Antibodies And Antigen-binding Fragments That Specifically Bind To Microtubule-associated Protein Tau
WADIA; Jehangir ; et al.
U.S. patent application number 16/514770 was filed with the patent office on 2020-12-24 for antibodies and antigen-binding fragments that specifically bind to microtubule-associated protein tau. The applicant listed for this patent is Janssen Vaccines & Prevention B.V.. Invention is credited to Jaap GOUDSMIT, Gabriel PASCUAL, Katarina RADOSEVIC, Jehangir WADIA, Robert Anthony WILLIAMS.
| Application Number | 20200399350 16/514770 |
| Document ID | / |
| Family ID | 1000005066198 |
| Filed Date | 2020-12-24 |




View All Diagrams
| United States Patent Application | 20200399350 |
| Kind Code | A1 |
| WADIA; Jehangir ; et al. | December 24, 2020 |
ANTIBODIES AND ANTIGEN-BINDING FRAGMENTS THAT SPECIFICALLY BIND TO MICROTUBULE-ASSOCIATED PROTEIN TAU
Abstract
The invention relates to antibodies and antigen-binding fragments that specifically bind to microtubule-associated protein tau. The invention also relates to diagnostic, prophylactic and therapeutic methods using anti-tau antibodies.
| Inventors: | WADIA; Jehangir; (San Diego, CA) ; PASCUAL; Gabriel; (San Diego, CA) ; WILLIAMS; Robert Anthony; (Leiden, NL) ; RADOSEVIC; Katarina; (Nootdorp, NL) ; GOUDSMIT; Jaap; (Amsterdam, NL) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000005066198 | ||||||||||
| Appl. No.: | 16/514770 | ||||||||||
| Filed: | July 17, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15321918 | Dec 23, 2016 | 10400034 | ||
| PCT/EP2015/064533 | Jun 26, 2015 | |||
| 16514770 | ||||
| 62017789 | Jun 26, 2014 | |||
| 62017812 | Jun 26, 2014 | |||
| 62017807 | Jun 26, 2014 | |||
| 62017746 | Jun 26, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2317/24 20130101; C07K 2317/30 20130101; C07K 2317/94 20130101; C07K 16/18 20130101; C07K 2317/565 20130101; C07K 2317/72 20130101; G01N 33/6896 20130101; A61K 39/395 20130101; C07K 2317/34 20130101; G01N 2800/2821 20130101; C07K 2317/21 20130101; G01N 2333/47 20130101; C07K 2317/56 20130101; C07K 2317/567 20130101; C07K 2317/52 20130101 |
| International Class: | C07K 16/18 20060101 C07K016/18; G01N 33/68 20060101 G01N033/68; A61K 39/395 20060101 A61K039/395 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Aug 4, 2014 | EP | 1417699.5 |
| Aug 4, 2014 | EP | 14178706.8 |
| Aug 4, 2014 | EP | 14179719.1 |
| Aug 4, 2014 | EP | 14179739.9 |
Claims
1. A monoclonal antibody, wherein the antibody binds tau deposits in human AD brain tissue, wherein the antibody binds tau deposits in human AD brain tissue, but does not bind tau in normal human brain tissue and does not bind tau deposits in Progressive Supranuclear Palsy ("PSP") brain tissue; and wherein the antibody comprises an antigen-binding site of a VH of SEQ ID NO:87 and an antigen-binding site of a VL of SEQ ID NO:88.
2. The antibody of claim 1, wherein the antibody comprises: a naturally occurring human antigen-binding variable region that binds specifically to tau, and a recombinant constant region of a human IgG1 antibody.
3. The antibody of claim 1, wherein the antibody comprises: naturally occurring human light and heavy chain variable regions from a human antibody, and recombinant human IgG1 heavy and light chain constant regions.
4. The antibody of claim 1, wherein the antibody comprises: heavy and light chain variable regions from a naturally occurring human antibody, and recombinant human IgG1 heavy and light chain constant regions.
5. The antibody of claim 1, wherein the antibody comprises: heavy and light chain variable regions from a human antibody, and recombinant human IgG1 heavy and light chain constant regions.
6. The antibody of claim 1, wherein the antibody is selected from the group consisting of a) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:175, a heavy chain CDR2 region comprising SEQ ID NO:176, and a heavy chain CDR3 region comprising SEQ ID NO:177, a light chain CDR1 region comprising SEQ ID NO:178, a light chain CDR2 region comprising SEQ ID NO:179 and a light chain CDR3 region comprising SEQ ID NO:180, b) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:181, a heavy chain CDR2 region comprising SEQ ID NO:182, and a heavy chain CDR3 region comprising SEQ ID NO:183, a light chain CDR1 region comprising SEQ ID NO:172, a light chain CDR2 region comprising SEQ ID NO:173 and a light chain CDR3 region comprising SEQ ID NO:184, c) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:185, a heavy chain CDR2 region comprising SEQ ID NO:186, and a heavy chain CDR3 region comprising SEQ ID NO:187, a light chain CDR1 region comprising SEQ ID NO:188, a light chain CDR2 region of comprising SEQ ID NO:173 and a light chain CDR3 region comprising SEQ ID NO:189, d) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:190, a heavy chain CDR2 region comprising SEQ ID NO:191, and a heavy chain CDR3 region comprising SEQ ID NO:192, a light chain CDR1 region comprising SEQ ID NO:193, a light chain CDR2 region comprising SEQ ID NO:194 and a light chain CDR3 region comprising SEQ ID NO:195, e) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:196, a heavy chain CDR2 region comprising SEQ ID NO:197, and a heavy chain CDR3 region comprising SEQ ID NO:198, a light chain CDR1 region comprising SEQ ID NO:199, a light chain CDR2 region comprising SEQ ID NO:173 and a light chain CDR3 region comprising SEQ ID NO:200, f) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:213, a heavy chain CDR2 region comprising SEQ ID NO:214, and a heavy chain CDR3 region comprising SEQ ID NO:215, a light chain CDR1 region comprising SEQ ID NO:216, a light chain CDR2 region comprising SEQ ID NO:173 and a light chain CDR3 region comprising SEQ ID NO:217; NO:217, g) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:213, a heavy chain CDR2 region comprising SEQ ID NO:214, and a heavy chain CDR3 region comprising SEQ ID NO:215, a light chain CDR1 region comprising SEQ ID NO:218, a light chain CDR2 region comprising SEQ ID NO:174 and a light chain CDR3 region comprising SEQ ID NO:217; NO:217, and h) an antibody comprising a heavy chain CDR1 region comprising SEQ ID NO:219, a heavy chain CDR2 region comprising SEQ ID NO:220, and a heavy chain CDR3 region comprising SEQ ID NO:221, a light chain CDR1 region comprising SEQ ID NO:218, a light chain CDR2 region comprising SEQ ID NO:173 and a light chain CDR3 region comprising SEQ ID NO:217.
7. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:95 and an antigen-binding site of a VL of SEQ ID NO:96.
8. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:99 and an antigen-binding site of a VL of SEQ ID NO:100.
9. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:103 and an antigen-binding site of a VL of SEQ ID NO:104.
10. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:107 and an antigen-binding site of a VL of SEQ ID NO:108.
11. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:111 and an antigen-binding site of a VL of SEQ ID NO:112.
12. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:123 and an antigen-binding site of a VL of SEQ ID NO:124.
13. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:127 and an antigen-binding site of a VL of SEQ ID NO:128.
14. The antibody of claim 1, comprising an antigen-binding site of a VH of SEQ ID NO:131 and an antigen-binding site of a VL of SEQ ID NO:132.
15. A method for the prophylaxis and/or treatment of a subject with Alzheimer's disease, or memory and/or cognitive disorders associated with tau, the method comprising utilizing the chimeric antibody of claim 1 or an antigen-binding fragment thereof for prophylaxis or treatment, or combination thereof, of Alzheimer's disease, or memory and/or cognitive disorders associated with tau.
16. A method for diagnosing a subject for Alzheimer's disease, or memory and/or cognitive disorders associated with tau, the method comprising: a) assaying the level of tau antigen in a biological sample from the subject utilizing the chimeric antibody of claim 1 or an antigen-binding fragment thereof.
17. The method of claim 16, wherein the biological sample comprises peripheral blood, serum, plasma, urine, cerebrospinal fluid, tissue biopsy, surgical specimen, fine needle aspirates, autopsy material, cell culture supernatant, isolated cells, fermentation supernatant, or tissue homogenate.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of application Ser. No. 15/321,918, filed on Dec. 23, 2016, which is the national phase entry under 35 U.S.C. .sctn. 371 of International Patent Application PCT/EP2015/064533, filed Jun. 26, 2015, designating the United States of America and published in English as International Patent Publication WO 2015/197823 A1 on Dec. 30, 2015, which claims the benefit under Article 8 of the Patent Cooperation Treaty to European Patent Application Serial Nos. 14179739.9, 14179719.1, 14179706.8 and 14179699.5, all filed on Aug. 4, 2014, and under 35 U.S.C. .sctn. 119 to U.S. Provisional Patent Application Ser. Nos. 62/017,746, 62/017,807, 62/017,789, and 62/017,812, all filed on Jun. 26, 2014. The entire disclosures of the prior applications are hereby incorporated by reference herein in their entirety.
STATEMENT ACCORDING TO 37 C.F.R. .sctn. 1.821(c) OR (e)--SEQUENCE LISTING SUBMITTED AS ASCII TEXT FILE
[0002] Pursuant to 37 C.F.R. .sctn. 1.821(c) or (e), this application contains a sequence listing, which is contained on an ASCII text file entitled "Sequence Listing" (SYT 3017-CON_Corrected_PatentIn Seq 535_ST25.txt, created Jul. 14, 2020, having a size of 262,774 bytes), which is herein incorporated by reference.
TECHNICAL FIELD
[0003] The application relates to medicine. The disclosure, in particular, relates to antibodies and antigen-binding fragments that specifically bind to microtubule-associated protein tau. The disclosure also relates to diagnostic, prophylactic and therapeutic methods using anti-tau antibodies.
BACKGROUND
[0004] Dementia is a syndrome that can be caused by a number of progressive disorders that affect memory, thinking, behavior and the ability to perform everyday activities. About 36 million people worldwide are suffering from dementia today. The number of people with dementia is projected to double by 2030, and more than triple to 115.4 million people by 2050. Alzheimer's disease (AD) is the most common type of dementia. Currently, one in nine people age 65 and older (11 percent) and nearly half of those over age 85 have Alzheimer's disease. According to Alzheimer's Disease International, current global costs of caring for these patients exceeds $600 billion annually. These costs are likely to rise even faster than the prevalence of disease, especially in the developing world, as more formal social care systems emerge, and rising incomes lead to higher opportunity costs (B. Winblad and L Jonsson, World Alzheimer Report 2010).
[0005] The brains of AD patients have an abundance of two abnormal structures, amyloid plaques and neurofibrillary tangles. This is especially true in certain regions of the brain that are important in memory. There is also a substantial loss of neurons and synapses in the cerebral cortex and certain subcortical regions. Both neurofibrillary tangles and neuronal loss increase in parallel with the duration and severity of illness (T. Gomez-Isla et al., Ann. Neurol. 1997; 41:17-24) and neurofibrillary load has been shown to correlate with cognitive decline (H. Braak and E. Braak, Neurobiol. Aging 1997 July-August; 18(4):351-7).
[0006] Neurofibrillary tangles are intraneuronal lesions that are composed of hyperphosphorylated and insoluble accumulations of the microtubule-associated protein, tau. These accumulations are a histopathological feature of many neurodegenerative diseases, which are collectively known as tauopathies. Tauopathies include, e.g., Alzheimer's disease (AD), Pick's disease (PiD), progressive supranuclear palsy (PSP), corticobasal degeneration (CBD), and frontotemporal lobar degeneration (FTLD). In human tauopathies, pathology progresses from one brain region to another in disease-specific patterns (H. Braak and E. Braak, Neurobiol. Aging 1997 July-August; 18(4):351-7; Raj et al., Neuron. 2012; 73:1204-1215; Seeley et al., Neuron. 2009; 62:42-52; and Zhou et al. Neuron. 2012; 73:1216-1227), the underlying mechanism of which is not yet clear.
[0007] Tau pathology is involved in and may be a cause of many tauopathies. In its normal form, tau is a highly soluble microtubule-associated protein (Jeganathan et al., Biochemistry 2008, 47:10526-10539) that binds and promotes the assembly of microtubules (Drechsel et al., Mol. Biol. Cell. 1992, 3:1141-1154). However, in tauopathies, tau becomes hyperphosphorylated, causing detachment from microtubules, and ultimately accumulation as neurofibrillary tangles that are visualized within dystrophic neurites and cell bodies (Mandelkow and Mandelkow, Cold Spring Harbor, Perspect. Med. 2, 2012: a006247). The amount of tau pathology correlates with progressive neuronal dysfunction, synaptic loss, and functional decline in humans and transgenic mouse models (Arriagada et al., Neurology 1992 March; 42(3 Pt 1):631-9; Bancher et al., Neurosci. Lett. 1993, 162:179-182; Polydoro et al., J. Neuroscience 2009, 29:10741-10749; and Small and Duff, Neuron. 2008 Nov. 26; 60(4):534-42). While there have been no tau mutations observed in Alzheimer's disease, mutations in the tau gene appear to cause some forms of frontotemporal dementia (Cairns et al., Am. J. Pathol. 2007, 171:227-40), presenting with tau-positive inclusions and signifying that tau dysfunction is sufficient to cause neurodegeneration. Moreover, pathological tau appears to be an integral part of AP-induced neurotoxicity in cell culture and transgenic animal models (M. Rapoport, PNAS 2002, 99:9, 6364-6369; E. D. Roberson et al., Science 2007, 316:750-754; A. M. Nicholson and A. Ferreira, J. Neurosci. 2009, 29:4640-4651; H. Oakley, J. Neurosci. 2006, 26(40):10129-10140).
[0008] Passive and active immunizations against tau have been analyzed in mice using several different mouse models, including different phospho-tau peptides for active immunizations and anti-tau antibodies for passive immunotherapy (A. A. Asuni et al., J. Neurosci. 2007, 27(34):9115-9129; E. M. Sigurdsson, Curr. Alzheimer Res. 2009, 6(5):446-450; A. Boutajangout et al., J. Neurosci. 2010, 30(49):16559-16566; H. Rosenmann et al., Arch. Neurol. 2006, 63(10):1459-1467; M. Boimel et al., Exp. Neurol. 2010, 224(2):472-485). In the first report describing immunizations with a 30-amino acid phosphorylated tau peptide, an effect on the ratios of soluble and insoluble tau, reduction of tangle formation in the immunized mice, and functional benefits observed in behavior testing for these mice were shown (A. Boutajangout et al., J. Neuroscience 2010, 30:16559-16566). Passive immunization with well-characterized anti-tau antibodies, which react with phosphorylated Ser396 and Ser404 of the hyperphosphorylated tau protein at an early pathologic conformational epitope on tau, confirmed the results seen in active immunization studies. Mice treated with these antibodies showed marked reductions in tau pathology, which was measured by biochemical methods and histology, as well as a significant delay in loss of motor-function decline that was assessed in behavioral testing (A. Boutajangout et al., J. Neurochem. 2011, 118(4):658-667; X. Chai et al., J. Biol. Chem. 2011, 286(39):34457-34467).
[0009] Tau-based therapies have only been analyzed in mouse models to date. But in view of the severity of tauopathies in general, and to the cost to society of Alzheimer's disease specifically, there is an ongoing need for effective means to diagnose, monitor, prevent and treat tauopathies.
BRIEF SUMMARY
[0010] Provided are antibodies comprising an antigen-binding variable region that binds specifically to tau. Provided in particular, are anti-tau antibodies, and antigen-binding fragments thereof, that detect tau in normal human brain tissue, or detect tau deposits in human Alzheimer's Disease (AD) and Progressive Supranuclear Palsy (PSP) brain tissue. In certain embodiments, the anti-tau antibodies, and antigen-binding fragments thereof, bind to recombinant tau and/or paired helical filament ("PHF")-tau by Western assay.
[0011] Further provided are chimeric antibodies comprising an antigen-binding variable region from a naturally occurring human antibody that binds specifically to tau, and a recombinant constant region of a human IgG1, wherein the constant region of the chimeric antibody is different from the naturally occurring antibody.
[0012] In certain embodiments, provided are chimeric anti-tau antibodies, and antigen-binding fragments thereof, that detect tau in normal human brain tissue, but do not detect tau deposits in human Alzheimer's Disease (AD) and Progressive Supranuclear Palsy (PSP) brain tissue.
[0013] In certain embodiments, the (chimeric) anti-tau antibodies, and antigen-binding fragments thereof, bind to recombinant tau and/or PHF-tau by Western assay. In certain embodiments, the anti-tau antibodies, and antigen-binding fragments thereof, do not bind to recombinant tau or PHF-tau in an ELISA. The antibodies and antigen-binding fragments are preferentially capable of specifically binding to a tau peptide. In some embodiments, the antibodies and antigen-binding fragments are preferentially capable of specifically binding to a tau phospho-peptide but do not bind the peptide when it is not phosphorylated.
[0014] In certain embodiments, the anti-tau antibodies are not naturally occurring. Preferably, the antibodies are human.
[0015] The antibodies and antigen-binding fragments of the disclosure are useful as diagnostic, prophylactic and/or therapeutic agents, both alone and in combination with other diagnostics, prophylactic and/or therapeutic agents.
[0016] In one aspect, the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:163, a heavy chain CDR2 region of SEQ ID NO:164, and a heavy chain CDR3 region of SEQ ID NO:165, a light chain CDR1 region of SEQ ID NO:166, a light chain CDR2 region of SEQ ID NO:167 and a light chain CDR3 region of SEQ ID NO:168.
[0017] Another embodiment of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:169, a heavy chain CDR2 region of SEQ ID NO:170, and a heavy chain CDR3 region of SEQ ID NO:171, a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:174.
[0018] In one aspect, the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:175, a heavy chain CDR2 region of SEQ ID NO:176, and a heavy chain CDR3 region of SEQ ID NO:177, a light chain CDR1 region of SEQ ID NO:178, a light chain CDR2 region of SEQ ID NO:179 and a light chain CDR3 region of SEQ ID NO:180.
[0019] Another embodiment of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:181, a heavy chain CDR2 region of SEQ ID NO:182, and a heavy chain CDR3 region of SEQ ID NO:183, a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:184.
[0020] Another embodiment of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:185, a heavy chain CDR2 region of SEQ ID NO:186, and a heavy chain CDR3 region of SEQ ID NO:187, a light chain CDR1 region of SEQ ID NO:188, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:189.
[0021] Yet another aspect of the disclosure relates to an anti-tau antibody, comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:190, a heavy chain CDR2 region of SEQ ID NO:191, and a heavy chain CDR3 region of SEQ ID NO:192, a light chain CDR1 region of SEQ ID NO:193, a light chain CDR2 region of SEQ ID NO:194 and a light chain CDR3 region of SEQ ID NO:195.
[0022] Yet another aspect of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:196, a heavy chain CDR2 region of SEQ ID NO:197, and a heavy chain CDR3 region of SEQ ID NO:198, a light chain CDR1 region of SEQ ID NO:199, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:200.
[0023] Yet another aspect of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, a light chain CDR1 region of SEQ ID NO:216, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:217.
[0024] A further aspect of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:174 and a light chain CDR3 region of SEQ ID NO:217.
[0025] A further aspect of the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain CDR1 region of SEQ ID NO:219, a heavy chain CDR2 region of SEQ ID NO:220, and a heavy chain CDR3 region of SEQ ID NO:221, a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:217.
[0026] The disclosure further relates to antigen-binding fragments of the above antibodies.
[0027] In another aspect, the disclosure relates to an anti-tau antibody comprising an antigen-binding site of a heavy chain variable region (VH) of SEQ ID NOS:87, 91, 95, 99, 103, 107, 111, 123, 127 or 131, and an antigen-binding site of a light chain variable region (VL) of SEQ ID NOS:88, 92, 96, 100, 104, 108, 112, 124, 128 or 132, and to antigen-binding fragments thereof.
[0028] In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:87 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:88 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:91 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:92 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:95 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:96 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:99 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:100 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:103 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:104 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:107 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:108 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:111 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:112 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:123 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:124 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:127 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:128 and to antigen-binding fragments thereof. In a further aspect, the disclosure relates to an anti-tau antibody comprising a heavy chain variable region comprising the amino acid sequence of SEQ ID NO:131 and a light chain variable region comprising an amino acid sequence of SEQ ID NO:132 and to antigen-binding fragments thereof.
[0029] In one embodiment, the IgG1 heavy chain constant region is comprised of the amino acid sequence of SEQ ID NO:83. In another embodiment, the IgG1 light chain constant region is comprised of the amino acid sequence of SEQ ID NO:84.
[0030] The disclosure also provides nucleic acid molecules encoding the antibodies or antigen-binding fragments thereof. Another aspect of the disclosure is a vector comprising the nucleic acid molecules of the disclosure. A further feature of the disclosure is a host cell comprising the vector of the disclosure.
[0031] The disclosure also provides a method of producing an anti-tau antibody comprising culturing the host cell of the disclosure and recovering the antibody produced by the host cell.
[0032] The disclosure further provides for functional variants of the antibodies and immunoconjugates comprising the antibody and/or antigen-binding fragment thereof
[0033] The disclosure further provides compositions and kits that comprise one or more antibodies of the disclosure and/or antigen-binding fragments thereof. The disclosure additionally provides diagnostic, prophylactic and therapeutic methods that employ the anti-tau antibodies. Prophylactic and therapeutic methods include administering to human subjects the anti-tau antibodies and/or antigen-binding fragments thereof for the prevention or treatment of a tauopathy and/or tau-mediated diseases or conditions, and/or amelioration of one or more symptoms of a tauopathy. Combinations of a plurality of different anti-tau antibodies and/or antigen-binding fragments thereof and/or with other anti-tau antibodies can be used for combination therapy. Compositions comprising the anti-tau antibodies and/or antigen-binding fragments thereof in combination with other prophylactic or therapeutic agents are also provided.
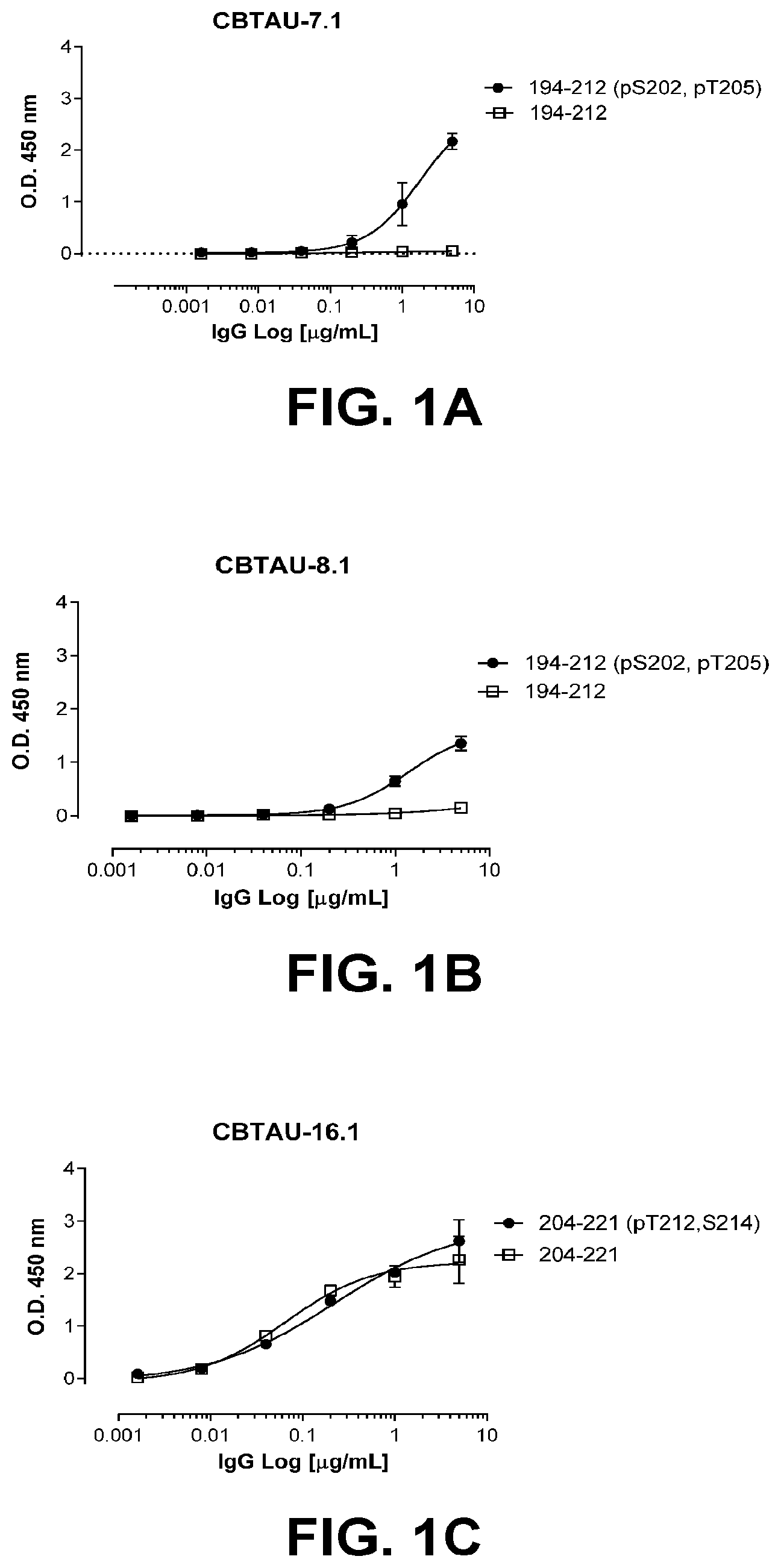
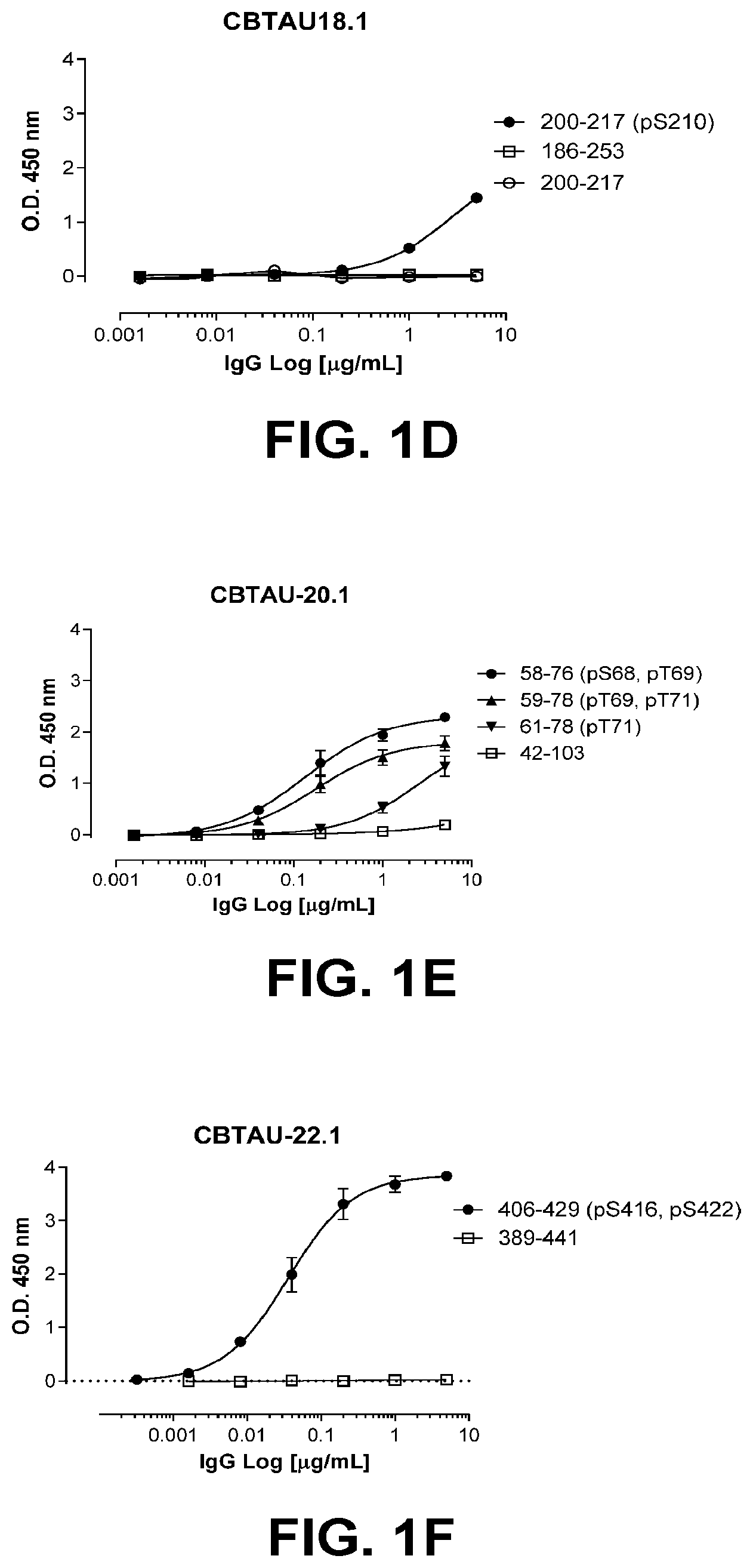
[0034] In some embodiments, the antibodies of the disclosure are unique in that the variable regions are recovered from anti-tau-specific memory B-cells from healthy individuals and detect tau deposits in human Alzheimer's brain, but do not detect tau in normal human brain, and are phospho-dependent and bind to tau peptide ptau 194-212 (SEQ ID NO:315) or ptau 200-217 (SEQ ID NO:319) or ptau 59-78 (SEQ ID NO:323) or ptau 406429 (SEQ ID NO:326). In other embodiments, the antibodies are unique in that the variable regions are recovered from anti-tau-specific memory B-cells from healthy individuals and detect tau deposits in human Alzheimer's brain, and detect tau in normal (i.e., healthy) human brain and are predominantly phospho-independent and bind to tau peptide tau 204-221 (SEQ ID NO:318) or tau 221-253 (SEQ ID NO:367). In still other embodiments, the antibodies of the disclosure are unique in that the variable regions are recovered from anti-tau-specific memory B-cells from individuals with Alzheimer's disease and are phospho-dependent and bind to phosphorylated tau peptide ptau 406429 (SEQ ID NO:326) and do not bind to unphosphorylated tau peptide tau 389-441 (SEQ ID NO:327).
BRIEF DESCRIPTION OF THE DRAWINGS
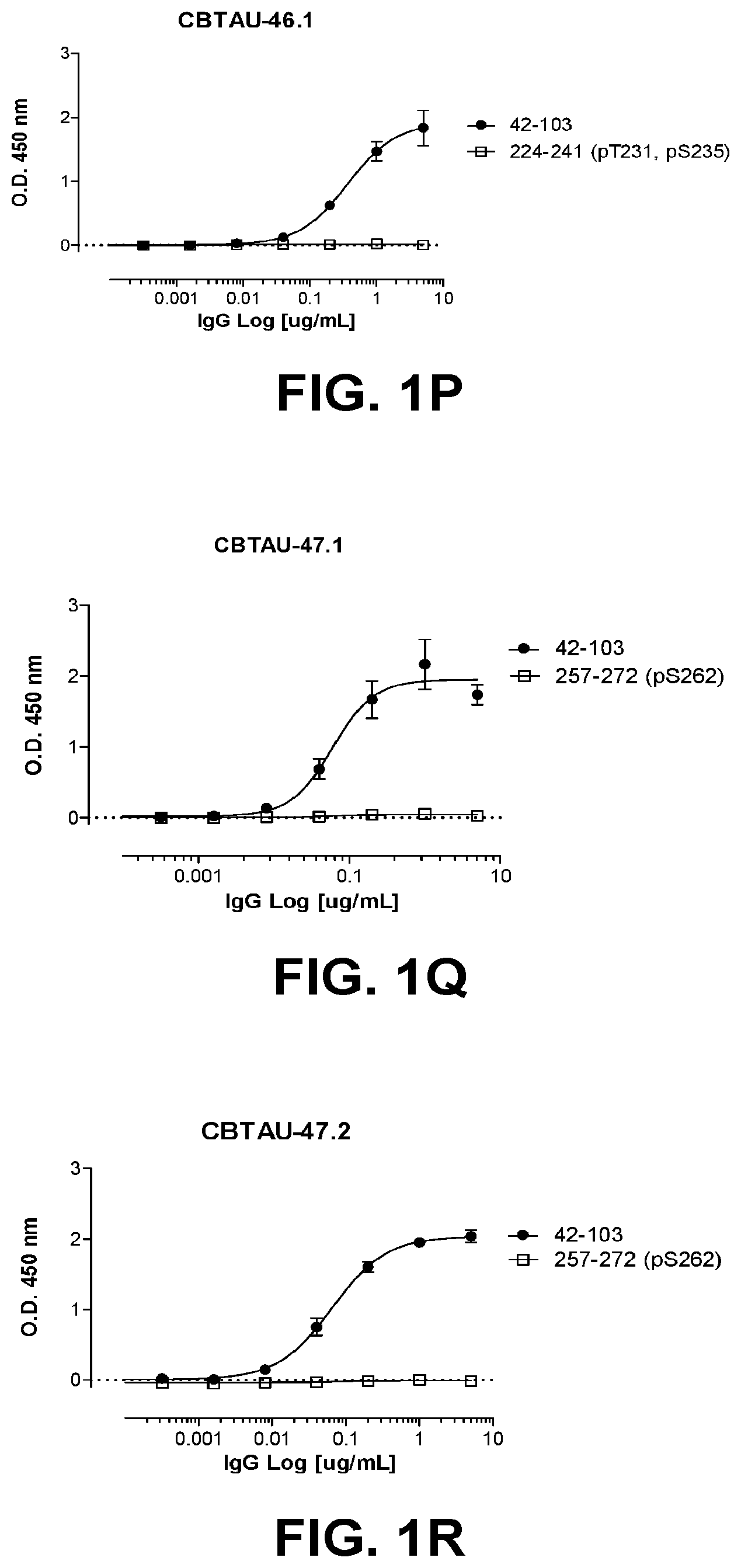
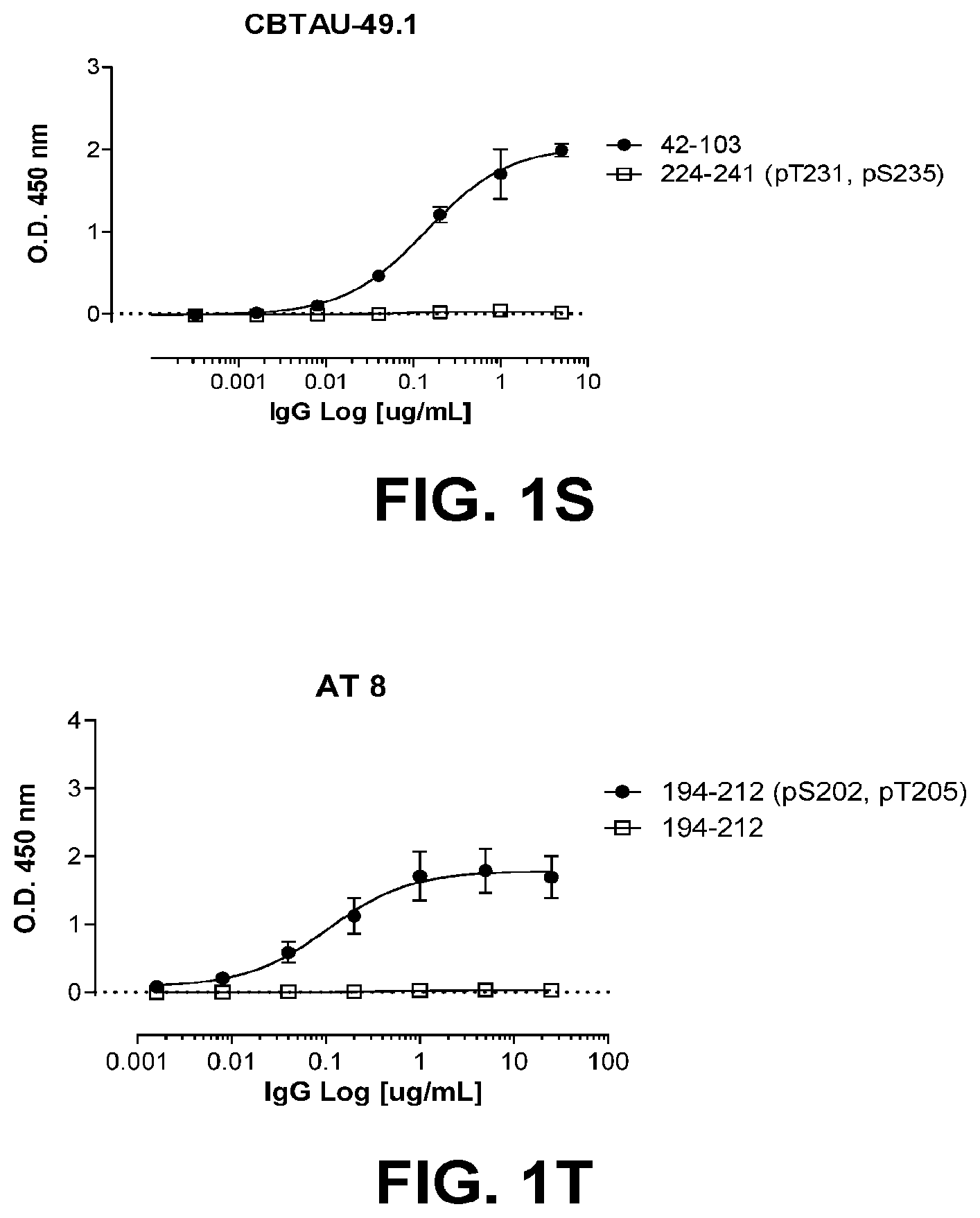
[0035] FIGS. 1A-1T show the reactivity of CBTAU-7.1, 8.1, 16.1, 18.1, 20.1, 22.1, 24.1, 27.1, 28.1, 41.1, 41.2, 42.1, 43.1, 44.1, 45.1, 46.1, 47.1, 47.2, and 49.1 against corresponding cognate and non-cognate peptide.
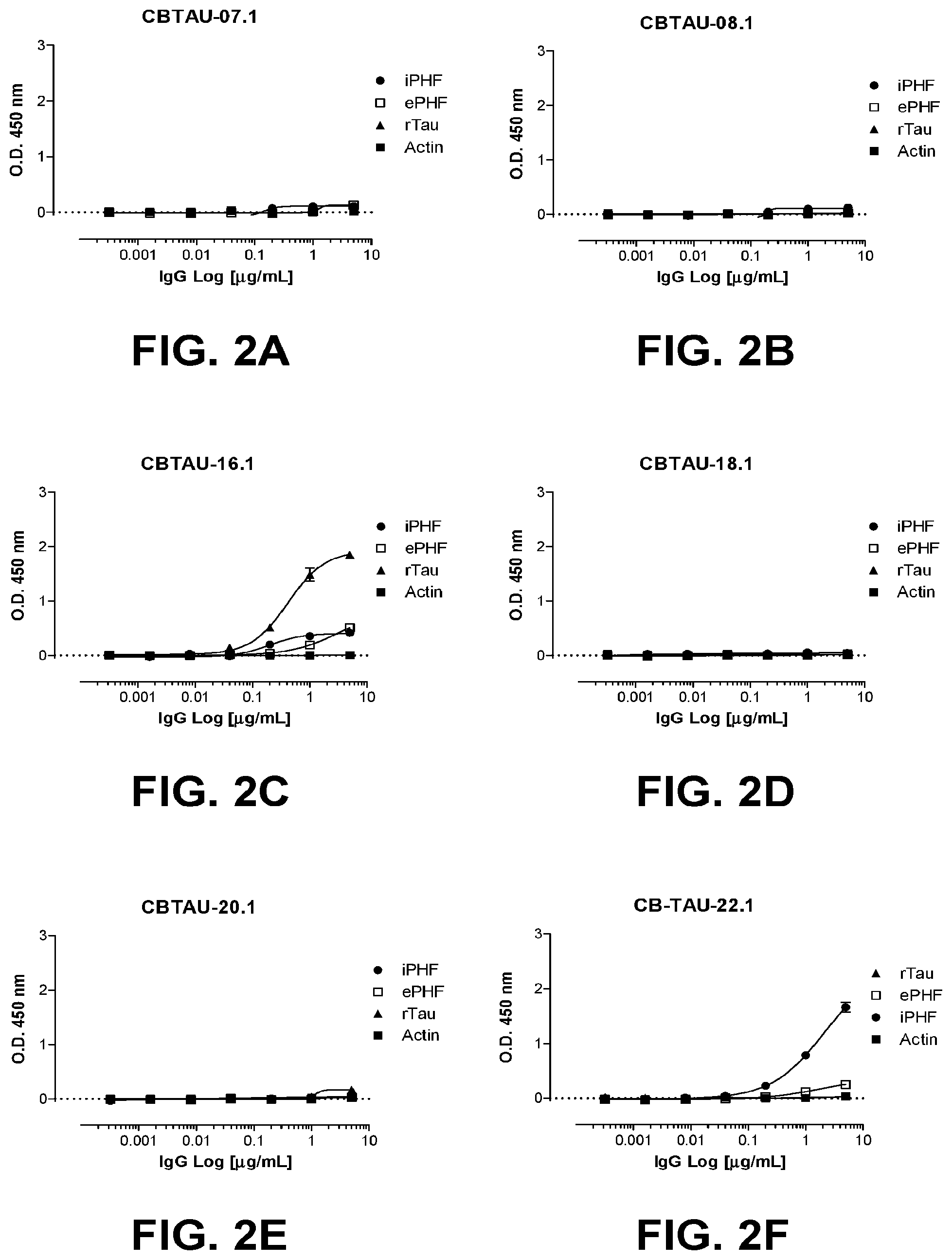
[0036] FIGS. 2A-2J show the reactivity of CBTAU-7.1, 8.1, 16.1, 18.1, 20.1, 22.1, 24.1, 27.1, and 28.1 against recombinant tau (rTau), enriched immunopurified paired helical filaments (ePHF), and immunopurified paired helical filaments (iPHF) by ELISA. Anti-tau mAb, ATB, was used as positive control.
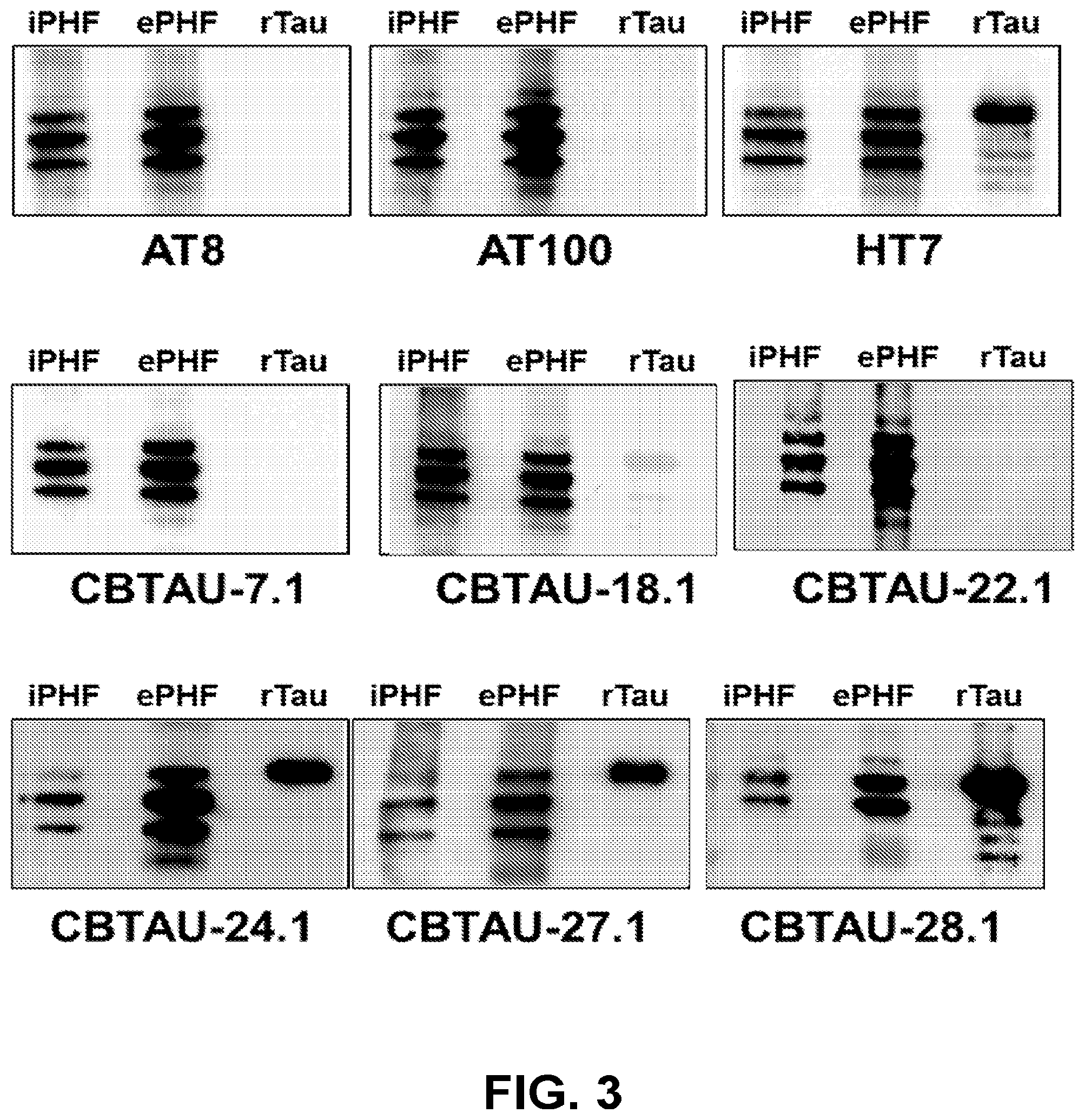
[0037] FIG. 3 shows the immunoreactivity of CBTAU-7.1, 18.1, 22.1, 24.1, 27.1, and 28.1 against rTau, ePHF, and iPHF by Western blot analysis.
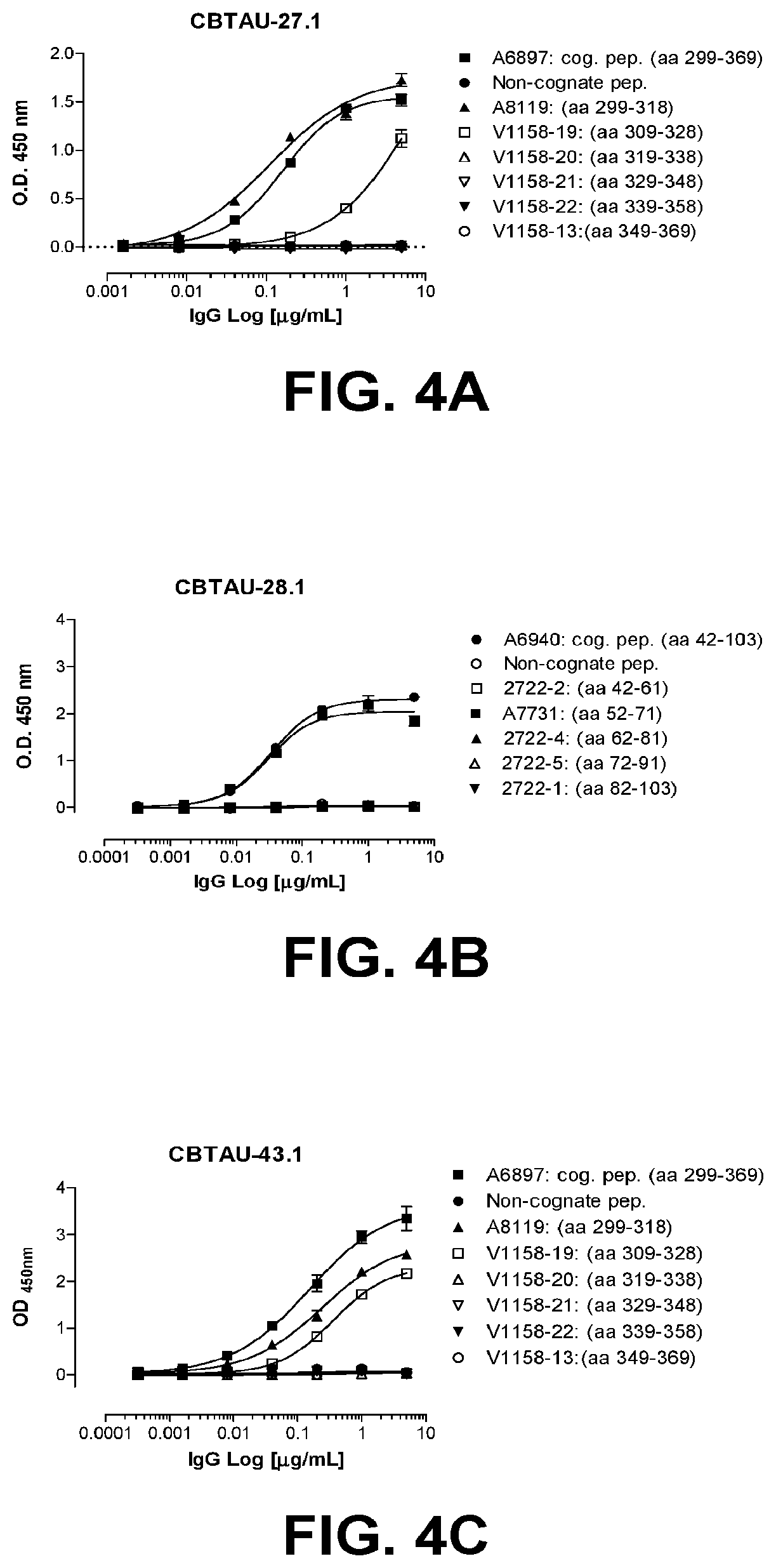
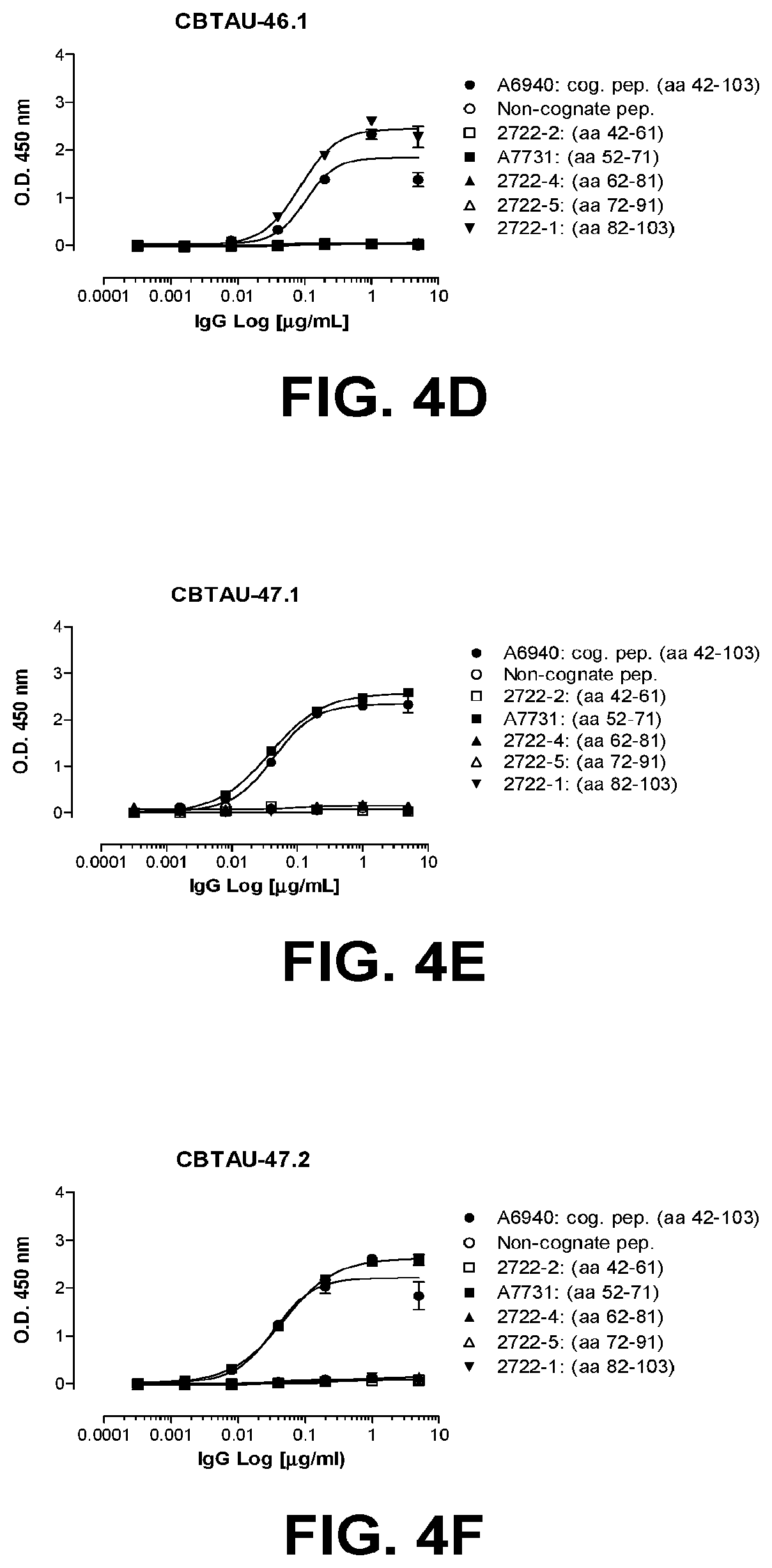
[0038] FIGS. 4A-4G show the epitope mapping of CBTAU-27.1, 28.1, 43.1, 46.1, 47.1, 47.2, and 49.1 using overlapping peptides that correspond to regions 42-103 and 299-369 on human tau441.
