Methods And Compositions For Use Of Recombinant Bacterial Effector Proteins As Anti-inflammatory Agents
Rauscher; Frank J. ; et al.
U.S. patent application number 16/964762 was filed with the patent office on 2021-02-25 for methods and compositions for use of recombinant bacterial effector proteins as anti-inflammatory agents. The applicant listed for this patent is Innate Biologics LLC, The Wistar Institute of Anatomy and Biology. Invention is credited to Peter Mondics, Frank J. Rauscher.
| Application Number | 20210054033 16/964762 |
| Document ID | / |
| Family ID | 1000005221403 |
| Filed Date | 2021-02-25 |



View All Diagrams
| United States Patent Application | 20210054033 |
| Kind Code | A1 |
| Rauscher; Frank J. ; et al. | February 25, 2021 |
METHODS AND COMPOSITIONS FOR USE OF RECOMBINANT BACTERIAL EFFECTOR PROTEINS AS ANTI-INFLAMMATORY AGENTS
Abstract
Provided herein are methods and compositions comprising a set of paired peptides comprising a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof. The paired peptides can be linked to a protein transduction domain. The compositions can be formulated as pharmaceuticals. The compositions are useful for the treatment of inflammatory disorders.
| Inventors: | Rauscher; Frank J.; (Wayne, PA) ; Mondics; Peter; (Newtown Square, PA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005221403 | ||||||||||
| Appl. No.: | 16/964762 | ||||||||||
| Filed: | January 25, 2019 | ||||||||||
| PCT Filed: | January 25, 2019 | ||||||||||
| PCT NO: | PCT/US2019/015224 | ||||||||||
| 371 Date: | July 24, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 62622068 | Jan 25, 2018 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2319/03 20130101; C07K 14/25 20130101; A61K 38/00 20130101; C07K 14/255 20130101; C07K 2319/30 20130101; C07K 2319/70 20130101 |
| International Class: | C07K 14/255 20060101 C07K014/255; C07K 14/25 20060101 C07K014/25 |
Claims
1. A composition comprising a set of paired peptides, wherein the set of paired peptides is linked to a protein transduction domain, and wherein the set of paired peptides comprises a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof.
2. The composition of claim 1, wherein the first bacterial effector polypeptide or fragment thereof and second bacterial effector polypeptide or fragment thereof are different.
3. The composition of claim 1, wherein a first bacterial effector polypeptide or fragment thereof and the second bacterial effector polypeptide or fragment thereof are immunomodulatory.
4. The composition of claim 1, wherein the first bacterial effector polypeptide or fragment thereof and the second bacterial effector polypeptide or fragment thereof recognize a different molecular target or modulate a different inflammatory pathway.
5. The composition of claim 4, wherein the inflammatory pathway is the NFkB pathway, the JNK pathway, the p38 pathway or the STING pathway.
6. The composition of claim 1, wherein the protein transduction domain and the set of paired peptides comprise a fusion protein.
7. The fusion protein of claim 6, further comprising one or more linkers.
8. The fusion protein of claim 7, wherein the linker is positioned between the first bacterial effector polypeptide or fragment thereof and the second bacterial effector polypeptide or fragment thereof.
9. The composition of claim 1, wherein the protein transduction domain is a YopM protein transduction domain, an SspH1 protein transduction domain, or an lpaH protein transduction domain.
10. The composition of claim 9, wherein the protein transduction domain is a YopM protein transduction domain.
11. The composition of claim 10, wherein the protein transduction domain comprises SECS ID NO. 5.
12. The composition of claim 1, wherein the protein transduction domain is selected from the group consisting of Poly-Arg, Tat and VP22, df Tat, a cyclic CPPs, IMT-P8, seven arginine (R7) and Streptolysin 0 (SLO)-mediated systems, elastin like polypeptide, CPP-adaptor system, 1, 2-Benzisothiazolin-3-one (BIT) and Tat, activatable cell-penetrating peptides, LDP12, BR2, POD, native protein independent of R11-CPP, Poly-arginine/Tat and Tat-PTO, Pep-1, CADY-2, R8, azo-R8, Penetratin, HR9 and IR9 peptides, or pVEC.
13. The composition of claim 1, wherein the first bacterial effector polypeptide or fragment thereof is a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, lpaH9.8, lpaH1.4, lpaH2.5, lpaH4.5, lpaH7.8 and SlrP, and the second bacterial effector polypeptide or fragment thereof is a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, lpaH9.8, lpaH1.4, lpaH2.5, lpaH4.5, lpaH7.8 and SlrP.
14. The composition of claim 13, wherein the first bacterial effector polypeptide or fragment thereof is a polypeptide having 90% sequence identity to an amino acid sequence set forth in the group consisting of SEQ ID NOs 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79 and the second bacterial effector polypeptide or fragment thereof is a polypeptide having 90% sequence identity to an amino acid sequence set forth in the group consisting of SEQ ID NOs.3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79.
15. The composition of claim 1, wherein (a) the first bacterial effector polypeptide or fragment thereof is a YopM polypeptide or a fragment thereof or an NLeE polypeptide or a fragment thereof; or (b) the second bacterial effector polypeptide or fragment thereof is a YopM polypeptide or a fragment thereof or an NLeE polypeptide or a fragment thereof.
16. (canceled)
17. The composition of claim 1, wherein the first bacterial effector polypeptide or fragment thereof is a YopM polypeptide or a fragment thereof and the second bacterial effector polypeptide or fragment thereof is an NLeE polypeptide or a fragment thereof.
18. The fusion protein of claim 6, wherein the amino acid sequence is at least 85% identical to the sequence set forth in SEQ ID NOs. 10, 13, 16, 19, 22, or 24.
19-35. (canceled)
36. A fusion protein comprising a set of paired peptides, wherein the set of paired peptides comprises a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof.
37-56. (canceled)
57. A composition comprising a protein transduction domain polypeptide linked to two or more bacterial effector polypeptides or fragments thereof.
58. (canceled)
Description
FIELD OF THE INVENTION
[0001] The present invention relates to methods and compositions for treating inflammatory disorders.
BACKGROUND OF THE INVENTION
[0002] Most treatments for acute inflammation, such as skin inflammatory conditions, only treat the symptoms (swelling, redness, pain, heat) by using non-specific drugs like corticosteroids and emollients/skin softeners. None of these non-specific drugs affect the underlying mechanism of activation of the inflammatory pathway, e.g., NFkB/JNK/p38. Moreover, normal aging processes chronically activate ectopically these three signal transduction pathways, resulting in inflammation. The inflammation makes these pathways a target for inhibition as part of an anti-aging/wellness program.
[0003] To date, the cosmeceutical industry's approach to reducing skin inflammation has many flaws. The "Active Compounds" contained in most cremes and topicals do not have defined, specific targets in the inflamed cell/tissue the cell and are extremely complex mixtures/extracts/serums containing millions of ingredients with no defined targets, or ingredients which are not absorbed by skin and have no effect at all. Because these compositions do not target the actual pathways which are causing inflammation, they have the potential to alter many processes non-specifically leading to toxic side effects.
[0004] A continuing need in the art exists for new and effective tools and methods for treating the causes of inflammation.
SUMMARY OF THE INVENTION
[0005] Disclosed herein are compositions of paired peptides comprising a first bacterial effector polypeptide linked to a second bacterial effector polypeptide that are useful for treating an inflammatory disorder. Accordingly, disclosed are compositions that include a set of paired peptides, wherein the set of paired peptides is linked to a protein transduction domain, and wherein the set of paired peptides comprises a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof. The first and second bacterial effector polypeptides can be different, that is they can recognize a different molecular targets or modulate different inflammatory pathways. In an embodiment, the protein transduction domain and the set of paired peptides can be a fusion protein. The fusion protein can include one or more linkers. The protein transduction domain can be a YopM protein transduction domain, an SspH1 protein transduction domain, or an IpaH protein transduction domain. The first bacterial effector polypeptide or fragment thereof can be a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP, and the second bacterial effector polypeptide or fragment thereof can be a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP. In some embodiments, the first bacterial effector polypeptide or fragment thereof can be a polypeptide having 90% sequence identity to an amino acid sequence set forth in the group consisting of SEQ ID NOs 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79 and the second bacterial effector polypeptide or fragment thereof can be a polypeptide having 90% sequence identity to an amino acid sequence set forth in the group consisting of SEQ ID NOs.3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79. In some embodiments, the first bacterial effector polypeptide or fragment thereof can be a YopM polypeptide or a fragment thereof and the second bacterial effector polypeptide or fragment thereof can be an NLeE polypeptide or a fragment thereof. In some embodiments, the fusion protein can have an amino acid sequence forth in SEQ ID NO. 10, 13, 16, 19, 22, or 24.
[0006] Also provided are fusion proteins comprising a set of paired peptides wherein the set of paired peptides comprises a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof. The first and second bacterial effector polypeptides can be different, that is they can recognize a different molecular targets or modulate different inflammatory pathways. The fusion protein can include one or more linkers. The first bacterial effector polypeptide or fragment thereof can be a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP, and the second bacterial effector polypeptide or fragment thereof can be a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP. In some embodiments, the first bacterial effector polypeptide or fragment thereof can be a polypeptide having 90% sequence identity to an amino acid sequence set forth in the group consisting of SEQ ID NOs 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79 and the second bacterial effector polypeptide or fragment thereof can be a polypeptide having 90% sequence identity to an amino acid sequence set forth in the group consisting of SEQ ID NOs.3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79. In some embodiments, the first bacterial effector polypeptide or fragment thereof can be a YopM polypeptide or a fragment thereof and the second bacterial effector polypeptide or fragment thereof can be an NLeE polypeptide or a fragment thereof. In some embodiments, the fusion protein comprising a set of paired peptides can have an amino acid sequence as set forth in SEQ ID NO. 80, 81, 82, 83, 84, 85, 86, 87, or 88.
[0007] Also provided are nucleic acids encoding a set of paired peptides, wherein the set of paired peptides is linked to a protein transduction domain, and wherein the set of paired peptides comprises a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof. Also provided are nucleic acids encoding fusion proteins comprising a set of paired peptides wherein the set of paired peptides comprises a first bacterial effector polypeptide or fragment thereof linked to a second bacterial effector polypeptide or fragment thereof. The nucleic acids can be contained within a vector, which can be expressed in a host cell. In one aspect, the compositions comprising a set of paired peptides can be formulated as pharmaceutical compositions.
[0008] Also provided are methods of treating a subject having or at risk for an inflammatory disorder, by administering to the subject a therapeutically effective amount of a pharmaceutical composition comprising the set of paired peptides. The inflammatory disorder can be a gastrointestinal disorder including inflammatory bowel disease, Crohn's disease and the ileocolitis, ileocecal, jeunoileitis, and gastroduodenal subtypes of-Crohn's disease, and ulcerative colitis. The inflammatory disorder can also be a skin disorder.
[0009] Also provided are articles of manufacture, e.g., a kit. The kit can include measured amount of one or more of the compositions of the paired peptides and one or more items selected from the group consisting of packaging material, a package insert comprising instructions for use, a sterile fluid, and a sterile container.
[0010] In one aspect, a composition comprises in a pharmaceutically acceptable carrier or excipient or formulation a first construct comprising a selected immunomodulatory effector protein or functional equivalent thereof that targets a first functional domain, optionally linked covalently or non-covalently to a selected protein transduction domain (PTD) or penetrating peptide (CPP). In one embodiment, the composition further comprises an additional construct comprising a different effector protein or a functional equivalent thereof that targets an additional functional domain, optionally linked to the same PTD or CPP or to an additional PTD or CPP. In one embodiment, a composition comprises multiple first and additional constructs. In another embodiment, the constructs are further associated with targeting moieties directing delivery of the constructs to a selected cell or tissue.
[0011] In another aspect, a recombinant polypeptide comprises a first construct comprising a selected immunomodulatory effector protein or functional equivalent thereof that targets a first functional domain, optionally linked covalently or non-covalently to a selected protein transduction domain (PTD) or penetrating peptide (CPP) and an additional construct comprising a different effector protein or a functional equivalent thereof that targets an additional functional domain, optionally linked to the same PTD or CPP or to an additional PTD or CPP. The first construct is linked covalently or non-covalently to one or more of the additional constructs in a single polypeptide. In another embodiment, the polypeptide comprises an optional linker amino acid sequence interposed between each first and additional construct. In another embodiment, the polypeptide is further associated with targeting moieties directing delivery of the polypeptide to a selected cell or tissue.
[0012] In another aspect, a recombinant nucleic acid molecule is provided which encodes one of the constructs or polypeptides described herein. These nucleic acid molecules can be further associated with regulatory sequences for expressing the constructs in vivo or in vitro.
[0013] In a further aspect, a pharmaceutical or cosmeceutical composition comprises as an active agent a polypeptide as described above, or a mixture of constructs as described above in a formulation suitable for delivery of the active agent into and through the layers of the skin. In one embodiment, the formulation contains a CAGE solvent (defined below) or other components suitable for topical administration.
[0014] In yet another aspect, a pharmaceutical or cosmeceutical composition comprises as an active agent a polypeptide as described above, or a mixture of constructs as described above in a formulation suitable for delivery to a selected cell or tissue.
[0015] In still other aspects, methods for making the compositions, constructs, polypeptides and nucleic acid molecules are provided.
[0016] In yet a further aspect, a method for treating or ameliorating or suppressing an inflammatory response comprises administering to a subject in need thereof a composition, construct, polypeptide or nucleic acid molecule described herein.
[0017] Still other aspects and advantages of these compositions and methods are described further in the following detailed description of the preferred embodiments thereof.
BRIEF DESCRIPTION OF THE DRAWINGS
[0018] These and other features and advantages of the present invention will be more fully disclosed in, or rendered obvious by, the following detailed description of the preferred embodiment of the invention, which is to be considered together with the accompanying drawings wherein like numbers refer to like parts and further wherein:
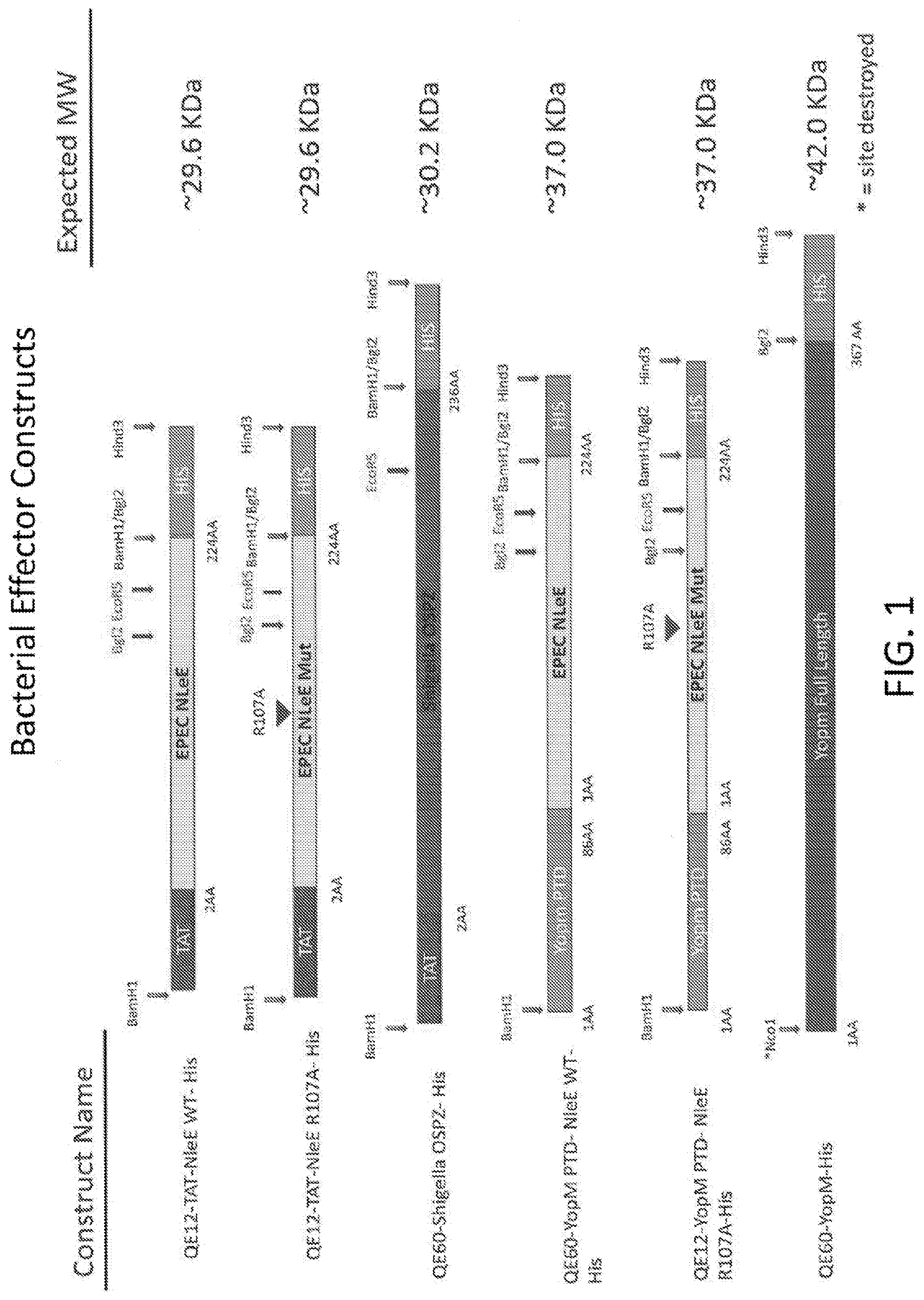
[0019] FIG. 1 is a schematic of bacterial effector constructs.
[0020] FIG. 2 is a schematic of bacterial effector constructs.
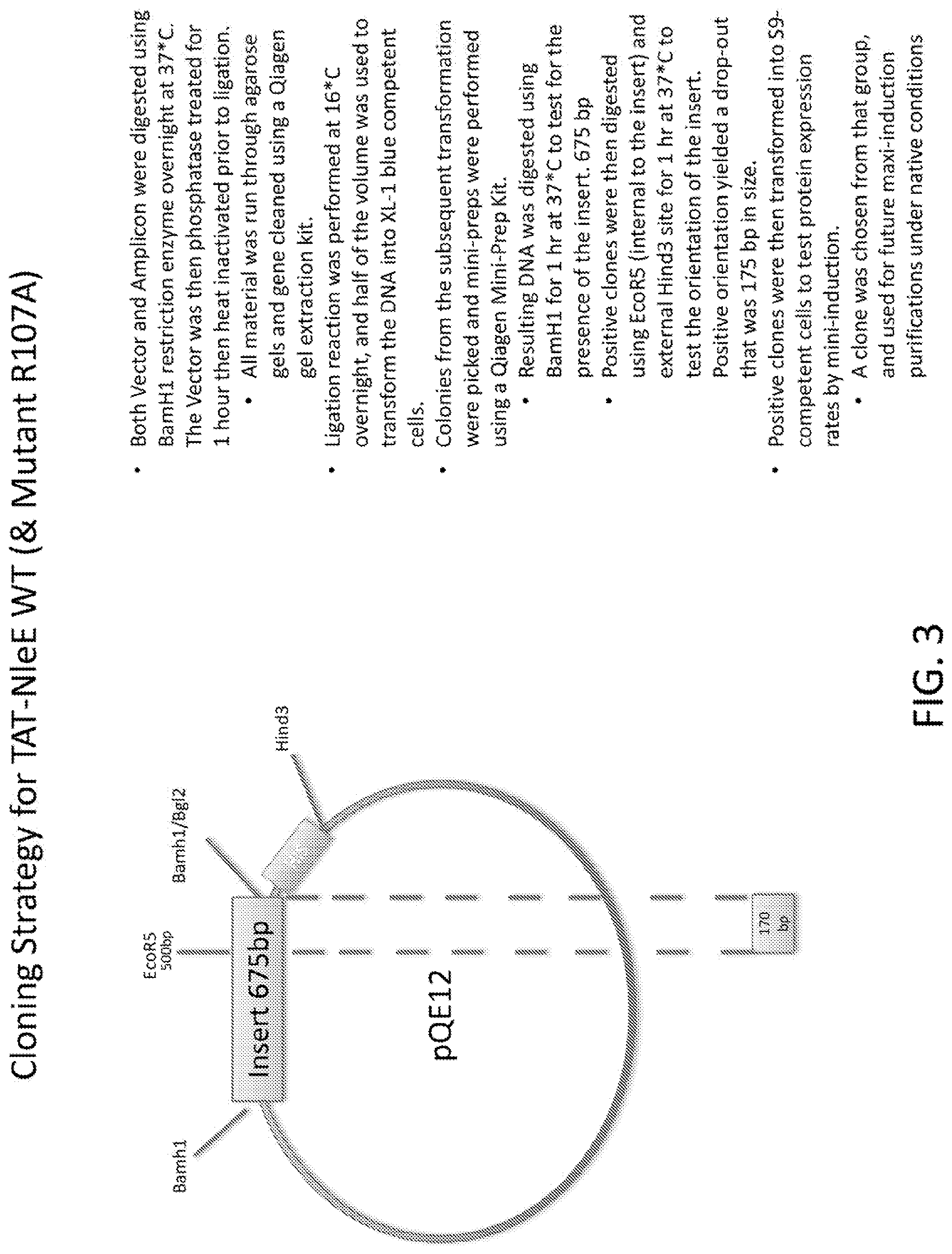
[0021] FIG. 3 is a schematic of the cloning strategy for TAT-NleE WT and Mutant R107A.
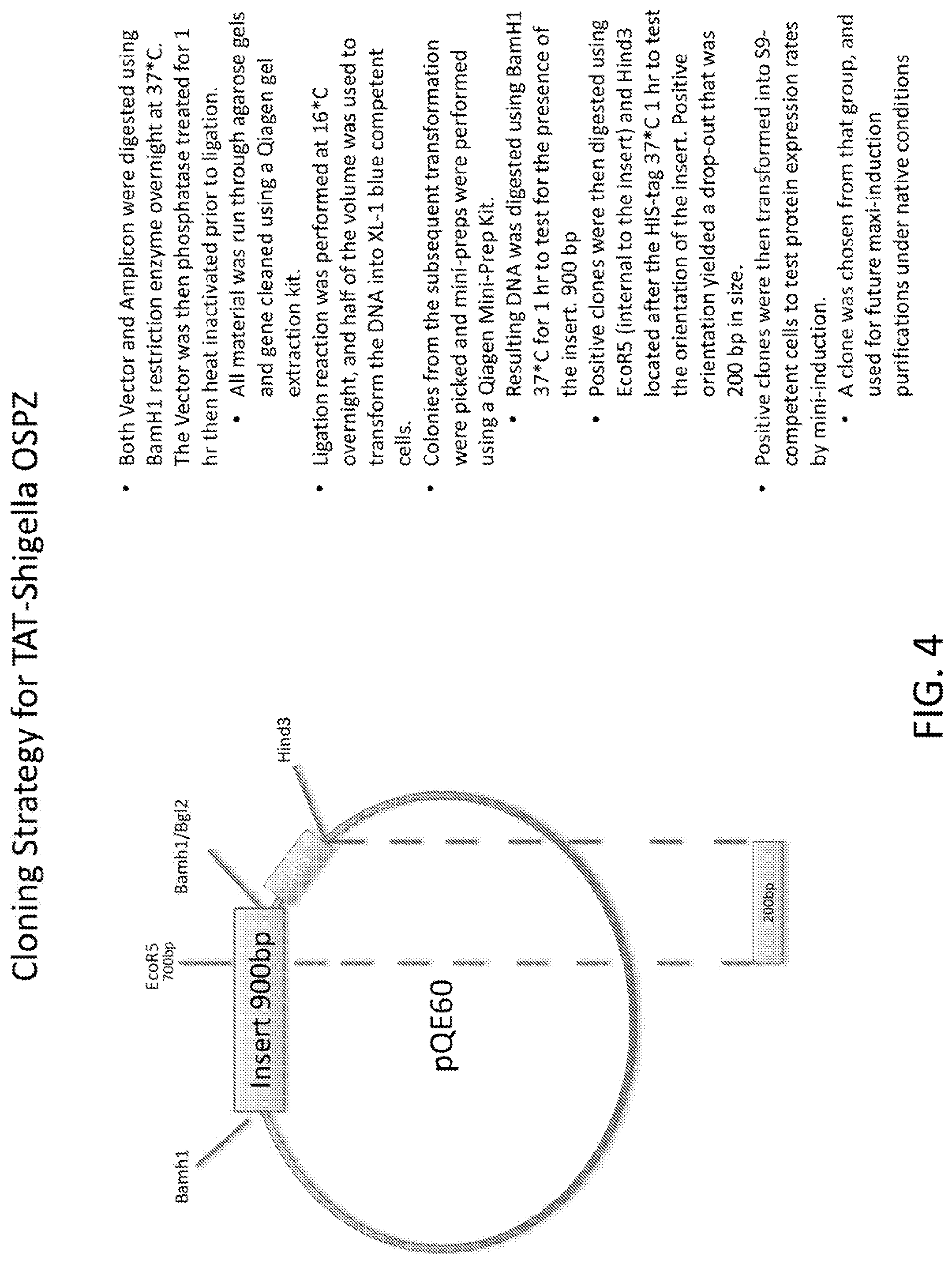
[0022] FIG. 4 is a schematic of the cloning strategy for TAT-Shigella OSPZ.
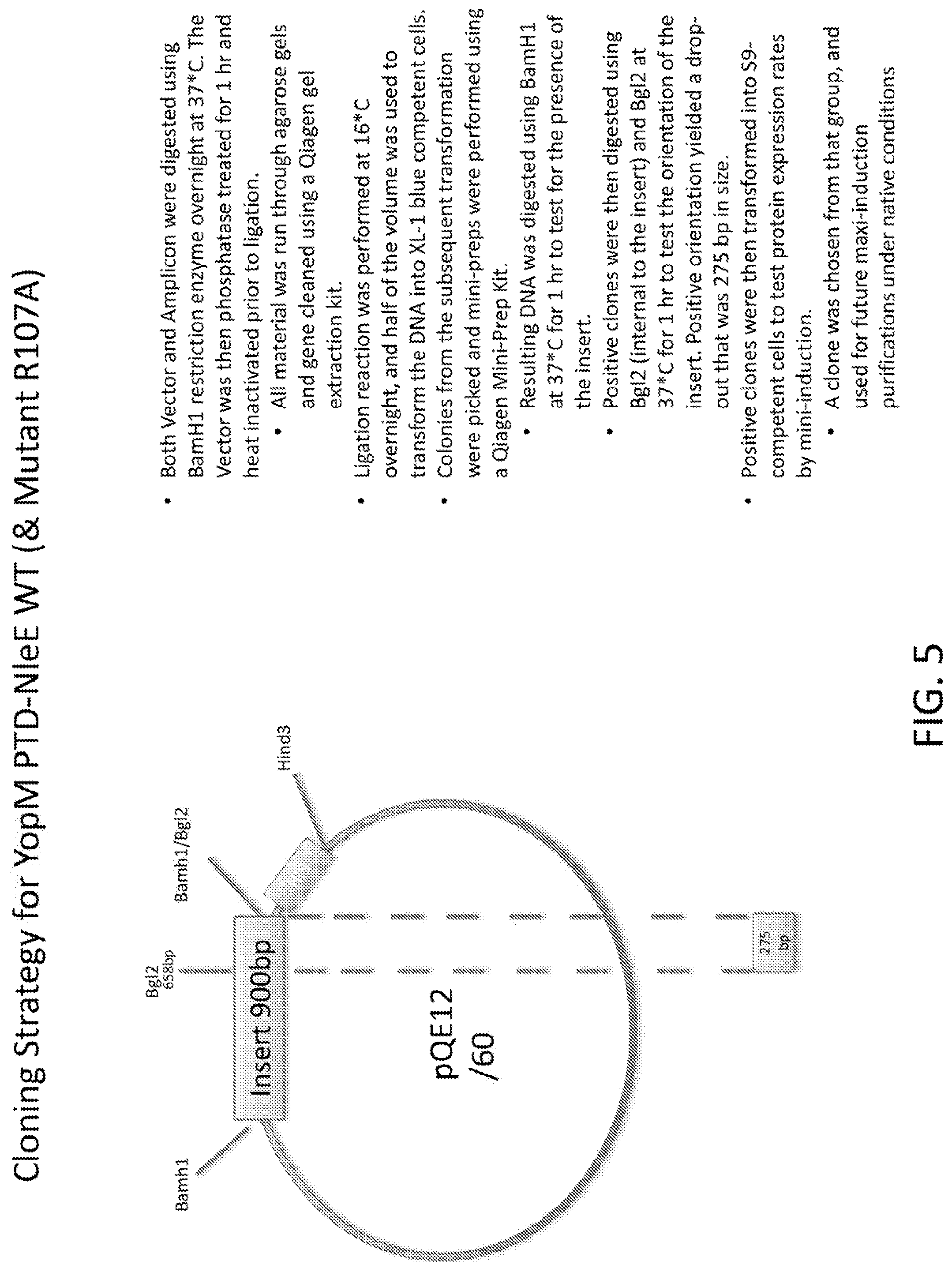
[0023] FIG. 5 is a schematic of the cloning strategy for YopM PTD-NleE WT and Mutant R107A.
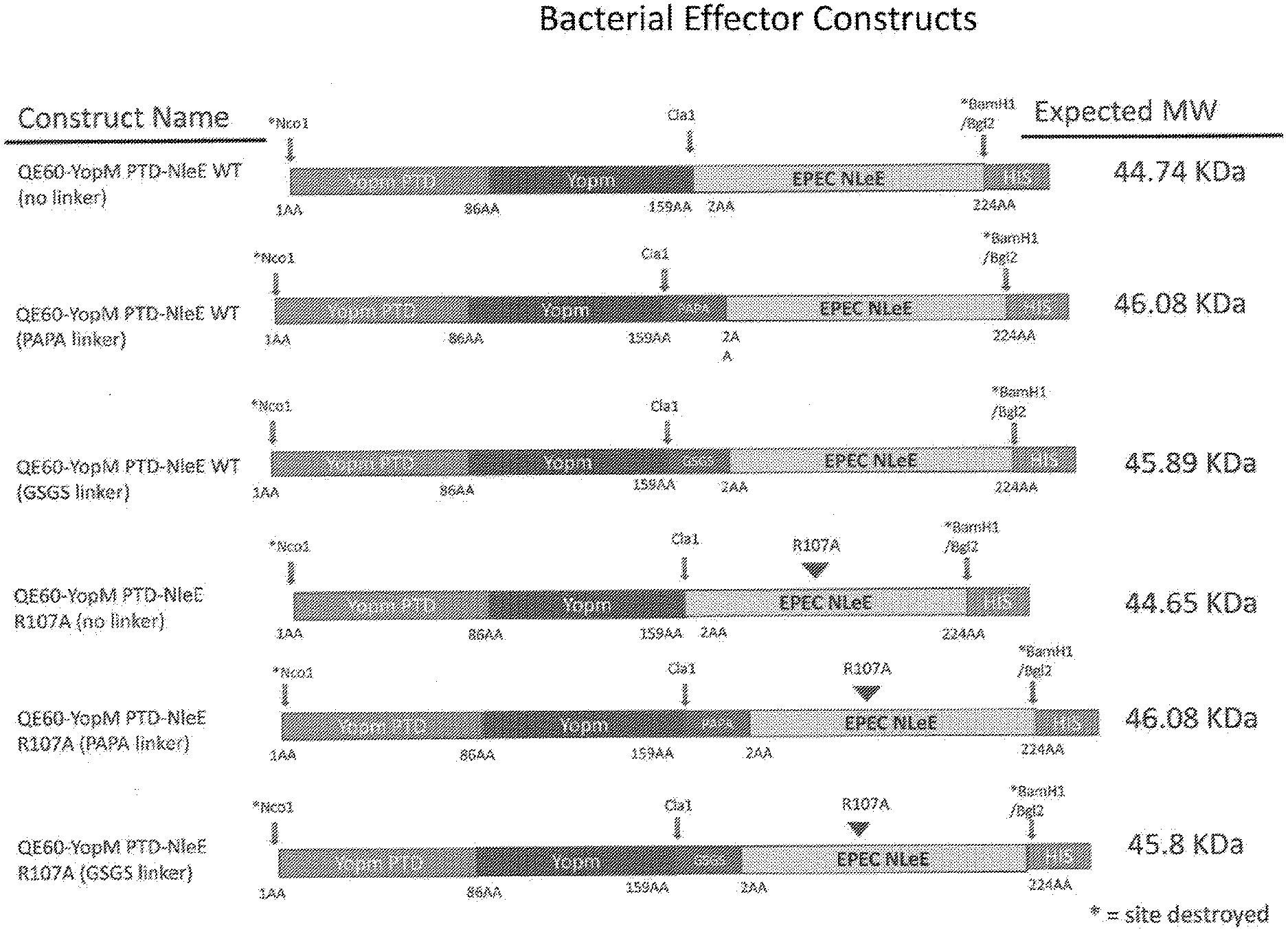
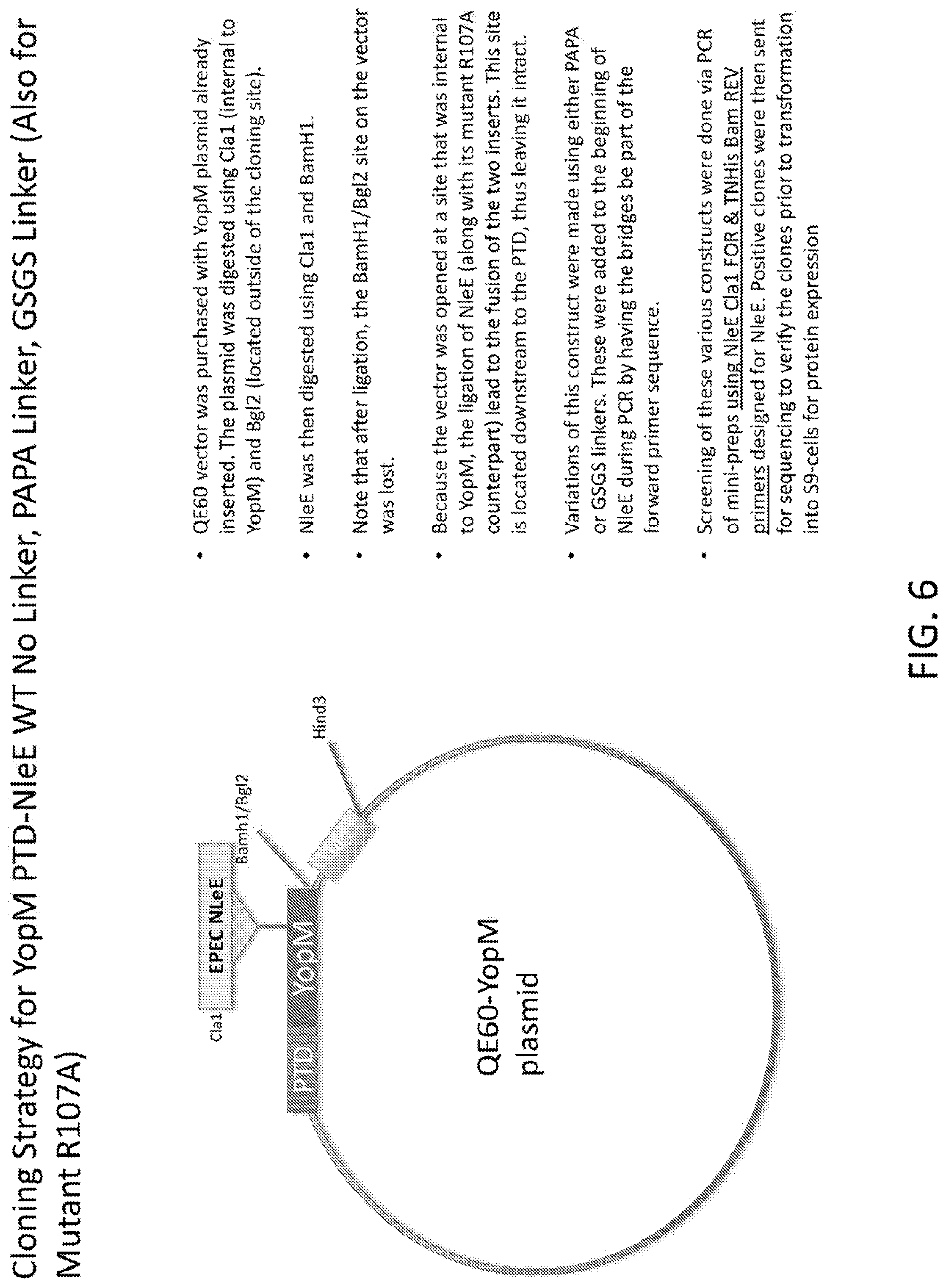
[0024] FIG. 6 is a schematic of the cloning strategy for YopM PTD-NleE WT No Linker, PAPA Linker, GSGS Linker and for Mutant R107A.
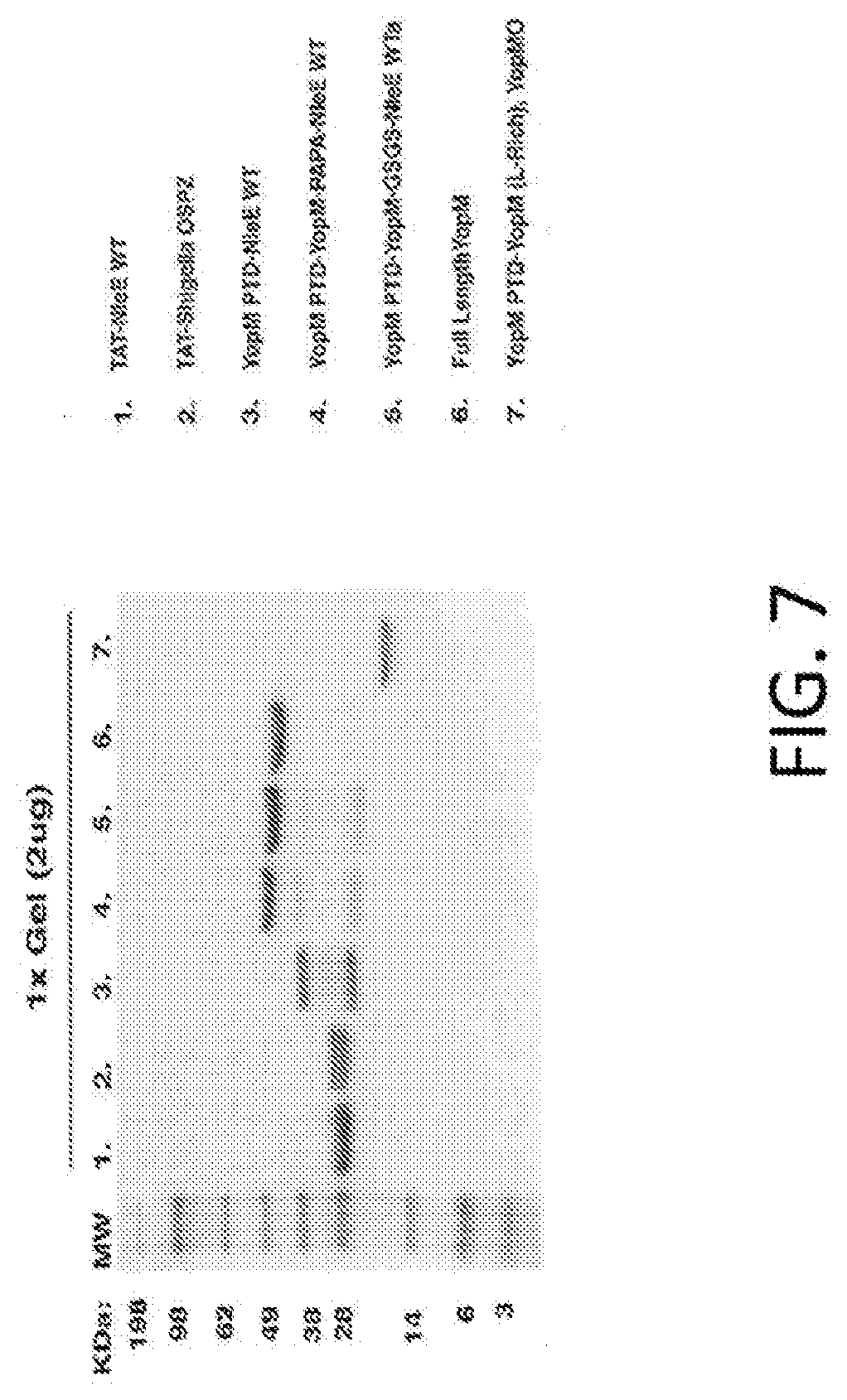
[0025] FIG. 7 is an exemplary SDS gel showing purified fusion proteins.
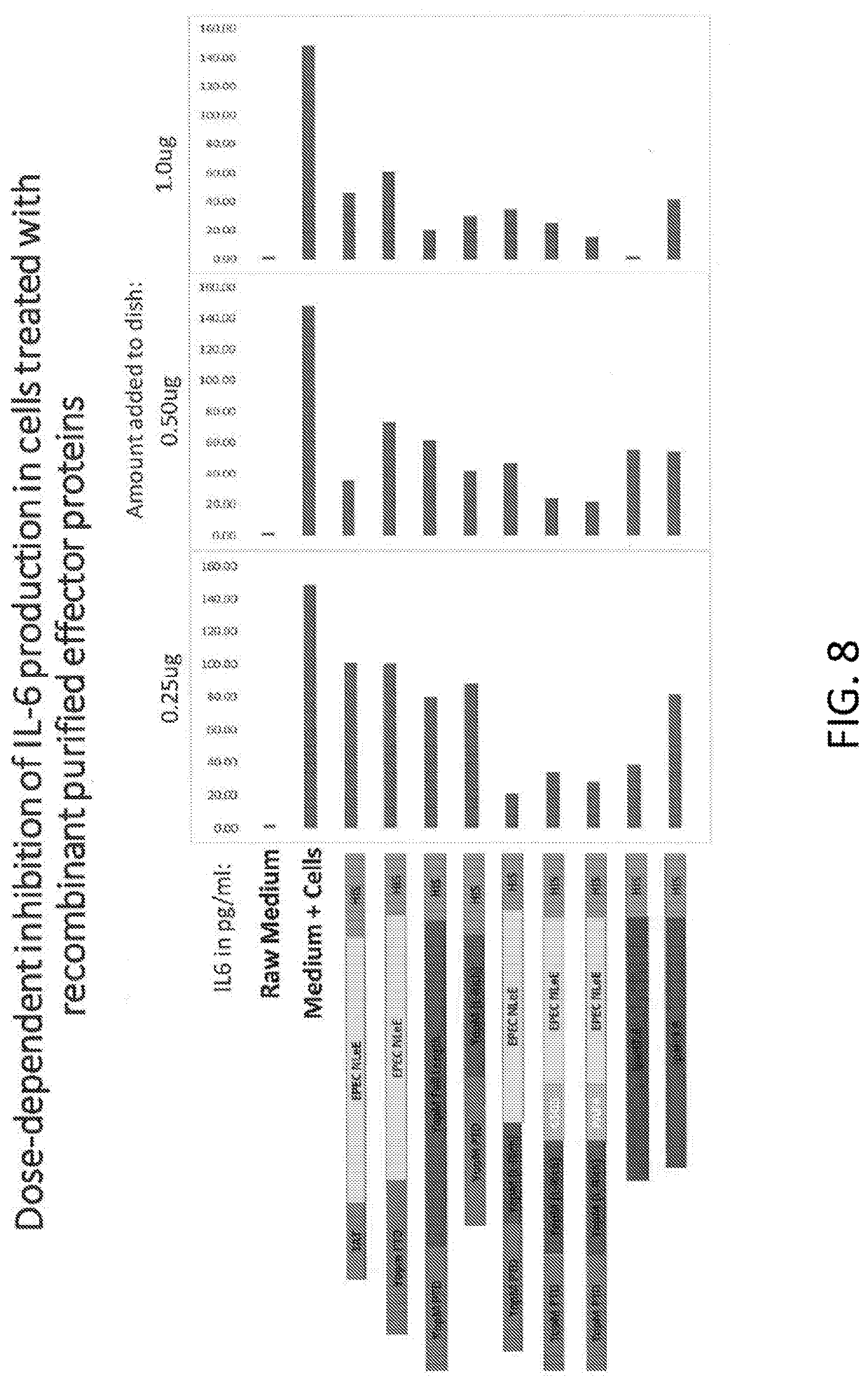
[0026] FIG. 8 is a graph showing the results of a dose-response analysis of IL-6 production in cells treated with recombinant purified effector proteins.
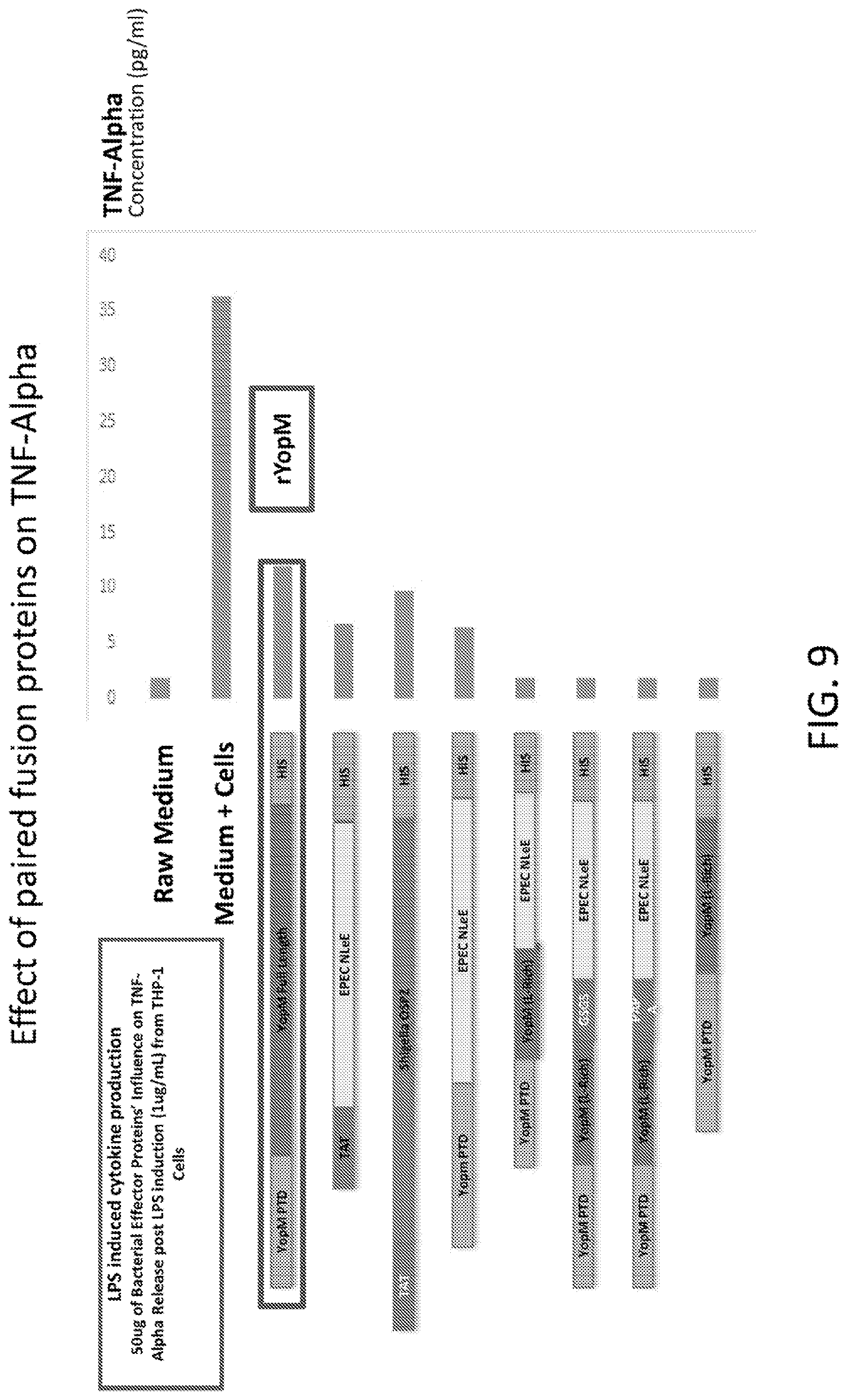
[0027] FIG. 9 is a graph showing the results of an analysis of the effect of paired fusion proteins on TNF-alpha.
