Complement Factor H Antibodies
Patz, Jr.; Edward F. ; et al.
U.S. patent application number 17/076042 was filed with the patent office on 2021-03-04 for complement factor h antibodies. The applicant listed for this patent is Duke University. Invention is credited to Michael J. Campa, Elizabeth Gottlin, Barton F. Haynes, Hua-Xin Liao, Michael Anthony Moody, Edward F. Patz, Jr..
| Application Number | 20210061891 17/076042 |
| Document ID | / |
| Family ID | 1000005164338 |
| Filed Date | 2021-03-04 |












View All Diagrams
| United States Patent Application | 20210061891 |
| Kind Code | A1 |
| Patz, Jr.; Edward F. ; et al. | March 4, 2021 |
COMPLEMENT FACTOR H ANTIBODIES
Abstract
Disclosed herein are Complement factor H (CFH) antibodies and methods of using the antibodies.
| Inventors: | Patz, Jr.; Edward F.; (Durham, NC) ; Campa; Michael J.; (Durham, NC) ; Gottlin; Elizabeth; (Durham, NC) ; Haynes; Barton F.; (Durham, NC) ; Liao; Hua-Xin; (Durham, NC) ; Moody; Michael Anthony; (Durham, NC) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005164338 | ||||||||||
| Appl. No.: | 17/076042 | ||||||||||
| Filed: | October 21, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15973345 | May 7, 2018 | 10865238 | ||
| 17076042 | ||||
| 62502448 | May 5, 2017 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 16/18 20130101; C07K 16/2887 20130101; A61P 35/00 20180101; C07K 2317/734 20130101; C07K 2317/21 20130101; A61K 2039/507 20130101; C07K 2317/34 20130101 |
| International Class: | C07K 16/18 20060101 C07K016/18; A61P 35/00 20060101 A61P035/00; C07K 16/28 20060101 C07K016/28 |
Claims
1. An isolated antibody or antibody fragment thereof which immunospecifically binds to Complement Factor H (CFH) protein, wherein the antibody comprises: i. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:11, a CDR2 comprising the amino acid sequence of SEQ ID NO:21, and a CDR comprising the amino acid sequence of SEQ ID NO:31, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:51, a CDR2 comprising the amino acid sequence of SEQ ID NO:61, and a CDR3 comprising the amino acid sequence of SEQ ID NO:71; ii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:12, a CDR2 comprising the amino acid sequence of SEQ ID NO:22, and a CDR comprising the amino acid sequence of SEQ ID NO:32, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:52, a CDR2 comprising the amino acid sequence of SEQ ID NO:62, and a CDR3 comprising the amino acid sequence of SEQ ID NO:72; iii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:13, a CDR2 comprising the amino acid sequence of SEQ ID NO:23, and a CDR comprising the amino acid sequence of SEQ ID NO:33, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:53, a CDR2 comprising the amino acid sequence of SEQ ID NO:63, and a CDR3 comprising the amino acid sequence of SEQ ID NO:73; iv. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:14, a CDR2 comprising the amino acid sequence of SEQ ID NO:24, and a CDR comprising the amino acid sequence of SEQ ID NO:34, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:54, a CDR2 comprising the amino acid sequence of SEQ ID NO:64, and a CDR3 comprising the amino acid sequence of SEQ ID NO:74; v. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:15, a CDR2 comprising the amino acid sequence of SEQ ID NO:25, and a CDR comprising the amino acid sequence of SEQ ID NO:35, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:55, a CDR2 comprising the amino acid sequence of SEQ ID NO:65, and a CDR3 comprising the amino acid sequence of SEQ ID NO:75; vi. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:16, a CDR2 comprising the amino acid sequence of SEQ ID NO:26, and a CDR comprising the amino acid sequence of SEQ ID NO:36, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:56, a CDR2 comprising the amino acid sequence of SEQ ID NO:66, and a CDR3 comprising the amino acid sequence of SEQ ID NO:76; vii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:17, a CDR2 comprising the amino acid sequence of SEQ ID NO:27, and a CDR comprising the amino acid sequence of SEQ ID NO:37, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:57, a CDR2 comprising the amino acid sequence of SEQ ID NO:67, and a CDR3 comprising the amino acid sequence of SEQ ID NO:77; viii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:18, a CDR2 comprising the amino acid sequence of SEQ ID NO:28, and a CDR comprising the amino acid sequence of SEQ ID NO:38, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:58, a CDR2 comprising the amino acid sequence of SEQ ID NO:68, and a CDR3 comprising the amino acid sequence of SEQ ID NO:78; ix. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:19, a CDR2 comprising the amino acid sequence of SEQ ID NO:29, and a CDR comprising the amino acid sequence of SEQ ID NO:39, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:59, a CDR2 comprising the amino acid sequence of SEQ ID NO:69, and a CDR3 comprising the amino acid sequence of SEQ ID NO:79; or x. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:20, a CDR2 comprising the amino acid sequence of SEQ ID NO:30, and a CDR comprising the amino acid sequence of SEQ ID NO: 40, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:60, a CDR2 comprising the amino acid sequence of SEQ ID NO:70, and a CDR3 comprising the amino acid sequence of SEQ ID NO:80.
2. The isolated antibody or antibody fragment of claim 1, wherein the isolated antibody or antibody fragment thereof binds to an epitope within short consensus repeat (SCR) 19 of CFH protein and the epitope is a cryptic epitope that is exposed in a tumor-bound form of CFH.
3. The isolated antibody or antibody fragment of claim 2, wherein the epitope comprises PIDNGDIT (SEQ ID NO: 3), or fragment or variant thereof.
4. The isolated antibody or antibody fragment of claim 1, wherein the antibody comprises: a) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:1 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:41; b) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:2 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:42; c) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:3 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:43; d) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:4 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:44; e) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:5 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:45; f) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:6 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:46; g) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:7 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:47; h) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:8 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:48; i) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:9 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:49; or j) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:10 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:50.
5. The isolated antibody or antibody fragment of claim 1, wherein the isolated antibody or antibody fragment does not cross-react with at least one of systemic lupus erythematosus autoantigens SSA, SSB, sphingomyelin (Sm), ribonucleoprotein (RNP), sclerosis autoantigen (Scl-70), histidine-tRNA ligase (Jo-1), double-stranded DNA (dsDNA), centromere B (CentB), and histones.
6. The isolated antibody or antibody fragment of claim 1, wherein the isolated antibody or antibody fragment is selected from the group consisting of a human antibody, an immunoglobulin molecule, a disulfide linked Fv, a monoclonal antibody, an affinity matured, a scFv, a chimeric antibody, a single domain antibody, a CDR-grafted antibody, a diabody, a humanized antibody, a multispecific antibody, a Fab, a dual specific antibody, a DVD, a TVD, a Fab', a bispecific antibody, a F(ab')2, and a Fv.
7. The isolated antibody or antibody fragment of claim 1, wherein the isolated antibody or antibody fragment comprises a heavy chain immunoglobulin constant domain selected from the group consisting of a human IgM constant domain, a human IgG4 constant domain, a human IgG1 constant domain, a human IgE constant domain, a human IgG2 constant domain, a human igG3 constant domain, and a human IgA constant domain.
8. A pharmaceutical composition comprising the isolated antibody or antibody fragment of claim 1.
9. A method of treating a subject in need thereof having cancer or inhibiting tumor growth in a subject, the method comprising administering to the subject the isolated antibody or antibody fragment of claim 1.
10. The method of claim 9, wherein the cancer is lung cancer.
11. The method of claim 9, further comprising administering an effective amount of at least one of Cetuximab, PERJETA.RTM., or HERCEPTIN.RTM..
12. A method of increasing complement dependent lysis of a cell, increasing C3b deposition on a cell, or inhibiting Complement Factor H (CFH) binding to C3b in a subject or a cell, the method comprising administering to the cell the isolated antibody or antibody fragment of claim 1.
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. patent application Ser. No. 15/973,345, filed May 7, 2018, which claims priority to U.S. Provisional Application No. 62/502,448, filed May 5, 2017, which is incorporated herein by reference in its entirety
STATEMENT OF GOVERNMENT INTEREST
[0002] Not applicable.
SEQUENCE LISTING
[0003] The instant application contains a Sequence Listing which has been submitted electronically in ASCII format and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Oct. 20, 2020, is named 028193-9232-US03_As_Filed_Sequence_Listing and is 66,036 bytes in size.
TECHNICAL FIELD
[0004] The present disclosure relates to Complement factor H (CFH) antibodies.
BACKGROUND
[0005] Lung cancer is a significant public health issue. In 2016, there were an estimated 1,685,210 new cases of cancer in the United States and 595,690 deaths. The majority of tumors are detected at an advanced stage when treatment options are limited and patients require systemic therapy. Even patients with resectable, early stage lung cancer have an almost 50% chance of developing recurrence and at some point need adjuvant treatment. Over the past several years new therapies targeting specific pathways have been introduced and, in select individuals, these produce an initial response. However, almost all patients develop resistance, which is most likely due to tumor heterogeneity and clonal evolution.
[0006] While activation of the humoral response against malignant cells has been investigated, humoral immunity per se has not been very well exploited for cancer therapy. Circulating antibodies against over 100 different tumor-associated antigens (TAAs) have been described, but very few are associated with tumor stage or outcome. Certain host antibodies may have the potential for anti-tumor activity, but this ability has not been fully realized for a number of possible reasons, including low concentration or low affinity of antibodies, or ineffective activation of B lymphocytes.
[0007] Engagement of both the innate and adaptive immune systems has been suggested as an approach that would improve anti-tumor immunity. A fundamental component of the innate response is the complement system, which when activated causes inflammation and modulates both T and B cell activity. CFH protects tumor cells from lysis by complement dependent cytotoxicity (CDC) by unleashing a proteolytic complement cascade, culminating in the formation of cell-lytic membrane attack complexes in the cell membrane. CFH is a regulatory protein that protects host cells from attack and destruction by the alternative pathway of complement-mediated cytotoxicity (CDC).
[0008] The alternative pathway is one of three complement pathways: The classical pathway is initiated by antibody-antigen complexes, the alternative pathway is spontaneous or can be initiated by pathogens or cell damage, and the lectin pathway is initiated spontaneously or by antigens. All three pathways converge at the cleavage of C3 into C3a and C3b by homologous proteases called convertases. In addition to being a subunit of the alternative pathway C3 convertase, C3b covalently attaches to the target cell membrane, acting as an opsonin that induces ingestion by phagocytes. The multi-protein complement cascade initiated by C3b deposition on the cell membrane culminates with the formation of cell-lytic membrane attack complexes (MAC). The cleavage of C5 during this process generates C5a and C5b. C5b participates in terminal MAC formation. C3a and C5a, the small proteolytic fragments of C3 and C5 cleavage, respectively, are anaphylatoxins; i.e., complement peptides with inflammatory and chemoattractant activity. The anaphylatoxins interact with many cell types of the innate and adaptive immune systems, including monocytes, macrophages, dendritic cells, B cells, and T cells, inducing chemotactic and inflammatory responses.
[0009] CFH prevents the deposition of complement protein C3b on the cell surface by promoting the cleavage of C3b by the complement factor I protease, and by preventing the formation of and accelerating the decay of the alternative pathway convertase that forms C3b from its precursor, C3. Thus, CFH inhibition of the deposition of C3b on the cell surface protects against cell lysis. Tumor cells take advantage of the protection conferred by CFH to evade destruction by the complement system and higher expression of CFH in NSCLC adenocarcinoma has been correlated with worse prognosis and poorer survival. By neutralizing this protective protein, antibodies to CFH may allow complement activation and tumor cell lysis, suppressing tumor growth. In addition, since the C3a and C5a that are generated during the complement cascade interact with immune cells, antibodies to CFH could potentially impact the adaptive immune response. There remains a need for a greater number and wider variety of effective therapies against cancer, such as lung cancer.
SUMMARY
[0010] The present invention is directed an isolated antibody or antibody fragment thereof which immunospecifically binds to Complement Factor H (CFH) protein. The antibody comprises: i. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:11, a CDR2 comprising the amino acid sequence of SEQ ID NO:21, and a CDR comprising the amino acid sequence of SEQ ID NO:31, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:51, a CDR2 comprising the amino acid sequence of SEQ ID NO:61, and a CDR3 comprising the amino acid sequence of SEQ ID NO:71; ii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:12, a CDR2 comprising the amino acid sequence of SEQ ID NO:22, and a CDR comprising the amino acid sequence of SEQ ID NO:32, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:52, a CDR2 comprising the amino acid sequence of SEQ ID NO:62, and a CDR3 comprising the amino acid sequence of SEQ ID NO:72; iii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:13, a CDR2 comprising the amino acid sequence of SEQ ID NO:23, and a CDR comprising the amino acid sequence of SEQ ID NO:33, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:53, a CDR2 comprising the amino acid sequence of SEQ ID NO:63, and a CDR3 comprising the amino acid sequence of SEQ ID NO:73; iv. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:14, a CDR2 comprising the amino acid sequence of SEQ ID NO:24, and a CDR comprising the amino acid sequence of SEQ ID NO:34, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:54, a CDR2 comprising the amino acid sequence of SEQ ID NO:64, and a CDR3 comprising the amino acid sequence of SEQ ID NO:74; v. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:15, a CDR2 comprising the amino acid sequence of SEQ ID NO:25, and a CDR comprising the amino acid sequence of SEQ ID NO:35, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:55, a CDR2 comprising the amino acid sequence of SEQ ID NO:65, and a CDR3 comprising the amino acid sequence of SEQ ID NO:75; vi. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:16, a CDR2 comprising the amino acid sequence of SEQ ID NO:26, and a CDR comprising the amino acid sequence of SEQ ID NO:36, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:56, a CDR2 comprising the amino acid sequence of SEQ ID NO:66, and a CDR3 comprising the amino acid sequence of SEQ ID NO:76; vii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:17, a CDR2 comprising the amino acid sequence of SEQ ID NO:27, and a CDR comprising the amino acid sequence of SEQ ID NO:37, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:57, a CDR2 comprising the amino acid sequence of SEQ ID NO:67, and a CDR3 comprising the amino acid sequence of SEQ ID NO:77; viii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:18, a CDR2 comprising the amino acid sequence of SEQ ID NO:28, and a CDR comprising the amino acid sequence of SEQ ID NO:38, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:58, a CDR2 comprising the amino acid sequence of SEQ ID NO:68, and a CDR3 comprising the amino acid sequence of SEQ ID NO:78; ix. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:19, a CDR2 comprising the amino acid sequence of SEQ ID NO:29, and a CDR comprising the amino acid sequence of SEQ ID NO:39, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:59, a CDR2 comprising the amino acid sequence of SEQ ID NO:69, and a CDR3 comprising the amino acid sequence of SEQ ID NO:79; or x. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:20, a CDR2 comprising the amino acid sequence of SEQ ID NO:30, and a CDR comprising the amino acid sequence of SEQ ID NO: 40, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:60, a CDR2 comprising the amino acid sequence of SEQ ID NO:70, and a CDR3 comprising the amino acid sequence of SEQ ID NO:80.
[0011] The present invention is directed a pharmaceutical composition comprising the isolated antibody or antibody fragment described above.
[0012] The present invention is directed a method of treating a subject in need thereof having cancer or inhibiting tumor growth in a subject, the method comprising administering to the subject the isolated antibody or antibody fragment described above.
[0013] The present invention is directed a method of increasing complement dependent lysis of a cell, increasing C3b deposition on a cell, or inhibiting Complement Factor H (CFH) binding to C3b in a subject or a cell, the method comprising administering to the cell the isolated antibody or antibody fragment described above.
BRIEF DESCRIPTION OF THE DRAWINGS
[0014] FIG. 1 shows the peptide ELISA signal data for representative overlapping peptides of a synthesized library of 115 overlapping peptides comprising all of SCR19-20 screened with three autoantibodies. The beginning and ending amino acid residue number within CFH (UniProt P08603) are noted at the beginning and end of each peptide. The residues in bold are those that are present in all peptides giving the highest signal. Residues comprising only part of this epitope are in italics. Data generated by PepScan, The Netherlands.
[0015] FIG. 2 shows CFH binding to A549 lung cancer cells. Antibodies tested are human IgG and human anti-CFH autoantibody from patient G (AbG). Experiments were performed in the presence of CFH depleted serum. Fluorescently labeled CFH was reduced using TCEP. CFH binding is reported as fluorescence intensity after subtraction of the baseline fluorescence intensity observed using CFH-depleted serum without fluorescently labeled CFH added to the reaction. P-values were calculated by Student's t-test
[0016] FIG. 3 shows deposition of C3-related fragments on A549 lung cancer cells. Antibodies are human IgG and human anti-CFH autoantibody from patient E (AbE), tested with or without blocking peptide (pep), in the presence of NHS or HI-NETS. Fold increase in C3 deposition is reported relative to the baseline observed in the absence of serum. P-values were calculated by Student's t-test.
[0017] FIG. 4 shows complement-dependent cytotoxicity of A549 lung cancer cells by the alternative pathway. Antibodies tested are human IgG and human anti-CFH autoantibody from patient E (AbE), with or without blocking peptide (pep). Fold increase in cytotoxicity is reported relative to the baseline observed in the presence of normal human serum (NHS). P-values were calculated by Student's t-test.
[0018] FIG. 5 shows the effect of reduction and denaturation of CFH on autoantibody binding. ELISA plate wells were coated with either native CFH or CFH treated with the reductant TCEP+/-the denaturant urea. Titration curves were generated using serum from a CFH-antibody positive individual and antibody binding was detected with anti-human IgG-HRP.
[0019] FIG. 6 shows immunoblot analysis of 7 recombinant CFH mAbs on native and reduced CFH and SCR19-20. Equimolar amounts of CFH and SCR19-20 proteins (.about.150 kDa and .about.15 kDa, respectively) were treated with Tris-HCl (oxidized) or TCEP (reduced) prior to electrophoresis. The proteins were transferred to PVDF, and the membranes were placed into a "Surf-Blot" apparatus for immunoblot analysis. Proteins on the membranes were reacted with recombinant CFH mAbs 1-6 and 8 or subtype-matched negative control mAbs 7, 9 and 10, loaded into individual slots. Binding was visualized with an antihuman IgG-HRP conjugate and chemiluminescent substrate, exposing the entire blot to one piece of film. (The relevant sections are shown.) Molecular mass markers are to the left, in kDa. Identity of CFH mAbs by well number: 1=7955; 2=7957; 3=7960; 4=7964; 5=7979; 6=7961; 8=7968.
[0020] FIGS. 7A-7B show the structure of Fab7968 in complex with CFH peptide. FIG. 7A shows Fab 7968 shown with the heavy chain in green and the light chain in blue. The paratope with bound peptide in red is oriented toward the top of the figure, and the CDRs are marked and labeled. FIG. 7B shows CFH peptide in its antibody-bound conformation (red) contrasted in a superposition to the same region in the natively folded CFH protein (gray). Here, only the 7968 epitope is highlighted in red (antibody-bound conformation) and yellow (on the natively folded protein).
[0021] FIG. 8 shows alanine scanning mutagenesis: binding of human CFH mAbs to alanine-substituted epitope peptides. Biotinylated 15mer peptides, mutated in one of 8 positions (CFH1114-1121) to create alanine substitutions, were immobilized on streptavidin and binding of each of 7 antibodies to each peptide was determined by ELISA. Signal vs. antibody concentration curves were generated and the area under the curve was determined for each peptide-antibody pair. Each number in the figure represents the percent of antibody binding to the Ala-containing peptide vs. the naturally occurring peptide. Colors were overlaid by the "Conditional Formatting" function in Microsoft Excel using a gradient from highest (green) to lowest (red).
[0022] FIG. 9 shows CDC mediated by the recombinant human CFH mAb. A549 (NSCLC--adenocarcinoma), H226 (NSCLC--squamous cell carcinoma), H460 (NSCLC--large cell carcinoma), DMS79 (small cell lung cancer), SKBR3 (breast adenocarcinoma), and KATOIII (gastric carcinoma) cell lines were treated with mAb7968 or a subtype-matched negative control mAb (Neg). Normal human serum (NHS) was added as a complement source; heat-inactivated NHS (HI-NHS) was added as a negative control. P-values were calculated by Student's t-test.
[0023] FIG. 10 shows CDC of rituximab-refractory B cells mediated by the combination of CFH mAb7968 and rituximab. B cells from a CLL patient were treated with CFH mAb7968 or a subtype-matched negative control mAb (Neg) at 200 .mu.g/ml, +/-rituximab (RTX) at 100 .mu.g/ml, in the presence of NHS as a source of complement. CDC was assayed by LDH release. The P-value was calculated by Student's t-test.
[0024] FIG. 11 shows augmentation of mAb7968-mediated complement-mediated cytotoxicity (CDC) by cetuximab. CDC was measured by lactate dehydrogenase (LDH) release, performed using the CytoTox 96 Non-radioactive Cytotoxicity Assay kit (Promega, Madison, Wis.). A549 cells (5.times.10.sup.3 per well) were plated overnight at 37.degree. C. in 96-well plates. A mixture of 1:8 diluted normal human serum (NETS), used as a source of complement (Complement Tech, Inc.), antibodies, and medium was then added to the cells. Cetuximab was used at 50 .mu.g/ml and anti-CFH mAb7968 or negative control antibodies at .about.100 .mu.g/ml. Cells were incubated overnight at 37.degree. C. after which LDH release into the medium was measured. All reactions were performed in triplicate. P-values were calculated by Student's t-test.
[0025] FIGS. 12A-12B show response to CFH mAb 7968 by tumor cells. FIG. 12A shows release of C3a from A549 or H226 lung cancer cells. Cells were incubated in the presence of either NHS alone, NHS plus mAb7968, or NHS plus human IgG. C3a was measured in cell supernatants at 1 hr and 4 hr by ELISA. FIG. 12B shows release of C5a from A549 or H226 cells. Using the same cell supernatants described in FIG. 12A, C5a was measured by ELISA. FIG. 12C shows C5b-9 deposition on A549 or H226 cells. After incubation with NHS, NHS plus mAb7968, or NHS plus a subtype-matched negative control mAb (Neg), C5b-9 deposition on cells was measured by flow cytometry. P-values were calculated by Student's t-test.
[0026] FIGS. 13A-13B show DC activation and migration by supernatant from CDC reactions. FIG. 13A shows DC Activation. PBMCs were incubated for 24 hr with supernatants from CDC reactions with CFH mAb7968 or IgG control mAb. Activated DC were gated as HLA-DR+CD25+. The percentage cells of each phenotypic class are shown in the quadrants. (The HLA-DR+CD25+ percentages are boxed.). FIG. 13B shows DC Migration. PBMCs were placed on one side of a transwell chamber, supernatant from CDC reactions carried out in the presence of CFH mAb7968 or control IgG was placed in the other. After PBMC migration, DCs that had migrated were counted by flow cytometry. Percent of the total migrated PBMCs in each chamber that were DCs is plotted.
[0027] FIG. 14 shows translocation of calreticulin to the plasma membrane during CFH-induced CDC. H460 or A549 cells were treated for 2.5 hr with 1, medium; 2, NHS and CFH mAb (200 .mu.g/ml); 3, NHS and IgG (200 .mu.g/ml); or 4, NHS and Doxorubicin (25 .mu.M). Panel A. shows plasma membranes were isolated, and proteins subjected to western blot analysis using a calreticulin mAb. Panel B shows whole cell lysates were prepared from cells undergoing the same treatments and analyzed as in panel A. Panel C shows the blot in panel A was stripped and reprobed with a pan-actin mAb.
[0028] FIGS. 15A-15C show in vivo inhibition of tumor growth with murine mAb7968. Groups of N=5 mice bearing human brain tumor xenografts were injected with either no mAb, murine mAb7968, or subtype-matched negative control murine mAb. FIG. 15A shows growth curve for all animals in each group. The mean tumor growth rate is reported for each group. FIG. 15B shows Kaplan-Meier survival curves by treatment group. FIG. 15C shows representative H&E images of the tumors. The tumor section in the control mouse shows confluent tumor cells, while a palpable subcutaneous mass in the mAb7968-treated mouse shows predominantly chronic inflammatory cells including numerous plasma cells.
[0029] FIGS. 16A-16B show inhibition of tumor growth in the KLN205-DBA/2 syngeneic lung cancer model with CFH mAb treatment. FIG. 16A shows growth curves. KLN205 tumor cells were injected s.c. on day 0 and murine CFH mAb7968 or subtype-matched control mAb was injected intraperitoneally every 3 days between days 1-13, after which treatment was stopped. The mean tumor volumes+/-SEM for t=7 mice treated with each mAb are plotted. P-values for the difference in tumor volumes between mAb7968 and control mAb on days 39, 42, and 45 were 0.027, 0.030, and 0.077, respectively. FIG. 16B shows representative hematoxylin and eosin images of the tumors. A lymphocytic component is present in residual mAb7968-treated tumor. Magnification is 100.times..
[0030] FIGS. 17A-17B show the sequence alignment (FIG. 17A) and percent identities (FIG. 17B) of the variable heavy chains (VH) of the CFH antibodies.
[0031] FIG. 18 shows the sequence alignment and percent identities of the heavy chain CDR1 region (HCDR1) of the CFH antibodies.
[0032] FIG. 19 shows the sequence alignment and percent identities of the heavy chain CDR2 region (HCDR2) of the CFH antibodies.
[0033] FIG. 20 shows the sequence alignment and percent identities of the heavy chain CDR3 region (HCDR3) of the CFH antibodies.
[0034] FIG. 21 shows the sequence alignment and percent identities of the variable light chains (VL) of the CFH antibodies.
[0035] FIG. 22 shows the sequence alignment and percent identities of the light chain CDR1 region (LCDR1) of the CFH antibodies.
[0036] FIG. 23 shows the sequence alignment and percent identities of the light chain CDR2 region (LCDR2) of the CFH antibodies.
[0037] FIG. 24 shows the sequence alignment and percent identities of the light chain CDR3 region (LCDR3) of the CFH antibodies.
DETAILED DESCRIPTION
[0038] The present disclosure is directed to Complement Factor H (CFH) antibodies, and uses of said antibodies, which recognize a reduced form of CFH in vitro, which may represent (or mimic) the tumor-bound form of CFH. Patients with stage I non-small cell lung cancer (NSCLC) have a significantly higher incidence of an antibody to Complement Factor H (CFH) than those with late-stage NSCLC and have no apparent side-effects attributable to the antibody. This disclosure relates to the discovery of an autoantibody to CFH in early stage lung cancer patients who do not develop metastasis. Although CFH is in the blood and binds to normal as well as cancer cells, CFH autoantibodies recognize a conformationally distinct epitope in CFH that appears to confer tumor specificity. "Exceptional outcomes patients," those with early stage lung cancer who never develop recurrence or metastasis, were studied to determine how these patients potentially used the host response to escape disease progression and to apply this knowledge to the development of new therapies. Administering CFH antibodies would not only activate the complement system causing direct tumor cell cytotoxicity, but produce a robust adaptive immune response for long term anti-tumor activity.
[0039] In an effort to develop a cancer therapy that would recapitulate the native immune response in cancer patients, human CFH mAbs were expressed and purified starting from the B cells of patients with CFH autoantibodies. The disclosed CFH antibodies target an epitope in the SCR 19 domain of CFH, which is the functional domain of CFH that interacts with C3b. The CFH antibodies inhibits tumor growth without side-effects. This particular epitope or region was discovered by characterizing the humoral immune response in cancer to develop therapeutic agents against lung cancer. The disclosed antibodies can increased C3b deposition on tumor cells and increased complement dependent lysis of tumor cells. This discovery provides a therapeutic target for cancer treatment. The antibodies inhibit CFH activity on tumor cells, activate complement, promote the formation of cell lytic complexes, and cause CDC of tumor cells. The disclosed antibodies caused CDC of a wide range of tumor cell types. By inactivating a tumor protective protein, the CFH autoantibodies can unleash the complement system on tumor cells and thereby forestall metastasis. In addition, the cytokines that are released upon CDC are expected to activate the adaptive immune response, provoking long-term tumor immunity. One particular CFH mAb recognizes a conformationally distinct epitope, kills tumor cells in vitro, and inhibits tumor growth in vivo with no apparent side effects.
[0040] The mechanism of this antibody may be that: i, the CFH mAb initially kills tumor cells by unleashing the alternative complement pathway from its constraint by CFH and ii, complement lysis releases signaling molecules that attract antigen presenting cells, leading to cross-priming of CD8+ T cells, and an adaptive immune response. Tumors of animals treated with the CFH mAb are infiltrated with lymphocytes.
1. DEFINITIONS
[0041] Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art. In case of conflict, the present document, including definitions, will control. Preferred methods and materials are described below, although methods and materials similar or equivalent to those described herein can be used in practice or testing of the present invention. All publications, patent applications, patents and other references mentioned herein are incorporated by reference in their entirety. The materials, methods, and examples disclosed herein are illustrative only and not intended to be limiting.
[0042] The terms "comprise(s)," "include(s)," "having," "has," "can," "contain(s)," and variants thereof, as used herein, are intended to be open-ended transitional phrases, terms, or words that do not preclude the possibility of additional acts or structures. The singular forms "a," "and" and "the" include plural references unless the context clearly dictates otherwise. The present disclosure also contemplates other embodiments "comprising," "consisting of" and "consisting essentially of," the embodiments or elements presented herein, whether explicitly set forth or not.
[0043] For the recitation of numeric ranges herein, each intervening number there between with the same degree of precision is explicitly contemplated. For example, for the range of 6-9, the numbers 7 and 8 are contemplated in addition to 6 and 9, and for the range 6.0-7.0, the number 6.0, 6.1, 6.2, 6.3, 6.4, 6.5, 6.6, 6.7, 6.8, 6.9, and 7.0 are explicitly contemplated.
[0044] The term "administration" or "administering," as used herein refers to providing, contacting, and/or delivery of the CFH antibody by any appropriate route to achieve the desired effect. These agents may be administered to a subject in numerous ways including, but not limited to, orally, ocularly, nasally, intravenously, topically, as aerosols, suppository, etc. and may be used in combination.
[0045] "Affinity Matured Antibody" is used herein to refer to an antibody with one or more alterations in one or more CDRs, which result in an improvement in the affinity (i.e. K.sub.D, k.sub.d or k.sub.a) of the antibody for a target antigen compared to a parent antibody, which does not possess the alteration(s). Exemplary affinity matured antibodies will have nanomolar or even picomolar affinities for the target antigen. Various procedures for producing affinity matured antibodies are known in the art, including the screening of a combinatory antibody library that has been prepared using bio-display. For example, Marks et al., BioTechnology, 10: 779-783 (1992) describes affinity maturation by VH and VL domain shuffling. Random mutagenesis of complementarity determining regions (CDRs) and/or framework residues is described by Barbas et al., Proc. Nat. Acad. Sci. USA, 91: 3809-3813 (1994); Schier et al., Gene, 169: 147-155 (1995); Yelton et al., J. Immunol., 155: 1994-2004 (1995); Jackson et al., J. Immunol., 154(7): 3310-3319 (1995); and Hawkins et al, J. Mol. Biol., 226: 889-896 (1992). Selective mutation at selective mutagenesis positions and at contact or hypermutation positions with an activity-enhancing amino acid residue is described in U.S. Pat. No. 6,914,128 B1.
[0046] "Alternative pathway", also known as "alternative complement pathway", as used herein refers to one of three complement pathways that opsonize and kill target cells. The alternative pathway is triggered when the C3b protein directly binds the target cell. The alternative complement pathway is able to distinguish self from non-self on the basis of the surface expression of complement regulatory proteins which limit the activation of the complement as host-cells do not accumulate cell surface C3b because this is prevented by the complement regulatory proteins. Foreign cells, pathogens and abnormal surfaces generally do not have complement regulatory proteins and thus may become heavily decorated with C3b, which eventually leads to the lysis of the cell.
[0047] "Antibody" and "antibodies" as used herein refers to monoclonal antibodies, multispecific antibodies, human antibodies, humanized antibodies (fully or partially humanized), animal antibodies such as, but not limited to, a bird (for example, a duck or a goose), a shark, a whale, and a mammal, including a non-primate (for example, a cow, a pig, a camel, a llama, a horse, a goat, a rabbit, a sheep, a hamster, a guinea pig, a cat, a dog, a rat, a mouse, etc.) or a non-human primate (for example, a monkey, a chimpanzee, etc.), recombinant antibodies, chimeric antibodies, single-chain Fvs ("scFv"), single chain antibodies, single domain antibodies, such as variable heavy chain domains ("VHH"; also known as "VHH fragments") derived from animals in the Camelidae family (VHH and methods of making them are described in Gottlin et al., Journal of Biomolecular Screening, 14:77-85 (2009)) and V.sub.NAR fragments, Fab fragments, F(ab') fragments, F(ab')2 fragments, disulfide-linked Fvs ("sdFv"), and anti-idiotypic ("anti-Id") antibodies, dual-domain antibodies, dual variable domain (DVD) or triple variable domain (TVD) antibodies (dual-variable domain immunoglobulins and methods for making them are described in Wu, C., et al., Nature Biotechnology, 25(11):1290-1297 (2007)) and PCT International Application WO 2001/058956, the contents of each of which are herein incorporated by reference), and functionally active epitope-binding fragments of any of the above. In particular, antibodies include immunoglobulin molecules and immunologically active fragments of immunoglobulin molecules, namely, molecules that contain an analyte-binding site. Immunoglobulin molecules can be of any type (for example, IgG, IgE, IgM, IgD, IgA, and IgY), class (for example, IgG1, IgG2, IgG3, IgG4, IgA1, and IgA2), or subclass. For simplicity sake, an antibody against an analyte is frequently referred to herein as being either an "anti-analyte antibody" or merely an "analyte antibody" (e.g., an anti-CFH antibody or a CFH antibody).
[0048] "Antibody fragment" as used herein refers to a portion of an intact antibody comprising the antigen-binding site or variable region. The portion does not include the constant heavy chain domains (i.e. CH2, CH3, or CH4, depending on the antibody isotype) of the Fc region of the intact antibody. Examples of antibody fragments include, but are not limited to, Fab fragments, Fab' fragments, Fab'-SH fragments, F(ab')2 fragments, Fd fragments, Fv fragments, diabodies, single-chain Fv (scFv) molecules, single-chain polypeptides containing only one light chain variable domain, single-chain polypeptides containing the three CDRs of the light-chain variable domain, single-chain polypeptides containing only one heavy chain variable region, single-chain polypeptides containing the three CDRs of the heavy chain variable region, and VHH.
[0049] "Autoantibody", "patient antibodies", "patient's CFH autoantibodies" or "patient's CFH antibodies" as used interchangeably herein refers to an immunoglobulin, antigen specific B cell surface receptor (surface immunoglobulin), or antigen specific T cell receptor produced by an individual that is directed against an individual's own self-protein, carbohydrate or nucleic acid.
[0050] An "autoantibody to the CFH protein" as used herein refers to an autoantibody capable of reacting with the CFH protein, or with a variant or with a fragment of said protein, provided that said variant or said fragment is functionally equivalent, i.e., susceptible of being recognized by said autoantibody. For example, an autoantibody to the CFH protein may be an IgG or an IgM.
[0051] "Binding Constants" are described herein. The term "association rate constant," "k.sub.on" or "k.sub.a" as used herein, refers to the value indicating the binding rate of an antibody to its target antigen or the rate of complex formation between an antibody and antigen as shown by the equation below:
Antibody (Ab)+Antigen (Ag).fwdarw.Ab-Ag.
