1 .mu.L dam methyltransferase (New England Biolabs), 12.5 .mu.L 80 .mu.M s-adenosylmethionine (New England Biolabs 80 .mu.M final concentration) and water to 50 .mu.l final volume and incubating at 37.degree. C. for two hours. Reaction mixtures were then purified using a QIAquick PCR purification kit (Qiagen) according to the manufacturer's instructions.
[0488] The relevant primers were employed in the PCR reaction (described in Tables 4 and 5) using dam-methylated pDB5155 as template and Q5 DNA polymerase (New England Biolabs).
TABLE-US-00005 TABLE 3 Oligonucleotides for mutagenic amplification with mutated codons underlined (R = reverse, = F Forward) and the resultant protein. Oligo Sequence (5' to 3') SEQ ID NO. C34A R TTGTTGCAAGTATTGAGCGAAAGCGATCAAGACCAA 31 C34A F TTCGCTCAATACTTGCAACAAGCTCCATTCGAAGATCACGTCAAG 32 L24C F GAAGAAAACTTCAAGGCTTTGGTCTGTATCGCTTTCGCTCAATACTTGCA 33 F49C F AGTTGGTCAACGAAGTTACCGAATGTGCTAAGACTTGTGTTGCTGACG 34 V54C F GTTACCGAATTCGCTAAGACTTGTTGTGCTGACGAATCCGCGGAAAAC 35 D56C F GAATTCGCTAAGACTTGTGTTGCTTGTGAATCCGCGGAAAACTGTGACA 36 L66C F CGCGGAAAACTGTGACAAGTCCTGTCACACCTTGTTCGGTGATAAGTT 37 A92C F CGGTGAAATGGCTGACTGTTGTTGTAAGCAAGAACCAGAAAGAAACGAA 38 K93C F GTGAAATGGCTGACTGTTGTGCTTGTCAAGAACCAGAAAGAAACGAATGT 39 Q94C F AAATGGCTGACTGTTGTGCTAAGTGTGAACCAGAAAGAAACGAATGTTTC 40 E97C F ACTGTTGTGCTAAGCAAGAACCATGTAGAAACGAATGTTTCTTGCAACAC 41 H128C F TTGACGTCATGTGTACTGCTTTCTGTGACAACGAAGAAACCTTCTTGAAG 42 F156C F ACTTCTACGCTCCAGAATTGTTGTGTTTCGCTAAGAGATACAAGGCTGC 43 A226C F AGATTGTCTCAAAGATTCCCAAAGTGTGAATTCGCTGAAGTTTCTAAGTTG 44 E227C F TGTCTCAAAGATTCCCAAAGGCTTGTTTCGCTGAAGTTTCTAAGTTGGTT 45 E230C F GATTCCCAAAGGCTGAATTCGCTTGTGTTTCTAAGTTGGTTACTGACTTG 46 D237C F GCTGAAGTTTCTAAGTTGGTTACTTGTTTGACTAAGGTTCACACTGAATGT 47 K240C F TCTAAGTTGGTTACTGACTTGACTTGTGTTCACACTGAATGTTGTCACGG 48 D259C F GGAATGTGCTGATGACAGAGCTTGTTTGGCTAAGTACATCTGTGAAAAC 49 K262C F TGATGACAGAGCTGACTTGGCTTGTTACATCTGTGAAAACCAAGACTCT 50 N267C F GACTTGGCTAAGTACATCTGTGAATGTCAAGACTCTATCTCTTCCAAGTTG 51 Q268C F TTGGCTAAGTACATCTGTGAAAACTGTGACTCTATCTCTTCCAAGTTGAAG 52 I271C F TACATCTGTGAAAACCAAGACTCTTGTTCTTCCAAGTTGAAGGAATGTTGT 53 L275C F ACCAAGACTCTATCTCTTCCAAGTGTAAGGAATGTTGTGAAAAGCCATTG 54 E277C F GACTCTATCTCTTCCAAGTTGAAGTGTTGTTGTGAAAAGCCATTGTTGGAA 55 L284C F AAGGAATGTTGTGAAAAGCCATTGTGTGAAAAGTCTCACTGTATTGCTGAA 56 E294C F AAGTCTCACTGTATTGCTGAAGTTTGTAACGATGAAATGCCAGCTGACTT 57 E311C F CATCTTTGGCTGCTGACTTCGTTTGTTCTAAGGACGTTTGTAAGAACTAC 58 K317C F TTCGTTGAATCTAAGGACGTTTGTTGTAACTACGCTGAAGCTAAGGACG 59 A322C F GACGTTTGTAAGAACTACGCTGAATGTAAGGACGTCTTCTTGGGTATGTT 60 E333C F GTCTTCTTGGGTATGTTCTTGTACTGTTACGCTAGAAGACACCCAGACT 61 D340C F CGAATACGCTAGAAGACACCCATGTTACTCCGTTGTCTTGTTGTTGAG 62 E354C F TGTTGAGATTGGCTAAGACCTACTGTACTACCCTCGAGAAGTGTTGTG 63 E358C F CTAAGACCTACGAAACTACCCTCTGTAAGTGTTGTGCTGCTGCTGACC 64 K359C F GACCTACGAAACTACCCTCGAGTGTTGTTGTGCTGCTGCTGACCCA 65 A362C F AAACTACCCTCGAGAAGTGTTGTTGTGCTGCTGACCCACACGAATGT 66 E382C F TCGATGAATTCAAGCCATTGGTCTGTGAACCACAAAACTTGATCAAGCAA 67 L398C F GCAAAACTGTGAATTGTTCGAACAATGTGGTGAATACAAGTTCCAAAACGC 68 L24C R GACCAAAGCCTTGAAGTTTTCTTCACCCAAGTCCT 69 F49C R TTCGGTAACTTCGTTGACCAACTTGACGTGATCTT 70 V54C R ACAAGTCTTAGCGAATTCGGTAACTTCGTTGACCAA 71 D56C R AGCAACACAAGTCTTAGCGAATTCGGTAACTTCGTT 72 L66C R GGACTTGTCACAGTTTTCCGCGGATTCGTCAGC 73 A92C R ACAACAGTCAGCCATTTCACCGTAGGTTTCTCTC 74 K93C R AGCACAACAGTCAGCCATTTCACCGTAGGTTTCTC 75 Q94C R CTTAGCACAACAGTCAGCCATTTCACCGTAGGTT 76 E97C R TGGTTCTTGCTTAGCACAACAGTCAGCCATTTCAC 77 H128C R GAAAGCAGTACACATGACGTCAACTTCTGGTCTAA 78 F156C R CAACAATTCTGGAGCGTAGAAGTATGGGTGTCTTC 79 A226C R CTTTGGGAATCTTTGAGACAATCTAGCGACAGCC 80 E227C R AGCCTTTGGGAATCTTTGAGACAATCTAGCGACAG 81 E230C R AGCGAATTCAGCCTTTGGGAATCTTTGAGACAATCT 82 D237C R AGTAACCAACTTAGAAACTTCAGCGAATTCAGCCTT 83 K240C R AGTCAAGTCAGTAACCAACTTAGAAACTTCAGCGAA 84 D259C R AGCTCTGTCATCAGCACATTCCAACAAGTCACCG 85 K262C R AGCCAAGTCAGCTCTGTCATCAGCACATTCCAAC 86 N267C R TTCACAGATGTACTTAGCCAAGTCAGCTCTGTCATC 87 Q268C R GTTTTCACAGATGTACTTAGCCAAGTCAGCTCTGT 88 I271C R AGAGTCTTGGTTTTCACAGATGTACTTAGCCAAGTC 89 L275C R CTTGGAAGAGATAGAGTCTTGGTTTTCACAGATGTA 90 E277C R CTTCAACTTGGAAGAGATAGAGTCTTGGTTTTCACAG 91 L284C R CAATGGCTTTTCACAACATTCCTTCAACTTGGAAGA 92 E294C R AACTTCAGCAATACAGTGAGACTTTTCCAACAATGG 93 E311C R AACGAAGTCAGCAGCCAAAGATGGCAAGTCAGCT 94 K317C R ACAAACGTCCTTAGATTCAACGAAGTCAGCAGCC 95 A322C R TTCAGCGTAGTTCTTACAAACGTCCTTAGATTCAACG 96 E333C R GTACAAGAACATACCCAAGAAGACGTCCTTAGCTTC 97 D340C R TGGGTGTCTTCTAGCGTATTCGTACAAGAACATAC 98 E354C R GTAGGTCTTAGCCAATCTCAACAACAAGACAACGG 99 E358C R GAGGGTAGTTTCGTAGGTCTTAGCCAATCTCAACA 100 K359C R CTCGAGGGTAGTTTCGTAGGTCTTAGCCAATCTC 101 A362C R ACAACACTTCTCGAGGGTAGTTTCGTAGGTCTTAG 102 E382C R GACCAATGGCTTGAATTCATCGAAAACCTTAGCGT 103 L398C R TTGTTCGAACAATTCACAGTTTTGCTTGATCAAGTTTTG 104
TABLE-US-00006 TABLE 4 PCR reaction components Template (5 ng/.mu.L) 1 .mu.L Forward primer (10 .mu.M) 2.5 .mu.L 5x buffer 10 .mu.L Reverse primer (10 .mu.M) 2.5 .mu.L dNTP (2.5 mM) 1 .mu.L Q5 polymerase 0.5 .mu.L Sterile water 32.5 .mu.L
TABLE-US-00007 TABLE 5 PCR reaction conditions Temperature Cycle Length Number of cycles 98.degree. C. 2 min 1 98.degree. C. 10 sec 30 60.degree. C. 30 sec 72.degree. C. 5 min 72.degree. C. 7 min 1
[0489] Amplification of the plasmid was confirmed by analysis of 5 .mu.l of PCR product on a 1% TBE agarose gel. The remaining PCR product was supplemented with 4 .mu.l buffer 4 (50 mM potassium acetate, 20 mM Tris-acetate, 10 mM magnesium acetate, 1 mM OTT, pH 7.9 at 25.degree. C. --New England Biolabs) and 1 .mu.l DpnI enzyme, followed by incubation at 37.degree. C. for one hour. The reaction mixtures were then purified using a QIAquick 96 PCR purification kit (Qiagen) according to the manufacturer's instructions. 2 .mu.l of purified plasmid was transformed into competent E. coli DH5-alpha cells and grown in a 96 deep well block in 1.2 mL LB media (1% w/v bacteriological tryptone, 0.5% w/v yeast extract, 0.5% w/v NaCl) supplemented with 50 .mu.g/mL ampicillin to repair nicks in the DNA backbone. Plasmids were isolated using a QiaPrep 96 turbo miniprep kit (Qiagen--according to manufacturer's instructions). The thio-albumin constructs are detailed in Table 6.
[0490] Plasmid DNA was prepared for transformation into S. cerevisiae as described in WO 2015/036579 (incorporated herein by reference), Method 4, except that 9723 bp Acc651-BamHI fragment from pDB4164 was used as the gapped vector fragment instead of the 9721 bp fragment from pDB3936, which has two additional bases GC next to the BamHI site to create a NotI restriction site GCGGCCGC (additional bases in bold). pDB3936 is described in WO 2011/124718 (incorporated herein by reference). pDB4164 also differs from pDB3936 in containing a 1368 bp sequence between the Acc65I and BamHI sites containing an apramycin resistance selectable marker which was excised by the Acc65I and BamHI digestion and was not used in the gap-repair transformation. The host strain for the constructs was S. cerevisiae BXP10 cir.sup.0 (WO 2015/036759, incorporated herein by reference). Transformed cells were grown as single colonies on selective agar plates (BMMD+CSM-Leu or BMMD) from which isolated colonies were patched out, also on selective agar plates, for the preparation of cryopreserved yeast stocks and samples for analysis. Cryopreserved stocks were made from 5 mL of a 48 hour BMMD+CSM-Leu shake flask culture mixed with an equal volume of 40% [w/v] trehalose and 1 mL aliquots transferred to cryovials for storage at -80.degree. C. 0.5 mL BMMD in 48-well microtitre plate wells was inoculated with yeast from the patch plates and grown for 4-days at 30.degree. C. with shaking as described in WO 2015/036579, Method 4 (incorporated herein by reference). Shake flask cultures were inoculated from trehalose stocks. Purification of these variants from shake flask was performed as described in WO 2012/150319 (incorporated herein by reference).
[0491] Preparation of the expression plasmids for the L24C, F49C, V54C, D56C, L66C, A92C, Q94C, E97C, H128C, F156C, E227C, D237C, K240C, D259C, K262C, N267C, Q268C, L275C, E277C, L284C, E311C, K317C, A322C, E333C, D340C, E354C, K359C, A362C, E382C, and L398C (all in C34A background) was slightly different to that described above:
[0492] Single amino acid mutations were introduced into the pDB5155 plasmid (encoding mutated C34A HSA, SEQ ID NO. 30) using a mutagenic forward primer and non-mutagenic reverse primer (Table 3).
[0493] Methylated template DNA was prepared by mixing about 2.5 .mu.g of plasmid DNA with 5 .mu.L 10x buffer (50 mM Tris-HCl mM .beta.-mercaptoethanol, 10 mM EDTA pH 7.5 at 25.degree. C.--New England Biolabs), 1 .mu.L dam methyltransferase (New England Biolabs), 12.5 .mu.L 80 .mu.M s-adenosylmethionine (New England Biolabs 80 .mu.M final concentration) and water to 50 .mu.l final volume and incubating at 37.degree. C. for one hour. Reaction mixtures were then purified using a QIAquick PCR purification kit (Qiagen) according to the manufacturer's instructions.
[0494] The relevant primers were employed in the PCR reaction (described in Tables 4 and 5, above) using dam-methylated pDB5155 as template and Q5 DNA polymerase (New England Biolabs).
[0495] Amplification of the plasmid was confirmed by analysis of 5 .mu.l of PCR product on a 1% TBE agarose gel. The remaining PCR product was supplemented with 4 .mu.l buffer 4 (50 mM potassium acetate, 20 mM Tris-acetate, 10 mM magnesium acetate, 1 mM DTT, pH 7.9 at 25.degree. C. --New England Biolabs) and 1 .mu.l DpnI enzyme, followed by incubation at 37.degree. C. for one hour. The reaction mixtures were then purified using a QIAquick 96 PCR purification kit (Qiagen) according to the manufacturer's instructions. 1 .mu.l of purified plasmid was transformed into competent E. coli DH5-alpha cells and grown in a 96 deep well block in 1.2 mL LB media (1% w/v bacteriological tryptone, 0.5% w/v yeast extract, 0.5% w/v NaCl) supplemented with 50 .mu.g/mL ampicillin to repair nicks in the DNA backbone. Plasmids were isolated using a QiaPrep 96 turbo miniprep kit (Qiagen--according to manufacturer's instructions). The thio-albumin constructs are detailed in Table 6.
[0496] Plasmid DNA was prepared for transformation into S. cerevisiae as described in WO 2015/036579, Method 4 (incorporated herein by reference). The host strain for the constructs was S. cerevisiae DYB7 (Payne et al. (2008) Applied and Environmental Microbiology Vol 74(24):7759-7766). The yeast microtitre plate growth diverged from the method as described in WO 2015/036579 in that transformations were performed in duplicate and the initial growth was for two days. Stocks were produced from the two days growth by transfer of 50 .mu.l culture to a fresh microtitre plate containing 50 .mu.l 40% (w/v) trehalose. 50 .mu.l of the two day culture was also added to a fresh microtitre plate containing 450 .mu.L of BMMD+CSM-leu and incubated at 30.degree. C. with shaking (200 rpm, 2.5 cm orbit at in a sealed chamber at 100% humidity in an Eppendorf Innova 44 incubated shaker) for a further four days. Culture supernatants were harvested by centrifugation at 3000 rpm for 5 minutes and 375 .mu.l of supernatant was transferred to a fresh 48-well microtitre plate.
Production of Expression Plasmid and Yeast Stocks.
[0497] Preparation of the expression plasmids and transformation of S. cerevisiae was performed as described in WO 2011/051489 and WO 2012/150319 (incorporated herein by reference) by the 48-hour stocking method, using equal volumes of culture and trehalose. The host strain for the constructs was S. cerevisiae BXP10 Cir.degree. (WO 2015/036759, incorporated herein by reference). Purification of variants from shake flask was performed as described in WO 2012/150319 unless otherwise stated.
The resultant albumin variants are summarized in Table 6.
TABLE-US-00008 TABLE 6 Albumin variant SEQ ID NO. C34A 30 C34A + L24C 105 C34A + F49C 106 C34A + V54C 107 C34A + D56C 108 C34A + L66C 109 C34A + A92C 110 C34A + K93C 111 C34A + Q94C 112 C34A + E97C 113 C34A + H128C 114 C34A + F156C 115 C34A + A226C 116 C34A + E227C 117 C34A + E230C 118 C34A + D237C 119 C34A + K240C 120 C34A + D259C 121 C34A + K262C 122 C34A + N267C 123 C34A + Q268C 124 C34A + I271C 125 C34A + L275C 126 C34A + E277C 127 C34A + L284C 128 C34A + E294C 129 C34A + E311C 130 C34A + K317C 131 C34A + A322C 132 C34A + E333C 133 C34A + D340C 134 C34A + E354C 135 C34A + E358C 136 C34A + K359C 137 C34A + A362C 138 C34A + E382C 139 C34A + L398C 140
Example 2. Thiol Determination of DTNB Incubated Thio-Albumin Variants
[0498] The free thiol content of thiol albumin variants was determined at small scale using microtitre plate (MTP) grown cultures. The tested thiol albumin variants included the C34A substitution, and thus should lack the thiol group of native albumin. As such, they were each expected to have only one free thiol.
[0499] The number of free thiols on a protein can be determined spectrophotometrically using Ellman's reagent. Ellman's reagent (5'5'-dithio-bis(2-nitrobenzoic acid) (DTNB)) is an aromatic disulphide which reacts with thiol groups to form a mixed disulphide of the protein and one mole of 5-thio-2-nitrobenzoic acid (TNB) (per mole of protein sulfhydryl group). This reaction also results in a yellow colour from free TNB being released in solution. Alternatively the number of free thiols on a protein can be determined using mass spectrometric analysis of protein sample treated with DTNB reagent. 5-thio-2-nitrobenzoic acid (TNB) has a molecular weight of 199 Da, thus an increase in mass of 197 Da (TNB minus H.sub.2 lost during disulphide bond formation with the free thiol group on the test protein) indicates the presence of one free thiol group on the protein sample.
[0500] 4 .mu.l Buffer 2 (4 mg/mL DTNB, 500 mM sodium phosphate, pH 7.0) was added to 200 .mu.L of the test protein culture sample in a 96-well MTP format. The preparation was allowed to incubate for 25 minutes at ambient temperature (20.+-.5.degree. C.) to allow TNB labelling. Protein intact mass was determined by UltraPerformance Liquid Chromatography Mass Spectrometry (UPLC-MS). UPLC separation was carried out on 10 .mu.L of sample using a Waters Acquity on a BEH 50.times.2.1 mm ACQUITY BEH 1.7 .mu.m 300 .ANG. C4 column and a 5 min analytical gradient of buffer A 0.1% formic acid and Buffer B 100% acetonitrile 0.1% formic acid. Eluted proteins were directly introduced to a Bruker MicrOTOF II mass spectrometer via an Electrospray Ionisation (ESI) source. All instrument control and sample tables were controlled using BioPharma Compass.TM.. All data were manually processed over the leading edge of the protein peak between 2.9-3.0 minutes in Data Analysis. This included spectral smoothing using a Gauss smoothing algorithm set at 0.0765 Da and a baseline correction setting of 0.8 flatness. Deconvoluted intact mass spectra were obtained using the Max. Entropy algorithm, all methods and parameters were set within BioPharma Compass.TM..
[0501] The results of the above thiol analysis of the thio-albumin samples are summarised in Table 7. An increase in mass of 197 Da upon DTNB incubation is predicted to be indicative of the presence of one free thiol group on the protein in the sample. A mass increase of 197.+-.15 Da as actually measured by MS was taken as indicative of the correct mass. All variants successfully bound a molecule of TNB.
TABLE-US-00009 TABLE 7 Mass Spectrometry DTNB thiol screening results Molecular weight (Da) Post DTNB treatment Variant Difference description Variant Actual (Actual minus (all C34A) Theoretical Theoretical measured theoretical) L24C 66397 66594 66599 5 F49C 66363 66560 66568 8 V54C 66411 66608 66613 5 D56C 66395 66592 66600 8 L66C 66397 66594 66599 5 A92C 66439 66636 66641 5 K93C 66382 66579 66588 9 Q94C 66382 66579 66581 2 E97C 66381 66578 66580 2 H128C 66373 66570 66572 2 F156C 66363 66560 66564 4 A226C 66439 66636 66637 1 E227C 66381 66578 66584 6 E230C 66381 66578 66582 4 D237C 66395 66592 66593 1 K240C 66382 66579 66584 5 D259C 66395 66592 66594 2 K262C 66382 66579 66584 5 N267C 66396 66593 66592 -1 Q268C 66382 66579 66584 5 I271C 66397 66594 66596 2 L275C 66397 66594 66597 3 E277C 66381 66578 66583 5 L284C 66397 66594 66592 -2 E294C 66381 66578 66581 3 E311C 66381 66578 66589 11 K317C 66382 66579 66582 3 A322C 66439 66636 66640 4 E333C 66381 66578 66582 4 D340C 66395 66592 66602 10 E354C 66381 66578 66583 5 E358C 66381 66578 66583 5 K359C 66382 66579 66583 4 A362C 66439 66636 66641 5 E382C 66381 66578 66586 8 L398C 66397 66594 66597 3
Example 3. Aggregation Screening of Thio-Albumin Variants
[0502] Variants were tested for tendency to remain as a monomer in solution. Each variant has a single free thiol group. Therefore, they were tested in comparison with wild-type HSA, which also has a single free thiol group.
[0503] Shake flask culturing of S. cerevisiae and purification was performed as described in WO 2012/150319 (incorporated herein by reference) with the following modifications. BMMS media (10 mL) was inoculated with S. cerevisiae and grown for 2 days at 30.degree. C. with orbital shaking at 200 rpm. An aliquot of each starter culture (5 mL) was used to inoculate 2.times.200 mL BMMS media and grown for 5 days at 30.degree. C. with orbital shaking at 200 rpm. Cells were harvested by filtration through a 0.2 .mu.m vacuum filter membrane (Nalgene Sterile Top Filter) and the supernatant retained for purification.
[0504] A single step chromatography procedure was used to prepare purified material from the thio-albumin variants. The purification step used a column (bed volume approximately 2 mL) packed with AlbuPure.RTM. matrix (ProMetic BioSciences Ltd, Cambridge UK or Albumedix Ltd (formerly Novozymes Biopharma UK Ltd)). This was equilibrated with 50 mM sodium acetate, pH 5.3, and loaded with neat shake flask culture supernatants, at approximately pH 5.5-6.5, to approximately 20 mg protein/mL matrix. The column was washed with approximately 10 column volumes each of 50 mM sodium acetate, pH 5.3, and 50 mM ammonium acetate, pH 8.0, respectively. Bound protein was eluted using approximately 10 column volumes of 50 mM ammonium acetate, 10 mM octanoate, pH 7.0. The flow rate throughout was 240 cm/h using an AKTA Explorer system (GE Healthcare). Eluate samples were approximately 20 mL in volume. The concentration and percentage monomer of the eluate samples was determined by Gel Permeation High Pressure Liquid Chromatography (GP-HPLC). Protein concentrations were determined using a LC2010 HPLC system (Shimadzu) equipped with UV detection under Shimadzu VP7.3 client server software control. Injections of 25 .mu.L were made onto a 7.8 mm internal diameter x 300 mm length TSK G3000SWXL column (Tosoh Bioscience), with a 6.0 mm internal diameter x 40 mm length TSK SW guard column (Tosoh Bioscience). Samples were chromatographed in 25 mM sodium phosphate, 100 mM sodium sulphate, 0.05% (w/v) sodium azide, pH 7.0 at 1 mLmin.sup.-1, with a run time of 15 minutes. Samples were quantified by UV detection at 280 nm, by peak area, relative to a recombinant human albumin standard of known concentration (10 mg/mL).
[0505] The samples were reanalysed to determine the change in percentage monomer post seven weeks storage at 2-8.degree. C., and post 6 months storage at 2-8.degree. C. The percentage monomer (in brackets) was determined for each sample relative to its wild type control under the same storage conditions. The results are summarised in Table 8A. Final eluate concentrations were in the range of 0.6-1.2 mg/mL, resulting in 12-24 mg protein recovered post purification. All variants had a monomer percentage equivalent to or higher than that of the wild type control at T=0, which had a monomer percentage of 87%. The variants maintained their monomeric protein percentage over 7 weeks' storage at 2-8.degree. C., with no significant evidence of aggregation propensity during 6 months storage at 2-8.degree. C. observed for at least four variants.
TABLE-US-00010 TABLE 8A GPHPLC aggregation screening results GPHPLC % Monomer .DELTA.% Monomer conc. T = 7 T = 6 0-7 0-6 Sample (mg/mL) T = 0 week month week month WT albumin 1.1 87 (100) 88 (100) 89 (100) 1 2 control C34A + K93C 0.7 91 (105) 92 (105) 92 (103) 1 1 C34A + A226C 1.1 93 (107) 93 (106) 93 (105) 0 0 C34A + E230C 0.6 90 (103) 91 (103) ND (ND) 1 ND C34A + I271C 1.2 91 (105) 91 (103) 91 (102) 0 0 C34A + E294C 0.9 96 (110) 96 (109) 96 (108) 0 0 C34A + E358C 1.0 89 (102) 83 (94) 80 (90) -6 -9 ND: Not determined
[0506] Further variants were analysed using the method previously described in Example 3, or alternatively using an Agilent 1260 isocratic UHPLC (Ultra-High Performance Liquid Chromatography) instrument. For the UHPLC method, injections of 4 .mu.L were made onto a 4.6 mm id.times.150 mm length BEH 200 .ANG., 1.7 .mu.m column (Waters), using the mobile phase described in Example 3, at 0.5 mLmin.sup.-1, with a run time of 5 minutes. Samples were quantified by UV detection at 280 nm, by peak height relative to a recombinant human albumin standard of known concentration (10 mg/mL).
[0507] The samples were reanalysed post eight weeks storage at 2-8.degree. C., and post 4 months storage at 2-8.degree. C. to determine the change in percentage monomer. The percentage monomer (in brackets) was determined for each sample relative to its wild type control under the same storage conditions. The results are summarised in Table 8B. Final eluate concentrations were in the range of 0.1-1.0 mg/mL, resulting in 2-20 mg protein recovered post purification. The majority of variants had a monomer percentage equivalent to or higher than that of the wild type control at T=0, which had a monomer percentage of 86%. These variants maintained their monomeric protein over 8 weeks' storage at 2-8.degree. C., with no significant evidence of aggregation propensity during 4 months storage at 2-8.degree. C. observed. However, it was evident that variants C34A+L66C, C34A+E277C, and C34A+E311C had a relatively low percentage monomer at T=0, and consequently had a propensity to form aggregates.
