System And Method For Propagating Information Using Modified Nucleic Acids
Kleinbaum; Daniel J.
U.S. patent application number 16/875641 was filed with the patent office on 2021-02-04 for system and method for propagating information using modified nucleic acids. The applicant listed for this patent is Emerald Therapeutics, Inc.. Invention is credited to Daniel J. Kleinbaum.
| Application Number | 20210034944 16/875641 |
| Document ID | / |
| Family ID | 1000005152333 |
| Filed Date | 2021-02-04 |



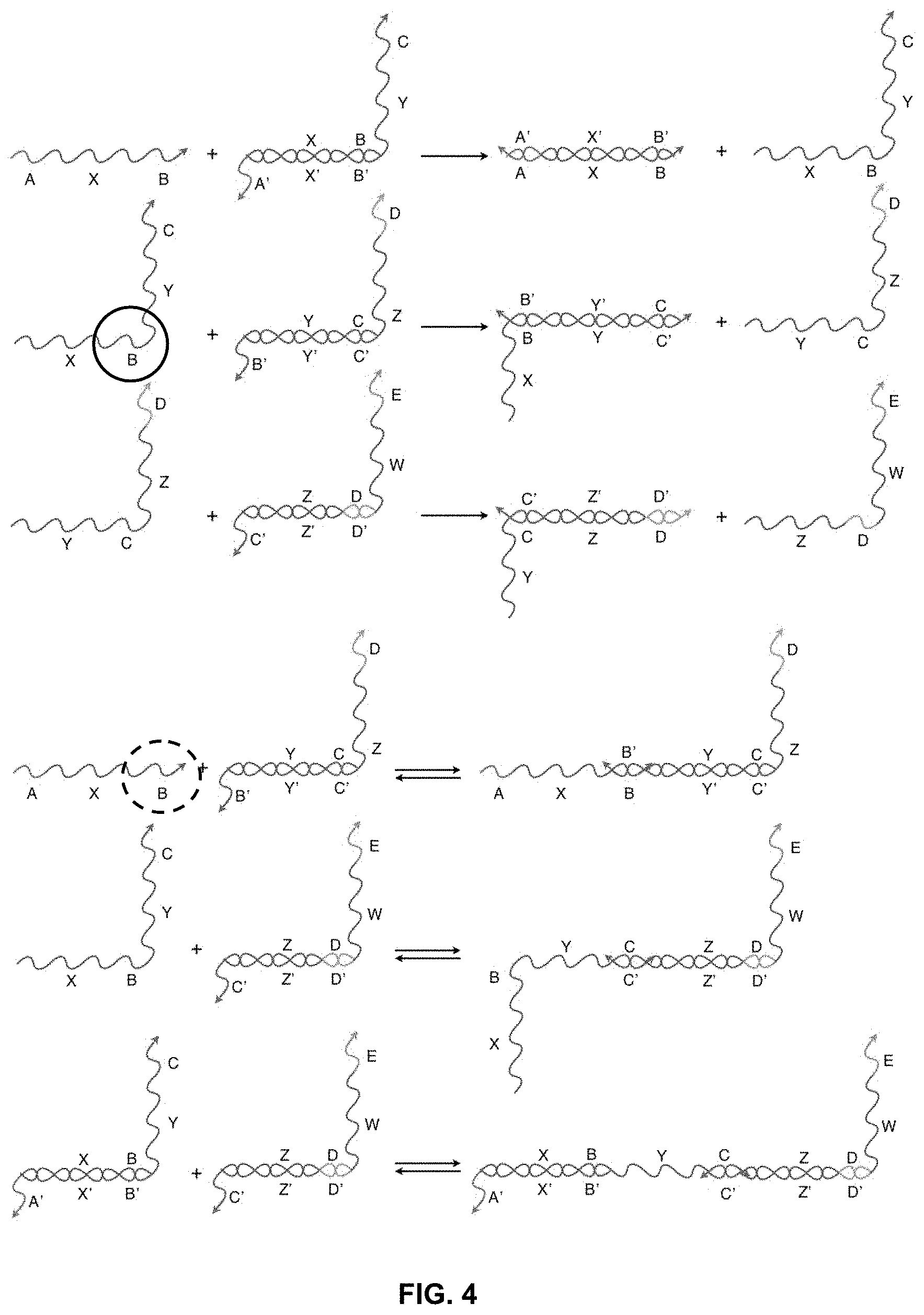

| United States Patent Application | 20210034944 |
| Kind Code | A1 |
| Kleinbaum; Daniel J. | February 4, 2021 |
SYSTEM AND METHOD FOR PROPAGATING INFORMATION USING MODIFIED NUCLEIC ACIDS
Abstract
A method is provided for improving a nucleic acid-based molecular computing system comprised of (i) a nucleic acid structure, (ii) at least one polynucleotide displacement molecule that can bind with the nucleic acid structure under hybridizing conditions, and (iii) a clashing polynucleotide molecule that competes with the polynucleotide displacement molecule for binding the nucleic acid structure under the hybridizing conditions The method entails incorporation of chemical modification that inhibits the binding of the clashing molecule and the nucleic acid structure or facilitates the binding of the displacement molecule and the nucleic structure.
| Inventors: | Kleinbaum; Daniel J.; (Redwood City, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005152333 | ||||||||||
| Appl. No.: | 16/875641 | ||||||||||
| Filed: | May 15, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15375629 | Dec 12, 2016 | |||
| 16875641 | ||||
| 14142531 | Dec 27, 2013 | 9547750 | ||
| 15375629 | ||||
| 13072438 | Mar 25, 2011 | 8630809 | ||
| 14142531 | ||||
| 61349012 | May 27, 2010 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12Q 1/6837 20130101; G06N 3/002 20130101; B82Y 10/00 20130101; G06N 3/123 20130101; G16B 30/00 20190201 |
| International Class: | G06N 3/00 20060101 G06N003/00; G16B 30/00 20060101 G16B030/00; B82Y 10/00 20060101 B82Y010/00; G06N 3/12 20060101 G06N003/12; C12Q 1/6837 20060101 C12Q001/6837 |
Claims
1. A method for improving a nucleic acid-based molecular computing system, comprising (A) identifying a computing system comprised of (i) a nucleic acid structure that comprises an incompletely base-paired duplex domain, (ii) at least one polynucleotide displacement molecule that can bind with said nucleic acid structure under hybridizing conditions, such that said nucleic acid structure undergoes a transition in energy state due to a branch migration reaction involving said duplex domain, and (iii) a clashing polynucleotide molecule that competes with said polynucleotide displacement molecule for binding said nucleic acid structure under said conditions but that cannot produce a branch migration reaction involving said duplex domain; then (B) reconfiguring at least one of said displacement molecule and said nucleic acid structure, respectively, to incorporate a chemical modification relative to a first reference molecule that comprises natural nucleotides and has the same sequence content as said displacement molecule or said nucleic acid structure, as the case may be, wherein said modification causes binding of said displacement molecule and said nucleic acid structure to have a hybridization free energy, differing from that of a first reference binding between said displacement molecule or said nucleic acid structure and said first reference molecule, such that said branch migration reaction is facilitated relative to said first reference binding; and/or (C) reconfiguring at least one of said clashing molecule and said nucleic acid structure, respectively, to incorporate a chemical modification relative to a second reference molecule that comprises natural nucleotides and has the same sequence content as said clashing molecule or said nucleic acid structure, as the case may be, wherein said modification causes binding of said clashing molecule and said nucleic acid structure to have a hybridization free energy, differing from that of a second reference binding between said clashing molecule or said nucleic acid structure and said second reference molecule, such that binding of said clashing molecule is impeded relative to said second reference binding.
2. The method of claim 1, wherein said branch migration reaction is facilitated by: increasing a probability of productive reactions in which binding of said nucleic acid structure by said displacement molecule has, by virtue of said chemical modification, a lower hybridization free energy than does a first reference binding reaction; or
3. The method of claim 1, wherein said branch migration reaction is facilitated by: reducing a probability of unproductive reactions in which binding of said nucleic acid structure by said clashing molecule has, by virtue of said chemical modification, a higher hybridization free energy than does a second reference binding reaction.
4. The method of claim 1, wherein said nucleic acid structure is part of an enzyme-free, nucleic acid logic gate selected from the group consisting of an AND gate, a NOT gate, an OR gate, a NAND gate, a NOR gate, an XOR gate, and an XNOR gate.
