Combinatorial Antibody Libraries And Uses Thereof
SMIDER; Vaughn ; et al.
U.S. patent application number 17/002749 was filed with the patent office on 2020-12-24 for combinatorial antibody libraries and uses thereof. This patent application is currently assigned to Taurus Biosciences, LLC. The applicant listed for this patent is Taurus Biosciences, LLC. Invention is credited to Omar BAZIRGAN, Tyson CHASE, James GRAZIANO, Helen Hongyuan MAO, Vaughn SMIDER, Byeong Doo SONG.
| Application Number | 20200399359 17/002749 |
| Document ID | / |
| Family ID | 1000005076690 |
| Filed Date | 2020-12-24 |





View All Diagrams
| United States Patent Application | 20200399359 |
| Kind Code | A1 |
| SMIDER; Vaughn ; et al. | December 24, 2020 |
COMBINATORIAL ANTIBODY LIBRARIES AND USES THEREOF
Abstract
Methods for making a combinatorial antibody library from human germline segments are provided. Also provided are libraries of nucleic acid molecules compiled from germline segments encoding VL chains and libraries of nucleic acid molecules encoding VH chains, and resulting antibody libraries. The libraries are provided as addressable libraries. Methods for screening antibody libraries against a target protein antigen, and the identified or selected antibodies are provided.
| Inventors: | SMIDER; Vaughn; (San Diego, CA) ; GRAZIANO; James; (San Diego, CA) ; MAO; Helen Hongyuan; (San Diego, CA) ; SONG; Byeong Doo; (Chuncheon, KR) ; CHASE; Tyson; (San Diego, CA) ; BAZIRGAN; Omar; (San Diego, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Taurus Biosciences, LLC San Diego CA |
||||||||||
| Family ID: | 1000005076690 | ||||||||||
| Appl. No.: | 17/002749 | ||||||||||
| Filed: | August 25, 2020 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14959940 | Dec 4, 2015 | 10774138 | ||
| 17002749 | ||||
| 13128219 | May 6, 2011 | 9221902 | ||
| PCT/US2009/063299 | Nov 4, 2009 | |||
| 14959940 | ||||
| 61211204 | Mar 25, 2009 | |||
| 61198764 | Nov 7, 2008 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2317/14 20130101; C07K 16/22 20130101; C07K 16/2863 20130101; C07K 2317/55 20130101; C40B 40/10 20130101; C40B 50/08 20130101; C07K 16/2887 20130101; C12N 15/1037 20130101; C07K 16/18 20130101; C07K 16/28 20130101; C07K 16/2884 20130101; C40B 40/08 20130101; C07K 2317/515 20130101; C07K 16/00 20130101; C07K 2317/56 20130101; C07K 16/32 20130101; C07K 2317/73 20130101; C07K 2317/92 20130101; C07K 2317/51 20130101; G01N 33/6845 20130101; C07K 16/2869 20130101; C07K 2317/74 20130101; C07K 2317/76 20130101; C07K 2317/24 20130101 |
| International Class: | C07K 16/22 20060101 C07K016/22; C40B 50/08 20060101 C40B050/08; C07K 16/00 20060101 C07K016/00; C07K 16/18 20060101 C07K016/18; C07K 16/28 20060101 C07K016/28; C40B 40/08 20060101 C40B040/08; C07K 16/32 20060101 C07K016/32; C12N 15/10 20060101 C12N015/10; G01N 33/68 20060101 G01N033/68 |
Claims
1. A combinatorial human antibody library, wherein the library is an addressable library and is selected from the group consisting of: A. a combinatorial human antibody addressable library comprising a plurality of antibody or antigen-binding antibody fragments, wherein each member in the library is a functional antibody or functional antigen-binding antibody fragment, and: a) each antibody or antigen-binding antibody fragment contains a variable light (VL) chain and a variable heavy (VH) chain or a sufficient portion thereof to form an antigen binding site; wherein: i) each VL chain is encoded by a nucleic acid molecule that comprises a V.sub..kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof, or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; and ii) each VH chain is encoded by a nucleic acid molecule that comprises a human V.sub.H and a human J.sub.H germline segment and any sequence of nucleotides between the V.sub.H and J.sub.H germline segments, wherein the segments are linked in-frame without a stop codon; and b) the library comprises at least about or 50 or 100 or more different antibody or antigen-binding antibody fragments; B. a combinatorial human antibody addressable library comprising a plurality of antibody or antigen-binding antibody fragments, wherein each member in the library is a functional antibody or functional antigen-binding antibody fragment, and: a) each antibody or antigen-binding antibody fragment contains a modified variable light (VL) chain and/or a modified variable heavy chain (VH) chain or a sufficient portion thereof to form an antigen binding site; wherein: i) each VL chain is encoded by a nucleic acid molecule that comprises a V.kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; ii) each VH chain is encoded by a nucleic acid molecule that comprises a V.sub.H, D.sub.H and a J.sub.H human germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; and b) the VL chain and/or VH chain is/are modified by replacement or insertion of at least one amino acid into at least one complementarity determining region (CDR); and c) the library comprises at least about or 50 or 100 or more different antibody or antigen-binding antibody fragments.
2. The combinatorial human antibody library according to claim 1, wherein: the sequence of nucleotides between the V.sub.H and J.sub.H germline segments encodes a peptide mimetic; or the amino acids that are inserted or replaced correspond to a peptide mimetic.
3. The combinatorial human antibody library according to claim 2, wherein: the peptide mimetic is selected from the group consisting of a peptide mimetic of TPO, EPO, G-CSF, IL-5, human brain natriuretic peptide (hBNP-32), exendin 4, GLP-1, GLP-2, glucagon, PACAP-38, CD209L, TNF, VEGF, MMP inhibitor, and CTLA-4; or the peptide mimetic has the sequence of amino acids selected from among any of SEQ ID NOS: 891 and 987-1014.
4. The combinatorial human antibody library according to claim 1, wherein the antibody or antigen-binding antibody fragment at each address is the same antibody or antigen-binding antibody fragment and is different from the antibody or antigen-binding antibody fragment at all other addresses.
5. The combinatorial human antibody library according to claim 1, wherein: all or a subset of germline V.sub.H segments are linked with all or a subset of D.sub.H segments which are linked with all or a subset of germline J.sub.H segments to generate the plurality of nucleic acid molecules encoding a VH chain; and all or a subset of germline V.sub..kappa. segments are linked with all or a subset of germline J.sub..kappa. segments, or all or a subset of germline V.sub..lamda. segments are linked to all or a subset of germline J.sub..lamda. segments to generate a plurality of nucleic acid molecules encoding a VL chain.
6. The combinatorial human antibody library according to claim 1, wherein each V.sub.H germline segment is selected from the group consisting of IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6, IGHV7, and an allelic variant of any of the foregoing; and/or each D.sub.H germline segment is selected from the group consisting of IGHD1, IGHD2, IGHD3, IGHD4, IGHD5, IGHD6, IGHD7, and an allelic variant of any of the foregoing; and/or each J.sub.H germline segment is selected from the group consisting of IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5, IGHJ6, and an allelic variant of any of the foregoing; and/or each V.sub..kappa. germline segment is selected from the group consisting of IGKV1, IGKV2, IGKV3, IGKV4, IGKV5, IGKV6, and an allelic variant of any of the foregoing; and/or each J.sub..kappa. germline segment is selected from the group consisting of IGKJ1, IGKJ2, IGKJ3, IGKJ4, IGKJ5, and an allelic variant of any of the foregoing and/or each V.sub..lamda. germline segment is selected from the group consisting of IGLV1, IGLV2, IGLV3, IGLV4, IGLV5, IGLV6, IGLV7, IGLV8, IGLV9, IGLV10, IGLV11, and an allelic variant of any of the foregoing; and/or each J.sub..lamda. germline segment selected from the group consisting of IGLJ1, IGLJ2, IGLJ3, IGLJ4, IGLJ5, IGLJ6, IGLJ7, and an allelic variant of any of the foregoing.
7. The combinatorial human antibody library according to claim 1, wherein the plurality of nucleic acid molecules encoding a VH chain and/or a VL chain is/are generated from a subset of germline segments selected based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group, or subgroup.
8. The combinatorial human antibody library according to claim 1 that comprises at or about or more than at or about 50, 10.sup.2, 10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9 different antibody or antigen-binding antibody fragments.
9. The combinatorial human antibody library of claim 1, wherein the CDR is CDRH3.
10. The combinatorial human antibody library of claim 1, wherein the sequence of nucleotides between the V.sub.H and J.sub.H germline segments is a DH germline segment or degenerate codons thereof or is an inverted D--H germline segment.
11. The combinatorial human antibody library of claim 1, wherein the antibodies or antigen-binding fragments are arranged in a spatial array.
12. The combinatorial human antibody library of claim 1, wherein the antibodies or antigen-binding fragments are immobilized on a solid support or are in solution.
13. The combinatorial human antibody library of claim 1, wherein the antibodies or antigen-binding fragments are identifiably labeled.
14. The combinatorial human antibody library of claim 6, wherein: each V.sub.H germline segment is selected from among any of SEQ ID NOS: 10-238; and/or each D.sub.H germline segment is selected from among any of SEQ ID NOS: 239-272; and/or each J.sub.H germline segment is selected from among any of SEQ ID NOS: 273-285; and/or each V.sub..kappa. germline segment is selected from among any of SEQ ID NOS: 286-355 and 868; and/or each J.sub..kappa. germline segment is selected from among any of SEQ ID NOS: 356-364; and/or each V.sub..lamda. germline segment is selected from among any of SEQ ID NOS: 365-441; and/or each J.sub..lamda. germline segment is selected from among any of SEQ ID NOS: 442-451.
15. The combinatorial human antibody library of claim 1, wherein the antibody or antigen-binding antibody fragments are full length antibodies.
16. The combinatorial human antibody library of claim 1, wherein the antibody or antigen-binding antibody fragments are antigen-binding antibody fragments.
17. The combinatorial human antibody library of claim 16, wherein the antigen-binding antibody fragment is a Fab.
18. A library of nucleic acid molecules that encodes the antibody and/or antigen-binding antibody fragments of a combinatorial human antibody library according to claim 1.
19. The library of nucleic acid molecules of claim 18, wherein the library of nucleic acid molecules are carried in vectors.
20. The library of nucleic acid molecules of claim 19, wherein the vector is an expression vector.
21. A library of nucleic acid molecules encoding a combinatorial human antibody library, wherein the encoded combinatorial human antibody library is an addressable library and comprises a plurality of antibody or antigen-binding antibody fragments, wherein each member in the library is a functional antibody or functional antigen-binding antibody fragment, and: a) each antibody or antigen-binding antibody fragment contains a variable light (VL) chain and a variable heavy (VH) chain or a sufficient portion thereof to form an antigen binding site; wherein: i) each VL chain is encoded by a nucleic acid molecule that comprises a V.sub..kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; ii) each VH chain is encoded by a nucleic acid molecule that comprises a human V.sub.H, D.sub.H, and a J.sub.H germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; and b) the library comprises at least about or 50 or 100 or more different antibody or antigen-binding antibody fragments.
22. The nucleic acid library of claim 21, wherein in the encoded combinatorial human antibody library the antibody or antigen-binding antibody fragment at each address is the same antibody or antigen-binding antibody fragment and is different from the antibody or antigen-binding antibody fragment at all other addresses.
23. The nucleic acid library according to claim 21, wherein in the encoded combinatorial human antibody library: all or a subset of germline V.sub.H segments are linked with all or a subset of D.sub.H segments which are linked with all or a subset of germline J.sub.H segments to generate the plurality of nucleic acid molecules encoding a VH chain; and all or a subset of germline V.sub..kappa. segments are linked with all or a subset of germline J.sub..kappa. segments, or all or a subset of germline V.sub..lamda. segments are linked to all or a subset of germline J.sub..lamda. segments to generate a plurality of nucleic acid molecules encoding a VL chain.
24. The nucleic acid library according to claim 21, wherein in the encoded combinatorial human antibody library: each V.sub.H germline segment is selected from the group consisting of IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6, IGHV7, and an allelic variant of any of the foregoing; and/or each D.sub.H germline segment is selected from the group consisting of IGHD1, IGHD2, IGHD3, IGHD4, IGHD5, IGHD6, IGHD7, and an allelic variant of any of the foregoing; and/or each J.sub.H germline segment is selected from the group consisting of IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5, IGHJ6, and an allelic variant of any of the foregoing; and/or each V.sub..kappa. germline segment is selected from the group consisting of IGKV1, IGKV2, IGKV3, IGKV4, IGKV5, IGKV6, and an allelic variant of any of the foregoing; and/or each J.sub..kappa. germline segment is selected from the group consisting of IGKJ1, IGKJ2, IGKJ3, IGKJ4, IGKJ5, and an allelic variant of any of the foregoing and/or each V.sub..lamda. germline segment is selected from the group consisting of IGLV1, IGLV2, IGLV3, IGLV4, IGLV5, IGLV6, IGLV7, IGLV8, IGLV9, IGLV10, IGLV11, and an allelic variant of any of the foregoing; and/or each J.sub..lamda. germline segment selected from the group consisting of IGLJ1, IGLJ2, IGLJ3, IGLJ4, IGLJ5, IGLJ6, IGLJ7, and an allelic variant of any of the foregoing.
25. The nucleic acid library of claim 21, wherein the encoded combinatorial human antibody library comprises at or about or more than at or about 50, 10.sup.2, 10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9 different antibodies or antigen-binding antibody fragments.
26. The nucleic acid library of claim 21, wherein in the encoded combinatorial human antibody library: each V.sub.H germline segment is selected from among any of SEQ ID NOS: 10-238; and/or each D.sub.H germline segment is selected from among any of SEQ ID NOS: 239-272; and/or each J.sub.H germline segment is selected from among any of SEQ ID NOS: 273-285; and/or each V.sub..kappa. germline segment is selected from among any of SEQ ID NOS: 286-355 and 868; and/or each J.sub..kappa. germline segment is selected from among any of SEQ ID NOS: 356-364; and/or each V.sub..lamda. germline segment is selected from among any of SEQ ID NOS: 365-441; and/or each J.sub..lamda. germline segment is selected from among any of SEQ ID NOS: 442-451.
27. The nucleic acid library of claim 21, wherein in the encoded combinatorial human antibody library each of the antibodies or antigen-binding antibody fragments are full length antibodies.
28. The nucleic acid library of claim 21, wherein in the encoded combinatorial human antibody library each of the antibody or antigen-binding antibody fragments are antigen-binding antibody fragments.
29. The nucleic acid library of claim 28, wherein the antigen-binding antibody fragment is a Fab.
30. A method of generating a combinatorial human antibody library, comprising: a) combining a V.sub.H, a D.sub.H, and a J.sub.H human germline segment or portion thereof in frame to generate a sequence of a nucleic acid molecule encoding a VH chain or a portion thereof; b) combining a V.sub..kappa. and a J.sub..kappa. human germline segment or portion thereof, or a V.sub..lamda. and a J.sub..lamda. human germline segment or portion thereof in frame to generate a sequence of a nucleic acid molecule encoding a VL chain or a portion thereof, wherein: in step a) and b) each of the portions of the V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. or J.sub..lamda. are sufficient to produce an antibody or portion thereof containing a VH or VL or portion thereof that forms a sufficient antigen binding site; c) repeating step a) and b) a plurality of times to generate sequences of a plurality of different nucleic acid molecules; d) synthesizing the nucleic acid molecules to produce two libraries, wherein: the first library comprises nucleic acid molecules encoding a VH chain or a portion thereof; and the second library comprises nucleic acid molecules encoding a VL chain or a portion thereof; e) expressing the nucleic acid molecules of the first library and the second library of part (d) and generating an antibody or antigen-binding antibody fragment formed by the VH chains and VL chains or portions thereof, thereby generating the combinatorial human antibody library.
31. The method of generating a combinatorial human antibody library of claim 30, wherein expressing the nucleic acid molecules comprises introducing a nucleic acid molecule from the first library and from the second library into a cell and growing the cells under conditions to express the antibodies or portions thereof in each cell, and repeating this a plurality of times to produce a library of cells, wherein each cell contains nucleic acid molecules encoding a different combination of VH and VL from every other cell in the library of cells.
32. The method of generating a combinatorial human antibody library of claim 30, wherein the library is an addressable library, wherein in step d) the synthesized nucleic acid sequences are individually addressed, thereby generating a first addressed nucleic acid library and a second addressed nucleic acid library.
33. The method of generating a combinatorial human antibody library of claim 30, wherein the expressed antibody is a full length antibody, or a fragment or portion thereof sufficient to form an antigen binding site.
34. The method of generating a combinatorial human antibody library of claim 30, wherein the expressed antibody is a Fab.
35. The method of generating a combinatorial human antibody library of claim 30, further purifying the antibodies or portions thereof.
36. A human combinatorial antibody library produced by the method of claim 30.
37. A method of screening a combinatorial human antibody library for binding or activity against a target protein, comprising: a) contacting one or more members of a combinatorial human antibody library according to claim 1 with a target protein, wherein the target protein is a membrane-bound protein, cell surface receptor (CSR), or a CSR ligand, a cytokine receptor, a receptor kinase, a receptor phosphatase, a receptor involved in cell-cell interactions, and a cellular adhesion molecule, wherein the target protein is selected from the group consisting of VEGFR-1, VEGFR-2, VEGFR-3, a epidermal growth factor receptor (EGFR), ErbB-2, ErbB-3, IGF-R1, C-Met, TNF-R1, TNF-R2, BTLA, HVEM, LT-13R, CD20, CD3, CD25, NOTCH, DLL4, G-CSF-R, GM-CSF-R, EPO-R, a cadherin, an integrin, CD52 and CD44, a VEGF-A, VEGF-B, VEGF-C, VEGF-D, PIGF, EGF, HGF, TNF-a, LIGHT, lymphotoxin (LT), IgE, G-CSF, GM-CSF, and EPO; and b) determining whether any member of the combinatorial human antibody library binds to or modulates a functional activity of the target protein, wherein the functional activity is selected from the group consisting of cellular proliferation, lymphoma apoptosis, chemotaxis, cancer cell invasion, matrigel, endothelial proliferation, tube formation, and signal transduction; wherein the method further comprises identifying the antibody or antigen-binding antibody fragment that binds the target protein.
38. A method of screening a combinatorial human antibody library for binding or activity against a target protein, comprising: a) contacting one or more members of a combinatorial human antibody library of claim 36 with a target protein, wherein the target protein is a membrane-bound protein, cell surface receptor (CSR), or a CSR ligand, a cytokine receptor, a receptor kinase, a receptor phosphatase, a receptor involved in cell-cell interactions, and a cellular adhesion molecule, wherein the target protein is selected from the group consisting of VEGFR-1, VEGFR-2, VEGFR-3, a epidermal growth factor receptor (EGFR), ErbB-2, ErbB-3, IGF-R1, C-Met, TNF-R1, TNF-R2, BTLA, HVEM, LT-13R, CD20, CD3, CD25, NOTCH, DLL4, G-CSF-R, GM-CSF-R, EPO-R, a cadherin, an integrin, CD52 and CD44, a VEGF-A, VEGF-B, VEGF-C, VEGF-D, PIGF, EGF, HGF, TNF-a, LIGHT, lymphotoxin (LT), IgE, G-CSF, GM-CSF, and EPO; and b) determining whether any member of the combinatorial human antibody library binds to or modulates a functional activity of the target protein, wherein the functional activity is selected from the group consisting of cellular proliferation, lymphoma apoptosis, chemotaxis, cancer cell invasion, matrigel, endothelial proliferation, tube formation, and signal transduction; wherein the method further comprises identifying the antibody or antigen-binding antibody fragment that binds the target protein.
39. A method of screening a combinatorial human antibody library for binding or activity against a target protein, comprising: 1) contacting one or more members of a combinatorial human antibody library with a target protein, wherein: the combinatorial human antibody library is an addressable library and comprises a plurality of antibody or antigen-binding antibody fragments, wherein each member in the library is a functional antibody or functional antigen-binding antibody fragment, and: a) each antibody or antigen-binding antibody fragment contains a variable light (VL) chain and a variable heavy (VH) chain or a sufficient portion thereof to form an antigen binding site; wherein: i) each VL chain is encoded by a nucleic acid molecule that comprises a V.sub..kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; ii) each VH chain is encoded by a nucleic acid molecule that comprises a human V.sub.H, D.sub.H, and a J.sub.H germline segment or degenerate codons thereof, wherein the segments are linked in-frame without a stop codon; and b) the library comprises at least about or 50 or 100 or more different antibody or antigen-binding antibody fragments; and the target protein is a membrane-bound protein, cell surface receptor (CSR), or a CSR ligand, a cytokine receptor, a receptor kinase, a receptor phosphatase, a receptor involved in cell-cell interactions, and a cellular adhesion molecule, wherein the target protein is selected from the group consisting of VEGFR-1, VEGFR-2, VEGFR-3, a epidermal growth factor receptor (EGFR), ErbB-2, ErbB-3, IGF-R1, C-Met, TNF-R1, TNF-R2, BTLA, HVEM, LT-13R, CD20, CD3, CD25, NOTCH, DLL4, G-CSF-R, GM-CSF-R, EPO-R, a cadherin, an integrin, CD52 and CD44, a VEGF-A, VEGF-B, VEGF-C, VEGF-D, PIGF, EGF, HGF, TNF-a, LIGHT, lymphotoxin (LT), IgE, G-CSF, GM-CSF, and EPO; and 2) determining whether any member of the combinatorial human antibody library binds to or modulates a functional activity of the target protein, wherein the functional activity is selected from the group consisting of cellular proliferation, lymphoma apoptosis, chemotaxis, cancer cell invasion, matrigel, endothelial proliferation, tube formation, and signal transduction; wherein the method further comprises identifying the antibody or antigen-binding antibody fragment that binds the target protein.
Description
RELATED APPLICATIONS
[0001] This application is a divisional of U.S. Non-Provisional application Ser. No. 14/959,940, entitled "Combinatorial Antibody Libraries And Uses Thereof," filed Dec. 4, 2015, which is a continuation of U.S. Non-Provisional application Ser. No. 13/128,219, entitled "Combinatorial Antibody Libraries And Uses Thereof," filed May 6, 2011, which is a U.S. National Stage Application under 35 U.S.C. .sctn. 371 of International Patent Application No. PCT/US2009/063299, entitled "Combinatorial Antibody Libraries And Uses Thereof," filed Nov. 4, 2009, which claims the benefit of priority to U.S. Provisional Application Ser. No. 61/198,764, entitled "Combinatorial Antibody Libraries and Uses Thereof," filed Nov. 7, 2008, and to U.S. Provisional Application Ser. No. 61/211,204, entitled "Combinatorial Antibody Libraries and Uses Thereof," filed Mar. 25, 2009, the entire contents of which are each incorporated herein by reference.
[0002] This application also is related to International PCT Application No. PCT/US2009/063303, entitled "Anti-DLL4 Antibodies and Uses Thereof," filed Nov. 4, 2009, which claims priority to U.S. Provisional Application No. 61/198,764 and to U.S. Provisional Application No. 61/211,204.
[0003] This application also is related to U.S. Provisional Application No. 61/280,618, entitled "Methods for Affinity-Maturation-Based Antibody Optimization," filed Nov. 4, 2009.
[0004] Where permitted, the subject matter of each of the above-noted related applications is incorporated by reference in its entirety.
INCORPORATION BY REFERENCE OF SEQUENCE LISTING PROVIDED ELECTRONICALLY
[0005] An electronic version of the Sequence Listing is filed herewith, the contents of which are incorporated by reference in their entirety. The electronic file was created on Aug. 25, 2020, is named 792472000110SeqList.TXT, and is 3,068,749 bytes in size.
FIELD OF THE INVENTION
[0006] Methods for making a combinatorial antibody library from human germline segments are provided. Also provided are libraries of nucleic acid molecules compiled from germline segments encoding VL chains and libraries of nucleic acid molecules encoding VH chains, and resulting antibody libraries. The libraries are provided as addressable libraries. Methods for screening antibody libraries against a target protein antigen, and the identified or selected antibodies are provided.
BACKGROUND
[0007] Numerous therapeutic and diagnostic monoclonal antibodies (MAbs) are used in the clinical setting to treat and diagnose human diseases, for example, cancer and autoimmune diseases. For example, exemplary therapeutic antibodies include Rituxan (Rituximab), Herceptin (Trastuzumab), Avastin (Bevacizumab) and Remicade (Infliximab). In designing antibody therapeutics, it is desirable to create antibodies, for example, antibodies that modulate a functional activity of a target, and/or improved antibodies such as antibodies with higher specificity and/or affinity and/or and antibodies that are more bioavailable, or stable or soluble in particular cellular or tissue environments.
[0008] Available techniques for generating antibody therapeutics are limited. Current methods include using antibody libraries to select variant proteins with desired properties in vitro. The libraries are generated to contain mutational diversity by targeted and non-targeted methods (e.g., Marks et al., J. Mol. Biol. (1991) 222, 581-597; Winters et al (1994) Annu Rev. Immunol. 12:433-55; Rosok et al (1996) J. Biol. Chem., 271:22611-22618; Kim et al. (2005) Mol. Cells 20:17-29; Mondon et al. (2008) Frontiers in Bioscience, 13:1117-1129; Benhar et al. (2007) Expert Opin. Biol. Ther., 7:763-779; and Knappik et al (2000) J. Mol. Biol., 296:57-86). Each of these antibody libraries has its limitations. Accordingly, it is among the objects herein is to provide methods for making antibody libraries, and antibodies produced by the methods.
SUMMARY
[0009] Provided herein are human combinatorial antibody libraries generated by rearrangement of human germline segments. Included among the combinatorial antibody libraries provided herein are libraries containing a plurality of antibodies, whereby each member antibody in the library contains a variable light (VL) chain and a variable heavy (VH) chain or a sufficient portion thereof to form an antigen binding site. Each VL chain of the antibodies in the library are encoded by a nucleic acid molecule that contains V.sub..kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof, or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, whereby the segments are linked in-frame. Each VH chain of the antibodies in the library are encoded by a nucleic acid molecule that contains a human V.sub.H and a human J.sub.H germline segment and any sequence of nucleotides between the V.sub.H and a J.sub.H germline segments, whereby the segments are linked in-frame. The human combinatorial antibody libraries contain at least about or 50 or 100 more different members. Each member in the library contains an antigen binding site and is a functional and productive antibody.
[0010] In such an example of a human combinatorial antibody library, the V.sub.H germline segment, the sequence of nucleotides between the V.sub.H and J.sub.H and the J.sub.H germline segment of the nucleic acid molecule encoding a VH chain are linked such that the V.sub.H segment is 5' to the sequence of nucleotides between the V.sub.H and J.sub.H which is 5' to the J.sub.H segment; and the V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. germline segments of the nucleic acid molecule encoding a VL chain are linked such that the V.sub..kappa. segment is 5' to the J.sub..kappa. segment or the V.sub..lamda. segment is 5' to the J.sub..lamda. segment. The sequence of nucleotides between the V.sub.H and J.sub.H germline segments is at or is about 5, 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100 nucleotides in length. In some examples, the sequence of nucleotides between the V.sub.H and J.sub.H germline segments encodes a peptide mimetic.
[0011] Also provided herein is a human combinatorial antibody library containing a plurality of antibodies, whereby each member antibody contains a modified variable light (VL) chain and/or a modified variable heavy chain (VH) chain or a sufficient portion thereof to form an antigen binding site. The VL chain in each library is encoded by a nucleic acid molecule that contains a V.sub..kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof, or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, whereby the segments are linked in-frame. Each VH chain of antibodies in the library are encoded by a nucleic acid molecule that contains a V.sub.H, D.sub.H and a J.sub.H human germline segment or degenerate codons thereof, whereby the segments are linked in-frame. The resultant protein of the VL chain and the VH chain are modified by amino acid replacement or insertion of amino acids into a CDR. The CDR can be any one or more up to all of a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 or a CDRL3, for example, a CDRH3. The amino acids that can be inserted or replaced correspond to a peptide mimetic.
[0012] In all of the examples above, the peptide mimetic can be a TPO, EPO, G-CSF, IL-5, human brain natriuretic peptide (hBNP-32), exendin 4, GLP-1, GLP-2, glucagon, PACAP-38, CD209L, TNF, VEGF, MMP inhibitor, or CTLA-4 peptide mimetic. In particular of a peptide mimietic is a mimetic that mimics Epo activiation of its receptor. The peptide mimetic further can include a flanking sequence at the carboxy and/ot N-terminal end, such as an amino acid or amino acids. For example, the flanking sequence can include glycine or a proline. Exemplary of peptide mimetics are any set forth in any of SEQ ID NOS: 891, and 987-1014.
[0013] In the libraries provided above, the nucleotides between the V.sub.H and J.sub.H germline segments is a human germline D.sub.H segment or degenerate codons thereof. Hence, a human combinatorial antibody library provided herein includes a library containing a plurality of antibodies, each member antibody contains a variable light (VL) chain and a variable heavy (VH) chain or a sufficient portion thereof to form an antigen binding site. Each VL chain of the antibodies in the library are encoded by a nucleic acid molecule that contains a V.sub..kappa. and a J.sub..kappa. human germline segment or degenerate codons thereof, or a V.sub..lamda. and a J.sub..lamda. human germline segment or degenerate codons thereof, whereby the segments are linked in-frame. Each VH chain of the antibodies in the library are encoded by nucleic acid molecule that contains a V.sub.H, D.sub.H and a J.sub.H germline segment, whereby the segments are linked in-frame. The human combinatorial antibody library contains at least about or 50 or 100 more different members. In such examples of a human combinatorial antibody library, the V.sub.H, D.sub.H and J.sub.H segments of the nucleic acid molecule encoding a VH chain are linked such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment; and the V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. germline segments of the nucleic acid molecule encoding a VL chain are linked such that the V.sub..kappa. segment is 5' to the J.sub..kappa. segment or the V.sub..lamda. segment is 5' to the J.sub..lamda. segment.
[0014] In all of the human combinatorial antibody libraries provided herein, the libraries can be provided as addressable libraries. In such addressable libraries, the antibody within each address is the same antibody and is different from the antibodies at all other addresses. For example, the addressable antibodies in the library are arranged in a spatial array. The spatial array can be a multiwall plate, wherein each individual locus of the plate corresponds to a different antibody member. The antibody members can be immobilized to the surface of the wells of the plate or can be present in solution. In another example, the addressable antibodies are attached to a solid support. In such an example, the solid support can be a filter, chip, slide, bead or cellulose, and the different antibody members are immobilized to the surface thereof. In some examples, the solid support is a biacore chip. In any of the addressable libraries provided herein, the members can be identifiably labeled. For example, the label can be colored, chromogenic, luminescent, chemical, fluorescent or electronic.
[0015] The human combinatorial libraries provided herein include members whereby a plurality of nucleic acid molecules encode a VH chain and a plurality of nucleic acid molecules encode a VL chain such that the plurality of nucleic acid molecules encoding the VH chain and the VL chain are sufficient to generate at least about or 50 or 100 different antibodies. Hence, in the libraries provided herein, in the nucleic acid molecules encoding a VH chain all or a subset of germline V.sub.H segments are linked with all or a subset of D.sub.H segments which are linked with all or a subset of germline J.sub.H segments to generate the plurality of nucleic acid molecules encoding a VH chain; and in the nucleic acid molecules encoding a VL chain all or a subset of germline V.sub..kappa. segments are linked with all or a subset of germline J.sub..kappa. segments, or all or a subset of germline V.sub..lamda. segments are linked to all or a subset of germline J.sub..lamda. segments to generate a plurality of nucleic acid molecules encoding a VL chain.
[0016] In the libraries provided herein, the VH chains in the members in the library are encoded by rearranged nucleic acid sequences combined by joining a human V.sub.H germline segment, a D.sub.H germline segment and a J.sub.H germline segment. The human V.sub.H germline segment is an IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6 or IGHV7 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS: 10-238. In examples of the combinatorial antibody libraries provided herein, the V.sub.H gene segment can have one nucleotide added or removed at the V-D joint of the nucleic acid molecule encoding the VH chain to maintain the reading frame of the VH chain. For example, a nucleotide is inserted at the 3' end of a V.sub.H nucleotide sequence to add a nucleotide between the V.sub.H and D.sub.H. The nucleotide can be any nucleotide, in particular, the nucleotide is a guanine (G). The D.sub.H segment is an IGHD1, IGHD2, IGHD3, IGHD4, IGHD5, IGHD6, or IGHD7 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS:239-272. In the examples of the combinatorial antibody libraries provided herein, the D.sub.H gene segment can have one nucleotide inserted or deleted at the V-D joint and/or the D-J joint of the nucleic acid encoding the VH chain to maintain the reading frame of the VH chain. The nucleotide insertion or deletion is chosen to maximize the hydrophilicity of the D.sub.H. For example, a nucleotide from the 5' end of a D.sub.H is deleted and/or a nucleotide from the 3' end of a D.sub.H is deleted. In another example, a nucleotide is inserted at the 3' end of a D.sub.H sequence to add a nucleotide between the D.sub.H and J.sub.H. The added nucleotide can be any nucleotide, in particular a guanine (G). The J.sub.H germline segment is an IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5 or IGHJ6 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS:273-285. In examples of the human combinatorial antibody library provided herein, the J.sub.H gene segment can have one or more nucleotides inserted or deleted at the D-J joint of the nucleic acid sequence encoding a VH chain to maintain the reading frame of the VH chain. For example, the nucleotide insertion or deletion is chosen to maintain the reading frame of the J.sub.H. In one examples, a nucleotide from the 5' end of the J.sub.H is deleted. In another example, a nucleotide from the 3' end of the J.sub.H is deleted.
[0017] In the libraries provided herein, the VL chains in the members in the library are encoded by rearranged nucleic acid sequences encoding a kappa light chain combined by joining a human V.sub..kappa. germline segment and a J.sub..kappa. germline segment or are encoded by rearranged nucleic acid sequences encoding a lambda light chain combined by joining a human V.sub..lamda. germline segment and a J.sub..lamda. germline segment. The human V.sub..kappa. is an IGKV1, IGKV2, IGKV3, IGKV4, IGKV5 and IGKV6, and genes or alleles thereof, for example, any set forth in any of SEQ ID NOS: 286-355 and 868. In examples of the combinatorial antibody libraries provided herein, the V.sub..kappa. gene segment can have one or more nucleotides inserted or deleted at the V-J joint of the nucleic acid molecule encoding the VL chain to maintain the reading frame of the VL chain. For example, a nucleotide is deleted at the 3' end of a V.sub..kappa.. In another example, a nucleotide is inserted at the 3' end of a of a V.sub..kappa. nucleotide sequence to add a nucleotide between the V.sub..kappa. and J.sub..kappa.. The nucleotide can be any nucleotide, in particular, the nucleotide is a guanine (G). The human J.sub..kappa. is an IGKJ1, IGKJ2, IGKJ3, IGKJ4 or IGKJ5 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS: 356-364. In examples of the combinatorial antibody libraries provided herein, the J.sub..kappa. gene segment has one or more nucleotides inserted or deleted at the V-J joint of the nucleic acid sequence encoding a VL chain to maintain the reading frame of the VL chain. The nucleotide insertion or deletion is chosen to maintain the reading frame of the J.sub..kappa. In some examples, a nucleotide from the 5' end of the J.sub..kappa. is deleted. The human V.sub..lamda. is an IGLV1, IGLV2, IGLV3, IGLV4, IGLV5, IGLV6, IGLV7, IGLV8, IGLV9, IGLV10 or IGLV11 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS:365-441. In examples of the combinatorial antibody libraries provided herein, the V.sub..lamda. has one or more nucleotides inserted or deleted at the V-J joint of the nucleic acid sequence encoding a VL to maintain the reading frame of the VL chain. For example, a nucleotide from the 3' end of a V.sub..lamda. nucleotide sequence is deleted. In another example, a nucleotide is inserted at the 3' end of a V.sub..lamda. nucleotide sequence to add a nucleotide between the V.sub..lamda. and J.sub..lamda.. The nucleotide can be any nucleotide, in particular a guanine (G). The human J.sub..lamda. is an IGLJ1, IGLJ2, IGLJ3, IGLJ4, IGLJ5, IGLJ6 or IGLJ7 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS:442-451. In examples of the combinatorial antibody library provided herein, the J.sub..lamda. has one nucleotide inserted or deleted at the V-J joint of the nucleic acid sequence encoding a VL chain to maintain the reading frame of the VL chain. The nucleotide insertion or deletion can be chosen to maintain the reading frame of the J.sub..lamda.. For example, a nucleotide from the 5' end of the J.sub..lamda. is deleted.
[0018] The human combinatorial antibody libraries contain a plurality of members each encoded by a plurality of nucleic acid molecules encoding a VH chain and a plurality of nucleic acid molecules encoding a VL chain. The plurality of nucleic acid molecules can correspond to all combinations or permutations of rearranged germline segments or a subset thereof. Generally, the libraries provided herein include libraries containing at or about 50, 10.sup.2, 10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9 or more different members. For example, libraries provided herein include those that contain 10.sup.3, 2.times.10.sup.3, 3.times.10.sup.3, 4.times.10.sup.3, 5.times.10.sup.3, 6.times.10.sup.3, 4.times.10.sup.3, 7.times.10.sup.3, 8.times.10.sup.3, 9.times.10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4 or more different members.
[0019] For example, the plurality of nucleic acid molecules encoding a VH chain are generated from a subset of germline segments selected based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group or subgroup. In one example, the plurality of nucleic acid molecules encoding a VH chain are generated from a subset of germline segments selected based on CDR and the CDR is CDR3. In another example, the plurality of nucleic acid molecules encoding a VH chain are selected based on gene family, whereby one germline segment from each of a V.sub.H, D.sub.H, and/or J.sub.H gene family is selected or one germline segment from a subset of a V.sub.H, D.sub.H, and/or J.sub.H gene family is selected. In such an example, the V.sub.H gene family is selected from among a IGHV1-18, IGHV1-2, IGHV1-24, IGHV1-3, IGHV1-45, IGHV1-46, IGHV1-58, IGHV1-69, IGHV1-8, IGHV2-26, IGHV2-5, IGHV2-70, IGHV3-11, IGHV3-13, IGHV3-15, IGHV3-16, IGHV3-20, IGHV3-21, IGHV3-23, IGHV3-30, IGHV3-33, IGHV3-35, IGHV3-38, IGHV3-43, IGHV3-48, IGHV3-49, IGHV3-53, IGHV3-64, IGHV3-66, IGHV3-7, IGHV3-72, IGHV3-73, IGHV3-74, IGHV3-9, IGHV4-28, IGHV4-31, IGHV4-34, IGHV4-39, IGHV4-4, IGHV4-59, IGHV4-61, IGHV5-51, IGHV6-1 and IGHV7-81; the D.sub.H gene family is selected from among a IGHD1-1, IGHD1-14, IGHD1-20, IGHD1-26, IGHD1-7, IGHD2-15, IGHD2-2, IGHD2-21, IGHD2-8, IGHD3-10, IGHD3- 16, IGHD3-22, IGHD3-3, IGHD3-9, IGHD4-11, IGHD4-17, IGHD4-23, IGHD4-4, IGHD5-12, IGHD5-18, IGHD5-24, IGHD5-5, IGHD6-13, IGHD6-19, IGHD6-25, IGHD6-6 and IGHD7-27; and the J.sub.H gene family is selected from among a IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5 and IGHJ6.
[0020] For example, the plurality of nucleic acid molecules encoding a VL chain are generated from a subset of germline segments selected based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group, subgroup. In one example, the plurality of nucleic acid molecules encoding a VL chain are generated from a subset of germline segments selected based on CDR and the CDR is CDR3. In another example, the plurality of nucleic acid molecules encoding a VH chain are selected based on gene family, whereby one germline segment from each of a V.sub..kappa. and/or J.sub..kappa. or V.sub..lamda. and/or J.sub..lamda. gene family is selected or one germline segment from a subset of a V.sub..kappa. and/or J.sub..kappa. or V.sub..lamda. and/or J.sub..lamda. gene family is selected. In such an example, the V.sub..kappa. gene family is selected from among a IGKV1-12, IGKV1-12, IGKV1-16, IGKV1-17, IGKV1-27, IGKV1-33, IGKV1-37, IGKV1-39, IGKV1-5, IGKV1-6, IGKV1-8, IGKV1-9, IGKV1-NL1, IGKV1/OR2, IGKV1D-12, IGKV1D-13, IGKV1D-16, IGKV1-D-17, IGKV1D-33, IGKV1D-37, IGKV1D-39, IGKV1D-42, IGKV1D-43, IGKV1D-8, IGKV2-24, IGKV2-28, IGKV2-29, IGKV2-30, IGKV2-30, IGKV2-40, IGKV2D-24, IGKV2D-26, IGKV2D-28, IGKV2D-29, IGKV2-D-30, IGKV2D-40, IGKV3-11, IGKV3-15, IGKV3-20, IGKV3-7, IGKV3-NL1, IGV3-NL2, IGKV3-NL3, IGKV3-NL4, IGKV3-NL5, IGKV3/OR2-268, IGKV3D-11, IGKV3D-15, IGKV3D-20, iGKV3D-7, IGKV4-1, IGKV5-2, IGKV6-21, IGKV6D-21, IGKV6D-41, and IGKV1-39; the J.sub..kappa. gene family is selected from among a IGKJ1, IGKJ2, IGKJ3, IGKJ4 and IGKJ5; the V.sub..lamda. gene family is selected from among a IGLV1-36, IGLV1-40, IGLV1-41, IGLV1-44, IGLV1-47, IGLV1-50, IGLV1-51, IGLV10-54, IGLV11-55, IGLV2-11, IGLV2-14, IGLV2-18, IGLV2-23, IGLV2-33, IGLV2-8, IGLV3-1, IGLV3-10, IGLV3-12, IGLV3-16, IGLV3-19, IGLV3-21, IGLV3-22, IGLV3-25, IGLV3-27, IGLV3-32, IGLV3-9, IGLV4-3, IGLV4-60, IGLV4-69, IGLV5-37, IGLV5-39, IGLV5-45, IGLV5-8, IGLV5-52, IGLV6-57, IGLV7- 43, IGLV7-46, IGLV8-61, IGLV8-61 and IGLV9-49; and the J.sub..lamda. gene family is selected from among a IGLJ1, IGLJ2, IGLJ4, IGLJ5, IGLJ6 and IGLJ7.
[0021] In any of the combinatorial antibody libraries provided herein, each antibody member in the library is productive and functional. Hence, in some examples, member antibodies in the library contains a VH chain and/or a VL chain that is encoded by a nucleic acid molecule that is modified to remove stop codons and/or restriction enzyme sites.
[0022] In any of the combinatorial antibody libraries provided herein, the VH chain is encoded by a nucleic acid molecule having a sequence of nucleotides set forth in any of SEQ ID NOS: 1059-1410, or a subset thereof and the VL chain is encoded by a nucleic acid molecule having a sequence of nucleotides set forth in any of SEQ ID NOS: 1411-1422, 1424-1439 and 1441-1471, or a subset thereof. The antibody libraries provided herein include libraries containing members whereby the VH chain has a sequence of amino acids set forth in any of SEQ ID NOS: 1475-1826 or a subset thereof and the VL chain has a sequence of amino acids set forth in any of SEQ ID NOS: 1827-1838, 1840-1855 and 1857-1888 or a subset thereof.
[0023] The human combinatorial antibody libraries provided herein include those having members that are full length antibodies or are fragments or portions thereof of antibodies, whereby the fragment or portion of the antibody is sufficient to form an antigen binding site. Thus, any of the combinatorial antibody libraries provided herein can further contain all of a portion of a constant region, such that the portion of a constant region is sufficient to permit association of a heavy and light chain. Included among fragments or portions of antibody members in the libraries provided herein are a Fab, Fab', F(ab').sub.2, single-chain Fvs (scFv), Fv, dsFv, diabody, Fd and Fd' fragments Fab fragments, Fd fragments, scFv fragments, or a scFab fragments. For example, combinatorial antibody libraries provided herein are Fab libraries, whereby antibody members of the library are Fabs.
[0024] Provided herein is a library of nucleic acid molecules containing a plurality of addressable nucleic acid molecules encoding a variable light (VL) chain. In such libraries, each VL chain is encoded by a nucleic acid molecule containing a V.sub..kappa. and a J.sub..kappa. human germline segment or a V.sub..lamda. and a J.sub..lamda. human germline segment linked in-frame, whereby the nucleic acid molecule within each address is the same and is different from the nucleic acid molecules at all other addresses. Each nucleic acid member are formed from combination of germline segments such that the V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. germline segments of the nucleic acid molecule encoding a VL chain are linked such that the V.sub..kappa. segment is 5' to the J.sub..kappa. segment or the V.sub..lamda. segment is 5' to the J.sub..lamda. segment. The library includes a plurality of nucleic acid members that can include all permutations of all combinations of germline segments. In some examples, the plurality of nucleic acid members includes a subset of all germline segments such that a subset of germline V.sub..kappa. segments are linked with all or a subset of germline J.sub..kappa. segments, or all or a subset of germline V.sub..lamda. segments are linked to all or a subset of germline J.sub..lamda. segments to generate a plurality of nucleic acid molecules encoding a VL chain.
[0025] In the VL nucleic acid libraries provided herein, nucleic acid molecules encoded a VL chain are generated by rearranged nucleic acid sequences combined by joining a human V.sub..kappa. germline segment and a J.sub..kappa. germline segment. The V.sub..kappa. is an IGKV1, IGKV2, IGKV3, IGKV4, IGKV5 or IGKV6, and genes and alleles thereof, for example any set forth in any of SEQ ID NOS:286-355 and 868. Included among the V.sub..kappa. germline segment contained in nucleic acid members in the libraries provided herein are any where the V.sub..kappa. has one or more nucleotides inserted or deleted a the V-J joint in the nucleic acid molecule encoding a VL to maintain the reading frame of the VL chain. For example, one or more nucleotides at the 3' end of the V.sub..kappa. nucleotide sequence is deleted. In other examples, one or more nucleotides is inserted at the 3' end of a V.sub..kappa. nucleotide sequence to add a nucleotide between the V.sub..kappa. and J.sub..kappa. germline segments. The nucleotide can be any nucleotide, and in particular is a guanine (G). The J.sub..kappa. germline segment is an IGKJ1, IGKJ2, IGKJ3, IGKJ4 and IGKJ5 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS: 356-364. Included among the J.sub..kappa. germline segment contained in nucleic acid members in the libraries provided herein are any where the J.sub..kappa. has one or more nucleotides inserted or deleted at the V-J joint to maintain the reading frame of the VL chain. The insertion or deletion is typically chosen to maintain the reading frame of the J K. For example, one or more nucleotides from the 5' end of the J.sub..kappa. is deleted.
[0026] In some examples, the VL nucleic acid libraries provided herein, nucleic acid molecules encoded a VL chain are generated by rearranged nucleic acid sequences combined by joining a human V.sub..lamda. germline segment and a J.sub..lamda. germline segment. The V.sub..lamda. germline segment is an IGLV1, IGLV2, IGLV3, IGLV4, IGLV5, IGLV6, IGLV7, IGLV8, IGLV9, IGLV10 and IGLV11 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS: 365-441. Included among the V.sub..lamda..theta.germline segment contained in nucleic acid members in the libraries provided herein are any where the V.sub..lamda. has one or more nucleotides inserted or deleted at the V-J joint of the nucleic acid molecule encoding a VL to maintain the reading frame of the VL chain. For example, one or more nucleotides is from the 3' end of a V.sub..lamda. nucleotide sequence is deleted. In another example, one or more nucleotides is inserted at the 3' end of a V.sub..lamda. nucleotide sequence to add a nucleotide between the V.sub..lamda. and J. The nucleotide can be any nucleotide, in particular a guanine (G). The J.sub..lamda. germline segment is an IGLJ1, IGLJ2, IGLJ3, IGLJ4, IGLJ5, IGLJ6 and IGLJ7 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS: 442-451. Included among J.sub..lamda. germline segments contained in nucleic acid members in the libraries provided herein are any where the J.sub..lamda. has one or more nucleotides inserted or deleted at the V-J join of the nucleic acid molecule encoding a VL to maintain the reading frame of the VL chain. The insertion or deletion is typically chosed to maintain the reading frame of the J.sub..lamda.. For example, a nucleotide from the 5' end of the J.sub..lamda. is deleted.
[0027] Any of the plurality of nucleic acid molecules encoding a VL chain in the libraries provided herein can be generated from a subset of germline segments selected based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group, subgroup. For example, the plurality of nucleic acid molecules encoding a VL chain are generated from a subset of germline segments selected based on CDR and the CDR is CDR3. In another example, the plurality of nucleic acid molecules encoding a VL chain are selected based on gene family, whereby one germline segment from each of a V.sub..kappa. and/or J.sub..kappa. or V.sub..lamda. and/or J.sub..lamda. gene family is selected or one germline segment from a subset of a V.sub..kappa. and/or J.sub..kappa. or V.sub..lamda. and/or J.sub..lamda. gene family is selected. In such an example, a V.sub..kappa. germline segment can include any one or more germline segments from a IGKV1-12, IGKV1-12, IGKV1-16, IGKV1-17, IGKV1-27, IGKV1-33, IGKV1-37, IGKV1-39, IGKV1-5, IGKV1-6, IGKV1-8, IGKV1-9, IGKV1-NL1, IGKV1/OR2, IGKV1D-12, IGKV1D-13, IGKV1D-16, IGKV1-D-17, IGKV1D-33, IGKV1D-37, IGKV1D-39, IGKV1D-42, IGKV1D-43, IGKV1D-8, IGKV2-24, IGKV2-28, IGKV2-29, IGKV2-30, IGKV2-30, IGKV2-40, IGKV2D-24, IGKV2D-26, IGKV2D-28, IGKV2D-29, IGKV2-D-30, IGKV2D-40, IGKV3-11, IGKV3-15, IGKV3-20, IGKV3-7, IGKV3-NL1, IGV3-NL2, IGKV3-NL3, IGKV3-NL4, IGKV3-NL5, IGKV3/OR2-268, IGKV3D-11, IGKV3D-15, IGKV3D-20, IGKV3D-7, IGKV4-1, IGKV5-2, IGKV6-21, IGKV6D-21, IGKV6D-41, or IGKV1-39 gene family; a J.sub..kappa. germline segments can include any one or more germline segments from a IGKJ1, IGKJ2, IGKJ3, IGKJ4 and IGKJ5 gene family; a V.sub..lamda. germline segment can include any one or more germline segments from a IGLV1-36, IGLV1-40, IGLV1-41, IGLV1-44, IGLV1-47, IGLV1-50, IGLV1-51, IGLV10-54, IGLV11-55, IGLV2-11, IGLV2-14, IGLV2-18, IGLV2-23, IGLV2-33, IGLV2-8, IGLV3-1, IGLV3-10, IGLV3-12, IGLV3-16, IGLV3-19, IGLV3-21, IGLV3-22, IGLV3-25, IGLV3-27, IGLV3-32, IGLV3-9, IGLV4-3, IGLV4-60, IGLV4-69, IGLV5-37, IGLV5-39, IGLV5-45, IGLV5-8, IGLV5-52, IGLV6-57, IGLV7-43, IGLV7-46, IGLV8-61, IGLV8-61 and IGLV9-49 gene family; and/or a J.sub..lamda. germline segment can include any one or more germline segments from a IGLJ1, IGLJ2, IGLJ4, IGLJ5, IGLJ6 and IGLJ7 gene family.
[0028] In all of the nucleic acid libraries encoding a VL chain provided herein, the nucleic acid molecule encoding a VL chain can be modified to remove stop codons and/or restriction enzyme sites. Exemplary of nucleic acid molecules in the libraries provided herein include any of SEQ ID NOS:1411-1422, 1424-1439 and 1441-1471, or a subset thereof.
[0029] Provided herein is a library of nucleic acid molecules containing a plurality of addressable nucleic acid molecules encoding a variable light (VH) chain. In such libraries, each VH chain is encoded by a nucleic acid molecule containing a V.sub.H, a D.sub.H and a J.sub.H human germline segment linked in-frame, whereby the nucleic acid molecule within each address is the same and is different from the nucleic acid molecules at all other addresses. Each nucleic acid member is formed from combination of germline segments such that the V.sub.H, a D.sub.H and a J.sub.H human germline segment of the nucleic acid molecule encoding a VH chain are linked such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment. The library includes a plurality of nucleic acid members that can include all permutations of all combinations of germline segments. In some examples, the plurality of nucleic acid members includes a subset of all germline segments such that a subset of germline V.sub.H segment are linked with all or a subset of germline D.sub.H segments which are linked with all or a subset of germline J.sub.H segments to generate the plurality of nucleic acid molecules encoding a VH chain.
[0030] In the VH nucleic acid libraries provided herein, nucleic acid molecules encoded a VH chain are generated by rearranged nucleic acid sequences combined by joining a human V.sub.H, D.sub.H and J.sub.H germline segments. The V.sub.H is an IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6 and IGHV7 and genes and alleles thereof, for example any set forth in any of SEQ ID NOS: 10-238. Included among the V.sub.H germline segment contained in nucleic acid members in the libraries provided herein are any where the V.sub.H has one or more nucleotides added or removed at the V-D joint in the nucleic acid molecule encoding a VH to maintain the reading frame of the VH chain. For example, one or more nucleotides at the 3' end of the V.sub.H nucleotide sequence is deleted. In other examples, one or more nucleotides is inserted or added at the 3' end of a V.sub.H nucleotide sequence to add a nucleotide between the V.sub.H and D.sub.H germline segments. The nucleotide can be any nucleotide, and in particular is a guanine (G). The D.sub.H germline segment is an IGHD1, IGHD2, IGHD3, IGHD4, IGHD5, IGHD6, and IGHD7 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS: 239-272. Included among the D.sub.H germline segment contained in nucleic acid members in the libraries provided herein are any where the D.sub.H has one or more nucleotides inserted or deleted at the V-D and/or the D-J joint to maintain the reading frame of the VH chain. The nucleotide insertion or deletion can be any nucleotide, but typically is chosen to maximize the hydrophilicity of the D.sub.H. For example, one or more nucleotides from the 5' end of the D.sub.H is deleted. In other examples, a nucleotide from the 3' end of a D.sub.H is deleted. In further examples, a nucleotide is inserted at the 3' end of a D.sub.H sequence to add a nucleotide been the D.sub.H and J.sub.H. The nucleotide can be any nucleotide, but typically is a guanine (G). The germline segment is an IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5, and IGHJ6 and genes and alleles thereof, for example, any set forth in any of SEQ ID NOS:273-285. Included among the J.sub.H germline segment contained in nucleic acid members in the libraries provided herein are any where the J.sub.H has one or more nucleotides inserted or deleted at the D-J joint to maintain the reading frame of the VH chain. Typically, the nucleotide insertion or deletion is chosen to maintain the reading frame of the J.sub.H. For example, one or more nucleotides from the 5' end of the J.sub.H is deleted. In another example, one or more nucleotides from the 3' end of the J.sub.H is deleted.
[0031] Any of the plurality of nucleic acid molecules encoding a VH chain in the libraries provided herein can be generated from a subset of germline segments selected based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group, subgroup. For example, the plurality of nucleic acid molecules encoding a VH chain are generated from a subset of germline segments selected based on CDR and the CDR is CDR3. In another example, the plurality of nucleic acid molecules encoding a VH chain are selected based on gene family, whereby one germline segment from each of a V.sub.H, D.sub.H, and/or J.sub.H gene family is selected or one germline segment from a subset of a V.sub.H, D.sub.H, and/or J.sub.H gene family is selected. In such an example, a V.sub.H germline segment can include any one or more germline segments from a IGHV1-18, IGHV1-2, IGHV1-24, IGHV1-3, IGHV1-45, IGHV1-46, IGHV1-58, IGHV1-69, IGHV1-8, IGHV2-26, IGHV2-5, IGHV2-70, IGHV3- 11, IGHV3-13, IGHV3-15, IGHV3-16, IGHV3-20, IGHV3-21, IGHV3-23, IGHV3-30, IGHV3-33, IGHV3-35, IGHV3-38, IGHV3-43, IGHV3-48, IGHV3-49, IGHV3-53, IGHV3-64, IGHV3-66, IGHV3-7, IGHV3-72, IGHV3-73, IGHV3-74, IGHV3-9, IGHV4-28, IGHV4-31, IGHV4-34, IGHV4-39, IGHV4-4, IGHV4-59, IGHV4-61, IGHV5-51, IGHV6-1 and IGHV7-81 gene family; a the D.sub.H germline segment can include any one ore more germline segments from a IGHD1-1, IGHD1-14, IGHD1-20, IGHD1-26, IGHD1-7, IGHD2-15, IGHD2-2, IGHD2-21, IGHD2-8, IGHD3-10, IGHD3-16, IGHD3-22, IGHD3-3, IGHD3-9, IGHD4-11, IGHD4-17, IGHD4-23, IGHD4-4, IGHD5-12, IGHD5-18, IGHD5-24, IGHD5-5, IGHD6-13, IGHD6-19, IGHD6-25, IGHD6-6 and IGHD7-27 gene family; and/or a J.sub.H germline segment can include any one or more germline segments from a IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5 and IGHJ6 gene family.
[0032] In all of the nucleic acid libraries encoding a VH chain provided herein, the nucleic acid molecule encoding a VH chain can be modified to remove stop codons and/or restriction enzyme sites. Exemplary of nucleic acid molecules in the libraries provided herein include any of SEQ ID NOS: 1059-1410, or a subset thereof.
[0033] Also provided herein are libraries of addressable vectors containing any of the above nucleic acid molecules encoding a variable light (VL) chain or a variable heavy (VH) chain. Also provided herein are addressable cells, whereby each cell in the library contains any of the above different vectors.
[0034] Also provided herein are a library of nucleic acid molecules containing a plurality of addressable nucleic acid molecules encoding a variable light (VL) chain and a plurality of addressable nucleic acid molecules encoding a variable heavy (VH) chain (i.e. paired nucleic acid libraries). In such libraries, each VL chain is encoded by a nucleic acid molecule that contains a V.sub..kappa. and a J.sub..kappa. human germline segments or V.sub..lamda. and J.sub..lamda. germline segments linked in-frame and each VH chain is encoded by a nucleic acid molecule that contains a V.sub.H, a D.sub.H and a J.sub.H human germline segment. The resulting nucleic acid members in the nucleic acid molecule encoding the VL chain can be any provided herein and the nucleic acid molecule encoding the VH chain can be any provided herein. In such addressable libraries, each locus contains a nucleic acid molecule encoding a VH chain and a nucleic acid molecule encoding a VL chain, such that the combination of VH nucleic acid molecules and VL nucleic acid molecules within each address is different from the combination of nucleic acid molecules at all other addresses, i.e. the pairs of nucleic acid libraries at each locus are different.
[0035] Provided herein is a method of generating human combinatorial antibody libraries. The method includes the steps of a) combining a V.sub.H, a D.sub.H and a J.sub.H human germline segment or portion thereof in frame to generate a sequence of a nucleic acid molecule encoding a VH chain or a portion thereof and b) combining a V.sub..kappa. and a J.sub..kappa. human germline segment or portion thereof, or a V.sub..lamda. and a J.sub..lamda. germline segment or portion thereof in frame to generate a sequence of a nucleic acid molecule encoding a VL chain or a portion thereof. In the method provided herein each of the portions of the V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. or J.sub..lamda. in step a) and b) are sufficient to produce an antibody or portion thereof containing a VH or VL or portion thereof that forms a sufficient antigen binding site. In the methods, steps a) and b) are repeated a plurality of times to generate sequences of a plurality of different nucleic acid molecules. The nucleic acid molecules are synthesized to produce two libraries, whereby the first library contains nucleic acid molecules encoding a VH chain or portion thereof and the second library contains nucleic acid molecules encoding a VL chain or a portion thereof. In the method, a nucleic acid molecule from the first library and from the second library are introduced into a cell (e.g. together such as by co-transformation). The step of introducing nucleic acids into cells is repeated a plurality of times with different pairs of nucleic acid molecules from the first library and the second library resulting in that contain nucleic acid molecules encoding a VH chain and a nucleic acid molecule encoding a VL chain such that the nucleic acid molecules encode a different combination of VH and VL chains from every other cell. The cells are grown to express the antibodies or portions thereof in each cell, thereby producing a plurality of antibodies or portion thereof. The plurality of produced antibody or portion thereof contains a VH and a VL or a sufficient portion thereof to form an antigen binding site and the antibodies or portions thereof are different from those at every other antibody or portions thereof in the library.
[0036] The human combinatorial library produced by the method provided herein can be provided as an addressable library. In such methods, each of the various steps can be performed in an addressed format so that throughout the steps of the method the identity of the germline segments, the recombined nucleic acid sequence and/or produced antibody or portion thereof are known by their address. For example, the synthesized nucleic acid sequences are individually addressed, thereby generating a first addressed nucleic acid library and a second addressed nucleic acid library. The nucleic acid molecules can be introduced into addressed cells, whereby each locus contains a cell that contains nucleic acid molecules encoding a different combination of a VH and a VL from every other cell in the addressed library of cells. Upon expression of the antibodies, addressed antibodies are produced whereby each locus contains an antibody containing a VH chain and a VL chain or a portion thereof sufficient to form an antigen binding site. The antibodies or portions thereof at each locus is the same and is different from those at each and every other locus. Hence, the identify of the antibody or portion thereof is known by its address.
[0037] In the method of generating a human combinatorial antibody library provided herein, in step a) the V.sub.H, D.sub.H and J.sub.H germline segments or portions thereof of the nucleic acid molecule encoding a VH chain are combined such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment; and in step b) the V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. germline segments or portions thereof of the nucleic acid molecule encoding a VL chain are linked such that the V.sub..kappa. segment is 5' to the J.sub..kappa. segment or the V.sub..lamda. segment is 5' to the J.sub..lamda. segment. Steps a) and/or b) can be performed manually or can be performed in silico, such as by a computer or computer system capable of or programmed to execute computer-readable instructions based on an algorithm for performing a method of combining human germline segments.
[0038] In the method provided herein for generating a human combinatorial antibody library, step a) includes the steps of selecting a V.sub.H, a D.sub.H and a J.sub.H germline segment or portion thereof, generating a V-D joint by modifying the germline sequence of the V.sub.H and/or D.sub.H germline segments by insertion or deletion of one or more nucleotides in order to maximize the hydrophilicity of the D.sub.H germline segment, generating a D-J joint by modifying the germline sequence of the D.sub.H and/or J.sub.H germline segments by insertion or deletion of one or more nucleotides to maintain the reading frame of the J.sub.H, and combining the resulting V.sub.H, D.sub.H, and J.sub.H, to generate a sequence of a nucleic acid molecule encoding a VH chain. In such a method, the V-D joint can be generated by deletion of one or more, for example one, nucleotide from the 5' end of the D.sub.H germline segment. In another example, the V-D joint can be generated by deleting one or more nucleotides from the 3' end of the V.sub.H germline segment. In a further example, the V-D joint can be generated by inserting one or more nucleotides at the 5' end of the D.sub.H germline segment. For example, the inserted or added nucleotide or nucleotides can be any nucleotide, and in particular is a guanine (G). Further, the D-J joint can be generated by deletion of one or more nucleotides from the 5' end of the J.sub.H. In another example, the D-J joint is generated by inserting a nucleotide from the 3' end of the D.sub.H germline segment. The inserted or added nucleotide or nucleotides can be any nucleotide, and in particular is a guanine (G).
[0039] In the method provided herein for generating a human combinatorial antibody library, step b) includes the steps of selecting V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. germline segment or portion thereof, generating a V-J joint by modifying the germline sequence of the V.sub..kappa. or J.sub..kappa. by insertion or deletion of one or more nucleotides to maintain the reading frame of the J.sub..kappa., or by modifying the germline sequence of the V.sub..lamda. or J.sub..lamda. by insertion or deletion of one or more nucleotides to maintain the reading frame of the J.sub..lamda., and combining the resulting V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. to generate a sequence of a nucleic acid molecule encoding a VL chain. In such a method, the V-J joint can be generated by deletion of one or more, for example one, nucleotide from the 5' end of the of the J.sub..kappa. or J.sub..lamda. germline segment. In another example, the V-J joint can be generated by deleting one or more nucleotides from the 3' end of the V.sub..kappa. or V.sub..lamda. germline segment. In a further example, the V-J joint can be generated by inserting one or more nucleotides at the 5' end of the J.sub..kappa. or J.sub..lamda.. germline segment. For example, the inserted or added nucleotide or nucleotides can be any nucleotide, and in particular is a guanine (G).
[0040] In the methods of generating a combinatorial library provided herein, steps a) and b) are repeated a plurality of times. Repeating step a) a plurality of times includes selecting N1 (i.e. a first number) of different V.sub.H germline segments, selecting N2 (i.e. a second number) of different D.sub.H germline segments and selecting N3 (a third number) of different J.sub.H sequences. The N1, N2 and N3 numbers can be the same or different, and can include all respective germline segments or a subset thereof. Generally, the N1, N2 and N3 are a number of germline segments that can be all or a subset of V.sub.H, D.sub.H or J.sub.H germline segments, respectively. In the method of repeating step a) a plurality of times, all possible combination of V.sub.H, D.sub.H and J.sub.H combinations are made to generate N1.times.N2.times.N3 different nucleic acid sequences encoding a VH chain.
[0041] For example, in the method of generating a human combinatorial antibody library provided herein, a V.sub.H germline segment (including N1 different V.sub.H germline segments) can be selected from all or a subset of an IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6 or IGHV7 and genes and alleles thereof, for example, a V.sub.H germline segment set forth in any of SEQ ID NOS: 10-238. A D.sub.H germline segment can be selected from all or a subset of an IGHD1, IGHD2, IGHD3, IGHD4, IGHD5, IGHD6, or IGHD7 and genes and alleles thereof, for example, a D.sub.H germline segment set forth in any of SEQ ID NOS: 239-272. A J.sub.H germline segment can be selected from all or a subset of IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5, or IGHJ6 and genes and alleles thereof, for example, a J.sub.H germline segment set forth in any of SEQ ID NOS: 273-285.
[0042] In any of the above examples, the method can include in step a) selecting a subset of germline segments based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group or subgroup. For example, the subset of germline segments can be selected based on gene family. In the methods, germline segments can be selected such that one germline segment from each of a V.sub.H, D.sub.H, and/or J.sub.H gene family is selected or one germline segment from a subset of a V.sub.H, D.sub.H, and/or J.sub.H gene family is selected. V.sub.H gene families include, but are not limited to, a IGHV1-18, IGHV1-2, IGHV1-24, IGHV1-3, IGHV1-45, IGHV1-46, IGHV1-58, IGHV1-69, IGHV1-8, IGHV2-26, IGHV2-5, IGHV2-70, IGHV3-11, IGHV3-13, IGHV3-15, IGHV3-16, IGHV3-20, IGHV3-21, IGHV3-23, IGHV3-30, IGHV3-33, IGHV3-35, IGHV3-38, IGHV3-43, IGHV3-48, IGHV3-49, IGHV3-53, IGHV3-64, IGHV3-66, IGHV3-7, IGHV3-72, IGHV3-73, IGHV3-74, IGHV3-9, IGHV4-28, IGHV4-31, IGHV4-34, IGHV4-39, IGHV4-4, IGHV4-59, IGHV4-61, IGHV5-51, IGHV6-1 and IGHV7-81 gene families including genes and alleles thereof. D.sub.H gene families include, but are not limited to, a IGHD1-1, IGHD1-14, IGHD1-20, IGHD1-26, IGHD1-7, IGHD2-15, IGHD2-2, IGHD2-21, IGHD2-8, IGHD3-10, IGHD3-16, IGHD3-22, IGHD3-3, IGHD3-9, IGHD4-11, IGHD4-17, IGHD4-23, IGHD4-4, IGHD5-12, IGHD5-18, IGHD5-24, IGHD5-5, IGHD6-13, IGHD6-19, IGHD6-25, IGHD6-6 and IGHD7-27 gene families including genes and alleles thereof. The J.sub.H gene families include, but are not limited to, a IGHJ1, IGHJ2, IGHJ3, IGHJ4, IGHJ5 and IGHJ6 gene families including genes and alleles thereof.
[0043] In the methods of generating a combinatorial library provided herein, steps a) and b) are repeated a plurality of times. Repeating step b) a plurality of times includes selecting N1 (i.e. a first number) of different V.sub..lamda. germline segments and selecting N2 (i.e. a second number) different J.sub..lamda. germline segments or selecting N3 (a third number) of different V.sub..kappa. germline segments and selecting N4 (i.e. a fourth number) of different J.sub..kappa. germline segments. The N1, N2, N3 and N4 numbers can be the same or different, and can include all respective germline segments or a subset thereof. Generally, the N1, N2, N3 and N4 are a number of germline segments that can be all or a subset of V.sub..lamda., J.sub..lamda. V.sub..kappa., J.sub..kappa. germline segments, respectively. In the method of repeating step b) a plurality of times, all possible combination of V.sub..lamda., J.sub..lamda., V.sub..kappa., J.sub..kappa. combinations are made to generate N1.times.N2 or N3.times.N4 different nucleic acid sequences encoding a VL chain.
[0044] For example, in the method of generating a human combinatorial antibody library provided herein, a V.sub..lamda. germline segment (including N1 different V.sub..lamda. germline segments) can be selected from all or a subset of an IGLV1, IGLV2, IGLV3, IGLV4, IGLV5, IGLV6, IGLV7, IGLV8, IGLV9, IGLV10 and IGLV11 and genes and alleles thereof, for example, a V.sub..lamda. germline segment set forth in any of SEQ ID NOS: 365-441. A J.sub..lamda. germline segment can be selected from all or a subset of an IGLJ1, IGLJ2, IGLJ3, IGLJ4, IGLJ5, IGLJ6 and IGLJ7 and genes and alleles thereof, for example, a J.sub..lamda. germline segment set forth in any of SEQ ID NOS: 442-451. A V.sub..kappa. germline segment can be selected from all or a subset of IGKV1, IGKV2, IGKV3, IGKV4, IGKV5 and IGKV6, and genes and alleles thereof, for example, a V.sub..kappa. germline segment set forth in any of SEQ ID NOS: 286-355 and 868. A J.sub..kappa. germline segment can be selected from all or a subset of a IGKJ1, IGKJ2, IGKJ3, IGKJ4 and IGKJ5 and genes and alleles thereof, for example, a J.sub..kappa. germline segment set forth in any of SEQ ID NOS: 356-364.
[0045] In any of the above examples, the method can include in step b) selecting a subset of germline segments based on sequence similarities or differences, gene family, length, composition, CDR length or composition, species, functionality, specificity, group or subgroup. For example, the subset of germline segments can be selected based on gene family. In the methods, germline segments can be selected such that one germline segment from each of a V.sub..kappa. and/or J.sub..kappa. or V.sub..lamda. and/or J.sub..lamda. gene family is selected or one germline segment from a subset of a V.sub..kappa. and/or J.sub..kappa. or V.sub..lamda. and/or J.sub..lamda. gene family is selected. V.sub..kappa. gene families include, but are not limited to, a IGKV1-12, IGKV1-12, IGKV1-16, IGKV1-17, IGKV1-27, IGKV1-33, IGKV1-37, IGKV1-39, IGKV1-5, IGKV1-6, IGKV1-8, IGKV1-9, IGKV1-NL1, IGKV1/OR2, IGKV1D-12, IGKV1D-13, IGKV1D-16, IGKV1-D-17, IGKV1D-33, IGKV1D-37, IGKV1D-39, IGKV1D-42, IGKV1D-43, IGKV1D-8, IGKV2-24, IGKV2-28, IGKV2-29, IGKV2-30, IGKV2-30, IGKV2-40, IGKV2D-24, IGKV2D-26, IGKV2D-28, IGKV2D-29, IGKV2-D-30, IGKV2D-40, IGKV3-11, IGKV3-15, IGKV3-20, IGKV3-7, IGKV3-NL1, IGV3-NL2, IGKV3-NL3, IGKV3-NL4, IGKV3-NL5, IGKV3/OR2-268, IGKV3D-11, IGKV3D-15, IGKV3D-20, IGKV3D-7, IGKV4-1, IGKV5-2, IGKV6-21, IGKV6D-21, IGKV6D-41, and IGKV1-39 gene families including genes and alleles thereof. V.sub..lamda. gene families include, but are not limited to, a IGLV1-36, IGLV1-40, IGLV1-41, IGLV1-44, IGLV1-47, IGLV1-50, IGLV1-51, IGLV10-54, IGLV11-55, IGLV2-11, IGLV2-14, IGLV2-18, IGLV2-23, IGLV2-33, IGLV2-8, IGLV3-1, IGLV3-10, IGLV3-12, IGLV3-16, IGLV3-19, IGLV3-21, IGLV3-22, IGLV3-25, IGLV3-27, IGLV3-32, IGLV3-9, IGLV4-3, IGLV4-60, IGLV4-69, IGLV5-37, IGLV5-39, IGLV5-45, IGLV5-8, IGLV5-52, IGLV6-57, IGLV7-43, IGLV7-46, IGLV8-61, IGLV8-61 and IGLV9-49 gene families including genes and alleles thereof. The J.sub..lamda. gene families include, but are not limited to, a IGLJ1, IGLJ2, IGLJ4, IGLJ5, IGLJ6 and IGLJ7 gene families including genes and alleles thereof.
[0046] In any of the methods above of generating a human combinatorial antibody library, the germline segments can be included in a user-created database, for example, to provide convenient access to such sequences. In practicing the method, the sequences of the J.sub.H, J.sub..kappa., and J.sub..lamda. germline segment in the database are set forth in their correct reading frame (e.g., such as is set forth in Table 13).
[0047] The methods provided herein can further include a step after steps a) and/or b) of modifying the nucleic acid sequences encoding a VH chain or a portion thereof and/or modifying the nucleic acid sequences encoding a VL chain or a portion thereof. For example, the nucleic acid sequences can be modified by removing any internal stop codons. Generally, modification of stop codon(s) is made by making only one or two nucleotide changes to the nucleic acid sequences. The codon triplet for a stop codon can be changed to any other codon triplet encoding an amino acid. For example, the stop codon TAA can be modified to be TAT, the stop codon TAG can be modified to be TAT and the stop codon TGA can be modified to be TCA. In another example, the nucleic acid sequences can be modified by removing any internal restriction sites. The nucleotides recognized by a restriction enzyme can be modified to any other nucleotide sequence so long as the sequence is not recognized by a restriction enzyme of interest, i.e. one used in subsequent cloning steps. Generally, only one or two nucleotides changes are made. Typically, modification of restriction sites are made to maximize codon usage in E. coli.
[0048] In the methods of generating a human combinatorial antibody library provided herein, the plurality of nucleic acid sequences encoding a VH chain include, but are not limited to, any set forth in any of SEQ ID NOS: 1059-1410, or a subset thereof. The plurality of nucleic acid sequences encoding a VL chain include, but are not limited to, any set forth in any of SEQ ID NOS: 1411-1422, 1424-1439 and 1441-1471, or a subset thereof.
[0049] In any of the methods provided herein of generating a human combinatorial antibody library, the plurality of nucleic acid sequences encoding a VH chain and/or the plurality of nucleic acid sequences encoding a VL chain can be ranked. For example, the sequences can be ranked based on sequence similarity (e.g. performed by sequence alignment or other methods known in the art and described herein). The sequence similarity between and among the plurality of different nucleic acid molecules encoding a VH chain and/or the sequence similarity between and among the plurality of different nucleic acid molecules encoding a VL chain can be determined. A subset of nucleic acid sequences encoding a VH chain and/or a VL chain can be selected (e.g. for synthesis and subsequent expression) such that the selected sequences include those that are the most similar or are the most different from other selected sequences.
[0050] In the method provided herein for generating a combinatorial antibody library, the synthesized nucleic acid sequences in the addressed libraries are contained in vectors. Hence, a vector from a first vector library containing nucleic acid sequences encoding a VH chain and a vector from a second vector library containing nucleic acid sequences encoding a VL chain are introduced into addressed cells for generation and production of antibody members of the library. The vector can further contain all or a portion of a constant region sufficient to permit association of heavy and light chains. For example, the vector can contain a C.sub.H1 and C.sub.L such that the resulting encoded antibody is a Fab. In the methods provided herein, the cells include prokaryotic or eukaryotic cells. For example, cells include E. coli cells. In the methods of generating a combinatorial antibody library provided herein, the addressed cells can be arranged in a spatial array. The spatial array includes, for example, a multi-well plate such that each individual locus of the plate corresponds to a cell that contains nucleic acid molecules encoding a different combination of a VH and a VL compared to every other cell in the addressed library of cells.
[0051] The antibody or portions thereof that are expressed in the method provided herein include a full length antibody or a fragment or portion thereof sufficient to form an antigen binding site. For example, the expressed antibody is a Fab. The methods provided herein, further can include a step of purifying the antibodies or portions thereof. The antibodies or portion thereof in the libraries include those that are 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% pure. Hence, the addressable library includes antibody members that are purified such that the purified antibodies or portions thereof are addressed, and the antibody within each address is the same antibody and is different from all other antibodies at all other addresses in the library.
[0052] Also provided herein, is an addressable combinatorial antibody library produced by the method provide herein of generating a combinatorial antibody library. Such an antibody library includes members each containing a VH chain having a sequence of amino acids selected from among any of SEQ ID NOS: 1475-836 and a VL chain having a sequence of amino acids selected from among any of SEQ ID NOS: 1827-1838, 1840-1855 and 1857-1888.
[0053] Provided herein is a method of screening a human combinatorial antibody library for binding or activity against a target protein to identify antibodies or portions thereof that bind to a target protein and/or modulate an activity of a target protein. In such methods, a human combinatorial antibody library is provided. The library includes any human combinatorial antibody library provided herein or any human combinatorial antibody library produced by the methods provided herein. In such methods, the an antibody or portion thereof in the library is contacted with a target protein and binding of the antibody or portion thereof with the target protein and/or modulation of a functional activity by an antibody or portion thereof in the library is assessed. Antibodies or portions thereof that bind to the target protein and/or modulate an activity of the target protein are identified, whereby the identified antibody or portion thereof is designated as a "HIT." In such methods, for example, the human combinatorial antibody library is an addressable library and contacting is performed in an addressable array, such that the identity of the "Hit" is known by its address. For example, screening is performed in a spatial array, such as a microwell plate.
[0054] In the methods of screening provided herein, the target protein is a membrane-bound protein, a cell surface receptor (CSR) or a CSR ligand. The membrane-bound protein or CSR includes, but is not limited to, a cytokine receptor, a receptor kinase, a receptor phosphatase, a receptor involved in cell-cell interactions or a cellular adhesion molecule. For example, the target protein includes, but is not limited to, VEGFR-1, VEGFR-2, VEGFR-3, a epidermal growth factor receptor (EGFR), ErbB-2, ErbB-b3, IGF-R1, C-Met, TNF-R1, TNF-R2, BTLA, HVEM, LT-.beta.R, CD20, CD3, CD25, NOTCH, DLL4, G-CSF-R, GM-CSF-R, EPO-R, a cadherin, an integrin, CD52 and CD44, a VEGF-A, VEGF-B, VEGF-C, VEGF-D, PIGF, EGF, HGF, TNF-.alpha., LIGHT, lymphotoxin (LT), IgE, G-CSF, GM-CSF and EPO.
[0055] In some examples, the binding of the antibody or portion thereof to the target protein is assessed. In other examples, modulation of a functional activity of the target protein by an antibody or portion thereof is assessed. The functional activity includes, but is not limited to, cellular proliferation, lymphoma apoptosis, chemotaxis, cancer cell invasion, matrigel, endothelial proliferation, tube formation and signal transduction.
[0056] In the methods of screening provided herein, binding can be assessed on cells and/or a functional activity can be assessed in a cell-based activity. In such examples, the cells express the target protein, typically on their surface as a membrane-bound or extracellular receptor, ligand or adhesion protein. For example, the cells can be transiently or stably expressed with a nucleic acid molecule encoding the target protein.
[0057] Provided herein is a method of screening that includes after identifying a "Hit", for example, in a previous iteration of the method of screening as provided above, a second library combinatorial antibody library is provided. In such examples, the second combinatorial antibody library is based on the identified "Hit." For example, the second library is generated by selecting V.sub.H, D.sub.H and J.sub.H human germline segments that are related by sequence similarity to the germline segments of the identified HIT(s), and combining all possible V.sub.H, D.sub.H and J.sub.H human germline segments or portions thereof in frame to generate a plurality of sequences of nucleic acid molecules each encoding a different VH chain or a portion thereof and selecting V.sub..kappa. and a J.sub..kappa. or V.sub..lamda. and a J.sub..lamda. human germline segments that are related by sequence similarity to the germline segments of the identified HIT(s), and combining all possible V.sub..kappa. and a J.sub..kappa. human germline segments or portions thereof, or all possible V.sub..lamda. and a J.sub..lamda. germline segment or portions thereof in frame to generate a plurality of sequences of nucleic acid molecule each encoding a different VL chain or a portion thereof. The portions of the V.sub.H, D.sub.H, J.sub.H, J.sub..kappa., V.sub..lamda. or J.sub..lamda. are sufficient to produce an antibody containing a VH or VL or portion thereof that is sufficient to bind to an antigen. Upon combination of the germline segments, the nucleic acid molecules are synthesized to produce a first library that contains nucleic acid molecules encoding a VH chain or a portion thereof and a second library that contains nucleic acid molecules encoding a VL chain or a portion thereof. A nucleic acid molecule from the first library and from the second library is introduced into a cell and the steps are repeated a plurality of times to produce a library of cells such that each cell contains nucleic acid molecules encoding a different combination of VH and VL from every other cell in the library of cells. The cells are grown to express the antibodies in each cell, thereby producing a plurality of antibodies or portion thereof such that each produced antibody or portion thereof in the library contains a different combination of a VH and a VL chain or a sufficient portion thereof to form an antigen binding site from all other antibodies or portions thereof in the library. The antibodies are further purified to generate a second human combinatorial antibody library. In such examples, the second human combinatorial antibody library is generated such that the sequence similarity between and among related germline segments is or is about 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more.
[0058] In another example, a second combinatorial antibody library is provided that is generated by selecting a plurality of nucleic acid molecules encoding a VH chain that contain a V.sub.H, a D.sub.H or a J.sub.H human germline segment that is the same as contained in the nucleic acid molecule encoding the identified HIT and selecting a plurality of nucleic acid molecules encoding a VL chain that contains a V.sub..kappa. and a J.sub..kappa. or a V.sub..lamda. and a J.sub..lamda. human germline segment that is the same as contained in the nucleic acid molecule encoding the identified HIT. A nucleic acid molecule from the plurality of nucleic acid molecules encoding a VL and a nucleic acid molecule from the plurality of nucleic acid molecules encoding a VH are introduced into a cells and the cells are grown. This is repeated a plurality of times to produce a library of cells, whereby each cells contains nucleic acid molecules encoding a different combination of a VH and a VL chain from every other cell in the library. Upon expression of the antibodies in each cell, a plurality of antibodies or portions thereof is produced. Each antibody or portion thereof in the library contains a different combination of a VH and a VL chain or a sufficient portion thereof to form an antigen binding site from all other antibodies or portions thereof in the library. The antibodies or portions thereof are purified to generate a second combinatorial antibody library.
[0059] In additional examples, a second combinatorial library is provided based on the identified HIT(s) that is generated by introducing amino acid mutations into the "HIT" to generate a plurality of antibodies or portions thereof. Each antibody member or portion thereof in the second combinatorial antibody library differs from the identified "HIT" by one or more amino acid mutations in its primary sequence. In such examples, the amino acid mutations are in the complementarity determining regions (CDRs) of the identified HIT.
[0060] In each of the methods above, the second human combinatorial library is contacted with a target protein and binding of the antibody or portion thereof with the target protein and/or modulation of a functional activity by an antibody or portion thereof in the library is assessed. Antibodies or portions thereof that bind to the target protein and/or modulate an activity of the target protein are identified, whereby the identified antibody or portion thereof is designated as a further "HIT." In some examples, the second combinatorial antibody library is an addressable library such that the purified antibodies or portions thereof are addressed and each purified antibody within each address is the same antibody and is different from the antibodies at all other addresses. In such examples, contacting is performed in an addressable array, such that the identity of the "Hit" is known by its address. For example, screening is performed in a spatial array, such as a microwell plate.
[0061] In any of the examples of screening provided above, the method is repeated iteratively until a further "HIT" is identified having an optimized binding affinity for a target protein and/or having an activity that is optimized against a target protein compared to previous "HITS" in earlier iterations of the method.
[0062] Provided herein is an anti-DLL4 antibody that contains a VH chain encoded by a sequence of nucleotides compiled from a V.sub.H, D.sub.H and J.sub.H germline segment and a VL chain encoded by a sequence of nucleotides compiled from a V.sub..kappa. and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. germline segments. The V.sub.H germline segment is an IGHV1, and IGHV5 or an IGHV6 or genes and alleles thereof; the D.sub.H germline segment is an IGHD6, and IGHD5 or an IGHD3 or genes and alleles thereof; and the J.sub.H germline segment is an IGHJ1 or an IGHJ4 or genes and alleles thereof. The V.sub..kappa. germline segment is a IGKV3 and the J.kappa. is a IGKJ1 or genes and alleles thereof; the V.sub..lamda. germline segment is a IGLV8 or an IGLV5 and the J.sub..lamda. germline segment is a IGLJ1 or a IGLJ4 or genes and alleles thereof. The anti-DLL4 antibodies provided herein bind DLL4 and/or modulates an activity of DLL4.
[0063] In some examples, the V.sub.H germline segment is an IGHV1-46*01, IGHV1-46*02 or an IGHV1-46*03 or an IGHV6-1*01 or an IGHV6-1*02. The D.sub.H germline segment is an IGHD6-6*01, IGHD5-18*01, IGHD3-3*01 or IGHD3-3*02. The J.sub.H germline segment is an IGHJ1*01, IGHJ4*01, IGHJ4*02 or IGHJ4*03. The V.sub..kappa. germline segment is an IGKV3-11*01 or IGKV3-11*02. The J.sub..kappa. germline segment is a IGKJ1*01. The V.sub..lamda. germline segment is a IGLV8-61*01, IGLV8-61*02, IGLV8-61*03 or IGLV5-48*01. The J.sub..lamda. germline segment that is a IGLJ1*01 or IGLJ4*01. For example, The anti-DLL4 antibody provided herein is an antibody or portion thereof containing a VH chain encoded by a sequence of nucleotides compiled from an IGHV1-46*01, IGHD6-6*01 and IGHJ1*01 and a VL chain encoded by a sequence of nucleotides compiled from IGKV3-11*01 and IGKJ1*01 germline segments; a VH encoded by a sequence of nucleotides compiled from IGHV5-51*03, IGHD5-18*01 and IGHJ4*01 germline segments and a VL chain encoded by a sequence of nucleotides compiled from an IGLV8-61*01 and IGLJ1*01 germline segments; or a VH chain encoded by a sequence of nucleotides compiled from an IGHV6-1*01, and IGHD3-3*01 and an IGHJ4*01 germline segments and a VL chain encoded by a sequence of nucleotides compiled from an IGLV5-48*01 and a IGLJ4*01 germline segments. Anti-DLL4 antibodies provided herein include, but are not limited to, antibodies containing a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1513 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1850; a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1803 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1881; or a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1812 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1884. For example, exemplary of an anti-DLL4 antibody provided herein is an antibody containing a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1513 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1850 Antibodies that include portions of any of the above antibodies that form a sufficient antigen binding site that bind to anti-DLL4 and/or modulate an activity of DLL4 also are provided. Also provided are any antibodies that contain conservative amino acid changes in their sequence compared to any of the antibodies provided herein.
[0064] Provided herein is an anti-EpoR antibody containing a VH chain encoded by a sequence of nucleotides compiled from a V.sub.H, D.sub.H and J.sub.H germline segment and a VL chain encoded by a sequence of nucleotides compiled from a V.sub..kappa. and J.sub..kappa. germline segments. The V.sub.H germline segment is an IGHV1 or genes and alleles thereof. The D.sub.H germline segment is an IGHD6 or an IGHD3 or genes and alleles thereof. The J.sub.H germline segment is an IGHJ1 or genes and alleles thereof. The V.sub..kappa. germline segment is an IGKV4. The J.kappa. is an IGKJ1. The anti-EpoR antibodies provided herein bind EpoR and/or modulate an activity of EpoR.
[0065] In some examples, the V.sub.H germline segment is an IGHV1-46*01, IGHV1-46*02 or an IGHV1-46*03. The D.sub.H germline segment is an IGHD6-6*01, IGHD3-10*01 or IGHD3-10*02. The J.sub.H germline segment is an IGHJ1*01, IGHJ4*01, IGHJ4*02 or IGHJ4*03. The V.kappa. germline segment is an IGKV4-1*01. the J.kappa. germline segment that is a IGKJ1*01. For example, the anti-EpoR antibody provided herein contains a VH chain encoded by a sequence of nucleotides compiled from an IGHV1-46*01, IGHD3-10*01 and IGHJ4*01 and a VL chain encoded by a sequence of nucleotides compiled from IGKV4-1*01 and IGKJ1*01 germline segments or contains a VH chain encoded by a sequence of nucleotides compiled from IGHV1-46*01, IGHD6-6*01 and IGHJ1*01 germline segments and a VL chain encoded by a sequence of nucleotides compiled from an IGKV4-1*01 and IGKJ1*01 germline segments. Anti-EpoR antibodies provided herein include, but are not limited to, an antibody containing a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1509 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1838; or a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1513 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1838. Antibodies that include portions of any of the above antibodies that form a sufficient antigen binding site that bind to anti-EpoR and/or modulate an activity of EpoR also are provided. Also provided are any antibodies that contain conservative amino acid changes in their sequence compared to any of the antibodies provided herein.
[0066] Provided herein is an anti-ErbB2 antibody containing a VH chain encoded by a sequence of nucleotides compiled from a V.sub.H, D.sub.H and J.sub.H germline segment and a VL chain encoded by a sequence of nucleotides compiled from a V.sub..kappa. and J.sub..kappa. germline segments. The V.sub.H germline segment is an IGHV4 or an IGHV1 or genes and alleles thereof. the D.sub.H germline segment is an IGHD6 or an IGHD1 or genes and alleles thereof. The J.sub.H germline segment is an IGHJ1 or an IGHJ2 or genes and alleles thereof. The V.sub..kappa. germline segment is a IGKV3 or IGKV4. The J.kappa. is an IGKJ1 or genes and alleles thereof The anti-ErbB2 antibodies provided herein bind ErbB2 and/or modulate an activity of ErbB2.
[0067] In some examples, the V.sub.H germline segment is an the V.sub.H germline segment is an IGHV1-46*01, IGHV1-46*02 or an IGHV1-46*03 or an IGHV4-31*01, IGHV4-31*02, IGHV4-31*03, IGHV4-31*04, IGHV4-31*05, IGHV4-31*06, IGHV4-31*07, IGHV4-31*08, IGHV4-31*09, IGHV4-3110. The D.sub.H germline segment is an IGHD6-6*01 or IGHD1-26*01. The J.sub.H germline segment is an IGHJ1*01 or an IGHJ2*01. The V.kappa. germline segment is an IGHV3-20*01, IGHV3-20*02 or IGKV4-1*01. The J.kappa. germline segment that is a IGKJ1*01. For example, the anti-ErbB2 antibody provided herein a VH chain encoded by a sequence of nucleotides compiled from an IGHV4-31*02, IGHD1-26*01 and IGHJ2*01 and a VL chain encoded by a sequence of nucleotides compiled from IGKV3-20*01 and IGKJ1*01 germline segments or contains a VH encoded by a sequence of nucleotides compiled from IGHV1-46*01, IGHD6-6*01 and IGHJ1*01 germline segments and a VL chain encoded by a sequence. Anti-ErbB2 antibodies provided herein include, but are not limited to, an antibody containing a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1760 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1833; or a VH chain having a sequence of amino acids set forth in SEQ ID NO: 1513 and a VL chain having a sequence of amino acids set forth in SEQ ID NO:1838. Antibodies that include portions of any of the above antibodies that form a sufficient antigen binding site that bind to anti-ErbB2 and/or modulate an activity of ErbB2 also are provided. Also provided are any antibodies that contain conservative amino acid changes in their sequence compared to any of the antibodies provided herein.
[0068] Any of the antibodies provided herein can further contain a constant region or a portion of a constant region sufficient to permit association of a heavy and light chain. For example, antibodies provided herein include Fab antibodies. In addition, any of the antibodies provided herein include antibodies having a binding affinity that is or is about 10.sup.-6M, 10.sup.-7M, 10.sup.-8M, 10.sup.-9M, 10.sup.-10M, 10.sup.-11M or 10.sup.-12M or lower. For example, any of the antibodies provided herein have a binding affinity that is or is about 1.times.10.sup.-9M, 2.times.10.sup.-9M, 3.times.10.sup.-9M, 4.times.10.sup.-9M, 5.times.10.sup.-9M, 6.times.10.sup.-9M, 7.times.10.sup.-9M, 8.times.10.sup.-9M, 9.times.10.sup.-9M, 1.times.10.sup.-10M, 2.times.10.sup.-10M, 3.times.10.sup.-10M, 4.times.10.sup.-10M, 5.times.10.sup.-10M, 6.times.10.sup.-10M, 7.times.10.sup.-10M, 8.times.10.sup.-10M, 9.times.10.sup.-10M or less.
[0069] Provided herein are methods of treatment using any of the antibodies provided herein, including any of the antibodies identified in the screening method provided herein. Such antibodies can be used to treat diseases or disorders associated with expression and/or activity of the target protein. In one example, provided herein are methods of treatment or uses of treating or formulating a medicament with any of the anti-DLL4 antibodies provided herein for treating a disease or disorder associated with expression and/or activity of DLL4. In another example, provided herein are methods of treatment or uses of treating or formulation of a medicament with any of the anti-EpoR antibodies provided herein for treating a disease or disorder associated with expression and/or activity of EpoR. In an additional example, provided herein are methods of treatment using anti-ErbB2 antibodies provided herein for treating a disease or disorder associated with expression and/or activity of ErbB2.
[0070] Provided herein is a computer system or a computer readable medium that contains computer-readable instructions executable by a computer device for performing a method of combining human germline segments. The method of combining human germline segments includes these steps of (a) accessing a user-created in silico database of all available human antibody germline segments (V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda.); (b) applying an algorithm to generate a collection of every possible recombined full length nucleic acid sequence encoding a heavy chain (5'-V.sub.H-D.sub.H-J.sub.H-3'); (c) applying an algorithm to generate a collection of every possible recombined full length nucleic acid sequence encoding a kappa light chains (5'-V.sub..kappa.-J.sub..kappa.-3') and/or every possible recombined full length nucleic acid sequence encoding a lambda light chains (5'-V.sub..kappa.-J.sub..kappa.-3'); (d) applying an algorithm to modify nucleotides at the V-D and/or D-J joints of the nucleic acid sequences of (b) and at the V-J joints of the nucleic acid sequences of (c) so that the resulting nucleic acids sequences are in frame; (e) modifying the nucleic acid sequences of (d) to remove any inadvertently generated stop codons; (f) assigning each recombined nucleic acid sequence to a unique locus of an addressable format; and (g) generating an output file that identifies the address of each recombined nucleic acid sequences. The method executed by the computer system or computer readable medium can further include (h) after step (e) adding nucleotides at the 5' and 3' termini of the recombined nucleic acid sequences containing a sequence recognized by a restriction enzyme; and (i) modifying the recombined nucleic acid sequence by nucleotide replacement to remove internal nucleotides that are recognized by a restriction enzyme. The method can further modify the recombined nucleic acid sequences to optimize codon usage for bacterial expression. In some examples, the method executed by the computer system or computer readable medium can include before step (f) selecting recombined nucleic acid sequence(s) from the library of recombined nucleic acid sequences based on sequence similarities or differences and assigning only the selected sequences to a locus in an addressable format in step f).
[0071] Provided herein is a method that includes execution of computer-readable instructions for performing a method of combining human germline segments by a computer device, whereby the method includes the steps of (a) accessing a user-created in silico database of all available human antibody germline segments (V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda.); (b) applying an algorithm to generate a collection of every possible recombined full length nucleic acid sequence encoding a heavy chain (5'-V.sub.H-D.sub.H-J.sub.H-3'); (c) applying an algorithm to generate a collection of every possible recombined full length nucleic acid sequence encoding a kappa light chains (5'-V.sub..kappa.-J.sub..kappa.-3') and/or every possible recombined full length nucleic acid sequence encoding a lambda light chains (5'-V.sub..lamda.-J.sub..lamda.-3'); (d) applying an algorithm to modify nucleotides at the V-D and/or D-J joints of the nucleic acid sequences of (b) and at the V-J joints of the nucleic acid sequences of (c) so that the resulting nucleic acids sequences are in frame; (e) modifying the nucleic acid sequences of (d) to remove any inadvertently generated stop codons; (f) assigning each recombined nucleic acid sequence to a unique locus of an addressable format; and (g) generating an output file that identifies the address of each recombined nucleic acid sequences. The method further includes DNA synthesis of the recombined nucleic acid sequences encoding a heavy chain, encoding a kappa light chain, and/or encoding a lambda light chain, or DNA synthesis of a subset of recombined nucleic acid sequences encoding a heavy chain, encoding a kappa light chain, and/or encoding a lambda light chain.
BRIEF DESCRIPTION OF THE FIGURES
[0072] FIG. 1: Schematic illustration of the method for generating antibody diversity
[0073] FIG. 1 illustrates how antibody diversity is generated, including combinatorial diversity, pairing diversity, and junctional diversity.
[0074] FIGS. 2A and 2B: Schematic illustration of iterative methods of antibody discovery
[0075] FIG. 2A is a depiction of the methods of 1) combining heavy and light germ line segments to generate nucleic acid sequences encoding a variable heavy and light chain; 2) synthesizing the nucleic acid molecules; 3) generating a plurality of antibodies by expression and association of the heavy and light chains; purifying the antibodies (e.g. using Piccolo and FLPC purification) and screening the antibodies for an activity (e.g. binding or a functional activity). FIG. 2B is a depiction of iterative methods to identify additional related antibodies involving repeating the method based on identified "HITS" to identify HITs that are optimized or improved for the activity.
[0076] FIG. 3: Plasmid A vector map
[0077] FIG. 3 is an illustrative vector map of plasmid A, provided and described in detail herein.
[0078] FIG. 4: Plasmid D vector map
[0079] FIG. 4 is an illustrative vector map of plasmid D, provided and described in detail herein.
[0080] FIG. 5: Plasmid C vector map
[0081] FIG. 5 is an illustrative vector map of plasmid C, provided and described in detail herein.
[0082] FIG. 6: Plasmid E vector map
[0083] FIG. 6 is an illustrative vector map of plasmid E, provided and described in detail herein.
[0084] FIG. 7: Schematic illustration of DNA Sequence Compilation Software modules
[0085] FIG. 7 illustrates the 10 modules that are contained within the DNA Sequence Compilation Software. As illustrated, there are four layers, including the GUI (Graphical User Interface), the GUI Controls, the Compilation rules and the NCBI tools. In addition, the associations between the modules are illustrated.
[0086] FIG. 8: Algorithm for heavy and light chain combinations
[0087] FIG. 8 is a schematic diagram of the algorithm for generating recombined nucleic acid sequences from germline segments encoding a variable heavy chain or a variable light chain.
[0088] FIG. 9: Algorithm for sequence compilation
[0089] FIG. 9 is a flow chart for modifying recombined nucleic acid sequences encoding a functional variable heavy chain or a variable light chain.
[0090] FIG. 10: Algorithm for ranking sequences
[0091] FIG. 10 is a flow chart for ranking recombined variable heavy and light chains based on diversity score and cluster information.
[0092] FIG. 11: Schematic illustration of sequence database file format
[0093] FIG. 11 illustrates the format of the sequence database file. Illustrated are exemplary sequences, including VH1-18 (SEQ ID NO:10), VH1-2 (SEQ ID NO:13), IGHD1-1*01 (SEQ ID NO:239), IGHJ1*01 (SEQ ID NO:273), AI (SEQ ID NO:330), A10 (SEQ ID NO:354), IGKJ1*01 (SEQ ID NO:356), V1-11 (SEQ ID NO:365), V1-13 (SEQ ID NO:366), IGLJ1*01 (SEQ ID NO:442), IGLJ2*01 (SEQ ID NO:443), and Mfe I restriction site (SEQ ID NO:1900).
[0094] FIG. 12: DNA Sequence Compilation Software initial startup screen
[0095] FIG. 12 depicts the initial startup screen for the DNA Sequence Compilation Software. Illustrated are 5' End sequence CCATGGCA (SEQ ID NO:1901), 5' End sequence CCATGGCG, (SEQ ID NO:1902), 3' End sequence CTAGC (SEQ ID NO:1903) and 3' End sequence GTACT (SEQ ID NO:1904).
[0096] FIG. 13: DNA Sequence Compilation Software splash screen
[0097] FIG. 13 depicts the splash screen that appears upon application startup.
[0098] FIG. 14: DNA Sequence Compilation Software 96-well plate screen
[0099] FIG. 14 depicts a model 96-well plate screen containing compiled sequences. The sequence of gnl|Fabrus|V3-4_IGLJ7*01 (SEQ ID NO:1906) is illustrated in the sequence information box.
[0100] FIG. 15: DNA Sequence Compilation Software light chain manual compilation screen
[0101] FIG. 15 depicts a model manual compilation screen, which allows the user to compile either a kappa or lambda light chain. Selected in the VL sequence box is V1-11 (SEQ ID NO:365). Selected in the JL sequence box is IGLJ1*01 (SEQ ID NO:442). The full length sequence (SEQ ID NO:1914) is depicted in the Dna sequence box.
[0102] FIG. 16: DNA Sequence Compilation Software heavy chain manual compilation screen
[0103] FIG. 16 depicts a model manual compilation screen, which allows the user to compile a heavy chain. Selected in the VH sequence box is VH1-18 (SEQ ID NO:10). Selected in the DH sequence box is IGHD1-1*01 (SEQ ID NO:239). Selected in the JH sequence box is IGHJ1*01 (SEQ ID NO:373). The full length sequence (SEQ ID NO:1915) is depicted in the Dna sequence box.
[0104] FIG. 17: DNA Sequence Compilation Software light chain auto compilation screen
[0105] FIG. 17 depicts a model light chain auto compilation screen. Illustrated are exemplary sequences of variable light chains set forth in SEQ ID NOS:1880-1881, 1883, 1905-1913 and 1916-1940.
[0106] FIG. 18: Light chain BLAST grid
[0107] FIG. 18 depicts a model BLAST grid for a light chain sequence. Illustrated are exemplary sequences of variable light chains set forth in SEQ ID NOS:1880-1881, 1883, 1905-1913, 1916-1939.
[0108] FIG. 19: DNA Sequence Compilation Software heavy chain auto compilation screen
[0109] FIG. 19 depicts a model heavy chain auto compilation screen. Illustrated are exemplary sequences of variable heavy chains set forth in SEQ ID NOS:1556, 1760, 1769, 1821, 1941-1973.
[0110] FIG. 20: Heavy chain BLAST grid
[0111] FIG. 20 depicts a model of the BLAST grid for a heavy chain sequence. Illustrated are exemplary sequences of variable heavy chains set forth in SEQ ID NOS:1530, 1539, 1974-1995.
DETAILED DESCRIPTION
Outline
[0112] A. Definitions
[0113] B. Overview [0114] 1. Methods of Generating Addressable Combinatorial Antibody Collections [0115] 2. The Resulting Libraries [0116] 3. Applications of the libraries
[0117] C. Antibodies [0118] 1. Antibody Polypeptides [0119] 2. Antibody structural and functional domains [0120] 3. Antibody Sequence and Specificity
[0121] D. Methods of Generating Members of the Combinatorial Antibody Library [0122] 1. Methods for Producing Functional Recombined Germline Variable Region [0123] a. Variable Gene Segments [0124] i. Germline Segments [0125] ii. Modified Germline Segments [0126] b. Choosing Germline Segments or Modified Segments Thereof [0127] c. Sequence Compilation [0128] d. Further Sequence Modification of Recombined Nucleic Acid Molecules [0129] i. Codon Usage [0130] ii. Adding or Removing Restriction Enzyme Sites [0131] iii. Linkers [0132] iv. Tags or detectable moieties [0133] v. Mutational Diversity [0134] vi. Directed Peptides [0135] e. Generating Variable Heavy and Light Chain Sequences and Nucleic Acid Molecules [0136] i. Storage and Collection [0137] ii. Determining Sequence Diversity of Collected [0138] Nucleic Acid Sequences [0139] iii. Generating Nucleic Acid Molecules [0140] a) Synthesis [0141] b) Recombinant Techniques [0142] f. Expressing and Producing Antibodies or Portions or Fragments Thereof [0143] Resulting Members in the Library [0144] 2. Automation [0145] a. User-Created Database [0146] b. Sequence Compilation [0147] c. Automation of protein expression and purification
[0148] E. Libraries [0149] 1. VH nucleic Acid Libraries and Vector Libraries Thereof [0150] 2. VL nucleic acid Libraries and Vector Libraries Thereof [0151] 3. Paired Nucleic Acid Libraries or Vector Libraries Thereof [0152] 4. Antibody Libraries [0153] 5. Addressable Formats [0154] a. Multiwell Plate [0155] b. Solid Support [0156] 6. Other Display Methods [0157] a. Cell Display [0158] b. Phage Display [0159] c. mRNA Display and Ribosome Display [0160] d. DNA display
[0161] F. Methods of Production of Antibodies [0162] 1. Vectors [0163] 2. Cells Expression Systems [0164] a. Prokaryotic Expression [0165] b. Yeast [0166] c. Insects [0167] d. Mammalian Cells [0168] e. Plants [0169] 3. Purification [0170] G. Application and Uses of the libraries [0171] 1. Binding Assays [0172] 2. Functional Activity [0173] a. Differentiation [0174] b. Alternation of Gene Expression [0175] c. Cytotoxicity Activity [0176] 3. Targets [0177] Membrane-bound proteins, receptors and ligands thereof [0178] i. Notch and Notch ligands a) Notch Proteins b) DLL4 [0179] ii. ErbB family a) Epidermal Growth Factor Receptor (EGFR) b) Human Epidermal Growth Factor Receptor 2 (HER2/neu) [0180] iii. IGF-R1 (Insulin-like Growth Factor 1 Receptor) [0181] iv. c-Met [0182] v. CD20--B lymphocyte antigen [0183] vii. Erythropoietin Receptor (Epo-R) [0184] viii. Cadherins a) P-Cadherin (P-cad/CDH3) [0185] ix. CD44 [0186] 4. Iterative Screening [0187] 5. Directed Evolution [0188] a. Random Mutagenesis [0189] i. Saturation Mutagenesis [0190] ii. Error Prone PCR [0191] iii. Cell lines [0192] iv. DNA shuffling/antibody chain shuffling [0193] v. CDR walking [0194] vi. Framework Stabilization [0195] 6. Epitope Mapping [0196] 7. In Vivo Assays of Identified Hits [0197] 8. Articles of Manufacture/Kits [0198] 9. Formulations/Administrations and uses of antibodies and polypeptides
[0199] H. Examples
A. DEFINITIONS
[0200] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as is commonly understood by one of skill in the art to which the invention(s) belong. All patents, patent applications, published applications and publications, Genbank sequences, databases, websites and other published materials referred to throughout the entire disclosure herein, unless noted otherwise, are incorporated by reference in their entirety. In the event that there are a plurality of definitions for terms herein, those in this section prevail. Where reference is made to a URL or other such identifier or address, it understood that such identifiers can change and particular information on the internet can come and go, but equivalent information can be found by searching the internet. Reference thereto evidences the availability and public dissemination of such information.
[0201] As used herein, "combinatorial library" refers to collections of compounds formed by reacting different combinations of interchangeable chemical "building blocks" to produce a collection of compounds based on permutations of the building blocks. For an antibody combinatorial library, the building blocks are the component V, D and J regions (or modified forms thereof) from which antibodies are formed. For purposes herein, the terms "library" or "collection" are used interchangeably.
[0202] As used herein, a combinatorial antibody library is a collection of antibodies (or portions thereof, such as Fabs), where the antibodies are encoded by nucleic acid molecules produced by the combination of V, D and J gene segments, particularly human V, D and J germline segments. The combinatorial libraries herein typically contain at least 50 different antibody (or antibody portions or fragment) members, typically at or about 50, 100, 500, 10.sup.3, 2.times.10.sup.3, 3.times.10.sup.3, 4.times.10.sup.3, 5.times.10.sup.3, 6.times.10.sup.3, 7.times.10.sup.3, 8.times.10.sup.3, 9.times.10.sup.3, 1.times.10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 1.times.10.sup.5, 2.times.10.sup.5, 3.times.10.sup.5, 4.times.10.sup.5, 5.times.10.sup.5, 6.times.10.sup.5, 7.times.10.sup.5, 8.times.10.sup.5, 9.times.10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9, 10.sup.10, or more different members. The resulting libraries or collections of antibodies or portions thereof, can be screened for binding to a target protein or modulation of a functional activity.
[0203] As used herein, a human combinatorial antibody library is a collection of antibodies or portions thereof, whereby each member contains a VL and VH chains or a sufficient portion thereof to form an antigen binding site encoded by nucleic acid containing human germline segments produced as described herein.
[0204] As used herein, germline gene segments refer to immunoglobulin (Ig) variable (V), diversity (D) and junction (J) or constant (C) genes from the germline that encode immunoglobulin heavy or light (kappa and lambda) chains. There are multiple V, D, J and C gene segments in the germline, but gene rearrangement results in only one segment of each occurring in each functional rearranged gene. For example, a functionally rearranged heavy chain contains one V, one D and one J and a functionally rearranged light chain gene contains one V and one J. Hence, these gene segments are carried in the germ cells but cannot be transcribed and translated into heavy and light chains until they are arranged into functional genes. During B-cell differentiation in the bone marrow, these gene segments are randomly shuffled by a dynamic genetic system capable of generating more than 10.sup.10 specificities. For purposes herein, the gene segments are rearranged in vitro by combination or compilation of the individual germline segments.
[0205] Reference to a variable germline segment herein refers to V, D and J groups, subgroups, genes or alleles thereof. Gene segment sequences are accessible from known database (e.g., National Center for Biotechnology Information (NCBI), the international ImMunoGeneTics information System.RTM. (IMGT), the Kabat database and the Tomlinson's VBase database (Lefranc (2003) Nucleic Acids Res., 31:307-310; Martin et al., Bioinformatics Tools for Antibody Engineering in Handbook of Therapeutic Antibodies, Wiley-VCH (2007), pp. 104-107). Tables 3-5 list exemplary human variable germline segments. Sequences of exemplary V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and or J.sub..lamda. germline segments are set forth in SEQ ID NOS: 10-451 and 868. For purposes herein, a germline segment includes modified sequences thereof, that are modified in accord with the rules of sequence compilation provided herein to permit practice of the method. For example, germline gene segments include those that contain one amino acid deletion or insertion at the 5' or 3' end compared to any of the sequences of nucleotides set forth in SEQ ID NOS:10-451, 868.
[0206] As used herein, modified form with reference to a germline segment refers to a sequence of nucleotides that is substantially the same as the sequence of nucleotides of a human germline segment (e.g. a V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda.) except that the sequence of nucleotides contains one or a few nucleotide differences, for example, 2, 3, 4, 5, 6, 7, 8, 9 or 10 nucleotide differences compared to the corresponding sequence in a germline segment sequence. In some instances, modified germline sequences are modified in accord with the rules herein to remove stop codons, restriction enzyme sites, or to add or delete nucleotides to maintain reading frames.
[0207] As used herein, inverted sequence with reference to nucleotides of a germline segment means that the gene segment has a sequence of nucleotides that is the reverse complement of a reference sequence of nucleotides. For purposes herein, the reference sequence of nucleotides is a germline segment, typically a D.sub.H germline segment.
[0208] As used herein, "compilation," "compile," "combine," "combination," "rearrange," "rearrangement," or other similar terms or grammatical variations thereof refers to the process by which germline segments are ordered or assembled into nucleic acid sequences representing genes. For example, variable heavy chain germline segments are assembled such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment, thereby resulting in a nucleic acid sequence encoding a VH chain. Variable light chain germline segments are assembled such that the V.sub.L segment is 5' to the J.sub.L segment, thereby resulting in a nucleic acid sequence encoding a VL chain. A constant gene segment or segments also can be assembled onto the 3' end of a nucleic acid encoding a VH or VL chain.
[0209] As used herein, "linked," or "linkage" or other grammatical variations thereof with reference to germline segments refers to the joining of germline segments. Linkage can be direct or indirect. Germline segments can be linked directly without additional nucleotides between segments, or additional nucleotides can be added to render the entire segment in-frame, or nucleotides can be deleted to render the resulting segment in-frame. It is understood that the choice of linker nucleotides is made such that the resulting nucleic acid molecule is in-frame and encodes a functional and productive antibody.
[0210] As used herein, "in-frame" or "linked in-frame" with reference to linkage of human germline segments means that there are insertions and/or deletions in the nucleotide germline segments at the joined junctions to render the resulting nucleic acid molecule in-frame with the 5' start codon (ATG), thereby producing a "productive" or functional full-length polypeptide. The choice of nucleotides inserted or deleted from germline segments, particularly at joints joining various VD, DJ and VJ segments, is in accord with the rules provided in the method herein for V(D)J joint generation. For example, germline segments are assembled such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment. At the junction joining the V.sub.H and the D.sub.H and at the junction joining the D.sub.H and J.sub.H segments, nucleotides can be inserted or deleted from the individual V.sub.H, D.sub.H or J.sub.H segments, such that the resulting nucleic acid molecule containing the joined VDJ segments are in-frame with the 5' start codon (ATG).
[0211] As used herein, a "functional antibody" or "productive antibody" with reference to a nucleic acid encoding an antibody or portion thereof refers to an antibody or portion thereof, such as Fab, that is encoded by the nucleic acid molecule produced by the methods as described herein. In a functional or productive antibody, the V(D)J germline segments are compiled (i.e. rearranged) such that the encoded antibody or portion thereof is not truncated and/or the amino acid sequence is not out of frame. This means that the nucleic acid molecule does not contain internal stop codons that result in the protein translation machinery terminating protein assembly prematurely.
[0212] As used herein, a portion of an antibody includes sufficient amino acids to form an antigen binding site.
[0213] As used herein, a reading frame refers to a contiguous and non-overlapping set of three-nucleotide codons in DNA or RNA. Because three codons encode one amino acid, there exist three possible reading frames for given nucleotide sequence, reading frames 1, 2 or 3. For example, the sequence ACTGGTCA will be ACT GGT CA for reading frame 1, A CTG GTC A for reading frame 2 and AC TGG TCA for reading frame 3. Generally for practice of the method described herein, nucleic acid sequences are combined so that the V sequence has reading frame 1.
[0214] As used herein, a stop codon is used to refer to a three-nucleotide sequence that signals a halt in protein synthesis during translation, or any sequence encoding that sequence (e.g. a DNA sequence encoding an RNA stop codon sequence), including the amber stop codon (UAG or TAG)), the ochre stop codon (UAA or TAA)) and the opal stop codon (UGA or TGA)). It is not necessary that the stop codon signal termination of translation in every cell or in every organism. For example, in suppressor strain host cells, such as amber suppressor strains and partial amber suppressor strains, translation proceeds through one or more stop codon (e.g. the amber stop codon for an amber suppressor strain), at least some of the time.
[0215] As used herein, reference to a variable heavy (VH) chain or a variable light (VL) chain (also termed VH domain or VL domain) refers to the polypeptide chains that make up the variable domain of an antibody. For purposes herein, heavy chain germline segments are designated as V.sub.H, D.sub.H and J.sub.H, and compilation thereof results in a nucleic acid encoding a VH chain. Light chain germline segments are designated as V.sub.L or J.sub.L, and include kappa and lambda light chains (V.sub..kappa. and J.sub..kappa.; V.sub..lamda. and J.sub..lamda.) and compilation thereof results in a nucleic acid encoding a VL chain. It is understood that a light chain is either a kappa or lambda light chain, but does not include a kappa/lambda combination by virtue of compilation of a V.sub..kappa. and J.sub..lamda..
[0216] As used herein, a "degenerate codon" refers to three-nucleotide codons that specifies the same amino acid as a codon in a parent nucleotide sequence. One of skill in the art is familiar with degeneracy of the genetic code and can identify degenerate codons.
[0217] As used herein, a "group" with reference to a germline segment refers to a core coding region from an immunoglobulin, i.e. a variable (V) gene, diversity (D) gene, joining (J) gene or constant (C) gene encoding a heavy or light chain. Exemplary of germline segment groups include V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda..
[0218] As used herein, a "subgroup" with reference to a germline segment refers to a set of sequences that are defined by nucleotide sequence similarity or identity. Generally, a subgroup is a set of genes that belong to the same group [V, D, J or C], in a given species, and that share at least 75% identity at the nucleotide level. Subgroups are classified based on IMGT nomenclature (imgt.cines.fr; see e.g., Lefranc et al. (2008) Briefings in Bioinformatics, 9:263-275). Generally, a subgroup represent a multigene family.
[0219] As used herein, an allele of a gene refer to germline sequences that have sequence polymorphism due to one or more nucleotide differences in the coding region compared to a reference gene sequence (e.g. substitutions, insertions or deletions). Thus, IG sequences that belong to the same subgroup can be highly similar in their coding sequence, but nonetheless exhibit high polymorphism. Subgroup alleles are classified based on IMGT nomenclature with an asterisk (*) followed by a two figure number. Exemplary allelic subgroup germline segments for V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda. are set forth in Tables 3-5.
[0220] As used herein, a "family" with reference to a germline segment refers to sets of germline segment sequences that are defined by amino acid sequence similarity or identity. Generally, a germline family includes all alleles of a gene.
[0221] As used herein, a "segment designated D.sub.H" refers to any sequence of nucleotides of at least or about 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 30, 40, 50, 60, 70, 80, 90 or 100 nucleotides in length. The sequence of nucleotides is sufficient to code for part of the CDR3 region of the VH chain.
[0222] As used herein, reference to a V, D or J gene segment "derived from a germline segment" refers to the corresponding nucleotides in a VH or VL nucleic acid sequence, that by recombination events, derived from a V, D or J germline gene.
[0223] As used herein, reference to a V region, D region or J region in an antibody or portion or fragment thereof refers to amino acids encoded by nucleotides that, by recombination events, derive from a corresponding V, D or J germline segment gene.
[0224] As used herein, "diversity" with respect to members in a collection refers to the number of unique members in a collection. Hence, diversity refers to the number of different amino acid sequences or nucleic acid sequences, respectively, among the analogous polypeptide members of that collection. For example, a collection of polynucleotides having a diversity of 10.sup.4 contains 10.sup.4 different nucleic acid sequences among the analogous polynucleotide members. In one example, the provided collections of polynucleotides and/or polypeptides have diversities of at least at or about 10.sup.2, 10.sup.3, 10.sup.4, 10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9, 10.sup.10 or more.
[0225] As used herein, "a diversity ratio" refers to a ratio of the number of different members in the library over the number of total members of the library. Thus, a library with a larger diversity ratio than another library contains more different members per total members, and thus more diversity per total members. The provided libraries include libraries having high diversity ratios, such as diversity ratios approaching 1, such as, for example, at or about 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 0.91, 0.92, 0.93, 0.94, 0.95, 0.96, 0.97, 0.98, or 0.99.
[0226] As used herein, "sequence diversity" refers to a representation of nucleic acid sequence similarity and is determined using sequence alignments, diversity scores, and/or sequence clustering. Any two sequences can be aligned by laying the sequences side-by-side and analyzing differences within nucleotides at every position along the length of the sequences. Sequence alignment can be assessed in silico using Basic Local Alignment Search Tool (BLAST), an NCBI tool for comparing nucleic acid and/or protein sequences. The use of BLAST for sequence alignment is well known to one of skill in the art. The Blast search algorithm compares two sequences and calculates the statistical significance of each match (a Blast score). Sequences that are most similar to each other will have a high Blast score, whereas sequences that are most varied will have a low Blast score.
[0227] As used herein, antibody refers to immunoglobulins and immunoglobulin portions, whether natural or partially or wholly synthetic, such as recombinantly, produced, including any portion thereof containing at least a portion of the variable region of the immunoglobulin molecule that is sufficient to form an antigen binding site. Hence, an antibody or portion thereof includes any protein having a binding domain that is homologous or substantially homologous to an immunoglobulin antigen binding site. For example, an antibody refers to an antibody that contains two heavy chains (which can be denoted H and H') and two light chains (which can be denoted L and L'), where each heavy chain can be a full-length immunoglobulin heavy chain or a portion thereof sufficient to form an antigen binding site (e.g. heavy chains include, but are not limited to, VH, chains VH-CH1 chains and VH-CH1-CH2-CH3 chains), and each light chain can be a full-length light chain or a portion thereof sufficient to form an antigen binding site (e.g. light chains include, but are not limited to, VL chains and VL-CL chains). Each heavy chain (H and H') pairs with one light chain (L and L', respectively). Typically, antibodies minimally include all or at least a portion of the variable heavy (VH) chain and/or the variable light (VL) chain. The antibody also can include all or a portion of the constant region.
[0228] For purposes herein, the term antibody includes full-length antibodies and portions thereof including antibody fragments, such as, but not limited to, Fab, Fab', F(ab').sub.2, single-chain Fvs (scFv), Fv, dsFv, diabody, Fd and Fd' fragments Fab fragments, Fd fragments and scFv fragments. Other known fragments include, but are not limited to, scFab fragments (Hust et al., BMC Biotechnology (2007), 7:14). Antibodies include members of any immunoglobulin class, including IgG, IgM, IgA, IgD and IgE.
[0229] As used herein, a full-length antibody is an antibody having two full-length heavy chains (e.g. VH-CH1-CH2-CH3 or VH-CH1-CH2-CH3-CH4) and two full-length light chains (VL-CL) and hinge regions, such as human antibodies produced by antibody secreting B cells and antibodies with the same domains that are produced synthetically.
[0230] As used herein, antibody fragment or antibody portion refers to any portion of a full-length antibody that is less than full length but contains at least a portion of the variable region of the antibody sufficient to form an antigen binding site (e.g. one or more CDRs) and thus retains the a binding specificity and/or an activity of the full-length antibody; antibody fragments include antibody derivatives produced by enzymatic treatment of full-length antibodies, as well as synthetically, e.g. recombinantly produced derivatives. Examples of antibody fragments include, but are not limited to, Fab, Fab', F(ab').sub.2, single-chain Fvs (scFv), Fv, dsFv, diabody, Fd and Fd' fragments (see, for example, Methods in Molecular Biology, Vol 207: Recombinant Antibodies for Cancer Therapy Methods and Protocols (2003); Chapter 1; p 3-25, Kipriyanov). The fragment can include multiple chains linked together, such as by disulfide bridges and/or by peptide linkers. An antibody fragment generally contains at least about 50 amino acids and typically at least 200 amino acids.
[0231] As used herein, an Fv antibody fragment is composed of one variable heavy domain (VH) and one variable light (VL) domain linked by noncovalent interactions.
[0232] As used herein, a dsFv refers to an Fv with an engineered intermolecular disulfide bond, which stabilizes the VH-VL pair.
[0233] As used herein, an Fd fragment is a fragment of an antibody containing a variable domain (VH) and one constant region domain (CH1) of an antibody heavy chain.
[0234] As used herein, "Fab fragment" is an antibody fragment that contains the portion of the full-length antibody that results from digestion of a full-length immunoglobulin with papain, or a fragment having the same structure that is produced synthetically, e.g. recombinantly. A Fab fragment contains a light chain (containing a VL and CL portion) and another chain containing a variable domain of a heavy chain (VH) and one constant region domain portion of the heavy chain (CH1); it can be recombinantly produced.
[0235] As used herein, a F(ab').sub.2 fragment is an antibody fragment that results from digestion of an immunoglobulin with pepsin at pH 4.0-4.5, or a synthetically, e.g. recombinantly, produced antibody having the same structure. The F(ab').sub.2 fragment contains two Fab fragments but where each heavy chain portion contains an additional few amino acids, including cysteine residues that form disulfide linkages joining the two fragments; it can be recombinantly produced.
[0236] A Fab' fragment is a fragment containing one half (one heavy chain and one light chain) of the F(ab').sub.2 fragment.
[0237] As used herein, an Fd' fragment is a fragment of an antibody containing one heavy chain portion of a
[0238] F(ab')2 fragment.
[0239] As used herein, an Fv' fragment is a fragment containing only the V.sub.H and V.sub.L domains of an antibody molecule.
[0240] As used herein, a scFv fragment refers to an antibody fragment that contains a variable light chain (VL) and variable heavy chain (VH), covalently connected by a polypeptide linker in any order. The linker is of a length such that the two variable domains are bridged without substantial interference. Exemplary linkers are (Gly-Ser).sub.n residues with some Glu or Lys residues dispersed throughout to increase solubility.
[0241] As used herein, diabodies are dimeric scFv; diabodies typically have shorter peptide linkers than scFvs, and they preferentially dimerize.
[0242] As used herein, hsFv refers to antibody fragments in which the constant domains normally present in a Fab fragment have been substituted with a heterodimeric coiled-coil domain (see, e.g., Arndt et al. (2001) J Mol Biol. 7:312:221-228).
[0243] As used herein, a polypeptide domain is a part of a polypeptide (a sequence of three or more, generally 5 or 7 or more amino acids) that is a structurally and/or functionally distinguishable or definable. Exemplary of a polypeptide domain is a part of the polypeptide that can form an independently folded structure within a polypeptide made up of one or more structural motifs (e.g. combinations of alpha helices and/or beta strands connected by loop regions) and/or that is recognized by a particular functional activity, such as enzymatic activity or antigen binding. A polypeptide can have one, typically more than one, distinct domains. For example, the polypeptide can have one or more structural domains and one or more functional domains. A single polypeptide domain can be distinguished based on structure and function. A domain can encompass a contiguous linear sequence of amino acids. Alternatively, a domain can encompass a plurality of non-contiguous amino acid portions, which are non-contiguous along the linear sequence of amino acids of the polypeptide. Typically, a polypeptide contains a plurality of domains. For example, each heavy chain and each light chain of an antibody molecule contains a plurality of immunoglobulin (Ig) domains, each about 110 amino acids in length.
[0244] As used herein, an Ig domain is a domain, recognized as such by those in the art, that is distinguished by a structure, called the Immunoglobulin (Ig) fold, which contains two beta-pleated sheets, each containing anti-parallel beta strands of amino acids connected by loops. The two beta sheets in the Ig fold are sandwiched together by hydrophobic interactions and a conserved intra-chain disulfide bond. Individual immunoglobulin domains within an antibody chain further can be distinguished based on function. For example, a light chain contains one variable region domain (VL) and one constant region domain (CL), while a heavy chain contains one variable region domain (VH) and three or four constant region domains (CH). Each VL, CL, VH, and CH domain is an example of an immunoglobulin domain.
[0245] As used herein, a "variable domain" with reference to an antibody is a specific Ig domain of an antibody heavy or light chain that contains a sequence of amino acids that varies among different antibodies. Each light chain and each heavy chain has one variable region domain (VL, and, VH). The variable domains provide antigen specificity, and thus are responsible for antigen recognition. Each variable region contains CDRs that are part of the antigen binding site domain and framework regions (FRs).
[0246] As used herein, "hypervariable region," "HV," "complementarity-determining region" and "CDR" and "antibody CDR" are used interchangeably to refer to one of a plurality of portions within each variable region that together form an antigen binding site of an antibody. Each variable region domain contains three CDRs, named CDR1, CDR2, and CDR3. The three CDRs are non-contiguous along the linear amino acid sequence, but are proximate in the folded polypeptide. The CDRs are located within the loops that join the parallel strands of the beta sheets of the variable domain.
[0247] As used herein, framework regions (FRs) are the domains within the antibody variable region domains that are located within the beta sheets; the FR regions are comparatively more conserved, in terms of their amino acid sequences, than the hypervariable regions.
[0248] As used herein, a constant region domain is a domain in an antibody heavy or light chain that contains a sequence of amino acids that is comparatively more conserved among antibodies than the variable region domain. Each light chain has a single light chain constant region (CL) domain and each heavy chain contains one or more heavy chain constant region (CH) domains, which include, CH1, CH2, CH3 and CH4. Full-length IgA, IgD and IgG isotypes contain CH1, CH2 CH3 and a hinge region, while IgE and IgM contain CH1, CH2 CH3 and CH4. CH1 and CL domains extend the Fab arm of the antibody molecule, thus contributing to the interaction with antigen and rotation of the antibody arms. Antibody constant regions can serve effector functions, such as, but not limited to, clearance of antigens, pathogens and toxins to which the antibody specifically binds, e.g. through interactions with various cells, biomolecules and tissues.
[0249] As used herein, humanized antibodies refer to antibodies that are modified to include "human" sequences of amino acids so that administration to a human does not provoke an immune response. Methods for preparation of such antibodies are known. For example, the antibody in which the amino acid composition of the non-variable regions can be based on human antibodies. Computer programs have been designed to identify such regions.
[0250] As used herein, "antigen-binding site" refers to the interface formed by one or more complementary determining regions (CDRs; also called hypervariable region). Each antigen binding site contains three CDRs from the heavy chain variable region and three CDRs from the light chain variable region. An antibody molecule has two antigen combining sites, each containing portions of a heavy chain variable region and portions of a light chain variable region. The antigen combining sites can contain other portions of the variable region domains in addition to the CDRs.
[0251] As used herein, reference to an "antibody or portion thereof that is sufficient to form an antigen binding site" means that the antibody or portion thereof contains at least 1 or 2, typically 3, 4, 5 or all 6 CDRs of the VH and VL sufficient to retain at least a portion of the binding specificity of the corresponding full-length antibody containing all 6 CDRs. Generally, a sufficient antigen binding site at least requires CDR3 of the heavy chain (CDRH3). It typically further requires the CDR3 of the light chain (CDRL3). As described herein, one of skill in the art knows and can identify the CDRs based on kabat or Chothia numbering (see e.g., Kabat, E. A. et al. (1991) Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242, and Chothia, C. et al. (1987) J. Mol. Biol. 196:901-917). For example, based on Kabat numbering, CDR-LI corresponds to residues L24-L34; CDR-L2 corresponds to residues L50-L56; CDR-L3 corresponds to residues L89-L97; CDR-H1 corresponds to residues H31-H35, 35a or 35b depending on the length; CDR-H2 corresponds to residues H50-H65; and CDR-H3 corresponds to residues H95-H102.
[0252] As used herein, a "peptide mimetic" is a peptide that mimics the activity of a polypeptide. For example, an erythropoietin (EPO) peptide mimetic is a peptide that mimics the activity of Epo, such as for binding and activation of the EPO receptor.
[0253] As used herein, an optimized antibody refers to an antibody, or portion thereof, that has an improved binding affinity for a target protein and/or an improved functional activity compared to a reference antibody. Typically, the antibody is optimized by virtue of one or more amino acid modifications (amino acid deletion, replacement or insertion) compared to a parent antibody not containing the one or more amino acid modifications. Generally, an activity or binding affinity is increased by at or about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 30-fold, 40-fold, 50-fold, 60-fold, 70-fold, 80-fold, 90-fold, 100-fold, 200-fold, 300-fold, 400-fold, 500-fold, 600-fold, 700-fold, 800-fold, 900-fold, 1000-fold or more compared to an activity or binding affinity of the parent antibody (e.g. germline Hit not containing the modification(s)).
[0254] As used herein, corresponding with reference to corresponding residues, for example "amino acid residues corresponding to", refers to residues compared among or between two polypeptides that are related sequences (e.g. allelic variants, genes of the same family, species variants). One of skill in the art can readily identify residues that correspond between or among polypeptides. For example, by aligning the sequence of regions encoded by germline segments, one of skill in the art can identify corresponding residues, using conserved and identical amino acids as guides. One of skill in the art can manually align a sequence or can use any of the numerous alignment programs available (for example, BLAST). Hence, an amino acid residues or positions that correspond to each other are those residues that are determined to correspond to one another based on sequence and/or structural alignments with a specified reference polypeptide.
[0255] As used herein, a consensus sequence is a sequence containing residues that are the most frequently occurring residues at each position when a plurality of related sequences (e.g. allelic variants, genes of the same family, species variants) are aligned. Hence a consensus sequence represents the residues that are the most abundant in the alignment at each position. For purposes herein, for example, germline sequences, or portions thereof, can be aligned to generate a consensus germline sequence.
[0256] As used herein, a locus in a library refers to a location or position, that can contain a member or members of library. The position does not have to be a physical position. For example, if the collection is provided as an array on a solid support, the support contains loci that can or do present members of the array.
[0257] As used herein, an address refers to a unique identifier for each locus in a collection whereby an addressed member (e.g. an antibody) can be identified. An addressed moiety is one that can be identified by virtue of its locus or location. Addressing can be effected by position on a surface, such as a well of a microplate. For example, an address for a protein in a microwell plate that is F9 means that the protein is located in row F, column 9 of the microwell plate. Addressing also can be effected by other identifiers, such as a tag encoded with a bar code or other symbology, a chemical tag, an electronic, such RF tag, a color-coded tag or other such identifier.
[0258] As used herein, an array refers to a collection of elements, such as antibodies, containing three or more members.
[0259] As used herein, a "spatial array" is an array where members are separated or occupy a distinct space in an array. Hence, spatial arrays are a type of addressable array. Examples of spatial arrays include microtiter plates where each well of a plate is an address in the array. Spatial arrays include any arrangement wherein a plurality of different molecules, e.g, polypeptides, are held, presented, positioned, situated, or supported. Arrays can include microtiter plates, such as 48-well, 96-well, 144-well, 192-well, 240-well, 288-well, 336-well, 384-well, 432-well, 480-well, 576-well, 672-well, 768-well, 864-well, 960-well, 1056-well, 1152-well, 1248-well, 1344-well, 1440-well, or 1536-well plates, tubes, slides, chips, flasks, or any other suitable laboratory apparatus. Furthermore, arrays can also include a plurality of sub-arrays. A plurality of sub-arrays encompasses an array where more than one arrangement is used to position the polypeptides. For example, multiple 96-well plates can constitute a plurality of sub-arrays and a single array.
[0260] As used herein, an addressable library is a collection of molecules such as nucleic acid molecules or protein agents, such as antibodies, in which each member of the collection is identifiable by virtue of its address.
[0261] As used herein, an addressable array is one in which the members of the array are identifiable by their address, the position in a spatial array, such as a well of a microtiter plate, or on a solid phase support, or by virtue of an identifiable or detectable label, such as by color, fluorescence, electronic signal (i.e. RF, microwave or other frequency that does not substantially alter the interaction of the molecules of interest), bar code or other symbology, chemical or other such label. Hence, in general the members of the array are located at identifiable loci on the surface of a solid phase or directly or indirectly linked to or otherwise associated with the identifiable label, such as affixed to a microsphere or other particulate support (herein referred to as beads) and suspended in solution or spread out on a surface.
[0262] As used herein, "an addressable combinatorial antibody library" refers to a collection of antibodies in which member antibodies are identifiable and all antibodies with the same identifier, such as position in a spatial array or on a solid support, or a chemical or RF tag, bind to the same antigen, and generally are substantially the same in amino acid sequence. For purposes herein, reference to an "addressable arrayed combinatorial antibody library" means that the antibody members are addressed in an array.
[0263] As used herein, a support (also referred to as a matrix support, a matrix, an insoluble support or solid support) refers to any solid or semisolid or insoluble support to which a molecule of interest, typically a biological molecule, organic molecule or biospecific ligand is linked or contacted. Such materials include any materials that are used as affinity matrices or supports for chemical and biological molecule syntheses and analyses, such as, but are not limited to: polystyrene, polycarbonate, polypropylene, nylon, glass, dextran, chitin, sand, pumice, agarose, polysaccharides, dendrimers, buckyballs, polyacrylamide, silicon, rubber, and other materials used as supports for solid phase syntheses, affinity separations and purifications, hybridization reactions, immunoassays and other such applications. The matrix herein can be particulate or can be in the form of a continuous surface, such as a microtiter dish or well, a glass slide, a silicon chip, a nitrocellulose sheet, nylon mesh, or other such materials. When particulate, typically the particles have at least one dimension in the 5-10 mm range or smaller. Such particles, referred collectively herein as "beads", are often, but not necessarily, spherical. Such reference, however, does not constrain the geometry of the matrix, which can be any shape, including random shapes, needles, fibers, and elongated. Roughly spherical "beads", particularly microspheres that can be used in the liquid phase, also are contemplated. The "beads" can include additional components, such as magnetic or paramagnetic particles (see, e.g., Dynabeads.RTM. (Dynal, Oslo, Norway)) for separation using magnets, as long as the additional components do not interfere with the methods and analyses herein.
[0264] As used herein, matrix or support particles refers to matrix materials that are in the form of discrete particles. The particles have any shape and dimensions, but typically have at least one dimension that is 100 mm or less, 50 mm or less, 10 mm or less, 1 mm or less, 100 .mu.m or less, 50 .mu.m or less and typically have a size that is 100 mm.sup.3 or less, 50 mm.sup.3 or less, 10 mm.sup.3 or less, and 1 mm.sup.3 or less, 100 .mu.m.sup.3 or less and can be on the order of cubic microns. Such particles are collectively called "beads."
[0265] As used herein, in silico refers to research and experiments performed using a computer. In silico methods include, but are not limited to, molecular modeling studies, biomolecular docking experiments, and virtual representations of molecular structures and/or processes, such as molecular interactions. For purposes herein, the antibody members of a library can be designed using a computer program that selects component V, D and J germline segments from among those input into the computer and joins them in-frame to output a list of nucleic acid molecules for synthesis. Thus, the recombination of the components of the antibodies in the collections or libraries provided herein, can be performed in silico by combining the nucleotide sequences of each building block in accord with software that contains rules for doing so. The process can be performed manually without a computer, but the computer provides the convenience of speed.
[0266] As used herein, a database refers to a collection of data items. For purposes herein, reference to a database is typically with reference to antibody databases, which provide a collection of sequence and structure information for antibody genes and sequences. Exemplary antibody databases include, but are not limited to, IMGT.RTM., the international ImMunoGeneTics information system (imgt.cines.fr; see e.g., Lefranc et al. (2008) Briefings in Bioinformatics, 9:263-275), National Center for Biotechnology Information (NCBI), the Kabat database and the Tomlinson's VBase database (Lefranc (2003) Nucleic Acids Res., 31:307-310; Martin et al., Bioinformatics Tools for Antibody Engineering in Handbook of Therapeutic Antibodies, Wiley-VCH (2007), pp. 104-107). A database also can be created by a user to include any desired sequences. The database can be created such that the sequences are inputted in a desired format (e.g., in a particular reading frame; lacking stop codons; lacking signal sequences). The database also can be created to include sequences in addition to antibody sequences.
[0267] As used herein, "a computer-based system" refers to the hardware, software, and data storage media and methods used to recombine germline segments. The minimum hardware of the computer-based systems provided herein include a central processing unit (CPU), input mean, output means and data storage means. A skilled artisan can select a suitable computer-based systems for use in the methods and systems provided herein.
[0268] As used herein, "recorded" refers to a process for storing information on computer readable medium. A skilled artisan can readily adopt any of the presently known methods for recording information on computer readable medium to array image data. The choice of the data storage structure can generally be based on the media and platforms chosen to access the stored information. In addition, a variety of data processor programs and formats can be used to store the array image information on computer readable medium. The image information can be represented in a word processing text file, formatted in commercially-available software such as MICROSOFT Word.RTM., graphics files or represented in the form of an ASCII file, stored in a database application, such as DB2.RTM., Sybase.RTM. and Oracle.RTM.. A skilled artisan can adapt any number of data processor structuring formats (e.g., text file or database) in order to obtain computer readable medium having recorded thereon the information or instructions as described herein.
[0269] As used herein, "screening" refers to identification or selection of an antibody or portion thereof from a collection or library of antibodies and/or portions thereof, based on determination of the activity or property of an antibody or portion thereof. Screening can be performed in any of a variety of ways, including, for example, by assays assessing direct binding (e.g. binding affinity) of the antibody to a target protein or by functional assays assessing modulation of an activity of a target protein.
[0270] As used herein, activity towards a target protein refers to binding specificity and/or modulation of a functional activity of a target protein, or other measurements that reflects the activity of an antibody or portion thereof towards a target protein.
[0271] As used herein the term assessing is intended to include quantitative and qualitative determination in the sense of obtaining an absolute value for the binding of an antibody or portion thereof with a target protein and/or modulation of an activity of a target protein by an antibody or portion thereof, and also of obtaining an index, ratio, percentage, visual or other value indicative of the level of the binding or activity. Assessment can be direct or indirect. For example, binding can be determined by directly labeling of an antibody or portion thereof with a detectable label and/or by using a secondary antibody that itself is labeled. In addition, functional activities can be determined using any of a variety of assays known to one of skill in the art, for example, proliferation, cytotoxicity and others as described herein, and comparing the activity of the target protein in the presence versus the absence of an antibody or portion thereof.
[0272] As used herein, a "target protein" refers to candidate proteins or peptides that are specifically recognized by an antibody or portion thereof and/or whose activity is modulated by an antibody or portion thereof. A target protein includes any peptide or protein that contains an epitope for antibody recognition. Target proteins include proteins involved in the etiology of a disease or disorder by virtue of expression or activity. Exemplary target proteins are described herein.
[0273] As used herein, "Hit" refers to an antibody or portion thereof identified, recognized or selected as having an activity in a screening assay.
[0274] As used herein, "iterative" with respect to screening means that the screening is repeated a plurality of times, such as 2, 3, 4, 5 or more times, until a "Hit" is identified whose activity is optimized or improved compared to prior iterations.
[0275] As used herein, "high-throughput" refers to a large-scale method or process that permits manipulation of large numbers of molecules or compounds, generally tens to hundred to thousands of compounds. For example, methods of purification and screening can be rendered high-throughput. High-throughput methods can be performed manually. Generally, however, high-throughput methods involve automation, robotics or software.
[0276] As used herein, "structure/activity relationship (SAR)" refers to the relationship between structure and function of a molecule. For purposes herein, structure is with reference to sequence, for example, a sequence of nucleotides encoding an antibody. By virtue of addressing library members, the identify of each antibody by its sequence is known based on its address. Hence, structure is known and can be correlated to a particular activity. Hence, SAR can be used to assess the affects of changes in structure on an activity.
[0277] As used herein, "functional activity" refer to activities of a polypeptide (e.g. target protein) or portion thereof associated with a full-length (complete) protein. Functional activities include, but are not limited to, biological activity, catalytic or enzymatic activity, antigenicity (ability to bind to or compete with a polypeptide for binding to an anti-polypeptide antibody), immunogenicity, ability to form multimers, the ability to specifically bind to a receptor or ligand for the polypeptide and signaling and downstream effector functions. For purposes herein, modulation (i.e. activation or inhibition) of a functional activity of a polypeptide by an antibody or portion thereof in the libraries herein means that a functional activity of the polypeptide is changed or altered in the presence of the antibody compared to the absence of the antibody or portion thereof.
[0278] As used herein, "modulate" or "modulation" and other various grammatical forms thereof with reference to the effect of an antibody or portion thereof on the functional activity of a target protein refers to increased activity such as induction or potentiation of activity, as well as inhibition of one or more activities of the target protein. Hence, modulation can include an increase in the activity (i.e., up-regulation or agonist activity) a decrease in activity (i.e., down-regulation or inhibition) or any other alteration in an activity (such as a change in periodicity, frequency, duration, kinetics or other parameter). Modulation can be context dependent and typically modulation is compared to a designated state, for example, the wildtype protein, the protein in a constitutive state, or the protein as expressed in a designated cell type or condition. The functional activity of a target protein by an antibody or portion thereof can be modulated by 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90% or more compared to the activity of the target protein in the absence of the antibody or portion thereof.
[0279] As used herein, "agonist" refers to an antibody or portion thereof that modulates signal transduction or other functional activity of a receptor by potentiating, inducing or otherwise enhancing the signal transduction activity or other functional activity of a receptor. Agonists can modulate signal transduction or other functional activity when used alone or can alter signal transduction or other functional activity in the presence of the natural ligand of the receptor or other receptor stimulator to enhance signaling by the receptor compared to the ligand alone.
[0280] As used herein, "antagonist" refers to an antibody or portion thereof that modulates signal transduction or other functional activity of a receptor by blocking or decreasing the signal transduction activity or other functional activity of a receptor
[0281] As used herein, a label is a detectable marker that can be attached or linked directly or indirectly to a molecule or associated therewith. The detection method can be any method known in the art.
[0282] As used herein, binding activity refer to characteristics of a molecule, e.g. a polypeptide, relating to whether or not, and how, it binds one or more binding partners. Binding activities include ability to bind the binding partner(s), the affinity with which it binds to the binding partner (e.g. high affinity), the avidity with which it binds to the binding partner, the strength of the bond with the binding partner and specificity for binding with the binding partner.
[0283] As used herein, "affinity" or "binding affinity" refers to the strength with which an antibody molecule or portion thereof binds to an epitope on a target protein or antigen. Affinity is often measured by equilibrium association constant (K.sub.A) or equilibrium dissociation constant (K.sub.D). Low-affinity antibody-antigen interaction is weak, and the molecules tend to dissociate rapidly, while high affinity antibody-antigen binding is strong and the molecules remain bound for a longer amount of time. A high antibody affinity means that the antibody specifically binds to a target protein with an equilibrium association constant (K.sub.A) of greater than or equal to about 10.sup.6 M.sup.-1, greater than or equal to about 10.sup.7 M.sup.-1, greater than or equal to about 10.sup.8 M.sup.-1, or greater than or equal to about 10.sup.9 M.sup.-1, 10.sup.10 M.sup.-1, 10.sup.11 M.sup.-1 or 10.sup.12 M.sup.-1. Antibodies also can be characterized by an equilibrium dissociation constant (K.sub.D), for example, 10.sup.-4 M, 10.sup.-5 M, 10.sup.-6 M, 10.sup.-7 M, 10.sup.-8 M, 10.sup.-10 M, 10.sup.-11 M or 10.sup.-12 M or lower. Generally, antibodies having a nanomolar or sub-nanomolar dissociation constant are deemed to be high affinity antibodies. Such affinities can be readily determined using conventional techniques, such as by equilibrium dialysis; by using the BIAcore 2000 instrument, using general procedures outlined by the manufacturer; by radioimmunoassay using radiolabeled target antigen; or by another method known to the skilled artisan. The affinity data can be analyzed, for example, by the method of Scatchard et al., Ann N.Y. Acad. ScL, 51:660 (1949).