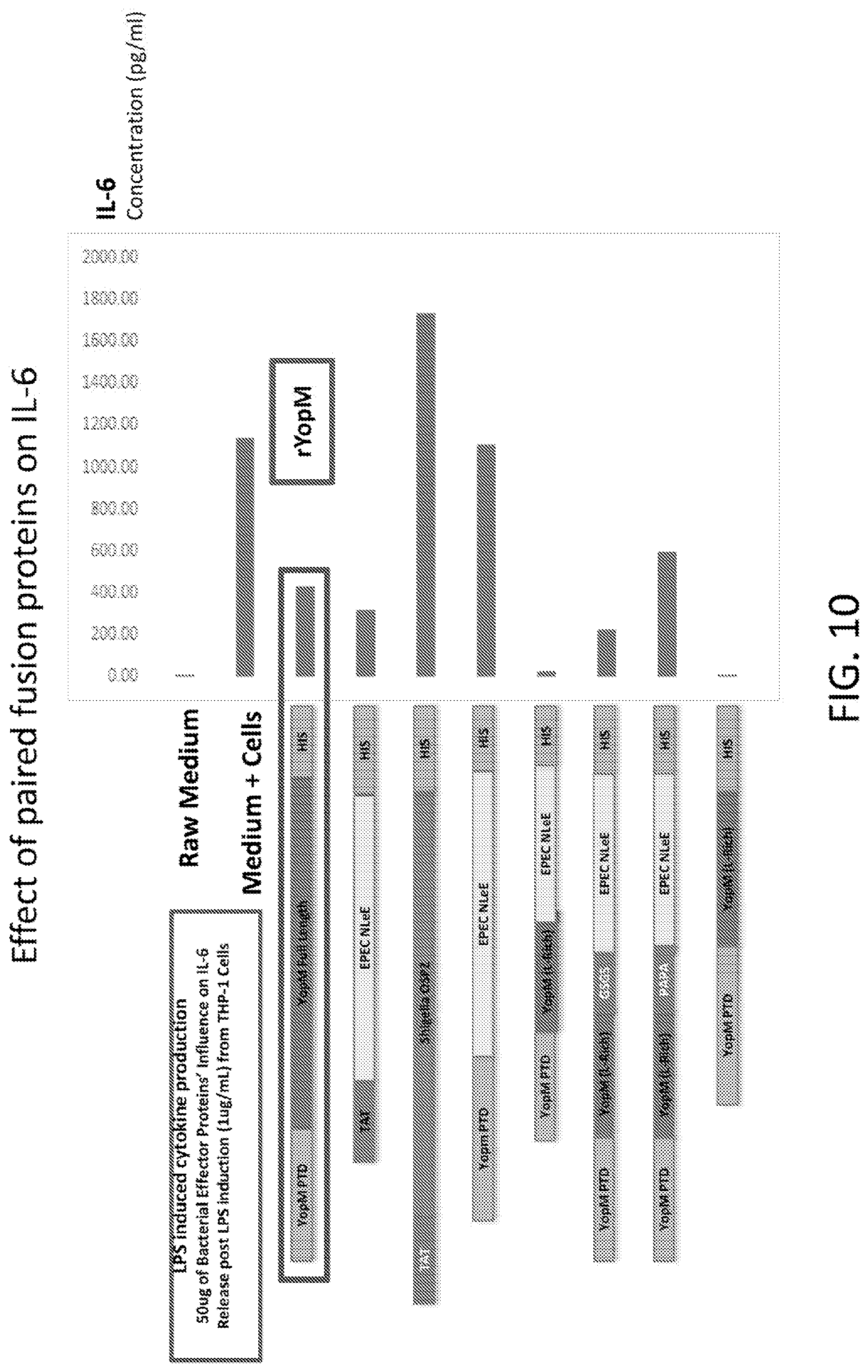
[0028] FIG. 10 is a graph showing the results of an analysis of the effect of paired fusion proteins on IL-6.
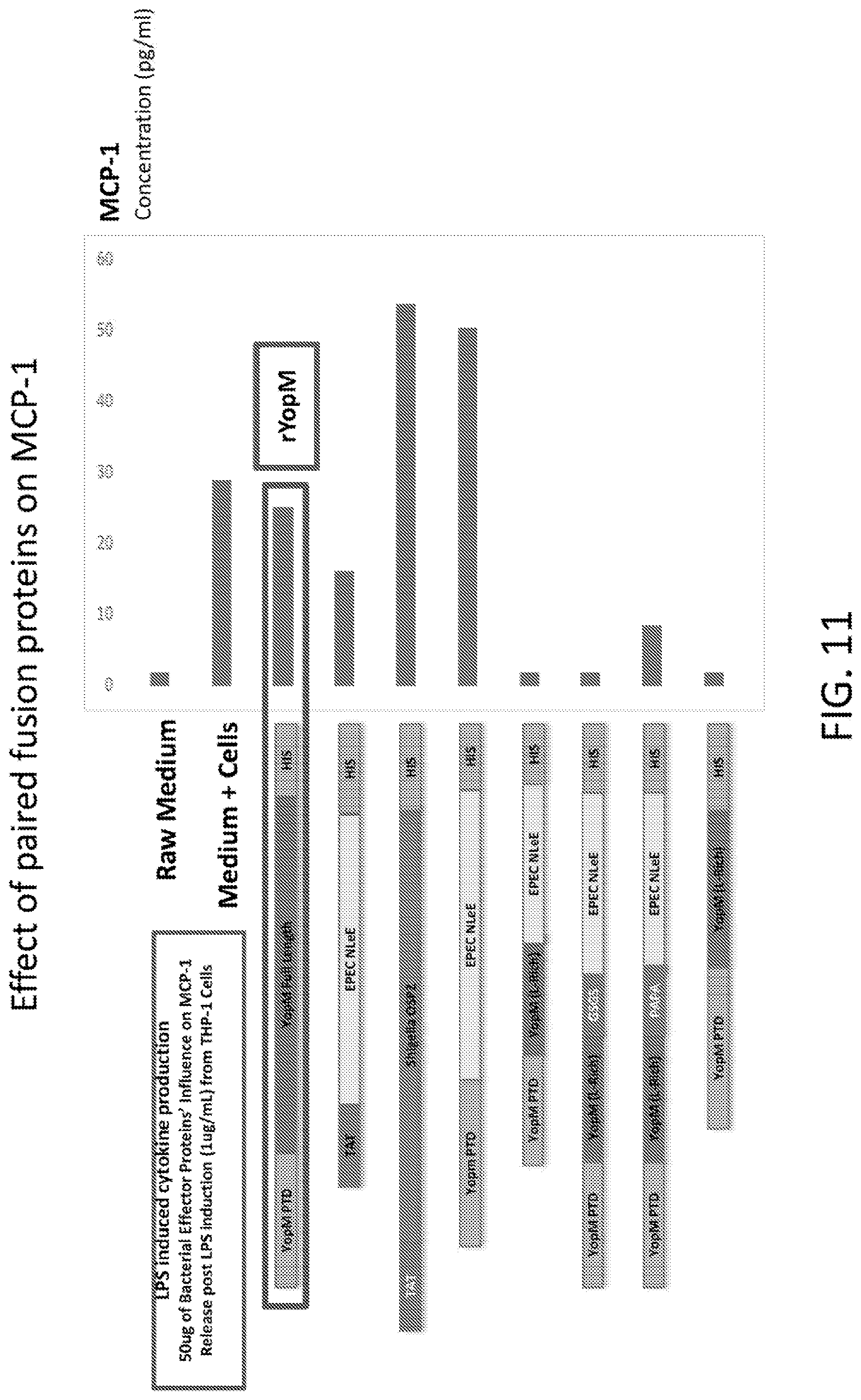
[0029] FIG. 11 is a graph showing the results of an analysis of the effect of paired fusion proteins on MCP-1.
[0030] FIG. 12 is a graph showing the results of an analysis of the effect of paired fusion proteins on IL-23.
[0031] FIG. 13 is 12 is a graph showing the results of a dose response analysis of rYopM and YopMo on caspase 1 activity.
[0032] FIG. 14 shows the uptake of FITC-TAT-NleE-WT-His protein by Hacat cells.
[0033] FIG. 15 shows the uptake of FITC-YopM PTD-YopM (L-Rich)-GSGS Linker NleE-WT-His protein by Hacat cells.
[0034] FIG. 16 shows the uptake of FITC-YopM PTD-YopM (L-Rich)-PAPA Linker-NleE-WT-His protein by Hacat cells
[0035] FIG. 17 shows the uptake of FITC-YopM PTD-YopM (L-Rich)-GSGS Linker-NleE-WT-His protein by Hacat cells.
[0036] FIG. 18 shows the uptake of TAT-NleE fusion polypeptide into intact mouse skin.
[0037] FIG. 19 shows is an two-photon microscopy image of uptake of TAT-NleE fusion polypeptide into intact mouse skin.
[0038] FIG. 20 shows a two-photon microscopy image of a 10 micron slice of mouse skin showing uptake of TAT-NleE fusion polypeptide into intact mouse skin.
[0039] FIG. 21 shows graphs illustrating an analysis of methylase activity of YopM PTD-YopM (L-Rich)-PAPA/GSGS-NleE-WT-His fusion proteins.
[0040] FIG. 22 shows graphs illustrating an analysis of methyltransferase activity NleE-R107A mutant and YopM.
[0041] FIG. 23 shows graphs illustrating a NleE and Shigella Methyltransferase activity assay.
[0042] FIG. 24 depicts the results of an experiment comparing the effect of formulations on NleE methylation activity.
[0043] FIG. 25 shows four micrograph panels showing transdermal penetration of CAGE-NleE protein.
[0044] FIG. 26 is a schematic showing how inflammation develops from cell to tissue in the skin.
[0045] FIG. 27 is a schematic showing the impact of a skin irritant on the NFKB pathways.
[0046] FIG. 28 shows the amino acid sequence of NleE SEQ ID NO: 1.
[0047] FIG. 29 shows the publicly available UniProt P17778 amino acid sequence of YopM SEQ ID NO: 2.
DETAILED DESCRIPTION OF THE PREFERRED EMBODIMENT
[0048] This description of preferred embodiments is intended to be read in connection with the accompanying drawings, which are to be considered part of the entire written description of this invention. The drawing figures are not necessarily to scale and certain features of the invention may be shown exaggerated in scale or in somewhat schematic form in the interest of clarity and conciseness. In the description, relative terms such as "horizontal," "vertical," "up," "down," "top" and "bottom" as well as derivatives thereof (e.g., "horizontally," "downwardly," "upwardly," etc.) should be construed to refer to the orientation as then described or as shown in the drawing figure under discussion. These relative terms are for convenience of description and normally are not intended to require a particular orientation. Terms including "inwardly" versus "outwardly," "longitudinal" versus "lateral" and the like are to be interpreted relative to one another or relative to an axis of elongation, or an axis or center of rotation, as appropriate. Terms concerning attachments, coupling and the like, such as "connected" and "interconnected," refer to a relationship wherein structures are secured or attached to one another either directly or indirectly through intervening structures, as well as both movable or rigid attachments or relationships, unless expressly described otherwise. The term "operatively connected" is such an attachment, coupling or connection that allows the pertinent structures to operate as intended by virtue of that relationship. When only a single machine is illustrated, the term "machine" shall also be taken to include any collection of machines that individually or jointly execute a set (or multiple sets) of instructions to perform any one or more of the methodologies discussed herein. In the claims, means-plus-function clauses, if used, are intended to cover the structures described, suggested, or rendered obvious by the written description or drawings for performing the recited function, including not only structural equivalents but also equivalent structures.
[0049] Technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs and by reference to published texts, which provide one skilled in the art with a general guide to many of the terms used in the present application. The definitions contained in this specification are provided for clarity in describing the components and compositions herein and are not intended to limit the claimed invention.
[0050] The present invention is based in part on the inventors' finding that combinations of bacterial effector polypeptides can have synergistic immunomodulatory activity. Many bacterial pathogens, including enteropathogenic Escherichia coli (EPEC), enterohemorrhagic E. coli, and Shigella, utilize a type III secretion system (T3SS) to deliver multiple virulence proteins directly into host cells. These virulence proteins, also referred to as effector proteins, are produced by bacteria during the infection of a eukaryotic host. The effector proteins down-regulate the host's immune system, typically at the site of the infection. Thus, effector proteins that target specific inflammatory pathways can function as immunomodulators, for example, in the treatment of inflammatory disorders. The inventors have found that combinations of effector proteins that target multiple inflammatory pathways produced augmented immunomodulatory effects.
[0051] Accordingly, the invention features compositions and methods that are useful for the treatment of inflammatory disorders. The compositions can include fusion proteins comprising bacterial effector polypeptides or a fragment of a bacterial effector polypeptide. More specifically, the compositions can include a set of paired peptides configured as a first bacterial effector polypeptide or a fragment thereof and a second bacterial effector polypeptides or a fragment thereof. For ease of reading, we will not repeat the phrase "or a fragment thereof" on every occasion. It is to be understood that where we refer to a first bacterial effector polypeptide, we refer to the first bacterial like and effector polypeptide or a fragment thereof. Similarly, it is to be understood that where we refer to a second bacterial effector polypeptide, we refer to the second bacterial effector polypeptide or a fragment thereof. Also featured are compositions comprising a set of paired peptides configured as a first bacterial effector polypeptide or a fragment thereof and a second bacterial effector polypeptides or a fragment thereof, and linked to a protein transduction domain. The methods can include a method of treating a subject at risk for or having an inflammatory disorder.
[0052] The compositions and methods described herein deliver a combinatorial construct of small recombinant protein effectors that directly target cellular pathways involved in inflammation, e.g., the NFKB pathway activated in skin inflammation. As described in detail below, the delivery of a recombinant construct comprising one or multiple effector proteins linked or fused to a PTD or CPP is useful to treat inflammation. Still other embodiments include the association of an optional targeting moiety, directing the construct to a specific cell or tissue type, with the effector protein and/or formulation in a topical carrier. While these compositions and methods of treatment have a number of advantages, a significant advantage is that the protein effectors used are not able to enter into the circulation. In one embodiment, as discussed in detail below, the compositions and methods involve the use of NleE incorporated into a cream or oil formulation that reduces skin inflammation and may be widely used for many applications, both as a cosmetic beauty creme to reduce redness and irritation and as a treatment of a disease or cause of skin irritation.
[0053] The present compositions and methods using multiple e.g., NFkB/MAPK inhibitors, for treatment of inflammatory responses (e.g., pain, redness, swelling, heat) is based upon the fact that inflammation caused by infection, injury, auto-immunity, sunburn, aging, etc is detected at the cell membrane. Signals that are received by the skin cell are funneled thru the NFKB (sometimes MAPK) pathway. This is a relay mechanism in the cell that must be highly regulated. Shutting off the NFKB pathway is the key to controlling skin inflammation. Naturally occurring bacterial effector proteins (e.g., NleE, YopM, SSPH1) are very potent inhibitors of the NFKB system. Their sole target and purpose is to dampen the inflammatory response.
[0054] As described herein, when these bacterial proteins are recombinantly engineered into a selected polypeptide and delivered to inflamed skin via fusion with a PTD/CPP and optional targeting moiety, they abolish the inflammatory response. The combination of two or more effector proteins, each with a different substrate in the cell, may be combinatorially fused together in a single polypeptide. In some embodiments, such combination achieves a synergistic effect, which is a significant improvement in activity beyond that accomplished by delivery of a single effector.
[0055] The bacterial effector, NleE, is characterized by specificity, potency and efficiency in shutting down NFkB and hence inflammatory reactions. The presence of NleE and/or a combination with other bacterial effectors in an anti-inflammatory formulation has a number of advantages. Among the advantages are extremely high substrate specificity, which results in only a very low, if any, chance of off-target effects and toxicity for therapeutic use. In one embodiment, fusing PTDs to NleE and its related effectors is useful to deliver the effector to sites of inflammation. We and others have shown that a number of recombinant effector proteins, when fused to PTDs can indeed cross the cell and tissue boundary and be taken up by cells resulting in NFkB/JNK/p38 pathway inhibition
[0056] The methods and compositions described below provide combinations of multiple effectors, or single or multiple effector(s) fused with a PTD, or single or multiple effector(s) fused with a targeting moiety, or single or multiple effector(s) fused with a PTD and a targeting moiety, in a chimeric recombinant protein, along with an emollient compound (such as CAGE) for use as a topical anti-inflammatory creme for many different ailments.
Compositions
[0057] Provided herein are compositions comprising engineered bacterial effector polypeptides for use in the treatment of inflammation. The engineered bacterial effector polypeptides can be configured as a set of paired peptides. More specifically, a set of paired peptides can be a construct comprising a first bacterial effector polypeptide or a fragment thereof and a second bacterial effector polypeptides or a fragment thereof. The set of paired peptides can be linked to one or more polypeptide sequences that facilitate intracellular delivery of the paired peptides, for example, a protein transduction domain (PTD) or a cell penetrating peptide (CPP).
[0058] Bacterial effector polypeptides. The first bacterial effector polypeptide and the second bacterial effector polypeptide can be a bacterial effector polypeptide selected from the exemplary bacterial effector polypeptides from a variety of bacteria as shown in Tables 1 and 2. Their enzymatic activity and host targets are also shown in Tables 1 and 2. Representative Uniprot or Genbank references for the polypeptides are shown in Tables 3 and 4. Additional amino acid sequences for, and nucleic acid sequences encoding, these bacterial effector polypeptides can be identified from databases such as UniProt, NCBI, GenBank and publications extant in the art.
TABLE-US-00001 TABLE 1 BACTERIAL T3SS EFFECTORS Effector Bacteria Intracellular Activity Host Target OspF Shigella Phosphothreonine lyase ERK, p38 flexneri MAPKs OspG Shigella Serine/threonine kinase E2 ubiquitin flexneri ligases NIeH1 EPEC-EHEC Serine/threonine kinase RPS3 NIeE/OspZ EPEC-EHEC/ Cysteine methylase TAB2/NfKB Shigella NIeB EPEC-EHEC O-GIcNAc transferase FADD, GAPDH, RIPK1, TRADD NIeC EPEC-EHEC Zinc metalloprotease NFkB YopH Yersinia Phosphotyrosine Akt/FAK phosphatase YopE Yersinia Rho GAP Rho GTPases/ caspases YopP/YopJ Yersinia Acetyltransferase MAP Ks YopM Yersinia LRR motif PKN/RSK
TABLE-US-00002 TABLE 2 BACTERIAL T3SS E3 UBIQUITIN LIGASE EFFECTORS E3 Ligase Factor Type/ Family Factor Bacteria Intracellular Activity Host Target HECT SopA Salmonella Regulation of host TRIM65/56 typhimurium inflammation NIeL EPEC/EHEC Formation of actin Unknown pedestal RING NIeG EPEC/EHEC Unknown Unknown U-Box LubX Legionella Regulation of another Cdh1, SidH pneumophila effector function Gob X L. pneumophila Unknown Unknown NEL IpaH1.4 Shigella Inhibition of NF-Kb HOIP flexneri activation IpaH2.5 S. flexneri Inhibition of NF-kB HOIP activation IaH4.5 S. flexneri Inhibition of NF-kB p65, TBK1 and I-IFN activation IpaH7.8 S. flexneri Induction of GLMN pyroptosis IpaH9.8 S. flexneri Inhibition of NF-kB NEMO activation IpaH0722 S. flexneri Inhibition of NF-kB TRAF2 activation SspH1 S. typhimurium Inhibition of androgen PKN1 receptor SspH2 S. typhimurium Promotion of IL-8 Nodi, SGT1 secretion SIrP S. typhimurium Induction of host cell Trx death SidE SidC L. pneumophila Unknown Unknown family (SdcA)
TABLE-US-00003 TABLE 3 Representative Amino Acid Sequences of Bacterial T355 Effectors UniProt or Genbank Effector Bacteria Host Target Reference ID OspF Shigella ERK, p38 Q8VSP9 flexneri MAPKs (OSPF_SHIFL) OspG Shigella E2 ubiquitin Q99PZ6 flexneri ligases (OSPG_SHIFL) NIeH1 EPEC- RPS3 Q8X831 EHEC (Q8X831_ECO57) NIeE/ EPEC- TAB2/NfKB Q7DBA6 OspZ EHEC/ (Q7DBA6_ECO57) Shigella NIeB EPEC- FADD, GAPDH, VEC94465.1 EHEC RIPK1, TRADD (Genbank) NIeC EPEC- NFkB CBG88408.1 EHEC (Genbank) YopH Yersinia Akt/FAK P15273 (YOPH_YEREN) YopE Yersinia Rho P31492 GTPases/caspases (YOPE_YEREN) YopP/YopJ Yersinia MAPKs O34336 (YOPP_BACSU) YopM Yersinia PKN/RSK P17778 (YOPM_YERPE)
TABLE-US-00004 TABLE 4 Representative Amino Acid Sequences of Bacterial T3ss E3 Ubiquitin Ligase Effectors E3 Ligase Factor UniProt or Genbank Type/Family Factor Bacteria Host Target Reference ID HECT SopA Salmonella TRIM65/56 Q8ZNR3 typhimurium (SOPA_SALTY) NIeL EPEC/EHEC Unknown A0A0D6ZN92 (A0A0D6ZN92_ECOLX) RING NIeG EPEC/EHEC Unknown A0A023YUN6 (A0A023YUN6_ECOLX) U-Box LubX Legionella Cdh1, Q5ZRQ0 pneumophila SidH (LUBX_LEGPH) Gob X L. pneumophila Unknown NEL IpaH1.4 Shigella HOIP A0A380D014 flexneri (A0A380D014_SHIFL) IpaH2.5 S. flexneri HOIP Q99Q42 (Q99Q42_SHIFL) IaH4.5 S. flexneri p65, TBK1 P18009 (IPA4_SHIFL) IpaH7.8 S. flexneri GLMN P18014 (IPA7_SHIFL) IpaH9.8 S. flexneri NEMO Q8VSC3 (IPA9_SHIFL) IpaH0722 S. flexneri TRAF2 SspH1 S. typhimurium PKN1 D0ZVG2 (SSPH1_SALT1) SspH2 S. typhimurium Nod1, P0CE12 SGT1 (SSPH2_SALTY) SIrP S. typhimurium Trx Q8ZQQ2 (SLRP_SALTY) SidE family SidC L. pneumophila Unknown Q6RCR3 (SdcA) (Q6RCR3_LEGPN)
[0059] Thus a bacterial effector peptide can be an SspH1; SspH2; SIrP; IpaH1.4; IpaH2.5; IpaH3; IpaH4.5; IpaH7.8; IpaH9.8; NleE; NleC; NleD; NleB; NleH; NleH1; YopM; YopE; YopH; YopJ; YopP; OspG; OspF; OspZ; OspI; SopE; SopB; SopE2; SipA; AvrA; SseL; EspT; or a TiR polypeptide.
[0060] In some embodiments, the bacterial effector polypeptide can have an amino acid sequence at least 90% identical to an amino acid sequence set forth in any of SEQ ID NOs. 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79. In some embodiments, the bacterial effector polypeptide have an amino acid sequence as set forth in any of SEQ ID NOs. 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79.
[0061] The term "type III secretion system or T355" refers to a highly specialized molecular needle construct containing a Yersiniae injectisome spanning the bacterial membranes, Yersinia outer protein (Yop) effectors and Yop translocators needed to deliver the effectors across the membrane (Camelis G R., Int J Med Microbial. 2002 February; 291(6-7):455-62). Pathogenic Yersiniae require this T3SS to survive and replicate extracellularly within lymphoid tissues of their animal or human hosts. See, also, U.S. Pat. No. 8,840,901.
[0062] The term "immodulatory effector protein" refers to small proteins, generally bacterial in origin, that suppress the human innate immune system during infection. These effector proteins activate the NFkB, JNK, and p38 signaling pathways during infection. These effector proteins are often secreted into the cells targeted for infection by T3SS. Once inside the cell, each effector protein targets a single host protein required for innate immunity, which it inactivates using a myriad of mechanisms including acetylation, methylation, action of phosphatases on P04 proteins, induced protein degradation etc.
[0063] All T3SS bacterial effector proteins are very small, globular, highly stable, highly catalytic, have high substrate specificity, bind co-factors very tightly, and can be injected into the cell in denatured form. They modify every substrate molecule in the cell. For instance, the EPEC effector NleE is a cysteine methyltransferase which has a single target in the cell, i.e., the TAB2 scaffold protein in the NFkB pathway. EPEC are mildly infective gut bacteria that attach to colon cells and directly inject virulence proteins thru a T3SS to control innate immune pathways as a survival strategy. During EPEC infection, the injected NleE protein methylates every molecule of cellular TAB2; thereby completely shutting off NFkB signaling, as described in Yao, Q. et al., Structure and Specificity of the Bacterial Cysteine Methyltransferase Effector NleE Suggests a Novel Substrate in Human DNA Repair Pathway., PLoS Pathogens (November 2014) 10(11):e1004522; doi:10.1371/journal.ppat.1004522.
[0064] Still another effector protein is YopM, discussed in U.S. Pat. No. 8,840,901 and Ruter, C & Hardwidge, PR, Drugs from Bugs': bacterial effector proteins as promising biological (immune-) therapeutics. FEMS Microbiol Lett 351 (December 2013/January 2014) 126-132.
[0065] In some embodiments, the first and second bacterial effector polypeptides are different. For example, they can have different amino acid sequences, different structures, different functions, different molecular targets, or have non-overlapping redundant roles in inhibiting an inflammatory pathway, for example, the NFkB, JNK, p38, and STING pathways.
[0066] Also included as effectors or effector proteins of the compositions and methods described herein are functional equivalents of the proteins described above. By the term "functional equivalent" is meant any amino acid sequence or modification thereof that has the same targeting and immune suppressing function of the naturally occurring effector protein. In one embodiment, such functional equivalents can have modifications of one or more amino acids from the known sequences. In one embodiment, such functional equivalents can be a smaller fragment of the known sequences. In one embodiment, such functional equivalents can be a derivative of the naturally occurring sequences or be derived from other than human sources. In one embodiment, such functional equivalents can be altered by chemical modification or be altered by recombinant production to be associated with sequences with which the effector proteins are not associated in nature. Similarly, chemical or structural changes or fragments of the nucleic acid sequences that encode the effector proteins are also considered functional equivalents herein.
[0067] The paired peptides can be joined by a linker. A linker can be any reagent, molecule or macromolecule that connects the first and second bacterial effector polypeptides such that the linker does not substantially alter the physiological activity of the effector polypeptides. A linker can be a peptide bond. That is, the first and second bacterial effector polypeptides or fragments thereof can be a fusion polypeptide comprising one or more amino acid segments from the first bacterial effector polypeptide and one or more amino acid segments from second bacterial effector polypeptide. The term "amino acid segment" as used herein refers to a contiguous stretch of amino acids within a polypeptide. For example, the amino acid residues 30 to 40 within a 100 amino acid polypeptide would be considered an amino acid segment. An amino acid segment can be a length greater than eight amino acid residues (e.g., greater than about nine, ten, 15, 20, 25, 30, 40, 50, 75, 100, 150, 200, 500, 1000, or more amino acid residues). In some embodiments, an amino acid segment can have a length less than 1000 amino acid residues (e.g., less than 500, less than 400, less than 350, less than 300, less than 200, or less than 100 amino acid residues). In other embodiments, an amino acid segment can have a length from about 20 to about 200 amino acid residues (e.g., about 30 to about 180 amino acid residues, or about 40 to about 150 amino acid residues).
[0068] The amino acid segments of the first bacterial effector polypeptide can be contiguous with the amino acid segments of the second or they can be separated by amino acids inserted as a structural spacer. A spacer segment can be one or more amino acids. The one or more amino acids can include amino acids that are the same or that are different. For example, a spacer can be a repeating series of a neutral amino acid (e.g., glycine, alanine, valine, isoleucine or leucine) ranging in number from 1 to 10 or more (e.g., 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more). Another example of a spacer configuration can be a series of interspersed amino acids that may be neutral (e.g., glycine-alanine-glycine-alanine-glycine-alanine, or glycine-glycine-glycine-valine-valine-valine) or charged amino acids (e.g., glutamate-glutamate-glutamate-arginine-arginine-arginine, or aspartate-lysine-aspartate-lysine-aspartate-lysine) or amino acids with other functional groups (e.g., proline-proline-proline-serine-serine-serine or tyrosine-glutamine-cysteine-methionine-tryptophan) ranging in number from 1 to 10 or more (e.g., 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more). In another embodiment, a spacer configuration can be a sequence of amino acids derived from a naturally occurring protein such as the hinge region joining the heavy chain CH1 and CH2 domains of immunoglobulin G. In some embodiments, the linker can be a GSGS linker (SEQ ID NO. ______) or a PAPA linker (SEQ ID NO. ______). In some embodiments, the fusion protein comprising a set of paired peptides can exclude a linker.
[0069] A fusion protein can be produced in vitro by continuous peptide synthesis according to standard chemical methods know to those in the art. Synthetic polypeptides can also be purchased from commercial sources. A fusion protein can also be produced by recombinant DNA techniques. Nucleic acid segments encoding the first bacterial effector polypeptide can be operably linked in the same open reading frame to nucleic acid sequences encoding the second bacterial effector polypeptide in a vector that includes the requisite regulatory elements, e.g., promoter sequences, transcription initiation sequences, and enhancer sequences, for expression in prokaryotic or eukaryotic cells.
[0070] The paired peptide constructs can include a combination of any of an SspH1; SspH2; SIrP; IpaH1.4; IpaH2.5; IpaH3; IpaH4.5; IpaH7.8; IpaH9.8; NleE; NleC; NleD; NleB; NleH; NleH1; YopM; YopE; YopH; YopJ; YopP; OspG; OspF; OspZ; OspI; SopE; SopB; SopE2; SipA; AvrA; SseL; EspT; or a TiR polypeptide. Thus, the the first bacterial effector polypeptide can be a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP, and the second bacterial effector polypeptide can be a polypeptide selected from the group consisting of NleE, NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP.
[0071] In some embodiments, the paired peptide construct can include a first bacterial effector polypeptide or fragment thereof is a YopM polypeptide or a fragment thereof or an NLeE polypeptide or a fragment thereof and a second bacterial effector polypeptide or fragment thereof is a YopM polypeptide or a fragment thereof or an NLeE polypeptide or a fragment thereof. In some embodiments, the first bacterial effector polypeptide or fragment thereof is a YopM polypeptide or a fragment thereof and the second bacterial effector polypeptide or fragment thereof is an NLeE polypeptide or a fragment thereof.
[0072] In some embodiments, the paired peptide constructs can be configured as summarized in Table 5 below.
TABLE-US-00005 TABLE 5 Paired Peptide Fusion Constructs Construct SEQ ID name Effector 1 Linker Effector 2 NO YopM PTD- YopM (L-rich) -- EPEC NLeE 80 NIeE (No linker) YopM PTD- YopM (L-rich) GSGS EPEC NLeE 82 NIeE (GSGS linker) YopM PTD- YopM (L-rich) PAPA EPEC NLeE 81 NIeE (PAPA linker)
[0073] In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 85% identical to the sequence set forth in SEQ ID NO. 80, 81, 82, 83, 84, 85, 86, 87, or 88. In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 90% identical to the sequence set forth in SEQ ID NO. 80, 81, 82, 83, 84, 85, 86, 87, or 88. In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 95% identical to the sequence set forth in SEQ ID NO. 80, 81, 82, 83, 84, 85, 86, 87, or 88. In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 99% identical to the sequence set forth in SEQ ID NO. 80, 81, 82, 83, 84, 85, 86, 87, or 88. In some embodiments, the paired peptide fusion protein can have an amino acid sequence as set forth in SEQ ID NOs. 80, 81, 82, 83, 84, 85, 86, 87, or 88.
[0074] Protein transduction domains. The set of paired peptides can be linked to one or more polypeptide sequences that facilitate intracellular delivery of the paired peptide. The terms "protein transduction domain (PTDs)" and/or "cell-penetrating peptide (CPPs)" refers to powerful sequences that allow intracellular delivery of conjugated cargoes to modify cell behavior. These small peptides can transport a wide variety of biologically active conjugates into the cell. Heterologous CPP coding sequences are added to effectors or effector-fusions to facilitate cellular uptake of the proteins into cells and tissues, including use of endogenous CPPs encoded in native effector proteins. This includes addition of CPP sequences or modules to the effector via chemical crosslinking, attachment to a nano-particle or other scaffold chemically or via PPIs for the purpose of transporting the effector across tissue and cell membranes. Among useful PTD or CPPs for the present methods and compositions are those known and identified in the art, including, without limitation, HIV Tat protein basic domain, (HIV Tat amino acids 48-60 or 49-57), poly-Arg or polyLys, penetratin, MPG, Pep-1, MAP, and transportan. See, e.g., Table 1 of Guidotti, G. et al, Trends in Pharmacological Sciences (April 2017), 38(4):406-424, which includes additional examples of CPPs and sequences origins and properties. Other CPPs are described in Norkowski, S. et al, Bacterial LPX motif-harboring virulence factors constitute a species-spanning family of cell-penetrating effectors, Cellular and Molecular Life Sciences (December 2017) doi.org/10.1007/s00018-017-2733-4, which described prototypes of such bacteria-derived cell-penetrating effectors (CPEs) including the Yersinia enterocolitica-derived YopM, the Salmonella typhimurium effector SspH1, and the Shigella IpaH proteins. Still other protein transporter molecules include those previously described in Dixon, J E et al, Proc. Natl Acad Sci, (January 2016), E291-299; as well as synthetic protein mimics described by Tezgel, A O et al BioMacromolecules (2017) 16:819-825. See, also, Bolhassani, A. et al, In vitro and in vivo delivery of therapeutic proteins using cell penetrating peptides. Peptides (November 2016), 87:50-63, which discusses useful CPPs for the present compositions and methods, including without limitation, covalent bonded CPPs, such as Poly-Arg peptides, Tat and VP22, df Tat, Cyclic CPPs, IMT-P8 (particularly useful for transdermal delivery), seven arginine (R7) and Streptolysin O (SLO)-mediated systems and elastin like polypeptide, CPP-adaptor system, 1, 2-Benzisothiazolin-3-one (BIT) and Tat, activatable cell-penetrating peptides, LDP12, M918, BR2, peptide for ocular delivery (POD), native protein independent of R11-CPP, Poly-arginine/Tat and Tat-PTD among others. Also identified are non-covalent bonded CPPs such as Pep-1, CADY-2, R8 and azo-R8, Penetratin, HR9 and IR9 peptides and pVEC. All of these documents are incorporated by reference herein for detailed descriptions of known CPPs and PTDs. It is also anticipated that novel PTD/CPPs will prove useful with the compositions described herein.
[0075] Exemplary protein transduction domains include a YopM protein transduction domain, an SspH1 protein transduction domain, or an IpaH protein transduction domain. A useful YopM protein transduction domain can have an amino acid sequence as set forth in SEQ ID NO 5.
[0076] In some embodiments, a fusion protein comprising a set of paired peptides linked to a protein transduction domain can have an amino acid sequence is at least 85% identical to the sequence set forth in SEQ ID NO. 10, 13, 16, 19, 22, or 24. In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 90% identical to the sequence set forth in SEQ ID NO. 10, 13, 16, 19, 22, or 24. In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 95% identical to the sequence set forth in SEQ ID NO. 10, 13, 16, 19, 22, or 24. In some embodiments, paired peptide fusion protein can have an amino acid sequence is at least 99% identical to the sequence set forth in SEQ ID NO. 10, 13, 16, 19, 22, or 24. In some embodiments, the paired peptide fusion protein can have an amino acid sequence as set forth in SEQ ID NOs. 10, 13, 16, 19, 22, or 24.
[0077] Polypeptides. We tend to use the term "protein" to refer to longer or larger amino acid polymers, and we tend to use the term "polypeptide" to refer to shorter sequences or to a chain of amino acid residues within a larger molecule (e.g., within a fusion protein) or complex. Both terms, however, are meant to describe an entity of two or more subunit amino acids, amino acid analogs, or other peptidomimetics, regardless of post-translational modification (e.g., amidation, phosphorylation or glycosylation). The subunits can be linked by peptide bonds or other bonds such as, for example, dicysteine, ester or ether bonds. The terms "amino acid" and "amino acid residue" refer to natural and/or unnatural or synthetic amino acids, which may be D- or L-form optical isomers. Full-length proteins, analogs, mutants, and fragments thereof are encompassed by this definition.
[0078] The amino acid sequence of the bacterial effector polypeptides disclosed herein can be identical to the wild-type sequences of appropriate components. Alternatively, any of the components can contain mutations such as deletions, additions, or substitutions. All that is required is that the variant bacterial effector polypeptide have at least 5% (e.g., 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, 99%, 100%, or even more) of the ability of the bacterial effector polypeptide containing only wild-type sequences to specifically bind the target. Substitutions will preferably be conservative substitutions. Conservative substitutions typically include substitutions within the following groups: glycine and alanine; valine, isoleucine, and leucine; aspartic acid and glutamic acid; asparagine, glutamine, serine and threonine; lysine, histidine and arginine; and phenylalanine and tyrosine.
[0079] Variant bacterial effector polypeptides, e.g., those having one or more amino acid substitutions relative to a native bacterial effector polypeptide amino acid sequence, can be prepared and modified as described herein. Amino acid substitutions can be made, in some cases, by selecting substitutions that do not differ significantly in their effect on maintaining (a) the structure of the peptide backbone in the area of the substitution, (b) the charge or hydrophobicity of the molecule at the target site, or (c) the bulk of the side chain. For example, naturally occurring residues can be divided into groups based on side-chain properties: (1) hydrophobic amino acids (norleucine, methionine, alanine, valine, leucine, and isoleucine); (2) neutral hydrophilic amino acids (cysteine, serine, and threonine); (3) acidic amino acids (aspartic acid and glutamic acid); (4) basic amino acids (asparagine, glutamine, histidine, lysine, and arginine); (5) amino acids that influence chain orientation (glycine and proline); and (6) aromatic amino acids (tryptophan, tyrosine, and phenylalanine) Substitutions made within these groups can be considered conservative substitutions. Non-limiting examples of useful substitutions include, without limitation, substitution of valine for alanine, lysine for arginine, glutamine for asparagine, glutamic acid for aspartic acid, serine for cysteine, asparagine for glutamine, aspartic acid for glutamic acid, proline for glycine, arginine for histidine, leucine for isoleucine, isoleucine for leucine, arginine for lysine, leucine for methionine, leucine for phenyalanine, glycine for proline, threonine for serine, serine for threonine, tyrosine for tryptophan, phenylalanine for tyrosine, and/or leucine for valine. Variant bacterial effector polypeptides having conservative and/or non-conservative substitutions (e.g., with respect to any of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, 79, 10, 13, 16, 19, 22, or 24), as well as fragments of any of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88, fragments of variants of any of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79. 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88 and polypeptides comprising any of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88, variants or fragments of any of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88, or fragments of variants of any of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88, can be screened for biological activity using suitable assays, including those described herein. For example, the activity of a bacterial effector polypeptide, for example, NLeE or a mutant or fragment thereof, can be evaluated in vitro by assaying for methylase activity or in cell based systems to characterize its effect on cytokine release.
[0080] In some embodiments, a bacterial effector polypeptide can comprise an amino acid sequence as set forth in SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, and 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88, but with a particular number of amino acid substitutions. For example, a bacterial effector polypeptides can have the amino acid sequence of any one of SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88, but with one, two, three, four, or five amino acid substitutions.
[0081] In some embodiments, a bacterial effector polypeptide as provided herein can include an amino acid sequence with at least 85% (e.g., 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 97.5%, 98%, 98.5%, 99.0%, 99.5%, 99.6%, 99.7%, 99.8%, 99.9%, or 100%) sequence identity to a region of a reference bacterial effector polypeptide sequence (e.g., SEQ ID NOS: 3, 89, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 78, 79, 10, 13, 16, 19, 22, or 24, 80, 81, 82, 83, 84, 85, 86, 87, or 88). Methods of determining percent sequence identity are discussed below.
[0082] In some embodiments, a polypeptide provided herein can be a substantially pure polypeptide. As used herein, the term "substantially pure" with reference to a polypeptide means that the polypeptide is substantially free of other polypeptides, lipids, carbohydrates, and nucleic acid with which it is naturally associated. Thus, a substantially pure polypeptide is any polypeptide that is removed from its natural environment and is at least 60 percent pure or is any chemically synthesized polypeptide. A substantially pure polypeptide can be at least about 60, 65, 70, 75, 80, 85, 90, 95, or 99 percent pure. Typically, a substantially pure polypeptide will yield a single major band on a non-reducing polyacrylamide gel.
[0083] A variety of methods can be used to to make a polypeptide including, for example, expression by prokaryotic systems, expression by eukaryotic systems, and chemical synthesis techniques. Exemplary methods for polypeptide purification purificinclude, without limitation, fractionation, centrifugation, and chromatography, e.g., gel filtration, ion exchange chromatography, reverse-phase HPLC and immunoaffinity purification.
[0084] A polypeptide can be modified by linkage to a polymer such as polyethylene glycol (PEG), or by fusion to another polypeptide such as albumin, for example. For example, one or more PEG moieties can be conjugated to a bacterial effector polypeptide or fusion protein via lysine residues. Linkage to PEG or another suitable polymer, or fusion to albumin or another suitable polypeptide can result in a modified bacterial effector polypeptide or fusion protein having an increased half life as compared to an unmodified bacterial effector polypeptide or fusion protein. Without being bound by a particular mechanism, an increased serum half life can result from reduced proteolytic degradation, immune recognition, or cell scavanging of the modified bacterial effector polypeptide or fusion protein. Methods for modifying a polypeptide by linkage to PEG (also referred to as "PEGylation") or other polymers include those set forth in U.S. Pat. No. 6,884,780; Cataliotti et al. ((2007) Trends Cardiovasc. Med. 17:10-14; Veronese and Mero (2008) BioDrugs 22:315-329; Miller et al. (2006) Bioconjugate Chem. 17:267-274; and Veronese and Pasut (2005) Drug Discov. Today 10:1451-1458, all of which are incorporated herein by reference in their entirety. Methods for modifying a polypeptide by fusion to albumin include those set forth in U.S. Patent Publication No. 20040086976, and Wang et al. (2004) Pharm. Res. 21:2105-2111, both of which are incorporated herein by reference in their entirety.
[0085] Nucleic acids. We may use the terms "nucleic acid" and "polynucleotide" interchangeably to refer to both RNA and DNA, including cDNA, genomic DNA, synthetic DNA, and DNA (or RNA) containing nucleic acid analogs, any of which may encode a polypeptide of the invention and all of which are encompassed by the invention. Polynucleotides can have essentially any three-dimensional structure. A nucleic acid can be double-stranded or single-stranded (i.e., a sense strand or an antisense strand). Non-limiting examples of polynucleotides include genes, gene fragments, exons, introns, messenger RNA (mRNA) and portions thereof, transfer RNA, ribosomal RNA, siRNA, micro-RNA, ribozymes, cDNA, recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes, and primers, as well as nucleic acid analogs. In the context of the present invention, nucleic acids can encode a bacterial effector polypeptide, paired peptide fusion protein, or construct comprising a paired peptide fusion protein linked to a protein transduction domain.
[0086] An "isolated" nucleic acid can be, for example, a naturally-occurring DNA molecule or a fragment thereof, provided that at least one of the nucleic acid sequences normally found immediately flanking that DNA molecule in a naturally-occurring genome is removed or absent. Thus, an isolated nucleic acid includes, without limitation, a DNA molecule that exists as a separate molecule, independent of other sequences (e.g., a chemically synthesized nucleic acid, or a cDNA or genomic DNA fragment produced by the polymerase chain reaction (PCR) or restriction endonuclease treatment). An isolated nucleic acid also refers to a DNA molecule that is incorporated into a vector, an autonomously replicating plasmid, a virus, or into the genomic DNA of a prokaryote or eukaryote. In addition, an isolated nucleic acid can include an engineered nucleic acid such as a DNA molecule that is part of a hybrid or fusion nucleic acid. A nucleic acid existing among many (e.g., dozens, or hundreds to millions) of other nucleic acids within, for example, cDNA libraries or genomic libraries, or gel slices containing a genomic DNA restriction digest, is not an isolated nucleic acid.
[0087] Isolated nucleic acid molecules can be produced by in several ways. For example, polymerase chain reaction (PCR) techniques can be used to obtain an isolated nucleic acid containing a nucleotide sequence described herein, including nucleotide sequences encoding a polypeptide described herein. PCR can be used to amplify specific sequences from DNA as well as RNA, including sequences from total genomic DNA or total cellular RNA. Generally, sequence information from the ends of the region of interest or beyond is employed to design oligonucleotide primers that are identical or similar in sequence to opposite strands of the template to be amplified. Various PCR strategies also are available by which site-specific nucleotide sequence modifications can be introduced into a template nucleic acid.
[0088] Isolated nucleic acids also can be chemically synthesized, either as a single nucleic acid molecule (e.g., using automated DNA synthesis in the 3' to 5' direction using phosphoramidite technology) or as a series of oligonucleotides. For example, one or more pairs of long oligonucleotides (e.g., >50-100 nucleotides) can be synthesized that contain the desired sequence, with each pair containing a short segment of complementarity (e.g., about 15 nucleotides) such that a duplex is formed when the oligonucleotide pair is annealed. DNA polymerase is used to extend the oligonucleotides, resulting in a single, double-stranded nucleic acid molecule per oligonucleotide pair, which then can be ligated into a vector.
[0089] Two nucleic acids or the polypeptides they encode may be described as having a certain degree of identity to one another. For example, a bacterial effector polypeptide and a biologically active variant thereof may be described as exhibiting a certain degree of identity. Alignments may be assembled by locating short sequences in the Protein Information Research (PIR) site (http://pir.georgetown.edu), followed by analysis with the "short nearly identical sequences" Basic Local Alignment Search Tool (BLAST) algorithm on the NCBI website (http://www.ncbi.nlm.nih.gov/blast).
[0090] As used herein, the term "percent sequence identity" refers to the degree of identity between any given query sequence and a subject sequence. For example, a bacterial effector polypeptide disclosed herein can be the query sequence and a fragment of a bacterial effector polypeptide can be the subject sequence. Similarly, a fragment of bacterial effector polypeptide can be the query sequence and a biologically active variant thereof can be the subject sequence.
[0091] To determine sequence identity, a query nucleic acid or amino acid sequence can be aligned to one or more subject nucleic acid or amino acid sequences, respectively, using the computer program ClustalW (version 1.83, default parameters), which allows alignments of nucleic acid or protein sequences to be carried out across their entire length (global alignment).
[0092] ClustalW calculates the best match between a query and one or more subject sequences and aligns them so that identities, similarities and differences can be determined. Gaps of one or more residues can be inserted into a query sequence, a subject sequence, or both, to maximize sequence alignments. For fast pair wise alignment of nucleic acid sequences, the following default parameters are used: word size: 2; window size: 4; scoring method: percentage; number of top diagonals: 4; and gap penalty: 5. For multiple alignments of nucleic acid sequences, the following parameters are used: gap opening penalty: 10.0; gap extension penalty: 5.0; and weight transitions: yes. For fast pair wise alignment of protein sequences, the following parameters are used: word size: 1; window size: 5; scoring method: percentage; number of top diagonals: 5; gap penalty: 3. For multiple alignment of protein sequences, the following parameters are used: weight matrix: blosum; gap opening penalty: 10.0; gap extension penalty: 0.05; hydrophilic gaps: on; hydrophilic residues: Gly, Pro, Ser, Asn, Asp, Gln, Glu, Arg, and Lys; residue-specific gap penalties: on. The output is a sequence alignment that reflects the relationship between sequences. ClustalW can be run, for example, at the Baylor College of Medicine Search Launcher site (searchlauncher.bcm.tmc.edu/multi-align/multi-align.html) and at the European Bioinformatics Institute site on the World Wide Web (ebi.ac.uk/clustalw).