5. The method of claim 1, wherein said chemical modification is selected from the group consisting of (i) replacing the sugar-phosphodiester backbone of said nucleotides with a pseudo-peptide backbone, (ii) modifying the sugar moiety of said nucleotides, and (iii) substituting an analogue for the nitrogenous base of at least one of said nucleotides.
6. The method of claim 5, wherein said chemical modification comprises replacing the sugar-phosphodiester backbone of said nucleotides with a pseudo-peptide backbone into which a guanidinium functional group is incorporated.
7. The method of claim 5, wherein said chemical modification comprises substituting a tricyclic cytosine analogue for the nitrogenous base of at least one of said nucleotides.
8. The method of claim 5, wherein said chemical modification comprises introducing a heteroatom at the 2'-position of said sugar moiety.
9. The method of claim 1, wherein said branch migration reaction comprises a toe-hold-mediated strand displacement.
10. A nucleic acid-based system for propagating information, the system comprising: (A) a nucleic acid structure that comprises an incompletely base-paired duplex domain; (B) at least one polynucleotide displacement molecule to effect binding with said nucleic acid structure under hybridizing conditions, such that said nucleic acid structure undergoes a transition in energy state due to a branch migration reaction involving said duplex domain; and (C) at least one polynucleotide clashing molecule capable of binding with said nucleic acid structure under hybridizing conditions, such that said nucleic acid structure and said clashing molecule are bound and prevent the binding of the polynucleotide displacement molecule, wherein at least one of said displacement molecule and said nucleic acid structure, respectively, has a chemical modification relative to a reference molecule that comprises natural nucleotides and has the same sequence content as said displacement molecule or said nucleic acid structure, as the case may be, wherein said modification causes said binding to have a hybridization free energy, differing from that of a reference binding between said displacement molecule or said nucleic acid structure and a reference molecule, such that said branch migration reaction is facilitated relative to said reference binding, and/or wherein at least one of said clashing molecule and said nucleic acid structure, respectively, has a chemical modification relative to a reference molecule that comprises natural nucleotides and has the same sequence content as said displacement molecule or said nucleic acid structure, as the case may be, wherein said modification causes said binding to have a hybridization free energy, differing from that of a reference binding between said displacement molecule or said nucleic acid structure and a reference molecule, such that binding of said clashing molecule is impeded relative to said reference binding.
11. The system of claim 10, wherein said nucleic acid structure is part of an enzyme-free, nucleic acid logic gate selected from the group consisting of an AND gate, a NOT gate, an OR gate, a NAND gate, a NOR gate, an XOR gate, and an XNOR gate.
12. The system of claim 11, wherein said system comprises one or more circuits comprised of a plurality of enzyme-free, nucleic acid logic gates.
13. The system of claim 10, wherein said chemical modification is selected from the group consisting of (i) replacing the sugar-phosphodiester backbone of said nucleotides with a pseudo-peptide backbone, (ii) modifying the sugar moiety of said nucleotides, and (iii) substituting an analogue for the nitrogenous base of at least one of said nucleotides.
14. The system of claim 13, wherein said chemical modification comprises replacing the sugar-phosphodiester backbone of said nucleotides with a pseudo-peptide backbone into which a guanidinium functional group is incorporated.
15. The system of claim 14, wherein said chemical modification comprises substituting a tricyclic cytosine analogue for the nitrogenous base of at least one of said nucleotides.
16. The system of claim 14, wherein said chemical modification comprises introducing a heteroatom at the 2'-position of said sugar moiety.
17. The system of claim 10, wherein said branch migration reaction comprises a toe-hold-mediated strand displacement.
18. A system for propagating information, the system comprising a first molecule, anda second molecule configured to bind with the first molecule, wherein (A) at least one of the first molecule or the second molecule, respectively, has a chemical modification relative to a reference molecule that has the same sequence content as the first molecule or the second molecule, as the case may be, and (B) the modification causes said binding to have a free energy different from that of a reference binding between the reference molecule and the first or second molecule, such that at least one of the following is realized in the system: (i) the probability of a productive binding is raised; and (ii) the probability of an unproductive binding is reduced.
19. The system of claim 18, wherein the productive binding comprises a binding that propagates information, and wherein the unproductive binding comprises a binding that does not propagate information.
20. The system of claim 18, wherein the free energy difference as a result of the modification decreases for the productive binding, and increases for the unproductive binding.
Description
CROSS-REFERENCE TO RELATED PATENT APPLICATIONS
[0001] This application is a continuation of U.S. patent application Ser. No. 15/375,629, filed Dec. 12, 2016, which is a continuation of U.S. patent application Ser. No. 14/142,531, filed Dec. 27, 2013, now Pat. No. 9,547,750, issued Jan. 17, 2017, which is a divisional of U.S. patent application Ser. No. 13/072,438, filed Mar. 25, 2011, now Pat. No. 8,630,809, issued Jan. 14, 2014, which claims priority to U.S. Provisional Application No. 61/349,012, filed May 27, 2010, the entire contents of which are incorporated herein by reference.
BACKGROUND OF THE INVENTION
[0002] Nano-scale computers can potentially be realized using molecules. Such computers may be suited for solving certain computation problems. In particular, computers employing biomolecules can be compatible with biological environments, and can potentially be used for complex disease diagnostics or even treatments.
[0003] The ability to translate one nucleic acid sequence into another can be employed to build logic gates and networks with nucleic acids. These gates and networks are driven by two events: hybridization and strand displacement. Both events are generally thermodynamically favorable; that is, they involve a transition from a higher to a lower-energy state. Thus, both events can occur spontaneously in a system.
[0004] Hybridization involves free, single-stranded stretches of nucleic acids. Accordingly, a nucleic-acid network may be regulated by the availability of these free strands.
[0005] A "sequestering event" allows certain sequences to be available conditionally to the rest of the network. Such events empower the construction of translators, which convert one single-stranded nucleic acid sequence into a different single-stranded nucleic acid sequence.
[0006] These translators are the foundation on which basic logic operators, such as AND, NOT, OR, NAND, NOR, XOR and XNOR, can be built with nucleic acids. From these and other logic components, larger networks can be constructed that include components such as amplifiers. As a result, these translation events are important for information processing with nucleic acids and molecular computing.
SUMMARY OF THE INVENTION
[0007] In accordance with one aspect of the present invention, a method is provided for improving a nucleic acid-based molecular computing system. The method includes: (A) identifying a computing system comprised of (i) a nucleic acid structure that includes an incompletely base-paired duplex domain, (ii) at least one polynucleotide displacement molecule that can bind with the nucleic acid structure under hybridizing conditions, such that the nucleic acid structure undergoes a transition in energy state due to a branch migration reaction involving the duplex domain, and (iii) a clashing polynucleotide molecule that competes with the polynucleotide displacement molecule for binding the nucleic acid structure under the hybridizing conditions but that cannot produce a branch migration reaction involving the duplex domain; then (B) reconfiguring at least one of the displacement molecule and the nucleic acid structure, respectively, to incorporate a chemical modification relative to a first reference molecule that comprises natural nucleosides and has the same sequence content as the displacement molecule or the nucleic acid structure, as the case may be. The aforementioned the modification causes binding of the displacement molecule and the nucleic acid structure to have a hybridization free energy, differing from that of a first reference binding between the displacement molecule or the nucleic acid structure and the first reference molecule, such that the branch migration reaction is facilitated relative to the first reference binding. After or in place of step (B) is a step (C) of reconfiguring at least one of the clashing molecule and the nucleic acid structure, respectively, to incorporate a chemical modification relative to a second reference molecule that comprises natural nucleosides and has the same sequence content as the clashing molecule or the nucleic acid structure, as the case may be. The modification causes binding of the clashing molecule and the nucleic acid structure to have a hybridization free energy, differing from that of a second reference binding between the clashing molecule or the nucleic acid structure and the second reference molecule, such that binding of the clashing molecule is impeded relative to the second reference binding.