[0284] As used herein, "epitope" refers to the localized region on the surface of an antigen or protein that is recognized by an antibody. Peptide epitopes include those that are continuous epitopes or discontinuous epitopes. An epitope is generally determined by the three dimensional structure of a protein as opposed to the linear amino acid sequence.
[0285] As used herein, "epitope mapping" is the process of identification of the molecular determinants for antibody-antigen recognition.
[0286] As used herein, Basic Local Alignment Search Tool (BLAST) is a search algorithm developed by Altschul et al (1990) to separately search protein or DNA databases, for example, based on sequence identity. For example, blastn is a program that compares a nucleotide query sequence against a nucleotide sequence database (e.g. GenBank). BlastP is a program that compares an amino acid query sequence against a protein sequence database.
[0287] As used herein, a BLAST bit score is a value calculated from the number of gaps and substitutions associated with each aligned sequence. The higher the score, the more significant the alignment.
[0288] As used herein, a human protein is one encoded by a nucleic acid molecule, such as DNA, present in the genome of a human, including all allelic variants and conservative variations thereof. A variant or modification of a protein is a human protein if the modification is based on the wildtype or prominent sequence of a human protein.
[0289] As used herein, "naturally occurring amino acids" refer to the 20 L-amino acids that occur in polypeptides. The residues are those 20 .alpha.-amino acids found in nature which are incorporated into protein by the specific recognition of the charged tRNA molecule with its cognate mRNA codon in humans.
[0290] As used herein, non-naturally occurring amino acids refer to amino acids that are not genetically encoded. For example, a non-natural amino acid is an organic compound that has a structure similar to a natural amino acid but has been modified structurally to mimic the structure and reactivity of a natural amino acid. Non-naturally occurring amino acids thus include, for example, amino acids or analogs of amino acids other than the 20 naturally-occurring amino acids and include, but are not limited to, the D-isostereomers of amino acids. Exemplary non-natural amino acids are known to those of skill in the art.
[0291] As used herein, nucleic acids include DNA, RNA and analogs thereof, including peptide nucleic acids (PNA) and mixtures thereof. Nucleic acids can be single or double-stranded. When referring to probes or primers, which are optionally labeled, such as with a detectable label, such as a fluorescent or radiolabel, single-stranded molecules are contemplated. Such molecules are typically of a length such that their target is statistically unique or of low copy number (typically less than 5, generally less than 3) for probing or priming a library. Generally a probe or primer contains at least 14, 16 or 30 contiguous nucleotides of sequence complementary to or identical to a gene of interest. Probes and primers can be 10, 20, 30, 50, 100 or more nucleic acids long.
[0292] As used herein, a peptide refers to a polypeptide that is from 2 to 40 amino acids in length.
[0293] As used herein, the amino acids which occur in the various sequences of amino acids provided herein are identified according to their known, three-letter or one-letter abbreviations (Table 1). The nucleotides which occur in the various nucleic acid fragments are designated with the standard single-letter designations used routinely in the art.
[0294] As used herein, an "amino acid" is an organic compound containing an amino group and a carboxylic acid group. A polypeptide contains two or more amino acids. For purposes herein, amino acids include the twenty naturally-occurring amino acids, non-natural amino acids and amino acid analogs (i.e., amino acids wherein the .alpha.-carbon has a side chain).
[0295] As used herein, "amino acid residue" refers to an amino acid formed upon chemical digestion (hydrolysis) of a polypeptide at its peptide linkages. The amino acid residues described herein are presumed to be in the "L" isomeric form. Residues in the "D" isomeric form, which are so designated, can be substituted for any L-amino acid residue as long as the desired functional property is retained by the polypeptide. NH.sub.2 refers to the free amino group present at the amino terminus of a polypeptide. COOH refers to the free carboxy group present at the carboxyl terminus of a polypeptide. In keeping with standard polypeptide nomenclature described in J. Biol. Chem., 243: 3557-3559 (1968), and adopted 37 C.F.R. '.sctn..sctn. 1.821-1.822, abbreviations for amino acid residues are shown in Table 1:
TABLE-US-00001 TABLE 1 Table of Correspondence SYMBOL 1-Letter 3-Letter AMINO ACID Y Tyr Tyrosine G Gly Glycine F Phe Phenylalanine M Met Methionine A Ala Alanine S Ser Serine I Ile Isoleucine L Leu Leucine T Thr Threonine V Val Valine P Pro Proline K Lys Lysine H His Histidine Q Gln Glutamine E Glu Glutamic acid Z Glx Glu and/or Gln W Trp Tryptophan R Arg Arginine D Asp Aspartic acid N Asn Asparagine B Asx Asn and/or Asp C Cys Cysteine X Xaa Unknown or other
[0296] It should be noted that all amino acid residue sequences represented herein by formulae have a left to right orientation in the conventional direction of amino-terminus to carboxyl-terminus. In addition, the phrase "amino acid residue" is broadly defined to include the amino acids listed in the Table of Correspondence (Table 1) and modified and unusual amino acids, such as those referred to in 37 C.F.R. .sctn..sctn. 1.821-1.822, and incorporated herein by reference. Furthermore, it should be noted that a dash at the beginning or end of an amino acid residue sequence indicates a peptide bond to a further sequence of one or more amino acid residues, to an amino-terminal group such as NH.sub.2 or to a carboxyl-terminal group such as COOH. The abbreviations for any protective groups, amino acids and other compounds, are, unless indicated otherwise, in accord with their common usage, recognized abbreviations, or the IUPAC-IUB Commission on Biochemical Nomenclature (see, (1972) Biochem. 11:1726). Each naturally occurring L-amino acid is identified by the standard three letter code (or single letter code) or the standard three letter code (or single letter code) with the prefix "L-"; the prefix "D-" indicates that the stereoisomeric form of the amino acid is D.
[0297] As used herein, an isokinetic mixture is one in which the molar ratios of amino acids has been adjusted based on their reported reaction rates (see, e.g., Ostresh et al, (1994) Biopolymers 34:1681).
[0298] As used herein, modification is in reference to modification of a sequence of amino acids of a polypeptide or a sequence of nucleotides in a nucleic acid molecule and includes deletions, insertions, and replacements of amino acids and nucleotides, respectively. Methods of modifying a polypeptide are routine to those of skill in the art, such as by using recombinant DNA methodologies.
[0299] As used herein, suitable conservative substitutions of amino acids are known to those of skill in this art and can be made generally without altering the biological activity of the resulting molecule. Those of skill in this art recognize that, in general, single amino acid substitutions in non-essential regions of a polypeptide do not substantially alter biological activity (see, e.g., Watson et al. Molecular Biology of the Gene, 4th Edition, 1987, The Benjamin/Cummings Pub. co., p. 224). Such substitutions can be made in accordance with those set forth in TABLE 2 as follows:
TABLE-US-00002 TABLE 2 Original residue Exemplary conservative substitution Ala (A) Gly; Ser Arg (R) Lys Asn (N) Gln; His Cys (C) Ser Gln (Q) Asn Glu (E) Asp Gly (G) Ala; Pro His (H) Asn; Gln Ile (I) Leu; Val Leu (L) Ile; Val Lys (K) Arg; Gln; Glu Met (M) Leu; Tyr; Ile Phe (F) Met; Leu; Tyr Ser (S) Thr Thr (T) Ser Trp (W) Tyr Tyr (Y) Trp; Phe Val (V) Ile; Leu
Other substitutions also are permissible and can be determined empirically or in accord with known conservative substitutions.
[0300] As used herein, a DNA construct is a single or double stranded, linear or circular DNA molecule that contains segments of DNA combined and juxtaposed in a manner not found in nature. DNA constructs exist as a result of human manipulation, and include clones and other copies of manipulated molecules.
[0301] As used herein, a DNA segment is a portion of a larger DNA molecule having specified attributes. For example, a DNA segment encoding a specified polypeptide is a portion of a longer DNA molecule, such as a plasmid or plasmid fragment, which, when read from the 5' to 3' direction, encodes the sequence of amino acids of the specified polypeptide.
[0302] As used herein, the term "nucleic acid" refers to single-stranded and/or double-stranded polynucleotides such as deoxyribonucleic acid (DNA), and ribonucleic acid (RNA) as well as analogs or derivatives of either RNA or DNA. Also included in the term "nucleic acid" are analogs of nucleic acids such as peptide nucleic acid (PNA), phosphorothioate DNA, and other such analogs and derivatives or combinations thereof. Nucleic acid can refer to polynucleotides such as deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). The term also includes, as equivalents, derivatives, variants and analogs of either RNA or DNA made from nucleotide analogs, single (sense or antisense) and double-stranded polynucleotides. Deoxyribonucleotides include deoxyadenosine, deoxycytidine, deoxyguanosine and deoxythymidine. For RNA, the uracil base is uridine.
[0303] As used herein, "nucleic acid molecule encoding" refers to a nucleic acid molecule which directs the expression of a specific protein or peptide. The nucleic acid sequences include both the DNA strand sequence that is transcribed into RNA and the RNA sequence that is translated into protein or peptide. The nucleic acid molecule includes both the full length nucleic acid sequences as well as non-full length sequences derived from the full length mature polypeptide, such as for example a full length polypeptide lacking a precursor sequence. For purposes herein, a nucleic acid sequence also includes the degenerate codons of the native sequence or sequences which can be introduced to provide codon preference in a specific host.
[0304] As used herein, the term "polynucleotide" refers to an oligomer or polymer containing at least two linked nucleotides or nucleotide derivatives, including a deoxyribonucleic acid (DNA), a ribonucleic acid (RNA), and a DNA or RNA derivative containing, for example, a nucleotide analog or a "backbone" bond other than a phosphodiester bond, for example, a phosphotriester bond, a phosphoramidate bond, a phosphorothioate bond, a thioester bond, or a peptide bond (peptide nucleic acid). The term "oligonucleotide" also is used herein essentially synonymously with "polynucleotide," although those in the art recognize that oligonucleotides, for example, PCR primers, generally are less than about fifty to one hundred nucleotides in length.
[0305] Polynucleotides can include nucleotide analogs, including, for example, mass modified nucleotides, which allow for mass differentiation of polynucleotides; nucleotides containing a detectable label such as a fluorescent, radioactive, luminescent or chemiluminescent label, which allow for detection of a polynucleotide; or nucleotides containing a reactive group such as biotin or a thiol group, which facilitates immobilization of a polynucleotide to a solid support. A polynucleotide also can contain one or more backbone bonds that are selectively cleavable, for example, chemically, enzymatically or photolytically. For example, a polynucleotide can include one or more deoxyribonucleotides, followed by one or more ribonucleotides, which can be followed by one or more deoxyribonucleotides, such a sequence being cleavable at the ribonucleotide sequence by base hydrolysis. A polynucleotide also can contain one or more bonds that are relatively resistant to cleavage, for example, a chimeric oligonucleotide primer, which can include nucleotides linked by peptide nucleic acid bonds and at least one nucleotide at the 3' end, which is linked by a phosphodiester bond or other suitable bond, and is capable of being extended by a polymerase. Peptide nucleic acid sequences can be prepared using well-known methods (see, for example, Weiler et al. Nucleic acids Res. 25: 2792-2799 (1997)).
[0306] As used herein, "similarity" between two proteins or nucleic acids refers to the relatedness between the sequence of amino acids of the proteins or the nucleotide sequences of the nucleic acids. Similarity can be based on the degree of identity and/or homology of sequences of residues and the residues contained therein. Methods for assessing the degree of similarity between proteins or nucleic acids are known to those of skill in the art. For example, in one method of assessing sequence similarity, two amino acid or nucleotide sequences are aligned in a manner that yields a maximal level of identity between the sequences. "Identity" refers to the extent to which the amino acid or nucleotide sequences are invariant. Alignment of amino acid sequences, and to some extent nucleotide sequences, also can take into account conservative differences and/or frequent substitutions in amino acids (or nucleotides). Conservative differences are those that preserve the physico-chemical properties of the residues involved. Alignments can be global (alignment of the compared sequences over the entire length of the sequences and including all residues) or local (the alignment of a portion of the sequences that includes only the most similar region or regions).
[0307] "Identity" per se has an art-recognized meaning and can be calculated using published techniques. (See, e.g.: Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D. W., ed., Academic Press, New York, 1993; Computer Analysis of Sequence Data, Part I, Griffin, A. M., and Griffin, H. G., eds., Humana Press, New Jersey, 1994; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; and Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M Stockton Press, New York, 1991). While there exists a number of methods to measure identity between two polynucleotide or polypeptides, the term "identity" is well known to skilled artisans (Carillo, H. & Lipton, D., SIAM J Applied Math 48:1073 (1988)).
[0308] As used herein, homologous (with respect to nucleic acid and/or amino acid sequences) means about greater than or equal to 25% sequence homology, typically greater than or equal to 25%, 40%, 50%, 60%, 70%, 80%, 85%, 90% or 95% sequence homology; the precise percentage can be specified if necessary. For purposes herein the terms "homology" and "identity" are often used interchangeably, unless otherwise indicated. In general, for determination of the percentage homology or identity, sequences are aligned so that the highest order match is obtained (see, e.g.: Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D. W., ed., Academic Press, New York, 1993; Computer Analysis of Sequence Data, Part I, Griffin, A. M., and Griffin, H. G., eds., Humana Press, New Jersey, 1994; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; and Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M Stockton Press, New York, 1991; Carillo et al. (1988) SIAM J Applied Math 48:1073). By sequence homology, the number of conserved amino acids is determined by standard alignment algorithms programs, and can be used with default gap penalties established by each supplier. Substantially homologous nucleic acid molecules hybridize typically at moderate stringency or at high stringency all along the length of the nucleic acid of interest. Also contemplated are nucleic acid molecules that contain degenerate codons in place of codons in the hybridizing nucleic acid molecule.
[0309] Whether any two molecules have nucleotide sequences or amino acid sequences that are at least 60%, 70%, 80%, 85%, 90%, 95%, 96%, 97%, 98% or 99% "identical" or "homologous" can be determined using known computer algorithms such as the "FASTA" program, using for example, the default parameters as in Pearson et al. (1988) Proc. Natl. Acad. Sci. USA 85:2444 (other programs include the GCG program package (Devereux, J., et al., Nucleic Acids Research 12(I):387 (1984)), BLASTP, BLASTN, FASTA (Atschul, S. F., et al., J Molec Biol 215:403 (1990)); Guide to Huge Computers, Martin J. Bishop, ed., Academic Press, San Diego, 1994, and Carillo et al (1988) SIAM J Applied Math 48:1073). For example, the BLAST function of the National Center for Biotechnology Information database can be used to determine identity. Other commercially or publicly available programs include, DNAStar "MegAlign" program (Madison, Wis.) and the University of Wisconsin Genetics Computer Group (UWG) "Gap" program (Madison Wis.). Percent homology or identity of proteins and/or nucleic acid molecules can be determined, for example, by comparing sequence information using a GAP computer program (e.g., Needleman et al. (1970) J. Mol. Biol. 48:443, as revised by Smith and Waterman ((1981) Adv. Appl. Math. 2:482). Briefly, the GAP program defines similarity as the number of aligned symbols (i.e., nucleotides or amino acids), which are similar, divided by the total number of symbols in the shorter of the two sequences. Default parameters for the GAP program can include: (1) a unary comparison matrix (containing a value of 1 for identities and 0 for non-identities) and the weighted comparison matrix of Gribskov et al. (1986) Nucl. Acids Res. 14:6745, as described by Schwartz and Dayhoff, eds., ATLAS OF PROTEIN SEQUENCE AND STRUCTURE, National Biomedical Research Foundation, pp. 353-358 (1979); (2) a penalty of 3.0 for each gap and an additional 0.10 penalty for each symbol in each gap; and (3) no penalty for end gaps.
[0310] Therefore, as used herein, the term "identity" or "homology" represents a comparison between a test and a reference polypeptide or polynucleotide. As used herein, the term at least "90% identical to" refers to percent identities from 90 to 99.99 relative to the reference nucleic acid or amino acid sequence of the polypeptide. Identity at a level of 90% or more is indicative of the fact that, assuming for exemplification purposes a test and reference polypeptide length of 100 amino acids are compared. No more than 10% (i.e., 10 out of 100) of the amino acids in the test polypeptide differs from that of the reference polypeptide. Similar comparisons can be made between test and reference polynucleotides. Such differences can be represented as point mutations randomly distributed over the entire length of a polypeptide or they can be clustered in one or more locations of varying length up to the maximum allowable, e.g. 10/100 amino acid difference (approximately 90% identity). Differences are defined as nucleic acid or amino acid substitutions, insertions or deletions. At the level of homologies or identities above about 85-90%, the result should be independent of the program and gap parameters set; such high levels of identity can be assessed readily, often by manual alignment without relying on software.
[0311] As used herein, a polypeptide containing a specified percentage of amino acids set forth in a reference polypeptide refers to the proportion of contiguous identical amino acids shared between a polypeptide and a reference polypeptide. For example, an isoform that contains 70% of the amino acids set forth in a reference polypeptide having a sequence of amino acids set forth in SEQ ID NO:XX, which recites 147 amino acids, means that the reference polypeptide contains at least 103 contiguous amino acids set forth in the amino acid sequence of SEQ ID NO:XX.
[0312] As used herein, an aligned sequence refers to the use of homology (similarity and/or identity) to align corresponding positions in a sequence of nucleotides or amino acids. Typically, two or more sequences that are related by 50% or more identity are aligned. An aligned set of sequences refers to 2 or more sequences that are aligned at corresponding positions and can include aligning sequences derived from RNAs, such as ESTs and other cDNAs, aligned with genomic DNA sequence.
[0313] As used herein, "primer" refers to a nucleic acid molecule that can act as a point of initiation of template-directed DNA synthesis under appropriate conditions (e.g., in the presence of four different nucleoside triphosphates and a polymerization agent, such as DNA polymerase, RNA polymerase or reverse transcriptase) in an appropriate buffer and at a suitable temperature. It will be appreciated that a certain nucleic acid molecules can serve as a "probe" and as a "primer." A primer, however, has a 3' hydroxyl group for extension. A primer can be used in a variety of methods, including, for example, polymerase chain reaction (PCR), reverse-transcriptase (RT)-PCR, RNA PCR, LCR, multiplex PCR, panhandle PCR, capture PCR, expression PCR, 3' and 5' RACE, in situ PCR, ligation-mediated PCR and other amplification protocols.
[0314] As used herein, "primer pair" refers to a set of primers that includes a 5' (upstream) primer that hybridizes with the 5' end of a sequence to be amplified (e.g. by PCR) and a 3' (downstream) primer that hybridizes with the complement of the 3' end of the sequence to be amplified.
[0315] As used herein, "specifically hybridizes" refers to annealing, by complementary base-pairing, of a nucleic acid molecule (e.g. an oligonucleotide) to a target nucleic acid molecule. Those of skill in the art are familiar with in vitro and in vivo parameters that affect specific hybridization, such as length and composition of the particular molecule. Parameters particularly relevant to in vitro hybridization further include annealing and washing temperature, buffer composition and salt concentration. Exemplary washing conditions for removing non-specifically bound nucleic acid molecules at high stringency are 0.1.times.SSPE, 0.1% SDS, 65.degree. C., and at medium stringency are 0.2.times.SSPE, 0.1% SDS, 50.degree. C. Equivalent stringency conditions are known in the art. The skilled person can readily adjust these parameters to achieve specific hybridization of a nucleic acid molecule to a target nucleic acid molecule appropriate for a particular application.
[0316] As used herein, substantially identical to a product means sufficiently similar so that the property of interest is sufficiently unchanged so that the substantially identical product can be used in place of the product.
[0317] As used herein, it also is understood that the terms "substantially identical" or "similar" varies with the context as understood by those skilled in the relevant art.
[0318] As used herein, an allelic variant or allelic variation references any of two or more alternative forms of a gene occupying the same chromosomal locus. Allelic variation arises naturally through mutation, and can result in phenotypic polymorphism within populations. Gene mutations can be silent (no change in the encoded polypeptide) or can encode polypeptides having altered amino acid sequence. The term "allelic variant" also is used herein to denote a protein encoded by an allelic variant of a gene. Typically the reference form of the gene encodes a wildtype form and/or predominant form of a polypeptide from a population or single reference member of a species. Typically, allelic variants, which include variants between and among species typically have at least 80%, 90% or greater amino acid identity with a wildtype and/or predominant form from the same species; the degree of identity depends upon the gene and whether comparison is interspecies or intraspecies. Generally, intraspecies allelic variants have at least about 80%, 85%, 90% or 95% identity or greater with a wildtype and/or predominant form, including 96%, 97%, 98%, 99% or greater identity with a wildtype and/or predominant form of a polypeptide. Reference to an allelic variant herein generally refers to variations n proteins among members of the same species.
[0319] As used herein, "allele," which is used interchangeably herein with "allelic variant" refers to alternative forms of a gene or portions thereof. Alleles occupy the same locus or position on homologous chromosomes. When a subject has two identical alleles of a gene, the subject is said to be homozygous for that gene or allele. When a subject has two different alleles of a gene, the subject is said to be heterozygous for the gene. Alleles of a specific gene can differ from each other in a single nucleotide or several nucleotides, and can include substitutions, deletions and insertions of nucleotides. An allele of a gene also can be a form of a gene containing a mutation.
[0320] As used herein, species variants refer to variants in polypeptides among different species, including different mammalian species, such as mouse and human.
[0321] As used herein, a splice variant refers to a variant produced by differential processing of a primary transcript of genomic DNA that results in more than one type of mRNA.
[0322] As used herein, the term promoter means a portion of a gene containing DNA sequences that provide for the binding of RNA polymerase and initiation of transcription. Promoter sequences are commonly, but not always, found in the 5' non-coding region of genes.
[0323] As used herein, isolated or purified polypeptide or protein or biologically-active portion thereof is substantially free of cellular material or other contaminating proteins from the cell or tissue from which the protein is derived, or substantially free from chemical precursors or other chemicals when chemically synthesized. Preparations can be determined to be substantially free if they appear free of readily detectable impurities as determined by standard methods of analysis, such as thin layer chromatography (TLC), gel electrophoresis and high performance liquid chromatography (HPLC), used by those of skill in the art to assess such purity, or sufficiently pure such that further purification does not detectably alter the physical and chemical properties, such as enzymatic and biological activities, of the substance. Methods for purification of the compounds to produce substantially chemically pure compounds are known to those of skill in the art. A substantially chemically pure compound, however, can be a mixture of stereoisomers. In such instances, further purification might increase the specific activity of the compound.
[0324] The term substantially free of cellular material includes preparations of proteins in which the protein is separated from cellular components of the cells from which it is isolated or recombinantly-produced. In one embodiment, the term substantially free of cellular material includes preparations of protease proteins having less that about 30% (by dry weight) of non-protease proteins (also referred to herein as a contaminating protein), generally less than about 20% of non-protease proteins or 10% of non-protease proteins or less that about 5% of non-protease proteins. When the protease protein or active portion thereof is recombinantly produced, it also is substantially free of culture medium, i.e., culture medium represents less than about or at 20%, 10% or 5% of the volume of the protease protein preparation.
[0325] As used herein, the term substantially free of chemical precursors or other chemicals includes preparations of protease proteins in which the protein is separated from chemical precursors or other chemicals that are involved in the synthesis of the protein. The term includes preparations of protease proteins having less than about 30% (by dry weight) 20%, 10%, 5% or less of chemical precursors or non-protease chemicals or components.
[0326] As used herein, synthetic, with reference to, for example, a synthetic nucleic acid molecule or a synthetic gene or a synthetic peptide refers to a nucleic acid molecule or polypeptide molecule that is produced by recombinant methods and/or by chemical synthesis methods.
[0327] As used herein, production by recombinant means by using recombinant DNA methods means the use of the well known methods of molecular biology for expressing proteins encoded by cloned DNA.
[0328] As used herein, vector (or plasmid) refers to discrete elements that are used to introduce a heterologous nucleic acid into cells for either expression or replication thereof. The vectors typically remain episomal, but can be designed to effect integration of a gene or portion thereof into a chromosome of the genome. Also contemplated are vectors that are artificial chromosomes, such as yeast artificial chromosomes and mammalian artificial chromosomes. Selection and use of such vehicles are well known to those of skill in the art.
[0329] As used herein, an expression vector includes vectors capable of expressing DNA that is operatively linked with regulatory sequences, such as promoter regions, that are capable of effecting expression of such DNA fragments. Such additional segments can include promoter and terminator sequences, and optionally can include one or more origins of replication, one or more selectable markers, an enhancer, a polyadenylation signal, and the like. Expression vectors are generally derived from plasmid or viral DNA, or can contain elements of both. Thus, an expression vector refers to a recombinant DNA or RNA construct, such as a plasmid, a phage, recombinant virus or other vector that, upon introduction into an appropriate host cell, results in expression of the cloned DNA. Appropriate expression vectors are well known to those of skill in the art and include those that are replicable in eukaryotic cells and/or prokaryotic cells and those that remain episomal or those which integrate into the host cell genome.
[0330] As used herein, vector also includes "virus vectors" or "viral vectors." Viral vectors are engineered viruses that are operatively linked to exogenous genes to transfer (as vehicles or shuttles) the exogenous genes into cells.
[0331] As used herein, operably or operatively linked when referring to DNA segments means that the segments are arranged so that they function in concert for their intended purposes, e.g., transcription initiates in the promoter and proceeds through the coding segment to the terminator.
[0332] As used herein, biological sample refers to any sample obtained from a living or viral source and includes any cell type or tissue of a subject from which nucleic acid or protein or other macromolecule can be obtained. Biological samples include, but are not limited to, body fluids, such as blood, plasma, serum, cerebrospinal fluid, synovial fluid, urine and sweat, tissue and organ samples from animals and plants. Also included are soil and water samples and other environmental samples, viruses, bacteria, fungi, algae, protozoa and components thereof. Hence bacterial and viral and other contamination of food products and environments can be assessed. The methods herein are practiced using biological samples and in some embodiments, such as for profiling, also can be used for testing any sample.
[0333] As used herein, macromolecule refers to any molecule having a molecular weight from the hundreds up to the millions. Macromolecules include peptides, proteins, nucleotides, nucleic acids, and other such molecules that are generally synthesized by biological organisms, but can be prepared synthetically or using recombinant molecular biology methods.
[0334] As used herein, the term "biopolymer" is a biological molecule, including macromolecules, composed of two or more monomeric subunits, or derivatives thereof, which are linked by a bond or a macromolecule. A biopolymer can be, for example, a polynucleotide, a polypeptide, a carbohydrate, or a lipid, or derivatives or combinations thereof, for example, a nucleic acid molecule containing a peptide nucleic acid portion or a glycoprotein, respectively. Biopolymers include, but are not limited to, nucleic acids, proteins, polysaccharides, lipids and other macromolecules. Nucleic acids include DNA, RNA, and fragments thereof. Nucleic acids can be derived from genomic DNA, RNA, mitochondrial nucleic acid, chloroplast nucleic acid and other organelles with separate genetic material.
[0335] As used herein, a biomolecule is any compound found in nature, or derivatives thereof. Biomolecules include, but are not limited to: oligonucleotides, oligonucleosides, proteins, peptides, amino acids, peptide nucleic acids (PNAs), oligosaccharides and monosaccharides.
[0336] As used herein, a biological particle refers to a virus, such as a viral vector or viral capsid with or without packaged nucleic acid, phage, including a phage vector or phage capsid, with or without encapsulated nucleic acid, a single cell, including eukaryotic and prokaryotic cells or fragments thereof, a liposome or micellar agent or other packaging particle, and other such biological materials.
[0337] As used herein, a composition refers to any mixture. It can be a solution, a suspension, liquid, powder, a paste, aqueous, non-aqueous or any combination thereof.
[0338] As used herein, a combination refers to any association between or among two or more items. The combination can be two or more separate items, such as two compositions or two collections, can be a mixture thereof, such as a single mixture of the two or more items, or any variation thereof.
[0339] As used herein, kit refers to a packaged combination, optionally including instructions and/or reagents for their use.
[0340] As used herein, fluid refers to any composition that can flow. Fluids thus encompass compositions that are in the form of semi-solids, pastes, solutions, aqueous mixtures, gels, lotions, creams and other such compositions.
[0341] As used herein, antigenic means that a polypeptide induce an immune response. Highly antigenic polypeptides are those that reproducibly and predictably induce an immune response.
[0342] As used herein, a pharmaceutical effect or therapeutic effect refers to an effect observed upon administration of an agent intended for treatment of a disease or disorder or for amelioration of the symptoms thereof.
[0343] As used herein, "disease or disorder" refers to a pathological condition in an organism resulting from cause or condition including, but not limited to, infections, acquired conditions, genetic conditions, and characterized by identifiable symptoms. Diseases and disorders of interest herein are those involving a specific target protein including those mediated by a target protein and those in which a target protein plays a role in the etiology or pathology. Exemplary target proteins and associated diseases and disorders are described elsewhere herein.
[0344] As used herein, "treating" a subject with a disease or condition means that the subject's symptoms are partially or totally alleviated, or remain static following treatment. Hence treatment encompasses prophylaxis, therapy and/or cure. Prophylaxis refers to prevention of a potential disease and/or a prevention of worsening of symptoms or progression of a disease. Treatment also encompasses any pharmaceutical use of a modified interferon and compositions provided herein.
[0345] As used herein, a therapeutic agent, therapeutic regimen, radioprotectant, or chemotherapeutic mean conventional drugs and drug therapies, including vaccines, which are known to those skilled in the art. Radiotherapeutic agents are well known in the art.
[0346] As used herein, treatment means any manner in which the symptoms of a condition, disorder or disease or other indication, are ameliorated or otherwise beneficially altered.
[0347] As used herein therapeutic effect means an effect resulting from treatment of a subject that alters, typically improves or ameliorates the symptoms of a disease or condition or that cures a disease or condition. A therapeutically effective amount refers to the amount of a composition, molecule or compound which results in a therapeutic effect following administration to a subject.
[0348] As used herein, the term "subject" refers to an animal, including a mammal, such as a human being.
[0349] As used herein, a patient refers to a human subject.
[0350] As used herein, amelioration of the symptoms of a particular disease or disorder by a treatment, such as by administration of a pharmaceutical composition or other therapeutic, refers to any lessening, whether permanent or temporary, lasting or transient, of the symptoms that can be attributed to or associated with administration of the composition or therapeutic.
[0351] As used herein, prevention or prophylaxis refers to methods in which the risk of developing disease or condition is reduced.
[0352] As used herein, an effective amount is the quantity of a therapeutic agent necessary for preventing, curing, ameliorating, arresting or partially arresting a symptom of a disease or disorder.
[0353] As used herein, administration refers to any method in which an antibody or portion thereof is contacted with its target protein. Administration can be effected in vivo or ex vivo or in vitro. For example, for ex vivo administration a body fluid, such as blood, is removed from a subject and contacted outside the body with the antibody or portion thereof. For in vivo administration, the antibody or portion thereof can be introduced into the body, such as by local, topical, systemic and/or other route of introduction. In vitro administration encompasses methods, such as cell culture methods.
[0354] As used herein, unit dose form refers to physically discrete units suitable for human and animal subjects and packaged individually as is known in the art.
[0355] As used herein, a single dosage formulation refers to a formulation for direct administration.
[0356] As used herein, an "article of manufacture" is a product that is made and sold. As used throughout this application, the term is intended to encompass compiled germline antibodies or antibodies obtained therefrom contained in articles of packaging.
[0357] As used herein, fluid refers to any composition that can flow. Fluids thus encompass compositions that are in the form of semi-solids, pastes, solutions, aqueous mixtures, gels, lotions, creams and other such compositions.
[0358] As used herein, animal includes any animal, such as, but are not limited to primates including humans, gorillas and monkeys; rodents, such as mice and rats; fowl, such as chickens; ruminants, such as goats, cows, deer, sheep; ovine, such as pigs and other animals. Non-human animals exclude humans as the contemplated animal. The germline segments, and resulting antibodies, provided herein are from any source, animal, plant, prokaryotic and fungal. Most germline segments, and resulting antibodies, are of animal origin, including mammalian origin.
[0359] As used herein, a control refers to a sample that is substantially identical to the test sample, except that it is not treated with a test parameter, or, if it is a sample plasma sample, it can be from a normal volunteer not affected with the condition of interest. A control also can be an internal control.
[0360] As used herein, the singular forms "a," "an" and "the" include plural referents unless the context clearly dictates otherwise. Thus, for example, reference to a compound, comprising "an extracellular domain" includes compounds with one or a plurality of extracellular domains.
[0361] As used herein, ranges and amounts can be expressed as "about" a particular value or range. About also includes the exact amount. Hence "about 5 bases" means "about 5 bases" and also "5 bases."
[0362] As used herein, "optional" or "optionally" means that the subsequently described event or circumstance does or does not occur, and that the description includes instances where said event or circumstance occurs and instances where it does not. For example, an optionally substituted group means that the group is unsubstituted or is substituted.
[0363] As used herein, the abbreviations for any protective groups, amino acids and other compounds, are, unless indicated otherwise, in accord with their common usage, recognized abbreviations, or the IUPAC-IUB Commission on Biochemical Nomenclature (see, (1972) Biochem. 11:1726).
B. OVERVIEW
[0364] Provided are methods for generating combinatorial libraries (i.e. collections) of functional antibodies, and the resulting libraries. The collections or libraries provided are addressable, where antibodies within each address have the same sequence, are known a priori, and are different from the antibodies at each other address in the collection. The collections can be provided as physical arrays or the members can be otherwise identified so that they can be sorted. The arrayed collections of antibodies can represent the complete repertoire of combined germline portions, a selected portion thereof, or collections of modified forms thereof. The members of the libraries are individually designed and addressed. Because of this, the libraries are highly diverse, permitting creation of libraries with far fewer members than other libraries, but having higher diversity. The libraries provided herein contain as few as 10.sup.2 members and typically contain about or 10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5 and more unique members, including about or 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9 and more unique members.
[0365] The collections of antibodies are addressable, such as in arrays or other addressable format, such that each member is identifiable and each locus has antibodies that are the same or that have the same binding specificity at each locus. The locus can be a physical locus or can be otherwise identifiable and sortable, such as an RF tag, attachment to label with a bar code, attachment to a chemical tag, in manners described for chemical libraries.
[0366] In contrast, other antibody libraries are produced such that they contain mixtures of antibodies at a locus, or contain unidentified members of libraries. Exemplary of such libraries are those described in any of the following: European Patent Application Nos. EP0368684 and EP89311731; International Published Patent Application Nos. WO92/001047, WO 02/38756, WO 97/08320, WO 2005/023993, WO 07/137616 and WO 2007/054816; U.S. Pat. Nos. 6,593,081 and 6,989,250; United States Published Patent Application No. US 2002/0102613, US 2003/153038, US 2003/0022240, US 2005/0119455, US 2005/0079574 and US 2006/0234302; and Orlandi et al. (1989) Proc Natl. Acad. Sci. U.S.A., 86:3833-3837; Ward et al. (1989) Nature, 341:544-546; Huse et al. (1989) Science, 246:1275-1281; Burton et al. (1991) Proc. Natl. Acad. Sci., U.S.A., 88:10134-10137; Marks et al. (1991) J Mol Biol, 222:581-591; Hoogenboom et al. (1991) J Mol Biol, 227:381-388; Nissim et al. (1994) EMBO J, 13:692-698; Barbas et al. (1992) Proc. Natl. Acad. Sci., U.S.A., 89:4457-4461; Akamatsu et al. (1993) J. Immunol., 151:4651-1659; Griffiths et al. (1994) EMBO J, 13:3245-3260; Fellouse (2004) PNAS, 101:12467-12472; Persson et al. (2006) J. Mol. Biol. 357:607-620; Knappik et al. (2000) J. Mol. Biol. 296:57-86; Rothe et al. (2008) J. Mol. Biol. 376:1182-1200; Mondon et al (2008) Frontiers in Bioscience, 13:1117-1129; and Behar, I (2007) Expert Opin. Ther., 7:763-779.
[0367] Although many of these libraries contain large numbers of members (e.g. 10.sup.8-10.sup.10 members), there is no mechanism to ensure that all members are functional, nor to maximize diversity nor to represent the complete repertoire of germline sequences or a selected portion thereof. Thus, the composition and diversity of the library is not optimal. For example, many existing libraries are developed by PCR amplification of germline sequences. PCR amplification introduces errors into the resulting amplified products. In addition, in some methods hybrid primers are used to facilitate recombination of individual V(D)J segments. This can result in recombination events that are "out-of-frame" resulting in non-functional members. Also, in practicing such methods, members either are pooled (such as in a tube or via phage display) and screened together for binding to a target substrate or are introduced into host cells as mixtures and then colonies are individually picked and grown. Upon identification of a positive interaction or other selected events, any "Hits" must be further characterized in order to be identified.
[0368] The combinatorial addressable libraries of antibodies provided herein do not share these problems. Each member of the collection is addressed, such that each member occupies a unique locus, for example, a spatial array or other array or other identifiable address (e.g., presentation in well-plates; being bound to a support or chip, bar-coded, color-coded, RF-tag labeled support or other such addressable format). Displaying members on an address is facilitated because each member is individually generated, and thus the sequence of each member is known. Display of the members can be achieved on any desired format, which permits screening the members not only for binding but also for function. The "Hits" can be quickly identified coincident with the screening results. Hence, structure/activity relationships (SAR) between members of the collections can be performed to identify similarities in characteristics or sequences between and among identified "Hits". Pharmacokinetics and dose-responses also are available on screening or immediately following "Hit" identification. Further optimization of "Hits" can be performed such as by mutagenesis and iterative screening. Accordingly, the methods provided herein for generating addressable combinatorial antibody collections and the resulting collections offer a robust alternative to identification of antibodies with desired specificities and/or activities, for example, for use as therapeutic antibodies.
[0369] 1. Methods of Generating Addressable Combinatorial Antibody Collections
[0370] In one example of the methods provided herein, variable heavy (VH) and variable light (VL) chain members of the libraries are generated, recombinantly or synthetically by DNA synthesis, from known germline antibody sequences or modified sequences thereof. Thus, the members can represent the entire repertoire of the naive germline, and are not restricted based on selection against "self" proteins. Combinatorial diversity in the collection exists from recombination (e.g. such as performed in silico by computer software as described herein) of individual V, D and J segments that make up the variable heavy chain and of individual V (V.sub..kappa. or V.sub..lamda.) and J (J.sub..kappa., or J.sub..lamda.) segments that make up the variable light chains (see FIG. 1). The sequences can be joined together by random recombination of all combinations of V(D)J segments generating maximal combinatorial diversity. Alternatively, the V(D)J segments can be recombined using rational or semi-rational approaches such that a specific sequence or subset of sequences are used in generating the members of the library. For example, germline segments can be selected based on sequence similarities or differences or based on shared characteristics (e.g., a V region family, CDR3 length or composition or other biochemical attribute).
[0371] In the methods herein, the combinatorial diversity of the resulting members is optimized for functioning sequences that encode a full length polypeptide. Although all combinations of V(D)J segments can be recombined, the joints in the compiled sequences between the different V(D)J sequences are selected so that the resulting sequences are in-frame. Each functioning member occupies an address (e.g. a position in a well or chip) of a collection. In vivo, however, junctional diversity exists upon V(D)J recombination such that nucleotides are often inserted at junction regions, which can result in new amino acids at the junctions. Hence, in some example of the methods herein, the resulting in-frame members can be subjected to mutagenesis, for example, to introduce diversity at the junction regions (e.g., junctional diversity). In such examples, each locus can contain a pool of antibodies with the same V(D)J segments, but differing from each other by one or more mutations (e.g. insertions, deletions or replacements of amino acids).
[0372] In addition to generating naive antibody libraries, the methods provided herein can be used to generate directed antibody libraries, whereby the resulting members are optimized against known targets. For example, the starting sequences of individual V(D)J segments of heavy and light chains can be generated to contain a known binding peptide against a target. The goal of such a library format is to generate a platform that allows the discovery of agonist or antagonist antibodies that mimic therapeutic targets, for example, growth factors, cytokines, hormones or other cellular activators.
[0373] Generally, the members of the collections provided herein contain all or a portion of a variable light (VL) and variable heavy (VH) chain, so long as the resulting antibody is sufficient to form an antigen binding site. Hence, in addition to combinatorial diversity, diversity in the collections provided herein is achieved by pairing diversity by combining heavy and light chains (FIG. 1). Thus, the individually recombined VL and VH chains as discussed above can themselves be paired in varying combinations to generate a scalable collection of all possible combinations of VL and VH chains, or a subset thereof. For example, a library can be generated where all members have the same VH chains, but are paired with all or a subset of the individually recombined VL members. The heavy and light chain members can be paired by direct or indirect linkage. The resulting pairs of heavy and light chains can be presented in any desired library format, such as a complete antibody, Fab forms, single chain (sc) Fv forms, or any multimeric form thereof.
[0374] 2. The Resulting Libraries
[0375] Provided herein are libraries of nucleic acid molecules encoding VL chains and libraries of nucleic acid molecules encoding VH chains. Also provided herein are combinatorial antibody libraries that are paired antibody libraries containing at a minimum all or a portion of a VL and VH chains, such that each resulting member in the library is sufficient to form an antigen binding site. The libraries can be naive libraries representing all or a portion of all possible germline antibodies, or can be modified forms thereof. The resulting members of the paired antibody collections include, but are not limited to, Fab, single chain (sc) Fv, disulfide-stabilized Fv and multimeric formats such as minibodies, bis-scFv, diabodies, triabodies and tetrabodies. The libraries provided herein differ from existing antibody collections because each individual member of the collection is known and, in the case of the antibody libraries, each member is "productive" or "functional" because the encoding nucleic acid molecules lack stop codons that can otherwise truncate the resulting protein before a full length polypeptide can be produced. Typically, all libraries provided herein are in an addressable format, such that the identity of each member of the library is known based on its locus or "address". Exemplary of antibody collections provided herein are combinatorial Fab libraries, such as addressable combinatorial Fab libraries. Any of the above libraries can include 10.sup.2, 10.sup.3, 10.sup.4 or 10.sup.5, or more different members.
[0376] 3. Applications of the Libraries
[0377] The resulting libraries can be used for any application or purpose as desired. Because of their diversity, specificity and effector functions, antibodies are attractive candidates for protein-based therapeutics. Thus, the libraries can be used in methods of screening for various activities to identify antibodies with unique functions, such as for use as therapeutic antibodies. For example, the antibody libraries provided herein can be used in screening assays based on function or binding against unknown or known targets. In particular, it is contemplated herein that the resulting libraries can be used in functional assays, such as cell based assays, to discover new MAbs (e.g., Fabs) against selected targets. Hence, the libraries provided herein offer advantages over existing libraries because they permit identification of antibodies that perhaps are low affinity binders, but functionally are ideal therapeutic antibody candidates. Accordingly, both agonist and antagonist antibodies can be easily discovered.
[0378] The resulting identified "Hits" can be further optimized against a desired target by iterative screening methods of antibody discovery (FIGS. 2A-2B). For example, antibody "Hits" identified from binding or activity-based screening assays, can be used to generate further libraries containing V(D)J germline segments related (e.g., by sequence identity or similarity) to the V(D)J germline segments in the identified Hit(s). By virtue of the fact that the collections are arrayed, such that the identity of each individual member in the collection is known, iterative approaches can be used for the rapid expansion of "Hits" to identify antibody "Hits" with improved therapeutic applications. In addition, antibody "Hits" can be used as a scaffold for mutagenesis to generate modified heavy and light chains, and for successive generations of libraries of nucleic acid molecules encoding modified VL chains, libraries of nucleic acid molecules encoding modified VH chains and antibody libraries minimally containing modified VL and VH chains.
[0379] Finally, antibody "Hits" identified from the libraries herein and/or further optimized by iterative screening and/or other mutagenesis methods, can be used in a variety of in vitro and in vivo applications by virtue of the specificity for one or more target proteins. For example, the antibodies can be used in diagnostic methods. In another example, the antibodies can be used in methods of treatment and other uses for treating a disease or disorder which is associated with expression or activation of a particular target protein, and for which the antibody can modulate.
[0380] The following sections describe exemplary components of the methods and libraries, the methods of generating combinatorial antibody libraries, including arrayed libraries, the resulting libraries and applications of the libraries.
C. ANTIBODIES
[0381] Provided herein are methods of generating libraries of addressable combinatorial antibodies, and the resulting libraries and antibodies. The antibodies in the libraries minimally include all or a portion of a variable heavy chain (VH) and/or a variable light (VL) chain so long as the antibody contains a sufficient antibody binding site. For example, the VH and VL chains of the antibodies provided herein typically include one or more, generally two or more, and up to all of the three CDRs making up the antigen binding site. In some examples, the antibodies can be generated to contain a synthetic CDR, whereby a peptide against a known target is grafted into the CDR regions of the variable region to effect directed binding and activation of the target (see e.g., Frederickson et al. (2006) PNAS 103: 14307-14312).
[0382] Optionally, the antibodies can include all or a portion of the constant heavy chain (e.g. one or more CH domains such as CH1, CH2, CH3 and CH4 and/or a constant heavy chain (CL)). Hence, the antibodies included in the libraries herein include those that are full-length antibodies, and also include fragments or portions thereof including, for example, Fab, Fab', F(ab').sub.2, single-chain Fvs (scFv), Fv, dsFv, diabody, Fd and Fd' fragments Fab fragments, Fd fragments, scFv fragments, and scFab fragments. For example, antibodies in the libraries provided herein include Fabs.
[0383] A description of the structure, sequence and function of antibodies are known to one of skill in the art and one of skill in the art is familiar with the mechanisms that give rise to diversity in the germline. It is contemplated herein that libraries of combinatorial antibodies can be made by recombination of germline DNA sequences that mimic the process of germline recombination during B cell differentiation. Such recombination can be performed in silico (e.g., by a computer) as described herein or can be performed manually using molecular biology techniques. Recombined sequences can be individually generated, such as by DNA synthesis or by recombinant DNA techniques, to generate all permutations of variable heavy and light chain sequences. The antibodies can be expressed in any desired form, and in some instances, pairing of variable and constant regions can be achieved. The result is that the libraries of combinatorial antibodies provided herein can represent the entire naive antibody repertoire or a subset thereof.
[0384] 1. Antibody Polypeptides
[0385] Antibodies are produced naturally by B cells in membrane-bound and secreted forms. Antibodies specifically recognize and bind antigen epitopes through cognate interactions. Antibody binding to cognate antigens can initiate multiple effector functions, which cause neutralization and clearance of toxins, pathogens and other infectious agents. Diversity in antibody specificity arises naturally due to recombination events during B cell development. Through these events, various combinations of multiple antibody V, D and J gene segments, which encode variable regions of antibody molecules, are joined with constant region genes to generate a natural antibody repertoire with large numbers of diverse antibodies. A human antibody repertoire contains more than 10.sup.10 different antigen specificities and thus theoretically can specifically recognize any foreign antigen. Antibodies include such naturally produced antibodies, as well as synthetically, i.e. recombinantly, produced antibodies, such as antibody fragments.
[0386] In folded antibody polypeptides, binding specificity is conferred by antigen binding site domains, which contain portions of heavy and/or light chain variable region domains. Other domains on the antibody molecule serve effector functions by participating in events such as signal transduction and interaction with other cells, polypeptides and biomolecules. These effector functions cause neutralization and/or clearance of the infecting agent recognized by the antibody.
[0387] 2. Antibody Structural and Functional Domains A full-length antibody contains four polypeptide chains, two identical heavy (H) chains (each usually containing about 440 amino acids) and two identical light (L) chains (each containing about 220 amino acids). The light chains exist in two distinct forms called kappa (.kappa.) and lambda (.lamda.). Each chain is organized into a series of domains organized as immunoglobulin (Ig) domains, including variable (V) and constant (C) region domains. Light chains have two domains, corresponding to the C region (CL) and the V region (VL). Heavy chains have four domains, the V region (VH) and three or four domains in the C region (CH1, CH2, CH3 and CH4), and, in some cases, hinge region. The four chains (two heavy and two light) are held together by a combination of covalent and non-covalent (disulfide) bonds.
[0388] Antibodies include those that are full-lengths and those that are fragments thereof, namely Fab, Fab', F(ab').sub.2, single-chain Fvs (scFv), Fv, dsFv, diabody, Fd and Fd' fragments. The fragments include those that are in single-chain or dimeric form. The Fv fragment, which contains only the VH and VL domain, is the smallest immunoglobulin fragment that retains the whole antigen-binding site (see, for example, Methods in Molecular Biology, Vol 207: Recombinant Antibodies for Cancer Therapy Methods and Protocols (2003); Chapter 1; p 3-25, Kipriyanov). Stabilization of Fv are achieved by direct linkage of the VH and VL chains, such as for example, by linkage with peptides (to generate single-chain Fvs (scFv)), disulfide bridges or knob-into-hole mutations. Fab fragments, in contrast, are stable because of the presence of the CH1 and CL domains that hold together the variable chains. Fd antibodies, which contain only the VH domain, lack a complete antigen-binding site and can be insoluble.
[0389] 3. Antibody Sequence and Specificity
[0390] The variable region of the heavy and light chains are encoded by multiple germline gene segments separated by non-coding regions, or introns, and often are present on different chromosomes. For example, the genes for the immunoglobulin heavy chain region contains approximately 65 variable (V.sub.H) genes, 27 Diversity (D.sub.H) genes, and 6 Joining (J.sub.H) genes. The kappa (.kappa.) and lambda (.lamda.) light chains are also each encoded by a similar number of V.sub.L and J.sub.L gene segments, but do not include any D gene segments. Exemplary V.sub.H, D.sub.H, J.sub.H and V.sub.L (V.sub..kappa. or V.sub..lamda.) and J.sub.L (J.sub..kappa. or J.sub..lamda.) germline gene segments are set forth in Tables 3-5.
[0391] During B cell differentiation germline DNA is rearranged whereby one D.sub.H and one J.sub.H gene segment of the heavy chain locus are recombined, which is followed by the joining of one V.sub.H gene segment forming a rearranged VDJ gene that encodes a VH chain. The rearrangement occurs only on a single heavy chain allele by the process of allelic exclusion. Allelic exclusion is regulated by in-frame or "productive" recombination of the VDJ segments, which occurs in only about one-third of VDJ recombinations of the variable heavy chain. When such productive recombination events first occur in a cell, this results in production of a .mu. heavy chain that gets expressed on the surface of a pre-B cell and transmits a signal to shut off further heavy chain recombination, thereby preventing expression of the allelic heavy chain locus. The surface-expressed .mu. heavy chain also acts to activate the kappa (.kappa.) locus for rearrangement. The lambda (.lamda.) locus is only activated for rearrangement if the .kappa. recombination is unproductive on both loci. The light chain rearrangement events are similar to heavy chain, except that only the V.sub.L and J.sub.L segments are recombined. Before primary transcription of each, the corresponding constant chain gene is added. Subsequent transcription and RNA splicing leads to mRNA that is translated into an intact light chain or heavy chain.
[0392] The variable regions of antibodies confer antigen binding and specificity due to recombination events of individual germline V, D and J segments, whereby the resulting recombined nucleic acid sequences encoding the variable region domains differ among antibodies and confer antigen-specificity to a particular antibody. The variation, however, is limited to three complementarity determining regions (CDR1, CDR2, and CDR3) found within the N-terminal domain of the heavy (H) and (L) chain variable regions. The CDRs are interspersed with regions that are more conserved, termed "framework regions" (FR). The extent of the framework region and CDRs has been precisely defined (see e.g., Kabat, E. A. et al. (1991) Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242, and Chothia, C. et al. (1987) J. Mol. Biol. 196:901-917). Each VH and VL is typically composed of three CDRs and four FRs arranged from the amino terminus to carboxy terminus in the following order: FR1, CDR1, FR2, CDR2, FR3, CDR3 and FR4. Sequence variability among VL and VH domains is generally limited to the CDRs, which are the regions that form the antigen binding site. For example, for the heavy chain, generally, V.sub.H genes encode the N-terminal three framework regions, the first two complete CDRs and the first part of the third CDR; the D.sub.H gene encodes the central portion of the third CDR, and the J.sub.H gene encodes the last part of the third CDR and the fourth framework region. For the light chain, the V.sub.L genes encode the first CDR and second CDR. The third CDR (CDRL3) is formed by the joining of the V.sub.L and J.sub.L gene segments. Hence, CDRs 1 and 2 are exclusively encoded by germline V gene segment sequences. The VH and VL chain CDR3s form the center of the Ag-binding site, with CDRs 1 and 2 form the outside boundaries; the FRs support the scaffold by orienting the H and L CDRs. On average, an antigen binding site typically requires that at least four of the CDRs make contact with the antigen's epitope, with CDR3 of both the heavy and light chain being the most variable and contributing the most specificity to antigen binding (see e.g., Janis Kuby, Immunology, Third Edition, New York, W.H. Freeman and Company, 1998, pp. 115-118). CDRH3, which includes all of the D gene segment, is the most diverse component of the Ab-binding site, and typically plays a critical role in defining the specificity of the Ab. In addition to sequence variation, there is variation in the length of the CDRs between the heavy and light chains (see Table 26 in Example 12).
[0393] The constant regions, on the other hand, are encoded by sequences that are more conserved among antibodies. These domains confer functional properties to antibodies, for example, the ability to interact with cells of the immune system and serum proteins in order to cause clearance of infectious agents. Different classes of antibodies, for example IgM, IgD, IgG, IgE and IgA, have different constant regions, allowing them to serve distinct effector functions.
[0394] These natural recombination events of V, D, and J, can provide nearly 2.times.10.sup.7 different antibodies with both high affinity and specificity. Additional diversity is introduced by nucleotide insertions and deletions in the joining segments and also by somatic hypermutation of V regions. The result is that there are approximately 10.sup.10 antibodies present in an individual with differing antigen specificities.
[0395] The methods provided herein take advantage of the mechanisms responsible for generating diversity between and among germline antibodies, thereby permitting generation of collections of antibodies that can be tested for varied functional or other properties.
D. METHODS OF GENERATING MEMBERS OF THE COMBINATORIAL ANTIBODY LIBRARY
[0396] Provided herein are methods of producing combinatorial antibody libraries and the resulting libraries. Typically, each member in the library contains a variable heavy chain and a variable light chain, or portions thereof sufficient to form an antigen binding site. In the methods provided herein, each antibody member of the library is generated by mimicking natural recombination events by combining known V(D)J gene segment sequences (e.g. from publicly available databases of germline sequences), or modified forms thereof, in various permutations, in-frame, to generate a plurality of nucleic acid sequences encoding functional VH and VL chains. For example, in the steps of the methods nucleic acid molecules encoding the variable heavy (VH) chain are generated by recombining individual V, D and J segments. Nucleic acid molecules encoding the variable light (VL) chain are generated by recombining individual V (V.sub..kappa. or V.sub..lamda.) and J (J.sub..kappa., or J.sub..lamda.) segments that make up the variable light chains. The segments can be germline segments, or degenerate sequences thereof. In such examples, the resulting antibodies produced by the method are naive antibodies. It is contemplated herein, however, that the method can be performed using any modified form of a known germline segment, for example, to introduce further diversity into the library. For example, the method can be performed using reverse complement (i.e. inverted) sequences of D.sub.H germline segments. The process of recombining germline segments in-frame can be performed manually using molecular biology techniques or in silico (e.g. using a computer programmed to perform an algorithm).
[0397] In the methods, the recombination is effected so that each gene segment is in-frame, such that resulting recombined nucleic acid molecules encodes a functional VH or VL polypeptide. Also, in the methods, each nucleic acid molecule is individually generated and synthesized. In the methods, resulting members of the library are produced by co-expression of nucleic acid molecules encoding the recombined variable region genes together, such that when expressed, a combinatorial antibody member is generated minimally containing a VH and VL chain, or portions thereof. In some examples of the methods, the nucleic acid molecule encoding the VH and VL chain can be expressed as a single nucleic acid molecule, whereby the genes encoding the heavy and light chain are joined by a linker. In another example of the methods, the nucleic acid molecules encoding the VH and VL chain can be separately provided for expression together. Thus, upon expression from the recombined nucleic acid molecules, each different member of the library represents a germline encoded antibody, whereby diversity is achieved by combinatorial diversity of V(D)J segments and pairing diversity of heavy and light chains. In the method, additional diversity can be introduced into the library using any of a number of approaches known in the art, including but not limited to, random mutagenesis, semi-rational or rational mutagenesis.
[0398] One or more or all steps of the method can be performed in an addressable format, such that the identity of each member in the process is known by its location at an addressed locus. Hence, provided herein are addressable libraries of germline recombined nucleic acid sequences encoding VH chains, addressable libraries of germline recombined nucleic acid sequences encoding VL chains, and addressable libraries formed by combinations of nucleic acids molecules encoding VL chains and nucleic acid molecules encoding VH chains at each locus. Also provided are addressable cells, each cell at a locus containing different combinations of a recombined nucleic acid molecule encoding a VL and a recombined nucleic acid encoding a VH. Resulting antibody libraries also can be addressable. Such addressable antibody libraries permit rapid identification of "Hits" and assessment of structure/activity relationships between and among "Hits." The resulting libraries of antibodies can be screened for a variety of activities, such as but not limited to binding, proliferation, cytotoxicity and low affinity leads against difficult antigens, such as self antigens, ion channels, G-protein coupled receptors, novel epitopes, non-protein antigens and the discovery of agonist antibodies.
[0399] 1. Methods for Producing Functional Recombined Germline Variable Region Genes
[0400] Provided herein is a method for generating nucleic acid molecules generated by recombination of germline segments or modified forms thereof, each nucleic acid molecule encoding a different and functional variable region of the heavy or light chains. Variable gene segments include V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda.. Germline segments can be selected from but not limited to human, mouse, rat, sheep, pig, goat horse, rabbit or dog germline segments. Exemplary germline segments are of human origin.
[0401] a. Variable Gene Segments
[0402] i. Germline Segments
[0403] In practicing the methods herein, germline segment sequences are obtained from any source that provides antibody germlines gene segments. These include any databases or published literature that sets forth sequences of germline gene segments. Exemplary antibody germline sources include but are not limited to databases at the National Center for Biotechnology Information (NCBI), the international ImMunoGeneTics information System.RTM. (IMGT), the Kabat database and the Tomlinson's VBase database (Lefranc (2003) Nucleic Acids Res., 31:307-310; Martin et al., Bioinformatics Tools for Antibody Engineering in Handbook of Therapeutic Antibodies, Wiley-VCH (2007), pp. 104-107). If desired, nucleic acid sequences for non-human germline segments also can be obtained from published literature or publicly available databases. For example, an exemplary mouse germline databases is ABG database available at ibt.unam.mx/vir/v_mice.html. The Sequence Listing provided herein provides sequences of exemplary human germline segment sequences collected from the IMGT database and other public database (see e.g., SEQ ID NOS:10-451 and 868).
[0404] For example, exemplary human Heavy Chain Germline Segments (SEQ ID NO. 10-285) are listed in Table 3. Exemplary human Light Chain Kappa Germline Segments (SEQ ID NO. 286-364 and SEQ ID NO. 868) are listed in Table 4. Exemplary human Light Chain Lambda Germline Segments (SEQ ID NO. 365-451) are listed in Table 5. Germline segments are listed using IMGT gene names and definitions previously approved by the Human Genome Organization (HUGO) nomenclature committee. The segments are named using IMGT nomenclature, whereby the first three letters indicate the locus (IGH, IGK or IGL), the fourth letter represents the gene (e.g., V for V-gene, D for D-gene, J for J-gene), the fifth position indicates the number of the subgroup, followed by a hyphen indicating the gene number classification. For alleles, the IMGT name is followed by an asterisk and a two figure number.
[0405] Tables 6-8 list alternative nomenclature for Human Heavy Chain V Genes, Human Light Chain Kappa V Genes, and Human Light Chain Lambda V Genes, respectively (see e.g. Lefranc, M.-P. Exp Clin Immunogenet, 18:100-116 (2001), Zachau, H. G. Immunologist, 4:49-54 (1996), Lefranc, M.-P. Exp Clin Immunogenet, 18:161-174 (2000), Kawasaki et al, Genome Res, 7:250-261 (1997), Lefranc, M.-P. Exp Clin Immunogenet, 18:242-254 (2001). Any desired naming convention can be used to identify antibody germline segments. One of skill in the art can identify a nucleic acid sequence using any desired naming convention. For purposes herein when describing recombined nucleic acid sequences (see e.g., Table 22), VH germline segments are named using IMGT nomenclature without any allele identified. Table 6 lists the IMGT nomenclature and corresponding IMGT nomenclature with the respective allele. VK germline segments are named using Zachau nomenclature. Table 7 lists the Zachau nomenclature and the corresponding IMGT nomenclature. VL germline segments are identified using Kawasaki nomenclature. Table 8 lists the Kawasaki nomenclature and the corresponding IMGT nomenclature. DH, JH, JK and JL germline segments are named using IMGT nomenclature.
TABLE-US-00003 TABLE 3 Human Heavy Chain Germline Segments SEQ ID NO. V Segments 10 IGHV1-18*01 11 IGHV1-18*02 12 IGHV1-2*01 13 IGHV1-2*02 14 IGHV1-2*03 15 IGHV1-2*04 16 IGHV1-24*01 17 IGHV1-3*01 18 IGHV1-3*02 19 IGHV1-45*01 20 IGHV1-45*02 21 IGHV1-45*03 22 IGHV1-46*01 23 IGHV1-46*02 24 IGHV1-46*03 25 IGHV1-58*01 26 IGHV1-58*02 27 IGHV1-69*01 28 IGHV1-69*02 29 IGHV1-69*03 30 IGHV1-69*04 31 IGHV1-69*05 32 IGHV1-69*06 33 IGHV1-69*07 34 IGHV1-69*08 35 IGHV1-69*09 36 IGHV1-69*10 37 IGHV1-69*11 38 IGHV1-69*12 39 IGHV1-69*13 40 IGHV1-8*01 41 IGHV1-c*01 42 IGHV1-f*01 43 IGHV1-f*02 44 IGHV2-26*01 45 IGHV2-5*01 46 IGHV2-5*02 47 IGHV2-5*03 48 IGHV2-5*04 49 IGHV2-5*05 50 IGHV2-5*06 51 IGHV2-5*07 52 IGHV2-5*08 53 IGHV2-5*09 54 IGHV2-5*10 55 IGHV2-70*01 56 IGHV2-70*02 57 IGHV2-70*03 58 IGHV2-70*04 59 IGHV2-70*05 60 IGHV2-70*06 61 IGHV2-70*07 62 IGHV2-70*08 63 IGHV2-70*09 64 IGHV2-70*10 65 IGHV2-70*11 66 IGHV2-70*12 67 IGHV2-70*13 68 IGHV3-11*01 69 IGHV3-11*03 70 IGHV3-13*01 71 IGHV3-13*02 72 IGHV3-13*02 73 IGHV3-15*01 74 IGHV3-15*02 75 IGHV3-15*03 76 IGHV3-15*04 77 IGHV3-15*05 78 IGHV3-15*06 79 IGHV3-15*07 80 IGHV3-15*08 81 IGHV3-16*01 82 IGHV3-16*02 83 IGHV3-20*01 84 IGHV3-21*01 85 IGHV3-21*02 86 IGHV3-23*01 87 IGHV3-23*02 88 IGHV3-23*03 89 IGHV3-23*04 90 IGHV3-23*05 91 IGHV3-30*01 92 IGHV3-30*02 93 IGHV3-30*03 94 IGHV3-30*04 95 IGHV3-30*05 96 IGHV3-30*06 97 IGHV3-30*07 98 IGHV3-30*08 99 IGHV3-30*09 100 IGHV3-30*10 101 IGHV3-30*11 102 IGHV3-30*12 103 IGHV3-30*13 104 IGHV3-30*14 105 IGHV3-30*15 106 IGHV3-30*16 107 IGHV3-30*17 108 IGHV3-30*18 109 IGHV3-30*19 110 IGHV3-30-3*01 111 IGHV3-30-3*02 112 IGHV3-33*01 113 IGHV3-33*02 114 IGHV3-33*03 115 IGHV3-33*04 116 IGHV3-33*05 117 IGHV3-35*01 118 IGHV3-38*01 119 IGHV3-38*02 120 IGHV3-43*01 121 IGHV3-43*02 122 IGHV3-48*01 123 IGHV3-48*02 124 IGHV3-48*03 125 IGHV3-49*01 126 IGHV3-49*02 127 IGHV3-49*03 128 IGHV3-49*04 129 IGHV3-49*05 130 IGHV3-53*01 131 IGHV3-53*02 132 IGHV3-53*03 133 IGHV3-64*01 134 IGHV3-64*02 135 IGHV3-64*03 136 IGHV3-64*04 137 IGHV3-64*05 138 IGHV3-66*01 139 IGHV3-66*02 140 IGHV3-66*03 141 IGHV3-66*04 142 IGHV3-7*01 143 IGHV3-7*02 144 IGHV3-72*01 145 IGHV3-72*02 146 IGHV3-73*01 147 IGHV3-73*02 148 IGHV3-74*01 149 IGHV3-74*02 150 IGHV3-74*03 151 IGHV3-9*01 152 IGHV3-d*0153 153 IGHV4-28*01 154 IGHV4-28*02 155 IGHV4-28*03 156 IGHV4-28*04 157 IGHV4-28*05 158 IGHV4-30-2*01 159 IGHV4-30-2*02 160 IGHV4-30-2*03 161 IGHV4-30-2*04 162 IGHV4-30-4*01 163 IGHV4-30-4*02 164 IGHV4-30-4*03 165 IGHV4-30-4*04 166 IGHV4-30-4*05 167 IGHV4-30-4*06 168 IGHV4-31*01 169 IGHV4-31*02 170 IGHV4-31*03 171 IGHV4-31*04 172 IGHV4-31*05 173 IGHV4-31*06 174 IGHV4-31*07 175 IGHV4-31*08 176 IGHV4-31*09 177 IGHV4-31*10 178 IGHV4-34*01 179 IGHV4-34*02 180 IGHV4-34*03 181 IGHV4-34*04 182 IGHV4-34*05 183 IGHV4-34*06 184 IGHV4-34*07 185 IGHV4-34*08 186 IGHV4-34*09 187 IGHV4-34*10 188 IGHV4-34*11 189 IGHV4-34*12 190 IGHV4-34*13 191 IGHV4-39*01 192 IGHV4-39*02 193 IGHV4-39*03 194 IGHV4-39*04 195 IGHV4-39*05 196 IGHV4-39*06 197 IGHV4-39*07 V, D, or J Segments 198 IGHV4-4*01 199 IGHV4-4*02 200 IGHV4-4*03 201 IGHV4-4*04 202 IGHV4-4*05 203 IGHV4-4*06 204 IGHV4-4*07 205 IGHV4-59*01 206 IGHV4-59*02 207 IGHV4-59*03 208 IGHV4-59*04 209 IGHV4-59*05 210 IGHV4-59*06 211 IGHV4-59*07 212 IGHV4-59*08 213 IGHV4-59*09 214 IGHV4-59*10 215 IGHV4-61*01 216 IGHV4-61*02 217 IGHV4-61*03 218 IGHV4-61*04 219 IGHV4-61*05 220 IGHV4-61*06 221 IGHV4-61*07 222 IGHV4-61*08 223 IGHV4-b*01 224 IGHV4-b*02 225 IGHV5-51*01 226 IGHV5-51*02 227 IGHV5-51*03 228 IGHV5-51*04 229 IGHV5-51*05 230 IGHV5-a*01 231 IGHV5-a*03 232 IGHV5-a*04 233 IGHV6-1*01 234 IGHV6-1*02 235 IGHV7-4-1*01 236 IGHV7-4-1*02 237 IGHV7-4-1*03 238 IGHV7-81*01 -- D Segments 239 IGHD1-1*01 240 IGHD1-14*01 241 IGHD1-20*01 242 IGHD1-26*01 243 IGHD1-7*01 244 IGHD2-15*01 245 IGHD2-2*01 246 IGHD2-2*02 247 IGHD2-2*03 248 IGHD2-21*01 249 IGHD2-21*02 250 IGHD2-8*01 251 IGHD2-8*02
252 IGHD3-10*01 253 IGHD3-10*02 254 IGHD3-16*01 255 IGHD3-16*02 256 IGHD3-22*01 257 IGHD3-3*01 258 IGHD3-3*02 259 IGHD3-9*01 260 IGHD4-11*01 261 IGHD4-17*01 262 IGHD4-23*01 263 IGHD4-4*01 264 IGHD5-12*01 265 IGHD5-18*01 266 IGHD5-24*01 267 IGHD5-5*01 268 IGHD6-13*01 269 IGHD6-19*01 270 IGHD6-25*01 271 IGHD6-6*01 272 IGHD7-27*01 -- J Segments 273 IGHJ1*01 274 IGHJ2*01 275 IGHJ3*01 276 IGHJ3*02 277 IGHJ4*01 278 IGHJ4*02 279 IGHJ4*03 280 IGHJ5*01 281 IGHJ5*02 282 IGHJ6*01 283 IGHJ6*02 284 IGHJ6*03 285 IGHJ6*04
TABLE-US-00004 TABLE 4 Human Light Chain Kappa Germline Segments SEQ ID NO. V Segments 286 IGKV1-12*01 287 IGKV1-12*02 288 IGKV1-13*02 289 IGKV1-16*01 290 IGKV1-17*01 291 IGKV1-17*02 292 IGKV1-27*01 293 IGKV1-33*01 294 IGKV1-37*01 295 IGKV1-39*01 296 IGKV1-5*01 297 IGKV1-5*02 298 IGKV1-5*03 299 IGKV1-6*01 300 IGKV1-8*01 301 IGKV1-9*01 302 IGKV1-NL1*01 303 IGKV1/OR2-0*01 304 IGKV1/OR2-108*01 305 IGKV1D-12*01 306 IGKV1D-12*02 307 IGKV1D-13*01 308 IGKV1D-16*01 309 IGKV1D-16*02 310 IGKV1D-17*01 311 IGKV1D-33*01 312 IGKV1D-37*01 313 IGKV1D-39*01 314 IGKV1D-42*01 315 IGKV1D-43*01 316 IGKV1D-8*01 317 IGKV2-24*01 318 IGKV2-28*01 319 IGKV2-29*02 320 IGKV2-29*03 321 IGKV2-30*01 322 IGKV2-40*01 323 IGKV2-40*02 324 IGKV2D-24*01 325 IGKV2D-26*01 326 IGKV2D-26*02 327 IGKV2D-28*01 328 IGKV2D-29*01 329 IGKV2D-29*02 330 IGKV2D-30*01 331 IGKV2D-40*01 332 IGKV3-11*01 333 IGKV3-11*02 334 IGKV3-15*01 335 IGKV3-20*01 336 IGKV3-20*02 337 IGKV3-7*01 338 IGKV3-7*02 339 IGKV3-7*03 340 IGKV3-NL1*01 341 IGKV3-NL2*01 V or J Segments 342 IGKV3-NL3*01 343 IGKV3-NL4*01 344 IGKV3-NL5*01 345 IGKV3/OR2-268*01 346 IGKV3/OR2-268*02 347 IGKV3D-11*01 348 IGKV3D-15*01 349 IGKV3D-20*01 350 IGKV3D-7*01 351 IGKV4-1*01 352 IGKV5-2*01 353 IGKV6-21*01 354 IGKV6D-21*01 355 IGKV6D-41*01 868 IGKV1-39*02 -- J Segments 356 IGKJ1*01 357 IGKJ2*01 358 IGKJ2*02 359 IGKJ2*03 360 IGKJ2*04 361 IGKJ3*01 362 IGKJ4*01 363 IGKJ4*02 364 IGKJ5*01
TABLE-US-00005 TABLE 5 Human Light Chain Lambda Germline Segments SEQ ID NO. V Segments 365 IGLV1-36*01 366 IGLV1-40*01 367 IGLV1-40*02 368 IGLV1-40*03 369 IGLV1-41*01 370 IGLV1-44*01 371 IGLV1-47*01 372 IGLV1-47*02 373 IGLV1-50*01 374 IGLV1-51*01 375 IGLV1-51*02 376 IGLV10-54*01 377 IGLV10-54*02 378 IGLV10-54*03 379 IGLV11-55*01 380 IGLV2-11*01 381 IGLV2-11*02 382 IGLV2-11*03 383 IGLV2-14*01 384 IGLV2-14*02 385 IGLV2-14*03 386 IGLV2-14*04 387 IGLV2-18*01 388 IGLV2-18*02 389 IGLV2-18*03 390 IGLV2-18*04 391 IGLV2-23*01 392 IGLV2-23*02 393 IGLV2-23*03 394 IGLV2-33*01 395 IGLV2-33*02 396 IGLV2-33*03 397 IGLV2-8*01 398 IGLV2-8*02 399 IGLV2-8*03 400 IGLV3-1*01 401 IGLV3-10*01 402 IGLV3-10*02 403 IGLV3-12*01 404 IGLV3-12*02 405 IGLV3-16*01 406 IGLV3-19*01 407 IGLV3-21*01 408 IGLV3-21*02 409 IGLV3-21*03 410 IGLV3-22*01 411 IGLV3-25*01 412 IGLV3-25*02 413 IGLV3-25*03 414 IGLV3-27*01 415 IGLV3-32*01 416 IGLV3-9*01 417 IGLV3-9*02 418 IGLV4-3*01 419 IGLV4-60*01 420 IGLV4-60*02 421 IGLV4-60*03 422 IGLV4-69*01 423 IGLV4-69*02 424 IGLV5-37*01 V or J Segments 425 IGLV5-39*01 426 IGLV5-39*02 427 IGLV5-45*01 428 IGLV5-45*02 429 IGLV5-45*03 430 IGLV5-48*01 431 IGLV5-52*01 432 IGLV6-57*01 433 IGLV7-43*01 434 IGLV7-46*01 435 IGLV7-46*02 436 IGLV8-61*01 437 IGLV8-61*02 438 IGLV8-61*03 439 IGLV9-49*01 440 IGLV9-49*02 441 IGLV9-49*03 -- J Segments 442 IGLJ1*01 443 IGLJ2*01 444 IGLJ3*01 445 IGLJ3*02 446 IGLJ4*01 447 IGLJ5*01 448 IGLJ5*02 449 IGLJ6*01 450 IGLJ7*01 451 IGLJ7*02
TABLE-US-00006 TABLE 6 Human Heavy Chain V Genes IMGT Nomenclature SEQ ID IMGT Nomenclature with alleles No. VH1-18 IGHV1-18*01 10 VH1-2 IGHV1-2*02 13 VH1-24 IGHV1-24*01 16 VH1-3 IGHV1-3*02 18 VH1-45 IGHV1-45*02 20 VH1-46 IGHV1-46*01 22 VH1-58 IGHV1-58*02 26 VH1-69 IGHV1-69*06 32 VH1-8 IGHV1-8*01 40 VH2-26 IGHV2-26*01 44 VH2-5 IGHV2-5*01 45 VH2-70 IGHV2-70*13 67 VH3-11 IGHV3-11*01 68 VH3-13 IGHV3-13*01 70 VH3-15 IGHV3-15*01 73 VH3-16 IGHV3-16*02 82 VH3-20 IGHV3-20*01 83 VH3-21 IGHV3-21*01 84 VH3-23 IGHV3-23*01 86 VH3-30 IGHV3-30*03 93 VH3-33 IGHV3-33*01 112 VH3-35 IGHV3-35*01 117 VH3-38 IGHV3-38*02 119 VH3-43 IGHV3-43*01 120 VH3-48 IGHV3-48*02 123 VH3-49 IGHV3-49*03 127 VH3-53 IGHV3-53*01 130 VH3-64 IGHV3-64*02 134 VH3-66 IGHV3-66*03 140 VH3-7 IGHV3-7*01 142 VH3-72 IGHV3-72*01 144 VH3-73 IGHV3-73*02 147 VH3-74 IGHV3-74*01 148 VH3-9 IGHV3-9*01 151 VH4-28 IGHV4-28*01 153 VH4-31 IGHV4-31*02 169 VH4-34 IGHV4-34*01 178 VH4-39 IGHV4-39*01 191 VH4-4 IGHV4-4*07 204 VH4-59 IGHV4-59*01 205 VH4-61 IGHV4-61*08 222 VH5-51 IGHV5-51*03 227 VH6-1 IGHV6-1*01 233 VH7-81 IGHV7-81*01 238
TABLE-US-00007 TABLE 7 Human Light Chain Kappa V Genes Zachau IMGT SEQ ID Nomencalture Nomenclature No. A1 IGKV2D-30*01 330 A10 IGKV6D-21*01 354 A11 IGKV3D-20*01 349 A14 IGKV6D-41*01 355 A17 IGKV2-30*01 321 A18b IGKV2-29*02 319 A19 IGKV2-28*01 318 A2 IGKV2D-29*01 328 A20 IGKV1-27*01 292 A23 IGKV2-24*01 317 A26 IGKV6D-21*01 354 A27 IGKV3-20*01 335 A3 IGKV2D-28*01 327 A30 IGKV1-17*01 290 A5 IGKV2D-26*01 325 A7 IGKV2D-24*01 324 B2 IGKV5-2*01 352 B3 IGKV4-1*01 351 L1 IGKV1-16*01 289 L10 IGKV3-7*01 337 L10a IGKV3-7*02 338 L11 IGKV1-6*01 299 L12 IGKV1-5*01 296 L12a IGKV1-5*03 298 L14 IGKV1D-17*01 310 L15 IGKV1D-16*01 308 L15a IGKV1D-16*02 309 L16 IGKV3D-15*01 348 L18 IGKV1D-13*01 307 L19 IGKV1D-12*01 305 L2 IGKV3-15*01 334 L20 IGKV3D-11*01 347 L22 IGKV1D-42*01 314 L23 IGKV1D-43*01 315 L24 IGKV1D-8*01 316 L25 IGKV3D-7*01 350 L4/18a IGKV1-13*02 288 L5 IGKV1-12*01 286 L6 IGKV3-11*01 332 L8 IGKV1-9*01 301 L9 IGKV1-8*01 300 O1 IGKV2D-40*01 331 O11 IGKV2-40*01 322 O11a IGKV2-40*02 323 O12 IGKV1-39*01 295 O12a IGKV1-39*02 868 O14 IGKV1-37*01 294 O18 IGKV1-33*01 293 O2 IGKV1D-39*01 313 O4 IGKV1D-37*01 312 O8 IGKV1D-33*01 311 Z0 IGKV1/OR2-0*01 303
TABLE-US-00008 TABLE 8 Human Light Chain Lambda V Genes Kawasaki IMGT SEQ ID Nomenclature Nomenclature No. V1-11 IGLV1-36*01 365 V1-13 IGLV1-40*01 366 V1-16 IGLV1-44*01 369 V1-17 IGLV1-47*02 372 V1-18 IGLV1-50*01 373 V1-19 IGLV1-51*01 374 V1-2 IGLV2-8*01 397 V1-20 IGLV10-54*02 377 V1-22 IGLV6-57*01 432 V1-3 IGLV2-11*01 380 V1-4 IGLV2-14*01 383 V1-5 IGLV2-18*01 387 V1-7 IGLV2-23*03 393 V1-9 IGLV2-33*01 394 V2-1 IGLV3-1*01 400 V2-11 IGLV3-16*01 405 V2-13 IGLV3-19*01 406 V2-14 IGLV3-21*02 408 V2-15 IGLV3-22*01 410 V2-17 IGLV3-25*02 412 V2-19 IGLV3-27*01 414 V2-6 IGLV3-9*01 416 V2-7 IGLV3-10*01 401 V2-8 IGLV3-12*02 404 V3-2 IGLV7-43*01 433 V3-3 IGLV7-46*02 435 V3-4 IGLV8-61*01 436 V4-1 IGLV5-37*01 424 V4-2 IGLV5-45*03 429 V4-3 IGLV5-48*01 430 V4-4 IGLV5-52*01 431 V4-6 IGLV11-55*01 379 V5-1 IGLV4-3*01 418 V5-2 IGLV9-49*2 440 V5-4 IGLV4-60*02 420 V5-6 IGLV4-69*01 422
[0406] ii. Modified Germline Segments
[0407] It is contemplated herein that the practice of the method is not limited to germline segment sequences. Hence, any modified V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and or J.sub..lamda. segment sequences, or any sequence analogous thereto, can be used in the practice of the method. By virtue of adding to the repertoire of segment sequences by modification thereto, the diversity of the library and the permutations of compiled segments can be further increased. The germline segments can be modified randomly or empirically. The germline segments can be modified to generate further diversity in the library. Alternatively or in addition, the germline segments can be modified to facilitate generation of the recombined full-length nucleic acid molecules by the introduction of linkers, restriction enzyme sites or other sequences of nucleotides required for practice of the method described herein.
[0408] Generally, the modified germline segments include those that are derived from germline sequences. The germline segments can be modified by introducing mutations into the germline sequence, randomly or empirically, or can be modified to generate germline consensus sequences. For example, modified J.sub.H germline segments are set forth in SEQ ID NOS: 3450-3455.
[0409] In another example, additional modifications of the germline segments include the addition of flanking sequences at one or both of the 5' and 3' termini of individual germline segments that provide restriction sites. Such modifications can be incorporated into the germline sequences using DNA synthesis, or by PCR, for example using primers that incorporate the restriction enzyme sites. In one example, as discussed below, the addition of such restriction sites facilitate joining of germline segments. In some cases, however, modifications of germline segments include the removal of restrictions sites. Restriction sites include any restriction site known in the art. Exemplary restriction site sequences are set forth in Table 15. Generally, the restriction site chosen is compatible with the subsequent compilation of germline segment sequences and can be chosen to facilitate blunt-ended ligation or sticky-ended ligation. The choice of restriction enzyme is routine and is well within the level of one of skill in the art.
[0410] In some examples, sequences of known antibodies, including monoclonal antibodies, particularly therapeutic antibodies, that are derived from germline sequences can be used in the methods herein. Since monoclonal antibodies already have a recognized antigen specificity, it is contemplated herein that incorporation of such derived sequences into the methods will permit the identification of antibodies with improved specificity and functionality against a target antigen. The nucleotide sequences derived from germline sequences, for example corresponding to any one or more of a V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and or J.sub..lamda. can themselves be combined with germline segment sequences. One of skill in the art can identify the corresponding sequences in a nucleic acid molecule encoding a particular antibody that are derived from germline sequences. Table 9 below identifies V, D and J regions that correlate with the derived germline sequences.
TABLE-US-00009 TABLE 9 Exemplary Monoclonal Antibody Sequences (V regions, normal font; D regions, underlined; J regions, boldface) Protein SEQ ID Nucleotide TARGET ANTIBODY HEAVY CHAIN SEQUENCE NO: SEQ ID NO: CD20 Rituxan QVQLQQPGAELVKPGASVKMSCKASGYTFTSYNMHW 1027 1043 (rituximab) VKQTPGRGLEWIGAIYPGNGDTSYNQKFKGKATLTADK SSSTAYMQLSSLTSEDSAVYYCARSTYYGGDVVYFNVW GAGTTVTVSA EGFR Erbitux (cetuximab QVQLKQSGPGLVQPSQSLSITCTVSGFSLTNYGVHWV 1028 1044 RQSPGKGLEWLGVIWSGGNTDYNTPFTSRLSINKDNSK SQVFFKMNSLQSNDTAIYYCARALTYYDYEFAYWGQG TLVTVSA Her2/Neu Herceptin EVQLVESGGGLVQPGGSLRLSCAASGFNIKDTYIHWVR (trastuzumab) QAPGKGLEWVARIYPTNGYTRYADSVKGRFTISADTSK 1029 452 NTAYLQMNSLRAEDTAVYYCSRWGGDGFYAMDYWGQ GTLVTVSS VEGFA Avastin EVQLVESGGGLVQPGGSLRLSCAASGYTFTNYGMNW 1030 1045 (bevacizumab) VRQAPGKGLEWVGWINTYTGEPTYAADFKRRFTFSLD TSKSTAYLQMNSLRAEDTAVYYCAKYPHYYGSSHWYF DVWGQGTLVTVSS VEGFA Lucentis EVQLVESGGGLVQPGGSLRLSCAASGYDFTHYGMNW 1031 1046 (ranibizumab) VRQAPGKGLEWVGWINTYTGEPTYAADFKRRFTFSLD TSKSTAYLQMNSLRAEDTAVYYCAKYPYYYGTSHWYF DVWGQGTLVTVSS CD52 Campath QVQLQESGPGLVRPSQTLSLTCTVSGFTFTDFYMNWV 1032 1047 (alemtuzumab) RQPPGRGLEWIGFIRDKAKGYTTEYNPSVKGRVTMLVD TSKNQFSLRLSSVTAADTAVYYCAREGHTAAPFDYWG QGSLVTVSS TNF.alpha. Humira (adalimumab) EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWV 1033 1048 RQAPGKGLEWVSAITWNSGHIDYADSVEGRFTISRDNA KNSLYLQMNSLRAEDTAVYYCAKVSYLSTASSLDYWG QGTLVTVSS EGFR Vectibix QVQLQESGPGLVKPSETLSLTCTVSGGSVSSGDYYWT 1034 1049 (panitumumab) WIRQSPGKGLEWIGHIYYSGNTNYNPSLKSRLTISIDTS KTQFSLKLSSVTAADTAIYYCVRDRVTGAFDIWGQGTM VTSS Protein SEQ ID Nucleotide TARGET ANTIBODY LIGHT CHAIN SEQUENCE NO: SEQ ID NO: CD20 Rituxan QIVLSQSPAILSASPGEKVTMTCRASSSVSYIHWFQQKP 1035 1050 (rituximab) GSSPKPWIYATSNLASGVPVRFSGSGSGTSYSLTISRV EAEDAATYYCQQVVTSNPPTFGGGTKLEIK EGFR Erbitux (cetuximab DILLTQSPVILSVSPGERVSFSCRASQSIGTNIHWYQQR 1036 1051 TNGSPRLLIKYASESISGIPSRFSGSGSGTDFTLSINSVE SEDIADYYCQQNNNWPTTFGAGTKLELK Her2/Neu Herceptin DIQMTQSPSSLSASVGDRVTITCRASQDVNTAVAWYQ 1037 818 (trastuzumab) QKPGKAPKLLIYSASFLYSGVPSRFSGSRSGTDFTLTIS SLQPEDFATYYCQQHYTTPPTFGQGTKVEIK VEGFA Avastin DIQMTQSPSSLSASVGDRVTITCSASQDISNYLNWYQQ 1038 1052 (bevacizumab) KPGKAPKVLIYFTSSLHSGVPSRFSGSGSGTDFTLTISS LQPEDFATYYCQQYSTVPWTFGQGTKVEIK VEGFA Lucentis DIQLTQSPSSLSASVGDRVTITCSASQDISNYLNWYQQK 1039 1053 (ranibizumab) PGKAPKVLIYFTSSLHSGVPSRFSGSGSGTDFTLTISSL QPEDFATYYCQQYSTVPWTFGQGTKVEIK CD52 Campath DIQMTQSPSSLSASVGDRVTITCKASQNIDKYLNWYQQ 1040 1054 (alemtuzumab) KPGKAPKLLIYNTNNLQTGVPSRFSGSGSGTDFTFTISS LQPEDIATYYCLQHISRPRTFGQGTKVEIK TNF.alpha. Humira (adalimumab) DIQMTQSPSSLSASVGDRVTITCRASQGIRNYLAWYQQ 1041 1055 KPGKAPKLLIYAASTLQSGVPSRFSGSGSGTDFTLTISS LQPEDVATYYCQRYNRAPYTFGQGTKVEIK EGFR Vectibix DIQMTQSPSSLSASVGDRVTITCQASQDISNYLNWYQQ 1042 1056 (panitumumab) KPGKAPKLLIYDASNLETGVPSRFSGSGSGTDFTFTISS LQPEDIATYFCQHFDHLPLAFGGGTKVEIK
[0411] In some examples, the modified germline sequence can include a sequence of nucleotides that replaces all or some of the nucleotides of a germline segment. For example, it is further contemplated herein, that the modified germline sequences designated D.sub.H can be any sequence of nucleotides. The D.sub.H segment of a nucleic acid molecule encodes the central portion of the CDRH3 and is largely responsible for the antigen specificity and variability among antibodies. Since this region is the most variable among antibodies, it can tolerate more modification. Also, it is the region most responsible for antigen specificity. Generally, a segment designated D.sub.H includes 5, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 30, 40, 50, 60, 70, 80, 90, 100 or more nucleotides. The sequence of nucleotides is chosen such that once compiled, in-frame, with a V.sub.H and J.sub.H as discussed below, an antibody molecule or fragment or portion thereof is produced that contains a sufficient antigen-binding site. The nucleotides are chosen randomly or empirically. In some examples, a segment of nucleotides designated D.sub.H can include a random sequence of nucleotides. In other examples, the segment of nucleotides can be selected to be targeted against a specific antigen. For example, the segment of nucleotides can encode a peptide mimetic (see e.g. Table 16 below). In additional examples, a segment of nucleotides designated D.sub.H can include nucleotides that are the reverse complement (i.e. inverted) compared to a known D.sub.H germline segment. This is exemplified in Example 14.
[0412] In other examples, germline segment sequences can be modified to provide a consensus sequence between and among germline segments. Generally, due to the variability between and among the CDR regions, consensus sequences are generated in framework regions. Such modifications aid in the practice of the method by facilitating manipulation of a common sequence, for example, where the method of generating a combinatorial antibody library is performed manually. This is exemplified in Example 1 where each of the J.sub.H contain a common F4 framework region.
[0413] b. Choosing Germline Segments or Modified Segments Thereof
[0414] As described herein above, each VH and VL chain is encoded by a nucleic acid molecule combinatorially generated from gene segments, generally germline segments or modified forms thereof. The members of the resulting library can be chosen by selecting, randomly or empirically, the gene segments that can be recombined. One of skill in the art can select any desired V(D)J gene segment or subsets thereof for recombination to generate in-frame nucleic acid molecules encoding VH or VL.
[0415] In one example, the germline V(D)J segment sequences can be recombined randomly, whereby all known germline sequences (e.g. any described in the Sequence Listing herein or any available in public databases or known to those of skill in the art, and any modified forms thereof) are recombined together in all possible permutations. In such an example, every V.sub.H gene segment is recombined with every D.sub.H which is recombined with every J.sub.H. Similarly, every V.sub.L(.sub..kappa. or .sub..lamda.) is recombined with every J.sub.L(.sub..kappa. or .sub..lamda.). In such an example, the resulting recombined germline nucleic acid molecules represent the complete repertoire of naive VH and VL. For example, if germline segments are recombined based on known germline segment sequences set forth in Tables 3-5, greater then or about 100,000 different recombined nucleic acid molecules encoding VH can be generated, greater then or about 600 different recombined nucleic acids encoding VL.sub..kappa. can be generated, and greater then or about 700 different recombined nucleic acid molecules encoding VL.sub..lamda. can be generated. Thus, libraries of nucleic acids encoding variable heavy and light chains provided herein can encode for every possible recombined antibody variable region. In addition, further diversity can be introduced by modification, such as by mutagenesis, by introducing directed peptides, or by using inverted D.sub.H sequences, as described herein above.
[0416] Alternatively, the V(D)J segments can be recombined using rational or semi-rational approaches such that a specific germline segment sequence or subset of sequences used are restricted in generating the members of the library. For example, as described in Example 14 herein, all members of the library contain a V.sub.H germline segment that is an IGHV3-23*01. In other examples, germline segment sequences can be selected that contain modifications, for example, those that contain mutations to a specific region or region generated randomly (e.g. by site-directed mutagenesis to a particular CDR) or empirically (e.g. modified to contain directed peptide mimetics). By permitting selection of germline segment sequences, the libraries provided herein are versatile and can be rationally designed based on the application of the library.
[0417] For example, antibody germline segments can be selected wherein the resulting nucleic acid sequences are restricted based on sequence similarities or differences or other shared characteristics. For example, germline segment sequences can be selected based on sequence similarities or differences or based on shared characteristics (e.g., a V region family, length, CDR3 length or composition, species, functionality, specificity, group, subgroup, pattern within the CDR, specific amino acids or other biochemical attribute). Antibody structure databases (e.g. CATH database: available at cathwww.biochem.ucl.ac.uk/; SACS database: available at bioinf.org.uk/abs/sacs/; IMGT 3D structure database: available at imgt3d.igh.cnrs.fr/) or other databases are available to sort germline segments based on a selected criteria. Alternatively, such selection can be done manually, for example, using sequence alignments or other manual sorting.
[0418] In one example, germline segments can be selected based on their sequence similarity or differences. One of skill in the art knows or can determine the sequence identity between and among germline segments, and identify germline segments that have a particular sequence identity. In one example, germline segment sequences from one or more of a V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and or J.sub..lamda. group can be selected based on sequence similarity. Sequence similarity between selected segments can include, but is not limited to, 60%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more. For example, subsets of germline segment belonging to the same subgroup or gene family can be selected, which generally share a high degree (e.g. greater then 70%, typically 75% or more) sequence identity. Tables 3-5 above identify germline segments belonging to the same subgroup or gene family. For example, in Table 3, IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6 and IGHV7 each represent a subgroup of a V.sub.H segment, and germline segments within a subgroup share at least 75% sequence identity. Thus, all germline segments in the IGHV1 can be selected, or all germline segments in IGHV2 can be selected, or all germline segments in IGHV3 can be selected, etc. In another example, in Table 3 IGHV1-18*01 and IGHV1-18*02 represent a gene family having germline segments that are alleles. Thus, all germline segments that are related by virtue of being in the same family can be selected as a subset of germline sequences.
[0419] In another example, germline segments can be selected based on sequence differences so that the resulting subset represents a diverse repertoire of sequences. One of skill in the art knows or can identify germline segments that have a particular sequence identity. Sequence differences between selected segments can include those that exhibit 60%, 50%, 40%, 30%, 25%, 20%, 15%, 10%, or less sequence similarity. For example, subsets of germline segments, each from a different subgroup can be selected. Thus, in one example, V.sub.H segments can be selected from each of the IGHV1, IGHV2, IGHV3, IGHV4, IGHV5, IGHV6 and IGHV7 subgroups; for example, a subset can include IGHV1-18*01, IGHV1-26*01, IGHV3-11*01, IGHV4-28*01, IGHV6-1*01, IGHV7-4-1*03. In another example, a V.sub.H segment can be selected from each gene family; for example, a subset can include IGHV1-18*01, IGHV1-2*01, IGHV1-3*01, IGHV1-45*01, IGHV1-46*01, IGHV1-58*01, IGHV1-69*01, IGHV1-8*01, IGHV1-c*01, IGHV1-f*01, IGHV2-26*01, IGHV2-5*01, IGHV2-70*01, IGHV3-11*01, IGHV3-13*01, IGHV3-15*01, IGHV3-16*01, IGHV3-20*01, IGHV3-21*01, IGHV3-23*01, IGHV3-30*01, IGHV3-33*01, IGHV3-38*01, IGHV3-43*01, IGHV3-48*01, IGHV3-49*01, IGHV3-53*01, IGHV3-64*01, IGHV3-66*01, IGHV3-7*01, IGHV3-72*01, IGHV3-73*01, IGHV3-74*01, IGHV3-9*01; IGHV4-28*01, IGHV4-30-2*01, IGHV4-31*01, IGHV4-34*01, IGHV4-39*01, IGHV4-59*01, IGHV4-61*01, IGHV5-51*01, IGHV6-1*01 and IGHV7-4-1*01. One of skill of the art is able to select any subset of germline sequences as desired based on sequence differences. Subsets for other germline segments sequences also can be selected based on sequence differences. Tables 10-12 sets forth exemplary selected V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and or J.sub..lamda. germline segments representing selection of at least one germline segment from each gene family.
TABLE-US-00010 TABLE 10 Selected Human Heavy Chain Germline Segments SEQ ID NO. V Segments IGHV1-18*01 10 IGHV1-2*02 13 IGHV1-24*01 16 IGHV1-3*02 18 IGHV1-45*02 20 IGHV1-46*01 22 IGHV1-58*02 26 IGHV1-69*06 32 IGHV1-8*01 40 IGHV2-26*01 44 IGHV2-5*01 45 IGHV2-70*13 67 IGHV3-11*01 68 IGHV3-13*01 70 IGHV3-15*01 73 IGHV3-16*02 82 IGHV3-20*01 83 IGHV3-21*01 84 IGHV3-23*01 86 IGHV3-30*03 93 IGHV3-35*01 117 IGHV3-38*02 119 IGHV3-43*01 120 IGHV3-48*02 123 IGHV3-49*03 127 IGHV3-53*01 130 V or D Segments IGHV3-64*02 134 IGHV3-66*03 140 IGHV3-7*01 142 IGHV3-72*01 144 IGHV3-73*02 147 IGHV3-74*01 148 IGHV3-9*01 151 IGHV4-28*01 153 IGHV4-31*02 169 IGHV4-34*01 178 IGHV4-39*01 191 IGHV4-4*07 204 IGHV4-59*01 205 IGHV5-51*03 227 IGHV6-1*01 233 IGHV7-81*01 238 D Segments -- IGHD1-1*01 239 IGHD1-14*01 240 IGHD1-20*01 241 IGHD1-26*01 242 IGHD1-7*01 243 IGHD2-15*01 244 IGHD2-2*01 245 IGHD2-21*01 248 IGHD2-8*01 250 D or J Segments IGHD3-10*01 252 IGHD3-16*01 254 IGHD3-22*01 256 IGHD3-3*01 257 IGHD3-9*01 259 IGHD4-11*01 260 IGHD4-17*01 261 IGHD4-23*01 262 IGHD4-4*01 263 IGHD5-12*01 264 IGHD5-18*01 265 IGHD5-24*01 266 IGHD5-5*01 267 IGHD6-13*01 268 IGHD6-19*01 269 IGHD6-25*01 270 IGHD6-6*01 271 IGHD7-27*01 272 J Segments -- IGHJ1*01 273 IGHJ2*01 274 IGHJ3*01 275 IGHJ4*01 277 IGHJ5*01 280 IGHJ6*01 282
TABLE-US-00011 TABLE 11 Human Light Chain Kappa Germline Segments Selected for Manual Compilation SEQ SEQ SEQ ID ID ID V Segments NO. V Segments NO. V or J Segments NO. IGKV2D-30*01 330 IGKV1-6*01 299 IGKV1-9*01 301 IGKV2-30*01 321 IGKV1-5*01 296 IGKV1-8*01 300 IGKV2D-29*01 328 IGKV1D-17*01 310 IGKV2D-40*01 331 IGKV1-27*01 292 IGKV3-15*01 334 IGKV1-39*01 295 IGKV2-24*01 317 IGKV1D-42*01 314 IGKV1-33*01 293 IGKV6D-21*01 354 IGKV1D-43*01 315 J Segments -- IGKV3-20*01 335 IGKV3D-7*01 350 IGKJ1*01 356 IGKV1-17*01 290 IGKV1-13*02 288 IGKJ2*01 357 IGKV5-2*01 352 IGKV1-12*01 286 IGKJ3*01 361 IGKV4-1*01 351 IGKV3-11*01 332
TABLE-US-00012 TABLE 12 Human Light Chain Lambda Germline Segments Selected for Manual Compilation SEQ SEQ SEQ ID ID ID V Segments NO. J Segments NO. J Segments NO. IGLV1-36*01 365 IGLV3-16*01 405 IGLV5-45*03 429 IGLV1-40*01 366 IGLV3-19*01 406 IGLV5-48*01 430 IGLV1-44*01 369 IGLV3-21*02 408 IGLV5-52*01 431 IGLV1-50*01 373 IGLV3-22*01 410 IGLV11-55*01 379 IGLV2-8*01 397 IGLV3-25*02 412 IGLV4-60*02 420 IGLV10-54*02 377 IGLV3-27*01 414 IGLV4-69*01 422 IGLV6-57*01 432 IGLV3-9*01 416 J Segments -- IGLV2-11*01 380 IGLV3-10*01 401 IGLJ1*01 442 IGLV2-14*01 383 IGLV3-12*02 404 IGLJ2*01 443 IGLV2-18*01 387 IGLV7-43*01 433 IGLJ4*01 446 IGLV2-23*03 393 IGLV7-46*02 435 IGLJ5*01 447 IGLV2-33*01 394 IGLV8-61*01 436 IGLJ6*01 449 IGLV3-1*01 400 IGLV5-37*01 424 IGLJ7*01 450
[0420] In all of the examples above, selection of germline segments based on sequence similarity or differences or other characteristics can be restricted for only one group of germline segments (from among any of V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda.), or 2 groups, 3 groups, 4 groups, 5 groups, 6 groups or all 7 groups. Thus, for example, in recombining the gene segments to encode for a plurality of VH chains, only the V.sub.H germline segment sequences can be restricted based on sequence similarity or differences or other characteristic, and the D.sub.H and J.sub.H segment sequences can represent all known D.sub.H and J.sub.H germline segment sequences. In another example, each of the V.sub.H, D.sub.H and J.sub.H segment sequences can be selected based on sequence similarity or differences or other characteristic, thereby resulting in a restricted subset of germline segment sequences for compilation herein. In yet another example, in recombining the gene segments to encode for a plurality of VH chains, the D.sub.H segment is restricted based on its modification to include nucleotides encoding a particular peptide mimetic or mimetics against a target. In such an example, the V.sub.H and J.sub.H segment sequences can represent all known germline segment sequences to be recombined with the restricted subset of D.sub.H (e.g. modified) segment. The choice of germline segments selected for use in the compilation method provided herein depends on a variety of factors including, but not limited to, the diversity of the resulting library, knowledge regarding preference for a particular germline segment sequence for a target from an initial screen of a library (as described herein below under Section G.4 entitled iterative screening), and the size of the library.
[0421] c. Sequence Compilation
[0422] In the methods provided herein, the variable gene segment sequences are recombined to generate heavy chain variable regions (5'-V.sub.HD.sub.HJ.sub.H-3'), kappa light chain variable regions (5'-V.sub..kappa.J.sub..kappa.-3'), and lambda light chain variable regions as described below. The gene segments can be recombined to generate full-length variable regions, or variable regions that are less then full length (i.e. portion thereof of full length), so long as the portion is sufficient to form an antigen binding site when expressed. The nucleic acid sequences are combined so that the resulting nucleic acid molecule is in-frame and encodes a functional VH or VL polypeptide, i.e. a full-length polypeptide or a portion thereof that is sufficient to form an antigen binding site.
[0423] The compilation method provided herein can be implemented by any procedures known to one of skill in the art. For example, it can be implemented manually, in silico (e.g. through computer software) or combinations thereof. In some examples, as described elsewhere herein below, the method can be implemented using sequence compilation software. In addition, public databases, such as those providing germline segment sequences or other public bioinformatics tools can be use to aid practice of the method.
[0424] Generally, full length heavy chain variable regions (5'-V.sub.HD.sub.HJ.sub.H-3'), or portions thereof, are recombined such that a V.sub.H segment is combined with a D.sub.H segment which is combined with a J.sub.H segment. Heavy chain segments are always combined such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment. The exact V.sub.HD.sub.HJ.sub.H can be chosen randomly or empirically and can represent germline segments, or modified forms thereof as discussed above. It is understood that when the method is performed manually using molecular biology techniques, restriction enzymes can be added to the ends of germline segments to facilitate joining of segments. Hence, in some examples, the resulting VH chain can contain additional amino acids between germline segments. For example, as described in the examples herein, nucleic acid molecule library members encoding a VH chain can encode sequences that contain amino acids SY at 5'end of the J.sub.H region between the joined D.sub.H region.
[0425] In one example of the methods herein, all permutations of V.sub.HD.sub.HJ.sub.H gene segments can be recombined to generate a nucleic acid molecule encoding a variable heavy chain. Thus, every V.sub.H segment (e.g., set forth in any of SEQ ID NOS:10-238), is combined with every D.sub.H segment (e.g., any set forth in SEQ ID NOS: 239-272), which is combined with every J.sub.H segment (e.g. any set forth in SEQ ID NOS: 273-285). In such an example, based on the exemplary heavy chain germline segments set forth in Table 3, greater then or about 100,000 nucleic acid molecules encoding a variable heavy (VH) chain can be generated. In other examples, the V.sub.HD.sub.HJ.sub.H gene segments can be recombined empirically (e.g. using rational or semi-rational approaches as discussed below). For example, as discussed below any subset of V.sub.H, D.sub.H and/or J.sub.H gene segment can be chosen to generate a recombined nucleic acid molecule. In some examples, individual gene segments are selected because of a shared characteristic including, but not limited to, diversity, same V region family, CDR3 length, composition or other biochemical attribute.
[0426] Full length kappa light chain variable regions (5'-V.sub..kappa.J.sub..kappa.-3'), or portions thereof, are recombined such that a V.sub..kappa. segment is combined with a J.sub..kappa. segment. Full length lambda light chain variable regions are recombined such that a V.sub..lamda. segment is combined with a J.sub..lamda. segment. Light chain segments are always combined such that the V.sub.L segment is 5' to the J.sub.L segment. The exact V.sub..kappa.J.sub..kappa. or V.sub..lamda.J.sub..lamda. can be chosen randomly or empirically and can represent germline segments, or modified forms thereof as discussed above. It is understood that when the method is performed manually using molecular biology techniques, restriction enzymes can be added to the ends of germline segments to facilitate joining of segments. Hence, in some examples, the resulting VL chain can contain additional amino acids between germline segments.
[0427] In one example of the methods herein, all permutations of V.sub..kappa.J.sub..kappa. can be recombined to generate a nucleic acid molecule encoding a variable kappa light chain. Thus, every V.sub..kappa. (e.g. any set forth in SEQ ID NOS:286-355, 868) is combined with every J.sub..kappa. (e.g. any set forth in SEQ ID NOS: 356-364). In such an example, based on the exemplary kappa light chain germ line segments set forth in Table 4, greater then or about 600 nucleic acid molecules encoding a variable kappa light chain can be generated. In another example, all permutations of V.sub..lamda.J.sub..lamda. can be recombined to generate a nucleic acid molecule encoding a variable lambda light chain. Thus, every V.sub..lamda. (e.g. any set forth in any of SEQ ID NOS:365-441) is combined with every J.sub..lamda. (e.g. any set forth in any of SEQ ID NOS: 442-451). In such an example, based on the exemplary lambda light chain germline segment set forth in Table 5, greater then or about 700 nucleic acid molecules encoding a variable lambda light chain can be generated. In another example, the V.sub..kappa.J.sub..kappa. or V.sub..lamda.J.sub..lamda. gene segments can be recombined empirically as described herein below.
[0428] In all of the examples above, recombined segments are joined such that the recombined full length nucleic acid is in frame with the 5' start codon (ATG), thereby allowing expression of a full length polypeptide. Any combination of a V(D)J can be made, and junctions modified accordingly in order to generate a compiled V(D)J sequence that is in-frame, while preserving reading frames of each segment. The choice of junction modification is a function of the combination of V(D)J that will be joined, and the proper reading frame of each gene segment. For example, any of the variable gene segments can exist in reading frame 1, 2 or 3 when compiled. Generally, however, for the practice of the method herein, the V sequence (V.sub.H, V.sub..kappa. or V.sub..lamda.) is always reading frame 1. Also, the reading frame of the J sequence is set to be either reading frame 1, 2 or 3 such that the resulting gene segment encodes the correct amino acids. Table 13 below sets forth the reading frames of the exemplary J germline sequences.
TABLE-US-00013 TABLE 13 J Germline Segment Sequences in Coding Frames SEQ ID SEQUENCE NO Heavy IGHJ1*01 GCT GAA TAC TTC CAG CAC TGG GGC CAG GGC ACC CTG 273 GTC ACC GTC TCC TCA G IGHJ2*01 C TAC TGG TAC TTC GAT CTC TGG GGC CGT GGC ACC CTG 274 GTC ACT GTC TCC TCA G IGHJ3*01 T GAT GCT TTT GAT GTC TGG GGC CAA GGG ACA ATG GTC 275 ACC GTC TCT TCA G IGHJ3*02 T GAT GCT TTT GAT ATC TGG GGC CAA GGG ACA ATG GTC 276 ACC GTC TCT TCA G IGHJ4*01 AC TAC TTT GAC TAC TGG GGC CAA GGA ACC CTG GTC 277 ACC GTC TCC TCA G IGHJ4*02 AC TAC TTT GAC TAC TGG GGC CAG GGA ACC CTG GTC 278 ACC GTC TCC TCA G IGHJ4*03 GC TAC TTT GAC TAC TGG GGC CAA GGG ACC CTG GTC 279 ACC GTC TCC TCA G IGHJ5*01 AC AAC TGG TTC GAC TCC TGG GGC CAA GGA ACC CTG 280 GTC ACC GTC TCC TCA G IGHJ5*02 AC AAC TGG TTC GAC CCC TGG GGC CAG GGA ACC CTG 281 GTC ACC GTC TCC TCA G IGHJ6*01 AT TAC TAC TAC TAC TAC GGT ATG GAC GTC TGG GGG 282 CAA GGG ACC ACG GTC ACC GTC TCC TCA G IGHJ6*02 AT TAC TAC TAC TAC TAC GGT ATG GAC GTC TGG GGC 283 CAA GGG ACC ACG GTC ACC GTC TCC TCA IGHJ6*03 AT TAC TAC TAC TAC TAC TAC ATG GAC GTC TGG GGC 284 AAA GGG ACC ACG GTC ACC GTC TCC TCA IGHJ6*04 AT TAC TAC TAC TAC TAC GGT ATG GAC GTC TGG GGC 285 AAA GGG ACC ACG GTC ACC GTC TCC TCA G Light Kappa IGKJ1*01 G TGG ACG TTC GGC CAA GGG ACC AAG GTG GAA ATC AAA C 356 IGKJ2*01 TG TAC ACT TTT GGC CAG GGG ACC AAG CTG GAG ATC AAA C 357 IGKJ2*02 G TGC ACT TTT GGC CAG GGG ACC AAG CTG GAG ATC AAA C 358 IGKJ2*03 TG TAC AGT TTT GGC CAG GGG ACC AAG CTG GAG ATC AAA C 359 IGKJ2*04 TG TGC AGT TTT GGC CAG GGG ACC AAG CTG GAG ATC AAA C 360 IGKJ3*01 A TTC ACT TTC GGC CCT GGG ACC AAA GTG GAT ATC AAA C 361 IGKJ4*01 G CTC ACT TTC GGC GGA GGG ACC AAG GTG GAG ATC AAA C 362 IGKJ4*02 G CTC ACG TTC GGC GGA GGG ACC AAG GTG GAG ATC AAA C 363 IGKJ5*01 G ATC ACC TTC GGC CAA GGG ACA CGA CTG GAG ATT AAA C 364 Light Lambda IGLJ1*01 T TAT GTC TTC GGA ACT GGG ACC AAG GTC ACC GTC CTA G 442 IGLJ2*01 T GTG GTA TTC GGC GGA GGG ACC AAG CTG ACC GTC CTA G 443 IGLJ3*01 T GTG GTA TTC GGC GGA GGG ACC AAG CTG ACC GTC CTA G 444 IGLJ3*02 T TGG GTG TTC GGC GGA GGG ACC AAG CTG ACC GTC CTA G 445 IGLJ4*01 T TTT GTA TTT GGT GGA GGA ACC CAG CTG ATC ATT TTA G 446 IGLJ5*01 C TGG GTG TTT GGT GAG GGG ACC GAG CTG ACC GTC CTA G 447 IGLJ5*02 C TGG GTG TTT GGT GAG GGG ACG GAG CTG ACC GTC CTA G 448 IGLJ6*01 T AAT GTG TTC GGC AGT GGC ACC AAG GTG ACC GTC CTC G 449 IGLJ7*01 T GCT GTG TTC GGA GGA GGC ACC CAG CTG ACC GTC CTC G 450 IGLJ7*02 T GCT GTG TTC GGA GGA GGC ACC CAG CTG ACC GCC CTC G 451
[0429] For the heavy chain, the reading frame of the D variable gene segment sequence chosen is less rigid then for the V or J germline segments. This is because the D gene sequence is responsible for encoding the central portion of the CDRH3, which plays a prominent role in antigen specificity. Hence, variation of amino acids is expected in the D gene segment sequence. Thus, for example, the D.sub.H gene segment can be any D.sub.H gene segment in any reading frame, an inverted or reverse complement thereof, or a modified form thereof, or any sequence of nucleotides designated as the D.sub.H. In some examples, however, the reading frame of the D germline sequence is chosen so that the resulting encoded amino acids are predominately hydrophilic. CDR3 is an antigen-binding site, and thereby is rich in hydrophilic residues that are surface exposed (see e.g., Zanetti and Billetta, Antigenized Antibodies from Concepts to Applications (1996), In The Antibodies, Volume 2 (pp. 75-122), Harwood Academic Publishers; Pommie et al. (2004) J Mol. Recognition, 17:17-32). One of skill in the art is familiar with techniques to assess the hydrophobicity/hydrophilicity of sequences. For example, hydrophilicity can be measured using protein grand average of hydropathy (GRAVY), which gives hydropathy value for a sequence by adding the hydropathy value for each residue and dividing by the length of the sequence (see e.g., Kyte and Doolittle (1982 and bioinformatics.org/sms2/protein_gravy.html). The lower the GRAVY value, the more hydrophilic a sequence is.
[0430] In some instances, compilation of variable gene segments in-frame, while preserving reading frames, requires no manipulation, i.e. no modification of joint regions. In other instances, however, simply compiling V(D)J sequences does not conserve reading frames. Thus, where the junctions between gene segments are not in the desired frame, modifications are made to nucleotides within the junctions between the segments so that each gene segment is in its desired reading frame, and the full length sequence is in-frame. Nucleic acid modifications include replacements or substitutions, insertions, or deletions of nucleotides, or any combination thereof. For example, at the V-D junction, one or more nucleotide can be deleted from the 5' end of the D, one or more nucleotide can be deleted from the 3' end of the V or one or more nucleotides can be inserted between the V and D (e.g. a nucleotide can be added to the 3' end of the V). In another example, at the D-J junction, one or more nucleotides can be deleted from the 5' end of the J, one or more nucleotides can be deleted from the 3' end of the D, or one or more nucleotides can be inserted between the D and J (e.g., a nucleotide can be added to the 3' end of the D). In a further example, at the V-J junction, as occurs in generation of a light chain, one or more nucleotides can be deleted from the 5' end of the J, one or more nucleotides can be deleted from the 3' end of the V or one or more nucleotides can be inserted between the V and J (e.g. a nucleotide can be added to the 3' end of the V). In such examples where nucleotides are inserted, any nucleotide insertion from among one or more of a guanine (G), adenine (A), cytosine (C) and thymine (T) is contemplated. In some examples, guanine (G) is chosen as the inserted nucleotide because of the slight preference of terminal deoxynucleotidyl transferase (TdT) for guanine residues (Alt et al. 1982).
[0431] In the methods, heavy chain segments are recombined separately from light chain gene segment sequences. Thus, an individual nucleic acid molecule encodes for either a heavy chain (VH) or a light chain (VL) variable region. In the methods, a plurality of VH nucleic acid molecules encoding a VH chain and a plurality of nucleic acid molecules encoding a VL chain are generated. The number of such sequences can be up to all possible permutations depending on the number of V, D or J gene segments available for combination. For example, where all known germline segment sequences are used for practice of the method, a fully naive antibody library is generated. In other examples, modified gene segments can also be used for practice of the method. Alternatively, the number of permutations is a function of the selected V, D and J, which can be a subset of all germline segments or modified forms thereof.
[0432] Once a nucleic acid sequence is compiled, it is further modified to remove stop codons so that the resulting molecule is functional, i.e. encodes a polypeptide that is not truncated early. For example, modifications to remove stop codons include substitutions of nucleotides. Exemplary of such modifications, include, but are not limited to, stop codon TAA replaced by codons TAT; stop codon TAG replaced by codons TAT, and stop codon TGA replaced by codons TCA.
[0433] d. Further Sequence Modification of Recombined Nucleic Acid Sequences
[0434] As discussed above, germline segment sequences can be modified before performing compilation as described herein. In addition or alternatively, modification can be made directly to the recombined nucleic acid sequence. Hence, it is understood that any of the modifications described below also can be made to individual germline segment sequences before compilation so long as the reading frames are maintained and the rules governing compilation as described herein are observed to generate in-frame recombined nucleic acid sequences.
[0435] Thus, any of the plurality of recombined nucleic acids encoding a VH chain or a VL chain can be further modified. Modifications of the nucleic acid sequences include replacements or substitutions, insertions, or deletions of nucleotides, or any combination thereof. Any modification contemplated by one of skill in the art can be made to the nucleic acid molecule, so long as the modification(s) do not interfere with or alter the junction joints made to maintain reading frames of the V(D)J segments achieved by virtue of practice of the method (as discussed in the Section entitled "Sequence Compilation" above). Any modification should be checked to confirm that all reading frames are intact to ensure that the resulting full length nucleic acid is in frame with the 5' start codon (ATG) thereby allowing expression of a full length VH or VL polypeptide, or a portion thereof that is sufficient to form an antigen binding site.
[0436] The resulting recombined germline variable heavy and light chain nucleic acid sequences can be further modified through DNA synthesis (i.e. modifications introduced upon synthesis of the nucleic acid molecule) or by using standard molecular biology techniques. Thus, in one example, any desired modification contemplated can be made to a nucleic acid molecule encoding a recombined variable heavy or variable light chain and the resulting nucleic acid molecule including any modifications synthesized as described in sub-section e.iii below. Due to the degeneracy of the genetic code nucleic acid sequences can be designed to avoid unwanted nucleotide sequences, including unwanted restriction sites, splicing donor or acceptor sites, or other nucleotide sequences potentially detrimental to efficient translation. Additionally, organisms sometimes favor particular codon usage and/or a defined ratio of GC to AT nucleotides. Thus, degeneracy of the genetic code permits design of nucleic acid sequences tailored for expression in particular organisms or groups of organisms. Additionally, nucleic acid molecules can be designed for different levels of expression based on optimizing (or non-optimizing) of the sequences. In another example, generated recombined germline VH and VL nucleic acid molecules as described in sub-section e.iii below, can be further modified using standard molecular biology techniques, such as PCR, site-directed mutagenesis, restriction enzyme digestion, ligation, cloning and any combination thereof. The choice of whether to generate such modifications during DNA synthesis or using molecular biology techniques is dependent on the end user and can be influenced by factors such as the purpose of the modification, the extent of the modification and timing considerations.
[0437] Modifications of recombined germline nucleic acid molecules encoding VH or VL can be generated randomly or empirically. For example, random mutation of one or more regions can increase diversity of the library, particularly where modifications are made to any of the CDR-loop regions, which contribute to the specificity and affinity of the antibody. This library with increased diversity permits the generation of antibodies, derivatives thereof or portions or fragments thereof, which potentially can bind to any desired antigen with a high affinity. In another example, modifications can be empirically generated using rational or semi-rational approaches. Among such empirical modifications of nucleic acid molecules encoding VH and VL chains contemplated herein include, but are not limited to, modifications of the CDR regions, for example for the generation of directed libraries, modifications to optimize codon usage, and/or modifications to introduce restriction sites or detectable moieties. Modifications also can include a combinations of random and empirical modifications.
[0438] i. Codon Usage
[0439] For example, nucleic acid sequences can be modified to adapt the codon usage for expression such as, for example, bacterial expression. Codon usage is degenerate in that multiple codons encode for the same amino acid. Thus a single amino acid is therefore encoded by multiple codons, however within any organism, codon usage varies for any given amino acid. The full length nucleic acids provided herein are modified to replace rare codons with more abundant codons utilized in the particular expression system. Typically, modifications include silent mutations, such that the substitutions do not alter the specificity of the codon. Codon usage tables are known to those of skill in the art, particularly for common expression systems. For example, for expression in bacteria E. coli K12, codon usage Tables are known (see, e.g., Grantham, R. et al., Nuc. Acids Res., 8:1892-1912 (1980); Grantham, R. et al, Nuc. Acids Res., 9:r43-r74 (1981) and also Table 14). The codon usage table lists all of the sixty four possible three nucleotide codons for DNA or RNA with their frequency of usage in the bacteria E. coli K12. The Table shows that while a single amino acid is encoded by multiple codons (redundancy), these codons are not used at the same rate for any given amino acid. For example, the amino acid arginine is coded for by six different codons: CGT, CGC, CGA, CGG, AGA and AGG. The codon AGA has a frequency of 2.0% while the codon CGC has a frequency of 22%.
TABLE-US-00014 TABLE 14 Codon Usage in E. coli K12 fields: [triplet] [frequency: per thousand] ([number]) TTT 22.2 (30361) TCT 8.4 (11498) TAT 16.1 (22071) TGT 5.1 (7020) TTC 16.6 (22649) TCC 8.6 (11804) TAC 12.2 (16734) TGC 6.4 (8787) TTA 13.8 (18915) TCA 7.1 (9706) TAA 2.0 (2752) TGA 0.9 (1261) TTG 13.6 (18601) TCG 8.9 (12156) TAG 0.2 (321) TGG 15.2 (20835) CTT 11.0 (15043) CCT 7.0 (9554) CAT 12.9 (17656) CGT 21.0 (28700) CTC 11.1 (15183) CCC 5.4 (7448) CAC 9.7 (13329) CGC 22.0 (30159) CTA 3.9 (5303) CCA 8.4 (11518) CAA 15.3 (20970) CGA 3.5 (4787) CTG 52.9 (72403) CCG 23.3 (31869) CAG 28.9 (39560) CGG 5.4 (7320) ATT 30.4 (41551) ACT 8.9 (12197) AAT 17.7 (24192) AGT 8.7 (11917) ATC 25.2 (34426) ACC 23.5 (32101) AAC 21.7 (29656) AGC 16.1 (21961) ATA 4.3 (5827) ACA 7.0 (9564) AAA 33.7 (46044) AGA 2.0 (2783) ATG 27.8 (38012) AGC 14.4 (19743) AAG 10.3 (14043) AGG 1.1 (1533) GTT 18.3 (25079) GCT 15.3 (20863) GAT 32.2 (44103) GGT 24.9 (34009) GTC 15.3 (20913) GCC 25.6 (35018) GAC 19.2 (26201) GGC 29.8 (40725) GTA 10.9 (14885) GCA 20.2 (27638) GAA 39.7 (54267) GGA 7.9 (10817) GTG 26.3 (35960) GCG 33.8 (46222) GAG 17.8 (24414) GGG 11.0 (15116)
[0440] ii. Adding or Removing Restriction Enzyme Sites
[0441] In another example, additional modifications of the nucleic acids include the addition of flanking sequences at one or both of the 5' and 3' termini of recombined VH or VL nucleic acid sequences that provide restriction sites. Such modifications can be incorporated into the germline recombined nucleic acid molecules during DNA synthesis, or by PCR, for example using primers that incorporate the restriction enzyme sites. In some examples, addition of such restriction sites facilitate cloning of the nucleic acids into a selected vector. For example, restriction sites include any restriction site known in the art. Exemplary restriction site sequences are set forth in Table 15. Generally, the restriction site chosen is compatible with the expression vector and can be chosen to facilitate blunt-ended ligation or sticky-ended ligation. The choice of restriction enzyme is routine and is well within the level of one of skill in the art.
TABLE-US-00015 TABLE 15 Common Restriction Enzyme Cleavage Recognition Sites Restriction Enzyme Cleavage Sequence SEQ ID NO NcoI CCATGG 977 NheI GCTAGC 978 AvrII CCTAGG 979 BsiWI CGTACG 980 SfiI GGCCNNNNNGGCC 1889 NotI GCGGCCGC 1890 HirdIII AAGCTT 1891 EcoRI GAATTC 1892 BamHI GGATCC 1893 EcoRV GATATC 1894 PstI CTGCAG 1895 SalI GTCGAC 1896 SmaI CCCGGG 1897 XmaI CCCGGG 1898 BglI GCCNNNNNGGC 1899 MfeI CAATTG 1900 BsaI GGTCTCN 3719
[0442] In some examples, nucleic acids can be modified to remove any restriction sites that occur within the nucleic acid sequence. In particular, removal of restriction sites is desired so that such sites do not interfere with subsequent digestion, ligation and cloning procedures. For example, as discussed above, recombined nucleic acid molecules can be modified to contain terminal flanking restriction sites to facilitate cloning into expression vectors. Generally, such restriction sites are chosen to be unique so that the presence of the site exists only at the terminal flanking end(s). If the site is not unique, modifications can be made to the sequence of the nucleic acid molecule to remove any conflicting restriction sites. One of skill in the art is familiar with restriction sites and can identify such sites within a nucleic acid sequence. Table 15 lists exemplary restriction sites that can be removed.
[0443] In some instances, a single nucleotide change is possible to effect change of the restriction site. In other instances, two or three nucleotide changes are necessary to remove a restriction site. Typically, modification of restriction sites existing internally in a recombined nucleic acid molecule are made in view of the codon usage as discussed above. For example, if a Sal I restriction sites (GTCGAC; SEQ ID NO:1896) exists internally in a nucleic acid molecule, the GTC codon that codes for valine (V) can be modified to GTA, GTG, GTT or GTC codons. Simply changing the last C to G correlates to changing the GTC codon (15.3% frequency of usage) to GTG (26.3% frequency of usage), which is an 11% increase in frequency of codon usage. Alternatively, the GAC codon (19.2% frequency of usage) that codes for asparagine (D) can be modified to GAT (32.2% frequency of usage) by changing the last C to T, which is a 13% increase in codon usage. In this example, either of the above modifications can be made. Typically, modifications are made to convey the highest absolute beneficial increase in frequency of codon usage.
[0444] iii. Linkers
[0445] In additional examples, nucleic acid molecules can be modified with a linker sequence. For example, where a single-chain antibody is desired (e.g. an scFv antibody) the variable heavy and light chains can first be joined by a linker. The linkage can be direct or via a linker. For example, nucleic acids encoding peptide linkers can be added during DNA synthesis or using molecular biology techniques to the 5' end of a first sequence (e.g. variable heavy chain) and the 3' terminus of a second nucleic acid sequence (e.g. variable light chain). Typically, the linker is of sufficient length so that the resulting polypeptide is soluble. Nucleic acid sequences for use as linkers can encode peptide linkers from about 2 or 2 to about 60 or 60 amino acid residues, for example from about 5 to 40, or from about 10 to 30, 2 to 6, 7, or 8 amino acid residues. Examples of known linker moieties include, but are not limited to, peptides, such as (Gly.sub.mSer)n and (Ser.sub.mGly)n, in which n is 1 to 6, including 1 to 4 and 2 to 4, and m is 1 to 6, including 1 to 4, and 2 to 4. Exemplary of such linkers include any that encode peptide linkers such as glycine serine polypeptides, such as -Gly-Gly-, GGGGG (SEQ ID NO:981), GGGGS (SEQ ID NO:982) or (GGGGS)n (SEQ ID NO:985), SSSSG (SEQ ID NO:983) or (SSSSG)n (SEQ ID NO:1996). Linking moieties are described, for example, in Huston et al, (1988) PNAS 85:5879-5883, Whitlow et al, (1993) Protein Engineering 6:989-995, and Newton et al., (1996) Biochemistry 35:545-553. Other suitable linkers include any encoding a peptide linker, such as any of those described in U.S. Pat. No. 4,751,180 or 4,935,233, which are hereby incorporated by reference. A polynucleotide encoding a desired peptide linker can be inserted anywhere in variable heavy or light chain sequence or at the 5'- or 3'-terminus, in frame, using any suitable conventional technique. For example, restriction sites can be added to the 5' terminus of the heavy chain sequence and to the 3' terminus of the light chain sequence while a nucleic acid encoding a linker segment (e.g. (Gly.sub.4Ser).sub.3; SEQ ID NO:984) can be added to the 3' terminus of the heavy chain sequence connecting it to the 5' terminus of the light chain sequence. Upon expression, such a nucleic acid molecule encodes an scFv antibody where the heavy chain variable region is operably linked to the light chain variable region.
[0446] iv. Tags or Detectable Moieties
[0447] Additionally, a small epitope tag, such as a myc tag, His tag, Flag tag or other small epitope tag, and/or any other additional DNA sequence can be added for incorporation into a nucleic acid sequence encoding a variable heavy chain or variable light chain (Arnau et al, (2006) Protein Expression and Purification, 48:1-13). In some instances, for example, a tag that permit attachment, for example, an LPETG tag, can be added that allows for site specific modification using the protein ligase, sortase (Chan et al, (2007) PLoS ONE, 2:e1164). Hence, inclusion of such a tag permits immobilization (e.g. on a BIAcore chip) and/or selective sorting in the presence of a sortase. Generally, the additional DNA sequence is added to the 3' or 5' terminus of the nucleic acid molecule encoding the recombined variable sequence directly or indirectly using a linker. Alternatively, the additional DNA sequence can be included in the expression vector of choice, such that, upon expression, the resulting antibody contains the additional sequence. For example, plasmid A set forth in SEQ ID NO:1 contains a His-Flag Tag corresponding to nucleotides 3265-3306 (Flag corresponds to nucleotides 3265-3288; His corresponds to nucleotides 3289-3306). In another example, Plasmid D set forth in SEQ ID NO: 2 contains a Flag tag corresponding to nucleotides 3265-3288, an LPETG tag corresponding to nucleotides 3289-3303. Thus, upon expression of the heavy chains, alone or together with a variable light chain, resulting antibodies can be detected using anti-Flag or anti-His tag reagents. This is described in Example 10. One of skill in the art can add any desired detectable sequence or other identifiable moiety to a nucleic acid molecule encoding a recombined variable heavy or light chain sequence to facilitate identification and/or purification of the resulting antibodies.
[0448] v. Mutational Diversity
[0449] In other examples, modifications can be made to introduce mutational diversity into the resulting nucleic acid molecules. Any modification can be made, such as by replacement, substitution, deletion or addition of amino acids, either randomly or empirically (i.e. into any region or segment of the recombined nucleic acid molecule). The modifications can be made during DNA synthesis or using routine molecular biology techniques such as site-directed mutagenesis, digestion with restriction enzymes and/or cloning.
[0450] For example, modification(s) can be introduced into a nucleic acid molecule encoding the VH chain, a nucleic acid molecule encoding the VL chain, or both. The modification(s) can be introduced in the region of one or more of CDR1, CDR2, CDR3, FR1, FR2, FR3 or FR4. For example, modifications can be introduced into one, two or all three of the three CDRs of a given variable domain (VH, VL or both). In one example, modifications are introduced into CDR1 and CDR2, e.g. of a heavy chain variable domain. Typically, modification(s) are introduced into the CDR3 of the heavy chain (CDRH3). Any combination is contemplated. One of skill in the art knows and can identify CDR1, CDR2, CDR3, FR1, FR2, FR3 and FR4 regions in a nucleic acid molecule encoding a VH or VL (see e.g., Chothia et al. (1989) Nature 342:877-883; Al-Lazikani et al. (1997) J Mol. Biol., 273:927-948); WO/2007/137616; bioinf.org.uk/abs/; bioc.unizh.ch/antibody/Numbering/NumFrame.html; Martin et al., Bioinformatics Tools for Antibody Engineering in Handbook of Therapeutic Antibodies, Wiley-VCH (2007), pp. 96-103). For example, CDRs can be identified in VH and VL chains using Kabat numbering based on sequence alignment or the Chothia numbering scheme based on structural topology. Since the Kabat numbering scheme was developed from sequence alignment, insertions in the sequence relative to the numbering scheme by alignment are indicated by letters (e.g. 27, 27A, 27B, 27C, etc. . . . ) and deletions have the corresponding number skipped. The residues corresponding to the six CDRs of the light and heavy chains based on Kabat numbering are CDR-L1: L24-L34; CDR-L2: L50-L56; CDR-L3: L89-L97; CDR-H1: H31-H35B; CDR-H2: H50-H65; CDR-H3: H95-H102. One of skill in the art knows that CDR lengths can vary and can identify corresponding residues, for example, by alignment and use of kabat numbering.
[0451] vi. Directed Peptides
[0452] In some cases, modifications include rationally generated modifications to generate antibodies and portions or fragments thereof that mimic the activity of biologically active peptides against known targets (see e.g., International published PCT Application No. WO 2004/050017). Important biological functions, such as receptor binding, activation and enzymatic activity, are often attributable to discrete regions of larger protein molecules, containing a limited number of amino acid residues termed peptide epitopes and mimitopes. These peptide epitopes and mimitopes can be used as therapeutics, but due to their small size, are typically unstable in vivo due to rapid degradation. The peptide epitopes, however, can be introduced into variable regions of antibodies, which can act to mimic the activity of the biologically active peptide. Such antibodies are more stable, and exhibit increased half-life. Thus, antibodies or portions thereof can be directed toward a known target or function by incorporating sequences into the variable regions of an antibody that correspond to a polynucleotide target of interest. Often, structure and or function information of the known targets is available. These libraries are useful in providing lead antibodies for future antibody libraries.
[0453] Hence, included in the modifications herein are nucleic acid sequences encoding germline recombined VH and VL, wherein nucleotides corresponding to one or more CDR is replaced with nucleotides encoding one or more amino acid residues for a peptide of choice. In one example, the modifications in a nucleic acid molecule encoding a germline recombined VH and/or VL can be generated during DNA synthesis. Alternatively, the modification can be introduced into a nucleic acid molecule encoding a germline recombined VH and/or VL by restriction digestion followed by ligation with a peptide of choice. If necessary, restriction sites can be created in a CDR, such as by site-directed mutagenesis or PCR, to facilitate ligation of the peptide. This latter method is described in Example 12 herein.
[0454] The nucleotides can encode for peptides of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 16, 20, 25 or more amino acids. Any peptide that exhibits a useful property is suitable for insertion into an antibody scaffold. Generally, the peptide is one that specifically binds a target molecule. The peptides also include those that exhibit a specific activity, for example, an agonist or antagonist activity upon binding to the target. Peptide activities and uses include, but are not limited to, binding a receptor, binding a membrane bound surface molecule, binding a ligand, binding an enzyme or structural protein, activating or inhibiting a receptor, target drug delivery or any enzymatic activity. Exemplary of peptides are those that bind to a cell surface receptor such as a receptor for a cytokine, growth factor or growth inhibitor. Peptide mimetics for incorporation into a recombined germline VH or VL include any set forth in U.S. Pat. Nos. 7,169,905; 7,396,917, 7,272,508, 7,019,017; U.S. published Patent Appl. No. US200701344; published International Appl. No. WO2005060642; Johnson et al. (2000) Nephrol Dial. Transplant, 15:1274-1277. Exemplary of such peptides are set forth in Table 16. Other peptides for incorporation into the VH and VL encoded by the recombined germline nucleic acids provided herein are known in the art (see e.g., any of the references cited above) and/or can be identified depending on the desired target.
TABLE-US-00016 TABLE 16 Exemplary Peptide Mimetics SEQ ID Peptide Sequence NO: TPO IEGPTLRQWLAARA 987 GGCADGPTLREWISFCGGK 988 GGCADGPTLREWISFCGG 989 LAIEGPTLRQWLHGNGRDT 990 GNADGPTLRQWLEGRRPKN 991 TIKGPTLRQWLKSREHTS 992 EPO TYSCHFGPLTWVCKPQ 891 DYHCRMGPLTWVCKPLGG 993 GGTYSCHFGPLTWVCKPQGG 994 DREGCRRGWVGQCKAWFN 995 QRVEILEGRTECVLSNLRGRTRY 996 G-CSF EEDCK 997 IL-5 TGGGDGYVCVEWARCPTCK 998 EGYVCVEWAACPTCR 999 human brain CFGRKMDRISSSSGLGC 1000 natriuretic FGRKMDRISSSSGLG 1001 peptide (hBNP-32) Exendin 4 HGEGRFTSDLSKQMEEEAVRLFIE 1002 WLKNGGPSSGAPPPS GLP-1 HAEGTFTSDVSSYLEGQMKEFIAW 1003 LVKGR GLP-2 HADGSFSDEMNTILDNLAARDFIN 1004 WLIQTKITDR Glucagon HSQGTFTSDYSKYLDSRRAQDRVQ 1005 WLMNT PACAP-38 HSDGIFTDSYSRYRKQMAVKKYLA 1006 AVLGKRYKQRVKNK CD209L RYWNSGEPNNSGNEDCAEFSGSGW 1007 NCNRCDVDN TNF YCFTASENHCY 1008 YCFTNSENHCY 1009 VEGF VEPNCDIHVMWEWECFERL 1010 GERWCFDGPLTWVCGEES 1011 MMP inhibitor CTTHWGFTLC 1012 CTLA-4 CSLHWGFWWC 1013 GFVCSGIFAVGVGRC 1014
[0455] Nucleic acid molecules encoding for recombined germline VH or VL can be modified by replacement or introduction of nucleotides encoding a peptide into one or more of CDR1, CDR2, CDR3, FR1, FR2, FR3 or FR4. For example, nucleic acid molecules encoding for recombined germline VH or VL can be modified by replacement of an entire CDR with nucleotides encoding a peptide. The CDR replaced by a peptide can be CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and/or CDRL3. For example, in the resulting VH or VL chain one or more CDRs is replaced by a peptide. The peptides can be the same or different. In another example, nucleic acid molecules encoding for recombined human germline VH and VL are modified by replacement of a portion of a CDR with nucleotides encoding a peptide. The portion of the CDR replaced by the nucleotides is a portion of CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and/or CDRL3. The portion of the CDR replaced by the nucleotides can encode for a peptide that is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 16, 20 or 25 or more amino acids. In an additional example, one or more portions of two or more CDRs are replaced by nucleotides encoding a peptide. The resulting peptides can be the same or different. In a further example, nucleic acid molecules encoding for recombined human germline VH or VL can be modified by insertion of nucleotides encoding a peptide between two nucleotides of a CDR of the antibody. The CDR with a peptide inserted is CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and/or CDRL3. In some instances, the resulting VH or VL chain includes one or more peptides inserted in one or more CDRs. The peptides inserted into the CDRs can be the same or different.
[0456] The addition of flanking sequences at the carboxy or N-terminal ends of a peptide have been shown to increase biological activity, by altering the presentation of the peptide within the antibody scaffold. Hence, nucleic acid molecules can be modified to encode peptides having adding flanking sequences at the carboxy or N-terminal ends of the peptides. Flanking sequences can encode for 1, 2, 3, 4, 5 or more amino acids. Flanking sequences can encode for any amino acid or any combinations of amino acids. Glycine is the smallest and simplest of the amino acids, containing only a single hydrogen atom in its side chain. Due to its small size, glycine can fit into small spaces and can adopt particular conformations that other amino acids can not. Proline is a sterically constrained amino acid that has been shown to increase activity of a peptide when flanking the peptide sequence (REF). Generally, flanking sequences encode for glycine or proline. Typically, flanking sequences encode for proline. For example, a nucleic acid molecule can encode a peptide containing proline and/or glycine added to the N- or C-terminus of the EPO peptide set forth in SEQ ID NO:891. Exemplary of nucleic acid molecules containing flanking sequences encode any of the EPO peptides set forth in SEQ ID NOS: 874-895.
[0457] e. Generating Variable Heavy and Light Chain Sequences and Nucleic Acid Molecules
[0458] The sequences for recombined nucleic acid molecules encoding VH and VL chain compiled by practice of the method herein are collected and stored. The collected sequences can be analyzed for any particular characteristic, such as for example, sequence similarity between and among other recombined sequences. The sequences then can be ranked based on sequence diversity. All recombined sequences, or a subset thereof, can be generated into recombined nucleic acid molecules using DNA synthesis and/or recombinant DNA technology. For example, a subset of sequences can be selected based on their sequence similarity or difference for generation of an antibody library.
[0459] i. Storage and Collection
[0460] Sequences recombined by the method herein are collected and stored. Typically, collection and storage is in an addressable format, such that the identity of each sequence is known by its locus. For example, the sequences can be stored in a database or in a list. Further, the individual gene segment components of each nucleic acid sequence are known, and the recombined nucleic acid sequence identified by the component segments. For example, a nucleic acid sequence named VH1-18_IGHD1-26*01_IGHJ2*01 identifies a nucleic acid sequence encoding a variable heavy chain containing the VH germline segment VH1-18 (also called VH1-18*01 by some nomenclature standards), the DH germline segment IGHD1-26*01, and the JH germline segment IGHJ2*01. One of skill in the art can identify a nucleic acid sequence using any desired naming convention, so long as the component segments are easily identified.
[0461] Generally sequences encoding VH chains are recombined, collected and stored separately from VL chains.
[0462] Further, among VL chains, sequences encoding V-kappa light chains are recombined, collected and stored separate from sequences encoding V-lambda chains. The identity of the nucleic acid sequence at each locus is known and can be mapped to an output file that contains the sequences for all the nucleic acid molecules within the addressable format.
[0463] For purposes herein, the sequences are addressably stored such that each sequence can be easily identified, including by its component parts (e.g. the individual compiled segments). By practice of the methods above, a plurality of different recombined nucleic acid sequences encoding a VH chain can be generated, which can represent all possible permutations of recombined segments or subsets thereof. For example, 10, 100, 500, 1000 (10.sup.3), 2.times.10.sup.3, 4.times.10.sup.3, 6.times.10.sup.3, 8.times.10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 2.times.10.sup.5, 3.times.10.sup.5, 4.times.10.sup.5, 5.times.10.sup.5, 6.times.10.sup.5, 7.times.10.sup.5, 8.times.10.sup.5, 9.times.10.sup.5, 10.sup.6, 10.sup.7 or more VH nucleic acid sequences can be generated. By practice of the methods above, a plurality of different recombined nucleic acid sequences encoding a VL chain can be generated, which can represent all possible permutations of recombined segments or subsets thereof. For example, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000 (10.sup.3), 2.times.10.sup.3, 3.times.10.sup.3, 4.times.10.sup.3, 5.times.10.sup.3, 6.times.10.sup.3, 7.times.10.sup.3, 8.times.10.sup.3, 9.times.10.sup.3, 10.sup.4, 5.times.10.sup.4, 10.sup.5 or more VL nucleic acid sequences can be generated.
[0464] The examples exemplify collection and storage of sequences compiled by practice of the method in a SequenceHistory.txt file. Such a file represents sequences generated in the Examples by the exemplified software and ordered for DNA synthesis. Sequences also can be stored manually, for example, in spreadsheets or lists.
[0465] ii. Determining Sequence Diversity of Collected Nucleic Acid Sequences
[0466] In some examples, recombined nucleic acid molecules can be collected and stored based on their sequence diversity. It is contemplated herein that knowledge of the sequence diversity of library members can be employed to select a restricted subset of nucleic acid sequences encoding VH chain and VL chain for synthesis and expression as described herein below. Hence, resulting antibody libraries can be made to maximize sequence diversity among members due to sequence differences. Alternatively, resulting antibody libraries can be made to minimize sequence diversity among members due to sequence similarities. Thus, for example, the sequence of a selected recombined nucleic acid can be compared to all other sequences in the libraries, and those sequences that are different (e.g. having sequence similarity that is less then 70%, e.g., 10%, 20%, 30%, 40%, 50%, 60%, 65%) can be selected. In another example, if a "Hit" is identified in an initial screen, a further library can be created where all members have a high sequence similarity (e.g. 70%, 75%, 80%, 85%, 90%, 95% or more) to the identified "Hit." The percentages given are for exemplification only. One of skill in the art can choose any desired limit of sequence similarity by which to select sequences for inclusion in a particular library.
[0467] To determine the sequence similarity or difference between and among recombined nucleic acid sequences sequence diversity based on sequence similarity of all collected nucleic acid sequences is assessed. Typically, due to the degeneracy of the genetic code, recombined nucleic acid sequences are first translated to give an amino acid sequence, and then sequence similarity between and among the resulting amino acid sequences is determined. Translation is performed based on the genetic code, whereby 64 codons encode the 20 amino acids plus three stop codons (see Table 20). Translation of each sequence can be performed manually or by other computer-based or automated methods. One of skill in the art is familiar with methods of translating proteins. The sequences can be grouped or stored based on their sequence diversity.
[0468] Typically, sequence diversity is assessed based on sequence similarity of two or more sequences, such as for example, as determined by alignment. One of skill in the art is familiar with various techniques to determine the sequence similarity (e.g. identity) between and among sequences. For example, sequence similarity can be determined manually by determining nucleotide differences between and among sequences. Sequence similarity or sequence identity of nucleotide or amino acid sequences also can be determined using conventional software or computer programs. Such algorithms are well known to one of skill in the art. For example, to find the best segment of identity or similarity of sequences, BLAST (Altschul et al (1990) J. Mol. Biol. 215:403-410 and Lipman et al (1990) J. Mol. Biol. 215:403-410), FASTA (Lipman et al (1985) Science 227:1435-1441), or Smith and Waterman (Smith and Waterman (1981) J. Mol. Biol. 147:195-197) homology search programs can be used. To perform global alignments, sequence alignment programs such as the CLUSTAL W (Thompson et al (1994) Nucleic Acids Research 22:4673-4680) can be used.
[0469] For example, nucleic acid or amino acid sequences can be assessed for sequence similarity using BLAST. Parameters for sufficient similarity to determine relatedness are computed based on well known methods for calculating statistical similarity. Exemplary parameters for determining relatedness of two or more sequences using the BLAST algorithm, for example, can be as set forth below. Briefly, amino acid sequence alignments can be performed using BLASTP version 2.0.8 (Jan. 5, 1999) and the following parameters: Matrix: 0 BLOSUM62; gap open: 11; gap extension: 1; x_dropoff: 50; expect: 10.0; wordsize: 3; filter: on. Nucleic acid sequence alignments can be performed using BLASTN version 2.0.6 (Sep. 16, 1998) and the following parameters: Match: 1; mismatch: 2; gap open: 5; gap extension: 2; x_dropoff: 50; expect: 10.0; wordsize: I 1; filter: off. Those skilled in the art will know what modifications can be made to the above parameters to either increase or decrease the stringency of the comparison, for example, and determine the relatedness of two or more sequences. The BLAST program provides an output indicator, the BLAST bit score, which is a value calculated from the number of gaps and substitutions associated with each aligned sequence. The higher the score, the more significant the alignment. The bit score can be used to select sequences that have either the most sequence diversity or alternatively, the least sequence diversity to every other selected sequence.