[0039] FIGS. 5A-5D show the immunohistochemical results for the CBTAU mAbs detailed in this application. FIGS. 5A and 5B show the immunostaining of CBTAU-7.1, 8.1, 16.1, 18.1, 20.1, 22.1, 24.1, 27.1, and 28.1 on non-AD versus AD hippocampal and cortical tissue sections, respectively. FIG. 5C shows the immunostaining of CBTAU-7.1, 8.1, 16.1, 18.1, 20.1, 22.1, and 24.1 on non-PSP and PSP cortical tissue sections. FIG. 5D shows the immunostaining of CBTAU43.1, 46.1, 47.2, and 49.1 against non-AD and AD cortical tissue sections.
[0040] FIG. 6A shows immunoreactivity of CBTAU-28.1 and control mAbs against non-AD (54 y.o. male; no clinical symptoms) and AD (93 y.o. Hispanic female) hippocampal tissue sections.
[0041] FIG. 6B shows reactivity of CBTAU-28.1 and control mAbs to AD (93 y.o. Hispanic female) hippocampal tissue sections with and without calf intestinal phosphatase treatment.
[0042] FIG. 7 shows reactivity of CBTAU-28.1 and phospho-tau mAb, ATB, against iPHF (immunopurified paired helical filaments) and calf intestinal phosphatase-treated iPHF samples.
[0043] FIGS. 8A-8F show reactivity of CBTAU-27.1, 28.1, 43.1, 46.1, 47.1, 47.2, and 49.1 to the tau phosphopeptides detailed on Tables 30-34.
DETAILED DESCRIPTION OF THE INVENTION
Definitions
[0044] The term "included" or "including" as used herein is deemed to be followed by the words "without limitation."
[0045] The term "tau" as used herein, is used interchangeably to specifically refer to the native monomer form of tau. The term "tau" is also used to generally identify other conformers of tau, for example, oligomers or aggregates of tau. The term "tau" is also used to refer collectively to all types and forms of tau. Due to alternative splicing, six tau isoforms are present in the human brain. These isoforms differ by the absence or presence of one or two 29-amino acid inserts encoded by exon 2 and 3 in the amino-terminal part, in combination with either three (R1, R3 and R4) or four (R1-R4) repeat-regions in the carboxy-terminal part. The microtubule-binding domain is encoded by exon 10. The adult tau isoforms include the longest 441-amino acid component (SEQ ID NO:1), or 4R/2N, the 410-amino acids component (SEQ ID NO:2), or 3R/2N, the 412-amino acids component (SEQ ID NO:3), or 4R/1N, the 381-amino acids component (SEQ ID NO:4), or 3R/1N and the 383-amino acids component (SEQ ID NO:5) or 4R/ON. The shortest 352-amino acids isoform (SEQ ID NO:6), or 3R/ON, is found in the fetal brain, and thus is referred to as fetal tau isoform.
[0046] The "wild type" tau amino acid sequence is represented by the 441 amino acid isoform (SEQ ID NO:1) also referred to as "tau441," "4R/2N," "hTau40," "TauF," "Tau-4" or "full-length tau."
[0047] The term "recombinant tau" herein, refers to the longest isoform of human brain Tau (SEQ ID NO:1) expressed in E. coli and purified to homogeneity or near homogeneity (S. Barghorn, Meth. Mol. Biol. 2004 299:35-51). Recombinant tau is soluble and is not phosphorylated.
[0048] The term "neurofibrillary tangle" (NFT) refers to the pathological structures first described by Alzheimer in the brain of dementia patients. NFT are composed of orderly arranged subunits called paired helical filament aggregates of hyperphosphorylated tau protein that are most commonly known as a primary marker of Alzheimer's Disease.
[0049] The term "paired helical filament-tau" or "PHF-tau" as used herein refers to well-known tau aggregates that make up the pathological structures called neurofibrillary tangles (NFT), first described by Alzheimer in the brain of dementia patients. Their presence is also found in numerous other diseases known as tauopathies.
[0050] "Enriched PHF-tau" or "ePHF-tau," is prepared according to the protocol of Greenberg and Davies as detailed in the Examples. PHF-tau is enriched from 27,200.times.g supernatants containing 0.8 M NaCl by taking advantage of their insolubility in zwitterionic detergents (K. S. Kosik et al. (1986) PNAS USA 83:4044-4048; R. Rubenstein et al. (1986) Brain Res. 372:80-88) and mercaptoethanol. PHFs isolated with zwitterionic detergents appear to maintain antigenic sites that may be lost during the isolation of SDS-insoluble NeuroFibular Tangles and are similar in structure and contain many antigenic properties to PHF in NFTs. "Immunopurified PHF-tau" or "iPHF-tau" is affinity purified with an anti-tau monoclonal antibody. Such protocols have provided PHF-tau preparations that retain the classical paired helical filament structure by electron microscopy and are completely soluble in low concentrations of SDS (G. Jicha, 1997, 48(2):128-32). PHF-tau is also formed from recombinant tau by induction of polymerization in vitro with heparin (Mandelkow, et al., Methods in Molecular Biology 299:35-51(2004). Alternatively, PHF-tau is isolated by various other methods from brains of patients having AD using protocols, such as described in Rostagna and Ghiso (A. Rostagna and J. Ghiso, Curr. Protoc. Cell. Biol. September 2009; CHAPTER: Unit-3.3333). The isolated PHF-tau is characterized for its purity and hyperphosphorylation status with antibodies known to react with PHF-tau. In a typical PHF-tau preparation, the hyperphosphorylated bands migrating at about 60, 64, 68 and 72 kDa in Western blot (Spillantini and Goedert, Trends Neurosci. 21:428-33, 1998) are detected by an AT8 antibody that specifically binds hyperphosphorylated PHF-tau but not dephosphorylated PHF-tau.
[0051] The term "antibodies" as used herein is meant in a broad sense and includes immunoglobulin or antibody molecules including polyclonal antibodies, monoclonal antibodies including murine, human, human-adapted, humanized and chimeric monoclonal antibodies, bispecific or multi-specific antibodies and antibody fragments. In general, antibodies are proteins or peptide chains that exhibit binding specificity to a defined antigen. Antibody structures are well known. Immunoglobulins can be assigned to five major classes, namely IgA, IgD, IgE, IgG and IgM, depending on the heavy chain constant domain amino acid sequence. IgA and IgG are further sub-classified as the isotypes IgA1, IgA2, IgG1, IgG2, IgG3 and IgG4. Antibody light chains of any vertebrate species can be assigned to one of two clearly distinct types, namely kappa (K) and lambda (X), based on the amino acid sequences of their constant domains.
[0052] The term "antigen-binding fragments" means a portion of an intact antibody. Examples of antibody fragments include Fab, Fab', F(ab')2 and Fv fragments, CDR, antigen-binding site, heavy or light chain variable region, diabodies, triabodies single chain antibody molecules (scFv) and multi-specific antibodies formed from at least two intact antibodies or fragments thereof or (poly) peptides that contain at least a fragment of an immunoglobin that is sufficient to confer antigen binding to the (poly) peptide, etc. An antigen-binding fragment may comprise a peptide or polypeptide comprising an amino acid sequence of at least 2, 5, 10, 15, 20, 25, 30, 35, 40, 50, 60, 70, 80, 90, 100, 125, 150, 175, 200, or 250 contiguous amino acid residues of the amino acid sequence of the antibody. The antigen-binding fragments may be produced synthetically or by enzymatic or chemical cleavage of intact immunoglobulins or they may be genetically engineered by recombinant DNA techniques. The methods of production are well known in the art and are described, for example, in Antibodies: A Laboratory Manual, edited by E. Harlow and D. Lane (1988), Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., which is incorporated herein by reference. An antibody or antigen-binding fragment thereof may have one or more binding sites. If there is more than one binding site, the binding sites may be identical to one another or they may be different.
[0053] An immunoglobulin light or heavy chain variable region consists of a "framework" region interrupted by "antigen-binding sites." The antigen-binding sites are defined using various terms as follows: (i) Complementarity-Determining Regions (CDRs) are based on sequence variability (Wu and Kabat, J. Exp. Med. 132:211-50, 1970). Generally, the antigen-binding site has three CDRs in each variable region (HCDR1, HCDR2 and HCDR3 in heavy chain variable region (VH) and LCDR1, LCDR2 and LCDR3 in light chain variable region (VL)) (Kabat et al., Sequences of Proteins of Immunological Interest, 5th Ed. Public Health Service, National Institute of Health, Bethesda, Md., 1991). (ii) The term "hypervariable region," "HVR," or "HV" refers to the regions of an antibody variable domain that are hypervariable in structure as defined by Chothia and Lesk (Chothia and Lesk, J. Mol. Biol. 96:901-17, 1987). Generally, the antigen-binding site has three hypervariable regions in each VH (H1, H2, H3) and VL (L1, L2, L3). Chothia and Lesk refer to structurally conserved HVs as "canonical structures." Numbering systems as well as annotation of CDRs and HVs have recently been revised by Abhinandan and Martin (Abhinandan and Martin, Mol. Immunol. 45:3832-9, 2008). (iii) Another definition of the regions that form the antigen-binding site has been proposed by Lefranc (Lefranc, et al., Dev. Camp. Immunol. 27:55-77, 2003) based on the comparison of V domains from immunoglobulins and T-cell receptors. The International ImMunoGeneTics (IMGT) database (http:_//www_imgt_org) provides a standardized numbering and definition of these regions. The correspondence between CDRs, HVs and IMGT delineations is described in Lefranc et al. The antigen-binding site can also be delineated based on Specificity-Determining Residue Usage (SDRU) (Almagro, J. Mol. Recognit. 17:13243, 2004), where Specificity-Determining Residues (SDR), refers to amino acid residues of an immunoglobulin that are directly involved in antigen contact.
[0054] Kabat et al. also defined a numbering system for variable domain sequences that is applicable to any antibody. One of ordinary skill in the art can unambiguously assign this system of "Kabat numbering" to any variable domain sequence, without reliance on any experimental data beyond the sequence itself. As used herein, "Kabat numbering" refers to the numbering system set forth by Kabat et al., U.S. Dept. of Health and Human Services, "Sequence of Proteins of Immunological Interest" (1983). Unless otherwise specified, references to the numbering of specific amino acid residue positions in an antibody or antigen-binding fragment, variant, or derivative thereof of the present disclosure are according to the Kabat numbering system, which, however, is theoretical and may not equally apply every antibody of the present disclosure. For example, depending on the position of the first CDR, the following CDRs might be shifted in either direction.
[0055] "Framework" or "framework sequence" are the remaining sequences within the variable region of an antibody other than those defined to be antigen-binding site sequences. Because the exact definition of an antigen-binding site can be determined by various delineations as described above, the exact framework sequence depends on the definition of the antigen-binding site.
[0056] The term "monoclonal antibody" (mAb) as used herein means an antibody (or antibody fragment) obtained from a population of substantially homogeneous antibodies. Monoclonal antibodies are highly specific, typically being directed against a single antigenic determinant.
[0057] In one aspect, the antibody of the present disclosure is a chimeric human antibody. Thus, in accordance with the present disclosure, the terms "human chimeric antibody" or "human recombinant antibody" and the like are used to denote a binding molecule, which antigen-binding features originated from a human cell, i.e., which antigen-binding site is derived from nucleic acids produced from a human cell such as a B cell, or the partial cDNA of which has been cloned from mRNA of a human cell, for example, a human memory B cell. A chimeric antibody is still "human" even if amino acid substitutions are made in the antibody, e.g., to improve biophysical or pharmacokinetic characteristics. Compared to artificially generated human-like antibodies such as single chain antibody fragments (scFvs) from a phage displayed antibody library or xenogeneic mice, the chimeric human antibody of the present disclosure is characterized by (i) the antigen-binding region being obtained using the human immune response rather than that of animal surrogates, i.e., the antigen-binding region has been generated in response to natural tau in its relevant conformation in the human body, and/or (ii) having protected the individual or is at least significant for the presence of tau.
[0058] Antibodies originating from human immunoglobulin libraries or from animals transgenic for one or more human immunoglobulins and that do not express endogenous immunoglobulins, as described infra and, for example, in, U.S. Pat. No. 5,939,598 by Kucherlapati et al., are denoted human-like antibodies in order to distinguish them from human-derived antibodies of the present disclosure.
[0059] For example, the pairing of heavy and light chains of human-like antibodies such as synthetic and semi-synthetic antibodies typically isolated from phage display do not necessarily reflect the original paring as it occurred in the original human B cell. Accordingly, Fab and scFv fragments obtained from recombinant expression libraries as commonly used in the prior art can be considered as being artificial with all possible associated effects on immunogenicity and stability. In contrast, the present disclosure provides antigen-binding regions of affinity-matured anti-tau antibodies from selected human subjects, which, in certain embodiments, are recombinantly expressed as chimeras with a common IgG1 constant region.
[0060] The term "functional variant," as used herein, refers to an antibody that comprises a nucleotide and/or amino acid sequence that is altered by one or more nucleotides and/or amino acids compared to the nucleotide and/or amino acid sequences of a reference antibody and that is capable of competing for specific binding to the binding partner, i.e., tau, with the reference antibody. In other words, the modifications in the amino acid and/or nucleotide sequence of the reference antibody do not significantly affect or alter the binding characteristics of the antibody encoded by the nucleotide sequence or containing the amino acid sequence, i.e., the antibody is still able to specifically recognize and bind its target. The functional variant may have conservative sequence modifications including nucleotide and amino acid substitutions, additions and deletions. Examples of functional variants include de-risking a free Cysteine or amino acid with potential post-translational modification in the hypervariable region, as well as Fc engineering to increase/decrease the binding affinity of IgG antibodies to FcRn, and increase/decrease serum half-life. A functional variant can also be generation of the antibody as a human chimeric or non-chimeric IgG2, IgG3 or IgG4 isotype, or as a chimeric or non-chimeric isotype of a different species. A functional variant can also be a mutation or mutations of the constant regions for enhancement of bispecific antibody formation. These modifications can be introduced by standard techniques known in the art, such as PCR, site-directed mutagenesis and random PCR-mediated mutagenesis, and may comprise natural as well as non-natural nucleotides and amino acids.
[0061] The term "specifically binding" or "specifically recognize," as used herein, in reference to the interaction of an antibody and its binding partner, e.g., an antigen, means that the interaction is dependent upon the presence of a particular amino acid sequence or structure, e.g., an antigenic determinant or epitope, on the binding partner. In other words, the antibody preferentially binds or recognizes the binding partner, even when the binding partner is present in a mixture of other molecules or organisms. The binding may be mediated by covalent or noncovalent interactions or a combination of both. In yet other words, the term "specifically binding" or "specifically recognizes" means that the antibody is specifically immunoreactive with an antigenic determinant or epitope and is not immunoreactive with other antigenic determinants or epitopes. An antibody that (immuno)specifically binds to an antigen may bind to other peptides or polypeptides with lower affinity as determined by, e.g., radioimmunoassays (MA), enzyme-linked immunosorbent assays (ELISA), BIACORE, or other assays known in the art. Antibodies or fragments thereof that specifically bind to an antigen may be cross-reactive with related antigens, carrying the same epitope. Preferably, antibodies or fragments thereof that specifically bind to an antigen do not cross-react with other antigens.
[0062] The term "epitope" as used herein means that part of the antigen that is contacted by the CDR loops of antibody. A "structural epitope" comprises about 15-22 contact residues on the antigen surface and involves many amino acid residues that make contact with a large group of residues on CDRs collectively referred to as the paratope of antibody. Direct contact between epitope and paratope residues is established through electrostatic forces such as hydrogen bonds, salt bridges, van der Waals forces of hydrophobic surfaces and shape complementarity. The interface has also bound water molecules or other co-factors that contribute to the specificity and affinity of antigen-antibody interactions. The binding energy of an antigen-antibody complex is primarily mediated by a small subset of contact residues in the epitope-paratope interface. These "energetic residues" are often located in the center of the epitope-paratope interface and make up the functional epitope. Contact residues in the periphery of the interface make generally minor contributions to the binding energy; their replacements have frequently little effect on the binding with antigen. Thus, the binding or functional activity of an epitope involves a small subset of energetic residues centrally located in the structural epitope and contacted by the specificity-determining CDRs. The assignment of a functional epitope on an antigenic protein can be made using several methods including Alanine scan mutagenesis or by solving the crystal structure of the antigen with the antibody.
[0063] An epitope can be linear in nature or can be a discontinuous epitope, e.g., a conformational epitope, which is formed by a spatial relationship between non-contiguous amino acids of an antigen rather than a linear series of amino acids. A conformational epitope includes epitopes resulting from folding of an antigen, where amino acids from differing portions of the linear sequence of the antigen come in close proximity in three-dimensional space. For discontinuous epitopes, it may be possible to obtain binding of one or more linear peptides with decreased affinity to a so-called partial epitope, e.g., dispersed at different regions of the protein sequence (M. S. Cragg, (2011) Blood 118 (2):219-20).
[0064] As used herein, the term "affinity" refers to a measure of the strength of the binding of an individual epitope or partial epitope with the CDRs of a binding molecule, e.g., an immunoglobulin molecule; see, e.g., Harlow et al., Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory Press, 2nd ed. (1988) at pages 27-28. As used herein, the term "avidity" refers to the overall stability of the complex between a population of immunoglobulins and an antigen, that is, the functional combining strength of an immunoglobulin mixture with the antigen; see, e.g., Harlow at pages 29-34. Avidity is related to both the affinity of individual immunoglobulin molecules in the population with specific epitopes, and also the valences of the immunoglobulins and the antigen. For example, the interaction between a bivalent monoclonal antibody and an antigen with a highly repeating epitope structure, such as a polymer, would be one of high avidity. The affinity or avidity of an antibody for an antigen can be determined experimentally using any suitable method; see, for example, Berzofsky et al., "Antibody-Antigen Interactions" in Fundamental Immunology, W. E. Paul, Ed., Raven Press New York, N.Y. (1984); J. Kuby, Immunology, W.H. Freeman and Company New York, N.Y. (1992), and methods described herein. General techniques for measuring the affinity of an antibody for an antigen include ELISA, RIA, and surface plasmon resonance. The measured affinity of a particular antibody-antigen interaction can vary if measured under different conditions, e.g., salt concentration, pH. Thus, measurements of affinity and other antigen-binding parameters, e.g., KD, IC50, are preferably made with standardized solutions of antibody and antigen, and a standardized buffer.
[0065] Antibodies or antigen-binding fragments or variants thereof of the disclosure may also be described or specified in terms of their ability to specifically detect the presence of antigen. The term "detect" or "detecting" is used in the broadest sense to include quantitative, semi-quantitative or qualitative measurements of a target molecule. In one aspect, antibodies described herein may only determine the presence or absence of tau polypeptide in a biological sample, e.g., by immunohistochemistry and, thus, the tau polypeptide is detectable or, alternatively, undetectable in the sample as determined by the method.
[0066] The term "phospho-specific antibody" or "phospho-dependent antibody" herein used means a specific antibody in which at least part or the entire epitope relies on a phosphorylated amino acid residue. A phospho-specific or phospho-dependent antibody does not detect un-phosphorylated antigen. The term "phospho-selective antibody" means a specific antibody that preferentially binds to the phosphorylated residue and has higher affinity to the phosphorylated versus the non-phosphorylated antigen. The term "non-phospho-selective antibody" or "phospho-independent" means a specific antibody that preferentially binds to the non-phosphorylated residue and has higher affinity to the non-phosphorylated versus the phosphorylated antigen. In certain embodiments, the anti-tau antibodies of the disclosure, or antigen-binding fragments thereof, are phospho-dependent. In certain other embodiments, the anti-tau antibodies of the disclosure, or antigen-binding fragments thereof, are phospho-independent.
[0067] The term "polynucleotide" is intended to encompass a singular nucleic acid as well as plural nucleic acids, and refers to an isolated nucleic acid molecule or construct, e.g., messenger RNA (mRNA) or plasmid DNA (pDNA). A polynucleotide may comprise a conventional phosphodiester bond or a non-conventional bond (e.g., an amide bond, such as found in peptide nucleic acids (PNA)). The term "nucleic acid molecule" refers to any one or more nucleic acid segments, e.g., DNA or RNA fragments, present in a polynucleotide. By "isolated" nucleic acid or polynucleotide is intended a nucleic acid molecule, DNA or RNA that has been removed from its native environment. For example, a recombinant polynucleotide encoding an antibody contained in a vector is considered isolated for the purposes of this disclosure.
[0068] In certain embodiments, the polynucleotide or nucleic acid is DNA. In the case of DNA, a polynucleotide comprising a nucleic acid that encodes a polypeptide normally may include a promoter and/or other transcription or translation control elements operably associated with one or more coding regions. An operable association is when a coding region for a gene product, e.g., a polypeptide, is associated with one or more regulatory sequences in such a way as to place expression of the gene product under the influence or control of the regulatory sequence(s). Thus, a promoter region would be operably associated with a nucleic acid encoding a polypeptide if the promoter was capable of effecting transcription of that nucleic acid. The promoter may be a cell-specific promoter that directs substantial transcription of the DNA only in predetermined cells. Other transcription control elements, besides a promoter, for example, enhancers, operators, repressors, and transcription termination signals, can be operably associated with the polynucleotide to direct cell-specific transcription. Suitable promoters and other transcription control regions are disclosed herein.
[0069] As used herein, the terms "treat" or "treatment" refer to both therapeutic treatment and prophylactic or preventative measures, wherein the object is to prevent or slow down (lessen) an undesired physiological change or disorder, such as the development of Parkinsonism or Alzheimer's Disease. Beneficial or desired clinical results include, but are not limited to, alleviation of symptoms, diminishment of extent of disease, stabilized (i.e., not worsening) state of disease, delay or slowing of disease progression, amelioration or palliation of the disease state, and remission (whether partial or total), whether detectable or undetectable. "Treatment" can also mean prolonging survival as compared to expected survival if not receiving treatment. Those in need of treatment include those already with the condition or disorder as well as those prone to have the condition or disorder or those in which the manifestation of the condition or disorder is to be prevented. A "medicament" as used herein, is an agent used in the treatment of an undesirable physiological change or disorder.
[0070] By "subject" or "individual" or "animal" or "patient" or "mammal" is meant any subject, particularly a mammalian subject, e.g., a human patient, for whom diagnosis, prognosis, prevention, or therapy is desired.
DESCRIPTION
[0071] Tau is an abundant central and peripheral nervous system protein having multiple well-known isoforms. In the human CNS, six major tau isoforms ranging in size from 352 to 441 exist due to alternative splicing (Hanger, et al., Trends Mol. Med. 15:112-9, 2009). These isoforms differ from each other by the regulated inclusion of 0-2 N-terminal inserts, and 3 or 4 tandemly arranged microtubule-binding repeats, and are referred to as ON3R (SEQ ID NO:6), 1N3R (SEQ ID NO:4), 2N3R (SEQ ID NO:2), ON4R (SEQ ID NO:5), 1N4R (SEQ ID NO:3) and 2N4R (SEQ ID NO:1). The term "recombinant tau" as used herein refers to the tau isoform of SEQ ID NO:1 that is devoid of phosphorylation and other post-translational modifications. The tau protein can be recombinantly expressed in high quantities, for example, in E. coli, baculovirus, mammalian or cell-free systems. "Recombinant tau" may be recombinantly expressed and purified using standard methods. (Barghorn, et al. 2004, Meth. Mol. Biol. 35-51).
[0072] Tau binds microtubules and regulates transport of cargo through cells, a process that can be modulated by tau phosphorylation that occurs at many of the 79 potential serine (Ser) and threonine (Thr) phosphorylation sites. Tau is highly phosphorylated during brain development. The degree of phosphorylation declines in adulthood. Some of the phosphorylation sites are located within the microtubule binding domains of tau, and it has been shown that an increase of tau phosphorylation negatively regulates the binding of microtubules. For example, Ser262 and Ser396, which lie within or adjacent to microtubule binding motifs, are hyperphosphorylated in the tau proteins of the abnormal paired helical filaments (PHFs), a major component of the neurofibrillary tangles (NFTs) in the brain of AD patients. PHFs are filamentous aggregates of tau proteins that are abnormally hyperphosphorylated and can be stained with specific anti-tau antibodies and detected by light microscopy. The same holds true for so-called straight tau filaments. PHFs form twisted ribbons consisting of two filaments twisted around one another with a periodicity of about 80 nm.
[0073] These pathological features are commonly referred to as "tau-pathology," "tauopathology" or "tau-related pathology." For a more detailed description of neuropathological features of tauopathies, refer to Le et al., Annu. Rev. Neurosci. 24 (2001), 1121-1159, and Gotz, Brain. Res. Rev. 35 (2001), 266-286, the disclosure content of which is incorporated herein by reference. Physiological tau protein stabilizes microtubules in neurons. Pathological phosphorylation leads to abnormal tau localization and aggregation, which causes destabilization of microtubules and impaired cellular transport. Aggregated tau is neurotoxic in vitro (Khlistunova et al., J. Biol. Chem. 281 (2006), 1205-1214). The exact neurotoxic species remains unclear, however, as do the mechanism(s) by which they lead to neuronal death. Aggregates of tau can be observed as the main component of neurofibrillary tangles (NFT) in many tauopathies, such as Alzheimer's disease (AD), Frontotemporal dementias, supranuclear palsy, Pick's disease, Argyrophilic grain disease (AGD), corticobasal degeneration, FTDP-17, Parkinson's disease, Dementia pugilistica (reviewed in Gendron and Petrucelli, Mol. Neurodegener. 4:13 (2009)). Besides these observations, evidence emerges that tau-mediated neuronal death can occur even in the absence of tangle formation. Soluble phospho-tau species are present in CSF (Aluise et al., Biochim. Biophys. Acta. 1782 (2008), 549-558). Tau aggregates can transmit a misfolded state from the outside to the inside of a cell and transfer between co-cultured cells (Frost et al., J. Biol. Chem. 284 (2009), 12845-12852).
[0074] In addition to the involvement in neurodegenerative tauopathies, observed alterations in tau phosphorylation during and after ischemia/reperfusion and after concussive head injury suggest tau plays a crucial role in neuronal damage and clinical pathophysiology of neurovascular disorders such as ischemic stroke (Zheng et al., J. Cell. Biochem. 109 (2010), 26-29), as well as changes in tau found in chronic traumatic encephalopathy, a tauopathy in concussed athletes and military veterans with Traumatic Brain Injury (BI).
[0075] This disclosure provides monoclonal antibodies, wherein the antibodies bind tau deposits in human AD brain tissue.
[0076] In certain embodiments, the antibodies a) bind tau deposits in human AD brain tissue and b) do not bind tau in normal brain tissue.
[0077] In certain embodiments, the antibodies a) bind tau deposits in human AD brain tissue, b) do not bind tau in normal human brain tissue, and c) do not bind tau deposits in Progressive Supranuclear Palsy (PSP) brain tissue.
[0078] In certain embodiments, the antibodies a) form an immunological complex with tau deposits in human AD brain tissue and b) do not form an immunological complex with tau in normal human brain tissue.
[0079] In certain embodiments, the antibodies are chimeric antibodies.
[0080] In certain embodiments, the antibodies are chimeric antibodies comprising an antigen-binding variable region from a human antibody that binds specifically to tau and a recombinant constant region of a human IgG1, and wherein the chimeric antibodies are different from the human antibody.
[0081] In certain embodiments, the antibodies are chimeric antibodies comprising an antigen-binding variable region from a human antibody that binds specifically to tau and a recombinant constant region of a human IgG1, wherein the constant region of the chimeric antibody differs from the constant region of the human antibody.
[0082] In certain embodiments, the antibodies are chimeric antibodies comprising a naturally occurring human antigen-binding variable region that binds specifically to tau and a recombinant constant region of a human IgG1 antibody.
[0083] In certain embodiments, the antibodies are chimeric antibodies comprising naturally occurring human light and heavy chain variable regions from a human antibody and recombinant human IgG1 heavy and light chain constant regions.
[0084] In certain embodiments, the antibodies are chimeric antibodies comprising heavy and light chain variable regions from a naturally occurring human antibody and recombinant human IgG1 heavy and light chain constant regions.
[0085] In certain embodiments, the antibodies are chimeric antibodies comprising heavy and light chain variable regions from a human antibody and recombinant human IgG1 heavy and light chain constant regions.
[0086] In certain embodiments, the antibodies are not naturally occurring.
[0087] The human anti-tau antibodies disclosed herein specifically bind tau and epitopes thereof and to various conformations of tau and epitopes thereof. For example, disclosed herein are antibodies that specifically bind pathologically modified tau isoforms and tau aggregates. In one example, a tau antibody disclosed herein binds to tau or an epitope thereof and shows no binding above about three times background for other proteins. An antibody that "specifically binds" or "selectively binds" a tau conformer refers to an antibody that does not bind all conformations of tau, i.e., does not bind at least one other tau conformer such as recombinant tau.
[0088] The variable domains of the chimeric anti-tau monoclonal antibodies of this disclosure may have origin from a pool of healthy human subjects exhibiting a tau-specific immune response or from subject affected with Alzheimer's Disease. The tau antibodies of this disclosure may also be called "human-derived antibodies" in order to emphasize that those antibody antigen-binding regions were indeed expressed by the subjects and have not been isolated from, for example, a human immunoglobulin-expressing phage library, which hitherto represented one common method for trying to provide human-like antibodies. For example, the antibodies of this disclosure differ from mAb AT8, MCI, and AT100, in that they are human-derived antibodies.
[0089] The anti-tau antibodies of this disclosure can be characterized by their binding properties to recombinant tau in an ELISA. Recombinant tau purified from E. coli is highly soluble owing to its hydrophilic character. It lacks phosphorylation of Ser, Thr and Tyr residues characteristic of tau found in tauopathies. In one embodiment of the disclosure, the anti-tau monoclonal antibodies disclosed herein do not specifically bind recombinant tau in an ELISA. In another embodiment of the disclosure, the anti-tau monoclonal antibodies bind to recombinant tau in an ELISA.
[0090] In an embodiment, the anti-tau antibody of the disclosure has been shown to specifically bind to a phosphorylated-tau peptide of SEQ ID NO:315 or SEQ ID NO:365. In another embodiment, the anti-tau antibody of the disclosure has been shown to bind to both phosphorylated tau peptide of SEQ ID NO:317 and non-phosphorylated peptide of tau SEQ ID NO:318 or phosphorylated peptide of SEQ ID NO:329 and non-phosphorylated tau peptide of SEQ ID NO:367.
[0091] The anti-tau antibodies disclosed herein in certain embodiments specifically bind tau peptide in a peptide ELISA. In one embodiment, an anti-tau antibody binds to a tau peptide, e.g., GTPGSRSRTPSLPTPPTR (SEQ ID NO:318), corresponding to amino acids 204-221 of tau441. In another embodiment, an anti-tau antibody binds to a tau peptide, e.g., GEPPK SGDRSGYS SP GSP GTP GSRSRTP SLP TPPTREPKKVAVVRTPPK SP S SAKSRLQTAPVP MPDL (SEQ ID NO:367), corresponding to amino acids 221-253 of tau441. The anti-tau antibody of the disclosure is unlike previously disclosed human anti-tau monoclonal antibodies (US 2013/0295021), which have been reported to bind to different tau peptides. Monoclonal antibodies NI-105.4E4, NI-105.4A3 and NI-105.4E4 bind to peptides 329-351+387-397, 337-343, and 35-49 of tau441, respectively.
[0092] Anti-tau antibodies of the disclosure can be characterized by their binding properties to PHF-tau in an ELISA. Anti-PHF-tau antibody clone AT8 binds to PHF-tau and has been used extensively to detect PHF-tau in neurofibrillary tangles in samples from Alzheimer's patients. AT8 is a phospho-specific monoclonal antibody and binds to phosphorylated Ser202 and Thr 205 of PHF-tau and is well-published for use in ELISA, immunohistochemistry, immunoblot, Western blot, and related applications. Clone AT8 recognizes Alzheimer Disease tau, as well as PHF-tau by ELISA and does not bind non-phosphorylated-tau from healthy individuals or recombinant tau. In one example, the anti-tau antibody of the disclosure does not bind to PHF-tau by ELISA. In another example, the anti-tau antibody of the disclosure binds to PHF-tau by ELISA.
[0093] Anti-tau antibodies of the disclosure can be characterized by their binding properties to PHF-tau and recombinant tau by Western blot. In neurodegenerative disorders, several mechanisms (phosphorylation, ubiquitination, oxidation, glycation) are involved in the aggregation of tau proteins into PHF. These pathological tau proteins are visualized by Western blotting as three major bands between 55 and 69 kDa, and a minor band at 74 kDa. Tau 55 results from the phosphorylation of the shortest isoform (SEQ ID NO:6), tau 64 from the phosphorylation of tau variants with one cassette exon (SEQ ID NO:4 and/or SEQ ID NO:5), tau 69 from the phosphorylation of tau variants with two cassette exons (SEQ ID NO:2 and/or SEQ ID NO:3). Phosphorylation of the longest tau isoform (SEQ ID NO:1) induces the formation of the additional hyperphosphorylated tau74 variant. In certain embodiments, the anti-tau antibodies bind to PHF-tau by Western analysis.
[0094] Anti-tau antibodies can be utilized in and characterized by immunohistochemistry (IHC) of tissue sections from normal or AD brain. Phospho-tau antibodies, in particular, highlight neurofibrillary pathology with a high degree of sensitivity and specificity, whereas, no detection of tau in normal healthy brain is observed. Clinicopathological studies have demonstrated that phospho-tau deposits or accumulations correspond more closely to clinical signs compared to amyloid-f3 accumulations, and progress in a stepwise fashion from transentorhinal, to limbic, to isocortical areas, forming the basis for AD staging [R. J. Castellani, et al., Acta Neuropathol. (Berl) 111:503(2006); H. Braak and E. Braak, Acta Neuropathol. (Berl) 82:239 (1991 Tau monoclonal antibodies that are commonly used in immunohistochemistry include AT8 (p202/p205 tau), AT180 (p231 tau), AT270 (p181 tau), AT100 (pT212 and S214), and MC-1, (M. Mercken et al., 1992 Acta Neuropathol. 84:265-272; Zheng-Fischhofer 1998 Eur. I Biochem. 252:542-552, M. Goedert et. al., 1994 Biochem. 1301:871-877). In one embodiment, the anti-tau monoclonal antibodies of the disclosure detect tau in normal (i.e., healthy) human brain tissue and detect tau deposits in human
[0095] AD brain tissue. In another example, the anti-tau antibodies of the disclosure detect tau deposits in human AD brain but do not detect tau in normal human brain.
[0096] Anti-tau antibodies of the disclosure can also be utilized in and characterized by immunohistochemistry of additional tauopathies, including Progressive Supranuclear Palsy, Pick's Disease and others. The pathological filamentous-tau inclusions in PSP are composed of aberrantly phosphorylated-tau proteins, but there is a preferential accumulation of abnormal 4R tau isoforms. A panel of anti-tau monoclonal antibodies, including Alz50, Tau-2, T46, PHF-1, PHF-6, 12E8, PHF-1, RD4 and ATB, has been used to characterize PSP deposits (J. Neuropathol. Exp. Neurol. 1998 (6):588-601). All of the monoclonal antibodies stained intraneuronal and glial inclusions, however, 12E8 and PHF-6 stained with less intensity. These antibodies detect different epitopes of tau, e.g., phospho-specific, isoform-specific, and also detect tau deposits in AD brain. RD3, an anti-tau monoclonal antibody that specifically detects the 3-repeat tau isoform, shows limited IHC detection of PSP, yet intensely stains tau deposits in human AD brain tissue. The limited detection of PSP by this antibody is due to reduced levels of the 3-repeat tau isoform in PSP (R. De Silva et al., Neuropath. and Appl. Neurobio. (2003) 29(3)288-302). In one embodiment, the anti-tau antibodies of the disclosure detect tau deposits in human AD brain but do not detect tau in normal human brain and do not detect tau deposits in human PSP brain.
[0097] In certain embodiments, the antibody comprises a heavy chain comprising a) a heavy chain CDR1 region of SEQ ID NO:163, a heavy chain CDR2 region of SEQ ID NO:164, and a heavy chain CDR3 region of SEQ ID NO:165, b) a heavy chain CDR1 region of SEQ ID NO:169, a heavy chain CDR2 region of SEQ ID NO:170, and a heavy chain CDR3 region of SEQ ID NO:171, c) a heavy chain CDR1 region of SEQ ID NO:175, a heavy chain CDR2 region of SEQ ID NO:176, and a heavy chain CDR3 region of SEQ ID NO:177, d) a heavy chain CDR1 region of SEQ ID NO:181, a heavy chain CDR2 region of SEQ ID NO:182, and a heavy chain CDR3 region of SEQ ID NO:183, e) a heavy chain CDR1 region of SEQ ID NO:185, a heavy chain CDR2 region of SEQ ID NO:186, and a heavy chain CDR3 region of SEQ ID NO:187, f a heavy chain CDR1 region of SEQ ID NO:190, a heavy chain CDR2 region of SEQ ID NO:191, and a heavy chain CDR3 region of SEQ ID NO:192, g) a heavy chain CDR1 region of SEQ ID NO:196, a heavy chain CDR2 region of SEQ ID NO:197, and a heavy chain CDR3 region of SEQ ID NO:198, h) a heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, i) a heavy chain CDR1 region of SEQ ID NO:219, a heavy chain CDR2 region of SEQ ID NO:220, and a heavy chain CDR3 region of SEQ ID NO:221.
[0098] In certain embodiments, the antibody comprises a light chain comprising: a) a light chain CDR1 region of SEQ ID NO:166, a light chain CDR2 region of SEQ ID NO:167, and a light chain CDR3 region of SEQ ID NO:168, b) a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:174, c) a light chain CDR1 region of SEQ ID NO:178, a light chain CDR2 region of SEQ ID NO:179, and a light chain CDR3 region of SEQ ID NO:180, d) a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:184, e) a light chain CDR1 region of SEQ ID NO:188, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:189, f a light chain CDR1 region of SEQ ID NO:193, a light chain CDR2 region of SEQ ID NO:194, and a light chain CDR3 region of SEQ ID NO:195, g) a light chain CDR1 region of SEQ ID NO:199, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:200, h) a light chain CDR1 region of SEQ ID NO:216, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:217, and i) a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:217.
[0099] In certain embodiments, the antibody is selected from the group consisting of a) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:163, a heavy chain CDR2 region of SEQ ID NO:164, and a heavy chain CDR3 region of SEQ ID NO:165, a light chain CDR1 region of SEQ ID NO:166, a light chain CDR2 region of SEQ ID NO:167 and a light chain CDR3 region of SEQ ID NO:168; b) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:169, a heavy chain CDR2 region of SEQ ID NO:170, and a heavy chain CDR3 region of SEQ ID NO:171, a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:174; c) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:175, a heavy chain CDR2 region of SEQ ID NO:176, and a heavy chain CDR3 region of SEQ ID NO:177, a light chain CDR1 region of SEQ ID NO:178, a light chain CDR2 region of SEQ ID NO:179, and alight chain CDR3 region of SEQ ID NO:180; d) a heavy chain CDR1 region of SEQ ID NO:181, a heavy chain CDR2 region of SEQ ID NO:182, and a heavy chain CDR3 region of SEQ ID NO:183, a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:184; e) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:185, a heavy chain CDR2 region of SEQ ID NO:186, and a heavy chain CDR3 region of SEQ ID NO:187, a light chain CDR1 region of SEQ ID NO:188, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:189; f an antibody comprising a heavy chain CDR1 region of SEQ ID NO:190, a heavy chain CDR2 region of SEQ ID NO:191, and a heavy chain CDR3 region of SEQ ID NO:192, a light chain CDR1 region of SEQ ID NO:193, a light chain CDR2 region of SEQ ID NO:194 and a light chain CDR3 region of SEQ ID NO:195; g) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:196, a heavy chain CDR2 region of SEQ ID NO:197, and a heavy chain CDR3 region of SEQ ID NO:198, a light chain CDR1 region of SEQ ID NO:199, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:200; h) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, a light chain CDR1 region of SEQ ID NO:216, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:217; i) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:174, and a light chain CDR3 region of SEQ ID NO:217; j) an antibody comprising a heavy chain CDR1 region of SEQ ID NO:219, a heavy chain CDR2 region of SEQ ID NO:220, and a heavy chain CDR3 region of SEQ ID NO:221, a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:173, and a light chain CDR3 region of SEQ ID NO:217.
[0100] In certain embodiments, the antibody comprises a heavy chain variable region selected from the group consisting of the amino acid sequence of SEQ ID NOS:87, 91, 95, 99, 103, 107, 111, 123, 127 and 131. In certain embodiments, the antibody comprises a light chain variable region selected from the group consisting of the amino acid sequence of SEQ ID NOS:88, 92, 96, 100, 104, 108, 112, 124, 128, 132.
[0101] In certain embodiments, antigen-binding fragments of the above-described antibodies are provided. The antigen-binding fragments preferably bind to the same epitope. The anti-tau monoclonal antibodies and antigen-binding fragments of the present disclosure bind to different epitopes as compared to the epitopes of known human anti-tau antibodies, such as, e.g., NI-105.4E4, and NI-105.4A3. With "binding to a different epitope," it is meant that the antibody binds to different critical amino acid residues as compared to known antibodies.
[0102] In certain embodiments, the antibodies act synergistically when used in combination with other antibodies binding to tau protein. As used herein, the term "synergistic" means that the combined effect of the antibodies or antigen-binding fragments when used in combination is greater than their additive effects when used individually. A way of calculating synergy is by means of the combination index. The concept of the combination index (CI) has been described by Chou and Talalay (Adv. Enzyme Regul. 22:27-55, 1984).
[0103] In certain embodiments, the antibodies and antigen-binding fragments are for use as a medicament, and preferably for use in the diagnostic, therapeutic and/or prophylactic treatment of neurodegenerative diseases. Human anti-tau antibodies of the disclosure or fragments thereof, including Fab, (Fab')2, scFv fragments, or antibodies comprising antigen-binding sites of the antibodies of the disclosure can be used to treat, reduce or prevent symptoms in patients having a neurodegenerative disease that involves accumulation of tau or pathological tau or tau aggregation within the brain, such as patients suffering from AD, as well as any other tauopathy or other tau-related pathologies in which tau may be overexpressed. While not wishing to be bound by any particular theory, the antibodies of the disclosure may exert their beneficial effect by reducing or eliminating pathological tau or tau aggregation and hence the amount of PHF-tau in the brain. The antibodies of the disclosure may be used to treat an animal patient belonging to any classification. Examples of such animals include mammals such as humans, rodents, dogs, cats and farm animals. For example, the antibodies of the disclosure are useful in the preparation of a medicament for treatment of AD wherein the medicament is prepared for administration in dosages defined herein.
[0104] Another embodiment of the disclosure is a method of reducing aggregation of tau in patients in need thereof comprising administering to the patient a therapeutically effective amount of the isolated antibody of the disclosure for a time sufficient to reduce the aggregation of tau. Another embodiment of the disclosure is a method of treating or reducing symptoms of a neurodegenerative disease that involves aggregation of tau in a patient comprising administering to the patient a therapeutically effective amount of the isolated antibody of the disclosure for a time sufficient to treat or reduce symptoms of the neurodegenerative disease. Another embodiment of the disclosure is a method of reducing tau in patients in need thereof comprising administering to the patient a therapeutically effective amount of the isolated antibody of the disclosure for a time sufficient to reduce tau.
[0105] In any of the embodiments above, the neurodegenerative disease that involves aggregation of tau is a tauopathy. As used herein, a "tauopathy" encompasses any neurodegenerative disease that involves the pathological aggregation of tau within the brain. In addition to familial and sporadic AD, other exemplary tauopathies are frontotemporal dementia with Parkinsonism linked to chromosome 17 (FTDP-17), progressive supranuclear palsy, corticobasal degeneration, Pick's disease, progressive subcortical gliosis, tangle only dementia, diffuse neurofibrillary tangles with calcification, argyrophilic grain dementia, amyotrophic lateral sclerosis, Parkinsonism-dementia complex, Down syndrome, Gerstmann-Straussler Scheinker disease, Hallervorden-Spatz disease, inclusion body myositis, Creutzfeld-Jakob disease, multiple system atrophy, Niemann-Pick disease type C, prion protein cerebral amyloid angiopathy, subacute sclerosing panencephalitis, myotonic dystrophy, nonguanamian motor neuron disease with neurofibrillary tangles, post-encephalitic Parkinsonism, and chronic traumatic encephalopathy, such as dementia pugilistica (boxing disease) (Morris, et al., Neuron. 70:410-26, 2011).
[0106] A tauopathy-related behavioral phenotype includes cognitive impairments, early personality change and disinhibition, apathy, abulia, mutism, apraxia, perseveration, stereotyped movements/behaviors, hyperorality, disorganization, inability to plan or organize sequential tasks, selfishness/callousness, antisocial traits, a lack of empathy, halting, agrammatic speech with frequent paraphasic errors but relatively preserved comprehension, impaired comprehension and word-finding deficits, slowly progressive gait instability, retropulsion, freezing, frequent falls, non-levodopa responsive axial rigidity, supranuclear gaze palsy, square wave jerks, slow vertical saccades, pseudobulbar palsy, limb apraxia, dystonia, cortical sensory loss, and tremor.
[0107] Patients amenable to treatment include asymptomatic individuals at risk of AD or other tauopathy, as well as patients presently showing symptoms. Patients amenable to treatment include individuals who have a known genetic risk of AD, such as a family history of AD or presence of genetic risk factors in the genome. Exemplary risk factors are mutations in the amyloid precursor protein (APP), especially at position 717 and positions 670 and 671 (Hardy and Swedish mutations, respectively). Other risk factors are mutations in the presenilin genes, PS1 and PS2, and ApoE4, family history of hypercholesterolemia or atherosclerosis. Individuals presently suffering from AD can be recognized from characteristic dementia by the presence of risk factors described above. In addition, a number of diagnostic tests are available to identify individuals who have AD. These include measurement of cerebrospinal fluid tau and Ar342 levels. Elevated tau and decreased Af342 levels signify the presence of AD. Individuals suffering from AD can also be diagnosed by AD and Related Disorders Association criteria.