[0008] The invention also provides, in another of its aspects, a system that includes (A) a nucleic acid structure that comprises an incompletely base-paired duplex domain; (B) at least one polynucleotide displacement molecule to effect binding with the nucleic acid structure under hybridizing conditions, such that the nucleic acid structure undergoes a transition in energy state due to a branch migration reaction involving the duplex domain; and (C) at least one polynucleotide clashing molecule capable of binding with the nucleic acid structure under hybridizing conditions, such that the nucleic acid structure and the clashing molecule are bound and prevent the binding of the polynucleotide displacement molecule. At least one of the displacement molecule and the nucleic acid structure, respectively, has a chemical modification relative to a reference molecule that comprises natural nucleosides and has the same sequence content as the displacement molecule or the nucleic acid structure, as the case may be. The modification causes the binding to have a hybridization free energy, differing from that of a reference binding between the displacement molecule or the nucleic acid structure and a reference molecule, such that the branch migration reaction is facilitated relative to the reference binding. In addition or in the alternative, at least one of the clashing molecule and the nucleic acid structure, respectively, has a chemical modification relative to a reference molecule that comprises natural nucleosides and has the same sequence content as the displacement molecule or the nucleic acid structure, as the case may be. The modification causes the binding to have a hybridization free energy, differing from that of a reference binding between the displacement molecule or the nucleic acid structure and a reference molecule, such that binding of the clashing molecule is impeded relative to the reference binding.
[0009] In yet another aspect, a system is provided for propagating information. The system includes a first molecule and a second molecule configured to bind with the first molecule. At least one of the first molecule and the second molecule, respectively, has a chemical modification relative to a reference molecule that has the same sequence content as the first molecule or the second molecule, as the case may be. The modification causes the binding to have a free energy different from that of a reference binding between the reference molecule and the first or second molecule, such that at least one of the following is realized in the system: a probability of a productive binding is raised, or a probability of an unproductive binding is reduced.
BRIEF DESCRIPTION OF THE DRAWINGS
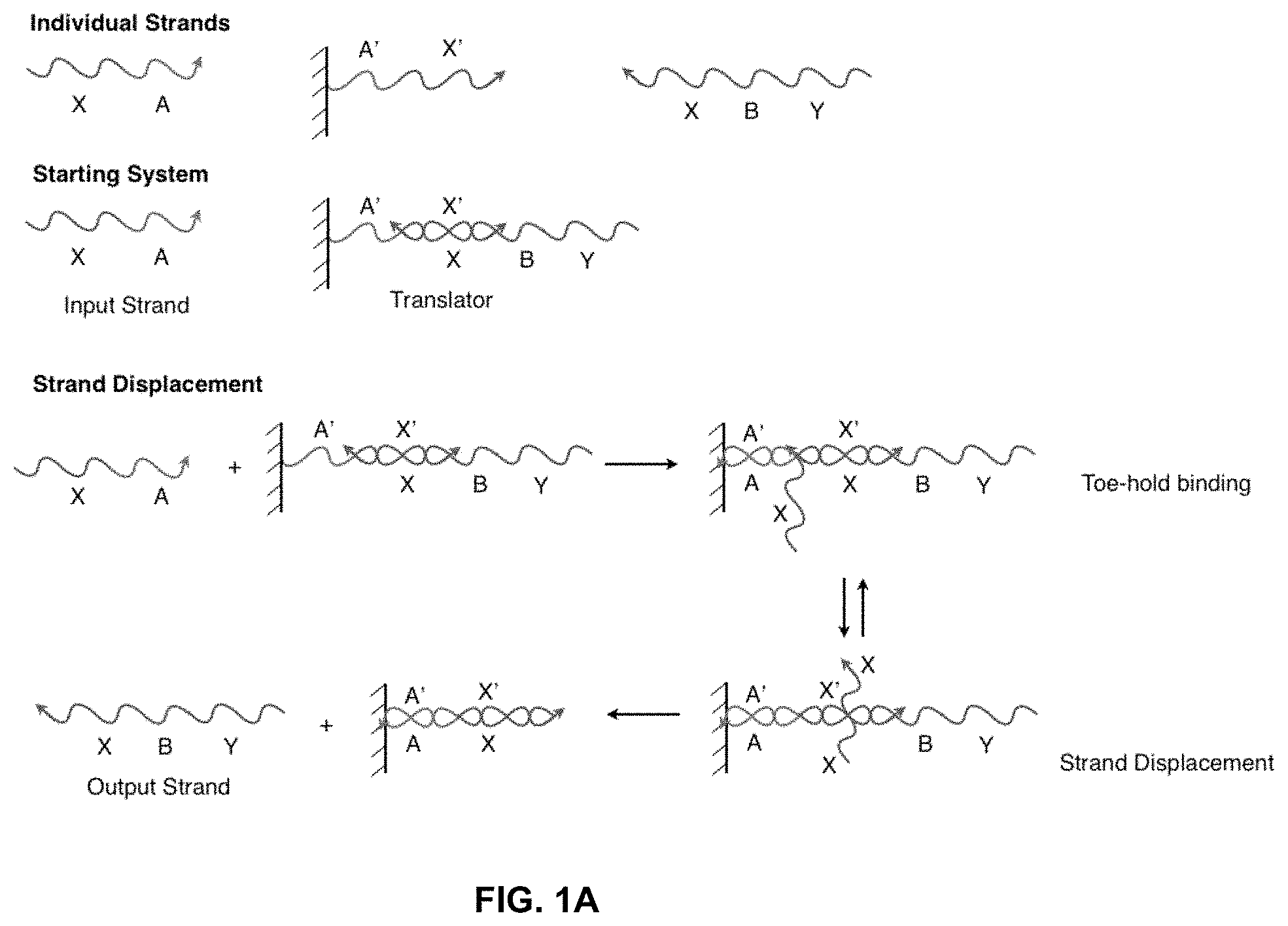
[0010] FIG. 1A illustrates a solid-phase sequestering implementation of a nucleic acid translator, where individual section (A, B, etc) represent stretches of oligonucleotides of arbitrary length and sequence.
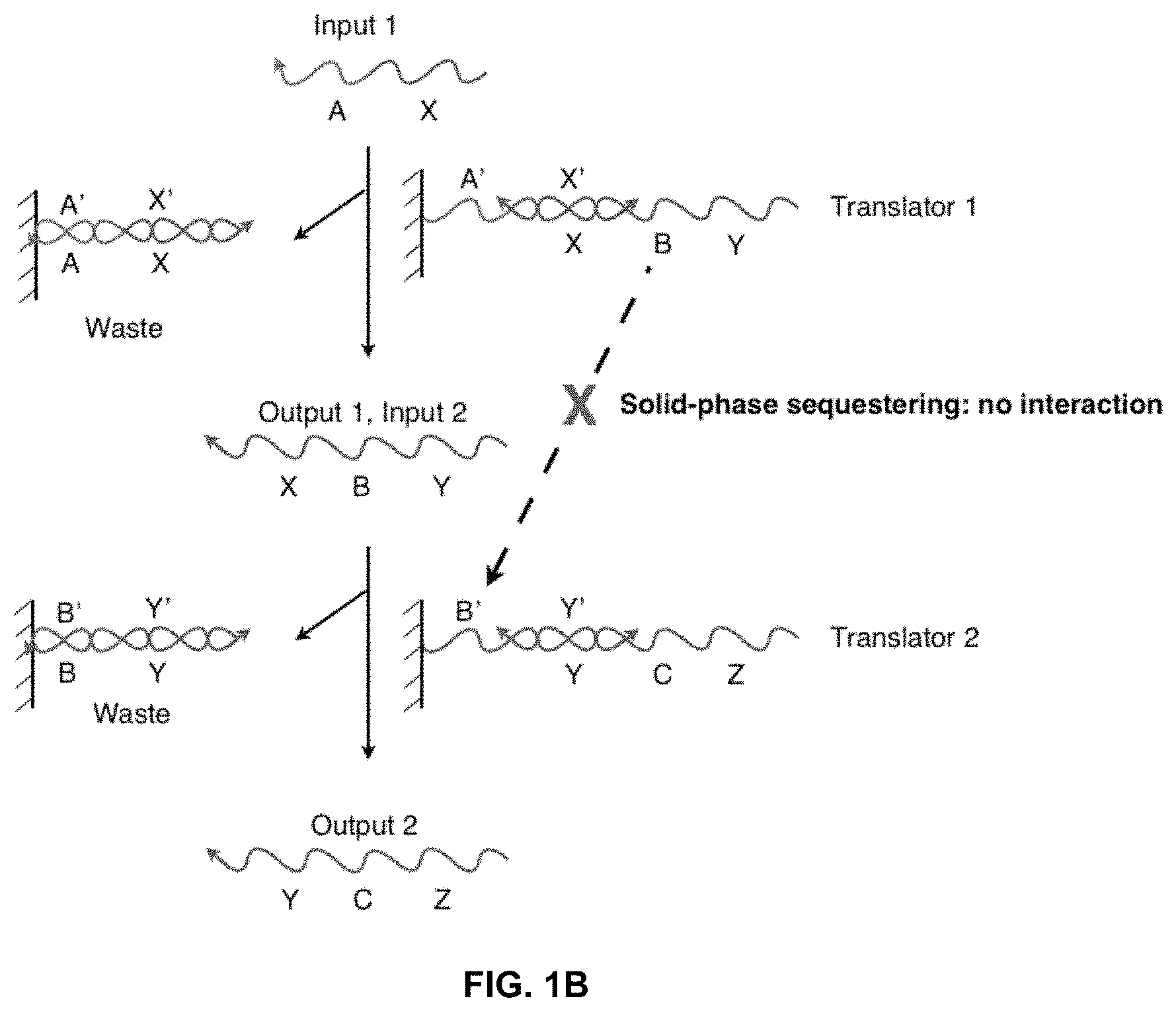
[0011] FIG. 1B illustrates how a solid-phase sequestering implementation of a nucleic acid translator sequesters stretches of oligonucleotides.