[0470] In another example, sequence diversity also can be assessed by comparison of two or more amino acid or nucleic acid sequences by alignment methods, e.g., the CLUSTAL method. (See, e.g., Higgins, D. G. and P. M. Sharp (1988) Gene 73:237-244.) The CLUSTAL algorithm groups sequences into clusters by examining the distances between all pairs. The clusters are aligned pairwise and then in groups. The percentage similarity between two amino acid sequences, e.g., sequence A and sequence B, is calculated by dividing the length of sequence A, minus the number of gap residues in sequence A, minus the number of gap residues in sequence B, into the sum of the residue matches between sequence A and sequence B, times one hundred. Gaps of low or of no similarity between the two amino acid sequences are not included in determining percentage similarity. Sequence similarity (e.g. sequence identity) between nucleic acid sequences can also be counted or calculated by other methods known in the art, e.g., the Jotun Hein method. (See, e.g., Hein, J. (1990) Methods Enzymol. 183:626-645.) Identity between sequences can also be determined by other methods known in the art, e.g., by varying hybridization conditions. BLASTclust is another program that can be used for cluster analysis. BLASTclust is used in the software compilation program described in the Examples.
[0471] Diversity and cluster information as well as BLAST bit score information provide the user with several options when selecting sequences. For example, the user can create an antibody library where the selected sequences are as diverse as possible. To do this, the user can use the diversity score and cluster information, and select sequences from different clusters that have the highest diversity. Alternatively, for example, the user can create a an antibody library where one sequence is initially selected and all subsequent sequences are as similar as possible to the first sequence. This can be accomplished by using the BLAST function. The user can BLAST the selected first sequence, and then select all the other sequences for the library using the BLAST bit score, choosing sequences with the highest score and therefore the highest sequence similarity. For example, Example 5 describes implementation of assessing sequence diversity between and among recombined sequences using Software Compilation software. The Example illustrates that BLAST can be performed on all sequences and Blast bit scores calculated to identify the sequence similarity or differences between sequences.
[0472] iii. Generating Nucleic Acid Molecules from Recombined Sequences
[0473] a) Synthesis
[0474] Where desired, the sequences can be individually synthesized into nucleic acid molecules. All collected sequences can be synthesized, or a subset of sequences synthesized. Nucleic acid molecules encoding VH or VL chain can be synthesized by methods known to one of skill in the art using synthetic gene synthesis (see e.g., U.S. Pat. Nos. 4,652,639; 5,132,215; 5,093,251; 6,110,668; 6,472,184; published U.S. application Nos. US20060281113; US20070004041; US20070122817; and International PCT Published Application Nos. WO98/15567; WO99/47536; WO00/75364; WO2004035781; WO2005071077). These include standard solid phase polypeptide synthesis methods involving synthesis of single stranded oligos that are ligated together. Methods also include methods using standard triplets that act as universal building blocks that represent all possible sequence combinations, and can be combined in a series of reaction steps (Slonomics.RTM.).
[0475] Nucleic acids can be synthesized that are 20, 50, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000 or more base pairs in length. Gene synthesis can be performed by automated methods. Any of the known synthesis methods can be used to produce the nucleic acid molecules. For example, companies exist for the purpose of synthesizing oligonucleotides and genes, for example, Integrated DNA Technologies (IDT) (Coralville, Iowa), TriLink Biotechnologies (San Diego, Calif.), Blue Heron Gene Synthesis (Bothell, Wash.), and Sloning Biotechnology (Puchheim, Germany).
[0476] The nucleotide monomers used in the synthesis can be purine and pyrimidine deoxyribonucleotides (adenosine (A), cytidine (C), guanosine (G) and thymidine (T)) or ribonucleotides (A, G, C and U (uridine)), or they can analogs or derivatives of these nucleotides, such as peptide nucleic acid (PNA), phosphorothioate DNA, and other such analogs and derivatives or combinations thereof. Other nucleotide analogs are well known in the art and can be used in synthesizing the oligonucleotides provided herein.
[0477] The nucleic acid molecules can be synthesized with nucleotide modifications. In one example, each oligonucleotide contains a terminal phosphate group, for example, a 5' phosphate group. For example, when it is desired to seal nicks between two adjacent oligonucleotides, e.g. following hybridization of the two oligonucleotides to a common opposite strand polynucleotide according to the methods herein, a 5' phosphate group is added to the end of the oligonucleotide whose 5' terminus will be joined with the 3' terminus of another oligonucleotide to seal the nick. In one example, a 5' phosphate (PO.sub.4) group is added during oligonucleotide synthesis. In another example, a kinase, such as T4 polynucleotide kinase (T4 PK) is added to the oligonucleotide for addition of the 5' phosphate group. Other oligonucleotide modifications are well-known and can be used with the provided methods.
[0478] The synthetic oligonucleotides can be chemically synthesized. Methods for chemical synthesis of oligonucleotides are well-known and involve the addition of nucleotide monomers or trimers to a growing oligonucleotide chain. Typically, synthetic oligonucleotides are made by chemically joining single nucleotide monomers or nucleotide trimers containing protective groups. For example, phosphoramidites, single nucleotides containing protective groups, can be added one at a time. Synthesis typically begins with the 3' end of the oligonucleotide. The 3' most phosphoramidite is attached to a solid support and synthesis proceeds by adding each phosphoramidite to the 5' end of the last. After each addition, the protective group is removed from the 5' phosphate group on the most recently added base, allowing addition of another phosphoramidite. See, for example, Behlke et al. "Chemical Synthesis of Oligonucleotides" Integrated DNA Technologies (2005), 1-12; Allen et al. "Ultramers.TM.--The Longest Oligonucleotides Available with Mass Spectrometry" Integrated DNA Technologies, Technical Report (2007); and McBride and Caruthers Tetrahedron Lett. 24:245-248, which describe synthesizing oligonucleotides using standard cyanoethyl chemistry (using phosphoramidite monomers and tetrazole catalysis). Such methods typically result in generation of oligonucleotides of 100-200 bases.
[0479] Thus, to synthesize larger genes, methods include annealing of a series of smaller oligonucleotides. In such a method, individually designed oligonucleotides are made, such as by using an automated DNA synthesizer, purified and connected by specific annealing using standard ligation or polymerase reactions. Generally, the oligos are designed with overlapping stretches of common sequence to permit annealing. Several methods of gene synthesis have been described, including, but not limited to the ligation of phosphorylated overlapping oligonucleotides (Gupta, N. K. et al. (1968) Studies on polynucleotides, 88. Enzymatic joining of chemically synthesized segments corresponding to the gene for alanine-tRNA. Proc. Natl Acad. Sci. USA, 60, 1338-1344; Fuhrmann M et al., A synthetic gene coding for the green fluorescent protein (GFP) is a versatile reporter in Chlamydomonas reinhardtii. Plant J. 1999 August; 19(3):353-61); de novo gene construction using Ultramers (Allen et al. "Ultramers.TM.--The Longest Oligonucleotides Available with Mass Spectrometry" Integrated DNA Technologies, Technical Report (2007); the Fok I method (Mandecki, W. and Bolling, T. J. (1988) FokI method of gene synthesis. Gene, 68, 101-107); a modified form of ligase chain reaction for gene synthesis; PCR assembly whereby the full-length molecule is generated progressively by overlap extension (Stemmer, W. P., Crameri, A., Ha, K. D., Brennan, T. M. and Heyneker, H. L. (1995) Single-step assembly of a gene and entire plasmid from large numbers of oligodeoxyribonucleotides. Gene, 164, 49-53), thermodynamically balanced inside-out (Gao X, Yo P, Keith A, Ragan T J, Harris T K. Thermodynamically balanced inside-out (TBIO) PCR-based gene synthesis: a novel method of primer design for high-fidelity assembly of longer gene sequences. Nucleic Acids Res. 2003 Nov. 15; 31(22):e143) or combined approaches (Young L, Dong Q. Two-step total gene synthesis method. Nucleic Acids Res. 2004 Apr. 15; 32(7):e59). Since the error frequency increases with longer oligonucleotides, methods typically include using shorter oligonucleotides (200 base pairs or less) assembled together.
[0480] The synthesized molecules can be purified by a number of well-known methods, for example, high-performance liquid chromatography (HPLC), thin layer chromatography (TLC), Polyacrylamide Gel Electrophoresis (PAGE) and desalting.
[0481] In one embodiment, the synthesized nucleic acids are arrayed in multiwell plates, with each individual well of a plate corresponding to one individual nucleic acid. More specifically, each individual locus of a plate contains an nucleic acid encoding an antibody variable region, either heavy or light. The identity of the nucleic acid contained within each well of the multiwell plate is known and mapped to an output file that contains the nucleic acid sequences for all of the nucleic acids within the plate. Multiwell plates can include but are not limited to 96-well plates, 384-well plates, and 1536-well plates. In an exemplary embodiment, the nucleic acids are spatially arrayed in a 96-well plate.
[0482] Upon synthesis, the resulting nucleic acid molecules are individually addressed into a locus (e.g. a well, chip, tag, and other addressable formats). Each individual locus of a plate can contain a different recombined and synthesized nucleic acid molecule encoding for either a heavy chain variable region or a light chain variable region or portion thereof compared to all other addresses. The identity of the nucleic acid molecule at each locus is known and can be mapped to an output file that contains the sequences for all the nucleic acid molecules within the addressable format. For example, nucleic acid molecules can be addressed by spatial array into multiwell plates, with each individual locus of a plate containing one individual nucleic acid molecule. Multiwell plates can include but are not limited to 12-well plates, 24-well plates, 96-well plates, 384-well plates, and 1536-well plates.
[0483] b) Recombinant Generation
[0484] In some examples, recombined VH and/or VL sequences, or a subset thereof, can be generated into recombined nucleic acid molecules using recombinant DNA technology. One of skill in the art is familiar with general recombinant DNA techniques, including but not limited to, PCR, cloning and restriction enzyme digestion. Such techniques can be used to combine germline segments as discussed herein above to generate recombined nucleic acid molecules that are in-frame. Generally, each vector is generated individually, such that the identity of the sequence of each vector is known through the cloning process. Thus, the recombinant generation of a combinatorial antibody library is addressable.
[0485] In the methods of generating combinatorial antibody libraries using recombinant DNA techniques, germline segments can be linked directly or indirectly by a linker so long as the resulting nucleic acid molecule is in-frame, resulting in a functional and productive antibody. The linker can be a peptide, polypeptide or an amino acid. For example, it is understood that by virtue of using recombinant DNA technologies, including the use of restriction enzymes, that amino acids can be inserted between V-D, D-J and V-J junctions in order to facilitate joining of germline segments. Exemplary of a linker as described herein in Example 14 is a sequence of nucleotides encoding an SY between the 3'end of the D.sub.H germline segment and the 5'end of the J.sub.H germline segment.
[0486] In methods of generating a combinatorial antibody library by recombinant DNA techniques, a parent vector or vectors can be generated that contain common nucleic acid sequences between and among members of the library. For example, a vector can be generated that contains nucleic acid sequence for a V.sub.H, D.sub.H and/or J.sub.H germline segment, modified forms thereof, or portions thereof and/or a V.sub.L and/or J.sub.L that are common between all members of the library. It is understood that introduction of segments is with reference to the reading frames as described herein above, such that the resulting compiled nucleic acid molecule is in-frame. The description below provides a general summary of a method of generating a combinatorial antibody library using recombinant DNA techniques. It is understood that the reference to the examples is for exemplification only. Using the description provided herein one of skill in the art can generate similar vectors containing nucleic acid compiled from germline segments or modified forms thereof to generate recombined VH or VL chains that are in-frame. For example, it is understood that the order of addition of V.sub.H/D.sub.H/J.sub.H or V.sub.L/J.sub.L segments or portions thereof to the recombinant vectors can occur in any order, so long as the resulting cloned nucleotide sequence encodes a recombined VH or VL chain that is in-frame.
[0487] Thus, a parent vector is typically generated containing a sequence common to all members of the library. For example, if all vectors share a common V.sub.H germline sequence, a vector can be generated carrying the V.sub.H germline sequence in its correct reading frame, which can be manipulated for subsequent inclusion of other germline segments. For example, the V.sub.H germline sequence can be modified to include restriction enzyme sites on the 3'end for subsequent joining with a D.sub.H germline sequence. In another example, vectors can be generated containing a portion of a V.sub.H, D.sub.H or J.sub.H germline sequence. For example, a vector can be generated containing a common framework consensus sequence as described elsewhere herein. This is exemplified in Example 14 where a modified Plasmid A vector was generated to contain a common framework 4 region of a J.sub.H germline segment. An exemplary parent vector for use in generating a combinatorial antibody library using the methods herein is set forth in SEQ ID NO:2051, which contains a common V.sub.H germline segment (VH3-23 (IGHV3-23*01) that is modified to remove an internal restriction site and add additional restriction sites at the 3'end) and a common framework 4 region of a JH germline segment.
[0488] The parent vector can then be used to further introduce remaining germline segments, modified forms thereof, or portions thereof such that a plurality of final vectors are generated each containing a nucleic acid sequence encoding a recombined VH and/or VL chain. The generation from a parent vector to a final vector can occur in steps, thereby resulting in intermediate vectors, generally at least one intermediate vector. Generally, nucleic acid sequences for subsequent germline segments, modified forms thereof or portions thereof are generated as oligonucleotides for subsequent cloning into the parent vector or an intermediate vector. It is understood that if a stop codon is inserted at any step, the stop codon is either removed as described herein above, or the particular segment containing the stop codon is not cloned. To facilitate joining with adjacent nucleic acid sequences, the oligonucleotides are generated to contain complementary restriction enzyme sites at the 3' and/or 5'ends.
[0489] For example, depending on the components contained in the parent vector, an intermediate vector can be generated to contain remaining germline segments, modified forms thereof or portions thereof. For example, intermediate vectors can be generated from the parent vector above (set forth in SEQ ID NO:2051), whereby each intermediate vector contains a different J.sub.H segment in its correct reading frame (see e.g. Table 13). The J.sub.H segment can be a germline segment or a modified form thereof. Exemplary of modified J.sub.H segments are any set forth in SEQ ID NOS: 3450-3455 and encoding J.sub.H regions set forth in any of SEQ ID NOS: 3456-3461. The entire J.sub.H segment or a portion of a J.sub.H segment can be added to an existing parent or intermediate vector. For example, if a parent vector is made to contain a consensus framework 4 region as described above, a portion of a J.sub.H segment containing nucleotides corresponding to the last portion of a CDR3 in the J.sub.H segment can be introduced. By virtue of the addition of different segments, for example different J.sub.H segments, in the intermediate vectors, the diversity of the library can be increased. Thus, generally, a plurality of intermediate vectors are generated. For example, Example 14 describes the generation of six intermediate vectors (having a sequence set forth in any of SEQ ID NOS: 2064-2069).
[0490] A plurality of final vectors are generated that contain the all components of a compiled germline sequence. As above, the remaining nucleotide to be inserted into the vector are generated as oligonucleotides, and typically contain complementary restriction enzyme sites at the 3' and/or 5'ends. As noted, the oligonucleotides are generated to provide the correct reading frame for the inserted segment and do not contain stop codons. In addition, the oligonucleotides are generated to preserve existing reading frames for the segments contained in the parent or intermediate vectors. For example, as described elsewhere herein, it is understood that the reading frame of the D.sub.H region is not critical. Thus, D.sub.H segments, including D.sub.H germline segments, can be inserted in any reading frame, or can be a random sequence of nucleotides. Example 14 exemplifies generation of a plurality of final vectors by introduction of D.sub.H germline segments (e.g., any set forth in any of SEQ ID NOS: 239-245, 248, 250, 252, 254, 256, 258-272), or inverted segments thereof (e.g., any set forth in any of SEQ ID NOS: 3462-3488), in all three reading frames. In generating the oligonucleotides, however, one or more nucleotides are removed or added from the 3' or 5'ends in order to preserve reading frames of the adjacent J.sub.H segments. This is exemplified in Example 14, which sets forth conditions for removing or adding nucleotides in order to preserve reading frames.
[0491] The resulting final vectors contain compiled V.sub.H/D.sub.H/J.sub.H or V.sub.L/J.sub.L germline segments, or modified forms thereof, that encode a recombined VH chain or VL chain. Each final vector is different and contains a different nucleic acid sequence encoding a different recombined VH chain or VL chain. Since the nucleic acid molecules are already cloned into an expression vector, they can be directly transformed into an expression system as discussed in Section f below.
[0492] f. Expressing and Producing Antibodies or Portions or Fragments Thereof
[0493] In the methods provided herein, recombined nucleic acid molecules, such as synthetic recombined nucleic acid molecules or recombined nucleic acid molecules generated recombinantly, are cloned into an expression vector. The polynucleotides typically are inserted into the vectors using restriction digest and ligation. Any conventional vector known to one of skill in the art can be used for expression in eukaryotic or prokaryotic cells. Exemplary vectors include plasmid A, C and D described herein below. The vector can be used to transform any expression system compatible therewith for amplification of the nucleic acid and/or expression of the encoded variable heavy or variable light chain polypeptide.
[0494] Typically, ligation into a vector is in an addressable format such that the identity of the recombined polypeptide expressed therefrom is known. For example, the vectors containing nucleic acids are spatially arrayed in multiwell plates, with each individual locus of a plate containing a vector with one individual nucleic acid inserted. More specifically, each individual locus of a plate contains a vector encoding for either a heavy chain or a light chain. The identity of the nucleic acid contained within each well of the multiwell plate is known, for example, by mapping to stored sequences collected from the compilation or synthesis above. For example, ligation into vectors can be performed directly into multiwall plates already containing synthesized nucleic acid molecules from above. Multiwell plates can include but are not limited to 96-well plates, 384-well plates, and 1536-well plates. In an exemplary embodiment, the nucleic acids are spatially arrayed in a 96-well plate.
[0495] Generally in practicing the methods, a nucleic acid molecules encoding a variable heavy chain is ligated into a first vector. A nucleic acid molecule encoding a variable light chain is ligated into a second vector. The first vector and second vector can be co-transformed into the same expression host for co-expression of a variable heavy chain and a variable light chain. The polypeptides, upon expression, will become operably joined by virtue of interactions between the heavy and light chain polypeptides. In some examples, it is possible to operably join the nucleic acid molecules directly prior to expression, such as by including a linker. In such examples, a single nucleic acid molecule encodes a variable heavy chain and a variable light chain, and can be ligated into a single vector for expression thereof.
[0496] In all methods herein, the expressed antibodies minimally include a VH chain and a VL chain, or portions thereof sufficient to form an antigen-binding site. In addition, if desired, a constant chain can be included for expression in operative linkage with the variable chains. In all examples of the methods, the recombined nucleic acid molecules, upon expression and ligation, encode for antibodies or fragments thereof, including, but not limited to an IgG, a Fab fragment, a F(ab').sub.2 fragment or a Fv fragment, such as a disulfide-linked Fv or a single chain Fv. An exemplary antibody is a Fab fragment. Such antibodies or fragments thereof can be purified by any methods known to one of skill in the art.
[0497] Section F describes methods of expressing and purifying antibodies or fragments thereof.
[0498] 2. Automation
[0499] Any of the steps of the method described above can be automated and/or made high-throughput and/or otherwise rendered more efficient or fast. One of skill in the art is familiar with methods of automation of systems and processes, including the implementation of in silico databases, application of computer programs, robotics and/or other high-throughput methods that can be used in practice of the method. It is contemplated that the entire process of the method can be automated or only a few steps can be automated. The choice of automation is up to the user. The description below and examples exemplify automation of various processes of the method.
[0500] a. User-Created Database
[0501] To practice the methods herein, sequences of germline segments or modified forms thereof must be obtained. Such sequences are known to one of skill in the art and can be obtained from commercially available databases, such as described above. Such germline segment sequences are set forth in the sequence listing as set forth in Tables 3-5 above. Exemplary of modified J.sub.H germline segments are set forth in SEQ ID NOS: 3450-3455. The sequences can be compiled into a user-created database for ease of access. Generation of a file or database containing all of the sequence information provides immediate access to these sequences. In addition the sequence file can be linked to other systems and processes to facilitate performance of the method. For example, as exemplified in FIG. 9, in Example 4, a database file is linked to the Sequence Compilation Alogrithm as an input file for identification of V(D)J heavy and light chain sequences for sequence compilation.
[0502] The database file contains sequences for germline V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda. segments. In particular, the database file can contain sequences of nucleic acids set forth in any of SEQ ID NOS:10-451, 868, or a subset thereof. It is helpful if the sequences are specified using FASTA format and all sequences contain a blank line between them. For purposes of practice of the method, the J.sub.H, J.sub..kappa. and J.sub..lamda. segment sequences are set forth in the database file in coding frame triplets corresponding to their optimal reading frame, which is set forth in Table 13 above. The sequences in the database file are named for identification. For example, germline segments are identified by section title headings [VH], [DH], [JH], [VK], [JK], [VL], and [JL]. Such a databased file is described in the Examples (e.g. Example 3) as a SequenceDatabase.txt.file. FIG. 11 provides a schematic illustration of a Sequence Database file format.
[0503] In addition, the database can contain other sequences used in practicing the method. For example, the database can contain nucleic acid sequences for restriction sites, and can be identified in the database under the section title heading [Restriction Sites]. These sequences can be accessed by particular program processes as described below to identify nucleic acid sequences corresponding to restriction sites within a recombined nucleic acid molecule. Restriction site sequences contained in the database include any of SEQ ID NOS:977-980, 1889-1900. Any restriction site sequence known to one of skill in the art can be contained in the database file. For example, the schematic illustration of a database file in FIG. 11 includes a sequence for the restriction enzyme Mfe I.
[0504] It is contemplated that the database file can be periodically updated to contain additional sequences. The database file also can be updated to include any sequence contemplated for practice of the method. For example, nucleic acid sequences that encode for proteins other than antibody germline segments can be entered into the database, using FASTA format, under an appropriate heading. These sequences are then available to be recombined into the germline antibody sequences. For example, one can insert peptide sequences into an antibody at D.sub.H by including nucleic acid sequences encoding for the peptide under the section title [DH].
[0505] b. Sequence Compilation
[0506] The method of compilation of sequence can be performed in silico, for example, using software. Any software programmed to perform an algorithm or process allowing for compiling germline segments in accordance with the method herein or any suitable method, can be used. One of skill in the art familiar with software programming can generate a computer program capable of performing such an algorithm or process. Generally, the software is programmed to perform any one or more of the following processes:
(a) accessing a user-created in silico database of all available antibody germline segments (V.sub.H, D.sub.H, J.sub.H, V.sub..kappa., J.sub..kappa., V.sub..lamda. and J.sub..lamda.); (b) applying an algorithm to generate every possible recombined full length nucleic acid sequence for heavy chains (5'-V.sub.H-D.sub.H-J.sub.H-3' combinations), every possible recombined full length nucleic acid sequence for kappa light chains (5'-V.sub..kappa.-J.sub..kappa.-3' combinations) and every possible recombined full length nucleic acid sequence for lambda light chains (5'-V.sub..lamda.-J.sub..lamda.-3' combinations); (c) applying an algorithm to modify the nucleic acid sequences of the joints so that the resulting nucleic acids sequences are in frame; (d) modifying the resulting nucleic acid sequences of the joints to remove any inadvertently generated stop codons; (e) modifying the resulting full length nucleic acid to optimize codon usage for bacterial expression; (f) modifying the resulting nucleic acid to remove any undesired restriction sites; (g) inserting flanking nucleic acids containing restriction sites for cloning at the 5' and 3' termini of the optimized full length nucleic acid sequences; (h) ranking recombined nucleic acid sequences based on their sequence diversity; (g) selecting recombined nucleic acid sequence(s) (encoding either a heavy chain variable region or a light chain variable region) from the library of recombined nucleic acid sequences; (h) assigning the selected nucleic acid sequence to a unique locus of an addressable format; (i) generating an output file that contains all recombined nucleic acid sequences in the form of the addressed format that lists the distinct heavy chain or light chain sequences such that each locus is addressed and corresponds to a locus of the addressed format (e.g. 96-well plate).
[0507] Provided herein are software, computer-readable media, and computer systems for performing the method as described herein. The Examples describe an exemplary software, computer-readable media, computer system and systems. It is understood that those of skill in the art can modify such software, computer-readable media, computer systems and systems based upon this disclosure and that such modifications are included herein.
[0508] For example, each of these processes of the method described herein is performed by an exemplary computer software exemplified in the Examples herein. For example, Example 2 and FIGS. 8 and 9 describe an exemplary process for sequence compilation of germline segments. The flow chart in FIG. 9 describes processes for each of steps (a)-(d) and (f) above. In addition, Example 2 and FIG. 10 describes an exemplary process for ranking sequences based on sequence diversity. The flow chart in FIG. 10 describes the process used to rank sequences after determination of diversity scores and an example of the ranking is set forth in FIG. 17.
[0509] c. Automation of Protein Expression and Purification Methods of automating protein expression and purification are known to one of skill in the art (see, e.g., Lesley et al. (2001) Protein Expression and Purification, 22:159-164; Acton T B et al. (2005) Methods Enzymol., 394:210-43; Nguyen et al. (2004) Journal of Structural and Functional Genomics, 5:23-27). Such processes typically include robotic methods.
[0510] Exemplary of a high-throughput automated method of protein expression and purification is Piccolo.TM. (Wollerton et al. (2006) JALA, 11:291-303). The Piccolo.TM. system automates protein expression and purification of proteins from both E. coli and baculovirus mediated insect cell expression systems. Piccolo is able to perform multiple different protein expression and purifications in parallel. The Piccolo system utilizes a 24-position culture vessel block (CVB) in an aeration assembly that supports the expression and purification of multiple samples at once. The Piccolo system contains four modules that perform these functions: a liquid handling module, CVB incubators, a centrifuge and storage carousels. A rail mounted 6-axis RX60L Spinal Transfer Robot (ST Robot; Staubli, Horgen, Switzerland) moves the lab ware between the liquid handling module, incubators, centrifuge and storage carousels. The system is controlled by software that permits the user to control expression and purification conditions.
[0511] Expression can be initiated by inoculation of CVB plates containing appropriate growth medium with an input inoculum, such as a bacterial culture. A total of 576 individual cultures can be grown at any one time, corresponding to 24 culture vessel blocks. The plates can be incubated under user-specified periods and conditions. Typical growth and induction temperatures range from 16.degree. C. to 37.degree. C. Selection of optimal temperatures for growth and induction is well within the level of skill of the skilled artisan. Bacterial growth can be monitored. If desired, protein expression can be induced by adding an appropriate amount of inducer into the CVB plate assembly and further grown under appropriate conditions. Protein expression can be induced by addition of any inducer compatible with the expression vector, including isopropyl .beta.-D-1-thiogalactopyranoside (IPTG) and arabinose. Expression times range from 2 hours to 48 hours. Selection of optimal expression times is well within the level of skill of the skilled artisan. Following expression, plates can be stored under cooling conditions. For example, as set forth in Example 9, spatially arrayed transformed cells are mapped to a 24-well culture vessel block for cell growth and protein expression. For each 96-well plate of transformed cells, four culture vessel blocks are generated, thereby allowing the growth of Fabs corresponding to every well of the 96-well plate.
[0512] Following expression of the desired protein, the Piccolo.TM. system also can be used to purify the resulting proteins. The Piccolo machine is programmed to perform lysis and purification steps. The cells are harvested and lysed using an appropriate lysis buffer that is compatible with the purification technique. Selection of a lysis buffer is well within the level of skill of the skilled artisan. The resulting supernatant is then purified by column chromatography with an appropriately modified resin, such as an anti-flag resin or Ni-charged resin. One of skill in the art can identify an appropriate resin for protein purification as described elsewhere herein. The resin should be manually equilibrated in an appropriate wash buffer before starting the run. The bound protein can be eluted with an appropriate elution buffer and the eluate collected in an output plate. The output plate can be stored at cool temperatures (e.g. 6.degree. C.) until collected by the operator.
[0513] Purity can be assessed by any method known in the art including gel electrophoresis and staining and spectrophotometric techniques. Additional purification methods can be combined with Piccolo such as are described elsewhere herein. For example, proteins can be further purified using an orthogonal secondary high throughput method of protein purification (see e.g. Example 10). Additional column chromatography can be performed using a compatible resin, an Aktapurifier (GE Healthcare) and an autosampler. Exemplary of purifying antibodies, a protein G resin can be utilized.
E. LIBRARIES
[0514] Provided herein are libraries. The libraries include nucleic acid libraries encoding VH or VL chains, vector libraries transformed with recombined nucleic acid molecules, and antibody libraries. In some examples, the members of each of the libraries are addressed in an addressable format, such as any discussed in Section E.2. The members of the libraries and the resulting libraries can be produced by the methods described herein above.
[0515] 1. VH Nucleic Acid Libraries and Vector Libraries Thereof
[0516] Provided herein are recombined nucleic acid libraries encoding VH chains. The libraries provided herein include recombined nucleic acid molecules made up entirely of V.sub.H, D.sub.H and J.sub.H germline segments or modified forms thereof. The V.sub.H, D.sub.H and J.sub.H germline segments include any set forth in Table 3 above, modified forms thereof, or a subset thereof. Any permutation is possible. The resulting nucleic acid molecule in the library have a sequence such that the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment. The segments can be linked directly or indirectly by a peptide linker.
[0517] Because the nucleic acid molecules in the library are derived from germline segments, members of such a nucleic acid library are capable of encoding a naive antibody when co-expressed with a nucleic acid encoding a VL chain. It is understood that the library is considered to be naive and derived from germline even though, in practicing the method herein, the joint regions of the segments are altered to render the resulting encoding nucleic acid molecules in frame. Such alterations, however, are minor and variously include insertion or deletion generally of only a single nucleotide of a germline segment. In addition, other modification made to the recombined nucleic acid sequence by virtue of practice of the method herein, such as removal of stop codons and restriction enzyme site sequences, also result in naive antibodies.
[0518] It is understood that libraries can be generated that are compiled from sequences that include modified germline segments. In some examples of the libraries, the libraries include recombined nucleic acid molecules made up entirely of a V.sub.H and a J.sub.H germline segment, and also any sequence of nucleotides between the V.sub.H and J.sub.H germline segment. This is the region that includes the central portion of the CDRH3, which is largely responsible for the antigen specificity of the resulting antibody. The sequence of nucleotides can be any random sequence of nucleotides. In some instances, the sequence of nucleotides is a sequence that encodes a peptide mimetic against any desired target, for example, a cell surface receptor. Exemplary peptide mimetics are set forth in Table 16. Generally, the sequence of nucleotides is or is about 5, 10, 15, 20, 25, 30, 40, 50, 60, 70, 80, 90 or more nucleotides in length. The resulting nucleic acid molecule in the library have a sequence such that the V.sub.H segment is 5' to the random sequence of nucleotides which is 5' to the J.sub.H segment. In some examples, the random sequence of nucleotides is a D.sub.H germline segment.
[0519] In other examples, the libraries provided herein include recombined nucleic acid molecules where at least one, two or all three of a V.sub.H, D.sub.H and J.sub.H germline segment contained therein are modified, for example, due to modification by insertion, deletion or addition of amino acids. For example, the libraries include nucleic acid molecules containing sequences encoding a directed peptide. The libraries also include recombined nucleic acid molecules containing nucleotide mutations encoding amino acid replacements, for example, of one or more amino acids of a CDR. In an additional example, the libraries provided herein include recombined nucleic acid molecules where at least a portion of the nucleic acid molecule, such as the entire nucleic acid molecule encoding the VH chain, or at least one or more of a V.sub.H, D.sub.H and J.sub.H are derived from an existing monoclonal antibody, including, but not limited to, any monoclonal antibody set forth in Table 9. For example, exemplary libraries provided herein can include a nucleic acid molecule encoding the VH chain of an anti-CD20 antibody such as is set forth in SEQ ID NO:1043 or SEQ ID NO:1058 (SEQ ID NO:453 including terminal restriction site sequences) or Herceptin such as is set forth in SEQ ID NO:1057 (SEQ ID NO:452 including terminal restriction site sequences)
[0520] Libraries of recombined nucleic acid molecules provided herein can include members that represent one, some or all of the above examples. Any of the libraries provided herein also can include members whose sequences include heterologous sequence, for example, restriction site sequences, linker sequences, sequences encoding tags or other detectable moieties or other sequences.
[0521] In the VH nucleic acid libraries provided herein, each recombined nucleic acid molecule member of the library is productive and, when co-expressed with a nucleic acid molecule encoding a VH chain, generates a functional antibody or portion thereof that is sufficient to form an antigen-binding site. In addition, in the VH nucleic acid libraries provided herein, each nucleic acid member of the library is different. The VH nucleic acid libraries provided herein can contain 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 2000, 3000, 4000, 5000, 6000, 7000, 8000, 9000, 10,000 (10.sup.4), 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 2.times.10.sup.5, 3.times.10.sup.5, 4.times.10.sup.5, 5.times.10.sup.5, 6.times.10.sup.5, 7.times.10.sup.5, 8.times.10.sup.5, 9.times.10.sup.5, 10.sup.6 or more different members. The nucleic acid members are provided in addressed formats, such that when addressed the identity of each nucleic acid is known by its location in the array.
[0522] For example, an exemplary VH nucleic acid library includes members set forth in SEQ ID NOS: 454-805, each representing a different recombined nucleic acid molecule of V.sub.H, D.sub.H and J.sub.H germline segments. Such a library includes members containing heterologous sequence for restriction sites at the 3' and 5' ends. It is understood that members of the library also can include those having sequences not including the heterologous sequence, such as is set forth in any of SEQ ID NOS: 1059-1410.
[0523] In an additional example, an exemplary VH nucleic acid library includes members set forth in SEQ ID NOS: 2070-2759, each representing a different recombined nucleic acid molecule of V.sub.H, D.sub.H and J.sub.H germline segments or modified forms thereof. Such a library includes members containing heterologous sequence for restriction sites at the 3'end corresponding to CTAGC (set forth in SEQ ID NO:1903) and at the 5'end corresponding to CCATGGCA (set forth in SEQ ID NO:1901). It is understood that members of the library also can include those having sequences set forth in any of SEQ ID NOS: 2070-2759 that do no include the heterologous sequences at one or both of the 3' and 5'ends.
[0524] A VH nucleic acid library can include members from any of the libraries provided herein, or a subset thereof. For example, a VH nucleic acid library includes members set forth in any of SEQ ID NOS:454-805 and 2070-2759, or a subset thereof. The library members can include those containing heterologous sequences at the 3' or 5'ends, and/or members that do not include heterologous sequences.
[0525] In some examples, any of the nucleic acid sequences in the libraries provided herein can be included in a vector to generate vector libraries. Exemplary of vector libraries are libraries of recombined nucleic acid molecules encoding VH chain included in backbone Plasmid A or Plasmid D.
[0526] 2. VL Nucleic Acid Libraries and Vector Libraries Thereof
[0527] Provided herein are recombined nucleic acid libraries encoding VL chains. The libraries include those encoding for lambda or gamma light chains, or combinations thereof. Thus, the libraries provided herein include recombined nucleic acid molecules made up entirely of V.sub..kappa. and J.sub..kappa. germline segments and/or V.sub..lamda. and J.sub..lamda. germline segments. The V.sub..kappa. and J.sub..kappa. germline segments and/or V.sub..lamda. and J.sub..lamda. germline segments include any set forth in Tables 4-5 above, or a subset thereof. Any permutation is possible. The resulting nucleic acid molecules in the library have a sequence such that the V.sub.L segment (V.sub..kappa. or V.sub..lamda.) is 5' to the J.sub.L segment (J.sub..kappa. or J.sub..lamda.).
[0528] Because the nucleic acid molecules in the library are derived from germline segments, members of such a nucleic acid library are capable of encoding a naive antibody when co-expressed with a nucleic acid encoding a VH chain. It is understood that the library is considered to be naive and derived from germline even though, in practicing the method herein, the joint regions of the segments are altered to render the resulting encoding nucleic acid molecules in frame. Such alterations, however, are minor and variously include insertion or deletion generally of only a single nucleotide of a germline segment. In addition, other modification made to the recombined nucleic acid sequence by virtue of practice of the method herein, such as removal of stop codons and restriction enzyme site sequences, also result in naive antibodies.
[0529] In some examples, the libraries provided herein include recombined nucleic acid molecules where at least one or both a V.sub..kappa., and J.sub..kappa. germline segment or a V.sub..lamda. and J.sub..lamda. germline segment contained therein are modified, for example, due to modification by insertion, deletion or addition of amino acids. For example, the libraries include nucleic acid molecules containing sequences encoding a directed peptide. The libraries also include recombined nucleic acid molecules containing nucleotide mutations encoding amino acid replacements, for example, of one or more amino acids of a CDR. In an additional example, the libraries provided herein include recombined nucleic acid molecules where at least a portion of the nucleic acid molecule, such as the entire nucleic acid molecule encoding the VH chain, or at least one or more of a V.sub..kappa., and J.sub..kappa. or V.sub..lamda. and J.sub..lamda. are derived from an existing monoclonal antibody including, but not limited to, any monoclonal antibody set forth in Table 9. For example, exemplary libraries provided herein can include a nucleic acid molecule encoding the VL chain of Herceptin such as is set forth in SEQ ID NO: 1423 (SEQ ID NO:818 including terminal restriction site sequences) or the VL chain of an anti-CD20 antibody such as is set forth in SEQ ID NO: 1050 or SEQ ID NO:1440 (SEQ ID NO:835 including terminal restriction site sequences).
[0530] Libraries of recombined nucleic acid molecules provided herein can include members that represent one, some or all of the above examples. Any of the libraries provided herein also can include members whose sequences include heterologous sequence, for example, restriction site sequences, linker sequences, sequences encoding tags or other detectable moieties or other sequences.
[0531] In the VL nucleic acid libraries provided herein, each recombined nucleic acid molecule member of the library is productive and, when co-expressed with a nucleic acid molecule encoding a VH chain, generates a functional antibody or portion thereof that contains a sufficient antigen-binding site. In addition, in the VL nucleic acid libraries provided herein, each nucleic acid member of the library is different. The VL nucleic acid libraries provided herein can contain 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 2000, 3000, 4000, 5000, 6000, 7000, 8000, 9000, 10,000 (10.sup.4), 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 2.times.10.sup.5, 3.times.10.sup.5, 4.times.10.sup.5, 5.times.10.sup.5, 6.times.10.sup.5, 7.times.10.sup.5, 8.times.10.sup.5, 9.times.10.sup.5, 10.sup.6 or more different members. The nucleic acid members are provided in addressed formats, such that when addressed the identity of each nucleic acid is known by its location in the array.
[0532] For example, an exemplary VL nucleic acid library includes members set forth in SEQ ID NOS: 806-815, 817, 819-834, 836-867 each representing a different recombined nucleic acid molecule of V.sub..kappa., and J.sub..kappa. germline segments. Such a library includes members containing heterologous sequence for restriction sites at the 3' and 5 ends. It is understood that members of the library also can include those having sequences not including the heterologous sequence, such as is set forth in any of SEQ ID NOS: 1411-1422, 1424-1439, 1441-1472.
[0533] In some examples, any of the nucleic acid sequences in the libraries provided herein can be included in a vector to generate vector libraries. Exemplary of vector libraries are libraries of recombined nucleic acid molecules encoding VL chain included in backbone Plasmid C or Plasmid E.
[0534] 3. Paired Nucleic Acid Libraries or Vector Libraries Thereof
[0535] Also provided herein are libraries containing both recombined nucleic acid molecules encoding a VH chain and nucleic acid molecules encoding a VL chain, i.e. paired nucleic acid libraries. The paired libraries provided herein can contain a first nucleic acid molecule that is any of the nucleic acid members of the VH nucleic acid library in Section E.1 above and a second nucleic acid molecule that is any of the nucleic acid members of the VL nucleic acid library in Section E.2 above. The nucleic acid members in the paired libraries include those having heterologous sequence and those not having heterologous sequence. In some examples, one of the nucleic acid molecules in the pair can contain a heterologous sequence (e.g. a tag or other detectable moiety), while the other paired molecule at the locus in the library does not contain any heterologous sequence.
[0536] The paired nucleic acid libraries can be provided as addressed libraries. In such libraries, each locus of an addressed format contains one nucleic acid molecule encoding a VH chain and one nucleic acid molecule encoding a VL chain. Each nucleic acid pair (i.e. the combination of the nucleic acid molecule encoding the VH chain and the nucleic acid molecule encoding the VL chain) is different compared to all other pairs at all other addressed loci.
[0537] In some examples, the nucleic acid molecules can be contained in vectors to generate paired vector libraries. In such libraries, each locus of the addressed library includes a vector containing a nucleic acid molecule encoding a VL chain and a vector containing a nucleic acid molecule encoding a VH chain.
[0538] In some examples, the paired nucleic acid libraries can contain a common nucleic acid molecule encoding a VL chain such that each locus in the library contains the same nucleic acid molecule. In other examples, the paired nucleic acid libraries can contain a common nucleic acid molecule encoding a VH chain such that each locus in the library contains the same nucleic acid molecule. Generally, a library contains 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 5,000, 10,000 or more nucleic acid molecules encoding a VL chain and 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 5,000, 10,000 or more nucleic acid molecules encoding a VH chain. The resulting paired library contains 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 2000, 3000, 4000, 5000, 6000, 7000, 8000, 9000, 10,000 (10.sup.4), 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5, 2.times.10.sup.5, 3.times.10.sup.5, 4.times.10.sup.5, 5.times.10.sup.5, 6.times.10.sup.5, 7.times.10.sup.5, 8.times.10.sup.5, 9.times.10.sup.5, 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9, 10.sup.10 or more different paired members.
[0539] As described herein below, upon co-expression of a first and second nucleic acid molecule (e.g a nucleic acid molecule encoding a VH chain and a nucleic acid encoding a light chain), a library of antibodies can be generated. If the nucleic acid molecule further contains a sequence encoding a CH, a Fab library can be generated, whereby each member in the library contains a VH chain and a VL chain linked by a CH.
[0540] For example, an exemplary nucleic acid paired library includes those where a first nucleic acid is any one or more nucleic acids molecules set forth in any of SEQ ID NOS: 454-805 or 2070-2759 (each encoding a VH chain), and a second nucleic acid molecule set forth in any of SEQ ID NOS: 806-815, 817, 819-834, and 836-867 (each encoding a VL chain). The sequences set forth above contain heterologous sequence for restriction sites at the 3' and 5' ends. It is understood that nucleic acid libraries can be generated without the heterologous sequences. Thus, in some examples, members of the library also can include those having sequences not including a heterologous sequence. For example, an exemplary nucleic acid paired library includes nucleic acid sequences not containing heterologous sequence for restriction sites at the 3' and 5' ends from a first nucleic acid set forth in any of SEQ ID NOS: 1059-1410 or 2070-2759 (not including the 3' and 5'restriction sites) (each encoding a VH chain) and a second nucleic acid molecule set forth in SEQ ID NOS: 1411-1422, 1424-1439 and 1441-1471. Such a library can include all permutations of any of the above paired nucleic acid sequences. Thus, the paired library can contain at or about 1.5.times.10.sup.5, 2.1.times.10.sup.5, 2.5.times.10.sup.5, 3.5.times.10.sup.5, 4.times.10.sup.5, 4.2.times.10.sup.5, 4.4.times.10.sup.5, 4.6.times.10.sup.5, 4.8.times.10.sup.5, 5.times.10.sup.5, 5.2.times.10.sup.5, 5.4.times.10.sup.5, 5.6.times.105, 5.8.times.10.sup.5, 6.times.10.sup.5, or more members, or a subset thereof such as 500, 600, 700, 800, 900, 10.sup.3, 5.times.10.sup.3, 10.sup.4, 5.times.10.sup.4, 10.sup.5 or more members.
[0541] Exemplary of a paired library is set forth in Table 17, where each row sets forth a different loci of the library. In the Table, SEQ ID NOS for the nucleic acid molecules are set forth as "RS" (containing a heterologous restriction site sequence) and "NO RS" (not containing a heterologous restriction site sequence).
TABLE-US-00017 TABLE 17 Exemplary Nucleic Acid Paired Library RS NO RS RS NO RS SEQ SEQ SEQ SEQ Heavy Chain Name ID NO ID NO Light Chain Name ID NO ID NO 1 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|O12_IGKJ1*01 833 1438 2 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|O12_IGKJ1*01 833 1438 3 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|O12_IGKJ1*01 833 1438 4 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|O12_IGKJ1*01 833 1438 5 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|O12_IGKJ1*01 833 1438 6 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|O12_IGKJ1*01 833 1438 7 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|O12_IGKJ1*01 833 1438 8 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|O12_IGKJ1*01 833 1438 9 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|O12_IGKJ1*01 833 1438 10 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|O12_IGKJ1*01 833 1438 11 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|O12_IGKJ1*01 833 1438 12 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|O12_IGKJ1*01 833 1438 13 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|O12_IGKJ1*01 833 1438 14 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|O12_IGKJ1*01 833 1438 15 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|O12_IGKJ1*01 833 1438 16 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|O12_IGKJ1*01 833 1438 17 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|O12_IGKJ1*01 833 1438 18 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|O12_IGKJ1*01 833 1438 19 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|O12_IGKJ1*01 833 1438 20 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|O12_IGKJ1*01 833 1438 21 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|O12_IGKJ1*01 833 1438 22 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|O12_IGKJ1*01 833 1438 23 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|O12_IGKJ1*01 833 1438 24 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|O12_IGKJ1*01 833 1438 25 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|O12_IGKJ1*01 833 1438 26 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|O12_IGKJ1*01 833 1438 27 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|O12_IGKJ1*01 833 1438 28 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|O12_IGKJ1*01 833 1438 29 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|O12_IGKJ1*01 833 1438 30 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|O12_IGKJ1*01 833 1438 31 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|O12_IGKJ1*01 833 1438 32 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|O12_IGKJ1*01 833 1438 33 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|O12_IGKJ1*01 833 1438 34 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|O12_IGKJ1*01 833 1438 35 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|O12_IGKJ1*01 833 1438 36 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|O12_IGKJ1*01 833 1438 37 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|O12_IGKJ1*01 833 1438 38 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|O12_IGKJ1*01 833 1438 39 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|O12_IGKJ1*01 833 1438 40 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|O12_IGKJ1*01 833 1438 41 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|O12_IGKJ1*01 833 1438 42 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|O12_IGKJ1*01 833 1438 43 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|O12_IGKJ1*01 833 1438 44 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|O12_IGKJ1*01 833 1438 45 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|O12_IGKJ1*01 833 1438 46 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|O12_IGKJ1*01 833 1438 47 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|O12_IGKJ1*01 833 1438 48 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|O12_IGKJ1*01 833 1438 49 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|O12_IGKJ1*01 833 1438 50 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|O12_IGKJ1*01 833 1438 51 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|O12_IGKJ1*01 833 1438 52 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|O12_IGKJ1*01 833 1438 53 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|O12_IGKJ1*01 833 1438 54 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|O12_IGKJ1*01 833 1438 55 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|O12_IGKJ1*01 833 1438 56 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|O12_IGKJ1*01 833 1438 57 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|O12_IGKJ1*01 833 1438 58 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|O12_IGKJ1*01 833 1438 59 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|O12_IGKJ1*01 833 1438 60 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|O12_IGKJ1*01 833 1438 61 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|O12_IGKJ1*01 833 1438 62 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|O12_IGKJ1*01 833 1438 63 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|O12_IGKJ1*01 833 1438 64 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|O12_IGKJ1*01 833 1438 65 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|O12_IGKJ1*01 833 1438 66 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|O12_IGKJ1*01 833 1438 67 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|O12_IGKJ1*01 833 1438 68 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|O12_IGKJ1*01 833 1438 69 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|O12_IGKJ1*01 833 1438 70 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|O12_IGKJ1*01 833 1438 71 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|O12_IGKJ1*01 833 1438 72 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|O12_IGKJ1*01 833 1438 73 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|O12_IGKJ1*01 833 1438 74 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|O12_IGKJ1*01 833 1438 75 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|O12_IGKJ1*01 833 1438 76 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|O12_IGKJ1*01 833 1438 77 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|O12_IGKJ1*01 833 1438 78 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|O12_IGKJ1*01 833 1438 79 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|O12_IGKJ1*01 833 1438 80 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|O12_IGKJ1*01 833 1438 81 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|O12_IGKJ1*01 833 1438 82 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|O12_IGKJ1*01 833 1438 83 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|O12_IGKJ1*01 833 1438 84 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|O12_IGKJ1*01 833 1438 85 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|O12_IGKJ1*01 833 1438 86 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|O12_IGKJ1*01 833 1438 87 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|O12_IGKJ1*01 833 1438 88 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|O12_IGKJ1*01 833 1438 89 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|O12_IGKJ1*01 833 1438 90 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|O12_IGKJ1*01 833 1438 91 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|O12_IGKJ1*01 833 1438 92 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|O12_IGKJ1*01 833 1438 93 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|O12_IGKJ1*01 833 1438 94 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|O12_IGKJ1*01 833 1438 95 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|O12_IGKJ1*01 833 1438 96 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|O12_IGKJ1*01 833 1438 97 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|O18_IGKJ1*01 834 1439 98 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|O18_IGKJ1*01 834 1439 99 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|O18_IGKJ1*01 834 1439 100 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|O18_IGKJ1*01 834 1439 101 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|O18_IGKJ1*01 834 1439 102 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|O18_IGKJ1*01 834 1439 103 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|O18_IGKJ1*01 834 1439 104 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|O18_IGKJ1*01 834 1439 105 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|O18_IGKJ1*01 834 1439 106 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|O18_IGKJ1*01 834 1439 107 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|O18_IGKJ1*01 834 1439 108 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|O18_IGKJ1*01 834 1439 109 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|O18_IGKJ1*01 834 1439 110 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|O18_IGKJ1*01 834 1439 111 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|O18_IGKJ1*01 834 1439 112 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|O18_IGKJ1*01 834 1439 113 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|O18_IGKJ1*01 834 1439 114 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|O18_IGKJ1*01 834 1439 115 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|O18_IGKJ1*01 834 1439 116 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|O18_IGKJ1*01 834 1439 117 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|O18_IGKJ1*01 834 1439 118 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|O18_IGKJ1*01 834 1439 119 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|O18_IGKJ1*01 834 1439 120 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|O18_IGKJ1*01 834 1439 121 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|O18_IGKJ1*01 834 1439 122 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363
gnl|Fabrus|O18_IGKJ1*01 834 1439 123 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|O18_IGKJ1*01 834 1439 124 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|O18_IGKJ1*01 834 1439 125 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|O18_IGKJ1*01 834 1439 126 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|O18_IGKJ1*01 834 1439 127 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|O18_IGKJ1*01 834 1439 128 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|O18_IGKJ1*01 834 1439 129 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|O18_IGKJ1*01 834 1439 130 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|O18_IGKJ1*01 834 1439 131 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|O18_IGKJ1*01 834 1439 132 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|O18_IGKJ1*01 834 1439 133 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|O18_IGKJ1*01 834 1439 134 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|O18_IGKJ1*01 834 1439 135 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|O18_IGKJ1*01 834 1439 136 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|O18_IGKJ1*01 834 1439 137 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|O18_IGKJ1*01 834 1439 138 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|O18_IGKJ1*01 834 1439 139 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|O18_IGKJ1*01 834 1439 140 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|O18_IGKJ1*01 834 1439 141 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|O18_IGKJ1*01 834 1439 142 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|O18_IGKJ1*01 834 1439 143 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|O18_IGKJ1*01 834 1439 144 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|O18_IGKJ1*01 834 1439 145 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|O18_IGKJ1*01 834 1439 146 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|O18_IGKJ1*01 834 1439 147 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|O18_IGKJ1*01 834 1439 148 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|O18_IGKJ1*01 834 1439 149 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|O18_IGKJ1*01 834 1439 150 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|O18_IGKJ1*01 834 1439 151 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|O18_IGKJ1*01 834 1439 152 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|O18_IGKJ1*01 834 1439 153 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|O18_IGKJ1*01 834 1439 154 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|O18_IGKJ1*01 834 1439 155 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|O18_IGKJ1*01 834 1439 156 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|O18_IGKJ1*01 834 1439 157 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|O18_IGKJ1*01 834 1439 158 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|O18_IGKJ1*01 834 1439 159 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|O18_IGKJ1*01 834 1439 160 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|O18_IGKJ1*01 834 1439 161 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|O18_IGKJ1*01 834 1439 162 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|O18_IGKJ1*01 834 1439 163 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|O18_IGKJ1*01 834 1439 164 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|O18_IGKJ1*01 834 1439 165 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|O18_IGKJ1*01 834 1439 166 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|O18_IGKJ1*01 834 1439 167 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|O18_IGKJ1*01 834 1439 168 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|O18_IGKJ1*01 834 1439 169 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|O18_IGKJ1*01 834 1439 170 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|O18_IGKJ1*01 834 1439 171 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|O18_IGKJ1*01 834 1439 172 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|O18_IGKJ1*01 834 1439 173 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|O18_IGKJ1*01 834 1439 174 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|O18_IGKJ1*01 834 1439 175 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|O18_IGKJ1*01 834 1439 176 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|O18_IGKJ1*01 834 1439 177 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|O18_IGKJ1*01 834 1439 178 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|O18_IGKJ1*01 834 1439 179 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|O18_IGKJ1*01 834 1439 180 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|O18_IGKJ1*01 834 1439 181 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|O18_IGKJ1*01 834 1439 182 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|O18_IGKJ1*01 834 1439 183 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|O18_IGKJ1*01 834 1439 184 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|O18_IGKJ1*01 834 1439 185 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|O18_IGKJ1*01 834 1439 186 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|O18_IGKJ1*01 834 1439 187 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|O18_IGKJ1*01 834 1439 188 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|O18_IGKJ1*01 834 1439 189 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|O18_IGKJ1*01 834 1439 190 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|O18_IGKJ1*01 834 1439 191 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|O18_IGKJ1*01 834 1439 192 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|O18_IGKJ1*01 834 1439 193 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|A20_IGKJ1*01 809 1414 194 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|A20_IGKJ1*01 809 1414 195 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|A20_IGKJ1*01 809 1414 196 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|A20_IGKJ1*01 809 1414 197 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|A20_IGKJ1*01 809 1414 198 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|A20_IGKJ1*01 809 1414 199 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|A20_IGKJ1*01 809 1414 200 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|A20_IGKJ1*01 809 1414 201 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|A20_IGKJ1*01 809 1414 202 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|A20_IGKJ1*01 809 1414 203 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|A20_IGKJ1*01 809 1414 204 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|A20_IGKJ1*01 809 1414 205 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|A20_IGKJ1*01 809 1414 206 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|A20_IGKJ1*01 809 1414 207 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|A20_IGKJ1*01 809 1414 208 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|A20_IGKJ1*01 809 1414 209 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|A20_IGKJ1*01 809 1414 210 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|A20_IGKJ1*01 809 1414 211 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|A20_IGKJ1*01 809 1414 212 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|A20_IGKJ1*01 809 1414 213 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|A20_IGKJ1*01 809 1414 214 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|A20_IGKJ1*01 809 1414 215 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|A20_IGKJ1*01 809 1414 216 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|A20_IGKJ1*01 809 1414 217 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|A20_IGKJ1*01 809 1414 218 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|A20_IGKJ1*01 809 1414 219 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|A20_IGKJ1*01 809 1414 220 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|A20_IGKJ1*01 809 1414 221 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|A20_IGKJ1*01 809 1414 222 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|A20_IGKJ1*01 809 1414 223 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|A20_IGKJ1*01 809 1414 224 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|A20_IGKJ1*01 809 1414 225 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|A20_IGKJ1*01 809 1414 226 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|A20_IGKJ1*01 809 1414 227 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|A20_IGKJ1*01 809 1414 228 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|A20_IGKJ1*01 809 1414 229 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|A20_IGKJ1*01 809 1414 230 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|A20_IGKJ1*01 809 1414 231 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|A20_IGKJ1*01 809 1414 232 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|A20_IGKJ1*01 809 1414 233 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|A20_IGKJ1*01 809 1414 234 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|A20_IGKJ1*01 809 1414 235 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|A20_IGKJ1*01 809 1414 236 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|A20_IGKJ1*01 809 1414 237 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|A20_IGKJ1*01 809 1414 238 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|A20_IGKJ1*01 809 1414 239 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|A20_IGKJ1*01 809 1414 240 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|A20_IGKJ1*01 809 1414 241 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|A20_IGKJ1*01 809 1414 242 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|A20_IGKJ1*01 809 1414 243 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|A20_IGKJ1*01 809 1414 244 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|A20_IGKJ1*01 809 1414 245 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|A20_IGKJ1*01 809 1414 246 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|A20_IGKJ1*01 809 1414 247 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|A20_IGKJ1*01 809 1414
248 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|A20_IGKJ1*01 809 1414 249 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|A20_IGKJ1*01 809 1414 250 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|A20_IGKJ1*01 809 1414 251 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|A20_IGKJ1*01 809 1414 252 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|A20_IGKJ1*01 809 1414 253 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|A20_IGKJ1*01 809 1414 254 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|A20_IGKJ1*01 809 1414 255 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|A20_IGKJ1*01 809 1414 256 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|A20_IGKJ1*01 809 1414 257 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|A20_IGKJ1*01 809 1414 258 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|A20_IGKJ1*01 809 1414 259 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|A20_IGKJ1*01 809 1414 260 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|A20_IGKJ1*01 809 1414 261 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|A20_IGKJ1*01 809 1414 262 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|A20_IGKJ1*01 809 1414 263 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|A20_IGKJ1*01 809 1414 264 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|A20_IGKJ1*01 809 1414 265 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|A20_IGKJ1*01 809 1414 266 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|A20_IGKJ1*01 809 1414 267 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|A20_IGKJ1*01 809 1414 268 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|A20_IGKJ1*01 809 1414 269 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|A20_IGKJ1*01 809 1414 270 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|A20_IGKJ1*01 809 1414 271 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|A20_IGKJ1*01 809 1414 272 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|A20_IGKJ1*01 809 1414 273 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|A20_IGKJ1*01 809 1414 274 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|A20_IGKJ1*01 809 1414 275 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|A20_IGKJ1*01 809 1414 276 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|A20_IGKJ1*01 809 1414 277 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|A20_IGKJ1*01 809 1414 278 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|A20_IGKJ1*01 809 1414 279 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|A20_IGKJ1*01 809 1414 280 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|A20_IGKJ1*01 809 1414 281 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|A20_IGKJ1*01 809 1414 282 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|A20_IGKJ1*01 809 1414 283 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|A20_IGKJ1*01 809 1414 284 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|A20_IGKJ1*01 809 1414 285 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|A20_IGKJ1*01 809 1414 286 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|A20_IGKJ1*01 809 1414 287 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|A20_IGKJ1*01 809 1414 288 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|A20_IGKJ1*01 809 1414 289 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|A30_IGKJ1*01 814 1419 290 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|A30_IGKJ1*01 814 1419 291 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|A30_IGKJ1*01 814 1419 292 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|A30_IGKJ1*01 814 1419 293 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|A30_IGKJ1*01 814 1419 294 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|A30_IGKJ1*01 814 1419 295 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|A30_IGKJ1*01 814 1419 296 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|A30_IGKJ1*01 814 1419 297 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|A30_IGKJ1*01 814 1419 298 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|A30_IGKJ1*01 814 1419 299 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|A30_IGKJ1*01 814 1419 300 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|A30_IGKJ1*01 814 1419 301 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|A30_IGKJ1*01 814 1419 302 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|A30_IGKJ1*01 814 1419 303 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|A30_IGKJ1*01 814 1419 304 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|A30_IGKJ1*01 814 1419 305 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|A30_IGKJ1*01 814 1419 306 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|A30_IGKJ1*01 814 1419 307 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|A30_IGKJ1*01 814 1419 308 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|A30_IGKJ1*01 814 1419 309 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|A30_IGKJ1*01 814 1419 310 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|A30_IGKJ1*01 814 1419 311 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|A30_IGKJ1*01 814 1419 312 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|A30_IGKJ1*01 814 1419 313 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|A30_IGKJ1*01 814 1419 314 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|A30_IGKJ1*01 814 1419 315 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|A30_IGKJ1*01 814 1419 316 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|A30_IGKJ1*01 814 1419 317 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|A30_IGKJ1*01 814 1419 318 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|A30_IGKJ1*01 814 1419 319 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|A30_IGKJ1*01 814 1419 320 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|A30_IGKJ1*01 814 1419 321 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|A30_IGKJ1*01 814 1419 322 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|A30_IGKJ1*01 814 1419 323 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|A30_IGKJ1*01 814 1419 324 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|A30_IGKJ1*01 814 1419 325 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|A30_IGKJ1*01 814 1419 326 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|A30_IGKJ1*01 814 1419 327 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|A30_IGKJ1*01 814 1419 328 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|A30_IGKJ1*01 814 1419 329 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|A30_IGKJ1*01 814 1419 330 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|A30_IGKJ1*01 814 1419 331 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|A30_IGKJ1*01 814 1419 332 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|A30_IGKJ1*01 814 1419 333 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|A30_IGKJ1*01 814 1419 334 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|A30_IGKJ1*01 814 1419 335 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|A30_IGKJ1*01 814 1419 336 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|A30_IGKJ1*01 814 1419 337 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|A30_IGKJ1*01 814 1419 338 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|A30_IGKJ1*01 814 1419 339 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|A30_IGKJ1*01 814 1419 340 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|A30_IGKJ1*01 814 1419 341 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|A30_IGKJ1*01 814 1419 342 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|A30_IGKJ1*01 814 1419 343 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|A30_IGKJ1*01 814 1419 344 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|A30_IGKJ1*01 814 1419 345 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|A30_IGKJ1*01 814 1419 346 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|A30_IGKJ1*01 814 1419 347 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|A30_IGKJ1*01 814 1419 348 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|A30_IGKJ1*01 814 1419 349 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|A30_IGKJ1*01 814 1419 350 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|A30_IGKJ1*01 814 1419 351 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|A30_IGKJ1*01 814 1419 352 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|A30_IGKJ1*01 814 1419 353 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|A30_IGKJ1*01 814 1419 354 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|A30_IGKJ1*01 814 1419 355 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|A30_IGKJ1*01 814 1419 356 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|A30_IGKJ1*01 814 1419 357 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|A30_IGKJ1*01 814 1419 358 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|A30_IGKJ1*01 814 1419 359 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|A30_IGKJ1*01 814 1419 360 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|A30_IGKJ1*01 814 1419 361 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|A30_IGKJ1*01 814 1419 362 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|A30_IGKJ1*01 814 1419 363 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|A30_IGKJ1*01 814 1419 364 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|A30_IGKJ1*01 814 1419 365 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|A30_IGKJ1*01 814 1419 366 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|A30_IGKJ1*01 814 1419 367 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|A30_IGKJ1*01 814 1419 368 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|A30_IGKJ1*01 814 1419 369 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|A30_IGKJ1*01 814 1419 370 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|A30_IGKJ1*01 814 1419 371 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|A30_IGKJ1*01 814 1419 372 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|A30_IGKJ1*01 814 1419 373 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140
gnl|Fabrus|A30_IGKJ1*01 814 1419 374 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|A30_IGKJ1*01 814 1419 375 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|A30_IGKJ1*01 814 1419 376 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|A30_IGKJ1*01 814 1419 377 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|A30_IGKJ1*01 814 1419 378 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|A30_IGKJ1*01 814 1419 379 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|A30_IGKJ1*01 814 1419 380 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|A30_IGKJ1*01 814 1419 381 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|A30_IGKJ1*01 814 1419 382 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|A30_IGKJ1*01 814 1419 383 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|A30_IGKJ1*01 814 1419 384 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|A30_IGKJ1*01 814 1419 385 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 386 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 387 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 388 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 389 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 390 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 391 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 392 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 393 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 394 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 395 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 396 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 397 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 398 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 399 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 400 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 401 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 402 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 403 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 404 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 405 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 406 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 407 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 408 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 409 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 410 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 411 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 412 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 413 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 414 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 415 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 416 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 417 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 418 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 419 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 420 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 421 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 422 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 423 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 424 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 425 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 426 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 427 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 428 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 429 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 430 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 431 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 432 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 433 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 434 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 435 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 436 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 437 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 438 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 439 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 440 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 441 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 442 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 443 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 444 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 445 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 446 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 447 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 448 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 449 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 450 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 451 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 452 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 453 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 454 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 455 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 456 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 457 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 458 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 459 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 460 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 461 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 462 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 463 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 464 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 465 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 466 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 467 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 468 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 469 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 470 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 471 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 472 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 473 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 474 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 475 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 476 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 477 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 478 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 479 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 480 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 481 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|L5_IGKJ1*01 828 1433 482 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|L5_IGKJ1*01 828 1433 483 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|L5_IGKJ1*01 828 1433 484 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|L5_IGKJ1*01 828 1433 485 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|L5_IGKJ1*01 828 1433 486 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|L5_IGKJ1*01 828 1433 487 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|L5_IGKJ1*01 828 1433 488 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|L5_IGKJ1*01 828 1433 489 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|L5_IGKJ1*01 828 1433 490 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|L5_IGKJ1*01 828 1433 491 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|L5_IGKJ1*01 828 1433 492 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|L5_IGKJ1*01 828 1433 493 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|L5_IGKJ1*01 828 1433 494 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|L5_IGKJ1*01 828 1433 495 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|L5_IGKJ1*01 828 1433 496 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|L5_IGKJ1*01 828 1433 497 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|L5_IGKJ1*01 828 1433 498 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|L5_IGKJ1*01 828 1433
499 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|L5_IGKJ1*01 828 1433 500 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|L5_IGKJ1*01 828 1433 501 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|L5_IGKJ1*01 828 1433 502 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|L5_IGKJ1*01 828 1433 503 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|L5_IGKJ1*01 828 1433 504 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|L5_IGKJ1*01 828 1433 505 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|L5_IGKJ1*01 828 1433 506 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|L5_IGKJ1*01 828 1433 507 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|L5_IGKJ1*01 828 1433 508 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|L5_IGKJ1*01 828 1433 509 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|L5_IGKJ1*01 828 1433 510 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|L5_IGKJ1*01 828 1433 511 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|L5_IGKJ1*01 828 1433 512 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|L5_IGKJ1*01 828 1433 513 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|L5_IGKJ1*01 828 1433 514 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|L5_IGKJ1*01 828 1433 515 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|L5_IGKJ1*01 828 1433 516 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|L5_IGKJ1*01 828 1433 517 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|L5_IGKJ1*01 828 1433 518 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|L5_IGKJ1*01 828 1433 519 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|L5_IGKJ1*01 828 1433 520 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|L5_IGKJ1*01 828 1433 521 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|L5_IGKJ1*01 828 1433 522 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|L5_IGKJ1*01 828 1433 523 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|L5_IGKJ1*01 828 1433 524 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|L5_IGKJ1*01 828 1433 525 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|L5_IGKJ1*01 828 1433 526 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|L5_IGKJ1*01 828 1433 527 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|L5_IGKJ1*01 828 1433 528 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|L5_IGKJ1*01 828 1433 529 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|L5_IGKJ1*01 828 1433 530 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|L5_IGKJ1*01 828 1433 531 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|L5_IGKJ1*01 828 1433 532 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|L5_IGKJ1*01 828 1433 533 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|L5_IGKJ1*01 828 1433 534 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|L5_IGKJ1*01 828 1433 535 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|L5_IGKJ1*01 828 1433 536 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|L5_IGKJ1*01 828 1433 537 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|L5_IGKJ1*01 828 1433 538 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|L5_IGKJ1*01 828 1433 539 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|L5_IGKJ1*01 828 1433 540 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|L5_IGKJ1*01 828 1433 541 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|L5_IGKJ1*01 828 1433 542 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|L5_IGKJ1*01 828 1433 543 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|L5_IGKJ1*01 828 1433 544 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|L5_IGKJ1*01 828 1433 545 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|L5_IGKJ1*01 828 1433 546 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|L5_IGKJ1*01 828 1433 547 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|L5_IGKJ1*01 828 1433 548 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|L5_IGKJ1*01 828 1433 549 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|L5_IGKJ1*01 828 1433 550 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|L5_IGKJ1*01 828 1433 551 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|L5_IGKJ1*01 828 1433 552 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|L5_IGKJ1*01 828 1433 553 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|L5_IGKJ1*01 828 1433 554 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|L5_IGKJ1*01 828 1433 555 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|L5_IGKJ1*01 828 1433 556 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|L5_IGKJ1*01 828 1433 557 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|L5_IGKJ1*01 828 1433 558 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|L5_IGKJ1*01 828 1433 559 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|L5_IGKJ1*01 828 1433 560 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|L5_IGKJ1*01 828 1433 561 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|L5_IGKJ1*01 828 1433 562 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|L5_IGKJ1*01 828 1433 563 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|L5_IGKJ1*01 828 1433 564 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|L5_IGKJ1*01 828 1433 565 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|L5_IGKJ1*01 828 1433 566 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|L5_IGKJ1*01 828 1433 567 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|L5_IGKJ1*01 828 1433 568 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|L5_IGKJ1*01 828 1433 569 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|L5_IGKJ1*01 828 1433 570 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|L5_IGKJ1*01 828 1433 571 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|L5_IGKJ1*01 828 1433 572 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|L5_IGKJ1*01 828 1433 573 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|L5_IGKJ1*01 828 1433 574 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|L5_IGKJ1*01 828 1433 575 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|L5_IGKJ1*01 828 1433 576 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|L5_IGKJ1*01 828 1433 577 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|L8_IGKJ1*01 830 1435 578 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|L8_IGKJ1*01 830 1435 579 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|L8_IGKJ1*01 830 1435 580 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|L8_IGKJ1*01 830 1435 581 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|L8_IGKJ1*01 830 1435 582 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|L8_IGKJ1*01 830 1435 583 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|L8_IGKJ1*01 830 1435 584 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|L8_IGKJ1*01 830 1435 585 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|L8_IGKJ1*01 830 1435 586 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|L8_IGKJ1*01 830 1435 587 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|L8_IGKJ1*01 830 1435 588 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|L8_IGKJ1*01 830 1435 589 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|L8_IGKJ1*01 830 1435 590 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|L8_IGKJ1*01 830 1435 591 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|L8_IGKJ1*01 830 1435 592 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|L8_IGKJ1*01 830 1435 593 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|L8_IGKJ1*01 830 1435 594 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|L8_IGKJ1*01 830 1435 595 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|L8_IGKJ1*01 830 1435 596 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|L8_IGKJ1*01 830 1435 597 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|L8_IGKJ1*01 830 1435 598 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|L8_IGKJ1*01 830 1435 599 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|L8_IGKJ1*01 830 1435 600 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|L8_IGKJ1*01 830 1435 601 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|L8_IGKJ1*01 830 1435 602 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|L8_IGKJ1*01 830 1435 603 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|L8_IGKJ1*01 830 1435 604 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|L8_IGKJ1*01 830 1435 605 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|L8_IGKJ1*01 830 1435 606 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|L8_IGKJ1*01 830 1435 607 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|L8_IGKJ1*01 830 1435 608 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|L8_IGKJ1*01 830 1435 609 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|L8_IGKJ1*01 830 1435 610 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|L8_IGKJ1*01 830 1435 611 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|L8_IGKJ1*01 830 1435 612 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|L8_IGKJ1*01 830 1435 613 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|L8_IGKJ1*01 830 1435 614 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|L8_IGKJ1*01 830 1435 615 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|L8_IGKJ1*01 830 1435 616 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|L8_IGKJ1*01 830 1435 617 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|L8_IGKJ1*01 830 1435 618 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|L8_IGKJ1*01 830 1435 619 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|L8_IGKJ1*01 830 1435 620 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|L8_IGKJ1*01 830 1435 621 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|L8_IGKJ1*01 830 1435 622 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|L8_IGKJ1*01 830 1435 623 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|L8_IGKJ1*01 830 1435 624 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400
gnl|Fabrus|L8_IGKJ1*01 830 1435 625 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|L8_IGKJ1*01 830 1435 626 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|L8_IGKJ1*01 830 1435 627 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|L8_IGKJ1*01 830 1435 628 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|L8_IGKJ1*01 830 1435 629 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|L8_IGKJ1*01 830 1435 630 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|L8_IGKJ1*01 830 1435 631 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|L8_IGKJ1*01 830 1435 632 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|L8_IGKJ1*01 830 1435 633 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|L8_IGKJ1*01 830 1435 634 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|L8_IGKJ1*01 830 1435 635 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|L8_IGKJ1*01 830 1435 636 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|L8_IGKJ1*01 830 1435 637 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|L8_IGKJ1*01 830 1435 638 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|L8_IGKJ1*01 830 1435 639 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|L8_IGKJ1*01 830 1435 640 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|L8_IGKJ1*01 830 1435 641 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|L8_IGKJ1*01 830 1435 642 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|L8_IGKJ1*01 830 1435 643 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|L8_IGKJ1*01 830 1435 644 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|L8_IGKJ1*01 830 1435 645 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|L8_IGKJ1*01 830 1435 646 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|L8_IGKJ1*01 830 1435 647 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|L8_IGKJ1*01 830 1435 648 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|L8_IGKJ1*01 830 1435 649 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|L8_IGKJ1*01 830 1435 650 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|L8_IGKJ1*01 830 1435 651 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|L8_IGKJ1*01 830 1435 652 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|L8_IGKJ1*01 830 1435 653 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|L8_IGKJ1*01 830 1435 654 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|L8_IGKJ1*01 830 1435 655 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|L8_IGKJ1*01 830 1435 656 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|L8_IGKJ1*01 830 1435 657 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|L8_IGKJ1*01 830 1435 658 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|L8_IGKJ1*01 830 1435 659 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|L8_IGKJ1*01 830 1435 660 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|L8_IGKJ1*01 830 1435 661 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|L8_IGKJ1*01 830 1435 662 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|L8_IGKJ1*01 830 1435 663 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|L8_IGKJ1*01 830 1435 664 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|L8_IGKJ1*01 830 1435 665 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|L8_IGKJ1*01 830 1435 666 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|L8_IGKJ1*01 830 1435 667 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|L8_IGKJ1*01 830 1435 668 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|L8_IGKJ1*01 830 1435 669 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|L8_IGKJ1*01 830 1435 670 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|L8_IGKJ1*01 830 1435 671 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|L8_IGKJ1*01 830 1435 672 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|L8_IGKJ1*01 830 1435 673 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|L11_IGKJ1*01 819 1424 674 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|L11_IGKJ1*01 819 1424 675 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|L11_IGKJ1*01 819 1424 676 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|L11_IGKJ1*01 819 1424 677 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|L11_IGKJ1*01 819 1424 678 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|L11_IGKJ1*01 819 1424 679 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|L11_IGKJ1*01 819 1424 680 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|L11_IGKJ1*01 819 1424 681 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|L11_IGKJ1*01 819 1424 682 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|L11_IGKJ1*01 819 1424 683 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|L11_IGKJ1*01 819 1424 684 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|L11_IGKJ1*01 819 1424 685 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|L11_IGKJ1*01 819 1424 686 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|L11_IGKJ1*01 819 1424 687 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|L11_IGKJ1*01 819 1424 688 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|L11_IGKJ1*01 819 1424 689 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|L11_IGKJ1*01 819 1424 690 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|L11_IGKJ1*01 819 1424 691 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|L11_IGKJ1*01 819 1424 692 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|L11_IGKJ1*01 819 1424 693 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|L11_IGKJ1*01 819 1424 694 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|L11_IGKJ1*01 819 1424 695 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|L11_IGKJ1*01 819 1424 696 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|L11_IGKJ1*01 819 1424 697 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|L11_IGKJ1*01 819 1424 698 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|L11_IGKJ1*01 819 1424 699 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|L11_IGKJ1*01 819 1424 700 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|L11_IGKJ1*01 819 1424 701 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|L11_IGKJ1*01 819 1424 702 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|L11_IGKJ1*01 819 1424 703 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|L11_IGKJ1*01 819 1424 704 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|L11_IGKJ1*01 819 1424 705 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|L11_IGKJ1*01 819 1424 706 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|L11_IGKJ1*01 819 1424 707 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|L11_IGKJ1*01 819 1424 708 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|L11_IGKJ1*01 819 1424 709 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|L11_IGKJ1*01 819 1424 710 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|L11_IGKJ1*01 819 1424 711 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|L11_IGKJ1*01 819 1424 712 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|L11_IGKJ1*01 819 1424 713 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|L11_IGKJ1*01 819 1424 714 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|L11_IGKJ1*01 819 1424 715 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|L11_IGKJ1*01 819 1424 716 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|L11_IGKJ1*01 819 1424 717 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|L11_IGKJ1*01 819 1424 718 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|L11_IGKJ1*01 819 1424 719 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|L11_IGKJ1*01 819 1424 720 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|L11_IGKJ1*01 819 1424 721 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|L11_IGKJ1*01 819 1424 722 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|L11_IGKJ1*01 819 1424 723 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|L11_IGKJ1*01 819 1424 724 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|L11_IGKJ1*01 819 1424 725 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|L11_IGKJ1*01 819 1424 726 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|L11_IGKJ1*01 819 1424 727 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|L11_IGKJ1*01 819 1424 728 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|L11_IGKJ1*01 819 1424 729 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|L11_IGKJ1*01 819 1424 730 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|L11_IGKJ1*01 819 1424 731 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|L11_IGKJ1*01 819 1424 732 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|L11_IGKJ1*01 819 1424 733 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|L11_IGKJ1*01 819 1424 734 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|L11_IGKJ1*01 819 1424 735 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|L11_IGKJ1*01 819 1424 736 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|L11_IGKJ1*01 819 1424 737 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|L11_IGKJ1*01 819 1424 738 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|L11_IGKJ1*01 819 1424 739 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|L11_IGKJ1*01 819 1424 740 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|L11_IGKJ1*01 819 1424 741 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|L11_IGKJ1*01 819 1424 742 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|L11_IGKJ1*01 819 1424 743 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|L11_IGKJ1*01 819 1424 744 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|L11_IGKJ1*01 819 1424 745 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|L11_IGKJ1*01 819 1424 746 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|L11_IGKJ1*01 819 1424 747 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|L11_IGKJ1*01 819 1424 748 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|L11_IGKJ1*01 819 1424 749 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|L11_IGKJ1*01 819 1424
750 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|L11_IGKJ1*01 819 1424 751 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|L11_IGKJ1*01 819 1424 752 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|L11_IGKJ1*01 819 1424 753 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|L11_IGKJ1*01 819 1424 754 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|L11_IGKJ1*01 819 1424 755 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|L11_IGKJ1*01 819 1424 756 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|L11_IGKJ1*01 819 1424 757 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|L11_IGKJ1*01 819 1424 758 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|L11_IGKJ1*01 819 1424 759 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|L11_IGKJ1*01 819 1424 760 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|L11_IGKJ1*01 819 1424 761 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|L11_IGKJ1*01 819 1424 762 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|L11_IGKJ1*01 819 1424 763 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|L11_IGKJ1*01 819 1424 764 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|L11_IGKJ1*01 819 1424 765 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|L11_IGKJ1*01 819 1424 766 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|L11_IGKJ1*01 819 1424 767 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|L11_IGKJ1*01 819 1424 768 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|L11_IGKJ1*01 819 1424 769 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|L12_IGKJ1*01 820 1425 770 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|L12_IGKJ1*01 820 1425 771 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|L12_IGKJ1*01 820 1425 772 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|L12_IGKJ1*01 820 1425 773 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|L12_IGKJ1*01 820 1425 774 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|L12_IGKJ1*01 820 1425 775 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|L12_IGKJ1*01 820 1425 776 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|L12_IGKJ1*01 820 1425 777 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|L12_IGKJ1*01 820 1425 778 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|L12_IGKJ1*01 820 1425 779 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|L12_IGKJ1*01 820 1425 780 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|L12_IGKJ1*01 820 1425 781 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|L12_IGKJ1*01 820 1425 782 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|L12_IGKJ1*01 820 1425 783 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|L12_IGKJ1*01 820 1425 784 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|L12_IGKJ1*01 820 1425 785 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|L12_IGKJ1*01 820 1425 786 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|L12_IGKJ1*01 820 1425 787 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|L12_IGKJ1*01 820 1425 788 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|L12_IGKJ1*01 820 1425 789 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|L12_IGKJ1*01 820 1425 790 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|L12_IGKJ1*01 820 1425 791 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|L12_IGKJ1*01 820 1425 792 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|L12_IGKJ1*01 820 1425 793 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|L12_IGKJ1*01 820 1425 794 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|L12_IGKJ1*01 820 1425 795 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|L12_IGKJ1*01 820 1425 796 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|L12_IGKJ1*01 820 1425 797 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|L12_IGKJ1*01 820 1425 798 gnl|Fabrus|VH2-26_IGHD4-1*01_IGHJ4*01 526 1131 gnl|Fabrus|L12_IGKJ1*01 820 1425 799 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|L12_IGKJ1*01 820 1425 800 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|L12_IGKJ1*01 820 1425 801 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|L12_IGKJ1*01 820 1425 802 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|L12_IGKJ1*01 820 1425 803 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|L12_IGKJ1*01 820 1425 804 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|L12_IGKJ1*01 820 1425 805 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|L12_IGKJ1*01 820 1425 806 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|L12_IGKJ1*01 820 1425 807 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|L12_IGKJ1*01 820 1425 808 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|L12_IGKJ1*01 820 1425 809 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|L12_IGKJ1*01 820 1425 810 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|L12_IGKJ1*01 820 1425 811 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|L12_IGKJ1*01 820 1425 812 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|L12_IGKJ1*01 820 1425 813 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|L12_IGKJ1*01 820 1425 814 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|L12_IGKJ1*01 820 1425 815 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|L12_IGKJ1*01 820 1425 816 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|L12_IGKJ1*01 820 1425 817 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|L12_IGKJ1*01 820 1425 818 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|L12_IGKJ1*01 820 1425 819 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|L12_IGKJ1*01 820 1425 820 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|L12_IGKJ1*01 820 1425 821 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|L12_IGKJ1*01 820 1425 822 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|L12_IGKJ1*01 820 1425 823 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|L12_IGKJ1*01 820 1425 824 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|L12_IGKJ1*01 820 1425 825 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|L12_IGKJ1*01 820 1425 826 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|L12_IGKJ1*01 820 1425 827 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|L12_IGKJ1*01 820 1425 828 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|L12_IGKJ1*01 820 1425 829 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|L12_IGKJ1*01 820 1425 830 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|L12_IGKJ1*01 820 1425 831 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|L12_IGKJ1*01 820 1425 832 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|L12_IGKJ1*01 820 1425 833 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|L12_IGKJ1*01 820 1425 834 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|L12_IGKJ1*01 820 1425 835 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|L12_IGKJ1*01 820 1425 836 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|L12_IGKJ1*01 820 1425 837 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|L12_IGKJ1*01 820 1425 838 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|L12_IGKJ1*01 820 1425 839 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|L12_IGKJ1*01 820 1425 840 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|L12_IGKJ1*01 820 1425 841 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|L12_IGKJ1*01 820 1425 842 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|L12_IGKJ1*01 820 1425 843 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|L12_IGKJ1*01 820 1425 844 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|L12_IGKJ1*01 820 1425 845 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|L12_IGKJ1*01 820 1425 846 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|L12_IGKJ1*01 820 1425 847 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|L12_IGKJ1*01 820 1425 848 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|L12_IGKJ1*01 820 1425 849 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|L12_IGKJ1*01 820 1425 850 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|L12_IGKJ1*01 820 1425 851 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|L12_IGKJ1*01 820 1425 852 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|L12_IGKJ1*01 820 1425 853 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|L12_IGKJ1*01 820 1425 854 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|L12_IGKJ1*01 820 1425 855 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|L12_IGKJ1*01 820 1425 856 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|L12_IGKJ1*01 820 1425 857 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|L12_IGKJ1*01 820 1425 858 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|L12_IGKJ1*01 820 1425 859 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|L12_IGKJ1*01 820 1425 860 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|L12_IGKJ1*01 820 1425 861 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|L12_IGKJ1*01 820 1425 862 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|L12_IGKJ1*01 820 1425 863 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|L12_IGKJ1*01 820 1425 864 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|L12_IGKJ1*01 820 1425 865 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|O1_IGKJ1*01 832 1437 866 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|O1_IGKJ1*01 832 1437 867 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|O1_IGKJ1*01 832 1437 868 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|O1_IGKJ1*01 832 1437 869 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|O1_IGKJ1*01 832 1437 870 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|O1_IGKJ1*01 832 1437 871 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|O1_IGKJ1*01 832 1437 872 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|O1_IGKJ1*01 832 1437 873 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|O1_IGKJ1*01 832 1437 874 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|O1_IGKJ1*01 832 1437 875 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108
gnl|Fabrus|O1_IGKJ1*01 832 1437 876 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|O1_IGKJ1*01 832 1437 877 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|O1_IGKJ1*01 832 1437 878 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|O1_IGKJ1*01 832 1437 879 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|O1_IGKJ1*01 832 1437 880 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|O1_IGKJ1*01 832 1437 881 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|O1_IGKJ1*01 832 1437 882 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|O1_IGKJ1*01 832 1437 883 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|O1_IGKJ1*01 832 1437 884 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|O1_IGKJ1*01 832 1437 885 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|O1_IGKJ1*01 832 1437 886 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|O1_IGKJ1*01 832 1437 887 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|O1_IGKJ1*01 832 1437 888 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|O1_IGKJ1*01 832 1437 889 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|O1_IGKJ1*01 832 1437 890 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|O1_IGKJ1*01 832 1437 891 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|O1_IGKJ1*01 832 1437 892 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|O1_IGKJ1*01 832 1437 893 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|O1_IGKJ1*01 832 1437 894 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|O1_IGKJ1*01 832 1437 895 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|O1_IGKJ1*01 832 1437 896 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|O1_IGKJ1*01 832 1437 897 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|O1_IGKJ1*01 832 1437 898 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|O1_IGKJ1*01 832 1437 899 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|O1_IGKJ1*01 832 1437 900 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|O1_IGKJ1*01 832 1437 901 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|O1_IGKJ1*01 832 1437 902 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|O1_IGKJ1*01 832 1437 903 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|O1_IGKJ1*01 832 1437 904 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|O1_IGKJ1*01 832 1437 905 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|O1_IGKJ1*01 832 1437 906 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|O1_IGKJ1*01 832 1437 907 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|O1_IGKJ1*01 832 1437 908 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|O1_IGKJ1*01 832 1437 909 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|O1_IGKJ1*01 832 1437 910 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|O1_IGKJ1*01 832 1437 911 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|O1_IGKJ1*01 832 1437 912 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|O1_IGKJ1*01 832 1437 913 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|O1_IGKJ1*01 832 1437 914 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|O1_IGKJ1*01 832 1437 915 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|O1_IGKJ1*01 832 1437 916 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|O1_IGKJ1*01 832 1437 917 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|O1_IGKJ1*01 832 1437 918 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|O1_IGKJ1*01 832 1437 919 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|O1_IGKJ1*01 832 1437 920 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|O1_IGKJ1*01 832 1437 921 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|O1_IGKJ1*01 832 1437 922 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|O1_IGKJ1*01 832 1437 923 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|O1_IGKJ1*01 832 1437 924 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|O1_IGKJ1*01 832 1437 925 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|O1_IGKJ1*01 832 1437 926 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|O1_IGKJ1*01 832 1437 927 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|O1_IGKJ1*01 832 1437 928 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|O1_IGKJ1*01 832 1437 929 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|O1_IGKJ1*01 832 1437 930 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|O1_IGKJ1*01 832 1437 931 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|O1_IGKJ1*01 832 1437 932 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|O1_IGKJ1*01 832 1437 933 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|O1_IGKJ1*01 832 1437 934 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|O1_IGKJ1*01 832 1437 935 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|O1_IGKJ1*01 832 1437 936 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|O1_IGKJ1*01 832 1437 937 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|O1_IGKJ1*01 832 1437 938 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|O1_IGKJ1*01 832 1437 939 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|O1_IGKJ1*01 832 1437 940 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|O1_IGKJ1*01 832 1437 941 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|O1_IGKJ1*01 832 1437 942 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|O1_IGKJ1*01 832 1437 943 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|O1_IGKJ1*01 832 1437 944 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|O1_IGKJ1*01 832 1437 945 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|O1_IGKJ1*01 832 1437 946 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|O1_IGKJ1*01 832 1437 947 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|O1_IGKJ1*01 832 1437 948 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|O1_IGKJ1*01 832 1437 949 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|O1_IGKJ1*01 832 1437 950 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|O1_IGKJ1*01 832 1437 951 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|O1_IGKJ1*01 832 1437 952 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|O1_IGKJ1*01 832 1437 953 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|O1_IGKJ1*01 832 1437 954 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|O1_IGKJ1*01 832 1437 955 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|O1_IGKJ1*01 832 1437 956 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|O1_IGKJ1*01 832 1437 957 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|O1_IGKJ1*01 832 1437 958 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|O1_IGKJ1*01 832 1437 959 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|O1_IGKJ1*01 832 1437 960 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|O1_IGKJ1*01 832 1437 961 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|L25_IGKJ3*01 826 1431 962 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|L25_IGKJ3*01 826 1431 963 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|L25_IGKJ3*01 826 1431 964 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|L25_IGKJ3*01 826 1431 965 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|L25_IGKJ3*01 826 1431 966 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|L25_IGKJ3*01 826 1431 967 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|L25_IGKJ3*01 826 1431 968 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|L25_IGKJ3*01 826 1431 969 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|L25_IGKJ3*01 826 1431 970 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|L25_IGKJ3*01 826 1431 971 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|L25_IGKJ3*01 826 1431 972 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|L25_IGKJ3*01 826 1431 973 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|L25_IGKJ3*01 826 1431 974 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|L25_IGKJ3*01 826 1431 975 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|L25_IGKJ3*01 826 1431 976 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|L25_IGKJ3*01 826 1431 977 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|L25_IGKJ3*01 826 1431 978 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|L25_IGKJ3*01 826 1431 979 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|L25_IGKJ3*01 826 1431 980 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|L25_IGKJ3*01 826 1431 981 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|L25_IGKJ3*01 826 1431 982 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|L25_IGKJ3*01 826 1431 983 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|L25_IGKJ3*01 826 1431 984 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|L25_IGKJ3*01 826 1431 985 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|L25_IGKJ3*01 826 1431 986 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|L25_IGKJ3*01 826 1431 987 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|L25_IGKJ3*01 826 1431 988 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|L25_IGKJ3*01 826 1431 989 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|L25_IGKJ3*01 826 1431 990 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|L25_IGKJ3*01 826 1431 991 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|L25_IGKJ3*01 826 1431 992 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|L25_IGKJ3*01 826 1431 993 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|L25_IGKJ3*01 826 1431 994 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|L25_IGKJ3*01 826 1431 995 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|L25_IGKJ3*01 826 1431 996 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|L25_IGKJ3*01 826 1431 997 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|L25_IGKJ3*01 826 1431 998 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|L25_IGKJ3*01 826 1431 999 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|L25_IGKJ3*01 826 1431 1000 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|L25_IGKJ3*01 826 1431
1001 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|L25_IGKJ3*01 826 1431 1002 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|L25_IGKJ3*01 826 1431 1003 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|L25_IGKJ3*01 826 1431 1004 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|L25_IGKJ3*01 826 1431 1005 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|L25_IGKJ3*01 826 1431 1006 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|L25_IGKJ3*01 826 1431 1007 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|L25_IGKJ3*01 826 1431 1008 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|L25_IGKJ3*01 826 1431 1009 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|L25_IGKJ3*01 826 1431 1010 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|L25_IGKJ3*01 826 1431 1011 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|L25_IGKJ3*01 826 1431 1012 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|L25_IGKJ3*01 826 1431 1013 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|L25_IGKJ3*01 826 1431 1014 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|L25_IGKJ3*01 826 1431 1015 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|L25_IGKJ3*01 826 1431 1016 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|L25_IGKJ3*01 826 1431 1017 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|L25_IGKJ3*01 826 1431 1018 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|L25_IGKJ3*01 826 1431 1019 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|L25_IGKJ3*01 826 1431 1020 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|L25_IGKJ3*01 826 1431 1021 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|L25_IGKJ3*01 826 1431 1022 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|L25_IGKJ3*01 826 1431 1023 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|L25_IGKJ3*01 826 1431 1024 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|L25_IGKJ3*01 826 1431 1025 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|L25_IGKJ3*01 826 1431 1026 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|L25_IGKJ3*01 826 1431 1027 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|L25_IGKJ3*01 826 1431 1028 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|L25_IGKJ3*01 826 1431 1029 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|L25_IGKJ3*01 826 1431 1030 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|L25_IGKJ3*01 826 1431 1031 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|L25_IGKJ3*01 826 1431 1032 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|L25_IGKJ3*01 826 1431 1033 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|L25_IGKJ3*01 826 1431 1034 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|L25_IGKJ3*01 826 1431 1035 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|L25_IGKJ3*01 826 1431 1036 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|L25_IGKJ3*01 826 1431 1037 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|L25_IGKJ3*01 826 1431 1038 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|L25_IGKJ3*01 826 1431 1039 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|L25_IGKJ3*01 826 1431 1040 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|L25_IGKJ3*01 826 1431 1041 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|L25_IGKJ3*01 826 1431 1042 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|L25_IGKJ3*01 826 1431 1043 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|L25_IGKJ3*01 826 1431 1044 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|L25_IGKJ3*01 826 1431 1045 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|L25_IGKJ3*01 826 1431 1046 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|L25_IGKJ3*01 826 1431 1047 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|L25_IGKJ3*01 826 1431 1048 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|L25_IGKJ3*01 826 1431 1049 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|L25_IGKJ3*01 826 1431 1050 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|L25_IGKJ3*01 826 1431 1051 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|L25_IGKJ3*01 826 1431 1052 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|L25_IGKJ3*01 826 1431 1053 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|L25_IGKJ3*01 826 1431 1054 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|L25_IGKJ3*01 826 1431 1055 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|L25_IGKJ3*01 826 1431 1056 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|L25_IGKJ3*01 826 1431 1057 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|A27_IGKJ1*01 812 1417 1058 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|A27_IGKJ1*01 812 1417 1059 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|A27_IGKJ1*01 812 1417 1060 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|A27_IGKJ1*01 812 1417 1061 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|A27_IGKJ1*01 812 1417 1062 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|A27_IGKJ1*01 812 1417 1063 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|A27_IGKJ1*01 812 1417 1064 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|A27_IGKJ1*01 812 1417 1065 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|A27_IGKJ1*01 812 1417 1066 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|A27_IGKJ1*01 812 1417 1067 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|A27_IGKJ1*01 812 1417 1068 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|A27_IGKJ1*01 812 1417 1069 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|A27_IGKJ1*01 812 1417 1070 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|A27_IGKJ1*01 812 1417 1071 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|A27_IGKJ1*01 812 1417 1072 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|A27_IGKJ1*01 812 1417 1073 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|A27_IGKJ1*01 812 1417 1074 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|A27_IGKJ1*01 812 1417 1075 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|A27_IGKJ1*01 812 1417 1076 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|A27_IGKJ1*01 812 1417 1077 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|A27_IGKJ1*01 812 1417 1078 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|A27_IGKJ1*01 812 1417 1079 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|A27_IGKJ1*01 812 1417 1080 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|A27_IGKJ1*01 812 1417 1081 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|A27_IGKJ1*01 812 1417 1082 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|A27_IGKJ1*01 812 1417 1083 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|A27_IGKJ1*01 812 1417 1084 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|A27_IGKJ1*01 812 1417 1085 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|A27_IGKJ1*01 812 1417 1086 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|A27_IGKJ1*01 812 1417 1087 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|A27_IGKJ1*01 812 1417 1088 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|A27_IGKJ1*01 812 1417 1089 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|A27_IGKJ1*01 812 1417 1090 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|A27_IGKJ1*01 812 1417 1091 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|A27_IGKJ1*01 812 1417 1092 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|A27_IGKJ1*01 812 1417 1093 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|A27_IGKJ1*01 812 1417 1094 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|A27_IGKJ1*01 812 1417 1095 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|A27_IGKJ1*01 812 1417 1096 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|A27_IGKJ1*01 812 1417 1097 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|A27_IGKJ1*01 812 1417 1098 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|A27_IGKJ1*01 812 1417 1099 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|A27_IGKJ1*01 812 1417 1100 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|A27_IGKJ1*01 812 1417 1101 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|A27_IGKJ1*01 812 1417 1102 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|A27_IGKJ1*01 812 1417 1103 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|A27_IGKJ1*01 812 1417 1104 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|A27_IGKJ1*01 812 1417 1105 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|A27_IGKJ1*01 812 1417 1106 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|A27_IGKJ1*01 812 1417 1107 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|A27_IGKJ1*01 812 1417 1108 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|A27_IGKJ1*01 812 1417 1109 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|A27_IGKJ1*01 812 1417 1110 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|A27_IGKJ1*01 812 1417 1111 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|A27_IGKJ1*01 812 1417 1112 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|A27_IGKJ1*01 812 1417 1113 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|A27_IGKJ1*01 812 1417 1114 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|A27_IGKJ1*01 812 1417 1115 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|A27_IGKJ1*01 812 1417 1116 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|A27_IGKJ1*01 812 1417 1117 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|A27_IGKJ1*01 812 1417 1118 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|A27_IGKJ1*01 812 1417 1119 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|A27_IGKJ1*01 812 1417 1120 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|A27_IGKJ1*01 812 1417 1121 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|A27_IGKJ1*01 812 1417 1122 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|A27_IGKJ1*01 812 1417 1123 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|A27_IGKJ1*01 812 1417 1124 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|A27_IGKJ1*01 812 1417 1125 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|A27_IGKJ1*01 812 1417 1126 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332
gnl|Fabrus|A27_IGKJ1*01 812 1417 1127 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|A27_IGKJ1*01 812 1417 1128 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|A27_IGKJ1*01 812 1417 1129 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|A27_IGKJ1*01 812 1417 1130 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|A27_IGKJ1*01 812 1417 1131 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|A27_IGKJ1*01 812 1417 1132 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|A27_IGKJ1*01 812 1417 1133 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|A27_IGKJ1*01 812 1417 1134 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|A27_IGKJ1*01 812 1417 1135 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|A27_IGKJ1*01 812 1417 1136 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|A27_IGKJ1*01 812 1417 1137 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|A27_IGKJ1*01 812 1417 1138 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|A27_IGKJ1*01 812 1417 1139 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|A27_IGKJ1*01 812 1417 1140 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|A27_IGKJ1*01 812 1417 1141 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|A27_IGKJ1*01 812 1417 1142 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|A27_IGKJ1*01 812 1417 1143 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|A27_IGKJ1*01 812 1417 1144 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|A27_IGKJ1*01 812 1417 1145 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|A27_IGKJ1*01 812 1417 1146 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|A27_IGKJ1*01 812 1417 1147 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|A27_IGKJ1*01 812 1417 1148 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|A27_IGKJ1*01 812 1417 1149 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|A27_IGKJ1*01 812 1417 1150 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|A27_IGKJ1*01 812 1417 1151 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|A27_IGKJ1*01 812 1417 1152 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|A27_IGKJ1*01 812 1417 1153 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|A2_IGKJ1*01 808 1413 1154 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|A2_IGKJ1*01 808 1413 1155 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|A2_IGKJ1*01 808 1413 1156 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|A2_IGKJ1*01 808 1413 1157 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|A2_IGKJ1*01 808 1413 1158 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|A2_IGKJ1*01 808 1413 1159 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|A2_IGKJ1*01 808 1413 1160 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|A2_IGKJ1*01 808 1413 1161 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|A2_IGKJ1*01 808 1413 1162 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|A2_IGKJ1*01 808 1413 1163 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|A2_IGKJ1*01 808 1413 1164 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|A2_IGKJ1*01 808 1413 1165 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|A2_IGKJ1*01 808 1413 1166 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|A2_IGKJ1*01 808 1413 1167 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|A2_IGKJ1*01 808 1413 1168 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|A2_IGKJ1*01 808 1413 1169 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|A2_IGKJ1*01 808 1413 1170 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|A2_IGKJ1*01 808 1413 1171 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|A2_IGKJ1*01 808 1413 1172 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|A2_IGKJ1*01 808 1413 1173 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|A2_IGKJ1*01 808 1413 1174 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|A2_IGKJ1*01 808 1413 1175 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|A2_IGKJ1*01 808 1413 1176 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|A2_IGKJ1*01 808 1413 1177 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|A2_IGKJ1*01 808 1413 1178 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|A2_IGKJ1*01 808 1413 1179 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|A2_IGKJ1*01 808 1413 1180 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|A2_IGKJ1*01 808 1413 1181 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|A2_IGKJ1*01 808 1413 1182 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|A2_IGKJ1*01 808 1413 1183 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|A2_IGKJ1*01 808 1413 1184 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|A2_IGKJ1*01 808 1413 1185 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|A2_IGKJ1*01 808 1413 1186 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|A2_IGKJ1*01 808 1413 1187 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|A2_IGKJ1*01 808 1413 1188 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|A2_IGKJ1*01 808 1413 1189 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|A2_IGKJ1*01 808 1413 1190 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|A2_IGKJ1*01 808 1413 1191 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|A2_IGKJ1*01 808 1413 1192 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|A2_IGKJ1*01 808 1413 1193 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|A2_IGKJ1*01 808 1413 1194 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|A2_IGKJ1*01 808 1413 1195 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|A2_IGKJ1*01 808 1413 1196 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|A2_IGKJ1*01 808 1413 1197 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|A2_IGKJ1*01 808 1413 1198 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|A2_IGKJ1*01 808 1413 1199 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|A2_IGKJ1*01 808 1413 1200 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|A2_IGKJ1*01 808 1413 1201 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|A2_IGKJ1*01 808 1413 1202 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|A2_IGKJ1*01 808 1413 1203 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|A2_IGKJ1*01 808 1413 1204 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|A2_IGKJ1*01 808 1413 1205 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|A2_IGKJ1*01 808 1413 1206 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|A2_IGKJ1*01 808 1413 1207 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|A2_IGKJ1*01 808 1413 1208 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|A2_IGKJ1*01 808 1413 1209 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|A2_IGKJ1*01 808 1413 1210 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|A2_IGKJ1*01 808 1413 1211 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|A2_IGKJ1*01 808 1413 1212 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|A2_IGKJ1*01 808 1413 1213 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|A2_IGKJ1*01 808 1413 1214 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|A2_IGKJ1*01 808 1413 1215 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|A2_IGKJ1*01 808 1413 1216 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|A2_IGKJ1*01 808 1413 1217 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|A2_IGKJ1*01 808 1413 1218 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|A2_IGKJ1*01 808 1413 1219 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|A2_IGKJ1*01 808 1413 1220 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|A2_IGKJ1*01 808 1413 1221 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|A2_IGKJ1*01 808 1413 1222 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|A2_IGKJ1*01 808 1413 1223 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|A2_IGKJ1*01 808 1413 1224 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|A2_IGKJ1*01 808 1413 1225 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|A2_IGKJ1*01 808 1413 1226 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|A2_IGKJ1*01 808 1413 1227 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|A2_IGKJ1*01 808 1413 1228 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|A2_IGKJ1*01 808 1413 1229 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|A2_IGKJ1*01 808 1413 1230 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|A2_IGKJ1*01 808 1413 1231 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|A2_IGKJ1*01 808 1413 1232 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|A2_IGKJ1*01 808 1413 1233 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|A2_IGKJ1*01 808 1413 1234 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|A2_IGKJ1*01 808 1413 1235 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|A2_1GKJ1*01 808 1413 1236 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|A2_1GKJ1*01 808 1413 1237 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|A2_1GKJ1*01 808 1413 1238 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|A2_1GKJ1*01 808 1413 1239 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|A2_1GKJ1*01 808 1413 1240 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|A2_1GKJ1*01 808 1413 1241 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|A2_1GKJ1*01 808 1413 1242 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|A2_1GKJ1*01 808 1413 1243 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|A2_1GKJ1*01 808 1413 1244 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|A2_1GKJ1*01 808 1413 1245 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|A2_1GKJ1*01 808 1413 1246 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|A2_1GKJ1*01 808 1413 1247 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|A2_1GKJ1*01 808 1413 1248 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|A2_1GKJ1*01 808 1413 1249 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 1200 gnl|Fabrus|HerceptinLC 818 1423 1250 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 1203 gnl|Fabrus|HerceptinLC 818 1423 1251 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 1207 gnl|Fabrus|HerceptinLC 818 1423
1252 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 1209 gnl|Fabrus|HerceptinLC 818 1423 1253 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 1211 gnl|Fabrus|HerceptinLC 818 1423 1254 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 1213 gnl|Fabrus|HerceptinLC 818 1423 1255 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 1214 gnl|Fabrus|HerceptinLC 818 1423 1256 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 1217 gnl|Fabrus|HerceptinLC 818 1423 1257 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 1218 gnl|Fabrus|HerceptinLC 818 1423 1258 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 1107 gnl|Fabrus|HerceptinLC 818 1423 1259 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 1108 gnl|Fabrus|HerceptinLC 818 1423 1260 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 1109 gnl|Fabrus|HerceptinLC 818 1423 1261 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 1110 gnl|Fabrus|HerceptinLC 818 1423 1262 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 1111 gnl|Fabrus|HerceptinLC 818 1423 1263 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 1113 gnl|Fabrus|HerceptinLC 818 1423 1264 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 1114 gnl|Fabrus|HerceptinLC 818 1423 1265 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 1116 gnl|Fabrus|HerceptinLC 818 1423 1266 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 1118 gnl|Fabrus|HerceptinLC 818 1423 1267 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 1354 gnl|Fabrus|HerceptinLC 818 1423 1268 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 1355 gnl|Fabrus|HerceptinLC 818 1423 1269 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 1356 gnl|Fabrus|HerceptinLC 818 1423 1270 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 1358 gnl|Fabrus|HerceptinLC 818 1423 1271 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 1359 gnl|Fabrus|HerceptinLC 818 1423 1272 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 1360 gnl|Fabrus|HerceptinLC 818 1423 1273 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 1361 gnl|Fabrus|HerceptinLC 818 1423 1274 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 1363 gnl|Fabrus|HerceptinLC 818 1423 1275 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 1126 gnl|Fabrus|HerceptinLC 818 1423 1276 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 1128 gnl|Fabrus|HerceptinLC 818 1423 1277 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 1129 gnl|Fabrus|HerceptinLC 818 1423 1278 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 1131 gnl|Fabrus|HerceptinLC 818 1423 1279 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 1133 gnl|Fabrus|HerceptinLC 818 1423 1280 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 1134 gnl|Fabrus|HerceptinLC 818 1423 1281 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 1135 gnl|Fabrus|HerceptinLC 818 1423 1282 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 1381 gnl|Fabrus|HerceptinLC 818 1423 1283 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 1383 gnl|Fabrus|HerceptinLC 818 1423 1284 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 1385 gnl|Fabrus|HerceptinLC 818 1423 1285 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 1386 gnl|Fabrus|HerceptinLC 818 1423 1286 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 1387 gnl|Fabrus|HerceptinLC 818 1423 1287 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 1388 gnl|Fabrus|HerceptinLC 818 1423 1288 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 1389 gnl|Fabrus|HerceptinLC 818 1423 1289 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 1390 gnl|Fabrus|HerceptinLC 818 1423 1290 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 1391 gnl|Fabrus|HerceptinLC 818 1423 1291 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 1393 gnl|Fabrus|HerceptinLC 818 1423 1292 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 1396 gnl|Fabrus|HerceptinLC 818 1423 1293 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 1398 gnl|Fabrus|HerceptinLC 818 1423 1294 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 1397 gnl|Fabrus|HerceptinLC 818 1423 1295 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 1399 gnl|Fabrus|HerceptinLC 818 1423 1296 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 1400 gnl|Fabrus|HerceptinLC 818 1423 1297 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 1401 gnl|Fabrus|HerceptinLC 818 1423 1298 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 1402 gnl|Fabrus|HerceptinLC 818 1423 1299 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 1380 gnl|Fabrus|HerceptinLC 818 1423 1300 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 1260 gnl|Fabrus|HerceptinLC 818 1423 1301 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 1230 gnl|Fabrus|HerceptinLC 818 1423 1302 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 1286 gnl|Fabrus|HerceptinLC 818 1423 1303 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 1275 gnl|Fabrus|HerceptinLC 818 1423 1304 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 1141 gnl|Fabrus|HerceptinLC 818 1423 1305 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 1148 gnl|Fabrus|HerceptinLC 818 1423 1306 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 1172 gnl|Fabrus|HerceptinLC 818 1423 1307 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 1170 gnl|Fabrus|HerceptinLC 818 1423 1308 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 1267 gnl|Fabrus|HerceptinLC 818 1423 1309 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 1268 gnl|Fabrus|HerceptinLC 818 1423 1310 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 1304 gnl|Fabrus|HerceptinLC 818 1423 1311 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 1306 gnl|Fabrus|HerceptinLC 818 1423 1312 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 1314 gnl|Fabrus|HerceptinLC 818 1423 1313 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 1313 gnl|Fabrus|HerceptinLC 818 1423 1314 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 1255 gnl|Fabrus|HerceptinLC 818 1423 1315 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 1258 gnl|Fabrus|HerceptinLC 818 1423 1316 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 1329 gnl|Fabrus|HerceptinLC 818 1423 1317 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 1326 gnl|Fabrus|HerceptinLC 818 1423 1318 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 1332 gnl|Fabrus|HerceptinLC 818 1423 1319 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 1367 gnl|Fabrus|HerceptinLC 818 1423 1320 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 1371 gnl|Fabrus|HerceptinLC 818 1423 1321 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 1065 gnl|Fabrus|HerceptinLC 818 1423 1322 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 1072 gnl|Fabrus|HerceptinLC 818 1423 1323 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 1066 gnl|Fabrus|HerceptinLC 818 1423 1324 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 1080 gnl|Fabrus|HerceptinLC 818 1423 1325 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 1085 gnl|Fabrus|HerceptinLC 818 1423 1326 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 1091 gnl|Fabrus|HerceptinLC 818 1423 1327 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 1405 gnl|Fabrus|HerceptinLC 818 1423 1328 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 1147 gnl|Fabrus|HerceptinLC 818 1423 1329 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 1101 gnl|Fabrus|HerceptinLC 818 1423 1330 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 1404 gnl|Fabrus|HerceptinLC 818 1423 1331 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 1339 gnl|Fabrus|HerceptinLC 818 1423 1332 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 1345 gnl|Fabrus|HerceptinLC 818 1423 1333 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 1140 gnl|Fabrus|HerceptinLC 818 1423 1334 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 1120 gnl|Fabrus|HerceptinLC 818 1423 1335 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 1145 gnl|Fabrus|HerceptinLC 818 1423 1336 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 1244 gnl|Fabrus|HerceptinLC 818 1423 1337 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 1175 gnl|Fabrus|HerceptinLC 818 1423 1338 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 1311 gnl|Fabrus|HerceptinLC 818 1423 1339 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 1153 gnl|Fabrus|HerceptinLC 818 1423 1340 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 1157 gnl|Fabrus|HerceptinLC 818 1423 1341 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 1189 gnl|Fabrus|HerceptinLC 818 1423 1342 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 1176 gnl|Fabrus|HerceptinLC 818 1423 1343 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 1297 gnl|Fabrus|HerceptinLC 818 1423 1344 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 1181 gnl|Fabrus|HerceptinLC 818 1423 1345 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|O12_IGKJ1*01 833 1438 1346 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|O18_IGKJ1*01 834 1439 1347 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A20_IGKJ1*01 809 1414 1348 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A30_IGKJ1*01 814 1419 1349 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L14_IGKJ1*01 821 1426 1350 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 1351 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L5_IGKJ1*01 828 1433 1352 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L6_IGKJ1*01 830 1435 1353 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L23_IGKJ1*01 824 1429 1354 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L11_IGKJ1*01 819 1424 1355 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L12_IGKJ1*01 820 1425 1356 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|O1_IGKJ1*01 832 1437 1357 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A17_IGKJ1*01 807 1412 1358 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A2_IGKJ1*01 808 1413 1359 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A23_IGKJ1*01 810 1415 1360 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A27_IGKJ3*01 813 1418 1361 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L2_IGKJ1*01 822 1427 1362 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L6_IGKJ1*01 829 1434 1363 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L25_IGKJ1*01 825 1430 1364 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|B3_IGKJ1*01 817 1422 1365 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|B2_IGKJ1*01 815 1420 1366 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A26_IGKJ1*01 811 1416 1367 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A14_IGKJ1*01 806 1411 1368 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L9_IGKJ2*01 831 1436 1369 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|A27_IGKJ1*01 812 1417 1370 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|B2_IGKJ3*01 816 1421 1371 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L25_IGKJ3*01 826 1431 1372 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|RituxanLC 835 1440 1373 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|L22_IGKJ3*01 823 1428 1374 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 1205 gnl|Fabrus|HerceptinLC 818 1423 1375 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|O12_IGKJ1*01 833 1438 1376 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|O18_IGKJ1*01 834 1439 1377 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352
gnl|Fabrus|A20_IGKJ1*01 809 1414 1378 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A30_IGKJ1*01 814 1419 1379 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L14_IGKJ1*01 821 1426 1380 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 1381 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L5_IGKJ1*01 828 1433 1382 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L8_IGKJ1*01 830 1435 1383 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L23_IGKJ1*01 824 1429 1384 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L11_IGKJ1*01 819 1424 1385 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L12_IGKJ1*01 820 1425 1386 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|O1_IGKJ1*01 832 1437 1387 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A17_IGKJ1*01 807 1412 1388 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A2_IGKJ1*01 808 1413 1389 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A23_IGKJ1*01 810 1415 1390 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A27_IGKJ3*01 813 1418 1391 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L2_IGKJ1*01 822 1427 1392 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L6_IGKJ1*01 829 1434 1393 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L25_IGKJ1*01 825 1430 1394 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|B3_IGKJ1*01 817 1422 1395 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|B2_IGKJ1*01 815 1420 1396 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A26_IGKJ1*01 811 1416 1397 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A14_IGKJ1*01 806 1411 1398 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L9_IGKJ2*01 831 1436 1399 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|A27_IGKJ1*01 812 1417 1400 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|B2_IGKJ3*01 816 1421 1401 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L25_IGKJ3*01 826 1431 1402 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|RituxanLC 835 1440 1403 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|L22_IGKJ3*01 823 1428 1404 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 1352 gnl|Fabrus|HerceptinLC 818 1423 1405 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|O12_IGKJ1*01 833 1438 1406 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|O18_IGKJ1*01 834 1439 1407 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A20_IGKJ1*01 809 1414 1408 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A30_IGKJ1*01 814 1419 1409 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L14_IGKJ1*01 821 1426 1410 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 1411 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L5_IGKJ1*01 828 1433 1412 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L8_IGKJ1*01 830 1435 1413 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L23_IGKJ1*01 824 1429 1414 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L11_IGKJ1*01 819 1424 1415 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L12_IGKJ1*01 820 1425 1416 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|O1_IGKJ1*01 832 1437 1417 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A17_IGKJ1*01 807 1412 1418 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A2_IGKJ1*01 808 1413 1419 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A23_IGKJ1*01 810 1415 1420 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A27_IGKJ3*01 813 1418 1421 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L2_IGKJ1*01 822 1427 1422 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L6_IGKJ1*01 829 1434 1423 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L25_IGKJ1*01 825 1430 1424 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|B3_IGKJ1*01 817 1422 1425 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|B2_IGKJ1*01 815 1420 1426 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A26_IGKJ1*01 811 1416 1427 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A14_IGKJ1*01 806 1411 1428 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L9_IGKJ2*01 831 1436 1429 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|A27_IGKJ1*01 812 1417 1430 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|B2_IGKJ3*01 816 1421 1431 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L25_IGKJ3*01 826 1431 1432 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|RituxanLC 835 1440 1433 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|L22_IGKJ3*01 823 1428 1434 gnl|Fabrus|RituxanHC 453 1058 gnl|Fabrus|HerceptinLC 818 1423 1435 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|O12_IGKJ1*01 833 1438 1436 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|O18_IGKJ1*01 834 1439 1437 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A20_IGKJ1*01 809 1414 1438 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A30_IGKJ1*01 814 1419 1439 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L14_IGKJ1*01 821 1426 1440 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L4/18a_IGKJ1*01 827 1432 1441 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L5_IGKJ1*01 828 1433 1442 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L8_IGKJ1*01 830 1435 1443 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L23_IGKJ1*01 824 1429 1444 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L11_IGKJ1*01 819 1424 1445 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L12_IGKJ1*01 820 1425 1446 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|O1_IGKJ1*01 832 1437 1447 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A17_IGKJ1*01 807 1412 1448 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A2_IGKJ1*01 808 1413 1449 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A23_IGKJ1*01 810 1415 1450 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A27_IGKJ3*01 813 1418 1451 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L2_IGKJ1*01 822 1427 1452 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L6_IGKJ1*01 829 1434 1453 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L25_IGKJ1*01 825 1430 1454 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|B3_IGKJ1*01 817 1422 1455 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|B2_IGKJ1*01 815 1420 1456 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A26_IGKJ1*01 811 1416 1457 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A14_IGKJ1*01 806 1411 1458 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L9_IGKJ2*01 831 1436 1459 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|A27_IGKJ1*01 812 1417 1460 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|B32_IGKJ3*01 816 1421 1461 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L25_IGKJ3*01 826 1431 1462 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|RituxanLC 835 1440 1463 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|L22_IGKJ3*01 823 1428 1464 gnl|Fabrus|HerceptinHC 452 1057 gnl|Fabrus|HerceptinLC 818 1423
[0542] An additional exemplary paired library is set forth in Table 17A, where each row sets forth a different loci of the library.
