Bispecific CD33 and CD3 Binding Proteins
Ellwanger; Kristina ; et al.
U.S. patent application number 16/822528 was filed with the patent office on 2021-01-28 for bispecific cd33 and cd3 binding proteins. The applicant listed for this patent is Amphivena Therapeutics, Inc.. Invention is credited to Kristina Ellwanger, Luke Evnin, Judith A. Fox, Ivica Fucek, Jeanmarie Guenot, Stefan Knackmuss, Lori Kunkel, Melvyn Little, Vera Molkenthin, Erich Rajkovic, Uwe Reusch, Claudia Wall, Michael Weichel, Eugene Zhukovsky.
| Application Number | 20210024654 16/822528 |
| Document ID | / |
| Family ID | 1000005150081 |
| Filed Date | 2021-01-28 |










View All Diagrams
| United States Patent Application | 20210024654 |
| Kind Code | A1 |
| Ellwanger; Kristina ; et al. | January 28, 2021 |
Bispecific CD33 and CD3 Binding Proteins
Abstract
Described herein are binding proteins that specifically bind to human CD33, and in particular to bispecific binding proteins that specifically bind to human CD33 and human CD3. Also described herein are bispecific tandem diabodies that bind to CD33 and CD33, and their uses for immunotherapy of CD33.sup.+ cancers, diseases and conditions such as acute myeloid leukemia (AML).
| Inventors: | Ellwanger; Kristina; (Heidelberg, DE) ; Evnin; Luke; (San Francisco, CA) ; Fox; Judith A.; (San Francisco, CA) ; Fucek; Ivica; (Frankfurt, DE) ; Guenot; Jeanmarie; (San Francisco, CA) ; Knackmuss; Stefan; (Plankstadt, DE) ; Kunkel; Lori; (San Francisco, CA) ; Little; Melvyn; (St. Peter-Ording, DE) ; Molkenthin; Vera; (Tannesberg, DE) ; Rajkovic; Erich; (Schriesheim, DE) ; Reusch; Uwe; (Maikammer, DE) ; Wall; Claudia; (Bruehl, DE) ; Weichel; Michael; (Bischofsheim, DE) ; Zhukovsky; Eugene; (Mannheim, DE) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005150081 | ||||||||||
| Appl. No.: | 16/822528 | ||||||||||
| Filed: | March 18, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15796556 | Oct 27, 2017 | 10626190 | ||
| 16822528 | ||||
| 14937494 | Nov 10, 2015 | 9803029 | ||
| 15796556 | ||||
| 14642497 | Mar 9, 2015 | 9212225 | ||
| 14937494 | ||||
| 62111470 | Feb 3, 2015 | |||
| 62019795 | Jul 1, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61K 45/06 20130101; C07K 2317/31 20130101; C07K 16/468 20130101; A61K 31/198 20130101; A61K 31/4035 20130101; C07K 16/2809 20130101; C07K 2317/565 20130101; C07K 2317/732 20130101; C07K 2317/626 20130101; A61K 31/706 20130101; A61K 31/704 20130101; C07K 2317/92 20130101; C07K 2317/73 20130101; A61K 31/69 20130101; C07K 2317/622 20130101; A61K 39/39558 20130101; A61K 31/519 20130101; A61K 31/7068 20130101; A61K 31/7076 20130101; A61K 2039/505 20130101; C07K 16/2803 20130101; A61K 31/454 20130101; C07K 2317/56 20130101 |
| International Class: | C07K 16/46 20060101 C07K016/46; C07K 16/28 20060101 C07K016/28; A61K 45/06 20060101 A61K045/06; A61K 31/198 20060101 A61K031/198; A61K 31/4035 20060101 A61K031/4035; A61K 31/454 20060101 A61K031/454; A61K 31/519 20060101 A61K031/519; A61K 31/69 20060101 A61K031/69; A61K 31/704 20060101 A61K031/704; A61K 31/706 20060101 A61K031/706; A61K 31/7068 20060101 A61K031/7068; A61K 31/7076 20060101 A61K031/7076; A61K 39/395 20060101 A61K039/395 |
Claims
1.-30. (canceled)
31. A method for treating acute myelogenous leukemia (AML) comprising administering to an individual a protein comprising a first polypeptide and a second polypeptide, each polypeptide having at least four variable chain domains linked one after another, wherein each polypeptide comprises (i) a variable heavy chain (VH) domain specific to human CD33; (ii) a variable light chain (VL) domain specific to human CD33; (iii) a VH domain specific for human CD3, and (iv) a VL domain specific for human CD3, and wherein in each polypeptide, the four variable chain domains are linked with one after another by peptide linkers L1, L2 and L3 in the order: VL(CD3)-L1-VH(CD33)-L2-VL(CD33)-L3-VH(CD3); VH(CD3)-L1-VL(CD33)-L2-VH(CD33)-L3-VL(CD3); VL(CD33)-L1-VH(CD3)-L2-VL(CD3)-L3-VH(CD33); or VH(CD33)-L1-VL(CD3)-L2-VH(CD3)-L3-VL(CD33), wherein the VL domain specific to human CD33 comprises a CDR1 comprising a sequence selected from the group consisting of SEQ ID NOs:21-27, a CDR2 comprising a sequence selected from the group consisting of SEQ ID NOs:28-34 and a CDR3 comprising a sequence selected from the of the group consisting of SEQ ID NOs:35-41, and wherein the VH domain specific to human CD33 comprises a CDR1 comprising a sequence selected from the group consisting of SEQ ID NOs:42-48, a CDR2 comprising a sequence selected from the group consisting of SEQ ID NOs:49-55 and a CDR3 comprising a sequences selected from the group consisting of SEQ ID NOs:56-63.
32. The method of claim 31, wherein the CDR1, CDR2 and CDR3 of the VL domain specific to human CD33 comprise sequences selected from the group consisting of: (i) SEQ ID NOs: 21, 28 and 35, respectively; (ii) SEQ ID NOs: 22, 29 and 36, respectively; (iii) SEQ ID NOs: 23, 30 and 37, respectively; (iv) SEQ ID NOs: 24, 31 and 38, respectively; (v) SEQ ID NOs: 25, 32 and 39, respectively; (vi) SEQ ID NOs: 26, 33 and 40, respectively; and (vii) SEQ ID NOs: 27, 34 and 41, respectively.
33. The method of claim 31, wherein the CDR1, CDR2 and CDR3 of the VH domain specific to CD33 comprise sequences selected from the group consisting of: (i) SEQ ID NOs: 42, 49 and 56, respectively; (ii) SEQ ID NOs: 43, 50 and 57, respectively; (iii) SEQ ID NOs: 43, 50 and 58, respectively; (iv) SEQ ID NOs: 43, 50 and 59, respectively; (v) SEQ ID NOs: 43, 50 and 60, respectively; (vi) SEQ ID NOs: 44, 51 and 61, respectively; (vii) SEQ ID NOs: 45, 52 and 62, respectively; (viii) SEQ ID NOs: 46, 53 and 63, respectively; (ix) SEQ ID NOs: 47, 54 and 63, respectively; and (x) SEQ ID NOs: 48, 55 and 63, respectively.
34. The method of claim 31, wherein the VL and VH domains specific to CD33 comprise sequences selected from the group consisting of: (i) SEQ ID NO:1 and SEQ ID NO:11, respectively; (ii) SEQ ID NO:3 and SEQ ID NO:13, respectively; (iii) SEQ ID NO:4 and SEQ ID NO:14, respectively; (iv) SEQ ID NO:5 and SEQ ID NO:15, respectively; (v) SEQ ID NO:6 and SEQ ID NO:16, respectively; (vi) SEQ ID NO:7 and SEQ ID NO:17, respectively; (vii) SEQ ID NO:8 and SEQ ID NO:18, respectively; (viii) SEQ ID NO:9 and SEQ ID NO:19, respectively; and (ix) SEQ ID NO:10 and SEQ ID NO:20, respectively.
35. The method of claim 31, wherein the VH domain specific for human CD3 comprises a CDR1 comprising the sequence STYAMN (SEQ ID NO:72), a CDR2 comprising the sequence RIRSKYNNYATYYADSVKD (SEQ ID NO:73) and a CDR3 comprising the sequence HGNFGNSYVSWFAY (SEQ ID NO:74) or comprising the sequence HGNFGNSYVSYFAY (SEQ ID NO:75).
36. The method of claim 31, wherein the VL domain specific for human CD3 comprises a CDR1 comprising the sequence RSSTGAVTTSNYAN (SEQ ID NO:90), a CDR2 comprising the sequence GTNKRAP (SEQ ID NO:91), and a CDR3 comprising the sequence ALWYSNL (SEQ ID NO:92).
37. The method of claim 31, wherein the VL and VH domains specific to CD3 comprise sequences selected from the group consisting of: (i) SEQ ID NO:64 and SEQ ID NO:68, respectively; (ii) SEQ ID NO:65 and SEQ ID NO:69, respectively; (iii) SEQ ID NO:66 and SEQ ID NO:70, respectively; and (iv) SEQ ID NO:67 and SEQ ID NO:71, respectively.
38. The method of claim 31, wherein a VL domain specific to human CD33, the VH domain specific to human CD33, the VH domain specific for human CD3, and the VL domain specific for human CD3 comprise sequences selected from the group consisting of: (i) SEQ ID NOs:3, 13, 65 and 69, respectively; (ii) SEQ ID NOs:4, 14, 65 and 69, respectively; (iii) SEQ ID NOs:5, 15, 65 and 69, respectively; (iv) SEQ ID NOs:1, 11, 64 and 68, respectively: (v) SEQ ID NOs:4, 14, 66 and 70, respectively; (vi) SEQ ID NOs:5, 15, 66 and 70, respectively; (vii) SEQ ID NOs:3, 13, 64 and 68, respectively; (viii) SEQ ID NOs:3, 13, 67 and 71, respectively; (ix) SEQ ID NOs:4, 14, 64 and 68, respectively; (x) SEQ ID NOs:5, 15, 64 and 68, respectively; (xi) SEQ ID NOs:7, 17, 64 and 68, respectively; (xii) SEQ ID NOs:6, 16, 64 and 68, respectively; (xiii) SEQ ID NOs:6, 16, 67 and 71, respectively; (xiv) SEQ ID NOs:8, 18, 64 and 68, respectively; (xv) SEQ ID NOs:9, 19, 64 and 68, respectively; (xvi) SEQ ID NOs:9, 19, 67 and 71, respectively; and (xvii) SEQ ID NOs:10, 20, 64 and 68, respectively.
39. The method of claim 31, wherein linkers L1, L2 and L3 consist of about 12 or less amino acid residues.
40. The method of claim 31, wherein linkers L1, L2 and L3 are each independently selected from GGSGGS (SEQ ID NO:95), GGSG (SEQ ID NO:96) or GGSGG (SEQ ID NO:97).
41. The method of claim 31, wherein linkers L1 and L3 are GGSGGS (SEQ ID NO:95) and linker L2 is GGSG (SEQ ID NO:96) or GGSGG (SEQ ID NO:97).
42. The method of claim 31, wherein the four variable chain domains are linked with one after another by peptide linkers L1, L2 and L3 in the order: VL(CD3)-L1-VH(CD33)-L2-VL(CD33)-L3-VH(CD3).
43. The method of claim 31, wherein the AML is AML with Recurrent Genetic Abnormalities, AML with myelodysplasia-related changes, Therapy-related myeloid neoplasms, Myeloid sarcoma, Myeloid proliferations related to Down syndrome, Blastic plasmacytoid dendritic cell neoplasm, or AML not otherwise categorized.
44. The method of claim 31, wherein the AML is AML-M0, AML-M1, AML-M2, AML-M3, AML-M4, AML-M5, AML-M6, or AML-M7.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. patent application Ser. No. 14/937,494, filed Nov. 10, 2015, which is a continuation of U.S. patent application Ser. No. 14/642,497, filed Mar. 9, 2015 (now U.S. Pat. No. 9,212,225, issued Jan. 7, 2016), and claims the benefit of U.S. Provisional Application No. 62/111,470, filed Feb. 3, 2015, and U.S. Provisional Application No. 62/019,795, filed Jul. 1, 2014, all of which are incorporated herein by reference in their entirety.
SEQUENCE LISTING
[0002] The instant application contains a Sequence Listing which has been submitted electronically in ASCII format and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Jun. 29, 2020, is named Sequence Listing.txt and is 144 KB in size.
BACKGROUND OF THE INVENTION
[0003] Acute myeloid leukemia (AML) is an acute leukemia in adults and children. CD33 is expressed on the majority of myeloblasts in AML. CD33, in some reports, is generally restricted to early multilineage myeloid progenitors and absent from normal pluripotent hematopoietic stem cells.
SUMMARY OF THE INVENTION
[0004] Provided herein are binding proteins that specifically bind to human CD33, and bispecific binding proteins that specifically bind to human CD33 and human CD3. Also provided herein are anti-CD33 variable domains and anti-CD3 variable domains for generating a number of bispecific CD33/CD3 binding proteins, such as, for example, tandem diabodies. Also further provided herein are bispecific tandem diabodies that bind to CD33 and CD3 and their use for immunotherapy of acute myeloid leukemia (AML) and other hematologic malignancies, disorders or conditions.
[0005] In particular, the binding proteins are provided that show binding to both human as well as cynomolgus monkey CD33. It is demonstrated in the examples that these CD33/CD3 tandem diabodies can re-direct polyclonal CD3.sup.+ T-cells from healthy donors, as well as autologous T-cells from AML patients, to effectively lyse CD33.sup.+ AML cells at low E:T cell ratios. In this process, which is dependent on the presence of both CD33.sup.+ target cells and T-cells, re-directed T-cells are activated, as shown by induction of CD25 and CD69, and stimulated to proliferate. The anti-AML effect of these tandem diabodies is shown to be dependent on the concentration of the antibodies used as well as on the E:T cell ratio. The tandem diabody is tetravalent and has two binding sites for CD33 and two binding sites for CD3. A particular feature of the CD33/CD3 tandem diabodies described herein is that they facilitate potent and efficient apoptosis as a result of bivalent binding that confers avidity to each antigen, namely CD33 and CD3.
[0006] In summary, the provided CD33/CD3 binding proteins described herein, in particular tandem diabodies, induce potent cytolysis of CD33.sup.+ leukemic cells and primary AML cells in vitro. Examples of bispecific CD33/CD3 binding proteins in the antibody format of tandem diabodies demonstrate cytolytic activity in vivo in cell lines, primary AML cells and in in vivo models with AML cell lines and with patient derived primary AML cells. This indicates high in vivo activity especially noteworthy in the stringent AML PDX model. Further, examples of bispecific CD33/CD3 binding proteins in the antibody format of tandem diabodies demonstrate cytolytic activity ex vivo in samples from patients at all stages of AML, including newly diagnosed, relapsed and refractory patients.
[0007] Furthermore, these CD33/CD3 binding proteins described herein are able to achieve a significant lysis of CD33 expressing cells within about four hours. CD33/CD3 binding proteins accordingly exhibit high cytotoxicity at low CD33 densities on the cell surface as well as a high cytotoxicity at low effector: target (E:T) ratios. In addition, CD33/CD3 binding proteins described herein exhibit not only potent CD33 and CD3 binding affinities to the human proteins, but show also excellent crossreactivity with the respective cynomolgus monkey proteins, for example with human:cynomolgous K.sub.D ratios between 5 and 0.2. Furthermore, the CD33/CD3 binding proteins described herein show no significant induction of cytokine release in the absence of CD33.sup.+ target cells which is an essential component of the safety profile of these molecules. Moreover, the CD33/CD3 tandem diabodies described herein belong to the class of molecules that have half-lives in the approximate range of 8-24 h, which should allow convenient dosing.
[0008] In one aspect, provided herein are CD33 binding proteins that specifically bind to an epitope of human CD33. In some embodiments, the binding proteins comprise a heavy chain variable domain and a light chain variable domain that is derived from human.
[0009] In some embodiments, a CD33 binding protein has at least one binding site comprising a light chain variable domain and a heavy chain variable domain, wherein the light chain variable domain comprises a CDR1 consisting of the sequence selected from the group consisting of SEQ ID NOs:21-27, a CDR2 consisting of the sequence selected from the group consisting of SEQ ID NOs:28-34 and a CDR3 consisting of the sequence of the group consisting of SEQ ID NOs:35-41.
[0010] In some embodiments, a CD33 binding protein has at least one binding site comprising a light chain variable domain and a heavy chain variable domain, wherein the heavy chain variable domain comprises a CDR1 consisting of the sequence selected from the group consisting of SEQ ID NOs:42-48, a CDR2 consisting of the sequence selected from the group consisting of SEQ ID NOs:49-55 and a CDR3 consisting of a sequences selected from the group consisting of SEQ ID NOs:56-63.
[0011] In certain instances, the CDR1, CDR2 and CDR3 of the light chain variable domain is selected from the group consisting of SEQ ID NOs:21, 28 and 35; SEQ ID NOs:22, 29 and 36; SEQ ID NOs:23, 30 and 37; SEQ ID NOs:24, 31 and 38; SEQ ID NOs:25, 32 and 39; SEQ ID NOs:26, 33 and 40; and SEQ ID NOs:27, 34 and 41.
[0012] In certain instances, the CDR1, CDR2 and CD3 of the heavy chain variable domain is selected from the group consisting of SEQ ID NOs:42, 49 and 56; SEQ ID NOs:43, 50 and 57; SEQ ID NOs:43, 50 and 58; SEQ ID NOs:43, 50 and 59; SEQ ID NOs:43, 50 and 60; SEQ ID NOs:44, 51 and 61; SEQ ID NOs:45, 52 and 62; SEQ ID NOs:46, 53 and 63; SEQ ID NOs:47, 54 and 63; and SEQ ID NOs:48, 55 and 63.
[0013] In certain instances, the human CD33 binding site of a variable heavy chain domain and a variable light chain domain is selected from the group consisting of SEQ ID NO:1 and SEQ ID NO:11; SEQ ID NO:2 and SEQ ID NO:12; SEQ ID NO:3 and SEQ ID NO:13; SEQ ID NO:4 and SEQ ID NO:14; SEQ ID NO:5 and SEQ ID NO:15; SEQ ID NO:6 and SEQ ID NO:16; SEQ ID NO:7 and SEQ ID NO:17; SEQ ID NO:8 and SEQ ID NO:18; SEQ ID NO:9 and SEQ ID NO:19; and SEQ ID NO:10 and SEQ ID NO:20.
[0014] In some embodiments, the CD33 epitope is within .sub.62DQEVQEETQ.sub.70 (SEQ ID NO:94) amino acid residues 62-70 of SEQ ID NO:93) of human CD33.
[0015] In any of the above embodiments, the CD33 binding protein comprises at least one further functional domain. In some instances, the functional domain is an effector domain that binds to an effector cell. In certain instances, the effector domain is a CD3 binding site comprising at least one antibody variable heavy chain domain and at least one variable light chain domain forming an antigen binding site for human CD3.
[0016] In certain instances, the CD3 binding site comprises a heavy chain variable domain comprising a CDR1 sequence of STYAMN (SEQ ID NO:72), a CDR2 sequence of RIRSKYNNYATYYADSVKD (SEQ ID NO:73) and a CDR3 sequence of HGNFGNSYVSWFAY (SEQ ID NO:74). In other instances, the CD3 binding site comprises a light chain variable domain comprising a CDR1 sequence of RSSTGAVTTSNYAN (SEQ ID NO:90), a CDR2 sequence of GTNKRAP (SEQ ID NO:91), and a CDR3 sequence of ALWYSNL (SEQ ID NO:92).
[0017] In certain instances, the CD3 binding site comprises a heavy chain variable domain of SEQ ID NO:64 and a variable light chain domain of SEQ ID NO:68; a heavy chain variable domain of SEQ ID NO:65 and a variable light chain domain of SEQ ID NO:69; a heavy chain variable domain of SEQ ID NO:66 and a variable light chain domain of SEQ ID NO:70; or a heavy chain variable domain of SEQ ID NO:67 and a variable light chain domain of SEQ ID NO:71.
[0018] In any of the above embodiments, the CD33 binding protein is a dimeric protein. In any of the above embodiments, the CD33 binding protein is multifunctional.
[0019] In certain instances, the multifunctional CD33 binding protein has bispecificity for CD33 and CD3, wherein the binding specificities are provided by heavy chain variable domain and light chain variable domains for CD33 and CD3 selected from the group consisting of SEQ ID NOs:2, 12, 65 and 69; SEQ ID NOs:3, 13, 65 and 69; SEQ ID NOs:4, 14, 65 and 69; SEQ ID NOs:5, 15, 65 and 69; SEQ ID NOs:1, 11, 64 and 68; SEQ ID NOs:2, 12, 64 and 68; SEQ ID NOs:2, 12, 66 and 70; SEQ ID NOs:4, 14, 66 and 70; SEQ ID NOs:5, 15, 66 and 70; SEQ ID NOs:3, 13, 64 and 68; SEQ ID NOs:3, 13, 67 and 71; SEQ ID NOs:4, 14, 64 and 68; SEQ ID NOs:5, 15, 64 and 68; SEQ ID NOs:7, 17, 64 and 68; SEQ ID NOs:6, 16, 64 and 68; SEQ ID NOs:6, 16, 67 and 71; SEQ ID NOs:8, 18, 64 and 68; SEQ ID NOs:9, 19, 64 and 68; SEQ ID NOs:9, 19, 67 and 71; and SEQ ID NOs:10, 20, 64 and 68.
[0020] In another aspect, provided herein are bispecific, antigen-binding tandem diabodies specific to human CD3 and human CD33. In some embodiments, the tandem diabodies comprise a first polypeptide and a second polypeptide, each polypeptide having at least four variable chain domains linked one after another, wherein each polypeptide comprises a variable heavy chain domain specific for human CD33; a variable light chain domain specific for human CD33; a variable heavy chain domain specific for human CD3, and a variable light chain domain specific for human CD3 and wherein in each polypeptide the four variable chain domains are linked with one after another by peptide linkers L1, L2 and L3 in the order of VL(CD3)-L1-VH(CD33)-L2-VL(CD33)-L3-VH(CD3); VH(CD3)-L1-VL(CD33)-L2-VH(CD33)-L3-VL(CD3); VL(CD33)-L1-VH(CD3)-L2-VL(CD3)-L3-VH(CD33); or VH(CD33)-L1-VL(CD3)-L2-VH(CD3)-L3-VL(CD33).
[0021] In some embodiments, the VL domain specific to human CD33 comprises a CDR1 consisting of the sequence selected from the group consisting of SEQ ID NOs:21-27, a CDR2 consisting of the sequence selected from the group consisting of SEQ ID NOs:28-34 and a CDR3 consisting of the sequence of the group consisting of SEQ ID NOs:35-41.
[0022] In some embodiments, the VH domain specific to human CD33 comprises a CDR1 consisting of the sequence selected from the group consisting of SEQ ID NOs:42-48, a CDR2 consisting of the sequence selected from the group consisting of SEQ ID NOs:49-55 and a CDR3 consisting of a sequences selected from the group consisting of SEQ ID NOs:56-63.
[0023] In some embodiments, the CDR1, CDR2 and CDR3 of the VL domain specific to human CD33 are sequences selected from the group consisting of SEQ ID NOs:21, 28 and 35; SEQ ID NOs:22, 29 and 36; SEQ ID NOs:23, 30 and 37; SEQ ID NOs:24, 31 and 38; SEQ ID NOs:25, 32 and 39; SEQ ID NOs:26, 33 and 40; and SEQ ID NOs:27, 34 and 41.
[0024] In some embodiments, the CDR1, CDR2 and CDR3 of the VH domain specific to human CD33 are sequences selected from the group consisting of SEQ ID NOs:42, 49 and 56; SEQ ID NOs:43, 50 and 57; SEQ ID NOs:43, 50 and 58; SEQ ID NOs:43, 50 and 59; SEQ ID NOs:43, 50 and 60; SEQ ID NOs:44, 51 and 61; SEQ ID NOs:45, 52 and 62; SEQ ID NOs:46, 53 and 63; SEQ ID NOs:47, 54 and 63; and SEQ ID NOs:48, 55 and 63.
[0025] In some embodiments, the VL and VH domains specific to CD33 are sequences selected from the group consisting of SEQ ID NO:1 and SEQ ID NO:11; SEQ ID NO:2 and SEQ ID NO:12; SEQ ID NO:3 and SEQ ID NO:13; SEQ ID NO:4 and SEQ ID NO:14; SEQ ID NO:5 and SEQ ID NO:15; SEQ ID NO:6 and SEQ ID NO:16; SEQ ID NO:7 and SEQ ID NO:17; SEQ ID NO:8 and SEQ ID NO:18; SEQ ID NO:9 and SEQ ID NO:19; and SEQ ID NO:10 and SEQ ID NO:20.
[0026] In some embodiments, the VH domain specific for human CD3 comprises a CDR1 sequence of STYAMN (SEQ ID NO:72), a CDR2 sequence of RIRSKYNNYATYYADSVKD (SEQ ID NO:73) and a CDR3 sequence of HGNFGNSYVSWFAY (SEQ ID NO:74) or HGNFGNSYVSYFAY (SEQ ID NO:75).
[0027] In some embodiments, the VL domain specific for human CD3 comprises a CDR1 sequence of RSSTGAVTTSNYAN (SEQ ID NO:90), a CDR2 sequence of GTNKRAP (SEQ ID NO:91), and a CDR3 sequence of ALWYSNL (SEQ ID NO:92).
[0028] In some embodiments, the VL and VH domains specific to CD3 are sequences selected from the group consisting of SEQ ID NO:64 and SEQ ID NO:68; SEQ ID NO:65 and SEQ ID NO:69; SEQ ID NO:66 and SEQ ID NO:70; and SEQ ID NO:67 and SEQ ID NO:71.
[0029] In some embodiments, each polypeptide comprises four variable chain domains selected from the group consisting of SEQ ID NOs:2, 12, 65 and 69; SEQ ID NOs:3, 13, 65 and 69; SEQ ID NOs:4, 14, 65 and 69; SEQ ID NOs:5, 15, 65 and 69; SEQ ID NOs:1, 11, 64 and 68; SEQ ID NOs:2, 12, 64 and 68; SEQ ID NOs:2, 12, 66 and 70; SEQ ID NOs:4, 14, 66 and 70; SEQ ID NOs:5, 15, 66 and 70; SEQ ID NOs:3, 13, 64 and 68; SEQ ID NOs:3, 13, 67 and 71; SEQ ID NOs:4, 14, 64 and 68; SEQ ID NOs:5, 15, 64 and 68; SEQ ID NOs:7, 17, 64 and 68; SEQ ID NOs:6, 16, 64 and 68; SEQ ID NOs:6, 16, 67 and 71; SEQ ID NOs:8, 18, 64 and 68; SEQ ID NOs:9, 19, 64 and 68; SEQ ID NOs:9, 19, 67 and 71; and SEQ ID NOs:10, 20, 64 and 68.