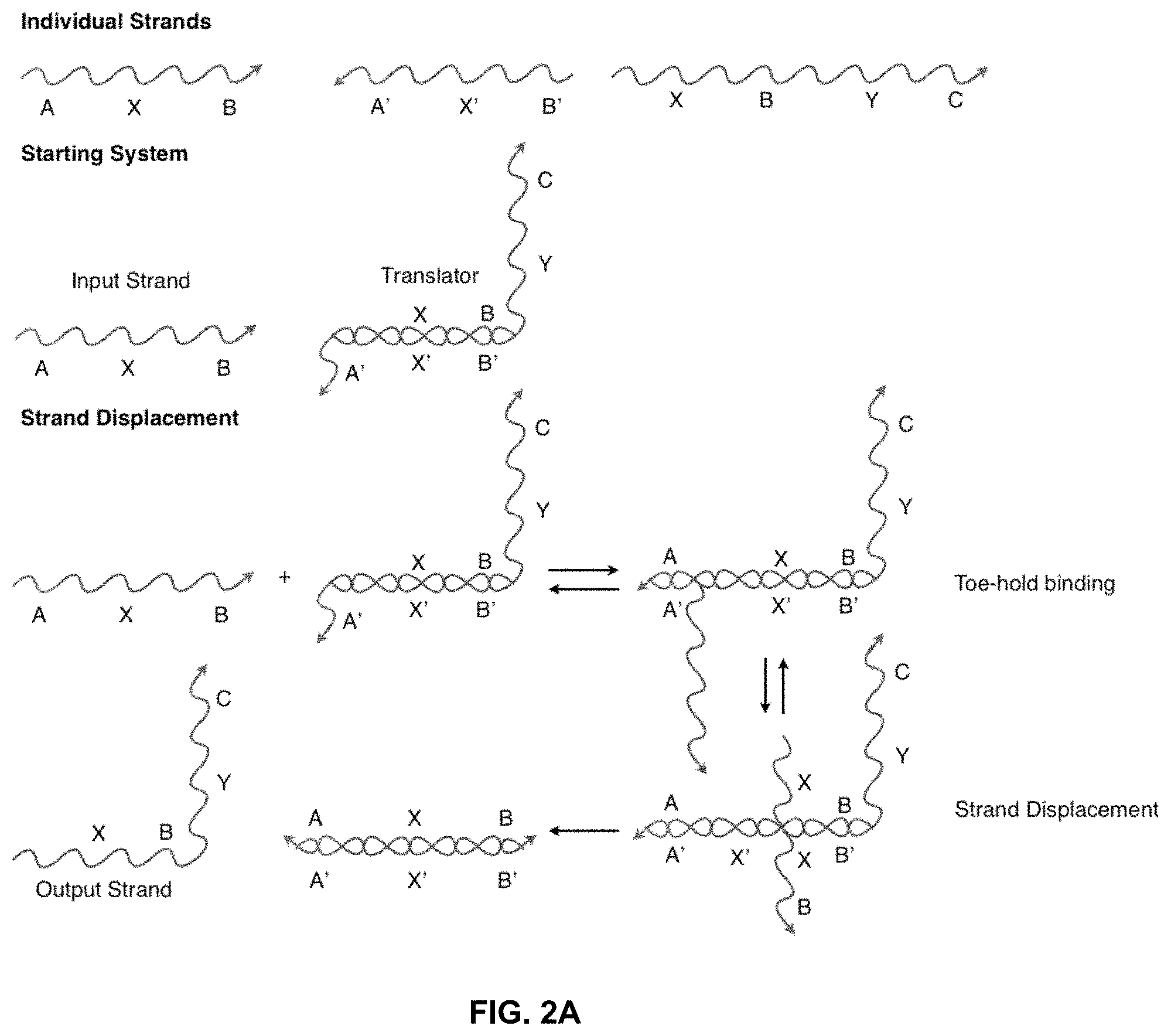
[0012] FIG. 2A illustrates a "toe-hold" sequestering implementation of a nucleic acid translator. Again, individual sections represent stretches of oligonucleotides of arbitrary length and sequence.
[0013] FIG. 2B illustrates how a "toe-hold"-sequestered implementation of a nucleic acid translator sequesters stretches of oligonucleotides.
[0014] FIG. 3A shows a system of three toe-hold sequestered nucleic acid translators. The reactions shown are all strand-displacement reactions that proceed by the same branch-migration mechanism as in FIG. 2A.
[0015] FIG. 3B shows a system with the same oligonucleotides as are shown in FIG. 3A but, instead of strand displacement reactions, toe-hold "clashes" are shown, where the toe-hold is bound by a sequence that cannot produce a strand displacement reaction. This binding event occupies the toe-hold such that the desired strand cannot bind.
[0016] FIG. 3C illustrates different architectures for incompletely base-paired nucleic acid structures including: (a) terminal; (b) internal loop; and (c) and (d) multi-part complex.
[0017] FIG. 4 presents a schematic depiction of how, in the system of nucleic acid translators shown in FIG. 3A, the equilibrium can be shifted to favor strand displacement reactions over toe-hold clashes by replacing the stretches of oligonucleotides with chemical modifications that increase the binding affinity and/or the stretches of oligonucleotides with modifications that decrease the binding affinity.
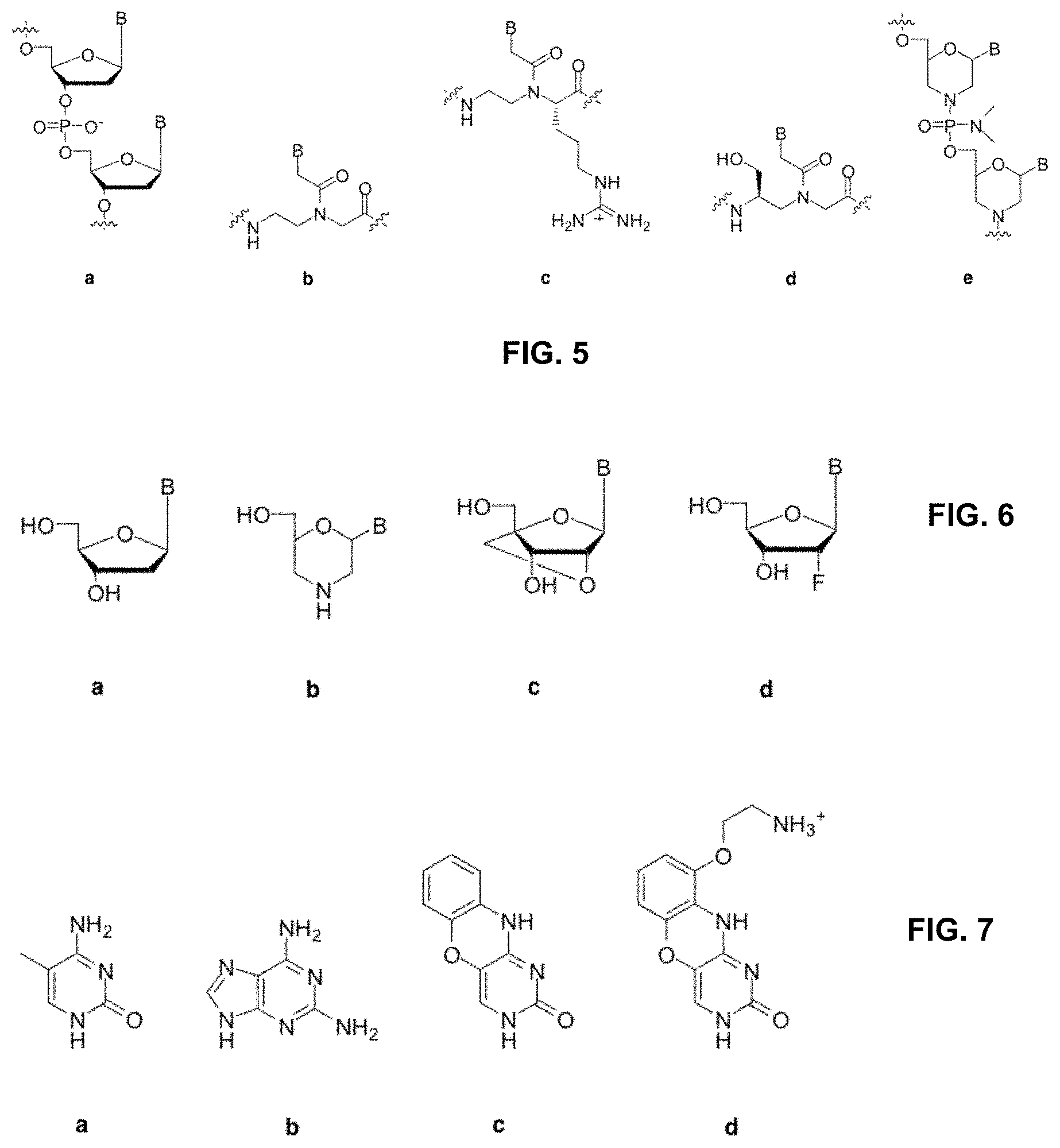
[0018] FIG. 5 shows modified backbone structures for nucleic acid analogs, where "B" represents an arbitrary nucelobase, and (a) shows a natural phosphodiester backbone found in DNA, (b) shows peptide nucleic acids, (c) shows guanidinium peptide nucleic acids, (d) shows L-serine derived gamma-PNAs, and (e) shows phosphorodiamidates (here with a morpholino sugar).
