Single-Chain Trail-Receptor Agonist Proteins
Buchanan; Fritz G. ; et al.
U.S. patent application number 16/993569 was filed with the patent office on 2021-02-11 for single-chain trail-receptor agonist proteins. The applicant listed for this patent is AbbVie Inc., Apogenix AG. Invention is credited to Fritz G. Buchanan, Christian Gieffers, Oliver Hill, Susan E. Lappe, Darren C. Phillips, Meinholf Thiemann.
| Application Number | 20210040178 16/993569 |
| Document ID | / |
| Family ID | 1000005170044 |
| Filed Date | 2021-02-11 |











View All Diagrams
| United States Patent Application | 20210040178 |
| Kind Code | A1 |
| Buchanan; Fritz G. ; et al. | February 11, 2021 |
Single-Chain Trail-Receptor Agonist Proteins
Abstract
Provided herein are specific TRAIL receptor agonist proteins, nucleic acids encoding the same, and methods of treating a subject having a TRAIL-associated disease or disorder. The TRAIL receptor agonist proteins provided herein comprise three soluble TRAIL domains and an Fc fragment. The TRAIL receptor agonist proteins are substantially non-aggregating and suitable for therapeutic, diagnostic and/or research applications.
| Inventors: | Buchanan; Fritz G.; (Salem, WI) ; Gieffers; Christian; (Dossenheim, DE) ; Hill; Oliver; (Neckarsteinach, DE) ; Lappe; Susan E.; (Riverwoods, IL) ; Phillips; Darren C.; (Glenview, IL) ; Thiemann; Meinholf; (Schriesheim, DE) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005170044 | ||||||||||
| Appl. No.: | 16/993569 | ||||||||||
| Filed: | August 14, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15887509 | Feb 2, 2018 | |||
| 16993569 | ||||
| 14694358 | Apr 23, 2015 | 9908927 | ||
| 15887509 | ||||
| 61983152 | Apr 23, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2319/30 20130101; C07K 14/70575 20130101; C07K 2319/00 20130101; A61K 38/00 20130101 |
| International Class: | C07K 14/705 20060101 C07K014/705 |
Claims
1. A TRAIL receptor agonist protein comprising a polypeptide having the amino acid sequence set forth in SEQ ID NO: 19.
2. A TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19.
3. The TRAIL receptor agonist protein of claim 2, wherein the two polypeptides are covalently linked through three interchain disulfide bonds formed between cysteine residues 513, 519, and 522 of each polypeptide.
4. The TRAIL receptor agonist protein of claim 2, wherein one or more of the asparagine residues at positions 168 and 337 of the polypeptide(s) are N-glycosylated.
5. The TRAIL receptor agonist protein of claim 2, wherein the asparagine residues at positions 168 and 337 of the polypeptide(s) are both N-glycosylated.
6. The TRAIL receptor agonist protein of claim 5, wherein the polypeptide(s) are further post-translationally modified.
7. The TRAIL receptor agonist protein of claim 6, wherein the post-translational modification comprises modification of the N-terminal glutamine to pyroglutamate.
8. A pharmaceutical composition comprising the TRAIL receptor agonist protein of any one of claim 6 and one or more pharmaceutically acceptable carriers, diluents, excipients, and/or adjuvants.
9. A nucleic acid molecule encoding the TRAIL receptor agonist protein of claim 1.
10. An expression vector comprising the nucleic acid molecule of claim 9.
11. A cell comprising the nucleic acid molecule of claim 9.
12. The cell of claim 11, which is a eukaryotic cell.
13. The cell of claim 11, wherein the cell is a mammalian cell.
14. The cell of claim 11, wherein the cell is a Chinese Hamster Ovary (CHO) cell.
15. A method of treating a subject having a TRAIL-associated disease or disorder, the method comprising administering to the subject an effective amount of the TRAIL receptor agonist protein of any one of claim 6.
16. The method of claim 15, wherein the disease or disorder is selected from the group consisting of: tumors, infectious diseases, inflammatory diseases, metabolic diseases, autoimmune disorders, degenerative diseases, apoptosis-associated diseases, and transplant rejections.
17. The method of claim 16, wherein the tumors are solid tumors.
18. The method of claim 16, wherein the tumors are lymphatic tumors.
19. The method of claim 16, wherein the autoimmune disorders are rheumatoid diseases, arthritic diseases, or rheumatoid and arthritic diseases.
20. The method of claim 16, wherein the disease or disorder is rheumatoid arthritis.
21. The method of claim 16, wherein the degenerative disease is a neurodegenerative disease.
22. The method of claim 16, wherein the neurodegenerative disease is multiple sclerosis.
Description
RELATED APPLICATIONS
[0001] This application is a continuation of U.S. application Ser. No. 15/887,509, filed Feb. 2, 2018 which is a continuation of U.S. application Ser. No. 14/694,358, filed Apr. 23, 2015, now U.S. Pat. No. 9,908,927, which claims the benefit of priority to U.S. Provisional Patent Application No. 61/983,152, filed Apr. 23, 2014. All of which are incorporated by reference herein in its entirety.
FIELD OF THE INVENTION
[0002] The present invention provides specific TRAIL receptor agonist proteins comprising three soluble TRAIL domains and an Fc fragment, nucleic acid molecules encoding the TRAIL receptor agonist proteins, and uses thereof. The TRAIL receptor agonist proteins are substantially non-aggregating and suitable for therapeutic, diagnostic and/or research applications.
BACKGROUND OF THE INVENTION
[0003] It is known that trimerization of TNF superfamily (TNFSF) cytokines is required for efficient receptor binding and activation. Trimeric complexes of TNF superfamily cytokines, however, are difficult to prepare from recombinant monomeric units.
[0004] WO 01/49866 and WO 02/09055 disclose recombinant fusion proteins comprising a TNF cytokine and a multimerization component, particularly a protein from the C1q protein family or a collectin. A disadvantage of these fusion proteins is, however, that the trimerization domain usually has a large molecular weight and/or that the trimerization is rather inefficient.
[0005] Schneider et al. (J Exp Med 187 (1989), 1205-1213) describe that trimers of TNF cytokines are stabilized by N-terminally positioned stabilization motifs. In CD95L, the stabilization of the receptor binding domain trimer is presumably caused by N-terminal amino acid domains which are located near the cytoplasmic membrane.
[0006] Shiraishi et al. (Biochem Biophys Res Commun 322 (2004), 197-202) describe that the receptor binding domain of CD95L may be stabilized by N-terminally positioned artificial .alpha.-helical coiled-coil (leucine zipper) motifs. It was found, however, that the orientation of the polypeptide chains to each other, e.g. parallel or antiparallel orientation, can hardly be predicted. Further, the optimal number of heptad-repeats in the coiled-coil zipper motif are difficult to determine. In addition, coiled-coil structures have the tendency to form macromolecular aggregates after alteration of pH and/or ionic strength.
[0007] WO 01/25277 relates to single-chain oligomeric polypeptides which bind to an extracellular ligand binding domain of a cellular receptor, wherein the polypeptide comprises at least three receptor binding sites of which at least one is capable of binding to a ligand binding domain of the cellular receptor and at least one is incapable of effectively binding to a ligand binding domain of the cellular receptor, whereby the single-chain oligomeric polypeptides are capable of binding to the receptor, but incapable of activating the receptor. For example, the monomers are derived from cytokine ligands of the TNF family, particularly from TNF-.alpha..
[0008] WO 2005/103077 discloses single-chain fusion polypeptides comprising at least three monomers of a TNF family ligand member and at least two peptide linkers that link the monomers of the TNF ligand family members to one another. Recent experiments, however, have shown that these single-chain fusion polypeptides show undesired aggregation.
[0009] WO 2010/010051 discloses single-chain fusion polypeptides comprising three soluble TNF family cytokine domains and at least two peptide linkers. The described fusion polypeptides are substantially non-aggregating.
[0010] Moreover, previous work, including that of Papadopoulos et al. (Cancer Chemother Pharmacol, 2015, DOI 10.1007/s00280-015-2712-0), has demonstrated that TRAIL receptor superclustering can result in toxicity.
[0011] Accordingly, there is a need in the art for novel TRAIL receptor agonists that exhibit high biological activity, high stability, low toxicity, and allow for efficient recombinant manufacturing.
SUMMARY OF THE INVENTION
[0012] The present invention provides specific TRAIL receptor agonist proteins that exhibit low proteolytic degradation, long half-life, and low TRAIL receptor superclustering in vivo (along with concomitant toxicity).
[0013] The TRAIL receptor agonist proteins of the instant invention generally comprise: (i) a first soluble TRAIL cytokine domain; (ii) a first peptide linker; (iii) a second soluble TRAIL domain; (iv) a second peptide linker; (v) a third soluble TRAIL domain; and (vi) an antibody Fc fragment.
[0014] In one aspect, the present invention provides a single-chain fusion polypeptide comprising: (i) a first soluble TRAIL domain, (ii) a first peptide linker, (iii) a second soluble TRAIL domain, (iv) a second peptide linker, (v) a third soluble TRAIL domain, and (vi) an antibody Fc fragment. In one embodiment, the antibody Fc fragment (vi) is located N terminal to the first TRAIL domain (i) and/or C-terminal to the third TRAIL domain (v). In another embodiment the antibody Fc fragment is located C-terminally to the third TRAIL domain (v). In one embodiment, the polypeptide is substantially non aggregating. In another embodiment, the second and/or third soluble TRAIL domain is an N-terminally shortened domain which optionally comprises amino acid sequence mutations.
[0015] In one embodiment, at least one of the soluble TRAIL domains, particularly at least one of the soluble TRAIL domains (iii) and (v), is a soluble TRAIL domain with an N-terminal sequence which starts between amino acid GIn120 and Val122 of human TRAIL and wherein Arg121 may be replaced by a neutral amino acid, e.g., Ser or Gly. In another embodiment, at least one of the soluble TRAIL domains, particularly at least one of the soluble TRAIL domains (iii) and (v), is a soluble TRAIL domain with an N-terminal sequence selected from (a) Arg121-Val122-Ala123 and (b) (Gly/Ser)121-Val122-Ala123. In one embodiment, the soluble TRAIL domain ends with amino acid Gly281 of human TRAIL and/or optionally comprises a mutation at positions R130, G160, H168, R170, H177, Y189, R191, Q193, E195, N199, K201, Y213, T214, S215, H264, 1266, D267 or D269 or at two or more of said positions. In one embodiment, the soluble TRAIL domain (i) consists of amino acids GIn120-Gly281 of human TRAIL according to SEQ ID NO: 1 and the soluble TRAIL domains (iii) and (v) consist of amino acids Arg121-Gly281 of human TRAIL according to SEQ ID NO: 1.
[0016] In one embodiment, the first and second peptide linkers (ii) and (iv) independently have a length of 3-8 amino acids, particularly a length of 3, 4, 5, 6, 7, or 8 amino acids, and preferably are glycine/serine linkers, optionally comprising an asparagine residue which may be glycosylated. In one embodiment, the first and the second peptide linkers (ii) and (iv) consist of the amino acid sequence according to SEQ ID NO: 2. In another embodiment, the polypeptide additionally comprises an N-terminal signal peptide domain, e.g., of SEQ ID NO: 12, which may comprise a protease cleavage site, and/or which additionally comprises a C-terminal element which may comprise and/or connect to a recognition/purification domain, e.g., a Strep-tag according to SEQ ID NO: 13.
[0017] In one embodiment, the antibody Fc fragment (vi) is fused to the soluble TRAIL domain (i) and/or (v) via a hinge-linker, preferably of SEQ ID NO: 11. In another embodiment, the antibody Fc fragment (vi) consists of the amino acid sequence as shown in SEQ ID NO: 10 or 17. In one embodiment, the polypeptide comprises the amino acid sequence of SEQ ID NO: 14, 15 or 18.
[0018] In another aspect, the present invention provides a TRAIL receptor agonist protein comprising a polypeptide having the amino acid sequence set forth in SEQ ID NO: 19, 20 or 21.
[0019] In another aspect, the present invention provides a TRAIL receptor agonist protein comprising a polypeptide having the amino acid sequence set forth in SEQ ID NO: 26, 27, 28, 29, or 30.
[0020] In another aspect, the present invention provides a TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO:
[0021] 19. In one embodiment, the two polypeptides are covalently linked through three interchain disulfide bonds formed between cysteine residues 513, 519, and 522 of each polypeptide.
[0022] In one embodiment, one or more of the asparagine residues at positions 168 and 337 of the polypeptide(s) are N-glycosylated. In another embodiment, the asparagine residues at positions 168 and 337 of the polypeptide(s) are both N-glycosylated.
[0023] In another embodiment, the polypeptide(s) are further post-translationally modified. In another embodiment, the post-translational modification comprises the N-terminal glutamine modified to pyroglutamate.
[0024] In another aspect, the present invention provides a pharmaceutical composition comprising a TRAIL receptor agonist protein disclosed herein and one or more pharmaceutically acceptable carriers, diluents, excipients, and/or adjuvants.
[0025] In another aspect, the present invention provides a nucleic acid molecule encoding the TRAIL receptor agonist protein. In another embodiment, the present invention provides an expression vector comprising the nucleic acid molecule. In another embodiment, the present invention provides a cell comprising the nucleic acid molecule. In a further embodiment, the cell is a eukaryotic cell. In another embodiment, the cell is a mammalian cell. In another embodiment, the cell is a Chinese Hamster Ovary (CHO) cell. In other embodiments, the cell is selected from the group consisting of CHO-DBX11, CHO-DG44, CHO-S, and CHO-K1 cells. In other embodiments, the cell is selected from the group consisting of Vero, BHK, HeLa, COS, MDCK, HEK-293, NIH-3T3, W138, BT483, Hs578T, HTB2, BT20, T47D, NSO, CRL7030, HsS78Bst, PER.C6, SP2/0-Agl4, and hybridoma cells.
[0026] In another aspect, the present invention provides a method of treating a subject having a TRAIL-associated disease or disorder, the method comprising administering to the subject an effective amount of the TRAIL receptor agonist protein. In one embodiment, the TRAIL receptor agonist protein is administered alone. In another embodiment, the TRAIL receptor agonist protein is administered before, concurrently, or after the administration of a second agent. In another embodiment, the disease or disorder is selected from the group consisting of: tumors, infectious diseases, inflammatory diseases, metabolic diseases, autoimmune disorders, degenerative diseases, apoptosis-associated diseases, and transplant rejections. In one embodiment, the tumors are solid tumors. In one embodiment, the tumors arise from the group of cancers consisting of sarcoma, esophageal cancer, and gastric cancer. In another embodiment, the tumors arise from Ewing's sarcoma or fibrosarcoma, In another embodiment, the tumors arise from the group of cancers consisting of Non-Small Cell Lung Carcinoma (NSCLC), pancreatic cancer, colorectal cancer, breast cancer, ovarian cancer, head and neck cancers, and Small Cell Lung Cancer (SCLC). In another embodiment, the tumors are lymphatic tumors. In one embodiment, the tumors are hematologic tumors. In another embodiment, the tumors arise from non-Hodgkin's lymphoma, leukemia, acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), B cell lymphoma, Burkitt's lymphoma, chronic myelocytic leukemia (CML), chronic lymphocytic leukemia (CLL), or hairy cell leukemia. In another embodiment, the autoimmune disorders are rheumatoid diseases, arthritic diseases, or rheumatoid and arthritic diseases. In a further embodiment, the disease or disorder is rheumatoid arthritis. In another embodiment, the degenerative disease is a neurodegenerative disease. In a further embodiment, the neurodegenerative disease is multiple sclerosis.
[0027] In one embodiment, the second agent is a chemotherapeutic, radiotherapeutic, or biological agent. In one embodiment, the second agent is selected from the group consisting of Duvelisib, Ibrutinib, Navitoclax, and Venetoclax, In another embodiment, the second agent is an apoptotic agent. In one embodiment, the apoptotic second agent is selected from the group consisting of Bortezomib, Azacitidine, Dasatinib, and Gefitinib. In a particular embodiment, the pharmaceutical compositions disclosed herein are administered to a patient by intravenous or subcutaneous administration. In other embodiments, the disclosed pharmaceutical compositions are administered to a patient byoral, parenteral, intramuscular, intrarticular, intrabronchial, intraabdominal, intracapsular, intracartilaginous, intracavitary, intracelial, intracerebellar, intracerebroventricular, intracolic, intracervical, intragastric, intrahepatic, intramyocardial, intraosteal, intrapelvic, intrapericardiac, intraperitoneal, intrapleural, intraprostatic, intrapulmonary, intrarectal, intrarenal, intraretinal, intraspinal, intrasynovial, intrathoracic, intrauterine, intravesical, bolus, vaginal, rectal, buccal, sublingual, intranasal, or transdermal administration.
[0028] In one embodiment, the TRAIL receptor agonist protein is administered as a single bolus. In another embodiment, TRAIL receptor agonist protein may be administered over several divided doses. The TRAIL receptor agonist protein can be administered at about 0.1-100 mg/kg. In one embodiment, the TRAIL receptor agonist protein can be administered at a dosage selected from the group consisting of: about 0.1-0.5, 0.1-1, 0.1-10, 0.1-20, 0.1-50, 0.1-75, 1-10, 1-15, 1-7.5, 1.25-15, 1.25-7.5, 2.5-7.5, 2.5-15, 5-15, 5-7.5, 1-20, 1-50, 7-75, 1-100, 5-10, 5-15, 5-20, 5-25, 5-50, 5-75, 10-20, 10-50, 10-75, and 10-100 mg/kg. In other embodiments, the TRAIL receptor agonist protein is present in pharmaceutical compositions at about 0.1-100 mg/ml. In one embodiment, the TRAIL receptor agonist protein is present in pharmaceutical compositions at an amount selected from the group consisting of: about 0.1-0.5, 0.1-1, 0.1-10, 0.1-20, 0.1-50, 0.1-75, 1-10, 1-20, 1-50, 1-75, 1-100, 5-10, 5-15, 5-20, 5-25, 5-50, 5-75, 10-20, 10-50, 10-75, or 10-100 mg/ml. In other embodiments, a therapeutically effective amount of TRAIL receptor agonist protein is administered to a subject. In another embodiment, a prophylactically effective amount of TRAIL receptor agonist protein is administered to a subject.
DESCRIPTION OF THE FIGURES
[0029] FIG. 1 Domain structure of a single-chain fusion polypeptide comprising three TRAIL domains. I., II., III. Soluble TRAIL domains.
[0030] FIG. 2 Schematic picture representing the general structure of TRAIL. [0031] .box-solid. .box-solid. .box-solid. Cell membrane, N-terminus located within the cell, [0032] 1. anti-parallel .beta.-fold of receptor-binding domain (RBD), [0033] 2. interface of RBD and cell membrane, [0034] 3. protease cleavage site.
[0035] FIG. 3 Schematic picture representing the structure of the native TRAIL trimer. Cylindric structures represent RBDs. N-termini connect RBDs with the cell membrane.
[0036] FIG. 4 Schematic picture representing the structure of three soluble domains comprising the receptor-binding domain of a TRAIL. I., II., III. soluble TRAIL domains.
[0037] FIG. 5 Trimerization of the soluble domains comprising the RBD of TRAIL, characterized in that the N- and C-termini of the three soluble domains form a surface.
[0038] FIG. 6 Schematic picture representing the structure of the single-chain TRAIL comprising all or a part of the stalk-region illustrating the requirement of longer linkers to compensate for the distance to the N-terminus of the next soluble domain.
[0039] FIG. 7 scFv-TRAIL fusion protein known from the art.
[0040] FIG. 8 Fc-TRAIL fusion protein known from the art.