[0052] The term "dissociation rate constant," "k.sub.off" or "k.sub.d" as used interchangeably herein, refers to the value indicating the dissociation rate of an antibody form its target antigen or separation of Ab-Ag complex over time into free antibody and antigen as shown by the equation below:
Antibody (Ab)+Antigen (Ag).rarw.Ab-Ag.
[0053] Methods for determining association and dissociation rate constants are well known in the art. Using fluorescence-based techniques offers high sensitivity and the ability to examine samples in physiological buffers at equilibrium. Other experimental approaches and instruments such as a BIAcore.RTM. (biomolecular interaction analysis) assay can be used (e.g., instrument available from BIAcore International AB, a GE Healthcare company, Uppsala, Sweden). Additionally, a KinExA.RTM. (Kinetic Exclusion Assay) assay, available from Sapidyne Instruments (Boise, Id.) can also be used.
[0054] The term "effective dosage" or "effective amount" as used interchangeably herein means a dosage of a drug effective for periods of time necessary, to achieve the desired therapeutic result. An effective dosage may be determined by a person skilled in the art and may vary according to factors such as the disease state, age, sex, and weight of the individual, and the ability of the drug to elicit a desired response in the individual. This term as used herein may also refer to an amount effective at bringing about a desired in vivo effect in an animal, mammal, or human, such as reducing and/or inhibiting the function of the estrogen receptor. A therapeutically effective amount may be administered in one or more administrations (e.g., the agent may be given as a preventative treatment or therapeutically at any stage of disease progression, before or after symptoms, and the like), applications or dosages and is not intended to be limited to a particular formulation, combination or administration route. It is within the scope of the present disclosure that the SERM may be administered at various times during the course of treatment of the subject. The times of administration and dosages used will depend on several factors, such as the goal of treatment (e.g., treating v. preventing), condition of the subject, etc. and can be readily determined by one skilled in the art.
[0055] The term "equilibrium dissociation constant", "Kd", "K.sub.d" or "K.sub.D" as used interchangeably, herein, refers to the value obtained by dividing the dissociation rate (koff) by the association rate (kon). The association rate, the dissociation rate and the equilibrium dissociation constant are used to represent the binding affinity of an antibody to an antigen.
[0056] "Binding Protein" is used herein to refer to a monomeric or multimeric protein that binds to and forms a complex with a binding partner, such as, for example, a polypeptide, an antigen, a chemical compound or other molecule, or a substrate of any kind. A binding protein specifically binds a binding partner. Binding proteins include antibodies, as well as antigen-binding fragments thereof and other various forms and derivatives thereof as are known in the art and described herein below and other molecules comprising one or more antigen-binding domains that bind to an antigen molecule or a particular site (epitope) on the antigen molecule. Accordingly, a binding protein includes, but is not limited to, an antibody a tetrameric immunoglobulin, an IgG molecule, an IgG1 molecule, a monoclonal antibody, a chimeric antibody, a CDR-grafted antibody, a humanized antibody, an affinity matured antibody, and fragments of any such antibodies that retain the ability to bind to an antigen.
[0057] "Bispecific antibody" is used herein to refer to a full-length antibody that is generated by quadroma technology (see Milstein et al., Nature, 305(5934): 537-540 (1983)), by chemical conjugation of two different monoclonal antibodies (see, Staerz et al., Nature, 314(6012): 628-631 (1985)), or by knob-into-hole or similar approaches, which introduce mutations in the Fc region (see Holliger et al., Proc. Natl. Acad. Sci. USA, 90(14): 6444-6448 (1993)), resulting in multiple different immunoglobulin species of which only one is the functional bispecific antibody. A bispecific antibody binds one antigen (or epitope) on one of its two binding arms (one pair of HC/LC), and binds a different antigen (or epitope) on its second arm (a different pair of HC/LC). By this definition, a bispecific antibody has two distinct antigen-binding arms (in both specificity and CDR sequences), and is monovalent for each antigen to which it binds to.
[0058] "C3b" as used herein refers to the larger of two elements formed by the cleavage of complement component 3 (C3) by C3 convertase enzyme complex or by spontaneous cleavage in the blood. C3b covalently bonds to microbial cell surfaces within an organism's body, leading to the production of surface-bound C3-convertase and more C3b components and opsonization of the microbe by macrophages. C3b that is generated from C3 by a C3 convertase enzyme complex in the fluid phase is rapidly inactivated by factor H and factor I. When the internal thioester of C3 reacts with a hydroxyl or amine group of a molecule on the surface of a cell or pathogen, the C3b that is now covalently bound to the surface is protected from factor H-mediated inactivation and may now bind factor B to form C3bB.
[0059] "Cancer" or "tumor" as used interchangeably herein refers to the uncontrolled and unregulated growth of abnormal cells in the body. Cancer may invade nearby parts of the body and may also spread to more distant parts of the body through the lymphatic system or bloodstream. "Cancer cell" or "tumor cell" as used interchangeably herein refers to a cell that divides and reproduces abnormally with uncontrolled growth. A cancer cell can break away and travel to other parts of the body and set up another site, referred to as metastasis. Cancer cells or cancerous cells are also called malignant cells. A cancer cell or cancer cell line may originate from a cancer. For examples, a cancer cell line may be A549 cell line ("A549"), which is a human lung adenocarcinoma epithelial cell line.
[0060] Cancers may include Adrenocortical Carcinoma, Anal Cancer, Bladder Cancer, Brain Tumor, Breast Cancer, Carcinoid Tumor, Gastrointestinal, Carcinoma of Unknown Primary, Cervical Cancer, Colon Cancer, Endometrial Cancer, Esophageal Cancer, Extrahepatic Bile Duct Cancer, Ewings Family of Tumors (PNET), Extracranial Germ Cell Tumor, Intraocular Melanoma Eye Cancer, Gallbladder Cancer, Gastric Cancer (Stomach), Extragonadal Germ Cell Tumor, Gestational Trophoblastic Tumor, Head and Neck Cancer, Hypopharyngeal Cancer, Islet Cell Carcinoma, Kidney Cancer (renal cell cancer), Laryngeal Cancer, Acute Lymphoblastic Leukemia, Leukemia, Acute Myeloid, Chronic Lymphocytic Leukemia, Chronic Myelogenous Leukemia, Hairy Cell Leukemia, Lip and Oral Cavity Cancer, Liver Cancer, Non-Small Cell Lung Cancer, Small Cell Lung Cancer, AIDS-Related Lymphoma, Central Nervous System (Primary) Lymphoma, Cutaneous T-Cell Lymphoma, Hodgkin's Disease Lymphoma, Non-Hodgkin's Disease Lymphoma, Malignant Mesothelioma, Melanoma, Merkel Cell Carcinoma, Metasatic Squamous Neck Cancer with Occult Primary, Multiple Myeloma and Other Plasma Cell Neoplasms, Mycosis Fungoides, Myelodysplastic Syndrome, Myeloproliferative Disorders, Nasopharyngeal Cancer, euroblastoma, Oral Cancer, Oropharyngeal Cancer, Osteosarcoma, Ovarian Epithelial Cancer, Ovarian Germ Cell Tumor, Pancreatic Cancer, Exocrine, Pancreatic Cancer, Islet Cell Carcinoma, Paranasal Sinus and Nasal Cavity Cancer, Parathyroid Cancer, Penile Cancer, Pituitary Cancer, Plasma Cell Neoplasm, Prostate Cancer, Rhabdomyosarcoma, Rectal Cancer, Renal Cell Cancer (cancer of the kidney), Transitional Cell Renal Pelvis and Ureter, Salivary Gland Cancer, Sezary Syndrome, Skin Cancer, Small Intestine Cancer, Soft Tissue Sarcoma, Testicular Cancer, Malignant Thymoma, Thyroid Cancer, Urethral Cancer, Uterine Cancer, Unusual Cancer of Childhood, Vaginal Cancer, Vulvar Cancer, and Wilms' Tumor.
[0061] "CDR" is used herein to refer to the "complementarity determining region" within an antibody variable sequence. There are three CDRs in each of the variable regions of the heavy chain and the light chain, which are designated "CDR1", "CDR2", and "CDR3", for each of the variable regions. The term "CDR set" as used herein refers to a group of three CDRs that occur in a single variable region that binds the antigen. The exact boundaries of these CDRs have been defined differently according to different systems. The system described by Kabat (Kabat et al., Sequences of Proteins of Immunological Interest (National Institutes of Health, Bethesda, Md. (1987) and (1991)) not only provides an unambiguous residue numbering system applicable to any variable region of an antibody, but also provides precise residue boundaries defining the three CDRs. These CDRs may be referred to as "Kabat CDRs". Chothia and coworkers (Chothia and Lesk, J. Mol. Biol., 196: 901-917 (1987); and Chothia et al., Nature, 342: 877-883 (1989)) found that certain sub-portions within Kabat CDRs adopt nearly identical peptide backbone conformations, despite having great diversity at the level of amino acid sequence. These sub-portions were designated as "L1", "L2", and "L3", or "H1", "H2", and "H3", where the "L" and the "H" designate the light chain and the heavy chain regions, respectively. These regions may be referred to as "Chothia CDRs", which have boundaries that overlap with Kabat CDRs. Other boundaries defining CDRs overlapping with the Kabat CDRs have been described by Padlan, FASEB J., 9: 133-139 (1995), and MacCallum, J. Mol. Biol., 262(5): 732-745 (1996). Still other CDR boundary definitions may not strictly follow one of the herein systems, but will nonetheless overlap with the Kabat CDRs, although they may be shortened or lengthened in light of prediction or experimental findings that particular residues or groups of residues or even entire CDRs do not significantly impact antigen binding. The methods used herein may utilize CDRs defined according to any of these systems, although certain embodiments use Kabat- or Chothia-defined CDRs. In some embodiments, the CDRs can be determined using Paratome-Antigen Binding Regions Identification Tool ((Kunik et al. (2012) PLoS Comput Biol 8(2): e1002388. doi:10.1371/journal.pcbi.1002388; Kunik et al. (2012) Nucleic Acids Res. 2012 July; 40(Web Server issue):W521-4. doi: 10.1093/nar/gks480. Epub 2012 Jun. 6)).
[0062] "Complement Factor H protein", "CFH protein", or "CFH" as used herein refers to a protein of approximately 150 kDa (UniProt P08603) that is a member of the regulators of complement activation family and is a complement control protein. CFH is a large soluble glycoprotein that circulates in human plasma and serves to regulate the alternative pathway of the complement system, ensuring that the complement system is directed towards pathogens or other dangerous material and does not damage host tissue. CFH is a cofactor in the inactivation of C3b by factor I and functions to increase the rate of dissociation of the C3bBb complex (C3 convertase) and the (C3b)NBB complex (C5 convertase) in the alternative complement pathway. CFH binds to glycosaminoglycans that are generally present on host cells but not, normally, on pathogen surfaces.
[0063] CFH is composed of 20 short consensus repeats (SCRs), some of which function in cell attachment, while others function to eliminate C3b from the cell surface. The 20 SCRs that comprise CFH are each approximately 60 amino acids long, are arranged head to tail, and contain 4 cysteine residues forming 2 disulfide bonds per module. The C3b binding domain may refer to the part of the CFH that binds to C3b. SCRs 19 and 20 are involved in C3b binding.
[0064] "Derivative" of an antibody as used herein may refer to an antibody having one or more modifications to its amino acid sequence when compared to a genuine or parent antibody and exhibit a modified domain structure. The derivative may still be able to adopt the typical domain configuration found in native antibodies, as well as an amino acid sequence, which is able to bind to targets (antigens) with specificity. Typical examples of antibody derivatives are antibodies coupled to other polypeptides, rearranged antibody domains, or fragments of antibodies. The derivative may also comprise at least one further compound, e.g. a protein domain, said protein domain being linked by covalent or non-covalent bonds. The linkage can be based on genetic fusion according to the methods known in the art. The additional domain present in the fusion protein comprising the antibody employed in accordance with the invention may preferably be linked by a flexible linker, advantageously a peptide linker, wherein said peptide linker comprises plural, hydrophilic, peptide-bonded amino acids of a length sufficient to span the distance between the C-terminal end of the further protein domain and the N-terminal end of the antibody or vice versa. The antibody may be linked to an effector molecule having a conformation suitable for biological activity or selective binding to a solid support, a biologically active substance (e.g. a cytokine or growth hormone), a chemical agent, a peptide, a protein, or a drug, for example.
[0065] "Dual-specific antibody" is used herein to refer to a full-length antibody that can bind two different antigens (or epitopes) in each of its two binding arms (a pair of HC/LC) (see PCT publication WO 02/02773). Accordingly, a dual-specific binding protein has two identical antigen binding arms, with identical specificity and identical CDR sequences, and is bivalent for each antigen to which it binds.
[0066] "Dual variable domain" or "DVD" as used interchangeably herein to refer to two or more antigen binding sites on a binding protein, which may be divalent (two antigen binding sites), tetravalent (four antigen binding sites), or multivalent binding proteins. DVDs may be monospecific, i.e., capable of binding one antigen (or one specific epitope), or multispecific, i.e., capable of binding two or more antigens (i.e., two or more epitopes of the same target antigen molecule or two or more epitopes of different target antigens). A preferred DVD binding protein comprises two heavy chain DVD polypeptides and two light chain DVD polypeptides and is referred to as a "DVD immunoglobulin" or "DVD-Ig". Such a DVD-Ig binding protein is thus tetrameric and reminiscent of an IgG molecule, but provides more antigen binding sites than an IgG molecule. Thus, each half of a tetrameric DVD-Ig molecule is reminiscent of one half of an IgG molecule and comprises a heavy chain DVD polypeptide and a light chain DVD polypeptide, but unlike a pair of heavy and light chains of an IgG molecule that provides a single antigen binding domain, a pair of heavy and light chains of a DVD-Ig provide two or more antigen binding sites.
[0067] Each antigen binding site of a DVD-Ig binding protein may be derived from a donor ("parental") monoclonal antibody and thus comprises a heavy chain variable domain (VH) and a light chain variable domain (VL) with a total of six CDRs involved in antigen binding per antigen binding site. Accordingly, a DVD-Ig binding protein that binds two different epitopes (i.e., two different epitopes of two different antigen molecules or two different epitopes of the same antigen molecule) comprises an antigen binding site derived from a first parental monoclonal antibody and an antigen binding site of a second parental monoclonal antibody.
[0068] In a preferred embodiment, a DVD-Ig binding protein according to the invention not only binds the same target molecules bound by its parental monoclonal antibodies, but also possesses one or more desirable properties of one or more of its parental monoclonal antibodies. Preferably, such an additional property is an antibody parameter of one or more of the parental monoclonal antibodies. Antibody parameters that may be contributed to a DVD-Ig binding protein from one or more of its parental monoclonal antibodies include, but are not limited to, antigen specificity, antigen affinity, potency, biological function, epitope recognition, protein stability, protein solubility, production efficiency, immunogenicity, pharmacokinetics, bioavailability, tissue cross reactivity, and orthologous antigen binding.
[0069] A DVD-Ig binding protein binds at least one epitope of a CFH. Non-limiting examples of a DVD-Ig binding protein include a DVD-Ig binding protein that binds one or more epitopes of CFH, a DVD-Ig binding protein that binds an epitope of a human CFH and an epitope of a CFH of another species (for example, mouse), and a DVD-Ig binding protein that binds an epitope of a human CFH and an epitope of another target molecule.
[0070] "Epitope," or "epitopes," or "epitopes of interest" refer to a site(s) on any molecule that is recognized and can bind to a complementary site(s) on its specific binding partner. The molecule and specific binding partner are part of a specific binding pair. For example, an epitope can be on a polypeptide, a protein, a hapten, a carbohydrate antigen (such as, but not limited to, glycolipids, glycoproteins or lipopolysaccharides), or a polysaccharide. Its specific binding partner can be, but is not limited to, an antibody.
[0071] "F(ab')2 fragment" as used herein refers to antibodies generated by pepsin digestion of whole IgG antibodies to remove most of the Fc region while leaving intact some of the hinge region. F(ab')2 fragments have two antigen-binding F(ab) portions linked together by disulfide bonds, and therefore are divalent with a molecular weight of about 110 kDa. Divalent antibody fragments (F(ab')2 fragments) are smaller than whole IgG molecules and enable a better penetration into tissue thus facilitating better antigen recognition in immunohistochemistry. The use of F(ab')2 fragments also avoids unspecific binding to Fc receptor on live cells or to Protein A/G. F(ab')2 fragments can both bind and precipitate antigens.
[0072] "Framework" (FR) or "Framework sequence" as used herein may mean the remaining sequences of a variable region minus the CDRs. Because the exact definition of a CDR sequence can be determined by different systems (for example, see above), the meaning of a framework sequence is subject to correspondingly different interpretations. The six CDRs (CDR-L1, -L2, and -L3 of light chain and CDR-H1, -H2, and -H3 of heavy chain) also divide the framework regions on the light chain and the heavy chain into four sub-regions (FR1, FR2, FR3, and FR4) on each chain, in which CDR1 is positioned between FR1 and FR2, CDR2 between FR2 and FR3, and CDR3 between FR3 and FR4. Without specifying the particular sub-regions as FR1, FR2, FR3, or FR4, a framework region, as referred by others, represents the combined FRs within the variable region of a single, naturally occurring immunoglobulin chain. As used herein, a FR represents one of the four sub-regions, and FRs represents two or more of the four sub-regions constituting a framework region.
[0073] Human heavy chain and light chain FR sequences are known in the art that can be used as heavy chain and light chain "acceptor" framework sequences (or simply, "acceptor" sequences) to humanize a non-human antibody using techniques known in the art. In one embodiment, human heavy chain and light chain acceptor sequences are selected from the framework sequences listed in publicly available databases such as V-base (hypertext transfer protocol://vbase.mrc-cpe.cam.ac.uk/) or in the international ImMunoGeneTics.RTM. (IMGT.RTM.) information system (hypertext transfer protocol://imgt.cines.fr/texts/IMGTrepertoire/LocusGenes/).
[0074] "Functional antigen binding site" as used herein may mean a site on a binding protein (e.g. an antibody) that is capable of binding a target antigen. The antigen binding affinity of the antigen binding site may not be as strong as the parent binding protein, e.g., parent antibody, from which the antigen binding site is derived, but the ability to bind antigen must be measurable using any one of a variety of methods known for evaluating protein, e.g., antibody, binding to an antigen. Moreover, the antigen binding affinity of each of the antigen binding sites of a multivalent protein, e.g., multivalent antibody, herein need not be quantitatively the same.
[0075] "Humanized antibody" is used herein to describe an antibody that comprises heavy and light chain variable region sequences from a non-human species (e.g. a mouse) but in which at least a portion of the VH and/or VL sequence has been altered to be more "human-like," i.e., more similar to human germline variable sequences. A "humanized antibody" is an antibody or a variant, derivative, analog, or fragment thereof, which immunospecifically binds to an antigen of interest and which comprises a framework (FR) region having substantially the amino acid sequence of a human antibody and a complementarity determining region (CDR) having substantially the amino acid sequence of a non-human antibody. As used herein, the term "substantially" in the context of a CDR refers to a CDR having an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 98%, or at least 99% identical to the amino acid sequence of a non-human antibody CDR. A humanized antibody comprises substantially all of at least one, and typically two, variable domains (Fab, Fab', F(ab')2, FabC, Fv) in which all or substantially all of the CDR regions correspond to those of a non-human immunoglobulin (i.e., donor antibody) and all or substantially all of the framework regions are those of a human immunoglobulin consensus sequence. In an embodiment, a humanized antibody also comprises at least a portion of an immunoglobulin constant region (Fc), typically that of a human immunoglobulin. In some embodiments, a humanized antibody contains the light chain as well as at least the variable domain of a heavy chain. The antibody also may include the CH1, hinge, CH2, CH3, and CH4 regions of the heavy chain. In some embodiments, a humanized antibody only contains a humanized light chain. In some embodiments, a humanized antibody only contains a humanized heavy chain. In specific embodiments, a humanized antibody only contains a humanized variable domain of a light chain and/or humanized heavy chain.
[0076] A humanized antibody can be selected from any class of immunoglobulins, including IgM, IgG, IgD, IgA, IgY, and IgE, and any isotype, including without limitation IgG1, IgG2, IgG3, and IgG4. A humanized antibody may comprise sequences from more than one class or isotype, and particular constant domains may be selected to optimize desired effector functions using techniques well-known in the art.
[0077] The framework regions and CDRs of a humanized antibody need not correspond precisely to the parental sequences, e.g., the donor antibody CDR or the consensus framework may be mutagenized by substitution, insertion, and/or deletion of at least one amino acid residue so that the CDR or framework residue at that site does not correspond to either the donor antibody or the consensus framework. In a preferred embodiment, such mutations, however, will not be extensive. Usually, at least 80%, preferably at least 85%, more preferably at least 90%, and most preferably at least 95% of the humanized antibody residues will correspond to those of the parental FR and CDR sequences. As used herein, the term "consensus framework" refers to the framework region in the consensus immunoglobulin sequence. As used herein, the term "consensus immunoglobulin sequence" refers to the sequence formed from the most frequently occurring amino acids (or nucleotides) in a family of related immunoglobulin sequences (see, e.g., Winnaker, From Genes to Clones (Verlagsgesellschaft, Weinheim, 1987)). A "consensus immunoglobulin sequence" may thus comprise a "consensus framework region(s)" and/or a "consensus CDR(s)". In a family of immunoglobulins, each position in the consensus sequence is occupied by the amino acid occurring most frequently at that position in the family. If two amino acids occur equally frequently, either can be included in the consensus sequence.
[0078] "Identical" or "identity," as used herein in the context of two or more polypeptide or polynucleotide sequences, can mean that the sequences have a specified percentage of residues that are the same over a specified region. The percentage can be calculated by optimally aligning the two sequences, comparing the two sequences over the specified region, determining the number of positions at which the identical residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the specified region, and multiplying the result by 100 to yield the percentage of sequence identity. In cases where the two sequences are of different lengths or the alignment produces one or more staggered ends and the specified region of comparison includes only a single sequence, the residues of the single sequence are included in the denominator but not the numerator of the calculation.
[0079] "Linking sequence" or "linking peptide sequence" refers to a natural or artificial polypeptide sequence that is connected to one or more polypeptide sequences of interest (e.g., full-length, fragments, etc.). The term "connected" refers to the joining of the linking sequence to the polypeptide sequence of interest. Such polypeptide sequences are preferably joined by one or more peptide bonds. Linking sequences can have a length of from about 4 to about 50 amino acids. Preferably, the length of the linking sequence is from about 6 to about 30 amino acids. Natural linking sequences can be modified by amino acid substitutions, additions, or deletions to create artificial linking sequences. Exemplary linking sequences include, but are not limited to: (i) Histidine (His) tags, such as a 6.times. His tag, which has an amino acid sequence of HHHEIHE (SEQ ID NO:119), are useful as linking sequences to facilitate the isolation and purification of polypeptides and antibodies of interest; (ii) Enterokinase cleavage sites, like His tags, are used in the isolation and purification of proteins and antibodies of interest. Often, enterokinase cleavage sites are used together with His tags in the isolation and purification of proteins and antibodies of interest. Various enterokinase cleavage sites are known in the art. Examples of enterokinase cleavage sites include, but are not limited to, the amino acid sequence of DDDDK (SEQ ID NO:120) and derivatives thereof (e.g., ADDDDK (SEQ ID NO:121), etc.); (iii) Miscellaneous sequences can be used to link or connect the light and/or heavy chain variable regions of single chain variable region fragments. Examples of other linking sequences can be found in Bird et al., Science 242: 423-426 (1988); Huston et al., PNAS USA 85: 5879-5883 (1988); and McCafferty et al., Nature 348: 552-554 (1990). Linking sequences also can be modified for additional functions, such as attachment of drugs or attachment to solid supports. In the context of the present disclosure, the CFH antibody for example, can contain a linking sequence, such as a His tag, an enterokinase cleavage site, or both.
[0080] "Lung cancer" as used herein refers to cancer that originates in the lung. For example, lung cancer may be cancer of the lung, such as small-cell lung cancer, also known as small-cell lung carcinoma and oat cell cancer, non-small-cell lung carcinoma ("NSCLC"), glandular tumors, carcinoid tumors and undifferentiated carcinomas.
[0081] "Non-small-cell lung carcinoma" or "NSCLC" as used interchangeably herein refers to any type of epithelial lung cancer other than small cell lung carcinoma. The three main subtypes of NSCLC are adenocarcinoma, including bronchioloalveolar carcinoma, squamous-cell lung carcinoma, and large-cell lung carcinoma. NSCLCs are relatively insensitive to chemotherapy.
[0082] "Monoclonal antibody" as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, i.e., the individual antibodies comprising the population are identical except for possible naturally occurring mutations that may be present in minor amounts. Monoclonal antibodies are highly specific, being directed against a single antigen. Furthermore, in contrast to polyclonal antibody preparations that typically include different antibodies directed against different determinants (epitopes), each monoclonal antibody is directed against a single determinant on the antigen. The monoclonal antibodies herein specifically include "chimeric" antibodies in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies derived from a particular species or belonging to a particular antibody class or subclass, while the remainder of the chain(s) is identical with or homologous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies, so long as they exhibit the desired biological properties.
[0083] "Multivalent binding protein" is used herein to refer to a binding protein comprising two or more antigen binding sites (also referred to herein as "antigen binding domains"). A multivalent binding protein is preferably engineered to have three or more antigen binding sites, and is generally not a naturally occurring antibody. The term "multispecific binding protein" refers to a binding protein that can bind two or more related or unrelated targets, including a binding protein capable of binding two or more different epitopes of the same target molecule.
[0084] "Recombinant antibody" and "recombinant antibodies" refer to antibodies prepared by one or more steps, including cloning nucleic acid sequences encoding all or a part of one or more monoclonal antibodies into an appropriate expression vector by recombinant techniques and subsequently expressing the antibody in an appropriate host cell. The terms include, but are not limited to, recombinantly produced monoclonal antibodies, chimeric antibodies, humanized antibodies (fully or partially humanized), multi-specific or multi-valent structures formed from antibody fragments, bifunctional antibodies, heteroconjugate Abs, DVD-Ig.RTM.s, and other antibodies as described in (i) herein. (Dual-variable domain immunoglobulins and methods for making them are described in Wu, C., et al., Nature Biotechnology, 25:1290-1297 (2007)). The term "bifunctional antibody," as used herein, refers to an antibody that comprises a first arm having a specificity for one antigenic site and a second arm having a specificity for a different antigenic site, i.e., the bifunctional antibodies have a dual specificity.
[0085] "Sample," "test sample," "specimen," "sample from a subject," and "patient sample" as used herein may be used interchangeable and may be a sample of blood, tissue, urine, serum, plasma, amniotic fluid, cerebrospinal fluid, placental cells or tissue, endothelial cells, leukocytes, or monocytes. The sample can be used directly as obtained from a patient or can be pre-treated, such as by filtration, distillation, extraction, concentration, centrifugation, inactivation of interfering components, addition of reagents, and the like, to modify the character of the sample in some manner as discussed herein or otherwise as is known in the art.
[0086] Any cell type, tissue, or bodily fluid may be utilized to obtain a sample. Such cell types, tissues, and fluid may include sections of tissues such as biopsy and autopsy samples, frozen sections taken for histologic purposes, blood (such as whole blood), plasma, serum, sputum, stool, tears, mucus, saliva, bronchoalveolar lavage (BAL) fluid, hair, skin, red blood cells, platelets, interstitial fluid, ocular lens fluid, cerebral spinal fluid, sweat, nasal fluid, synovial fluid, menses, amniotic fluid, semen, etc. Cell types and tissues may also include lymph fluid, ascetic fluid, gynecological fluid, urine, peritoneal fluid, cerebrospinal fluid, a fluid collected by vaginal rinsing, or a fluid collected by vaginal flushing. A tissue or cell type may be provided by removing a sample of cells from an animal, but can also be accomplished by using previously isolated cells (e.g., isolated by another person, at another time, and/or for another purpose). Archival tissues, such as those having treatment or outcome history, may also be used. Protein or nucleotide isolation and/or purification may not be necessary.
[0087] "Small consensus repeat" or "SCR" as used interchangeably herein refers to a structure based on a beta-sandwich arrangement where one face is made up of three beta-strands hydrogen bonded to form a triple stranded region at its center and the other face formed from two separate beta-strands. SCRs are also called complement control protein (CCP) modules and sushi domains. SCRs exist in a wide variety of complement and adhesion proteins. As used herein, "SCR19" refers to short consensus repeat domain 19 and "SCR19-20" refers to short consensus repeat domains 19 and 20, covalently linked to one another as in the parent molecule, CFH.
[0088] "Specific binding" or "specifically binding" as used herein may refer to the interaction of an antibody, a protein, or a peptide with a second chemical species, wherein the interaction is dependent upon the presence of a particular structure (e.g., an antigenic determinant or epitope) on the chemical species; for example, an antibody recognizes and binds to a specific protein structure rather than to proteins generally. If an antibody is specific for epitope "A", the presence of a molecule containing epitope A (or free, unlabeled A), in a reaction containing labeled "A" and the antibody, will reduce the amount of labeled A bound to the antibody.
[0089] "Subject" and "patient" as used herein interchangeably refers to any vertebrate, including, but not limited to, a mammal (e.g., cow, pig, camel, llama, horse, goat, rabbit, sheep, hamsters, guinea pig, cat, dog, rat, and mouse, a non-human primate (for example, a monkey, such as a cynomolgous or rhesus monkey, chimpanzee, etc.) and a human). In some embodiments, the subject may be a human or a non-human. The subject or patient may be undergoing other forms of treatment.
[0090] "Target region" or "molecular target" as used interchangeably herein refers to a region of CFH to which, for example, CFH antibodies, such as the anti-CFH antibodies, may bind. For example, the target region may include SCR 19 and/or the amino acid sequence of PIDNGDIT (SEQ ID NO:83). The target region may include a 15-mer peptide of GPPPPIDNGDITSFP (SEQ ID NO:84).
[0091] "Treat", "treating" or "treatment" are each used interchangeably herein to describe reversing, alleviating, or inhibiting the progress of a disease, or one or more symptoms of such disease, to which such term applies. Depending on the condition of the subject, the term also refers to preventing a disease, and includes preventing the onset of a disease, or preventing the symptoms associated with a disease. A treatment may be either performed in an acute or chronic way. The term also refers to reducing the severity of a disease or symptoms associated with such disease prior to affliction with the disease. Such prevention or reduction of the severity of a disease prior to affliction refers to administration of an antibody or pharmaceutical composition of the present invention to a subject that is not at the time of administration afflicted with the disease. "Preventing" also refers to preventing the recurrence of a disease or of one or more symptoms associated with such disease. "Treatment" and "therapeutically," refer to the act of treating, as "treating" is defined above.
[0092] "Variant" is used herein to describe a peptide or polypeptide that differs in amino acid sequence by the insertion, deletion, or conservative substitution of amino acids, but retains at least one biological activity. Representative examples of "biological activity" include the ability to be bound by a specific antibody or to promote an immune response. Variant is also used herein to describe a protein with an amino acid sequence that is substantially identical to a referenced protein with an amino acid sequence that retains at least one biological activity. A conservative substitution of an amino acid, i.e., replacing an amino acid with a different amino acid of similar properties (e.g., hydrophilicity, degree, and distribution of charged regions) is recognized in the art as typically involving a minor change. These minor changes can be identified, in part, by considering the hydropathic index of amino acids, as understood in the art. Kyte et al., J. Mol. Biol. 157:105-132 (1982). The hydropathic index of an amino acid is based on a consideration of its hydrophobicity and charge. It is known in the art that amino acids of similar hydropathic indexes can be substituted and still retain protein function. In one aspect, amino acids having hydropathic indexes of .+-.2 are substituted. The hydrophilicity of amino acids can also be used to reveal substitutions that would result in proteins retaining biological function. A consideration of the hydrophilicity of amino acids in the context of a peptide permits calculation of the greatest local average hydrophilicity of that peptide, a useful measure that has been reported to correlate well with antigenicity and immunogenicity. U.S. Pat. No. 4,554,101, incorporated fully herein by reference. Substitution of amino acids having similar hydrophilicity values can result in peptides retaining biological activity, for example immunogenicity, as is understood in the art. Substitutions may be performed with amino acids having hydrophilicity values within .+-.2 of each other. Both the hydrophobicity index and the hydrophilicity value of amino acids are influenced by the particular side chain of that amino acid. Consistent with that observation, amino acid substitutions that are compatible with biological function are understood to depend on the relative similarity of the amino acids, and particularly the side chains of those amino acids, as revealed by the hydrophobicity, hydrophilicity, charge, size, and other properties. "Variant" also can be used to refer to an antigenically reactive fragment of an anti-CFH antibody that differs from the corresponding fragment of anti-CFH antibody in amino acid sequence but is still antigenically reactive and can compete with the corresponding fragment of anti-CFH antibody for binding with CFH. "Variant" also can be used to describe a polypeptide or a fragment thereof that has been differentially processed, such as by proteolysis, phosphorylation, or other post-translational modification, yet retains its antigen reactivity.
[0093] Unless otherwise defined herein, scientific and technical terms used in connection with the present disclosure shall have the meanings that are commonly understood by those of ordinary skill in the art. For example, any nomenclatures used in connection with, and techniques of, cell and tissue culture, molecular biology, immunology, microbiology, genetics and protein and nucleic acid chemistry and hybridization described herein are those that are well known and commonly used in the art. The meaning and scope of the terms should be clear; in the event, however of any latent ambiguity, definitions provided herein take precedent over any dictionary or extrinsic definition. Further, unless otherwise required by context, singular terms shall include pluralities and plural terms shall include the singular.
2. CFH-RECOGNIZING ANTIBODIES
[0094] Provided herein are antibodies that bind CFH and can be used in methods of treating cancer, such as lung cancer. Complement factor H is one of a class of complement inhibitory factors that protect both normal and tumor cells from attack and destruction by the alternative complement pathway by inactivating C3b, a protein that is essential for formation of a cell lytic complex on a cell surface. The primary function of CFH is to inhibit the alternative pathway of complement-mediated lysis. CFH prevents the deposition of complement protein C3b on the cell surface by (a) acting as a cofactor for complement factor I (CFI), a protease that cleaves C3b, and (b) preventing the formation of and accelerating the decay of the enzyme that forms C3b from its precursor, C3. Deposition of C3b initiates the formation of the cell-lytic membrane attack complex, leading to cell lysis; thus, control of the deposition of C3b on the cell surface by CFH protects against cell lysis. CFH engages with C3b (or degraded C3b, named C3d) on mammalian cell surfaces that contain glycosaminoglycans and sialic acid, as opposed to bacterial surfaces lacking these groups, thus mediating target discrimination. Besides protecting normal host cells, CFH has been shown to protect tumor cells, including those from NSCLC, glioblastoma, and colon cancer cells, from complement attack.