[0030] In some embodiments, linkers L1, L2 and L3 consist of about 12 or less amino acid residues. In certain instances, linkers L1, L2 and L3 are each independently GGSGGS (SEQ ID NO:95), GGSG (SEQ ID NO:96) or GGSGG (SEQ ID NO:97). In other instances, linkers L1 and L3 are GGSGGS (SEQ ID NO:95) and linker L2 is GGSG (SEQ ID NO:96) or GGSGG (SEQ ID NO:97).
[0031] In some embodiments, a bispecific tandem diabody has a sequence selected from the group consisting of SEQ ID NOs:98-121. In other embodiments, a bispecific tandem diabody is tandem diabody 01 (SEQ ID NO:98), 02 (SEQ ID NO:99), 03 (SEQ ID NO:100), 04 (SEQ ID NO:101), 05 (SEQ ID NO:102), 06 (SEQ ID NO:103), 07 (SEQ ID NO:104), 08 (SEQ ID NO:105), 09 (SEQ ID NO:106), 10 (SEQ ID NO:107), 11 (SEQ ID NO:108), 12 (SEQ ID NO:109), 13 (SEQ ID NO:110), 14 (SEQ ID NO:111), 15 (SEQ ID NO:112), 16 (SEQ ID NO:113), 17 (SEQ ID NO:114), 18 (SEQ ID NO:115), 19 (SEQ ID NO:116), 20 (SEQ ID NO:117), 21 (SEQ ID NO:118), 22 (SEQ ID NO:119), 23 (SEQ ID NO:120), or 24 (SEQ ID NO:121).
[0032] In some embodiments, the bispecific, antigen-binding tandem diabodies possess binding K.sub.D of 10 nM or less to CD33 on CD33.sup.+ tumor cells selected from HL-60, KG-1, and U-937.
[0033] In some embodiments, the bispecific, antigen-binding tandem diabodies specifically binds to an epitope of human CD33 which is within .sub.62DQEVQEETQ.sub.70 (SEQ ID NO:94) (amino acid residues 62-70 of SEQ ID NO:93) of human CD33.
[0034] In another aspect, provided herein are polynucleotides encoding a CD33 binding protein or bispecific, tandem diabody of any of the above embodiments. In another aspect, provided herein are vectors comprising the described polynucleotides. In another aspect, provided herein are host cells transformed with the described vectors.
[0035] In yet another aspect, provided herein are pharmaceutical compositions comprising a CD33 binding protein or bispecific, tandem diabody of any of the above embodiments and a pharmaceutically acceptable carrier.
[0036] In yet another aspect, provided herein methods of producing a CD33 binding protein or bispecific, tandem diabody of any of the above embodiments comprising introducing into a host cell a polynucleotide encoding a CD33 binding protein or bispecific, tandem diabody of any of the above embodiments, or a vector comprising the described polynucleotides, culturing the host cell under conditions whereby the CD33 binding protein or the bispecific tandem diabody is expressed, and purifying the expressed CD33 binding protein or the bispecific tandem diabody.
[0037] Also provided herein are methods for the treatment of a CD33.sup.+ cancer comprising the administration of a bispecific, tandem diabody of any of the above embodiments to an individual suffering from CD33.sup.+ cancer. In some embodiments, the CD33.sup.+ cancer is acute myeloid leukemia (AML), acute lymphoblastic leukemia (ALL), precursor B-cell lymphoblastic leukemia, myeloid sarcoma, multiple myeloma, acute lymphoma, acute lymphoblastic lymphoma or chronic myelomonocytic leukemia (CMML). In some embodiments, the CD33.sup.+ cancer is acute myeloid leukemia (AML). In some embodiments, the CD33.sup.+ cancer is multiple myeloma. In some embodiments, the CD33.sup.+ cancer is acute lymphoblastic leukemia (ALL).
[0038] Also provided herein are methods for the treatment of acute myeloid leukemia (AML) comprising the administration of a bispecific, tandem diabody of any of the above embodiments to an individual suffering from AML. In some embodiments, the AML is AML with Recurrent Genetic Abnormalities, AML with myelodysplasia-related changes, Therapy-related myeloid neoplasms, Myeloid sarcoma, Myeloid proliferations related to Down syndrome, Blastic plasmacytoid dendritic cell neoplasm, or AML not otherwise categorized. In some embodiments, the AML is AML-M0, AML-M1, AML-M2, AML-M3, AML-M4, AML-M5, AML-M6, or AML-M7. In further embodiments, the AML is newly diagnosed, relapsed, or refractory.
[0039] Also provided herein are methods for the treatment of myeloid dysplastic syndrome (MDS) comprising the administration of a bispecific, tandem diabody of any of the above embodiments to an individual suffering from MDS.
[0040] Also provided herein are methods for the treatment of myeloproliferative disease (MPD) comprising the administration of a bispecific, tandem diabody of any of the above embodiments to an individual suffering from MPD.
[0041] Also provided herein are methods for the treatment of chronic myelomonocytic leukemia (CMML) comprising the administration of a bispecific, tandem diabody of any of the above embodiments to an individual suffering from CMML.
[0042] Also provided herein are methods for the treatment of immune suppression by myeloid derived suppressor cells (MDSCs) comprising the administration of a bispecific, tandem diabody in any of the above embodiments to an individual suffering from immune suppression.
[0043] In the above methods for the treatment, in certain instances, the methods further comprise administering cytarabine, azacitidine, decitabine, an anthracycline (e.g., daunorubicin, idarubicin, doxorubicin, and the like), amsacrine, fludarabine, clofarabine, cladribine, nelarabine, methotrexate, bortezomib, carfilzomib, melphalan, ibrutinib, thalidomide, lenalidomide, pomalidomide, apremilast, an epipodophyllotoxin (e.g., etoposide, teniposide, and the like), an anthracenedione (e.g., mitoxantrone, pixantrone, losoxantrone, piroxantrone, ametantrone and the like), an anti-CD20 agent (e.g., rituximab, ocrelizumab, ofatumumab, and the like) or combinations thereof.
INCORPORATION BY REFERENCE
[0044] All publications, patents, and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication, patent, or patent application was specifically and individually indicated to be incorporated by reference.
BRIEF DESCRIPTION OF THE DRAWINGS
[0045] The novel features of the invention are set forth with particularity in the appended claims. A better understanding of the features and advantages of the present invention will be obtained by reference to the following detailed description that sets forth illustrative embodiments, in which the principles of the invention are utilized, and the accompanying drawings of which:
[0046] FIG. 1 Schematic representation of the gene organization and a domain order of CD3/CD33 tandem diabodies (TandAb.RTM.). Tandem diabodies are expressed as a single polypeptide comprised of four variable domains connected via short peptide linkers L1, L2 and L3. Following expression, two monomeric polypeptides associate non-covalently head-to-tail to form the functional homodimeric tandem diabody molecule. L1, L2, L3: Linker; VH: Heavy chain variable domain; VL: Light chain variable domain.
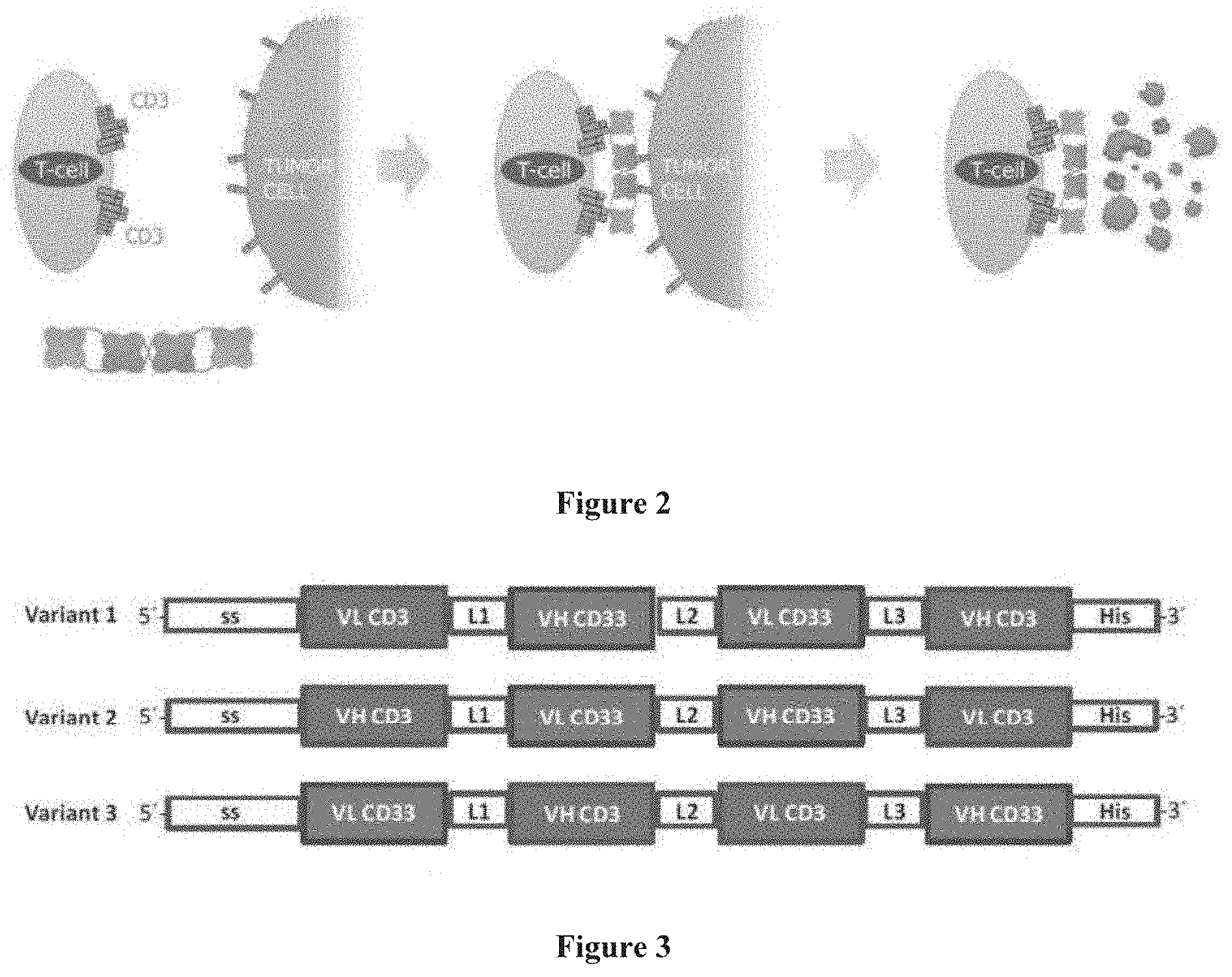
[0047] FIG. 2 CD3 engaging tandem diabody and its mode of action. Tandem diabodies are tetravalent bispecific proteins that can engage cytotoxic T-cells via binding to CD3. The tandem diabody binds to a CD33.sup.+ tumor cell with two of four binding domains and to CD3 with the other two binding domains. This T-cell/target cell binding (crosslinking) event promotes activation of the T-cell and promotes the subsequent destruction of the tumor cell via ADCC.
[0048] FIG. 3 Domain order variants of CD33/CD3 tandem diabodies. Variations of domain order of variable heavy (VH) and variable light (VL) chains within gene sequences encoding tandem diabodies allows production of antibodies with CD33 and CD3 specificities located on the inside or outside of the molecule. Domain specificities, location of signal sequences (ss) and linkers (L1, L2, L3) and affinity tags (His) as well as 5'- and 3'-ends are indicated.
[0049] FIG. 4 Comparison of positively enriched vs. negatively selected healthy donor T-cells. KG-1a cells were incubated with 10 pM (approx. 1 ng/mL) and 25 pM (approx. 2.5 ng/mL) of one of 10 selected tandem diabodies and either negatively selected healthy donor T-cells or positively selected healthy donor T-cells at an E:T cell ratio of 1:1 or 3:1, as indicated. After 48 hours, cell counts were determined and cytotoxicity was assessed with DAPI staining. Results are shown as mean.+-.SEM for the percentage of dead cells (upper panels) and the percentage of specific cytotoxicity (lower panels) from 3 independent experiments performed in duplicate wells.
[0050] FIG. 5 Analysis strategy. Scatter and histogram plots from one healthy donor T-cell aliquot and 1 representative AML cell line (HL-60) and primary AML specimen (AMP002) each illustrating the strategy pursued to determine tandem diabody-induced cytotoxicity. FSC, forward scatter; SSC, side scatter.
[0051] FIGS. 6A-6D Screening cytotoxicity assays in CD33+ AML, cell lines. Parental HL-60 (FIG. 6A, FIG. 6B) and KG-1a (FIG. 6C, FIG. 6D) cells were incubated with 10 pM (approx. 1 ng/mL) and 25 pM (approx. 2.5 ng/mL) of one of 22 CD33/CD3 tandem diabody molecules or a non-binding control tandem diabody (00) and healthy donor T-cells at an E:T cell ratio of either 1:1 (FIG. 6A, FIG. 6C) or 5:1 (FIG. 6B, FIG. 6D) as indicated. After 48 hours, cell counts were determined and cytotoxicity was assessed with DAPI staining to quantify drug-specific cytotoxicity. Results are shown as mean.+-.SEM for the percentage of DAPI cells from 3 independent experiments performed in duplicate wells. Qualitatively similar results were obtained when cytotoxicity was expressed as the percentage of specific cytotoxicity.
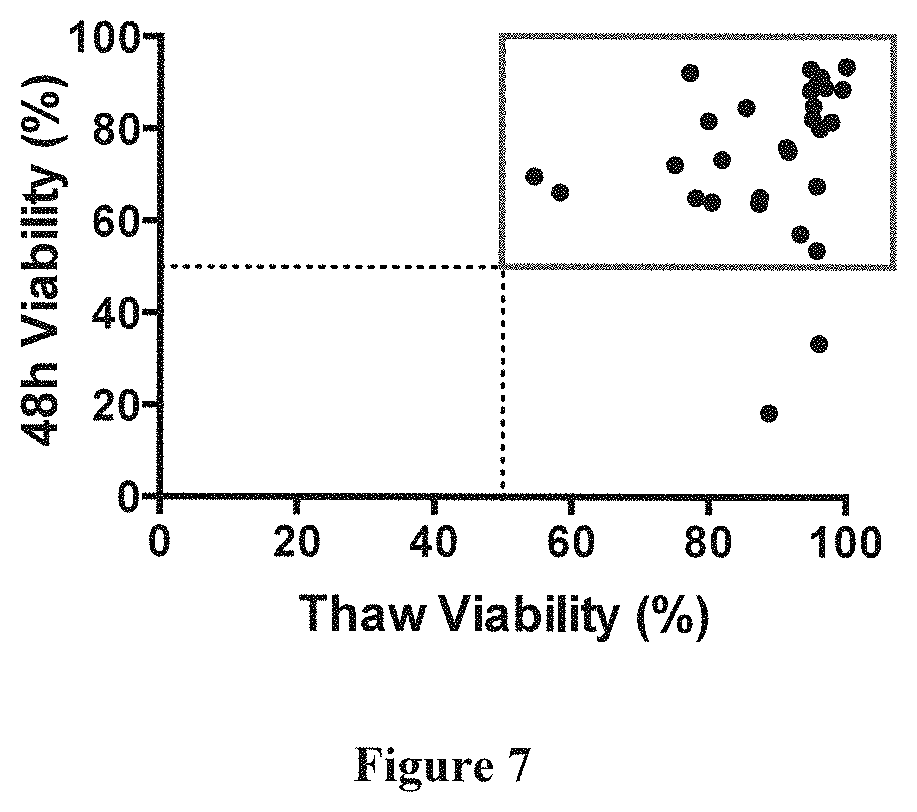
[0052] FIG. 7 Selection of primary AML specimens for study. Frozen aliquots from a total of primary human AML specimens were obtained for analysis. The percentage of AML blasts upon thaw was determined by flow cytometry based on CD45/side-scatter properties. Viability of the specimens was determined upon thaw as well after 48 hours in cytokine-containing liquid culture (without addition of tandem diabody molecules or healthy donor T-cells) via flow cytometry using DAPI as live/dead cell marker. Results for viability after thawing as well as after 48 hours are depicted for all specimens, which had >58% AML blasts. Square: Primary AML specimens that showed a viability of >50% at thaw as well as >50% after 48 hours in cytokine-containing liquid culture which were included in the final analyses.
[0053] FIGS. 8A-8C Tandem diabody-induced cytotoxicity in primary AML specimens. Primary AML specimens were incubated with 2.5 pM (approx. 250 pg/mL), 10 pM (approx. 1 ng/mL), and 25 pM (approx.2.5 ng/mL) of one of 9 tandem diabody molecules without healthy donor T-cells added (FIG. 8A) or with healthy donor T-cells at an E:T cell ratio of either 1:3 (FIG. 8B) or 1:1 (FIG. 8C) as indicated. After 48 hours, cell counts were determined and cytotoxicity was assessed with DAPI staining to quantify drug-specific cytotoxicity. Results are shown as mean.+-.SEM for the percentage of specific cytotoxicity from experiments performed in duplicate wells.
[0054] FIGS. 9A-9Y Amino acid sequences
[0055] (FIG. 9A) sequence of extracellular domain of human CD33 (aa 18-259) (SEQ ID NO: 93);
[0056] (FIG. 9B) complete sequence of tandem diabody 1 (SEQ ID NO:98);
[0057] (FIG. 9C) complete sequence of tandem diabody 2 (SEQ ID NO:99);
[0058] (FIG. 9D) complete sequence of tandem diabody 3 (SEQ ID NO:100);
[0059] (FIG. 9E) complete sequence of tandem diabody 4 (SEQ ID NO:101);
[0060] (FIG. 9F) complete sequence of tandem diabody 5 (SEQ ID NO:102);
[0061] (FIG. 9G) complete sequence of tandem diabody 6 (SEQ ID NO:103);
[0062] (FIG. 9H) complete sequence of tandem diabody 7 (SEQ ID NO:104);
[0063] (FIG. 9I) complete sequence of tandem diabody 8 (SEQ ID NO:105);
[0064] (FIG. 9J) complete sequence of tandem diabody 9 (SEQ ID NO:106);
[0065] (FIG. 9K) complete sequence of tandem diabody 10 (SEQ ID NO:107);
[0066] (FIG. 9L) complete sequence of tandem diabody 11 (SEQ ID NO:108);
[0067] (FIG. 9M) complete sequence of tandem diabody 12 (SEQ ID NO:109);
[0068] (FIG. 9N) complete sequence of tandem diabody 13 (SEQ ID NO:110);
[0069] (FIG. 9O) complete sequence of tandem diabody 14 (SEQ ID NO:111);
[0070] (FIG. 9P) complete sequence of tandem diabody 15 (SEQ ID NO:112);
[0071] (FIG. 9Q) complete sequence of tandem diabody 16 (SEQ ID NO:113);
[0072] (FIG. 9R) complete sequence of tandem diabody 17 (SEQ ID NO:114);
[0073] (FIG. 9S) complete sequence of tandem diabody 18 (SEQ ID NO:115);
[0074] (FIG. 9T) complete sequence of tandem diabody 19 (SEQ ID NO:116);
[0075] (FIG. 9U) complete sequence of tandem diabody 20 (SEQ ID NO:117);
[0076] (FIG. 9V) complete sequence of tandem diabody 21 (SEQ ID NO:118);
[0077] (FIG. 9W) complete sequence of tandem diabody 22 (SEQ ID NO:119);
[0078] (FIG. 9X) complete sequence of tandem diabody 23 (SEQ ID NO:120); and
[0079] (FIG. 9Y) complete sequence of tandem diabody 24 (SEQ ID NO:121). Underlined sequences represent linkers L1, L2 and L3.
[0080] FIG. 10 Effect of tandem diabodies 16 and 12 on the growth of HL-60 cells in NOD/scid mice. Eight experimental groups of immunodeficient NOD/scid mice were xenotransplanted by subcutaneous injection with a suspension of 4.times.10.sup.6 HL-60 cells on day 0. Prior to injection HL-60 cells were mixed with 3.times.10.sup.6 purified T-cells from healthy donors. All animals of the experimental groups transplanted with tumor cells and T-cells received an intravenous bolus on days 0, 1, 2, 3 and 4 of either vehicle (control) or tandem diabody 16 or 12 at three different dose levels as indicated (0.1 .mu.g, 1 .mu.g, and 10 .mu.g). One group without effector cells and vehicle treatment served as an additional negative control.
[0081] FIG. 11 Anti-tumor activity of tandem diabody 16 in an AML Xenograft Model. NOD/scid mice were sublethally irradiated (2 Gy) and subcutaneously inoculated with 4.times.10.sup.6 HL-60 cells. On day 9 the animals received a single bolus injection of anti-asialo GM1 rabbit Ab. When tumors reached a volume between 50-150 mm.sup.3 (mean 73.+-.11 mm.sup.3) on day 10 animals were allocated to 3 treatment groups. Groups 2 and 3 (n=8) were intraperitoneally injected with 1.5.times.10.sup.7 expanded and activated human T-cells. From day 13 to day 21 (qd.times.d9) animals received either tandem diabody 16 (Group 3) or vehicle into the lateral tail vein (Group 1 and Group 2).
[0082] FIGS. 12A-12B Relative amount (FIG. 12A) and absolute counts (FIG. 12B) of human AML blasts in the bone marrow (BM) and spleen of NSG mice at day 38 after treatment with 5 .mu.g (0.25 mg/kg) or 50 .mu.g (2.5 mg/kg) CD33/CD3 tandem diabody 12 and 16.
[0083] FIG. 13 Kinetics of CD33/CD3 tandem diabody 16-mediated target cell lysis. 1.times.10.sup.4 calcein-labeled HL-60 target cells were incubated with primary human T-cells as effector cells at an E:T ratio of 25:1 in the presence of serial dilutions of tandem diabody 16 or without antibody (w/o) for 30 min, 1 h, 2 h, 3 h, 4 h, or 5 h. At each time point, the fluorescent calcein released from lysed target cells was used to calculated specific lysis. Mean and SD of three replicates are plotted.
[0084] FIG. 14 Kinetics of EC.sub.50 and specific lysis values for CD33/CD3 tandem diabody 16. EC.sub.50 values (black solid circles) and tandem diabody 16-mediated target cell lysis (open squares) were determined in calcein-release cytotoxicity assays at the indicated incubation times by non-linear regression/sigmoidal dose-response and plotted.
[0085] FIG. 15 Cytotoxic activity in newly diagnosed, relapsed and refractory AML patient samples.
DETAILED DESCRIPTION OF THE INVENTION
[0086] According to a first aspect, described herein are binding proteins having specificity for at least CD33, preferably human CD33. In some embodiments, the CD33 binding proteins have specificity for human and cynomolgus CD33, i.e. are cross-reactive. In some embodiments, these cross-reactive binding proteins bind to human and cynomolgous CD33 with similar affinity.
[0087] CD33 is expressed on myeloid cells, for example, such as the blasts of acute myeloid leukemia (AML). For the isolation of antibody domains specific for CD33, such as human CD33, antibody libraries may be screened. For example IgM phage display libraries can be screened by employing, for example, a recombinant CD33-Fc fusion protein containing amino acids 1-243 of the extracellular domain of human CD33 (FIG. 9A, SEQ ID NO:93).
[0088] In some embodiments the CD33 binding protein has at least one CD33 binding site comprising a light chain variable domain and a heavy chain variable domain. The light chain variable domain comprises the light chain CDR1, CDR2 and CDR3 and the heavy chain variable domain comprises the heavy chain CDR1, CDR2 and CDR3. In some embodiments these light chain CDRs (CDR1, CDR2 and CDR3) are selected from the human CDR sequences shown in Table 1 (SEQ ID NOs:21-41). In certain instances, the light chain CDR1 is selected from SEQ ID NOs:21-27. In certain instances, the light chain CDR2 is selected from SEQ ID NOs:28-34. In certain instances, the light chain CDR3 is selected from SEQ ID NOs:35-41.
[0089] In some embodiments these heavy chain CDRs (heavy chain CDR1, CDR2 and CDR3) are selected from the human CDR sequences shown in Table 2 (SEQ ID NOs:42-63). In certain instances, the heavy chain CDR1 is selected from SEQ ID NOs:42-48. In certain instances, the heavy chain CDR2 is selected from SEQ ID NOs:49-55. In certain instances, the heavy chain CDR3 is selected from SEQ ID NOs:56-63.
[0090] In some embodiments, the light and heavy CDRs are selected without the surrounding framework sequences of the respective variable domains, which include framework sequences from other immunoglobulins or consensus framework regions, optionally are further mutated and/or replaced by other suitable framework sequences. Therefore provided herein in some embodiments, is a CD33 binding protein comprising a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:21; the light chain CDR2 is SEQ ID NO:28 and the light chain CDR3 is SEQ ID NO:35. In some embodiments, a CD33 binding protein comprises a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:22; the light chain CDR2 is SEQ ID NO:29 and the light chain CDR3 is SEQ ID NO:36. In some embodiments, a CD33 binding protein comprises a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:23; the light chain CDR2 is SEQ ID NO:30 and the light chain CDR3 is SEQ ID NO:37. In some embodiments, a CD33 binding protein comprises a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:24; the light chain CDR2 is SEQ ID NO:31 and the light chain CDR3 is SEQ ID NO:38. In some embodiments, a CD33 binding protein comprises a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:25; the light chain CDR2 is SEQ ID NO:32 and the light chain CDR3 is SEQ ID NO:39. In some embodiments, a CD33 binding protein comprises a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:26; the light chain CDR2 is SEQ ID NO:33 and the light chain CDR3 is SEQ ID NO:40. In some embodiments, a CD33 binding protein comprises a light chain variable domain, wherein the light chain CDR1 is SEQ ID NO:27; the light chain CDR2 is SEQ ID NO:34 and the light chain CDR3 is SEQ ID NO:41.
[0091] Also provided herein in some embodiments, is a CD33 binding protein comprising a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:42; the heavy chain CDR2 is SEQ ID NO:49 and the heavy chain CDR3 is SEQ ID NO:56. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:43; the heavy chain CDR2 is SEQ ID NO:50 and the heavy chain CDR3 is SEQ ID NO:57. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:43; the heavy chain CDR2 is SEQ ID NO:50 and the heavy chain CDR3 is SEQ ID NO:58. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:43; the heavy chain CDR2 is SEQ ID NO:50 and the heavy chain CDR3 is SEQ ID NO:59. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:43; the heavy chain CDR2 is SEQ ID NO:50 and the heavy chain CDR3 is SEQ ID NO:60. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:44; the heavy chain CDR2 is SEQ ID NO:51 and the heavy chain CDR3 is SEQ ID NO:61. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:45; the heavy chain CDR2 is SEQ ID NO:52 and the heavy chain CDR3 is SEQ ID NO:62. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:46; the heavy chain CDR2 is SEQ ID NO:53 and the heavy chain CDR3 is SEQ ID NO:63. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:47; the heavy chain CDR2 is SEQ ID NO:54 and the heavy chain CDR3 is SEQ ID NO:63. In some embodiments, a CD33 binding protein comprises a heavy chain variable domain, wherein the heavy chain CDR1 is SEQ ID NO:48; the heavy chain CDR2 is SEQ ID NO:55 and the heavy chain CDR3 is SEQ ID NO:63.