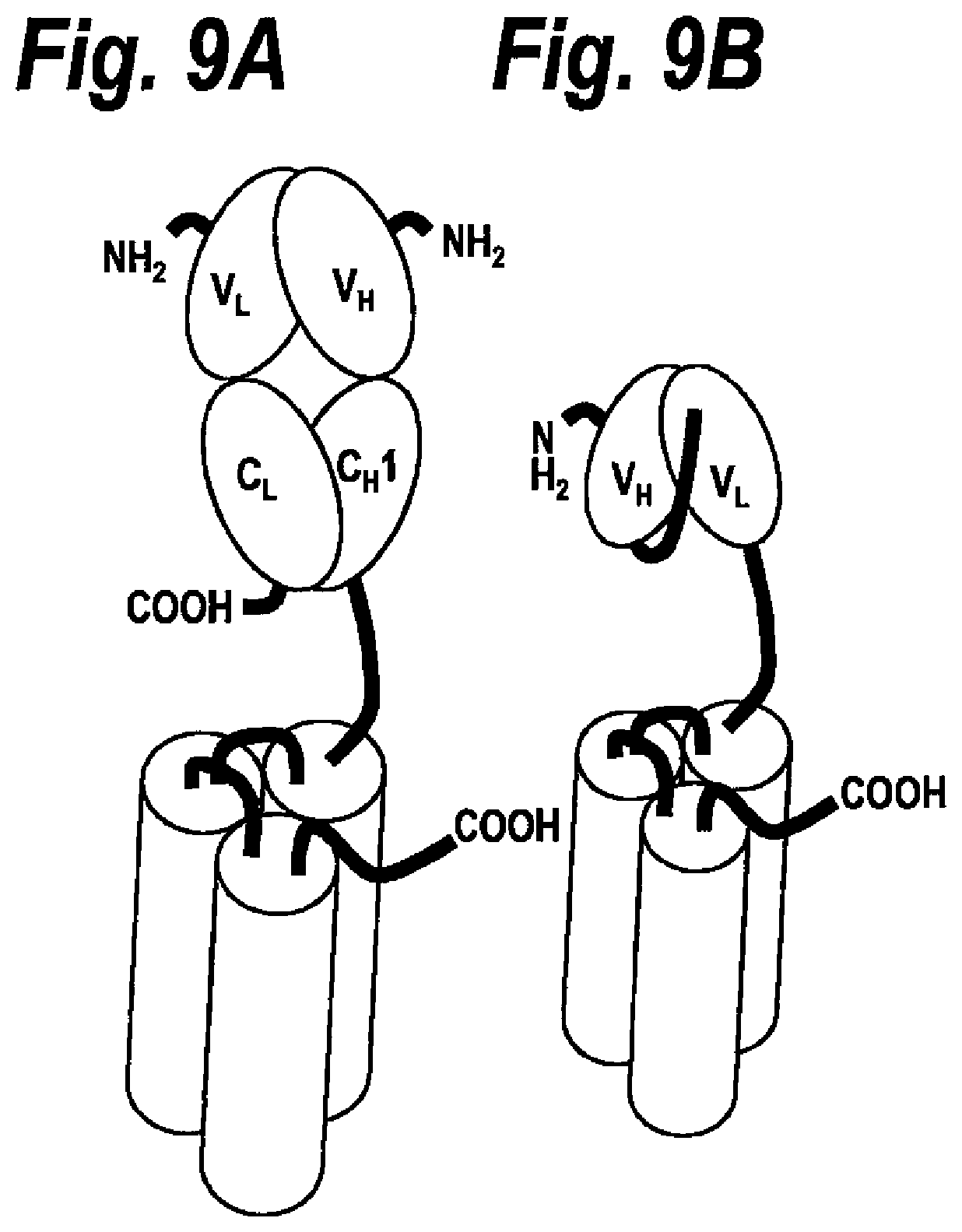
[0041] FIG. 9A Single-chain fusion polypeptide comprising an additional Fab antibody fragment.
[0042] FIG. 9B Single-chain fusion polypeptide comprising an additional scFv antibody fragment.
[0043] FIG. 10 Dimerization of two N-terminally fused scFc fusion polypeptides via disulfide bridges.
[0044] FIG. 11 Dimerization of two C-terminally fused scFc fusion polypeptides via disulfide bridges.
[0045] FIG. 12 Dimerization of single-chain fusion polypeptides via a linker.
[0046] FIG. 13 Single-chain fusion polypeptide comprising an additional Fab antibody fragment further fused to a second fusion polypeptide or to a scFv fusion polypeptide.
[0047] FIG. 14 Dimerization of two scFab fusion polypeptides via disulfide bridges.
[0048] FIG. 15 N-terminally fused scFc fusion polypeptides further comprising a Fv and/or Fab antibody fragment.
[0049] FIG. 16 C-terminally fused scFc fusion polypeptides further comprising a Fv and/or Fab antibody fragment.
[0050] FIG. 17A The exemplary TRAIL receptor agonist protein as shown with the N-terminal signal peptide domain is set forth in SEQ ID NO: 14. The mature protein (which does not include the N-terminal signal peptide domain) is set forth in SEQ ID NO: 19.
[0051] FIG. 17B Schematic picture representing the overall structure and annotated sequence of an exemplary TRAIL receptor agonist protein.
[0052] FIG. 18 Assay setup of the ELISA for the quantitation of the TRAIL-receptor agonists containing an FC-domain.
[0053] FIG. 19 A TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 induces cell death in human tumor cell lines in vitro. SKM-1, Colo205 or Jurkat cells were treated with increasing concentrations the TRAIL receptor agonist protein for 24 hours and cell viability assessed.
[0054] FIGS. 20(A-C) A TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 synergizes with anti-tumorigenic agents in vitro. SU-DHL-4 cells were incubated with increasing concentrations of the TRAIL receptor agonist protein in the presence or absence of the indicated concentrations of venetoclax (FIG. 20A) or navitoclax (FIG. 20B) for 24 hours. Alternatively, (FIG. 20C) NCI-H596 cells were treated with increasing concentrations of the TRAIL receptor agonist protein in the presence or absence of the indicated concentrations of docetaxel (DTX) for 72 hours. Cell viability was assessed and synergy determined by Bliss sum.
[0055] FIG. 21 Effect of TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 on tumor growth in the Colo205 colorectal carcinoma xenograft model.
[0056] FIG. 22 Effect of TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 on tumor growth in the SKM-1 acute myeloid leukemia xenograft model.
[0057] FIG. 23 Effect of TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 on tumor growth in the H460LM non-small cell lung xenograft model.
[0058] FIGS. 24(A-G) Effect of TRAIL receptor agonist protein comprising two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 on tumor growth in PDX models. Diamonds, TRAIL receptor agonist protein-treated; Squares, untreated. Tumor volumes are shown for (A) CTG-0069, (B) CTG-0167, (C) CTG-0293, (D) CTG-0785, (E) CTG-0714, (F) CTG-0136, and (G) CTG-0485.
DETAILED DESCRIPTION OF THE INVENTION
[0059] According to the present invention, it was found that fusing a single-chain TRAIL receptor-binding domain to an Fc domain results in a hexavalent TRAIL receptor agonist providing high biological activity combined with good stability. Accordingly, a single-chain fusion polypeptide comprising at least three soluble TRAIL domains connected by two peptide linkers and N-terminally and/or C-terminally an antibody Fc fragment, is provided.
[0060] Preferably, the single-chain fusion polypeptide is non-aggregating. The term "non-aggregating" refers to a monomer content of the preparation of 50%, preferably 70% and more preferably 90%. The ratio of monomer content to aggregate content may be determined by examining the amount of aggregate formation using size-exclusion chromatography (SEC). The stability concerning aggregation may be determined by SEC after defined time periods, e.g. from a few to several days, to weeks and months under different storage conditions, e.g. at 4.degree. C. or 25.degree. C. For the fusion protein, in order to be classified as substantially non-aggregating, it is preferred that the monomer content is as defined above after a time period of several days, e.g. 10 days, more preferably after several weeks, e.g. 2, 3 or 4 weeks, and most preferably after several months, e.g. 2 or 3 months of storage at 4.degree. C., or 25.degree. C.
[0061] The single-chain fusion polypeptide may comprise additional domains which may be located at the N- and/or C-termini thereof. Examples for additional fusion domains are e.g. an N-terminal signal peptide domain which may comprise a protease cleave site or a C-terminal element which may comprise and/or connect to a recognition/purification domain. According to a preferred embodiment, the fusion polypeptide comprises a Strep-tag at its C-terminus that is fused via a linker. An exemplary Strep-tag including a short serine linker is shown in SEQ ID NO: 13.
[0062] The TRAIL receptor agonist protein of the present invention comprises three soluble domains derived from TRAIL. Preferably, those soluble domains are derived from a mammalian, particularly human TRAIL including allelic variants and/or derivatives thereof. The soluble domains comprise the extracellular portion of TRAIL including the receptor binding domain without membrane located domains. Like other proteins of the TNF superfamily, TRAIL is anchored to the membrane via an N-terminal portion of 15-30 amino acids, the so-called stalk-region. The stalk region contributes to trimerization and provides a certain distance to the cell membrane. However, the stalk region is not part of the receptor binding domain (RBD).
[0063] Importantly, the RBD is characterized by a particular localization of its N- and C-terminal amino acids. Said amino acids are immediately adjacent and are located centrally to the axis of the trimer. The first N-terminal amino acids of the RBD form an anti-parallel beta-strand with the C-terminal amino acids of the RBD (FIGS. 2 and 3).
[0064] Thus, the anti-parallel beta-strand of the RBD forms an interface with the cell membrane, which is connected to and anchored within the cell membrane via the amino acids of the stalk region. It is highly preferred that the soluble TRAIL domains of the TRAIL receptor agonist protein comprise a receptor binding domain of the TRAIL lacking any amino acids from the stalk region (FIGS. 4 and 5). Otherwise, a long linker connecting the C-terminus of one of the soluble domains with the N-terminus of the next soluble domain would be required to compensate for the N-terminal stalk-region of the next soluble domain (FIG. 6), which might result in instability and/or formation of aggregates.
[0065] A further advantage of such soluble domains is that the N- and C-terminal amino acids of the RBD are not accessible for any anti-drug antibodies. Preferably, the single-chain fusion polypeptide is capable of forming an ordered trimeric structure comprising at least one functional binding site for the respective TRAIL receptor.
[0066] The TRAIL receptor agonist protein comprises three functional TRAIL receptor binding sites, i.e. amino acid sequences capable of forming a complex with a TRAIL receptor. Thus, the soluble domains are capable of binding to the corresponding TRAIL receptor. In one embodiment, at least one of the soluble domains is capable of receptor activation, whereby apoptotic and/or proliferative activity may be affected. In a further embodiment, one or more of the soluble domains are selected as not being capable of receptor activation.
[0067] The soluble TRAIL domain may be derived from human TRAIL as shown in SEQ ID NO: 1. Preferably, the soluble TRAIL domains are derived from human TRAIL, particularly starting from amino acids 120-122 and comprise particularly amino acids 120-281, 121-281 or 122-281 of SEQ ID NO: 1. Optionally, amino acid Arg121 of SEQ ID NO: 1 may be replaced by a non-charged amino acid, e.g. Ser or Gly.
TABLE-US-00001 TABLE 1 Sequence of Human TRAIL Protein SEQ ID NO Sequence 1 MAMMEVQGGPSLGQTCVLIVIFTVLLQSLCVAVTYVYFTNE LKQMQDKYSKSGIACFLKEDDSYWDPNDEESMNSPCWQVKW QLRQLVRKMILRTSEETISTVQEKQQNISPLVRERGPQRVA AHITGTRGRSNTLSSPNSKNEKALGRKINSWESSRSGHSFL SNLHLRNGELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQ MVQYIYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVG
[0068] As indicated above, the soluble TRAIL domains may comprise the wild-type sequences as indicated in SEQ ID NO: 1. It should be noted, however, that it is possible to introduce mutations in one or more of these soluble domains, e.g. mutations which alter (e.g. increase or decrease) the binding properties of the soluble domains. In one embodiment, soluble domains may be selected which cannot bind to the corresponding cytokine receptor.
[0069] In a further preferred embodiment of the invention, the soluble TRAIL domain (i) comprises a mutant of TRAIL or a receptor binding domain thereof which binds and/or activates TRAIL-receptor 1 (TRAILR1) and/or TRAIL-receptor 2 (TRAILR2). The binding and/or activity of the mutant may be, e.g., determined by the assays as described in van der Sloot et al. (PNAS, 2006, 103:8634-8639), Kelley et al. (J. Biol. Chem., 2005, 280:2205-2215), or MacFarlane et al. (Cancer Res., 2005, 65: 11265-11270).
[0070] The mutant may be generated by any technique and is known by the skilled person, e.g., the techniques described in van der Sloot et al. (PNAS, 2006, 103:8634-8639), Kelley et al. (J. Biol. Chem., 2005, 280:2205-2215), or MacFarlane et al. (Cancer Res., 2005, 65: 11265-11270) and may comprise any type of structural mutations, e.g., substitution, deletion, duplication and/or insertion of an amino acid. A preferred embodiment is the generation of substitutions. The substitution may affect at least one amino acid of TRAIL or a receptor binding domain thereof as described herein. In a preferred embodiment, the substitution may affect at least one of the amino acids of TRAIL, e.g., human TRAIL (e.g., SEQ ID NO: 1). Preferred substitutions in this regard affect at least one of the following amino acids of human TRAIL of SEQ ID NO: 1: R130, G160, Y189, R191, Q193, E195, N199, K201, Y213, T214, S215, H264, 1266, D267, D269. Preferred amino acid substitutions of human TRAIL of SEQ ID NO:1 are at least one of the following substitutions: R130E, G160M, Y189A, Y189Q, R191K, Q193S, Q193R, E195R, N199V, N199R, K201R, Y213W, T214R, S215D, H264R, I266L, D267Q, D269H, D269R, or D269K.
[0071] The amino acid substitution(s) may affect the binding and/or activity of TRAIL, e.g., human TRAIL, to or on either the TRAILR1 or the TRAILR2. Alternatively, the amino acid substitution(s) may affect the binding and/or activity of TRAIL, e.g., human TRAIL, to or on both, the TRAILR1 and the TRAILR2. The binding and/or activity of the TRAILR1 and/or TRAILR2 may be affected positively, i.e., stronger, more selective or more specific binding and/or more activation of the receptor. Alternatively, the binding and/or activity of the TRAILR1 and/or TRAILR2 may be affected negatively, i.e., weaker, less selective or less specific binding and/or less or no activation of the receptor.
[0072] Examples of mutants of TRAIL with amino acid substitution(s) of the invention that affect binding and/or activation of both TRAILR1 and TRAILR2 may be found, e.g., in Table 1 of MacFarlane et al. (cf. above) and may comprise a human TRAIL mutant with the following two amino acid substitutions of SEQ ID NO: 1 Y213W and S215D or with the following single amino acid substitution: Y189A.
[0073] Examples of mutants of TRAIL with amino acid substitution(s) of the invention that affect binding and/or activation of TRAILR1 may be found, e.g., in Table 1 of MacFarlane et al. (cf. above) and may comprise a human TRAIL mutant with the following four amino acid substitutions of SEQ ID NO: 1 N 199V, K201R, Y213W and S215D or with the following five amino acid substitutions: Q193S, N199V, K201R, Y213W and S215D, or may be found in Table 2 of Kelley et al. (cf. above) and may comprise a human TRAIL mutant with the following six amino acid substitutions: Y213W, S215D, Y189A, Q193S, N199V, and K201R, or with Y213W, S215D, Y189A, Q193S, N199R, and K201R.
[0074] Examples of mutants of TRAIL with amino acid substitution(s) of the invention that affect binding and/or activation of TRAILR2 may be found, e.g., in Table 1 of MacFarlane et al. (cf. above) or in Table 2 of Kelley et al. (cf. above) and may comprise a human TRAIL mutant with the following six amino acid substitutions of SEQ ID NO: 1: Y189Q, R191 K, Q193R, H264R, I266L, and D267Q, or may be found in Table 2 of van der Sloot et al. (cf. above) and may comprise a human TRAIL mutant with the following single amino acid substitution: D269H, or with the following two amino acid substitutions: D269H and E195R or D269H and T214R.
[0075] Thus one preferred embodiment is a TRAIL receptor agonist protein as described herein wherein at least one of the soluble domains comprises a mutant of TRAIL or of a receptor binding domain thereof which binds and/or activates TRAILR1 and/or TRAILR2.
[0076] Further examples of mutants of TRAIL, which show reduced TRAIL induced receptor aggregation are H168 (S, T, Q), R170 (E, S, T, Q) and H177 (S, T).
[0077] One preferred embodiment of a TRAIL receptor agonist protein comprising a mutant of TRAIL or of a receptor binding domain as described herein is a TRAIL receptor agonist protein wherein component (i) comprises at least one amino acid substitution, particularly as indicated below.
[0078] Such an amino acid substitution affects at least one of the following amino acid positions of human TRAIL (SEQ ID NO: 1): R130, G160, H168, R170, H177, Y189, R191, Q193, E195, N199, K201, Y213, T214, S215, H264, 1266, D267, D269.
[0079] Such an amino acid substitution is at least one of the following: R130E, G160M, H168 (S, T, Q), R170 (E, S, T, Q), H177 (SJ)1 Y189A, Y189Q, R191K, Q193S, Q193R, E195R, N199V, N199R, K201R, Y213W, T214R, S215D, H264R, I266L, D267Q, D269H, D269R, or D269K.
[0080] A preferred TRAIL-R2 selective domain comprises amino acid substitutions Y189Q, R191K, Q193R, H264R, I266L and D267Q.
[0081] A preferred TRAIL-R1 selective domain comprises amino acid substitutions Y189A, Q193S, N199V, K201R, Y213W and S215D.
[0082] The single-chain fusion molecule of the present invention comprises three soluble TRAIL domains, namely components (i), (iii) and (v). The stability of a single-chain TRAIL fusion polypeptide against aggregation is enhanced, if the second and/or third soluble TRAIL domain is an N-terminally shortened domain which optionally comprises amino acid sequence mutations. Thus, preferably, both the second and the third soluble TRAIL domain are N-terminally shortened domains which optionally comprise amino acid sequence mutations in the N-terminal regions, preferably within the first five amino acids of the N-terminus of the soluble TRAIL domain. These mutations may comprise replacement of charged, e.g. acidic or basic amino acids, by neutral amino acids, particularly serine or glycine.
[0083] In contrast thereto, the selection of the first soluble TRAIL domain is not as critical. Here, a soluble domain having a full-length N-terminal sequence may be used. It should be noted, however, that also the first soluble TRAIL domain may have an N-terminally shortened and optionally mutated sequence.
[0084] In a further preferred embodiment of the present invention, the soluble TRAIL domains (i), (iii) and (v) are soluble human TRAIL domains. The first soluble TRAIL domain (i) may be selected from native, shortened and/or mutated sequences. Thus, the first soluble TRAIL domain (i) has an N-terminal sequence which may start between amino acid Glu116 and Val122 of human TRAIL, and wherein Arg121 may be replaced by a neutral amino acid, e.g. by Ser or Gly. The second and third soluble TRAIL domains (iii) and (v) have a shortened N-terminal sequence which preferably starts between amino acid GIn120 and Val122 of human TRAIL and wherein Arg121 may be replaced by another amino acid, e.g. Ser or Gly.
[0085] Preferably, the N-terminal sequence of the soluble TRAIL domains (iii) and (v) is selected from:
(a) Arg121-Val122-Ala123 and
(b) (Gly/Ser) 121.
[0086] The soluble TRAIL domain preferably ends with amino acid Gly281 of human TRAIL. In certain embodiments, the TRAIL domain may comprise internal mutations as described above.
[0087] Components (ii) and (iv) of the TRAIL receptor agonist protein are peptide linker elements located between components (i) and (iii) or (iii) and (v), respectively. The flexible linker elements have a length of 3-8 amino acids, particularly a length of 3, 4, 5, 6, 7, or 8 amino acids. The linker elements are preferably glycine/serine linkers, i.e. peptide linkers substantially consisting of the amino acids glycine and serine. In cases in which the soluble cytokine domain terminates with S or G (C-terminus), e.g. human TRAIL, the linker starts after S or G. In cases in which the soluble cytokine domain starts with S or G (N-terminus), the linker ends before this S or G.
[0088] It should be noted that linker (ii) and linker (iv) do not need to be of the same length. In order to decrease potential immunogenicity, it may be preferred to use shorter linkers. In addition it turned out that shorter linkers lead to single chain molecules with reduced tendency to form aggregates. Whereas linkers that are substantially longer than the ones disclosed here may exhibit unfavorable aggregations properties.
[0089] If desired, the linker may comprise an asparagine residue which may form a glycosylate site Asn-Xaa-Ser. In certain embodiments, one of the linkers, e.g. linker (ii) or linker (iv) comprises a glycosylation site. In other embodiments, both linkers (iv) comprise glycosylation sites. In order to increase the solubility of the sc TRAIL proteins and/or in order to reduce the potential immunogenicity, it may be preferred that linker (ii) or linker (iv) or both comprise a glycosylation site.
[0090] Preferred linker sequences are selected from GSGSGSGS (SEQ ID NO: 3), GSGSGNGS (SEQ ID NO: 2), GGSGSGSG (SEQ ID NO: 4), GGSGSG (SEQ ID NO: 5), GGSG (SEQ ID NO: 6), GGSGNGSG (SEQ ID NO: 7), GGNGSGSG (SEQ ID NO: 8), GGNGSG (SEQ ID NO: 9), and GSGS (SEQ ID NO: 23).
[0091] According to a most preferred embodiment, the linker sequences are each GSGSGNGS according to SEQ ID NO: 2. Example linker sequences are shown in Table 2.
TABLE-US-00002 TABLE 2 Example Linker Sequences SEQ ID NO Sequence 2 GSGSGNGS 3 GSGSGSGS 4 GGSGSGSG 5 GGSGSG 6 GGSG 7 GGSGNGSG 8 GGNGSGSG 9 GGNGSG 22 GSGSGS 23 GSGS 24 GSG
[0092] The TRAIL receptor agonist protein additionally comprises an antibody Fc fragment domain which may be located N-terminal to the first TRAIL domain (i) and/or C-terminal to the third TRAIL domain (v). Preferably, the antibody Fc fragment domain comprises or consists of an amino acid sequence as shown in SEQ ID NO: 10. Alternatively, the Fc fragment domain comprises or consists of an amino acid sequence as shown in SEQ ID NO: 17. Example Fc fragment domains are shown in Table 3.
TABLE-US-00003 TABLE 3 Example Fc Fragment Domains SEQ ID NO Sequence 10 PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPE VKFNWYVDGVEVHNAKTKPREEQYSSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRE EMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVL DSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQK SLSLSPGK 17 PAPPVAGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPE VKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLPPSRE EMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVL DSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQK SLSLSPGK
[0093] The total number of glycosites and the individual position of the carbohydrates in three dimensions impacts the in-vivo stability of TRAIL receptor agonist proteins. Further, carbohydrate recognition depends on local density of the terminal saccharides, the branching of the carbohydrate tree and the relative position of the carbohydrates matter.