TABLE-US-00011 TABLE 8B GPHPLC aggregation screening results GPHPLC % Monomer .DELTA. % Monomer conc. T = 8 T = 4 0-8 0-4 Sample (mg/mL) T = 0 week month week month WT albumin 0.6 86 (100) 88 (100) 87 (100) 2 1 control C34A + L24C 0.7 94 (109) 96 (109) 97 (112) 2 3 C34A + F49C 0.5 94 (109) 95 (108) 94 (108) 1 0 C34A + V54C 0.5 93 (108) 94 (107) 93 (107) 1 0 C34A + D56C 0.3 85 (99) 77 (88) 75 (86) -8 -10 C34A + L66C 0.2 7 (8) 12 (14) 6 (7) 5 -1 C34A + A92C 0.9 93 (108) 94 (107) 94 (108) 1 1 C34A + Q94C 0.1 95 (111) 96 (109) 95 (109) 1 0 C34A + E97C 0.5 88 (102) 85 (97) 85 (98) -3 -3 C34A + H128C 0.6 92 (107) 93 (106) 93 (107) 1 1 C34A + F156C 1.0 92 (107) 94 (107) 94 (108) 2 2 C34A + E227C 0.5 86 (100) 88 (100) 88 (101) 2 2 C34A + D237C 0.5 93 (108) 95 (108) 94 (108) 2 1 C34A + K240C 0.6 93 (108) 94 (107) 94 (108) 1 1 C34A + D259C 0.5 93 (108) 95 (108) 94 (108) 2 1 C34A + K262C 0.6 92 (107) 93 (106) 93 (107) 1 1 C34A + N267C 0.6 94 (109) 95 (108) 95 (109) 1 1 C34A + Q268C 0.8 95 (111) 96 (109) 96 (110) 1 1 C34A + L275C 0.5 94 (109) 95 (108) 94 (108) 1 0 C34A + E277C 0.7 65 (76) 60 (68) 59 (68) -5 -6 C34A + L284C 0.7 92 (107) 94 (107) 94 (108) 2 2 C34A + E311C 0.7 54 (63) 48 (55) 46 (53) -6 -8 C34A + K317C 0.6 83 (97) 82 (93) 82 (94) -1 -1 C34A + A322C 0.8 81 (94) 84 (96) 83 (95) 3 2 C34A + E333C 0.3 94 (109) 97 (110) 95 (109) 3 1 C34A + D340C 0.6 93 (108) 94 (107) 94 (108) 1 1 C34A + E354C 0.7 89 (104) 90 (102) 90 (103) 1 1 C34A + K359C 0.6 86 (100) 87 (99) 87 (100) 1 1 C34A + A362C 0.6 89 (104) 89 (101) 88 (101) 0 -1 C34A + E382C 0.6 86 (100) 84 (96) 84 (97) -2 -2 C34A + L398C 0.7 90 (105) 92 (105) 87 (100) 2 -3
Example 4. Conjugation Efficiency and Controlled Hydrolysis of Thio-Albumin Variants
[0508] Thio-albumin variants from Example 3 were conjugated with biotin (Thermo Scientific, EZ-Link Maleimide-PEG2-Biotin) using a 3.2 fold molar excess of maleimide-PEG2-biotin to protein. A reaction schematic is shown in FIG. 4. The thio-albumin AlbuPure.RTM. eluates were diluted with phosphate buffered saline (PBS buffer), pH 7.4 to give 10 mL solutions at 0.3 mg/mL (45.15 nmol) and conjugated as described below Table 9A.
[0509] The MS spectrum for the thio-albumin variant C34A+A226C indicated that no conjugation had occurred post an overnight incubation with maleimide-PEG2-biotin. The results are summarised in Table 9A. The MS spectra for the thio-albumin variants C34A+E230C, and C34A+I271C indicated that conjugation had occurred post an overnight incubation, giving approximately 72% or 72% monoconjugate respectively (i.e. the same level of monoconjugate) when comparing the relative peak heights of conjugated and unconjugated species. The MS spectrum for C34A+I271C is shown in FIG. 5A. The MS spectrum for thio-albumin variant C34A+K93C shown in FIG. 5B, exhibited a single species at 66908 Da indicating the correct molecular weight for the thio-albumin variant plus a single addition of maleimide-PEG2-biotin (+525 Da). This confirmed the variant had a single free thiol available for conjugation. Comparable results were obtained for thio-albumin variants C34A+E294C and C34A+E358C.
TABLE-US-00012 TABLE 9A Conjugation efficiency results Reference Mr Theoretical Conjugate % Sample unconjugated conjugate intact mass conju- Description (Da) mass (Da) result (Da) gation WT control 66439 66964 * * C34A + K93C 66382 66907 66908 100 C34A + A226C 66439 66964 66440 0 C34A + E230C 66381 66906 66908 72 C34A + I271C 66397 66922 66924 72 C34A + E294C 66381 66906 66909 >95 C34A + E358C 66381 66906 66909 100 * WT control sample failed to inject on MS during sequence run.
[0510] Further variants were analysed and the results are shown in Table 9B. For samples C34A+L66C and C34A+Q94C the protein concentrations were low, hence 10 mL solutions at 0.15 mg/mL (22.58 nmol) were used. Stock solutions of 2 mg/mL biotin were prepared by the addition of 5.times.200 .mu.L aliquots of PBS buffer, pH 7.4, to each of two 2 mg pre-weighed EZ-Link micotubes, the vials were rinsed to maximise recovery of the lyophilised product. The two 1 mL volumes were pooled into a 7 mL container with a lid. From the biotin stock solution, 38 .mu.L (144.5 nmol) was added to the 10 mL albumin samples to give approximately a 3.2-fold molar excess of biotin over albumin. However, for the C34A+L66C and C34A+Q94C samples only 19 .mu.L biotin was added to maintain a 3.2 fold excess of maleimide-PEG2-biotin to protein. Samples were gently mixed and incubated at ambient temperature overnight. Post incubation, the samples were subjected to mass spectrometry to determine the intact protein mass post conjugation according to the method described in Example 2, but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The MS spectra results summarised in Table 9B indicated that thio-albumin variants C34A+L66C, C34A+A92C, C34A+Q94C, C34A+D259C, C34A+L275C, and C34A+L284C did not conjugate post an overnight incubation with maleimide-PEG2-biotin. The MS spectra for the WT control, and the thio-albumin variants C34A+L24C, C34A+V54C, C34A+H128C, C34A+E227C, C34A+K240C, C34A+K262C, C34A+Q268C, C34A+E277C, C34A+K317C, C34A+A322C, C34A+K359C and C34A+A362C indicated 90% conjugation or greater with maleimide-PEG2-biotin.
TABLE-US-00013 TABLE 9B Conjugation efficiency results Reference Mr Theoretical Conjugate % Sample unconjugated conjugate intact mass conju- Description (Da) mass (Da) result (Da) gation WT control 66439 66964 66966 93 C34A + L24C 66397 66922 66924 96 C34A + F49C 66363 66888 66889 84 C34A + V54C 66411 66936 66938 100 C34A + D56C 66395 66920 66922 79 C34A + L66C 66397 66922 66400 0 C34A + A92C 66439 66964 66407 0 C34A + Q94C 66382 66907 66409 0 C34A + E97C 66381 66906 66907 9 C34A + H128C 66373 66898 66899 100 C34A + F156C 66363 66888 66890 76 C34A + E227C 66381 66906 66907 95 C34A + D237C 66395 66920 66921 73 C34A + K240C 66382 66907 66908 100 C34A + D259C 66395 66920 67424 0 C34A + K262C 66382 66907 66908 100 C34A + N267C 66396 66921 66922 47 C34A + Q268C 66382 66907 66908 92 C34A + L275C 66397 66922 66897 0 C34A + E277C 66381 66906 66908 90 C34A + L284C 66397 66922 67427 0 C34A + E311C 66381 66906 66909 76 C34A + K317C 66382 66907 66909 91 C34A + A322C 66439 66964 66965 94 C34A + E333C 66381 66906 66907 83 C34A + D340C 66395 66920 66923 12 C34A + E354C 66381 66906 66908 32 C34A + K359C 66382 66907 66908 95 C34A + A362C 66439 66964 66966 94 C34A + E382C 66381 66906 66909 83 C34A + L398C 66397 66922 66925 36
[0511] The stability of maleimide conjugate bonds is not robust. The succinimide can revert back to maleimide and free thiol via a retro-Michael pathway (FIG. 4). Thus, highly undesirably, the released maleimide may react with other thiol reactive species and the released thiol may react with other compounds in vivo. To avoid retro-Michael reactivity, the succinimide may be hydrolysed to succinic acid, effectively taking on H.sub.2O (+18 Da) and locking the conjugate to be thiol-stable. The property of thiol-stability by hydrolysis is desirable as it would ensure that there was no unwanted thiol transfer taking place in various environments in vivo. Therefore, controlled hydrolysis of the succinimide was performed by increasing the pH and temperature. Post conjugation the samples were transferred to Vivaspin 20 centrifugal concentrators (Sartorius) and balanced with PBS buffer pH 7.4. The samples were centrifuged at 4,500.times.g for 15 minutes to reduce the volume to approximately 200 .mu.L. A diafiltration cup was fitted to the Vivaspin 20 vessels and subsequently filled with 15 mL of PBS buffer pH 9.0. The samples were centrifuged at 4,500.times.g for 15 minutes a second time. A further 15 mL PBS buffer pH 9.0 was added and the samples centrifuged a third time to ensure that all the free maleimide-PEG2-biotin was removed from solution. The remaining retentate was removed and made up to a final volume of 10 mL with PBS buffer pH 9.0 (i.e. assuming no losses then to a concentration of 0.3 mg/mL). The samples were incubated at 37.degree. C. for at least 24 hours for controlled hydrolysis to occur to determine the stability of the thio ether conjugate bond. The results are summarised in Table 10.
[0512] The yield of the hydrolysed thiol stable wild type control conjugate was in the order of 53%, likely due to the competing retro-Michael deconjugation during hydrolysis (FIG. 6A). Also observed was an average conjugate mass shift of +14 Da indicating that partial hydrolysis had occurred. It was apparent that the thio-albumin variants that had the highest conjugation efficiency also had improved conjugate stability upon controlled hydrolysis. Specifically the reaction favoured the hydrolysis of the succinimide rather than the retro-Michael deconjugation pathway. An example of C34A+E294C is shown in FIG. 6B indicating no conjugate losses following incubation at pH 9.0, 37.degree. C. Comparable results were obtained for thio-albumin variants C34A+K93C, C34A+E294C and C34A+E358C with no significant losses during controlled hydrolysis.
TABLE-US-00014 TABLE 10 Controlled hydrolysis stability results Reference Mr Theoretical Conjugate Conjugate % conjugation Sample unconjugated conjugate intact mass mass increase post Description (Da) mass (Da) result (Da) (Da) hydrolysis WT control 66439 66964 66978 14 53 C34A + K93C 66382 66907 66911 4 100 C34A + A226C 66439 66964 66441 2 0 C34A + E230C 66381 66906 66926 20 63 C34A + I271C 66397 66922 66939 6 61 C34A + E294C 66381 66906 66926 20 100 C34A + E358C 66381 66906 66927 21 100
[0513] The combined aggregation results and conjugation results are summarised together in Table 11. It was apparent that the variants C34A+K93C and C34A+E294C had improved aggregation profiles compared to wild type albumin, conjugated to a high percentage with maleimide-PEG2-biotin, and had minimal loss of conjugate following controlled hydrolysis at pH 9.0, 37.degree. C. These variants were selected for further evaluation.
TABLE-US-00015 TABLE 11 Thio-albumin variant aggregation screen and conjugation results summary No losses Improved during Sample aggregation Conjugation controlled Variant Description profile efficiency >95% hydrolysis selected WT control C34A + K93C C34A + A226C C34A + E230C C34A + I271C C34A + E294C C34A + E358C
Example 5: Combination Variants
Method 2.
[0514] Combination variants (Table 12) were produced to combine the mutations K93C and E294C described both with and without the HSA C34A mutation. Briefly, plasmids comprising the individual mutations were prepared, and the mutations combined by restriction enzyme digestion and ligation.
[0515] 1 .mu.l of purified plasmid produced in Method 1 corresponding to the mutations K93C or E294C was transformed into E. coli NEB 5-alpha (New England Biolabs) and plated onto LB plates (as described above) supplemented with 50 .mu.g/mL ampicillin. Plasmids were isolated using a Qiagen Plasmid Plus Kit (Qiagen--according to manufacturer's instructions) and sequenced to confirm the presence of the desired mutation within the HSA sequence. These plasmids were named pDB5623 (C34A+K93C) and pDB5624 (C34A+E294C).
[0516] A fragment was removed from plasmid pDB5624 using the NheI and SphI restriction sites and was purified using a QIAquick Gel Extraction Kit (Qiagen) and ligated into pDB5623 digested with the same enzymes to produce construct pDB5625. pDB5626 and pDB5627 were constructed by insertion of the fragment produced by digestion of pDB5102 with SacII and PstI restriction enzymes into similarly digested pDB5623 and pDB5624. pDB5102 is described in WO 2015/036579 (incorporated herein by reference). The ligated plasmids were all transformed into E. coli NEB 5-alpha and plated onto LB plates (as described above) supplemented with 50 .mu.g/mL ampicillin. Plasmids were isolated using a Qiagen Plasmid Plus Kit (Qiagen--according to manufacturer's instructions) and sequenced to confirm the presence of the desired mutation within the HSA sequence.
[0517] To produce pDB5628 a fragment was removed from plasmid pDB5102 using the SacII and PstI restriction sites and was purified using a QIAquick Gel Extraction Kit (Qiagen) and ligated into pDB5625 digested with the same enzymes. The ligated plasmids were all transformed into E. coli NEB 5-alpha and plated onto LB plates supplemented with 50 .mu.g/mL ampicillin. Plasmids were isolated using a Qiagen Plasmid Plus Kit (Qiagen--according to manufacturer's instructions) and sequenced to confirm the presence of the desired mutation within the HSA sequence.
TABLE-US-00016 TABLE 12 Summary information for combination variants Protein Variant Number of thiols Plasmid SEQ ID NO. C34A + K93C 1 pDB5623 111 C34A + E294C 1 pDB5624 129 C34A + K93C + E294C 2 pDB5625 141 K93C 2 pDB5626 142 E294C 2 pDB5627 143 K93C + E294C 3 pDB5628 144
Production of Expression Plasmid and Yeast Stocks.
[0518] Preparation of the expression plasmids and transformation of S. cerevisiae was performed as described in WO 2012/150319 by the 48-hour stocking method (incorporated herein by reference). The host strain for the constructs was S. cerevisiae BXP10 Cir.degree. (WO 2015/036579, incorporated herein by reference). Purification of variants from shake flask was performed as described in WO 2012/150319 (incorporated herein by reference) unless otherwise stated.
Example 6. Production, Purification and Conjugation of Thio-Albumin Variants
[0519] Cryopreserved yeast stocks each in 1 mL aliquots were inoculated into separate shake flasks containing 100 mL BMMS growth medium (yeast nitrogen base without amino acids or (NH.sub.4).sub.2SO.sub.4, Difco 1.7 g/L; citric acid monohydrate 6.09 g/L; Na.sub.2HPO.sub.4.2H.sub.2O 25.27 g/L; (NH.sub.4).sub.2SO.sub.4 5.0 g/L; pH 6.5.+-.0.2; sucrose added to 20 g/L). Cells were transferred from the shake flask to the fermenter (10 L working volume, Sartorius Biostat C 10-3 fermenter) when the concentration of cells in the shake flask reached 0.8-1.2 mg/mL achieving a cell inoculum concentration of .gtoreq.10 mg/L (greater than or equal to 10 mg/L) in the fermenter.
[0520] The thio-albumin variants were produced by axenic culture of each of the yeast strains in high cell density (HCD) fed-batch fermentation. The aim of the fermentation was to achieve maximum biomass and productivity by controlling feed rate addition so that formation of by-products such as ethanol and acetate were avoided. Further details of the fermentation process are described in WO 96/37515 (incorporated herein by reference). The temperature and pH were controlled at 30.degree. C. and pH 6.2 respectively. Culture supernatant was harvested by centrifugation using a Sorvall RC 3C centrifuge (DuPont) to provide materials for immediate purification and the remaining materials were frozen (-20.degree. C.) for storage, before being thawed for subsequent purifications. Final product concentrations were determined by GP-HPLC using a LC2010 HPLC system (Shimadzu) equipped with UV detection under Shimadzu VP7.3 client server software control as described in Example 3. Table 13 provides the yields of each thio-albumin variant (in mg/mL culture supernatant) and shows that high product titres of greater than 1 mg/mL culture supernatant were obtained in all cases.
TABLE-US-00017 TABLE 13 Thio-albumin variant protein concentration by GP-HPLC Concentration Number of by GPHPLC Sample Description thiols (mg/mL) SEQ ID NO. C34A + K93C 1 3.1 111 C34A + E294C 1 4.6 129 C34A + K93C + E294C 2 2.3 141 K93C 2 1.8 142 E294C 2 3.9 143 K93C + E294C 3 1.6 144
[0521] The variants were purified at scale by a two-step chromatography process. The first purification step was using AlbuPure.RTM. chromatography as previously described in Example 3 but washing the column with approximately 4 column volumes of 50 mM sodium acetate, pH 5.3, 10 column volumes of 50 mM sodium phosphate, pH 8.0, and 10 column volumes of 50 mM ammonium acetate pH 8.0 respectively. Bound protein was eluted using between 1 and 3 column volumes of 50 mM ammonium acetate, 10 mM sodium octanoate, pH 7.0. The AlbuPure.RTM. eluates were then further purified using ion exchange chromatography via DE-FF as described in Evans et al. (2010), Protein Expression and Purification Volume 73, Issue 2, Pages 113-124. Post purification, the DE-FF eluate samples were concentrated and buffer exchanged by ultrafiltration/diafiltration using 10,000 molecular weight cut-off Vivacell 100 centrifugal concentrators (Sartorius). The samples were centrifuged at 2,000.times.g for 30 minutes (multiple times) to reduce the volume to below 10 mL before diafiltration against 10 volumes of 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5. Post diafiltration, sample concentrations were in the range of 124 to 177 mg/mL. The samples were diluted to a final formulation concentration of 50 mg/mL in 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5.
[0522] The thio-albumin variants were conjugated with maleimide-PEG2-biotin as described in Example 4, but with a 3.2-fold molar excess of biotin over the free thiol content (number of free thiols). Due to some variants having multiple free thiol sites available for conjugation, the expected molecular weights for all biotin conjugation permutations are summarised in Table 14. The variants with two or three thiol groups increased by 2.times.525 Da, and 3.times.525 Da respectively. The relative peak heights of each peak species were used to calculate the percentage of target conjugate, i.e. the correct percentage of a single, double or triple biotin labelled thio-albumin variant. The K93C+E294C variant had a total of 3 free thiol residues, the MS spectrum for this variant is shown in FIG. 7A. It was evident from the single peak species on the MS spectrum that the variant has successfully conjugated 3 moles of maleimide-PEG2-biotin per mole protein, as indicated by a mass increase of 1575 Da (3.times.525 Da) to 67968 Da (Table 15). The samples were incubated at 37.degree. C., pH 9, for at least 24 hours for controlled hydrolysis to occur to determine the stability of the thio ether conjugate bond as previously described in Example 4. The results are summarised in Table 15. The yield of the hydrolysed thiol stable K93C+E294C conjugate was in the order of 20% triple conjugate, due to the competing retro-Michael deconjugation of the C34 conjugate during hydrolysis (FIG. 7B). The main species was now a hydrolysed thiol stable double conjugate with a mass of 67476 Da indicating that hydrolysis had occurred to the double conjugate species. It was evident that the variants containing a cysteine at position C34 had significant deconjugation during hydrolysis compared to the variants with a C34A mutation. The double thiol variant C34A+K93C+E294C was 62% double conjugated pre hydrolysis and 56% post hydrolysis. An observed peak species with a mass 66443 Da confirmed that hydrolysis had occurred with minimal conjugate loss (FIG. 7C) compared to the K93C+E294C conjugate which contained a cysteine at C34 (FIG. 7B) highlighting that the K93C and E294C variants had improved conjugate stability when using a maleimide linker.
TABLE-US-00018 TABLE 14 Expected molecular weights post conjugation and hydrolysis Single Double Triple conjugate Mr conjugate Mr conjugate Mr No. Free +biotin Hydro- +2x Hydro- +3x Hydro- Sample thiols Mr (525 Da) lysed biotin lysed biotin lysed C34A + K93C 1 66382 66907 66925 n/a n/a n/a n/a C34A + E294C 1 66381 66906 66924 n/a n/a n/a n/a C34A + K93C + 2 66356 66881 66899 67406 67442 n/a n/a E294C K93C 2 66414 66939 66957 67464 67500 n/a n/a E294C 2 66413 66938 66956 67463 67499 n/a n/a K93C + E294C 3 66388 66913 66931 67438 67474 67963 68017 n/a: not applicable
TABLE-US-00019 TABLE 15 Conjugation efficiency and controlled hydrolysis results Post conjugation Post hydrolysis Conjugate Conjugate Number intact % target intact % target Sample of mass con- mass con- Description thiols result (Da) jugate result (Da) jugate C34A + K93C 1 66910 68 66927 70 C34A + E294C 1 66908 52 66927 46 C34A + 2 67410 62 67443 56 K93C + E294C K93C 2 67467 99 67502 36 E294C 2 67467 87 67501 40 K93C + E294C 3 67968 98 68020 20
[0523] The formulated samples were subjected to a six month stability assessment at 2-8.degree. C. by GPHPLC, using the method described in Example 3. The percentage monomer (in brackets) was determined for each sample relative to its wild type control under the same storage conditions. The percentage monomer results are summarized in Table 16, and indicated that aggregation levels were within acceptable limits when the albumin variants were formulated at 50 mg/mL and stored for six months at 2-8.degree. C.
TABLE-US-00020 TABLE 16 GPHPLC protein stability assessment at 50 mg/mL, post storage at 2-8.degree. C. .DELTA. % Protein Sample Number % Monomer at Monomer SEQ Description of thiols T = 0 T = 1 m T = 2 m T = 3 m T = 6 m 0-6 month ID NO. WT control 1 93.8 (100) 94.5 (100) 94.5 (100) 94.4 (100) 94.9 (100) 1.1 2 C34A + K93C 1 92.1 (98) 91.2 (97) 90.2 (95) 89.3 (95) 88.1 (93) -4.0 111 C34A + E294C 1 92.4 (99) 92.7 (98) 91.6 (97) 91.0 (96) 90.8 (96) -1.6 129 C34A + K93C + 2 84.4 (90) 81.8 (87) 79.7 (84) 78.4 (83) 75.9 (80) -8.5 141 E294C K93C 2 88.2 (94) 86.9 (92) 85.5 (91) 84.9 (90) 84.2 (89) -4.0 142 E294C 2 89.3 (95) 90.2 (95) 89.7 (95) 89.0 (94) 88.9 (94) -0.4 143 K93C + E294C 3 82.1 (88) 79.8 (84) 78.4 (83) 77.3 (82) 76.8 (81) -5.3 144 m = month
Example 7: Combination Variants Having Altered FcRn Binding
[0524] HSA having the K573P substitution, as described in WO 2011/051489 (incorporated herein by reference), has a higher affinity for FcRn than does wild type HSA. Constructs were produced to combine the mutations in the variants described in Table 12 with the HSA K573P mutation from plasmid pDB4673.
Method 3.
[0525] A fragment was removed from plasmids pDB5623, 5624, 5625, 5626, 5627 and 5628 using the PstI and XhoI restriction sites and was purified using a QIAquick Gel Extraction Kit (Qiagen) and ligated into pDB4673 digested with the same enzymes to produce constructs pDB5704, 5707, 5710, 5713, 5716 and 5719 (Table 17). The ligated plasmids were all transformed into E. coli NEB 5-alpha (New England Biolabs) and plated onto LB plates supplemented with 50 .mu.g/mL ampicillin. Plasmids were isolated using a Qiagen Plasmid Plus Kit (Qiagen--according to manufacturer's instructions) and sequenced to confirm the presence of the desired mutations within the HSA sequence.
TABLE-US-00021 TABLE 17 Summary information for variants having altered FcRn binding Variant Plasmid Protein SEQ ID NO. K573P pDB4673 145 C34A + K93C + K573P pDB5704 146 C34A + E294C + K573P pDB5707 147 C34A + K93C + E294C + K573P pDB5710 148 K93C + K573P pDB5713 149 E294C + K573P pDB5716 150 K93C + E294C + K573P pDB5719 151
Production of Expression Plasmid and Yeast Stocks.
[0526] Preparation of the expression plasmids and transformation of S. cerevisiae was performed as described in WO 2012/150319 by the 48-hour stocking method (incorporated herein by reference). The host strain for the constructs was S. cerevisiae BXP10 Cir.degree. (WO 2015/036579, incorporated herein by reference)). Purification of variants from shake flask was performed as described in WO 2012/150319 (incorporated herein by reference) unless otherwise stated.
Example 8. Aggregation Screening of Combination Variants Having Altered FcRn Binding
[0527] Shake flask culturing of S. cerevisiae and purification was performed as described in Example 3. A single step AlbuPure chromatography procedure was used to prepare purified material from 6 variants as described in Example 3. Post purification the 20 mL eluates were concentrated to less than 200 .mu.L using Vivaspin centrifugal concentrators as described in Example 4. Post concentration the samples were buffer exchanged by the addition of 10 mL of 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5 and the samples centrifuged as before. The final volumes recovered were between 75 .mu.L and 200 .mu.L. The concentration and percentage monomer of the eluate samples was determined by Gel Permeation High Pressure Liquid Chromatography (GP-HPLC) as described in Example 3. The results are summarised in Table 18. Final product concentrations were in the range of 47 to 154 mg/mL. A typical wild type albumin control in Example 4 resulted in a monomer percentage of 87% at 1.1 mg/mL (Table 8A). All variants analysed had monomer percentages equal to or greater than 87% even at significantly higher protein concentrations. This indicated that all variants had minimal propensity to aggregate.
TABLE-US-00022 TABLE 18 GPHPLC aggregation screening results GPHPLC monomer concentration % Monomer Sample description (mg/mL) at T = 0 C34A + K93C + K573P 48.2 91.8 C34A + E294C + K573P 153.5 90.8 C34A + K93C + E294C + 98.8 88.4 K573P K93C + K573P 101.7 86.9 E294C + K573P 115.5 87.6 K93C + E294C + K573P 46.5 91.2
Example 9. Conjugation Efficiency and Controlled Hydrolysis of Combination Variants Having Altered FcRn Binding
[0528] The thio-albumin combination variants (Table 17) were conjugated with a 3.2 fold excess of maleimide-PEG2-biotin as described in Example 6. Due to some variants having multiple free thiol sites available for conjugation, the expected molecular weights for all biotin conjugation permutations are summarised in Table 19.
TABLE-US-00023 TABLE 19 Expected molecular weights of albumin variants post conjugation and hydrolysis Single Double Triple conjugate Mr conjugate Mr conjugate Mr No. Free +biotin Hydro- +2x Hydro- +3x Hydro- Sample thiols Mr (525 Da) lysed biotin lysed biotin lysed C34A + K93C + 1 66351 66876 66894 n/a n/a n/a n/a K573P C34A + E294C + 1 66350 66875 66893 n/a n/a n/a n/a K573P C34A + K93C + 2 66325 66850 66868 67375 67411 n/a n/a E294C + 573P K93C + K573P 2 66383 66908 66926 67433 67469 n/a n/a E294C + K573P 2 66382 66907 66925 67432 67468 n/a n/a K93C + E294C + 3 66357 66882 66900 67407 67443 67932 67986 K573P n/a: not applicable
[0529] The molecular weight of the variants with two or three thiol groups increased by 2.times.525 Da, and 3.times.525 Da respectively. The relative peak heights of each peak species were used to calculate the percentage of target conjugate, i.e. the percentage of a single, double or triple biotin labelled thio-albumin variant. The K93C+E294C+K573P variant had a total of 3 free thiol residues (the third thiol being provided by native Cys34); the MS spectrum for this variant is shown in FIG. 8A. It was evident from the single peak species on the MS spectrum that the variant has successfully conjugated with 3 moles of maleimide-PEG2-biotin per mole protein, as indicated by a mass increase of 1575 Da (3.times.525 Da) to 67940.8 Da. The samples were incubated at 37.degree. C., pH 9, for at least 18 hours for controlled hydrolysis to occur to determine the stability of the thio ether conjugate bond as previously described in Example 4. The results are summarized in Table 20.
[0530] The yield of the hydrolysed thiol stable K93C+E294C+K573P conjugate was in the order of 23% triple conjugate, likely due to the competing retro-Michael deconjugation of the C34 conjugate during hydrolysis (FIG. 8B). The main species was now a hydrolysed thiol stable double conjugate with a mass of 67447.3 Da indicating that hydrolysis had occurred to this double conjugate species. It was evident that the variants containing a cysteine at position C34 underwent more pronounced deconjugation during hydrolysis compared to the variants with a C34A mutation.
TABLE-US-00024 TABLE 20 Conjugation efficiency and controlled hydrolysis results Post conjugation Post hydrolysis Conjugate Conjugate Number intact % target intact % target Sample of mass con- mass con- Description thiols result (Da) jugate result (Da) jugate C34A + K93C + 1 66878 100 66896 100 K573P C34A + E294C + 1 66880 85 66897 89 K573P C34A + K93C + 2 67382 90 * * E294C + K573P K93C + K573P 2 67438 100 67473 32 E294C + K573P 2 67441 100 67474 25 K93C + E294C + 3 67941 100 67989 23 K573P * low intensity MS spectrum, unable to accurately quantify data
Example 10. Surface Plasmon Resonance (SPR) Analysis of Combination Variants Having Altered FcRn Binding, Pre and Post Conjugation with Maleimide-PEG2-Biotin
[0531] Thio-albumin combination variants detailed in Tables 12 and 17 were produced by fed-batch fermentation and purified according to Example 6. Post purification, the samples were concentrated and the buffer was exchanged against a minimum of 7 continuous volumes of 25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5 using 10,000 molecular weight cut-off Centramate Tangential Flow Filtration Membrane cassettes (PALL) before final formulation at 20 mg/mL in buffer (25 mM sodium phosphate, 215 mM sodium chloride, pH 6.5). Subsequently, a size exclusion chromatography step (Sephacryl.RTM. 5200, GE Healthcare) was performed. For each sample 25 mL was split equally between two Vivaspin 20 centrifugal concentrators and centrifuged at 4,500.times.g for two 20 minute time periods to reduce the total volume to 5 mL. The concentrated material was loaded onto a 483 mL S200 column and the monomer peak collected to generate monomeric protein at greater than 98% for FcRn binding analysis by SPR. Post purification, eluates were diluted to 5 mg/mL (.+-.5%). The binding affinity of each variant for the human FcRn receptor was determined both pre and post conjugation with maleimide-PEG2-biotin. Variants were conjugated with a 3.2 fold excess of maleimide-PEG2-biotin as described in Example 6. The percentage conjugation was determined by MS as described in Example 2, but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The results are shown in Table 21 and indicated all samples had conjugated to varying extent, depending on the number of thiols.