[0093] To determine a percent identity between a query sequence and a subject sequence, ClustalW divides the number of identities in the best alignment by the number of residues compared (gap positions are excluded), and multiplies the result by 100. The output is the percent identity of the subject sequence with respect to the query sequence. It is noted that the percent identity value can be rounded to the nearest tenth. For example, 78.11, 78.12, 78.13, and 78.14 are rounded down to 78.1, while 78.15, 78.16, 78.17, 78.18, and 78.19 are rounded up to 78.2.
[0094] The nucleic acids and polypeptides described herein may be referred to as "exogenous". The term "exogenous" indicates that the nucleic acid or polypeptide is part of, or encoded by, a recombinant nucleic acid construct, or is not in its natural environment. For example, an exogenous nucleic acid can be a sequence from one species introduced into another species, i.e., a heterologous nucleic acid. Typically, such an exogenous nucleic acid is introduced into the other species via a recombinant nucleic acid construct. An exogenous nucleic acid can also be a sequence that is native to an organism and that has been reintroduced into cells of that organism. An exogenous nucleic acid that includes a native sequence can often be distinguished from the naturally occurring sequence by the presence of non-natural sequences linked to the exogenous nucleic acid, e.g., non-native regulatory sequences flanking a native sequence in a recombinant nucleic acid construct. In addition, stably transformed exogenous nucleic acids typically are integrated at positions other than the position where the native sequence is found.
[0095] Nucleic acids of the invention, that is, nucleic acids having a nucleotide sequence of any paired peptides fusion proteins and constructs disclosed herein, can include nucleic acids sequences that are at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 99% identical to the reference sequences disclosed herein.
[0096] A nucleic acid, i.e., an oligonucleotide (e.g., a probe or a primer) that is specific for a target nucleic acid will hybridize to the target nucleic acid under suitable conditions. We may refer to hybridization or hybridizing as the process by which an oligonucleotide single strand anneals with a complementary strand through base pairing under defined hybridization conditions. It is a specific, i.e., non-random, interaction between two complementary polynucleotides. Hybridization and the strength of hybridization (i.e., the strength of the association between the nucleic acids) is influenced by such factors as the degree of complementary between the nucleic acids, stringency of the conditions involved, and the melting temperature (Tm) of the formed hybrid. The hybridization products can be duplexes or triplexes formed with targets in solution or on solid supports.
[0097] Vectors. Vectors containing nucleic acids such as those described herein also are provided. A "vector" is a replicon, such as a plasmid, phage, or cosmid, into which another DNA segment may be inserted so as to bring about the replication of the inserted segment. Generally, a vector is capable of replication when associated with the proper control elements. Suitable vector backbones include, for example, those routinely used in the art such as plasmids, viruses, artificial chromosomes, BACs, YACs, or PACs. The term "vector" includes cloning and expression vectors, as well as viral vectors and integrating vectors. An "expression vector" is a vector that includes a regulatory region. A wide variety of host/expression vector combinations may be used to express the nucleic acid sequences described herein. Suitable expression vectors include, without limitation, plasmids and viral vectors derived from, for example, bacteriophage, baculoviruses, and retroviruses. Numerous vectors and expression systems are commercially available from such corporations as Novagen (Madison, Wis.), Clontech (Palo Alto, Calif.), Stratagene (La Jolla, Calif.), and Invitrogen/Life Technologies (Carlsbad, Calif.).
[0098] The vectors provided herein also can include, for example, origins of replication, scaffold attachment regions (SARs), and/or markers. A marker gene can confer a selectable phenotype on a host cell. For example, a marker can confer biocide resistance, such as resistance to an antibiotic (e.g., kanamycin, G418, bleomycin, or hygromycin). As noted above, an expression vector can include a tag sequence designed to facilitate manipulation or detection (e.g., purification or localization) of the expressed polypeptide. Tag sequences, such as green fluorescent protein (GFP), glutathione S-transferase (GST), polyhistidine, c-myc, hemagglutinin, or Flag.TM. tag sequences typically are expressed as a fusion with the encoded polypeptide. Such tags can be inserted anywhere within the polypeptide, including at either the carboxyl or amino terminus.
[0099] Additional expression vectors also can include, for example, segments of chromosomal, non-chromosomal and synthetic DNA sequences. Suitable vectors include derivatives of SV40 and known bacterial plasmids, e.g., E. coli plasmids col E1, pCR1, pBR322, pMal-C2, pET, pGEX, pMB9 and their derivatives, plasmids such as RP4; phage DNAs, e.g., the numerous derivatives of phage 1, e.g., NM989, and other phage DNA, e.g., M13 and filamentous single stranded phage DNA; yeast plasmids such as the 2p plasmid or derivatives thereof, vectors useful in eukaryotic cells, such as vectors useful in insect or mammalian cells; vectors derived from combinations of plasmids and phage DNAs, such as plasmids that have been modified to employ phage DNA or other expression control sequences.
[0100] Yeast expression systems can also be used. For example, the non-fusion pYES2 vector (XbaI, SphI, ShoI, NotI, GstXI, EcoRI, BstXI, BamH1, SacI, Kpn1, and HindIII cloning sites; Invitrogen) or the fusion pYESHisA, B, C (XbaI, SphI, ShoI, NotI, BstXI, EcoRI, BamH1, SacI, KpnI, and HindIII cloning sites.
[0101] The vector can also include a regulatory region. The term "regulatory region" refers to nucleotide sequences that influence transcription or translation initiation and rate, and stability and/or mobility of a transcription or translation product. Regulatory regions include, without limitation, promoter sequences, enhancer sequences, response elements, protein recognition sites, inducible elements, protein binding sequences, 5' and 3' untranslated regions (UTRs), transcriptional start sites, termination sequences, polyadenylation sequences, nuclear localization signals, and introns.
[0102] As used herein, the term "operably linked" refers to positioning of a regulatory region and a sequence to be transcribed in a nucleic acid so as to influence transcription or translation of such a sequence. For example, to bring a coding sequence under the control of a promoter, the translation initiation site of the translational reading frame of the polypeptide is typically positioned between one and about fifty nucleotides downstream of the promoter. A promoter can, however, be positioned as much as about 5,000 nucleotides upstream of the translation initiation site or about 2,000 nucleotides upstream of the transcription start site. A promoter typically comprises at least a core (basal) promoter. A promoter also may include at least one control element, such as an enhancer sequence, an upstream element or an upstream activation region (UAR). The choice of promoters to be included depends upon several factors, including, but not limited to, efficiency, selectability, inducibility, desired expression level, and cell- or tissue-preferential expression.
[0103] Vectors include, for example, viral vectors (such as adenoviruses ("Ad"), adeno-associated viruses (AAV), and vesicular stomatitis virus (VSV) and retroviruses), liposomes and other lipid-containing complexes, and other macromolecular complexes capable of mediating delivery of a polynucleotide to a host cell. Vectors can also comprise other components or functionalities that further modulate gene delivery and/or gene expression, or that otherwise provide beneficial properties to the targeted cells.
[0104] A "recombinant viral vector" refers to a viral vector comprising one or more heterologous gene products or sequences. Since many viral vectors exhibit size-constraints associated with packaging, the heterologous gene products or sequences are typically introduced by replacing one or more portions of the viral genome. Such viruses may become replication-defective, requiring the deleted function(s) to be provided in trans during viral replication and encapsidation (by using, e.g., a helper virus or a packaging cell line carrying gene products necessary for replication and/or encapsidation).
[0105] Suitable nucleic acid delivery systems include recombinant viral vector, typically sequence from at least one of an adenovirus, adenovirus-associated virus (AAV), helper-dependent adenovirus, retrovirus, or hemagglutinating virus of Japan-liposome (HVJ) complex. In such cases, the viral vector comprises a strong eukaryotic promoter operably linked to the polynucleotide e.g., a cytomegalovirus (CMV) promoter. The recombinant viral vector can include one or more of the polynucleotides therein, preferably about one polynucleotide. In some embodiments, the viral vector used in the invention methods has a pfu (plague forming units) of from about 10.sup.8 to about 5.times.10.sup.10 pfu. In embodiments in which the polynucleotide is to be administered with a non-viral vector, use of between from about 0.1 nanograms to about 4000 micrograms will often be useful e.g., about 1 nanogram to about 100 micrograms.
[0106] Additional vectors include retroviral vectors such as Moloney murine leukemia viruses and HIV-based viruses. One HIV-based viral vector comprises at least two vectors wherein the gag and pol genes are from an HIV genome and the env gene is from another virus. DNA viral vectors include pox vectors such as orthopox or avipox vectors, herpesvirus vectors such as a herpes simplex I virus (HSV) vector.
[0107] Pox viral vectors introduce the gene into the cells cytoplasm. Avipox virus vectors result in only a short term expression of the nucleic acid. Adenovirus vectors, adeno-associated virus vectors and herpes simplex virus (HSV) vectors may be an indication for some invention embodiments. The adenovirus vector results in a shorter term expression (e.g., less than about a month) than adeno-associated virus, in some embodiments, may exhibit much longer expression. The particular vector chosen will depend upon the target cell and the condition being treated. The selection of appropriate promoters can readily be accomplished. An example of a suitable promoter is the 763-base-pair cytomegalovirus (CMV) promoter. Other suitable promoters which may be used for gene expression include, but are not limited to, the Rous sarcoma virus (RSV), the SV40 early promoter region, the herpes thymidine kinase promoter, the regulatory sequences of the metallothionein (MMT) gene, prokaryotic expression vectors such as the .beta.-lactamase promoter, the tac promoter, promoter elements from yeast or other fungi such as the Gal 4 promoter, the ADC (alcohol dehydrogenase) promoter, PGK (phosphoglycerol kinase) promoter, alkaline phosphatase promoter; and the animal transcriptional control regions, which exhibit tissue specificity and have been utilized in transgenic animals: elastase I gene control region which is active in pancreatic acinar cells, insulin gene control region which is active in pancreatic beta cells, immunoglobulin gene control region which is active in lymphoid cells, mouse mammary tumor virus control region which is active in testicular, breast, lymphoid and mast cells, albumin gene control region which is active in liver, alpha-fetoprotein gene control region which is active in liver, alpha 1-antitrypsin gene control region which is active in the liver, beta-globin gene control region which is active in myeloid cells, myelin basic protein gene control region which is active in oligodendrocyte cells in the brain, myosin light chain-2 gene control region which is active in skeletal muscle, and gonadotropic releasing hormone gene control region which is active in the hypothalamus. Certain proteins can expressed using their native promoter. Other elements that can enhance expression can also be included such as an enhancer or a system that results in high levels of expression such as a tat gene and tar element. This cassette can then be inserted into a vector, e.g., a plasmid vector such as, pUC19, pUC118, pBR322, or other known plasmid vectors, that includes, for example, an E. coli origin of replication. The plasmid vector may also include a selectable marker such as the .beta.-lactamase gene for ampicillin resistance, provided that the marker polypeptide does not adversely affect the metabolism of the organism being treated.
[0108] Pharmaceutical carriers. The compositions also include a pharmaceutically acceptable carrier. We use the terms "pharmaceutically acceptable" (or "pharmacologically acceptable") to refer to molecular entities and compositions that do not produce an adverse, allergic or other untoward reaction when administered to an animal or a human, as appropriate. The term "pharmaceutically acceptable carrier," as used herein, includes any and all solvents, dispersion media, coatings, antibacterial, isotonic and absorption delaying agents, buffers, excipients, binders, lubricants, gels, surfactants and the like, that may be used as media for a pharmaceutically acceptable substance.
[0109] Thus, the invention also includes pharmaceutical compositions which contain, as the active ingredient, a fusion protein comprising a set of paired peptides or a fusion protein comprising a set of paired peptides linked a protein transduction domain, in combination with one or more pharmaceutically acceptable carriers. An active ingredient can be a composition comprising a set of paired peptides linked to a protein transduction domain and wherein prepared peptides comprise a first bacterial effector polypeptide linked to a second bacterial effector polypeptide. In some embodiments, the polypeptide compositions can be sterilized using conventional sterilization techniques before or after it is combined with the pharmaceutically acceptable carrier. In making the compositions of the invention, the polypeptide compositions are typically mixed with an excipient, diluted by an excipient or enclosed within such a carrier in the form of, for example, a capsule, tablet, sachet, paper, or other container. When the excipient serves as a diluent, it can be a solid, semisolid, or liquid material (e.g., normal saline), which acts as a vehicle, carrier or medium for the active ingredient. Thus, the compositions can be in the form of tablets, pills, powders, lozenges, sachets, cachets, elixirs, suspensions, emulsions, solutions, syrups, aerosols (as a solid or in a liquid medium), ointments, soft and hard gelatin capsules, suppositories, sterile injectable solutions, and sterile packaged powders. As is known in the art, the type of diluent can vary depending upon the intended route of administration. The resulting compositions can include additional agents, such as preservatives. The excipient or carrier is selected on the basis of the mode and route of administration. Suitable pharmaceutical carriers, as well as pharmaceutical necessities for use in pharmaceutical formulations, are described in Remington's Pharmaceutical Sciences (E. W. Martin), a well-known reference text in this field, and in the USP/NF (United States Pharmacopeia and the National Formulary). Some examples of suitable excipients include lactose, dextrose, sucrose, sorbitol, mannitol, starches, gum acacia, calcium phosphate, alginates, tragacanth, gelatin, calcium silicate, microcrystalline cellulose, polyvinylpyrrolidone, cellulose, water, syrup, and methyl cellulose. The formulations can additionally include: lubricating agents such as talc, magnesium stearate, and mineral oil; wetting agents; emulsifying and suspending agents; preserving agents such as methyl- and propylhydroxy-benzoates; sweetening agents; and flavoring agents. The pharmaceutical compositions can also be formulated so as to provide quick, sustained or delayed release of the active ingredient after administration to the patient.
[0110] Pharmaceutically acceptable compositions for use in the present methods, including those in which the polypeptides are entrapped in a colloid for oral delivery, can be prepared according to standard techniques. The polypeptides can be dried and compacted by grinding or pulverizing and inserted into a capsule for oral administration. In some embodiments, the polypeptides can be combined one or more excipients, for example, a disintegrant, a filler, a glidant, or a preservative. Suitable capsules include both hard shell capsules or soft-shelled capsules. Any lipid-based or polymer-based colloid may be used to form the capusule. Exemplary polymers useful for colloid preparations include gelatin, plant polysaccharides or their derivatives such as carrageenans and modified forms of starch and cellulose, e.g., hypromellose. Optionally, other ingredients may be added to the gelling agent solution, for example plasticizers such as glycerin and/or sorbitol to decrease the capsule's hardness, coloring agents, preservatives, disintegrants, lubricants and surface treatment. In some embodiments, the capsule does not include gelatin. In other embodiments, the capsule does not include plant polysaccharides or their derivatives.
[0111] Regardless of their original source or the manner in which they are obtained, the polypeptides of the invention can be formulated in accordance with their use. These compositions can be prepared in a manner well known in the pharmaceutical art, and can be administered by a variety of routes, depending upon whether local or systemic treatment is desired and upon the area to be treated. Administration may be oral or topical (including ophthalmic and to mucous membranes including intranasal, vaginal and rectal delivery). In some embodiments, administration can be pulmonary (e.g., by inhalation or insufflation of powders or aerosols, including by nebulizer; intratracheal, intranasal, epidermal and transdermal) or ocular. Methods for ocular delivery can include topical administration (eye drops), subconjunctival, periocular or intravitreal injection or introduction by balloon catheter or ophthalmic inserts surgically placed in the conjunctival sac. Parenteral administration includes intravenous, intraarterial, subcutaneous, intraperitoneal or intramuscular injection or infusion; or intracranial, e.g., intrathecal or intraventricular administration. Parenteral administration can be in the form of a single bolus dose, or may be, for example, by a continuous perfusion pump. Pharmaceutical compositions and formulations for topical administration may include transdermal patches, ointments, lotions, creams, gels, drops, suppositories, sprays, liquids, powders, and the like. Conventional pharmaceutical carriers, aqueous, powder or oily bases, thickeners and the like may be necessary or desirable.
[0112] The compositions can be formulated in a unit dosage form, each dosage containing, for example, from about 0.005 mg to about 2000 mg of polypeptides per daily dose. The term "unit dosage forms" refers to physically discrete units suitable as unitary dosages for human subjects and other mammals, each unit containing a predetermined quantity of active material calculated to produce the desired therapeutic effect, in association with a suitable pharmaceutical excipient. For preparing solid compositions such as tablets, the principal active ingredient is mixed with a pharmaceutical excipient to form a solid preformulation composition containing a homogeneous mixture of a compound of the present invention. When referring to these preformulation compositions as homogeneous, the active ingredient is typically dispersed evenly throughout the composition so that the composition can be readily subdivided into equally effective unit dosage forms such as tablets, pills and capsules. This solid preformulation is then subdivided into unit dosage forms of the type described above containing from, for example, 0.005 mg to about 1000 mg of the compositions of the present invention.
[0113] The compositions can be formulated in a unit dosage form, each dosage containing, for example, from about 0.1 mg to about 50 mg, from about 0.1 mg to about 40 mg, from about 0.1 mg to about 20 mg, from about 0.1 mg to about 10 mg, from about 0.2 mg to about 20 mg, from about 0.3 mg to about 15 mg, from about 0.4 mg to about 10 mg, from about 0.5 mg to about 1 mg; from about 0.5 mg to about 100 mg, from about 0.5 mg to about 50 mg, from about 0.5 mg to about 30 mg, from about 0.5 mg to about 20 mg, from about 0.5 mg to about 10 mg, from about 0.5 mg to about 5 mg; from about 1 mg from to about 50 mg, from about 1 mg to about 30 mg, from about 1 mg to about 20 mg, from about 1 mg to about 10 mg, from about 1 mg to about 5 mg; from about 5 mg to about 50 mg, from about 5 mg to about 20 mg, from about 5 mg to about 10 mg; from about 10 mg to about 100 mg, from about 20 mg to about 200 mg, from about 30 mg to about 150 mg, from about 40 mg to about 100 mg, from about 50 mg to about 100 mg of the active ingredient.
[0114] In some embodiments, tablets or pills of the present invention can be coated or otherwise compounded to provide a dosage form affording the advantage of prolonged action. For example, the tablet or pill can comprise an inner dosage and an outer dosage component, the latter being in the form of an envelope over the former. The two components can be separated by an enteric layer which serves to resist disintegration in the stomach and permit the inner component to pass intact into the duodenum or to be delayed in release. A variety of materials can be used for such enteric layers or coatings, such materials including a number of polymeric acids and mixtures of polymeric acids with such materials as shellac, cetyl alcohol, and cellulose acetate.
[0115] The liquid forms in which the compositions of the present invention can be incorporated for administration orally or by injection include aqueous solutions, suitably flavored syrups, aqueous or oil suspensions, and flavored emulsions with edible oils such as cottonseed oil, sesame oil, coconut oil, or peanut oil, as well as elixirs and similar pharmaceutical vehicles.
[0116] The proportion or concentration of the compositions of the invention in a pharmaceutical composition can vary depending upon a number of factors including dosage, chemical characteristics (e.g., hydrophobicity), and the route of administration.
Methods of Treatment
[0117] The compositions disclosed herein are generally and variously useful for treatment of inflammatory disorders and conditions. Inflammatory disorders and conditions encompass a wide range of disorders covering many different systems and organs, including the gastrointestinal tract, the skin, the lungs, and the musculoskeletal system. Exemplary inflammatory disorders include inflammatory bowel disease, rheumatoid arthritis, allergy, asthma, autoimmune diseases, coeliac disease, glomerulonephritis, hepatitis, preperfusion injury and transplant rejection. An inflammatory disorder can be a gastrointestinal disorder, for example, inflammatory bowel disease, Crohn's disease, and the ileocolitis, ileocecal, jeunoileitis, and gastroduodenal subtypes of-Crohn's disease, and ulcerative colitis and subtypes of ulcerative colitis.
[0118] A subject is effectively treated whenever a clinically beneficial result ensues. This may mean, for example, a complete resolution of the symptoms associated with an inflammatory disorder, a decrease in the severity of the symptoms associated with an inflammatory disorder, or a slowing of the progression of symptoms associated with an inflammatory disorder. These methods can further include the steps of a) identifying a subject (e.g., a patient and, more specifically, a human patient) who has an inflammatory disorder; and b) providing to the subject a composition comprising a paired peptide composition disclosed herein in a physiologically acceptable carrier. An amount of such a composition provided to the subject that results in a complete resolution of the symptoms associated with an inflammatory disorder, a decrease in the severity of the symptoms associated with an inflammatory disorder, or a slowing of the progression of symptoms associated with an inflammatory disorder considered a therapeutically effective amount. The present methods may also include a monitoring step to help optimize dosing and scheduling as well as predict outcome.
[0119] The methods disclosed herein can be applied to a wide range of species, e.g., humans, non-human primates (e.g., monkeys), horses, pigs, cows or other livestock, dogs, cats or other mammals kept as pets, rats, mice, or other laboratory animals. The compositions described herein are useful in therapeutic compositions and regimens or for the manufacture of a medicament for use in treatment of conditions as described herein (e.g., inflammatory disorders and conditions.)
[0120] When formulated as pharmaceuticals, the compositions can be administered to any part of the host's body for subsequent delivery to a target cell. A composition can be delivered to, without limitation, the brain, the cerebrospinal fluid, joints, nasal mucosa, blood, lungs, intestines, muscle tissues, skin, or the peritoneal cavity of a mammal. In terms of routes of delivery, a composition can be administered by intravenous, intracranial, intraperitoneal, intramuscular, subcutaneous, intramuscular, intrarectal, intravaginal, intrathecal, intratracheal, intradermal, or transdermal injection, by oral or nasal administration, or by gradual perfusion over time. In a further example, an aerosol preparation of a composition can be given to a host by inhalation.
[0121] Regardless of how the compositions are formulated, the dosage required will depend on the route of administration, the nature of the formulation, the nature of the subject's condition, e.g., a gastrointestinal disorder or a skin disorder, the subject's size, weight, surface area, age, and sex, other drugs being administered, and the judgment of the attending clinicians. Suitable dosages are in the range of 0.01-1,000 mg/kg. Some typical dose ranges are from about 1 .mu.g/kg to about 1 g/kg of body weight per day. In some embodiments, the dose range is from about 0.01 mg/kg to about 100 mg/kg of body weight per day. In some embodiments, the dose can be, for example, 1 mg/kg, 2 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg, 50 mg/kg or 100 mg/kg. The dosage is likely to depend on such variables as the type and extent of progression of the disease or disorder, the overall health status of the particular patient, the relative biological efficacy of the compound selected, formulation of the excipient, and its route of administration.
[0122] Effective doses can be extrapolated from dose-response curves derived from in vitro or animal model test systems. For example, in vitro analysis of cytokine production by peripheral blood mononuclear cells (PBMCs) can be a useful for assaying pro- and anti-inflammatory responses, e.g., secretion of IL-1beta, IL-12, IL-4 or IL-10, IL-6, IL-23, and TNF-alpha. Compositions can also be analyzed for effects in animal models, for example, IgA production, cytokine production by explants of Peyer's patches, and dendritic cell and T-cell responses.
[0123] Wide variations in the needed dosage are to be expected in view of the variety of cellular targets and the differing efficiencies of various routes of administration. Variations in these dosage levels can be adjusted using standard empirical routines for optimization, as is well understood in the art. Administrations can be single or multiple (e.g., 2- or 3-, 4-, 6-, 8-, 10-, 20-, 50-, 100-, 150-, or more fold). Encapsulation of the compounds in a suitable delivery vehicle (e.g., polymeric microparticles or implantable devices) may increase the efficiency of delivery.
[0124] The duration of treatment with any composition provided herein can be any length of time from as short as one day to as long as the life span of the host (e.g., many years). For example, a composition can be administered once a week (for, for example, 4 weeks to many months or years); once a month (for example, three to twelve months or for many years); or once a year for a period of 5 years, ten years, or longer. It is also noted that the frequency of treatment can be variable. For example, the present compositions can be administered once (or twice, three times, etc.) daily, weekly, monthly, or yearly.
[0125] Any method known to those in the art can be used to determine if a particular response is induced. Clinical methods that can assess the degree of a particular disease state can be used to determine if a response is induced. For example, a subject can be monitored for symptomatic relief, e.g., relief from colic, diarrhea, constipation, nausea, vomiting, abdominal pain, cramping, heartburn, abdominal distention, flatulence, or incontinence, dermatitis, redness, pain, swelling. Alternatively or in addition, serum markers, imaging techniques, e.g., ultrasound, x-rays, and endoscopic methods can be used.
[0126] The compositions may also be administered in conjunction with other therapeutic modalities. Other therapeutic modalities will vary according to the particular disorder, but can include, for example, anti-inflammatory agents, antibiotics and other dietary treatments, anti-diarrhea medications, anti-emetics, anti-cholinergic agents, Concurrent administration of two or more therapeutic agents does not require that the agents be administered at the same time or by the same route, as long as there is an overlap in the time period during which the agents are exerting their therapeutic effect. Simultaneous or sequential administration is contemplated, as is administration on different days or weeks.
Articles of Manufacture
[0127] The invention also features kits for administering the compositions. Accordingly, packaged products (e.g., sterile containers containing one or more of the compositions described herein and packaged for storage, shipment, or sale at concentrated or ready-to-use concentrations) and kits, are also within the scope of the invention. A product can include a container (e.g., a vial, jar, bottle, bag, microplate or beads) containing one or more compositions of the invention. In addition, an article of manufacture further may include, for example, packaging materials, instructions for use, syringes, delivery devices, buffers or other control reagents.
[0128] For example, the kit can include a measured amount of a composition comprising a set of paired peptides wherein the set of paired peptides comprises a first bacterial effect or polypeptide went to a second bacterial effector polypeptide and a protein transduction domain. The compounds, agents, and/or reagents can be packaged in a suitable container. The kit can further comprise instructions for administering the compositions. For example, the kit can include: a fusion protein comprising a protein transduction domain and a set of paired peptides. The kit can also include a buffering agent, a preservative, and/or a protein stabilizing agent. Each component of the kit can be enclosed within an individual container and all of the various containers can be within a single package. The product may also include a legend (e.g., a printed label or insert or other medium describing the product's use (e.g., an audio- or videotape or computer readable medium)). The legend can be associated with the container (e.g., affixed to the container) and can describe the manner in which the reagents can be used. The reagents can be ready for use (e.g., present in appropriate units), and may include one or more additional adjuvants, carriers or other diluents.
[0129] Also included as effectors or effector proteins of the compositions and methods described herein are functional equivalents of the proteins described above. By the term "functional equivalent" is meant any amino acid sequence or modification thereof that has the same targeting and immune suppressing function of the naturally occurring effector protein. In one embodiment, such functional equivalents can have modifications of one or more amino acids from the known sequences. In one embodiment, such functional equivalents can be a smaller fragment of the known sequences. In one embodiment, such functional equivalents can be a derivative of the naturally occurring sequences or be derived from other than human sources. In one embodiment, such functional equivalents can be altered by chemical modification or be altered by recombinant production to be associated with sequences with which the effector proteins are not associated in nature. Similarly, chemical or structural changes or fragments of the nucleic acid sequences that encode the effector proteins are also considered functional equivalents herein.
[0130] As used herein, the term "construct" as described herein refers to a chemically synthesized or genetically engineered assemblage that comprises one or more PTD/CPP associated with one or more effector proteins and further optionally associated with one or more targeting moieties. The construct can be in the form of a polypeptide or a nucleic acid molecule encoding the polypeptide.
[0131] As used herein the term(s) "cosmeceutically or pharmaceutically acceptable carrier, excipient or formulation" refer to the components of a composition that provide a vehicle for delivery. For example, where the cosmeceutical or pharmaceutical product is a topical composition, the carrier or formulation can contain typical components such as cremes, saline, vitamins, oils that are normally found in cosmetic or pharmaceutical preparations for skin. See, for example, U.S. Pat. No. 5,635,497 which discloses an oil-in-water, fatty cream composition for topical administration comprising from 60 to 80 percent by weight of fatty components, from 1.5 to 5 percent by weight of at least one non-ionic, hydrophilic surfactant having an HLB of at least 14, about 6% of fatty alcohols and esters, a therapeutically effective amount of at least one topically active therapeutic agent, and water, provided the topically active therapeutic agent is not dithranol or its derivatives. See, also International Patent Publication No. WO2014/076642. Pharmaceutically or cosmetically acceptable excipients suitable for the compositions described herein can be selected from plasticizers, disintegrants, glidants, coloring agents, lubricants, stabilizers, adsorbents, preservatives, delivery retarders and mixtures thereof. Such a composition may contain a transepidermal or transdermal carrier agent consisting of acidic electrolyzed water having a pH of 1.0 to 4.0 and comprise clusters of water having 5 to 10 molecules of water per cluster, and a polyacrylate. Other suitable formulations may include oils, emollients, lotions for topical and transdermal applications along with buffered/aqueous and saline solutions. See, also, texts such as Topical Drug Delivery Formulation (eds. Osborne and Amann), 2000, publishers Taylor & Francis, Drugs and the Pharmaceutical Sciences Series #42.
[0132] The term "CAGE" as used herein refers to deep eutectic synthetic solvent, a choline-based oil that has antimicrobial activity, which has been shown to penetrate deep into the dermis. CAGE is described by Zakrewsky, M. et al, Adv. Healthcare Mater. (March 2016), 5, 1282-1289, incorporated by reference herein. CAGE has been shown to be useful in transdermal protein delivery, wherein the protein is carried 15-20 cell layers into model skin preparations. See, e.g., Banerjee, A. et al, Adv. Healthcare Mater. 2017, 1601411 DOI: 10.1002/adhm.201601411, incorporated by reference herein.
[0133] Still other pharmaceutical strategies to enhance dermal delivery of peptides or proteins including carrier peptides, signaling peptides neurotransmitter-inhibiting peptides and enzyme-inhibiting peptides, include chemical and physical penetration enhancers, such as listed in Table 2 of Badenhorst, T. et al, Pharmaceutical Strategies for the Topical Dermal Delivery of Peptides/Proteins for Cosmetic and Therapeutic Applications. Austin Journal of Pharmacology and Therapeutics (2014), 2(6):10. Also as discussed in this document are coupling with lipophilic moiety, such as lauric, palmitic and other acids, using CPP conjugates, formulation with microemulsions, encapsulation in liposomal vesicles and use of lipid particles, as well as combinations of these formulations.
[0134] The term "targeting moiety" refers to constructs useful in fusion with the effector proteins and/or PTD/CPPs described herein to direct the fusion protein or a nucleic acid sequence encoding it to a specific cell or tissue type within the body. Alterations to the fusion constructs described by the addition of amino acid segments to naturally occurring effector or fusion effector sequence, which enables the protein to bind and to specifically target cells, tissues or other target or physiologic compartment in the human body create "targeted-effectors" and "targeted effector fusions". Such targeting moieties include amino acid segments that enhance the efficacy of the preparation by its ability to be activated due to conditions in a specific compartment of the body could be cleaved, for instance by proteolytic cleavage, addition of post-translational modifications, or forming of PPIs with cell and tissue specific host proteins. Examples of tissue-specific targeting peptides for this use include those described in Jung, E. et al., Identification of tissue-specific targeting peptide. J Comput Aided Mol Des (October 2012) 26:1267-1275, incorporated by reference herein. Targeting moieties can also be antibodies, antibody fragments, aptamers, amino acid sequences, nucleic acid sequences that are complementary to or capable of binding a complementary sequence on a cell or tissue or chemical moieties that have a three-dimensional structure that can fit into a three-dimensional pocket on the targeted cell or tissue. For example, a targeting sequence can be a hormone, or fragment thereof, that targets or binds its naturally occurring cell surface receptor, or tissue specific markers, etc.
[0135] The term "polypeptide," when used in singular or plural form, generally refers to a polymer of amino acids joined together by peptide bonds and may include unmodified or naturally occurring amino acids or modified or unnatural amino acids. In certain embodiment, the term polypeptide refers to a construct formed by multiple shorter peptides joined directly, or indirectly via linkers, to form a single peptide. In one embodiment, as described herein, a polypeptide is formed by the fusion of a PTD or CPP and an effector protein. In another embodiment, as described herein, a polypeptide is formed by the fusion of a PTD attached to an effector protein, wherein the effector protein is further attached to another effector protein (with or without its own PTD). In still another embodiment, the polypeptide of this invention is formed by the covalent association of a first PTD fused to a first effector protein, a linker followed by an additional (e.g., second, third, fourth, etc) effector protein. The term "first" is used only to distinguish among the effectors. In yet another embodiment, each effector protein of the polypeptide is associated with its own PTD or CPP. In yet another embodiment, only one PTD/CPP is present. In another embodiment, the single PTD/CPP is located at the N terminus of the polypeptide. In yet another embodiment, multiple PTD/CPP are present in the polypeptide, each located at the N terminus of its associated effector protein. In another embodiment, each PTD/CPP is separated from its effector protein by a linker. In another embodiment, each effector protein is separated from each additional effector protein by a linker. The first and additional effector constructs in a single polypeptide may occur in any order.
[0136] In still another embodiment of a polypeptide as described herein, the PTD/CPP are fused directly to the effector protein and each effector protein is fused to each additional effector protein. In certain embodiments, the polypeptides contain two or more different effector proteins, each having its own target. In certain embodiments, the polypeptides contain two or more different effector proteins, each targeting related targets. In still further embodiments, each polypeptide is further associated with a targeting moiety to target the polypeptide to a specific tissue or cell type, e.g., skin, epidermis, dermis.
[0137] By "homologous protein" is meant a protein having a percent sequence similarity or identity of greater than 80%, greater than 85%, greater than 90%, greater than 95%, greater than 97%, or greater than 99% and sharing the same function as the effector protein.
[0138] As used herein, the term "polynucleotide," when used in singular or plural form, generally refers to any polyribonucleotide or polydeoxribonucleotide, which may be unmodified RNA or DNA or modified RNA or DNA that encode any of the polypeptide constructs as described above. Thus, for instance, polynucleotides as defined herein include, without limitation, single- and double-stranded DNA, DNA including single- and double-stranded regions, single- and double-stranded RNA, and RNA including single- and double-stranded regions, hybrid molecules comprising DNA and RNA that may be single-stranded or, more typically, double-stranded or include single- and double-stranded regions. In addition, the term "polynucleotide" as used herein refers to triple-stranded regions comprising RNA or DNA or both RNA and DNA. The term "polynucleotide" specifically includes cDNAs. The term includes DNAs (including cDNAs) and RNAs that contain one or more modified bases. In general, the term "polynucleotide" embraces all chemically, enzymatically and/or metabolically modified forms of unmodified polynucleotides, as well as the chemical forms of DNA and RNA characteristic of viruses and cells, including simple and complex cells.
[0139] By "nucleic acid molecule" as used herein is meant the nucleic acid sequence that encodes a construct or polypeptide as described above. The nucleic acid molecule can include other operative components, such as regulatory sequences directing expression of the construct or polypeptide in a cell in vivo or in vitro. The nucleic acid molecule can be in a vector.
[0140] By "vector" is meant an entity that delivers the nucleic acid molecule to cells, for therapeutic or cosmetic purposes. As used herein, a vector may include any genetic element including, without limitation, naked DNA, a phage, transposon, cosmid, episome, plasmid, or a virus. Vectors are generated using the techniques and sequences provided herein, in conjunction with techniques known to those of skill in the art. Such techniques include conventional cloning techniques of cDNA such as those described in texts such as Sambrook et al, Molecular Cloning: A Laboratory Manual, 3rd edition, 2001 Cold Spring Harbor Press, Cold Spring Harbor, N.Y., and current editions thereof, use of overlapping oligonucleotide sequences of the adenovirus genomes, polymerase chain reaction, CRISPR, gene editing, and any suitable method which provides the desired nucleotide sequence.
[0141] By the term "attachment" or "attach" as used herein to describe the interaction between the components of the constructs is meant covalent attachments or a variety of non-covalent types of attachment. Still another useful attachment mechanism involves via "affinity interactions", i.e., one domain fused to an antibody fragment that recognizes an epitope on the second domain to be used instead of the two domains fused together. Other attachment chemistries useful in assembling the constructs described herein include, but are not limited to, thiol-maleimide, thiol-haloacetate, amine-NHS, am ine-isothiocyanate, azide-alkyne (CuAAC), tetrazole-cyclooctene (iEDDA).
[0142] The "linker" refers to any moiety used to attach or associate different elements of the polypeptide//polynucleotide sequence components of the constructs (i.e., the effector, the PTD, the targeting moiety) to each other. Thus in one embodiment, the linker is a covalent bond. In another embodiment, the linker is a non-covalent bond. In an embodiment of a polynucleotide described herein, the linker is composed of at least one to about 20 nucleic acids. Thus, in various embodiments, the linker is formed of a sequence of at least 3, 6, 9, 12, 15, 18, 21, 24, 27, 30, 33, 36, 39, 42, 45, 48, 51, 54, 57 up to about 60 nucleic acids. In yet another embodiment of a polypeptide as described herein, the linker refers to at least one to about 20 amino acids. Thus, in various embodiments, the linker is formed of a sequence of at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or about 20 amino acids. In still other embodiments, the linker can be a larger compound or two or more compounds that associate covalently or non-covalently. In still other embodiment, the linker can be a combination of the linkers defined herein. The linkers used in the constructs of the compositions and methods are in one embodiment chemically or enzymatically cleavable, such as by redox, pH, and the like. The linkers used in the constructs of the compositions and methods are in one embodiment non-cleavable.
[0143] The term "nucleic acid molecule" refers to a recombinant assembled nucleic acid sequence encoding a construct or polypeptide described herein. The nucleic acid molecule may be naked DNA or RNA. Alternatively the nucleic acid molecule may be associated operatively with regulatory sequences permitting expression of the construct or polypeptide in vivo or in vitro. The nucleic acid molecule may be a vector, plasmid vector, or be presented in a viral vector for delivery to the subject. Generation of such nucleic acid molecules with resort to the teachings of this specification can utilize known recombinant and genetic engineering techniques. See, e.g., "Inflammatory conditions" as used herein refer, in one aspect to inflammatory skin diseases, which are the most common problem in dermatology and cause pain, redness, swelling and the sensation of heat. Such inflammatory skin conditions include, without limitation, non-specific rashes accompanied by skin itching and redness, sunburn, dermatitis, eczema, rosacea, seborrheic dermatitis, psoriasis, infection, skin injury or wounds, autoimmunity or aging effects. Other inflammatory conditions (non-skin) include, without limitation, autoimmune conditions, asthma, chronic peptic ulcer, tuberculosis, rheumatoid arthritis, periodontitis, ulcerative colitis and Crohn's disease, sinusitis, active hepatitis, gut dysbiosis syndromes and any other disease caused by NFkB/JNK/p38 pathway activation.
[0144] "Patient" or "subject" as used herein means a mammalian animal, including a human, a veterinary or farm animal, a domestic animal or pet, and animals normally used for clinical research. In one embodiment, the subject of these methods and compositions is a human.
[0145] The terms "a" or "an" refers to one or more. For example, "an expression cassette" is understood to represent one or more such cassettes. As such, the terms "a" (or "an"), "one or more," and "at least one" are used interchangeably herein.
[0146] As used herein, the term "about" means a variability of plus or minus 10% from the reference given, unless otherwise specified.
[0147] The words "comprise", "comprises", and "comprising" are to be interpreted inclusively rather than exclusively, i.e., to include other unspecified components or process steps. The words "consist", "consisting", and its variants, are to be interpreted exclusively, rather than inclusively, i.e., to exclude components or steps not specifically recited.
[0148] With resort to the definitions of the components above, in one aspect, the inventors provide a composition comprising a first construct comprising a selected immunomodulatory effector protein or its functional equivalent thereof that targets a first functional domain optionally linked covalently or non-covalently to a selected protein transduction domain (PTD) or penetrating peptide (CPP); an additional construct comprising a different effector protein or a functional equivalent thereof that targets an additional functional domain, optionally linked to the same PTD or CPP of (a) or to an additional PTD or CPP, or a combination of a first and one or more additional constructs.
[0149] In one embodiment, the first and additional constructs are further linked with a targeting moiety to direct activity of the composition to a specific cell or tissue, e.g., skin. In another embodiment, the constructs are admixed in a pharmaceutically or cosmeceutically acceptable carrier or excipient or formulation, such as a formulation suitable for topical administration to the skin.
[0150] One embodiment of such a composition is a single polypeptide comprising a fusion of two or more effectors. Another embodiment is a single polypeptide comprising a first effector-PTD fused construct linked covalently or non-covalently to one or more of additional constructs. The polypeptide in a further embodiment contains one or more optional linker amino acid sequences interposed between each construct of the polypeptide. In one embodiment, the polypeptide contains fused to the first construct and/or additional construct, a targeting moiety to direct the polypeptide to a specific cell or tissue. In a further embodiment, the single polypeptide is in a pharmaceutically or cosmeceutically acceptable carrier or excipient or formulation.
[0151] Whether the constructs are admixed in a composition or present in one or more polypeptides, the effector can be one or more of the effectors identified above, particularly in Tables 1 or 2. In one embodiment of the admixture or polypeptide, the first effector protein is NleE or a functional equivalent thereof. In yet other embodiment, the additional effector protein is one or more of NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP. In other embodiments, the admixture of constructs in the composition or present in the single polypeptide can be selected from the following embodiments. In one embodiment, the first effector is NleE and the additional effector is NleC. In another embodiment, the first effector is NleE and the additional effector is NleD. In another embodiment, the first effector is NleE and the additional effector is NleB. In another embodiment, the first effector is NleE and the additional effector is NleH. In another embodiment, the first effector is NleE and the additional effector is YopB. In another embodiment, the first effector is NleE and the additional effector is YopH. In another embodiment, the first effector is NleE and the additional effector is YopJ. In another embodiment, the first effector is NleE and the additional effector is YopP. In still another embodiment, the first effector is NleE and the additional effector is SspH1. In a further embodiment, the first effector is NleE and the additional effector is OspG. In another embodiment, the first effector is NleE and the additional effector is OspF. In another embodiment, the first effector is NleE and the additional effector is IpaH9.8. In still a further embodiment, the first effector is NleE and the additional effector is IpaH1.4. In another embodiment, the first effector is NleE and the additional effector is IpaH2.5. In another embodiment, the first effector is NleE and the additional effector is IpaH4.5. In another embodiment, the first effector is NleE and the additional effector is IpaH7.8. In yet a further embodiment, the first effector is NleE and the additional effector is SIrP. In still other embodiments of the constructs or polypeptides, three or more effectors can be delivered in a single mixture of constructs or single polypeptide. The first and additional effectors or effector constructs may occur in any order in the fusions or in the polypeptides described herein.
[0152] In the above-listed constructs and polypeptides, a suitable PTD or CPP linked to the first construct and optionally to each additional construct or polypeptide is the naturally occurring PTD of the selected effector protein, e.g., the YopM PTD (SEQ ID NO: 2; amino acids 1-50 of YopM). See U.S. Pat. No. 8,840,901.
[0153] The PTD/CPP can also be a functional equivalent, e.g., a mutated or modified version of a naturally occurring PTD, such as the naturally occurring PTD sequence of SspH1, or of Shigella IpaH protein or any other of the effectors. The PTD useful in the constructs and polypeptides of the compositions can be a completely novel sequence which is developed to transport the construct or polypeptide or composition across cell membranes.
[0154] In one specific embodiment, the first construct is the YopM PTD fused to NleE. In another specific embodiment, the first construct is the SspH1 PTD fused to NleE. In still another specific embodiment, the first construct is the IpaH PTD fused to NleE. In still other embodiments, the additional construct comprises YopM, optionally associated with the YopM PTD. In still other embodiments of the single polypeptide, the polypeptide further contains at least one linker sequence as defined above interposed between the first and the one or more additional constructs. In still other embodiments of the constructs and single polypeptide, each construct or polypeptide can further contain a fused targeting moiety, such as a skin cell targeting peptide or other cell or tissue targeting peptide. Given the number of effectors, PTD/CPPs, linkers and targeting moieties identified herein and in the cited publications, any number of constructs or polypeptides may be prepared according to the teachings contained herein.
[0155] In still other embodiments of the above-listed constructs and polypeptides, the admixture of protein constructs or constructs contained on the single polypeptide is such that the targeted functional domain of each effector protein in the construct is expressed in the same cell. In another embodiment, each construct in admixture or in the single polypeptide has a non-overlapping redundant role in inhibiting NFkB, JNK and p38 pathways when present in a mammalian cell.
[0156] As described above, any of the compositions described herein, whether a mixture of constructs, a single polypeptide or nucleic acid molecules encoding them, can be prepared in a formulation comprises ingredients suitable for application to, and absorption through, the cells of the skin. One such formulation employs the above-described CAGE solvent. Other suitable formulations may include oils, emollients, lotions for topical and transdermal applications along with buffered/aqueous and saline solutions.
[0157] In yet another aspect, the compositions described herein include nucleic acid molecules comprising a nucleic acid sequence encoding a first construct, an additional construct, or the single polypeptide of any one of the above-described embodiments. Such nucleic acid molecules can also comprise operatively associated regulatory sequences, such as promoters, enhancer, etc (see, e.g., Sambrook et al) necessary to express the construct or polypeptide in a suitable cell. In certain embodiments, the nucleic acid molecule is naked DNA or RNA. In certain embodiments, the molecule is part of a plasmid or contained in a recombinant vector or virus. Methods for generating such nucleic acid molecules are within the skill of the art given the teachings herein. The nucleic acid molecules may also be delivered in a cosmeceutically or pharmaceutically acceptable carrier or excipient or formulation. Such formulations are described in detail in available texts as described above.
[0158] These constructs and/or polypeptides may be formulation into a wide variety of cosmeceutically or pharmaceutically acceptable carrier or excipient or formulation for many different uses and routes of administration. While topical administration is preferred for uses on skin inflammation, it is contemplated that other conventional routes of administration will be used for treating other inflammatory conditions. In some embodiments, routes of administration include transdermal (including patch formulation), intra-dermal injection (including infusion and subcutaneous injection). Other pharmaceutically acceptable routes of administration include, but are not limited to, systemic routes, such as intraperitoneal, intravenous, intranasal, intramuscular, intratracheal, subcutaneous, epidural, and oral routes and other parenteral routes of administration or intratumoral or intranodal administration. The composition may be administered by any convenient route, for example by infusion or bolus injection, by absorption through epithelial or mucocutaneous linings (e.g., oral mucosa, rectal and intestinal mucosa, etc.) and may be administered together with other biologically active agents. Administration can be systemic or local. In addition, it may be desirable to introduce the pharmaceutical compounds or compositions of the invention into the central nervous system by any suitable route, including intraventricular and intrathecal injection; intraventricular injection may be facilitated by an intraventricular catheter, for example, attached to a reservoir, such as an Ommaya reservoir. Pulmonary administration can also be employed, e.g., by use of an inhaler or nebulizer, and formulation with an aerosolizing agent. Routes of administration may be combined, if desired.
[0159] Other methods of delivery of the effectors include via an attenuated bacterial strain which expresses a functional Type-Three Secretion System (TTSS)-expressing microorganism and is engineered to contain nucleic acid sequences encoding at least one of the effector immuno-modulatory proteins or fusions or polypeptide, wherein amino acid sequences also code for functional TTSS secretion signal sequences. These include attenuated bacteria engineered or induced to shed outer membrane vesicles (OMV) or other type of exosome-like, bacterial or cell-derived vesicle containing the proteins described herein.
[0160] In still other embodiments, the compositions, constructs, fusions and polypeptides and nucleic acid molecules described herein may be further manipulated by encapsulation in liposomes, micro particles, microcapsules, or in recombinant cells capable of expressing the compound, receptor-mediated endocytosis construction of a nucleic acid as part of a retroviral or other vector, etc.