[0094] Depletion of CH2-domain carbohydrates is necessary in order to avoid Fc-receptor based crosslinking in vivo and potential TRAIL-receptor superclustering-based toxicity. Further, partially degraded carbohydrates reduce the in vivo half-life of TRAIL receptor agonist proteins through lectin-driven mechanisms. By reducing the total number of glycosylation sites on the molecule, the resulting compound is less accessible to these mechanisms, increasing half-life. Accordingly, in one embodiment, the overall number of glycosites on the TRAIL receptor agonist proteins of the instant invention was reduced through the depletion of CH2 glycosites, resulting in TRAIL receptor agonist proteins comprising N297S equivalent mutations (according to the EU numbering system) creating aglycosl-CH2 domains.
[0095] CH2-glycosites present on the inner surface areas normally shield the subdomain from proteases during "open Fc-conformation transits" wherein hinge-interchain disulfide bonds are reduced and the covalent interchain linkage is disrupted. This enables CH2-dissociation and exposure of the inner surface area towards proteases. TRAIL receptor agonist proteins comprising an N297S equivalent mutation (according to the EU numbering system) creating an aglycosl-CH2 are therefore likely to be less proteolytically stable that equivalent structures with wild-type CH2 glycosylation. This would impact the compound's stability during USP/DSP/storage, where host cell proteases are present and have long-term access to the structure. Accordingly, in certain embodiments, the TRAIL receptor agonist lacks CH2 glycosites, but comprises glycosites in the linker sequences of each polypeptide chain (e.g., GSGSGNGS according to SEQ ID NO: 2). In certain exemplary embodiments, the TRAIL receptor agonist comprises two glycosites per polypeptide chain, for a total of four glycosites, According to a preferred embodiment of the invention, the antibody Fc fragment domain is fused via a hinge-linker element. The hinge-linker element has a length of 10-30 amino acids, particularly a length of 15-25 amino acids, e.g. 22 amino acids. The hinge-linker element preferably comprises the hinge-region sequence of an immunoglobulin, herein referred to as "Ig hinge-region". The term "Ig hinge-region" means any polypeptide comprising an amino acid sequence that shares sequence identity or similarity with a portion of a naturally occurring Ig hinge-region sequence which includes the cysteine residues at which the disulfide bonds link the two heavy chains of the immunoglobulin.
[0096] Derivatives and analogues of the hinge-region can be obtained by mutations. A derivative or analogue as referred to herein is a polypeptide comprising an amino acid sequence that shares sequence identity or similarity with the full length sequence of the wild type (or naturally occurring protein) except that it has one or more amino acid sequence differences attributable to a deletion, insertion and/or substitution. According to the present invention, however, the term "hinge-linker" is not limited to those linkers comprising an Ig hinge-region or a derivative thereof, but any linkers long enough to allow the domains attached by the hinge-linker element to attain a biologically active confirmation.
[0097] The number of molecules with open Fc-conformation in an individual TRAIL receptor agonist protein depends on the number of interchain-disulfide bonds present in the hinge region. Accordingly, in one embodiment a third cysteine was introduced into the hinge region of the TRAIL receptor agonist proteins of the instant invention in order to ameliorate the effect of depleting the CH2-glycosites.
[0098] Further, the TRAIL receptor agonist proteins of the invention additionally comprise mutation of the upper-hinge lysine to a glycine to reduce proteolytic processing at this site.
[0099] A particularly preferred hinge-linker element comprises or consists of the amino acid sequence as shown in SEQ ID NO: 11 (Table 4).
[0100] The TRAIL receptor agonist protein may additionally comprise an N-terminal signal peptide domain, which allows processing, e.g. extracellular secretion, in a suitable host cell. Preferably, the N-terminal signal peptide domain comprises a protease cleavage site, e.g. a signal peptidase cleavage site and thus may be removed after or during expression to obtain the mature protein. A particularly preferred N-terminal signal peptide domain comprises the amino acid sequence as shown in SEQ ID NO: 12 (Table 4).
[0101] Further, the TRAIL receptor agonist protein may additionally comprise a C-terminal element, having a length of e.g. 1-50, preferably 10-30 amino acids which may include or connect to a recognition/purification domain, e.g. a FLAG domain, a Strep-tag or
[0102] Strep-tag II domain and/or a poly-His domain. According to a particularly preferred embodiment, the fusion polypeptide comprises a Strep-tag fused to the C-terminus via a short serine linker as shown in SEQ ID NO: 13 (Table 4).
[0103] An exemplary hinge-linker element, N-terminal signal peptide domain, and short serine linker are shown in Table 4.
TABLE-US-00004 TABLE 4 Exemplary domains and linkers SEQ ID NO Sequence 11 GPGSSSSSSSGSCDKTHTCPPC 12 METDTLLVFVLLVWVPAGNG 13 SSSSSSAWSHPQFEK 25 GPGSSSSSSGSCDKTHTCPPC
[0104] According to a particularly preferred embodiment of the invention, the fusion polypeptide comprises three soluble TRAIL domains fused by peptide linker elements of SEQ ID NO: 2. The first soluble TRAIL domain (i) consists of amino acids 120-281 of human TRAIL according to SEQ ID NO: 1 and the soluble TRAIL domains (iii) and (v) consist of amino acids 121-281 of human TRAIL according to SEQ ID NO: 1. Additionally, the fusion polypeptide comprises an antibody Fc fragment domain according to SEQ ID NO: 10 that is fused C-terminally to the soluble TRAIL domain (v) via a hinge-linker according to SEQ ID NO: 11. The inventors surprisingly found that this particular fusion polypeptide provides improved biological activity and is particularly stable. The amino acid sequence of an exemplary embodiment of a TRAIL receptor agonist protein of the invention is set forth in SEQ ID NO: 19.
[0105] Further, the fusion polypeptide may comprise an N-terminal signal peptide domain e.g. according to SEQ ID NO: 12. A specific example of a TRAIL receptor agonist protein of the invention is shown in SEQ ID NO: 14.
[0106] According to another preferred embodiment, the fusion polypeptide may additionally comprise a C-terminal Strep-tag that is fused to the polypeptide of the invention via a short serine linker as shown in SEQ ID NO: 13. According to this aspect of the invention, the Fc fragment preferably consists of the amino acid sequence as shown in SEQ ID NO: 10 or 17. Further, the Fc fragment may consist of a shorter Fc fragment, for example including amino acids 1-217 of SEQ ID NO: 10. Particularly preferred examples of fusion polypeptides comprising a C-terminal Strep-tag are shown in SEQ ID NOs: 15 and 18.
[0107] The exemplary TRAIL receptor agonist proteins as shown in SEQ ID NOs: 14, 15 and 18 each comprise an N-terminal signal peptide domain. The signal peptide domain includes amino acids 1-20. In each case, the mature protein starts with amino acid 21. Mature exemplary TRAIL receptor agonist proteins of the instant invention are set forth in SEQ ID NO: 19, 20, 21, 26, 27, 28, 29, and 30. Exemplary TRAIL receptor agonist proteins described above are shown in Table 5.
[0108] The TRAIL receptor agonist as set forth in SEQ ID NO: 19 has a reduced total number of glycosylation sites (the N297S mutation in the CH2 region providing an aglycosylated CH2 domain), an increased number of inter-chain disulfide bonds in the hinge region, and the mutation of an upper-hinge lysine to a glycine. These alterations provide a decrease in potential degradation and TRAIL receptor superclustering (along with concomitant toxicity) while increasing the half-life of the molecule. In some embodiments, the N-terminal glutamine is modified to pyroglutamate (Liu et al. 2011, J. Biol. Chem. 286:11211-11217).
TABLE-US-00005 TABLE 5 Exemplary TRAIL Receptor Agonist Proteins SEQ ID NO Sequence 14 METDTLLVFVLLVWVPAGNGQRVAAHITGTRGRSNTL SSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNGE LVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQYI YKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGGI FELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGSG SGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKIN SWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTY FRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKSA RNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTNE HLIDMDHEASFFGAFLVGGSGSGNGSRVAAHITGTRG RSNTLSSPNSKNEKALGRKINSWESSRSGHSFLSNLH LRNGELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQ MVQYIYKYTSYPDPILLMKSARNSCWSKDAEYGLYSI YQGGIFELKENDRIFVSVTNEHLIDMDHEASFFGAFL VGGPGSSSSSSSGSCDKTHTCPPCPAPELLGGPSVFL FPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV DGVEVHNAKTKPREEQYSSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPP SREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENN YKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK 15 METDTLLVFVLLVVVVPAGNGQRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKI NSWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQT YFRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKS ARNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTN EHLIDMDHEASFFGAFLVGGSGSGNGSRVAAHITGTR GRSNTLSSPNSKNEKALGRKINSWESSRSGHSFLSNL HLRNGELVIHEKGFYYIYSQTYFRFQEEIKENTKNDK QMVQYIYKYTSYPDPILLMKSARNSCWSKDAEYGLYS IYQGGIFELKENDRIFVSVTNEHLIDMDHEASFFGAF LVGGPGSSSSSSSGSCDKTHTCPPCPAPELLGGPSVF LFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNVW YVDGVEVHNAKTKPREEQYSSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTL PPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSC SVMHEALHNHYTQKSLSLSPGSSSSSSAWSHPQFEK 18 METDTLLVFVLLVWVPAGNGQRVAAHITGTRGRSNTL SSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNGE LVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQYI YKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGGI FELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGSG SGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKIN SWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTY FRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKSA RNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTNE HLIDMDHEASFFGAFLVGGSGSGNGSRVAAHITGTRG RSNTLSSPNSKNEKALGRKINSWESSRSGHSFLSNLH LRNGELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQ MVQYIYKYTSYPDPILLMKSARNSCWSKDAEYGLYSI YQGGIFELKENDRIFVSVTNEHLIDMDHEASFFGAFL VGGPGSSSSSSSGSCDKTHTCPPCPAPPVAGPSVFLF PPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVD GVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGK EYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLPPS REEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNY KTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVM HEALHNHYTQKSLSLSPGK 19 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGNGSRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKI NSWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQT YFRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKS ARNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTN EHLIDMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHT CPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCV VVDVSHEDPEVKFNVVYVDGVEVHNAKTKPREEQYSS TYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKT ISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGF YPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSK LTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPG K 20 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGNGSRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKI NSWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQT YFRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKS ARNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTN EHLIDMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHT CPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCV VVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYSST YRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTI SKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFY PSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKL TVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGS SSSSSAWSHPQFEK 21 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGNGSRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKI NSWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQT YFRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKS ARNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTN EHLIDMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHT CPPCPAPPVAGPSVFLFPPKPKDTLMISRTPEVTCVV VDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTY RVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTIS KAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLT VDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 26 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGNGSRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSGNGSRVAAHITGTRGRSNTLSSPNSKNEKALGRKI NSWESSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQT YFRFQEEIKENTKNDKQMVQYIYKYTSYPDPILLMKS ARNSCWSKDAEYGLYSIYQGGIFELKENDRIFVSVTN EHLIDMDHEASFFGAFLVGGPGSSSSSSGSDKTHTCP PCPAPPVAGPSVFLFPPKPKDTLMISRTPEVTCVVVD VSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRV VSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKA KGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSD IAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGSSSS SSAWSHPQFEK 27 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGNGSRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWE SSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRF QEEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNS CWSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLI DMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDV SHEDPEVKFNWYVDGVEVHNAKTKPREEQYSSTYRVV SVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDI AVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDK SRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGSSSSS SAWSHPQFEK 28 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGSRVAAHITGTRGRSNTLS SPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNGEL VIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQYIY KYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGGIF ELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGSGS GSRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWE SSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRF QEEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNS CWSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLI DMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDV SHEDPEVKFNWYVDGVEVHNAKTKPREEQYSSTYRVV SVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDI AVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDK SRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGSSSSS SAWSHPQFEK 29 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGNGSRVAAHITGTRGRSNT LSSPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNG ELVIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQY IYKYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGG IFELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGS GSRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWE SSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRF QEEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNS CWSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLI DMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDV SHEDPEVKFNWYVDGVEVHNAKTKPREEQYSSTYRVV SVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDI AVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDK SRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 30 QRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWES SRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRFQ EEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNSC WSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLID MDHEASFFGAFLVGGSGSGSRVAAHITGTRGRSNTLS SPNSKNEKALGRKINSWESSRSGHSFLSNLHLRNGEL VIHEKGFYYIYSQTYFRFQEEIKENTKNDKQMVQYIY KYTSYPDPILLMKSARNSCWSKDAEYGLYSIYQGGIF ELKENDRIFVSVTNEHLIDMDHEASFFGAFLVGGSGS GSRVAAHITGTRGRSNTLSSPNSKNEKALGRKINSWE SSRSGHSFLSNLHLRNGELVIHEKGFYYIYSQTYFRF QEEIKENTKNDKQMVQYIYKYTSYPDPILLMKSARNS CWSKDAEYGLYSIYQGGIFELKENDRIFVSVTNEHLI DMDHEASFFGAFLVGGPGSSSSSSSGSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDV SHEDPEVKFNWYVDGVEVHNAKTKPREEQYSSTYRVV SVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDI AVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDK SRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK
[0109] A further aspect of the present invention relates to a nucleic acid molecule encoding a TRAIL receptor agonist protein as described herein. The nucleic acid molecule may be a DNA molecule, e.g. a double-stranded or single-stranded DNA molecule, or an RNA molecule. The nucleic acid molecule may encode the TRAIL receptor agonist protein or a precursor thereof, e.g. a pro- or pre-proform of the TRAIL receptor agonist protein which may comprise a signal sequence or other heterologous amino acid portions for secretion or purification which are preferably located at the N- and/or C-terminus of the TRAIL receptor agonist protein. The heterologous amino acid portions may be linked to the first and/or second domain via a protease cleavage site, e.g. a Factor X3, thrombin or IgA protease cleavage site. A specific example of a nucleic acid sequence of the invention is shown in Table 6 as SEQ ID NO: 16. This nucleic acid molecule encodes the fusion polypeptide of SEQ ID NO: 14.
TABLE-US-00006 TABLE 6 Nucleic Acid Sequence of Exemplary TRAIL Receptor Agonist Protein SEQ ID NO Sequence 16 gatatcggtaccgccaccatggaaaccgacaccctgctggtgt tcgtgctgctcgtgtgggtgccagccggcaatggacagagagt ggccgctcatatcaccggcacccggggcagatctaacaccctg tccagccccaactccaagaacgagaaggccctgggccggaaga tcaactcctgggagtcctccagatccggccactcctttctgtc caacctgcacctgagaaacggcgagctggtcatccacgagaag ggcttctactacatctactcccagacctacttcaggtttcagg aagagatcaaagagaacacaaagaacgacaagcagatggtgca gtatatctacaagtacacctcctaccccgaccccatcctgctg atgaagtccgcccggaactcctgctggtccaaggatgctgagt acggcctgtacagcatctaccagggcggcatcttcgagctgaa agagaacgaccggatcttcgtgtccgtgaccaacgagcacctg atcgacatggaccacgaggccagctttttcggcgcctttctcg tgggcggatccggaagcggaaacggcagtagagtggctgccca cattaccggaaccagaggccggtccaacaccctgagcagccct aacagcaaaaatgagaaagctctcgggcgcaagatcaacagct gggaatctagcagaagcggccacagctttctgagcaatctgca tctgcggaacggcgaactcgtgattcatgagaaggggttttat tatatctatagccagacatactttcgattccaggaggaaatca aggaaaacaccaaaaatgataaacagatggtccagtacattta taagtataccagctaccctgatcctatcctcctcatgaagtct gccagaaactcttgttggagcaaggacgccgagtatggactgt actctatctatcagggggggatctttgaactcaaagaaaacga tcgcatctttgtcagcgtcaccaatgagcatctcattgatatg gatcatgaagctagtttcttcggggcattcctcgtgggaggct ccggctctggcaacggatctagagtcgccgcacacatcacagg gaccagaggcagaagcaataccctgtcctccccaaatagtaaa aacgaaaaggcactcggccgcaaaattaattcctgggagagca gcagatccgggcacagttttctgtctaatctccatctgaggaa tggggagctggtgattcacgaaaaaggattttactacatttac agtcagacttactttcgttttcaggaagagattaaggaaaata ccaaaaacgacaagcagatggtccagtacatctataaatacac ctcttatcctgacccaattctgctcatgaagagtgcccgcaac agctgctggtctaaagacgccgaatacgggctgtattccattt accaggggggaatttttgagctgaaggaaaatgatcggatttt tgtctctgtcacaaacgaacacctcatcgatatggatcacgaa gcctctttctttggcgccttcctggtcggaggccctggctcga gttccagctcctcttctggctcctgcgacaagacccacacctg tcccccttgtcctgcccctgaactgctgggcggaccttccgtg ttcctgttccccccaaagcccaaggacaccctgatgatctccc ggacccccgaagtgacctgcgtggtggtggatgtgtctcacga ggaccctgaagtgaagttcaattggtacgtggacggcgtggaa gtgcacaacgccaagaccaagcccagagaggaacagtactcct ccacctaccgggtggtgtctgtgctgaccgtgctgcaccagga ctggctgaacggcaaagagtacaagtgcaaggtgtccaacaag gccctgcctgcccccatcgaaaagaccatctccaaggccaagg gccagccccgggaaccccaggtgtacacactgccccctagccg ggaagagatgaccaagaaccaggtgtccctgacctgcctggtc aagggcttttacccctccgacattgccgtggaatgggagtcca acggccagcctgagaacaactacaagaccaccccccctgtgct ggactccgacggctcattcttcctgtactccaagctgacagtg gacaagtcccggtggcagcagggcaacgtgttctcctgctccg tgatgcacgaggccctgcacaaccactacacccagaagtccct gtccctgagccccggcaaatgatagaagcttgatatc
[0110] The nucleic acid molecule may be operatively linked to an expression control sequence, e.g. an expression control sequence which allows expression of the nucleic acid molecule in a desired host cell. The nucleic acid molecule may be located on a vector, e.g. a plasmid, a bacteriophage, a viral vector, a chromosomal integration vector, etc. Examples of suitable expression control sequences and vectors are described for example by Sambrook et al. (1989) Molecular Cloning, A Laboratory Manual, Cold Spring Harbor Press, and Ausubel et al. (1989), Current Protocols in Molecular Biology, John Wiley & Sons or more recent editions thereof.
[0111] Various expression vector/host cell systems may be used to express the nucleic acid sequences encoding the TRAIL receptor agonist proteins of the present invention. Suitable host cells include, but are not limited to, prokaryotic cells such as bacteria, e.g. E. coli, eukaryotic host cells such as yeast cells, insect cells, plant cells or animal cells, preferably mammalian cells and, more preferably, human cells. Further, the invention relates to a non-human organism transformed or transfected with a nucleic acid molecule as described above. Such transgenic organisms may be generated by known methods of genetic transfer including homologous recombination.
[0112] A further aspect of the present invention relates to a pharmaceutical or diagnostic composition comprising as the active agent at least one TRAIL receptor agonist protein, a respective nucleic acid encoding therefore, or a transformed or transfected cell, all as described herein.
[0113] The term "TRAIL-associated disease or disorder" as used herein is any disease or disorder which may be ameliorated by addition of a TRAIL receptor agonist. At least one TRAIL receptor agonist protein, respective nucleic acid encoding therefore, or transformed or transfected cell, all as described herein may be used in therapy, e.g., in the prophylaxis and/or treatment of disorders caused by, associated with and/or accompanied by dysfunction of TRAIL, particularly proliferative disorders, such as tumors, e.g. solid or lymphatic tumors; infectious diseases; inflammatory diseases; metabolic diseases; autoimmune disorders, e.g. rheumatoid and/or arthritic diseases; degenerative diseases, e.g. neurodegenerative diseases such as multiple sclerosis; apoptosis-associated diseases or transplant rejections.
[0114] The term "dysfunction of TRAIL" as used herein is to be understood as any function or expression of TRAIL that deviates from the normal function or expression of TRAIL, e.g., overexpression of the TRAIL gene or protein, reduced or abolished expression of the TRAIL gene or protein compared to the normal physiological expression level of TRAIL, increased activity of TRAIL, reduced or abolished activity of TRAIL, increased binding of TRAIL to any binding partners, e.g., to a receptor, particularly a TRAIL receptor or another cytokine molecule, reduced or abolished binding to any binding partner, e.g. to a receptor, particularly a TRAIL receptor or another cytokine molecule, compared to the normal physiological activity or binding of TRAIL.