TABLE-US-00025 TABLE 21 Conjugation efficiency for samples for SPR Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 0 100 n/a n/a 2 C34A + K93C 1 0 100 n/a n/a 111 C34A + E294C 1 74 26 n/a n/a 129 C34A + K93C + 2 0 74 26 n/a 141 E294C K93C 2 0 26 74 n/a 142 E294C 2 0 81 19 n/a 143 K93C + E294C 3 0 0 80 20 144 K573P 1 8 93 n/a n/a 145 C34A + K93C + 1 0 100 n/a n/a 146 K573P C34A + E294C + 1 20 80 n/a n/a 147 K573P C34A + K93C + 2 0 10 90 n/a 148 E294C + K573P K93C + K573P 2 0 0 100 n/a 149 E294C + K573P 2 0 18 82 n/a 150 K93C + E294C + 3 0 0 40 60 151 K573P n/a: not applicable
[0532] SPR analyses were carried out using a Biacore 3000 instrument (GE Healthcare). Flow cells of CM5 sensor chips were coupled with soluble human FcRn (1200-1600 RU) using amine coupling chemistry as described in the protocol provided by the manufacturer (GE Healthcare). The coupling was performed by injecting 5 .mu.g/mL of the protein in 10 mM sodium acetate pH 4.5 (GE healthcare). Phosphate buffer (67 mM phosphate buffer, 0.15 M NaCl, 0.005% Tween 20) at pH 5.5 was used as running buffer and dilution buffer. Regeneration of the surfaces were performed using injections of HBS-EP buffer (0.01 M HEPES, 0.15 M NaCl, 3 mM EDTA, 0.005% surfactant P20) at pH 7.4 (GE Healthcare). Post immobilisation, the chip was left to stabilise with a constant flow (5 .mu.L/min) of running buffer. Chip surface was conditioned by injecting 3.times. injections of running buffer followed by 3.times. injections of regeneration buffer. Surfaces were checked for activity with native sequence HSA control. For determination of binding kinetics, serial dilutions of albumin variants (10-0 .mu.M) were injected over immobilized receptor at a constant flow rate (30 .mu.L/min) at 25.degree. C. In all analyses, data were zero adjusted and the reference cell subtracted. Data evaluations were performed using BIAevaluation 4.1 software (BIAcore AB). The results pre and post conjugation are shown in Tables 22 and 23 respectively.
[0533] The thio-albumin variants screened over human FcRn bound to the receptor in a reversible, pH-dependent manner.
[0534] The thio-albumin variants in a wild type background (i.e. the only amino acid alterations were those that were introduced to affect the number of conjugatable cysteine residues) gave a similar fold increase in binding affinity to FcRn compared to the wild type control (SEQ ID NO. 2) both pre and post conjugation. The thio-albumin variants which also included a K573P mutation (to increase the affinity of the albumin variant to FcRn) maintained their increase in FcRn affinity, compared to the K573P control (SEQ ID NO. 145), pre and post conjugation indicating that neither the change in conjugatable cysteine residues nor the conjugation partner had an observable influence on the binding affinity of the albumin variant to FcRn.
TABLE-US-00026 TABLE 22 FcRn affinity for variants pre conjugation Protein Sample Number Ka Kd KD SEQ Description of thiols (10.sup.3/Ms) (10.sup.3/Ms) (.mu.M) Fold > WT ID NO. WT control 1 7.38 54.0 7.32 n/a 2 C34A + K93C 1 12.4 44.9 3.62 2.02 111 C34A + E294C 1 6.13 46.7 7.61 0.96 129 C34A + K93C + E294C 2 10.41 39.5 3.79 1.93 141 K93C 2 8.19 43.45 5.30 1.38 142 E294C 2 5.0 46.3 9.25 0.79 143 K93C + E294C 3 7.65 37.9 4.95 1.48 144 K573P 1 5.70 3.76 0.66 11.10 145 C34A + K93C + K573P 1 8.06 3.83 0.48 15.25 146 C34A + E294C + K573P 1 6.07 3.96 0.65 11.26 147 C34A + K93C + E294C + K573P 2 8.11 3.67 0.45 16.27 148 K93C + K573P 2 8.37 4.07 0.48 15.25 149 E294C + K573P 2 5.65 4.17 0.74 9.89 150 K93C + E294C + K573P 3 6.8 3.82 0.56 13.07 151 n/a: not applicable Fold > WT = KD (.mu.M) WT control/KD (.mu.M) variant
TABLE-US-00027 TABLE 23 FcRn affinity for samples post conjugation with maleimide-PEG2-biotin Protein Sample Number Ka Kd KD SEQ Description of thiols (10.sup.3/Ms) (10.sup.3/Ms) (.mu.M) Fold > WT ID NO. WT control 1 9.26 25.4 2.74 n/a 2 C34A + K93C 1 12.95 19.85 1.53 1.79 111 C34A + E294C 1 8.07 20.15 2.50 1.09 129 C34A + K93C + E294C 2 11.55 17.4 1.51 1.80 141 K93C 2 10.2 21.0 2.06 1.33 142 E294C 2 9.38 19.9 2.12 1.29 143 K93C + E294C 3 11.8 19.3 1.63 1.68 144 K573P 1 9.25 3.12 0.337 8.13 145 C34A + K93C + K573P 1 14.35 2.97 0.207 13.24 146 C34A + E294C + K573P 1 10.42 2.97 0.285 9.61 147 C34A + K93C + E294C + K573P 2 13.7 2.71 0.198 13.84 148 K93C + K573P 2 13.7 2.93 0.214 12.80 149 E294C + K573P 2 11.4 3.22 0.283 9.68 150 K93C + E294C + K573P 3 13.05 3.31 0.254 10.79 151 n/a: not applicable Fold > WT = KD (.mu.M) WT control/KD (.mu.M) variant
Example 11. Aggregation Analysis of Combination Variants Having Altered FcRn Binding
[0535] The thio-albumin combination variants formulated at 5 mg/mL in Example 10 were analysed for their tendency to remain as a monomer in solution. WT HSA, the variant K573P and the variant C34A+L302C were prepared as described in Example 10 and included as controls. The free thiol content for each variant was determined at T=0 and following 3 months storage at 5.degree. C. by mass spectrometric analysis of protein sample treated with DTNB reagent, similar to the method of Example 2. For this example, 80 .mu.L of each variant sample was diluted with 920 .mu.L of buffer 1 (100 mM Tris-HCl, 10 mM EDTA, pH 8.0). To each variant sample, 50 .mu.L of buffer 2 (4 mg/mL DTNB, 500 mM sodium phosphate, pH 7.0) was added. The resultant preparation incubated for at least 25 minutes at ambient temperature (20.+-.5.degree. C.) to allow TNB labelling. Post incubation, the samples were subjected to mass spectrometry to determine the intact protein mass post conjugation as per the method described in Example 2, but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The results are summarised in Table 24A and Table 24B. An increase in mass of 197 Da upon DTNB incubation is indicative of the presence of one free thiol group on the protein in the sample. An increase of 394 Da and 591 Da is indicative of two or three free thiol groups respectively. All samples had high levels of free thiol at the start of the stability study, and the majority maintained a high free thiol level following 3 months storage at 5.degree. C.
TABLE-US-00028 TABLE 24A Mass Spectrometry DTNB free thiol results Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 0 91 0 9 2 C34A + L302C 1 0 100 0 0 152 C34A + K93C 1 0 91 0 9 111 C34A + E294C 1 6 94 0 0 129 C34A + K93C + 2 0 39 61 0 141 E294C K93C 2 16 0 84 0 142 E294C 2 0 28 84 0 143 K93C + E294C 3 0 0 51 49 144 K573P 1 0 91 0 9 145 C34A + K93C + 1 0 93 0 7 146 K573P C34A + E294C + 1 7 87 0 6 147 K573P C34A + K93C + 2 0 5 89 0 148 E294C + K573P K93C + K573P 2 0 0 92 0 149 E294C + K573P 2 0 7 87 0 150 K93C + E294C + 3 0 0 22 78 151 K573P
TABLE-US-00029 TABLE 24B Mass Spectrometry DTNB free thiol results, post storage at 5.degree. C. Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 0 94 6 0 2 C34A + L302C 1 0 100 0 0 152 C34A + K93C 1 0 95 0 5 111 C34A + E294C 1 55 45 0 0 129 C34A + K93C + 2 0 54 46 0 141 E294C K93C 2 23 0 77 0 142 E294C 2 0 50 50 0 143 K93C + E294C 3 0 0 73 27 144 K573P 1 0 94 0 6 145 C34A + K93C + 1 0 95 0 5 146 K573P C34A + E294C + 1 14 86 0 0 147 K573P C34A + K93C + 2 0 10 90 0 148 E294C + K573P K93C + K573P 2 0 0 94 0 149 E294C + K573P 2 0 13 87 0 150 K93C + E294C + 3 0 0 32 68 151 K573P
[0536] Samples were stored for 3 months at 5.degree. C. and 40.degree. C. and the aggregation profile determined at various time points by GPHPLC as described in Example 3. The percentage monomer (in brackets) was determined for each sample relative to its wild type control under the same storage conditions. The results for 5.degree. C. and 40.degree. C. are provided in Tables 25 and 26 respectively. All variants had a monomer greater than 98% at T=0. The majority of thio-albumin variants maintained a monomeric protein percentage equal to or greater than 97% during 3 month's storage at 5.degree. C. Relative to the other variants analysed, variant C34A+L302C was more prone to aggregation. It was also evident that the majority of thio-albumin variants maintained a monomeric protein percentage equal to or greater than 80% during 3 months storage at 40.degree. C., even those containing two or three thiol residues. However, it was evident that variant C34A+L302C was more prone to aggregation with a monomer percentage of 73.2% following 3 months storage at 40.degree. C.
TABLE-US-00030 TABLE 25 GPHPLC protein stability assessment at 5 mg/mL, post storage at 5.degree. C. % Monomer at Protein Sample Number T = 1 T = 2 T = 3 SEQ Description of thiols T = 0 month month month ID NO. WT control 1 99.8 (100) 99.3 (100) 99.7 (100) 99.7 (100) 2 C34A + L302C 1 98.3 (99) 95.5 (96) 94.9 (95) 95.0 (95) 152 C34A + K93C 1 99.5 (100) 99.2 (100) 98.9 (99) 98.7 (99) 111 C34A + E294C 1 99.5 (100) 99.4 (100) 99.2 (100) 99.2 (100) 129 C34A + K93C + 2 99.1 (99) 98.5 (99) 97.9 (98) 97.5 (98) 141 E294C K93C 2 99.5 (100) 99.3 (100) 99.0 (99) 98.8 (99) 142 E294C 2 99.6 (100) 99.4 (100) 99.2 (100) 99.1 (99) 143 K93C + E294C 3 99.2 (99) 98.8 (100) 98.3 (99) 97.9 (98) 144 K573P 1 99.7 (100) 99.7 (100) 99.6 (100) 98.5 (99) 145 C34A + K93C + 1 99.6 (100) 99.2 (100) 99.0 (99) 98.6 (99) 146 K573P C34A + E294C + 1 99.4 (100) 99.1 (100) 98.8 (99) 98.5 (99) 147 K573P C34A + K93C + 2 99.3 (100) 98.5 (99) 97.7 (98) 96.9 (97) 148 E294C + K573P K93C + K573P 2 99.8 (100) 99.7 (100) 99.6 (100) 98.5 (100) 149 E294C + K573P 2 99.7 (100) 98.9 (99) 99.0 (99) 98.8 (99) 150 K93C + E294C + 3 99.5 (100) 99.1 (100) 98.8 (99) 98.5 (99) 151 K573P
TABLE-US-00031 TABLE 26 GPHPLC protein stability assessment at 5 mg/mL, post storage at 40.degree. C. % Monomer at Protein Sample Number T = 0.5 T = 1 T = 2 T = 3 SEQ Description of thiols T = 0 month month month month ID NO. WT control 1 99.8 (100) 99.4 (100) 99.3 (100) 99.0 (100) 97.6 (100) 2 C34A + L302C 1 98.3 (99) 87.6 (88) 80.9 (82) 75.6 (76) 73.2 (75) 152 C34A + K93C 1 99.5 (100) 96.1 (97) 93.4 (94) 90.3 (91) 86.6 (89) 111 C34A + E294C 1 99.5 (100) 98.6 (99) 96.6 (97) 96.0 (97) 95.4 (98) 129 C34A + K93C + 2 99.1 (99) 93.8 (94) 89.3 (90) 85.1 (86) 80.4 (82) 141 E294C K93C 2 99.5 (100) 96.6 (97) 94.5 (95) 90.4 (91) 88.8 (91) 142 E294C 2 99.6 (100) 98.1 (99) 95.7 (96) 95.1 (96) 94.2 (97) 143 K93C + E294C 3 99.2 (99) 93.9 (95) 89.3 (90) 82.9 (84) 80.0 (82) 144 K573P 1 99.7 (100) 99.3 (100) 99.0 (100) 98.7 (100) 97.4 (100) 145 C34A + K93C + 1 99.6 (100) 95.5 (96) 93.0 (94) 89.5 (90) 86.5 (89) 146 K573P C34A + E294C + 1 99.4 (100) 97.3 (98) 95.9 (97) 95.0 (96) 94.4 (97) 147 K573P C34A + K93C + 2 99.3 (100) 91.8 (92) 88.3 (89) 82.7 (84) 80.9 (83) 148 E294C + K573P K93C + K573P 2 99.8 (100) 98.1 (99) 96.7 (97) 94.5 (96) 92.6 (95) 149 E294C + K573P 2 99.7 (100) 97.5 (98) 95.8 (97) 94.5 (96) 94.4 (97) 150 K93C + E294C + 3 99.5 (100) 94.9 (96) 92.0 (93) 88.7 (90) 84.1 (86) 151 K573P
Example 12. Conjugation of Combination Variants Having Altered FcRn Binding, with Fluorescent Probes
[0537] Thio-albumin combination variants formulated at 5 mg/mL in Example 10, following 6 weeks storage at 2-8.degree. C., were conjugated using a 3-fold excess of Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye (FIG. 9A) or 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye (FIG. 10A) (Almac Group Ltd., UK, custom synthesis). Variants were diluted with PBS buffer, pH 7.4 to give 1 mL solutions at 1 mg/mL (15.05 nmol). A 1 mg/mL stock solution of Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye was prepared by reconstituting 1.6 mg material with 1.6 mL PBS buffer pH 7.4. From the Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye stock solution, 51.5 .mu.L (45.15 nmol) was added to the single thiol variants, 103 .mu.L (90.3 nmol) dye stock solution was added to the double thiol variants, and 154.5 .mu.L (135.3 nmol) dye stock solution was added to the triple thiol variants to give a threefold excess of Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye over the number of free thiols. A 0.5 mg/mL stock solution of 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye was prepared by reconstituting 1.7 mg material with 1.7 mL dimethyl sulfoxide (DMSO) and 1.7 mL PBS pH 7.4 buffer. From the 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye stock solution 44.3 .mu.L (45.15 nmol) was added to the single thiol variants, 88.6 .mu.L (90.3 nmol) dye stock solution was added to the double thiol variants, and 132.9 .mu.L (135.3 nmol) dye stock solution was added to the triple thiol variants to give a threefold excess of 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye over the number of free thiols. Samples were gently mixed and incubated at ambient temperature overnight in the dark. Post incubation the samples were analysed by mass spectrometry to determine the intact protein mass post conjugation as per the MS method described in Example 2, but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The results are summarised in Table 27 and Table 28.
[0538] The MS spectrum for the altered FcRn binding variant K573P shown in FIG. 9B, exhibited a single species at 67468.5 Da indicating the correct molecular weight for a K573P variant plus a single addition of Alexa Fluor.RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye (+1058 Da). This confirmed the variant had a single free thiol located at Cys34 available for conjugation. The thio-albumin variant K93C+E294C+K573P shown in FIG. 9C indicated that conjugation had occurred post an overnight incubation, giving approximately 42% diconjugate and 58% triconjugate species respectively, when comparing the relative peak heights of conjugated species. It was evident that the main peak species had increased by approximately 3174 Da (3.times.1058 Da) to 69536 Da. This indicated the variant had three free thiols available for conjugation.
[0539] The MS spectrum for the altered FcRn binding variant K573P shown in FIG. 10B, exhibited a single species at 67310.6 Da indicating the correct molecular weight for a K573P variant plus a single addition of 5-carboxyfluorescein-PEG4-Lys(monobromomaleimide)-NH2 dye (+901 Da). The thio-albumin variant C34A+K93C+E294C+K573P shown in FIG. 10C indicated that conjugation had occurred post an overnight incubation, giving approximately 9% monoconjugate and 91% diconjugate species respectively, when comparing the relative peak heights of conjugated species. It was evident that the main peak species had increased by approximately 1802 Da (2.times.901 Da) to 68129.7 Da. This indicated the variant had two free thiols available for conjugation.
TABLE-US-00032 TABLE 27 Conjugation efficiency results of thio-albumin variants with Alexa Fluor .RTM. 488-PEG4-Lys(monobromomaleimide)-NH2 dye Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 0 93 n/a n/a 2 C34A + K93C 1 0 100 n/a n/a 111 C34A + E294C 1 100 0 n/a n/a 129 C34A + K93C + 2 * * * n/a 141 E294C K93C 2 17 0 83 n/a 142 E294C 2 0 89 11 n/a 143 K93C + E294C 3 * * * * 144 K573P 1 0 100 n/a n/a 145 C34A + K93C + 1 0 100 n/a n/a 146 K573P C34A + E294C + 1 8 92 n/a n/a 147 K573P C34A + K93C + 2 0 7 93 n/a 148 E294C + K573P K93C + K573P 2 0 0 91 n/a 149 E294C + K573P 2 0 0 100 n/a 150 K93C + E294C + 3 0 0 42 58 151 K573P n/a: not applicable * low intensity MS spectrum, unable to accurately quantify data
TABLE-US-00033 TABLE 28 Conjugation efficiency results of thio-albumin variants with 5-carboxyfluorescein-PEG4-Lys(Bromomaleimide)-NH2 dye Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 0 96 n/a n/a 2 C34A + K93C 1 0 100 n/a n/a 111 C34A + E294C 1 100 0 n/a n/a 129 C34A + K93C + 2 * * * n/a 141 E294C K93C 2 30 0 70 n/a 142 E294C 2 * * * n/a 143 K93C + E294C 3 * * * 144 K573P 1 0 100 n/a n/a 145 C34A + K93C + 1 0 100 n/a n/a 146 K573P C34A + E294C + 1 18 82 n/a n/a 147 K573P C34A + K93C + 2 0 9 91 n/a 148 E294C + K573P K93C + K573P 2 1 0 99 n/a 149 E294C + K573P 2 * * * n/a 150 K93C + E294C + 3 * * * * 151 K573P n/a: not applicable * low intensity MS spectrum, unable to accurately quantify data
Example 13. Conjugation of Combination Variants Having Altered FcRn Binding, with Paclitaxel
[0540] Thio-albumin combination variants formulated at 5 mg/mL in Example 10, following 3 months storage at 2-8.degree. C., were conjugated using a 1.5 fold excess of paclitaxel which was via an ester group activated with a monobromomaleimide moiety, as shown in FIG. 11A, resulting in the molecule monobromomaleimide-paclitaxel (Almac Group Ltd., UK custom synthesis). Variants were diluted with PBS buffer, pH 7.4 to give 1 mL solutions at 1 mg/mL (15.05 nmol). A 2 mg/mL stock solution of monobromomaleimide-paclitaxel was prepared by reconstituting 6.6 mg material with 3.3 mL DMSO. From the monobromomaleimide-paclitaxel stock solution, 12.24 .mu.L (22.58 nmol) was added to the single thiol variants, 24.47 .mu.L (45.15 nmol) stock solution was added to the double thiol variants, and 36.71 .mu.L (67.73 nmol) stock solution was added to the triple thiol variants to give a threefold excess of monobromomaleimide-paclitaxel over the number of free thiols. Samples were gently mixed and incubated at ambient temperature overnight. Post incubation the samples were subjected to mass spectrometry to determine the intact protein mass post conjugation as per the MS method described in Example 2, but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The results are summarised in Table 29.
[0541] The MS spectrum for the altered FcRn binding variant K573P shown in FIG. 11B indicated that conjugation had occurred post an overnight incubation, giving approximately 77% monoconjugated and 23% unconjugated species respectively, when comparing the relative peak heights of the protein species. It was evident that the main peak species at 67412.2 Da had increased by approximately 1004 Da due to a single addition of monobromomaleimide-paclitaxel. The MS spectrum for the thio-albumin variant K93C+E294C+K573P shown in FIG. 11C indicated that conjugation had occurred post an overnight incubation, giving approximately 6% monoconjugate, approximately 60% diconjugate and 30% triconjugate species respectively, when comparing the relative peak heights of conjugated species. It was evident that the main peak species had increased by approximately 2008 Da (2.times.1004 Da) to 68364.2 Da, with a 69383.7 Da species indicative of a 3012 Da triple addition.
TABLE-US-00034 TABLE 29 Conjugation efficiency results of thio-albumin variants with monobromomaleimide-paclitaxel Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 24 76 n/a n/a 2 C34A + K93C 1 50 50 n/a n/a 111 C34A + E294C 1 100 0 n/a n/a 129 C34A + K93C + 2 * * * n/a 141 E294C K93C 2 30 26 44 n/a 142 E294C 2 0 100 0 n/a 143 K93C + E294C 3 * * * 0 144 K573P 1 23 77 n/a n/a 145 C34A + K93C + 1 59 41 n/a n/a 146 K573P C34A + E294C + 1 34 66 n/a n/a 147 K573P C34A + K93C + 2 10 50 40 n/a 148 E294C + K573P K93C + K573P 2 8 40 52 n/a 149 E294C + K573P 2 0 18 68 n/a 150 K93C + E294C + 3 0 6 60 30 151 K573P n/a: not applicable * low intensity MS spectrum, unable to accurately quantify data
Example 14. Conjugation of Combination Variants Having Altered FcRn Binding, with Exenatide Peptide
[0542] Thio-albumin combination variants formulated at 5 mg/mL in Example 10, following 3 months storage at 2-8.degree. C., were conjugated using a 1.5 fold excess of monobromomaleimide-PEG2-exenatide peptide as shown in FIG. 12A (Almac Group Ltd., UK, custom synthesis). Variants were diluted with PBS buffer, pH 7.4 to give 1 mL solutions at 1 mg/mL (15.05 nmol). A 5 mg/mL stock solution of monobromomaleimide-PEG2-exenatide peptide was prepared by reconstituting 5 mg material with 1 mL PBS buffer pH 7.4. From the monobromomaleimide-PEG2-exenatide peptide stock solution, 21.17 .mu.L (22.58 nmol) was added to the single thiol variants, 42.35 .mu.L (45.15 nmol) peptide stock solution was added to the double thiol variants, and 63.52 .mu.L (67.73 nmol) peptide stock solution was added to the triple thiol variants to give a threefold excess of monobromomaleimide-PEG2-exenatide peptide over the number of free thiols. Samples were gently mixed and incubated at ambient temperature overnight. Post incubation the samples were subjected to mass spectrometry to determine the intact protein mass post conjugation as per the MS method described in Example 2, but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The results are summarised in Table 30.
[0543] The MS spectrum for the altered FcRn binding variant K573P shown in FIG. 12B indicated that conjugation had occurred post an overnight incubation, giving approximately 33% monoconjugate and 67% unconjugated species respectively, when comparing the relative peak heights of protein species. It was evident that the main peak species at 66409.2 Da was unconjugated K573P variant. The second species had increased by approximately 4609 Da due to a single addition of monobromomaleimide-PEG2-exenatide peptide. The thio-albumin variant C34A+K93C+E294C+K573P shown in FIG. 12C indicated that conjugation had occurred post an overnight incubation, giving approximately 33% diconjugate species, approximately 45% monoconjugate species, and approximately 22% unconjugated species respectively, when comparing the relative peak heights of protein species. It was evident that the main peak species had increased by approximately 4609 Da to 70941.7 Da, with a 75557.3 Da species indicative of a 9218 Da addition representing a double conjugation of monobromomaleimide-PEG2-exenatide peptide.
TABLE-US-00035 TABLE 30 Conjugation efficiency results of thio-albumin variants with exenatide peptide Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 71 29 n/a n/a 2 C34A + K93C 1 74 26 n/a n/a 111 C34A + E294C 1 100 0 n/a n/a 129 C34A + K93C + 2 * * * n/a 141 E294C K93C 2 79 0 21 n/a 142 E294C 2 * * * n/a 143 K93C + E294C 3 * * * * 144 K573P 1 67 33 n/a n/a 145 C34A + K93C + 1 74 26 n/a n/a 146 K573P C34A + E294C + 1 51 49 n/a n/a 147 K573P C34A + K93C + 2 22 45 33 n/a 148 E294C + K573P K93C + K573P 2 60 0 39 n/a 149 E294C + K573P 2 21 33 47 n/a 150 K93C + E294C + 3 * * * * 151 K573P n/a: not applicable * low intensity MS spectrum, unable to accurately quantify data
Example 15. Conjugation of Combination Variants Having Altered FcRn Binding Affinity, with FLAG Peptide
[0544] Thio-albumin combination variants formulated at 5 mg/mL in Example 10, following 3 months storage at 2-8.degree. C., were conjugated using a 1.5 fold excess of maleimide-propyl-FLAG peptide as shown in FIG. 13A (Peptide Protein Research Ltd., UK, custom synthesis). Variants were diluted with PBS buffer, pH 7.4 to give 1 mL solutions at 1 mg/mL (15.05 nmol). A 1 mg/mL stock solution of maleimide-propyl-FLAG peptide was prepared by reconstituting 5.4 mg material with 5.4 mL PBS buffer pH 7.4. From the maleimide-propyl-FLAG peptide stock solution, 26.28 .mu.L (22.58 nmol) was added to the single thiol variants, 52.56 .mu.L (45.15 nmol) peptide stock solution was added to the double thiol variants, and 78.84 .mu.L (67.73 nmol) peptide stock solution was added to the triple thiol variants to give a threefold excess of maleimide-propyl-FLAG peptide over the number of free thiols. Samples were gently mixed and incubated at ambient temperature overnight. Post incubation the samples were subjected to mass spectrometry to determine the intact protein mass post conjugation as per the MS method described in Example 2 but using a 15 minute analytical gradient, and processing data for the protein peak between approximately 7 and 10 minutes. The results are summarised in Table 31.
[0545] The MS spectrum for the altered FcRn binding variant K573P shown in FIG. 13B indicated that conjugation had occurred post an overnight incubation, giving approximately 29% monoconjugate and 71% unconjugated species respectively, when comparing the relative peak heights of protein species. It was evident that the main peak species at 66409.1 Da was unconjugated K573P variant. The second most abundant peak species had increased by approximately 1164 Da due to a single addition of maleimide-propyl-FLAG peptide. The MS spectrum for the thio-albumin variant K93C+E294C+K573P shown in FIG. 13C indicated that conjugation had occurred post an overnight incubation, giving approximately 29% triconjugate species, approximately 50% diconjugate species, approximately 20% monoconjugate species, and approximately 2% unconjugated species respectively, when comparing the relative peak heights of the protein species. It was evident that the main peak species had increased by approximately 2328 Da to 68685.5 Da, with a 69850.5 Da species indicative of a 3492 Da addition representing a triple conjugation of maleimide-propyl-FLAG peptide.