[0161] Still other embodiments of compositions are provided herein. In certain embodiments, a composition of single effectors, or fusions of multiple effectors, or fusions of single effector-PTD, or fusions of multiple effectors and one or more PTD, or single/multiple effector(s)-PTD-targeting moiety fusions are prepared. The polypeptide linker regions and other non-native peptide sequences are engineered into fusion-effectors and can contain active segments which can lead to their enhanced ability to be purified in active form. In another embodiment, the effector fusions can be modified recombinantly or chemically, or by e.g., editing methodologies to provide altered subcellular localization, altered stability, or altered ability to participate in protein-protein interactions. In one embodiment, the effector fusions are modified to include amino acid segments normally used as tags for efficient purification of proteins from complex mixtures (e.g., 6HIS, GST, and Maltose BP).
[0162] In still a further specific embodiment, each individual effector protein or multiple effector fusion has an amino acid sequence that is deliberately altered by designed site directed mutagenesis. In one embodiment, such mutagenesis alters the activity of the effector in terms of its potency as an anti-inflammatory effector. In another embodiment, the altered effector amino acid sequence alters the effector's specificity for a target, catalytic activity, antigenicity, stability, ability to bind substrate, and/or ability to work with other effectors in a preparation. Such mutagenic techniques can alter any other common property of an effector protein for the benefit of the efficacy of the resulting therapeutic preparation.
[0163] In another aspect, the compositions containing the effector fusions can lead to the inhibition of at least one or more of an NF-.kappa.B, JNK and p38 mediated signal transduction pathway protein in selected cells and tissues. Such inhibition provides an anti-inflammatory response, an anti-apoptotic effect or a pro-apoptotic effect in a target cell.
[0164] Still other embodiments of compositions can be obtained by selection of the components as taught herein resulting in a pharmaceutical/cosmeceutical preparation, containing active and therapeutic levels of the effector fusions and constructs as described herein for use in preventing, treating, or ameliorating an immune-related disorder whose pathology stems from aberrant activation of the NFkB, JNK or P38 pathways.
[0165] In still other aspects, methods for designing the various constructs and polypeptides described herein can be made using techniques well-known in the art of recombinant genetic engineering and manipulation of the nucleic acid techniques. Suitable techniques are known to those of skill in the art, and as provided by many of the publications incorporated by reference herein. Selection of the appropriate techniques to create the various embodiments will depend upon the effectors, PTDs, linkers, targeting moieties and other components as set out above. Molecular biology strategies to alter the proteins to make them more stable, specific, catalytic and robust are known. Further such known techniques can be used to optimize the design, construction and delivery of effector fusions based upon knowledge of the specific pathways and enzymes used by the cell in a selected inflammatory stimulus.
[0166] For example, in one embodiment the selected effectors are subject to random, high throughput mutagenesis in the catalytic active sites or the sites of substrate binding. Selection of effectors of these mutagenized populations for variants with novel substrate specificities, rates of catalysis and their combination with other known or mutagenized effectors in the fusions, constructs and polypeptides described herein can result in compositions with novel signal transduction inhibition functions.
[0167] Still other known and available methods can be employed for creating recombinant fusion effector genes/proteins and for their expression, purification, stabilization as single effectors or fusion effectors. Such methods include producing/expressing the effector protein or effector fusion proteins from cloned genes in any suitable cell based systems, including, without limitation, bacteria, yeast, insect, or mammalian cells. Methods for purification of the resulting fusion constructs resulting in highly active, stable proteins which retain the ability to target their native substrate also employ known techniques. Additional techniques for producing compositions of this invention include, without limitation, optimization of the effector preparation by incorporation into liposomes, PEGylation of the effector constructs or fusions or polypeptides. The effector constructs may be subjected to chemical derivatization to associate or conjugate the construct physically or in admixture with additional entities. Such entities include without limitation, lipids, liposomes, other drugs, and cargo molecules including nucleic acids, polypeptides, organic molecules, small organic molecules, metals, nano-particles, viruses, modified viruses, viral vectors, antibodies and/or plasmids as targets for conjugation.
[0168] Yet further aspects provide a variety of methods of using the compositions, fusions, constructs and polypeptides described above in methods for treating cosmetic conditions, such as inflammation or irritation due to normal skin agent, to the treatment of other inflammatory conditions, such as cancer, gut dysbiosis syndromes, and other conditions identified herein caused by NFkB/JNK/p38 pathway activation. For example one therapeutic use is to ameliorate the inflammation in the tumor microenvironment (TME) in any malignant or non-malignant condition that involves activation of the NFkB/JNK/p38 pathway.
[0169] The various compositions defined above may be employed therapeutically in the down regulation/normalization of the immune response elicited by the ectopic or pathologic activation of NFkB, JNK and P38 pathways. In one embodiment, such methods of treating or suppressing pathway activation can occur in subjects suffering from the consequences of normal, naturally occurring conditions such as aging associated inflammation. In yet another method, the compositions are administered for treatment of numerous conditions wherein the three pathways are aberrantly activated resulting in an inflammatory response. Such aberrant responses include without limitation, infection, wound healing, reactive dermatitis, auto-immune disease, and malignant or non-malignant proliferative disorder. These diseases, conditions and syndromes wherein the three pathways are aberrantly activated also include, but are not limited to diseases caused by autoimmunity of the patient, topical inflammation, chronic inflammation, gastroenteritis, chronic gastritis, inflammatory bowel diseases (IBD), colitis ulcerosa, psoriasis, allergic reactions, Crohns disease, dysbiosis syndromes, cancer (including gliobastoma) rheumatoid arthritis related bone diseases characterized by changes in bone resorption, reduction and relief of the signs and symptoms associated with treating inflammation, and/or suppression of the immune system.
[0170] These methods involve administering via a suitable route of administration (as described above) an amount of the composition (i.e., the effector fusions, effector-PTD fusions, effector-PTD-Targeting fusions, and the other embodiments described above) in an amount sufficient to reduce activation of the desired pathway.
[0171] Desirably, the methods further involve, in one aspect, administering a suitable dose or doses of the fusion construct(s) in a therapeutic regimen to the subject in need thereof. In one embodiment, such administration can occur once or more prior to, simultaneously with, or after any conventional additional treatment for the conditions. In one embodiment, where the condition is an infection, the additional component is an antibiotic. In another embodiment, wherein the condition is a cancer, the additional component is radiation or chemotherapy. Still other known "additional components" may be selected by one of skill in the art or the attending physician depending on the condition being treated and the physical status of the subject.
[0172] Methods for determining the timing of frequency of administration will include an assessment of subject's response to the first administration of the composition. The dose is generally the lowest dose of the composition that is effective to suppress activation of the NFKB, JNK/p38 pathway. In still other embodiments, a suboptimal dose is delivered in a continuous infusion or a slow release formulation. The dosage required will depend primarily on factors such as the condition being treated, the age, weight and health of the patient, and may thus vary among patients. In one embodiment, where the composition comprises an admixture of peptide constructs or the single polypeptide, one such dose is about 1 to 25 .mu.M protein/polypeptide. In another embodiment, the dose is less than 10 .mu.M protein/polypeptide. In still another embodiment, the dose is between 1 .mu.M and 5 .mu.M protein/polypeptide. In another embodiment, the suboptimal dose is less than 1 .mu.M protein/polypeptide.
[0173] When the composition is in the form of a nucleic acid or vector or nucleic acid molecule, it is administered in sufficient amounts to transduce the targeted cells and to provide sufficient levels of gene transfer and expression to reduce or inhibit activation of the NFKB, JNK/p38 pathway and provide a therapeutic benefit without undue adverse or with medically acceptable physiological effects, which can be determined by those skilled in the medical arts. Dosages of these therapeutic compositions will depend primarily on factors such as the condition being treated, the age, weight and health of the patient, and may thus vary among patients. For example, a therapeutically effective adult dosage of a viral vector or siRNA nanoparticle is generally in the range of from about 100 .mu.L to about 100 mL of a carrier containing concentrations of from about 1.times.10.sup.6 to about 1.times.10.sup.15 particles, about 1.times.10.sup.11 to 1.times.10.sup.13 particles, or about 1.times.10.sup.9 to 1.times.10.sup.12 virus particles.
[0174] In still another aspect, methods for use of the compositions described herein involve veterinary use for the treatment of inflammatory conditions in animals, e.g., for auto-immune diseases, reactive dermatitis, pruritis, alopecia and any other skin ailment for which the underlying pathogenesis involves alteration of NFKB, JNK/p38 pathway. Selection of administration routes, dosages and therapeutic regimens may be selected by a veterinarian.
[0175] In one aspect, a composition in a pharmaceutically acceptable carrier or excipient or formulation is provided. The composition includes: (a) a first construct comprising a selected immunomodulatory effector protein or functional equivalent thereof that targets a first functional domain optionally linked covalently or non-covalently or by affinity to a selected protein transduction domain (PTD) or penetrating peptide (CPP); (b) an additional construct comprising a different effector protein or a functional equivalent thereof that targets an additional functional domain, optionally linked to the same PTD or CPP of (a) or to an additional PTD or CPP, or (c) a combination of constructs (a) and (b) in any order.
[0176] In one embodiment of the composition, the first construct is linked covalently or non-covalently to one or more of the additional constructs in a single polypeptide. Additionally, the composition further includes an optional linker amino acid sequence interposed between each construct.
[0177] In one embodiment, the functional equivalent includes a chemically or recombinantly modified amino acid sequence of the effector protein, or a fragment of the naturally-occurring effector amino acid sequence, or of the derivative of said chemically or recombinantly modified amino acid sequence of the effector protein that shares the functional activity of the effector protein.
[0178] In one embodiment, the first effector protein is NleE or a functional equivalent thereof.
[0179] In one embodiment, the additional effector protein is one or more of NleC, NleD, NleB, NleH, YopM, YopE, YopH, YopJ, YopP, SspH1, OspG, OspF, IpaH9.8, IpaH1.4, IpaH2.5, IpaH4.5, IpaH7.8 and SIrP. In one embodiment, the PTD or CPP is the naturally occurring PTD of the selected effector protein. In one embodiment, the PTD is a sequence of YopM, of SspH1, or of Shigella IpaH protein. In one embodiment, the PTD is amino acids 1-50 SEQ ID NO: 2.
[0180] In one embodiment, the PTD or CPP is one or more of Poly-Arg, Tat and VP22, df Tat, a cyclic CPPs, IMT-P8, seven arginine (R7) and Streptolysin 0 (SLO)-mediated systems, elastin like polypeptide, CPP-adaptor system, 1, 2-Benzisothiazolin-3-one (BIT) and Tat, activatable cell-penetrating peptides, LDP12, M918, BR2, POD, native protein independent of R11-CPP, Poly-arginine/Tat and Tat-PTD, Pep-1, CADY-2, R8, azo-R8, Penetratin, HR9 and IR9 peptides, or pVEC.
[0181] In one embodiment, each targeted functional domain of each effector protein in the construct is expressed in the same cell. In one embodiment, each construct has a non-overlapping redundant role in inhibiting NFkB, JNK and p38 pathways when present in a mammalian cell. In one embodiment, the formulation comprises ingredients suitable for application to, and absorption through, the cells of the skin. In one embodiment, the formulation comprises CAGE solvent. In one embodiment, the first construct is the YopM PTD fused to NleE or the SspH1 PTD fused to NleE or the IpaH PTD fused to NleE. In one embodiment, the additional construct comprises YopM, optionally associated with the YopM PTD. In one embodiment, the composition further includes at least one linker between the first and additional constructs.
[0182] In one embodiment, the first effector is NleE and the additional effector is NleC. In another embodiment, the first effector is NleE and the additional effector is NleD. In another embodiment, the first effector is NleE and the additional effector is NleB. In another embodiment, the first effector is NleE and the additional effector is NleH. In another embodiment, the first effector is NleE and the additional effector is YopB. In another embodiment, the first effector is NleE and the additional effector is YopH. In another embodiment, the first effector is NleE and the additional effector is YopJ. In another embodiment, the first effector is NleE and the additional effector is YopP. In another embodiment, the first effector is NleE and the additional effector is SspH1. In another embodiment, the first effector is NleE and the additional effector is OspG. In another embodiment, the first effector is NleE and the additional effector is OspF. In another embodiment, the first effector is NleE and the additional effector is IpaH9.8. In another embodiment, the first effector is NleE and the additional effector is IpaH1.4. In another embodiment, the first effector is NleE and the additional effector is IpaH2.5. In another embodiment, the first effector is NleE and the additional effector is IpaH4.5. In another embodiment, the first effector is NleE and the additional effector is IpaH7.8. In another embodiment, the first effector is NleE and the additional effector is SIrP. In any of the embodiments above, the constructs may be in any order.
[0183] In one embodiment, the composition further includes a skin cell targeting peptide.
[0184] In another aspect, a nucleic acid construct is provided. The nucleic acid sequence includes a nucleic acid sequence encoding the polypeptide of any of the compositions described above.
[0185] In one embodiment, the nucleic acid construct further includes regulatory sequences necessary to express the polypeptide in a suitable cell.
[0186] In one embodiment, the nucleic acid construct is a DNA, RNA, a plasmid or a recombinant vector or virus.
[0187] In one embodiment, the nucleic acid construct is in a pharmaceutically acceptable carrier or excipient or formulation.
[0188] In another aspect, a method is provided for treating, preventing or ameliorating an NFkB, JNK or p38 mediated inflammatory disorder in a subject, or a subject at risk from developing such disorder. The method includes administering to the subject any of the compositions or nucleic acid constructs described above.
[0189] In one embodiment of the method, the composition is administered in an amount sufficient to down-regulate the innate inflammatory response in a targeted cell or tissue of the subject mediated by the pathologic or non-pathologic activation of intracellular NFkB, JNK or P38 signal transduction pathway.
[0190] In one embodiment, the method reduces symptoms, and inhibits progression, of damage to the subject's cells or tissue caused by inflammation.
[0191] In one embodiment of the method, a composition containing multiple constructs targets multiple targeted functional domains that are expressed in the same cell and each construct plays multiple, non-overlapping redundant roles in inhibiting NFkB, JNK and p38 pathways.
[0192] In one embodiment of the method, the presence of multiple constructs in the compositions produces a synergistic therapeutic effect.
EXAMPLES
Example 1: Preparation of Recombinant Constructs
[0193] Sequences. The native YopM, NleE, OSPZ, and IpA7.8 nucleotide sequences were codon optimized for efficient bacterial expression of the encoded polypeptides. The optimized nucleotide sequences were synthesized de novo as long single stranded nucleotide chains. The overlapping and complementary single strand DNA sequences were annealed and the mixture subject to polymerize chain reaction (PCR) to generate full-length double-stranded (ds) DNA. The ds DNA was gel isolated and cloned into plasmid pQE60. The sequence of the inserted DNA was confirmed by nucleotide sequence analysis.
[0194] TAT-NleE wild type and NleE Mutant R107A. Both Vector and Amplicon were digested using BamH1 restriction enzyme overnight at 37.degree. C. The Vector was then phosphatase treated for 1 hour then heat inactivated prior to ligation. All material was run through agarose gels and gene cleaned using a Qiagen gel extraction kit. Ligation reaction was performed at 16.degree. C. overnight, and half of the volume was used to transform the DNA into XL-1 blue competent cells. Colonies from the subsequent transformation were picked and mini-preps were performed using a Qiagen Mini-Prep Kit. Resulting DNA was digested using BamH1 for 1 hr at 37.degree. C. to test for the presence of the insert. 675 bp Positive clones were then digested using EcoR5 (internal to the insert) and external Hind3 site for 1 hr at 37.degree. C. to test the orientation of the insert. Positive orientation yielded a drop-out that was 175 bp in size. Positive clones were then transformed into S9-competent cells to test protein expression rates by mini-induction. A clone was chosen from that group, and used for future maxi-induction purifications under native conditions. A schematic of the construct (based upon the backbone plasmid pQE60 available from Qiagen Inc, Catalogue number: #32903) is shown in FIG. 3.
[0195] TAT-Shigella OSPZ. Both Vector and Amplicon were digested using BamH1 restriction enzyme overnight at 37*C. The Vector was then phosphatase treated for 1 hr then the enzyme was heat inactivated prior to ligation. All material was electrophoresed through agarose gels and the DNA isolated from the gel using a Qiagen DNA gel extraction kit. Ligation reaction was performed at 16*C overnight, and half of the volume was used to transform the DNA into XL-1 blue competent cells. Colonies from the subsequent transformation were picked and mini-preps were performed using a Qiagen Mini-Prep Kit. Resulting DNA was digested using BamH1 37*C for 1 hr to test for the presence of the 900 bp insert. Positive clones were then digested using EcoR5 (internal to the insert) and Hind3 located after the HIS-tag 37*C 1 hr to test the orientation of the insert. Positive orientation yielded a drop-out DNA fragment that was 200 bp in size. Positive clones were then transformed into S9-competent cells to test protein expression rates by mini-induction. A clone was chosen from that group, and used for future maxi-induction of proteins and purification of protein under native conditions. A schematic of the construct is shown in FIG. 4.
[0196] YopM PTD-NleE wild type and Mutant R107A. Both Vector and Amplicon were digested using BamH1 restriction enzyme overnight at 37*C. The Vector was then phosphatase treated for 1 hr and heat inactivated prior to ligation. All material was run through agarose gels and gene cleaned using a Qiagen gel extraction kit. Ligation reaction was performed at 16*C overnight, and half of the volume was used to transform the DNA into XL-1 blue competent cells. Colonies from the subsequent transformation were picked and mini-preps were performed using a Qiagen Mini-Prep Kit. Resulting DNA was digested using Barn H1 at 37*C for 1 hr to test for the presence of the insert. Positive clones were then digested using Bgl2 (internal to the insert) and Bgl2 at 37*C for 1 hr to test the orientation of the insert. Positive orientation yielded a drop-out that was 275 bp in size. Positive clones were then transformed into S9-competent cells to test protein expression rates by mini-induction. A clone was chosen from that group, and used for future maxi-induction purifications under native conditions. A schematic of the construct is shown in FIG. 5.
[0197] YopM PTD-NleE wild type and Mutant R107A, No Linker, PAPA Linker, and GSGS Linker. A QE60 vector containing a codon optimized YopM sequence was purchased from Epoch Life Science Inc. The plasmid was digested using Cla1 (internal to YopM) and Bgl2 (located outside of the cloning site). NleE was then digested using Cla1 and Bam H1. Note that after ligation, the BamH1/Bgl2 site on the vector was lost. Because the vector was opened at a site that was internal to YopM, the ligation of NleE (along with its mutant R107A counterpart) lead to the fusion of the two inserts. This site is located downstream to the PTD, thus leaving it intact. Variations of this construct were made using either PAPA or GSGS linkers. These were added to the beginning of NleE during PCR by having the bridges be part of the forward primer sequence. Screening of these various constructs were done via PCR of mini-preps using NleE Cla1 FOR and TNHis Barn REV primers designed for NleE. Positive clones were then sent for sequencing to verify the clones prior to transformation into S9-cells for protein expression. A schematic of the construct is shown in FIG. 6.
[0198] Schematics of the constructs used are shown in FIG. 1 and FIG. 2.
[0199] Purification of recombinant polypeptides. The transformed SG13009 [pREP4] were plated on an agar plate containing both ampicillin and kanamycin. A single colony was used to inoculate a 200 mL culture, shaking at 200 RPM at 37.degree. C. overnight. A volume of the overnight culture was added to 1 L of 2YT medium containing ampicillin and kanamycin and grown as described above in an incubator shaker at 200 RPM 37.degree. C. to an OD 600 nm of 0.6-0.8. Then, 1 mL of 1M Isopropyl .beta.-D-1-thiogalactopyranoside (IPTG) was added to the culture and growth was continued overnight with shaking at 200 RPM at 20.degree. C. The following morning, cells were collected by centrifugation at 6,000 RPM at 4.degree. C. The pelleted cells were resuspended in 40 mL Sonication Buffer and sonicated on ice for two cycles of 4-5 minutes each. The sonification buffer had a pH of 7.5 and 1 L of sonification buffer included one bottle of PBS, 300 mM NaCl, 10% Glycerol, 5 mM Imidazole. The remainder of the volume was filled with ddH.sub.2O. The buffer was then filtered and 1 ml 0.1M PMSF and 1 ml (2 mg/ml) each of Aprotinin, Leupeptine, and Pepstatin were added fresh. The sonicate was centrifuged 12,000 RPM at 4.degree. C. for 30 minutes to separate lysed cells from soluble protein.
[0200] The supernatant was collected and incubated with pre-equilibrated Ni-NTA beads (available from BioRad) for 1 hour at 4.degree. C. by rotation. The binding capacity of polypeptides for Ni-NTA beads was about 5-10 mg/ml. Following the binding step, bound materials were centrifuged 4,000 RPM 4.degree. C. for 10 minutes to pellet the Ni-NTA beads and separate from unwanted materials or "flow-through". Beads were then washed twice in 40 mL wash buffer by rotation in 4.degree. C. for 30 minutes per wash cycle. The was buffer had a pH of 7.5 and 1 L of wash buffer included one bottle of PBS, 300 mM NaCl, 10% Glycerol, and 20 mM Imidazole. The remainder of the volume was filled with ddH.sub.2O In addition, 1 ml 0.1 M PMSF was added fresh. The washed Ni-Beads were loaded on to a disposable gravity drip column and washed with residual Wash Buffer.
[0201] Bound proteins were eluted with elution buffer, using 2.times. elution buffer per 1 mL of packed volume of beads, i.e., 2 ml elution buffer for 1 ml of Ni-NTA beads. The elution buffer had a pH of 7.5 and 100 ml of elution buffer contained one bottle of PBS, 300 mM NaCl, 10% Glycerol, and 500 mM Imidazole. The remainder of the volume was filled with ddH.sub.2O. The elution buffer was filtered before use. Eluate was collected in 2 mL fractions. The protein content of the fractions was analyzed using Bradford reagent. Individual fractions were dialyzed overnight in dialysis buffer at 4.degree. C. The dialysis buffer had a pH of 7.5 and 4 L of elution buffer included one bottle of PBS, 300 mM NaCl, and 10% Glycerol. The remainder of the volume was then filled with ddH.sub.2O. The following were freshly added to the elution buffer: 1 ml 0.1 M PMSF and 1 ml 1 M DTT. In the morning, the dialysis buffer was discarded and replaced with fresh buffer and dialysis was continued for an additional 3 hours. After dialysis, the individual fractions were pooled and centrifuged at 12,000 RPM at 4.degree. C. for 20 minutes to remove any precipitation. The soluble protein was concentrated in the final concentration was determined using Bradford reagent.
[0202] Purified polypeptides were analyzed by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) analysis using known volumes of bovine serum albumin (BSA) as standards. Concentrated samples were aliquoted and stored at -80.degree. C. Proteins were used directly after thawing on ice. An SDS-PAGE analysis of purified recombinant chimeric effector proteins is shown in FIG. 7. Each lane contained 2 .mu.g of protein. Proteins were visualized by Coomassie blue staining.
Example 3: Cytokine Release
[0203] The effect of the bacterial effector proteins on cytokine release was analyzed in a cell-based assay using the human monocyte cell line, THP-1. Cytokine release was measured in vitro using the LEGENDplex Multi-Analyte Flow Assay Kit (BioLegend, Cat no. 740118) according to the supplier's directions. For culturing THP-1 cells, RPMI with glutamine and 20% FBS was used. These cells were incubated at 37.degree. C. in humidified air 5% CO2 atmosphere. Finally, cell medium was changed every 2-3 days, and the cells were passaged at 95% confluence. To allow the cells to adapt to their environment, 3.times.10.sup.5 cells were added to a 24-well plate the night before. The next day, protein was added to these cells for a period of 3 hours. The protein was then washed out with PBS. After cells were stimulated overnight with LPS stimulation, a concentration of 1 ug/mL was used. The next day, cells were washed again and cytokine were measured. All 13 inflammatory cytokines were measured using the LEGENDplex Human Inflammation Plane (13-plex) assay from Biolegend.
[0204] We analyzed the effect of effector protein constructs on the release of the pro-inflammatory cytokines IL-1beta, TNF-alpha, 11-6, MCP-1, IL 23 and IL-8. The effector protein constructs are shown in the table below:
TABLE-US-00006 Protein SEQ Construct transduction ID name domain Effector 1 Linker Effector 2 NO YopM Full- YopM YopM 3 length TAT-NIeE TAT EPEC NLeE -- -- 28 TAT Shigella TAT Shigella 34 OSPZ OSPZ YopM PTD- YopM EPEC NLeE -- -- 37 NIeE YopM PTD- YopM YopM -- EPEC NLeE 10 NIeE (No (L-rich) linker) YopM PTD- YopM YopM GSGS EPEC NLeE 16 NIeE (GSGS (L-rich) linker) YopM PTD- YopM YopM PAPA EPEC NLeE 13 NIeE (PAPA (L-rich) linker) YopM PTD- YopM YopM -- -- 7 YopM (L-rich) (L-rich) IpaH 7.8 IpaH 7.8 IpaH 9.8 IpaH 9.8
[0205] The results of these experiments are shown in FIGS. 8, 9, 10, 11 and 12. As shown in FIGS. 8, 9, 10, 11 and 12, the effector proteins reduced the levels of cytokines released into the medium relative to control samples that did not receive effector proteins. This effect was dose-dependent, with higher doses of effector proteins resulting in a steeper reduction in levels of released cytokines. These dose-dependent effects are shown in FIG. 8 with respect to IL-6. Similar dose-dependent effects were observed for IL-1beta, TNF-alpha, 11-6, MCP-1, IL 23, and IL-8.
[0206] The effector fusion proteins that included more than one effector protein domain produced a steeper reduction in the level of all released cytokines than did native effector proteins. As shown in FIG. 9, the paired effector fusion proteins YopM PTD-YopM (L-rich)-NLeE; YopM PTD-YopM (L-rich)-GSGS-NLeE; YopM PTD-YopM (L-rich)-PAPA NLeE; and YopM PTD-YopM (L-rich) produced a greater reduction in levels of TNF alpha released into the medium shown than did the full length YopM (rYopM) or Shigella OSPZ. This reduction was observed for effector fusion proteins that did not include a linker sequence and for effector proteins that included either a GSGS or a PAPA linker sequence.
[0207] As shown in FIG. 10, the paired effector fusion proteins YopM PTD-YopM (L-rich)-NLeE; YopM PTD-YopM (L-rich)-GSGS-NLeE; YopM PTD-YopM (L-rich)-PAPA NLeE; and YopM PTD-YopM (L-rich) produced a greater reduction in levels of IL-6 released into the medium shown than did the full length YopM (rYopM) or Shigella OSPZ. This reduction was observed for effector fusion proteins that did not include a linker sequence and for effector proteins that included either a GSGS or a PAPA linker sequence.
[0208] As shown in FIG. 11, the paired effector fusion proteins YopM PTD-YopM (L-rich)-NLeE; YopM PTD-YopM (L-rich)-GSGS-NLeE; YopM PTD-YopM (L-rich)-PAPA NLeE; and YopM PTD-YopM (L-rich) produced a greater reduction in levels of MCP-1 released into the medium shown than did the full length YopM (rYopM) or Shigella OSPZ. This reduction was observed for effector fusion proteins that did not include a linker sequence and for effector proteins that included either a GSGS or a PAPA linker sequence.
[0209] As shown in FIG. 12, the paired effector fusion proteins YopM PTD-YopM (L-rich)-NLeE; YopM PTD-YopM (L-rich)-GSGS-NLeE; YopM PTD-YopM (L-rich)-PAPA NLeE; and YopM PTD-YopM (L-rich) produced a greater reduction in levels of IL-23 released into the medium shown then did the full length YopM (rYopM) or Shigella OSPZ. This reduction was observed for effector fusion proteins that did not include a linker sequence and for effector proteins that included either a GSGS or a PAPA linker sequence.
[0210] Similar results were observed for IL-8. Taken together, the results shown in FIGS. 8, 9, 10, 11, and 12 indicate that fusion proteins that target multiple inflammatory pathways are more effective at reducing an inflammatory response than are proteins that target a single inflammatory pathway.
Example 4: Caspase Activity
[0211] The effect of YopM on caspase activity was measured using R&D System's Caspase-1/ICE Colorimetric Assay Kit (K111-100) according to the supplier's instructions. THP-1. cells were incubated with rYopM or a truncated YopM (YopMo) and then stimulated with LPS (lipopolysaccharide) and ATP to induce activation of caspase 1. More specifically, 2.times.10.sup.6 THP-1 cells were seeded in 6-well cell culture plates in triplicate. The cells were incubated for 2 hours with rYopM (25 .mu.g/ml) or a truncated YopM (YopMo) and then LPS was added (1 .mu.g/mL) and the cells were incubated for an additional for 4 hours. Finally, ATP (5 mM) was added and the cells were incubated for 45 minutes. To pellet the cells, centrifugation was carried out at 250.times.g for 10 min. The cell pellet (2.times.10.sup.6 cells) was resuspended in 50 .mu.l lysis buffer (Caspase 1/ICE Colorimetric Assay Kit R & D Systems) and incubated on ice for 10 min. Cell debris were pelleted at 10,000.times.g for 1 min. The colorimetric assay was carried out in 96-well plates. For this purpose, 50 ul cell lysate/well were placed and mixed with 50 ul 2.times. reaction buffer (Caspase 1/ICE Colorimetric Assay Kit R & D Systems). After addition of 5 .mu.l of the substrate Ac-YVAD-pNA, incubation was carried out at 37.degree. C. for 3 h. Subsequently, the absorbance at 405 nm was measured.
[0212] The results of this experiment are shown in FIG. 13. As shown in FIG. 13, a dose-dependent reduction of caspase 1 activity was observed in THP-1 cells that had been treated with rYopM. A dose-dependent reduction of caspase 1 activity was also observed in THP-1 cells that had been treated with the truncated YopM (YopMo). These data showed that the truncated YopM (YopMo) that contained only the L-rich region of the YopM retained the caspase I reducing function of the full length YopM.
Examples 5: Effector Polypeptide Uptake
[0213] Confocal microscopy. Cell uptake of effector polypeptides was analyzed using confocal microscopy. We analyzed single cell uptake of the following fusion protein constructs:
TABLE-US-00007 Protein SEQ Construct transduction ID name domain Effector 1 Linker Effector 2 NO TAT-NIeE TAT EPEC NLeE -- -- 28 YopM PTD- YopM YopM -- EPEC NLeE 10 NIeE (No (L-rich) linker) YopM PTD- YopM YopM GSGS EPEC NLeE 16 NIeE (GSGS (L-rich) linker) YopM PTD- YopM YopM PAPA EPEC NLeE 13 NIeE (PAPA (L-rich) linker)
[0214] Fusion proteins were labeled with FITC using the Sigma-Aldrich FluoroTag.TM. FITC Conjugation Kit according to the supplier's instructions as follows. Protein and FITC were dissolved in carbonate-bicarbonate buffer. The FITC was slowly added to the protein with stirring, then covered with foil and stirred for two hours at room temperature. The conjugate was separated from free FITC on a G-25 column in the fractions were collected. Fractions containing conjugate were pooled and the F/P ratio of conjugate was determined spectrophotometrically. The labeled protein was stabilized with 1% bovine serum albumin and 0.1% sodium azide and stored at 0-5.degree. C.
[0215] HaCat cells were cultured using DMEM High glucose medium with 10% FBS. These cells were incubated at 37.degree. C. in humidified air 5% CO2 atmosphere. Finally, cell medium was changed every 2-3 days, and the cells were passaged at 95% confluence. Cells were grown to appropriate confluency. Next, appropriate coated coverslips were added to well plates and the cells were seeded in the well plates containing the coverslips. For uptake analysis, polypeptides were added to the cells to a final concentration of 50 ug/mL and incubated for two hours. The protein with the cells were incubated and the protein was aspirated after the incubation period was complete. The cells were fixed with 4% formaldehyde and washed once again. The membrane was permeabilized with 0.2% triton-x-100 and washed again. The cells were blocked for 30 minutes and incubated in primary antibody (either Cy5 or Actin) overnight. The antibody was then aspirated, washed, and incubated in appropriate secondary antibody for 30 minutes. The cells were then washed and DAPI solution was applied for 5 minutes. The cells were then washed and mounted using Prolong Gold Anti-Fade Reagent. The mounted cells were allowed to cure overnight and then imaged. The DAPI was imaged using a wavelength of 490 nm. The Cy5 and Actin were imaged using 594 nm wavelength.
[0216] Detection of protein from fixed cells was performed using Nikon 80i upright fluorescent microscope. A filter cube specific for the relative green range (.about.488 nm) was used to detect FITC labelled protein within cells.
[0217] The results of this experiment are shown in FIGS. 14, 15, 16, and 17. As shown in FIG. 14, incubation of cells with QE12-TAT-NleE resulted in nuclear fluorescence. As shown in FIGS. 15 and 16, both QE60-YopM PTD-NleE (GSGS linker) and QE60-YopM PTD-NleE (PAPA linker), respectively, produced punctate cytoplasmic fluorescence that was not observed in untreated control cells (FIG. 17). Taken together, these data show that the fusion polypeptides were taken up by individual cells.
[0218] Inverted microscopy. Uptake of QE12-TAT-NleE was analyzed by inverted microscopy as follows. A segment of skin was isolated from a shaved section of a mouse, and the tissue was preserved in a saline solution to maintain the cell viability within the tissue. The tissue was stained with 50 ug of FITC labeled TAT-NleE that was applied to the top of the sample and allowed to diffuse over a 2 hour period. Afterwards, the tissue was fixed with paraformaldehyde and stained with DAPI. The skin segment was then sliced in 10 uM increments, and the top slice was observed in an inverted microscope was used with varying objectives. As shown in FIG. 18, the TAT-NleE fusion polypeptide entered intact mouse skin and penetrated through multiple cell layers.
[0219] Two-photon microscopy. Penetration of TAT-NleE-into mouse skin was analyzed by two-photon microscopy. A 1 cm.times.1 cm segment of mouse skin was stained with 50 .mu.g of FITC-labeled TAT-NleE for two hours and counter stained with DAPI. the sample was analyzed by two-photon microscopy. More specifically, mouse hair was removed post-euthanasia and a 1 cm.times.1 cm segment of skin was removed and submerged in 1.times.PBS pH 7.4 to maintain tissue viability. Afterwards, the PBS was removed, and 50 ug FITC-TAT-NleE was applied to the top of the skin and allowed to incubate at room temperature for two hours. Afterwards, the tissue was washed three times for five minutes each with 1.times.PBS pH 7.4 to remove all waste. From there the tissue was permeabilized with Triton-x and fixed with 4% PFA. DAPI was then added to solution of PBS at a concentration of 1:1000 and incubated in the tissue for 3-5 minutes. Then the washing period was repeated (three times for five minutes each in 1 ml of PBS). The tissue was then mounted on a dish submerged in PBS ready for 2-photon imaging.
[0220] A 3 dimensional image is shown in FIG. 19 and a 10 uM slice of the stained tissue is shown in FIG. 20. As shown in FIGS. 19 and 20, the TAT-NleE penetrated the skin into the epidermis, but not the dermis.
Example 6: Methylase Activity
[0221] Methylase activity of recombinant proteins that included NleE effectors was analyzed using the MTase glo assay from Promega. We assayed the methylase activity of wild-type NleE, mutant NleE R107A and fusion proteins YopM PTD-NleE (GSGS linker), YopM PTD-NleE (PAPA linker). We also analyzed the methylase activity of NLeE from Shigella.
[0222] The in vitro methylase assay was developed using as a base, the MTase glo assay from Promega Inc. The primary assay buffer for enzyme activity was (25 mM Tris, pH 8, 50 mM NaCl, 1 mM EDTA, 3 mM MgCL2, 0.1% BSA, 0.005% Tween20, 5 mM DTT). To determine the effect of various vehicles and buffers on NLeE a 40 uL aliquot of vehicle was spiked with 1 uL of 20 mg/mL NLeE (0.5 mg/mL final). After .about.30 mins, 5 uL of spiked vehicle was diluted into 5000 uL of primary assay buffer--(termed `SPIKED` in graphs). To determine the effect of diluted vehicle on NLeE activity, 5 uL of vehicle diluted to 5000 uL of assay buffer was used. 1000 uL was then removed and used for the no enzyme controls. 1 uL of 0.5 mg/mL NLEE was then added to 1000 uL of diluted vehicle and the assay commenced. The methyltransferase assay itself was performed in 4.times.1:2 serial dilutions of each sample generated above in assay buffer and 2.5 uL added to 384-well assay plate. Reactions were initiated at various time points by the addition of 2.5 uL of 2.times. MTaseGlo, 1 uM GST-TAB2 protein substrate and 20 uM SAM in assay buffer. After the time course, reactions were terminated by the addition of 5 uL Methyltransferase Detection Reagent and the multiwell plates read for quantitative O.D. using a standard plate reader.
[0223] The results of this analysis are shown in FIGS. 21, 22 and 23. As shown in FIG. 21, both fusion constructs retained the methylase functionality of native NLeE. As shown in FIG. 22, the presence of an R107A mutation in NLeE abolished methylase activity. As shown in FIG. 23, the Shigella NleE showed methylase activity at levels comparable to that seen for the EPEC NleE
Example 7: Effect of Formulations on Nlee Methylation Activity
[0224] We analyzed the effective three different commercial formulations/delivery agents on NleE methylation activity in vitro. These were designated: Formula A, Formula B, and Formula C
[0225] The experimental setup was exactly as described above except that the buffer controls contained an equivalent amount of Formula A, Formula B or Formula C as controls. The results of this experiment are summarized in FIG. 24. As shown in FIG. 24a, NLeE methylase activity was retained in the presence of all three formulations. As shown in FIG. 24b, NLeE was stable in the presence of each of the formulations, although the degree of stability varied.
Example 8: How Specific is NleE for Inhibiting NFKB?
[0226] We cloned expressed and tested 64 other human C4 ZF containing proteins, among them the sequences identified in the table below. But few of them were modified by NleE. Thus, the specificity of NleE is extremely high.
TABLE-US-00008 CAN15_HUMAN/7-26 WsCvr..CtflNpagqrqCsiC CAN15_HUMAN/48-67 WpCar..CtfrNflgkeaCevC CAN15_HUMAN/147-166 WaCpr..CtlhNtpvassCsvC CAN15_HUMAN/344-363 WsCak..CtlrNptvaprCsaC CAN15_HUMAN/416-435 WaCpa..CtllNalrakhCaaC EWS_HUMAN/522-543 WqCpnpgCgnqNfawrteCnqC Oncogene RNA Binder FUS_HUMAN/426-447 WkCpnptCenmNfswrneCnqC Oncogene Annealing HOIL1_HUMAN/197-216 WqCpg..CtfiNkptrpgCemC Binds Ub Chains MDM2_HUMAN/303-322 WkCts..CnemNpplpshCnrC P53 regulation MDM4_HUMAN/304-323 WqCte..CkkfNspskryCfrC P53 regulation NEIL3_HUMAN/321-340 WtCvv..CtliNkpsskaCdaC Endonuclease8 Repair NPL4_HUMAN/584-603 WaCqh..CtfmNqpgtghCemC Ub Chaperone Erad NRP1_YEAST/359-378 WnCps..CgfsNfqrrtaCfrC Rna Binder No Fcn NRP1_YEAST/585-604 WkCst..CtyhNfaknvvClrC NU153_HUMAN/662-681 WqCdt..CllqNkvtdnkCiaC NUP153 Nuc Pore NU153_HUMAN/726-745 WdCdt..ClvqNkpeaikCvaC NU153_HUMAN/797-816 WeCsv..CcvsNnaednkCvsC NU153_HUMAN/855-874 WdCel..ClvqNkadstkClaC RBM10_HUMAN/217-236 WlCnk..CgvqNfkrrekCfkC RNA Binding RBM5_HUMAN/185-204 WlCnk..CclnNfrkrlkCfrC RNA Binding RBP2_HUMAN/1356-1375 WhCns..CslkNastakkCvsC RANBP2 Huge Nuc Pore RBP2_HUMAN/1419-1438 WdCsi..ClvrNeptvsrCiaC RBP2_HUMAN/1483-1502 WdCsa..ClvqNegsstkCaaC RBP2_HUMAN/1547-1566 WdCss..ClvrNeanatrCvaC RBP2_HUMAN/1610-1629 WdCsv..ClvrNeasatkCiaC RBP2_HUMAN/1669-1688 WdCsv..ClvrNeasatkCiaC RBP2_HUMAN/1728-1747 WdCsv..ClvrNeasatkCiaC RBP2_HUMAN/1785-1804 WdCsv..CcvqNessslkCvaC RBP56 HUMAN/358-379 WvCpnpsCgnmNfarrnsCnqC RNF31 HUMAN/304-323 WhCaa..CamlNepwavlCvaC RNF31 HUMAN/354-373 WaCqs..CtfeNeaaavlCsiC RNF31 HUMAN/413-432 WyCih..CtfcNsspgwvCvmC RYBP_HUMAN/25-44 WdCsv..CtfrNsaeafkCsiC SHRPN_HUMAN/352-371 WsCps..CtfiNapdrpgCemC TAB2_HUMAN/668-687 WnCta..CtflNhpalirCeqC NFKB pathway TAB3_HUMAN/687-706 WnCds..CtflNhpalnrCeqC NFKB pathway TX13A_HUMAN/380-399 WdCpw..CnavNfsrrdtCfdC
[0227] A cysteine methylation assay was performed and shows that NleE does not utilize zinc fingers of known highly related zinc fingers. NleE targets NFKB signaling at a key step in TAK1 kinase activation. The CYS-4 Zinc Finger of TAB2 was required to bind free K63 Ub chains to activate TAK1 kinase, which thereby triggers NFKB signaling. The TAB2/3 Cys 4 Zinc-Finger binds K63 linked Ub chains: The metal chelated structure of the ZF required binding. NleE targets NFKB signaling by methylating a single Cysteine in the C4 ZF TAB2 of TAB2 abolishing K63 Ub sensing. Cys 673 in TAB2 is the target of NleE. This highly specific activity of NleE is repurposed in the compositions described herein to shut down the NFKB pathway and reduce inflammation.
Example 9: Penetration of Recombinant NleE into the Skin in an Ex Vivo Pig Skin Model
[0228] A key claim and potential stumbling block to use of NleE as a skin anti-inflammatory compound is to show that this globular folded protein of MW 38 kDa is able to penetrate the dermis and epidermis in order to access cells in which the NFKB system is aberrantly activated. A recombinant construct was prepared by fusing the amino acid sequence of NleE to an N-terminal histidine, and covalently labelling that peptide with FITC dye, resulting in the peptide His-NleE--FITC.
[0229] This construct was mixed with the eutectic, cholinate-based solvent CAGE, as described above at a concentration of 1-10 mg/ml. The amount of peptide in CAGE or PBS was 1 mg/ml. The final CAGE concentration was 90% v/v. The mixture was applied to live pig skin being perfused in a Franz diffusion chamber at 37 degrees C. for 24 or 48 hours. The Franz diffusion chamber is described by Technical Brief 2009 vol. 10, Particle Sciences--Drug Development Services (Bethlehem, Pa.). After these time periods, skin was harvested, fixed and sectioned followed by microscopy to visualize the FITC dye. The results (FIG. 4) show that CAGE-solubilized NleE FITC was able penetrate the stratum corneum, epidermis and dermis and that virtually every cellular compartment of the skin. Up to 20-30 cell layers were contacted.
[0230] In comparison to published reports by Banerjee et. al 2017, particularly FIG. 1 using CAGE to transport the globular proteins ovalbumin and bovine serum albumin, the NleE penetrates as well if not better than these test proteins. NleE solubilized in PBS and applied to the pig skin showed no transduction past the stratum corneum.
[0231] In summary, this pre-clinical data provides support that recombinant NleE protein is stable in the eutectic solvent CAGE and can be delivered to multi layers of the skin locally in a topical manner. It also provides support for using NleE protein and the combinations described herein that contain an intrinsic PTD domain which renders the protein capable of penetrating cells and tissues and turning off inflammatory cytokines. The ability of these combinations of proteins to penetrate, in an apparently active form and inhibit production of a multitude of inflammatory cytokines, is reflected in its therapeutic effect on psoriatic lesions in a mouse skin model. These compositions delivered in a cream topically, in therapeutic levels are not anticipated to appreciably enter the regional lymphatics.
[0232] These observations strongly support the use of recombinant bacterial effector proteins containing PTDs (Table1) are useful as a platform for inhibiting various accessible inflammatory disease processes in the body.
[0233] Thus, methods and compositions for treating inflammation related to activation of the NFkB, JNK and p38 pathways, e.g., inflammatory conditions of skin, include a composition comprising a first construct comprising a selected immunomodulatory effector protein or functional equivalent thereof that targets a first functional domain; and at least one additional construct comprising a different effector protein or a functional equivalent thereof that targets an additional functional domain. The first construct is optionally linked to a selected protein transduction domain (PTD) or penetrating peptide (CPP) or to a targeting moiety. The additional construct is optionally linked to the same PTD or CPP of the first construct or to an additional PTD or CPP. In one embodiment, a combination of these constructs is provided. In another embodiment the first construct is fused to one or more additional constructs in a single polypeptide. The composition further comprise an optional linker amino acid sequence interposed between each construct and an optional targeting moiety.
[0234] Each and every patent, patent application, and publication, including websites cited throughout specification are incorporated herein by reference. Similarly, the SEQ ID NOs which are reference herein and which appear in the appended Sequence Listing are incorporated by reference. While the invention has been described with reference to particular embodiments, it will be appreciated that modifications can be made without departing from the spirit of the invention. Such modifications are intended to fall within the scope of the appended claims.