[0019] FIG. 6 illustrates modified sugar structures for nucleic acid analogs, where "B" represents an arbitrary nucleobase, and (a) shows a natural deoxyribose sugar found in DNA, (b) shows morpholinos, (c) shows locked nucleic acids, and (d) shows fluorine-modified RNA derivative.
[0020] FIG. 7 illustrates modified nucelobase structures for the nucleic acid analogs methylcytosine (a), diaminopurine (b), phenoxazine (c), and G-clamp (d).
[0021] FIG. 8 illustrates a fluorescence kinetics assay for indicating the reaction rate change due to a chemical modification according to the invention.
DETAILED DESCRIPTION
[0022] Various approaches, such as solid-phase sequestering, toe-hold sequestering, and toe-hold exchange, can be used in translating nucleic acid sequences to build logic operators and networks. These three particular approaches, described in greater detail below, are exemplified via geometries that utilize three-way, toe-hold-mediated branch migration reactions. Additional mechanisms are possible for branch migration reactions, however, including but not limited to four-way branch migration, four-way accelerated migration, and multi-strand complex migration.
[0023] Thus, while the following embodiments are described using three-way branch migration for illustration purposes, the present invention contemplates DNA logic gates and networks built to utilize other branch migration pathways. Conversely, the embodiments of the invention can be applied to any branch migration reaction.
[0024] Solid-phase sequestering uses physically separating the relevant sequences/strands in space, using beads, nanoparticles, or surfaces to do so. This approach employs principles of site isolation, which has been used extensively in the organic chemistry context. In solid-phase sequestered geometries, when these displacement events occur can be controlled by regulating whether the necessary strands are in the solution or in the solid phase of the system.
[0025] FIG. 1A shows a basic solid-phase sequestering setup for a translator--a component that allows a system to substitute one nucleic acid sequence for another. Here, the strand A'-X' (where A and X each represent a stretch of oligonucleotides of arbitrary length and sequence and X' and A' represent their respective reverse complements) is bound to a solid support and is initially hybridized to Y-B-X, forming a nucleic acid structure in the form of an incompletely base-paired duplex, which can function as a translator. In this configuration, the strand Y-B-X is solid-phase sequestered and cannot interact with the rest of the system. However, in the presence of the strand X-A, referred to as a "polynucleotide displacement molecule," strand Y-B-X can be displaced from the solid support and exposed to the solution phase of the system, while strand X-A is bound to the support. This operation involves two steps, the first of which is the hybridization of complementary sequences A and A' (often referred to as toe-hold binding). In the second step, the X region of strand X-A binds to the X' region of A'-X', displacing the X region of Y-B-X and releasing this strand into solution while leaving X-A bound to the solid support (this step is often referred to as a branch migration reaction). This two-step process effectively allows for the translation of a free X-A strand into a free Y-B-X strand.
[0026] FIG. 1B shows a system having an input strand X-A interacting with an incompletely base-paired duplex, "Translator 1." The output includes a completely base-paired duplex, A'-X'/X-A, which is considered a "waste" product, and an output strand Y-B-X, which is referred to as "Output 1" and can be used as "Input 2" in a further reaction. "Input 2" interacts with "Translator 2" and produces the "Output 2" and another waste product. In this figure, the B region of strand Y-B-X illustrates the sequestering of sequences in this network. At the start, Y-B-X cannot hybridize with the B' region of Translator 2 because both are isolated on separate solid-supports. When Input 1 binds to Translator 1 and releases Y-B-X into solution, Y-B-X can then interact with Translator 2. Therefore, the ability of Y-B-X and Translator 2 to interact is conditional on the presence of Input 1.
[0027] The strands bound to one solid surface interact extremely slowly with strands on another solid surface due to steric effects. Consequently, the strands in the solution phase are the only components that can interact with the solid phase operators.
[0028] Toe-hold sequestering and toe-hold exchange are separate approaches that use similar pairing interactions, but with different geometries. Both can accomplish the same operations as solid-phase translators, but function by keeping stretches of sequence bound up in a duplex. Just as in the solid-phase translator, a displacement event can free the sequence of interest. For both toe-hold geometries, all of the strands can be in solution together, by the consequence of which displacement events are regulated by the availability of toe-holds, i.e., short stretches of single-strand nucleic acid sequences that provide a starting point for a displacement event.
[0029] FIG. 2A shows a toe-hold sequestered translator similar to the one in FIG. 1A, but based on toe-hold rather than solid-phase sequestering. In this example, the A' region of the translator is the toe-hold that binds the input strand and allows the strand displacement reaction to proceed.
[0030] FIG. 2B shows a toe-hold-based system having an input strand A-X-B interacting with an incompletely base-paired duplex, "Translator 1." The output includes a waste product, i.e., a completely base-paired duplex, B'-X'-A'/A-X-B, and "Output 1" strand X-B-Y-C, which can be used as "Input 2" in a further reaction. "Input 2" interacts with "Translator 2" and produces the "Output 2" and another waste product. In this figure, the B region of X-B-Y-C is sequestered in Translator 1 by being hybridized to a complementary B' region and therefore unable to interact with the B' region of Translator 2. The ability of X-B-Y-C to interact with Translator 2 is conditional on the presence of Input 1 (A-X-B) in the system.
[0031] The toe-hold geometries have the potential to be very useful, but their use to date has been limited by the rate at which a system containing such toe-hold geometries can propagate information. This is due to inherent limitations, present in the current toe-hold sequestered approaches, which slow them down below a biologically useful timescale. These kinetic bottlenecks are a result of unproductive reactions, referred to here as "toe-hold clashes," that occur when a toe-hold is bound by a molecule having a complementary sequence or "clashing strand" that cannot produce a displacement reaction.
[0032] FIG. 3A shows a system of three toe-hold sequestered nucleic acid translators, much like the one in FIG. 2A. If all three strands are in solution together, however, there are other binding events that can take place. FIG. 3B illustrates some of the non-productive binding events or clashes that can take place. By involving a "polynucleotide clashing molecule," these events do not lead to a displacement reaction but can slow the system down, because the incidence of a clashing strand blocks strands from binding that can produce a displacement reaction.
[0033] Toe-holds can be kept short to mitigate the effect of these clashes on the system: the shorter the toe-hold is, the faster the on/off rate of the complementary sequence can be. Thus, five or six nucleotide-long toe-holds are common because at these lengths, if a non-productive binding event occurs, the time spent in the double-stranded, "clashed" state is short. This approach creates the aforementioned kinetic bottleneck, however, because the productive binding event is constrained by the same thermodynamic parameters; hence, the incoming strand likewise does not bind strongly to these toe-holds. Consequently, the desired displacement does not always occur when the correct incoming strand binds, as it needs to be in the bound state long enough to initiate the displacement reaction. The use of short toe-holds thus increases the amount of time required for a given operation to occur and produce an output. Put another way, the displacement reaction cannot take place before the occurrence of many binding events, both by clashing strands and by desired strands. This inefficiency limits the utility of the system by slowing down the propagation of information to time scales that are too extended to be useful.