TABLE-US-00018 TABLE 17A Exemplary Nucleic Acid Paired Library RS RS NO RS HEAVY CHAIN SEQ LIGHT CHAIN SEQ SEQ Name ID NO Name ID NO ID NO 1 VH3-23_IGHD1-1*01 > 1_IGHJ1*01 2070 O12_IGKJ1*01 833 1438 2 VH3-23_IGHD1-1*01 > 2_IGHJ1*01 2071 O12_IGKJ1*01 833 1438 3 VH3-23_IGHD1-1*01 > 3_IGHJ1*01 2072 O12_IGKJ1*01 833 1438 4 VH3-23_IGHD1-7*01 > 1_IGHJ1*01 2073 O12_IGKJ1*01 833 1438 5 VH3-23_IGHD1-7*01 > 3_IGHJ1*01 2074 O12_IGKJ1*01 833 1438 6 VH3-23_IGHD1-14*01 > 1_IGHJ1*01 2075 O12_IGKJ1*01 833 1438 7 VH3-23_IGHD1-14*01 > 3_IGHJ1*01 2076 O12_IGKJ1*01 833 1438 8 VH3-23_IGHD1-20*01 > 1_IGHJ1*01 2077 O12_IGKJ1*01 833 1438 9 VH3-23_IGHD1-20*01 > 3_IGHJ1*01 2078 O12_IGKJ1*01 833 1438 10 VH3-23_IGHD1-26*01 > 1_IGHJ1*01 2079 O12_IGKJ1*01 833 1438 11 VH3-23_IGHD1-26*01 > 3_IGHJ1*01 2080 O12_IGKJ1*01 833 1438 12 VH3-23_IGHD2-2*01 > 2_IGHJ1*01 2081 O12_IGKJ1*01 833 1438 13 VH3-23_IGHD2-2*01 > 3_IGHJ1*01 2082 O12_IGKJ1*01 833 1438 14 VH3-23_IGHD2-8*01 > 2_IGHJ1*01 2083 O12_IGKJ1*01 833 1438 15 VH3-23_IGHD2-8*01 > 3_IGHJ1*01 2084 O12_IGKJ1*01 833 1438 16 VH3-23_IGHD2-15*01 > 2_IGHJ1*01 2085 O12_IGKJ1*01 833 1438 17 VH3-23_IGHD2-15*01 > 3_IGHJ1*01 2086 O12_IGKJ1*01 833 1438 18 VH3-23_IGHD2-21*01 > 2_IGHJ1*01 2087 O12_IGKJ1*01 833 1438 19 VH3-23_IGHD2-21*01 > 3_IGHJ1*01 2088 O12_IGKJ1*01 833 1438 20 VH3-23_IGHD3-3*01 > 1_IGHJ1*01 2089 O12_IGKJ1*01 833 1438 21 VH3-23_IGHD3-3*01 > 2_IGHJ1*01 2090 O12_IGKJ1*01 833 1438 22 VH3-23_IGHD3-3*01 > 3_IGHJ1*01 2091 O12_IGKJ1*01 833 1438 23 VH3-23_IGHD3-9*01 > 2_IGHJ1*01 2092 O12_IGKJ1*01 833 1438 24 VH3-23_IGHD3-10*01 > 2_IGHJ1*01 2093 O12_IGKJ1*01 833 1438 25 VH3-23_IGHD3-10*01 > 3_IGHJ1*01 2094 O12_IGKJ1*01 833 1438 26 VH3-23_IGHD3-16*01 > 2_IGHJ1*01 2095 O12_IGKJ1*01 833 1438 27 VH3-23_IGHD3-16*01 > 3_IGHJ1*01 2096 O12_IGKJ1*01 833 1438 28 VH3-23_IGHD3-22*01 > 2_IGHJ1*01 2097 O12_IGKJ1*01 833 1438 29 VH3-23_IGHD3-22*01 > 3_IGHJ1*01 2098 O12_IGKJ1*01 833 1438 30 VH3-23_IGHD4-4*01 (1) > 2_IGHJ1*01 2099 O12_IGKJ1*01 833 1438 31 VH3-23_IGHD4-4*01 (1) > 3_IGHJ1*01 2100 O12_IGKJ1*01 833 1438 32 VH3-23_IGHD4-11*01 (1) > 2_IGHJ1*01 2101 O12_IGKJ1*01 833 1438 33 VH3-23_IGHD4-11*01 (1) > 3_IGHJ1*01 2102 O12_IGKJ1*01 833 1438 34 VH3-23_IGHD4-17*01 > 2_IGHJ1*01 2103 O12_IGKJ1*01 833 1438 35 VH3-23_IGHD4-17*01 > 3_IGHJ1*01 2104 O12_IGKJ1*01 833 1438 36 VH3-23_IGHD4-23*01 > 2_IGHJ1*01 2105 O12_IGKJ1*01 833 1438 37 VH3-23_IGHD4-23*01 > 3_IGHJ1*01 2106 O12_IGKJ1*01 833 1438 38 VH3-23_IGHD5-5*01 (2) > 1_IGHJ1*01 2107 O12_IGKJ1*01 833 1438 39 VH3-23_IGHD5-5*01 (2) > 2_IGHJ1*01 2108 O12_IGKJ1*01 833 1438 40 VH3-23_IGHD5-5*01 (2) > 3_IGHJ1*01 2109 O12_IGKJ1*01 833 1438 41 VH3-23_IGHD5-12*01 > 1_IGHJ1*01 2110 O12_IGKJ1*01 833 1438 42 VH3-23_IGHD5-12*01 > 3_IGHJ1*01 2111 O12_IGKJ1*01 833 1438 43 VH3-23_IGHD5-18*01 (2) > 1_IGHJ1*01 2112 O12_IGKJ1*01 833 1438 44 VH3-23_IGHD5-18*01 (2) > 2_IGHJ1*01 2113 O12_IGKJ1*01 833 1438 45 VH3-23_IGHD5-18*01 (2) > 3_IGHJ1*01 2114 O12_IGKJ1*01 833 1438 46 VH3-23_IGHD5-24*01 > 1_IGHJ1*01 2115 O12_IGKJ1*01 833 1438 47 VH3-23_IGHD5-24*01 > 3_IGHJ1*01 2116 O12_IGKJ1*01 833 1438 48 VH3-23_IGHD6-6*01 > 1_IGHJ1*01 2117 O12_IGKJ1*01 833 1438 49 VH3-23_IGHD1-1*01 > 1'_IGHJ1*01 2127 O12_IGKJ1*01 833 1438 50 VH3-23_IGHD1-1*01 > 2'_IGHJ1*01 2128 O12_IGKJ1*01 833 1438 51 VH3-23_IGHD1-1*01 > 3'_IGHJ1*01 2129 O12_IGKJ1*01 833 1438 52 VH3-23_IGHD1-7*01 > 1'_IGHJ1*01 2130 O12_IGKJ1*01 833 1438 53 VH3-23_IGHD1-7*01 > 3'_IGHJ1*01 2131 O12_IGKJ1*01 833 1438 54 VH3-23_IGHD1-14*01 > 1'_IGHJ1*01 2132 O12_IGKJ1*01 833 1438 55 VH3-23_IGHD1-14*01 > 2'_IGHJ1*01 2133 O12_IGKJ1*01 833 1438 56 VH3-23_IGHD1-14*01 > 3'_IGHJ1*01 2134 O12_IGKJ1*01 833 1438 57 VH3-23_IGHD1-20*01 > 1'_IGHJ1*01 2135 O12_IGKJ1*01 833 1438 58 VH3-23_IGHD1-20*01 > 2'_IGHJ1*01 2136 O12_IGKJ1*01 833 1438 59 VH3-23_IGHD1-20*01 > 3'_IGHJ1*01 2137 O12_IGKJ1*01 833 1438 60 VH3-23_IGHD1-26*01 > 1'_IGHJ1*01 2138 O12_IGKJ1*01 833 1438 61 VH3-23_IGHD1-26*01 > 3'_IGHJ1*01 2139 O12_IGKJ1*01 833 1438 62 VH3-23_IGHD2-2*01 > 1'_IGHJ1*01 2140 O12_IGKJ1*01 833 1438 63 VH3-23_IGHD2-2*01 > 3'_IGHJ1*01 2141 O12_IGKJ1*01 833 1438 64 VH3-23_IGHD2-8*01 > 1'_IGHJ1*01 2142 O12_IGKJ1*01 833 1438 65 VH3-23_IGHD2-15*01 > 1'_IGHJ1*01 2143 O12_IGKJ1*01 833 1438 66 VH3-23_IGHD2-15*01 > 3'_IGHJ1*01 2144 O12_IGKJ1*01 833 1438 67 VH3-23_IGHD2-21*01 > 1'_IGHJ1*01 2145 O12_IGKJ1*01 833 1438 68 VH3-23_IGHD2-21*01 > 3'_IGHJ1*01 2146 O12_IGKJ1*01 833 1438 69 VH3-23_IGHD3-3*01 > 1'_IGHJ1*01 2147 O12_IGKJ1*01 833 1438 70 VH3-23_IGHD3-3*01 > 3'_IGHJ1*01 2148 O12_IGKJ1*01 833 1438 71 VH3-23_IGHD3-9*01 > 1'_IGHJ1*01 2149 O12_IGKJ1*01 833 1438 72 VH3-23_IGHD3-9*01 > 3'_IGHJ1*01 2150 O12_IGKJ1*01 833 1438 73 VH3-23_IGHD3-10*01 > 1'_IGHJ1*01 2151 O12_IGKJ1*01 833 1438 74 VH3-23_IGHD3-10*01 > 3'_IGHJ1*01 2152 O12_IGKJ1*01 833 1438 75 VH3-23_IGHD3-16*01 > 1'_IGHJ1*01 2153 O12_IGKJ1*01 833 1438 76 VH3-23_IGHD3-16*01 > 3'_IGHJ1*01 2154 O12_IGKJ1*01 833 1438 77 VH3-23_IGHD3-22*01 > 1'_IGHJ1*01 2155 O12_IGKJ1*01 833 1438 78 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ1*01 2156 O12_IGKJ1*01 833 1438 79 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ1*01 2157 O12_IGKJ1*01 833 1438 80 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ1*01 2158 O12_IGKJ1*01 833 1438 81 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ1*01 2159 O12_IGKJ1*01 833 1438 82 VH3-23_IGHD4-17*01 > 1'_IGHJ1*01 2160 O12_IGKJ1*01 833 1438 83 VH3-23_IGHD4-17*01 > 3'_IGHJ1*01 2161 O12_IGKJ1*01 833 1438 84 VH3-23_IGHD4-23*01 > 1'_IGHJ1*01 2162 O12_IGKJ1*01 833 1438 85 VH3-23_IGHD4-23*01 > 3'_IGHJ1*01 2163 O12_IGKJ1*01 833 1438 86 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ1*01 2164 O12_IGKJ1*01 833 1438 87 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ1*01 2165 O12_IGKJ1*01 833 1438 88 VH3-23_IGHD5-12*01 > 1'_IGHJ1*01 2166 O12_IGKJ1*01 833 1438 89 VH3-23_IGHD5-12*01 > 3'_IGHJ1*01 2167 O12_IGKJ1*01 833 1438 90 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ1*01 2168 O12_IGKJ1*01 833 1438 91 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ1*01 2169 O12_IGKJ1*01 833 1438 92 VH3-23_IGHD5-24*01 > 1'_IGHJ1*01 2170 O12_IGKJ1*01 833 1438 93 VH3-23_IGHD5-24*01 > 3'_IGHJ1*01 2171 O12_IGKJ1*01 833 1438 94 VH3-23_IGHD6-6*01 > 1'_IGHJ1*01 2172 O12_IGKJ1*01 833 1438 95 VH3-23_IGHD6-6*01 > 2'_IGHJ1*01 2173 O12_IGKJ1*01 833 1438 96 VH3-23_IGHD6-6*01 > 3'_IGHJ1*01 2174 O12_IGKJ1*01 833 1438 97 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 O12_IGKJ1*01 833 1438 98 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 O12_IGKJ1*01 833 1438 99 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 O12_IGKJ1*01 833 1438 100 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 O12_IGKJ1*01 833 1438 101 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 O12_IGKJ1*01 833 1438 102 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 O12_IGKJ1*01 833 1438 103 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 O12_IGKJ1*01 833 1438 104 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 O12_IGKJ1*01 833 1438 105 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 O12_IGKJ1*01 833 1438 106 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 O12_IGKJ1*01 833 1438 107 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 O12_IGKJ1*01 833 1438 108 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 O12_IGKJ1*01 833 1438 109 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 O12_IGKJ1*01 833 1438 110 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 O12_IGKJ1*01 833 1438 111 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 O12_IGKJ1*01 833 1438 112 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 O12_IGKJ1*01 833 1438 113 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 O12_IGKJ1*01 833 1438 114 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 O12_IGKJ1*01 833 1438 115 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 O12_IGKJ1*01 833 1438 116 VH3-23_IGHD1-1*01 > 1_IGHJ2*01 2185 O12_IGKJ1*01 833 1438 117 VH3-23_IGHD1-1*01 > 2_IGHJ2*01 2186 O12_IGKJ1*01 833 1438 118 VH3-23_IGHD1-1*01 > 3_IGHJ2*01 2187 O12_IGKJ1*01 833 1438 119 VH3-23_IGHD1-7*01 > 1_IGHJ2*01 2188 O12_IGKJ1*01 833 1438 120 VH3-23_IGHD1-7*01 > 3_IGHJ2*01 2189 O12_IGKJ1*01 833 1438 121 VH3-23_IGHD1-14*01 > 1_IGHJ2*01 2190 O12_IGKJ1*01 833 1438 122 VH3-23_IGHD1-14*01 > 3_IGHJ2*01 2191 O12_IGKJ1*01 833 1438 123 VH3-23_IGHD1-20*01 > 1_IGHJ2*01 2192 O12_IGKJ1*01 833 1438 124 VH3-23_IGHD1-20*01 > 3_IGHJ2*01 2193 O12_IGKJ1*01 833 1438 125 VH3-23_IGHD1-26*01 > 1_IGHJ2*01 2194 O12_IGKJ1*01 833 1438 126 VH3-23_IGHD1-26*01 > 3_IGHJ2*01 2195 O12_IGKJ1*01 833 1438 127 VH3-23_IGHD2-2*01 > 2_IGHJ2*01 2196 O12_IGKJ1*01 833 1438 128 VH3-23_IGHD2-2*01 > 3_IGHJ2*01 2197 O12_IGKJ1*01 833 1438 129 VH3-23_IGHD2-8*01 > 2_IGHJ2*01 2198 O12_IGKJ1*01 833 1438 130 VH3-23_IGHD2-8*01 > 3_IGHJ2*01 2199 O12_IGKJ1*01 833 1438 131 VH3-23_IGHD2-15*01 > 2_IGHJ2*01 2200 O12_IGKJ1*01 833 1438 132 VH3-23_IGHD2-15*01 > 3_IGHJ2*01 2201 O12_IGKJ1*01 833 1438 133 VH3-23_IGHD2-21*01 > 2_IGHJ2*01 2202 O12_IGKJ1*01 833 1438 134 VH3-23_IGHD2-21*01 > 3_IGHJ2*01 2203 O12_IGKJ1*01 833 1438 135 VH3-23_IGHD3-3*01 > 1_IGHJ2*01 2204 O12_IGKJ1*01 833 1438 136 VH3-23_IGHD3-3*01 > 2_IGHJ2*01 2205 O12_IGKJ1*01 833 1438 137 VH3-23_IGHD3-3*01 > 3_IGHJ2*01 2206 O12_IGKJ1*01 833 1438 138 VH3-23_IGHD3-9*01 > 2_IGHJ2*01 2207 O12_IGKJ1*01 833 1438 139 VH3-23_IGHD3-10*01 > 2_IGHJ2*01 2208 O12_IGKJ1*01 833 1438 140 VH3-23_IGHD3-10*01 > 3_IGHJ2*01 2209 O12_IGKJ1*01 833 1438 141 VH3-23_IGHD3-16*01 > 2_IGHJ2*01 2210 O12_IGKJ1*01 833 1438 142 VH3-23_IGHD3-16*01 > 3_IGHJ2*01 2211 O12_IGKJ1*01 833 1438 143 VH3-23_IGHD3-22*01 > 2_IGHJ2*01 2212 O12_IGKJ1*01 833 1438 144 VH3-23_IGHD3-22*01 > 3_IGHJ2*01 2213 O12_IGKJ1*01 833 1438 145 VH3-23_IGHD4-4*01 (1) > 2_IGHJ2*01 2214 O12_IGKJ1*01 833 1438 146 VH3-23_IGHD4-4*01 (1) > 3_IGHJ2*01 2215 O12_IGKJ1*01 833 1438 147 VH3-23_IGHD4-11*01 (1) > 2_IGHJ2*01 2216 O12_IGKJ1*01 833 1438 148 VH3-23_IGHD4-11*01 (1) > 3_IGHJ2*01 2217 O12_IGKJ1*01 833 1438 149 VH3-23_IGHD4-17*01 > 2_IGHJ2*01 2218 O12_IGKJ1*01 833 1438 150 VH3-23_IGHD4-17*01 > 3_IGHJ2*01 2219 O12_IGKJ1*01 833 1438 151 VH3-23_IGHD4-23*01 > 2_IGHJ2*01 2220 O12_IGKJ1*01 833 1438 152 VH3-23_IGHD4-23*01 > 3_IGHJ2*01 2221 O12_IGKJ1*01 833 1438 153 VH3-23_IGHD5-5*01 (2) > 1_IGHJ2*01 2222 O12_IGKJ1*01 833 1438 154 VH3-23_IGHD5-5*01 (2) > 2_IGHJ2*01 2223 O12_IGKJ1*01 833 1438 155 VH3-23_IGHD5-5*01 (2) > 3_IGHJ2*01 2224 O12_IGKJ1*01 833 1438 156 VH3-23_IGHD5-12*01 > 1_IGHJ2*01 2225 O12_IGKJ1*01 833 1438 157 VH3-23_IGHD5-12*01 > 3_IGHJ2*01 2226 O12_IGKJ1*01 833 1438 158 VH3-23_IGHD5-18*01 (2) > 1_IGHJ2*01 2227 O12_IGKJ1*01 833 1438 159 VH3-23_IGHD5-18*01 (2) > 2_IGHJ2*01 2228 O12_IGKJ1*01 833 1438 160 VH3-23_IGHD5-18*01 (2) > 3_IGHJ2*01 2229 O12_IGKJ1*01 833 1438 161 VH3-23_IGHD5-24*01 > 1_IGHJ2*01 2230 O12_IGKJ1*01 833 1438 162 VH3-23_IGHD5-24*01 > 3_IGHJ2*01 2231 O12_IGKJ1*01 833 1438 163 VH3-23_IGHD6-6*01 > 1_IGHJ2*01 2232 O12_IGKJ1*01 833 1438 164 VH3-23_IGHD1-1*01 > 1'_IGHJ2*01 2242 O12_IGKJ1*01 833 1438 165 VH3-23_IGHD1-1*01 > 2'_IGHJ2*01 2243 O12_IGKJ1*01 833 1438 166 VH3-23_IGHD1-1*01 > 3'_IGHJ2*01 2244 O12_IGKJ1*01 833 1438 167 VH3-23_IGHD1-7*01 > 1'_IGHJ2*01 2245 O12_IGKJ1*01 833 1438 168 VH3-23_IGHD1-7*01 > 3'_IGHJ2*01 2246 O12_IGKJ1*01 833 1438 169 VH3-23_IGHD1-14*01 > 1'_IGHJ2*01 2247 O12_IGKJ1*01 833 1438 170 VH3-23_IGHD1-14*01 > 2'_IGHJ2*01 2248 O12_IGKJ1*01 833 1438 171 VH3-23_IGHD1-14*01 > 3'_IGHJ2*01 2249 O12_IGKJ1*01 833 1438 172 VH3-23_IGHD1-20*01 > 1'_IGHJ2*01 2250 O12_IGKJ1*01 833 1438 173 VH3-23_IGHD1-20*01 > 2'_IGHJ2*01 2251 O12_IGKJ1*01 833 1438 174 VH3-23_IGHD1-20*01 > 3'_IGHJ2*01 2252 O12_IGKJ1*01 833 1438 175 VH3-23_IGHD1-26*01 > 1'_IGHJ2*01 2253 O12_IGKJ1*01 833 1438 176 VH3-23_IGHD1-26*01 > 1_IGHJ2*01_B 2254 O12_IGKJ1*01 833 1438 177 VH3-23_IGHD2-2*01 > 1'_IGHJ2*01 2255 O12_IGKJ1*01 833 1438 178 VH3-23_IGHD2-2*01 > 3'_IGHJ2*01 2256 O12_IGKJ1*01 833 1438 179 VH3-23_IGHD2-8*01 > 1'_IGHJ2*01 2257 O12_IGKJ1*01 833 1438 180 VH3-23_IGHD2-15*01 > 1'_IGHJ2*01 2258 O12_IGKJ1*01 833 1438 181 VH3-23_IGHD2-15*01 > 3'_IGHJ2*01 2259 O12_IGKJ1*01 833 1438 182 VH3-23_IGHD2-21*01 > 1'_IGHJ2*01 2260 O12_IGKJ1*01 833 1438 183 VH3-23_IGHD2-21*01 > 3'_IGHJ2*01 2261 O12_IGKJ1*01 833 1438 184 VH3-23_IGHD3-3*01 > 1'_IGHJ2*01 2262 O12_IGKJ1*01 833 1438 185 VH3-23_IGHD3-3*01 > 3'_IGHJ2*01 2263 O12_IGKJ1*01 833 1438 186 VH3-23_IGHD3-9*01 > 1'_IGHJ2*01 2264 O12_IGKJ1*01 833 1438 187 VH3-23_IGHD3-9*01 > 3'_IGHJ2*01 2265 O12_IGKJ1*01 833 1438 188 VH3-23_IGHD3-10*01 > 1'_IGHJ2*01 2266 O12_IGKJ1*01 833 1438 189 VH3-23_IGHD3-10*01 > 3'_IGHJ2*01 2267 O12_IGKJ1*01 833 1438 190 VH3-23_IGHD3-16*01 > 1'_IGHJ2*01 2268 O12_IGKJ1*01 833 1438 191 VH3-23_IGHD3-16*01 > 3'_IGHJ2*01 2269 O12_IGKJ1*01 833 1438 192 VH3-23_IGHD3-22*01 > 1'_IGHJ2*01 2270 O12_IGKJ1*01 833 1438 193 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ2*01 2271 O12_IGKJ1*01 833 1438 194 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ2*01 2272 O12_IGKJ1*01 833 1438 195 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ2*01 2273 O12_IGKJ1*01 833 1438 196 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ2*01 2274 O12_IGKJ1*01 833 1438 197 VH3-23_IGHD4-17*01 > 1'_IGHJ2*01 2275 O12_IGKJ1*01 833 1438 198 VH3-23_IGHD4-17*01 > 3'_IGHJ2*01 2276 O12_IGKJ1*01 833 1438 199 VH3-23_IGHD4-23*01 > 1'_IGHJ2*01 2277 O12_IGKJ1*01 833 1438 200 VH3-23_IGHD4-23*01 > 3'_IGHJ2*01 2278 O12_IGKJ1*01 833 1438 201 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ2*01 2279 O12_IGKJ1*01 833 1438 202 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ2*01 2280 O12_IGKJ1*01 833 1438 203 VH3-23_IGHD5-12*01 > 1'_IGHJ2*01 2281 O12_IGKJ1*01 833 1438 204 VH3-23_IGHD5-12*01 > 3'_IGHJ2*01 2282 O12_IGKJ1*01 833 1438 205 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ2*01 2283 O12_IGKJ1*01 833 1438 206 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ2*01 2284 O12_IGKJ1*01 833 1438 207 VH3-23_IGHD5-24*01 > 1'_IGHJ2*01 2285 O12_IGKJ1*01 833 1438 208 VH3-23_IGHD5-24*01 > 3'_IGHJ2*01 2286 O12_IGKJ1*01 833 1438 209 VH3-23_IGHD6-6*01 > 1'_IGHJ2*01 2287 O12_IGKJ1*01 833 1438 210 VH3-23_IGHD6-6*01 > 2'_IGHJ2*01 2288 O12_IGKJ1*01 833 1438 211 VH3-23_IGHD6-6*01 > 3'_IGHJ2*01 2289 O12_IGKJ1*01 833 1438 212 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 O12_IGKJ1*01 833 1438 213 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 O12_IGKJ1*01 833 1438 214 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 O12_IGKJ1*01 833 1438 215 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 O12_IGKJ1*01 833 1438 216 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 O12_IGKJ1*01 833 1438 217 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 O12_IGKJ1*01 833 1438 218 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 O12_IGKJ1*01 833 1438 219 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 O12_IGKJ1*01 833 1438 220 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 O12_IGKJ1*01 833 1438 221 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 O12_IGKJ1*01 833 1438 222 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 O12_IGKJ1*01 833 1438 223 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 O12_IGKJ1*01 833 1438 224 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 O12_IGKJ1*01 833 1438 225 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 O12_IGKJ1*01 833 1438 226 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 O12_IGKJ1*01 833 1438 227 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 O12_IGKJ1*01 833 1438 228 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 O12_IGKJ1*01 833 1438 229 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 O12_IGKJ1*01 833 1438 230 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 O12_IGKJ1*01 833 1438 231 VH3-23_IGHD1-1*01 > 1_IGHJ3*01 2300 O12_IGKJ1*01 833 1438 232 VH3-23_IGHD1-1*01 > 2_IGHJ3*01 2301 O12_IGKJ1*01 833 1438 233 VH3-23_IGHD1-1*01 > 3_IGHJ3*01 2302 O12_IGKJ1*01 833 1438 234 VH3-23_IGHD1-7*01 > 1_IGHJ3*01 2303 O12_IGKJ1*01 833 1438 235 VH3-23_IGHD1-7*01 > 3_IGHJ3*01 2304 O12_IGKJ1*01 833 1438 236 VH3-23_IGHD1-14*01 > 1_IGHJ3*01 2305 O12_IGKJ1*01 833 1438 237 VH3-23_IGHD1-14*01 > 3_IGHJ3*01 2306 O12_IGKJ1*01 833 1438 238 VH3-23_IGHD1-20*01 > 1_IGHJ3*01 2307 O12_IGKJ1*01 833 1438 239 VH3-23_IGHD1-20*01 > 3_IGHJ3*01 2308 O12_IGKJ1*01 833 1438 240 VH3-23_IGHD1-26*01 > 1_IGHJ3*01 2309 O12_IGKJ1*01 833 1438 241 VH3-23_IGHD1-26*01 > 3_IGHJ3*01 2310 O12_IGKJ1*01 833 1438 242 VH3-23_IGHD2-2*01 > 2_IGHJ3*01 2311 O12_IGKJ1*01 833 1438 243 VH3-23_IGHD2-2*01 > 3_IGHJ3*01 2312 O12_IGKJ1*01 833 1438
244 VH3-23_IGHD2-8*01 > 2_IGHJ3*01 2313 O12_IGKJ1*01 833 1438 245 VH3-23_IGHD2-8*01 > 3_IGHJ3*01 2314 O12_IGKJ1*01 833 1438 246 VH3-23_IGHD2-15*01 > 2_IGHJ3*01 2315 O12_IGKJ1*01 833 1438 247 VH3-23_IGHD2-15*01 > 3_IGHJ3*01 2316 O12_IGKJ1*01 833 1438 248 VH3-23_IGHD2-21*01 > 2_IGHJ3*0 1 2317 O12_IGKJ1*01 833 1438 249 VH3-23_IGHD2-21*01 > 3_IGHJ3*01 2318 O12_IGKJ1*01 833 1438 250 VH3-23_IGHD3-3*01 > 1_IGHJ3*01 2319 O12_IGKJ1*01 833 1438 251 VH3-23_IGHD3-3*01 > 2_IGHJ3*01 2320 O12_IGKJ1*01 833 1438 252 VH3-23_IGHD3-3*01 > 3_IGHJ3*01 2321 O12_IGKJ1*01 833 1438 253 VH3-23_IGHD3-9*01 > 2_IGHJ3*01 2322 O12_IGKJ1*01 833 1438 254 VH3-23_IGHD3-10*01 > 2_IGHJ3*01 2323 O12_IGKJ1*01 833 1438 255 VH3-23_IGHD3-10*01 > 3_IGHJ3*01 2324 O12_IGKJ1*01 833 1438 256 VH3-23_IGHD3-16*01 > 2_IGHJ3*01 2325 O12_IGKJ1*01 833 1438 257 VH3-23_IGHD3-16*01 > 3_IGHJ3*01 2326 O12_IGKJ1*01 833 1438 258 VH3-23_IGHD3-22*01 > 2_IGHJ3*01 2327 O12_IGKJ1*01 833 1438 259 VH3-23_IGHD3-22*01 > 3_IGHJ3*01 2328 O12_IGKJ1*01 833 1438 260 VH3-23_IGHD4-4*01 (1) > 2_IGHJ3*01 2329 O12_IGKJ1*01 833 1438 261 VH3-23_IGHD4-4*01 (1) > 3_IGHJ3*01 2330 O12_IGKJ1*01 833 1438 262 VH3-23_IGHD4-11*01 (1) > 2_IGHJ3*01 2331 O12_IGKJ1*01 833 1438 263 VH3-23_IGHD4-11*01 (1) > 3_IGHJ3*01 2332 O12_IGKJ1*01 833 1438 264 VH3-23_IGHD4-17*01 > 2_IGHJ3*01 2333 O12_IGKJ1*01 833 1438 265 VH3-23_IGHD4-17*01 > 3_IGHJ3*01 2334 O12_IGKJ1*01 833 1438 266 VH3-23_IGHD4-23*01 > 2_IGHJ3*01 2335 O12_IGKJ1*01 833 1438 267 VH3-23_IGHD4-23*01 > 3_IGHJ3*01 2336 O12_IGKJ1*01 833 1438 268 VH3-23_IGHD5-5*01 (2) > 1_IGHJ3*01 2337 O12_IGKJ1*01 833 1438 269 VH3-23_IGHD5-5*01 (2) > 2_IGHJ3*01 2338 O12_IGKJ1*01 833 1438 270 VH3-23_IGHD5-5*01 (2) > 3_IGHJ3*01 2339 O12_IGKJ1*01 833 1438 271 VH3-23_IGHD5-12*01 > 1_IGHJ3*01 2340 O12_IGKJ1*01 833 1438 272 VH3-23_IGHD5-12*01 > 3_IGHJ3*01 2341 O12_IGKJ1*01 833 1438 273 VH3-23_IGHD5-18*01 (2) > 1_IGHJ3*01 2342 O12_IGKJ1*01 833 1438 274 VH3-23_IGHD5-18*01 (2) > 2_IGHJ3*01 2343 O12_IGKJ1*01 833 1438 275 VH3-23_IGHD5-18*01 (2) > 3_IGHJ3*01 2344 O12_IGKJ1*01 833 1438 276 VH3-23_IGHD5-24*01 > 1_IGHJ3*01 2345 O12_IGKJ1*01 833 1438 277 VH3-23_IGHD5-24*01 > 3_IGHJ3*01 2346 O12_IGKJ1*01 833 1438 278 VH3-23_IGHD6-6*01 > 1_IGHJ3*01 2347 O12_IGKJ1*01 833 1438 279 VH3-23_IGHD1-1*01 > 1'_IGHJ3*01 2357 O12_IGKJ1*01 833 1438 280 VH3-23_IGHD1-1*01 > 2'_IGHJ3*01 2358 O12_IGKJ1*01 833 1438 281 VH3-23_IGHD1-1*01 > 3'_IGHJ3*01 2359 O12_IGKJ1*01 833 1438 282 VH3-23_IGHD1-7*01 > 1'_IGHJ3*01 2360 O12_IGKJ1*01 833 1438 283 VH3-23_IGHD1-7*01 > 3'_IGHJ3*01 2361 O12_IGKJ1*01 833 1438 284 VH3-23_IGHD1-14*01 > 1'_IGHJ3*01 2362 O12_IGKJ1*01 833 1438 285 VH3-23_IGHD1-14*01 > 2'_IGHJ3*01 2363 O12_IGKJ1*01 833 1438 286 VH3-23_IGHD1-14*01 > 3'_IGHJ3*01 2364 O12_IGKJ1*01 833 1438 287 VH3-23_IGHD1-20*01 > 1'_IGHJ3*01 2365 O12_IGKJ1*01 833 1438 288 VH3-23_IGHD1-20*01 > 2'_IGHJ3*01 2366 O12_IGKJ1*01 833 1438 289 VH3-23_IGHD1-20*01 > 3'_IGHJ3*01 2367 O12_IGKJ1*01 833 1438 290 VH3-23_IGHD1-26*01 > 1'_IGHJ3*01 2368 O12_IGKJ1*01 833 1438 291 VH3-23_IGHD1-26*01 > 3'_IGHJ3*01 2369 O12_IGKJ1*01 833 1438 292 VH3-23_IGHD2-2*01 > 1'_IGHJ3*01 2370 O12_IGKJ1*01 833 1438 293 VH3-23_IGHD2-2*01 > 3'_IGHJ3*01 2371 O12_IGKJ1*01 833 1438 294 VH3-23_IGHD2-8*01 > 1'_IGHJ3*01 2372 O12_IGKJ1*01 833 1438 295 VH3-23_IGHD2-15*01 > 1'_IGHJ3*01 2373 O12_IGKJ1*01 833 1438 296 VH3-23_IGHD2-15*01 > 3'_IGHJ3*01 2374 O12_IGKJ1*01 833 1438 297 VH3-23_IGHD2-21*01 > 1'_IGHJ3*01 2375 O12_IGKJ1*01 833 1438 298 VH3-23_IGHD2-21*01 > 3'_IGHJ3*01 2376 O12_IGKJ1*01 833 1438 299 VH3-23_IGHD3-3*01 > 1'_IGHJ3*01 2377 O12_IGKJ1*01 833 1438 300 VH3-23_IGHD3-3*01 > 3'_IGHJ3*01 2378 O12_IGKJ1*01 833 1438 301 VH3-23_IGHD3-9*01 > 1'_IGHJ3*01 2379 O12_IGKJ1*01 833 1438 302 VH3-23_IGHD3-9*01 > 3'_IGHJ3*01 2380 O12_IGKJ1*01 833 1438 303 VH3-23_IGHD3-10*01 > 1'_IGHJ3*01 2381 O12_IGKJ1*01 833 1438 304 VH3-23_IGHD3-10*01 > 3'_IGHJ3*01 2382 O12_IGKJ1*01 833 1438 305 VH3-23_IGHD3-16*01 > 1'_IGHJ3*01 2383 O12_IGKJ1*01 833 1438 306 VH3-23_IGHD3-16*01 > 3'_IGHJ3*01 2384 O12_IGKJ1*01 833 1438 307 VH3-23_IGHD3-22*01 > 1'_IGHJ3*01 2385 O12_IGKJ1*01 833 1438 308 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ3*01 2386 O12_IGKJ1*01 833 1438 309 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ3*01 2387 O12_IGKJ1*01 833 1438 310 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ3*01 2388 O12_IGKJ1*01 833 1438 311 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ3*01 2389 O12_IGKJ1*01 833 1438 312 VH3-23_IGHD4-17*01 > 1'_IGHJ3*01 2390 O12_IGKJ1*01 833 1438 313 VH3-23_IGHD4-17*01 > 3'_IGHJ3*01 2391 O12_IGKJ1*01 833 1438 314 VH3-23_IGHD4-23*01 > 1'_IGHJ3*01 2392 O12_IGKJ1*01 833 1438 315 VH3-23_IGHD4-23*01 > 3'_IGHJ3*01 2393 O12_IGKJ1*01 833 1438 316 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ3*01 2394 O12_IGKJ1*01 833 1438 317 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ3*01 2395 O12_IGKJ1*01 833 1438 318 VH3-23_IGHD5-12*01 > 1'_IGHJ3*01 2396 O12_IGKJ1*01 833 1438 319 VH3-23_IGHD5-12*01 > 3'_IGHJ3*01 2397 O12_IGKJ1*01 833 1438 320 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ3*01 2398 O12_IGKJ1*01 833 1438 321 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ3*01 2399 O12_IGKJ1*01 833 1438 322 VH3-23_IGHD5-24*01 > 1'_IGHJ3*01 2400 O12_IGKJ1*01 833 1438 323 VH3-23_IGHD5-24*01 > 3'_IGHJ3*01 2401 O12_IGKJ1*01 833 1438 324 VH3-23_IGHD6-6*01 > 1'_IGHJ3*01 2402 O12_IGKJ1*01 833 1438 325 VH3-23_IGHD6-6*01 > 2'_IGHJ3*01 2403 O12_IGKJ1*01 833 1438 326 VH3-23_IGHD6-6*01 > 3'_IGHJ3*01 2404 O12_IGKJ1*01 833 1438 327 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 O12_IGKJ1*01 833 1438 328 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 O12_IGKJ1*01 833 1438 329 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 O12_IGKJ1*01 833 1438 330 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 O12_IGKJ1*01 833 1438 331 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 O12_IGKJ1*01 833 1438 332 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 O12_IGKJ1*01 833 1438 333 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 O12_IGKJ1*01 833 1438 334 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 O12_IGKJ1*01 833 1438 335 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 O12_IGKJ1*01 833 1438 336 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 O12_IGKJ1*01 833 1438 337 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 O12_IGKJ1*01 833 1438 338 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 O12_IGKJ1*01 833 1438 339 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 O12_IGKJ1*01 833 1438 340 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 O12_IGKJ1*01 833 1438 341 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 O12_IGKJ1*01 833 1438 342 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 O12_IGKJ1*01 833 1438 343 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 O12_IGKJ1*01 833 1438 344 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 O12_IGKJ1*01 833 1438 345 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 O12_IGKJ1*01 833 1438 346 VH3-23_IGHD1-1*01 > 1_IGHJ4*01 2415 O12_IGKJ1*01 833 1438 347 VH3-23_IGHD1-1*01 > 2_IGHJ4*01 2416 O12_IGKJ1*01 833 1438 348 VH3-23_IGHD1-1*01 > 3_IGHJ4*01 2417 O12_IGKJ1*01 833 1438 349 VH3-23_IGHD1-7*01 > 1_IGHJ4*01 2418 O12_IGKJ1*01 833 1438 350 VH3-23_IGHD1-7*01 > 3_IGHJ4*01 2419 O12_IGKJ1*01 833 1438 351 VH3-23_IGHD1-14*01 > 1_IGHJ4*01 2420 O12_IGKJ1*01 833 1438 352 VH3-23_IGHD1-14*01 > 3_IGHJ4*01 2421 O12_IGKJ1*01 833 1438 353 VH3-23_IGHD1-20*01 > 1_IGHJ4*01 2422 O12_IGKJ1*01 833 1438 354 VH3-23_IGHD1-20*01 > 3_IGHJ4*01 2423 O12_IGKJ1*01 833 1438 355 VH3-23_IGHD1-26*01 > 1_IGHJ4*01 2424 O12_IGKJ1*01 833 1438 356 VH3-23_IGHD1-26*01 > 3_IGHJ4*01 2425 O12_IGKJ1*01 833 1438 357 VH3-23_IGHD2-2*01 > 2_IGHJ4*01 2426 O12_IGKJ1*01 833 1438 358 VH3-23_IGHD2-2*01 > 3_IGHJ4*01 2427 O12_IGKJ1*01 833 1438 359 VH3-23_IGHD2-8*01 > 2_IGHJ4*01 2428 O12_IGKJ1*01 833 1438 360 VH3-23_IGHD2-8*01 > 3_IGHJ4*01 2429 O12_IGKJ1*01 833 1438 361 VH3-23_IGHD2-15*01 > 2_IGHJ4*01 2430 O12_IGKJ1*01 833 1438 362 VH3-23_IGHD2-15*01 > 3_IGHJ4*01 2431 O12_IGKJ1*01 833 1438 363 VH3-23_IGHD2-21*01 > 2_IGHJ4*01 2432 O12_IGKJ1*01 833 1438 364 VH3-23_IGHD2-21*01 > 3_IGHJ4*01 2433 O12_IGKJ1*01 833 1438 365 VH3-23_IGHD3-3*01 > 1_IGHJ4*01 2434 O12_IGKJ1*01 833 1438 366 VH3-23_IGHD3-3*01 > 2_IGHJ4*01 2435 O12_IGKJ1*01 833 1438 367 VH3-23_IGHD3-3*01 > 3_IGHJ4*01 2436 O12_IGKJ1*01 833 1438 368 VH3-23_IGHD3-9*01 > 2_IGHJ4*01 2437 O12_IGKJ1*01 833 1438 369 VH3-23_IGHD3-10*01 > 2_IGHJ4*01 2438 O12_IGKJ1*01 833 1438 370 VH3-23_IGHD3-10*01 > 3_IGHJ4*01 2439 O12_IGKJ1*01 833 1438 371 VH3-23_IGHD3-16*01 > 2_IGHJ4*01 2440 O12_IGKJ1*01 833 1438 372 VH3-23_IGHD3-16*01 > 3_IGHJ4*01 2441 O12_IGKJ1*01 833 1438 373 VH3-23_IGHD3-22*01 > 2_IGHJ4*01 2442 O12_IGKJ1*01 833 1438 374 VH3-23_IGHD3-22*01 > 3_IGHJ4*01 2443 O12_IGKJ1*01 833 1438 375 VH3-23_IGHD4-4*01 (1) > 2_IGHJ4*01 2444 O12_IGKJ1*01 833 1438 376 VH3-23_IGHD4-4*01 (1) > 3_IGHJ4*01 2445 O12_IGKJ1*01 833 1438 377 VH3-23_IGHD4-11*01 (1) > 2_IGHJ4*01 2446 O12_IGKJ1*01 833 1438 378 VH3-23_IGHD4-11*01 (1) > 3_IGHJ4*01 2447 O12_IGKJ1*01 833 1438 379 VH3-23_IGHD4-17*01 > 2_IGHJ4*01 2448 O12_IGKJ1*01 833 1438 380 VH3-23_IGHD4-17*01 > 3_IGHJ4*01 2449 O12_IGKJ1*01 833 1438 381 VH3-23_IGHD4-23*01 > 2_IGHJ4*01 2450 O12_IGKJ1*01 833 1438 382 VH3-23_IGHD4-23*01 > 3_IGHJ4*01 2451 O12_IGKJ1*01 833 1438 383 VH3-23_IGHD5-5*01 (2) > 1_IGHJ4*01 2452 O12_IGKJ1*01 833 1438 384 VH3-23_IGHD5-5*01 (2) > 2_IGHJ4*01 2453 O12_IGKJ1*01 833 1438 385 VH3-23_IGHD5-5*01 (2) > 3_IGHJ4*01 2454 O12_IGKJ1*01 833 1438 386 VH3-23_IGHD5-12*01 > 1_IGHJ4*01 2455 O12_IGKJ1*01 833 1438 387 VH3-23_IGHD5-12*01 > 3_IGHJ4*01 2456 O12_IGKJ1*01 833 1438 388 VH3-23_IGHD5-18*01 (2) > 1_IGHJ4*01 2457 O12_IGKJ1*01 833 1438 389 VH3-23_IGHD5-18*01 (2) > 2_IGHJ4*01 2458 O12_IGKJ1*01 833 1438 390 VH3-23_IGHD5-18*01 (2) > 3_IGHJ4*01 2459 O12_IGKJ1*01 833 1438 391 VH3-23_IGHD5-24*01 > 1_IGHJ4*01 2460 O12_IGKJ1*01 833 1438 392 VH3-23_IGHD5-24*01 > 3_IGHJ4*01 2461 O12_IGKJ1*01 833 1438 393 VH3-23_IGHD6-6*01 > 1_IGHJ4*01 2462 O12_IGKJ1*01 833 1438 394 VH3-23_IGHD1-1*01 > 1'_IGHJ4*01 2472 O12_IGKJ1*01 833 1438 395 VH3-23_IGHD1-1*01 > 2'_IGHJ4*01 2473 O12_IGKJ1*01 833 1438 396 VH3-23_IGHD1-1*01 > 3'_IGHJ4*01 2474 O12_IGKJ1*01 833 1438 397 VH3-23_IGHD1-7*01 > 1'_IGHJ4*01 2475 O12_IGKJ1*01 833 1438 398 VH3-23_IGHD1-7*01 > 3'_IGHJ4*01 2476 O12_IGKJ1*01 833 1438 399 VH3-23_IGHD1-14*01 > 1'_IGHJ4*01 2477 O12_IGKJ1*01 833 1438 400 VH3-23_IGHD1-14*01 > 2'_IGHJ4*01 2478 O12_IGKJ1*01 833 1438 401 VH3-23_IGHD1-14*01 > 3'_IGHJ4*01 2479 O12_IGKJ1*01 833 1438 402 VH3-23_IGHD1-20*01 > 1'_IGHJ4*01 2480 O12_IGKJ1*01 833 1438 403 VH3-23_IGHD1-20*01 > 2'_IGHJ4*01 2481 O12_IGKJ1*01 833 1438 404 VH3-23_IGHD1-20*01 > 3'_IGHJ4*01 2482 O12_IGKJ1*01 833 1438 405 VH3-23_IGHD1-26*01 > 1'_IGHJ4*01 2483 O12_IGKJ1*01 833 1438 406 VH3-23_IGHD1-26*01 > 1_IGHJ4*01_B 2484 O12_IGKJ1*01 833 1438 407 VH3-23_IGHD2-2*01 > 1'_IGHJ4*01 2485 O12_IGKJ1*01 833 1438 408 VH3-23_IGHD2-2*01 > 3'_IGHJ4*01 2486 O12_IGKJ1*01 833 1438 409 VH3-23_IGHD2-8*01 > 1'_IGHJ4*01 2487 O12_IGKJ1*01 833 1438 410 VH3-23_IGHD2-15*01 > 1'_IGHJ4*01 2488 O12_IGKJ1*01 833 1438 411 VH3-23_IGHD2-15*01 > 3'_IGHJ4*01 2489 O12_IGKJ1*01 833 1438 412 VH3-23_IGHD2-21*01 > 1'_IGHJ4*01 2490 O12_IGKJ1*01 833 1438 413 VH3-23_IGHD2-21*01 > 3'_IGHJ4*01 2491 O12_IGKJ1*01 833 1438 414 VH3-23_IGHD3-3*01 > 1'_IGHJ4*01 2492 O12_IGKJ1*01 833 1438 415 VH3-23_IGHD3-3*01 > 3'_IGHJ4*01 2493 O12_IGKJ1*01 833 1438 416 VH3-23_IGHD3-9*01 > 1'_IGHJ4*01 2494 O12_IGKJ1*01 833 1438 417 VH3-23_IGHD3-9*01 > 3'_IGHJ4*01 2495 O12_IGKJ1*01 833 1438 418 VH3-23_IGHD3-10*01 > 1'_IGHJ4*01 2496 O12_IGKJ1*01 833 1438 419 VH3-23_IGHD3-10*01 > 3'_IGHJ4*01 2497 O12_IGKJ1*01 833 1438 420 VH3-23_IGHD3-16*01 > 1'_IGHJ4*01 2498 O12_IGKJ1*01 833 1438 421 VH3-23_IGHD3-16*01 > 3'_IGHJ4*01 2499 O12_IGKJ1*01 833 1438 422 VH3-23_IGHD3-22*01 > 1'_IGHJ4*01 2500 O12_IGKJ1*01 833 1438 423 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ4*01 2501 O12_IGKJ1*01 833 1438 424 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ4*01 2502 O12_IGKJ1*01 833 1438 425 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ4*01 2503 O12_IGKJ1*01 833 1438 426 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ4*01 2504 O12_IGKJ1*01 833 1438 427 VH3-23_IGHD4-17*01 > 1'_IGHJ4*01 2505 O12_IGKJ1*01 833 1438 428 VH3-23_IGHD4-17*01 > 3'_IGHJ4*01 2506 O12_IGKJ1*01 833 1438 429 VH3-23_IGHD4-23*01 > 1'_IGHJ4*01 2507 O12_IGKJ1*01 833 1438 430 VH3-23_IGHD4-23*01 > 3'_IGHJ4*01 2508 O12_IGKJ1*01 833 1438 431 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ4*01 2509 O12_IGKJ1*01 833 1438 432 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ4*01 2510 O12_IGKJ1*01 833 1438 433 VH3-23_IGHD5-12*01 > 1'_IGHJ4*01 2511 O12_IGKJ1*01 833 1438 434 VH3-23_IGHD5-12*01 > 3'_IGHJ4*01 2512 O12_IGKJ1*01 833 1438 435 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ4*01 2513 O12_IGKJ1*01 833 1438 436 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ4*01 2514 O12_IGKJ1*01 833 1438 437 VH3-23_IGHD5-24*01 > 1'_IGHJ4*01 2515 O12_IGKJ1*01 833 1438 438 VH3-23_IGHD5-24*01 > 3'_IGHJ4*01 2516 O12_IGKJ1*01 833 1438 439 VH3-23_IGHD6-6*01 > 1'_IGHJ4*01 2517 O12_IGKJ1*01 833 1438 440 VH3-23_IGHD6-6*01 > 2'_IGHJ4*01 2518 O12_IGKJ1*01 833 1438 441 VH3-23_IGHD6-6*01 > 3'_IGHJ4*01 2519 O12_IGKJ1*01 833 1438 442 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 O12_IGKJ1*01 833 1438 443 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 O12_IGKJ1*01 833 1438 444 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 O12_IGKJ1*01 833 1438 445 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 O12_IGKJ1*01 833 1438 446 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 O12_IGKJ1*01 833 1438 447 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 O12_IGKJ1*01 833 1438 448 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 O12_IGKJ1*01 833 1438 449 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 O12_IGKJ1*01 833 1438 450 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 O12_IGKJ1*01 833 1438 451 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 O12_IGKJ1*01 833 1438 452 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 O12_IGKJ1*01 833 1438 453 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 O12_IGKJ1*01 833 1438 454 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 O12_IGKJ1*01 833 1438 455 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 O12_IGKJ1*01 833 1438 456 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 O12_IGKJ1*01 833 1438 457 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 O12_IGKJ1*01 833 1438 458 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 O12_IGKJ1*01 833 1438 459 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 O12_IGKJ1*01 833 1438 460 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 O12_IGKJ1*01 833 1438 461 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 O12_IGKJ1*01 833 1438 462 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 O12_IGKJ1*01 833 1438 463 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 O12_IGKJ1*01 833 1438 464 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 O12_IGKJ1*01 833 1438 465 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 O12_IGKJ1*01 833 1438 466 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 O12_IGKJ1*01 833 1438 467 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 O12_IGKJ1*01 833 1438 468 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 O12_IGKJ1*01 833 1438 469 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 O12_IGKJ1*01 833 1438 470 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 O12_IGKJ1*01 833 1438 471 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 O12_IGKJ1*01 833 1438 472 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 O12_IGKJ1*01 833 1438 473 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 O12_IGKJ1*01 833 1438 474 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 O12_IGKJ1*01 833 1438 475 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 O12_IGKJ1*01 833 1438 476 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 O12_IGKJ1*01 833 1438 477 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 O12_IGKJ1*01 833 1438 478 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 O12_IGKJ1*01 833 1438 479 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 O12_IGKJ1*01 833 1438 480 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 O12_IGKJ1*01 833 1438 481 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 O12_IGKJ1*01 833 1438 482 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 O12_IGKJ1*01 833 1438 483 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 O12_IGKJ1*01 833 1438 484 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 O12_IGKJ1*01 833 1438 485 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 O12_IGKJ1*01 833 1438 486 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 O12_IGKJ1*01 833 1438 487 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 O12_IGKJ1*01 833 1438 488 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 O12_IGKJ1*01 833 1438 489 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 O12_IGKJ1*01 833 1438 490 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 O12_IGKJ1*01 833 1438 491 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 O12_IGKJ1*01 833 1438 492 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 O12_IGKJ1*01 833 1438 493 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 O12_IGKJ1*01 833 1438 494 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 O12_IGKJ1*01 833 1438
495 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 O12_IGKJ1*01 833 1438 496 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 O12_IGKJ1*01 833 1438 497 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 O12_IGKJ1*01 833 1438 498 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 O12_IGKJ1*01 833 1438 499 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 O12_IGKJ1*01 833 1438 500 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 O12_IGKJ1*01 833 1438 501 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 O12_IGKJ1*01 833 1438 502 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 O12_IGKJ1*01 833 1438 503 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 O12_IGKJ1*01 833 1438 504 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 O12_IGKJ1*01 833 1438 505 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 O12_IGKJ1*01 833 1438 506 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 O12_IGKJ1*01 833 1438 507 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 O12_IGKJ1*01 833 1438 508 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 O12_IGKJ1*01 833 1438 509 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 O12_IGKJ1*01 833 1438 510 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 O12_IGKJ1*01 833 1438 511 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 O12_IGKJ1*01 833 1438 512 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 O12_IGKJ1*01 833 1438 513 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 O12_IGKJ1*01 833 1438 514 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 O12_IGKJ1*01 833 1438 515 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 O12_IGKJ1*01 833 1438 516 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 O12_IGKJ1*01 833 1438 517 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 O12_IGKJ1*01 833 1438 518 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 O12_IGKJ1*01 833 1438 519 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 O12_IGKJ1*01 833 1438 520 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 O12_IGKJ1*01 833 1438 521 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 O12_IGKJ1*01 833 1438 522 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 O12_IGKJ1*01 833 1438 523 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 O12_IGKJ1*01 833 1438 524 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 O12_IGKJ1*01 833 1438 525 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 O12_IGKJ1*01 833 1438 526 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 O12_IGKJ1*01 833 1438 527 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 O12_IGKJ1*01 833 1438 528 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 O12_IGKJ1*01 833 1438 529 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 O12_IGKJ1*01 833 1438 530 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 O12_IGKJ1*01 833 1438 531 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 O12_IGKJ1*01 833 1438 532 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 O12_IGKJ1*01 833 1438 533 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 O12_IGKJ1*01 833 1438 534 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 O12_IGKJ1*01 833 1438 535 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 O12_IGKJ1*01 833 1438 536 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 O12_IGKJ1*01 833 1438 537 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 O12_IGKJ1*01 833 1438 538 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 O12_IGKJ1*01 833 1438 539 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 O12_IGKJ1*01 833 1438 540 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 O12_IGKJ1*01 833 1438 541 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 O12_IGKJ1*01 833 1438 542 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 O12_IGKJ1*01 833 1438 543 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 O12_IGKJ1*01 833 1438 544 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 O12_IGKJ1*01 833 1438 545 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 O12_IGKJ1*01 833 1438 546 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 O12_IGKJ1*01 833 1438 547 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 O12_IGKJ1*01 833 1438 548 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 O12_IGKJ1*01 833 1438 549 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 O12_IGKJ1*01 833 1438 550 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 O12_IGKJ1*01 833 1438 551 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 O12_IGKJ1*01 833 1438 552 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 O12_IGKJ1*01 833 1438 553 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 O12_IGKJ1*01 833 1438 554 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 O12_IGKJ1*01 833 1438 555 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 O12_IGKJ1*01 833 1438 556 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 O12_IGKJ1*01 833 1438 557 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 O12_IGKJ1*01 833 1438 558 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 O12_IGKJ1*01 833 1438 559 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 O12_IGKJ1*01 833 1438 560 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 O12_IGKJ1*01 833 1438 561 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 O12_IGKJ1*01 833 1438 562 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 O12_IGKJ1*01 833 1438 563 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 O12_IGKJ1*01 833 1438 564 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 O12_IGKJ1*01 833 1438 565 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 O12_IGKJ1*01 833 1438 566 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 O12_IGKJ1*01 833 1438 567 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 O12_IGKJ1*01 833 1438 568 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 O12_IGKJ1*01 833 1438 569 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 O12_IGKJ1*01 833 1438 570 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 O12_IGKJ1*01 833 1438 571 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 O12_IGKJ1*01 833 1438 572 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 O12_IGKJ1*01 833 1438 573 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 O12_IGKJ1*01 833 1438 574 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 O12_IGKJ1*01 833 1438 575 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 O12_IGKJ1*01 833 1438 576 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 O12_IGKJ1*01 833 1438 577 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 O12_IGKJ1*01 833 1438 578 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 O12_IGKJ1*01 833 1438 579 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 O12_IGKJ1*01 833 1438 580 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 O12_IGKJ1*01 833 1438 581 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 O12_IGKJ1*01 833 1438 582 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 O12_IGKJ1*01 833 1438 583 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 O12_IGKJ1*01 833 1438 584 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 O12_IGKJ1*01 833 1438 585 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 O12_IGKJ1*01 833 1438 586 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 O12_IGKJ1*01 833 1438 587 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 O12_IGKJ1*01 833 1438 588 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 O12_IGKJ1*01 833 1438 589 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 O12_IGKJ1*01 833 1438 590 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 O12_IGKJ1*01 833 1438 591 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 O12_IGKJ1*01 833 1438 592 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 O12_IGKJ1*01 833 1438 593 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 O12_IGKJ1*01 833 1438 594 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 O12_IGKJ1*01 833 1438 595 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 O12_IGKJ1*01 833 1438 596 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 O12_IGKJ1*01 833 1438 597 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 O12_IGKJ1*01 833 1438 598 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 O12_IGKJ1*01 833 1438 599 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 O12_IGKJ1*01 833 1438 600 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 O12_IGKJ1*01 833 1438 601 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 O12_IGKJ1*01 833 1438 602 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 O12_IGKJ1*01 833 1438 603 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 O12_IGKJ1*01 833 1438 604 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 O12_IGKJ1*01 833 1438 605 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 O12_IGKJ1*01 833 1438 606 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 O12_IGKJ1*01 833 1438 607 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 O12_IGKJ1*01 833 1438 608 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 O12_IGKJ1*01 833 1438 609 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 O12_IGKJ1*01 833 1438 610 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 O12_IGKJ1*01 833 1438 611 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 O12_IGKJ1*01 833 1438 612 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 O12_IGKJ1*01 833 1438 613 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 O12_IGKJ1*01 833 1438 614 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 O12_IGKJ1*01 833 1438 615 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 O12_IGKJ1*01 833 1438 616 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 O12_IGKJ1*01 833 1438 617 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 O12_IGKJ1*01 833 1438 618 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 O12_IGKJ1*01 833 1438 619 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 O12_IGKJ1*01 833 1438 620 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 O12_IGKJ1*01 833 1438 621 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 O12_IGKJ1*01 833 1438 622 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 O12_IGKJ1*01 833 1438 623 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 O12_IGKJ1*01 833 1438 624 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 O12_IGKJ1*01 833 1438 625 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 O12_IGKJ1*01 833 1438 626 VH3-23_IGHD5-18*01(2) > 1'_IGHJ6*01 2743 O12_IGKJ1*01 833 1438 627 VH3-23_IGHD5-18*01(2) > 3'_IGHJ6*01 2744 O12_IGKJ1*01 833 1438 628 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 O12_IGKJ1*01 833 1438 629 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 O12_IGKJ1*01 833 1438 630 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 O12_IGKJ1*01 833 1438 631 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 O12_IGKJ1*01 833 1438 632 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 O12_IGKJ1*01 833 1438 633 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 O12_IGKJ1*01 833 1438 634 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 O12_IGKJ1*01 833 1438 635 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 O12_IGKJ1*01 833 1438 636 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 O12_IGKJ1*01 833 1438 637 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 O12_IGKJ1*01 833 1438 638 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 O12_IGKJ1*01 833 1438 639 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 O12_IGKJ1*01 833 1438 640 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 O12_IGKJ1*01 833 1438 641 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 O12_IGKJ1*01 833 1438 642 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 O12_IGKJ1*01 833 1438 643 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 O12_IGKJ1*01 833 1438 644 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 O12_IGKJ1*01 833 1438 645 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 O12_IGKJ1*01 833 1438 646 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 O12_IGKJ1*01 833 1438 647 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 O12_IGKJ1*01 833 1438 648 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 O12_IGKJ1*01 833 1438 649 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 O12_IGKJ1*01 833 1438 650 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 O12_IGKJ1*01 833 1438 651 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 O12_IGKJ1*01 833 1438 652 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 O12_IGKJ1*01 833 1438 653 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 O12_IGKJ1*01 833 1438 654 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 O12_IGKJ1*01 833 1438 655 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 O12_IGKJ1*01 833 1438 656 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 O12_IGKJ1*01 833 1438 657 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 O12_IGKJ1*01 833 1438 658 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 O12_IGKJ1*01 833 1438 659 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 O12_IGKJ1*01 833 1438 660 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 O12_IGKJ1*01 833 1438 661 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 O12_IGKJ1*01 833 1438 662 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 O12_IGKJ1*01 833 1438 663 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 O12_IGKJ1*01 833 1438 664 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 O12_IGKJ1*01 833 1438 665 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 O12_IGKJ1*01 833 1438 666 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 O12_IGKJ1*01 833 1438 667 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 O12_IGKJ1*01 833 1438 668 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 O12_IGKJ1*01 833 1438 669 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 O12_IGKJ1*01 833 1438 670 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 O12_IGKJ1*01 833 1438 671 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 O12_IGKJ1*01 833 1438 672 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 O12_IGKJ1*01 833 1438 673 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 O12_IGKJ1*01 833 1438 674 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 O12_IGKJ1*01 833 1438 675 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 O12_IGKJ1*01 833 1438 676 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 O12_IGKJ1*01 833 1438 677 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 O12_IGKJ1*01 833 1438 678 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 O12_IGKJ1*01 833 1438 679 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 O12_IGKJ1*01 833 1438 680 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 O12_IGKJ1*01 833 1438 681 VH3-23_IGHD6-13*01 > 1'_IGHJ6*01 2750 O12_IGKJ1*01 833 1438 682 VH3-23_IGHD6-13*01 > 2'_IGHJ6*01 2751 O12_IGKJ1*01 833 1438 683 VH3-23_IGHD6-13*01 > 3'_IGHJ6*01 2752 O12_IGKJ1*01 833 1438 684 VH3-23_IGHD6-19*01 > 1'_IGHJ6*01 2753 O12_IGKJ1*01 833 1438 685 VH3-23_IGHD6-19*01 > 2'_IGHJ6*01 2754 O12_IGKJ1*01 833 1438 686 VH3-23_IGHD6-19*01 > 3'_IGHJ6*01 2755 O12_IGKJ1*01 833 1438 687 VH3-23_IGHD6-25*01 > 1'_IGHJ6*01 2756 O12_IGKJ1*01 833 1438 688 VH3-23_IGHD6-25*01 > 3'_IGHJ6*01 2757 O12_IGKJ1*01 833 1438 689 VH3-23_IGHD7-27*01 > 1'_IGHJ6*01 2758 O12_IGKJ1*01 833 1438 690 VH3-23_IGHD7-27*01 > 2'_IGHJ6*01 2759 O12_IGKJ1*01 833 1438 691 VH3-23_IGHD1-1*01 > 1_IGHJ1*01 2070 O18_IGKJ1*01 834 1439 692 VH3-23_IGHD1-1*01 > 2_IGHJ1*01 2071 O18_IGKJ1*01 834 1439 693 VH3-23_IGHD1-1*01 > 3_IGHJ1*01 2072 O18_IGKJ1*01 834 1439 694 VH3-23_IGHD1-7*01 > 1_IGHJ1*01 2073 O18_IGKJ1*01 834 1439 695 VH3-23_IGHD1-7*01 > 3_IGHJ1*01 2074 O18_IGKJ1*01 834 1439 696 VH3-23_IGHD1-14*01 > 1_IGHJ1*01 2075 O18_IGKJ1*01 834 1439 697 VH3-23_IGHD1-14*01 > 3_IGHJ1*01 2076 O18_IGKJ1*01 834 1439 698 VH3-23_IGHD1-20*01 > 1_IGHJ1*01 2077 O18_IGKJ1*01 834 1439 699 VH3-23_IGHD1-20*01 > 3_IGHJ1*01 2078 O18_IGKJ1*01 834 1439 700 VH3-23_IGHD1-26*01 > 1_IGHJ1*01 2079 O18_IGKJ1*01 834 1439 701 VH3-23_IGHD1-26*01 > 3_IGHJ1*01 2080 O18_IGKJ1*01 834 1439 702 VH3-23_IGHD2-2*01 > 2_IGHJ1*01 2081 O18_IGKJ1*01 834 1439 703 VH3-23_IGHD2-2*01 > 3_IGHJ1*01 2082 O18_IGKJ1*01 834 1439 704 VH3-23_IGHD2-8*01 > 2_IGHJ1*01 2083 O18_IGKJ1*01 834 1439 705 VH3-23_IGHD2-8*01 > 3_IGHJ1*01 2084 O18_IGKJ1*01 834 1439 706 VH3-23_IGHD2-15*01 > 2_IGHJ1*01 2085 O18_IGKJ1*01 834 1439 707 VH3-23_IGHD2-15*01 > 3_IGHJ1*01 2086 O18_IGKJ1*01 834 1439 708 VH3-23_IGHD2-21*01 > 2_IGHJ1*01 2087 O18_IGKJ1*01 834 1439 709 VH3-23_IGHD2-21*01 > 3_IGHJ1*01 2088 O18_IGKJ1*01 834 1439 710 VH3-23_IGHD3-3*01 > 1_IGHJ1*01 2089 O18_IGKJ1*01 834 1439 711 VH3-23_IGHD3-3*01 > 2_IGHJ1*01 2090 O18_IGKJ1*01 834 1439 712 VH3-23_IGHD3-3*01 > 3_IGHJ1*01 2091 O18_IGKJ1*01 834 1439 713 VH3-23_IGHD3-9*01 > 2_IGHJ1*01 2092 O18_IGKJ1*01 834 1439 714 VH3-23_IGHD3-10*01 > 2_IGHJ1*01 2093 O18_IGKJ1*01 834 1439 715 VH3-23_IGHD3-10*01 > 3_IGHJ1*01 2094 O18_IGKJ1*01 834 1439 716 VH3-23_IGHD3-16*01 > 2_IGHJ1*01 2095 O18_IGKJ1*01 834 1439 717 VH3-23_IGHD3-16*01 > 3_IGHJ1*01 2096 O18_IGKJ1*01 834 1439 718 VH3-23_IGHD3-22*01 > 2_IGHJ1*01 2097 O18_IGKJ1*01 834 1439 719 VH3-23_IGHD3-22*01 > 3_IGHJ1*01 2098 O18_IGKJ1*01 834 1439 720 VH3-23_IGHD4-4*01 (1) > 2_IGHJ1*01 2099 O18_IGKJ1*01 834 1439 721 VH3-23_IGHD4-4*01 (1) > 3_IGHJ1*01 2100 O18_IGKJ1*01 834 1439 722 VH3-23_IGHD4-11*01 (1) > 2_IGHJ1*01 2101 O18_IGKJ1*01 834 1439 723 VH3-23_IGHD4-11*01 (1) > 3_IGHJ1*01 2102 O18_IGKJ1*01 834 1439 724 VH3-23_IGHD4-17*01 > 2_IGHJ1*01 2103 O18_IGKJ1*01 834 1439 725 VH3-23_IGHD4-17*01 > 3_IGHJ1*01 2104 O18_IGKJ1*01 834 1439 726 VH3-23_IGHD4-23*01 > 2_IGHJ1*01 2105 O18_IGKJ1*01 834 1439 727 VH3-23_IGHD4-23*01 > 3_IGHJ1*01 2106 O18_IGKJ1*01 834 1439 728 VH3-23_IGHD5-5*01 (2) > 1_IGHJ1*01 2107 O18_IGKJ1*01 834 1439 729 VH3-23_IGHD5-5*01 (2) > 2_IGHJ1*01 2108 O18_IGKJ1*01 834 1439 730 VH3-23_IGHD5-5*01 (2) > 3_IGHJ1*01 2109 O18_IGKJ1*01 834 1439 731 VH3-23_IGHD5-12*01 > 1_IGHJ1*01 2110 O18_IGKJ1*01 834 1439 732 VH3-23_IGHD5-12*01 > 3_IGHJ1*01 2111 O18_IGKJ1*01 834 1439 733 VH3-23_IGHD5-18*01 (2) > 1_IGHJ1*01 2112 O18_IGKJ1*01 834 1439 734 VH3-23_IGHD5-18*01 (2) > 2_IGHJ1*01 2113 O18_IGKJ1*01 834 1439 735 VH3-23_IGHD5-18*01 (2) > 3_IGHJ1*01 2114 O18_IGKJ1*01 834 1439 736 VH3-23_IGHD5-24*01 > 1_IGHJ1*01 2115 O18_IGKJ1*01 834 1439 737 VH3-23_IGHD5-24*01 > 3_IGHJ1*01 2116 O18_IGKJ1*01 834 1439 738 VH3-23_IGHD6-6*01 > 1_IGHJ1*01 2117 O18_IGKJ1*01 834 1439 739 VH3-23_IGHD1-1*01 > 1'_IGHJ1*01 2127 O18_IGKJ1*01 834 1439 740 VH3-23_IGHD1-1*01 > 2'_IGHJ1*01 2128 O18_IGKJ1*01 834 1439 741 VH3-23_IGHD1-1*01 > 3'_IGHJ1*01 2129 O18_IGKJ1*01 834 1439 742 VH3-23_IGHD1-7*01 > 1'_IGHJ1*01 2130 O18_IGKJ1*01 834 1439 743 VH3-23_IGHD1-7*01 > 3'_IGHJ1*01 2131 O18_IGKJ1*01 834 1439 744 VH3-23_IGHD1-14*01 > 1'_IGHJ1*01 2132 O18_IGKJ1*01 834 1439 745 VH3-23_IGHD1-14*01 > 2'_IGHJ1*01 2133 O18_IGKJ1*01 834 1439
746 VH3-23_IGHD1-14*01 > 3'_IGHJ1*01 2134 O18_IGKJ1*01 834 1439 747 VH3-23_IGHD1-20*01 > 1'_IGHJ1*01 2135 O18_IGKJ1*01 834 1439 748 VH3-23_IGHD1-20*01 > 2'_IGHJ1*01 2136 O18_IGKJ1*01 834 1439 749 VH3-23_IGHD1-20*01 > 3'_IGHJ1*01 2137 O18_IGKJ1*01 834 1439 750 VH3-23_IGHD1-26*01 > 1'_IGHJ1*01 2138 O18_IGKJ1*01 834 1439 751 VH3-23_IGHD1-26*01 > 3'_IGHJ1*01 2139 O18_IGKJ1*01 834 1439 752 VH3-23_IGHD2-2*01 > 1'_IGHJ1*01 2140 O18_IGKJ1*01 834 1439 753 VH3-23_IGHD2-2*01 > 3'_IGHJ1*01 2141 O18_IGKJ1*01 834 1439 754 VH3-23_IGHD2-8*01 > 1'_IGHJ1*01 2142 O18_IGKJ1*01 834 1439 755 VH3-23_IGHD2-15*01 > 1'_IGHJ1*01 2143 O18_IGKJ1*01 834 1439 756 VH3-23_IGHD2-15*01 > 3'_IGHJ1*01 2144 O18_IGKJ1*01 834 1439 757 VH3-23_IGHD2-21*01 > 1'_IGHJ1*01 2145 O18_IGKJ1*01 834 1439 758 VH3-23_IGHD2-21*01 > 3'_IGHJ1*01 2146 O18_IGKJ1*01 834 1439 759 VH3-23_IGHD3-3*01 > 1'_IGHJ1*01 2147 O18_IGKJ1*01 834 1439 760 VH3-23_IGHD3-3*01 > 3'_IGHJ1*01 2148 O18_IGKJ1*01 834 1439 761 VH3-23_IGHD3-9*01 > 1'_IGHJ1*01 2149 O18_IGKJ1*01 834 1439 762 VH3-23_IGHD3-9*01 > 3'_IGHJ1*01 2150 O18_IGKJ1*01 834 1439 763 VH3-23_IGHD3-10*01 > 1'_IGHJ1*01 2151 O18_IGKJ1*01 834 1439 764 VH3-23_IGHD3-10*01 > 3'_IGHJ1*01 2152 O18_IGKJ1*01 834 1439 765 VH3-23_IGHD3-16*01 > 1'_IGHJ1*01 2153 O18_IGKJ1*01 834 1439 766 VH3-23_IGHD3-16*01 > 3'_IGHJ1*01 2154 O18_IGKJ1*01 834 1439 767 VH3-23_IGHD3-22*01 > 1'_IGHJ1*01 2155 O18_IGKJ1*01 834 1439 768 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ1*01 2156 O18_IGKJ1*01 834 1439 769 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ1*01 2157 O18_IGKJ1*01 834 1439 770 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ1*01 2158 O18_IGKJ1*01 834 1439 771 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ1*01 2159 O18_IGKJ1*01 834 1439 772 VH3-23_IGHD4-17*01 > 1'_IGHJ1*01 2160 O18_IGKJ1*01 834 1439 773 VH3-23_IGHD4-17*01 > 3'_IGHJ1*01 2161 O18_IGKJ1*01 834 1439 774 VH3-23_IGHD4-23*01 > 1'_IGHJ1*01 2162 O18_IGKJ1*01 834 1439 775 VH3-23_IGHD4-23*01 > 3'_IGHJ1*01 2163 O18_IGKJ1*01 834 1439 776 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ1*01 2164 O18_IGKJ1*01 834 1439 777 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ1*01 2165 O18_IGKJ1*01 834 1439 778 VH3-23_IGHD5-12*01 > 1'_IGHJ1*01 2166 O18_IGKJ1*01 834 1439 779 VH3-23_IGHD5-12*01 > 3'_IGHJ1*01 2167 O18_IGKJ1*01 834 1439 780 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ1*01 2168 O18_IGKJ1*01 834 1439 781 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ1*01 2169 O18_IGKJ1*01 834 1439 782 VH3-23_IGHD5-24*01 > 1'_IGHJ1*01 2170 O18_IGKJ1*01 834 1439 783 VH3-23_IGHD5-24*01 > 3'_IGHJ1*01 2171 O18_IGKJ1*01 834 1439 784 VH3-23_IGHD6-6*01 > 1'_IGHJ1*01 2172 O18_IGKJ1*01 834 1439 785 VH3-23_IGHD6-6*01 > 2'_IGHJ1*01 2173 O18_IGKJ1*01 834 1439 786 VH3-23_IGHD6-6*01 > 3'_IGHJ1*01 2174 O18_IGKJ1*01 834 1439 787 VH3-23_IGHD7-27*01 > 1'_IGHJ6*01 2758 O18_IGKJ1*01 834 1439 788 VH3-23_IGHD6-13*01 > 2_IGHJ6*01 2695 O18_IGKJ1*01 834 1439 789 VH3-23_IGHD6-19*01 > 1_IGHJ6*01 2696 O18_IGKJ1*01 834 1439 790 VH3-23_IGHD6-19*01 > 2_IGHJ6*01 2697 O18_IGKJ1*01 834 1439 791 VH3-23_IGHD6-25*01 > 1_IGHJ6*01 2698 O18_IGKJ1*01 834 1439 792 VH3-23_IGHD6-25*01 > 2_IGHJ6*01 2699 O18_IGKJ1*01 834 1439 793 VH3-23_IGHD7-27*01 > 1_IGHJ6*01 2700 O18_IGKJ1*01 834 1439 794 VH3-23_IGHD7-27*01 > 3_IGHJ6*01 2701 O18_IGKJ1*01 834 1439 795 VH3-23_IGHD6-13*01 > 1'_IGHJ6*01 2750 O18_IGKJ1*01 834 1439 796 VH3-23_IGHD6-13*01 > 2'_IGHJ6*01 2751 O18_IGKJ1*01 834 1439 797 VH3-23_IGHD6-13*01 > 2_IGHJ6*01_B 2695 O18_IGKJ1*01 834 1439 798 VH3-23_IGHD6-19*01 > 1'_IGHJ6*01 2753 O18_IGKJ1*01 834 1439 799 VH3-23_IGHD6-19*01 > 2'_IGHJ6*01 2754 O18_IGKJ1*01 834 1439 800 VH3-23_IGHD6-25*01 > 1_IGHJ6*01_B 2698 O18_IGKJ1*01 834 1439 801 VH3-23_IGHD6-25*01 > 1'_IGHJ6*01 2756 O18_IGKJ1*01 834 1439 802 VH3-23_IGHD6-25*01 > 3'_IGHJ6*01 2757 O18_IGKJ1*01 834 1439 803 VH3-23_IGHD7-27*01 > 1'_IGHJ6*01 2758 O18_IGKJ1*01 834 1439 804 VH3-23_IGHD7-27*01 > 2'_IGHJ6*01 2759 O18_IGKJ1*01 834 1439 805 VH3-23_IGHD7-27*01 > 1'_IGHJ6*01 2758 A20_IGKJ1*01 809 1414 806 VH3-23_IGHD6-13*01 > 2_IGHJ6*01 2695 A20_IGKJ1*01 809 1414 807 VH3-23_IGHD6-19*01 > 1_IGHJ6*01 2696 A20_IGKJ1*01 809 1414 808 VH3-23_IGHD6-19*01 > 2_IGHJ6*01 2697 A20_IGKJ1*01 809 1414 809 VH3-23_IGHD6-25*01 > 1_IGHJ6*01 2698 A20_IGKJ1*01 809 1414 810 VH3-23_IGHD6-25*01 > 2_IGHJ6*01 2699 A20_IGKJ1*01 809 1414 811 VH3-23_IGHD7-27*01 > 1_IGHJ6*01 2700 A20_IGKJ1*01 809 1414 812 VH3-23_IGHD7-27*01 > 3_IGHJ6*01 2701 A20_IGKJ1*01 809 1414 813 VH3-23_IGHD6-13*01 > 1'_IGHJ6*01 2750 A20_IGKJ1*01 809 1414 814 VH3-23_IGHD6-13*01 > 2'_IGHJ6*01 2751 A20_IGKJ1*01 809 1414 815 VH3-23_IGHD6-13*01 > 2_IGHJ6*01_B 2695 A20_IGKJ1*01 809 1414 816 VH3-23_IGHD6-19*01 > 1'_IGHJ6*01 2753 A20_IGKJ1*01 809 1414 817 VH3-23_IGHD6-19*01 > 2'_IGHJ6*01 2754 A20_IGKJ1*01 809 1414 818 VH3-23_IGHD6-25*01 > 1_IGHJ6*01_B 2698 A20_IGKJ1*01 809 1414 819 VH3-23_IGHD6-25*01 > 1'_IGHJ6*01 2756 A20_IGKJ1*01 809 1414 820 VH3-23_IGHD6-25*01 > 3'_IGHJ6*01 2757 A20_IGKJ1*01 809 1414 821 VH3-23_IGHD7-27*01 > 1'_IGHJ6*01 2758 A20_IGKJ1*01 809 1414 822 VH3-23_IGHD7-27*01 > 2'_IGHJ6*01 2759 A20_IGKJ1*01 809 1414 823 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 L11_IGKJ1*01 819 1424 824 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 L11_IGKJ1*01 819 1424 825 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 L11_IGKJ1*01 819 1424 826 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 L11_IGKJ1*01 819 1424 827 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 L11_IGKJ1*01 819 1424 828 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 L11_IGKJ1*01 819 1424 829 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 L11_IGKJ1*01 819 1424 830 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 L11_IGKJ1*01 819 1424 831 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 L11_IGKJ1*01 819 1424 832 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 L11_IGKJ1*01 819 1424 833 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 L11_IGKJ1*01 819 1424 834 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 L11_IGKJ1*01 819 1424 835 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 L11_IGKJ1*01 819 1424 836 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 L11_IGKJ1*01 819 1424 837 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 L11_IGKJ1*01 819 1424 838 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 L11_IGKJ1*01 819 1424 839 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 L11_IGKJ1*01 819 1424 840 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 L11_IGKJ1*01 819 1424 841 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 L11_IGKJ1*01 819 1424 842 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 L11_IGKJ1*01 819 1424 843 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 L11_IGKJ1*01 819 1424 844 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 L11_IGKJ1*01 819 1424 845 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 L11_IGKJ1*01 819 1424 846 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 L11_IGKJ1*01 819 1424 847 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 L11_IGKJ1*01 819 1424 848 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 L11_IGKJ1*01 819 1424 849 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 L11_IGKJ1*01 819 1424 850 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 L11_IGKJ1*01 819 1424 851 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 L11_IGKJ1*01 819 1424 852 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 L11_IGKJ1*01 819 1424 853 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 L11_IGKJ1*01 819 1424 854 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 L11_IGKJ1*01 819 1424 855 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 L11_IGKJ1*01 819 1424 856 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 L11_IGKJ1*01 819 1424 857 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 L11_IGKJ1*01 819 1424 858 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 L11_IGKJ1*01 819 1424 859 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 L11_IGKJ1*01 819 1424 860 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 L11_IGKJ1*01 819 1424 861 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 L11_IGKJ1*01 819 1424 862 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 L11_IGKJ1*01 819 1424 863 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 L11_IGKJ1*01 819 1424 864 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 L11_IGKJ1*01 819 1424 865 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 L11_IGKJ1*01 819 1424 866 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 L11_IGKJ1*01 819 1424 867 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 L11_IGKJ1*01 819 1424 868 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 L11_IGKJ1*01 819 1424 869 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 L11_IGKJ1*01 819 1424 870 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 L11_IGKJ1*01 819 1424 871 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 L11_IGKJ1*01 819 1424 872 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 L11_IGKJ1*01 819 1424 873 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 L11_IGKJ1*01 819 1424 874 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 L11_IGKJ1*01 819 1424 875 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 L11_IGKJ1*01 819 1424 876 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 L11_IGKJ1*01 819 1424 877 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 L11_IGKJ1*01 819 1424 878 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 L11_IGKJ1*01 819 1424 879 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 L11_IGKJ1*01 819 1424 880 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 L11_IGKJ1*01 819 1424 881 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 L11_IGKJ1*01 819 1424 882 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 L11_IGKJ1*01 819 1424 883 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 L11_IGKJ1*01 819 1424 884 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 L11_IGKJ1*01 819 1424 885 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 L11_IGKJ1*01 819 1424 886 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 L11_IGKJ1*01 819 1424 887 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 L11_IGKJ1*01 819 1424 888 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 L11_IGKJ1*01 819 1424 889 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 L11_IGKJ1*01 819 1424 890 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 L11_IGKJ1*01 819 1424 891 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 L11_IGKJ1*01 819 1424 892 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 L11_IGKJ1*01 819 1424 893 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 L11_IGKJ1*01 819 1424 894 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 L11_IGKJ1*01 819 1424 895 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 L11_IGKJ1*01 819 1424 896 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 L11_IGKJ1*01 819 1424 897 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 L11_IGKJ1*01 819 1424 898 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 L11_IGKJ1*01 819 1424 899 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 L11_IGKJ1*01 819 1424 900 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 L11_IGKJ1*01 819 1424 901 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 L11_IGKJ1*01 819 1424 902 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 L11_IGKJ1*01 819 1424 903 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 L11_IGKJ1*01 819 1424 904 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 L11_IGKJ1*01 819 1424 905 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 L11_IGKJ1*01 819 1424 906 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 L11_IGKJ1*01 819 1424 907 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 L11_IGKJ1*01 819 1424 908 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 L11_IGKJ1*01 819 1424 909 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 L11_IGKJ1*01 819 1424 910 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 L11_IGKJ1*01 819 1424 911 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 L11_IGKJ1*01 819 1424 912 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 L11_IGKJ1*01 819 1424 913 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 L11_IGKJ1*01 819 1424 914 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 L11_IGKJ1*01 819 1424 915 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 L11_IGKJ1*01 819 1424 916 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 L11_IGKJ1*01 819 1424 917 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 L11_IGKJ1*01 819 1424 918 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 L11_IGKJ1*01 819 1424 919 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 L12_IGKJ1*01 820 1425 920 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 L12_IGKJ1*01 820 1425 921 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 L12_IGKJ1*01 820 1425 922 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 L12_IGKJ1*01 820 1425 923 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 L12_IGKJ1*01 820 1425 924 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 L12_IGKJ1*01 820 1425 925 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 L12_IGKJ1*01 820 1425 926 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 L12_IGKJ1*01 820 1425 927 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 L12_IGKJ1*01 820 1425 928 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 L12_IGKJ1*01 820 1425 929 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 L12_IGKJ1*01 820 1425 930 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 L12_IGKJ1*01 820 1425 931 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 L12_IGKJ1*01 820 1425 932 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 L12_IGKJ1*01 820 1425 933 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 L12_IGKJ1*01 820 1425 934 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 L12_IGKJ1*01 820 1425 935 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 L12_IGKJ1*01 820 1425 936 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 L12_IGKJ1*01 820 1425 937 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 L12_IGKJ1*01 820 1425 938 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 L12_IGKJ1*01 820 1425 939 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 L12_IGKJ1*01 820 1425 940 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 L12_IGKJ1*01 820 1425 941 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 L12_IGKJ1*01 820 1425 942 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 L12_IGKJ1*01 820 1425 943 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 L12_IGKJ1*01 820 1425 944 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 L12_IGKJ1*01 820 1425 945 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 L12_IGKJ1*01 820 1425 946 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 L12_IGKJ1*01 820 1425 947 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 L12_IGKJ1*01 820 1425 948 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 L12_IGKJ1*01 820 1425 949 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 L12_IGKJ1*01 820 1425 950 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 L12_IGKJ1*01 820 1425 951 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 L12_IGKJ1*01 820 1425 952 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 L12_IGKJ1*01 820 1425 953 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 L12_IGKJ1*01 820 1425 954 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 L12_IGKJ1*01 820 1425 955 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 L12_IGKJ1*01 820 1425 956 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 L12_IGKJ1*01 820 1425 957 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 L12_IGKJ1*01 820 1425 958 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 L12_IGKJ1*01 820 1425 959 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 L12_IGKJ1*01 820 1425 960 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 L12_IGKJ1*01 820 1425 961 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 L12_IGKJ1*01 820 1425 962 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 L12_IGKJ1*01 820 1425 963 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 L12_IGKJ1*01 820 1425 964 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 L12_IGKJ1*01 820 1425 965 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 L12_IGKJ1*01 820 1425 966 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 L12_IGKJ1*01 820 1425 967 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 L12_IGKJ1*01 820 1425 968 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 L12_IGKJ1*01 820 1425 969 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 L12_IGKJ1*01 820 1425 970 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 L12_IGKJ1*01 820 1425 971 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 L12_IGKJ1*01 820 1425 972 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 L12_IGKJ1*01 820 1425 973 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 L12_IGKJ1*01 820 1425 974 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 L12_IGKJ1*01 820 1425 975 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 L12_IGKJ1*01 820 1425 976 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 L12_IGKJ1*01 820 1425 977 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 L12_IGKJ1*01 820 1425 978 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 L12_IGKJ1*01 820 1425 979 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 L12_IGKJ1*01 820 1425 980 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 L12_IGKJ1*01 820 1425 981 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 L12_IGKJ1*01 820 1425 982 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 L12_IGKJ1*01 820 1425 983 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 L12_IGKJ1*01 820 1425 984 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 L12_IGKJ1*01 820 1425 985 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 L12_IGKJ1*01 820 1425 986 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 L12_IGKJ1*01 820 1425 987 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 L12_IGKJ1*01 820 1425 988 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 L12_IGKJ1*01 820 1425 989 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 L12_IGKJ1*01 820 1425 990 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 L12_IGKJ1*01 820 1425 991 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 L12_IGKJ1*01 820 1425 992 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 L12_IGKJ1*01 820 1425 993 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 L12_IGKJ1*01 820 1425 994 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 L12_IGKJ1*01 820 1425 995 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 L12_IGKJ1*01 820 1425 996 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 L12_IGKJ1*01 820 1425
997 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 L12_IGKJ1*01 820 1425 998 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 L12_IGKJ1*01 820 1425 999 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 L12_IGKJ1*01 820 1425 1000 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 L12_IGKJ1*01 820 1425 1001 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 L12_IGKJ1*01 820 1425 1002 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 L12_IGKJ1*01 820 1425 1003 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 L12_IGKJ1*01 820 1425 1004 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 L12_IGKJ1*01 820 1425 1005 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 L12_IGKJ1*01 820 1425 1006 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 L12_IGKJ1*01 820 1425 1007 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 L12_IGKJ1*01 820 1425 1008 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 L12_IGKJ1*01 820 1425 1009 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 L12_IGKJ1*01 820 1425 1010 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 L12_IGKJ1*01 820 1425 1011 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 L12_IGKJ1*01 820 1425 1012 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 L12_IGKJ1*01 820 1425 1013 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 L12_IGKJ1*01 820 1425 1014 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 L12_IGKJ1*01 820 1425 1015 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 O1_IGKJ1*01 832 1437 1016 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 O1_IGKJ1*01 832 1437 1017 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 O1_IGKJ1*01 832 1437 1018 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 O1_IGKJ1*01 832 1437 1019 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 O1_IGKJ1*01 832 1437 1020 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 O1_IGKJ1*01 832 1437 1021 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 O1_IGKJ1*01 832 1437 1022 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 O1_IGKJ1*01 832 1437 1023 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 O1_IGKJ1*01 832 1437 1024 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 O1_IGKJ1*01 832 1437 1025 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 O1_IGKJ1*01 832 1437 1026 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 O1_IGKJ1*01 832 1437 1027 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 O1_IGKJ1*01 832 1437 1028 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 O1_IGKJ1*01 832 1437 1029 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 O1_IGKJ1*01 832 1437 1030 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 O1_IGKJ1*01 832 1437 1031 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 O1_IGKJ1*01 832 1437 1032 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 O1_IGKJ1*01 832 1437 1033 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 O1_IGKJ1*01 832 1437 1034 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 O1_IGKJ1*01 832 1437 1035 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 O1_IGKJ1*01 832 1437 1036 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 O1_IGKJ1*01 832 1437 1037 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 O1_IGKJ1*01 832 1437 1038 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 O1_IGKJ1*01 832 1437 1039 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 O1_IGKJ1*01 832 1437 1040 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 O1_IGKJ1*01 832 1437 1041 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 O1_IGKJ1*01 832 1437 1042 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 O1_IGKJ1*01 832 1437 1043 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 O1_IGKJ1*01 832 1437 1044 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 O1_IGKJ1*01 832 1437 1045 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 O1_IGKJ1*01 832 1437 1046 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 O1_IGKJ1*01 832 1437 1047 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 O1_IGKJ1*01 832 1437 1048 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 O1_IGKJ1*01 832 1437 1049 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 O1_IGKJ1*01 832 1437 1050 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 O1_IGKJ1*01 832 1437 1051 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 O1_IGKJ1*01 832 1437 1052 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 O1_IGKJ1*01 832 1437 1053 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 O1_IGKJ1*01 832 1437 1054 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 O1_IGKJ1*01 832 1437 1055 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 O1_IGKJ1*01 832 1437 1056 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 O1_IGKJ1*01 832 1437 1057 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 O1_IGKJ1*01 832 1437 1058 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 O1_IGKJ1*01 832 1437 1059 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 O1_IGKJ1*01 832 1437 1060 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 O1_IGKJ1*01 832 1437 1061 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 O1_IGKJ1*01 832 1437 1062 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 O1_IGKJ1*01 832 1437 1063 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 O1_IGKJ1*01 832 1437 1064 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 O1_IGKJ1*01 832 1437 1065 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 O1_IGKJ1*01 832 1437 1066 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 O1_IGKJ1*01 832 1437 1067 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 O1_IGKJ1*01 832 1437 1068 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 O1_IGKJ1*01 832 1437 1069 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 O1_IGKJ1*01 832 1437 1070 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 O1_IGKJ1*01 832 1437 1071 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 O1_IGKJ1*01 832 1437 1072 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 O1_IGKJ1*01 832 1437 1073 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 O1_IGKJ1*01 832 1437 1074 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 O1_IGKJ1*01 832 1437 1075 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 O1_IGKJ1*01 832 1437 1076 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 O1_IGKJ1*01 832 1437 1077 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 O1_IGKJ1*01 832 1437 1078 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 O1_IGKJ1*01 832 1437 1079 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 O1_IGKJ1*01 832 1437 1080 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 O1_IGKJ1*01 832 1437 1081 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 O1_IGKJ1*01 832 1437 1082 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 O1_IGKJ1*01 832 1437 1083 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 O1_IGKJ1*01 832 1437 1084 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 O1_IGKJ1*01 832 1437 1085 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 O1_IGKJ1*01 832 1437 1086 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 O1_IGKJ1*01 832 1437 1087 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 O1_IGKJ1*01 832 1437 1088 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 O1_IGKJ1*01 832 1437 1089 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 O1_IGKJ1*01 832 1437 1090 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 O1_IGKJ1*01 832 1437 1091 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 O1_IGKJ1*01 832 1437 1092 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 O1_IGKJ1*01 832 1437 1093 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 O1_IGKJ1*01 832 1437 1094 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 O1_IGKJ1*01 832 1437 1095 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 O1_IGKJ1*01 832 1437 1096 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 O1_IGKJ1*01 832 1437 1097 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 O1_IGKJ1*01 832 1437 1098 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 O1_IGKJ1*01 832 1437 1099 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 O1_IGKJ1*01 832 1437 1100 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 O1_IGKJ1*01 832 1437 1101 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 O1_IGKJ1*01 832 1437 1102 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 O1_IGKJ1*01 832 1437 1103 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 O1_IGKJ1*01 832 1437 1104 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 O1_IGKJ1*01 832 1437 1105 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 O1_IGKJ1*01 832 1437 1106 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 O1_IGKJ1*01 832 1437 1107 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 O1_IGKJ1*01 832 1437 1108 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 O1_IGKJ1*01 832 1437 1109 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 O1_IGKJ1*01 832 1437 1110 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 O1_IGKJ1*01 832 1437 1111 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 A2_IGKJ1*01 808 1413 1112 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 A2_IGKJ1*01 808 1413 1113 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 A2_IGKJ1*01 808 1413 1114 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 A2_IGKJ1*01 808 1413 1115 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 A2_IGKJ1*01 808 1413 1116 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 A2_IGKJ1*01 808 1413 1117 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 A2_IGKJ1*01 808 1413 1118 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 A2_IGKJ1*01 808 1413 1119 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 A2_IGKJ1*01 808 1413 1120 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 A2_IGKJ1*01 808 1413 1121 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 A2_IGKJ1*01 808 1413 1122 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 A2_IGKJ1*01 808 1413 1123 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 A2_IGKJ1*01 808 1413 1124 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 A2_IGKJ1*01 808 1413 1125 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 A2_IGKJ1*01 808 1413 1126 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 A2_IGKJ1*01 808 1413 1127 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 A2_IGKJ1*01 808 1413 1128 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 A2_IGKJ1*01 808 1413 1129 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 A2_IGKJ1*01 808 1413 1130 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 A2_IGKJ1*01 808 1413 1131 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 A2_IGKJ1*01 808 1413 1132 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 A2_IGKJ1*01 808 1413 1133 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 A2_IGKJ1*01 808 1413 1134 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 A2_IGKJ1*01 808 1413 1135 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 A2_IGKJ1*01 808 1413 1136 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 A2_IGKJ1*01 808 1413 1137 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 A2_IGKJ1*01 808 1413 1138 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 A2_IGKJ1*01 808 1413 1139 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 A2_IGKJ1*01 808 1413 1140 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 A2_IGKJ1*01 808 1413 1141 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 A2_IGKJ1*01 808 1413 1142 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 A2_IGKJ1*01 808 1413 1143 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 A2_IGKJ1*01 808 1413 1144 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 A2_IGKJ1*01 808 1413 1145 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 A2_IGKJ1*01 808 1413 1146 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 A2_IGKJ1*01 808 1413 1147 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 A2_IGKJ1*01 808 1413 1148 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 A2_IGKJ1*01 808 1413 1149 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 A2_IGKJ1*01 808 1413 1150 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 A2_IGKJ1*01 808 1413 1151 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 A2_IGKJ1*01 808 1413 1152 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 A2_IGKJ1*01 808 1413 1153 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 A2_IGKJ1*01 808 1413 1154 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 A2_IGKJ1*01 808 1413 1155 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 A2_IGKJ1*01 808 1413 1156 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 A2_IGKJ1*01 808 1413 1157 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 A2_IGKJ1*01 808 1413 1158 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 A2_IGKJ1*01 808 1413 1159 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 A2_IGKJ1*01 808 1413 1160 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 A2_IGKJ1*01 808 1413 1161 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 A2_IGKJ1*01 808 1413 1162 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 A2_IGKJ1*01 808 1413 1163 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 A2_IGKJ1*01 808 1413 1164 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 A2_IGKJ1*01 808 1413 1165 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 A2_IGKJ1*01 808 1413 1166 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 A2_IGKJ1*01 808 1413 1167 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 A2_IGKJ1*01 808 1413 1168 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 A2_IGKJ1*01 808 1413 1169 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 A2_IGKJ1*01 808 1413 1170 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 A2_IGKJ1*01 808 1413 1171 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 A2_IGKJ1*01 808 1413 1172 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 A2_IGKJ1*01 808 1413 1173 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 A2_IGKJ1*01 808 1413 1174 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 A2_IGKJ1*01 808 1413 1175 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 A2_IGKJ1*01 808 1413 1176 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 A2_IGKJ1*01 808 1413 1177 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 A2_IGKJ1*01 808 1413 1178 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 A2_IGKJ1*01 808 1413 1179 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 A2_IGKJ1*01 808 1413 1180 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 A2_IGKJ1*01 808 1413 1181 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 A2_IGKJ1*01 808 1413 1182 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 A2_IGKJ1*01 808 1413 1183 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 A2_IGKJ1*01 808 1413 1184 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 A2_IGKJ1*01 808 1413 1185 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 A2_IGKJ1*01 808 1413 1186 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 A2_IGKJ1*01 808 1413 1187 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 A2_IGKJ1*01 808 1413 1188 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 A2_IGKJ1*01 808 1413 1189 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 A2_IGKJ1*01 808 1413 1190 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 A2_IGKJ1*01 808 1413 1191 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 A2_IGKJ1*01 808 1413 1192 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 A2_IGKJ1*01 808 1413 1193 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 A2_IGKJ1*01 808 1413 1194 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 A2_IGKJ1*01 808 1413 1195 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 A2_IGKJ1*01 808 1413 1196 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 A2_IGKJ1*01 808 1413 1197 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 A2_IGKJ1*01 808 1413 1198 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 A2_IGKJ1*01 808 1413 1199 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 A2_IGKJ1*01 808 1413 1200 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 A2_IGKJ1*01 808 1413 1201 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 A2_IGKJ1*01 808 1413 1202 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 A2_IGKJ1*01 808 1413 1203 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 A2_IGKJ1*01 808 1413 1204 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 A2_IGKJ1*01 808 1413 1205 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 A2_IGKJ1*01 808 1413 1206 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 A2_IGKJ1*01 808 1413 1207 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 L2_IGKJ1*01 822 1427 1208 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 L2_IGKJ1*01 822 1427 1209 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 L2_IGKJ1*01 822 1427 1210 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 L2_IGKJ1*01 822 1427 1211 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 L2_IGKJ1*01 822 1427 1212 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 L2_IGKJ1*01 822 1427 1213 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 L2_IGKJ1*01 822 1427 1214 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 L2_IGKJ1*01 822 1427 1215 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 L2_IGKJ1*01 822 1427 1216 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 L2_IGKJ1*01 822 1427 1217 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 L2_IGKJ1*01 822 1427 1218 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 L2_IGKJ1*01 822 1427 1219 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 L2_IGKJ1*01 822 1427 1220 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 L2_IGKJ1*01 822 1427 1221 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 L2_IGKJ1*01 822 1427 1222 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 L2_IGKJ1*01 822 1427 1223 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 L2_IGKJ1*01 822 1427 1224 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 L2_IGKJ1*01 822 1427 1225 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 L2_IGKJ1*01 822 1427 1226 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 L2_IGKJ1*01 822 1427 1227 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 L2_IGKJ1*01 822 1427 1228 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 L2_IGKJ1*01 822 1427 1229 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 L2_IGKJ1*01 822 1427 1230 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 L2_IGKJ1*01 822 1427 1231 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 L2_IGKJ1*01 822 1427 1232 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 L2_IGKJ1*01 822 1427 1233 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 L2_IGKJ1*01 822 1427 1234 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 L2_IGKJ1*01 822 1427 1235 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 L2_IGKJ1*01 822 1427 1236 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 L2_IGKJ1*01 822 1427 1237 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 L2_IGKJ1*01 822 1427 1238 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 L2_IGKJ1*01 822 1427 1239 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 L2_IGKJ1*01 822 1427 1240 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 L2_IGKJ1*01 822 1427 1241 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 L2_IGKJ1*01 822 1427 1242 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 L2_IGKJ1*01 822 1427 1243 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 L2_IGKJ1*01 822 1427 1244 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 L2_IGKJ1*01 822 1427 1245 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 L2_IGKJ1*01 822 1427 1246 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 L2_IGKJ1*01 822 1427 1247 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 L2_IGKJ1*01 822 1427
1248 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 L2_IGKJ1*01 822 1427 1249 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 L2_IGKJ1*01 822 1427 1250 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 L2_IGKJ1*01 822 1427 1251 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 L2_IGKJ1*01 822 1427 1252 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 L2_IGKJ1*01 822 1427 1253 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 L2_IGKJ1*01 822 1427 1254 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 L2_IGKJ1*01 822 1427 1255 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 L2_IGKJ1*01 822 1427 1256 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 L2_IGKJ1*01 822 1427 1257 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 L2_IGKJ1*01 822 1427 1258 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 L2_IGKJ1*01 822 1427 1259 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 L2_IGKJ1*01 822 1427 1260 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 L2_IGKJ1*01 822 1427 1261 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 L2_IGKJ1*01 822 1427 1262 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 L2_IGKJ1*01 822 1427 1263 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 L2_IGKJ1*01 822 1427 1264 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 L2_IGKJ1*01 822 1427 1265 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 L2_IGKJ1*01 822 1427 1266 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 L2_IGKJ1*01 822 1427 1267 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 L2_IGKJ1*01 822 1427 1268 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 L2_IGKJ1*01 822 1427 1269 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 L2_IGKJ1*01 822 1427 1270 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 L2_IGKJ1*01 822 1427 1271 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 L2_IGKJ1*01 822 1427 1272 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 L2_IGKJ1*01 822 1427 1273 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 L2_IGKJ1*01 822 1427 1274 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 L2_IGKJ1*01 822 1427 1275 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 L2_IGKJ1*01 822 1427 1276 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 L2_IGKJ1*01 822 1427 1277 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 L2_IGKJ1*01 822 1427 1278 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 L2_IGKJ1*01 822 1427 1279 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 L2_IGKJ1*01 822 1427 1280 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 L2_IGKJ1*01 822 1427 1281 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 L2_IGKJ1*01 822 1427 1282 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 L2_IGKJ1*01 822 1427 1283 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 L2_IGKJ1*01 822 1427 1284 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 L2_IGKJ1*01 822 1427 1285 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 L2_IGKJ1*01 822 1427 1286 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 L2_IGKJ1*01 822 1427 1287 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 L2_IGKJ1*01 822 1427 1288 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 L2_IGKJ1*01 822 1427 1289 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 L2_IGKJ1*01 822 1427 1290 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 L2_IGKJ1*01 822 1427 1291 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 L2_IGKJ1*01 822 1427 1292 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 L2_IGKJ1*01 822 1427 1293 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 L2_IGKJ1*01 822 1427 1294 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 L2_IGKJ1*01 822 1427 1295 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 L2_IGKJ1*01 822 1427 1296 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 L2_IGKJ1*01 822 1427 1297 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 L2_IGKJ1*01 822 1427 1298 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 L2_IGKJ1*01 822 1427 1299 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 L2_IGKJ1*01 822 1427 1300 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 L2_IGKJ1*01 822 1427 1301 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 L2_IGKJ1*01 822 1427 1302 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 L2_IGKJ1*01 822 1427 1303 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 L6_IGKJ1*01 829 1434 1304 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 L6_IGKJ1*01 829 1434 1305 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 L6_IGKJ1*01 829 1434 1306 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 L6_IGKJ1*01 829 1434 1307 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 L6_IGKJ1*01 829 1434 1308 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 L6_IGKJ1*01 829 1434 1309 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 L6_IGKJ1*01 829 1434 1310 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 L6_IGKJ1*01 829 1434 1311 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 L6_IGKJ1*01 829 1434 1312 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 L6_IGKJ1*01 829 1434 1313 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 L6_IGKJ1*01 829 1434 1314 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 L6_IGKJ1*01 829 1434 1315 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 L6_IGKJ1*01 829 1434 1316 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 L6_IGKJ1*01 829 1434 1317 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 L6_IGKJ1*01 829 1434 1318 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 L6_IGKJ1*01 829 1434 1319 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 L6_IGKJ1*01 829 1434 1320 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 L6_IGKJ1*01 829 1434 1321 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 L6_IGKJ1*01 829 1434 1322 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 L6_IGKJ1*01 829 1434 1323 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 L6_IGKJ1*01 829 1434 1324 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 L6_IGKJ1*01 829 1434 1325 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 L6_IGKJ1*01 829 1434 1326 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 L6_IGKJ1*01 829 1434 1327 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 L6_IGKJ1*01 829 1434 1328 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 L6_IGKJ1*01 829 1434 1329 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 L6_IGKJ1*01 829 1434 1330 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 L6_IGKJ1*01 829 1434 1331 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 L6_IGKJ1*01 829 1434 1332 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 L6_IGKJ1*01 829 1434 1333 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 L6_IGKJ1*01 829 1434 1334 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 L6_IGKJ1*01 829 1434 1335 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 L6_IGKJ1*01 829 1434 1336 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 L6_IGKJ1*01 829 1434 1337 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 L6_IGKJ1*01 829 1434 1338 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 L6_IGKJ1*01 829 1434 1339 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 L6_IGKJ1*01 829 1434 1340 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 L6_IGKJ1*01 829 1434 1341 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 L6_IGKJ1*01 829 1434 1342 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 L6_IGKJ1*01 829 1434 1343 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 L6_IGKJ1*01 829 1434 1344 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 L6_IGKJ1*01 829 1434 1345 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 L6_IGKJ1*01 829 1434 1346 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 L6_IGKJ1*01 829 1434 1347 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 L6_IGKJ1*01 829 1434 1348 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 L6_IGKJ1*01 829 1434 1349 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 L6_IGKJ1*01 829 1434 1350 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 L6_IGKJ1*01 829 1434 1351 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 L6_IGKJ1*01 829 1434 1352 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 L6_IGKJ1*01 829 1434 1353 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 L6_IGKJ1*01 829 1434 1354 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 L6_IGKJ1*01 829 1434 1355 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 L6_IGKJ1*01 829 1434 1356 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 L6_IGKJ1*01 829 1434 1357 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 L6_IGKJ1*01 829 1434 1358 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 L6_IGKJ1*01 829 1434 1359 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 L6_IGKJ1*01 829 1434 1360 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 L6_IGKJ1*01 829 1434 1361 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 L6_IGKJ1*01 829 1434 1362 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 L6_IGKJ1*01 829 1434 1363 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 L6_IGKJ1*01 829 1434 1364 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 L6_IGKJ1*01 829 1434 1365 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 L6_IGKJ1*01 829 1434 1366 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 L6_IGKJ1*01 829 1434 1367 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 L6_IGKJ1*01 829 1434 1368 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 L6_IGKJ1*01 829 1434 1369 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 L6_IGKJ1*01 829 1434 1370 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 L6_IGKJ1*01 829 1434 1371 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 L6_IGKJ1*01 829 1434 1372 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 L6_IGKJ1*01 829 1434 1373 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 L6_IGKJ1*01 829 1434 1374 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 L6_IGKJ1*01 829 1434 1375 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 L6_IGKJ1*01 829 1434 1376 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 L6_IGKJ1*01 829 1434 1377 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 L6_IGKJ1*01 829 1434 1378 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 L6_IGKJ1*01 829 1434 1379 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 L6_IGKJ1*01 829 1434 1380 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 L6_IGKJ1*01 829 1434 1381 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 L6_IGKJ1*01 829 1434 1382 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 L6_IGKJ1*01 829 1434 1383 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 L6_IGKJ1*01 829 1434 1384 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 L6_IGKJ1*01 829 1434 1385 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 L6_IGKJ1*01 829 1434 1386 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 L6_IGKJ1*01 829 1434 1387 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 L6_IGKJ1*01 829 1434 1388 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 L6_IGKJ1*01 829 1434 1389 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 L6_IGKJ1*01 829 1434 1390 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 L6_IGKJ1*01 829 1434 1391 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 L6_IGKJ1*01 829 1434 1392 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 L6_IGKJ1*01 829 1434 1393 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 L6_IGKJ1*01 829 1434 1394 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 L6_IGKJ1*01 829 1434 1395 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 L6_IGKJ1*01 829 1434 1396 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 L6_IGKJ1*01 829 1434 1397 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 L6_IGKJ1*01 829 1434 1398 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 L6_IGKJ1*01 829 1434 1399 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 L25_IGKJ1*01 826 1431 1400 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 L25_IGKJ1*01 826 1431 1401 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 L25_IGKJ1*01 826 1431 1402 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 L25_IGKJ1*01 826 1431 1403 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 L25_IGKJ1*01 826 1431 1404 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 L25_IGKJ1*01 826 1431 1405 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 L25_IGKJ1*01 826 1431 1406 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 L25_IGKJ1*01 826 1431 1407 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 L25_IGKJ1*01 826 1431 1408 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 L25_IGKJ1*01 826 1431 1409 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 L25_IGKJ1*01 826 1431 1410 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 L25_IGKJ1*01 826 1431 1411 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 L25_IGKJ1*01 826 1431 1412 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 L25_IGKJ1*01 826 1431 1413 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 L25_IGKJ1*01 826 1431 1414 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 L25_IGKJ1*01 826 1431 1415 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 L25_IGKJ1*01 826 1431 1416 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 L25_IGKJ1*01 826 1431 1417 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 L25_IGKJ1*01 826 1431 1418 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 L25_IGKJ1*01 826 1431 1419 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 L25_IGKJ1*01 826 1431 1420 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 L25_IGKJ1*01 826 1431 1421 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 L25_IGKJ1*01 826 1431 1422 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 L25_IGKJ1*01 826 1431 1423 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 L25_IGKJ1*01 826 1431 1424 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 L25_IGKJ1*01 826 1431 1425 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 L25_IGKJ1*01 826 1431 1426 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 L25_IGKJ1*01 826 1431 1427 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 L25_IGKJ1*01 826 1431 1428 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 L25_IGKJ1*01 826 1431 1429 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 L25_IGKJ1*01 826 1431 1430 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 L25_IGKJ1*01 826 1431 1431 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 L25_IGKJ1*01 826 1431 1432 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 L25_IGKJ1*01 826 1431 1433 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 L25_IGKJ1*01 826 1431 1434 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 L25_IGKJ1*01 826 1431 1435 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 L25_IGKJ1*01 826 1431 1436 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 L25_IGKJ1*01 826 1431 1437 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 L25_IGKJ1*01 826 1431 1438 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 L25_IGKJ1*01 826 1431 1439 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 L25_IGKJ1*01 826 1431 1440 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 L25_IGKJ1*01 826 1431 1441 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 L25_IGKJ1*01 826 1431 1442 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 L25_IGKJ1*01 826 1431 1443 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 L25_IGKJ1*01 826 1431 1444 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 L25_IGKJ1*01 826 1431 1445 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 L25_IGKJ1*01 826 1431 1446 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 L25_IGKJ1*01 826 1431 1447 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 L25_IGKJ1*01 826 1431 1448 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 L25_IGKJ1*01 826 1431 1449 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 L25_IGKJ1*01 826 1431 1450 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 L25_IGKJ1*01 826 1431 1451 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 L25_IGKJ1*01 826 1431 1452 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 L25_IGKJ1*01 826 1431 1453 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 L25_IGKJ1*01 826 1431 1454 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 L25_IGKJ1*01 826 1431 1455 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 L25_IGKJ1*01 826 1431 1456 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 L25_IGKJ1*01 826 1431 1457 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 L25_IGKJ1*01 826 1431 1458 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 L25_IGKJ1*01 826 1431 1459 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 L25_IGKJ1*01 826 1431 1460 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 L25_IGKJ1*01 826 1431 1461 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 L25_IGKJ1*01 826 1431 1462 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 L25_IGKJ1*01 826 1431 1463 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 L25_IGKJ1*01 826 1431 1464 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 L25_IGKJ1*01 826 1431 1465 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 L25_IGKJ1*01 826 1431 1466 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 L25_IGKJ1*01 826 1431 1467 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 L25_IGKJ1*01 826 1431 1468 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 L25_IGKJ1*01 826 1431 1469 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 L25_IGKJ1*01 826 1431 1470 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 L25_IGKJ1*01 826 1431 1471 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 L25_IGKJ1*01 826 1431 1472 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 L25_IGKJ1*01 826 1431 1473 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 L25_IGKJ1*01 826 1431 1474 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 L25_IGKJ1*01 826 1431 1475 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 L25_IGKJ1*01 826 1431 1476 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 L25_IGKJ1*01 826 1431 1477 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 L25_IGKJ1*01 826 1431 1478 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 L25_IGKJ1*01 826 1431 1479 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 L25_IGKJ1*01 826 1431 1480 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 L25_IGKJ1*01 826 1431 1481 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 L25_IGKJ1*01 826 1431 1482 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 L25_IGKJ1*01 826 1431 1483 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 L25_IGKJ1*01 826 1431 1484 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 L25_IGKJ1*01 826 1431 1485 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 L25_IGKJ1*01 826 1431 1486 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 L25_IGKJ1*01 826 1431 1487 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 L25_IGKJ1*01 826 1431 1488 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 L25_IGKJ1*01 826 1431 1489 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 L25_IGKJ1*01 826 1431 1490 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 L25_IGKJ1*01 826 1431 1491 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 L25_IGKJ1*01 826 1431 1492 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 L25_IGKJ1*01 826 1431 1493 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 L25_IGKJ1*01 826 1431 1494 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 L25_IGKJ1*01 826 1431 1495 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 B3_IGKJ1*01 817 1422 1496 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 B3_IGKJ1*01 817 1422 1497 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 B3_IGKJ1*01 817 1422 1498 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 B3_IGKJ1*01 817 1422
1499 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 B3_IGKJ1*01 817 1422 1500 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 B3_IGKJ1*01 817 1422 1501 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 B3_IGKJ1*01 817 1422 1502 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 B3_IGKJ1*01 817 1422 1503 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 B3_IGKJ1*01 817 1422 1504 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 B3_IGKJ1*01 817 1422 1505 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 B3_IGKJ1*01 817 1422 1506 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 B3_IGKJ1*01 817 1422 1507 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 B3_IGKJ1*01 817 1422 1508 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 B3_IGKJ1*01 817 1422 1509 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 B3_IGKJ1*01 817 1422 1510 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 B3_IGKJ1*01 817 1422 1511 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 B3_IGKJ1*01 817 1422 1512 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 B3_IGKJ1*01 817 1422 1513 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 B3_IGKJ1*01 817 1422 1514 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 B3_IGKJ1*01 817 1422 1515 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 B3_IGKJ1*01 817 1422 1516 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 B3_IGKJ1*01 817 1422 1517 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 B3_IGKJ1*01 817 1422 1518 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 B3_IGKJ1*01 817 1422 1519 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 B3_IGKJ1*01 817 1422 1520 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 B3_IGKJ1*01 817 1422 1521 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 B3_IGKJ1*01 817 1422 1522 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 B3_IGKJ1*01 817 1422 1523 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 B3_IGKJ1*01 817 1422 1524 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 B3_IGKJ1*01 817 1422 1525 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 B3_IGKJ1*01 817 1422 1526 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 B3_IGKJ1*01 817 1422 1527 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 B3_IGKJ1*01 817 1422 1528 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 B3_IGKJ1*01 817 1422 1529 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 B3_IGKJ1*01 817 1422 1530 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 B3_IGKJ1*01 817 1422 1531 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 B3_IGKJ1*01 817 1422 1532 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 B3_IGKJ1*01 817 1422 1533 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 B3_IGKJ1*01 817 1422 1534 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 B3_IGKJ1*01 817 1422 1535 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 B3_IGKJ1*01 817 1422 1536 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 B3_IGKJ1*01 817 1422 1537 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 B3_IGKJ1*01 817 1422 1538 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 B3_IGKJ1*01 817 1422 1539 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 B3_IGKJ1*01 817 1422 1540 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 B3_IGKJ1*01 817 1422 1541 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 B3_IGKJ1*01 817 1422 1542 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 B3_IGKJ1*01 817 1422 1543 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 B3_IGKJ1*01 817 1422 1544 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 B3_IGKJ1*01 817 1422 1545 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 B3_IGKJ1*01 817 1422 1546 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 B3_IGKJ1*01 817 1422 1547 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 B3_IGKJ1*01 817 1422 1548 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 B3_IGKJ1*01 817 1422 1549 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 B3_IGKJ1*01 817 1422 1550 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 B3_IGKJ1*01 817 1422 1551 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 B3_IGKJ1*01 817 1422 1552 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 B3_IGKJ1*01 817 1422 1553 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 B3_IGKJ1*01 817 1422 1554 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 B3_IGKJ1*01 817 1422 1555 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 B3_IGKJ1*01 817 1422 1556 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 B3_IGKJ1*01 817 1422 1557 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 B3_IGKJ1*01 817 1422 1558 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 B3_IGKJ1*01 817 1422 1559 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 B3_IGKJ1*01 817 1422 1560 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 B3_IGKJ1*01 817 1422 1561 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 B3_IGKJ1*01 817 1422 1562 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 B3_IGKJ1*01 817 1422 1563 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 B3_IGKJ1*01 817 1422 1564 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 B3_IGKJ1*01 817 1422 1565 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 B3_IGKJ1*01 817 1422 1566 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 B3_IGKJ1*01 817 1422 1567 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 B3_IGKJ1*01 817 1422 1568 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 B3_IGKJ1*01 817 1422 1569 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 B3_IGKJ1*01 817 1422 1570 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 B3_IGKJ1*01 817 1422 1571 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 B3_IGKJ1*01 817 1422 1572 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 B3_IGKJ1*01 817 1422 1573 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 B3_IGKJ1*01 817 1422 1574 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 B3_IGKJ1*01 817 1422 1575 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 B3_IGKJ1*01 817 1422 1576 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 B3_IGKJ1*01 817 1422 1577 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 B3_IGKJ1*01 817 1422 1578 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 B3_IGKJ1*01 817 1422 1579 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 B3_IGKJ1*01 817 1422 1580 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 B3_IGKJ1*01 817 1422 1581 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 B3_IGKJ1*01 817 1422 1582 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 B3_IGKJ1*01 817 1422 1583 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 B3_IGKJ1*01 817 1422 1584 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 B3_IGKJ1*01 817 1422 1585 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 B3_IGKJ1*01 817 1422 1586 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 B3_IGKJ1*01 817 1422 1587 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 B3_IGKJ1*01 817 1422 1588 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 B3_IGKJ1*01 817 1422 1589 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 B3_IGKJ1*01 817 1422 1590 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 B3_IGKJ1*01 817 1422 1591 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 A26_IGKJ1*01 811 1416 1592 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 A26_IGKJ1*01 811 1416 1593 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 A26_IGKJ1*01 811 1416 1594 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 A26_IGKJ1*01 811 1416 1595 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 A26_IGKJ1*01 811 1416 1596 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 A26_IGKJ1*01 811 1416 1597 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 A26_IGKJ1*01 811 1416 1598 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 A26_IGKJ1*01 811 1416 1599 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 A26_IGKJ1*01 811 1416 1600 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 A26_IGKJ1*01 811 1416 1601 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 A26_IGKJ1*01 811 1416 1602 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 A26_IGKJ1*01 811 1416 1603 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 A26_IGKJ1*01 811 1416 1604 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 A26_IGKJ1*01 811 1416 1605 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 A26_IGKJ1*01 811 1416 1606 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 A26_IGKJ1*01 811 1416 1607 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 A26_IGKJ1*01 811 1416 1608 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 A26_IGKJ1*01 811 1416 1609 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 A26_IGKJ1*01 811 1416 1610 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 A26_IGKJ1*01 811 1416 1611 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 A26_IGKJ1*01 811 1416 1612 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 A26_IGKJ1*01 811 1416 1613 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 A26_IGKJ1*01 811 1416 1614 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 A26_IGKJ1*01 811 1416 1615 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 A26_IGKJ1*01 811 1416 1616 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 A26_IGKJ1*01 811 1416 1617 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 A26_IGKJ1*01 811 1416 1618 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 A26_IGKJ1*01 811 1416 1619 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 A26_IGKJ1*01 811 1416 1620 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 A26_IGKJ1*01 811 1416 1621 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 A26_IGKJ1*01 811 1416 1622 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 A26_IGKJ1*01 811 1416 1623 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 A26_IGKJ1*01 811 1416 1624 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 A26_IGKJ1*01 811 1416 1625 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 A26_IGKJ1*01 811 1416 1626 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 A26_IGKJ1*01 811 1416 1627 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 A26_IGKJ1*01 811 1416 1628 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 A26_IGKJ1*01 811 1416 1629 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 A26_IGKJ1*01 811 1416 1630 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 A26_IGKJ1*01 811 1416 1631 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 A26_IGKJ1*01 811 1416 1632 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 A26_IGKJ1*01 811 1416 1633 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 A26_IGKJ1*01 811 1416 1634 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 A26_IGKJ1*01 811 1416 1635 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 A26_IGKJ1*01 811 1416 1636 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 A26_IGKJ1*01 811 1416 1637 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 A26_IGKJ1*01 811 1416 1638 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 A26_IGKJ1*01 811 1416 1639 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 A26_IGKJ1*01 811 1416 1640 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 A26_IGKJ1*01 811 1416 1641 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 A26_IGKJ1*01 811 1416 1642 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 A26_IGKJ1*01 811 1416 1643 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 A26_IGKJ1*01 811 1416 1644 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 A26_IGKJ1*01 811 1416 1645 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 A26_IGKJ1*01 811 1416 1646 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 A26_IGKJ1*01 811 1416 1647 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 A26_IGKJ1*01 811 1416 1648 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 A26_IGKJ1*01 811 1416 1649 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 A26_IGKJ1*01 811 1416 1650 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 A26_IGKJ1*01 811 1416 1651 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 A26_IGKJ1*01 811 1416 1652 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 A26_IGKJ1*01 811 1416 1653 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 A26_IGKJ1*01 811 1416 1654 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 A26_IGKJ1*01 811 1416 1655 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 A26_IGKJ1*01 811 1416 1656 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 A26_IGKJ1*01 811 1416 1657 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 A26_IGKJ1*01 811 1416 1658 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 A26_IGKJ1*01 811 1416 1659 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 A26_IGKJ1*01 811 1416 1660 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 A26_IGKJ1*01 811 1416 1661 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 A26_IGKJ1*01 811 1416 1662 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 A26_IGKJ1*01 811 1416 1663 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 A26_IGKJ1*01 811 1416 1664 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 A26_IGKJ1*01 811 1416 1665 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 A26_IGKJ1*01 811 1416 1666 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 A26_IGKJ1*01 811 1416 1667 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 A26_IGKJ1*01 811 1416 1668 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 A26_IGKJ1*01 811 1416 1669 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 A26_IGKJ1*01 811 1416 1670 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 A26_IGKJ1*01 811 1416 1671 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 A26_IGKJ1*01 811 1416 1672 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 A26_IGKJ1*01 811 1416 1673 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 A26_IGKJ1*01 811 1416 1674 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 A26_IGKJ1*01 811 1416 1675 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 A26_IGKJ1*01 811 1416 1676 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 A26_IGKJ1*01 811 1416 1677 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 A26_IGKJ1*01 811 1416 1678 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 A26_IGKJ1*01 811 1416 1679 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 A26_IGKJ1*01 811 1416 1680 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 A26_IGKJ1*01 811 1416 1681 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 A26_IGKJ1*01 811 1416 1682 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 A26_IGKJ1*01 811 1416 1683 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 A26_IGKJ1*01 811 1416 1684 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 A26_IGKJ1*01 811 1416 1685 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 A26_IGKJ1*01 811 1416 1686 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 A26_IGKJ1*01 811 1416 1687 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 A14_IGKJ1*01 806 1411 1688 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 A14_IGKJ1*01 806 1411 1689 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 A14_IGKJ1*01 806 1411 1690 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 A14_IGKJ1*01 806 1411 1691 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 A14_IGKJ1*01 806 1411 1692 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 A14_IGKJ1*01 806 1411 1693 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 A14_IGKJ1*01 806 1411 1694 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 A14_IGKJ1*01 806 1411 1695 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 A14_IGKJ1*01 806 1411 1696 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 A14_IGKJ1*01 806 1411 1697 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 A14_IGKJ1*01 806 1411 1698 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 A14_IGKJ1*01 806 1411 1699 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 A14_IGKJ1*01 806 1411 1700 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 A14_IGKJ1*01 806 1411 1701 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 A14_IGKJ1*01 806 1411 1702 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 A14_IGKJ1*01 806 1411 1703 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 A14_IGKJ1*01 806 1411 1704 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 A14_IGKJ1*01 806 1411 1705 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 A14_IGKJ1*01 806 1411 1706 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 A14_IGKJ1*01 806 1411 1707 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 A14_IGKJ1*01 806 1411 1708 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 A14_IGKJ1*01 806 1411 1709 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 A14_IGKJ1*01 806 1411 1710 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 A14_IGKJ1*01 806 1411 1711 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 A14_IGKJ1*01 806 1411 1712 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 A14_IGKJ1*01 806 1411 1713 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 A14_IGKJ1*01 806 1411 1714 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 A14_IGKJ1*01 806 1411 1715 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 A14_IGKJ1*01 806 1411 1716 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 A14_IGKJ1*01 806 1411 1717 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 A14_IGKJ1*01 806 1411 1718 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 A14_IGKJ1*01 806 1411 1719 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 A14_IGKJ1*01 806 1411 1720 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 A14_IGKJ1*01 806 1411 1721 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 A14_IGKJ1*01 806 1411 1722 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 A14_IGKJ1*01 806 1411 1723 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 A14_IGKJ1*01 806 1411 1724 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 A14_IGKJ1*01 806 1411 1725 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 A14_IGKJ1*01 806 1411 1726 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 A14_IGKJ1*01 806 1411 1727 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 A14_IGKJ1*01 806 1411 1728 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 A14_IGKJ1*01 806 1411 1729 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 A14_IGKJ1*01 806 1411 1730 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 A14_IGKJ1*01 806 1411 1731 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 A14_IGKJ1*01 806 1411 1732 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 A14_IGKJ1*01 806 1411 1733 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 A14_IGKJ1*01 806 1411 1734 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 A14_IGKJ1*01 806 1411 1735 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 A14_IGKJ1*01 806 1411 1736 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 A14_IGKJ1*01 806 1411 1737 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 A14_IGKJ1*01 806 1411 1738 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 A14_IGKJ1*01 806 1411 1739 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 A14_IGKJ1*01 806 1411 1740 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 A14_IGKJ1*01 806 1411 1741 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 A14_IGKJ1*01 806 1411 1742 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 A14_IGKJ1*01 806 1411 1743 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 A14_IGKJ1*01 806 1411 1744 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 A14_IGKJ1*01 806 1411 1745 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 A14_IGKJ1*01 806 1411 1746 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 A14_IGKJ1*01 806 1411 1747 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 A14_IGKJ1*01 806 1411 1748 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 A14_IGKJ1*01 806 1411 1749 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 A14_IGKJ1*01 806 1411
1750 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 A14_IGKJ1*01 806 1411 1751 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 A14_IGKJ1*01 806 1411 1752 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 A14_IGKJ1*01 806 1411 1753 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 A14_IGKJ1*01 806 1411 1754 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 A14_IGKJ1*01 806 1411 1755 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 A14_IGKJ1*01 806 1411 1756 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 A14_IGKJ1*01 806 1411 1757 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 A14_IGKJ1*01 806 1411 1758 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 A14_IGKJ1*01 806 1411 1759 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 A14_IGKJ1*01 806 1411 1760 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 A14_IGKJ1*01 806 1411 1761 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 A14_IGKJ1*01 806 1411 1762 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 A14_IGKJ1*01 806 1411 1763 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 A14_IGKJ1*01 806 1411 1764 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 A14_IGKJ1*01 806 1411 1765 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 A14_IGKJ1*01 806 1411 1766 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 A14_IGKJ1*01 806 1411 1767 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 A14_IGKJ1*01 806 1411 1768 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 A14_IGKJ1*01 806 1411 1769 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 A14_IGKJ1*01 806 1411 1770 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 A14_IGKJ1*01 806 1411 1771 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 A14_IGKJ1*01 806 1411 1772 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 A14_IGKJ1*01 806 1411 1773 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 A14_IGKJ1*01 806 1411 1774 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 A14_IGKJ1*01 806 1411 1775 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 A14_IGKJ1*01 806 1411 1776 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 A14_IGKJ1*01 806 1411 1777 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 A14_IGKJ1*01 806 1411 1778 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 A14_IGKJ1*01 806 1411 1779 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 A14_IGKJ1*01 806 1411 1780 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 A14_IGKJ1*01 806 1411 1781 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 A14_IGKJ1*01 806 1411 1782 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 A14_IGKJ1*01 806 1411 1783 VH3-23_IGHD1-1*01 > 1_IGHJ5*01 2530 A27_IGKJ1*01 812 1417 1784 VH3-23_IGHD1-1*01 > 2_IGHJ5*01 2531 A27_IGKJ1*01 812 1417 1785 VH3-23_IGHD1-1*01 > 3_IGHJ5*01 2532 A27_IGKJ1*01 812 1417 1786 VH3-23_IGHD1-7*01 > 1_IGHJ5*01 2533 A27_IGKJ1*01 812 1417 1787 VH3-23_IGHD1-7*01 > 3_IGHJ5*01 2534 A27_IGKJ1*01 812 1417 1788 VH3-23_IGHD1-14*01 > 1_IGHJ5*01 2535 A27_IGKJ1*01 812 1417 1789 VH3-23_IGHD1-14*01 > 3_IGHJ5*01 2536 A27_IGKJ1*01 812 1417 1790 VH3-23_IGHD1-20*01 > 1_IGHJ5*01 2537 A27_IGKJ1*01 812 1417 1791 VH3-23_IGHD1-20*01 > 3_IGHJ5*01 2538 A27_IGKJ1*01 812 1417 1792 VH3-23_IGHD1-26*01 > 1_IGHJ5*01 2539 A27_IGKJ1*01 812 1417 1793 VH3-23_IGHD1-26*01 > 3_IGHJ5*01 2540 A27_IGKJ1*01 812 1417 1794 VH3-23_IGHD2-2*01 > 2_IGHJ5*01 2541 A27_IGKJ1*01 812 1417 1795 VH3-23_IGHD2-2*01 > 3_IGHJ5*01 2542 A27_IGKJ1*01 812 1417 1796 VH3-23_IGHD2-8*01 > 2_IGHJ5*01 2543 A27_IGKJ1*01 812 1417 1797 VH3-23_IGHD2-8*01 > 3_IGHJ5*01 2544 A27_IGKJ1*01 812 1417 1798 VH3-23_IGHD2-15*01 > 2_IGHJ5*01 2545 A27_IGKJ1*01 812 1417 1799 VH3-23_IGHD2-15*01 > 3_IGHJ5*01 2546 A27_IGKJ1*01 812 1417 1800 VH3-23_IGHD2-21*01 > 2_IGHJ5*01 2547 A27_IGKJ1*01 812 1417 1801 VH3-23_IGHD2-21*01 > 3_IGHJ5*01 2548 A27_IGKJ1*01 812 1417 1802 VH3-23_IGHD3-3*01 > 1_IGHJ5*01 2549 A27_IGKJ1*01 812 1417 1803 VH3-23_IGHD3-3*01 > 2_IGHJ5*01 2550 A27_IGKJ1*01 812 1417 1804 VH3-23_IGHD3-3*01 > 3_IGHJ5*01 2551 A27_IGKJ1*01 812 1417 1805 VH3-23_IGHD3-9*01 > 2_IGHJ5*01 2552 A27_IGKJ1*01 812 1417 1806 VH3-23_IGHD3-10*01 > 2_IGHJ5*01 2553 A27_IGKJ1*01 812 1417 1807 VH3-23_IGHD3-10*01 > 3_IGHJ5*01 2554 A27_IGKJ1*01 812 1417 1808 VH3-23_IGHD3-16*01 > 2_IGHJ5*01 2555 A27_IGKJ1*01 812 1417 1809 VH3-23_IGHD3-16*01 > 3_IGHJ5*01 2556 A27_IGKJ1*01 812 1417 1810 VH3-23_IGHD3-22*01 > 2_IGHJ5*01 2557 A27_IGKJ1*01 812 1417 1811 VH3-23_IGHD3-22*01 > 3_IGHJ5*01 2558 A27_IGKJ1*01 812 1417 1812 VH3-23_IGHD4-4*01 (1) > 2_IGHJ5*01 2559 A27_IGKJ1*01 812 1417 1813 VH3-23_IGHD4-4*01 (1) > 3_IGHJ5*01 2560 A27_IGKJ1*01 812 1417 1814 VH3-23_IGHD4-11*01 (1) > 2_IGHJ5*01 2561 A27_IGKJ1*01 812 1417 1815 VH3-23_IGHD4-11*01 (1) > 3_IGHJ5*01 2562 A27_IGKJ1*01 812 1417 1816 VH3-23_IGHD4-17*01 > 2_IGHJ5*01 2563 A27_IGKJ1*01 812 1417 1817 VH3-23_IGHD4-17*01 > 3_IGHJ5*01 2564 A27_IGKJ1*01 812 1417 1818 VH3-23_IGHD4-23*01 > 2_IGHJ5*01 2565 A27_IGKJ1*01 812 1417 1819 VH3-23_IGHD4-23*01 > 3_IGHJ5*01 2566 A27_IGKJ1*01 812 1417 1820 VH3-23_IGHD5-5*01 (2) > 1_IGHJ5*01 2567 A27_IGKJ1*01 812 1417 1821 VH3-23_IGHD5-5*01 (2) > 2_IGHJ5*01 2568 A27_IGKJ1*01 812 1417 1822 VH3-23_IGHD5-5*01 (2) > 3_IGHJ5*01 2569 A27_IGKJ1*01 812 1417 1823 VH3-23_IGHD5-12*01 > 1_IGHJ5*01 2570 A27_IGKJ1*01 812 1417 1824 VH3-23_IGHD5-12*01 > 3_IGHJ5*01 2571 A27_IGKJ1*01 812 1417 1825 VH3-23_IGHD5-18*01 (2) > 1_IGHJ5*01 2572 A27_IGKJ1*01 812 1417 1826 VH3-23_IGHD5-18*01 (2) > 2_IGHJ5*01 2573 A27_IGKJ1*01 812 1417 1827 VH3-23_IGHD5-18*01 (2) > 3_IGHJ5*01 2574 A27_IGKJ1*01 812 1417 1828 VH3-23_IGHD5-24*01 > 1_IGHJ5*01 2575 A27_IGKJ1*01 812 1417 1829 VH3-23_IGHD5-24*01 > 3_IGHJ5*01 2576 A27_IGKJ1*01 812 1417 1830 VH3-23_IGHD6-6*01 > 1_IGHJ5*01 2577 A27_IGKJ1*01 812 1417 1831 VH3-23_IGHD1-1*01 > 1'_IGHJ5*01 2587 A27_IGKJ1*01 812 1417 1832 VH3-23_IGHD1-1*01 > 2'_IGHJ5*01 2588 A27_IGKJ1*01 812 1417 1833 VH3-23_IGHD1-1*01 > 3'_IGHJ5*01 2589 A27_IGKJ1*01 812 1417 1834 VH3-23_IGHD1-7*01 > 1'_IGHJ5*01 2590 A27_IGKJ1*01 812 1417 1835 VH3-23_IGHD1-7*01 > 3'_IGHJ5*01 2591 A27_IGKJ1*01 812 1417 1836 VH3-23_IGHD1-14*01 > 1'_IGHJ5*01 2592 A27_IGKJ1*01 812 1417 1837 VH3-23_IGHD1-14*01 > 2'_IGHJ5*01 2593 A27_IGKJ1*01 812 1417 1838 VH3-23_IGHD1-14*01 > 3'_IGHJ5*01 2594 A27_IGKJ1*01 812 1417 1839 VH3-23_IGHD1-20*01 > 1'_IGHJ5*01 2595 A27_IGKJ1*01 812 1417 1840 VH3-23_IGHD1-20*01 > 2'_IGHJ5*01 2596 A27_IGKJ1*01 812 1417 1841 VH3-23_IGHD1-20*01 > 3'_IGHJ5*01 2597 A27_IGKJ1*01 812 1417 1842 VH3-23_IGHD1-26*01 > 1'_IGHJ5*01 2598 A27_IGKJ1*01 812 1417 1843 VH3-23_IGHD1-26*01 > 3'_IGHJ5*01 2599 A27_IGKJ1*01 812 1417 1844 VH3-23_IGHD2-2*01 > 1'_IGHJ5*01 2600 A27_IGKJ1*01 812 1417 1845 VH3-23_IGHD2-2*01 > 3'_IGHJ5*01 2601 A27_IGKJ1*01 812 1417 1846 VH3-23_IGHD2-8*01 > 1'_IGHJ5*01 2602 A27_IGKJ1*01 812 1417 1847 VH3-23_IGHD2-15*01 > 1'_IGHJ5*01 2603 A27_IGKJ1*01 812 1417 1848 VH3-23_IGHD2-15*01 > 3'_IGHJ5*01 2604 A27_IGKJ1*01 812 1417 1849 VH3-23_IGHD2-21*01 > 1'_IGHJ5*01 2605 A27_IGKJ1*01 812 1417 1850 VH3-23_IGHD2-21*01 > 3'_IGHJ5*01 2606 A27_IGKJ1*01 812 1417 1851 VH3-23_IGHD3-3*01 > 1'_IGHJ5*01 2607 A27_IGKJ1*01 812 1417 1852 VH3-23_IGHD3-3*01 > 3'_IGHJ5*01 2608 A27_IGKJ1*01 812 1417 1853 VH3-23_IGHD3-9*01 > 1'_IGHJ5*01 2609 A27_IGKJ1*01 812 1417 1854 VH3-23_IGHD3-9*01 > 3'_IGHJ5*01 2610 A27_IGKJ1*01 812 1417 1855 VH3-23_IGHD3-10*01 > 1'_IGHJ5*01 2611 A27_IGKJ1*01 812 1417 1856 VH3-23_IGHD3-10*01 > 3'_IGHJ5*01 2612 A27_IGKJ1*01 812 1417 1857 VH3-23_IGHD3-16*01 > 1'_IGHJ5*01 2613 A27_IGKJ1*01 812 1417 1858 VH3-23_IGHD3-16*01 > 3'_IGHJ5*01 2614 A27_IGKJ1*01 812 1417 1859 VH3-23_IGHD3-22*01 > 1'_IGHJ5*01 2615 A27_IGKJ1*01 812 1417 1860 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ5*01 2616 A27_IGKJ1*01 812 1417 1861 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ5*01 2617 A27_IGKJ1*01 812 1417 1862 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ5*01 2618 A27_IGKJ1*01 812 1417 1863 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ5*01 2619 A27_IGKJ1*01 812 1417 1864 VH3-23_IGHD4-17*01 > 1'_IGHJ5*01 2620 A27_IGKJ1*01 812 1417 1865 VH3-23_IGHD4-17*01 > 3'_IGHJ5*01 2621 A27_IGKJ1*01 812 1417 1866 VH3-23_IGHD4-23*01 > 1'_IGHJ5*01 2622 A27_IGKJ1*01 812 1417 1867 VH3-23_IGHD4-23*01 > 3'_IGHJ5*01 2623 A27_IGKJ1*01 812 1417 1868 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ5*01 2624 A27_IGKJ1*01 812 1417 1869 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ5*01 2625 A27_IGKJ1*01 812 1417 1870 VH3-23_IGHD5-12*01 > 1'_IGHJ5*01 2626 A27_IGKJ1*01 812 1417 1871 VH3-23_IGHD5-12*01 > 3'_IGHJ5*01 2627 A27_IGKJ1*01 812 1417 1872 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ5*01 2628 A27_IGKJ1*01 812 1417 1873 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ5*01 2629 A27_IGKJ1*01 812 1417 1874 VH3-23_IGHD5-24*01 > 1'_IGHJ5*01 2630 A27_IGKJ1*01 812 1417 1875 VH3-23_IGHD5-24*01 > 3'_IGHJ5*01 2631 A27_IGKJ1*01 812 1417 1876 VH3-23_IGHD6-6*01 > 1'_IGHJ5*01 2632 A27_IGKJ1*01 812 1417 1877 VH3-23_IGHD6-6*01 > 2'_IGHJ5*01 2633 A27_IGKJ1*01 812 1417 1878 VH3-23_IGHD6-6*01 > 3'_IGHJ5*01 2634 A27_IGKJ1*01 812 1417 1879 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 V1-11_IGLJ2*01 836 1441 1880 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 V1-11_IGLJ2*01 836 1441 1881 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 V1-11_IGLJ2*01 836 1441 1882 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 V1-11_IGLJ2*01 836 1441 1883 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 V1-11_IGLJ2*01 836 1441 1884 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 V1-11_IGLJ2*01 836 1441 1885 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 V1-11_IGLJ2*01 836 1441 1886 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 V1-11_IGLJ2*01 836 1441 1887 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 V1-11_IGLJ2*01 836 1441 1888 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 V1-11_IGLJ2*01 836 1441 1889 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 V1-11_IGLJ2*01 836 1441 1890 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 V1-11_IGLJ2*01 836 1441 1891 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 V1-11_IGLJ2*01 836 1441 1892 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 V1-11_IGLJ2*01 836 1441 1893 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 V1-11_IGLJ2*01 836 1441 1894 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 V1-11_IGLJ2*01 836 1441 1895 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 V1-11_IGLJ2*01 836 1441 1896 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 V1-11_IGLJ2*01 836 1441 1897 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 V1-11_IGLJ2*01 836 1441 1898 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 V1-11_IGLJ2*01 836 1441 1899 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 V1-11_IGLJ2*01 836 1441 1900 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 V1-11_IGLJ2*01 836 1441 1901 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 V1-11_IGLJ2*01 836 1441 1902 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 V1-11_IGLJ2*01 836 1441 1903 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 V1-11_IGLJ2*01 836 1441 1904 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 V1-11_IGLJ2*01 836 1441 1905 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 V1-11_IGLJ2*01 836 1441 1906 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 V1-11_IGLJ2*01 836 1441 1907 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 V1-11_IGLJ2*01 836 1441 1908 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 V1-11_IGLJ2*01 836 1441 1909 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 V1-11_IGLJ2*01 836 1441 1910 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 V1-11_IGLJ2*01 836 1441 1911 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 V1-11_IGLJ2*01 836 1441 1912 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 V1-11_IGLJ2*01 836 1441 1913 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 V1-11_IGLJ2*01 836 1441 1914 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 V1-11_IGLJ2*01 836 1441 1915 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 V1-11_IGLJ2*01 836 1441 1916 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 V1-11_IGLJ2*01 836 1441 1917 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 V1-11_IGLJ2*01 836 1441 1918 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 V1-11_IGLJ2*01 836 1441 1919 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 V1-11_IGLJ2*01 836 1441 1920 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 V1-11_IGLJ2*01 836 1441 1921 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 V1-11_IGLJ2*01 836 1441 1922 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 V1-11_IGLJ2*01 836 1441 1923 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 V1-11_IGLJ2*01 836 1441 1924 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 V1-11_IGLJ2*01 836 1441 1925 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 V1-11_IGLJ2*01 836 1441 1926 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 V1-11_IGLJ2*01 836 1441 1927 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 V1-11_IGLJ2*01 836 1441 1928 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 V1-11_IGLJ2*01 836 1441 1929 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 V1-11_IGLJ2*01 836 1441 1930 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 V1-11_IGLJ2*01 836 1441 1931 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 V1-11_IGLJ2*01 836 1441 1932 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 V1-11_IGLJ2*01 836 1441 1933 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 V1-11_IGLJ2*01 836 1441 1934 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 V1-11_IGLJ2*01 836 1441 1935 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 V1-11_IGLJ2*01 836 1441 1936 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 V1-11_IGLJ2*01 836 1441 1937 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 V1-11_IGLJ2*01 836 1441 1938 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 V1-11_IGLJ2*01 836 1441 1939 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 V1-11_IGLJ2*01 836 1441 1940 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 V1-11_IGLJ2*01 836 1441 1941 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 V1-11_IGLJ2*01 836 1441 1942 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 V1-11_IGLJ2*01 836 1441 1943 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 V1-11_IGLJ2*01 836 1441 1944 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 V1-11_IGLJ2*01 836 1441 1945 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 V1-11_IGLJ2*01 836 1441 1946 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 V1-11_IGLJ2*01 836 1441 1947 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 V1-11_IGLJ2*01 836 1441 1948 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 V1-11_IGLJ2*01 836 1441 1949 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 V1-11_IGLJ2*01 836 1441 1950 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 V1-11_IGLJ2*01 836 1441 1951 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 V1-11_IGLJ2*01 836 1441 1952 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 V1-11_IGLJ2*01 836 1441 1953 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 V1-11_IGLJ2*01 836 1441 1954 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 V1-11_IGLJ2*01 836 1441 1955 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 V1-11_IGLJ2*01 836 1441 1956 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 V1-11_IGLJ2*01 836 1441 1957 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 V1-11_IGLJ2*01 836 1441 1958 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 V1-11_IGLJ2*01 836 1441 1959 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 V1-11_IGLJ2*01 836 1441 1960 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 V1-11_IGLJ2*01 836 1441 1961 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 V1-11_IGLJ2*01 836 1441 1962 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 V1-11_IGLJ2*01 836 1441 1963 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 V1-11_IGLJ2*01 836 1441 1964 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 V1-11_IGLJ2*01 836 1441 1965 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 V1-11_IGLJ2*01 836 1441 1966 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 V1-11_IGLJ2*01 836 1441 1967 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 V1-11_IGLJ2*01 836 1441 1968 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 V1-11_IGLJ2*01 836 1441 1969 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 V1-11_IGLJ2*01 836 1441 1970 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 V1-11_IGLJ2*01 836 1441 1971 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 V1-11_IGLJ2*01 836 1441 1972 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 V1-11_IGLJ2*01 836 1441 1973 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 V1-11_IGLJ2*01 836 1441 1974 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 V1-11_IGLJ2*01 836 1441 1975 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 V1-13_IGLJ5*01 837 1442 1976 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 V1-13_IGLJ5*01 837 1442 1977 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 V1-13_IGLJ5*01 837 1442 1978 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 V1-13_IGLJ5*01 837 1442 1979 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 V1-13_IGLJ5*01 837 1442 1980 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 V1-13_IGLJ5*01 837 1442 1981 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 V1-13_IGLJ5*01 837 1442 1982 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 V1-13_IGLJ5*01 837 1442 1983 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 V1-13_IGLJ5*01 837 1442 1984 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 V1-13_IGLJ5*01 837 1442 1985 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 V1-13_IGLJ5*01 837 1442 1986 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 V1-13_IGLJ5*01 837 1442 1987 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 V1-13_IGLJ5*01 837 1442 1988 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 V1-13_IGLJ5*01 837 1442 1989 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 V1-13_IGLJ5*01 837 1442 1990 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 V1-13_IGLJ5*01 837 1442 1991 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 V1-13_IGLJ5*01 837 1442 1992 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 V1-13_IGLJ5*01 837 1442 1993 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 V1-13_IGLJ5*01 837 1442 1994 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 V1-13_IGLJ5*01 837 1442 1995 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 V1-13_IGLJ5*01 837 1442 1996 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 V1-13_IGLJ5*01 837 1442 1997 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 V1-13_IGLJ5*01 837 1442 1998 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 V1-13_IGLJ5*01 837 1442 1999 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 V1-13_IGLJ5*01 837 1442 2000 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 V1-13_IGLJ5*01 837 1442
2001 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 V1-13_IGLJ5*01 837 1442 2002 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 V1-13_IGLJ5*01 837 1442 2003 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 V1-13_IGLJ5*01 837 1442 2004 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 V1-13_IGLJ5*01 837 1442 2005 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 V1-13_IGLJ5*01 837 1442 2006 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 V1-13_IGLJ5*01 837 1442 2007 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 V1-13_IGLJ5*01 837 1442 2008 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 V1-13_IGLJ5*01 837 1442 2009 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 V1-13_IGLJ5*01 837 1442 2010 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 V1-13_IGLJ5*01 837 1442 2011 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 V1-13_IGLJ5*01 837 1442 2012 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 V1-13_IGLJ5*01 837 1442 2013 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 V1-13_IGLJ5*01 837 1442 2014 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 V1-13_IGLJ5*01 837 1442 2015 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 V1-13_IGLJ5*01 837 1442 2016 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 V1-13_IGLJ5*01 837 1442 2017 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 V1-13_IGLJ5*01 837 1442 2018 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 V1-13_IGLJ5*01 837 1442 2019 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 V1-13_IGLJ5*01 837 1442 2020 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 V1-13_IGLJ5*01 837 1442 2021 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 V1-13_IGLJ5*01 837 1442 2022 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 V1-13_IGLJ5*01 837 1442 2023 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 V1-13_IGLJ5*01 837 1442 2024 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 V1-13_IGLJ5*01 837 1442 2025 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 V1-13_IGLJ5*01 837 1442 2026 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 V1-13_IGLJ5*01 837 1442 2027 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 V1-13_IGLJ5*01 837 1442 2028 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 V1-13_IGLJ5*01 837 1442 2029 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 V1-13_IGLJ5*01 837 1442 2030 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 V1-13_IGLJ5*01 837 1442 2031 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 V1-13_IGLJ5*01 837 1442 2032 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 V1-13_IGLJ5*01 837 1442 2033 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 V1-13_IGLJ5*01 837 1442 2034 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 V1-13_IGLJ5*01 837 1442 2035 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 V1-13_IGLJ5*01 837 1442 2036 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 V1-13_IGLJ5*01 837 1442 2037 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 V1-13_IGLJ5*01 837 1442 2038 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 V1-13_IGLJ5*01 837 1442 2039 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 V1-13_IGLJ5*01 837 1442 2040 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 V1-13_IGLJ5*01 837 1442 2041 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 V1-13_IGLJ5*01 837 1442 2042 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 V1-13_IGLJ5*01 837 1442 2043 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 V1-13_IGLJ5*01 837 1442 2044 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 V1-13_IGLJ5*01 837 1442 2045 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 V1-13_IGLJ5*01 837 1442 2046 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 V1-13_IGLJ5*01 837 1442 2047 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 V1-13_IGLJ5*01 837 1442 2048 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 V1-13_IGLJ5*01 837 1442 2049 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 V1-13_IGLJ5*01 837 1442 2050 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 V1-13_IGLJ5*01 837 1442 2051 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 V1-13_IGLJ5*01 837 1442 2052 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 V1-13_IGLJ5*01 837 1442 2053 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 V1-13_IGLJ5*01 837 1442 2054 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 V1-13_IGLJ5*01 837 1442 2055 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 V1-13_IGLJ5*01 837 1442 2056 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 V1-13_IGLJ5*01 837 1442 2057 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 V1-13_IGLJ5*01 837 1442 2058 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 V1-13_IGLJ5*01 837 1442 2059 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 V1-13_IGLJ5*01 837 1442 2060 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 V1-13_IGLJ5*01 837 1442 2061 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 V1-13_IGLJ5*01 837 1442 2062 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 V1-13_IGLJ5*01 837 1442 2063 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 V1-13_IGLJ5*01 837 1442 2064 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 V1-13_IGLJ5*01 837 1442 2065 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 V1-13_IGLJ5*01 837 1442 2066 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 V1-13_IGLJ5*01 837 1442 2067 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 V1-13_IGLJ5*01 837 1442 2068 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 V1-13_IGLJ5*01 837 1442 2069 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 V1-13_IGLJ5*01 837 1442 2070 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 V1-13_IGLJ5*01 837 1442 2071 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 V1-16_IGLJ6*01 838 1443 2072 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 V1-16_IGLJ6*01 838 1443 2073 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 V1-16_IGLJ6*01 838 1443 2074 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 V1-16_IGLJ6*01 838 1443 2075 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 V1-16_IGLJ6*01 838 1443 2076 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 V1-16_IGLJ6*01 838 1443 2077 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 V1-16_IGLJ6*01 838 1443 2078 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 V1-16_IGLJ6*01 838 1443 2079 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 V1-16_IGLJ6*01 838 1443 2080 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 V1-16_IGLJ6*01 838 1443 2081 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 V1-16_IGLJ6*01 838 1443 2082 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 V1-16_IGLJ6*01 838 1443 2083 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 V1-16_IGLJ6*01 838 1443 2084 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 V1-16_IGLJ6*01 838 1443 2085 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 V1-16_IGLJ6*01 838 1443 2086 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 V1-16_IGLJ6*01 838 1443 2087 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 V1-16_IGLJ6*01 838 1443 2088 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 V1-16_IGLJ6*01 838 1443 2089 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 V1-16_IGLJ6*01 838 1443 2090 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 V1-16_IGLJ6*01 838 1443 2091 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 V1-16_IGLJ6*01 838 1443 2092 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 V1-16_IGLJ6*01 838 1443 2093 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 V1-16_IGLJ6*01 838 1443 2094 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 V1-16_IGLJ6*01 838 1443 2095 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 V1-16_IGLJ6*01 838 1443 2096 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 V1-16_IGLJ6*01 838 1443 2097 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 V1-16_IGLJ6*01 838 1443 2098 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 V1-16_IGLJ6*01 838 1443 2099 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 V1-16_IGLJ6*01 838 1443 2100 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 V1-16_IGLJ6*01 838 1443 2101 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 V1-16_IGLJ6*01 838 1443 2102 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 V1-16_IGLJ6*01 838 1443 2103 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 V1-16_IGLJ6*01 838 1443 2104 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 V1-16_IGLJ6*01 838 1443 2105 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 V1-16_IGLJ6*01 838 1443 2106 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 V1-16_IGLJ6*01 838 1443 2107 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 V1-16_IGLJ6*01 838 1443 2108 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 V1-16_IGLJ6*01 838 1443 2109 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 V1-16_IGLJ6*01 838 1443 2110 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 V1-16_IGLJ6*01 838 1443 2111 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 V1-16_IGLJ6*01 838 1443 2112 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 V1-16_IGLJ6*01 838 1443 2113 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 V1-16_IGLJ6*01 838 1443 2114 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 V1-16_IGLJ6*01 838 1443 2115 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 V1-16_IGLJ6*01 838 1443 2116 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 V1-16_IGLJ6*01 838 1443 2117 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 V1-16_IGLJ6*01 838 1443 2118 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 V1-16_IGLJ6*01 838 1443 2119 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 V1-16_IGLJ6*01 838 1443 2120 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 V1-16_IGLJ6*01 838 1443 2121 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 V1-16_IGLJ6*01 838 1443 2122 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 V1-16_IGLJ6*01 838 1443 2123 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 V1-16_IGLJ6*01 838 1443 2124 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 V1-16_IGLJ6*01 838 1443 2125 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 V1-16_IGLJ6*01 838 1443 2126 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 V1-16_IGLJ6*01 838 1443 2127 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 V1-16_IGLJ6*01 838 1443 2128 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 V1-16_IGLJ6*01 838 1443 2129 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 V1-16_IGLJ6*01 838 1443 2130 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 V1-16_IGLJ6*01 838 1443 2131 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 V1-16_IGLJ6*01 838 1443 2132 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 V1-16_IGLJ6*01 838 1443 2133 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 V1-16_IGLJ6*01 838 1443 2134 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 V1-16_IGLJ6*01 838 1443 2135 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 V1-16_IGLJ6*01 838 1443 2136 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 V1-16_IGLJ6*01 838 1443 2137 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 V1-16_IGLJ6*01 838 1443 2138 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 V1-16_IGLJ6*01 838 1443 2139 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 V1-16_IGLJ6*01 838 1443 2140 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 V1-16_IGLJ6*01 838 1443 2141 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 V1-16_IGLJ6*01 838 1443 2142 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 V1-16_IGLJ6*01 838 1443 2143 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 V1-16_IGLJ6*01 838 1443 2144 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 V1-16_IGLJ6*01 838 1443 2145 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 V1-16_IGLJ6*01 838 1443 2146 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 V1-16_IGLJ6*01 838 1443 2147 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 V1-16_IGLJ6*01 838 1443 2148 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 V1-16_IGLJ6*01 838 1443 2149 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 V1-16_IGLJ6*01 838 1443 2150 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 V1-16_IGLJ6*01 838 1443 2151 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 V1-16_IGLJ6*01 838 1443 2152 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 V1-16_IGLJ6*01 838 1443 2153 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 V1-16_IGLJ6*01 838 1443 2154 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 V1-16_IGLJ6*01 838 1443 2155 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 V1-16_IGLJ6*01 838 1443 2156 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 V1-16_IGLJ6*01 838 1443 2157 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 V1-16_IGLJ6*01 838 1443 2158 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 V1-16_IGLJ6*01 838 1443 2159 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 V1-16_IGLJ6*01 838 1443 2160 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 V1-16_IGLJ6*01 838 1443 2161 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 V1-16_IGLJ6*01 838 1443 2162 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 V1-16_IGLJ6*01 838 1443 2163 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 V1-16_IGLJ6*01 838 1443 2164 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 V1-16_IGLJ6*01 838 1443 2165 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 V1-16_IGLJ6*01 838 1443 2166 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 V1-16_IGLJ6*01 838 1443 2167 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 V1-2_IGLJ7*01 840 1445 2168 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 V1-2_IGLJ7*01 840 1445 2169 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 V1-2_IGLJ7*01 840 1445 2170 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 V1-2_IGLJ7*01 840 1445 2171 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 V1-2_IGLJ7*01 840 1445 2172 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 V1-2_IGLJ7*01 840 1445 2173 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 V1-2_IGLJ7*01 840 1445 2174 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 V1-2_IGLJ7*01 840 1445 2175 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 V1-2_IGLJ7*01 840 1445 2176 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 V1-2_IGLJ7*01 840 1445 2177 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 V1-2_IGLJ7*01 840 1445 2178 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 V1-2_IGLJ7*01 840 1445 2179 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 V1-2_IGLJ7*01 840 1445 2180 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 V1-2_IGLJ7*01 840 1445 2181 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 V1-2_IGLJ7*01 840 1445 2182 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 V1-2_IGLJ7*01 840 1445 2183 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 V1-2_IGLJ7*01 840 1445 2184 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 V1-2_IGLJ7*01 840 1445 2185 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 V1-2_IGLJ7*01 840 1445 2186 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 V1-2_IGLJ7*01 840 1445 2187 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 V1-2_IGLJ7*01 840 1445 2188 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 V1-2_IGLJ7*01 840 1445 2189 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 V1-2_IGLJ7*01 840 1445 2190 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 V1-2_IGLJ7*01 840 1445 2191 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 V1-2_IGLJ7*01 840 1445 2192 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 V1-2_IGLJ7*01 840 1445 2193 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 V1-2_IGLJ7*01 840 1445 2194 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 V1-2_IGLJ7*01 840 1445 2195 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 V1-2_IGLJ7*01 840 1445 2196 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 V1-2_IGLJ7*01 840 1445 2197 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 V1-2_IGLJ7*01 840 1445 2198 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 V1-2_IGLJ7*01 840 1445 2199 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 V1-2_IGLJ7*01 840 1445 2200 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 V1-2_IGLJ7*01 840 1445 2201 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 V1-2_IGLJ7*01 840 1445 2202 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 V1-2_IGLJ7*01 840 1445 2203 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 V1-2_IGLJ7*01 840 1445 2204 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 V1-2_IGLJ7*01 840 1445 2205 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 V1-2_IGLJ7*01 840 1445 2206 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 V1-2_IGLJ7*01 840 1445 2207 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 V1-2_IGLJ7*01 840 1445 2208 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 V1-2_IGLJ7*01 840 1445 2209 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 V1-2_IGLJ7*01 840 1445 2210 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 V1-2_IGLJ7*01 840 1445 2211 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 V1-2_IGLJ7*01 840 1445 2212 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 V1-2_IGLJ7*01 840 1445 2213 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 V1-2_IGLJ7*01 840 1445 2214 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 V1-2_IGLJ7*01 840 1445 2215 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 V1-2_IGLJ7*01 840 1445 2216 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 V1-2_IGLJ7*01 840 1445 2217 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 V1-2_IGLJ7*01 840 1445 2218 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 V1-2_IGLJ7*01 840 1445 2219 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 V1-2_IGLJ7*01 840 1445 2220 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 V1-2_IGLJ7*01 840 1445 2221 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 V1-2_IGLJ7*01 840 1445 2222 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 V1-2_IGLJ7*01 840 1445 2223 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 V1-2_IGLJ7*01 840 1445 2224 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 V1-2_IGLJ7*01 840 1445 2225 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 V1-2_IGLJ7*01 840 1445 2226 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 V1-2_IGLJ7*01 840 1445 2227 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 V1-2_IGLJ7*01 840 1445 2228 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 V1-2_IGLJ7*01 840 1445 2229 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 V1-2_IGLJ7*01 840 1445 2230 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 V1-2_IGLJ7*01 840 1445 2231 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 V1-2_IGLJ7*01 840 1445 2232 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 V1-2_IGLJ7*01 840 1445 2233 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 V1-2_IGLJ7*01 840 1445 2234 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 V1-2_IGLJ7*01 840 1445 2235 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 V1-2_IGLJ7*01 840 1445 2236 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 V1-2_IGLJ7*01 840 1445 2237 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 V1-2_IGLJ7*01 840 1445 2238 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 V1-2_IGLJ7*01 840 1445 2239 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 V1-2_IGLJ7*01 840 1445 2240 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 V1-2_IGLJ7*01 840 1445 2241 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 V1-2_IGLJ7*01 840 1445 2242 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 V1-2_IGLJ7*01 840 1445 2243 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 V1-2_IGLJ7*01 840 1445 2244 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 V1-2_IGLJ7*01 840 1445 2245 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 V1-2_IGLJ7*01 840 1445 2246 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 V1-2_IGLJ7*01 840 1445 2247 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 V1-2_IGLJ7*01 840 1445 2248 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 V1-2_IGLJ7*01 840 1445 2249 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 V1-2_IGLJ7*01 840 1445 2250 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 V1-2_IGLJ7*01 840 1445 2251 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 V1-2_IGLJ7*01 840 1445