[0092] In further embodiments, a CD33 binding protein comprises a variable light chain domain selected from amino acid sequences SEQ ID NOs.:1-10 shown in Table 3. In further embodiments, a CD33 binding protein comprises a variable heavy chain domain selected from amino acid sequences SEQ ID NO:11-20 shown in Table 4. In yet further embodiments, a CD33 binding protein comprises a variable light chain domain selected from amino acid sequences SEQ ID NOs.:1-10 shown in Table 3 and a variable heavy chain domain selected from amino acid sequences SEQ ID NO:11-20 shown in Table 4.
[0093] The term "binding protein" refers to an immunoglobulin derivative with antigen binding properties, i.e. immunoglobulin polypeptides or fragments thereof that contain an antigen binding site. The binding protein comprises variable domains of an antibody or fragments thereof. Each antigen-binding domain is formed by an antibody, i.e. immunoglobulin, variable heavy chain domain (VH) and an antibody variable light chain domain (VL) binding to the same epitope, whereas the variable heavy chain domain (VH) comprises three heavy chain complementarity determining regions (CDR): CDR1, CDR2 and CDR3; and the variable light chain domain (VL) comprises three light chain complementarity determining regions (CDR): CDR1, CDR2 and CDR3. In some instances, the binding protein according to some embodiments herein is devoid of immunoglobulin constant domains. In some instances, the variable light and heavy chain domains forming the antigen binding site is covalently linked with one another, e.g. by a peptide linker, or in other instances, the variable light and heavy chain domains non-covalently associate with one another to form the antigen binding site. The term "binding protein" refers also to antibody fragments or antibody derivatives including, for example, Fab, Fab', F(ab').sub.2, Fv fragments, single-chain Fv, tandem single-chain Fv ((scFv).sub.2, Bi-specific T-cell engagers (BiTE.RTM.), dual affinity retargeting antibodies (DART.TM.), diabody and tandem diabody (TandAb.RTM.). Furthermore, in certain instances, the binding protein is multivalent, i.e. has two, three or more binding sites for CD33.
TABLE-US-00001 TABLE 1 Amino acid sequences of anti-CD33 variable light chain CDR1, CDR2 and CDR3 Sequence CDR identifier Light Chain CDR Sequence CDR1 SEQ ID NO: 21 GGNNIGSTTVH SEQ ID NO: 22 SGSRSNIGSNTVN SEQ ID NO: 23 SGSSSNIGSNTVN SEQ ID NO: 24 TGSSSNIGAGYDVH SEQ ID NO: 25 SGSSSNIGSNIVN SEQ ID NO: 26 SGSSSNIGSNTVK SEQ ID NO: 27 SGSSSNIGDNVVN CDR2 SEQ ID NO: 28 DDNERPS SEQ ID NO: 29 GNNQRPS SEQ ID NO: 30 SDNQRPS SEQ ID NO: 31 GNSNRPS SEQ ID NO: 32 SNNQRPS SEQ ID NO: 33 SNNQRSS SEQ ID NO: 34 STNKRPS CDR3 SEQ ID NO: 35 QVWDSGSDH SEQ ID NO: 36 ATWDDSLIG SEQ ID NO: 37 ATWDDSLNG SEQ ID NO: 38 QSYDSSLSD SEQ ID NO: 39 AAWDDSLKG SEQ ID NO: 40 AAWDDSLNG SEQ ID NO: 41 AAWDDSLSA
TABLE-US-00002 TABLE 2 Amino acid sequences of anti-CD33 variable heavy chain CDR1, CDR2 and CDR3 Sequence CDR identifier Heavy Chain CDR Sequence CDR1 SEQ ID NO: 42 SNYGIH SEQ ID NO: 43 TSYDIN SEQ ID NO: 44 TSYYMH SEQ ID NO: 45 TSYWIG SEQ ID NO: 46 SSYAIS SEQ ID NO: 47 SSYGIS SEQ ID NO: 48 DSYAIS CDR2 SEQ ID NO: 49 LISYDGNKKFYADSVKG SEQ ID NO: 50 WMNPNSGNTGFAQKFQG SEQ ID NO: 51 GIINPSGGSTSYAQKFQG SEQ ID NO: 52 HYPGDSDTRYSPSFQG SEQ ID NO: 53 GIYPIFGSANYAQKFQG SEQ ID NO: 54 GIIPIFGSAHYAQKFQG SEQ ID NO: 55 GIIPIFGSAHYSQKFQG CDR3 SEQ ID NO: 56 DRLESAAFDY SEQ ID NO: 57 DRANTDFSYGMDV SEQ ID NO: 58 DRAVTDYYYGMDV SEQ ID NO: 59 DRANTDYSFGMDV SEQ ID NO: 60 DRANTDYSLGMDV SEQ ID NO: 61 DVVPAAIDYYGMDV SEQ ID NO: 62 HKRGSDAFDI SEQ ID NO: 63 EYYYDSSEWAFDI
TABLE-US-00003 TABLE 3 Amino acid sequences of all anti-CD33 variable light chain domains (amino acid sequences of variable light chain CDR1 CDR2 and CDR3 are in bold and underlined) anti- CD33 Sequence clone identifier Variable light chain (VL) domain Sequence 01 SEQ ID NO: 1 SYELTQPPSVSVAPGQTAMITCGGNNIGSTTVHWYQQKPGQAPVLVV YDDNERPSGIPERFSGSNSGSTATLTINRVEAGDEADYYCQVWDSGSD HVVFGGGTKLTVL 02 SEQ ID NO: 2 QSVLTQPPSASGTPGQRVTISCSGSRSNIGSNTVNWYQQLPGTAPKLLI YGNNQRPSGVPDRFSGSKSGSSASLAISGLQSEDEADYYCATWDDSLI GWVFGGGTKLTVL 03 SEQ ID NO: 3 QSVLTQPPSASGTPGQRVTISCSGSRSNIGSNTVNWYQQLPGTAPKLLI YGNNQRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCATWDDSLI GWVFGGGTKLTVL 04 SEQ ID NO: 4 QSVLTQPPSASGTPGQRVTISCSGSRSNIGSNTVNWYQQLPGTAPKLLI YGNNQRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCATWDDSLI GWVFGGGTKLTVL 05 SEQ ID NO: 5 QSVLTQPPSASGTPGQRVTISCSGSRSNIGSNTVNWYQQLPGTAPKLLI YGNNQRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCATWDDSLI GWVFGGGTKLTVL 06 SEQ ID NO: 6 QSVLTQPPSASGTPGQRVTISCSGSSSNIGSNTVNWYQQLPGTAPKLLI YSDNQRPSGVPDRFSGSKSGSSASLAISGLQSDDEADYYCATWDDSLN GAVFGGGTKLTVL 07 SEQ ID NO: 7 QSVLTQPPSVSGAPGQRVTISCTGSSSNIGAGYDVHWYQQLPGTAPKL LIYGNSNRPSGVPDRFSGSKSGTSASLAITGLQAEDEADYYCQSYDSSL SDVVFGGGTKLTVL 08 SEQ ID NO: 8 QSVLTQPPSASGTPGQRVTISCSGSSSNIGSNIVNWYQQLPGTAPKLLIY SNNQRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCAAWDDSLKG YVFGGGTKLTVL 09 SEQ ID NO: 9 QSVLTQPPSASGTPGQRVTISCSGSSSNIGSNTVKWYQQLPGTAPKLLI YSNNQRSSGVPDRFSGSKSGSSASLAISGLQSEDEADYYCAAWDDSLN GYVFGGGTKLTVL 10 SEQ ID NO: 10 QSVLTQPPSASGTPGQRVTISCSGSSSNIGDNVVNWYQQLPGTAPKLLI YSTNKRPSGVPDRFSGSKSGSSASLAISGLQSEDEADYYCAAWDDSLS AYVFGGGTKLTVL
TABLE-US-00004 TABLE 4 Amino acid sequence of anti-CD33 variable heavy chain domain (amino acid sequences of variable heavy chain CDR1, CDR2 and CDR3 are in bold and underlined) anti- CD33 Sequence clone identifier Variable heavy chain (VH) domain Sequence 01 SEQ ID NO: 11 QVQLQESGGGVVQPGRSLRLSCAASGFSFSNYGIHWVRQAPGKGLEWVA LISYDGNKKFYADSVKGRFAISRDTSKNTVDLQMTSLRPEDTAVYYCAK DRLESAAFDYWGQGTLVTVSS 02 SEQ ID NO: 12 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYDINWVRQAPGQGLEWM GWMNPNSGNTGFAQKFQGRVTMTRDTSTSTVYMELSSLRSEDTAVYYC ARDRANTDFSYGMDVWGQGTLVTVSS 03 SEQ ID NO: 13 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYDINWVRQAPGQGLEWM GWMNPNSGNTGFAQKFQGRVTMTRDTSTSTVYMELSSLRSEDTAVYYC ARDRAVTDYYYGMDVWGQGTLVTVSS 04 SEQ ID NO: 14 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYDINWVRQAPGQGLEWM GWMNPNSGNTGFAQKFQGRVTMTRDTSTSTVYMELSSLRSEDTAVYYC ARDRANTDYSFGMDVWGQGTLVTVSS 05 SEQ ID NO: 15 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYDINWVRQAPGQGLEWM GWMNPNSGNTGFAQKFQGRVTMTRDTSTSTVYMELSSLRSEDTAVYYC ARDRANTDYSLGMDVWGQGTLVTVSS 06 SEQ ID NO: 16 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYYMHWVRQAPGQGLEW MGIINPSGGSTSYAQKFQGRVTMTRDTSTSTVYMELSSLRSEDTAVYYC ARDVVPAAIDYYGMDVWGQGTTVTVSS 07 SEQ ID NO: 17 QVQLVQSGAEVKKPGESLKISCKGSGYSFTSYWIGWVRQMPGKGLEWM GIIYPGDSDTRYSPSFQGQVTISADKSISTAYLQWSSLKASDTAMYYCAR HKRGSDAFDIWGQGTTVTVSS 08 SEQ ID NO: 18 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYAISWVRQAPGQGLEWMG GIYPIFGSANYAQKFQGRVTITADESTSTAYMELSSLRSEDTAVYYCARE YYYDSSEWAFDIWGQGTLVTVSS 09 SEQ ID NO: 19 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYGISWVRQAPGQGLEWM GGIIPIFGSAHYAQKFQGRVTITADESTSTAYMELSSLRSEDTAVYYCAR EYYYDSSEWAFDIWGQGTLVTVSS 10 SEQ ID NO: 20 QVQLVQSGAEVKKPGSSVKVSCKASGGTFDSYAISWVRQAPGQGLEWM GGIIPIFGSAHYSQKFQGRVTITADESTSTAYMELSSLRSEDTAVYYCAR EYYYDSSEWAFDIWGQGTLVTVSS
[0094] In some embodiments, a binding protein conferring specificity to CD33 is selected from one of the following combinations of a variable heavy chain domain and a variable light chain domain forming the human CD33 binding site shown in Table 3 and in Table 4. Non-limiting examples include (i) SEQ ID NO:1 and SEQ ID NO:11, (ii) SEQ ID NO:2 and SEQ ID NO:12, (iii) SEQ ID NO:3 and SEQ ID NO:13, (iv) SEQ ID NO:4 and SEQ ID NO:14, (v) SEQ ID NO:5 and SEQ ID NO:15, (vi) SEQ ID NO:6 and SEQ ID NO:16, (vii) SEQ ID NO:7 and SEQ ID NO:17, (viii) SEQ ID NO:8 and SEQ ID NO:18, (ix) SEQ ID NO:9 and SEQ ID NO:19, and (x) SEQ ID NO:10 and SEQ ID NO:20.
[0095] Also described herein are binding proteins that not only have specificity for CD33, but which also have at least one further functional domain. In a further embodiment at least one further functional domain is an effector domain. An "effector domain" comprises a binding site of an antibody specific for an effector cell, which can stimulate or trigger cytotoxicity, phagocytosis, antigen presentation, cytokine release. Such effector cells are, for example, but not limited to, T-cells. In particular, the effector domain comprises at least one antibody variable heavy chain domain and at least one variable light chain domain forming an antigen binding site for an antigen on T-cells, such as, for example, human CD3.
[0096] Thus, in some embodiments, the CD33 binding protein is multifunctional. The term multifunctional as used herein means that a binding protein exhibits two or more different biological functions. For example, the different biological functions are different specificities for different antigens. In certain instances, the multifunctional CD33 binding protein is multispecific, i.e. has binding specificity to CD33 and one or more further antigens. In certain instances, the binding protein is bispecific with specificities for CD33 and CD3. Such bispecific binding proteins include, for example, bispecific monoclonal antibodies of the classes IgA, IgD, IgE, IgG or IgM, diabodies, single-chain diabodies (scDb), tandem single chain Fv (scFv)2, for example Bi-specific T-cell engagers (BiTE.RTM.), dual affinity retargeting antibodies (DART.TM.), tandem diabodies (TandAb.RTM.), and flexibodies.
[0097] In certain embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has specificity for human CD3 and, in some instances, cynomolgus CD3. Examples of such a binding site are polypeptides comprising the VH domain CDR1, CDR2 and CDR3 from the sequences shown in Table 5 (SEQ ID NOs:64-67) and VL domain CDR1, CDR2 and CDR3 from the sequence shown in Table 6 (SEQ ID NOs:68-71). In certain instances, a CD3 binding site is the combination of the variable heavy chain domain of SEQ ID NO:64 and the variable light chain domain of SEQ ID NO:68. In certain instances, a CD3 binding site is the combination of the variable heavy chain domain of SEQ ID NO:65 and the variable light chain domain of SEQ ID NO:69. In certain instances, a CD3 binding site is the combination of the variable heavy chain domain of SEQ ID NO:66 and the variable light chain domain of SEQ ID NO:70. In certain instances, a CD3 binding site is the combination of the variable heavy chain domain of SEQ ID NO:67 and the variable light chain domain of SEQ ID NO:71.
TABLE-US-00005 TABLE 5 Amino acid sequence of an anti-CD3 variable heavy chain domain (amino acid sequences of variable heavy chain CDR1, CDR2 and CDR3 are in bold and underlined) anti-CD3 VH domain Sequence SEQ ID NO: 64 EVQLVESGGGLVQPGGSLRLSCAASGFTFSTYAMNWVRQAPGKGLEWVGRIRSKYNNY CD3-01 ATYYADSVKDRFTISRDDSKNSLYLQMNSLKTEDTAVYYCARHGNFGNSWSYFAYWG QGTLVTVSS SEQ ID NO: 65 EVQLVESGGGLVQPGGSLRLSCAASGFTFSTYAMNWVRQAPGKGLEWVGRIRSKYNNY CD3-02 ATYYADSVKDRFTISRDDSKNSLYLQMNSLKTEDTAVYYCARHGNFGNSWSWFAYWG QGTLVTVSS SEQ ID NO: 66 EVQLVESGGGLVQPGGSLRLSCAASGFTFSTYAMNWVRQAPGKGLEWVGRIRSKYNNY CD3-03 ATYYADSVKDRFTISRDDSKNSLYLQMNSLKTEDTAVYYCARHGNFGNSWSWFAYWG QGTLVTVSS SEQ ID NO: 67 EVQLVESGGGLVQPGGSLRLSCAASGFTFSTYAMNWVRQAPGKGLEWVGRIRSKYNNY CD3-04 ATYYADSVKDRFTISRDDSKNSLYLQMNSLKTEDTAVYYCARHGNFGNSWSWFAYWG QGTLVTVSS
TABLE-US-00006 TABLE 6 Amino acid sequence of an anti-CD3 variable light chain domain (amino acid sequences of variable light chain CDR1, CDR2 and CDR3 are in bold and underlined) anti-CD3 VL domain Sequence SEQ ID NO:68 DIQMTQSPSSLSASVGDRVTITCRSSTGAVTTSNYANWVQQKPGKAPKALIGGTNKRAP CD3-01 GVPSRFSGSLIGDKATLTISSLQPEDFATYYCALWYSNLWVFGQGTKVEIK SEQ ID NO:69 DIQMTQSPSSLSASVGDRVTITCRSSTGAVTTSNYANWVQQKPGKAPKGLIGGTNKRAP CD3-02 GVPARFSGSGSGTDFTLTISSLQPEDFATYYCALWYSNLWVFGQGTKVEIK SEQ ID NO:70 DIQMTQSPSSLSASVGDRVTITCRSSTGAVTTSNYANWVQQKPGKAPKGLIGGTNKRAP CD3-03 GVPSRFSGSLIGDKATLTISSLQPEDFATYYCALWYSNLWVFGQGTKVEIK SEQ ID NO:71 DIQMTQSPSSLSASVGDRVTITCRSSTGAVTTSNYANWVQQKPGKAPKGLIGGTNKRAP CD3-04 GVPSRFSGSLIGTDFTLTISSLQPEDFATYYCALWYSNLWVFGQGTKVEIK
[0098] In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR1 sequence of STYAMN (SEQ ID NO:72). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR2 sequence of RIRSKYNNYATYYADSVKD (SEQ ID NO:73). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR3 sequence of HGNFGNSYVSWFAY (SEQ ID NO:74). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR3 sequence of HGNFGNSYVSYFAY (SEQ ID NO:75). In yet further embodiments, the CD3 binding site has a variable heavy chain domain comprising a CDR1, CDR2 and CDR3 sequence of SEQ ID NOs:72-74 respectively. In yet further embodiments, the CD3 binding site has a variable heavy chain domain comprising a CDR1, CDR2 and CDR3 sequence of SEQ ID NOs:72, 73 and 75 respectively.
[0099] In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR1 sequence selected from the group consisting of NTYAMN (SEQ ID NO:76), NTYAMH (SEQ ID NO:77) and NKYAMN (SEQ ID NO:78). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR2 sequence selected from the group consisting of RIRNKYNNYATYYADSVKD (SEQ ID NO:79), RIRNKYNNYATEYADSVKD (SEQ ID NO:80), RIRSKYNNYATEYAASVKD (SEQ ID NO:81), RIRNKYNNYATEYAASVKD (SEQ ID NO:82), RIRSKYNNYATYYADSVKG (SEQ ID NO:83) and RIRSKYNNYATEYADSVKS (SEQ ID NO:84). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable heavy chain domain comprising a CDR3 sequence selected from the group consisting of HGNFGDSYVSWFAY (SEQ ID NO:85), HGNFGNTYVSWFAY (SEQ ID NO:86), HGNFGCSYVSWFAY (SEQ ID NO:87), HGNFGNSYISYWAY (SEQ ID NO:88) and HGNFGNSYVSFFAY (SEQ ID NO:89).
[0100] . In yet further embodiments, the CD3 binding site has a variable heavy chain domain comprising a CDR1, CDR2 and CDR3 sequence of SEQ ID NOs:76, 73 and 74 respectively, SEQ ID NOs:76, 79 and 74 respectively, SEQ ID NOs:76, 80 and 74 respectively, SEQ ID NOs:76, 81 and 74 respectively, SEQ ID NOs:76, 82 and 74 respectively, SEQ ID NOs:76, 83 and 74 respectively, SEQ ID NOs:72, 83 and 74 respectively, SEQ ID NOs:72, 83 and 85 respectively, SEQ ID NOs:76, 83 and 86 respectively, SEQ ID NOs:77, 83 and 74 respectively, SEQ ID NOs:72, 83 and 87 respectively, SEQ ID NOs:78, 73 and 88 respectively or SEQ ID NOs:78, 84 and 89 respectively.
[0101] In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable light chain domain comprising a CDR1 sequence of RSSTGAVTTSNYAN (SEQ ID NO:90). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable light chain domain comprising a CDR2 sequence of GTNKRAP (SEQ ID NO:91). In further embodiments, the CD3 binding site of a bispecific CD33 and CD3 binding protein has a variable light chain domain comprising a CDR3 sequence of ALWYSNL (SEQ ID NO:92). In yet further embodiments, the CD3 binding site has a variable light chain domain comprising a CDR1, CD2 and CD3 sequence of SEQ ID NOs:90-92 respectively.
[0102] In certain instances, the CD3 binding site has a high affinity to CD3. Alternatively, in other instances, the CDR1, CDR2, CDR3 from the heavy-chain domain as well as the light-chain domain or, optionally, the variable light-chain domains and variable heavy-chain domains is derived from other CD3 antibodies, such as, for example UCHT1, muromonab-CD3 (OKT3), otelixizumab (TRX4), teplizumab (MGA031), visilizumab (Nuvion), and the like.
[0103] In another aspect, described herein are CD33 binding proteins as well as the bispecific CD33 and CD3 binding proteins that are humanized or fully human, i.e. of human origin.
[0104] In some embodiments, a bispecific CD33 and CD3 binding protein has one of the following combinations providing CD33 and CD3 specificity by variable light and heavy chain domains for CD33 and CD3: include, but are not limited to, (i) SEQ ID NOs:2, 12, 65 and 69, (ii) SEQ ID NOs:3, 13, 65 and 69, (iii) SEQ ID NOs:4, 14, 65 and 69, (iv) SEQ ID NOs:5, 15, 65 and 69, (v) SEQ ID NOs:1, 11, 64 and 68, (vi) SEQ ID NOs:2, 12, 64 and 68, (vii) SEQ ID NOs:2, 12, 66 and 70, (viii) SEQ ID NOs:4, 14, 66 and 70, (ix) SEQ ID NOs:5, 15, 66 and 70, and (x) SEQ ID NOs:3, 13, 64 and 68, (xi) SEQ ID NOs:3, 13, 67 and 71, (xii) SEQ ID NOs:4, 14, 64 and 68, (xiii) SEQ ID NOs:5, 15, 64 and 68, (xiv) SEQ ID NOs:7, 17, 64 and 68, (xv) SEQ ID NOs:6, 16, 64 and 68, (xvi) SEQ ID NOs:6, 16, 67 and 71, (xvii) SEQ ID NOs:8, 18, 64 and 68, (xviii) SEQ ID NOs:9, 19, 64 and 68; (xix) SEQ ID NOs:9, 19, 67 and 71, and (xx) SEQ ID NOs:10, 20, 64 and 68.
Conserved Variants of CDR Sequences and Heavy and Light Chain Domains
[0105] In alternative embodiments, the heavy and light chain domains incorporate immunologically active homologues or variants of the CDR sequences described herein. Accordingly in some embodiments, a CDR sequence in a heavy or light chain domain that binds to CD33 or CD3 is similar to, but not identical to, the amino acid sequence depicted in SEQ ID NOs: 21-63 or 72-92. In certain instances, a CDR variant sequence has a sequence identity of 99%, 98%, 97%, 96%, 95%, 94%, 93%, 92%, 91%, 90%, 89%, 88%, 87%, 86%, 85%, 84%, 83%, 82%, 81%, or 80% compared to the sequence of SEQ ID NOs: 21-63 or 72-90 and which is immunologically active.
[0106] In further instances, a CDR variant sequence incorporates 1, 2, 3, 4, or 5 conserved amino acid substitutions. Conservative substitutions include amino acid substitutions that substitute a given amino acid with another amino acid of similar characteristics and further include, among the aliphatic amino acids interchange of alanine, valine, leucine, and isoleucine; interchange of the hydroxyl residues serine and threonine, exchange of the acidic residues aspartate and glutamate, substitution between the amide residues asparagine and glutamine, exchange of the basic residues lysine and arginine, and replacements among the aromatic residues phenylalanine and tyrosine.
[0107] In yet further instances, a CDR variant sequence incorporates substitutions that enhance properties of the CDR such as increase in stability, resistance to proteases and/or binding affinities to CD33 or CD3.
[0108] In other instances, a CDR variant sequence is modified to change non-critical residues or residues in non-critical regions. Amino acids that are not critical can be identified by known methods, such as affinity maturation, CDR walking, site-directed mutagenesis, crystallization, nuclear magnetic resonance, photoaffinity labeling, or alanine-scanning mutagenesis.
[0109] In further alternative embodiments, the CD33 and CD3 binding proteins comprise heavy and light chain domains that are immunologically active homologues or variants of heavy and light chain domain sequences provided herein. Accordingly, in some embodiments, a CD33 and CD3 binding protein comprises a heavy or light chain domain sequence that is similar to, but not identical to, the amino acid sequence depicted in SEQ ID NOs:1-20 or 64-71. In certain instances, a variant heavy or light chain domain sequence has a sequence identity of 99%, 98%, 97%, 96%, 95%, 94%, 93%, 92%, 91%, 90%, 89%, 88%, 8'7%, 86%, 85%, 84%, 83%, 82%, 81%, or 80% compared to the sequence of SEQ ID NOs:1-20 or 64-71 and which is immunologically active.
[0110] In further instances, a variant heavy or light chain domain sequence incorporates 1, 2, 3, 4, or 5 conserved amino acid substitutions. Conservative substitutions include amino acid substitutions that substitute a given amino acid with another amino acid of similar characteristics and further include, among the aliphatic amino acids interchange of alanine, valine, leucine, and isoleucine; interchange of the hydroxyl residues serine and threonine, exchange of the acidic residues aspartate and glutamate, substitution between the amide residues asparagine and glutamine, exchange of the basic residues lysine and arginine, and replacements among the aromatic residues phenylalanine and tyrosine.
[0111] In yet further instances, a variant heavy or light chain domain sequence incorporates substitutions that enhance properties of the CDR such as increase in stability, resistance to proteases and/or binding affinities to CD33 or CD3.
[0112] In other instances, a variant heavy or light chain domain sequence is modified to change non-critical residues or residues in non-critical regions. Amino acids that are not critical can be identified by known methods, such as affinity maturation, CDR walking, site-directed mutagenesis, crystallization, nuclear magnetic resonance, photoaffinity labeling, or alanine-scanning mutagenesis.
CD33 and CD3 Bispecific and Tandem Diabodies
[0113] In another aspect, a CD33 binding protein or the bispecific CD33 and CD3 binding protein is a dimer, i.e. comprises two polypeptides with antigen binding sites for CD33 and CD3.