[0034] Pursuant to one aspect of the invention, by virtue of chemical modification of the nucleic acid structure and/or the polynucleotide displacement molecule, the productive interactions are facilitated. This makes it feasible for chaining individual logic gates together into networks of arbitrary size for biological applications. More specifically, an approach is provided to determine how to disfavor clashing interactions and favor productive interactions without altering the information content of the sequences.
[0035] In accordance with the invention, these approaches improve the thermodynamics of binding for the desired strand and/or disfavor the thermodynamics for the clashing strands, all without altering the sequence content. In particular, using chemically-modified structures, the Gibbs free energy (AG) for the desired hybridization reaction between a given oligonucleotide and its DNA or RNA complement is reduced and/or the AG of the clashing interaction is increased. On a molecular level, these respectively correspond to a tighter binding (higher binding affinity) of desired hybridization reaction and to a less tight binding (lower binding affinity) for clashing strands, resulting from the modification of the structures.
[0036] The former shifts the equilibrium for desired binding events with the toe-hold toward the duplex (bound) state, creating a better chance for the displacement reaction to occur without affecting the equilibrium of binding between the toe-hold and clashing strands. The latter disfavors the binding of the clashing strand(s), making these interactions favor the unbound state. These two changes can be used separately or together, in order to favor the binding of the desired strand and disfavor the binding of the clashing strand(s).
[0037] Embodiments in accordance with the invention take advantage of the fact that the reaction rate of any toe-hold mediated strand displacement is related to the thermodynamic favorability of the two nucleic acid strands or complexes being bound. By making these thermodynamics more favorable for the non-clashing binding events and/or less favorable for clashing binding events, one can drive the system toward productive displacement reactions. This will speed up the rate at which information is propagated to the point that it can be used on biologically relevant timescales. The approach of the invention, described here, accomplishes this through chemical modifications to stretches of oligonucleotides in the system.
[0038] By virtue of such chemical modification of the nucleic acid structure and/or of the polynucleotide molecule, pursuant to invention, the effective concentration range is widened over which the reaction is optimal. This happens because the percentage of nucleic acid in a duplex at a given temperature is a function of concentration and of the AG of the hybridization reaction. Thus, the more concentrated a set of complementary oligonucleotides is, the higher the percentage of duplex formation and the lower (more favorable) the AG is, the higher the percentage of duplex formation. Changing the AG for a given interaction therefore changes the percent duplex at a given concentration. Lowering the AG (i.e., making the interaction more favorable) means that there will be a higher percentage of duplex at a given concentration, while raising the AG (making the interaction less favorable) means there will be a lower percentage of duplex at the same concentration. This effect widens the optimal concentration range at which these reactions can be practically used, as desired duplexes can be formed at lower concentrations and undesired duplexes will not be formed at higher concentrations, relative to a reference set of unmodified oligonucleotides.
[0039] These approaches apply to any toe-hold-mediated reaction geometry, including but not limited to the 3-way branch migration reactions discussed above. Different architectures can be used for the incompletely base-paired duplex where branching or loops are located at different points in the structure (see FIG. 3C, for instance). The orientation of the branches or loops is not predicated on the directionality of the strand. Thus, these structures can include (a) a terminal structure, (b) an internal loop, or (c) and (d) a multi-part complex, or any other possible architectures for an incompletely base-paired duplex.
[0040] These nucleic acid structures can translate an active "input" sequence (the displacement polynucleotide) into an active "output" sequence (the polynucleotide which is released).
[0041] The nucleic acid structures called "translators" above can be at their respective thermodynamic minimum; i.e., they are the most stable structures that the particular set of nucleic acid sequences can form. These structures can be formed by annealing the two or more individual nucleic acid strands. For instance, all of the strands can be mixed together, heated to well above the melting point for any structure to form, and then slowly cooled down. This allows the strands to hybridize in the lowest possible energy state (thermodynamic minimum).
[0042] This procedure can be the same for natural nucleic acids like DNA and RNA, nucleic acids with modified backbones, sugars, or bases, and for chimeras made up of natural and modified nucleic acids.
[0043] Selective incorporation of specific chemical modifications can favor the binding of strands capable of productive displacement reactions over toe-hold clash binding events. For example, FIG. 4 shows the same strands as FIG. 3A and 3B. However, if certain stretches of nucleic acids are replaced with chemically modified analogs that improve the thermodynamics of binding, the kinetics for clashes shifts in favor of the displacement reactions. For example, if the "B" stretch of oligonucleotides in the strand X-B-Y-C (circled with a solid line) has a higher binding affinity for B' than the "B" stretch of oligonucleotides in the strand A-X-B, the equilibrium shifts in favor of the productive, strand displacement reaction, which increases the probability or frequency with which this reaction occurs. Both B's have the same Watson-Crick base pairing sequence, but the interaction between X-B-Y-C and the B' toe-hold is more favorable due to chemical modifications.
[0044] Similarly, if certain stretches of nucleic acids are replaced with analogs that disfavor binding, the equilibrium will shift away from the clashes as well. For instance, if "B" in the strand A-X-B (circled with a dashed line) has a lower binding for B' than "B" in X-B-Y-C, then the equilibrium will shift away from the toe-hold clash, improving the kinetics for a strand displacement reaction.
[0045] Both approaches accelerate the speed at which a nucleic acid network can process signals because both increase the residence time of the desired strand on the toe-hold relative to the residence time of the clashing strand or strands. This provides a better chance that the strand displacement reaction will occur in a given period of time and thus increases the rate at which the network evaluates a given input or set of inputs. These modifications have not previously been used to facilitate or disfavor displacement reactions, to build significant DNA logic gates and networks, or to propagate information.
[0046] There are many well-characterized nucleic acid modifications that can be used in accordance with embodiments of the invention to improve or reduce the thermodynamic properties of binding to natural DNA or RNA. These include changes to the backbone, sugar, or nucleobase of the oligonucleotide. These modifications also can be used separately or in conjunction with one another; that is, using a modified backbone does not preclude the use of a modified nucleobase in the same strand.
[0047] Nucleic acid backbone analogs can be used to improve the binding of strands capable of producing a displacement reaction. There are a number of different analogs that could be used, all of which offer tighter binding affinities to DNA and RNA than natural nucleic acids. These analogs include but are not limited to those with uncharged backbones (peptide nucleic acids or phosphorodiamidates), positively charged backbones (guanidinium peptide nucleic acids), and hydrogen-bonding groups that allow for pre-organization (gamma peptide nucleic acids).
[0048] The general structures for certain analogs are shown in FIG. 5. These analogs all improve the thermodynamics of nucleic acid hybridization reactions, allowing for tighter toe-hold binding and therefore faster displacement. By using these analogs in specific places of the logic network, the desired displacement reactions can be strongly favored in comparison to the clashing interactions, thus accelerating the rate of information processing.
[0049] The use of modified sugar rings can also alter the thermodynamics of binding to DNA or RNA for an oligonucleotide. The most widely used analogs are morpholinos, locked nucleic acids (LNAs), and LNA derivatives. Other modified sugars are documented in the literature that also could produce a similar result, in terms of altering binding thermodynamics. Illustrative of these are sugars with modifications at the 1', 2', 3' or 4' position and sugars with different atoms substituted for the oxygen in the ribose cyclopentane ring. These analogs are illustrated in FIG. 6.
[0050] Nucleobase modifications can also be used to achieve the same effect as backbone and sugar analogs; namely, altering the thermodynamics of specific hybridization reactions. These bases include methylcytosine, diaminopurine, G-clamp, and phenoxazine (FIG. 7), all of which improve the binding affinity of a strand for its reverse complement. Another opportunity that exists with nucleobase modification involves pseudocomplementary bases. This class of base analogs forms weak base pairs with one another but forms strong base pairs with standard bases. One such pair of bases is 2-aminoadenine (nA) and 2-thiothymine (sT).
[0051] These bases could be used to favor one strand binding while disfavoring another, an example of increasing the likelihood of a productive binding event while decreasing the likelihood of a clash at the same time.
[0052] Another chemical modification that could be used to alter the thermodynamics of binding interactions is the incorporation of charged polymers like chitosan, which has been shown in the literature to accelerate the rate of displacement reactions. However, since these polymers speed up reactions non-specifically, they would have to be used in conjugation with one of the other modifications mentioned above to allow for discrimination between desired and undesired binding events.