2252 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 V1-2_IGLJ7*01 840 1445 2253 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 V1-2_IGLJ7*01 840 1445 2254 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 V1-2_IGLJ7*01 840 1445 2255 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 V1-2_IGLJ7*01 840 1445 2256 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 V1-2_IGLJ7*01 840 1445 2257 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 V1-2_IGLJ7*01 840 1445 2258 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 V1-2_IGLJ7*01 840 1445 2259 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 V1-2_IGLJ7*01 840 1445 2260 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 V1-2_IGLJ7*01 840 1445 2261 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 V1-2_IGLJ7*01 840 1445 2262 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 V1-2_IGLJ7*01 840 1445 2263 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 V1-20_IGLJ6*01 841 1446 2264 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 V1-20_IGLJ6*01 841 1446 2265 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 V1-20_IGLJ6*01 841 1446 2266 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 V1-20_IGLJ6*01 841 1446 2267 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 V1-20_IGLJ6*01 841 1446 2268 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 V1-20_IGLJ6*01 841 1446 2269 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 V1-20_IGLJ6*01 841 1446 2270 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 V1-20_IGLJ6*01 841 1446 2271 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 V1-20_IGLJ6*01 841 1446 2272 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 V1-20_IGLJ6*01 841 1446 2273 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 V1-20_IGLJ6*01 841 1446 2274 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 V1-20_IGLJ6*01 841 1446 2275 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 V1-20_IGLJ6*01 841 1446 2276 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 V1-20_IGLJ6*01 841 1446 2277 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 V1-20_IGLJ6*01 841 1446 2278 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 V1-20_IGLJ6*01 841 1446 2279 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 V1-20_IGLJ6*01 841 1446 2280 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 V1-20_IGLJ6*01 841 1446 2281 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 V1-20_IGLJ6*01 841 1446 2282 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 V1-20_IGLJ6*01 841 1446 2283 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 V1-20_IGLJ6*01 841 1446 2284 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 V1-20_IGLJ6*01 841 1446 2285 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 V1-20_IGLJ6*01 841 1446 2286 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 V1-20_IGLJ6*01 841 1446 2287 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 V1-20_IGLJ6*01 841 1446 2288 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 V1-20_IGLJ6*01 841 1446 2289 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 V1-20_IGLJ6*01 841 1446 2290 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 V1-20_IGLJ6*01 841 1446 2291 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 V1-20_IGLJ6*01 841 1446 2292 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 V1-20_IGLJ6*01 841 1446 2293 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 V1-20_IGLJ6*01 841 1446 2294 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 V1-20_IGLJ6*01 841 1446 2295 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 V1-20_IGLJ6*01 841 1446 2296 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 V1-20_IGLJ6*01 841 1446 2297 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 V1-20_IGLJ6*01 841 1446 2298 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 V1-20_IGLJ6*01 841 1446 2299 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 V1-20_IGLJ6*01 841 1446 2300 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 V1-20_IGLJ6*01 841 1446 2301 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 V1-20_IGLJ6*01 841 1446 2302 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 V1-20_IGLJ6*01 841 1446 2303 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 V1-20_IGLJ6*01 841 1446 2304 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 V1-20_IGLJ6*01 841 1446 2305 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 V1-20_IGLJ6*01 841 1446 2306 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 V1-20_IGLJ6*01 841 1446 2307 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 V1-20_IGLJ6*01 841 1446 2308 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 V1-20_IGLJ6*01 841 1446 2309 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 V1-20_IGLJ6*01 841 1446 2310 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 V1-20_IGLJ6*01 841 1446 2311 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 V1-20_IGLJ6*01 841 1446 2312 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 V1-20_IGLJ6*01 841 1446 2313 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 V1-20_IGLJ6*01 841 1446 2314 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 V1-20_IGLJ6*01 841 1446 2315 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 V1-20_IGLJ6*01 841 1446 2316 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 V1-20_IGLJ6*01 841 1446 2317 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 V1-20_IGLJ6*01 841 1446 2318 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 V1-20_IGLJ6*01 841 1446 2319 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 V1-20_IGLJ6*01 841 1446 2320 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 V1-20_IGLJ6*01 841 1446 2321 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 V1-20_IGLJ6*01 841 1446 2322 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 V1-20_IGLJ6*01 841 1446 2323 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 V1-20_IGLJ6*01 841 1446 2324 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 V1-20_IGLJ6*01 841 1446 2325 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 V1-20_IGLJ6*01 841 1446 2326 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 V1-20_IGLJ6*01 841 1446 2327 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 V1-20_IGLJ6*01 841 1446 2328 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 V1-20_IGLJ6*01 841 1446 2329 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 V1-20_IGLJ6*01 841 1446 2330 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 V1-20_IGLJ6*01 841 1446 2331 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 V1-20_IGLJ6*01 841 1446 2332 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 V1-20_IGLJ6*01 841 1446 2333 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 V1-20_IGLJ6*01 841 1446 2334 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 V1-20_IGLJ6*01 841 1446 2335 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 V1-20_IGLJ6*01 841 1446 2336 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 V1-20_IGLJ6*01 841 1446 2337 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 V1-20_IGLJ6*01 841 1446 2338 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 V1-20_IGLJ6*01 841 1446 2339 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 V1-20_IGLJ6*01 841 1446 2340 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 V1-20_IGLJ6*01 841 1446 2341 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 V1-20_IGLJ6*01 841 1446 2342 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 V1-20_IGLJ6*01 841 1446 2343 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 V1-20_IGLJ6*01 841 1446 2344 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 V1-20_IGLJ6*01 841 1446 2345 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 V1-20_IGLJ6*01 841 1446 2346 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 V1-20_IGLJ6*01 841 1446 2347 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 V1-20_IGLJ6*01 841 1446 2348 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 V1-20_IGLJ6*01 841 1446 2349 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 V1-20_IGLJ6*01 841 1446 2350 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 V1-20_IGLJ6*01 841 1446 2351 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 V1-20_IGLJ6*01 841 1446 2352 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 V1-20_IGLJ6*01 841 1446 2353 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 V1-20_IGLJ6*01 841 1446 2354 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 V1-20_IGLJ6*01 841 1446 2355 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 V1-20_IGLJ6*01 841 1446 2356 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 V1-20_IGLJ6*01 841 1446 2357 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 V1-20_IGLJ6*01 841 1446 2358 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 V1-20_IGLJ6*01 841 1446 2359 VH3-23_IGHD6-6*01 > 2_IGHJ1*01 2118 V1-3_IGLJ1*01 842 1447 2360 VH3-23_IGHD6-13*01 > 1_IGHJ1*01 2119 V1-3_IGLJ1*01 842 1447 2361 VH3-23_IGHD6-13*01 > 2_IGHJ1*01 2120 V1-3_IGLJ1*01 842 1447 2362 VH3-23_IGHD6-19*01 > 1_IGHJ1*01 2121 V1-3_IGLJ1*01 842 1447 2363 VH3-23_IGHD6-19*01 > 2_IGHJ1*01 2122 V1-3_IGLJ1*01 842 1447 2364 VH3-23_IGHD6-25*01 > 1_IGHJ1*01 2123 V1-3_IGLJ1*01 842 1447 2365 VH3-23_IGHD6-25*01 > 2_IGHJ1*01 2124 V1-3_IGLJ1*01 842 1447 2366 VH3-23_IGHD7-27*01 > 1_IGHJ1*01 2125 V1-3_IGLJ1*01 842 1447 2367 VH3-23_IGHD7-27*01 > 3_IGHJ1*01 2126 V1-3_IGLJ1*01 842 1447 2368 VH3-23_IGHD6-13*01 > 1'_IGHJ1*01 2175 V1-3_IGLJ1*01 842 1447 2369 VH3-23_IGHD6-13*01 > 2'_IGHJ1*01 2176 V1-3_IGLJ1*01 842 1447 2370 VH3-23_IGHD6-13*01 > 2_IGHJ1*01_B 2177 V1-3_IGLJ1*01 842 1447 2371 VH3-23_IGHD6-19*01 > 1'_IGHJ1*01 2178 V1-3_IGLJ1*01 842 1447 2372 VH3-23_IGHD6-19*01 > 2'_IGHJ1*01 2179 V1-3_IGLJ1*01 842 1447 2373 VH3-23_IGHD6-19*01 > 2_IGHJ1*01_B 2180 V1-3_IGLJ1*01 842 1447 2374 VH3-23_IGHD6-25*01 > 1'_IGHJ1*01 2181 V1-3_IGLJ1*01 842 1447 2375 VH3-23_IGHD6-25*01 > 3'_IGHJ1*01 2182 V1-3_IGLJ1*01 842 1447 2376 VH3-23_IGHD7-27*01 > 1'_IGHJ1*01_B 2183 V1-3_IGLJ1*01 842 1447 2377 VH3-23_IGHD7-27*01 > 2'_IGHJ1*01 2184 V1-3_IGLJ1*01 842 1447 2378 VH3-23_IGHD6-6*01 > 2_IGHJ2*01 2233 V1-3_IGLJ1*01 842 1447 2379 VH3-23_IGHD6-13*01 > 1_IGHJ2*01 2234 V1-3_IGLJ1*01 842 1447 2380 VH3-23_IGHD6-13*01 > 2_IGHJ2*01 2235 V1-3_IGLJ1*01 842 1447 2381 VH3-23_IGHD6-19*01 > 1_IGHJ2*01 2236 V1-3_IGLJ1*01 842 1447 2382 VH3-23_IGHD6-19*01 > 2_IGHJ2*01 2237 V1-3_IGLJ1*01 842 1447 2383 VH3-23_IGHD6-25*01 > 1_IGHJ2*01 2238 V1-3_IGLJ1*01 842 1447 2384 VH3-23_IGHD6-25*01 > 2_IGHJ2*01 2239 V1-3_IGLJ1*01 842 1447 2385 VH3-23_IGHD7-27*01 > 1_IGHJ2*01 2240 V1-3_IGLJ1*01 842 1447 2386 VH3-23_IGHD7-27*01 > 3_IGHJ2*01 2241 V1-3_IGLJ1*01 842 1447 2387 VH3-23_IGHD6-13*01 > 1'_IGHJ2*01 2290 V1-3_IGLJ1*01 842 1447 2388 VH3-23_IGHD6-13*01 > 2'_IGHJ2*01 2291 V1-3_IGLJ1*01 842 1447 2389 VH3-23_IGHD6-13*01 > 2_IGHJ2*01_B 2292 V1-3_IGLJ1*01 842 1447 2390 VH3-23_IGHD6-19*01 > 1'_IGHJ2*01 2293 V1-3_IGLJ1*01 842 1447 2391 VH3-23_IGHD6-19*01 > 2'_IGHJ2*01 2294 V1-3_IGLJ1*01 842 1447 2392 VH3-23_IGHD6-19*01 > 2_IGHJ2*01_B 2295 V1-3_IGLJ1*01 842 1447 2393 VH3-23_IGHD6-25*01 > 1'_IGHJ2*01 2296 V1-3_IGLJ1*01 842 1447 2394 VH3-23_IGHD6-25*01 > 3'_IGHJ2*01 2297 V1-3_IGLJ1*01 842 1447 2395 VH3-23_IGHD7-27*01 > 1'_IGHJ2*01 2298 V1-3_IGLJ1*01 842 1447 2396 VH3-23_IGHD7-27*01 > 2'_IGHJ2*01 2299 V1-3_IGLJ1*01 842 1447 2397 VH3-23_IGHD6-6*01 > 2_IGHJ3*01 2348 V1-3_IGLJ1*01 842 1447 2398 VH3-23_IGHD6-13*01 > 1_IGHJ3*01 2349 V1-3_IGLJ1*01 842 1447 2399 VH3-23_IGHD6-13*01 > 2_IGHJ3*01 2350 V1-3_IGLJ1*01 842 1447 2400 VH3-23_IGHD6-19*01 > 1_IGHJ3*01 2351 V1-3_IGLJ1*01 842 1447 2401 VH3-23_IGHD6-19*01 > 2_IGHJ3*01 2352 V1-3_IGLJ1*01 842 1447 2402 VH3-23_IGHD6-25*01 > 1_IGHJ3*01 2353 V1-3_IGLJ1*01 842 1447 2403 VH3-23_IGHD6-25*01 > 2_IGHJ3*01 2354 V1-3_IGLJ1*01 842 1447 2404 VH3-23_IGHD7-27*01 > 1_IGHJ3*01 2355 V1-3_IGLJ1*01 842 1447 2405 VH3-23_IGHD7-27*01 > 3_IGHJ3*01 2356 V1-3_IGLJ1*01 842 1447 2406 VH3-23_IGHD6-13*01 > 1'_IGHJ3*01 2405 V1-3_IGLJ1*01 842 1447 2407 VH3-23_IGHD6-13*01 > 2'_IGHJ3*01 2406 V1-3_IGLJ1*01 842 1447 2408 VH3-23_IGHD6-13*01 > 1_IGHJ6*01 2407 V1-3_IGLJ1*01 842 1447 2409 VH3-23_IGHD6-19*01 > 1'_IGHJ3*01 2408 V1-3_IGLJ1*01 842 1447 2410 VH3-23_IGHD6-19*01 > 2'_IGHJ3*01 2409 V1-3_IGLJ1*01 842 1447 2411 VH3-23_IGHD6-19*01 > 3'_IGHJ3*01 2410 V1-3_IGLJ1*01 842 1447 2412 VH3-23_IGHD6-25*01 > 1'_IGHJ3*01 2411 V1-3_IGLJ1*01 842 1447 2413 VH3-23_IGHD6-25*01 > 3'_IGHJ3*01 2412 V1-3_IGLJ1*01 842 1447 2414 VH3-23_IGHD7-27*01 > 1'_IGHJ3*01 2413 V1-3_IGLJ1*01 842 1447 2415 VH3-23_IGHD7-27*01 > 2'_IGHJ3*01 2414 V1-3_IGLJ1*01 842 1447 2416 VH3-23_IGHD6-6*01 > 2_IGHJ4*01 2463 V1-3_IGLJ1*01 842 1447 2417 VH3-23_IGHD6-13*01 > 1_IGHJ4*01 2464 V1-3_IGLJ1*01 842 1447 2418 VH3-23_IGHD6-13*01 > 2_IGHJ4*01 2465 V1-3_IGLJ1*01 842 1447 2419 VH3-23_IGHD6-19*01 > 1_IGHJ4*01 2466 V1-3_IGLJ1*01 842 1447 2420 VH3-23_IGHD6-19*01 > 2_IGHJ4*01 2467 V1-3_IGLJ1*01 842 1447 2421 VH3-23_IGHD6-25*01 > 1_IGHJ4*01 2468 V1-3_IGLJ1*01 842 1447 2422 VH3-23_IGHD6-25*01 > 2_IGHJ4*01 2469 V1-3_IGLJ1*01 842 1447 2423 VH3-23_IGHD7-27*01 > 1_IGHJ4*01 2470 V1-3_IGLJ1*01 842 1447 2424 VH3-23_IGHD7-27*01 > 3_IGHJ4*01 2471 V1-3_IGLJ1*01 842 1447 2425 VH3-23_IGHD6-13*01 > 1'_IGHJ4*01 2520 V1-3_IGLJ1*01 842 1447 2426 VH3-23_IGHD6-13*01 > 2'_IGHJ4*01 2521 V1-3_IGLJ1*01 842 1447 2427 VH3-23_IGHD6-13*01 > 2_IGHJ4*01_B 2522 V1-3_IGLJ1*01 842 1447 2428 VH3-23_IGHD6-19*01 > 1'_IGHJ4*01 2523 V1-3_IGLJ1*01 842 1447 2429 VH3-23_IGHD6-19*01 > 2'_IGHJ4*01 2524 V1-3_IGLJ1*01 842 1447 2430 VH3-23_IGHD6-19*01 > 2_IGHJ4*01_B 2525 V1-3_IGLJ1*01 842 1447 2431 VH3-23_IGHD6-25*01 > 1'_IGHJ4*01 2526 V1-3_IGLJ1*01 842 1447 2432 VH3-23_IGHD6-25*01 > 3'_IGHJ4*01 2527 V1-3_IGLJ1*01 842 1447 2433 VH3-23_IGHD7-27*01 > 1'_IGHJ4*01 2528 V1-3_IGLJ1*01 842 1447 2434 VH3-23_IGHD7-27*01 > 2'_IGHJ4*01 2529 V1-3_IGLJ1*01 842 1447 2435 VH3-23_IGHD6-6*01 > 2_IGHJ5*01 2578 V1-3_IGLJ1*01 842 1447 2436 VH3-23_IGHD6-13*01 > 1_IGHJ5*01 2579 V1-3_IGLJ1*01 842 1447 2437 VH3-23_IGHD6-13*01 > 2_IGHJ5*01 2580 V1-3_IGLJ1*01 842 1447 2438 VH3-23_IGHD6-19*01 > 1_IGHJ5*01 2581 V1-3_IGLJ1*01 842 1447 2439 VH3-23_IGHD6-19*01 > 2_IGHJ5*01 2582 V1-3_IGLJ1*01 842 1447 2440 VH3-23_IGHD6-25*01 > 1_IGHJ5*01 2583 V1-3_IGLJ1*01 842 1447 2441 VH3-23_IGHD6-25*01 > 2_IGHJ5*01 2584 V1-3_IGLJ1*01 842 1447 2442 VH3-23_IGHD7-27*01 > 1_IGHJ5*01 2585 V1-3_IGLJ1*01 842 1447 2443 VH3-23_IGHD7-27*01 > 3_IGHJ5*01 2586 V1-3_IGLJ1*01 842 1447 2444 VH3-23_IGHD6-13*01 > 1'_IGHJ5*01 2635 V1-3_IGLJ1*01 842 1447 2445 VH3-23_IGHD6-13*01 > 2'_IGHJ5*01 2636 V1-3_IGLJ1*01 842 1447 2446 VH3-23_IGHD6-13*01 > 3'_IGHJ5*01 2637 V1-3_IGLJ1*01 842 1447 2447 VH3-23_IGHD6-19*01 > 1'_IGHJ5*01 2638 V1-3_IGLJ1*01 842 1447 2448 VH3-23_IGHD6-19*01 > 2'_IGHJ5*01 2639 V1-3_IGLJ1*01 842 1447 2449 VH3-23_IGHD6-19*01 > 2_IGHJ5*01_B 2640 V1-3_IGLJ1*01 842 1447 2450 VH3-23_IGHD6-25*01 > 1'_IGHJ5*01 2641 V1-3_IGLJ1*01 842 1447 2451 VH3-23_IGHD6-25*01 > 3'_IGHJ5*01 2642 V1-3_IGLJ1*01 842 1447 2452 VH3-23_IGHD7-27*01 > 1'_IGHJ5*01 2643 V1-3_IGLJ1*01 842 1447 2453 VH3-23_IGHD7-27*01 > 2'_IGHJ5*01 2644 V1-3_IGLJ1*01 842 1447 2454 VH3-23_IGHD6-6*01 > 2_IGHJ6*01 2693 V1-3_IGLJ1*01 842 1447 2455 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 V2-13_IGLJ2*01 849 1454 2456 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 V2-13_IGLJ2*01 849 1454 2457 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 V2-13_IGLJ2*01 849 1454 2458 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 V2-13_IGLJ2*01 849 1454 2459 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 V2-13_IGLJ2*01 849 1454 2460 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 V2-13_IGLJ2*01 849 1454 2461 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 V2-13_IGLJ2*01 849 1454 2462 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 V2-13_IGLJ2*01 849 1454 2463 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 V2-13_IGLJ2*01 849 1454 2464 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 V2-13_IGLJ2*01 849 1454 2465 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 V2-13_IGLJ2*01 849 1454 2466 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 V2-13_IGLJ2*01 849 1454 2467 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 V2-13_IGLJ2*01 849 1454 2468 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 V2-13_IGLJ2*01 849 1454 2469 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 V2-13_IGLJ2*01 849 1454 2470 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 V2-13_IGLJ2*01 849 1454 2471 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 V2-13_IGLJ2*01 849 1454 2472 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 V2-13_IGLJ2*01 849 1454 2473 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 V2-13_IGLJ2*01 849 1454 2474 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 V2-13_IGLJ2*01 849 1454 2475 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 V2-13_IGLJ2*01 849 1454 2476 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 V2-13_IGLJ2*01 849 1454 2477 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 V2-13_IGLJ2*01 849 1454 2478 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 V2-13_IGLJ2*01 849 1454 2479 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 V2-13_IGLJ2*01 849 1454 2480 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 V2-13_IGLJ2*01 849 1454 2481 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 V2-13_IGLJ2*01 849 1454 2482 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 V2-13_IGLJ2*01 849 1454 2483 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 V2-13_IGLJ2*01 849 1454 2484 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 V2-13_IGLJ2*01 849 1454 2485 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 V2-13_IGLJ2*01 849 1454 2486 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 V2-13_IGLJ2*01 849 1454 2487 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 V2-13_IGLJ2*01 849 1454 2488 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 V2-13_IGLJ2*01 849 1454 2489 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 V2-13_IGLJ2*01 849 1454 2490 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 V2-13_IGLJ2*01 849 1454 2491 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 V2-13_IGLJ2*01 849 1454 2492 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 V2-13_IGLJ2*01 849 1454 2493 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 V2-13_IGLJ2*01 849 1454 2494 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 V2-13_IGLJ2*01 849 1454 2495 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 V2-13_IGLJ2*01 849 1454 2496 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 V2-13_IGLJ2*01 849 1454 2497 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 V2-13_IGLJ2*01 849 1454 2498 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 V2-13_IGLJ2*01 849 1454 2499 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 V2-13_IGLJ2*01 849 1454 2500 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 V2-13_IGLJ2*01 849 1454 2501 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 V2-13_IGLJ2*01 849 1454 2502 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 V2-13_IGLJ2*01 849 1454
2503 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 V2-13_IGLJ2*01 849 1454 2504 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 V2-13_IGLJ2*01 849 1454 2505 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 V2-13_IGLJ2*01 849 1454 2506 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 V2-13_IGLJ2*01 849 1454 2507 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 V2-13_IGLJ2*01 849 1454 2508 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 V2-13_IGLJ2*01 849 1454 2509 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 V2-13_IGLJ2*01 849 1454 2510 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 V2-13_IGLJ2*01 849 1454 2511 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 V2-13_IGLJ2*01 849 1454 2512 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 V2-13_IGLJ2*01 849 1454 2513 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 V2-13_IGLJ2*01 849 1454 2514 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 V2-13_IGLJ2*01 849 1454 2515 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 V2-13_IGLJ2*01 849 1454 2516 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 V2-13_IGLJ2*01 849 1454 2517 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 V2-13_IGLJ2*01 849 1454 2518 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 V2-13_IGLJ2*01 849 1454 2519 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 V2-13_IGLJ2*01 849 1454 2520 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 V2-13_IGLJ2*01 849 1454 2521 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 V2-13_IGLJ2*01 849 1454 2522 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 V2-13_IGLJ2*01 849 1454 2523 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 V2-13_IGLJ2*01 849 1454 2524 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 V2-13_IGLJ2*01 849 1454 2525 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 V2-13_IGLJ2*01 849 1454 2526 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 V2-13_IGLJ2*01 849 1454 2527 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 V2-13_IGLJ2*01 849 1454 2528 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 V2-13_IGLJ2*01 849 1454 2529 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 V2-13_IGLJ2*01 849 1454 2530 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 V2-13_IGLJ2*01 849 1454 2531 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 V2-13_IGLJ2*01 849 1454 2532 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 V2-13_IGLJ2*01 849 1454 2533 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 V2-13_IGLJ2*01 849 1454 2534 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 V2-13_IGLJ2*01 849 1454 2535 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 V2-13_IGLJ2*01 849 1454 2536 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 V2-13_IGLJ2*01 849 1454 2537 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 V2-13_IGLJ2*01 849 1454 2538 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 V2-13_IGLJ2*01 849 1454 2539 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 V2-13_IGLJ2*01 849 1454 2540 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 V2-13_IGLJ2*01 849 1454 2541 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 V2-13_IGLJ2*01 849 1454 2542 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 V2-13_IGLJ2*01 849 1454 2543 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 V2-13_IGLJ2*01 849 1454 2544 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 V2-13_IGLJ2*01 849 1454 2545 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 V2-13_IGLJ2*01 849 1454 2546 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 V2-13_IGLJ2*01 849 1454 2547 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 V2-13_IGLJ2*01 849 1454 2548 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 V2-13_IGLJ2*01 849 1454 2549 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 V2-13_IGLJ2*01 849 1454 2550 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 V2-13_IGLJ2*01 849 1454 2551 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 V2-14_IGLJ4*01 850 1455 2552 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 V2-14_IGLJ4*01 850 1455 2553 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 V2-14_IGLJ4*01 850 1455 2554 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 V2-14_IGLJ4*01 850 1455 2555 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 V2-14_IGLJ4*01 850 1455 2556 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 V2-14_IGLJ4*01 850 1455 2557 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 V2-14_IGLJ4*01 850 1455 2558 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 V2-14_IGLJ4*01 850 1455 2559 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 V2-14_IGLJ4*01 850 1455 2560 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 V2-14_IGLJ4*01 850 1455 2561 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 V2-14_IGLJ4*01 850 1455 2562 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 V2-14_IGLJ4*01 850 1455 2563 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 V2-14_IGLJ4*01 850 1455 2564 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 V2-14_IGLJ4*01 850 1455 2565 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 V2-14_IGLJ4*01 850 1455 2566 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 V2-14_IGLJ4*01 850 1455 2567 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 V2-14_IGLJ4*01 850 1455 2568 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 V2-14_IGLJ4*01 850 1455 2569 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 V2-14_IGLJ4*01 850 1455 2570 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 V2-14_IGLJ4*01 850 1455 2571 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 V2-14_IGLJ4*01 850 1455 2572 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 V2-14_IGLJ4*01 850 1455 2573 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 V2-14_IGLJ4*01 850 1455 2574 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 V2-14_IGLJ4*01 850 1455 2575 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 V2-14_IGLJ4*01 850 1455 2576 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 V2-14_IGLJ4*01 850 1455 2577 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 V2-14_IGLJ4*01 850 1455 2578 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 V2-14_IGLJ4*01 850 1455 2579 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 V2-14_IGLJ4*01 850 1455 2580 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 V2-14_IGLJ4*01 850 1455 2581 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 V2-14_IGLJ4*01 850 1455 2582 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 V2-14_IGLJ4*01 850 1455 2583 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 V2-14_IGLJ4*01 850 1455 2584 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 V2-14_IGLJ4*01 850 1455 2585 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 V2-14_IGLJ4*01 850 1455 2586 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 V2-14_IGLJ4*01 850 1455 2587 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 V2-14_IGLJ4*01 850 1455 2588 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 V2-14_IGLJ4*01 850 1455 2589 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 V2-14_IGLJ4*01 850 1455 2590 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 V2-14_IGLJ4*01 850 1455 2591 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 V2-14_IGLJ4*01 850 1455 2592 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 V2-14_IGLJ4*01 850 1455 2593 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 V2-14_IGLJ4*01 850 1455 2594 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 V2-14_IGLJ4*01 850 1455 2595 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 V2-14_IGLJ4*01 850 1455 2596 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 V2-14_IGLJ4*01 850 1455 2597 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 V2-14_IGLJ4*01 850 1455 2598 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 V2-14_IGLJ4*01 850 1455 2599 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 V2-14_IGLJ4*01 850 1455 2600 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 V2-14_IGLJ4*01 850 1455 2601 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 V2-14_IGLJ4*01 850 1455 2602 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 V2-14_IGLJ4*01 850 1455 2603 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 V2-14_IGLJ4*01 850 1455 2604 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 V2-14_IGLJ4*01 850 1455 2605 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 V2-14_IGLJ4*01 850 1455 2606 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 V2-14_IGLJ4*01 850 1455 2607 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 V2-14_IGLJ4*01 850 1455 2608 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 V2-14_IGLJ4*01 850 1455 2609 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 V2-14_IGLJ4*01 850 1455 2610 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 V2-14_IGLJ4*01 850 1455 2611 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 V2-14_IGLJ4*01 850 1455 2612 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 V2-14_IGLJ4*01 850 1455 2613 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 V2-14_IGLJ4*01 850 1455 2614 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 V2-14_IGLJ4*01 850 1455 2615 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 V2-14_IGLJ4*01 850 1455 2616 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 V2-14_IGLJ4*01 850 1455 2617 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 V2-14_IGLJ4*01 850 1455 2618 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 V2-14_IGLJ4*01 850 1455 2619 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 V2-14_IGLJ4*01 850 1455 2620 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 V2-14_IGLJ4*01 850 1455 2621 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 V2-14_IGLJ4*01 850 1455 2622 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 V2-14_IGLJ4*01 850 1455 2623 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 V2-14_IGLJ4*01 850 1455 2624 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 V2-14_IGLJ4*01 850 1455 2625 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 V2-14_IGLJ4*01 850 1455 2626 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 V2-14_IGLJ4*01 850 1455 2627 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 V2-14_IGLJ4*01 850 1455 2628 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 V2-14_IGLJ4*01 850 1455 2629 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 V2-14_IGLJ4*01 850 1455 2630 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 V2-14_IGLJ4*01 850 1455 2631 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 V2-14_IGLJ4*01 850 1455 2632 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 V2-14_IGLJ4*01 850 1455 2633 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 V2-14_IGLJ4*01 850 1455 2634 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 V2-14_IGLJ4*01 850 1455 2635 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 V2-14_IGLJ4*01 850 1455 2636 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 V2-14_IGLJ4*01 850 1455 2637 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 V2-14_IGLJ4*01 850 1455 2638 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 V2-14_IGLJ4*01 850 1455 2639 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 V2-14_IGLJ4*01 850 1455 2640 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 V2-14_IGLJ4*01 850 1455 2641 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 V2-14_IGLJ4*01 850 1455 2642 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 V2-14_IGLJ4*01 850 1455 2643 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 V2-14_IGLJ4*01 850 1455 2644 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 V2-14_IGLJ4*01 850 1455 2645 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 V2-14_IGLJ4*01 850 1455 2646 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 V2-14_IGLJ4*01 850 1455 2647 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 V2-15_IGLJ7*01 850 1455 2648 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 V2-15_IGLJ7*01 851 1456 2649 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 V2-15_IGLJ7*01 851 1456 2650 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 V2-15_IGLJ7*01 851 1456 2651 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 V2-15_IGLJ7*01 851 1456 2652 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 V2-15_IGLJ7*01 851 1456 2653 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 V2-15_IGLJ7*01 851 1456 2654 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 V2-15_IGLJ7*01 851 1456 2655 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 V2-15_IGLJ7*01 851 1456 2656 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 V2-15_IGLJ7*01 851 1456 2657 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 V2-15_IGLJ7*01 851 1456 2658 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 V2-15_IGLJ7*01 851 1456 2659 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 V2-15_IGLJ7*01 851 1456 2660 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 V2-15_IGLJ7*01 851 1456 2661 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 V2-15_IGLJ7*01 851 1456 2662 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 V2-15_IGLJ7*01 851 1456 2663 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 V2-15_IGLJ7*01 851 1456 2664 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 V2-15_IGLJ7*01 851 1456 2665 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 V2-15_IGLJ7*01 851 1456 2666 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 V2-15_IGLJ7*01 851 1456 2667 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 V2-15_IGLJ7*01 851 1456 2668 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 V2-15_IGLJ7*01 851 1456 2669 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 V2-15_IGLJ7*01 851 1456 2670 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 V2-15_IGLJ7*01 851 1456 2671 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 V2-15_IGLJ7*01 851 1456 2672 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 V2-15_IGLJ7*01 851 1456 2673 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 V2-15_IGLJ7*01 851 1456 2674 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 V2-15_IGLJ7*01 851 1456 2675 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 V2-15_IGLJ7*01 851 1456 2676 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 V2-15_IGLJ7*01 851 1456 2677 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 V2-15_IGLJ7*01 851 1456 2678 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 V2-15_IGLJ7*01 851 1456 2679 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 V2-15_IGLJ7*01 851 1456 2680 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 V2-15_IGLJ7*01 851 1456 2681 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 V2-15_IGLJ7*01 851 1456 2682 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 V2-15_IGLJ7*01 851 1456 2683 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 V2-15_IGLJ7*01 851 1456 2684 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 V2-15_IGLJ7*01 851 1456 2685 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 V2-15_IGLJ7*01 851 1456 2686 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 V2-15_IGLJ7*01 851 1456 2687 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 V2-15_IGLJ7*01 851 1456 2688 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 V2-15_IGLJ7*01 851 1456 2689 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 V2-15_IGLJ7*01 851 1456 2690 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 V2-15_IGLJ7*01 851 1456 2691 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 V2-15_IGLJ7*01 851 1456 2692 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 V2-15_IGLJ7*01 851 1456 2693 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 V2-15_IGLJ7*01 851 1456 2694 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 V2-15_IGLJ7*01 851 1456 2695 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 V2-15_IGLJ7*01 851 1456 2696 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 V2-15_IGLJ7*01 851 1456 2697 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 V2-15_IGLJ7*01 851 1456 2698 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 V2-15_IGLJ7*01 851 1456 2699 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 V2-15_IGLJ7*01 851 1456 2700 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 V2-15_IGLJ7*01 851 1456 2701 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 V2-15_IGLJ7*01 851 1456 2702 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 V2-15_IGLJ7*01 851 1456 2703 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 V2-15_IGLJ7*01 851 1456 2704 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 V2-15_IGLJ7*01 851 1456 2705 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 V2-15_IGLJ7*01 851 1456 2706 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 V2-15_IGLJ7*01 851 1456 2707 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 V2-15_IGLJ7*01 851 1456 2708 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 V2-15_IGLJ7*01 851 1456 2709 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 V2-15_IGLJ7*01 851 1456 2710 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 V2-15_IGLJ7*01 851 1456 2711 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 V2-15_IGLJ7*01 851 1456 2712 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 V2-15_IGLJ7*01 851 1456 2713 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 V2-15_IGLJ7*01 851 1456 2714 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 V2-15_IGLJ7*01 851 1456 2715 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 V2-15_IGLJ7*01 851 1456 2716 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 V2-15_IGLJ7*01 851 1456 2717 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 V2-15_IGLJ7*01 851 1456 2718 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 V2-15_IGLJ7*01 851 1456 2719 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 V2-15_IGLJ7*01 851 1456 2720 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 V2-15_IGLJ7*01 851 1456 2721 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 V2-15_IGLJ7*01 851 1456 2722 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 V2-15_IGLJ7*01 851 1456 2723 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 V2-15_IGLJ7*01 851 1456 2724 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 V2-15_IGLJ7*01 851 1456 2725 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 V2-15_IGLJ7*01 851 1456 2726 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 V2-15_IGLJ7*01 851 1456 2727 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 V2-15_IGLJ7*01 851 1456 2728 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 V2-15_IGLJ7*01 851 1456 2729 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 V2-15_IGLJ7*01 851 1456 2730 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 V2-15_IGLJ7*01 851 1456 2731 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 V2-15_IGLJ7*01 851 1456 2732 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 V2-15_IGLJ7*01 851 1456 2733 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 V2-15_IGLJ7*01 851 1456 2734 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 V2-15_IGLJ7*01 851 1456 2735 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 V2-15_IGLJ7*01 851 1456 2736 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 V2-15_IGLJ7*01 851 1456 2737 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 V2-15_IGLJ7*01 851 1456 2738 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 V2-15_IGLJ7*01 851 1456 2739 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 V2-15_IGLJ7*01 851 1456 2740 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 V2-15_IGLJ7*01 851 1456 2741 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 V2-15_IGLJ7*01 851 1456 2742 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 V2-15_IGLJ7*01 851 1456 2743 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 V2-17_IGLJ2*01 852 1457 2744 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 V2-17_IGLJ2*01 852 1457 2745 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 V2-17_IGLJ2*01 852 1457 2746 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 V2-17_IGLJ2*01 852 1457 2747 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 V2-17_IGLJ2*01 852 1457 2748 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 V2-17_IGLJ2*01 852 1457 2749 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 V2-17_IGLJ2*01 852 1457 2750 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 V2-17_IGLJ2*01 852 1457 2751 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 V2-17_IGLJ2*01 852 1457 2752 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 V2-17_IGLJ2*01 852 1457 2753 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 V2-17_IGLJ2*01 852 1457
2754 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 V2-17_IGLJ2*01 852 1457 2755 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 V2-17_IGLJ2*01 852 1457 2756 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 V2-17_IGLJ2*01 852 1457 2757 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 V2-17_IGLJ2*01 852 1457 2758 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 V2-17_IGLJ2*01 852 1457 2759 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 V2-17_IGLJ2*01 852 1457 2760 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 V2-17_IGLJ2*01 852 1457 2761 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 V2-17_IGLJ2*01 852 1457 2762 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 V2-17_IGLJ2*01 852 1457 2763 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 V2-17_IGLJ2*01 852 1457 2764 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 V2-17_IGLJ2*01 852 1457 2765 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 V2-17_IGLJ2*01 852 1457 2766 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 V2-17_IGLJ2*01 852 1457 2767 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 V2-17_IGLJ2*01 852 1457 2768 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 V2-17_IGLJ2*01 852 1457 2769 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 V2-17_IGLJ2*01 852 1457 2770 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 V2-17_IGLJ2*01 852 1457 2771 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 V2-17_IGLJ2*01 852 1457 2772 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 V2-17_IGLJ2*01 852 1457 2773 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 V2-17_IGLJ2*01 852 1457 2774 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 V2-17_IGLJ2*01 852 1457 2775 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 V2-17_IGLJ2*01 852 1457 2776 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 V2-17_IGLJ2*01 852 1457 2777 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 V2-17_IGLJ2*01 852 1457 2778 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 V2-17_IGLJ2*01 852 1457 2779 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 V2-17_IGLJ2*01 852 1457 2780 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 V2-17_IGLJ2*01 852 1457 2781 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 V2-17_IGLJ2*01 852 1457 2782 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 V2-17_IGLJ2*01 852 1457 2783 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 V2-17_IGLJ2*01 852 1457 2784 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 V2-17_IGLJ2*01 852 1457 2785 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 V2-17_IGLJ2*01 852 1457 2786 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 V2-17_IGLJ2*01 852 1457 2787 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 V2-17_IGLJ2*01 852 1457 2788 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 V2-17_IGLJ2*01 852 1457 2789 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 V2-17_IGLJ2*01 852 1457 2790 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 V2-17_IGLJ2*01 852 1457 2791 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 V2-17_IGLJ2*01 852 1457 2792 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 V2-17_IGLJ2*01 852 1457 2793 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 V2-17_IGLJ2*01 852 1457 2794 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 V2-17_IGLJ2*01 852 1457 2795 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 V2-17_IGLJ2*01 852 1457 2796 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 V2-17_IGLJ2*01 852 1457 2797 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 V2-17_IGLJ2*01 852 1457 2798 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 V2-17_IGLJ2*01 852 1457 2799 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 V2-17_IGLJ2*01 852 1457 2800 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 V2-17_IGLJ2*01 852 1457 2801 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 V2-17_IGLJ2*01 852 1457 2802 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 V2-17_IGLJ2*01 852 1457 2803 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 V2-17_IGLJ2*01 852 1457 2804 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 V2-17_IGLJ2*01 852 1457 2805 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 V2-17_IGLJ2*01 852 1457 2806 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 V2-17_IGLJ2*01 852 1457 2807 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 V2-17_IGLJ2*01 852 1457 2808 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 V2-17_IGLJ2*01 852 1457 2809 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 V2-17_IGLJ2*01 852 1457 2810 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 V2-17_IGLJ2*01 852 1457 2811 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 V2-17_IGLJ2*01 852 1457 2812 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 V2-17_IGLJ2*01 852 1457 2813 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 V2-17_IGLJ2*01 852 1457 2814 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 V2-17_IGLJ2*01 852 1457 2815 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 V2-17_IGLJ2*01 852 1457 2816 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 V2-17_IGLJ2*01 852 1457 2817 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 V2-17_IGLJ2*01 852 1457 2818 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 V2-17_IGLJ2*01 852 1457 2819 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 V2-17_IGLJ2*01 852 1457 2820 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 V2-17_IGLJ2*01 852 1457 2821 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 V2-17_IGLJ2*01 852 1457 2822 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 V2-17_IGLJ2*01 852 1457 2823 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 V2-17_IGLJ2*01 852 1457 2824 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 V2-17_IGLJ2*01 852 1457 2825 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 V2-17_IGLJ2*01 852 1457 2826 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 V2-17_IGLJ2*01 852 1457 2827 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 V2-17_IGLJ2*01 852 1457 2828 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 V2-17_IGLJ2*01 852 1457 2829 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 V2-17_IGLJ2*01 852 1457 2830 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 V2-17_IGLJ2*01 852 1457 2831 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 V2-17_IGLJ2*01 852 1457 2832 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 V2-17_IGLJ2*01 852 1457 2833 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 V2-17_IGLJ2*01 852 1457 2834 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 V2-17_IGLJ2*01 852 1457 2835 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 V2-17_IGLJ2*01 852 1457 2836 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 V2-17_IGLJ2*01 852 1457 2837 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 V2-17_IGLJ2*01 852 1457 2838 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 V2-17_IGLJ2*01 852 1457 2839 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 V2-6_IGLJ4*01 854 1459 2840 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 V2-6_IGLJ4*01 854 1459 2841 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 V2-6_IGLJ4*01 854 1459 2842 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 V2-6_IGLJ4*01 854 1459 2843 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 V2-6_IGLJ4*01 854 1459 2844 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 V2-6_IGLJ4*01 854 1459 2845 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 V2-6_IGLJ4*01 854 1459 2846 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 V2-6_IGLJ4*01 854 1459 2847 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 V2-6_IGLJ4*01 854 1459 2848 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 V2-6_IGLJ4*01 854 1459 2849 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 V2-6_IGLJ4*01 854 1459 2850 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 V2-6_IGLJ4*01 854 1459 2851 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 V2-6_IGLJ4*01 854 1459 2852 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 V2-6_IGLJ4*01 854 1459 2853 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 V2-6_IGLJ4*01 854 1459 2854 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 V2-6_IGLJ4*01 854 1459 2855 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 V2-6_IGLJ4*01 854 1459 2856 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 V2-6_IGLJ4*01 854 1459 2857 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 V2-6_IGLJ4*01 854 1459 2858 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 V2-6_IGLJ4*01 854 1459 2859 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 V2-6_IGLJ4*01 854 1459 2860 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 V2-6_IGLJ4*01 854 1459 2861 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 V2-6_IGLJ4*01 854 1459 2862 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 V2-6_IGLJ4*01 854 1459 2863 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 V2-6_IGLJ4*01 854 1459 2864 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 V2-6_IGLJ4*01 854 1459 2865 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 V2-6_IGLJ4*01 854 1459 2866 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 V2-6_IGLJ4*01 854 1459 2867 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 V2-6_IGLJ4*01 854 1459 2868 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 V2-6_IGLJ4*01 854 1459 2869 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 V2-6_IGLJ4*01 854 1459 2870 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 V2-6_IGLJ4*01 854 1459 2871 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 V2-6_IGLJ4*01 854 1459 2872 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 V2-6_IGLJ4*01 854 1459 2873 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 V2-6_IGLJ4*01 854 1459 2874 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 V2-6_IGLJ4*01 854 1459 2875 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 V2-6_IGLJ4*01 854 1459 2876 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 V2-6_IGLJ4*01 854 1459 2877 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 V2-6_IGLJ4*01 854 1459 2878 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 V2-6_IGLJ4*01 854 1459 2879 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 V2-6_IGLJ4*01 854 1459 2880 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 V2-6_IGLJ4*01 854 1459 2881 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 V2-6_IGLJ4*01 854 1459 2882 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 V2-6_IGLJ4*01 854 1459 2883 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 V2-6_IGLJ4*01 854 1459 2884 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 V2-6_IGLJ4*01 854 1459 2885 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 V2-6_IGLJ4*01 854 1459 2886 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 V2-6_IGLJ4*01 854 1459 2887 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 V2-6_IGLJ4*01 854 1459 2888 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 V2-6_IGLJ4*01 854 1459 2889 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 V2-6_IGLJ4*01 854 1459 2890 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 V2-6_IGLJ4*01 854 1459 2891 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 V2-6_IGLJ4*01 854 1459 2892 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 V2-6_IGLJ4*01 854 1459 2893 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 V2-6_IGLJ4*01 854 1459 2894 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 V2-6_IGLJ4*01 854 1459 2895 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 V2-6_IGLJ4*01 854 1459 2896 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 V2-6_IGLJ4*01 854 1459 2897 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 V2-6_IGLJ4*01 854 1459 2898 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 V2-6_IGLJ4*01 854 1459 2899 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 V2-6_IGLJ4*01 854 1459 2900 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 V2-6_IGLJ4*01 854 1459 2901 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 V2-6_IGLJ4*01 854 1459 2902 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 V2-6_IGLJ4*01 854 1459 2903 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 V2-6_IGLJ4*01 854 1459 2904 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 V2-6_IGLJ4*01 854 1459 2905 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 V2-6_IGLJ4*01 854 1459 2906 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 V2-6_IGLJ4*01 854 1459 2907 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 V2-6_IGLJ4*01 854 1459 2908 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 V2-6_IGLJ4*01 854 1459 2909 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 V2-6_IGLJ4*01 854 1459 2910 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 V2-6_IGLJ4*01 854 1459 2911 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 V2-6_IGLJ4*01 854 1459 2912 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 V2-6_IGLJ4*01 854 1459 2913 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 V2-6_IGLJ4*01 854 1459 2914 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 V2-6_IGLJ4*01 854 1459 2915 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 V2-6_IGLJ4*01 854 1459 2916 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 V2-6_IGLJ4*01 854 1459 2917 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 V2-6_IGLJ4*01 854 1459 2918 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 V2-6_IGLJ4*01 854 1459 2919 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 V2-6_IGLJ4*01 854 1459 2920 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 V2-6_IGLJ4*01 854 1459 2921 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 V2-6_IGLJ4*01 854 1459 2922 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 V2-6_IGLJ4*01 854 1459 2923 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 V2-6_IGLJ4*01 854 1459 2924 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 V2-6_IGLJ4*01 854 1459 2925 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 V2-6_IGLJ4*01 854 1459 2926 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 V2-6_IGLJ4*01 854 1459 2927 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 V2-6_IGLJ4*01 854 1459 2928 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 V2-6_IGLJ4*01 854 1459 2929 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 V2-6_IGLJ4*01 854 1459 2930 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 V2-6_IGLJ4*01 854 1459 2931 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 V2-6_IGLJ4*01 854 1459 2932 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 V2-6_IGLJ4*01 854 1459 2933 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 V2-6_IGLJ4*01 854 1459 2934 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 V2-6_IGLJ4*01 854 1459 2935 VH3-23_IGHD1-1*01 > 1_IGHJ6*01 2645 V2-7_IGLJ2*01 855 1460 2936 VH3-23_IGHD1-1*01 > 2_IGHJ6*01 2646 V2-7_IGLJ2*01 855 1460 2937 VH3-23_IGHD1-1*01 > 3_IGHJ6*01 2647 V2-7_IGLJ2*01 855 1460 2938 VH3-23_IGHD1-7*01 > 1_IGHJ6*01 2648 V2-7_IGLJ2*01 855 1460 2939 VH3-23_IGHD1-7*01 > 3_IGHJ6*01 2649 V2-7_IGLJ2*01 855 1460 2940 VH3-23_IGHD1-14*01 > 1_IGHJ6*01 2650 V2-7_IGLJ2*01 855 1460 2941 VH3-23_IGHD1-14*01 > 3_IGHJ6*01 2651 V2-7_IGLJ2*01 855 1460 2942 VH3-23_IGHD1-20*01 > 1_IGHJ6*01 2652 V2-7_IGLJ2*01 855 1460 2943 VH3-23_IGHD1-20*01 > 3_IGHJ6*01 2653 V2-7_IGLJ2*01 855 1460 2944 VH3-23_IGHD1-26*01 > 1_IGHJ6*01 2654 V2-7_IGLJ2*01 855 1460 2945 VH3-23_IGHD1-26*01 > 3_IGHJ6*01 2655 V2-7_IGLJ2*01 855 1460 2946 VH3-23_IGHD2-2*01 > 2_IGHJ6*01 2656 V2-7_IGLJ2*01 855 1460 2947 VH3-23_IGHD2-2*01 > 3_IGHJ6*01 2657 V2-7_IGLJ2*01 855 1460 2948 VH3-23_IGHD2-8*01 > 2_IGHJ6*01 2658 V2-7_IGLJ2*01 855 1460 2949 VH3-23_IGHD2-8*01 > 3_IGHJ6*01 2659 V2-7_IGLJ2*01 855 1460 2950 VH3-23_IGHD2-15*01 > 2_IGHJ6*01 2660 V2-7_IGLJ2*01 855 1460 2951 VH3-23_IGHD2-15*01 > 3_IGHJ6*01 2661 V2-7_IGLJ2*01 855 1460 2952 VH3-23_IGHD2-21*01 > 2_IGHJ6*01 2662 V2-7_IGLJ2*01 855 1460 2953 VH3-23_IGHD2-21*01 > 3_IGHJ6*01 2663 V2-7_IGLJ2*01 855 1460 2954 VH3-23_IGHD3-3*01 > 1_IGHJ6*01 2664 V2-7_IGLJ2*01 855 1460 2955 VH3-23_IGHD3-3*01 > 2_IGHJ6*01 2665 V2-7_IGLJ2*01 855 1460 2956 VH3-23_IGHD3-3*01 > 3_IGHJ6*01 2666 V2-7_IGLJ2*01 855 1460 2957 VH3-23_IGHD3-9*01 > 2_IGHJ6*01 2667 V2-7_IGLJ2*01 855 1460 2958 VH3-23_IGHD3-10*01 > 2_IGHJ6*01 2668 V2-7_IGLJ2*01 855 1460 2959 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 2669 V2-7_IGLJ2*01 855 1460 2960 VH3-23_IGHD3-16*01 > 2_IGHJ6*01 2670 V2-7_IGLJ2*01 855 1460 2961 VH3-23_IGHD3-16*01 > 3_IGHJ6*01 2671 V2-7_IGLJ2*01 855 1460 2962 VH3-23_IGHD3-22*01 > 2_IGHJ6*01 2672 V2-7_IGLJ2*01 855 1460 2963 VH3-23_IGHD3-22*01 > 3_IGHJ6*01 2673 V2-7_IGLJ2*01 855 1460 2964 VH3-23_IGHD4-4*01 (1) > 2_IGHJ6*01 2674 V2-7_IGLJ2*01 855 1460 2965 VH3-23_IGHD4-4*01 (1) > 3_IGHJ6*01 2675 V2-7_IGLJ2*01 855 1460 2966 VH3-23_IGHD4-11*01 (1) > 2_IGHJ6*01 2676 V2-7_IGLJ2*01 855 1460 2967 VH3-23_IGHD4-11*01 (1) > 3_IGHJ6*01 2677 V2-7_IGLJ2*01 855 1460 2968 VH3-23_IGHD4-17*01 > 2_IGHJ6*01 2678 V2-7_IGLJ2*01 855 1460 2969 VH3-23_IGHD4-17*01 > 3_IGHJ6*01 2679 V2-7_IGLJ2*01 855 1460 2970 VH3-23_IGHD4-23*01 > 2_IGHJ6*01 2680 V2-7_IGLJ2*01 855 1460 2971 VH3-23_IGHD4-23*01 > 3_IGHJ6*01 2681 V2-7_IGLJ2*01 855 1460 2972 VH3-23_IGHD5-5*01 (2) > 1_IGHJ6*01 2682 V2-7_IGLJ2*01 855 1460 2973 VH3-23_IGHD5-5*01 (2) > 2_IGHJ6*01 2683 V2-7_IGLJ2*01 855 1460 2974 VH3-23_IGHD5-5*01 (2) > 3_IGHJ6*01 2684 V2-7_IGLJ2*01 855 1460 2975 VH3-23_IGHD5-12*01 > 1_IGHJ6*01 2685 V2-7_IGLJ2*01 855 1460 2976 VH3-23_IGHD5-12*01 > 3_IGHJ6*01 2686 V2-7_IGLJ2*01 855 1460 2977 VH3-23_IGHD5-18*01 (2) > 1_IGHJ6*01 2687 V2-7_IGLJ2*01 855 1460 2978 VH3-23_IGHD5-18*01 (2) > 2_IGHJ6*01 2688 V2-7_IGLJ2*01 855 1460 2979 VH3-23_IGHD5-18*01 (2) > 3_IGHJ6*01 2689 V2-7_IGLJ2*01 855 1460 2980 VH3-23_IGHD5-24*01 > 1_IGHJ6*01 2690 V2-7_IGLJ2*01 855 1460 2981 VH3-23_IGHD5-24*01 > 3_IGHJ6*01 2691 V2-7_IGLJ2*01 855 1460 2982 VH3-23_IGHD6-6*01 > 1_IGHJ6*01 2692 V2-7_IGLJ2*01 855 1460 2983 VH3-23_IGHD1-1*01 > 1'_IGHJ6*01 2702 V2-7_IGLJ2*01 855 1460 2984 VH3-23_IGHD1-1*01 > 2'_IGHJ6*01 2703 V2-7_IGLJ2*01 855 1460 2985 VH3-23_IGHD1-1*01 > 3'_IGHJ6*01 2704 V2-7_IGLJ2*01 855 1460 2986 VH3-23_IGHD1-7*01 > 1'_IGHJ6*01 2705 V2-7_IGLJ2*01 855 1460 2987 VH3-23_IGHD1-7*01 > 3'_IGHJ6*01 2706 V2-7_IGLJ2*01 855 1460 2988 VH3-23_IGHD1-14*01 > 1'_IGHJ6*01 2707 V2-7_IGLJ2*01 855 1460 2989 VH3-23_IGHD1-14*01 > 2'_IGHJ6*01 2708 V2-7_IGLJ2*01 855 1460 2990 VH3-23_IGHD1-14*01 > 3'_IGHJ6*01 2709 V2-7_IGLJ2*01 855 1460 2991 VH3-23_IGHD1-20*01 > 1'_IGHJ6*01 2710 V2-7_IGLJ2*01 855 1460 2992 VH3-23_IGHD1-20*01 > 2'_IGHJ6*01 2711 V2-7_IGLJ2*01 855 1460 2993 VH3-23_IGHD1-20*01 > 3'_IGHJ6*01 2712 V2-7_IGLJ2*01 855 1460 2994 VH3-23_IGHD1-26*01 > 1'_IGHJ6*01 2713 V2-7_IGLJ2*01 855 1460 2995 VH3-23_IGHD1-26*01 > 1_IGHJ6*01_B 2714 V2-7_IGLJ2*01 855 1460 2996 VH3-23_IGHD2-2*01 > 2_IGHJ6*01_B 2715 V2-7_IGLJ2*01 855 1460 2997 VH3-23_IGHD2-2*01 > 3'_IGHJ6*01 2716 V2-7_IGLJ2*01 855 1460 2998 VH3-23_IGHD2-8*01 > 1'_IGHJ6*01 2717 V2-7_IGLJ2*01 855 1460 2999 VH3-23_IGHD2-15*01 > 1'_IGHJ6*01 2718 V2-7_IGLJ2*01 855 1460 3000 VH3-23_IGHD2-15*01 > 3'_IGHJ6*01 2719 V2-7_IGLJ2*01 855 1460 3001 VH3-23_IGHD2-21*01 > 1'_IGHJ6*01 2720 V2-7_IGLJ2*01 855 1460 3002 VH3-23_IGHD2-21*01 > 3'_IGHJ6*01 2721 V2-7_IGLJ2*01 855 1460 3003 VH3-23_IGHD3-3*01 > 1'_IGHJ6*01 2722 V2-7_IGLJ2*01 855 1460 3004 VH3-23_IGHD3-3*01 > 3'_IGHJ6*01 2723 V2-7_IGLJ2*01 855 1460
3005 VH3-23_IGHD3-9*01 > 1'_IGHJ6*01 2724 V2-7_IGLJ2*01 855 1460 3006 VH3-23_IGHD3-9*01 > 3'_IGHJ6*01 2725 V2-7_IGLJ2*01 855 1460 3007 VH3-23_IGHD3-10*01 > 1'_IGHJ6*01 2726 V2-7_IGLJ2*01 855 1460 3008 VH3-23_IGHD3-10*01 > 3'_IGHJ6*01 2727 V2-7_IGLJ2*01 855 1460 3009 VH3-23_IGHD3-16*01 > 1'_IGHJ6*01 2728 V2-7_IGLJ2*01 855 1460 3010 VH3-23_IGHD3-16*01 > 3'_IGHJ6*01 2729 V2-7_IGLJ2*01 855 1460 3011 VH3-23_IGHD3-22*01 > 1'_IGHJ6*01 2730 V2-7_IGLJ2*01 855 1460 3012 VH3-23_IGHD4-4*01 (1) > 1'_IGHJ6*01 2731 V2-7_IGLJ2*01 855 1460 3013 VH3-23_IGHD4-4*01 (1) > 3'_IGHJ6*01 2732 V2-7_IGLJ2*01 855 1460 3014 VH3-23_IGHD4-11*01 (1) > 1'_IGHJ6*01 2733 V2-7_IGLJ2*01 855 1460 3015 VH3-23_IGHD4-11*01 (1) > 3'_IGHJ6*01 2734 V2-7_IGLJ2*01 855 1460 3016 VH3-23_IGHD4-17*01 > 1'_IGHJ6*01 2735 V2-7_IGLJ2*01 855 1460 3017 VH3-23_IGHD4-17*01 > 3'_IGHJ6*01 2736 V2-7_IGLJ2*01 855 1460 3018 VH3-23_IGHD4-23*01 > 1'_IGHJ6*01 2737 V2-7_IGLJ2*01 855 1460 3019 VH3-23_IGHD4-23*01 > 3'_IGHJ6*01 2738 V2-7_IGLJ2*01 855 1460 3020 VH3-23_IGHD5-5*01 (2) > 1'_IGHJ6*01 2739 V2-7_IGLJ2*01 855 1460 3021 VH3-23_IGHD5-5*01 (2) > 3'_IGHJ6*01 2740 V2-7_IGLJ2*01 855 1460 3022 VH3-23_IGHD5-12*01 > 1'_IGHJ6*01 2741 V2-7_IGLJ2*01 855 1460 3023 VH3-23_IGHD5-12*01 > 3'_IGHJ6*01 2742 V2-7_IGLJ2*01 855 1460 3024 VH3-23_IGHD5-18*01 (2) > 1'_IGHJ6*01 2743 V2-7_IGLJ2*01 855 1460 3025 VH3-23_IGHD5-18*01 (2) > 3'_IGHJ6*01 2744 V2-7_IGLJ2*01 855 1460 3026 VH3-23_IGHD5-24*01 > 1'_IGHJ6*01 2745 V2-7_IGLJ2*01 855 1460 3027 VH3-23_IGHD5-24*01 > 3'_IGHJ6*01 2746 V2-7_IGLJ2*01 855 1460 3028 VH3-23_IGHD6-6*01 > 1'_IGHJ6*01 2747 V2-7_IGLJ2*01 855 1460 3029 VH3-23_IGHD6-6*01 > 2'_IGHJ6*01 2748 V2-7_IGLJ2*01 855 1460 3030 VH3-23_IGHD6-6*01 > 3'_IGHJ6*01 2749 V2-7_IGLJ2*01 855 1460
[0543] 4. Antibody Libraries
[0544] Provided herein are antibody libraries containing antibodies or portions thereof minimally containing a VH chain and a VL chain or a portion thereof containing a sufficient antigen binding site. The VH chain or portion thereof of the antibody members in the libraries provided herein are encoded by any of the nucleic acid members of the library set forth in Section E.1 above. The VL chain or portion thereof of the antibody members in the libraries provided herein are encoded by any of the nucleic acid members set forth in Section E.2 above. Thus, each antibody member of the library is derived in full or in part from germline segment sequences and/or are derived from modified sequences thereof. In some examples, the libraries are provided as addressed libraries.
[0545] In addition, each antibody in the library is productive and functional by virtue of containing a sufficient antigen binding site. The antibodies in the library are different, and when provided in an addressed format, each locus of the library contains a different antibody from all other loci in the library. Thus, the libraries provided herein exhibit high antibody diversity. The antibody libraries provided herein contain as few as 10.sup.2 different members and typically contain about or 10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4, 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5 and more unique members, including about or 10.sup.6, 10.sup.7, 10.sup.8, 10.sup.9 and more unique members. The antibody libraries provided herein can be produced by the methods herein, whereby the natural recombination process and natural structural diversity of the antibody repertoire is mimicked.
[0546] Besides containing a VH chain and a VL chain, or a portion thereof containing a sufficient antigen-binding site, the resulting antibodies in the library provided herein can contain all or a portion of a constant region. For example, the antibodies can contain one or more of a CH1, CH2, CH3 or C.sub.L portion. Generally, the antibodies or portions thereof contain a CH1 portion. The resulting antibodies or portions thereof include, but are not limited to a full-length antibody, Fab, Fab', F(ab').sub.2, single-chain Fvs (scFv), Fv, dsFv, diabody, Fd and Fd' fragments. Exemplary addressed antibody libraries provided herein are Fab libraries.
[0547] The number of different members in the antibody libraries provided herein can be restricted such that each member of the library is selected based on sequence similarities or differences or based on shared characteristics (e.g., a V region family, CDR3 length or composition or other biochemical attribute). For example, members of the library can be selected so that individual encoding germline segments of the VL or VH chains of the resulting antibodies have shared characteristics (e.g. are of the same subgroup or gene family) or otherwise contain similar or different sequence identity. In another example, the members of the library can be selected based on the sequence diversity of the VH or VL chains of the resulting antibody members. The antibody members of the library can be selected to be diverse in the represented sequences or to be similar. Thus, in some instances, library members represent a group of highly diverse antibodies. In other instances, library members represent a group similar, non-diverse antibodies. For example, library members can have 40%, 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% sequence similarity of the VH or VL to all other members in the library. The choice of antibody library is a function of the application and is within the level of one of skill in the art.
[0548] In some examples, the addressed libraries provided herein are human naive libraries. That means that every member of the library is derived completely from human germline segment sequences. For example, the VH chain of each antibody member is encoded by a nucleic acid molecule made up of a combination of a V.sub.H, D.sub.H and J.sub.H germline segment, for example, any set forth in Table 3 above, or a subset thereof, such that the resulting nucleic acid molecule has a sequence where the V.sub.H segment is 5' to the D.sub.H segment which is 5' to the J.sub.H segment. The VL chain of each antibody member is encoded by a combination of a V.sub..kappa. and J.sub..kappa. germline segments and/or V.sub..lamda. and J.sub..lamda. germline segments, for example, any set forth in Tables 3-4 above, or a subset thereof, such that the resulting nucleic acid molecule has a sequence where the V.sub.L segment (V.sub..kappa. or V.sub..lamda.) is 5' to the J.sub.L segment (J.sub..kappa. or J.sub..lamda.). It is understood that the library is considered to be naive and derived from germline even though, in practicing the method herein, the joint regions of the segments are altered to render the resulting encoding nucleic acid molecules in frame. Such alterations, however, are minor and variously include insertion or deletion generally of only a single nucleotide of a germline segment. In addition, other modification made to the recombined nucleic acid sequence by virtue of practice of the method herein, such as removal of stop codons and restriction enzyme site sequences, also result in naive antibodies. Naive antibody libraries provided herein can include the entire repertoire of naive antibodies, or a subset thereof. For example, a naive library provided herein can include 10.sup.3, 10.sup.4, 2.times.10.sup.4, 3.times.10.sup.4 4.times.10.sup.4, 5.times.10.sup.4, 6.times.10.sup.4, 4.times.10.sup.4, 7.times.10.sup.4, 8.times.10.sup.4, 9.times.10.sup.4, 10.sup.5 and more unique members, including about or 10.sup.6, 10.sup.7 or more members.
[0549] In particular examples, the VH chain of members of the antibody library is encoded by a sequence of nucleotides made up entirely of a V.sub.H and a J.sub.H germline segment, and also any sequence of nucleotides between the V.sub.H and J.sub.H germline segment. The V.sub.H segment is 5' to the random sequence of nucleotides which is 5' to the J.sub.H segment in the encoding nucleic acid molecule. Thus, the resulting antibody members of the library contain a random sequence of amino acids in the region that includes the central portion of the CDRH3, which is largely responsible for the antigen specificity of the resulting antibody. The sequence of nucleotides can be any random sequence of nucleotides. In some instances, the sequence of nucleotides is a sequence that encodes a peptide mimetic against any desired target, for example, a cell surface receptor. Exemplary peptide mimetics are set forth in Table 16. In other examples, the random sequence of nucleotides is a D.sub.H germline segment, or modified form thereof. Generally, the sequence of nucleotides is or is about 5, 10, 15, 20, 25, 30, 40, 50, 60, 70, 80, 90 or more nucleotides in length.
[0550] Also provided herein are antibody libraries where the antibodies in the libraries are derived at least in part from modified germline segment sequences. In some examples, all encoding germline segments are modified, for example, by random mutation. In other examples, particular regions of germline segments are targeted for modification. For example, the modified germline segment sequences can include modifications, such as by amino acid mutation, of one or more CDR. In particular, resulting antibody members of the libraries provided herein can contain modification of CDR3, such as of CDRH3. Thus, the resulting members of the library can contain one or more amino acid replacement of a CDR as compared to a naive antibody.
[0551] In some examples, the members of the antibody libraries provided herein can be directed toward a desired target or function by incorporation of sequences into the variable regions of a naive antibody that correspond to a polynucleotide target of interest. Thus, except for the incorporated sequence, which is added by replacement or insertion into the antibody sequence, the remaining antibody represents germline segment sequences. These incorporated sequences include, but are not limited to, peptides as well as portions of monoclonal antibodies.
[0552] For example, the antibody can contain a directed peptide that acts as a mimetic against a particular target. Generally, such antibodies act as agonists toward the target, but in some instances, can be antagonists. The peptides can be included in any region of the antibody, but generally are included in the variable region, and more generally in one or more of the CDR regions. In particular, directed peptides are included in the CDR3 region of the antibody. Each member antibody member of the library can be modified with the same directed peptide and/or with different directed peptides.
[0553] In related examples, the antibody can contain a sequence portion from a known monoclonal antibodies. The portion can include a portion corresponding to one or more CDRs of a known monoclonal antibody. In other cases, antibodies in the addressed library can contain entire variable region (for example, VL or VH) of a monoclonal antibody. For example, antibodies in the library can contain a VH or VL of a known monoclonal antibody, which can be combined with a VL or VH derived from germline segments as described herein. Exemplary of such an antibody library is one where one or more of the antibody members contain a VH or VL chain from any known monoclonal antibody set forth in Table 9 above.
[0554] The antibody libraries of provided herein can include members that represent one, some or all of the above examples.
[0555] In the libraries provided herein, the individual VH and VL chains or portions thereof can be the same of different from the VH and VL chains of other antibodies in the library. It is the combination of the VH and VL chain that renders each antibody in the library different, for example, each addressed antibody at a locus. Thus, for example, libraries provided herein can all contain a common VH chain, but can each contain a different VL chain. Thus, the resulting antibody members of the library each contain a different combination of VH and VL chain.
[0556] An antibody library provided herein can include a VH encoded from any of nucleic acid sequences SEQ ID NOS: 454-805 (each encoding a VH chain), and a VL chain encoded by any of nucleic acid molecule set forth in any of SEQ ID NOS: 806-815, 817, 819-834, and 836-867 (each encoding a VL chain), or sequences thereof that do not contain heterologous sequence for restriction sites at the 3' and 5' ends, and subsets thereof. An antibody library provided herein also can include a VH encoded from any of nucleic acid sequences SEQ ID NOS: 2070-2759 (each encoding a VH chain), and a VL chain encoded by any of nucleic acid molecule set forth in any of SEQ ID NOS: 806-815, 817, 819-834, and 836-867 (each encoding a VL chain), or sequences thereof not do not contain heterologous sequence for restriction sites at the 3' and 5' ends. Any combination of VH and VL from any of the above recombined nucleic acids provided herein can be paired to generate an antibody library. The components of the library, the size of the library and the type of library (e.g. Fab, full-length antibody, etc) all can be varied
[0557] Exemplary of such an antibody library provided herein includes a VH and a VL encoded by a sequence of nucleic acids set forth in Table 17. An additional exemplary antibody library includes a subset of members set forth in Table 17, including a VH and a VL chain encoded from nucleic acid sequences set forth in Table 22 below. For example, in such an exemplary library any of the VH chain sequences set forth in any of SEQ ID NOS: 1475-1826 is paired with any of the VL light chain sequences set forth in SEQ ID NOS:1827-1838, 1840-1855, 1857-1888. The resulting library, when expressed with a CH as exemplified in Example 9, is a Fab library. By virtue of the pairing of VH and VL chain the resulting library can contain at or about 2.1.0.times.10.sup.5 or more members, or a subset thereof. Each member pair is encoded by germline segments and represents a naive antibody.
[0558] Exemplary of such an antibody library provided herein includes a VH and a VL encoded by a sequence of nucleic acids set forth in 17.A above.
[0559] 5. Addressable Formats
[0560] The libraries provided herein are provided as addressed libraries such that each loci is a distinct address containing a different member of the library compared to all other loci of the library, and each can be identified by virtue of its address. For example, members at a loci can be identified by prior recording of their distinct location or by an identifiable tag. Where the library is a nucleic acid library, each loci of the library contains a nucleic acid molecule that is different from all other nucleic acid molecules at other loci in the library. Where the library is an antibody library, each loci of the library contains an antibody or portion thereof that is different from all other antibodies or portion thereof at other loci in the library.
[0561] The libraries provided herein can be presented in any addressable format known to one of skill in the art. Addressing can be effected by position on a surface or other physical locus or can be otherwise identifiable and/or sortable by other identifiers, such as a tag encoded with a bar code or other symbology, a chemical tag, an electronic, such RF tag, a color-coded tag or other such identifier. One of skill in the art is familiar with various addressable format. Exemplary of such formats is as an array. The library members can be provided on or in the array by immobilization or attachment, or can be provided in solution.
[0562] Any nucleic acid molecule or antibody provided herein can be provided in overlapping areas or at random locations on an array, such as a multiwell plate. Alternatively the library members can be provided in spatial array, such that each nucleic acid molecule or antibody is provided in a distinct region of the array which does not overlap with the any other region containing any other antibody or nucleic acid molecules. Any addressable array technology known in the art can be employed with the antibodies.
[0563] a. Multiwell Plate
[0564] Nucleic acid molecules or antibodies can be spatially arrayed in multiwell plates, such that each individual locus of the plate corresponds to one individual antibody. Multiwell plates can include, but are not limited to, 12-well, 24-well, 48-well, 96-well plates, 384-well plates, and 1536-well plates. In this instance, the identity of each member in each well is known. The members can be attached to the support or provided in solution. For example, one advantage of this technique is that antibodies are presented in solution and are therefore fully folded and functional thereby eliminating the loss of activity observed when proteins are immobilized on filters, chip surfaces or slides. In addition, in the case of antibodies, the antibodies can be screened for any desired activity, including but not limited to binding, cytotoxicity, differentiation or proliferation of cells, and alteration of gene expression. Since the identity of each antibody is known, information on structure activity relationships (SAR) is immediately available. Finally, pharmacokinetics and/or dose response experiments can be performed during screening or immediately following identification of a "hit" or lead compound.
[0565] In another instance, nucleic acid molecule and antibody members can be spatially arrayed in multiwell plates, such that each individual locus of the plate corresponds to a group of members. In this instance, the identity of every member within the group in any particular well is known. Groups of members can contain 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100 or more antibodies or nucleic acid molecules. For example, antibodies or nucleic acid molecules can be grouped randomly or by any desired characteristic, such as belonging to the same V-region family, containing similar CDR3 region amino acid composition or length, or any other biochemical attributes. For the case of antibodies, the group of addressed antibodies can be screened for any desired activity, including but not limited to binding, cytotoxicity, differentiation or proliferation of cells, and alteration of gene expression. Screening groups of libraries allows for screening a greater number of library members at any given time thereby covering a larger part of the available antibody diversity. Furthermore, following the identification of a "hit" or lead group, since the identity of every antibody within a group is known and the antibodies are all readily available individually, individual antibodies can be immediately screened to identify "active" library members within the group.
[0566] ii. Solid Support
[0567] Nucleic acid and antibody arrays include those in which members are immobilized on a solid support, such as in a microarray. For example, solid supports for attachment or immobilization include, but are not limited to, blotting on filters, on chip surfaces, or on cellulose and by attachment to slides using affinity tags. In some examples, cells expressing variant polypeptides can be naturally adsorbed to a bead, such that a population of beads contains a single cell per bead (Freeman et al. Biotechnol. Bioeng. (2004) 86:196-200). Following immobilization to a glass support, microcolonies can be grown and screened with a chromogenic or fluorogenic substrate. See, e.g., U.S. Pat. Nos. 6,372,483, 6,352,842, 6,346,416 and 6,242,266.
[0568] For example, it is contemplated herein that antibody members of the libraries can be immobilized on a chip. Exemplary of chips include those used for BIAcore. Determining the ability of the protein to bind to a target molecule can be accomplished, e.g., using a technology such as real-time Biomolecular Interaction Analysis (BIA). Sjolander, S. and Urbaniczky, C. (1991) Anal. Chem. 63:2338-2345 and Szabo et al. (1995) Curr. Opin. Struct. Biol. 5:699-705. Biacore is a method for measuring protein-protein interaction and binding affinity. The technology is based on surface plasmon resonance (SPR), an optical phenomenon that enables detection of unlabeled interactants in real time (Welford K. 1991, Opt. Quant. Elect. 23:1; Morton and Myszka, 1998, Methods in Enzymology 295: 268). The SPR-based biosensors can be used to monitor biomolecular interactions in real time to determine active concentration, screening and characterization in terms of both affinity and kinetics. BIAcore analysis conveniently generates association rate constants, dissociation rate constants, equilibrium dissociation constants, and affinity constants.
[0569] The methodology relies on immobilization of ligands onto the surface of a sensor chip consisting of a glass substrate having a gold film covered by a monolayer of a long hydroxyalkyl thiol to which is covalently attached a thin layer of carboxymethylated dextran. The immobilization procedure is performed with the sensor chip in place in the instrument and is continuously monitored by the SPR detector. An unknown sample or ligate solution is introduced into the apparatus to contact the immobilized ligand. The interaction between ligand and ligate is observed directly by surface plasmon resonance techniques and the measurements recorded on a computer via a program such as Bialogue [Pharmacia].
[0570] 6. Other Display Methods
[0571] The libraries also can be provided in other non-addressable formats. Exemplary of such other non-addressable formats include by display, in particular, any display format that facilitates screening of the members of the libraries for an activity or activities. Generally, such formats are used for the antibody libraries, but if desired can also be provided for the nucleic acid or vector libraries. Typically libraries are screened using a display technique, such that there is a physical link between individual molecules of the library (phenotype) and the genetic information encoding them (genotype). These methods include, but are not limited to, cell display, phage display, mRNA display, ribosome display and DNA display.
[0572] a. Cell Display
[0573] Antibody libraries for screening can be expressed on the surfaces of cells, including bacteria E. coli, yeast S. cerevisiae, and mammalian cells, by fusing them with a protein that is expressed on the surface of the cell. Cell display is a technology used to screen antibody libraries wherein immobilization of the target antigen is unnecessary. Instead a technique, such as fluorescence-activated cell sorting (FACS), can be used to identify desired antibodies. FACS permits the separation of subpopulations of cells on the basis of their light scatter properties as they pass through a laser beam. See e.g. United States Published Patent Application Nos. US 2003/0100023 and US 2003/0036092. Single chain antibodies can be expressed on the external surface of E. coli by fusing them to a protein previously shown to direct heterologous proteins to the bacterial surface (Francisco et al, (1993) Proc. Natl. Acad. Sci., USA, 90:10444-10448). Single chain and Fab antibodies can be displayed on the surface of a yeast cell, and homologous recombination in yeast can be exploited to generate libraries of transformants (see e.g. Kieke et al., (1997) Prot. Eng., 10:1303-1310; Weaver-Feldhaus et al., (2004) FEBS Lett., 564:24-34; and Swers et al., (2004) Nucleic Acids Res., 32:e36). Mammalian cell display has been utilized to screen scFv libraries as well as IgGs (Ho et al., (2005) J. Biol. Chem., 280:07-617).
[0574] b. Phage Display
[0575] Phage display is a widely used method for screening antibody libraries for their ability to bind to a particular antigen. Phage display is a cell based method in which proteins or peptides are expressed individually on the surface of phage as fusions to a coat protein, while the same phage particle carries the DNA encoding the protein or peptide (Smith, G. P. (1985) Science 228:1315-1317). Selection of the phage is achieved through a specific binding reaction involving recognition of the protein or peptide, enabling the particular phage to be isolated and cloned and the DNA for the protein or peptide to be recovered and propagated or expressed. Use of phage display is rapid and facile due to its reliance upon E. coli for amplification and propagation. Typical use involves panning phage libraries against an immobilized antigen.
[0576] c. mRNA Display and Ribosome Display
[0577] The use of mRNA display and ribosome display allow for totally in vitro construction of antibody libraries. mRNA display is a method of displaying proteins or peptides in which the nascent protein is caused to bind covalently to its mRNA through a puromycin link (Roberts et al (1997) Proc. Natl. Acad. Sci, U.S.A. 64:12297-12302). Puromycin acts as a mimic of aminacyl tRNA, enters the ribosome A site, and the nascent protein is bound covalently to it by the peptidyl-transferase activity of the ribosome. Selection is carried out on these protein-mRNA fusions after dissociation of the ribosome. Alternatively, ribosome display is a method of displaying proteins or peptides in nascent form on the surface of ribosomes, such that a stable complex with the encoding mRNA is formed; the complexes are selected with a ligand for the protein or peptide and the genetic information is obtained by reverse transcription of the isolated mRNA (see e.g. U.S. Pat. Nos. 5,643,768 and 5,658,754). Selection techniques are similar to that of phage display wherein an ribosome display libraries are panned against an immobilized antigen.
[0578] d. DNA Display
[0579] In DNA display the DNA encoding the peptide is linked to the peptide. In non-covalent DNA display, the DNA-protein linkage is promoted by the recognition of the bacterial RepA protein as well as its own origin of replication sequence integrated into the template DNA (Odegrip et al. (2004) Proc. Natl. Acad. Sci, U.S.A. 101:2806-2810). Alternatively, a biotin-streptavidin interaction can be utilized. In covalent DNA display a bacteriophage P2 protein genetically fused to an antibody fragment binds to its own DNA sequence (Reiersen et al (2005) Nucl. Acids Res. 33:e10). Alternatively, the DNA and the peptide can be compartmentalized, such as in an oil-in-water emulsion. Selection techniques are similar to that of phage display wherein DNA display libraries are panned against an immobilized antigen. See e.g. International Patent Publication No. WO 98/037186.
F. METHODS OF PRODUCTION OF ANTIBODIES
[0580] Nucleic acid molecules and antibody members of the libraries provided herein can be made by any method known to one of skill in the art. Such procedures are routine and are well known to the skill artisan. They include routine molecular biology techniques including gene synthesis, PCR, ligation, cloning, transfection and purification techniques. A description of such procedures is provided below.
[0581] For example, nucleic acid sequences can be constructed using gene synthesis techniques as discussed herein above. Gene synthesis or routine molecular biology techniques also can be used to effect insertion, deletion, addition or replacement of nucleotides. For example, additional nucleotide sequences can be joined to a nucleic acid sequence. In one example linker sequences can be added, such as sequences containing restriction endonuclease sites for the purpose of cloning the synthetic gene into a vector, for example, a protein expression vector or a vector designed for the amplification of the antibody constant region coding DNA sequences. Furthermore, additional nucleotide sequences specifying functional DNA elements can be operatively linked to a recombined germline encoding nucleic acid molecule. Examples of such sequences include, but are not limited to, promoter sequences designed to facilitate intracellular protein expression, and leader peptide sequences designed to facilitate protein secretion. Additional nucleotide sequences such as sequences specifying protein binding regions also can be linked to nucleic acid sequences. Such regions include, but are not limited to, sequences to facilitate uptake of recombined antibodies or fragments thereof into specific target cells, or otherwise enhance the pharmacokinetics of the synthetic gene.
[0582] Nucleic acid sequences can be further engineered as described herein, such as by mutagenesis, to generate mutant antibodies. Mutagenesis can be effected entirely through gene synthesis. For example, nucleic acid molecules can be designed manually or in silico for synthesis to encode mutant antibodies. The benefit of using gene synthesis methods is that the mutations can be effected so that the resulting nucleic acid molecules are in-frame and are "productive" as discussed herein above. Other methods of synthesis exist where randomization can be achieved during the gene synthesis. For example, a protocol has been developed by which synthesis of an oligonucleotide is "doped" with non-native phosphoramidites, resulting in randomization of the gene section targeted for random mutagenesis (Wang and Hoover (1997) J. Bacteriol., 179:5812-9). This method allows control of position selection while retaining a random substitution rate. Alternatively, mutagenesis can be effected through other molecular biology techniques. Generally, site-directed mutagenesis strategies can be employed.
[0583] Other current methods can be used to create mutant antibody libraries from a template nucleic acid molecule or molecules, such as a germline recombined nucleic acid molecule encoding a naive antibody. Such methods include, but are not limited to, error-prone polymerase chain reaction (Caldwell and Joyce (1992); Gram et al. (1992) Proc. Natl. Acad. Sci., 89:3576-80); cassette mutagenesis in which the specific region to be optimized is replaced with a synthetically mutagenized oligonucleotide (Stemmer and Morris (1992) Biotechniques, 13:214-20); Arkin and Youvan (1992) Proc. Natl. Acad. Sci., 89:7811-7815; Oliphant et al. (1986) Gene, 44:177-83; Hermes et al. (1990) Proc. Natl. Acad. Sci, 87:696-700); the use of mutator strains of hosts cells to add mutational frequency (Greener et al. (1997) Mol. Biotechnol., 7:189-95); DNA shuffling (Crameri et al. (1998) Nature, 391:288-291; U.S. Pat. Nos. 6,177,263; 5,965,408; Ostermeier et al. (1999) Nat. Biotechnol., 17:1205-1209); and other random mutagenesis methods.
[0584] 1. Vectors
[0585] Provided herein are vectors that contain nucleic acid encoding the recombined antibodies or portions thereof. The nucleic acids encoding antibody polypeptides are typically cloned into a intermediate vector before transformation into prokaryotic or eukaryotic cells. Choice of vector can depend on the desired application. For example, after insertion of the nucleic acid, the vectors typically are used to transform host cells, for example, to amplify the recombined antibody genes for replication and/or expression thereof. In such examples, a vector suitable for high level expression is used. In other cases, a vector is chosen that is compatible with display of the expressed polypeptide on the surface of the cell.
[0586] Many expression vectors are available and known to those of skill in the art for the expression of recombined antibodies or portions thereof. The choice of an expression vector is influenced by the choice of host expression system. Such selection is well within the level of skill of the skilled artisan. In general, expression vectors can include transcriptional promoters and optionally enhancers, translational signals, and transcriptional and translational termination signals. Expression vectors that are used for stable transformation typically have a selectable marker which allows selection and maintenance of the transformed cells. In some cases, an origin of replication can be used to amplify the copy number of the vectors in the cells. Vectors also generally can contain additional nucleotide sequences operably linked to the ligated nucleic acid molecule (e.g. His tag, Flag tag). For purposes herein, vectors generally include sequences encoding the constant region. Thus, recombined antibodies or portions thereof also can be expressed as protein fusions. For example, a fusion can be generated to add additional functionality to a polypeptide. Examples of fusion proteins include, but are not limited to, fusions of a signal sequence, an epitope tag such as for localization, e.g. a hiss tag or a myc tag, or a tag for purification, for example, a GST fusion, and a sequence for directing protein secretion and/or membrane association.
[0587] For example, expression of the proteins can be controlled by any promoter/enhancer known in the art. Suitable bacterial promoters are well known in the art and described herein below. Other suitable promoters for mammalian cells, yeast cells and insect cells are well known in the art and some are exemplified below. Selection of the promoter used to direct expression of a heterologous nucleic acid depends on the particular application. Promoters which can be used include but are not limited to eukaryotic expression vectors containing the SV40 early promoter (Bernoist and Chambon, Nature 290:304-310 (1981)), the promoter contained in the 3' long terminal repeat of Rous sarcoma virus (Yamamoto et al. Cell 22:787-797 (1980)), the herpes thymidine kinase promoter (Wagner et al., Proc. Natl. Acad. Sci. USA 78:1441-1445 (1981)), the regulatory sequences of the metallothionein gene (Brinster et al., Nature 296:39-42 (1982)); prokaryotic expression vectors such as the -lactamase promoter (Jay et al., (1981) Proc. Natl. Acad. Sci. USA 78:5543) or the tac promoter (DeBoer et al., Proc. Natl. Acad. Sci. USA 8021-25 (1983)); see also "Useful Proteins from Recombinant Bacteria": in Scientific American 242:79-94 (1980)); plant expression vectors containing the nopaline synthetase promoter (Herrara-Estrella et al., Nature 303:209-213 (1984)) or the cauliflower mosaic virus 35S RNA promoter (Gardner et al., Nucleic Acids Res. 9:2871 (1981)), and the promoter of the photosynthetic enzyme ribulose bisphosphate carboxylase (Herrera-Estrella et al., Nature 310:115-120 (1984)); promoter elements from yeast and other fungi such as the Gal4 promoter, the alcohol dehydrogenase promoter, the phosphoglycerol kinase promoter, the alkaline phosphatase promoter, and the following animal transcriptional control regions that exhibit tissue specificity and have been used in transgenic animals: elastase I gene control region which is active in pancreatic acinar cells (Swift et al., Cell 38:639-646 (1984); Ornitz et al., Cold Spring Harbor Symp. Quant. Biol. 50:399-409 (1986); MacDonald, Hepatology 7:425-515 (1987)); insulin gene control region which is active in pancreatic beta cells (Hanahan et al., Nature 315:115-122 (1985)), immunoglobulin gene control region which is active in lymphoid cells (Grosschedl et al., Cell 38:647-658 (1984); Adams et al., Nature 318:533-538 (1985); Alexander et al., Mol. Cell Biol. 7:1436-1444 (1987)), mouse mammary tumor virus control region which is active in testicular, breast, lymphoid and mast cells (Leder et al., Cell 45:485-495 (1986)), albumin gene control region which is active in liver (Pinckert et al., Genes and Devel. 1:268-276 (1987)), alpha-fetoprotein gene control region which is active in liver (Krumlauf et al., Mol. Cell. Biol. 5:1639-1648 (1985); Hammer et al., Science 235:53-58 1987)), alpha-1 antitrypsin gene control region which is active in liver (Kelsey et al., Genes and Devel. 1:161-171 (1987)), beta globin gene control region which is active in myeloid cells (Magram et al., Nature 315:338-340 (1985); Kollias et al., Cell 46:89-94 (1986)), myelin basic protein gene control region which is active in oligodendrocyte cells of the brain (Readhead et al., Cell 48:703-712 (1987)), myosin light chain-2 gene control region which is active in skeletal muscle (Shani, Nature 314:283-286 (1985)), and gonadotrophic releasing hormone gene control region which is active in gonadotrophs of the hypothalamus (Mason et al, Science 234:1372-1378 (1986)).
[0588] In addition to the promoter, the expression vector typically contains a transcription unit or expression cassette that contains all the additional elements required for the expression of the antibody, or portion thereof, in host cells. A typical expression cassette contains a promoter operably linked to the nucleic acid sequence encoding the germline antibody chain and signals required for efficient polyadenylation of the transcript, ribosome binding sites and translation termination. Additional elements of the cassette can include enhancers. In addition, the cassette typically contains a transcription termination region downstream of the structural gene to provide for efficient termination. The termination region can be obtained from the same gene as the promoter sequence or can be obtained from different genes.
[0589] Some expression systems have markers that provide gene amplification such as thymidine kinase and dihydrofolate reductase. Alternatively, high yield expression systems not involving gene amplification are also suitable, such as using a baculovirus vector in insect cells, with a nucleic acid sequence encoding a germline antibody chain under the direction of the polyhedron promoter or other strong baculovirus promoter.
[0590] For purposes herein, vectors are provided that contain a sequence of nucleotides that encodes a constant region of an antibody operably linked to the nucleic acid sequence encoding the recombined variable region of the antibody. The vector can include the sequence for one or all of a CH1, CH2, CH3 or CH4 and/or CL. Generally, such as for expression of Fabs, the vector contains the sequence for a CH1 or CL. Exemplary of such vectors containing a heavy chain constant region gene (e.g. CH1) are plasmids A and D, described herein. Exemplary of such vectors containing a light chain constant region genes are plasmids C and E, described herein.
[0591] Exemplary expression vectors include any mammalian expression vector such as, for example, pCMV. For bacterial expression, such vectors include pBR322, pUC, pSKF, pET23D, and fusion vectors such as MBP, GST and LacZ. Exemplary of such a vector are bacterial expression vectors such as, for example, plasmid A, plasmid C, plasmid D and plasmid E, described herein. Other eukaryotic vectors, for example any containing regulatory elements from eukaryotic viruses can be used as eukaryotic expression vectors. These include, for example, SV40 vectors, papilloma virus vectors, and vectors derived from Epstein-Bar virus. Other exemplary eukaryotic vectors include pMSG, pAV009/A+, pMT010/A+, pMAMneo-5, baculovirus pDSCE, and any other vector allowing expression of proteins under the direction of the CMV promoter, SV40 early promoter, SV40 late promoter, metallothionein promoter, murine mammary tumor virus promoter, Rous sarcoma virus promoter, polyhedron promoter, or other promoters shown effective for expression in eukaryotes.
[0592] Exemplary plasmid vectors for transformation of E. coli cells, include, for example, the ColE1 replication vectors described herein. Several features common to all these vectors include (a) a pBAD inducible promoter; (b) an AraC gene, which controls the pBAD promoter; (c) a synthetic ribosomal binding site (RBS) for efficient translation; (d) a ColE1 origin of replication, allowing for high copy expression; (e) a STII leader sequence, allowing for expressed proteins to be translocated to the periplasm; (f) a f1 origin of replication; and (g) a gene for conferring antibiotic resistance. Such plasmids include plasmid A (FIG. 3), plasmid C (FIG. 5), plasmid D (FIG. 4) and plasmid E (FIG. 6). Plasmid A and Plasmid D are utilized for expression of heavy chain antibody genes in as they contain a gene for the heavy chain constant region (CH1) operably linked to the inserted gene for the heavy chain variable region. The vectors contain NheI and NcoI restriction sites to allow for cloning of the recombined antibody genes described herein. Both vectors contain a pUC origin of replication, a ColE1 type origin of replication, and an aminoglycoside phosphotransferase gene conferring kanamycin resistance. Plasmid A contains a (His).sub.6 Tag and a Flag Tag for protein purification. Plasmid D contains both a (His).sub.6 Tag and a Flag Tag, and an additional LPETG tag, which allows for covalent attachment of the resulting protein using a sortase. Plasmid C and Plasmid E are utilized for expression of light chain antibody genes in as they contain a gene for the light chain constant region (CL) operably linked to the inserted gene for the light chain variable region. Plasmid C is specific for kappa light chains and contains BseWI and NcoI restriction sites to allow for cloning of the recombined antibody genes described herein. Plasmid E is specific for lambda light chains and contains AcrII and NcoI restriction sites to allow for cloning of the recombined antibody genes described herein. Both vectors contain a 3.3 origin of replication, a ColE1 type origin of replication, and a gene conferring chloramphenicol resistance. The vectors described above are designed to be utilized in a dual vector system, in which a light chain vector and a heavy chain vector are co-transformed. Thus, they contain two different but compatible ColE1 origins of replication utilized, one for heavy chains and one light chain. This allows for efficient expression of both chains of the antibody when the vectors are co-transformed and expressed.
[0593] Any methods known to those of skill in the art for the insertion of DNA fragments into a vector can be used to construct expression vectors containing a nucleic acid encoding an antibody chain. These methods can include in vitro recombinant DNA and synthetic techniques and in vivo recombinants (genetic recombination). The insertion into a cloning vector can, for example, be accomplished by ligating the DNA fragment into a cloning vector which has complementary cohesive termini. If the complementary restriction sites used to fragment the DNA are not present in the cloning vector, the ends of the DNA molecules can be enzymatically modified. Alternatively, any site desired can be produced by ligating nucleotide sequences (linkers) onto the DNA termini; these ligated linkers can contain specific chemically synthesized nucleic acids encoding restriction endonuclease recognition sequences.
[0594] 2. Cells and Expression Systems
[0595] Cells containing the vectors also are provided. Generally, any cell type that can be engineered to express heterologous DNA and has a secretory pathway is suitable. Expression hosts include prokaryotic and eukaryotic organisms such as bacterial cells (e.g. E. cob), yeast cells, fungal cells, Archea, plant cells, insect cells and animal cells including human cells. Expression hosts can differ in their protein production levels as well as the types of post-translational modifications that are present on the expressed proteins. Further, the choice of expression host is often related to the choice of vector and transcription and translation elements used. For example, the choice of expression host is often, but not always, dependent on the choice of precursor sequence utilized. For example, many heterologous signal sequences can only be expressed in a host cell of the same species (i.e., an insect cell signal sequence is optimally expressed in an insect cell). In contrast, other signal sequences can be used in heterologous hosts such as, for example, the human serum albumin (hHSA) signal sequence which works well in yeast, insect, or mammalian host cells and the tissue plasminogen activator pre/pro sequence which has been demonstrated to be functional in insect and mammalian cells (Tan et al., (2002) Protein Eng. 15:337). The choice of expression host can be made based on these and other factors, such as regulatory and safety considerations, production costs and the need and methods for purification. Thus, the vector system must be compatible with the host cell used.
[0596] Expression in eukaryotic hosts can include expression in yeasts such as Saccharomyces cerevisiae and Pichia pastoris, insect cells such as Drosophila cells and lepidopteran cells, plants and plant cells such as tobacco, corn, rice, algae, and lemna. Eukaryotic cells for expression also include mammalian cells lines such as Chinese hamster ovary (CHO) cells or baby hamster kidney (BHK) cells. Eukaryotic expression hosts also include production in transgenic animals, for example, including production in serum, milk and eggs.
[0597] Recombinant molecules can be introduced into host cells via, for example, transformation, transfection, infection, electroporation and sonoporation, so that many copies of the gene sequence are generated. Generally, standard transfection methods are used to produce bacterial, mammalian, yeast, or insect cell lines that express large quantity of antibody chains, which is then purified using standard techniques (see e.g., Colley et al. (1989) J. Biol. Chem., 264:17619-17622; Guide to Protein Purification, in Methods in Enzymology, vol. 182 (Deutscher, ed.), 1990). Transformation of eukaryotic and prokaryotic cells are performed according to standard techniques (see, e.g., Morrison (1977) J. Bact. 132:349-351; Clark-Curtiss and Curtiss (1983) Methods in Enzymology, 101, 347-362). For example, any of the well-known procedures for introducing foreign nucleotide sequences into host cells can be used. These include the use of calcium phosphate transfection, polybrene, protoplast fusion, electroporation, biolistics, liposomes, microinjection, plasma vectors, viral vectors and any other the other well known methods for introducing cloned genomic DNA, cDNA, synthetic DNA or other foreign genetic material into a host cell. Generally, for purposes herein, host cells are transfected with a first vector encoding at least a VH chain and a second vector encoding at least a VL chain. Thus, it is only necessary that the particular genetic engineering procedure used be capable of successfully introducing at least both genes into the host cell capable of expressing germline, or modified form thereof, antibody polypeptide.
[0598] Transformation of host cells with recombinant DNA molecules that incorporate the isolated recombined variable region gene, cDNA, or synthesized DNA sequence enables generation of multiple copies of the gene. Thus, the gene can be obtained in large quantities by growing transformants, isolating the recombinant DNA molecules from the transformants and, when necessary, retrieving the inserted gene from the isolated recombinant DNA. Generally, After the expression vector is introduced into the cells, the transfected cells are cultured under conditions favoring expression of the germline chain, which is recovered from the culture using standard purification techniques identified below.
[0599] Antibodies and portions thereof can be produced using a high throughput approach by any methods known in the art for protein production including in vitro and in vivo methods such as, for example, the introduction of nucleic acid molecules encoding recombined antibodies or portions thereof into a host cell or host animal and expression from nucleic acid molecules encoding recombined antibodies in vitro. Prokaryotes, especially E. coli, provide a system for producing large amounts of recombined antibodies or portions thereof, and are particularly desired in applications of high-throughput expression and purification of proteins. Transformation of E. coli is a simple and rapid technique well known to those of skill in the art. E. coli host strains for high throughput expression include, but are not limited to, BL21 (EMD Biosciences) and LMG194 (ATCC). Exemplary of such an E. coli host strain is BL21. Vectors for high throughput expression include, but are not limited to, pBR322 and pUC vectors. Exemplary of such vectors are the vectors described herein, including plasmid A, plasmid C, plasmid D and plasmid E. Automation of expression and purification can facilitate high-throughput expression. For example, use of a Piccolo.TM. system, a fully automatic system that combines cell culture with automated harvesting, lysing and purification units, or other similar robotic system can be employed.
[0600] a. Prokaryotic Expression
[0601] Prokaryotes, especially E. coli, provide a system for producing large amounts of recombined antibodies or portions thereof. Transformation of E. coli is a simple and rapid technique well known to those of skill in the art. Expression vectors for E. coli can contain inducible promoters that are useful for inducing high levels of protein expression and for expressing proteins that exhibit some toxicity to the host cells. Examples of inducible promoters include the lac promoter, the trp promoter, the hybrid tac promoter, the T7 and SP6 RNA promoters and the temperature regulated .lamda.P.sub.L promoter.
[0602] Recombined antibodies or portions thereof can be expressed in the cytoplasmic environment of E. coli. The cytoplasm is a reducing environment and for some molecules, this can result in the formation of insoluble inclusion bodies. Reducing agents such as dithiothreitol and 3-mercaptoethanol and denaturants (e.g., such as guanidine-HCl and urea) can be used to resolubilize the proteins. An exemplary alternative approach is the expression of recombined antibodies or fragments thereof in the periplasmic space of bacteria which provides an oxidizing environment and chaperonin-like and disulfide isomerases leading to the production of soluble protein. Typically, a leader sequence is fused to the protein to be expressed which directs the protein to the periplasm. The leader is then removed by signal peptidases inside the periplasm. There are three major pathways to translocate expressed proteins into the periplasm, namely the Sec pathway, the SRP pathway and the TAT pathway. Examples of periplasmic-targeting leader sequences include the pelB leader from the pectate lyase gene, the StII leader sequence, and the DsbA leader sequence. An exemplary leader sequence is a DsbA leader sequence. In some cases, periplasmic expression allows leakage of the expressed protein into the culture medium. The secretion of proteins allows quick and simple purification from the culture supernatant. Proteins that are not secreted can be obtained from the periplasm by osmotic lysis. Similar to cytoplasmic expression, in some cases proteins can become insoluble and denaturants and reducing agents can be used to facilitate solubilization and refolding. Temperature of induction and growth also can influence expression levels and solubility. Typically, temperatures between 25.degree. C. and 37.degree. C. are used. Mutations also can be used to increase solubility of expressed proteins. Typically, bacteria produce aglycosylated proteins. Thus, if proteins require glycosylation for function, glycosylation can be added in vitro after purification from host cells.
[0603] b. Yeast
[0604] Yeasts such as Saccharomyces cerevisiae, Schizosaccharomyces pombe, Yarrowia lipolytica, Kluyveromyces lactis, and Pichia pastoris are useful expression hosts for recombined antibodies or portions thereof. Yeast can be transformed with episomal replicating vectors or by stable chromosomal integration by homologous recombination. Typically, inducible promoters are used to regulate gene expression. Examples of such promoters include AOX1, GAL1, GAL7, and GALS and metallothionein promoters such as CUP1. Expression vectors often include a selectable marker such as LEU2, TRP1, HIS3, and URA3 for selection and maintenance of the transformed DNA. Proteins expressed in yeast are often soluble. Co-expression with chaperonins such as Bip and protein disulfide isomerase can improve expression levels and solubility. Additionally, proteins expressed in yeast can be directed for secretion using secretion signal peptide fusions such as the yeast mating type alpha-factor secretion signal from Saccharomyces cerevisae and fusions with yeast cell surface proteins such as the Aga2p mating adhesion receptor or the Arxula adeninivorans glucoamylase. A protease cleavage site such as for the Kex-2 protease, can be engineered to remove the fused sequences from the expressed polypeptides as they exit the secretion pathway. Yeast also is capable of glycosylation at Asn-X-Ser/Thr motifs.
[0605] c. Insects
[0606] Insect cells, particularly using baculovirus expression, are useful for expressing antibodies or portions thereof. Insect cells express high levels of protein and are capable of most of the post-translational modifications used by higher eukaryotes. Baculovirus have a restrictive host range which improves the safety and reduces regulatory concerns of eukaryotic expression. Typical expression vectors use a promoter for high level expression such as the polyhedrin promoter and p10 promoter of baculovirus. Commonly used baculovirus systems include the baculoviruses such as Autographa californica nuclear polyhedrosis virus (AcNPV), and the Bombyx mori nuclear polyhedrosis virus (BmNPV) and an insect cell line such as Sf9 derived from Spodoptera frugiperda and TN derived from Trichoplusia ni. For high-level expression, the nucleotide sequence of the molecule to be expressed is fused immediately downstream of the polyhedrin initiation codon of the virus. To generate baculovirus recombinants capable of expressing human antibodies, a dual-expression transfer, such as pAcUW51 (PharMingen) is utilized. Mammalian secretion signals are accurately processed in insect cells and can be used to secrete the expressed protein into the culture medium
[0607] An alternative expression system in insect cells is the use of stably transformed cells. Cell lines such as Sf9 derived cells from Spodoptera frugiperda and TN derived cells from Trichoplusia ni can be used for expression. The baculovirus immediate early gene promoter IE1 can be used to induce consistent levels of expression. Typical expression vectors include the pIE1-3 and pI31-4 transfer vectors (Novagen). Expression vectors are typically maintained by the use of selectable markers such as neomycin and hygromycin.
[0608] d. Mammalian Cells
[0609] Mammalian expression systems can be used to express antibodies or portions thereof. Expression constructs can be transferred to mammalian cells by viral infection such as adenovirus or by direct DNA transfer such as liposomes, calcium phosphate, DEAE-dextran and by physical means such as electroporation and microinjection. Expression vectors for mammalian cells typically include an mRNA cap site, a TATA box, a translational initiation sequence (Kozak consensus sequence) and polyadenylation elements. Such vectors often include transcriptional promoter-enhancers for high-level expression, for example the SV40 promoter-enhancer, the human cytomegalovirus (CMV) promoter and the long terminal repeat of Rous sarcoma virus (RSV). These promoter-enhancers are active in many cell types. Tissue and cell-type promoters and enhancer regions also can be used for expression. Exemplary promoter/enhancer regions include, but are not limited to, those from genes such as elastase I, insulin, immunoglobulin, mouse mammary tumor virus, albumin, alpha fetoprotein, alpha 1 antitrypsin, beta globin, myelin basic protein, myosin light chain 2, and gonadotropic releasing hormone gene control. Selectable markers can be used to select for and maintain cells with the expression construct. Examples of selectable marker genes include, but are not limited to, hygromycin B phosphotransferase, adenosine deaminase, xanthine-guanine phosphoribosyl transferase, aminoglycoside phosphotransferase, dihydrofolate reductase and thymidine kinase. Antibodies are typically produced using a NEO.sup.R/G418 system, a dihydrofolate reductase (DHFR) system or a glutamine synthetase (GS) system. The GS system uses joint expression vectors, such as pEE12/pEE6, to express both heavy chain and light chain. Fusion with cell surface signaling molecules such as TCR-.zeta. and Fc.sub..epsilon.RI-.gamma. can direct expression of the proteins in an active state on the cell surface.
[0610] Many cell lines are available for mammalian expression including mouse, rat human, monkey, chicken and hamster cells. Exemplary cell lines include but are not limited to CHO, Balb/3T3, HeLa, MT2, mouse NS0 (nonsecreting) and other myeloma cell lines, hybridoma and heterohybridoma cell lines, lymphocytes, fibroblasts, Sp2/0, COS, NIH3T3, HEK293, 293S, 2B8, and HKB cells. Cell lines also are available adapted to serum-free media which facilitates purification of secreted proteins from the cell culture media. One such example is the serum free EBNA-1 cell line (Pham et al., (2003) Biotechnol. Bioeng. 84:332-42.)
[0611] e. Plants
[0612] Transgenic plant cells and plants can be used to express proteins such as any antibody or portion thereof described herein. Expression constructs are typically transferred to plants using direct DNA transfer such as microprojectile bombardment and PEG-mediated transfer into protoplasts, and with agrobacterium-mediated transformation. Expression vectors can include promoter and enhancer sequences, transcriptional termination elements and translational control elements. Expression vectors and transformation techniques are usually divided between dicot hosts, such as Arabidopsis and tobacco, and monocot hosts, such as corn and rice. Examples of plant promoters used for expression include the cauliflower mosaic virus CaMV 35S promoter, the nopaline synthase promoter, the ribose bisphosphate carboxylase promoter and the maize ubiquitin-1 (ubi-1) promoter promoters. Selectable markers such as hygromycin, phosphomannose isomerase and neomycin phosphotransferase are often used to facilitate selection and maintenance of transformed cells. Transformed plant cells can be maintained in culture as cells, aggregates (callus tissue) or regenerated into whole plants. Transgenic plant cells also can include algae engineered to produce proteases or modified proteases (see for example, Mayfield et al, (2003) PNAS 100:438-442). Because plants have different glycosylation patterns than mammalian cells, this can influence the choice of protein produced in these hosts.
[0613] 3. Purification
[0614] Antibodies and portions thereof are purified by any procedure known to one of skill in the art. The recombined germline antibodies can be purified to substantial purity using standard protein purification techniques known in the art including but not limited to, SDS-PAGE, size fraction and size exclusion chromatography, ammonium sulfate precipitation, chelate chromatography, ionic exchange chromatography or column chromatography. For example, antibodies can be purified by column chromatography. Exemplary of a method to purify antibodies is by using column chromatography, wherein a solid support column material is linked to Protein G, a cell surface-associated protein from Streptococcus, that binds immunoglobulins with high affinity. The antibodies can be purified to 60%, 70%, 80% purity and typically at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% purity. Purity can be assessed by standard methods such as by SDS-PAGE and coomassie staining.
[0615] Methods for purification of recombined antibodies or portions thereof from host cells depend on the chosen host cells and expression systems. For secreted molecules, proteins are generally purified from the culture media after removing the cells. For intracellular expression, cells can be lysed and the proteins purified from the extract. When transgenic organisms such as transgenic plants and animals are used for expression, tissues or organs can be used as starting material to make a lysed cell extract. Additionally, transgenic animal production can include the production of polypeptides in milk or eggs, which can be collected, and if necessary further the proteins can be extracted and further purified using standard methods in the art.
[0616] When antibodies are expressed by transformed bacteria in large amounts, typically after promoter induction, although expression can be constitutive, the polypeptides can form insoluble aggregates. There are several protocols that are suitable for purification of polypeptide inclusion bodies known to one of skill in the art. Numerous variations will be apparent to those of skill in the art.
[0617] For example, in one method, the cell suspension is generally centrifuged and the pellet containing the inclusion bodies resuspended in buffer which does not dissolve but washes the inclusion bodies, e.g., 20 mM Tris-HCL (pH 7.2), 1 mM EDTA, 150 mM NaCl and 2% Triton-X 100, a non-ionic detergent. It can be necessary to repeat the wash step to remove as much cellular debris as possible. The remaining pellet of inclusion bodies can be resuspended in an appropriate buffer (e.g., 20 mM sodium phosphate, pH 6.8, 150 mM NaCl). Other appropriate buffers are apparent to those of skill in the art.
[0618] Alternatively, antibodies can be purified from bacteria periplasm. Where the polypeptide is exported into the periplasm of the bacteria, the periplasmic fraction of the bacteria can be isolated by cold osmotic shock in addition to other methods known to those of skill in the art. For example, in one method, to isolate recombinant polypeptides from the periplasm, the bacterial cells are centrifuged to form a pellet. The pellet is resuspended in a buffer containing 20% sucrose. To lyse the cells, the bacteria are centrifuged and the pellet is resuspended in ice-cold 5 mM MgSO.sub.4 and kept in an ice bath for approximately 10 minutes. The cell suspension is centrifuged and the supernatant decanted and saved. The recombinant polypeptides present in the supernatant can be separated from the host proteins by standard separation techniques well known to those of skill in the art. These methods include, but are not limited to, the following steps: solubility fractionation, size differential filtration, and column chromatography.
G. APPLICATIONS AND USES OF THE LIBRARIES
[0619] Provided herein are methods of using the antibody libraries provided herein for screening to identify or select an antibody or antibodies that alter or modulate (increase or decrease) an activity of a target. As discussed above, the antibody libraries provided herein contain members 1) that are each different from all other members in the library; and 2) that are each productive and functional. By virtue of these characteristics the libraries are both diverse and robust in their screening capabilities. For example, using the libraries provided herein, it is possible to screen a small library (e.g. containing 1000 or fewer members) and identify antibodies having desired functions or activities. This is exemplified in Example 13 where a library of only 960 members was used in a screening assay for binding to various targets, resulting in the identification of several high affinity (nanomolar affinity) antibodies.
[0620] In the method of screening herein, any desired activity can be assayed for, including but not limited to binding, cytotoxicity, differentiation or proliferation of cells, cell migration, apoptosis, angiogenesis and alteration of gene expression. For example, the resulting libraries can be screened for the discovery of agonist or antagonist antibodies against therapeutic targets, such as, for example, targets involved in cell proliferation and differentiation, cell migration, apoptosis and angiogenesis. Such targets include, but not limited to, growth factors, cytokines, lymphocytic antigens, other cellular activators and receptors thereof.
[0621] In other examples, the libraries can be screened using "blind" functional screens to measure an observable biological processes of interest without requiring specific knowledge of the target in advance. Any Hit that is identified in such screening assays, for example by virtue of a strong functional activity, can be further analyzed to identify the specific target. The "blind" assay approach can be applied to any observable biological outcome. For example, apoptosis of B-cells can be readily assayed, and a screen for increased B-cell apoptosis can yield antibody Hits that promote B-cell killing. Such antibodies can bind known B-cell surface-proteins like CD-20, or bind a novel target. In another example, the increased (or decreased) secretion of specific cytokines, for example interferon alpha, from a population of cells such as bone marrow cells can be assayed by ELISA, allowing screening for antibodies which modify the secretion of these proteins. No a priori knowledge of the mechanism that achieves the screened outcome is necessary, and the specific antibody target can later be pursued if the effect is strong. The blind functional assays can also be merged with high-content screening approaches to find antibodies that induce morphological changes in a certain cell type or promote cell differentiation in a progenitor cell population.
[0622] The methods of screening provided herein involve contacting each member of an antibody library provided herein with a target protein, peptide or antigen or cell expressing a target protein, peptide or antigen or simply a cell population and identifying antibody members that modulate (increase or decrease) the activity of the target protein or cell. For example, the members of the library can be contacted to identify antibodies that bind to the target protein. In another example, antibodies can be screened for modulation of a functional activity of a target. Functional assays permit identification of agonist and antagonist antibodies. It is contemplated herein that this screening assay is particularly useful against membrane-bound proteins, such as receptor targets and other signaling molecules.
[0623] The antibody libraries provided herein can be presented in a format that permits screening, such as, for example, as addressable libraries, including spatial arrays; and cell display libraries, including yeast display and mammalian display, phage display, mRNA display, including ribosome display, and DNA display. For example, screening is performed on addressable antibody libraries, such as a spatial arrayed library, whereby each antibody member at a locus (e.g. well) is separately contacted and screened against a target protein or proteins from all other members in the library. Thus, upon identification of a "HIT" the identity of the antibody member is immediately known from its location in the array without any requirement for further enrichment, amplification or sequencing thereof. Library screening can be high-throughput by screening hundreds to thousands or more antibodies in the same screening assay.
[0624] Any technique known to one of skill in the art can be used for antibody screening herein to identify antibody members having a desired activity. For a typical high-throughput screen, sets of microtiter plates are generated, whereby each well in the microplate contains a different library member. For example, plates can be created where every well of a plate contains an antibody containing the same antibody heavy chain, but each well contains a different light chain such that the paired antibody (heavy and light chain) in each well is different from the antibody in all other wells of the plate. All wells are then loaded with the constant components necessary for performing the screening assay (for example, target protein or cell expressing a target protein, buffer, assay reagent), incubated for the appropriate time, and assayed for a readout of choice. This method can be used for screening for activities such as binding or other functional assays where reaction with a single purified target molecule or cell(s) is sufficient to give a readout.
[0625] For example, in such methods, a cell-based assay can be performed to effect activation of the target (e.g. in the presence of a ligand thereof) and activation (such as by assessing signaling, differentiation, proliferation, chemotaxis, apoptosis or other downstream process) is assessed in the presence of the antibody. Generally, the performance of such functional assays require the presence of soluble protein. Hence, it is an advantage of the antibody libraries provided herein that they permit screening in a soluble format. For example, the antibodies can be addressed, for example, in wells of a multiwall plate. Each antibody at each locus is different. Thus, upon assessing activity the identity of a "Hit" can be immediately determined.
[0626] Following an initial round of screening, identified "HITS" can be further optimized by iterative screening and/or by directed evolution methods. For example, following identification of a "Hit" `focused` libraries can be created using information gained from the primary screening libraries. Focused libraries represent substructures of the original primary library because they are created from germline segment components that are related to those of the "Hit." They can allow one to refine the properties of a selected "Hit", such as by increasing its affinity of binding or associated functional activity.
[0627] The identity of the antibody is known by virtue of its address. Once identified, the antibody Hits can be generated as a full-length IgG or as Fab, Fab', Fab'-SH and F(ab').sub.2 fragments. The antibody or fragments thereof can be created by traditional means, such as using recombinant techniques and enzymatic digestion. Antibodies or fragments thereof can be chimeric or humanized. The antibodies or fragments thereof can be used for diagnostic and therapeutic purposes as set forth below.
[0628] 1. Binding Assays
[0629] The antibody libraries provided herein can be screened for their ability to bind a selected target by any method known to one of skill in the art. Exemplary target antigens are described below. Binding assays can be performed in solution, suspension or on a solid support. For example, target antigens can be immobilized to a solid support (e.g. a carbon or plastic surface or chip) and contacted with antibody. Unbound antibody or target protein can be washed away and bound complexes can then be detected. Binding assays can be performed under conditions to reduce nonspecific binding, such as by using a high ionic strength buffer (e.g. 0.3-0.4 M NaCl) with nonionic detergent (e.g. 0.1% Triton X-100 or Tween 20) and/or blocking proteins (e.g. bovine serum albumin or gelatin). Negative controls also can be including in such assays as a measure of background binding. Binding affinities can be determined using Scatchard analysis (Munson et al, Anal. Biochem., 107:220 (1980)), BIACore or other methods known to one of skill in the art.
[0630] Exemplary binding assays include, but are not limited to immunoassays such as competitive and non-competitive assay systems using techniques such as western blots, radioimmunoassays, ELISA (enzyme linked immunosorbent assay), "sandwich" immunoassays, Meso Scale Discovery (MSD, Gaithersburg, Md.), immunoprecipitation assays, ELISPOT, precipitin reactions, gel diffusion precipitin reactions, immunodiffusion assays, agglutination assays, complement-fixation assays, immunoradiometric assays, fluorescent immunoassays, and protein A immunoassays. Such assays are routine and well known in the art (see, e.g., Ausubel et al., eds, 1994, Current Protocols in Molecular Biology, Vol. 1, John Wiley & Sons, Inc., New York, which is incorporated by reference herein in its entirety). Other assay formats include liposome immunoassays (LIA), which use liposomes designed to bind specific molecules (e.g., antibodies) and release encapsulated reagents or markers. The released chemicals are then detected according to standard techniques (see Monroe et al, (1986) Amer. Clin. Prod. Rev. 5:34-41).
[0631] Generally, binding is detected using a detectable moiety or label (e.g. an enzyme, a radionuclide, a fluorescent probe, electrochemiluminescent label, or a color dye) typically attached to the target or, if desired, directly to the antibody members in the library. Alternatively, binding can be detected by a further third reagent that itself is labeled or detectable. For example, detection of an antibody bound to a target protein can be achieved using a labeled capture molecule in a sandwich assay format. Other proteins capable of specifically binding immunoglobulin constant regions, such as protein A or protein G also can be used as the label agent. These proteins exhibit a strong non-immunogenic reactivity with immunoglobulin constant regions from a variety of species (see, e.g., Kronval et al, (1973) J. Immunol. 111:1401-1406; Akerstrom et al, (1985) J. Immunol. 135:2589-2542). The detection agent can be modified with a detectable moiety, such as biotin, to which another molecule can specifically bind, such as streptavidin. A variety of detectable moieties are well known to those skilled in the art.
[0632] The choice of label or detectable group used in the assay is not critical, as long as it does not significantly interfere with the specific binding of the antibody used in the assay. Generally, the choice depends on sensitivity required, ease of conjugation with the compound, stability requirements, available instrumentation, and disposal provisions. One of skill in the art is familiar with labels and can identify a detectable label suitable for and compatible with the assay employed.
[0633] The detectable group can be any material having a detectable physical or chemical property. Such detectable labels have been well-developed in the field of immunoassays and, in general, most any label useful in such methods can be applied to the present invention. Thus, a label is any composition detectable by spectroscopic, photochemical, biochemical, immunochemical, electrical, optical or chemical means. Useful labels in the present invention include magnetic beads (e.g., DYNABEADS.TM.), fluorescent dyes (e.g., fluorescein isothiocyanate, Texas red, rhodamine, and the like), radiolabels (e.g., 3H, 125I, 35S, 14C, or 32P), enzymes (e.g., horse radish peroxidase, alkaline phosphatase and others commonly used in an ELISA), chemiluminescent labels (luciferin and 2,3-dihydrophtahlazinediones, e.g., luminol), and colorimetric labels such as colloidal gold or colored glass or plastic beads (e.g., polystyrene, polypropylene, latex, etc.). For a review of various labeling or signal producing systems that can be used, see e.g. U.S. Pat. No. 4,391,904.
[0634] Means of detecting labels are well known to those of skill in the art. Thus, for example, where the label is a radioactive label, means for detection include a scintillation counter or photographic film as in autoradiography. Where the label is a fluorescent label, it can be detected by exciting the fluorochrome with the appropriate wavelength of light and detecting the resulting fluorescence. The fluorescence can be detected visually, by the use of electronic detectors such as charge coupled devices (CCDs) or photomultipliers and the like. Similarly, enzymatic labels can be detected by providing the appropriate substrates for the enzyme and detecting the resulting reaction product. Finally simple colorimetric labels can be detected simply by observing the color associated with the label.
[0635] Some assay formats do not require the use of labeled components. For instance, agglutination assays can be used to detect the presence of the target antibodies. In this case, antigen-coated particles are agglutinated by samples containing the target antibodies. In this format, none of the components need be labeled and the presence of the target antibody is detected by simple visual inspection.
[0636] Alternatively, the antibody libraries provided herein can be screened for their ability to bind to cells, using whole cell panning, with or without subtractive panning. Screening can be done against live cells or against intact, mildly fixed target cells. Methods for whole cell panning have been described previously (see e.g. Siegel et al. (1997) J. Immunol. Methods 206:73-85 incorporated herein by reference). Other techniques for screening which can be applied include fluorescent activated cell sorting (FACs).
[0637] For high-throughput screening, assays can be multiplexed. Thus, the binding affinities of antibodies to a number of different target proteins can be determined at once. In one example, different target proteins can be separately labeled with different detectable moieties. For example, different antigens can be coupled to color-coded beads (Schwenk et al. (2007) Mol. Cell. Prot., 6:125-132). In another example, multi-spot plates can be used that permit assay multiplexing by absorption of up to 100 proteins in a locus of the plate (e.g. using Multi-Array or Multi-Spot plates from Meso Scale Discovery; MSD, Gaithersburg, Md.). In such an example, addressed antibody libraries provided herein can be screened by addition of a different antibody member to each well of a multi-spot plate. The assay readily permits the screening of thousands of antibodies at once against numerous target proteins. This is exemplified herein in Example 13.
[0638] In the methods of screening herein, the binding affinity of the antibodies is determined to identify or select antibodies that have high affinity for a target protein. Typically, antibodies are selected or identified that have a binding affinity that is or is about 10.sup.-6M, 10.sup.-7M, 10.sup.-8M, 10.sup.-9M, 10.sup.-10M, 10.sup.-11M, 10.sup.-12M or lower. Generally, antibodies are screened until antibodies are identified having nanomolar or sub-nanomolar binding affinity. A "Hit" identified in a first round of screening that does not have the desired binding affinity can be optimized by iterative screening and/or directed evolution methods and further screened for binding to a target antigen to identify an antibody that has a high binding affinity.
[0639] Any method known to one of skill in the art can be used to measure the binding affinity of an antibody. For example, the binding properties of an antibody can be assessed by performing a saturation binding assay, for example, a saturation ELISA, whereby binding to a target protein is assessed with increasing amounts of antibody. In such experiments, it is possible to assess whether the binding is dose-dependent and/or saturable. In addition, the binding affinity can be extrapolated from the 50% binding signal. Typically, apparent binding affinity is measured in terms of its association constant (Ka) or dissociation constant (Kd) and determined using Scatchard analysis (Munson et al, Anal. Biochem., 107:220 (1980). For example, binding affinity to a target protein can be assessed in a competition binding assay in where increasing concentrations of unlabeled protein is added, such as by radioimmunoassay (RIA) or ELISA. Binding affinity also can be analyzed using BIAcore kinetic analysis. This involves analyzing the binding and dissociation of an antibody member from chips containing immobilized target proteins on their surface. The Biacore evaluation software generates the values of Ka and Kd by fitting the data to interaction models. It is understood that the binding affinity of an antibody can vary depending on the assay and conditions employed, although all assays for binding affinity provide a rough approximation. By performing various assays under various conditions it is possible to estimate the binding affinity of an antibody.
[0640] In addition, binding affinities can differ depending on the structure of an antibody. For example, generally a bivalent antibody, for example a bivalent F(ab')2 fragment or full-length IgG, has a better binding affinity then a monovalent Fab antibody. Hence, it is understood that where a Fab has a specified binding affinity for a particular target, it is excepted that the binding affinity is even greater for a full-length IgG that is bivalent.
[0641] 2. Functional Activity
[0642] The antibody libraries provided herein can be screened for their ability to modulate the functional activity of a target by any method known to one of skill in the art. Assays can be designed to identify antibodies capable of binding and/or modulating cell surface receptors. Such antibodies can either be agonists, mimicking the normal effects of receptor binding, or antagonists, inhibiting the normal effects of receptor binding. Of particular interest is the identification of agents which bind to the receptors and modulate intracellular signalling.
[0643] In some example, such assays are cell-based assays. Generally, assays are performed using cell lines known to express the target of interest. Such cells are known to one of skill in the art. For example, one can consult the ATCC Catalog (atcc.org) to identify cell lines. Also, if a particular cell type is desired, the means for obtaining such cells, and/or their instantly available source is known to those in the art. An analysis of the scientific literature can readily reveal appropriate choice of cells expressing any desired target. Table 18 lists exemplary cells lines that express targets of interest that can be screened in functional activities herein against antibody libraries provided herein.
TABLE-US-00019 TABLE 18 Cell lines expressing targets Target Cell Lines References GP IIb/IIIa MEG-01 chronic myelogenous Ogura et al. Establishment of a novel human megakaryoblastic leukemia cell leukemia megakaryoblast cells line, MEG- 01, with positive Philadelphia chromosome. Blood 66: 1384-1392, (ATCC CRL-2021); 1985; Komatsu et al. Establishment and Characterization of a Human UT-7 human leukemia cell ine Leukemic Cell Line with Megakaryocytic Features: Dependency on Granulocyte-Macrophage Colony-stimulating Factor, Interleukin 3, or Erythropoietin for Growth and Survival. Cancer Research 51: 341-348 (1991) GM-CSF-R VA-ES-BJ epitheloid sarcoma Int J Oncol 1995; 7: 51-56; Ali Habib et al. A urokinase-activated recombinant cells (ATCC CRL-2138); diphtheria toxin targeting the granulocyte-macrophage colony-stimulating factor TF1-HaRas; receptor is selectively cytotoxic to human acute myeloid leukemia blasts. Blood TF1-vRaf; 104(7): 2143-2148 (2004); Kiser et al. Oncogene-dependent engraftment of TF1-vSrc; human myeloid leukemia cells in immunosuppressed mice. Leukemia HL-60 (ATCC CCL-240); 15(5): 814-818 (2001) U-937 (ATCC CRL-1593.2); ML-2 VEGFA Human A673 Gerber et al. Complete inhibition of rhabdomyosarcoma xenograft growth and rhabdomyosarcoma cells (ATCC neovascularization requires blockade of both tumor and host vascular CRL-1598); endothelial growth factor. Cancer Res. 60(22): 6253-8 (2000); Presta et al. Breast carcinoma MDA-MB-435 Humanization of an anti-vascular endothelial growth factor monoclonal cells (ATCC); Bovine adrenal antibody for the therapy of solid tumors and other disorders. Cancer Research, cortex-derived capillary 57(20): 4593-4599 (1997) endothelial cells CD3 Jurkat E6.1 Human leukemic T Buhler et al. A bispecific diabody directed against prostate-specific membrane cell lymphoblast (Sigma Aldrich antigen and CD3 induces T-cell mediated lysis of prostate cancer cells. Cancer 88042803) Immunol Immunother. 57(1): 43-52 (2008) EGFR DiFi human colorectal carcinoma Olive et al. Characterization of the DiFi rectal carcinoma cell line derived from a cells; familial adenomatous polyposis patient. In Vitro Cell Dev Biol. 29A(3 Pt 1): 239- A431 cells (ATCC CRL-1555); 248 (1993); Wu et al. Apoptosis induced by an anti-epidermal growth factor Caco-2 colorectal receptor monoclonal antibody in a human colorectal carcinoma cell line and its adenocarcinoma cells (ATCC delay by insulin. Clin. Invest. 95(4): 1897-1905 (1995) HTB-37); HRT-18 colorectal adenocarcinoma cells (ATCC CCL-244); HT-29 colorectal adenocarcinoma cells (ATCC HTB-38) EPO receptor A2780 ovarian cancer cells; Jeong et al. Characterization of erythropoietin receptor and erythropoietin UT-7 human leukemia cell ine expression and function in human ovarian cancer cells. Int J Cancer. 122(2): 274-280 (2008); Elliott et al. Activation of the Erythropoietin (EPO) Receptor by Bivalent Anti-EPO Receptor Antibodies. J Biol Chem. 271(40): 24691-24697 (1996) Her2/Neu BT-474 ductal carcinoma breast Le et al. Roles of human epidermal growth factor receptor 2, c-jun NH2- receptor cancer cell (ATCC HTB-20); terminal kinase, phosphoinositide 3-kinase, and p70 S6 kinase pathways in SK-BR-3 adenocarcinoma regulation of cyclin G2 expression in human breast cancer cells. Mol Cancer breast cancer cell (ATCC Ther. 6(11): 2843-2857 (2007) HTB-30); MDA-MB-453 metastatic carcinoma cell line (ATCC HTB-131) cMet H1993 lung adenocarcinoma Ma et al. Functional expression and mutations of c-Met and its therapeutic cells (ATCC CRL-5909); inhibition with SU11274 and small interfering RNA in non-small cell lung H1838 lung adenocarcinoma cancer. Cancer Res. 65(4): 1479-1488 (2005); cells (ATCC CRL-5899); Ma et al. A selective small molecule c-MET Inhibitor, PHA665752, cooperates SW 900 lung squamous cell with rapamycin. Clin Cancer Res 11(6): 2312-2319 (2005) carcinoma cells (ATCC HTB-59); H358 lung bronchioalveolar carcinoma cells (ATCC CRL-5807); SK-Lu-1 lung adenocarcinoma cells (ATCC HTB-57); H441 Non-small cell lung cancer cells (ATCC HTB-174) CD20 Ramos Burkitt's lymphoma B Jazirehi et al. Rituximab (anti-CD20) selectively modifies Bcl-xL and apoptosis cells (ATCC CRL-1596); protease activating factor-1 (Apaf-1) expression and sensitizes human non- Raji Burkitt's lymphoma B Hodgkin's lymphoma B cell lines to paclitaxel-induced apoptosis. Mol Cancer cells (ATCC CCL-86): Ther. 2(11): 1183-1193 (2003) Daudi Burkitt's lymphoma B cells (ATCC CCL-213); 2F7 Burkitt's lymphoma B cells
[0644] In addition, cells lines expressing a target of interest can be generated by transient or stable transfection with an expression vector expressing a target of interest. Methods of transfection and expression are known to those of skill in the art (see e.g., Kaufman R. J. (1990) Methods in Enzymology 185:537-566; Kaufman et al. (1990) Methods in Enzymology 185:537-566). In addition, any primary cell or cell line can be assessed for expression of a particular target (e.g. cell surface marker). Cell surface markers can be assayed using fluorescently labeled antibodies and FACS. Suitable cell lines include A549 (lung), HeLa, Jurkat, BJAB, Colo205, H1299, MCF7, MDA-MB-231, PC3, HUMEC, HUVEC, and PrEC.
[0645] Any suitable functional effect can be measured, as described herein. For example, cellular morphology (e.g., cell volume, nuclear volume, cell perimeter, and nuclear perimeter), ligand binding, substrate binding, nuclease activity, apoptosis, chemotaxis or cell migrations, cell surface marker expression, cellular proliferation, GFP positivity and dye dilution assays (e.g., cell tracker assays with dyes that bind to cell membranes), DNA synthesis assays (e.g., 3H-thymidine and fluorescent DNA-binding dyes such as BrdU or Hoechst dye with FACS analysis) and nuclear foci assays, are all suitable assays to identify potential modulators using a cell based system. Other functional activities that can be measured include, but are not limited to, ligand binding, substrate binding, endonuclease and/or exonuclease activity, transcriptional changes to both known and uncharacterized genetic markers (e.g., northern blots), changes in cell metabolism, changes related to cellular proliferation, cell surface marker expression, DNA synthesis, marker and dye dilution assays (e.g., GFP and cell tracker assays), contact inhibition, tumor growth in nude mice, and others.
[0646] For example, antibodies or portions thereof in the libraries provided herein can be assessed for their modulation of one or more phenotypes of a cell known to express a target protein. Phenotypic assays, kits and reagents for their use are well known to those skilled in the art and are herein used to screen antibody libraries. Representative phenotypic assays, which can be purchased from any one of several commercial vendors, include those for determining cell viability, cytotoxicity, proliferation or cell survival (Molecular Probes, Eugene, Oreg.; PerkinElmer, Boston, Mass.), protein-based assays including enzymatic assays (Panvera, LLC, Madison, Wis.; BD Biosciences, Franklin Lakes, N.J.; Oncogene Research Products, San Diego, Calif.), cell regulation, signal transduction, inflammation, oxidative processes and apoptosis (Assay Designs Inc., Ann Arbor, Mich.), triglyceride accumulation (Sigma-Aldrich, St. Louis, Mo.), angiogenesis assays, tube formation assays, cytokine and hormone assays and metabolic assays (Chemicon International Inc., Temecula, Calif.; Amersham Biosciences, Piscataway, N.J.).
[0647] Cells determined to be appropriate for a particular phenotypic assay (i.e., A549, HeLa, Jurkat, BJAB, Colo205, H1299, MCF7, MDA-MB-231, PC3, HUMEC, HUVEC, and PrEC and any others known to express the target of interest) are treated with antibodies as well as control compounds. If necessary, a ligand for the receptor target is included so that activation of the receptor is effected. At the end of the treatment period, treated and untreated cells are analyzed by one or more methods specific for the assay to determine phenotypic outcomes and endpoints.
[0648] Phenotypic endpoints include changes in cell morphology over time or treatment dose as well as changes in levels of cellular components such as proteins, lipids, nucleic acids, hormones, saccharides or metals. Measurements of cellular status which include pH, stage of the cell cycle, intake or excretion of biological indicators by the cell, are also endpoints of interest.
[0649] The assays can be performed to assess the direct effects of an antibody on a target protein. For example, if the target protein is a cell surface receptor, an antibody can be added to assess whether the target protein directly modulates, such as by stimulation, the activity or function of the receptor. In such instances, the antibody is deemed an agonist antibody. In other examples, if the target protein is a cell surface receptor, the activity of the receptor can be stimulated in the presence of a ligand or other stimulating agent in the presence or absence of the antibody to determine if the antibody modulates (e.g. inhibits) the actions of the antibody. For example, the antibody can act by blocking the ability of the ligand to interact with the receptor and/or otherwise induce a negative stimulatory signal. In such instances, the antibody is deemed to be an antagonist of the receptor. Thus, the methods of screening herein by functional activity permits identification of agonist and antagonist antibodies.
[0650] a. Differentiation
[0651] Cellular differentiation can be analyzed using any assay that allows a detection of a physical, chemical or phenotypic change. Various assays are used to quantitatively determine cellular proliferation and activation in response to an external stimuli. Cell proliferation assays are used to quantitatively determine cellular proliferation by incorporating a reagent into the DNA of newly synthesized cells upon cell division. Such reagents include, but are not limited to .sup.3H-thymidine, 5-bromo-2'-deoxyuridine (BrdU) and fluorescent Hoechst dyes. Cell viability assays are used to determine the number of healthy cells in a sample by staining cells with a dye and measuring how many cells uptake the dye based on the fact that living cells will exclude the dye. Such dyes include but are not limited to 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), 2,3-Bis(2-methoxy-4-nitro-5-sulfophenyl)-2H-tetrazolium-5-carboxanilide inner salt (XTT), and 4-[3-(4-iodophenyl)-2-(4-nitrophenyl)-2H-5-tetrazolio]-1,3-benzene disulfonate (WST-1). Uptake of the reagent is measured either colorimetrically using a spectrophotometer or by measuring radiation with a scintillation counter. Details of these methods are well-known to one skilled in the art. For example, Example 12 exemplifies an MTT proliferation assay to assess cell proliferation.
[0652] Fluorescent dyes are commonly used for the detection of live cells and key functional activities in a variety of cell-based assays. There are several non-radioactive, fluorescence-based assays that are not dependent on cellular metabolism. The fluorescent dye binds nucleic acids and the fluorescence can then be measured quantitatively or qualitatively. Such dyes include, but are not limited to, propidium iodide and Hoechst 33342. The cell number can then be quantitated based on the fluorescence. DNA content can also be quantitated using the tools available in the imaging instruments. Details of these methods are well known to one skilled in the art.
[0653] The degree of invasiveness into Matrigel or some other extracellular matrix constituent can be used as an assay to identify antibodies that are capable of inhibiting abnormal cell proliferation and tumor growth. Tumor cells exhibit a good correlation between malignancy and invasiveness of cells into Matrigel or some other extracellular matrix constituent. In this assay, tumorigenic cells are typically used as host cells. Therefore, antibodies can be identified by measuring changes in the level of invasiveness between the host cells before and after the introduction of potential modulators.
[0654] Briefly, the level of invasion of host cells can be measured by using filters coated with Matrigel or some other extracellular matrix constituent. Penetration into the gel, or through to the distal side of the filter, is rated as invasiveness, and rated histologically by number of cells and distance moved, or by prelabeling the cells with 125I and counting the radioactivity on the distal side of the filter or bottom of the dish. (see, e.g., Freshney, Culture of Animal Cells a Manual of Basic Technique, 3rd ed., Wiley-Liss, New York (1994), herein incorporated by reference).
[0655] b. Alteration of Gene Expression
[0656] Detection of binding and/or modulation of a target by an antibody can be accomplished by detecting a biological response, such as, for example, measuring Cat.sup.2+ ion flux, cAMP, IP3, PIP3 or transcription of reporter genes. Analysis of the genotype of the cell (measurement of the expression of one or more of the genes of the cell using a reporter gene assay) after treatment is also used as an indicator of the efficacy or potency of the antibody. Hallmark genes, or those genes suspected to be associated with a signal transduction pathway are measured in both treated and untreated cells.
[0657] Assays can be performed that measure the activation of a reporter gene. Suitable reporter genes include endogenous genes as well as exogenous genes that are introduced into a cell by any of the standard methods familiar to the skilled artisan, such as transfection, electroporation, lipofection and viral infection. For example, cells expressing a recombinant receptor can be transfected with a reporter gene (e.g., chloramphenicol acetyltransferase, firefly luciferase, bacterial luciferase, 3-galactosidase and alkaline phosphatase) operably linked to a response element. The cells are then incubated with antibodies and the expression of the reporter gene is compared to expression in control cells that do not express the recombinant receptor but that are essentially identical in other respects. A statistically significant change in reporter gene expression in the receptor-expressing cells is indicative of a test compound that interacts with the receptor. Furthermore, the protein of interest can be used as an indirect reporter via attachment to a second reporter such as red or green fluorescent protein (see, e.g., Mistili & Spector, (1997) Nature Biotechnology 15:961-964).
[0658] The reporter construct is typically transfected into a cell. After treatment with a potential modulator, the amount of reporter gene transcription, translation, or activity is measured according to standard techniques known to those of skill in the art. The use of a reporter gene assay using luciferase to measure activiation of STAT5 directly or by induction of cyclin-D promoter is exemplified in Example 12.
[0659] c. Cytotoxicity Activity
[0660] Antibodies can be screened for their ability to directly induce apoptosis or programmed cell death or to indirectly induce apoptosis by blocking growth factor receptors, thereby effectively arresting proliferation. Antibodies also bind complement, leading to direct cell toxicity, known as complement dependent cytotoxicity (CDC). Thus, assays can be performed to assess complement-dependent cytotoxicity.
[0661] A variety of assays to assess apoptosis are known to one of skill in the art. For example, apoptosis assays include those that assay for the activation of a caspase, which are enzymes involved in apoptosis. Caspase assays are based on the measurement of zymogen processing to an active enzyme and proteolytic activity. A number of commercial kits and reagents are available to assess apoptosis based on caspase function including, but not limited to, PhiPhiLux (Oncolmmunin, Inc.), Caspase 3 activity assay (Roche Applied science), Homogenous Caspase assay (Roche Applied Science), Caspase-Glo Assays (Promega), Apo-ONE Homogeneous Caspase-3/7 Assay (Promega), CaspACE Assay System Colorimetric or Fluormetric (Promega), EnzChek Caspase-3 Assay Kit (Invitrogen), Imag-iT LIVE green Caspase-3 and 7 Detection Kit (Invitrogen), Active Caspase-3 Detection Kits (Stratagene), Caspase-mediated Apoptosis Products (BioVision) and CasPASE Apoptosis Assay Kit (Genotech). Example 11 exemplifies a assaying for apoptosis using a caspase assay.
[0662] Assays for apoptosis include TUNEL and DNA fragmentation assays that measure the activation of nucleases and subsequent cleavage of DNA into 180 to 200 base pair increments. Such assays and kits are commercially available and include, but are not limited to, Apoptotic DNA Ladder Kit (Roche Applied Science), Cellular DNA Fragmentation ELISA (Roche Applied Science), Cell Death Detection ELISAPLUS (Roche Applied Science), In Situ Cell Death Detection Kit (Roche Applied Science), DeadEnd Fluorometirc or Colorimetric TUNEL System (Promega), APO-BrdU TUNEL Assay Kit (Invitrogen), and TUNEL Apoptosis Detection Kit (Upstate).
[0663] Other assays to assess apoptosis include, for example, cell permeability assays that evaluate the loss of membrane integrity. For example, to determine whether the antibody is able to induce cell death, loss of membrane integrity as evaluated by uptake of propidium iodide (PI), trypan blue, or 7-aminoactinomycin D (7AAD) can be assessed relative to untreated cells. In addition, commercial kits such as APOPercentage Assay (Biocolor Assays) can be used to measure apoptosis. Annexin V assays also can be employed. Annexin V binds to phosphatidylserine, which is normally found on the inner surface of the cytoplasmic membrane. During apoptosis, phosphatidylserine is translocated to the outer surface and can be detected by Annexin V. For example, standard binding assays using a fluorescent labeled Annexin V can be used (e.g. Annexin V, Alex Fluor 350 Conjugate from Invitrogen). Apoptosis also can be measured by assessing the presence of other markers of apoptosis, assessing protein cleavage, and/or by mitochondrial and ATP/ADP assays. Such assays are routine and known to one of skill in the art.
[0664] For example, apoptosis analysis can be used as an assay to identify functional antibodies using cell lines, such as RKO or HCT116, or other cells expressing a target protein of interest. The cells can be co-transfected with a construct containing a marker gene, such as a gene that encodes green fluorescent protein, or a cell tracker dye. The apoptotic change can be determined using methods known in the art, such as DAPI staining and TUNEL assay using fluorescent microscope. For TUNEL assay, commercially available kit can be used (e.g., Fluorescein FragEL DNA Fragmentation Detection Kit (Oncogene Research Products, Cat. #QIA39) and Tetramethyl-rhodamine-5-dUTP (Roche, Cat. #1534 378)). Cells contacted with an antibody can exhibit, e.g., an increased apoptosis compared to control.
[0665] Cell death in vitro can be determined in the absence of complement and immune effector cells to distinguish cell death induced by antibody dependent cellular cytotoxicity (ADCC) or complement dependent cytotoxicity (CDC). Thus, the assay for cell death can be performed using heat inactivated serum (i.e. in the absence of complement) and in the absence of immune effector cells.
[0666] 3. Targets
[0667] In the screening methods provided herein, the antibodies are screened for any activity, such as binding or other functional activity, against a target. The activity can be an agonist or antagonist activity of the target. The screening assays can be designed based on any contemplated target. Exemplary of such targets include membrane-bound proteins, receptors and ligands thereof; ion channels; G-protein coupled receptors; novel epitopes; and non-protein antigens. Any activity can be assessed and is a function of the target of interest. One of skill in the art is familiar with the activities of various targets, and can choose a screening assay based on such known activities. Many of these activities for exemplary targets are exemplified herein below. Binding activity can be assessed for all targets.
[0668] Membrane-Bound Proteins, Receptors and Ligands Thereof
[0669] Exemplary targets are membrane-bound proteins, receptors and ligands thereof that can play important roles in, among other things, the formation, differentiation and maintenance of multicellular organisms. Identifying antibodies that interfere with these functions is contemplated. For example, the fate of many individual cells, e.g., proliferation, migration, differentiation, or interaction with other cells, is typically governed by information received from other cells and/or the immediate environment. This information is often transmitted by secreted polypeptides (for instance, mitogenic factors, survival factors, cytotoxic factors, differentiation factors, neuropeptides, and hormones) which are, in turn, received and interpreted by diverse cell receptors or membrane-bound proteins. These secreted polypeptides or signaling molecules normally pass through the cellular secretory pathway to reach their site of action in the extracellular environment. Thus, identification of antibodies against any one or more of such targets (receptor or ligand) can modulate a pathway important in a disease, thereby modulating the disease and ameliorating symptoms thereof.
[0670] Such membrane-bound proteins and cell receptors include, but are not limited to, cytokine receptors, receptor kinases, receptor phosphatases, receptors involved in cell-cell interactions, and cellular adhesin molecules like cadherins and integrins, and ligands of any such receptors. Exemplary of such targets include, membrane bound receptors, such as cell surface receptors, including, but are not limited to, a VEGFR-1, VEGFR-2, VEGFR-3 (vascular endothelial growth factor receptors 1, 2, and 4), a epidermal growth factor receptor (EGFR), ErbB-2, ErbB-b3, IGF-R1, C-Met (also known as hepatocyte growth factor receptor; HGFR), DLL4, DDR1 (discoidin domain receptor), KIT (recetpr for c-kit), FGFR1, FGFR2, FGFR4 (fibroblast growth factor receptors 1, 2, and 4), RON (recepteur d'origine nantais; also known as macrophage stimulating 1 receptor), TEK (endothelial-specific receptor tyrosine kinase), TIE (tyrosine kinase with immunoglobulin and epidermal growth factor homology domains receptor), CSF1R (colngly stimulating factor 1 receptor), PDGFRB (platelet-derived growth factor receptor B), EPHA1, EPHA2, EPHB1 (erythropoietin-producing hepatocellular receptor A1, A2 and B1), TNF-R1, TNF-R2, HVEM, LT-.beta.R, CD20, CD3, CD25, NOTCH, G-CSF-R, GM-CSF-R and EPO-R. Other targets include membrane-bound proteins such as selected from among a cadherin, integrin, CD52 or CD44. Exemplary ligands that can be targets of the screening methods herein, include, but are not limited to, VEGF-A, VEGF-B, VEGF-C, VEGF-D, PIGF, EGF, HGF, TNF-.alpha., LIGHT, BTLA, lymphotoxin (LT), IgE, G-CSF, GM-CSF and EPO.
[0671] One of skill in the art is familiar with the activity and functions of various target proteins. Thus, screening assays (e.g. binding or functional assays), such as any described above, for example, binding, proliferation or apoptosis assays, can be chosen to screen antibody libraries herein. Table 18A provides a summary of the activities and known function of exemplary target proteins, and exemplary assays that can be employed to screen such target proteins. In another example, antibodies or fragments thereof can be screened for binding affinity for a target protein. Binding can be to a known epitope in a target protein or to a novel epitope, which can be identified as described herein below. Table 18B provides examples of known epitopes for exemplary target proteins recognized by therapeutic antibodies. The sections that follow also further exemplify the activity and function of target proteins, including exemplary screening assays and antibodies identified thereby. It is understood that similar assays can be employed against any target protein of interest to identify antibodies that exhibit high affinity to the target protein and/or otherwise modulate the activity of a target protein.
TABLE-US-00020 TABLE 18A TARGET PROTEINS Existing Target SEQ ID NO Screening Assay Functions/Activity Antibody Drugs CD20 2011 Apoptosis of B cell lymphoma cell lines B-cell lymphoma Rituxan expressing CD20 on the surface HER2/Neu 1999 Binding to HER2/Neu receptor and Breast cancer Herceptin receptor inhibition of cancer cell growth VEGFA 2012 Binding Colorectal cancer, Avastin, non-small lung cancer. Lucentis Neovascular age- related macular degeneration EGFR 2000 Binding Colorectal, head and Erbitux, neck, metastatic cancers Vectibix CD52 2013 Binding to CD52 on T and B cells B-cell chronic Campath lymphocytic leukemia TNF.alpha. 2014 Binding to TNF.alpha. blocks its interaction Rheumatoid arthritis Humira, with p55 and p75 receptors, reducing the Remicade level of inflammation markers, CRP, ESR, IL6 CD25 2015 Binding to CD25 (IL2 receptor) inhibits Prophylaxis against Simulect, lymphocyte activation allograft rejection Zenapax CD3 2016 (delta) Binding blocks T cell function Allograft rejection, Orthoclone 2017 (gamma) acute renal and 2018 (zeta) hepatic 2019 (epsilon) IgE 2020 (constant) Binding to high affinity IgE reduces Asthma Xolair 2021 (FcER1) activation of mast cells, basophils and 2022 (FcERII) release of inflammatory mediators IIb/IIIa 2023 (IIIa) Binding to the receptor inhibits platelet Prevention of ReoPro integrin 2024 (IIb) aggregation cardiac receptor ischaemia EPO-R 2009 EPO binding activates STAT and Anaemia Epogen stimulates erythropoiesis G-CSF 2025 G-CSF binding stimulates neutrophill Neutropaenia Neupogen, receptor proliferation Neulasta GM-CSF 2026 GM-CSF binding stiumulates proliferation Leukopaenia Lukine receptor and differentiation of neutrophils, eosinophils and monocytes
TABLE-US-00021 TABLE 18B Epitopes SEQ TARGET ANTIBODY AMINO ACID SEQUENCE ID NO DLL4 YW26.82 ECIPHNGCRHGTCSTPWQCTCDEGWGGLFCD 2029 (252-282) EpoR Mab 71 PGNYSFSYQLEDEPWKLCRLHQAPTARGAV 2030 and Mab 73 (49-78 ErbB2 702 and 7F3 STQVCTGTDMKLRLPASPETHLDMLRHLYQG 2031 C (22-53) 4D5 LPCHPECQPQNGSVTCFGPEADQCVACAHYK 2032 DPPFCVARCPSGVKPDLSYMPIWKFPDEEGA CQP (561-625) 3H4 VEECRVLQGLPREYVNARHCLPCHPECQPQN 2033 GSVTCFGPEADQCVACAHYKDPPFCVAR (541-599) Her2/ Herceptin PEADQ(557-561)+DPPF(570-573)+KF 2034 Neu PDEEGACQP (593-603) C24 Q156+H245+YF(252,257)+T268+DVGS 2035 (Pertuzumab) CTPLH (285-290, 294-296)+K311 EGFR IMC-C225 R353+Q384+QHFVS(408-409, 412, 2036 (Erbitux) 417-418)+ISK(438, 440, 443)+ KIISN(465-468, 473) EGFR antibody KK(293-294)+G+NYVVTD(298-303) 2037 [1i8k] VEGFA Fab12 (Avastin) [fy(17, 21)]+[yk(45, 48)+qiMRIkh 2038 qGQhiGEM(79-84, 86-94)] G6 [FMYQY(17, 18, 21, 2 2, 25)+Del 2039 [(63, 64, 66)+cp(104, 106)]+ k48+mlHqgQhl(81, 83, 86-91) B20-4 [kFMDYqRYCH(16-19, 21-23, 25-27)+ 2040 cndI(61-63, 66)+nkec(100, 101, 103, 104)]+[e30+k48+qmiQi (79, 81, 83, 89, 91) Fab 12 variant Y(71)+K(74)+QIMRIK(105-110)+ 2041 Y0317 HQGQHIGEM(112-120)+F(43)+Y(47) Fab 12 variant F(43)+Y(47)+Y(71)+K(74)+QIMRIK 2042 Y0317 (105-110)+HQGQHIGEM(112-120) CD52 Campath G+TSSPSA(32-37)+D 2043 CD20 C2h7 EPANPSEK 168-175 2044 C2b8 (Rituxan) I(164)+NCEPANPSE(166-174) 2045 C2b8 (Rituxan) IYNCEPANP(164-172)+K(175) 2046 CD40 5c8 EASS(129-132)+EKGYY(142-146)+ 2047 C178+0218+SQVSHG(245-250) CD41 10E5 RNVGSQ(77-82)+N149+N158+SRLWH 2048 (GPIIb) (206, 208, 213-215) TPO TN1 ETKAQ(57-61)+RGALQSLLGTQLPPQ 2049 (98, 102-115)
[0672] i. Notch and Notch Ligands
[0673] a) Notch Proteins
[0674] The Notch proteins (Notch1, set forth in SEQ ID NO:2002; Notch 2, set forth in SEQ ID NO:2003; Notch 3, set forth in SEQ ID NO: 2004; and Notch 4 set forth in SEQ ID NO:2005) are single-pass transmembrane receptor proteins that play a crucial role in cell-to-cell communication. Cell surface receptors in the Notch family are expressed on numerous types of cells, including many types of stem cells and undifferentiated progenitor cells, either in the embryo or in self-renewing tissues after birth, see e.g. Artavanis-Tsakonas, et al., (1995) Science 268:225-32. For example, human primary macophages express all Notch receptors, with Notch 3 being selectively increased during macrophage differentiation (see e.g., Fung et al. (2007) Circulation, 115: 2948-2956). Notch 4 also is expressed specifically in endothelial cells and plays a role in angiogenesis. Notch also is expressed on lymphocytes where Notch signaling participates in lymphocyte development, maturation, activation and transformation.
[0675] There are five Notch ligands, designated Delta Like-1 (DLL-1), Delta Like-3 (DLL-3), Delta Like-4 (DLL-4), Jagged-1 and Jagged-2. When Notch is activated by a ligand, its intracellular domain is proteolytically cleaved and transported to the nucleus, along with CSL (CBF-1/Su(H)/Lag-1/RBP-J.sub..kappa.) transcription factor to activate transcription of downstream effectors. The resulting effector can repress the transcriptional activity of other genes encoding transcription factors for entry into terminal differentiation.
[0676] The Notch Signaling Pathway (NSP) is involved in many cellular processes, such as differentiation, cell fate decisions, maintenance of stem cells, cell motility, proliferation, and apoptosis in various cell types during development and tissue homeostasis. Notch signalling is dysregulated in many cancers, and faulty Notch signalling is implicated in many diseases including T-ALL (T-cell acute lymphoblastic leukemia), CADASIL, (Cerebral Autosomal Dominant Arteriopathy with Sub-cortical Infarcts and Leukoencephalopathy), Multiple Sclerosis (MS), Tetralogy of Fallot, Alagille Syndrome, multiple myeloma and other disease states thereby making Notch an important target in protein therapeutics (see e.g. U.S. Pat. No. 6,083,904).
[0677] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to a Notch receptor and/or assaying for a functional activity, for example, proliferation, cell motility or apoptosis of cells known to express a Notch receptor. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to Notch-1. Assays also include signal transduction assays such as by using an RBP-J.kappa./CBF-1 luciferase reporter assay (Fung et al. (2007) Circulation, 115: 2948-2956.) Assays can be performed in the presence or absence of a Notch ligand, for example, by coincubation of cells that express DLL4 (see e.g., Fung et al. (2007) Circulation, 115: 2948-2956) or by immobilization of a ligand (see e.g., Lefort et al. (2003) Experimental Hematology, 34:1720-1729). For example, as described in Fung et al., cells can be stably transfected with a construct expressing DLL4, and can be overlaid on human primary macrophages that express Notch and assayed in the presence of antibody library members.
[0678] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of a Notch receptor (Notch 1, Notch 2, Notch 3 and/or Notch 4) can be used for the treatment or prevention of a disease associated with expression and/or activity of Notch. The antibodies can be used as agonist antibodies or antagonist antibodies in such treatments. For example, agonist antibodies to Notch can be used to inhibit or decrease cancer cell growth or proliferation, such as in the treatment of various cancers including, but not limited to, prostate disorders and leukemia (see e.g., U.S. Pat. No. 6,689,744).
[0679] Antagonist antibodies provided herein also can be used in the treatment of a T-cell acute lymphoblastic leukemia; lymphoma; liver disease involving aberrant bascularization; diabetes; ovarian cancer; diseases involving vascular cell fate; rheumatoid arthritis; pancreatic cancer; non-small cell lung carcinoma; plasma cell neoplasmas such as multiple myeloma, plasma cell leukemia and extramedullary plasmacytoma; and neuroblastoma; and in treatment of plasma cell disorders such as multiple myeloma; angiogenesis; cancers such as sarcomas and carcinomas including but not limited to, breast cancer, colorectal cancer, lung cancer, pancreatic cancer, prostate cancer, liver cancer, ovarian cancer, head and neck cancer, skin cancer, brain cancer or blood cancer (see e.g., US20080226621, WO2008/091641, WO2005/054434). Notch signaling is associated in the development of skin, blood vessels and fat, and activated receptor can transform mammary epithelium. Hence, antagonist antibodies to Notch can be used in the treatment of breast tumors (see e.g. US20080206753).
[0680] Provided herein are antibodies that modulate the activity of Notch-1 and therefore can be used in the treatment of diseases or conditions associated with expression or activity of Notch-1. Such antibodies include those that have a VH chain and a VL chain encoded by a sequence of nucleotides compiled from germline segments, or any antibodies optimized therefrom. Exemplary of such antibodies are Fab antibodies. The antibodies further can contain a constant region. The antibodies include those that have a binding affinity for Notch-1 that is or is about 10.sup.-6 M, 10.sup.-7 M, 10.sup.-8 M, 10.sup.-6 M, 10.sup.-10 M, 10.sup.-11 M, 10.sup.-12 M or lower, in particular any that have a nanomolar or sub-nanomolar binding affinity.
[0681] Anti-Notch-1 antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GYTFTSYYMH (amino acids 26-35 of SEQ ID NO:1512); a CDRH2 is IINPSGGSTSYAQKFQG (amino acids 50-66 of SEQ ID NO:1512); a CDRH3 is EGYSSSWYDYFDY (amino acids 99-111 of SEQ ID NO:1512); a CDRH3 is EYYYGSGSYYNDYFDY (amino acids 99-114 of SEQ ID NO:1509); a CDRL1 is RASQSVSSNLA (amino acids 24-34 of SEQ ID NO:1843); a CDRL1 is RASQSVSSSYLA (amino acids 24-35 of SEQ ID NO:1833); a CDRL1 is RASQSISSWLA (amino acids 24-34 of SEQ ID NO:1841); a CDRL2 is GASTRAT (amino acids 50-56 of SEQ ID NO:1843); a CDRL2 is GASSRAT (amino acids 51-57 of SEQ ID NO:1833); a CDRL2 is DASSLES (amino acids 50-56 of SEQ ID NO:1841); a CDRL3 is QQYNNWPPWT (amino acids 8-98 of SEQ ID NO:1843); a CDRL3 is QQYGSSPPWT (amino acids 90-99 of SEQ ID NO:1833); and a CDRL3 is QQYNSYSPWT (amino acids 89-98 of SEQ ID NO:1841). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0682] For example, antibodies that modulate an activity of Notch-1 include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV1 (e.g. any set forth in any of SEQ ID NOS:1-43); a D.sub.H germline segment that is an IGHD6 (e.g. any set forth in SEQ ID NOS: 268-271) or an IGHD3 (e.g. any set forth in SEQ ID NOS: 252-259); and a J.sub.H germline segment that is an IGHJ4 (e.g. set forth in SEQ ID NO: 278 or 279). Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.kappa. germline segment that is an IGKV1 (set forth in any of SEQ ID NOS: 286-316) or an IGKV3 (e.g. any set forth in any of SEQ ID NOS:332-350); and a J.kappa. germline segment that is an IGKJ1 (e.g. set forth in SEQ ID NO: 356). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other mutations, so long as the resulting antibody is a functional and productive antibody and binds to Notch-1 and/or modulates a functional activity.
[0683] Exemplary of antibodies against Notch-1 include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is a IGHV1-46 (e.g. an IGHV1-46*01, IGHV1-46*02, or an IGHV1-46*03); a D.sub.H germline segment that is an IGHD3-10 (e.g. IGHD3-10*01, IGHD3-10*02) or an IGHD6-13 (e.g. IGHD6-13*01); and a JH germline segment that is an IGHJ4 (e.g. IGHJ4*01, IGHJ4*02, IGHJ4*03). The VL chain is encoded by a sequence of nucleotides compiled from a V.kappa. germline segment that is an IGKV3-15 (e.g. IGKV3-15*01), IGKV3-20 (e.g. IGKV3-20*01, IGKV3-20*02) or an IGKV1-5 (e.g. IGKV1-5*01, IGKV1-5*02, IGKV1-5*03); and a J.kappa. germline segment that is a IGKJ1*01. Exemplary antibodies provided herein that modulate an activity of Notch-1 are set forth in Table 18C.