[0108] Another embodiment of the disclosure is a method of reducing tau in patients in need thereof comprising administering to the patient a therapeutically effective amount of the isolated anti-tau antibody of the disclosure for a time sufficient to reduce tau. Patients amenable to treatment may suffer from an ailment associated with overexpression of tau. Some mutations, including mutations in intron 10, induce increased levels of the functionally normal four-repeat tau protein isoform, leading to neurodegeneration. Overexpression of the four-repeat human tau protein isoform specifically in neurons in a transgenic mouse led to development of axonal degeneration in brain and spinal cord. In the model, axonal dilations with accumulation ofneurofilaments, mitochondria, and vesicles were documented. The axonopathy and the accompanying dysfunctional sensorimotor capacities were transgene-dosage related. These findings proved that merely increasing the concentration of the four-repeat tau protein isoform is sufficient to injure neurons in the central nervous system, without formation of intraneuronal neurofibrillary tangles (Spittaels, et al., Am. J. Pathology 155(6):2153-2165, 1999).
[0109] Administration/Pharmaceutical Compositions
[0110] Anti-tau antibodies of the disclosure are suitable, both as therapeutic and prophylactic agents for treating or preventing neurodegenerative diseases that involves accumulation of tau, and/or pathological aggregation of tau, such as AD or other tauopathies or tau-associated ailments. In asymptomatic patients, treatment can begin at any age (e.g., at about 10, 15, 20, 25, 30 years). Usually, however, it is not necessary to begin treatment until a patient reaches about 40, 50, 60, or 70 years. Treatment typically entails multiple dosages over a period of time. Treatment can be monitored by assaying antibody, or activated T-cell or B-cell responses to the therapeutic agent over time. If the response falls, a booster dosage is indicated.
[0111] In prophylactic applications, pharmaceutical compositions or medicaments are administered to a patient susceptible to, or otherwise at risk of, AD or other ailments involving tau, in an amount sufficient to eliminate or reduce the risk, lessen the severity, or delay the outset of the disease, including biochemical, histologic and/or behavioral symptoms of a disease, its complications and intermediate pathological phenotypes presented during development of the disease. In therapeutic applications, compositions or medicaments are administered to a patient suspected of, or already suffering from, such a disease in an amount sufficient to reduce, arrest, or delay any of the symptoms of the disease (biochemical, histologic and/or behavioral). Administration of a therapeutic may reduce or eliminate mild cognitive impairment in patients that have not yet developed characteristic Alzheimer's pathology. An amount adequate to accomplish therapeutic or prophylactic treatment is defined as a therapeutically or prophylactically effective dose. In both prophylactic and therapeutic regimes, compositions or medicaments are usually administered in several dosages until a sufficient immune response has been achieved.
[0112] Anti-tau antibodies or fragments thereof of the disclosure may be administered in combination with other agents that are effective for treatment of related neurodegenerative diseases. In the case of AD, antibodies of the disclosure may be administered in combination with agents that reduce or prevent the deposition of amyloid beta (A(3). It is possible that PHF-tau and AP pathologies are synergistic. Therefore, combination therapy targeting the clearance of both PHF-tau and AP-elated pathologies at the same time may be more effective than targeting each individually.
[0113] In the case of Parkinson's Disease and related neurodegenerative diseases, immune modulation to clear aggregated forms of the a-synuclein protein is also an emerging therapy. A combination therapy that targets the clearance of both tau and a-synuclein proteins simultaneously may be more effective than targeting either protein individually. In the methods of the disclosure, the "therapeutically effective amount" of the antibody in the treatment or ameliorating symptoms of a tauopathy can be determined by standard research techniques. For example, the dosage of the antibody can be determined by administering the agent to relevant animal models well known in the art.
[0114] In addition, in vitro assays can be optionally employed to help identify optimal dosage ranges. Selection of a particular effective dose can be determined (e.g., via clinical trials) by those skilled in the art based upon the consideration of several factors. Such factors include the disease to be treated or prevented, the symptoms involved, the patient's body mass, the patient's immune status and other factors known by the skilled artisan. The precise dose to be employed in the formulation will also depend on the route of administration, and the severity of disease, and should be decided according to the judgment of the practitioner and each patient's circumstances. Effective doses can be extrapolated from dose-response curves derived from in vitro or animal model test systems. The mode of administration for therapeutic use of the antibodies of the disclosure may be any suitable route that delivers the agent to the host. Pharmaceutical compositions of these antibodies are useful for parenteral administration, e.g., intradermal, intramuscular, intraperitoneal, intravenous, subcutaneous, intranasal or intracranial, or they can be administered into the cerebrospinal fluid of the brain or spine.
[0115] The antibodies of the disclosure may be prepared as pharmaceutical compositions containing an effective amount of the antibody as an active ingredient in a pharmaceutically acceptable carrier. The term "carrier" refers to a diluent, adjuvant, excipient, or vehicle with which the antibody is administered. Such pharmaceutical vehicles can be liquids, such as water and oils, including those of petroleum, animal, vegetable or synthetic origin, such as peanut oil, soybean oil, mineral oil, sesame oil and the like. For example, 0.4% saline and 0.3% glycine can be used. These solutions are sterile and generally free of particulate matter. They may be sterilized by conventional, well-known sterilization techniques (e.g., filtration). The compositions may contain pharmaceutically acceptable auxiliary substances as required to approximate physiological conditions such as pH adjusting and buffering agents, stabilizing, thickening, lubricating and coloring agents, etc. The concentration of the antibodies of the disclosure in such pharmaceutical formulation can vary widely, i.e., from less than about 0.5%, usually at or at least about 1% to as much as 15% or 20% by weight and will be selected primarily based on required dose, fluid volumes, viscosities, etc., according to the particular mode of administration selected.
[0116] The treatment may be given in a single dose schedule, or as a multiple dose schedule in which a primary course of treatment may be with 1-10 separate doses, followed by other doses given at subsequent time intervals required to maintain and or reinforce the response, for example, at 1-4 months for a second dose, and if needed, a subsequent dose(s) after several months. Examples of suitable treatment schedules include: (i) 0, 1 month and 6 months, (ii) 0, 7 days and 1 month, (iii) 0 and 1 month, (iv) 0 and 6 months, or other schedules sufficient to elicit the desired responses expected to reduce disease symptoms, or reduce severity of disease. Thus, a pharmaceutical composition of the disclosure for intramuscular injection could be prepared to contain 1 ml sterile buffered water, and between about 1 ng to about 100 mg, about 50 ng to about 30 mg or about 5 mg to about 25 mg of an antibody of the disclosure. Similarly, a pharmaceutical composition of the disclosure for intravenous infusion could be made up to contain about 250 ml of sterile Ringer's solution, and about 1 mg to about 30 mg or about 5 mg to about 25 mg of an antibody of the disclosure. Actual methods for preparing parenterally administrable compositions are well known and are described in more detail in, for example, Remington's Pharmaceutical Science, 15th ed., Mack Publishing Company, Easton, Pa.
[0117] The antibodies of the disclosure can be lyophilized for storage and reconstituted in a suitable carrier prior to use. This technique has been shown to be effective with antibody and other protein preparations and known lyophilization and reconstitution techniques can be employed.
[0118] Diagnostic Methods and Kits
[0119] Antibodies of the disclosure may be used in methods of diagnosing AD or other tauopathy in a subject. This method involves detecting, in the subject, the presence of tau using a diagnostic reagent such as an antibody or a fragment thereof of this disclosure. Tau may be detected in a biological sample from a subject (e.g., blood, urine, cerebral spinal fluid) by contacting the biological sample with the diagnostic antibody reagent, and detecting binding of the diagnostic antibody reagent to PHF-tau in the sample from the subject. Assays for carrying out the detection include well-known methods such as ELISA, immunohistochemistry, Western blot, or in vivo imaging. Exemplary diagnostic antibodies are antibodies CBTAU-7.1, CBTAU-8.1, CBTAU-16.1, CBTAU-18.1, CBTAU-20.1, CBTAU-22.1, CBTAU-24.1, CBTAU-41.1, CBTAU-41.2, and CBTAU-42.1 of the disclosure, and are of IgG1, K type.
[0120] Diagnostic antibodies or similar reagents can be administered by intravenous injection into the body of the patient, or directly into the brain by any suitable route that delivers the agent to the host as exemplified above. The dosage of antibody should be within the same ranges as for treatment methods. Typically, the antibody is labeled, although in some methods, the primary antibody with affinity for tau is not labeled and a secondary labeling agent is used to bind to the primary antibody. The choice of label depends on the means of detection. For example, a fluorescent label is suitable for optical detection. Use of paramagnetic labels is suitable for tomographic detection without surgical intervention. Radioactive labels can also be detected using PET or SPECT.
[0121] Diagnosis is performed by comparing the number, size, and/or intensity of labeled tau, tau accumulation, tau aggregates, and/or neurofibrillary tangles in a sample from the subject or in the subject, to corresponding baseline values. The baseline values can represent the mean levels in a population of non-diseased individuals. Baseline values can also represent previous levels determined in the same subject.
[0122] The diagnostic methods described above can also be used to monitor a subject's response to therapy by detecting the presence of tau in a subject before, during or after the treatment. A change in values relative to baseline signals a response to treatment. Values can also change temporarily in biological fluids as pathological tau is being cleared from the brain.
[0123] The present disclosure is further directed to a kit for performing the above-described diagnostic and monitoring methods. Typically, such kits contain a diagnostic reagent such as the antibodies of the disclosure and, optionally, a detectable label. The diagnostic antibody itself may contain the detectable label (e.g., fluorescent molecule, biotin, etc.), which is directly detectable or detectable via a secondary reaction (e.g., reaction with streptavidin). Alternatively, a second reagent containing the detectable label may be utilized, where the second reagent has binding specificity for the primary antibody. In a diagnostic kit suitable for measuring tau in a biological sample, the antibodies of the kit may be supplied pre-bound to a solid phase, such as to the wells of a microtiter dish.
[0124] The contents of all cited references (including literature references, issued patents, published patent applications, and co-pending patent applications) cited throughout this application are hereby expressly incorporated by reference.
EXAMPLES
Example 1
[0125] Tau Peptide Design and Labeling
[0126] Hyperphosphorylation of tau protein resulting in release from microtubules and leading to depolymerization is a pathological hallmark occurring in Alzheimer's disease (AD) and other related tauopathies. As the equilibrium of free tau to microtubule-bound tau shifts in favor of the former, unassociated tau protein is thought to accumulate in a misfolded, aggregated state. During the disease process, tau is thought to adopt a variety of conformations, progressing from soluble dimeric and oligomeric forms to higher-order insoluble aggregates such as paired helical filaments (PHFs) and neurofibrillary tangles (NFTs). However, the exact forms of tau that contribute to pathology and, hence, optimal for therapeutic targeting, remain unknown. Consequently, attempts to trget disease-promoting tau are often limited by the choice of target. In an effort to prepare novel anti-tau binding molecules, antibody variable regions to tau were recovered from human memory B-cells using phosphorylated and non-phosphorylated-tau peptides as bait antigens using a single-cell-based approach.
[0127] Human memory B-cells to tau are likely rare in the human repertoire; therefore, it was decided to label the tau baits with the brightest fluorophores. All tau peptides were synthesized with an amino-terminal biotin group to aid labeling with two bright fluorophores, streptavidin-APC or streptavidin-PE (aka, tau peptide tetramers). Each tau peptide was labeled with both fluorophores to increase the signal-to-noise during the screening of human memory B-cells (detailed in Example 2) from donor samples. Labeled tau peptide tetramers were prepared by mixing biotinylated peptide at a 35:1 molar ratio of peptide to streptavidin label overnight at 4.degree. C. with gentle mixing. Free peptide was removed by separation over a BioSpin 30 column (Biorad). All tau peptide tetramers were stored at 4.degree. C. for up to 2 months.
Example 2
[0128] Recovery of Anti-Tau-Specific Memory B-cells by Facs Sorting With Labelled Peptide Tetramers
[0129] Monoclonal antibodies against tau were recovered from memory B-cells (CD22+CD19+CD27+IgG+) isolated from peripheral blood mononuclear cells (PBMCs) obtained from presumably asymptomatic (non-AD) human blood donors obtained from San Diego Blood Bank and TSRI Normal Blood Donor Services. In addition, AD patient blood samples were obtained through the CRO, Quintiles, from which three antibodies detailed here were recovered. PBMCs were isolated on FICOLL-PAQUE PLUS.RTM. (GE Healthcare) and cryopreserved at 50 million cells per ml in 90% FBS and 10% DMSO. An aliquot of plasma was heat inactivated at 56.degree. C. and stored at -20.degree. C. for downstream assessment of plasma reactivity.
[0130] For each sorting experiment, PBMCs from three to four donors were thawed and transferred to tubes containing pre-warmed RPMI complete (RPMI, 10% heat inactivated FBS and 1% penicillin/streptomycin), washed and incubated separately at 37.degree. C. for 16 hours. Pooled PBMCs were enriched for mature B-cells by positive selection using CD22+magnetic beads (Miltenyi Biotec). Cells were resuspended in Tris-buffered saline pH 7.4, containing 2 mM EDTA and 0.25% bovine serum albumin Fraction V (TBS Buffer). The cells were stained with the extracellular markers IgG-FITC, CD19-PerCPCy5.5 and CD27-PECy7 (all from BD Biosciences) to label B-cells. Ten million cells were removed and as a negative control, biotin streptavidin-labeled conjugates were used. The remaining cells were incubated with a pool of ten dual-labeled tau peptide tetramers (SA-APC and SA-PE) at 16.8 nM each. Cells were incubated for 60 minutes at 4.degree. C. with gentle mixing, washed twice and re-suspended at 20 million cells per ml in TBS buffer. Prior to sorting, DAPI (Thermo Fisher) was added as a live cell marker and cells were sorted on a Beckman Coulter MoFlo XDP. Negative control samples were used to determine nonspecific binding and signal-to-noise ratio. CD19+, IgG+, CD27hi, and antigen double-positive cells were collected and deposited into individual wells of a 96-well PCR plate and stored at -80.degree. C.
Example 3
[0131] Recovery of Heavy and Light Chain Genes from Tau-Specific Single B-Cells
[0132] As detailed in Example 2, memory B-cells with reactivity to tau peptide tetramers were identified, isolated and sorted into individual microtiter wells. Heavy and light chain cDNAs were then recovered by a two-step PCR approach from individual B-cells, and variable domain sequences were cloned and expressed in vitro as full-length recombinant IgG1 antibodies and are thus human chimeric antibodies.
[0133] First Strand cDNA Synthesis
[0134] First-strand complementary DNA (cDNA) was generated from single sorted cells according to manufacturer's protocol (Superscript III, Invitrogen Corp.) with the following modifications: to each well containing a single B-cell, 0.5 .mu.l of 10% NP-40, 1.0 .mu.l of oligo dT, 1.0 .mu.l of dNTP was added and samples were incubated at 65.degree. C. for 5 minutes. After incubation, samples were placed on ice for 1 minute. The following was then added to each well: 2.0 .mu.l of DTI, 4.0 .mu.l of MgCl2, 1.0 .mu.l of SUPERSCRIPT.RTM. RT, and 0.5 .mu.l of RNaseOut. Samples were incubated at 50.degree. C. for 50 minutes, followed by incubation at 85.degree. C. for 5 minutes.
[0135] Step I Amplification
[0136] For the initial PCR (Step I), 2.5 .mu.l of cDNA preparation was used as a template to amplify heavy and kappa or lambda light chains. Primer pools specific to the leader regions of antibody heavy (CB-5'LVH primers, Table 1), kappa light chain (CB-5'LVk primers, Table 2), and lambda light chain (CB-5' LVlam primers, Table 3) were used. A single reverse primer specific to the CH1 region, CK, and CL regions of the heavy, kappa light and lambda light chains, respectively, were used in the Step I PCR reaction.
TABLE-US-00001 TABLE 1 VH STEP I FORWARD PRIMERS SEQ ID Primer ID DNA SEQUENCE (5'-3') NO: CB-5'LVH1a ATGGACTGGACCTGGAGGTTCCTC 7 CB-5'LVH1b ATGGACTGGACCTGGAGGATCCTC 8 CB-5'LVH1c ATGGACTGGACCTGGAGGGTCTTC 9 CB-5'LVH1d ATGGACTGGACCTGGAGCATCC 10 CB-5'LVH2 GGACATACTTTGTTCCACGCTCCTGC 11 CB-5'LVH3a AGGTGTCCAGTGTCAGGTGCAGC 12 CB-5'LVH3b AGGTGTCCAGTGTGAGGTGCAGC 13 CB-5'LVH3c AGGTGTCCAGTGTCAGGTACAGC 14 CB-5'LVH4 GCAGCTCCCAGATGGGTCCTG 15 CB-5'LVH5 TCAACCGCCATCCTCGCCCTC 16 CB-5'LVH6 GTCTGTCTCCTTCCTCATCTTCCTGC 17 3'CgCH1 GGAAGGTGTGCACGCCGCTGGTC 18
TABLE-US-00002 TABLE 2 VK STEP I FORWARD PRIMERS SEQ ID Primer ID DNA SEQUENCE (5'-3') NO: CB-5'LVk1a ATGAGGGTCCCCGCTCAGCTC 19 CB-5'LVk1b ATGAGGGTCCCTGCTCAGCTC 20 CB-5'LVk1c ATGAGAGTCCTCGCTCAGCTC 21 CB-5'LVk2 TGGGGCTGCTAATGCTCTGG 22 CB-5'LVk3 CCTCCTGCTACTCTGGCTCCCAG 23 CB-5'LVk4 TCTCTGTTGCTCTGGATCTCTGGTGC 24 CB-5'LVk5 CTCCTCAGCTTCCTCCTCCTTTGG 25 CB-5'LVk6 AACTCATTGGGTTTCTGCTGCTCTGG 26 3'Ck-Rev543 GTTTCTCGTAGTCTGCTTTGCTCAGC 27 3'Ck-Rev494 GTGCTGTCCTTGCTGTCCTGCTC 28 3'Ck-Rev GCACTCTCCCCTGTTGAAGCTCTTTG 29
TABLE-US-00003 TABLE 3 VL STEP I FORWARD PRIMERS (5'-3') SEQ ID Primer ID DNA SEQUENCE (5'-3') NO: CB-5'L Vlam1 CTCCTCGCTCACTGCACAGG 30 CB-5'L Vlam2 CTCCTCTCTCACTGCACAGG 31 CB-5'L Vlam3 CTCCTCACTCGGGACACAGG 32 CB-5'L Vlam4 ATGGCCTGGACCCCTCTCTG 33 CB-5'L Vlam5 ATGGCATGGATCCCTCTCTTCCTC 34 3'Cl-Rev CACTAGTGTGGCCTTGTTGGCTTG 35
[0137] Step II Amplification
[0138] For Step II, 2.5 .mu.l of Step I PCR product was used as a template to amplify heavy, and kappa or lambda light chain variable regions. A pool of forward and reverse primers specifically designed to the framework 1 region of antibody heavy chain (pCB-IgG-VH and 3'SalIJH primers, Table 4), kappa light chain (pCB-IgG-VK and 3'Jk primers, Table 5), and lambda light chain (CB-VL and 3'Clam-Step II primers, Table 6) were used to prepare DNA from the variable regions. Furthermore, Step II primers were designed to introduce XbaI (VK and VL forward primers) and XhoI (3'SalIJH primers) restriction sites for downstream cloning. Following the Step II amplification reactions, heavy and light chain variable domain PCR products were run on a 1% agarose gel. Heavy and light chain variable region fragments were purified according to the manufacturer's protocol (Qiagen) and used in the Step III PCR reaction.
TABLE-US-00004 TABLE 4 VH Step II Forward and Reverse Primers SEQ ID Primer ID DNA SEQUENCE (5' - 3') NO: pCB-IgG-VH1a CCTGTCTGGAATTCAGCATGGCCCAGGTGCAGCTGGTGCAGTC 36 pCB-IgG-VH1b CCTGTCTGGAATTCAGCATGGCCCAGGTCCAGCTGGTGCAGTC 37 pCB-IgG-VH1c CCTGTCTGGAATTCAGCATGGCCCAGGTTCAGCTGGTGCAGTC 38 pCB-IgG-VH1d CCTGTCTGGAATTCAGCATGGCCCAGGTCCAGCTTGTGCAGTC 39 pCB-IgG-VH2a CCTGTCTGGAATTCAGCATGGCCCAGGTCACCTTGAGGGAGTCTGG 40 pCB-IgG-VH2b CCTGTCTGGAATTCAGCATGGCCCAGGTCACCTTGAAGGAGTCTGG 41 pCB-IgG-VH3a CCTGTCTGGAATTCAGCATGGCCCAGGTGCAGCTGGTGGAGTC 42 pCB-IgG-VH3b CCTGTCTGGAATTCAGCATGGCCGAGGTGCAGCTGTTGGAGTC 43 pCB-IgG-VH3c CCTGTCTGGAATTCAGCATGGCCGAGGTGCAGCTGGTGGAGTC 44 pCB-IgG-VH3d CCTGTCTGGAATTCAGCATGGCCCAGGTACAGCTGGTGGAGTCTG 45 pCB-IgG-VH4a CCTGTCTGGAATTCAGCATGGCCCAGSTGCAGCTGCAGGAG 46 pCB-IgG-VH4b CCTGTCTGGAATTCAGCATGGCCCAGGTGCAGCTACAGCAGTGG 47 pCB-IgG-VH5 CCTGTCTGGAATTCAGCATGGCCGAGGTGCAGCTGGTGCAGTC 48 pCB-IgG-VH6 CCTGTCTGGAATTCAGCATGGCCCAGGTACAGCTGCAGCAGTCAG 49 pCB-IgG-VH7 CCTGTCTGGAATTCAGCATGGCCCAGGTGCAGCTGGTGCAATCTG 50 3'SalIJH 1/2/4/5 TCGGGCCTCGAGACTCACCTGAGGAGACGGTGACCAG 51 3'SalIJH3 TCGGGCCTCGAGACTCACCTGAAGAGACGGTGACCATTG 52 3'SalIJH6 TCGGGCCTCGAGACTCACCTGAGGAGACGGTGACCGTG 53
TABLE-US-00005 TABLE 5 VK STEP II FORWARD and REVERSE PRIMERS SEQ ID Primer ID DNA SEQUENCE (5'-3') NO: pCB-IgG- CCGGTCTAGAGTTTTCCATGGCGGACATCCAGATGACCCAGTCTCC 54 VK1a pCB-IgG- CCGGTCTAGAGTTTTCCATGGCGGACATCCAGTTGACCCAGTCTCC 55 VK1b pCB-IGG- CCGGTCTAGAGTTTTCCATGGCGGCCATCCAGTTGACCCAGTCTCC 56 VK1c pCB-IGG- CCGGTCTAGAGTTTTCCATGGCGGATRTTGTGATGACTCAGTCTCCACTC 57 VK2a pCB-IgG- CCGGTCTAGAGTTTTCCATGGCGGAAATTGTGTTGACGCAGTCTCCAG 58 VK3a pCB-IgG- CCGGTCTAGAGTTTTCCATGGCGGAAATTGTGTTGACACAGTCTCCAG 59 VK3b pCB-IgG- CCGGTCTAGAGTTTTCCATGGCGGAAATAGTGATGACGCAGTCTCCAG 60 VK3c pCB-IgG-VK4 CCGGTCTAGAGTTTTCCATGGCGGACATCGTGATGACCCAGTCTCC 61 pCB-IgG-VK5 CCGGTCTAGAGTTTTCCATGGCGGAAACGACACTCACGCAGTCTCC 62 pCB-IgG-VK6 CCGGTCTAGAGTTTTCCATGGCGGAAATTGTGCTGACTCAGTCTCCAG 63 3'Jk1 Rev IL CGCAAAGTGCACTTACGTTTGATTTCCACCTTGGTCCCTTGGC 64 3'Jk2 Rev Iib CGCAAAGTGCACTTACGTTTGATCTCCAGCTTGGTCCCCTGGC 65 3'Jk4 Rev Ilc CGCAAAGTGCACTTACGTTTGATATCCACTTTGGTCCCAGGGC 66 3'Jk3 Rev Ilc CGCAAAGTGCACTTACGTTTGATCTCCACCTTGGTCCCTCCGC 67 3'Jk5 Rev Ild CGCAAAGTGCACTTACGTTTAATCTCCAGTCGTGTCCCTTGGC 68
TABLE-US-00006 TABLE 6 VL STEP II FORWARD and REVERSE PRIMERS SEQ ID Primer ID DNA SEQUENCE (5'-3') NO: CB-VL1 CCGGTCTAGAGTTTTCCATGGCGAATTTTATGCTGACTCAGCCCCACTC 69 CB-VL2 CCGGTCTAGAGTTTTCCATGGCGTCCTATGTGCTGACTCAGCC 70 CB-VL3 CCGGTCTAGAGTTTTCCATGGCGCAGTCTGTGCTGACGCAGCC 71 CB-VL4 CCGGTCTAGAGTTTTCCATGGCGCAGTCTGTCGTGACGCAGCC 72 CB-VL5 CCGGTCTAGAGTTTTCCATGGCGCAGTCTGCCCTGACTCAGCC 73 CB-VL6 CCGGTCTAGAGTTTTCCATGGCGTCTTCTGAGCTGACTCAGGACC 74 CB-VL7 CCGGTCTAGAGTTTTCCATGGCGTCCTATGAGCTGACTCAGCCACC 75 3'Clam-Step II CTCAGAGGAGGGYGGGAACAGAGTGAC 76
[0139] Step III Amplification: Overlap Extension PCR
[0140] For Step III, the heavy and light chain variable region DNA fragments produced in Step II were linked into a single cassette via overlap extension PCR using 1) a kappa linker or lambda linker (see linker preparation method below), which anneals to the 3' end of the light chain Step II fragment and the 5' end of the heavy chain Step II fragment, and contains either the kappa or lambda constant region, 2) a forward overlap primer containing an XbaI restriction site, and 3) a reverse primer containing an XhoI restriction site. This reaction results in an approximate 2400 bp or 2200 bp amplicon (i.e., cassette) for the kappa or lambda chains, respectively, consisting of the light chain variable region, linker, and heavy chain variable region. Following amplification, the overlap extension PCR reaction product was PCR-purified according to the manufacturer's instructions (Qiagen PCR Purification Kit).
[0141] Linker Preparation
[0142] The linker fragment was amplified using pCB-IgG, a dual-CMV promoter vector generated in house and used to express both heavy and light chain genes, as template and the primers listed in Table 7. The linker fragment is 1765 or 1536 base pairs in length for kappa or lambda linker, respectively. The kappa linker contains from a 5' to 3' intron sequence followed by the kappa constant region, poly(A) termination sequence, and cytomegalovirus promoter sequence, allowing for one vector expression of the recombinant antibodies. The lambda linker contains the lambda constant region, poly(A) termination sequence, and cytomegalovirus promoter sequence. A common reverse primer (Linker VH_HAVT20_pCB-IgG-R) and kappa-specific forward primer (Linker_CK_intronpCB-IgG-F) were used (Table 7). The amplified fragment was separated on a 1% agarose gel and purified according to the manufacturer's protocol (Qiagen Gel Extraction Kit).
TABLE-US-00007 TABLE 7 LINKER AND OVERLAP PRIMERS SEQ ID Primer ID SEQUENCE (5'-3') NO: Linker_VH_HAVT20_ GGCCATGCTGAATTCCAGACAGG 78 pCB-IgG-R Linker_CK_intron_ AAACGTAAGTGCACTTTGCGGCCGC 79 pCB-IgG-F TAGG Linker_CL_intron_ ACTCTGTTCCCRCCCTCCTCTGAGG 80 pCB-IgG-F_opt pCB-overlap F CCGGTCTAGAGTTTTCCATGGCG 81 pCB-overlap R TCGGGCCTCGAGACTCACC 82
[0143] Cloning into Mammalian Expression Vector
[0144] Following purification of the overlap extension PCR product, the fragment was digested with XhoI and XbaI and subsequently separated on a 1% agarose gel. The band corresponding to the overlap cassette (.sup..about.2.4 kb) was purified and ligated into an IgG1 expression vector, pCB-IgG. Antibody variable genes were subcloned into this vector and antibodies were recombinantly expressed as IgG1, regardless of their original (native) isotype. (An example of an IgG1 heavy chain constant region amino acid sequence is shown in SEQ ID NO:83 and light chain kappa constant region amino acid sequence is shown in SEQ ID NO:84.) All transformations were carried out using DH5a Max Efficiency cells (Invitrogen Corp.) and recovered in 250 IA of SOC for 1 hour at 37.degree. C. Approximately 100 .mu.l of recovered cells were plated onto a carbenicillin plate supplemented with 20 mM glucose. Plates were incubated overnight at 37.degree. C. to allow for colony growth. The remaining recovered cell mixture was cultured with 4 ml of Super Broth (SB) media supplemented with 50 .mu.g/ml carbenicillin and incubated overnight at 37.degree. C. with shaking at 250 rpm. The following day, five colonies were picked per plate and grown in 3 ml of SB media supplemented with 50 .mu.g/ml carbenicillin overnight at 37.degree. C. Overnight cultures were used for DNA plasmid preparation (Qiagen).
Example 4
[0145] Antibody Sequencing, Germline Identification and Confirmation of Anti-Tau Peptide Reactivity in Transfection Supernatant
[0146] To express IgG1 s, DNA plasmid minipreps of the aforementioned 4 ml cultures were prepared (Qiagen) and used to transfect 293Expi cells using EXPIFECTAMINE.RTM. according to the manufacturer's instructions (Invitrogen, Corp.). Transfections were carried out for a minimum of 72 hours in 10 ml cultures to allow for sufficient IgG expression. Cell media was harvested post-transfection and centrifuged to remove the cells and debris. Supernatants were quantified using Protein A sensor tips on an Octet Red system (ForteBio). Each supernatant was subsequently tested by ELISA with the bait peptide in order to confirm the presence of anti-tau reactive antibodies. Plasmid MINIPREP.TM. DNA (Qiagen) from the four individually picked cultures in Example 3 was prepared and heavy and light chains were sequenced with primers pC9_seq_HC-R (5'CATGTCACCGGGGTGTGG 3') (SEQ ID NO:85) and pC9_seq_LC-R (5'TCACAGGGGATGTTAGGGACA3') (SEQ ID NO:86). One clone of the four was selected for subsequent experiments.
[0147] Heavy and Light chain variable region protein and nucleic acid sequences of antibody clones CBTAU-7.1 (SEQ ID NOS:87, 88, 89, 90), CBTAU-8.1 (SEQ ID NOS:91, 92, 93, 94), CBTAU-16.1 (SEQ ID NOS:95, 96, 97, 98), CBTAU-18.1 (SEQ ID NOS: 99, 100, 101, 102), CBTAU-20.1 (SEQ ID NOS:103, 104, 105, 106), CBTAU-22.1 (SEQ ID NOS:107, 108, 109, 110), CBTAU-24.1 (SEQ ID NO:111, 112, 113, 114), CBTAU-27.1 (SEQ ID NOS:115, 116, 117, 118), CBTAU-28.1 (SEQ ID NOS:119, 120, 121, 122), CBTAU-41.1 (SEQ ID NOS:123, 124, 125, 126), CBTAU-41.2 (SEQ ID NOS:127, 128, 129, 130), CBTAU-42.1 (SEQ ID NOS:131, 132, 133, 134), CBTAU-43.1 (SEQ ID NOS:135, 136, 137, 138), CBTAU-44.1 (SEQ ID NOS:139, 140, 141, 142), CBTAU-45.1 (SEQ ID NOS:143, 144, 145, 146), CBTAU-46.1 (SEQ ID NOS:147, 148, 149, 150), CBTAU-47.1 (SEQ ID NOS:151, 152, 153, 154), CBTAU-47.2 (SEQ ID NOS:155, 156, 157, 158), and CBTAU-49.1 (SEQ ID NOS:159, 160, 161, 162) define novel CDRs for the selected anti-tau antibodies (Table 8).
[0148] An anti-tau antibody CBTAU-7.1 was generated comprising the VH of SEQ ID NO:87 and the VL of SEQ ID NO:88 and a human IgG1 constant region. An anti-tau antibody CBTAU-8.1 was generated comprising the VH of SEQ ID NO:91 and the VL of SEQ ID NO:92 and a human IgG1 constant region. An anti-tau antibody CBTAU-16.1 was generated comprising the VH of SEQ ID NO:95 and the VL of SEQ ID NO:96 and a human IgG1 constant region. An anti-tau antibody CBTAU-18.1 was generated comprising the VH of SEQ ID NO:99 and the VL of SEQ ID NO:100 and a human IgG1 constant region. An anti-tau antibody CBTAU-20.1 was generated comprising the VH of SEQ ID NO:103 and the VL of SEQ ID NO:104 and a human IgG1 constant region. An anti-tau antibody CBTAU-22.1 was generated comprising the VH of SEQ ID NO:107 and the VL of SEQ ID NO:108 and a human IgG constant region. An anti-tau antibody CBTAU-24.1 was generated comprising the VH of SEQ ID NO:111 and the VL of SEQ ID NO:112 and a human IgG1 constant region. An anti-tau antibody CBTAU-27.1 was generated comprising the VH of SEQ ID NO:115 and the VL of SEQ ID NO:116 and a human IgG1 constant region. An anti-tau antibody CBTAU-28.1 was generated comprising the VH of SEQ ID NO:119 and the VL of SEQ ID NO:120 and a human IgG1 constant region. An anti-tau antibody CBTAU-41.1 was generated comprising the VH of SEQ ID NO:123 and the VL of SEQ ID NO:124 and a human IgG1 constant region. An anti-tau antibody CBTAU-41.2 was generated comprising the VH of SEQ ID NO:127 and the VL of SEQ ID NO:128 and a human IgG1 constant region. An anti-tau antibody CBTAU-42.1 was generated comprising the VH of SEQ ID NO:131 and the VL of SEQ ID NO:132 and a human IgG1 constant region. An anti-tau antibody CBTAU-43.1 was generated comprising the VH of SEQ ID NO:135 and the VL of SEQ ID NO:136 and a human IgG1 constant region. An anti-tau antibody CBTAU-44.1 was generated comprising the VH of SEQ ID NO:139 and the VL of SEQ ID NO:140 and a human IgG1 constant region. An anti-tau antibody CBTAU-45.1 was generated comprising the VH of SEQ ID NO:143 and the VL of SEQ ID NO:144 and a human IgG1 constant region. An anti-tau antibody CBTAU-46.1 was generated comprising the VH of SEQ ID NO:147 and the VL of SEQ ID NO:148 and a human IgG1 constant region. An anti-tau antibody CBTAU-47.1 was generated comprising the VH of SEQ ID NO:151 and the VL of SEQ ID NO:152 and a human IgG1 constant region. An anti-tau antibody CBTAU-47.2 was generated comprising the VH of SEQ ID NO:155 and the VL of SEQ ID NO:156 and a human IgG1 constant region. An anti-tau antibody CBTAU-49.1 was generated comprising the VH of SEQ ID NO:159 and the VL of SEQ ID NO:160 and a human IgG1 constant region.
TABLE-US-00008 TABLE 8 Amino acid sequences of heavy and light chain variable region CDRs Tau aa CDR1 CDR2 CDR3 Clone region (SEQ ID NO:) (SEQ ID NO:) (SEQ ID NO:) CBTA 194-212 SYWMH RINSDGSDTNYAD GRSYGFFDY U 7.1 (163) SVKG (165) (164) RASQIISSNYLA GASSRAT QQYGTSPRT (166) (167) (168) CBTA 194-212 TYGMH VIWFDGNNKYYA DWWEAGCRPCYFF U 8.1 (169) DSVKG DY (170) (171) KSSQSVLYSSNNK WASTRES QQYYSPPLT NYLA (172) (173) (174) CBTA 204-221 DYWMS NINQDGSAAYYVD DAHYYDRNRNNYY U 16.1 (175) SVRG YYFDF (176) (177) RASQSVGANLA SASTRAT QQYNNWPRT (178) (179) (180) CBTA 200-217 SGNYYWS RMSSSGSTNYNPSL ESGSSWQNHYYYYG U 18.1 (181) KS MDV (182) (183 KSSQSVLYSSNNK WASTRES QQYYSTPLT NYLA (172) (173) (184) CBTA 58-78 NYAMS GISSDGNTFYADSV ESGRWGGGTLYGA U 20.1 (185) KG HY (186) (187) KSSQSLLYNSNNK WASTRES QQYYSSPLT NYLT (188) (173) (189) CBTA 406-429 DYNVH RISPNSGGTKYAQ GHCDGTTCSRAY U 22.1 (190) KFQG (192) (191) RSSQSLLHRSGHK LGSNRAS MQTLQTPWT YLH (194) (195) (193) CBTA 221-245 GYYLH WVNPRSGGTSYPP GRIPDVTAFDI U 24.1 (196) KFQG (198) (197) KSSESLLYDSNNK WASTRES QQYFSTPWT NYLA(199) (173) (200) CBTA 299-328 DYWTA IIYSGDSDTRYHPS LDARVDAGWQLDS U 27.1 (201) VQG(202) (203) KSSQSVFSRDNNK WASSRES QHYFNTPHN NYLA (204) (205) (206) CBTA 52-71 NYWIG IIYPGDSDTRYSPPF VGRPSKGGWFDP U 28.1 (207) QG (209) (208) ESSQTLLYSSNEK WASTPES QQYYNSPYT NYLA (211) (212) (210) CBTA 406-429 DSYMS YISRSSSHTNYADS VQTTMIEGKTKLNY U 41.1 (213) VKG FDY (214) (215) ESSHSLLYRSNNR WASTRES QQFYTTPYT NYLA (173) (217) (216) CBTA 406-429 DSYMS YISRSSSHTNYADS VQTTMIEGKTKLNY U 41.2 (213) VKG FDY (214) (215) ESSHSLLYRSNNK WASTRES QQFYTTPYT NYLA (173) (217) (218) CBTA 406-429 KAWMS RIKSKVDGETTDY LIHCDLSACLPHF U 42.1 (219) AAPVRG (220) (221) ESSHSLLYRSNNK WASTRES QQFYTTPYT NYLA (173) (217) (218) CBTA 299-328 NYWIA IIYPGDSDTTYSPSF LPRTDGDNSIGYFEY U 43.1 (222) QG (224) (223) KSSQSVLYSSNSE WASTRES QQYYSTPFT NYLA (173) (226) (225) CBTA 406-429 SYSMN YISSSTTTIYYADSV VPAPRLGGSYTY U 44.1 (227) KG (229) (228) RASQSVSSSYLA GASSRAT QQYGTSPLT (230) (167) (231) CBTA 406-429 DAWMS RIKSKNVGEIIDY GLGGGTYG U 45.1 (232) AEHVRG (233) (234) RSSAGLRNNDGDI RVSRRDS MRGPY LLS (236) (237) (235) CBTA 82-103 IYEMN YITNRGSTIYYADS PRIGARVFDV U 46.1 (238) VKG (240) (239) KSSQTLLYKSNNE WASTRES QQYFTTALT NYLA (173) (242) (241) CBTA 52-71 DHWIG IIFPEDSDTRYSGSF VSVVRKGGWFDP U 47.1 (243) EG (245) (244) KSSQSLLYTSNNK WASTRES QQYYNSPYT NYLA (173) (212) (246) CBTA 52-71 DHWIG IIFPGDSDIRYSPSF VAVVRKGGWFDS U 47.2 (243) EG (248) (247) KSTQSLLWSANNK WASTRES QQYYNSPYT NYLA (173) (212) (249) CBTA 52-71 SYWIG IIYPDDSDTRYNAS RDRNCSGTTCYPRW U 49.1 (250) LEG FDS (251) (252) KSSQSLFYSGNSK WASTRDS HQYHSTPLS DFLA (253) (254) (255)
[0149] CBTAU-7.1 antibody comprises a heavy chain CDR region of SEQ ID NO:163, a heavy chain CDR2 region of SEQ ID NO:164, and a heavy chain CDR3 region of SEQ ID NO:165, alight chain CDR1 region of SEQ ID NO:166, alight chain CDR2 region of SEQ ID NO:167 and alight chain CDR3 region of SEQ ID NO:168. CBTAU-8.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:169, a heavy chain CDR2 region of SEQ ID NO:170, and a heavy chain CDR3 region of SEQ ID NO:171, alight chain CDR1 region of SEQ ID NO:172, alight chain CDR2 region of SEQ ID NO:173 and alight chain CDR3 region of SEQ ID NO:174. CBTAU-16.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:175, a heavy chain CDR2 region of SEQ ID NO:176, and a heavy chain CDR3 region of SEQ ID NO:177, alight chain CDR1 region of SEQ ID NO:178, alight chain CDR2 region of SEQ ID NO:179 and alight chain CDR3 region of SEQ ID NO:180. CBTAU-18.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:181, a heavy chain CDR2 region of SEQ ID NO:182, and a heavy chain CDR3 region of SEQ ID NO:183, a light chain CDR1 region of SEQ ID NO:172, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:184. CBTAU-20.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:185, a heavy chain CDR2 region of SEQ ID NO:186, and a heavy chain CDR3 region of SEQ ID NO:187, a light chain CDR1 region of SEQ ID NO:188, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:189. CBTAU-22.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:190, a heavy chain CDR2 region of SEQ ID NO:191, and a heavy chain CDR3 region of SEQ ID NO:192, a light chain CDR1 region of SEQ ID NO:193, a light chain CDR2 region of SEQ ID NO:194 and a light chain CDR3 region of SEQ ID NO:195. CBTAU-24.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:196, a heavy chain CDR2 region of SEQ ID NO:197, and a heavy chain CDR3 region of SEQ ID NO:198, a light chain CDR1 region of SEQ ID NO:199, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:200. CBTAU-27.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:201, a heavy chain CDR2 region of SEQ ID NO:202, and a heavy chain CDR3 region of SEQ ID NO:203, a light chain CDR1 region of SEQ ID NO:204, a light chain CDR2 region of SEQ ID NO:205 and a light chain CDR3 region of SEQ ID NO:206. CBTAU-28.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:207, a heavy chain CDR2 region of SEQ ID NO:208, and a heavy chain CDR3 region of SEQ ID NO:209, a light chain CDR1 region of SEQ ID NO:210, a light chain CDR2 region of SEQ ID NO:211 and a light chain CDR3 region of SEQ ID NO:212. CBTAU-41.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, a light chain CDR1 region of SEQ ID NO:216, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:217. CBTAU-41.2 antibody comprises a heavy chain CDR1 region of SEQ ID NO:213, a heavy chain CDR2 region of SEQ ID NO:214, and a heavy chain CDR3 region of SEQ ID NO:215, a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:174 and a light chain CDR3 region of SEQ ID NO:217. CBTAU-42.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:219, a heavy chain CDR2 region of SEQ ID NO:220, and a heavy chain CDR3 region of SEQ ID NO:221, a light chain CDR1 region of SEQ ID NO:218, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:217. CBTAU-43.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:222, a heavy chain CDR2 region of SEQ ID NO:223, and a heavy chain CDR3 region of SEQ ID NO:224, a light chain CDR1 region of SEQ ID NO:225, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:226. CBTAU-44.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:227, a heavy chain CDR2 region of SEQ ID NO:228, and a heavy chain CDR3 region of SEQ ID NO:229, a light chain CDR1 region of SEQ ID NO:230, a light chain CDR2 region of SEQ ID NO:167 and a light chain CDR3 region of SEQ ID NO:231. CBTAU-45.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:232, a heavy chain CDR2 region of SEQ ID NO:233, and a heavy chain CDR3 region of SEQ ID NO:234, a light chain CDR1 region of SEQ ID NO:235, a light chain CDR2 region of SEQ ID NO:236 and a light chain CDR3 region of SEQ ID NO:237. CBTAU-46.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:238, a heavy chain CDR2 region of SEQ ID NO:239, and a heavy chain CDR3 region of SEQ ID NO:240, a light chain CDR1 region of SEQ ID NO:241, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:242. CBTAU-47.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:243, a heavy chain CDR2 region of SEQ ID NO:244, and a heavy chain CDR3 region of SEQ ID NO:245, a light chain CDR1 region of SEQ ID NO:246, a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:212. CBTAU-47.2 antibody comprises a heavy chain CDR1 region of SEQ ID NO:243, a heavy chain CDR2 region of SEQ ID NO:247, and a heavy chain CDR3 region of SEQ ID NO:248, a light chain CDR1 region of SEQ ID NO:249 a light chain CDR2 region of SEQ ID NO:173 and a light chain CDR3 region of SEQ ID NO:212. CBTAU-49.1 antibody comprises a heavy chain CDR1 region of SEQ ID NO:250, a heavy chain CDR2 region of SEQ ID NO:251, and a heavy chain CDR3 region of SEQ ID NO:252, a light chain CDR1 region of SEQ ID NO:254, a light chain CDR2 region of SEQ ID NO:254 and a light chain CDR3 region of SEQ ID NO:255.
[0150] Nucleic acid sequences of heavy and light chain variable regions of the anti-tau monoclonal antibodies were compared to known germline sequences using IgBLAST, an immunoglobulin variable domain sequence analysis tool, available at the NCBI (Nucleic Acids Res. 2013 July; 41 (Web Server issue):W34-40). Sequence alignment of heavy and light chain framework H1 and L1 regions, aligned with their respective proposed germline sequence and PCR primer, are shown in Table 9. Confirmed sequences were scaled up for expression and purification (detailed in Example 5). Selected clones were expanded into a 50 ml culture and plasmid midiprep DNA was prepared (Machery Nagel Midi Prep kit). Plasmid DNA was then used to transfect a 30 ml culture of 293Expi cells as detailed in Example 5.