[0053] Different approaches can be used to synthesize the molecules with the chemical modifications discussed above. For example, the backbone chemistry can be taken into consideration of the design of the modified molecules. Backbone chemistry is what is used to put together individual monomers into a longer strand. Modifications that involve the nucleobase or the sugar but that keep the natural phosphodiester backbone of DNA/RNA can be synthesized via standard phosphoramidite chemistry, as employed for natural monomers. Illustrations of these methods are found, for example, in Beaucage, S., and R. Iyer, Tetrahedron 48: 2223 (1992), in Brown, D. M. A, "Brief history of oligonucleotide synthesis," 20 METHODS IN MOLECULAR BIOLOGY (Protocols for Oligonucleotides and Analogs) 1-17 (1993), in Reese, Colin B., Organic & Biomolecular Chemistry 3: 3851 (2005), and in Iyer, R. P.; and S. L. Beaucage, "7.05. Oligonucleotide synthesis," 7 COMPREHENSIVE NATURAL PRODUCTS CHEMISTRY (DNA and Aspects of Molecular Biology) 105-52 (1999), the respective contents of which are hereby incorporated by reference in their entirety.
[0054] If the backbone is being changed in a particular modification, different chemistry will be employed. Such modification chemistry is generally known in the scientific literature. Thus, peptide nucleic acids (PNAs) and their derivatives rely on amide bonds to link the individual monomers together. Instead of using phosphoramidite chemistry, therefore, strands of these monomers are made with amide bonding forming conditions and coupling reagents like HBTU. An exploration of the methods used to make PNA or PNA-like oligonucleotides can be found, for instance, in F. Beck, "Solid Phase Synthesis of PNA Oligomers," METHODS IN MOLECULAR BIOLOGY SERIES (Peptide Nucleic Acids), Humana Press, http://www.springerlink.com/content/mr571738x7t65067/.
[0055] Another backbone modification approach involves chimeric oligonucleotides. These are oligonucleotide strands that contain different backbone chemistries in the same molecule. For example, if one needed a strand that was half PNA backbone and half DNA backbone, one would need a way to join these two different backbone chemistries. Making these chimeric strands is also generally known in the art. In the above example of a PNA/DNA chimera, the difference in chemistries can be bridged by using modified DNA or PNA monomers. For DNA, the 5'-dimethoxytrityl (DMT) protected hydroxyl is replaced with a monomethoxytrityl (MMT)-protected amine that can react with the carboxylic acid of a PNA after deprotection. For PNA, the protected N-terminal nitrogen is replaced with a DMT-protected hydroxyl that can react with the phosphoramidite group on DNA after de-protection. These approaches are more described in E. Uhlmann, et al., Angew. Chem. (Int'l ed.) 37: 2796-823 (1998), for example.
[0056] All of these modifications, whether used individually or in conjunction with one another, can affect the thermodynamic conditions of specific interactions in an arbitrary nucleic acid network such that the binding of desired strands or complexes is favored over clashing interactions without altering sequence content. All of these interactions can apply to any branch-mediated migration reaction, whether they are 3-way branch migrations, such as solid-phase sequestering, toe-hold sequestering, or toe-hold exchange, or are branch migrations that take place by other mechanisms, e.g., four-way branch migration, four-way accelerated branch migration, or multi-strand complex migration.
[0057] In accordance with some embodiments, the kinetics of the chemically modified molecules for improved branch migration reaction can be tested. One approach is to measure the improvement that different modifications provide in accelerating or retarding the branch migration reaction versus a natural polynucleotide using a fluorescence kinetics assay (see FIG. 8, for example). Here, the translator structure has a quencher on one strand and a fluorophore on the other. In the absence of the displacement strand, the fluorescence of the fluorophore is quenched, providing a baseline for the reaction. When the displacement molecule, which could be modified in the toe-hold and/or non-toe-hold regions, is added to the translator, it will displace the strand with the quencher and in doing so produce a fluorescent signal. Therefore, the rate of fluorescence "turn-on" in the system indicates the kinetics of the displacement reaction. This assay can be made more complex by requiring a larger circuit to evaluate before a final, fluorescence-producing displacement reaction or clashing strands could be added to compete with the displacement strand for the toe-hold.
[0058] The approaches in accordance with the invention also can apply to any molecules, natural or artificial, that are suitable for propagating information, so long as a first molecule and a second molecule are involved, with the latter configured to bind with the former. At least one of the first and second molecules, respectively, has a chemical modification relative to a reference molecule that has the same sequence content as the first molecule or the second molecule, as the case may be. The modification causes the binding to have a free energy different from that of a reference binding between the reference molecule and the first or second molecule, such that at least one of the following is realized in the system: a probability of a productive binding is raised; or a probability of an unproductive binding is reduced.
[0059] Pursuant to the invention, as indicated above, a multistep process could be employed to achieve an improved nucleic acid network. This would entail examining the full network of nucleic acid translators under consideration, including any clashing interactions, and then identifying the stretches of sequence that need to be modified in order best to facilitate displacement events and to minimize clashing.
[0060] With respect to any given strand displacement reaction, these modifications could be made on the desired displacement molecule, on the clashing molecule, or on both. This is so because, for any translator or logic gate with an available toe-hold, there will be a competition between the desired displacement molecule and any clashing molecule to bind the toe-hold. Furthermore, increasing the ability of the displacement molecule to bind the toe-hold relative to the clashing molecule will increase the rate at which the displacement reaction takes place. Thus, increasing the ability of the displacement molecule to bind (lowering .DELTA.G of binding) or decreasing the ability of the clashing molecule(s) to bind (raising .DELTA.G of binding) will have the effect of increasing the rate of the reaction. Since both of these changes in .DELTA.G can be accomplished via chemical modifications, pursuant to the invention, it is important when considering which stretches of sequence to modify that one should explore the possibility of modifying the displacement molecule alone, the clashing molecule alone, and both of these together.
[0061] According to embodiments of the invention, two parameters that could be varied in this analysis are (i) the rate constants for the interaction between any two oligonucleotides and (ii) the length of the toe-holds for all of the oligonucleotides in the network. The former parameter would be dictated by the chemical composition of the oligonucleotide and the .DELTA.G of its interactions with other oligonucleotides. The latter parameter would be the number of nucleotides in the toe-hold region of sequence under consideration.
[0062] The kinetic rate constants needed to model a nucleic acid network could be calculated from the .DELTA.G of hybridization between any two oligonucleotides. For the natural bases and many of the modifications, these values, which come from nearest-neighbor parameters, have been reported in the literature. If they are not known already, they can be experimentally determined through Van't Hoff analysis of melting temperature data, differential scanning calorimetry, or isothermal titration calorimetry. Each of these methods would yield the thermodynamic parameters for the nucleic acid interaction in question, which then can be used to determine the .DELTA.G of the reaction at any temperature. The .DELTA.G value then would be used to determine the equilibrium constant (K.sub.eq) of the interaction. The equilibrium constant would be proportional to the ratio of rate constants; hence, one could use known constants to solve for the necessary kinetic rate constant for a given reaction.
[0063] Thus, the full set of rate constants would be obtained for the modifications under consideration for use in a nucleic acid logic network, along with the lengths for all of the toe-holds. With this information the system could be modeled to determine the time course behavior of the network, and also to determine where specific modifications could be employed to optimize the productive displacement reactions relative to clashing interactions. Additionally, the length of specific toe-hold regions could be varied in the model. The network would be optimized by varying modifications at specific locations and by simulating the reactions in the network with different length toe-holds. These calculations would be employed to determine the optimal modification and toe-hold length for each component that produces the most favorable time course behavior of the network.
[0064] The network also could be optimized by experimentally changing the modifications and toe-hold lengths and then studying the time course behavior in the laboratory, as opposed to simulating the behavior with a mathematical model.
[0065] Given a fully optimized network, the necessary oligonucleotides could be synthesized with standard phosphoramidite chemistry or with the approach(es) reported in the literature for specific modifications. Any translator or gate structure that consists of more than one oligonucleotide could be constructed by mixing the individual oligonucleotides together and then annealing them, first by heating the mixture to above the melting temperature of all possible duplexes and then slowly cooling it. This would cause the oligonucleotides to hybridize, forming the most stable structure that the particular set of nucleic acid sequences can adopt. These structures then could be purified before they were used in the network. Once all of the translators, gates, and other components are designed, synthesized, and annealed, the network could be applied to the target assay, diagnostic, or biological system.