[0115] In various embodiments, a method is provided for diagnosing and/or treating a human subject suffering from a disorder which can be diagnosed and/or treated by targeting TRAIL receptors comprising administering to the human subject a TRAIL receptor agonist protein disclosed herein such that the effect on the activity of the target, or targets, in the human subject is agonistic, one or more symptoms is alleviated, and/or treatment is achieved. The TRAIL receptor agonist proteins provided herein can be used to diagnose and/or treat humans suffering from primary and metastatic cancers, including carcinomas of breast, colon, rectum, lung (e.g., small cell lung cancer "SCLC" and non-small cell lung cancer "NSCLC"), oropharynx, hypopharynx, esophagus, stomach, pancreas, liver, gallbladder and bile ducts, small intestine, urinary tract (including kidney, bladder and urothelium), female genital tract (including cervix, uterus, and ovaries as well as choriocarcinoma and gestational trophoblastic disease), male genital tract (including prostate, seminal vesicles, testes and germ cell tumors), endocrine glands (including the thyroid, adrenal, and pituitary glands), and skin, as well as hemangiomas, melanomas, sarcomas (including those arising from bone and soft tissues as well as Kaposi's sarcoma), tumors of the brain, nerves, eyes, and meninges (including astrocytomas, gliomas, glioblastomas, retinoblastomas, neuromas, neuroblastomas, Schwannomas, and meningiomas), tumors arising from hematopoietic malignancies, acute leukemia, acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), B cell lymphoma, Burkitt's lymphoma, chronic myelocytic leukemia (CML), chronic lymphocytic leukemia (CLL), hairy cell leukemia, Hodgkin's and non-Hodgkin's lymphomas, DLBCL, follicular lymphomas, hematopoietic malignancies, Kaposi's sarcoma, malignant lymphoma, malignant histiocytosis, malignant melanoma, multiple myeloma, paraneoplastic syndrome/hypercalcemia of malignancy, or solid tumors.
[0116] A pharmaceutical composition comprising a TRAIL receptor agonist protein disclosed herein and a pharmaceutically acceptable carrier is provided. In some embodiments, the pharmaceutical composition comprises at least one additional therapeutic agent for treating a disorder. For example, the additional agent may be a therapeutic agent, a chemotherapeutic agent; an imaging agent, a cytotoxic agent, an angiogenesis inhibitor, a kinase inhibitor (including but not limited to a KDR and a TIE-2 inhibitor), a co-stimulation molecule modulator or an immune checkpoint inhibitor (including but not limited to anti-B7.1, anti-B7.2, anti-B7.3, anti-B7.4, anti-CD28, anti-B7RP1, CTLA4-Ig, anti-CTLA-4, anti-PD-1, anti-PD-L1, anti-PD-L2, anti-ICOS, anti-LAG-3, anti-Tim3, anti-VISTA, anti-HVEM, anti-BTLA, LIGHT fusion protein, anti-CD137, anti-CD137L, anti-OX40, anti-OX4OL, anti-CD70, anti-CD27, anti-GAL9, anti-A2AR, anti-KIR, anti-IDO-1, anti-CD20), a dendritic cell/antigen-presenting cell modulator (including but not limited to anti-CD40 antibody, anti-CD40 L, anti-DC-SIGN, anti-Dectin-1, anti-CD301, anti-CD303, anti-CD123, anti-CD207, anti-DNGR1, anti-CD205, anti-DCIR, anti-CD206, anti-ILT7), a modulator for Toll-like receptors (including but not limited to anti-TLR-1, anti-TLR-2, anti-TLR-3, anti-TLR-4, anti-TLR-4, anti-TLR-5, anti-TLR-6, anti-TLR-7, anti-TLR-8, anti-TLR-9), an adhesion molecule blocker (including but not limited to an anti-LFA-1 antibody, an anti-E/L selectin antibody, a small molecule inhibitor), an anti-cytokine antibody or functional fragment thereof (including but not limited to an anti-IL-18, an anti-TNF, or an anti-IL-6/cytokine receptor antibody), a bispecific redirected T cell or NK cell cytotoxicity (including but not limited to a BiTE.RTM.), a chimeric T cell receptor (CAR-T) based therapy, a T cell receptor (TCR)-based therapy, a therapeutic cancer vaccine, methotrexate, cyclosporin, rapamycin, FK506, a detectable label or reporter, a TNF antagonist, an anti-rheumatic, a muscle relaxant, a narcotic, a non-steroid anti-inflammatory drug (NSAID), an analgesic, an anesthetic, a sedative, a local anesthetic, a neuromuscular blocker, an antimicrobial, an antipsoriatic, a corticosteriod, an anabolic steroid, an erythropoietin, an immunization, an immunoglobulin, an immunosuppressive, a growth hormone, a hormone replacement drug, a radiopharmaceutical, an antidepressant, an antipsychotic, a stimulant, an asthma medication, a beta agonist, an inhaled steroid, an epinephrine or analog, a cytokine, or a cytokine antagonist.
[0117] In an embodiment, a method of treating a cancer or in the prevention or inhibition of metastases from the tumors described herein, the TRAIL receptor agonist protein(s) can be used alone or in combination with one or more additional agents, e.g., a chemotherapeutic, radiotherapy, or biological agent. In some embodiments, the agent can include the following: 13-cis-Retinoic Acid; 2-CdA; 2-Chlorodeoxyadenosine; 5-Azacitidine; 5-Fluorouracil; 5-FU; 6-Mercaptopurine; 6-MP; 6-TG; 6-Thioguanine; Abraxane; Accutane.RTM.; Actinomycin-D; Adriamycin.RTM.; Adrucil.RTM.; Afinitor.RTM.; Agrylin.RTM.; Ala-Cod.RTM.; Aldesleukin; Alemtuzumab; ALIMTA; Alitretinoin; Alkaban-AQ.RTM.; Alkeran.RTM.; All-transretinoic Acid; Alpha Interferon; Altretamine; Amethopterin; Amifostine; Aminoglutethimide; Anagrelide; Anandron.RTM.; Anastrozole; Arabinosylcytosine; Ara-C Aranesp.RTM.; Aredia.RTM.; Arimidex.RTM.; Aromasin.RTM.; Arranon.RTM.; Arsenic Trioxide; Arzerra.TM.; Asparaginase; ATRA; Avastin.RTM.; Azacitidine; BCG; BCNU; Bendamustine; Bevacizumab; Bexarotene; BEXXAR.RTM.; Bicalutamide; BiCNU; Blenoxane.RTM.; Bleomycin; Bortezomib; Busulfan; Busulfex.RTM.; C225; Calcium Leucovorin; Campath.RTM.; Camptosar.RTM.; Camptothecin-11; Capecitabine Carac.TM.; Carboplatin; Carmustine; Carmustine Wafer; Casodex.RTM.; CC-5013; CCI-779; CCNU; CDDP; CeeNU; Cerubidine.RTM.; Cetuximab; Chlorambucil; Cisplatin; Citrovorum Factor; Cladribine; Cortisone; Cosmegen.RTM.; CPT-11; Cyclophosphamide; Cytadren.RTM.; Cytarabine; Cytarabine Liposomal; Cytosar-U.RTM.; Cytoxan.RTM.; Dacarbazine; Dacogen; Dactinomycin; Darbepoetin Alfa; Dasatinib; Daunomycin; Daunorubicin; Daunorubicin Hydrochloride; Daunorubicin Liposomal; DaunoXome.RTM.; Decadron; Decitabine; Delta-Cortef.RTM.; Deltasone.RTM.; Denileukin; Diftitox; DepoCyt.TM.; Dexamethasone; Dexamethasone Acetate; Dexamethasone Sodium Phosphate; Dexasone; Dexrazoxane; DHAD; DIC; Diodex; Docetaxel; Doxil.RTM.; Doxorubicin; Doxorubicin Liposomal; Droxia.TM.; DTIC; DTIC-Dome.RTM.; Duralone.RTM.; Duvelisib; Efudex.RTM.; Eligard.TM.; Ellence.TM.; Eloxatin.TM.; Elspar.RTM.; Emcyt.RTM.; Epirubicin; Epoetin Alfa; Erbitux; Erlotinib; Erwinia L-asparaginase; Estramustine; Ethyol Etopophos.RTM.; Etoposide; Etoposide Phosphate; Eulexin.RTM.; Everolimus; Evista.RTM.; Exemestane; Fareston.RTM.; Faslodex.RTM.; Femara.RTM.; Filgrastim; Floxuridine; Fludara.RTM.; Fludarabine; Fluoroplex.RTM.; Fluorouracil; Fluorouracil (cream); Fluoxymesterone; Flutamide; Folinic Acid; FUDR.RTM.; Fulvestrant; Gefitinib; Gemcitabine; Gemtuzumab ozogamicin; Gemzar; Gleevec.TM.; Gliadel.RTM.; Wafer; GM-CSF; Goserelin; Granulocyte-Colony Stimulating Factor (G-CSF); Granulocyte Macrophage Colony Stimulating Factor (G-MCSF); Halotestin.RTM.; Herceptin.RTM.; Hexadrol; Hexalen.RTM.; Hexamethylmelamine; HMM; Hycamtin.RTM.; Hydrea.RTM.; Hydrocort Acetate.RTM.; Hydrocortisone; Hydrocortisone Sodium Phosphate; Hydrocortisone Sodium Succinate; Hydrocortone Phosphate; Hydroxyurea; Ibrutinib; Ibritumomab; Ibritumomab Tiuxetan; Idamycin.RTM.; Idarubicin Ifex.RTM.; Interferon-alpha; Interferon-alpha-2b (PEG Conjugate); Ifosfamide; Interleukin-11 (IL-11); Interleukin-2 (IL-2); Imatinib mesylate; Imidazole Carboxamide; Intron A.RTM.; ipilimumab, Iressa.RTM.; Irinotecan; Isotretinoin; Ixabepilone; Ixempra.TM.; KADCYCLA.RTM.; Kidrolase (t) Lanacort.RTM.; Lapatinib; L-asparaginase; LCR; Lenalidomide; Letrozole; Leucovorin; Leukeran; Leukine.TM.; Leuprolide; Leurocristine; Leustatin.TM.; Lirilumab; Liposomal Ara-C; Liquid Pred.RTM.; Lomustine; L-PAM; L-Sarcolysin; Lupron.RTM.; Lupron Depot.RTM.; Matulane.RTM.; Maxidex; Mechlorethamine; Mechlorethamine Hydrochloride; Medralone.RTM.; Medrol.RTM.; Megace.RTM.; Megestrol; Megestrol Acetate; MEK inhibitors; Melphalan; Mercaptopurine; Mesna; Mesnex.TM.; Methotrexate; Methotrexate Sodium; Methylprednisolone; Meticorten.RTM.; Mitomycin; Mitomycin-C; Mitoxantrone M-Prednisol.RTM.; MTC; MTX; Mustargen.RTM.; Mustine; Mutamycin.RTM.; Myleran.RTM.; Mylocel.TM.; Mylotarg.RTM.; Navitoclax; Navelbine.RTM.; Nelarabine; Neosar.RTM.; Neulasta.TM.; Neumega.RTM.; Neupogen.RTM.; Nexavar.RTM.; Nilandron.RTM.; Nilotinib; Nilutamide; Nipent.RTM.; Nitrogen Mustard Novaldex.RTM.; Nivolumab; Novantrone.RTM.; Nplate; Octreotide; Octreotide acetate; Ofatumumab; Oncospar.RTM.; Oncovin.RTM.; Ontak.RTM.; Onxal.TM.; Oprelvekin; Orapred.RTM.; Orasone.RTM.; Oxaliplatin; Paclitaxel; Paclitaxel Protein-bound; Pamidronate; Panitumumab; Panretin.RTM.; Paraplatin.RTM.; Pazopanib; Pediapred.RTM.; PEG Interferon; Pegaspargase; Pegfilgrastim; PEG-INTRON.TM.; PEG-L-asparaginase; PEMETREXED; Pembrolizumab; Pentostatin; Pertuzumab; Phenylalanine Mustard; Pidilizumab; Platinol.RTM.; Platinol-AQ.RTM.; Prednisolone; Prednisone; Prelone.RTM.; Procarbazine; PROCRIT.RTM.; Proleukin.RTM.; Prolifeprospan 20 with Carmustine Implant; Purinethol.RTM.; BRAF inhibitors; Raloxifene; Revlimid.RTM.; Rheumatrex.RTM.; Rituxan.RTM.; Rituximab; Roferon-A.RTM.; Romiplostim; Rubex.RTM.; Rubidomycin hydrochloride; Sandostatin.RTM.; Sandostatin LAR.RTM.; Sargramostim; Solu-Cortef.RTM.; Solu-Medrol.RTM.; Sorafenib; SPRYCEL.TM.; STI-571; STIVAGRA.TM., Streptozocin; SU11248; Sunitinib; Sutent.RTM.; Tamoxifen Tarceva.RTM.; Targretin.RTM.; Tasigna.RTM.; Taxol.RTM.; Taxotere.RTM.; Temodar.RTM.; Temozolomide Temsirolimus; Teniposide; TESPA; Thalidomide; Thalomid.RTM.; TheraCys.RTM.; Thioguanine; Thioguanine Tabloid.RTM.; Thiophosphoamide; Thioplex.RTM.; Thiotepa; TICE.RTM.; Toposar.RTM.; Topotecan; Toremifene; Torisel.RTM.; Tositumomab; Trastuzumab; Treanda.RTM.; Tremelimumab; Tretinoin; Trexall.TM.; Trisenox.RTM.; TSPA; TYKERB.RTM.; Urelumab; VCR; Vectibix.TM. Velban.RTM.; Velcade.RTM.; Venetoclax; VePesid.RTM.; Vesanoid.RTM.; Viadur.TM.; Vidaza.RTM.; Vinblastine; Vinblastine Sulfate; Vincasar Pfs.RTM.; Vincristine; Vinorelbine; Vinorelbine tartrate; VLB; VM-26; Vorinostat; Votrient; VP-16; Vumon.RTM.; Xeloda.RTM.; Zanosar.RTM.; Zevalin.TM. Zinecard.RTM.; Zoladex.RTM.; Zoledronic acid; Zolinza; or Zometa.RTM., and/or any other agent not specifically listed here that target similar pathways.
[0118] When two or more substances or principles are to be used as part of a combined treatment regimen, they can be administered via the same route of administration or via different routes of administration, at essentially the same time or at different times (e.g. essentially simultaneously, consecutively, or according to an alternating regime). When the substances or principles are to be administered simultaneously via the same route of administration, they may be administered as different pharmaceutical formulations or compositions or part of a combined pharmaceutical formulation or composition, as will be clear to the skilled person.
[0119] Also, when two or more active substances or principles are to be used as part of a combined treatment regimen, each of the substances or principles may be administered in the same amount and according to the same regimen as used when the compound or principle is used on its own, and such combined use may or may not lead to a synergistic effect. However, when the combined use of the two or more active substances or principles leads to a synergistic effect, it may also be possible to reduce the amount of one, more than one, or all of the substances or principles to be administered, while still achieving the desired therapeutic action. This may, e.g., be useful for avoiding, limiting or reducing any unwanted side-effects that are associated with the use of one or more of the substances or principles when they are used in their usual amounts, while still obtaining the desired pharmaceutical or therapeutic effect.
[0120] The effectiveness of the treatment regimen used according to the invention may be determined and/or followed in any manner known per se for the disease or disorder involved, as will be clear to the clinician. The clinician will also be able, where appropriate and on a case-by-case basis, to change or modify a particular treatment regimen, so as to achieve the desired therapeutic effect, to avoid, limit or reduce unwanted side-effects, and/or to achieve an appropriate balance between achieving the desired therapeutic effect on the one hand and avoiding, limiting or reducing undesired side effects on the other hand.
[0121] Generally, the treatment regimen will be followed until the desired therapeutic effect is achieved and/or for as long as the desired therapeutic effect is to be maintained. Again, this can be determined by the clinician.
[0122] In various embodiments, pharmaceutical compositions comprising one or more TRAIL receptor agonist proteins, either alone or in combination with prophylactic agents, therapeutic agents, and/or pharmaceutically acceptable carriers are provided herein. In various embodiments, nonlimiting examples of the uses of the pharmaceutical compositions disclosed herein include diagnosing, detecting, and/or monitoring a disorder, preventing, treating, managing, and/or ameliorating a disorder or one or more symptoms thereof, and/or in research. The formulation of pharmaceutical compositions, either alone or in combination with prophylactic agents, therapeutic agents, and/or pharmaceutically acceptable carriers, are known to one skilled in the art (US Patent Publication No. 20090311253 A1).
[0123] In various embodiments, a pharmaceutical formulation can comprise one or more amino acid, one or more polysaccharide and/or polysorbate, and a TRAIL receptor agonist protein present at a concentration of between about 0.1 and 100 mg/ml, inclusive of endpoints (e.g., 0.1-10, 1-10, 0.01-50, 1-50, 1-100, 10-100, 25-100, 25-50, or 50-100 mg/ml), where the formulation is at a pH between about 5.0 and 7.0, inclusive of endpoints (e.g., a pH of about 5.0-6.0, 5.5-6.0, 5.0-6.5, 5.5-6.5, or 6.0-7.0). In an embodiment, at least one amino acid in the formulation is histidine and is present at a concentration of about 10-20 mM, 10-15 mM, 15-20 mM, or about 15 mM. In an embodiment, at least one polysaccharide in the formulation is sucrose and is present at a concentration of about 0-8.0% weight/volume (w/v). In an embodiment, the polysorbate in the formulation is polysorbate 80 and is at a concentration of about 0-0.06% w/v. In an embodiment, at least one amino acid in the formulation is arginine and is present at a concentration of about 0-1.5% w/v (e.g., 0.5-1.5, 1.0-1.5, or 0.5-1.0 w/v). In an embodiment, the TRAIL receptor agonist protein is present in the formulation at a concentration of about 0.1-100 mg/ml, (e.g., about 1-100 mg/ml, or about 1-15 mg/ml, or about 1-7.5 mg/ml, or about 2.5-7.5 mg/ml, or about 5-7.5 mg/ml, or about 25-100 mg/ml, or about 20-60 mg/ml, or about 25-50 mg/ml, or about 25 mg/ml, or about 50 mg/ml, or about 0.1-60 mg/ml, or about 0.1-25 mg/ml, or about 1.0-60 mg/ml, or about 0.5-60 mg/ml, or about 0.1-2.0 mg/ml, or about 0.5-2.0 mg/ml, or about 1-5 mg/ml, or about 1-7.5 mg/ml, or about 1-15 mg/ml, or about 0.5 mg/ml, or about 1.0 mg/m).
[0124] As used herein, the phrase "effective amount" means an amount of TRAIL agonist protein that results in a detectable improvement (e.g., at least about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, or more from baseline) in one or more parameters associated with a dysfunction of TRAIL or with a TRAIL-associated disease or disorder.
[0125] In various embodiments, the pharmaceutical formulation is an aqueous formulation, a lyophilized formulation, or a lyophilized and rehydrated formulation. In an embodiment, the hydrating solution is dextrose and/or saline (e.g., dextrose at a concentration of about 5% w/v and/or the saline at a concentration of about 0.9% w/v). In an embodiment, the pharmaceutical formulation comprises about 15 mM histidine, about 0.03% (w/v) polysorbate 80, about 4% (w/v) sucrose, and about 0.1-25 mg/ml of the TRAIL receptor agonist protein, or about 1-15 mg/ml of TRAIL receptor agonist protein, and is at a pH of about 6. In an embodiment, the formulation further comprises at least one additional agent.
[0126] In various embodiments, a formulation is used containing about 25 mg/ml TRAIL receptor agonist protein, about 15 mM histidine, 0.03% polysorbate 80 (weight/volume, w/v), 4.0% sucrose (w/v), and a pH of about 6.0. In some embodiments, the formulation does not comprise arginine. In some embodiments, the formulation exhibits unexpectedly improved freeze-thaw stability, liquid formulation stability, and/or lyophilized formulation stability, as compared to other formulations comprising other components or concentrations.
[0127] Methods of administering a therapeutic agent provided herein include, but are not limited to, oral administration, parenteral administration (e.g., intradermal, intramuscular, intraperitoneal, intravenous and subcutaneous), epidural administration, intratumoral administration, mucosal administration (e.g., intranasal and oral routes) and pulmonary administration (e.g., aerosolized compounds administered with an inhaler or nebulizer). The formulation of pharmaceutical compositions for specific routes of administration, and the materials and techniques necessary for the various methods of administration are available and known to one skilled in the art (US Patent Publication No. 20090311253 A1).
[0128] In various embodiments, dosage regimens may be adjusted to provide for an optimum desired response (e.g., a therapeutic or prophylactic response). For example, a single bolus may be administered, several divided doses may be administered over time or the dose may be proportionally reduced or increased as indicated by the exigencies of the therapeutic situation. In some embodiments, parenteral compositions are formulated in dosage unit form for ease of administration and uniformity of dosage. The term "dosage unit form" refers to physically discrete units suited as unitary dosages for the mammalian subjects to be treated; each unit containing a predetermined quantity of active compound calculated to produce the desired therapeutic effect in association with the required pharmaceutical carrier.