[0114] Also provided herein in another aspect, is a dimeric and bispecific CD33 and CD3 binding protein in the format of a tandem diabody (TandAb.RTM.). Such tandem diabodies are constructed by linking four antibody variable binding domains (two heavy-chain variable domains (VH) and two light-chain variable domains (VL) in a single gene construct (FIG. 1) enabling homo-dimerization. In such tandem diabodies the linker length is such that it prevents intramolecular pairing of the variable domains so that the molecule cannot fold back upon itself to form a single-chain diabody, but rather is forced to pair with the complementary domains of another chain. The domains are also arranged such that the corresponding VH and VL domains pair during this dimerization. Following expression from a single gene construct, two identical polypeptide chains fold head-to-tail forming a functional non-covalent homodimer of approximately 105 kDa (FIG. 1). Despite the absence of intermolecular covalent bonds, the homodimer is highly stable once formed, remains intact and does not revert back to the monomeric form.
[0115] Tandem diabodies have a number of properties that provide advantages over traditional monoclonal antibodies and other smaller bispecific molecules. Tandem diabodies contain only antibody variable domains and therefore are contemplated to lack side effects or non-specific interactions that may be associated with an Fc moiety. For example, Fc receptors which can bind to Fc domains are found on numerous cell types such as white blood cells (e.g., basophils, B-cells, eosinophils, natural killer cells, neutrophils and the like) or Kuppfer cells. Because tandem diabodies allow for bivalent binding to each of CD33 and CD3, the avidity is the same as that of an IgG. The size of a tandem diabody, at approximately 105 kDa, is smaller than that of an IgG, which may allow for enhanced tumor penetration. However, this size is well above the renal threshold for first-pass clearance, offering a pharmacokinetic advantage compared with smaller bispecific formats based on antibody-binding domains or non-antibody scaffolds. Moreover tandem diabodies are advantageous over other bispecific binding proteins such as BiTE or DART molecules based on this pharmacokinetic and avidity properties resulting in longer intrinsic half-lives and rapid cytotoxicity. Tandem diabodies are well expressed in host cells, for example, mammalian CHO cells. It is contemplated that robust upstream and downstream manufacturing process is available for tandem diabodies.
[0116] The CD33 and CD3 bispecific tandem diabodies described herein are designed to allow specific targeting of CD33.sup.+ tumor cells by recruiting cytotoxic T-cells. This improves ADCC (antibody dependent cell-mediated cytotoxicity) as compared to full length antibodies directed to a sole antigen and are not capable of directly recruiting cytotoxic T-cells. In contrast, by engaging CD3 molecules expressed specifically on these cells, the tandem diabody can crosslink cytotoxic T-cells with CD33.sup.+ tumor cells in a highly specific fashion, thereby significantly increasing the cytotoxic potential of such molecules. This mechanism is outlined in FIG. 2. The tandem diabody displays strong, specific and efficient ADCC. It is reported that T-cells can play a role in controlling tumor growth. For example, the presence of cytotoxic T-cells in colorectal tumors as well as lymph nodes from NHL patients was shown to correlate with a better clinical outcome. Furthermore, the potential of therapies designed to induce T-cell responses has been demonstrated for melanoma vaccines, as well as the antibody directed against CTLA-4, a negative regulator of T-cell activation. The tandem diabodies described herein engage cytotoxic T-cells via binding to the surface-expressed CD3, which forms part of the T-cell receptor. Simultaneous binding of this tandem diabody to CD3 and to CD33 expressed on the surface of particular tumor cells causes T-cell activation and mediates the subsequent lysis of the tumor cell (FIG. 2).
[0117] Therefore, in a further aspect is a multispecific, tandem diabody. In some embodiments, a multispecific tandem diabody has specificities to two, three or more different epitopes, wherein two or more epitopes can be of the same antigen target or of different antigen targets. In certain embodiments the multispecific, tandem diabody is bispecific and tetravalent, i.e. comprises four antigen-binding sites. Such a bispecific tandem diabody binds with at least one antigen-binding site, to human CD3 and to human CD33, wherein in certain instances, the tandem diabody binds with two antigen-binding sites to human CD3 and with two other antigen-binding sites to human CD33, i.e. the tandem diabody binds bivalently to each antigen.
[0118] In some embodiments, a bispecific, antigen-binding tandem diabody is specific to human CD33 and human CD3, wherein said tandem diabody comprises a first polypeptide and a second polypeptide, each polypeptide having at least four variable chain domains linked one after another, wherein each polypeptide comprises a variable heavy chain (VH) domain specific to human CD33;
[0119] (ii) a variable light chain (VL) domain specific to human CD33;
[0120] (iii) a VH domain specific for human CD3, and
[0121] (iv) a VL domain specific for human CD3.
[0122] In particular embodiments, a bispecific tandem diabody specifically binds to an epitope of human CD33 which is within .sub.62DQEVQEETQ.sub.70 (SEQ ID NO:94) (amino acid residues 62-70 of SEQ ID NO:93) of human CD33. In particular instances, such a tandem diabody comprises a first polypeptide and a second polypeptide, each polypeptide having at least four variable chain domains linked one after another, wherein each polypeptide comprises [0123] (i) a variable heavy chain domain specific to an epitope of human CD33 which is within .sub.62DQEVQEETQ.sub.70 (SEQ ID NO:94) (amino acid residues 62-70 of SEQ ID NO:93) of human CD33; [0124] (ii) a variable light chain domain specific to an epitope of human CD33 which is within .sub.62DQEVQEETQ.sub.70 (SEQ ID NO:94) (amino acid residues 62-70 of SEQ ID NO:93) of human CD33; [0125] (iii) a variable heavy chain domain specific for human CD3, and [0126] (iv) a variable light chain domain specific for human CD3.
[0127] In other embodiments, described herein are CD33/CD3 tandem diabodies that have an affinity to CD33 on CD33.sup.+ cells with a K.sub.D of 10 nM or less, 5 nM or less, 1 nM or less, or 0.5 nM or less. The CD33.sup.+ cells can be selected from tumor cells such as, for example, HL-60 or KG-1.
[0128] In a further embodiment a CD33/CD3 tandem diabody described herein binds CD3 and in certain instances, the epsilon chain of CD3 on CD3.sup.+ cells, particularly T-cells, with a K.sub.D of 10 nM or less, 5 nM or less or 2 nM or less.
[0129] In some embodiments, each polypeptide of a bispecific tandem diabody comprises one of the following combinations of the four variable chain domains: (i) SEQ ID NOs:2, 12, 65 and 69, (ii) SEQ ID NOs:3, 13, 65 and 69, (iii) SEQ ID NOs:4, 14, 65 and 69, (iv) SEQ ID NOs:5, 15, 65 and 69, (v) SEQ ID NOs:1, 11, 64 and 68, (vi) SEQ ID NOs:2, 12, 64 and 68, (vii) SEQ ID NOs:2, 12, 66 and 70, (viii) SEQ ID NOs:4, 14, 66 and 70, (ix) SEQ ID NOs:5, 15, 66 and 70, and (x) SEQ ID NOs:3, 13, 64 and 68, (xi) SEQ ID NOs:3, 13, 67 and 71, (xii) SEQ ID NOs:4, 14, 64 and 68, (xiii) SEQ ID NOs:5, 15, 64 and 68, (xiv) SEQ ID NOs:7, 17, 64 and 68, (xv) SEQ ID NOs:6, 16, 64 and 68, (xvi) SEQ ID NOs:6, 16, 67 and 71, (xvii) SEQ ID NOs:8, 18, 64 and 68, (xviii) SEQ ID NOs:9, 19, 64 and 68; (xix) SEQ ID NOs:9, 19, 67 and 71, and (xx) SEQ ID NOs:10, 20, 64 and 68.
[0130] As used herein, "dimer" refers to a complex of two polypeptides. In certain embodiments, the two polypeptides are non-covalently associated with each other, in particular with the proviso that there is no covalent bond between the two polypeptides. In certain instances, the two polypeptides have covalent associations such as disulfide bonds that form to aid in stabilization of the dimer. In certain embodiments, the dimer is homodimeric, i.e. comprises two identical polypeptides. The term "polypeptide" refers to a polymer of amino acid residues linked by amide bonds. The polypeptide is, in certain instances, a single chain fusion protein, which is not branched. In the polypeptide the variable antibody domains are linked one after another. The polypeptide, in other instances, may have contiguous amino acid residues in addition to the variable domain N-terminal and/or C-terminal residues. For example, such contiguous amino acid residues may comprise a Tag sequence, in some instances at the C-terminus, which is contemplated to be useful for the purification and detection of the polypeptide.
[0131] In one aspect, each polypeptide of the bispecific tandem diabody comprises four variable domains, a variable light chain (VL) and a variable heavy chain (VH) of a CD3 binding protein as well as a variable light chain (VL) and a variable heavy chain (VH) of a CD33 binding protein. In certain embodiments, four variable domains are linked by peptide linkers L1, L2 and L3 and in some instances arranged from the N- to the C-terminus as follows:
Domain Order:
(1) VL(CD3)-L1-VH(CD33)-L2-VL(CD33)-L3-VH(CD3); or
(2) VH(CD3)-L1-VL(CD33)-L2-VH(CD33)-L3-VL(CD3); or
(3) VL(CD33)-L1-VH(CD3)-L2-VL(CD3)-L3-VH(CD33); or
(4) VH(CD33)-L1-VL(CD3)-L2-VH(CD3)-L3-VL(CD33).
[0132] The length of the linkers influences the flexibility of the antigen-binding tandem diabody according to reported studies. Accordingly, in some embodiments, the length of the peptide linkers L1, L2 and L3 is such that the domains of one polypeptide can associate intermolecularly with the domains of another polypeptide to form the dimeric antigen-binding tandem diabody. In certain embodiments, such linkers are "short", i.e. consist of 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 or 12 amino acid residues. Thus, in certain instances, the linkers consist of about 12 or less amino acid residues. In the case of 0 amino acid residues, the linker is a peptide bond. Such short linkers favor the intermolecular dimerization of the two polypeptides by binding and forming correct antigen-binding sites between antibody variable light chain domains and antibody variable heavy chain domains of different polypeptides. Shortening the linker to about 12 or less amino acid residues generally prevents adjacent domains of the same polypeptide chain from intramolecular interaction with each other. In some embodiments, these linkers consist of about 3 to about 10, for example 4, 5 or 6 contiguous amino acid residues.
[0133] Regarding the amino acid composition of the linkers, peptides are selected that do not interfere with the dimerization of the two polypeptides. For example, linkers comprising glycine and serine residues generally provide protease resistance. The amino acid sequence of the linkers can be optimized, for example, by phage-display methods to improve the antigen binding and production yield of the antigen-binding polypeptide dimer. Examples of peptide linkers suitable for a tandem diabody in some embodiments are GGSGGS (SEQ ID NO:95), GGSG (SEQ ID NO:96), or GGSGG (SEQ ID NO:97).
[0134] Non-limiting examples of tandem diabodies as described herein are tandem diabodies having an anti-CD33 VL and VH domain, an anti-CD3 VL and VH domain, domain order and linker according to Table 7.
TABLE-US-00007 TABLE 7 Exemplary CD33/CD3 Tandem Diabodies (TandAbs) Anti-CD33 Anti-CD3 Tandem domain domain Domain Linker Diabody VL VH VH VL Order L1/L3 L2 01 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 2 NO: 12 NO: 65 NO: 69 02 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 3 NO: 13 NO: 65 NO: 69 03 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 4 NO: 14 NO: 65 NO: 69 04 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 5 NO: 15 NO: 65 NO: 69 05 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSGG NO: 4 NO: 14 NO: 65 NO: 69 06 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSGG NO: 5 NO: 15 NO: 65 NO: 69 07 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSGGS NO: 1 NO: 11 NO: 64 NO: 68 08 SEQ ID SEQ ID SEQ ID SEQ ID 3 GGSGGS GGSGGS NO: 2 NO: 12 NO: 64 NO: 68 09 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 2 NO: 12 NO: 66 NO: 70 10 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 4 NO: 14 NO: 66 NO: 70 11 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 5 NO: 15 NO: 66 NO: 70 12 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 3 NO: 13 NO: 64 NO: 68 13 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 3 NO: 13 NO: 67 NO: 71 14 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 2 NO: 12 NO: 64 NO: 68 15 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 4 NO: 14 NO: 64 NO: 68 16 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 5 NO: 15 NO: 64 NO: 68 17 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 7 NO: 17 NO: 64 NO: 68 18 SEQ ID SEQ ID SEQ ID SEQ ID 2 GGSGGS GGSG NO: 7 NO: 17 NO: 64 NO: 68 19 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 6 NO: 16 NO: 64 NO: 68 20 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 6 NO: 16 NO: 67 NO: 71 21 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 8 NO: 18 NO: 64 NO: 68 22 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 9 NO: 19 NO: 64 NO: 68 23 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 9 NO: 19 NO: 67 NO: 71 24 SEQ ID SEQ ID SEQ ID SEQ ID 1 GGSGGS GGSG NO: 10 NO: 20 NO: 64 NO: 68
[0135] In some embodiments, a tandem diabody is tandem diabody 01 (SEQ ID NO:98), 02 (SEQ ID NO:99), 03 (SEQ ID NO:100), 04 (SEQ ID NO:101), 05 (SEQ ID NO:102), 06 (SEQ ID NO:103), 07 (SEQ ID NO:104), 08 (SEQ ID NO:105), 09 (SEQ ID NO:106), 10 (SEQ ID NO:107), 11 (SEQ ID NO:108), 12 (SEQ ID NO:109), 13 (SEQ ID NO:110), 14 (SEQ ID NO:111), 15 (SEQ ID NO:112), 16 (SEQ ID NO:113), 17 (SEQ ID NO:114), 18 (SEQ ID NO:115), 19 (SEQ ID NO:116), 20 (SEQ ID NO:117), 21 (SEQ ID NO:118), 22 (SEQ ID NO:119), 23 (SEQ ID NO:120), or 24 (SEQ ID NO:121) as depicted in FIG. 9B to 9Y.
[0136] The CD33 binding protein and the CD33/CD3 bispecific binding protein (e.g., CD33/CD3 bispecific tandem diabody) described herein is produced, in some embodiments, by expressing polynucleotides encoding the polypeptide of the tandem diabody which associates with another identical polypeptide to form the antigen-binding tandem diabody. Therefore, another aspect is a polynucleotide, e.g. DNA or RNA, encoding the polypeptide of an antigen-binding tandem diabody as described herein.
[0137] The polynucleotide is constructed by known methods such as by combining the genes encoding at least four antibody variable domains either separated by peptide linkers or, in other embodiments, directly linked by a peptide bond, into a single genetic construct operably linked to a suitable promoter, and optionally a suitable transcription terminator, and expressing it in bacteria or other appropriate expression system such as, for example CHO cells. Depending on the vector system and host utilized, any number of suitable transcription and translation elements, including constitutive and inducible promoters, may be used. The promoter is selected such that it drives the expression of the polynucleotide in the respective host cell.
[0138] In some embodiments, the polynucleotide is inserted into a vector, preferably an expression vector, which represents a further embodiment. This recombinant vector can be constructed according to known methods.
[0139] A variety of expression vector/host systems may be utilized to contain and express the polynucleotide encoding the polypeptide of the described antigen-binding tandem diabody. Examples of expression vectors for expression in E. coli are pSKK (Le Gall et al., J Immunol Methods. (2004) 285(1):111-27) or pcDNA5 (Invitrogen) for expression in mammalian cells.
[0140] Thus, the antigen-binding tandem diabody as described herein, in some embodiments, is produced by introducing a vector encoding the polypeptide as described above into a host cell and culturing said host cell under conditions whereby the polypeptide chains are expressed, may be isolated and, optionally, further purified.
[0141] In other aspects, the CD33 binding protein or the CD33/CD3 bispecific binding protein (e.g., CD33/CD3 bispecific tandem diabody) described herein has a modification. Typical modifications include, but are not limited to, acetylation, acylation, ADP-ribosylation, amidation, covalent attachment of flavin, covalent attachment of a heme moiety, covalent attachment of a nucleotide or nucleotide derivative, covalent attachment of a lipid or lipid derivative, covalent attachment of phosphatidylinositol, drug conjugation, cross-linking, cyclization, disulfide bond formation, demethylation, formation of covalent crosslinks, formation of cystine, formation of pyroglutamate, formylation, gamma carboxylation, glycosylation, GPI anchor formation, hydroxylation, iodination, methylation, myristoylation, oxidation, proteolytic processing, phosphorylation, prenylation, racemization, selenoylation, sulfation, transfer-RNA mediated addition of amino acids to proteins such as arginylation, and ubiquitination. In further embodiments, the CD33 binding protein or the CD33/CD3 bispecific binding protein is modified with additional amino acids, such as a leader or secretory sequence or a sequence for purification of the polypeptide.
[0142] In other aspects, provided herein are pharmaceutical compositions comprising the CD33 binding protein, an antigen-binding tandem diabody, a vector comprising the polynucleotide encoding the polypeptide of the antigen binding tandem diabody or a host cell transformed by this vector and at least one pharmaceutically acceptable carrier. The term "pharmaceutically acceptable carrier" includes, but is not limited to, any carrier that does not interfere with the effectiveness of the biological activity of the ingredients and that is not toxic to the patient to whom it is administered. Examples of suitable pharmaceutical carriers are well known in the art and include phosphate buffered saline solutions, water, emulsions, such as oil/water emulsions, various types of wetting agents, sterile solutions etc. Such carriers can be formulated by conventional methods and can be administered to the subject at a suitable dose. Preferably, the compositions are sterile. These compositions may also contain adjuvants such as preservative, emulsifying agents and dispersing agents. Prevention of the action of microorganisms may be ensured by the inclusion of various antibacterial and antifungal agents.
[0143] Bispecific CD33/CD3 binding proteins with high-affinity binding to CD33 and CD3 are highly active in a large number of primary AML, specimens, suggesting that these molecules could be active against human AML across the entire cytogenetic/molecular disease spectrum, even in cases of minimal CD33 expression. Of note, drug-specific cytotoxicity is also observed in the presence of residual autologous T-cells and is significantly augmented by the addition of controlled amounts of healthy donor T-cells (see Example 6).
[0144] The CD33/CD3 bispecific binding proteins, in particular tandem diabodies, can induce potent cytolysis of CD33.sup.+ leukemic cells in vitro. The data indicate that high-affinity binding to both CD33 and CD3 maximizes bispecific protein-induced T-cell activation and anti-AML efficacy. High-affinity CD33/CD3-directed bispecific binding proteins, such as the tandem diabodies described herein display cytolytic activity in primary AML in vitro. Thus, these bispecific binding proteins and tandem diabodies are suitable for a therapeutic approach for the treatment of acute myeloid leukemia (AML) or other hematologic malignancies, for example, myeloid dysplastic syndrome (MDS) or myeloproliferative disease (MPD).
[0145] Therefore, provided herein are methods wherein the antigen-binding tandem diabody as described herein above is administered in an effective dose to a subject, e.g., a patient, for the treatment of a CD33.sup.+ cancer (e.g. acute myeloid leukemia (AML)), disease or condition. CD33.sup.+ cancers include, but are not limited to, acute leukemias such as acute myeloid leukemia, acute lymphoblastic leukemia (ALL) including precursor B-cell lymphoblastic leukemia, myeloid sarcoma, multiple myeloma, acute lymphomas such as acute lymphoblastic lymphoma, chronic myelomonocytic leukemia and the like. CD33.sup.+ diseases and conditions include immune suppressive states or environments attributed by myeloid derived suppressor cells (MDSCs) in certain cancers and chronic inflammation.
[0146] In some embodiments, the antigen-binding tandem diabody as described herein is administered for the treatment of acute myeloid leukemia (AML). In certain embodiments, the antigen-binding tandem diabody as described herein is administered for the treatment of an acute myeloid leukemia subtype.
[0147] The French-American-British classification system divides AML into eight subtypes: AML-M0 (minimally differentiated), AML-M1 (without maturation), AML-M2 (with granulocytic maturation), AML-M3 (promyelocytic or acute promyelocytic leukemia), AML-4 (acute myelomonocytic leukemia), AML-M5 (acute monoblastic or monocytic leukemia), AML-M6 (acute erythroid leukemia), and AML-M7 (acute megakaryoblastic leukemia). In certain instances, the antigen-binding tandem diabody as described herein is administered for the treatment of AML-M0, AML-M1, AML-M2, AML-M3, AML-M4, AML-M5, AML-M6, or AML-M7.
[0148] The WHO AML classification scheme organizes AML according to the following subtypes: AML with Recurrent Genetic Abnormalities, AML with myelodysplasia-related changes, Therapy-related myeloid neoplasms, Myeloid sarcoma, Myeloid proliferations related to Down syndrome, Blastic plasmacytoid dendritic cell neoplasm, and AML not otherwise categorized. In certain other instances, the antigen-binding tandem diabody as described herein is administered for the treatment of AML with Recurrent Genetic Abnormalities, AML with myelodysplasia-related changes, Therapy-related myeloid neoplasms, Myeloid sarcoma, Myeloid proliferations related to Down syndrome, Blastic plasmacytoid dendritic cell neoplasm, or AML not otherwise categorized.
[0149] In some other embodiments, the antigen-binding tandem diabody as described herein is administered for the treatment of a newly diagnosed, recurrent or refractory AML.
[0150] In further embodiments, the antigen-binding tandem diabody as described herein is administered for the treatment of a preleukemia blood disorder such as myeloid dysplastic syndrome (MDS) or myeloproliferative disease (MPD). In certain instances, the antigen-binding tandem diabody as described herein is administered for the treatment of MDS. In certain instances, the antigen-binding tandem diabody as described herein is administered for the treatment of MPD.
[0151] In other embodiments, the antigen-binding tandem diabody as described herein is administered for the treatment of multiple myeloma. In further embodiments, the antigen-binding tandem diabody as described herein is administered for the treatment of chronic myelomonocytic leukemia(CMML).
[0152] In other embodiments, the antigen-binding tandem diabody as described herein is administered for inhibiting or eliminating myeloid derived suppressor cells (MDSCs). MDSCs highly overexpress CD33 in certain isolated diseased tissues and possess strong immunosuppressive activities. In certain human cancers (CD33.sup.+ as well as non-CD33.sup.+), MDSCs proliferate and are activated to suppress tumor-specific CD4.sup.+ T-cell responses and induce T.sub.reg cells, allowing the tumor or cancer to flourish in a microenvironment. In chronic inflammation, MDSCs are reportedly expanded and found at inflammation sites to suppress T cell immune function. In other embodiments, the antigen-binding tandem diabody as described herein is administered for treating a condition associated with MDSCs. In yet other embodiments, the antigen-binding tandem diabody as described herein is administered to treat immune suppression. In yet other embodiments, the antigen-binding tandem diabody as described herein is administered to treat inflammation suppressed by MDSCs. In yet other embodiments, the antigen-binding tandem diabody as described herein is administered to treat a decreased immune response caused by MDSCs. In yet other embodiments, the antigen-binding tandem diabody as described herein is administered to treat angiogenesis, tumor invasion, or metastasis of cancers that are promoted by MDSCs. In yet other embodiments, the antigen-binding tandem diabody as described herein is administered to treat a cancer or tumor that is enhanced, augmented, aggravated or increased by MDSCs.
[0153] The antigen-binding tandem diabody described herein is contemplated for use as a medicament. Administration is effected by different ways, e.g. by intravenous, intraperitoneal, subcutaneous, intramuscular, topical or intradermal administration. In some embodiments, the route of administration depends on the kind of therapy and the kind of compound contained in the pharmaceutical composition. The dosage regimen will be determined by the attending physician and other clinical factors. Dosages for any one patient depends on many factors, including the patient's size, body surface area, age, sex, the particular compound to be administered, time and route of administration, the kind of therapy, general health and other drugs being administered concurrently. An "effective dose" refers to amounts of the active ingredient that are sufficient to affect the course and the severity of the disease, leading to the reduction or remission of such pathology. An "effective dose" useful for treating and/or preventing AML, may be determined using known methods.
[0154] In further embodiments, the antigen-binding tandem diabody described herein is administered in combination with a standard therapy to CD33.sup.+ cancers, diseases or conditions. Standard therapies include chemotherapies, immunotherapies, hormone therapies, radiation, surgery, gene therapies and the like. In certain instances, the antigen-binding tandem diabody described herein is administered in combination with a standard AML therapy. In certain instances, the antigen-binding tandem diabody described herein is administered in combination with cytarabine, azacitidine, decitabine, an anthracycline (e.g., daunorubicin, idarubicin, doxorubicin, and the like), amsacrine, fludarabine, clofarabine, cladribine, nelarabine, methotrexate, bortezomib, carfilzomib, melphalan, ibrutinib, thalidomide, lenalidomide, pomalidomide, apremilast, an epipodophyllotoxin (e.g., etoposide, teniposide, and the like), an anthracenedione (e.g., mitoxantrone, pixantrone, losoxantrone, piroxantrone, ametantrone and the like) an anti-CD20 agent (e.g., rituximab, ocrelizumab, ofatumumab, or combinations thereof. In certain instances, the antigen-binding tandem diabody described herein is administered in combination with cytarabine (ara-C). In certain instances, the antigen-binding tandem diabody described herein is administered in combination with azacitidine. In certain instances, the antigen-binding tandem diabody described herein is administered in combination with decitabine. In further instances, the antigen-binding tandem diabody described herein is administered in combination with an anthracycline (e.g., daunorubicin, idarubicin, doxorubicin, and the like). In other instances, the antigen-binding tandem diabody described herein is administered in combination with a checkpoint inhibitor (e.g., PD-1 inhibitor, CTLA-4 inhibitor, and the like). In yet other instances, the antigen-binding tandem diabody described herein is administered in combination with an epipodophyllotoxin (e.g., etoposide, teniposide, and the like). In yet other instances, the antigen-binding tandem diabody described herein is administered in combination with an anthracenedione (e.g., mitoxantrone, pixantrone, losoxantrone, piroxantrone, ametantrone and the like).
[0155] The examples below further illustrate the described embodiments without limiting the scope of the invention.
Example 1
Cloning of DNA Expression Constructs Encoding Single-Chain Fv Antibodies
[0156] For bacterial expression of anti-CD33 single-chain Fv (scFv) antibodies in E. coli, DNA coding sequences of all molecules were cloned into a bacterial expression vector. All expression constructs were designed to contain coding sequences for an N-terminal signal peptide and C-terminal hexa-histidine (6.times.His)-tag to facilitate antibody secretion into the periplasm and purification, respectively. The amino acid sequences of the VL and VH-domains from all anti-CD33 scFv clones are shown in Table 3 and Table 4.
Expression of Recombinant Anti-CD33 scFv Antibodies in E. coli
[0157] Recombinant scFv antibodies were expressed as soluble secreted proteins in the E. coli periplasm. In a first step a small medium culture supplemented with ampicillin was inoculated with transformed bacteria and incubated for 16 h at 28.degree. C. Subsequently, optical density was adjusted by adding a second medium supplemented with ampicillin and incubated once more at 28.degree. C. until an optical density in the range of 0.6-0.8 at 600 nm was reached. Protein expression was induced through addition of 50 .mu.M IPTG and incubation of cultures at 21-28.degree. C. and 200 rpm for up to 16 h. Following incubation, cells were pelleted (30 min, 4.degree. C., 7500 rpm) and stored at -20.degree. C. until further processing.