[0066] While particular embodiments of the subject invention have been discussed, they are illustrative only and not restrictive of the invention. A review of this specification will make many variations of the invention apparent to those skilled in the field of the invention. The full scope of the invention should be determined by reference both to the claims below, along with their full range of equivalents, and to the specification, with such variations.
Cited Publications
[0067] The following publications may aid understanding or practicing embodiments of the invention. Each cited publication is incorporated by reference in its entirety. [0068] (1) Picuri, J. M.; Frezza, B. M.; Ghadiri, M. R. J Am Chem Soc 2009, 131, 9368-77. [0069] (2) Li, Q.; Luan, G.; Guo, Q.; Liang, J. Nucleic Acids Res 2002, 30, E5. [0070] (3) Zhang, D. Y.; Turberfield, A. J.; Yurke, B.; Winfree, E. Science 2007, 318, 1121-5. [0071] (4) Frezza, B. M.; Cockroft, S. L.; Ghadiri, M. R. J Am Chem Soc 2007, 129, 14875-9. [0072] (5) Voelcker, N. H.; Guckian, K. M.; Saghatelian, A.; Ghadiri, M. R. Small 2008, 4, 427-31. [0073] (6) Yashin, R.; Rudchenko, S.; Stojanovic, M. N. J Am Chem Soc 2007, 129, 15581-4. [0074] (7) Seelig, G.; Soloveichik, D.; Zhang, D. Y.; Winfree, E. Science 2006, 314, 1585-8. [0075] (8) Zhang, D. Y.; Winfree, E. J Am Chem Soc 2009, 131, 17303-14. [0076] (9) Biswas, I.; Yamamoto, A.; Hsieh, P. J Mol Biol 1998, 279, 795-806. [0077] (10) Panyutin, I. G.; Hsieh, P. J Mol Biol 1993, 230, 413-24. [0078] (11) Panyutin, I. G.; Hsieh, P. Proc Natl Acad Sci USA 1994, 91, 2021-5. [0079] (12) Maugh, T. H., 2nd Science 1982, 217, 719-720. [0080] (13) Tajima, T.; Nakajima, A. Journal of the American Chemical Society 2008, 130, 10496-7. [0081] (14) Voit, B. Angew Chem Int Ed Engl 2006, 45, 4238-40. [0082] (15) Kahan, M.; Gil, B.; Adar, R.; Shapiro, E. Physica D-Nonlinear Phenomena 2008, 237, 1165-1172. [0083] (16) Nielsen, P. E.; Egholm, M. Curr Issues Mol Biol 1999,1, 89-104. [0084] (17) Dragulescu-Andrasi, A.; Zhou, P.; He, G.; Ly, D. H. Chem Commun (Camb) 2005, 244-6. [0085] (18) Sahu, B.; Chenna, V.; Lathrop, K. L.; Thomas, S. M.; Zon, G.; Livak, K. J.; Ly, D. H. J Org Chem 2009, 74, 1509-16. [0086] (19) Zhou, P.; Wang, M.; Du, L.; Fisher, G. W.; Waggoner, A.; Ly, D. H. J Am Chem Soc 2003, 125, 6878-9. [0087] (20) Dragulescu-Andrasi, A.; Rapireddy, S.; Frezza, B. M.; Gayathri, C.; Gil, R. R.; Ly, D. H. J Am Chem Soc 2006, 128, 10258-67. [0088] (21) Freier, S. M.; Altmann, K H Nucleic Acids Research 1997, 25, 4429-4443. [0089] (22) Braasch, D. A.; Corey, D. R. Chemistry & Biology 2001, 8, 1-7. [0090] (23) Summerton, J.; Weller, D. Antisense & Nucleic Acid Drug Development 1997, 7, 187-195. [0091] (24) Ortega, J. A.; Blas, J. R.; Orozco, M.; Grandas, A.; Pedroso, E.; Robles, J. Org Lett 2007, 9, 4503-6. [0092] (25) Kutyavin, I. V.; Rhinehart, R. L.; Lukhtanov, E. A.; Gorn, V. V.; Meyer, R. B., Jr.; Gamper, H. B., Jr. Biochemistry 1996, 35, 11170-6. [0093] (26) Lee, D.; Singha, K.; Jang, M. K.; Nah, J. W.; Park, I. K.; Kim, W. J. Mol Biosyst 2009, 5, 391-6. [0094] (27) U.S. Pat. No. 7,538,202. [0095] (28) U.S. Patent Application Pub. No. 2005/0112614. [0096] (29) U.S. Patent Application Pub. No. 2007/0072215. [0097] (30) U.S. Patent Application Pub. No. 2009/0191546.
* * * * *
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007
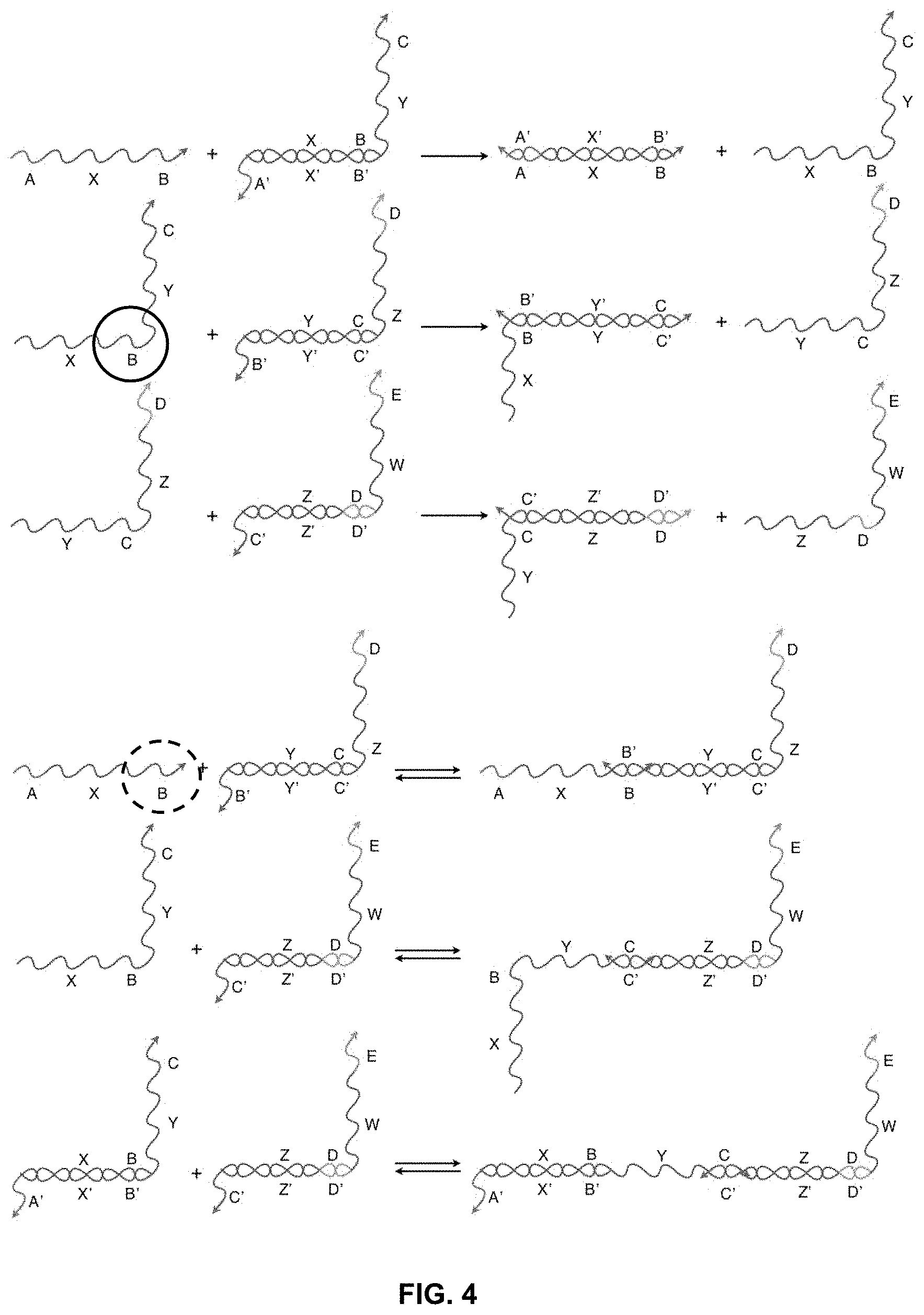
D00008

D00009

XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.