Sequence CWU 1
1
931224PRTUnknownDescription of Unknown NleE sequence 1Met Ile Asn
Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr1 5 10 15Lys Tyr
Ala Glu His Val Val Lys Asn Ile Tyr Pro Glu Ile Lys His 20 25 30Asp
Tyr Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys Lys Tyr Ile Ser 35 40
45Gly Ile Thr Arg Gly Val Ala Glu Leu Lys Gln Glu Glu Phe Val Asn
50 55 60Glu Lys Ala Arg Arg Phe Ser Tyr Met Lys Thr Met Tyr Ser Val
Cys65 70 75 80Pro Glu Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser
Thr Pro Glu 85 90 95Gly Ser Trp Leu Thr Val Ile Ser Gly Lys Arg Pro
Met Gly Gln Phe 100 105 110Ser Val Asp Ser Leu Tyr Asn Pro Asp Leu
His Ala Leu Cys Glu Leu 115 120 125Pro Asp Ile Cys Cys Lys Ile Phe
Pro Lys Glu Asn Asn Asp Phe Leu 130 135 140Tyr Ile Val Val Val Tyr
Arg Asn Asp Ser Pro Leu Gly Glu Gln Arg145 150 155 160Ala Asn Arg
Phe Ile Glu Leu Tyr Asn Ile Lys Arg Asp Ile Met Gln 165 170 175Glu
Leu Asn Tyr Glu Leu Pro Glu Leu Lys Ala Val Lys Ser Glu Met 180 185
190Ile Ile Ala Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro Gly Glu
195 200 205Ile Asp Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys
Ile Glu 210 215 2202409PRTYersinia pestis 2Met Phe Ile Asn Pro Arg
Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1 5 10 15Leu Arg His Ser Ser
Asn Leu Thr Glu Met Pro Val Glu Ala Glu Asn 20 25 30Val Lys Ser Lys
Thr Glu Tyr Tyr Asn Ala Trp Ser Glu Trp Glu Arg 35 40 45Asn Ala Pro
Pro Gly Asn Gly Glu Gln Arg Glu Met Ala Val Ser Arg 50 55 60Leu Arg
Asp Cys Leu Asp Arg Gln Ala His Glu Leu Glu Leu Asn Asn65 70 75
80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro Pro His Leu Glu Ser Leu
85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu Leu Pro Glu Leu Pro Gln
Ser 100 105 110Leu Lys Ser Leu Leu Val Asp Asn Asn Asn Leu Lys Ala
Leu Ser Asp 115 120 125Leu Pro Pro Leu Leu Glu Tyr Leu Gly Val Ser
Asn Asn Gln Leu Glu 130 135 140Lys Leu Pro Glu Leu Gln Asn Ser Ser
Phe Leu Lys Ile Ile Asp Val145 150 155 160Asp Asn Asn Ser Leu Lys
Lys Leu Pro Asp Leu Pro Pro Ser Leu Glu 165 170 175Phe Ile Ala Ala
Gly Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 180 185 190Asn Leu
Pro Phe Leu Thr Ala Ile Tyr Ala Asp Asn Asn Ser Leu Lys 195 200
205Lys Leu Pro Asp Leu Pro Leu Ser Leu Glu Ser Ile Val Ala Gly Asn
210 215 220Asn Ile Leu Glu Glu Leu Pro Glu Leu Gln Asn Leu Pro Phe
Leu Thr225 230 235 240Thr Ile Tyr Ala Asp Asn Asn Leu Leu Lys Thr
Leu Pro Asp Leu Pro 245 250 255Pro Ser Leu Glu Ala Leu Asn Val Arg
Asp Asn Tyr Leu Thr Asp Leu 260 265 270Pro Glu Leu Pro Gln Ser Leu
Thr Phe Leu Asp Val Ser Glu Asn Ile 275 280 285Phe Ser Gly Leu Ser
Glu Leu Pro Pro Asn Leu Tyr Tyr Leu Asn Ala 290 295 300Ser Ser Asn
Glu Ile Arg Ser Leu Cys Asp Leu Pro Pro Ser Leu Glu305 310 315
320Glu Leu Asn Val Ser Asn Asn Lys Leu Ile Glu Leu Pro Ala Leu Pro
325 330 335Pro Arg Leu Glu Arg Leu Ile Ala Ser Phe Asn His Leu Ala
Glu Val 340 345 350Pro Glu Leu Pro Gln Asn Leu Lys Gln Leu His Val
Glu Tyr Asn Pro 355 360 365Leu Arg Glu Phe Pro Asp Ile Pro Glu Ser
Val Glu Asp Leu Arg Met 370 375 380Asn Ser Glu Arg Val Val Asp Pro
Tyr Glu Phe Ala His Glu Thr Thr385 390 395 400Asp Lys Leu Glu Asp
Asp Val Phe Glu 4053367PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 3Met Phe Ile Thr Pro Arg
Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1 5 10 15Leu Arg His Ser Ser
Asp Leu Thr Glu Met Pro Val Glu Ala Glu Asn 20 25 30Val Lys Ser Lys
Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp Glu Arg 35 40 45Asn Ala Pro
Pro Gly Asn Gly Glu Gln Arg Glu Met Ala Val Ser Arg 50 55 60Leu Arg
Asp Cys Leu Asp Arg Gln Ala His Glu Leu Glu Leu Asn Asn65 70 75
80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro Pro His Leu Glu Ser Leu
85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu Leu Pro Glu Leu Pro Gln
Ser 100 105 110Leu Lys Ser Leu Gln Val Asp Asn Asn Asn Leu Lys Ala
Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu Glu Phe Leu Ala Ala Gly
Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro Glu Leu Gln Asn Ser Ser
Phe Leu Lys Ile Ile Asp Val145 150 155 160Asp Asn Asn Ser Leu Lys
Lys Leu Pro Asp Leu Pro Pro Ser Leu Glu 165 170 175Phe Leu Ala Ala
Gly Asn Asn Gln Leu Glu Glu Leu Ser Glu Leu Gln 180 185 190Asn Leu
Pro Phe Leu Thr Glu Ile His Ala Asp Asn Asn Ser Leu Lys 195 200
205Thr Leu Pro Asp Leu Pro Pro Ser Leu Lys Thr Leu Asn Val Arg Glu
210 215 220Asn Tyr Leu Thr Asp Leu Pro Glu Leu Pro Gln Ser Leu Thr
Phe Leu225 230 235 240Asp Val Ser Asp Asn Ile Phe Ser Gly Leu Ser
Glu Leu Pro Pro Asn 245 250 255Leu Tyr Tyr Leu Asp Ala Ser Ser Asn
Gly Ile Arg Ser Leu Cys Asp 260 265 270Leu Pro Pro Ser Leu Val Glu
Leu Asp Val Arg Asp Asn Gln Leu Ile 275 280 285Glu Leu Pro Ala Leu
Pro Pro His Leu Glu Arg Leu Ile Ala Ser Leu 290 295 300Asn His Leu
Ala Glu Val Pro Glu Leu Pro Gln Asn Leu Lys Gln Leu305 310 315
320His Val Glu His Asn Ala Leu Arg Glu Phe Pro Asp Ile Pro Glu Ser
325 330 335Val Glu Asp Leu Arg Met Asp Ser Glu Arg Val Thr Asp Thr
Tyr Glu 340 345 350Phe Ala His Glu Thr Thr Asp Lys Leu Glu Asp Asp
Val Phe Glu 355 360 36541104DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 4atgtttataa ctccaagaaa
tgtatctaat acttttttgc aagaaccatt acgtcattct 60tctgatttaa ctgagatgcc
ggttgaagca gaaaatgtta aatctaagac tgaatattat 120aatgcatggg
cggtatggga acgaaatgcc cctccgggga atggtgaaca gagggaaatg
180gcggtttcaa ggttacgcga ttgcctggac cgacaagccc atgagctaga
actaaataat 240ctggggctga gttctttgcc ggaattacct ccgcatttag
agagtttagt ggcgtcatgt 300aattctctta cagaattacc ggaattgccg
cagagcctga aatcacttca agttgataat 360aacaatctga aggcattatc
cgatttacct ccttcactgg aatttcttgc tgctggtaat 420aatcagctgg
aagaattgcc agagttgcaa aactcgtcct tcttgaaaat tattgatgtt
480gataacaatt cactgaaaaa actacctgat ttacctcctt cactggaatt
tcttgctgct 540ggtaataatc agctggaaga attgtcagag ttacaaaact
tgccattctt gactgagatt 600catgctgata acaattcact gaaaacatta
cccgatttac ccccttccct gaaaacactt 660aatgtcagag aaaattattt
aactgatctg ccagaattac cgcagagttt aaccttctta 720gatgtttctg
ataatatttt ttctggatta tcggaattgc caccaaactt gtattatctc
780gatgcatcca gcaatggaat aagatcctta tgcgatttac ccccttcact
ggtagaactt 840gatgtcagag ataatcagtt gatcgaactg ccagcgttac
ctccacactt agaacgttta 900atcgcttcac ttaatcatct tgctgaagta
cctgaattgc cgcaaaacct gaaacagctc 960cacgtagagc acaacgctct
aagagagttt cccgatatac ctgagtcagt ggaagatctt 1020cggatggact
ctgaacgtgt aactgataca tatgaatttg ctcatgagac tacagacaaa
1080cttgaagatg atgtatttga gtag 1104586PRTYersinia pestis 5Met Phe
Ile Asn Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1 5 10 15Leu
Arg His Ser Ser Asn Leu Thr Glu Met Pro Val Glu Ala Glu Asn 20 25
30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ser Glu Trp Glu Arg
35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala Val Ser
Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu Glu Leu
Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu 856258DNAYersinia pestis
6atgttcatca ctccacgcaa tgtatctaac acctttctgc aggaaccgct gcgtcattct
60agcgacctga ccgaaatgcc agttgaagcg gagaacgtga aatctaagac tgaatactac
120aacgcgtggg cagtatggga gcgcaatgca ccaccaggta acggtgaaca
gcgtgaaatg 180gcagtaagcc gtctgcgtga ttgcctggat cgccaggctc
acgagctgga gctgaacaac 240ctgggtctgt ctagcctg 2587131PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
7Met Phe Ile Asn Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1 5
10 15Leu Arg His Ser Ser Asn Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ser Glu Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Leu Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro
1308393DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 8atgtttatta acccgcgcaa cgtgagcaac
acctttctgc aggaaccgct gcgccatagc 60agcaacctga ccgaaatgcc ggtggaagcg
gaaaacgtga aaagcaaaac cgaatattat 120aacgcgtgga gcgaatggga
acgcaacgcg ccgccgggca acggcgaaca gcgcgaaatg 180gcggtgagcc
gcctgcgcga ttgcctggat cgccaggcgc atgaactgga actgaacaac
240ctgggcctga gcagcctgcc ggaactgccg ccgcatctgg aaagcctggt
ggcgagctgc 300aacagcctga ccgaactgcc ggaactgccg cagagcctga
aaagcctgct ggtggataac 360aacaacctga aagcgctgag cgatctgccg ccg
3939390PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 9Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr
Phe Leu Gln Glu Pro1 5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met
Pro Val Glu Ala Glu Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn
Ala Trp Ala Val Trp Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu
Gln Arg Glu Met Ala Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg
Gln Ala His Glu Leu Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser
Leu Pro Glu Leu Pro Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys
Asn Ser Leu Thr Glu Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys
Ser Leu Gln Val Asp Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120
125Leu Pro Pro Ser Leu Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu
130 135 140Glu Leu Pro Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile
Asp Ile145 150 155 160Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro
Ile Asn Thr Lys Tyr 165 170 175Ala Glu His Val Val Lys Asn Ile Tyr
Pro Lys Ile Lys His Asp Tyr 180 185 190Phe Asn Glu Ser Pro Asn Ile
Tyr Asp Lys Lys Tyr Ile Ser Gly Ile 195 200 205Thr Arg Gly Val Ala
Glu Leu Lys Gln Glu Glu Phe Val Asn Glu Lys 210 215 220Ala Arg Arg
Phe Ser Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu225 230 235
240Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser
245 250 255Trp Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln Phe
Ser Val 260 265 270Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys
Glu Leu Pro Asp 275 280 285Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn
Asn Asp Phe Leu Tyr Ile 290 295 300Val Val Val Tyr Arg Asn Asp Ser
Pro Leu Gly Glu Gln Arg Ala Asn305 310 315 320Arg Phe Ile Glu Leu
Tyr Asn Ile Lys Arg Asp Ile Met Gln Glu Leu 325 330 335Asn Tyr Glu
Leu Pro Glu Leu Lys Ala Val Lys Ser Glu Met Ile Ile 340 345 350Ala
Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 355 360
365Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser
370 375 380His His His His His His385 39010384PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
10Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Ile145 150 155
160Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr
165 170 175Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile Lys His
Asp Tyr 180 185 190Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys Lys Tyr
Ile Ser Gly Ile 195 200 205Thr Arg Gly Val Ala Glu Leu Lys Gln Glu
Glu Phe Val Asn Glu Lys 210 215 220Ala Arg Arg Phe Ser Tyr Met Lys
Thr Met Tyr Ser Val Cys Pro Glu225 230 235 240Ala Phe Glu Pro Ile
Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 245 250 255Trp Leu Thr
Val Ile Ser Gly Lys Arg Pro Met Gly Gln Phe Ser Val 260 265 270Asp
Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp 275 280
285Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile
290 295 300Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly Glu Gln Arg
Ala Asn305 310 315 320Arg Phe Ile Glu Leu Tyr Asn Ile Lys Arg Asp
Ile Met Gln Glu Leu 325 330 335Asn Tyr Glu Leu Pro Glu Leu Lys Ala
Val Lys Ser Glu Met Ile Ile 340 345 350Ala Arg Glu Met Gly Glu Ile
Phe Ser Tyr Met Pro Gly Glu Ile Asp 355 360 365Ser Tyr Met Lys Tyr
Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser 370 375
380111173DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 11atgttcatca ctccacgcaa tgtatctaac
acctttctgc aggaaccgct gcgtcattct 60agcgacctga ccgaaatgcc agttgaagcg
gagaacgtga aatctaagac tgaatactac 120aacgcgtggg cagtatggga
gcgcaatgca ccaccaggta acggtgaaca gcgtgaaatg 180gcagtaagcc
gtctgcgtga ttgcctggat cgccaggctc acgagctgga gctgaacaac
240ctgggtctgt ctagcctgcc agagctccca ccacatctgg aaagcctggt
ggctagctgt 300aactctctga ctgaactgcc agagctgcca caaagcctga
aatccctgca ggtggacaac 360aacaacctga aagcgctgtc cgatctgcca
ccgtctctgg agtttctggc agctggtaac 420aaccaactgg aagaactgcc
agagctgcag aactcctcct ttctgaagat catcgatatt 480aatcctgtta
ctaatactca gggcgtgtcc cctataaata ctaaatatgc tgaacatgtg
540gtgaaaaata tttacccgaa aattaaacat gattacttta atgaatcacc
caatatatat 600gataagaagt atatatccgg tataaccaga ggagtagctg
aactaaaaca ggaagaattt 660gttaacgaga aagccagacg gttttcttat
atgaagacta tgtattctgt atgtccagaa 720gcgtttgaac ctatttccag
aaatgaagcc agtacaccgg aaggaagctg gctaacagtt 780atatccggaa
aagccccaat ggggcagttt tctgtagata gtttatacaa tcctgattta
840catgcattat gtgagcttcc ggacatttgt tgtaagatct tccctaaaga
aaataatgat 900tttttataca tagttgttgt gtacagaaat gacagccctc
taggagaaca acgggcaaat 960agatttatag aattatataa tataaaaaga
gatatcatgc aggaattaaa ttatgagtta 1020ccagagttaa aggcagtaaa
atctgaaatg attatcgcac gtgaaatggg agaaatcttt 1080agctacatgc
ctggggaaat agacagttat atgaaataca taaataataa actttctaaa
1140attgagggat ctcatcacca tcaccatcac taa 117312406PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
12Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Pro145 150 155
160Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400His His His His His His 40513400PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
13Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Pro145 150 155
160Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400141221DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 14atgttcatca ctccacgcaa tgtatctaac
acctttctgc aggaaccgct gcgtcattct 60agcgacctga ccgaaatgcc agttgaagcg
gagaacgtga aatctaagac tgaatactac 120aacgcgtggg cagtatggga
gcgcaatgca ccaccaggta acggtgaaca gcgtgaaatg 180gcagtaagcc
gtctgcgtga ttgcctggat cgccaggctc acgagctgga gctgaacaac
240ctgggtctgt ctagcctgcc agagctccca ccacatctgg aaagcctggt
ggctagctgt 300aactctctga ctgaactgcc agagctgcca caaagcctga
aatccctgca ggtggacaac 360aacaacctga aagcgctgtc cgatctgcca
ccgtctctgg agtttctggc agctggtaac 420aaccaactgg aagaactgcc
agagctgcag aactcctcct ttctgaagat catcgatcca 480gcaccagcac
cagcaccagc accagcacca gcaccagcac cagcaattaa tcctgttact
540aatactcagg gcgtgtcccc tataaatact aaatatgctg aacatgtggt
gaaaaatatt 600tacccgaaaa ttaaacatga ttactttaat gaatcaccca
atatatatga taagaagtat 660atatccggta taaccagagg agtagctgaa
ctaaaacagg aagaatttgt taacgagaaa 720gccagacggt tttcttatat
gaagactatg tattctgtat gtccagaagc gtttgaacct 780atttccagaa
atgaagccag tacaccggaa ggaagctggc taacagttat atccggaaaa
840cgcccaatgg ggcagttttc tgtagatagt ttatacaatc ctgatttaca
tgcattatgt 900gagcttccgg acatttgttg taagatcttc cctaaagaaa
ataatgattt tttatacata 960gttgttgtgt acagaaatga cagccctcta
ggagaacaac gggcaaatag atttatagaa 1020ttatataata taaaaagaga
tatcatgcag gaattaaatt atgagttacc agagttaaag 1080gcagtaaaat
ctgaaatgat tatcgcacgt gaaatgggag aaatctttag ctacatgcct
1140ggggaaatag acagttatat gaaatacata aataataaac tttctaaaat
tgagggatct 1200catcaccatc accatcacta a 122115406PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
15Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Gly145 150 155
160Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400His His His His His His 40516400PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
16Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Gly145 150 155
160Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400171221DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 17atgttcatca ctccacgcaa tgtatctaac
acctttctgc aggaaccgct gcgtcattct 60agcgacctga ccgaaatgcc agttgaagcg
gagaacgtga aatctaagac tgaatactac 120aacgcgtggg cagtatggga
gcgcaatgca ccaccaggta acggtgaaca gcgtgaaatg 180gcagtaagcc
gtctgcgtga ttgcctggat cgccaggctc acgagctgga gctgaacaac
240ctgggtctgt ctagcctgcc agagctccca ccacatctgg aaagcctggt
ggctagctgt 300aactctctga ctgaactgcc agagctgcca caaagcctga
aatccctgca ggtggacaac 360aacaacctga aagcgctgtc cgatctgcca
ccgtctctgg agtttctggc agctggtaac 420aaccaactgg aagaactgcc
agagctgcag aactcctcct ttctgaagat catcgatggt 480agcggtagcg
gtagcggtag cggtagcggt agcggtagcg gtagcattaa tcctgttact
540aatactcagg gcgtgtcccc tataaatact aaatatgctg aacatgtggt
gaaaaatatt 600tacccgaaaa ttaaacatga ttactttaat gaatcaccca
atatatatga taagaagtat 660atatccggta taaccagagg agtagctgaa
ctaaaacagg aagaatttgt taacgagaaa 720gccagacggt tttcttatat
gaagactatg tattctgtat gtccagaagc gtttgaacct 780atttccagaa
atgaagccag tacaccggaa ggaagctggc taacagttat atccggaaaa
840gccccaatgg ggcagttttc tgtagatagt ttatacaatc ctgatttaca
tgcattatgt 900gagcttccgg acatttgttg taagatcttc cctaaagaaa
ataatgattt tttatacata 960gttgttgtgt acagaaatga cagccctcta
ggagaacaac gggcaaatag atttatagaa 1020ttatataata taaaaagaga
tatcatgcag gaattaaatt atgagttacc agagttaaag 1080gcagtaaaat
ctgaaatgat tatcgcacgt gaaatgggag aaatctttag ctacatgcct
1140ggggaaatag acagttatat gaaatacata aataataaac tttctaaaat
tgagggatct 1200catcaccatc accatcacta a 122118390PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
18Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Ile145 150 155
160Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr
165 170 175Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile Lys His
Asp Tyr 180 185 190Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys Lys Tyr
Ile Ser Gly Ile 195 200 205Thr Arg Gly Val Ala Glu Leu Lys Gln Glu
Glu Phe Val Asn Glu Lys 210 215 220Ala Arg Arg Phe Ser Tyr Met Lys
Thr Met Tyr Ser Val Cys Pro Glu225 230 235
240Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser
245 250 255Trp Leu Thr Val Ile Ser Gly Lys Ala Pro Met Gly Gln Phe
Ser Val 260 265 270Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys
Glu Leu Pro Asp 275 280 285Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn
Asn Asp Phe Leu Tyr Ile 290 295 300Val Val Val Tyr Arg Asn Asp Ser
Pro Leu Gly Glu Gln Arg Ala Asn305 310 315 320Arg Phe Ile Glu Leu
Tyr Asn Ile Lys Arg Asp Ile Met Gln Glu Leu 325 330 335Asn Tyr Glu
Leu Pro Glu Leu Lys Ala Val Lys Ser Glu Met Ile Ile 340 345 350Ala
Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 355 360
365Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser
370 375 380His His His His His His385 39019384PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
19Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Ile145 150 155
160Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr
165 170 175Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile Lys His
Asp Tyr 180 185 190Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys Lys Tyr
Ile Ser Gly Ile 195 200 205Thr Arg Gly Val Ala Glu Leu Lys Gln Glu
Glu Phe Val Asn Glu Lys 210 215 220Ala Arg Arg Phe Ser Tyr Met Lys
Thr Met Tyr Ser Val Cys Pro Glu225 230 235 240Ala Phe Glu Pro Ile
Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 245 250 255Trp Leu Thr
Val Ile Ser Gly Lys Ala Pro Met Gly Gln Phe Ser Val 260 265 270Asp
Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp 275 280
285Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile
290 295 300Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly Glu Gln Arg
Ala Asn305 310 315 320Arg Phe Ile Glu Leu Tyr Asn Ile Lys Arg Asp
Ile Met Gln Glu Leu 325 330 335Asn Tyr Glu Leu Pro Glu Leu Lys Ala
Val Lys Ser Glu Met Ile Ile 340 345 350Ala Arg Glu Met Gly Glu Ile
Phe Ser Tyr Met Pro Gly Glu Ile Asp 355 360 365Ser Tyr Met Lys Tyr
Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser 370 375
380201173DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 20atgttcatca ctccacgcaa tgtatctaac
acctttctgc aggaaccgct gcgtcattct 60agcgacctga ccgaaatgcc agttgaagcg
gagaacgtga aatctaagac tgaatactac 120aacgcgtggg cagtatggga
gcgcaatgca ccaccaggta acggtgaaca gcgtgaaatg 180gcagtaagcc
gtctgcgtga ttgcctggat cgccaggctc acgagctgga gctgaacaac
240ctgggtctgt ctagcctgcc agagctccca ccacatctgg aaagcctggt
ggctagctgt 300aactctctga ctgaactgcc agagctgcca caaagcctga
aatccctgca ggtggacaac 360aacaacctga aagcgctgtc cgatctgcca
ccgtctctgg agtttctggc agctggtaac 420aaccaactgg aagaactgcc
agagctgcag aactcctcct ttctgaagat catcgatatt 480aatcctgtta
ctaatactca gggcgtgtcc cctataaata ctaaatatgc tgaacatgtg
540gtgaaaaata tttacccgaa aattaaacat gattacttta atgaatcacc
caatatatat 600gataagaagt atatatccgg tataaccaga ggagtagctg
aactaaaaca ggaagaattt 660gttaacgaga aagccagacg gttttcttat
atgaagacta tgtattctgt atgtccagaa 720gcgtttgaac ctatttccag
aaatgaagcc agtacaccgg aaggaagctg gctaacagtt 780atatccggaa
aagccccaat ggggcagttt tctgtagata gtttatacaa tcctgattta
840catgcattat gtgagcttcc ggacatttgt tgtaagatct tccctaaaga
aaataatgat 900tttttataca tagttgttgt gtacagaaat gacagccctc
taggagaaca acgggcaaat 960agatttatag aattatataa tataaaaaga
gatatcatgc aggaattaaa ttatgagtta 1020ccagagttaa aggcagtaaa
atctgaaatg attatcgcac gtgaaatggg agaaatcttt 1080agctacatgc
ctggggaaat agacagttat atgaaataca taaataataa actttctaaa
1140attgagggat ctcatcacca tcaccatcac taa 117321406PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
21Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Gly145 150 155
160Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Ala Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400His His His His His His 40522400PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
22Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Gly145 150 155
160Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Gly Ser Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Ala Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400231221DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 23atgttcatca ctccacgcaa tgtatctaac
acctttctgc aggaaccgct gcgtcattct 60agcgacctga ccgaaatgcc agttgaagcg
gagaacgtga aatctaagac tgaatactac 120aacgcgtggg cagtatggga
gcgcaatgca ccaccaggta acggtgaaca gcgtgaaatg 180gcagtaagcc
gtctgcgtga ttgcctggat cgccaggctc acgagctgga gctgaacaac
240ctgggtctgt ctagcctgcc agagctccca ccacatctgg aaagcctggt
ggctagctgt 300aactctctga ctgaactgcc agagctgcca caaagcctga
aatccctgca ggtggacaac 360aacaacctga aagcgctgtc cgatctgcca
ccgtctctgg agtttctggc agctggtaac 420aaccaactgg aagaactgcc
agagctgcag aactcctcct ttctgaagat catcgatggt 480agcggtagcg
gtagcggtag cggtagcggt agcggtagcg gtagcattaa tcctgttact
540aatactcagg gcgtgtcccc tataaatact aaatatgctg aacatgtggt
gaaaaatatt 600tacccgaaaa ttaaacatga ttactttaat gaatcaccca
atatatatga taagaagtat 660atatccggta taaccagagg agtagctgaa
ctaaaacagg aagaatttgt taacgagaaa 720gccagacggt tttcttatat
gaagactatg tattctgtat gtccagaagc gtttgaacct 780atttccagaa
atgaagccag tacaccggaa ggaagctggc taacagttat atccggaaaa
840gccccaatgg ggcagttttc tgtagatagt ttatacaatc ctgatttaca
tgcattatgt 900gagcttccgg acatttgttg taagatcttc cctaaagaaa
ataatgattt tttatacata 960gttgttgtgt acagaaatga cagccctcta
ggagaacaac gggcaaatag atttatagaa 1020ttatataata taaaaagaga
tatcatgcag gaattaaatt atgagttacc agagttaaag 1080gcagtaaaat
ctgaaatgat tatcgcacgt gaaatgggag aaatctttag ctacatgcct
1140ggggaaatag acagttatat gaaatacata aataataaac tttctaaaat
tgagggatct 1200catcaccatc accatcacta a 122124406PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
24Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Pro145 150 155
160Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225 230 235 240Ala Arg Arg Phe Ser
Tyr Met Lys Thr Met Tyr Ser Val Cys Pro Glu 245 250 255Ala Phe Glu
Pro Ile Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 260 265 270Trp
Leu Thr Val Ile Ser Gly Lys Ala Pro Met Gly Gln Phe Ser Val 275 280
285Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp
290 295 300Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile305 310 315 320Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn 325 330 335Arg Phe Ile Glu Leu Tyr Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu 340 345 350Asn Tyr Glu Leu Pro Glu Leu
Lys Ala Val Lys Ser Glu Met Ile Ile 355 360 365Ala Arg Glu Met Gly
Glu Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp 370 375 380Ser Tyr Met
Lys Tyr Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser385 390 395
400His His His His His His 40525400PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
25Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln Glu Pro1
5 10 15Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu Ala Glu
Asn 20 25 30Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala Val Trp
Glu Arg 35 40 45Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu Met Ala
Val Ser Arg 50 55 60Leu Arg Asp Cys Leu Asp Arg Gln Ala His Glu Leu
Glu Leu Asn Asn65 70 75 80Leu Gly Leu Ser Ser Leu Pro Glu Leu Pro
Pro His Leu Glu Ser Leu 85 90 95Val Ala Ser Cys Asn Ser Leu Thr Glu
Leu Pro Glu Leu Pro Gln Ser 100 105 110Leu Lys Ser Leu Gln Val Asp
Asn Asn Asn Leu Lys Ala Leu Ser Asp 115 120 125Leu Pro Pro Ser Leu
Glu Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu 130 135 140Glu Leu Pro
Glu Leu Gln Asn Ser Ser Phe Leu Lys Ile Ile Asp Pro145 150 155
160Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Ile
165 170 175Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro Ile Asn Thr
Lys Tyr 180 185 190Ala Glu His Val Val Lys Asn Ile Tyr Pro Lys Ile
Lys His Asp Tyr 195 200 205Phe Asn Glu Ser Pro Asn Ile Tyr Asp Lys
Lys Tyr Ile Ser Gly Ile 210 215 220Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys225
230 235 240Ala Arg Arg Phe Ser Tyr Met Lys Thr Met Tyr Ser Val Cys
Pro Glu 245 250 255Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr
Pro Glu Gly Ser 260 265 270Trp Leu Thr Val Ile Ser Gly Lys Ala Pro
Met Gly Gln Phe Ser Val 275 280 285Asp Ser Leu Tyr Asn Pro Asp Leu
His Ala Leu Cys Glu Leu Pro Asp 290 295 300Ile Cys Cys Lys Ile Phe
Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile305 310 315 320Val Val Val
Tyr Arg Asn Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn 325 330 335Arg
Phe Ile Glu Leu Tyr Asn Ile Lys Arg Asp Ile Met Gln Glu Leu 340 345
350Asn Tyr Glu Leu Pro Glu Leu Lys Ala Val Lys Ser Glu Met Ile Ile
355 360 365Ala Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro Gly Glu
Ile Asp 370 375 380Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys
Ile Glu Gly Ser385 390 395 400261221DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
26atgttcatca ctccacgcaa tgtatctaac acctttctgc aggaaccgct gcgtcattct
60agcgacctga ccgaaatgcc agttgaagcg gagaacgtga aatctaagac tgaatactac
120aacgcgtggg cagtatggga gcgcaatgca ccaccaggta acggtgaaca
gcgtgaaatg 180gcagtaagcc gtctgcgtga ttgcctggat cgccaggctc
acgagctgga gctgaacaac 240ctgggtctgt ctagcctgcc agagctccca
ccacatctgg aaagcctggt ggctagctgt 300aactctctga ctgaactgcc
agagctgcca caaagcctga aatccctgca ggtggacaac 360aacaacctga
aagcgctgtc cgatctgcca ccgtctctgg agtttctggc agctggtaac
420aaccaactgg aagaactgcc agagctgcag aactcctcct ttctgaagat
catcgatcca 480gcaccagcac cagcaccagc accagcacca gcaccagcac
cagcaattaa tcctgttact 540aatactcagg gcgtgtcccc tataaatact
aaatatgctg aacatgtggt gaaaaatatt 600tacccgaaaa ttaaacatga
ttactttaat gaatcaccca atatatatga taagaagtat 660atatccggta
taaccagagg agtagctgaa ctaaaacagg aagaatttgt taacgagaaa
720gccagacggt tttcttatat gaagactatg tattctgtat gtccagaagc
gtttgaacct 780atttccagaa atgaagccag tacaccggaa ggaagctggc
taacagttat atccggaaaa 840gccccaatgg ggcagttttc tgtagatagt
ttatacaatc ctgatttaca tgcattatgt 900gagcttccgg acatttgttg
taagatcttc cctaaagaaa ataatgattt tttatacata 960gttgttgtgt
acagaaatga cagccctcta ggagaacaac gggcaaatag atttatagaa
1020ttatataata taaaaagaga tatcatgcag gaattaaatt atgagttacc
agagttaaag 1080gcagtaaaat ctgaaatgat tatcgcacgt gaaatgggag
aaatctttag ctacatgcct 1140ggggaaatag acagttatat gaaatacata
aataataaac tttctaaaat tgagggatct 1200catcaccatc accatcacta a
122127252PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 27Gly Ser Met Arg Lys Lys Arg Arg Gln Arg Arg
Arg Ile Asn Pro Val1 5 10 15Thr Asn Thr Gln Ile Asn Pro Val Thr Asn
Thr Gln Gly Val Ser Pro 20 25 30Ile Asn Thr Lys Tyr Ala Glu His Val
Val Lys Asn Ile Tyr Pro Lys 35 40 45Ile Lys His Asp Tyr Phe Asn Glu
Ser Pro Asn Tyr Asp Lys Lys Tyr 50 55 60Ile Ser Gly Ile Thr Arg Gly
Val Ala Glu Leu Lys Gln Glu Glu Phe65 70 75 80Val Asn Glu Lys Ala
Arg Arg Phe Ser Tyr Met Lys Thr Met Tyr Ser 85 90 95Val Cys Pro Glu
Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr 100 105 110Pro Glu
Gly Ser Trp Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly 115 120
125Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys
130 135 140Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn
Asn Asp145 150 155 160Phe Leu Tyr Ile Val Val Val Tyr Arg Asn Asp
Ser Pro Leu Gly Glu 165 170 175Gln Arg Ala Asn Arg Phe Ile Glu Leu
Tyr Asn Ile Lys Arg Asp Ile 180 185 190Met Gln Glu Leu Asn Tyr Glu
Leu Pro Glu Leu Lys Ala Val Lys Ser 195 200 205Glu Met Ile Ile Ala
Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro 210 215 220Gly Glu Ile
Asp Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys225 230 235
240Ile Glu Gly Ser Arg Ser His His His His His His 245
25028246PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 28Gly Ser Met Arg Lys Lys Arg Arg Gln Arg Arg
Arg Ile Asn Pro Val1 5 10 15Thr Asn Thr Gln Ile Asn Pro Val Thr Asn
Thr Gln Gly Val Ser Pro 20 25 30Ile Asn Thr Lys Tyr Ala Glu His Val
Val Lys Asn Ile Tyr Pro Lys 35 40 45Ile Lys His Asp Tyr Phe Asn Glu
Ser Pro Asn Tyr Asp Lys Lys Tyr 50 55 60Ile Ser Gly Ile Thr Arg Gly
Val Ala Glu Leu Lys Gln Glu Glu Phe65 70 75 80Val Asn Glu Lys Ala
Arg Arg Phe Ser Tyr Met Lys Thr Met Tyr Ser 85 90 95Val Cys Pro Glu
Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr 100 105 110Pro Glu
Gly Ser Trp Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly 115 120
125Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys
130 135 140Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn
Asn Asp145 150 155 160Phe Leu Tyr Ile Val Val Val Tyr Arg Asn Asp
Ser Pro Leu Gly Glu 165 170 175Gln Arg Ala Asn Arg Phe Ile Glu Leu
Tyr Asn Ile Lys Arg Asp Ile 180 185 190Met Gln Glu Leu Asn Tyr Glu
Leu Pro Glu Leu Lys Ala Val Lys Ser 195 200 205Glu Met Ile Ile Ala
Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro 210 215 220Gly Glu Ile
Asp Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys225 230 235
240Ile Glu Gly Ser Arg Ser 24529762DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
29ggatccatga ggaagaagcg gagacagcga cgaagaatta atcctgttac taatactcag
60attaatcctg ttactaatac tcagggcgtg tcccctataa atactaaata tgctgaacat
120gtggtgaaaa atatttaccc gaaaattaaa catgattact ttaatgaatc
acccaatata 180tatgataaga agtatatatc cggtataacc agaggagtag
ctgaactaaa acaggaagaa 240tttgttaacg agaaagccag acggttttct
tatatgaaga ctatgtattc tgtatgtcca 300gaagcgtttg aacctatttc
cagaaatgaa gccagtacac cggaaggaag ctggctaaca 360gttatatccg
gaaaacgccc aatggggcag ttttctgtag atagtttata caatcctgat
420ttacatgcat tatgtgagct tccggacatt tgttgtaaga tcttccctaa
agaaaataat 480gattttttat acatagttgt tgtgtacaga aatgacagcc
ctctaggaga acaacgggca 540aatagattta tagaattata taatataaaa
agagatatca tgcaggaatt aaattatgag 600ttaccagagt taaaggcagt
aaaatctgaa atgattatcg cacgtgaaat gggagaaatc 660tttagctaca
tgcctgggga aatagacagt tatatgaaat acataaataa taaactttct
720aaaattgagg gatccagatc tcatcaccat caccatcact aa
76230252PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 30Gly Ser Met Arg Lys Lys Arg Arg Gln Arg Arg
Arg Ile Asn Pro Val1 5 10 15Thr Asn Thr Gln Ile Asn Pro Val Thr Asn
Thr Gln Gly Val Ser Pro 20 25 30Ile Asn Thr Lys Tyr Ala Glu His Val
Val Lys Asn Ile Tyr Pro Lys 35 40 45Ile Lys His Asp Tyr Phe Asn Glu
Ser Pro Asn Tyr Asp Lys Lys Tyr 50 55 60Ile Ser Gly Ile Thr Arg Gly
Val Ala Glu Leu Lys Gln Glu Glu Phe65 70 75 80Val Asn Glu Lys Ala
Arg Arg Phe Ser Tyr Met Lys Thr Met Tyr Ser 85 90 95Val Cys Pro Glu
Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr 100 105 110Pro Glu
Gly Ser Trp Leu Thr Val Ile Ser Gly Lys Ala Pro Met Gly 115 120
125Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys
130 135 140Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn
Asn Asp145 150 155 160Phe Leu Tyr Ile Val Val Val Tyr Arg Asn Asp
Ser Pro Leu Gly Glu 165 170 175Gln Arg Ala Asn Arg Phe Ile Glu Leu
Tyr Asn Ile Lys Arg Asp Ile 180 185 190Met Gln Glu Leu Asn Tyr Glu
Leu Pro Glu Leu Lys Ala Val Lys Ser 195 200 205Glu Met Ile Ile Ala
Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro 210 215 220Gly Glu Ile
Asp Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys225 230 235