TABLE-US-00036 TABLE 31 Conjugation efficiency results of albumin variants with FLAG peptide Protein Sample Number Unconju- Mono- Di- Tri- SEQ Description of thiols gated % conjugate % conjugate % conjugate % ID NO. WT control 1 73 27 n/a n/a 2 C34A + K93C 1 48 52 n/a n/a 111 C34A + E294C 1 80 20 n/a n/a 129 C34A + K93C + 2 12 77 10 n/a 141 E294C K93C 2 45 30 25 n/a 142 E294C 2 26 63 11 n/a 143 K93C + E294C 3 * * * * 144 K573P 1 71 29 n/a n/a 145 C34A + K93C + 1 47 53 n/a n/a 146 K573P C34A + E294C + 1 22 78 n/a n/a 147 K573P C34A + K93C + 2 5 34 61 n/a 148 E294C + K573P K93C + K573P 2 23 50 27 n/a 149 E294C + K573P 2 10 51 39 n/a 150 K93C + E294C + 3 2 20 50 29 151 K573P n/a: not applicable * low intensity MS spectrum, unable to accurately quantify data
Example 16. Immunogenicity Assessment of Thio-Albumin Variants Using EpiScreen.TM. Time Course T Cell Assay
[0546] Thio-albumin variants K93C (SEQ ID NO. 142) and E294C (SEQ ID NO. 143) were prepared as described in Example 10 along with a wild type albumin control (SEQ ID NO. 2). In contrast to Example 10, the size exclusion chromatography eluates were diluted to 4 mg/mL (.+-.5%). Albumin test samples were assessed for their ability to induce CD4+ T cell responses using the EpiScreen.TM. time course T cell assay (Abzena, Cambridge UK). Briefly, the EpiScreen.TM. assay was carried out as follows: peripheral blood mononuclear cells from a cohort of 50 healthy donors representing the European and North American population (based on HLA allotypes) were incubated with the test samples. T cell responses were measured using proliferation assays ([3H]-Thymidine uptake) and cytokine secretion assays (IL-2 ELISpot).
[0547] The frequency of positive responses in the proliferation assay were low for all samples (ranges from 0% to 8%) and no positive responses were observed in the IL-2 ELISpot assay suggesting a low risk of clinical immunogenicity for all three samples.
Sequence CWU 1
1
15211761DNAHomo sapiens 1gacgctcaca agtccgaagt cgctcacaga
ttcaaggact tgggtgaaga aaacttcaag 60gctttggtct tgatcgcttt cgctcaatac
ttgcaacaat gtccattcga agatcacgtc 120aagttggtca acgaagttac
cgaattcgct aagacttgtg ttgctgacga atccgcggaa 180aactgtgaca
agtccttgca caccttgttc ggtgataagt tgtgtactgt tgctaccttg
240agagaaacct acggtgaaat ggctgactgt tgtgctaagc aagaaccaga
aagaaacgaa 300tgtttcttgc aacacaagga cgacaaccca aacttgccaa
gattggttag accagaagtt 360gacgtcatgt gtactgcttt ccacgacaac
gaagaaacct tcttgaagaa gtacttgtac 420gaaattgcta gaagacaccc
atacttctac gctccagaat tgttgttctt cgctaagaga 480tacaaggctg
ctttcaccga atgttgtcaa gctgctgata aggctgcttg tttgttgcca
540aagttggatg aattgagaga cgaaggtaag gctagctccg caaagcaaag
attgaagtgt 600gcttccttgc aaaagttcgg tgaaagagct ttcaaggctt
gggctgtcgc tagattgtct 660caaagattcc caaaggctga attcgctgaa
gtttctaagt tggttactga cttgactaag 720gttcacactg aatgttgtca
cggtgacttg ttggaatgtg ctgatgacag agctgacttg 780gctaagtaca
tctgtgaaaa ccaagactct atctcttcca agttgaagga atgttgtgaa
840aagccattgt tggaaaagtc tcactgtatt gctgaagttg aaaacgatga
aatgccagct 900gacttgccat ctttggctgc tgacttcgtt gaatctaagg
acgtttgtaa gaactacgct 960gaagctaagg acgtcttctt gggtatgttc
ttgtacgaat acgctagaag acacccagac 1020tactccgttg tcttgttgtt
gagattggct aagacctacg aaactaccct cgagaagtgt 1080tgtgctgctg
ctgacccaca cgaatgttac gctaaggttt tcgatgaatt caagccattg
1140gtcgaagaac cacaaaactt gatcaagcaa aactgtgaat tgttcgaaca
attgggtgaa 1200tacaagttcc aaaacgcttt gttggttaga tacactaaga
aggtcccaca agtctccacc 1260ccaactttgg ttgaagtctc tagaaacttg
ggtaaggtcg gttctaagtg ttgtaagcac 1320ccagaagcta agagaatgcc
atgtgctgaa gattacttgt ccgtcgtttt gaaccaattg 1380tgtgttttgc
acgaaaagac cccagtctct gatagagtca ccaagtgttg tactgaatct
1440ttggttaaca gaagaccatg tttctctgct ttggaagtcg acgaaactta
cgttccaaag 1500gaattcaacg ctgaaacttt caccttccac gctgatatct
gtaccttgtc cgaaaaggaa 1560agacaaatta agaagcaaac tgctttggtt
gaattggtca agcacaagcc aaaggctact 1620aaggaacaat tgaaggctgt
catggatgat ttcgctgctt tcgttgaaaa gtgttgtaag 1680gctgatgata
aggaaacttg tttcgctgaa gaaggtaaga agttggtcgc tgcttcccaa
1740gctgccttag gtttgtaata a 17612585PRTHomo sapiens 2Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Cys Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
5853609PRTHomo
sapiensSIGNAL(1)..(19)PROPEP(20)..(24)mat_peptide(25)..(609) 3Met
Lys Trp Val Thr Phe Ile Ser Leu Leu Phe Leu Phe Ser Ser Ala -20 -15
-10Tyr Ser Arg Gly Val Phe Arg Arg Asp Ala His Lys Ser Glu Val Ala
-5 -1 1 5His Arg Phe Lys Asp Leu Gly Glu Glu Asn Phe Lys Ala Leu
Val Leu 10 15 20Ile Ala Phe Ala Gln Tyr Leu Gln Gln Cys Pro Phe Glu
Asp His Val25 30 35 40Lys Leu Val Asn Glu Val Thr Glu Phe Ala Lys
Thr Cys Val Ala Asp 45 50 55Glu Ser Ala Glu Asn Cys Asp Lys Ser Leu
His Thr Leu Phe Gly Asp 60 65 70Lys Leu Cys Thr Val Ala Thr Leu Arg
Glu Thr Tyr Gly Glu Met Ala 75 80 85Asp Cys Cys Ala Lys Gln Glu Pro
Glu Arg Asn Glu Cys Phe Leu Gln 90 95 100His Lys Asp Asp Asn Pro
Asn Leu Pro Arg Leu Val Arg Pro Glu Val105 110 115 120Asp Val Met
Cys Thr Ala Phe His Asp Asn Glu Glu Thr Phe Leu Lys 125 130 135Lys
Tyr Leu Tyr Glu Ile Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro 140 145
150Glu Leu Leu Phe Phe Ala Lys Arg Tyr Lys Ala Ala Phe Thr Glu Cys
155 160 165Cys Gln Ala Ala Asp Lys Ala Ala Cys Leu Leu Pro Lys Leu
Asp Glu 170 175 180Leu Arg Asp Glu Gly Lys Ala Ser Ser Ala Lys Gln
Arg Leu Lys Cys185 190 195 200Ala Ser Leu Gln Lys Phe Gly Glu Arg
Ala Phe Lys Ala Trp Ala Val 205 210 215Ala Arg Leu Ser Gln Arg Phe
Pro Lys Ala Glu Phe Ala Glu Val Ser 220 225 230Lys Leu Val Thr Asp
Leu Thr Lys Val His Thr Glu Cys Cys His Gly 235 240 245Asp Leu Leu
Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala Lys Tyr Ile 250 255 260Cys
Glu Asn Gln Asp Ser Ile Ser Ser Lys Leu Lys Glu Cys Cys Glu265 270
275 280Lys Pro Leu Leu Glu Lys Ser His Cys Ile Ala Glu Val Glu Asn
Asp 285 290 295Glu Met Pro Ala Asp Leu Pro Ser Leu Ala Ala Asp Phe
Val Glu Ser 300 305 310Lys Asp Val Cys Lys Asn Tyr Ala Glu Ala Lys
Asp Val Phe Leu Gly 315 320 325Met Phe Leu Tyr Glu Tyr Ala Arg Arg
His Pro Asp Tyr Ser Val Val 330 335 340Leu Leu Leu Arg Leu Ala Lys
Thr Tyr Glu Thr Thr Leu Glu Lys Cys345 350 355 360Cys Ala Ala Ala
Asp Pro His Glu Cys Tyr Ala Lys Val Phe Asp Glu 365 370 375Phe Lys
Pro Leu Val Glu Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys 380 385
390Glu Leu Phe Glu Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu
395 400 405Val Arg Tyr Thr Lys Lys Val Pro Gln Val Ser Thr Pro Thr
Leu Val 410 415 420Glu Val Ser Arg Asn Leu Gly Lys Val Gly Ser Lys
Cys Cys Lys His425 430 435 440Pro Glu Ala Lys Arg Met Pro Cys Ala
Glu Asp Tyr Leu Ser Val Val 445 450 455Leu Asn Gln Leu Cys Val Leu
His Glu Lys Thr Pro Val Ser Asp Arg 460 465 470Val Thr Lys Cys Cys
Thr Glu Ser Leu Val Asn Arg Arg Pro Cys Phe 475 480 485Ser Ala Leu
Glu Val Asp Glu Thr Tyr Val Pro Lys Glu Phe Asn Ala 490 495 500Glu
Thr Phe Thr Phe His Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu505 510
515 520Arg Gln Ile Lys Lys Gln Thr Ala Leu Val Glu Leu Val Lys His
Lys 525 530 535Pro Lys Ala Thr Lys Glu Gln Leu Lys Ala Val Met Asp
Asp Phe Ala 540 545 550Ala Phe Val Glu Lys Cys Cys Lys Ala Asp Asp
Lys Glu Thr Cys Phe 555 560 565Ala Glu Glu Gly Lys Lys Leu Val Ala
Ala Ser Gln Ala Ala Leu Gly 570 575 580Leu5854621PRTPan troglodytes
4Met Asn Glu Ser Ser Cys Cys Ser Thr Ser Leu Pro Ala Phe Gly Val1 5
10 15Ser Val Leu Asp Ser Gly His Ser Ser Ser Ser Ala Tyr Ser Arg
Gly 20 25 30Val Phe Arg Arg Asp Ala His Lys Ser Glu Val Ala His Arg
Phe Lys 35 40 45Asp Leu Gly Glu Glu Asn Phe Lys Ala Leu Val Leu Val
Ala Phe Ala 50 55 60Gln Tyr Leu Gln Gln Cys Pro Phe Glu Asp His Val
Lys Leu Val Asn65 70 75 80Glu Val Thr Glu Phe Ala Lys Thr Cys Val
Ala Asp Glu Ser Ala Glu 85 90 95Asn Cys Asp Lys Ser Leu His Thr Leu
Phe Gly Asp Lys Leu Cys Thr 100 105 110Val Ala Thr Leu Arg Glu Lys
Tyr Gly Glu Met Ala Asp Cys Cys Ala 115 120 125Lys Gln Glu Pro Glu
Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp 130 135 140Asn Pro Asn
Leu Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys145 150 155
160Thr Ala Phe His Asp Asn Glu Gly Thr Phe Leu Lys Lys Tyr Leu Tyr
165 170 175Glu Val Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu
Leu Phe 180 185 190Phe Ala Glu Arg Tyr Lys Ala Ala Phe Thr Glu Cys
Cys Gln Ala Ala 195 200 205Asp Lys Ala Ala Cys Leu Leu Pro Lys Leu
Asp Glu Leu Arg Asp Glu 210 215 220Gly Lys Ala Ser Ser Ala Lys Gln
Arg Leu Lys Cys Ala Ser Leu Gln225 230 235 240Lys Phe Gly Glu Arg
Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser 245 250 255Gln Arg Phe
Pro Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr 260 265 270Asp
Leu Thr Lys Val His Thr Glu Cys Cys His Gly Asp Leu Leu Glu 275 280
285Cys Ala Asp Asp Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln
290 295 300Asp Ser Ile Ser Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro
Leu Leu305 310 315 320Glu Lys Ser His Cys Leu Ala Glu Val Glu Asn
Asp Glu Met Pro Ala 325 330 335Asp Leu Pro Ser Leu Ala Ala Asp Phe
Val Glu Ser Lys Glu Val Cys 340 345 350Lys Asn Tyr Ala Glu Ala Lys
Asp Val Phe Leu Gly Met Phe Leu Tyr 355 360 365Glu Tyr Ala Arg Arg
His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg 370 375 380Leu Ala Lys
Thr Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala385 390 395
400Asp Pro His Glu Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu
405 410 415Val Glu Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu
Phe Glu 420 425 430Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu
Val Arg Tyr Thr 435 440 445Lys Lys Val Pro Gln Val Ser Thr Pro Thr
Leu Val Glu Val Ser Arg 450 455 460Asn Leu Gly Lys Val Gly Ser Lys
Cys Cys Lys His Pro Glu Ala Lys465 470 475 480Arg Met Pro Cys Ala
Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu 485 490 495Cys Val Leu
His Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys 500 505 510Cys
Thr Glu Ser Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu 515 520
525Val Asp Glu Thr Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr
530 535 540Phe His Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln
Ile Lys545 550 555 560Lys Gln Thr Ala Leu Val Glu Leu Val Lys His
Lys Pro Lys Ala Thr 565 570 575Lys Glu Gln Leu Lys Ala Val Met Asp
Asp Phe Ala Ala Phe Val Glu 580 585 590Lys Cys Cys Lys Ala Asp Asp
Lys Glu Thr Cys Phe Ala Glu Glu Gly 595 600 605Lys Lys Leu Val Ala
Ala Ser Gln Ala Ala Leu Gly Leu 610 615 6205608PRTMacaca mulatta
5Met Lys Trp Val Thr Phe Ile Ser Leu Leu Phe Leu Phe Ser Ser Ala1 5
10 15Tyr Ser Arg Gly Val Phe Arg Arg Asp Thr His Lys Ser Glu Val
Ala 20 25 30His Arg Phe Lys Asp Leu Gly Glu Glu His Phe Lys Gly Leu
Val Leu 35 40 45Val Ala Phe Ser Gln Tyr Leu Gln Gln Cys Pro Phe Glu
Glu His Val 50 55 60Lys Leu Val Asn Glu Val Thr Glu Phe Ala Lys Thr
Cys Val Ala Asp65 70 75 80Glu Ser Ala Glu Asn Cys Asp Lys Ser Leu
His Thr Leu Phe Gly Asp 85 90 95Lys Leu Cys Thr Val Ala Thr Leu Arg
Glu Thr Tyr Gly Glu Met Ala 100 105 110Asp Cys Cys Ala Lys Gln Glu
Pro Glu Arg Asn Glu Cys Phe Leu Gln 115 120 125His Lys Asp Asp Asn
Pro Asn Leu Pro Pro Leu Val Arg Pro Glu Val 130 135 140Asp Val Met
Cys Thr Ala Phe His Asp Asn Glu Ala Thr Phe Leu Lys145 150 155
160Lys Tyr Leu Tyr Glu Val Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro
165 170 175Glu Leu Leu Phe Phe Ala Ala Arg Tyr Lys Ala Ala Phe Ala
Glu Cys 180 185 190Cys Gln Ala Ala Asp Lys Ala Ala Cys Leu Leu Pro
Lys Leu Asp Glu 195 200 205Leu Arg Asp Glu Gly Lys Ala Ser Ser Ala
Lys Gln Arg Leu Lys Cys 210 215 220Ala Ser Leu Gln Lys Phe Gly Asp
Arg Ala Phe Lys Ala Trp Ala Val225 230 235 240Ala Arg Leu Ser Gln
Lys Phe Pro Lys Ala Glu Phe Ala Glu Val Ser 245 250 255Lys Leu Val
Thr Asp Leu Thr Lys Val His Thr Glu Cys Cys His Gly 260 265 270Asp
Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala Lys Tyr Met 275 280
285Cys Glu Asn Gln Asp Ser Ile Ser Ser Lys Leu Lys Glu Cys Cys Asp
290 295 300Lys Pro Leu Leu Glu Lys Ser His Cys Leu Ala Glu Val Glu
Asn Asp305 310 315 320Glu Met Pro Ala Asp Leu Pro Ser Leu Ala Ala
Asp Tyr Val Glu Ser 325 330
335Lys Asp Val Cys Lys Asn Tyr Ala Glu Ala Lys Asp Val Phe Leu Gly
340 345 350Met Phe Leu Tyr Glu Tyr Ala Arg Arg His Pro Asp Tyr Ser
Val Met 355 360 365Leu Leu Leu Arg Leu Ala Lys Ala Tyr Glu Ala Thr
Leu Glu Lys Cys 370 375 380Cys Ala Ala Ala Asp Pro His Glu Cys Tyr
Ala Lys Val Phe Asp Glu385 390 395 400Phe Gln Pro Leu Val Glu Glu
Pro Gln Asn Leu Val Lys Gln Asn Cys 405 410 415Glu Leu Phe Glu Gln
Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu 420 425 430Val Arg Tyr
Thr Lys Lys Val Pro Gln Val Ser Thr Pro Thr Leu Val 435 440 445Glu
Val Ser Arg Asn Leu Gly Lys Val Gly Ala Lys Cys Cys Lys Leu 450 455
460Pro Glu Ala Lys Arg Met Pro Cys Ala Glu Asp Tyr Leu Ser Val
Val465 470 475 480Leu Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro
Val Ser Glu Lys 485 490 495Val Thr Lys Cys Cys Thr Glu Ser Leu Val
Asn Arg Arg Pro Cys Phe 500 505 510Ser Ala Leu Glu Leu Asp Glu Ala
Tyr Val Pro Lys Ala Phe Asn Ala 515 520 525Glu Thr Phe Thr Phe His
Ala Asp Met Cys Thr Leu Ser Glu Lys Glu 530 535 540Lys Gln Val Lys
Lys Gln Thr Ala Leu Val Glu Leu Val Lys His Lys545 550 555 560Pro
Lys Ala Thr Lys Glu Gln Leu Lys Gly Val Met Asp Asn Phe Ala 565 570
575Ala Phe Val Glu Lys Cys Cys Lys Ala Asp Asp Lys Glu Ala Cys Phe
580 585 590Ala Glu Glu Gly Pro Lys Phe Val Ala Ala Ser Gln Ala Ala
Leu Ala 595 600 6056608PRTMesocricetus auratus 6Met Lys Trp Val Thr
Phe Leu Leu Leu Leu Phe Val Ser Asp Ser Ala1 5 10 15Phe Ser Arg Gly
Leu Phe Arg Arg Asp Ala His Lys Ser Glu Ile Ala 20 25 30His Arg Phe
Lys Asp Leu Gly Glu Gln His Phe Lys Gly Leu Val Leu 35 40 45Ile Ala
Phe Ser Gln Phe Leu Gln Lys Cys Pro Tyr Glu Glu His Val 50 55 60Lys
Leu Val Asn Glu Val Thr Asp Phe Ala Lys Thr Cys Val Ala Asp65 70 75
80Glu Ser Ala Glu Asn Cys Asp Lys Ser Leu His Thr Leu Phe Gly Asp
85 90 95Lys Leu Cys Ala Ile Pro Thr Leu Arg Asp Ser Tyr Gly Glu Leu
Ala 100 105 110Asp Cys Cys Ala Lys Lys Glu Pro Glu Arg Asn Glu Cys
Phe Leu Lys 115 120 125His Lys Asp Asp His Pro Asn Leu Pro Pro Phe
Val Arg Pro Asp Ala 130 135 140Glu Ala Met Cys Thr Ser Phe Gln Glu
Asn Ala Val Thr Phe Met Gly145 150 155 160His Tyr Leu His Glu Val
Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro 165 170 175Glu Leu Leu Tyr
Tyr Ala Glu Lys Tyr Ser Ala Ile Met Thr Glu Cys 180 185 190Cys Gly
Glu Ala Asp Lys Ala Ala Cys Ile Thr Pro Lys Leu Asp Ala 195 200
205Leu Lys Glu Lys Ala Leu Ala Ser Ser Val Asn Gln Arg Leu Lys Cys
210 215 220Ser Ser Leu Gln Arg Phe Gly Gln Arg Ala Phe Lys Ala Trp
Ala Val225 230 235 240Ala Arg Met Ser Gln Lys Phe Pro Lys Ala Asp
Phe Ala Glu Ile Thr 245 250 255Lys Leu Ala Thr Asp Leu Thr Lys Leu
Thr Glu Glu Cys Cys His Gly 260 265 270Asp Leu Leu Glu Cys Ala Asp
Asp Arg Ala Glu Leu Ala Lys Tyr Met 275 280 285Cys Glu Asn Gln Ala
Ser Ile Ser Ser Lys Leu Gln Ala Cys Cys Asp 290 295 300Lys Pro Val
Leu Lys Lys Ser His Cys Leu Ser Glu Val Glu Asn Asp305 310 315
320Asp Leu Pro Ala Asp Leu Pro Ser Leu Ala Ala Asp Phe Val Glu Asp
325 330 335Lys Glu Val Cys Lys Asn Tyr Ala Glu Ala Lys Asp Val Phe
Leu Gly 340 345 350Thr Phe Leu Tyr Glu Tyr Ala Arg Arg His Pro Asp
Tyr Ser Val Ala 355 360 365Leu Leu Leu Arg Leu Ala Lys Lys Tyr Glu
Ala Thr Leu Glu Lys Cys 370 375 380Cys Ala Glu Ala Asp Pro Ser Ala
Cys Tyr Gly Lys Val Leu Asp Glu385 390 395 400Phe Gln Pro Leu Val
Glu Glu Pro Lys Asn Leu Val Lys Ala Asn Cys 405 410 415Glu Leu Phe
Glu Lys Leu Gly Glu Tyr Gly Phe Gln Asn Ala Leu Ile 420 425 430Val
Arg Tyr Thr Gln Lys Ala Pro Gln Val Ser Thr Pro Thr Leu Val 435 440
445Glu Ala Ala Arg Asn Leu Gly Lys Val Gly Ser Lys Cys Cys Val Leu
450 455 460Pro Glu Ala Gln Arg Leu Pro Cys Val Glu Asp Tyr Ile Ser
Ala Ile465 470 475 480Leu Asn Arg Val Cys Val Leu His Glu Lys Thr
Pro Val Ser Glu Gln 485 490 495Val Thr Lys Cys Cys Thr Gly Ser Val
Val Glu Arg Arg Pro Cys Phe 500 505 510Ser Ala Leu Pro Val Asp Glu
Thr Tyr Val Pro Lys Glu Phe Lys Ala 515 520 525Glu Thr Phe Thr Phe
His Ala Asp Ile Cys Ser Leu Pro Glu Lys Glu 530 535 540Lys Gln Met
Lys Lys Gln Ala Ala Leu Val Glu Leu Val Lys His Lys545 550 555
560Pro Lys Ala Thr Gly Pro Gln Leu Arg Thr Val Leu Gly Glu Phe Thr
565 570 575Ala Phe Leu Asp Lys Cys Cys Lys Ala Glu Asp Lys Glu Ala
Cys Phe 580 585 590Ser Glu Asp Gly Pro Lys Leu Val Ala Ser Ser Gln
Ala Ala Leu Ala 595 600 6057608PRTCavia porcellus 7Met Lys Trp Val
Thr Phe Ile Ser Leu Leu Phe Leu Phe Ser Ser Val1 5 10 15Tyr Ser Arg
Gly Val Phe Arg Arg Glu Ala His Lys Ser Glu Ile Ala 20 25 30His Arg
Phe Asn Asp Leu Gly Glu Gly His Phe Lys Gly Leu Val Leu 35 40 45Ile
Thr Leu Ser Gln His Leu Gln Lys Ser Pro Phe Glu Glu His Val 50 55
60Lys Leu Val Asn Glu Val Thr Asp Phe Ala Lys Ala Cys Val Ala Asp65
70 75 80Glu Ser Ala Gln Asn Cys Gly Lys Ala Ile Ala Thr Leu Phe Gly
Asp 85 90 95Lys Val Cys Ala Ile Pro Ser Leu Arg Glu Thr Tyr Gly Glu
Leu Ala 100 105 110Asp Cys Cys Ala Lys Glu Asp Pro Asp Arg Val Glu
Cys Phe Leu Gln 115 120 125His Lys Asp Asp Asn Pro Asn Leu Pro Pro
Phe Glu Arg Pro Glu Pro 130 135 140Glu Ala Leu Cys Thr Ala Phe Lys
Glu Asn Asn Asp Arg Phe Ile Gly145 150 155 160His Tyr Leu Tyr Glu
Val Ser Arg Arg His Pro Tyr Phe Tyr Ala Pro 165 170 175Glu Leu Leu
Tyr Tyr Ala Glu Lys Tyr Lys Asn Ala Leu Thr Glu Cys 180 185 190Cys
Glu Ala Ala Asp Lys Ala Ala Cys Leu Thr Pro Lys Leu Asp Ala 195 200
205Ile Lys Glu Lys Ala Leu Val Ser Ser Ala Gln Gln Arg Leu Lys Cys
210 215 220Ala Ser Leu Gln Lys Phe Gly Glu Arg Ala Phe Lys Ala Trp
Ser Val225 230 235 240Ala Arg Leu Ser Gln Lys Phe Pro Lys Ala Glu
Phe Ala Glu Ile Ser 245 250 255Thr Ile Val Thr Ser Leu Thr Lys Val
Thr Lys Glu Cys Cys His Gly 260 265 270Asp Leu Leu Glu Cys Ala Asp
Asp Arg Gln Glu Leu Ala Lys Tyr Met 275 280 285Cys Glu His Gln Asp
Ser Ile Ser Ser Lys Leu Lys Glu Cys Cys Val 290 295 300Lys Pro Thr
Leu Gln Lys Ala His Cys Ile Leu Glu Ile Gln Arg Asp305 310 315
320Glu Leu Pro Thr Glu Leu Pro Asp Leu Ala Val Asp Phe Val Glu Asp
325 330 335Lys Glu Val Cys Lys Asn Phe Ala Glu Ala Lys Asp Val Phe
Leu Gly 340 345 350Thr Phe Leu Tyr Glu Tyr Ser Arg Arg His Pro Glu
Tyr Ser Ile Gly 355 360 365Met Leu Leu Arg Ile Ala Lys Gly Tyr Glu
Ala Lys Leu Glu Lys Cys 370 375 380Cys Ala Glu Ala Asp Pro His Ala
Cys Tyr Ala Lys Val Phe Asp Glu385 390 395 400Leu Gln Pro Leu Ile
Asp Glu Pro Lys Lys Leu Val Gln Gln Asn Cys 405 410 415Glu Leu Phe
Asp Lys Leu Gly Glu Tyr Gly Phe Gln Asn Ala Leu Ala 420 425 430Val
Arg Tyr Thr Gln Lys Ala Pro Gln Val Ser Thr Pro Thr Leu Val 435 440
445Glu Tyr Ala Arg Lys Leu Gly Ser Val Gly Thr Lys Cys Cys Ser Leu
450 455 460Pro Glu Thr Glu Arg Leu Ser Cys Thr Glu Asn Tyr Leu Ala
Leu Ile465 470 475 480Leu Asn Arg Leu Cys Ile Leu His Glu Lys Thr
Pro Val Ser Glu Arg 485 490 495Val Thr Lys Cys Cys Thr Glu Ser Leu
Val Asn Arg Arg Pro Cys Phe 500 505 510Ser Ala Leu His Val Asp Glu
Thr Tyr Val Pro Lys Pro Phe His Ala 515 520 525Asp Ser Phe Thr Phe
His Ala Asp Ile Cys Thr Leu Pro Glu Lys Glu 530 535 540Lys Gln Val
Lys Lys Gln Met Ala Leu Val Glu Leu Val Lys His Lys545 550 555
560Pro Lys Ala Ser Glu Glu Gln Met Lys Thr Val Met Gly Asp Phe Ala
565 570 575Ala Phe Leu Lys Lys Cys Cys Asp Ala Asp Asn Lys Glu Ala
Cys Phe 580 585 590Thr Glu Asp Gly Pro Lys Leu Val Ala Lys Cys Gln
Ala Thr Leu Ala 595 600 6058608PRTMus musculus 8Met Lys Trp Val Thr
Phe Leu Leu Leu Leu Phe Val Ser Gly Ser Ala1 5 10 15Phe Ser Arg Gly
Val Phe Arg Arg Glu Ala His Lys Ser Glu Ile Ala 20 25 30His Arg Tyr
Asn Asp Leu Gly Glu Gln His Phe Lys Gly Leu Val Leu 35 40 45Ile Ala
Phe Ser Gln Tyr Leu Gln Lys Cys Ser Tyr Asp Glu His Ala 50 55 60Lys
Leu Val Gln Glu Val Thr Asp Phe Ala Lys Thr Cys Val Ala Asp65 70 75
80Glu Ser Ala Ala Asn Cys Asp Lys Ser Leu His Thr Leu Phe Gly Asp
85 90 95Lys Leu Cys Ala Ile Pro Asn Leu Arg Glu Asn Tyr Gly Glu Leu
Ala 100 105 110Asp Cys Cys Thr Lys Gln Glu Pro Glu Arg Asn Glu Cys
Phe Leu Gln 115 120 125His Lys Asp Asp Asn Pro Ser Leu Pro Pro Phe
Glu Arg Pro Glu Ala 130 135 140Glu Ala Met Cys Thr Ser Phe Lys Glu
Asn Pro Thr Thr Phe Met Gly145 150 155 160His Tyr Leu His Glu Val
Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro 165 170 175Glu Leu Leu Tyr
Tyr Ala Glu Gln Tyr Asn Glu Ile Leu Thr Gln Cys 180 185 190Cys Ala
Glu Ala Asp Lys Glu Ser Cys Leu Thr Pro Lys Leu Asp Gly 195 200
205Val Lys Glu Lys Ala Leu Val Ser Ser Val Arg Gln Arg Met Lys Cys
210 215 220Ser Ser Met Gln Lys Phe Gly Glu Arg Ala Phe Lys Ala Trp
Ala Val225 230 235 240Ala Arg Leu Ser Gln Thr Phe Pro Asn Ala Asp
Phe Ala Glu Ile Thr 245 250 255Lys Leu Ala Thr Asp Leu Thr Lys Val
Asn Lys Glu Cys Cys His Gly 260 265 270Asp Leu Leu Glu Cys Ala Asp
Asp Arg Ala Glu Leu Ala Lys Tyr Met 275 280 285Cys Glu Asn Gln Ala
Thr Ile Ser Ser Lys Leu Gln Thr Cys Cys Asp 290 295 300Lys Pro Leu
Leu Lys Lys Ala His Cys Leu Ser Glu Val Glu His Asp305 310 315
320Thr Met Pro Ala Asp Leu Pro Ala Ile Ala Ala Asp Phe Val Glu Asp
325 330 335Gln Glu Val Cys Lys Asn Tyr Ala Glu Ala Lys Asp Val Phe
Leu Gly 340 345 350Thr Phe Leu Tyr Glu Tyr Ser Arg Arg His Pro Asp
Tyr Ser Val Ser 355 360 365Leu Leu Leu Arg Leu Ala Lys Lys Tyr Glu
Ala Thr Leu Glu Lys Cys 370 375 380Cys Ala Glu Ala Asn Pro Pro Ala
Cys Tyr Gly Thr Val Leu Ala Glu385 390 395 400Phe Gln Pro Leu Val
Glu Glu Pro Lys Asn Leu Val Lys Thr Asn Cys 405 410 415Asp Leu Tyr
Glu Lys Leu Gly Glu Tyr Gly Phe Gln Asn Ala Ile Leu 420 425 430Val
Arg Tyr Thr Gln Lys Ala Pro Gln Val Ser Thr Pro Thr Leu Val 435 440
445Glu Ala Ala Arg Asn Leu Gly Arg Val Gly Thr Lys Cys Cys Thr Leu
450 455 460Pro Glu Asp Gln Arg Leu Pro Cys Val Glu Asp Tyr Leu Ser
Ala Ile465 470 475 480Leu Asn Arg Val Cys Leu Leu His Glu Lys Thr
Pro Val Ser Glu His 485 490 495Val Thr Lys Cys Cys Ser Gly Ser Leu
Val Glu Arg Arg Pro Cys Phe 500 505 510Ser Ala Leu Thr Val Asp Glu
Thr Tyr Val Pro Lys Glu Phe Lys Ala 515 520 525Glu Thr Phe Thr Phe
His Ser Asp Ile Cys Thr Leu Pro Glu Lys Glu 530 535 540Lys Gln Ile
Lys Lys Gln Thr Ala Leu Ala Glu Leu Val Lys His Lys545 550 555
560Pro Lys Ala Thr Ala Glu Gln Leu Lys Thr Val Met Asp Asp Phe Ala
565 570 575Gln Phe Leu Asp Thr Cys Cys Lys Ala Ala Asp Lys Asp Thr
Cys Phe 580 585 590Ser Thr Glu Gly Pro Asn Leu Val Thr Arg Cys Lys
Asp Ala Leu Ala 595 600 6059608PRTRattus norvegicus 9Met Lys Trp
Val Thr Phe Leu Leu Leu Leu Phe Ile Ser Gly Ser Ala1 5 10 15Phe Ser
Arg Gly Val Phe Arg Arg Glu Ala His Lys Ser Glu Ile Ala 20 25 30His
Arg Phe Lys Asp Leu Gly Glu Gln His Phe Lys Gly Leu Val Leu 35 40
45Ile Ala Phe Ser Gln Tyr Leu Gln Lys Cys Pro Tyr Glu Glu His Ile
50 55 60Lys Leu Val Gln Glu Val Thr Asp Phe Ala Lys Thr Cys Val Ala
Asp65 70 75 80Glu Asn Ala Glu Asn Cys Asp Lys Ser Ile His Thr Leu
Phe Gly Asp 85 90 95Lys Leu Cys Ala Ile Pro Lys Leu Arg Asp Asn Tyr
Gly Glu Leu Ala 100 105 110Asp Cys Cys Ala Lys Gln Glu Pro Glu Arg
Asn Glu Cys Phe Leu Gln 115 120 125His Lys Asp Asp Asn Pro Asn Leu
Pro Pro Phe Gln Arg Pro Glu Ala 130 135 140Glu Ala Met Cys Thr Ser
Phe Gln Glu Asn Pro Thr Ser Phe Leu Gly145 150 155 160His Tyr Leu
His Glu Val Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro 165 170 175Glu
Leu Leu Tyr Tyr Ala Glu Lys Tyr Asn Glu Val Leu Thr Gln Cys 180 185
190Cys Thr Glu Ser Asp Lys Ala Ala Cys Leu Thr Pro Lys Leu Asp Ala
195 200 205Val Lys Glu Lys Ala Leu Val Ala Ala Val Arg Gln Arg Met
Lys Cys 210 215 220Ser Ser Met Gln Arg Phe Gly Glu Arg Ala Phe Lys
Ala Trp Ala Val225 230 235 240Ala Arg Met Ser Gln Arg Phe Pro Asn
Ala Glu Phe Ala Glu Ile Thr 245 250 255Lys Leu Ala Thr Asp Val Thr
Lys Ile Asn Lys Glu Cys Cys His Gly 260 265 270Asp Leu Leu Glu Cys
Ala Asp Asp Arg Ala Glu Leu Ala Lys Tyr Met 275 280 285Cys Glu Asn
Gln Ala Thr Ile Ser Ser Lys Leu Gln Ala Cys Cys Asp 290 295 300Lys
Pro Val Leu Gln Lys Ser Gln Cys Leu Ala Glu Ile Glu His Asp305 310
315 320Asn Ile Pro Ala Asp Leu Pro Ser Ile Ala Ala Asp Phe Val Glu
Asp 325 330 335Lys Glu Val Cys Lys Asn Tyr Ala Glu Ala Lys Asp Val
Phe Leu Gly 340 345 350Thr Phe Leu Tyr Glu Tyr Ser Arg Arg His Pro
Asp Tyr Ser Val Ser 355 360 365Leu
Leu Leu Arg Leu Ala Lys Lys Tyr Glu Ala Thr Leu Glu Lys Cys 370 375
380Cys Ala Glu Gly Asp Pro Pro Ala Cys Tyr Gly Thr Val Leu Ala
Glu385 390 395 400Phe Gln Pro Leu Val Glu Glu Pro Lys Asn Leu Val
Lys Thr Asn Cys 405 410 415Glu Leu Tyr Glu Lys Leu Gly Glu Tyr Gly
Phe Gln Asn Ala Val Leu 420 425 430Val Arg Tyr Thr Gln Lys Ala Pro
Gln Val Ser Thr Pro Thr Leu Val 435 440 445Glu Ala Ala Arg Asn Leu
Gly Arg Val Gly Thr Lys Cys Cys Thr Leu 450 455 460Pro Glu Ala Gln
Arg Leu Pro Cys Val Glu Asp Tyr Leu Ser Ala Ile465 470 475 480Leu
Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro Val Ser Glu Lys 485 490
495Val Thr Lys Cys Cys Ser Gly Ser Leu Val Glu Arg Arg Pro Cys Phe
500 505 510Ser Ala Leu Thr Val Asp Glu Thr Tyr Val Pro Lys Glu Phe
Lys Ala 515 520 525Glu Thr Phe Thr Phe His Ser Asp Ile Cys Thr Leu
Pro Asp Lys Glu 530 535 540Lys Gln Ile Lys Lys Gln Thr Ala Leu Ala
Glu Leu Val Lys His Lys545 550 555 560Pro Lys Ala Thr Glu Asp Gln
Leu Lys Thr Val Met Gly Asp Phe Ala 565 570 575Gln Phe Val Asp Lys
Cys Cys Lys Ala Ala Asp Lys Asp Asn Cys Phe 580 585 590Ala Thr Glu
Gly Pro Asn Leu Val Ala Arg Ser Lys Glu Ala Leu Ala 595 600
60510607PRTBos taurus 10Met Lys Trp Val Thr Phe Ile Ser Leu Leu Leu
Leu Phe Ser Ser Ala1 5 10 15Tyr Ser Arg Gly Val Phe Arg Arg Asp Thr
His Lys Ser Glu Ile Ala 20 25 30His Arg Phe Lys Asp Leu Gly Glu Glu
His Phe Lys Gly Leu Val Leu 35 40 45Ile Ala Phe Ser Gln Tyr Leu Gln
Gln Cys Pro Phe Asp Glu His Val 50 55 60Lys Leu Val Asn Glu Leu Thr
Glu Phe Ala Lys Thr Cys Val Ala Asp65 70 75 80Glu Ser His Ala Gly
Cys Glu Lys Ser Leu His Thr Leu Phe Gly Asp 85 90 95Glu Leu Cys Lys
Val Ala Ser Leu Arg Glu Thr Tyr Gly Asp Met Ala 100 105 110Asp Cys
Cys Glu Lys Gln Glu Pro Glu Arg Asn Glu Cys Phe Leu Ser 115 120
125His Lys Asp Asp Ser Pro Asp Leu Pro Lys Leu Lys Pro Asp Pro Asn
130 135 140Thr Leu Cys Asp Glu Phe Lys Ala Asp Glu Lys Lys Phe Trp
Gly Lys145 150 155 160Tyr Leu Tyr Glu Ile Ala Arg Arg His Pro Tyr
Phe Tyr Ala Pro Glu 165 170 175Leu Leu Tyr Tyr Ala Asn Lys Tyr Asn
Gly Val Phe Gln Glu Cys Cys 180 185 190Gln Ala Glu Asp Lys Gly Ala
Cys Leu Leu Pro Lys Ile Glu Thr Met 195 200 205Arg Glu Lys Val Leu
Ala Ser Ser Ala Arg Gln Arg Leu Arg Cys Ala 210 215 220Ser Ile Gln
Lys Phe Gly Glu Arg Ala Leu Lys Ala Trp Ser Val Ala225 230 235
240Arg Leu Ser Gln Lys Phe Pro Lys Ala Glu Phe Val Glu Val Thr Lys
245 250 255Leu Val Thr Asp Leu Thr Lys Val His Lys Glu Cys Cys His
Gly Asp 260 265 270Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala
Lys Tyr Ile Cys 275 280 285Asp Asn Gln Asp Thr Ile Ser Ser Lys Leu
Lys Glu Cys Cys Asp Lys 290 295 300Pro Leu Leu Glu Lys Ser His Cys
Ile Ala Glu Val Glu Lys Asp Ala305 310 315 320Ile Pro Glu Asn Leu
Pro Pro Leu Thr Ala Asp Phe Ala Glu Asp Lys 325 330 335Asp Val Cys
Lys Asn Tyr Gln Glu Ala Lys Asp Ala Phe Leu Gly Ser 340 345 350Phe
Leu Tyr Glu Tyr Ser Arg Arg His Pro Glu Tyr Ala Val Ser Val 355 360
365Leu Leu Arg Leu Ala Lys Glu Tyr Glu Ala Thr Leu Glu Glu Cys Cys
370 375 380Ala Lys Asp Asp Pro His Ala Cys Tyr Ser Thr Val Phe Asp
Lys Leu385 390 395 400Lys His Leu Val Asp Glu Pro Gln Asn Leu Ile
Lys Gln Asn Cys Asp 405 410 415Gln Phe Glu Lys Leu Gly Glu Tyr Gly
Phe Gln Asn Ala Leu Ile Val 420 425 430Arg Tyr Thr Arg Lys Val Pro
Gln Val Ser Thr Pro Thr Leu Val Glu 435 440 445Val Ser Arg Ser Leu
Gly Lys Val Gly Thr Arg Cys Cys Thr Lys Pro 450 455 460Glu Ser Glu
Arg Met Pro Cys Thr Glu Asp Tyr Leu Ser Leu Ile Leu465 470 475
480Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro Val Ser Glu Lys Val
485 490 495Thr Lys Cys Cys Thr Glu Ser Leu Val Asn Arg Arg Pro Cys
Phe Ser 500 505 510Ala Leu Thr Pro Asp Glu Thr Tyr Val Pro Lys Ala
Phe Asp Glu Lys 515 520 525Leu Phe Thr Phe His Ala Asp Ile Cys Thr
Leu Pro Asp Thr Glu Lys 530 535 540Gln Ile Lys Lys Gln Thr Ala Leu
Val Glu Leu Leu Lys His Lys Pro545 550 555 560Lys Ala Thr Glu Glu
Gln Leu Lys Thr Val Met Glu Asn Phe Val Ala 565 570 575Phe Val Asp
Lys Cys Cys Ala Ala Asp Asp Lys Glu Ala Cys Phe Ala 580 585 590Val
Glu Gly Pro Lys Leu Val Val Ser Thr Gln Thr Ala Leu Ala 595 600