Purification of Anti-CD33 Single-Chain Fv Antibodies
[0158] Recombinant scFv were extracted from E. coli periplasm following centrifugation of bacterial cell cultures by resuspending cell pellets in buffer and incubation for 30 min at room temperature with gentle agitation. Cells were pelleted and supernatants containing recombinant proteins were kept. The procedure was repeated once more before supernatants were pooled and homogenized by ultrasonication. Homogenates were diluted, supplemented with low concentrations of imidazole and loaded onto a prepacked immobilized metal affinity chromatography (IMAC) column (GE Healthcare). The column was washed until baseline was reached and bound protein was then eluted with an imidazole buffer. Antibody containing fractions were pooled and subsequently purified by size-exclusion chromatography (SEC). Finally, protein eluates were concentrated by ultrafiltration and dialysed against storage buffer. Subsequent to low pH treatment (incubation at pH 3.0 for 20-24 h at 37.degree. C.), samples were neutralized using Tris. Purified proteins were stored as aliquots at -80.degree. C. until use.
Example 2
Cloning of DNA Expression Constructs Encoding Tandem Diabodies (TandAb.RTM.)
[0159] For expression of bispecific tandem diabodies in CHO cells, coding sequences of all molecules were cloned into a mammalian expression vector system. The anti-CD33 scFv domains of Example 1 were used to construct CD33/CD3 tandem diabodies in combination with an anti-CD3 scFv domain, with domains organized as shown in Table 7 and FIG. 3. In brief, gene sequences encoding anti-CD33 VH and VL domains separated by a peptide linker (VH-linker-VL or VL-linker-VH) were synthesized and subcloned. The resulting construct was digested to generate separate VH and VL coding sequences utilizing a Bam HI restriction site located within the linker sequence. These VH and VL fragments were then ligated with a DNA fragment encoding VH and VL domains of anti-CD3 (VH-linker-VL or VL-linker-VH) to yield the final construct. Domain order variants 1 to 3 of CD33/CD3 tandem diabodies are shown in FIG. 3. All expression constructs were designed to contain coding sequences for an N-terminal signal peptide and a C-terminal hexahistidine (6.times.His)-tag to facilitate antibody secretion and purification, respectively.
Expression of Tandem Diabodies in Stably Transfected CHO Cells
[0160] A CHO cell expression system (Flp-In.RTM., Life Technologies), a derivative of CHO-K1 Chinese Hamster ovary cells (ATCC, CCL-61) (Kao and Puck, Proc. Natl. Acad Sci USA 1968; 60(4):1275-81), was used. Adherent cells were subcultured according to standard cell culture protocols provided by Life Technologies.
[0161] For adaption to growth in suspension, cells were detached from tissue culture flasks and placed in serum-free medium. Suspension-adapted cells were cryopreserved in medium with 10% DMSO.
[0162] Recombinant CHO cell lines stably expressing secreted tandem diabodies were generated by transfection of suspension-adapted cells. During selection with the antibiotic Hygromycin B viable cell densities were measured twice a week, and cells were centrifuged and resuspended in fresh selection medium at a maximal density of 0.1.times.10.sup.6 viable cells/mL. Cell pools stably expressing tandem diabodies were recovered after 2-3 weeks of selection at which point cells were transferred to standard culture medium in shake flasks. Expression of recombinant secreted proteins was confirmed by performing protein gel electrophoresis or flow cytometry. Stable cell pools were cryopreserved in DMSO containing medium.
[0163] Tandem diabodies were produced in 10-day fed-batch cultures of stably transfected CHO cell lines by secretion into the cell culture supernatant. Cell culture supernatants were harvested after 10 days at culture viabilities of typically >75%. Samples were collected from the production cultures every other day and cell density and viability were assessed. On day of harvest, cell culture supernatants were cleared by centrifugation and vacuum filtration before further use.
[0164] Protein expression titers and product integrity in cell culture supernatants were analyzed by SDS-PAGE.
Purification of Tandem Diabodies
[0165] Tandem diabodies were purified from CHO cell culture supernatants in a two-step procedure. The His6-tagged constructs were subjected to Ni-NTA Superflow chromatography in a first step followed by preparative size exclusion chromatography (SEC) on Superdex 200 in a second step. Eluted tandem diabodies were characterized with regards to their homodimer (tandem diabody) content and pooled if the homodimer content was 90% or higher. Finally, pooled samples were buffer-exchanged and concentrated by ultrafiltration to a typical concentration of >1 mg/mL. Purity and homogeneity (typically >90%) of final samples were assessed by SDS PAGE under reducing and non-reducing conditions, followed by immunoblotting using an anti-His-Tag antibody as well as by analytical SEC, respectively. Purified proteins were stored at aliquots at -80.degree. C. until use.
[0166] Polypeptides of CD33/CD3 tandem diabodies are shown in Table 7 and FIG. 3. Each tandem diabody consists of two identical polypeptides (FIG. 1). Both outer linkers L1 and L3 were comprised of six amino acids GGSGGS (SEQ ID NO:95), whereas the central peptide linker 2 varied in length (4-6 amino acids) with the sequences GGSG (SEQ ID NO:96), GGSGG (SEQ ID NO:97), or GGSGGS (SEQ ID NO:95), respectively.
[0167] Using a series of anti-CD33 variable domains and anti-CD3 variable domains a large number of tandem diabody molecules was generated that could be stably produced in transfected cell lines and that maintained stability at body temperature as well as after repeated freeze/thaw cycles. To facilitate further development and preclinical toxicology studies, emphasis was placed on the selection of tandem diabody molecules that showed binding to both human and cynomolgus monkey CD33. Examples of complete amino acid sequences are shown for the single-chain of tandem diabodies 12 (SEQ ID NO:109), 14 (SEQ ID NO:111) and 16 (SEQ ID NO:113) in FIGS. 9M, 90 and 9Q, respectively. In this example the order of the variable domains and their linkers for the structures is: VL (CD3)-L1-VH (CD33)-L2-VL (CD33)-L3-VH (CD3).
Example 3
Determination of Antibody Affinity by Flow Cytometry
[0168] Cells were incubated with 100 .mu.L of serial dilutions of CD33/CD3 tandem diabodies. After washing three times with FACS buffer the cells were incubated with 0.1 mL of 10 .mu.g/mL mouse monoclonal anti-His antibody in the same buffer for 45 min on ice. After a second washing cycle, the cells were incubated with 0.1 mL of 15 .mu.g/mL FITC-conjugated goat anti-mouse IgG antibodies under the same conditions as before. As a control, cells were incubated with the anti-His IgG followed by the FITC-conjugated goat anti-mouse IgG antibodies without anti-CD33 tandem diabodies. The cells were then washed again and resuspended in 0.2 mL of FACS buffer containing 2 .mu.g/mL propidium iodide (PI) in order to exclude dead cells. The fluorescence of 1.times.10.sup.4 living cells was measured using a Beckman-Coulter FC500 MPL flow cytometer using the MXP software (Beckman-Coulter, Krefeld, Germany) or a Millipore Guava EasyCyte flow cytometer using the Incyte software (Merck Millipore, Schwalbach, Germany). Mean fluorescence intensities of the cell samples were calculated using CXP software (Beckman-Coulter, Krefeld, Germany) or Incyte software (Merck Millipore, Schwalbach, Germany). After subtracting the fluorescence intensity values of the cells stained with the secondary and tertiary reagents alone the values were used for calculation of the K.sub.D values with the equation for one-site binding (hyperbola) of the GraphPad Prism (version 6.00 for Windows, GraphPad Software, La Jolla Calif. USA).
[0169] The tandem diabodies were tested for their binding affinities to human CD3.sup.+ and CD33.sup.+ cells and cynomolgus CD3.sup.+ and CD33.sup.+ cells. Exemplary binding data for selected tandem diabodies are summarized in Table 8:
TABLE-US-00008 TABLE 8 CD3 and CD33 binding characteristics of CD33/CD3 tandem diabodies: K.sub.D on K.sub.D on K.sub.D on K.sub.D on K.sub.D ratio EC.sub.50 on T cells HL-60 KG-1 U-937 CynoCD33/ HL-60 TandAb [nm] [nM] [nM] [nM] huCD33 [pM] 01 94.2 0.6 0.9 7.1 0.7 1.9 02 69.8 0.2 0.3 0.9 1.1 0.5 03 81.9 1.1 1.8 8.9 0.6 3.6 04 79.3 0.5 0.5 1.7 1.1 1.8 05 69.5 1.0 1.2 6.2 0.8 2.7 06 86.3 0.4 0.5 1.6 0.8 1.6 07 49.7 13.7 47.9 47.1 45.8 17.8 08 2.4 0.3 0.5 1.8 0.6 1.8 09 2.4 0.5 0.3 2.2 1.0 6.8 10 1.9 0.5 1.0 1.7 0.8 7.0 11 2.6 0.3 0.5 0.6 1.2 5.9 12 1.5 0.3 0.9 0.5 1.7 1.3 13 55.7 0.2 0.3 0.5 1.6 1.1 14 2.1 0.3 0.3 1.2 1.0 1.6 15 1.3 0.4 0.3 0.9 1.1 1.8 16 2.1 0.3 0.2 0.3 1.4 1.5 17 3.3 5.0 52.5 24.4 1.9 18.4 18 1.9 3.4 16.3 15.1 3.1 6.3 19 6.3 2.8 3.6 5.4 37.3 5.7 20 143.8 4.1 7.0 7.2 33.8 10.0 21 2.1 9.7 25.1 80.2 0.9 7.6 22 4.1 0.7 2.0 8.6 0.6 3.2 23 97.2 0.4 1.0 5.1 1.9 2.8 24 2.3 5.6 12.4 39.5 1.8 9.6 #K.sub.D ratio cyno CD33/human CD33 was calculated based on the K.sub.D values measured on CHO cells expressing cynomolgus CD33 and human CD33, respectively. .sup..dagger-dbl.KD ratio hu CD3/hu CD33 was calculated based on the K.sub.D values measured on Jurkat cells (hu CD3) and the mean K.sub.D of three human CD33.sup.+ tumor cell lines (HL-60, KG-1, U937).
[0170] CD3 binding affinity and crossreactivity were evaluated in titration and flow cytometric experiments on CD3.sup.+ Jurkat cells (provided by Dr. Moldenhauer, DKFZ Heidelberg; human acute T-cell leukemia) and the cynomolgus CD3.sup.+ HSC-F cell line (JCRB, cat.:JCRB1164). CD33 binding and crossreactivity were assessed on the human CD33.sup.+ tumor cell lines: HL-60 (DSMZ, cat.:ACC 3, human B cell precursor leukemia), U-937 (DSMZ, cat.: ACC5; human histiocytic lymphoma), and KG-1 (DSMZ, cat.:ACC14; acute myeloid leukemia). The K.sub.D ratio of crossreactivity was calculated using the K.sub.D values determined on the CHO cell lines expressing either recombinant human or recombinant cynomolgus antigens.
[0171] The tandem diabodies exhibited a relatively high affinity to human CD33.sup.+ on most of the tested tumor cell lines below 1 nM. Affinities to human CD3 were determined to be equal or less than 2 nM.
Example 4
Cytotoxicity Assay
[0172] For the cytoxicity assay target cells cultured under standard conditions were harvested, washed and resuspended in diluent C, provided in the PKH67 Green Fluorescent Cell Linker Mini Kit, to a density of 2.times.10.sup.7 cells/mL. The cell suspension was then mixed with an equal volume of a double concentrated PKH67-labeling solution and incubated for 2-5 min at RT. The staining reaction was performed by adding an equal volume of FCS and incubating for 1 min. After washing the labeled target cells with complete RPMI medium, cells were counted and resuspended to a density of 2.times.10.sup.5 cells/mL in complete RPMI medium. 2.times.10.sup.4 target cells were then seeded together with enriched human T-cells as effector cells at an E:T ratio of 5:1, in the presence of increasing concentrations of the indicated tandem diabodies in individual wells of a microtiter plate, in a total volume of 200 .mu.L/well. Spontaneous cell death and killing of targets by T-cells in the absence of antibodies were determined for at least three replicates on each plate. After centrifugation the assay plates were incubated for the indicated periods of time at 37.degree. C. in a humidified atmosphere with 5% CO.sub.2. After incubation, cultures were washed once with FACS buffer and then resuspended in 150 .mu.L FACS buffer supplemented with 2 .mu.g/mL PI. The absolute amount of living target cells was measured by a positive green staining with PKH67 and negative staining for PI using a Beckman-Coulter FC500 MPL flow cytometer (Beckman-Coulter) or a Millipore Guava EasyCyte flow cytometer (Merck Millipore). Based on the measured remaining living target cells, the percentage of specific cell lysis was calculated according to the following formula: [1-(number of living targets.sub.(sample)/number of living targets.sub.(spontaneous))].times.100%. Sigmoidal dose response curves and EC.sub.50 values were calculated by non-linear regression/4-parameter logistic fit using the GraphPad Software. The lysis values obtained for a given antibody concentration were used to calculate sigmoidal dose-response curves by 4 parameter logistic fit analysis using the Prism software.
[0173] EC.sub.50 values were determined in 20-24 hour assay on CD33.sup.+ U-937 (DSMZ, cat.: ACC5; human histiocytic lymphoma) target cells with enriched human T-cells as effector cells at a ratio of 5:1. Some tandem diabodies were also tested in cytotoxicity assays on CD33.sup.+ KG-1 (DSMZ, cat.:ACC14; acute myeloid leukemia) and HL-60 target cells. Specifically, HL-60 cells were chosen as a model of an AML with relatively high cell surface expression of CD33 (arbitrary MFI [mean.+-.SEM]: 3,133.+-.215; n=3), and KG-1a was chosen as a model of an AML with very limited CD33 expression (arbitrary MFI: 277.+-.11; n=3). Exemplary cytotoxicity data for selected tandem diabodies are summarized in Table 9. Additional cytotoxicity data for HL-60 cell lines is found on Table 8, last column.
TABLE-US-00009 TABLE 9 In vitro potency of CD33/CD3 tandem diabodies on different CD33.sup.+ tumor cell lines: Tandem EC.sub.50 [pM (pg/mL)] on human CD33.sup.+ target cell lines diabody HL-60 U-937 KG-1 mean 12 1.3 (137) 0.8 (84) 1.2 (126) 1.1 (116) 14 1.6 (168) 3.6 (378) 2.6 (273) 2.6 (273) 16 1.5 (158) 1.9 (200) 1.8 (189) 1.7 (179)
[0174] EC.sub.50 values were determined in FACS-based cytotoxicity assays with primary human T-cells as effector cells at an E:T ratio of 5:1 on the indicated target cell lines incubated for 20-24 hours Each tandem diabody was tested on each tumor cell line in at least two independent experiments. Mean values are presented.
Example 5
Further Cytotoxicity Screening Experiments in Human CD33+ AML, Cell Lines at 48 Hours
[0175] As described above significant cytotoxicity was detected as early as 24 hours, however higher levels of toxicity can be detected at 48 hours. For the subsequent assays a 48-hour time point was chosen. The impact of T-cell selection on tandem diabody-induced cytotoxicity was tested. To accomplish this, unstimulated PBMCs from a healthy volunteer donor were obtained, and CD3.sup.+ cells were isolated both by simple "positive enrichment" via use of CD3 microbeads as well as by more complex "negative selection" via a microbead cocktail of antibodies against CD14, CD15, CD16, CD19, CD34, CD36, CD56, CD123, and CD235a. As depicted in FIG. 4, tandem diabody-induced cytotoxicity was greater with negatively selected healthy donor T-cells than positively selected T-cells. However, the relative cytotoxic activities of individual tandem diabodies were unaffected by the method of T-cell selection. Therefore the subsequent assays were performed with positively enriched healthy donor T-cells.
[0176] Unstimulated mononuclear cells were collected from healthy adult volunteers via leukapheresis by the Fred Hutchinson Cancer Research Center (FHCRC) Hematopoietic Cell Processing Core (Core Center of Excellence) under research protocols approved by the FHCRC Institutional Review Board. T-cells were enriched through magnetic cell sorting either via CD3 Microbeads ("positive enrichment") or via Pan T-Cell Isolation Kit ("negative selection"; both from Miltenyi Biotec, Auburn, Calif.), and then frozen in aliquots and stored in liquid nitrogen. Thawed cell aliquots were labeled with 3 .mu.M CellVue Burgundy (eBioscience, San Diego, Calif.) according to the manufacturer's instructions. Purified PBMCs were cultured in the presence of various concentrations of tandem diabody molecules.
[0177] For the quantification of drug-induced cytotoxicity cells were incubated at 37.degree. C. (in 5% CO.sub.2 and air), as in Example 4, at different E:T cell ratios. After 24-72 hours, cell numbers and drug-induced cytotoxicity, using DAPI to detect non-viable cells, were determined using a LSRII cytometer (BD Biosciences) and analyzed with FlowJo. AML cells were identified by forward/side scatter properties and, in experiments where healthy donor T-cells were added, negativity for CellVue Burgundy dye (FIG. 5). Drug-induced specific cytotoxicity is presented as: % cytotoxicity=100.times.(1-live target cells.sub.treated/live target cells.sub.control). Results from cytotoxicity assays are presented as mean values.+-.standard error of the mean (SEM). Spearman nonparametric correlation was used to compute correlations between continuous sample characteristics. All P-values are two-sided. Statistical analyses were performed using GraphPad Prism software.
[0178] In the absence of healthy donor T-cells, neither of the CD33/CD tandem diabodies exerted any noticeable cytotoxic effect on AML cell lines in the absence of T-cells, confirming the absolute requirement for T-cells for their cytotoxic effects (data not shown). In the presence of T-cells, the extent of tandem diabody-induced specific cytotoxicity was dependent on the concentration of the tandem diabody as well as the E:T cell ratio. Direct head-to-head comparisons between the CD33/CD3-directed tandem diabody molecules and one control tandem diabody (00) indicated considerable differences in antibody-induced cytotoxicity in both HL-60 cells (FIG. 6A/B and Table 10) and KG-1a cells (FIG. 6C/D and Table 10), with results being highly reproducible in repeat experiments. Overall, the degree of tandem diabody-induced cytotoxicity correlated with the binding affinity for CD3 on primary human T-cells (for cytotoxicity in KG-1a cells at 25 pM (approx. 2.5 ng/mL) and E:T=5:1: r=-0.542, p=0.009; for cytotoxicity in HL-60 cells at 25 pM and E:T=5:1: r=-0.391, p=0.07). The tandem diabodies 12, 14, 16 were highly cytotoxic for both HL-60 and KG-1a cells.
TABLE-US-00010 TABLE 10 CD25 and CD69 induction and cytotoxicity at 48 h of CD33/CD3 tandem diabodies CD3 K.sub.D CD33 K.sub.D CD25 CD69 T cell (nM) (nM) Induction Induction Proliferation in Cytotoxicity Cytotoxicity Tandem Human HL-60 EC.sub.50 EC.sub.50 PBMC EC.sub.50 HL-60 cells KG-1a cells Diabody.sup.1 T-cells cells (pM).sup.2 (pM).sup.2 (pM).sup.3 (% .+-. SEM).sup.4 (% .+-. SEM).sup.4 15 1.3 0.4 6 7 7 82.9 .+-. 3.7 80.2 .+-. 1.9 12 1.5 0.3 6 3 2 84.7 .+-. 2.3 85.6 .+-. 1.6 10 1.9 0.5 10 6 6 48.0 .+-. 2.4 78.6 .+-. 2.3 14 2.1 0.3 10 7 6 86.0 .+-. 0.4 69.8 .+-. 5.7 21 2.1 9.7 ND 225 500 12.4 .+-. 1.0 0.0 .+-. 0.2 24 2.3 5.6 ND 57 264 24.5 .+-. 1.9 1.1 .+-. 0.2 09 2.4 0.5 11 7 9 43.2 .+-. 15.8 74.6 .+-. 3.2 11 2.6 0.3 11 5 6 52.7 .+-. 8.1 84.7 .+-. 1.4 17 3.3 5.0 30 114 30 4.2 .+-. 0.2 0.7 .+-. 0.4 22 4.1 0.7 10 4 7 74.2 .+-. 7.4 44.4 .+-. 5.3 16 5.1 0.3 1 2 3 86.0 .+-. 1.4 81.3 .+-. 1.5 19 6.3 2.8 9 5 6 79.4 .+-. 3.5 83.8 .+-. 2.9 07 49.7 13.7 134 65 50 6.3 .+-. 3.3 2.1 .+-. 0.7 13 55.7 0.2 30 22 23 70.4 .+-. 2.5 1.3 .+-. 0.4 05 69.5 1 116 74 74 23.8 .+-. 6.9 0.3 .+-. 0.3 02 69.8 0.2 42 27 4 80.9 .+-. 3.6 4.6 .+-. 2.1 04 79.3 0.5 94 62 44 24.1 .+-. 4.0 0.7 .+-. 0.8 03 81.9 1.1 117 87 63 13.1 .+-. 3.6 0.0 .+-. 0.5 06 86.3 0.4 39 21 48 45.7 .+-. 6.4 1.4 .+-. 0.2 01 94.2 0.6 92 91 89 8.0 .+-. 1.6 0.4 .+-. 0.4 23 97.2 0.4 41 17 37 73.7 .+-. 2.6 1.5 .+-. 0.3 20 143.8 4.1 98 75 38 31.2 .+-. 3.9 1.1 .+-. 0.3 .sup.1Tandem Diabodies (TandAbs) are listed in order of increasing CD3 affinity. .sup.2CD25 and CD69 induction was measured after 24 hours in unfractionated PBMC cultures. .sup.3T cell proliferation induced by CD33/CD3 tandem diabodies in unfractionated PBMC with CD33+ cells present. .sup.4Cytotoxicity (%) after 48 hours of DAPI+ cells at a tandem diabodies concentration of 25 pM in the presence of healthy donor T-cells at an E:T cell ratio of 5:1 from 3 independent experiments performed in duplicate wells. ND: no CD25 activation detectable
Example 6
Further Characterization of Tandem Diabodies in Primary Human AML Specimens
[0179] For a comprehensive characterization of the cytotoxic properties of these candidates, specimens from AML patients were obtained for the studies from a FHCRC specimen repository.
[0180] Frozen aliquots of Ficoll-isolated mononuclear cells from pretreatment ("diagnostic") peripheral blood or bone marrow specimens from adult patients with AML were obtained from repositories at FHCRC. We used the 2008 WHO criteria to define AML (Vardiman et al.; Blood. 2009; 114(5):937-951) and the refined United Kingdom Medical Research Council (MRC) criteria to assign cytogenetic risk (Grimwalde et al.; Blood. 2010; 116(3):354-365). Patients provided written informed consent for the collection and use of their biospecimens for research purposes under protocols approved by the FHCRC Institutional Review Board. Clinical data were de-identified in compliance with Health Insurance Portability and Accountability Act regulations. After thawing, cells were stained with directly labeled antibodies recognizing CD33 (clone P67.6; PE-Cy7-conjugated), CD3 (clone SK7; PerCP-conjugated), CD34 (clone 8G12; APC-conjugated; all from BD Biosciences, San Jose, Calif.), and CD45 (clone HI30; APC-eFluor.RTM.780-conjugated; eBioscience). To identify nonviable cells, samples were stained with 4',6-diamidino-2-phenylindole (DAPI). At least 10,000 events were acquired on a Canto II flow cytometer (BD Biosciences), and DAPI-cells analyzed using FlowJo (Tree Star, Ashland, Oreg.).
[0181] After thawing, specimens had >58% AML blasts, as determined by flow cytometry based on CD45/side-scatter properties. Specimens had >50% viable cells immediately after thawing and >50% viable cells after 48 hours in cytokine-containing liquid culture (FIG. 7). Median age of the patients was 58.1 (range: 23.9-76.2) years; cytogenetic disease risk was favorable in 2, intermediate in 18, and adverse in 7. Information on the mutation status of NPM1, FLT3, and CEBPA was incomplete; however, one sample was known to be CEBPA.sup.double-mutant, and another sample was NPM1.sup.pos/FLT3-ITD.sup.neg. The median percentage of myeloid blasts and CD3.sup.+ T-cells in the studied specimens was 86.1% (range: 58.4-97.0%) and 2.0% (range: 0-11.9%), respectively, and the median sample viability after 48 hours in culture was 80.1% (range: 53.6-93.6%). Fifteen of the patients had newly diagnosed AML, whereas 12 either had relapsed (n=7) or refractory (n=5) disease at the time of specimen collection. As summarized in Table 11, basic characteristics of the specimens from patients with newly diagnosed AML were similar to those with relapsed/refractory disease with regard to CD33 expression on myeloid blasts, amount of autologous T-cells, proportion of myeloid blasts, and culture viability.
[0182] The addition of tandem diabody molecules to AML specimen cultures resulted in modest, dose-dependent cytotoxicity (FIG. 8A), demonstrating that autologous T-cells, contained in the specimens from patients with active AML, can be engaged to lyse leukemic cells. In the presence of healthy donor T-cells, the cytotoxic activity of individual tandem diabodies was strictly dependent on the drug dose and the E:T cell ratio (FIG. 8B/C). However, high activity of tandem diabodies was observed even in some specimens with very low CD33 expression on AML blasts. Among the tandem diabody molecules, 12 appeared to be the most active, since it had the highest cytotoxicity at low concentrations (2.5 pM (approx. 250 ng/mL) and, to a less pronounced degree, also 10 pM (approx. 1 ng/mL)) at both E:T=1:3 and E:T=1:1.
[0183] The CD33/CD3 tandem diabodies have been screened in representative AML patient blood samples, which varied in terms of patient sex, age, disease stage (newly diagnosed, relapsed, refractory), degree of CD33 expression and cytogenic risk (Table 11). Remarkably, a number of examined tandem diabodies (e.g., 02, 08, 09, 11, 12, 14, 16, 19, 22 and 23) were highly active in nearly all patient samples across the disease spectrum as shown in FIG. 15. Moreover, the extent and scope of activity is similar in all stages of AML, including newly-diagnosed, relapsed and refractory patients.