[0129] An exemplary, non-limiting range for a therapeutically or prophylactically effective amount of a TRAIL receptor agonist protein provided herein is about 0.1-100 mg/kg, (e.g., about 0.1-0.5, 0.1-1, 0.1-10, 0.1-20, 0.1-50, 0.1-75, 1-10, 1-15, 1-7.5, 1.25-15, 1.25-7.5, 2.5-7.5, 2.5-15, 5-15, 5-7.5, 1-20, 1-50, 7-75, 1-100, 5-10, 5-15, 5-20, 5-25, 5-50, 5-75, 10-20, 10-50, 10-75, or 10-100 mg/kg, or any concentration in between). In some embodiments, the TRAIL receptor agonist protein is present in a pharmaceutical composition at a therapeutically effective concentration, e.g., a concentration of about 0.1-100 mg/ml (e.g., about 0.1-0.5, 0.1-1, 0.1-10, 0.1-20, 0.1-50, 0.1-75, 1-10, 1-20, 1-50, 1-75, 1-100, 5-10, 5-15, 5-20, 5-25, 5-50, 5-75, 10-20, 10-50, 10-75, or 10-100 mg/ml, or any concentration in between). Note that dosage values may vary with the type and/or severity of the condition to be alleviated. It is to be further understood that for any particular subject, specific dosage regimens may be adjusted over time according to the individual need and/or the professional judgment of the person administering or supervising the administration of the compositions, and that dosage ranges set forth herein are exemplary only and are not intended to limit the scope or practice of the claimed composition.
Examples
1. Manufacture of a TRAIL Receptor Agonist Protein (Sc TRAIL Wt)
1.1 Polypeptide Structure
[0130] A) Amino acids Met1-Gly20 [0131] Ig-Kappa-signal peptide, assumed signal peptidase cleavage site after amino acid Gly 20. B) Amino acids GIn21-Gly182 [0132] First soluble cytokine domain of the human TRAIL ligand (TRAIL, amino acid 120-281 of SEQ ID NO: 1). C) Amino acids Gly 183-Ser 190 [0133] First peptide linker element of SEQ ID NO: 2. D) Amino acids Arg191-Gly351 [0134] Second soluble cytokine domain of the human TRAIL ligand (TRAIL, amino acids 121-281 of SEQ ID NO: 1). E) Amino acids Gly352-Ser359. [0135] Second peptide linker element of SEQ ID NO: 2. F) Amino acids Arg360-Gly520 [0136] Third soluble cytokine domain of the human TRAIL ligand (TRAIL, amino acids 121-281 of SEQ ID NO: 1). G) Amino acids Gly521-Cys542 [0137] Hinge-linker element of SEQ ID NO: 11. H) Amino acids Pro543-Lys760 [0138] Antibody Fc fragment domain of SEQ ID NO: 10.
[0139] The above TRAIL receptor agonist protein is shown in SEQ ID NO: 14.
[0140] The indicated linkers may be replaced by other preferred linkers, e.g. as shown in SEQ ID NOs: 3-9.
[0141] It should be noted that the first and second peptide linkers do not need to be identical.
[0142] The signal peptide sequence (A) may be replaced by any other suitable, e.g. mammalian signal peptide sequence.
1.2 Gene Cassette Encoding the Polypeptide
[0143] The synthetic gene may be optimized in view of its codon usage for the expression in suitable host cells, e.g. insect cells or mammalian cells. A preferred nucleic acid sequence is shown in SEQ ID NO: 16.
2. Expression and Purification
Cloning, Expression and Purification of Fusion Polypeptides
[0144] The aforementioned fusion proteins were expressed recombinantly in two different eukaryotic host cells:
[0145] For initial analysis of aforementioned TRAIL receptor agonist fusion proteins, Hek293T cells grown in DMEM+GlutaMAX (GibCo) supplemented with 10% FBS, 100 units/ml Penicillin and 100 [mu]g/ml Streptomycin were transiently transfected with a plasmid containing an expression cassette for a fusion polypeptide and an appropriate selection marker, e.g. a functional expression cassette comprising a blasticidine, puromycin or hygromycin resistance gene. In those cases, where a plurality of polypeptide chains is necessary to achieve the final product, the expression cassettes were either combined on one plasmid or positioned on different plasmids during the transfection. Cell culture supernatant containing recombinant fusion polypeptide was harvested three days post transfection and clarified by centrifugation at 300.times.g followed by filtration through a 0.22 .mu.m sterile filter.
[0146] For larger scale expression of TRAIL receptor agonist fusion proteins to be used in vivo, synthetic DNA cassettes encoding the aforementioned proteins were inserted into eukaryotic expression vectors comprising appropriate selection markers (e.g. a functional expression cassette comprising a blasticidin, puromycin or hygromycin resistance gene) and genetic elements suitable to enhance the number of transcriptionally active insertion sites within the host cells genome. The sequence verified expression vectors were introduced by electroporation into suspension adapted Chinese Hamster Ovary cells (CHO-S, Invitrogen). Appropriate selection pressure was applied three days post-transfection to the transfected cells. Surviving cells carrying the vector derived resistance gene(s) were recovered by subsequent cultivation under selection pressure. Upon stable growth of the selected cell pools in chemically defined medium (PowerCHO2-CD, Lonza) at 37.degree. C. and 7% CO2 atmosphere in an orbital shaker incubator (100 rpm, 50 mm shaking throw), the individual supernatants were analysed by ELISA-assays detecting the aforementioned proteins and the cell pools with the highest specific productivity were expanded in shake flasks prior to protein production (orbital shaker, 100 rpm, shaking throw 50 mm).
[0147] For lab-scale protein production, individual cell pools were cultured for 7-12 days in chemically defined medium (PowerCHO2-CD, Lonza) at 37.degree. C. and 7% CO2 atmosphere in a Wave bioreactor 20/50 EHT (GE-Healthcare). The basal medium was PowerCHO2-CD supplemented with 4 mM Glutamax. Wave culture started with a viable cell concentration of 0.3 to 0.4.times.10e6 cells/ml and the following settings (for a five- or ten liter bag): shaking frequency 18 rpm, shaking ankle 7.degree., gas current 0.2-0.3 L/min, 7% CO2, 36.5.degree. C. During the Wave run, the cell culture were fed twice with PowerFeed A (Lonza), usually on day 2 (20% feed) and day 5 (30% feed). After the second feed, shaking frequency was increased to 22 rpm, as well as the shaking ankle to 8.degree..
[0148] The bioreactor was usually harvested in between day 7 to day 12 when the cell viability dropped below 80%. First, the culture supernatant was clarified using a manual depth filtration system (Millipore Millistak Pod, MCOHC 0.054 m.sup.2). For Strep-tagged proteins, Avidin was added to a final concentration of 0.5 mg/L. Finally, the culture supernatant containing the TRAIL receptor agonist fusion protein was sterile filtered using a bottle top filter (0.22 .mu.m, PES, Corning) and stored at 2-8.degree. C. until further processing.
[0149] For affinity purification Streptactin Sepharose was packed to a column (gel bed 1 ml), equilibrated with 15 ml buffer W (100 mM Tris-HCl, 150 mM NaCl, pH 8.0) or PBS pH 7.4 and the cell culture supernatant was applied to the column with a flow rate of 4 ml/min. Subsequently, the column was washed with 15 ml buffer W and bound polypeptide was eluted stepwise by addition of 7.times.1 ml buffer E (100 mM Tris HCl, 150 mM NaCl, 2.5 mM Desthiobiotin, pH 8.0). Alternately, PBS pH 7.4 containing 2.5 mM Desthiobiotin can be used for this step.
[0150] Alternately to the Streptactin Sepharose based method, the affinity purification was performed employing a column with immobilized Protein-A as affinity ligand and a Akta chromatography system (GE-Healthcare). A solid phase material with high affinity for the FC-domain of the fusion protein was chosen: MABSelect Sure.TM. (GE Healthcare). Briefly, the clarified cell culture supernatant was loaded on a HiTrap MabSelectSure column (CV=5 ml) equilibrated in wash-buffer-1 (20 mM Pi, 95 mM NaCl, pH7.2) not exceeding a load of 10 mg fusion protein per ml column-bed. The column was washed with ten column-volumes (10CV) of aforementioned equilibration buffer followed by four column-volumes (4CV) of wash-buffer-2 (20 mM Pi, 95 mM NaCl, pH 8.0) to deplete host-cell protein and host-cell DNA. The column was then eluted with elution buffer (20 mM Pi, 95 mM NaCl, pH 3.5) and the eluate was collected in up to ten fractions with each fraction having a volume equal to column-bed volume (5 ml). Each fraction was neutralized with an equal volume of aforementioned wash-buffer-2. The linear velocity was set to 150 cm/h and kept constant during the aforementioned affinity chromatography method.
[0151] The protein amount of the eluate fractions was quantitated and peak fractions were concentrated by ultrafiltration and further purified by size exclusion chromatography (SEC).
[0152] SEC was performed on Superdex 200 10/300 GL or HiLoad 26/60 columns using an Akta chromatography system (GE-Healthcare). The columns were equilibrated with phosphate buffered saline and the concentrated, affinity-purified polypeptide was loaded onto the SEC column with the sample volume not exceeding 2% (v/v) of the column-volume. In the case of Superdex200 10/300 GL columns (GE Healthcare), a flow rate of 0.5 ml per minute was applied. In the case of HiLoad 26/60 Superdex200 columns, a flow rate of 2.5 ml per minute was applied. The elution profile of the polypeptide was monitored by absorbance at 280 nm
[0153] For determination of the apparent molecular weight of purified fusion polypeptide under native conditions a Superdex 200 column was loaded with standard proteins of known molecular weight. Based on the elution volume of the standard proteins a calibration curve was plotted and the apparent molecular weight of purified fusion polypeptide was determined. The FC-domain comprising TRAIL receptor agonist fusion proteins typically eluted from the Supoerdex200 columns with an apparent molecular weight for the homodimer of approx. 160-180 kDa.
3. Apoptosis Assay
[0154] A cellular assay with a Jurkat A3 permanent T-cell line was used to determine the apoptosis inducing activity of the TRAIL-receptor agonist fusion proteins. Jurkat cells were grown in flasks with RPMI 1640-medium+GlutaMAX (GibCo) supplemented with 10% FBS, 100 units/ml Penicillin and 100 .mu.g/ml Streptomycin. Prior to the assay, 100,000 cells were seeded per well into a 96-well microtiterplate. The addition of different concentrations of fusion peptides to the wells was followed by a 3 hour incubation at 37.degree. C. Cells were lysed by adding lysis buffer (250 mM HEPES, 50 mM MgCl2, 10 mM EGTA, 5% Triton-X-100, 100 mM DTT, 10 mM AEBSF, pH 7.5) and plates were put on ice for 30 minutes to 2 hours. Apoptosis is paralleled by an increased activity of caspases, e.g. Caspase-3. Hence, cleavage of the specific caspase substrate Ac-DEVD-AFC (Biomol) was used to determine the extent of apoptosis. In fact, Caspase activity correlates with the percentage of apoptotic cells determined morphologically after staining the cells with propidium iodide and Hoechst-33342. For the caspase activity assay, 20 .mu.l cell lysate was transferred to a black 96-well microtiterplate. After the addition of 80 .mu.l buffer containing 50 mM HEPES, 1% Sucrose, 0.1% CHAPS, 50 .mu.M Ac-DEVD-AFC, and 25 mM DTT, pH 7.5, the plate was transferred to a Tecan Infinite 500 microtiterplate reader and the increase in fluorescence intensity was monitored (excitation wavelength 400 nm, emission wavelength 505 nm).
3.1 Cell Death Assay
[0155] For the determination of cell death in HT1080 fibrosarcoma cells 15,000 cells were plated in 96-well plates overnight in RPM1 1640-medium+GlutaMAX (GibCo) supplemented with 10% FBS (Biochrom). Cells were coincubated with cycloheximide (Sigma) at a final concentration of 2.5 g/ml. Cell death was quantified by staining with buffer KV (0.5% crystal violet, 20% methanol). After staining, the wells were washed with water and air-dried. The dye was eluted with methanol and optical density at 595 nm was measured with an ELISA reader.
4. Stability/Aggregation Test
4.1 Principle of the Aggregation Analysis (Definition for Soluble Protein)
[0156] The content of monomers (defined trimeric assembly of TRAIL receptor binding modules) and aggregates is determined by analytical SEC as described in Example 2. For this particular purpose the analysis is performed in buffers containing physiological salt concentrations at physiological pH (e.g. 0.9% NaCl, pH 7.4; PBS pH 7.4). A typical aggregation analysis is done on a Superdex200 column (GE Healthcare). This column separates proteins in the range between 10 to 800 kDa.
[0157] For determination of the apparent molecular weight of purified fusion polypeptide under native conditions a Superdex 200 column is loaded with standard proteins of known molecular weight. Based on the elution volume of the standard proteins a calibration curve is plotted and the apparent molecular weight of purified fusion polypeptide is calculated based on the elution volume.
[0158] SEC analysis of soluble, non-aggregated proteins, e.g. trimeric TRAIL, typically shows a distinct single protein peak at a defined elution volume. This elution volume corresponds to the apparent native molecular weight of the particular protein and approximately complies to the theoretical molecular weight calculated on the basis of the primary amino acid sequence.
[0159] If protein aggregation occurs the SEC analysis shows additional protein peaks with lower retention volumes. For TRAIL, the aggregation of soluble proteins occurs in a characteristic manner. The proteins tend to form oligomers of the "trimers", forming nonamers (3.times.3) and 27mers (3.times.9). These oligomers serve as aggregation seeds and a high content of oligomers potentially leads to aggregation of the protein. Oligomers of large molecular weight and aggregates elute in the void volume of the Superdex200 column and cannot be analysed by SEC with respect to their native molecular weight.
[0160] Due to the induction of (complete) aggregation, purified preparations of TRAIL-SF fusion proteins should preferably contain only defined trimeric proteins and only a very low amount of oligomerised protein. The degree of aggregation/oligomerisation of a particular TRAIL-SF protein preparation is determined on basis of the SEC analysis by calculating the peak areas of the OD280 diagram for the defined trimeric and the oligomer/aggregate fraction, respectively. Based on the total peak area the percentage of defined trimeric protein is calculated as follows:
(% Trimer content=[Peak area trimer]/[Total peak area].times.100)
[0161] The definition for soluble protein as used in this text, describes a protein preparation of purified TRAIL protein in a buffer of physiological salt concentrations at physiological pH that contains a defined soluble protein (trimeric assembly of TRAIL domains) content of >90% within a typical protein concentration range from 0.2 to 10.0 mg/ml.
5. Half-Life Determination
[0162] Molecules A-D are each made up of two polypeptides covalently linked by interchain disulfide bonds. The number of glycosites and hinge cysteines (resulting in interchain disulfide bonds between proteins) were tested in order to determine the effect that altering these characteristics has on the half-life of these compounds.
[0163] Female NMRI mice were treated with 1.2 mg/kg bw and/or with 4 mg/kg bw of the specified compounds as a single intravenous bolus injection. Whole blood was collected before application (predose), and up to 168 hours after test item administration. Serum was prepared and samples were stored at -80.degree. C. until determination of serum concentrations. Pharmacokinetic parameters were calculated using the mean serum concentrations and the pharmacokinetic evaluation program PK Solutions Version 2.0 for non-compartmental pharmacokinetic data analysis (Summit Research Services, Montrose, Colo.). PK Solutions is an automated, Excel-based application, which computes pharmacokinetic parameters from concentration-time data obtained from analysis of e.g. biological samples following intravenous or extra-vascular routes of administration. PK Solutions calculates results without presuming any specific compartmental model.
[0164] Quantitation of the test items in serum was performed with an ELISA-assay detecting the individual TRAIL-receptor agonists shown in Table 7 independent of a Strep-Tag being part of the molecules. The general layout is shown in FIG. 18. The results are summarized in Table 7.
[0165] Molecule A (made up of two polypeptides of SEQ ID NO:26) has two hinge cysteines (forming two interchain disulfide bonds) and an N residue at position 297 of the Fc region (according to the EU index), resulting in wild-type CH2 glycosylation. Molecule A also has glycosites at positions 168 and 337. Molecule B (made up of two polypeptides of SEQ ID NO: 19) has three hinge cysteines (forming three interchain disulfide bonds) (at positions 513, 519, and 522) and an N297S mutation at position 297 of the Fc region (according to the EU index), resulting in aglycosylation of the CH2 domain. Molecule B also has glycosites at positions 168 and 337. Molecule C (made up of two polypeptides of SEQ ID NO: 27) has three hinge cysteines (forming three interchain disulfide bonds) and an N297S mutation at position 297 of the Fc region (according to the EU index), resulting in aglycosylation of the CH2 domain. Further, there is a glycosite at position 168 (linker 1), but not at position 337 (linker 2). Molecule D (made up of two polypeptides of SEQ ID NO:28) has three hinge cysteines (forming three interchain disulfide bonds) and an N297S mutation at position 297 of the Fc region (according to the EU index), resulting in aglycosylation of the CH2 domain. Further, the glycosites on both linker 1 and linker 2 (positions 168 and 337, respectively) have been depleted in Molecule D.
[0166] The in vivo stability (as judged by compound half-life) of Molecule B (both linkers glycosylated, CH2 glycosites depleted, and the addition of a third hinge cysteine) was enhanced when compared to Molecule A. Further, the depletion of all glycosites in from the compound (Molecule D) resulted in reduced in vivo stability and low productivity during transient expression. Molecule C (first linker glycosylated, second linker aglycosylated, CH2 glycosites depleted) demonstrated an intermediate in vivo stability when compared to Molecules B and D (see results in Table 7).
TABLE-US-00007 TABLE 7 Results of Compound Half-life Testing in NMRI-mice Terminal Number of Terminal Half- Half-life glycosylation Number of life 4 mg/kg 1.2 mg/kg Molecule sites hinge cysteines i.v. (hour) i.v. (hour) A 6 2 23.1 17.7 B 4 3 33.94 28.28 C 2 3 21.03 -- D 0 3 8.81 --
[0167] These experimental results demonstrate that combining linker glycosylation (in both linkers 1 and 2) with a third interchain disulfide bond (through the addition of a hinge cysteine) and the deglycosylation of the CH2 domain in the Fc region results in greater in vivo stability in the molecules of the instant invention.
6. In Vitro Demonstration of Efficacy
6.1 TRAIL Receptor Agonist Protein of SEQ ID NO: 19 Inhibits Human Hematologic and Solid Tumor Cell Survival In Vitro
[0168] Tumor cells were seeded at 10,000 cells/well in 96-well plates in the recommended media containing 10% FBS and treated with a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 for 24 hours at 37.degree. C. in a humidified, 5% CO2 atmosphere. Cell viability was subsequently assessed using CellTiter-Glo.RTM. reagent as described by the manufacturer's instructions (Promega; Madison, Wis.). IC.sub.50 values were determined by non-linear regression analysis of the concentration response data normalized to non-treated control cells. Examples of resulting concentration response curves for Colo205, Jurkat and SKM-1 cells that demonstrate a loss in cell viability in response to TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 treatment are shown in FIG. 19. Table 8 shows the results of hematologic (A; (n=40; Non-Hodgkin's Lymphoma, NHL; Acute Myeloid Lymphoma, AML; Acute Lymphoblastic Leukemia, ALL) and solid tumor (B; (n=44; Non-Small Cell Lung Carcinoma, NSCLC; Pancreatic; Colorectal Cancer, CRC; Breast Cancer, BrCa; Ovarian, Fibrosarcoma; Head and Neck, H&N; Small Cell Lung Cancer, SCLC) cell lines treated with a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 for 24 hours and viability assessed by CellTiter-Glo.RTM.. Resulting IC.sub.50s for TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19-mediated effects on tumor cell viability are presented.