[0095] a. CFH
[0096] Human CFH may have the following amino acid sequence:
TABLE-US-00001 (SEQ ID NO: 81 MRLLAKIICLMLWAICVAEDCNELPPRRNTEILTGSWSDQTYPEGTQAIY KCRPGYRSLGNVIMVCRKGEWVALNPLRKCQKRPCGHPGDTPFGTFTLTG GNVFEYGVKAVYTCNEGYQLLGEINYRECDTDGWTNDIPICEVVKCLPVT APENGKIVSSAMEPDREYHFGQAVRFVCNSGYKIEGDEEMHCSDDGFWSK EKPKCVEISCKSPDVINGSPISQKIIYKENERFQYKCNMGYEYSERGDAV CTESGWRPLPSCEEKSCDNPYIPNGDYSPLRIKHRTGDEITYQCRNGFYP ATRGNTAKCTSTGWIPAPRCTLKPCDYPDIKHGGLYHENMRRPYFPVAVG KYYSYYCDEHFETPSGSYWDHIFICTQDGWSPAVPCLRKCYFPYLENGYN QNYGRKFVQGKSIDVACHPGYALPKAQTTVTCMENGWSPTPRCIRVKTCS KSSIDIENGFISESQYTYALKEKAKYQCKLGYVTADGETSGSITCGKDGW SAQPTCIKSCDIPVFMNARTKNDFTWFKLNDTLDYECHDGYESNTGSTTG SIVCGYNGWSDLPICYERECELPKIDVHLVPDRKKDQYKVGEVLKFSCKP GFTIVGPNSVQCYHFGLSPDLPICKEQVQScGPPPELLNGNVKEKTKEEY GHSEVVEYYCNPRFLMKGPNKIQCVDGEWTTLPVCIVEESTCGDIPELEH GWAQLSSPPYYYGDSVEFNCSESFTMIGHRSITCIHGVWTQLPQCVAIDK LKKCKSSNLIILEEHLKNKKEFDHNSNIRYRCRGKEGWIHTVCINGRWDP EVNCSMAQIQLCPPPPQIPNSHNMTTTLNYRDGEKVSVLCQENYLIQEGE EITCKDGRWQSIPLCVEKIPCSQPPQIEHGTINSSRSSQESYAHGTKLSY TCEGGFRISEENETTCYMGKWSSPPQCEGLPCKSPPEISHGVVAHMSDSY QYGEEVTYKCFEGFGIDGPAIAKCLGEKWSEIPPSCIKTDCLSLPSFENA IPMGEKKDVYKAGEQVTYTCATYYKMDGASNVTCINSRWTGRPTCRDTSC VNPPTVQNAYIVSRQMSKYPSGERVRYQCRSPYEMFGDEEVMCLNGNWTE PPQCKDSTGKCGPPPPIDNGDITSFPLSVYAPASSVEYQCQNLYQLEGNK RITCRNGQWSEPPKCLHPCVISREIMENYNIALRWTAKQKLYSRTGESVE FVCKRGYRLSSRSHTLRTTCWDGKLEYPTCAKR;
UniProt P08603). The CFH may be reduced or not reduced.
[0097] The human CFH may be a fragment or variant of SEQ ID NO:81. The fragment or variant may be reduced or not reduced form. The fragment of CFH may be between 5 and 1230 amino acids, between 10 and 1000 amino acids, between 10 and 750 amino acids, between 10 and 500 amino acids, between 50 and 400 amino acids, between 60 and 400 amino acids, between 65 and 400 amino acids, between 100 and 400 amino acids, between 150 and 400 amino acids, between 100 and 300 amino acids, or between 200 and 300 amino acids in length. The fragment may comprise a contiguous number of amino acids from SEQ ID NO:81.
[0098] The fragment of human CFH may have the following amino acid sequence: GKCGPPPPIDNGDITSFPLSVYAPASSVEYQCQNLYQLEGNKRITCRNGQWSEPPKCLH (SEQ ID NO:82), which correspond to amino acids 1107-1165 of SEQ ID NO:81. The fragment of human CFH may have the following amino acid sequence GPPPPIDNGDITSFP (SEQ ID NO:84).
[0099] The reduced form of CFH may reveal a cryptic epitope or cryptic target region. This epitope or target region may be revealed only on the surface of tumor cells. CFH may be an autoantigen due to the presentation of the cryptic epitope in the tumor microenvironment. NSCLC tumors exhibit elevated levels of thioredoxin, the disulfide reductase macrophage migration inhibitory factor, and non-protein thiols such as reduced cysteine and glutathione. These factors contribute to the production of a more reducing environment in the tumor than in normal tissues. Thus, the anti-CFH epitope(s) may be hidden and only exposed upon reduction of the protein in the intratumoral space. Alternatively, once the soluble form of CFH binds to the tumor cell, the protein may unfold and bind in a tumor cell-specific conformation so that it becomes antigenic; reduction in vitro may simply put CFH in a conformation that mimics this state.
[0100] b. CFH Antibodies
[0101] The antibody is an antibody that binds to the reduced form of CFH, a fragment thereof, an epitope of CFH, or a variant thereof. The antibody may be a fragment of the anti-CFH antibody or a variant or a derivative thereof. The antibody may be a polyclonal or monoclonal antibody. The antibody may be a chimeric antibody, a single chain antibody, an affinity matured antibody, a human antibody, a humanized antibody, a fully human antibody or an antibody fragment, such as a Fab fragment, or a mixture thereof. Antibody fragments or derivatives may comprise F(ab').sub.2, Fv or scFv fragments. The antibody derivatives can be produced by peptidomimetics. The anti-CFH antibodies may be human-derived antibodies. Further, techniques described or the production of single chain antibodies can be adapted to produce single chain antibodies. The antibody may or may not be generated from a human in vivo immune response. For example, the antibody may or may not be an autoantibody.
[0102] The anti-CFH antibodies may be a chimeric anti-CFH or humanized anti-CFH antibody. In one embodiment, both the humanized antibody and chimeric antibody are monovalent. In one embodiment, both the humanized antibody and chimeric antibody comprise a single Fab region linked to an Fc region.
[0103] Human antibodies may be derived from phage-display technology or from transgenic mice that express human immunoglobulin genes. The antibody may be generated as a result of a human in vivo immune response and isolated. See, for example, Funaro et al., BMC Biotechnology, 2008(8):85. Therefore, the antibody may be a product of the human and not animal repertoire. Because it is of human origin, the risks of reactivity against self-antigens may be minimized. Alternatively, standard yeast display libraries and display technologies may be used to select and isolate human anti-CFH antibodies. For example, libraries of naive human single chain variable fragments (scFv) may be used to select human anti-CFH antibodies. Transgenic animals may be used to express human antibodies.
[0104] Humanized antibodies may be antibody molecules from non-human species antibody that binds the desired antigen having one or more complementarity determining regions (CDRs) from the non-human species and framework regions from a human immunoglobulin molecule.
[0105] The antibody is distinguishable from known antibodies in that it possesses different biological function(s) than those known in the art.
[0106] (1) Epitope
[0107] The epitope for the CFH antibody, which may be cryptic, may be SCR 19 (SEQ ID NO: 82), which is involved with CFH function. SCR19 contains binding sites for C3b/C3d and polyanions typical of self- or auto-surfaces. SCR19 is a domain that is involved in the host cell-protective function of CFH as it is involved in binding to the C3d portion of C3b. The target region may be an epitope of PIDNGDIT (SEQ ID NO:83), which resides in SCR19. The target region may include the D1119 residue of SEQ ID NO:81, which is also residue 6 of SEQ ID NO:83. The target region may include the 15-mer peptide of GPPPPIDNGDITSFP (SEQ ID NO:84)
[0108] A CFH antibody may immunospecifically bind to a reduced or non-reduced form of any one or more of epitopes within SEQ ID NOs:81-84, a fragment thereof, or a variant thereof. The antibody may immunospecifically recognize and bind at least three amino acids, at least four amino acids, at least five amino acids, at least six amino acids, or at least seven amino acids within the epitope peptide of PIDNGDIT (SEQ ID NO:83) or GPPPPIDNGDITSFP (SEQ ID NO:84). The antibody may immunospecifically recognize and bind to an epitope that has at least three contiguous amino acids, at least four contiguous amino acids, at least five contiguous amino acids, at least six contiguous amino acids, at least seven contiguous amino acids, at least eight contiguous amino acids, at least nine contiguous amino acids, or at least ten contiguous amino acids of SEQ ID NO: 81 or 82. The contiguous amino acids may include amino acid D1119 of SEQ ID NO:81.
[0109] (2) Antibody Binding Characteristics
[0110] The antibody may immunospecifically bind to CFH (SEQ ID NO:81), SCR19 (SEQ ID NO:82), the amino acid sequence of PIDNGDIT (SEQ ID NO:83), the amino acid sequence of GPPPPIDNGDITSFP (SEQ ID NO:84), a fragment thereof, or a variant thereof and have an off-rate (kd) of about 1.0.times.10.sup.-4 s.sup.-1 or less, about 1.0.times.10.sup.-5 s.sup.-1 or less, about 5.0.times.10.sup.-6 s.sup.-1 or less, about 1.0.times.10.sup.-6 s.sup.-1 or less, about 5.0.times.10.sup.-7 s.sup.-1 or less, about 1.0.times.10.sup.-7 s.sup.-1 or less, about 5.0.times.10.sup.-8 s.sup.-1 or less, about 1.0.times.10.sup.-8 s.sup.-1 or less, about 1.0.times.10.sup.-9 s.sup.-1 or less, about 1.0.times.10.sup.-10 s.sup.-1 or less, about 1.0.times.10.sup.-11 s.sup.-1 or less, about 1.0.times.10.sup.-12 s.sup.-1 or less, or has a kd ranging from about 1.0.times.10.sup.-12 s.sup.-1 to about 1.0.times.10.sup.-4 s.sup.-1, about 1.0.times.10.sup.-12 s.sup.-1 to about 1.0.times.10.sup.-5 s.sup.-1, about 1.0.times.10.sup.-12 s.sup.-1 to about 1.0.times.10.sup.-6 s.sup.-1, about 1.0.times.10.sup.-2 s.sup.-1 to about 1.0.times.10.sup.-7 s.sup.-1, about 1.0.times.10.sup.-12 s.sup.-1 to about 1.0.times.10.sup.-8 s.sup.-1, about 1.0.times.10.sup.-12 s.sup.-1 to about 1.0.times.10.sup.-9 s.sup.-1, about 1.0.times.10.sup.-12 s.sup.-1 to about 1.0.times.10.sup.-10 s.sup.-1, about 1.0.times.10.sup.-10 s.sup.-1 to about 1.0.times.10.sup.-4 s.sup.-1, about 1.0.times.10.sup.-10 s.sup.-1 to about 1.0.times.10.sup.-5 s.sup.-1, about 1.0.times.10.sup.-10 s.sup.-1 to about 1.0.times.10.sup.-6 s.sup.-1, about 1.0.times.10.sup.-10 s.sup.-1 to about 1.0.times.10.sup.-7 s.sup.-1, about 1.0.times.10.sup.-10 s.sup.-1 to about 1.0.times.10.sup.-8 s.sup.-1, about 1.0.times.10.sup.-8 s.sup.-1 to about 1.0.times.10.sup.-4 s.sup.-1s, about 1.0.times.10.sup.-8 s.sup.-1 to about 1.0.times.10.sup.-5 s.sup.-1s, about 1.0.times.10.sup.-8 s.sup.-1s to about 1.0.times.10.sup.-6 s.sup.-1, about 1.0.times.10.sup.-8 s.sup.-1 to about 1.0.times.10.sup.-7 s.sup.-1, about 1.0.times.10.sup.-7 s.sup.-1 to about 1.0.times.10.sup.-4 s.sup.-1, about 1.0.times.10.sup.-7 s.sup.-1 to about 1.0.times.10.sup.-5 s.sup.-1, or about 1.0.times.10.sup.-7 s.sup.-1 to about 1.0.times.10.sup.-6 s.sup.-1. The fragment may be SEQ ID NO:84 or SEQ ID NO: 122.
[0111] The antibody may immunospecifically bind to CFH (SEQ ID NO:81), SCR19 (SEQ ID NO:82), the amino acid sequence of PIDNGDIT (SEQ ID NO:83), the amino acid sequence of GPPPPIDNGDITSFP (SEQ ID NO:84), a fragment thereof, or a variant thereof and have an on-rate (ka) of at least about 1.0.times.10.sup.3 M.sup.-1s.sup.-1, at least about 1.0.times.10.sup.4 M.sup.-1s.sup.-1, at least about 5.0.times.10.sup.4 M.sup.-1s.sup.-1, at least about 1.0.times.10.sup.5 M.sup.-1s.sup.-1, at least about 2.0.times.10.sup.5 M.sup.-1s.sup.-1, at least about 3.0.times.10.sup.5 M.sup.-1s.sup.-1, at least about 4.0.times.10.sup.5 M.sup.-1s.sup.-1, at least about 5.0.times.10.sup.5 M.sup.-1s.sup.-1, least about 6.0.times.10.sup.5 M.sup.-1s.sup.-1, at least about 1.0.times.10.sup.6 M.sup.-1s.sup.-1, at least about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, at least about 1.0.times.10.sup.8 M.sup.-1s.sup.-1, or has a ka ranging from about 1.0.times.10.sup.3 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.8 M.sup.-1s.sup.-1, about 1.0.times.10.sup.4 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.8 M.sup.-1s.sup.-1, about 1.0.times.10.sup.5 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.8 M.sup.-1s.sup.-1, about 1.0.times.10.sup.6 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.8 M.sup.-1s.sup.-1, about 1.0.times.10.sup.7 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.8 M.sup.-1s.sup.-1, about 1.0.times.10.sup.3 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, about 1.0.times.10.sup.4 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, about 1.0.times.10.sup.5 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, about 1.0.times.10.sup.6 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, about 1.0.times.10.sup.4 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, about 1.0.times.10.sup.4 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.6 M.sup.-1s.sup.-1, about 1.0.times.10.sup.4 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.5 M.sup.-1s.sup.-1, about 1.0.times.10.sup.5 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.7 M.sup.-1s.sup.-1, or about 1.0.times.10.sup.5 M.sup.-1s.sup.-1 to about 1.0.times.10.sup.6 M.sup.-1s.sup.-1. The fragment may be SEQ ID NO:84 or SEQ ID NO: 122.
[0112] The antibody may immunospecifically bind to CFH (SEQ ID NO:81), SCR19 (SEQ ID NO:82), the amino acid sequence of PIDNGDIT (SEQ ID NO:83), the amino acid sequence of GPPPPIDNGDITSFP (SEQ ID NO:84), a fragment thereof, or a variant thereof and have an affinity (K.sub.D) of at least about 1.0.times.10.sup.-15 M, at least about 1.0.times.10.sup.-14 M, at least about 1.0.times.10.sup.-13 M, at least about 1.5.times.10.sup.-13 M, at least about 1.0.times.10.sup.-12 M, at least about 1.6.times.10.sup.-12 M, at least about 1.7.times.10.sup.-12 M, at least about 1.8.times.10.sup.-12 M, at least about 1.9.times.10.sup.-12 M, at least about 2.0.times.10.sup.-12 M, at least about 2.1.times.10.sup.-12 M, at least about 2.2.times.10.sup.-12 M, at least about 2.3.times.10.sup.-12 M, at least about 2.4.times.10.sup.-12 M, at least about 2.5.times.10.sup.-12 M, at least about 2.6.times.10.sup.-12 M, at least about 2.7.times.10.sup.-12 M, at least about 2.8.times.10.sup.-12 M, at least about 2.9.times.10.sup.-12 M, at least about 3.0.times.10.sup.-12 M, at least about 5.0.times.10.sup.-12 M, at about least 1.0.times.10.sup.-11 M, at least about 1.5.times.10.sup.-11 M, at least about 5.0.times.10.sup.-11 M, at least about 1.0.times.10.sup.-10 M, at least about 5.0.times.10.sup.-10 M, at least about 1.0.times.10.sup.-9 M, or has a K.sub.D ranging from about 1.0.times.10.sup.-15 M to about 1.0.times.10.sup.-9 M, about 1.0.times.10.sup.-15 M to about 1.0.times.10.sup.-10 M about 1.0.times.10.sup.-15 M to about 1.0.times.10.sup.-11 M, about 1.0.times.10.sup.-15 M to about 1.0.times.10.sup.-12 M, about 1.0.times.10.sup.-15 M to about 1.0.times.10.sup.-13 M, about 1.0.times.10.sup.-14 M to about 1.0.times.10.sup.-9 M, about 1.0.times.10.sup.-14 M to about 1.0.times.10.sup.-10 M, about 1.0.times.10.sup.-14 M to about 1.0.times.10.sup.-11 M, about 1.0.times.10.sup.-14 M to about 1.0.times.10.sup.-12 M, about 1.0.times.10.sup.-14 M to about 1.0.times.10.sup.-13 M, about 1.0.times.10.sup.-13 M to about 1.0.times.10.sup.-9 M, about 1.0.times.10.sup.-13 M to about 1.0.times.10.sup.-10 M, about 1.0.times.10.sup.-13 M to about 1.0.times.10.sup.-11 M, about 1.0.times.10.sup.-13 M to about 1.0.times.10.sup.-12 M, about 1.0.times.10.sup.-12 M to about 1.0.times.10.sup.-9 M, about 1.0.times.10.sup.-12 M to about 1.0.times.10.sup.-10 M or about 1.0.times.10.sup.-12 M to about 1.0.times.10.sup.-11 M. The fragment may be SEQ ID NO:84 or SEQ ID NO: 122.
[0113] The binding of the antibody to CFH may be sensitive to the reduced form of CFH. An antibody that is sensitive to the reduced form of CFH means that the antibody's binding affinity to CFH changes depending on whether the CFH is in the reduced form or not reduced form. For example, an antibody whose binding is sensitive to the CFH being in the reduced form or not reduced form may have lower binding affinity to CFH if the CFH is not in the reduced form. Alternatively, an antibody whose binding is sensitive to the CFH being in the reduced form or not reduced form may have lower binding affinity to CFH if the CFH is in the reduced form. An antibody that is insensitive to the CFH being in the reduced form or not reduced form means that the antibody's binding affinity to CFH does not change if the CFH is in the reduced form or not reduced form.
[0114] (1) Antibody Structure
[0115] (a) Heavy Chain and Light Chain CDRs
[0116] The antibody may immunospecifically bind to reduced CFH, a fragment thereof, or a variant thereof and comprise a variable heavy chain (VH) and/or variable light chain (VL) shown in Table 1. The antibody may immunospecifically bind to reduced CFH, a fragment thereof, or a variant thereof, and comprises one or more of the heavy chain or light chain CDR sequences also shown in Table 1. The light chain of the antibody may be a kappa chain or a lambda chain. For example, see Table 1.
TABLE-US-00002 TABLE 1 SEQ ID Antibody NO: Sequence Ab7955 (VH) 1 LVESGGGVVRPGRSLRLSCVASG FTFNAYGMHWVRQGPGKGLEWLA VISYEGKTVYYADSVKDRFTISR DNSRNTVSLHLNNLRGEDTAVYY CAKGSASAAVLQHWGQGTLVSVT S Ab7955 (VH) 11 FTFNAYGMH CDR-H1 Ab7955 (VH) 21 WLAVISYEGKTVYY CDR-H2 Ab7955 (VH) 31 KGSASAAVLQH CDR-H3 Ab7955 (VL) 41 MTQSPGSLAVSLGSRATINCKSS RSLLYRSNNKNYLAWYQQKPGQS PRLLIYWASSRESGVPDRFSGGG SGTSFTLTISSLQAEDVAVYYCQ QYFNPPWTFGQGTKVEIK Ab7955 (VL) 51 RSLLYRSNNKNYLA CDR-L1 Ab7955 (VL) 61 LLIYWASSRES CDR-L2 Ab7955 (VL) 71 QQYFNPPW CDR-L3 Ab7957/293i 2 LVESGGGVVPPGKSLRLSCAASG (VH) FTFSLYGIHWVRQAPGKGLEWVA VISYDGNTKYYTDSVKGRFTISR DNAKNTIYLQMNSLRLDDTAVYY CAKGAANSATFDFWGRGTMVTVS S Ab7957/293i 12 FTFSLYGIH (VH) CDR- H1 Ab7957/293i 22 WVAVISYDGNTKYYT (VH) CDR- H2 Ab7957/293i 32 KGAANSATFDF (VH) CDR- H3 Ab7957/293i 42 MTQSPDSLTLSLGERATINCRSS (VL) RTVLYRSNNKNYLAWYQHKPGQP PKLLMSWASTRETGVPDRFSGSG SGTHFTLTITSLQPEDVAVYYCQ QYYSPPWTFGQGTKVEIR Ab7957/293i 52 RTVLYRSNNKNYLA (VL) CDR-L1 Ab7957/293i 62 LLMSWASTRE (VL) CDR-L2 Ab7957/293i 72 QQYYSPPW (VL) CDR-L3 Ab7960/293i 3 LVESGGGVVQPGKSLRLSCVASG (VH) FSFSTYGMHWVRQAPGKGLEWVA VMSFDGKTKYYADSVKGRFTISR DNPKNTLYLQMNSLRSEDTAVYY CAKGGAAAAVFDSWGPGILLTVS S Ab7960/293i 13 FSFSTYGMH (VH) CDR- H1 Ab7960/293i 23 WVAVMSFDGKTKYY (VH) CDR- H2 Ab7960/293i 33 KGGAAAAVFDS (VH) CDR- H3 Ab7960/293i 43 MTQSPNSLAVSLGGRATINCKAS (VL) QSILYRSNNKNYLAWYQHKAGQP PKLLIYWASTRESGVPERFSGSG SRTDFTLTINGLQAEDVAVYYCQ QYYNPPWTFGQGTKVEIK Ab7960/293i 53 QSILYRSNNKNYLA (VL) CDR-L1 Ab7960/293i 63 LLIYWASTRES (VL) CDR-L2 Ab7960/293i 73 QQYYNPPW (VL) CDR-L3 Ab7964 (VH) 4 LVESGGGVVQPGRSLRLSCAASG VTFSRYGMHWVRQAPGKGLEWVA VISYDEKTKYYADSVKGRFTISR DNSKNTLFLHMNRLRYEDTAVYY CAKGASSGAYFDYWGQGTLVTVS S Ab7964 (VH) 14 VTFSRYGMH CDR-H1 Ab7964 (VH) 24 WVAVISYDEKTKYY CDR-H2 Ab7964 (VH) 34 KGASSGAYFDY CDR-H3 Ab7964 (VL) 44 MTQSPDSLAVSLGERATIKCKSS QSVLYSSNNKNYLAWYQHKPGQP PKVLVYWASTRESGVPDRFSGSG SGTDFTLTISSLQAEDVAVYYCQ QYYNPPWTFGQGTKVAIK Ab7964 (VL) 54 QSVLYSSNNKNYLA CDR-L1 Ab7964 (VL) 64 VLVYWASTRES CDR-L2 Ab7964 (VL) 74 QQYYNPPW CDR-L3 Ab7979 (VH) 5 LVESGGGVVQPGKSLRLSCVASG FTFSTYGMHWVRQAPGKGLEWVA VMSFDGKTKYYADSVKGRFTISR DNPKNTLYLQMNSLRSEDTAVYY CAKGGAAAAVFDSWGQGILLTVS S Ab7979 (VH) 15 FTFSTYGMH CDR-H1 Ab7979 (VH) 25 WVAVMSFDGKTKYY CDR-H2 Ab7979 (VH) 35 KGGAAAAVFDS CDR-H3 Ab7979 (VL) 45 MTQSPNSLAVSLGGRATINCKTS QSILYRSNNKNYLAWYQHKPGQP PKLLIYWASTRESRVPDRFSGSG SRTDFTLTISGLQAEDVAVYYCQ QYYNPPWTFGQGTKVEIK Ab7979 (VL) 55 QSILYRSNNKNYLA CDR-L1 Ab7979 (VL) 65 LLIYWASTRESR CDR-L2 Ab7979 (VL) 75 QQYYNPPW CDR-L3 Ab7961/293i 6 LVESGGGVVQPGKSLRLSCVASG (VH) FTFSSYGMHWVRQAPGKGLEWVA VMSFDGKTKYYADSVKGRFTISR DNPKNTLYLQMNSLRSEDTAVYY CAKGGAAAAVFDSWGQGILLTVS S Ab7961/293i 16 FTFSSYGMH (VH) CDR- H1 Ab7961/293i 26 WVAVMSFDGKTKYY (VH) CDR- H2 Ab7961/293i 36 KGGAAAAVFDS (VH) CDR- H3 Ab7961/293i 46 MTQSPNSLAVSLGGRATINCKTS (VL) QSILYRSNNKNYLAWYQHKSGQP PKLLIYWASTRESGVPDRFSGSG SRTDFTLTISGLQAEDVAVYYCQ QYYNPPWTFGQGTKVEIK Ab7961/293i 56 QSILYRSNNKNYLA (VL) CDR-L1 Ab7961/293i 66 LLIYWASTRES (VL) CDR-L2 Ab7961/293i 76 QQYYNPPW (VL) CDR-L3 Ab7962/293i 7 LVESGGGLVQPGGSLRLSCAASG (VH) FTFSSYEMNWVRQAPGKGLEWVS YISSSGSTIYYADSVKGRFTISR DNAKNSLYLQMNSLRAEDTAVYY CARVEQLAPSPYMDVWGKGTTVT VSS Ab7962/293i 17 FTFSSYEMN (VH) CDR- H1 Ab7962/293i 27 WVSYISSSGSTIYY (VH) CDR- H2 Ab7962/293i 37 RVEQLAPSPYMDV (VH) CDR- H3 Ab7962/293i 47 MTQSLDSLTVSLGERATINCKSS (VL) QSLLYTSNNKNYLAWYQQKSGQP PKLLIYWASIRDSGVPDRFSGSG SATDFTLTINNLQAEDVAVYFCQ QYYKTPLTFGGGTKVEVR Ab7962/293i 57 QSLLYTSNNKNYLA (VL) CDR-L1 Ab7962/293i 67 LLIYWASIRDS (VL) CDR-L2 Ab7962/293i 77 QQYYKTPL (VL) CDR-L3 Ab7968 (VH) 8 LVESGGGVVQPGRSLRLSCAASG FTFSRYGMHWVRQAPGKGLEWVA VMSYDGSTKYYADSVKGRFAISR DNPKNTLFLQMNSLRPDDTAVYY CAKGGAAAAVMDVWGKGTTVTVS S Ab7968 (VH) 18 FTFSRYGMH CDR-H1 Ab7968 (VH) 28 WVAVMSYDGSTKYY CDR-H2 Ab7968 (VH) 38 KGGAAAAVMDV
CDR-H3 Ab7968 (VL) 48 MTQSPDSLTVSLGERATISCKSS QRLLYSSNNKNYLAWYQQKPGQP PKLLMYWASTRESGVPDRFSGSG SGTDFSLTISSLQAEDVAVYYCQ QYYNPPWTFGQGTKVEVK Ab7968 (VL) 58 QRLLYSSNNKNYLA CDR-L1 Ab7968 (VL) 68 LLMYWASTRES CDR-L2 Ab7968 (VL) 78 QQYYNPPW CDR-L3 Ab7966 (VH) 9 QVQLVQSGAEVKKPGESLKISCK GSGYSFTSYWIGWVRQMPGKGLE WMGIIYPGDSDTRYSPSFQGQVT ISADKSISTAYLQWSSLKASDTA MYYCARRGLRGAYYYYYGMDVWG QGTTVTVSS Ab7966 (VH) 19 CKGSGYSFTSYWIG CDR-H1 Ab7966 (VH) 29 WMGIIYPGDSDTRY CDR-H2 Ab7966 (VH) 39 RRGLRGAYYYYYGMDV CDR-H3 Ab7966 (VL) 49 DIQXTQSPSTLSASVGDRVTITC RASQSISSWLAWYQQKPGKAPKL LIYKASSLESGVPSRFSGSGSGT EFTLTISSLQPDDFATYYCQQYN SYSWTFGQGTKVEIK Ab7966 (VL) 59 QSISSWLA CDR-L1 Ab7966 (VL) 69 LLIYKASSLES CDR-L2 Ab7966 (VL) 79 QQYNSYSW CDR-L3 Ab7970 (VH) 10 LVESGGGVVQPGRSLRLSCAASG LTFSFYNFHWVRQTPGKGLEWVA GISYDATRTNYAGSVTGRFTISR DNSKKMLYLQMSSLGPQDTAVYH CARDRSDGQLHKVAFDSWGQGAL VTVSS Ab7970 (VH) 20 LTFSFYNFH CDR-H1 Ab7970 (VH) 30 WVAGISYDATRTNY CDR-H2 Ab7970 (VH) 40 RDRSDGQLHKVAFDS CDR-H3 Ab7970 (VL) 50 MTQSPDSLTLSLGERATINCRSS RTVLYRSNNKNYLAWYQHKPGQP PKLLMSWASTRETGVPDRFSGSG SGTHFTLTITSLQPEDVAVYYCQ QYYSPPWTFGQGTKVEIR Ab7970 (VL) 60 RTVLYRSNNKNYLA CDR-L1 Ab7970 (VL) 70 LLMSWASTRET CDR-L2 Ab7970 (VL) 80 QQYYSPPW CDR-L3
[0117] The antibody or variant or derivative thereof may contain one or more amino acid sequences that are greater than 95%, 90%, 85%, 80%, 75%, 70%, 65%, 60%, 55%, or 50% identical to one or more of SEQ ID NOs:1-80. The antibody or variant or derivative thereof may be encoded by one or more nucleic acid sequences that are greater than 95%, 90%, 85%, 80%, 75%, 70%, 65%, 60%, 55%, or 50% identical to one or more of SEQ ID NOs:85-118. Polypeptide identity and homology can be determined, for example, by the algorithm described in the report: Wilbur, W. J. and Lipman, D. J. Proc. Natl. Acad. Sci. USA 80, 726-730 (1983). The herein described antibody, variant, or derivative thereof may be encoded by a nucleic acid that hybridizes under stringent conditions with the complement of one or more of SEQ ID NOs: 17-32. The herein described antibody, variant, or derivative thereof may be encoded by a nucleic acid that hybridizes under highly stringent conditions with the complement of one or more nucleic acids that encode one or more of SEQ ID NOs:1-80.
[0118] The antibody may be an IgG, IgE, IgM, IgD, IgA, and IgY molecule class (for example, IgG1, IgG2, IgG3, IgG4, IgA1 and IgA2) or subclass.
[0119] c. Nucleotide Sequences
[0120] Provided herein is an isolated nucleic acid encoding an antibody that immunospecifically binds to CFH, a fragment thereof, or a variant thereof. The isolated nucleic acid may comprise a nucleotide sequence that hybridizes, under stringent conditions, to the nucleic acid molecule that encodes an antibody comprising a heavy chain or light chain sequence, as shown in Table 1. The isolated nucleic acid may comprise a nucleotide sequence, as shown in Table 2.