60511607PRTEquus caballus 11Met Lys Trp Val Thr Phe Val Ser Leu Leu
Phe Leu Phe Ser Ser Ala1 5 10 15Tyr Ser Arg Gly Val Leu Arg Arg Asp
Thr His Lys Ser Glu Ile Ala 20 25 30His Arg Phe Asn Asp Leu Gly Glu
Lys His Phe Lys Gly Leu Val Leu 35 40 45Val Ala Phe Ser Gln Tyr Leu
Gln Gln Cys Pro Phe Glu Asp His Val 50 55 60Lys Leu Val Asn Glu Val
Thr Glu Phe Ala Lys Lys Cys Ala Ala Asp65 70 75 80Glu Ser Ala Glu
Asn Cys Asp Lys Ser Leu His Thr Leu Phe Gly Asp 85 90 95Lys Leu Cys
Thr Val Ala Thr Leu Arg Ala Thr Tyr Gly Glu Leu Ala 100 105 110Asp
Cys Cys Glu Lys Gln Glu Pro Glu Arg Asn Glu Cys Phe Leu Thr 115 120
125His Lys Asp Asp His Pro Asn Leu Pro Lys Leu Lys Pro Glu Pro Asp
130 135 140Ala Gln Cys Ala Ala Phe Gln Glu Asp Pro Asp Lys Phe Leu
Gly Lys145 150 155 160Tyr Leu Tyr Glu Val Ala Arg Arg His Pro Tyr
Phe Tyr Gly Pro Glu 165 170 175Leu Leu Phe His Ala Glu Glu Tyr Lys
Ala Asp Phe Thr Glu Cys Cys 180 185 190Pro Ala Asp Asp Lys Leu Ala
Cys Leu Ile Pro Lys Leu Asp Ala Leu 195 200 205Lys Glu Arg Ile Leu
Leu Ser Ser Ala Lys Glu Arg Leu Lys Cys Ser 210 215 220Ser Phe Gln
Asn Phe Gly Glu Arg Ala Val Lys Ala Trp Ser Val Ala225 230 235
240Arg Leu Ser Gln Lys Phe Pro Lys Ala Asp Phe Ala Glu Val Ser Lys
245 250 255Ile Val Thr Asp Leu Thr Lys Val His Lys Glu Cys Cys His
Gly Asp 260 265 270Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala
Lys Tyr Ile Cys 275 280 285Glu His Gln Asp Ser Ile Ser Gly Lys Leu
Lys Ala Cys Cys Asp Lys 290 295 300Pro Leu Leu Gln Lys Ser His Cys
Ile Ala Glu Val Lys Glu Asp Asp305 310 315 320Leu Pro Ser Asp Leu
Pro Ala Leu Ala Ala Asp Phe Ala Glu Asp Lys 325 330 335Glu Ile Cys
Lys His Tyr Lys Asp Ala Lys Asp Val Phe Leu Gly Thr 340 345 350Phe
Leu Tyr Glu Tyr Ser Arg Arg His Pro Asp Tyr Ser Val Ser Leu 355 360
365Leu Leu Arg Ile Ala Lys Thr Tyr Glu Ala Thr Leu Glu Lys Cys Cys
370 375 380Ala Glu Ala Asp Pro Pro Ala Cys Tyr Arg Thr Val Phe Asp
Gln Phe385 390 395 400Thr Pro Leu Val Glu Glu Pro Lys Ser Leu Val
Lys Lys Asn Cys Asp 405 410 415Leu Phe Glu Glu Val Gly Glu Tyr Asp
Phe Gln Asn Ala Leu Ile Val 420 425 430Arg Tyr Thr Lys Lys Ala Pro
Gln Val Ser Thr Pro Thr Leu Val Glu 435 440 445Ile Gly Arg Thr Leu
Gly Lys Val Gly Ser Arg Cys Cys Lys Leu Pro 450 455 460Glu Ser Glu
Arg Leu Pro Cys Ser Glu Asn His Leu Ala Leu Ala Leu465 470 475
480Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro Val Ser Glu Lys Ile
485 490 495Thr Lys Cys Cys Thr Asp Ser Leu Ala Glu Arg Arg Pro Cys
Phe Ser 500 505 510Ala Leu Glu Leu Asp Glu Gly Tyr Val Pro Lys Glu
Phe Lys Ala Glu 515 520 525Thr Phe Thr Phe His Ala Asp Ile Cys Thr
Leu Pro Glu Asp Glu Lys 530 535 540Gln Ile Lys Lys Gln Ser Ala Leu
Ala Glu Leu Val Lys His Lys Pro545 550 555 560Lys Ala Thr Lys Glu
Gln Leu Lys Thr Val Leu Gly Asn Phe Ser Ala 565 570 575Phe Val Ala
Lys Cys Cys Gly Arg Glu Asp Lys Glu Ala Cys Phe Ala 580 585 590Glu
Glu Gly Pro Lys Leu Val Ala Ser Ser Gln Leu Ala Leu Ala 595 600
60512607PRTEquus asinus 12Met Lys Trp Val Thr Phe Val Ser Leu Leu
Phe Leu Phe Ser Ser Ala1 5 10 15Tyr Phe Arg Gly Val Leu Arg Arg Asp
Thr His Lys Ser Glu Ile Ala 20 25 30His Arg Phe Asn Asp Leu Gly Glu
Lys His Phe Lys Gly Leu Val Leu 35 40 45Val Ala Phe Ser Gln Tyr Leu
Gln Gln Cys Pro Phe Glu Asp His Val 50 55 60Lys Leu Val Asn Glu Val
Thr Glu Phe Ala Lys Lys Cys Ala Ala Asp65 70 75 80Glu Ser Ala Glu
Asn Cys Asp Lys Ser Leu His Thr Leu Phe Gly Asp 85 90 95Lys Leu Cys
Thr Val Ala Thr Leu Arg Ala Thr Tyr Gly Glu Leu Ala 100 105 110Asp
Cys Cys Glu Lys Gln Glu Pro Glu Arg Asn Glu Cys Phe Leu Thr 115 120
125His Lys Asp Asp His Pro Asn Leu Pro Lys Leu Lys Pro Glu Pro Asp
130 135 140Ala Gln Cys Ala Ala Phe Gln Glu Asp Pro Asp Lys Phe Leu
Gly Lys145 150 155 160Tyr Leu Tyr Glu Val Ala Arg Arg His Pro Tyr
Phe Tyr Gly Pro Glu 165 170 175Leu Leu Phe His Ala Glu Glu Tyr Lys
Ala Asp Phe Thr Glu Cys Cys 180 185 190Pro Ala Asp Asp Lys Ala Gly
Cys Leu Ile Pro Lys Leu Asp Ala Leu 195 200 205Lys Glu Arg Ile Leu
Leu Ser Ser Ala Lys Glu Arg Leu Lys Cys Ser 210 215 220Ser Phe Gln
Lys Phe Gly Glu Arg Ala Phe Lys Ala Trp Ser Val Ala225 230 235
240Arg Leu Ser Gln Lys Phe Pro Lys Ala Asp Phe Ala Glu Val Ser Lys
245 250 255Ile Val Thr Asp Leu Thr Lys Val His Lys Glu Cys Cys His
Gly Asp 260 265 270Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp Leu Thr
Lys Tyr Ile Cys 275 280 285Glu His Gln Asp Ser Ile Ser Gly Lys Leu
Lys Ala Cys Cys Asp Lys 290 295 300Pro Leu Leu Gln Lys Ser His Cys
Ile Ala Glu Val Lys Glu Asp Asp305 310 315 320Leu Pro Ser Asp Leu
Pro Ala Leu Ala Ala Asp Phe Ala Glu Asp Lys 325 330 335Glu Ile Cys
Lys His Tyr Lys Asp Ala Lys Asp Val Phe Leu Gly Thr 340 345 350Phe
Leu Tyr Glu Tyr Ser Arg Arg His Pro Asp Tyr Ser Val Ser Leu 355 360
365Leu Leu Arg Ile Ala Lys Thr Tyr Glu Ala Thr Leu Glu Lys Cys Cys
370 375 380Ala Glu Ala Asp Pro Pro Ala Cys Tyr Ala Thr Val Phe Asp
Gln Phe385 390 395 400Thr Pro Leu Val Glu Glu Pro Lys Ser Leu Val
Lys Lys Asn Cys Asp 405 410 415Leu Phe Glu Glu Val Gly Glu Tyr Asp
Phe Gln Asn Ala Leu Ile Val 420 425 430Arg Tyr Thr Lys Lys Ala Pro
Gln Val Ser Thr Pro Thr Leu Val Glu 435 440 445Ile Gly Arg Thr Leu
Gly Lys Val Gly Ser Arg Cys Cys Lys Leu Pro 450 455 460Glu Ser Glu
Arg Leu Pro Cys Ser Glu Asn His Leu Ala Leu Ala Leu465 470 475
480Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro Val Ser Glu Lys Ile
485 490 495Thr Lys Cys Cys Thr Asp Ser Leu Ala Glu Arg Arg Pro Cys
Phe Ser 500 505 510Ala Leu Glu Leu Asp Glu Gly Tyr Ile Pro Lys Glu
Phe Lys Ala Glu 515 520 525Thr Phe Thr Phe His Ala Asp Ile Cys Thr
Leu Pro Glu Asp Glu Lys 530 535 540Gln Ile Lys Lys Gln Ser Ala Leu
Ala Glu Leu Val Lys His Lys Pro545 550 555 560Lys Ala Thr Lys Glu
Gln Leu Lys Thr Val Leu Gly Asn Phe Ser Ala 565 570 575Phe Val Ala
Lys Cys Cys Gly Ala Glu Asp Lys Glu Ala Cys Phe Ala 580 585 590Glu
Glu Gly Pro Lys Leu Val Ala Ser Ser Gln Leu Ala Leu Ala 595 600
60513608PRTOryctolagus cuniculus 13Met Lys Trp Val Thr Phe Ile Ser
Leu Leu Phe Leu Phe Ser Ser Ala1 5 10 15Tyr Ser Arg Gly Val Phe Arg
Arg Glu Ala His Lys Ser Glu Ile Ala 20 25 30His Arg Phe Asn Asp Val
Gly Glu Glu His Phe Ile Gly Leu Val Leu 35 40 45Ile Thr Phe Ser Gln
Tyr Leu Gln Lys Cys Pro Tyr Glu Glu His Ala 50 55 60Lys Leu Val Lys
Glu Val Thr Asp Leu Ala Lys Ala Cys Val Ala Asp65 70 75 80Glu Ser
Ala Ala Asn Cys Asp Lys Ser Leu His Asp Ile Phe Gly Asp 85 90 95Lys
Ile Cys Ala Leu Pro Ser Leu Arg Asp Thr Tyr Gly Asp Val Ala 100 105
110Asp Cys Cys Glu Lys Lys Glu Pro Glu Arg Asn Glu Cys Phe Leu His
115 120 125His Lys Asp Asp Lys Pro Asp Leu Pro Pro Phe Ala Arg Pro
Glu Ala 130 135 140Asp Val Leu Cys Lys Ala Phe His Asp Asp Glu Lys
Ala Phe Phe Gly145 150 155 160His Tyr Leu Tyr Glu Val Ala Arg Arg
His Pro Tyr Phe Tyr Ala Pro 165 170 175Glu Leu Leu Tyr Tyr Ala Gln
Lys Tyr Lys Ala Ile Leu Thr Glu Cys 180 185 190Cys Glu Ala Ala Asp
Lys Gly Ala Cys Leu Thr Pro Lys Leu Asp Ala 195 200 205Leu Glu Gly
Lys Ser Leu Ile Ser Ala Ala Gln Glu Arg Leu Arg Cys 210 215 220Ala
Ser Ile Gln Lys Phe Gly Asp Arg Ala Tyr Lys Ala Trp Ala Leu225 230
235 240Val Arg Leu Ser Gln Arg Phe Pro Lys Ala Asp Phe Thr Asp Ile
Ser 245 250 255Lys Ile Val Thr Asp Leu Thr Lys Val His Lys Glu Cys
Cys His Gly 260 265 270Asp Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp
Leu Ala Lys Tyr Met 275 280 285Cys Glu His Gln Glu Thr Ile Ser Ser
His Leu Lys Glu Cys Cys Asp 290 295 300Lys Pro Ile Leu Glu Lys Ala
His Cys Ile Tyr Gly Leu His Asn Asp305 310 315 320Glu Thr Pro Ala
Gly Leu Pro Ala Val Ala Glu Glu Phe Val Glu Asp 325 330 335Lys Asp
Val Cys Lys Asn Tyr Glu Glu Ala Lys Asp Leu Phe Leu Gly 340 345
350Lys Phe Leu Tyr Glu Tyr Ser Arg Arg His Pro Asp Tyr Ser Val Val
355 360 365Leu Leu Leu Arg Leu Gly Lys Ala Tyr Glu Ala Thr Leu Lys
Lys Cys 370 375 380Cys Ala Thr Asp Asp Pro His Ala Cys Tyr Ala Lys
Val Leu Asp Glu385 390 395
400Phe Gln Pro Leu Val Asp Glu Pro Lys Asn Leu Val Lys Gln Asn Cys
405 410 415Glu Leu Tyr Glu Gln Leu Gly Asp Tyr Asn Phe Gln Asn Ala
Leu Leu 420 425 430Val Arg Tyr Thr Lys Lys Val Pro Gln Val Ser Thr
Pro Thr Leu Val 435 440 445Glu Ile Ser Arg Ser Leu Gly Lys Val Gly
Ser Lys Cys Cys Lys His 450 455 460Pro Glu Ala Glu Arg Leu Pro Cys
Val Glu Asp Tyr Leu Ser Val Val465 470 475 480Leu Asn Arg Leu Cys
Val Leu His Glu Lys Thr Pro Val Ser Glu Lys 485 490 495Val Thr Lys
Cys Cys Ser Glu Ser Leu Val Asp Arg Arg Pro Cys Phe 500 505 510Ser
Ala Leu Gly Pro Asp Glu Thr Tyr Val Pro Lys Glu Phe Asn Ala 515 520
525Glu Thr Phe Thr Phe His Ala Asp Ile Cys Thr Leu Pro Glu Thr Glu
530 535 540Arg Lys Ile Lys Lys Gln Thr Ala Leu Val Glu Leu Val Lys
His Lys545 550 555 560Pro His Ala Thr Asn Asp Gln Leu Lys Thr Val
Val Gly Glu Phe Thr 565 570 575Ala Leu Leu Asp Lys Cys Cys Ser Ala
Glu Asp Lys Glu Ala Cys Phe 580 585 590Ala Val Glu Gly Pro Lys Leu
Val Glu Ser Ser Lys Ala Thr Leu Gly 595 600 60514583PRTCapra hircus
14Asp Thr His Lys Ser Glu Ile Ala His Arg Phe Asn Asp Leu Gly Glu1
5 10 15Glu Asn Phe Gln Gly Leu Val Leu Ile Ala Phe Ser Gln Tyr Leu
Gln 20 25 30Gln Cys Pro Phe Asp Glu His Val Lys Leu Val Lys Glu Leu
Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser His Ala Gly
Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Glu Leu Cys Lys
Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Asp Met Ala Asp Cys
Cys Glu Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Lys His
Lys Asp Asp Ser Pro Asp Leu 100 105 110Pro Lys Leu Lys Pro Glu Pro
Asp Thr Leu Cys Ala Glu Phe Lys Ala 115 120 125Asp Glu Lys Lys Phe
Trp Gly Lys Tyr Leu Tyr Glu Val Ala Arg Arg 130 135 140His Pro Tyr
Phe Tyr Ala Pro Glu Leu Leu Tyr Tyr Ala Asn Lys Tyr145 150 155
160Asn Gly Val Phe Gln Glu Cys Cys Gln Ala Glu Asp Lys Gly Ala Cys
165 170 175Leu Leu Pro Lys Ile Glu Thr Met Arg Glu Lys Val Leu Ala
Ser Ser 180 185 190Ala Arg Gln Arg Leu Arg Cys Ala Ser Ile Gln Lys
Phe Gly Glu Arg 195 200 205Ala Leu Lys Ala Trp Ser Val Ala Arg Leu
Ser Gln Lys Phe Pro Lys 210 215 220Ala Asp Phe Thr Asp Val Thr Lys
Ile Val Thr Asp Leu Thr Lys Val225 230 235 240His Lys Glu Cys Cys
His Gly Asp Leu Leu Glu Cys Ala Asp Asp Arg 245 250 255Ala Asp Leu
Ala Lys Tyr Ile Cys Asp His Gln Asp Thr Leu Ser Ser 260 265 270Lys
Leu Lys Glu Cys Cys Asp Lys Pro Val Leu Glu Lys Ser His Cys 275 280
285Ile Ala Glu Ile Asp Lys Asp Ala Val Pro Glu Asn Leu Pro Pro Leu
290 295 300Thr Ala Asp Phe Ala Glu Asp Lys Glu Val Cys Lys Asn Tyr
Gln Glu305 310 315 320Ala Lys Asp Val Phe Leu Gly Ser Phe Leu Tyr
Glu Tyr Ser Arg Arg 325 330 335His Pro Glu Tyr Ala Val Ser Val Leu
Leu Arg Leu Ala Lys Glu Tyr 340 345 350Glu Ala Thr Leu Glu Asp Cys
Cys Ala Lys Glu Asp Pro His Ala Cys 355 360 365Tyr Ala Thr Val Phe
Asp Lys Leu Lys His Leu Val Asp Glu Pro Gln 370 375 380Asn Leu Ile
Lys Lys Asn Cys Glu Leu Phe Glu Lys His Gly Glu Tyr385 390 395
400Gly Phe Gln Asn Ala Leu Ile Val Arg Tyr Thr Arg Lys Ala Pro Gln
405 410 415Val Ser Thr Pro Thr Leu Val Glu Ile Ser Arg Ser Leu Gly
Lys Val 420 425 430Gly Thr Lys Cys Cys Ala Lys Pro Glu Ser Glu Arg
Met Pro Cys Thr 435 440 445Glu Asp Tyr Leu Ser Leu Ile Leu Asn Arg
Leu Cys Val Leu His Glu 450 455 460Lys Thr Pro Val Ser Glu Lys Val
Thr Lys Cys Cys Thr Glu Ser Leu465 470 475 480Val Asn Arg Arg Pro
Cys Phe Ser Asp Leu Thr Leu Asp Glu Thr Tyr 485 490 495Val Pro Lys
Pro Phe Asp Gly Glu Ser Phe Thr Phe His Ala Asp Ile 500 505 510Cys
Thr Leu Pro Asp Thr Glu Lys Gln Ile Lys Lys Gln Thr Ala Leu 515 520
525Val Glu Leu Leu Lys His Lys Pro Lys Ala Thr Asp Glu Gln Leu Lys
530 535 540Thr Val Met Glu Asn Phe Val Ala Phe Val Asp Lys Cys Cys
Ala Ala545 550 555 560Asp Asp Lys Glu Gly Cys Phe Leu Leu Glu Gly
Pro Lys Leu Val Ala 565 570 575Ser Thr Gln Ala Ala Leu Ala
58015607PRTOvis aries 15Met Lys Trp Val Thr Phe Ile Ser Leu Leu Leu
Leu Phe Ser Ser Ala1 5 10 15Tyr Ser Arg Gly Val Phe Arg Arg Asp Thr
His Lys Ser Glu Ile Ala 20 25 30His Arg Phe Asn Asp Leu Gly Glu Glu
Asn Phe Gln Gly Leu Val Leu 35 40 45Ile Ala Phe Ser Gln Tyr Leu Gln
Gln Cys Pro Phe Asp Glu His Val 50 55 60Lys Leu Val Lys Glu Leu Thr
Glu Phe Ala Lys Thr Cys Val Ala Asp65 70 75 80Glu Ser His Ala Gly
Cys Asp Lys Ser Leu His Thr Leu Phe Gly Asp 85 90 95Glu Leu Cys Lys
Val Ala Thr Leu Arg Glu Thr Tyr Gly Asp Met Ala 100 105 110Asp Cys
Cys Glu Lys Gln Glu Pro Glu Arg Asn Glu Cys Phe Leu Asn 115 120
125His Lys Asp Asp Ser Pro Asp Leu Pro Lys Leu Lys Pro Glu Pro Asp
130 135 140Thr Leu Cys Ala Glu Phe Lys Ala Asp Glu Lys Lys Phe Trp
Gly Lys145 150 155 160Tyr Leu Tyr Glu Val Ala Arg Arg His Pro Tyr
Phe Tyr Ala Pro Glu 165 170 175Leu Leu Tyr Tyr Ala Asn Lys Tyr Asn
Gly Val Phe Gln Glu Cys Cys 180 185 190Gln Ala Glu Asp Lys Gly Ala
Cys Leu Leu Pro Lys Ile Asp Ala Met 195 200 205Arg Glu Lys Val Leu
Ala Ser Ser Ala Arg Gln Arg Leu Arg Cys Ala 210 215 220Ser Ile Gln
Lys Phe Gly Glu Arg Ala Leu Lys Ala Trp Ser Val Ala225 230 235
240Arg Leu Ser Gln Lys Phe Pro Lys Ala Asp Phe Thr Asp Val Thr Lys
245 250 255Ile Val Thr Asp Leu Thr Lys Val His Lys Glu Cys Cys His
Gly Asp 260 265 270Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala
Lys Tyr Ile Cys 275 280 285Asp His Gln Asp Ala Leu Ser Ser Lys Leu
Lys Glu Cys Cys Asp Lys 290 295 300Pro Val Leu Glu Lys Ser His Cys
Ile Ala Glu Val Asp Lys Asp Ala305 310 315 320Val Pro Glu Asn Leu
Pro Pro Leu Thr Ala Asp Phe Ala Glu Asp Lys 325 330 335Glu Val Cys
Lys Asn Tyr Gln Glu Ala Lys Asp Val Phe Leu Gly Ser 340 345 350Phe
Leu Tyr Glu Tyr Ser Arg Arg His Pro Glu Tyr Ala Val Ser Val 355 360
365Leu Leu Arg Leu Ala Lys Glu Tyr Glu Ala Thr Leu Glu Asp Cys Cys
370 375 380Ala Lys Glu Asp Pro His Ala Cys Tyr Ala Thr Val Phe Asp
Lys Leu385 390 395 400Lys His Leu Val Asp Glu Pro Gln Asn Leu Ile
Lys Lys Asn Cys Glu 405 410 415Leu Phe Glu Lys His Gly Glu Tyr Gly
Phe Gln Asn Ala Leu Ile Val 420 425 430Arg Tyr Thr Arg Lys Ala Pro
Gln Val Ser Thr Pro Thr Leu Val Glu 435 440 445Ile Ser Arg Ser Leu
Gly Lys Val Gly Thr Lys Cys Cys Ala Lys Pro 450 455 460Glu Ser Glu
Arg Met Pro Cys Thr Glu Asp Tyr Leu Ser Leu Ile Leu465 470 475
480Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro Val Ser Glu Lys Val
485 490 495Thr Lys Cys Cys Thr Glu Ser Leu Val Asn Arg Arg Pro Cys
Phe Ser 500 505 510Asp Leu Thr Leu Asp Glu Thr Tyr Val Pro Lys Pro
Phe Asp Glu Lys 515 520 525Phe Phe Thr Phe His Ala Asp Ile Cys Thr
Leu Pro Asp Thr Glu Lys 530 535 540Gln Ile Lys Lys Gln Thr Ala Leu
Val Glu Leu Leu Lys His Lys Pro545 550 555 560Lys Ala Thr Asp Glu
Gln Leu Lys Thr Val Met Glu Asn Phe Val Ala 565 570 575Phe Val Asp
Lys Cys Cys Ala Ala Asp Asp Lys Glu Gly Cys Phe Val 580 585 590Leu
Glu Gly Pro Lys Leu Val Ala Ser Thr Gln Ala Ala Leu Ala 595 600
60516608PRTcanis lupus familiaris 16Met Lys Trp Val Thr Phe Ile Ser
Leu Phe Phe Leu Phe Ser Ser Ala1 5 10 15Tyr Ser Arg Gly Leu Val Arg
Arg Glu Ala Tyr Lys Ser Glu Ile Ala 20 25 30His Arg Tyr Asn Asp Leu
Gly Glu Glu His Phe Arg Gly Leu Val Leu 35 40 45Val Ala Phe Ser Gln
Tyr Leu Gln Gln Cys Pro Phe Glu Asp His Val 50 55 60Lys Leu Ala Lys
Glu Val Thr Glu Phe Ala Lys Ala Cys Ala Ala Glu65 70 75 80Glu Ser
Gly Ala Asn Cys Asp Lys Ser Leu His Thr Leu Phe Gly Asp 85 90 95Lys
Leu Cys Thr Val Ala Ser Leu Arg Asp Lys Tyr Gly Asp Met Ala 100 105
110Asp Cys Cys Glu Lys Gln Glu Pro Asp Arg Asn Glu Cys Phe Leu Ala
115 120 125His Lys Asp Asp Asn Pro Gly Phe Pro Pro Leu Val Ala Pro
Glu Pro 130 135 140Asp Ala Leu Cys Ala Ala Phe Gln Asp Asn Glu Gln
Leu Phe Leu Gly145 150 155 160Lys Tyr Leu Tyr Glu Ile Ala Arg Arg
His Pro Tyr Phe Tyr Ala Pro 165 170 175Glu Leu Leu Tyr Tyr Ala Gln
Gln Tyr Lys Gly Val Phe Ala Glu Cys 180 185 190Cys Gln Ala Ala Asp
Lys Ala Ala Cys Leu Gly Pro Lys Ile Glu Ala 195 200 205Leu Arg Glu
Lys Val Leu Leu Ser Ser Ala Lys Glu Arg Phe Lys Cys 210 215 220Ala
Ser Leu Gln Lys Phe Gly Asp Arg Ala Phe Lys Ala Trp Ser Val225 230
235 240Ala Arg Leu Ser Gln Arg Phe Pro Lys Ala Asp Phe Ala Glu Ile
Ser 245 250 255Lys Val Val Thr Asp Leu Thr Lys Val His Lys Glu Cys
Cys His Gly 260 265 270Asp Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp
Leu Ala Lys Tyr Met 275 280 285Cys Glu Asn Gln Asp Ser Ile Ser Thr
Lys Leu Lys Glu Cys Cys Asp 290 295 300Lys Pro Val Leu Glu Lys Ser
Gln Cys Leu Ala Glu Val Glu Arg Asp305 310 315 320Glu Leu Pro Gly
Asp Leu Pro Ser Leu Ala Ala Asp Phe Val Glu Asp 325 330 335Lys Glu
Val Cys Lys Asn Tyr Gln Glu Ala Lys Asp Val Phe Leu Gly 340 345
350Thr Phe Leu Tyr Glu Tyr Ala Arg Arg His Pro Glu Tyr Ser Val Ser
355 360 365Leu Leu Leu Arg Leu Ala Lys Glu Tyr Glu Ala Thr Leu Glu
Lys Cys 370 375 380Cys Ala Thr Asp Asp Pro Pro Thr Cys Tyr Ala Lys
Val Leu Asp Glu385 390 395 400Phe Lys Pro Leu Val Asp Glu Pro Gln
Asn Leu Val Lys Thr Asn Cys 405 410 415Glu Leu Phe Glu Lys Leu Gly
Glu Tyr Gly Phe Gln Asn Ala Leu Leu 420 425 430Val Arg Tyr Thr Lys
Lys Ala Pro Gln Val Ser Thr Pro Thr Leu Val 435 440 445Glu Val Ser
Arg Lys Leu Gly Lys Val Gly Thr Lys Cys Cys Lys Lys 450 455 460Pro
Glu Ser Glu Arg Met Ser Cys Ala Glu Asp Phe Leu Ser Val Val465 470
475 480Leu Asn Arg Leu Cys Val Leu His Glu Lys Thr Pro Val Ser Glu
Arg 485 490 495Val Thr Lys Cys Cys Ser Glu Ser Leu Val Asn Arg Arg
Pro Cys Phe 500 505 510Ser Gly Leu Glu Val Asp Glu Thr Tyr Val Pro
Lys Glu Phe Asn Ala 515 520 525Glu Thr Phe Thr Phe His Ala Asp Leu
Cys Thr Leu Pro Glu Ala Glu 530 535 540Lys Gln Val Lys Lys Gln Thr
Ala Leu Val Glu Leu Leu Lys His Lys545 550 555 560Pro Lys Ala Thr
Asp Glu Gln Leu Lys Thr Val Met Gly Asp Phe Gly 565 570 575Ala Phe
Val Glu Lys Cys Cys Ala Ala Glu Asn Lys Glu Gly Cys Phe 580 585
590Ser Glu Glu Gly Pro Lys Leu Val Ala Ala Ala Gln Ala Ala Leu Val
595 600 60517615PRTGallus gallus 17Met Lys Trp Val Thr Leu Ile Ser
Phe Ile Phe Leu Phe Ser Ser Ala1 5 10 15Thr Ser Arg Asn Leu Gln Arg
Phe Ala Arg Asp Ala Glu His Lys Ser 20 25 30Glu Ile Ala His Arg Tyr
Asn Asp Leu Lys Glu Glu Thr Phe Lys Ala 35 40 45Val Ala Met Ile Thr
Phe Ala Gln Tyr Leu Gln Arg Cys Ser Tyr Glu 50 55 60Gly Leu Ser Lys
Leu Val Lys Asp Val Val Asp Leu Ala Gln Lys Cys65 70 75 80Val Ala
Asn Glu Asp Ala Pro Glu Cys Ser Lys Pro Leu Pro Ser Ile 85 90 95Ile
Leu Asp Glu Ile Cys Gln Val Glu Lys Leu Arg Asp Ser Tyr Gly 100 105
110Ala Met Ala Asp Cys Cys Ser Lys Ala Asp Pro Glu Arg Asn Glu Cys
115 120 125Phe Leu Ser Phe Lys Val Ser Gln Pro Asp Phe Val Gln Pro
Tyr Gln 130 135 140Arg Pro Ala Ser Asp Val Ile Cys Gln Glu Tyr Gln
Asp Asn Arg Val145 150 155 160Ser Phe Leu Gly His Phe Ile Tyr Ser
Val Ala Arg Arg His Pro Phe 165 170 175Leu Tyr Ala Pro Ala Ile Leu
Ser Phe Ala Val Asp Phe Glu His Ala 180 185 190Leu Gln Ser Cys Cys
Lys Glu Ser Asp Val Gly Ala Cys Leu Asp Thr 195 200 205Lys Glu Ile
Val Met Arg Glu Lys Ala Lys Gly Val Ser Val Lys Gln 210 215 220Gln
Tyr Phe Cys Gly Ile Leu Lys Gln Phe Gly Asp Arg Val Phe Gln225 230
235 240Ala Arg Gln Leu Ile Tyr Leu Ser Gln Lys Tyr Pro Lys Ala Pro
Phe 245 250 255Ser Glu Val Ser Lys Phe Val His Asp Ser Ile Gly Val
His Lys Glu 260 265 270Cys Cys Glu Gly Asp Met Val Glu Cys Met Asp
Asp Met Ala Arg Met 275 280 285Met Ser Asn Leu Cys Ser Gln Gln Asp
Val Phe Ser Gly Lys Ile Lys 290 295 300Asp Cys Cys Glu Lys Pro Ile
Val Glu Arg Ser Gln Cys Ile Met Glu305 310 315 320Ala Glu Phe Asp
Glu Lys Pro Ala Asp Leu Pro Ser Leu Val Glu Lys 325 330 335Tyr Ile
Glu Asp Lys Glu Val Cys Lys Ser Phe Glu Ala Gly His Asp 340 345
350Ala Phe Met Ala Glu Phe Val Tyr Glu Tyr Ser Arg Arg His Pro Glu
355 360 365Phe Ser Ile Gln Leu Ile Met Arg Ile Ala Lys Gly Tyr Glu
Ser Leu 370 375 380Leu Glu Lys Cys Cys Lys Thr Asp Asn Pro Ala Glu
Cys Tyr Ala Asn385 390 395 400Ala Gln Glu Gln Leu Asn Gln His Ile
Lys Glu Thr Gln Asp Val Val 405 410 415Lys Thr Asn Cys Asp Leu Leu
His Asp His Gly Glu Ala Asp Phe Leu 420 425 430Lys Ser Ile Leu Ile
Arg Tyr Thr Lys Lys Met Pro Gln Val Pro Thr 435 440 445Asp Leu Leu
Leu Glu Thr Gly Lys Lys Met Thr Thr Ile Gly Thr Lys 450
455 460Cys Cys Gln Leu Gly Glu Asp Arg Arg Met Ala Cys Ser Glu Gly
Tyr465 470 475 480Leu Ser Ile Val Ile His Asp Thr Cys Arg Lys Gln
Glu Thr Thr Pro 485 490 495Ile Asn Asp Asn Val Ser Gln Cys Cys Ser
Gln Leu Tyr Ala Asn Arg 500 505 510Arg Pro Cys Phe Thr Ala Met Gly
Val Asp Thr Lys Tyr Val Pro Pro 515 520 525Pro Phe Asn Pro Asp Met
Phe Ser Phe Asp Glu Lys Leu Cys Ser Ala 530 535 540Pro Ala Glu Glu
Arg Glu Val Gly Gln Met Lys Leu Leu Ile Asn Leu545 550 555 560Ile
Lys Arg Lys Pro Gln Met Thr Glu Glu Gln Ile Lys Thr Ile Ala 565 570
575Asp Gly Phe Thr Ala Met Val Asp Lys Cys Cys Lys Gln Ser Asp Ile
580 585 590Asn Thr Cys Phe Gly Glu Glu Gly Ala Asn Leu Ile Val Gln
Ser Arg 595 600 605Ala Thr Leu Gly Ile Gly Ala 610 61518607PRTSus
scrofa 18Met Lys Trp Val Thr Phe Ile Ser Leu Leu Phe Leu Phe Ser
Ser Ala1 5 10 15Tyr Ser Arg Gly Val Phe Arg Arg Asp Thr Tyr Lys Ser
Glu Ile Ala 20 25 30His Arg Phe Lys Asp Leu Gly Glu Gln Tyr Phe Lys
Gly Leu Val Leu 35 40 45Ile Ala Phe Ser Gln His Leu Gln Gln Cys Pro
Tyr Glu Glu His Val 50 55 60Lys Leu Val Arg Glu Val Thr Glu Phe Ala
Lys Thr Cys Val Ala Asp65 70 75 80Glu Ser Ala Glu Asn Cys Asp Lys
Ser Ile His Thr Leu Phe Gly Asp 85 90 95Lys Leu Cys Ala Ile Pro Ser
Leu Arg Glu His Tyr Gly Asp Leu Ala 100 105 110Asp Cys Cys Glu Lys
Glu Glu Pro Glu Arg Asn Glu Cys Phe Leu Gln 115 120 125His Lys Asn
Asp Asn Pro Asp Ile Pro Lys Leu Lys Pro Asp Pro Val 130 135 140Ala
Leu Cys Ala Asp Phe Gln Glu Asp Glu Gln Lys Phe Trp Gly Lys145 150
155 160Tyr Leu Tyr Glu Ile Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro
Glu 165 170 175Leu Leu Tyr Tyr Ala Ile Ile Tyr Lys Asp Val Phe Ser
Glu Cys Cys 180 185 190Gln Ala Ala Asp Lys Ala Ala Cys Leu Leu Pro
Lys Ile Glu His Leu 195 200 205Arg Glu Lys Val Leu Thr Ser Ala Ala
Lys Gln Arg Leu Lys Cys Ala 210 215 220Ser Ile Gln Lys Phe Gly Glu
Arg Ala Phe Lys Ala Trp Ser Leu Ala225 230 235 240Arg Leu Ser Gln
Arg Phe Pro Lys Ala Asp Phe Thr Glu Ile Ser Lys 245 250 255Ile Val
Thr Asp Leu Ala Lys Val His Lys Glu Cys Cys His Gly Asp 260 265
270Leu Leu Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala Lys Tyr Ile Cys
275 280 285Glu Asn Gln Asp Thr Ile Ser Thr Lys Leu Lys Glu Cys Cys
Asp Lys 290 295 300Pro Leu Leu Glu Lys Ser His Cys Ile Ala Glu Ala
Lys Arg Asp Glu305 310 315 320Leu Pro Ala Asp Leu Asn Pro Leu Glu
His Asp Phe Val Glu Asp Lys 325 330 335Glu Val Cys Lys Asn Tyr Lys
Glu Ala Lys His Val Phe Leu Gly Thr 340 345 350Phe Leu Tyr Glu Tyr
Ser Arg Arg His Pro Asp Tyr Ser Val Ser Leu 355 360 365Leu Leu Arg
Ile Ala Lys Ile Tyr Glu Ala Thr Leu Glu Asp Cys Cys 370 375 380Ala
Lys Glu Asp Pro Pro Ala Cys Tyr Ala Thr Val Phe Asp Lys Phe385 390
395 400Gln Pro Leu Val Asp Glu Pro Lys Asn Leu Ile Lys Gln Asn Cys
Glu 405 410 415Leu Phe Glu Lys Leu Gly Glu Tyr Gly Phe Gln Asn Ala
Leu Ile Val 420 425 430Arg Tyr Thr Lys Lys Val Pro Gln Val Ser Thr
Pro Thr Leu Val Glu 435 440 445Val Ala Arg Lys Leu Gly Leu Val Gly
Ser Arg Cys Cys Lys Arg Pro 450 455 460Glu Glu Glu Arg Leu Ser Cys
Ala Glu Asp Tyr Leu Ser Leu Val Leu465 470 475 480Asn Arg Leu Cys
Val Leu His Glu Lys Thr Pro Val Ser Glu Lys Val 485 490 495Thr Lys
Cys Cys Thr Glu Ser Leu Val Asn Arg Arg Pro Cys Phe Ser 500 505
510Ala Leu Thr Pro Asp Glu Thr Tyr Lys Pro Lys Glu Phe Val Glu Gly
515 520 525Thr Phe Thr Phe His Ala Asp Leu Cys Thr Leu Pro Glu Asp
Glu Lys 530 535 540Gln Ile Lys Lys Gln Thr Ala Leu Val Glu Leu Leu
Lys His Lys Pro545 550 555 560His Ala Thr Glu Glu Gln Leu Arg Thr
Val Leu Gly Asn Phe Ala Ala 565 570 575Phe Val Gln Lys Cys Cys Ala
Ala Pro Asp His Glu Ala Cys Phe Ala 580 585 590Val Glu Gly Pro Lys
Phe Val Ile Glu Ile Arg Gly Ile Leu Ala 595 600
60519205PRTArtificial sequenceArtificial albumin variant human
serum albumin domain 3 19Val Glu Glu Pro Gln Asn Leu Ile Lys Gln
Asn Cys Glu Leu Phe Glu1 5 10 15Gln Leu Gly Glu Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr 20 25 30Lys Lys Val Pro Gln Val Ser Thr
Pro Thr Leu Val Glu Val Ser Arg 35 40 45Asn Leu Gly Lys Val Gly Ser
Lys Cys Cys Lys His Pro Glu Ala Lys 50 55 60Arg Met Pro Cys Ala Glu
Asp Tyr Leu Ser Val Val Leu Asn Gln Leu65 70 75 80Cys Val Leu His
Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys 85 90 95Cys Thr Glu
Ser Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu 100 105 110Val
Asp Glu Thr Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr 115 120
125Phe His Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys
130 135 140Lys Gln Thr Ala Leu Val Glu Leu Val Lys His Lys Pro Lys
Ala Thr145 150 155 160Lys Glu Gln Leu Lys Ala Val Met Asp Asp Phe
Ala Ala Phe Val Glu 165 170 175Lys Cys Cys Lys Ala Asp Asp Lys Glu
Thr Cys Phe Ala Glu Glu Gly 180 185 190Lys Lys Leu Val Ala Ala Ser
Gln Ala Ala Leu Gly Leu 195 200 20520403PRTArtificial
sequenceArtificial albumin variant human serum albumin domain 2 and
human serum albumin domain 3 20Asp Glu Leu Arg Asp Glu Gly Lys Ala
Ser Ser Ala Lys Gln Arg Leu1 5 10 15Lys Cys Ala Ser Leu Gln Lys Phe
Gly Glu Arg Ala Phe Lys Ala Trp 20 25 30Ala Val Ala Arg Leu Ser Gln
Arg Phe Pro Lys Ala Glu Phe Ala Glu 35 40 45Val Ser Lys Leu Val Thr
Asp Leu Thr Lys Val His Thr Glu Cys Cys 50 55 60His Gly Asp Leu Leu
Glu Cys Ala Asp Asp Arg Ala Asp Leu Ala Lys65 70 75 80Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser Ser Lys Leu Lys Glu Cys 85 90 95Cys Glu
Lys Pro Leu Leu Glu Lys Ser His Cys Ile Ala Glu Val Glu 100 105
110Asn Asp Glu Met Pro Ala Asp Leu Pro Ser Leu Ala Ala Asp Phe Val
115 120 125Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala Glu Ala Lys Asp
Val Phe 130 135 140Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg Arg His
Pro Asp Tyr Ser145 150 155 160Val Val Leu Leu Leu Arg Leu Ala Lys
Thr Tyr Glu Thr Thr Leu Glu 165 170 175Lys Cys Cys Ala Ala Ala Asp
Pro His Glu Cys Tyr Ala Lys Val Phe 180 185 190Asp Glu Phe Lys Pro
Leu Val Glu Glu Pro Gln Asn Leu Ile Lys Gln 195 200 205Asn Cys Glu
Leu Phe Glu Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala 210 215 220Leu
Leu Val Arg Tyr Thr Lys Lys Val Pro Gln Val Ser Thr Pro Thr225 230
235 240Leu Val Glu Val Ser Arg Asn Leu Gly Lys Val Gly Ser Lys Cys
Cys 245 250 255Lys His Pro Glu Ala Lys Arg Met Pro Cys Ala Glu Asp
Tyr Leu Ser 260 265 270Val Val Leu Asn Gln Leu Cys Val Leu His Glu
Lys Thr Pro Val Ser 275 280 285Asp Arg Val Thr Lys Cys Cys Thr Glu
Ser Leu Val Asn Arg Arg Pro 290 295 300Cys Phe Ser Ala Leu Glu Val
Asp Glu Thr Tyr Val Pro Lys Glu Phe305 310 315 320Asn Ala Glu Thr
Phe Thr Phe His Ala Asp Ile Cys Thr Leu Ser Glu 325 330 335Lys Glu
Arg Gln Ile Lys Lys Gln Thr Ala Leu Val Glu Leu Val Lys 340 345
350His Lys Pro Lys Ala Thr Lys Glu Gln Leu Lys Ala Val Met Asp Asp
355 360 365Phe Ala Ala Phe Val Glu Lys Cys Cys Lys Ala Asp Asp Lys
Glu Thr 370 375 380Cys Phe Ala Glu Glu Gly Lys Lys Leu Val Ala Ala
Ser Gln Ala Ala385 390 395 400Leu Gly Leu21399PRTArtificial
sequenceArtificial albumin variant human serum albumin domain 1 and
human serum albumin domain 3 21Asp Ala His Lys Ser Glu Val Ala His
Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu
Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Cys Pro Phe Glu Asp His
Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val
Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu
Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr
Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg
Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105
110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His
115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile
Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe
Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys
Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp
Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Val Glu Glu
Pro Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu 195 200 205Phe Glu Gln
Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg 210 215 220Tyr
Thr Lys Lys Val Pro Gln Val Ser Thr Pro Thr Leu Val Glu Val225 230
235 240Ser Arg Asn Leu Gly Lys Val Gly Ser Lys Cys Cys Lys His Pro
Glu 245 250 255Ala Lys Arg Met Pro Cys Ala Glu Asp Tyr Leu Ser Val
Val Leu Asn 260 265 270Gln Leu Cys Val Leu His Glu Lys Thr Pro Val
Ser Asp Arg Val Thr 275 280 285Lys Cys Cys Thr Glu Ser Leu Val Asn