TABLE-US-00009 TABLE 9 Framework nucleic acids of H1 and L1 aligned with germline and primer (Amino acids above) Differences marked in lower case lettering. Native refers to the antibody SEQ mAb ID Amino Terminal Protein and N-terminal Nucleic Acid Sequences NO CBTAU-7.1 VH 87 (Q V Q L V E S) pCB-IgG-VH1b 37 CAGGTCCAGCTGGTGCAGTC CBTAU-7.1VH 89 CAGGTCCAGCTGGTGGAGTCCGGGGGAGGCTTAGTTCAGCC TGGGGGGTCCCTGAGACTCTCCT IGHV3-74*01. 256 gAGGTgCAGCTGGTGGAGTCCGGGGGAGGCTTAGTTCAGCCTG GGGGGTCCCTGAGACTCTCCT Native 7.1 VH 257 (e V Q L V E) VL 88 (D I V M T Q S P) pCB-IgG-Vk4 61 GACATCGTGATGACCCAGTCTCC CBTAU-7.1 VL 90 GACATCGTGATGACCCAGTCTCCAGACACCCTGTCTTTGTCTC CAGGGGAGAGAGCCACCCTCT IGKV3-NL5*01 258 GAaATtGTGtTGACgCAGTCTCCAGcCACCCTGTCTTTGTCTCC AGGGGAaAGAGCCACCCTCT Native 7.1 VL 259 (e I V l T Q S P) CBTAU-8.1 VH 91 (Q V A L V E S) pCB-IgG- 42 CAGGTGCAGCTGGTGGAGTC VH3a CBTAU-8.1 93 CAGGTGCAGCTGGTGGAGTCGAGGGGAGGCGTGGTCCAGCCTGGG VH ACGTCCCTGAGACTCTCCT IGHV3- 260 CAGGTGCAGCTGGTGGAGTCtgGGGGAGGCGTGGTCCAGCCTGGG 33*01 ACGTCCCTGAGACTCTCCT Native 8.1 261 (Q V A L V E S) VH VL 92 (E T T L T Q S P) pCB-IgG- 62 GAAACGACACTCACGCAGTCTCC Vk5 CBTAU-8.1 94 GAAACGACACTCACGCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGG VL CGAGAGGGCCACCATCA IGKV4- 262 GAcAtcgtgaTgACcCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGGG 1*01 CCACCATCA Native 8.1 263 (d i v m T Q S P) VL CBTAU-16.1 VH 95 (E V Q L V Q) pCB-IgG- 48 GAGGTGCAGCTGGTGCAGTC VH5 CBTAU- 97 GAGGTGCAGCTGGTGCAGTCTGGGGGAGGCTTGGTCCAGCCTGGG 16.1 VH GGGTCCCTGAGACTCTCCT IGHV3- 264 GAGGTGCAGCTGGTGgAGTCTGGGGGAGGCTTGGTCCAGCCTGGG 64*01 GGGTCCCTGAGACTCTCCT Native 16.1 265 (E V Q L V e S) VH VL 96 (E I V M T Q S P) pCB-IgG- 60 GAAATAGTGATGACGCAGTCTCCGG VK3c CBTAU- 98 GAAATAGTGATGACGCAGTCTCCGGCCACCCTGTCTGTGTCTCCAGG 16.1VL GGAAAGAGCCACCCTCT IGKV3- 266 GAAATAGTGATGACGCAGTCTCCaGCCACCCTGTCTGTGTCTCCAGG 15*01 GGAAAGAGCCACCCTCT Native 16.1 267 (E I V M T Q S P) VL CBTAU-18.1 VH 99 (Q V Q L L E S) No exact primer match CBTAU- 101 CAGGTGCAGCTMIGGAGTCGGGCCCAGGACTGGTGAACCCTTCACAG 18.1 VH ACCCTGTCCCTCACCT IGHV4- 268 CAGGTGCAGCTGcaGGAGTCGGGCCCAGGACTGGTGAAGCCTTCACAGA 31*05 CCCTGTCCCTCACCT Native 18.1 269 (Q V Q L q E S) VH VL 100 (E I V L T Q S P) pCB-IgG- 59 GAAATTGTGTTGACACAGTCTCCAG VK3b CBTAU- 102 GAAATTGTGTTGACACAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCG 18.1 VL AGAGGGCCAACATTA IGKV4- 262 GAcATcGTGaTGACcCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAG 1*01 AGGGCCAcCATcA Native 18.1 270 (d I V m T Q S P) VL CBTAU-20.1 VH 103 (Q V Q L V E S) pCB-IgG- 45 CAGGTACAGCTGGTGGAGTCTG VH3d CBTAU- 105 CAGGTACAGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGG 20.1 VH GGGTCCCTGAGACTCTCCT IGHV3- 271 gAGGTgCAGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGGG 23*04 GGTCCCTGAGACTCTCCT Native 20.1 272 (e V Q L V E S) VH VL 104 (D I Q M T Q S P) pCB-IgG- 54 GACATCCAGATGACCCAGTCTCC VK1a CBTAU- 106 GACATCCAGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCG 20.1 VL AGAGGGCCACCATCA IGKV4- 262 GACATCgtGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAG 1*01 AGGGCCACCATCA Native 20.1 273 (D I v M T Q S P) VL CBTAU-22.1 VH 107 (Q V Q L V Q S) pCB-IgG- 36 CAGGTGCAGCTGGTGCAGTC VH1a CBTAU- 109 CAGGTGCAGCTGGTGCAGTCTGGGGCTGAGGTGAAGAAGCCTGGGG 22.1 VH CCCCAGTGAAGGTCTC IGHV1- 274 CAGGTGCAGCTGGTGCAGTCTGGGGCTGAGGTGAAGAAGCCTGGGG 2*02 CCtCAGTGAAGGTCTC Native 22.1 275 (Q V Q L V Q S) VH VL 108 (D V V M T Q S P L) No Exact Primer match CBTAU- 110 GATGTTGTGATGACGCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAG 22.1 VL AGCCGGCCTCCATC IGKV2- 276 GATaTTGTGATGACtCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAG 28*01 CCGGCCTCCATC Native 22.1 277 (D i V M T Q S P L) VL CBTAU-24.1 VH 111 (Q V Q L V S G) pCB-IgG- 39 CAGGTCCAGCTTGTGCAGTC VH1d CBTAU- 113 CAGGTCCAGCTTGTGCAGTCTGGGGCTGAGGTGAAGAAGCCTGGGG 24.1 VH CCTCAGTGAAGGTCTCC IGHV1- 278 CAGGTCCAGCTTGTGCAGTCTGGGGCTGAGGTGAAGAAGCCTGGGG 3*01 CCTCAGTGAAGGTtTCC Native 24.1 279 (Q V Q L V S G) VH VL 112 (D I Q M T Q S P) pCB-IgG- 54 GACATCCAGATGACCCAGTCTCC VK1a CBTAU- 114 GACATCCAGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGG 24.1 VL CGAGAGGGCCACCATC IGKV4- 262 GACATCgtGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCG 1*01 AGAGGGCCACCATC Native 24.1 280 (D I v M T Q S P) VL CBTAU27.1 VH 115 (Q V Q L V E S) pCB-IgG- 42 CAGGTGCAGCTGGTGGAGTC VH3a CBTAU27.1 117 CAGGTTCAGCTGGTGGAGTCTGGACCGGAGATGAGAAAGCCCGGGGAG VH TCTCTGAAAATTTCC IGHV5- 281 gAGGTgCAGCTGGTGcAGTCTGGAgCaGAGgTGAaAAAGCCCGGGGAGTCTC 51*01 TGAAgATcTCC Native 27.1 282 (e V Q L V q S) VH VL 116 (D I Q L T Q S P) pCB-IgG- 55 GACATCCAGTTGACCCAGTCTCC VK1b CBTAU27.1 118 GACATCCAGTTGACCCAGTCTCCAGATTCCCTGGCTGTGTCTCTGGG VL CGAGCGGGCCACCATC IGKV4- 262 GACATCgtGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGG 1*01 CGAGAGGGCCACCATC Native 27.1 283 (D I v m T Q S)P VL CBTAU28.1 VH 119 (Q V Q L Q Q S) pCB-IgG- 49 CAGGTaCAGCTgCAGCAGTCAG VH6 CBTAU28.1 121 CAGGTGCAGCTACAGCAGTCAGGAGCAGAAGTGAAAAAGCCCGGGG VH AGTCTCTGAAGATCTCC IGHV5- 281 gAGGTGCAGCTggtGCAGTCtGGAGCAGAAGTGAAAAAGCCCGGGGAG 51*01 TCTCTGAAGATCTCC Native 28.1 284 (e V Q L v Q S) VH VL 120 (D I Q M T Q S P) pCB-IgG- 54 GACATCCAGATGACCCAGTCTCC VK1a CBTAU28.1 122 GACATCCAGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCG VL AGAGGGCCACCATC IGKV4- 262 GACATCgtGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAG 1*01 AGGGCCACCATC Native 28.1 285 (D I v M T Q S P) VL CBTAU41.1 VH 123 (E V Q L L E S) pCB-IgG- 43 GAGGTGCAGCTGTTGGAGTC VH3b CBTAU41.1 125 GAGGTGCAGCTGTTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGG VH GTCCCTGAGACTCTCC IGHV3- 286 cAGGTGCAGCTGgTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGG 11*06 GTCCCTGAGACTCTCC Native 41.1 287 (q V Q L v E S) VH VL 124 (D I Q M T Q S P) pCB-IgG- 54 GACATCCAGATGACCCAGTCTCC VK1a CBTAU41.1 126 GACATCCAGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGG VL CGAGAGGGTCACCATC IGKV4- 262 GACATCgtGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCG 101 AGAGGGT cAccATc Native 41.1 288 (D I v M T Q S P) VL CBTAU41.2 VH 127 (E V Q L V Q S) pCB-IgG- 43 GAGGTGCAGCTGGTGCAGTC VH3b CBTAU41.2 129 GAGGTGCAGCTGGTGCAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGG VH TCCCTGAGACTCTCC IGHV3- 286 cAGGTGCAGCTGGTGgAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGG 11*06 TCCCTGAGACTCTCC Native 41.2 289 (q V Q L v E S) VH VL 128 (A I Q L T Q S P) pCB-IgG- 56 GCCATCCAGTTGACCCAGTCTCC VK1c CBTAU41.2 130 GCCATCCAGTTGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAG VL AGGGTCACCATC IGKV4- 262 gaCATCgtGaTGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAG 1*01 GGTCACCATC Native 41.2 290 (d I v m T Q S P) VL CBTAU42.1 VH 131 (Q L V Q S E G) pCB-IgG- 36 CAGCTGGTGCAGTC VH1a-c CBTAU42.1 133 CAGCTGGTGCAGTCTGAGGGAGGCCTGGCAGAGCCTGGGGGGTCCC VH TTAGACTC IGHV3- 291 CAGGTGGTGgAGTCTGgGGGAGGCCTGGCAGAGCCTGGGGGGTCCC 15*01 TTAGACTC Native 42.1 292 (Q L V e S g g) VH VL 132 (E I V L T Q S P) pCB-IgG- 58 GAAATTGTGTTGACGCAGTCTCCAG VK3a CBTAU41.1 134 GAAATTGTGTTGACGCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGA VL GAGGGTCACCATC IGKV4- 262 GAcATcGTGaTGACcCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGA 1*01 GGGTCACCATC Native 42.1 293 (d I V m T Q S P) VL CBTAU43.1 VH 135 (Q V Q L V Q S) pCB-IgG- 36 CAGGTGCAGCTGGTGCAGTC VH1a CBTAU43.1 137 CAGGTGCAGCTGGTGCAGTCTGGAGGAGAGGTGAAAAAGCCGGGGGAG VH TCTCTGAAGATCTCC IGHV5- 294 gAGGTGCAGCTGGTGCAGTCTGGAGGAGAGGTGAAAAAGCCGGGGGAG
51*03 TCTCTGAAGATCTCC Native 43.1 295 (e V Q L V Q S) VH VL 136 (E I V L T Q S P) pCB-IgG- 59 GAAATTGTGTTGACACAGTCTCCAG VK3b CBTAU43.1 138 GAAATTGTGTTGACACAGTCTCCAGCCTCCCTGGCTGTGTCTCTGGGCGAG VL AGGGCCACCATC IGKV4- 262 GAcATcGTGaTGACcCAGTCTCCAGCCTCCCTGGCTGTGTCTCTGGGCGAGAG 1*01 GGCCACCATC Native 43.1 296 (d I V m T Q S P) VL CBTAU44.1 VH 139 (E V Q L V E S) pCB-IgG- 44 GAGGTGCAGCTGGTGGAGTC VH3c CBTAU44.1 141 GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGGGGGTC VH CCTGAGACTCTCC IGHV3- 297 GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGGGGGTC 48*01 CCTGAGACTCTCC Native 44.1 298 (E V Q L V E S) VH VL 140 (D I Q M T Q S) pCB-IgG- 54 VK1a GACATCCAGATGACCCAGTCTCC CBTAU44.1 142 GACATCCAGATGACCCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAG VL AG CCACCCTC IGKV3- 299 GAaATtgtGtTGACgCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGAG 20*01 CCACCCTC Native 44.1 300 (e I v L T Q S) VL CBTAU45.1 VH 143 (E V Q L V E S) pCB-IgG- 44 GAGGTGCAGCTGGTGGAGTC VH3c CBTAU45.1 145 GAAATTGTGTTGACACAGTCTCCACTCTCCCTGCCCGCCACCCTTGGACAGC VH CGGCCTCCATC IGHV3- 301 GAGGTGCAGCTGGTGGAGTCTGGGGGAGACTTGGTAAAGCCTGGGGGGTC 15*02 CCTTAGACTCTCC Native 45.1 302 (E V Q L V E S) VH VL 144 (E I V L T Q S P) pCB-IgG- 59 GAAATTGTGTTGACACAGTCTCCAG VK3b CBTAU45.1 146 GAAATTGTGTTGACACAGTCTCCACTCTCCCTGCCCGCCACCCTTGGACAG VL CCGGCCTCCATC IGKV2- 303 GAtgTTGTGaTGACtCAGTCTCCACTCTCCCTGCCCGCCACCCTTGGACAGCC 30*01 GGCCTCCATC Native 45.1 304 (d v V m T Q S P) VL CBTAU46.1 VH 147 (Q V Q L V E S) pCB-IgG- 45 CAGGTACAGCTGGTGGAGTCTG VH3d CBTAU46.1 149 CAGGTACAGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGAGAGT VH CCCTGAGACTCTCC IGHV3- 305 gAGGTgCAGCTGGTGGAGTCTGGGGGAGGGTTGGTACAGCCTGGAGAGTC 48*03 CCTGAGACTCTCC Native 46.1 306 (E V Q L V E S) VH VL 148 (D I Q L T Q S P) pCB-IgG- 55 GACATCCAGTTGACCCAGTCTCC VK1b CBTAU46.1 150 GACATCCAGTTGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGA VL GGGCCACCATC IGKV4- 262 GACATCgtGaTGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGG 1*01 GCCACCATC Native 46.1 307 (D I v m T Q S P) VL CBTAU47.1 VH 151 (Q V Q L V Q S) pCB-IgG- 36 CAGGTGCAGCTGGTGCAGTC VH1a CBTAU47.1 153 CAGGTGCAGCTGGTGCAGTCTGGAGCAGTGGTGAAAAAGCCCGGGGAG VH TCTCTGAAGATCTC IGHV5- 308 gAGGTGCAGCTGGTGCAGTCTGGAGCAGTGGTGAAAAAGCCCGGGGAGTC 51*01 TCTGAAGATCTC Native 47.1 309 (E V Q L V Q S) VH VL 152 (A I Q L T Q S P) pCB-IgG- 56 GCCATCCAGTTGACCCAGTCTCC VK1c CBTAU44.1 154 GCCATCCAGTTGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGA VL GGGCCACCATC IGKV4- 262 GaCATCgtGaTGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAG 1*01 GGCCACCATC Native 47.1 310 (d I v m T Q S P) VL CBTAU47.2 VH 155 (Q V Q L V E S) pCB-IgG- 45 CAGGTACAGCTGGTGGAGTCTG VH3d CBTAU47.1 157 CAGGTACAGCTGGTGGAGTCTGGAGCAGAACTGAAAAAGCCCGGGGAGTCT VH CTGAAGATCTCC IGHV5- 281 gAGGTgCAGCTGGTGcAGTCTGGAGCAGAACTGAAAAAGCCCGGGGAGTCTCT 51*01 GAAGATCTCC Native 47.2 311 (e V Q L V q S) VH VL 156 (E I V M T Q S P) pCB-IgG- 60 GAAATAGTGATGACGCAGTCTCCAG VK3c CBTAU44.1 158 GAAATTGTGATGACCCAGTCTCCAGAGTCCCTGGCTGTGTCTCTGGGCGAGA VL GGGCCACCATC IGKV4- 262 GAcATcGTGATGACCCAGTCTCCAGAGTCCCTGGCTGTGTCTCTGGGCGAGA 1*01 GGGCCACCATC Native 47.2 312 (d I V M T Q S P) VL CBTAU49.1 VH 159 (Q V Q L V Q S) pCB-IgG- 36 CAGGTGCAGCTGGTGCAGTC VH1a CBTAU49.1 161 CAGGTGCAGCTGGTGCAGTCTGGGGCAGAGGTGAAAAAGCCGTGGGAGTC VH TCTGAAGATCTCC IGHV5- 294 gAGGTGCAGCTGGTGCAGTCTGGGGCAGAGGTGAAAAAGCCGTGGGAGTCT 51*03 CTGAAGATCTCC Native 49.1 313 (e V Q L V Q S) VH VL 160 (E I V L T Q S P) pCB-IgG- 63 GAAATTGTGCTGACTCAGTCTCCAG VK6 CBTAU49.1 162 GAAATTGTGCTGACTCAGTCTCCAGACTTCCTGGCTGTGTCTCTGGGCGAG VL AGGGCCACCATC IGKV4- 262 GAcATcGTGaTGACcCAGTCTCCAGACTTCCTGGCTGTGTCTCTGGGCGAGAGG 1*01 GCCACCATC Native 49.1 314 (d I V m T Q S P) VL
[0151] Transfected IgG1 supernatants were assayed for reactivity to tau peptides by ELISA. First, 96 half-well ELISA plates (Costar) were coated with 50 IA of bovine actin (1 .mu.g/ml, Sigma) as a negative control and Affinipure goat anti-human F(ab)2 (2 .mu.g/ml, Jackson Immunoresearch) to confirm antibody production. Plates were coated in TBS overnight at 4.degree. C. The following day, plates were washed five times with TB S/0.05% TWEEN.RTM. (TBS-T) and blocked with 150 IA of TBS-T plus 2.5% BSA (blocking buffer) for 2 hour. Tau peptides were captured on streptavidin-coated plates (Pierce) at a concentration of 0.43 M in 100 .mu.l of TBS. Tau peptides used to set up ELISA assays were the same used as baits in corresponding sorting experiment. Tau peptide-coated plates were then incubated at RT for 1.5 hours. All plates were then washed five times with TBS/0.05% TWEEN.RTM. and blocked with 150 .mu.l and 300 .mu.l (tau peptide plates only) of blocking buffer and incubated at RT for 2 hours. IgG transfection supernatants were diluted to 5 .mu.g/ml (based on quantitation by Octet Red) and titrated five-fold in TBS/0.25% BSA. Mouse anti-actin (Sigma, Cat. No. A3853) was used at 1.25 .mu.g/ml as a positive control for bovine actin-coated plates. Commercial grade antibodies were used at 1 .mu.g/ml as positive controls for ELISA assays, including AT8 monoclonal antibody (Thermo, MN1020), AT100 monoclonal antibody (Thermo, MN1060) and AT180 monoclonal antibody (Thermo, MN1040). Primary antibodies were incubated for 2 hours at RT and washed five times in TBS-T. Finally, goat-anti human IgG Fab or goat anti-mouse HRP (Jackson Labs) was used at 1:2000 and 1:4000, respectively, and incubated for 1 hour at RT. Plates were washed five times in TBS-T and developed with SureBlue Reserve TMB Microwell Peroxidase Substrate (KPL). The reaction was halted by the addition of 50 .mu.l and 100 .mu.l (peptide plates) of TMB Stop Solution (KPL) and the absorbance at 450 nm was measured using an ELISA plate reader. Supernatants with the aforementioned binding activities were subsequently reconfirmed in an independent ELISA experiment. Once reconfirmed, a clone was selected for downstream IgG expression and purification (Example 5).
Example 5
[0152] IgG1 Expression and Purification of Cloned Anti-Tau Chimeric mAbs
[0153] After ELISA screening and confirmation of antibody reactivity, selected clones were expressed as IgG1 s as indicated in Example 4. Cell culture media was harvested and centrifuged to remove the cells after a minimum of 72 hours and up to a maximum of 168 hours. Clarified supernatants were subsequently passed twice through a Protein A Sepharose column (GE Healthcare Life Sciences) and washed with 50 ml of PBS. IgGs were subsequently eluted with 10 ml of IgG elution buffer (Pierce) and neutralized with Tris pH 8.0 and subsequently dialyzed overnight against PBS. Dialyzed samples were concentrated using a 10,000 MWCO ultra-centrifugal unit (Amicon) to a final volume of about 1 mL, and antibody concentrations were determined with Protein A sensor tips using a human IgG standard on the Octet Red384 (ForteBio). Purified antibodies were further quality controlled by performing SDS-PAGE under non-reducing and reducing conditions and by size exclusion chromatography.
Example 6
[0154] IgG Binding
[0155] Reactivity to Tau Peptides
[0156] IgG1 s generated and quality controlled as described above were tested by ELISA for their ability to bind to specified cognate peptide(s), as well as non-cognate peptide (Table 10). 96-well ELISA plates (Costar) or streptavidin-coated plates (Pierce) were coated with antigen (bovine actin and affinipure goat anti-human F(ab)2) or tau peptides, respectively, as detailed in Example 4. Purified anti-tau IgGs were diluted to 5 .mu.g/ml in TBS containing 0.25% BSA, and titrated five-fold. Antibody controls and secondary antibodies were used as detailed in Example 4. antibody reactivity at 1 .mu.g/mL was determined by ELISA and scored as no binding (-), weak (-/+), moderate (+), or strong (++). (-) for average of two O.D. 450 nm readings <0.3; (-/+) for >0.5 and <1.0; (+) for >1.0 and <1.5; (++) for >1.5.
TABLE-US-00010 TABLE 10 Cognate and non-cognate peptides used in ELISAs Peptide sequence SEQ mAb Peptide (pX) denotes phosphorylated amino acid ID NO Results CBTAU-7.1 ptau 194-212 RSGYSSPG(pS)PG(pT)PGSRSRT 315 + (pS202, pT205) tau 194-212 RSGYSSPGSPGTPGSRSRT 316 - CBTAU-8.1 ptau 194-212 RSGYSSPG(pS)PG(pT)PGSRSRT 315 -/+ (pS202, pT205) tau 194-212 RSGYSSPGSPGTPGSRSRT 316 - CBTAU-16.1 ptau 204-221 GTPGSRSR(pT)P(pS)LPTPPTR 317 ++ (pT212, pS214) tau 204-221 GTPGSRSRTPSLPTPPTR 318 ++ CBTAU-18.1 patu 200-217 PGSPGTPGSR(pS)RTPSLPT 319 -/+ (pS210) tau 200-217 PGSPGTPGSRSRTPSLPT 320 - tau 186-253 GEPPKSGDRSGYSSPGSPGTPGSRSRTPSLPTPPTREP 321 - KKVAVVRTPPKSPSSAKSRLQTAPVPMPDL CBTAU-20.1 ptau 58-76 EPGSETSDAK(pS)(pT)PTAEDVT 322 ++ (pS68, pT69) ptau 59-78 PGSETSDAKS(pT)P(pT)AEDVTAP 323 ++ (pT69, pT71) ptau 61-78 SETSDAKSTP(pT)AEDVTAP 324 -/+ (pT71) tau 42-103 GLKESPLQTPTEDGSEEPGSETSDAKSTPTAEDVTAPL 325 - VDEGAPGKQAAAQPHTEIPEGTTA CBTAU-22.1 ptau 406-429 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA 326 ++ (pS416, pS422) tau 389-441 GAEIVYKSPVVSGDTSPRHLSNVSSTGSIDMVDSPQLA 327 - TLADEVSASLAKQGL CBTAU-24.1 ptau 221-245 REPKKVAVVR(pT)PPKSPS(pS)AKSRLQT 328 ++ (pT231, pS238) ptau 228-245 VVRTPPKSPS(pS)AKSRLQT 329 ++ (pS238) ptau 225-245 KVAVVRTPPK(pS)PS(pS)AKSRLQT 330 ++ (pS235, pS238) tau 186-253 GEPPKSGDRSGYSSPGSPGTPGSRSRTPSLPTPPTREP 321 ++ KKVAVVRTPPKSPSSAKSRLQTAPVPMPDL CBTAU-27.1 tau 299-369 HVPGGGSVQIVYKPVDLSKVTSKCGSLGNIHHKPGGGQ 331 + VEVKSEKLDFKDRVQSKIGSLDNITHVPGGGNK ptau 194-212 RSGYSSPG(pS)PG(pT)PGSRSRT 315 - (pS202, pT205) CBTAU-28.1 tau 42-103 GLKESPLQTPTEDGSEEPGSETSDAKSTPTAEDVTAPL 325 ++ VDEGAPGKQAAAQPHTEIPEGTTA ptau 257-272 KSKIG(pS)TENLKHQPGG 332 - CBTAU-41.1 ptau 406-429 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA 326 + (p416, p422) tau 389-441 GAEIVYKSPVVSGDTSPRHLSNVSSTGSIDMVDSPQLA 327 - TLADEVSASLAKQGL CBTAU-41.2 ptau 406-429 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA 326 + (p416, p422) tau 389-441 GAEIVYKSPVVSGDTSPRHLSNVSSTGSIDMVDSPQLA 327 - TLADEVSASLAKQGL CBTAU-42.1 ptau 406-429 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA 326 ++ (p416, p422) tau 389-441 GAEIVYKSPVVSGDTSPRHLSNVSSTGSIDMVDSPQLA 327 - TLADEVSASLAKQGL CBTAU-43.1 tau 299-369 HVPGGGSVQIVYKPVDLSKVTSKCGSLGNIHHKPGGGQ 331 + VEVKSEKLDFKDRVQSKIGSLDNITHVPGGGNK ptau 194-212 RSGYSSPG(pS)PG(pT)PGSRSRT 315 - (pS202, pT205) CBTAU-44.1 ptau 406-429 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA 326 -/+ (p416, p422) tau 389-441 GAEIVYKSPVVSGDTSPRHLSNVSSTGSIDMVDSPQLA 327 - TLADEVSASLAKQGL CBTAU-45.1 ptau 406-429 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA 326 ++ (p416, p422) tau 389-441 GAEIVYKSPVVSGDTSPRHLSNVSSTGSIDMVDSPQLA 327 - TLADEVSASLAKQGL CBTAU-46.1 tau 42-103 GLKESPLQTPTEDGSEEPGSETSDAKSTPTAEDVTAPL 325 + VDEGAPGKQAAAQPHTEIPEGTTA ptau 224-241 KKVAVVR(pT)PPK(pS)PSSAKS 333 - (pT231, pS235) CBTAU-47.1 tau 42-103 GLKESPLQTPTEDGSEEPGSETSDAKSTPTAEDVTAPL 325 ++ VDEGAPGKQAAAQPHTEIPEGTTA ptau 257-272 KSKIG(pS)TENLKHQPGG 332 - (pS262) CBTAU-47.2 tau 42-103 GLKESPLQTPTEDGSEEPGSETSDAKSTPTAEDVTAPL 325 ++ VDEGAPGKQAAAQPHTEIPEGTTA ptau 257-272 KSKIG(pS)TENLKHQPGG 332 - (pS262) CBTAU-49.1 tau 42-103 GLKESPLQTPTEDGSEEPGSETSDAKSTPTAEDVTAPL 325 ++ VDEGAPGKQAAAQPHTEIPEGTTA ptau 224-241 KKVAVVR(pT)PPK(pS)PSSAKS 333 - (pT231, pS235) AT8 ptau 194-121 RSGYSSPG(pS)PG(pT)PGSRSRT 315 ++ (pS202, pT205) tau 194-212 RSGYSSPGSPGTPGSRSRT 316 - *Amino acid region on human tau441 isoform
[0157] Results are shown in FIGS. 1A-1T. The anti-tau mAbs described herein can be classified into two main groups: those that react only to phosphorylated peptides (phospho-dependent mAbs) and those that react to both phosphorylated and non-phosphorylated peptides (phospho-independent mAbs). Anti-tau mAbs CBTAU-7.1, CBTAU-8.1, CBTAU-18.1, and CBTAU-22.1 were recovered from non-AD individuals using the approach detailed in Example 3. These mAbs are phospho-dependent and, as shown by ELISA (FIGS. 1A-1T), react only with the phosphorylated peptide but not to either a non-phosphorylated peptide spanning that region or a non-phosphorylated version of the peptide. Anti-tau mAbs CBTAU-7.1 and CBTAU-8.1 react specifically to a phosphorylated peptide containing the AT8-binding epitope. This peptide spans amino acids 194 to 212 and contains phosphorylated residues at positions 202 and 205. CBTAU-18.1 reacts to a phosphorylated peptide spanning amino acids 200-217 with a phosphorylated serine residue at position 210. Lastly, CBTAU-22.1 reacts to a peptide spanning amino acids 406-429, with two phosphorylated serines at positions 416 and 422.
[0158] Similarly, CBTAU-20.1 was identified from a non-AD individual and is predominately phospho-dependent as it reacts to three different phosphorylated peptides spanning amino acids 59-77. Two of these peptides are dually phosphorylated, one at positions 68 and 69, and the second at positions 69 and 71. CBTAU-20.1 also reacts to a third peptide that is singly phosphorylated at position 71, suggesting that phosphorylation at threonine 71 is sufficient and important for CBTAU-20.1 reactivity. CBTAU-20.1 shows weak reactivity to a non-phosphorylated peptide spanning region 42-103.
[0159] Like the aforementioned mAbs, CBTAU-16.1 and CBTAU-24.1 were also recovered from non-AD individuals; however, both mAbs are phospho-independent and, as observed by ELISA, react to both a phosphorylated and non-phosphorylated peptide spanning the specified region. CBTAU-16.1 reacts to amino acid region 204-221, whereas CABTAU-24.1 reacts to three different peptides spanning amino acids 221-245. In addition, two additional anti-tau mAbs
[0160] (CBTAU-27.1 and CBTAU-28.1) were identified from screens conducted with non-AD donor samples using 60-70 amino acid-length non-phosphorylated peptides corresponding to amino acid regions 42-103 and 299-369, respectively; therefore, both mAbs are specific to non-phosphorylated-tau.
[0161] Finally, CBTAU mAbs 41.1, 41.2, 42.1, 43.1, 44.1, 45.1, 46.1, 47.1, 47.2, and 49.1 were identified from a small study where 25 young non-AD (18-27 y.o.), 25 non-AD (55+y.o.), and 25 AD (55+ y.o.) individuals were screened. The peptide set that was used for this study included eight phosphorylated peptides (including CBTAU-22.1 cognate peptide) and two non-phosphorylated peptides (CBTAU-27.1 and CBTAU-28.1 cognate peptides). CBTAU mAbs 41.1, 41.2, and 42.1 were recovered from AD donors and react with the CBTAU-22.1 cognate peptide. Similar to CBTAU-22.1, these mAbs are phospho-dependent as shown in FIGS. 1J-1L Two additional mAbs (CBTAU-44.1 and CBTAU-45.1) were identified from non-AD (55+y.o.) individuals with reactivity to the CBTAU-22.1 cognate peptide. As expected, these two were also phospho-dependent (FIGS. 1N and 1O). CBTAU-43.1 was also identified from screens conducted in non-AD (55+y.o.) individuals; however, the mAb was recovered with the CBTAU-27.1 cognate peptide and is specific to non-phosphorylated-tau (FIG. 1M). Lastly, CBTAU-46.1, 47.1, 47.2, and 49.1 were recovered from non-AD (18-27 y.o.) individuals with reactivity to the CBTAU-28.1 peptide and, similar to CBTAU-28.1, are specific to non-phosphorylated-tau (FIGS. 1P-1S).
Example 7
[0162] Reactivity to Paired Helical Filaments and Recombinant Tau by ELISA
[0163] To further characterize the specificity of some of the chimeric antibodies, their reactivity to recombinant tau, enriched and immunopurified paired helical filaments was tested by ELISA.
[0164] PHF-tau was immunopurified according to the protocol of Greenberg and Davies. Briefly, cortical tissues corresponding to Alzheimer's disease individuals were homogenized with 10 volumes of cold buffer (10 mM Tris, pH 7.4, 1 mM EGTA, 0.8 M NaCL and 10% sucrose) and centrifuged at 27,200.times.g for 20 minutes at 4.degree. C. N-lauroylsarcosine and 2-mercaptoethanol were added to the supernatant to reach a final concentration of 1% (wt/vol) and 1% (vol/vol), respectively. The mixture was incubated at 37.degree. C. for 2-2.5 hours with constant rocking, followed by centrifugation at 108,000.times.g for 30 minutes at room temperature. The pellet containing PHF-tau was washed three times with PBS and dissolved in PBS without protein inhibitors and further centrifuged at 12,000.times.g for 5 minutes. The recovered supernatant containing enriched PHF-tau (ePHF-tau) was immunoaffinity purified over an hTau10 affinity column and eluted with 3M or 4 M KSCN overnight at 4.degree. C., followed by dialysis against 1 L PBS at 4.degree. C. with three changes of buffer. hTau10 is an antibody generated in house by immunizing with recombinant tau. It binds to both recombinant and PHF-tau at an amino-terminal epitope. The immunopurified PHF-tau (iPHF-tau) was concentrated with a Sartorius centrifugal filtering device.
[0165] For the ELISA, half-area 96-well binding plates (Costar) were coated with 50 .mu.l of antigen in TBS (2 .mu.g/ml recombinant tau, 2 .mu.g/ml bovine action affinipure goat anti-human F(ab)2, 1 .mu.g/ml of affinity-purified paired helical filaments, and 1 .mu.g/ml of monoclonal anti-tau antibody, HT7 (Thermo Scientific, MN1000). The next day, plates were washed with TBS-T and subsequently blocked with 150 .mu.l of TBS plus 2.5% BSA for 2 hours at RT. Following blocking, ePHF-tau was captured for 2 hours at RT on the anti-tau antibody-coated plate. Purified anti-tau IgGs were diluted to 10 .mu.g/ml in TBS plus 0.25% BSA, and IgGs were titrated five-fold at RT for 2 hours. AT8 (10 .mu.g/ml) was used as a positive control for iPHF-tau and captured ePHF-tau. Plates were washed five times with TBS-T and secondary antibodies, diluted in TBS plus 0.25% BSA, were added and incubated at RT for 1 hour. Goat Anti-Human IgG F(ab')2 (Jackson Labs) was used at a 1:2000 dilution and goat anti-mouse HRP (Jackson Labs) was used at 1:4000 (used for anti-actin control). Following incubation, plates were washed four times in TBS-T and developed with SureBlue Reserve TMB Microwell Peroxidase Substrate (KPL) for approximately 2 minutes. The reaction was immediately halted by the addition of TMB Stop Solution (KPL) and the absorbance at 450 nm was measured using an ELISA plate reader.
[0166] Results are shown in FIGS. 2A-2J. As expected, phospho-dependent mAbs CBTAU-7.1, CBTAU-8.1, and CBTAU-18.1 do not react to recombinant tau by ELISA (FIGS. 2A, 2B, and 2D). CBTAU-20.1 shows minor reactivity to recombinant tau consistent with its weak reactivity to a non-phosphorylated peptide spanning region 42-103. Interestingly, these phospho-dependent mAbs do not show any reactivity to paired helical filaments (i.e., ePHF-tau and iPHF-tau) with the exception of CBTAU-7.1, which shows minor reactivity to ePHF-tau at higher antibody concentrations. Lastly, phospho-dependent CBTAU-22.1 shows no reactivity to recombinant tau, but does react to both iPHF-tau and ePHF-tau (FIG. 2F).
[0167] Phospho-independent anti-tau mAbs, CBTAU-16.1 and CBTAU-24.1 react to both recombinant tau and both formats of paired helical filaments (i.e., iPHF-tau and ePHF-tau; FIGS. 2C and 2G). CBTAU-28.1 shows strong binding to recombinant tau, with weak immunoreactivity to both PHF-tau formats (FIG. 2I). Finally, CBTAU-27.1 shows weak immunoreactivity to both recombinant tau and PHF-tau (FIG. 1H).
Example 8
[0168] Reactivity to Paired Helical Filaments and Recombinant Tau by Western Blot Analysis
[0169] To extend the observations of the rTau- and PHF-binding ELISAs and to examine if secondary structure plays a role in reactivity, recombinant tau, enriched and immunopurified paired helical filaments were tested by Western blot analysis. Approximately 0.5 .mu.g of iPHF, ePHF, and 1 .mu.g of rTau at a final concentration of 1.times. NuPAGE.RTM. LDS Sample buffer (0.5% LDS final) (Novex, NP0007) was heated at 70.degree. C. for 10 minutes. Samples were loaded onto a 26-well, 4-12% Bis-Tris NOVEX.RTM. NuPAGE.RTM. gel (Invitrogen) with MOPS SDS running buffer (Novagen, NP0001), and subsequently transferred onto a nitrocellulose membrane. Membrane was blocked overnight in 1.times. Tris Buffered Saline (TBS) with 0.05% TWEEN.RTM.-20 and 4% non-fat dry milk. CBTAU mAbs were used as primary at 25 .mu.g/mL in 1.times.TBS with 0.05% TWEEN.RTM.-20 and 4% non-fat dry milk and incubated for 2 hours at room temperature. The membrane was then washed three times for 5 minutes each in 1.times.TBS with 0.05% TWEEN.RTM.-20. Peroxidase AffiniPure goat anti-human IgG, Fcy fragment specific (Jackson ImmunoResearch) was then used as secondary at a 1:2000 dilution in 1.times.TBS with 0.05% TWEEN.RTM.-20 and 4% non-fat dry milk and incubated for 45 minutes at RT. The membrane was washed three times for 5 minutes each and developed using the SUPERSIGNAL.RTM. West Pico kit (Pierce).
[0170] The results for the Western blot analysis are shown in FIG. 3. FIG. 3 shows reactivity of three control antibodies, AT8, AT100, and HT7 (two phospho-tau-specific and total tau-specific, respectively). Both AT8 and AT100 show the triple bands characteristic of PHF-tau, which correspond to approximately 68, 64, and 60 kDa. Contrary to the ELISA results, phospho-dependent mAbs, CBTAU-7.1 and CBTAU-18.1 react to both iPHF-tau and ePHF-tau by Western blot, suggesting that the epitopes for these mAbs are not accessible when tau adopts higher order conformations present in PHF-tau. However, these epitopes become accessible under the strong denaturing conditions of SDS-PAGE. CBTAU-27.1 shows binding to recombinant tau and PHF by Western blot yet weak reactivity to each by ELISA, suggesting that the epitope for this antibody is only exposed under strong denaturing conditions. CBTAU-28.1 reacts strongly to recombinant tau by both Western blot and ELISA, and also shows reactivity to PHF-tau by both assays. CBTAU-28.1 reacts to the E1/E2 region of tau (amino acids 42-103), which is not present in all tau isoforms;
[0171] therefore, only the 68 and 64 kDa bands on PHF-tau are detected by CBTAU-28.1. Finally, CBTAU-22.1 and CBTAU-24.1 show similar results to the ELISA assay, reacting to either PHF-tau but not recombinant tau and to both PHF-tau and recombinant tau, respectively.
Example 9
[0172] Reactivity to Tau Fragment Peptides by ELISA
[0173] To characterize the specificity of the recovered antibodies, their reactivity to tau phosphorylated and non-phosphorylated peptides (Tables 11-21, FIGS. 4A-4G) was tested by ELISA. Biotinylated tau peptides were synthesized commercially and dissolved in water at 1 mg/ml and frozen at -80.degree. C. Briefly, 96-well streptavidin-binding plates (Thermo-Fisher) were coated with 2 .mu.g/ml of tau peptides diluted in TBS and incubated overnight at 4.degree. C. The following day, plates were washed with TBS-T and subsequently blocked with 2.5% BSA in TBS for 2 hours at RT. Following blocking, purified anti-tau IgGs were diluted to 2 .mu.g/ml (or to 5 .mu.g/ml and titrated five-fold for finer mapping of CBTAU-27.1, 28.1, 43.1, 46.1, 47.1, 47.2, and 49.1 using peptide sequences in Tables 15-20) in TBS plus 0.25% BSA and incubated at RT for 2 hours. The human-chimerized version of AT8 IgG (at 2 .mu.g/ml) described in Example 11 was used as a positive control in each of the mapping experiments. Plates were washed five times with TBS-T followed by the addition of secondary antibody [goat Anti-Human IgG F(ab')2 (ackson Labs) at 1:2000 dilution], diluted in TBS plus 0.25% BSA, and incubated at RT for 1 hour. Following incubation, plates were washed four times in TBS-T and developed with SureBlue Reserve TMB Microwell Peroxidase Substrate (KPL) for approximately 90 seconds. The reaction was immediately halted by the addition of TMB Stop Solution (KPL) and the absorbance at 450 nm was measured using an ELISA plate reader. Each experiment was conducted in triplicate across three different days. Reactivity was considered positive when values were equal to or higher than an OD of 0.4 in the ELISA assay. For determining the reactivity of each mAb to tau-phosphorylated and non-phosphorylated peptides, antibody reactivity at 2 .mu.g/mL was determined by ELISA and scored as no binding (-), weak (-/+), moderate (+), or strong (++). (-) for average of two O.D. 450 nm readings <0.3; (-/+) for >0.5 and <1.0; (+) for >1.0 and <1.5; (++) for >1.5. For finer mapping of CBTAU-27.1, 28.1, 43.1, 46.1, 47.1, 47.2, and 49.1 (detailed on Tables 15-20), antibody reactivity at 1 .mu.g/mL was determined by ELISA and scored as no binding (-), weak (-/+), moderate (+), or strong (++). (-) for average of three O.D. 450 nm readings <0.3; (-/+) for >0.5 and <1.0; (+) for >1.0 and <1.5; (++) for >1.5.
TABLE-US-00011 TABLE 11 CBTAU-7.1: Peptides for reactivity by ELISA Peptide sequence Peptide SEQ ID NO. (pX) denotes phosphorylated amino acid Results tau 186-253 321 GEPPKSGDRSGYSSPGSPGTPGSRSRTPSLPTPPTREP - KKVAVVRTPPKSPSSAKSRLQTAPVPMPDL ptau 187-212 334 EPPKSGDRSG(pY)SSPGSPG(pT)PGSRSRT -/+ ptau 188-205 335 PPKSGDRSGY(pS)SPGSPGT - ptau 188-206 336 PPKSGDRSGY(pS(pS)PGSPGTP - ptau 188-209 337 PPKSGDRSGY(pS)SPG(pS)PGTPGSR ++ ptau 188-212 338 PPKSGDRSGY(pS)SPGSPG(pT)PGSRSRT -/+ ptau 189-206 339 PKSGDRSGYS(pS)PGSPGTP - ptau 189-209 340 PKSGDRSGYS(pS)PG(pS)PGTPGSR - ptau 189-212 341 PKSGDRSGYS(pS)PGSPG(pT)PGSRSRT + tau 190-209 342 KSGDRSGYSSPGSPGTPGSR - ptau 192-209 343 GDRSGYSSPG(pS)PGTPGSR - ptau 192-212 344 GDRSGYSSPG(pS)PG(pT)PGSRSRT + ptau 192-215 345 GDRSGYSSPG(pS)PGTPG(pS)RSRTPSL - ptau 192-217 346 GDRSGYSSPG(pS)PGTPGSR(pS)RTPSLPT - ptau 194-212 315 RSGYSSPG(pS)PG(pT)PGSRSRT + tau 194-212 316 RSGYSSPGSPGTPGSRSRT -
TABLE-US-00012 TABLE 12 CBTAU-18.1: Peptides for reactivity by ELISA Peptide sequence Peptide SEQ ID NO. (pX) denotes phosphorylated amino acid Results ptau 186-253 321 GEPPKSGDRSGYSSPGSPGTPGSRSRTPSLPTPPTREP - KKVAVVRTPPKSPSSAKSRLQTAPVPMPDL ptau 192-217 346 GDRSGYSSPG(pS)PGTPGSR(pS)RTPSLPT ++ ptau 194-212 315 RSGYSSPG(pS)PG(pT)GSRSRT - tau 194-212* 316 RSGYSSPGSPGTPGSRSRT - ptau 195-212 347 SGYSSPGSPG(pT)PGSRSRT - ptau 195-215 348 SGYSSPGSPG(pT)PG(pS)RSRTPSL - ptau 195-217 349 SGYSSPGSPG(pT)PGSR(pS)RTPSLPT ++ ptau 195-219 350 SGYSSPGSPG(pT)PGSRSR(pT)PSLPTPP - tau 195-214 351 SGYSSPGSPGTPGSRSRTPS - ptau 198-215 352 SSPGSPGTPG(pS)RSRTPSL - ptau 198-217 353 SSPGSPGTPG(pS)R(pS)RTPSLPT ++ ptau 198-219 354 SSPGSPGTPG(pS)RSR(pT)PSLPTPP -/+ ptau 198-221 355 SSPGSPGTPG(pS)RSRTP(pS)LPTPPTR - tau 198-217 356 SSPGSPGTPGSRSRTPSLPT - ptau 200-217 319 PGSPGTPGSR(pS)RTPSLPT + tau 200-217 320 PGSPGTPGSRSRTPSLPT - ptau 200-219 357 PGSPGTPGSR(pS)R(pT)PSLPTPP -/+ ptau 200-221 358 PGSPGTPGSR(pS)RTP(pS)LPTPPTR - ptau 200-224 359 PGSPGTPGSR(pS)RTPSLP(pT)PPTREPK -
TABLE-US-00013 TABLE 13 CBTAU-22.1: Peptides for reactivity by ELISA Peptide sequence Peptide SEQ ID NO. (pX) denotes phosphorylated amino acid Results ptau 404-421 360 SPRHLSNVSS(pT)GSIDMVD - ptau 404-429 361 SPRHLSNVSS(pT)GSIDMVD(pS)PQLATLA ++ tau 405-423 362 PRHLSNVSSTGSIDMVDSP - ptau 406-423 363 RHLSNVSSTG(pS)IDMVDSP - ptau 406-429 326 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA ++ tau 409-428 364 SNVSSTGSIDMVDSPQLATL - ptau 412-429 365 SSTGSIDMVD(pS)PQLATLA ++ ptau 412-434 366 SSTGSIDMVD(pS)PQLA(pT)LADEVSA ++
TABLE-US-00014 TABLE 14 CBTAU-24.1: Peptides for reactivity by ELISA Peptidesequence Peptide SEQ ID NO. (pX) denotes phosphorylated amino acid Results tau 221-253 367 GEPPKSGDRSGYSSPGSPGTPGSRSRTPSLPTPPTREP ++ KKVAVVRTPPKSPSSAKSRLQTAPVPMPDL ptau 221-238 368 REPKKVAVVR(pT)PPKSPSS - ptau 221-242 369 REPKKVAVVR(pT)PPK(pS)PSSAKSR - ptau 221-244 370 REPKKVAVVR(pT)PPKSP(pS)SAKSRLQ - ptau 221-245 328 REPKKVAVVR(pT)PPKSPS(pS)AKSRLQT ++ ptau 225-242 371 KVAVVRTPPK(pS)PSSAKSR - ptau 225-244 372 KVAVVRTPPK(pS)P(pS)SAKSRLQ -/+ ptau 225-245 330 KVAVVRTPPK(pS)PS(pS)AKSRLQT ++ ptau 227-244 373 AVVRTPPKSP(pS)SAKSRLQ ++ ptau 227-245 374 AVVRTPPKSP(pS)(pS)AKSRLQT ++ ptau 228-245 329 VVRTPPKSPS(pS)AKSRLQT ++
TABLE-US-00015 TABLE 15 CBTAU-27.1: Peptides for reactivity by ELISA Peptide sequence Peptide SEQ ID NO. (pX) denotes phosphorylated amino acid Results Cluster 1 tau 299-369 331 HVPGGGSVQIVYKPVDLSKVTSKCGSLGNIHH ++ KPGGGQVEVKSEKIDFKDRVQSKIGSLDNIT HVPGGGNK Cluster 2 ptau 404-421 p414 360 SPRHLSNVSS(pT)GSIDMVD - ptau 404-429 p414, 422 361 SPRHLSNVSS(pT)GSIDMVD(pS)PQLATLA -/+ ptau 406-423 p416 363 RHLSNVSSTG(pS)DMVDSP - ptau 406-429 p416, 422 326 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA -/+ ptau 412-429 p422 365 SSTGSIDMVD(pS)PQLATLA -/+ ptau 412-434 p422, 427 366 SSTGSIDMVD(pS)PQLA(pT)LADEVSA -/+ tau 299-318 375 HVPGGGSVQIVYKPVDLSKV + tau 309-328 376 VYKPVDLSKVTSKCGSLGNI -/+ tau 319-338 377 TSKCGSLGNIHHKPGGGQVE - tau 329-348 378 HHKPGGGQVEVKSEKIDFKD - tau 339-358 379 VKSEKIDFKDRVQSKIGSLD - tau 349-369 380 RVQSKIGSLDNITHVPGGGNK -
TABLE-US-00016 TABLE 16 CBTAU-28.1: Peptides for reactivity by ELISA SEQ Peptide ID NO. Peptide sequence Results tau 42-103 325 GLKFSPLQTPTEDGSEEPGSETSD ++ AKSTPTAEDVTAPLVDEGAPGKQA AAQPHTEIPEGTTA tau 42-61 381 GLKFSPLQTPTEDGSEEPGS - tau 52-71 382 TEDGSEEPGSETSDAKSTPT ++ ptau 58-75 383 EPGSETSDAK(pS)TPTAEDV - tau 62-81 384 ETSDAKSTPTAEDVTAPLVD - tau 72-91 385 AEDVTAPLVDEGAPGKQAAA - tau 82-103 386 EGAPGKQAAAQPHTEIPEGTTA -
TABLE-US-00017 TABLE 17 CBTAU-43.1: Peptides for reactivity by ELISA SEQ Peptide ID NO. Peptide sequence Results tau 299-369 331 HVPGGGSVQIVYKPVDLSKVTS ++ KCGSLGNIHHKPGGGQVEVKSE KIDFKDRVQSKIGSLDNITHVP GGGNK tau 299-318 375 HVPGGGSVQIVYKPVDLSKV ++ tau 309-328 376 VYKPVDLSKVTSKCGSLGNI ++ tau 319-338 377 TSKCGSLGNIHHKPGGGQVE - tau 329-348 378 HHKPGGGQVEVKSEKIDFKD - tau 339-358 379 VKSEKIDFKDRVQSKIGSLD - tau 349-369 380 RVQSKIGSLDNITHVPGGGNK -
TABLE-US-00018 TABLE 18 CBTAU-46.1: Peptides for reactivity by ELISA SEQ Peptide ID NO. Peptide sequence Results tau 42-103 325 GLKFSPLQTPTEDGSEEPGSETSD ++ AKSTPTAEDVTAPLVDEGAPGKQA AAQPHTEIPEGTA tau 42-61 381 GLKFSPLQTPTEDGSEEPGS - tau 52-71 382 TEDGSEEPGSETSDAKSTPT - tau 62-81 384 ETSDAKSTPTAEDVTAPLVD - tau 72-91 385 AEDVTAPLVDEGAPGKQAAA - tau 82-103 386 EGAPGKQAAAQPHTEIPEGTTA ++
TABLE-US-00019 TABLE 19 CBTAU-47.1 and CBTAU-47.2: Peptides for reactivity by ELISA SEQ Peptide ID NO. Peptide sequence Results tau 42-103 325 GLKFSPLQTPTEDGSEEPGSETSD ++ AKSTPTAEDVTAPLVDEGAPGKQA AAQPHTEIPEGTIA tau 42-61 381 GLKFSPLQTPTEDGSEEPGS - tau 52-71 382 TEDGSEEPGSETSDAKSTPT ++ tau 62-81 384 ETSDAKSTPTAEDVTAPLVD - tau 72-91 385 AEDVTAPLVDEGAPGKQAAA - tau 82-103 386 EGAPGKQAAAQPHTEIPEGTTA -
TABLE-US-00020 TABLE 20 CBTAU-49.1: Peptides for reactivity by ELISA SEQ Peptide ID NO. Peptide sequence Results tau 42-103 325 GLKFSPLQTPTEDGSEEPGSETSDA ++ KSTPTAEDVTAPLVDEGAPGKQAAA QPHTEIPEGTIA tau 42-61 381 GLKFSPLQTPTEDGSEEPGS - tau 52-71 382 TEDGSEEPGSETSDAKSTPT ++ tau 62-81 384 ETSDAKSTPTAEDVTAPLVD - tau 72-91 385 AEDVTAPLVDEGAPGKQAAA - tau 82-103 386 EGAPGKQAAAQPHTEIPEGTTA -
TABLE-US-00021 TABLE 20 AT8 Peptides for reactivity by ELISA Peptide sequence Peptide SEQ ID NO. (pX) denotes phosphorylated amino acid Result ptau 189-212 341 PKSGDRSGYS(pS)PGSPG(pT)PGSRSRT - tau 192-211 387 GDRSGYSSPGSPGTPGSRSR - ptau 192-209 343 GDRSGYSSPG(pS)PGTPGSR - ptau 192-212 344 GDRSGYSSPG(pS)PG(pT)PGSRSRT ++ ptau 192-215 345 GDRSGYSSPG(pS)PGTPG(pS)RSRTPSL - ptau 192-217 346 GDRSGYSSPG(pS)PGTPGSR(pS)RTPSLPT - ptau 194-212 315 RSGYSSPG(pS)PG(pT)PGSRSRT ++ tau 194-212* 316 RSGYSSPGSPGTPGSRSRT - ptau 195-212 347 SGYSSPGSPG(pT)PGSRSRT -
[0174] Although CBTAU-7.1 was recovered using a peptide that contains the AT8 epitope (Table 21; 192-212; pS202, pT205), the phospho-residues contributing to binding for CBTAU-7.1 appeared to be promiscuous involving positions S202+T205, but also combinations of S198+S202, S198+T205, S199+T205 and possibly Y197+T205. Unphosphorylated peptides showed no reactivity to CBTAU-7.1. For CBTAU-18.1, the minimal epitope was found to consist of amino acids 198-217 and dependent on pS210 but not when T212, S214 or T217 were also phosphorylated. CBTAU-22.1 reactivity was found to be dependent on pS422, while antibody CBTAU-24.1 revealed strong binding to its corresponding unphosphorylated peptide and, thus, unaffected by phosphorylation.
[0175] CBTAU-27.1 and CBTAU-43.1 were recovered using an unphosphorylated peptide spanning amino acids 299-369. Interestingly, overlapping peptides within this region revealed similar binding requirements for both mAbs (i.e., CBTAU-27.1 and CBTAU-43.1 reacted to peptides spanning amino acids 299-318 and 309-328, respectively), suggesting that the epitope for both mAbs is within regions 299-328 on tau441 (FIGS. 4A and 4C).