TABLE-US-00003 TABLE 2 SEQ Nucleo- ID tide NO. Sequence H007970 85 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGGAGGTCCCTGAGACTCTCCTGCGCAGCCTC TGGACTCACCTTCAGTTTCTATAATTTCCACTGGGTC CGCCAGACTCCAGGCAAGGGGCTGGAGTGGGTGGCAG GCATCTCATACGATGCAACCAGGACGAACTACGCAGG CTCGTCACGGGCCGATTCACCATTTCCAGAGACAATT CCAAGAAAATGCTGTATCTGCAAATGAGCAGCCTGGG ACCTCAAGACACGGCTGTATATCATTGTGCGAGAGAT CGTTCTGACGGGCAACTGCATAAAGTGGCTTTTGACT CCTGGGGCCAGGGAGCCCTGGTCACCGTCTCATCA H007955 86 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) GGCCTGGGCGGTCCCTGAGACTCTCCTGTGTTGCCTC TGGTTTCACCTTCAATGCTTATGGCATGCATTGGGTC CGCCAGGGTCCAGGCAAGGGCCTTGAGTGGCTGGCGG TCATTTCATATGAAGGAAAGACTGTTTATTATGCAGA TTCCGTTAAGGACCGTTTCACCATCTCCAGAGACAAT TCCAGGAACACGGTGTCTCTACATCTGAACAACCTGA GAGGTGAGGACACGGCTGTCTATTACTGTGCGAAGGG GTCGGCTTCAGCAGCAGTCCTCCAACACTGGGGTCAG GGCACCCTGGTCAGCGTCACGTCA H007957 87 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) CGCCTGGGAAGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTCTCTATGGCATACACTGGGTC CGCCAGGCTCCCGGCAAGGGACTGGAGTGGGTGGCAG TTATCTCATATGATGGAAATACTAAATACTATACAGA CTCTGTAAAGGGTCGATTCACCATCTCCAGAGACAAT GCCAAGAACACAATTTATCTGCAAATGAACAGTCTAA GACTTGACGACACGGCTGTTTATTACTGTGCGAAAGG AGCGGCGAATAGCGCTACTTTTGATTTCTGGGGCCGA GGGACAATGGTCACCGTCTCTTCA H007958 88 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) CGCCTGGGAAGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTCTCTATGGCATACACTGGGTC CGCCAGGCTCCCGGCAAGGGACTGGAGTGGGTGGCAG TTATCTCATATGATGGAAATACTAAATACTATACAGA CTCTGTAAAGGGTCGATTCACCATCTCCAGAGACAAT GCCAAGAACACAATTTATCTGCAAATGAACAGTCTAA GACTTGACGACACGGCTGTTTATTACTGTGCGAAAGG AGCGGCGAATAGCGCTACTTTTGATTTCTGGGGCCGA GGGACAATGGTCACCGTCTCTTCA H007963 89 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) CGCCTGGGAAGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTCTCTATGGCATACACTGGGTC CGCCAGGCTCCCGGCAAGGGACTGGAGTGGGTGGCAG TTATCTCATATGATGGAAATACTAAATACTATACAGA CTCTGTAAAGGGTCGATTCACCATCTCCAGAGACAAT GCCAAGAACACAATTTATCTGCAAATGAACAGTCTAA GACTTGACGACACGGCTGTTTATTACTGTGCGAAAGG AGCGGCGAATAGCGCTACTTTTGATTTCTGGGGCCGA GGGACAATGGTCACCGTCTCTTCA H007982 90 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) CGCCTGGGAAGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTCTCTATGGCATACACTGGGTC CGCCAGGCTCCCGGCAAGGGACTGGAGTGGGTGGCAG TTATCTCATATGATGGAAATACTAAATACTATACAGA CTCTGTAAAGGGTCGATTCACCATCTCCAGAGACAAT GCCAAGAACACAATTTATCTGCAAATGAACAGTCTAA GACTTGACGACACGGCTGTTTATTACTGTGCGAAAGG AGCGGCGAATAGCGCTACTTTTGATTTCTGGGGCCGA GGGACAATGGTCACCGTCTCTTCA H007960 91 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGAAAGTCTCTGAGACTCTCCTGTGTAGCCTC TGGATTCAGCTTCAGTACTTATGGCATGCACTGGGTC CGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCGG TTATGTCATTTGATGGAAAGACTAAATACTATGCAGA CTCCGTGAAGGGCCGATTCACCATCTCCAGAGACAAT CCCAAGAACACACTATATCTGCAA H007967 92 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGAAAGTCTCTGAGACTCTCCTGTGTAGCCTC TGGATTCAGCTTCAGTACTTATGGCATGCACTGGGTC CGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCGG TTATGTCATTTGATGGAAAGACTAAATACTATGCAGA CTCCGTGAAGGGCCGATTCACCATCTCCAGAGACAAT CCCAAGAACACACTATATCTGCAAATGAACAGCCTGA GAAGCGAAGACACGGCTGTGTATTATTGTGCGAAGGG GGGTGCAGCAGCGGCCGTCTTTGACTCCTGGGGCCCG GGAATACTGCTCACCGTCTCCTCA H007964 93 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGGAGGTCCCTAAGACTCTCCTGTGCAGCCTC TGGAGTCACCTTCAGTAGATATGGCATGCACTGGGTC CGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCAG TTATATCATATGATGAAAAGACTAAATACTATGCAGA CTCCGTGAAGGGCCGATTCACCATCTCCAGAGACAAT TCCAAGAACACACTGTTTCTGCACATGAACAGACTGA GATATGAGGACACGGCTGTATATTATTGTGCGAAAGG GGCCAGTAGCGGTGCGTACTTTGACTACTGGGGCCAG GGTACCCTGGTCACCGTCTCCTCA H007979 94 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGAAAGTCTCTGAGACTCTCCTGTGTAGCCTC TGGATTCACCTTCAGTACTTATGGCATGCACTGGGTC CGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCAG TTATGTCATTTGATGGAAAGACTAAATACTATGCAGA CTCCGTGAAGGGCCGATTCACCATCTCCAGAGACAAT CCCAAGAACACACTATATCTGCAAATGAACAGCCTGA GAAGCGAAGACACGGCTGTGTATTATTGTGCGAAGGG GGGTGCAGCAGCGGCCGTCTTTGACTCCTGGGGCCAG GGAATACTGCTCACCGTCTCCTCA H007961 95 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGAAAGTCTCTGAGACTCTCCTGTGTAGCCTC TGGATTCACCTTCAGTAGTTATGGCATGCACTGGGTC CGCCAGGCTCCGGGCAAGGGGCTGGAGTGGGTGGCGG TTATGTCATTTGATGGAAAGACTAAATACTATGCAGA CTCCGTGAAGGGCCGATTCACCATCTCCAGAGACAAC CCCAAGAACACACTATATCTGCAAATGAACAGCCTGA GAAGCGAAGACACGGCTGTCTATTATTGTGCGAAGGG GGGTGCAGCAGCGGCCGTCTTTGACTCCTGGGGCCAG GGAATACTGCTCACCGTCTCCTCA H007965 96 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGAAAGTCTCTGAGACTCTCCTGTGTAGCCTC TGGATTCACCTTCAGTAGTTATGGCATGCACTGGGTC CGCCAGGCTCCGGGCAAGGGGCTGGAGTGGGTGGCGG TTATGTCATTTGATGGAAAGACTAAATACTATGCAGA CTCCGTGAAGGGCCGATTCACCATCTCCAGAGACAAC CCCAAGAACACACTATATCTGCAAATGAACAGCCTGA GAAGCGAAGACACGGCTGTCTATTATTGTGCGAAGGG GGGTGCAGCAGCGGCCGTCTTTGACTCCTGGGGCCAG GGAATACTGCTCACCGTCTCCTCA H007968 97 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGGAGGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTAGATATGGCATGCACTGGGTC CGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCAG TTATGTCATATGATGGAAGTACTAAATACTATGCAGA CTCCGTGAAGGGCCGCTTCGCCATCTCCAGAGACAAT CCCAAGAACACGCTATTTCTGCAAATGAACAGCCTGA GACCTGACGACACGGCTGTATATTACTGTGCGAAAGG GGGGGCGGCA8CAGCTGTCATGGACGTCTGGGGCAAA GGGACCACGGTCACCGTCTCCTCA H007971 98 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGGAGGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTAGATATGGCATGCACTGGGTC CGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCAG TTATGTCATATGATGGAAGTACTAAATACTATGCAGA CTCCGTGAAGGGCCGCTTCGCCATCTCCAGAGACAAT CCCAAGAACACGCTATTTCTGCAAATGAACAGCCTGA GACCTGACGACACGGCTGTATATTACTGTGCGAAAGG GGGGGCGGCAGCAGCTGTCATGGACGTCTGGGGCAAA GGGACCACGGTCACCGTCTCCTCA H007983 99 GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCC (VH) AGCCTGGGAGGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAATAGGTTTGGCATGCACTGGGTC CGCCAGCGCCAGGTTCCAGGCAAGGGGCTGGAGTGGG TGGCAGTTATCTCATATGACGACAACACTAAATATTA TGCGGACTCCGTGAAGGGCCGTTTCACCATCTCCAGA GACAATAACAAGAGCACTCTCTATCTGCAAATGAGCA GCCTGAGAGTTGAGGACACGGCTGTCTATTTCTGTGC GAAGGGGTCGACAGCGGCAGCTGTTCTTGACTACTGG GGCCAGGGAACCCTTGTCACCGTCTCCTCA H007962 100 GAGGTgCAGCTGGTGGAGTCTGGGGGAGGCTTGGTAC (VH) AGCCTGGAGGGTCCCTGAGACTCTCCTGTGCAGCCTC TGGATTCACCTTCAGTAGTTATGAAATGAACTGGGTC CGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTTTCAT ACATTAGTAGTAGTGGTAGTACCATATACTACGCAGA CTCTGTGAAGGGCCGATTCACCATCTCCAGAGACAAC GCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGA GAGCCGAGGACACGGCTGTTTATTACTGTGCGAGAGT AGAGCAGCTCGCCCCCTCCCCCTACATGGACGTCTGG GGCAAAGGGACCACGGTCACCGTCTCCTCA H007966 101 CAGGTGCAGCTGGTGCAGTCTGGGGCAGAGGTGAAAA (VH) AGCCCGGGGAGTCTCTGAAGATCTCCTGTAAGGGTTC TGGATACAGCTTTACCAGCTACTGGATCGGCTGGGTG CGCCAGATGCCCGGGAAAGGCCTGGAGTGGATGGGGA TCATCTATCCTGGTGACTCTGATACCAGATACAGCCC GTCCTTCCAAGGCCAGGTCACCATCTCAGCCGACAAG TCCATCAGCACCGCCTACCTGCAGTGGAGCAGCCTGA AGGCCTCGGACACCGCCATGTATTACTGTGCGAGACG GGGTCTTCGAGGGGCCTACTACTACTACTACGGTATG GACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCT CA K006004 102 GACATCGTGATGACCCAGTCTCCAGACTCCCTGACTC (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAGATC CAGCCGGACTGTTTTATACAGGTCCAACAATAAAAAT TACTTAGCTTGGTATCAACATAAACCAGGACAGCCTC CTAAGTTGCTCATGTCCTGGGCATCTACCCGGGAAAC CGGGGTCCCTGACCGATTCAGTGGCAGCGGTTCTGGG ACACATTTCACTCTCACCATCACCAGCCTGCAGCCTG AAGATGTGGCAGTTTATTACTGTCAACAGTATTATAG TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAGA K005989 103 GACATCGTGATGACCCAGTCTCCAGGCTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCTCGAGGGCCACCATCAACTGCAAGTC CAGCCGGAGTCTTTTATACAGGTCCAACAATAAGAAT TATTTAGCTTGGTATCAACAGAAACCAGGACAGTCTC CTCGGCTTCTCATTTATTGGGCATCTTCCCGGGAATC CGGGGTCCCTGACCGATTCAGTGGCGGCGGGTCTGGG ACAAGTTTCACTCTCACCATCAGC K005991 104 GACATCGTGATGACCCAGTCTCCAGACTCCCTGACTC (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAGATC CAGCCGGACTGTTTTATACAGGTCCAACAATAAAAAT TACTTAGCTTGGTATCAACATAAACCAGGACAGCCTC CTAAGTTGCTCATGTCCTGGGCATCTACCCGGGAAAC CGGGGTCCCTGACCGATTCAGTGGCAGCGGTTCTGGG ACACATTTCACTCTCACCATCACCAGCCTGCAGCCTG AAGATGTGGCAGTTTATTACTGTCAACAGTATTATAG TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAGA K005992 105 GACATCGTGATGACCCAGTCTCCAGACTCCCTGACTC (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAGATC CAGCCGGACTGTTTTATACAGGTCCAACAATAAAAAT TACTTAGCTTGGTATCAACATAAACCAGGACAGCCTC CTAAGTTGCTCATGTCCTGGGCATCTACCCGGGAAAC CGGGGTCCCTGACCGATTCAGTGGCAGCGGTTCTGGG ACACATTTCACTCTCACCATCACCAGCCTGCAGCCTG AAGATGTGGCAGTTTATTACTGTCAACAGTATTATAG TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAGA K005998 106 GACATCGTGATGACCCAGTCTCCAGACTCCCTGACTC (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAGATC CAGCCGGACTGTTTTATACAGGTCCAACAATAAAAAT TACTTAGCTTGGTATCAACATAAACCAGGACAGCCTC CTAAGTTGCTCATGTCCTGGGCATCTACCCGGGAAAC CGGGGTCCCTGACCGATTCAGTGGCAGCGGTTCTGGG ACACATTTCACTCTCACCATCACCAGCCTGCAGCCTG AAGATGTGGCAGTTTATTACTGTCAACAGTATTATAG TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAGA K006018 107 GACATCGTGATGACCCAGTCTCCAGACTCCCTGACTC (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAGATC CAGCCGGACTGTTTTATACAGGTCCAACAATAAAAAT TACTTAGCTTGGTATCAACATAAACCAGGACAGCCTC CTAAGTTGCTCATGTCCTGGGCATCTACCCGGGAAAC CGGGGTCCCTGACCGATTCAGTGGCAGCGGTTCTGGG
ACACATTTCACTCTCACCATCACCAGCCTGCAGCCTG AAGATGTGGCAGTTTATTACTGTCAACAGTATTATAG TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAGA K005994 108 GACATCGTGATGACCCAGTCTCCAAACTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCGGGAGGGCCACCATCAACTGCAAGGC CAGCCAGAGTATTTTATACAGGTCCAACAATAAGAAC TATTTAGCTTGGTACCAACATAAAGCAGGACAGCCTC CCAAGCTGCTCATTTACTGGGCATCTACCCGGGAATC CGGGGTCCCTGAGCGATTCAGTGGCAGCGGGTCTAGG ACAGATTTCACTCTCACCATCAACGGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAAA K006002 109 GACATCGTGATGACCCAGTCTCCAAACTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCGGGAGGGCCACCATCAACTGCAAGGC CAGCCAGAGTATTTTATACAGGTCCAACAATAAGAAC TATTTAGCTTGGTACCAACATAAAGCAGGACAGCCTC CCAAGCTGCTCATTTACTGGGCATCTACCCGGGAATC CGGGGTCCCTGAGCGATTCAGTGGCAGCGGGTCTAGG ACAGATTTCACTCTCACCATCAACGGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAAA K005999 110 GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAAGTGCAAGTC CAGCCAGAGTGTCTTGTACAGCTCCAACAATAAGAAC TACTTAGCTTGGTACCAGCATAAACCAGGACAGCCTC CTAAGGTACTCGTTTACTGGGCATCCACCCGGGAATC CGGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTGGG ACAGATTTCACTCTCACCATCAGCAGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAACAATATTATAA TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTAGCA ATCAAG K006015 111 GACATCGTGATGACCCAGTCTCCAAACTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCGGGAGGGCCACCATCAACTGCAAGAC CAGCCAGAGTATTTTATACAGGTCCAACAATAAGAAC TACTTAGCTTGGTACCAGCATAAACCAGGACAGCCTC CCAAGCTGCTCATTTACTGGGCATCTACCCGGGAATC CAGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTAGG ACAGATTTCACTCTCACCATCAGCGGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCGTGGACGTTCGGCCAGGGGACCAAGGTGGAA ATCAAA K005995 112 GACATCGTGATGACCCAGTCTCCAAACTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCGGGAGGGCCACCATCAACTGCAAGAC CAGCCAGAGTATTTTATACAGGTCCAACAATAAGAAC TACTTAGCTTGGTACCAGCATAAATCAGGACAGCCTC CCAAGCTGCTCATTTACTGGGCATCTACCCGGGAATC CGGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTAGG ACAGATTTCACTCTCACCATCAGCGGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAAA K006000 113 GACATCGTGATGACCCAGTCTCCAAACTCCCTGGCTG (VL.sub.K) TGTCTCTGGGCGGGAGGGCCACCATCAACTGCAAGAC CAGCCAGAGTATTTTATACAGGTCCAACAATAAGAAC TACTTAGCTTGGTACCAGCATAAATCAGGACAGCCTC CCAAGCTGCTCATTTACTGGGCATCTACCCGGGAATC CGGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTAGG ACAGATTTCACTCTCACCATCAGCGGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCGTGGACGTTCGGCCAAGGGACCAAGGTGGAA ATCAAA K006003 114 GACATCGTGATGACCCAGTCTCCGGACTCCCTGACTG (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAGCTGCAAGTC CAGCCAGCGTCTTTTGTATAGTTCCAACAATAAGAAC TACTTAGCTTGGTACCAGCAGAAACCTGGACAGCCTC CTAAACTGCTCATGTACTGGGCGTCCACCCGGGAATC CGGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTGGG ACAGATTTCTCTCTCACCATCAGCAGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCCTGGACGTTCGGCCAAGGGACCAAGGTGGAA GTCAAA K006005 115 GACATCGTGATGACCCAGTCTCCGGACTCCCTGACTG (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAGCTGCAAGTC CAGCCAGCGTCTTTTGTATAGTTCCAACAATAAGAAC TACTTAGCTTGGTACCAGCAGAAACCTGGACAGCCTC CTAAACTGCTCATGTACTGGGCGTCCACCCGGGAATC CGGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTGGG ACAGATTTCTCTCTCACCATCAGCAGCCTGCAGGCTG AAGATGTGGCAGTTTATTACTGTCAGCAATATTATAA TCCTCCCTGGACGTTCGGCCAAGGGACCAAGGTGGAA GTCAAA K006019 116 GACATCGTGATGACCCAGTCTCCAGATTCCCTGACTC (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAAGTC CAGCCAGAGTCTGTTTTACAGGTCCAACAATAAGAGC TACTTAGCTTGGTATCAGCAAAAACCAGGGCAGCCTC CTAAACTGCTCATTTACTGGGCCTCTGTCCGGGAATC CGGGGTCCCTGACCGATTCACTGGCAGCGGGTCTGTA ACAGATTTCACTCTCACCATCAGCAGCCTGCGGGCTG AGGATGTGGCTGTTTATTATTGTCAACAGTATTTTAC TACTCCTCTCACTTTCGGCGGGGGGACCAAGGTGGCG ATCAAA K005996 117 GACATCGTGATGACCCAGTCTCTAGACTCCCTGACTG (VL.sub.K) TGTCTCTGGGCGAGAGGGCCACCATCAACTGCAAGTC CAGCCAGAGTCTTTTATACACCTCCAACAATAAGAAT TACTTAGCTTGGTACCAGCAGAAATCAGGACAGCCTC CTAAGTTACTCATTTACTGGGCGTCTATTCGGGATTC CGGGGTCCCTGACCGATTCAGTGGCAGCGGGTCTGCG ACAGATTTCACTCTCACCATCAACAACCTGCAGGCTG AAGATGTGGCAGTTTACTTCTGTCAGCAATATTACAA GACTCCTCTCACTTTCGGCGGGGGGACCAAGGTGGAG GTCAGA K006001 118 GACATCCAGWTGACCCAGTCTCCTTCCACCCTGTCTG (VL.sub.K) CATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGC CAGTCAGAGTATTAGTAGCTGGTTGGCCTGGTATCAG CAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATA AGGCGTCTAGTTTAGAAAGTGGGGTCCCATCAAGGTT CAGCGGCAGTGGATCTGGGACAGAATTCACTCTCACC ATCAGCAGCCTGCAGCCTGATGATTTTGCAACTTATT ACTGCCAACAGTATAATAGTTATTCTTGGACGTTCGG CCAAGGGACCAAGGTGGAAATCAAA
[0121] d. Antibody Preparation/Production
[0122] Antibodies may be prepared by any of a variety of techniques. In general, antibodies can be produced by cell culture techniques, including the generation of monoclonal antibodies via conventional techniques, or via transfection of antibody genes, heavy chains, and/or light chains into suitable bacterial or mammalian cell hosts, in order to allow for the production of antibodies, wherein the antibodies may be recombinant. The various forms of the term "transfection" are intended to encompass a wide variety of techniques commonly used for the introduction of exogenous DNA into a prokaryotic or eukaryotic host cell, e.g., electroporation, calcium-phosphate precipitation, DEAE-dextran transfection and the like. Although it is possible to express the antibodies of the invention in either prokaryotic or eukaryotic host cells, expression of antibodies in eukaryotic cells is preferable, and most preferable in mammalian host cells, because such eukaryotic cells (and in particular mammalian cells) are more likely than prokaryotic cells to assemble and secrete a properly folded and immunologically active antibody.
[0123] Exemplary mammalian host cells for expressing the recombinant antibodies of the invention include Chinese Hamster Ovary (CHO cells) (including dhfr-CHO cells, described in Urlaub and Chasin, Proc. Natl. Acad. Sci. USA, 77: 4216-4220 (1980)), used with a DHFR selectable marker, e.g., as described in Kaufman and Sharp, J. Mol. Biol., 159: 601-621 (1982), NSO myeloma cells, COS cells, HEK 293T cells, and SP2 cells. When recombinant expression vectors encoding antibody genes are introduced into mammalian host cells, the antibodies are produced by culturing the host cells for a period of time sufficient to allow for expression of the antibody in the host cells or, more preferably, secretion of the antibody into the culture medium in which the host cells are grown. Antibodies can be recovered from the culture medium using standard protein purification methods.
[0124] Host cells can also be used to produce functional antibody fragments, such as Fab fragments or scFv molecules. It will be understood that variations on the above procedure are within the scope of the present invention. For example, it may be desirable to transfect a host cell with DNA encoding functional fragments of either the light chain and/or the heavy chain of an antibody of this invention. Recombinant DNA technology may also be used to remove some, or all, of the DNA encoding either or both of the light and heavy chains that is not necessary for binding to the antigens of interest. The molecules expressed from such truncated DNA molecules are also encompassed by the antibodies of the invention. In addition, bifunctional antibodies may be produced in which one heavy and one light chain are an antibody of the invention (i.e., binds human CFH) and the other heavy and light chain are specific for an antigen other than human CFH by crosslinking an antibody of the invention to a second antibody by standard chemical crosslinking methods.
[0125] In a preferred system for recombinant expression of an antibody, or antigen-binding portion thereof, of the invention, a recombinant expression vector encoding both the antibody heavy chain and the antibody light chain is introduced into dhfr-CHO cells by calcium phosphate-mediated transfection. Within the recombinant expression vector, the antibody heavy and light chain genes are each operatively linked to CMV enhancer/AdMLP promoter regulatory elements to drive high levels of transcription of the genes. The recombinant expression vector also carries a DHFR gene, which allows for selection of CHO cells that have been transfected with the vector using methotrexate selection/amplification. The selected transformant host cells are cultured to allow for expression of the antibody heavy and light chains and intact antibody is recovered from the culture medium. Standard molecular biology techniques are used to prepare the recombinant expression vector, transfect the host cells, select for transformants, culture the host cells, and recover the antibody from the culture medium. Still further, the invention provides a method of synthesizing a recombinant antibody of the invention by culturing a host cell of the invention in a suitable culture medium until a recombinant antibody of the invention is synthesized. The method can further comprise isolating the recombinant antibody from the culture medium.
[0126] Methods of preparing monoclonal antibodies involve the preparation of immortal cell lines capable of producing antibodies having the desired specificity. Such cell lines may be produced from spleen cells obtained from an immunized animal. The animal may be immunized with CFH or a fragment and/or variant thereof. For example, any of SEQ ID NOs:81-83, or a variant of SEQ ID NOs:81-83 may be used to immunize the animal. The immunizing antigen may be reduced or not reduced. The spleen cells may then be immortalized by, for example, fusion with a myeloma cell fusion partner. A variety of fusion techniques may be employed. For example, the spleen cells and myeloma cells may be combined with a nonionic detergent for a few minutes and then plated at low density on a selective medium that supports that growth of hybrid cells, but not myeloma cells. One such technique uses hypoxanthine, aminopterin, thymidine (HAT) selection. Another technique includes eletrofusion. After a sufficient time, usually about 1 to 2 weeks, colonies of hybrids are observed. Single colonies are selected and their culture supernatants tested for binding activity against the polypeptide. Hybridomas having high reactivity and specificity may be used.
[0127] Monoclonal antibodies may be isolated from the supernatants of growing hybridoma colonies. In addition, various techniques may be employed to enhance the yield, such as injection of the hybridoma cell line into the peritoneal cavity of a suitable vertebrate host, such as a mouse. Monoclonal antibodies may then be harvested from the ascites fluid or the blood. Contaminants may be removed from the antibodies by conventional techniques, such as chromatography, gel filtration, precipitation, and extraction. Affinity chromatography is an example of a method that can be used in a process to purify the antibodies.
[0128] The proteolytic enzyme papain preferentially cleaves IgG molecules to yield several fragments, two of which (the F(ab) fragments) each comprise a covalent heterodimer that includes an intact antigen-binding site. The enzyme pepsin is able to cleave IgG molecules to provide several fragments, including the F(ab')2 fragment, which comprises both antigen-binding sites.
[0129] The Fv fragment can be produced by preferential proteolytic cleavage of an IgM, and on rare occasions IgG or IgA immunoglobulin molecules. The Fv fragment may be derived using recombinant techniques. The Fv fragment includes a non-covalent VH::VL heterodimer including an antigen-binding site that retains much of the antigen recognition and binding capabilities of the native antibody molecule.
[0130] The antibody, antibody fragment, or derivative may comprise a heavy chain and a light chain complementarity determining region ("CDR") set, respectively interposed between a heavy chain and a light chain framework ("FR") set which provide support to the CDRs and define the spatial relationship of the CDRs relative to each other. The DR set may contain three hypervariable regions of a heavy or light chain V region. Proceeding from the N-terminus of a heavy or light chain, these regions are denoted as "CDR1," "CDR2," and "CDR3," respectively. An antigen-binding site, therefore, may include six CDRs, comprising the CDR set from each of a heavy and a light chain V region. A polypeptide comprising a single CDR, (e.g., a CDR1, CDR2, or CDR3) may be referred to as a "molecular recognition unit." Crystallographic analyses of antigen-antibody complexes have demonstrated that the amino acid residues of CDRs form extensive contact with bound antigen, wherein the most extensive antigen contact is with the heavy chain CDR3. Thus, the molecular recognition units may be primarily responsible for the specificity of an antigen-binding site. In general, the CDR residues are directly and most substantially involved in influencing antigen binding.
[0131] Other suitable methods of producing or isolating antibodies of the requisite specificity can be used, including, but not limited to, methods that select recombinant antibody from a peptide or protein library (e.g., but not limited to, a bacteriophage, ribosome, oligonucleotide, RNA, cDNA, yeast or the like, display library); e.g., as available from various commercial vendors such as Cambridge Antibody Technologies (Cambridgeshire, UK), MorphoSys (Martinsreid/Planegg, Del.), Biovation (Aberdeen, Scotland, UK) Biolnvent (Lund, Sweden), using methods known in the art. See U.S. Pat. Nos. 4,704,692; 5,723,323; 5,763,192; 5,814,476; 5,817,483; 5,824,514; 5,976,862. Alternative methods rely upon immunization of transgenic animals (e.g., SCID mice, Nguyen et al. (1997) Microbiol. Immunol. 41:901-907; Sandhu et al. (1996) Crit. Rev. Biotechnol. 16:95-118; Eren et al. (1998) Immunol. 93:154-161) that are capable of producing a repertoire of human antibodies, as known in the art and/or as described herein. Such techniques, include, but are not limited to, ribosome display (Hanes et al. (1997) Proc. Natl. Acad. Sci. USA, 94:4937-4942; Hanes et al. (1998) Proc. Natl. Acad. Sci. USA, 95:14130-14135); single cell antibody producing technologies (e.g., selected lymphocyte antibody method ("SLAM") (U.S. Pat. No. 5,627,052, Wen et al. (1987) J. Immunol. 17:887-892; Babcook et al. (1996) Proc. Natl. Acad. Sci. USA 93:7843-7848); gel microdroplet and flow cytometry (Powell et al. (1990) Biotechnol. 8:333-337; One Cell Systems, (Cambridge, Mass).; Gray et al. (1995) J. Imm. Meth. 182:155-163; Kenny et al. (1995) Bio/Technol. 13:787-790); B-cell selection (Steenbakkers et al. (1994) Molec. Biol. Reports 19:125-134 (1994)). In particular, human antibodies against CFH may be derived, sequenced and characterized from peripheral human B lymphocytes using methods as described in Liao et al. (2013) Immunity 38(1): 176-186; Bonsignori et al. (2012) J Virol 86(21): 11521-11532; Moody et al. (2012) J Virol 86(14): 7496-7507; Gray et al. (2011) J Virol 85(15): 7719-7729; Morris et al. (2011) PLoS ONE 6(9): e23532; and Liao et al. (2009) J Virol Methods 158(1-2): 171-179.
[0132] An affinity matured antibody may be produced by any one of a number of procedures that are known in the art. For example, see Marks et al., BioTechnology, 10: 779-783 (1992) describes affinity maturation by VH and VL domain shuffling. Random mutagenesis of CDR and/or framework residues is described by Barbas et al., Proc. Nat. Acad. Sci. USA, 91: 3809-3813 (1994); Schier et al., Gene, 169: 147-155 (1995); Yelton et al., J. Immunol., 155: 1994-2004 (1995); Jackson et al., J. Immunol., 154(7): 3310-3319 (1995); Hawkins et al, J. Mol. Biol., 226: 889-896 (1992). Selective mutation at selective mutagenesis positions and at contact or hypermutation positions with an activity enhancing amino acid residue is described in U.S. Pat. No. 6,914,128 B1.
[0133] Antibody variants of the present invention can also be prepared by delivering a polynucleotide encoding an antibody of this invention to a suitable host such as to provide transgenic animals or mammals, such as goats, cows, horses, sheep, and the like, that produce such antibodies in their milk. These methods are known in the art and are described for example in U.S. Pat. Nos. 5,827,690; 5,849,992; 4,873,316; 5,849,992; 5,994,616; 5,565,362; and 5,304,489.
[0134] Antibody variants also can be prepared by delivering a polynucleotide of this invention to provide transgenic plants and cultured plant cells (e.g., but not limited to tobacco, maize, and duckweed) that produce such antibodies, specified portions or variants in the plant parts or in cells cultured therefrom. For example, Cramer et al. (1999) Curr. Top. Microbiol. Immunol. 240:95-118 and references cited therein, describe the production of transgenic tobacco leaves expressing large amounts of recombinant proteins, e.g., using an inducible promoter. Transgenic maize have been used to express mammalian proteins at commercial production levels, with biological activities equivalent to those produced in other recombinant systems or purified from natural sources. See, e.g., Hood et al., Adv. Exp. Med. Biol. (1999) 464:127-147 and references cited therein. Antibody variants have also been produced in large amounts from transgenic plant seeds including antibody fragments, such as single chain antibodies (scFv's), including tobacco seeds and potato tubers. See, e.g., Conrad et al. (1998) Plant Mol. Biol. 38:101-109 and reference cited therein. Thus, antibodies of the present invention can also be produced using transgenic plants, according to known methods.
[0135] Antibody derivatives can be produced, for example, by adding exogenous sequences to modify immunogenicity or reduce, enhance or modify binding, affinity, on-rate, off-rate, avidity, specificity, half-life, or any other suitable characteristic. Generally, part or all of the non-human or human CDR sequences are maintained while the non-human sequences of the variable and constant regions are replaced with human or other amino acids.
[0136] Small antibody fragments may be diabodies having two antigen-binding sites, wherein fragments comprise a heavy chain variable domain (VH) connected to a light chain variable domain (VL) in the same polypeptide chain (VH VL). See for example, EP 404,097; WO 93/11161; and Hollinger et al., (1993) Proc. Natl. Acad. Sci. USA 90:6444-6448. By using a linker that is too short to allow pairing between the two domains on the same chain, the domains are forced to pair with the complementary domains of another chain and create two antigen-binding sites. See also, U.S. Pat. No. 6,632,926 to Chen et al. which is hereby incorporated by reference in its entirety and discloses antibody variants that have one or more amino acids inserted into a hypervariable region of the parent antibody and a binding affinity for a target antigen which is at least about two fold stronger than the binding affinity of the parent antibody for the antigen.
[0137] The antibody may be a linear antibody. The procedure for making a linear antibody is known in the art and described in Zapata et al. (1995) Protein Eng. 8(10):1057-1062. Briefly, these antibodies comprise a pair of tandem Fd segments (VH-CH1-VH-CH1) which form a pair of antigen binding regions. Linear antibodies can be bispecific or monospecific.
[0138] The antibodies may be recovered and purified from recombinant cell cultures by known methods including, but not limited to, protein A purification, ammonium sulfate or ethanol precipitation, acid extraction, anion or cation exchange chromatography, phosphocellulose chromatography, hydrophobic interaction chromatography, affinity chromatography, hydroxylapatite chromatography and lectin chromatography. High performance liquid chromatography ("HPLC") can also be used for purification.
[0139] It may be useful to detectably or therapeutically label the antibody. Methods for conjugating antibodies to these agents are known in the art. For the purpose of illustration only, antibodies can be labeled with a detectable moiety such as a radioactive atom, a chromophore, a fluorophore, or the like. Such labeled antibodies can be used for diagnostic techniques, either in vivo, or in an isolated test sample. Antibodies can also be conjugated, for example, to a pharmaceutical agent, such as chemotherapeutic drug or a toxin. They can be linked to a cytokine, to a ligand, to another antibody. Suitable agents for coupling to antibodies to achieve an anti-tumor effect include cytokines, such as interleukin 2 (IL-2) and Tumor Necrosis Factor (TNF); photosensitizers, for use in photodynamic therapy, including aluminum (III) phthalocyanine tetrasulfonate, hematoporphyrin, and phthalocyanine; radionuclides, such as iodine-131 (131I), yttrium-90 (90Y), bismuth-212 (212Bi), bismuth-213 (213Bi), technetium-99m (99mTc), rhenium-186 (186Re), and rhenium-188 (188Re); antibiotics, such as doxorubicin, adriamycin, daunorubicin, methotrexate, daunomycin, neocarzinostatin, and carboplatin; bacterial, plant, and other toxins, such as diphtheria toxin, pseudomonas exotoxin A, staphylococcal enterotoxin A, abrin-A toxin, ricin A (deglycosylated ricin A and native ricin A), TGF-alpha toxin, cytotoxin from chinese cobra (naj a naj a atra), and gelonin (a plant toxin); ribosome inactivating proteins from plants, bacteria and fungi, such as restrictocin (a ribosome inactivating protein produced by Aspergillus restrictus), saporin (a ribosome inactivating protein from Saponaria officinalis), and RNase; tyrosine kinase inhibitors; ly207702 (a difluorinated purine nucleoside); liposomes containing anti cystic agents (e.g., antisense oligonucleotides, plasmids which encode for toxins, methotrexate, etc.); and other antibodies or antibody fragments, such as F(ab).
[0140] The antibodies can be sequenced and replicated by recombinant or synthetic means. They also can be further sequenced down to the linear sequence of nucleotides that encode them. Accordingly, this invention provides these polynucleotides, alone or in combination with a carrier, vector or host cell, as described above, that encode a sequence of an antibody of this invention.
[0141] Antibody production via the use of hybridoma technology, the selected lymphocyte antibody method (SLAM), transgenic animals, and recombinant antibody libraries is described in more detail below.
[0142] (1) Anti-CFH Monoclonal Antibodies Using Hybridoma Technology
[0143] Monoclonal antibodies can be prepared using a wide variety of techniques known in the art including the use of hybridoma, recombinant, and phage display technologies, or a combination thereof. For example, monoclonal antibodies can be produced using hybridoma techniques including those known in the art and taught, for example, in Harlow et al., Antibodies: A Laboratory Manual, second edition, (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1988); Hammerling, et al., In Monoclonal Antibodies and T-Cell Hybridomas, (Elsevier, N.Y., 1981). It is also noted that the term "monoclonal antibody" as used herein is not limited to antibodies produced through hybridoma technology. The term "monoclonal antibody" refers to an antibody that is derived from a single clone, including any eukaryotic, prokaryotic, or phage clone, and not the method by which it is produced.
[0144] In an embodiment, the present invention provides methods of generating monoclonal antibodies as well as antibodies produced by the method. The method may comprise culturing a hybridoma cell secreting an antibody of the invention wherein, preferably, the hybridoma is generated by fusing splenocytes isolated from an animal, e.g., a rat or a mouse, immunized with CFH with myeloma cells and then screening the hybridomas resulting from the fusion for hybridoma clones that secrete an antibody able to bind a polypeptide, such as GPPPPIDNGDITSFP(GGGK-biotin) (SEQ ID NO:122). Briefly, rats can be immunized with a CFH antigen. In a preferred embodiment, the CFH antigen is administered with an adjuvant to stimulate the immune response. Such adjuvants include complete or incomplete Freund's adjuvant, RIBI (muramyl dipeptides) or ISCOM (immunostimulating complexes). Such adjuvants may protect the polypeptide from rapid dispersal by sequestering it in a local deposit, or they may contain substances that stimulate the host to secrete factors that are chemotactic for macrophages and other components of the immune system. Preferably, if a polypeptide is being administered, the immunization schedule will involve two or more administrations of the polypeptide, spread out over several weeks; however, a single administration of the polypeptide may also be used.
[0145] After immunization of an animal with a CFH antigen, antibodies and/or antibody-producing cells may be obtained from the animal. An anti-CFH antibody-containing serum is obtained from the animal by bleeding or sacrificing the animal. The serum may be used as it is obtained from the animal, an immunoglobulin fraction may be obtained from the serum, or the anti-CFH antibodies may be purified from the serum. Serum or immunoglobulins obtained in this manner are polyclonal, thus having a heterogeneous array of properties.
[0146] Once an immune response is detected, e.g., antibodies specific for the antigen CFH are detected in the rat serum, the rat spleen is harvested and splenocytes isolated. The splenocytes are then fused by well-known techniques to any suitable myeloma cells, for example, cells from cell line SP20 available from the American Type Culture Collection (ATCC, Manassas, Va., US). Hybridomas are selected and cloned by limited dilution. The hybridoma clones are then assayed by methods known in the art for cells that secrete antibodies capable of binding CFH. Ascites fluid, which generally contains high levels of antibodies, can be generated by immunizing rats with positive hybridoma clones.
[0147] In another embodiment, antibody-producing immortalized hybridomas may be prepared from the immunized animal. After immunization, the animal is sacrificed and the splenic B cells are fused to immortalized myeloma cells as is well known in the art. See, e.g., Harlow and Lane, supra. In a preferred embodiment, the myeloma cells do not secrete immunoglobulin polypeptides (a non-secretory cell line). After fusion and antibiotic selection, the hybridomas are screened using CFH, or a portion thereof, such as GPPPPIDNGDITSFP(GGGK-biotin) (SEQ ID NO:122), or a cell expressing CFH. In a preferred embodiment, the initial screening is performed using an enzyme-linked immunosorbent assay (ELISA) or a radioimmunoassay (MA), preferably an ELISA. An example of ELISA screening is provided in PCT Publication No. WO 00/37504.
[0148] Anti-CFH antibody-producing hybridomas are selected, cloned, and further screened for desirable characteristics, including robust hybridoma growth, high antibody production, and desirable antibody characteristics. Hybridomas may be cultured and expanded in vivo in syngeneic animals, in animals that lack an immune system, e.g., nude mice, or in cell culture in vitro. Methods of selecting, cloning and expanding hybridomas are well known to those of ordinary skill in the art.