TABLE-US-00008 TABLE 8 Potency of TRAIL receptor agonist protein of SEQ ID NO: 19 in human tumor cancer cell lines in vitro Tumor cell SEQ ID NO: 19 line Tumor Type IC.sub.50 (ng/ml) A SU-DHL-8 NHL 1.36 NUDHL-1 NHL 6.50 OCl-Ly8 NHL 7.49 ULA NHL 8.44 OCl-Ly2 NHL 18.98 OCl-LY19 NHL 26.34 WSU-NHL NHL 31.60 OCl-Ly7 NHL 63.76 SU-DHL-5 NHL 82.07 OCl-Ly18 NHL 196.20 OCl-Ly1 NHL 416.95 SU-DHL-16 NHL 545.55 SU-DHL-2 NHL 1000.00 WSU-DLCL2 NHL 1000.00 Toledo NHL 1000.00 OCl-LY3 NHL 1000.00 RL NHL 1000.00 SU-DHL-4 NHL 1000.00 U2932 NHL 1000.00 HT NHL 1000.00 RC-K8 NHL 1000.00 SKM-1 AML 0.95 PL-21 AML 10.67 EOL-1 AML 18.31 HL-60 AML 76.62 OCl-AML2 AML 124.32 UKE-1 AML 205.35 MV4-11 AML 312.55 SET-2 AML 384.80 MOLM-13 AML 722.10 OCl-AML5 AML 1032.60 Kasumi-1 AML 1000.00 KG-1 AML 1000.00 OCl-AML3 AML 1000.00 SHI-1 AML 1000.00 SKNO-1 AML 1000.00 TF-1 AML 1000.00 THP-1 AML 1000.00 HEL AML 1000.00 Jurkat ALL 3.08 B NCl-H847 NSCLC 14.53 NCl-H647 NSCLC 24.75 NCl-H2444 NSCLC 27.75 NCl-H2170 NSCLC 30.16 NCl-H460 NSCLC 36.85 NCl-H838 NSCLC 44.48 NCl-H1792 NSCLC 61.09 NCl-H2347 NSCLC 81.06 NCl-H1373 NSCLC 125.15 NCl-H522 NSCLC 259.87 NCl-H2110 NSCLC 314.20 NCl-H596 NSCLC 397.80 HCC4006 NSCLC 407.24 NCl-H2122 NSCLC 480.55 NCl-H1299 NSCLC 716.00 NCl-H1975 NSCLC 741.50 HCC827 NSCLC 2824.50 NCl-H727 NSCLC 3178.00 NCl-H1944 NSCLC 4068.75 NCl-H1299 NSCLC 4214.87 Calu-6 NSCLC 4757.00 NCl-H1693 NSCLC 5000.00 HCC2935 NSCLC 5000.00 A549 NSCLC 5000.00 NCl-H1395 NSCLC 5000.00 NCl-H2172 NSCLC 5000.00 Calu-1 NSCLC 5000.00 NCl-H441 NSCLC 5000.00 NCl-H23 NSCLC 5000.00 NCl-H661 NSCLC 5000.00 NCl-H1650-GFP NSCLC >3 BxPC3 Pancreatic 16.00 Capan-1 Pancreatic 393.00 MIA PaCa-2 Pancreatic 158.00 PANC-1 Pancreatic >1000 SW48 CRC 6.10 Colo205 CRC 1.30 SW480 CRC 132.00 HCT 116-GFP CRC 337.00 HCC38 BrCa 3.00 HCC1569 BrCa 219.00 MCF7 BrCa >3000 MDA-MB-231 BRCa 235.00 HeyA8-GFP Ovarian 141.00 Fadu-GFP H&N >3 HT-1080 Fibrosarcoma 377.00 NCl-H211 SCLC 72.58
6.2 TRAIL Receptor Agonist Protein of SEQ ID NO: 19 Synergizes with Anti-Tumorigenic Agents to Induce Tumor Cell Death
[0169] Tumor cells were seeded at 10,000 cells/well in 96-well plates in the recommended media containing 10% FBS and co-treated with a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 and venetoclax (ABT-199), navitoclax (ABT-263) or docetaxel (DTX) for 24 hrs at 37.degree. C. in a humidified, 5% CO2 atmosphere. Cell viability was subsequently assessed using CellTiter-Glo.RTM. reagent as described by the manufacturer's instructions. The Bliss independence model (Wong et al., 2012; Mol. Cancer Ther. 11:1026-1035; Bernebaum, 1981 Adv. Cancer Res. 35:269-335; Borisy et al., 2003 Proc. Natl. Acad. Sci. USA 100:7977-7982) was employed to assess combination activity, with negative integers indicating antagonism, a value of zero indicating additive activity, and positive integers indicating synergy. Bliss scores were calculated for each combination in the dose matrix and totaled to give a "Bliss sum" value. An example of synergistic tumor cell death induced by co-treating human tumor cells with a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO:19 and venetoclax, navitoclax or DTX, with the associated Bliss sum is shown in FIGS. 20(A-C). Bliss sums determined for these combinations in a number of tumor cell lines are depicted in Table 9.
TABLE-US-00009 TABLE 9 Bliss synergy assessment of cell killing by TRAIL receptor agonist protein of SEQ ID NO: 19 in combination with DTX in NSCLC cell lines (A) and venetoclax or navitoclax in NHL & AML cell lines (B) in vitro. Bliss Sum Bliss Sum Bliss Sum (SEQ ID (SEQ ID (SEQ ID Tumor Cell NO: 19 + Tumor Cell NO: 19 + NO: 19 + line DTX) line venitoclax) navitoclax) LG0552 748.2 WSU-DLCL2 1292 560 NCl-H522 549.1 SU-DHL-4 898 617 NCl-H647 452 OCl-AML3 831.9 456.4 NCl-H727 429.4 OCl-AML5 777.8 174.5 NCl-H1373 387.1 U2932 736.2 636.1 NCl-H596 261 PL-21 600.8 244.9 HCC2935 224.2 ULA 343.8 79.4 NCl-H2347 154 OCl-Ly18 309.8 8.3 NCl-H2444 135 MV4; 11 301.1 351 A549 118.1 RL 286.1 446.7 NCl-H23 70.6 MOLM-13 270.6 264.8 NCl-H847 64.2 SKM-1 222.9 88.1 HCC4006 15.75 OCl-Ly1 218.8 69.1 NCl-H2170 -97.1 SU-DHL-16 217.5 142.5 LG0567 -105.2 OCl-AMI2 160.2 145.5 HCC2935 -183 OCl-Ly8 154.9 177 HCC827 -292.8 THP-1 152.7 43.1 NCl-H661 -344.5 OCl-Ly3 146.5 -242.2 NCl-H441 -362 OCl-Ly2 145 127.2 NCl-H1395 -512 OCl-Ly19 114.9 37.3 NCl-H1944 -565 SKNO-1 104.7 -138.9 NCl-H1693 -584 UKE-1 80.5 28.9 Calu-6 -628.7 WSU-NHK 79.8 84 LG0481 -803 EOL-1 69.7 -6.3 NCl-H2172 -1404 SU-DHL-2 53.5 -31.8 Toledo 51.4 -68.2 HEL 21.5 -92.4 NuDHL-1 -28.4 18.7 TF-1 -100.6 -50.2 RC-K8 -131 -68 HT -173 12.1 HL-60 -176 -112.7 SHI-1 -208.4 -122.6 SU-DHL-8 -210.6 -37 SU-DHL-5 -233.8 -280.6 SET-2 -248.4 71.2 KG-1 -260 -20.3 Kasumi-1 -356.4 -241.2
7. TRAIL Receptor Agonist Protein of SEQ ID NO:19 Treatment Inhibits Tumor Growth In Vivo
[0170] The effect of a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO:19 on tumor growth was evaluated in subcutaneous Colo205 (colorectal), SKM-1 (acute myeloid leukemia), and H460LM (non-small cell lung) xenograft tumors implanted in SCID female mice (Charles Rivers Laboratories; Wilmington, Mass.). Briefly, human cancer cells were inoculated subcutaneously into the right hind flank of female SCID mice on study day 0. Administration of TRAIL receptor agonist protein of SEQ ID NO:19 (0.3, 1, or 3 mkd dosed IV, QDx5 or IP, Q2Dx5 as indicated) was initiated at the time of size match. Tumor volume was measured for the duration of the experiment until the mean tumor volume in each group reached an endpoint of >2000 mm.sup.3 for Colo205 and SKM-1 or >2500 mm.sup.3 for H460LM. Results are shown in FIGS. 21-23. Administration of a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO:19 induced significant tumor growth inhibition in the Colo205, SKM-1, and H460LM xenograft tumor models.
[0171] The effect of a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO:19 on tumor growth was also evaluated in patient-derived xenograft models CTG-0069 (colorectal), CTG-0167 (NSCLC), CTG-0293 (pancreatic), CTG-0714 (sarcoma), CTG-0136 (esophageal), CTG-485 (gastric), and CTG-0785 (Ewing's sarcoma) implanted in NSG female mice (Champions Oncology; Hackensack, N.J.). Briefly, tumor fragments were propagated subcutaneously into the right hind flank of female NSG mice on study day 0. Administration of a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 (3 mkd dosed IP, Q2Dx5) was initiated at the time of size match. Tumor volume was measured for the duration of the experiment until the mean tumor volume in each group reached an endpoint of >2000 mm.sup.3 or 60 days. Results are shown in FIGS. 24(A-G). Administration of a TRAIL receptor agonist protein made up of two polypeptides having the amino acid sequence set forth in SEQ ID NO: 19 induced significant tumor growth inhibition in the CTG-0069 (colorectal), CTG-0167 (NSCLC), CTG-0293 (pancreatic), CTG-0714 (sarcoma), CTG-0136 (esophageal), CTG-485 (gastric), and CTG-0785 (Ewing's sarcoma) PDX models.
Sequence CWU 1
1
311281PRTHomo sapienssource/note="human TRAIL" 1Met Ala Met Met Glu
Val Gln Gly Gly Pro Ser Leu Gly Gln Thr Cys1 5 10 15Val Leu Ile Val
Ile Phe Thr Val Leu Leu Gln Ser Leu Cys Val Ala 20 25 30Val Thr Tyr
Val Tyr Phe Thr Asn Glu Leu Lys Gln Met Gln Asp Lys 35 40 45Tyr Ser
Lys Ser Gly Ile Ala Cys Phe Leu Lys Glu Asp Asp Ser Tyr 50 55 60Trp
Asp Pro Asn Asp Glu Glu Ser Met Asn Ser Pro Cys Trp Gln Val65 70 75
80Lys Trp Gln Leu Arg Gln Leu Val Arg Lys Met Ile Leu Arg Thr Ser
85 90 95Glu Glu Thr Ile Ser Thr Val Gln Glu Lys Gln Gln Asn Ile Ser
Pro 100 105 110Leu Val Arg Glu Arg Gly Pro Gln Arg Val Ala Ala His
Ile Thr Gly 115 120 125Thr Arg Gly Arg Ser Asn Thr Leu Ser Ser Pro
Asn Ser Lys Asn Glu 130 135 140Lys Ala Leu Gly Arg Lys Ile Asn Ser
Trp Glu Ser Ser Arg Ser Gly145 150 155 160His Ser Phe Leu Ser Asn
Leu His Leu Arg Asn Gly Glu Leu Val Ile 165 170 175His Glu Lys Gly
Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe 180 185 190Gln Glu
Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met Val Gln 195 200
205Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys
210 215 220Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu Tyr Gly
Leu Tyr225 230 235 240Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu Lys
Glu Asn Asp Arg Ile 245 250 255Phe Val Ser Val Thr Asn Glu His Leu
Ile Asp Met Asp His Glu Ala 260 265 270Ser Phe Phe Gly Ala Phe Leu
Val Gly 275 28028PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 2Gly Ser Gly Ser Gly
Asn Gly Ser1 538PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 3Gly Ser Gly Ser Gly
Ser Gly Ser1 548PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 4Gly Gly Ser Gly Ser
Gly Ser Gly1 556PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 5Gly Gly Ser Gly Ser
Gly1 564PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 6Gly Gly Ser
Gly178PRTArtificial Sequencesource/note="Description of Artificial
Sequence Synthetic linker peptide" 7Gly Gly Ser Gly Asn Gly Ser
Gly1 588PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 8Gly Gly Asn Gly Ser
Gly Ser Gly1 596PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic linker peptide" 9Gly Gly Asn Gly Ser
Gly1 510218PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic Fc fragment polypeptide" 10Pro Ala
Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro1 5 10 15Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys 20 25
30Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp
35 40 45Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg
Glu 50 55 60Glu Gln Tyr Ser Ser Thr Tyr Arg Val Val Ser Val Leu Thr
Val Leu65 70 75 80His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn 85 90 95Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly 100 105 110Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Glu Glu 115 120 125Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr 130 135 140Pro Ser Asp Ile Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn145 150 155 160Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe 165 170
175Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
180 185 190Val Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn His
Tyr Thr 195 200 205Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 210
2151122PRTArtificial Sequencesource/note="Description of Artificial
Sequence Synthetic hinge-linker peptide" 11Gly Pro Gly Ser Ser Ser
Ser Ser Ser Ser Gly Ser Cys Asp Lys Thr1 5 10 15His Thr Cys Pro Pro
Cys 201220PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic signal peptide" 12Met Glu Thr Asp Thr
Leu Leu Val Phe Val Leu Leu Val Trp Val Pro1 5 10 15Ala Gly Asn Gly
201315PRTArtificial Sequencesource/note="Description of Artificial
Sequence Synthetic strep-tag + serine linker peptide" 13Ser Ser Ser
Ser Ser Ser Ala Trp Ser His Pro Gln Phe Glu Lys1 5 10
1514760PRTArtificial Sequencesource/note="Description of Artificial
Sequence Synthetic single-chain fusion polypeptide" 14Met Glu Thr
Asp Thr Leu Leu Val Phe Val Leu Leu Val Trp Val Pro1 5 10 15Ala Gly
Asn Gly Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly 20 25 30Arg
Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu 35 40
45Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser Phe
50 55 60Leu Ser Asn Leu His Leu Arg Asn Gly Glu Leu Val Ile His Glu
Lys65 70 75 80Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe
Gln Glu Glu 85 90 95Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met Val
Gln Tyr Ile Tyr 100 105 110Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg 115 120 125Asn Ser Cys Trp Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr 130 135 140Gln Gly Gly Ile Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser145 150 155 160Val Thr Asn
Glu His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe 165 170 175Gly
Ala Phe Leu Val Gly Gly Ser Gly Ser Gly Asn Gly Ser Arg Val 180 185
190Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser Asn Thr Leu Ser Ser
195 200 205Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile Asn
Ser Trp 210 215 220Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn
Leu His Leu Arg225 230 235 240Asn Gly Glu Leu Val Ile His Glu Lys
Gly Phe Tyr Tyr Ile Tyr Ser 245 250 255Gln Thr Tyr Phe Arg Phe Gln
Glu Glu Ile Lys Glu Asn Thr Lys Asn 260 265 270Asp Lys Gln Met Val
Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp 275 280 285Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp 290 295 300Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu305 310
315 320Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu His Leu
Ile 325 330 335Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe Leu
Val Gly Gly 340 345 350Ser Gly Ser Gly Asn Gly Ser Arg Val Ala Ala
His Ile Thr Gly Thr 355 360 365Arg Gly Arg Ser Asn Thr Leu Ser Ser
Pro Asn Ser Lys Asn Glu Lys 370 375 380Ala Leu Gly Arg Lys Ile Asn
Ser Trp Glu Ser Ser Arg Ser Gly His385 390 395 400Ser Phe Leu Ser
Asn Leu His Leu Arg Asn Gly Glu Leu Val Ile His 405 410 415Glu Lys
Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe Gln 420 425
430Glu Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met Val Gln Tyr
435 440 445Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met
Lys Ser 450 455 460Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu Tyr
Gly Leu Tyr Ser465 470 475 480Ile Tyr Gln Gly Gly Ile Phe Glu Leu
Lys Glu Asn Asp Arg Ile Phe 485 490 495Val Ser Val Thr Asn Glu His
Leu Ile Asp Met Asp His Glu Ala Ser 500 505 510Phe Phe Gly Ala Phe
Leu Val Gly Gly Pro Gly Ser Ser Ser Ser Ser 515 520 525Ser Ser Gly
Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala 530 535 540Pro
Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro545 550
555 560Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val
Val 565 570 575Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn
Trp Tyr Val 580 585 590Asp Gly Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln 595 600 605Tyr Ser Ser Thr Tyr Arg Val Val Ser
Val Leu Thr Val Leu His Gln 610 615 620Asp Trp Leu Asn Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Ala625 630 635 640Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro 645 650 655Arg Glu
Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr 660 665
670Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
675 680 685Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr 690 695 700Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser
Phe Phe Leu Tyr705 710 715 720Ser Lys Leu Thr Val Asp Lys Ser Arg
Trp Gln Gln Gly Asn Val Phe 725 730 735Ser Cys Ser Val Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys 740 745 750Ser Leu Ser Leu Ser
Pro Gly Lys 755 76015774PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
single-chain fusion polypeptide" 15Met Glu Thr Asp Thr Leu Leu Val
Phe Val Leu Leu Val Trp Val Pro1 5 10 15Ala Gly Asn Gly Gln Arg Val
Ala Ala His Ile Thr Gly Thr Arg Gly 20 25 30Arg Ser Asn Thr Leu Ser
Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu 35 40 45Gly Arg Lys Ile Asn
Ser Trp Glu Ser Ser Arg Ser Gly His Ser Phe 50 55 60Leu Ser Asn Leu
His Leu Arg Asn Gly Glu Leu Val Ile His Glu Lys65 70 75 80Gly Phe
Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu 85 90 95Ile
Lys Glu Asn Thr Lys Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr 100 105
110Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg
115 120 125Asn Ser Cys Trp Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser
Ile Tyr 130 135 140Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp Arg
Ile Phe Val Ser145 150 155 160Val Thr Asn Glu His Leu Ile Asp Met
Asp His Glu Ala Ser Phe Phe 165 170 175Gly Ala Phe Leu Val Gly Gly
Ser Gly Ser Gly Asn Gly Ser Arg Val 180 185 190Ala Ala His Ile Thr
Gly Thr Arg Gly Arg Ser Asn Thr Leu Ser Ser 195 200 205Pro Asn Ser
Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile Asn Ser Trp 210 215 220Glu
Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn Leu His Leu Arg225 230
235 240Asn Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr Ile Tyr
Ser 245 250 255Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys Glu Asn
Thr Lys Asn 260 265 270Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr
Thr Ser Tyr Pro Asp 275 280 285Pro Ile Leu Leu Met Lys Ser Ala Arg
Asn Ser Cys Trp Ser Lys Asp 290 295 300Ala Glu Tyr Gly Leu Tyr Ser
Ile Tyr Gln Gly Gly Ile Phe Glu Leu305 310 315 320Lys Glu Asn Asp
Arg Ile Phe Val Ser Val Thr Asn Glu His Leu Ile 325 330 335Asp Met
Asp His Glu Ala Ser Phe Phe Gly Ala Phe Leu Val Gly Gly 340 345
350Ser Gly Ser Gly Asn Gly Ser Arg Val Ala Ala His Ile Thr Gly Thr
355 360 365Arg Gly Arg Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys Asn
Glu Lys 370 375 380Ala Leu Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser
Arg Ser Gly His385 390 395 400Ser Phe Leu Ser Asn Leu His Leu Arg
Asn Gly Glu Leu Val Ile His 405 410 415Glu Lys Gly Phe Tyr Tyr Ile
Tyr Ser Gln Thr Tyr Phe Arg Phe Gln 420 425 430Glu Glu Ile Lys Glu
Asn Thr Lys Asn Asp Lys Gln Met Val Gln Tyr 435 440 445Ile Tyr Lys
Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser 450 455 460Ala
Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser465 470
475 480Ile Tyr Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp Arg Ile
Phe 485 490 495Val Ser Val Thr Asn Glu His Leu Ile Asp Met Asp His
Glu Ala Ser 500 505 510Phe Phe Gly Ala Phe Leu Val Gly Gly Pro Gly
Ser Ser Ser Ser Ser 515 520 525Ser Ser Gly Ser Cys Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala 530 535 540Pro Glu Leu Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro545 550 555 560Lys Asp Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val 565 570 575Val Asp
Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val 580 585
590Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
595 600 605Tyr Ser Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln 610 615 620Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser Asn Lys Ala625 630 635 640Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro 645 650 655Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Glu Glu Met Thr 660 665 670Lys Asn Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser 675 680 685Asp Ile Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr 690 695 700Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr705 710
715 720Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe 725 730 735Ser Cys Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys 740 745 750Ser Leu Ser Leu Ser Pro Gly Ser Ser Ser Ser
Ser Ser Ala Trp Ser 755 760 765His Pro Gln Phe Glu Lys
770162316DNAArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic DNA polynucleotide sequence encoding
a single-chain fusion polypeptide" 16gatatcggta ccgccaccat
ggaaaccgac accctgctgg tgttcgtgct gctcgtgtgg 60gtgccagccg gcaatggaca
gagagtggcc gctcatatca ccggcacccg gggcagatct 120aacaccctgt
ccagccccaa ctccaagaac gagaaggccc tgggccggaa gatcaactcc