Arg Arg Pro Cys Phe Ser Ala 290 295 300Leu Glu Val Asp Glu Thr Tyr
Val Pro Lys Glu Phe Asn Ala Glu Thr305 310 315 320Phe Thr Phe His
Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln 325 330 335Ile Lys
Lys Gln Thr Ala Leu Val Glu Leu Val Lys His Lys Pro Lys 340 345
350Ala Thr Lys Glu Gln Leu Lys Ala Val Met Asp Asp Phe Ala Ala Phe
355 360 365Val Glu Lys Cys Cys Lys Ala Asp Asp Lys Glu Thr Cys Phe
Ala Glu 370 375 380Glu Gly Lys Lys Leu Val Ala Ala Ser Gln Ala Ala
Leu Gly Leu385 390 39522410PRTArtificial sequenceArtificial albumin
variant two consecutive copies of human serum albumin domain 3
22Val Glu Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu1
5 10 15Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr
Thr 20 25 30Lys Lys Val Pro Gln Val Ser Thr Pro Thr Leu Val Glu Val
Ser Arg 35 40 45Asn Leu Gly Lys Val Gly Ser Lys Cys Cys Lys His Pro
Glu Ala Lys 50 55 60Arg Met Pro Cys Ala Glu Asp Tyr Leu Ser Val Val
Leu Asn Gln Leu65 70 75 80Cys Val Leu His Glu Lys Thr Pro Val Ser
Asp Arg Val Thr Lys Cys 85 90 95Cys Thr Glu Ser Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu 100 105 110Val Asp Glu Thr Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr 115 120 125Phe His Ala Asp Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys 130 135 140Lys Gln Thr
Ala Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr145 150 155
160Lys Glu Gln Leu Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu
165 170 175Lys Cys Cys Lys Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu
Glu Gly 180 185 190Lys Lys Leu Val Ala Ala Ser Gln Ala Ala Leu Gly
Leu Val Glu Glu 195 200 205Pro Gln Asn Leu Ile Lys Gln Asn Cys Glu
Leu Phe Glu Gln Leu Gly 210 215 220Glu Tyr Lys Phe Gln Asn Ala Leu
Leu Val Arg Tyr Thr Lys Lys Val225 230 235 240Pro Gln Val Ser Thr
Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly 245 250 255Lys Val Gly
Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro 260 265 270Cys
Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu 275 280
285His Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu
290 295 300Ser Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val
Asp Glu305 310 315 320Thr Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr
Phe Thr Phe His Ala 325 330 335Asp Ile Cys Thr Leu Ser Glu Lys Glu
Arg Gln Ile Lys Lys Gln Thr 340 345 350Ala Leu Val Glu Leu Val Lys
His Lys Pro Lys Ala Thr Lys Glu Gln 355 360 365Leu Lys Ala Val Met
Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys 370 375 380Lys Ala Asp
Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu385 390 395
400Val Ala Ala Ser Gln Ala Ala Leu Gly Leu 405
41023615PRTArtificial SequenceHSA Domain III + HSA Domain III + HSA
Domain III 23Val Glu Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys Glu
Leu Phe Glu1 5 10 15Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu
Val Arg Tyr Thr 20 25 30Lys Lys Val Pro Gln Val Ser Thr Pro Thr Leu
Val Glu Val Ser Arg 35 40 45Asn Leu Gly Lys Val Gly Ser Lys Cys Cys
Lys His Pro Glu Ala Lys 50 55 60Arg Met Pro Cys Ala Glu Asp Tyr Leu
Ser Val Val Leu Asn Gln Leu65 70 75 80Cys Val Leu His Glu Lys Thr
Pro Val Ser Asp Arg Val Thr Lys Cys 85 90 95Cys Thr Glu Ser Leu Val
Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu 100 105 110Val Asp Glu Thr
Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr 115 120 125Phe His
Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys 130 135
140Lys Gln Thr Ala Leu Val Glu Leu Val Lys His Lys Pro Lys Ala
Thr145 150 155 160Lys Glu Gln Leu Lys Ala Val Met Asp Asp Phe Ala
Ala Phe Val Glu 165 170 175Lys Cys Cys Lys Ala Asp Asp Lys Glu Thr
Cys Phe Ala Glu Glu Gly 180 185 190Lys Lys Leu Val Ala Ala Ser Gln
Ala Ala Leu Gly Leu Val Glu Glu 195 200 205Pro Gln Asn Leu Ile Lys
Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly 210 215 220Glu Tyr Lys
Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val225 230 235
240Pro Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly
245 250 255Lys Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg
Met Pro 260 265 270Cys Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln
Leu Cys Val Leu 275 280 285His Glu Lys Thr Pro Val Ser Asp Arg Val
Thr Lys Cys Cys Thr Glu 290 295 300Ser Leu Val Asn Arg Arg Pro Cys
Phe Ser Ala Leu Glu Val Asp Glu305 310 315 320Thr Tyr Val Pro Lys
Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala 325 330 335Asp Ile Cys
Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr 340 345 350Ala
Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln 355 360
365Leu Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys
370 375 380Lys Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys
Lys Leu385 390 395 400Val Ala Ala Ser Gln Ala Ala Leu Gly Leu Val
Glu Glu Pro Gln Asn 405 410 415Leu Ile Lys Gln Asn Cys Glu Leu Phe
Glu Gln Leu Gly Glu Tyr Lys 420 425 430Phe Gln Asn Ala Leu Leu Val
Arg Tyr Thr Lys Lys Val Pro Gln Val 435 440 445Ser Thr Pro Thr Leu
Val Glu Val Ser Arg Asn Leu Gly Lys Val Gly 450 455 460Ser Lys Cys
Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys Ala Glu465 470 475
480Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His Glu Lys
485 490 495Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser
Leu Val 500 505 510Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp
Glu Thr Tyr Val 515 520 525Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr
Phe His Ala Asp Ile Cys 530 535 540Thr Leu Ser Glu Lys Glu Arg Gln
Ile Lys Lys Gln Thr Ala Leu Val545 550 555 560Glu Leu Val Lys His
Lys Pro Lys Ala Thr Lys Glu Gln Leu Lys Ala 565 570 575Val Met Asp
Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys Ala Asp 580 585 590Asp
Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val Ala Ala 595 600
605Ser Gln Ala Ala Leu Gly Leu 610 61524604PRTArtificial
SequenceHSA Domain I + HSA Domain III + HSA Domain III 24Asp Ala
His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu
Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25
30Gln Cys Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu
35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp
Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala
Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala
Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp
Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp
Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu
Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe
Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys
Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170
175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser
180 185 190Ser Ala Val Glu Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu 195 200 205Phe Glu Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala
Leu Leu Val Arg 210 215 220Tyr Thr Lys Lys Val Pro Gln Val Ser Thr
Pro Thr Leu Val Glu Val225 230 235 240Ser Arg Asn Leu Gly Lys Val
Gly Ser Lys Cys Cys Lys His Pro Glu 245 250 255Ala Lys Arg Met Pro
Cys Ala Glu Asp Tyr Leu Ser Val Val Leu Asn 260 265 270Gln Leu Cys
Val Leu His Glu Lys Thr Pro Val Ser Asp Arg Val Thr 275 280 285Lys
Cys Cys Thr Glu Ser Leu Val Asn Arg Arg Pro Cys Phe Ser Ala 290 295
300Leu Glu Val Asp Glu Thr Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr305 310 315 320Phe Thr Phe His Ala Asp Ile Cys Thr Leu Ser Glu
Lys Glu Arg Gln 325 330 335Ile Lys Lys Gln Thr Ala Leu Val Glu Leu
Val Lys His Lys Pro Lys 340 345 350Ala Thr Lys Glu Gln Leu Lys Ala
Val Met Asp Asp Phe Ala Ala Phe 355 360 365Val Glu Lys Cys Cys Lys
Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu 370 375 380Glu Gly Lys Lys
Leu Val Ala Ala Ser Gln Ala Ala Leu Gly Leu Val385 390 395 400Glu
Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln 405 410
415Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys
420 425 430Lys Val Pro Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser
Arg Asn 435 440 445Leu Gly Lys Val Gly Ser Lys Cys Cys Lys His Pro
Glu Ala Lys Arg 450 455 460Met Pro Cys Ala Glu Asp Tyr Leu Ser Val
Val Leu Asn Gln Leu Cys465 470 475 480Val Leu His Glu Lys Thr Pro
Val Ser Asp Arg Val Thr Lys Cys Cys 485 490 495Thr Glu Ser Leu Val
Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val 500 505 510Asp Glu Thr
Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe 515 520 525His
Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys 530 535
540Gln Thr Ala Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys545 550 555 560Glu Gln Leu Lys Ala Val Met Asp Asp Phe Ala Ala
Phe Val Glu Lys 565 570 575Cys Cys Lys Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys 580 585 590Lys Leu Val Ala Ala Ser Gln Ala
Ala Leu Gly Leu 595 600251758DNAArtificial SequenceHSA coding
sequence mutated to introduce restriction enzyme sites 25gatgcacaca
agagtgaggt tgctcatcgg tttaaagatt tgggagaaga aaatttcaaa 60gccttggtgt
tgattgcctt tgctcagtat cttcagcagt gtccatttga agatcatgta
120aaattagtga atgaagtaac tgaatttgca aaaacatgtg ttgctgatga
gtccgcggaa 180aattgtgaca aatcacttca tacccttttt ggagacaaat
tatgcacagt tgcaactctt 240cgtgaaacct atggtgaaat ggctgactgc
tgtgcaaaac aagaacctga gagaaatgaa 300tgcttcttgc aacacaaaga
tgacaaccca aacctccccc gattggtgag accagaggtt 360gatgtgatgt
gcactgcttt tcatgacaat gaagagacat ttttgaaaaa atacttatat
420gaaattgcca gaagacatcc ttacttttat gccccggaac tccttttctt
tgctaaaagg 480tataaagctg cttttacaga atgttgccaa gctgctgata
aagctgcctg cctgttgcca 540aagctcgatg aacttcggga tgaagggaag
gctagctctg ccaaacagag actcaagtgt 600gccagtctcc aaaaatttgg
agaaagagct ttcaaagcat gggcagtagc tcgcctgagc 660cagagatttc
ccaaagctga gtttgcagaa gtttccaagt tagtgacaga tcttaccaaa
720gtccacacgg aatgctgcca tggagatctg ctcgagtgtg ctgatgacag
ggcggacctt 780gccaagtata tctgtgaaaa tcaagattcg atctccagta
aactgaagga atgctgtgaa 840aaacctctgt tggaaaaatc ccactgcatt
gccgaagtgg aaaatgatga gatgcctgct 900gacttgcctt cattagctgc
tgattttgtt gaaagtaagg atgtttgcaa aaactatgct 960gaggcaaagg
atgtcttcct gggcatgttt ttgtatgaat atgcaagaag gcatcctgat
1020tactctgtcg tgctgctgct gagacttgcc aagacatatg aaaccactct
agagaagtgc 1080tgtgccgctg ctgatcctca tgaatgctat gccaaagtgt
tcgatgaatt taaacctctt 1140gtggaagagc ctcagaattt aatcaaacaa
aattgtgagc tttttgagca gcttggagag 1200tacaaattcc agaatgcgct
attagttcgt tacaccaaga aagtacccca agtgtcaact 1260ccaactcttg
tagaggtctc aagaaaccta ggaaaagtgg gatccaaatg ttgtaaacat
1320cctgaagcaa aaagaatgcc ctgtgcagaa gactatctat ccgtggtcct
gaaccagtta 1380tgtgtgttgc atgagaaaac gccagtaagt gacagagtca
ccaaatgctg cacagaatcc 1440ttggtgaaca ggcgaccatg cttttcagct
ctggaagtcg acgaaacata cgttcccaaa 1500gagtttaatg ctgaaacatt
caccttccat gcagatatat gcacactttc tgagaaggag 1560agacaaatca
agaaacaaac tgcacttgtt gagctcgtga aacacaagcc caaggcaaca
1620aaagagcaac tgaaagctgt tatggatgat ttcgcagctt ttgtagagaa
gtgctgcaag 1680gctgacgata aggagacctg ctttgccgag gagggtaaaa
aacttgttgc tgcaagtcaa 1740gctgccttag gcttataa 175826290PRTHomo
sapiensmisc_feature(1)..(290)Truncated heavy chain of the major
histocompatibility complex class I-like Fc receptor (FCGRT)
(together, SEQ ID No. 30 and SEQ ID No. 31 form
FcRN)misc_feature(1)..(290)Truncated heavy chain of the major
histocompatibility complex class I-like Fc receptor (FCGRT)
(together, SEQ ID No. 26 and SEQ ID No. 27 form FcRN) 26Met Gly Val
Pro Arg Pro Gln Pro Trp Ala Leu Gly Leu Leu Leu Phe1 5 10 15Leu Leu
Pro Gly Ser Leu Gly Ala Glu Ser His Leu Ser Leu Leu Tyr 20 25 30His
Leu Thr Ala Val Ser Ser Pro Ala Pro Gly Thr Pro Ala Phe Trp 35 40
45Val Ser Gly Trp Leu Gly Pro Gln Gln Tyr Leu Ser Tyr Asn Ser Leu
50 55 60Arg Gly Glu Ala Glu Pro Cys Gly Ala Trp Val Trp Glu Asn Gln
Val65 70 75 80Ser Trp Tyr Trp Glu Lys Glu Thr Thr Asp Leu Arg Ile
Lys Glu Lys 85 90 95Leu Phe Leu Glu Ala Phe Lys Ala Leu Gly Gly Lys
Gly Pro Tyr Thr 100 105 110Leu Gln Gly Leu Leu Gly Cys Glu Leu Gly
Pro Asp Asn Thr Ser Val 115 120 125Pro Thr Ala Lys Phe Ala Leu Asn
Gly Glu Glu Phe Met Asn Phe Asp 130 135 140Leu Lys Gln Gly Thr Trp
Gly Gly Asp Trp Pro Glu Ala Leu Ala Ile145 150 155 160Ser Gln Arg
Trp Gln Gln Gln Asp Lys Ala Ala Asn Lys Glu Leu Thr 165 170 175Phe
Leu Leu Phe Ser Cys Pro His Arg Leu Arg Glu His Leu Glu Arg 180 185
190Gly Arg Gly Asn Leu Glu Trp Lys Glu Pro Pro Ser Met Arg Leu Lys
195 200 205Ala Arg Pro Ser Ser Pro Gly Phe Ser Val Leu Thr Cys Ser
Ala Phe 210 215 220Ser Phe Tyr Pro Pro Glu Leu Gln Leu Arg Phe Leu
Arg Asn Gly Leu225 230 235 240Ala Ala Gly Thr Gly Gln Gly Asp Phe
Gly Pro Asn Ser Asp Gly Ser 245 250 255Phe His Ala Ser Ser Ser Leu
Thr Val Lys Ser Gly Asp Glu His His 260 265 270Tyr Cys Cys Ile Val
Gln His Ala Gly Leu Ala Gln Pro Leu Arg Val 275 280 285Glu Leu
29027119PRTHomo sapiensmisc_feature(1)..(119)Beta-2-microglobulin
(together, SEQ ID No. 30 and SEQ ID No. 31 form
FcRN)misc_feature(1)..(119)Beta-2-microglobulin (together, SEQ ID
No. 26 and SEQ ID No. 27 form FcRN) 27Met Ser Arg Ser Val Ala Leu
Ala Val Leu Ala Leu Leu Ser Leu Ser1 5 10 15Gly Leu Glu Ala Ile Gln
Arg Thr Pro Lys Ile Gln Val Tyr Ser Arg 20 25 30His Pro Ala Glu Asn
Gly Lys Ser Asn Phe Leu Asn Cys Tyr Val Ser 35 40 45Gly Phe His Pro
Ser Asp Ile Glu Val Asp Leu Leu Lys Asn Gly Glu 50 55 60Arg Ile Glu
Lys Val Glu His Ser Asp Leu Ser Phe Ser Lys Asp Trp65 70 75 80Ser
Phe Tyr Leu Leu Tyr Tyr Thr Glu Phe Thr Pro Thr Glu Lys Asp 85 90
95Glu Tyr Ala Cys Arg Val Asn His Val Thr Leu Ser Gln Pro Lys Ile
100 105 110Val Lys Trp Asp Arg Asp Met 115281758DNAArtificial
SequencePolynucleotide coding sequence (codon optimised for
expression in yeast) for human serum albumin 28gacgctcaca
agtccgaagt cgctcacaga ttcaaggact tgggtgaaga aaacttcaag 60gctttggtct
tgatcgcttt cgctcaatac ttgcaacaat gtccattcga agatcacgtc
120aagttggtca acgaagttac cgaattcgct aagacttgtg ttgctgacga
atctgctgaa 180aactgtgaca agtccttgca caccttgttc ggtgataagt
tgtgtactgt tgctaccttg 240agagaaacct acggtgaaat ggctgactgt
tgtgctaagc aagaaccaga aagaaacgaa 300tgtttcttgc aacacaagga
cgacaaccca aacttgccaa gattggttag accagaagtt 360gacgtcatgt
gtactgcttt ccacgacaac gaagaaacct tcttgaagaa gtacttgtac
420gaaattgcta gaagacaccc atacttctac gctccagaat tgttgttctt
cgctaagaga 480tacaaggctg ctttcaccga atgttgtcaa gctgctgata
aggctgcttg tttgttgcca 540aagttggatg aattgagaga cgaaggtaag
gcttcttccg ctaagcaaag attgaagtgt 600gcttccttgc aaaagttcgg
tgaaagagct ttcaaggctt gggctgtcgc tagattgtct 660caaagattcc
caaaggctga attcgctgaa gtttctaagt tggttactga cttgactaag
720gttcacactg aatgttgtca cggtgacttg ttggaatgtg ctgatgacag
agctgacttg 780gctaagtaca tctgtgaaaa ccaagactct atctcttcca
agttgaagga atgttgtgaa 840aagccattgt tggaaaagtc tcactgtatt
gctgaagttg aaaacgatga aatgccagct 900gacttgccat ctttggctgc
tgacttcgtt gaatctaagg acgtttgtaa gaactacgct 960gaagctaagg
acgtcttctt gggtatgttc ttgtacgaat acgctagaag acacccagac
1020tactccgttg tcttgttgtt gagattggct aagacctacg aaactacctt
ggaaaagtgt 1080tgtgctgctg ctgacccaca cgaatgttac gctaaggttt
tcgatgaatt caagccattg 1140gtcgaagaac cacaaaactt gatcaagcaa
aactgtgaat tgttcgaaca attgggtgaa 1200tacaagttcc aaaacgcttt
gttggttaga tacactaaga aggtcccaca agtctccacc 1260ccaactttgg
ttgaagtctc tagaaacttg ggtaaggtcg gttctaagtg ttgtaagcac
1320ccagaagcta agagaatgcc atgtgctgaa gattacttgt ccgtcgtttt
gaaccaattg 1380tgtgttttgc acgaaaagac cccagtctct gatagagtca
ccaagtgttg tactgaatct 1440ttggttaaca gaagaccatg tttctctgct
ttggaagtcg acgaaactta cgttccaaag 1500gaattcaacg ctgaaacttt
caccttccac gctgatatct gtaccttgtc cgaaaaggaa 1560agacaaatta
agaagcaaac tgctttggtt gaattggtca agcacaagcc aaaggctact
1620aaggaacaat tgaaggctgt catggatgat ttcgctgctt tcgttgaaaa
gtgttgtaag 1680gctgatgata aggaaacttg tttcgctgaa gaaggtaaga
agttggtcgc tgcttcccaa 1740gctgctttgg gtttgtaa
175829609PRTArtificial SequenceHSA with fusion leader
sequencesignal(1)..(24)mat_peptide(25)..(609) 29Met Lys Trp Val Ser
Phe Ile Ser Leu Leu Phe Leu Phe Ser Ser Ala -20 -15 -10Tyr Ser Arg
Ser Leu Asp Lys Arg Asp Ala His Lys Ser Glu Val Ala -5 -1 1 5His
Arg Phe Lys Asp Leu Gly Glu Glu Asn Phe Lys Ala Leu Val Leu 10 15
20Ile Ala Phe Ala Gln Tyr Leu Gln Gln Cys Pro Phe Glu Asp His Val25
30 35 40Lys Leu Val Asn Glu Val Thr Glu Phe Ala Lys Thr Cys Val Ala
Asp 45 50 55Glu Ser Ala Glu Asn Cys Asp Lys Ser Leu His Thr Leu Phe
Gly Asp 60 65 70Lys Leu Cys Thr Val Ala Thr Leu Arg Glu Thr Tyr Gly
Glu Met Ala 75 80 85Asp Cys Cys Ala Lys Gln Glu Pro Glu Arg Asn Glu
Cys Phe Leu Gln 90 95 100His Lys Asp Asp Asn Pro Asn Leu Pro Arg
Leu Val Arg Pro Glu Val105 110 115 120Asp Val Met Cys Thr Ala Phe
His Asp Asn Glu Glu Thr Phe Leu Lys 125 130 135Lys Tyr Leu Tyr Glu
Ile Ala Arg Arg His Pro Tyr Phe Tyr Ala Pro 140 145 150Glu Leu Leu
Phe Phe Ala Lys Arg Tyr Lys Ala Ala Phe Thr Glu Cys 155 160 165Cys
Gln Ala Ala Asp Lys Ala Ala Cys Leu Leu Pro Lys Leu Asp Glu 170 175
180Leu Arg Asp Glu Gly Lys Ala Ser Ser Ala Lys Gln Arg Leu Lys
Cys185 190 195 200Ala Ser Leu Gln Lys Phe Gly Glu Arg Ala Phe Lys
Ala Trp Ala Val 205 210 215Ala Arg Leu Ser Gln Arg Phe Pro Lys Ala
Glu Phe Ala Glu Val Ser 220 225 230Lys Leu Val Thr Asp Leu Thr Lys
Val His Thr Glu Cys Cys His Gly 235 240 245Asp Leu Leu Glu Cys Ala
Asp Asp Arg Ala Asp Leu Ala Lys Tyr Ile 250 255 260Cys Glu Asn Gln
Asp Ser Ile Ser Ser Lys Leu Lys Glu Cys Cys Glu265 270 275 280Lys
Pro Leu Leu Glu Lys Ser His Cys Ile Ala Glu Val Glu Asn Asp 285 290
295Glu Met Pro Ala Asp Leu Pro Ser Leu Ala Ala Asp Phe Val Glu
Ser
300 305 310Lys Asp Val Cys Lys Asn Tyr Ala Glu Ala Lys Asp Val Phe
Leu Gly 315 320 325Met Phe Leu Tyr Glu Tyr Ala Arg Arg His Pro Asp
Tyr Ser Val Val 330 335 340Leu Leu Leu Arg Leu Ala Lys Thr Tyr Glu
Thr Thr Leu Glu Lys Cys345 350 355 360Cys Ala Ala Ala Asp Pro His
Glu Cys Tyr Ala Lys Val Phe Asp Glu 365 370 375Phe Lys Pro Leu Val
Glu Glu Pro Gln Asn Leu Ile Lys Gln Asn Cys 380 385 390Glu Leu Phe
Glu Gln Leu Gly Glu Tyr Lys Phe Gln Asn Ala Leu Leu 395 400 405Val
Arg Tyr Thr Lys Lys Val Pro Gln Val Ser Thr Pro Thr Leu Val 410 415
420Glu Val Ser Arg Asn Leu Gly Lys Val Gly Ser Lys Cys Cys Lys
His425 430 435 440Pro Glu Ala Lys Arg Met Pro Cys Ala Glu Asp Tyr
Leu Ser Val Val 445 450 455Leu Asn Gln Leu Cys Val Leu His Glu Lys
Thr Pro Val Ser Asp Arg 460 465 470Val Thr Lys Cys Cys Thr Glu Ser
Leu Val Asn Arg Arg Pro Cys Phe 475 480 485Ser Ala Leu Glu Val Asp
Glu Thr Tyr Val Pro Lys Glu Phe Asn Ala 490 495 500Glu Thr Phe Thr
Phe His Ala Asp Ile Cys Thr Leu Ser Glu Lys Glu505 510 515 520Arg
Gln Ile Lys Lys Gln Thr Ala Leu Val Glu Leu Val Lys His Lys 525 530
535Pro Lys Ala Thr Lys Glu Gln Leu Lys Ala Val Met Asp Asp Phe Ala
540 545 550Ala Phe Val Glu Lys Cys Cys Lys Ala Asp Asp Lys Glu Thr
Cys Phe 555 560 565Ala Glu Glu Gly Lys Lys Leu Val Ala Ala Ser Gln
Ala Ala Leu Gly 570 575 580Leu58530585PRTArtificialHSA C34A 30Asp
Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10
15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln
20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr
Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys
Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val
Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys
Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys
Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val
Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe
Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr
Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr
Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170
175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser
180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe
Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser
Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu
Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His
Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala
Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu
Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys
Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295
300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr
Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr
Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu
Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys
Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe
Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile
Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr
Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410
415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys
420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met
Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu
Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr
Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys
Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu
Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr
Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu
Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535
540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys
Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly
Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
5853136DNAArtificial SequenceC34A reverse primer 31ttgttgcaag
tattgagcga aagcgatcaa gaccaa 363245DNAArtificial SequenceC34A
forward primer 32ttcgctcaat acttgcaaca agctccattc gaagatcacg tcaag
453350DNAArtificial SequenceL24C forward primer 33gaagaaaact
tcaaggcttt ggtctgtatc gctttcgctc aatacttgca 503448DNAArtificial
SequenceF49C forward primer 34agttggtcaa cgaagttacc gaatgtgcta
agacttgtgt tgctgacg 483548DNAArtificial SequenceV54C forward primer
35gttaccgaat tcgctaagac ttgttgtgct gacgaatccg cggaaaac
483649DNAArtificial SequenceD56C forward primer 36gaattcgcta
agacttgtgt tgcttgtgaa tccgcggaaa actgtgaca 493748DNAArtificial
SequenceL66C forward primer 37cgcggaaaac tgtgacaagt cctgtcacac
cttgttcggt gataagtt 483849DNAArtificial SequenceA92C forward primer
38cggtgaaatg gctgactgtt gttgtaagca agaaccagaa agaaacgaa
493950DNAArtificial SequenceK93C forward primer 39gtgaaatggc
tgactgttgt gcttgtcaag aaccagaaag aaacgaatgt 504050DNAArtificial
SequenceQ94C forward primer 40aaatggctga ctgttgtgct aagtgtgaac
cagaaagaaa cgaatgtttc 504150DNAArtificial SequenceE97C forward
primer 41actgttgtgc taagcaagaa ccatgtagaa acgaatgttt cttgcaacac
504250DNAArtificial SequenceH128C forward primer 42ttgacgtcat
gtgtactgct ttctgtgaca acgaagaaac cttcttgaag 504349DNAArtificial
SequenceF156C forward primer 43acttctacgc tccagaattg ttgtgtttcg
ctaagagata caaggctgc 494451DNAArtificial SequenceA226C forward
primer 44agattgtctc aaagattccc aaagtgtgaa ttcgctgaag tttctaagtt g
514550DNAArtificial SequenceA227C forward primer 45tgtctcaaag
attcccaaag gcttgtttcg ctgaagtttc taagttggtt 504650DNAArtificial
SequenceE230C forward primer 46gattcccaaa ggctgaattc gcttgtgttt
ctaagttggt tactgacttg 504751DNAArtificial SequenceD237C forward
primer 47gctgaagttt ctaagttggt tacttgtttg actaaggttc acactgaatg t
514850DNAArtificial SequenceK240C forward primer 48tctaagttgg
ttactgactt gacttgtgtt cacactgaat gttgtcacgg 504949DNAArtificial
SequenceD259C forward primer 49ggaatgtgct gatgacagag cttgtttggc
taagtacatc tgtgaaaac 495049DNAArtificial SequenceK262C forward
primer 50tgatgacaga gctgacttgg cttgttacat ctgtgaaaac caagactct
495151DNAArtificial SequenceN267C forward primer 51gacttggcta
agtacatctg tgaatgtcaa gactctatct cttccaagtt g 515251DNAArtificial
SequenceQ268C forward primer 52ttggctaagt acatctgtga aaactgtgac
tctatctctt ccaagttgaa g 515351DNAArtificial SequenceI271C forward
primer 53tacatctgtg aaaaccaaga ctcttgttct tccaagttga aggaatgttg t
515450DNAArtificial SequenceL275C forward primer 54accaagactc
tatctcttcc aagtgtaagg aatgttgtga aaagccattg 505551DNAArtificial
SequenceE277C forward primer 55gactctatct cttccaagtt gaagtgttgt
tgtgaaaagc cattgttgga a 515651DNAArtificial SequenceL284C forward
primer 56aaggaatgtt gtgaaaagcc attgtgtgaa aagtctcact gtattgctga a
515750DNAArtificial SequenceE294C forward primer 57aagtctcact
gtattgctga agtttgtaac gatgaaatgc cagctgactt 505850DNAArtificial
SequenceE311C forward primer 58catctttggc tgctgacttc gtttgttcta
aggacgtttg taagaactac 505949DNAArtificial SequenceK317C forward
primer 59ttcgttgaat ctaaggacgt ttgttgtaac tacgctgaag ctaaggacg
496050DNAArtificial SequenceA322C forward primer 60gacgtttgta
agaactacgc tgaatgtaag gacgtcttct tgggtatgtt 506149DNAArtificial
SequenceE333C 61gtcttcttgg gtatgttctt gtactgttac gctagaagac
acccagact 496248DNAArtificial SequenceD340C forward primer
62cgaatacgct agaagacacc catgttactc cgttgtcttg ttgttgag
486348DNAArtificial SequenceE354C forward primer 63tgttgagatt
ggctaagacc tactgtacta ccctcgagaa gtgttgtg 486448DNAArtificial
SequenceE358C forward primer 64ctaagaccta cgaaactacc ctctgtaagt
gttgtgctgc tgctgacc 486546DNAArtificial SequenceK359C forward
primer 65gacctacgaa actaccctcg agtgttgttg tgctgctgct gaccca
466647DNAArtificial SequenceA362C forward primer 66aaactaccct
cgagaagtgt tgttgtgctg ctgacccaca cgaatgt 476750DNAArtificial
SequenceE382C forward primer 67tcgatgaatt caagccattg gtctgtgaac
cacaaaactt gatcaagcaa 506851DNAArtificial SequenceL398C forward
primer 68gcaaaactgt gaattgttcg aacaatgtgg tgaatacaag ttccaaaacg c
516935DNAArtificial SequenceL24C reverse primer 69gaccaaagcc
ttgaagtttt cttcacccaa gtcct 357035DNAArtificial SequenceF49C
reverse primer 70ttcggtaact tcgttgacca acttgacgtg atctt
357136DNAArtificial SequenceV54C reverse primer 71acaagtctta
gcgaattcgg taacttcgtt gaccaa 367236DNAArtificial SequenceD56C
reverse primer 72agcaacacaa gtcttagcga attcggtaac ttcgtt
367333DNAArtificial SequenceL66C reverse primer 73ggacttgtca
cagttttccg cggattcgtc agc 337434DNAArtificial SequenceA92C reverse
primer 74acaacagtca gccatttcac cgtaggtttc tctc 347535DNAArtificial
SequenceK93C reverse primer 75agcacaacag tcagccattt caccgtaggt
ttctc 357634DNAArtificial SequenceQ94C reverse primer 76cttagcacaa
cagtcagcca tttcaccgta ggtt 347735DNAArtificial SequenceE97C reverse
primer 77tggttcttgc ttagcacaac agtcagccat ttcac 357835DNAArtificial
SequenceH128C reverse primer 78gaaagcagta cacatgacgt caacttctgg
tctaa 357935DNAArtificial SequenceF156C reverse primer 79caacaattct
ggagcgtaga agtatgggtg tcttc 358034DNAArtificial SequenceA226C
reverse primer 80ctttgggaat ctttgagaca atctagcgac agcc
348135DNAArtificial SequenceE227C reverse primer 81agcctttggg
aatctttgag acaatctagc gacag 358236DNAArtificial SequenceE230C
reverse primer 82agcgaattca gcctttggga atctttgaga caatct
368336DNAArtificial SequenceE237C reverse primer 83agtaaccaac
ttagaaactt cagcgaattc agcctt 368436DNAArtificial SequenceK240C
reverse primer 84agtcaagtca gtaaccaact tagaaacttc agcgaa
368534DNAArtificial SequenceD259C reverse primer 85agctctgtca
tcagcacatt ccaacaagtc accg 348634DNAArtificial SequenceK262C
reverse primer 86agccaagtca gctctgtcat cagcacattc caac
348736DNAArtificial SequenceN267C reverse primer 87ttcacagatg
tacttagcca agtcagctct gtcatc 368835DNAArtificial SequenceQ268C
reverse primer 88gttttcacag atgtacttag ccaagtcagc tctgt
358936DNAArtificial SequenceI271C reverse primer 89agagtcttgg
ttttcacaga tgtacttagc caagtc 369036DNAArtificial SequenceL275C
reverse primer 90cttggaagag atagagtctt ggttttcaca gatgta
369137DNAArtificial SequenceE277C reverse primer 91cttcaacttg
gaagagatag agtcttggtt ttcacag 379236DNAArtificial SequenceE284C
reverse primer 92caatggcttt tcacaacatt ccttcaactt ggaaga
369336DNAArtificial SequenceE294C reverse primer 93aacttcagca
atacagtgag acttttccaa caatgg 369434DNAArtificial SequenceE311C
reverse primer 94aacgaagtca gcagccaaag atggcaagtc agct
349534DNAArtificial SequenceK317C reverse primer 95acaaacgtcc
ttagattcaa cgaagtcagc agcc 349637DNAArtificial SequenceA322C
reverse primer 96ttcagcgtag ttcttacaaa cgtccttaga ttcaacg
379736DNAArtificial SequenceE333C reverse primer 97gtacaagaac
atacccaaga agacgtcctt agcttc 369835DNAArtificial SequenceD340C
reverse primer 98tgggtgtctt ctagcgtatt cgtacaagaa catac
359935DNAArtificial SequenceE354C reverse primer 99gtaggtctta
gccaatctca acaacaagac aacgg 3510035DNAArtificial SequenceE358C
reverse primer 100gagggtagtt tcgtaggtct tagccaatct caaca
3510134DNAArtificial SequenceK359C reverse primer 101ctcgagggta
gtttcgtagg tcttagccaa tctc 3410235DNAArtificial SequenceA362C