240Ile Glu Gly Ser Arg Ser His His His His His His 245
25031246PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 31Gly Ser Met Arg Lys Lys Arg Arg Gln Arg Arg
Arg Ile Asn Pro Val1 5 10 15Thr Asn Thr Gln Ile Asn Pro Val Thr Asn
Thr Gln Gly Val Ser Pro 20 25 30Ile Asn Thr Lys Tyr Ala Glu His Val
Val Lys Asn Ile Tyr Pro Lys 35 40 45Ile Lys His Asp Tyr Phe Asn Glu
Ser Pro Asn Tyr Asp Lys Lys Tyr 50 55 60Ile Ser Gly Ile Thr Arg Gly
Val Ala Glu Leu Lys Gln Glu Glu Phe65 70 75 80Val Asn Glu Lys Ala
Arg Arg Phe Ser Tyr Met Lys Thr Met Tyr Ser 85 90 95Val Cys Pro Glu
Ala Phe Glu Pro Ile Ser Arg Asn Glu Ala Ser Thr 100 105 110Pro Glu
Gly Ser Trp Leu Thr Val Ile Ser Gly Lys Ala Pro Met Gly 115 120
125Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro Asp Leu His Ala Leu Cys
130 135 140Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn
Asn Asp145 150 155 160Phe Leu Tyr Ile Val Val Val Tyr Arg Asn Asp
Ser Pro Leu Gly Glu 165 170 175Gln Arg Ala Asn Arg Phe Ile Glu Leu
Tyr Asn Ile Lys Arg Asp Ile 180 185 190Met Gln Glu Leu Asn Tyr Glu
Leu Pro Glu Leu Lys Ala Val Lys Ser 195 200 205Glu Met Ile Ile Ala
Arg Glu Met Gly Glu Ile Phe Ser Tyr Met Pro 210 215 220Gly Glu Ile
Asp Ser Tyr Met Lys Tyr Ile Asn Asn Lys Leu Ser Lys225 230 235
240Ile Glu Gly Ser Arg Ser 24532762DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
32ggatccatga ggaagaagcg gagacagcga cgaagaatta atcctgttac taatactcag
60attaatcctg ttactaatac tcagggcgtg tcccctataa atactaaata tgctgaacat
120gtggtgaaaa atatttaccc gaaaattaaa catgattact ttaatgaatc
acccaatata 180tatgataaga agtatatatc cggtataacc agaggagtag
ctgaactaaa acaggaagaa 240tttgttaacg agaaagccag acggttttct
tatatgaaga ctatgtattc tgtatgtcca 300gaagcgtttg aacctatttc
cagaaatgaa gccagtacac cggaaggaag ctggctaaca 360gttatatccg
gaaaagcccc aatggggcag ttttctgtag atagtttata caatcctgat
420ttacatgcat tatgtgagct tccggacatt tgttgtaaga tcttccctaa
agaaaataat 480gattttttat acatagttgt tgtgtacaga aatgacagcc
ctctaggaga acaacgggca 540aatagattta tagaattata taatataaaa
agagatatca tgcaggaatt aaattatgag 600ttaccagagt taaaggcagt
aaaatctgaa atgattatcg cacgtgaaat gggagaaatc 660tttagctaca
tgcctgggga aatagacagt tatatgaaat acataaataa taaactttct
720aaaattgagg gatccagatc tcatcaccat caccatcact aa
76233257PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 33Gly Ser Met Arg Lys Lys Arg Arg Gln Arg Arg
Arg Ser Ile Glu Ile1 5 10 15Lys Met Ile Ser Pro Ile Lys Asn Ile Lys
Asn Val Phe Pro Ile Asn 20 25 30Thr Ala Asn Thr Glu Tyr Ile Val Arg
Asn Ile Tyr Pro Arg Val Glu 35 40 45His Gly Tyr Phe Asn Glu Ser Pro
Asn Ile Tyr Asp Lys Lys Tyr Ile 50 55 60Ser Gly Ile Thr Arg Ser Met
Ala Gln Leu Lys Ile Glu Glu Phe Ile65 70 75 80Asn Glu Lys Ser Arg
Arg Leu Asn Tyr Met Lys Thr Met Tyr Ser Pro 85 90 95Cys Pro Glu Asp
Phe Gln Pro Ile Ser Arg Asp Glu Ala Ser Thr Pro 100 105 110Glu Gly
Ser Trp Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln 115 120
125Phe Ser Val Asp Ser Leu Tyr His Pro Asp Leu His Ala Leu Cys Glu
130 135 140Leu Pro Glu Ile Ser Cys Lys Ile Phe Pro Lys Glu Asn Ser
Asp Phe145 150 155 160Leu Tyr Ile Ile Val Val Phe Arg Asn Asp Ser
Pro Gln Gly Glu Leu 165 170 175Arg Ala Asn Arg Phe Ile Glu Leu Tyr
Asp Ile Lys Arg Glu Ile Met 180 185 190Gln Val Leu Arg Asp Glu Ser
Pro Glu Leu Lys Ser Ile Lys Ser Glu 195 200 205Ile Ile Ile Ala Arg
Glu Met Gly Glu Leu Phe Ser Tyr Ala Ser Glu 210 215 220Glu Ile Asp
Ser Tyr Ile Lys Gln Met Asn Asp Arg Leu Ser Gln Ile225 230 235
240Lys Ala Arg Met Pro Val Thr Gly Ser Arg Ser His His His His His
245 250 255His34251PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 34Gly Ser Met Arg Lys Lys Arg Arg
Gln Arg Arg Arg Ser Ile Glu Ile1 5 10 15Lys Met Ile Ser Pro Ile Lys
Asn Ile Lys Asn Val Phe Pro Ile Asn 20 25 30Thr Ala Asn Thr Glu Tyr
Ile Val Arg Asn Ile Tyr Pro Arg Val Glu 35 40 45His Gly Tyr Phe Asn
Glu Ser Pro Asn Ile Tyr Asp Lys Lys Tyr Ile 50 55 60Ser Gly Ile Thr
Arg Ser Met Ala Gln Leu Lys Ile Glu Glu Phe Ile65 70 75 80Asn Glu
Lys Ser Arg Arg Leu Asn Tyr Met Lys Thr Met Tyr Ser Pro 85 90 95Cys
Pro Glu Asp Phe Gln Pro Ile Ser Arg Asp Glu Ala Ser Thr Pro 100 105
110Glu Gly Ser Trp Leu Thr Val Ile Ser Gly Lys Arg Pro Met Gly Gln
115 120 125Phe Ser Val Asp Ser Leu Tyr His Pro Asp Leu His Ala Leu
Cys Glu 130 135 140Leu Pro Glu Ile Ser Cys Lys Ile Phe Pro Lys Glu
Asn Ser Asp Phe145 150 155 160Leu Tyr Ile Ile Val Val Phe Arg Asn
Asp Ser Pro Gln Gly Glu Leu 165 170 175Arg Ala Asn Arg Phe Ile Glu
Leu Tyr Asp Ile Lys Arg Glu Ile Met 180 185 190Gln Val Leu Arg Asp
Glu Ser Pro Glu Leu Lys Ser Ile Lys Ser Glu 195 200 205Ile Ile Ile
Ala Arg Glu Met Gly Glu Leu Phe Ser Tyr Ala Ser Glu 210 215 220Glu
Ile Asp Ser Tyr Ile Lys Gln Met Asn Asp Arg Leu Ser Gln Ile225 230
235 240Lys Ala Arg Met Pro Val Thr Gly Ser Arg Ser 245
25035774DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 35ggatccatga ggaagaagcg gagacagcga
cgaagatcca tcgagatcaa gatgatctct 60ccgatcaaga acatcaagaa cgtattcccg
atcaacactg cgaataccga gtacatcgtt 120cgcaacatct acccacgcgt
agaacacggc tacttcaacg agagcccaaa catctacgac 180aagaagtaca
tcagcggtat cactcgctct atggctcaac tgaagatcga agagttcatc
240aacgagaagt cccgtcgtct gaactacatg aagaccatgt actctccgtg
tccggaagat 300ttccaaccga tctctcgtga tgaagctagc actccagaag
gcagctggct gaccgtgatc 360tctggcaaac gtccgatggg tcagttctcc
gttgactctc tgtaccatcc agacctgcac 420gctctgtgcg aactgccaga
gatttcttgc aagatctttc cgaaagagaa ctctgatttc 480ctgtacatca
tcgttgtgtt ccgcaacgat tctccacaag gtgaactgcg tgctaaccgc
540ttcatcgaac tgtacgatat caagcgtgag atcatgcagg tgctgcgcga
cgagtctcca 600gaactgaaga gcatcaaatc cgagatcatc attgcccgtg
aaatgggcga actgttctct 660tacgcatctg aagaaatcga ctcttacatc
aagcagatga acgatcgtct gtcccagatc 720aaggctcgta tgccggtaac
cggatccaga tctcatcacc atcaccatca ctaa 77436322PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
36Gly Ser Met Phe Ile Thr Pro Arg Asn Val Ser Asn Thr Phe Leu Gln1
5 10 15Glu Pro Leu Arg His Ser Ser Asp Leu Thr Glu Met Pro Val Glu
Ala 20 25 30Glu Asn Val Lys Ser Lys Thr Glu Tyr Tyr Asn Ala Trp Ala
Val Trp 35 40 45Glu Arg Asn Ala Pro Pro Gly Asn Gly Glu Gln Arg Glu
Met Ala Val 50 55 60Ser Arg Leu Arg Asp Cys
Leu Asp Arg Gln Ala His Glu Leu Glu Leu65 70 75 80Asn Asn Leu Gly
Leu Ser Ser Leu Met Ile Asn Pro Val Thr Asn Thr 85 90 95Gln Gly Val
Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys 100 105 110Asn
Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro Asn 115 120
125Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly Val Ala Glu
130 135 140Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala Arg Arg Phe
Ser Tyr145 150 155 160Met Lys Thr Met Tyr Ser Val Cys Pro Glu Ala
Phe Glu Pro Ile Ser 165 170 175Arg Asn Glu Ala Ser Thr Pro Glu Gly
Ser Trp Leu Thr Val Ile Ser 180 185 190Gly Lys Arg Pro Met Gly Gln
Phe Ser Val Asp Ser Leu Tyr Asn Pro 195 200 205Asp Leu His Ala Leu
Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe 210 215 220Pro Lys Glu
Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn225 230 235
240Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu Tyr
245 250 255Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu Leu
Pro Glu 260 265 270Leu Lys Ala Val Lys Ser Glu Met Ile Ile Ala Arg
Glu Met Gly Glu 275 280 285Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp
Ser Tyr Met Lys Tyr Ile 290 295 300Asn Asn Lys Leu Ser Lys Ile Glu
Gly Ser Arg Ser His His His His305 310 315 320His
His37316PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 37Gly Ser Met Phe Ile Thr Pro Arg Asn Val Ser
Asn Thr Phe Leu Gln1 5 10 15Glu Pro Leu Arg His Ser Ser Asp Leu Thr
Glu Met Pro Val Glu Ala 20 25 30Glu Asn Val Lys Ser Lys Thr Glu Tyr
Tyr Asn Ala Trp Ala Val Trp 35 40 45Glu Arg Asn Ala Pro Pro Gly Asn
Gly Glu Gln Arg Glu Met Ala Val 50 55 60Ser Arg Leu Arg Asp Cys Leu
Asp Arg Gln Ala His Glu Leu Glu Leu65 70 75 80Asn Asn Leu Gly Leu
Ser Ser Leu Met Ile Asn Pro Val Thr Asn Thr 85 90 95Gln Gly Val Ser
Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys 100 105 110Asn Ile
Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro Asn 115 120
125Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly Val Ala Glu
130 135 140Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala Arg Arg Phe
Ser Tyr145 150 155 160Met Lys Thr Met Tyr Ser Val Cys Pro Glu Ala
Phe Glu Pro Ile Ser 165 170 175Arg Asn Glu Ala Ser Thr Pro Glu Gly
Ser Trp Leu Thr Val Ile Ser 180 185 190Gly Lys Arg Pro Met Gly Gln
Phe Ser Val Asp Ser Leu Tyr Asn Pro 195 200 205Asp Leu His Ala Leu
Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe 210 215 220Pro Lys Glu
Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn225 230 235
240Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu Tyr
245 250 255Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu Leu
Pro Glu 260 265 270Leu Lys Ala Val Lys Ser Glu Met Ile Ile Ala Arg
Glu Met Gly Glu 275 280 285Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp
Ser Tyr Met Lys Tyr Ile 290 295 300Asn Asn Lys Leu Ser Lys Ile Glu
Gly Ser Arg Ser305 310 31538969DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 38ggatccatgt
tcatcactcc acgcaatgta tctaacacct ttctgcagga accgctgcgt 60cattctagcg
acctgaccga aatgccagtt gaagcggaga acgtgaaatc taagactgaa
120tactacaacg cgtgggcagt atgggagcgc aatgcaccac caggtaacgg
tgaacagcgt 180gaaatggcag taagccgtct gcgtgattgc ctggatcgcc
aggctcacga gctggagctg 240aacaacctgg gtctgtctag cctgatgatt
aatcctgtta ctaatactca gggcgtgtcc 300cctataaata ctaaatatgc
tgaacatgtg gtgaaaaata tttacccgaa aattaaacat 360gattacttta
atgaatcacc caatatatat gataagaagt atatatccgg tataaccaga
420ggagtagctg aactaaaaca ggaagaattt gttaacgaga aagccagacg
gttttcttat 480atgaagacta tgtattctgt atgtccagaa gcgtttgaac
ctatttccag aaatgaagcc 540agtacaccgg aaggaagctg gctaacagtt
atatccggaa aacgcccaat ggggcagttt 600tctgtagata gtttatacaa
tcctgattta catgcattat gtgagcttcc ggacatttgt 660tgtaagatct
tccctaaaga aaataatgat tttttataca tagttgttgt gtacagaaat
720gacagccctc taggagaaca acgggcaaat agatttatag aattatataa
tataaaaaga 780gatatcatgc aggaattaaa ttatgagtta ccagagttaa
aggcagtaaa atctgaaatg 840attatcgcac gtgaaatggg agaaatcttt
agctacatgc ctggggaaat agacagttat 900atgaaataca taaataataa
actttctaaa attgagggat ccagatctca tcaccatcac 960catcactaa
96939322PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 39Gly Ser Met Phe Ile Thr Pro Arg Asn Val Ser
Asn Thr Phe Leu Gln1 5 10 15Glu Pro Leu Arg His Ser Ser Asp Leu Thr
Glu Met Pro Val Glu Ala 20 25 30Glu Asn Val Lys Ser Lys Thr Glu Tyr
Tyr Asn Ala Trp Ala Val Trp 35 40 45Glu Arg Asn Ala Pro Pro Gly Asn
Gly Glu Gln Arg Glu Met Ala Val 50 55 60Ser Arg Leu Arg Asp Cys Leu
Asp Arg Gln Ala His Glu Leu Glu Leu65 70 75 80Asn Asn Leu Gly Leu
Ser Ser Leu Met Ile Asn Pro Val Thr Asn Thr 85 90 95Gln Gly Val Ser
Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys 100 105 110Asn Ile
Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro Asn 115 120
125Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly Val Ala Glu
130 135 140Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala Arg Arg Phe
Ser Tyr145 150 155 160Met Lys Thr Met Tyr Ser Val Cys Pro Glu Ala
Phe Glu Pro Ile Ser 165 170 175Arg Asn Glu Ala Ser Thr Pro Glu Gly
Ser Trp Leu Thr Val Ile Ser 180 185 190Gly Lys Ala Pro Met Gly Gln
Phe Ser Val Asp Ser Leu Tyr Asn Pro 195 200 205Asp Leu His Ala Leu
Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe 210 215 220Pro Lys Glu
Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn225 230 235
240Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu Tyr
245 250 255Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu Leu
Pro Glu 260 265 270Leu Lys Ala Val Lys Ser Glu Met Ile Ile Ala Arg
Glu Met Gly Glu 275 280 285Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp
Ser Tyr Met Lys Tyr Ile 290 295 300Asn Asn Lys Leu Ser Lys Ile Glu
Gly Ser Arg Ser His His His His305 310 315 320His
His40316PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 40Gly Ser Met Phe Ile Thr Pro Arg Asn Val Ser
Asn Thr Phe Leu Gln1 5 10 15Glu Pro Leu Arg His Ser Ser Asp Leu Thr
Glu Met Pro Val Glu Ala 20 25 30Glu Asn Val Lys Ser Lys Thr Glu Tyr
Tyr Asn Ala Trp Ala Val Trp 35 40 45Glu Arg Asn Ala Pro Pro Gly Asn
Gly Glu Gln Arg Glu Met Ala Val 50 55 60Ser Arg Leu Arg Asp Cys Leu
Asp Arg Gln Ala His Glu Leu Glu Leu65 70 75 80Asn Asn Leu Gly Leu
Ser Ser Leu Met Ile Asn Pro Val Thr Asn Thr 85 90 95Gln Gly Val Ser
Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys 100 105 110Asn Ile
Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro Asn 115 120
125Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly Val Ala Glu
130 135 140Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala Arg Arg Phe
Ser Tyr145 150 155 160Met Lys Thr Met Tyr Ser Val Cys Pro Glu Ala
Phe Glu Pro Ile Ser 165 170 175Arg Asn Glu Ala Ser Thr Pro Glu Gly
Ser Trp Leu Thr Val Ile Ser 180 185 190Gly Lys Ala Pro Met Gly Gln
Phe Ser Val Asp Ser Leu Tyr Asn Pro 195 200 205Asp Leu His Ala Leu
Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe 210 215 220Pro Lys Glu
Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn225 230 235
240Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu Tyr
245 250 255Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu Leu
Pro Glu 260 265 270Leu Lys Ala Val Lys Ser Glu Met Ile Ile Ala Arg
Glu Met Gly Glu 275 280 285Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp
Ser Tyr Met Lys Tyr Ile 290 295 300Asn Asn Lys Leu Ser Lys Ile Glu
Gly Ser Arg Ser305 310 31541969DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 41ggatccatgt
tcatcactcc acgcaatgta tctaacacct ttctgcagga accgctgcgt 60cattctagcg
acctgaccga aatgccagtt gaagcggaga acgtgaaatc taagactgaa
120tactacaacg cgtgggcagt atgggagcgc aatgcaccac caggtaacgg
tgaacagcgt 180gaaatggcag taagccgtct gcgtgattgc ctggatcgcc
aggctcacga gctggagctg 240aacaacctgg gtctgtctag cctgatgatt
aatcctgtta ctaatactca gggcgtgtcc 300cctataaata ctaaatatgc
tgaacatgtg gtgaaaaata tttacccgaa aattaaacat 360gattacttta
atgaatcacc caatatatat gataagaagt atatatccgg tataaccaga
420ggagtagctg aactaaaaca ggaagaattt gttaacgaga aagccagacg
gttttcttat 480atgaagacta tgtattctgt atgtccagaa gcgtttgaac
ctatttccag aaatgaagcc 540agtacaccgg aaggaagctg gctaacagtt
atatccggaa aagccccaat ggggcagttt 600tctgtagata gtttatacaa
tcctgattta catgcattat gtgagcttcc ggacatttgt 660tgtaagatct
tccctaaaga aaataatgat tttttataca tagttgttgt gtacagaaat
720gacagccctc taggagaaca acgggcaaat agatttatag aattatataa
tataaaaaga 780gatatcatgc aggaattaaa ttatgagtta ccagagttaa
aggcagtaaa atctgaaatg 840attatcgcac gtgaaatggg agaaatcttt
agctacatgc ctggggaaat agacagttat 900atgaaataca taaataataa
actttctaaa attgagggat ccagatctca tcaccatcac 960catcactaa
96942240PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 42Met Thr Lys Ile Thr Leu Ser Pro Gln Asn Phe
Arg Ile Gln Lys Gln1 5 10 15Glu Thr Thr Leu Leu Lys Glu Lys Ser Thr
Glu Lys Asn Ser Leu Ala 20 25 30Lys Ser Ile Leu Ala Val Lys Asn His
Phe Ile Glu Leu Arg Ser Lys 35 40 45Leu Ser Glu Arg Phe Ile Ser His
Lys Asn Thr Glu Ser Ser Ala Thr 50 55 60His Phe His Arg Gly Ser Ala
Ser Glu Gly Arg Ala Val Leu Thr Asn65 70 75 80Lys Val Val Lys Asp
Phe Met Leu Gln Thr Leu Asn Asp Ile Asp Ile 85 90 95Arg Gly Ser Ala
Ser Lys Asp Pro Ala Tyr Ala Ser Gln Thr Arg Glu 100 105 110Ala Ile
Leu Ser Ala Val Tyr Ser Lys Asn Lys Asp Gln Cys Cys Asn 115 120
125Leu Leu Ile Ser Lys Gly Ile Asn Ile Ala Pro Phe Leu Gln Glu Ile
130 135 140Gly Glu Ala Ala Lys Asn Ala Gly Leu Pro Gly Thr Thr Lys
Asn Asp145 150 155 160Val Phe Thr Pro Ser Gly Ala Gly Ala Asn Pro
Phe Ile Thr Pro Leu 165 170 175Ile Ser Ser Ala Asn Ser Lys Tyr Pro
Arg Met Phe Ile Asn Gln His 180 185 190Gln Gln Ala Ser Phe Lys Ile
Tyr Ala Glu Lys Ile Ile Met Thr Glu 195 200 205Val Ala Pro Leu Phe
Asn Glu Cys Ala Met Pro Thr Pro Gln Gln Phe 210 215 220Gln Leu Ile
Leu Glu Asn Ile Ala Asn Lys Tyr Ile Gln Asn Thr Pro225 230 235
240431011DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 43tttctaattc atcaatcaga tggacatagc
atttgctata aaaaataaaa gtattcctgc 60tatctatata taaatgagtt atgtacatat
aaaaggagca ttaccgtgac aaaaataact 120ttatttcccc ataactttag
aatccaaaaa caggaagcca caccactaaa agaaaaatca 180accgagaaaa
attctttagc aaaaagtatt ctcgcagtaa aaaatcactt catcaaatta
240aattcaaaat tatcggaacg ttttatttcg cataagaaca ctgaatcttc
tgcaacacac 300tttcaccgag gaagcgcatc tgagggccgg gcagtgttga
caaataaagt cgttaaaaac 360tttatgcttc aaacgctcca tgatatagat
attagaggta gcgcgagtaa agaccccgca 420tacgccagcc agacccgtga
agctatacta tcggcagttt acagcaagta taaagatcag 480tattgtaact
tgctcatcag caaaggaatc gacatagcgc cttttcttaa ggaaattggc
540gaggctgcgc aaaatgcagg tctgcccgga gcaaccaaga atgacgtttt
tagcccaagc 600ggcgcaggag ccaatccttt tataactccg ttgattacat
cagcatacag taagtatcca 660catatgttta ccagtcaaca tcagaaggca
tcctttaaca tctatgcgga gaagatcatt 720atgacagaag ttgtaccgct
gtttaatgag tgtgctatgc cgactccaca gcaattccaa 780caaatactag
aaaacattgc taataaatat atccaaaaca ctccctgaac acagaaacac
840caaaaaatat gcgagcctct tcctgattaa tatgaaccaa tagtatccat
aatttttccc 900aggaactaac tctggagcta aaccgtcatt taccagtgct
aaaattatac actcaaccat 960caaaataata gccattgctg ctatataaca
tatagcagca atctctacta c 101144561PRTArtificial SequenceDescription
of Artificial Sequence Synthetic polypeptide 44Met Gln Ile Gln Ser
Phe Tyr His Ser Ala Ser Leu Lys Thr Gln Glu1 5 10 15Ala Phe Lys Ser
Leu Gln Lys Thr Leu Tyr Asn Gly Met Gln Ile Leu 20 25 30Ser Gly Gln
Gly Lys Ala Pro Ala Lys Ala Pro Asp Ala Arg Pro Glu 35 40 45Ile Ile
Val Leu Arg Glu Pro Gly Ala Thr Trp Gly Asn Tyr Leu Gln 50 55 60His
Gln Lys Ala Ser Asn His Ser Leu His Asn Leu Tyr Asn Leu Gln65 70 75
80Arg Asp Leu Leu Thr Val Ala Ala Thr Val Leu Gly Lys Gln Asp Pro
85 90 95Val Leu Thr Ser Met Ala Asn Gln Met Glu Leu Ala Lys Val Lys
Ala 100 105 110Asp Arg Pro Ala Thr Lys Gln Glu Glu Ala Ala Ala Lys
Ala Leu Lys 115 120 125Lys Asn Leu Ile Glu Leu Ile Ala Ala Arg Thr
Gln Gln Gln Asp Gly 130 135 140Leu Pro Ala Lys Glu Ala His Arg Phe
Ala Ala Val Ala Phe Arg Asp145 150 155 160Ala Gln Val Lys Gln Leu
Asn Asn Gln Pro Trp Gln Thr Ile Lys Asn 165 170 175Thr Leu Thr His
Asn Gly His His Tyr Thr Asn Thr Gln Leu Pro Ala 180 185 190Ala Glu
Met Lys Ile Gly Ala Lys Asp Ile Phe Pro Ser Ala Tyr Glu 195 200
205Gly Lys Gly Val Cys Ser Trp Asp Thr Lys Asn Ile His His Ala Asn
210 215 220Asn Leu Trp Met Ser Thr Val Ser Val His Glu Asp Gly Lys
Asp Lys225 230 235 240Thr Leu Phe Cys Gly Ile Arg His Gly Val Leu
Ser Pro Tyr His Glu 245 250 255Lys Asp Pro Leu Leu Arg His Val Gly
Ala Glu Asn Lys Ala Lys Glu 260 265 270Val Leu Thr Ala Ala Leu Phe
Ser Lys Pro Glu Leu Leu Asn Lys Ala 275 280 285Leu Ala Gly Glu Ala
Val Ser Leu Lys Leu Val Ser Val Gly Leu Leu 290 295 300Thr Ala Ser
Asn Ile Phe Gly Lys Glu Gly Thr Met Val Glu Asp Gln305 310 315
320Met Arg Ala Trp Gln Ser Leu Thr Gln Pro Gly Lys Met Ile His Leu
325 330 335Lys Ile Arg Asn Lys Asp Gly Asp Leu Gln Thr Val Lys Ile
Lys Pro 340 345 350Asp Val Ala Ala Phe Asn Val Gly Val Asn Glu Leu
Ala Leu Lys Leu 355 360 365Gly Phe Gly Leu Lys Ala Ser Asp Ser Tyr
Asn Ala Glu Ala Leu His 370 375 380Gln Leu Leu Gly Asn Asp Leu Arg
Pro Glu Ala Arg Pro Gly Gly Trp385 390 395 400Val Gly Glu Trp Leu
Ala Gln Tyr Pro Asp Asn Tyr Glu Val Val Asn 405 410 415Thr Leu Ala
Arg Gln Ile Lys
Asp Ile Trp Lys Asn Asn Gln His His 420 425 430Lys Asp Gly Gly Glu
Pro Tyr Lys Leu Ala Gln Arg Leu Ala Met Leu 435 440 445Ala His Glu
Ile Asp Ala Val Pro Ala Trp Asn Cys Lys Ser Gly Lys 450 455 460Asp
Arg Thr Gly Met Met Asp Ser Glu Ile Lys Arg Glu Ile Ile Ser465 470
475 480Leu His Gln Thr His Met Leu Ser Ala Pro Gly Ser Leu Pro Asp
Ser 485 490 495Gly Gly Gln Lys Ile Phe Gln Lys Val Leu Leu Asn Ser
Gly Asn Leu 500 505 510Glu Ile Gln Lys Gln Asn Thr Gly Gly Ala Gly
Asn Lys Val Met Lys 515 520 525Asn Leu Ser Pro Glu Val Leu Asn Leu
Ser Tyr Gln Lys Arg Val Gly 530 535 540Asp Glu Asn Ile Trp Gln Ser
Val Lys Gly Ile Ser Ser Leu Ile Thr545 550 555
560Ser451253DNAArtificial SequenceDescription of Artificial
Sequence Synthetic polynucleotide 45tgcctgcaaa ggaggctcgc
cgccttgcgg cagcagattt taagagtgcc caggtcaagc 60agcttaataa tcaaccgtgg
cagaccataa aaaatacact aacgcataat gggcatcaat 120ataccagcac
gcaggttcct gccgcagaga tgaaaatcgg cgcacaggat atttttccta
180aagcctatca gggaaagggc gtatgcagtt gggataccca aaatattcat
cacgctacta 240acctgtggat gtccacaata agtgtacatg aggacggtga
agataaaacg ctttttagtg 300ggatacgtca tggcgtgctt tcaccctatc
atgtggaaga tccgcttctg cgtcagaccg 360gcgctgaaag cagagccaaa
gaagtattaa ctgcagcgct ctttagtaaa cctgagttgc 420ttaccagggc
cttaaagggc gaagcggtaa gcctgaaact ggtatctgtc tgtttactca
480ccgcgtcgaa tgttctaggc caggagggaa caatggtcaa ggagcaaatg
agagcatggc 540aatcgttgac ccagccggga aaaatgattc atttaaaaat
ccgcaatgac gatggcgaac 600tacagacggt aaaaataaag ccggaagtcg
ccgcgttcaa tgtgggtgtc aatgagttag 660cactcaagtt cggctttggc
cttaaggcat ctgatagtta taatatcgag gcgctacagc 720agttattagg
caatgattta cgccctgagg ccagaccagg cggttgggtt ggcgagtggc
780tggcgcgata tccggataac gatgaaagcg taaatacatt agcacgccag
attaaagata 840tctggcaaaa taagctgcat cacaaagatg gcggcgaacc
ctataaatta gcacaacgcc 900ttgcaatgtt agccaatgaa atcgatgtgg
tacccgcctg gaattgtaaa agcggcaaag 960atcgcacagg aatgatggac
tcagaaacca agcgagaagc catttctttc catcagaccc 1020atactttgag
ttctccaggc agccttccgg atcgcagcgg acagcaaatc ttccaaaaag
1080tattacttaa tagcggaaac ctggaaattc aaaaacaaaa tacaagcggg
gcgggaaaca 1140aagtaataaa aaacttgtcg ccagaagtgc ttaatctttc
ctatcataaa cgaattggag 1200atgaaaatac ctggcaatcg gtaaagggaa
tttctacatt aatcatttct tga 125346240PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
46Met Thr Asn Ile Thr Leu Ser Thr Gln His Tyr Arg Ile His Arg Ser1
5 10 15Asp Val Glu Pro Val Lys Glu Lys Thr Thr Glu Lys Asp Ile Phe
Ala 20 25 30Lys Ser Ile Thr Ala Val Arg Asn Ser Phe Ile Ser Leu Ser
Thr Ser 35 40 45Leu Ser Asp Arg Phe Ser Leu His Gln Gln Thr Asp Ile
Pro Thr Thr 50 55 60His Phe His Arg Gly Asn Ala Ser Glu Gly Arg Ala
Val Leu Thr Ser65 70 75 80Lys Thr Val Lys Asp Phe Met Leu Gln Lys
Leu Asn Ser Leu Asp Ile 85 90 95Lys Gly Asn Ala Ser Lys Asp Pro Ala
Tyr Ala Arg Gln Thr Cys Glu 100 105 110Ala Ile Leu Ser Ala Val Tyr
Ser Asn Asn Lys Asp Gln Cys Cys Lys 115 120 125Leu Leu Ile Ser Lys
Gly Val Ser Ile Thr Pro Phe Leu Lys Glu Ile 130 135 140Gly Glu Ala
Ala Gln Asn Ala Gly Leu Pro Gly Glu Ile Lys Asn Gly145 150 155
160Val Phe Thr Pro Gly Gly Ala Gly Ala Asn Pro Phe Val Val Pro Leu
165 170 175Ile Ala Ser Ala Ser Ile Lys Tyr Pro His Met Phe Ile Asn
His Asn 180 185 190Gln Gln Val Ser Phe Lys Ala Tyr Ala Glu Lys Ile
Val Met Lys Glu 195 200 205Val Thr Pro Leu Phe Asn Lys Gly Thr Met
Pro Thr Pro Gln Gln Phe 210 215 220Gln Leu Thr Ile Glu Asn Ile Ala
Asn Lys Tyr Leu Gln Asn Ala Ser225 230 235 240471316DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
47taagtgacag aagaacaaaa tccatcagga aaataaaatt tataaatatc aatgagtaaa
60aatggttgtg gagaaggtgg ctattttttg aaagcaagaa atataaacaa agtgtagcta
120tgcatagtta tctaaaagga gaactaccgt gactaacata acactatcca
cccagcacta 180cagaatccat agaagtgacg ttgaaccagt aaaagaaaaa
acaacggaga aggacatttt 240tgcaaaaagt attactgccg ttagaaatag
ctttatcagc ctgtcgacga gtctgtcaga 300tcgttttagc ctgcatcaac
aaacagacat accgactacc cattttcatc gtgggaacgc 360ttctgagggt
agggcggtat taaccagtaa aactgttaaa gattttatgc tgcaaaagct
420caatagtctg gatatcaaag gtaatgcgag taaagatccg gcctatgctc
gtcagacatg 480cgaagccata ttatcagccg tgtacagtaa taataaagat
caatgttgta aattactcat 540cagtaaaggg gtcagtatta cccccttttt
gaaagagata ggagaggctg cgcagaatgc 600ggggctacct ggggagataa
aaaatggcgt atttactcca ggtggggcag gggcgaatcc 660ttttgtcgtc
cccctcattg cttccgcgag tattaaatat ccgcatatgt ttataaatca
720taatcagcag gtatctttta aagcgtatgc tgagaaaatc gttatgaaag
aggttacgcc 780gctgtttaat aaggggacga tgccaacgcc acaacaattt
cagttaacta tagaaaatat 840tgcgaataag tatcttcaga atgcctcctg
atggtagtaa atacaatact gctattaacc 900atatgaatta atagcagatt
ttctgaaact atatattcgt ttttatggcg acgcttttaa 960caaaacgtcg
tcatctttat tttaatataa aaatcaaatg gataacctta aaacttaaaa
1020taattgttat tactctaaat acctgcacaa ccagacagct ttttttgtaa
cgctgcgtgt 1080tgttaagtct atgatacccg cgcacggatg atgtctcgat
aaggattaat ggtctgttaa 1140ctgtcaggag aggttatgcc gcctttaaaa
aaaattgtgc tgcgcctgtt tgtgggggcg 1200atggtcgcca cggtaacgac
acctgcattg gcgctggttt gccttgagga tcactccgct 1260aaagagtgtg
ctatatcgtg tgctgaggtg atgtggttta tgtctcgaga tcggat
131648900PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 48Met Pro Met Trp Ala Gly Gly Val Gly Ser Pro
Arg Arg Gly Met Ala1 5 10 15Pro Ala Ser Thr Asp Asp Leu Phe Ala Arg
Lys Leu Arg Gln Pro Ala 20 25 30Arg Pro Pro Leu Thr Pro His Thr Phe
Glu Pro Arg Pro Val Arg Gly 35 40 45Pro Leu Leu Arg Ser Gly Ser Asp
Ala Gly Glu Ala Arg Pro Pro Thr 50 55 60Pro Ala Ser Pro Arg Ala Arg
Ala His Ser His Glu Glu Ala Ser Arg65 70 75 80Pro Ala Ala Thr Ser
Thr Arg Leu Phe Thr Asp Pro Leu Ala Leu Leu 85 90 95Gly Leu Pro Ala
Glu Glu Pro Glu Pro Ala Phe Pro Pro Val Leu Glu 100 105 110Pro Arg
Trp Phe Ala His Tyr Asp Val Gln Ser Leu Leu Phe Asp Trp 115 120
125Ala Pro Arg Ser Gln Gly Met Gly Ser His Ser Glu Ala Ser Ser Gly
130 135 140Thr Leu Ala Ser Ala Glu Asp Gln Ala Ala Ser Ser Asp Leu
Leu His145 150 155 160Gly Ala Pro Gly Phe Val Cys Glu Leu Gly Gly
Glu Gly Glu Leu Gly 165 170 175Leu Gly Gly Pro Ala Ser Pro Pro Val
Pro Pro Ala Leu Pro Asn Ala 180 185 190Ala Val Ser Ile Leu Glu Glu
Pro Gln Asn Arg Thr Ser Ala Tyr Ser 195 200 205Leu Glu His Ala Asp
Leu Gly Ala Gly Tyr Tyr Arg Lys Tyr Phe Tyr 210 215 220Gly Lys Glu
His Gln Asn Phe Phe Gly Met Asp Glu Ser Leu Gly Pro225 230 235
240Val Ala Val Ser Leu Arg Arg Glu Glu Lys Glu Gly Ser Gly Gly Gly
245 250 255Thr Leu His Ser Tyr Arg Val Ile Val Arg Thr Thr Gln Leu
Arg Thr 260 265 270Leu Arg Gly Thr Ile Ser Glu Asp Ala Leu Pro Pro
Gly Pro Pro Arg 275 280 285Gly Leu Ser Pro Arg Lys Leu Leu Glu His
Val Ala Pro Gln Leu Ser 290 295 300Pro Ser Cys Leu Arg Leu Gly Ser
Ala Ser Pro Lys Val Pro Arg Thr305 310 315 320Leu Leu Thr Leu Asp
Glu Gln Val Leu Ser Phe Gln Arg Lys Val Gly 325 330 335Ile Leu Tyr
Cys Arg Ala Gly Gln Gly Ser Glu Glu Glu Met Tyr Asn 340 345 350Asn
Gln Glu Ala Gly Pro Ala Phe Met Gln Phe Leu Thr Leu Leu Gly 355 360
365Asp Val Val Arg Leu Lys Gly Phe Glu Ser Tyr Arg Ala Gln Leu Asp
370 375 380Thr Lys Thr Asp Ser Thr Gly Thr His Ser Leu Tyr Thr Thr
Tyr Gln385 390 395 400Asp His Glu Ile Met Phe His Val Ser Thr Met
Leu Pro Tyr Thr Pro 405 410 415Asn Asn Gln Gln Gln Leu Leu Arg Lys
Arg His Ile Gly Asn Asp Ile 420 425 430Val Thr Ile Val Phe Gln Glu
Pro Gly Ser Lys Pro Phe Cys Pro Thr 435 440 445Thr Ile Arg Ser His
Phe Gln His Val Phe Leu Val Val Arg Ala His 450 455 460Thr Pro Cys
Thr Pro His Thr Thr Tyr Arg Val Ala Val Ser Arg Thr465 470 475
480Gln Asp Thr Pro Ala Phe Gly Pro Ala Leu Pro Ala Gly Gly Gly Pro
485 490 495Phe Ala Ala Asn Ala Asp Phe Arg Ala Phe Leu Leu Ala Lys
Ala Leu 500 505 510Asn Gly Glu Gln Ala Ala Gly His Ala Arg Gln Phe
His Ala Met Ala 515 520 525Thr Arg Thr Arg Gln Gln Tyr Leu Gln Asp
Leu Ala Thr Asn Glu Val 530 535 540Thr Thr Thr Ser Leu Asp Ser Ala
Ser Arg Phe Gly Leu Pro Ser Leu545 550 555 560Gly Gly Arg Arg Arg
Ala Ala Pro Arg Gly Pro Gly Ala Glu Leu Gln 565 570 575Ala Ala Gly
Ser Leu Val Trp Gly Val Arg Ala Ala Pro Gly Ala Arg 580 585 590Val
Ala Ala Gly Ala Gln Ala Ser Gly Pro Glu Gly Ile Glu Val Pro 595 600
605Cys Leu Leu Gly Ile Ser Ala Glu Ala Leu Val Leu Val Ala Pro Arg
610 615 620Asp Gly Arg Val Val Phe Asn Cys Ala Cys Arg Asp Val Leu
Ala Trp625 630 635 640Thr Phe Ser Glu Gln Gln Leu Asp Leu Tyr His
Gly Arg Gly Glu Ala 645 650 655Ile Thr Leu Arg Phe Asp Gly Ser Pro
Gly Gln Ala Val Gly Glu Val 660 665 670Val Ala Arg Leu Gln Leu Val
Ser Arg Gly Cys Glu Thr Arg Glu Leu 675 680 685Ala Leu Pro Arg Asp
Gly Gln Gly Arg Leu Gly Phe Glu Val Asp Ala 690 695 700Glu Gly Phe
Val Thr His Val Glu Arg Phe Thr Phe Ala Glu Thr Ala705 710 715
720Gly Leu Arg Pro Gly Ala Arg Leu Leu Arg Val Cys Gly Gln Thr Leu
725 730 735Pro Ser Leu Arg Pro Glu Ala Ala Ala Gln Leu Leu Arg Ser
Ala Pro 740 745 750Lys Val Cys Val Thr Val Leu Pro Pro Asp Glu Ser
Gly Arg Pro Arg 755 760 765Arg Ser Phe Ser Glu Leu Tyr Thr Leu Ser
Leu Gln Glu Pro Ser Arg 770 775 780Arg Gly Ala Pro Asp Pro Val Gln
Asp Glu Val Gln Gly Val Thr Leu785 790 795 800Leu Pro Thr Thr Lys
Gln Leu Leu His Leu Cys Leu Gln Asp Gly Gly 805 810 815Ser Pro Pro
Gly Pro Gly Asp Leu Ala Glu Glu Arg Thr Glu Phe Leu 820 825 830His
Ser Gln Asn Ser Leu Ser Pro Arg Ser Ser Leu Ser Asp Glu Ala 835 840
845Pro Val Leu Pro Asn Thr Thr Pro Asp Leu Leu Leu Ala Thr Thr Ala
850 855 860Lys Pro Ser Val Pro Ser Ala Asp Ser Glu Thr Pro Leu Thr
Gln Asp865 870 875 880Arg Pro Gly Ser Pro Ser Gly Ser Glu Asp Lys
Gly Asn Pro Ala Pro 885 890 895Glu Leu Arg Ala
90049720DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 49atggttacaa gtgtaaggac tcagcccccc
gtcataatgc caggtatgca gaccgagatc 60aaaacgcagg ccacgaatct tgcggcgaat
ctttccgcag tcagagaaag tgccacaacg 120acgctgtcag gggaaattaa
aggcccgcaa ctggaagatt ttcccgcgct gatcaaacag 180gcgagtctgg
atgcattgtt taaatgcggg aaagacgctg aggcgttaaa agaagttttt
240accaattcaa ataatgtcgc cggtaagaaa gcgataatgg agtttgccgg
gctctttcgt 300tcagcgctca acgccaccag tgattctcct gaggcgaaga
cgctactgat gaaggtgggg 360gcagagtata ccgcgcaaat cataaaagat
ggcctgaaag aaaagtcagc ttttgggcca 420tggctgccag aaacaaagaa
agcggaagcg aagctggaaa acctggaaaa gcagctgtta 480gatattatca
aaaataacac tggtggtgaa ttaagtaaat tatcgacgaa tcttgttatg
540caggaggtga tgccctatat tgccagctgc attgaacata actttggctg
tacgttagat 600ccgttaaccc gcagcaatct tacgcacctt gttgacaaag
cggcggcgaa ggcggttgag 660gcgcttgata tgtgccacca aaaattaacg
caagagcagg gtaccagcgt aggacgggaa 72050907PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
50Met Trp Asn Val Ser Lys Ser Ser Asn Asn Leu Gly Ala Tyr Lys Leu1
5 10 15Pro Leu Glu Ala Gln Thr Pro Pro Glu Lys Ile Ser Pro Phe Asp
Ala 20 25 30Met Ser Ala Ala Gln Pro Glu Gly Lys Ala Pro His Asp Gln
Leu Gln 35 40 45Asn Asp Gln Tyr Pro Ile Gln Gln Ala Glu Asp Arg Gly
Arg His Leu 50 55 60Val Glu Gln Ala Glu Ile Gln Ala His Val Gln His
Cys His Ser Lys65 70 75 80Ala Pro Glu Ile Gly Asp Ala Thr Lys Thr
Gln Ser Val Ser Glu Lys 85 90 95Leu Gly Thr Ala Lys Asn Ser Ser Cys
Asp Ala Ser Gln Ile Leu Ile 100 105 110Gly Ser Lys Asn Asp Asp Phe
His Lys Asn Lys Ala Gly Ser Asn Gly 115 120 125Asp Ile Asn Lys Ser
Ser Asp Pro Ser Ala Leu Arg Cys Ser Leu Ser 130 135 140Pro Ala Pro
Arg Arg Val Pro Lys Ser Lys Lys Ser Tyr Gly Ala Ala145 150 155
160Thr Ile Gly Gly Lys Val Tyr His Pro His Glu Lys Thr Asp Ser Thr
165 170 175Ile Ala Asp Phe Leu Ser Arg Ser Leu Ser Asn Asn Ala Tyr
Arg Ser 180 185 190Glu Arg His Leu Arg Lys Arg Ala Leu Ala Tyr Leu
Asn His Ile Ser 195 200 205Ala Glu Lys Glu Ile Thr Ser Asn Ala Cys
Phe Ala Met Lys Asp Val 210 215 220Asn Ser Phe Ala His Lys Gln Ser
Glu Trp Leu Cys His Leu Glu Arg225 230 235 240Ser Leu Trp Arg Asp
Glu Pro Ala Leu Gln Phe His Asp Arg Gln Gln 245 250 255Leu Gly Asn
Glu Val Leu Gly Leu Lys Lys Pro Asp Asp Gln Ser Pro 260 265 270Tyr
Phe Lys Pro Arg Ala Trp Lys Ile Ser Asp Glu Ala Ala Ser Ala 275 280
285Phe Ala Met Met Leu Lys Gly Glu Ser Gly Pro Phe Thr Gln Asp Gln
290 295 300Val Lys Val Gly Phe Glu Ile Cys Gln Glu Gly Glu Leu Leu
Ala Gly305 310 315 320Arg Leu Lys Ile Gln Pro Arg Met Ala Phe Arg
Leu Lys Asn Arg His 325 330 335Asp Ala Asn Arg Ser Gly Thr His Ser
Val Lys Ser Leu Ser Gly Leu 340 345 350Asp Leu Ser Ala Asp Val Gly
Thr Asp Ile Arg Glu Phe Phe Gln Val 355 360 365Pro Val Met Ser Gly
Thr Ser Gly Thr Ser Ser Asp Val Val Ile Ala 370 375 380Ala Arg Tyr
Ala Ala Met His Ala Gly Leu Lys Trp Ser Ala Pro Glu385 390 395
400Leu Thr Ile Asp Gln Ala Lys His Ala Leu Ile Asp Leu Ser Met Asp
405 410 415Phe Phe Arg Arg Asn Gly Pro Ala Val Val Met Ala Leu Arg
Met Asn 420 425 430Ser Leu Arg Lys Asn Gln Gly Leu Pro Tyr Lys Glu
Val Asp Arg Cys 435 440 445Glu Val Phe Thr His Ser Tyr Ala Glu Ile
His Gly Ala Ile Ser Leu 450 455 460Thr Ile Asp Gly Val Asp Pro Ala
Asp Lys Val Glu Val Lys Asn Arg465 470 475 480Leu Tyr Gly Tyr Thr
Leu Asp Ala Lys Ala Thr Leu Met Lys Ile Ala 485 490 495Asp Arg Ser