[0149] In a preferred embodiment, hybridomas are rat hybridomas. In another embodiment, hybridomas are produced in a non-human, non-rat species such as mice, sheep, pigs, goats, cattle, rabbits, or horses. In yet another preferred embodiment, the hybridomas are human hybridomas, in which a human non-secretory myeloma is fused with a human cell expressing an anti-CFH antibody.
[0150] Antibody fragments that recognize specific epitopes may be generated by known techniques. For example, Fab and F(ab')2 fragments of the invention may be produced by proteolytic cleavage of immunoglobulin molecules, using enzymes such as papain (to produce two identical Fab fragments) or pepsin (to produce an F(ab')2 fragment). A F(ab')2 fragment of an IgG molecule retains the two antigen-binding sites of the larger ("parent") IgG molecule, including both light chains (containing the variable light chain and constant light chain regions), the CH1 domains of the heavy chains, and a disulfide-forming hinge region of the parent IgG molecule. Accordingly, an F(ab')2 fragment is still capable of crosslinking antigen molecules like the parent IgG molecule.
[0151] (2) Anti-CFH Monoclonal Antibodies Using SLAM
[0152] In another aspect of the invention, recombinant antibodies are generated from single, isolated lymphocytes using a procedure referred to in the art as the selected lymphocyte antibody method (SLAM), as described in U.S. Pat. No. 5,627,052; PCT Publication No. WO 92/02551; and Babcook et al., Proc. Natl. Acad. Sci. USA, 93: 7843-7848 (1996). In this method, single cells secreting antibodies of interest, e.g., lymphocytes derived from any one of the immunized animals are screened using an antigen-specific hemolytic plaque assay, wherein the antigen CFH, a subunit of CFH, or a fragment thereof, is coupled to sheep red blood cells using a linker, such as biotin, and used to identify single cells that secrete antibodies with specificity for CFH. Following identification of antibody-secreting cells of interest, heavy- and light-chain variable region cDNAs are rescued from the cells by reverse transcriptase-PCR (RT-PCR) and these variable regions can then be expressed, in the context of appropriate immunoglobulin constant regions (e.g., human constant regions), in mammalian host cells, such as COS or CHO cells. The host cells transfected with the amplified immunoglobulin sequences, derived from in vivo selected lymphocytes, can then undergo further analysis and selection in vitro, for example, by panning the transfected cells to isolate cells expressing antibodies to CFH. The amplified immunoglobulin sequences further can be manipulated in vitro, such as by in vitro affinity maturation methods. See, for example, PCT Publication No. WO 97/29131 and PCT Publication No. WO 00/56772.
[0153] (3) Anti-CFH Monoclonal Antibodies Using Transgenic Animals
[0154] In another embodiment of the invention, antibodies are produced by immunizing a non-human animal comprising some, or all, of the human immunoglobulin locus with a CFH antigen. In an embodiment, the non-human animal is a XENOMOUSE.RTM. transgenic mouse, an engineered mouse strain that comprises large fragments of the human immunoglobulin loci and is deficient in mouse antibody production. See, e.g., Green et al., Nature Genetics, 7: 13-21 (1994) and U.S. Pat. Nos. 5,916,771; 5,939,598; 5,985,615; 5,998,209; 6,075,181; 6,091,001; 6,114,598; and 6,130,364. See also PCT Publication Nos. WO 91/10741; WO 94/02602; WO 96/34096; WO 96/33735; WO 98/16654; WO 98/24893; WO 98/50433; WO 99/45031; WO 99/53049; WO 00/09560; and WO 00/37504. The XENOMOUSE.RTM. transgenic mouse produces an adult-like human repertoire of fully human antibodies, and generates antigen-specific human monoclonal antibodies. The XENOMOUSE.RTM. transgenic mouse contains approximately 80% of the human antibody repertoire through introduction of megabase sized, germline configuration YAC fragments of the human heavy chain loci and x light chain loci. See Mendez et al., Nature Genetics, 15: 146-156 (1997), Green and Jakobovits, J. Exp. Med., 188: 483-495 (1998), the disclosures of which are hereby incorporated by reference.
[0155] (4) Anti-CFH Monoclonal Antibodies Using Recombinant Antibody Libraries
[0156] In vitro methods also can be used to make the antibodies of the invention, wherein an antibody library is screened to identify an antibody having the desired CFH-binding specificity. Methods for such screening of recombinant antibody libraries are well known in the art and include methods described in, for example, U.S. Pat. No. 5,223,409 (Ladner et al.); PCT Publication No. WO 92/18619 (Kang et al.); PCT Publication No. WO 91/17271 (Dower et al.); PCT Publication No. WO 92/20791 (Winter et al.); PCT Publication No. WO 92/15679 (Markland et al.); PCT Publication No. WO 93/01288 (Breitling et al.); PCT Publication No. WO 92/01047 (McCafferty et al.); PCT Publication No. WO 92/09690 (Garrard et al.); Fuchs et al., Bio/Technology, 9: 1369-1372 (1991); Hay et al., Hum. Antibod. Hybridomas, 3: 81-85 (1992); Huse et al., Science, 246: 1275-1281 (1989); McCafferty et al., Nature, 348: 552-554 (1990); Griffiths et al., EMBO J., 12: 725-734 (1993); Hawkins et al., J. Mol. Biol., 226: 889-896 (1992); Clackson et al., Nature, 352: 624-628 (1991); Gram et al., Proc. Natl. Acad. Sci. USA, 89: 3576-3580 (1992); Garrard et al., Bio/Technology, 9: 1373-1377 (1991); Hoogenboom et al., Nucl. Acids Res., 19: 4133-4137 (1991); Barbas et al., Proc. Natl. Acad. Sci. USA, 88: 7978-7982 (1991); US Patent Application Publication No. 2003/0186374; and PCT Publication No. WO 97/29131, the contents of each of which are incorporated herein by reference.
[0157] The recombinant antibody library may be from a subject immunized with CFH, or a portion of CFH. Alternatively, the recombinant antibody library may be from a naive subject, i.e., one who has not been immunized with CFH, such as a human antibody library from a human subject who has not been immunized with human CFH. Antibodies of the invention are selected by screening the recombinant antibody library with the peptide comprising human CFH to thereby select those antibodies that recognize CFH. Methods for conducting such screening and selection are well known in the art, such as described in the references in the preceding paragraph. To select antibodies of the invention having particular binding affinities for CFH, such as those that dissociate from human CFH with a particular K.sub.off rate constant, the art-known method of surface plasmon resonance can be used to select antibodies having the desired K.sub.off rate constant. To select antibodies of the invention having a particular neutralizing activity for hCFH, such as those with a particular IC.sub.50, standard methods known in the art for assessing the inhibition of CFH activity may be used.
[0158] In one aspect, the invention pertains to an isolated antibody, or an antigen-binding portion thereof, that binds human CFH. Preferably, the antibody is a neutralizing antibody. In various embodiments, the antibody is a recombinant antibody or a monoclonal antibody.
[0159] For example, antibodies of the present invention can also be generated using various phage display methods known in the art. In phage display methods, functional antibody domains are displayed on the surface of phage particles which carry the polynucleotide sequences encoding them. Such phage can be utilized to display antigen-binding domains expressed from a repertoire or combinatorial antibody library (e.g., human or murine). Phage expressing an antigen binding domain that binds the antigen of interest can be selected or identified with antigen, e.g., using labeled antigen or antigen bound or captured to a solid surface or bead. Phage used in these methods are typically filamentous phage including fd and M13 binding domains expressed from phage with Fab, Fv, or disulfide stabilized Fv antibody domains recombinantly fused to either the phage gene III or gene VIII protein. Examples of phage display methods that can be used to make the antibodies of the present invention include those disclosed in Brinkmann et al., J. Immunol. Methods, 182: 41-50 (1995); Ames et al., J. Immunol. Methods, 184:177-186 (1995); Kettleborough et al., Eur. J. Immunol., 24: 952-958 (1994); Persic et al., Gene, 187: 9-18 (1997); Burton et al., Advances in Immunology, 57: 191-280 (1994); PCT Publication No. WO 92/01047; PCT Publication Nos. WO 90/02809; WO 91/10737; WO 92/01047; WO 92/18619; WO 93/11236; WO 95/15982; WO 95/20401; and U.S. Pat. Nos. 5,698,426; 5,223,409; 5,403,484; 5,580,717; 5,427,908; 5,750,753; 5,821,047; 5,571,698; 5,427,908; 5,516,637; 5,780,225; 5,658,727; 5,733,743; and 5,969,108.
[0160] As described in the above references, after phage selection, the antibody coding regions from the phage can be isolated and used to generate whole antibodies including human antibodies or any other desired antigen binding fragment, and expressed in any desired host, including mammalian cells, insect cells, plant cells, yeast, and bacteria, e.g., as described in detail below. For example, techniques to recombinantly produce Fab, Fab', and F(ab').sub.2 fragments can also be employed using methods known in the art such as those disclosed in PCT publication No. WO 92/22324; Mullinax et al., BioTechniques, 12(6): 864-869 (1992); Sawai et al., Am. J. Reprod. Immunol., 34: 26-34 (1995); and Better et al., Science, 240: 1041-1043 (1988). Examples of techniques which can be used to produce single-chain Fvs and antibodies include those described in U.S. Pat. Nos. 4,946,778 and 5,258,498; Huston et al., Methods in Enzymology, 203: 46-88 (1991); Shu et al., Proc. Natl. Acad. Sci. USA, 90: 7995-7999 (1993); and Skerra et al., Science, 240: 1038-1041 (1988).
[0161] Alternative to screening of recombinant antibody libraries by phage display, other methodologies known in the art for screening large combinatorial libraries can be applied to the identification of antibodies of the invention. One type of alternative expression system is one in which the recombinant antibody library is expressed as RNA-protein fusions, as described in PCT Publication No. WO 98/31700 (Szostak and Roberts), and in Roberts and Szostak, Proc. Natl. Acad. Sci. USA, 94: 12297-12302 (1997). In this system, a covalent fusion is created between an mRNA and the peptide or protein that it encodes by in vitro translation of synthetic mRNAs that carry puromycin, a peptidyl acceptor antibiotic, at their 3' end. Thus, a specific mRNA can be enriched from a complex mixture of mRNAs (e.g., a combinatorial library) based on the properties of the encoded peptide or protein, e.g., antibody, or portion thereof, such as binding of the antibody, or portion thereof, to the dual specificity antigen. Nucleic acid sequences encoding antibodies, or portions thereof, recovered from screening of such libraries can be expressed by recombinant means, as described above (e.g., in mammalian host cells) and, moreover, can be subjected to further affinity maturation by either additional rounds of screening of mRNA-peptide fusions in which mutations have been introduced into the originally selected sequence(s), or by other methods for affinity maturation in vitro of recombinant antibodies, as described above. A preferred example of this methodology is PROfusion display technology.
[0162] In another approach, the antibodies of the present invention can also be generated using yeast display methods known in the art. In yeast display methods, genetic methods are used to tether antibody domains to the yeast cell wall and display them on the surface of yeast. In particular, such yeast can be utilized to display antigen-binding domains expressed from a repertoire or combinatorial antibody library (e.g., human or murine). Examples of yeast display methods that can be used to make the antibodies of the present invention include those disclosed in U.S. Pat. No. 6,699,658 (Wittrup et al.) incorporated herein by reference.
[0163] e. Production of Recombinant CFH Antibodies
[0164] Antibodies of the present invention may be recombinant antibodies and may be produced by any of a number of techniques known in the art. For example, expression from host cells, wherein expression vector(s) encoding the heavy and light chains is (are) transfected into a host cell by standard techniques. The various forms of the term "transfection" are intended to encompass a wide variety of techniques commonly used for the introduction of exogenous DNA into a prokaryotic or eukaryotic host cell, e.g., electroporation, calcium-phosphate precipitation, DEAE-dextran transfection, and the like. Although it is possible to express the antibodies of the invention in either prokaryotic or eukaryotic host cells, expression of antibodies in eukaryotic cells is preferable, and most preferable in mammalian host cells, because such eukaryotic cells (and in particular mammalian cells) are more likely than prokaryotic cells to assemble and secrete a properly folded and immunologically active antibody. The recombinant antibody may be a humanized antibody.
[0165] Exemplary mammalian host cells for expressing the recombinant antibodies of the invention include Chinese Hamster Ovary (CHO cells) (including dhfr-CHO cells, described in Urlaub and Chasin, Proc. Natl. Acad. Sci. USA, 77: 4216-4220 (1980), used with a DHFR selectable marker, e.g., as described in Kaufman and Sharp, J. Mol. Biol., 159: 601-621 (1982), NSO myeloma cells, COS cells, and SP2 cells. When recombinant expression vectors encoding antibody genes are introduced into mammalian host cells, the antibodies are produced by culturing the host cells for a period of time sufficient to allow for expression of the antibody in the host cells or, more preferably, secretion of the antibody into the culture medium in which the host cells are grown. Antibodies can be recovered from the culture medium using standard protein purification methods.
[0166] Host cells can also be used to produce functional antibody fragments, such as Fab fragments or scFv molecules. It will be understood that variations on the above procedure are within the scope of the present invention. For example, it may be desirable to transfect a host cell with DNA encoding functional fragments of either the light chain and/or the heavy chain of an antibody of this invention. Recombinant DNA technology may also be used to remove some, or all, of the DNA encoding either or both of the light and heavy chains that is not necessary for binding to the antigens of interest. The molecules expressed from such truncated DNA molecules are also encompassed by the antibodies of the invention. In addition, bifunctional antibodies may be produced in which one heavy and one light chain are an antibody of the invention (i.e., binds human CFH) and the other heavy and light chain are specific for an antigen other than human CFH by crosslinking an antibody of the invention to a second antibody by standard chemical crosslinking methods.
[0167] In a preferred system for recombinant expression of an antibody, or antigen-binding portion thereof, of the invention, a recombinant expression vector encoding both the antibody heavy chain and the antibody light chain is introduced into dhfr-CHO cells by calcium phosphate-mediated transfection. Within the recombinant expression vector, the antibody heavy and light chain genes are each operatively linked to CMV enhancer/AdMLP promoter regulatory elements to drive high levels of transcription of the genes. The recombinant expression vector also carries a DHFR gene, which allows for selection of CHO cells that have been transfected with the vector using methotrexate selection/amplification. The selected transformant host cells are cultured to allow for expression of the antibody heavy and light chains and intact antibody is recovered from the culture medium. Standard molecular biology techniques are used to prepare the recombinant expression vector, transfect the host cells, select for transformants, culture the host cells, and recover the antibody from the culture medium. Still further, the invention provides a method of synthesizing a recombinant antibody of the invention by culturing a host cell of the invention in a suitable culture medium until a recombinant antibody of the invention is synthesized. The method can further comprise isolating the recombinant antibody from the culture medium.
[0168] (1) Humanized Antibody
[0169] The humanized antibody may be an antibody or a variant, derivative, analog or portion thereof which immunospecifically binds to an antigen of interest and which comprises a framework (FR) region having substantially the amino acid sequence of a human antibody and a complementary determining region (CDR) having substantially the amino acid sequence of a non-human antibody. The humanized antibody may be from a non-human species antibody that binds the desired antigen having one or more complementarity determining regions (CDRs) from the non-human species and framework regions from a human immunoglobulin molecule.
[0170] As used herein, the term "substantially" in the context of a CDR refers to a CDR having an amino acid sequence at least 90%, at least 95%, at least 98% or at least 99% identical to the amino acid sequence of a non-human antibody CDR. A humanized antibody comprises substantially all of at least one, and typically two, variable domains (Fab, Fab', F(ab')2, FabC, Fv) in which all or substantially all of the CDR regions correspond to those of a non-human immunoglobulin (i.e., donor antibody) and all or substantially all of the framework regions are those of a human immunoglobulin consensus sequence. According to one aspect, a humanized antibody also comprises at least a portion of an immunoglobulin constant region (Fc), typically that of a human immunoglobulin. In some embodiments, a humanized antibody contains both the light chain as well as at least the variable domain of a heavy chain. The antibody also may include the CH1, hinge, CH2, CH3, and CH4 regions of the heavy chain. In some embodiments, a humanized antibody only contains a humanized light chain. In some embodiments, a humanized antibody only contains a humanized heavy chain. In specific embodiments, a humanized antibody only contains a humanized variable domain of a light chain and/or of a heavy chain.
[0171] The humanized antibody can be selected from any class of immunoglobulins, including IgM, IgG, IgD, IgA and IgE, and any isotype, including without limitation IgG 1, IgG2, IgG3, and IgG4. The humanized antibody may comprise sequences from more than one class or isotype, and particular constant domains may be selected to optimize desired effector functions using techniques well-known in the art.
[0172] The framework and CDR regions of a humanized antibody need not correspond precisely to the parental sequences, e.g., the donor antibody CDR or the consensus framework may be mutagenized by substitution, insertion and/or deletion of at least one amino acid residue so that the CDR or framework residue at that site does not correspond to either the donor antibody or the consensus framework. In one embodiment, such mutations, however, will not be extensive. Usually, at least 90%, at least 95%, at least 98%, or at least 99% of the humanized antibody residues will correspond to those of the parental FR and CDR sequences. As used herein, the term "consensus framework" refers to the framework region in the consensus immunoglobulin sequence. As used herein, the term "consensus immunoglobulin sequence" refers to the sequence formed from the most frequently occurring amino acids (or nucleotides) in a family of related immunoglobulin sequences (See e.g., Winnaker, From Genes to Clones (Verlagsgesellschaft, Weinheim, Germany 1987)). In a family of immunoglobulins, each position in the consensus sequence is occupied by the amino acid occurring most frequently at that position in the family. If two amino acids occur equally frequently, either can be included in the consensus sequence.
[0173] The humanized antibody may be designed to minimize unwanted immunological response toward rodent anti-human antibodies, which limits the duration and effectiveness of therapeutic applications of those moieties in human recipients. The humanized antibody may have one or more amino acid residues introduced into it from a source that is non-human. These non-human residues are often referred to as "import" residues, which are typically taken from a variable domain. Humanization may be performed by substituting hypervariable region sequences for the corresponding sequences of a human antibody. Accordingly, such "humanized" antibodies are chimeric antibodies wherein substantially less than an intact human variable domain has been substituted by the corresponding sequence from a non-human species. For example, see U.S. Pat. No. 4,816,567, the contents of which are herein incorporated by reference. The humanized antibody may be a human antibody in which some hypervariable region residues, and possibly some FR residues are substituted by residues from analogous sites in rodent antibodies. Humanization or engineering of antibodies of the present invention can be performed using any known method, such as but not limited to those described in U.S. Pat. Nos. 5,723,323; 5,976,862; 5,824,514; 5,817,483; 5,814,476; 5,763,192; 5,723,323; 5,766,886; 5,714,352; 6,204,023; 6,180,370; 5,693,762; 5,530,101; 5,585,089; 5,225,539; and 4,816,567.
[0174] The humanized antibody may retain high affinity for CFH and other favorable biological properties. The humanized antibody may be prepared by a process of analysis of the parental sequences and various conceptual humanized products using three-dimensional models of the parental and humanized sequences. Three-dimensional immunoglobulin models are commonly available. Computer programs are available that illustrate and display probable three-dimensional conformational structures of selected candidate immunoglobulin sequences. Inspection of these displays permits analysis of the likely role of the residues in the functioning of the candidate immunoglobulin sequence, i.e., the analysis of residues that influence the ability of the candidate immunoglobulin to bind its antigen. In this way, FR residues can be selected and combined from the recipient and import sequences so that the desired antibody characteristics, such as increased affinity for CFH, is achieved. In general, the hypervariable region residues may be directly and most substantially involved in influencing antigen binding.
[0175] As an alternative to humanization, human antibodies (also referred to herein as "fully human antibodies") can be generated. For example, it is possible to isolate human antibodies from libraries via PROfusion and/or yeast related technologies. It is also possible to isolate antibody producing B cells from patients producing a relevant antibody, sequence, and then clone the immunoglobulin. It is also possible to produce transgenic animals (e.g. mice that are capable, upon immunization, of producing a full repertoire of human antibodies in the absence of endogenous immunoglobulin production. For example, the homozygous deletion of the antibody heavy-chain joining region (J.sub.H) gene in chimeric and germ-line mutant mice results in complete inhibition of endogenous antibody production. Transfer of the human germ-line immunoglobulin gene array in such germ-line mutant mice will result in the production of human antibodies upon antigen challenge. The humanized or fully human antibodies may be prepared according to the methods described in U.S. Pat. Nos. 5,770,429; 5,833,985; 5,837,243; 5,922,845; 6,017,517; 6,096,311; 6,111,166; 6,270,765; 6,303,755; 6,365,116; 6,410,690; 6,682,928; and 6,984,720, the contents each of which are herein incorporated by reference.
3. PHARMACEUTICAL COMPOSITIONS
[0176] The CFH antibody may be a component in a pharmaceutical composition. The pharmaceutical composition may also contain a pharmaceutically acceptable carrier. The pharmaceutical compositions comprising a CFH antibody of the invention are for use in, but not limited to, diagnosing, detecting, or monitoring a disorder, in preventing, treating, managing, or ameliorating of a disorder, or one or more symptoms thereof, and/or in research. In a specific embodiment, a composition comprises one or more CFH antibodies of the invention. In another embodiment, the pharmaceutical composition comprises one or more CFH antibodies of the invention and one or more prophylactic or therapeutic agents other than CFH antibodies of the invention for treating a disorder in which activity of a targeted CFH is detrimental. In a further embodiment, the prophylactic or therapeutic agents are known to be useful for, or have been, or are currently being used in the prevention, treatment, management, or amelioration of a disorder, or one or more symptoms thereof In accordance with these embodiments, the composition may further comprise of a carrier, diluent, or excipient.
[0177] The CFH antibody of the invention can be incorporated into pharmaceutical compositions suitable for administration to a subject. Typically, the pharmaceutical composition comprises a CFH antibody of the invention and a pharmaceutically acceptable carrier. As used herein, "pharmaceutically acceptable carrier" includes any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents, and the like that are physiologically compatible. Examples of pharmaceutically acceptable carriers include one or more of water, saline, phosphate buffered saline, dextrose, glycerol, ethanol and the like, as well as combinations thereof. In many cases, it will be preferable to include isotonic agents, for example, sugars, polyalcohols such as mannitol, sorbitol, or sodium chloride in the composition. Pharmaceutically acceptable carriers may further comprise minor amounts of auxiliary substances such as wetting or emulsifying agents, preservatives, or buffers, which enhance the shelf life or effectiveness of the CFH antibody.
[0178] Various delivery systems are known and can be used to administer one or more CFH antibodies of the invention or the combination of one or more CFH antibodies of the invention and a prophylactic agent or therapeutic agent useful for preventing, managing, treating, or ameliorating a disorder or one or more symptoms thereof, e.g., encapsulation in liposomes, microparticles, microcapsules, recombinant cells capable of expressing the CFH antibody, receptor-mediated endocytosis (see, e.g., Wu and Wu, J. Biol. Chem. 262:4429-4432 (1987)), construction of a nucleic acid as part of a retroviral or other vector, etc. Methods of administering a prophylactic or therapeutic agent of the invention include, but are not limited to, parenteral administration (e.g., intradermal, intramuscular, intraperitoneal, intravenous and subcutaneous), epidural administration, intratumoral administration, and mucosal administration (e.g., intranasal and oral routes). In addition, pulmonary administration can be employed, e.g., by use of an inhaler or nebulizer, and formulation with an aerosolizing agent. See, e.g., U.S. Pat. Nos. 6,019,968; 5,985,320; 5,985,309; 5,934,272; 5,874,064; 5,855,913; 5,290,540; and 4,880,078; and PCT Publication Nos. WO 92/19244; WO97/32572; WO97/44013; WO98/31346; and WO99/66903, each of which is incorporated herein by reference in their entireties. In one embodiment, a CFH antibody of the invention or a composition of the invention is administered using Alkermes AIR.RTM. pulmonary drug delivery technology (Alkermes, Inc., Cambridge, Mass.). In a specific embodiment, prophylactic or therapeutic agents of the invention are administered intramuscularly, intravenously, intratumorally, orally, intranasally, pulmonary, or subcutaneously. The prophylactic or therapeutic agents may be administered by any convenient route, for example by infusion or bolus injection, by absorption through epithelial or mucocutaneous linings (e.g., oral mucosa, rectal and intestinal mucosa, etc.) and may be administered together with other biologically active agents. Administration can be systemic or local.
[0179] In a specific embodiment, it may be desirable to administer the CFH antibodies of the invention locally to the area in need of treatment; this may be achieved by, for example, and not by way of limitation, local infusion, by injection, or by means of an implant, said implant being of a porous or non-porous material, including membranes and matrices, such as sialastic membranes, polymers, fibrous matrices (e.g., Tissuel.RTM.), or collagen matrices. In one embodiment, an effective amount of one or more CFH antibodies of the invention is administered locally to the affected area to a subject to prevent, treat, manage, and/or ameliorate a disorder or a symptom thereof.
[0180] In another embodiment, the CFH antibodies can be delivered in a controlled release or sustained release system. In one embodiment, a pump may be used to achieve controlled or sustained release (see Langer, supra; Sefton, 1987, CRC Crit. Ref. Biomed. Eng. 14:20; Buchwald et al., 1980, Surgery 88:507; Saudek et al., 1989, N. Engl. J. Med. 321:574). In another embodiment, polymeric materials can be used to achieve controlled or sustained release of the therapies of the invention (see e.g., Medical Applications of Controlled Release, Langer and Wise (eds.), CRC Pres., Boca Raton, Fla. (1974); Controlled Drug Bioavailability, Drug Product Design and Performance, Smolen and Ball (eds.), Wiley, New York (1984); Ranger and Peppas, 1983, J., Macromol. Sci. Rev. Macromol. Chem. 23:61; see also Levy et al., 1985, Science 228:190; During et al., 1989, Ann. Neurol. 25:351; Howard et al., 1989, J. Neurosurg. 7 1:105); U.S. Pat. Nos. 5,679,377; 5,916,597; 5,912,015; 5,989,463; 5,128,326; PCT Publication No. WO99/15154; and PCT Publication No. WO99/20253. Examples of polymers used in sustained release formulations include, but are not limited to, poly(-hydroxy ethyl methacrylate), poly(methyl methacrylate), poly(acrylic acid), poly(ethylene-co-vinyl acetate), poly(methacrylic acid), polyglycolides (PLG), polyanhydrides, poly(N-vinyl pyrrolidone), poly(vinyl alcohol), polyacrylamide, poly(ethylene glycol), polylactides (PLA), poly(lactide-co-glycolides) (PLGA), and polyorthoesters. In a particular embodiment, the polymer used in a sustained release formulation is inert, free of leachable impurities, stable on storage, sterile, and biodegradable. In yet another embodiment, a controlled or sustained release system can be placed in proximity of the prophylactic or therapeutic target, thus requiring only a fraction of the systemic dose (see, e.g., Goodson, in Medical Applications of Controlled Release, supra, vol. 2, pp. 115-138 (1984)).
[0181] Controlled release systems are discussed in the review by Langer (1990, Science 249:1527-1533). Any technique known to one of skill in the art can be used to produce sustained release formulations comprising one or more CFH antibodies of the invention. See, e.g., U.S. Pat. No. 4,526, 938, PCT publication WO91/05548, PCT publication WO96/20698, Ning et al., 1996, "Intratumoral Radioimmunotherapy of a Human Colon Cancer Xenograft Using a Sustained-Release Gel," Radiotherapy &Oncology 39:179-189; Song et al., 1995, "Antibody Mediated Lung Targeting of Long-Circulating Emulsions," PDA Journal of Pharmaceutical Science & Technology 50:372-397; Cleek et al., 1997, "Biodegradable Polymeric Carriers for a bFGF Antibody for Cardiovascular Application," Pro. Int'l. Symp. Control. Rel. Bioact. Mater. 24:853-854; and Lam et al., 1997, "Microencapsulation of Recombinant Humanized Monoclonal Antibody for Local Delivery," Proc. Int'l. Symp. Control Rel. Bioact. Mater. 24:759-760, each of which is incorporated herein by reference in their entireties.
[0182] A pharmaceutical composition of the invention is formulated to be compatible with its intended route of administration. Examples of routes of administration include, but are not limited to, parenteral, e.g., intravenous, intradermal, subcutaneous, oral, intranasal (e.g., inhalation), transdermal (e.g., topical), transmucosal, and rectal administration. In a specific embodiment, the composition is formulated in accordance with routine procedures as a pharmaceutical composition adapted for intravenous, subcutaneous, intramuscular, oral, intranasal, or topical administration to human beings. Typically, compositions for intravenous administration are solutions in sterile isotonic aqueous buffer. Where necessary, the composition may also include a solubilizing agent and a local anesthetic such as lignocaine to ease pain at the site of the injection.
[0183] If the compositions of the invention are to be administered topically, the compositions can be formulated in the form of an ointment, cream, transdermal patch, lotion, gel, shampoo, spray, aerosol, solution, emulsion, or other form well-known to one of skill in the art. See, e.g., Remington's Pharmaceutical Sciences and Introduction to Pharmaceutical Dosage Forms, 19th ed., Mack Pub. Co., Easton, Pa. (1995). For non-sprayable topical dosage forms, viscous to semi-solid or solid forms comprising a carrier or one or more excipients compatible with topical application and having a dynamic viscosity greater than water are typically employed. Suitable formulations include, without limitation, solutions, suspensions, emulsions, creams, ointments, powders, liniments, salves, and the like, which are, if desired, sterilized or mixed with auxiliary agents (e.g., preservatives, stabilizers, wetting agents, buffers, or salts) for influencing various properties, such as, for example, osmotic pressure. Other suitable topical dosage forms include sprayable aerosol preparations wherein the active ingredient, for example in combination with a solid or liquid inert carrier, is packaged in a mixture with a pressurized volatile (e.g., a gaseous propellant, such as freon) or in a squeeze bottle. Moisturizers or humectants can also be added to pharmaceutical compositions and dosage forms if desired. Examples of such additional ingredients are well-known in the art.
[0184] If the method of the invention comprises intranasal administration of a composition, the composition can be formulated in an aerosol form, spray, mist or in the form of drops. In particular, prophylactic or therapeutic agents for use according to the present invention can be conveniently delivered in the form of an aerosol spray presentation from pressurized packs or a nebulizer, with the use of a suitable propellant (e.g., dichlorodifluoromethane, trichlorofluoromethane, dichlorotetrafluoroethane, carbon dioxide or other suitable gas). In the case of a pressurized aerosol the dosage unit may be determined by providing a valve to deliver a metered amount. Capsules and cartridges (composed of, e.g., gelatin) for use in an inhaler or insufflator may be formulated containing a powder mix of the compound and a suitable powder base such as lactose or starch.
[0185] If the method of the invention comprises oral administration, compositions can be formulated orally in the form of tablets, capsules, cachets, gelcaps, solutions, suspensions, and the like. Tablets or capsules can be prepared by conventional means with pharmaceutically acceptable excipients such as binding agents (e.g., pregelatinised maize starch, polyvinylpyrrolidone, or hydroxypropyl methylcellulose); fillers (e.g., lactose, microcrystalline cellulose, or calcium hydrogen phosphate); lubricants (e.g., magnesium stearate, talc, or silica); disintegrants (e.g., potato starch or sodium starch glycolate); or wetting agents (e.g., sodium lauryl sulphate). The tablets may be coated by methods well-known in the art. Liquid preparations for oral administration may take the form of, but not limited to, solutions, syrups or suspensions, or they may be presented as a dry product for constitution with water or other suitable vehicle before use. Such liquid preparations may be prepared by conventional means with pharmaceutically acceptable additives such as suspending agents (e.g., sorbitol syrup, cellulose derivatives, or hydrogenated edible fats); emulsifying agents (e.g., lecithin or acacia); non-aqueous vehicles (e.g., almond oil, oily esters, ethyl alcohol, or fractionated vegetable oils); and preservatives (e.g., methyl or propyl-p-hydroxybenzoates or sorbic acid). The preparations may also contain buffer salts, flavoring, coloring, and sweetening agents as appropriate. Preparations for oral administration may be suitably formulated for slow release, controlled release, or sustained release of a prophylactic or therapeutic agent(s).
[0186] The method of the invention may comprise pulmonary administration, e.g., by use of an inhaler or nebulizer, of a composition formulated with an aerosolizing agent. See, e.g., U.S. Pat. Nos. 6,019, 968; 5,985, 320; 5, 985,309; 5,934,272; 5,874,064; 5,855,913; 5,290,540; and 4,880,078; and PCT Publication Nos. WO 92/19244; WO 97/32572; WO 97/44013; WO 98/31346; and WO 99/66903, each of which is incorporated herein by reference their entireties. In a specific embodiment, a CFH antibody of the invention and/or composition of the invention is administered using Alkermes AIR.RTM. pulmonary drug delivery technology (Alkermes, Inc., Cambridge, Mass.).
[0187] The method of the invention may comprise administration of a composition formulated for parenteral administration by injection (e.g., by bolus injection or continuous infusion). Formulations for injection may be presented in unit dosage form (e.g., in ampoules or in multi-dose containers) with an added preservative. The compositions may take such forms as suspensions, solutions or emulsions in oily or aqueous vehicles, and may contain formulatory agents such as suspending, stabilizing and/or dispersing agents. Alternatively, the active ingredient may be in powder form for constitution with a suitable vehicle (e.g., sterile pyrogen-free water) before use. The methods of the invention may additionally comprise of administration of compositions formulated as depot preparations. Such long acting formulations may be administered by implantation (e.g., subcutaneously or intramuscularly) or by intramuscular injection. Thus, for example, the compositions may be formulated with suitable polymeric or hydrophobic materials (e.g., as an emulsion in an acceptable oil) or ion exchange resins, or as sparingly soluble derivatives (e.g., as a sparingly soluble salt).
[0188] The methods of the invention encompass administration of compositions formulated as neutral or salt forms. Pharmaceutically acceptable salts include those formed with anions such as those derived from hydrochloric, phosphoric, acetic, oxalic, tartaric acids, etc., and those formed with cations such as those derived from sodium, potassium, ammonium, calcium, ferric hydroxides, isopropylamine, triethylamine, 2-ethylamino ethanol, histidine, procaine, etc.
[0189] Generally, the ingredients of compositions are supplied either separately or mixed together in unit dosage form, for example, as a dry lyophilized powder or water free concentrate in a hermetically sealed container such as an ampoule or sachette indicating the quantity of active agent. Where the mode of administration is infusion, composition can be dispensed with an infusion bottle containing sterile pharmaceutical grade water or saline. Where the mode of administration is by injection, an ampoule of sterile water for injection or saline can be provided so that the ingredients may be mixed prior to administration.