reverse primer 102acaacacttc tcgagggtag tttcgtaggt cttag
3510335DNAArtificial SequenceE382C reverse primer 103gaccaatggc
ttgaattcat cgaaaacctt agcgt 3510439DNAArtificial SequenceL398C
reverse primer 104ttgttcgaac aattcacagt tttgcttgat caagttttg
39105585PRTArtificial SequenceHSA C34A_L24C 105Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Cys Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585106585PRTArtificial SequenceHSA C34A_F49C 106Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Cys Ala
Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585107585PRTArtificial SequenceHSA C34A_V54C 107Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Cys Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585108585PRTArtificial SequenceHSA C34A_D56C 108Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Val Ala Cys Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585109585PRTArtificial SequenceHSA C34A_L66C 109Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Cys His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu
Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val
Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser
Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn
Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr
Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505
510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala
515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu
Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu
Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala
Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu
Gly Leu 580 585110585PRTArtificial SequenceHSA C34A_A92C 110Asp Ala
His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu
Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25
30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu
35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp
Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala
Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Cys
Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp
Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp
Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu
Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe
Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys
Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170
175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser
180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe
Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser
Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu
Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His
Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala
Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu
Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys
Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295
300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr
Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr
Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu
Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys
Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe
Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile
Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr
Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410
415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys
420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met
Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu
Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr
Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys
Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu
Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr
Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu
Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535
540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys
Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly
Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585111585PRTArtificial SequenceHSA C34A_K93C 111Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Cys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585112585PRTArtificial SequenceHSA C34A_Q94C 112Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Cys Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585113585PRTArtificial SequenceHSA C34A_E97C 113Asp Ala His Lys Ser
Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys
Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro
Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala
Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser
Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Cys Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp
Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp
Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570
575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580 585114585PRTArtificial
SequenceHSA C34A_H128C 114Asp Ala His Lys Ser Glu Val Ala His Arg
Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile
Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val
Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala
Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe
Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr
Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn
Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro
Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe Cys 115 120
125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg
130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala
Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala
Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu
Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys
Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala
Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu
Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235
240Val His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp
245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser
Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu
Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met
Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser
Lys Asp Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val
Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro
Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr
Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360
365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro
370 375 380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu
Gly Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr
Thr Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu
Val Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys
His Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu
Ser Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr
Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475
480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr
485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His
Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys
Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys
Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala
Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu
Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser
Gln Ala Ala Leu Gly Leu 580 585115585PRTArtificial SequenceHSA
C34A_F156C 115Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp
Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala
Gln Tyr Leu Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val
Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser
Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys
Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met
Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe
Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val
Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn
Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135
140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Cys Phe Ala Lys
Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala
Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg
Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys
Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp
Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe
Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val
His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250
255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser
260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys
Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala
Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp
Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu
Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr
Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr
Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys
Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375
380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly
Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr
Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val
Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His
Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser
Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro
Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu
Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490
495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp
500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln
Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe
Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala
Ala Leu Gly Leu 580 585116585PRTArtificial SequenceHSA C34A_A226C
116Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1
5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu
Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val
Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn
Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr
Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys
Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His
Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu
Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr
Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro
Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155
160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala
165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys
Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln
Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg
Leu Ser Gln Arg Phe Pro 210 215 220Lys Cys Glu Phe Ala Glu Val Ser
Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys
Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp
Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser
Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280
285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser
290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn
Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu
Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu
Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys
Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val
Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu
Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395
400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro
405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu
Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys
Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn
Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg
Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520
525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu
530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys
Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu
Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu
580 585117585PRTArtificial SequenceHSA C34A_E227C 117Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Cys Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585118585PRTArtificial SequenceHSA C34A_E230C 118Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50
55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Cys Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585119585PRTArtificial SequenceHSA C34A_D237C 119Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Cys Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585120585PRTArtificial SequenceHSA C34A_K240C 120Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Cys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585121585PRTArtificial SequenceHSA C34A_D259C 121Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Cys Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585122585PRTArtificial SequenceHSA C34A_K262C 122Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala
165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys
Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln
Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg
Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser
Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys
Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp
Leu Ala Cys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser
Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280
285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser
290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn
Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu
Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu
Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys
Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val
Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu
Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395
400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro
405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu
Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys
Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn
Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg
Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520
525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu
530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys
Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu
Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu
580 585123585PRTArtificial SequenceHSA C34A_N267C 123Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Cys Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585124585PRTArtificial SequenceHSA C34A_Q268C 124Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Cys Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585125585PRTArtificial SequenceHSA C34A_I271C 125Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Cys Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585126585PRTArtificial SequenceHSA C34A_L275C 126Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Cys Lys Glu Cys Cys
Glu Lys Pro Leu Leu
Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met
Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser
Lys Asp Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val
Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro
Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr
Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360
365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro
370 375 380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu
Gly Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr
Thr Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu
Val Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys
His Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu
Ser Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr
Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475
480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr
485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His
Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys
Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys
Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala
Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu
Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser
Gln Ala Ala Leu Gly Leu 580 585127585PRTArtificial SequenceHSA
C34A_E277C 127Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp
Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala
Gln Tyr Leu Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val
Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser
Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys
Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met
Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe
Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val
Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn
Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135
140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys
Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala
Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg
Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys
Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp
Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe
Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val
His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250
255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser
260 265 270Ser Lys Leu Lys Cys Cys Cys Glu Lys Pro Leu Leu Glu Lys
Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala
Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp
Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu
Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr
Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr
Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys
Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375
380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly
Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr
Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val
Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His
Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser
Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro
Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu
Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490
495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp
500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln
Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe
Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala
Ala Leu Gly Leu 580 585128585PRTArtificial SequenceHSA C34A_L284C
128Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1
5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu
Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val
Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn
Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr
Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys
Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His
Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu
Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr
Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro
Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155
160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala
165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys
Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln
Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg
Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser
Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys
Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp
Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser
Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Cys Glu Lys Ser His 275 280
285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser
290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn
Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu
Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu
Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys
Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val
Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu
Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395
400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro
405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu
Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys
Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn
Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg
Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520
525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu
530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys
Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu
Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu
580 585129585PRTArtificial SequenceHSA C34A_E294C 129Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Cys Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585130585PRTArtificial SequenceHSA C34A_E311C 130Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Cys Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu
Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu
Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro
Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser
Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala
Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His 450 455
460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu
Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu
Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr
Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys Glu
Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val Lys
His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val Met
Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala
Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570
575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580 585131585PRTArtificial
SequenceHSA C34A_K317C 131Asp Ala His Lys Ser Glu Val Ala His Arg
Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile
Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val
Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala
Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe
Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr
Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn
Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro
Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120
125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg
130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala
Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala
Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu
Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys
Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala
Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu
Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235
240Val His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp
245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser
Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu
Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met
Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser
Lys Asp Val Cys Cys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val
Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro
Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr
Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360
365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro
370 375 380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu
Gly Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr
Thr Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu
Val Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys
His Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu
Ser Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr
Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475
480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr
485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His
Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys
Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys
Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala
Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu
Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser
Gln Ala Ala Leu Gly Leu 580 585132585PRTArtificial SequenceHSA
C34A_A322C 132Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp
Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala
Gln Tyr Leu Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val
Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser
Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys
Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met
Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe
Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val
Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn
Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135
140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys
Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala
Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg
Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys
Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp
Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe
Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val
His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250
255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser
260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys
Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala
Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp
Val Cys Lys Asn Tyr Ala305 310 315 320Glu Cys Lys Asp Val Phe Leu
Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr
Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr
Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys
Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375
380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly
Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr
Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val
Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His
Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser
Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro
Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu
Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490
495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp
500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln
Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe
Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala
Ala Leu Gly Leu 580 585133585PRTArtificial SequenceHSA C34A_E333C
133Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1
5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu
Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val
Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn
Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr
Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys
Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His
Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu
Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr
Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro
Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155
160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala
165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys
Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln
Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg
Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser
Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys
Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp
Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser
Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280
285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser
290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn
Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu
Tyr Cys Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu
Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys
Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val
Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu
Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395
400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro
405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu
Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys
Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn
Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg
Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520
525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu
530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys
Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu
Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu
580 585134585PRTArtificial SequenceHSA C34A_D340C 134Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Cys Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490
495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp
500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln
Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe
Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala
Ala Leu Gly Leu 580 585135585PRTArtificial SequenceHSA C34A_E354C
135Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1
5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu
Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val
Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn
Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr
Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys
Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His
Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu
Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr
Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro
Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155
160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala
165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys
Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln
Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg
Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser
Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys
Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp
Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser
Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280
285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser
290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn
Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu
Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu
Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Cys Thr Thr Leu Glu Lys
Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val
Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu
Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395
400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro
405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu
Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys
Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn
Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg
Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520
525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu
530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys
Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu
Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu
580 585136585PRTArtificial SequenceHSA C34A_E358C 136Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Cys Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585137585PRTArtificial SequenceHSA C34A_K359C 137Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Cys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585138585PRTArtificial SequenceHSA C34A_A362C 138Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Ala
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Cys Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585139585PRTArtificial SequenceHSA C34A_E382C 139Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5
10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu
Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val
Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn
Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr
Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys
Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His
Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu
Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr
Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro
Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155
160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala
165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys
Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln
Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg
Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser
Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys
Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp
Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser
Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280
285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser
290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn
Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu
Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu
Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys
Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val
Phe Asp Glu Phe Lys Pro Leu Val Cys Glu Pro 370 375 380Gln Asn Leu
Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395
400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro
405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu
Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys
Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn
Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg
Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg
Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro
Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile
Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520
525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu
530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys
Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu
Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu
580 585140585PRTArtificial SequenceHSA C34A_L398C 140Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Cys Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585141585PRTArtificial SequenceHSA C34A_K93C_E294C 141Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Cys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Cys Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585142585PRTArtificial SequenceHSA K93C 142Asp Ala His Lys Ser Glu
Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala
Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Cys Pro Phe
Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala Lys
Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu
His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Cys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu
Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu
Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr
Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro
Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala
Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585143585PRTArtificial SequenceHSA E294C 143Asp Ala His Lys Ser Glu
Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala
Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Cys Pro Phe
Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe Ala Lys
Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu
His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65 70 75
80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu Pro
85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro Asn
Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys Thr
Ala Phe His 115
120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala
Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe
Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln
Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu
Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu
Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys
Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala
Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235
240Val His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp
245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser
Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu
Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val Cys Asn Asp Glu Met
Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser
Lys Asp Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val
Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro
Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr
Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360
365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro
370 375 380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu
Gly Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr
Thr Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu
Val Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys
His Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu
Ser Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr
Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475
480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr
485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His
Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys
Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys
Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala
Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu
Thr Cys Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser
Gln Ala Ala Leu Gly Leu 580 585144585PRTArtificial SequenceHSA
K93C_E294C 144Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp
Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala
Gln Tyr Leu Gln 20 25 30Gln Cys Pro Phe Glu Asp His Val Lys Leu Val
Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser
Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys
Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met
Ala Asp Cys Cys Ala Cys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe
Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val
Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn
Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135
140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys
Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala
Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg
Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys
Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp
Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe
Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val
His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250
255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser
260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys
Ser His 275 280 285Cys Ile Ala Glu Val Cys Asn Asp Glu Met Pro Ala
Asp Leu Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp
Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu
Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr
Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr
Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys
Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375
380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly
Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr
Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val
Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His
Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser
Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro
Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu
Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490
495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp
500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln
Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe
Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala
Ala Leu Gly Leu 580 585145585PRTArtificial SequenceHSA K573P 145Asp
Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10
15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln
20 25 30Gln Cys Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr
Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys
Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val
Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys
Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys
Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val
Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe
Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr
Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr
Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170
175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser
180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe
Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser
Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu
Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His
Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala
Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu
Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys
Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295
300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr
Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr
Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu
Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys
Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe
Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile
Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr
Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410
415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys
420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met
Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu
Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr
Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys
Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu
Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr
Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu
Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535
540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys
Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly
Pro Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585146585PRTArtificial SequenceHSA C34A_K93C_K573P 146Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Cys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Pro Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585147585PRTArtificial SequenceHSA C34A_E294C_K573P 147Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val
Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Cys Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu
Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala
Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe
Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn
Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln
Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val
Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425
430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys
435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val
Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys
Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser
Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn
Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser
Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu
Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys
Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550
555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Pro Lys Leu
Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585148585PRTArtificial SequenceHSA C34A_K93C_E294C_K573P 148Asp Ala
His Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu
Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25
30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu
35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp
Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala
Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala
Cys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp
Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp
Val Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu
Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe
Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys
Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170
175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser
180 185 190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe
Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser
Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu
Val Thr Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His
Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala
Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu
Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys
Ile Ala Glu Val Cys Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295
300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr
Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr
Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu
Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys
Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe
Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile
Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr
Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410
415Gln Val Ser Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys
420 425 430Val Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met
Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu
Cys Val Leu His 450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr
Lys Cys Cys Thr Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys
Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu
Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr
Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu
Val Glu Leu Val Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535
540Lys Ala Val Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys
Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly
Pro Lys Leu Val 565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585149585PRTArtificial SequenceHSA K93C_K573P 149Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Cys
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Cys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Glu Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Pro Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585150585PRTArtificial SequenceHSA E294C_K573P 150Asp Ala His Lys
Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn Phe
Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln Cys
Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40 45Phe
Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys 50 55
60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr Leu65
70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Lys Gln Glu
Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp Asn Pro
Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val Met Cys
Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys Lys Tyr
Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr Ala Pro
Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala Ala Phe
Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys Leu Leu
Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185 190Ser
Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu 195 200
205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg Phe Pro
210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr Asp Leu
Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp Leu Leu
Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr Ile Cys
Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu Cys Cys
Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala Glu Val
Cys Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu Ala Ala
Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310 315
320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala Arg
325 330 335Arg His Pro Asp Tyr Ser Val Val Leu Leu Leu Arg Leu Ala
Lys Thr 340 345 350Tyr Glu Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala
Asp Pro His Glu 355 360 365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys
Pro Leu Val Glu Glu Pro 370 375 380Gln Asn Leu Ile Lys Gln Asn Cys
Glu Leu Phe Glu Gln Leu Gly Glu385 390 395 400Tyr Lys Phe Gln Asn
Ala Leu Leu Val Arg Tyr Thr Lys Lys Val Pro 405 410 415Gln Val Ser
Thr Pro Thr Leu Val Glu Val Ser Arg Asn Leu Gly Lys 420 425 430Val
Gly Ser Lys Cys Cys Lys His Pro Glu Ala Lys Arg Met Pro Cys 435 440
445Ala Glu Asp Tyr Leu Ser Val Val Leu Asn Gln Leu Cys Val Leu His
450 455 460Glu Lys Thr Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr
Glu Ser465 470 475 480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu
Glu Val Asp Glu Thr 485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu
Thr Phe Thr Phe His Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys
Glu Arg Gln Ile Lys Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val
Lys His Lys Pro Lys Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val
Met Asp Asp Phe Ala Ala Phe Val Glu Lys Cys Cys Lys545 550 555
560Ala Asp Asp Lys Glu Thr Cys Phe Ala Glu Glu Gly Pro Lys Leu Val
565 570 575Ala Ala Ser Gln Ala Ala Leu Gly Leu 580
585151585PRTArtificial SequenceHSA K93C_E294C_K573P 151Asp Ala His
Lys Ser Glu Val Ala His Arg Phe Lys Asp Leu Gly Glu1 5 10 15Glu Asn
Phe Lys Ala Leu Val Leu Ile Ala Phe Ala Gln Tyr Leu Gln 20 25 30Gln
Cys Pro Phe Glu Asp His Val Lys Leu Val Asn Glu Val Thr Glu 35 40
45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser Ala Glu Asn Cys Asp Lys
50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys Leu Cys Thr Val Ala Thr
Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met Ala Asp Cys Cys Ala Cys
Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe Leu Gln His Lys Asp Asp
Asn Pro Asn Leu 100 105 110Pro Arg Leu Val Arg Pro Glu Val Asp Val
Met Cys Thr Ala Phe His 115 120 125Asp Asn Glu Glu Thr Phe Leu Lys
Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135 140Arg His Pro Tyr Phe Tyr
Ala Pro Glu Leu Leu Phe Phe Ala Lys Arg145 150 155 160Tyr Lys Ala
Ala Phe Thr Glu Cys Cys Gln Ala Ala Asp Lys Ala Ala 165 170 175Cys
Leu Leu Pro Lys Leu Asp Glu Leu Arg Asp Glu Gly Lys Ala Ser 180 185
190Ser Ala Lys Gln Arg Leu Lys Cys Ala Ser Leu Gln Lys Phe Gly Glu
195 200 205Arg Ala Phe Lys Ala Trp Ala Val Ala Arg Leu Ser Gln Arg
Phe Pro 210 215 220Lys Ala Glu Phe Ala Glu Val Ser Lys Leu Val Thr
Asp Leu Thr Lys225 230 235 240Val His Thr Glu Cys Cys His Gly Asp
Leu Leu Glu Cys Ala Asp Asp 245 250 255Arg Ala Asp Leu Ala Lys Tyr
Ile Cys Glu Asn Gln Asp Ser Ile Ser 260 265 270Ser Lys Leu Lys Glu
Cys Cys Glu Lys Pro Leu Leu Glu Lys Ser His 275 280 285Cys Ile Ala
Glu Val Cys Asn Asp Glu Met Pro Ala Asp Leu Pro Ser 290 295 300Leu
Ala Ala Asp Phe Val Glu Ser Lys Asp Val Cys Lys Asn Tyr Ala305 310
315 320Glu Ala Lys Asp Val Phe Leu Gly Met Phe Leu Tyr Glu Tyr Ala
Arg 325 330 335Arg His Pro Asp
Tyr Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu
Thr Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360
365Cys Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro
370 375 380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu
Gly Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr
Thr Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu
Val Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys
His Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu
Ser Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr
Pro Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475
480Leu Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr
485 490 495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His
Ala Asp 500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys
Lys Gln Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys
Ala Thr Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala
Ala Phe Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu
Thr Cys Phe Ala Glu Glu Gly Pro Lys Leu Val 565 570 575Ala Ala Ser
Gln Ala Ala Leu Gly Leu 580 585152585PRTArtificial SequenceHSA
C34A_L302C 152Asp Ala His Lys Ser Glu Val Ala His Arg Phe Lys Asp
Leu Gly Glu1 5 10 15Glu Asn Phe Lys Ala Leu Val Leu Ile Ala Phe Ala
Gln Tyr Leu Gln 20 25 30Gln Ala Pro Phe Glu Asp His Val Lys Leu Val
Asn Glu Val Thr Glu 35 40 45Phe Ala Lys Thr Cys Val Ala Asp Glu Ser
Ala Glu Asn Cys Asp Lys 50 55 60Ser Leu His Thr Leu Phe Gly Asp Lys
Leu Cys Thr Val Ala Thr Leu65 70 75 80Arg Glu Thr Tyr Gly Glu Met
Ala Asp Cys Cys Ala Lys Gln Glu Pro 85 90 95Glu Arg Asn Glu Cys Phe
Leu Gln His Lys Asp Asp Asn Pro Asn Leu 100 105 110Pro Arg Leu Val
Arg Pro Glu Val Asp Val Met Cys Thr Ala Phe His 115 120 125Asp Asn
Glu Glu Thr Phe Leu Lys Lys Tyr Leu Tyr Glu Ile Ala Arg 130 135
140Arg His Pro Tyr Phe Tyr Ala Pro Glu Leu Leu Phe Phe Ala Lys
Arg145 150 155 160Tyr Lys Ala Ala Phe Thr Glu Cys Cys Gln Ala Ala
Asp Lys Ala Ala 165 170 175Cys Leu Leu Pro Lys Leu Asp Glu Leu Arg
Asp Glu Gly Lys Ala Ser 180 185 190Ser Ala Lys Gln Arg Leu Lys Cys
Ala Ser Leu Gln Lys Phe Gly Glu 195 200 205Arg Ala Phe Lys Ala Trp
Ala Val Ala Arg Leu Ser Gln Arg Phe Pro 210 215 220Lys Ala Glu Phe
Ala Glu Val Ser Lys Leu Val Thr Asp Leu Thr Lys225 230 235 240Val
His Thr Glu Cys Cys His Gly Asp Leu Leu Glu Cys Ala Asp Asp 245 250
255Arg Ala Asp Leu Ala Lys Tyr Ile Cys Glu Asn Gln Asp Ser Ile Ser
260 265 270Ser Lys Leu Lys Glu Cys Cys Glu Lys Pro Leu Leu Glu Lys
Ser His 275 280 285Cys Ile Ala Glu Val Glu Asn Asp Glu Met Pro Ala
Asp Cys Pro Ser 290 295 300Leu Ala Ala Asp Phe Val Glu Ser Lys Asp
Val Cys Lys Asn Tyr Ala305 310 315 320Glu Ala Lys Asp Val Phe Leu
Gly Met Phe Leu Tyr Glu Tyr Ala Arg 325 330 335Arg His Pro Asp Tyr
Ser Val Val Leu Leu Leu Arg Leu Ala Lys Thr 340 345 350Tyr Glu Thr
Thr Leu Glu Lys Cys Cys Ala Ala Ala Asp Pro His Glu 355 360 365Cys
Tyr Ala Lys Val Phe Asp Glu Phe Lys Pro Leu Val Glu Glu Pro 370 375
380Gln Asn Leu Ile Lys Gln Asn Cys Glu Leu Phe Glu Gln Leu Gly
Glu385 390 395 400Tyr Lys Phe Gln Asn Ala Leu Leu Val Arg Tyr Thr
Lys Lys Val Pro 405 410 415Gln Val Ser Thr Pro Thr Leu Val Glu Val
Ser Arg Asn Leu Gly Lys 420 425 430Val Gly Ser Lys Cys Cys Lys His
Pro Glu Ala Lys Arg Met Pro Cys 435 440 445Ala Glu Asp Tyr Leu Ser
Val Val Leu Asn Gln Leu Cys Val Leu His 450 455 460Glu Lys Thr Pro
Val Ser Asp Arg Val Thr Lys Cys Cys Thr Glu Ser465 470 475 480Leu
Val Asn Arg Arg Pro Cys Phe Ser Ala Leu Glu Val Asp Glu Thr 485 490
495Tyr Val Pro Lys Glu Phe Asn Ala Glu Thr Phe Thr Phe His Ala Asp
500 505 510Ile Cys Thr Leu Ser Glu Lys Glu Arg Gln Ile Lys Lys Gln
Thr Ala 515 520 525Leu Val Glu Leu Val Lys His Lys Pro Lys Ala Thr
Lys Glu Gln Leu 530 535 540Lys Ala Val Met Asp Asp Phe Ala Ala Phe
Val Glu Lys Cys Cys Lys545 550 555 560Ala Asp Asp Lys Glu Thr Cys
Phe Ala Glu Glu Gly Lys Lys Leu Val 565 570 575Ala Ala Ser Gln Ala
Ala Leu Gly Leu 580 585
References
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.