Ile Arg Arg Gly Val Arg Ser Lys Val Asp Ile Arg Ser 500 505 510Thr
Ser Thr Ser Leu Gln Thr Pro Gln Leu Arg Arg Val Leu Glu Lys 515 520
525Lys Lys Ile Val Gln Lys Val Ala Glu Leu Tyr Ser Glu Met Gly Lys
530 535 540Ala Gly Asn Ser Ala Thr Leu Lys Glu Ala Ile Thr Lys Ser
Ser Val545 550 555 560Lys Glu Leu Leu Val Asn Asp Lys Pro Val Val
Ser Arg Asp Tyr Ala 565
570 575Leu Gly Glu Pro Leu Met Val Arg Ser Leu Arg Phe Ser His Asp
His 580 585 590Glu Ala Thr Ser Ser Phe Gly Ser Ala Gly Lys Thr Pro
Ala Lys Arg 595 600 605Glu Val Asp Thr Leu Cys Asp Asn Ser Thr Ala
Phe Asp Ile Val Met 610 615 620Thr Pro Phe Ser Val Ile Asn Ala Lys
Ala Lys Gly Asp Thr Ile Ser625 630 635 640Glu Met Lys Val Pro His
Arg Pro Lys Trp Lys Gly Leu Pro Ser Val 645 650 655Leu Tyr Lys Val
Thr Ala Ser Val Asp Leu Pro Glu Tyr Ala Val Ala 660 665 670Arg Pro
Gly Phe Gly Asp Ile His Ser Phe Asn Ser Asn Lys Ala Phe 675 680
685Ser Ser Glu Phe Ser Ser Val Arg Asn Ser Leu Ser His Ala Glu Lys
690 695 700Met Gly Phe Ile Glu Asn Ser Leu Lys Pro Tyr Ile Lys His
Asp Pro705 710 715 720Asp Arg Glu Ser Phe Asp Phe Lys His Ser Ile
Asp Glu Leu Ala Asp 725 730 735Ala Gln Cys Met Leu Gln Ser Arg Lys
Pro Asn Ser Thr Leu Arg His 740 745 750Asn Glu Tyr Cys Ala Lys Leu
Glu Leu Trp Asp Ala Lys Ala Ile Glu 755 760 765Val Gly Met Ser Arg
Pro Val Ala Val Ala Thr Leu Ile Glu Phe Asn 770 775 780Leu Glu Met
Leu Ser Ala Ala Arg Tyr Ile Glu Asp Glu Gly Tyr Asp785 790 795
800Gly Lys Leu Ile Thr Asn Phe Leu Glu Arg Gln Leu Ser Trp Phe Gly
805 810 815Gln Asn Ala Ala Leu Asn Lys Glu Val Thr Leu Lys Lys Leu
Trp Gly 820 825 830Leu Pro Phe Asp Glu Arg Lys Ala Val Ala Glu Lys
Val Cys Glu Ala 835 840 845Leu Arg Gln Gly Val Ser Leu Cys Val Tyr
Glu Lys Asn Val Glu Gly 850 855 860Ser Arg Ile Arg Glu Leu Ser Leu
Leu Asn Phe Asn Ala Tyr Asp Ile865 870 875 880Met Arg Gly Ile Glu
Leu Phe Leu Ser Ser Lys Leu Leu Gln Pro Pro 885 890 895Thr Gly Ala
Gly Pro Thr Val Lys Ser Arg Leu 900 905511080DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
51accgctatcg ctgttccggg gactgaaact agacgcgcat ttacagcgca ctgcttttgc
60acataaccct gctgtacctg acgataattg cgcgttaccg attgcggtag gaatgaatta
120aaatattttt tatcatttat gaccatgtgt tgagctttta ttataaaaaa
gattttttga 180gtagtaattc ttatatataa tcatccggag gtggttggta
gcctggctca atcattgagg 240catatttttg caggcaatat attgaatctg
aaaagttaaa gatgatattt tcggtgcagg 300agctatcatg tggagggaaa
agtatgctaa gtcctacgac tcgtaatatg ggggcgagtt 360tatcgcctca
gcctgacgtc agcggggagc taaacaccga agcattgacc tgtattgttg
420agcgtctgga aagtgaaatt atagatggca gctggattca tatcagttac
gaggaaaccg 480atctcgaaat gatgcctttt cttgttgcac aggccaataa
gaagtatcca gagttaaatc 540ttaaatttgt tatgtcagtc catgagcttg
tttcctctat aaaggagacc agaatggaag 600gcgttgaatc tgcccgattt
ctcgtaaata tgggaagttc aggtatccat atttcagtcg 660tcgattttag
agttatggac ggaaagacat cggtgatttt gttcgaacca gcagcgtgta
720gcgcttttgg acctgcactg gcgttgagga ccaaagcagc tcttgaacgt
gaacaactgc 780ctgattgtta ttttgctatg gtcgagctgg acattcaacg
aagctcttct gaatgcggta 840tttttagcct ggcgctcgcc aaaaaacttc
agcttgaatt tatgaactta gtaaaaattc 900atgaagataa tatttgtgaa
cgtctgtgtg gtgaagaacc ttttctcccg tccgataaag 960cagaccgcta
tctgccggtg agtttttaca aacatactca aggcgcacaa cgattaaatg
1020aatatgtgga ggccaatccg gcggcgggaa gcagtatagt aaacaaaaag
aatgaaacgc 108052700PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 52Met Phe Asn Ile Arg Asn Thr Gln
Pro Ser Val Ser Met Gln Ala Ile1 5 10 15Ala Gly Ala Ala Ala Pro Glu
Ala Ser Pro Glu Glu Ile Val Trp Glu 20 25 30Lys Ile Gln Val Phe Phe
Pro Gln Glu Asn Tyr Glu Glu Ala Gln Gln 35 40 45Cys Leu Ala Glu Leu
Cys His Pro Ala Arg Gly Met Leu Pro Asp His 50 55 60Ile Ser Ser Gln
Phe Ala Arg Leu Lys Ala Leu Thr Phe Pro Ala Trp65 70 75 80Glu Glu
Asn Ile Gln Cys Asn Arg Asp Gly Ile Asn Gln Phe Cys Ile 85 90 95Leu
Asp Ala Gly Ser Lys Glu Ile Leu Ser Ile Thr Leu Asp Asp Ala 100 105
110Gly Asn Tyr Thr Val Asn Cys Gln Gly Tyr Ser Glu Ala His Asp Phe
115 120 125Ile Met Asp Thr Glu Pro Gly Glu Glu Cys Thr Glu Phe Ala
Glu Gly 130 135 140Ala Ser Gly Thr Ser Leu Arg Pro Ala Thr Thr Val
Ser Gln Lys Ala145 150 155 160Ala Glu Tyr Asp Ala Val Trp Ser Lys
Trp Glu Arg Asp Ala Pro Ala 165 170 175Gly Glu Ser Pro Gly Arg Ala
Ala Val Val Gln Glu Met Arg Asp Cys 180 185 190Leu Asn Asn Gly Asn
Pro Val Leu Asn Val Gly Ala Ser Gly Leu Thr 195 200 205Thr Leu Pro
Asp Arg Leu Pro Pro His Ile Thr Thr Leu Val Ile Pro 210 215 220Asp
Asn Asn Leu Thr Ser Leu Pro Glu Leu Pro Glu Gly Leu Arg Glu225 230
235 240Leu Glu Val Ser Gly Asn Leu Gln Leu Thr Ser Leu Pro Ser Leu
Pro 245 250 255Gln Gly Leu Gln Lys Leu Trp Ala Tyr Asn Asn Trp Leu
Ala Ser Leu 260 265 270Pro Thr Leu Pro Pro Gly Leu Gly Asp Leu Ala
Val Ser Asn Asn Gln 275 280 285Leu Thr Ser Leu Pro Glu Met Pro Pro
Ala Leu Arg Glu Leu Arg Val 290 295 300Ser Gly Asn Asn Leu Thr Ser
Leu Pro Ala Leu Pro Ser Gly Leu Gln305 310 315 320Lys Leu Trp Ala
Tyr Asn Asn Arg Leu Thr Ser Leu Pro Glu Met Ser 325 330 335Pro Gly
Leu Gln Glu Leu Asp Val Ser His Asn Gln Leu Thr Arg Leu 340 345
350Pro Gln Ser Leu Thr Gly Leu Ser Ser Ala Ala Arg Val Tyr Leu Asp
355 360 365Gly Asn Pro Leu Ser Val Arg Thr Leu Gln Ala Leu Arg Asp
Ile Ile 370 375 380Gly His Ser Gly Ile Arg Ile His Phe Asp Met Ala
Gly Pro Ser Val385 390 395 400Pro Arg Glu Ala Arg Ala Leu His Leu
Ala Val Ala Asp Trp Leu Thr 405 410 415Ser Ala Arg Glu Gly Glu Ala
Ala Gln Ala Asp Arg Trp Gln Ala Phe 420 425 430Gly Leu Glu Asp Asn
Ala Ala Ala Phe Ser Leu Val Leu Asp Arg Leu 435 440 445Arg Glu Thr
Glu Asn Phe Lys Lys Asp Ala Gly Phe Lys Ala Gln Ile 450 455 460Ser
Ser Trp Leu Thr Gln Leu Ala Glu Asp Ala Ala Leu Arg Ala Lys465 470
475 480Thr Phe Ala Met Ala Thr Glu Ala Thr Ser Thr Cys Glu Asp Arg
Val 485 490 495Thr His Ala Leu His Gln Met Asn Asn Val Gln Leu Val
His Asn Ala 500 505 510Glu Lys Gly Glu Tyr Asp Asn Asn Leu Gln Gly
Leu Val Ser Thr Gly 515 520 525Arg Glu Met Phe Arg Leu Ala Thr Leu
Glu Gln Ile Ala Arg Glu Lys 530 535 540Ala Gly Thr Leu Ala Leu Val
Asp Asp Val Glu Val Tyr Leu Ala Phe545 550 555 560Gln Asn Lys Leu
Lys Glu Ser Leu Glu Leu Thr Ser Val Thr Ser Glu 565 570 575Met Arg
Phe Phe Asp Val Ser Gly Val Thr Val Ser Asp Leu Gln Ala 580 585
590Ala Glu Leu Gln Val Lys Thr Ala Glu Asn Ser Gly Phe Ser Lys Trp
595 600 605Ile Leu Gln Trp Gly Pro Leu His Ser Val Leu Glu Arg Lys
Val Pro 610 615 620Glu Arg Phe Asn Ala Leu Arg Glu Lys Gln Ile Ser
Asp Tyr Glu Asp625 630 635 640Thr Tyr Arg Lys Leu Tyr Asp Glu Val
Leu Lys Ser Ser Gly Leu Val 645 650 655Asp Asp Thr Asp Ala Glu Arg
Thr Ile Gly Val Ser Ala Met Asp Ser 660 665 670Ala Lys Lys Glu Phe
Leu Asp Gly Leu Arg Ala Leu Val Asp Glu Val 675 680 685Leu Gly Ser
Tyr Leu Thr Ala Arg Trp Arg Leu Asn 690 695 700532091DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
53gccggaacgc tttaacgcgc ttcgtgaaaa gcaaatatcg gattatgaag acacgtaccg
60gaagctgtat gacgaagtgc tgaaatcgtc cgggctggtc gacgataccg atgcagaacg
120tactatcgga gtaagtgcga tggatagtgc gaaaaaagaa tttctggatg
gcctgcgcgc 180tcttgtggat gaggtgctgg gtagctatct gacagcccgg
tggcgtctta actgagcacg 240atattcaccg caccaggcga atgtggtgcg
gtgaacaaag atattcctgg acaaacaaca 300tcagacagca ctgatgatgc
acaggtgaaa caggggagac ttcttcagtc agggcgtagc 360gcaactcaac
cttttcgacg ataacgcgcc gcgcgccgat agtgcgaagt tgatggaagt
420actggaccat cttaatgcaa aagacgggaa ggggacgctg tacttcgccg
ggtaggggat 480gtcgcaacag tgggctatga agcgagaaat gctttcacct
cggtatacga ccagattttc 540cgatctgcca atagtcaggt aacgggtttg
atcagctctt ccccttgatt tttcacattg 600ccaacggcgc gcttcacggc
gtgccaggta aatttatctg tcggcacggc accatcagta 660attatctcct
ctacctcctt cccgcttatg ccctggcgca tccattttcg cgcggtgcca
720ggtgacagaa cgagaggacg acggtcgtga atgtctacca gacctttatc
agctgcggag 780gtaacaatca ggaatccctc tgcgtcatcg ccgcgctcaa
acggtgtact gccaatggca 840gccatgaata tcggcttccc gtcctttctg
tgaatgaaat acggctgttt cttgtcgcct 900tccttcttcc actcaaacca
accatcagca aacacgatag ctcggccatg ttgccatagc 960ggtttaaaca
ttctgctggt ggccgcagtc tcaacccgtg cattaatcag cggtggttta
1020tcccaccatc cgggcgcaaa tccccagaat accggatcca gatgcagttg
ctcgtcgcgt 1080tcactgagca gcagaacttt ggtaccgggc gccacgttgt
accggcctat aggttcaggg 1140tcataagcga tatcgcgctc ggcttcatcg
gccagatatg ccaggtattc ttcgcgggtc 1200tgtgcttgtg caaagcgtcc
acacatatga aacctccagt cggtcagact gaaagtatag 1260aagaggatat
gcgagtggct gttccggtgt ttcttcgagc gacctcaacc aatgtagaag
1320cttcactatt gggggttgcc actagtagca tcatgttgaa tgtactggcg
tgaaaaaatt 1380ggaatcttga agaaaactct tccccaaaac tataatcaac
gttttgataa tcaatgagtt 1440gtaaaagaca gttactggat ttttttgata
gtaggaagaa tgataatttc aactttatca 1500aatggttgat atgtttttgg
caatgtaatg ctgcgccaca tgcagtggtt cgaagccgca 1560gacctgattg
ttaaaggtat ggaaggcgcg attgccgcga agaccgtgac ctatgacttt
1620gaacgcctga tggaaggcgc taagctgctg aaatgtagtg agtttggtga
cgcgattatc 1680gcgaatatgt aataacgata attgttaaaa acaaaaacgg
ggacttaacg tccccgtttt 1740tattattagt attcgaacgg ttatcaaaac
tttatcaaaa ctccctcaat tcagaccgca 1800atagtagttc atcctttacc
ccgatcgtca tgataacctg agacccccct gaaagctgat 1860agttaccgat
tttgattgta gtggttttct taataatgtg tgatttggct attttttgaa
1920agtgccttac cagatttatt gttattatca ggagaatttt ttaataaaaa
gtgatggttt 1980attgactaca atagtgggta ggttaagtat tttataaaat
tattagcatg ctttatttgc 2040tcttctacaa ggctgctaag aagtgttgaa
atatcagcct gataaagaat a 209154340PRTArtificial SequenceDescription
of Artificial Sequence Synthetic polypeptide 54Met Asn Ile Cys Val
Asn Ser Leu Tyr Arg Leu Ser Ile Pro Gln Phe1 5 10 15His Ser Leu Tyr
Thr Glu Glu Val Ser Asp Glu Ala Leu Thr Leu Leu 20 25 30Phe Ser Ala
Val Glu Asn Gly Asp Gln Asn Cys Ile Asp Leu Leu Cys 35 40 45Asn Leu
Ala Leu Arg Asn Asp Asp Leu Gly His Arg Val Glu Lys Phe 50 55 60Leu
Phe Asp Leu Phe Ser Gly Lys Arg Thr Gly Ser Ser Asp Ile Asp65 70 75
80Lys Lys Ile Asn Gln Ala Cys Leu Val Leu His Gln Ile Ala Asn Asn
85 90 95Asp Ile Thr Lys Asp Asn Thr Glu Trp Lys Lys Leu His Ala Pro
Ser 100 105 110Arg Leu Leu Tyr Met Ala Gly Ser Ala Thr Thr Asp Leu
Ser Lys Lys 115 120 125Ile Gly Ile Ala His Lys Ile Met Gly Asp Gln
Phe Ala Gln Thr Asp 130 135 140Gln Glu Gln Val Gly Val Glu Asn Leu
Trp Cys Gly Ala Arg Met Leu145 150 155 160Ser Ser Asp Glu Leu Ala
Ala Ala Thr Gln Gly Leu Val Gln Glu Ser 165 170 175Pro Leu Leu Ser
Val Asn Tyr Pro Ile Gly Leu Ile His Pro Thr Thr 180 185 190Lys Glu
Asn Ile Leu Ser Thr Gln Leu Leu Glu Lys Ile Ala Gln Ser 195 200
205Gly Leu Ser His Asn Glu Val Phe Leu Val Asn Thr Gly Asp His Trp
210 215 220Leu Leu Cys Leu Phe Tyr Lys Leu Ala Glu Lys Ile Lys Cys
Leu Ile225 230 235 240Phe Asn Thr Tyr Tyr Asp Leu Asn Glu Asn Thr
Lys Gln Glu Ile Ile 245 250 255Glu Ala Ala Lys Ile Ala Gly Ile Ser
Glu Ser Asp Glu Val Asn Phe 260 265 270Ile Glu Met Asn Leu Gln Asn
Asn Val Pro Asn Gly Cys Gly Leu Phe 275 280 285Cys Tyr His Thr Ile
Gln Leu Leu Ser Asn Ala Gly Gln Asn Asp Pro 290 295 300Ala Thr Thr
Leu Arg Glu Phe Ala Glu Asn Phe Leu Thr Leu Ser Val305 310 315
320Glu Glu Gln Ala Leu Phe Asn Thr Gln Thr Arg Arg Gln Ile Tyr Glu
325 330 335Tyr Ser Leu Gln 34055954DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
55gtgagcgatg agacgcttgc gttgttgttt agcgccgtag agaacggtga tcagaattgt
60attgatctgt tatgcaatct tgcgttacac aatgataacc tgggacatag agttgagaaa
120tttctttttg aactctttag cggaaaaaga tcgggctcac cagatataga
taaaaaaatc 180aatcaggctt gccttgtatt acatcaaatt gccaataacg
atataacaaa aaataatact 240gagtggaaaa agctacatac cccttccaga
ttactttata tggccggttc cgcgacaacc 300gacctttcta aaaaaataga
aatagcacat aaaattatgg gcaaccagtt cgctcagaca 360gataaagaac
aggttggagt tgaaaatctt tggtgtggtg tgcgaatgat gtcgtcagat
420gagctggcag ctgcaacgca aggtctggtt caagaatcac cttttctctc
ggtaaactat 480cccattggac ttattcatcc taccaccaaa gaaaatatat
taagcactca gctacttgaa 540aagattgctc aatcaggatt atgtgaaaat
gaaatctttc tgataaatac aggagatcac 600tggcttctct gtttatttta
taaacttgca gaaaaaataa aatgtctcat atttaacagt 660tatcatgatt
taaatgaaaa tactaagcaa gagattatag aagcagcaaa gattgcaggt
720atatcagaaa gcgatgaggt taattttatc gaaattaatt tacagaataa
tgtacccaac 780ggctgtggtc tattttgtta ccatgcaatt caactcttat
cgaatgccgg gcaaaacgat 840cctgttacca cactacgaga atttgcggaa
aatttcttaa cgcttccagt agaggaacaa 900acactattta acacccaaac
ccggcgacaa atatatgaat acagtctcca gtaa 95456189PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
56Met Pro Gly Thr Ile Ser Ser Ser Gly Phe Gly Phe Ser Ile Ala Lys1
5 10 15Gln Pro His Ser Ser Gly Gln Lys Thr Val Ile Asp Gly Phe Phe
Leu 20 25 30Gly Thr Arg Lys Ile Ser Phe Ser Tyr Leu Arg Leu Glu Ser
Glu Leu 35 40 45Met Gln Cys Ile Asn Leu Lys Asn Glu Gly Lys Met Asn
Glu Trp Met 50 55 60Arg Glu Glu Cys Ile Cys Phe Val Ser Arg Asp Val
Asn Lys Gln Leu65 70 75 80Asp Ile Phe Ala Lys Asn Asn Gln Thr Thr
Ile Pro Gly Cys Val Arg 85 90 95Glu Arg Val Phe Gln Arg Ala Ser Phe
His Cys Gly Phe Ser Leu Asp 100 105 110Val Arg Cys Ala Gln Thr Ser
Thr His His Met Ile Leu Asn Ser Leu 115 120 125Tyr Phe Gln Lys Lys
Met Asp Thr Leu Phe Gly Ser Ala Asp Val Glu 130 135 140Val Arg Asn
Gln Cys Val Arg Thr Ala Leu Ser Ser Leu Ala Asp Ile145 150 155
160Phe Phe Glu Arg Asn Val Asn Ser Ile Asp Met Asn Lys Phe Arg Asp
165 170 175Lys Val Tyr Asp Ala Ile Val Gln Glu Ala Gln Arg Thr 180
18557551DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 57acagtaaact cctgtggttt tggatttagc
attacaaaag ccccctattc ttccggacaa 60aaaccattta tagatggttt ctttttaggc
gcgaggaaaa tctcattctc ttatcctcga 120ctggaaagcg agttaataca
gtgcattaac ctgaaaaatg aaggaaaaaa gaatgagtgg 180atgaaggagg
agggtatttg ttttgtttcg cgggatgtca ataaactcct ggatatgttt
240gctaaaaaca accagacaaa catacctgag ggggtccggg agcgggtttt
tcagctcgca 300agtttttatt gcggtttctc gttggatgca agatgcgccc
agacatccac tcatcacatg 360attttaaata gtcagtattt tcagaaaaaa
atggatactc ttttgacttc agtagatata 420aatgtcagaa atcagtgtgt
ccgtacagcg ctaagtagtc tagcggatac tttttttgag 480aataatgtta
acaatataga tatgaataaa ttccgtgaca gggttcataa cactattgta
540caggaggttc a 55158293PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 58Met Leu Ser Pro Tyr Ser
Val Asn Leu Gly Cys Ser Trp Asn Ser Leu1 5 10 15Thr Arg Asn Leu Thr
Ser
Pro Asp Asn Arg Val Leu Ser Ser Val Arg 20 25 30Asp Ala Ala Val His
Ser Asp Asn Gly Ala Gln Val Lys Val Gly Asn 35 40 45Arg Thr Tyr Arg
Val Val Ala Thr Asp Asn Lys Phe Cys Val Thr Arg 50 55 60Glu Ser His
Ser Gly Cys Phe Thr Asn Leu Leu His Arg Leu Gly Trp65 70 75 80Pro
Lys Gly Glu Ile Ser Arg Lys Ile Glu Val Met Leu Asn Ala Ser 85 90
95Pro Val Ser Ala Ala Met Glu Arg Gly Ile Val His Ser Asn Arg Pro
100 105 110Asp Leu Pro Pro Val Asp Tyr Ala Pro Pro Glu Leu Pro Ser
Val Asp 115 120 125Tyr Asn Arg Leu Ser Val Pro Gly Asn Val Ile Gly
Lys Gly Gly Asn 130 135 140Ala Val Val Tyr Glu Asp Ala Glu Asp Ala
Thr Lys Val Leu Lys Met145 150 155 160Phe Thr Thr Ser Gln Ser Asn
Glu Glu Val Thr Ser Glu Val Arg Cys 165 170 175Phe Asn Gln Tyr Tyr
Gly Ala Gly Ser Ala Glu Lys Ile Tyr Gly Asn 180 185 190Asn Gly Asp
Ile Ile Gly Ile Arg Met Asp Lys Ile Asn Gly Glu Ser 195 200 205Leu
Leu Asn Ile Ser Ser Leu Pro Ala Gln Ala Glu His Ala Ile Tyr 210 215
220Asp Met Phe Asp Arg Leu Glu Gln Lys Gly Ile Leu Phe Val Asp
Thr225 230 235 240Thr Glu Thr Asn Ile Leu Tyr Asp Arg Ala Lys Asn
Glu Phe Asn Pro 245 250 255Ile Asp Ile Ser Ser Tyr Asn Val Ser Asp
Arg Ser Trp Ser Glu Ser 260 265 270Gln Ile Met Gln Ser Tyr His Gly
Gly Lys Gln Asp Leu Ile Ser Val 275 280 285Val Leu Ser Lys Ile
29059795DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 59atgctgagcc cgtatagcgt gaacctgggc
tgcagctgga acagcctgac ccgcaacctg 60accagcccgg ataaccgcgt gctgagcagc
gtgcgcgatg cggcggtgca tagcgataac 120ggcgcgcagg tgaaagtggg
caaccgcacc tatcgcgtgg tggcgaccga taacaaattt 180tgcgtgaccc
gcgaaagcca tagcggctgc tttaccaacc tgctgcatcg cctgggctgg
240ccgaaaggcg aaattagccg caaaattgaa gtgatgctga acgcgagccc
ggtgagcgcg 300gcgatggaac gcggcattgt gcatagcaac cgcccggatc
tgccgccggt ggattatgcg 360ccgccggaac tgccgagcgt ggattataac
cgcctgagcg tgccgggcaa cgtgattggc 420aaaggcggca acgcggtggt
gtatgaagat gcggaagatg cgaccaaagt gctgaaaatg 480tttaccacca
gccagagcaa cgaagaagtg accagcgaag tgcgctgctt taaccagtat
540tatggcgcgg gcagcgcgga aaaaatttat ggcaacaacg gcgatattat
tggcattcgc 600atggataaaa ttaacggcga aagcctgctg aacattagca
gcctgccggc gcaggcggaa 660catgcgattt atgatatgtt tgatcgcctg
gaacagaaag gcattctgtt tgtggatacc 720accgaaacca acattctgta
tgatcgcgcg catggcggca aacaggatct gattagcgtg 780gtgctgagca aaatt
79560336PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 60Met Ile Pro Pro Leu Asn Arg Tyr Val Pro Ala
Leu Ser Lys Asn Glu1 5 10 15Leu Val Lys Thr Val Thr Asn Arg Asp Ile
Gln Phe Thr Ser Phe Asn 20 25 30Gly Lys Asp Tyr Pro Leu Cys Phe Leu
Asp Glu Lys Thr Pro Leu Leu 35 40 45Phe Gln Trp Phe Glu Arg Asn Pro
Ala Arg Phe Gly Lys Asn Asp Ile 50 55 60Pro Ile Ile Asn Thr Glu Lys
Asn Pro Tyr Leu Asn Asn Ile Ile Lys65 70 75 80Ala Ala Thr Ile Glu
Lys Glu Arg Leu Ile Gly Ile Phe Val Asp Gly 85 90 95Asp Phe Phe Pro
Gly Gln Lys Asp Ala Phe Ser Lys Leu Glu Tyr Asp 100 105 110Tyr Glu
Asn Ile Lys Val Ile Tyr Arg Asn Asp Ile Asp Phe Ser Met 115 120
125Tyr Asp Lys Lys Leu Ser Glu Ile Tyr Met Glu Asn Ile Ser Lys Gln
130 135 140Glu Ser Met Pro Glu Glu Lys Arg Asp Cys His Leu Leu Gln
Leu Leu145 150 155 160Lys Lys Glu Leu Ser Asp Ile Gln Glu Gly Asn
Asp Ser Leu Ile Lys 165 170 175Ser Tyr Leu Leu Asp Lys Gly His Gly
Trp Phe Asp Phe Tyr Arg Asn 180 185 190Met Ala Met Leu Lys Ala Gly
Gln Leu Phe Leu Glu Ala Asp Lys Val 195 200 205Gly Cys Tyr Asp Leu
Ser Thr Asn Ser Gly Cys Ile Tyr Leu Asp Ala 210 215 220Asp Met Ile
Ile Thr Glu Lys Leu Gly Gly Ile Tyr Ile Pro Asp Gly225 230 235
240Ile Ala Val His Val Glu Arg Ile Asp Gly Arg Ala Ser Met Glu Asn
245 250 255Gly Ile Ile Ala Val Asp Arg Asn Asn His Pro Ala Leu Leu
Ala Gly 260 265 270Leu Glu Ile Met His Thr Lys Phe Asp Ala Asp Pro
Tyr Ser Asp Gly 275 280 285Val Cys Asn Gly Ile Arg Lys His Phe Asn
Tyr Ser Leu Asn Glu Asp 290 295 300Tyr Asn Ser Phe Cys Asp Phe Ile
Glu Phe Lys His Asp Asn Ile Ile305 310 315 320Met Asn Thr Ser Gln
Phe Thr Gln Ser Ser Trp Ala Arg His Val Gln 325 330
33561540DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 61atgttatctc cattaaatgt tcttcaattt
aatttcagag gagagaccgc tttatcagat 60agtgctcctc tccagactgt ttcctttgct
ggaaaagatt attctatgga acccattgat 120gaaaaaacac ccattctttt
tcagtggttt gaagcaaggc cagagcgata cggaaaaggt 180gaagtaccga
tattgaatac caaagagcat ccgtatttga gcaatattat aaatgctgca
240aaaatagaaa atgagcgcgt aataggagta ctggtagacg gagactttac
ttatgagcaa 300agaaaagaat ttctcagtct tgaagatgaa catcaaaata
taaagataat atatcgggaa 360aatgttgatt tcagtatgta tgataaaaaa
ctgtctgata tttatcttga aaatattcat 420gaacaagaat catatccagc
gagtgagaga gataattatc tgttaggctt attaagagaa 480gagttaaaaa
atattccata cggaaaggac tctttgattg aatcatatgc agaaaaaaga
54062330PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 62Met Lys Ile Pro Ser Leu Gln Ser Asn Phe Asn
Phe Ser Ala Pro Ala1 5 10 15Gly Tyr Ser Ala Pro Ile Ala Pro Asn Arg
Ala Glu Asn Ala Tyr Ala 20 25 30Asp Tyr Val Leu Asp Ile Gly Lys Arg
Ile Pro Leu Ser Ala Ala Asp 35 40 45Leu Ser Asn Val Tyr Glu Ser Val
Ile Arg Ala Val His Asp Ser Arg 50 55 60Ser Arg Leu Ile Asp Gln His
Thr Val Asp Met Ile Gly Asn Thr Val65 70 75 80Leu Asp Ala Leu Ser
Arg Ser Gln Thr Phe Arg Asp Ala Val Ser Tyr 85 90 95Gly Ile His Asn
Glu Lys Val His Ile Gly Cys Ile Lys Tyr Arg Asn 100 105 110Glu Tyr
Glu Leu Asn Glu Glu Ser Ser Val Lys Ile Asp Asp Ile Gln 115 120
125Ser Leu Thr Cys Asn Glu Leu Tyr Glu Tyr Asp Val Gly Gln Glu Pro
130 135 140Ile Phe Pro Ile Cys Glu Ala Gly Glu Asn Asp Asn Glu Glu
Pro Tyr145 150 155 160Val Ser Phe Ser Val Ala Pro Asp Thr Asp Ser
Tyr Glu Met Pro Ser 165 170 175Trp Gln Glu Gly Leu Ile His Glu Ile
Ile His His Val Thr Gly Ser 180 185 190Ser Asp Pro Ser Gly Asp Ser
Asn Ile Glu Leu Gly Pro Thr Glu Ile 195 200 205Leu Ala Arg Arg Val
Ala Gln Glu Leu Gly Trp Ser Val Pro Asp Phe 210 215 220Lys Gly Tyr
Ala Glu Pro Glu Arg Glu Ala His Leu Arg Leu Arg Asn225 230 235
240Leu Asn Ala Leu Arg Gln Ala Ala Met Arg His Glu Glu Asn Glu Arg
245 250 255Ala Phe Phe Glu Arg Leu Gly Thr Ile Ser Asp Arg Tyr Glu
Ala Ser 260 265 270Pro Asp Phe Thr Glu Tyr Ser Ala Val Ser Asn Ile
Gly Tyr Gly Phe 275 280 285Ile Gln Gln His Asp Phe Pro Gly Leu Ala
Ile Asn Asp Asn Leu Gln 290 295 300Asp Ala Asn Gln Ile Gln Leu Tyr
His Gly Ala Pro Tyr Ile Phe Thr305 310 315 320Phe Gly Asp Val Asp
Lys His Asn Gln Arg 325 330631464DNAArtificial SequenceDescription
of Artificial Sequence Synthetic polynucleotide 63gaattcatcc
ggtcaaacgg cttctttttg caggaaagga atatgagtta aaggtcattg 60atgaaaaaac
gcctattatc ctgctctact tgagggattg tcttttatgc acagaagagt
120agatgctcat ccatattatg atggtttagg taaagggata aagaaatatt
ttgattttac 180tcaattacat gattacaatc atttttatga ctttattgag
tttaaacatc caaatattat 240tatgaacaca agtcagtata caggcagttc
atggtaaatg gtttttacat agtttattct 300gttgtaataa atgattagca
tggtattagg tatcaacatg aaaattccct cactccagcc 360cagcttcaac
tttttcgccc cagcaggata ctctgctgcc gttgctccca atcgttcgga
420caatgcctat gctgattacg tattggatat aggcaagcga ataccacttt
ccgcggaaga 480tttaggcaac ctatatgaaa atgtcattcg cgccgttcgt
gacagccgta gcaagctcat 540agatcagcat acggtcgata tgattggtaa
cactatactt gatgctttga gccgatcaca 600aacctttcgt gatgccgtaa
gctatggcat tcataataag gaggtacaca ttggttgcat 660taaatacaga
aacgaatacg agctcaacgg agaatccccc gtcaaagttg atgatattca
720atcactaacc tgtaccgaat tatatgaata cgatgtcggg caagaaccaa
ttttacccat 780ttgcgaggca ggagaaaacg ataacgaaga gccttatgtc
agttttagtg ttgcgccaga 840tactgactct tatgagatgc catcgtggca
ggaagggctg attcacgaga ttattcatca 900tgtgactgga gctagcgatc
cgtctggaga tagtaatata gagctaggac ccacggagat 960tctcgcacgt
cgtgtcgctc aagagctggg atggactgtc cccgacttca taggatatgc
1020agagccagat cgtgaagctc atcttagggg acgtaacctg aatgcccttc
gacaggcggc 1080catgcgacat gaagataatg agaggacttt cttcgaaagg
ctgggtatga tcagtgatcg 1140atatgaggcg agtcctgatt tcacagagta
ttccgctgtg tctaacatag aatatggatt 1200tatccagcaa catgattttc
ccgggttggc tatcgacgat aatttacagg atgcaaatca 1260gatccaactc
tatcatggag caccttatat ctttacattc ggggatgtgg acaaacacaa
1320tcagcgctga cgcgtctttg cagcgacaca aggctactac tcttgcattt
taacggagtt 1380gatgatggaa aatcgtgcaa ccttgtatgt aaaggcgaaa
aaccaaattt tacggtagta 1440agtgagcctg gcgggaatgg tacc
146464558PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 64Met Pro Ile Gly Asn Leu Gly His Asn Pro Asn
Val Asn Asn Ser Ile1 5 10 15Pro Pro Ala Pro Pro Leu Pro Ser Gln Thr
Asp Gly Ala Gly Gly Arg 20 25 30Gly Gln Leu Ile Asn Ser Thr Gly Pro
Leu Gly Ser Arg Ala Leu Phe 35 40 45Thr Pro Val Arg Asn Ser Met Ala
Asp Ser Gly Asp Asn Arg Ala Ser 50 55 60Asp Val Pro Gly Leu Pro Val
Asn Pro Met Arg Leu Ala Ala Ser Glu65 70 75 80Ile Thr Leu Asn Asp
Gly Phe Glu Val Leu His Asp His Gly Pro Leu 85 90 95Asp Thr Leu Asn
Arg Gln Ile Gly Ser Ser Val Phe Arg Val Glu Thr 100 105 110Gln Glu
Asp Gly Lys His Ile Ala Val Gly Gln Arg Asn Gly Val Glu 115 120
125Thr Ser Val Val Leu Ser Asp Gln Glu Tyr Ala Arg Leu Gln Ser Ile
130 135 140Asp Pro Glu Gly Lys Asp Lys Phe Val Phe Thr Gly Gly Arg
Gly Gly145 150 155 160Ala Gly His Ala Met Val Thr Val Ala Ser Asp
Ile Thr Glu Ala Arg 165 170 175Gln Arg Ile Leu Glu Leu Leu Glu Pro
Lys Gly Thr Gly Glu Ser Lys 180 185 190Gly Ala Gly Glu Ser Lys Gly
Val Gly Glu Leu Arg Glu Ser Asn Ser 195 200 205Gly Ala Glu Asn Thr
Thr Glu Thr Gln Thr Ser Thr Ser Thr Ser Ser 210 215 220Leu Arg Ser
Asp Pro Lys Leu Trp Leu Ala Leu Gly Thr Val Ala Thr225 230 235
240Gly Leu Ile Gly Leu Ala Ala Thr Gly Ile Val Gln Ala Leu Ala Leu
245 250 255Thr Pro Glu Pro Asp Ser Pro Thr Thr Thr Asp Pro Asp Ala
Ala Ala 260 265 270Ser Ala Thr Glu Thr Ala Thr Arg Asp Gln Leu Thr
Lys Glu Ala Phe 275 280 285Gln Asn Pro Asp Asn Gln Lys Val Asn Ile
Asp Glu Leu Gly Asn Ala 290 295 300Ile Pro Ser Gly Val Leu Lys Asp
Asp Val Val Ala Asn Ile Glu Glu305 310 315 320Gln Ala Lys Ala Ala
Gly Glu Glu Ala Lys Gln Gln Ala Ile Glu Asn 325 330 335Asn Ala Gln
Ala Gln Lys Lys Tyr Asp Glu Gln Gln Ala Lys Arg Gln 340 345 350Glu
Glu Leu Lys Val Ser Ser Gly Ala Gly Tyr Gly Leu Ser Gly Ala 355 360
365Leu Ile Leu Gly Gly Gly Ile Gly Val Ala Val Thr Ala Ala Leu His
370 375 380Arg Lys Asn Gln Pro Val Glu Gln Thr Thr Thr Thr Thr Thr
Thr Thr385 390 395 400Thr Thr Thr Ser Ala Arg Thr Val Glu Asn Lys
Pro Ala Asn Asn Thr 405 410 415Pro Ala Gln Gly Asn Val Asp Thr Pro
Gly Ser Glu Asp Thr Met Glu 420 425 430Ser Arg Arg Ser Ser Met Ala
Ser Thr Ser Ser Thr Phe Phe Asp Thr 435 440 445Ser Ser Ile Gly Thr
Val Gln Asn Pro Tyr Ala Asp Val Lys Thr Ser 450 455 460Leu His Asp
Ser Gln Val Pro Thr Ser Asn Ser Asn Thr Ser Val Gln465 470 475
480Asn Met Gly Asn Thr Asp Ser Val Val Tyr Ser Thr Ile Gln His Pro
485 490 495Pro Arg Asp Thr Thr Asp Asn Gly Ala Arg Leu Leu Gly Asn
Pro Ser 500 505 510Ala Gly Ile Gln Ser Thr Tyr Ala Arg Leu Ala Leu
Ser Gly Gly Leu 515 520 525Arg His Asp Met Gly Gly Leu Thr Gly Gly
Ser Asn Ser Ala Val Asn 530 535 540Thr Ser Asn Asn Pro Pro Ala Pro
Gly Ser His Arg Phe Val545 550 555651689DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
65taaataaaag gatatatgta tgcctattgg taaccttggt aataatgtaa atagcaatca
60tttaattccc cctgcgccgc cactaccttc acaaacagac ggcgcggcac ggggaggaac
120tggtcatcta attagctcta caggagcatt aggatctcgt tcattgtttt
ctcccctgag 180aaattctgtg gctgattctg tcgattccag agatattcca
ggacttcctg caaacccatc 240gaggcttgct gcagctacat ctgagacatg
cttgcttgga ggatttgaag ttctccatga 300taaggggcca cttgatactc
ttaatcagca aattggaccc tctgcatttc gtgttgaagc 360gcaggcagat
ggtactcatg ccgctattgg agaaaaaaat ggtttggagg ttagcgttgc
420attaagtcct caagaattgc aaagcttgca atctattgat attgagggga
aaaacagatt 480tgtttttacc gggggacgtg gcggtagtgg gcatccgatg
gtcactgtcg catcagatat 540cgcggaagct cgtatgaaaa tactggccaa
attagaccca gacaatcatg gaggacgtca 600acccaaggac gttgatacgc
gttctgttgg tgttggcagc gcttcgggaa tagatgatgg 660cgttgttagc
gaaacccata cttcaacaac aaattccagc gttcgctcag atcctaaatt
720ctgggtttct gtcggcgcaa ttgctgctgg tttagcggga ctggcggcaa
ctggtattgc 780acaggcgttg gctttgacac cggaaccgga tgatcctaca
accaccgatc ctgatcaggc 840cgcaaatgct gcagaaagtg caacaaaaga
tcagttaacg caagaagcat tcaagaaccc 900tgagaaccag aaagttaaca
tcgatgcgaa cggaaatgct attccgtctg gggaattaaa 960agatgatatt
gttgagcaaa tagcacaaca agctaaagag gctggtgagg tggccagaca
1020gcaggctgtt gaaagcaatg cacaggcgca gcagcgatat gaggatcagt
atgccagacg 1080tcaggaggaa ttacagcttt catcgggtat tggttacggc
ctcagcagtg cattgattgg 1140tgctggggga attggtgctg gtgtaacgac
tgcgctccat agacgaaatc agccggcaga 1200acagacaact actacaacaa
cacatacggt agtgcagcag cagaccggag ggaatacccc 1260agcacaaggt
ggcactgatg ccacaagagc agaagatgct tctctgaata gacgtgattc
1320gcaggggagt gttgcatcga cacactggtc agattcctct agcgaagtgg
ttaatccata 1380tgctgaagtt ggggagcctc ggaatagtct atcgactcgt
cagcaagaag agcatattta 1440cgatgaggtc gctgcagatc ctgtttatag
cgtcattcag aatttttcac ggaatgctcc 1500agttaccgga aggttaatgg
gaagcccagg gcaaggtatc caaagtactt atgcgcttct 1560ggcaaacagc
gctggattgc gtttaggtat gggaggatta acggggagtg gcgagagcgc
1620agtaaatact gcaaatgcaa atgccgcacc aacgccggga ccagtacgtt
tcgtttaaat 1680atatctgtg 168966358PRTArtificial SequenceDescription
of Artificial Sequence Synthetic polypeptide 66Met Gln Asn Lys Ile
Lys Gln Leu Lys Asn Tyr Ala Val Tyr Asp Asp1 5 10 15Ile Glu Gly Phe
Leu Ile Asn Lys Asp Ile Arg Ser Ser Ser Gly Asn 20 25 30Ser Asn Tyr
Met Met Pro Ser Ser Thr Arg Arg Val Ser Asn Thr Arg 35 40 45Lys Asn
Tyr Glu Gly Asp Ile Lys Gln Phe Phe Ser Val Ile Lys Gly 50 55 60Lys
Asp Val Lys Ser Leu Val Pro Asp Asp Leu Val Val Ser Lys Ser65 70 75
80Glu Leu Ser Asn Tyr Val Lys Tyr Leu Gln Glu Lys Gly Leu Val Asn
85 90 95Asn Ser Ile Asn Arg Lys Met Thr Ser Leu Lys Met Leu Tyr Thr
Tyr 100 105 110Leu Glu His Asp Tyr Lys Asp Tyr Ile Asp Leu Ser
Val Phe Asn Thr 115 120 125Val Glu Arg Leu Lys Thr Val Thr Lys Asn
Trp Asp Lys Thr Thr Gln 130 135 140Thr Glu Ala Glu Arg Ile Ala Gln
Asp Met Tyr Ile Asn Glu Arg Gln145 150 155 160Lys Pro Leu Met Lys
Lys Leu Phe Val Lys Phe Ala Ile Arg Thr Ser 165 170 175Phe Arg Val
Ser Ala Ile Leu Arg Val Arg Trp Lys Asp Ile Gln Leu 180 185 190Asp
Glu Ser Thr Gly His Tyr Ile Val Thr Val Ile Asp Lys Gly Ser 195 200
205Gln Val Val Ser Thr Gly Ile Asn Gln Val Phe Tyr Glu Glu Leu Leu
210 215 220Gln Leu Lys Glu Glu Asp Asp Ser Glu Thr Glu Leu Val Phe
Gln Gly225 230 235 240Leu Ser Glu Gln Ser Leu Arg His Ser Leu Lys
Arg Ser Lys Lys Arg 245 250 255Leu Gly Ile Pro Pro Glu Arg Glu Leu
Val Leu His Ser Phe Lys Gly 260 265 270Val Gly Ile Asp Tyr Val Tyr
Glu Asn Ser Gly His Asp Leu Leu Ala 275 280 285Ala Lys Glu Gln Gly
Asn His Lys Asn Thr Leu Thr Thr Glu Arg Tyr 290 295 300Met Ser Arg
Lys Ile Asn Ile Ala Asn Ser Ala Gly Val Thr Met Asp305 310 315
320Glu Lys Ile Asp Leu Asn Pro Leu Tyr Glu Ala Thr Gln Glu Asp Phe
325 330 335Ile Ser Phe Phe Glu Asn Ala Asp Leu Val Thr Leu Lys Lys
Phe Ile 340 345 350Lys His Val Asn Glu Arg 35567867DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
67atgattgggc caatatcaca aataaacagc ttcggtggct tatcagaaaa agagacccgt
60tctttaatca gtaatgaaga gcttaaaaat atcataatac agttggaaac tgatatagcg
120gatggatcct ggttccataa aaattattca cgcctggata tagaagtcat
gcccgcatta 180gtaattcagg cgaacaataa atatccggaa atgaatctta
attttgttac atctccccag 240gacctttcga tagaaataaa aaatgtcata
gaaaatggag ttggatcttc ccgcttcata 300attaacatgg gggagggtgg
aatacatttc agtgtaattg attacaaaca tataaatggg 360aaaacatctc
tgatattatt tgaaccagta aactttaata gtatggggcc agcgatactg
420gcaataagta caaaaacggc cattgaacgt tatcaattac ctgattgcca
tttttccatg 480gtggaaatgg atattcagcg aagctcatct gaatgtggta
tttttagttt ggcactggca 540aaaaaacttt acaccgagag agatagcctg
ttgaaaatac atgaagataa tataaaaggt 600atattaagtg atagtgaaaa
tcctttaccc cacaataagt tggatccgta tctcccggta 660actttttaca
aacatactca aggtaaaaaa cgtcttaatg aatatttaaa tactaacccg
720cagggagttg gtactgttgt taacaaaaaa aatgaaacca tctttaatag
gtttgataac 780aataaatcca ttatagatgg aaaggaatta tcagtttcgg
tacataaaaa gagaatagct 840gaatataaaa cacttctcaa agtataa
86768288PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 68Met Ile Gly Pro Ile Ser Gln Ile Asn Ile Ser
Gly Gly Leu Ser Glu1 5 10 15Lys Glu Thr Ser Ser Leu Ile Ser Asn Glu
Glu Leu Lys Asn Ile Ile 20 25 30Thr Gln Leu Glu Thr Asp Ile Ser Asp
Gly Ser Trp Phe His Lys Asn 35 40 45Tyr Ser Arg Met Asp Val Glu Val
Met Pro Ala Leu Val Ile Gln Ala 50 55 60Asn Asn Lys Tyr Pro Glu Met
Asn Leu Asn Leu Val Thr Ser Pro Leu65 70 75 80Asp Leu Ser Ile Glu
Ile Lys Asn Val Ile Glu Asn Gly Val Arg Ser 85 90 95Ser Arg Phe Ile
Ile Asn Met Gly Glu Gly Gly Ile His Phe Ser Val 100 105 110Ile Asp
Tyr Lys His Ile Asn Gly Lys Thr Ser Leu Ile Leu Phe Glu 115 120
125Pro Ala Asn Phe Asn Ser Met Gly Pro Ala Met Leu Ala Ile Arg Thr
130 135 140Lys Thr Ala Ile Glu Arg Tyr Gln Leu Pro Asp Cys His Phe
Ser Met145 150 155 160Val Glu Met Asp Ile Gln Arg Ser Ser Ser Glu
Cys Gly Ile Phe Ser 165 170 175Phe Ala Leu Ala Lys Lys Leu Tyr Ile
Glu Arg Asp Ser Leu Leu Lys 180 185 190Ile His Glu Asp Asn Ile Lys
Gly Ile Leu Ser Asp Gly Glu Asn Pro 195 200 205Leu Pro His Asp Lys
Leu Asp Pro Tyr Leu Pro Val Thr Phe Tyr Lys 210 215 220His Thr Gln
Gly Lys Lys Arg Leu Asn Glu Tyr Leu Asn Thr Asn Pro225 230 235
240Gln Gly Val Gly Thr Val Val Asn Lys Lys Asn Glu Thr Ile Val Asn
245 250 255Arg Phe Asp Asn Asn Lys Ser Ile Val Asp Gly Lys Glu Leu
Ser Val 260 265 270Ser Val His Lys Lys Arg Ile Ala Glu Tyr Lys Thr
Leu Leu Lys Val 275 280 28569867DNAArtificial SequenceDescription
of Artificial Sequence Synthetic polynucleotide 69atgatcggac
caatatcaca aataaatatc tccggtggct tatcagaaaa agagaccagt 60tctttaatca
gtaatgaaga gcttaaaaat atcataacac agttggaaac tgatatatcg
120gatggatcct ggttccataa aaattattca cgtatggatg tagaagtcat
gcccgcattg 180gtaatccagg cgaacaataa atatccggaa atgaatctta
atcttgttac atctccattg 240gacctttcaa tagaaataaa aaacgtcata
gaaaatggag ttagatcttc ccgcttcata 300attaacatgg gggaaggtgg
aatacatttc agtgtaattg attacaaaca tataaatggg 360aaaacatctc
tgatattgtt tgaaccagca aactttaaca gtatggggcc agcgatgctg
420gcaataagga caaaaacggc tattgaacgt tatcaattac ctgattgcca
tttctccatg 480gtggaaatgg atattcagcg aagctcatct gaatgtggta
tttttagttt tgcactggca 540aaaaaacttt acatcgagag agatagcctg
ttgaaaatac atgaagataa tataaaaggt 600atattaagtg atggtgaaaa
tcctttaccc cacgataagt tggacccgta tctcccggta 660actttttaca