[0176] CBTAU-28.1, 46.1, 47.1, 47.2, and 49.1 were recovered from human donor samples using a peptide spanning regions 42-103 on tau441. Testing the reactivity of each mAb against a smaller overlapping set of peptides showed similar binding for CBTAU-47.1, 47.2, and 49.1 as CBTAU-28.1 (i.e., reactivity to a peptide spanning regions 52-71), suggesting comparable binding requirements; however, CBTAU-46.1 bound to a region C-terminal to the aforementioned mAbs (i.e., 82-1031; FIGS. 4B and 4D-4G).
Example 10
[0177] Alanine Scanning of Peptide Epitopes
[0178] To further characterize the specificity and amino acid contribution to binding of each of the recover mAbs, their reactivity to tau peptides with each position replaced with Alanine was tested by ELISA. All experimental protocols were identical to Example 9. Antibody reactivity at 1 .mu.g/mL was determined by ELISA and scored as no binding (-), weak (-/+), moderate (+), or strong (++). (-) for average of two O.D. 450 nm readings <0.3; (-/+) for >0.5 and <1.0; (+) for >1.0 and <1.5; (++) for >1.5. Results for each antibody are shown in Tables 22-29.
TABLE-US-00022 TABLE 22 Alanine scanning results for CBTAU-7.1 and CBTAU-8.1 Region SEQ Peptide sequence Results Results (Tau441) ID NO (pX) denotes phosphorylated amino acid 7.1 8.1 ptau 187-212 334 EPPKSGDRSGYSSPG(pS)PG(pT)PGSRSRT ++ ++ ptau 187-212 388 APPKSGDRSGYSSPG(pS)PG(pT)PGSRSRT + ++ (A187) ptau 187-212 389 EAPKSGDRSGYSSPG(pS)PG(pT)PGSRSRT + ++ (A18) ptau 187-212 390 EPAKSGDRSGYSSPG(pS)PG(pT)PGSRSRT + ++ (A19) ptau 187-212 391 EPPASGDRSGYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A10) ptau 187-212 392 EPPKAGDRSGYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A191) ptau 187-212 393 EPPKSADRSGYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A12) ptau 187-212 394 EPPKSGARSGYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A193) ptau 187-212 395 EPPKSGDASGYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A194) ptau 187-212 396 EPPKSGDRAGYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A195) ptau 187-212 397 EPPKSGDRSAYSSPG(pS)PG(pT)PGSRSRT ++ ++ (A196) ptau 187-212 398 EPPKSGDRSGASSPG(pS)PG(pT)PGSRSRT ++ ++ (A197) ptau 187-212 399 EPPKSGDRSGYASPG(pS)PG(pT)PGSRSRT ++ ++ (A198) ptau 187-212 400 EPPKSGDRSGYSAPG(pS)PG(pT)PGSRSRT + ++ (A199) ptau 187-212 401 EPPKSGDRSGYSSAG(pS)PG(pT)PGSRSRT + ++ (A200) ptau 187-212 402 EPPKSGDRSGYSSPA(pS)PG(pT)PGSRSRT ++ ++ (A201) ptau 187-212 403 EPPKSGDRSGYSSPGAPG(pT)PGSRSRT + ++ (A202) ptau 187-212 404 EPPKSGDRSGYSSPG(pS)AG(pT)PGSRSRT ++ ++ (A203) ptau 187-212 405 EPPKSGDRSGYSSPG(pS)PA(pT)PGSRSRT -/+ + (A204) ptau 187-212 406 EPPKSGDRSGYSSPG(pS)PGAPGSRSRT -/+ -/+ (A205) ptau 187-212 407 EPPKSGDRSGYSSPG(pS)PG(pT)AGSRSRT -/+ - (A206) ptau 187-212 408 EPPKSGDRSGYSSPG(pS)PG(pT)PASRSRT ++ -/+ (A207) ptau 187-212 409 EPPKSGDRSGYSSPG(pS)PG(pT)PGARSRT ++ ++ (A208) ptau 187-212 410 EPPKSGDRSGYSSPG(pS)PG(pT)PGSASRT ++ - (A209) ptau 187-212 411 EPPKSGDRSGYSSPG(pS)PG(pT)PGSRART ++ ++ (A210) ptau 187-212 412 EPPKSGDRSGYSSPG(pS)PG(pT)PGSRSAT ++ ++ (A211) ptau 187-212 413 EPPKSGDRSGYSSPG(pS)PG(pT)PGSRSRA ++ ++ (A212)
TABLE-US-00023 TABLE 23 Alanine scanning results for CBTAU-22.1 Region SEQ Peptide sequence (Tau441) ID NO (pX) denotes phosphorylated amino acid Results ptau 406-429 326 RHLSNVSSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A406) 414 AHLSNVSSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A407) 415 RALSNVSSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A408) 416 RHASNVSSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A409) 417 RHLANVSSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A410) 418 RHLSAVSSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A411) 419 RHLSNASSTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A412) 420 RHLSNVASTG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A413) 421 RHLSNVSATG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A414) 422 RHLSNVSSAG(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A415) 423 RHLSNVSSTA(pS)IDMVD(pS)PQLATLA ++ ptau 406-429 (A416) 424 RHLSNVSSTGAIDMVD(pS)PQLATLA ++ ptau 406-429 (A417) 425 RHLSNVSSTG(pS)ADMVD(pS)PQLATLA ++ ptau 406-429 (A418) 426 RHLSNVSSTG(pS)IAMVD(pS)PQLATLA ++ ptau 406-429 (A419) 427 RHLSNVSSTG(pS)IDAVD(pS)PQLATLA ++ ptau 406-429 (A420) 428 RHLSNVSSTG(pS)IDMAD(pS)PQLATLA ++ ptau 406-429 (A421) 429 RHLSNVSSTG(pS)IDMVA(pS)PQLATLA -/+ ptau 406-429 (A422) 430 RHLSNVSSTG(pS)IDMVDAPQLATLA - ptau 406-429 (A423) 431 RHLSNVSSTG(pS)IDMVD(pS)AQLATLA + ptau 406-429 (A424) 432 RHLSNVSSTG(pS)IDMVD(pS)PALATLA ++ ptau 406-429 (A425) 433 RHLSNVSSTG(pS)IDMVD(pS)PQAATLA + ptau 406-429 (A427) 434 RHLSNVSSTG(pS)IDMVD(pS)PQLAALA ++ ptau 406-429 (A428) 435 RHLSNVSSTG(pS)IDMVD(pS)PQLATAA ++
TABLE-US-00024 TABLE 24 CBTAU-24.1 Alanine Scan Results SEQ Peptide sequence (pX) Region ID denotes phosphorylated Re- (Tau441) NO amino acid sults ptau 221-245 328 REPKKVAVVR(pT)PPKSPS ++ (pS)AKSRLQT ptau 221-245 436 RAPKKVAVVR(pT)PPKSPS ++ (A222) (pS)AKSRLQT ptau 221-245 437 REAKKVAVVR(pT)PPKSPS ++ (A223) (pS)AKSRLQT ptau 221-245 438 REPAKVAVVR(pT)PPKSPS ++ (A224) (pS)AKSRLQT ptau 221-245 439 REPKAVAVVR(pT)PPKSPS ++ (A225) (pS)AKSRLQT ptau 221-245 440 REPKKAAVVR(pT)PPKSPS ++ (A226) (pS)AKSRLQT ptau 221-245 441 REPKKVAAVR(pT)PPKSPS ++ (A228) (pS)AKSRLQT ptau 221-245 442 REPKKVAVAR(pT)PPKSPS ++ (A229) (pS)AKSRLQT ptau 221-245 443 REPKKVAVVA(pT)PPKSPS ++ (A230) (pS)AKSRLQT ptau 221-245 444 REPKKVAVVRAPPKSPS ++ (A231) (pS)AKSRLQT ptau 221-245 445 REPKKVAVVR(pT)APKSPS ++ (A232) (pS)AKSRLQT ptau 221-245 446 REPKKVAVVR(pT)PAKSPS ++ (A233) (pS)AKSRLQT ptau 221-245 447 REPKKVAVVR(pT)PPASPS ++ (A234) (pS)AKSRLQT ptau 221-245 448 REPKKVAVVR(pT)PPKAPS ++ (A235) (pS)AKSRLQT ptau 221-245 449 REPKKVAVVR(pT)PPKSAS - (A236) (pS)AKSRLQT ptau 221-245 450 REPKKVAVVR(pT)PPKSPA ++ (A237) (pS)AKSRLQT ptau 221-245 451 REPKKVAVVR(pT) ++ (A238) PPKSPSAAKSRLQT ptau 221-245 452 REPKKVAVVR(pT) ++ (A240) PPKSPS(pS)AASRLQT ptau 221-245 453 REPKKVAVVR(pT) ++ (A241) PPKSPS(pS)AKARLQT ptau 221-245 454 REPKKVAVVR(pT) ++ (A242) PPKSPS(pS)AKSALQT ptau 221-245 455 REPKKVAVVR(pT) ++ (A243) PPKSPS(pS)AKSRAQT ptau 221-245 456 REPKKVAVVR(pT) ++ (A244) PPKSPS(pS)AKSRLAT ptau 221-245 457 REPKKVAVVR(pT) ++ (A245) PPKSPS(pS)AKSRLQA
TABLE-US-00025 TABLE 25 CBTAU-27.1 Alanine Scan Results SEQ Region ID Re- (Tau441) NO: Peptide sequence sults tau 299-323 458 HVPGGGSVQIVYKPVDLSKVTSKC ++ G tau 299-323 459 AVPGGGSVQIVYKPVDLSKVTSKCG ++ (A299) tau 299-323 460 HAPGGGSVQIVYKPVDLSKVTSKC ++ (A300) G tau 299-323 461 HVAGGGSVQIVYKPVDLSKVTSKC ++ (A301) G tau 299-323 462 HVPAGGSVQIVYKPVDLSKVTSKCG ++ (A302) tau 299-323 463 HVPGAGSVQIVYKPVDLSKVTSKCG ++ (A303) tau 299-323 464 HVPGGASVQIVYKPVDLSKVTSKCG ++ (A304) tau 299-323 465 HVPGGGAVQIVYKPVDLSKVTSKC ++ (A305) G tau 299-323 466 HVPGGGSAQIVYKPVDLSKVTSKC ++ (A306) G tau 299-323 467 HVPGGGSVAIVYKPVDLSKVTSKCG ++ (A307) tau 299-323 468 HVPGGGSVQAVYKPVDLSKVTSKC ++ (A308) G tau 299-323 469 HVPGGGSVQIAYKPVDLSKVTSKC ++ (A309) G tau 299-323 470 HVPGGGSVQIVAKPVDLSKVTSKC ++ (A310) G tau 299-323 471 HVPGGGSVQIVYAPVDLSKVTSKCG ++ (A311) tau 299-323 472 HVPGGGSVQIVYKAVDLSKVTSKC ++ (A312) G tau 299-323 473 HVPGGGSVQIVYKPADLSKVTSKC ++ (A313) G tau 299-323 474 HVPGGGSVQIVYKPVALSKVTSKCG -/+ (A314) tau 299-323 475 HVPGGGSVQIVYKPVDASKVTSKC - (A315) G tau 299-323 476 HVPGGGSVQIVYKPVDLAKVTSKC ++ (A316) G tau 299-323 477 HVPGGGSVQIVYKPVDLSAVTSKCG - (A317) tau 299-323 478 HVPGGGSVQIVYKPVDLSKATSKC ++ (A318) G tau 299-323 479 HVPGGGSVQIVYKPVDLSKVASKC ++ (A319) G tau 299-323 480 HVPGGGSVQIVYKPVDLSKVTAKC ++ (A320) G tau 299-323 481 HVPGGGSVQIVYKPVDLSKVTSACG ++ (A321) tau 299-323 482 HVPGGGSVQIVYKPVDLSKVTSKA ++ (A322) G tau 299-323 483 HVPGGGSVQIVYKPVDLSKVTSKCA ++ (A323)
TABLE-US-00026 TABLE 26 CBTAU-28.1 Alanine Scan Results SEQ Region ID Re- (Tau441) NO Peptide sequence sults tau 52-71 382 TEDGSEEPGSETSDAKSTPT ++ tau 52-71 (A52) 484 AEDGSEEPGSETSDAKSTPT ++ tau 52-71 (A53) 485 TADGSEEPGSETSDAKSTPT ++ tau 52-71 (A54) 486 TEAGSEEPGSETSDAKSTPT ++ tau 52-71 (A55) 487 TEDASEEPGSETSDAKSTPT ++ tau 52-71 (A56) 488 TEDGAEEPGSETSDAKSTPT ++ tau 52-71 (A57) 489 TEDGSAEPGSETSDAKSTPT ++ tau 52-71 (A58) 490 TEDGSEAPGSETSDAKSTPT ++ tau 52-71 (A59) 491 TEDGSEEAGSETSDAKSTPT -/+ tau 52-71 (A60) 492 TEDGSEEPASETSDAKSTPT ++ tau 52-71 (A61) 493 TEDGSEEPGAETSDAKSTPT ++ tau 52-71 (A62) 494 TEDGSEEPGSATSDAKSTPT - tau 52-71 (A63) 495 TEDGSEEPGSEASDAKSTPT -/+ tau 52-71 (A64) 496 TEDGSEEPGSETADAKSTPT ++ tau 52-71 (A65) 497 TEDGSEEPGSETSAAKSTPT - tau 52-71 (A67) 498 TEDGSEEPGSETSDAASTPT - tau 52-71 (A68) 499 TEDGSEEPGSETSDAKATPT ++ tau 52-71 (A69) 500 TEDGSEEPGSETSDAKSAPT ++ tau 52-71 (A70) 501 TEDGSEEPGSETSDAKSTAT ++ tau 52-71 (A71) 502 TEDGSEEPGSETSDAKSTPA ++
TABLE-US-00027 TABLE 27 CBTAU-43.1 Alanine Scan Results SEQ Region ID Re- (Tau441) NO: Peptide sequence sults tau 299-323 458 HVPGGGSVQIVYKPVDLSKVTSKCG ++ tau 299-323(A299) 459 AVPGGGSVQIVYKPVDLSKVTSKCG ++ tau 299-323(A300) 460 HAPGGGSVQIVYKPVDLSKVTSKCG ++ tau 299-323(A301) 461 HVAGGGSVQIVYKPVDLSKVTSKCG ++ tau 299-323(A302) 462 HVPAGGSVQIVYKPVDLSKVTSKCG ++ tau 299-323(A303) 463 HVPGAGSVQIVYKPVDLSKVTSKCG ++ tau 299-323(A304) 464 HVPGGASVQIVYKPVDLSKVTSKCG ++ tau 299-323(A305) 465 HVPGGGAVQIVYKPVDLSKVTSKCG ++ tau 299-323(A306) 466 HVPGGGSAQIVYKPVDLSKVTSKCG ++ tau 299-323(A307) 467 HVPGGGSVAIVYKPVDLSKVTSKCG ++ tau 299-323(A308) 468 HVPGGGSVQAVYKPVDLSKVTSKCG ++ tau 299-323(A309) 469 HVPGGGSVQIAYKPVDLSKVTSKCG ++ tau 299-323(A310) 470 HVPGGGSVQIVAKPVDLSKVTSKCG ++ tau 299-323(A311) 471 HVPGGGSVQIVYAPVDLSKVTSKCG ++ tau 299-323(A312) 472 HVPGGGSVQIVYKAVDLSKVTSKCG -/+ tau 299-323(A313) 473 HVPGGGSVQIVYKPADLSKVTSKCG ++ tau 299-323(A314) 474 HVPGGGSVQIVYKPVALSKVTSKCG ++ tau 299-323(A315) 475 HVPGGGSVQIVYKPVDASKVTSKCG - tau 299-323(A316) 476 HVPGGGSVQIVYKPVDLAKVTSKCG ++ tau 299-323(A317) 477 HVPGGGSVQIVYKPVDLSAVTSKCG - tau 299-323(A318) 478 HVPGGGSVQIVYKPVDLSKATSKCG ++ tau 299-323(A319) 479 HVPGGGSVQIVYKPVDLSKVASKCG ++ tau 299-323(A320) 480 HVPGGGSVQIVYKPVDLSKVTAKCG ++ tau 299-323(A321) 481 HVPGGGSVQIVYKPVDLSKVTSACG ++ tau 299-323(A322) 482 HVPGGGSVQIVYKPVDLSKVTSKAG ++ tau 299-323(A323) 483 HVPGGGSVQIVYKPVDLSKVTSKCA ++
TABLE-US-00028 TABLE 28 CBTAU-47.1 and 47.2 Alanine Scan Results SEQ Results Results Region ID CBTAU- CBTAU- (Tau441) NO Peptide sequence 47.1 47.2 tau 52-71 382 TEDGSEEPGSETSDAKSTPT ++ ++ tau 52-71 484 AEDGSEEPGSETSDAKSTPT ++ ++ (A52) tau 52-71 485 TADGSEEPGSETSDAKSTPT ++ ++ (A53) tau 52-71 486 TEAGSEEPGSETSDAKSTPT ++ ++ (A54) tau 52-71 487 TEDASEEPGSETSDAKSTPT ++ ++ (A55) tau 52-71 488 TEDGAEEPGSETSDAKSTPT ++ ++ (A56) tau 52-71 489 TEDGSAEPGSETSDAKSTPT ++ ++ (A57) tau 52-71 490 TEDGSEAPGSETSDAKSTPT ++ ++ (A58) tau 52-71 491 TEDGSEEAGSETSDAKSTPT - - (A59) tau 52-71 492 TEDGSEEPASETSDAKSTPT ++ ++ (A60) tau 52-71 493 TEDGSEEPGAETSDAKSTPT -/+ ++ (A61) tau 52-71 494 TEDGSEEPGSATSDAKSTPT - - (A62) tau 52-71 495 TEDGSEEPGSEASDAKSTPT - - (A63) tau 52-71 496 TEDGSEEPGSETADAKSTPT ++ ++ (A64) tau 52-71 497 TEDGSEEPGSETSAAKSTPT - - (A65) tau 52-71 498 TEDGSEEPGSETSDAASTPT - - (A67) tau 52-71 499 TEDGSEEPGSETSDAKATPT ++ ++ (A68) tau 52-71 500 TEDGSEEPGSETSDAKSAPT ++ ++ (A69) tau 52-71 501 TEDGSEEPGSETSDAKSTAT ++ ++ (A70) tau 52-71 502 TEDGSEEPGSETSDAKSTPA ++ ++ (A71)
TABLE-US-00029 TABLE 29 CBTAU-49.1 Alanine Scan Results SEQ Region ID Re- (Tau441) NO Peptide sequence sults tau 52-71 382 TEDGSEEPGSETSDAKSTPT ++ tau 52-71 (A52) 484 AEDGSEEPGSETSDAKSTPT ++ tau 52-71 (A53) 485 TADGSEEPGSETSDAKSTPT ++ tau 52-71 (A54) 486 TEAGSEEPGSETSDAKSTPT ++ tau 52-71 (A55) 487 TEDASEEPGSETSDAKSTPT ++ tau 52-71 (A56) 488 TEDGAEEPGSETSDAKSTPT ++ tau 52-71 (A57) 489 TEDGSAEPGSETSDAKSTPT ++ tau 52-71 (A58) 490 TEDGSEAPGSETSDAKSTPT ++ tau 52-71 (A59) 491 TEDGSEEAGSETSDAKSTPT - tau 52-71 (A60) 492 TEDGSEEPASETSDAKSTPT ++ tau 52-71 (A61) 493 TEDGSEEPGAETSDAKSTPT - tau 52-71 (A62) 494 TEDGSEEPGSATSDAKSTPT - tau 52-71 (A63) 495 TEDGSEEPGSEASDAKSTPT + tau 52-71 (A64) 496 TEDGSEEPGSETADAKSTPT + tau 52-71 (A65) 497 TEDGSEEPGSETSAAKSTPT - tau 52-71 (A67) 498 TEDGSEEPGSETSDAASTPT - tau 52-71 (A68) 499 TEDGSEEPGSETSDAKATPT + tau 52-71 (A69) 500 TEDGSEEPGSETSDAKSAPT + tau 52-71 (A70) 501 TEDGSEEPGSETSDAKSTAT + tau 52-71 (A71) 502 TEDGSEEPGSETSDAKSTPA +
[0179] Although CBTAU-7.1 and CBTAU-8.1 were recovered using a tau phosphopeptide containing the AT8 epitope (i.e., pS202, pT205), both mAbs exhibited different epitope requirements according to the alanine scan results (Table 22). In addition to S202 and T205, substitutions at positions G204 and P206 resulted in reduced binding for CBTAU-7.1. In contrast, alanine substitutions at positions G204, T205, P206, and R209 reduced the reactivity of CBTAU-8.1 to the peptide, yet the S202A substitution had no effect. Like AT8, both mAbs are phospho-dependent, but require additional (non-phosphorylated residues) for binding. The alanine scan results for CBTAU-22.1 showed a dependency on phosphorylation at S422 (Table 23), as substitution at this position completed inhibited binding. Substitution at D421 resulted in a reduction but not a complete inhibition in binding. Finally, alanine scan results for CBTAU-24.1 showed P236 to be the only critical residue for binding (Table 24).
[0180] To map the critical contact residues for CBTAU-27.1 and 43.1, alanine scanning was also conducted within regions 299-323 of tau (Table 25 and Table 27, respectively). The critical contact residues for CBTAU-27.1 binding were shown to be D314, L315, and K317. The results suggest that residues D314 and K317 may form salt bridge interactions between the epitope and CDR residues on the mAb. While CBTAU-43.1 was recovered using the cognate peptide for CBAU-27.1, the critical residues according to the alanine scan were different. In addition to L315 and K317, the proline at position 312 was shown to be an important contact for CBTAU-43.1 binding. Lastly, alanine scanning was also conducted for CBTAU-28.1 as well as to CBTAU-47.1, 47.1, and 49.1 (Tables 26, 28, 29). As shown in Example 9, CBTAU mAbs 47.1, 47.2, 49.1 mapped to the same peptide region as CBTAU-28.1 (i.e., 52-71). Interestingly, all mAbs shared identical binding requirements as to CBTAU-28.1. The critical contact residues were shown to be P59, S61, E62, T63, D65, and K67. Several of these residues were found to be charged, implicating important salt bridge interactions between the epitope and the mAbs.
Example 11
[0181] Immunohistochemistry
[0182] Tau pathology is believed to initiate within the entorhinal cortex (EC) and spread along connected neuronal pathways in the hippocampus before progressing into the cortex. To determine the reactivity of the recovered IgGs to pathogenic deposits of tau along these neuronal pathways, hippocampal tissues were obtained from an 82-year-old, non-diseased (non-AD; Abcam, Cat No. ab4305) male and an 88-year-old Alzheimer's disease (AD; Abcam, Cat. No. ab4583) male (Abcam). Cortical tissues were obtained from a 71-year-old non-diseased (non-AD) and 71-year-old Alzheimer's disease (AD) individual (Banner Sun Health). In addition to AD, there are many neurological disorders that are characterized by tau pathology, also known as tauopathies. To extend the findings, the recovered mAbs were tested in tissues obtained from progressive supranuclear palsy (PSP) and non-progressive supranuclear palsy (non-PSP) frontal lobes obtained from a 73-year-old male and an 81-year-old female, respectively (Biochain). Brain tissues were de-paraffinized and rehydrated by washing twice for 10 minutes in xylene (VWR International), followed by washing twice for 3 minutes in 100% ethanol, twice for 3 minutes in 95% ethanol, twice for 3 minutes in 70% ethanol, and once for 30 seconds in distilled H2O using TISSUE-TEK.RTM. Slide Staining Set (VWR Intemational). Tissue sections underwent heat-mediated antigen retrieval using citrate buffer (10 mM citric acid, pH 6.0) to expose antigenic sites. Sections were then incubated with blocking buffer [10% normal goat serum (Jackson ImmunoResearch, Inc.), 1% BSA and 0.3% TRITON.RTM. X-100 in PBS)] at RT for 1 hour. Excess water was removed and tissue sections were circled with an ImmEdge Hydrophobic Barrier Pen (Vector Labs). A humidified chamber was prepared by covering the bottom of a staining tray with H2O, and sections were then washed with PBS three times for 5 minutes by aspiration. Endogenous peroxidase activity was quenched in 10% H2O2 for 30 minutes at RT. Following quenching, slides were washed with PBS three times for 5 minutes by aspiration. Slides were then blocked for 1 hour at RT with a solution of 10% normal goat serum, 0.3% TRITON.RTM. X-100, 1% BSA in 1.times.PBS. Primary antibodies were labeled with biotin using the Zenon Human IgG Labeling Kit (Life Technologies) per manufacturer's instructions. As a negative control, a human anti-RSV-specific antibody was used. An Fc-region human-chimerized version of AT8 IgG was used as a positive control. After labeling, primary antibodies were diluted separately in blocking buffer at concentrations of 5 .mu.g/ml and 20 .mu.g/ml. For peptide competition experiments, 13.3 .mu.M of cognate peptide (i.e., peptide used to recover the mAb in sorting experiments) was pre-incubated with the primary antibody for 30 minutes at RT prior to incubation with tissue sections. Tissue sections were incubated at RT for two hours with 100 .mu.l of diluted biotin-labeled primary antibody or peptide competed antibody. After antibody was removed by aspiration, a second fixation of the tissue section was performed in 4% formaldehyde in PBS and incubated for 15 minutes at RT. The section was washed with PBS three times for 5 minutes by aspiration. Sections were then incubated for 30 minutes with streptavidin substrate VECTASTAIN.RTM. ABC Reagent (Vector Labs) before washing with PBS. Tissues were then developed with DAB substrate (Vector Labs) in the presence of nickel. Sections were then washed two times with ddH2O and allowed to completely dry at RT before mounting with 50 .mu.l of VectaMount Permanent Mounting Medium (Vector Labs). Finally, tissue sections were counterstained with hematoxylin (Vector Labs). Representative images were acquired with Olympus BX-41 upright microscope using METAMORPH.RTM. software.
[0183] Results from the immunohistochemistry are shown in FIGS. 5A-5D. CBTAU-7.1 and CBTAU-8.1 showed positive immunoreactivity on AD brain tissues specifically, and not to healthy brain tissues, which suggests binding to pathogenic tau deposits present in diseased brain tissues. These antibodies recognize AT8-positive tau tangles and neutrophil threads in subregions of the hippocampus (FIG. 5A; entorhinal cortex) and cerebral cortex (FIG. 5B). Furthermore, the positive immunoreactivity was consistently found across multiple experiments in the neuronal cytoplasm and processes. In addition, CBTAU-18.1, 22.1, and 24.1 were also tested against hippocampal and cortical tissue sections (FIGS. 5A and 5B). Similar to CBTAU-7.1 and CBTAU-8.1, all mAbs reacted specifically to tau on AD tissue sections but not to tau on non-AD tissue sections. Of interest, CBTAU-24.1, which is not specific to phosphorylated tau, reacts specifically to diseased tau on AD tissue sections but not to tau on non-AD sections. Finally, CBTAU-16.1 and CBTAU-20.1 show reactivity to tau on both non-AD and AD tissue sections.
[0184] In addition, CBTAU-7.1, 8.1, 16.1, 18.1, 20.1, 22.1, and 24.1 were tested on cortical tissue sections corresponding to progressive supranuclear palsy (FIG. 5C). Unlike AT8, CBTAU-7.1 and CBTAU-8.1 failed to detect tau tangles in the human PSP brain, suggesting that the epitope for both mAbs is not present on PSP. CBTAU-16.1 and CBTAU-20.1 showed positive immunoreactivity to tau on non-PSP and PSP cortical brain sections, suggesting binding to both normal tau and pathogenic forms of tau. In non-AD brain sections, these antibodies showed positive immunostaining of tau in neuronal cytoplasm and processes (FIGS. 5A and 5B), yet both mAbs detected tangles and neutrophil threads in AD brain sections similar to AT8. Similar immunoreactivity as AT8 to tau tangles was also detected in PSP brain tissue sections, suggesting that both CBTAU-16.1 and CBTAU-20.1 recognize common pathogenic tau forms in other non-AD tauopathies. Furthermore, CBTAU-22.1 and CBTAU-24.1 showed immunoreactivity exclusively in AD brain tissues, with positive immunoreactivity to tangles and neutrophil threads. CBTAU-18.1 showed weak immunoreactivity in non-AD brain tissues, yet reacted stronger to AD tissue samples. CBTAU-18.1, CBTAU-22.1 and CBTAU-24.1 were also positive for tau tangles in PSP brain tissue sections (FIG. 5C).
[0185] CBTAU-27.1 and CBTAU-28.1 showed selective immunostaining in non-AD tissue sections with diffuse immunostaining in neuronal cytoplasm and processes. Interestingly, both antibodies failed to show immunoreactivity in AD tissue sections (both hippocampal and cortical), defining a novel epitope that is lost during disease progression. Unlike the majority of the human anti-tau mAbs that were identified, CBTAU-27.1 and CBTAU-28.1 were recovered by screening donor samples using an unphosphorylated peptide set spanning the entire region of human tau441. These antibodies do not require phosphorylation for binding (FIGS. 1A-1T) and, as shown in FIGS. 2A-2J, do not react to PHF by ELISA. Therefore, the diffuse immunostaining pattern observed for these two mAbs was expected. In addition, CBTAU-43.1, which was originally recovered using the CBTAU-27.1 cognate peptide, was tested against cortical tissue sections. CBTAU-43.1 reacted similar to CBTAU-27.1, staining tau on non-AD but not tau on AD tissue sections. Likewise, CBTAU-46.1, 47.2 (only one variant tested), and 49.1, which were recovered using the CBTAU-28.1 cognate peptide, reacted specifically to tau on non-AD but not AD tissue sections (FIG. 5D). It is interesting to note that these mAbs all share common heavy and light chain germlines (i.e., VH5-51 and VK4-1), bind to the same regions on tau and, as shown in FIG. 5D, share similar immunohistochemical properties.
[0186] The immunohistochemistry results presented here for CBTAU-7.1, 8.1, 18.1, 22.1, 24.1, 27.1, and 28.1 have been confirmed on multiple regions of the brain and tissue samples corresponding to several non-AD and AD individuals. Immunoreactivity of CBTAU mAbs 43.1, 46.1, 47.2, and 49.1 has been confirmed once using the same tissue sample and has not yet been confirmed on samples corresponding to other AD and non-AD individuals.
Example 12
[0187] Dephosphorylation IHC
[0188] Given that the results of the IHC for CBTAU-28.1 showed immunoreactivity to tau on non-AD tissue sections but not to tau on AD tissue sections, it was hypothesized that the loss of this epitope during disease progression was a result of modification(s) (i.e., phosphorylation). To test this hypothesis, human brain tissue sections were dephosphorylated prior to assessing immunoreactivity of CBTAU-28.1. Paraffin-embedded human brain tissue sections (Abcam, cat #:ab4305, 54-year-old male, no clinical symptom vs. Abcam, cat #:ab4583, 93-year-old Hispanic female, Alzheimer disease) were deparaffinized and rehydrated by washing twice for 10 minutes in xylene (VWR International), followed by washing twice for 3 minutes in 100% ethanol, twice for 3 minutes in 95% ethanol, twice for 3 minutes in 70% ethanol, and once for 30 seconds in distilled H2O using TISSUE-TEK.RTM. Slide Staining Set (VWR International). To minimize non-specific antibody binding, tissues were never allowed to dry during washes. Tissue sections underwent heat-mediated antigen retrieval using citrate buffer {citric acid, pH 6.0) to expose antigenic sites. Excess water was removed and tissue sections were circled with an ImmEdge Hydrophobic Barrier Pen (Vector Labs). A humidified chamber was prepared by covering the bottom of a staining tray with H2O, and sections were then washed with PBS three times for 5 minutes by aspiration. Endogenous peroxidase activity was quenched in H2O2 for 15 minutes at RT. Following quenching, slides were washed with PBS three times for 5 minutes by aspiration. Sections were subsequently treated with 130 units/mL of calf intestinal alkaline phosphatase (CIAP) for 2.5 hours at 32 degrees. Slides were then blocked for 1 hour at RT with a solution of 10% normal goat serum, 0.3% TRITON.RTM. X-100, 1% BSA in 1.times.PBS. Murinized CBTAU-28.1 (Fc region murinized), and control mAbs AT8 and isotype control (anti-RSV mAb 4.1) were incubated overnight on hippocampal sections at a final concentration of 1 .mu.g/mL Sections were washed and incubated with anti-mouse, Fcy fragment-specific antibody for 2 hours at room temperature. Samples were developed with peroxidase substrate solution DAB in the presence of Nickel. Samples were counterstained with hematoxylin (Vector Labs). Representative images were acquired with Olympus BX-41 upright microscope using METAMORPH.RTM. software.
[0189] Results are shown in FIGS. 6A and 6B. As expected, CBTAU-28.1 reacts to tau present in the non-AD hippocampal tissue sections but does not react to tau in the AD tissue sections. In contrast, control mAb, AT8, does not react to tau in non-AD sections, but clearly reacts to pathogenic tau deposits present in the AD sections (FIG. 6A). However, pretreatment of AD tissue sections with phosphatase restores reactivity of CBTAU-28.1, allowing it to stain pathogenic tau deposits present in these sections. As expected, the reactivity of AT8 was reduced with pretreatment of the AD tissue sections with phosphatase (FIG. 6B).
Example 13
[0190] Dephosphorylation ELISA
[0191] To confirm the results of Example 12, dephosphorylation of paired helical filaments was tested for reactivity to CBTAU-28.1 by ELISA. Half-area 96-well binding plates (Costar) were coated with 50 .mu.l of antigen in TBS (2 .mu.g/ml bovine action affinipure goat anti-human F(ab)2, and 1 .mu.g/ml of affinity-purified paired helical filaments, iPHF, pretreated with and without calf intestinal phosphatase, CIP). Phosphatase-treated iPHF was prepared as follows. iPHF samples were resuspended in 1.times.NEB buffer 4 (50 mM potassium acetate, 20 mM Tris-acetate, 10 mM magnesium acetate, and 1 mM DTT) at a final concentration of 0.05 .mu.g/ml. One unit of CIP per .mu.g of iPHF was added (CIP, NEB Cat. No. M0290S). iPHF samples were incubated with CIP for 90 minutes at 37.degree. C. prior to coating on ELISA binding plates. Following antigen binding overnight, the plates were washed with TBS-T and subsequently blocked with 150 .mu.l of TBS plus 2.5% BSA for 2 hours at RT. Purified control and anti-tau IgG, CBTAU-28.1), were titrated in five-fold dilutions starting at 25 .mu.g/ml in TBS/0.25% BSA, and IgGs and incubated for 1.5 hours. Plates were washed four times with TBS-T and secondary antibody (anti-human Fab HRP, Jackson Immunoresearch, Cat. No. 109-036-097) was added and incubated at RT for 45 minutes. Following incubation, plates were washed four times in TBS-T and developed with SureBlue Reserve TMB Microwell Peroxidase Substrate (KPL) for approximately 2 minutes. The reaction was immediately halted by the addition of TMB Stop Solution (KPL) and the absorbance at 450 nm was measured using an ELISA plate reader. Each experimental point was performed in triplicate.
[0192] Results are shown in FIG. 7. As previously shown in Example 7, CBTAU-28.1 reacts poorly to iPHF by ELISA in contrast to AT8. However, dephosphorylation of iPHF with CIP restores the reactivity of CBTAU-28.1 to the filamentous sample. As expected, the reactivity of the phospho-tau control mAb, AT8, is abolished after dephosphorylation of iPHF with CIP.
Example 14
[0193] Reactivity of CBTAU-27.1, 28.1, 43.1, 47.1, 47.2 and 49.1 to Phosphopeptides
[0194] The immunohistochemical results for CBTAU-27.1 (and CBTAU-43.1) and CBTAU-28.1 (and CBTAU-46.1, 47.2, 49.1) suggest that these mAbs react with an epitope on tau that is present in normal, non-AD, tissue sections but is lost or masked during the disease setting (FIGS. 5A-5D). It was hypothesized that this was a result of a phosphorylation event within the region that results in masking of the epitope(s). The experiments highlighted in Examples 12 and 13 showed that this was indeed true for 28.1. Therefore, it was desired to specifically identify the site(s) that could potentially be targeted for phosphorylation and account for loss of reactivity for CBTAU-28.1. Because 47.1, 47.2, and 49.1 bound to the same region on tau as CBTAU-28.1 (i.e., 52-71), it was decided to test these mAbs as well in these experiments. In addition, the same exercise was carried out for CBTAU-43.1 and CBTAU-27.1 as they behave similarly to CBTAU-28.1 by IHC.
[0195] Singly and dually phosphorylated-tau peptides were designed to cover all potential phosphorylation sites with regions 52-71 and 299-323 (CBTAU-28.1 and CBTAU-27.1 binding region, respectively). CBTAU-27.1 and CBTAU-43.1 mAbs were tested against the peptides listed on Tables 30 and 32.96-well streptavidin-coated ELISA plates (Pierce) were coated with phosphorylated tau peptides as detailed in Example 9. Purified anti-tau IgGs were diluted to 5 .mu.g/ml in TBS containing 0.25% BSA, and titrated five-fold. Antibody control and secondary antibodies were used as detailed in Example 9. Antibody reactivity at 1 .mu.g/mL was determined by ELISA and scored as no binding (-), weak (-1+), moderate (+), or strong (++). (-) for average of two O.D. 450 nm readings <0.3; (-/+) for >0.5 and <1.0; (+) for >1.0 and <1.5; (++) for >1.5. Results for each antibody are shown in Tables 30-34 and FIGS. 8A-8F.
TABLE-US-00030 TABLE 30 CBTAU-27.1: Peptides for reactivity by ELISA SEQ ID Re- Peptide NO. sult tau 299-369 331 HVPGGGSVQIVYKPVDLSKVTSKCGSL ++ GNIHHKPGGGQVEVKSEKLDFKDRVQ SKIGSLDNITHVPGGGNK tau 299-323 458 HVPGGGSVQIVYKPVDLSKVTSKCG ++ ptau 299-323 503 HVPGGG(pS)VQIVYKPVDLSKVTSKCG ++ p305 ptau 299-323 504 HVPGGGSVQIV(pY)KPVDLSKVTSKCG ++ p310 ptau 299-323 505 HVPGGGSVQIVYKPVDL(pS)KVTSKCG - p316 ptau 299-323 506 HVPGGGSVQIVYKPVDLSKV(pT)SKCG ++ p319 ptau 299-323 507 HVPGGGSVQIVYKPVDLSKVT(pS)KCG ++ p320 ptau 299-323 508 HVPGGG(pS)VQIV(pY)KPVDLSKVTSKC ++ p305, 310 G ptau 299-323 509 HVPGGG(pS)VQIVYKPVDL(pS)KVTSKC - p305, 316 G ptau 299-323 510 HVPGGG(pS)VQIVYKPVDLSKV(pT)SKC ++ p305, 320 G ptau 299-323 511 HVPGGG(pS)VQIVYKPVDLSKVT(pS)KC ++ p305, 321 G ptau 299-323 512 HVPGGGSVQIV(pY)KPVDL(pS)KVTSKC - p310, 316 G ptau 299-323 513 HVPGGGSVQIV(pY)KPVDLSKV(pT)SKC ++ p310, 320 G ptau 299-323 514 HVPGGGSVQIV(pY)KPVDLSKVT(pS)KC ++ p310, 321 G ptau 299-323 515 HVPGGGSVQIVYKPVDL(pS)KV(pT)SKC - p316, 320 G ptau 299-323 516 HVPGGGSVQIVYKPVDL(pS)KVT(pS)KC - p316, 321 G ptau 299-323 517 HVPGGGSVQIVYKPVDLSKV(pT)(pS)KC -/+ p320, 321 G
TABLE-US-00031 TABLE 31 CBTAU-28.1: Peptides for reactivity by ELISA SEQ ID Re- Peptide NO. sult tau 42-103 325 GLKESPLQTPTEDGSEEPGSETSDAKST ++ PTAEDVTAPLVDEGAPGKQAAAQPHT EIPEGTTA tau 48-71 518 LQTPTEDGSEEPGSETSDAKSTPT ++ ptau 48-71 519 LQTPTEDGSEEPGSETSDAKSTP(pT) ++ (p71) ptau 48-71 520 LQTPTEDGSEEPGSE(pT)SDAKSTPT - (p63) ptau 48-71 521 LQTPTEDGSEEPG(pS)ETSDAKSTPT - (p61) ptau 48-71 522 LQTPTEDG(pS)EEPGSETSDAKSTPT ++ (p56) ptau 48-71 523 LQTP(pT)EDGSEEPGSETSDAKSTPT ++ (p52) ptau 48-71 524 LQTPTEDGSEEPGSETSDAK(pS)TPT ++ (p68) ptau 48-71 525 LQTPTEDGSEEPGSETSDAKS(pT)PT ++ (p69) ptau 48-71 526 LQTPTEDGSEEPGSET(pS)DAKSTPT ++ (p64) ptau 48-71 527 LQTPTEDGSEEPG(pS)ET(pS)DAKSTPT - (p61, p64) ptau 48-71 528 LQTPTEDGSEEPG(pS)E(pT)SDAKSTPT - (p61, p63) ptau 48-71 529 LQTPTEDGSEEPGSE(pT)(pS)DAKSTPT - (p63, p64)
TABLE-US-00032 TABLE 32 CBTAU-43.1: Peptides for reactivity by ELISA SEQ ID Re- Peptide NO. sult tau 299-369 331 HVPGGGSVQIVYKPVDLSKVTSKCGSL ++ GNIHHKPGGGQVEVKSEKIDFKDRVQ SKIGSLDNITHVPGGGNK tau 299-323 458 HVPGGGSVQIVYKPVDLSKVTSKCG ++ ptau 299-323 503 HVPGGG(pS)VQIVYKPVDLSKVTSKCG ++ p305 ptau 299-323 504 HVPGGGSVQIV(pY)KPVDLSKVTSKCG ++ p310 ptau 299-323 505 HVPGGGSVQIVYKPVDL(pS)KVTSKCG - p316 ptau 299-323 506 HVPGGGSVQIVYKPVDLSKV(pT)SKCG ++ p319 ptau 299-323 507 HVPGGGSVQIVYKPVDLSKVT(pS)KCG ++ p320 ptau 299-323 508 HVPGGG(pS)VQIV(pY)KPVDLSKVTSKC ++ p305, 310 G ptau 299-323 509 HVPGGG(pS)VQIVYKPVDL(pS)KVTSKC - p305, 316 G ptau 299-323 510 HVPGGG(pS)VQIVYKPVDLSKV(pT)SKC ++ p305, 320 G ptau 299-323 511 HVPGGG(pS)VQIVYKPVDLSKVT(pS)KC ++ p305, 321 G ptau 299-323 512 HVPGGGSVQIV(pY)KPVDL(pS)KVTSKC - p310, 316 G ptau 299-323 513 HVPGGGSVQIV(pY)KPVDLSKV(pT)SKC ++ p310, 320 G ptau 299-323 514 HVPGGGSVQIV(pY)KPVDLSKVT(pS)KC ++ p310, 321 G ptau 299-323 515 HVPGGGSVQIVYKPVDL(pS)KV(pT)SKC -/+ p316, 320 G ptau 299-323 516 HVPGGGSVQIVYKPVDL(pS)KVT(pS)KC - p316, 321 G ptau 299-323 517 HVPGGGSVQIVYKPVDLSKV(pT)(pS)KC ++ p320, 321 G
TABLE-US-00033 TABLE 33 CBTAU-47.1 and CBTAU-47.2: Peptides for reactivity by ELISA SEQ Result Result Pep- ID CBTAU- CBTAU- tide NO. 47.1 47.2 tau 325 GLKESPLQTPTEDGSEEPGSETSDA ++ ++ 42-103 KSTPTAEDVTAPLVDEGAPGKQAA AQPHTIPEGTTA tau 518 LQTPTEDGSEEPGSETSDAKSTPT ++ ++ 48-71 ptau 519 LQTPTEDGSEEPGSETSDAKSTP ++ ++ 48-71 (pT) (p71) ptau 520 LQTPTEDGSEEPGSE(pT)SDAK - - 48-71 STPT (p63) ptau 521 LQTPTEDGSEEPG(pS)ETSDAK - - 48-71 STPT (p61) ptau 522 LQTPTEDG(pS)EEPGSETSDAK ++ ++ 48-71 STPT (p56) ptau 523 LQTP(pT)EDGSEEPGSETSDAK ++ ++ 48-71 STPT (p52) ptau 524 LQTPTEDGSEEPGSETSDAK ++ ++ 48-71 (pS)TPT (p68) ptau 525 LQTPTEDGSEEPGSETSDAKS ++ ++ 48-71 (pT)PT (p69) ptau 526 LQTPTEDGSEEPGSET(pS) ++ ++ 48-71 DAKSTPT (Ser64) ptau 527 LQTPTEDGSEEPG(pS)ET(pS) - - 48-71 DAKSTPT (p61, Ser64) ptau 528 LQTPTEDGSEEPG(pS)E(pT) - - 48-71 SDAKSTPT (p61, p63) ptau 529 LQTPTEDGSEEPGSE(pT)(pS) - - 48-71 DAKSTPT (p63, p64)
TABLE-US-00034 TABLE 34 CBTAU-49.1: Peptides for reactivity by ELISA SEQ ID Re- Peptide NO. sult tau 42-103 325 GLKESPLQTPTEDGSEEPGSETSDA ++ KSTPTAEDVTAPLVDEGAPGKQAA AQPHT EIPEGTTA tau 48-71 518 LQTPTEDGSEEPGSETSDAKSTPT ++ ptau 48-71 519 LQTPTEDGSEEPGSETSDAKSTP(pT) ++ (p71) ptau 48-71 520 LQTPTEDGSEEPGSE(pT)SDAKSTPT - (p63) ptau 48-71 521 LQTPTEDGSEEPG(pS)ETSDAKSTPT - (p61) ptau 48-71 522 LQTPTEDG(pS)EEPGSETSDAKSTPT ++ (p56) ptau 48-71 523 LQTP(pT)EDGSEEPGSETSDAKSTPT ++ (p52) ptau 48-71 524 LQTPTEDGSEEPGSETSDAK(pS)TPT ++ (p68) ptau 48-71 525 LQTPTEDGSEEPGSETSDAKS(pT)PT -/+ (p69) ptau 48-71 526 LQTPTEDGSEEPGSET(pS)DAKSTPT ++ (Ser64) ptau 48-71 527 LQTPTEDGSEEPG(pS)ET(pS)DAKST - (p61, Ser64) PT ptau 48-71 528 LQTPTEDGSEEPG(pS)E(pT)SDAKST - (p61 ,p63) PT ptau 48-71 529 LQTPTEDGSEEPGSE(pT)(pS)DAKST - (p63, p64) PT
[0196] The results for CBTAU-27.1 and CBTAU-43.1 show that phosphorylation at S316 is sufficient to completely inhibit reactivity (Table 30 and Table 32). This suggests that the loss in reactivity to tau on AD tissue sections (Example 11) may be due to phosphorylation at S316, an event that may occur early during the course of the disease. For CBTAU-28.1, 47.1, 47.2, 49.1, phosphorylation at either S61 or T63 is sufficient to completely inhibit reactivity. Taken together, the results for CBTAU-28.1, 47.1, 47.2, and 49.1 suggest that phosphorylation at S61 and/or T63 is a mechanism that accounts for loss of this epitope during the course of the disease.
Example 15
[0197] Generation of Anti-Tau mAb Mouse-Human Chimeras and Human Isotypes
[0198] To test the efficacy of the human anti-tau mAbs in a mouse model of tauopathy, mouse-human antibody chimeras were generated by replacing the human Fc region with a mouse IgG1 Fc. Briefly, the human IgG1 CH1 region was amplified from the pCB-IgG vector using primers SteplHMchim-Fwd and SteplHMchim-Rev (Table 35) to generate a 0.95 kb fragment containing a 5'-XhoI site (frag. 1). Mouse IgG1 CH2 and CH3 domains (Fc region) are amplified from a gene-synthesized construct using primers Step2HMchim-Fwd and Step2HMchim-Rev
[0199] (Table 30) to generate a 0.82 kb fragment (frag. 2). A third fragment (frag. 3) was generated by amplifying the polyA region of the pCB-IgG vector using primers Step3HMchim-Fwd and Step3HMchim-Rev (Table 30), which includes a 3'-DraIII site. The three fragments were linked into a single cassette by overlap extension PCR to generate a 2.3 kb overlap fragment harboring the human CH1 followed by the mouse CH2-CH3 domains. The overlap fragment was subsequently cloned via the XhoI and DraIII sites into the pCB-IgG CBTAU-7.1 vector to generate a mouse-human chimera of CBTAU-7.1, containing the human variable, CH1, hinge and Ck regions followed by the mouse CH2 and CH3 regions. CBTAU-22.1, 24.1, 27.1, 28.1, 47.1, 47.2, 46.1, 49.1, and 43.1 chimeras were then generated by digesting the CBTAU-7.1 chimera construct and pCB-IgG CBTAU human mAb constructs with XhoI and XbaI, and the corresponding fragments were subcloned. Nucleotide sequences for all constructs are verified according to standard techniques known to the skilled artisan. Chimeric antibodies were subsequently expressed and purified as detailed in Example 5 using Protein G agarose instead of Protein A.
TABLE-US-00035 TABLE 35 PRIMERS FOR MOUSE-HUMAN CHIMERIZATION SEQ ID Primer ID DNA SEQUENCE (5'-3') NO: Step1HMchim- TCTCCGCCGGTGAGTCTCGAGGC 530 Fwd Step1HMchim- TGTCCCTGGATGCAGGCTACTCTAGG 531 Rev Step2HMchim- AGAGTAGCCTGCATCCAGGGACAG 532 Fwd Step2HMchim- TCTAGATCATTTACCAGGAGAGTGGGAGAG 533 Rev Step3HMchim- TCTCCTGGTAAATGATCTAGAGTTTAAACC 534 Fwd GCTG Step3HMchim- ATGGCCCACTACGTGAACCATCACC 535 Rev
Example 16
[0200] Preparation of IgG2, 3 and 4 Isotypes
[0201] As mentioned in Example 3, all CBTAU mAbs were cloned and expressed as chimeric human IgG1 regardless of their native isotype. To generate additional human isotype versions (i.e., IgG2/3/4), the CH1 through CH3 region corresponding to each of the human IgG isotypes is PCR amplified from gene-synthesized constructs containing the corresponding constant regions, hinge, and intron sequences. The PCR amplicons contain 5'-XhoI and 3'-DraIII sites, which are used to subclone the fragments into the corresponding pCB-IgG CBTAU antibody construct. In this manner, human IgG2, 3 and 4 isotype versions are generated for each of the anti-tau mAbs.