TABLE-US-00011 TABLE 11 Characteristics of primary AML specimens All patients Newly diagnosed Relapsed/refractory (n = 27) AML (n = 15) AML (n = 12) Median age (range), years 58.1 (23.9-76.2) 64.0 (40.2-76.2) 44.4 (23.9-67.4) Cytogenetic/molecular risk Favorable 2 2 -- Intermediate 18 10 8 CEBPA.sup.double-mutant 1 1 -- NPM1.sup.pos/FLT3-ITD.sup.neg 1 -- 1 NPM1.sup.pos/FLT3-ITD.sup.pos or NPM1.sup.neg/FLT3-ITD.sup.pos 10 5 5 Adverse 7 3 4 Specimen source Bone marrow 11 4 7 Peripheral blood 16 11 5 Median % blasts (range) 86.1 (58.4-97.0) 86.1 (66.7-95.5) 86.7 (58.4-97.0) Median CD33 expression on blasts (range) 849 (5-5,356) 849 (5-5,356) 788 (7-2,242) Median % T-cells (range) 2.0 (0-11.9).sup. 1.6 (0-11.9).sup. 2.1 (0.7-8.7) Median % viability at 48 hours (range) 80.1 (53.6-93.6) 76.0 (53.6-93.6) 83.5 (63.9-93.1)
Example 7
Potency and Efficacy of CD33/CD3 Tandem Diabody 12 and Tandem Diabody 16 on Different CD33.sup.+ Cell Lines of Various Origin Expressing Different Levels of CD33
[0184] In order to assess whether potency and efficacy of CD33/CD3 tandem diabodies depend on the CD33 density on the target cells, various human CD33.sup.+ tumor cell lines and CHO cells expressing recombinant human CD33 were tested for their CD33 expression levels using the QIFIKIT quantification kit and anti-CD33 mAb WM53. The results in Table 12 show that the CD33 densities on the tumor cell lines were in the range between .about.1300 SABC (standardized antibody binding capacity) and .about.46000 SABC. The expression on CHO-CD33 cells was .about.197000 SABC, substantially higher than on the tumor cell lines. All tested CD33.sup.+ cell lines were used as target cells in at least 3 independent FACS-based cytotoxicity assays with human T-cells as effector cells at an effector-to-target ratio of 5:1 in the presence of serial dilutions of CD33/CD3 tandem diabody 12 and tandem diabody 16. In each assay EC.sub.50 and tandem diabody-mediated lysis values were calculated by non-linear regression. The results demonstrate that neither the potency (EC.sub.50 values) nor the efficacy (% lysis) of 12 and 16 correlates with the CD33 density on the surface of target cells.
[0185] Noteworthy, at least 12 and 16 exhibit their cytotoxic activity also against cells like SEM with very low CD33 densities of below 1500 SABC.
TABLE-US-00012 TABLE 12 CD33 target cell surface expression and cytotoxic potency of CD33/CD3 tandem diabody 12 and tandem diabody 16: CD33 density 12 16 [SABC] EC.sub.50 [pM] EC.sub.50 [pM] Cell line mean SD mean SD mean SD CHO-CD33 196990 28053 11.8 11.2 24.0 19.5 HL-60 45948 4478 1.4 0.5 1.6 0.4 KG-1 42828 6923 1.0 0.6 1.9 2.0 KASUMI-1 25922 6484 1.3 0.6 2.4 1.4 THP-1 22065 415 1.9 0.2 6.0 1.2 RPMI-8226 19931 2604 14.0 17.8 2.8 2.0 U-937 17669 4593 0.9 0.1 1.3 0.6 K562 13789 2156 4.5 1.3 4.8 2.7 BV-173 8518 1231 1.4 0.6 3.2 1.6 SEM 1306 144.2 2.2 0.5 5.1 3.0
[0186] The standardized antibody binding capacity (SABC) on CD33.sup.+ cell lines was determined using QIFIKIT and the anti-CD33 mAb WM53. EC.sub.50 values for tandem diaboody 12 and tandem diabody 16 redirected target cell lysis were determined in FACS-based cytotoxicity assays with human primary T-cells as effector cells at E:T ratios of 5:1 and 20-24 h incubation; assays with CD33-expressing CHO cells were incubated for 40-48 h. Mean and SD of at least 3 independent assays are shown.
Example 8
TandAb-Activation of T-Cells and In Vitro Killing of AML Cells
[0187] TandAbs were incubated with purified human T cells and a VPD-450-labeled human CD33.sup.+ leukemia cell line, KG-1, or the CD33.sup.- human ALL cell line, G2 (E:T 5:1). Flow cytometry was used to evaluate target cell lysis by TandAbs (10.sup.15 to 10.sup.-8M; 24 h, 37.degree. C.).
[0188] Incubation of TandAbs 12, 16, and 19 with human T cells efficiently lysed KG-1 cells (IC50.about.0.01, 0.5, and 5 pM respectively). Up to 40% of T cells were activated (CD25+) rising with cytotoxic activity. A control TandAb with an irrelevant target, 00 (>10.sup.-7M), did not result in significant killing of KG-1 in vitro. Separately, 16 induced lysis of KG-1 cells (IC50=5.times.10.sup.-12M) while 1.times.10.sup.-8M had no effect on CD33- G2 cells. The results indicate thats T cells become activated and potently lyse tumor cells when targeted to CD33+ leukemic cells (KG-1) and primary CD33+ AML blasts by CD33/CD3 TandAbs.
Example 9
Epitope Mapping
[0189] Tandem diabodies containing different CD33 binding moieties were subjected to epitope mapping using CLIPS Technology (Pepscan) in order to identify CD33-binding epitopes.
[0190] CLIPS Technology facilitates the structuring of peptides into single loops, double-loops, triple loops, sheet-like folds, helix-like folds, and combinations thereof, offering the possibility to map discontinuous epitopes of the target molecule.
[0191] An array of more than 7000 independent peptides was synthesized and the binding of each antibody to the peptides was tested in an ELISA.
[0192] The tandem diabodies 12, 14, 16 and 22 bind to the stretch .sub.62DQEVQEETQ.sub.70 (SEQ ID NO:94) in the first Ig like domain of human CD33. The respective amino acid stretches are shown underlined and in bold in FIG. 9A. It is contemplated that tandem diabodies 01, 02, 04, 06, 08, 09, 13 and 23 also bind to this epitope as these tandem diabodies share the same CD33 binding domains (SEQ ID NOs:2 and 12, 3 and 13, 5 and 15, 9 and 19) as tandem diabodies 12, 14 16 and 12.
Example 10
Dose-Response in a Prophylactic In Vivo Tumor Model
[0193] Tandem diabodies 12 and 16 are compared at different dose levels in a prophylactic HL-60 tumor xenograft model in NOD/scid mice reconstituted with human T-cells. In order to achieve a dose-response three dose levels at 10, 1 and 0.1 .mu.g (0.5, 0.05, and 0.005 mg/kg) were selected.
[0194] Eight experimental groups of immunodeficient NOD/scid mice were xenotransplanted by subcutaneous injection with a suspension of 4.times.10.sup.6 HL-60 cells. Prior to injection cells were mixed with 3.times.10.sup.6 T-cells isolated from buffy coats (healthy donors) employing negative selection. To account for potential donor variability of the T-cells, each of the experimental groups was subdivided into three cohorts each receiving T-cells of one individual donor only. All animals of the experimental groups transplanted with tumor cells and T-cells received an intravenous bolus on days 0, 1, 2, 3 and 4 (qd.times.d5) of either vehicle (control) or 16 or 12 at three different dose levels as indicated (0.1 .mu.g, 1 .mu.g, and 10 .mu.g). One group without effector cells and vehicle treatment served as an additional control. Table 13 summarizes group allocation and dosing schedule.
TABLE-US-00013 TABLE 13 Schedule Group treatment dose Cell concentration/animal Cohort (iv) n 1 Vehicle -- 4 .times. 106 HL-60 4 2 Vehicle -- 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1, 2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3 3 16 10 .mu.g 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1, 2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3 4 16 1 .mu.g 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1, 2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3 5 16 0.1 .mu.g 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1,2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3 6 12 10 .mu.g 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1, 2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3 7 12 1 .mu.g 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1, 2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3 8 12 0.1 .mu.g 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 1 Day 0, 1, 2, 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 2 3, 4 3 4 .times. 10.sup.6 HL-60 + 3 .times. 10.sup.6 T-cells Cohort 3 3
[0195] Treatment groups for the in vivo dose-response study in a HL-60 xenograft model. All animals in the control groups reliably developed a tumor and exhibited homogeneous tumor growth. The presence of T-cells had no influence on tumor development. No difference in HL-60 growth was observed in the presence or absence of T-cells in the vehicle-treated control groups.
[0196] Treatment with both test items revealed a clear dose-dependent anti-tumor effect (FIG. 10). No substantial difference was found between the two tandem diabodies. Plotting of mean tumor volumes in FIG. 10 was restricted to day 29 when most of the treatment groups were complete. The study was continued until day 45 and animals were observed for tumor-free survival. In the groups treated with 10 or 1 .mu.g of 16, 6 of 9 animals were tumor-free at the end of the observation period and 5 of 9 animals receiving 10 .mu.g of 12 were tumor-free on day 45. One animal remained tumor-free when treated with 1 .mu.g of 12.
[0197] All animals in the control groups reliably developed a tumor and exhibited homogeneous tumor growth. Treatment with either of the tandem diabodies revealed a dose-dependent anti-tumor effect and no substantial difference was found between the two tandem diabodies until day 29.
[0198] Detectable differences were observed only after prolonged observation (day 45), at which time the low dose and control groups had already been terminated due to the growth of large tumors. Groups treated with 16 had more tumor-free animals.
Example 11
Established Tumor Model
[0199] A xenograft model in NOD/scid mice with pre-established HL-60 tumors employing 16 was developed to demonstrate proof of concept.
[0200] In brief, female immune-deficient NOD/scid mice were sub-lethally irradiated (2 Gy) and subcutaneously inoculated with 4.times.10.sup.6 HL-60 cells. On day 9 the animals received a single bolus injection of anti-asialo GM1 rabbit antibody (Wako, Neuss, Germany) to deplete murine natural killer (NK) cells. On day 10, when the tumor reached a volume between 50-150 mm.sup.3 (mean 73.+-.11 mm.sup.3) animals were allocated to 3 treatment groups. Groups 2 and 3 (8 animals each) were intraperitoneally injected with 1.5.times.10.sup.7 activated human T-cells. Prior to injection T-cells were isolated from buffy coats (healthy donors) employing negative selection. T-cells were expanded and activated with the T-Cell Activation/Expansion Kit according to the manufacturer's specification (Miltenyi Biotech). In order to address potential donor variability Groups 2 and 3 were subdivided into two cohorts each receiving expanded and activated T-cells from an individual donor. Each cohort received T-cells from one individual T-cell donor only.
TABLE-US-00014 TABLE 14 Treatment groups for the established HL-60 xenograft model. Animals Inoculated cells Treatment Group (n) Day 0, sc. Day 10, ip. Cohort Day 13 to 21, once daily 1 5 4 .times. 10.sup.6 HL-60 Vehicle (iv) 2 4 4 .times. 10.sup.6 HL-60 1.5 .times. 10.sup.7 T-cells (Donor 1) 1 Vehicle (iv) 4 4 .times. 10.sup.6 HL-60 1.5 .times. 10.sup.7 T-cells (Donor 2) 2 3 4 4 .times. 10.sup.6 HL-60 1.5 .times. 10.sup.7 T-cells (Donor 1) 1 TandAb 16 (iv) 50 .mu.g 4 4 .times. 10.sup.6 HL-60 1.5 .times. 10.sup.7 T-cells (Donor 2) 2
[0201] Starting on day 13 animals in Group 3 displayed a mean tumor volume of 105 mm.sup.3 and were treated with a total of 9 intravenous doses of 50 .mu.g tandem diabody 16 (qd.times.9d). Table 14 illustrates group allocation and dosing schedule. Groups 1 and 2 were only treated with the vehicle. Body weight and tumor volume were determined until day 27.
[0202] All animals reliably developed a tumor, which was palpable on day 6. The mean tumor volume of vehicle-treated Group 1 and 2 (HL-60) animals continually increased until study termination on day 27 (FIG. 11). In Group 2 animals that received primary activated human T-cells in addition to HL-60 tumor cells, the mean tumor volume increased faster compared to Group 1 (HL-60 only).
[0203] Repeated intravenous treatment from days 13 to 21 (qd.times.d9) with tandem diabody 16 (50 .mu.g/animal; 2.5 mg/kg) in the presence of human T-cells (Group 3) rapidly delayed tumor growth relative to Group 1 and Group 2. Tandem diabody 16 delayed tumor growth in Group 3 by approximately 4-5 days compared to vehicle-treated control group (Group 2). Statistically significant differences in the time period from day 6 to day 27 were identified between Group 2 (HL-60, T-cells, vehicle) and Group 3 (HL-60, T-cells, 16) on day 22 (p<0.05), day 23 (p<0.01) and day 27 (p<0.01) (Two-way Repeated Measures ANOVA with Bonferroni post-tests). No statistically significant differences were present between Group 1 and Group 3 due to unusual slow growth of the tumor in Group 1.
[0204] No donor variability with regard to T-cell activity was observed, when comparing tumor development in Cohort 1 and Cohort 2 within a group, which received T-cells from different donors (see Table 14).
[0205] Example 10 shows that a xenograft model in NOD/scid mice with a pre-established HL-60 tumor (AML) and intraperitoneally-engrafted human T-cells was successfully developed. Repeated dosing with tandem diabody 16 at a single dose level lead to a statistically significant delay in tumor growth in comparison to the respective vehicle-treated control group. The data generated are comparable to results published for a similar study with a CD33/CD3 BiTE.TM. (Aigner et al., 2012; Leukemia, 2013, April; 27(5):1107-15).
Example 12
Efficacy of CD33/CD3 Tandem Diabodies in an AML PDX Model in NSG Mice
[0206] Cryopreserved cells from an AML patient whose CD33.sup.+ leukemia contained 2-4% CD3.sup.+ T-cells were used to establish an AML PDX model in NSG mice. One hour post-injection of tumor cells into irradiated (250 cGy) NSG mice, CD33/CD3 tandem diabodies, 16 or 12, at either of two i. v. doses (50 .mu.g or 5 .mu.g; n=8 mice/group) were injected in a 200 .mu.L bolus. Additional injections of tandem diabodies were performed on each of the following 4 days. Mice were weighed once weekly, and subsequently were sacrificed on day 38 to permit collection of peripheral blood, bone marrow, and spleen for analysis by flow cytometry (huCD33, huCD34, huCD45, muCD45, huCD14, huCD3, huCD4, huCD8, and 7AAD). The results are shown in FIG. 12.
[0207] FIG. 12 shows that untreated mice had substantial amounts of human blasts in the bone marrow and spleen after 38 days. In contrast, mice treated with daily i. v. injections of tandem diabodies 12 or 16 exhibited substantially lower numbers of human AML blasts in the bone marrow and in the spleen. The strong anti-AML effect of the CD33/CD3 tandem diabody was observed at both dose levels (5 and 50 .mu.g/injection).
[0208] The observed anti-AML effect for both CD33/CD3 tandem diabodies, 12 and 16, was much stronger than the effect of a CD123/CD3 DART.RTM. antibody targeting AML in an identical mouse model (Hussaini et al.:" Targeting CD123 In Leukemic Stem Cells Using Dual Affinity ReTargeting Molecules (DARTs.RTM.) Nov. 15, 2013; Blood: 122 (21)). In contrast to the CD33/CD3 tandem diabodies which eliminated nearly all AML blasts in bone marrow and spleen, Hussaini et al. reported that the CD123/CD3 DART.RTM. reduced the number of AML blasts in the bone marrow and spleen in the PDX model only by factor 50-1000 at 2.5 and 0.25 mg/kg, the authors further reported that the CD123/CD3 DART.TM. reduced the number of AML blasts in bone marrow and spleen in the PDX model only by 40-78% at 0.5 mg/kg.
Example 13
Fast Onset of CD33/CD3 Tandem Diabody 16-Mediated Target Cell Lysis
[0209] In order to assess the kinetics of CD33/CD3 tandem diabody-mediated target cell lysis, calcein-release cytotoxicity assays with different incubation times were performed. Calcein-labeled CD33.sup.+ HL-60 target cells were incubated with serial dilutions of tandem diabody 16 in the presence of primary human T cells as effector cells at an E:T ratio of 25:1 for 30 min, 1 h, 2 h, 3 h, 4 h, or 5 h. At each time point the calcein that was released from lysed target cells was used to calculate the EC.sub.50 value and tandem diabody 16-mediated target cell lysis using non-linear regression/sigmoidal dose-response. FIG. 13 shows an unexpected fast onset of tandem diabody-mediated target cells lysis with more than 40% lysis after 30 min incubation at saturating tandem diabody concentrations. After 4 hours incubation more than 90% target cell lysis was reached. Table 15 and FIG. 14 summarize the EC.sub.50 and specific lysis values determined for tandem diabody 16 at incubation times between 30 min and 5 hours. The results further demonstrate that under the used assay conditions maximal potency (lowest EC.sub.50 value) was reached after 2 hours incubation and that after 5 hours incubation almost all target cells were lysed. Altogether these results demonstrate a very fast, potent and efficacious target cell lysis mediated by CD33/CD3 tandem diabodies.
TABLE-US-00015 TABLE 15 Kinetics of EC.sub.50 and lysis values determined for tandem diabody 16 incubation tandem diabody- time [min] EC.sub.50 [pM] mediated lysis [%] 30 4.8 44.1 60 2.5 59.8 120 1.6 75.1 180 1.6 88.8 240 1.5 93.7 300 1.6 97.4
Example 14
Proof-of-Concept Clinical Trial Protocol for Administration of CD33/CD3 Tandem Diabodies to AML Patients
[0210] This Phase I/II clinical trial for studying CD33/CD3 tandem diabody 16 as a treatment for with acute myeloid leukemia (AML).
[0211] Study Outcomes:
[0212] Primary: Maximum tolerated dose of CD33/CD3 tandem diabody 16
[0213] Secondary: To determine whether in vitro response of CD33/CD3 tandem diabody 16 is associated with clinical response
[0214] Phase I
[0215] The maximum tolerated dose (MTD) will be determined in the phase I section of the trial. [0216] 1.1 The maximum tolerated dose (MTD) will be determined in the phase I section of the trial. [0217] 1.2 Patients who fulfill eligibility criteria will be entered into the trial to CD33/CD3 tandem diabody 16. [0218] 1.3 The goal is to identify the highest dose of CD33/CD3 tandem diabody 16 that can be administered safely without severe or unmanageable side effects in participants. The dose given will depend on the number of participants who have been enrolled in the study prior and how well the dose was tolerated. Not all participants will receive the same dose.
[0219] Phase II [0220] 2.1 A subsequent phase II section will be treated at the MTD with a goal of determining if therapy with therapy of CD33/CD3 tandem diabody 16 results in at least a 20% response rate. [0221] Primary Outcome for the Phase II--To determine if therapy of CD33/CD3 tandem diabody 16 results in at least 20% of patients achieving a clinical response (blast response, minor response, partial response, or complete response)
[0222] Eligibility: [0223] Documented AML by peripheral blood and bone marrow analyses meeting WHO criteria, excluding patients with acute promyelocytic leukemia (APL) [0224] Patients with AML refractory to primary induction chemotherapy, relapsed disease, or age.gtoreq.60 and not appropriate for standard cytotoxic therapy due to age, performance status, and/or adverse risk factors according to the treating physician [0225] Age.gtoreq.18 years [0226] Karnofsky performance status.gtoreq.50% or ECOG performance status 0-2 [0227] Life expectancy.gtoreq.6 weeks
[0228] While certain embodiments have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the invention. It should be understood that various alternatives to the embodiments described herein may be employed in practicing the embodiments. It is intended that the following claims define the scope of the invention and that methods and structures within the scope of these claims and their equivalents be covered thereby.