180tgggagtcct ccagatccgg ccactccttt ctgtccaacc tgcacctgag
aaacggcgag 240ctggtcatcc acgagaaggg cttctactac atctactccc
agacctactt caggtttcag 300gaagagatca aagagaacac aaagaacgac
aagcagatgg tgcagtatat ctacaagtac 360acctcctacc ccgaccccat
cctgctgatg aagtccgccc ggaactcctg ctggtccaag 420gatgctgagt
acggcctgta cagcatctac cagggcggca tcttcgagct gaaagagaac
480gaccggatct tcgtgtccgt gaccaacgag cacctgatcg acatggacca
cgaggccagc 540tttttcggcg cctttctcgt gggcggatcc ggaagcggaa
acggcagtag agtggctgcc 600cacattaccg gaaccagagg ccggtccaac
accctgagca gccctaacag caaaaatgag 660aaagctctcg ggcgcaagat
caacagctgg gaatctagca gaagcggcca cagctttctg 720agcaatctgc
atctgcggaa cggcgaactc gtgattcatg agaaggggtt ttattatatc
780tatagccaga catactttcg attccaggag gaaatcaagg aaaacaccaa
aaatgataaa 840cagatggtcc agtacattta taagtatacc agctaccctg
atcctatcct cctcatgaag 900tctgccagaa actcttgttg gagcaaggac
gccgagtatg gactgtactc tatctatcag 960ggggggatct ttgaactcaa
agaaaacgat cgcatctttg tcagcgtcac caatgagcat 1020ctcattgata
tggatcatga agctagtttc ttcggggcat tcctcgtggg aggctccggc
1080tctggcaacg gatctagagt cgccgcacac atcacaggga ccagaggcag
aagcaatacc 1140ctgtcctccc caaatagtaa aaacgaaaag gcactcggcc
gcaaaattaa ttcctgggag 1200agcagcagat ccgggcacag ttttctgtct
aatctccatc tgaggaatgg ggagctggtg 1260attcacgaaa aaggatttta
ctacatttac agtcagactt actttcgttt tcaggaagag 1320attaaggaaa
ataccaaaaa cgacaagcag atggtccagt acatctataa atacacctct
1380tatcctgacc caattctgct catgaagagt gcccgcaaca gctgctggtc
taaagacgcc 1440gaatacgggc tgtattccat ttaccagggg ggaatttttg
agctgaagga aaatgatcgg 1500atttttgtct ctgtcacaaa cgaacacctc
atcgatatgg atcacgaagc ctctttcttt 1560ggcgccttcc tggtcggagg
ccctggctcg agttccagct cctcttctgg ctcctgcgac 1620aagacccaca
cctgtccccc ttgtcctgcc cctgaactgc tgggcggacc ttccgtgttc
1680ctgttccccc caaagcccaa ggacaccctg atgatctccc ggacccccga
agtgacctgc 1740gtggtggtgg atgtgtctca cgaggaccct gaagtgaagt
tcaattggta cgtggacggc 1800gtggaagtgc acaacgccaa gaccaagccc
agagaggaac agtactcctc cacctaccgg 1860gtggtgtctg tgctgaccgt
gctgcaccag gactggctga acggcaaaga gtacaagtgc 1920aaggtgtcca
acaaggccct gcctgccccc atcgaaaaga ccatctccaa ggccaagggc
1980cagccccggg aaccccaggt gtacacactg ccccctagcc gggaagagat
gaccaagaac 2040caggtgtccc tgacctgcct ggtcaagggc ttttacccct
ccgacattgc cgtggaatgg 2100gagtccaacg gccagcctga gaacaactac
aagaccaccc cccctgtgct ggactccgac 2160ggctcattct tcctgtactc
caagctgaca gtggacaagt cccggtggca gcagggcaac 2220gtgttctcct
gctccgtgat gcacgaggcc ctgcacaacc actacaccca gaagtccctg
2280tccctgagcc ccggcaaatg atagaagctt gatatc 231617217PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
human Fc mutein polypeptide" 17Pro Ala Pro Pro Val Ala Gly Pro Ser
Val Phe Leu Phe Pro Pro Lys1 5 10 15Pro Lys Asp Thr Leu Met Ile Ser
Arg Thr Pro Glu Val Thr Cys Val 20 25 30Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp Tyr 35 40 45Val Asp Gly Val Glu Val
His Asn Ala Lys Thr Lys Pro Arg Glu Glu 50 55 60Gln Tyr Asn Ser Thr
Tyr Arg Val Val Ser Val Leu Thr Val Leu His65 70 75 80Gln Asp Trp
Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys 85 90 95Gly Leu
Pro Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln 100 105
110Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met
115 120 125Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro 130 135 140Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn145 150 155 160Tyr Lys Thr Thr Pro Pro Val Leu Asp
Ser Asp Gly Ser Phe Phe Leu 165 170 175Tyr Ser Lys Leu Thr Val Asp
Lys Ser Arg Trp Gln Gln Gly Asn Val 180 185 190Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn His Tyr Thr Gln 195 200 205Lys Ser Leu
Ser Leu Ser Pro Gly Lys 210 21518759PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
polypeptide" 18Met Glu Thr Asp Thr Leu Leu Val Phe Val Leu Leu Val
Trp Val Pro1 5 10 15Ala Gly Asn Gly Gln Arg Val Ala Ala His Ile Thr
Gly Thr Arg Gly 20 25 30Arg Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys
Asn Glu Lys Ala Leu 35 40 45Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser
Arg Ser Gly His Ser Phe 50 55 60Leu Ser Asn Leu His Leu Arg Asn Gly
Glu Leu Val Ile His Glu Lys65 70 75 80Gly Phe Tyr Tyr Ile Tyr Ser
Gln Thr Tyr Phe Arg Phe Gln Glu Glu 85 90 95Ile Lys Glu Asn Thr Lys
Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr 100 105 110Lys Tyr Thr Ser
Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg 115 120 125Asn Ser
Cys Trp Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr 130 135
140Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe Val
Ser145 150 155 160Val Thr Asn Glu His Leu Ile Asp Met Asp His Glu
Ala Ser Phe Phe 165 170 175Gly Ala Phe Leu Val Gly Gly Ser Gly Ser
Gly Asn Gly Ser Arg Val 180 185 190Ala Ala His Ile Thr Gly Thr Arg
Gly Arg Ser Asn Thr Leu Ser Ser 195 200 205Pro Asn Ser Lys Asn Glu
Lys Ala Leu Gly Arg Lys Ile Asn Ser Trp 210 215 220Glu Ser Ser Arg
Ser Gly His Ser Phe Leu Ser Asn Leu His Leu Arg225 230 235 240Asn
Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr Ile Tyr Ser 245 250
255Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys Asn
260 265 270Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr
Pro Asp 275 280 285Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser Cys
Trp Ser Lys Asp 290 295 300Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln
Gly Gly Ile Phe Glu Leu305 310 315 320Lys Glu Asn Asp Arg Ile Phe
Val Ser Val Thr Asn Glu His Leu Ile 325 330 335Asp Met Asp His Glu
Ala Ser Phe Phe Gly Ala Phe Leu Val Gly Gly 340 345 350Ser Gly Ser
Gly Asn Gly Ser Arg Val Ala Ala His Ile Thr Gly Thr 355 360 365Arg
Gly Arg Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys 370 375
380Ala Leu Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser Arg Ser Gly
His385 390 395 400Ser Phe Leu Ser Asn Leu His Leu Arg Asn Gly Glu
Leu Val Ile His 405 410 415Glu Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln
Thr Tyr Phe Arg Phe Gln 420 425 430Glu Glu Ile Lys Glu Asn Thr Lys
Asn Asp Lys Gln Met Val Gln Tyr 435 440 445Ile Tyr Lys Tyr Thr Ser
Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser 450 455 460Ala Arg Asn Ser
Cys Trp Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser465 470 475 480Ile
Tyr Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe 485 490
495Val Ser Val Thr Asn Glu His Leu Ile Asp Met Asp His Glu Ala Ser
500 505 510Phe Phe Gly Ala Phe Leu Val Gly Gly Pro Gly Ser Ser Ser
Ser Ser 515 520 525Ser Ser Gly Ser Cys Asp Lys Thr His Thr Cys Pro
Pro Cys Pro Ala 530 535 540Pro Pro Val Ala Gly Pro Ser Val Phe Leu
Phe Pro Pro Lys Pro Lys545 550 555 560Asp Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val 565 570 575Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp 580 585 590Gly Val Glu
Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr 595 600 605Asn
Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp 610 615
620Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly
Leu625 630 635 640Pro Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys
Gly Gln Pro Arg 645 650 655Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser
Arg Glu Glu Met Thr Lys 660 665 670Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr Pro Ser Asp 675 680 685Ile Ala Val Glu Trp Glu
Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys 690 695 700Thr Thr Pro Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser705 710 715 720Lys
Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser 725 730
735Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser
740 745 750Leu Ser Leu Ser Pro Gly Lys 75519740PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
polypeptide" 19Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg
Ser Asn Thr1 5 10 15Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu
Gly Arg Lys Ile 20 25 30Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser
Phe Leu Ser Asn Leu 35 40 45His Leu Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr 50 55 60Ile Tyr Ser Gln Thr Tyr Phe Arg Phe
Gln Glu Glu Ile Lys Glu Asn65 70 75 80Thr Lys Asn Asp Lys Gln Met
Val Gln Tyr Ile Tyr Lys Tyr Thr Ser 85 90 95Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser Cys Trp 100 105 110Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile 115 120 125Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu 130 135
140His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe
Leu145 150 155 160Val Gly Gly Ser Gly Ser Gly Asn Gly Ser Arg Val
Ala Ala His Ile 165 170 175Thr Gly Thr Arg Gly Arg Ser Asn Thr Leu
Ser Ser Pro Asn Ser Lys 180 185 190Asn Glu Lys Ala Leu Gly Arg Lys
Ile Asn Ser Trp Glu Ser Ser Arg 195 200 205Ser Gly His Ser Phe Leu
Ser Asn Leu His Leu Arg Asn Gly Glu Leu 210 215 220Val Ile His Glu
Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe225 230 235 240Arg
Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met 245 250
255Val Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu
260 265 270Met Lys Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu
Tyr Gly 275 280 285Leu Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu
Lys Glu Asn Asp 290 295 300Arg Ile Phe Val Ser Val Thr Asn Glu His
Leu Ile Asp Met Asp His305 310 315 320Glu Ala Ser Phe Phe Gly Ala
Phe Leu Val Gly Gly Ser Gly Ser Gly 325 330 335Asn Gly Ser Arg Val
Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser 340 345 350Asn Thr Leu
Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg 355 360 365Lys
Ile Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser 370 375
380Asn Leu His Leu Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly
Phe385 390 395 400Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe Gln
Glu Glu Ile Lys 405 410 415Glu Asn Thr Lys Asn Asp Lys Gln Met Val
Gln Tyr Ile Tyr Lys Tyr 420 425 430Thr Ser Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser 435 440 445Cys Trp Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly 450 455 460Gly Ile Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr465 470 475 480Asn
Glu His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala 485 490
495Phe Leu Val Gly Gly Pro Gly Ser Ser Ser Ser Ser Ser Ser Gly Ser
500 505 510Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu
Leu Leu 515 520 525Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro
Lys Asp Thr Leu 530 535 540Met Ile Ser Arg Thr Pro Glu Val Thr Cys
Val Val Val Asp Val Ser545 550 555 560His Glu Asp Pro Glu Val Lys
Phe Asn Trp Tyr Val Asp Gly Val Glu 565 570 575Val His Asn Ala Lys
Thr Lys Pro Arg Glu Glu Gln Tyr Ser Ser Thr 580 585 590Tyr Arg Val
Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn 595 600 605Gly
Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro 610 615
620Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro
Gln625 630 635 640Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr
Lys Asn Gln Val 645 650 655Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val 660 665 670Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro 675 680 685Pro Val Leu Asp Ser Asp
Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr 690 695 700Val Asp Lys Ser
Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val705 710 715 720Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu 725 730
735Ser Pro Gly Lys 74020754PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
polypeptide" 20Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg
Ser Asn Thr1 5 10 15Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu
Gly Arg Lys Ile 20 25 30Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser
Phe Leu Ser Asn Leu 35 40 45His Leu Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr 50 55 60Ile Tyr Ser Gln Thr Tyr Phe Arg Phe
Gln Glu Glu Ile Lys Glu Asn65 70 75 80Thr Lys Asn Asp Lys Gln Met
Val Gln Tyr Ile Tyr Lys Tyr Thr Ser 85 90 95Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser Cys Trp 100 105 110Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile 115 120 125Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu 130 135
140His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe
Leu145 150 155 160Val Gly Gly Ser Gly Ser Gly Asn Gly Ser Arg Val
Ala Ala His Ile 165 170 175Thr Gly Thr Arg Gly Arg Ser Asn Thr Leu
Ser Ser Pro Asn Ser Lys 180 185 190Asn Glu Lys Ala Leu Gly Arg Lys
Ile Asn Ser Trp Glu Ser Ser Arg 195 200 205Ser Gly His Ser Phe Leu
Ser Asn Leu His Leu Arg Asn Gly Glu Leu 210 215 220Val Ile His Glu
Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe225 230 235 240Arg
Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met 245 250
255Val Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu
260 265 270Met Lys Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu
Tyr Gly 275 280 285Leu Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu
Lys Glu Asn Asp 290 295 300Arg Ile Phe Val Ser Val Thr Asn Glu His
Leu Ile Asp Met Asp His305 310 315 320Glu Ala Ser Phe Phe Gly Ala
Phe Leu Val Gly Gly Ser Gly Ser Gly 325 330 335Asn Gly Ser Arg Val
Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser 340 345 350Asn Thr Leu
Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg 355 360 365Lys
Ile Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser 370 375
380Asn Leu His Leu Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly
Phe385 390 395 400Tyr Tyr Ile Tyr
Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys 405 410 415Glu Asn
Thr Lys Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr 420 425
430Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser
435 440 445Cys Trp Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr
Gln Gly 450 455 460Gly Ile Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe
Val Ser Val Thr465 470 475 480Asn Glu His Leu Ile Asp Met Asp His
Glu Ala Ser Phe Phe Gly Ala 485 490 495Phe Leu Val Gly Gly Pro Gly
Ser Ser Ser Ser Ser Ser Ser Gly Ser 500 505 510Cys Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu 515 520 525Gly Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu 530 535 540Met
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser545 550
555 560His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val
Glu 565 570 575Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr
Ser Ser Thr 580 585 590Tyr Arg Val Val Ser Val Leu Thr Val Leu His
Gln Asp Trp Leu Asn 595 600 605Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala Pro 610 615 620Ile Glu Lys Thr Ile Ser Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln625 630 635 640Val Tyr Thr Leu
Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln Val 645 650 655Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val 660 665
670Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
675 680 685Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu Thr 690 695 700Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe
Ser Cys Ser Val705 710 715 720Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu 725 730 735Ser Pro Gly Ser Ser Ser Ser
Ser Ser Ala Trp Ser His Pro Gln Phe 740 745 750Glu
Lys21739PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic polypeptide" 21Gln Arg Val Ala Ala
His Ile Thr Gly Thr Arg Gly Arg Ser Asn Thr1 5 10 15Leu Ser Ser Pro
Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile 20 25 30Asn Ser Trp
Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn Leu 35 40 45His Leu
Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr 50 55 60Ile
Tyr Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys Glu Asn65 70 75
80Thr Lys Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr Thr Ser
85 90 95Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser Cys
Trp 100 105 110Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln
Gly Gly Ile 115 120 125Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe Val
Ser Val Thr Asn Glu 130 135 140His Leu Ile Asp Met Asp His Glu Ala
Ser Phe Phe Gly Ala Phe Leu145 150 155 160Val Gly Gly Ser Gly Ser
Gly Asn Gly Ser Arg Val Ala Ala His Ile 165 170 175Thr Gly Thr Arg
Gly Arg Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys 180 185 190Asn Glu
Lys Ala Leu Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser Arg 195 200
205Ser Gly His Ser Phe Leu Ser Asn Leu His Leu Arg Asn Gly Glu Leu
210 215 220Val Ile His Glu Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr
Tyr Phe225 230 235 240Arg Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys
Asn Asp Lys Gln Met 245 250 255Val Gln Tyr Ile Tyr Lys Tyr Thr Ser
Tyr Pro Asp Pro Ile Leu Leu 260 265 270Met Lys Ser Ala Arg Asn Ser
Cys Trp Ser Lys Asp Ala Glu Tyr Gly 275 280 285Leu Tyr Ser Ile Tyr
Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp 290 295 300Arg Ile Phe
Val Ser Val Thr Asn Glu His Leu Ile Asp Met Asp His305 310 315
320Glu Ala Ser Phe Phe Gly Ala Phe Leu Val Gly Gly Ser Gly Ser Gly
325 330 335Asn Gly Ser Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly
Arg Ser 340 345 350Asn Thr Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys
Ala Leu Gly Arg 355 360 365Lys Ile Asn Ser Trp Glu Ser Ser Arg Ser
Gly His Ser Phe Leu Ser 370 375 380Asn Leu His Leu Arg Asn Gly Glu
Leu Val Ile His Glu Lys Gly Phe385 390 395 400Tyr Tyr Ile Tyr Ser
Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys 405 410 415Glu Asn Thr
Lys Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr 420 425 430Thr
Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser 435 440
445Cys Trp Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly
450 455 460Gly Ile Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe Val Ser
Val Thr465 470 475 480Asn Glu His Leu Ile Asp Met Asp His Glu Ala
Ser Phe Phe Gly Ala 485 490 495Phe Leu Val Gly Gly Pro Gly Ser Ser
Ser Ser Ser Ser Ser Gly Ser 500 505 510Cys Asp Lys Thr His Thr Cys
Pro Pro Cys Pro Ala Pro Pro Val Ala 515 520 525Gly Pro Ser Val Phe
Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met 530 535 540Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser His545 550 555
560Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val
565 570 575His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser
Thr Tyr 580 585 590Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp
Trp Leu Asn Gly 595 600 605Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Gly Leu Pro Ser Ser Ile 610 615 620Glu Lys Thr Ile Ser Lys Ala Lys
Gly Gln Pro Arg Glu Pro Gln Val625 630 635 640Tyr Thr Leu Pro Pro
Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser 645 650 655Leu Thr Cys
Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu 660 665 670Trp
Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro 675 680
685Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val
690 695 700Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser
Val Met705 710 715 720His Glu Ala Leu His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu Ser 725 730 735Pro Gly Lys226PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
peptide" 22Gly Ser Gly Ser Gly Ser1 5234PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
peptide" 23Gly Ser Gly Ser1243PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
peptide" 24Gly Ser Gly12521PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