Example 17
[0202] Generation of De-Risked and Fc-Engineered Anti-Tau Chimeric Monoclonal Antibody Variants
[0203] The heavy and light chain variable regions (VH and VL) for each anti-tau antibody clone isolated in Example 3 are analyzed for the presence of free cysteines and potential post-translational modification sites, which include glycosylation, deamidation and oxidation sites. Amino acid mutations consisting of structurally conserved and/or germline-based substitutions are used to change these sites. Non-conserved cysteines in the variable regions are mutated to serine. For glycosylation sites, several mutations are used, including replacement of asparagine for the conservative glutamine or germline mutations. Modifications to the deamidation sites include replacement of aspartic acid for asparagine and serine or alanine for glycine. Sites of potential oxidation are not modified. To increase the binding affinity to FcRn and thus increase the half-life of IgG1 mAbs in vivo, several mutations located at the boundary between the CH2 and CH3 regions are generated. These mutations included M252Y/S254T/T256E plus H433K/N434F (C. Vaccaro et al., 2005) or T250Q/M428L (P. R. Hinton et al., 2004), which have been shown to increase IgG1 binding to FcRn. All substitutions are generated by site-directed mutagenesis per manufacturer's instructions (QuicKCHANGE.TM. II, Agilent Technologies, cat. no. 200521). Nucleotide sequences for all constructs are verified according to standard techniques known to the skilled artisan.
Sequence CWU 1
1
5351441PRTHomo sapiens 1Met Ala Glu Pro Arg Gln Glu Phe Glu Val Met
Glu Asp His Ala Gly1 5 10 15Thr Tyr Gly Leu Gly Asp Arg Lys Asp Gln
Gly Gly Tyr Thr Met His 20 25 30Gln Asp Gln Glu Gly Asp Thr Asp Ala
Gly Leu Lys Glu Ser Pro Leu 35 40 45Gln Thr Pro Thr Glu Asp Gly Ser
Glu Glu Pro Gly Ser Glu Thr Ser 50 55 60Asp Ala Lys Ser Thr Pro Thr
Ala Glu Asp Val Thr Ala Pro Leu Val65 70 75 80Asp Glu Gly Ala Pro
Gly Lys Gln Ala Ala Ala Gln Pro His Thr Glu 85 90 95Ile Pro Glu Gly
Thr Thr Ala Glu Glu Ala Gly Ile Gly Asp Thr Pro 100 105 110Ser Leu
Glu Asp Glu Ala Ala Gly His Val Thr Gln Ala Arg Met Val 115 120
125Ser Lys Ser Lys Asp Gly Thr Gly Ser Asp Asp Lys Lys Ala Lys Gly
130 135 140Ala Asp Gly Lys Thr Lys Ile Ala Thr Pro Arg Gly Ala Ala
Pro Pro145 150 155 160Gly Gln Lys Gly Gln Ala Asn Ala Thr Arg Ile
Pro Ala Lys Thr Pro 165 170 175Pro Ala Pro Lys Thr Pro Pro Ser Ser
Gly Glu Pro Pro Lys Ser Gly 180 185 190Asp Arg Ser Gly Tyr Ser Ser
Pro Gly Ser Pro Gly Thr Pro Gly Ser 195 200 205Arg Ser Arg Thr Pro
Ser Leu Pro Thr Pro Pro Thr Arg Glu Pro Lys 210 215 220Lys Val Ala
Val Val Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala Lys225 230 235
240Ser Arg Leu Gln Thr Ala Pro Val Pro Met Pro Asp Leu Lys Asn Val
245 250 255Lys Ser Lys Ile Gly Ser Thr Glu Asn Leu Lys His Gln Pro
Gly Gly 260 265 270Gly Lys Val Gln Ile Ile Asn Lys Lys Leu Asp Leu
Ser Asn Val Gln 275 280 285Ser Lys Cys Gly Ser Lys Asp Asn Ile Lys
His Val Pro Gly Gly Gly 290 295 300Ser Val Gln Ile Val Tyr Lys Pro
Val Asp Leu Ser Lys Val Thr Ser305 310 315 320Lys Cys Gly Ser Leu
Gly Asn Ile His His Lys Pro Gly Gly Gly Gln 325 330 335Val Glu Val
Lys Ser Glu Lys Leu Asp Phe Lys Asp Arg Val Gln Ser 340 345 350Lys
Ile Gly Ser Leu Asp Asn Ile Thr His Val Pro Gly Gly Gly Asn 355 360
365Lys Lys Ile Glu Thr His Lys Leu Thr Phe Arg Glu Asn Ala Lys Ala
370 375 380Lys Thr Asp His Gly Ala Glu Ile Val Tyr Lys Ser Pro Val
Val Ser385 390 395 400Gly Asp Thr Ser Pro Arg His Leu Ser Asn Val
Ser Ser Thr Gly Ser 405 410 415Ile Asp Met Val Asp Ser Pro Gln Leu
Ala Thr Leu Ala Asp Glu Val 420 425 430Ser Ala Ser Leu Ala Lys Gln
Gly Leu 435 4402410PRTHomo sapiens 2Met Ala Glu Pro Arg Gln Glu Phe
Glu Val Met Glu Asp His Ala Gly1 5 10 15Thr Tyr Gly Leu Gly Asp Arg
Lys Asp Gln Gly Gly Tyr Thr Met His 20 25 30Gln Asp Gln Glu Gly Asp
Thr Asp Ala Gly Leu Lys Glu Ser Pro Leu 35 40 45Gln Thr Pro Thr Glu
Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr Ser 50 55 60Asp Ala Lys Ser
Thr Pro Thr Ala Glu Asp Val Thr Ala Pro Leu Val65 70 75 80Asp Glu
Gly Ala Pro Gly Lys Gln Ala Ala Ala Gln Pro His Thr Glu 85 90 95Ile
Pro Glu Gly Thr Thr Ala Glu Glu Ala Gly Ile Gly Asp Thr Pro 100 105
110Ser Leu Glu Asp Glu Ala Ala Gly His Val Thr Gln Ala Arg Met Val
115 120 125Ser Lys Ser Lys Asp Gly Thr Gly Ser Asp Asp Lys Lys Ala
Lys Gly 130 135 140Ala Asp Gly Lys Thr Lys Ile Ala Thr Pro Arg Gly
Ala Ala Pro Pro145 150 155 160Gly Gln Lys Gly Gln Ala Asn Ala Thr
Arg Ile Pro Ala Lys Thr Pro 165 170 175Pro Ala Pro Lys Thr Pro Pro
Ser Ser Gly Glu Pro Pro Lys Ser Gly 180 185 190Asp Arg Ser Gly Tyr
Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser 195 200 205Arg Ser Arg
Thr Pro Ser Leu Pro Thr Pro Pro Thr Arg Glu Pro Lys 210 215 220Lys
Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala Lys225 230
235 240Ser Arg Leu Gln Thr Ala Pro Val Pro Met Pro Asp Leu Lys Asn
Val 245 250 255Lys Ser Lys Ile Gly Ser Thr Glu Asn Leu Lys His Gln
Pro Gly Gly 260 265 270Gly Lys Val Gln Ile Val Tyr Lys Pro Val Asp
Leu Ser Lys Val Thr 275 280 285Ser Lys Cys Gly Ser Leu Gly Asn Ile
His His Lys Pro Gly Gly Gly 290 295 300Gln Val Glu Val Lys Ser Glu
Lys Leu Asp Phe Lys Asp Arg Val Gln305 310 315 320Ser Lys Ile Gly
Ser Leu Asp Asn Ile Thr His Val Pro Gly Gly Gly 325 330 335Asn Lys
Lys Ile Glu Thr His Lys Leu Thr Phe Arg Glu Asn Ala Lys 340 345
350Ala Lys Thr Asp His Gly Ala Glu Ile Val Tyr Lys Ser Pro Val Val
355 360 365Ser Gly Asp Thr Ser Pro Arg His Leu Ser Asn Val Ser Ser
Thr Gly 370 375 380Ser Ile Asp Met Val Asp Ser Pro Gln Leu Ala Thr
Leu Ala Asp Glu385 390 395 400Val Ser Ala Ser Leu Ala Lys Gln Gly
Leu 405 4103412PRTHomo sapiens 3Met Ala Glu Pro Arg Gln Glu Phe Glu
Val Met Glu Asp His Ala Gly1 5 10 15Thr Tyr Gly Leu Gly Asp Arg Lys
Asp Gln Gly Gly Tyr Thr Met His 20 25 30Gln Asp Gln Glu Gly Asp Thr
Asp Ala Gly Leu Lys Glu Ser Pro Leu 35 40 45Gln Thr Pro Thr Glu Asp
Gly Ser Glu Glu Pro Gly Ser Glu Thr Ser 50 55 60Asp Ala Lys Ser Thr
Pro Thr Ala Glu Ala Glu Glu Ala Gly Ile Gly65 70 75 80Asp Thr Pro
Ser Leu Glu Asp Glu Ala Ala Gly His Val Thr Gln Ala 85 90 95Arg Met
Val Ser Lys Ser Lys Asp Gly Thr Gly Ser Asp Asp Lys Lys 100 105
110Ala Lys Gly Ala Asp Gly Lys Thr Lys Ile Ala Thr Pro Arg Gly Ala
115 120 125Ala Pro Pro Gly Gln Lys Gly Gln Ala Asn Ala Thr Arg Ile
Pro Ala 130 135 140Lys Thr Pro Pro Ala Pro Lys Thr Pro Pro Ser Ser
Gly Glu Pro Pro145 150 155 160Lys Ser Gly Asp Arg Ser Gly Tyr Ser
Ser Pro Gly Ser Pro Gly Thr 165 170 175Pro Gly Ser Arg Ser Arg Thr
Pro Ser Leu Pro Thr Pro Pro Thr Arg 180 185 190Glu Pro Lys Lys Val
Ala Val Val Arg Thr Pro Pro Lys Ser Pro Ser 195 200 205Ser Ala Lys
Ser Arg Leu Gln Thr Ala Pro Val Pro Met Pro Asp Leu 210 215 220Lys
Asn Val Lys Ser Lys Ile Gly Ser Thr Glu Asn Leu Lys His Gln225 230
235 240Pro Gly Gly Gly Lys Val Gln Ile Ile Asn Lys Lys Leu Asp Leu
Ser 245 250 255Asn Val Gln Ser Lys Cys Gly Ser Lys Asp Asn Ile Lys
His Val Pro 260 265 270Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro
Val Asp Leu Ser Lys 275 280 285Val Thr Ser Lys Cys Gly Ser Leu Gly
Asn Ile His His Lys Pro Gly 290 295 300Gly Gly Gln Val Glu Val Lys
Ser Glu Lys Leu Asp Phe Lys Asp Arg305 310 315 320Val Gln Ser Lys
Ile Gly Ser Leu Asp Asn Ile Thr His Val Pro Gly 325 330 335Gly Gly
Asn Lys Lys Ile Glu Thr His Lys Leu Thr Phe Arg Glu Asn 340 345
350Ala Lys Ala Lys Thr Asp His Gly Ala Glu Ile Val Tyr Lys Ser Pro
355 360 365Val Val Ser Gly Asp Thr Ser Pro Arg His Leu Ser Asn Val
Ser Ser 370 375 380Thr Gly Ser Ile Asp Met Val Asp Ser Pro Gln Leu
Ala Thr Leu Ala385 390 395 400Asp Glu Val Ser Ala Ser Leu Ala Lys
Gln Gly Leu 405 4104383PRTHomo sapiens 4Thr Ala Met Ala Glu Pro Arg
Gln Glu Phe Glu Val Met Glu Asp His1 5 10 15Ala Gly Thr Tyr Gly Leu
Gly Asp Arg Lys Asp Gln Gly Gly Tyr Thr 20 25 30Met His Gln Asp Gln
Glu Gly Asp Thr Asp Ala Gly Leu Lys Glu Ser 35 40 45Pro Leu Gln Thr
Pro Thr Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu 50 55 60Thr Ser Asp
Ala Lys Ser Thr Pro Thr Ala Glu Ala Glu Glu Ala Gly65 70 75 80Ile
Gly Asp Thr Pro Ser Leu Glu Asp Glu Ala Ala Gly His Val Thr 85 90
95Gln Ala Arg Met Val Ser Lys Ser Lys Asp Gly Thr Gly Ser Asp Asp
100 105 110Lys Lys Ala Lys Gly Ala Asp Gly Lys Thr Lys Ile Ala Thr
Pro Arg 115 120 125Gly Ala Ala Pro Pro Gly Gln Lys Gly Gln Ala Asn
Ala Thr Arg Ile 130 135 140Pro Ala Lys Thr Pro Pro Ala Pro Lys Thr
Pro Pro Ser Ser Gly Glu145 150 155 160Pro Pro Lys Ser Gly Asp Arg
Ser Gly Tyr Ser Ser Pro Gly Ser Pro 165 170 175Gly Thr Pro Gly Ser
Arg Ser Arg Thr Pro Ser Leu Pro Thr Pro Pro 180 185 190Thr Arg Glu
Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser 195 200 205Pro
Ser Ser Ala Lys Ser Arg Leu Gln Thr Ala Pro Val Pro Met Pro 210 215
220Asp Leu Lys Asn Val Lys Ser Lys Ile Gly Ser Thr Glu Asn Leu
Lys225 230 235 240His Gln Pro Gly Gly Gly Lys Val Gln Ile Val Tyr
Lys Pro Val Asp 245 250 255Leu Ser Lys Val Thr Ser Lys Cys Gly Ser
Leu Gly Asn Ile His His 260 265 270Lys Pro Gly Gly Gly Gln Val Glu
Val Lys Ser Glu Lys Leu Asp Phe 275 280 285Lys Asp Arg Val Gln Ser
Lys Ile Gly Ser Leu Asp Asn Ile Thr His 290 295 300Val Pro Gly Gly
Gly Asn Lys Lys Ile Glu Thr His Lys Leu Thr Phe305 310 315 320Arg
Glu Asn Ala Lys Ala Lys Thr Asp His Gly Ala Glu Ile Val Tyr 325 330
335Lys Ser Pro Val Val Ser Gly Asp Thr Ser Pro Arg His Leu Ser Asn
340 345 350Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp Ser Pro Gln
Leu Ala 355 360 365Thr Leu Ala Asp Glu Val Ser Ala Ser Leu Ala Lys
Gln Gly Leu 370 375 3805385PRTHomo sapiens 5Thr Ala Met Ala Glu Pro
Arg Gln Glu Phe Glu Val Met Glu Asp His1 5 10 15Ala Gly Thr Tyr Gly
Leu Gly Asp Arg Lys Asp Gln Gly Gly Tyr Thr 20 25 30Met His Gln Asp
Gln Glu Gly Asp Thr Asp Ala Gly Leu Lys Ala Glu 35 40 45Glu Ala Gly
Ile Gly Asp Thr Pro Ser Leu Glu Asp Glu Ala Ala Gly 50 55 60His Val
Thr Gln Ala Arg Met Val Ser Lys Ser Lys Asp Gly Thr Gly65 70 75
80Ser Asp Asp Lys Lys Ala Lys Gly Ala Asp Gly Lys Thr Lys Ile Ala
85 90 95Thr Pro Arg Gly Ala Ala Pro Pro Gly Gln Lys Gly Gln Ala Asn
Ala 100 105 110Thr Arg Ile Pro Ala Lys Thr Pro Pro Ala Pro Lys Thr
Pro Pro Ser 115 120 125Ser Gly Glu Pro Pro Lys Ser Gly Asp Arg Ser
Gly Tyr Ser Ser Pro 130 135 140Gly Ser Pro Gly Thr Pro Gly Ser Arg
Ser Arg Thr Pro Ser Leu Pro145 150 155 160Thr Pro Pro Thr Arg Glu
Pro Lys Lys Val Ala Val Val Arg Thr Pro 165 170 175Pro Lys Ser Pro
Ser Ser Ala Lys Ser Arg Leu Gln Thr Ala Pro Val 180 185 190Pro Met
Pro Asp Leu Lys Asn Val Lys Ser Lys Ile Gly Ser Thr Glu 195 200
205Asn Leu Lys His Gln Pro Gly Gly Gly Lys Val Gln Ile Ile Asn Lys
210 215 220Lys Leu Asp Leu Ser Asn Val Gln Ser Lys Cys Gly Ser Lys
Asp Asn225 230 235 240Ile Lys His Val Pro Gly Gly Gly Ser Val Gln
Ile Val Tyr Lys Pro 245 250 255Val Asp Leu Ser Lys Val Thr Ser Lys
Cys Gly Ser Leu Gly Asn Ile 260 265 270His His Lys Pro Gly Gly Gly
Gln Val Glu Val Lys Ser Glu Lys Leu 275 280 285Asp Phe Lys Asp Arg
Val Gln Ser Lys Ile Gly Ser Leu Asp Asn Ile 290 295 300Thr His Val
Pro Gly Gly Gly Asn Lys Lys Ile Glu Thr His Lys Leu305 310 315
320Thr Phe Arg Glu Asn Ala Lys Ala Lys Thr Asp His Gly Ala Glu Ile
325 330 335Val Tyr Lys Ser Pro Val Val Ser Gly Asp Thr Ser Pro Arg
His Leu 340 345 350Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val
Asp Ser Pro Gln 355 360 365Leu Ala Thr Leu Ala Asp Glu Val Ser Ala
Ser Leu Ala Lys Gln Gly 370 375 380Leu3856352PRTHomo sapiens 6Met
Ala Glu Pro Arg Gln Glu Phe Glu Val Met Glu Asp His Ala Gly1 5 10
15Thr Tyr Gly Leu Gly Asp Arg Lys Asp Gln Gly Gly Tyr Thr Met His
20 25 30Gln Asp Gln Glu Gly Asp Thr Asp Ala Gly Leu Lys Ala Glu Glu
Ala 35 40 45Gly Ile Gly Asp Thr Pro Ser Leu Glu Asp Glu Ala Ala Gly
His Val 50 55 60Thr Gln Ala Arg Met Val Ser Lys Ser Lys Asp Gly Thr
Gly Ser Asp65 70 75 80Asp Lys Lys Ala Lys Gly Ala Asp Gly Lys Thr
Lys Ile Ala Thr Pro 85 90 95Arg Gly Ala Ala Pro Pro Gly Gln Lys Gly
Gln Ala Asn Ala Thr Arg 100 105 110Ile Pro Ala Lys Thr Pro Pro Ala
Pro Lys Thr Pro Pro Ser Ser Gly 115 120 125Glu Pro Pro Lys Ser Gly
Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser 130 135 140Pro Gly Thr Pro
Gly Ser Arg Ser Arg Thr Pro Ser Leu Pro Thr Pro145 150 155 160Pro
Thr Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys 165 170
175Ser Pro Ser Ser Ala Lys Ser Arg Leu Gln Thr Ala Pro Val Pro Met
180 185 190Pro Asp Leu Lys Asn Val Lys Ser Lys Ile Gly Ser Thr Glu
Asn Leu 195 200 205Lys His Gln Pro Gly Gly Gly Lys Val Gln Ile Val
Tyr Lys Pro Val 210 215 220Asp Leu Ser Lys Val Thr Ser Lys Cys Gly
Ser Leu Gly Asn Ile His225 230 235 240His Lys Pro Gly Gly Gly Gln
Val Glu Val Lys Ser Glu Lys Leu Asp 245 250 255Phe Lys Asp Arg Val
Gln Ser Lys Ile Gly Ser Leu Asp Asn Ile Thr 260 265 270His Val Pro
Gly Gly Gly Asn Lys Lys Ile Glu Thr His Lys Leu Thr 275 280 285Phe
Arg Glu Asn Ala Lys Ala Lys Thr Asp His Gly Ala Glu Ile Val 290 295
300Tyr Lys Ser Pro Val Val Ser Gly Asp Thr Ser Pro Arg His Leu
Ser305 310 315 320Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp
Ser Pro Gln Leu 325 330 335Ala Thr Leu Ala Asp Glu Val Ser Ala Ser
Leu Ala Lys Gln Gly Leu 340 345 350724DNAartificialsynthetic
construct 7atggactgga cctggaggtt cctc 24824DNAartificialsynthetic
construct 8atggactgga cctggaggat cctc 24924DNAartificialsynthetic
construct 9atggactgga cctggagggt cttc 241022DNAUnknownsynthetic
construct 10atggactgga cctggagcat cc 221126DNAartificialsynthetic
construct 11ggacatactt tgttccacgc tcctgc
261223DNAartificialsynthetic construct 12aggtgtccag tgtcaggtgc agc
231323DNAartificialsynthetic construct 13aggtgtccag tgtgaggtgc agc
231423DNAartificialsynthetic construct 14aggtgtccag tgtcaggtac agc
231521DNAartificialsynthetic construct 15gcagctccca gatgggtcct g
211621DNAartificialsynthetic construct 16tcaaccgcca tcctcgccct c
211726DNAartificialsynthetic construct 17gtctgtctcc ttcctcatct
tcctgc 261823DNAartificialsynthetic construct 18ggaaggtgtg
cacgccgctg gtc 231921DNAartificialsynthetic construct 19atgagggtcc
ccgctcagct c 212021DNAartificialsynthetic construct 20atgagggtcc
ctgctcagct c 212121DNAartificialsynthetic construct 21atgagagtcc
tcgctcagct c 212220DNAartificialsynthetic construct 22tggggctgct
aatgctctgg 202323DNAartificialsynthetic construct 23cctcctgcta
ctctggctcc cag 232426DNAartificialsynthetic construct 24tctctgttgc
tctggatctc tggtgc 262524DNAartificialsynthetic construct
25ctcctcagct tcctcctcct ttgg 242626DNAartificialsynthetic construct
26aactcattgg gtttctgctg ctctgg 262726DNAartificialsynthetic
construct 27gtttctcgta gtctgctttg ctcagc
262823DNAartificialsynthetic construct 28gtgctgtcct tgctgtcctg ctc
232926DNAartificialsynthetic construct 29gcactctccc ctgttgaagc
tctttg 263020DNAartificialsynthetic construct 30ctcctcgctc
actgcacagg 203120DNAartificialsynthetic construct 31ctcctctctc
actgcacagg 203220DNAartificialsynthetic construct 32ctcctcactc
gggacacagg 203320DNAartificialsynthetic construct 33atggcctgga
cccctctctg 203424DNAartificialsynthetic construct 34atggcatgga
tccctctctt cctc 243524DNAartificialsynthetic construct 35cactagtgtg
gccttgttgg cttg 243643DNAartificialsynthetic construct 36cctgtctgga
attcagcatg gcccaggtgc agctggtgca gtc 433743DNAartificialsynthetic
construct 37cctgtctgga attcagcatg gcccaggtcc agctggtgca gtc
433843DNAartificialsynthetic construct 38cctgtctgga attcagcatg
gcccaggttc agctggtgca gtc 433943DNAartificialsynthetic construct
39cctgtctgga attcagcatg gcccaggtcc agcttgtgca gtc
434046DNAartificialsynthetic construct 40cctgtctgga attcagcatg
gcccaggtca ccttgaggga gtctgg 464146DNAartificialsynthetic construct
41cctgtctgga attcagcatg gcccaggtca ccttgaagga gtctgg
464243DNAartificialsynthetic construct 42cctgtctgga attcagcatg
gcccaggtgc agctggtgga gtc 434343DNAartificialsynthetic construct
43cctgtctgga attcagcatg gccgaggtgc agctgttgga gtc
434443DNAartificialsynthetic construct 44cctgtctgga attcagcatg
gccgaggtgc agctggtgga gtc 434545DNAartificialsynthetic construct
45cctgtctgga attcagcatg gcccaggtac agctggtgga gtctg
454641DNAartificialsynthetic construct 46cctgtctgga attcagcatg
gcccagstgc agctgcagga g 414744DNAartificialsynthetic construct
47cctgtctgga attcagcatg gcccaggtgc agctacagca gtgg
444843DNAartificialsynthetic construct 48cctgtctgga attcagcatg
gccgaggtgc agctggtgca gtc 434945DNAartificialsynthetic construct
49cctgtctgga attcagcatg gcccaggtac agctgcagca gtcag
455045DNAartificialsynthetic construct 50cctgtctgga attcagcatg
gcccaggtgc agctggtgca atctg 455137DNAartificialsynthetic construct
51tcgggcctcg agactcacct gaggagacgg tgaccag
375239DNAartificialsynthetic construct 52tcgggcctcg agactcacct
gaagagacgg tgaccattg 395338DNAartificialsynthetic construct
53tcgggcctcg agactcacct gaggagacgg tgaccgtg
385446DNAartificialsynthetic construct 54ccggtctaga gttttccatg
gcggacatcc agatgaccca gtctcc 465546DNAartificialsynthetic construct
55ccggtctaga gttttccatg gcggacatcc agttgaccca gtctcc
465646DNAartificialsynthetic construct 56ccggtctaga gttttccatg
gcggccatcc agttgaccca gtctcc 465750DNAartificialsynthetic
constructmisc_feature(7)..(7)A or Gmisc_feature(7)..(7)a or g
57ccggtctaga gttttccatg gcggatrttg tgatgactca gtctccactc
505848DNAartificialsynthetic construct 58ccggtctaga gttttccatg
gcggaaattg tgttgacgca gtctccag 485948DNAartificialsynthetic
construct 59ccggtctaga gttttccatg gcggaaattg tgttgacaca gtctccag
486048DNAartificialsynthetic construct 60ccggtctaga gttttccatg
gcggaaatag tgatgacgca gtctccag 486146DNAartificialsynthetic
construct 61ccggtctaga gttttccatg gcggacatcg tgatgaccca gtctcc
466246DNAartificialsynthetic construct 62ccggtctaga gttttccatg
gcggaaacga cactcacgca gtctcc 466348DNAartificialsynthetic construct
63ccggtctaga gttttccatg gcggaaattg tgctgactca gtctccag
486443DNAartificialsynthetic construct 64cgcaaagtgc acttacgttt
gatttccacc ttggtccctt ggc 436543DNAartificialsynthetic construct
65cgcaaagtgc acttacgttt gatctccagc ttggtcccct ggc
436643DNAartificialsynthetic construct 66cgcaaagtgc acttacgttt
gatatccact ttggtcccag ggc 436743DNAartificialsynthetic construct
67cgcaaagtgc acttacgttt gatctccacc ttggtccctc cgc
436843DNAartificialsynthetic construct 68cgcaaagtgc acttacgttt
aatctccagt cgtgtccctt ggc 436949DNAartificialsynthetic construct
69ccggtctaga gttttccatg gcgaatttta tgctgactca gccccactc
497043DNAartificialsynthetic construct 70ccggtctaga gttttccatg
gcgtcctatg tgctgactca gcc 437143DNAartificialsynthetic construct
71ccggtctaga gttttccatg gcgcagtctg tgctgacgca gcc
437243DNAartificialsynthetic construct 72ccggtctaga gttttccatg
gcgcagtctg tcgtgacgca gcc 437343DNAartificialsynthetic construct
73ccggtctaga gttttccatg gcgcagtctg ccctgactca gcc
437445DNAartificialsynthetic construct 74ccggtctaga gttttccatg
gcgtcttctg agctgactca ggacc 457546DNAartificialsynthetic construct
75ccggtctaga gttttccatg gcgtcctatg agctgactca gccacc
467627DNAartificialsynthetic constructmisc_feature(13)..(13)c or t
76ctcagaggag ggygggaaca gagtgac 27771766DNAartificialsynthetic
construct 77aaacgtaagt gcactttgcg gccgctagga agaaactcaa aacatcaaga
ttttaaatac 60gcttcttggt ctccttgcta taattatctg ggataagcat gctgttttct
gtctgtccct 120aacatgccct gtgattatcc gcaaacaaca cacccaaggg
cagaactttg ttacttaaac 180accatcctgt ttgcttcttt cctcaggaac
tgtggctgca ccatctgtct tcatcttccc 240gccatctgat gagcagttga
aatctggaac tgcctctgtt gtgtgcctgc tgaataactt 300ctatcccaga
gaggccaaag tacagtggaa ggtggataac gccctccaat cgggtaactc
360ccaggagagt gtcacagagc aggacagcaa ggacagcacc tacagcctca
gcagcaccct 420gacgctgagc aaagcagact acgagaaaca caaagtctac
gcctgcgaag tcacccatca 480gggcctgagc tcgcccgtca caaagagctt
caacagggga gagtgttagt taacggatcg 540atccgagctc ggtaccaagc
ttaagtttaa accgctgatc agcctcgact gtgccttcta 600gttgccagcc
atctgttgtt tgcccctccc ccgtgccttc cttgaccctg gaaggtgcca
660ctcccactgt cctttcctaa taaaatgagg aaattgcatc gcattgtctg
agtaggtgtc 720attctattct ggggggtggg gtggggcagg acagcaaggg
ggaggattgg gaagacaata 780gcaggcatgc tggggatgcg gtaatctgct
tagggttagg cgttttgcgc tgcttcgcta 840ggtggtcaat attggccatt
agccatatta ttcattggtt atatagcata aatcaatatt 900ggctattggc
cattgcatac gttgtatcca tatcataata tgtacattta tattggctca
960tgtccaacat taccgccatg ttgacattga ttattgacta gttattaata
gtaatcaatt 1020acggggtcat tagttcatag cccatatatg gagttccgcg
ttacataact tacggtaaat 1080ggcccgcctg gctgaccgcc caacgacccc
cgcccattga cgtcaataat gacgtatgtt 1140cccatagtaa cgccaatagg
gactttccat tgacgtcaat gggtggagta tttacggtaa 1200actgcccact
tggcagtaca tcaagtgtat catatgccaa gtacgccccc tattgacgtc
1260aatgacggta aatggcccgc ctggcattat gcccagtaca tgaccttatg
ggactttcct 1320acttggcagt acatctacgt attagtcatc gctattacca
tggtgatgcg gttttggcag 1380tacatcaatg ggcgtggata gcggtttgac
tcacggggat ttccaagtct ccaccccatt 1440gacgtcaatg ggagtttgtt
ttggcaccaa aatcaacggg actttccaaa atgtcgtaac 1500aactccgccc
cattgacgca aatgggcggt aggcgtgtac ggtgggaggt ctatataagc
1560agagctcgtt tagtgaaccg tcagatcgcc tggagacgcc atccacgctg
ttttgacctc 1620catagaagac accgggaccg atccagcctc cgcggccggg
aacggtgcat tggaagcttg 1680gtaccgagct cggatccgcc accatggcct
gccccggatt tctgtgggcc ctggtcatca 1740gcacctgtct ggaattcagc atggcc
17667823DNAartificialsynthetic construct 78ggccatgctg aattccagac
agg 237929DNAartificialsynthetic construct 79aaacgtaagt gcactttgcg
gccgctagg 298025DNAartificialsynthetic construct 80actctgttcc
crccctcctc tgagg 258123DNAartificialsynthetic construct
81ccggtctaga gttttccatg gcg 238219DNAartificialsynthetic construct
82tcgggcctcg agactcacc 1983330PRTHomo sapiens 83Ala Ser Thr Lys Gly
Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1 5 10 15Ser Thr Ser Gly
Gly Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu
Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val
His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu
Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr65 70 75
80Tyr Ile Cys Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys
85 90 95Arg Val Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro
Cys 100 105 110Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu
Phe Pro Pro 115 120 125Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr
Pro Glu Val Thr Cys 130 135 140Val Val Val Asp Val Ser His Glu Asp
Pro Glu Val Lys Phe Asn Trp145 150 155 160Tyr Val Asp Gly Val Glu
Val His Asn Ala Lys Thr Lys Pro Arg Glu 165 170 175Glu Gln Tyr Asn
Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu 180 185 190His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn 195 200
205Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly
210 215 220Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg
Glu Glu225 230 235 240Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr 245 250 255Pro Ser Asp Ile Ala Val Glu Trp Glu
Ser Asn Gly Gln Pro Glu Asn 260 265 270Asn Tyr Lys Thr Thr Pro Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe 275 280 285Leu Tyr Ser Lys Leu
Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn 290 295 300Val Phe Ser
Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr305 310 315
320Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 325 33084107PRTHomo
sapiens 84Arg Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser
Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu
Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu
Ser Lys Ala Asp Tyr Glu65 70 75 80Lys His Lys Val Tyr Ala Cys Glu
Val Thr His Gln Gly Leu Ser Ser 85 90 95Pro Val Thr Lys Ser Phe Asn
Arg Gly Glu Cys 100 1058518DNAartificialsynthetic construct
85catgtcaccg gggtgtgg 188621DNAartificialsynthetic construct
86tcacagggga tgttagggac a 2187117PRTartificialCombination of
PCR-generated and human sequence 87Gln Val Gln Leu Val Glu Ser Gly
Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala
Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Trp Met His Trp Val Arg
Gln Ala Pro Gly Lys Gly Leu Val Trp Val 35 40 45Ser Arg Ile Asn Ser
Asp Gly Ser Asp Thr Asn Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe
Thr Phe Ser Arg Asp Asn Ala Lys Asn Thr Leu Tyr65 70 75 80Leu Gln
Met Thr Ser Leu Arg Ala Glu Asp Thr Ala Ile Tyr Tyr Cys 85 90 95Thr
Arg Gly Arg Ser Tyr Gly Phe Phe Asp Tyr Trp Gly Gln Gly Ala 100 105
110Leu Val Thr Val Ser 11588108PRTartificialCombination of
PCR-generated and human sequence 88Asp Ile Val Met Thr Gln Ser Pro
Asp Thr Leu Ser Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ile Ile Ser Ser Asn 20 25 30Tyr Leu Ala Trp Tyr Gln
Gln Gln Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser
Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60Gly Ser Gly Ser
Ala Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu65 70 75 80Pro Glu
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Gly Thr Ser Pro 85 90 95Arg
Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100
10589354DNAartificialCombination of PCR-generated and human
sequence 89caggtccagc tggtggagtc cgggggaggc ttagttcagc ctggggggtc
cctgagactc 60tcctgtgcag cgtctggatt caccttcagc agctactgga tgcactgggt
ccgccaagct 120ccagggaagg ggctggtgtg ggtctcccgt attaatagtg
atgggagtga cacaaactac 180gcggactccg tgaagggccg attcaccttc
tccagagaca acgccaagaa cacattgtat 240ctgcagatga ccagtctgcg
agccgaggac acggctatat attactgtac aagggggcgc 300agttatggtt
tctttgacta ctggggccag ggagccctgg tcaccgtctc ctca
35490324DNAartificialCombination of PCR-generated and human
sequence 90gacatcgtga tgacccagtc tccagacacc ctgtctttgt ctccagggga
gagagccacc 60ctctcctgca gggccagtca gattattagc agcaactact tagcctggta
ccaacaacaa 120cctggccagg ctcccaggct cctcatctat ggtgcatcca
gcagggccac aggcatccca 180gacaggttca gtggcagtgg gtctgcgaca
gacttcactc tcaccatcag cagactggag 240cctgaagatt ttgcagtgta
ttactgtcag cagtatggta cctcacctcg gacgttcggc 300caggggacca
agctggagat caaa 32491124PRTartificialCombination of PCR-generated
and human sequence 91Gln Val Gln Leu Val Glu Ser Arg Gly Gly Val
Val Gln Pro Gly Thr1 5 10 15Ser Leu Arg Leu Ser Cys Lys Ala Ser Gly
Phe Thr Phe Ser Thr Tyr 20 25 30Gly Met His Trp Val Arg Gln Ala Pro
Gly Lys Ala Leu Glu Trp Val 35 40 45Ala Val Ile Trp Phe Asp Gly Asn
Asn Lys Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser
Arg Asp Asn Ser Lys Asn Thr Val Tyr65 70 75 80Leu Gln Met Asn Ser
Leu Arg Gly Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Trp
Trp Glu Ala Gly Cys Arg Pro Cys Tyr Phe Phe Asp 100 105 110Tyr Trp
Gly Gln Gly Ser Leu Val Thr Val Ser Ser 115
12092113PRTartificialCombination of PCR-generated and human
sequence 92Glu Thr Thr Leu Thr Gln Ser Pro Asp Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val
Leu Tyr Ser 20
25 30Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly
Gln 35 40 45Pro Pro Lys Val Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser
Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val
Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Pro Pro Leu Thr Phe Gly Gly
Gly Thr Lys Leu Glu Ile 100 105 110Lys93372DNAartificialCombination
of PCR-generated and human sequence 93caggtgcagc tggtggagtc
gaggggaggc gtggtccagc ctgggacgtc cctgagactc 60tcctgtaaag cgtctggatt
caccttcagc acttatggca tgcactgggt ccgccaggct 120ccaggcaagg
cgctggagtg ggtggcagtt atatggtttg atggaaataa caaatattat
180gcagactccg tgaagggccg attcaccatc tccagagaca attccaagaa
cacggtgtat 240ctgcaaatga acagcctgag aggcgaggac acggctgttt
attactgtgc gagagattgg 300tgggaagccg ggtgccggcc ctgttatttc
tttgactact ggggccaggg aagcctggtc 360accgtctcct ca
37294339DNAartificialCombination of PCR-generated and human
sequence 94gaaacgacac tcacgcagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccagcca gagtgtttta tacagctcca acaataagaa
ctacttagct 120tggtaccagc agaaaccagg acagcctcct aaggtgctca
tttactgggc ttctacccgg 180gaatccgggg tccctgaccg attcagtggc
agcgggtctg ggaccgattt cactctcacc 240atcagcagcc tgcaggctga
agatgtggca gtttattact gtcagcaata ttatagtcct 300ccgctcactt
tcggcggagg gaccaagctg gagatcaaa 33995127PRTartificialCombination of
PCR-generated and human sequence 95Glu Val Gln Leu Val Gln Ser Gly
Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala
Ala Ser Gly Phe Thr Phe Ser Asp Tyr 20 25 30Trp Met Ser Trp Val Arg
Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Asn Ile Asn Gln
Asp Gly Ser Ala Ala Tyr Tyr Val Asp Ser Val 50 55 60Arg Gly Arg Phe
Thr Ile Ser Arg Asp Ser Ala Glu Asn Ser Leu Asn65 70 75 80Leu Gln
Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala
Lys Asp Ala His Tyr Tyr Asp Arg Asn Arg Asn Asn Tyr Tyr Tyr 100 105
110Tyr Phe Asp Phe Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115
120 12596107PRTartificialCombination of PCR-generated and human
sequence 96Glu Ile Val Met Thr Gln Ser Pro Ala Thr Leu Ser Val Ser
Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val
Gly Ala Asn 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro
Arg Leu Leu Ile 35 40 45Tyr Ser Ala Ser Thr Arg Ala Thr Gly Val Pro
Ala Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu Phe Thr Leu Thr
Ile Ser Ser Leu Gln Ser65 70 75 80Glu Asp Phe Ala Val Tyr Tyr Cys
Gln Gln Tyr Asn Asn Trp Pro Arg 85 90 95Thr Phe Gly Gln Gly Thr Lys
Val Glu Ile Lys 100 10597381DNAartificialCombination of
PCR-generated and human sequence 97gaggtgcagc tggtgcagtc tgggggaggc
ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt
gactattgga tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtggccaac ataaatcaag atggaagtgc tgcatactat 180gtggactctg
tgaggggccg attcaccatc tccagagaca gcgccgagaa ctcactgaat
240ctgcaaatga acagcctgag agccgaggac acggctgtgt attactgtgc
gaaagatgca 300cattactacg atcgtaatcg taataattat tactactact
ttgacttctg gggccaggga 360accctggtca ccgtctcctc a
38198321DNAartificialCombination of PCR-generated and human
sequence 98gaaatagtga tgacgcagtc tccggccacc ctgtctgtgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttggc gccaacttag cctggtacca
gcagaaacct 120gggcaggctc ccaggctcct catctattct gcatccacca
gggccactgg tgtcccagcc 180aggttcagtg gcagtgggtc tgggacagag
ttcactctca ccatcagcag cctgcagtct 240gaagattttg cagtttatta
ctgtcagcag tataataact ggcctcggac gttcggccaa 300gggaccaagg
tggaaatcaa a 32199127PRTartificialCombination of PCR-generated and
human sequence 99Gln Val Gln Leu Leu Glu Ser Gly Pro Gly Leu Val
Asn Pro Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly
Ser Ile Ser Ser Gly 20 25 30Asn Tyr Tyr Trp Ser Trp Ile Arg Gln Pro
Ala Gly Lys Gly Leu Glu 35 40 45Trp Ile Gly Arg Met Ser Ser Ser Gly
Ser Thr Asn Tyr Asn Pro Ser 50 55 60Leu Lys Ser Arg Val Thr Met Ser
Glu Asp Thr Ser Lys Asn Gln Phe65 70 75 80Ser Leu Lys Leu Ser Ser
Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr 85 90 95Cys Ala Arg Glu Ser
Gly Ser Ser Trp Gln Asn His Tyr Tyr Tyr Tyr 100 105 110Gly Met Asp
Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120
125100113PRTartificialCombination of PCR-generated and human
sequence 100Glu Ile Val Leu Thr Gln Ser Pro Asp Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Asn Ile Asn Cys Lys Ser Ser Gln Ser Val
Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln
Lys Ser Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr
Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala Glu Asp
Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Thr Pro Leu Thr
Phe Gly Gly Gly Thr Lys Val Asp Ile 100 105
110Lys101381DNAartificialCombination of PCR-generated and human
sequence 101caggtgcagc tgttggagtc gggcccagga ctggtgaacc cttcacagac
cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtggtaatt actactggag
ctggatccgg 120cagcccgccg ggaagggact ggagtggatt gggcgtatgt
ctagtagtgg gagcaccaac 180tacaacccct ccctcaagag tcgagtcacc
atgtcagaag acacgtccaa gaaccagttc 240tccctgaagt tgagctctgt
gaccgccgca gacacggccg tgtattactg tgccagagaa 300tcgggtagca
gctggcaaaa tcactactac tactacggta tggacgtctg gggccaaggg
360accacggtca ccgtctcctc a 381102339DNAartificialCombination of
PCR-generated and human sequence 102gaaattgtgt tgacacagtc
tccagactcc ctggctgtgt ctctgggcga gagggccaac 60attaattgca agtccagcca
gagtgtttta tacagctcca acaataagaa ctacttagct 120tggtaccagc
agaaatcagg acagcctcct aagctgctca tttactgggc atctacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtctg ggacagattt
cactctcacc 240atcagcagcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttatagtact 300cctctcactt tcggcggagg gaccaaagtg gatatcaaa
339103123PRTartificialCombination of PCR-generated and human
sequence 103Gln Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Arg Asn Tyr 20 25 30Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Ser Asp Gly Asn Thr Phe Tyr
Ala Asp Ser Val Lys 50 55 60Gly Arg Phe Thr Val Ser Arg Asp Asn Ser
Lys Asn Thr Leu Tyr Leu65 70 75 80Gln Val Asn Ser Leu Arg Ala Asp
Asp Thr Ala Val Tyr Tyr Cys Ala 85 90 95Lys Glu Ser Gly Arg Trp Gly
Gly Gly Thr Leu Tyr Gly Ala His Tyr 100 105 110Trp Gly Gln Gly Thr
Pro Val Thr Val Ser Ser 115 120104113PRTartificialCombination of
PCR-generated and human sequence 104Asp Ile Gln Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Leu Leu Tyr Asn 20 25 30Ser Asn Asn Lys Asn Tyr
Leu Thr Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr
Tyr Ser Ser Pro Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105
110Lys105369DNAartificialCombination of PCR-generated and human
sequence 105caggtacagc tggtggagtc tgggggaggc ttggtacagc ctggggggtc
cctgagactc 60tcctgtgcag cctctggatt cacctttagg aactatgcca tgagttgggt
ccgccaggct 120ccagggaagg ggctggagtg ggtctcaggc attagtagtg
acggtaatac attctacgca 180gactccgtga agggccggtt caccgtctcc
agagacaatt ccaagaacac gttgtatctg 240caagtgaaca gtctgagagc
cgacgacacg gccgtatatt actgtgcgaa agagagtggc 300cgttggggtg
gtggaacctt atacggggcg cactactggg gccagggaac cccggtcacc 360gtctcctca
369106339DNAartificialCombination of PCR-generated and human
sequence 106gacatccaga tgacccagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccagcca gagtttgtta tacaactcca ataataagaa
ctacttaact 120tggtaccagc aaaaaccagg acagcctcct aagttgctca
tttactgggc atctacccgg 180gaatccgggg tccctgaccg attcagtggc
agcgggtctg ggacagattt cactctcacc 240atcagcagcc tgcaggctga
agatgtggca gtttattact gtcagcaata ttatagtagt 300cctctcactt
tcggcggagg gaccaaggtg gagatcaaa 339107121PRTartificialCombination
of PCR-generated and human sequence 107Gln Val Gln Leu Val Gln Ser
Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Pro Val Lys Val Ser Cys
Glu Thr Ser Gly Tyr Arg Phe Ser Asp Tyr 20 25 30Asn Val His Trp Val
Arg Gln Ala Pro Gly Gln Gly Pro Glu Trp Ile 35 40 45Gly Arg Ile Ser
Pro Asn Ser Gly Gly Thr Lys Tyr Ala Gln Lys Phe 50 55 60Gln Gly Arg
Val Thr Met Thr Arg Asp Met Ser Met Asn Thr Ala Tyr65 70 75 80Met
Glu Leu Ser Gly Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys 85 90
95Val Arg Gly His Cys Asp Gly Thr Thr Cys Ser Arg Ala Tyr Trp Gly
100 105 110Gln Gly Thr Leu Val Thr Val Ser Ser 115
120108112PRTartificialCombination of PCR-generated and human
sequence 108Asp Val Val Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr
Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser Cys Arg Ser Ser Gln Ser Leu
Leu His Arg 20 25 30Ser Gly His Lys Tyr Leu His Trp Tyr Leu Gln Arg
Pro Gly Gln Ser 35 40 45Pro Gln Val Leu Ile Tyr Leu Gly Ser Asn Arg
Ala Ser Gly Val Pro 50 55 60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr
Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu Ala Glu Asp Val
Gly Leu Tyr Tyr Cys Met Gln Thr 85 90 95Leu Gln Thr Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
110109363DNAartificialCombination of PCR-generated and human
sequence 109caggtgcagc tggtgcagtc tggggctgag gtgaagaagc ctggggcccc
agtgaaggtc 60tcctgcgaga cttctggata caggttcagc gactacaatg tacactgggt
gcgacaggcc 120cctggacaag ggcctgagtg gataggacgg atcagcccta
acagcggtgg cacaaagtat 180gcacagaagt ttcaaggcag ggtcaccatg
accagggaca tgtccatgaa cacagcctac 240atggagctga gcgggctgag
atctgacgac acggccgtgt attattgtgt aagaggacat 300tgtgatggta
ccacttgctc tcgtgcctac tggggccagg gaaccctggt caccgtctcc 360tca
363110336DNAartificialCombination of PCR-generated and human
sequence 110gatgttgtga tgacgcagtc tccactctcc ctgcccgtca cccctggaga
gccggcctcc 60atctcctgca ggtctagtca gagcctcctg catagaagtg gacacaagta
tttgcattgg 120tacctgcaga ggccagggca gtctccacag gtcctgatct
atttgggttc taatcgggcc 180tccggggtcc ctgacaggtt cagtggcagt
ggatcaggca cagattttac actgaaaatc 240agcagagtgg aggctgagga
tgttgggctt tattactgca tgcaaactct acaaaccccc 300tggacattcg
gccaagggac caaggtggaa atcaaa 336111120PRTartificialCombination of
PCR-generated and human sequence 111Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Thr
Ala Ser Gly Tyr Thr Phe Thr Gly Tyr 20 25 30Tyr Leu His Trp Val Arg
Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Trp Val Asn Pro
Arg Ser Gly Gly Thr Ser Tyr Pro Pro Lys Phe 50 55 60Gln Gly Arg Val
Thr Met Thr Arg Asp Thr Ser Ile Asn Thr Ala Tyr65 70 75 80Met Asp
Leu Thr Trp Leu Thr Ser Asp Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala
Val Gly Arg Ile Pro Asp Val Thr Ala Phe Asp Ile Trp Gly Gln 100 105
110Gly Thr Pro Val Thr Val Ser Ser 115
120112113PRTartificialCombination of PCR-generated and human
sequence 112Asp Ile Gln Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Glu Ser Leu
Leu Tyr Asp 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln
Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr
Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Glu
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala Glu Asp
Val Ala Val Tyr His Cys Gln Gln 85 90 95Tyr Phe Ser Thr Pro Trp Thr
Phe Gly Gln Gly Thr Lys Leu Glu Ile 100 105
110Lys113360DNAartificialCombination of PCR-generated and human
sequence 113caggtccagc ttgtgcagtc tggggctgag gtgaagaagc ctggggcctc
agtgaaggtc 60tcctgtacgg cctctggata cactttcacc ggctattatt tacactgggt
gcgacaggcc 120cctggacaag gacttgagtg gatggggtgg gtcaacccta
ggagtggtgg cacaagctat 180ccaccgaagt ttcagggcag ggtcaccatg
accagggaca cgtccatcaa cacagcctac 240atggacctga cctggctgac
atctgacgac acggccgtct attattgtgc ggtcggaaga 300atacctgatg
taactgcttt cgatatctgg ggccagggga caccggtcac cgtctcctca
360114339DNAartificialCombination of PCR-generated and human
sequence 114gacatccaga tgacccagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccagcga gagtctttta tacgactcca acaataagaa
ctacttagct 120tggtaccagc agaaaccagg acagcctcct aagttgctca
tttattgggc atctacccgg 180gaatccgggg tccctgaccg attcagtggc
agcgggtctg agacagattt cactctcacc 240atcagcagcc tgcaggctga
agatgtggca gtttatcact gtcaacaata tttcagtact 300ccttggacgt
tcggccaggg gaccaagctg gagatcaaa 339115122PRTartificialCombination
of PCR-generated and human sequence 115Gln Val Gln Leu Val Glu Ser
Gly Pro Glu Met Arg Lys Pro Gly Glu1 5 10 15Ser Leu Lys Ile Ser Cys
Lys Thr Ser Gly Tyr Ile Phe Ser Asp Tyr 20 25 30Trp Thr Ala Trp Val
Arg Gln Leu Pro Gly Lys Gly Leu Gln Trp Met 35 40 45Gly Ile Ile Tyr
Ser Gly Asp Ser Asp Thr Arg Tyr His Pro Ser Val 50 55 60Gln Gly His
Val Thr Met Ser Thr Asp Ser Ser Leu Thr Thr Ala Tyr65 70 75 80Leu
Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr Gly Ile Tyr Tyr Cys 85 90
95Ala Arg Leu Asp Ala Arg Val Asp Ala Gly Trp Gln Leu Asp Ser Trp
100 105 110Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115
120116113PRTartificialCombination of PCR-generated and human
sequence 116Asp Ile Gln Leu Thr Gln Ser Pro Asp Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val
Phe Ser Arg 20 25 30Asp Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln His
Lys Ser Gly Gln 35 40 45Pro Pro Lys Leu Leu Phe Phe Trp Ala Ser Ser
Arg Glu Ser Gly Val 50 55 60Ser Asp Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Asp Asn Leu Gln Ala Glu Asp
Val Ala Leu Tyr Tyr Cys Gln His
85 90 95Tyr Phe Asn Thr Pro His Asn Phe Gly Gln Gly Thr Lys Leu Glu
Ile 100 105 110Lys117366DNAartificialCombination of PCR-generated
and human sequence 117caggttcagc tggtggagtc tggaccggag atgagaaagc
ccggggagtc tctgaaaatt 60tcctgcaaga cttctggcta catatttagc gactactgga
ctgcctgggt gcgccagctg 120cccgggaagg gccttcagtg gatgggaatc
atctattctg gtgactctga taccagatat 180catccgtccg tccaaggcca
cgtcaccatg tcaaccgaca gttccctcac caccgcctac 240ctgcagtgga
gcagcctgaa ggcctcggac accggcatat attactgtgc gcgccttgat
300gcaagagttg atgctggatg gcaattagat tcgtggggcc aggggaccct
ggtcaccgtc 360tcttca 366118339DNAartificialCombination of
PCR-generated and human sequence 118gacatccagt tgacccagtc
tccagattcc ctggctgtgt ctctgggcga gcgggccacc 60atcaactgca agtccagcca
gagtgttttc tccagggaca acaataaaaa ctacctagct 120tggtaccagc
acaaatcagg gcagcctcct aagttgctct ttttctgggc gtccagtcgt
180gaatctgggg tctcagaccg attcagcggc agcgggtctg ggacagattt
cactctcacc 240atcgacaacc tgcaggctga agatgtggca ctttattact
gtcaacatta ttttaatact 300ccccacaatt ttggccaggg gaccaagctg gagatcaaa
339119121PRTartificialCombination of PCR-generated and human
sequence 119Gln Val Gln Leu Gln Gln Ser Gly Ala Glu Val Lys Lys Pro
Gly Glu1 5 10 15Ser Leu Lys Ile Ser Cys Glu Ala Ser Gly Tyr Ser Phe
Thr Asn Tyr 20 25 30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly
Leu Glu Trp Met 35 40 45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg
Tyr Ser Pro Pro Phe 50 55 60Gln Gly Gln Val Thr Ile Thr Ala Asp Arg
Ser Ile Thr Thr Ala Tyr65 70 75 80Leu Glu Trp Ser Ser Leu Lys Ala
Ser Asp Thr Ala Met Tyr Tyr Cys 85 90 95Ala Arg Val Gly Arg Pro Ser
Lys Gly Gly Trp Phe Asp Pro Trp Gly 100 105 110Gln Gly Thr Leu Val
Thr Val Ser Ser 115 120120113PRTartificialCombination of
PCR-generated and human sequence 120Asp Ile Gln Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Glu Ser Ser Gln Thr Leu Leu Tyr Ser 20 25 30Ser Asn Glu Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu
Ile Ser Trp Ala Ser Thr Pro Glu Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Ser Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr
Tyr Asn Ser Pro Tyr Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile 100 105
110Lys121363DNAartificialCombination of PCR-generated and human
sequence 121caggtgcagc tacagcagtc aggagcagaa gtgaaaaagc ccggggagtc
tctgaagatc 60tcctgtgagg cctctggata cagcttcacc aattactgga tcggctgggt
gcgccagatg 120cccggtaaag gcctggagtg gatgggaatc atctatcctg
gtgactctga caccagatac 180agtccgccct tccaaggcca ggtcaccatc
acagccgaca ggtccatcac caccgcctac 240ttggagtgga gcagtctgaa
ggcctcggac accgccatgt attactgtgc aagagttggg 300agaccttcta
aaggaggctg gttcgacccc tggggccagg gaaccctggt caccgtctct 360tca
363122339DNAartificialCombination of PCR-generated and human
sequence 122gacatccaga tgacccagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgtg agtccagcca gactctttta tacagttcca acgaaaagaa
ctacctagct 120tggtaccagc agaaaccagg acagcctcct aagttgctca
tttcctgggc ttctaccccg 180gaatccgggg tccctgaccg attcagtggc
agtgggtctg ggacaagttt cactctcacc 240atcagcagcc tgcaggctga
ggatgtggca gtttattact gtcagcaata ttataacagt 300ccatacactt
ttggccaagg gacacgactg gagattaaa 339123126PRTartificialCombination
of PCR-generated and human sequence 123Glu Val Gln Leu Leu Glu Ser
Gly Gly Gly Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys
Val Ala Ser Gly Ile Thr Phe Ser Asp Ser 20 25 30Tyr Met Ser Trp Ile
Arg Gln Thr Pro Gly Lys Gly Leu Glu Trp Leu 35 40 45Ser Tyr Ile Ser
Arg Ser Ser Ser His Thr Asn Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg
Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu
Gln Met Asn Ser Leu Arg Gly Glu Asp Thr Ala Val Tyr Tyr Cys 85 90
95Ala Arg Val Gln Thr Thr Met Ile Glu Gly Lys Thr Lys Leu Asn Tyr
100 105 110Phe Asp Tyr Trp Gly Gln Gly Thr Gln Val Thr Val Ser Ser
115 120 125124113PRTartificialCombination of PCR-generated and
human sequence 124Asp Ile Gln Met Thr Gln Ser Pro Asp Ser Leu Ala
Val Ser Leu Gly1 5 10 15Glu Arg Val Thr Ile Thr Cys Glu Ser Ser His
Ser Leu Leu Tyr Arg 20 25 30Ser Asn Asn Arg Asn Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Arg Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala
Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly
Ser Glu Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Arg Ala
Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Phe Tyr Thr Thr Pro
Tyr Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys125378DNAartificialCombination of PCR-generated and human
sequence 125gaggtgcagc tgttggagtc tgggggaggc ttggtcaagc ctggagggtc
cctgagactc 60tcctgtgtcg cctctggaat caccttcagt gactcctaca tgagctggat
ccgccagact 120ccagggaagg ggctagagtg gctttcatac attagtcgta
gtagttctca cacaaattac 180gcagactctg tgaagggccg attcaccatc
tccagagaca acgccaagaa ttcactatat 240ctccaaatga atagcctgag
aggcgaggac acggctgtgt attactgtgc gagagtccag 300acaactatga
tagaagggaa aacgaaactt aactactttg actactgggg ccagggaacc
360caggtcaccg tctcctca 378126339DNAartificialCombination of
PCR-generated and human sequence 126gacatccaga tgacccagtc
tccagactcc ctggctgtgt ctctgggcga gagggtcacc 60atcacctgcg agtccagcca
cagtctttta tacaggtcca acaataggaa ctacttagct 120tggtaccagc
aaaaaccaag acagcctcct aagttgctca tttactgggc atctacccgg
180gagtccgggg tcccagaccg attcagcggc agcgggtctg agacagattt
cactctcacc 240atcagcagcc tgcgggctga agatgtggcg gtatattact
gtcaacaatt ttatactact 300ccttacactt ttggccaagg gaccaaggtg gagatcaaa
339127126PRTartificialCombination of PCR-generated and human
sequence 127Glu Val Gln Leu Val Gln Ser Gly Gly Gly Leu Val Lys Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Val Ala Ser Gly Ile Thr Phe
Ser Asp Ser 20 25 30Tyr Met Ser Trp Ile Arg Gln Thr Pro Gly Lys Gly
Leu Glu Trp Leu 35 40 45Ser Tyr Ile Ser Arg Ser Ser Ser His Thr Asn
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Gly
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Val Gln Thr Thr Met
Ile Glu Gly Lys Thr Lys Leu Asn Tyr 100 105 110Phe Asp Tyr Trp Gly
Gln Gly Thr Leu Val Thr Val Ser Ser 115 120
125128113PRTartificialCombination of PCR-generated and human
sequence 128Ala Ile Gln Leu Thr Gln Ser Pro Asp Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Val Thr Ile Ser Cys Glu Ser Ser His Ser Leu
Leu Tyr Arg 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln
Lys Pro Arg Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr
Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Glu
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Arg Ala Glu Asp
Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Phe Tyr Thr Thr Pro Tyr Thr
Phe Gly Gln Gly Thr Lys Leu Glu Ile 100 105
110Lys129378DNAartificialCombination of PCR-generated and human
sequence 129gaggtgcagc tggtgcagtc tgggggaggc ttggtcaagc ctggagggtc
cctgagactc 60tcctgtgtcg cctctggaat caccttcagt gactcctaca tgagctggat
ccgccagact 120ccagggaagg ggctagagtg gctttcatac attagtcgta
gtagttctca cacaaattac 180gcagactctg tgaagggccg attcaccatc
tccagagaca acgccaagaa ttcactatat 240ctccaaatga atagcctgag
aggcgaggac acggctgtgt attactgtgc gagagtccag 300acaactatga
tagaagggaa aacgaaactt aactactttg actactgggg ccagggaacc
360ctggtcaccg tctcctca 378130339DNAartificialCombination of
PCR-generated and human sequence 130gccatccagt tgacccagtc
tccagactcc ctggctgtgt ctctgggcga gagggtcacc 60atcagctgcg agtccagcca
cagtctttta tacaggtcca acaataagaa ctacttagct 120tggtaccagc
agaaaccaag acagcctcct aagttgctca tttactgggc atctacccgg
180gagtccgggg tcccagaccg attcagcggc agcgggtctg agacagattt
cactctcacc 240atcagcagcc tgcgggctga agatgtggca gtgtattact
gtcaacaatt ttatactact 300ccttacactt ttggccaggg gaccaagctg gagatcaaa
339131122PRTartificialCombination of PCR-generated and human
sequence 131Gln Leu Val Gln Ser Glu Gly Gly Leu Ala Glu Pro Gly Gly
Ser Leu1 5 10 15Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Thr Lys
Ala Trp Met 20 25 30Ser Trp Val Arg Gln Thr Pro Gly Arg Gly Leu Glu
Trp Leu Gly Arg 35 40 45Ile Lys Ser Lys Val Asp Gly Glu Thr Thr Asp
Tyr Ala Ala Pro Val 50 55 60Arg Gly Arg Phe Thr Ile Ser Arg Asp Asp
Ser Lys Ser Thr Val Tyr65 70 75 80Leu Gln Met Thr Gly Leu Arg Thr
Glu Asp Thr Gly Val Tyr Phe Cys 85 90 95Thr Thr Leu Ile His Cys Asp
Leu Ser Ala Cys Leu Pro His Phe Trp 100 105 110Gly Arg Gly Thr Leu
Val Thr Val Ser Ser 115 120132113PRTartificialCombination of
PCR-generated and human sequence 132Glu Ile Val Leu Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Val Thr Ile Ser Cys
Glu Ser Ser His Ser Leu Leu Tyr Arg 20 25 30Ser Asn Asn Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Arg Gln 35 40 45Pro Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Glu Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Arg Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Phe
Tyr Thr Thr Pro Tyr Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile 100 105
110Lys133378DNAartificialCombination of PCR-generated and human
sequence 133gaggtgcagc tggtgcagtc tgggggaggc ttggtcaagc ctggagggtc
cctgagactc 60tcctgtgtcg cctctggaat caccttcagt gactcctaca tgagctggat
ccgccagact 120ccagggaagg ggctagagtg gctttcatac attagtcgta
gtagttctca cacaaattac 180gcagactctg tgaagggccg attcaccatc
tccagagaca acgccaagaa ttcactatat 240ctccaaatga atagcctgag
aggcgaggac acggctgtgt attactgtgc gagagtccag 300acaactatga
tagaagggaa aacgaaactt aactactttg actactgggg ccagggaacc
360ctggtcaccg tctcctca 378134339DNAartificialCombination of
PCR-generated and human sequence 134gaaattgtgt tgacgcagtc
tccagactcc ctggctgtgt ctctgggcga gagggtcacc 60atcagctgcg agtccagcca
cagtctttta tacaggtcca acaataagaa ctacttagct 120tggtaccagc
agaaaccaag acagcctcct aagttgctca tttactgggc atctacccgg
180gagtccgggg tcccagaccg attcagcggc agcgggtctg agacagattt
cactctcacc 240atcagcagcc tgcgggctga agatgtggca gtgtattact
gtcaacaatt ttatactact 300ccttacactt ttggccaggg gaccaagctg gagatcaaa
339135124PRTartificialCombination of PCR-generated and human
sequence 135Gln Val Gln Leu Val Gln Ser Gly Gly Glu Val Lys Lys Pro
Gly Glu1 5 10 15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe
Ser Asn Tyr 20 25 30Trp Ile Ala Trp Val Arg Gln Met Pro Gly Lys Gly
Leu Glu Cys Met 35 40 45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Thr
Tyr Ser Pro Ser Phe 50 55 60Gln Gly Gln Val Thr Ile Ser Ala Asp Lys
Ser Val Ser Thr Thr Tyr65 70 75 80Leu Gln Trp Ser Ser Leu Lys Ala
Ser Asp Ser Ala Met Tyr Tyr Cys 85 90 95Ala Arg Leu Pro Arg Thr Asp
Gly Asp Asn Ser Ile Gly Tyr Phe Glu 100 105 110Tyr Trp Gly Gln Gly
Thr Met Val Thr Val Ser Ser 115 120136113PRTartificialCombination
of PCR-generated and human sequence 136Glu Ile Val Leu Thr Gln Ser
Pro Ala Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn
Cys Lys Ser Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Ser Glu Asn
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu
Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg
Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile
Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90
95Tyr Tyr Ser Thr Pro Phe Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile
100 105 110Lys137372DNAartificialCombination of PCR-generated and
human sequence 137caggtgcagc tggtgcagtc tggaggagag gtgaaaaagc
cgggggagtc tctgaagatc 60tcctgtaagg gttctgggta ctcctttagt aactactgga
tcgcctgggt gcgccagatg 120cccgggaaag gcctggagtg catgggaatc
atctatcctg gtgactctga taccacatac 180agcccgtcct tccaaggcca
ggtcaccatt tctgccgaca agtccgtcag taccacctac 240ctacaatgga
gcagcctgaa ggcctcggac agcgccatgt attattgtgc gaggctaccc
300cgtacagatg gcgacaattc catcggctac tttgaatatt ggggccaggg
aaccatggtc 360accgtctctt ca 372138345DNAartificialCombination of
PCR-generated and human sequence 138gaaattgtgt tgacacagtc
tccagcctcc ctggctgtgt ctctgggcga gagggccacc 60atcaactgca agtccagcca
gagtgtttta tacagctcca acagtgagaa ctacttagct 120tggtatcagc
agaaaccagg acagcctccc aagttgctca tttactgggc gtctacccgg
180gaatccgggg tccctgaccg attcagtggc agcgggtctg ggacagattt
cactctcacc 240atcagcagcc tgcaggctga agatgtggca gtttattact
gtcagcaata ttatagtact 300ccattcactt tcggccaagg gacacgactg
gagattaaac gtaag 345139118PRTartificialCombination of PCR-generated
and human sequence 139Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
Phe Thr Phe Ser Ser Tyr 20 25 30Ser Met Asn Trp Val Arg Gln Ala Pro
Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Tyr Ile Ser Ser Ser Thr Thr
Thr Ile Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser
Arg Asp Asn Val Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met His Ser
Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Val Pro Ala Pro
Arg Leu Gly Gly Ser Tyr Thr Tyr Trp Gly Gln Gly 100 105 110Thr Leu
Val Thr Val Ser 115140108PRTartificialCombination of PCR-generated
and human sequence 140Asp Ile Gln Met Thr Gln Ser Pro Gly Thr Leu
Ser Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser
Gln Ser Val Ser Ser Ser 20 25 30Tyr Leu Ala Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser Ser Arg Ala
Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60Gly Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr Ile Ser Arg Leu Glu65 70 75 80Pro Glu Asp Phe Ala
Val Tyr Tyr Cys Gln Gln Tyr Gly Thr Ser Pro 85 90 95Leu Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile Lys 100
105141357DNAartificialCombination of PCR-generated and human
sequence 141gaggtgcagc tggtggagtc tgggggaggc ttggtacagc ctggggggtc
cctgagactc 60tcctgtgcag cctctggatt caccttcagt agttatagca tgaactgggt
ccgccaggct 120ccagggaagg ggctggagtg ggtttcatac attagtagta
gtactactac catatactac
180gcagactctg tgaagggccg attcaccatc tccagggaca atgtcaagaa
ctctctgtat 240ctgcaaatgc acagcctgag agccgaggac acggctgtgt
attactgtgt cccggccccc 300cggttgggtg ggagctacac ttactggggc
cagggaaccc tggtcaccgt ctcctca 357142324DNAartificialCombination of
PCR-generated and human sequence 142gacatccaga tgacccagtc
tccaggcacc ctgtctttgt ctccagggga aagagccacc 60ctctcctgca gggccagtca
gagtgttagc agcagctact tagcctggta ccagcagaaa 120cctggccagg
ctcccaggct cctcatctat ggtgcatcca gcagggccac tggcatccca
180gacaggttca gtggcagtgg gtctgggaca gacttcactc tcaccatcag
cagactggag 240cctgaagatt ttgcagtgta ttactgtcag cagtatggca
cctcaccgct cactttcggc 300ggagggacca aggtggaaat caaa
324143119PRTartificialCombination of PCR-generated and human
sequence 143Glu Val Gln Leu Val Glu Ser Gly Gly Asp Leu Val Lys Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Val Phe
Thr Asp Ala 20 25 30Trp Met Ser Trp Val Arg Gln Ser Pro Gly Lys Gly
Pro Glu Trp Leu 35 40 45Gly Arg Ile Lys Ser Lys Asn Val Gly Glu Thr
Thr Asp Tyr Ala Glu 50 55 60His Val Arg Gly Arg Phe Thr Ile Ala Arg
Asp Asp Ser Asn Arg Thr65 70 75 80Leu Tyr Leu Gln Met Ser Asn Leu
Lys Ile Asp Asp Thr Ala Val Tyr 85 90 95Tyr Cys Thr Thr Gly Leu Gly
Gly Gly Thr Tyr Gly Trp Gly Arg Gly 100 105 110Thr Arg Val Thr Val
Ser Ser 115144108PRTartificialCombination of PCR-generated and
human sequence 144Glu Ile Val Leu Thr Gln Ser Pro Leu Ser Leu Pro
Ala Thr Leu Gly1 5 10 15Gln Pro Ala Ser Ile Ser Cys Arg Ser Ser Ala
Gly Leu Arg Asn Asn 20 25 30Asp Gly Asp Ile Leu Leu Ser Trp Phe His
Gln Arg Pro Gly Gln Ser 35 40 45Pro Arg Arg Leu Phe Tyr Arg Val Ser
Arg Arg Asp Ser Gly Val Pro 50 55 60Asp Arg Phe Asn Gly Ser Gly Ser
Ala Thr Asp Phe Thr Leu Arg Ile65 70 75 80Asn Ser Val Glu Ala Glu
Asp Val Gly Ile Tyr Tyr Cys Met Arg Gly 85 90 95Pro Tyr Trp Gly Gln
Gly Thr Arg Leu Glu Ile Lys 100 105145357DNAartificialCombination
of PCR-generated and human sequence 145gaggtgcagc tggtggagtc
tgggggagac ttggtaaagc ctggggggtc ccttagactc 60tcctgtgcag cctctggatt
cgttttcact gacgcctgga tgagctgggt ccgccagagt 120cccgggaagg
ggccggagtg gcttggccgt atcaaaagta aaaatgtcgg tgagacaaca
180gactacgctg aacacgtgag aggcagattt accatcgcaa gagatgattc
caaccgcact 240ctatatctac aaatgagcaa cctgaaaatc gacgacacag
ccgtctatta ttgtaccact 300ggactgggag gagggaccta cggatggggc
cggggaaccc gggtcaccgt ctcttca 357146324DNAartificialCombination of
PCR-generated and human sequence 146gaaattgtgt tgacacagtc
tccactctcc ctgcccgcca cccttggaca gccggcctcc 60atctcctgca ggtcgagtgc
aggcctccga aacaacgatg gtgacatcct cttgagttgg 120tttcatcagc
ggccaggcca gtctccgagg cgcctatttt atagagtttc taggcgtgac
180tctggagtcc cagacagatt caacggcagt gggtcagcca ctgatttcac
actgagaatc 240aattctgtgg aggctgaaga tgttggcatt tactactgca
tgcgaggacc atattggggc 300caagggacac gactggagat taaa
324147119PRTartificialCombination of PCR-generated and human
sequence 147Gln Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Glu1 5 10 15Ser Leu Arg Leu Ser Cys Val Ala Ser Gly Phe Asn Phe
Ser Ile Tyr 20 25 30Glu Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Tyr Ile Thr Asn Arg Gly Ser Thr Ile Tyr
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Gln Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Lys Pro Arg Ile Gly Ala
Arg Val Phe Asp Val Trp Gly Gln Gly 100 105 110Thr Met Val Thr Val
Ser Ser 115148113PRTartificialCombination of PCR-generated and
human sequence 148Asp Ile Gln Leu Thr Gln Ser Pro Asp Ser Leu Ala
Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln
Thr Leu Leu Tyr Lys 20 25 30Ser Asn Asn Glu Asn Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala
Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala
Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Phe Thr Thr Ala
Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105
110Lys149357DNAartificialCombination of PCR-generated and human
sequence 149caggtacagc tggtggagtc tgggggaggc ttggtacagc ctggagagtc
cctgagactc 60tcctgtgtag cctctggatt caacttcagt atttatgaga tgaattgggt
ccgccaggct 120ccagggaagg ggctggagtg ggtttcatac attaccaatc
gaggtagtac catatactac 180gcagactctg tgaagggccg attcaccatc
tccagggaca acgcccagaa ctcactgtat 240ctgcaaatga acagcctgag
agccgaagac acggctgttt attactgtgc gaaaccccgt 300ataggagctc
gtgtatttga tgtctggggc caagggacaa tggtcaccgt ctcctca
357150339DNAartificialCombination of PCR-generated and human
sequence 150gacatccagt tgacccagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccagcca gactctttta tacaagtcca acaatgagaa
ctacttagct 120tggtaccagc agaaaccagg acagcctcca aagctgctca
tttactgggc atctactcgg 180gaatccgggg tccctgaccg attcagtggc
agcgggtctg ggacagattt cactctcacc 240atcagcagcc tgcaggctga
ggatgtggca gtttactact gtcagcaata ttttactact 300gcgctcactt
tcggcggagg gaccaaggtg gagatcaaa 339151121PRTartificialCombination
of PCR-generated and human sequence 151Gln Val Gln Leu Val Glu Ser
Gly Ala Val Val Lys Lys Pro Gly Glu1 5 10 15Ser Leu Lys Ile Ser Cys
Glu Ala Ser Gly Tyr Met Phe Leu Asp His 20 25 30Trp Ile Gly Trp Val
Arg Gln Met Pro Gly Lys Gly Leu Glu Trp Met 35 40 45Gly Ile Ile Phe
Pro Glu Asp Ser Asp Thr Arg Tyr Ser Gly Ser Phe 50 55 60Glu Gly Gln
Val Thr Ile Ser Ala Asp Arg Ser Val Asn Thr Val Tyr65 70 75 80Leu
Glu Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Tyr Cys 85 90
95Ala Arg Val Ser Val Val Arg Lys Gly Gly Trp Phe Asp Pro Trp Gly
100 105 110Gln Gly Thr Leu Val Thr Val Ser Ser 115
120152113PRTartificialCombination of PCR-generated and human
sequence 152Ala Ile Gln Leu Thr Gln Ser Pro Asp Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Leu
Leu Tyr Thr 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln
Lys Pro Gly Gln 35 40 45Thr Pro Lys Leu Leu Ile Ile Trp Ala Ser Thr
Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Thr Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala Glu Asp
Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Asn Ser Pro Tyr Thr
Phe Gly Gln Gly Thr Arg Leu Glu Ile 100 105
110Lys153363DNAartificialCombination of PCR-generated and human
sequence 153caggtgcagc tggtgcagtc tggagcagtg gtgaaaaagc ccggggagtc
tctgaagatc 60tcctgtgagg cttctggata catgttcctc gatcactgga tcggctgggt
gcgccagatg 120cccgggaaag gcctggagtg gatgggaatc atctttcctg
aggactctga taccagatat 180agtgggtcct tcgaaggcca ggtcaccatc
tcagccgaca ggtccgtcaa caccgtctac 240ctggagtgga gcagcctgaa
ggcctcggac accgccatgt attattgtgc gagagtctca 300gtagttcgta
aagggggctg gttcgaccca tggggccagg gaaccacggt caccgtctcc 360tca
363154339DNAartificialCombination of PCR-generated and human
sequencemisc_feature(5)..(6)n is a, c, g, or t 154gaaanncaac
tgacgcagtc tccagactcc ctggctgtgt ctctgggcga gagggccacc 60atcaactgca
agtccagcca gagtctttta tacacctcca acaataagaa ctacttagct
120tggtaccagc agaaaccagg acagactcct aaactgctca ttatctgggc
ctctacccga 180gaatccgggg tccctgaccg attcactggc agcgggtctg
ggacagattt cactctcacc 240atcagcagcc tgcaggctga agatgtggca
gtttattact gtcagcaata ttataatagt 300ccgtacactt ttggccaagg
gacacgactg gagattaaa 339155121PRTartificialCombination of
PCR-generated and human sequence 155Gln Val Gln Leu Val Glu Ser Gly
Ala Glu Leu Lys Lys Pro Gly Glu1 5 10 15Ser Leu Lys Ile Ser Cys Glu
Ala Ser Gly Tyr Ile Phe Ala Asp His 20 25 30Trp Ile Gly Trp Val Arg
Gln Met Pro Gly Lys Gly Leu Glu Trp Met 35 40 45Gly Ile Ile Phe Pro
Gly Asp Ser Asp Ile Arg Tyr Ser Pro Ser Phe 50 55 60Glu Gly Gln Val
Thr Ile Ser Val Asp Arg Ser Val Ser Thr Ala Phe65 70 75 80Leu Gln
Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Phe Cys 85 90 95Ala
Arg Val Ala Val Val Arg Lys Gly Gly Trp Phe Asp Ser Trp Gly 100 105
110Gln Gly Thr Arg Val Thr Val Ser Ser 115
120156113PRTartificialCombination of PCR-generated and human
sequence 156Glu Ile Val Met Thr Gln Ser Pro Glu Ser Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Thr Gln Ser Leu
Leu Trp Ser 20 25 30Ala Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Arg Gln
Lys Pro Arg Gln 35 40 45Thr Pro Glu Leu Leu Ile Thr Trp Ala Ser Thr
Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Thr Ser Leu Gln Ala Glu Asp
Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Asn Ser Pro Tyr Thr
Phe Gly Gln Gly Thr Arg Leu Glu Ile 100 105
110Lys157363DNAartificialCombination of PCR-generated and human
sequence 157caggtacagc tggtggagtc tggagcagaa ctgaaaaagc ccggggagtc
tctgaagatc 60tcctgtgagg catctggata catctttgcc gatcactgga tcggctgggt
gcgccagatg 120cccgggaaag gcctggagtg gatgggaatc atctttcctg
gtgactctga tatcagatat 180agtccgtcct tcgaaggcca ggtcaccatc
tcagtcgaca ggtccgtcag taccgccttc 240ctgcagtgga gcagcctgaa
ggcctcggac accgccatgt atttttgtgc gagagtcgca 300gtagtgcgta
aagggggctg gttcgactcc tggggccagg gaacccgggt caccgtctcc 360tca
363158339DNAartificialCombination of PCR-generated and human
sequence 158gaaattgtga tgacccagtc tccagagtcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccaccca gagtctttta tggagcgcca acaacaagaa
ctacttagct 120tggtaccggc agaaaccacg acagactcct gaactgctca
ttacgtgggc ttccacccgg 180gaatccgggg tccctgaccg attcagtggc
agcgggtctg ggacagattt cactctcacc 240atcaccagcc tgcaggctga
agatgtggca gtttattatt gtcaacaata ttataatagt 300ccgtacactt
ttggccaagg gacacgactg gagattaaa 339159126PRTartificialCombination
of PCR-generated and human sequence 159Gln Val Gln Leu Val Gln Ser
Gly Ala Glu Val Lys Lys Pro Trp Glu1 5 10 15Ser Leu Lys Ile Ser Cys
Leu Ala Ser Gly Tyr Asp Phe Ala Ser Tyr 20 25 30Trp Ile Gly Trp Val
Arg Gln Met Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Gly Ile Ile Tyr
Pro Asp Asp Ser Asp Thr Arg Tyr Asn Ala Ser Leu 50 55 60Glu Gly Arg
Val Thr Met Ser Val Asp Thr Ser Thr Asn Thr Ala Tyr65 70 75 80Leu
Gln Trp Thr Ser Leu Lys Val Ser Asp Thr Gly Met Tyr Tyr Cys 85 90
95Ala Arg Arg Asp Arg Asn Cys Ser Gly Thr Thr Cys Tyr Pro Arg Trp
100 105 110Phe Asp Ser Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser
115 120 125160113PRTartificialCombination of PCR-generated and
human sequence 160Glu Ile Val Leu Thr Gln Ser Pro Asp Phe Leu Ala
Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln
Ser Leu Phe Tyr Ser 20 25 30Gly Asn Ser Lys Asp Phe Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Arg Leu Leu Val Tyr Trp Ala
Ser Thr Arg Asp Ser Gly Val 50 55 60Pro Glu Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Arg Leu Gln Ala
Glu Asp Val Ala Leu Tyr Tyr Cys His Gln 85 90 95Tyr His Ser Thr Pro
Leu Ser Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105
110Lys161378DNAartificialCombination of PCR-generated and human
sequence 161caggtgcagc tggtgcagtc tggggcagag gtgaaaaagc cgtgggagtc
tctgaagatc 60tcctgtttgg cttctggata cgactttgcc tcctactgga tcggctgggt
gcgccagatg 120cccgggaaag gcctggagtg ggtggggatc atctatcctg
atgactctga taccagatac 180aatgcgtcac tagaaggccg ggtcaccatg
tcagtcgaca cgtccaccaa taccgcctac 240ctgcagtgga ccagcctgaa
ggtctcggac accggcatgt attactgtgc gagacgggat 300cgcaattgta
gtgggactac gtgttatccg aggtggttcg actcctgggg ccagggaacc
360ctggtcaccg tctcttca 378162339DNAartificialCombination of
PCR-generated and human sequence 162gaaattgtgc tgactcagtc
tccagacttc ctggctgtgt ctctgggcga gagggccacc 60atcaactgca agtccagcca
gagtcttttc tacagcggca acagtaagga cttcttagct 120tggtaccagc
agaaaccagg acagcctcct cgcttgctcg tttactgggc atctacccgg
180gattccgggg tccctgagcg attcagtggc agcgggtctg ggacagattt
tactctcacc 240atcagccgcc tgcaggctga agatgtggct ctttattact
gtcaccaata tcatagtact 300cctctctctt tcggcggagg gaccaaggtg gaaatcaaa
3391635PRTHomo sapiens 163Ser Tyr Trp Met His1 516417PRTHomo
sapiens 164Arg Ile Asn Ser Asp Gly Ser Asp Thr Asn Tyr Ala Asp Ser
Val Lys1 5 10 15Gly1659PRTHomo sapiens 165Gly Arg Ser Tyr Gly Phe
Phe Asp Tyr1 516612PRTHomo sapiens 166Arg Ala Ser Gln Ile Ile Ser
Ser Asn Tyr Leu Ala1 5 101677PRTHomo sapiens 167Gly Ala Ser Ser Arg
Ala Thr1 51689PRTHomo sapiens 168Gln Gln Tyr Gly Thr Ser Pro Arg
Thr1 51695PRTHomo sapiens 169Thr Tyr Gly Met His1 517017PRTHomo
sapiens 170Val Ile Trp Phe Asp Gly Asn Asn Lys Tyr Tyr Ala Asp Ser
Val Lys1 5 10 15Gly17115PRTHomo sapiens 171Asp Trp Trp Glu Ala Gly
Cys Arg Pro Cys Tyr Phe Phe Asp Tyr1 5 10 1517217PRTHomo sapiens
172Lys Ser Ser Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys Asn Tyr Leu1
5 10 15Ala1737PRTHomo sapiens 173Trp Ala Ser Thr Arg Glu Ser1
51749PRTHomo sapiens 174Gln Gln Tyr Tyr Ser Pro Pro Leu Thr1
51755PRTHomo sapiens 175Asp Tyr Trp Met Ser1 517617PRTHomo sapiens
176Asn Ile Asn Gln Asp Gly Ser Ala Ala Tyr Tyr Val Asp Ser Val Arg1
5 10 15Gly17718PRTHomo sapiens 177Asp Ala His Tyr Tyr Asp Arg Asn
Arg Asn Asn Tyr Tyr Tyr Tyr Phe1 5 10 15Asp Phe17811PRTHomo sapiens
178Arg Ala Ser Gln Ser Val Gly Ala Asn Leu Ala1 5 101797PRTHomo
sapiens 179Ser Ala Ser Thr Arg Ala Thr1 51809PRTHomo sapiens 180Gln
Gln Tyr Asn Asn Trp Pro Arg Thr1 51817PRTHomo sapiens 181Ser Gly
Asn Tyr Tyr Trp Ser1 518216PRTHomo sapiens 182Arg Met Ser Ser Ser
Gly Ser Thr Asn Tyr Asn Pro Ser Leu Lys Ser1 5 10 1518317PRTHomo
sapiens 183Glu Ser Gly Ser Ser Trp Gln Asn His Tyr Tyr Tyr Tyr Gly
Met Asp1 5 10 15Val1849PRTHomo sapiens 184Gln Gln Tyr Tyr Ser Thr
Pro Leu Thr1 51855PRTHomo sapiens 185Asn Tyr Ala Met Ser1
518616PRTHomo sapiens 186Gly Ile Ser Ser Asp Gly Asn Thr Phe Tyr
Ala Asp Ser Val Lys Gly1 5 10 1518715PRTHomo sapiens 187Glu Ser Gly
Arg Trp Gly Gly Gly Thr Leu Tyr Gly Ala His Tyr1 5 10
1518817PRTHomo sapiens 188Lys Ser Ser Gln Ser Leu Leu Tyr Asn Ser
Asn Asn Lys Asn Tyr Leu1 5 10 15Thr1899PRTHomo sapiens 189Gln Gln
Tyr Tyr Ser Ser Pro Leu Thr1 51905PRTHomo sapiens 190Asp Tyr Asn
Val
His1 519117PRTHomo sapiens 191Arg Ile Ser Pro Asn Ser Gly Gly Thr
Lys Tyr Ala Gln Lys Phe Gln1 5 10 15Gly19212PRTHomo sapiens 192Gly
His Cys Asp Gly Thr Thr Cys Ser Arg Ala Tyr1 5 1019316PRTHomo
sapiens 193Arg Ser Ser Gln Ser Leu Leu His Arg Ser Gly His Lys Tyr
Leu His1 5 10 151947PRTHomo sapiens 194Leu Gly Ser Asn Arg Ala Ser1
51959PRTHomo sapiens 195Met Gln Thr Leu Gln Thr Pro Trp Thr1
51965PRTHomo sapiens 196Gly Tyr Tyr Leu His1 519717PRTHomo sapiens
197Trp Val Asn Pro Arg Ser Gly Gly Thr Ser Tyr Pro Pro Lys Phe Gln1
5 10 15Gly19811PRTHomo sapiens 198Gly Arg Ile Pro Asp Val Thr Ala
Phe Asp Ile1 5 1019917PRTHomo sapiens 199Lys Ser Ser Glu Ser Leu
Leu Tyr Asp Ser Asn Asn Lys Asn Tyr Leu1 5 10 15Ala2009PRTHomo
sapiens 200Gln Gln Tyr Phe Ser Thr Pro Trp Thr1 52015PRTHomo
sapiens 201Asp Tyr Trp Thr Ala1 520217PRTHomo sapiens 202Ile Ile
Tyr Ser Gly Asp Ser Asp Thr Arg Tyr His Pro Ser Val Gln1 5 10
15Gly20313PRTHomo sapiens 203Leu Asp Ala Arg Val Asp Ala Gly Trp
Gln Leu Asp Ser1 5 1020417PRTHomo sapiens 204Lys Ser Ser Gln Ser
Val Phe Ser Arg Asp Asn Asn Lys Asn Tyr Leu1 5 10 15Ala2057PRTHomo
sapiens 205Trp Ala Ser Ser Arg Glu Ser1 52069PRTHomo sapiens 206Gln
His Tyr Phe Asn Thr Pro His Asn1 52075PRTHomo sapiens 207Asn Tyr
Trp Ile Gly1 520817PRTHomo sapiens 208Ile Ile Tyr Pro Gly Asp Ser
Asp Thr Arg Tyr Ser Pro Pro Phe Gln1 5 10 15Gly20912PRTHomo sapiens
209Val Gly Arg Pro Ser Lys Gly Gly Trp Phe Asp Pro1 5
1021017PRTHomo sapiens 210Glu Ser Ser Gln Thr Leu Leu Tyr Ser Ser
Asn Glu Lys Asn Tyr Leu1 5 10 15Ala2117PRTHomo sapiens 211Trp Ala
Ser Thr Pro Glu Ser1 52129PRTHomo sapiens 212Gln Gln Tyr Tyr Asn
Ser Pro Tyr Thr1 52135PRTHomo sapiens 213Asp Ser Tyr Met Ser1
521417PRTHomo sapiens 214Tyr Ile Ser Arg Ser Ser Ser His Thr Asn
Tyr Ala Asp Ser Val Lys1 5 10 15Gly21517PRTHomo sapiens 215Val Gln
Thr Thr Met Ile Glu Gly Lys Thr Lys Leu Asn Tyr Phe Asp1 5 10
15Tyr21617PRTHomo sapiens 216Glu Ser Ser His Ser Leu Leu Tyr Arg
Ser Asn Asn Arg Asn Tyr Leu1 5 10 15Ala2179PRTHomo sapiens 217Gln
Gln Phe Tyr Thr Thr Pro Tyr Thr1 521817PRTHomo sapiens 218Glu Ser
Ser His Ser Leu Leu Tyr Arg Ser Asn Asn Lys Asn Tyr Leu1 5 10
15Ala2195PRTHomo sapiens 219Lys Ala Trp Met Ser1 522019PRTHomo
sapiens 220Arg Ile Lys Ser Lys Val Asp Gly Glu Thr Thr Asp Tyr Ala
Ala Pro1 5 10 15Val Arg Gly22113PRTHomo sapiens 221Leu Ile His Cys
Asp Leu Ser Ala Cys Leu Pro His Phe1 5 102225PRTHomo sapiens 222Asn
Tyr Trp Ile Ala1 522317PRTHomo sapiens 223Ile Ile Tyr Pro Gly Asp
Ser Asp Thr Thr Tyr Ser Pro Ser Phe Gln1 5 10 15Gly22415PRTHomo
sapiens 224Leu Pro Arg Thr Asp Gly Asp Asn Ser Ile Gly Tyr Phe Glu
Tyr1 5 10 1522517PRTHomo sapiens 225Lys Ser Ser Gln Ser Val Leu Tyr
Ser Ser Asn Ser Glu Asn Tyr Leu1 5 10 15Ala2269PRTHomo sapiens
226Gln Gln Tyr Tyr Ser Thr Pro Phe Thr1 52275PRTHomo sapiens 227Ser
Tyr Ser Met Asn1 522817PRTHomo sapiens 228Tyr Ile Ser Ser Ser Thr
Thr Thr Ile Tyr Tyr Ala Asp Ser Val Lys1 5 10 15Gly22912PRTHomo
sapiens 229Val Pro Ala Pro Arg Leu Gly Gly Ser Tyr Thr Tyr1 5
1023012PRTHomo sapiens 230Arg Ala Ser Gln Ser Val Ser Ser Ser Tyr
Leu Ala1 5 102319PRTHomo sapiens 231Gln Gln Tyr Gly Thr Ser Pro Leu
Thr1 52325PRTHomo sapiens 232Asp Ala Trp Met Ser1 523319PRThomo
sapiens 233Arg Ile Lys Ser Lys Asn Val Gly Glu Thr Thr Asp Tyr Ala
Glu His1 5 10 15Val Arg Gly2348PRThomo sapiens 234Gly Leu Gly Gly
Gly Thr Tyr Gly1 523516PRTHomo sapiens 235Arg Ser Ser Ala Gly Leu
Arg Asn Asn Asp Gly Asp Ile Leu Leu Ser1 5 10 152367PRThomo sapiens
236Arg Val Ser Arg Arg Asp Ser1 52375PRThomo sapiens 237Met Arg Gly
Pro Tyr1 52385PRTHomo sapiens 238Ile Tyr Glu Met Asn1 523917PRTHomo
sapiens 239Tyr Ile Thr Asn Arg Gly Ser Thr Ile Tyr Tyr Ala Asp Ser
Val Lys1 5 10 15Gly24010PRTHomo sapiens 240Pro Arg Ile Gly Ala Arg
Val Phe Asp Val1 5 1024117PRTHomo sapiens 241Lys Ser Ser Gln Thr
Leu Leu Tyr Lys Ser Asn Asn Glu Asn Tyr Leu1 5 10 15Ala2429PRTHomo
sapiens 242Gln Gln Tyr Phe Thr Thr Ala Leu Thr1 52435PRTHomo
sapiens 243Asp His Trp Ile Gly1 524417PRTHomo sapiens 244Ile Ile
Phe Pro Glu Asp Ser Asp Thr Arg Tyr Ser Gly Ser Phe Glu1 5 10
15Gly24512PRTHomo sapiens 245Val Ser Val Val Arg Lys Gly Gly Trp
Phe Asp Pro1 5 1024617PRTHomo sapiens 246Lys Ser Ser Gln Ser Leu
Leu Tyr Thr Ser Asn Asn Lys Asn Tyr Leu1 5 10 15Ala24717PRTHomo
sapiens 247Ile Ile Phe Pro Gly Asp Ser Asp Ile Arg Tyr Ser Pro Ser
Phe Glu1 5 10 15Gly24812PRTHomo sapiens 248Val Ala Val Val Arg Lys
Gly Gly Trp Phe Asp Ser1 5 1024917PRTHomo sapiens 249Lys Ser Thr
Gln Ser Leu Leu Trp Ser Ala Asn Asn Lys Asn Tyr Leu1 5 10
15Ala2505PRTHomo sapiens 250Ser Tyr Trp Ile Gly1 525117PRTHomo
sapiens 251Ile Ile Tyr Pro Asp Asp Ser Asp Thr Arg Tyr Asn Ala Ser
Leu Glu1 5 10 15Gly25217PRTHomo sapiens 252Arg Asp Arg Asn Cys Ser
Gly Thr Thr Cys Tyr Pro Arg Trp Phe Asp1 5 10 15Ser25317PRTHomo
sapiens 253Lys Ser Ser Gln Ser Leu Phe Tyr Ser Gly Asn Ser Lys Asp
Phe Leu1 5 10 15Ala2547PRTHomo sapiens 254Trp Ala Ser Thr Arg Asp
Ser1 52559PRTHomo sapiens 255His Gln Tyr His Ser Thr Pro Leu Ser1
5256296DNAHomo sapiens 256gaggtgcagc tggtggagtc cgggggaggc
ttagttcagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt
agctactgga tgcactgggt ccgccaagct 120ccagggaagg ggctggtgtg
ggtctcacgt attaatagtg atgggagtag cacaagctac 180gcggactccg
tgaagggccg attcaccatc tccagagaca acgccaagaa cacgctgtat
240ctgcaaatga acagtctgag agccgaggac acggctgtgt attactgtgc aagaga
296257107PRTHomo sapiens 257Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Ser Tyr 20 25 30Trp Met His Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Val Trp Val 35 40 45Ser Arg Ile Asn Ser Asp Gly
Ser Asp Thr Asn Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Phe
Ser Arg Asp Asn Ala Lys Asn Thr Leu Tyr65 70 75 80Leu Gln Met Thr
Ser Leu Arg Ala Glu Asp Thr Ala Ile Tyr Tyr Cys 85 90 95Thr Arg Gly
Arg Ser Tyr Gly Phe Phe Asp Tyr 100 105258288DNAHomo sapiens
258gaaattgtgt tgacgcagtc tccagccacc ctgtctttgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttagc agcagctact tagcctggta
ccagcagaaa 120cctggccagg ctcccaggct cctcatctat gatgcatcca
gcagggccac tggcatccca 180gacaggttca gtggcagtgg gtctgggaca
gacttcactc tcaccatcag cagactggag 240cctgaagatt ttgcagtcta
ttactgtcag cagcgtagca actggcat 28825998PRTHomo sapiens 259Glu Ile
Val Leu Thr Gln Ser Pro Asp Thr Leu Ser Leu Ser Pro Gly1 5 10 15Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ile Ile Ser Ser Asn 20 25
30Tyr Leu Ala Trp Tyr Gln Gln Gln Pro Gly Gln Ala Pro Arg Leu Leu
35 40 45Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe
Ser 50 55 60Gly Ser Gly Ser Ala Thr Asp Phe Thr Leu Thr Ile Ser Arg
Leu Glu65 70 75 80Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr
Gly Thr Ser Pro 85 90 95Arg Thr260296DNAHomo sapiens 260caggtgcagc
tggtggagtc tgggggaggc gtggtccagc ctgggaggtc cctgagactc 60tcctgtgcag
cgtctggatt caccttcagt agctatggca tgcactgggt ccgccaggct
120ccaggcaagg ggctggagtg ggtggcagtt atatggtatg atggaagtaa
taaatactat 180gcagactccg tgaagggccg attcaccatc tccagagaca
attccaagaa cacgctgtat 240ctgcaaatga acagcctgag agccgaggac
acggctgtgt attactgtgc gagaga 296261113PRTHomo sapiens 261Gln Val
Gln Leu Val Glu Ser Arg Gly Gly Val Val Gln Pro Gly Thr1 5 10 15Ser
Leu Arg Leu Ser Cys Lys Ala Ser Gly Phe Thr Phe Ser Thr Tyr 20 25
30Gly Met His Trp Val Arg Gln Ala Pro Gly Lys Ala Leu Glu Trp Val
35 40 45Ala Val Ile Trp Phe Asp Gly Asn Asn Lys Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr
Val Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Gly Glu Asp Thr Ala
Val Tyr Tyr Cys 85 90 95Ala Arg Asp Trp Trp Glu Ala Gly Cys Arg Pro
Cys Tyr Phe Phe Asp 100 105 110Tyr262305DNAHomo sapiens
262gacatcgtga tgacccagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccagcca gagtgtttta tacagctcca acaataagaa
ctacttagct 120tggtaccagc agaaaccagg acagcctcct aagctgctca
tttactgggc atctacccgg 180gaatccgggg tccctgaccg attcagtggc
agcgggtctg ggacagattt cactctcacc 240atcagcagcc tgcaggctga
agatgtggca gtttattact gtcagcaata ttatagtact 300cctcc
305263103PRTHomo sapiens 263Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Val Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser
Pro Pro Leu Thr 100264296DNAHomo sapiens 264gaggtgcagc tggtggagtc
tgggggaggc ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt
caccttcagt agctatgcta tgcactgggt ccgccaggct 120ccagggaagg
gactggaata tgtttcagct attagtagta atgggggtag cacatattat
180gcaaactctg tgaagggcag attcaccatc tccagagaca attccaagaa
cacgctgtat 240cttcaaatgg gcagcctgag agctgaggac atggctgtgt
attactgtgc gagaga 296265116PRTHomo sapiens 265Glu Val Gln Leu Val
Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu
Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asp Tyr 20 25 30Trp Met Ser
Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Asn
Ile Asn Gln Asp Gly Ser Ala Ala Tyr Tyr Val Asp Ser Val 50 55 60Arg
Gly Arg Phe Thr Ile Ser Arg Asp Ser Ala Glu Asn Ser Leu Asn65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Lys Asp Ala His Tyr Tyr Asp Arg Asn Arg Asn Asn Tyr Tyr
Tyr 100 105 110Tyr Phe Asp Phe 115266287DNAHomo sapiens
266gaaatagtga tgacgcagtc tccagccacc ctgtctgtgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttagc agcaacttag cctggtacca
gcagaaacct 120ggccaggctc ccaggctcct catctatggt gcatccacca
gggccactgg tatcccagcc 180aggttcagtg gcagtgggtc tgggacagag
ttcactctca ccatcagcag cctgcagtct 240gaagattttg cagtttatta
ctgtcagcag tataataact ggcctcc 28726797PRTHomo sapiens 267Glu Ile
Val Met Thr Gln Ser Pro Ala Thr Leu Ser Val Ser Pro Gly1 5 10 15Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Gly Ala Asn 20 25
30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45Tyr Ser Ala Ser Thr Arg Ala Thr Gly Val Pro Ala Arg Phe Ser
Gly 50 55 60Ser Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu
Gln Ser65 70 75 80Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Asn
Asn Trp Pro Arg 85 90 95Thr268291DNAHomo sapiens 268caggtgcagc
tgcaggagtc gggcccagga ctggtgaagc cttcacagac cctgtccctc 60acctgcactg
tctctggtgg ctccatcagc agtggtggtt actactggag ctggatccgc
120cagcacccag ggaagggcct ggagtggatt gggtacatct attacagtgg
gagcacctac 180tacaacccgt ccctcaagag tcgagttacc atatcagtag
acacgtctaa gaaccagttc 240tccctgaagc tgagctctgt gaccgcggac
gcggccgtgt attactgtgc g 291269116PRTHomo sapiens 269Gln Val Gln Leu
Gln Glu Ser Gly Pro Gly Leu Val Asn Pro Ser Gln1 5 10 15Thr Leu Ser
Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Gly 20 25 30Asn Tyr
Tyr Trp Ser Trp Ile Arg Gln Pro Ala Gly Lys Gly Leu Glu 35 40 45Trp
Ile Gly Arg Met Ser Ser Ser Gly Ser Thr Asn Tyr Asn Pro Ser 50 55
60Leu Lys Ser Arg Val Thr Met Ser Glu Asp Thr Ser Lys Asn Gln Phe65
70 75 80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr
Tyr 85 90 95Cys Ala Arg Glu Ser Gly Ser Ser Trp Gln Asn His Tyr Tyr
Tyr Tyr 100 105 110Gly Met Asp Val 115270103PRTHomo sapiens 270Asp
Ile Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10
15Glu Arg Ala Asn Ile Asn Cys Lys Ser Ser Gln Ser Val Leu Tyr Ser
20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Ser Gly
Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser
Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val
Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Thr Pro Leu Thr
100271296DNAHomo sapiens 271gaggtgcagc tggtggagtc tgggggaggc
ttggtacagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc
agctatgcca tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtctcagct attagtggta gtggtggtag cacatactac 180gcagactccg
tgaagggccg gttcaccatc tccagagaca attccaagaa cacgctgtat
240ctgcaaatga acagcctgag agccgaggac acggccgtat attactgtgc gaaaga
296272112PRTHomo sapiens 272Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Arg Asn Tyr 20 25 30Ala Met Ser Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Ser Asp Gly
Asn Thr Phe Tyr Ala Asp Ser Val Lys 50 55 60Gly Arg Phe Thr Val Ser
Arg Asp Asn Ser Lys Asn Thr Leu Tyr Leu65 70 75 80Gln Val Asn Ser
Leu Arg Ala Asp Asp Thr Ala Val Tyr Tyr Cys Ala 85 90 95Lys Glu Ser
Gly Arg Trp Gly Gly Gly Thr Leu Tyr Gly Ala His Tyr 100 105
110273103PRTHomo sapiens 273Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Leu Leu Tyr Asn 20 25 30Ser Asn Asn Lys Asn Tyr Leu Thr
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val
Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Ser Pro Leu Thr
100274296DNAHomo sapiens 274caggtgcagc tggtgcagtc tggggctgag
gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggata caccttcacc
ggctactata tgcactgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcaacccta acagtggtgg cacaaactat 180gcacagaagt
ttcagggcag ggtcaccatg accagggaca cgtccatcag cacagcctac
240atggagctga gcaggctgag atctgacgac acggccgtgt attactgtgc gagaga
296275110PRTHomo sapiens 275Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Pro Val Lys Val Ser Cys Glu Thr Ser
Gly Tyr Arg Phe Ser Asp Tyr 20 25 30Asn Val His Trp Val Arg Gln Ala
Pro Gly Gln Gly Pro Glu Trp Ile 35 40 45Gly Arg Ile Ser Pro Asn Ser
Gly Gly Thr Lys Tyr Ala Gln Lys Phe 50 55 60Gln Gly Arg Val Thr Met
Thr Arg Asp Met Ser Met Asn Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Gly Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys 85 90 95Val Arg Gly
His Cys Asp Gly Thr Thr Cys Ser Arg Ala Tyr 100 105
110276302DNAHomo sapiens 276gatattgtga tgactcagtc tccactctcc
ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca gagcctcctg
catagtaatg gatacaacta tttggattgg 120tacctgcaga agccagggca
gtctccacag ctcctgatct atttgggttc taatcgggcc 180tccggggtcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc
240agcagagtgg aggctgagga tgttggggtt tattactgca tgcaagctct
acaaactcct 300cc 302277102PRTHomo sapiens 277Asp Ile Val Met Thr
Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser
Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu His Arg 20 25 30Ser Gly His
Lys Tyr Leu His Trp Tyr Leu Gln Arg Pro Gly Gln Ser 35 40 45Pro Gln
Val Leu Ile Tyr Leu Gly Ser Asn Arg Ala Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75
80Ser Arg Val Glu Ala Glu Asp Val Gly Leu Tyr Tyr Cys Met Gln Thr
85 90 95Leu Gln Thr Pro Trp Thr 100278296DNAHomo sapiens
278caggtccagc ttgtgcagtc tggggctgag gtgaagaagc ctggggcctc
agtgaaggtt 60tcctgcaagg cttctggata caccttcact agctatgcta tgcattgggt
gcgccaggcc 120cccggacaaa ggcttgagtg gatgggatgg atcaacgctg
gcaatggtaa cacaaaatat 180tcacagaagt tccagggcag agtcaccatt
accagggaca catccgcgag cacagcctac 240atggagctga gcagcctgag
atctgaagac acggctgtgt attactgtgc gagaga 296279109PRTHomo sapiens
279Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Thr Ala Ser Gly Tyr Thr Phe Thr Gly
Tyr 20 25 30Tyr Leu His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Trp Val Asn Pro Arg Ser Gly Gly Thr Ser Tyr Pro
Pro Lys Phe 50 55 60Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Ile
Asn Thr Ala Tyr65 70 75 80Met Asp Leu Thr Trp Leu Thr Ser Asp Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Val Gly Arg Ile Pro Asp Val Thr
Ala Phe Asp Ile 100 105280103PRTHomo sapiens 280Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Glu Ser Leu Leu Tyr Asp 20 25 30Ser Asn Asn
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Glu Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr His Cys Gln Gln
85 90 95Tyr Phe Ser Thr Pro Trp Thr 100281296DNAHomo sapiens
281gaggtgcagc tggtgcagtc tggagcagag gtgaaaaagc ccggggagtc
tctgaagatc 60tcctgtaagg gttctggata cagctttacc agctactgga tcggctgggt
gcgccagatg 120cccgggaaag gcctggagtg gatggggatc atctatcctg
gtgactctga taccagatac 180agcccgtcct tccaaggcca ggtcaccatc
tcagccgaca agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa
ggcctcggac accgccatgt attactgtgc gagaca 296282111PRTHomo sapiens
282Glu Val Gln Leu Val Gln Ser Gly Pro Glu Met Arg Lys Pro Gly Glu1
5 10 15Ser Leu Lys Ile Ser Cys Lys Thr Ser Gly Tyr Ile Phe Ser Asp
Tyr 20 25 30Trp Thr Ala Trp Val Arg Gln Leu Pro Gly Lys Gly Leu Gln
Trp Met 35 40 45Gly Ile Ile Tyr Ser Gly Asp Ser Asp Thr Arg Tyr His
Pro Ser Val 50 55 60Gln Gly His Val Thr Met Ser Thr Asp Ser Ser Leu
Thr Thr Ala Tyr65 70 75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp
Thr Gly Ile Tyr Tyr Cys 85 90 95Ala Arg Leu Asp Ala Arg Val Asp Ala
Gly Trp Gln Leu Asp Ser 100 105 110283103PRTHomo sapiens 283Asp Ile
Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu
Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val Phe Ser Arg 20 25
30Asp Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln His Lys Ser Gly Gln
35 40 45Pro Pro Lys Leu Leu Phe Phe Trp Ala Ser Ser Arg Glu Ser Gly
Val 50 55 60Ser Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr
Leu Thr65 70 75 80Ile Asp Asn Leu Gln Ala Glu Asp Val Ala Leu Tyr
Tyr Cys Gln His 85 90 95Tyr Phe Asn Thr Pro His Asn
100284110PRTHomo sapiens 284Glu Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Glu1 5 10 15Ser Leu Lys Ile Ser Cys Glu Ala Ser
Gly Tyr Ser Phe Thr Asn Tyr 20 25 30Trp Ile Gly Trp Val Arg Gln Met
Pro Gly Lys Gly Leu Glu Trp Met 35 40 45Gly Ile Ile Tyr Pro Gly Asp
Ser Asp Thr Arg Tyr Ser Pro Pro Phe 50 55 60Gln Gly Gln Val Thr Ile
Thr Ala Asp Arg Ser Ile Thr Thr Ala Tyr65 70 75 80Leu Glu Trp Ser
Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Tyr Cys 85 90 95Ala Arg Val
Gly Arg Pro Ser Lys Gly Gly Trp Phe Asp Pro 100 105
110285103PRTHomo sapiens 285Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Glu Ser
Ser Gln Thr Leu Leu Tyr Ser 20 25 30Ser Asn Glu Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Ser
Trp Ala Ser Thr Pro Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Ser Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Asn
Ser Pro Tyr Thr 100286296DNAHomo sapiens 286caggtgcagc tggtggagtc
tgggggaggc ttggtcaagc ctggagggtc cctgagactc 60tcctgtgcag cctctggatt
caccttcagt gactactaca tgagctggat ccgccaggct 120ccagggaagg
ggctggagtg ggtttcatac attagtagta gtagtagtta cacaaactac
180gcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa
ctcactgtat 240ctgcaaatga acagcctgag agccgaggac acggctgtgt
attactgtgc gagaga 296287115PRTHomo sapiens 287Gln Val Gln Leu Val
Glu Ser Gly Gly Gly Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Arg Leu
Ser Cys Val Ala Ser Gly Ile Thr Phe Ser Asp Ser 20 25 30Tyr Met Ser
Trp Ile Arg Gln Thr Pro Gly Lys Gly Leu Glu Trp Leu 35 40 45Ser Tyr
Ile Ser Arg Ser Ser Ser His Thr Asn Tyr Ala Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Gly Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Val Gln Thr Thr Met Ile Glu Gly Lys Thr Lys Leu Asn
Tyr 100 105 110Phe Asp Tyr 115288103PRTHomo sapiens 288Asp Ile Val
Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg
Val Thr Ile Thr Cys Glu Ser Ser His Ser Leu Leu Tyr Arg 20 25 30Ser
Asn Asn Arg Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Arg Gln 35 40
45Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val
50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Glu Thr Asp Phe Thr Leu
Thr65 70 75 80Ile Ser Ser Leu Arg Ala Glu Asp Val Ala Val Tyr Tyr
Cys Gln Gln 85 90 95Phe Tyr Thr Thr Pro Tyr Thr 100289115PRTHomo
sapiens 289Gln Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Lys Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Val Ala Ser Gly Ile Thr Phe
Ser Asp Ser 20 25 30Tyr Met Ser Trp Ile Arg Gln Thr Pro Gly Lys Gly
Leu Glu Trp Leu 35 40 45Ser Tyr Ile Ser Arg Ser Ser Ser His Thr Asn
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Gly
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Val Gln Thr Thr Met
Ile Glu Gly Lys Thr Lys Leu Asn Tyr 100 105 110Phe Asp Tyr
115290103PRTHomo sapiens 290Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Val Thr Ile Ser Cys Glu Ser
Ser His Ser Leu Leu Tyr Arg 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Arg Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Glu Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Arg Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Phe Tyr Thr
Thr Pro Tyr Thr 100291302DNAHomo sapiens 291gaggtgcagc tggtggagtc
tgggggaggc ttggtaaagc ctggggggtc ccttagactc 60tcctgtgcag cctctggatt
cactttcagt aacgcctgga tgagctgggt ccgccaggct 120ccagggaagg
ggctggagtg ggttggccgt attaaaagca aaactgatgg tgggacaaca
180gactacgctg cacccgtgaa aggcagattc accatctcaa gagatgattc
aaaaaacacg 240ctgtatctgc aaatgaacag cctgaaaacc gaggacacag
ccgtgtatta ctgtaccaca 300ga 302292111PRTHomo sapiens 292Gln Leu Val
Glu Ser Gly Gly Gly Leu Ala Glu Pro Gly Gly Ser Leu1 5 10 15Arg Leu
Ser Cys Ala Ala Ser Gly Phe Thr Phe Thr Lys Ala Trp Met 20 25 30Ser
Trp Val Arg Gln Thr Pro Gly Arg Gly Leu Glu Trp Leu Gly Arg 35 40
45Ile Lys Ser Lys Val Asp Gly Glu Thr Thr Asp Tyr Ala Ala Pro Val
50 55 60Arg Gly Arg Phe Thr Ile Ser Arg Asp Asp Ser Lys Ser Thr Val
Tyr65 70 75 80Leu Gln Met Thr Gly Leu Arg Thr Glu Asp Thr Gly Val
Tyr Phe Cys 85 90 95Thr Thr Leu Ile His Cys Asp Leu Ser Ala Cys Leu
Pro His Phe 100 105 110293103PRTHomo sapiens 293Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Val Thr
Ile Ser Cys Glu Ser Ser His Ser Leu Leu Tyr Arg 20 25 30Ser Asn Asn
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Arg Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Glu Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Arg Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Phe Tyr Thr Thr Pro Tyr Thr 100294294DNAHomo sapiens