TABLE-US-00022 TABLE 18C Anti-Notch-1 Antibodies SEQ ID NO SEQ ID NO Heavy Chain Germline nucleo- Amino Light Chain Germline nucleo- Amino Segments tide acid Segments tide acid IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ4*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-20*01; IGKJ1*01 1417 1833 IGHJ4*01 IGHV1-46*01; IGHD3-10*01; 1093 1509 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ4*01
[0684] b) DLL4
[0685] DLL4 (set forth in SEQ ID NO:2010) is a transmembrane protein ligand for Notch transmembrane receptors. It is expressed widely in a variety of tissues, but its expression is predominantly localized to the vasculature. DLL4 activates Notch-1 and Notch-4 receptors. It is required for normal vascular development and is expressed on tumor vessels. It is upregulated in blood vessels during tumor angiogenesis and expression is dependent on VEGF signaling. DLL4 expression on angiogenic endothelial cells acts as a negative regulator of vascular growth by acting to allow angiogenesis to productively proceed (Ridgway et al. (2006) Nature, 444:1083; Noguera-Troise et al. (2006) Nature, 444:1032). It acts to inhibit endothelial cell proliferation. Blockage of DLL4, however, is associated with increased angiogenesis characterized by sprouting and branching of blood vessels, but a decrease in vessel function, thereby resulting in decreased tumor growth (Ridgway et al. (2006) Nature, 444:1083; Noguera-Troise et al. (2006) Nature, 444:1032). Thus, DLL4 function is associated with an uncoupling of tumor growth from tumor vascular density. DLL4 also is expressed on activated macrophages exposed to proinflammatory stimuli such as lipopolysaccharide, interleukin-1.beta., Toll-like receptor 4 ligands and other proinflammatory stimuli and it's signaling through the Notch pathway plays a role in inflammatory states characterized by macrophage activation (Fung et al. (2007) Circulation, 115: 2948-2956).
[0686] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to DLL4 and/or assaying for a functional activity. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to DLL4. Assays also include binding assays to assess the inhibition of DLL4-Notch interaction in the presence of an anti-DLL4 antibody. Such assays can be used to identify antagonist antibodies. This is exemplified in Example 17. Assays for functional activity include those that assess activation of Notch signaling by DLL4 by assaying for signal transduction and/or down stream functional activities such as are described above. Activiation of Notch can be achieved, for example, by coincubation with cells that express DLL4 and/or immobilization of DLL4, and assays performed in the presence of antibody members. In such assays, for example, the effects of antibodies on endothelial cell proliferation (e.g. HUVECs) induced by DLL4 can be assessed (see e.g., Ridgway et al. (2006) Nature, 444:1083). In some examples, antibodies can be used to assess effects on cell differentiation of a cell expressing Notch. The cells can be co-cultured with cells expressing a ligand for Notch, for example, DLL4 or Jag1. To identify antibodies that promote differentiation (i.e. interfere with Notch activation), antibodies can be added to the assay. An exemplary assay is set forth in Example 18.
[0687] Hence, the antibodies identified from the libraries provided herein or the antibodies provided herein bind (such as specifically bind) DLL4, and in some embodiments, can modulate one or more aspects of DLL4-associated effects, including but not limited to any one or more of reduction or blocking of Notch receptor activation, reduction or blocking of Notch receptor downstream molecular signaling, disruption or blocking of Notch receptor binding to DLL4, and/or promotion of endothelial cell proliferation, and/or inhibition of endothelial cell differentiation, and/or inhibition of arterial differentiation, and/or inhibition of tumor vascular perfusion, and/or treatment and/or prevention of a tumor, cell proliferative disorder or a cancer; and/or treatment or prevention of a disorder associated with DLL4 expression and/or activity and/or treatment or prevention of a disorder associated with Notch receptor expression and/or activity. In some embodiments, the antibody specifically binds to DLL4. In some embodiments, the antibody specifically binds to the DLL4 extracellular domain (ECD). In some embodiments, an antibody reduces, inhibits, and/or blocks DLL4 activity in vivo and/or in vitro. In some embodiments, the antibody competes for binding with DLL4-ligand (reduces and/or blocks Notch receptor binding to DLL4).
[0688] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of DLL4 can be used for the treatment or prevention of disease states associated with expression and/or activity of DLL4, such as increased expression and/or activity or undesired expression and/or activity (see e.g., U.S. Published Application Serial No. US20080175847 and International Published PCT Appl. No. WO2008060705, WO2008091222). Treatment includes neoplastic and non-neoplastic disorders. For example, the antibodies or portions thereof can be used to treat a tumor, a cancer (e.g. colon cancer, lung cancer or breast cancer) and/or a cell proliferative disorder and/or conditions associated with angiogenesis (e.g. intraocular neovascular disease). In particular, the antibodies or portions thereof can be used in combination with anti-VEGF therapies and/or in treatments that are resistant to anti-VEGF treatment.
[0689] Angiogenesis is implicated in the pathogenesis of a variety of disorders. These include solid tumors and metastasis, atherosclerosis, retrolental fibroplasia, hemangiomas, chronic inflammation, intraocular neovascular diseases such as proliferative retinopathies, e.g., diabetic retinopathy, age-related macular degeneration (AMD), neovascular glaucoma, immune rejection of transplanted corneal tissue and other tissues, rheumatoid arthritis, and psoriasis. Folkman et al., J. Biol. Chem. 267:10931-34 (1992); Klagsbrun et al., Annu. Rev. Physiol. 53:217-39 (1991); and Garner A., "Vascular diseases," In: Pathobiology of Ocular Disease. A Dynamic Approach, Garner A., Klintworth G K, eds., 2nd Edition (Marcel Dekker, N Y, 1994), pp 1625-1710.
[0690] In the case of tumor growth, angiogenesis appears to be crucial for the transition from hyperplasia to neoplasia, and for providing nourishment for the growth and metastasis of the tumor. Folkman et al., Nature 339:58 (1989). The neovascularization allows the tumor cells to acquire a growth advantage and proliferative autonomy compared to the normal cells. A tumor usually begins as a single aberrant cell which can proliferate only to a size of a few cubic millimeters due to the distance from available capillary beds, and it can stay `dormant` without further growth and dissemination for a long period of time. Some tumor cells then switch to the angiogenic phenotype to activate endothelial cells, which proliferate and mature into new capillary blood vessels. These newly formed blood vessels not only allow for continued growth of the primary tumor, but also for the dissemination and recolonization of metastatic tumor cells. Accordingly, a correlation has been observed between density of microvessels in tumor sections and patient survival in breast cancer as well as in several other tumors. Weidner et al., N. Engl. J. Med. 324:1-6 (1991); Horak et al., Lancet 340:1120-24 (1992); Macchiarini et al., Lancet 340:145-46 (1992). The precise mechanisms that control the angiogenic switch is not well understood, but it is believed that neovascularization of tumor mass results from the net balance of a multitude of angiogenesis stimulators and inhibitors (Folkman, Nat. Med. 1(1):27-31 (1995)).
[0691] In addition, antibodies or portions thereof can be used to treat non-neoplastic disorders including, but not limited to, undesired or aberrant hypertrophy, arthritis, rheumatoid arthritis (RA), psoriasis, psoriatic plaques, sarcoidosis, atherosclerosis, atherosclerotic plaques, edema from myocardial infarction, diabetic and other proliferative retinopathies including retinopathy of prematurity, retrolental fibroplasia, neovascular glaucoma, age-related macular degeneration, diabetic macular edema, corneal neovascularization, corneal graft neovascularization, corneal graft rejection, retinal/choroidal neovascularization, neovascularization of the angle (rubeosis), ocular neovascular disease, vascular restenosis, arteriovenous malformations (AVM), meningioma, hemangioma, angiofibroma, thyroid hyperplasias (including Grave's disease), corneal and other tissue transplantation, chronic inflammation, lung inflammation, acute lung injury/ARDS, sepsis, primary pulmonary hypertension, malignant pulmonary effusions, cerebral edema (e.g., associated with acute stroke/closed head injury/trauma), synovial inflammation, pannus formation in RA, myositis ossificans, hypertrophic bone formation, osteoarthritis (OA), refractory ascites, polycystic ovarian disease, endometriosis, 3rd spacing of fluid diseases (pancreatitis, compartment syndrome, burns, bowel disease), uterine fibroids, premature labor, chronic inflammation such as IBD (Crohn's disease and ulcerative colitis), renal allograft rejection, inflammatory bowel disease, nephrotic syndrome, undesired or aberrant tissue mass growth (non-cancer), obesity, adipose tissue mass growth, hemophilic joints, hypertrophic scars, inhibition of hair growth, Osier-Weber syndrome, pyogenic granuloma retrolental fibroplasias, scleroderma, trachoma, vascular adhesions, synovitis, dermatitis, preeclampsia, ascites, pericardial effusion (such as that associated with pericarditis), and pleural effusion.
[0692] Provided herein are antibodies that modulate the activity of DLL4 and therefore can be used in the treatment of diseases or conditions associated with expression or activity of DLL4. Such antibodies include those that have a VH chain and a VL chain encoded by a sequence of nucleotides compiled from germline segments, or any antibodies optimized therefrom. Exemplary of such antibodies are Fab antibodies. The antibodies further can contain a constant region. The antibodies include those that have a binding affinity for DLL4 that is or is about 10.sup.-6 M, 10.sup.-7M, 10.sup.-8 M, 10.sup.-9M, 10.sup.-10 M, 10.sup.-11M, 10.sup.-12 M or lower, in particular any that have a nanomolar or sub-nanomolar binding affinity.
[0693] Anti-DLL4 antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GYTFTSYYMH (amino acids 26-35 of SEQ ID NO:1513); a CDRH1 is GYSFTSYWIG (amino acids 26-35 of SEQ ID NO:1803); a CDRH1 is GDSVSSNSAAWN (amino acids 26-37 of SEQ ID NO:1812); a CDRH1 is GGSFSGYYWS (amino acids 26-35 of SEQ ID NO:1779); a CDRH1 is GYTFTSYAMH (amino acids 26-35 of SEQ ID NO:1494); a CDRH1 is GYTFTSYDIN (amino acids 26-35 of SEQ ID NO:1537); a CDRH1 is GGSISSGGYYWS (amino acids 26-37 of SEQ ID NO:1761); a CDRH2 is IINPSGGSTSYAQKFQG (amino acids 50-66 of SEQ ID NO:1513); a CDRH2 is IIYPGDSDTRYSPSFQG (amino acids 50-66 of SEQ ID NO:1803); a CDRH2 is RTYYRSKWYNDYAVSVKS (amino acids 52-69 of SEQ ID NO:1812); a CDRH2 is EINHSGSTNYNPSLKS (amino acids 50-65 of SEQ ID NO:1779); a CDRH2 is WSNAGNGNTKYSQEFQG (amino acids 50-66 of SEQ ID NO:1494); a CDRH2 is WMNPNSGNTGYAQKFQG (amino acids 50-66 of SEQ ID NO:1537); a CDRH2 is (amino acids 52-67 of SEQ ID NO:1761); a CDRH3 is EEYSSSSAEYKQH (amino acids 99-111 of SEQ ID NO:1513); a CDRH3 is RGYSYGYDYFDY (amino acids 99-110 of SEQ ID NO:1803); a CDRH3 is EYYDFWSGYYTDYFD (amino acids 102-117 of SEQ ID NO:1812); a CDRH3 is EGYSSSWYDYFDY (amino acids 99-111 of SEQ ID NO:1512); a CDRH3 is ANWGDYFDY (amino acids 89-106 of SEQ ID NO:1779); a CDRH3 is ANWGYWYFDL (amino acids 99-108 of SEQ ID NO:1514); a CDRH3 is DDYGGNSDYFDY (amino acids 99-110 of SEQ ID NO:1494); a CDRH3 is EGYCSGGSCYSYWYFDL (amino acids 99-115 of SEQ ID NO:1508); a CDRH3 is EYYYGSGSYYNDYFDY (amino acids 99-114 of SEQ ID NO:1509); a CDRH3 is GGYCSSTSCYADYYYYYGMDV (amino acids 99-119 of SEQ ID NO:1537); a CDRH3 is EGYCSGGSCYSYWYFDL (amino acids 100-116 of SEQ ID NO:1761); a CDRL1 is RASQSVSSYLA (amino acids 24-34 of SEQ ID NO:1850); a CDRL1 is GLSSGSVSTSYYPS (amino acids 23-36 of SEQ ID NO:1881); a CDRL1 is TLRSGINLGSYRIF (amino acids 23-36 of SEQ ID NO:1884); a CDRL1 is RASQSVSSNLA (amino acids 24-34 of SEQ ID NO:1843); a CDRL1 isRASQGISSWLA (amino acids 24-34 of SEQ ID NO:1849); a CDRL1 is RASQSVSSSYLA (amino acids 24-35 of SEQ ID NO:1833); a CDRL1 is RASQSISSWLA (amino acids 24-34 of SEQ ID NO:1841); a CDRL1 is RSSQSLLDSDDGNTYLD (amino acids 24-40 of SEQ ID NO:1853); a CDRL1 is TGTSSDVGGYNYVS (amino acids 23-36 of SEQ ID NO:1864); a CDRL1 is TLSSDLSVGGKNMF (amino acids 23-36 of SEQ ID NO:1886); a CDRL2 is DASNRAT (amino acids 50-56 of SEQ ID NO:1850); a CDRL2 is STNTRSS (amino acids 52-58 of SEQ ID NO:1881); a CDRL2 is YYSDSSK (amino acids 52-58 of SEQ ID NO:1884); a CDRL2 is GASTRAT (amino acids 50-56 of SEQ ID NO:1843); a CDRL2 is AASSLQS (amino acids 50-56 of SEQ ID NO:1849); a CDRL2 is GASSRAT (amino acids 51-57 of SEQ ID NO:1833); a CDRL2 is DASSLES (amino acids 50-56 of SEQ ID NO:1841); a CDRL2 is TLSYRAS (amino acids 56-62 of SEQ ID NO:1853); a CDRL2 is EVSNRPS (amino acids 52-58 of SEQ ID NO:1864); a CDRL2 is HYSDSDK (amino acids 52-58 of SEQ ID NO:1886); a CDRL3 is QQRSNWPPWT (amino acids 89-98 of SEQ ID NO:1850); a CDRL3 is VLYMGSGISYV (amino acids 91-101 of SEQ ID NO:1881); a CDRL3 is MIWHSSASFV (amino acids 97-106 of SEQ ID NO:1884); a CDRL3 is QQYNNWPPWT (amino acids 89-98 of SEQ ID NO:1843); a CDRL3 is QQANSFPPWT (amino acids 89-98 of SEQ ID NO:1849); a CDRL3 is QQYGSSPPWT (amino acids 90-99 of SEQ ID NO:1833); a CDRL3 is QQYNSYSPWT (amino acids 89-98 of SEQ ID NO:1841); a CDRL3 is MQRIEFPSWT (amino acids 95-104 of SEQ ID NO:1853); a CDRL3 is SSYTSSSTLFV (amino acids 91-101 of SEQ ID NO:1864); and a CDRL3 is QVYESSANFV (amino acids 89-98 of SEQ ID NO:1886). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0694] For example, antibodies that modulate an activity of DLL4 include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV1 (e.g. any set forth in any of SEQ ID NOS:10-43), an IGHV4 (e.g. any set forth in SEQ ID NOS: 153-224), an IGHV5 (e.g. any set forth in SEQ ID NOS: 225-232) or an IGHV6 (e.g., any set forth in any of SEQ ID NOS: 233 or 234); a D.sub.H germline segment that is an IGHD6 (e.g. any set forth in SEQ ID NOS: 268-271), an IGHD5 (e.g. any set forth in any of SEQ ID NOS: 264-267); an IGHD4 (e.g. any set forth in SEQ ID NOS: 260-263); an IGHD2 (e.g. any set forth in SEQ ID NOS: 244-251), an IGHD3 (e.g. any set forth in any of SEQ ID NOS: 252-259) an IGHD6 (e.g. any set forth in SEQ ID NO: 268-271), or an IGHD7 (e.g. set forth in SEQ ID NO:272); and a J.sub.H germline segment that is an IGHJ1 (e.g., set forth in SEQ ID NO:273), an IGHJ2 (set forth in SEQ ID NO:274), an IGHJ4 (e.g. any set forth in any of SEQ ID NOS: 277-279), or an IGHJ6 (e.g. any set forth in SEQ ID NOS: 282-285). Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.kappa. germline segment that is an IGKV1 (e.g. any set forth in any of SEQ ID NOS: 286-316), an IGKV2 (e.g. any set forth in SEQ ID NOS: 317-331), or an IGKV3 (e.g. any set forth in any of SEQ ID NOS:332-350) and a J.kappa. germline segment that is an IGKJ1 (e.g. set forth in SEQ ID NO: 356); or from a V.sub..lamda. germline segment that is an IGLV2 (e.g. any set forth in any of SEQ ID NOS:380-399), IGLV8 (e.g. any set forth in any of SEQ ID NOS: 436-438), IGLV11 (e.g. any set forth in any of SEQ ID NO: 379), or a IGLV5 (e.g. any set forth in any of SEQ ID NOS: 424-431) and a J.sub..lamda. germline segment that is a IGLJ1 (e.g. set forth in SEQ ID NO:442) or an IGLJ4 (e.g. set forth in SEQ ID NO:446). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other nucleotide mutations, so long as the resulting antibody is a functional and productive antibody and binds to DLL4 and/or modulates a functional activity.
[0695] Exemplary of antibodies against DLL4 include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is an IGHV1-3 (e.g. IGHV1-3*01, IGHV1-3*02), an IGHV1-8*01, an IGHV1-46 (e.g. an IGHV1-46*01, IGHV1-46*02 or an IGHV1-46*03), an IGHV4-31 (e.g. IGHV4-31*01, IGHV4-31*02, IGHV4-31*03, IGHV4-31*04, IGHV4-31*05, IGHV4-31*06, IGHV4-31*07, IGHV4-31*08, IGHV4-31*09, IGHV4-3110), an IGHV4-34 (e.g. IGHV4-34*01, IGHV4-34*02, IGHV4-34*03, IGHV4-34*04, IGHV4-34*05, IGHV4-34*06, IGHV4-34*07, IGHV4-34*08, IGHV4-34*09, IGHV4-34*10, IGHV4-34*11, IGHV4-34*12, IGHV4-34*13), an IGHV5-51 (e.g., an IGHV1-5-51*01, IGHV1-5-51*02, IGHV1-5-51*03, IGHV1-5-51*04 or IGHV1-5-51*05) or is an IGHV6-1 (e.g. an IGHV6-1*01 or an IGHV6-1*02); a D.sub.H germline segment that is an IGHD2-2 (e.g. IGHD2-2*01, IGHD2-2*02), an IGHD2-15*01, an IGHD4-23*01, an IGHD6-6 (e.g. IGHD6-6*01), an IGHD6-13*01, an IGHD5-18 (e.g. an IGHD5-18*01), an IGHD3-3 (e.g. an IGHD3-3*01 or IGHD3-3*02), an IGHD3-10 (e.g. IGHD3-10*01, IGHD3-10*02), or an IGHD7-27*01; and a J.sub.H germline segment that is a IGHJ1*01, IGHJ2*01, IGHJ4*01, IGHJ4*02, IGHJ4*03, or an IGHJ6 (e.g. IGHJ6*01, IGHJ6*02, IGHJ6*03, IGHJ6*04). The VL chain is encoded by a sequence of nucleotides compiled from a V.kappa. germline segment that is an IGKV1-5 (e.g. IGKV1-5*01, IGKV1-5*02, IGKV1-5*03), an IGKV1-12 (e.g IGKV1-12*01, IGKV1-12*02), an IGKV2-D-40*01, an IGKV3-11 (e.g. IGKV3-11*01 or IGKV3-11*02), an IGKV3-15*01, an IGKV3-20 (e.g. IGKV3-20*01, IGKV3-20*02) and a J.kappa. germline segment that is a IGKJ1*01; or is compiled from a V.sub..lamda. germline segment that is an IGLV2-14 (e.g. IGLV2-14*01, IGLV2-14*02, IGLV2-14*03, IGLV2-14*04), an IGLV8-61 (e.g. IGLV8-61*01, IGLV8-61*02 or IGLV8-61*03), an IGLV5 (e.g. IGLV5-48*01), or an IGLV11-55*01 and a J.sub..lamda. germline segment that is a IGLJ1*01 or IGLJ4*01. Exemplary antibodies provided herein that modulate an activity of DLL4 are set forth in Table 18D.
TABLE-US-00023 TABLE 18D Anti-DLL4 Antibodies SEQ ID NO SEQ ID NO Heavy Chain Germline nucleo- Amino Light Chain Germline nucleo- Amino Segments tide acid Segments tide acid IGHV1-46*01; IGHD6-6*01; 1097 1513 IGKV3-11*01; IGKJ1*01 1434 1850 IGHJ1*01 IGHV5-51*03; IGHD5-18*01 > 3; 1387 1803 IGLV8-61*01; IGLJ1*01 1465 1881 IGHJ4*01 IGHV6-1*01; IGHD3-3*01; 1396 1812 IGLV5-48*01; IGLJ4*01 1468 1884 IGHJ4*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ4*01 IGHV4-34*01; IGHD7-27*01; 1363 1779 IGKV1-12*01; IGKJ1*01 1433 1849 IGHJ4*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-20*01; IGKJ1*01 1417 1833 IGHJ4*01 IGHV1-3*02; IGHD4-23*01; 1078 1494 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ4*01 IGHV1-46*01; IGHD2-15*01; 1092 1508 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ2*01 IGHV1-46*01; IGHD3-10*01; 1093 1509 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ4*01 IGHV1-8*01; IGHD2-2*01; 1121 1537 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ6*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV2D-40*01; IGKJ1*01 1437 1853 IGHJ4*01 IGHV4-34*01; IGHD7-27*01; 1363 1779 IGLV2-14*01; IGLJ4*01 1448 1864 IGHJ4*01 IGHV4-31*02; IGHD2-15*01; 1345 1761 IGLV2-14*01; IGLJ4*01 1448 1864 IGHJ2*01 IGHV4-34*01; IGHD7-27*01; 1363 1779 IGLV11-55*01; IGLJ4*01 1470 1886 IGHJ4*01
[0696] Anti-DLL4 antibodies provided herein include antibodies that are optimized compared to any of the identified anti-DLL4 germline Hits. The antibodies include one or more mutations in the VH chain and/or one or more mutations in the VL chain compared to an identified germline Hit. For example, the antibodies can contain 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 or more amino acid replacements compared to the corresponding antibody germline Hit. The mutations can be in the VH chain, for example, in any one or more of the amino acid residues of a V.sub.H, D.sub.H or J.sub.H region. Alternatively, or in addition, the mutations can be in the VL chain, for example, in any one or more of the amino acid residues of the V.sub.L or J.sub.L region. Optimized antibodies containing one or more mutations exhibit improved activity compared to the parent antibody (e.g. germline Hit not containing the modifications). The antibodies are optimized to exhibit an improved functional activity, either agonistic or antagonistic, against the DLL4 target protein. In other examples, the antibodies are optimized to exhibit an improved binding affinity for DLL4. Generally, an activity or binding affinity is increased by at or about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 30-fold, 40-fold, 50-fold, 60-fold, 70-fold, 80-fold, 90-fold, 100-fold, 200-fold, 300-fold, 400-fold, 500-fold, 600-fold, 700-fold, 800-fold, 900-fold, 1000-fold or more compared to an activity or binding affinity of the parent antibody (e.g. germline Hit not containing the modification(s)). For example, as described in the Examples, optimized anti-DLL4 antibodies provided herein exhibit a binding affinity that is improved by at least 100-fold compared to the parent antibody. Such antibodies exhibit a nanomolar binding affinity.
[0697] For example, provided herein are optimized anti-DLL4 antibodies that contain one or more mutations in the VH chain of an anti-DLL4 Hit, for example, any set forth in any of SEQ ID NOS: 1494, 1508-1509, 1512-1513, 1537, 1761, 1779, 1803 and 1812. In one example, the one or more mutations include mutations in the D.sub.H region. For example, an anti-DLL4 antibody provided herein can include a VH chain containing a germline segment compiled from a variant of a IGHD5-18*01 that encodes a D.sub.H region that contains a mutation(s) at position G1 and/or G5 corresponding to amino acid residues set forth in SEQ ID NO:3736 (also corresponding to amino acid residues G100 and/or G104 set forth in the VH chain of the anti-DLL4 Hit set forth in SEQ ID NO: 1803). The mutation can be to any other amino acid residue, in particular, the mutation is a lysine (K), Arginine (R), threonine (T). Exemplary of such mutations are G1K, G1R, and/or G5T corresponding to amino acid replacements in the D.sub.H region set forth in SEQ ID NO:3736 (also corresponding to amino acid replacements G100K, G100R, and/or G104T in the VH chain of the anti-DLL4 Hit set forth in SEQ ID NO:1803). In another example, an anti-DLL4 antibody provided herein can include a VH chain containing germline segments compiled from a variant of IGHD6-6*01 that encodes a D.sub.H region that contains a mutations(s) at positions S3, S4 and/or S5 corresponding to amino acid residues set forth in SEQ ID NO:3737 (also corresponding to amino acid residues S102, S103 and/or S104 in the VH chain of the anti-DLL4 Hit set forth in SEQ ID NO: 1513. The mutation can be to any other amino acid residue, in particular, the mutation is a alanine (A), proline (P) or phenylalanine (F). Exemplary of such mutations are 53A, S4P, S5F and/or S5A corresponding to amino acid replacements in the D.sub.H region set forth in SEQ ID NO:3737 (also corresponding to amino acid replacements S102A, S103P, S104F and S104A in the VH chain of the anti-DLL4 Hit set forth in SEQ ID NO:1513. Table 18E lists exemplary anti-DLL4 antibody variants containing one or more mutations in the D.sub.H region of the VH chain of anti-DLL4 Hits.
[0698] Also provided herein are optimized anti-DLL4 antibodies containing one or more mutations in the J.sub.H region of the VH chain of an anti-DLL4 hit. For example, an anti-DLL4 antibody provided herein can include a VH chain containing a germline segment compiled from a variant of an IGHJ1*01 that encodes a J.sub.H region that contains a mutation at position H6 corresponding to amino acid residue set forth in SEQ ID NO:3738 (also corresponding to amino acid residue H111 in the VH chain of the anti-DLL4 Hit set forth in SEQ ID NO:1513). The mutation can be to any other amino acid residue, in particular, the mutation is a phenylalanine (F) or a tyrosine (Y). Exemplary of such mutations are H6F and H6Y corresponding to amino acid replacements in the J.sub.H region set forth in SEQ ID NO:3738 (also corresponding to amino acid replacements H111F and H111Y in the J.sub.H region of the VH chain of the anti-DLL4 Hit set forth in SEQ ID NO:1513). Table 18E lists exemplary anti-DLL4 antibody variants containing one or more mutations in the J.sub.H region of the VH chain of anti-DLL4 Hits.
[0699] Optimized anti-DLL4 antibodies provided herein also can contain one or more amino acid mutations in the VL chain. For example, provided herein are optimized anti-DLL4 antibodies containing one or more mutation in the V.kappa. region of the VL chain of an anti-DLL4 hit. For example, an anti-DLL4 antibody provided herein can include a VL chain containing a germline segment compiled from a variant of an IGKV3-11 (e.g. IGKV3-11*01 or IGKV3-11*02) that encodes a V.kappa. region that contains a mutation at position S28, S30 and/or S31 corresponding to amino acid residues set forth in SEQ ID NO:3739 (also corresponding to amino acid residues S28, S30 and/or S31 in the VL chain of the anti-DLL4 Hit set forth in SEQ ID NO:1850). The mutation can be to any other amino acid residue, in particular, the mutation is a proline (P), asparagine (N), or lysine (K). Exemplary of such mutations are S28P, S30N and/or S31K corresponding to amino acid replacements in the V.kappa. region set forth in SEQ ID NO:3739 (also corresponding to amino acid replacements S28P, S30N and/or S31K in the V.kappa. region of the VL chain of the anti-DLL4 Hit set forth in SEQ ID NO:1850). Table 18E lists exemplary anti-DLL4 antibody variants containing one or more mutations in the V.kappa. region of the VL chain of anti-DLL4 Hits.
TABLE-US-00024 TABLE 18E Anti-DLL4 antibody variants SEQ ID NO Parent anti SEQ ID NO Parent anti-DLL4 Muta- Nucleo- Amino DLL4 VL Muta- Nucleo- Amino VH chain Hit tion(s) tide acid chain Hit tion(s) tide acid IGHV5-51*03, G100K 3741 3720 IGLV8-61*01; n/a 1465 1881 IGHD5-18*01 > 3; IGLJ1*01 IGHJ4*01 IGHV5-51*03, G100R 3742 3721 IGLV8-61*01; n/a 1465 1881 IGHD5-18*01 > 3, IGLJ1*01 IGHJ4*01 IGHV5-51*03, G104T 3745 3724 IGLV8-61*01; n/a 1465 1881 IGHD5-18*01 > 3, IGLJ1*01 IGHJ4*01 IGHV5-51*03, G100K/ 3749 3728 IGLV8-61*01; n/a 1465 1881 IGHD5-18*01 > 3, G104T IGLJ1*01 IGHJ4*01 IGHV1-46*01, S104F 3743 3722 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, IGKJ1*01 IGHJ1*01 IGHV1-46*01, S104A 3744 3723 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, IGKJ1*01 IGHJ1*01 IGHV1-46*01, S103P 3746 3725 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, IGKJ1*01 IGHJ1*01 IGHV1-46*01, S102A 3747 3726 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, IGKJ1*01 IGHJ1*01 IGHV1-46*01, S102A/ 3748 3727 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104F IGHV1-46*01, S102A/ 3750 3729 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104F/ H111F IGHV1-46*01, S102A/ 3751 3730 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104F/ H111Y IGHV1-46*01, S102A/ 3752 3731 IGKV3-11*01; n/a 1434 1850 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104Y/ H111Y IGHV1-46*01, S102A/ 3748 3727 IGKV3-11*01; S28P 3753 3735 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104F IGHV1-46*01, S102A/ 3748 3727 IGKV3-11*01; S30N 3754 3733 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104F IGHV1-46*01, S102A/ 3748 3727 IGKV3-11*01; S31K 3755 3734 IGHD6-6*01, S103P/ IGKJ1*01 IGHJ1*01 S104F
[0700] ii. ErbB Family
[0701] Group I receptor tyrosine kinases, including EGFR (ErbB-1), HER2/c-neu (ErbB-2), Her 3 (ErbB-3) and Her 4 (ErbB-4), are widely expressed in epithelial, mesenchymal, and neuronal tissues and play fundamental roles in proliferation and differentiation. They are activated by a family of ligands that variously bind to the receptors. For example, epidermal growth factor (EGF), transforming growth factor alpha (TGF-alpha) and amphiregulin bind to ErbB1, but not to other receptors. The neuregulins bind to ErbB3 and ErbB4. Finally, b-cellulin (BTC), heparin-binding EGF and epiregulin bind to ErbB1 and ErbB4. ErbB2 has no characterized ligand, but can be activated by homodimerization in trans by heterodimerization with another ErbB family member.
[0702] a) Epidermal Growth Factor Receptor
[0703] (EGFR)
[0704] The epidermal growth factor receptor (EGFR; ErbB-1; HER1 in humans; set forth in SEQ ID NO:2000) is a cell-surface receptor for members of the epidermal growth factor family (EGF-family) of extracellular protein ligands. Upon binding by the ligand epidermal growth factor (EGF), EGFR dimerizes stimulating its intrinsic intracellular protein-tyrosine kinase activity. This autophosphorylation elicits downstream activation and signaling by several other proteins that associate with the phosphorylated tyrosines through their own phosphotyrosine-binding SH2 domains. These downstream signaling proteins initiate several signal transduction cascades, including MAPK, Akt and JNK pathways, leading to DNA synthesis and cell proliferation. Such proteins modulate phenotypes such as cell migration, adhesion, and proliferation. Therefore, mutations affecting EGFR expression or activity can result in cancer
[0705] Upregulation of EGFR is associated with poor cancer prognosis. ERBITUX.RTM. (cetuximab) is a chimeric monoclonal antibody approved for the treatment of colorectal and/or head and neck cancers. ERBITUX.RTM. binds the EGFR and thereby prevents intracellular signaling associated with DNA synthesis and cell proliferation. VECTIBIX.RTM. (panitumumab) is a fully human monoclonal antibody approved for the treatment of EGFR-expressing, metastatic colorectal carcinoma. Both ERBITUX.RTM. and VECTIBIX.RTM. can be used alone or in conjunction with a chemotherapeutic agent.
[0706] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to EGFR and/or assaying for a functional activity, for example, signal transduction, cell migration, adhesion, and proliferation. Functional assays can be performed in the presence or absence of the EGF ligand. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to EGFR.
[0707] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of EGFR can be used for the treatment or prevention of disease states associated with expression and/or activity of EGFR. For example, the antibodies or portions thereof can be used in the treatment of cancers including, but not limited to, glioblastoma, head and neck cancer, pancreatic cancer, colorectal cancer, lung cancer, cancer of the nervous system, gastrointestinal cancer, prostate cancer, ovarian cancer, breast cancer, kidney cancer, retina cancer, skin cancer, liver cancer, genital-urinary cancer, bladder cancer, or a lung cancer such as a lun adenocarcinoma, lung squamous cell carcinoma or non-small cell lung cancer.
[0708] Provided herein are antibodies that modulate the activity of EGFR and therefore can be used in the treatment of diseases or conditions associated with expression or activity of EGFR. Such antibodies include those that have a VH chain and a VL chain encoded by a sequence of nucleotides compiled from germline segments, or any antibodies optimized therefrom. Exemplary of such antibodies are Fab antibodies. The antibodies further can contain a constant region. The antibodies include those that have a binding affinity for EGFR that is or is about 10.sup.-6 M, 10.sup.-7 M, 10.sup.-8 M, 10.sup.-9 M, 10.sup.-10 M, 10.sup.-11 M, 10.sup.-12 M or lower, in particular any that have a nanomolar or sub-nanomolar binding affinity.
[0709] Anti-EGFR antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GYTFTSYYMH (amino acids 26-35 of SEQ ID NO:1508); a CDRH2 is IINPSGGSTSYAQKFQG (amino acids 50-66 of SEQ ID NO:1508); a CDRH3 is EGYCSGGSCYSYWYFDL (amino acids 99-115 of SEQ ID NO:1508); a CDRH3 is EGYSSSWYDYFDY (amino acids 99-111 of SEQ ID NO:1512); a CDRL1 is RASQSVSSNLA (amino acids 24-34 of SEQ ID NO:1843); a CDRL1 is RASQSISSWLA (amino acids 24-34 of SEQ ID NO:1841); a CDRL2 is GASTRAT (amino acids 50-56 of SEQ ID NO:1843); a CDRL2 is DASSLES (amino acids 50-56 of SEQ ID NO:1841); a CDRL3 is QQYNNWPPWT (amino acids 89-98 of SEQ ID NO:1843); and a CDRL3 is QQYNSYSPWT (amino acids 89-98 of SEQ ID NO:1841). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0710] For example, antibodies that modulate an activity of EGFR include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV1 (e.g. any set forth in any of SEQ ID NOS:1-43); a D.sub.H germline segment that is an IGHD6 (e.g. any set forth in SEQ ID NOS: 268-271) or an IGHD2 (e.g. any set forth in SEQ ID NOS: 244-251); and a J.sub.H germline segment that is an IGHJ2 (e.g., set forth in SEQ ID NO:274) or an IGHJ4 (e.g. set forth in SEQ ID NO: 278 or 279). Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.kappa. germline segment that is an IGKV1 (set forth in any of SEQ ID NOS: 286-316) or an IGKV3 (e.g. any set forth in any of SEQ ID NOS:332-350); and a J.kappa. germline segment that is an IGKJ1 (e.g. set forth in SEQ ID NO: 356). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other mutations, so long as the resulting antibody is a functional and productive antibody and binds to EGFR and/or modulates a functional activity.
[0711] Exemplary of antibodies against EGFR include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is a IGHV1-46 (e.g. an IGHV1-46*01, IGHV1-46*01, or an IGHV1-46*03); a D.sub.H germline segment that is an IGHD2-15 (e.g. IGHD2-15*01) or an IGHD6-13 (e.g. IGHD6-13*01); and a JH germline segment that is an IGHJ2*01 or an IGHJ4 (e.g. IGHJ4*01, IGHJ4*02, IGHJ4*03). The VL chain is encoded by a sequence of nucleotides compiled from a V.kappa. germline segment that is an IGKV1-5 (e.g. IGKV1-5*01, IGKV1-5*02, IGKV1-5*03) or an IGKV3-15 (e.g. IGHV3-15*01); and a J.kappa. germline segment that is a IGKJ1*01. Exemplary antibodies provided herein that modulate an activity of EGFR are set forth in Table 18F.
TABLE-US-00025 TABLE 18F Anti-EGFR Antibodies SEQ ID NO SEQ ID NO Heavy Chain Germline nucleo- Amino Light Chain Germline nucleo- Amino Segments tide acid Segments tide acid IGHV1-46*01; IGHD2-15*01; 1092 1508 IGKV3-15*01; IGKJ1*01 1427 1841 IGHJ2*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-15*01; IGKJ1*01 1427 1841 IGHJ4*01 IGHV1-46*01; IGHD2-15*01; 1092 1508 IGKV1-5*01; IGKJ1*01 1425 1843 IGHJ2*01
[0712] b) Human Epidermal Growth Factor Receptor 2 (HER2/Neu)
[0713] HER2/neu (ErbB-2; set forth in SEQ ID NO: 1999) is a cell membrane surface-bound receptor tyrosine kinase normally involved in the signal transduction pathways leading to cell growth and differentiation. ErbB-2 is thought to be an orphan receptor, with none of the EGF family of ligands able to activate it. However, ErbB receptors dimerize on ligand binding, and ErbB-2 is the preferential dimerization partner of other members of the ErbB family. ErbB2 activation leads to kinase activation and cell proliferation. ErbB-2 is notable for its role in the pathogenesis of breast cancer and therefore as a target for treatment. In fact, ErbB-2 protein overexpression is observed in 25-30% of primary breast cancers. HERCEPTIN.RTM. (trastuzumab) is a recombinant DNA-derived humanized monoclonal antibody, used to treat breast cancer, that selectively binds to the extracellular domain of ErbB-2 Thus, ErbB-2 is an attractive target for additional protein therapeutics. See e.g. Carter et al, (1992) Proc. Natl. Acad. Sci. USA, 89:4285-4289; and U.S. Pat. No. 5,725,856.
[0714] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to ErbB-2 and/or assaying for a functional activity, for example, signal transduction, cell migration, adhesion, and proliferation. For example, cells that are known to express ErbB-2 can be assessed for proliferation in the presence or absence of an antibody or portion thereof. Alternatively, reporter system assays can be constructed, whereby the expression of a reporter protein, such as luciferase, is dependent on the activiation of ErbB2 (see e.g. Ueda et al. (2004) J Biol. Chem., 279:24505-24513). Assays can be performed in the presence of EGF, TGF or other ErbB ligands that are ligands for ErbB-2 binding partners. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to ErbB-2.
[0715] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of ErbB-2 can be used for the treatment or prevention of disease states associated with expression and/or activity of ErbB-2. For example, the antibodies or portions thereof can be used in the treatment of proliferative diseases such as cancers, including, but not limited to, breast cancer, ovarian cancer, colon cancer, lung cancer and prostate cancer.
[0716] Provided herein are antibodies that modulate the activity of ErbB-2 and therefore can be used in the treatment of diseases or conditions associated with expression or activity of ErbB-2. Such antibodies include those that have a VH chain and a VL chain encoded by a sequence of nucleotides compiled from germline segments, or any antibodies optimized therefrom. Exemplary of such antibodies are Fab antibodies. The antibodies further can contain a constant region. The antibodies include those that have a binding affinity for ErbB-2 that is or is about 10.sup.-6 M, 10.sup.-7 M, 10.sup.-8M, 10.sup.-9 M, 10.sup.-10 M, 10.sup.-11 M, 10.sup.-12 M or lower, in particular any that have a nanomolar or sub-nanomolar binding affinity.
[0717] Anti-ErbB-2 antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GGSISSGGYYWS (amino acids 26-37 of SEQ ID NO:1760); a CDRH1 is GYTFTSYYMH (amino acids 26-35 of SEQ ID NO:1512); a CDRH1 is GFSLSTSGVGVG (amino acids 26-37 of SEQ ID NO:1559); a CDRH1 is GGTFSSYAIS (amino acids 26-35 of SEQ ID NO:1522); a CDRH2 is YIYYSGSTYYNPSLKS (amino acids 52-67 of SEQ ID NO:1760); a CDRH2 is IINPSGGSTSYAQKFQG (amino acids 50-66 of SEQ ID NO:1512); a CDRH2 is LIYWNDDKRYSPSLKS (amino acids 52-67 of SEQ ID NO:1559); a CDRH2 is GIIPIFGTANYAQKFQG (amino acids 50-66 of SEQ ID NO:1522); a CDRH3 is EGYSSSWYDYFDY (amino acids 100-112 of SEQ ID NO:1760); a CDRH3 is GYSGSYYYWYFDL (amino acids 99-111 of SEQ ID NO:1512); a CDRH3 is EEYSSSSAEYKQH (amino acids 99-111 of SEQ ID NO:1513); a CDRH3 is RPNWGYWYFDL (amino acids 100-110 of SEQ ID NO:1559); a CDRH3 is GYNWNDDYYYYYGMDV (amino acids 99-114 of SEQ ID NO:1522); a CDRL1 is RASQSVSSSYLA (amino acids 24-35 of SEQ ID NO:1833); a CDRL1 is KSSQSVLYSSNNKNYLA (amino acids 24-40 of SEQ ID NO:1838); a CDRL1 is RASQSVSSNLA (amino acids 24-34 of SEQ ID NO:1843); a CDRL1 is RSSQSLVYSDGNTYLN (amino acids 24-39 of SEQ ID NO:1828); a CDRL2 is GASSRAT (amino acids 51-57 of SEQ ID NO:1833); a CDRL2 is WASTRES (amino acids 56-62 of SEQ ID NO:1838); a CDRL2 is GASTRAT (amino acids 50-56 of SEQ ID NO:1843); a CDRL2 is KVSNDRS (amino acids 55-61 of SEQ ID NO:1828); a CDRL3 is QQYGSSPPWT (amino acids 90-99 of SEQ ID NO:1833); a CDRL3 is QQYYSTPPWT (amino acids 95-104 of SEQ ID NO:1838); a CDRL3 is QQYNNWPPWT (amino acids 89-98 of SEQ ID NO:1843); and a CDRL3 is MQGTHWPPWT (amino acids 94-103 of SEQ ID NO:1828). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0718] For example, antibodies that modulate an activity of ErbB-2 include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV4 (e.g. any set forth in any of SEQ ID NOS:153-224), an IGHV1 (e.g. any set forth in SEQ ID NOS: 10-43), or an IGHV2 (e.g. any set forth in SEQ ID NOS:44-67); a D.sub.H germline segment that is an IGHD6 (e.g. any set forth in SEQ ID NOS: 268-271); an IGHD1 (e.g. any set forth in any of SEQ ID NOS: 239-243), or an IGHD7 (e.g. set forth in SEQ ID NO:272); and a J.sub.H germline segment that is an IGHJ1 (e.g., set forth in SEQ ID NO:273), an IGHJ2 (e.g. set forth in SEQ ID NO:274), an IGHJ4 (e.g. set forth in any of SEQ ID NOS: 277-279), or an IGHJ6 (e.g. set forth in any of SEQ ID NOS:282-285). Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.kappa. germline segment that is a IGKV3 (e.g. any set forth in any of SEQ ID NOS:332-350), a IGKV4 (e.g. set forth in SEQ ID NO:351), or an IGKV2 (e.g. any set forth in SEQ ID NOS:317-331); and a J.kappa. germline segment that is an IGKJ1 (e.g. set forth in SEQ ID NO: 356). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other mutations, so long as the resulting antibody is a functional and productive antibody and binds to ErbB-2 and/or modulates a functional activity.
[0719] Exemplary of antibodies against ErbB-2 include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is a IGHV4-31 (e.g. an IGHV4-31*01, IGHV4-31*02, IGHV4-31*03, IGHV4-31*04, IGHV4-31*05, IGHV4-31*06, IGHV4-31*07, IGHV4-31*08, IGHV4-31*09, IGHV4-3110), an IGHV1-46 (e.g., IGHV1-46*01, IGHV1-46*02, IGHV1-46*03), an IGHV2-5 (e.g. IGHV2-5*01, IGHV2-5*02; IGHV2-5*03, IGHV2-5*04, IGHV2-5*05, IGHV2-5*06, IGHV2-5*07, IGHV2-5*08, IGHV2-5*09, IGHV2-510) IGHV1-69 (e.g. IGHV1-69*01, IGHV1-69*02, IGHV1-69*03, IGHV1-69*04, IGHV1-69*05, IGHV1-69*06, IGHV1-69*07, IGHV1-69*08, IGHV1-69*09, IGHV1-69*10, IGHV1-69*11, IGHV1-69*12, IGHV1-69*13); a D.sub.H germline segment that is a IGHD6-6 (e.g. IGHD6-6*01), an IGHD6-13 (e.g. IGHD6-13*01), an IGHD1-26 (e.g. IGHD1-26*01), an IGHD7-27*01, or an IGHD1-1*01; and a JH germline segment that is an IGHJ1*01, an IGHJ2*01, an IGHJ4 (e.g. IGHJ4*01, IGHJ4*02, IGHJ4*03) or an IGHJ6 (e.g. IGHJ6*01, IGHJ6*02, IGHJ6*03, IGHJ6*04). The VL chain is encoded by a sequence of nucleotides compiled from a V.kappa. germline segment that is a IGKV3-20 (e.g. IGHV3-20*01 or IGHV3-20*02), a IGKV4-1 (e.g. IGKV4-1*01), an IGKV3-15*01, or an IGKV2-30*01; and a J.kappa. germline segment that is a IGKJ1*01. Exemplary antibodies provided herein that modulate an activity of ErbB-2 are set forth in Table 18G.
TABLE-US-00026 TABLE 18G Anti-ErbB2 Antibodies SEQ ID NO SEQ ID NO Heavy Chain Germline nucleo- Amino Light Chain Germline nucleo- Amino Segments tide acid Segments tide acid IGHV4-31*02; IGHD1-26*01; 1344 1760 IGKV3-20*01; IGKJ1*01 1417 1833 IGHJ2*01 IGHV1-46*01, IGHD6-6*01 1097 1513 IGKV4-1*01; IGKJ1*01 1422 1838 and IGHJ1*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV4-1*01; IGKJ1*01 1422 1838 IGHJ4*01 IGHV2-5*01; IGHD7-27*01; 1143 1559 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ2*01 IGHV1-69*06; IGHD1-1*01; 1106 1522 IGKV2-30*01; IGKJ1*01 1412 1828 IGHJ6*01
[0720] iii. IGF-R1 (Insulin-Like Growth Factor 1 Receptor)
[0721] Insulin-like Growth Factor 1 Receptor (IGF-R1; set forth in SEQ ID NO:2007) is a transmembrane receptor activated by Insulin-like Growth Factor 1 (IGF-1) and Insulin-like Growth Factor 2 (IFG-2). Overexpression of insulin-like Growth Factor Receptor-I has been demonstrated in several cancer cell lines and tumor tissues. IGFR1 is overexpressed in 40% of all breast cancer cell lines (Pandini, et al., (1999) Cancer Res. 5: 1935) and in 15% of lung cancer cell lines. In breast cancer tumor tissue, IGFR1 is overexpressed 6-14 fold and IGFR1 exhibits 2-4 fold higher kinase activity as compared to normal tissue (Webster, et al., (1996) Cancer Res, 56: 2781 and Pekonen, et al., (1998) Cancer Res. 48: 1343). Moreover, colorectal cancer tissue has been reported to exhibit strongly elevated IGFR1 levels (Weber et al., Cancer 95 (10): 2086-95 (2002)). Analysis of primary cervical cancer cell cultures and cervical cancer cell lines revealed 3- and 5-fold overexpression of IGFR1, respectively, as compared to normal ectocervical cells (Steer, et al., (1996) Cancer Res. 56: 1762). Expression of IGFR1 in synovial sarcoma cells also correlated with an aggressive phenotype (i.e., metastasis and high rate of proliferation; Xie, et al., (1999) Cancer Res. 59: 3588).
[0722] Activation of IGF-R1 causes survival and proliferation of mitosis-competent cells and growth in tissues such as skeletal muscle and cardiac muscle. The IGF-1 receptor is implicated in several cancers, most notably breast cancer. IGF-R1 can serve to increase the metastatic potential of the tumor by inferring the ability of the tumor to promote vascularization. In addition, IGF-R1's anti-apoptotic properties allow cancer cells to evade the cytotoxic properties of chemotherapeutic drugs or radiation. Crosstalk can occur between IGF-R1 and EGFR allowing EGFR signaling to resume, even in the presence of EGFR inhibitors. Inhibition of IGF-1R mediated signaling has been shown to reduce tumor growth rate, increase apoptosis, and increase killing of tumors by chemotherapy and other molecular target therapies.
[0723] Experimental approaches undertaken to inhibit IGF-1R function in tumors have provided encouraging but limited success, and their effectiveness in treating cancer is yet to be determined in the clinic. The ability of an antibody to inhibit IGF-R1 function was first demonstrated with a mouse monoclonal antibody (.alpha.-IR3) targeting an unknown epitope in the a subunit of IGF-1R (Kull et al, (1983) J. Biol. Chem. 258:6561-66). Subsequently other antibodies developed to the a subunit of IGF-1R have been shown to inhibit IGF-R1 function to varying degrees in different experimental cancer models. There remains a need for IGF-1R antibodies with different or improved binding, efficacy, and safety characteristics for the treatment of various neoplastic diseases including cancer and metastases thereof.
[0724] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to IGF-R1 and/or assaying for a functional activity, for example, cell proliferation. Assays can be performed in the presence of IGF-1 or IGF-2. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to IGF-R1.
[0725] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of IGF-R1 can be used for the treatment or prevention of disease states associated with expression and/or activity of IGF-R1. For example, the antibodies or portions thereof can be used in the treatment of rheumatoid arthritis, Grave's disease, multiple sclerosis, systemic lupus erythematosus, Hashimoto's Thyroiditis, Myasthenia Gravis, autoimmune thyroiditis; Bechet's disease; acromegaly; bladder cancer, Wilm's cancer, ovarian cancer, pancreatic cancer, benign prostatic hyperplasia, breast cancer, prostate cancer, bone cancer, lung cancer, colorectal cancer, cervical cancer, synovial sarcoma, diarrhea associated with metastatic carcinoid, vasoactive intestinal peptide secreting tumors; gigantism, psoriasis; atherosclerosis, smooth muscle restenosis of blood vessels or inappropriate microvascular proliferation.
[0726] iv. C-Met
[0727] C-Met (or hepatocyte growth factor receptor, HGFR; set forth in SEQ ID NO: 2001) is a membrane receptor found in cells of epithelial origin, including stem cells and progenitor cells. Upon stimulation of c-Met by its ligand, hepatocyte growth factor (HGF), c-Met induces several biological responses that trigger invasive growth, including mitogenesis, motogenesis and morphogenesis. C-Met also is expressed in tumor cell lines and in various human solid tumors. Abnormal c-Met activation in cancer cells correlates with poor prognosis and triggers tumor growth, angiogenesis and metastasis. C-Met engages multiple oncogenic signal transduction pathways, including RAS, which leads to morphogenesis; P13K, which is associated with cell motility; STAT, which induces branching morphogenesis; and beta catenin, which participates in transcriptional regulation of numerous genes. HGF, through c-Met, has also been shown to be a mitogen for certain cell types, including melanocytes, renal tubular cells, keratinocytes, certain endothelial cells and cells of epithelial origin [Matsumoto et al, Biochem. Biophys. Res. Commun. 176: 45-51 (1991); Igawa et al., Biochem. Biophys. Res. Commun. 174: 831-838 (1991); Han et al, Biochem., 30: 9768-9780 (1991); Rubin et al., Proc. Natl. Acad. Sci. USA, 88: 415-419 (1991)].
[0728] Several cancer therapies involve interference of c-Met signaling. These therapies include kinase inhibitors, that prevent ATP from binding to c-Met preventing transphosphorylation; HGF inhibitors, that prevent HGF activation of c-Met; decoy MET inhibitors, that prevent ligand binding and homodimerization; and immunotherapy, including passive immunotherapy which activates CDC or ADCC and active immunotherapy with cytokines triggering nonspecific stimulation of immune cells. In view of the important role that this pathway plays in the etiology of various pathological conditions, however, it is clear that there continues to be a need for agents that have clinical attributes that are optimal for development as therapeutic agents.
[0729] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to c-Met and/or assaying for a functional activity, for example, cell proliferation or cell signaling. Assays can be performed in the presence of HGF. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to c-Met.
[0730] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of c-Met can be used for the treatment or prevention of disease states associated with expression and/or activity of c-Met. For example, the antibodies or portions thereof can be used in the treatment of lung cancer, bone cancer, pancreatic cancer, skin cancer, cancer of the head and neck, cutaneous or intraocular melanoma, uterine cancer, ovarian cancer, rectal cancer, cancer of the anal region, stomach cancer, colon cancer, breast cancer, gynecologic tumors (e. g., uterine sarcomas, carcinoma of the fallopian tubes, carcinoma of the endometrium, carcinoma of the cervix, carcinoma of the vagina or carcinoma of the vulva), Hodgkin's disease, cancer of the esophagus, cancer of the small intestine, cancer of the endocrine system (e.g., cancer of the thyroid, parathyroid or adrenal glands), sarcomas of soft tissues, cancer of the urethra, cancer of the penis, prostate cancer, chronic or acute leukemia, solid tumors of childhood, lymphocytic lymphomas, cancer of the bladder, cancer of the kidney or ureter (e. g., renal cell carcinoma, carcinoma of the renal pelvis), or neoplasms of the central nervous system (e. g., primary CNS lymphoma, spinal axis tumors, brain stem gliomas or pituitary adenomas).
[0731] Anti-c-Met antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GFTFSSYAMS (amino acids 26-35 of SEQ ID NO:3353); a CDRH2 is SISGSGGSTYYADSVKG (amino acids 50-66 of SEQ ID NO:3353); a CDRH3 is EHIVWIAISYYYYYYGMDV (amino acids 99-118 of SEQ ID NO:3353); a CDRH3 is EDIVVVPAAMSYYYYYYGMDV (amino acids 99-119 of SEQ ID NO:3347); a CDRH3 is EDIVLMVYAISYYYYYYGMDV (amino acids 99-119 of SEQ ID NO:3349); a CDRH3 is EDIVVVVAATSYYYYYYGMDV (amino acids 99-119 of SEQ ID NO:3351); a CDRL1 is QGDSLRSYYAS (amino acids 22-33 of SEQ ID NO:1870); a CDRL2 is GKNNRPS (amino acids 49-55 of SEQ ID NO:1870); and a CDRL3 is NSRDSSGNHLW (amino acids 88-99 of SEQ ID NO:1870). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0732] For example, antibodies that modulate an activity of c-Met include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV3 (e.g. any set forth in SEQ ID NOS: 68-152); a D.sub.H germline segment that is an IGHD2 (e.g., any set forth in any of SEQ ID NOS:244-251); and a J.sub.H germline segment that is an IGHJ6 (e.g. any set forth in SEQ ID NOS: 282-285), or is a modified form of a J.sub.H germline segment, for example, set forth in SEQ ID NO: 3455. Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.lamda. germline segment that is an IGLV3 (e.g. set forth in SEQ ID NO: 400-417); and a J.lamda. germline segment that is an IGLJ2 (e.g. set forth in SEQ ID NO: 443). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other nucleotide mutations, so long as the resulting antibody is a functional and productive antibody and binds to c-Met and/or modulates a functional activity.
[0733] Exemplary of antibodies against c-Met include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is an IGHV3-23 (e.g. IGHV3-23*01, IGHV3-23*02, IGHV3-23*03, IGHV3-23*04 or IGHV3-23*05); a D.sub.H germline segment that is an IGHD2-15*01, an IGHD2-2 (e.g. IGHD2-2*01, IGHD2-2*02, or IGHD2-2*03), an IGHD2-8 (e.g. IGHD2-8*01 or IGHD2-8*02), or an IGHD2-21 (e.g. IGHD2-21*01 or IGHD2-21*02); and a JH germline segment that is an IGHJ6 (e.g. IGHJ6*01, IGHJ6*02, IGHJ6*03, IGHJ6*04), or any modified form thereof, for example, set forth in SEQ ID NOS: 3455. The VL chain is encoded by a sequence of nucleotides compiled from a V.lamda. germline segment that is an IGLV3-19*01; and a J.lamda. germline segment that is an ILGJ2*01. Exemplary antibodies provided herein that modulate an activity of EpoR are set forth in Table 18H.
TABLE-US-00027 TABLE 18H Anti-HGFR Antibodies SEQ ID NO SEQ ID NO Heavy Chain Germline nucleo- Amino Light Chain Germline nucleo- Amino Segments tide acid Segments tide acid IGHV3-23*01; IGHD2-21*01 > 3; 2663 3353 IGLV3-19*01; IGLJ2*01 1454 1870 IGHJ6*01 IGHV3-23*01; IGHD2-2*01 > 3; 2657 3347 IGLV3-19*01; IGLJ2*01 1454 1870 IGHJ6*01 IGHV3-23*01; IGHD2-8*01 > 3; 2659 3349 IGLV3-19*01; IGLJ2*01 1454 1870 IGHJ6*01 IGHV3-23*01; IGHD2-15*01 > 3; 2661 3351 IGLV3-19*01; IGLJ2*01 1454 1870 IGHJ6*01
[0734] v. CD20--B-Lymphocyte Antigen
[0735] CD20 (human B-lymphocyte-restricted differentiation antigen, Bp35; set forth in SEQ ID NO: 2011), is a hydrophobic transmembrane protein located on pre-B and mature B lymphocytes. CD20 is found on the surface of greater than 90% of B cells from peripheral blood or lymphoid organs and is expressed during early pre-B cell development and remains until plasma cell differentiation. CD20 is present on both normal B cells as well as malignant B cells. In particular, CD20 is expressed on greater than 90% of B cell non-Hodgkin's lymphomas (NHL) (Anderson et al. (1984) Blood 63(6):1424-1433), but is not found on hematopoietic stem cells, pro-B cells, normal plasma cells, or other normal tissue (Tedder et al (1985) J. Immunol. 135(2):973-979). CD20 also is expressed on tumor cells, e.g. NHL.
[0736] CD20 regulates an early step in the activation process for cell cycle initiation and differentiation. CD20 functions as an ion channel and operates as a store of calcium facilitating entry of extracellular calcium following BCR-induced emptying of intracellular stores (see e.g., Teeling et al. (2006) J Immunol., 177:362-371). Due to the expression of CD20 on almost all B-cells, but not stem cells, it is a target for antigenic modulation via mAb induced antibody-dependent cellular cytotoxicity and complement-dependent cytotoxicity. In addition, when engaged by antibody, CD20 initiates signaling that can control growth and triggering cell death in tumors (Teeling et al. (2006) J Immunol., 177:362-371).
[0737] CD20 has been validated as a cancer cell target by RITUXAN.RTM. (Rituximab), a genetically engineered chimeric murine/human monoclonal antibody directed against the CD20 antigen found on the surface of normal and malignant B lymphocytes (see e.g. U.S. Pat. No. 5,736,137). RITUXAN.RTM. has been shown to induce B-cell lysis through apoptosis, complement-dependent cytotoxicity (CDC) and ADCC.
[0738] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to CD20 and/or assaying for a functional activity, for example, complement-dependent cell-mediate cytotoxicity and killing of cells and apoptosis assays of cells expressing CD20. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to CD20. Example 11 exemplifies a lymphoma apoptosis assay for assessing the function of cross-linked Fab antibodies.
[0739] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of CD20 can be used for the treatment or prevention of disease states associated with expression and/or activity of CD20. For example, the antibodies or portions thereof can be used in the treatment of lymphomas, autoimmune diseases and transplant rejections (e.g, to prevent rejection of organ and tissue grafts by suppressing autoimmune responses.) Lymphomas include, but are not limited to, non-Hodgkin's lymphomas (high-grade lymphomas, intermediate grade lymphomas, and low grade lymphomas), Hodgkin's disease, acute lymphoblastic leukemias, myelomas, chronic lymphocytic leukemias, and myeloblastic leukemias. Autoimmune diseases include, but are not limited to, systemic lupus erythematosus (SLE), rheumatoid arthritis, Crohn's disease, psoriasis, autoimmune thrombocytopenic purpura, multiple sclerosis, ankylosing spondylitis, myasthenia gravis, and pemphigus vulgaris.
[0740] vii. Erythropoietin Receptor (Epo-R)
[0741] Erythropoietin (Epo; set forth in SEQ ID NO: 2009) is a glycoprotein hormone that induces proliferation and differentiation of erythroid progenitor cells. Epo is responsible for promoting the growth, differentiation and survival of erythroid progenitors, which give rise to mature red blood cells. In response to changes in the level of oxygen in the blood and tissues, erythropoietin appears to stimulate both proliferation and differentiation of immature erythroblasts. It also functions as a growth factor, stimulating the mitotic activity of erythroid progenitor cells, such as erythrocyte burst forming and colony-forming units. It also acts as a differentiation factor, triggering transformation of an erythrocyte colony-forming-unit into a proerythroblast (See Erslev, A., New Eng. J. Med., 316:101-103 (1987)).
[0742] The activity of Epo is mediated through the binding and activation of a cell surface receptor referred to as the erythropoietin receptor (EpoR). In the absence of ligands the Epo receptor exists in a preformed dimer. The binding of Epo to its receptor causes a conformational change such that the cytoplasmic domains are placed in close proximity. While not completely understood, it is believed that this "dimerization" plays a role in the activation of the receptor. The activation of the Epo receptor results in a number of biological effects. Some of these activities include stimulation of proliferation, stimulation of differentiation and inhibition of apoptosis (See U.S. Pat. No. 6,319,499, Liboi et al., PNAS USA, 90:11351 (1993), Koury, Science, 248:378 (1990)). Defects in the erythropoietin receptor can produce erythroleukemia and familial erythrocytosis.
[0743] Erythropoietin is an important pharmaceutical for use in a variety of therapies where stimulation of red blood cell proliferation (RBC) is desired. Epogen.RTM. (epoetin alfa) is a recombinant erythropoietin used to stimulate red blood cell proliferation and thereby treat anemia (see e.g. U.S. Pat. Nos. 4,703,008 and 5,955,422).
[0744] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to EpoR and/or assaying for a functional activity, for example, proliferation, apoptosis or cell signaling. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to EpoR. Example 12 and 18 exemplify assays to assess modulation of proliferation or apoptosis induced by EpoR. Hence, such assays can be used to identify agonist antibodies.
[0745] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of EpoR can be used for the treatment or prevention of disease states associated with expression and/or activity of EpoR. For example, the antibodies or portions thereof can be used in the treatment of disorders characterized by low red blood cell levels and/or decreased hemoglobin levels (e.g. anemia). In addition, such antibodies or portions thereof can be used for treating disorders characterized by decreased or subnormal levels of oxygen in the blood or tissue, such as, for example, hypoxemia or chronic tissue hypoxia and/or diseases characterized by inadequate blood circulation or reduced blood flow. Antibodies or antigen-binding portions thereof also can be useful in promoting wound healing or for protecting against neural cell and/or tissue damage, resulting from brain/spinal cord injury, stroke and the like. Non-limiting examples of conditions that can be treatable by the antibodies include anemia, such as chemotherapy-induced anemia, cancer associated anemia, anemia of chronic disease, HIV-associated anemia, bone marrow transplant-associated anemia and the like, heart failure, ischemic heart disease and renal failure.
[0746] Provided herein are antibodies that modulate the activity of EpoR and therefore can be used in the treatment of diseases or conditions associated with expression or activity of EpoR. Such antibodies include those that have a VH chain and a VL chain encoded by a sequence of nucleotides compiled from germline segments, or any antibodies optimized therefrom. Exemplary of such antibodies are Fab antibodies. The antibodies further can contain a constant region. The antibodies include those that have a binding affinity for EpoR that is or is about 10.sup.-6 M, 10.sup.-7 M, 10.sup.-8 M, 10.sup.-9 M, 10.sup.-10 M, 10.sup.-11M, 10.sup.-12 M or lower, in particular any that have a nanomolar or sub-nanomolar binding affinity.
[0747] Anti-EpoR antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GYTFTSYYMH (amino acids 26-35 of SEQ ID NO:1509); a CDRH1 is SGYSISSSNWWG (amino acids $26-37 of SEQ ID NO:1759); a CDRH1 is GGSISSGGYYWS (amino acids 26-37 of SEQ ID NO:1769); a CDRH1 is GFTFSSYAMS (amino acids 26-35 of SEQ ID NO:3359); a CDRH2 is IINPSGGSTSYAQKFQG (amino acids 50-66 of SEQ ID NO:1509); a CDRH2 is YIYYSGSTYYNPSLKS (amino acids 51-66 of SEQ ID NO:1759); a CDRH2 is YIYYSGSTYYNPSLKS (amino acids 52-67 of SEQ ID NO:1769); a CDRH2 is SISGSGGSTYYADSVKG (amino acids 50-66 of SEQ ID NO:3359); a CDRH3 is EYYYGSGSYYNDYFDY (amino acids 99-114 of SEQ ID NO:1509); a CDRH3 is EGYSSSWYDYFDY (amino acids 99-111 of SEQ ID NO:1512); a CDRH3 is TNWGAEYFQH (amino acids 99-108 of SEQ ID NO:1759); a CDRH3 is ANWGDNWFDS (amino acids 100-109 of SEQ ID NO:1769); a CDRH3 is ANWGYWYFDL (amino acids 99-108 of SEQ ID NO:1514); a CDRH3 is EGYCSGGSCYSYWYFDL (amino acids 99-115 of SEQ ID NO:1508); a CDRH3 is GITMVRGVIISYYYYYYGMDV (amino acids 99-119 of SEQ ID NO:3359); a CDRL1 is RASQSVSSSYLA (amino acids 24-35 of SEQ ID NO:1833); a CDRL1 is KSSQSVLYSSNNKNYLA (amino acids 24-40 of SEQ ID NO:1838); a CDRL1 is RASQSVSSNLA (amino acids 24-34 of SEQ ID NO:1843); a CDRL1 is RASQSISSWLA (amino acids 24-34 of SEQ ID NO:1841); a CDRL1 is RSSQSLLDSDDGNTYLD (amino acids 24-40 of SEQ ID NO:1853); a CDRL1 is RASQSISSYLN (amino acids 24-34 of SEQ ID NO:1854); a CDRL1 is QGDSLRSYYAS (amino acids 23-33 of SEQ ID NO:1870); a CDRL2 is GASSRAT (amino acids 51-57 of SEQ ID NO:1833); a CDRL2 is WASTRES (amino acids 56-62 of SEQ ID NO:1838); a CDRL2 is GASTRAT (amino acids 50-56 of SEQ ID NO:1843); a CDRL2 is DASSLES (amino acids 50-56 of SEQ ID NO:1841); a CDRL2 is TLSYRAS (amino acids 56-62 of SEQ ID NO:1853); a CDRL2 is AASSLQS (amino acids 50-56 of SEQ ID NO:1854); a CDRL2 is GKNNRPS (amino acids 49-55 of SEQ ID NO:1870); a CDRL3 is QQYGSSPPWT (amino acids 90-99 of SEQ ID NO:1833); a CDRL3 is QQYYSTPPWT (amino acids 95-104 of SEQ ID NO:1838); a CDRL3 is QQYNNWPPWT (amino acids 89-98 of SEQ ID NO:1843); a CDRL3 is QQYNSYSPWT (amino acids 89-98 of SEQ ID NO:1841); a CDRL3 is MQRIEFPSWT (amino acids 95-104 of SEQ ID NO:1853); a CDRL3 is QQSYSTPPWT (amino acids 89-98 of SEQ ID NO:1854); and a CDRL3 is NSRDSSGNHLW (amino acids 88-99 of SEQ ID NO:1870). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0748] For example, antibodies that modulate an activity of EpoR include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV1 (e.g. any set forth in SEQ ID NOS: 10-43), IGHV3 (e.g. any set forth in SEQ ID NOS: 68-152), or an IGHV4 (e.g. any set forth in SEQ ID NOS: 153-224); a D.sub.H germline segment that is an IGHD6 (e.g. any set forth in SEQ ID NOS: 268-271), an IGHD3 (e.g. any set forth in any of SEQ ID NOS: 252-259), an IGHD7 (e.g. any set forth in SEQ ID NO: 272); an IGHD2 (e.g., any set forth in any of SEQ ID NOS:244-251); and a J.sub.H germline segment that is an IGHJ1 (e.g., set forth in SEQ ID NO:273), IGHJ4 (e.g. set forth in SEQ ID NO:277-279), an IGHJ5 (e.g. set forth in SEQ ID NOS: 280 or 281), an IGHJ2 (e.g. set forth in SEQ ID NOS: 274), or an IGHJ6 (e.g. any set forth in SEQ ID NOS: 282-285), or is a modified form of a JH germline segment, for example, set forth in any of SEQ ID NOS: 3450-3455. Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.kappa. germline segment that is an IGKV4 (e.g. set forth in SEQ ID NO:351), an IGKV3 (e.g. set forth in SEQ ID NOS:332-350); an IGKV1 (e.g. set forth in SEQ ID NOS:286-316 and 868); an IGKV2 (e.g. set forth in SEQ ID NOS:317-331); a J.kappa. germline segment that is an IGKJ1 (e.g. set forth in SEQ ID NO: 356); a V.lamda. germline segment that is an IGLV3 (e.g. set forth in SEQ ID NO: 400-417); and a J.lamda. germline segment that is an IGLJ2 (e.g. set forth in SEQ ID NO: 443). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other nucleotide mutations, so long as the resulting antibody is a functional and productive antibody and binds to EpoR and/or modulates a functional activity.
[0749] Exemplary of antibodies against EpoR include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is a IGHV1-46 (e.g., IGHV1-46*01, IGHV1-46*02, IGHV1-46*03), an IGHV3-23 (e.g. IGHV3-23*01, IGHV3-23*02, IGHV3-23*03, IGHV3-23*04, IGHV3-23*05), an IGHV4-28 (e.g. IGHV4-28*01, IGHV4-28*02, IGHV4-28*03, IGHV4-28*04, IGHV4-28*05), or an IGHV4-31 (e.g. IGHV4-31*01, IGHV4-31*02, IGHV4-31*03; IGHV4-31*04, IGHV4-31*05, IGHV4-31*06, IGHV4-31*07, IGHV4-31*08, IGHV4-31*09, IGHV4-3110); a D.sub.H germline segment that is a IGHD6-6 (e.g. IGHD6-6*01), an IGHD6-13 (e.g. IGHD6-13*01), an IGHD3-10 (e.g. IGHD3-10*01 or IGHD3-10*02), an IGHD7-27*01, an IGHD2-15*01, or an IGHD6-13*01; and a JH germline segment that is an IGHJ1*01, an IGHJ4 (e.g. IGHJ4*01, IGHJ4*02 or IGHJ4*03), an IGHJ5 (e.g. IGHJ5*01, IGHJ5*02), an IGHJ6 (e.g. IGHJ6*01, IGHJ6*02, IGHJ6*03, IGHJ6*04), or an IGHJ2*01, or any modified form thereof, for example, set forth in SEQ ID NOS: 3455. The VL chain is encoded by a sequence of nucleotides compiled from a V.kappa. germline segment that is an IGKV4-1 (e.g. IGKV4-1*01), an IGKV3-15*01, an IGKV3-20 (e.g. IGKV3*01, IGKV3*02), an IGKV1-5 (e.g. IGKV1-5*01, IGKV1-5*02, IGKV1-5*03), an IGKV1-39*01, or an IGKV2D-40*01; a J.kappa. germline segment that is a IGKJ1*01; a V.lamda. germline segment that is an IGLV3-19*01; and a J.lamda. germline segment that is an ILGJ2*01. Exemplary antibodies provided herein that modulate an activity of EpoR are set forth in Table 181.
TABLE-US-00028 TABLE 18I Anti-EpoR Antibodies SEQ ID NO SEQ ID NO Heavy Chain Germline nucleo- Amino Light Chain Germline nucleo- Amino Segments tide acid Segments tide acid IGHV1-46*01; IGHD3-10*01; 1093 1509 IGKV4-1*01; IGKJ1*01 1422 1838 IGHJ4*01 IGHV1-46*01; IGHD6-6*01; 1097 1513 IGKV4-1*01; IGKJ1*01 1422 1838 IGHJ1*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV4-1*01; IGKJ1*01 1422 1838 IGHJ4*01 IGHV4-28*01; IGHD7-27*01; 1343 1759 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ1*01 IGHV4-31*02; IGHD7-27*01; 1353 1769 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ5*01 IGHV1-46*01; IGHD7-27*01; 1098 1514 IGKV3-20*01; IGKJ1*01 1417 1833 IGHJ2*01 IGHV1-46*01; IGHD2-15*01; 1092 1508 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ2*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ4*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-20*01; IGKJ1*01 1417 1833 IGHJ4*01 IGHV1-46*01; IGHD2-15*01; 1092 1508 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ2*01 IGHV1-46*01; IGHD3-10*01; 1093 1509 IGKV2D-40*01; IGKJ1*01 1437 1853 IGHJ4*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV2D-40*01; IGKJ1*01 1437 1853 IGHJ4*01 IGHV3-23*01; IGHD3-10*01 > 3; 2669 3359 IGKV1-39*01; IGKJ1*01 1438 1854 IGHJ6*01 IGHV3-23*01; IGHD3-10*01 > 3; 2669 3359 IGLV3-19*01; IGLJ2*01 1454 1870 IGHJ6*01
[0750] viii. Cadherins
[0751] Cadherins are a class of calcium dependent, type-1 transmembrane glycoproteins that are involved in cell adhesion, helping to ensure that cells within tissues are bound together. There are multiple classes of cadherin molecules, including but not limited to N-cadherin, E-cadherin and P-cadherin. Members of this family exhibit calcium-dependent homophilic interactions and are responsible for the selective cell-cell recognition and adhesion, which is necessary for allocating different cell types to their proper places during organ development. Cadherins also play an important role in maintaining the integrity of multicellular structures. During embryonic morphogenesis the expression of diverse members of the cadherin family is spatially and temporally regulated facilitating the orderly assembly of various cell types into functional structures. Cadherins are considered to play a significant role in the cellular connections of cancer and metastatic cells.
[0752] a) P-Cadherin (P-cad/CDH3)
[0753] P-cadherin (P-cad; set forth in SEQ ID NO: 2008) is a single-span type-1 transmembrane glycoprotein found in the placenta that is homologous to E-cadherin (epithelial cadherin, E-cad). Both P-cad and E-cad interact with the cytoskeleton by alpha-catenin. Like other cadherins, p-cadherin plays a role in epithelial cell-cell adhesion. Other major roles include the determination of cell phenotypes and involvement in cell dynamics, including migration and the dissemination of tumor cells. The expression of P-cad in epithelial tissues appears to identify cell populations with proliferative activity, and its expression decreases as cells differentiate.
[0754] Expression of P-cadherin, a calcium-dependent cellular adhesion protein, has been reported in poorly differentiated and invasive bladder carcinoma cells. Such bladder carcinoma cells exhibit reduced E-cadherin expression. (Mialhe, A. et al., J. Urol. 164:826 (2000)). Down-regulation of E-cadherin and P-cadherin has also been associated with cultured neoplastic prostate cells. (Wang, J. et al, Urol. Res. 5:308 (2000)). The development of human colorectal cancer has been attributed, at least in part, to a decrease in cellular levels of the E-cadherin/catenin complex. (Debruyne, P. et al., Acta Gastroenterol. Belg. 62(4):393 (1999)). Aberrant up-regulation of P-cadherin was recently reported to be associated with proliferative cell phenotypes that can be related to neoplastic transformation of tissues of the gastrointestinal tract, particularly metaplastic and adenomatous polyps. (Sanders, D. S., et al., J. Pathol. 190(5):526 (2000)). Hence, certain cancer types, particularly some digestive cancer types, e.g., colon cancer, are characterized by the upregulation and the overexpression of P-cadherin relative to normal cells. P-cadherin is a valid target for cancer diagnosis, prophylaxis or therapy (see e.g. United States Published Patent Application No. 2003/0194406 and International Published Patent Application No. WO 02/097395).
[0755] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to pCad and/or assaying for a functional activity. Assays for functional activity, include but are not limited to, assays that assess proliferation and/or adhesion of tumor cells, ADCC or CDC activity, anti-apoptotic assays, or cell cycle checkpoint assays. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to pCad. Example 20 exemplifies an assay to assess effects of antibodies on cell to cell adhesion induced by p-cadherin. For example, an antibody that inhibits p-cadherin function can be identified based on the failure of cells to clump in the presence of the antibody.
[0756] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of pCad can be used for the treatment or prevention of disease states associated with expression and/or activity of pCad. For example, the antibodies or portions thereof can be used in the treatment of digestive cancers such as colon cancer, stomach cancer and liver cancer, and other cancers, such as lung cancer and breast cancer.
[0757] Provided herein are antibodies that modulate the activity of p-cadherin and therefore can be used in the treatment of diseases or conditions associated with expression or activity of p-cadherin. Such antibodies include those that have a VH chain and a VL chain encoded by a sequence of nucleotides compiled from germline segments, or any antibodies optimized therefrom. Exemplary of such antibodies are Fab antibodies. The antibodies further can contain a constant region. The antibodies include those that have a binding affinity for p-cadherin that is or is about 10.sup.-6 M, 10.sup.-7 M, 10.sup.-8 M, 10.sup.-9 M, 10.sup.-10 M, 10.sup.-11M, 10.sup.-12 M or lower, in particular any that have a nanomolar or sub-nanomolar binding affinity.
[0758] Anti-p-cadherin antibodies provided herein include antibodies having at least one CDR that is a CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and/or CDRL3. For example, a CDRH1 is GYTFTSYYMH (amino acids 26-35 of SEQ ID NO:1512); a CDRH1 is GFTFSSYAMS (amino acids 26-35 of SEQ ID NO:3559); a CDRH2 is IINPSGGSTSYAQKFQG (amino acids 50-66 of SEQ ID NO:1512); a CDRH2 is SISGSGGSTYYADSVKG (amino acids 50-66 of SEQ ID NO:3359); a CDRH3 is EGYSSSWYDYFDY (amino acids 99-111 of SEQ ID NO:1512); a CDRH3 is ANWGYWYFDL (amino acids 99-108 of SEQ ID NO:1514); a CDRH3 is EYYYGSGSYYNDYFDY (amino acids 99-114 of 1509); a CDRH3 is GITMVRGVIISYYYYYYGMDV (amino acids 99-119 of SEQ ID NO:3359); a CDRH3 is VIITPRTIVISYAFDV (amino acids 99-114 of SEQ ID NO:3071; a CDRL1 is RASQSVSSNLA (amino acids 24-35 of SEQ ID NO:1843); a CDRL1 is RASQSVSSSYLA (amino acids 24-35 of SEQ ID NO:1833); a CDRL1 is RASQSISSWLA (amino acids 24-34 of SEQ ID NO:1841); a CDRL1 is RASQSISSYLN (amino acids 24-34 of SEQ ID NO:1854); a CDRL1 is QGDSLRSYYAS (amino acids 23-33 of SEQ ID NO:1870; a CDRL2 is GASTRAT (amino acids 50-56 of SEQ ID NO:1843); a CDRL2 is GASSRAT (amino acids 51-57 of SEQ ID NO:1833); a CDRL2 is DASSLES (amino acids 50-56 of SEQ ID NO:1841); a CDRL2 is AASSLQS (amino acids 50-56 of SEQ ID NO:1854); a CDRL2 is GKNNRPS (amino acids 49-55 of SEQ ID NO:1870); a CDRL3 is QQYNNWPPWT (amino acids 89-98 of SEQ ID NO:1843); a CDRL3 is QQYGSSPPWT (amino acids 90-99 of SEQ ID NO:1833); a CDRL3 is QQYNSYSPWT (amino acids 89-98 of SEQ ID NO:1841); a CDRL3 is QQSYSTPPWT (amino acids 89-98 of SEQ ID NO:1854); and a CDRL3 is NSRDSSGNHLW (amino acids 88-99 of SEQ ID NO:1870). Also provided herein is a CDR that exhibits 60%, 65%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to any of the above CDRs.
[0759] For example, antibodies that modulate an activity of p-cadherin include any that contain a VH chain encoded by a sequence of nucleotides containing germline components compiled from a V.sub.H germline segment that is an IGHV1 (e.g. any set forth in any of SEQ ID NOS:1-43) or an IGHV3 (e.g. any set forth in any of SEQ ID NOS:68-152); a D.sub.H germline segment that is an IGHD3 (e.g. any set forth in SEQ ID NOS:252-259), an IGHD6 (e.g. any set forth in SEQ ID NOS: 268-271) or an IGHD7 (e.g. set forth in SEQ ID NO: 272); and a J.sub.H germline segment that is an IGHJ2 (e.g. set forth in SEQ ID NO:274), IGHJ3 (e.g. set forth in SEQ ID NO:275 or 276), is an IGHJ4 (e.g. set forth in SEQ ID NO: 278 or 279), an IGHJ6 (e.g. set forth in any of SEQ ID NOS: 282-285), or is a modified form of a JH germline segment, for example, set forth in any of SEQ ID NOS: 3450-3455. Such antibodies also include any that contain a VL chain encoded by a sequence of nucleotides containing germline components compiled from a V.kappa. germline segment that is an IGKV1 (e.g. any set forth in SEQ ID NOS: 286-316), or an IGKV3 (e.g. any set forth in any of SEQ ID NOS:332-350); a J.kappa. germline segment that is an IGKJ1 (e.g. set forth in SEQ ID NO: 356); a V.lamda. germline segment that is an IGLV3 (e.g. set forth in SEQ ID NO: 400-417); and a J.lamda. germline segment that is an IGLJ2 (e.g. set forth in SEQ ID NO: 443). Such antibodies also include any that are encoded by a sequence of nucleotides containing germline segments that are variants of any of the above germline segments, for example due to conservative mutations or other mutations, so long as the resulting antibody is a functional and productive antibody and binds to p-cadherin and/or modulates a functional activity.
[0760] Exemplary of antibodies against p-cadherin include an antibody where the VH chain is encoded by a sequence of nucleotides compiled from a V.sub.H germline segment that is a IGHV1-46 (e.g. an IGHV1-46*01, IGHV1-46*01, or an IGHV1-46*03) or an IGHV3-23 (e.g. IGHV3-23*01, IGHV3-23*02, IGHV3-3*03, IGHV3-23*04, IGHV3-23*05); a D.sub.H germline segment that is an IGHD3-10 (e.g. IGHD3-10*01, IGHD3-10*02) or an IGHD6-13 (e.g. IGHD6-13*01) or an IGHD7-27*01; and a JH germline segment that is an IGHJ3 (e.g. IGHJ3*01, IGHJ3*02), IGHJ4 (e.g. IGHJ4*01, IGHJ4*02, IGHJ4*03), an IGHJ6 (e.g. IGHJ6*01, IGHJ6*02, IGHJ6*03, IGHJ6*04), or an IGHJ2*01, or any modified form thereof, for example, set forth in SEQ ID NO: 3452 and 3455 and encoding a J.sub.H region set forth in SEQ ID NO:3458 and 3461, respectively. The VL chain is encoded by a sequence of nucleotides compiled from a V.kappa. germline segment that is an IGKV1-5 (e.g. IGKV1-5*01, IGKV1-5*02, IGKV1-5*03), IGKV1-39*01, an IGKV3-15 (e.g. IGKV3-15*01), IGKV3-20 (e.g. IGKV3-20*01, IGKV3-20*02) or an IGKV3-11 (e.g. IGKV3-11*01, IGKV3-11*02); a J.kappa. germline segment that is a IGKJ1*01; a V.lamda. germline segment that is an IGLV3-19*01; and a J.lamda. germline segment that is an ILGJ2*01. Exemplary antibodies provided herein that modulate an activity of p-cadherin are set forth in Table 18J.
TABLE-US-00029 TABLE 18J Anti-p-cadherin Antibodies Heavy Chain Germline SEQ ID NO SEQ ID NO Segments or Modified nucleo- Amino Light Chain Germline nucleo- Amino forms thereof tide acid Segments tide acid IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-15*01; IGKJ1*01 1427 1843 IGHJ4*01 IGHV1-46*01; IGHD6-13*01; 1096 1512 IGKV3-20*01; IGKJ1*01 1417 1833 IGHJ4*01 IGHV1-46*01; IGHD7-27*01; 1098 1514 IGKV3-11*01; IGKJ1*01 1434 1850 IGHJ2*01 IGHV1-46*01; IGHD3-10*01; 1093 1509 IGKV1-5*01; IGKJ1*01 1425 1841 IGHJ4*01 IGHV3-23*01; IGHD3-10*01 > 3; 2669 3359 IGKV1-39*01; IGKJ1*01 1438 1854 IGHJ6*01 IGHV3-23*01; IGHD3-10*01 > 1'; 2381 3071 IGKV1-39*01; IGKJ1*01 1438 1854 IGHJ3*01 IGHV3-23*01; IGHD3-10*01 > 3; 2669 3359 IGLV3-19*01; IGLJ2*01 1454 1870 IGHJ6*01
[0761] ix. CD44
[0762] CD44 (set forth in SEQ ID NO: 2006) is a cell-surface integral membrane glycoprotein involved in cell-to-cell interactions, cell adhesion and migration. Transcripts of the CD44 gene undergo complex alternative splicing resulting in many distinct functional isoform variants. CD44 is a protein which is expressed in several different isoforms on the surface of a wide variety of cell types. The smallest isoform, standard CD44 (CD44s), which is expressed by a variety of different cells, is thought to mediate cell attachment to extracellular matrix components and can transmit a co-stimulus in lymphocyte and monocyte activation. In contrast, expression of splice variants of CD44 which contain the domain v6 (CD44v6) in the extracellular region, is restricted to a subset of epithelia.
[0763] CD44 participates in a wide variety of cellular functions, including lymphocyte activation, recirculation and homing, hematopoiesis and tumor metastasis. CD44 is a receptor for hyaluronic acid and can also interact with other ligands such as E-selectin and L-selectin and other ligands such as osteopontin, collagens and matrix metalloproteinases (MMPs). MMPs degrade proteins that keep vessel walls solid allowing endothelial cells to escape into the interstitial matrix causing sprouting angiogenesis. Inhibition of MMPs prevents formation of new capillaries. Interactions of CD44 with hyaluronan mediates cell adhesion interactions. Treatments that disrupt such interactions can be used in the treatment of a number of pathologies.
[0764] Some CD44 variations are cell surface markers for breast and prostate cancer cells. Overexpression of CD44 has been linked to the growth and spread of a range of different types of malignancies, particularly lymphomas. CD44v6, as well as other variant exons (CD44v3, CD44v5, CD44v7/v8, CD44v10) has been shown to be a tumor-associated antigen with a favorable expression pattern in human tumors and normal tissues (Heider K H et al. Eur. J. Cancer 31A:2385-2391, 1995; Heider K H et al. Cancer Immunology Immunotherapy 43:245-253, 1996; Dall et al., 1996; Beham-Schmid et al., 1998; Tempfer et al., 1998; Wagner et al., 1998) and has been subject to antibody-based diagnostic and therapeutic approaches, in particular radioimmunotherapy (RIT) of tumors (Stroomer J W et al. Clin Cancer Res 6(8):3046-55, 2000, WO 95/33771, WO 97/21104). CD44 has been the target of the development of anti-cancer therapeutics (see e.g. U.S. Pat. No. 5,990,299).
[0765] Antibody libraries provided herein can be screened for modulation of an activity by assaying for binding to CD44 and/or assaying for a functional activity. Example 13 exemplifies a binding assay to screen antibody libraries provided herein to select or identify antibodies that bind to pCad.
[0766] The antibodies or portions thereof identified from the libraries provided herein that modulate an activity of CD44 can be used for the treatment or prevention of disease states associated with expression and/or activity of CD44 or a variant thereof. For example, the antibodies or portions thereof can be used in the treatment of a cancer that is a head and neck squamous cell carcinoma (SCC), esophagus SCC, lung SCC, skin SCC, breast adenocarcinoma (AC), lung AC, cervix SCC, pancreas AC, colon AC, or stomach AC. The antibodies or portions thereof also can be used in the treatment of rheumatoid arthritis.
[0767] 4. Iterative Screening
[0768] Upon identification of a "Hit" the methods described in Section C can be repeated to generate a further library for use in the screening methods herein. By virtue of the fact that antibodies are provided in an addressable format, identification of a "Hit" permits instant identification and assessment of the "structure/activity" relationship. For example, upon identification of a "Hit" the component germline segments of the encoded antibody can be immediately envisaged. A subsequent library can be generated that includes antibodies encoded by nucleic acid molecules derived from germline segments "related" to the identified "Hit". Typically, the germline segments are related by sequence similarity, but can also be related by some other shared characteristic such as a CDR, directed peptide, or other biochemical attribute.
[0769] Generally, for the generation of a further antibody library in the iterative screening method, all gene families of the germline segments contained in a "Hit" are identified. All V.sub.H, D.sub.H and J.sub.H gene family segments thereof are combined in all possible permutations, or a subset thereof as desired, to generate a recombined nucleic acid molecule encoding a VH chain. All V.sub..kappa. and J.sub..kappa., or V.sub..lamda. and J.sub..lamda. gene family segments thereof are combined in all possible permutation, or a subset thereof as desired, to generate a recombined nucleic acid molecule encoding a VL chain. Vectors and cells expressing the various paired VH and VL members, in all permutations, are generated, proteins expressed, and antibody members purified to generate a further antibody library. The further antibody library can be provided in an addressed format. The further library can be used in screening assays as described herein above to optimize the "Hit".
[0770] In addition, it is contemplated herein that because libraries can be created whereby the identify of each member is known, a library might exist already containing the exact desired "related" library members. Thus, those members can be collected to create a sub-library of related "Hits" for screening without having to reperform the method herein.
[0771] It also is contemplated that the "Hits" or any subsequent members of a further library can be mutagenized, such as by directed evolution, to further optimize the "Hit".
[0772] 5. Directed Evolution
[0773] Antibody "Hits" identified by the screening methods herein can be further modified, e.g., by mutagenesis, to provide a library of modified antibodies. The modified antibodies are then evaluated to identify one or more antibodies which have altered functional properties (e.g., improved binding, improved stability, lengthened stability in vivo, and/or improved modulation of a functional activity). A further library of modified "Hits" can be generated to select or screen for an improved activity against a target protein. The library can be screened in an addressable format or other display format as described herein above. For example, higher affinity binding proteins can be identified from the second library, e.g., by using higher stringency or more competitive binding and washing conditions. Other screening techniques can also be used.
[0774] Various approaches have been used to create mutated antibody repertoires, including chain shuffling (Marks et al. (1992) Bio/Technology 10:779 and Clackson et al. (1991) Nature 352:624), error prone PCR (Hawkins et al. (1992) J. Mol. Biol. 226:889; and Leung et al. (1989) Technique 1:11-15), use of E. coli mutator strains (Low et al. (1996) J. Mol. Biol. 260,359), recombination (see, e.g., United States Publication No. 2004-0005709), DNA shuffling using random cleavage (Stemmer (1994) Nature 389-391; termed "nucleic acid shuffling"), RACHITT.RTM. (Coco et al. (2001) Nature Biotech. 19:354), or approaches more specifically directed to the complementarity determining regions (CDRs) of the antibody molecule, like CDR walking (Barbas et al. (1994) Proc. Natl. Acad. Sci. USA 91:3809 and Yang et al. (1995) J. Mol. Biol. 254:392), site-directed mutagenesis (Zoller et al. (1987) Nucl Acids Res 10:6487-6504), cassette mutagenesis (Reidhaar-Olson (1991) Methods Enzymol. 208:564-586) and incorporation of degenerate oligonucleotides (Griffiths et al. (1994) EMBO J 13:3245).
[0775] In some implementations, the mutagenesis is targeted to regions known or likely to be at the binding interface. For example, mutagenesis can be directed to the CDR regions of the heavy or light chains as described herein. Further, mutagenesis can be directed to framework regions near or adjacent to the CDRs, e.g., framework regions, particular within ten, five, or three amino acids of a CDR junction. Additionally, mutagenesis can also be limited to one or a few of the CDRs, e.g., to make step-wise improvements.
[0776] a. Random Mutagenesis
[0777] Antibodies bind to antigens via residues in their CDRs. Consequently, mutagenesis of CDRs is widely used to improve the affinity of Fab and Fv fragments of antibodies. Random mutagenesis methods include, for example, use of E. coli XL1red, UV irradiation, chemical modification such as by deamination, alkylation, or base analog mutagens, or PCR methods such as DNA shuffling, cassette mutagenesis, site-directed random mutagenesis, error prone PCR (see e.g. U.S. Application No.: 2006-0115874) or the use of commercially available random mutagenesis kits such as, for example, GeneMorph PCR-based random mutagenesis kits (Stratagene) or Diversify random mutagenesis kits (Clontech). The Diversify random mutagenesis kit allows the selection of a desired mutation rate for a given DNA sequence (from 2 to 8 mutations/1000 base pairs) by varying the amounts of manganese (Mn.sup.2+) and dGTP in the reaction mixture. Raising manganese levels initially increases the mutation rate, with a further mutation rate increase provided by increased concentration of dGTP. Even higher rates of mutation can be achieved by performing additional rounds of PCR. All of these approaches involve the construction of expression libraries of antibodies with mutations in the CDRs and selection for better binders. Any of a variety of general approaches for directed protein evolution based on mutagenesis can be employed. Any of these, alone or in combination can be used to modify a lead antibody to achieve a desired property. Any method known in the art can be used to modify or alter a lead antibody.
[0778] i. Saturation Mutagenesis
[0779] In one exemplary embodiment, a saturation mutagenesis technique is used in which the CDR residue(s) are mutated to each of the 20 possible amino acids (see e.g., the Kunkle method, Current Protocols in Molecular Biology, John Wiley and Sons, Inc., Media Pa. and U.S. Pat. No. 6,562,594). In such a technique, a degenerate mutagenic oligonucleotide primer can be synthesized which contains randomization of nucleotides at the desired codon(s) encoding the selected amino acid(s). Exemplary randomization schemes include NNS- or NNK-randomization, where N represents any nucleotide, S represents guanine or cytosine and K represents guanine or thymine. The degenerate mutagenic primer is annealed to the single stranded DNA template and DNA polymerase is added to synthesize the complementary strand of the template. After ligation, the double stranded DNA template is transformed into E. coli for amplification. Alternatively, single amino acid changes are made using standard, commercially available site-directed mutagenesis kits such as QuikChange (Stratagene). In another embodiment, any method commonly known in the art for site specific amino acid mutation can be used.
[0780] ii. Error Prone PCR
[0781] Error prone PCR can introduce random mutations into nucleic acid sequences (See, e.g., Hawkins et al., (1992) J. Mol. Biol. 226(3):889-96). Briefly, PCR is run under conditions which compromise the fidelity of replication, thus introducing random mutations in sequences as those skilled in the art can accomplish. After generation of such random mutants, they can be placed into genetic display formats, panned and thus evaluated for activity.
[0782] iii. Cell Lines
[0783] A mutator strain of bacteria, such as E. coli mutD5 or Epicurian Coli.RTM. XL1-Red Competent cells (Stratagene, La Jolla, Calif.), can be used during plasmid replication to generate a repertoire of antibodies that include single nucleotide point mutations. The libraries of random mutants are then screened for biological activity using a genetic display format.
[0784] iv. DNA Shuffling/Antibody Chain Shuffling
[0785] DNA shuffling can be employed to modulate the activities of lead antibodies. In DNA shuffling, variant DNA molecules are generated by in vitro homologous recombination by random fragmentation of a parent DNA followed by reassembly using PCR, resulting in randomly introduced point mutations (see e.g. Stemmer, (1994) Nature 370:389-391; Stemmer (1994) Proc. Natl Acad. Sci. USA 91:10747-10751; U.S. Pat. Nos. 5,605,793, 5,811,238, and 5,830,721; and International Publication Nos. WO 95/022625 and WO 97/20078). This technique can be modified by using a family of parent DNA molecules, such as allelic variants or DNA molecules from different species, to introduce additional variability into the process. Selection or screening for the desired activity, followed by additional iterations of mutagenesis and assay provides for rapid "evolution" of sequences by selecting for desirable mutations while simultaneously selecting against detrimental changes.
[0786] In antibody chain shuffling, one selected VH sequence is paired with variety of VL sequences to create new VH/VL partners (see e.g. Kang et al. (1991) Proc. Natl. Acad. Sci. USA 88:11120-11123; Collet et al. (1992) Proc. Natl. Acad. Sci. USA 89:10026-10030; and Marks et al. (1992) Bio/Technology 10:779)
[0787] v. CDR Walking
[0788] In another example, the generation or selection of higher affinity antibodies can be carried out by CDR walking mutagenesis, which mimics the tertiary immune selection process. For example, saturation mutagenesis of the CDR's of an antibody can be used to generate one or more libraries of antibody fragments which are displayed on the surface of filamentous bacteriophage followed by the subsequent selection of the relevant antibody using immobilized antigen. Sequential and parallel optimization strategies can be used to then select the higher affinity antibody (see e.g. Barbas et al. (1994) Proc. Natl. Acad. Sci. USA 91:3809-3813; and Yang et al. (1995) J. Mol. Biol 254(3):392-403).
[0789] vi. Framework Stabilization
[0790] A mutation can be made in a framework region or constant domain to increase the half-life of the antibody. A mutation in a framework region or constant domain also can be made to alter the immunogenicity of the antibody, to provide a site for covalent or non-covalent binding to another molecule, or to alter such properties as complement fixation, FcR binding and antibody-dependent cell-mediated cytotoxicity (ADCC). A single antibody can have mutations in any one or more of the CDRs or framework regions of the variable domain or in the constant domain. See, e.g., PCT Publication No. WO 00/09560; Ewert et al. (2003) Biochemistry 42(6):1517-1528; Ewert et al. (2004) Methods 23(2):184-99; and Knappik et al. (1995) Protein Engineering 8:81-89.
[0791] 6. Epitope Mapping
[0792] Once a "Hit" is identified its binding epitope can be determined. Hence, antibody libraries provided herein can be used to identify novel epitopes, including but not limited to, previously unidentified epitopes on known antigens or novel epitopes of unknown antigens, for example, those presented by a carcinoma cell line. Methods of epitope mapping are known to one skill in the art (see e.g., Olwyn M. R. Westwood and Frank C. Hay. Epitope Mapping. Oxford University Press, 2001; Sigma Catalog #Z373990 Epitope Mapping Protocols: Methods in Molecular Biology, Vol. 66). Methods for mapping of epitopes recognized by an antibody include, but are not limited to, binding assays such as by using BIAcore or ELISA (Reineke et al. 1999), ELISPOT, prediction software, combinatorial synthesis of peptide libraries onto a planar support (i.e. protein chip) followed by exposure of the surface to the antibody, phage display of a library of peptides derived from the sequence of the protein antigen, methods using mass spectrometry methods such as methods using MALDI or amide H/D exchange (see e.g. Baerga-Ortiz et al. (2002) Protein Science, 11:1300-1308) and surface plasma resonance.
[0793] 7. In Vivo Assays of Identified Hits
[0794] Once a "Hit" is identified against a target of interest, it can be assessed in vivo assays associated with aberrant activity of the target. In general, the method involves administering an antibody to a subject, generally a non-human animal model for a disease or condition and determining the effect of the antibody on the on the disease or condition of the model animal. In vivo assays include controls, where suitable controls include a sample in the absence of the antibody. Generally a plurality of assay mixtures is run in parallel with different antibody concentrations to obtain a differential response to the various concentrations. Typically, one of these concentrations serves as a negative control, i.e. at zero concentration or below the level of detection. An antibody of interest is one that modulates, i.e., reduces or increases a symptom of the animal model disease or condition by at least about 10%, at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 80%, at least about 90%, or more, when compared to a control in the absence of the antibody. In general, an antibody of interest will cause a subject animal to be more similar to an equivalent animal that is not suffering from the disease or condition.
[0795] Non-human animals models include those induced to have a disease such as by injection with disease and/or phenotype-inducing substances prior to administration of the antibodies to monitor the effects on disease progression. Genetic models also are useful. Animals, such as mice, can be generated which mimic a disease or condition by the overexpression, underexpression or knock-out of one or more genes. Such animals can be generated by transgenic animal production techniques well-known in the art or using naturally-occurring or induced mutant strains. One of skill in the art is familiar with various animal models associated with particular targets.
[0796] Such animal model systems include, but are not limited to, mice, rats, rabbits, guinea pigs, sheep, goats, pigs, and non-human primates, e.g. baboons, chimpanzees and monkey. Any animal system well-known in the art can be used. Several aspects of the procedure can vary; said aspects include, but are not limited to, the temporal regime of administering the antibodies (e.g., prophylactic and/or therapeutic agents), whether such antibodies are administered separately or as an admixture, and the frequency of administration of the antibodies.
[0797] Recombinant (transgenic) animal models can be engineered by introducing the coding portion of the genes identified herein into the genome of animals of interest, using standard techniques for producing transgenic animals. Animals that can serve as a target for transgenic manipulation include, without limitation, mice, rats, rabbits, guinea pigs, sheep, goats, pigs, and non-human primates, e.g. baboons, chimpanzees and monkeys. Techniques known in the art to introduce a transgene into such animals include pronucleic microinjection (U.S. Pat. No. 4,873,191); retrovirus-mediated gene transfer into germ lines (e.g., Van der Putten et al., (1985) Proc. Natl. Acad. Sci. USA 82:6148-615); gene targeting in embryonic stem cells (Thompson et al., (1989) Cell 56:313-321); electroporation of embryos (Lo, (1983) Mol. Cel. Biol. 3:1803-1814); sperm-mediated gene transfer (Lavitrano et al, (1989) Cell 57:717-73). For review, see, for example, U.S. Pat. No. 4,736,866.
[0798] Animal models can be used to assess the efficacy of an antibody, a composition, or a combination therapy provided herein. Examples of animal models for lung cancer include, but are not limited to, lung cancer animal models (see e.g. Zhang et al., (1994) In Vivo 8(5):755-69) and a transgenic mouse model with disrupted p53 function (see, e.g., Morris et al, (1998) J La State Med Soc 150(4):179-85). An example of an animal model for breast cancer includes, but is not limited to, a transgenic mouse that overexpresses cyclin D1 (see, e.g., Hosokawa et al, (2001) Transgenic Res 10(5):471-8). An example of an animal model for colon cancer includes, but is not limited to, a TCR b and p53 double knockout mouse (see, e.g., Kado et al, (2001), Cancer Res 61(6):2395-8). Examples of animal models for pancreatic cancer include, but are not limited to, a metastatic model of Panc02 murine pancreatic adenocarcinoma (see, e.g., Wang et al., (2001) Int J Pancreatol 29(1):37-46) and nu-nu mice generated in subcutaneous pancreatic tumors (see, e.g., Ghaneh et al., (2001) Gene Ther 8(3):199-208). Examples of animal models for non-Hodgkin's lymphoma include, but are not limited to, a severe combined immunodeficiency ("SCID") mouse (see, e.g., Bryant et al., (2000) Lab Invest 80(4):553-73) and an IgHmu-HOX11 transgenic mouse (see, e.g., Hough et al., (1998) Proc Nati Acad Sci USA 95(23):13853-8). An example of an animal model for esophageal cancer includes, but is not limited to, a mouse transgenic for the human papillomavirus type 16 E7 oncogene (see, e.g., Herber et al., (1996) J Virol 70(3):1873-81). Examples of animal models for colorectal carcinomas include, but are not limited to, Apc mouse models (see, e.g., Fodde & Smits, (2001) Trends Mol Med 7(8):369-73 and Kuraguchi et al., (2000) Oncogene 19(50):5755-63).
[0799] Animal models for arthritis include, but are not limited to, rheumatoid arthritis rats (see e.g. Pearson, (1956) Proc. Soc. Exp. Biol. Med., 91:95-101) and collagen induced arthritis in mice and rats (see e.g. Current Protocols in Immunology, Eds. J. E. Cologan, A. M. Kruisbeek, D. H. Margulies, E. M. Shevach and W. Strober, John Wiley & Sons, Inc., 1994). An example of an animal model for asthma, includes but is not limited to, a mouse model of pulmonary hypersensitivity (see e.g. Riese et al. (1998) J. Clin. Invest. 101:2351-2363 and Shi, et al. (1999) Immunity 10:197-206). Animal models for allogenic rejection include, but are not limited to, rat allogeneic heart transplant models (see e.g. Tanabe et al (1994) Transplantation 58:23-27 and Tinubu et al (1994) J. Immunol. 153:4330-4338) and rat heterocardiac allograft rejection (Jae-Hyuck Sim et al (2002) Proc Natl Acad Sci U.S.A. 99(16):10617-10622). Steel mice are used as a model of human aplastic anemia (see e.g. Jones, (1983) Exp. Hematol., 11:571-580). An example of an animal model for anemia, includes but is not limited to, hemolytic anemia guinea pigs (see e.g. Schreiber, et al. (1972) J. Clin. Invest. 51:575). An example of an animal model for neutropenia, includes but is not limited to, neutropenia neutropenic CD rats (see, e.g. Nohynek et al. (1997) Cancer Chemother. Pharmacol. 39:259-266).
[0800] 8. Articles of Manufacture/Kits
[0801] Pharmaceutical compounds of selected antibodies or nucleic acids encoding selected antibodies, or a derivative or a biologically active portion thereof can be packaged as articles of manufacture containing packaging material, a pharmaceutical composition which is effective for treating the disease or disorder, and a label that indicates that selected antibody or nucleic acid molecule is to be used for treating the disease or disorder.
[0802] The articles of manufacture provided herein contain packaging materials. Packaging materials for use in packaging pharmaceutical products are well known to those of skill in the art. See, for example, U.S. Pat. Nos. 5,323,907, 5,052,558 and 5,033,252, each of which is incorporated herein in its entirety. Examples of pharmaceutical packaging materials include, but are not limited to, blister packs, bottles, tubes, inhalers, pumps, bags, vials, containers, syringes, bottles, and any packaging material suitable for a selected formulation and intended mode of administration and treatment. A wide array of formulations of the compounds and compositions provided herein are contemplated as are a variety of treatments for any EPO-mediated disease or disorder or therapeutic polypeptide-mediated disease or disorder.
[0803] Antibodies and nucleic acid molecules encoding the antibodies thereof also can be provided as kits. Kits can include a pharmaceutical composition described herein and an item for administration. For example, a selected antibody can be supplied with a device for administration, such as a syringe, an inhaler, a dosage cup, a dropper, or an applicator. The kit can, optionally, include instructions for application including dosages, dosing regimens and instructions for modes of administration. Kits also can include a pharmaceutical composition described herein and an item for diagnosis. For example, such kits can include an item for measuring the concentration, amount or activity of the antibody in a subject.
[0804] 9. Formulations/Administration and Uses of Antibodies and Polypeptides
[0805] The antibodies provided herein can be provided as a formulation for administration. While it is possible for the active ingredient to be administered alone, it is preferable to present it as a pharmaceutical formulation. Formulations contain at least one active ingredient, together with one or more acceptable carriers thereof. Each carrier must be both pharmaceutically and physiologically acceptable in the sense of being compatible with the other ingredients and not injurious to the patient. Formulations include those suitable for oral, rectal, nasal, or parenteral (including subcutaneous, intramuscular, intravenous and intradermal) administration. The formulations can conveniently be presented in unit dosage form and can be prepared by methods well known in the art of pharmacy. See, e.g., Gilman, et al. (eds. 1990) Goodman and Gilman's: The Pharmacological Bases of Therapeutics, 8th Ed., Pergamon Press; and Remington's Pharmaceutical Sciences, 17th ed. (1990), Mack Publishing Co., Easton, Pa.; Avis, et al. (eds. 1993) Pharmaceutical Dosage Forms: Parenteral Medications Dekker, NY; Lieberman, et al. (eds. 1990) Pharmaceutical Dosage Forms: Tablets Dekker, NY; and Lieberman, et al. (eds. 1990) Pharmaceutical Dosage Forms: Disperse Systems Dekker, NY.
[0806] The route of antibody administration is in accord with known methods, e.g., injection or infusion by intravenous, intraperitoneal, intracerebral, intramuscular, subcutaneous, intraocular, intraarterial, intrathecal, inhalation or intralesional routes, topical or by sustained release systems as noted below. The antibody is preferably administered continuously by infusion or by bolus injection. One can administer the antibodies in a local or systemic manner.
[0807] The antibodies provided herein can be prepared in a mixture with a pharmaceutically acceptable carrier. Techniques for formulation and administration of the compounds of the instant application can be found in "Remington's Pharmaceutical Sciences," Mack Publishing Co., Easton, Pa., latest edition. This therapeutic composition can be administered intravenously or through the nose or lung, preferably as a liquid or powder aerosol (lyophilized). The composition also can be administered parenterally or subcutaneously as desired. When administered systematically, the therapeutic composition should be sterile, pyrogen-free and in a parenterally acceptable solution having due regard for pH, isotonicity, and stability. These conditions are known to those skilled in the art.
[0808] Therapeutic formulations can be administered in many conventional dosage formulations. Briefly, dosage formulations of the antibodies provided herein are prepared for storage or administration by mixing the compound having the desired degree of purity with physiologically acceptable carriers, excipients, or stabilizers. Such materials are non-toxic to the recipients at the dosages and concentrations employed, and can include buffers such as TRIS HCl, phosphate, citrate, acetate and other organic acid salts; antioxidants such as ascorbic acid; low molecular weight (less than about ten residues) peptides such as polyarginine, proteins, such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidinone; amino acids such as glycine, glutamic acid, aspartic acid, or arginine; monosaccharides, disaccharides, and other carbohydrates including cellulose or its derivatives, glucose, mannose, or dextrins; chelating agents such as EDTA; sugar alcohols such as mannitol or sorbitol; counterions such as sodium and/or nonionic surfactants such as TWEEN, PLURONICS or polyethyleneglycol.
[0809] When used for in vivo administration, the antibody formulation must be sterile and can be formulated according to conventional pharmaceutical practice. This is readily accomplished by filtration through sterile filtration membranes, prior to or following lyophilization and reconstitution. The antibody ordinarily will be stored in lyophilized form or in solution. Other vehicles such as naturally occurring vegetable oil like sesame, peanut, or cottonseed oil or a synthetic fatty vehicle like ethyl oleate or the like can be desired. Buffers, preservatives, antioxidants and the like can be incorporated according to accepted pharmaceutical practice.
[0810] Pharmaceutical compositions suitable for use include compositions wherein one or more rationally designed antibodies are contained in an amount effective to achieve their intended purpose. Determination of a therapeutically effective amount is well within the capability of those skilled in the art, especially in light of the detailed disclosure provided herein. Therapeutically effective dosages can be determined by using in vitro and in vivo methods.
[0811] An effective amount of antibody to be employed therapeutically will depend, for example, upon the therapeutic objectives, the route of administration, and the condition of the patient. In addition, the attending physician takes into consideration various factors known to modify the action of drugs including severity and type of disease, body weight, sex, diet, time and route of administration, other medications and other relevant clinical factors. Accordingly, it will be necessary for the therapist to titer the dosage and modify the route of administration as required to obtain the optimal therapeutic effect. Typically, the clinician will administer antibody until a dosage is reached that achieves the desired effect. The progress of this therapy is easily monitored by conventional assays.
[0812] For any antibody containing a peptide, the therapeutically effective dose can be estimated initially from cell culture assays. For example, a dose can be formulated in animal models to achieve a circulating concentration range that includes the EC50 as determined in cell culture (e.g., the concentration of the test molecule which promotes or inhibits cellular proliferation or differentiation). Such information can be used to more accurately determine useful doses in humans.
[0813] Toxicity and therapeutic efficacy of the antibody molecules described herein can be determined by standard pharmaceutical procedures in cell cultures or experimental animals, e.g., for determining the LD50 (the dose lethal to 50% of the population) and the ED50 (the dose therapeutically effective in 50% of the population). The dose ratio between toxic and therapeutic effects is the therapeutic index and it can be expressed as the ratio between LD50 and ED50. Molecules which exhibit high therapeutic indices are preferred. The data obtained from these cell culture assays and animal studies can be used in formulating a range of dosage for use in human. The dosage of such molecules lies within a range of circulating concentrations that include the ED50 with little or no toxicity. The dosage can vary within this range depending upon the dosage form employed and the route of administration utilized. The exact formulation, route of administration and dosage can be chosen by the individual physician in view of the patient's condition. (See e.g., Fingl et al, 1975, in "The Pharmacological Basis of Therapeutics", Ch. 1, p. 1).
[0814] Dosage amount and interval can be adjusted individually to provide plasma levels of the antibody which are sufficient to promote or inhibit cellular proliferation or differentiation or minimal effective concentration (MEC). The MEC will vary for each antibody, but can be estimated from in vitro data using described assays. Dosages necessary to achieve the MEC will depend on individual characteristics and route of administration. However, HPLC assays or bioassays can be used to determine plasma concentrations.
[0815] Dosage intervals can also be determined using MEC value. Antibody molecules should be administered using a regimen which maintains plasma levels above the MEC for 10-90% of the time, typically between 30-90% and most typically between 50-90%.
[0816] In cases of local administration or selective uptake, the effective local concentration of the antibody can not be related to plasma concentration.
[0817] A typical daily dosage might range from about 1 .mu./kg to up to 1000 mg/kg or more, depending on the factors mentioned above. Typically, the clinician will administer the molecule until a dosage is reached that achieves the desired effect. The progress of this therapy is easily monitored by conventional assays.
[0818] Depending on the type and severity of the disease, from about 0.001 mg/kg to abut 1000 mg/kg, more typically about 0.01 mg to 100 mg/kg, more typically about 0.010 to 20 mg/kg of the antibody, for example an antagonist or agonist antibody, is an initial candidate dosage for administration to the patient, whether, for example, by one or more separate administrations, or by continuous infusion. For repeated administrations over several days or longer, depending on the condition, the treatment is repeated until a desired suppression of disease symptoms occurs or the desired improvement in the patient's condition is achieved. However, other dosage regimes also are contemplated.
H. EXAMPLES
[0819] The following examples are included for illustrative purposes only and are not intended to limit the scope of the invention.
Example 1
Materials and Methods
1. Plasmids A and C
[0820] Plasmid A (SEQ ID NO:1), Plasmid C (SEQ ID NO:3), Plasmid D (SEQ ID NO:2) and Plasmid E (SEQ ID NO:4), set forth in FIGS. 3-6, are used to produce recombinant Fab antibodies. They contain high-copy, ColE1 replication origins that are compatible with E. coli. In addition, the ColE1 replication origins of plasmids A and D are compatible with the replication origins of plasmids C and E. Moreover, they replicate at similar copy numbers when co-cultured in the same E. coli cell. The plasmids contain a STII leader sequence that allows for the Fab to be translocated to the periplasm, to allow for better folding, and an Ara promoter for arabinose inducible gene expression. Plasmid A contains a heavy chain constant region for production of a Fab heavy chain and includes Flag and His tags for protein purification. Plasmid D contains a heavy chain constant region for production of a Fab heavy chain, and includes Flag, LPETG, and His tags for both protein purification and site specific modification using the protein ligase, sortase. Plasmid C contains a kappa light chain constant region for production of a kappa Fab light chain while Plasmid E contains a lambda light chain constant region for production of a lambda Fab light chain. Thus Plasmids A and D encode Fab heavy chain and Plasmids C and E encode Fab light chain. A Fab is produced upon co-transformation of E. coli with a plasmid encoding a heavy chain and a plasmid encoding a light chain, and induction of the promoter to express the heavy or light chain genes.
2. Cell Lines
[0821] E. coli strains BL21 (EMD Biosciences) and LMG194 (ATCC) were used for expression of the Fabs. Ramos B-Lymphocyte cells (ATCC) and Jurkat T-Lymphocyte cells (ATCC) were used for the Apo-ONE homogenous Caspase-3/7 Assay. The BaF3 mouse peripheral blood cell line (Harvey Lodish, Massachusetts Institute of Technology) was used for the EPO Fab library cell based assays.
3. DNA Sequence Compilation Software
[0822] The DNA Sequence Compilation Software is provided herein. It implements an in silico method for recombining heavy chain V, D, and J segment sequences and recombining light chain V and J segments while maintaining functional germline antibody sequences. The software systematically combines DNA encoding the V.sub.H, D.sub.H and J.sub.H and V.sub.L and J.sub.L germline segment sequences, or alternatively a subset of V.sub.H, D.sub.H and J.sub.H and V.sub.L and J.sub.L germline segment sequences, to produce sequences of recombined heavy chain and light chain variable regions from which nucleic acid molecules are produced for production of a Fab library. The germline segment sequences are inputted from a database. Example 2 provides a description of the software. Example 3 describes how to use the software, and Example 4 describes implementation of the software to compile recombined germline sequences and output the sequences for DNA synthesis.
4. Piccolo.TM. Automated System
[0823] Piccolo.TM. system is a fully automated system for the rapid optimization of recombinant protein production, sold by The Automation Partnership (TAP) (see e.g., Wollerton et al. (2006) JALA, 11:291-303, which describes the system). The system combines cell culture with automated harvesting, lysis, and purification units. Example 9 provides details about the specifics of the Piccolo.TM. system operation.
Example 2
DNA Sequence Compilation Software Design
[0824] This example describes the DNA Sequence Compilation Software.
A. General Description:
[0825] Sequences for V.sub.H, D.sub.H and J.sub.H and V.sub.L and J.sub.L germline segment sequences are inputted from a database file, i.e. SequenceDatabase.txt file described in detail below. The software 1) generates every possible recombination of V.sub.H, D.sub.H and J.sub.H and V.sub.L and J.sub.L germline segments to generate sequences encoding a variable heavy (VH) chain and sequences encoding a variable light (VL) chain; 2) translates and frame-checks the sequences; 3) stores the sequences in a SequenceHistory.txt file to keep track of sequences in nucleotide format; and 4) outputs sequences as files that represent 96-well plates for ordering for DNA synthesis. Output is generated in the form of a 96-well plate file that lists the distinct heavy chain or light chain sequences as an array designating each locus in the array corresponding to a well in a 96-well plate. DNA molecules can be synthesized in a 96-well plate based on this output. In addition, the software can rank the sequences and generate diversity scores. The software automatically generates all permutions of recombined nucleic acid sequences encoding VH chains and VL chains (kappa and lambda). If desired, the output file can be manually restricted by the user by selecting germline segments to be recombined by the software, or by choosing sequences that are sufficiently diverse from one another using the ranking function.
B. Software Description:
[0826] The DNA Sequence Compilation Software was written in the C# programming language using the Visual Studio 2005 Professional Edition development tool. The DNA Sequence compilation Software require the Microsoft .NET framework 2.0 to run under the Windows XP and Vista 32-bit Edition Operating System. The software is composed of ten modules (see FIG. 7.) These modules include (a) Blast to Diversity Converter; (b) DNA Sequence Compilation GUI; (c) DNA Sequence Compilation Controls; (d) DNA Sequence Compilation Rules; and (e) DNA Sequence Compilation Rules Test, which are provided herein. The remaining five modules are available from public domain software. They include (f) NUnit; (g) formatdb; (h) BLAST; (i) BLASTClust; and (j) XP Explorer Bar.
[0827] The modules contain classes. A class is the formal definition of an object. The class acts as the template from which an instance of an object is created at run time. The class defines the properties of the object and the methods used to control the object's behavior. A singleton class is a class that is globally available to be accessed by all other classes. In this case the singleton class is the sequence compiler module, which has every rule for sequence compilation. A static class contains data and functions that can be accessed without creating an instance of the class. Static class members can be used to separate data and behavior that is independent of any object identity; the data and functions do not change regardless of what happens to the object.
[0828] Additionally, the software is grouped into 4 layers: 1) the graphical user interface (GUI); 2) GUI controls; 3) Compilation rules; and 4) NCBI tools. The 4 layers are: [0829] GUI--The GUI is the presentation of information to and interaction with the computer operator. The GUI is made up of the Blast to Diversity Converter, the DNA Sequence Compilation GUI and NUnit. [0830] GUI controls--Custom visible Windows controls that are re-used by the GUI. GUI controls is made up of the XP Explorer Bar and the DNA Sequence Compilation Controls. [0831] Compilation Rules--Implementation of the rules for DNA sequence compilation. The compilation rules are made up of DNA Sequence Compilation Rules and DNA Sequence Compilation Rules Test. [0832] NCBI tools--A set of tools available from NCBI that allows the user to run searches on a local computer.
1. Modules
[0833] a. DNA Sequence Compilation GUI
[0834] DNA Sequence Compilation GUI is the main application. It interacts with the user and enables all the functionality to be available to the user. As a whole, the DNA Sequence Compilation GUI permits the user to selectively restrict germline segments to compile a sequence; automatically compile all sequences; automatically rank all the sequences in terms of sequence diversity, clusters or similarity; and select compiled DNA sequences to be placed in a 96-well plate.
[0835] The DNA Sequence Compilation GUI contains the following classes: [0836] Program--a static class automatically generated that contains the execution entry point to the application; [0837] SplashScreen--a user control that displays a splash screen when the application starts and exits; [0838] MainForm--a form that encapsulates all the functionality that is available to the user. The MainForm also displays the user controls that are implemented in the DNA Sequence Compilation Controls module and initializes the SequenceCompiler singleton defined in the DNA Sequence Compilation Rules module; [0839] Resources--a resource file containing the image for the splash screen; [0840] Settings--an automatically generated class that contains the user settings for the 5' restriction site and 3' restriction site to be placed with the sequence for the IDT order; and [0841] AboutBox--a form that displays the application information to the user. It interacts with the user and enables all the functionality to be available to the user.
[0842] b. DNA Sequence Compilation Controls
[0843] The DNA Sequence Compilation Controls module contains custom window controls to be used by the DNA Sequence Compilation GUI. It contains the following classes: [0844] FabrusDataGridView--a derived class of .NET framework DataGridView that displays the row number in the row header. A .NET framework is part of windows operating systems that contains a large library of pre-coded solutions to common programming problems and manages the execution of programs written specifically for the framework. The DataGridView control provides a customizable table for displaying data. [0845] WellPlateControl--a custom control that displays 96-well plate information; [0846] AutoHeavySequenceControl--a custom control that displays all of the automatically compiled DNA sequences in a data grid view for user selection into the 96-well plate, while also displaying rank, diversity score, cluster, V(D)J sequence identifier and DNA sequence protein sequence; [0847] AutoLightSequenceControl--a custom control that displays all the automatically compiled DNA sequences in a data grid view for user selection into the 96-well plate, while also displaying rank, diversity score, cluster, V(D)J sequence identifier and DNA sequence protein sequence; [0848] ManualHeavySequenceControl--a custom control that allows the user to manually select a V.sub.H, D.sub.H and J.sub.H sequence with which to compile into a DNA heavy chain sequence; [0849] Manual LightSequenceControl--a custom control that allows the user to manually select a V.sub.L and J.sub.L sequence with which to compile into a DNA light chain sequence; [0850] LightSequenceBlastForm--a custom control that displays in grid format the BLAST results of a light sequence against all other light chains; and [0851] HeavySequenceBlastForm--a custom control that displays in grid format the BLAST results of a heavy chain sequence against all the other heavy chains.
[0852] c. DNA Sequence Compilation Rules Module
[0853] The DNA Sequence Compilation Rules module contains all of the business logic for the compilation of the DNA sequences. This module includes all the rules for compiling functional recombined VDJ and VJ antibody sequences, including compilation, joint generation, stop codon removal, and sequence ranking.
[0854] There are six main classes within the DNA Sequence Compilation Module: [0855] SequenceCompiler is a singleton class that provides all the DNA sequence compilation functionality. This class provides the ability to automatically or manually generate combined DNA sequences from the V(D)J sequences and all its associated information; [0856] BlastTable provides all the information regarding diversity scores, cluster number and performs the BLAST search for a specified sequence; [0857] SequenceHistorian is a class that keeps track of the DNA sequences in nucleotide format that have been ordered from the DNA synthesis company (e.g., IDT, Genscript, Invitrogen, and the like). Persistence is achieved by using a text file named SequenceHistory.txt, which keeps track of all of the sequences in nucleotide format ordered from the DNA Synthesis Company and [0858] SequenceContainer is a class that holds all the V(D)J sequence information and restriction sites as specified in the text file SequencesDatabase.txt. [0859] DnaSequence is the parent class that models the heavy and light chain sequences through the use of HeavyChainDnaSequence and LightChainDnaSequence classes, respectively. [0860] WellPlate_8.times.12 is a class that models a 96-well plate. It holds references to the DNA sequence instances and allows them to be saved into an order file.
[0861] Additionally, there are three helper classes within the DNA Sequence Compilation Rules: [0862] CodonUsageTable is a class that encapsulates the codon usage table for E. coli K12; [0863] Translator is a static class that performs the translation of the sixty four codons to their respective amino acid equivalents based on the genetic code; and [0864] ProtoParam is a static class that computes the GRAVY value.
[0865] The following is a discussion of each of the six main classes (i-vi) and the three helper classes (vii-ix) of the DNA Sequence Compilation Rules Module.
[0866] i. SequenceCompiler
[0867] SequenceCompiler is a singleton class that provides all the DNA sequence compilation functionality. This class provides the ability to automatically or manually generate combined DNA sequences from the V(D)J sequences and all its associated information. Sequence Compiler includes the following functionalities:
[0868] 1) Algorithm for Automatic DNA Sequence Compilation
[0869] SequenceCompiler uses the algorithm shown in FIG. 8 to generate compiled sequences by computing all the possible combinations of the available V(D)J sequences for the heavy and light chains. The algorithm equates to simply computing all the possible combinations of the available V(D)J sequences for the heavy and light sequences, respectively.
[0870] For example, if there are three V.sub.H sequences, three D.sub.H sequences and three J.sub.H sequences, the algorithm computes 3*3*3 combinations, thereby producing all 27 possible combined sequences. Similarly, if there are four V.sub..kappa. sequences and four J.sub..kappa. sequences, the algorithm computes 4*4 combinations, thereby producing 16 possible combined sequences. In addition, the algorithm computes lambda light chains and kappa light chains separately.
[0871] 2) Sequence Compilation
[0872] Actual sequence compilation starts with the identification of a V.sub.H, D.sub.H, and J.sub.H sequence and a V.sub.L and J.sub.L sequence for a heavy and light chain respectively from the SequenceDatabase.txt file. Individual segments are identified as described in Example 4. Once the individual segments are identified, V(D)J joints are created, and finally all nucleotide pieces (the segments and the joints) are joined together thus compiling the sequence (see FIG. 9). The rules for joint generation, removal of STOP codons and restriction sites are set forth below under Section Headings "V(D)J Joint Generation," "Stop Codon Removal," and "Codon Usage Table," respectively.
[0873] 3) V(D)J Joint Generation
[0874] The joints in the compiled sequences between the different V(D)J sequences are selected so that the resulting sequences are in frame. SequenceCompiler uses the following three rules for determining reading frames. [0875] 1. The V Sequence is always reading frame 1. [0876] 2. The D Sequence has a frame that yields the most negative GRAVY (Grand Average of Hydropathy) value. [0877] 3. The J Sequence has a reading frame as specified in the SequenceDatabase.txt file (see Example 4.1 entitled "Creating the V(D)J Sequence Database File," which describes that the J sequences are set forth in three letter codons to designate the reading frame; also see Table 13).
[0878] Based on the rules listed above, in the joints between sequences, a codon will not automatically be generated by simply recombining the bases from either of the sequences. SequenceCompiler uses the rules listed in Table 19 to generate a new codon. For example, when creating a light chain V-J joint where the V sequence is in reading frame 1 and the J sequence is in reading frame 1, a "G" nucleotide will be inserted between the V-J joint, thereby maintaining the reading frame of the entire segment.
TABLE-US-00030 TABLE 19 V(D)J joint generation. V-D Joints (Heavy Chains) V D Frame Rule 0 0 0 NO MANIPULATION 0 1 1 Delete 1 nucleotide from 5' end of D 0 2 2 Add a "G" between V-D 1 0 1 Delete 1 nucleotide from 3' end of V 1 1 2 Add a "G" between V-D 1 2 3 NO MANIPULATION 2 0 2 Add a "G" between V-D 2 1 3 NO MANIPULATION 2 2 4 Delete 1 nucleotide from 3' end of V D-J Joints (Heavy Chains) D J Frame Rule 0 0 0 NO MANIPULATION 0 1 1 Delete 1 nucleotide from 5' end of J 0 2 2 Add a "G" between D-J 1 0 1 Delete 1 nucleotide from 3' end of D 1 1 2 Add a "G" between D-J 1 2 3 NO MANIPULATION 2 0 2 Add a "G" between D-J 2 1 3 NO MANIPULATION 2 2 4 Delete 1 nucleotide from 3' end of D V-J Joints (Light Chains) V J Frame Rule 0 0 0 NO MANIPULATION 0 1 1 Delete 1 nucleotide from 5' end of J 0 2 2 Add a "G" between V-J 1 0 1 Delete 1 nucleotide from 3' end of V 1 1 2 Add a "G" between V-J 1 2 3 NO MANIPULATION 2 0 2 Add a "G" between V-J 2 1 3 NO MANIPULATION 2 2 4 Delete 1 nucleotide from 3' end of V
[0879] 4) Stop Codon Removal
[0880] There are situations where V(D)J joint generation will generate a STOP codon (TAA, TAG or TGA). SequenceCompiler removes detrimental STOP codons according to the following rules, whereby nucleotides encoding a Stop Codon are replaced with the designated new codon:
TABLE-US-00031 STOP Codon New Codon TAA TAT TAG TAT TGA TCA
[0881] Once SequenceCompiler has completed the successive steps of V(D)J joint generation and STOP codon removal, the protein sequence is complete and functional.
[0882] 5) Algorithm for Ranking Sequences
[0883] Once sequences are compiled, each sequence is ranked and presented to the user in order of highest diversity to lowest diversity (See Table 20 and FIGS. 17 and 19). This presentation of ranked sequences allows the user several options when selecting sequences to be placed in the 96-well plate order file. The user can select the most diverse sequences, or sequences that are most similar to each other.
[0884] Compiled sequences are ranked by the software by using a combination of the diversity score and cluster number information for the sequences. Diversity scores are generated by the blast to diversity converter module, as described in Section B.1.e, and cluster information is generated by BLASTClust, as described in Section B.1.i. The sequence ranking algorithm flowchart is shown in FIG. 10. The algorithm picks from each cluster the sequence with the lowest diversity score, orders those sequences in terms of diversity from the lowest diversity number to the highest diversity number (a low diversity score indicates lack of similarity, and therefore highest diversity), ranks the sequences in that order, eliminates those sequences from the respective clusters, and then iteratively applies the previous procedure until all the sequences are ranked.
[0885] An example of the ranking of Sequences using this algorithm is set forth in Table 20. The sequences in the table are ordered from highest diversity to lowest diversity. The sequence with the lowest diversity score indicates the highest diversity and receives the rank of 1. The second sequence belongs to the same cluster as the first sequences, and receives the rank of 64. This data also indicates to the user that there are a total of 63 clusters defined, since sequence 001 and sequence 002 are both in cluster 39 and they have consecutive diversity scores.
TABLE-US-00032 TABLE 20 Ranking of Sequences Diversity V J Amino Acid SEQ Selected Rank Cluster Score Sequence Sequence Sequence ID NO 001 1 39 73.202 V3-4 IGLJ6*01 QTVVTQEPSFSVSP . . . 1905 002 64 39 73.305 V3-4 IGLJ7*01 QTVVTQEPSFSVSP . . . 1906 003 2 21 73.560 V4-2 IGLJ6*01 QAVLTQPSSLSASP . . . 1907 004 65 32 73.603 V4-2 IGLJ7*01 QAVLTQPSSLSASP . . . 1908 005 3 32 73.876 V1-22 IGLJ6*01 NFMLTQPHSVSES . . . 1909 006 66 24 74.017 V1-22 IGLJ7*01 NFMLTQPHSVSES . . . 1910 007 4 24 74.040 V5-1 IGLJ7*01 LPVLTQPPSASALL . . . 1911 008 67 39 74.056 V5-1 IGLJ6*01 LPVLTQPPSASALL . . . 1912 009 127 39 74.081 V3-4 IGLJ1*01 QTVVTQEPSFSVSP . . . 1881 010 190 21 74.399 V3-4 IGLJ4*01 QTVVTQEPSFSVSP . . . 1913 011 128 39 74.458 V4-2 IGLJ1*01 QAVLTQPSSLSASP . . . 1914 012 253 21 74.571 V3-4 IGLJ5*01 QTVVTQEPSFSVSP . . . 1915 013 191 21 74.577 V4-2 IGLJ4*01 QAVLTQPSSLSASP . . . 1883
[0886] ii. BlastTable
[0887] BlastTable provides all the information regarding diversity scores, cluster number and performs the BLAST search for a specified sequence;
[0888] iii. SequenceHistorian
[0889] SequenceHistorian is a class that keeps track of the DNA sequences in nucleotide format that have been ordered for DNA synthesis. The persistence is achieved by using a text file named SequenceHistory.txt.
[0890] iv. SequenceContainer
[0891] SequenceContainer is a class that reads V(D)J sequences and restriction sites from a text file and stores them in memory for later retrieval for either manual (i.e. restricted selection) or automatic compilation.
[0892] v. DnaSequence
[0893] DnaSequence is the parent class that models the heavy and light chain sequences through the use of HeavyChainDnaSequence and LightChainDnaSequence classes, respectively. The only difference between the HeavyChainDnaSequence and LightChainDnaSequence classes is in the way the sequence is compiled which is implemented in the constructor of each class.
[0894] Each class contains Fields, Properties and Methods. In addition, the parent DnaSequence class contains events and Nested Types. Thus, DnaSequence implements all the functionality that is common to heavy chain and light chain sequences. This includes the fields such as AminoAcids, Cluster, DiversityScore, _SEQ_ID, Name, Notes, Nucleotides, Rank, and RestrictionSites and methods such as removing restriction sites, removing stop codons and silencing restriction sites. HeavyChainDnaSequence class contains fields for the V.sub.H, J.sub.H and D.sub.H sequences, the V.sub.H-J.sub.H and J.sub.H-D.sub.H joints and methods to generate the V.sub.H-J.sub.H and J.sub.H-D.sub.H joints. LightChainDnaSequence class contains fields for the V.sub.L and J.sub.L sequences, the V.sub.L-J.sub.L joint and methods to generate the V.sub.L-J.sub.L joint.
[0895] vi. WellPlate_8.times.12
[0896] WellPlate_8.times.12 is a class that models a 96-well plate. It holds references to the DNA sequence instances and allows them to be saved into an order file.
[0897] vii CodonUsageTable
[0898] CodonUsageTable is a class that encapsulates the codon usage table for E. coli K12 (see, Table 14). The CodonUsageTable is used by the software to modify reading frame problems as well as restriction site silencing. The software is programmed to recognize restriction sites that will be used to insert the DNA into the desired plasmid. When these restriction sites are observed in the compiled DNA sequence, the software modifies the nucleotide sequence to modify the undesired restriction site while maintaining the appropriate amino acid sequence. The above mentioned codon redundancy is used to modify codons. A codon change is used to provide a beneficial increase in the frequency of usage of the possible newly used codon. In addition, modifying the last base of a codon (the degenerate position) is preferred over modifying the first or second base of the codon.
[0899] viii. Translator
[0900] Translator is a static class that performs the translation of the sixty four codons to their respective amino acid equivalents based on the genetic code.
TABLE-US-00033 TABLE 20 DNA Codon table. Amino Acid SLC DNA codons Isoleucine I ATT, ATC, ATA Leucine L CTT, CTC, CTA, CTG, TTA, TTG Valine V GTT, GTC, GTA, GTG Phenylalanine F TTT, TTC Methionine M ATG Cysteine C TGT, TGC Alanine A GCT, GCC, GCA, GCG Glycine G GGT, GGC, GGA, GGG Proline P CCT, CCC, CCA, CCG Threonine T ACT, ACC, ACA, ACG Serine S TCT, TCC, TCA, TCG, AGT, AGC Tyrosine Y TAT, TAC Tryptophan W TGG Glutamine Q CAA, CAG Asparagine N AAT, AAC Histidine H CAT, CAC Glutamic acid E GAA, GAG Aspartic acid D GAT, GAC Lysine K AAA, AAG Arginine R CGT, CGC, CGA, CGG, AGA, AGG Stop codons Stop TAA, TAG, TGA In this table, the twenty amino acids found in proteins are listed, along with the single-letter code used to represent these amino acids in protein data bases. The DNA codons representing each amino acid are also listed. All 64 possible 3-letter combinations of the DNA coding units T, C, A and G are used either to encode one of these amino acids or as one of the three stop codons that signals the end of a sequence. While DNA can be decoded unambiguously, it is not possible to predict a DNA sequence from its protein sequence. Because most amino acids have multiple codons, a number of possible DNA sequences might represent the same protein sequence.
[0901] ix. ProtoParam
[0902] ProtoParam is a static class that computes the GRAVY value. The GRAVY (Grand Average of Hydropathy) value for a peptide or protein is calculated as the sum of hydropathy values of all the amino acids divided by the number of residues in the sequence (Table 21). In essence, the GRAVY score is the relative value for the hydrophobic residues of the protein. This information is used by the V(D)J joint generation software when computing the D sequence reading frame. This reading frame is determined by the most negative GRAVY value.
TABLE-US-00034 TABLE 21 Hydropathy Values Amino Acid Hydropathy Value Ala 1.800 Arg -4.500 Asn -3.500 Asp -3.500 Cys 2.500 Gln -3.500 Glu -3.500 Gly -0.400 His -3.200 Ile 4.500 Leu 3.800 Lys -3.900 Met 1.900 Phe 2.800 Pro -1.600 Ser -0.800 Thr -0.700 Trp -0.900 Tyr -1.300 Val 4.200
[0903] d. DNA Sequence Compilation Rules Test Module
[0904] The DNA Sequence Compilation Rules Test module is a collection of NUnit automated tests that perform unit tests for each class. These tests help to ensure the individual modules and classes listed are running properly. These nine classes include:
[0905] BlastTableTest;
[0906] CodonUsageTableTest;
[0907] HeavyChainDnaSequenceTester;
[0908] LightChainDnaSequenceTester;
[0909] ProtoParamTest;
[0910] SequenceCompilerTest,
[0911] SequenceContainerTest;
[0912] TranslatorTest; and
[0913] TranslatorWrapper.
[0914] e. Blast to Diversity Converter Module
[0915] The Blast to Diversity Converter is an executable that takes a BLAST output file and outputs a diversity score. It contains five classes: [0916] Program--a static class that is automatically generated and contains the execution entry point to the application; [0917] Main Form--a button for the operator to click on to select a file to be converted; [0918] Resources--an empty resource file that is created by default; [0919] Settings--an automatically generated class that contains the user settings; and [0920] BlastConverter--a class that contains the implementation of the conversion of the BLAST output file to a diversity score file.
[0921] i. BLAST Converter Algorithm
[0922] The BLAST converter algorithm takes a BLAST output file and applies the following equation to compute a sequence's diversity score.
DiversityScore ( seq i ) = 1 N k = 1 , k .noteq. i N BLAST ( i , k ) 2 ##EQU00001##
[0923] The algorithm computes a standard deviation for the BLAST scores by taking all the individual BLAST numbers, squaring them, adding up all the squares, normalizing by the number of samples (N), and then taking the square root.
[0924] For example, a sequence's diversity score was computed by applying the algorithm to the following BLAST output file (-m 9 BLAST switch, i.e. Alignment view options, tabular with comment lines; see Section B.1.h.):
TABLE-US-00035 # BLASTP 2.2.17 # Query: gnl|Fabrus|V4-2_IGLJ2*01 # Database: AllLightChainSequences.txt # Fields: Query id, Subject id, % identity, alignment length, mismatches, gap openings, q. start, q. end, s. start, s. end, e-value, bit score gnl|Fabrus|V4-2_IGLJ2*01 gnl|Fabrus|V4-2_IGLJ2*01 100.00 116 0 0 1 116 1 116 1e-050 186 gnl|Fabrus|V4-2_IGLJ2*01 gnl|Fabrus|V4-2_IGLJ7*01 98.28 116 2 0 1 116 1 116 2e-04 182 . . . gnl|Fabrus|V4-2_IGLJ2*01 gnl|Fabrus|A5-IGKJ1*01 38.82 85 46 3 31 115 34 112 2e-007 43.1 # BLASTP 2.2.17 # Query: gnl|Fabrus|V4-2_IGLJ1*01 # Database: AllLightChainSequences.txt # Fields: Query id, Subject id, % identity, alignment length, mismatches, gap openings, q. start, q. end, s. start, s. end, e-value, bit score gnl|Fabrus|V4-2_IGLJ1*01 gnl|Fabrus|V4-2_IGLJ1*01 100.00 116 0 0 1 116 1 116 3e-050 185 gnl|Fabrus|V4-2_IGLJ2*01 gnl|Fabrus|V4-2_IGLJ6*01 98.2 116 2 0 1 116 1 116 4e-049 181 . . . indicates data missing or illegible when filed
[0925] The following output file is generated, which provide diversity scores for the respective variable heavy chain sequences. The lowest diversity score indicates the least similarity between the sequence and the next most related sequence, and therefore has the highest diversity.
TABLE-US-00036 gnl | Fabrus | VH1-69_IGHD7-27*01_IGHJ2*01 119.675348536301 gnl | Fabrus | VH3-15_IGHD5-24*01_IGHJ2*01 136.102291820977 gnl | Fabrus | VH2-26_IGHD1-7*01_IGHJ4*01 111.073223319952 gnl | Fabrus | VH3-13_IGHD1-14*01_IGHJ1*01 136.760646289454 gnl | Fabrus | VH3-30_IGHD6-13*01_IGHJ6*01 139.423052600581
[0926] f. NUnit
[0927] NUnit is a unit-testing framework for all .Net languages. NUnit testing tools provide a means to test an object model, and is useful in ensuring that the application and its logic work correctly. Initially ported from JUnit, the current production release, version 2.4.2, is the fifth major release of this xUnit based unit testing tool for Microsoft .NET. It is written entirely in C# and has been completely redesigned to take advantage of many .NET language features, for example custom attributes and other reflection related capabilities. NUnit brings xUnit to all .NET languages. NUnit is available free to the public from the organization NUnit.
[0928] g. Formatdb
[0929] Formatdb must be used in order to format protein or nucleotide source databases before these databases can be searched by blastall, blastpgp or MegaBLAST. The source database can be in either FASTA or ASN.1 format. Once a source database has been formatted by formatdb it is not needed by BLAST.
[0930] The following Command Line Options are used when generating output files with formatdb: [0931] i Input File for formatting [0932] p Type of File (T--protein; F--nucleotide; Default=T) [0933] o Parse options [0934] T--True: Parse SegID and create indexes [0935] F--False: Do not parse SegID. Do not create indexes.
[0936] The version win32-ia32 (2.2.17) is currently used. Formatdb can be downloaded from the National Center for Biotechnology Information.
[0937] h. The Basic Local Alignment Search Tool (BLAST)
[0938] The Basic Local Alignment Search Tool (BLAST) finds regions of local similarity between sequences. The program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches. BLAST can be used to infer functional and evolutionary relationships between sequences as well as help identify members of gene families.
[0939] The following Command Line Options are used when generating output files with blastall: [0940] p Program Name ("blastp", "blastn", "blastx", "tblastn", or "tblastx") [0941] d Database [0942] l Query File [0943] o BLAST report Output File [0944] m Alignment View Options (0=default; 9=tabular with comment lines) [0945] b Number of Database Sequence to show alignments for (B) [Integer]
[0946] The version win32-ia32 (2.2.17) is currently used. BLAST can be downloaded from the National Center for Biotechnology Information.
[0947] i. BLASTClust
[0948] BLASTClust is a program within the stand alone BLAST package used to cluster either protein or nucleotide sequences. The program begins with pairwise matches and places a sequence in a cluster if the sequence matches at least one sequence already in the cluster. In the case of proteins, the blastp algorithm is used to compute the pairwise matches; in the case of nucleotide sequences, the Megablast algorithm is used.
[0949] In the simplest case, BLASTClust takes as input a file containing concatenated FASTA format sequences, each with a unique identifier at the start of the definition line. BLASTClust formats the input sequence to produce a temporary BLAST database, performs the clustering, and removes the database at completion. Hence, there is no need to run formatdb in advance to use BLASTClust. The output of BLASTClust is a file, with one cluster to a line, of sequence identifiers separated by spaces. The clusters are sorted from the largest cluster to the smallest.
[0950] The following Command Line Options are used when generating output files with blastclust: [0951] d Sequence Database Name [0952] o Output File to save cluster list [0953] S Similarity Threshold [0954] if <3 then the threshold is set as a BLAST score density (0.0 to 0.3; default=1.75) [0955] if >3 then the threshold is set as a percent of identical residues (3 to 100)
[0956] The version win32-ia32 (2.2.17) is currently used. BLASTClust can be downloaded from the National Center for Biotechnology Information
[0957] j. XPExplorerBar
[0958] The XPExplorerBar is a fully customizable Windows XP style Explorer Bar that supports Windows XP themes and animated expand/collapse with transparency. The XP Explorer Bar is available as free public domain software.
Example 3
Sequence Database
[0959] The DNA Sequence compilation software requires a file containing sequence information used by the program to perform sequence compilation in accordance with sequence compilation rules. This is called the SequenceDatabase.txt file. All recombined sequences are generated from the sequence data in the file. Hence, the file contains all sequences that are used in performing the sequence compilation by the software. For example, the file contains nucleotide sequences for all desired germline segments and also contains nucleotide recognition sequences for restriction enzyme. The sequences in the database include any known sequence, and can be updated manually at any time. V, D, and J antibody germline sequences of human origin were obtained from the NCBI IgBLAST database (available from the National Center for Biotechnology Information) and the International ImMunoGeneTics Information system (IMGT) and were entered into a sequence database file. The database file served as an input file for the Sequence Compilation Software.