[0190] In particular, the invention also provides that one or more of the CFH antibodies, or pharmaceutical compositions, of the invention is packaged in a hermetically sealed container such as an ampoule or sachette indicating the quantity of the CFH antibody. In one embodiment, one or more of the CFH antibodies, or pharmaceutical compositions of the invention is supplied as a dry sterilized lyophilized powder or water free concentrate in a hermetically sealed container and can be reconstituted (e.g., with water or saline) to the appropriate concentration for administration to a subject. In one embodiment, one or more of the CFH antibodies or pharmaceutical compositions of the invention is supplied as a dry sterile lyophilized powder in a hermetically sealed container at a unit dosage of at least 5 mg, for example at least 10 mg, at least 15 mg, at least 25 mg, at least 35 mg, at least 45 mg, at least 50 mg, at least 75 mg, or at least 100 mg. The lyophilized CFH antibodies or pharmaceutical compositions of the invention should be stored at between 2.degree. C. and 8.degree. C. in its original container and the CFH antibodies, or pharmaceutical compositions of the invention should be administered within 1 week, for example within 5 days, within 72 hours, within 48 hours, within 24 hours, within 12 hours, within 6 hours, within 5 hours, within 3 hours, or within 1 hour after being reconstituted. In an alternative embodiment, one or more of the CFH antibodies or pharmaceutical compositions of the invention is supplied in liquid form in a hermetically sealed container indicating the quantity and concentration of the CFH antibody. In a further embodiment, the liquid form of the administered composition is supplied in a hermetically sealed container at least 0.25 mg/ml, for example at least 0.5 mg/ml, at least 1 mg/ml, at least 2.5 mg/ml, at least 5 mg/ml, at least 8 mg/ml, at least 10 mg/ml, at least 15 mg/ml, at least 25 mg/ml, at least 50 mg/ml, at least 75 mg/ml or at least 100 mg/ml. The liquid form should be stored at between 2.degree. C. and 8.degree. C. in its original container.
[0191] The CFH antibodies of the invention can be incorporated into a pharmaceutical composition suitable for parenteral administration. In one aspect, CFH antibodies will be prepared as an injectable solution containing 0.1-250 mg/ml CFH antibody. The injectable solution can be composed of either a liquid or lyophilized dosage form in a flint or amber vial, ampule or pre-filled syringe. The buffer can be L-histidine (1-50 mM), optimally 5-10 mM, at pH 5.0 to 7.0 (optimally pH 6.0). Other suitable buffers include but are not limited to, sodium succinate, sodium citrate, sodium phosphate or potassium phosphate. Sodium chloride can be used to modify the tonicity of the solution at a concentration of 0-300 mM (optimally 150 mM for a liquid dosage form). Cryoprotectants can be included for a lyophilized dosage form, principally 0-10% sucrose (optimally 0.5-1.0%). Other suitable cryoprotectants include trehalose and lactose. Bulking agents can be included for a lyophilized dosage form, principally 1-10% mannitol (optimally 2-4%). Stabilizers can be used in both liquid and lyophilized dosage forms, principally 1-50 mM L-Methionine (optimally 5-10 mM). Other suitable bulking agents include glycine, arginine, can be included as 0-0.05% polysorbate-80 (optimally 0.005-0.01%). Additional surfactants include but are not limited to polysorbate 20 and BRIJ surfactants. The pharmaceutical composition comprising the CFH antibodies of the invention prepared as an injectable solution for parenteral administration, can further comprise an agent useful as an adjuvant, such as those used to increase the absorption, or dispersion of the CFH antibody. A particularly useful adjuvant is hyaluronidase, such as Hylenex.RTM. (recombinant human hyaluronidase). Addition of hyaluronidase in the injectable solution improves human bioavailability following parenteral administration, particularly subcutaneous administration. It also allows for greater injection site volumes (i.e. greater than 1 ml) with less pain and discomfort, and minimum incidence of injection site reactions. (See International Appin. Publication No. WO 04/078140 and U.S. Patent Appin. Publication No. US2006104968, incorporated herein by reference.)
[0192] The compositions of this invention may be in a variety of forms. These include, for example, liquid, semi-solid and solid dosage forms, such as liquid solutions (e.g., injectable and infusible solutions), dispersions or suspensions, tablets, pills, powders, liposomes and suppositories. The preferred form depends on the intended mode of administration and therapeutic application. Compositions can be in the form of injectable or infusible solutions, such as compositions similar to those used for passive immunization of humans with other CFH antibodies. In one embodiment, the CFH antibody is administered by intravenous infusion or injection. In another embodiment, the CFH antibody is administered by intramuscular or subcutaneous injection.
[0193] Therapeutic compositions typically must be sterile and stable under the conditions of manufacture and storage. The composition can be formulated as a solution, microemulsion, dispersion, liposome, or other ordered structure suitable to high drug concentration. Sterile injectable solutions can be prepared by incorporating the active compound (i.e., a CFH antibody of the present invention) in the required amount in an appropriate solvent with one or a combination of ingredients enumerated above, as required, followed by filtered sterilization. Generally, dispersions are prepared by incorporating the active compound into a sterile vehicle that contains a basic dispersion medium and the required other ingredients from those enumerated above. In the case of sterile, lyophilized powders for the preparation of sterile injectable solutions, methods of preparation comprise vacuum drying and spray-drying that yields a powder of the active ingredient plus any additional desired ingredient from a previously sterile-filtered solution thereof. The proper fluidity of a solution can be maintained, for example, by the use of a coating such as lecithin, by the maintenance of the required particle size in the case of dispersion and by the use of surfactants. Prolonged absorption of injectable compositions can be brought about by including, in the composition, an agent that delays absorption, for example, monostearate salts and gelatin.
[0194] The CFH antibodies of the present invention can be administered by a variety of methods known in the art. For many therapeutic applications, the route/mode of administration may be subcutaneous injection, intravenous injection or infusion. As will be appreciated by the skilled artisan, the route and/or mode of administration will vary depending upon the desired results. In certain embodiments, the active compound may be prepared with a carrier that will protect the compound against rapid release, such as a controlled release formulation, including implants, transdermal patches, and microencapsulated delivery systems. Biodegradable, biocompatible polymers can be used, such as ethylene vinyl acetate, polyanhydrides, polyglycolic acid, collagen, polyorthoesters, and polylactic acid. Many methods for the preparation of such formulations are patented or generally known to those skilled in the art. See, e.g., Sustained and Controlled Release Drug Delivery Systems, J. R. Robinson, ed., Marcel Dekker, Inc., New York, 1978.
[0195] In certain embodiments, a CFH antibody of the invention may be orally administered, for example, with an inert diluent or an assimilable edible carrier. The CFH antibody (and other ingredients, if desired) may also be enclosed in a hard or soft shell gelatin capsule, compressed into tablets, or incorporated directly into the subject's diet. For oral therapeutic administration, the CFH antibody may be incorporated with excipients and used in the form of ingestible tablets, buccal tablets, troches, capsules, elixirs, suspensions, syrups, wafers, and the like. To administer a CFH antibody of the invention by other than parenteral administration, it may be necessary to coat the CFH antibody with, or co-administer the CFH antibody with, a material to prevent its inactivation.
[0196] In certain embodiments, a CFH antibody of the invention is linked to a half-life extending vehicle known in the art. Such vehicles include, but are not limited to, the Fc domain, polyethylene glycol, and dextran. Such vehicles are described, e.g., in U.S. application Ser. No. 09/428,082 and published PCT Application No. WO 99/25044, which are hereby incorporated by reference for any purpose.
[0197] The pharmaceutical compositions may include a "therapeutically effective amount" or a "prophylactically effective amount" of a CFH antibody. A "therapeutically effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired therapeutic result. A therapeutically effective amount of the CFH antibody may be determined by a person skilled in the art and may vary according to factors such as the disease state, age, sex, and weight of the individual, and the ability of the CFH antibody to elicit a desired response in the individual. A therapeutically effective amount is also one in which toxic or detrimental effects, if any, of the CFH antibody are outweighed by the therapeutically beneficial effects. A "prophylactically effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired prophylactic result. Typically, since a prophylactic dose is used in subjects prior to or at an earlier stage of disease, the prophylactically effective amount will be less than the therapeutically effective amount.
[0198] Dosage regimens may be adjusted to provide the optimum desired response (e.g., a therapeutic or prophylactic response). For example, a single bolus may be administered, several divided doses may be administered over time or the dose may be proportionally reduced or increased as indicated by the exigencies of the therapeutic situation. It is especially advantageous to formulate parenteral compositions in dosage unit form for ease of administration and uniformity of dosage. Dosage unit form as used herein refers to physically discrete units suited as unitary dosages for the mammalian subjects to be treated; each unit containing a predetermined quantity of active compound calculated to produce the desired therapeutic effect in association with the required pharmaceutical carrier. The specification for the dosage unit forms are dictated by and directly dependent on (a) the unique characteristics of the active compound and the particular therapeutic or prophylactic effect to be achieved, and (b) the limitations inherent in the art of compounding such an active compound for the treatment of sensitivity in individuals.
[0199] An exemplary, non-limiting range for a therapeutically or prophylactically effective amount of the CFH antibody is a dose of between 0.1 and 200 mg/kg, for example between 0.1 and 10 mg/kg. The therapeutically or prophylactically effective amount of the CFH antibody may be between 1 and 200 mg/kg, 10 and 200 mg/kg, 20 and 200 mg/kg, 50 and 200 mg/kg, 75 and 200 mg/kg, 100 and 200 mg/kg, 150 and 200 mg/kg, 50 and 100 mg/kg, 5 and 10 mg/kg, or 1 and 10 mg/kg. It is to be noted that dosage values may vary with the type and severity of the condition to be alleviated. Further, the CFH antibody dose may be determined by a person skilled in the art and may vary according to factors such as the disease state, age, sex, and weight of the individual, and the ability of the CFH antibody to elicit a desired response in the individual. The dose is also one in which toxic or detrimental effects, if any, of the CFH antibody are outweighed by the therapeutically beneficial effects. It is to be further understood that for any particular subject, specific dosage regimens should be adjusted over time according to the individual need and the professional judgment of the person administering or supervising the administration of the compositions, and that dosage ranges set forth herein are exemplary only and are not intended to limit the scope or practice of the claimed composition.
4. METHODS OF TREATING
[0200] Provided herein is a method of treating cancer in a subject. The method may include administering to the subject in need thereof an anti-CFH antibody described above. The CFH antibody may be administered in a therapeutically effective amount.
[0201] In general, the dosage of administered CFH antibody will vary depending upon such factors as the patient's age, weight, height, sex, general medical condition and previous medical history. Typically, it is desirable to provide the recipient with a dosage of CFH antibody component, immunoconjugate or fusion protein which is in the range of from about 1 pg/kg to 10 mg/kg (amount of agent/body weight of patient), although a lower or higher dosage also may be administered as circumstances dictate. Dosage regimens may be adjusted to provide the optimum desired response (e.g., a therapeutic or prophylactic response). For example, a single bolus may be administered, several divided doses may be administered over time or the dose may be proportionally reduced or increased as indicated by the exigencies of the therapeutic situation. It is especially advantageous to formulate parenteral compositions in dosage unit form for ease of administration and uniformity of dosage. Dosage unit form as used herein refers to physically discrete units suited as unitary dosages for the mammalian subjects to be tested; each unit containing a predetermined quantity of active compound calculated to produce the desired therapeutic effect in association with the required pharmaceutical carrier. The specification for the dosage unit forms of the present invention are dictated by and directly dependent on (a) the unique characteristics of the active compound and the particular therapeutic or prophylactic effect to be achieved and (b) the limitations inherent in the art of compounding such an active compound for the treatment of sensitivity in individuals.
[0202] An exemplary, non-limiting range for a therapeutically or prophylactically effective amount of a CFH antibody of the invention is 0.1-20 mg/kg, more preferably 0.5-10 mg/kg. It is to be noted that dosage values may vary with the type and severity of the condition to be alleviated. It is to be further understood that for any particular subject, specific dosage regimens should be adjusted over time according to the individual need and the professional judgment of the person administering or supervising the administration of the compositions, and that dosage ranges set forth herein are exemplary only and are not intended to limit the scope or practice of the claimed composition.
[0203] Administration of CFH antibodies to a patient can be intravenous, intraarterial, intraperitoneal, intramuscular, subcutaneous, intrapleural, intrathecal, intraocular, intravitreal, by perfusion through a regional catheter, or by direct intralesional injection. When administering therapeutic proteins by injection, the administration may be by continuous infusion or by single or multiple boluses. Intravenous injection provides a useful mode of administration due to the thoroughness of the circulation in rapidly distributing CFH antibodies. The CFH antibody may be administered orally, for example, with an inert diluent or an assimilable edible carrier. The antibody and other ingredients, if desired, may be enclosed in a hard or soft shell gelatin capsule, compressed into tablets, buccal tablets, troches, capsules, elixirs, suspensions, syrups, wafers, and the like.
[0204] CFH antibodies may be administered at low protein doses, such as 20 milligrams to 2 grams protein per dose, given once, or repeatedly, parenterally. Alternatively, the CFH antibodies may be administered in doses of 20 to 1000 milligrams protein per dose, or 20 to 500 milligrams protein per dose, or 20 to 100 milligrams protein per dose.
[0205] The CFH antibodies may be administered alone or they may be conjugated to liposomes, and can be formulated according to known methods to prepare pharmaceutically useful compositions, whereby the CFH antibodies are combined in a mixture with a pharmaceutically acceptable carrier. A "pharmaceutically acceptable carrier" may be tolerated by a recipient patient. Sterile phosphate-buffered saline is one example of a pharmaceutically acceptable carrier. Other suitable carriers are well known to those in the art. See, for example, REMINGTON'S PHARMACEUTICAL SCIENCES, 19th Ed. (1995).
[0206] For purposes of therapy, CFH antibodies are administered to a patient in a therapeutically effective amount in a pharmaceutically acceptable carrier. A "therapeutically effective amount" is one that is physiologically significant. The CFH antibody is physiologically significant if its presence results in a detectable change in the physiology of a recipient patient. In the present context, the CFH antibodies may be physiologically significant if its presence results in, for example, increased complement dependent lysis of a cell, increased C3b deposition on a cell, and/or inhibition of CFH binding to C3b.
[0207] Additional treatment methods may be employed to control the duration of action of an antibody in a therapeutic application. Control release preparations can be prepared through the use of polymers to complex or adsorb the antibody. For example, biocompatible polymers include matrices of poly(ethylene-co-vinyl acetate) and matrices of a polyanhydride copolymer of a stearic acid dimer and sebacic acid. Sherwood et al., Bio/Technology 10:1446 (1992). The rate of release of an antibody from such a matrix depends upon the molecular weight of the protein, the amount of antibody within the matrix, and the size of dispersed particles. Saltzman et al., Biophys. J. 55:163 (1989); Sherwood et al., supra. Other solid dosage forms are described in REMINGTON'S PHARMACEUTICAL SCIENCES, 19th ed. (1995).
[0208] a. CFH Antibodies
[0209] The CFH antibodies described herein may interfere with CFH binding to tumor cells, such as lung cancer cells, and may be used to treat cancer in a subject. The interference of CFH binding to the tumor cells decreases the number of CFH on the tumor cells and enhances complement-dependent lysis of the tumor cells. The CFH antibodies may cause an increase in the deposition of C3b on lung cancer cells. C3b deposition is required for complement dependent cytotoxicity (CDC) and is frequently used as evidence for complement activation.
[0210] An effective amount of the CFH antibody or fragment thereof may be administered to the cell. For example, an effective amount between about 1 .mu.g/mL to about 250 .mu.g/mL, between about 10 .mu.g/mL to about 250 .mu.g/mL, between about 25 .mu.g/mL to about 250 .mu.g/mL, between about 40 .mu.g/mL to about 250 .mu.g/mL, between about 45 .mu.g/mL to about 250 .mu.g/mL, between about 50 .mu.g/mL to about 250 .mu.g/mL, between about 60 .mu.g/mL to about 250 .mu.g/mL, between about 75 .mu.g/mL to about 250 .mu.g/mL, between about 100 .mu.g/mL to about 250 .mu.g/mL, between about 10 .mu.g/mL to about 200 .mu.g/mL, between about 25 .mu.g/mL to about 200 .mu.g/mL, between about 40 .mu.g/mL to about 200 .mu.g/mL, between about 45 .mu.g/mL to about 200 .mu.g/mL, between about 50 .mu.g/mL to about 200 .mu.g/mL, between about 60 .mu.g/mL to about 200 .mu.g/mL, between about 75 .mu.g/mL to about 200 .mu.g/mL, between about 100 .mu.g/mL to about 200 .mu.g/mL, between about 10 .mu.g/mL to about 150 .mu.g/mL, between about 25 .mu.g/mL to about 150 .mu.g/mL, between about 40 .mu.g/mL to about 150 .mu.g/mL, between about 45 .mu.g/mL to about 150 .mu.g/mL, between about 50 .mu.g/mL to about 150 .mu.g/mL, between about 60 .mu.g/mL to about 150 .mu.g/mL, between about 75 .mu.g/mL to about 150 .mu.g/mL, between about 100 .mu.g/mL to about 150 .mu.g/mL, between about 10 .mu.g/mL to about 120 .mu.g/mL, between about 25 .mu.g/mL to about 120 .mu.g/mL, between about 40 .mu.g/mL to about 120 .mu.g/mL, between about 45 .mu.g/mL to about 120 .mu.g/mL, between about 50 .mu.g/mL to about 120 .mu.g/mL, between about 60 .mu.g/mL to about 120 .mu.g/mL, between about 75 .mu.g/mL to about 120 .mu.g/mL, between about 100 .mu.g/mL to about 120 .mu.g/mL, between about 10 .mu.g/mL to about 100 .mu.g/mL, between about 25 .mu.g/mL to about 100 .mu.g/mL, between about 40 .mu.g/mL to about 100 .mu.g/mL, between about 45 .mu.g/mL to about 100 .mu.g/mL, between about 50 .mu.g/mL to about 100 .mu.g/mL, between about 60 .mu.g/mL to about 100 .mu.g/mL, or between about 75 .mu.g/mL to about 100 .mu.g/mL of the CFH antibody or fragment thereof may be administered to the cell. The CFH antibodies may be Ab7960/293i or Ab7968.
[0211] b. Cancer
[0212] The method described herein may be used to treat a subject having any form of cancer. The method may include administering to the subject in need thereof an anti-CFH antibody, as described above. The cancer may be any cancer that uses CFH as a protective mechanism. The cancer may be Adrenocortical Carcinoma, Anal Cancer, Bladder Cancer, Brain Tumor, Breast Cancer, Carcinoid Tumor, Gastrointestinal, Carcinoma of Unknown Primary, Cervical Cancer, Colon Cancer, Endometrial Cancer, Esophageal Cancer, Extrahepatic Bile Duct Cancer, Ewings Family of Tumors (PNET), Extracranial Germ Cell Tumor, Intraocular Melanoma Eye Cancer, Gallbladder Cancer, Gastric Cancer (Stomach), Extragonadal Germ Cell Tumor, Gestational Trophoblastic Tumor, Head and Neck Cancer, Hypopharyngeal Cancer, Islet Cell Carcinoma, Kidney Cancer (renal cell cancer), Laryngeal Cancer, Acute Lymphoblastic Leukemia, Leukemia, Acute Myeloid, Chronic Lymphocytic Leukemia, Chronic Myelogenous Leukemia, Hairy Cell Leukemia, Lip and Oral Cavity Cancer, Liver Cancer, Non-Small Cell Lung Cancer, Small Cell Lung Cancer, AIDS-Related Lymphoma, Central Nervous System (Primary) Lymphoma, Cutaneous T-Cell Lymphoma, Hodgkin's Disease Lymphoma, Non-Hodgkin's Disease Lymphoma, Malignant Mesothelioma, Melanoma, Merkel Cell Carcinoma, Metasatic Squamous Neck Cancer with Occult Primary, Multiple Myeloma and Other Plasma Cell Neoplasms, Mycosis Fungoides, Myelodysplastic Syndrome, Myeloproliferative Disorders, Nasopharyngeal Cancer, euroblastoma, Oral Cancer, Oropharyngeal Cancer, Osteosarcoma, Ovarian Epithelial Cancer, Ovarian Germ Cell Tumor, Pancreatic Cancer, Exocrine, Pancreatic Cancer, Islet Cell Carcinoma, Paranasal Sinus and Nasal Cavity Cancer, Parathyroid Cancer, Penile Cancer, Pituitary Cancer, Plasma Cell Neoplasm, Prostate Cancer, Rhabdomyosarcoma, Rectal Cancer, Renal Cell Cancer (cancer of the kidney), Transitional Cell Renal Pelvis and Ureter, Salivary Gland Cancer, Sezary Syndrome, Skin Cancer, Small Intestine Cancer, Soft Tissue Sarcoma, Testicular Cancer, Malignant Thymoma, Thyroid Cancer, Urethral Cancer, Uterine Cancer, Unusual Cancer of Childhood, Vaginal Cancer, Vulvar Cancer, and Wilms' Tumor.
[0213] (1) Lung Cancer
[0214] The method described herein can be used to treat a subject having lung cancer. The method may include administering to the subject in need thereof an anti-CFH antibody, as described above. The lung cancer may be small-cell lung cancer, also known as small-cell lung carcinoma and oat cell cancer, non-small-cell lung carcinoma ("NSCLC"), glandular tumors, carcinoid tumors and/or undifferentiated carcinomas.
[0215] (2) Breast Cancer
[0216] The method described herein can be used to treat a subject having breast cancer. The method may include administering to the subject in need thereof an anti-CFH antibody, as described above. Breast cancer may be any cancer that starts in the tissues of the breast. The two main types of breast cancer are ductal carcinoma, which starts in the tubes (ducts) that move milk from the breast to the nipple, and lobular carcinoma, which starts in the parts of the breast, called lobules, that produce milk. Breast cancer may also start in other areas of the breast. Breast cancer may be invasive or noninvasive (in situ).
[0217] c. Combination Therapy
[0218] The methods described above may include a combination treatment of the CFH antibody with other drugs and/or other conventional cancer therapies.
[0219] (1) Combination Drugs
[0220] The methods may further include administering an effective amount of at least one anti-cancer compound or chemotherapeutic agent. The CFH antibodies may be used in conjunction with an anti-cancer drug or chemotherapeutic agent to increase tumor cell killing, i.e., enhance antibody-dependent cell-mediated cytotoxicity (ADCC) and cell mediated toxicity. Examples of anti-cancer compounds and chemotherapeutic agents include anthracyclines, such as doxorubicin (Adriamycin, Doxil), epirubicin (Ellence), and daunorubicin (Cerubidine, DaunoXome), capecitabine (Xeloda), carboplatin (Paraplatin), cisplatin, cyclophosphamide (Cytoxan), eribulin (Halaven), fluorouracil (also called 5-fluorouracil or 5-FU; Adrucil), gemcitabine (Gemzar), ixabepilone (Ixempra), methotrexate (Amethopterin, Mexate, Folex), mitoxantrone (Novantrone), mutamycin (Mitomycin), taxanes, such as paclitaxel (Taxol, Abraxane), and docetaxel (Taxotere), thiotepa (Thioplex), vincristine (Oncovin, Vincasar PES, Vincrex), and vinorelbine (Navelbine). Examples of targeted therapy include trastuzumab (Herceptin), lapatinib (Tykerb), onartuzumab, rilotumumab (AMG102), ficlatuzumab (AV-299), bevacizumab (Avastin), pertuzumab (Perjeta), Rituximab, panatumamab, and everolimus (Afinitor). The CFH antibodies may be used in conjunction with Cetuximab, Perjeta, and Herceptin.
[0221] (2) Conventional Cancer Therapies
[0222] Conventional cancer therapies may include surgery, radiation therapy, hormone therapy, and targeted therapy. Examples of surgery include open craniotomy with maximal excision, which may be followed by radiation therapy. Examples of radiation therapy include whole-brain irradiation, fractionated radiotherapy, and radiosurgery, such as stereotactic radiosurgery, e.g., Gamma Knife radiosurgery.
[0223] d. Subject
[0224] The subject may be a mammal, which may be a human. The subject may have, or be at risk of developing a cancer. The subject may have cancer. The subject may already be undergoing treatment for a cancer.
5. METHODS OF INCREASING COMPLEMENT DEPENDENT LYSIS OF CELLS
[0225] The methods described herein can also be used to increase complement-dependent lysis of a cell. The method described herein may include administering to the cell an anti-CFH antibody, as described above. The cell may be a tumor cell. For example, the tumor cell may be MCF7 breast cancer cell, SKBR3 breast cancer cell, MDA-MB-231 breast cancer cell, or A549 lung carcinoma cell.
[0226] As disclosed below, purified CFH antibodies had a statistically significant effect on both C3 deposition on A549 lung carcinoma cells and cytotoxicity by the alternative pathway. It should be noted that lung tumor cells, as well as other types of tumor cells, are protected from complement attack by other membrane bound inhibitors including MCP (CD46), CR1 (CD35), and DAF (CD55) in addition to CFH. Efficiency of cytotoxicity could conceivably be increased by combining patient antibodies to CFH with monoclonal antibodies to these proteins (See Example 2).
6. METHODS OF INHIBITING COMPLEMENT FACTOR H BINDING TO C3B
[0227] The methods described herein can also be used to inhibit CFH binding to C3b in a subject or a cell. The method may include administering to the subject or the cell an anti-CFH antibody, as described above. The cell may be a tumor cell. For example, the tumor cell may be MCF7 breast cancer cell, SKBR3 breast cancer cell, MDA-MB-231 breast cancer cell, or A549 lung carcinoma cell.
7. METHODS OF INCREASING C3B DEPOSITION ON CELLS
[0228] The methods described herein can also be used to increase C3b deposition on a cell. The method may include administering to the subject or the cell an anti-CFH antibody, as described above. The cell may be a tumor cell. For example, the tumor cell may be MCF7 breast cancer cell, SKBR3 breast cancer cell, MDA-MB-231 breast cancer cell, or A549 lung carcinoma cell.
8. METHODS OF INHIBITING TUMOR GROWTH
[0229] The methods described herein can also be used to inhibit tumor growth in a subject. The method may include may include administering to the subject or the cell an anti-CFH antibody, as described above. The tumor may be a solid tumor or a hematologic malignancy. For example, the tumor may be a lung tumor.
9. MECHANISMS OF DELIVERY
[0230] The CFH antibody may be formulated to be compatible with its intended route of administration. Examples of routes of administration include, but are not limited to, parenteral, e.g., intravenous, intradermal, subcutaneous, oral, intranasal (e.g., inhalation), transdermal (e.g., topical), transmucosal, and rectal administration. In a specific embodiment, the CFH antibody is formulated in accordance with routine procedures as a pharmaceutical composition adapted for intravenous, subcutaneous, intramuscular, oral, intranasal, or topical administration to human beings. Typically, compositions for intravenous administration are solutions in sterile isotonic aqueous buffer. Where necessary, the composition may also include a solubilizing agent and a local anesthetic such as lignocaine to ease pain at the site of the injection.
[0231] Various delivery systems are known and can be used to administer one or more SERMs or the combination of one or more CFH antibodies and a prophylactic agent or therapeutic agent useful for preventing, managing, treating, or ameliorating a disorder or one or more symptoms thereof, e.g., encapsulation in liposomes, microparticles, microcapsules, receptor-mediated endocytosis (see, e.g., Wu and Wu, J. Biol. Chem. 262:4429-4432 (1987)), etc. Methods of administering a prophylactic or therapeutic agent of the SERM include, but are not limited to, parenteral administration (e.g., intradermal, intramuscular, intraperitoneal, intravenous and subcutaneous), epidural administration, intratumoral administration, and mucosal administration (e.g., intranasal and oral routes).
10. CELL TYPES
[0232] The methods described herein may be utilized with a cell from a sample or subject. The cell may be a tumor or cancer cell. The cell may be a breast cancer cell or a lung cancer cell. For example, the cell may be MCF7 breast cancer cell, SKBR3 breast cancer cell, MDA-MB-231 breast cancer cell, A549 lung carcinoma cell, DMS79, or H226 cell lines.
11. KIT
[0233] Provided herein is a kit, which may be used for using the CFH antibodies. The kit comprises at least one component for using the CFH antibodiesand instructions for using the CFH antibodies. Instructions included in kits can be affixed to packaging material or can be included as a package insert. While the instructions are typically written or printed materials they are not limited to such. Any medium capable of storing such instructions and communicating them to an end user is contemplated by this disclosure. Such media include, but are not limited to, electronic storage media (e.g., magnetic discs, tapes, cartridges, chips), optical media (e.g., CD ROM), and the like. As used herein, the term "instructions" can include the address of an internet site that provides the instructions.
[0234] The component may include at least one composition comprising one or more isolated antibodies or antibody fragments thereof that specifically bind to CFH or reduced form of CFH. The antibody may be a CFH or reduced form of CFH capture antibody and/or a CFH or reduced form of CFH detection antibody. Preferably, the kit comprises all components, i.e., reagents, standards, buffers, diluents, etc., which are necessary to perform the assay. The kit may also include other drugs for treating cancer.
12. EXAMPLES
[0235] The present invention has multiple aspects, illustrated by the following non-limiting examples.
Example 1
[0236] Cloning a human CFH mAb. Antibodies were generated to target the same epitope that is recognized by the CFH autoantibodies of cancer patients. CFH is a ubiquitous protein that binds to the surface of host cells and is particularly important in protecting the kidney. CFH is a multifunctional 150 kDa protein that is composed of 20 short consensus repeats (SCRs), each 60 amino acids long. The C-terminal SCR domains SCR19 and SCR20 bind to glycosaminoglycan and sialic acid polyanions, as well as to cell-bound C3b and its proteolytic fragments, on the mammalian cell surface. CFH antibodies were affinity purified from the sera of lung cancer patients and shown to bind an epitope within SCR19, PIDNGDIT (FIG. 1). This epitope comprises residues that are predicted on mutational and structural grounds to be critical for the CFH-C3b interaction. In vitro, the CFH autoantibodies prevented CFH binding to tumor cells (FIG. 2), increased C3b binding (FIG. 3), and promoted CDC (FIG. 4). Although CFH is abundant in the blood, it is notable that patients who have these antibodies show no apparent side-effects or off-target effects, and no kidney disease. This observation, and the fact that the autoantibodies bind preferentially to reduced over native CFH in vitro (FIG. 5), led to the discovery that the autoantibodies bind to a conformationally distinct form of CFH that may exist on tumor cells.
[0237] In order to clone human mAbs that recognize the same epitope recognized by the autoantibodies, the peripheral blood mononuclear cells (PBMCs) of 11 patients who were shown by immunoblot to express CFH autoantibody were pooled. Using a biotinylated CFH peptide containing the mapped 8 amino acid binding segment as bait, single memory B cells were sorted from the pooled PBMCs by flow cytometry for isolation of the immunoglobulin variable region of heavy and light gene segments (V.sub.HDJ.sub.H and V.sub.LJ.sub.L). V.sub.HDJ.sub.H and V.sub.LJ.sub.L gene pairs were amplified by RT/PCR from individual CFH antigen-specific memory B cells, isolated, inserted into expression vectors, and recombinant mAbs were generated.
[0238] The mAbs were sequenced and the CDR regions were determined using Paratome-Antigen Binding Regions Identification Tool ((Kunik et al. (2012) PLoS Comput Biol 8(2): e1002388. doi:10.1371/journal.pcbi.1002388; Kunik et al. (2012) Nucleic Acids Res. 2012 July; 40(Web Server issue):W521-4. doi: 10.1093/nar/gks480. Epub 2012 Jun. 6)). The variable heavy (VH) and variable light (VL) chain sequences as well as the CDR regions in each are shown in Table 1. The VH sequences, CDR1 sequences of the heavy chain (HCDR1), CDR2 sequences of the heavy chain (HCDR2), CDR3 of the heavy chain (HCDR3), VL sequences, CDR1 sequences of the light chain (LCDR1), CDR2 sequences of the light chain (LCDR2), and CDR3 sequences of the light chain (LCDR3) from each of the antibodies was aligned using Clustal Omega (McWilliam et al., Nucleic Acids Research (2013) 41(W1):W597-W600; Li et al., Nucleic Acids Research (2015) 43(W1):W580-W584; Sievers et al., Molecular Systems Biology (2011) 7:539). See FIGS. 17A and 18-24. The percent identities between the sequences are shown in FIGS. 17B and 18-24.
[0239] Specificity of the CFH mAbs. The recombinant CFH mAbs were demonstrated by western blot analysis to bind CFH with specificity for the conformationally distinct form, thus recapitulating the specificity of the autoantibodies identified in sera (FIG. 6). A mAb that has a low nM affinity for the epitope-containing peptide, CFH mAb7968 (also known as CBAb1), was chosen for further development. A co-crystal structure of a CFH mAb-peptide complex showed a conformational change in the peptide relative to the folding of that peptide sequence within the native structure of CFH (FIG. 7). In particular, residues 1117-1120 (sequence element NGDI) near the C-terminus of the peptide adopted an .alpha.-helical conformation in the antibody-bound complex, whereas the same region exhibited a .beta.-strand conformation in natively folded structures of CFH. The presence of the helical element was also consistent with alanine scanning results showing a discontinuous epitope for mAb7968 (FIG. 8). These findings supported the hypothesis that mAb7968 binds a conformationally distinct form of CFH. CFH mAb7968 does not bind to normal human tissues in a tissue microarray. In Biacore experiments, mAb7968 does not bind native CFH but does bind a peptide containing the epitope with an affinity of 2 nM. In ELISA format, CFH mAb7968 binds to a target peptide equally well in the absence or presence of a vast excess of full-length native CFH as might be found in the blood, indicating no competition by native CFH with the epitope peptide for the antibody. Taken together, these data suggest that the binding of CFH mAb7968 to CFH has specificity for a non-native form of the protein.
[0240] CDC of cancer cell lines by a human CFH mAb. CFH mAb7968 were tested in CDC assays in vitro using a variety of cancer cell lines. In this assay, cells are mixed with antibodies and normal human serum (NHS) as a source of complement, incubated at 37.degree. C., and cytolysis is measured by lactate dehydrogenase release. A significant increase in CDC over that seen with control antibody was observed with CFH mAb7968 for lung (three NSCLC and one small cell), breast, and gastric cancer cell lines (FIG. 9).
[0241] CFH mAb potentiation of CDC by other agents. Rituximab therapy for B cell chronic lymphocytic leukemia (B-CLL) has met with mixed success. Among several factors to which resistance can be attributed is failure to activate CDC due to protective complement regulatory proteins, including CFH. We hypothesized that rituximab killing of non-responsive B-CLL cells could be augmented by inactivating CFH with the CFH mAb. B cells from 12 patients were tested with B-CLL cells ex vivo in CDC assays with combinations of mAb7968, rituximab, and a negative control antibody. In 6 of 12 cases, it was found that CDC of rituximab non-responsive B-CLL cells could be augmented by the CFH mAb (shown for one patient in FIG. 10). Antibody-mediated cytotoxicity of cells was dependent upon functional complement. In one case where B-CLL cells were refractory to CDC by the combination of rituximab plus CFH mAb, additionally neutralizing the membrane complement regulatory protein CD59 allowed CDC to occur. Inhibiting CDC regulatory proteins such as CFH could hold promise for overcoming resistance to rituximab therapy in B-CLL.
[0242] Cetuximab is an anti-EGFR antibody that induces CDC of EGFR-positive tumor cells by the classical complement pathway in vivo. The CDC activity of cetuximab was previously shown to be significantly higher on cancer cells in which CFH was down-regulated. In order to examine whether a CFH mAb could potentiate CDC by cetuximab, a CDC assay was performed with A431 cells and a combination of cetuximab, mAb7968, and negative control antibodies. This experiment showed that mAb7968 enhanced CDC by cetuximab (FIG. 11).
[0243] These experiments demonstrate that the CFH mAb can potentiate CDC by other antibodies.