Sequence CWU 1
1
1221108PRTHomo sapiens 1Ser Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser
Val Ala Pro Gly Gln1 5 10 15Thr Ala Met Ile Thr Cys Gly Gly Asn Asn
Ile Gly Ser Thr Thr Val 20 25 30His Trp Tyr Gln Gln Lys Pro Gly Gln
Ala Pro Val Leu Val Val Tyr 35 40 45Asp Asp Asn Glu Arg Pro Ser Gly
Ile Pro Glu Arg Phe Ser Gly Ser 50 55 60Asn Ser Gly Ser Thr Ala Thr
Leu Thr Ile Asn Arg Val Glu Ala Gly65 70 75 80Asp Glu Ala Asp Tyr
Tyr Cys Gln Val Trp Asp Ser Gly Ser Asp His 85 90 95Val Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu 100 1052110PRTHomo sapiens 2Gln Ser
Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro Gly Gln1 5 10 15Arg
Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn Ile Gly Ser Asn 20 25
30Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu
35 40 45Ile Tyr Gly Asn Asn Gln Arg Pro Ser Gly Val Pro Asp Arg Phe
Ser 50 55 60Gly Ser Lys Ser Gly Ser Ser Ala Ser Leu Ala Ile Ser Gly
Leu Gln65 70 75 80Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp
Asp Asp Ser Leu 85 90 95Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu
Thr Val Leu 100 105 1103110PRTHomo sapiens 3Gln Ser Val Leu Thr Gln
Pro Pro Ser Ala Ser Gly Thr Pro Gly Gln1 5 10 15Arg Val Thr Ile Ser
Cys Ser Gly Ser Arg Ser Asn Ile Gly Ser Asn 20 25 30Thr Val Asn Trp
Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu 35 40 45Ile Tyr Gly
Asn Asn Gln Arg Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55 60Gly Ser
Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70 75
80Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp Ser Leu
85 90 95Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 100
105 1104110PRTHomo sapiens 4Gln Ser Val Leu Thr Gln Pro Pro Ser Ala
Ser Gly Thr Pro Gly Gln1 5 10 15Arg Val Thr Ile Ser Cys Ser Gly Ser
Arg Ser Asn Ile Gly Ser Asn 20 25 30Thr Val Asn Trp Tyr Gln Gln Leu
Pro Gly Thr Ala Pro Lys Leu Leu 35 40 45Ile Tyr Gly Asn Asn Gln Arg
Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55 60Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70 75 80Ser Glu Asp Glu
Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp Ser Leu 85 90 95Ile Gly Trp
Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 100 105 1105110PRTHomo
sapiens 5Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
Gly Gln1 5 10 15Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn Ile
Gly Ser Asn 20 25 30Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala
Pro Lys Leu Leu 35 40 45Ile Tyr Gly Asn Asn Gln Arg Pro Ser Gly Val
Pro Asp Arg Phe Ser 50 55 60Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu
Ala Ile Ser Gly Leu Gln65 70 75 80Ser Glu Asp Glu Ala Asp Tyr Tyr
Cys Ala Thr Trp Asp Asp Ser Leu 85 90 95Ile Gly Trp Val Phe Gly Gly
Gly Thr Lys Leu Thr Val Leu 100 105 1106110PRTHomo sapiens 6Gln Ser
Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro Gly Gln1 5 10 15Arg
Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn 20 25
30Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu
35 40 45Ile Tyr Ser Asp Asn Gln Arg Pro Ser Gly Val Pro Asp Arg Phe
Ser 50 55 60Gly Ser Lys Ser Gly Ser Ser Ala Ser Leu Ala Ile Ser Gly
Leu Gln65 70 75 80Ser Asp Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp
Asp Asp Ser Leu 85 90 95Asn Gly Ala Val Phe Gly Gly Gly Thr Lys Leu
Thr Val Leu 100 105 1107111PRTHomo sapiens 7Gln Ser Val Leu Thr Gln
Pro Pro Ser Val Ser Gly Ala Pro Gly Gln1 5 10 15Arg Val Thr Ile Ser
Cys Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly 20 25 30Tyr Asp Val His
Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu 35 40 45Leu Ile Tyr
Gly Asn Ser Asn Arg Pro Ser Gly Val Pro Asp Arg Phe 50 55 60Ser Gly
Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Thr Gly Leu65 70 75
80Gln Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser Ser
85 90 95Leu Ser Asp Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu
100 105 1108110PRTHomo sapiens 8Gln Ser Val Leu Thr Gln Pro Pro Ser
Ala Ser Gly Thr Pro Gly Gln1 5 10 15Arg Val Thr Ile Ser Cys Ser Gly
Ser Ser Ser Asn Ile Gly Ser Asn 20 25 30Ile Val Asn Trp Tyr Gln Gln
Leu Pro Gly Thr Ala Pro Lys Leu Leu 35 40 45Ile Tyr Ser Asn Asn Gln
Arg Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55 60Gly Ser Lys Ser Gly
Thr Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70 75 80Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Ala Trp Asp Asp Ser Leu 85 90 95Lys Gly
Tyr Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 100 105
1109110PRTHomo sapiens 9Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser
Gly Thr Pro Gly Gln1 5 10 15Arg Val Thr Ile Ser Cys Ser Gly Ser Ser
Ser Asn Ile Gly Ser Asn 20 25 30Thr Val Lys Trp Tyr Gln Gln Leu Pro
Gly Thr Ala Pro Lys Leu Leu 35 40 45Ile Tyr Ser Asn Asn Gln Arg Ser
Ser Gly Val Pro Asp Arg Phe Ser 50 55 60Gly Ser Lys Ser Gly Ser Ser
Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70 75 80Ser Glu Asp Glu Ala
Asp Tyr Tyr Cys Ala Ala Trp Asp Asp Ser Leu 85 90 95Asn Gly Tyr Val
Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 100 105 11010110PRTHomo
sapiens 10Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
Gly Gln1 5 10 15Arg Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile
Gly Asp Asn 20 25 30Val Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala
Pro Lys Leu Leu 35 40 45Ile Tyr Ser Thr Asn Lys Arg Pro Ser Gly Val
Pro Asp Arg Phe Ser 50 55 60Gly Ser Lys Ser Gly Ser Ser Ala Ser Leu
Ala Ile Ser Gly Leu Gln65 70 75 80Ser Glu Asp Glu Ala Asp Tyr Tyr
Cys Ala Ala Trp Asp Asp Ser Leu 85 90 95Ser Ala Tyr Val Phe Gly Gly
Gly Thr Lys Leu Thr Val Leu 100 105 11011119PRTHomo sapiens 11Gln
Val Gln Leu Gln Glu Ser Gly Gly Gly Val Val Gln Pro Gly Arg1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Ser Phe Ser Asn Tyr
20 25 30Gly Ile His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp
Val 35 40 45Ala Leu Ile Ser Tyr Asp Gly Asn Lys Lys Phe Tyr Ala Asp
Ser Val 50 55 60Lys Gly Arg Phe Ala Ile Ser Arg Asp Thr Ser Lys Asn
Thr Val Asp65 70 75 80Leu Gln Met Thr Ser Leu Arg Pro Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95Ala Lys Asp Arg Leu Glu Ser Ala Ala Phe
Asp Tyr Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val Ser Ser
11512122PRTHomo sapiens 12Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Asp Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe Ala Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Met
Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp
Arg Ala Asn Thr Asp Phe Ser Tyr Gly Met Asp Val Trp 100 105 110Gly
Gln Gly Thr Leu Val Thr Val Ser Ser 115 12013122PRTHomo sapiens
13Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
Tyr 20 25 30Asp Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Trp Met Asn Pro Asn Ser Gly Asn Thr Gly Phe Ala
Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr
Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Arg Ala Val Thr Asp Tyr
Tyr Tyr Gly Met Asp Val Trp 100 105 110Gly Gln Gly Thr Leu Val Thr
Val Ser Ser 115 12014122PRTHomo sapiens 14Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser
Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Asp Ile Asn Trp
Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Trp Met
Asn Pro Asn Ser Gly Asn Thr Gly Phe Ala Gln Lys Phe 50 55 60Gln Gly
Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Arg Ala Asn Thr Asp Tyr Ser Phe Gly Met Asp Val
Trp 100 105 110Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115
12015122PRTHomo sapiens 15Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Asp Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe Ala Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Met
Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp
Arg Ala Asn Thr Asp Tyr Ser Leu Gly Met Asp Val Trp 100 105 110Gly
Gln Gly Thr Leu Val Thr Val Ser Ser 115 12016123PRTHomo sapiens
16Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
Tyr 20 25 30Tyr Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Ile Ile Asn Pro Ser Gly Gly Ser Thr Ser Tyr Ala
Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr
Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Val Val Pro Ala Ala Ile
Asp Tyr Tyr Gly Met Asp Val 100 105 110Trp Gly Gln Gly Thr Thr Val
Thr Val Ser Ser 115 12017119PRTHomo sapiens 17Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Glu1 5 10 15Ser Leu Lys Ile
Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr 20 25 30Trp Ile Gly
Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu Trp Met 35 40 45Gly Ile
Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe 50 55 60Gln
Gly Gln Val Thr Ile Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70 75
80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Tyr Cys
85 90 95Ala Arg His Lys Arg Gly Ser Asp Ala Phe Asp Ile Trp Gly Gln
Gly 100 105 110Thr Thr Val Thr Val Ser Ser 11518122PRTHomo sapiens
18Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser
Tyr 20 25 30Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Gly Ile Tyr Pro Ile Phe Gly Ser Ala Asn Tyr Ala
Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Glu Tyr Tyr Tyr Asp Ser Ser
Glu Trp Ala Phe Asp Ile Trp 100 105 110Gly Gln Gly Thr Leu Val Thr
Val Ser Ser 115 12019122PRTHomo sapiens 19Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ser1 5 10 15Ser Val Lys Val Ser
Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30Gly Ile Ser Trp
Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Gly Ile
Ile Pro Ile Phe Gly Ser Ala His Tyr Ala Gln Lys Phe 50 55 60Gln Gly
Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Glu Tyr Tyr Tyr Asp Ser Ser Glu Trp Ala Phe Asp Ile
Trp 100 105 110Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115
12020122PRTHomo sapiens 20Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ser1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Gly Thr Phe Asp Ser Tyr 20 25 30Ala Ile Ser Trp Val Arg Gln Ala
Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Gly Ile Ile Pro Ile Phe
Gly Ser Ala His Tyr Ser Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Ile
Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Glu
Tyr Tyr Tyr Asp Ser Ser Glu Trp Ala Phe Asp Ile Trp 100 105 110Gly
Gln Gly Thr Leu Val Thr Val Ser Ser 115 1202111PRTHomo sapiens
21Gly Gly Asn Asn Ile Gly Ser Thr Thr Val His1 5 102213PRTHomo
sapiens 22Ser Gly Ser Arg Ser Asn Ile Gly Ser Asn Thr Val Asn1 5
102313PRTHomo sapiens 23Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn Thr
Val Asn1 5 102414PRTHomo sapiens 24Thr Gly Ser Ser Ser Asn Ile Gly
Ala Gly Tyr Asp Val His1 5 102513PRTHomo sapiens 25Ser Gly Ser Ser
Ser Asn Ile Gly Ser Asn Ile Val Asn1 5 102613PRTHomo sapiens 26Ser
Gly Ser Ser Ser Asn Ile Gly Ser Asn Thr Val Lys1 5 102713PRTHomo
sapiens 27Ser Gly Ser Ser Ser Asn Ile Gly Asp Asn Val Val Asn1 5
10287PRTHomo sapiens 28Asp Asp Asn Glu Arg Pro
Ser1 5297PRTHomo sapiens 29Gly Asn Asn Gln Arg Pro Ser1 5307PRTHomo
sapiens 30Ser Asp Asn Gln Arg Pro Ser1 5317PRTHomo sapiens 31Gly
Asn Ser Asn Arg Pro Ser1 5327PRTHomo sapiens 32Ser Asn Asn Gln Arg
Pro Ser1 5337PRTHomo sapiens 33Ser Asn Asn Gln Arg Ser Ser1
5347PRTHomo sapiens 34Ser Thr Asn Lys Arg Pro Ser1 5359PRTHomo
sapiens 35Gln Val Trp Asp Ser Gly Ser Asp His1 5369PRTHomo sapiens
36Ala Thr Trp Asp Asp Ser Leu Ile Gly1 5379PRTHomo sapiens 37Ala
Thr Trp Asp Asp Ser Leu Asn Gly1 5389PRTHomo sapiens 38Gln Ser Tyr
Asp Ser Ser Leu Ser Asp1 5399PRTHomo sapiens 39Ala Ala Trp Asp Asp
Ser Leu Lys Gly1 5409PRTHomo sapiens 40Ala Ala Trp Asp Asp Ser Leu
Asn Gly1 5419PRTHomo sapiens 41Ala Ala Trp Asp Asp Ser Leu Ser Ala1
5426PRTHomo sapiens 42Ser Asn Tyr Gly Ile His1 5436PRTHomo sapiens
43Thr Ser Tyr Asp Ile Asn1 5446PRTHomo sapiens 44Thr Ser Tyr Tyr
Met His1 5456PRTHomo sapiens 45Thr Ser Tyr Trp Ile Gly1 5466PRTHomo
sapiens 46Ser Ser Tyr Ala Ile Ser1 5476PRTHomo sapiens 47Ser Ser
Tyr Gly Ile Ser1 5486PRTHomo sapiens 48Asp Ser Tyr Ala Ile Ser1
54917PRTHomo sapiens 49Leu Ile Ser Tyr Asp Gly Asn Lys Lys Phe Tyr
Ala Asp Ser Val Lys1 5 10 15Gly5017PRTHomo sapiens 50Trp Met Asn
Pro Asn Ser Gly Asn Thr Gly Phe Ala Gln Lys Phe Gln1 5 10
15Gly5118PRTHomo sapiens 51Gly Ile Ile Asn Pro Ser Gly Gly Ser Thr
Ser Tyr Ala Gln Lys Phe1 5 10 15Gln Gly5217PRTHomo sapiens 52Ile
Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe Gln1 5 10
15Gly5317PRTHomo sapiens 53Gly Ile Tyr Pro Ile Phe Gly Ser Ala Asn
Tyr Ala Gln Lys Phe Gln1 5 10 15Gly5417PRTHomo sapiens 54Gly Ile
Ile Pro Ile Phe Gly Ser Ala His Tyr Ala Gln Lys Phe Gln1 5 10
15Gly5517PRTHomo sapiens 55Gly Ile Ile Pro Ile Phe Gly Ser Ala His
Tyr Ser Gln Lys Phe Gln1 5 10 15Gly5610PRTHomo sapiens 56Asp Arg
Leu Glu Ser Ala Ala Phe Asp Tyr1 5 105713PRTHomo sapiens 57Asp Arg
Ala Asn Thr Asp Phe Ser Tyr Gly Met Asp Val1 5 105813PRTHomo
sapiens 58Asp Arg Ala Val Thr Asp Tyr Tyr Tyr Gly Met Asp Val1 5
105913PRTHomo sapiens 59Asp Arg Ala Asn Thr Asp Tyr Ser Phe Gly Met
Asp Val1 5 106013PRTHomo sapiens 60Asp Arg Ala Asn Thr Asp Tyr Ser
Leu Gly Met Asp Val1 5 106114PRTHomo sapiens 61Asp Val Val Pro Ala
Ala Ile Asp Tyr Tyr Gly Met Asp Val1 5 106210PRTHomo sapiens 62His
Lys Arg Gly Ser Asp Ala Phe Asp Ile1 5 106313PRTHomo sapiens 63Glu
Tyr Tyr Tyr Asp Ser Ser Glu Trp Ala Phe Asp Ile1 5 1064125PRTHomo
sapiens 64Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ser Thr Tyr 20 25 30Ala Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Gly Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala
Thr Tyr Tyr Ala Asp 50 55 60Ser Val Lys Asp Arg Phe Thr Ile Ser Arg
Asp Asp Ser Lys Asn Ser65 70 75 80Leu Tyr Leu Gln Met Asn Ser Leu
Lys Thr Glu Asp Thr Ala Val Tyr 85 90 95Tyr Cys Ala Arg His Gly Asn
Phe Gly Asn Ser Tyr Val Ser Tyr Phe 100 105 110Ala Tyr Trp Gly Gln
Gly Thr Leu Val Thr Val Ser Ser 115 120 12565125PRTHomo sapiens
65Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Thr
Tyr 20 25 30Ala Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu
Trp Val 35 40 45Gly Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala Thr Tyr
Tyr Ala Asp 50 55 60Ser Val Lys Asp Arg Phe Thr Ile Ser Arg Asp Asp
Ser Lys Asn Ser65 70 75 80Leu Tyr Leu Gln Met Asn Ser Leu Lys Thr
Glu Asp Thr Ala Val Tyr 85 90 95Tyr Cys Ala Arg His Gly Asn Phe Gly
Asn Ser Tyr Val Ser Trp Phe 100 105 110Ala Tyr Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Ser 115 120 12566125PRTHomo sapiens 66Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser
Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Thr Tyr 20 25
30Ala Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Gly Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala Thr Tyr Tyr Ala
Asp 50 55 60Ser Val Lys Asp Arg Phe Thr Ile Ser Arg Asp Asp Ser Lys
Asn Ser65 70 75 80Leu Tyr Leu Gln Met Asn Ser Leu Lys Thr Glu Asp
Thr Ala Val Tyr 85 90 95Tyr Cys Ala Arg His Gly Asn Phe Gly Asn Ser
Tyr Val Ser Trp Phe 100 105 110Ala Tyr Trp Gly Gln Gly Thr Leu Val
Thr Val Ser Ser 115 120 12567125PRTHomo sapiens 67Glu Val Gln Leu
Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg
Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Thr Tyr 20 25 30Ala Met
Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Gly
Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala Thr Tyr Tyr Ala Asp 50 55
60Ser Val Lys Asp Arg Phe Thr Ile Ser Arg Asp Asp Ser Lys Asn Ser65
70 75 80Leu Tyr Leu Gln Met Asn Ser Leu Lys Thr Glu Asp Thr Ala Val
Tyr 85 90 95Tyr Cys Ala Arg His Gly Asn Phe Gly Asn Ser Tyr Val Ser
Trp Phe 100 105 110Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser
Ser 115 120 12568110PRTHomo sapiens 68Asp Ile Gln Met Thr Gln Ser
Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr
Cys Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn
Trp Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly
Gly Thr Asn Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly
Ser Leu Ile Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu
Gln Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90
95Asn Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
11069110PRTHomo sapiens 69Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser Gly Ser
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 11070110PRTHomo
sapiens 70Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser
Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser Ser Thr Gly Ala
Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln Gln Lys Pro Gly
Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn Lys Arg Ala Pro
Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile Gly Asp Lys Ala
Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu Asp Phe Ala Thr
Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp Val Phe Gly Gln
Gly Thr Lys Val Glu Ile Lys 100 105 11071110PRTHomo sapiens 71Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ser Ser Thr Gly Ala Val Thr Thr
20 25 30Ser Asn Tyr Ala Asn Trp Val Gln Gln Lys Pro Gly Lys Ala Pro
Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn Lys Arg Ala Pro Gly Val Pro
Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile Gly Thr Asp Phe Thr Leu Thr
Ile Ser Ser65 70 75 80Leu Gln Pro Glu Asp Phe Ala Thr Tyr Tyr Cys
Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp Val Phe Gly Gln Gly Thr Lys
Val Glu Ile Lys 100 105 110726PRTHomo sapiens 72Ser Thr Tyr Ala Met
Asn1 57319PRTHomo sapiens 73Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala
Thr Tyr Tyr Ala Asp Ser1 5 10 15Val Lys Asp7414PRTHomo sapiens
74His Gly Asn Phe Gly Asn Ser Tyr Val Ser Trp Phe Ala Tyr1 5
107514PRTHomo sapiens 75His Gly Asn Phe Gly Asn Ser Tyr Val Ser Tyr
Phe Ala Tyr1 5 10766PRTHomo sapiens 76Asn Thr Tyr Ala Met Asn1
5776PRTHomo sapiens 77Asn Thr Tyr Ala Met His1 5786PRTHomo sapiens
78Asn Lys Tyr Ala Met Asn1 57919PRTHomo sapiens 79Arg Ile Arg Asn
Lys Tyr Asn Asn Tyr Ala Thr Tyr Tyr Ala Asp Ser1 5 10 15Val Lys
Asp8019PRTHomo sapiens 80Arg Ile Arg Asn Lys Tyr Asn Asn Tyr Ala
Thr Glu Tyr Ala Asp Ser1 5 10 15Val Lys Asp8119PRTHomo sapiens
81Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala Thr Glu Tyr Ala Ala Ser1
5 10 15Val Lys Asp8219PRTHomo sapiens 82Arg Ile Arg Asn Lys Tyr Asn
Asn Tyr Ala Thr Glu Tyr Ala Ala Ser1 5 10 15Val Lys Asp8319PRTHomo
sapiens 83Arg Ile Arg Ser Lys Tyr Asn Asn Tyr Ala Thr Tyr Tyr Ala
Asp Ser1 5 10 15Val Lys Gly8419PRTHomo sapiens 84Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr Ala Thr Glu Tyr Ala Asp Ser1 5 10 15Val Lys
Ser8514PRTHomo sapiens 85His Gly Asn Phe Gly Asp Ser Tyr Val Ser
Trp Phe Ala Tyr1 5 108614PRTHomo sapiens 86His Gly Asn Phe Gly Asn
Thr Tyr Val Ser Trp Phe Ala Tyr1 5 108714PRTHomo sapiens 87His Gly
Asn Phe Gly Cys Ser Tyr Val Ser Trp Phe Ala Tyr1 5 108814PRTHomo
sapiens 88His Gly Asn Phe Gly Asn Ser Tyr Ile Ser Tyr Trp Ala Tyr1
5 108914PRTHomo sapiens 89His Gly Asn Phe Gly Asn Ser Tyr Val Ser
Phe Phe Ala Tyr1 5 109014PRTHomo sapiens 90Arg Ser Ser Thr Gly Ala
Val Thr Thr Ser Asn Tyr Ala Asn1 5 10917PRTHomo sapiens 91Gly Thr
Asn Lys Arg Ala Pro1 5927PRTHomo sapiens 92Ala Leu Trp Tyr Ser Asn
Leu1 593242PRTHomo sapiens 93Asp Pro Asn Phe Trp Leu Gln Val Gln
Glu Ser Val Thr Val Gln Glu1 5 10 15Gly Leu Cys Val Leu Val Pro Cys
Thr Phe Phe His Pro Ile Pro Tyr 20 25 30Tyr Asp Lys Asn Ser Pro Val
His Gly Tyr Trp Phe Arg Glu Gly Ala 35 40 45Ile Ile Ser Arg Asp Ser
Pro Val Ala Thr Asn Lys Leu Asp Gln Glu 50 55 60Val Gln Glu Glu Thr
Gln Gly Arg Phe Arg Leu Leu Gly Asp Pro Ser65 70 75 80Arg Asn Asn
Cys Ser Leu Ser Ile Val Asp Ala Arg Arg Arg Asp Asn 85 90 95Gly Ser
Tyr Phe Phe Arg Met Glu Arg Gly Ser Thr Lys Tyr Ser Tyr 100 105
110Lys Ser Pro Gln Leu Ser Val His Val Thr Asp Leu Thr His Arg Pro
115 120 125Lys Ile Leu Ile Pro Gly Thr Leu Glu Pro Gly His Ser Lys
Asn Leu 130 135 140Thr Cys Ser Val Ser Trp Ala Cys Glu Gln Gly Thr
Pro Pro Ile Phe145 150 155 160Ser Trp Leu Ser Ala Ala Pro Thr Ser
Leu Gly Pro Arg Thr Thr His 165 170 175Ser Ser Val Leu Ile Ile Thr
Pro Arg Pro Gln Asp His Gly Thr Asn 180 185 190Leu Thr Cys Gln Val
Lys Phe Ala Gly Ala Gly Val Thr Thr Glu Arg 195 200 205Thr Ile Gln
Leu Asn Val Thr Tyr Val Pro Gln Asn Pro Thr Thr Gly 210 215 220Ile
Phe Pro Gly Asp Gly Ser Gly Lys Gln Glu Thr Arg Ala Gly Val225 230
235 240Val His949PRTHomo sapiens 94Asp Gln Glu Val Gln Glu Glu Thr
Gln1 5956PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 95Gly Gly Ser Gly Gly Ser1 5964PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 96Gly
Gly Ser Gly1975PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 97Gly Gly Ser Gly Gly1
598494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 98Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser Ser
Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln Gln
Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn Lys
Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu Asp
Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp Val
Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser Gly
Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Phe Ser Tyr Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Ser
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp
Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu Tyr
Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val Tyr
Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val Ser
Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475 480Val
Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
49099494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 99Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser Ser
Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln Gln
Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn Lys
Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu Asp
Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp Val
Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser Gly
Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Val Thr Asp Tyr Tyr Tyr Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490100494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 100Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser Gly Ser
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Phe Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490101494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 101Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser Gly Ser
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Leu Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490102495PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 102Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser Gly Ser
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Phe Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr
245 250 255Pro Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser
Asn Ile 260 265 270Gly Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro
Gly Thr Ala Pro 275 280 285Lys Leu Leu Ile Tyr Gly Asn Asn Gln Arg
Pro Ser Gly Val Pro Asp 290 295 300Arg Phe Ser Gly Ser Lys Ser Gly
Thr Ser Ala Ser Leu Ala Ile Ser305 310 315 320Gly Leu Gln Ser Glu
Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp 325 330 335Asp Ser Leu
Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val 340 345 350Leu
Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly 355 360
365Gly Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser
370 375 380Gly Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln
Ala Pro385 390 395 400Gly Lys Gly Leu Glu Trp Val Gly Arg Ile Arg
Ser Lys Tyr Asn Asn 405 410 415Tyr Ala Thr Tyr Tyr Ala Asp Ser Val
Lys Asp Arg Phe Thr Ile Ser 420 425 430Arg Asp Asp Ser Lys Asn Ser
Leu Tyr Leu Gln Met Asn Ser Leu Lys 435 440 445Thr Glu Asp Thr Ala
Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly 450 455 460Asn Ser Tyr
Val Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val465 470 475
480Thr Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490 495103495PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 103Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp
Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly
Thr Asn Lys Arg Ala Pro Gly Val Pro Ala Arg 50 55 60Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln
Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn
Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105
110Ser Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
115 120 125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly
Tyr Thr 130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro
Asn Ser Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg
Val Thr Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met
Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr
Cys Ala Arg Asp Arg Ala Asn Thr Asp Tyr Ser Leu Gly 210 215 220Met
Asp Val Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230
235 240Ser Gly Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly
Thr 245 250 255Pro Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg
Ser Asn Ile 260 265 270Gly Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu
Pro Gly Thr Ala Pro 275 280 285Lys Leu Leu Ile Tyr Gly Asn Asn Gln
Arg Pro Ser Gly Val Pro Asp 290 295 300Arg Phe Ser Gly Ser Lys Ser
Gly Thr Ser Ala Ser Leu Ala Ile Ser305 310 315 320Gly Leu Gln Ser
Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp 325 330 335Asp Ser
Leu Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val 340 345
350Leu Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly
355 360
365Gly Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser
370 375 380Gly Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln
Ala Pro385 390 395 400Gly Lys Gly Leu Glu Trp Val Gly Arg Ile Arg
Ser Lys Tyr Asn Asn 405 410 415Tyr Ala Thr Tyr Tyr Ala Asp Ser Val
Lys Asp Arg Phe Thr Ile Ser 420 425 430Arg Asp Asp Ser Lys Asn Ser
Leu Tyr Leu Gln Met Asn Ser Leu Lys 435 440 445Thr Glu Asp Thr Ala
Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly 450 455 460Asn Ser Tyr
Val Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val465 470 475
480Thr Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490 495104491PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 104Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp
Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly
Thr Asn Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser
Leu Ile Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln
Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn
Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105
110Ser Gly Gly Ser Gln Val Gln Leu Gln Glu Ser Gly Gly Gly Val Val
115 120 125Gln Pro Gly Arg Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
Phe Ser 130 135 140Phe Ser Asn Tyr Gly Ile His Trp Val Arg Gln Ala
Pro Gly Lys Gly145 150 155 160Leu Glu Trp Val Ala Leu Ile Ser Tyr
Asp Gly Asn Lys Lys Phe Tyr 165 170 175Ala Asp Ser Val Lys Gly Arg
Phe Ala Ile Ser Arg Asp Thr Ser Lys 180 185 190Asn Thr Val Asp Leu
Gln Met Thr Ser Leu Arg Pro Glu Asp Thr Ala 195 200 205Val Tyr Tyr
Cys Ala Lys Asp Arg Leu Glu Ser Ala Ala Phe Asp Tyr 210 215 220Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly Ser Gly Gly225 230
235 240Ser Ser Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser Val Ala Pro
Gly 245 250 255Gln Thr Ala Met Ile Thr Cys Gly Gly Asn Asn Ile Gly
Ser Thr Thr 260 265 270Val His Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro Val Leu Val Val 275 280 285Tyr Asp Asp Asn Glu Arg Pro Ser Gly
Ile Pro Glu Arg Phe Ser Gly 290 295 300Ser Asn Ser Gly Ser Thr Ala
Thr Leu Thr Ile Asn Arg Val Glu Ala305 310 315 320Gly Asp Glu Ala
Asp Tyr Tyr Cys Gln Val Trp Asp Ser Gly Ser Asp 325 330 335His Val
Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly Gly Ser 340 345
350Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln
355 360 365Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe
Thr Phe 370 375 380Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala Pro
Gly Lys Gly Leu385 390 395 400Glu Trp Val Gly Arg Ile Arg Ser Lys
Tyr Asn Asn Tyr Ala Thr Tyr 405 410 415Tyr Ala Asp Ser Val Lys Asp
Arg Phe Thr Ile Ser Arg Asp Asp Ser 420 425 430Lys Asn Ser Leu Tyr