peptide" 25Gly Pro Gly Ser Ser Ser Ser Ser Ser Gly Ser Cys Asp Lys
Thr His1 5 10 15Thr Cys Pro Pro Cys 2026751PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
polypeptide" 26Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg
Ser Asn Thr1 5 10 15Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu
Gly Arg Lys Ile 20 25 30Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser
Phe Leu Ser Asn Leu 35 40 45His Leu Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr 50 55 60Ile Tyr Ser Gln Thr Tyr Phe Arg Phe
Gln Glu Glu Ile Lys Glu Asn65 70 75 80Thr Lys Asn Asp Lys Gln Met
Val Gln Tyr Ile Tyr Lys Tyr Thr Ser 85 90 95Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser Cys Trp 100 105 110Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile 115 120 125Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu 130 135
140His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe
Leu145 150 155 160Val Gly Gly Ser Gly Ser Gly Asn Gly Ser Arg Val
Ala Ala His Ile 165 170 175Thr Gly Thr Arg Gly Arg Ser Asn Thr Leu
Ser Ser Pro Asn Ser Lys 180 185 190Asn Glu Lys Ala Leu Gly Arg Lys
Ile Asn Ser Trp Glu Ser Ser Arg 195 200 205Ser Gly His Ser Phe Leu
Ser Asn Leu His Leu Arg Asn Gly Glu Leu 210 215 220Val Ile His Glu
Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe225 230 235 240Arg
Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met 245 250
255Val Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu
260 265 270Met Lys Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu
Tyr Gly 275 280 285Leu Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu
Lys Glu Asn Asp 290 295 300Arg Ile Phe Val Ser Val Thr Asn Glu His
Leu Ile Asp Met Asp His305 310 315 320Glu Ala Ser Phe Phe Gly Ala
Phe Leu Val Gly Gly Ser Gly Ser Gly 325 330 335Asn Gly Ser Arg Val
Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser 340 345 350Asn Thr Leu
Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg 355 360 365Lys
Ile Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser 370 375
380Asn Leu His Leu Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly
Phe385 390 395 400Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe Gln
Glu Glu Ile Lys 405 410 415Glu Asn Thr Lys Asn Asp Lys Gln Met Val
Gln Tyr Ile Tyr Lys Tyr 420 425 430Thr Ser Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser 435 440 445Cys Trp Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly 450 455 460Gly Ile Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr465 470 475 480Asn
Glu His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala 485 490
495Phe Leu Val Gly Gly Pro Gly Ser Ser Ser Ser Ser Ser Gly Ser Asp
500 505 510Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Pro Val Ala
Gly Pro 515 520 525Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr
Leu Met Ile Ser 530 535 540Arg Thr Pro Glu Val Thr Cys Val Val Val
Asp Val Ser His Glu Asp545 550 555 560Pro Glu Val Lys Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn 565 570 575Ala Lys Thr Lys Pro
Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val 580 585 590Val Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu 595 600 605Tyr
Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys 610 615
620Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr625 630 635 640Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln
Val Ser Leu Thr 645 650 655Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu 660 665 670Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val Leu 675 680 685Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys 690 695 700Ser Arg Trp Gln
Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu705 710 715 720Ala
Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly 725 730
735Ser Ser Ser Ser Ser Ser Ala Trp Ser His Pro Gln Phe Glu Lys 740
745 75027750PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic polypeptide" 27Gln Arg Val Ala Ala
His Ile Thr Gly Thr Arg Gly Arg Ser Asn Thr1 5 10 15Leu Ser Ser Pro
Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile 20 25 30Asn Ser Trp
Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn Leu 35 40 45His Leu
Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr 50 55 60Ile
Tyr Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys Glu Asn65 70 75
80Thr Lys Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr Thr Ser
85 90 95Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser Cys
Trp 100 105 110Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln
Gly Gly Ile 115 120 125Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe Val
Ser Val Thr Asn Glu 130 135 140His Leu Ile Asp Met Asp His Glu Ala
Ser Phe Phe Gly Ala Phe Leu145 150 155 160Val Gly Gly Ser Gly Ser
Gly Asn Gly Ser Arg Val Ala Ala His Ile 165 170 175Thr Gly Thr Arg
Gly Arg Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys 180 185 190Asn Glu
Lys Ala Leu Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser Arg 195 200
205Ser Gly His Ser Phe Leu Ser Asn Leu His Leu Arg Asn Gly Glu Leu
210 215 220Val Ile His Glu Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr
Tyr Phe225 230 235 240Arg Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys
Asn Asp Lys Gln Met 245 250 255Val Gln Tyr Ile Tyr Lys Tyr Thr Ser
Tyr Pro Asp Pro Ile Leu Leu 260 265 270Met Lys Ser Ala Arg Asn Ser
Cys Trp Ser Lys Asp Ala Glu Tyr Gly 275 280 285Leu Tyr Ser Ile Tyr
Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp 290 295 300Arg Ile Phe
Val Ser Val Thr Asn Glu His Leu Ile Asp Met Asp His305 310 315
320Glu Ala Ser Phe Phe Gly Ala Phe Leu Val Gly Gly Ser Gly Ser Arg
325 330 335Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser Asn Thr
Leu Ser 340 345 350Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg
Lys Ile Asn Ser 355 360 365Trp Glu Ser Ser Arg Ser Gly His Ser Phe
Leu Ser Asn Leu His Leu 370 375 380Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr Ile Tyr385 390 395 400Ser Gln Thr Tyr Phe
Arg Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys 405 410 415Asn Asp Lys
Gln Met Val Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro 420 425 430Asp
Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser Cys Trp Ser Lys 435 440
445Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu
450 455 460Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu
His Leu465 470 475 480Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly
Ala Phe Leu Val Gly 485 490 495Gly Pro Gly Ser Ser Ser Ser
Ser Ser Ser Gly Ser Cys Asp Lys Thr 500 505 510His Thr Cys Pro Pro
Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser 515 520 525Val Phe Leu
Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 530 535 540Thr
Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro545 550
555 560Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn
Ala 565 570 575Lys Thr Lys Pro Arg Glu Glu Gln Tyr Ser Ser Thr Tyr
Arg Val Val 580 585 590Ser Val Leu Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr 595 600 605Lys Cys Lys Val Ser Asn Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr 610 615 620Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu625 630 635 640Pro Pro Ser Arg
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 645 650 655Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 660 665
670Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp
675 680 685Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp
Lys Ser 690 695 700Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val
Met His Glu Ala705 710 715 720Leu His Asn His Tyr Thr Gln Lys Ser
Leu Ser Leu Ser Pro Gly Ser 725 730 735Ser Ser Ser Ser Ser Ala Trp
Ser His Pro Gln Phe Glu Lys 740 745 75028750PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
polypeptide" 28Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg
Ser Asn Thr1 5 10 15Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu
Gly Arg Lys Ile 20 25 30Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser
Phe Leu Ser Asn Leu 35 40 45His Leu Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr 50 55 60Ile Tyr Ser Gln Thr Tyr Phe Arg Phe
Gln Glu Glu Ile Lys Glu Asn65 70 75 80Thr Lys Asn Asp Lys Gln Met
Val Gln Tyr Ile Tyr Lys Tyr Thr Ser 85 90 95Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser Cys Trp 100 105 110Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile 115 120 125Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu 130 135
140His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe
Leu145 150 155 160Val Gly Gly Ser Gly Ser Gly Ser Arg Val Ala Ala
His Ile Thr Gly 165 170 175Thr Arg Gly Arg Ser Asn Thr Leu Ser Ser
Pro Asn Ser Lys Asn Glu 180 185 190Lys Ala Leu Gly Arg Lys Ile Asn
Ser Trp Glu Ser Ser Arg Ser Gly 195 200 205His Ser Phe Leu Ser Asn
Leu His Leu Arg Asn Gly Glu Leu Val Ile 210 215 220His Glu Lys Gly
Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe225 230 235 240Gln
Glu Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met Val Gln 245 250
255Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys
260 265 270Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu Tyr Gly
Leu Tyr 275 280 285Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu Lys Glu
Asn Asp Arg Ile 290 295 300Phe Val Ser Val Thr Asn Glu His Leu Ile
Asp Met Asp His Glu Ala305 310 315 320Ser Phe Phe Gly Ala Phe Leu
Val Gly Gly Ser Gly Ser Gly Ser Arg 325 330 335Val Ala Ala His Ile
Thr Gly Thr Arg Gly Arg Ser Asn Thr Leu Ser 340 345 350Ser Pro Asn
Ser Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile Asn Ser 355 360 365Trp
Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn Leu His Leu 370 375
380Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr Ile
Tyr385 390 395 400Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys
Glu Asn Thr Lys 405 410 415Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr
Lys Tyr Thr Ser Tyr Pro 420 425 430Asp Pro Ile Leu Leu Met Lys Ser
Ala Arg Asn Ser Cys Trp Ser Lys 435 440 445Asp Ala Glu Tyr Gly Leu
Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu 450 455 460Leu Lys Glu Asn
Asp Arg Ile Phe Val Ser Val Thr Asn Glu His Leu465 470 475 480Ile
Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe Leu Val Gly 485 490
495Gly Pro Gly Ser Ser Ser Ser Ser Ser Ser Gly Ser Cys Asp Lys Thr
500 505 510His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly
Pro Ser 515 520 525Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu
Met Ile Ser Arg 530 535 540Thr Pro Glu Val Thr Cys Val Val Val Asp
Val Ser His Glu Asp Pro545 550 555 560Glu Val Lys Phe Asn Trp Tyr
Val Asp Gly Val Glu Val His Asn Ala 565 570 575Lys Thr Lys Pro Arg
Glu Glu Gln Tyr Ser Ser Thr Tyr Arg Val Val 580 585 590Ser Val Leu
Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 595 600 605Lys
Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr 610 615
620Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu625 630 635 640Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys 645 650 655Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser 660 665 670Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp 675 680 685Ser Asp Gly Ser Phe Phe
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser 690 695 700Arg Trp Gln Gln
Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala705 710 715 720Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Ser 725 730
735Ser Ser Ser Ser Ser Ala Trp Ser His Pro Gln Phe Glu Lys 740 745
75029736PRTArtificial Sequencesource/note="Description of
Artificial Sequence Synthetic polypeptide" 29Gln Arg Val Ala Ala
His Ile Thr Gly Thr Arg Gly Arg Ser Asn Thr1 5 10 15Leu Ser Ser Pro
Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile 20 25 30Asn Ser Trp
Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn Leu 35 40 45His Leu
Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr 50 55 60Ile
Tyr Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys Glu Asn65 70 75
80Thr Lys Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr Lys Tyr Thr Ser
85 90 95Tyr Pro Asp Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser Cys
Trp 100 105 110Ser Lys Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln
Gly Gly Ile 115 120 125Phe Glu Leu Lys Glu Asn Asp Arg Ile Phe Val
Ser Val Thr Asn Glu 130 135 140His Leu Ile Asp Met Asp His Glu Ala
Ser Phe Phe Gly Ala Phe Leu145 150 155 160Val Gly Gly Ser Gly Ser
Gly Asn Gly Ser Arg Val Ala Ala His Ile 165 170 175Thr Gly Thr Arg
Gly Arg Ser Asn Thr Leu Ser Ser Pro Asn Ser Lys 180 185 190Asn Glu
Lys Ala Leu Gly Arg Lys Ile Asn Ser Trp Glu Ser Ser Arg 195 200
205Ser Gly His Ser Phe Leu Ser Asn Leu His Leu Arg Asn Gly Glu Leu
210 215 220Val Ile His Glu Lys Gly Phe Tyr Tyr Ile Tyr Ser Gln Thr
Tyr Phe225 230 235 240Arg Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys
Asn Asp Lys Gln Met 245 250 255Val Gln Tyr Ile Tyr Lys Tyr Thr Ser
Tyr Pro Asp Pro Ile Leu Leu 260 265 270Met Lys Ser Ala Arg Asn Ser
Cys Trp Ser Lys Asp Ala Glu Tyr Gly 275 280 285Leu Tyr Ser Ile Tyr
Gln Gly Gly Ile Phe Glu Leu Lys Glu Asn Asp 290 295 300Arg Ile Phe
Val Ser Val Thr Asn Glu His Leu Ile Asp Met Asp His305 310 315
320Glu Ala Ser Phe Phe Gly Ala Phe Leu Val Gly Gly Ser Gly Ser Arg
325 330 335Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg Ser Asn Thr
Leu Ser 340 345 350Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu Gly Arg
Lys Ile Asn Ser 355 360 365Trp Glu Ser Ser Arg Ser Gly His Ser Phe
Leu Ser Asn Leu His Leu 370 375 380Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr Ile Tyr385 390 395 400Ser Gln Thr Tyr Phe
Arg Phe Gln Glu Glu Ile Lys Glu Asn Thr Lys 405 410 415Asn Asp Lys
Gln Met Val Gln Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro 420 425 430Asp
Pro Ile Leu Leu Met Lys Ser Ala Arg Asn Ser Cys Trp Ser Lys 435 440
445Asp Ala Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu
450 455 460Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu
His Leu465 470 475 480Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly
Ala Phe Leu Val Gly 485 490 495Gly Pro Gly Ser Ser Ser Ser Ser Ser
Ser Gly Ser Cys Asp Lys Thr 500 505 510His Thr Cys Pro Pro Cys Pro
Ala Pro Glu Leu Leu Gly Gly Pro Ser 515 520 525Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 530 535 540Thr Pro Glu
Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro545 550 555
560Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
565 570 575Lys Thr Lys Pro Arg Glu Glu Gln Tyr Ser Ser Thr Tyr Arg
Val Val 580 585 590Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn
Gly Lys Glu Tyr 595 600 605Lys Cys Lys Val Ser Asn Lys Ala Leu Pro
Ala Pro Ile Glu Lys Thr 610 615 620Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu625 630 635 640Pro Pro Ser Arg Glu
Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 645 650 655Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 660 665 670Asn
Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 675 680
685Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
690 695 700Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His
Glu Ala705 710 715 720Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 725 730 73530736PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
polypeptide" 30Gln Arg Val Ala Ala His Ile Thr Gly Thr Arg Gly Arg
Ser Asn Thr1 5 10 15Leu Ser Ser Pro Asn Ser Lys Asn Glu Lys Ala Leu
Gly Arg Lys Ile 20 25 30Asn Ser Trp Glu Ser Ser Arg Ser Gly His Ser
Phe Leu Ser Asn Leu 35 40 45His Leu Arg Asn Gly Glu Leu Val Ile His
Glu Lys Gly Phe Tyr Tyr 50 55 60Ile Tyr Ser Gln Thr Tyr Phe Arg Phe
Gln Glu Glu Ile Lys Glu Asn65 70 75 80Thr Lys Asn Asp Lys Gln Met
Val Gln Tyr Ile Tyr Lys Tyr Thr Ser 85 90 95Tyr Pro Asp Pro Ile Leu
Leu Met Lys Ser Ala Arg Asn Ser Cys Trp 100 105 110Ser Lys Asp Ala
Glu Tyr Gly Leu Tyr Ser Ile Tyr Gln Gly Gly Ile 115 120 125Phe Glu
Leu Lys Glu Asn Asp Arg Ile Phe Val Ser Val Thr Asn Glu 130 135
140His Leu Ile Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe
Leu145 150 155 160Val Gly Gly Ser Gly Ser Gly Ser Arg Val Ala Ala
His Ile Thr Gly 165 170 175Thr Arg Gly Arg Ser Asn Thr Leu Ser Ser
Pro Asn Ser Lys Asn Glu 180 185 190Lys Ala Leu Gly Arg Lys Ile Asn
Ser Trp Glu Ser Ser Arg Ser Gly 195 200 205His Ser Phe Leu Ser Asn
Leu His Leu Arg Asn Gly Glu Leu Val Ile 210 215 220His Glu Lys Gly
Phe Tyr Tyr Ile Tyr Ser Gln Thr Tyr Phe Arg Phe225 230 235 240Gln
Glu Glu Ile Lys Glu Asn Thr Lys Asn Asp Lys Gln Met Val Gln 245 250
255Tyr Ile Tyr Lys Tyr Thr Ser Tyr Pro Asp Pro Ile Leu Leu Met Lys
260 265 270Ser Ala Arg Asn Ser Cys Trp Ser Lys Asp Ala Glu Tyr Gly
Leu Tyr 275 280 285Ser Ile Tyr Gln Gly Gly Ile Phe Glu Leu Lys Glu
Asn Asp Arg Ile 290 295 300Phe Val Ser Val Thr Asn Glu His Leu Ile
Asp Met Asp His Glu Ala305 310 315 320Ser Phe Phe Gly Ala Phe Leu
Val Gly Gly Ser Gly Ser Gly Ser Arg 325 330 335Val Ala Ala His Ile
Thr Gly Thr Arg Gly Arg Ser Asn Thr Leu Ser 340 345 350Ser Pro Asn
Ser Lys Asn Glu Lys Ala Leu Gly Arg Lys Ile Asn Ser 355 360 365Trp
Glu Ser Ser Arg Ser Gly His Ser Phe Leu Ser Asn Leu His Leu 370 375
380Arg Asn Gly Glu Leu Val Ile His Glu Lys Gly Phe Tyr Tyr Ile
Tyr385 390 395 400Ser Gln Thr Tyr Phe Arg Phe Gln Glu Glu Ile Lys
Glu Asn Thr Lys 405 410 415Asn Asp Lys Gln Met Val Gln Tyr Ile Tyr
Lys Tyr Thr Ser Tyr Pro 420 425 430Asp Pro Ile Leu Leu Met Lys Ser
Ala Arg Asn Ser Cys Trp Ser Lys 435 440 445Asp Ala Glu Tyr Gly Leu
Tyr Ser Ile Tyr Gln Gly Gly Ile Phe Glu 450 455 460Leu Lys Glu Asn
Asp Arg Ile Phe Val Ser Val Thr Asn Glu His Leu465 470 475 480Ile
Asp Met Asp His Glu Ala Ser Phe Phe Gly Ala Phe Leu Val Gly 485 490
495Gly Pro Gly Ser Ser Ser Ser Ser Ser Ser Gly Ser Cys Asp Lys Thr
500 505 510His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly
Pro Ser 515 520 525Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu
Met Ile Ser Arg 530 535 540Thr Pro Glu Val Thr Cys Val Val Val Asp
Val Ser His Glu Asp Pro545 550 555 560Glu Val Lys Phe Asn Trp Tyr
Val Asp Gly Val Glu Val His Asn Ala 565 570 575Lys Thr Lys Pro Arg
Glu Glu Gln Tyr Ser Ser Thr Tyr Arg Val Val 580 585 590Ser Val Leu
Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 595 600 605Lys
Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr 610 615
620Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu625 630 635 640Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys 645 650 655Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser 660 665 670Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp 675 680
685Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
690 695 700Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His
Glu Ala705 710 715 720Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 725 730 735314PRTArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic
peptide" 31Asp Glu Val Asp1
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015
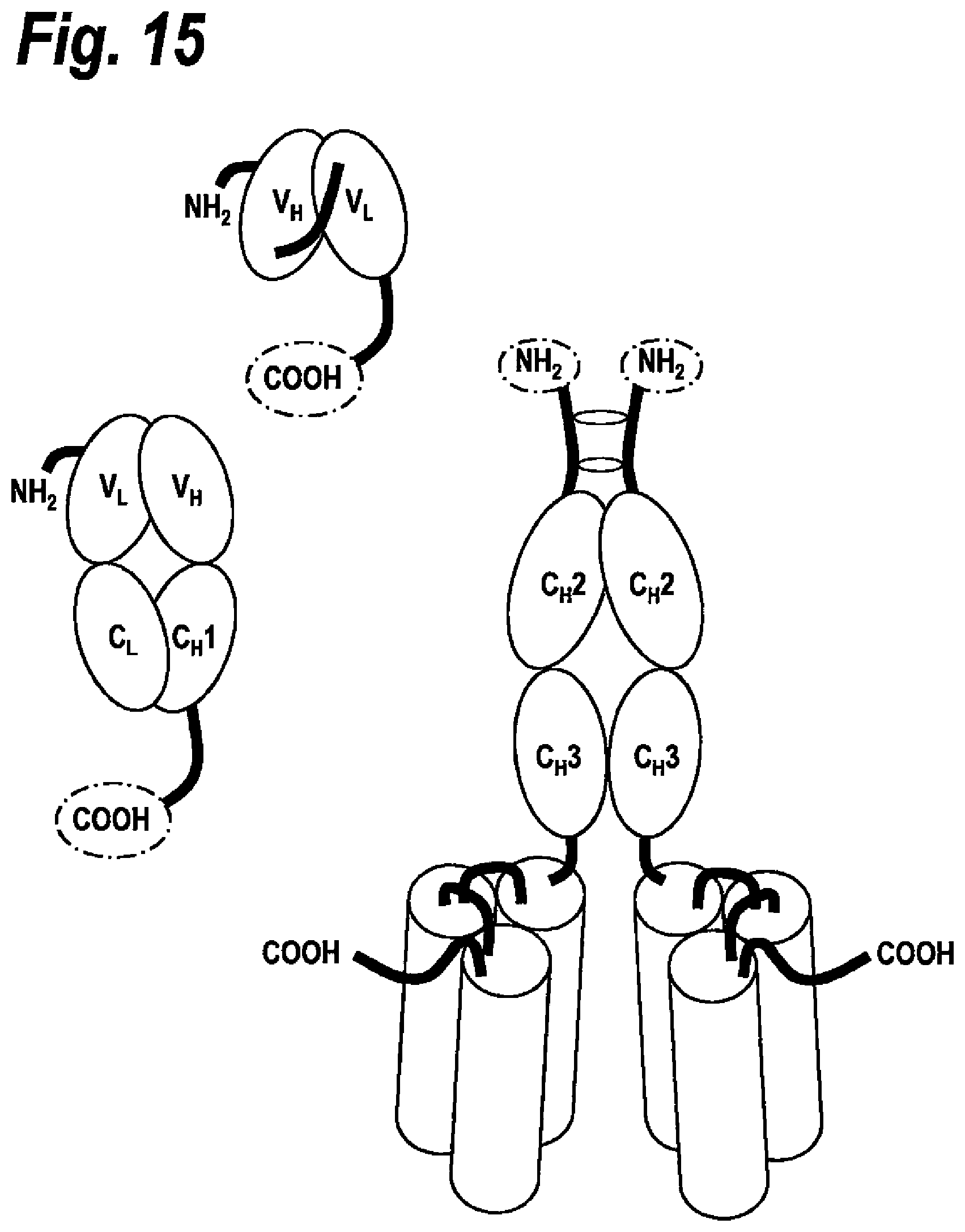
D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025
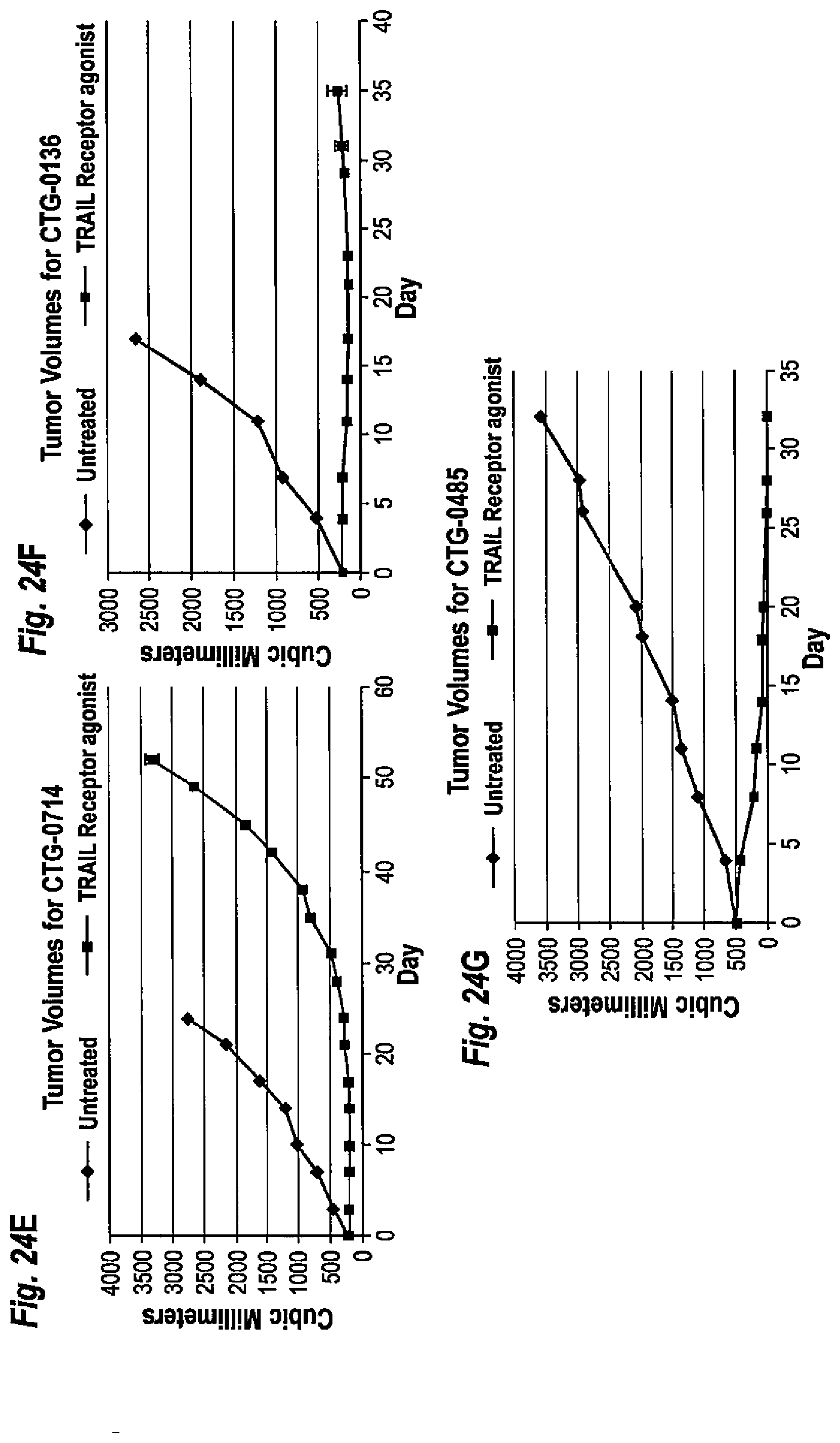
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.