[0244] Confirmation of complement activation by CFH mAb. To confirm complement activation as a mechanism of CFH mAb-induced cell killing, products of the CDC reaction--the anaphylatoxins C3a and C5a, and the terminal MAC C5b-9--were assayed on two lung cancer cell lines treated with the antibody. Addition of CFH mAb7968 to A549 or H226 cells in the presence of NHS resulted in significant increases in C3a release (FIG. 12A), C5a release (FIG. 12B) and C5b-9 deposition (FIG. 12C) over levels of each measured with an IgG negative control. Note that under the conditions that this experiment was done, either of two mechanisms could account for these findings: i, the CFH mAb could inactivate CFH, leading to activation of the alternative complement pathway, and/or; ii, the CFH mAb could bind to C1q via its Fc domain, leading to activation of the classical complement pathway.
[0245] Release of cytokines from tumor cells by CFH mAb treatment. To ask whether other signaling molecules are released upon CDC of tumor cells, release of known cytokines from complement-lysed cells were measured using a cytokine array. A549 cells were incubated for 4 hr with serum and CFH mAb or a matched control mAb, then assayed the conditioned media on the R&D Systems array ARY022B (105 targets). Increased levels of 6 targets were observed in the complement-lysed samples compared to the control in two independent experiments. These were the two adipokines adipsin and leptin (cytokines that promote inflammation in adipose tissue); CD14, a pattern recognition receptor; and three cell surface proteins with pro-metastatic activities: EMMPRIN (CD147), ICAM-1 (CD54) and thrombospondin-1 (TSP-1). This experiment demonstrated that cells lysed by CDC release molecules that potentially interact with the innate or adaptive immune systems (in addition to C3a and C5a).
[0246] Effect of the CFH mAb on activation and migration of DC. C3a and C5a bind receptors on DC, activating them to mature and cross-prime T cells. Since tumor cells treated with the CFH mAb release C3a and C5a, whether mAb treatment of tumor cells could cause DC activation was investigated. In this experiment, A549 cells were treated with mAb or control mAb in the presence of NHS to allow CDC, then the supernatants from these reactions were incubated with PBMCs to investigate activation and maturation of immune cells. The cells were labeled with a variety of antibodies for immune cell typing and sorted by flow cytometry. DCs were gated as the population of lineage minus (CD3.sup.-CD14.sup.-CD16.sup.-CD19.sup.-CD20.sup.-CD56.sup.-) CD25.sup.+ HLA-DR.sup.+ cells. A 41% increase in DC activation was seen when supernatants generated by incubation of tumor cells with the CFH mAb were used for PBMC activation compared to when the negative control mAb was used (FIG. 13A).
[0247] To see if CDC supernatants could induce DC migration, a DC migration assay was performed in 5 .mu.m transwell plates. Peripheral blood mononuclear cells (PBMCs) from a normal donor were used as a source of DCs and placed on one side of the chamber. Conditioned medium from CFH mAb or control mAb treated cells was placed in the recipient chamber. After incubation to allow migration, flow cytometry was performed on the migrated cells with the DCs gated as lineage minus HLA-DR positive cells. The DC population comprised 4.4% of the PBMCs that migrated using supernatant from the CDC reaction with the CFH mAb, but only 2.1% of the cells that migrated using the supernatant from the control reaction with IgG (FIG. 13B).
[0248] Evidence for immunogenic cell death signaling. Dying tumor cells display on their surfaces or release damage-associated molecular pattern molecules (DAMPs), molecular hallmarks of "immunogenic cell death" (ICD) that stimulate DC cross-priming of T cells with tumor cell antigens. However, it is not known if CDC is associated with expression or release of DAMPs or other immunostimulatory molecules. Cells undergoing CDC show features of programmed necrotic cell death involving Bid and JNK and this death pathway is counteracted by the NF-.kappa.B signaling pathway. Little else is known about death pathways in CDC.
[0249] In order to see if lung tumor cells undergoing CFH mAb-induced CDC send an ICD signal, human lung tumor cell lines (H460 and A549) were incubated in medium containing complement from the addition of NHS, and treated with CFH mAb, control antibody, or the known ICD inducer doxorubicin (Kepp et al. Oncoimmunology 2014; 3:e955691; Krysko et al. Nat Rev Cancer 2012; 12:860-75.). Cells were harvested, plasma membranes prepared, and probed for the appearance of calreticulin, a surface-displayed DAMP. In both cell lines, increases in plasma membrane calreticulin were seen in CFH mAb and doxorubicin-treated cells, compared to negative controls (FIG. 14). Total calreticulin in whole cell lysates was the same under all conditions. These results implicate CDC as an ICD signal generating event.
[0250] Effect of the CFH mAb on tumor growth. For in vivo studies, a murine version of mAb7968 was developed. To test the effect of antibody on tumor growth, an adult patient-derived brain tumor xenograft grown s.c. in nude mice was first used (Bigner et al. Cancer Res 1990; 50:8017-2). After allowing tumors to grow to 200 mm.sup.3 in volume we performed intratumoral injections of mAb7968 or a murine subtype-matched negative control mAb. Injections were repeated biweekly for 3 weeks with tumor measurements recorded throughout the study. By the end of the study, there was significant tumor growth inhibition and prolonged survival in the group of animals that received mAb7968 (FIGS. 15A and 15B). Stained sections of tumor excised from mice receiving the negative control mAb show densely packed tumor cells and no lymphocytic infiltrate where sections from the small tumor mass from a mAb7968-treated mouse show a dense lymphocytic infiltrate (FIG. 15C). The primary concern for side effects from inhibition of CFH is renal toxicity; however, there were no observed adverse reactions at necropsy in any animal treated with the CFH mAb.
[0251] Tumor growth experiments were performed in the immunocompetent, syngeneic KLN205 lung cancer-DBA/2 mouse model, this time injecting mAb7968 i.p. on days 1, 4, 7, 10 and 13 after s.c. tumor initiation. Again, there was significant tumor growth inhibition in the group of animals that received the CFH mAb (FIG. 16A). As was the case with brain tumors, stained sections of s.c. lung tumor excised from mice receiving the negative control mAb show densely packed tumor cells and no lymphocytic infiltrate where sections from the small tumor mass from a mAb7968-treated mouse show a dense lymphocytic infiltrate (FIG. 16B). Again, there were no observed adverse reactions at necropsy in any of the animals treated with the CFH mAb.
[0252] It is understood that the foregoing detailed description and accompanying examples are merely illustrative and are not to be taken as limitations upon the scope of the invention, which is defined solely by the appended claims and their equivalents.
[0253] Various changes and modifications to the disclosed embodiments will be apparent to those skilled in the art. Such changes and modifications, including without limitation those relating to the chemical structures, substituents, derivatives, intermediates, syntheses, compositions, formulations, or methods of use of the invention, may be made without departing from the spirit and scope thereof.
[0254] For reasons of completeness, various aspects of the invention are set out in the following numbered clause:
[0255] Clause 1. An isolated antibody or antibody fragment thereof which immunospecifically binds to Complement Factor H (CFH) protein, wherein the antibody comprises: i. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:11, a CDR2 comprising the amino acid sequence of SEQ ID NO:21, and a CDR comprising the amino acid sequence of SEQ ID NO:31, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:51, a CDR2 comprising the amino acid sequence of SEQ ID NO:61, and a CDR3 comprising the amino acid sequence of SEQ ID NO:71; ii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:12, a CDR2 comprising the amino acid sequence of SEQ ID NO:22, and a CDR comprising the amino acid sequence of SEQ ID NO:32, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:52, a CDR2 comprising the amino acid sequence of SEQ ID NO:62, and a CDR3 comprising the amino acid sequence of SEQ ID NO:72; iii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:13, a CDR2 comprising the amino acid sequence of SEQ ID NO:23, and a CDR comprising the amino acid sequence of SEQ ID NO:33, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:53, a CDR2 comprising the amino acid sequence of SEQ ID NO:63, and a CDR3 comprising the amino acid sequence of SEQ ID NO:73; iv. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:14, a CDR2 comprising the amino acid sequence of SEQ ID NO:24, and a CDR comprising the amino acid sequence of SEQ ID NO:34, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:54, a CDR2 comprising the amino acid sequence of SEQ ID NO:64, and a CDR3 comprising the amino acid sequence of SEQ ID NO:74; v. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:15, a CDR2 comprising the amino acid sequence of SEQ ID NO:25, and a CDR comprising the amino acid sequence of SEQ ID NO:35, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:55, a CDR2 comprising the amino acid sequence of SEQ ID NO:65, and a CDR3 comprising the amino acid sequence of SEQ ID NO:75; vi. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:16, a CDR2 comprising the amino acid sequence of SEQ ID NO:26, and a CDR comprising the amino acid sequence of SEQ ID NO:36, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:56, a CDR2 comprising the amino acid sequence of SEQ ID NO:66, and a CDR3 comprising the amino acid sequence of SEQ ID NO:76; vii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:17, a CDR2 comprising the amino acid sequence of SEQ ID NO:27, and a CDR comprising the amino acid sequence of SEQ ID NO:37, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:57, a CDR2 comprising the amino acid sequence of SEQ ID NO:67, and a CDR3 comprising the amino acid sequence of SEQ ID NO:77; viii. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:18, a CDR2 comprising the amino acid sequence of SEQ ID NO:28, and a CDR comprising the amino acid sequence of SEQ ID NO:38, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:58, a CDR2 comprising the amino acid sequence of SEQ ID NO:68, and a CDR3 comprising the amino acid sequence of SEQ ID NO:78; ix. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:19, a CDR2 comprising the amino acid sequence of SEQ ID NO:29, and a CDR comprising the amino acid sequence of SEQ ID NO:39, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:59, a CDR2 comprising the amino acid sequence of SEQ ID NO:69, and a CDR3 comprising the amino acid sequence of SEQ ID NO:79; or x. a variable heavy domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:20, a CDR2 comprising the amino acid sequence of SEQ ID NO:30, and a CDR comprising the amino acid sequence of SEQ ID NO: 40, and a variable light domain chain comprising a CDR1 comprising the amino acid sequence of SEQ ID NO:60, a CDR2 comprising the amino acid sequence of SEQ ID NO:70, and a CDR3 comprising the amino acid sequence of SEQ ID NO:80.
[0256] Clause 2. The isolated antibody or antibody fragment of clause 1, wherein the isolated antibody or antibody fragment thereof binds to an epitope within short consensus repeat (SCR) 19 of CFH protein and the epitope is a cryptic epitope that is exposed in a tumor-bound form of CFH.
[0257] Clause 3. The isolated antibody or antibody fragment of clause 2, wherein the epitope comprises PIDNGDIT (SEQ ID NO: 3), or fragment or variant thereof.
[0258] Clause 4. The isolated antibody or antibody fragment of clause 1, wherein the antibody comprises: a) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:1 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:41; b) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:2 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:42; c) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:3 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:43; d) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:4 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:44; e) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:5 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:45; f) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:6 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:46; g) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:7 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:47; h) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:8 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:48; i) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:9 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:49; or j) a variable heavy domain comprising the amino acid sequence of SEQ ID NO:10 and a variable light domain region comprising the amino acid sequence of SEQ ID NO:50.
[0259] Clause 5. The isolated antibody or antibody fragment of clause 1, wherein the isolated antibody or antibody fragment does not cross-react with at least one of systemic lupus erythematosus autoantigens SSA, SSB, sphingomyelin (Sm), ribonucleoprotein (RNP), sclerosis autoantigen (Scl-70), histidine-tRNA ligase (Jo-1), double-stranded DNA (dsDNA), centromere B (CentB), and histones.
[0260] Clause 6. The isolated antibody or antibody fragment of clause 1, wherein the isolated antibody or antibody fragment is selected from the group consisting of a human antibody, an immunoglobulin molecule, a disulfide linked Fv, a monoclonal antibody, an affinity matured, a scFv, a chimeric antibody, a single domain antibody, a CDR-grafted antibody, a diabody, a humanized antibody, a multispecific antibody, a Fab, a dual specific antibody, a DVD, a TVD, a Fab', a bispecific antibody, a F(ab')2, and a Fv.
[0261] Clause 7. The isolated antibody or antibody fragment of clause 1, wherein the isolated antibody or antibody fragment comprises a heavy chain immunoglobulin constant domain selected from the group consisting of a human IgM constant domain, a human IgG4 constant domain, a human IgG1 constant domain, a human IgE constant domain, a human IgG2 constant domain, a human igG3 constant domain, and a human IgA constant domain.
[0262] Clause 8. A pharmaceutical composition comprising the isolated antibody or antibody fragment of clause 1.
[0263] Clause 9. A method of treating a subject in need thereof having cancer or inhibiting tumor growth in a subject, the method comprising administering to the subject the isolated antibody or antibody fragment of clause 1.
[0264] Clause 10. The method of clause 9, wherein the cancer is lung cancer.
[0265] Clause 11. The method of clause 9, further comprising administering an effective amount of at least one of Cetuximab, PERJETA.RTM., or HERCEPTIN.RTM..
[0266] Clause 12. A method of increasing complement dependent lysis of a cell, increasing C3b deposition on a cell, or inhibiting Complement Factor H (CFH) binding to C3b in a subject or a cell, the method comprising administering to the cell the isolated antibody or antibody fragment of clause 1.
Sequence CWU 1
1
1221116PRTArtificial sequenceSynthetic 1Leu Val Glu Ser Gly Gly Gly
Val Val Arg Pro Gly Arg Ser Leu Arg1 5 10 15Leu Ser Cys Val Ala Ser
Gly Phe Thr Phe Asn Ala Tyr Gly Met His 20 25 30Trp Val Arg Gln Gly
Pro Gly Lys Gly Leu Glu Trp Leu Ala Val Ile 35 40 45Ser Tyr Glu Gly
Lys Thr Val Tyr Tyr Ala Asp Ser Val Lys Asp Arg 50 55 60Phe Thr Ile
Ser Arg Asp Asn Ser Arg Asn Thr Val Ser Leu His Leu65 70 75 80Asn
Asn Leu Arg Gly Glu Asp Thr Ala Val Tyr Tyr Cys Ala Lys Gly 85 90
95Ser Ala Ser Ala Ala Val Leu Gln His Trp Gly Gln Gly Thr Leu Val
100 105 110Ser Val Thr Ser 1152116PRTArtificial sequenceSynthetic
2Leu Val Glu Ser Gly Gly Gly Val Val Pro Pro Gly Lys Ser Leu Arg1 5
10 15Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Leu Tyr Gly Ile
His 20 25 30Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val Ala
Val Ile 35 40 45Ser Tyr Asp Gly Asn Thr Lys Tyr Tyr Thr Asp Ser Val
Lys Gly Arg 50 55 60Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Ile
Tyr Leu Gln Met65 70 75 80Asn Ser Leu Arg Leu Asp Asp Thr Ala Val
Tyr Tyr Cys Ala Lys Gly 85 90 95Ala Ala Asn Ser Ala Thr Phe Asp Phe
Trp Gly Arg Gly Thr Met Val 100 105 110Thr Val Ser Ser
1153116PRTArtificial sequenceSynthetic 3Leu Val Glu Ser Gly Gly Gly
Val Val Gln Pro Gly Lys Ser Leu Arg1 5 10 15Leu Ser Cys Val Ala Ser
Gly Phe Ser Phe Ser Thr Tyr Gly Met His 20 25 30Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val Ala Val Met 35 40 45Ser Phe Asp Gly
Lys Thr Lys Tyr Tyr Ala Asp Ser Val Lys Gly Arg 50 55 60Phe Thr Ile
Ser Arg Asp Asn Pro Lys Asn Thr Leu Tyr Leu Gln Met65 70 75 80Asn
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys Ala Lys Gly 85 90
95Gly Ala Ala Ala Ala Val Phe Asp Ser Trp Gly Pro Gly Ile Leu Leu
100 105 110Thr Val Ser Ser 1154116PRTArtificial sequenceSynthetic
4Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly Arg Ser Leu Arg1 5
10 15Leu Ser Cys Ala Ala Ser Gly Val Thr Phe Ser Arg Tyr Gly Met
His 20 25 30Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val Ala
Val Ile 35 40 45Ser Tyr Asp Glu Lys Thr Lys Tyr Tyr Ala Asp Ser Val
Lys Gly Arg 50 55 60Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu
Phe Leu His Met65 70 75 80Asn Arg Leu Arg Tyr Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Lys Gly 85 90 95Ala Ser Ser Gly Ala Tyr Phe Asp Tyr
Trp Gly Gln Gly Thr Leu Val 100 105 110Thr Val Ser Ser
1155116PRTArtificial sequenceSynthetic 5Leu Val Glu Ser Gly Gly Gly
Val Val Gln Pro Gly Lys Ser Leu Arg1 5 10 15Leu Ser Cys Val Ala Ser
Gly Phe Thr Phe Ser Thr Tyr Gly Met His 20 25 30Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val Ala Val Met 35 40 45Ser Phe Asp Gly
Lys Thr Lys Tyr Tyr Ala Asp Ser Val Lys Gly Arg 50 55 60Phe Thr Ile
Ser Arg Asp Asn Pro Lys Asn Thr Leu Tyr Leu Gln Met65 70 75 80Asn
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys Ala Lys Gly 85 90
95Gly Ala Ala Ala Ala Val Phe Asp Ser Trp Gly Gln Gly Ile Leu Leu
100 105 110Thr Val Ser Ser 1156116PRTArtificial sequenceSynthetic
6Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly Lys Ser Leu Arg1 5
10 15Leu Ser Cys Val Ala Ser Gly Phe Thr Phe Ser Ser Tyr Gly Met
His 20 25 30Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val Ala
Val Met 35 40 45Ser Phe Asp Gly Lys Thr Lys Tyr Tyr Ala Asp Ser Val
Lys Gly Arg 50 55 60Phe Thr Ile Ser Arg Asp Asn Pro Lys Asn Thr Leu
Tyr Leu Gln Met65 70 75 80Asn Ser Leu Arg Ser Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Lys Gly 85 90 95Gly Ala Ala Ala Ala Val Phe Asp Ser
Trp Gly Gln Gly Ile Leu Leu 100 105 110Thr Val Ser Ser
1157118PRTArtificial sequenceSynthetic 7Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly Ser Leu Arg1 5 10 15Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Ser Tyr Glu Met Asn 20 25 30Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val Ser Tyr Ile 35 40 45Ser Ser Ser Gly
Ser Thr Ile Tyr Tyr Ala Asp Ser Val Lys Gly Arg 50 55 60Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr Leu Gln Met65 70 75 80Asn
Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg Val 85 90
95Glu Gln Leu Ala Pro Ser Pro Tyr Met Asp Val Trp Gly Lys Gly Thr
100 105 110Thr Val Thr Val Ser Ser 1158116PRTArtificial
sequenceSynthetic 8Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly
Arg Ser Leu Arg1 5 10 15Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser
Arg Tyr Gly Met His 20 25 30Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val Ala Val Met 35 40 45Ser Tyr Asp Gly Ser Thr Lys Tyr Tyr
Ala Asp Ser Val Lys Gly Arg 50 55 60Phe Ala Ile Ser Arg Asp Asn Pro
Lys Asn Thr Leu Phe Leu Gln Met65 70 75 80Asn Ser Leu Arg Pro Asp
Asp Thr Ala Val Tyr Tyr Cys Ala Lys Gly 85 90 95Gly Ala Ala Ala Ala
Val Met Asp Val Trp Gly Lys Gly Thr Thr Val 100 105 110Thr Val Ser
Ser 1159124PRTArtificial sequenceSynthetic 9Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Glu1 5 10 15Ser Leu Lys Ile Ser
Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr 20 25 30Trp Ile Gly Trp
Val Arg Gln Met Pro Gly Lys Gly Leu Glu Trp Met 35 40 45Gly Ile Ile
Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe 50 55 60Gln Gly
Gln Val Thr Ile Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70 75
80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Tyr Cys
85 90 95Ala Arg Arg Gly Leu Arg Gly Ala Tyr Tyr Tyr Tyr Tyr Gly Met
Asp 100 105 110Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115
12010120PRTArtificial sequenceSynthetic 10Leu Val Glu Ser Gly Gly
Gly Val Val Gln Pro Gly Arg Ser Leu Arg1 5 10 15Leu Ser Cys Ala Ala
Ser Gly Leu Thr Phe Ser Phe Tyr Asn Phe His 20 25 30Trp Val Arg Gln
Thr Pro Gly Lys Gly Leu Glu Trp Val Ala Gly Ile 35 40 45Ser Tyr Asp
Ala Thr Arg Thr Asn Tyr Ala Gly Ser Val Thr Gly Arg 50 55 60Phe Thr
Ile Ser Arg Asp Asn Ser Lys Lys Met Leu Tyr Leu Gln Met65 70 75
80Ser Ser Leu Gly Pro Gln Asp Thr Ala Val Tyr His Cys Ala Arg Asp
85 90 95Arg Ser Asp Gly Gln Leu His Lys Val Ala Phe Asp Ser Trp Gly
Gln 100 105 110Gly Ala Leu Val Thr Val Ser Ser 115
120119PRTArtificial sequenceSynthetic 11Phe Thr Phe Asn Ala Tyr Gly
Met His1 5129PRTArtificial sequenceSynthetic 12Phe Thr Phe Ser Leu
Tyr Gly Ile His1 5139PRTArtificial sequenceSynthetic 13Phe Ser Phe
Ser Thr Tyr Gly Met His1 5149PRTArtificial sequenceSynthetic 14Val
Thr Phe Ser Arg Tyr Gly Met His1 5159PRTArtificial
sequenceSynthetic 15Phe Thr Phe Ser Thr Tyr Gly Met His1
5169PRTArtificial sequenceSynthetic 16Phe Thr Phe Ser Ser Tyr Gly
Met His1 5179PRTArtificial sequenceSynthetic 17Phe Thr Phe Ser Ser
Tyr Glu Met Asn1 5189PRTArtificial sequenceSynthetic 18Phe Thr Phe
Ser Arg Tyr Gly Met His1 51914PRTArtificial sequenceSynthetic 19Cys
Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr Trp Ile Gly1 5
10209PRTArtificial sequenceSynthetic 20Leu Thr Phe Ser Phe Tyr Asn
Phe His1 52114PRTArtificial sequenceSynthetic 21Trp Leu Ala Val Ile
Ser Tyr Glu Gly Lys Thr Val Tyr Tyr1 5 102215PRTArtificial
sequenceSynthetic 22Trp Val Ala Val Ile Ser Tyr Asp Gly Asn Thr Lys
Tyr Tyr Thr1 5 10 152314PRTArtificial sequenceSynthetic 23Trp Val
Ala Val Met Ser Phe Asp Gly Lys Thr Lys Tyr Tyr1 5
102414PRTArtificial sequenceSynthetic 24Trp Val Ala Val Ile Ser Tyr
Asp Glu Lys Thr Lys Tyr Tyr1 5 102514PRTArtificial
sequenceSynthetic 25Trp Val Ala Val Met Ser Phe Asp Gly Lys Thr Lys
Tyr Tyr1 5 102614PRTArtificial sequenceSynthetic 26Trp Val Ala Val
Met Ser Phe Asp Gly Lys Thr Lys Tyr Tyr1 5 102714PRTArtificial
sequenceSynthetic 27Trp Val Ser Tyr Ile Ser Ser Ser Gly Ser Thr Ile
Tyr Tyr1 5 102814PRTArtificial sequenceSynthetic 28Trp Val Ala Val
Met Ser Tyr Asp Gly Ser Thr Lys Tyr Tyr1 5 102914PRTArtificial
sequenceSynthetic 29Trp Met Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr
Arg Tyr1 5 103014PRTArtificial sequenceSynthetic 30Trp Val Ala Gly
Ile Ser Tyr Asp Ala Thr Arg Thr Asn Tyr1 5 103111PRTArtificial
sequenceSynthetic 31Lys Gly Ser Ala Ser Ala Ala Val Leu Gln His1 5
103211PRTArtificial sequenceSynthetic 32Lys Gly Ala Ala Asn Ser Ala
Thr Phe Asp Phe1 5 103311PRTArtificial sequenceSynthetic 33Lys Gly
Gly Ala Ala Ala Ala Val Phe Asp Ser1 5 103411PRTArtificial
sequenceSynthetic 34Lys Gly Ala Ser Ser Gly Ala Tyr Phe Asp Tyr1 5
103511PRTArtificial sequenceSynthetic 35Lys Gly Gly Ala Ala Ala Ala
Val Phe Asp Ser1 5 103611PRTArtificial sequenceSynthetic 36Lys Gly
Gly Ala Ala Ala Ala Val Phe Asp Ser1 5 103713PRTArtificial
sequenceSynthetic 37Arg Val Glu Gln Leu Ala Pro Ser Pro Tyr Met Asp
Val1 5 103811PRTArtificial sequenceSynthetic 38Lys Gly Gly Ala Ala
Ala Ala Val Met Asp Val1 5 103916PRTArtificial sequenceSynthetic
39Arg Arg Gly Leu Arg Gly Ala Tyr Tyr Tyr Tyr Tyr Gly Met Asp Val1
5 10 154015PRTArtificial sequenceSynthetic 40Arg Asp Arg Ser Asp
Gly Gln Leu His Lys Val Ala Phe Asp Ser1 5 10 1541110PRTArtificial
sequenceSynthetic 41Met Thr Gln Ser Pro Gly Ser Leu Ala Val Ser Leu
Gly Ser Arg Ala1 5 10 15Thr Ile Asn Cys Lys Ser Ser Arg Ser Leu Leu
Tyr Arg Ser Asn Asn 20 25 30Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys
Pro Gly Gln Ser Pro Arg 35 40 45Leu Leu Ile Tyr Trp Ala Ser Ser Arg
Glu Ser Gly Val Pro Asp Arg 50 55 60Phe Ser Gly Gly Gly Ser Gly Thr
Ser Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Ala Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln Tyr Phe Asn 85 90 95Pro Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 11042110PRTArtificial
sequenceSynthetic 42Met Thr Gln Ser Pro Asp Ser Leu Thr Leu Ser Leu
Gly Glu Arg Ala1 5 10 15Thr Ile Asn Cys Arg Ser Ser Arg Thr Val Leu
Tyr Arg Ser Asn Asn 20 25 30Lys Asn Tyr Leu Ala Trp Tyr Gln His Lys
Pro Gly Gln Pro Pro Lys 35 40 45Leu Leu Met Ser Trp Ala Ser Thr Arg
Glu Thr Gly Val Pro Asp Arg 50 55 60Phe Ser Gly Ser Gly Ser Gly Thr
His Phe Thr Leu Thr Ile Thr Ser65 70 75 80Leu Gln Pro Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Ser 85 90 95Pro Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Arg 100 105 11043110PRTArtificial
sequenceSynthetic 43Met Thr Gln Ser Pro Asn Ser Leu Ala Val Ser Leu
Gly Gly Arg Ala1 5 10 15Thr Ile Asn Cys Lys Ala Ser Gln Ser Ile Leu
Tyr Arg Ser Asn Asn 20 25 30Lys Asn Tyr Leu Ala Trp Tyr Gln His Lys
Ala Gly Gln Pro Pro Lys 35 40 45Leu Leu Ile Tyr Trp Ala Ser Thr Arg
Glu Ser Gly Val Pro Glu Arg 50 55 60Phe Ser Gly Ser Gly Ser Arg Thr
Asp Phe Thr Leu Thr Ile Asn Gly65 70 75 80Leu Gln Ala Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Asn 85 90 95Pro Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 11044110PRTArtificial
sequenceSynthetic 44Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu
Gly Glu Arg Ala1 5 10 15Thr Ile Lys Cys Lys Ser Ser Gln Ser Val Leu
Tyr Ser Ser Asn Asn 20 25 30Lys Asn Tyr Leu Ala Trp Tyr Gln His Lys
Pro Gly Gln Pro Pro Lys 35 40 45Val Leu Val Tyr Trp Ala Ser Thr Arg
Glu Ser Gly Val Pro Asp Arg 50 55 60Phe Ser Gly Ser Gly Ser Gly Thr
Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Ala Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Asn 85 90 95Pro Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Ala Ile Lys 100 105 11045110PRTArtificial
sequenceSynthetic 45Met Thr Gln Ser Pro Asn Ser Leu Ala Val Ser Leu
Gly Gly Arg Ala1 5 10 15Thr Ile Asn Cys Lys Thr Ser Gln Ser Ile Leu
Tyr Arg Ser Asn Asn 20 25 30Lys Asn Tyr Leu Ala Trp Tyr Gln His Lys
Pro Gly Gln Pro Pro Lys 35 40 45Leu Leu Ile Tyr Trp Ala Ser Thr Arg
Glu Ser Arg Val Pro Asp Arg 50 55 60Phe Ser Gly Ser Gly Ser Arg Thr
Asp Phe Thr Leu Thr Ile Ser Gly65 70 75 80Leu Gln Ala Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Asn 85 90 95Pro Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 11046110PRTArtificial