aacatactca aggtaaaaaa cgtcttaatg aatatttaaa tactaacccg
720cagggagttg gtactgttgt taacaaaaaa aatgaaacca tcgttaatag
atttgataac 780aataaatcca ttgtagatgg aaaggaatta tcagtttcgg
tacataaaaa gagaatagct 840gaatataaaa cacttctcaa agtataa
86770219PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 70Met Lys Ile Ser Ser Phe Ile Ser Thr Ser Leu
Pro Leu Pro Ala Ser1 5 10 15Val Ser Gly Ser Ser Ser Val Gly Glu Met
Ser Gly Arg Ser Val Ser 20 25 30Gln Gln Lys Ser Asp Gln Tyr Ala Asn
Asn Leu Ala Gly Arg Thr Glu 35 40 45Ser Pro Gln Gly Ser Ser Leu Ala
Ser Arg Ile Ile Glu Arg Leu Ser 50 55 60Ser Met Ala His Ser Val Ile
Gly Phe Ile Gln Arg Met Phe Ser Glu65 70 75 80Gly Ser His Lys Pro
Val Val Thr Pro Ala Leu Thr Pro Ala Gln Met 85 90 95Pro Ser Pro Thr
Ser Phe Ser Asp Ser Ile Lys Gln Leu Ala Ala Glu 100 105 110Thr Leu
Pro Lys Tyr Met Gln Gln Leu Ser Ser Leu Asp Ala Glu Thr 115 120
125Leu Gln Lys Asn His Asp Gln Phe Ala Thr Gly Ser Gly Pro Leu Arg
130 135 140Gly Ser Ile Thr Gln Cys Gln Gly Leu Met Gln Phe Cys Gly
Gly Glu145 150 155 160Leu Gln Ala Glu Ala Ser Ala Ile Leu Asn Thr
Pro Val Cys Gly Ile 165 170 175Pro Phe Ser Gln Trp Gly Thr Val Gly
Gly Ala Ala Ser Ala Tyr Val 180 185 190Ala Ser Gly Val Asp Leu Thr
Gln Ala Ala Asn Glu Ile Lys Gly Leu 195 200 205Gly Gln Gln Met Gln
Gln Leu Leu Ser Leu Met 210 215711152DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
71gaattcccca actttgacac cgataaccgg ttcaatagta tctggaatag acagcgaaag
60ttgttgaaat aattgagtga tagcttgttc aaatgaatac atttgatctc ctaatagtta
120gataaaatat caacttaacc aaagcactct cggcagacca tcaattttag
cctataattt 180ttagttttta ttttgtctaa tataacaaca aaaacagcag
cggtttttta tataaccacc 240ggctattttc ccactaagat aaccttgttt
taatagccaa gggaataaat agtcatgaaa 300atatcatcat ttatttctac
atcactgccc ctgccggcat cagtgtcagg atctagcagc 360gtaggagaaa
tgtctgggcg ctcagtctca cagcaaaaaa gtgatcaata tgcaaacaat
420ctggccgggc gcactgaaag ccctcagggt tccagcttag ccagccgtat
cattgagagg 480ttatcatcaa tggcccactc tgtgattgga tttatccaac
gcatgttctc ggaggggagc 540cataaaccgg tggtgacacc agcactcacg
cctgcacaaa tgccaagccc tacgtctttc 600agtgatagta tcaagcaact
tgctgctgag acgctgccaa aatacatgca gcagttgagt 660agcttggatg
cagagacgct gcagaaaaat catgaccagt tcgccacggg cagcggccct
720cttcgtggca gtatcactca atgccaaggg ctgatgcagt tttgtggtgg
ggaattgcaa 780gctgaggcca gtgccatttt aaacacgcct gtttgtggta
ttcccttctc gcagtgggga 840actgttggtg gggcggccag cgcgtacgtc
gccagtggcg ttgatctaac gcaggcagca 900aatgagatca aagggctggg
gcaacagatg cagcaattac tgtcattgat gtgatatgga 960taaaaacaag
ggggtagtgt ttcccccttt ttctatcaat attgcgaata tcttcgtccc
1020tgatctttca ggggcgaatc gttttttagc atgctcattg ttagaatttc
tgacttatct 1080ctcttctgta ttactactca tactctggaa aatcctgagc
atttatatct atggattgat 1140gcagcactcg ag 115272545PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
72Met Leu Pro Ile Asn Asn Asn Phe Ser Leu Pro Gln Asn Ser Phe Tyr1
5 10 15Asn Thr Ile Ser Gly Thr Tyr Ala Asp Tyr Phe Ser Ala Trp Asp
Lys 20 25 30Trp Glu Lys Gln Ala Leu Pro Gly Glu Glu Arg Asp Glu Ala
Val Ser 35 40 45Arg Leu Lys Glu Cys Leu Ile Asn Asn Ser Asp Glu Leu
Arg Leu Asp 50 55 60Arg Leu Asn Leu Ser Ser Leu Pro Asp Asn Leu Pro
Ala Gln Ile Thr65 70 75 80Leu Leu Asn Val Ser Tyr Asn Gln Leu Thr
Asn Leu Pro Glu Leu Pro 85 90 95Val Thr Leu Lys Lys Leu Tyr Ser Ala
Ser Asn Lys Leu Ser Glu Leu 100 105 110Pro Val Leu Pro Pro Ala Leu
Glu Ser Leu Gln Val Gln His Asn Glu 115 120 125Leu Glu Asn Leu Pro
Ala Leu Pro Asp Ser Leu Leu Thr Met Asn Ile 130 135 140Ser Tyr Asn
Glu Ile Val Ser Leu Pro Ser Leu Pro Gln Ala Leu Lys145 150 155
160Asn Leu Arg Ala Thr Arg Asn Phe Leu Thr Glu Leu Pro Ala Phe Ser
165 170 175Glu Gly Asn Asn Pro Val Val Arg Glu Tyr Phe Phe Asp Arg
Asn Gln 180 185 190Ile Ser His Ile Pro Glu Ser Ile Leu Asn Leu Arg
Asn Glu Cys Ser 195 200 205Ile His Ile Ser Asp Asn Pro Leu Ser Ser
His Ala Leu Pro Ala Leu 210 215 220Gln Arg Leu Thr Ser Ser Pro Asp
Tyr His Gly Pro Arg Ile Tyr Phe225 230 235 240Ser Met Ser Asp Gly
Gln Gln Asn Thr Leu His Arg Pro Leu Ala Asp 245 250 255Ala Val Thr
Ala Trp Phe Pro Glu Asn Lys Gln Ser Asp Val Ser Gln 260 265 270Ile
Trp His Ala Phe Glu His Glu Glu His Ala Asn Thr Phe Ser Ala 275 280
285Phe Leu Asp Arg Leu Ser Asp Thr Val Ser Ala Arg Asn Thr Ser Gly
290 295 300Phe Arg Glu Gln Val Ala Ala Trp Leu Glu Lys Leu Ser Ala
Ser Ala305 310 315 320Glu Leu Arg Gln Gln Ser Phe Ala Val Ala Ala
Asp Ala Thr Glu Ser 325 330 335Cys Glu Asp Arg Val Ala Leu Thr Trp
Asn Asn Leu Arg Lys Thr Leu 340 345 350Leu Val His Gln Ala Ser Glu
Gly Leu Phe Asp Asn Asp Thr Gly Ala 355 360 365Leu Leu Ser Leu Gly
Arg Glu Met Phe Arg Leu Glu Ile Leu Glu Asp 370 375 380Ile Ala Arg
Asp Lys Val Arg Thr Leu His Phe Val Asp Glu Ile Glu385 390 395
400Val Tyr Leu Ala Phe Gln Thr Met Leu Ala Glu Lys Leu Gln Leu Ser
405 410 415Thr Ala Val Lys Glu Met Arg Phe Tyr Gly Val Ser Gly Val
Thr Ala 420 425 430Asn Asp Leu Arg Thr Ala Glu Ala Met Val Arg Ser
Arg Glu Glu Asn 435 440 445Glu Phe Thr Asp Trp Phe Ser Leu Trp Gly
Pro Trp His Ala Val Leu 450 455 460Lys Arg Thr Glu Ala Asp Arg Trp
Ala Gln Ala Glu Glu Gln Lys Tyr465 470 475 480Glu Met Leu Glu Asn
Glu Tyr Pro Gln Arg Val Ala Asp Arg Leu Lys 485 490 495Ala Ser Gly
Leu Ser Gly Asp Ala Asp Ala Glu Arg Glu Ala Gly Ala 500 505 510Gln
Val Met Arg Glu Thr Glu Gln Gln Ile Tyr Arg Gln Leu Thr Asp 515 520
525Glu Val Leu Ala Leu Arg Leu Pro Glu Asn Gly Ser Gln Leu His His
530 535 540Ser545731635DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 73atgctgccga
ttaacaacaa ctttagcctg ccgcagaaca gcttttataa caccattagc 60ggcacctatg
cggattattt tagcgcgtgg gataaatggg aaaaacaggc gctgccgggc
120gaagaacgcg atgaagcggt gagccgcctg aaagaatgcc tgattaacaa
cagcgatgaa 180ctgcgcctgg atcgcctgaa cctgagcagc ctgccggata
acctgccggc gcagattacc 240ctgctgaacg tgagctataa ccagctgacc
aacctgccgg aactgccggt gaccctgaaa 300aaactgtata gcgcgagcaa
caaactgagc gaactgccgg tgctgccgcc ggcgctggaa 360agcctgcagg
tgcagcataa cgaactggaa aacctgccgg cgctgccgga tagcctgctg
420accatgaaca ttagctataa cgaaattgtg agcctgccga gcctgccgca
ggcgctgaaa 480aacctgcgcg cgacccgcaa ctttctgacc gaactgccgg
cgtttagcga aggcaacaac 540ccggtggtgc gcgaatattt ttttgatcgc
aaccagatta gccatattcc ggaaagcatt 600ctgaacctgc gcaacgaatg
cagcattcat attagcgata acccgctgag cagccatgcg 660ctgccggcgc
tgcagcgcct gaccagcagc ccggattatc atggcccgcg catttatttt
720agcatgagcg atggccagca gaacaccctg catcgcccgc tggcggatgc
ggtgaccgcg 780tggtttccgg aaaacaaaca gagcgatgtg agccagattt
ggcatgcgtt tgaacatgaa 840gaacatgcga acacctttag cgcgtttctg
gatcgcctga gcgataccgt gagcgcgcgc 900aacaccagcg gctttcgcga
acaggtggcg gcgtggctgg aaaaactgag cgcgagcgcg 960gaactgcgcc
agcagagctt tgcggtggcg gcggatgcga ccgaaagctg cgaagatcgc
1020gtggcgctga cctggaacaa cctgcgcaaa accctgctgg tgcatcaggc
gagcgaaggc 1080ctgtttgata acgataccgg cgcgctgctg agcctgggcc
gcgaaatgtt tcgcctggaa 1140attctggaag atattgcgcg cgataaagtg
cgcaccctgc attttgtgga tgaaattgaa 1200gtgtatctgg cgtttcagac
catgctggcg gaaaaactgc agctgagcac cgcggtgaaa 1260gaaatgcgct
tttatggcgt gagcggcgtg accgcgaacg atctgcgcac cgcggaagcg
1320atggtgcgca gccgcgaaga aaacgaattt accgattggt ttagcctgtg
gggcccgtgg 1380catgcggtgc tgaaacgcac cgaagcggat cgctgggcgc
aggcggaaga acagaaatat 1440gaaatgctgg aaaacgaata tccgcagcgc
gtggcggatc gcctgaaagc gagcggcctg 1500agcggcgatg cggatgcgga
acgcgaagcg ggcgcgcagg tgatgcgcga aaccgaacag 1560cagatttatc
gccagctgac cgatgaagtg ctggcgctgc gcctgccgga aaacggcagc
1620cagctgcatc atagc 163574196PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 74Met Lys Ile Thr Ser Thr
Ile Ile Gln Thr Pro Phe Pro Phe Glu Asn1 5 10 15Asn Asn Ser His Ala
Gly Ile Val Thr Glu Pro Ile Leu Gly Lys Leu 20 25 30Ile Gly Gln Gly
Ser Thr Ala Glu Ile Phe Glu Asp Val Asn Asp Ser 35 40 45Ser Ala Leu
Tyr Lys Lys Tyr Asp Leu Ile Gly Asn Gln Tyr Asn Glu 50 55 60Ile Leu
Glu Met Ala Trp Gln Glu Ser Glu Leu Phe Asn Ala Phe Tyr65 70 75
80Gly Asp Glu Ala Ser Val Val Ile Gln Tyr Gly Gly Asp Val Tyr Leu
85 90 95Arg Met Leu Arg Val Pro Gly Thr Pro Leu Ser Asp Ile Asp Thr
Ala 100 105 110Asp Ile Pro Asp Asn Ile Glu Ser Leu Tyr Leu Gln Leu
Ile Cys Lys 115 120 125Leu Asn Glu Leu Ser Ile Ile His Tyr Asp Leu
Asn Thr Gly Asn Met 130 135 140Leu Tyr Asp Lys Glu Ser Glu Ser Leu
Phe Pro Ile Asp Phe Arg Asn145 150 155 160Ile Tyr Ala Glu Tyr Tyr
Ala Ala Thr Lys Lys Asp Lys Glu Ile Ile 165 170 175Asp Arg Arg Leu
Gln Met Arg Thr Asn Asp Phe Tyr Ser Leu Leu Asn 180 185 190Arg Lys
Tyr Leu 19575588DNAArtificial SequenceDescription of Artificial
Sequence Synthetic polynucleotide 75atgaaaatta ccagcaccat
tattcagacc ccgtttccgt ttgaaaacaa caacagccat 60gcgggcattg tgaccgaacc
gattctgggc aaactgattg gccagggcag caccgcggaa 120atttttgaag
atgtgaacga tagcagcgcg ctgtataaaa aatatgatct gattggcaac
180cagtataacg aaattctgga aatggcgtgg caggaaagcg aactgtttaa
cgcgttttat 240ggcgatgaag cgagcgtggt gattcagtat ggcggcgatg
tgtatctgcg catgctgcgc 300gtgccgggca ccccgctgag cgatattgat
accgcggata ttccggataa cattgaaagc 360ctgtatctgc agctgatttg
caaactgaac gaactgagca ttattcatta tgatctgaac 420accggcaaca
tgctgtatga taaagaaagc gaaagcctgt ttccgattga ttttcgcaac
480atttatgcgg aatattatgc ggcgaccaaa aaagataaag aaattattga
tcgccgcctg 540cagatgcgca ccaacgattt ttatagcctg ctgaaccgca
aatatctg
58876462PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 76Met Ser Pro Ile Leu Gly Tyr Trp Lys Ile Lys
Gly Leu Val Gln Pro1 5 10 15Thr Arg Leu Leu Leu Glu Tyr Leu Glu Glu
Lys Tyr Glu Glu His Leu 20 25 30Tyr Glu Arg Asp Glu Gly Asp Lys Trp
Arg Asn Lys Lys Phe Glu Leu 35 40 45Gly Leu Glu Phe Pro Asn Leu Pro
Tyr Tyr Ile Asp Gly Asp Val Lys 50 55 60Leu Thr Gln Ser Met Ala Ile
Ile Arg Tyr Ile Ala Asp Lys His Asn65 70 75 80Met Leu Gly Gly Cys
Pro Lys Glu Arg Ala Glu Ile Ser Met Leu Glu 85 90 95Gly Ala Val Leu
Asp Ile Arg Tyr Gly Val Ser Arg Ile Ala Tyr Ser 100 105 110Lys Asp
Phe Glu Thr Leu Lys Val Asp Phe Leu Ser Lys Leu Pro Glu 115 120
125Met Leu Lys Met Phe Glu Asp Arg Leu Cys His Lys Thr Tyr Leu Asn
130 135 140Gly Asp His Val Thr His Pro Asp Phe Met Leu Tyr Asp Ala
Leu Asp145 150 155 160Val Val Leu Tyr Met Asp Pro Met Cys Leu Asp
Ala Phe Pro Lys Leu 165 170 175Val Cys Phe Lys Lys Arg Ile Glu Ala
Ile Pro Gln Ile Asp Lys Tyr 180 185 190Leu Lys Ser Ser Lys Tyr Ile
Ala Trp Pro Leu Gln Gly Trp Gln Ala 195 200 205Thr Phe Gly Gly Gly
Asp His Pro Pro Lys Ser Asp Leu Val Pro Arg 210 215 220Gly Ser Met
Ser Ile Glu Ile Lys Met Ile Ser Pro Ile Lys Asn Ile225 230 235
240Lys Asn Val Phe Pro Ile Asn Thr Ala Asn Thr Glu Tyr Ile Val Arg
245 250 255Asn Ile Tyr Pro Arg Val Glu His Gly Tyr Phe Asn Glu Ser
Pro Asn 260 265 270Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg
Ser Met Ala Gln 275 280 285Leu Lys Ile Glu Glu Phe Ile Asn Glu Lys
Ser Arg Arg Leu Asn Tyr 290 295 300Met Lys Thr Met Tyr Ser Pro Cys
Pro Glu Asp Phe Gln Pro Ile Ser305 310 315 320Arg Asp Glu Ala Ser
Thr Pro Glu Gly Ser Trp Leu Thr Val Ile Ser 325 330 335Gly Lys Arg
Pro Met Gly Gln Phe Ser Val Asp Ser Leu Tyr His Pro 340 345 350Asp
Leu His Ala Leu Cys Glu Leu Pro Glu Ile Ser Cys Lys Ile Phe 355 360
365Pro Lys Glu Asn Ser Asp Phe Leu Tyr Ile Ile Val Val Phe Arg Asn
370 375 380Asp Ser Pro Gln Gly Glu Leu Arg Ala Asn Arg Phe Ile Glu
Leu Tyr385 390 395 400Asp Ile Lys Arg Glu Ile Met Gln Val Leu Arg
Asp Glu Ser Pro Glu 405 410 415Leu Lys Ser Ile Lys Ser Glu Ile Ile
Ile Ala Arg Glu Met Gly Glu 420 425 430Leu Phe Ser Tyr Ala Ser Glu
Glu Ile Asp Ser Tyr Ile Lys Gln Met 435 440 445Asn Asp Arg Leu Ser
Gln Ile Lys Ala Arg Met Pro Val Thr 450 455 460771386DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
77atgagcccga ttctgggcta ttggaaaatt aaaggcctgg tgcagccgac ccgcctgctg
60ctggaatatc tggaagaaaa atatgaagaa catctgtatg aacgcgatga aggcgataaa
120tggcgcaaca aaaaatttga actgggcctg gaatttccga acctgccgta
ttatattgat 180ggcgatgtga aactgaccca gagcatggcg attattcgct
atattgcgga taaacataac 240atgctgggcg gctgcccgaa agaacgcgcg
gaaattagca tgctggaagg cgcggtgctg 300gatattcgct atggcgtgag
ccgcattgcg tatagcaaag attttgaaac cctgaaagtg 360gattttctga
gcaaactgcc ggaaatgctg aaaatgtttg aagatcgcct gtgccataaa
420acctatctga acggcgatca tgtgacccat ccggatttta tgctgtatga
tgcgctggat 480gtggtgctgt atatggatcc gatgtgcctg gatgcgtttc
cgaaactggt gtgctttaaa 540aaacgcattg aagcgattcc gcagattgat
aaatatctga aaagcagcaa atatattgcg 600tggccgctgc agggctggca
ggcgaccttt ggcggcggcg atcatccgcc gaaaagcgat 660ctggtgccgc
gcggcagcat gagcattgaa attaaaatga ttagcccgat taaaaacatt
720aaaaacgtgt ttccgattaa caccgcgaac accgaatata ttgtgcgcaa
catttatccg 780cgcgtggaac atggctattt taacgaaagc ccgaacattt
atgataaaaa atatattagc 840ggcattaccc gcagcatggc gcagctgaaa
attgaagaat ttattaacga aaaaagccgc 900cgcctgaact atatgaaaac
catgtatagc ccgtgcccgg aagattttca gccgattagc 960cgcgatgaag
cgagcacccc ggaaggcagc tggctgaccg tgattagcgg caaacgcccg
1020atgggccagt ttagcgtgga tagcctgtat catccggatc tgcatgcgct
gtgcgaactg 1080ccggaaatta gctgcaaaat ttttccgaaa gaaaacagcg
attttctgta tattattgtg 1140gtgtttcgca acgatagccc gcagggcgaa
ctgcgcgcga accgctttat tgaactgtat 1200gatattaaac gcgaaattat
gcaggtgctg cgcgatgaaa gcccggaact gaaaagcatt 1260aaaagcgaaa
ttattattgc gcgcgaaatg ggcgaactgt ttagctatgc gagcgaagaa
1320attgatagct atattaaaca gatgaacgat cgcctgagcc agattaaagc
gcgcatgccg 1380gtgacc 138678212PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 78Met Ile Asn Gly Val Ser
Leu Gln Gly Thr Ala Gly Tyr Glu Ala His1 5 10 15Thr Glu Glu Gly Asn
Val Asn Val Lys Lys Leu Leu Glu Ser Leu Asn 20 25 30Ser Lys Ser Leu
Gly Asp Met Asp Lys Asp Ser Glu Leu Ala Ala Thr 35 40 45Leu Gln Lys
Met Ile Asn Pro Ser Gly Gly Asp Gly Asn Cys Ser Gly 50 55 60Cys Ala
Leu His Ala Cys Met Ala Met Leu Gly Tyr Gly Val Arg Glu65 70 75
80Ala Pro Val Pro Asn Glu Ile Ser Glu Tyr Met Thr Gly Phe Phe His
85 90 95Arg His Leu Glu Gln Ile Asp Ser Glu Gly Ile Val Ser His Pro
Asn 100 105 110Glu Thr Tyr Ser Lys Phe Arg Glu Arg Ile Ala Glu Asn
Ile Leu Gln 115 120 125Asn Thr Ser Lys Gly Ser Val Val Met Ile Ser
Ile Glu Gln Ala Thr 130 135 140His Trp Ile Ala Gly Phe Asn Asp Gly
Glu Lys Ile Met Phe Leu Asp145 150 155 160Val Gln Thr Gly Lys Gly
Phe Asn Leu Tyr Asp Pro Val Glu Lys Ser 165 170 175Pro Asp Ala Phe
Val Asp Glu Asn Ser Ser Val Gln Val Ile His Val 180 185 190Ser Asp
Gln Glu Phe Asp His Tyr Ala Asn Ser Ser Ser Trp Lys Ser 195 200
205Lys Arg Leu Cys 21079636DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 79atgattaacg
gcgtgagcct gcagggcacc gcgggctatg aagcgcatac cgaagaaggc 60aacgtgaacg
tgaaaaaact gctggaaagc ctgaacagca aaagcctggg cgatatggat
120aaagatagcg aactggcggc gaccctgcag aaaatgatta acccgagcgg
cggcgatggc 180aactgcagcg gctgcgcgct gcatgcgtgc atggcgatgc
tgggctatgg cgtgcgcgaa 240gcgccggtgc cgaacgaaat tagcgaatat
atgaccggct tttttcatcg ccatctggaa 300cagattgata gcgaaggcat
tgtgagccat ccgaacgaaa cctatagcaa atttcgcgaa 360cgcattgcgg
aaaacattct gcagaacacc agcaaaggca gcgtggtgat gattagcatt
420gaacaggcga cccattggat tgcgggcttt aacgatggcg aaaaaattat
gtttctggat 480gtgcagaccg gcaaaggctt taacctgtat gatccggtgg
aaaaaagccc ggatgcgttt 540gtggatgaaa acagcagcgt gcaggtgatt
catgtgagcg atcaggaatt tgatcattat 600gcgaacagca gcagctggaa
aagcaaacgc ctgtgc 63680298PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 80Pro Glu Leu Pro Pro His
Leu Glu Ser Leu Val Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu Pro
Glu Leu Pro Gln Ser Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn Asn
Leu Lys Ala Leu Ser Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu Ala
Ala Gly Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn Ser
Ser Phe Leu Lys Ile Ile Asp Ile Asn Pro Val Thr Asn Thr65 70 75
80Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys
85 90 95Asn Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro
Asn 100 105 110Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly
Val Ala Glu 115 120 125Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala
Arg Arg Phe Ser Tyr 130 135 140Met Lys Thr Met Tyr Ser Val Cys Pro
Glu Ala Phe Glu Pro Ile Ser145 150 155 160Arg Asn Glu Ala Ser Thr
Pro Glu Gly Ser Trp Leu Thr Val Ile Ser 165 170 175Gly Lys Arg Pro
Met Gly Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro 180 185 190Asp Leu
His Ala Leu Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe 195 200
205Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn
210 215 220Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu
Leu Tyr225 230 235 240Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn
Tyr Glu Leu Pro Glu 245 250 255Leu Lys Ala Val Lys Ser Glu Met Ile
Ile Ala Arg Glu Met Gly Glu 260 265 270Ile Phe Ser Tyr Met Pro Gly
Glu Ile Asp Ser Tyr Met Lys Tyr Ile 275 280 285Asn Asn Lys Leu Ser
Lys Ile Glu Gly Ser 290 29581315PRTArtificial SequenceDescription
of Artificial Sequence Synthetic polypeptide 81Pro Glu Leu Pro Pro
His Leu Glu Ser Leu Val Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu
Pro Glu Leu Pro Gln Ser Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn
Asn Leu Lys Ala Leu Ser Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu
Ala Ala Gly Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn
Ser Ser Phe Leu Lys Ile Ile Asp Pro Ala Pro Ala Pro Ala Pro65 70 75
80Ala Pro Ala Pro Ala Pro Ala Pro Pro Ala Ile Asn Pro Val Thr Asn
85 90 95Thr Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val
Val 100 105 110Lys Asn Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn
Glu Ser Pro 115 120 125Asn Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile
Thr Arg Gly Val Ala 130 135 140Glu Leu Lys Gln Glu Glu Phe Val Asn
Glu Lys Ala Arg Arg Phe Ser145 150 155 160Tyr Met Lys Thr Met Tyr
Ser Val Cys Pro Glu Ala Phe Glu Pro Ile 165 170 175Ser Arg Asn Glu
Ala Ser Thr Pro Glu Gly Ser Trp Leu Thr Val Ile 180 185 190Ser Gly
Lys Arg Pro Met Gly Gln Phe Ser Val Asp Ser Leu Tyr Asn 195 200
205Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile
210 215 220Phe Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile Val Val Val
Tyr Arg225 230 235 240Asn Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn
Arg Phe Ile Glu Leu 245 250 255Tyr Asn Ile Lys Arg Asp Ile Met Gln
Glu Leu Asn Tyr Glu Leu Pro 260 265 270Glu Leu Lys Ala Val Lys Ser
Glu Met Ile Ile Ala Arg Glu Met Gly 275 280 285Glu Ile Phe Ser Tyr
Met Pro Gly Glu Ile Asp Ser Tyr Met Lys Tyr 290 295 300Ile Asn Asn
Lys Leu Ser Lys Ile Glu Gly Ser305 310 31582314PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
82Pro Glu Leu Pro Pro His Leu Glu Ser Leu Val Ala Ser Cys Asn Ser1
5 10 15Leu Thr Glu Leu Pro Glu Leu Pro Gln Ser Leu Lys Ser Leu Gln
Val 20 25 30Asp Asn Asn Asn Leu Lys Ala Leu Ser Asp Leu Pro Pro Ser
Leu Glu 35 40 45Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu Glu Leu Pro
Glu Leu Gln 50 55 60Asn Ser Ser Phe Leu Lys Ile Ile Asp Gly Ser Gly
Ser Gly Ser Gly65 70 75 80Ser Gly Ser Gly Ser Gly Ser Gly Ser Ile
Asn Pro Val Thr Asn Thr 85 90 95Gln Gly Val Ser Pro Ile Asn Thr Lys
Tyr Ala Glu His Val Val Lys 100 105 110Asn Ile Tyr Pro Lys Ile Lys
His Asp Tyr Phe Asn Glu Ser Pro Asn 115 120 125Ile Tyr Asp Lys Lys
Tyr Ile Ser Gly Ile Thr Arg Gly Val Ala Glu 130 135 140Leu Lys Gln
Glu Glu Phe Val Asn Glu Lys Ala Arg Arg Phe Ser Tyr145 150 155
160Met Lys Thr Met Tyr Ser Val Cys Pro Glu Ala Phe Glu Pro Ile Ser
165 170 175Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser Trp Leu Thr Val
Ile Ser 180 185 190Gly Lys Arg Pro Met Gly Gln Phe Ser Val Asp Ser
Leu Tyr Asn Pro 195 200 205Asp Leu His Ala Leu Cys Glu Leu Pro Asp
Ile Cys Cys Lys Ile Phe 210 215 220Pro Lys Glu Asn Asn Asp Phe Leu
Tyr Ile Val Val Val Tyr Arg Asn225 230 235 240Asp Ser Pro Leu Gly
Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu Tyr 245 250 255Asn Ile Lys
Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu Leu Pro Glu 260 265 270Leu
Lys Ala Val Lys Ser Glu Met Ile Ile Ala Arg Glu Met Gly Glu 275 280
285Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp Ser Tyr Met Lys Tyr Ile
290 295 300Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser305
31083298PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 83Pro Glu Leu Pro Pro His Leu Glu Ser Leu Val
Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu Pro Glu Leu Pro Gln Ser
Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn Asn Leu Lys Ala Leu Ser
Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu Ala Ala Gly Asn Asn Gln
Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn Ser Ser Phe Leu Lys Ile
Ile Asp Ile Asn Pro Val Thr Asn Thr65 70 75 80Gln Gly Val Ser Pro
Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys 85 90 95Asn Ile Tyr Pro
Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro Asn 100 105 110Ile Tyr
Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly Val Ala Glu 115 120
125Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala Arg Arg Phe Ser Tyr
130 135 140Met Lys Thr Met Tyr Ser Val Cys Pro Glu Ala Phe Glu Pro
Ile Ser145 150 155 160Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser Trp
Leu Thr Val Ile Ser 165 170 175Gly Lys Ala Pro Met Gly Gln Phe Ser
Val Asp Ser Leu Tyr Asn Pro 180 185 190Asp Leu His Ala Leu Cys Glu
Leu Pro Asp Ile Cys Cys Lys Ile Phe 195 200 205Pro Lys Glu Asn Asn
Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn 210 215 220Asp Ser Pro
Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu Tyr225 230 235
240Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu Leu Pro Glu
245 250 255Leu Lys Ala Val Lys Ser Glu Met Ile Ile Ala Arg Glu Met
Gly Glu 260 265 270Ile Phe Ser Tyr Met Pro Gly Glu Ile Asp Ser Tyr
Met Lys Tyr Ile 275 280 285Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser
290 29584315PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 84Pro Glu Leu Pro Pro His Leu Glu
Ser Leu Val Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu Pro Glu Leu
Pro Gln Ser Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn Asn Leu Lys
Ala Leu Ser Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu Ala Ala Gly
Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn Ser Ser Phe
Leu Lys Ile Ile Asp Pro Ala Pro Ala Pro Ala Pro65 70 75 80Ala Pro
Ala Pro Ala Pro Ala Pro Pro Ala Ile Asn Pro Val Thr Asn 85 90 95Thr
Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val 100 105
110Lys Asn Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser
Pro
115 120 125Asn Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly
Val Ala 130 135 140Glu Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala
Arg Arg Phe Ser145 150 155 160Tyr Met Lys Thr Met Tyr Ser Val Cys
Pro Glu Ala Phe Glu Pro Ile 165 170 175Ser Arg Asn Glu Ala Ser Thr
Pro Glu Gly Ser Trp Leu Thr Val Ile 180 185 190Ser Gly Lys Ala Pro
Met Gly Gln Phe Ser Val Asp Ser Leu Tyr Asn 195 200 205Pro Asp Leu
His Ala Leu Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile 210 215 220Phe
Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg225 230
235 240Asn Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu
Leu 245 250 255Tyr Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr
Glu Leu Pro 260 265 270Glu Leu Lys Ala Val Lys Ser Glu Met Ile Ile
Ala Arg Glu Met Gly 275 280 285Glu Ile Phe Ser Tyr Met Pro Gly Glu
Ile Asp Ser Tyr Met Lys Tyr 290 295 300Ile Asn Asn Lys Leu Ser Lys
Ile Glu Gly Ser305 310 31585314PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 85Pro Glu Leu Pro Pro His
Leu Glu Ser Leu Val Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu Pro
Glu Leu Pro Gln Ser Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn Asn
Leu Lys Ala Leu Ser Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu Ala
Ala Gly Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn Ser
Ser Phe Leu Lys Ile Ile Asp Gly Ser Gly Ser Gly Ser Gly65 70 75
80Ser Gly Ser Gly Ser Gly Ser Gly Ser Ile Asn Pro Val Thr Asn Thr
85 90 95Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val
Lys 100 105 110Asn Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu
Ser Pro Asn 115 120 125Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr
Arg Gly Val Ala Glu 130 135 140Leu Lys Gln Glu Glu Phe Val Asn Glu
Lys Ala Arg Arg Phe Ser Tyr145 150 155 160Met Lys Thr Met Tyr Ser
Val Cys Pro Glu Ala Phe Glu Pro Ile Ser 165 170 175Arg Asn Glu Ala
Ser Thr Pro Glu Gly Ser Trp Leu Thr Val Ile Ser 180 185 190Gly Lys
Ala Pro Met Gly Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro 195 200
205Asp Leu His Ala Leu Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe
210 215 220Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr
Arg Asn225 230 235 240Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg
Phe Ile Glu Leu Tyr 245 250 255Asn Ile Lys Arg Asp Ile Met Gln Glu
Leu Asn Tyr Glu Leu Pro Glu 260 265 270Leu Lys Ala Val Lys Ser Glu
Met Ile Ile Ala Arg Glu Met Gly Glu 275 280 285Ile Phe Ser Tyr Met
Pro Gly Glu Ile Asp Ser Tyr Met Lys Tyr Ile 290 295 300Asn Asn Lys
Leu Ser Lys Ile Glu Gly Ser305 31086292PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
86Pro Glu Leu Pro Pro His Leu Glu Ser Leu Val Ala Ser Cys Asn Ser1
5 10 15Leu Thr Glu Leu Pro Glu Leu Pro Gln Ser Leu Lys Ser Leu Gln
Val 20 25 30Asp Asn Asn Asn Leu Lys Ala Leu Ser Asp Leu Pro Pro Ser
Leu Glu 35 40 45Phe Leu Ala Ala Gly Asn Asn Gln Leu Glu Glu Leu Pro
Glu Leu Gln 50 55 60Asn Ser Ser Phe Leu Lys Ile Ile Asp Ile Asn Pro
Val Thr Asn Thr65 70 75 80Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr
Ala Glu His Val Val Lys 85 90 95Asn Ile Tyr Pro Lys Ile Lys His Asp
Tyr Phe Asn Glu Ser Pro Asn 100 105 110Ile Tyr Asp Lys Lys Tyr Ile
Ser Gly Ile Thr Arg Gly Val Ala Glu 115 120 125Leu Lys Gln Glu Glu
Phe Val Asn Glu Lys Ala Arg Arg Phe Ser Tyr 130 135 140Met Lys Thr
Met Tyr Ser Val Cys Pro Glu Ala Phe Glu Pro Ile Ser145 150 155
160Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser Trp Leu Thr Val Ile Ser
165 170 175Gly Lys Arg Pro Met Gly Gln Phe Ser Val Asp Ser Leu Tyr
Asn Pro 180 185 190Asp Leu His Ala Leu Cys Glu Leu Pro Asp Ile Cys
Cys Lys Ile Phe 195 200 205Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile
Val Val Val Tyr Arg Asn 210 215 220Asp Ser Pro Leu Gly Glu Gln Arg
Ala Asn Arg Phe Ile Glu Leu Tyr225 230 235 240Asn Ile Lys Arg Asp
Ile Met Gln Glu Leu Asn Tyr Glu Leu Pro Glu 245 250 255Leu Lys Ala
Val Lys Ser Glu Met Ile Ile Ala Arg Glu Met Gly Glu 260 265 270Ile
Phe Ser Tyr Met Pro Gly Glu Ile Asp Ser Tyr Met Lys Tyr Ile 275 280
285Asn Asn Lys Leu 29087309PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 87Pro Glu Leu Pro Pro His
Leu Glu Ser Leu Val Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu Pro
Glu Leu Pro Gln Ser Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn Asn
Leu Lys Ala Leu Ser Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu Ala
Ala Gly Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn Ser
Ser Phe Leu Lys Ile Ile Asp Pro Ala Pro Ala Pro Ala Pro65 70 75
80Ala Pro Ala Pro Ala Pro Ala Pro Pro Ala Ile Asn Pro Val Thr Asn
85 90 95Thr Gln Gly Val Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val
Val 100 105 110Lys Asn Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn
Glu Ser Pro 115 120 125Asn Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile
Thr Arg Gly Val Ala 130 135 140Glu Leu Lys Gln Glu Glu Phe Val Asn
Glu Lys Ala Arg Arg Phe Ser145 150 155 160Tyr Met Lys Thr Met Tyr
Ser Val Cys Pro Glu Ala Phe Glu Pro Ile 165 170 175Ser Arg Asn Glu
Ala Ser Thr Pro Glu Gly Ser Trp Leu Thr Val Ile 180 185 190Ser Gly
Lys Arg Pro Met Gly Gln Phe Ser Val Asp Ser Leu Tyr Asn 195 200
205Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile
210 215 220Phe Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile Val Val Val
Tyr Arg225 230 235 240Asn Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn
Arg Phe Ile Glu Leu 245 250 255Tyr Asn Ile Lys Arg Asp Ile Met Gln
Glu Leu Asn Tyr Glu Leu Pro 260 265 270Glu Leu Lys Ala Val Lys Ser
Glu Met Ile Ile Ala Arg Glu Met Gly 275 280 285Glu Ile Phe Ser Tyr
Met Pro Gly Glu Ile Asp Ser Tyr Met Lys Tyr 290 295 300Ile Asn Asn
Lys Leu30588308PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 88Pro Glu Leu Pro Pro His Leu Glu
Ser Leu Val Ala Ser Cys Asn Ser1 5 10 15Leu Thr Glu Leu Pro Glu Leu
Pro Gln Ser Leu Lys Ser Leu Gln Val 20 25 30Asp Asn Asn Asn Leu Lys
Ala Leu Ser Asp Leu Pro Pro Ser Leu Glu 35 40 45Phe Leu Ala Ala Gly
Asn Asn Gln Leu Glu Glu Leu Pro Glu Leu Gln 50 55 60Asn Ser Ser Phe
Leu Lys Ile Ile Asp Gly Ser Gly Ser Gly Ser Gly65 70 75 80Ser Gly
Ser Gly Ser Gly Ser Gly Ser Ile Asn Pro Val Thr Asn Thr 85 90 95Gln
Gly Val Ser Pro Ile Asn Thr Lys Tyr Ala Glu His Val Val Lys 100 105
110Asn Ile Tyr Pro Lys Ile Lys His Asp Tyr Phe Asn Glu Ser Pro Asn
115 120 125Ile Tyr Asp Lys Lys Tyr Ile Ser Gly Ile Thr Arg Gly Val
Ala Glu 130 135 140Leu Lys Gln Glu Glu Phe Val Asn Glu Lys Ala Arg
Arg Phe Ser Tyr145 150 155 160Met Lys Thr Met Tyr Ser Val Cys Pro
Glu Ala Phe Glu Pro Ile Ser 165 170 175Arg Asn Glu Ala Ser Thr Pro
Glu Gly Ser Trp Leu Thr Val Ile Ser 180 185 190Gly Lys Arg Pro Met
Gly Gln Phe Ser Val Asp Ser Leu Tyr Asn Pro 195 200 205Asp Leu His
Ala Leu Cys Glu Leu Pro Asp Ile Cys Cys Lys Ile Phe 210 215 220Pro
Lys Glu Asn Asn Asp Phe Leu Tyr Ile Val Val Val Tyr Arg Asn225 230
235 240Asp Ser Pro Leu Gly Glu Gln Arg Ala Asn Arg Phe Ile Glu Leu
Tyr 245 250 255Asn Ile Lys Arg Asp Ile Met Gln Glu Leu Asn Tyr Glu
Leu Pro Glu 260 265 270Leu Lys Ala Val Lys Ser Glu Met Ile Ile Ala
Arg Glu Met Gly Glu 275 280 285Ile Phe Ser Tyr Met Pro Gly Glu Ile
Asp Ser Tyr Met Lys Tyr Ile 290 295 300Asn Asn Lys
Leu30589224PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 89Asn Pro Val Thr Asn Thr Gln Gly Val Ser Pro
Ile Asn Thr Lys Tyr1 5 10 15Ala Glu His Val Val Lys Asn Ile Tyr Pro
Lys Ile Lys His Asp Tyr 20 25 30Phe Asn Glu Ser Pro Asn Ile Tyr Asp
Lys Lys Tyr Ile Ser Gly Ile 35 40 45Thr Arg Gly Val Ala Glu Leu Lys
Gln Glu Glu Phe Val Asn Glu Lys 50 55 60Ala Arg Arg Phe Ser Tyr Met
Lys Thr Met Tyr Ser Val Cys Pro Glu65 70 75 80Ala Phe Glu Pro Ile
Ser Arg Asn Glu Ala Ser Thr Pro Glu Gly Ser 85 90 95Trp Leu Thr Val
Ile Ser Gly Lys Arg Pro Met Gly Gln Phe Ser Val 100 105 110Asp Ser
Leu Tyr Asn Pro Asp Leu His Ala Leu Cys Glu Leu Pro Asp 115 120
125Ile Cys Cys Lys Ile Phe Pro Lys Glu Asn Asn Asp Phe Leu Tyr Ile
130 135 140Val Val Val Tyr Arg Asn Asp Ser Pro Leu Gly Glu Gln Arg
Ala Asn145 150 155 160Arg Phe Ile Glu Leu Tyr Asn Ile Lys Arg Asp
Ile Met Gln Glu Leu 165 170 175Asn Tyr Glu Leu Pro Glu Leu Lys Ala
Val Lys Ser Glu Met Ile Ile 180 185 190Ala Arg Glu Met Gly Glu Ile
Phe Ser Tyr Met Pro Gly Glu Ile Asp 195 200 205Ser Tyr Met Lys Tyr
Ile Asn Asn Lys Leu Ser Lys Ile Glu Gly Ser 210 215
220906PRTArtificial SequenceDescription of Artificial Sequence
Synthetic 6xHis tag 90His His His His His His1 5914PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 91Pro
Ala Pro Ala1924PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 92Gly Ser Gly Ser193672DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
93atgattaatc ctgttactaa tactcagggc gtgtccccta taaatactaa atatgctgaa
60catgtggtga aaaatattta cccgaaaatt aaacatgatt actttaatga atcacccaat
120atatatgata agaagtatat atccggtata accagaggag tagctgaact
aaaacaggaa 180gaatttgtta acgagaaagc cagacggttt tcttatatga
agactatgta ttctgtatgt 240ccagaagcgt ttgaacctat ttccagaaat
gaagccagta caccggaagg aagctggcta 300acagttatat ccggaaaacg
cccaatgggg cagttttctg tagatagttt atacaatcct 360gatttacatg
cattatgtga gcttccggac atttgttgta agatcttccc taaagaaaat
420aatgattttt tatacatagt tgttgtgtac agaaatgaca gccctctagg
agaacaacgg 480gcaaatagat ttatagaatt atataatata aaaagagata
tcatgcagga attaaattat 540gagttaccag agttaaaggc agtaaaatct
gaaatgatta tcgcacgtga aatgggagaa 600atctttagct acatgcctgg
ggaaatagac agttatatga aatacataaa taataaactt 660tctaaaattg ag
672
References
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014
D00015

D00016
D00017
D00018

D00019

D00020

D00021

D00022
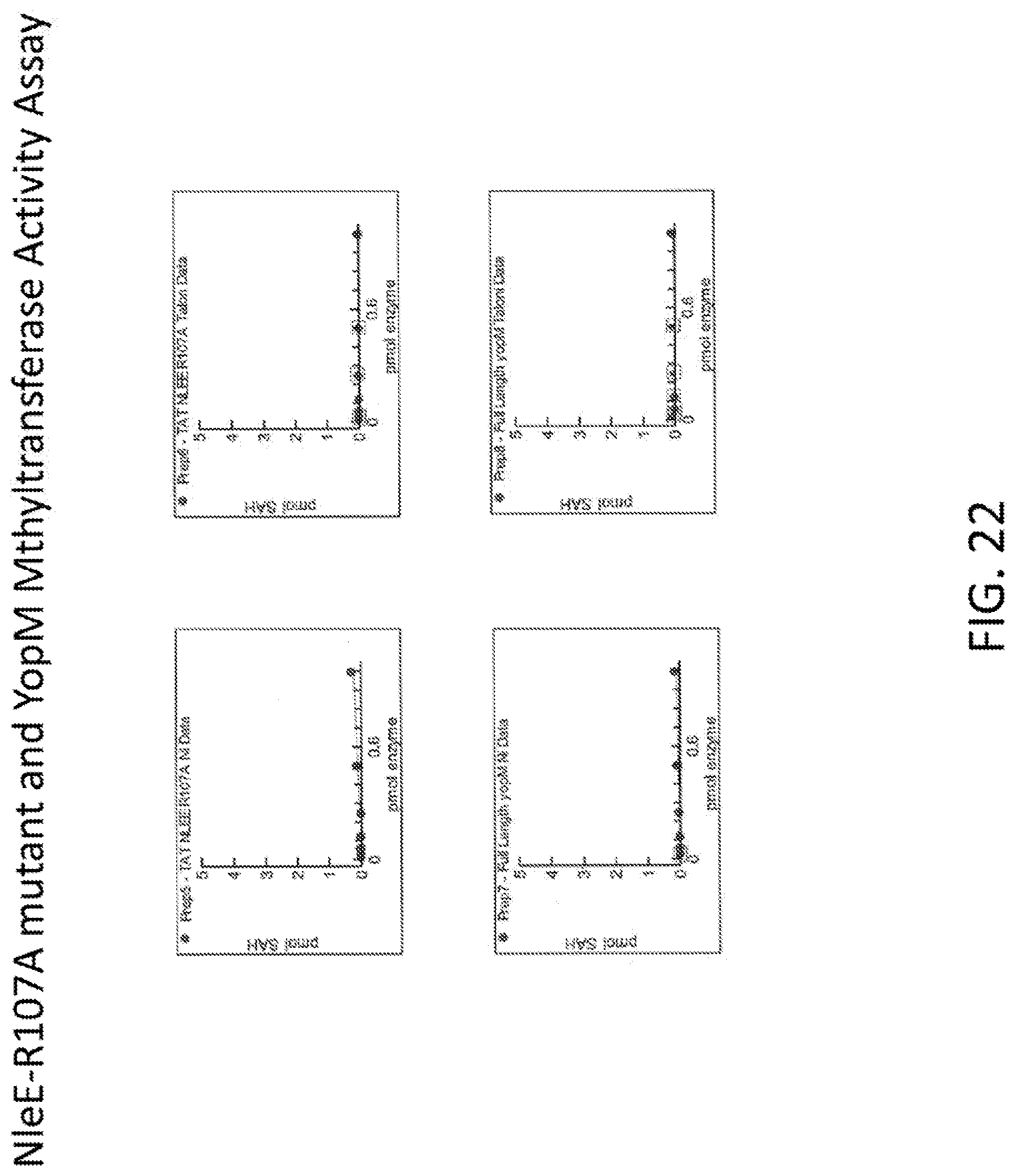
D00023
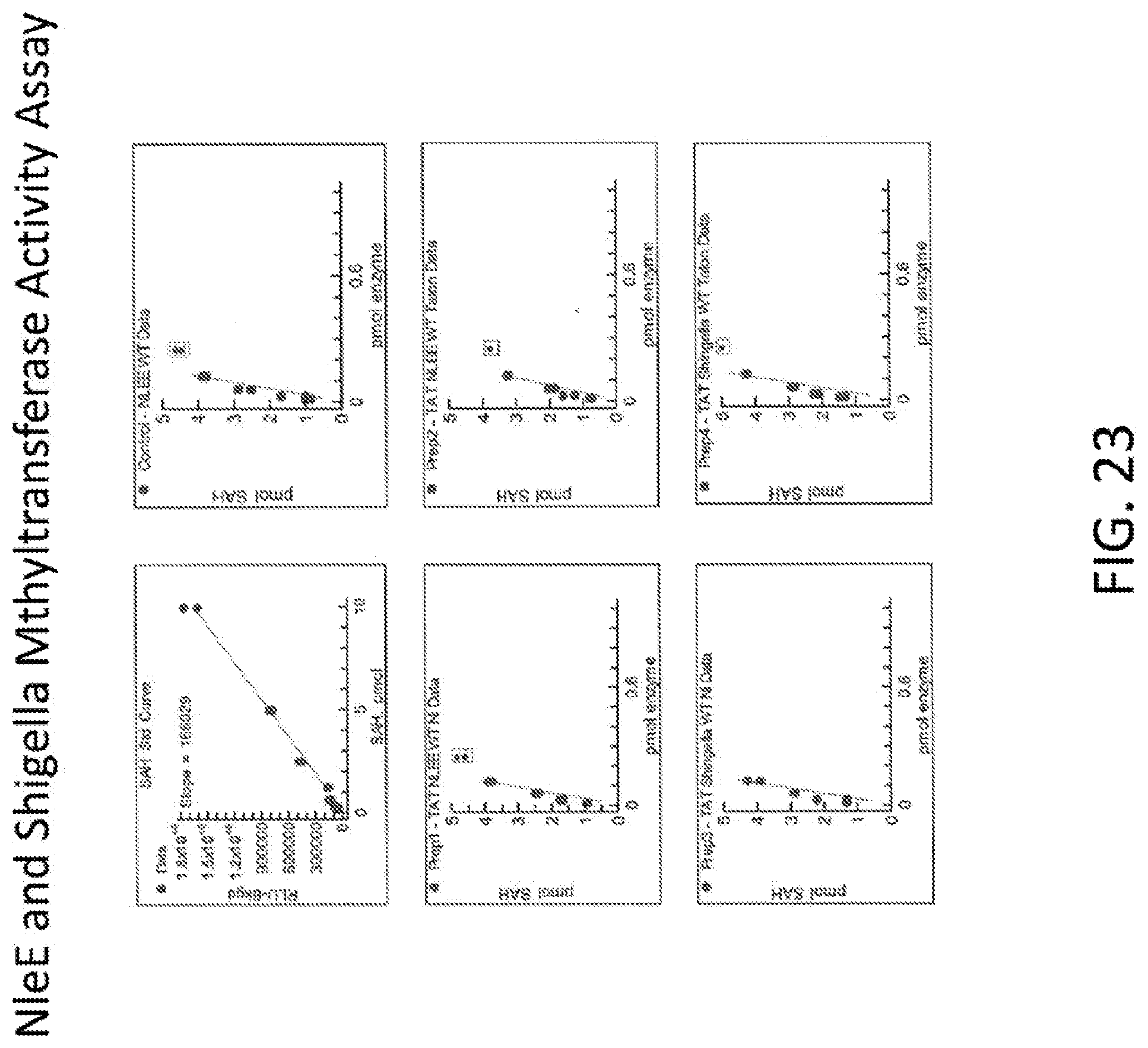

D00024

D00025

D00026
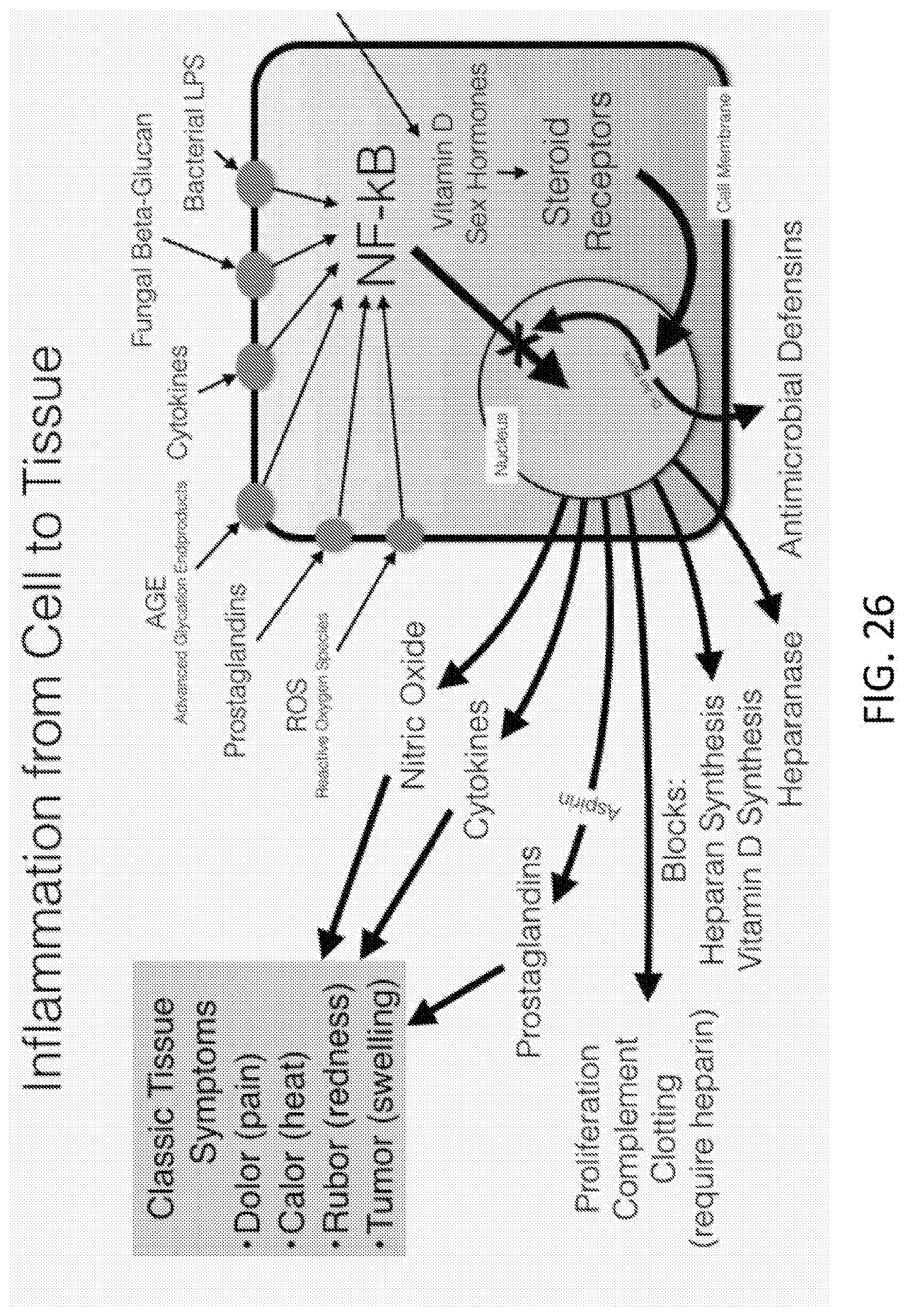
D00027
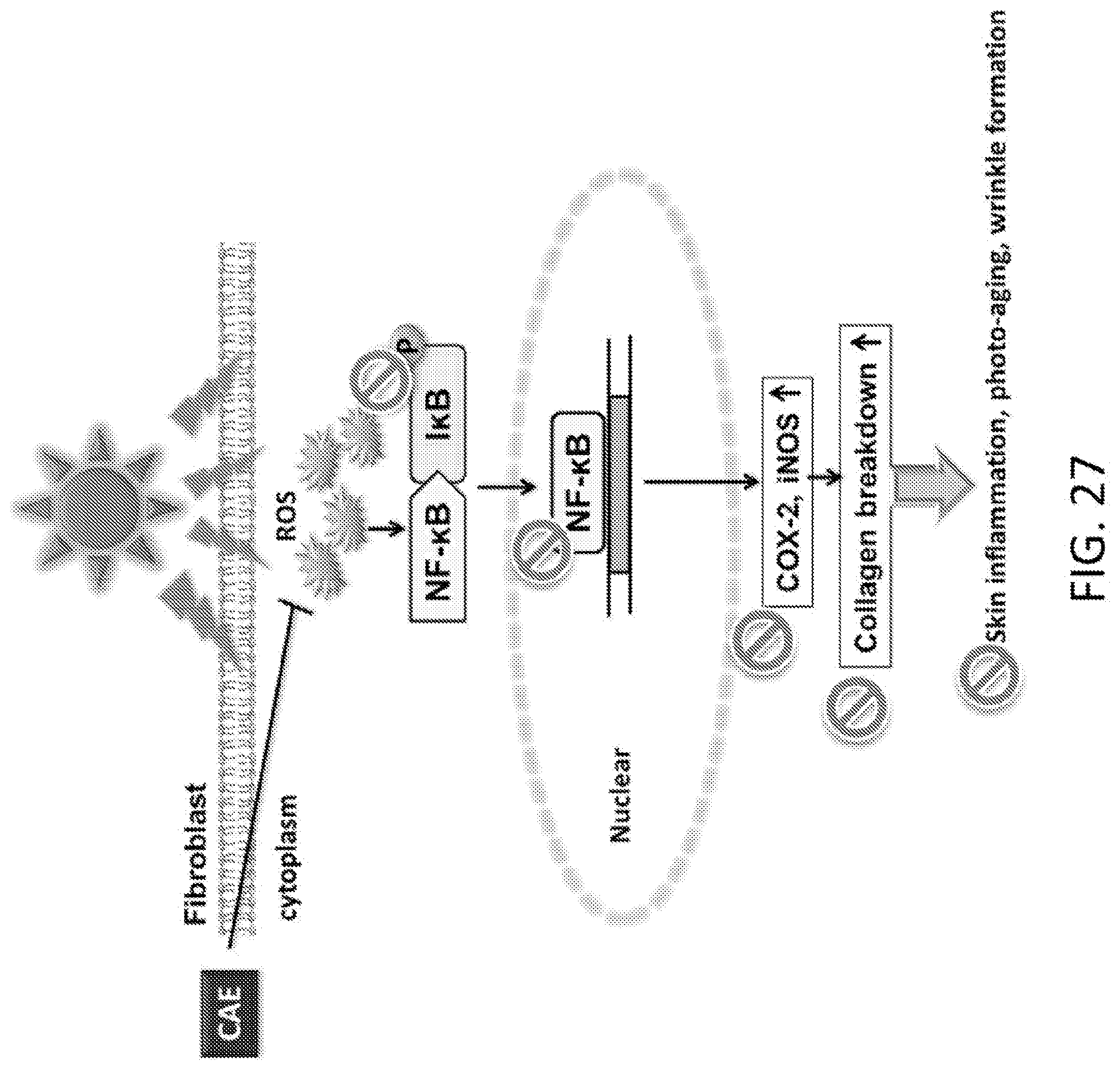
D00028

D00029

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.