Leu Gln Met Asn Ser Leu Lys Thr Glu Asp Thr 435 440 445Ala Val Tyr
Tyr Cys Ala Arg His Gly Asn Phe Gly Asn Ser Tyr Val 450 455 460Ser
Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser465 470
475 480Ala Ala Ala Gly Ser His His His His His His 485
490105496PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 105Gln Ser Val Leu Thr Gln Pro Pro Ser Ala
Ser Gly Thr Pro Gly Gln1 5 10 15Arg Val Thr Ile Ser Cys Ser Gly Ser
Arg Ser Asn Ile Gly Ser Asn 20 25 30Thr Val Asn Trp Tyr Gln Gln Leu
Pro Gly Thr Ala Pro Lys Leu Leu 35 40 45Ile Tyr Gly Asn Asn Gln Arg
Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55 60Gly Ser Lys Ser Gly Ser
Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70 75 80Ser Glu Asp Glu
Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp Ser Leu 85 90 95Ile Gly Trp
Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly Gly 100 105 110Ser
Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val 115 120
125Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr
130 135 140Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala Pro Gly
Lys Gly145 150 155 160Leu Glu Trp Val Gly Arg Ile Arg Ser Lys Tyr
Asn Asn Tyr Ala Thr 165 170 175Tyr Tyr Ala Asp Ser Val Lys Asp Arg
Phe Thr Ile Ser Arg Asp Asp 180 185 190Ser Lys Asn Ser Leu Tyr Leu
Gln Met Asn Ser Leu Lys Thr Glu Asp 195 200 205Thr Ala Val Tyr Tyr
Cys Ala Arg His Gly Asn Phe Gly Asn Ser Tyr 210 215 220Val Ser Tyr
Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser225 230 235
240Ser Gly Gly Ser Gly Gly Ser Asp Ile Gln Met Thr Gln Ser Pro Ser
245 250 255Ser Leu Ser Ala Ser Val Gly Asp Arg Val Thr Ile Thr Cys
Arg Ser 260 265 270Ser Thr Gly Ala Val Thr Thr Ser Asn Tyr Ala Asn
Trp Val Gln Gln 275 280 285Lys Pro Gly Lys Ala Pro Lys Ala Leu Ile
Gly Gly Thr Asn Lys Arg 290 295 300Ala Pro Gly Val Pro Ser Arg Phe
Ser Gly Ser Leu Ile Gly Asp Lys305 310 315 320Ala Thr Leu Thr Ile
Ser Ser Leu Gln Pro Glu Asp Phe Ala Thr Tyr 325 330 335Tyr Cys Ala
Leu Trp Tyr Ser Asn Leu Trp Val Phe Gly Gln Gly Thr 340 345 350Lys
Val Glu Ile Lys Gly Gly Ser Gly Gly Ser Gln Val Gln Leu Val 355 360
365Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala Ser Val Lys Val Ser
370 375 380Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr Asp Ile Asn
Trp Val385 390 395 400Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
Gly Trp Met Asn Pro 405 410 415Asn Ser Gly Asn Thr Gly Phe Ala Gln
Lys Phe Gln Gly Arg Val Thr 420 425 430Met Thr Arg Asp Thr Ser Thr
Ser Thr Val Tyr Met Glu Leu Ser Ser 435 440 445Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys Ala Arg Asp Arg Ala 450 455 460Asn Thr Asp
Phe Ser Tyr Gly Met Asp Val Trp Gly Gln Gly Thr Leu465 470 475
480Val Thr Val Ser Ser Ala Ala Ala Gly Ser His His His His His His
485 490 495106494PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 106Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp
Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly
Thr Asn Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser
Leu Ile Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln
Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn
Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105
110Ser Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
115 120 125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly
Tyr Thr 130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro
Asn Ser Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg
Val Thr Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met
Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr
Cys Ala Arg Asp Arg Ala Asn Thr Asp Phe Ser Tyr Gly 210 215 220Met
Asp Val Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230
235 240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr
Pro 245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser
Asn Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro
Gly Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg
Pro Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly
Ser Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu
Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu
Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345
350Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly
355 360 365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly 370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg
Gln Ala Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile
Arg Ser Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser
Val Lys Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn
Ser Leu Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr
Ala Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser
Tyr Val Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470
475 480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490107494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 107Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Phe Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490108494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 108Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Leu Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly
Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490109494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 109Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Val Thr Asp Tyr Tyr Tyr Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490110483PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 110Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Val Thr Asp Tyr Tyr Tyr Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser111494PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 111Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp
Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly
Thr Asn Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser
Leu Ile Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln
Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn
Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105
110Ser Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
115 120 125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly
Tyr Thr 130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro
Asn Ser Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg
Val Thr Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met
Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr
Cys Ala Arg Asp Arg Ala Asn Thr Asp Phe Ser Tyr Gly 210 215 220Met
Asp Val Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230
235 240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr
Pro 245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser
Asn Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro
Gly Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg
Pro Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly
Ser Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu
Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu
Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345
350Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly
355 360 365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly 370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg
Gln Ala Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile
Arg Ser Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser
Val Lys Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn
Ser Leu Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr
Ala Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser
Tyr Val Ser Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470
475 480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490112494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 112Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Phe Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg Ser Asn
Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser Leu Ile
Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490113494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 113Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Asp Ile Asn Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Trp Met Asn Pro Asn Ser
Gly Asn Thr Gly Phe 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Arg Ala Asn Thr Asp Tyr Ser Leu Gly 210 215 220Met Asp Val
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Arg
Ser Asn Ile Gly 260 265 270Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu
Pro Gly Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Gly Asn Asn Gln
Arg Pro Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser
Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser
Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp Asp 325 330 335Ser
Leu Ile Gly Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345
350Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly
355 360 365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly 370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg
Gln Ala Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile
Arg Ser Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser
Val Lys Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn
Ser Leu Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr
Ala Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser
Tyr Val Ser Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470
475 480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490114492PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 114Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Glu Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser
130 135 140Phe Thr Ser Tyr Trp Ile Gly Trp Val Arg Gln Met Pro Gly
Lys Gly145 150 155 160Leu Glu Trp Met Gly Ile Ile Tyr Pro Gly Asp
Ser Asp Thr Arg Tyr 165 170 175Ser Pro Ser Phe Gln Gly Gln Val Thr
Ile Ser Ala Asp Lys Ser Ile 180 185 190Ser Thr Ala Tyr Leu Gln Trp
Ser Ser Leu Lys Ala Ser Asp Thr Ala 195 200 205Met Tyr Tyr Cys Ala
Arg His Lys Arg Gly Ser Asp Ala Phe Asp Ile 210 215 220Trp Gly Gln
Gly Thr Thr Val Thr Val Ser Ser Gly Gly Ser Gly Gln225 230 235
240Ser Val Leu Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Gly Gln Arg
245 250 255Val Thr Ile Ser Cys Thr Gly Ser Ser Ser Asn Ile Gly Ala
Gly Tyr 260 265 270Asp Val His Trp Tyr Gln Gln Leu Pro Gly Thr Ala
Pro Lys Leu Leu 275 280 285Ile Tyr Gly Asn Ser Asn Arg Pro Ser Gly
Val Pro Asp Arg Phe Ser 290 295 300Gly Ser Lys Ser Gly Thr Ser Ala
Ser Leu Ala Ile Thr Gly Leu Gln305 310 315 320Ala Glu Asp Glu Ala
Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser Ser Leu 325 330 335Ser Asp Val
Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly Gly 340 345 350Ser
Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val 355 360
365Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr
370 375 380Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala Pro Gly
Lys Gly385 390 395 400Leu Glu Trp Val Gly Arg Ile Arg Ser Lys Tyr
Asn Asn Tyr Ala Thr 405 410 415Tyr Tyr Ala Asp Ser Val Lys Asp Arg
Phe Thr Ile Ser Arg Asp Asp 420 425 430Ser Lys Asn Ser Leu Tyr Leu
Gln Met Asn Ser Leu Lys Thr Glu Asp 435 440 445Thr Ala Val Tyr Tyr
Cys Ala Arg His Gly Asn Phe Gly Asn Ser Tyr 450 455 460Val Ser Tyr
Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser465 470 475
480Ser Ala Ala Ala Gly Ser His His His His His His 485
490115492PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 115Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Thr Tyr 20 25 30Ala Met Asn Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Gly Arg Ile Arg Ser Lys Tyr
Asn Asn Tyr Ala Thr Tyr Tyr Ala Asp 50 55 60Ser Val Lys Asp Arg Phe
Thr Ile Ser Arg Asp Asp Ser Lys Asn Ser65 70 75 80Leu Tyr Leu Gln
Met Asn Ser Leu Lys Thr Glu Asp Thr Ala Val Tyr 85 90 95Tyr Cys Ala
Arg His Gly Asn Phe Gly Asn Ser Tyr Val Ser Tyr Phe 100 105 110Ala
Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly Ser 115 120
125Gly Gly Ser Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser Gly Ala
130 135 140Pro Gly Gln Arg Val Thr Ile Ser Cys Thr Gly Ser Ser Ser
Asn Ile145 150 155 160Gly Ala Gly Tyr Asp Val His Trp Tyr Gln Gln
Leu Pro Gly Thr Ala 165 170 175Pro Lys Leu Leu Ile Tyr Gly Asn Ser
Asn Arg Pro Ser Gly Val Pro 180 185 190Asp Arg Phe Ser Gly Ser Lys
Ser Gly Thr Ser Ala Ser Leu Ala Ile 195 200 205Thr Gly Leu Gln Ala
Glu Asp Glu Ala Asp Tyr Tyr Cys Gln Ser Tyr 210 215 220Asp Ser Ser
Leu Ser Asp Val Val Phe Gly Gly Gly Thr Lys Leu Thr225 230 235
240Val Leu Gly Gly Ser Gly Gln Val Gln Leu Val Gln Ser Gly Ala Glu
245 250 255Val Lys Lys Pro Gly Glu Ser Leu Lys Ile Ser Cys Lys Gly
Ser Gly 260 265 270Tyr Ser Phe Thr Ser Tyr Trp Ile Gly Trp Val Arg
Gln Met Pro Gly 275 280 285Lys Gly Leu Glu Trp Met Gly Ile Ile Tyr
Pro Gly Asp Ser Asp Thr 290 295 300Arg Tyr Ser Pro Ser Phe Gln Gly
Gln Val Thr Ile Ser Ala Asp Lys305 310 315 320Ser Ile Ser Thr Ala
Tyr Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp 325 330 335Thr Ala Met
Tyr Tyr Cys Ala Arg His Lys Arg Gly Ser Asp Ala Phe 340 345 350Asp
Ile Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Gly Gly Ser 355 360
365Gly Gly Ser Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala
370 375 380Ser Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ser Ser Thr
Gly Ala385 390 395 400Val Thr Thr Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys 405 410 415Ala Pro Lys Ala Leu Ile Gly Gly Thr
Asn Lys Arg Ala Pro Gly Val 420 425 430Pro Ser Arg Phe Ser Gly Ser
Leu Ile Gly Asp Lys Ala Thr Leu Thr 435 440 445Ile Ser Ser Leu Gln
Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu 450 455 460Trp Tyr Ser
Asn Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile465 470 475
480Lys Ala Ala Ala Gly Ser His His His His His His 485
490116495PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 116Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
130 135 140Phe Thr Ser Tyr Tyr Met His Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Ile Ile Asn Pro Ser Gly
Gly Ser Thr Ser Tyr 165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr
Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Asp Val Val Pro Ala Ala Ile Asp Tyr Tyr 210 215 220Gly Met Asp
Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Gly225 230 235
240Gly Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr
245 250 255Pro Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Ser Ser
Asn Ile 260 265 270Gly Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu Pro
Gly Thr Ala Pro 275 280 285Lys Leu Leu Ile Tyr Ser Asp Asn Gln Arg
Pro Ser Gly Val Pro Asp 290 295 300Arg Phe Ser Gly Ser Lys Ser Gly
Ser Ser Ala Ser Leu Ala Ile Ser305 310 315 320Gly Leu Gln Ser Asp
Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp 325 330 335Asp Ser Leu
Asn Gly Ala Val Phe Gly Gly Gly Thr Lys Leu Thr Val 340 345 350Leu
Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly 355 360
365Gly Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser
370 375 380Gly Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln
Ala Pro385 390 395 400Gly Lys Gly Leu Glu Trp Val Gly Arg Ile Arg
Ser Lys Tyr Asn Asn 405 410 415Tyr Ala Thr Tyr Tyr Ala Asp Ser Val
Lys Asp Arg Phe Thr Ile Ser 420 425 430Arg Asp Asp Ser Lys Asn Ser
Leu Tyr Leu Gln Met Asn Ser Leu Lys 435 440 445Thr Glu Asp Thr Ala
Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly 450 455 460Asn Ser Tyr
Val Ser Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val465 470 475
480Thr Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
490 495117484PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 117Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp
Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Gly Leu Ile Gly Gly
Thr Asn Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser
Leu Ile Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln
Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn
Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105
110Ser Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
115 120 125Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly
Tyr Thr 130 135 140Phe Thr Ser Tyr Tyr Met His Trp Val Arg Gln Ala
Pro Gly Gln Gly145 150 155 160Leu Glu Trp Met Gly Ile Ile Asn Pro
Ser Gly Gly Ser Thr Ser Tyr 165 170 175Ala Gln Lys Phe Gln Gly Arg
Val Thr Met Thr Arg Asp Thr Ser Thr 180 185 190Ser Thr Val Tyr Met
Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr
Cys Ala Arg Asp Val Val Pro Ala Ala Ile Asp Tyr Tyr 210 215 220Gly
Met Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Gly225 230
235 240Gly Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly
Thr 245 250 255Pro Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Ser
Ser Asn Ile 260 265 270Gly Ser Asn Thr Val Asn Trp Tyr Gln Gln Leu
Pro Gly Thr Ala Pro 275 280 285Lys Leu Leu Ile Tyr Ser Asp Asn Gln
Arg Pro Ser Gly Val Pro Asp 290 295 300Arg Phe Ser Gly Ser Lys Ser
Gly Ser Ser Ala Ser Leu Ala Ile Ser305 310 315 320Gly Leu Gln Ser
Asp Asp Glu Ala Asp Tyr Tyr Cys Ala Thr Trp Asp 325 330 335Asp Ser
Leu Asn Gly Ala Val Phe Gly Gly Gly Thr Lys Leu Thr Val 340 345
350Leu Gly Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly
355 360 365Gly Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala
Ala Ser 370 375 380Gly Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val
Arg Gln Ala Pro385 390 395 400Gly Lys Gly Leu Glu Trp Val Gly Arg
Ile Arg Ser Lys Tyr Asn Asn 405 410 415Tyr Ala Thr Tyr Tyr Ala Asp
Ser Val Lys Asp Arg Phe Thr Ile Ser 420 425 430Arg Asp Asp Ser Lys
Asn Ser Leu Tyr Leu Gln Met Asn Ser Leu Lys 435 440 445Thr Glu Asp
Thr Ala Val Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly 450 455 460Asn
Ser Tyr Val Ser Trp Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val465 470
475 480Thr Val Ser Ser118494PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 118Asp Ile Gln Met Thr
Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr
Ile Thr Cys Arg Ser Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr
Ala Asn Trp Val Gln Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu
Ile Gly Gly Thr Asn Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe
Ser Gly Ser Leu Ile Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75
80Leu Gln Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser
85 90 95Asn Leu Trp Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly
Gly 100 105 110Ser Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys 115 120 125Lys Pro Gly Ser Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr 130 135 140Phe Ser Ser Tyr Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly145 150 155 160Leu Glu Trp Met Gly Gly
Ile Tyr Pro Ile Phe Gly Ser Ala Asn Tyr 165 170 175Ala Gln Lys Phe
Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr 180 185 190Ser Thr
Ala Tyr Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200
205Val Tyr Tyr Cys Ala Arg Glu Tyr Tyr Tyr Asp Ser
Ser Glu Trp Ala 210 215 220Phe Asp Ile Trp Gly Gln Gly Thr Leu Val
Thr Val Ser Ser Gly Gly225 230 235 240Ser Gly Gln Ser Val Leu Thr
Gln Pro Pro Ser Ala Ser Gly Thr Pro 245 250 255Gly Gln Arg Val Thr
Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly 260 265 270Ser Asn Ile
Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys 275 280 285Leu
Leu Ile Tyr Ser Asn Asn Gln Arg Pro Ser Gly Val Pro Asp Arg 290 295
300Phe Ser Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser
Gly305 310 315 320Leu Gln Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala
Ala Trp Asp Asp 325 330 335Ser Leu Lys Gly Tyr Val Phe Gly Gly Gly
Thr Lys Leu Thr Val Leu 340 345 350Gly Gly Ser Gly Gly Ser Glu Val
Gln Leu Val Glu Ser Gly Gly Gly 355 360 365Leu Val Gln Pro Gly Gly
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly 370 375 380Phe Thr Phe Ser
Thr Tyr Ala Met Asn Trp Val Arg Gln Ala Pro Gly385 390 395 400Lys
Gly Leu Glu Trp Val Gly Arg Ile Arg Ser Lys Tyr Asn Asn Tyr 405 410
415Ala Thr Tyr Tyr Ala Asp Ser Val Lys Asp Arg Phe Thr Ile Ser Arg
420 425 430Asp Asp Ser Lys Asn Ser Leu Tyr Leu Gln Met Asn Ser Leu
Lys Thr 435 440 445Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg His Gly
Asn Phe Gly Asn 450 455 460Ser Tyr Val Ser Tyr Phe Ala Tyr Trp Gly
Gln Gly Thr Leu Val Thr465 470 475 480Val Ser Ser Ala Ala Ala Gly
Ser His His His His His His 485 490119494PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
119Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser Ser Thr Gly Ala Val Thr
Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln Gln Lys Pro Gly Lys Ala
Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn Lys Arg Ala Pro Gly Val
Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile Gly Asp Lys Ala Thr Leu
Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu Asp Phe Ala Thr Tyr Tyr
Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp Val Phe Gly Gln Gly Thr
Lys Val Glu Ile Lys Gly Gly 100 105 110Ser Gly Gly Ser Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys 115 120 125Lys Pro Gly Ser Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr 130 135 140Phe Ser Ser
Tyr Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly145 150 155
160Leu Glu Trp Met Gly Gly Ile Ile Pro Ile Phe Gly Ser Ala His Tyr
165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr 180 185 190Ser Thr Ala Tyr Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala Arg Glu Tyr Tyr Tyr
Asp Ser Ser Glu Trp Ala 210 215 220Phe Asp Ile Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Ser Gly Gly225 230 235 240Ser Gly Gln Ser Val
Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro 245 250 255Gly Gln Arg
Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly 260 265 270Ser
Asn Thr Val Lys Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys 275 280
285Leu Leu Ile Tyr Ser Asn Asn Gln Arg Ser Ser Gly Val Pro Asp Arg
290 295 300Phe Ser Gly Ser Lys Ser Gly Ser Ser Ala Ser Leu Ala Ile
Ser Gly305 310 315 320Leu Gln Ser Glu Asp Glu Ala Asp Tyr Tyr Cys
Ala Ala Trp Asp Asp 325 330 335Ser Leu Asn Gly Tyr Val Phe Gly Gly
Gly Thr Lys Leu Thr Val Leu 340 345 350Gly Gly Ser Gly Gly Ser Glu
Val Gln Leu Val Glu Ser Gly Gly Gly 355 360 365Leu Val Gln Pro Gly
Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly 370 375 380Phe Thr Phe
Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala Pro Gly385 390 395
400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser Lys Tyr Asn Asn Tyr
405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys Asp Arg Phe Thr Ile
Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu Tyr Leu Gln Met Asn
Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg
His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val Ser Tyr Phe Ala Tyr
Trp Gly Gln Gly Thr Leu Val Thr465 470 475 480Val Ser Ser Ala Ala
Ala Gly Ser His His His His His His 485 490120483PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
120Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser Ser Thr Gly Ala Val Thr
Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln Gln Lys Pro Gly Lys Ala
Pro Lys 35 40 45Gly Leu Ile Gly Gly Thr Asn Lys Arg Ala Pro Gly Val
Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile Gly Thr Asp Phe Thr Leu
Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu Asp Phe Ala Thr Tyr Tyr
Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp Val Phe Gly Gln Gly Thr
Lys Val Glu Ile Lys Gly Gly 100 105 110Ser Gly Gly Ser Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys 115 120 125Lys Pro Gly Ser Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr 130 135 140Phe Ser Ser
Tyr Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly145 150 155
160Leu Glu Trp Met Gly Gly Ile Ile Pro Ile Phe Gly Ser Ala His Tyr
165 170 175Ala Gln Lys Phe Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr 180 185 190Ser Thr Ala Tyr Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala Arg Glu Tyr Tyr Tyr
Asp Ser Ser Glu Trp Ala 210 215 220Phe Asp Ile Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Ser Gly Gly225 230 235 240Ser Gly Gln Ser Val
Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro 245 250 255Gly Gln Arg
Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly 260 265 270Ser
Asn Thr Val Lys Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys 275 280
285Leu Leu Ile Tyr Ser Asn Asn Gln Arg Ser Ser Gly Val Pro Asp Arg
290 295 300Phe Ser Gly Ser Lys Ser Gly Ser Ser Ala Ser Leu Ala Ile
Ser Gly305 310 315 320Leu Gln Ser Glu Asp Glu Ala Asp Tyr Tyr Cys
Ala Ala Trp Asp Asp 325 330 335Ser Leu Asn Gly Tyr Val Phe Gly Gly
Gly Thr Lys Leu Thr Val Leu 340 345 350Gly Gly Ser Gly Gly Ser Glu
Val Gln Leu Val Glu Ser Gly Gly Gly 355 360 365Leu Val Gln Pro Gly
Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly 370 375 380Phe Thr Phe
Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala Pro Gly385 390 395
400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser Lys Tyr Asn Asn Tyr
405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys Asp Arg Phe Thr Ile
Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu Tyr Leu Gln Met Asn
Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg
His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val Ser Trp Phe Ala Tyr
Trp Gly Gln Gly Thr Leu Val Thr465 470 475 480Val Ser
Ser121494PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 121Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ser
Ser Thr Gly Ala Val Thr Thr 20 25 30Ser Asn Tyr Ala Asn Trp Val Gln
Gln Lys Pro Gly Lys Ala Pro Lys 35 40 45Ala Leu Ile Gly Gly Thr Asn
Lys Arg Ala Pro Gly Val Pro Ser Arg 50 55 60Phe Ser Gly Ser Leu Ile
Gly Asp Lys Ala Thr Leu Thr Ile Ser Ser65 70 75 80Leu Gln Pro Glu
Asp Phe Ala Thr Tyr Tyr Cys Ala Leu Trp Tyr Ser 85 90 95Asn Leu Trp
Val Phe Gly Gln Gly Thr Lys Val Glu Ile Lys Gly Gly 100 105 110Ser
Gly Gly Ser Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys 115 120
125Lys Pro Gly Ser Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr
130 135 140Phe Asp Ser Tyr Ala Ile Ser Trp Val Arg Gln Ala Pro Gly
Gln Gly145 150 155 160Leu Glu Trp Met Gly Gly Ile Ile Pro Ile Phe
Gly Ser Ala His Tyr 165 170 175Ser Gln Lys Phe Gln Gly Arg Val Thr
Ile Thr Ala Asp Glu Ser Thr 180 185 190Ser Thr Ala Tyr Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala 195 200 205Val Tyr Tyr Cys Ala
Arg Glu Tyr Tyr Tyr Asp Ser Ser Glu Trp Ala 210 215 220Phe Asp Ile
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly Gly225 230 235
240Ser Gly Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro
245 250 255Gly Gln Arg Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn
Ile Gly 260 265 270Asp Asn Val Val Asn Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys 275 280 285Leu Leu Ile Tyr Ser Thr Asn Lys Arg Pro
Ser Gly Val Pro Asp Arg 290 295 300Phe Ser Gly Ser Lys Ser Gly Ser
Ser Ala Ser Leu Ala Ile Ser Gly305 310 315 320Leu Gln Ser Glu Asp
Glu Ala Asp Tyr Tyr Cys Ala Ala Trp Asp Asp 325 330 335Ser Leu Ser
Ala Tyr Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 340 345 350Gly
Gly Ser Gly Gly Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly 355 360
365Leu Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
370 375 380Phe Thr Phe Ser Thr Tyr Ala Met Asn Trp Val Arg Gln Ala
Pro Gly385 390 395 400Lys Gly Leu Glu Trp Val Gly Arg Ile Arg Ser
Lys Tyr Asn Asn Tyr 405 410 415Ala Thr Tyr Tyr Ala Asp Ser Val Lys
Asp Arg Phe Thr Ile Ser Arg 420 425 430Asp Asp Ser Lys Asn Ser Leu
Tyr Leu Gln Met Asn Ser Leu Lys Thr 435 440 445Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg His Gly Asn Phe Gly Asn 450 455 460Ser Tyr Val
Ser Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr465 470 475
480Val Ser Ser Ala Ala Ala Gly Ser His His His His His His 485
4901226PRTArtificial SequenceDescription of Artificial Sequence
Synthetic 6xHis tag 122His His His His His His1 5
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016
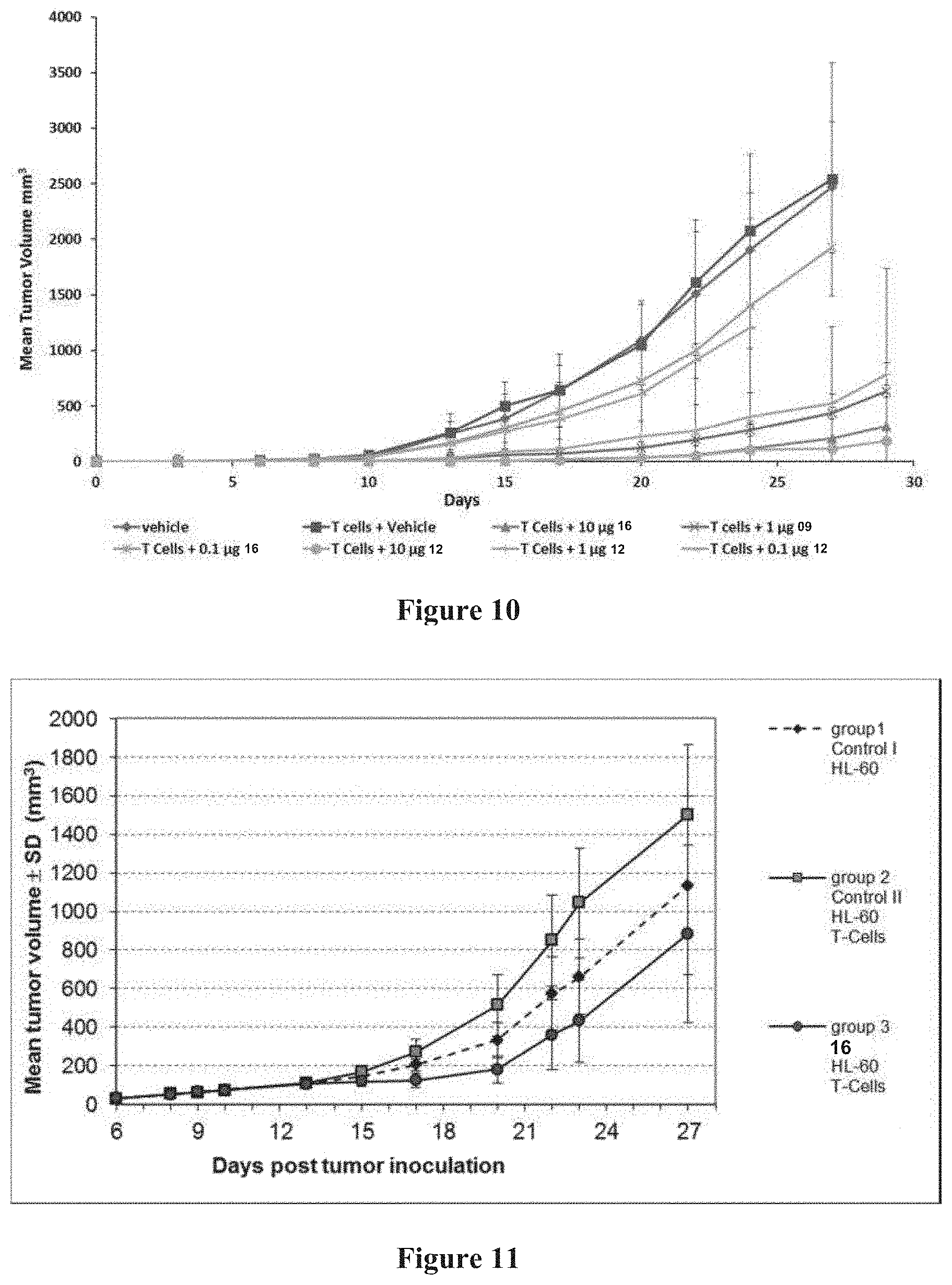
D00017

D00018

D00019
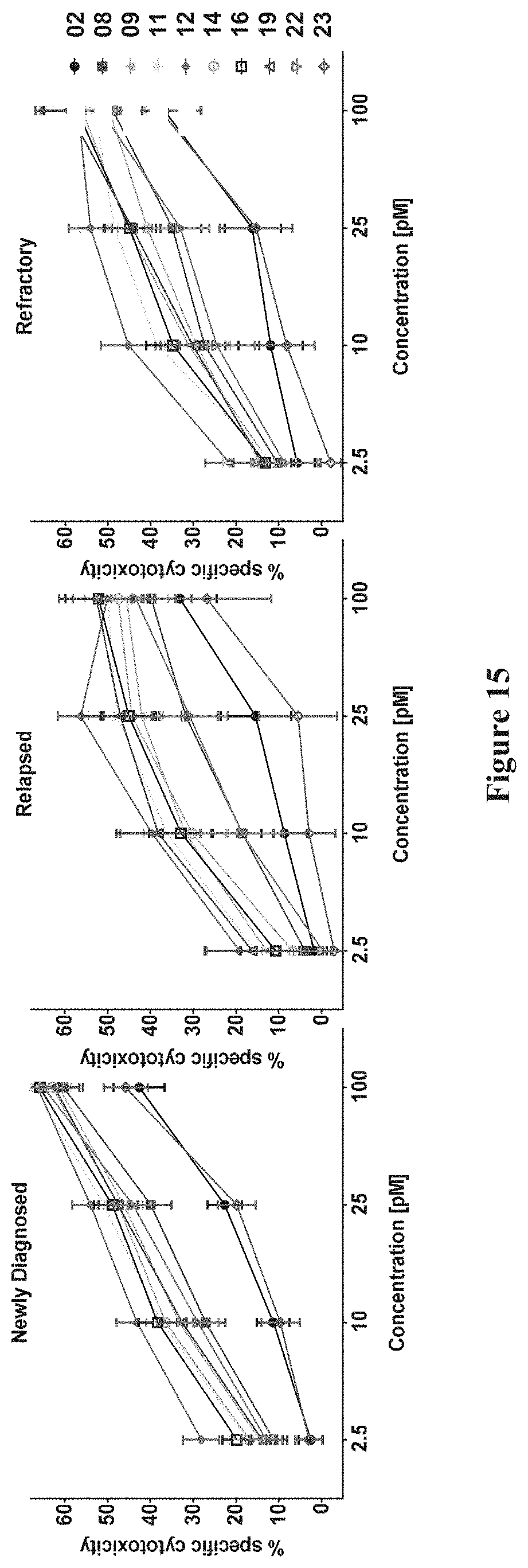
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.