sequenceSynthetic 46Met Thr Gln Ser Pro Asn Ser Leu Ala Val Ser Leu
Gly Gly Arg Ala1 5 10 15Thr Ile Asn Cys Lys Thr Ser Gln Ser Ile Leu
Tyr Arg Ser Asn Asn 20 25 30Lys Asn Tyr Leu Ala Trp Tyr Gln His Lys
Ser Gly Gln Pro Pro Lys 35 40 45Leu Leu Ile Tyr Trp Ala Ser Thr Arg
Glu Ser Gly Val Pro Asp Arg 50 55 60Phe Ser Gly Ser Gly Ser Arg Thr
Asp Phe Thr Leu Thr Ile Ser Gly65 70 75 80Leu Gln Ala Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Asn 85 90 95Pro Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 11047110PRTArtificial
sequenceSynthetic 47Met Thr Gln Ser Leu Asp Ser Leu Thr Val Ser Leu
Gly Glu Arg Ala1 5 10 15Thr Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu
Tyr Thr Ser Asn Asn 20 25 30Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Ser Gly Gln Pro Pro Lys 35 40 45Leu Leu
Ile Tyr Trp Ala Ser Ile Arg Asp Ser Gly Val Pro Asp Arg 50 55 60Phe
Ser Gly Ser Gly Ser Ala Thr Asp Phe Thr Leu Thr Ile Asn Asn65 70 75
80Leu Gln Ala Glu Asp Val Ala Val Tyr Phe Cys Gln Gln Tyr Tyr Lys
85 90 95Thr Pro Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Val Arg 100
105 11048110PRTArtificial sequenceSynthetic 48Met Thr Gln Ser Pro
Asp Ser Leu Thr Val Ser Leu Gly Glu Arg Ala1 5 10 15Thr Ile Ser Cys
Lys Ser Ser Gln Arg Leu Leu Tyr Ser Ser Asn Asn 20 25 30Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro Pro Lys 35 40 45Leu Leu
Met Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val Pro Asp Arg 50 55 60Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Ser Leu Thr Ile Ser Ser65 70 75
80Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Asn
85 90 95Pro Pro Trp Thr Phe Gly Gln Gly Thr Lys Val Glu Val Lys 100
105 11049107PRTArtificial sequenceSyntheticmisc_feature(4)..(4)Xaa
can be any naturally occurring amino acid 49Asp Ile Gln Xaa Thr Gln
Ser Pro Ser Thr Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Trp 20 25 30Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala
Ser Ser Leu Glu Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Asp Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Tyr Asn Ser Tyr Ser Trp
85 90 95Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100
10550110PRTArtificial sequenceSynthetic 50Met Thr Gln Ser Pro Asp
Ser Leu Thr Leu Ser Leu Gly Glu Arg Ala1 5 10 15Thr Ile Asn Cys Arg
Ser Ser Arg Thr Val Leu Tyr Arg Ser Asn Asn 20 25 30Lys Asn Tyr Leu
Ala Trp Tyr Gln His Lys Pro Gly Gln Pro Pro Lys 35 40 45Leu Leu Met
Ser Trp Ala Ser Thr Arg Glu Thr Gly Val Pro Asp Arg 50 55 60Phe Ser
Gly Ser Gly Ser Gly Thr His Phe Thr Leu Thr Ile Thr Ser65 70 75
80Leu Gln Pro Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Ser
85 90 95Pro Pro Trp Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Arg 100
105 1105114PRTArtificial sequenceSynthetic 51Arg Ser Leu Leu Tyr
Arg Ser Asn Asn Lys Asn Tyr Leu Ala1 5 105214PRTArtificial
sequenceSynthetic 52Arg Thr Val Leu Tyr Arg Ser Asn Asn Lys Asn Tyr
Leu Ala1 5 105314PRTArtificial sequenceSynthetic 53Gln Ser Ile Leu
Tyr Arg Ser Asn Asn Lys Asn Tyr Leu Ala1 5 105414PRTArtificial
sequenceSynthetic 54Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys Asn Tyr
Leu Ala1 5 105514PRTArtificial sequenceSynthetic 55Gln Ser Ile Leu
Tyr Arg Ser Asn Asn Lys Asn Tyr Leu Ala1 5 105614PRTArtificial
sequenceSynthetic 56Gln Ser Ile Leu Tyr Arg Ser Asn Asn Lys Asn Tyr
Leu Ala1 5 105714PRTArtificial sequenceSynthetic 57Gln Ser Leu Leu
Tyr Thr Ser Asn Asn Lys Asn Tyr Leu Ala1 5 105814PRTArtificial
sequenceSynthetic 58Gln Arg Leu Leu Tyr Ser Ser Asn Asn Lys Asn Tyr
Leu Ala1 5 10598PRTArtificial sequenceSynthetic 59Gln Ser Ile Ser
Ser Trp Leu Ala1 56014PRTArtificial sequenceSynthetic 60Arg Thr Val
Leu Tyr Arg Ser Asn Asn Lys Asn Tyr Leu Ala1 5 106111PRTArtificial
sequenceSynthetic 61Leu Leu Ile Tyr Trp Ala Ser Ser Arg Glu Ser1 5
106210PRTArtificial sequenceSynthetic 62Leu Leu Met Ser Trp Ala Ser
Thr Arg Glu1 5 106311PRTArtificial sequenceSynthetic 63Leu Leu Ile
Tyr Trp Ala Ser Thr Arg Glu Ser1 5 106411PRTArtificial
sequenceSynthetic 64Val Leu Val Tyr Trp Ala Ser Thr Arg Glu Ser1 5
106512PRTArtificial sequenceSynthetic 65Leu Leu Ile Tyr Trp Ala Ser
Thr Arg Glu Ser Arg1 5 106611PRTArtificial sequenceSynthetic 66Leu
Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser1 5 106711PRTArtificial
sequenceSynthetic 67Leu Leu Ile Tyr Trp Ala Ser Ile Arg Asp Ser1 5
106811PRTArtificial sequenceSynthetic 68Leu Leu Met Tyr Trp Ala Ser
Thr Arg Glu Ser1 5 106911PRTArtificial sequenceSynthetic 69Leu Leu
Ile Tyr Lys Ala Ser Ser Leu Glu Ser1 5 107011PRTArtificial
sequenceSynthetic 70Leu Leu Met Ser Trp Ala Ser Thr Arg Glu Thr1 5
10718PRTArtificial sequenceSynthetic 71Gln Gln Tyr Phe Asn Pro Pro
Trp1 5728PRTArtificial sequenceSynthetic 72Gln Gln Tyr Tyr Ser Pro
Pro Trp1 5738PRTArtificial sequenceSynthetic 73Gln Gln Tyr Tyr Asn
Pro Pro Trp1 5748PRTArtificial sequenceSynthetic 74Gln Gln Tyr Tyr
Asn Pro Pro Trp1 5758PRTArtificial sequenceSynthetic 75Gln Gln Tyr
Tyr Asn Pro Pro Trp1 5768PRTArtificial sequenceSynthetic 76Gln Gln
Tyr Tyr Asn Pro Pro Trp1 5778PRTArtificial sequenceSynthetic 77Gln
Gln Tyr Tyr Lys Thr Pro Leu1 5788PRTArtificial sequenceSynthetic
78Gln Gln Tyr Tyr Asn Pro Pro Trp1 5798PRTArtificial
sequenceSynthetic 79Gln Gln Tyr Asn Ser Tyr Ser Trp1
5808PRTArtificial sequenceSynthetic 80Gln Gln Tyr Tyr Ser Pro Pro
Trp1 5811231PRTHomo sapien 81Met Arg Leu Leu Ala Lys Ile Ile Cys
Leu Met Leu Trp Ala Ile Cys1 5 10 15Val Ala Glu Asp Cys Asn Glu Leu
Pro Pro Arg Arg Asn Thr Glu Ile 20 25 30Leu Thr Gly Ser Trp Ser Asp
Gln Thr Tyr Pro Glu Gly Thr Gln Ala 35 40 45Ile Tyr Lys Cys Arg Pro
Gly Tyr Arg Ser Leu Gly Asn Val Ile Met 50 55 60Val Cys Arg Lys Gly
Glu Trp Val Ala Leu Asn Pro Leu Arg Lys Cys65 70 75 80Gln Lys Arg
Pro Cys Gly His Pro Gly Asp Thr Pro Phe Gly Thr Phe 85 90 95Thr Leu
Thr Gly Gly Asn Val Phe Glu Tyr Gly Val Lys Ala Val Tyr 100 105
110Thr Cys Asn Glu Gly Tyr Gln Leu Leu Gly Glu Ile Asn Tyr Arg Glu
115 120 125Cys Asp Thr Asp Gly Trp Thr Asn Asp Ile Pro Ile Cys Glu
Val Val 130 135 140Lys Cys Leu Pro Val Thr Ala Pro Glu Asn Gly Lys
Ile Val Ser Ser145 150 155 160Ala Met Glu Pro Asp Arg Glu Tyr His
Phe Gly Gln Ala Val Arg Phe 165 170 175Val Cys Asn Ser Gly Tyr Lys
Ile Glu Gly Asp Glu Glu Met His Cys 180 185 190Ser Asp Asp Gly Phe
Trp Ser Lys Glu Lys Pro Lys Cys Val Glu Ile 195 200 205Ser Cys Lys
Ser Pro Asp Val Ile Asn Gly Ser Pro Ile Ser Gln Lys 210 215 220Ile
Ile Tyr Lys Glu Asn Glu Arg Phe Gln Tyr Lys Cys Asn Met Gly225 230
235 240Tyr Glu Tyr Ser Glu Arg Gly Asp Ala Val Cys Thr Glu Ser Gly
Trp 245 250 255Arg Pro Leu Pro Ser Cys Glu Glu Lys Ser Cys Asp Asn
Pro Tyr Ile 260 265 270Pro Asn Gly Asp Tyr Ser Pro Leu Arg Ile Lys
His Arg Thr Gly Asp 275 280 285Glu Ile Thr Tyr Gln Cys Arg Asn Gly
Phe Tyr Pro Ala Thr Arg Gly 290 295 300Asn Thr Ala Lys Cys Thr Ser
Thr Gly Trp Ile Pro Ala Pro Arg Cys305 310 315 320Thr Leu Lys Pro
Cys Asp Tyr Pro Asp Ile Lys His Gly Gly Leu Tyr 325 330 335His Glu
Asn Met Arg Arg Pro Tyr Phe Pro Val Ala Val Gly Lys Tyr 340 345
350Tyr Ser Tyr Tyr Cys Asp Glu His Phe Glu Thr Pro Ser Gly Ser Tyr
355 360 365Trp Asp His Ile His Cys Thr Gln Asp Gly Trp Ser Pro Ala
Val Pro 370 375 380Cys Leu Arg Lys Cys Tyr Phe Pro Tyr Leu Glu Asn
Gly Tyr Asn Gln385 390 395 400Asn Tyr Gly Arg Lys Phe Val Gln Gly
Lys Ser Ile Asp Val Ala Cys 405 410 415His Pro Gly Tyr Ala Leu Pro
Lys Ala Gln Thr Thr Val Thr Cys Met 420 425 430Glu Asn Gly Trp Ser
Pro Thr Pro Arg Cys Ile Arg Val Lys Thr Cys 435 440 445Ser Lys Ser
Ser Ile Asp Ile Glu Asn Gly Phe Ile Ser Glu Ser Gln 450 455 460Tyr
Thr Tyr Ala Leu Lys Glu Lys Ala Lys Tyr Gln Cys Lys Leu Gly465 470
475 480Tyr Val Thr Ala Asp Gly Glu Thr Ser Gly Ser Ile Thr Cys Gly
Lys 485 490 495Asp Gly Trp Ser Ala Gln Pro Thr Cys Ile Lys Ser Cys
Asp Ile Pro 500 505 510Val Phe Met Asn Ala Arg Thr Lys Asn Asp Phe
Thr Trp Phe Lys Leu 515 520 525Asn Asp Thr Leu Asp Tyr Glu Cys His
Asp Gly Tyr Glu Ser Asn Thr 530 535 540Gly Ser Thr Thr Gly Ser Ile
Val Cys Gly Tyr Asn Gly Trp Ser Asp545 550 555 560Leu Pro Ile Cys
Tyr Glu Arg Glu Cys Glu Leu Pro Lys Ile Asp Val 565 570 575His Leu
Val Pro Asp Arg Lys Lys Asp Gln Tyr Lys Val Gly Glu Val 580 585
590Leu Lys Phe Ser Cys Lys Pro Gly Phe Thr Ile Val Gly Pro Asn Ser
595 600 605Val Gln Cys Tyr His Phe Gly Leu Ser Pro Asp Leu Pro Ile
Cys Lys 610 615 620Glu Gln Val Gln Ser Cys Gly Pro Pro Pro Glu Leu
Leu Asn Gly Asn625 630 635 640Val Lys Glu Lys Thr Lys Glu Glu Tyr
Gly His Ser Glu Val Val Glu 645 650 655Tyr Tyr Cys Asn Pro Arg Phe
Leu Met Lys Gly Pro Asn Lys Ile Gln 660 665 670Cys Val Asp Gly Glu
Trp Thr Thr Leu Pro Val Cys Ile Val Glu Glu 675 680 685Ser Thr Cys
Gly Asp Ile Pro Glu Leu Glu His Gly Trp Ala Gln Leu 690 695 700Ser
Ser Pro Pro Tyr Tyr Tyr Gly Asp Ser Val Glu Phe Asn Cys Ser705 710
715 720Glu Ser Phe Thr Met Ile Gly His Arg Ser Ile Thr Cys Ile His
Gly 725 730 735Val Trp Thr Gln Leu Pro Gln Cys Val Ala Ile Asp Lys
Leu Lys Lys 740 745 750Cys Lys Ser Ser Asn Leu Ile Ile Leu Glu Glu
His Leu Lys Asn Lys 755 760 765Lys Glu Phe Asp His Asn Ser Asn Ile
Arg Tyr Arg Cys Arg Gly Lys 770 775 780Glu Gly Trp Ile His Thr Val
Cys Ile Asn Gly Arg Trp Asp Pro Glu785 790 795 800Val Asn Cys Ser
Met Ala Gln Ile Gln Leu Cys Pro Pro Pro Pro Gln 805 810 815Ile Pro
Asn Ser His Asn Met Thr Thr Thr Leu Asn Tyr Arg Asp Gly 820 825
830Glu Lys Val Ser Val Leu Cys Gln Glu Asn Tyr Leu Ile Gln Glu Gly
835 840 845Glu Glu Ile Thr Cys Lys Asp Gly Arg Trp Gln Ser Ile Pro
Leu Cys 850 855 860Val Glu Lys Ile Pro Cys Ser Gln Pro Pro Gln Ile
Glu His Gly Thr865 870 875 880Ile Asn Ser Ser Arg Ser Ser Gln Glu
Ser Tyr Ala His Gly Thr Lys 885 890 895Leu Ser Tyr Thr Cys Glu Gly
Gly Phe Arg Ile Ser Glu Glu Asn Glu 900 905 910Thr Thr Cys Tyr Met
Gly Lys Trp Ser Ser Pro Pro Gln Cys Glu Gly 915 920 925Leu Pro Cys
Lys Ser Pro Pro Glu Ile Ser His Gly Val Val Ala His 930 935 940Met
Ser Asp Ser Tyr Gln Tyr Gly Glu Glu Val Thr Tyr Lys Cys Phe945 950
955 960Glu Gly Phe Gly Ile Asp Gly Pro Ala Ile Ala Lys Cys Leu Gly
Glu 965 970 975Lys Trp Ser His Pro Pro Ser Cys Ile Lys Thr Asp Cys
Leu Ser Leu 980 985 990Pro Ser Phe Glu Asn Ala Ile Pro Met Gly Glu
Lys Lys Asp Val Tyr 995 1000 1005Lys Ala Gly Glu Gln Val Thr Tyr
Thr Cys Ala Thr Tyr Tyr Lys 1010 1015 1020Met Asp Gly Ala Ser Asn
Val Thr Cys Ile Asn Ser Arg Trp Thr 1025 1030 1035Gly Arg Pro Thr
Cys Arg Asp Thr Ser Cys Val Asn Pro Pro Thr 1040 1045 1050Val Gln
Asn Ala Tyr Ile Val Ser Arg Gln Met Ser Lys Tyr Pro 1055 1060
1065Ser Gly Glu Arg Val Arg Tyr Gln Cys Arg Ser Pro Tyr Glu Met
1070 1075 1080Phe Gly Asp Glu Glu Val Met Cys Leu Asn Gly Asn Trp
Thr Glu 1085 1090 1095Pro Pro Gln Cys Lys Asp Ser Thr Gly Lys Cys
Gly Pro Pro Pro 1100 1105 1110Pro Ile Asp Asn Gly Asp Ile Thr Ser
Phe Pro Leu Ser Val Tyr 1115 1120 1125Ala Pro Ala Ser Ser Val Glu
Tyr Gln Cys Gln Asn Leu Tyr Gln 1130 1135 1140Leu Glu Gly Asn Lys
Arg Ile Thr Cys Arg Asn Gly Gln Trp Ser 1145 1150 1155Glu Pro Pro
Lys Cys Leu His Pro Cys Val Ile Ser Arg Glu Ile 1160 1165 1170Met
Glu Asn Tyr Asn Ile Ala Leu Arg Trp Thr Ala Lys Gln Lys 1175 1180
1185Leu Tyr Ser Arg Thr Gly Glu Ser Val Glu Phe Val Cys Lys Arg
1190 1195 1200Gly Tyr Arg Leu Ser Ser Arg Ser His Thr Leu Arg Thr
Thr Cys 1205 1210 1215Trp Asp Gly Lys Leu Glu Tyr Pro Thr Cys Ala
Lys Arg 1220 1225 12308259PRTArtificial sequenceSynthetic 82Gly Lys
Cys Gly Pro Pro Pro Pro Ile Asp Asn Gly Asp Ile Thr Ser1 5 10 15Phe
Pro Leu Ser Val Tyr Ala Pro Ala Ser Ser Val Glu Tyr Gln Cys 20 25
30Gln Asn Leu Tyr Gln Leu Glu Gly Asn Lys Arg Ile Thr Cys Arg Asn
35 40 45Gly Gln Trp Ser Glu Pro Pro Lys Cys Leu His 50
55838PRTArtificial sequenceSynthetic 83Pro Ile Asp Asn Gly Asp Ile
Thr1 58415PRTArtificial sequenceSynthetic 84Gly Pro Pro Pro Pro Ile
Asp Asn Gly Asp Ile Thr Ser Phe Pro1 5 10 1585368DNAArtificial
sequenceSynthetic 85gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctgggaggtc cctgagactc 60tcctgcgcag cctctggact caccttcagt ttctataatt
tccactgggt ccgccagact 120ccaggcaagg ggctggagtg ggtggcaggc
atctcatacg atgcaaccag gacgaactac 180gcaggctcgt cacgggccga
ttcaccattt ccagagacaa ttccaagaaa atgctgtatc 240tgcaaatgag
cagcctggga cctcaagaca cggctgtata tcattgtgcg agagatcgtt
300ctgacgggca actgcataaa gtggcttttg actcctgggg ccagggagcc
ctggtcaccg 360tctcatca 36886357DNAArtificial sequenceSynthetic
86gaggtgcagc tggtggagtc tgggggaggc gtggtccggc ctgggcggtc cctgagactc
60tcctgtgttg cctctggttt caccttcaat gcttatggca tgcattgggt ccgccagggt
120ccaggcaagg gccttgagtg gctggcggtc atttcatatg aaggaaagac
tgtttattat 180gcagattccg ttaaggaccg tttcaccatc tccagagaca
attccaggaa cacggtgtct 240ctacatctga acaacctgag aggtgaggac
acggctgtct attactgtgc gaaggggtcg 300gcttcagcag cagtcctcca
acactggggt cagggcaccc tggtcagcgt cacgtca 35787357DNAArtificial
sequenceSynthetic 87gaggtgcagc tggtggagtc tgggggaggc gtggtcccgc
ctgggaagtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt ctctatggca
tacactgggt ccgccaggct 120cccggcaagg gactggagtg ggtggcagtt
atctcatatg atggaaatac taaatactat 180acagactctg taaagggtcg
attcaccatc tccagagaca atgccaagaa cacaatttat 240ctgcaaatga
acagtctaag acttgacgac acggctgttt attactgtgc gaaaggagcg
300gcgaatagcg ctacttttga tttctggggc cgagggacaa tggtcaccgt ctcttca
35788357DNAArtificial sequenceSynthetic 88gaggtgcagc tggtggagtc
tgggggaggc gtggtcccgc ctgggaagtc cctgagactc 60tcctgtgcag cctctggatt
caccttcagt ctctatggca tacactgggt ccgccaggct 120cccggcaagg
gactggagtg
ggtggcagtt atctcatatg atggaaatac taaatactat 180acagactctg
taaagggtcg attcaccatc tccagagaca atgccaagaa cacaatttat
240ctgcaaatga acagtctaag acttgacgac acggctgttt attactgtgc
gaaaggagcg 300gcgaatagcg ctacttttga tttctggggc cgagggacaa
tggtcaccgt ctcttca 35789357DNAArtificial sequenceSynthetic
89gaggtgcagc tggtggagtc tgggggaggc gtggtcccgc ctgggaagtc cctgagactc
60tcctgtgcag cctctggatt caccttcagt ctctatggca tacactgggt ccgccaggct
120cccggcaagg gactggagtg ggtggcagtt atctcatatg atggaaatac
taaatactat 180acagactctg taaagggtcg attcaccatc tccagagaca
atgccaagaa cacaatttat 240ctgcaaatga acagtctaag acttgacgac
acggctgttt attactgtgc gaaaggagcg 300gcgaatagcg ctacttttga
tttctggggc cgagggacaa tggtcaccgt ctcttca 35790357DNAArtificial
sequenceSynthetic 90gaggtgcagc tggtggagtc tgggggaggc gtggtcccgc
ctgggaagtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt ctctatggca
tacactgggt ccgccaggct 120cccggcaagg gactggagtg ggtggcagtt
atctcatatg atggaaatac taaatactat 180acagactctg taaagggtcg
attcaccatc tccagagaca atgccaagaa cacaatttat 240ctgcaaatga
acagtctaag acttgacgac acggctgttt attactgtgc gaaaggagcg
300gcgaatagcg ctacttttga tttctggggc cgagggacaa tggtcaccgt ctcttca
35791246DNAArtificial sequenceSynthetic 91gaggtgcagc tggtggagtc
tgggggaggc gtggtccagc ctggaaagtc tctgagactc 60tcctgtgtag cctctggatt
cagcttcagt acttatggca tgcactgggt ccgccaggct 120ccaggcaagg
ggctggagtg ggtggcggtt atgtcatttg atggaaagac taaatactat
180gcagactccg tgaagggccg attcaccatc tccagagaca atcccaagaa
cacactatat 240ctgcaa 24692357DNAArtificial sequenceSynthetic
92gaggtgcagc tggtggagtc tgggggaggc gtggtccagc ctggaaagtc tctgagactc
60tcctgtgtag cctctggatt cagcttcagt acttatggca tgcactgggt ccgccaggct
120ccaggcaagg ggctggagtg ggtggcggtt atgtcatttg atggaaagac
taaatactat 180gcagactccg tgaagggccg attcaccatc tccagagaca
atcccaagaa cacactatat 240ctgcaaatga acagcctgag aagcgaagac
acggctgtgt attattgtgc gaaggggggt 300gcagcagcgg ccgtctttga
ctcctggggc ccgggaatac tgctcaccgt ctcctca 35793357DNAArtificial
sequenceSynthetic 93gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctgggaggtc cctaagactc 60tcctgtgcag cctctggagt caccttcagt agatatggca
tgcactgggt ccgccaggct 120ccaggcaagg ggctggagtg ggtggcagtt
atatcatatg atgaaaagac taaatactat 180gcagactccg tgaagggccg
attcaccatc tccagagaca attccaagaa cacactgttt 240ctgcacatga
acagactgag atatgaggac acggctgtat attattgtgc gaaaggggcc
300agtagcggtg cgtactttga ctactggggc cagggtaccc tggtcaccgt ctcctca
35794357DNAArtificial sequenceSynthetic 94gaggtgcagc tggtggagtc
tgggggaggc gtggtccagc ctggaaagtc tctgagactc 60tcctgtgtag cctctggatt
caccttcagt acttatggca tgcactgggt ccgccaggct 120ccaggcaagg
ggctggagtg ggtggcagtt atgtcatttg atggaaagac taaatactat
180gcagactccg tgaagggccg attcaccatc tccagagaca atcccaagaa
cacactatat 240ctgcaaatga acagcctgag aagcgaagac acggctgtgt
attattgtgc gaaggggggt 300gcagcagcgg ccgtctttga ctcctggggc
cagggaatac tgctcaccgt ctcctca 35795357DNAArtificial
sequenceSynthetic 95gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctggaaagtc tctgagactc 60tcctgtgtag cctctggatt caccttcagt agttatggca
tgcactgggt ccgccaggct 120ccgggcaagg ggctggagtg ggtggcggtt
atgtcatttg atggaaagac taaatactat 180gcagactccg tgaagggccg
attcaccatc tccagagaca accccaagaa cacactatat 240ctgcaaatga
acagcctgag aagcgaagac acggctgtct attattgtgc gaaggggggt
300gcagcagcgg ccgtctttga ctcctggggc cagggaatac tgctcaccgt ctcctca
35796357DNAArtificial sequenceSynthetic 96gaggtgcagc tggtggagtc
tgggggaggc gtggtccagc ctggaaagtc tctgagactc 60tcctgtgtag cctctggatt
caccttcagt agttatggca tgcactgggt ccgccaggct 120ccgggcaagg
ggctggagtg ggtggcggtt atgtcatttg atggaaagac taaatactat
180gcagactccg tgaagggccg attcaccatc tccagagaca accccaagaa
cacactatat 240ctgcaaatga acagcctgag aagcgaagac acggctgtct
attattgtgc gaaggggggt 300gcagcagcgg ccgtctttga ctcctggggc
cagggaatac tgctcaccgt ctcctca 35797356DNAArtificial
sequenceSynthetic 97gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctgggaggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt agatatggca
tgcactgggt ccgccaggct 120ccaggcaagg ggctggagtg ggtggcagtt
atgtcatatg atggaagtac taaatactat 180gcagactccg tgaagggccg
cttcgccatc tccagagaca atcccaagaa cacgctattt 240ctgcaaatga
acagcctgag acctgacgac acggctgtat attactgtgc gaaagggggg
300gcggcacagc tgtcatggac gtctggggca aagggaccac ggtcaccgtc tcctca
35698357DNAArtificial sequenceSynthetic 98gaggtgcagc tggtggagtc
tgggggaggc gtggtccagc ctgggaggtc cctgagactc 60tcctgtgcag cctctggatt
caccttcagt agatatggca tgcactgggt ccgccaggct 120ccaggcaagg
ggctggagtg ggtggcagtt atgtcatatg atggaagtac taaatactat
180gcagactccg tgaagggccg cttcgccatc tccagagaca atcccaagaa
cacgctattt 240ctgcaaatga acagcctgag acctgacgac acggctgtat
attactgtgc gaaagggggg 300gcggcagcag ctgtcatgga cgtctggggc
aaagggacca cggtcaccgt ctcctca 35799363DNAArtificial
sequenceSynthetic 99gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctgggaggtc cctgagactc 60tcctgtgcag cctctggatt caccttcaat aggtttggca
tgcactgggt ccgccagcgc 120caggttccag gcaaggggct ggagtgggtg
gcagttatct catatgacga caacactaaa 180tattatgcgg actccgtgaa
gggccgtttc accatctcca gagacaataa caagagcact 240ctctatctgc
aaatgagcag cctgagagtt gaggacacgg ctgtctattt ctgtgcgaag
300gggtcgacag cggcagctgt tcttgactac tggggccagg gaacccttgt
caccgtctcc 360tca 363100363DNAArtificial sequenceSynthetic
100gaggtgcagc tggtggagtc tgggggaggc ttggtacagc ctggagggtc
cctgagactc 60tcctgtgcag cctctggatt caccttcagt agttatgaaa tgaactgggt
ccgccaggct 120ccagggaagg ggctggagtg ggtttcatac attagtagta
gtggtagtac catatactac 180gcagactctg tgaagggccg attcaccatc
tccagagaca acgccaagaa ctcactgtat 240ctgcaaatga acagcctgag
agccgaggac acggctgttt attactgtgc gagagtagag 300cagctcgccc
cctcccccta catggacgtc tggggcaaag ggaccacggt caccgtctcc 360tca
363101372DNAArtificial sequenceSynthetic 101caggtgcagc tggtgcagtc
tggggcagag gtgaaaaagc ccggggagtc tctgaagatc 60tcctgtaagg gttctggata
cagctttacc agctactgga tcggctgggt gcgccagatg 120cccgggaaag
gcctggagtg gatggggatc atctatcctg gtgactctga taccagatac
180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca agtccatcag
caccgcctac 240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt
attactgtgc gagacggggt 300cttcgagggg cctactacta ctactacggt
atggacgtct ggggccaagg gaccacggtc 360accgtctcct ca
372102339DNAArtificial sequenceSynthetic 102gacatcgtga tgacccagtc
tccagactcc ctgactctgt ctctgggcga gagggccacc 60atcaactgca gatccagccg
gactgtttta tacaggtcca acaataaaaa ttacttagct 120tggtatcaac
ataaaccagg acagcctcct aagttgctca tgtcctgggc atctacccgg
180gaaaccgggg tccctgaccg attcagtggc agcggttctg ggacacattt
cactctcacc 240atcaccagcc tgcagcctga agatgtggca gtttattact
gtcaacagta ttatagtcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaga
339103246DNAArtificial sequenceSynthetic 103gacatcgtga tgacccagtc
tccaggctcc ctggctgtgt ctctgggctc gagggccacc 60atcaactgca agtccagccg
gagtctttta tacaggtcca acaataagaa ttatttagct 120tggtatcaac
agaaaccagg acagtctcct cggcttctca tttattgggc atcttcccgg
180gaatccgggg tccctgaccg attcagtggc ggcgggtctg ggacaagttt
cactctcacc 240atcagc 246104339DNAArtificial sequenceSynthetic
104gacatcgtga tgacccagtc tccagactcc ctgactctgt ctctgggcga
gagggccacc 60atcaactgca gatccagccg gactgtttta tacaggtcca acaataaaaa
ttacttagct 120tggtatcaac ataaaccagg acagcctcct aagttgctca
tgtcctgggc atctacccgg 180gaaaccgggg tccctgaccg attcagtggc
agcggttctg ggacacattt cactctcacc 240atcaccagcc tgcagcctga
agatgtggca gtttattact gtcaacagta ttatagtcct 300ccgtggacgt
tcggccaagg gaccaaggtg gaaatcaga 339105339DNAArtificial
sequenceSynthetic 105gacatcgtga tgacccagtc tccagactcc ctgactctgt
ctctgggcga gagggccacc 60atcaactgca gatccagccg gactgtttta tacaggtcca
acaataaaaa ttacttagct 120tggtatcaac ataaaccagg acagcctcct
aagttgctca tgtcctgggc atctacccgg 180gaaaccgggg tccctgaccg
attcagtggc agcggttctg ggacacattt cactctcacc 240atcaccagcc
tgcagcctga agatgtggca gtttattact gtcaacagta ttatagtcct
300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaga
339106339DNAArtificial sequenceSynthetic 106gacatcgtga tgacccagtc
tccagactcc ctgactctgt ctctgggcga gagggccacc 60atcaactgca gatccagccg
gactgtttta tacaggtcca acaataaaaa ttacttagct 120tggtatcaac
ataaaccagg acagcctcct aagttgctca tgtcctgggc atctacccgg
180gaaaccgggg tccctgaccg attcagtggc agcggttctg ggacacattt
cactctcacc 240atcaccagcc tgcagcctga agatgtggca gtttattact
gtcaacagta ttatagtcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaga
339107339DNAArtificial sequenceSynthetic 107gacatcgtga tgacccagtc
tccagactcc ctgactctgt ctctgggcga gagggccacc 60atcaactgca gatccagccg
gactgtttta tacaggtcca acaataaaaa ttacttagct 120tggtatcaac
ataaaccagg acagcctcct aagttgctca tgtcctgggc atctacccgg
180gaaaccgggg tccctgaccg attcagtggc agcggttctg ggacacattt
cactctcacc 240atcaccagcc tgcagcctga agatgtggca gtttattact
gtcaacagta ttatagtcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaga
339108339DNAArtificial sequenceSynthetic 108gacatcgtga tgacccagtc
tccaaactcc ctggctgtgt ctctgggcgg gagggccacc 60atcaactgca aggccagcca
gagtatttta tacaggtcca acaataagaa ctatttagct 120tggtaccaac
ataaagcagg acagcctccc aagctgctca tttactgggc atctacccgg
180gaatccgggg tccctgagcg attcagtggc agcgggtcta ggacagattt
cactctcacc 240atcaacggcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaaa
339109339DNAArtificial sequenceSynthetic 109gacatcgtga tgacccagtc
tccaaactcc ctggctgtgt ctctgggcgg gagggccacc 60atcaactgca aggccagcca
gagtatttta tacaggtcca acaataagaa ctatttagct 120tggtaccaac
ataaagcagg acagcctccc aagctgctca tttactgggc atctacccgg
180gaatccgggg tccctgagcg attcagtggc agcgggtcta ggacagattt
cactctcacc 240atcaacggcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaaa
339110339DNAArtificial sequenceSynthetic 110gacatcgtga tgacccagtc
tccagactcc ctggctgtgt ctctgggcga gagggccacc 60atcaagtgca agtccagcca
gagtgtcttg tacagctcca acaataagaa ctacttagct 120tggtaccagc
ataaaccagg acagcctcct aaggtactcg tttactgggc atccacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtctg ggacagattt
cactctcacc 240atcagcagcc tgcaggctga agatgtggca gtttattact
gtcaacaata ttataatcct 300ccgtggacgt tcggccaagg gaccaaggta gcaatcaag
339111339DNAArtificial sequenceSynthetic 111gacatcgtga tgacccagtc
tccaaactcc ctggctgtgt ctctgggcgg gagggccacc 60atcaactgca agaccagcca
gagtatttta tacaggtcca acaataagaa ctacttagct 120tggtaccagc
ataaaccagg acagcctccc aagctgctca tttactgggc atctacccgg
180gaatccaggg tccctgaccg attcagtggc agcgggtcta ggacagattt
cactctcacc 240atcagcggcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccgtggacgt tcggccaggg gaccaaggtg gaaatcaaa
339112339DNAArtificial sequenceSynthetic 112gacatcgtga tgacccagtc
tccaaactcc ctggctgtgt ctctgggcgg gagggccacc 60atcaactgca agaccagcca
gagtatttta tacaggtcca acaataagaa ctacttagct 120tggtaccagc
ataaatcagg acagcctccc aagctgctca tttactgggc atctacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtcta ggacagattt
cactctcacc 240atcagcggcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaaa
339113339DNAArtificial sequenceSynthetic 113gacatcgtga tgacccagtc
tccaaactcc ctggctgtgt ctctgggcgg gagggccacc 60atcaactgca agaccagcca
gagtatttta tacaggtcca acaataagaa ctacttagct 120tggtaccagc
ataaatcagg acagcctccc aagctgctca tttactgggc atctacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtcta ggacagattt
cactctcacc 240atcagcggcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccgtggacgt tcggccaagg gaccaaggtg gaaatcaaa
339114339DNAArtificial sequenceSynthetic 114gacatcgtga tgacccagtc
tccggactcc ctgactgtgt ctctgggcga gagggccacc 60atcagctgca agtccagcca
gcgtcttttg tatagttcca acaataagaa ctacttagct 120tggtaccagc
agaaacctgg acagcctcct aaactgctca tgtactgggc gtccacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtctg ggacagattt
ctctctcacc 240atcagcagcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccctggacgt tcggccaagg gaccaaggtg gaagtcaaa
339115339DNAArtificial sequenceSynthetic 115gacatcgtga tgacccagtc
tccggactcc ctgactgtgt ctctgggcga gagggccacc 60atcagctgca agtccagcca
gcgtcttttg tatagttcca acaataagaa ctacttagct 120tggtaccagc
agaaacctgg acagcctcct aaactgctca tgtactgggc gtccacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtctg ggacagattt
ctctctcacc 240atcagcagcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttataatcct 300ccctggacgt tcggccaagg gaccaaggtg gaagtcaaa
339116339DNAArtificial sequenceSynthetic 116gacatcgtga tgacccagtc
tccagattcc ctgactctgt ctctgggcga gagggccacc 60atcaactgca agtccagcca
gagtctgttt tacaggtcca acaataagag ctacttagct 120tggtatcagc
aaaaaccagg gcagcctcct aaactgctca tttactgggc ctctgtccgg
180gaatccgggg tccctgaccg attcactggc agcgggtctg taacagattt
cactctcacc 240atcagcagcc tgcgggctga ggatgtggct gtttattatt
gtcaacagta ttttactact 300cctctcactt tcggcggggg gaccaaggtg gcgatcaaa
339117339DNAArtificial sequenceSynthetic 117gacatcgtga tgacccagtc
tctagactcc ctgactgtgt ctctgggcga gagggccacc 60atcaactgca agtccagcca
gagtctttta tacacctcca acaataagaa ttacttagct 120tggtaccagc
agaaatcagg acagcctcct aagttactca tttactgggc gtctattcgg
180gattccgggg tccctgaccg attcagtggc agcgggtctg cgacagattt
cactctcacc 240atcaacaacc tgcaggctga agatgtggca gtttacttct
gtcagcaata ttacaagact 300cctctcactt tcggcggggg gaccaaggtg gaggtcaga
339118321DNAArtificial sequenceSynthetic 118gacatccagw tgacccagtc
tccttccacc ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgcc gggccagtca
gagtattagt agctggttgg cctggtatca gcagaaacca 120gggaaagccc
ctaagctcct gatctataag gcgtctagtt tagaaagtgg ggtcccatca
180aggttcagcg gcagtggatc tgggacagaa ttcactctca ccatcagcag
cctgcagcct 240gatgattttg caacttatta ctgccaacag tataatagtt
attcttggac gttcggccaa 300gggaccaagg tggaaatcaa a
3211196PRTArtificial sequenceSynthetic 119His His His His His His1
51205DNAArtificial sequenceSynthetic 120ddddk 51216PRTArtificial
sequenceSynthetic 121Ala Asp Asp Asp Asp Lys1 512215PRTArtificial
sequenceSynthetic 122Gly Pro Pro Pro Pro Ile Asp Asn Gly Asp Ile
Thr Ser Phe Pro1 5 10 15
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
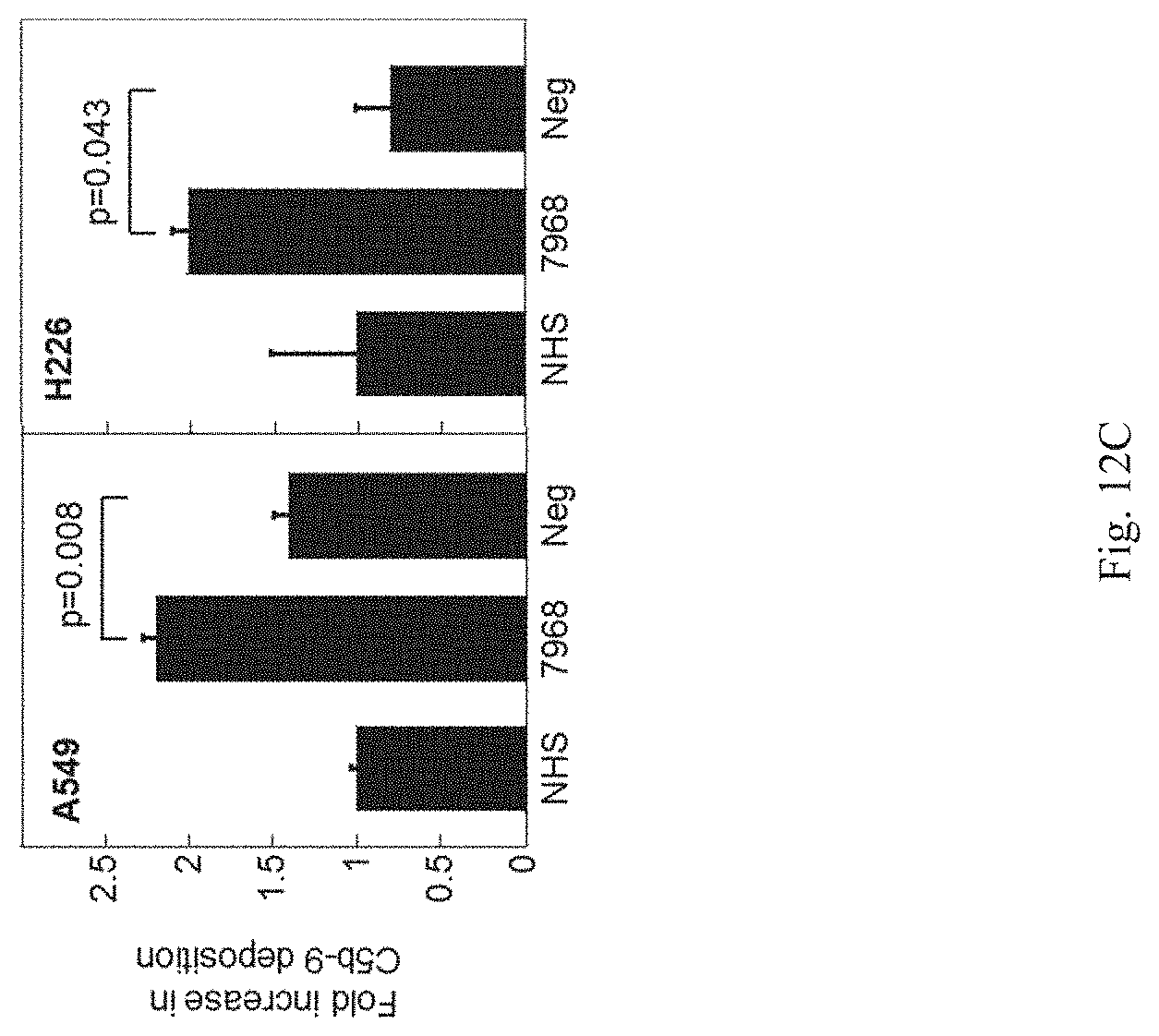
D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.