[0960] The format of the SequenceDatabast.txt file is illustrated in FIG. 11. The SequencesDatabase.txt file is updated by simply manually adding new sequences according to the following rules: [0961] All sequences must be specified using the FASTA format. [0962] Three letter codons are specified for J sequences. [0963] All sequences must have a "blank line" between them. [0964] // is used a the beginning of any line containing extraneous comments. [0965] Section titles are indicated by the following designations: [0966] [VH]--List of V.sub.H sequence to follow. [0967] [DH]--List of D.sub.H sequence to follow. [0968] [JH]--List of J.sub.H sequence to follow. [0969] [VK]--List of V.sub..kappa. sequence to follow. [0970] [JK]--List of J.sub..kappa. sequence to follow. [0971] [VL]--List of V.sub.L sequence to follow. [0972] [JL]--List of J.sub.L sequence to follow. [0973] [Restriction Sites]--List of restriction sites to follow.
Example 4
Generation of Heavy and Light Chain DNA Files and Ranking Analysis
[0974] This example describes methods for generating recombined heavy and light chain sequence files that will be used by the DNA Sequence Compilation Software. The files were created immediately upon initiating the software. The files are created from the SequenceDatabase.txt file and are analyzed using public domain software, such as NCBI's formatdb, BLAST and BLASTclust, to compute diversity and clustering information for the sequences included in the database file. The ranking function of the software is performed automatically, but can optionally be used by the user to rank the sequences for selection and ordering for DNA synthesis.
[0975] To generate sequence files, a database file that contains DNA sequences for all desired V, (D), and J variable regions sequences was generated as described in Example 3. The DNA Sequence Compilation Software mechanically recombined all heavy chain variable germline segments and light chain variable germline segment sequences in the database into nucleic acid molecules encoding functional amino acid sequences. The software translated the nucleic acid sequence into an amino acid sequence to create sequence files. The recombined amino acid sequences were compared to determine the similarity of each compiled sequence to every other compiled sequence, which was subsequently used to generate a diversity score. Finally, the compiled sequences were clustered by sequence similarity. These various steps of the software were performed immediately upon initiating the software and are described below.
[0976] 1. Compiling Heavy Chains and Light Chains
[0977] Once the database file was updated and contains all the desired V(D)J sequences, the database file was used as an input file by DNA Sequence Compilation software to mechanically compile full length heavy chain or light chain sequences in accordance with the DNA sequence compilation rules as described in Example 2, Section B.1.c. The software created files containing the full-length amino acid sequences encoded by compiled variable heavy and light chain germline segments. These are contained in the following two files: [0978] TempLightChainSequences.txt--This file contains all the lambda and kappa light chain sequence combinations in the amino-acid format. [0979] TempHeavyChainSequences.txt--This file contains all the heavy chain sequence combinations in the amino-acid format.
[0980] The compiled light and heavy chain amino acid sequences are each identified with a unique sequence ID (gnl|Fabrus|V_(D)_J), examples of which are illustrated below.
TABLE-US-00037 >gnl|Fabrus|V4-2_IGLJ2*01 (SEQ ID NO: 2027) QAVLTQPSSLSASPGASASLTCTLRSGINVGTYRIYWYQQKPGSPPQYLL RYKSDSDKQQGSGVPSRFSGSKDASANAGILLISGLQSEDEADYYCMIWN SSASWFGGGTKLTVL >gnl|Fabrus|VH1-2_IGHD2-21*01_IGHJ1*01 (SEQ ID NO: 2028) QVQLVQSGAEVKKPGASVKVSCKASGYTFTGYYMHWVRQAPGQGLEWMGW INPNSGGTNYAQKFQGRVTMTRDTSISTAYMELSRLRSDDTAVYYCAREA YCGGDCYSAEYFQHWGQGTLVTVSS
[0981] 2. Ranking Recombined Variable Heavy and Light Chains
[0982] Recombined variable heavy and light chain sequences were compared by the software for sequence similarity between and among all recombined sequences. This is performed using the NCBI utility BLAST. The Blast bit score is used to generate a diversity score. Recombined variable heavy and light chain sequences also were analyzed for sequence similarity using cluster analysis (e.g. BLASTclust).
[0983] As described in Example 2, under the heading "Alogrithm for ranking Sequences," compiled sequences were ranked by the software by using a combination of the diversity score and cluster number information for the sequences. The ranking occurs automatically upon initiation of the program, and can be viewed by the user using the auto compilation function as described in Example 5 below. For example, FIGS. 17 and 19 exemplify screen shots following auto compilation, and depict columns indicating the cluster, diversity score and rank for exemplary heavy and light chain sequences.
[0984] The following sections describe the various steps for determining the diversity score and the cluster in order to rank the sequences using the Alogrithm described in Example 2.
[0985] a. Calculating Compiled Sequence Similarity and Diversity Scores
[0986] The compiled sequences were compared in order to determine the similarity of each compiled sequence to every other compiled sequence. This information was used to generate a diversity score. To start, the TempLightChainSequences.txt and TempHeavyChainSequences.txt files were copied to a folder (as created by the user) and a DOS command prompt window was opened. The NCBI DNA utility formatdb was used to prepare the files for use by BLAST by first using the following command prompts for the respective light chain and heavy chain files:
[0987] Run formatdb -i TempLightChainSequences.txt -p T -o F
[0988] Run formatdb -i TempHeavyChainSequences.txt -p T -o F
[0989] Formatdb must be installed in the folder for which the DOS command prompt window is opened or the utility will not be found. Alternatively, a path can be added to direct the DOS command to utilize formatdb from the folder it was previously installed in. For example, if all the NCBI utilities are downloaded into a folder entitled "NCBIUtilities" then the command prompts are be as follows:
[0990] Run NCBIUtilities\bin\formatdb -i TempLightChainSequences.txt -p T -o F
[0991] Run NCBIUtilities\bin\formatdb -i TempHeavyChainSequences.txt -p T -o F
[0992] BLAST command prompts were then used to generate output files LightChainBlastResults.txt and HeavyChainBlastResults.txt. The command prompts are as follows: [0993] Run blastall -p blastp -d TempLightChainSequences.txt -I TempLightChainSequences.txt -o LightChainBlastResults.txt -m 9 -b 500 (The -b value must be equal to or greater than the number of sequences to BLAST.) [0994] Run blastall -p blastp -d TempHeavyChainSequences.txt -I TempHeavyChainSequences.txt -o HeavyChainBlastResults.txt -m 9 -b 8000 (The -b value must be equal to or greater than the number of sequences to BLAST.)
[0995] Once the BLAST output files were generated, the BLAST bit score is used to calculate a diversity score for each sequence using the algorithm shown in Example 2, under the sub-section heading "Blast Converter Algorithm" using the following commands:
[0996] (a) Run BlastToDiversityConverter.exe (This file runs automatically each time the software is started.)
[0997] (b) Open the file LightChainBlastResults.txt
[0998] (c) Save this file as AllLightChainSequencesDiversityScores.txt
[0999] (d) Open the file HeavyChainBlastResults.txt
[1000] (e) Save this file as AllHeavyChainSequencesDiversityScores.txt
[1001] b. Clustering Compiled Sequences by Sequence Similarity
[1002] Finally, the sequences are clustered based on sequence similarity by generating pairwise matches using the NCBI utility BLASTClust. BLASTClust is used to generate two output files, AllLightChainsBLastClust.txt and AllHeavyChainsBlastClust.txt. The current example dictates that two sequences must be either 95% or 96% identical to register a pairwise match. The command prompts are as follows: [1003] Run blastclust -d TempLightChainSequences.txt -o AllLightChainsBlastClust.txt -S 95 [1004] Run blastclust -d TempHeavyChainSequences.txt -o AllHeavyChainsBlastClust.txt -S 96
[1005] The BlastClust setting (-S) can be modified to in order to better suit any individual requirements that are necessary.
[1006] Once all the files are generated (as indicated above), the following files are manually copied into the installation folder of the DNA Sequence Compilation Tool.
[1007] (a) AllHeavyChainsBlastClust.txt
[1008] (b) AllLightChainsBlastClust.txt
[1009] (c) AllLightChainSequencesDiversityScores.txt
[1010] (d) AllHeavyChainSequencesDiversityScores.txt
Example 5
Installation and Graphical User Interface of the DNA Sequence Compilation Tool Software
[1011] The DNA Sequence Compilation Software is used to recombine heavy and light chain antibody sequences and subsequently select desired sequences to populate a 96-well plate output file, which serves as an order file for purchasing DNA sequences in nucleotide format from a DNA Synthesis Company. This example illustrates the software requirements as well as the controls the user encounters when using the DNA Sequence Compilation Software, including Main Menu Options and Explorer Bar Options.
A. Software Overview
[1012] The software is designed to run on a computer that uses Windows XP or Windows Vista as the operating system. The software is installed by running the executable file setup.exe and installing the program in a folder of the users choice. Once installed, the user encounters Main Menu Options as well as Explorer Bar Options. The details are described below.
[1013] 1. System Requirements
[1014] The DNA Sequence Compilation Software has been designed to run on a local machine as a stand alone application. [1015] Operating System: Windows XP/Vista 32-bit Edition [1016] Internet Connection: Not Required [1017] Database Server: None
[1018] 2. Installation of the DNA Sequence Compilation Tool Program
[1019] To install the program, run the executable file setup.exe and install the program in a folder of your choice.
[1020] 3. Main Menu Options (see FIG. 12)
[1021] There are three main menu options. [1022] File--The file menu is used to close the application. [1023] Well-Plate--The well-plate menu is used to generate an order, clear the entire plate or a selected well, or configure the restriction sites for the well-plate order generation. [1024] Help--The help menu is used to find out what version of the software you are running.
[1025] 4. Explorer Bar Options
[1026] There are three explorer bar options (see FIG. 12; left column): [1027] Manual Compilation--Manual compilation contains items used to display the light chain or heavy chain manual compilation form, which allows users selection of germline segments. [1028] Auto Compilation--Auto compilation contains items used to display the light chain or heavy chain automatic compilation form. [1029] Well Plate--Well plate contains items used to display the well plate form.
[1030] C. Starting the Application
[1031] The software is launched either by clicking on the Fabrus DNA Compilation Tool on the All Programs Windows menu or by double-clicking the Fabrus DNA Compilation Tool shortcut on the desktop. A Splash Screen (FIG. 13) appears upon application startup and then fades into the background.
D. Generating an Output Plate Using the DNA Sequence Compilation Software
[1032] The DNA Sequence Compilation Software is used to recombine heavy and light chain antibody sequences and subsequently select desired sequences to populate a 96-well plate output file which serves as an order file for purchasing DNA sequences in nucleotide format from a DNA Synthesis Company. In addition to the plate file, a SequenceHistory.txt file is generated to keep track of all sequences ordered. Users can select either manual compilation or auto compilation in order to generate sequences and subsequently select individual sequences for inclusion into the 96-well plate grid, as described below in sections C.2-4. The software program also computes a diversity score for all sequences. The following sections describe user options for selecting and compiling germline sequences encoding variable heavy and light chains for DNA synthesis ordering.
[1033] 1. The 96-Well Plate Screen
[1034] When the application launches, the 96 Well-Plate screen is displayed. As is shown in FIG. 12, the 96-well plate is empty at application startup. The Main Menu is observed horizontally along the top of the screen, underneath the title DNA Sequence Compilation. The Explorer Bar is observed vertically along the left side of the screen. The user must populate the empty 96-well plate with DNA sequences that will be selected for manufacture as described herein below using either the manual compilation function or the automatic compilation function. The operating functions within the 96-well plate screen that permit population of the screen include the following:
[1035] a. Selecting Well Plate Restriction Sites
[1036] Restriction enzyme sequences are selected by the user from the Sequencedatabase.txt file to add to the compiled DNA sequence at the 5' and 3' ends for cloning purposes. The following commands are used to specify the 5' and 3' restriction sites that are added to the plate order according to antibody chain type:
[1037] Heavy Chains: [1038] Well Plate>Configuration>Heavy Chains>5' End.fwdarw.choose; and Well Plate>Configuration>Heavy Chains>3' End.fwdarw.choose.
[1039] Kappa Light Chains: [1040] Well Plate>Configuration>Kappa Light Chains>5' End.fwdarw.choose; and Well Plate>Configuration>Kappa Light Chains>3' End.fwdarw.choose.
[1041] Lambda Light Chains: [1042] Well Plate>Configuration>Lambda Light Chains>5' End.fwdarw.choose; and WellPlate>Configuration>Lambda Light Chains>3' End.fwdarw.choose.
[1043] This defines the restriction site that is prefixed and suffixed to each sequence chain type in the order file and can be viewed in the IDT Order Well Plate Start/Stop Restriction Sites box (See FIG. 12 and FIG. 14). The compilation software includes a function to internally search compiled sequences for the chosen restriction site and modify the nucleotide sequence in accord with codon usage to remove any internal restriction sites that can be incompatible for cloning purposes. This is completed prior to placing the sequences in the well plate for ordering such that only those sequences "with restriction sites silenced" are ordered. This is described in Example 2 in the section entitled "Codon usage Tables," and in Section 2 below.
[1044] b. Placing Sequences in the Well Plate
[1045] The selection of sequences is performed from either manual compilation or auto compilation screens as described below in Sections C.2 and C.3. As described below, in the manual compilation or auto compilation screens, following compilation, sequences are selected by checking the "Selected" column (for auto compilation) or clicking the "Add to Well Plate" button (for manual compilation).
[1046] c. Displaying Well Sequence Information
[1047] The sequence of a compiled sequence placed in a well is displayed by clicking on the well. The well for the selected sequence is highlighted. For example, FIG. 14 shows a model screen for a single 96-well plate containing compiled sequences which the user has selected for ordering. In row A of the Figure, columns 1-3 (shaded in medium gray) indicate a cell with a compiled heavy chain sequence; columns 3-6 (shaded in light gray) indicate a cell with a compiled light chain sequence. The highlighted cell (row A, column 5, indicated in dark shading) indicates the cell for which the individual data is observed in the Sequence Information box at the bottom of the display.
[1048] The Sequence Information text box is then updated with the sequence information for that well. The Sequence Information box lists all the pertinent information for a particular sequence, including the name, the nucleotide sequence, the amino acid sequence, the diversity score and individual component titles for the V(D)J recombination. For example, in FIG. 14, the highlighted sequence is a variable light chain compiled from the V.sub.L germline segment V3-4 and the J.sub.L germline segment IGLJ7*01.
[1049] d. Generating an Order
[1050] An order is generated for the 96-well plate regardless of how many samples are placed in the plate, i.e. from one (1) to 96 sequences can be ordered from a 96-well plate. To generate an order, the option Well Plate>Generate Order is chosen and a filename is inputted when requested. The order file is saved in a comma separated variable (.csv) format. Example 4 describes compilation, output and ordering of exemplary sequences compiled using the Software compilation software.
[1051] e. Clearing a Selected Well of a Well Plate
[1052] In order to clear a specific well in the plate, click on the selected well, then choose WellPlate>Clear>Selected Well or, alternatively, right click on the desired well and select "Clear Selected Well".
[1053] f. Clearing all Sequences from an Entire Well Plate
[1054] The Well Plate is not automatically cleared when an order is generated. In order to clear the entire Well-Plate, choose Well Plate>Clear>Entire Plate.
[1055] 2. Manual Compilation of Sequences
[1056] Compiled sequences can be generated manually by user selection using the Manual Compilation option.
[1057] a. Manual Compilation of Light Chains
[1058] To manually compile a light chain, choose the option Manual Compilation>Light Chain from the explorer bar. The manual compilation screen is displayed in FIG. 15. The manual compilation light chain screen allows the user to compile either a Lambda or Kappa sequence. This option is available under the heading Type and is chosen by selecting the circle in front of either Lambda or Kappa at the top of the screen. The software does not allow the combination of a V and a J sequence or vice-versa.
[1059] i. Compiling Lambda Light Chains
[1060] The following steps show how to compile a Lambda Light Chain. [1061] Select Type>Lambda. [1062] Select the desired V.sub..lamda. Sequence from the VL Sequence Combo box. All of the available V.sub..lamda. nucleotide sequences that are entered in the SequenceDatabase.txt file appear in the drop down menu box underneath. [1063] Select the desired J.sub..lamda. Sequence from the JL Sequence Combo Box. All of the available J.sub..lamda. nucleotide sequences that are entered in the SequenceDatabase.txt file appear in the drop down menu box underneath. [1064] Click "Add to Well Plate" button to add the compiled sequence to the well plate.
[1065] Two boxes are observed under the Dna Sequence heading. The "No Silencing of restricted Sites" box displays the individual V.sub..lamda. and J.sub..lamda. nucleotide sequences plus any V-J joints generated in accord with the sequence compilation rules. The corresponding encoded amino acid sequence also is displayed. For example, in FIG. 15, the first sequence displayed in the "no silencing of restricted sites" box corresponds to the selected V.sub..lamda. sequence (in this case V1-11). Following this is a depiction of three nucleotides "cct" encoding proline (P), which correspond to the joint region created by performance of the compilation method in order to generate a functional nucleic acid molecule. The last sequence displayed corresponds to the selected J.sub..lamda. sequence (in this case IGLJ1*01). The "With Restriction Sites Silenced" box displays the compiled sequence containing any necessary nucleotide modifications to the sequence to silence restriction sites, while maintaining codon usage. Any restriction sites silenced by the software are displayed in the "Notes" box. The sequence with the restriction sites silenced is the sequence that is placed in the order file.
[1066] (ii) Compiling Kappa Light Chains
[1067] The following steps show how to compile a Kappa Light Chain. [1068] Select Type>Kappa. [1069] Select the desired V.sub..kappa. Sequence from the VK Sequence Combo box. All of the available V.sub..kappa. nucleotide sequences that are entered in the SequenceDatabase.txt file appear in the box underneath. [1070] Select the desired J.sub..kappa. Sequence from the JK Sequence Combo Box. All of the available J.sub..kappa. nucleotide sequences that are entered in the SequenceDatabase.txt file appear in the box underneath. [1071] Click "Add to Well Plate" button to add the compiled sequence to the well plate. All compiled sequences are added to the well plate for ordering for DNA synthesis.
[1072] As above for compiling lambda light chain sequences, two boxes are observed under the Dna Sequence heading. The "No Silencing of restricted Sites" box displays the individual V.kappa. and J.kappa. sequences plus any V-J joints generated. The "With Restriction Sites Silenced" box displays the compiled sequence with any sequence modifications to silence selected restriction sites. Any restriction sites silenced by the software are displayed in the "Notes" box. The sequence with the restriction sites silenced is the sequence that is placed in the order file.
[1073] b. Manual Compilation of Heavy Chains
[1074] To manually compile a heavy chain, choose the option Manual Compilation>Heavy Chain on the explorer bar. The manual compilation screen is displayed in FIG. 16. The manual compilation heavy chain screen allows the user to concatenate V.sub.H, D.sub.H and J.sub.H sequences. The following steps show how to compile a heavy chain. [1075] Select the desired V.sub.H Sequence from the VH Sequence Combo box. All of the available V.sub.H nucleotide sequences that are entered in the SequenceDatabase.txt file appear in the drop down menu box underneath. [1076] Select the desired D.sub.H Sequence from the DH Sequence Combo Box. All of the available D.sub.H nucleotide sequences that are entered in the SequencDatabase.txt file appear in the drop down menu box underneath. [1077] Select the desired D.sub.H reading frame from the DH Reading Frame Box. The GRAVY value is updated according to the reading frame selection. The user should select the reading frame that corresponds to the lowest GRAVY value. [1078] Select the desired J.sub.H Sequence from the JH Sequence Combo Box. All of the available J.sub.H nucleotide sequences that are entered in the SequenceDatabase.txt file appear in the box underneath. [1079] Click "Add to Well Plate" button to add the compiled sequence to the well plate. All compiled sequences are added to the well plate for ordering for DNA synthesis.
[1080] Two boxes are observed in under the Dna Sequence heading. The "No Silencing of restricted Sites" box displays the individual V.sub.H, D.sub.H and J.sub.H sequences plus any V-D-J joints generated in accord with the sequence compilation rules. The corresponding encoded amino acid sequence also is displayed. For example, in FIG. 16, the first sequence displayed in the "no silencing of restricted sites" box corresponds to the selected V.sub.H sequence (in this case VH1-18). Following this is a depiction of three nucleotides "gag" encoding glutamic acid (E), which correspond to the V-D joint region created by performance of the compilation method in order to generate a functional nucleic acid molecule. The next sequence displayed corresponds to the selected D.sub.H sequence (in this case IGHD1-1*01). That sequence is followed by depiction of three nucleotides "acg" encoding threonine (T), which correspond to the D-J joint region created by performance of the compilation method in order to generate a functional nucleic acid molecule. The last sequence displayed corresponds to the J.sub.H selected sequence (in this case IGHJ1*01). The "With Restriction Sites Silenced" box displays the compiled sequence with any restriction sites that are present silenced. Any restriction sites silenced are displayed in the "Notes" box. The sequence with the restriction sites silenced is the sequence that is placed in the order file.
[1081] 3. Auto Compilation of Sequences
[1082] Compiled chains are generated automatically by using the Auto Compilation option. As discussed in Example 4, compilation of all sequences is performed mechanically by the computer upon initiation and stored in files. The auto compilation function permits the user to view these sequences.
[1083] a. Auto Compilation of Light Chains
[1084] To automatically compile a light chain, choose the option Auto Compilation>Light Chain on the explorer bar. The auto compilation screen is displayed in FIG. 17. The auto compilation light chain screen displays all the possible combinations of V.sub..kappa. and J.sub..kappa., and V.sub..lamda. and J.sub..lamda. sequences.
[1085] The auto compilation screen shows a grid with the following columns: [1086] Selected--This column contains a box that is either checked or unchecked to add or remove a sequence to or from the well plate. [1087] Rank--This column indicates the rank order in terms of diversity. A value of 1 indicates the most diverse sequence. [1088] Cluster--This column lists the cluster that the sequence is grouped into. The cluster number is simply an identifier and its value is not associated to any sort of ranking. [1089] Diversity Score--This column lists a score based on the root mean square of the BLAST Bit score results of this sequence against all other sequences in this list. A low value indicates a more diverse sequence. [1090] V Sequence--This column lists the V sequence identifier that is used for this sequence. [1091] J Sequence--This column lists the J sequence identifier that is used for this sequence. [1092] Amino Acid Sequence--This column lists the compiled sequence in protein format.
[1093] b. Auto Compilation of Heavy Chains
[1094] To automatically compile heavy chains, choose the option Auto Compilation>Heavy Chain on the explorer bar. The auto compilation screen is displayed in FIG. 19. The auto compilation heavy chain screen displays all the possible combinations of V.sub.H, D.sub.H and J.sub.H sequences.
[1095] The auto compilation screen shows a grid with the following columns: [1096] Selected--This column contains a box that is either checked or unchecked to add or remove a sequence to or from the well plate. [1097] Rank--This column indicates the rank order in terms of diversity. A value of 1 indicates the most diverse sequence. [1098] Cluster--This column lists the cluster that the sequence is grouped into. The cluster number is simply an identifier and its value is not associated to any sort of ranking. [1099] Diversity Score--This column lists a score based on the root mean square of the BLAST Bit score results of this sequence against all other sequences in this list. A low value indicates a more diverse sequence. [1100] V Sequence--This column lists the V sequence identifier that is used for this sequence. [1101] D Sequence--This column lists the D sequence identifier that is used for this sequence. [1102] J Sequence--This column lists the J sequence identifier that is used for this sequence. [1103] Amino Acid Sequence--This column lists the compiled sequence in protein format.
[1104] 4. Other Functionalities Associated with Compilation of Light Chain and Heavy Chain Sequences
[1105] a. Changing the Sorting Option of the Auto Compilation Screen
[1106] The view of the sequences listed in the auto compilation screen can be reorganized by clicking on the desired column header. For example, to view sequences based on diversity score, the column header "Diversity Score" is clicked and the sequences are reordered in terms of lowest to highest diversity score.
[1107] b. Status Indicators
[1108] The auto compilation screen uses color conventions to identify the status of compiled sequences. For example, grey indicates the sequence has already been ordered, blue indicates the sequence has neither been ordered nor placed in the 96-Well Plate, and bold blue indicates the sequence has been placed in the 96-Well Plate. For example, the bolded items in rows 022, 023 and 024 in FIG. 17 indicate that the compiled sequences have been placed in the 96-well plate for ordering, which also is indicated by the check mark in the "selected" column. For example, the bolded items in rows 0103-0107 in FIG. 19 indicate that the compiled sequences have been placed in the 96-well plate for ordering, which also is indicated by the check mark in the "selected" column.
[1109] c. Placing Multiple Sequences in the 96-Well Plate
[1110] Multiple sequences can be placed into the 96-well plate by clicking on the box for multiple selected sequences. This can be down by selecting a first sequence with the mouse, and then, holding down the Shift key and clicking on the box for the last selected sequence. Alternatively, the Ctrl key can be used to select individual sequences by holding down the Ctrl key and clicking on the individual boxes for all selected sequences. Once VL sequences are selected to add to the 96-well plate, right Click on the row header and select "Copy to Well Plate". All selected sequences are added to the well plate for ordering for DNA synthesis.
[1111] d. Placing a Single Sequence in the 96-Well Plate
[1112] To place a single sequences into the 96-well plate, click the box in the "Selected" Column; or alternatively, right Click on the Row Header for the sequence and click "Copy to Well Plate". Only the selected sequence is added to the well place for ordering for DNA synthesis.
[1113] e. BLAST of a Selected Sequence Against all the Other Sequences in the Compilation.
[1114] Once the sequences are compiled by auto compilation, the software allows the user the option of performing a BLAST search of a single sequence against all the other sequences in the list. This function provides an output indicator, the BLAST bit score, which is a value calculated from the number of gaps and substitutions associated with each aligned sequence. The higher the score, the more significant the alignment. This data is useful if the user is trying to select sequences that have either the most diversity or alternatively, the least diversity to every other selected sequence. An example of generated data for selected VL sequences is illustrated in FIG. 18 and an example of generated data for selected VH sequences is illustrated in FIG. 20.
[1115] The compiled sequences are then compared in order to determine the similarity of each compiled sequence to every other compiled sequence and this information is used to generate a diversity score. For example, the NCBI utility BLAST can be used to find regions of local similarity between sequences, by comparing nucleotide or protein sequences and calculating the statistical significance of matches.
[1116] To perform BLAST, the sequence to be compared is selected by clicking on the sequence. The user can select the BLAST option by right clicking the sequence. For example, for VL sequences, FIG. 18 illustrates a BLAST grid form generated by selecting on the V4-2_IGLJ2*01 sequence on a manual compilation or auto compilation screen, followed by right clicking the sequence and selecting Blast. A new grid form was generated, illustrated in FIG. 18, which provides the Blast bit score of other compiled sequences aligned with the selected sequence. For example, for VH sequences, FIG. 20 illustrates a BLAST grid form generated by selecting on the VH4-39_IGHD5-24*01_IGHJ6*01 sequence on a manual compilation or auto compilation screen, followed by right clicking the sequence and selecting Blast. A new grid was generated, illustrated in FIG. 20, which provides the Blast bit score of other compiled sequences aligned with the selected sequence. Sequences can be selected from this new grid form for insertion into the 96-well plate file for ordering for DNA synthesis as indicated by checking the box in the "selected" column.
[1117] The BLAST form is a grid with the columns: [1118] Selected--This column contains a box that is either checked or unchecked to add or remove a sequence to or from the well plate. [1119] Rank--This column indicates the rank order in terms of diversity. A value of 1 indicates the most diverse sequence. [1120] BLAST Bit Score--This column indicates the BLAST Bit score for this particular sequence against the selected sequence. [1121] V Sequence--This column lists the V sequence identifier that is used for this sequence. [1122] J Sequence--This column lists the J sequence identifier that is used for this sequence. [1123] Amino Acid Sequence--This column lists the compiled sequence in protein format.
Example 6
Generation of Heavy and Light Chain Germline Recombined DNA Sequences
[1124] This example describes methods for generating heavy and light chain DNA sequences by the DNA Sequence Compilation Software. The first step in compilation is to select either manual compilation or automatic compilation of sequences as discussed in Example 5 above. Manual compilation is useful as it gives the user complete control over the selected sequences. This allows the user to cater a library to any desired circumstance, such as creating a library where every sequence contains a particular segment that is the same, or by controlling the diversity by selecting sequences that are different (e.g. from different gene families).
[1125] Using manual compilation germline segments were selected by selecting a germline segment from each gene family of a V.sub.H, D.sub.H, J.sub.H, V.kappa., J.kappa., V.sub..lamda. or J.sub..lamda. germline segment. Compilation of the variable heavy chain was performed separate from the light chain. Also compilation of the variable kappa light chain was performed separate from the variable lambda chain. Hence, separate 96-well plate files were created for the variable heavy chain, variable kappa light chain and variable lambda light chain.
[1126] For example, for a variable heavy chain, one V.sub.H germline segment from the IGHV1-18 gene family was selected, i.e. IGHV1-18*01; one V.sub.H germline segment from the IGHV1-2 gene family was selected, i.e. IGHV1-2*01; and so on. Similar selections were made for D.sub.H germline segments and J.sub.H germline segments. The D.sub.H germline segments were selected in a reading frame to give the lowest gravy score where possible. A combination of V.sub.HD.sub.HJ.sub.H germline segments were made one at time in the Manual compilation screen. In addition, the restriction enzyme sites NcoI and Nhe I were selected from the "well plate" menu for each V.sub.HD.sub.HJ.sub.H combination for inclusion at the 5' and 3' ends of the recombined segments to allow for subcloning into appropriate vectors (see Example 8). This resulted in the software automatically viewing in the DNA Sequence Window the sequence of the selected sequence with "no silencing of restricted sites" and "with restriction sites silenced," (i.e. in this case any NcoI or NheI restriction sites present internally in the sequence were silenced by modification using the codon usage table). Generated V-D and D-J joints also were created by the software in accordance with the sequence compilation rules and indicated in the DNA sequence window (an exemplary selection is set forth in FIG. 16). Once germline segment components for a rearranged sequence were selected, it was "added to the well plate." This was repeated for all variable heavy chain germline segment combinations.
[1127] Similar selections also were made for the V.kappa. and J.kappa. light chain germline by selecting a combination of V.kappa. and J.kappa. germline segments (only one gene family member from each germline segment was ever selected) one at a time in the manual compilation screen. In addition, the restriction enzyme sites Nco I and BsiW I were selected from the "well plate" menu for each V.kappa. and J.kappa. combination for inclusion at the 5' and 3' ends of the recombined segments to allow for subcloning into appropriate vectors (see Example 8). This resulted in the software automatically displaying in the DNA Sequence Window the sequence of the selected sequence with "no silencing of restricted sites" and "with restriction sites silenced," (i.e. in this case any Nco I or BsiW I restriction sites present internally in the sequence were silenced by nucleotide modification using the codon usage table.) Generated V-J joints also were created by the software in accordance with the sequence compilation rules and indicated in the DNA sequence window. Once germline segment components for a rearranged sequence were selected, it was "added to the well plate." This was repeated for all variable kappa light chain germline segment combinations.
[1128] Similar selections also were made for the V.sub..lamda. or J.sub..lamda. light chain germline segments by selecting a combination of V.sub..lamda. or J.sub..lamda. germline segments (only one gene family member from each germline segment was ever selected) one at at time in the manual compilation screen. In addition, the restriction enzyme sites Nco I and Avr II were selected from the "well plate" menu for each V.sub..lamda. and J.sub..lamda. combination for inclusion at the 5' and 3' ends of the recombined segments to allow for subcloning into appropriate vectors (see Example 8) This resulted in the software automatically displaying in the DNA Sequence Window the sequence of the selected sequence with "no silencing of restricted sites" and "with restriction sites silenced," (i.e. in this case any Nco I or Avr II restriction sites present internally in the sequence were silenced by nucleotide modification using the codon usage table.) Generated V-J joints also were created by the software in accordance with the sequence compilation rules and indicated in the DNA sequence window (an exemplary selection is set forth in FIG. 15). Once germline segment components for a rearranged sequence were selected, it was "added to the well plate." This was repeated for all variable lambda light chain germline segment combinations.
[1129] The choice of Nhe I, Bsi WI and Avr II sites allow for complete amino acid conservation for their respective heavy and light chains thereby maintaining the natural human V and C regions.
[1130] Once all sequences were selected, the selected sequences were viewed under the 96-well plate view (see e.g., FIG. 12 and FIG. 14). A DNA synthesis order was generated using the Generate Order function. The DNA Compilation Software outputted the selected sequences in arrays that mapped to the 96-well format. The sequences in this format were sent to a DNA synthesis vendor (Genscript Corp.) to generate synthetic antibody variable heavy and light chain sequences. The synthesized nucleic acid molecules were returned and identified by the 96-well format. Table 22 lists sequences generated by the Sequence Compilation Software, ordered and synthesized.
TABLE-US-00038 TABLE 22 Exemplary Sequences Generated by DNA Compilation Software Heavy Chain SEQ Number Name ID NO. 3 gnl|Fabrus|VH1-18_IGHD1-26*01_IGHJ2*01 454 4 gnl|Fabrus|VH1-18_IGHD2-21*01_IGHJ2*01 455 5 gnl|Fabrus|VH1-18_IGHD3-16*01_IGHJ6*01 456 6 gnl|Fabrus|VH1-18_IGHD3-22*01_IGHJ4*01 457 7 gnl|Fabrus|VH1-18_IGHD4-23*01_IGHJ1*01 458 8 gnl|Fabrus|VH1-18_IGHD5-12*01_IGHJ4*01 459 9 gnl|Fabrus|VH1-18_IGHD6-6*01_IGHJ1*01 460 10 gnl|Fabrus|VH1-2_IGHD1-1*01_IGHJ3*01 461 11 gnl|Fabrus|VH1-24_IGHD1-7*01_IGHJ4*01 462 12 gnl|Fabrus|VH1-24_IGHD2-15*01_IGHJ2*01 463 13 gnl|Fabrus|VH1-24_IGHD3-10*01_IGHJ4*01 464 14 gnl|Fabrus|VH1-24_IGHD3-16*01_IGHJ4*01 465 15 gnl|Fabrus|VH1-24_IGHD4-23*01_IGHJ2*01 466 16 gnl|Fabrus|VH1-24_IGHD5-12*01_IGHJ4*01 467 17 gnl|Fabrus|VH1-24_IGHD5-18*01_IGHJ6*01 468 18 gnl|Fabrus|VH1-24_IGHD6-19*01_IGHJ4*01 469 19 gnl|Fabrus|VH1-3_IGHD2-15*01_IGHJ2*01 470 20 gnl|Fabrus|VH1-3_IGHD2-2*01_IGHJ5*01 471 21 gnl|Fabrus|VH1-3_IGHD3-9*01_IGHJ6*01 472 22 gnl|Fabrus|VH1-3_IGHD4-23*01_IGHJ4*01 473 23 gnl|Fabrus|VH1-3_IGHD5-18*01_IGHJ4*01 474 24 gnl|Fabrus|VH1-3_IGHD6-6*01_IGHJ1*01 475 25 gnl|Fabrus|VH1-3_IGHD7-27*01_IGHJ4*01 476 26 gnl|Fabrus|VH1-45_IGHD1-26*01_IGHJ4*01 477 27 gnl|Fabrus|VH1-45_IGHD2-15*01_IGHJ6*01 478 28 gnl|Fabrus|VH1-45_IGHD2-8*01_IGHJ3*01 479 29 gnl|Fabrus|VH1-45_IGHD3-10*01_IGHJ4*01 480 30 gnl|Fabrus|VH1-45_IGHD3-16*01_IGHJ2*01 481 31 gnl|Fabrus|VH1-45_IGHD4-23*01_IGHJ4*01 482 32 gnl|Fabrus|VH1-45_IGHD5-24*01_IGHJ4*01 483 33 gnl|Fabrus|VH1-45_IGHD6-19*01_IGHJ4*01 484 34 gnl|Fabrus|VH1-45_IGHD7-27*01_IGHJ6*01 485 35 gnl|Fabrus|VH1-46_IGHD1-26*01_IGHJ4*01 486 36 gnl|Fabrus|VH1-46_IGHD2-15*01_IGHJ2*01 487 37 gnl|Fabrus|VH1-46_IGHD3-10*01_IGHJ4*01 488 38 gnl|Fabrus|VH1-46_IGHD4-17*01_IGHJ4*01 489 39 gnl|Fabrus|VH1-46_IGHD5-18*01_IGHJ4*01 490 40 gnl|Fabrus|VH1-46_IGHD6-13*01_IGHJ4*01 491 41 gnl|Fabrus|VH1-46_IGHD6-6*01_IGHJ1*01 492 42 gnl|Fabrus|VH1-46_IGHD7-27*01_IGHJ2*01 493 43 gnl|Fabrus|VH1-58_IGHD1-26*01_IGHJ4*01 494 44 gnl|Fabrus|VH1-58_IGHD2-15*01_IGHJ2*01 495 45 gnl|Fabrus|VH1-58_IGHD3-10*01_IGHJ6*01 496 46 gnl|Fabrus|VH1-58_IGHD4-17*01_IGHJ4*01 497 47 gnl|Fabrus|VH1-58_IGHD5-18*01_IGHJ4*01 498 48 gnl|Fabrus|VH1-58_IGHD6-6*01_IGHJ1*01 499 49 gnl|Fabrus|VH1-58_IGHD7-27*01_IGHJ5*01 500 50 gnl|Fabrus|VH1-69_IGHD1-1*01_IGHJ6*01 501 51 gnl|Fabrus|VH1-69_IGHD1-14*01_IGHJ4*01 502 52 gnl|Fabrus|VH1-69_IGHD2-2*01_IGHJ4*01 503 53 gnl|Fabrus|VH1-69_IGHD2-8*01_IGHJ6*01 504 54 gnl|Fabrus|VH1-69_IGHD3-16*01_IGHJ4*01 505 55 gnl|Fabrus|VH1-69_IGHD3-3*01_IGHJ4*01 506 56 gnl|Fabrus|VH1-69_IGHD3-9*01_IGHJ6*01 507 57 gnl|Fabrus|VH1-69_IGHD4-17*01_IGHJ4*01 508 58 gnl|Fabrus|VH1-69_IGHD5-12*01_IGHJ4*01 509 59 gnl|Fabrus|VH1-69_IGHD5-24*01_IGHJ6*01 510 60 gnl|Fabrus|VH1-69_IGHD6-19*01_IGHJ4*01 511 61 gnl|Fabrus|VH1-69_IGHD6-6*01_IGHJ1*01 512 62 gnl|Fabrus|VH1-69_IGHD7-27*01_IGHJ4*01 513 63 gnl|Fabrus|VH1-8_IGHD1-26*01_IGHJ4*01 514 64 gnl|Fabrus|VH1-8_IGHD2-15*01_IGHJ6*01 515 65 gnl|Fabrus|VH1-8_IGHD2-2*01_IGHJ6*01 516 66 gnl|Fabrus|VH1-8_IGHD3-10*01_IGHJ4*01 517 67 gnl|Fabrus|VH1-8_IGHD4-17*01_IGHJ4*01 518 68 gnl|Fabrus|VH1-8_IGHD5-5*01_IGHJ4*01 519 69 gnl|Fabrus|VH1-8_IGHD7-27*01_IGHJ4*01 520 70 gnl|Fabrus|VH2-26_IGHD1-20*01_IGHJ4*01 521 71 gnl|Fabrus|VH2-26_IGHD2-15*01_IGHJ2*01 522 72 gnl|Fabrus|VH2-26_IGHD2-2*01_IGHJ4*01 523 73 gnl|Fabrus|VH2-26_IGHD3-10*01_IGHJ4*01 524 74 gnl|Fabrus|VH2-26_IGHD3-9*01_IGHJ6*01 525 75 gnl|Fabrus|VH2-26_IGHD4-11*01_IGHJ4*01 526 76 gnl|Fabrus|VH2-26_IGHD5-12*01_IGHJ4*01 527 77 gnl|Fabrus|VH2-26_IGHD5-18*01_IGHJ4*01 528 78 gnl|Fabrus|VH2-26_IGHD6-13*01_IGHJ4*01 529 79 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 530 80 gnl|Fabrus|VH2-26_IGHD7-27*01_IGHJ4*01 531 81 gnl|Fabrus|VH2-5_IGHD1-1*01_IGHJ5*01 532 82 gnl|Fabrus|VH2-5_IGHD2-15*01_IGHJ6*01 533 83 gnl|Fabrus|VH2-5_IGHD3-16*01_IGHJ4*01 534 84 gnl|Fabrus|VH2-5_IGHD3-9*01_IGHJ6*01 535 85 gnl|Fabrus|VH2-5_IGHD5-12*01_IGHJ4*01 536 86 gnl|Fabrus|VH2-5_IGHD6-13*01_IGHJ4*01 537 87 gnl|Fabrus|VH2-5_IGHD7-27*01_IGHJ2*01 538 88 gnl|Fabrus|VH2-70_IGHD1-1*01_IGHJ2*01 539 89 gnl|Fabrus|VH2-70_IGHD2-15*01_IGHJ2*01 540 90 gnl|Fabrus|VH2-70_IGHD3-22*01_IGHJ4*01 541 91 gnl|Fabrus|VH2-70_IGHD3-9*01_IGHJ6*01 542 92 gnl|Fabrus|VH2-70_IGHD5-12*01_IGHJ4*01 543 93 gnl|Fabrus|VH2-70_IGHD7-27*01_IGHJ2*01 544 94 gnl|Fabrus|VH3-11_IGHD1-26*01_IGHJ4*01 545 95 gnl|Fabrus|VH3-11_IGHD2-2*01_IGHJ6*01 546 96 gnl|Fabrus|VH3-11_IGHD3-16*01_IGHJ4*01 547 97 gnl|Fabrus|VH3-11_IGHD3-9*01_IGHJ6*01 548 98 gnl|Fabrus|VH3-11_IGHD4-23*01_IGHJ5*01 549 99 gnl|Fabrus|VH3-11_IGHD5-18*01_IGHJ4*01 550 100 gnl|Fabrus|VH3-11_IGHD6-19*01_IGHJ6*01 551 101 gnl|Fabrus|VH3-11_IGHD6-6*01_IGHJ1*01 552 102 gnl|Fabrus|VH3-11_IGHD7-27*01_IGHJ4*01 553 103 gnl|Fabrus|VH3-13_IGHD1-26*01_IGHJ4*01 554 104 gnl|Fabrus|VH3-13_IGHD2-8*01_IGHJ5*01 555 105 gnl|Fabrus|VH3-13_IGHD3-3*01_IGHJ1*01 556 106 gnl|Fabrus|VH3-13_IGHD3-9*01_IGHJ6*01 557 107 gnl|Fabrus|VH3-13_IGHD4-23*01_IGHJ5*01 558 108 gnl|Fabrus|VH3-13_IGHD5-5*01_IGHJ4*01 559 109 gnl|Fabrus|VH3-13_IGHD6-6*01_IGHJ1*01 560 110 gnl|Fabrus|VH3-13_IGHD7-27*01_IGHJ5*01 561 111 gnl|Fabrus|VH3-15_IGHD1-26*01_IGHJ4*01 562 112 gnl|Fabrus|VH3-15_IGHD2-15*01_IGHJ2*01 563 113 gnl|Fabrus|VH3-15_IGHD2-15*01_IGHJ6*01 564 114 gnl|Fabrus|VH3-15_IGHD3-10*01_IGHJ4*01 565 115 gnl|Fabrus|VH3-15_IGHD3-9*01_IGHJ2*01 566 116 gnl|Fabrus|VH3-15_IGHD5-12*01_IGHJ4*01 567 117 gnl|Fabrus|VH3-15_IGHD6-6*01_IGHJ1*01 568 118 gnl|Fabrus|VH3-16_IGHD1-1*01_IGHJ1*01 569 119 gnl|Fabrus|VH3-16_IGHD1-7*01_IGHJ6*01 570 120 gnl|Fabrus|VH3-16_IGHD2-15*01_IGHJ2*01 571 121 gnl|Fabrus|VH3-16_IGHD2-2*01_IGHJ2*01 572 122 gnl|Fabrus|VH3-16_IGHD3-10*01_IGHJ4*01 573 123 gnl|Fabrus|VH3-16_IGHD4-4*01_IGHJ2*01 574 124 gnl|Fabrus|VH3-16_IGHD5-24*01_IGHJ4*01 575 125 gnl|Fabrus|VH3-16_IGHD6-13*01_IGHJ4*01 576 126 gnl|Fabrus|VH3-16_IGHD7-27*01_IGHJ2*01 577 127 gnl|Fabrus|VH3-20_IGHD1-14*01_IGHJ4*01 578 128 gnl|Fabrus|VH3-20_IGHD2-15*01_IGHJ2*01 579 129 gnl|Fabrus|VH3-20_IGHD2-8*01_IGHJ4*01 580 130 gnl|Fabrus|VH3-20_IGHD3-10*01_IGHJ4*01 581 131 gnl|Fabrus|VH3-20_IGHD3-9*01_IGHJ6*01 582 132 gnl|Fabrus|VH3-20_IGHD4-23*01_IGHJ4*01 583 133 gnl|Fabrus|VH3-20_IGHD5-12*01_IGHJ4*01 584 134 gnl|Fabrus|VH3-20_IGHD6-13*01_IGHJ4*01 585 135 gnl|Fabrus|VH3-20_IGHD7-27*01_IGHJ2*01 586 136 gnl|Fabrus|VH3-21_IGHD1-26*01_IGHJ4*01 587 137 gnl|Fabrus|VH3-21_IGHD2-2*01_IGHJ5*01 588 138 gnl|Fabrus|VH3-21_IGHD3-22*01_IGHJ4*01 589 139 gnl|Fabrus|VH3-21_IGHD4-23*01_IGHJ5*01 590 140 gnl|Fabrus|VH3-21_IGHD5-24*01_IGHJ5*01 591 141 gnl|Fabrus|VH3-21_IGHD6-19*01_IGHJ1*01 592 142 gnl|Fabrus|VH3-21_IGHD7-27*01_IGHJ4*01 593 143 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ1*01 594 144 gnl|Fabrus|VH3-23_IGHD1-1*01_IGHJ4*01 595 145 gnl|Fabrus|VH3-23_IGHD1-20*01_IGHJ3*01 596 146 gnl|Fabrus|VH3-23_IGHD1-26*01_IGHJ4*01 597 147 gnl|Fabrus|VH3-23_IGHD2-15*01_IGHJ4*01 598 148 gnl|Fabrus|VH3-23_IGHD2-21*01_IGHJ1*01 599 149 gnl|Fabrus|VH3-23_IGHD3-10*01_IGHJ4*01 600 150 gnl|Fabrus|VH3-23_IGHD3-16*01_IGHJ4*01 601 151 gnl|Fabrus|VH3-23_IGHD3-22*01_IGHJ4*01 602 152 gnl|Fabrus|VH3-23_IGHD3-3*01_IGHJ5*01 603 153 gnl|Fabrus|VH3-23_IGHD4-11*01_IGHJ4*01 604 154 gnl|Fabrus|VH3-23_IGHD4-23*01_IGHJ2*01 605 155 gnl|Fabrus|VH3-23_IGHD5-12*01_IGHJ4*01 606 156 gnl|Fabrus|VH3-23_IGHD5-24*01_IGHJ1*01 607 157 gnl|Fabrus|VH3-23_IGHD5-5*01_IGHJ4*01 608 158 gnl|Fabrus|VH3-23_IGHD6-13*01_IGHJ4*01 609 159 gnl|Fabrus|VH3-23_IGHD6-25*01_IGHJ2*01 610 160 gnl|Fabrus|VH3-23_IGHD6-6*01_IGHJ1*01 611 161 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ4*01 612 162 gnl|Fabrus|VH3-23_IGHD7-27*01_IGHJ6*01 613 163 gnl|Fabrus|VH3-30_IGHD1-1*01_IGHJ6*01 614 164 gnl|Fabrus|VH3-30_IGHD1-26*01_IGHJ1*01 615 165 gnl|Fabrus|VH3-30_IGHD1-26*01_IGHJ4*01 616 166 gnl|Fabrus|VH3-30_IGHD2-15*01_IGHJ2*01 617 167 gnl|Fabrus|VH3-30_IGHD2-2*01_IGHJ6*01 618 168 gnl|Fabrus|VH3-30_IGHD3-10*01_IGHJ1*01 619 169 gnl|Fabrus|VH3-30_IGHD3-16*01_IGHJ6*01 620 170 gnl|Fabrus|VH3-30_IGHD4-17*01_IGHJ4*01 621 171 gnl|Fabrus|VH3-30_IGHD5-12*01_IGHJ4*01 622 172 gnl|Fabrus|VH3-30_IGHD5-18*01_IGHJ1*01 623 173 gnl|Fabrus|VH3-30_IGHD6-13*01_IGHJ4*01 624 174 gnl|Fabrus|VH3-30_IGHD6-6*01_IGHJ1*01 625 175 gnl|Fabrus|VH3-35_IGHD1-1*01_IGHJ2*01 626 176 gnl|Fabrus|VH3-35_IGHD1-20*01_IGHJ6*01 627 177 gnl|Fabrus|VH3-35_IGHD2-15*01_IGHJ2*01 628 178 gnl|Fabrus|VH3-35_IGHD2-21*01_IGHJ6*01 629 179 gnl|Fabrus|VH3-35_IGHD3-10*01_IGHJ4*01 630 180 gnl|Fabrus|VH3-35_IGHD3-9*01_IGHJ6*01 631 181 gnl|Fabrus|VH3-35_IGHD5-12*01_IGHJ4*01 632 182 gnl|Fabrus|VH3-35_IGHD6-13*01_IGHJ4*01 633 183 gnl|Fabrus|VH3-35_IGHD7-27*01_IGHJ1*01 634 184 gnl|Fabrus|VH3-38_IGHD1-14*01_IGHJ5*01 635 185 gnl|Fabrus|VH3-38_IGHD1-20*01_IGHJ6*01 636 186 gnl|Fabrus|VH3-38_IGHD2-15*01_IGHJ6*01 637 187 gnl|Fabrus|VH3-38_IGHD2-2*01_IGHJ1*01 638 188 gnl|Fabrus|VH3-38_IGHD3-10*01_IGHJ4*01 639 189 gnl|Fabrus|VH3-38_IGHD3-16*01_IGHJ1*01 640 190 gnl|Fabrus|VH3-38_IGHD4-17*01_IGHJ2*01 641 191 gnl|Fabrus|VH3-38_IGHD5-24*01_IGHJ3*01 642 192 gnl|Fabrus|VH3-38_IGHD6-6*01_IGHJ1*01 643 193 gnl|Fabrus|VH3-38_IGHD7-27*01_IGHJ6*01 644 194 gnl|Fabrus|VH3-43_IGHD1-26*01_IGHJ5*01 645 195 gnl|Fabrus|VH3-43_IGHD1-7*01_IGHJ6*01 646 196 gnl|Fabrus|VH3-43_IGHD2-2*01_IGHJ3*01 647 197 gnl|Fabrus|VH3-43_IGHD2-21*01_IGHJ6*01 648 198 gnl|Fabrus|VH3-43_IGHD3-16*01_IGHJ6*01 649 199 gnl|Fabrus|VH3-43_IGHD3-22*01_IGHJ4*01 650 200 gnl|Fabrus|VH3-43_IGHD4-23*01_IGHJ3*01 651 201 gnl|Fabrus|VH3-43_IGHD5-18*01_IGHJ5*01 652 202 gnl|Fabrus|VH3-43_IGHD6-13*01_IGHJ4*01 653 203 gnl|Fabrus|VH3-43_IGHD7-27*01_IGHJ1*01 654 204 gnl|Fabrus|VH3-48_IGHD6-6*01_IGHJ1*01 655 205 gnl|Fabrus|VH3-49_IGHD1-26*01_IGHJ4*01 656 206 gnl|Fabrus|VH3-49_IGHD1-7*01_IGHJ6*01 657 207 gnl|Fabrus|VH3-49_IGHD2-2*01_IGHJ6*01 658 208 gnl|Fabrus|VH3-49_IGHD2-8*01_IGHJ4*01 659 209 gnl|Fabrus|VH3-49_IGHD3-22*01_IGHJ4*01 660 210 gnl|Fabrus|VH3-49_IGHD3-9*01_IGHJ6*01 661 211 gnl|Fabrus|VH3-49_IGHD5-18*01_IGHJ4*01 662 212 gnl|Fabrus|VH3-49_IGHD6-13*01_IGHJ4*01 663 213 gnl|Fabrus|VH3-49_IGHD7-27*01_IGHJ1*01 664 214 gnl|Fabrus|VH3-53_IGHD1-14*01_IGHJ6*01 665 215 gnl|Fabrus|VH3-53_IGHD1-7*01_IGHJ1*01 666 216 gnl|Fabrus|VH3-53_IGHD2-2*01_IGHJ2*01 667 217 gnl|Fabrus|VH3-53_IGHD3-22*01_IGHJ3*01 668 218 gnl|Fabrus|VH3-53_IGHD4-23*01_IGHJ1*01 669 219 gnl|Fabrus|VH3-53_IGHD5-5*01_IGHJ4*01 670 220 gnl|Fabrus|VH3-53_IGHD6-13*01_IGHJ3*01 671 221 gnl|Fabrus|VH3-53_IGHD7-27*01_IGHJ4*01 672 222 gnl|Fabrus|VH3-64_IGHD1-26*01_IGHJ4*01 673 223 gnl|Fabrus|VH3-64_IGHD1-7*01_IGHJ6*01 674 224 gnl|Fabrus|VH3-64_IGHD2-2*01_IGHJ5*01 675 225 gnl|Fabrus|VH3-64_IGHD3-3*01_IGHJ4*01 676 226 gnl|Fabrus|VH3-64_IGHD4-17*01_IGHJ4*01 677 227 gnl|Fabrus|VH3-64_IGHD5-12*01_IGHJ4*01 678 228 gnl|Fabrus|VH3-64_IGHD6-19*01_IGHJ1*01 679 229 gnl|Fabrus|VH3-64_IGHD7-27*01_IGHJ4*01 680 230 gnl|Fabrus|VH3-66_IGHD6-6*01_IGHJ1*01 681 231 gnl|Fabrus|VH3-7_IGHD1-20*01_IGHJ3*01 682 232 gnl|Fabrus|VH3-7_IGHD1-7*01_IGHJ6*01 683 233 gnl|Fabrus|VH3-7_IGHD2-21*01_IGHJ5*01 684 234 gnl|Fabrus|VH3-7_IGHD2-8*01_IGHJ6*01 685 235 gnl|Fabrus|VH3-7_IGHD3-22*01_IGHJ3*01 686 236 gnl|Fabrus|VH3-7_IGHD3-9*01_IGHJ6*01 687 237 gnl|Fabrus|VH3-7_IGHD4-17*01_IGHJ4*01 688 238 gnl|Fabrus|VH3-7_IGHD5-12*01_IGHJ4*01 689 239 gnl|Fabrus|VH3-7_IGHD5-24*01_IGHJ4*01 690 240 gnl|Fabrus|VH3-7_IGHD6-19*01_IGHJ6*01 691 241 gnl|Fabrus|VH3-7_IGHD6-6*01_IGHJ1*01 692 242 gnl|Fabrus|VH3-7_IGHD7-27*01_IGHJ2*01 693 243 gnl|Fabrus|VH3-72_IGHD1-1*01_IGHJ4*01 694 244 gnl|Fabrus|VH3-72_IGHD2-15*01_IGHJ1*01 695
245 gnl|Fabrus|VH3-72_IGHD3-22*01_IGHJ4*01 696 246 gnl|Fabrus|VH3-72_IGHD3-9*01_IGHJ6*01 697 247 gnl|Fabrus|VH3-72_IGHD4-23*01_IGHJ2*01 698 248 gnl|Fabrus|VH3-72_IGHD5-18*01_IGHJ4*01 699 249 gnl|Fabrus|VH3-72_IGHD5-24*01_IGHJ6*01 700 250 gnl|Fabrus|VH3-72_IGHD6-6*01_IGHJ1*01 701 251 gnl|Fabrus|VH3-72_IGHD7-27*01_IGHJ2*01 702 252 gnl|Fabrus|VH3-73_IGHD1-1*01_IGHJ5*01 703 253 gnl|Fabrus|VH3-73_IGHD2-8*01_IGHJ2*01 704 254 gnl|Fabrus|VH3-73_IGHD3-22*01_IGHJ4*01 705 255 gnl|Fabrus|VH3-73_IGHD3-9*01_IGHJ6*01 706 256 gnl|Fabrus|VH3-73_IGHD4-11*01_IGHJ6*01 707 257 gnl|Fabrus|VH3-73_IGHD4-23*01_IGHJ5*01 708 258 gnl|Fabrus|VH3-73_IGHD5-12*01_IGHJ4*01 709 259 gnl|Fabrus|VH3-73_IGHD6-19*01_IGHJ1*01 710 260 gnl|Fabrus|VH3-73_IGHD7-27*01_IGHJ5*01 711 261 gnl|Fabrus|VH3-74_IGHD1-1*01_IGHJ6*01 712 262 gnl|Fabrus|VH3-74_IGHD1-26*01_IGHJ4*01 713 263 gnl|Fabrus|VH3-74_IGHD2-2*01_IGHJ5*01 714 264 gnl|Fabrus|VH3-74_IGHD3-22*01_IGHJ5*01 715 265 gnl|Fabrus|VH3-74_IGHD4-17*01_IGHJ1*01 716 266 gnl|Fabrus|VH3-74_IGHD5-12*01_IGHJ4*01 717 267 gnl|Fabrus|VH3-74_IGHD6-6*01_IGHJ1*01 718 268 gnl|Fabrus|VH3-74_IGHD7-27*01_IGHJ4*01 719 269 gnl|Fabrus|VH3-9_IGHD1-1*01_IGHJ6*01 720 270 gnl|Fabrus|VH3-9_IGHD1-7*01_IGHJ5*01 721 271 gnl|Fabrus|VH3-9_IGHD2-2*01_IGHJ4*01 722 272 gnl|Fabrus|VH3-9_IGHD3-16*01_IGHJ6*01 723 273 gnl|Fabrus|VH3-9_IGHD3-22*01_IGHJ4*01 724 274 gnl|Fabrus|VH3-9_IGHD4-11*01_IGHJ4*01 725 275 gnl|Fabrus|VH3-9_IGHD5-24*01_IGHJ1*01 726 276 gnl|Fabrus|VH3-9_IGHD6-13*01_IGHJ4*01 727 277 gnl|Fabrus|VH3-9_IGHD6-25*01_IGHJ6*01 728 278 gnl|Fabrus|VH3-9_IGHD7-27*01_IGHJ2*01 729 279 gnl|Fabrus|VH4-28_IGHD1-20*01_IGHJ1*01 730 280 gnl|Fabrus|VH4-28_IGHD1-7*01_IGHJ6*01 731 281 gnl|Fabrus|VH4-28_IGHD2-15*01_IGHJ6*01 732 282 gnl|Fabrus|VH4-28_IGHD3-16*01_IGHJ2*01 733 283 gnl|Fabrus|VH4-28_IGHD3-9*01_IGHJ6*01 734 284 gnl|Fabrus|VH4-28_IGHD4-4*01_IGHJ4*01 735 285 gnl|Fabrus|VH4-28_IGHD5-5*01_IGHJ1*01 736 286 gnl|Fabrus|VH4-28_IGHD6-13*01_IGHJ4*01 737 287 gnl|Fabrus|VH4-28_IGHD7-27*01_IGHJ1*01 738 288 gnl|Fabrus|VH4-31_IGHD1-26*01_IGHJ2*01 739 289 gnl|Fabrus|VH4-31_IGHD2-15*01_IGHJ2*01 740 290 gnl|Fabrus|VH4-31_IGHD2-2*01_IGHJ6*01 741 291 gnl|Fabrus|VH4-31_IGHD3-10*01_IGHJ4*01 742 292 gnl|Fabrus|VH4-31_IGHD3-9*01_IGHJ6*01 743 293 gnl|Fabrus|VH4-31_IGHD4-17*01_IGHJ5*01 744 294 gnl|Fabrus|VH4-31_IGHD5-12*01_IGHJ4*01 745 295 gnl|Fabrus|VH4-31_IGHD6-13*01_IGHJ4*01 746 296 gnl|Fabrus|VH4-31_IGHD6-6*01_IGHJ1*01 747 297 gnl|Fabrus|VH4-31_IGHD7-27*01_IGHJ5*01 748 298 gnl|Fabrus|VH4-34_IGHD1-7*01_IGHJ4*01 749 299 gnl|Fabrus|VH4-34_IGHD2-2*01_IGHJ4*01 750 300 gnl|Fabrus|VH4-34_IGHD3-16*01_IGHJ4*01 751 301 gnl|Fabrus|VH4-34_IGHD3-22*01_IGHJ6*01 752 302 gnl|Fabrus|VH4-34_IGHD4-17*01_IGHJ4*01 753 303 gnl|Fabrus|VH4-34_IGHD5-12*01_IGHJ4*01 754 304 gnl|Fabrus|VH4-34_IGHD6-13*01_IGHJ4*01 755 305 gnl|Fabrus|VH4-34_IGHD6-25*01_IGHJ6*01 756 306 gnl|Fabrus|VH4-34_IGHD6-6*01_IGHJ6*01 757 307 gnl|Fabrus|VH4-34_IGHD7-27*01_IGHJ4*01 758 308 gnl|Fabrus|VH4-39_IGHD1-14*01_IGHJ1*01 759 309 gnl|Fabrus|VH4-39_IGHD1-20*01_IGHJ6*01 760 310 gnl|Fabrus|VH4-39_IGHD2-21*01_IGHJ3*01 761 311 gnl|Fabrus|VH4-39_IGHD3-10*01_IGHJ4*01 762 312 gnl|Fabrus|VH4-39_IGHD3-16*01_IGHJ2*01 763 313 gnl|Fabrus|VH4-39_IGHD3-9*01_IGHJ6*01 764 314 gnl|Fabrus|VH4-39_IGHD4-23*01_IGHJ2*01 765 315 gnl|Fabrus|VH4-39_IGHD5-12*01_IGHJ4*01 766 316 gnl|Fabrus|VH4-39_IGHD6-6*01_IGHJ1*01 767 317 gnl|Fabrus|VH4-4_IGHD1-20*01_IGHJ3*01 768 318 gnl|Fabrus|VH4-4_IGHD2-8*01_IGHJ4*01 769 319 gnl|Fabrus|VH4-4_IGHD3-22*01_IGHJ2*01 770 320 gnl|Fabrus|VH4-4_IGHD4-23*01_IGHJ4*01 771 321 gnl|Fabrus|VH4-4_IGHD5-12*01_IGHJ5*01 772 322 gnl|Fabrus|VH4-4_IGHD6-6*01_IGHJ4*01 773 323 gnl|Fabrus|VH4-4_IGHD7-27*01_IGHJ6*01 774 324 gnl|Fabrus|VH4-59_IGHD6-25*01_IGHJ3*01 775 325 gnl|Fabrus|VH5-51_IGHD1-14*01_IGHJ4*01 776 326 gnl|Fabrus|VH5-51_IGHD1-26*01_IGHJ6*01 777 327 gnl|Fabrus|VH5-51_IGHD2-8*01_IGHJ4*01 778 328 gnl|Fabrus|VH5-51_IGHD3-10*01_IGHJ6*01 779 329 gnl|Fabrus|VH5-51_IGHD3-3*01_IGHJ4*01 780 330 gnl|Fabrus|VH5-51_IGHD4-17*01_IGHJ4*01 781 331 gnl|Fabrus|VH5-51_IGHD5-18*01 > 3_IGHJ4*01 782 332 gnl|Fabrus|VH5-51_IGHD5-18*01 > 1_IGHJ4*01 783 333 gnl|Fabrus|VH5-51_IGHD6-25*01_IGHJ4*01 784 334 gnl|Fabrus|VH5-51_IGHD7-27*01_IGHJ4*01 785 335 gnl|Fabrus|VH6-1_IGHD1-1*01_IGHJ4*01 786 336 gnl|Fabrus|VH6-1_IGHD1-20*01_IGHJ6*01 787 337 gnl|Fabrus|VH6-1_IGHD2-15*01_IGHJ4*01 788 338 gnl|Fabrus|VH6-1_IGHD2-21*01_IGHJ6*01 789 339 gnl|Fabrus|VH6-1_IGHD3-16*01_IGHJ5*01 790 340 gnl|Fabrus|VH6-1_IGHD3-3*01_IGHJ4*01 791 341 gnl|Fabrus|VH6-1_IGHD4-11*01_IGHJ6*01 792 342 gnl|Fabrus|VH6-1_IGHD4-23*01_IGHJ4*01 793 343 gnl|Fabrus|VH6-1_IGHD5-5*01_IGHJ4*01 794 344 gnl|Fabrus|VH6-1_IGHD6-13*01_IGHJ4*01 795 345 gnl|Fabrus|VH6-1_IGHD6-25*01_IGHJ6*01 796 346 gnl|Fabrus|VH6-1_IGHD7-27*01_IGHJ4*01 797 347 gnl|Fabrus|VH7-81_IGHD1-14*01_IGHJ4*01 798 348 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ2*01 799 349 gnl|Fabrus|VH7-81_IGHD2-21*01_IGHJ6*01 800 350 gnl|Fabrus|VH7-81_IGHD3-16*01_IGHJ6*01 801 351 gnl|Fabrus|VH7-81_IGHD4-23*01_IGHJ1*01 802 352 gnl|Fabrus|VH7-81_IGHD5-12*01_IGHJ6*01 803 353 gnl|Fabrus|VH7-81_IGHD6-25*01_IGHJ4*01 804 354 gnl|Fabrus|VH7-81_IGHD7-27*01_IGHJ4*01 805 Light Chain SEQ Number Name ID NO. 1 gnl|Fabrus|A14_IGKJ1*01 806 2 gnl|Fabrus|A17_IGKJ1*01 807 3 gnl|Fabrus|A2_IGKJ1*01 808 4 gnl|Fabrus|A20_IGKJ1*01 809 5 gnl|Fabrus|A23_IGKJ1*01 810 6 gnl|Fabrus|A26_IGKJ1*01 811 7 gnl|Fabrus|A27_IGKJ1*01 812 8 gnl|Fabrus|A27_IGKJ3*01 813 9 gnl|Fabrus|A30_IGKJ1*01 814 10 gnl|Fabrus|B2_IGKJ1*01 815 11 gnl|Fabrus|B2_IGKJ3*01 816 12 gnl|Fabrus|B3_IGKJ1*01 817 14 gnl|Fabrus|L11_IGKJ1*01 819 15 gnl|Fabrus|L12_IGKJ1*01 820 16 gnl|Fabrus|L14_IGKJ1*01 821 17 gnl|Fabrus|L2_IGKJ1*01 822 18 gnl|Fabrus|L22_IGKJ3*01 823 19 gnl|Fabrus|L23_IGKJ1*01 824 20 gnl|Fabrus|L25_IGKJ1*01 825 21 gnl|Fabrus|L25_IGKJ3*01 826 22 gnl|Fabrus|L4/18a_IGKJ1*01 827 23 gnl|Fabrus|L5_IGKJ1*01 828 24 gnl|Fabrus|L6_IGKJ1*01 829 25 gnl|Fabrus|L8_IGKJ1*01 830 26 gnl|Fabrus|L9_IGKJ2*01 831 27 gnl|Fabrus|O1_IGKJ1*01 832 28 gnl|Fabrus|O12_IGKJ1*01 833 29 gnl|Fabrus|O18_IGKJ1*01 834 31 gnl|Fabrus|V1-11_IGLJ2*01 836 32 gnl|Fabrus|V1-13_IGLJ5*01 837 33 gnl|Fabrus|V1-16_IGLJ6*01 838 34 gnl|Fabrus|V1-18_IGLJ2*01 839 35 gnl|Fabrus|V1-2_IGLJ7*01 840 36 gnl|Fabrus|V1-20_IGLJ6*01 841 37 gnl|Fabrus|V1-3_IGLJ1*01 842 38 gnl|Fabrus|V1-4_IGLJ4*01 843 39 gnl|Fabrus|V1-5_IGLJ2*01 844 40 gnl|Fabrus|V1-7_IGLJ1*01 845 41 gnl|Fabrus|V1-9_IGLJ6*01 846 42 gnl|Fabrus|V2-1_IGLJ6*01 847 43 gnl|Fabrus|V2-11_IGLJ7*01 848 44 gnl|Fabrus|V2-13_IGLJ2*01 849 45 gnl|Fabrus|V2-14_IGLJ4*01 850 46 gnl|Fabrus|V2-15_IGLJ7*01 851 47 gnl|Fabrus|V2-17_IGLJ2*01 852 48 gnl|Fabrus|V2-19_IGLJ4*01 853 49 gnl|Fabrus|V2-6_IGLJ4*01 854 50 gnl|Fabrus|V2-7_IGLJ2*01 855 51 gnl|Fabrus|V2-7_IGLJ7*01 856 52 gnl|Fabrus|V2-8_IGLJ6*01 857 53 gnl|Fabrus|V3-2_IGLJ4*01 858 54 gnl|Fabrus|V3-3_IGLJ7*01 859 55 gnl|Fabrus|V3-4_IGLJ1*01 860 56 gnl|Fabrus|V4-1_IGLJ4*01 861 57 gnl|Fabrus|V4-2_IGLJ4*01 862 58 gnl|Fabrus|V4-3_IGLJ4*01 863 59 gnl|Fabrus|V4-4_IGLJ5*01 864 60 gnl|Fabrus|V4-6_IGLJ4*01 865 61 gnl|Fabrus|V5-4_IGLJ2*01 866 62 gnl|Fabrus|V5-6_IGLJ1*01 867
Example 7
Comparison of Fab Secretion by Different Leader Sequences
[1131] In this Example, the effect of different leader sequences on expression of encoded Fabs was assessed. In order to assure properly folded Fab proteins, disulfide bonds must form in an oxidizing environment and therefore it is necessary to translocate the Fab proteins into the periplasm by using a pathway such as Sec, SRP and TAT. The SRP pathway does not require unfolding of any proteins. SRP leader sequences are shown in Table 23.
TABLE-US-00039 TABLE 23 SRP Leader Sequences Name SEQ ID NO. Sequence DsbA 5 MKKIWLALAGLVLAFSASA SfmC 6 MMTKIKLLMLIIFYLIISASAHA TolB 7 MKQALRVAFGFLILWASVLHA TorT 8 MRVLLFLLLSLRMLPAFS
[1132] Plasmid A (SEQ ID NO:1) and plasmid C (SEQ ID NO:3) were modified to contain either a DsbA leader sequence (SEQ ID NO:5) or a mutant DsbA leader sequence (SEQ ID NO:965) and a ribosomal binding site (RBS) variation. Overlap PCR was performed using three forward primers containing RBS variations and the sequence corresponding to the N-terminus of DsbA (set forth in SEQ ID NOS:966-968) and two reverse primers corresponding to the C-terminus of either DsbA or mutant DsbA (set forth in SEQ ID NOS:969-970). The PCR resulted in six different leader sequences (Table 24). These leader sequences were then inserted into Plasmid A and Plasmid C between EcoRI and NcoI sites upstream of the start codon, ATG. Plasmids A and C containing the DsbA leader sequence with the RBS variations are set forth in SEQ ID NO:1015-1017 and 1021-1023, respectively. Plasmids A and C containing the mutant DsbA leader sequence with the RBS variations are set forth in SEQ ID NO:1018-1020 and 1024-1026, respectively.
TABLE-US-00040 TABLE 24 DsbA and mutant DsbA leader sequences with RBS variations SEQ Name Leader Sequence ID NO. DS1 GAATTCTAAGGAGGTTTCACCATGAAAAAGA 971 TTTGGCTGGCGCTGGCTGGTTTAGTTTTAGC GTTTAGC GCC ATG G D52 GAATTCTAAGGAGTATTCACCATGAAAAAGA 972 TTTGGCTGGCGCTGGCTGGTTTAGTTTTAGC GTTTAGC GCC ATG G D53 GAATTCTTAGGATTATTCACCATGAAAAAGAT 973 TTGGCTGGCGCTGGCTGGTTTAGTTTTAGCG TTTAGC GCC ATG G DM1 GAATTCTAAGGAGGTTTCACCATGAAAAAGA 974 TTTGGCTGGCGCTGGCTGGTTTAGTTTTAGC GTTTAGCGCATCG GCG GCC ATG GCA DM2 GAATTCTAAGGAGTATTCACCATGAAAAAGA 975 TTTGGCTGGCGCTGGCTGGTTTAGTTTTAGC GTTTAGCGCATCG GCG GCC ATG GCA DM3 GAATTCTTAGGATTATTCACCATGAAAAAGAT 976 TTGGCTGGCGCTGGCTGGTTTAGTTTTAGCG TTTAGCGCATCG GCG GCC ATG GCA
[1133] To assess the effect of the different leader sequences on the expression of a Fab antibody, expression of Rituxan was assessed. Rituxan heavy chain (SEQ ID NO:453) was cloned in-frame to the C.sub.H sequence present on the respective Plasmid A vector and Rituxan light chain (SEQ ID NO:835) was cloned in-frame to the C.kappa. sequence present in the respective Plasmid C vector. Briefly, the V.sub.H and V.kappa. chains were cloned into the Plasmid A or C vectors (containing an STII leader sequence, a DsbA leader sequence or a mutant DsbA leader sequence) by digestion of the plasmids with NheI and NcoI (for the heavy chain DNA) or with NcoI and BsiWI (for the light chain DNA) followed by ligation. Plasmids encoding a heavy and light chain were co-transformed into LMG194 cells as described in Example 8 below and grown for 36 hours at 20.degree. C. in terrific broth (TB) medium in the presence of 0.4% glucose and 0.008% arabinose. Expressed Fabs were extracted from the whole cells using Bugbuster.RTM. (Novagen), and purified using a Ni.sup.2+ affinity column (EMD), and analyzed by SDS-PAGE followed by Western Blot. The results show that in the absence of an inducer little to no protein was expressed. Upon induction, expression of the Fabs under the control of the DsbA leader sequence (for each of the three RBS variations) was equal to expression under the control of the STII leader sequence. Mutated DsbA resulted in lower expression. There was no significant difference in expression among the various RBS variations of the DsbA leader sequence.
Example 8
Cloning and Co-Transformation of Synthesized Variable Heavy and Light Chains
[1134] In this Example, a Fab library was created by cloning heavy or light chain DNA into their respective Plasmids followed by co-transformation and protein growth/purification. Following synthesis of the DNA molecules generated from the Sequence Compilation Software as described in Example 6, the DNA molecules were cloned into plasmids containing constant heavy or light chains as appropriate for co-transformation and expression of combinatorial Fabs. Plasmid A (SEQ ID NO:1) and plasmid D (SEQ ID NO:2) contain heavy chain constant regions sequences. Plasmid C (SEQ ID NO:3) contains a kappa light chain constant region sequence and Plasmid E (SEQ ID NO:4) contains a lambda light chain constant region sequence.
[1135] Synthetic recombined nucleic acid encoding a variable heavy chain were digested with Nhe I and Nco I and ligated into Plasmid A with a StII leader sequence using standard molecular techniques. Synthetic recombined nucleic acid encoding a variable kappa light chain were digested with NcoI and BsiWI and synthetic recombined nucleic acid encoding a variable lambda chain were digested with NcoI and AvrII, and were ligated into Plasmid C or Plasmid E, respectively, with a StII leader sequence, using standard molecular biology techniques.
[1136] Plasmid A and one of either Plasmid C or Plasmid E, each containing various combinations of variable heavy and light chains, were co-transformed into E. coli. The process was repeated for all combinations of heavy and light chains. Briefly, plasmid A (encoding a Fab heavy chain) and plasmid C or Plasmid E (encoding a Fab light chain) were resuspended separately in TE buffer to a final concentration of 1 ng/.mu.l. One (1) .mu.L of heavy chain plasmid and 1 .mu.L of light chain plasmid were combined in a PCR tube or a PCR plate and were mixed with 20 .mu.L ice cold LMG194 competent cells. The transformation reaction was incubated on ice for 10 minutes followed by heat shock in a preheated PCR block at 42.degree. C. for 45 seconds. The tube was then placed on ice for an additional 2 minutes followed by addition of 200 .mu.L SOC medium. The cells were allowed to recover for 1.5 hours at 37.degree. C. A 100 .mu.L aliquot of the transformation culture was used to inoculate 0.9 mL LB (Luria-Bertani Broth) containing 0.4% (w/v) glucose, 17 .mu.g/mL kanamycin (Sigma Aldrich) and 34 .mu.g/mL chloramphenicol (Sigma Aldrich). The culture was grown at 30.degree. C. with vigorous shaking for 20 hours. The transformation culture was grown and purified using the Piccolo.TM. system as described in Example 9.
Example 9
High Throughput Growth and Purification of Fab Libraries
[1137] In this Example, Fab libraries are generated and purified using high throughput techniques. High throughput transformation of pairs of heavy chains (plasmid A) and light chains (plasmid C or E) was performed as described in Example 8, except that a 96-well PCR plate was used instead of individual PCR tubes. After transformation, the cells were grown overnight in 2 ml deep well 96-well plates (VWR) block covered with breathable tape. The overnight culture was used directly for inoculation in Piccolo.TM. (Wollerton et al. (2006) JALA, 11:291-303.)
[1138] High throughput, parallel expression and purification of antibody Fab fragments was performed using Piccolo.TM. (The Automation Partnership (TAP)), which automates protein expression and purification. The expression and purification parameters for Piccolo.TM. were prepared using Run Composer software (TAP). A `Strain File` was generated mapping the location of each clone in the seed culture plate. This was submitted to the Run Composer software and the basic machine settings were set as follows: Pre-induction Incubator set at 30.degree. C.; Expression Incubator 1 set at 16.degree. C.; Centrifuge set at 6.degree. C. and 5000.times.g; Media Pump 1 primed with TB (Terrific Broth; per liter contains 12 g tryptone, 24 g yeast extract, 9.4 g potassium phosphate, dibasic, and 2.2 g potassium phosphate, monobasic) (EMD Biosciences; catalog No. 71754), 50 .mu.g/mL kanamycin (Sigma Aldrich), 35 .mu.g/mL chloramphenicol (Sigma Aldrich), 0.4% (w/v) glucose (Sigma Aldrich) and 0.015% (v/v) Antifoam 204 (Sigma Aldrich); Inducer Pump 1 primed with 0.2% (w/v) arabinose (EMD Biosciences); Incubator Gassing Rate set at 2 sec with 51% oxygen, 0.1 mL inoculation volume; Induction Statistic Mean set w/o Outliers (i.e. block mean 0D600 determined after excluding the 3 highest and 3 lowest values); culture vessel blocks (CVB) pre-induction delay set at 1 hr 20 min and Expression Incubator Acclimatization set at 30 min.
[1139] The seed cultures were prepared and loaded into Piccolo.TM. along with the necessary labware: 24-well culture vessel blocks (CVBs; The Automation Partnership), 24-well Filter Plates (The Automation Partnership), 24-well Output Plates (Seahorse Bioscience) and Pipette Tip Boxes (MBP) as specified by the manufacturer. The TB media supplemented as described above, arabinose inducer and associated pumps were prepared under sterile conditions and attached to the machine. The centrifuge counterbalance weight was set and placed inside the centrifuge. Lastly, purification reagents were prepared and attached to the system pumps (lysis buffer, resin, wash buffer and elution buffer as described below). Once this was complete, the machine was started and processing began.
[1140] Before inoculation, the inocula were mapped to specific wells of 24-well CVB, and expression and induction conditions were set as described in Table 25 below. Each well of the CVBs were filled with 10 mL of TB media supplemented as described above prior to inoculation from the seed plate. Each well of each CVB was inoculated with 0.1 mL seed culture and then returned to the storage carousel to await scheduled admission to pre-induction incubation. Once a CVB was queued to begin pre-induction incubation it was removed from the storage carousel and coupled to an aeration assembly (which provides agitation, well sealing and a means for controlled administration of oxygen/air) and then placed in the pre-induction incubator. 0D600 readings were taken upon commencement of incubation and approximately every 30 minutes thereafter. Piccolo operation control software monitors the 0D600 measurements to predict when each CVB will reach the 1.0 OD600 set point. Approximately 30 minutes prior to the CVB reaching the OD.sub.600 set point the assembly was moved to the expression incubator to equilibrate to the desired expression temperature, and then the cultures in the CVB were induced by addition of the predetermined volume of arabinose as set forth in Table 25. Cell growth was monitored by measuring the OD.sub.600 every 30 minutes after induction and plotting the data showing total incubation time versus optical density reading. The 6 CVB expressions/purifications schemes from Table 25 were analyzed. CVB01 through CVB04 exhibited the same growth patterns with the 0D600 maxing out around 20 after approximately 1700 minutes. In these plates, growth of the Fab cultures in each of the individual wells of the plates were similar, with a somewhat greater variability in growth rates per well observed in the CVB03 and CVB04 conditions. CVB05 and CVB06 showed slower growth, with the 0D600 still increasing at the last time point tested, i.e. total incubation time of 2300 minutes after induction. The growth of the Fab cultures varied between wells of each plate; the maximum OD.sub.600 between wells varied between OD.sub.600=10 to OD.sub.600=16-18.
TABLE-US-00041 TABLE 25 Growth Curves of CVB plates Under Various Conditions Post- Culture Inocula Wells establishment Pre- Vessel of 96-well chill Induction Induction Expression Induction Block plate Incubation Incubation Time Conditions Conditions CVB01 A1, 3, 5, 7, 9, 11 41 min 7 hr 56 min 509 min 30 hours 0.008% (w/v) B1, 3, 5, 7, 9, 11 47 s @ 30.degree. C. @ 16.degree. C. arabinose at C1, 3, 5, 7, 9, 11 OD = 2 D1, 3, 5, 7, 9, 11 CVB02 A1, 3, 5, 7, 9, 11 4 hr 26 min 7 hr 40 min 492 min 36 hours 0.008% (w/v) B1, 3, 5, 7, 9, 11 25 s @ 30.degree. C. @ 16.degree. C. arabinose at C1, 3, 5, 7, 9, 11 OD = 2 D1, 3, 5, 7, 9, 11 CVB03 E1, 3, 5, 7, 9, 11 1 hr 45 min 8 hr 11 min 525 min 30 hours 0.008% (w/v) F1, 3, 5, 7, 9, 11 40 s @ 30.degree. C. @ 16.degree. C. arabinose at G1, 3, 5, 7, 9, 11 OD = 2 H1, 3, 5, 7, 9, 11 CVB04 E1, 3, 5, 7, 9, 11 5 hr 31 min 8 hr 12 min 525 min 36 hours 0.008% (w/v) F1, 3, 5, 7, 9, 11 34 s @ 30.degree. C. @ 16.degree. C. arabinose at G1, 3, 5, 7, 9, 11 OD = 2 H1, 3, 5, 7, 9, 11 CVB05 E2, 4, 6, 8, 10, 12 3 hr 11 min 8 hr 18 min 532 min 30 hours 0.008% (w/v) F2, 4, 6, 8, 10, 12 23 s @ 30.degree. C. @ 16.degree. C. arabinose at G2, 4, 6, 8, 10, 12 OD = 2 H2, 4, 6, 8, 10, 12 CVB06 E2, 4, 6, 8, 10, 12 6 hr 46 min 8 hr 35 min 549 min 36 hours 0.008% (w/v) F2, 4, 6, 8, 10, 12 46 s @ 30.degree. C. @ 16.degree. C. arabinose at G2, 4, 6, 8, 10, 12 OD = 2 H2, 4, 6, 8, 10, 12
[1141] Following culture inoculation and growth induction of cultures, the cells were harvested and lysed for purification of Fabs. Piccolo.TM. was used for purification of the expressed Fab proteins using an automated expression and purification `lifecycle` of a whole culture purification. After controlled expression, CVBs were chilled for 30 minutes at 6.degree. C. in the storage carousel prior to lysis. The CVB was moved to the liquid handling bed and lysis buffer (2.5 mL of Popculture with 1:1000 Lysonase (EMD Biosciences)) was added to each well with thorough mixing. The lysis proceeded for 10 minutes and then the CVB was centrifuged for 10 minutes at 5000.times.g to pellet cell debris. During centrifugation, a Filter Plate was placed in the filter bed and resin (2 mL of a 50% slurry of Ni-charged His-Bind resin (EMD Biosciences)) was added to each well. Soluble lysate was added to the corresponding wells of the filter plate containing resin and allowed to bind for 10 minutes prior to draining to waste. Wash buffer (12 mL of wash buffer (50 mM Sodium Phosphate, 300 mM NaCl, 30 mM Imidazole, pH 8.0)) was added in two steps to each well and allowed to drain to waste. Finally, an Output Plate was placed under the Filter Plate in the filter bed and IMAC elution buffer ((50 mM Sodium Phosphate, 300 mM NaCl, 500 mM Imidazole)) was added in two steps draining into the output plate. The output plate was returned to the storage carousel as was all other labware. Once this process was complete for each CVB in the designed run, the machine was unloaded.
[1142] Using similar Piccolo.TM. runs with the same clones allowed for the optimization of incubation temperatures for expression, concentration of arabinose inducer and time of expression. Global optimal results were obtained by use of a two-step temperature incubation where the pre-induction samples are incubated at 30.degree. C. Following induction, protein expression was carried out at 20.degree. C. Optimal expression yields were obtained using 0.032% arabinose inducer followed by 45 hours of expression.
Example 10
Orthogonal Secondary Purification of Fab Antibodies
[1143] To rapidly further purify partially pure Fabs generated after the Piccolo.TM. process, an orthogonal method of purification was developed. Fabs were expressed and purified as described above in Example 9 using the Piccolo.TM. machine. Approximately 1.8 mL of the IMAC elution per Fab sample obtained from Piccolo.TM. purification was further purified on a 1 mL Hi-Trap Protein G column (GE Healthcare) at 4.degree. C. using the Akta purifier (GE Healthcare) and A-905 autosampler (GE Healthcare) according to the manufacturer's protocol. The protein samples were transferred to a deep well 96-well block (VWR), which was covered by aluminum foil tape to prevent evaporation. The autosampler was set for multiple injections (typically four injections of 450 .mu.L per sample) onto the Hi-Trap Protein G column. The column was then washed with 2 column volumes of 50 mM sodium phosphate pH7.2, 150 mM NaCl. The Fab was eluted with six column volumes of 100 mM glycine pH2.8. The elution peak fractions (approximately 0.8 mL) were collected in a deep well 96-well plate block. The eluted protein was immediately neutralized with 100 .mu.L saturated dibasic sodium phosphate pH9.0. Protein concentration was determined by measuring absorbance at A280 on a Molecular Dynamic plate reader and calculated from the extinction coefficient of the corresponding Fab. Extinction coefficients are calculated based on the total numbers of Tyrosine+Tryptophane+Phenylalanine in the Fab heavy and light chains. Following purification using the Piccolo.TM. system, expressed protein was generally less than 20% pure. After orthogonal purification with protein G, Fab purity was greater than 95% pure as indicated by SDS-PAGE.
[1144] Table 17 sets forth Fab antibodies generated by Piccolo and further purified in a secondary purification. The amino acid sequences of the Fab antibodies correspond to sequences containing variable heavy chain sequences set forth in any of SEQ ID NOS: 1475-1826 and variable light chain sequences set forth in SEQ ID NO: 1827-1838, 1840-1855 and 1857-188. The sequences of the heavy and light chain Fab library members also further contains a sequence for a constant region included in Plasmid A, C or E.
Example 11
Lymphoma Apoptosis Assay
[1145] To identify unique Fabs from an antibody library, assays can be performed to assess any desired function, property or activity. As an example of an assay for protein function, cross-linked anti-CD20 Fab (heavy chain set forth in SEQ ID NO:453 and light chain set forth in SEQ ID NO:835) was tested in a cell based assay for the ability to induce apoptosis in lymphoma cells. Anti-CD20 Fabs were cross-linked by adding equal molar concentration of polyclonal anti human kappa light chain antibodies that recognize different parts of the anti-CD20 kappa light chain.
[1146] Apoptosis was determined using the Apo-ONE homogenous Caspase-3/7 Assay (Promega). Ramos B-Lymphocyte cells were grown in RPM11640 media containing 10% FBS (fetal bovine serum), P-S (penicillin-streptomycin) and arrayed into a clear 96-well plate (Costar 3595, Corning, 2.5.times.10.sup.4 cells per well). Jurkat T-Lymphocyte cells were grown in RPM11640 with 10% FBS and P-S, and arrayed into a clear 96-well plate (Costar 3595, Corning, 2.5.times.10.sup.4 cells per well). Crosslinked anti-CD20 Fab was added to each well at concentrations of 0, 25, 50, 100 and 200 nM together with equal molar concentrations of polyclonal antibody against human kappa light chain (Sigma). Anti-Fas monoclonal antibody (20-100 ng/ml) or staurosporine (1.0 .mu.M) were added for positive controls and trastuzumab Fab was used as a negative control. An equal volume of Apo-ONE Caspase-3/7 Reagent (rhodamine 110, bis-(N-CBZ-L-aspartyl-L-glutamyl-L-valyl-L-aspartic acid amide, Z-DEVD-R110) was added and the plate was incubated for one hour at room temperature. The presence of rhodamine 110 was detected by measuring fluorescence in a fluorescent plate reader at 485 nm excitation/521 nm emission. The increase in the percentage of apoptosis was calculated according to the equation: 100.times.[(Fluorescence for Fab)-(Fluorescence for background)]/(Fluorescence for background).
Example 12
Erythropoietin Binding Fab Library
[1147] As an alternative to "naive" libraries, directed Fab libraries are constructed to a known target. In this example, a directed library was constructed in which a 16 amino acid erythropoietin peptide (EPO) was inserted into various CDRs of an antibody to identify Fabs that induce activation of recombinant EPO (EpoR). Because there are variations in the number of amino acid residues that occur in a CDR (see, e.g., Table 26), CDRs with larger inserts were selected for insertion of the EPO peptide. These CDRs include CDR-H2, CDR-H3, CDR-L1, and CDR-L3.
TABLE-US-00042 TABLE 26 Amino Acid Length of CDR Heavy Chain Light Chain CDR1 10-12 10-17 CDR2 16-19 7 CDR3 3-25 7-11
[1148] To randomize the orientation of the peptide insert to expose the active surface of the peptide, two extra amino acid residues, either proline or glycine, were added to the N-terminus and C-terminus of the EPO peptide, yielding sixteen different peptides (SEQ ID NO:874-889). A Fab, containing heavy chain VH3-23 (SEQ ID NO:869) and light chain A17 (SEQ ID NO:871), served as a parent antibody for the EPO peptide library. BsaI restriction sites were introduced into the nucleic acid sequences of the EPO peptides and the nucleic acid sequences of the heavy and light chain variable regions in order to allow the cloning of the DNA encoding the EPO peptides into the respective CDRs. Heavy chain VH3-23 DNA was modified to create VH3-23B (SEQ ID NO:896) containing BsaI sites at CDR2 and VH3-23R (SEQ ID NO:913) containing BsaI sites at CDR3. Light chain A17 DNA was modified to create A17P (SEQ ID NO:872) containing BsaI sites at CDR1 and A17Q (SEQ ID NO:873) containing BsaI sites at CDR3. Sixteen different genes encoding the EPO peptides were cloned into the respective heavy (CDR2 and CDR3) or light (CDR1 and CDR3) chain sequences and the resulting Fabs were expressed and purified as described in Example 9. Table 27 lists the resulting 64 EPO containing Fabs.
TABLE-US-00043 TABLE 27 EPO Peptide containing Fab Library HEAVY CHAIN LIGHT CHAIN SEQ SEQ NAME ID NO NAME ID NO H2-EP14 (VH3-23B) 897 A17 871 H2-EP18 (VH3-23B) 898 A17 871 H2-EP19 (VH3-23B) 899 A17 871 H2-EP20 (VH3-23B) 900 A17 871 H2-EP21 (VH3-23B) 901 A17 871 H2-EP22 (VH3-23B) 902 A17 871 H2-EP23 (VH3-23B) 903 A17 871 H2-EP24 (VH3-23B) 904 A17 871 H2-EP25 (VH3-23B) 905 A17 871 H2-EP26 (VH3-23B) 906 A17 871 H2-EP27 (VH3-23B) 907 A17 871 H2-EP28 (VH3-23B) 908 A17 871 H2-EP29 (VH3-23B) 909 A17 871 H2-EP30 (VH3-23B) 910 A17 871 H2-EP31 (VH3-23B) 911 A17 871 H2-EP32 (VH3-23B) 912 A17 871 H3-EP14 (VH3-23R) 914 A17 871 H3-EP18 (VH3-23R) 915 A17 871 H3-EP19 (VH3-23R) 916 A17 871 H3-EP20 (VH3-23R) 917 A17 871 H3-EP21 (VH3-23R) 918 A17 871 H3-EP22 (VH3-23R) 919 A17 871 H3-EP23 (VH3-23R) 920 A17 871 H3-EP24 (VH3-23R) 921 A17 871 H3-EP25 (VH3-23R) 922 A17 871 H3-EP26 (VH3-23R) 923 A17 871 H3-EP27 (VH3-23R) 924 A17 871 H3-EP28 (VH3-23R) 925 A17 871 H3-EP29 (VH3-23R) 926 A17 871 H3-EP30 (VH3-23R) 927 A17 871 H3-EP31 (VH3-23R) 928 A17 871 H3-EP32 (VH3-23R) 929 A17 871 VH3-23 869 L1-EP14 (A17P) 930 VH3-23 869 L1-EP18 (A17P) 931 VH3-23 869 L1-EP19 (A17P) 932 VH3-23 869 L1-EP20 (A17P) 933 VH3-23 869 L1-EP21 (A17P) 934 VH3-23 869 L1-EP22 (A17P) 935 VH3-23 869 L1-EP23 (A17P) 936 VH3-23 869 L1-EP24 (A17P) 937 VH3-23 869 L1-EP25 (A17P) 938 VH3-23 869 L1-EP26 (A17P) 939 VH3-23 869 L1-EP27 (A17P) 940 VH3-23 869 L1-EP28 (A17P) 941 VH3-23 869 L1-EP29 (A17P) 942 VH3-23 869 L1-EP30 (A17P) 943 VH3-23 869 L1-EP31 (A17P) 944 VH3-23 869 L1-EP32 (A17P) 945 VH3-23 869 L3-EP14 (A17Q) 946 VH3-23 869 L3-EP18 (A17Q) 947 VH3-23 869 L3-EP19 (A17Q) 948 VH3-23 869 L3-EP20 (A17Q) 949 VH3-23 869 L3-EP21 (A17Q) 950 VH3-23 869 L3-EP22 (A17Q) 951 VH3-23 869 L3-EP23 (A17Q) 952 VH3-23 869 L3-EP24 (A17Q) 953 VH3-23 869 L3-EP25 (A17Q) 954 VH3-23 869 L3-EP26 (A17Q) 955 VH3-23 869 L3-EP27 (A17Q) 956 VH3-23 869 L3-EP28 (A17Q) 957 VH3-23 869 L3-EP29 (A17Q) 958 VH3-23 869 L3-EP30 (A17Q) 959 VH3-23 869 L3-EP31 (A17Q) 960 VH3-23 869 L3-EP32 (A17Q) 961
[1149] Screening to identify Fabs that modulated activation of the Epo Receptor (EpoR, SEQ ID NO:962) was performed in BaF3 cells stably transfected with EpoR-encoding cDNA.
[1150] a. MIT Cell Proliferation Assay
[1151] The tetrazolium ring of MTT (3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyl tetrazolium bromide) is cleaved by mitochondrial dehydrogenase of viable cells to produce purple MTT formazan crystals that are insoluble in aqueous solution but can be solubilized in acidified isopropanol. BaF3/EpoR cells (5.times.10.sup.3-5.times.10.sup.5) were incubated for one or two days in the presence and in the absence of the Epo Fabs. MTT solution (10% of the culture volume) was added to cells and incubated 3-4 hrs at 37.degree. C. in a CO.sub.2 incubator. An equal volume of 0.1 N HCl in isopropanol was added. Absorbance was measured at 570 nm and the number of cells was calculated based on the standard curve.
[1152] b. Luciferase Reporter Assay
[1153] Activation of EpoR leads to activation of STAT5 transcription factor, which in turn induces transcription of genes including c-myc, bcl-2, pim-1 and cyclin-D. Transcription can be detected by linking the promotor of any one of the above genes to a reporter gene, thus creating a reporter plasmid. Two reporter plasmids were created to assess activation of the EpoR. The first reporter plasmid was constructed by placing the mouse cyclin-D promoter (SEQ ID NO:963) at the 5' end of the luciferase gene in the pGL4.70 vector (Promega; SEQ ID NO:1997). Briefly, pGL4.70 was digested with restriction enzymes NheI and HindIII and the cyclin D promoter was inserted between nucleotides 28-66 of the plasmid using standard molecular biology protocols.
[1154] An EPO Fab that is capable of activating EpoR will cause activation of STAT5, thereby activating the cyclin-D promoter causing induction and expression of the luciferase gene. Hence, a second reporter plasmid was created for STAT5 by using the pGL4.23 vector (Promega; SEQ ID NO: 1998), a vector that contains a minimal promoter at the 5' end of the translation start site for luciferase that allows for activation of a promoterless-response element to drive expression. Six repeats of a DNA element for STAT5 binding (SEQ ID NO:964) were cloned directly upstream of the minimal promoter. Briefly, pGL4.23 was digested with restriction enzymes NheI and HindIII and the DNA element repeats of STAT5 was inserted between nucleotides 28-66 of the plasmid using standard molecular biology protocols.
[1155] An EPO Fab capable of activating EpoR will cause the activation of STAT5 and the direct expression of the luciferase gene. BaF3 cells, previously transfected with EpoR-encoding cDNA, were transiently transfected with one of the reporter plasmids above. The resulting cells are incubated in the presence and absence of EPO Fabs. An equal volume of lysis buffer containing luciferase substrate is added to the cell culture. Relative luminescence was measured for 10 seconds using a luminometer after 5 min incubation with the substrate.
Example 13
Electrochemiluminescence Binding Assay
[1156] In this example, an electrochemiluminescence (ECL) binding assay was used to screen a 960 member Fab library for antibodies capable of binding to one of nine different antigens, including the human epidermal growth factor 2 receptor (ErbB2), epidermal growth factor receptor (EGF R), hepatocyte growth factor receptor (HGF R/c-Met), Notch-1, CD44, insulin-like growth factor-1 soluble receptor (IGF-1 sR), P-cadherin, erythropoietin receptor (Epo R) and delta-like protein 4 (DLL4). In an ECL assay, an antigen-antibody interaction is detected by addition of a detection antibody labeled with ruthenium tri-bispyridine-(4-methysulfone) (Ru(bpy).sub.2.sup.2+). Upon application of an electric current, the Ru(bpy).sub.2.sup.2+-label undergoes an oxidation-reduction cycle in the presence of a co-reactant and light is emitted. A signal is only generated when the Ru(bpy).sub.2.sup.2+-label is in close proximity to the electrode, eliminating the need for washing. Detected light intensity is proportional to the amount of captured protein.
[1157] Recombinant human proteins were obtained from R&D Systems and included: rHuman ErbB2/Fc Chimera, CF (Cat #1129-ER); rHuman EGF R/Fc Chimera, CF (Cat #344-ER); rHuman HGF R/c-MET/Fc Chimera, CF (Cat #358-MT/CF); rHuman Notch-1/Fc Chimera, CF (Cat #3647-TK); rHuman CD44/Fc Chimera, CF (Cat #3660-CD); rHuman IGF-1 sR, (IGF-1 sR), CF (Cat #391-GR); rHuman P-Cadherin/Fc Chimera, CF (Cat #861-PC); rHuman Erythropoietin R/Fc Chimera, CF (Cat #963-ER); and Recombinant Human DLL4 (Cat #1506-D4/CF). The proteins were immobilized onto each well of 10 plates by spotting 50 nanoliters (nl) of each protein (of a 60 .mu.g/mL antigen) on the surface of a 96-well Multi-Spot 10 Highbind plate (Meso Scale Discovery; Gaithersburg Md.). Spot 10 was left blank as a control.
[1158] An 150 .mu.l aliquot of 1% Bovine Serum Albumin (BSA) in Tris-buffered Saline Tween (TBST) was added to each well and allowed to incubate for 30 min at 20.degree. C. followed by washing and tap drying to completely remove any residual solution. Subsequently, a 12.5 .mu.l aliquot of 1% BSA TBST was added to each well followed by the addition of a 12.5 .mu.l aliquot of a purified Fab. The plate was sealed and incubated for 1 hour at 20.degree. C. with shaking.
[1159] Detection antibodies were prepared by individually conjugating both goat anti-human Kappa light chain polyclonal antibody (K3502-1MG, Sigma-Aldrich) and goat anti-human Lambda light chain polyclonal antibody (L1645-1ML, Sigma-Aldrich) with Ruthenium (II) tris-bipyridine-(4-methylsulfone)-N-hydroxysuccinimide (SULFO-TAG NHS-ester, Meso Scale Discovery) according to the manufacturer's instructions. TAG-detection antibody at 25 .mu.l was added to each well and allowed to incubate for 1 hour at 20.degree. C. with shaking. Finally, 15 .mu.l of Read Buffer P with Surfactant (Cat # R92PC-1, Meso Scale Discovery) was added to each well. The electrochemiluminescence was measured using a Sector Imager 2400 (Meso Scale Discovery). Data was analyzed by comparing the ECL signals for an antigen to the blank of each well. A signal to blank ratio of 4 or more was considered a "Hit" Fab.
[1160] Ten plates, each containing 96 different Fabs, were screened using the ECL assay. The results of the initial screen are shown in Tables 28-28B, below. Table 28, below, lists the 6 Fabs (including the heavy chain and light chain) that were identified as "hits" in the initial ECL screen. A "hit" was a Fab antibody with signal to blank ratio of greater than 4. Three Fabs were identified that bind to recombinant human delta-like protein 4 (DLL4). One Fab was identified that binds to recombinant human epidermal growth factor 2 (ErbB2) and one Fab was identified that binds to recombinant human erythropoietin receptor (Epo R). An additional Fab was identified that binds to both ErbB2 and EpoR. The results of the initial MSD assay screen at a single Fab concentration are listed in Table 28B below. Table 28B lists the 6 Fabs (the Fab No. corresponds each of the Fabs identified in Table 28), the Fab concentration, the 9 recombinant human target/protein antigens, and the ECL signals from the initial MSD assay screen at the given Fab concentration.
[1161] To confirm a "Hit" from the initial ECL screening, a Fab concentration dependent titration was carried out to determine the Fab-antigen binding affinity. The assay procedure was the same as described above, except that the concentration of Fab antibody was varied between wells from 0.1 nM to 2.4 .mu.M. The data are set forth in Tables 29-34 below. The data were graphed using Microsoft Excel and the binding affinity was estimated from the 50% binding signal. As noted, binding affinity can be assay dependent (see Example 16).
[1162] The results show that Fab VH1-46_IGHD6-6*01_IGHJ1*01 & L6_IGKJ1*01 specifically binds Human DLL4 with high affinity in the lower nanomolar range at or about 10 nM or lower, whereas Fabs VH5-51_IGHD5-18*01>3_IGHJ4*01 & V3-4_IGLJ1*01, and VH6-1_IGHD3-3*01_IGHJ4*01 & V4-3_IGLJ4*01 bind Human DLL4 in the micromolar range. Since the three Fabs contain different heavy and light chains, the results suggest that the binding epitopes on DL44 recognized by the antibodies can be different.
[1163] The results further show that Fab VH4-31_IGHD1-26*01_IGHJ2*01 & A27_IGKJ1*01 binds Human ErbB2/Fc chimera at a concentration of approximately 100 nM and Fab VH1-46_IGHD3-10*01_IGHJ4*01 & B3_IGKJ1*01 binds Human Erythropoietin R/Fc chimera at a concentration of approximately 100 nM. One Fab, VH1-46_IGHD6-13*01_IGH41*01 & B3_IGKJ1*01, shows affinity for both Human ErbB2/Fc and Human Erythropoietin R/Fc chimeras. It is unlikely that this Fab is binding to the Fc region of the chimera proteins, since no binding was observed to five other antigens that were Fc fusion proteins.
TABLE-US-00044 TABLE 28 Identified Fab "Hits" Fab SEQ SEQ No. Target Heavy Chain ID NO Light Chain ID NO 1 rHuman DLL4 VH1-46_IGHD6- 1513 L6_IGKJ1*01 1850 6*01_IGHJ1*01 2 rHuman DLL4 VH5-51_IGHD5- 1803 V3-4_IGLJ1*01 1881 18*01 > 3_IGHJ4*01 3 rHuman DLL4 VH6-1_IGHD3- 1812 V4-3_IGLJ4*01 1884 3*01_IGHJ4*01 4 rHuman ErbB2/Fc chimera VH4-31_IGHD1- 1760 A27_IGKJ1*01 1833 26*01_IGHJ2*01 5 rHuman Epo R/Fc chimera VH1-46_IGHD3- 1509 B3_IGKJ1*01 1838 10*01_IGHJ4*01 6 rHuman ErbB2/Fc chimera VH1-46_IGHD6- 1512 B3_IGKJ1*01 1838 and rHuman Epo R/Fc 13*01_IGHJ4*01 chimera
TABLE-US-00045 TABLE 28B ECL Signals for Identified Fab "Hits" from 960 Fab Library Screen rHuman Target Fab [Fab] Notch- No. uM ErbB2 EGFR HGFR 1 CD44 IGF-1 P-cad EPOR DLL4 Blank 1 0.14 870 823 848 614 515 663 423 693 19293 237 2 2.38 594 681 931 636 666 691 1224 834 12172 526 3 0.21 9229 815 744 949 837 763 890 1115 1069 529 4 0.23 1251 942 954 722 675 982 508 777 2808 363 5 0.74 562 652 633 980 682 593 1155 4108 1000 692 6 0.43 16499 1233 1138 1793 1497 1139 5245 12985 2538 879
TABLE-US-00046 TABLE 29 Binding affinity of Fab VH1-46_IGHD6-6*01_IGHJ1*01 & L6_IGKJ1*01 Fab [nM] 2383 595.8 148.9 37.2 9.3 2.3 0.6 0.1 ErbB2/Fc 454 321 247 384 354 291 215 306 EGF R/Fc 621 403 290 228 424 289 309 311 HGF R/Fc 762 353 205 207 324 253 256 286 Notch-1/Fc 690 306 375 402 492 333 337 378 CD44/Fc 559 372 348 356 396 317 238 323 IGF-1 sR 527 335 322 295 315 231 313 241 P-Cadherin/ 728 617 687 649 452 401 321 235 Fc EPO R/Fc 658 378 373 315 306 429 337 373 DLL4 11794 17203 16253 16717 13210 3055 508 317 Blank 344 285 218 199 287 234 226 201
TABLE-US-00047 TABLE 30 Binding affinity of Fab VH5-51_IGHD5-18*01 > 3_IGHJ4*01 & V3-4_IGLJ1*01 Fab[nM] 154 51 17 6 ErbB2/Fc 1593 1248 1033 873 EGF R/Fc 1398 816 805 742 HGF R/Fc 1520 1044 914 831 Notch-1/Fc 929 685 558 464 CD44/FC 960 651 518 547 IGF-1 sR 1396 1051 872 854 P-Cadherin/Fc 1733 854 542 358 EPO R/Fc 1195 750 620 548 DLL4 40392 17025 7158 1946 Blank 447 335 143 191
TABLE-US-00048 TABLE 31 Binding affinity of Fab VH6-1_IGHD3-3*01_IGHJ4*01 & V4-3_IGLJ4*01 Fab [nM] 480 240 120 60 30 15 7.5 3.8 ErbB2/Fc 965 833 822 777 726 713 695 714 EGF R/Fc 877 690 658 679 585 584 582 511 HGF R/Fc 951 834 785 623 640 694 558 519 Notch-1/Fc 545 368 472 415 425 508 392 383 CD44/Fc 541 470 442 434 484 454 444 419 IGF-1 sR 741 625 813 654 697 705 642 463 P-Cadherin/Fc 596 383 450 372 440 351 352 281 EPO R/Fc 621 478 431 423 325 397 443 407 DLL4 1532 1273 938 875 736 690 598 462 Blank 362 316 363 237 213 261 217 198
TABLE-US-00049 TABLE 32 Binding affinity of Fab VH4-31_IGHD1-26*01_IGHJ2*01 & A27_IGKJ1*01 Fab [nM] 410 205 102.5 51.3 25.6 12.8 6.4 3.2 ErbB2/Fc 5422 5260 4355 3588 2992 2255 1796 868 EGF R/Fc 734 595 455 379 373 320 249 254 HGF R/Fc 753 735 425 456 382 258 234 294 Notch-1/Fc 804 722 607 408 270 249 279 275 CD44/Fc 767 613 461 409 332 273 240 295 IGF-1 sR 600 565 443 316 311 323 209 313 P-Cadherin/Fc 814 769 714 424 323 245 197 206 EPO R/Fc 797 595 587 498 409 338 264 233 DLL4 859 599 550 474 384 268 256 242 Blank 637 430 437 337 345 227 133 172
TABLE-US-00050 TABLE 33 Binding affinity of Fab VH1-46_IGHD3-10*01_IGHJ4*01 & B3_IGKJ1*01 Fab [nM] 1410 705 352.5 176.3 88.1 44.1 22 11 ErbB2/Fc 932 671 514 448 200 347 363 216 EGF R/Fc 1071 692 769 428 376 428 312 201 HGF R/Fc 903 839 606 418 392 336 203 268 Notch-1/Fc 1034 958 715 664 440 331 389 404 CD44/Fc 885 693 556 376 340 302 317 296 IGF-1 sR 426 630 528 393 273 309 347 289 P-Cadherin/Fc 1059 827 649 532 278 343 215 270 EPO R/Fc 4314 4894 4105 3519 3368 2387 2241 1824 DLL4 1265 981 660 460 434 388 342 254 Blank 709 483 494 346 301 200 289 212
TABLE-US-00051 TABLE 34 Binding affinity of Fab VH1-46_IGHD6-13*01_IGHJ4*01 & B3_IGKJ1*01 Fab [nM] 1000 500 250 125 62.5 31.3 15.6 7.8 ErbB2/Fc 8731 10241 11026 12956 13124 13911 14791 13220 EGF R/Fc 2236 1468 1138 860 602 447 346 379 HGF R/Fc 2109 1371 1221 778 578 299 293 282 Notch-1/Fc 2267 1975 1241 802 536 563 418 486 CD44/Fc 1966 1685 1175 764 591 439 473 409 IGF-1 sR 1667 1334 993 654 491 385 349 353 P-Cadherin/ 4495 3447 2784 1481 1173 1105 971 695 Fc EPO R/Fc 8594 10305 8535 9237 7749 7878 8357 6765 DLL4 2785 2319 1560 912 715 528 525 407 Blank 1133 680 590 403 268 250 294 316
Example 14
VH3-23 Heavy Chain Library
[1164] In this example, a library of 690 different VH3-23 containing heavy chains was generated using standard molecular biology protocols. Library diversity was generated by 1) use of 6 different J.sub.H segments; 2) including the DIRECT sequence and INVERTED sequence (or reverse complement) of each D.sub.H segment; and 3) translating each DIRECT and INVERTED D.sub.H segment in all 3 reading frames. This resulted in the initial generation of six pF-VH3-23-IGHJ plasmids and 115 different D.sub.H segments. Simultaneous cloning of each individual D.sub.H segment into each pF-VH3-23-IGHJ plasmid resulted in the 690 member heavy chain library. The library was then transformed with various previously generated light chains (see Example 8) to generate a 3012 member Fab library.
[1165] A. Generation of pF-VH3-23-IGHJ1 to pF-VH3-23-IGHJ6 Plasmids
[1166] In this example, 6 plasmids, each encoding the VH3-23 V.sub.H segment and one of six J.sub.H segments, were generated. The plasmids were modified such that BsaI sites were incorporated to 1) allow the cloning of individual J.sub.H segments (see Table 35); and 2) subsequently allow the cloning of any individual D.sub.H segment (see Table 37). To this end, Plasmid A (SEQ ID NO:1) and V.sub.H segment VH3-23R (SEQ ID NO:2050) were first modified to remove internal BsaI sites (SEQ ID NO:3719). Subsequently, VH3-23R was further modified to add two BsaI sites, including 1) at the 3' end of the VH3-23 V.sub.H segment; and 2) at the 5' end of the nucleotides encoding framework region 4 (amino acids WGQGTLVTVSSAS of SEQ ID NOS:3456-3461, see Table 35 below) of the J.sub.H segment. VH3-23R (SEQ ID NO:2050) was then synthesized using standard DNA synthesis protocols. VH3-23R was digested with NcoI (SEQ ID NO:977) and NheI (SEQ ID NO:978) and ligated into the modified Plasmid A creating plasmid pF-VH3-23R (SEQ ID NO:2051).
[1167] A series of forward and reverse oligos (see Table 36, below) encoding segments IGHJ1-IGHJ6 (see Table 35, and SEQ ID NOS:3450-3455 and SEQ ID NOS:3456-3461) were generated by standard DNA synthesis protocols. Pairs of oligos (encoding a particular IGHJ segment) were digested with BsaI and ligated into similarly digested plasmid pF-VH3-23R (SEQ ID NO:2051), thereby generating 6 new plasmids: pF-VH3-23-IGHJ1 (SEQ ID NO:2064), pF-VH3-23-IGHJ2 (SEQ ID NO:2065), pF-VH3-23-IGHJ3 (SEQ ID NO:2066), pF-VH3-23-IGHJ4 (SEQ ID NO:2067), pF-VH3-23-IGHJ5 (SEQ ID NO:2068), and pF-VH3-23-IGHJ6 (SEQ ID NO:2069). For each of the new plasmids, this resulted in the removal of the BsaI site at the 5' end of the nucleotides encoding framework region 4 and the generation of a new BsaI site at the 5' end of the J.sub.H segment to allow subsequent cloning of a single D.sub.H segment into all six vectors simultaneously thereby creating six different genes per each D.sub.H segment.
TABLE-US-00052 TABLE 35 IGHJ1 to IGHJ6 Segments Segment Nucleotide Sequence IGHJ1 (SEQ ID NO: 3450) GCT GAA TAC TTC CAG CAC TGG GGC CAA GGA ACC CTG GTC ACC GTC TCC TCA G IGHJ2 (SEQ ID NO: 3451) C TAC TGG TAC TTC GAT TCC TGG GGC CAA GGA ACC CTG GTC ACC GTC TCC TCA G IGHJ3 (SEQ ID NO: 3452) T GCT TTT GAT GTC TGG GGC CAA GGA ACC CTG GTC ACC GTC TCC TCA G IGHJ4 (SEQ ID NO: 3453) AC TAC TTT GAC TAC TGG GGC CAA GGA ACC CTG GTC ACC GTC TCC TCA G IGHJ5 (SEQ ID NO: 3454) AC AAC TGG TTC GAC TAC TGG GGC CAA GGA ACC CTG GTC ACC GTC TCC TCA G IGHJ6 (SEQ ID NO: 3455) AT TAC TAC TAC TAC TAC GGT ATG GAC GTC TGG GGC CAA GGA ACC CTG GTC ACC GTC TCC TCA G Segment Protein Sequence IGHJ1 (SEQ ID NO: 3456) AEYFQHWGQGTLVTVSSAS IGHJ2 (SEQ ID NO: 3457) YVVYFDSWGQGTLVTVSSAS IGHJ3 (SEQ ID NO: 3458) AFDWVGQGTLVTVSSAS IGHJ4 (SEQ ID NO: 3459) YFDYVVGQGTLVTVSSAS IGHJ5 (SEQ ID NO: 3460) NWFDYVVGQGTLVTVSSAS IGHJ6 (SEQ ID NO: 3461) YYYYYGMDWVGQGTLVTVSSAS
TABLE-US-00053 TABLE 36 Oligos used to generate J1 to J6 plasmids Oligo Nucleotide Sequence J1_F CGAAAGAGACCTACTATGGTTCGGGTCTCTCC (SEQ ID NO: 2052) TACGCTGAATACTTCCAGCACT J1_R CCCCAGTGCTGGAAGTATTCAGCGTAGGAGAG (SEQ ID NO: 2053) ACCCGAACCATAGTAGGTCTCT J2_F CGAAAGAGACCTACTATGGTTCGGGTCTCTCC (SEQ ID NO: 2054) TACTACTGGTACTTCGATTCCT J2_R CCCCAGGAATCGAAGTACCAGTAGTAGGAGAG (SEQ ID NO: 2055) ACCCGAACCATAGTAGGTCTCT J3_F CGAAAGAGACCTACTATGGTTCGGGTCTCTCC (SEQ ID NO: 2056) TATGCTTTTGATGTCT J3_R CCCCAGACATCAAAAGCATAGGAGAGACCCGA (SEQ ID NO: 2057) ACCATAGTAGGTCTCT J4_F CGAAAGAGACCTACTATGGTTCGGGTCTCTCC (SEQ ID NO: 2058) TACTACTTTGACTACT J4_R CCCCAGTAGTCAAAGTAGTAGGAGAGACCCGA (SEQ ID NO: 2059) ACCATAGTAGGTCTCT J5_F CGAAAGAGACCTACTATGGTTCGGGTCTCTCC (SEQ ID NO: 2060) TACAACTGGTTCGACTACT J5_R CCCCAGGAGTCGAACCAGTTGTAGGAGAGACC (SEQ ID NO: 2061) CGAACCATAGTAGGTCTCT J6_F CGAAAGAGACCTACTATGGTTCGGGTCTCTCC (SEQ ID NO: 2062) TATTACTACTACTACTACGGTATGGACGTCT J6_R CCCCAGACGTCCATACCGTAGTAGTAGTAGTA (SEQ ID NO: 2063) ATAGGAGAGACCCGAACCATAGTAGGTCTCT
[1168] B. Generation and Cloning of Modified D.sub.H Oligos into pF-VH3-23-IGHJ Plasmids
[1169] In this example, pairs of D.sub.H oligos were generated and cloned into the pF-VH3-23-IGHJ plasmids thereby generating 690 new VH3-23 heavy chains as follows. Twenty seven (27) D.sub.H segments (see Table 37 below) were selected for cloning into each modified pF-VH3-23-IGHJ plasmid. Library diversity in the CDR3 region was generated by 1) including the DIRECT sequence and INVERTED sequence (or reverse complement) of each D.sub.H segment; and 2) translating each DIRECT and INVERTED D.sub.H segment in all 3 reading frames. If the translation of a reading frame for any given D.sub.H segment resulted in a stop codon, that particular sequence and reading frame were excluded from the library. Table 37 below indicates the open reading frames for each particular D.sub.H segment that were included in the library, thereby resulting in 115 different D.sub.H segments.
TABLE-US-00054 TABLE 37 Ph segments included in heavy chain library DIRECT INVERTED (SEQ Reading (SEQ Reading Number D segment ID NO) Frames ID NO) Frames 1 IGHD1-1*01 239 1, 2, 3 3462 1, 2, 3 2 IGHD1-7*01 243 1, 3 3463 1, 3 3 IGHD1-14*01 240 1, 3 3464 1, 2, 3 4 IGHD1-20*01 241 1, 3 3465 1, 2, 3 5 IGHD1-26*01 242 1, 3 3466 1, 3 6 IGHD2-2*01 245 2, 3 3467 1, 3 7 IGHD2-8*01 250 2, 3 3468 1 8 IGHD2-15*01 244 2, 3 3469 1, 3 9 IGHD2-21*01 248 2, 3 3470 1, 3 10 IGHD3-3*01 258 1, 2, 3 3471 1, 3 11 IGHD3-9*01 259 2 3472 1, 3 12 IGHD3-10*01 252 2, 3 3473 1, 3 13 IGHD3-16*01 254 2, 3 3474 1, 3 14 IGHD3-22*01 256 2, 3 3475 1, 3 15 IGHD4-4*01 263 2, 3 3476 1, 3 16 IGHD4-11*01 260 2, 3 3477 1, 3 17 IGHD4-17*01 261 2, 3 3478 1, 3 18 IGHD4-23*01 262 2, 3 3479 1, 3 19 IGHD5-5*01 267 1, 2, 3 3480 1, 3 20 IGHD5-12*01 264 1, 3 3481 1, 3 21 IGHD5-18*01 265 1, 2, 3 3482 1, 3 22 IGHD5-24*01 266 1, 3 3483 1, 3 23 IGHD6-6*01 271 1, 2 3484 1, 2, 3 24 IGHD6-13*01 268 1, 2 3485 1, 2, 3 25 IGHD6-19*01 269 1, 2 3486 1, 2, 3 26 IGHD6-25*01 270 1, 2 3487 1, 3 27 IGHD7-27*01 272 1, 3 3488 1, 2
[1170] The following rules (see Table 38) were used to design pairs of D oligos for cloning into the six different vectors to ensure that the resulting heavy chain sequences are in frame. The rules were applied to both the direct and inverted D.sub.H segments. In order to facilite cloning into the BsaI sites of the pF-VH3-23-IGHJ plasmids, the following nucleotides were added to the D.sub.H segment oligonucleotides:
[1171] DIRECT 5' End: CGAAA;
[1172] DIRECT 3' End: T;
[1173] INDIRECT 5' End: CGAAA;
[1174] INDIRECT 3' End: T.
[1175] The resulting oligos for each particular D.sub.H segment are listed in Tables 39-40 below. The nucleotides added to facilite cloning are in lower case letters. The nucleotides encoding the D.sub.H segment are in upper case letters. The oligos were synthesized by standard DNA synthesis technology. Pairs of oligos (encoding a particular D.sub.H segment) were digested with BsaI and ligated into similarly digested plasmids pF-VH3-23-IGHJ1 to pF-VH3-23-IGHJ6 thereby generating a VH3-23 library of 690 members, listed in Table 41 below. The VH3-23 heavy chain library was co-transformed with various light chains (as described in Example 8) creating a Fab library of 3012 members.
TABLE-US-00055 TABLE 38 Rules for generation of DH segment oligos No. of Frame nucleotides Manipulation 1 3N NONE 1 3N + 1 Remove nucleotide at 3' end 1 3N + 2 Add G to 3' end 2 3N Remove nucleotide at 5' end, add G to 3' end 2 3N + 1 Remove nucleotide at 5' end 2 3N + 2 Remove nucleotide at 5' end, add GG to 3' end 3 3N Add G to 5' end, remove nucleotide at 3' end 3 3N + 1 Add G to 5' end, remove 2 nucleotides at 3' end 3 3N + 2 Add G to 5' end
TABLE-US-00056 TABLE 39 Direct DH segment oligos SEQ SEQ ID ID D segment Forward NO Reverse NO 1_IGHD1-1*01 cgaaaGGTACAACTGGAACGACGt 3489 taggaCGTCGTTCCAGTTGTACCt 3490 2_IGHD1-1*01 cgaaaGTACAACTGGAACGACGGt 3491 taggaCCGTCGTTCCAGTTGTACt 3492 3_IGHD1-1*01 cgaaaGGGTACAACTGGAACGACt 3493 taggaGTCGTTCCAGTTGTACCCt 3494 1_IGHD1-7*01 cgaaaGGTATAACTGGAACTACGt 3495 taggaCGTAGTTCCAGTTATACCt 3496 3_IGHD1-7*01 cgaaaGGGTATAACTGGAACTACt 3497 taggaGTAGTTCCAGTTATACCCt 3498 1_IGHD1-14*01 cgaaaGGTATAACCGGAACCACGt 3499 taggaCGTGGTTCCGGTTATACCt 3500 3_IGHD1-14*01 cgaaaGGGTATAACCGGAACCACt 3501 taggaGTGGTTCCGGTTATACCCt 3502 1_IGHD1-20*01 cgaaaGGTATAACTGGAACGACGt 3503 taggaCGTCGTTCCAGTTATACCt 3504 3_IGHD1-20*01 cgaaaGGGTATAACTGGAACGACt 3505 taggaGTCGTTCCAGTTATACCCt 3506 1_IGHD1-26*01 cgaaaGGTATAGTGGGAGCTACT 3507 taggaCGTAGTAGCTCCCACTATAC 3508 ACGt Ct 3_IGHD1-26*01 cgaaaGGGTATAGTGGGAGCTAC 3509 taggaGTAGTAGCTCCCACTATACC 3510 TACt Ct 2_IGHD2-2*01 cgaaaGGATATTGTAGTAGTACCA 3511 taggaGGCATAGCAGCTGGTACTA 3512 GCTGCTATGCCt CTACAATATCCt 3_IGHD2-2*01 cgaaaGAGGATATTGTAGTAGTAC 3513 taggaCATAGCAGCTGGTACTACTA 3514 CAGCTGCTATGt CAATATCCTCt 2_IGHD2-8*01 cgaaaGGATATTGTACTAATGGTG 3515 taggaGGTATAGCATACACCATTAG 3516 TATGCTATACCt TACAATATCCt 3_IGHD2-8*01 cgaaaGAGGATATTGTACTAATGG 3517 taggaTATAGCATACACCATTAGTA 3518 TGTATGCTATAt CAATATCCTCt 2_IGHD2-15*01 cgaaaGGATATTGTAGTGGTGGT 3519 taggaGGAGTAGCAGCTACCACCA 3520 AGCTGCTACTCCt CTACAATATCCt 3_IGHD2-15*01 cgaaaGAGGATATTGTAGTGGTG 3521 taggaAGTAGCAGCTACCACCACTA 3522 GTAGCTGCTACTt CAATATCCTCt 2_IGHD2-21*01 cgaaaGCATATTGTGGTGGTGATT 3523 taggaGGAATAGCAATCACCACCAC 3524 GCTATTCCt AATATGCt 3_IGHD2-21*01 cgaaaGAGCATATTGTGGTGGTG 3525 taggaAATAGCAATCACCACCACAA 3526 ATTGCTATTt TATGCTCt 1_IGHD3-3*01 cgaaaGTATTACGATTTTTGGAGT 3527 taggaGTATAATAACCACTCCAAAA 3528 GGTTATTATACt ATCGTAATACt 2_IGHD3-3*01 cgaaaTATTACGATTTTTGGAGTG 3529 taggaGGTATAATAACCACTCCAAA 3530 GTTATTATACCt GAG AATCGTAATAt 3_IGHD3-3*01 cgaaaGGTATTACGATTTTTG 3531 taggaTATAATAACCACTCCAAAAAT 3532 TGGTTATTATAt CGTAATACCt 2_IGHD3-9*01 cgaaaTATTACGATATTTTGACTG 3533 taggaGTTATAATAACCAGTCAAAA 3534 GTTATTATAACt TATCGTAATAt 2_IGHD3-10*01 cgaaaTATTACTATGGTTCGGGGA 3535 taggaGTTATAATAACTCCCCGAAC 3536 GTTATTATAACt CATAGTAATAt 3_IGHD3-10*01 cgaaaGGTATTACTATGGTTCGG 3537 taggaTATAATAACTCCCCGAACCA 3538 GGAGTTATTATAt TAGTAATACCt 2_IGHD3-16*01 cgaaaTATTATGATTACGTTTGGG 3539 taggaGGTATAAGCATAACTCCCCC 3540 GGAGTTATGCTTATACCt AAACGTAATCATAATAt 3_IGHD3-16*01 cgaaaGGTATTATGATTACGTTTG 3541 taggaTATAAGCATAACTCCCCCAA 3542 GGGGAGTTATGCTTATAt ACGTAATCATAATACCt 2_IGHD3-22*01 cgaaaTATTACTATGATAGTAGTG 3543 taggaGTAGTAATAACCACTACTAT 3544 GTTATTACTACt CATAGTAATAt 3_IGHD3-22*01 cgaaaGGTATTACTATGATAGTAG 3545 taggaAGTAATAACCACTACTATCA 3546 TGGTTATTACTt TAGTAATACCt 2_IGHD4-4*01 (1) cgaaaGACTACAGTAACTACt 3547 taggaGTAGTTACTGTAGTCt 3548 3_IGHD4-4*01 (1) cgaaaGTGACTACAGTAACTt 3549 taggaAGTTACTGTAGTCACt 3550 2_IGHD4-11*01 (1) cgaaaGACTACAGTAACTACt 3551 taggaGTAGTTACTGTAGTCt 3552 3_IGHD4-11*01 (1) cgaaaGTGACTACAGTAACTt 3553 taggaAGTTACTGTAGTCACt 3554 2_IGHD4-17*01 cgaaaGACTACGGTGACTACt 3555 taggaGTAGTCACCGTAGTCt 3556 3_IGHD4-17*01 cgaaaGTGACTACGGTGACTt 3557 taggaAGTCACCGTAGTCACt 3558 2_IGHD4-23*01 cgaaaGACTACGGTGGTAACTCCt 3559 taggaGGAGTTACCACCGTAGTCt 3560 3_IGHD4-23*01 cgaaaGTGACTACGGTGGTAACTt 3561 taggaAGTTACCACCGTAGTCACt 3562 1_IGHD5-5*01 (2) cgaaaGTGGATACAGCTATGGTTA 3563 taggaCGTAACCATAGCTGTATCCA 3564 CGt Ct 2_IGHD5-5*01 (2) cgaaaTGGATACAGCTATGGTTAC 3565 taggaCCGTAACCATAGCTGTATCC 3566 GGt At 3_IGHD5-5*01 (2) cgaaaGGTGGATACAGCTATGGT 3567 taggaGTAACCATAGCTGTATCCAC 3568 TACt Ct 1_IGHD5-12*01 cgaaaGTGGATATAGTGGCTACG 3569 taggaCGTAATCGTAGCCACTATAT 3570 ATTACGt CCACt 3_IGHD5-12*01 cgaaaGGTGGATATAGTGGCTAC 3571 taggaGTAATCGTAGCCACTATATC 3572 GATTACt CACCt 1_IGHD5-18*01 (2) cgaaaGTGGATACAGCTATGGTTA 3573 taggaCGTAACCATAGCTGTATCCA 3574 CGt Ct 2_IGHD5-18*01 (2) cgaaaTGGATACAGCTATGGTTAC 3575 taggaCCGTAACCATAGCTGTATCC 3576 GGt At 3_IGHD5-18*01 (2) cgaaaGGTGGATACAGCTATGGT 3577 taggaGTAACCATAGCTGTATCCAC 3578 TACt Ct 1_IGHD5-24*01 cgaaaGTAGAGATGGCTACAATTA 3579 taggaCGTAATTGTAGCCATCTCTA 3580 CGt Ct 3_IGHD5-24*01 cgaaaGGTAGAGATGGCTACAAT 3581 taggaGTAATTGTAGCCATCTCTAC 3582 TACt Ct 1_IGHD6-6*01 cgaaaGAGTATAGCAGCTCGTCCt 3583 taggaGGACGAGCTGCTATACTCt 3584 2_IGHD6-6*01 cgaaaAGTATAGCAGCTCGTCCGt 3585 taggaCGGACGAGCTGCTATACTt 3586 1_IGHD6-13*01 cgaaaGGGTATAGCAGCAGCTGG 3587 taggaGTACCAGCTGCTGCTATACC 3588 TACt Ct 2_IGHD6-13*01 cgaaaGGTATAGCAGCAGCTGGT 3589 taggaCGTACCAGCTGCTGCTATAC 3590 ACGt Ct 1_IGHD6-19*01 cgaaaGGGTATAGCAGTGGCTGG 3591 taggaGTACCAGCCACTGCTATACC 3592 TACt Ct 2_IGHD6-19*01 cgaaaGGTATAGCAGTGGCTGGT 3593 taggaCGTACCAGCCACTGCTATAC 3594 ACGt Ct 1_IGHD6-25*01 cgaaaGGGTATAGCAGCGGCTACt 3595 taggaGTAGCCGCTGCTATACCCt 3596 2_IGHD6-25*01 cgaaaGGTATAGCAGCGGCTACGt 3597 taggaCGTAGCCGCTGCTATACCt 3598 1_IGHD7-27*01 cgaaaCTAACTGGGGAGt 3599 taggaCTCCCCAGTTAGt 3600 3_IGHD7-27*01 cgaaaGCTAACTGGGGAt 3601 taggaTCCCCAGTTAGCt 3602
TABLE-US-00057 TABLE 40 Inverted DH segment oligos SEQ SEQ ID ID D segment Forward NO Reverse NO 1'_IGHD1-1*01 cgaaaGTCGTTCCAGTTGTACCGt 3603 taggaCGGTACAACTGGAACGACt 3604 2'_IGHD1-1*01 cgaaaTCGTTCCAGTTGTACCGGt 3605 taggaCCGGTACAACTGGAACGAt 3606 3'_IGHD1-1*01 cgaaaGGTCGTTCCAGTTGTACCt 3607 taggaGGTACAACTGGAACGACCt 3608 1'_IGHD1-7*01 cgaaaGTAGTTCCAGTTATACCGt 3609 taggaCGGTATAACTGGAACTACt 3610 3'_IGHD1-7*01 cgaaaGGTAGTTCCAGTTATACCt 3611 taggaGGTATAACTGGAACTACCt 3612 1'_IGHD1-14*01 cgaaaGTGGTTCCGGTTATACCGt 3613 taggaCGGTATAACCGGAACCACt 3614 2'_IGHD1-14*01 cgaaaTGGTTCCGGTTATACCGGt 3615 taggaCCGGTATAACCGGAACCAt 3616 3'_IGHD1-14*01 cgaaaGGTGGTTCCGGTTATACCt 3617 taggaGGTATAACCGGAACCACCt 3618 1'_IGHD1-20*01 cgaaaGTCGTTCCAGTTATACCGt 3619 taggaCGGTATAACTGGAACGACt 3620 2'_IGHD1-20*01 cgaaaTCGTTCCAGTTATACCGGt 3621 taggaCCGGTATAACTGGAACGAt 3622 3'_IGHD1-20*01 cgaaaGGTCGTTCCAGTTATACCt 3623 taggaGGTATAACTGGAACGACCt 3624 1'_IGHD1-26*01 cgaaaGTAGTAGCTCCCACTATAC 3625 taggaCGGTATAGTGGGAGCTACTA 3626 CGt Ct 3'_IGHD1-26*01 cgaaaGGTAGTAGCTCCCACTATA 3627 taggaGGTATAGTGGGAGCTACTAC 3628 CCt Ct 1'_IGHD2-2*01 cgaaaGGCATAGCAGCTGGTACT 3629 taggaGGATATTGTAGTAGTACCAG 3630 ACTACAATATCCt CTGCTATGCCt 3'_IGHD2-2*01 cgaaaGGGCATAGCAGCTGGTAC 3631 taggaGATATTGTAGTAGTACCAGC 3632 TACTACAATATCt TGCTATGCCCt 1'_IGHD2-8*01 cgaaaGGTATAGCATACACCATTA 3633 taggaGGATATTGTACTAATGGTGT 3634 GTACAATATCCt ATGCTATACCt 1'_IGHD2-15*01 cgaaaGGAGTAGCAGCTACCACC 3635 taggaGGATATTGTAGTGGTGGTAG 3636 ACTACAATATCCt CTGCTACTCCt 3'_IGHD2-15*01 cgaaaGGGAGTAGCAGCTACCAC 3637 taggaGATATTGTAGTGGTGGTAGC 3638 CACTACAATATCt TGCTACTCCCt 1'_IGHD2-21*01 cgaaaGGAATAGCAATCACCACC 3639 taggaGCATATTGTGGTGGTGATTG 3640 ACAATATGCt CTATTCCt 3'_IGHD2-21*01 cgaaaGGGAATAGCAATCACCAC 3641 taggaCATATTGTGGTGGTGATTGC 3642 CACAATATGt TATTCCCt 1'_IGHD3-3*01 cgaaaGGTATAATAACCACTCCAA 3643 taggaTATTACGATTTTTGGAGTGG 3644 AAATCGTAATAt TTATTATACCt 3'_IGHD3-3*01 cgaaaGGGTATAATAACCACTCCA 3645 taggaATTACGATTTTTGGAGTGGT 3646 AAAATCGTAATt TATTATACCCt 1'_IGHD3-9*01 cgaaaGTTATAATAACCAGTCAAA 3647 taggaTATTACGATATTTTGACTGGT 3648 ATATCGTAATAt TATTATAACt 3'_IGHD3-9*01 cgaaaGGTTATAATAACCAGTCAA 3649 taggaATTACGATATTTTGACTGGTT 3650 AATATCGTAATt ATTATAACCt 1'_IGHD3-10*01 cgaaaGTTATAATAACTCCCCGAA 3651 taggaTATTACTATGGTTCGGGGAG 3652 CCATAGTAATAt TTATTATAACt 3'_IGHD3-10*01 cgaaaGGTTATAATAACTCCCCGA 3653 taggaATTACTATGGTTCGGGGAGT 3654 ACCATAGTAATt TATTATAACCt 1'_IGHD3-16*01 cgaaaGGTATAAGCATAACTCCCC 3655 taggaTATTATGATTACGTTTGGGG 3656 CAAACGTAATCATAATAt GAGTTATGCTTATACCt 3'_IGHD3-16*01 cgaaaGGGTATAAGCATAACTCC 3657 taggaATTATGATTACGTTTGGGGG 3658 CCCAAACGTAATCATAATt AGTTATGCTTATACCCt 1'_IGHD3-22*01 cgaaaGTAGTAATAACCACTACTA 3659 taggaTATTACTATGATAGTAGTGG 3660 TCATAGTAATAt TTATTACTACt 1'_IGHD4-4*01 (1) cgaaaGTAGTTACTGTAGTCt 3661 taggaGACTACAGTAACTACt 3662 3'_IGHD4-4*01 (1) cgaaaGGTAGTTACTGTAGTt 3663 taggaACTACAGTAACTACCt 3664 1'_IGHD4-11*01 (1) cgaaaGTAGTTACTGTAGTCt 3665 taggaGACTACAGTAACTACt 3666 3'_IGHD4-11*01 (1) cgaaaGGTAGTTACTGTAGTt 3667 taggaACTACAGTAACTACCt 3668 1'_IGHD4-17*01 cgaaaGTAGTCACCGTAGTCt 3669 taggaGACTACGGTGACTACt 3670 3'_IGHD4-17*01 cgaaaGGTAGTCACCGTAGTt 3671 taggaACTACGGTGACTACCt 3672 1'_IGHD4-23*01 cgaaaGGAGTTACCACCGTAGTCt 3673 taggaGACTACGGTGGTAACTCCt 3674 3'_IGHD4-23*01 cgaaaGGGAGTTACCACCGTAGTt 3675 taggaACTACGGTGGTAACTCCCt 3676 1'_IGHD5-5*01 (2) cgaaaGTAACCATAGCTGTATCCA 3677 taggaCGTGGATACAGCTATGGTTA 3678 CGt Ct 3'_IGHD5-5*01 (2) cgaaaGGTAACCATAGCTGTATCC 3679 taggaGTGGATACAGCTATGGTTAC 3680 ACt Ct 1'_IGHD5-12*01 cgaaaGTAATCGTAGCCACTATAT 3681 taggaCGTGGATATAGTGGCTACGA 3682 CCACGt TTACt 3'_IGHD5-12*01 cgaaaGGTAATCGTAGCCACTATA 3683 taggaGTGGATATAGTGGCTACGAT 3684 TCCACt TACCt 1'_IGHD5-18*01 (2) cgaaaGTAACCATAGCTGTATCCA 3685 taggaCGTGGATACAGCTATGGTTA 3686 CGt Ct 3'_IGHD5-18*01 (2) cgaaaGGTAACCATAGCTGTATCC 3687 taggaGTGGATACAGCTATGGTTAC 3688 ACt Ct 1'_IGHD5-24*01 cgaaaGTAATTGTAGCCATCTCTA 3689 taggaCGTAGAGATGGCTACAATTA 3690 CGt Ct 3'_IGHD5-24*01 cgaaaGGTAATTGTAGCCATCTCT 3691 taggaGTAGAGATGGCTACAATTAC 3692 ACt Ct 1'_IGHD6-6*01 cgaaaGGACGAGCTGCTATACTCt 3693 taggaGAGTATAGCAGCTCGTCCt 3694 2'_IGHD6-6*01 cgaaaGACGAGCTGCTATACTCGt 3695 taggaCGAGTATAGCAGCTCGTCt 3696 3'_IGHD6-6*01 cgaaaGGGACGAGCTGCTATACTt 3697 taggaAGTATAGCAGCTCGTCCCt 3698 1'_IGHD6-13*01 cgaaaGTACCAGCTGCTGCTATA 3699 taggaGGGTATAGCAGCAGCTGGT 3700 CCCt ACt 2'_IGHD6-13*01 cgaaaTACCAGCTGCTGCTATACC 3701 taggaCGGGTATAGCAGCAGCTGG 3702 CGt TAt 3'_IGHD6-13*01 cgaaaGGTACCAGCTGCTGCTAT 3703 taggaGGTATAGCAGCAGCTGGTA 3704 ACCt CCt 1'_IGHD6-19*01 cgaaaGTACCAGCCACTGCTATA 3705 taggaGGGTATAGCAGTGGCTGGT 3706 CCCt ACt 2'_IGHD6-19*01 cgaaaTACCAGCCACTGCTATACC 3707 taggaCGGGTATAGCAGTGGCTGG 3708 CGt TAt 3'_IGHD6-19*01 cgaaaGGTACCAGCCACTGCTAT 3709 taggaGGTATAGCAGTGGCTGGTA 3710 ACCt CCt 1'_IGHD6-25*01 cgaaaGTAGCCGCTGCTATACCCt 3711 taggaGGGTATAGCAGCGGCTACt 3712 3'_IGHD6-25*01 cgaaaGGTAGCCGCTGCTATACCt 3713 taggaGGTATAGCAGCGGCTACCt 3714 1'_IGHD7-27*01 cgaaaTCCCCAGTTAGGt 3715 taggaCCTAACTGGGGAt 3716 2'_IGHD7-27*01 cgaaaCCCCAGTTAGGGt 3717 taggaCCCTAACTGGGGt 3718
TABLE-US-00058 TABLE 41 VH3-23 Library Nuc. Prot. SEQ SEQ No. HeavyChain ID NO ID NO 1 >gi|Fabrus|VH3-23_IGHD1- 2070 2760 1*01 > 1_IGHJ1*01 2 >gi|Fabrus|VH3-23_IGHD1- 2071 2761 1*01 > 2_IGHJ1*01 3 >gi|Fabrus|VH3-23_IGHD1- 2072 2762 1*01 > 3_IGHJ1*01 4 >gi|Fabrus|VH3-23_IGHD1- 2073 2763 7*01 > 1_IGHJ1*01 5 >gi|Fabrus|VH3-23_IGHD1- 2074 2764 7*01 > 3_IGHJ1*01 6 >gi|Fabrus|VH3-23_IGHD1- 2075 2765 14*01 > 1_IGHJ1*01 7 >gi|Fabrus|VH3-23_IGHD1- 2076 2766 14*01 > 3_IGHJ1*01 8 >gi|Fabrus|VH3-23_IGHD1- 2077 2767 20*01 > 1_IGHJ1*01 9 >gi|Fabrus|VH3-23_IGHD1- 2078 2768 20*01 > 3_IGHJ1*01 10 >gi|Fabrus|VH3-23_IGHD1- 2079 2769 26*01 > 1_IGHJ1*01 11 >gi|Fabrus|VH3-23_IGHD1- 2080 2770 26*01 > 3_IGHJ1*01 12 >gi|Fabrus|VH3-23_IGHD2- 2081 2771 2*01 > 2_IGHJ1*01 13 >gi|Fabrus|VH3-23_IGHD2- 2082 2772 2*01 > 3_IGHJ1*01 14 >gi|Fabrus|VH3-23_IGHD2- 2083 2773 8*01 > 2_IGHJ1*01 15 >gi|Fabrus|VH3-23_IGHD2- 2084 2774 8*01 > 3_IGHJ1*01 16 >gi|Fabrus|VH3-23_IGHD2- 2085 2775 15*01 > 2_IGHJ1*01 17 >gi|Fabrus|VH3-23_IGHD2- 2086 2776 15*01 > 3_IGHJ1*01 18 >gi|Fabrus|VH3-23_IGHD2- 2087 2777 21*01 > 2_IGHJ1*01 19 >gi|Fabrus|VH3-23_IGHD2- 2088 2778 21*01 > 3_IGHJ1*01 20 >gi|Fabrus|VH3-23_IGHD3- 2089 2779 3*01 > 1_IGHJ1*01 21 >gi|Fabrus|VH3-23_IGHD3- 2090 2780 3*01 > 2_IGHJ1*01 22 >gi|Fabrus|VH3-23_IGHD3- 2091 2781 3*01 > 3_IGHJ1*01 23 >gi|Fabrus|VH3-23_IGHD3- 2092 2782 9*01 > 2_IGHJ1*01 24 >gi|Fabrus|VH3-23_IGHD3- 2093 2783 10*01 > 2_IGHJ1*01 25 >gi|Fabrus|VH3-23_IGHD3- 2094 2784 10*01 > 3_IGHJ1*01 26 >gi|Fabrus|VH3-23_IGHD3- 2095 2785 16*01 > 2_IGHJ1*01 27 >gi|Fabrus|VH3-23_IGHD3- 2096 2786 16*01 > 3_IGHJ1*01 28 >gi|Fabrus|VH3-23_IGHD3- 2097 2787 22*01 > 2_IGHJ1*01 29 >gi|Fabrus|VH3-23_IGHD3- 2098 2788 22*01 > 3_IGHJ1*01 30 >gi|Fabrus|VH3-23_IGHD4- 2099 2789 4*01(1) > 2_IGHJ1*01 31 >gi|Fabrus|VH3-23_IGHD4- 2100 2790 4*01(1) > 3_IGHJ1*01 32 >gi|Fabrus|VH3-23_IGHD4- 2101 2791 11*01(1) > 2_IGHJ1*01 33 >gi|Fabrus|VH3-23_IGHD4- 2102 2792 11*01(1) > 3_IGHJ1*01 34 >gi|Fabrus|VH3-23_IGHD4- 2103 2793 17*01 > 2_IGHJ1*01 35 >gi|Fabrus|VH3-23_IGHD4- 2104 2794 17*01 > 3_IGHJ1*01 36 >gi|Fabrus|VH3-23_IGHD4- 2105 2795 23*01 > 2_IGHJ1*01 37 >gi|Fabrus|VH3-23_IGHD4- 2106 2796 23*01 > 3_IGHJ1*01 38 >gi|Fabrus|VH3-23_IGHD5- 2107 2797 5*01(2) > 1_IGHJ1*01 39 >gi|Fabrus|VH3-23_IGHD5- 2108 2798 5*01(2) > 2_IGHJ1*01 40 >gi|Fabrus|VH3-23_IGHD5- 2109 2799 5*01(2) > 3_IGHJ1*01 41 >gi|Fabrus|VH3-23_IGHD5- 2110 2800 12*01 > 1_IGHJ1*01 42 >gi|Fabrus|VH3-23_IGHD5- 2111 2801 12*01 > 3_IGHJ1*01 43 >gi|Fabrus|VH3-23_IGHD5- 2112 2802 18*01(2) > 1_IGHJ1*01 44 >gi|Fabrus|VH3-23_IGHD5- 2113 2803 18*01(2) > 2_IGHJ1*01 45 >gi|Fabrus|VH3-23_IGHD5- 2114 2804 18*01(2) > 3_IGHJ1*01 46 >gi|Fabrus|VH3-23_IGHD5- 2115 2805 24*01 > 1_IGHJ1*01 47 >gi|Fabrus|VH3-23_IGHD5- 2116 2806 24*01 > 3_IGHJ1*01 48 >gi|Fabrus|VH3-23_IGHD6- 2117 2807 6*01 > 1_IGHJ1*01 49 >gi|Fabrus|VH3-23_IGHD6- 2118 2808 6*01 > 2_IGHJ1*01 50 >gi|Fabrus|VH3-23_IGHD6- 2119 2809 13*01 > 1_IGHJ1*01 51 >gi|Fabrus|VH3-23_IGHD6- 2120 2810 13*01 > 2_IGHJ1*01 52 >gi|Fabrus|VH3-23_IGHD6- 2121 2811 19*01 > 1_IGHJ1*01 53 >gi|Fabrus|VH3-23_IGHD6- 2122 2812 19*01 > 2_IGHJ1*01 54 >gi|Fabrus|VH3-23_IGHD6- 2123 2813 25*01 > 1_IGHJ1*01 55 >gi|Fabrus|VH3-23_IGHD6- 2124 2814 25*01 > 2_IGHJ1*01 56 >gi|Fabrus|VH3-23_IGHD7- 2125 2815 27*01 > 1_IGHJ1*01 57 >gi|Fabrus|VH3-23_IGHD7- 2126 2816 27*01 > 3_IGHJ1*01 58 >gi|Fabrus|VH3-23_IGHD1- 2127 2817 1*01 > 1'_IGHJ1*01 59 >gi|Fabrus|VH3-23_IGHD1- 2128 2818 1*01 > 2'_IGHJ1*01 60 >gi|Fabrus|VH3-23_IGHD1- 2129 2819 1*01 > 3'_IGHJ1*01 61 >gi|Fabrus|VH3-23_IGHD1- 2130 2820 7*01 > 1'_IGHJ1*01 62 >gi|Fabrus|VH3-23_IGHD1- 2131 2821 7*01 > 3'_IGHJ1*01 63 >gi|Fabrus|VH3-23_IGHD1- 2132 2822 14*01 > 1'_IGHJ1*01 64 >gi|Fabrus|VH3-23_IGHD1- 2133 2823 14*01 > 2'_IGHJ1*01 65 >gi|Fabrus|VH3-23_IGHD1- 2134 2824 14*01 > 3'_IGHJ1*01 66 >gi|Fabrus|VH3-23_IGHD1- 2135 2825 20*01 > 1'_IGHJ1*01 67 >gi|Fabrus|VH3-23_IGHD1- 2136 2826 20*01 > 2'_IGHJ1*01 68 >gi|Fabrus|VH3-23_IGHD1- 2137 2827 20*01 > 3'_IGHJ1*01 69 >gi|Fabrus|VH3-23_IGHD1- 2138 2828 26*01 > 1'_IGHJ1*01 70 >gi|Fabrus|VH3-23_IGHD1- 2139 2829 26*01 > 3'_IGHJ1*01 71 >gi|Fabrus|VH3-23_IGHD2- 2140 2830 2*01 > 1'_IGHJ1*01 72 >gi|Fabrus|VH3-23_IGHD2- 2141 2831 2*01 > 3'_IGHJ1*01 73 >gi|Fabrus|VH3-23_IGHD2- 2142 2832 8*01 > 1'_IGHJ1*01 74 >gi|Fabrus|VH3-23_IGHD2- 2143 2833 15*01 > 1'_IGHJ1*01 75 >gi|Fabrus|VH3-23_IGHD2- 2144 2834 15*01 > 3'_IGHJ1*01 76 >gi|Fabrus|VH3-23_IGHD2- 2145 2835 21*01 > 1'_IGHJ1*01 77 >gi|Fabrus|VH3-23_IGHD2- 2146 2836 21*01 > 3'_IGHJ1*01 78 >gi|Fabrus|VH3-23_IGHD3- 2147 2837 3*01 > 1'_IGHJ1*01 79 >gi|Fabrus|VH3-23_IGHD3- 2148 2838 3*01 > 3'_IGHJ1*01 80 >gi|Fabrus|VH3-23_IGHD3- 2149 2839 9*01 > 1'_IGHJ1*01 81 >gi|Fabrus|VH3-23_IGHD3- 2150 2840 9*01 > 3'_IGHJ1*01 82 >gi|Fabrus|VH3-23_IGHD3- 2151 2841 10*01 > 1'_IGHJ1*01 83 >gi|Fabrus|VH3-23_IGHD3- 2152 2842 10*01 > 3'_IGHJ1*01 84 >gi|Fabrus|VH3-23_IGHD3- 2153 2843 16*01 > 1'_IGHJ1*01 85 >gi|Fabrus|VH3-23_IGHD3- 2154 2844 16*01 > 3'_IGHJ1*01 86 >gi|Fabrus|VH3-23_IGHD3- 2155 2845 22*01 > 1'_IGHJ1*01 87 >gi|Fabrus|VH3-23_IGHD4- 2156 2846 4*01(1) > 1'_IGHJ1*01 88 >gi|Fabrus|VH3-23_IGHD4- 2157 2847 4*01(1) > 3'_IGHJ1*01 89 >gi|Fabrus|VH3-23_IGHD4- 2158 2848 11*01(1) > 1'_IGHJ1*01 90 >gi|Fabrus|VH3-23_IGHD4- 2159 2849 11*01(1) > 3'_IGHJ1*01 91 >gi|Fabrus|VH3-23_IGHD4- 2160 2850 17*01 > 1'_IGHJ1*01 92 >gi|Fabrus|VH3-23_IGHD4- 2161 2851 17*01 > 3'_IGHJ1*01 93 >gi|Fabrus|VH3-23_IGHD4- 2162 2852 23*01 > 1'_IGHJ1*01 94 >gi|Fabrus|VH3-23_IGHD4- 2163 2853 23*01 > 3'_IGHJ1*01 95 >gi|Fabrus|VH3-23_IGHD5- 2164 2854 5*01(2) > 1'_IGHJ1*01 96 >gi|Fabrus|VH3-23_IGHD5- 2165 2855 5*01(2) > 3'_IGHJ1*01 97 >gi|Fabrus|VH3-23_IGHD5- 2166 2856 12*01 > 1'_IGHJ1*01 98 >gi|Fabrus|VH3-23_IGHD5- 2167 2857 12*01 > 3'_IGHJ1*01 99 >gi|Fabrus|VH3-23_IGHD5- 2168 2858 18*01(2) > 1'_IGHJ1*01 100 >gi|Fabrus|VH3-23_IGHD5- 2169 2859 18*01(2) > 3'_IGHJ1*01 101 >gi|Fabrus|VH3-23_IGHD5- 2170 2860 24*01 > 1'_IGHJ1*01 102 >gi|Fabrus|VH3-23_IGHD5- 2171 2861 24*01 > 3'_IGHJ1*01 103 >gi|Fabrus|VH3-23_IGHD6- 2172 2862 6*01 > 1'_IGHJ1*01 104 >gi|Fabrus|VH3-23_IGHD6- 2173 2863 6*01 > 2'_IGHJ1*01 105 >gi|Fabrus|VH3-23_IGHD6- 2174 2864 6*01 > 3'_IGHJ1*01 106 >gi|Fabrus|VH3-23_IGHD6- 2175 2865 13*01 > 1'_IGHJ1*01 107 >gi|Fabrus|VH3-23_IGHD6- 2176 2866 13*01 > 2'_IGHJ1*01 108 >gi|Fabrus|VH3-23_IGHD6- 2177 2867 13*01 > 3'_IGHJ1*01 109 >gi|Fabrus|VH3-23_IGHD6- 2178 2868 19*01 > 1'_IGHJ1*01 110 >gi|Fabrus|VH3-23_IGHD6- 2179 2869 19*01 > 2'_IGHJ1*01 111 >gi|Fabrus|VH3-23_IGHD6- 2180 2870 19*01 > 3'_IGHJ1*01 112 >gi|Fabrus|VH3-23_IGHD6- 2181 2871 25*01 > 1'_IGHJ1*01 113 >gi|Fabrus|VH3-23_IGHD6- 2182 2872 25*01 > 3'_IGHJ1*01 114 >gi|Fabrus|VH3-23_IGHD7- 2183 2873 27*01 > 1'_IGHJ1*01 115 >gi|Fabrus|VH3-23_IGHD7- 2184 2874 27*01 > 2'_IGHJ1*01 116 >gi|Fabrus|VH3-23_IGHD1- 2185 2875 1*01 > 1_IGHJ2*01 117 >gi|Fabrus|VH3-23_IGHD1- 2186 2876 1*01 > 2_IGHJ2*01 118 >gi|Fabrus|VH3-23_IGHD1- 2187 2877 1*01 > 3_IGHJ2*01 119 >gi|Fabrus|VH3-23_IGHD1- 2188 2878 7*01 > 1_IGHJ2*01 120 >gi|Fabrus|VH3-23_IGHD1- 2189 2879 7*01 > 3_IGHJ2*01 121 >gi|Fabrus|VH3-23_IGHD1- 2190 2880 14*01 > 1_IGHJ2*01 122 >gi|Fabrus|VH3-23_IGHD1- 2191 2881
14*01 > 3_IGHJ2*01 123 >gi|Fabrus|VH3-23_IGHD1- 2192 2882 20*01 > 1_IGHJ2*01 124 >gi|Fabrus|VH3-23_IGHD1- 2193 2883 20*01 > 3_IGHJ2*01 125 >gi|Fabrus|VH3-23_IGHD1- 2194 2884 26*01 > 1_IGHJ2*01 126 >gi|Fabrus|VH3-23_IGHD1- 2195 2885 26*01 > 3_IGHJ2*01 127 >gi|Fabrus|VH3-23_IGHD2- 2196 2886 2*01 > 2_IGHJ2*01 128 >gi|Fabrus|VH3-23_IGHD2- 2197 2887 2*01 > 3_IGHJ2*01 129 >gi|Fabrus|VH3-23_IGHD2- 2198 2888 8*01 > 2_IGHJ2*01 130 >gi|Fabrus|VH3-23_IGHD2- 2199 2889 8*01 > 3_IGHJ2*01 131 >gi|Fabrus|VH3-23_IGHD2- 2200 2890 15*01 > 2_IGHJ2*01 132 >gi|Fabrus|VH3-23_IGHD2- 2201 2891 15*01 > 3_IGHJ2*01 133 >gi|Fabrus|VH3-23_IGHD2- 2202 2892 21*01 > 2_IGHJ2*01 134 >gi|Fabrus|VH3-23_IGHD2- 2203 2893 21*01 > 3_IGHJ2*01 135 >gi|Fabrus|VH3-23_IGHD3- 2204 2894 3*01 > 1_IGHJ2*01 136 >gi|Fabrus|VH3-23_IGHD3- 2205 2895 3*01 > 2_IGHJ2*01 137 >gi|Fabrus|VH3-23_IGHD3- 2206 2896 3*01 > 3_IGHJ2*01 138 >gi|Fabrus|VH3-23_IGHD3- 2207 2897 9*01 > 2_IGHJ2*01 139 >gi|Fabrus|VH3-23_IGHD3- 2208 2898 10*01 > 2_IGHJ2*01 140 >gi|Fabrus|VH3-23_IGHD3- 2209 2899 10*01 > 3_IGHJ2*01 141 >gi|Fabrus|VH3-23_IGHD3- 2210 2900 16*01 > 2_IGHJ2*01 142 >gi|Fabrus|VH3-23_IGHD3- 2211 2901 16*01 > 3_IGHJ2*01 143 >gi|Fabrus|VH3-23_IGHD3- 2212 2902 22*01 > 2_IGHJ2*01 144 >gi|Fabrus|VH3-23_IGHD3- 2213 2903 22*01 > 3_IGHJ2*01 145 >gi|Fabrus|VH3-23_IGHD4- 2214 2904 4*01(1) > 2_IGHJ2*01 146 >gi|Fabrus|VH3-23_IGHD4- 2215 2905 4*01(1) > 3_IGHJ2*01 147 >gi|Fabrus|VH3-23_IGHD4- 2216 2906 11*01(1) > 2_IGHJ2*01 148 >gi|Fabrus|VH3-23_IGHD4- 2217 2907 11*01(1) > 3_IGHJ2*01 149 >gi|Fabrus|VH3-23_IGHD4- 2218 2908 17*01 > 2_IGHJ2*01 150 >gi|Fabrus|VH3-23_IGHD4- 2219 2909 17*01 > 3_IGHJ2*01 151 >gi|Fabrus|VH3-23_IGHD4- 2220 2910 23*01 > 2_IGHJ2*01 152 >gi|Fabrus|VH3-23_IGHD4- 2221 2911 23*01 > 3_IGHJ2*01 153 >gi|Fabrus|VH3-23_IGHD5- 2222 2912 5*01(2) > 1_IGHJ2*01 154 >gi|Fabrus|VH3-23_IGHD5- 2223 2913 5*01(2) > 2_IGHJ2*01 155 >gi|Fabrus|VH3-23_IGHD5- 2224 2914 5*01(2) > 3_IGHJ2*01 156 >gi|Fabrus|VH3-23_IGHD5- 2225 2915 12*01 > 1_IGHJ2*01 157 >gi|Fabrus|VH3-23_IGHD5- 2226 2916 12*01 > 3_IGHJ2*01 158 >gi|Fabrus|VH3-23_IGHD5- 2227 2917 18*01(2) > 1_IGHJ2*01 159 >gi|Fabrus|VH3-23_IGHD5- 2228 2918 18*01(2) > 2_IGHJ2*01 160 >gi|Fabrus|VH3-23_IGHD5- 2229 2919 18*01(2) > 3_IGHJ2*01 161 >gi|Fabrus|VH3-23_IGHD5- 2230 2920 24*01 > 1_IGHJ2*01 162 >gi|Fabrus|VH3-23_IGHD5- 2231 2921 24*01 > 3_IGHJ2*01 163 >gi|Fabrus|VH3-23_IGHD6- 2232 2922 6*01 > 1_IGHJ2*01 164 >gi|Fabrus|VH3-23_IGHD6- 2233 2923 6*01 > 2_IGHJ2*01 165 >gi|Fabrus|VH3-23_IGHD6- 2234 2924 13*01 > 1_IGHJ2*01 166 >gi|Fabrus|VH3-23_IGHD6- 2235 2925 13*01 > 2_IGHJ2*01 167 >gi|Fabrus|VH3-23_IGHD6- 2236 2926 19*01 > 1_IGHJ2*01 168 >gi|Fabrus|VH3-23_IGHD6- 2237 2927 19*01 > 2_IGHJ2*01 169 >gi|Fabrus|E06_VH3-23_IGHD6- 2238 2928 25*01 > 1_IGHJ2*01 170 >gi|Fabrus|E07_VH3-23_IGHD6- 2239 2929 25*01 > 2_IGHJ2*01 171 >gi|Fabrus|E08_VH3-23_IGHD7- 2240 2930 27*01 > 1_IGHJ2*01 172 >gi|Fabrus|E09_VH3-23_IGHD7- 2241 2931 27*01 > 3_IGHJ2*01 173 >gi|Fabrus|A01_VH3-23_IGHD1- 2242 2932 1*01 > 1'_IGHJ2*01 174 >gi|Fabrus|A02_VH3-23_IGHD1- 2243 2933 1*01 > 2'_IGHJ2*01 175 >gi|Fabrus|A03_VH3-23_IGHD1- 2244 2934 1*01 > 3'_IGHJ2*01 176 >gi|Fabrus|A04_VH3-23_IGHD1- 2245 2935 7*01 > 1'_IGHJ2*01 177 >gi|Fabrus|A05_VH3-23_IGHD1- 2246 2936 7*01 > 3'_IGHJ2*01 178 >gi|Fabrus|A06_VH3-23_IGHD1- 2247 2937 14*01 > 1'_IGHJ2*01 179 >gi|Fabrus|A07_VH3-23_IGHD1- 2248 2938 14*01 > 2'_IGHJ2*01 180 >gi|Fabrus|A08_VH3-23_IGHD1- 2249 2939 14*01 > 3'_IGHJ2*01 181 >gi|Fabrus|A09_VH3-23_IGHD1- 2250 2940 20*01 > 1'_IGHJ2*01 182 >gi|Fabrus|A10_VH3-23_IGHD1- 2251 2941 20*01 > 2'_IGHJ2*01 183 >gi|Fabrus|A11_VH3-23_IGHD1- 2252 2942 20*01 > 3'_IGHJ2*01 184 >gi|Fabrus|A12_VH3-23_IGHD1- 2253 2943 26*01 > 1'_IGHJ2*01 185 >gi|Fabrus|B01_VH3-23_IGHD1- 2254 2944 26*01 > 3'_IGHJ2*01 186 >gi|Fabrus|B02_VH3-23_IGHD2- 2255 2945 2*01 > 1'_IGHJ2*01 187 >gi|Fabrus|B03_VH3-23_IGHD2- 2256 2946 2*01 > 3'_IGHJ2*01 188 >gi|Fabrus|B04_VH3-23_IGHD2- 2257 2947 8*01 > 1'_IGHJ2*01 189 >gi|Fabrus|B05_VH3-23_IGHD2- 2258 2948 15*01 > 1'_IGHJ2*01 190 >gi|Fabrus|B06_VH3-23_IGHD2- 2259 2949 15*01 > 3'_IGHJ2*01 191 >gi|Fabrus|B07_VH3-23_IGHD2- 2260 2950 21*01 > 1'_IGHJ2*01 192 >gi|Fabrus|B08_VH3-23_IGHD2- 2261 2951 21*01 > 3'_IGHJ2*01 193 >gi|Fabrus|B09_VH3-23_IGHD3- 2262 2952 3*01 > 1'_IGHJ2*01 194 >gi|Fabrus|B10_VH3-23_IGHD3- 2263 2953 3*01 > 3'_IGHJ2*01 195 >gi|Fabrus|B11_VH3-23_IGHD3- 2264 2954 9*01 > 1'_IGHJ2*01 196 >gi|Fabrus|B12_VH3-23_IGHD3- 2265 2955 9*01 > 3'_IGHJ2*01 197 >gi|Fabrus|C01_VH3-23_IGHD3- 2266 2956 10*01 > 1'_IGHJ2*01 198 >gi|Fabrus|C02_VH3-23_IGHD3- 2267 2957 10*01 > 3'_IGHJ2*01 199 >gi|Fabrus|C03_VH3-23_IGHD3- 2268 2958 16*01 > 1'_IGHJ2*01 200 >gi|Fabrus|C04_VH3-23_IGHD3- 2269 2959 16*01 > 3'_IGHJ2*01 201 >gi|Fabrus|C05_VH3-23_IGHD3- 2270 2960 22*01 > 1'_IGHJ2*01 202 >gi|Fabrus|C06_VH3-23_IGHD4- 2271 2961 4*01(1) > 1'_IGHJ2*01 203 >gi|Fabrus|C07_VH3-23_IGHD4- 2272 2962 4*01(1) > 3'_IGHJ2*01 204 >gi|Fabrus|C08_VH3-23_IGHD4- 2273 2963 11*01(1) > 1'_IGHJ2*01 205 >gi|Fabrus|C09_VH3-23_IGHD4- 2274 2964 11*01(1) > 3'_IGHJ2*01 206 >gi|Fabrus|C10_VH3-23_IGHD4- 2275 2965 17*01 > 1'_IGHJ2*01 207 >gi|Fabrus|C11_VH3-23_IGHD4- 2276 2966 17*01 > 3'_IGHJ2*01 208 >gi|Fabrus|C12_VH3-23_IGHD4- 2277 2967 23*01 > 1'_IGHJ2*01 209 >gi|Fabrus|D01_VH3-23_IGHD4- 2278 2968 23*01 > 3'_IGHJ2*01 210 >gi|Fabrus|D02_VH3-23_IGHD5- 2279 2969 5*01(2) > 1'_IGHJ2*01 211 >gi|Fabrus|D03_VH3-23_IGHD5- 2280 2970 5*01(2) > 3'_IGHJ2*01 212 >gi|Fabrus|D04_VH3-23_IGHD5- 2281 2971 12*01 > 1'_IGHJ2*01 213 >gi|Fabrus|D05_VH3-23_IGHD5- 2282 2972 12*01 > 3'_IGHJ2*01 214 >gi|Fabrus|D06_VH3-23_IGHD5- 2283 2973 18*01(2) > 1'_IGHJ2*01 215 >gi|Fabrus|D07_VH3-23_IGHD5- 2284 2974 18*01(2) > 3'_IGHJ2*01 216 >gi|Fabrus|D08_VH3-23_IGHD5- 2285 2975 24*01 > 1'_IGHJ2*01 217 >gi|Fabrus|D09_VH3-23_IGHD5- 2286 2976 24*01 > 3'_IGHJ2*01 218 >gi|Fabrus|D10_VH3-23_IGHD6- 2287 2977 6*01 > 1'_IGHJ2*01 219 >gi|Fabrus|D11_VH3-23_IGHD6- 2288 2978 6*01 > 2'_IGHJ2*01 220 >gi|Fabrus|D12_VH3-23_IGHD6- 2289 2979 6*01 > 3'_IGHJ2*01 221 >gi|Fabrus|E01_VH3-23_IGHD6- 2290 2980 13*01 > 1'_IGHJ2*01 222 >gi|Fabrus|E02_VH3-23_IGHD6- 2291 2981 13*01 > 2'_IGHJ2*01 223 >gi|Fabrus|E03_VH3-23_IGHD6- 2292 2982 13*01 > 3'_IGHJ2*01 224 >gi|Fabrus|E04_VH3-23_IGHD6- 2293 2983 19*01 > 1'_IGHJ2*01 225 >gi|Fabrus|E05_VH3-23_IGHD6- 2294 2984 19*01 > 2'_IGHJ2*01 226 >gi|Fabrus|E06_VH3-23_IGHD6- 2295 2985 19*01 > 3'_IGHJ2*01 227 >gi|Fabrus|E07_VH3-23_IGHD6- 2296 2986 25*01 > 1'_IGHJ2*01 228 >gi|Fabrus|E08_VH3-23_IGHD6- 2297 2987 25*01 > 3'_IGHJ2*01 229 >gi|Fabrus|E09_VH3-23_IGHD7- 2298 2988 27*01 > 1'_IGHJ2*01 230 >gi|Fabrus|E10_VH3-23_IGHD7- 2299 2989 27*01 > 2'_IGHJ2*01 231 >gi|Fabrus|A01_VH3-23_IGHD1- 2300 2990 1*01 > 1_IGHJ3*01 232 >gi|Fabrus|A02_VH3-23_IGHD1- 2301 2991 1*01 > 2_IGHJ3*01 233 >gi|Fabrus|A03_VH3-23_IGHD1- 2302 2992 1*01 > 3_IGHJ3*01 234 >gi|Fabrus|A04_VH3-23_IGHD1- 2303 2993 7*01 > 1_IGHJ3*01 235 >gi|Fabrus|A05_VH3-23_IGHD1- 2304 2994 7*01 > 3_IGHJ3*01 236 >gi|Fabrus|A06_VH3-23_IGHD1- 2305 2995 14*01 > 1_IGHJ3*01 237 >gi|Fabrus|A07_VH3-23_IGHD1- 2306 2996 14*01 > 3_IGHJ3*01 238 >gi|Fabrus|A08_VH3-23_IGHD1- 2307 2997 20*01 > 1_IGHJ3*01 239 >gi|Fabrus|A09_VH3-23_IGHD1- 2308 2998 20*01 > 3_IGHJ3*01 240 >gi|Fabrus|A10_VH3-23_IGHD1- 2309 2999 26*01 > 1_IGHJ3*01 241 >gi|Fabrus|A11_VH3-23_IGHD1- 2310 3000 26*01 > 3_IGHJ3*01 242 >gi|Fabrus|A12_VH3-23_IGHD2- 2311 3001 2*01 > 2_IGHJ3*01 243 >gi|Fabrus|B01_VH3-23_IGHD2- 2312 3002 2*01 > 3_IGHJ3*01 244 >gi|Fabrus|B02_VH3-23_IGHD2- 2313 3003 8*01 > 2_IGHJ3*01 245 >gi|Fabrus|B03_VH3-23_IGHD2- 2314 3004 8*01 > 3_IGHJ3*01 246 >gi|Fabrus|B04_VH3-23_IGHD2- 2315 3005 15*01 > 2_IGHJ3*01 247 >gi|Fabrus|B05_VH3-23_IGHD2- 2316 3006 15*01 > 3_IGHJ3*01
248 >gi|Fabrus|B06_VH3-23_IGHD2- 2317 3007 21*01 > 2_IGHJ3*01 249 >gi|Fabrus|B07_VH3-23_IGHD2- 2318 3008 21*01 > 3_IGHJ3*01 250 >gi|Fabrus|B08_VH3-23_IGHD3- 2319 3009 3*01 > 1_IGHJ3*01 251 >gi|Fabrus|B09_VH3-23_IGHD3- 2320 3010 3*01 > 2_IGHJ3*01 252 >gi|Fabrus|B10_VH3-23_IGHD3- 2321 3011 3*01 > 3_IGHJ3*01 253 >gi|Fabrus|B11_VH3-23_IGHD3- 2322 3012 9*01 > 2_IGHJ3*01 254 >gi|Fabrus|B12_VH3-23_IGHD3- 2323 3013 10*01 > 2_IGHJ3*01 255 >gi|Fabrus|C01_VH3-23_IGHD3- 2324 3014 10*01 > 3_IGHJ3*01 256 >gi|Fabrus|C02_VH3-23_IGHD3- 2325 3015 16*01 > 2_IGHJ3*01 257 >gi|Fabrus|C03_VH3-23_IGHD3- 2326 3016 16*01 > 3_IGHJ3*01 258 >gi|Fabrus|C04_VH3-23_IGHD3- 2327 3017 22*01 > 2_IGHJ3*01 259 >gi|Fabrus|C05_VH3-23_IGHD3- 2328 3018 22*01 > 3_IGHJ3*01 260 >gi|Fabrus|C06_VH3-23_IGHD4- 2329 3019 4*01(1) > 2_IGHJ3*01 261 >gi|Fabrus|C07_VH3-23_IGHD4- 2330 3020 4*01(1) > 3_IGHJ3*01 262 >gi|Fabrus|C08_VH3-23_IGHD4- 2331 3021 11*01(1) > 2_IGHJ3*01 263 >gi|Fabrus|C09_VH3-23_IGHD4- 2332 3022 11*01(1) > 3_IGHJ3*01 264 >gi|Fabrus|C10_VH3-23_IGHD4- 2333 3023 17*01 > 2_IGHJ3*01 265 >gi|Fabrus|C11_VH3-23_IGHD4- 2334 3024 17*01 > 3_IGHJ3*01 266 >gi|Fabrus|C12_VH3-23_IGHD4- 2335 3025 23*01 > 2_IGHJ3*01 267 >gi|Fabrus|D01_VH3-23_IGHD4- 2336 3026 23*01 > 3_IGHJ3*01 268 >gi|Fabrus|D02_VH3-23_IGHD5- 2337 3027 5*01(2) > 1_IGHJ3*01 269 >gi|Fabrus|D03_VH3-23_IGHD5- 2338 3028 5*01(2) > 2_IGHJ3*01 270 >gi|Fabrus|D04_VH3-23_IGHD5- 2339 3029 5*01(2) > 3_IGHJ3*01 271 >gi|Fabrus|D05_VH3-23_IGHD5- 2340 3030 12*01 > 1_IGHJ3*01 272 >gi|Fabrus|D06_VH3-23_IGHD5- 2341 3031 12*01 > 3_IGHJ3*01 273 >gi|Fabrus|D07_VH3-23_IGHD5- 2342 3032 18*01(2) > 1_IGHJ3*01 274 >gi|Fabrus|D08_VH3-23_IGHD5- 2343 3033 18*01(2) > 2_IGHJ3*01 275 >gi|Fabrus|D09_VH3-23_IGHD5- 2344 3034 18*01(2) > 3_IGHJ3*01 276 >gi|Fabrus|D10_VH3-23_IGHD5- 2345 3035 24*01 > 1_IGHJ3*01 277 >gi|Fabrus|D11_VH3-23_IGHD5- 2346 3036 24*01 > 3_IGHJ3*01 278 >gi|Fabrus|D12_VH3-23_IGHD6- 2347 3037 6*01 > 1_IGHJ3*01 279 >gi|Fabrus|E01_VH3-23_IGHD6- 2348 3038 6*01 > 2_IGHJ3*01 280 >gi|Fabrus|E02_VH3-23_IGHD6- 2349 3039 13*01 > 1_IGHJ3*01 281 >gi|Fabrus|E03_VH3-23_IGHD6- 2350 3040 13*01 > 2_IGHJ3*01 282 >gi|Fabrus|E04_VH3-23_IGHD6- 2351 3041 19*01 > 1_IGHJ3*01 283 >gi|Fabrus|E05_VH3-23_IGHD6- 2352 3042 19*01 > 2_IGHJ3*01 284 >gi|Fabrus|E06_VH3-23_IGHD6- 2353 3043 25*01 > 1_IGHJ3*01 285 >gi|Fabrus|E07_VH3-23_IGHD6- 2354 3044 25*01 > 2_IGHJ3*01 286 >gi|Fabrus|E08_VH3-23_IGHD7- 2355 3045 27*01 > 1_IGHJ3*01 287 >gi|Fabrus|E09_VH3-23_IGHD7- 2356 3046 27*01 > 3_IGHJ3*01 288 >gi|Fabrus|A01_VH3-23_IGHD1- 2357 3047 1*01 > 1'_IGHJ3*01 289 >gi|Fabrus|A02_VH3-23_IGHD1- 2358 3048 1*01 > 2'_IGHJ3*01 290 >gi|Fabrus|A03_VH3-23_IGHD1- 2359 3049 1*01 > 3'_IGHJ3*01 291 >gi|Fabrus|A04_VH3-23_IGHD1- 2360 3050 7*01 > 1'_IGHJ3*01 292 >gi|Fabrus|A05_VH3-23_IGHD1- 2361 3051 7*01 > 3'_IGHJ3*01 293 >gi|Fabrus|A06_VH3-23_IGHD1- 2362 3052 14*01 > 1'_IGHJ3*01 294 >gi|Fabrus|A07_VH3-23_IGHD1- 2363 3053 14*01 > 2'_IGHJ3*01 295 >gi|Fabrus|A08_VH3-23_IGHD1- 2364 3054 14*01 > 3'_IGHJ3*01 296 >gi|Fabrus|A09_VH3-23_IGHD1- 2365 3055 20*01 > 1'_IGHJ3*01 297 >gi|Fabrus|A10_VH3-23_IGHD1- 2366 3056 20*01 > 2'_IGHJ3*01 298 >gi|Fabrus|A11_VH3-23_IGHD1- 2367 3057 20*01 > 3'_IGHJ3*01 299 >gi|Fabrus|A12_VH3-23_IGHD1- 2368 3058 26*01 > 1'_IGHJ3*01 300 >gi|Fabrus|B01_VH3-23_IGHD1- 2369 3059 26*01 > 3'_IGHJ3*01 301 >gi|Fabrus|B02_VH3-23_IGHD2- 2370 3060 2*01 > 1'_IGHJ3*01 302 >gi|Fabrus|B03_VH3-23_IGHD2- 2371 3061 2*01 > 3'_IGHJ3*01 303 >gi|Fabrus|B04_VH3-23_IGHD2- 2372 3062 8*01 > 1'_IGHJ3*01 304 >gi|Fabrus|B05_VH3-23_IGHD2- 2373 3063 15*01 > 1'_IGHJ3*01 305 >gi|Fabrus|B06_VH3-23_IGHD2- 2374 3064 15*01 > 3'_IGHJ3*01 306 >gi|Fabrus|B07_VH3-23_IGHD2- 2375 3065 21*01 > 1'_IGHJ3*01 307 >gi|Fabrus|B08_VH3-23_IGHD2- 2376 3066 21*01 > 3'_IGHJ3*01 308 >gi|Fabrus|B09_VH3-23_IGHD3- 2377 3067 3*01 > 1'_IGHJ3*01 309 >gi|Fabrus|B10_VH3-23_IGHD3- 2378 3068 3*01 > 3'_IGHJ3*01 310 >gi|Fabrus|B11_VH3-23_IGHD3- 2379 3069 9*01 > 1'_IGHJ3*01 311 >gi|Fabrus|B12_VH3-23_IGHD3- 2380 3070 9*01 > 3'_IGHJ3*01 312 >gi|Fabrus|C01_VH3-23_IGHD3- 2381 3071 10*01 > 1'_IGHJ3*01 313 >gi|Fabrus|C02_VH3-23_IGHD3- 2382 3072 10*01 > 3'_IGHJ3*01 314 >gi|Fabrus|C03_VH3-23_IGHD3- 2383 3073 16*01 > 1'_IGHJ3*01 315 >gi|Fabrus|C04_VH3-23_IGHD3- 2384 3074 16*01 > 3'_IGHJ3*01 316 >gi|Fabrus|C05_VH3-23_IGHD3- 2385 3075 22*01 > 1'_IGHJ3*01 317 >gi|Fabrus|C06_VH3-23_IGHD4- 2386 3076 4*01(1) > 1'_IGHJ3*01 318 >gi|Fabrus|C07_VH3-23_IGHD4- 2387 3077 4*01(1) > 3'_IGHJ3*01 319 >gi|Fabrus|C08_VH3-23_IGHD4- 2388 3078 11*01(1) > 1'_IGHJ3*01 320 >gi|Fabrus|C09_VH3-23_IGHD4- 2389 3079 11*01(1) > 3'_IGHJ3*01 321 >gi|Fabrus|C10_VH3-23_IGHD4- 2390 3080 17*01 > 1'_IGHJ3*01 322 >gi|Fabrus|C11_VH3-23_IGHD4- 2391 3081 17*01 > 3'_IGHJ3*01 323 >gi|Fabrus|C12_VH3-23_IGHD4- 2392 3082 23*01 > 1'_IGHJ3*01 324 >gi|Fabrus|D01_VH3-23_IGHD4- 2393 3083 23*01 > 3'_IGHJ3*01 325 >gi|Fabrus|D02_VH3-23_IGHD5- 2394 3084 5*01(2) > 1'_IGHJ3*01 326 >gi|Fabrus|D03_VH3-23_IGHD5- 2395 3085 5*01(2) > 3'_IGHJ3*01 327 >gi|Fabrus|D04_VH3-23_IGHD5- 2396 3086 12*01 > 1'_IGHJ3*01 328 >gi|Fabrus|D05_VH3-23_IGHD5- 2397 3087 12*01 > 3'_IGHJ3*01 329 >gi|Fabrus|D06_VH3-23_IGHD5- 2398 3088 18*01(2) > 1'_IGHJ3*01 330 >gi|Fabrus|D07_VH3-23_IGHD5- 2399 3089 18*01(2) > 3'_IGHJ3*01 331 >gi|Fabrus|D08_VH3-23_IGHD5- 2400 3090 24*01 > 1'_IGHJ3*01 332 >gi|Fabrus|D09_VH3-23_IGHD5- 2401 3091 24*01 > 3'_IGHJ3*01 333 >gi|Fabrus|D10_VH3-23_IGHD6- 2402 3092 6*01 > 1'_IGHJ3*01 334 >gi|Fabrus|D11_VH3-23_IGHD6- 2403 3093 6*01 > 2'_IGHJ3*01 335 >gi|Fabrus|D12_VH3-23_IGHD6- 2404 3094 6*01 > 3'_IGHJ3*01 336 >gi|Fabrus|E01_VH3-23_IGHD6- 2405 3095 13*01 > 1'_IGHJ3*01 337 >gi|Fabrus|E02_VH3-23_IGHD6- 2406 3096 13*01 > 2'_IGHJ3*01 338 >gi|Fabrus|E03_VH3-23_IGHD6- 2407 3097 13*01 > 3'_IGHJ3*01 339 >gi|Fabrus|E04_VH3-23_IGHD6- 2408 3098 19*01 > 1'_IGHJ3*01 340 >gi|Fabrus|E05_VH3-23_IGHD6- 2409 3099 19*01 > 2'_IGHJ3*01 341 >gi|Fabrus|E06_VH3-23_IGHD6- 2410 3100 19*01 > 3'_IGHJ3*01 342 >gi|Fabrus|E07_VH3-23_IGHD6- 2411 3101 25*01 > 1'_IGHJ3*01 343 >gi|Fabrus|E08_VH3-23_IGHD6- 2412 3102 25*01 > 3'_IGHJ3*01 344 >gi|Fabrus|E09_VH3-23_IGHD7- 2413 3103 27*01 > 1'_IGHJ3*01 345 >gi|Fabrus|E10_VH3-23_IGHD7- 2414 3104 27*01 > 2'_IGHJ3*01 346 >gi|Fabrus|A01_VH3-23_IGHD1- 2415 3105 1*01 > 1_IGHJ4*01 347 >gi|Fabrus|A02_VH3-23_IGHD1- 2416 3106 1*01 > 2_IGHJ4*01 348 >gi|Fabrus|A03_VH3-23_IGHD1- 2417 3107 1*01 > 3_IGHJ4*01 349 >gi|Fabrus|A04_VH3-23_IGHD1- 2418 3108 7*01 > 1_IGHJ4*01 350 >gi|Fabrus|A05_VH3-23_IGHD1- 2419 3109 7*01 > 3_IGHJ4*01 351 >gi|Fabrus|A06_VH3-23_IGHD1- 2420 3110 14*01 > 1_IGHJ4*01 352 >gi|Fabrus|A07_VH3-23_IGHD1- 2421 3111 14*01 > 3_IGHJ4*01 353 >gi|Fabrus|A08_VH3-23_IGHD1- 2422 3112 20*01 > 1_IGHJ4*01 354 >gi|Fabrus|A09_VH3-23_IGHD1- 2423 3113 20*01 > 3_IGHJ4*01 355 >gi|Fabrus|A10_VH3-23_IGHD1- 2424 3114 26*01 > 1_IGHJ4*01 356 >gi|Fabrus|A11_VH3-23_IGHD1- 2425 3115 26*01 > 3_IGHJ4*01 357 >gi|Fabrus|A12_VH3-23_IGHD2- 2426 3116 2*01 > 2_IGHJ4*01 358 >gi|Fabrus|B01_VH3-23_IGHD2- 2427 3117 2*01 > 3_IGHJ4*01 359 >gi|Fabrus|B02_VH3-23_IGHD2- 2428 3118 8*01 > 2_IGHJ4*01 360 >gi|Fabrus|B03_VH3-23_IGHD2- 2429 3119 8*01 > 3_IGHJ4*01 361 >gi|Fabrus|B04_VH3-23_IGHD2- 2430 3120 15*01 > 2_IGHJ4*01 362 >gi|Fabrus|B05_VH3-23_IGHD2- 2431 3121 15*01 > 3_IGHJ4*01 363 >gi|Fabrus|B06_VH3-23_IGHD2- 2432 3122 21*01 > 2_IGHJ4*01 364 >gi|Fabrus|B07_VH3-23_IGHD2- 2433 3123 21*01 > 3_IGHJ4*01 365 >gi|Fabrus|B08_VH3-23_IGHD3- 2434 3124 3*01 > 1_IGHJ4*01 366 >gi|Fabrus|B09_VH3-23_IGHD3- 2435 3125 3*01 > 2_IGHJ4*01 367 >gi|Fabrus|B10_VH3-23_IGHD3- 2436 3126 3*01 > 3_IGHJ4*01 368 >gi|Fabrus|B11_VH3-23_IGHD3- 2437 3127 9*01 > 2_IGHJ4*01 369 >gi|Fabrus|B12_VH3-23_IGHD3- 2438 3128 10*01 > 2_IGHJ4*01 370 >gi|Fabrus|C01_VH3-23_IGHD3- 2439 3129 10*01 > 3_IGHJ4*01 371 >gi|Fabrus|C02_VH3-23_IGHD3- 2440 3130 16*01 > 2_IGHJ4*01 372 >gi|Fabrus|C03_VH3-23_IGHD3- 2441 3131 16*01 > 3_IGHJ4*01 373 >gi|Fabrus|C04_VH3-23_IGHD3- 2442 3132
22*01 > 2_IGHJ4*01 374 >gi|Fabrus|C05_VH3-23_IGHD3- 2443 3133 22*01 > 3_IGHJ4*01 375 >gi|Fabrus|C06_VH3-23_IGHD4- 2444 3134 4*01(1) > 2_IGHJ4*01 376 >gi|Fabrus|C07_VH3-23_IGHD4- 2445 3135 4*01(1) > 3_IGHJ4*01 377 >gi|Fabrus|C08_VH3-23_IGHD4- 2446 3136 11*01(1) > 2_IGHJ4*01 378 >gi|Fabrus|C09_VH3-23_IGHD4- 2447 3137 11*01(1) > 3_IGHJ4*01 379 >gi|Fabrus|C10_VH3-23_IGHD4- 2448 3138 17*01 > 2_IGHJ4*01 380 >gi|Fabrus|C11_VH3-23_IGHD4- 2449 3139 17*01 > 3_IGHJ4*01 381 >gi|Fabrus|C12_VH3-23_IGHD4- 2450 3140 23*01 > 2_IGHJ4*01 382 >gi|Fabrus|D01_VH3-23_IGHD4- 2451 3141 23*01 > 3_IGHJ4*01 383 >gi|Fabrus|D02_VH3-23_IGHD5- 2452 3142 5*01(2) > 1_IGHJ4*01 384 >gi|Fabrus|D03_VH3-23_IGHD5- 2453 3143 5*01(2) > 2_IGHJ4*01 385 >gi|Fabrus|D04_VH3-23_IGHD5- 2454 3144 5*01(2) > 3_IGHJ4*01 386 >gi|Fabrus|D05_VH3-23_IGHD5- 2455 3145 12*01 > 1_IGHJ4*01 387 >gi|Fabrus|D06_VH3-23_IGHD5- 2456 3146 12*01 > 3_IGHJ4*01 388 >gi|Fabrus|D07_VH3-23_IGHD5- 2457 3147 18*01(2) > 1_IGHJ4*01 389 >gi|Fabrus|D08_VH3-23_IGHD5- 2458 3148 18*01(2) > 2_IGHJ4*01 390 >gi|Fabrus|D09_VH3-23_IGHD5- 2459 3149 18*01(2) > 3_IGHJ4*01 391 >gi|Fabrus|D10_VH3-23_IGHD5- 2460 3150 24*01 > 1_IGHJ4*01 392 >gi|Fabrus|D11_VH3-23_IGHD5- 2461 3151 24*01 > 3_IGHJ4*01 393 >gi|Fabrus|D12_VH3-23_IGHD6- 2462 3152 6*01 > 1_IGHJ4*01 394 >gi|Fabrus|E01_VH3-23_IGHD6- 2463 3153 6*01 > 2_IGHJ4*01 395 >gi|Fabrus|E02_VH3-23_IGHD6- 2464 3154 13*01 > 1_IGHJ4*01 396 >gi|Fabrus|E03_VH3-23_IGHD6- 2465 3155 13*01 > 2_IGHJ4*01 397 >gi|Fabrus|E04_VH3-23_IGHD6- 2466 3156 19*01 > 1_IGHJ4*01 398 >gi|Fabrus|E05_VH3-23_IGHD6- 2467 3157 19*01 > 2_IGHJ4*01 399 >gi|Fabrus|E06_VH3-23_IGHD6- 2468 3158 25*01 > 1_IGHJ4*01 400 >gi|Fabrus|E07_VH3-23_IGHD6- 2469 3159 25*01 > 2_IGHJ4*01 401 >gi|Fabrus|E08_VH3-23_IGHD7- 2470 3160 27*01 > 1_IGHJ4*01 402 >gi|Fabrus|E09_VH3-23_IGHD7- 2471 3161 27*01 > 3_IGHJ4*01 403 >gi|Fabrus|A01_VH3-23_IGHD1- 2472 3162 1*01 > 1'_IGHJ4*01 404 >gi|Fabrus|A02_VH3-23_IGHD1- 2473 3163 1*01 > 2'_IGHJ4*01 405 >gi|Fabrus|A03_VH3-23_IGHD1- 2474 3164 1*01 > 3'_IGHJ4*01 406 >gi|Fabrus|A04_VH3-23_IGHD1- 2475 3165 7*01 > 1'_IGHJ4*01 407 >gi|Fabrus|A05_VH3-23_IGHD1- 2476 3166 7*01 > 3'_IGHJ4*01 408 >gi|Fabrus|A06_VH3-23_IGHD1- 2477 3167 14*01 > 1'_IGHJ4*01 409 >gi|Fabrus|A07_VH3-23_IGHD1- 2478 3168 14*01 > 2'_IGHJ4*01 410 >gi|Fabrus|A08_VH3-23_IGHD1- 2479 3169 14*01 > 3'_IGHJ4*01 411 >gi|Fabrus|A09_VH3-23_IGHD1- 2480 3170 20*01 > 1'_IGHJ4*01 412 >gi|Fabrus|A10_VH3-23_IGHD1- 2481 3171 20*01 > 2'_IGHJ4*01 413 >gi|Fabrus|A11_VH3-23_IGHD1- 2482 3172 20*01 > 3'_IGHJ4*01 414 >gi|Fabrus|A12_VH3-23_IGHD1- 2483 3173 26*01 > 1'_IGHJ4*01 415 >gi|Fabrus|B01_VH3-23_IGHD1- 2484 3174 26*01 > 3'_IGHJ4*01 416 >gi|Fabrus|B02_VH3-23_IGHD2- 2485 3175 2*01 > 1'_IGHJ4*01 417 >gi|Fabrus|B03_VH3-23_IGHD2- 2486 3176 2*01 > 3'_IGHJ4*01 418 >gi|Fabrus|B04_VH3-23_IGHD2- 2487 3177 8*01 > 1'_IGHJ4*01 419 >gi|Fabrus|B05_VH3-23_IGHD2- 2488 3178 15*01 > 1'_IGHJ4*01 420 >gi|Fabrus|B06_VH3-23_IGHD2- 2489 3179 15*01 > 3'_IGHJ4*01 421 >gi|Fabrus|B07_VH3-23_IGHD2- 2490 3180 21*01 > 1'_IGHJ4*01 422 >gi|Fabrus|B08_VH3-23_IGHD2- 2491 3181 21*01 > 3'_IGHJ4*01 423 >gi|Fabrus|B09_VH3-23_IGHD3- 2492 3182 3*01 > 1'_IGHJ4*01 424 >gi|Fabrus|B10_VH3-23_IGHD3- 2493 3183 3*01 > 3'_IGHJ4*01 425 >gi|Fabrus|B11_VH3-23_IGHD3- 2494 3184 9*01 > 1'_IGHJ4*01 426 >gi|Fabrus|B12_VH3-23_IGHD3- 2495 3185 9*01 > 3'_IGHJ4*01 427 >gi|Fabrus|C01_VH3-23_IGHD3- 2496 3186 10*01 > 1'_IGHJ4*01 428 >gi|Fabrus|C02_VH3-23_IGHD3- 2497 3187 10*01 > 3'_IGHJ4*01 429 >gi|Fabrus|C03_VH3-23_IGHD3- 2498 3188 16*01 > 1'_IGHJ4*01 430 >gi|Fabrus|C04_VH3-23_IGHD3- 2499 3189 16*01 > 3'_IGHJ4*01 431 >gi|Fabrus|C05_VH3-23_IGHD3- 2500 3190 22*01 > 1'_IGHJ4*01 432 >gi|Fabrus|C06_VH3-23_IGHD4- 2501 3191 4*01(1) > 1'_IGHJ4*01 433 >gi|Fabrus|C07_VH3-23_IGHD4- 2502 3192 4*01(1) > 3'_IGHJ4*01 434 >gi|Fabrus|C08_VH3-23_IGHD4- 2503 3193 11*01(1) > 1'_IGHJ4*01 435 >gi|Fabrus|C09_VH3-23_IGHD4- 2504 3194 11*01(1) > 3'_IGHJ4*01 436 >gi|Fabrus|C10_VH3-23_IGHD4- 2505 3195 17*01 > 1'_IGHJ4*01 437 >gi|Fabrus|C11_VH3-23_IGHD4- 2506 3196 17*01 > 3'_IGHJ4*01 438 >gi|Fabrus|C12_VH3-23_IGHD4- 2507 3197 23*01 > 1'_IGHJ4*01 439 >gi|Fabrus|D01_VH3-23_IGHD4- 2508 3198 23*01 > 3'_IGHJ4*01 440 >gi|Fabrus|D02_VH3-23_IGHD5- 2509 3199 5*01(2) > 1'_IGHJ4*01 441 >gi|Fabrus|D03_VH3-23_IGHD5- 2510 3200 5*01(2) > 3'_IGHJ4*01 442 >gi|Fabrus|D04_VH3-23_IGHD5- 2511 3201 12*01 > 1'_IGHJ4*01 443 >gi|Fabrus|D05_VH3-23_IGHD5- 2512 3202 12*01 > 3'_IGHJ4*01 444 >gi|Fabrus|D06_VH3-23_IGHD5- 2513 3203 18*01(2) > 1'_IGHJ4*01 445 >gi|Fabrus|D07_VH3-23_IGHD5- 2514 3204 18*01(2) > 3'_IGHJ4*01 446 >gi|Fabrus|D08_VH3-23_IGHD5- 2515 3205 24*01 > 1'_IGHJ4*01 447 >gi|Fabrus|D09_VH3-23_IGHD5- 2516 3206 24*01 > 3'_IGHJ4*01 448 >gi|Fabrus|D10_VH3-23_IGHD6- 2517 3207 6*01 > 1'_IGHJ4*01 449 >gi|Fabrus|D11_VH3-23_IGHD6- 2518 3208 6*01 > 2'_IGHJ4*01 450 >gi|Fabrus|D12_VH3-23_IGHD6- 2519 3209 6*01 > 3'_IGHJ4*01 451 >gi|Fabrus|E01_VH3-23_IGHD6- 2520 3210 13*01 > 1'_IGHJ4*01 452 >gi|Fabrus|E02_VH3-23_IGHD6- 2521 3211 13*01 > 2'_IGHJ4*01 453 >gi|Fabrus|E03_VH3-23_IGHD6- 2522 3212 13*01 > 3'_IGHJ4*01 454 >gi|Fabrus|E04_VH3-23_IGHD6- 2523 3213 19*01 > 1'_IGHJ4*01 455 >gi|Fabrus|E05_VH3-23_IGHD6- 2524 3214 19*01 > 2'_IGHJ4*01 456 >gi|Fabrus|E06_VH3-23_IGHD6- 2525 3215 19*01 > 3'_IGHJ4*01 457 >gi|Fabrus|E07_VH3-23_IGHD6- 2526 3216 25*01 > 1'_IGHJ4*01 458 >gi|Fabrus|E08_VH3-23_IGHD6- 2527 3217 25*01 > 3'_IGHJ4*01 459 >gi|Fabrus|E09_VH3-23_IGHD7- 2528 3218 27*01 > 1'_IGHJ4*01 460 >gi|Fabrus|E10_VH3-23_IGHD7- 2529 3219 27*01 > 2'_IGHJ4*01 461 >gi|Fabrus|A01_VH3-23_IGHD1- 2530 3220 1*01 > 1_IGHJ5*01 462 >gi|Fabrus|A02_VH3-23_IGHD1- 2531 3221 1*01 > 2_IGHJ5*01 463 >gi|Fabrus|A03_VH3-23_IGHD1- 2532 3222 1*01 > 3_IGHJ5*01 464 >gi|Fabrus|A04_VH3-23_IGHD1- 2533 3223 7*01 > 1_IGHJ5*01 465 >gi|Fabrus|A05_VH3-23_IGHD1- 2534 3224 7*01 > 3_IGHJ5*01 466 >gi|Fabrus|A06_VH3-23_IGHD1- 2535 3225 14*01 > 1_IGHJ5*01 467 >gi|Fabrus|A07_VH3-23_IGHD1- 2536 3226 14*01 > 3_IGHJ5*01 468 >gi|Fabrus|A08_VH3-23_IGHD1- 2537 3227 20*01 > 1_IGHJ5*01 469 >gi|Fabrus|A09_VH3-23_IGHD1- 2538 3228 20*01 > 3_IGHJ5*01 470 >gi|Fabrus|A10_VH3-23_IGHD1- 2539 3229 26*01 > 1_IGHJ5*01 471 >gi|Fabrus|A11_VH3-23_IGHD1- 2540 3230 26*01 > 3_IGHJ5*01 472 >gi|Fabrus|A12_VH3-23_IGHD2- 2541 3231 2*01 > 2_IGHJ5*01 473 >gi|Fabrus|B01_VH3-23_IGHD2- 2542 3232 2*01 > 3_IGHJ5*01 474 >gi|Fabrus|B02_VH3-23_IGHD2- 2543 3233 8*01 > 2_IGHJ5*01 475 >gi|Fabrus|B03_VH3-23_IGHD2- 2544 3234 8*01 > 3_IGHJ5*01 476 >gi|Fabrus|B04_VH3-23_IGHD2- 2545 3235 15*01 > 2_IGHJ5*01 477 >gi|Fabrus|B05_VH3-23_IGHD2- 2546 3236 15*01 > 3_IGHJ5*01 478 >gi|Fabrus|B06_VH3-23_IGHD2- 2547 3237 21*01 > 2_IGHJ5*01 479 >gi|Fabrus|B07_VH3-23_IGHD2- 2548 3238 21*01 > 3_IGHJ5*01 480 >gi|Fabrus|B08_VH3-23_IGHD3- 2549 3239 3*01 > 1_IGHJ5*01 481 >gi|Fabrus|B09_VH3-23_IGHD3- 2550 3240 3*01 > 2_IGHJ5*01 482 >gi|Fabrus|B10_VH3-23_IGHD3- 2551 3241 3*01 > 3_IGHJ5*01 483 >gi|Fabrus|B11_VH3-23_IGHD3- 2552 3242 9*01 > 2_IGHJ5*01 484 >gi|Fabrus|B12_VH3-23_IGHD3- 2553 3243 10*01 > 2_IGHJ5*01 485 >gi|Fabrus|C01_VH3-23_IGHD3- 2554 3244 10*01 > 3_IGHJ5*01 486 >gi|Fabrus|C02_VH3-23_IGHD3- 2555 3245 16*01 > 2_IGHJ5*01 487 >gi|Fabrus|C03_VH3-23_IGHD3- 2556 3246 16*01 > 3_IGHJ5*01 488 >gi|Fabrus|C04_VH3-23_IGHD3- 2557 3247 22*01 > 2_IGHJ5*01 489 >gi|Fabrus|C05_VH3-23_IGHD3- 2558 3248 22*01 > 3_IGHJ5*01 490 >gi|Fabrus|C06_VH3-23_IGHD4- 2559 3249 4*01(1) > 2_IGHJ5*01 491 >gi|Fabrus|C07_VH3-23_IGHD4- 2560 3250 4*01(1) > 3_IGHJ5*01 492 >gi|Fabrus|C08_VH3-23_IGHD4- 2561 3251 11*01(1) > 2_IGHJ5*01 493 >gi|Fabrus|C09_VH3-23_IGHD4- 2562 3252 11*01(1) > 3_IGHJ5*01 494 >gi|Fabrus|C10_VH3-23_IGHD4- 2563 3253 17*01 > 2_IGHJ5*01 495 >gi|Fabrus|C11_VH3-23_IGHD4- 2564 3254 17*01 > 3_IGHJ5*01 496 >gi|Fabrus|C12_VH3-23_IGHD4- 2565 3255 23*01 > 2_IGHJ5*01 497 >gi|Fabrus|D01_VH3-23_IGHD4- 2566 3256 23*01 > 3_IGHJ5*01 498 >gi|Fabrus|D02_VH3-23_IGHD5- 2567 3257 5*01(2) > 1_IGHJ5*01
499 >gi|Fabrus|D03_VH3-23_IGHD5- 2568 3258 5*01(2) > 2_IGHJ5*01 500 >gi|Fabrus|D04_VH3-23_IGHD5- 2569 3259 5*01(2) > 3_IGHJ5*01 501 >gi|Fabrus|D05_VH3-23_IGHD5- 2570 3260 12*01 > 1_IGHJ5*01 502 >gi|Fabrus|D06_VH3-23_IGHD5- 2571 3261 12*01 > 3_IGHJ5*01 503 >gi|Fabrus|D07_VH3-23_IGHD5- 2572 3262 18*01(2) > 1_IGHJ5*01 504 >gi|Fabrus|D08_VH3-23_IGHD5- 2573 3263 18*01(2) > 2_IGHJ5*01 505 >gi|Fabrus|D09_VH3-23_IGHD5- 2574 3264 18*01(2) > 3_IGHJ5*01 506 >gi|Fabrus|D10_VH3-23_IGHD5- 2575 3265 24*01 > 1_IGHJ5*01 507 >gi|Fabrus|D11_VH3-23_IGHD5- 2576 3266 24*01 > 3_IGHJ5*01 508 >gi|Fabrus|D12_VH3-23_IGHD6- 2577 3267 6*01 > 1_IGHJ5*01 509 >gi|Fabrus|E01_VH3-23_IGHD6- 2578 3268 6*01 > 2_IGHJ5*01 510 >gi|Fabrus|E02_VH3-23_IGHD6- 2579 3269 13*01 > 1_IGHJ5*01 511 >gi|Fabrus|E03_VH3-23_IGHD6- 2580 3270 13*01 > 2_IGHJ5*01 512 >gi|Fabrus|E04_VH3-23_IGHD6- 2581 3271 19*01 > 1_IGHJ5*01 513 >gi|Fabrus|E05_VH3-23_IGHD6- 2582 3272 19*01 > 2_IGHJ5*01 514 >gi|Fabrus|E06_VH3-23_IGHD6- 2583 3273 25*01 > 1_IGHJ5*01 515 >gi|Fabrus|E07_VH3-23_IGHD6- 2584 3274 25*01 > 2_IGHJ5*01 516 >gi|Fabrus|E08_VH3-23_IGHD7- 2585 3275 27*01 > 1_IGHJ5*01 517 >gi|Fabrus|E09_VH3-23_IGHD7- 2586 3276 27*01 > 3_IGHJ5*01 518 >gi|Fabrus|A01_VH3-23_IGHD1- 2587 3277 1*01 > 1'_IGHJ5*01 519 >gi|Fabrus|A02_VH3-23_IGHD1- 2588 3278 1*01 > 2'_IGHJ5*01 520 >gi|Fabrus|A03_VH3-23_IGHD1- 2589 3279 1*01 > 3'_IGHJ5*01 521 >gi|Fabrus|A04_VH3-23_IGHD1- 2590 3280 7*01 > 1'_IGHJ5*01 522 >gi|Fabrus|A05_VH3-23_IGHD1- 2591 3281 7*01 > 3'_IGHJ5*01 523 >gi|Fabrus|A06_VH3-23_IGHD1- 2592 3282 14*01 > 1'_IGHJ5*01 524 >gi|Fabrus|A07_VH3-23_IGHD1- 2593 3283 14*01 > 2'_IGHJ5*01 525 >gi|Fabrus|A08_VH3-23_IGHD1- 2594 3284 14*01 > 3'_IGHJ5*01 526 >gi|Fabrus|A09_VH3-23_IGHD1- 2595 3285 20*01 > 1'_IGHJ5*01 527 >gi|Fabrus|A10_VH3-23_IGHD1- 2596 3286 20*01 > 2'_IGHJ5*01 528 >gi|Fabrus|A11_VH3-23_IGHD1- 2597 3287 20*01 > 3'_IGHJ5*01 529 >gi|Fabrus|A12_VH3-23_IGHD1- 2598 3288 26*01 > 1'_IGHJ5*01 530 >gi|Fabrus|B01_VH3-23_IGHD1- 2599 3289 26*01 > 3'_IGHJ5*01 531 >gi|Fabrus|B02_VH3-23_IGHD2- 2600 3290 2*01 > 1'_IGHJ5*01 532 >gi|Fabrus|B03_VH3-23_IGHD2- 2601 3291 2*01 > 3'_IGHJ5*01 533 >gi|Fabrus|B04_VH3-23_IGHD2- 2602 3292 8*01 > 1'_IGHJ5*01 534 >gi|Fabrus|B05_VH3-23_IGHD2- 2603 3293 15*01 > 1'_IGHJ5*01 535 >gi|Fabrus|B06_VH3-23_IGHD2- 2604 3294 15*01 > 3'_IGHJ5*01 536 >gi|Fabrus|B07_VH3-23_IGHD2- 2605 3295 21*01 > 1'_IGHJ5*01 537 >gi|Fabrus|B08_VH3-23_IGHD2- 2606 3296 21*01 > 3'_IGHJ5*01 538 >gi|Fabrus|B09_VH3-23_IGHD3- 2607 3297 3*01 > 1'_IGHJ5*01 539 >gi|Fabrus|B10_VH3-23_IGHD3- 2608 3298 3*01 > 3'_IGHJ5*01 540 >gi|Fabrus|B11_VH3-23_IGHD3- 2609 3299 9*01 > 1'_IGHJ5*01 541 >gi|Fabrus|B12_VH3-23_IGHD3- 2610 3300 9*01 > 3'_IGHJ5*01 542 >gi|Fabrus|C01_VH3-23_IGHD3- 2611 3301 10*01 > 1'_IGHJ5*01 543 >gi|Fabrus|C02_VH3-23_IGHD3- 2612 3302 10*01 > 3'_IGHJ5*01 544 >gi|Fabrus|C03_VH3-23_IGHD3- 2613 3303 16*01 > 1'_IGHJ5*01 545 >gi|Fabrus|C04_VH3-23_IGHD3- 2614 3304 16*01 > 3'_IGHJ5*01 546 >gi|Fabrus|C05_VH3-23_IGHD3- 2615 3305 22*01 > 1'_IGHJ5*01 547 >gi|Fabrus|C06_VH3-23_IGHD4- 2616 3306 4*01(1) > 1'_IGHJ5*01 548 >gi|Fabrus|C07_VH3-23_IGHD4- 2617 3307 4*01(1) > 3'_IGHJ5*01 549 >gi|Fabrus|C08_VH3-23_IGHD4- 2618 3308 11*01(1) > 1'_IGHJ5*01 550 >gi|Fabrus|C09_VH3-23_IGHD4- 2619 3309 11*01(1) > 3'_IGHJ5*01 551 >gi|Fabrus|C10_VH3-23_IGHD4- 2620 3310 17*01 > 1'_IGHJ5*01 552 >gi|Fabrus|C11_VH3-23_IGHD4- 2621 3311 17*01 > 3'_IGHJ5*01 553 >gi|Fabrus|C12_VH3-23_IGHD4- 2622 3312 23*01 > 1'_IGHJ5*01 554 >gi|Fabrus|D01_VH3-23_IGHD4- 2623 3313 23*01 > 3'_IGHJ5*01 555 >gi|Fabrus|D02_VH3-23_IGHD5- 2624 3314 5*01(2) > 1'_IGHJ5*01 556 >gi|Fabrus|D03_VH3-23_IGHD5- 2625 3315 5*01(2) > 3'_IGHJ5*01 557 >gi|Fabrus|D04_VH3-23_IGHD5- 2626 3316 12*01 > 1'_IGHJ5*01 558 >gi|Fabrus|D05_VH3-23_IGHD5- 2627 3317 12*01 > 3'_IGHJ5*01 559 >gi|Fabrus|D06_VH3-23_IGHD5- 2628 3318 18*01(2) > 1'_IGHJ5*01 560 >gi|Fabrus|D07_VH3-23_IGHD5- 2629 3319 18*01(2) > 3'_IGHJ5*01 561 >gi|Fabrus|D08_VH3-23_IGHD5- 2630 3320 24*01 > 1'_IGHJ5*01 562 >gi|Fabrus|D09_VH3-23_IGHD5- 2631 3321 24*01 > 3'_IGHJ5*01 563 >gi|Fabrus|D10_VH3-23_IGHD6- 2632 3322 6*01 > 1'_IGHJ5*01 564 >gi|Fabrus|D11_VH3-23_IGHD6- 2633 3323 6*01 > 2'_IGHJ5*01 565 >gi|Fabrus|D12_VH3-23_IGHD6- 2634 3324 6*01 > 3'_IGHJ5*01 566 >gi|Fabrus|E01_VH3-23_IGHD6- 2635 3325 13*01 > 1'_IGHJ5*01 567 >gi|Fabrus|E02_VH3-23_IGHD6- 2636 3326 13*01 > 2'_IGHJ5*01 568 >gi|Fabrus|E03_VH3-23_IGHD6- 2637 3327 13*01 > 3'_IGHJ5*01 569 >gi|Fabrus|E04_VH3-23_IGHD6- 2638 3328 19*01 > 1'_IGHJ5*01 570 >gi|Fabrus|E05_VH3-23_IGHD6- 2639 3329 19*01 > 2'_IGHJ5*01 571 >gi|Fabrus|E06_VH3-23_IGHD6- 2640 3330 19*01 > 3'_IGHJ5*01 572 >gi|Fabrus|E07_VH3-23_IGHD6- 2641 3331 25*01 > 1'_IGHJ5*01 573 >gi|Fabrus|E08_VH3-23_IGHD6- 2642 3332 25*01 > 3'_IGHJ5*01 574 >gi|Fabrus|E09_VH3-23_IGHD7- 2643 3333 27*01 > 1'_IGHJ5*01 575 >gi|Fabrus|E10_VH3-23_IGHD7- 2644 3334 27*01 > 2'_IGHJ5*01 576 >gi|Fabrus|A01_VH3-23_IGHD1- 2645 3335 1*01 > 1_IGHJ6*01 577 >gi|Fabrus|A02_VH3-23_IGHD1- 2646 3336 1*01 > 2_IGHJ6*01 578 >gi|Fabrus|A03_VH3-23_IGHD1- 2647 3337 1*01 > 3_IGHJ6*01 579 >gi|Fabrus|A04_VH3-23_IGHD1- 2648 3338 7*01 > 1_IGHJ6*01 580 >gi|Fabrus|A05_VH3-23_IGHD1- 2649 3339 7*01 > 3_IGHJ6*01 581 >gi|Fabrus|A06_VH3-23_IGHD1- 2650 3340 14*01 > 1_IGHJ6*01 582 >gi|Fabrus|A07_VH3-23_IGHD1- 2651 3341 14*01 > 3_IGHJ6*01 583 >gi|Fabrus|A08_VH3-23_IGHD1- 2652 3342 20*01 > 1_IGHJ6*01 584 >gi|Fabrus|A09_VH3-23_IGHD1- 2653 3343 20*01 > 3_IGHJ6*01 585 >gi|Fabrus|A10_VH3-23_IGHD1- 2654 3344 26*01 > 1_IGHJ6*01 586 >gi|Fabrus|A11_VH3-23_IGHD1- 2655 3345 26*01 > 3_IGHJ6*01 587 >gi|Fabrus|A12_VH3-23_IGHD2- 2656 3346 2*01 > 2_IGHJ6*01 588 >gi|Fabrus|B01_VH3-23_IGHD2- 2657 3347 2*01 > 3_IGHJ6*01 589 >gi|Fabrus|B02_VH3-23_IGHD2- 2658 3348 8*01 > 2_IGHJ6*01 590 >gi|Fabrus|B03_VH3-23_IGHD2- 2659 3349 8*01 > 3_IGHJ6*01 591 >gi|Fabrus|B04_VH3-23_IGHD2- 2660 3350 15*01 > 2_IGHJ6*01 592 >gi|Fabrus|B05_VH3-23_IGHD2- 2661 3351 15*01 > 3_IGHJ6*01 593 >gi|Fabrus|B06_VH3-23_IGHD2- 2662 3352 21*01 > 2_IGHJ6*01 594 >gi|Fabrus|B07_VH3-23_IGHD2- 2663 3353 21*01 > 3_IGHJ6*01 595 >gi|Fabrus|B08_VH3-23_IGHD3- 2664 3354 3*01 > 1_IGHJ6*01 596 >gi|Fabrus|B09_VH3-23_IGHD3- 2665 3355 3*01 > 2_IGHJ6*01 597 >gi|Fabrus|B10_VH3-23_IGHD3- 2666 3356 3*01 > 3_IGHJ6*01 598 >gi|Fabrus|B11_VH3-23_IGHD3- 2667 3357 9*01 > 2_IGHJ6*01 599 >gi|Fabrus|B12_VH3-23_IGHD3- 2668 3358 10*01 > 2_IGHJ6*01 600 >gi|Fabrus|C01_VH3-23_IGHD3- 2669 3359 10*01 > 3_IGHJ6*01 601 >gi|Fabrus|C02_VH3-23_IGHD3- 2670 3360 16*01 > 2_IGHJ6*01 602 >gi|Fabrus|C03_VH3-23_IGHD3- 2671 3361 16*01 > 3_IGHJ6*01 603 >gi|Fabrus|C04_VH3-23_IGHD3- 2672 3362 22*01 > 2_IGHJ6*01 604 >gi|Fabrus|C05_VH3-23_IGHD3- 2673 3363 22*01 > 3_IGHJ6*01 605 >gi|Fabrus|C06_VH3-23_IGHD4- 2674 3364 4*01(1) > 2_IGHJ6*01 606 >gi|Fabrus|C07_VH3-23_IGHD4- 2675 3365 4*01(1) > 3_IGHJ6*01 607 >gi|Fabrus|C08_VH3-23_IGHD4- 2676 3366 11*01(1) > 2_IGHJ6*01 608 >gi|Fabrus|C09_VH3-23_IGHD4- 2677 3367 11*01(1) > 3_IGHJ6*01 609 >gi|Fabrus|C10_VH3-23_IGHD4- 2678 3368 17*01 > 2_IGHJ6*01 610 >gi|Fabrus|C11_VH3-23_IGHD4- 2679 3369 17*01 > 3_IGHJ6*01 611 >gi|Fabrus|C12_VH3-23_IGHD4- 2680 3370 23*01 > 2_IGHJ6*01 612 >gi|Fabrus|D01_VH3-23_IGHD4- 2681 3371 23*01 > 3_IGHJ6*01 613 >gi|Fabrus|D02_VH3-23_IGHD5- 2682 3372 5*01(2) > 1_IGHJ6*01 614 >gi|Fabrus|D03_VH3-23_IGHD5- 2683 3373 5*01(2) > 2_IGHJ6*01 615 >gi|Fabrus|D04_VH3-23_IGHD5- 2684 3374 5*01(2) > 3_IGHJ6*01 616 >gi|Fabrus|D05_VH3-23_IGHD5- 2685 3375 12*01 > 1_IGHJ6*01 617 >gi|Fabrus|D06_VH3-23_IGHD5- 2686 3376 12*01 > 3_IGHJ6*01 618 >gi|Fabrus|D07_VH3-23_IGHD5- 2687 3377 18*01(2) > 1_IGHJ6*01 619 >gi|Fabrus|D08_VH3-23_IGHD5- 2688 3378 18*01(2) > 2_IGHJ6*01 620 >gi|Fabrus|D09_VH3-23_IGHD5- 2689 3379 18*01(2) > 3_IGHJ6*01 621 >gi|Fabrus|D10_VH3-23_IGHD5- 2690 3380 24*01 > 1_IGHJ6*01 622 >gi|Fabrus|D11_VH3-23_IGHD5- 2691 3381 24*01 > 3_IGHJ6*01 623 >gi|Fabrus|D12_VH3-23_IGHD6- 2692 3382 6*01 > 1_IGHJ6*01 624 >gi|Fabrus|E01_VH3-23_IGHD6- 2693 3383
6*01 > 2_IGHJ6*01 625 >gi|Fabrus|E02_VH3-23_IGHD6- 2694 3384 13*01 > 1_IGHJ6*01 626 >gi|Fabrus|E03_VH3-23_IGHD6- 2695 3385 13*01 > 2_IGHJ6*01 627 >gi|Fabrus|E04_VH3-23_IGHD6- 2696 3386 19*01 > 1_IGHJ6*01 628 >gi|Fabrus|E05_VH3-23_IGHD6- 2697 3387 19*01 > 2_IGHJ6*01 629 >gi|Fabrus|E06_VH3-23_IGHD6- 2698 3388 25*01 > 1_IGHJ6*01 630 >gi|Fabrus|E07_VH3-23_IGHD6- 2699 3389 25*01 > 2_IGHJ6*01 631 >gi|Fabrus|E08_VH3-23_IGHD7- 2700 3390 27*01 > 1_IGHJ6*01 632 >gi|Fabrus|E09_VH3-23_IGHD7- 2701 3391 27*01 > 3_IGHJ6*01 633 >gi|Fabrus|A01_VH3-23_IGHD1- 2702 3392 1*01 > 1'_IGHJ6*01 634 >gi|Fabrus|A02_VH3-23_IGHD1- 2703 3393 1*01 > 2'_IGHJ6*01 635 >gi|Fabrus|A03_VH3-23_IGHD1- 2704 3394 1*01 > 3'_IGHJ6*01 636 >gi|Fabrus|A04_VH3-23_IGHD1- 2705 3395 7*01 > 1'_IGHJ6*01 637 >gi|Fabrus|A05_VH3-23_IGHD1- 2706 3396 7*01 > 3'_IGHJ6*01 638 >gi|Fabrus|A06_VH3-23_IGHD1- 2707 3397 14*01 > 1'_IGHJ6*01 639 >gi|Fabrus|A07_VH3-23_IGHD1- 2708 3398 14*01 > 2'_IGHJ6*01 640 >gi|Fabrus|A08_VH3-23_IGHD1- 2709 3399 14*01 > 3'_IGHJ6*01 641 >gi|Fabrus|A09_VH3-23_IGHD1- 2710 3400 20*01 > 1'_IGHJ6*01 642 >gi|Fabrus|A10_VH3-23_IGHD1- 2711 3401 20*01 > 2'_IGHJ6*01 643 >gi|Fabrus|A11_VH3-23_IGHD1- 2712 3402 20*01 > 3'_IGHJ6*01 644 >gi|Fabrus|A12_VH3-23_IGHD1- 2713 3403 26*01 > 1'_IGHJ6*01 645 >gi|Fabrus|B01_VH3-23_IGHD1- 2714 3404 26*01 > 3'_IGHJ6*01 646 >gi|Fabrus|B02_VH3-23_IGHD2- 2715 3405 2*01 > 1'_IGHJ6*01 647 >gi|Fabrus|B03_VH3-23_IGHD2- 2716 3406 2*01 > 3'_IGHJ6*01 648 >gi|Fabrus|B04_VH3-23_IGHD2- 2717 3407 8*01 > 1'_IGHJ6*01 649 >gi|Fabrus|B05_VH3-23_IGHD2- 2718 3408 15*01 > 1'_IGHJ6*01 650 >gi|Fabrus|B06_VH3-23_IGHD2- 2719 3409 15*01 > 3'_IGHJ6*01 651 >gi|Fabrus|B07_VH3-23_IGHD2- 2720 3410 21*01 > 1'_IGHJ6*01 652 >gi|Fabrus|B08_VH3-23_IGHD2- 2721 3411 21*01 > 3'_IGHJ6*01 653 >gi|Fabrus|B09_VH3-23_IGHD3- 2722 3412 3*01 > 1'_IGHJ6*01 654 >gi|Fabrus|B10_VH3-23_IGHD3- 2723 3413 3*01 > 3'_IGHJ6*01 655 >gi|Fabrus|B11_VH3-23_IGHD3- 2724 3414 9*01 > 1'_IGHJ6*01 656 >gi|Fabrus|B12_VH3-23_IGHD3- 2725 3415 9*01 > 3'_IGHJ6*01 657 >gi|Fabrus|C01_VH3-23_IGHD3- 2726 3416 10*01 > 1'_IGHJ6*01 658 >gi|Fabrus|C02_VH3-23_IGHD3- 2727 3417 10*01 > 3'_IGHJ6*01 659 >gi|Fabrus|C03_VH3-23_IGHD3- 2728 3418 16*01 > 1'_IGHJ6*01 660 >gi|Fabrus|C04_VH3-23_IGHD3- 2729 3419 16*01 > 3'_IGHJ6*01 661 >gi|Fabrus|C05_VH3-23_IGHD3- 2730 3420 22*01 > 1'_IGHJ6*01 662 >gi|Fabrus|C06_VH3-23_IGHD4- 2731 3421 4*01(1) > 1'_IGHJ6*01 663 >gi|Fabrus|C07_VH3-23_IGHD4- 2732 3422 4*01(1) > 3'_IGHJ6*01 664 >gi|Fabrus|C08_VH3-23_IGHD4- 2733 3423 11*01(1) > 1'_IGHJ6*01 665 >gi|Fabrus|C09_VH3-23_IGHD4- 2734 3424 11*01(1) > 3'_IGHJ6*01 666 >gi|Fabrus|C10_VH3-23_IGHD4- 2735 3425 17*01 > 1'_IGHJ6*01 667 >gi|Fabrus|C11_VH3-23_IGHD4- 2736 3426 17*01 > 3'_IGHJ6*01 668 >gi|Fabrus|C12_VH3-23_IGHD4- 2737 3427 23*01 > 1'_IGHJ6*01 669 >gi|Fabrus|D01_VH3-23_IGHD4- 2738 3428 23*01 > 3'_IGHJ6*01 670 >gi|Fabrus|D02_VH3-23_IGHD5- 2739 3429 5*01(2) > 1'_IGHJ6*01 671 >gi|Fabrus|D03_VH3-23_IGHD5- 2740 3430 5*01(2) > 3'_IGHJ6*01 672 >gi|Fabrus|D04_VH3-23_IGHD5- 2741 3431 12*01 > 1'_IGHJ6*01 673 >gi|Fabrus|D05_VH3-23_IGHD5- 2742 3432 12*01 > 3'_IGHJ6*01 674 >gi|Fabrus|D06_VH3-23_IGHD5- 2743 3433 18*01(2) > 1'_IGHJ6*01 675 >gi|Fabrus|D07_VH3-23_IGHD5- 2744 3434 18*01(2) > 3'_IGHJ6*01 676 >gi|Fabrus|D08_VH3-23_IGHD5- 2745 3435 24*01 > 1'_IGHJ6*01 677 >gi|Fabrus|D09_VH3-23_IGHD5- 2746 3436 24*01 > 3'_IGHJ6*01 678 >gi|Fabrus|D10_VH3-23_IGHD6- 2747 3437 6*01 > 1'_IGHJ6*01 679 >gi|Fabrus|D11_VH3-23_IGHD6- 2748 3438 6*01 > 2'_IGHJ6*01 680 >gi|Fabrus|D12_VH3-23_IGHD6- 2749 3439 6*01 > 3'_IGHJ6*01 681 >gi|Fabrus|E01_VH3-23_IGHD6- 2750 3440 13*01 > 1'_IGHJ6*01 682 >gi|Fabrus|E02_VH3-23_IGHD6- 2751 3441 13*01 > 2'_IGHJ6*01 683 >gi|Fabrus|E03_VH3-23_IGHD6- 2752 3442 13*01 > 3'_IGHJ6*01 684 >gi|Fabrus|E04_VH3-23_IGHD6- 2753 3443 19*01 > 1'_IGHJ6*01 685 >gi|Fabrus|E05_VH3-23_IGHD6- 2754 3444 19*01 > 2'_IGHJ6*01 686 >gi|Fabrus|E06_VH3-23_IGHD6- 2755 3445 19*01 > 3'_IGHJ6*01 687 >gi|Fabrus|E07_VH3-23_IGHD6- 2756 3446 25*01 > 1'_IGHJ6*01 688 >gi|Fabrus|E08_VH3-23_IGHD6- 2757 3447 25*01 > 3'_IGHJ6*01 689 >gi|Fabrus|E09_VH3-23_IGHD7- 2758 3448 27*01 > 1'_IGHJ6*01 690 >gi|Fabrus|E10_VH3-23_IGHD7- 2759 3449 27*01 > 2'_IGHJ6*01
Example 15
Electrochemiluminescence Binding Assay
[1176] The electrochemiluminescence (ECL) binding assay described in Example 13 was further used to screen a 5,376 member Fab library and 15 modified Fab antibodies set forth in rows 22-36 in Table 77 below for antibodies capable of binding to one of the nine different antigens. As noted in Example 13 above, data were analyzed by comparing the ECL signals for an antigen to the blank of each well. A signal to blank ratio of 4 or more was considered a "Hit" Fab.
[1177] Fifty-six (56) plates, each containing 96 different Fabs, were screened using the ECL assay. Thirty-six (36) Fabs were identified with specific binding affinity to one or more of the protein antigens, as indicated in Tables 42-44. Table 42, below, summarizes the results of the ECL assay, including the recombinant human target/protein antigen, the number of antibody hits per target and the % of hits per target (number of hits/5376 antibodies screened). Eleven Fabs were identified that bind to recombinant human delta-like protein 4 (DLL4) whereas only 2 hits were identified that bind to recombinant human epidermal growth factor 2 (ErbB2). Ten Fabs were identified that bind to recombinant human erythropoietin receptor (Epo R) and 6 Fabs were identified that bind to recombinant human P-cadherin. Additionally, 3 Fabs were identified that bind to recombinant human epidermal growth factor 2 (ErbB2) and 3 Fabs were identified that bind to recombinant human Notch-1. Table 43, below, lists the 21 Fabs (including the heavy chain and light chain) that were identified as "hits" in the initial ECL screen. A "hit" was a Fab antibody with signal to blank ratio of greater than 4. The results of the initial MSD assay screen at a single Fab concentration are listed in Table 44 below. Table 44 lists the 21 Fabs (the Fab No. corresponds each of the Fabs identified in Table 43), the Fab concentration, the 9 recombinant human target/protein antigens, and the ECL signals from the initial MSD assay screen at the given Fab concentration.
TABLE-US-00059 TABLE 42 Summary of 5376 Fab Library ECL Screen Number % Per rHuman Antigen of Hits Target ErbB2/Fc chimera 2 0.037 EGF R/Fc chimera 3 0.056 HGF R/Fc chimera 0 n/a Notch-1/Fc chimera 3 0.056 CD44/Fc chimera 0 n/a IGF-1 sR 0 n/a P-cadherin/Fc chimera 6 0.112 Epo R/Fc chimera 10 0.186 DLL4 11 0.205
TABLE-US-00060 TABLE 43 Fab Antibody "Hits" Identified In ECL Screen Fab No. Heavy Chain Light Chain 1 VH4-28_IGHD7-27*01_IGHJ1*01 L2_IGKJ1*01 2 VH4-31_IGHD7-27*01_IGHJ5*01 L2_IGKJ1*01 3 VH2-5_IGHD7-27*01_IGHJ2*01 L2_IGKJ1*01 4 VH1-46_IGHD7-27*01_IGHJ2*01 A27_IGKJ1*01 5 VH1-69_IGHD1-1*01_IGHJ6*01 A17_IGKJ1*01 6 VH1-46_IGHD2-15*01_IGHJ2*01 L2_IGKJ1*01 7 VH1-46_IGHD6-13*01_IGHJ4*01 L2_IGKJ1*01 8 VH4-34_IGHD7-27*01_IGHJ4*01 L5_IGKJ1*01 9 VH1-46_IGHD6-13*01_IGHJ4*01 A27_IGKJ1*01 10 VH1-46_IGHD7-27*01_IGHJ2*01 L6_IGKJ1*01 11 VH1-3_IGHD4-23*01_IGHJ4*01 L12_IGKJ1*01 12 VH1-46_IGHD2-15*01_IGHJ2*01 L12_IGKJ1*01 13 VH1-46_IGHD3-10*01_IGHJ4*01 L12_IGKJ1*01 14 VH1-8_IGHD2-2*01_IGHJ6*01 L12_IGKJ1*01 15 VH1-46_IGHD3-10*01_IGHJ4*01 O1_IGKJ1*01 16 VH1-46_IGHD6-13*01_IGHJ4*01 O1_IGKJ1*01 17 VH4-34_IGHD7-27*01_IGHJ4*01 V1-4_IGLJ4*01 18 VH4-31_IGHD2-15*01_IGHJ2*01 V1-4_IGLJ4*01 19 VH4-34_IGHD7-27*01_IGHJ4*01 V4-6_IGLJ4*01 20 VH3-23_IGHD3-10*01 > 3_IGHJ6*01 O12_IGKJ1*01 21 VH3-23_IGHD3-10*01 > 1'_IGHJ3*01 O12_IGKJ1*01
TABLE-US-00061 TABLE 44 ECL Signals for Identified Fab "Hits" from 5376 Fab Library Screen rHuman Target Fab [Fab] Notch- No. .mu.M ErbB2 EGFR HGFR 1 CD44 IGF-1 P-cad EPOR DLL4 Blank 1 0.36 2028 3803 2034 3813 2482 1703 2741 19375 3911 1948 2 0.61 581 1222 539 1195 1048 596 782 3065 1006 570 3 0.03 2556 381 311 563 421 344 352 492 476 373 4 0.36 1474 1405 1225 1921 1542 1081 1795 2434 9082 928 5 0.65 2940 662 437 1336 797 656 695 922 849 441 6 0.25 15169 88015 5912 10346 10436 4929 16145 30118 14728 3693 7 1.19 18514 32030 17322 41739 27596 14089 34551 141492 39645 11543 8 0.51 1705 1807 1738 2876 1879 1742 3176 2563 5648 1393 9 0.48 6940 9043 6313 12759 9979 5468 15301 67194 13726 2877 10 1.56 2452 2497 3116 3953 2281 1919 5418 4129 3770 1129 11 0.06 1569 1755 1910 2232 1683 2325 2432 1973 13614 842 12 0.28 4314 8327 5182 5281 4561 4643 8057 8071 8151 1773 13 0.93 10795 15559 13790 23657 10634 13930 26366 16729 74294 4760 14 0.13 840 859 969 1324 932 1167 1641 1106 9848 786 15 1.57 904 1403 1039 1256 836 985 2029 6393 1496 797 16 0.93 1610 1855 2596 1918 1453 1787 2422 4241 9632 859 17 0.58 1262 1022 1706 739 761 1198 725 768 3128 403 18 0.21 1308 1297 1531 809 868 1557 915 885 3332 489 19 0.34 900 738 914 912 694 871 935 907 2561 356 20 0.12 4578 4060 5472 5501 4213 3137 17528 11210 6646 2225 21 0.42 702 759 935 919 748 844 3178 1504 922 450
[1178] To confirm a "Hit" identified in the initial ECL screening (see Tables 42-44 above), a Fab concentration dependent titration was carried out to determine the Fab-antigen binding affinity. The assay procedure was the same as described Example 13 above, except that the concentration of Fab antibody was varied between wells from 0.0628 nM to 1.57 NM. The results of the dose response assays are set forth in Tables 45-61 below. Tables 62-76 below list the results of the dose response assays for 15 modified anti-DLL4 antibodies (set forth in rows 22-36 of Table 77, below). The modified anti-DLL4 antibodies have at least one mutation in the heavy or light chain as compared to the previously identified germline antibodies.
TABLE-US-00062 TABLE 45 Binding affinity of Fab VH4-28_IGHD7-27*01_IGHJ1*01 & L2_IGKJ1*01 Fab[nM] 360 36 ErbB2/Fc 647 600 EGF R/Fc 957 711 HGF R/Fc 581 613 Notch-1/Fc 1026 773 CD44/FC 740 679 IGF-1 sR 535 486 P-Cadherin/Fc 636 693 EPO R/Fc 4715 2977 DLL4 866 799 Blank 462 413
TABLE-US-00063 TABLE 46 Binding affinity of Fab VH1-46_IGHD2-15*01_IGHJ2*01 & L2_IGKJ1*01 Fab[.mu.M] 0.25 0.0625 0.01563 0.00391 ErbB2/Fc 29608 9033 4495 1667 EGF R/Fc 116674 94778 70836 35936 HGF R/Fc 13427 4108 1998 913 Notch-1/Fc 21447 5848 2800 1282 CD44/Fc 23015 6746 3182 1295 IGF-1 sR 11050 3150 1742 822 P-Cadherin/Fc 25459 7739 4945 1962 EPO R/Fc 49177 21136 11342 5022 DLL4 27691 8051 4015 1551 Blank 6344 1738 906 576
TABLE-US-00064 TABLE 47 Binding affinity of Fab VH1-46_IGHD6- 13*01_IGHJ4*01 & L2_IGKJ1*01 Fab[.mu.M] 1.19 0.2975 0.07438 0.01859 ErbB2/Fc 38410 15111 7551 5531 EGF R/Fc 62454 42213 16605 11750 HGF R/Fc 45494 17396 6611 4566 Notch-1/Fc 72018 37503 21990 17565 CD44/Fc 47145 28601 10922 7322 IGF-1 sR 35187 17389 5804 3779 P-Cadherin/Fc 69710 26043 14807 11672 EPO R/Fc 192967 167064 153692 188065 DLL4 74900 34726 20719 18888 Blank 24999 5019 2504 1776
TABLE-US-00065 TABLE 48 Binding affinity of Fab VH4-34_IGHD7- 27*01_IGHJ4*01 & L5_IGKJ1*01 Fab[.mu.M] 0.51 0.1275 0.03188 0.00797 ErbB2/Fc 1532 857 584 493 EGF R/Fc 2363 1061 694 530 HGF R/Fc 1989 853 693 419 Notch-1/Fc 2773 1497 849 654 CD44/Fc 2012 926 653 490 IGF-1 sR 2236 1045 765 564 P-Cadherin/Fc 2389 957 775 502 EPO R/Fc 2624 1067 789 566 DLL4 5183 2382 1282 872 Blank 1096 530 536 364
TABLE-US-00066 TABLE 49 Binding affinity of Fab VH1-46_IGHD6- 13*01_IGHJ4*01 & A27_IGKJ1*01 Fab[.mu.M] 0.48 0.096 0.0192 ErbB2/Fc 11287 3365 2313 EGF R/Fc 14638 4509 3115 HGF R/Fc 8002 2328 1582 Notch-1/Fc 15931 4802 3041 CD44/Fc 13445 4320 2915 IGF-1 sR 8927 2449 1826 P-Cadherin/Fc 15595 6654 5040 EPO R/Fc 70938 57356 62037 DLL4 16065 5586 3555 Blank 2945 917 751
TABLE-US-00067 TABLE 50 Binding affinity of Fab VH1-46_IGHD7- 27*01_IGHJ2*01 & L6_IGKJ1*01 Fab[.mu.M] 1.56 0.312 0.0624 ErbB2/Fc 7577 3659 2146 EGF R/Fc 7832 4328 2415 HGF R/Fc 10267 4691 2453 Notch-1/Fc 9447 4462 2352 CD44/Fc 7595 4171 2110 IGF-1 sR 6913 3508 2034 P-Cadherin/Fc 15016 7098 4226 EPO R/Fc 9480 5020 2678 DLL4 10897 5484 2585 Blank 4357 1977 960
TABLE-US-00068 TABLE 51 Binding affinity of Fab VH1-3_IGHD4- 23*01_IGHJ4*01 & L12_IGKJ1*01 Fab[nM] 60 15 3.75 0.9375 ErbB2/Fc 2155 740 291 268 EGF R/Fc 2563 842 371 224 HGF R/Fc 2298 743 394 243 Notch-1/Fc 2886 1058 375 348 CD44/Fc 2355 748 307 251 IGF-1 sR 2666 859 314 204 P-Cadherin/Fc 2662 837 331 191 EPO R/Fc 3214 970 358 238 DLL4 17270 7728 1569 453 Blank 1433 536 191 153
TABLE-US-00069 TABLE 52 Binding affinity of Fab VH1-46_IGHD2- 15*01_IGHJ2*01 & L12_IGKJ1*01 Fab[nM] 280 70 17.5 4.375 ErbB2/Fc 3953 1358 541 384 EGF R/Fc 6667 2574 1305 542 HGF R/Fc 3564 1289 565 193 Notch-1/Fc 4382 1492 680 480 CD44/Fc 4069 1370 664 424 IGF-1 sR 3533 1319 626 369 P-Cadherin/Fc 5400 1817 949 469 EPO R/Fc 8496 2485 1262 594 DLL4 8111 2747 1219 558 Blank 1691 635 304 305
TABLE-US-00070 TABLE 53 Binding affinity of Fab VH1-46_IGHD3- 10*01_IGHJ4*01 & L12_IGKJ1*01 Fab[nM] 920 230 57.5 14.375 ErbB2/Fc 10924 4078 2447 1594 EGF R/Fc 13406 5723 3858 2672 HGF R/Fc 10708 3934 2297 1600 Notch-1/Fc 20086 9737 5886 4206 CD44/Fc 9698 3817 2313 1488 IGF-1 sR 10246 4764 2833 1746 P-Cadherin/Fc 16666 6484 4110 2318 EPO R/Fc 16429 6949 4038 2718 DLL4 73638 119436 144126 125422 Blank 4082 1656 954 738
TABLE-US-00071 TABLE 54 Binding affinity of Fab VH1-8_IGHD2- 2*01_IGHJ6*01 & L12_IGKJ1*01 Fab[nM] 130 32.5 8.1 2.0 ErbB2/Fc 1533 556 557 382 EGF R/Fc 1746 645 560 424 HGF R/Fc 1882 525 551 356 Notch-1/Fc 1759 706 612 539 CD44/Fc 1754 573 528 447 IGF-1 sR 1973 561 518 367 P-Cadherin/Fc 1845 556 573 250 EPO R/Fc 2151 673 660 433 DLL4 7738 2989 1548 605 Blank 1153 473 435 316
TABLE-US-00072 TABLE 55 Binding affinity of Fab FabVH1-46_IGHD3- 10*01_IGHJ4*01 & O1_IGKJ1*01 Fab[nM] 1570 392.5 98.1 24.5 ErbB2/Fc 1263 539 247 241 EGF R/Fc 2481 744 4386 317 HGF R/Fc 1638 581 335 211 Notch-1/Fc 1639 749 313 434 CD44/Fc 1381 498 265 267 IGF-1 sR 1428 466 309 239 P-Cadherin/Fc 1793 459 347 257 EPO R/Fc 6121 5863 5628 4531 DLL4 2701 735 402 339 Blank 866 338 210 149
TABLE-US-00073 TABLE 56 Binding affinity of Fab VH1-46_IGHD6- 13*01_IGHJ4*01 & O1_IGKJ1*01 Fab[nM] 930 232.5 58.1 14.5 ErbB2/Fc 2225 779 322 274 EGF R/Fc 3110 803 444 357 HGF R/Fc 2344 790 432 373 Notch-1/Fc 2206 778 388 317 CD44/Fc 1917 607 375 212 IGF-1 sR 1915 569 343 234 P-Cadherin/Fc 2438 655 478 277 EPO R/Fc 3009 1472 829 660 DLL4 8162 3586 1876 1149 Blank 1206 460 225 117
TABLE-US-00074 TABLE 57 Binding affinity of Fab VH4-34_IGHD7- 27*01_IGHJ4*01 & V1-4_IGLJ4*01 Fab[nM] 580 145 36.3 9.1 ErbB2/Fc 1712 1123 1029 987 EGF R/Fc 1631 856 831 800 HGF R/Fc 2341 1173 1065 894 Notch-1/Fc 1585 860 633 754 CD44/Fc 1228 692 629 607 IGF-1 sR 1364 794 799 788 P-Cadherin/Fc 2240 850 684 589 EPO R/Fc 1579 845 722 697 DLL4 4420 2140 1399 1030 Blank 679 357 314 276
TABLE-US-00075 TABLE 58 Binding affinity of Fab VH4-31_IGHD2- 15*01_IGHJ2*01 & V1-4_IGLJ4*01 Fab[nM] 210 52.5 13.1 3.3 ErbB2/Fc 1977 1511 930 1031 EGF R/Fc 1617 1109 824 847 HGF R/Fc 2060 1286 981 849 Notch-1/Fc 1972 1323 669 726 CD44/Fc 1395 897 708 621 IGF-1 sR 1431 911 814 743 P-Cadherin/Fc 4410 2161 1062 678 EPO R/Fc 2123 1319 776 695 DLL4 4108 1951 1107 922 Blank 833 467 376 359
TABLE-US-00076 TABLE 59 Binding affinity of Fab VH4-34_IGHD7- 27*01_IGHJ4*01 & V4-6_IGLJ4*01 Fab[nM] 340 170 85.0 42.5 ErbB2/Fc 1226 964 844 866 EGF R/Fc 1208 826 1001 528 HGF R/Fc 1238 757 998 607 Notch-1/Fc 1209 816 780 649 CD44/Fc 959 660 693 522 IGF-1 sR 1042 832 891 646 P-Cadherin/Fc 1160 744 709 421 EPO R/Fc 1255 790 817 494 DLL4 2332 1462 1311 877 Blank 554 262 292 162
TABLE-US-00077 TABLE 60 Binding affinity of Fab VH3-23_IGHD3- 10*01 > 3_IGHJ6*01 & O12_IGKJ1*01 Fab[nM] 120 12 1.2 0.12 ErbB2/Fc 17294 4358 677 287 EGF R/Fc 14925 1984 464 272 HGF R/Fc 15917 2703 412 287 Notch-1/Fc 14382 2582 660 218 CD44/Fc 13519 1321 341 291 IGF-1 sR 13265 1135 181 175 P-Cadherin/Fc 61714 28490 1684 318 EPO R/Fc 33268 10966 1014 260 DLL4 20627 2510 319 210 Blank 6749 573 227 264
TABLE-US-00078 TABLE 61 Binding affinity of Fab VH3-23_IGHD3- 10*01 > 1'_IGHJ3*01 & O12_IGKJ1*01 Fab[nM] 421.12 42.112 ErbB2/Fc 868 524 EGF R/Fc 765 422 HGF R/Fc 1202 565 Notch-1/Fc 1061 437 CD44/Fc 903 360 IGF-1 sR 1065 364 P-Cadherin/Fc 2949 1546 EPO R/Fc 1299 759 DLL4 1090 404 Blank 639 323
TABLE-US-00079 TABLE 62 Binding affinity of Fab VH5-51_IGHD5- 18*01 > 3_IGHJ4*01_G100K & V3-4_IGLJ1*01 Fab[nM] 1000 200 40 8 1.6 ErbB2/Fc 1251 1467 1394 1232 1320 EGF R/Fc 1029 1371 1149 1033 1180 HGF R/Fc 1199 1428 1306 1260 1438 Notch-1/Fc 1176 1247 875 831 816 CD44/Fc 1025 1162 857 853 938 IGF-1 sR 1134 1320 1271 1203 1437 P-Cadherin/Fc 1043 1126 744 788 811 EPO R/Fc 1122 1226 979 881 899 DLL4 29554 45421 35876 11408 2423 Blank 630 588 518 489 565
TABLE-US-00080 TABLE 63 Binding affinity of Fab VH5-51_IGHD5-18*01 > 3_IGHJ4*01_G100R & V3-4_IGLJ1*01 Fab [nM] 1000 200 40 8 1.6 0.32 ErbB2/Fc 945 1448 1421 1326 1439 1485 EGF R/Fc 721 1017 1113 1078 1204 1221 HGF R/Fc 778 1164 1234 1240 1206 1287 Notch-1/Fc 761 1330 1014 916 1052 983 CD44/Fc 559 895 824 786 997 877 IGF-1 sR 733 1110 1243 1107 1329 1253 P-Cadherin/Fc 762 815 1020 669 666 656 EPO R/Fc 666 1175 994 911 920 851 DLL4 23398 29772 19481 5472 1541 1212 Blank 427 529 502 479 449 513
TABLE-US-00081 TABLE 64 Binding affinity of Fab VH1-46_IGHD6-6*01 > 1_IGHJ1*01_S104F & L6_IGKJ1*01 Fab [nM] 1000 200 40 8 1.6 0.32 ErbB2/Fc 7472 5434 3428 1775 1008 611 EGF R/Fc 3575 2921 2297 1473 987 709 HGF R/Fc 3314 2836 2110 1619 861 710 Notch-1/Fc 9798 6377 3990 2003 1114 684 CD44/Fc 5105 4071 2711 1523 1016 679 IGF-1 sR 3074 2778 2141 1517 966 630 P-Cadherin/Fc 5955 6263 5103 2634 1369 750 EPO R/Fc 5162 4593 3233 2040 1132 673 DLL4 35253 58743 62035 44781 5920 916 Blank 2186 1787 1442 1117 713 518
TABLE-US-00082 TABLE 65 Binding affinity of Fab VH1-46_IGHD6-6*01_IGHJ1*01_S104A & L6_IGKJ1*01 Fab [nM] 1000 200 40 8 1.6 0.32 ErbB2/Fc 7419 4038 2231 1902 1168 766 EGF R/Fc 3481 2951 1843 1298 862 660 HGF R/Fc 3531 2443 1545 1305 830 574 Notch-1/Fc 8058 5705 3479 2196 1166 739 CD44/Fc 4734 3189 2677 1487 1017 684 IGF-1 sR 3348 2603 1897 1287 815 611 P-Cadherin/Fc 5632 3993 2602 1916 1139 649 EPO R/Fc 8292 3997 3510 1982 1117 739 DLL4 14331 14838 13637 9329 2130 646 Blank 2142 1659 1229 1188 648 428
TABLE-US-00083 TABLE 66 Binding affinity of Fab VH5-51_IGHD5- 18*01 > 3_IGHJ4*01_G104T & V3-4_IGLJ1*01 Fab[nM] 1000 10 ErbB2/Fc 3532 1517 EGF R/Fc 3937 1188 HGF R/Fc 3769 1392 Notch-1/Fc 3342 933 CD44/Fc 3378 841 IGF-1 sR 3992 1118 P-Cadherin/Fc 3426 705 EPO R/Fc 3376 910 DLL4 29776 5229 Blank 1742 427
TABLE-US-00084 TABLE 67 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S103P & L6_IGKJ1*01 Fab[nM] 1000 10 ErbB2/Fc 1276 847 EGF R/Fc 1909 773 HGF R/Fc 1359 859 Notch-1/Fc 1210 792 CD44/Fc 1220 746 IGF-1 sR 1336 814 P-Cadherin/Fc 1376 846 EPO R/Fc 1511 802 DLL4 29752 55127 Blank 760 550
TABLE-US-00085 TABLE 68 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A & L6_IGKJ1*01 Fab[nM] 1000 10 ErbB2/Fc 946 670 EGF R/Fc 1601 677 HGF R/Fc 1658 698 Notch-1/Fc 1253 647 CD44/Fc 1087 681 IGF-1 sR 1167 657 P-Cadherin/Fc 1502 691 EPO R/Fc 1320 761 DLL4 22225 38724 Blank 1121 418
TABLE-US-00086 TABLE 69 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01 Fab[nM] 1000 10 ErbB2/Fc 1292 842 EGF R/Fc 2434 809 HGF R/Fc 1581 691 Notch-1/Fc 1620 1144 CD44/Fc 1314 887 IGF-1 sR 1175 720 P-Cadherin/Fc 2499 894 EPO R/Fc 1656 1109 DLL4 73307 345030 Blank 731 680
TABLE-US-00087 TABLE 70 Binding affinity of Fab VH5-51_IGHD5- 18*01 > 3_IGHJ4*01_G100K_G104T & V34_IGLJ1*01 Fab[nM] 500 25 1.25 0.0625 ErbB2/Fc 957 927 695 692 EGF R/Fc 1106 821 729 736 HGF R/Fc 1299 805 774 643 Notch-1/Fc 703 440 312 439 CD44/Fc 923 570 444 394 IGF-1 sR 1114 780 674 537 P-Cadherin/Fc 815 269 374 331 EPO R/Fc 828 597 448 332 DLL4 25941 77931 38837 1014 Blank 480 206 256 239
TABLE-US-00088 TABLE 71 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104F_H111F & L6_IGKJ1*01 Fab[nM] 500 50 5 ErbB2/Fc 2621 2114 1377 EGF R/Fc 2514 2042 1158 HGF R/Fc 2043 1282 1016 Notch-1/Fc 4376 2726 1170 CD44/Fc 2912 1610 1132 IGF-1 sR 1872 1226 917 P-Cadherin/Fc 3665 2491 1663 EPO R/Fc 5518 2462 1835 DLL4 29919 94200 149972 Blank 1016 774 626
TABLE-US-00089 TABLE 72 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104F_H111Y & L6_IGKJ1*01 Fab[nM] 500 50 5 0.5 ErbB2/Fc 713 589 315 331 EGF R/Fc 898 523 311 273 HGF R/Fc 523 325 323 319 Notch-1/Fc 644 475 383 263 CD44/Fc 664 433 413 279 IGF-1 sR 592 460 320 164 P-Cadherin/Fc 939 626 437 215 EPO R/Fc 1212 631 377 359 DLL4 93071 206919 84945 2971 Blank 410 433 367 365
TABLE-US-00090 TABLE 73 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104Y_H111Y & L6_IGKJ1*01 Fab[nM] 500 50 5 0.5 ErbB2/Fc 596 470 2855 1721 EGF R/Fc 758 489 2980 1751 HGF R/Fc 841 312 3101 1688 Notch-1/Fc 689 389 3232 1756 CD44/Fc 604 345 2756 1563 IGF-1 sR 747 399 3033 1662 P-Cadherin/Fc 1195 470 4162 2628 EPO R/Fc 1273 533 3582 1840 DLL4 105743 201050 73428 5150 Blank 436 329 2104 1309
TABLE-US-00091 TABLE 74 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01_S28P Fab[nM] 50 10 2 0.4 ErbB2/Fc 384 136 176 275 EGF R/Fc 328 207 352 246 HGF R/Fc 183 136 255 199 Notch-1/Fc 477 280 307 248 CD44/Fc 340 212 266 257 IGF-1 sR 355 152 205 172 P-Cadherin/Fc 431 216 223 191 EPO R/Fc 527 392 251 147 DLL4 37653 26851 6103 1140 Blank 274 146 130 138
TABLE-US-00092 TABLE 75 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01_S30N Fab[nM] 50 10 2 0.4 ErbB2/Fc 395 274 138 203 EGF R/Fc 294 245 171 173 HGF R/Fc 135 239 207 120 Notch-1/Fc 279 253 274 197 CD44/Fc 271 286 209 176 IGF-1 sR 240 233 296 141 P-Cadherin/Fc 143 283 395 111 EPO R/Fc 500 420 294 224 DLL4 37208 33374 11375 1039 Blank 299 165 205 179
TABLE-US-00093 TABLE 76 Binding affinity of Fab VH1-46_IGHD6- 6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01_S31K Fab[nM] 50 10 2 0.4 ErbB2/Fc 397 303 337 159 EGF R/Fc 350 297 197 175 HGF R/Fc 257 271 183 119 Notch-1/Fc 355 388 202 197 CD44/Fc 262 352 146 190 IGF-1 sR 257 214 222 65 P-Cadherin/Fc 411 240 351 151 EPO R/Fc 574 392 326 217 DLL4 35274 29781 10534 1259 Blank 183 267 197 255
[1179] Summary
[1180] Table 77, below, summarizes the results of the MSD assay. Table 77 lists the recombinant human target/protein antigen(s) and the Fabs, as designated by their respective heavy and light chains (including SEQ ID NOS). As is indicated in Table 77, below, several Fabs were identified that bind to multiple targets. For example, Fab VH1-46_IGHD2-15*01_IGHJ2*01 & L12_IGKJ1*01 binds to EGF R, Epo R and DLL4 while Fab VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01 binds to Notch-1, P-cadherin and DLL4. Table 77 below also lists the 15 additional modified Fabs (set forth in rows 22-36) that bind to DLL4.
TABLE-US-00094 TABLE 77 Selected Fabs and their targets Fab SEQ ID SEQ ID No. rHuman Target Heavy Chain NO Light Chain NO 1 Epo R/Fc chimera VH4-28_IGHD7-27*01_IGHJ1*01 1759 L2_IGKJ1*01 1843 2 Epo R/Fc chimera VH4-31_IGHD7-27*01_IGHJ5*01 1769 L2_IGKJ1*01 1843 3 ErbB2/Fc chimera VH2-5_IGHD7-27*01_IGHJ2*01 1559 L2_IGKJ1*01 1843 4 Epo R/Fc chimera VH1-46_IGHD7-27*01_IGHJ2*01 1514 A27_IGKJ1*01 1833 5 ErbB2/Fc chimera VH1-69_IGHD1-1*01_IGHJ6*01 1522 A17_IGKJ1*01 1828 6 Epo R/Fc chimera and VH1-46_IGHD2-15*01_IGHJ2*01 1508 L2_IGKJ1*01 1843 EGF R/Fc chimera 7 EGF R/Fc chimera, Notch- VH1-46_IGHD6-13*01_IGHJ4*01 1512 L2_IGKJ1*01 1843 1/Fc chimera, P-cadherin/ Fc chimera, Epo R/Fc chimera and DLL4 8 DLL4 VH4-34_IGHD7-27*01_IGHJ4*01 1779 L5_IGKJ1*01 1849 9 Notch-1/Fc chimera, P- VH1-46_IGHD6-13*01_IGHJ4*01 1512 A27_IGKJ1*01 1833 cadherin/Fc chimera, Epo R/Fc chimera and DLL4 10 P-cadherin/Fc chimera VH1-46_IGHD7-27*01_IGHJ2*01 1514 L6_IGKJ1*01 1850 11 DLL4 VH1-3_IGHD4-23*01_IGHJ4*01 1494 L12_IGKJ1*01 1841 12 EGF R/Fc chimera, Epo VH1-46_IGHD2-15*01_IGHJ2*01 1508 L12_IGKJ1*01 1841 R/Fc chimera and DLL4 13 Notch-1/Fc chimera, P- VH1-46_IGHD3-10*01_IGHJ4*01 1509 L12_IGKJ1*01 1841 cadherin/Fc chimera and DLL4 14 DLL4 VH1-8_IGHD2-2*01_IGHJ6*01 1537 L12_IGKJ1*01 1841 15 Epo R/Fc chimera VH1-46_IGHD3-10*01_IGHJ4*01 1509 O1_IGKJ1*01 1853 16 Epo R/Fc chimera and VH1-46_IGHD6-13*01_IGHJ4*01 1512 O1_IGKJ1*01 1853 DLL4 17 DLL4 VH4-34_IGHD7-27*01_IGHJ4*01 1779 V1-4_IGLJ4*01 1864 18 DLL4 VH4-31_IGHD2-15*01_IGHJ2*01 1761 V1-4_IGLJ4*01 1864 19 DLL4 VH4-34_IGHD7-27*01_IGHJ4*01 1779 V4-6_IGLJ4*01 1886 20 P-cadherin/Fc chimera and VH3-23_IGHD3-10*01>3_IGHJ6*01 3359 O12_IGKJ1*01 1854 Epo R/Fc chimera 21 P-cadherin/Fc chimera VH3-23_IGHD3-10*01 > 1'_IGHJ3*01 3071 O12_IGKJ1*01 1854 22 DLL4 VH5-51_IGHD5-18*01 > 3_IGHJ4*01_G100K 3720 V3-4_IGLJ1*01 1881 23 DLL4 VH5-51_IGHD5-18*01 > 3_IGHJ4*01_G100R 3721 V3-4_IGLJ1*01 1881 24 DLL4 VH1-46_IGHD6-6*01 > 1_IGHJ1*01_S104F 3722 L6_IGKJ1*01 1850 25 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S104A 3723 L6_IGKJ1*01 1850 26 DLL4 VH5-51_IGHD5-18*01 > 3_IGHJ4*01_G104T 3724 V3-4_IGLJ1*01 1881 27 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S103P 3725 L6_IGKJ1*01 1850 28 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A 3726 L6_IGKJ1*01 1850 29 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F 3727 L6_IGKJ1*01 1850 30 DLL4 VH5-51_IGHD5-18*01 > 3_IGHJ4*01_G100K_G104T 3728 V3-4_IGLJ1*01 1881 31 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111F 3729 L6_IGKJ1*01 1850 32 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111Y 3730 L6_IGKJ1*01 1850 33 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104Y_H111Y 3731 L6_IGKJ1*01 1850 34 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F 3727 L6_IGKJ1*01_S28P 3732 35 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F 3727 L6_IGKJ1*01_S30N 3733 36 DLL4 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F 3727 L6_IGKJ1*01_S31K 3734
Example 16
Surface Plasmon Resonance
[1181] In this example, the binding affinities of selected Fabs (see Tables 78-79) to recombinant human DLL4 (R&D Systems) were analyzed using Surface Plasmon Resonance (SPR) (Biosensor Tools, Salt Lake City, Utah). The Fabs (see Table 78) include germline antibodies identified in the initial ECL screen as binding to DLL4 (as shown in Example 13) and modified Fabs that contain one or more mutations in the heavy or light chain as compared to the initially identified anti-DLL4 Fabs.
[1182] The results are shown in Table 79 below. Table 79 lists Fab (by Fab No.), the k.sub.a (M.sup.-1s.sup.-1), the k.sub.d (s.sup.-1), and the K.sub.D (nM) and the standard deviation (in parentheses). The results indicate that the Fabs have binding affinity for DLL4 ranging from 48.5 nM to 38 uM. Germline Fab VH5-51_IGHD5-18*01>3_IGHJ4*01 & V3-4_IGLJ1*01 has an average K.sub.D of 4.8 uM while variant Fab VH5-51_IGHD5-18*01>3_IGHJ4*01_G100K_G104T & V3-4_IGLJ1*01 has an improved K.sub.D of 355 nM. Germline Fab VH1-46_IGHD6-6*01_IGHJ1*01 & L6_IGKJ1*01 binds DLL4 with an average K.sub.D of 730 nM while the variant Fabs (rows 5-6 and 8-10 of Tables 78 and 79 below) bind DLL4 with improved K.sub.Ds ranging from 70.6 nM to 388 nM. Germline Fab VH6-1_IGHD3-3*01_IGHJ4*01 & V4-3_IGLJ4*01 has an average binding affinity of 38 uM while germline Fab VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01 has an average K.sub.D of 500 nM.
TABLE-US-00095 TABLE 78 DLL4 Fabs for Surface Plasmon Resonance Fab No Heavy Chain Light Chain 1 VH5-51_IGHD5-18*01 > 3_IGHJ4*01 V3-4_IGLJ1*01 2 VH1-46_IGHD6-6*01_IGHJ1*01 L6_IGKJ1*01 3 VH6-1_IGHD3-3*01_IGHJ4*01 V4-3_IGLJ4*01 4 VH1-46_IGHD3-10*01_IGHJ4*01 L12_IGKJ1*01 5 VH1-46_IGHD6-6*01 > 1_IGHJ1*01_S104F L6_IGKJ1*01 6 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F L6_IGKJ1*01 7 VH5-51_IGHD5-18*01 > 3_IGHJ4*01_G100K_G104T V3-4_IGLJ1*01 8 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111F L6_IGKJ1*01 9 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111Y L6_IGKJ1*01 10 VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F L6_IGKJ1*01_S31K
TABLE-US-00096 TABLE 79 Binding affinity of DLL4 Fabs Fab No k.sub.a (M.sup.-1s.sup.-1) k.sub.d (s.sup.-1) K.sub.D (nM) 1 n/a n/a 4800(200) 2 1.63(3)e5 0.101(2) 730(130) 3 n/a n/a 38000(4000) 4 5(1)e5 0.29(2) 500(100) 5 5.0(8)e5 0.19(1) 380(60) 6 4.05(5)e5 0.0492(4) 122(1) 7 0.645(0.92)e5 0.023 355(7) 8 4.25(4)e5 0.0300(2) 70.6(7) 9 3.40(3)e5 0.0317(2) 93.1(9) 10 3.50(5)e5 0.0392(4) 112(2)
Example 17
Inhibition of DLL4-Notch Interaction
[1183] In this example, four Fabs previously identified as binding to DLL4 were functionally screened for their ability to block the binding of Notch-Fc to DLL4.
In this ELISA assay, recombinant human DLL4 bound to the plate followed by the addition of both the Fab and Notch-Fc. An anti-human FC-HRP conjugated antibody was used as a detection molecule therefore if Notch-Fc binds to DLL4, a strong signal will be observed at A450. Alternatively, if the Fab is capable of blocking the bind of Notch-Fc to DLL4, no signal should be observed. The Fabs that were assayed included Fab VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01, Fab VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111F & L6_IGKJ1*01, Fab VH5-51_IGHD5-18*01>3_IGHJ4*01_G100K_G104T &V3-4_1 GLJ1*01 and Fab VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01.
[1184] In short, Maxisorp Nunc 96-well plates were coated with 0.5 pg/ml recombinant human DLL4 extracellular domain (R&D Systems) for at least 2 hours. The wells were washed and then blocked with 4% BSA. Following blocking, Fabs VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01, VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111F & L6_IGKJ1*01, VH5-51_IGHD5-18*01>3_IGHJ4*01_G100K_G104T &V3-4_1 GLJ1*01 and VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01 at concentrations from 0.004 and 5 .mu.M were added together with recombinant human Fc-Notch extracellular domain (R&D Systems) at a concentration of 10 nM. After incubation for one to two hours, the wells were washed and Notch binding was measure using a mouse anti-human FC-HRP conjugated antibody (Southern Biotech) at an 1:1000 dilution. HRP activity was detected using TMB substrate (Pierce) followed by acid neutralization. The A450 was measured on a SpectraMax Plus 384.
[1185] Results show that the addition of Fabs VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01, VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111F & L6_IGKJ1*01 or VH5-51_IGHD5-18*01>3_IGHJ4*01_G100K_G104T & V3-4_IGLJ1*01 resulted in a decreased signal therefore indicating their ability to block the binding of Notch-Fc to DLL4. The addition of Fab VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01 did not result an any loss of activity, indicating that Fab VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01 does not block the Notch-DLL4 interaction. This result also indicates that Fabs VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F & L6_IGKJ1*01, VH1-46_IGHD6-6*01_IGHJ1*01_S102A_S103P_S104F_H111F & L6_IGKJ1*01 or VH5-51_IGHD5-18*01>3_IGHJ4*01_G100K_G104T & V3-4_IGLJ1*01 bind different epitopes of DLL4 than Fab VH1-46_IGHD3-10*01_IGHJ4*01 & L12_IGKJ1*01.
Example 18
EpoR Cell Based Assay for Receptor Stimulation
[1186] In this example, Fabs were analyzed for their ability to stimulate the erythropoietin receptor using a cell based assay. The cell lines used included Ba/F3 cells transfected with the human erythropoietin receptor (EpoR) and parental Ba/F3 cells lacking EpoR. The parental Ba/F3 cells do not respond to receptor agonist and both cell lines required IL-3 for growth.
[1187] In short, Ba/F3 cells (with and without EpoR) were propagated in RPMI 1640 media with 10% FBS, antibiotics, and 5 ng/L recombinant mouse IL-3, washed into equivalent media lacking IL-3 and plated into 96 well plates at 5000 cells/well in 50 .mu.l. Following plating, cells were treated with 10 .mu.l of Fabs, agonist control EMP16 (TYSCHFGPLTWVCKPQ, SEQ ID NO:3735), or vehicle, and grown for 4 days at 37.degree. C. in humid 5% CO.sub.2 atmosphere. To measure cell viability and proliferation, a resazurin-based viability assay reagent was added to test wells for 24 hours. Reduction of the reagent by metabolically active cells produced the readily-quantifiable fluorescent molecule resorufine. Average fluorescence for each treatment was divided by the average fluorescence of vehicle controls to give fold proliferation.
[1188] The results show that 40 nM Fab VH1-46_IGHD2-15*01_IGHJ2*01 & L2_IGKJ1*01 showed receptor dependent proliferation, while 54 nM Fab VH1-46_IGHD6-13*01_IGHJ4*01 & O1_IGKJ1*01 showed little proliferation over vehicle in both receptor-expressing cells and parental cells. The known receptor agonist peptide EMP16 (TYSCHFGPLTWVCKPQ, SEQ ID NO:3735), added at a concentration of 2.5 .mu.M, showed strong cell proliferation.
Example 19
Inhibition of DLL4/Jag1
[1189] In this example, a cellular assay is described in which activation of the Notch pathway prevents C2C12 myoblast cell differentiation (see e.g., Jarriault et al., 1998 Molecular and Cellular Biology, 18:7423-7431). In order to activate the Notch pathway, Notch ligands, such as DLL4 or Jag1, must be expressed as full-length proteins on the cell surface. To achieve this Notch activation, non-adherent cells naturally or ectopically expressing Notch ligands DLL4 or Jag1 are co-cultured with C2C12 cells and selected Fabs. Functional inhibition of DLL4 or Jag1 is assessed by the ability of the Fab to promote differentiation, indicating Notch pathway inactivation. The differentiation into tube-like structures is easily discernible morphologically and additionally can be detected with an antibody against troponin t (Sigma-Aldrich).
[1190] In short, C2C12 mouse myoblast cells are cultured in the presence and absence of Jag1-expressing IM9 cells (a human lymphoblast cell line) and Fabs. The cells are plated onto glass coverslips in 12 well dishes in DMEM containing 10% FBS (fetal bovine serum). The next day attached C2C12 cells are transferred into DMEM containing 1% FBS to induce differentiation. Following incubation, the cells are visualized to observe whether differentiation into myotubes occurred. Low serum conditions will induce the differentiation of myotubes while Jag1-expressing IM9 cells maintain C2C12 cells in an undifferentiated state in low serum conditions.
Example 20
Inhibition of p-Cadherin
[1191] In this example, a cellular assay is described in which the ability of an antibody to inhibit P-cadherin is observed by the failure of cells to "clump". P-cadherin is involved in cell-to-cell adhesion and therefore inhibition of P-cadherin leads to cell scattering.
[1192] In short, A431 epidermoid carcinoma cells are plated at 10,000 cells/well (96-well in DMEM with 10% FBS into 96-well dishes. The next day, Fab is added to wells at 100 g/ml. Function blocking p-cadherin antibody (Abcam) is used as a positive control and DMEM with 10% FBS alone is used as a negative control (see e.g. Shimoyama, Y. et al., 1989 Cancer Research, 49:2128-2133). After 3.5 hours cells are examined for "scattering" and photographed. Cells that are incubated with media only exhibit significant "clumping" while cells that are incubated with an anti-P-cadherin Fab or antibody are scattered.
[1193] Since modifications will be apparent to those of skill in this art, it is intended that this invention be limited only by the scope of the appended claims.
Sequence CWU 0 SQTB SEQUENCE LISTING The patent application
contains a lengthy "Sequence Listing" section. A copy of the
"Sequence Listing" is available in electronic form from the USPTO
web site
(https://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200399359A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
0 SQTB SEQUENCE LISTING The patent application contains a lengthy
"Sequence Listing" section. A copy of the "Sequence Listing" is
available in electronic form from the USPTO web site
(https://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200399359A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
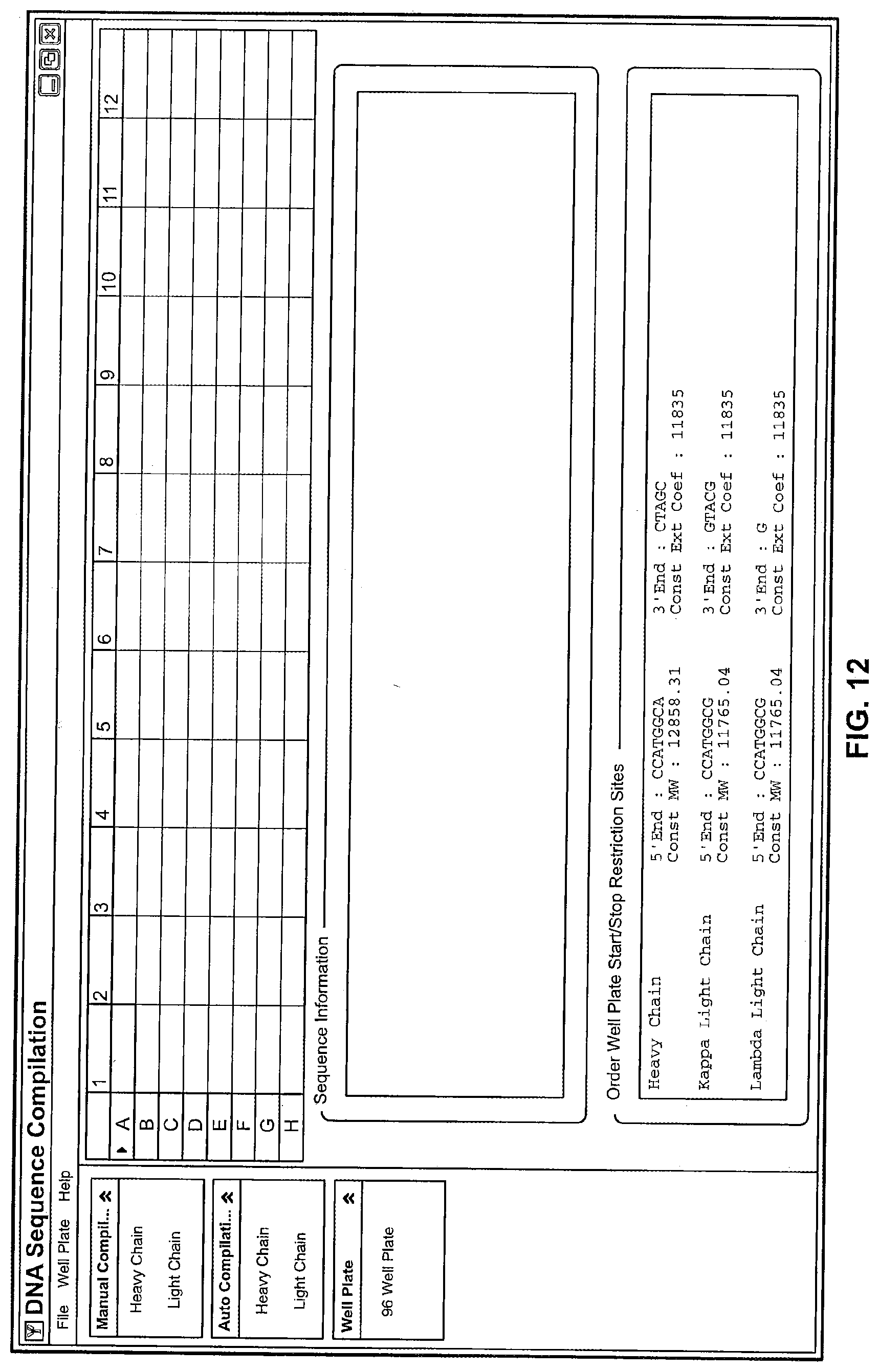
D00014
D00015

D00016
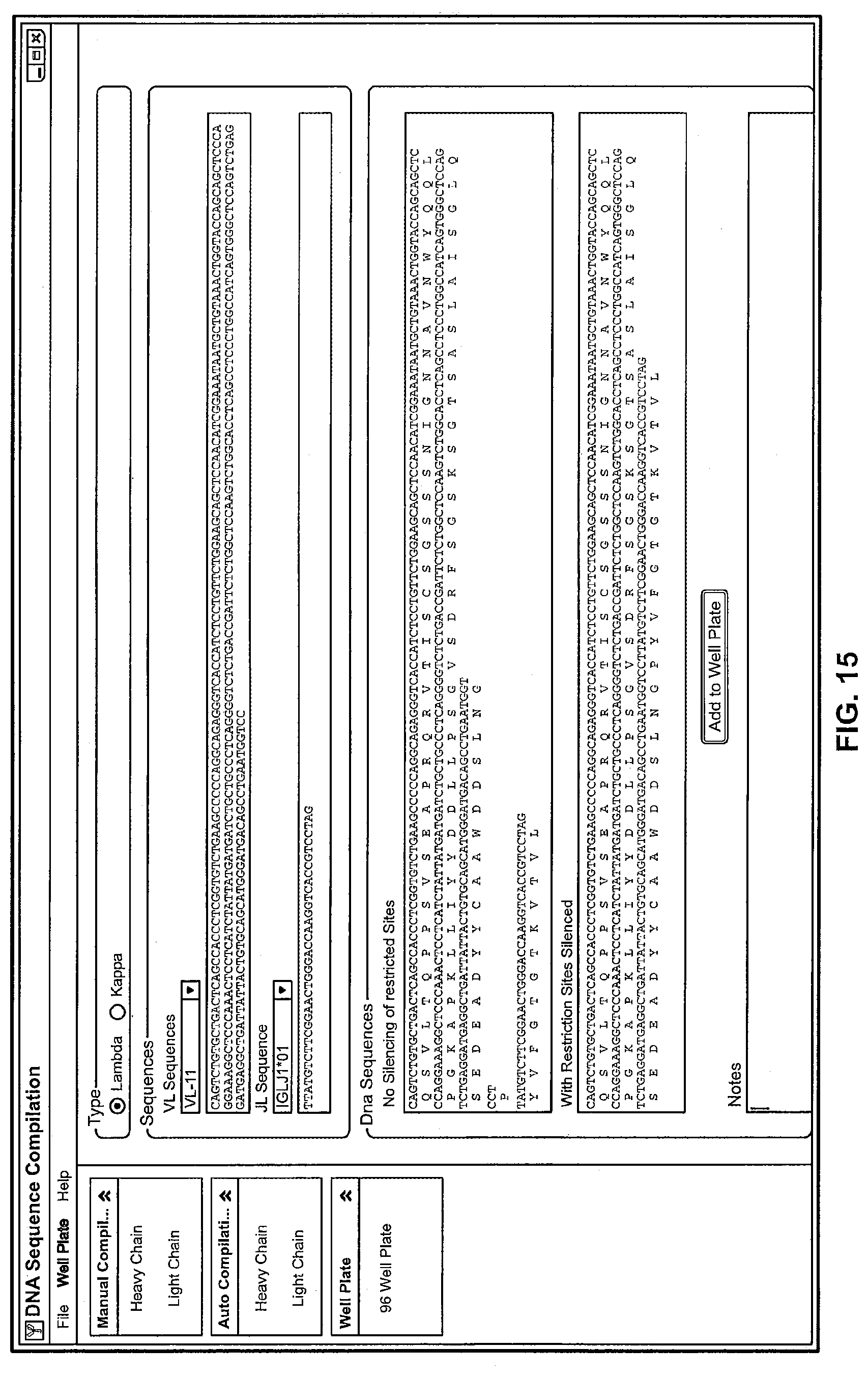
D00017
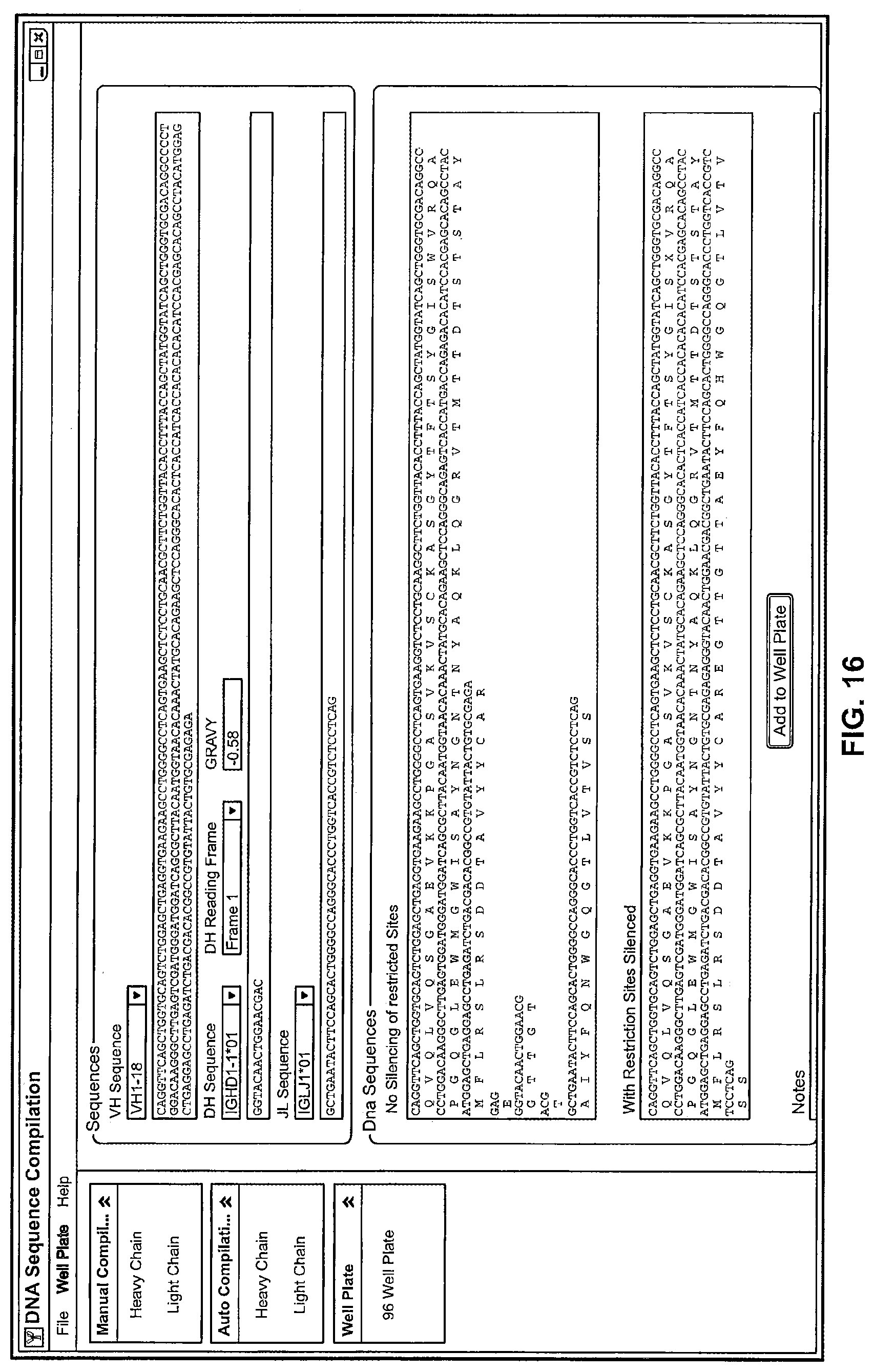

D00018
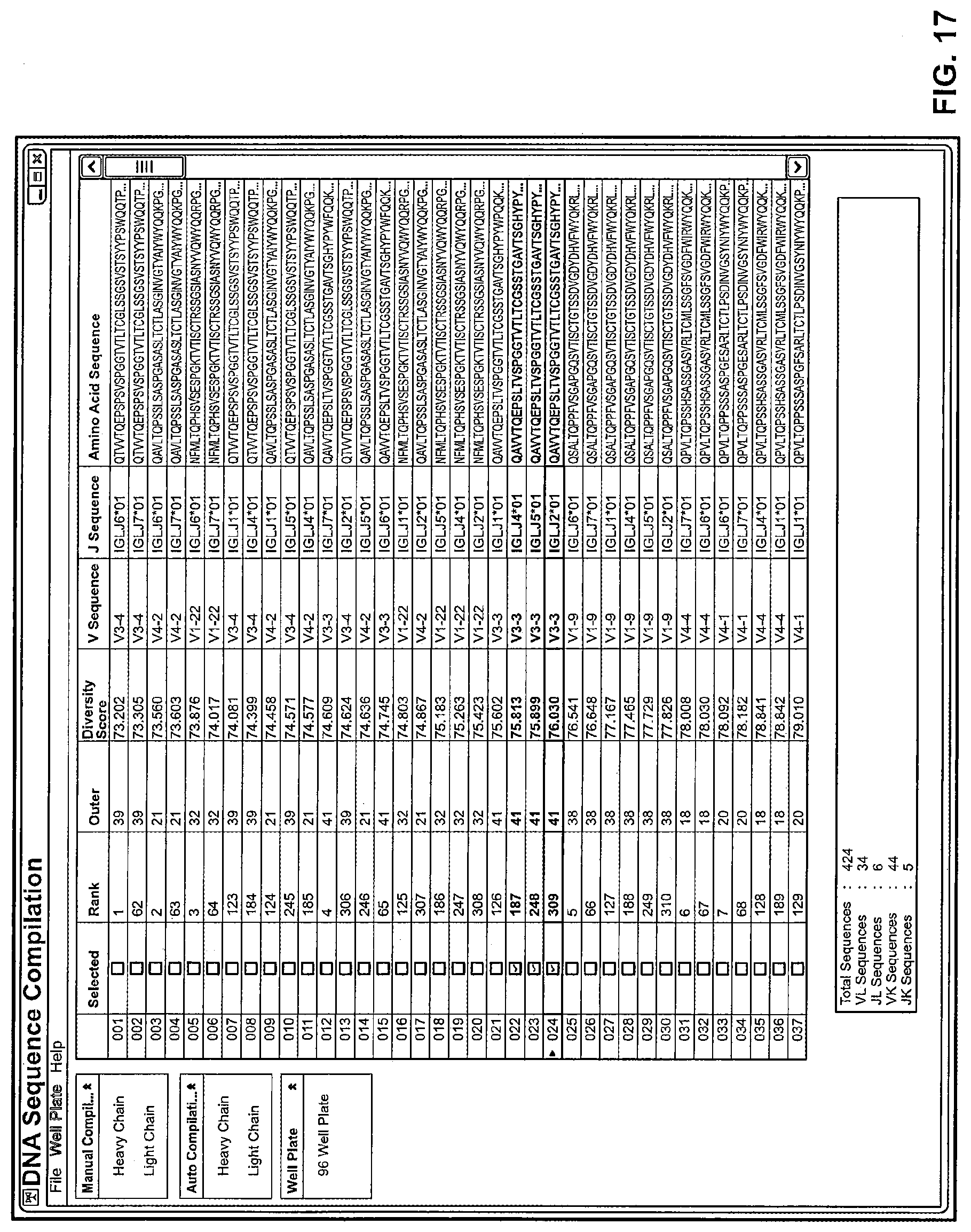

D00019
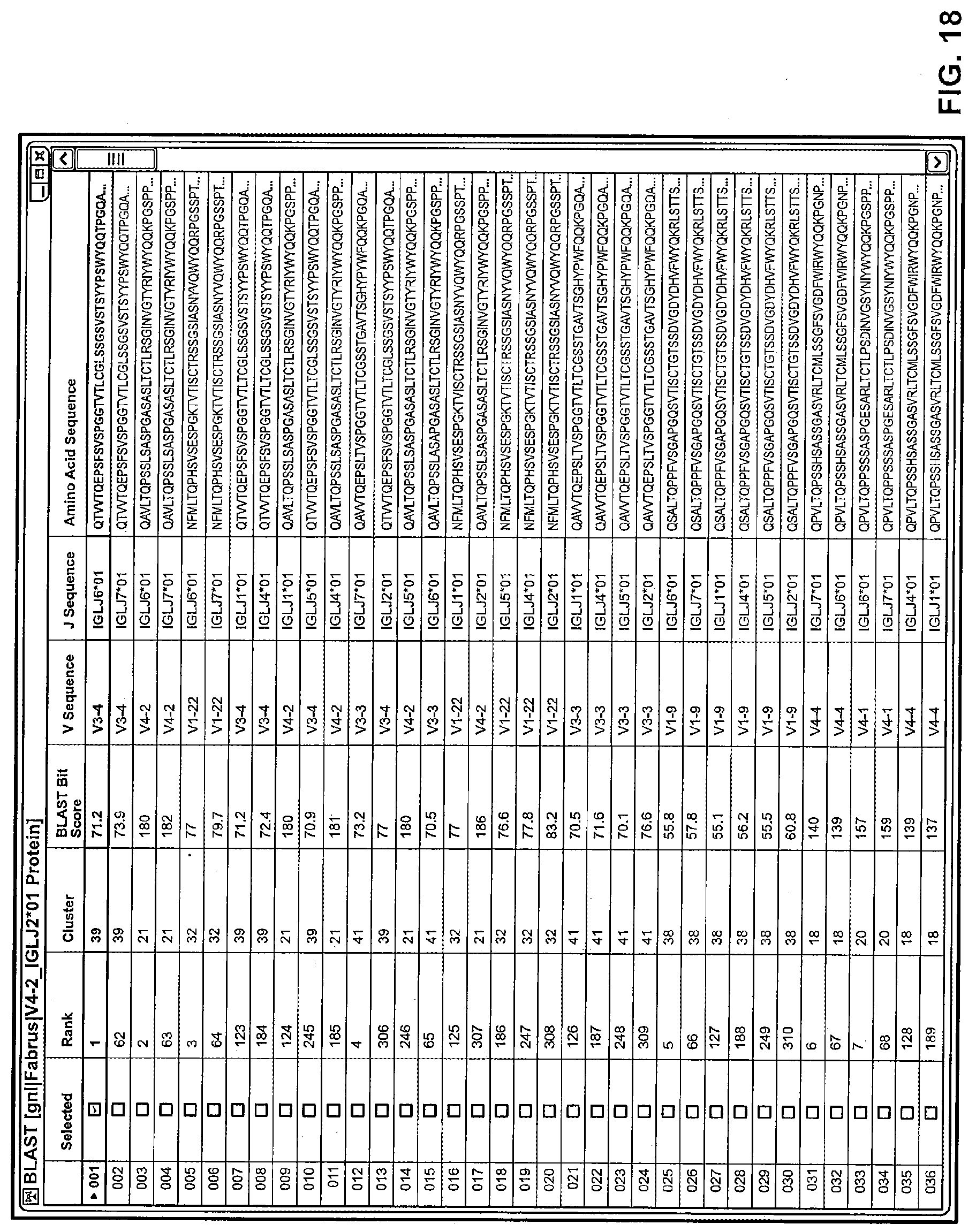
D00020
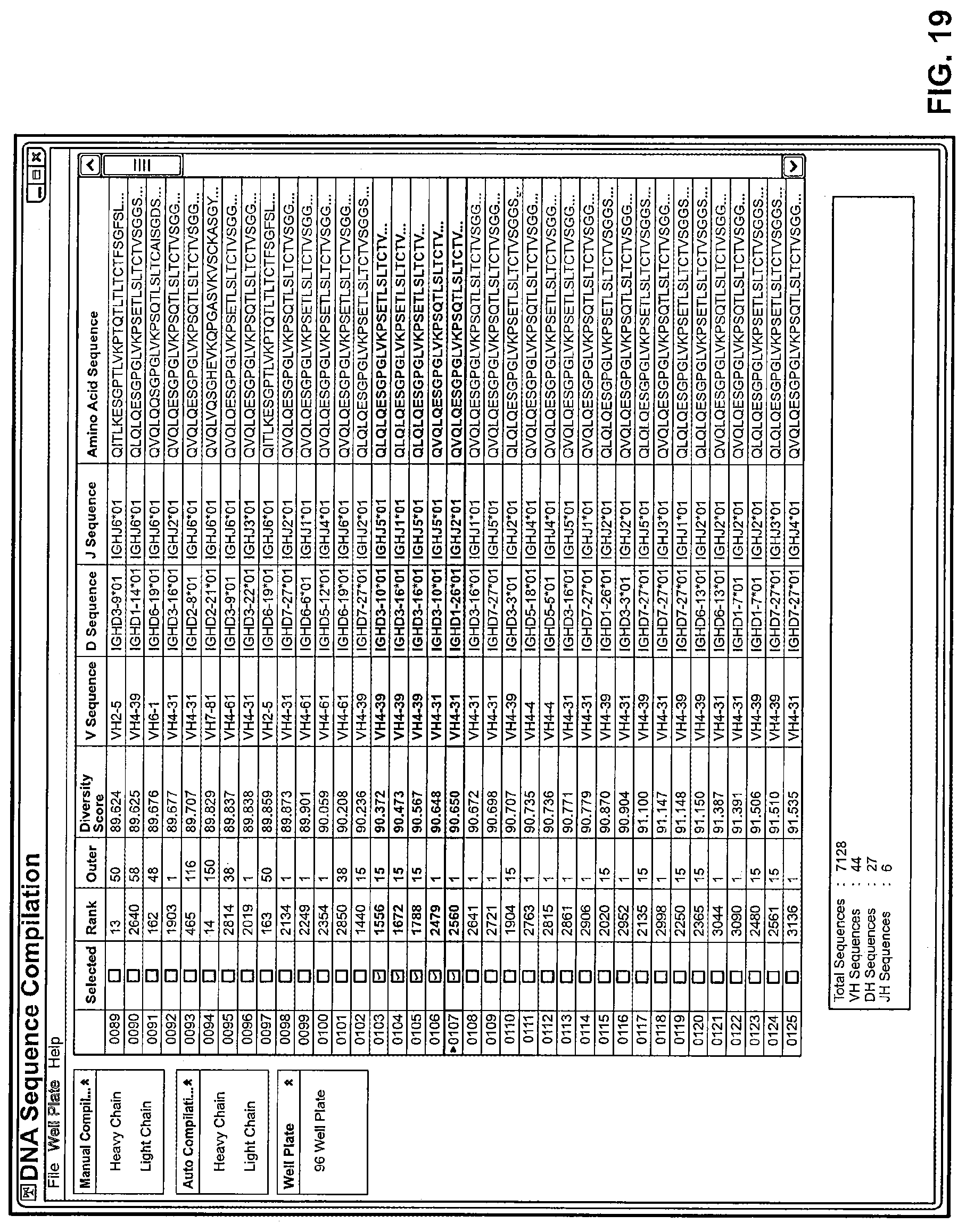

D00021

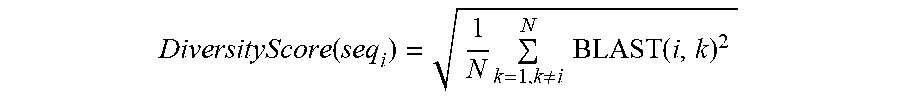
P00001
P00899
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.