294gaggtgcagc tggtgcagtc tggagcagag gtgaaaaagc cgggggagtc
tctgaagatc 60tcctgtaagg gttctggata cagctttacc agctactgga tcggctgggt
gcgccagatg 120cccgggaaag gcctggagtg gatggggatc atctatcctg
gtgactctga taccagatac 180agcccgtcct tccaaggcca ggtcaccatc
tcagccgaca agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa
ggcctcggac accgccatgt attactgtgc gaga 294295113PRTHomo sapiens
295Glu Val Gln Leu Val Gln Ser Gly Gly Glu Val Lys Lys Pro Gly Glu1
5 10 15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Ser Asn
Tyr 20 25 30Trp Ile Ala Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu
Cys Met 35 40 45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Thr Tyr Ser
Pro Ser Phe 50 55 60Gln Gly Gln Val Thr Ile Ser Ala Asp Lys Ser Val
Ser Thr Thr Tyr65 70 75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp
Ser Ala Met Tyr Tyr Cys 85 90 95Ala Arg Leu Pro Arg Thr Asp Gly Asp
Asn Ser Ile Gly Tyr Phe Glu 100 105 110Tyr296103PRTHomo sapiens
296Asp Ile Val Met Thr Gln Ser Pro Ala Ser Leu Ala Val Ser Leu Gly1
5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val Leu Tyr
Ser 20 25 30Ser Asn Ser Glu Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro
Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu
Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala
Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Thr Pro Phe Thr
100297296DNAHomo sapiens 297gaggtgcagc tggtggagtc tgggggaggc
ttggtacagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt
agctatagca tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtttcatac attagtagta gtagtagtac catatactac 180gcagactctg
tgaagggccg attcaccatc tccagagaca atgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agccgaggac acggctgtgt attactgtgc gagaga
296298108PRTHomo sapiens 298Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Ser Tyr 20 25 30Ser Met Asn Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Tyr Ile Ser Ser Ser Thr
Thr Thr Ile Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Val Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met His
Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Val Pro Ala
Pro Arg Leu Gly Gly Ser Tyr Thr Tyr 100 105299290DNAHomo sapiens
299gaaattgtgt tgacgcagtc tccaggcacc ctgtctttgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttagc agcagctact tagcctggta
ccagcagaaa 120cctggccagg ctcccaggct cctcatctat ggtgcatcca
gcagggccac tggcatccca 180gacaggttca gtggcagtgg gtctgggaca
gacttcactc tcaccatcag cagactggag 240cctgaagatt ttgcagtgta
ttactgtcag cagtatggta gctcacctcc 29030098PRTHomo sapiens 300Glu Ile
Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly1 5 10 15Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Ser 20 25
30Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu
35 40 45Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe
Ser 50 55 60Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg
Leu Glu65 70 75 80Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr
Gly Thr
Ser Pro 85 90 95Leu Thr301302DNAHomo sapiens 301gaggtgcagc
tggtggagtc tgggggagcc ttggtaaagc ctggggggtc ccttagactc 60tcctgtgcag
cctctggatt cactttcagt aacgcctgga tgagctgggt ccgccaggct
120ccagggaagg ggctggagtg ggttggccgt attaaaagca aaactgatgg
tgggacaaca 180gactacgctg cacccgtgaa aggcagattc accatctcaa
gagatgattc aaaaaacacg 240ctgtatctgc aaatgaacag cctgaaaacc
gaggacacag ccgtgtatta ctgtaccaca 300ga 302302108PRTHomo sapiens
302Glu Val Gln Leu Val Glu Ser Gly Gly Asp Leu Val Lys Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Val Phe Thr Asp
Ala 20 25 30Trp Met Ser Trp Val Arg Gln Ser Pro Gly Lys Gly Pro Glu
Trp Leu 35 40 45Gly Arg Ile Lys Ser Lys Asn Val Gly Glu Thr Thr Asp
Tyr Ala Glu 50 55 60His Val Arg Gly Arg Phe Thr Ile Ala Arg Asp Asp
Ser Asn Arg Thr65 70 75 80Leu Tyr Leu Gln Met Ser Asn Leu Lys Ile
Asp Asp Thr Ala Val Tyr 85 90 95Tyr Cys Thr Thr Gly Leu Gly Gly Gly
Thr Tyr Gly 100 105303302DNAHomo sapiens 303gatgttgtga tgactcagtc
tccactctcc ctgcccgtca cccttggaca gccggcctcc 60atctcctgca ggtctagtca
aagcctcgta tacagtgatg gaaacaccta cttgaattgg 120tttcagcaga
ggccaggcca atctccaagg cgcctaattt ataaggtttc taaccgggac
180tctggggtcc cagacagatt cagcggcagt gggtcaggca ctgatttcac
actgaaaatc 240agcagggtgg aggctgagga tgttggggtt tattactgca
tgcaaggtac acactggcct 300cc 30230498PRTHomo sapiens 304Asp Val Val
Met Thr Gln Ser Pro Leu Ser Leu Pro Ala Thr Leu Gly1 5 10 15Gln Pro
Ala Ser Ile Ser Cys Arg Ser Ser Ala Gly Leu Arg Asn Asn 20 25 30Asp
Gly Asp Ile Leu Leu Ser Trp Phe His Gln Arg Pro Gly Gln Ser 35 40
45Pro Arg Arg Leu Phe Tyr Arg Val Ser Arg Arg Asp Ser Gly Val Pro
50 55 60Asp Arg Phe Asn Gly Ser Gly Ser Ala Thr Asp Phe Thr Leu Arg
Ile65 70 75 80Asn Ser Val Glu Ala Glu Asp Val Gly Ile Tyr Tyr Cys
Met Arg Gly 85 90 95Pro Tyr305296DNAHomo sapiens 305gaggtgcagc
tggtggagtc tgggggaggc ttggtacagc ctggagggtc cctgagactc 60tcctgtgcag
cctctggatt caccttcagt agttatgaaa tgaactgggt ccgccaggct
120ccagggaagg ggctggagtg ggtttcatac attagtagta gtggtagtac
catatactac 180gcagactctg tgaagggccg attcaccatc tccagagaca
acgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agccgaggac
acggctgttt attactgtgc gagaga 296306108PRTHomo sapiens 306Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Glu1 5 10 15Ser
Leu Arg Leu Ser Cys Val Ala Ser Gly Phe Asn Phe Ser Ile Tyr 20 25
30Glu Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Ser Tyr Ile Thr Asn Arg Gly Ser Thr Ile Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Gln Asn Ser
Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala
Val Tyr Tyr Cys 85 90 95Ala Lys Pro Arg Ile Gly Ala Arg Val Phe Asp
Val 100 105307103PRTHomo sapiens 307Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Thr Leu Leu Tyr Lys 20 25 30Ser Asn Asn Glu Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr
Phe Thr Thr Ala Leu Thr 100308296DNAHomo sapiens 308gaggtgcagc
tggtgcagtc tggagcagag gtgaaaaagc ccggggagtc tctgaagatc 60tcctgtaagg
gttctggata cagctttacc agctactgga tcggctgggt gcgccagatg
120cccgggaaag gcctggagtg gatggggatc atctatcctg gtgactctga
taccagatac 180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca
agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa ggcctcggac
accgccatgt attactgtgc gagaca 296309110PRTHomo sapiens 309Glu Val
Gln Leu Val Gln Ser Gly Ala Val Val Lys Lys Pro Gly Glu1 5 10 15Ser
Leu Lys Ile Ser Cys Glu Ala Ser Gly Tyr Met Phe Leu Asp His 20 25
30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu Trp Met
35 40 45Gly Ile Ile Phe Pro Glu Asp Ser Asp Thr Arg Tyr Ser Gly Ser
Phe 50 55 60Glu Gly Gln Val Thr Ile Ser Ala Asp Arg Ser Val Asn Thr
Val Tyr65 70 75 80Leu Glu Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala
Met Tyr Tyr Cys 85 90 95Ala Arg Val Ser Val Val Arg Lys Gly Gly Trp
Phe Asp Pro 100 105 110310103PRTHomo sapiens 310Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr Thr 20 25 30Ser Asn Asn
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Thr Pro
Lys Leu Leu Ile Ile Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro
Asp Arg Phe Thr Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Tyr Tyr Asn Ser Pro Tyr Thr 100311110PRTHomo sapiens 311Glu
Val Gln Leu Val Gln Ser Gly Ala Glu Leu Lys Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Glu Ala Ser Gly Tyr Ile Phe Ala Asp His
20 25 30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu Trp
Met 35 40 45Gly Ile Ile Phe Pro Gly Asp Ser Asp Ile Arg Tyr Ser Pro
Ser Phe 50 55 60Glu Gly Gln Val Thr Ile Ser Val Asp Arg Ser Val Ser
Thr Ala Phe65 70 75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Phe Cys 85 90 95Ala Arg Val Ala Val Val Arg Lys Gly Gly
Trp Phe Asp Ser 100 105 110312103PRTHomo sapiens 312Asp Ile Val Met
Thr Gln Ser Pro Glu Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala
Thr Ile Asn Cys Lys Ser Thr Gln Ser Leu Leu Trp Ser 20 25 30Ala Asn
Asn Lys Asn Tyr Leu Ala Trp Tyr Arg Gln Lys Pro Arg Gln 35 40 45Thr
Pro Glu Leu Leu Ile Thr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55
60Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65
70 75 80Ile Thr Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln
Gln 85 90 95Tyr Tyr Asn Ser Pro Tyr Thr 100313115PRTHomo sapiens
313Glu Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Trp Glu1
5 10 15Ser Leu Lys Ile Ser Cys Leu Ala Ser Gly Tyr Asp Phe Ala Ser
Tyr 20 25 30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu
Trp Val 35 40 45Gly Ile Ile Tyr Pro Asp Asp Ser Asp Thr Arg Tyr Asn
Ala Ser Leu 50 55 60Glu Gly Arg Val Thr Met Ser Val Asp Thr Ser Thr
Asn Thr Ala Tyr65 70 75 80Leu Gln Trp Thr Ser Leu Lys Val Ser Asp
Thr Gly Met Tyr Tyr Cys 85 90 95Ala Arg Arg Asp Arg Asn Cys Ser Gly
Thr Thr Cys Tyr Pro Arg Trp 100 105 110Phe Asp Ser 115314103PRTHomo
sapiens 314Asp Ile Val Met Thr Gln Ser Pro Asp Phe Leu Ala Val Ser
Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Leu
Phe Tyr Ser 20 25 30Gly Asn Ser Lys Asp Phe Leu Ala Trp Tyr Gln Gln
Lys Pro Gly Gln 35 40 45Pro Pro Arg Leu Leu Val Tyr Trp Ala Ser Thr
Arg Asp Ser Gly Val 50 55 60Pro Glu Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Arg Leu Gln Ala Glu Asp
Val Ala Leu Tyr Tyr Cys His Gln 85 90 95Tyr His Ser Thr Pro Leu Ser
10031519PRTartificialsynthetic
constructMOD_RES(9)..(9)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION
315Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg1
5 10 15Ser Arg Thr31619PRTartificialsynthetic construct 316Arg Ser
Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg1 5 10 15Ser
Arg Thr31718PRTartificialsynthetic
constructMOD_RES(9)..(9)PHOSPHORYLATIONMOD_RES(11)..(11)PHOSPHORYLATION
317Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro Ser Leu Pro Thr Pro Pro1
5 10 15Thr Arg31818PRTartificialsynthetic construct 318Gly Thr Pro
Gly Ser Arg Ser Arg Thr Pro Ser Leu Pro Thr Pro Pro1 5 10 15Thr
Arg31918PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 319Pro Gly Ser Pro Gly
Thr Pro Gly Ser Arg Ser Arg Thr Pro Ser Leu1 5 10 15Pro
Thr32018PRTartificialsynthetic construct 320Pro Gly Ser Pro Gly Thr
Pro Gly Ser Arg Ser Arg Thr Pro Ser Leu1 5 10 15Pro
Thr32168PRTartificialsynthetic construct 321Gly Glu Pro Pro Lys Ser
Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly1 5 10 15Ser Pro Gly Thr Pro
Gly Ser Arg Ser Arg Thr Pro Ser Leu Pro Thr 20 25 30Pro Pro Thr Arg
Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro 35 40 45Lys Ser Pro
Ser Ser Ala Lys Ser Arg Leu Gln Thr Ala Pro Val Pro 50 55 60Met Pro
Asp Leu6532219PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION
322Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys Ser Thr Pro Thr Ala Glu1
5 10 15Asp Val Thr32320PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION-
MOD_RES(13)..(13)PHOSPHORYLATION 323Pro Gly Ser Glu Thr Ser Asp Ala
Lys Ser Thr Pro Thr Ala Glu Asp1 5 10 15Val Thr Ala Pro
2032418PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 324Ser Glu Thr Ser Asp
Ala Lys Ser Thr Pro Thr Ala Glu Asp Val Thr1 5 10 15Ala
Pro32562PRTartificialsynthetic construct 325Gly Leu Lys Glu Ser Pro
Leu Gln Thr Pro Thr Glu Asp Gly Ser Glu1 5 10 15Glu Pro Gly Ser Glu
Thr Ser Asp Ala Lys Ser Thr Pro Thr Ala Glu 20 25 30Asp Val Thr Ala
Pro Leu Val Asp Glu Gly Ala Pro Gly Lys Gln Ala 35 40 45Ala Ala Gln
Pro His Thr Glu Ile Pro Glu Gly Thr Thr Ala 50 55
6032624PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
326Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2032753PRTartificialsynthetic construct 327Gly Ala Glu Ile Val Tyr
Lys Ser Pro Val Val Ser Gly Asp Thr Ser1 5 10 15Pro Arg His Leu Ser
Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val 20 25 30Asp Ser Pro Gln
Leu Ala Thr Leu Ala Asp Glu Val Ser Ala Ser Leu 35 40 45Ala Lys Gln
Gly Leu 5032825PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
328Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2532918PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 329Val Val Arg Thr Pro
Pro Lys Ser Pro Ser Ser Ala Lys Ser Arg Leu1 5 10 15Gln
Thr33021PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(14)..(14)PHOSPHORYLATION
330Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala Lys1
5 10 15Ser Arg Leu Gln Thr 2033171PRTartificialsynthetic construct
331His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly Ser Leu Gly Asn Ile His
His 20 25 30Lys Pro Gly Gly Gly Gln Val Glu Val Lys Ser Glu Lys Leu
Asp Phe 35 40 45Lys Asp Arg Val Gln Ser Lys Ile Gly Ser Leu Asp Asn
Ile Thr His 50 55 60Val Pro Gly Gly Gly Asn Lys65
7033216PRTartificialsynthetic
constructMOD_RES(6)..(6)PHOSPHORYLATION 332Lys Ser Lys Ile Gly Ser
Thr Glu Asn Leu Lys His Gln Pro Gly Gly1 5 10
1533318PRTartificialsynthetic
constructMOD_RES(8)..(8)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION
333Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala1
5 10 15Lys Ser33426PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
334Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2533518PRTartificialsynthetic
constructPhosphorylation(11)..(12)Phosphorylation of serine at
amino acid 11MOD_RES(11)..(11)PHOSPHORYLATION 335Pro Pro Lys Ser
Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro1 5 10 15Gly
Thr33619PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION
336Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro1
5 10 15Gly Thr Pro33722PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(15)..(15)PHOSPHORYLATION
337Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro1
5 10 15Gly Thr Pro Gly Ser Arg 2033825PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
338Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro1
5 10 15Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2533918PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 339Pro Lys Ser Gly Asp
Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly1 5 10 15Thr
Pro34021PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(14)..(14)PHOSPHORYLATION
340Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly1
5 10 15Thr Pro Gly Ser Arg 2034124PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(14)..(14)PHOSPHORYLATION
341Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly1
5 10 15Thr Pro Gly Ser Arg Ser Arg Thr
2034220PRTartificialsynthetic construct 342Lys Ser Gly Asp Arg Ser
Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr1 5 10 15Pro Gly Ser Arg
2034318PRTartificialsynthetic constructMOD_RES(11)..(11) 343Gly Asp
Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly1 5 10 15Ser
Arg34421PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(14)..(14)PHOSPHORYLATION
344Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly1
5 10 15Ser Arg Ser Arg Thr 2034524PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
345Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly1
5 10 15Ser Arg Ser Arg Thr Pro Ser Leu
2034626PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
346Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly1
5 10 15Ser Arg Ser Arg Thr Pro Ser Leu Pro Thr 20
2534718PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 347Ser Gly Tyr Ser Ser
Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser1 5 10 15Arg
Thr34821PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(14)..(14)PHOSPHORYLATION
348Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser1
5 10 15Arg Thr Pro Ser Leu 2034923PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(16)..(16)PHOSPHORYLATION
349Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser1
5 10 15Arg Thr Pro Ser Leu Pro Thr 2035025PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
350Ser Gly Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser1
5 10 15Arg Thr Pro Ser Leu Pro Thr Pro Pro 20
2535120PRTartificialsynthetic construct 351Ser Gly Tyr Ser Ser Pro
Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser1 5 10 15Arg Thr Pro Ser
2035218PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 352Ser Ser Pro Gly Ser
Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro1 5 10 15Ser
Leu35320PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(13)..(13)PHOSPHORYLATION
353Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro1
5 10 15Ser Leu Pro Thr 2035422PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(15)..(15)PHOSPHORYLATION
354Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro1
5 10 15Ser Leu Pro Thr Pro Pro 2035524PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
355Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro1
5 10 15Ser Leu Pro Thr Pro Pro Thr Arg
2035620PRTartificialsynthetic construct 356Ser Ser Pro Gly Ser Pro
Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro1 5 10 15Ser Leu Pro Thr
2035720PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(13)..(13)PHOSPHORYLATION
357Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro Ser Leu1
5 10 15Pro Thr Pro Pro 2035822PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(15)..(15)PHOSPHORYLATION
358Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro Ser Leu1
5 10 15Pro Thr Pro Pro Thr Arg 2035925PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
359Pro Gly Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro Ser Leu1
5 10 15Pro Thr Pro Pro Thr Arg Glu Pro Lys 20
2536018PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 360Ser Pro Arg His Leu
Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met1 5 10 15Val
Asp36126PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
361Ser Pro Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met1
5 10 15Val Asp Ser Pro Gln Leu Ala Thr Leu Ala 20
2536219PRTartificialsynthetic construct 362Pro Arg His Leu Ser Asn
Val Ser Ser Thr Gly Ser Ile Asp Met Val1 5 10 15Asp Ser
Pro36318PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 363Arg His Leu Ser Asn
Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1 5 10 15Ser
Pro36420PRTartificialsynthetic construct 364Ser Asn Val Ser Ser Thr
Gly Ser Ile Asp Met Val Asp Ser Pro Gln1 5 10 15Leu Ala Thr Leu
2036518PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 365Ser Ser Thr Gly Ser
Ile Asp Met Val Asp Ser Pro Gln Leu Ala Thr1 5 10 15Leu
Ala36623PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(16)..(16)PHOSPHORYLATION
366Ser Ser Thr Gly Ser Ile Asp Met Val Asp Ser Pro Gln Leu Ala Thr1
5 10 15Leu Ala Asp Glu Val Ser Ala 2036768PRTartificialsynthetic
construct 367Gly Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser
Ser Pro Gly1 5 10 15Ser Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr Pro
Ser Leu Pro Thr 20 25 30Pro Pro Thr Arg Glu Pro Lys Lys Val Ala Val
Val Arg Thr Pro Pro 35 40 45Lys Ser Pro Ser Ser Ala Lys Ser Arg Leu
Gln Thr Ala Pro Val Pro 50 55 60Met Pro Asp
Leu6536818PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 368Arg Glu Pro Lys Lys
Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1 5 10 15Ser
Ser36922PRTartificialsynthetic
constructMOD_RES(15)..(15)PHOSPHORYLATION 369Arg Glu Pro Lys Lys
Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1 5 10 15Ser Ser Ala Lys
Ser Arg 2037024PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
370Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln
2037118PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 371Lys Val Ala Val Val
Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala Lys1 5 10 15Ser
Arg37220PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(13)..(13)PHOSPHORYLATION
372Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala Lys1
5 10 15Ser Arg Leu Gln 2037318PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 373Ala Val Val Arg Thr
Pro Pro Lys Ser Pro Ser Ser Ala Lys Ser Arg1 5 10 15Leu
Gln37419PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION
374Ala Val Val Arg Thr Pro Pro Lys Ser Pro Ser Ser Ala Lys Ser Arg1
5 10 15Leu Gln Thr37520PRTartificialsynthetic construct 375His Val
Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu
Ser Lys Val 2037620PRTartificialsynthetic construct 376Val Tyr Lys
Pro Val Asp Leu Ser Lys Val Thr Ser Lys Cys Gly Ser1 5 10 15Leu Gly
Asn Ile 2037720PRTartificialsynthetic construct 377Thr Ser Lys Cys
Gly Ser Leu Gly Asn Ile His His Lys Pro Gly Gly1 5 10 15Gly Gln Val
Glu 2037820PRTartificialsynthetic construct 378His His Lys Pro Gly
Gly Gly Gln Val Glu Val Lys Ser Glu Lys Leu1 5 10 15Asp Phe Lys Asp
2037920PRTartificialsynthetic construct 379Val Lys Ser Glu Lys Leu
Asp Phe Lys Asp Arg Val Gln Ser Lys Ile1 5 10 15Gly Ser Leu Asp
2038021PRTartificialsynthetic construct 380Arg Val Gln Ser Lys Ile
Gly Ser Leu Asp Asn Ile Thr His Val Pro1 5 10 15Gly Gly Gly Asn Lys
2038120PRTartificialsynthetic construct 381Gly Leu Lys Glu Ser Pro
Leu Gln Thr Pro Thr Glu Asp Gly Ser Glu1 5 10 15Glu Pro Gly Ser
2038220PRTartificialsynthetic construct 382Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2038318PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 383Glu Pro Gly Ser Glu
Thr Ser Asp Ala Lys Ser Thr Pro Thr Ala Glu1 5 10 15Asp
Val38420PRTartificialsynthetic construct 384Glu Thr Ser Asp Ala Lys
Ser Thr Pro Thr Ala Glu Asp Val Thr Ala1 5 10 15Pro Leu Val Asp
2038520PRTartificialsynthetic construct 385Ala Glu Asp Val Thr Ala
Pro Leu Val Asp Glu Gly Ala Pro Gly Lys1 5 10 15Gln Ala Ala Ala
2038622PRTartificialsynthetic construct 386Glu Gly Ala Pro Gly Lys
Gln Ala Ala Ala Gln Pro His Thr Glu Ile1 5 10 15Pro Glu Gly Thr Thr
Ala 2038720PRTartificialsynthetic construct 387Gly Asp Arg Ser Gly
Tyr Ser Ser Pro Gly Ser Pro Gly Thr Pro Gly1 5 10 15Ser Arg Ser Arg
2038826PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
388Ala Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2538926PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
389Glu Ala Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539026PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
390Glu Pro Ala Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539126PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
391Glu Pro Pro Ala Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539226PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
392Glu Pro Pro Lys Ala Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539326PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
393Glu Pro Pro Lys Ser Ala Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539426PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
394Glu Pro Pro Lys Ser Gly Ala Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539526PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
395Glu Pro Pro Lys Ser Gly Asp Ala Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539626PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
396Glu Pro Pro Lys Ser Gly Asp Arg Ala Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539726PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
397Glu Pro Pro Lys Ser Gly Asp Arg Ser Ala Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539826PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
398Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Ala Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2539926PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
399Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ala Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2540026PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
400Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ala Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2540126PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
401Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Ala Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2540226PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
402Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Ala Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2540326PRTartificialsynthetic
constructMOD_RES(19)..(19)PHOSPHORYLATION 403Glu Pro Pro Lys Ser
Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ala1 5 10 15Pro Gly Thr Pro
Gly Ser Arg Ser Arg Thr 20 2540426PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
404Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Ala Gly Thr Pro Gly Ser Arg Ser Arg Thr 20
2540526PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
405Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Ala Thr Pro Gly Ser Arg Ser Arg Thr 20
2540626PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATION 406Glu Pro Pro Lys Ser
Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1 5 10 15Pro Gly Ala Pro
Gly Ser Arg Ser Arg Thr 20 2540726PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
407Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Ala Gly Ser
Arg Ser Arg Thr 20 2540826PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
408Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Ala Ser Arg Ser Arg Thr 20
2540926PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
409Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ala Arg Ser Arg Thr 20
2541026PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
410Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Ala Ser Arg Thr 20
2541126PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
411Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ala Arg Thr 20
2541226PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
412Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Ala Thr 20
2541326PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(19)..(19)PHOSPHORYLATION
413Glu Pro Pro Lys Ser Gly Asp Arg Ser Gly Tyr Ser Ser Pro Gly Ser1
5 10 15Pro Gly Thr Pro Gly Ser Arg Ser Arg Ala 20
2541424PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
414Ala His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2041524PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
415Arg Ala Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2041624PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
416Arg His Ala Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2041724PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
417Arg His Leu Ala Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2041824PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
418Arg His Leu Ser Ala Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2041924PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
419Arg His Leu Ser Asn Ala Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042024PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
420Arg His Leu Ser Asn Val Ala Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042124PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
421Arg His Leu Ser Asn Val Ser Ala Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042224PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
422Arg His Leu Ser Asn Val Ser Ser Ala Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042324PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
423Arg His Leu Ser Asn Val Ser Ser Thr Ala Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042424PRTartificialsynthetic
constructMOD_RES(17)..(17)PHOSPHORYLATION 424Arg His Leu Ser Asn
Val Ser Ser Thr Gly Ala Ile Asp Met Val Asp1 5 10 15Ser Pro Gln Leu
Ala Thr Leu Ala 2042524PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
425Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ala Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042624PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
426Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Ala Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042724PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
427Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Ala Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042824PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
428Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Ala Asp1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2042924PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
429Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Ala1
5 10 15Ser Pro Gln Leu Ala Thr Leu Ala
2043024PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 430Arg His Leu Ser Asn
Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1 5 10 15Ala Pro Gln Leu
Ala Thr Leu Ala 2043124PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
431Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Ala Gln Leu Ala Thr Leu Ala
2043224PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
432Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Ala Leu Ala Thr Leu Ala
2043324PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
433Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Ala Ala Thr Leu Ala
2043424PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
434Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Ala Leu Ala
2043524PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
435Arg His Leu Ser Asn Val Ser Ser Thr Gly Ser Ile Asp Met Val Asp1
5 10 15Ser Pro Gln Leu Ala Thr Ala Ala
2043625PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
436Arg Ala Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2543725PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
437Arg Glu Ala Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2543825PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
438Arg Glu Pro Ala Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2543925PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
439Arg Glu Pro Lys Ala Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Pro Ser Ala Lys Ser Arg Leu Gln Thr 20
2544025PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
440Arg Glu Pro Lys Lys Ala Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544125PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
441Arg Glu Pro Lys Lys Val Ala Ala Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544225PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
442Arg Glu Pro Lys Lys Val Ala Val Ala Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544325PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
443Arg Glu Pro Lys Lys Val Ala Val Val Ala Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544425PRTartificialsynthetic
constructMOD_RES(18)..(18)PHOSPHORYLATION 444Arg Glu Pro Lys Lys
Val Ala Val Val Arg Ala Pro Pro Lys Ser Pro1 5 10 15Ser Ser Ala Lys
Ser Arg Leu Gln Thr 20 2544525PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
445Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Ala Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544625PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
446Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Ala1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544725PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
447Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Ala Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544825PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
448Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ala Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2544925PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
449Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Ala1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Thr 20
2545025PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
450Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ala Ser Ala Lys Ser Arg Leu Gln Thr 20
2545125PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATION 451Arg Glu Pro Lys Lys
Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1 5 10 15Ser Ala Ala Lys
Ser Arg Leu Gln Thr 20 2545225PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
452Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Ala Ser Arg Leu Gln Thr 20
2545325PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
453Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ala Arg Leu Gln Thr 20
2545425PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
454Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Ala Leu Gln Thr 20
2545525PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
455Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Ala Gln Thr 20
2545625PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
456Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Ala Thr 20
2545725PRTartificialsynthetic
constructMOD_RES(11)..(11)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
457Arg Glu Pro Lys Lys Val Ala Val Val Arg Thr Pro Pro Lys Ser Pro1
5 10 15Ser Ser Ala Lys Ser Arg Leu Gln Ala 20
2545825PRTartificialsynthetic construct 458His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2545925PRTartificialsynthetic construct 459Ala
Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2546025PRTartificialsynthetic construct 460His Ala Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2546125PRTartificialsynthetic construct 461His
Val Ala Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2546225PRTartificialsynthetic construct 462His Val Pro Ala Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2546325PRTartificialsynthetic construct 463His
Val Pro Gly Ala Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2546425PRTartificialsynthetic construct 464His Val Pro Gly Gly Ala
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2546525PRTartificialsynthetic construct 465His
Val Pro Gly Gly Gly Ala Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2546625PRTartificialsynthetic construct 466His Val Pro Gly Gly Gly
Ser Ala Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2546725PRTartificialsynthetic construct 467His
Val Pro Gly Gly Gly Ser Val Ala Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2546825PRTartificialsynthetic construct 468His Val Pro Gly Gly Gly
Ser Val Gln Ala Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2546925PRTartificialsynthetic construct 469His
Val Pro Gly Gly Gly Ser Val Gln Ile Ala Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2547025PRTartificialsynthetic construct 470His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Ala Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2547125PRTartificialsynthetic construct 471His
Val Pro Gly Gly Gly Ser
Val Gln Ile Val Tyr Ala Pro Val Asp1 5 10 15Leu Ser Lys Val Thr Ser
Lys Cys Gly 20 2547225PRTartificialsynthetic construct 472His Val
Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Ala Val Asp1 5 10 15Leu
Ser Lys Val Thr Ser Lys Cys Gly 20 2547325PRTartificialsynthetic
construct 473His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys
Pro Ala Asp1 5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2547425PRTartificialsynthetic construct 474His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Ala1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2547525PRTartificialsynthetic construct 475His
Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Ala Ser Lys Val Thr Ser Lys Cys Gly 20
2547625PRTartificialsynthetic construct 476His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ala Lys Val Thr
Ser Lys Cys Gly 20 2547725PRTartificialsynthetic construct 477His
Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Ala Val Thr Ser Lys Cys Gly 20
2547825PRTartificialsynthetic construct 478His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Ala Thr
Ser Lys Cys Gly 20 2547925PRTartificialsynthetic construct 479His
Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Ala Ser Lys Cys Gly 20
2548025PRTartificialsynthetic construct 480His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ala Lys Cys Gly 20 2548125PRTartificialsynthetic construct 481His
Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Ala Cys Gly 20
2548225PRTartificialsynthetic construct 482His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Ala Gly 20 2548325PRTartificialsynthetic construct 483His
Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10
15Leu Ser Lys Val Thr Ser Lys Cys Ala 20
2548420PRTartificialsynthetic construct 484Ala Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2048520PRTartificialsynthetic construct 485Thr Ala Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2048620PRTartificialsynthetic construct 486Thr Glu Ala Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2048720PRTartificialsynthetic construct 487Thr Glu Asp Ala Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2048820PRTartificialsynthetic construct 488Thr Glu Asp Gly Ala Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2048920PRTartificialsynthetic construct 489Thr Glu Asp Gly Ser Ala
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049020PRTartificialsynthetic construct 490Thr Glu Asp Gly Ser Glu
Ala Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049120PRTartificialsynthetic construct 491Thr Glu Asp Gly Ser Glu
Glu Ala Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049220PRTartificialsynthetic construct 492Thr Glu Asp Gly Ser Glu
Glu Pro Ala Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049320PRTartificialsynthetic construct 493Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ala Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049420PRTartificialsynthetic construct 494Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Ala Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049520PRTartificialsynthetic construct 495Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Ala Ser Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049620PRTartificialsynthetic construct 496Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ala Asp Ala Lys1 5 10 15Ser Thr Pro Thr
2049720PRTartificialsynthetic construct 497Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Ala Ala Lys1 5 10 15Ser Thr Pro Thr
2049820PRTartificialsynthetic construct 498Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Ala1 5 10 15Ser Thr Pro Thr
2049920PRTartificialsynthetic construct 499Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ala Thr Pro Thr
2050020PRTartificialsynthetic construct 500Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Ala Pro Thr
2050120PRTartificialsynthetic construct 501Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Ala Thr
2050220PRTartificialsynthetic construct 502Thr Glu Asp Gly Ser Glu
Glu Pro Gly Ser Glu Thr Ser Asp Ala Lys1 5 10 15Ser Thr Pro Ala
2050325PRTartificialsynthetic
constructMOD_RES(7)..(7)PHOSPHORYLATION 503His Val Pro Gly Gly Gly
Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val Thr
Ser Lys Cys Gly 20 2550425PRTartificialsynthetic
constructMOD_RES(12)..(12)PHOSPHORYLATION 504His Val Pro Gly Gly
Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val
Thr Ser Lys Cys Gly 20 2550525PRTartificialsynthetic
constructMOD_RES(18)..(18)PHOSPHORYLATION 505His Val Pro Gly Gly
Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val
Thr Ser Lys Cys Gly 20 2550625PRTartificialsynthetic
constructMOD_RES(21)..(21)PHOSPHORYLATION 506His Val Pro Gly Gly
Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val
Thr Ser Lys Cys Gly 20 2550725PRTartificialsynthetic
constructMOD_RES(22)..(22)PHOSPHORYLATION 507His Val Pro Gly Gly
Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1 5 10 15Leu Ser Lys Val
Thr Ser Lys Cys Gly 20 2550825PRTartificialsynthetic
constructMOD_RES(7)..(7)PHOSPHORYLATIONMOD_RES(12)..(12)PHOSPHORYLATION
508His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2550925PRTartificialsynthetic
constructMOD_RES(7)..(7)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
509His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551025PRTartificialsynthetic
constructMOD_RES(7)..(7)PHOSPHORYLATIONMOD_RES(21)..(21)PHOSPHORYLATION
510His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551125PRTartificialsynthetic
constructMOD_RES(7)..(7)PHOSPHORYLATIONMOD_RES(22)..(22)PHOSPHORYLATION
511His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551225PRTartificialsynthetic
constructMOD_RES(12)..(12)PHOSPHORYLATIONMOD_RES(18)..(18)PHOSPHORYLATION
512His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551325PRTartificialsynthetic
constructMOD_RES(12)..(12)PHOSPHORYLATIONMOD_RES(21)..(21)PHOSPHORYLATION
513His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551425PRTartificialsynthetic
constructMOD_RES(12)..(12)PHOSPHORYLATIONMOD_RES(22)..(22)PHOSPHORYLATION
514His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551525PRTartificialsynthetic
constructMOD_RES(18)..(18)PHOSPHORYLATIONMOD_RES(21)..(21)PHOSPHORYLATION
515His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551625PRTartificialsynthetic
constructMOD_RES(18)..(18)PHOSPHORYLATIONMOD_RES(22)..(22)PHOSPHORYLATION
516His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551725PRTartificialsynthetic
constructMOD_RES(21)..(21)PHOSPHORYLATIONMOD_RES(22)..(22)PHOSPHORYLATION
517His Val Pro Gly Gly Gly Ser Val Gln Ile Val Tyr Lys Pro Val Asp1
5 10 15Leu Ser Lys Val Thr Ser Lys Cys Gly 20
2551824PRTartificialsynthetic construct 518Leu Gln Thr Pro Thr Glu
Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys Ser
Thr Pro Thr 2051924PRTartificialsynthetic
constructMOD_RES(24)..(24)PHOSPHORYLATION 519Leu Gln Thr Pro Thr
Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys
Ser Thr Pro Thr 2052024PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATION 520Leu Gln Thr Pro Thr
Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys
Ser Thr Pro Thr 2052124PRTartificialsynthetic
constructMOD_RES(14)..(14)PHOSPHORYLATION 521Leu Gln Thr Pro Thr
Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys
Ser Thr Pro Thr 2052224PRTartificialsynthetic
constructMOD_RES(9)..(9)PHOSPHORYLATION 522Leu Gln Thr Pro Thr Glu
Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys Ser
Thr Pro Thr 2052324PRTartificialsynthetic
constructMOD_RES(5)..(5)PHOSPHORYLATION 523Leu Gln Thr Pro Thr Glu
Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys Ser
Thr Pro Thr 2052424PRTartificialsynthetic
constructMOD_RES(21)..(21)PHOSPHORYLATION 524Leu Gln Thr Pro Thr
Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys
Ser Thr Pro Thr 2052524PRTartificialsynthetic
constructMOD_RES(22)..(22)PHOSPHORYLATION 525Leu Gln Thr Pro Thr
Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys
Ser Thr Pro Thr 2052624PRTartificialsynthetic
constructMOD_RES(17)..(17)PHOSPHORYLATION 526Leu Gln Thr Pro Thr
Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1 5 10 15Ser Asp Ala Lys
Ser Thr Pro Thr 2052724PRTartificialsynthetic
constructMOD_RES(14)..(14)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
527Leu Gln Thr Pro Thr Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1
5 10 15Ser Asp Ala Lys Ser Thr Pro Thr
2052824PRTartificialsynthetic
constructMOD_RES(14)..(14)PHOSPHORYLATIONMOD_RES(16)..(16)PHOSPHORYLATION
528Leu Gln Thr Pro Thr Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1
5 10 15Ser Asp Ala Lys Ser Thr Pro Thr
2052924PRTartificialsynthetic
constructMOD_RES(16)..(16)PHOSPHORYLATIONMOD_RES(17)..(17)PHOSPHORYLATION
529Leu Gln Thr Pro Thr Glu Asp Gly Ser Glu Glu Pro Gly Ser Glu Thr1
5 10 15Ser Asp Ala Lys Ser Thr Pro Thr
2053023DNAartificialsynthetic construct 530tctccgccgg tgagtctcga
ggc 2353126DNAartificialsynthetic construct 531tgtccctgga
tgcaggctac tctagg 2653224DNAartificialsynthetic construct
532agagtagcct gcatccaggg acag 2453330DNAartificialsynthetic
construct 533tctagatcat ttaccaggag agtgggagag
3053434DNAartificialsynthetic construct 534tctcctggta aatgatctag
agtttaaacc gctg 3453525PRTartificialsynthetic construct 535Ala Thr
Gly Gly Cys Cys Cys Ala Cys Thr Ala Cys Gly Thr Gly Ala1 5 10 15Ala
Cys Cys Ala Thr Cys Ala Cys Cys 20 25
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
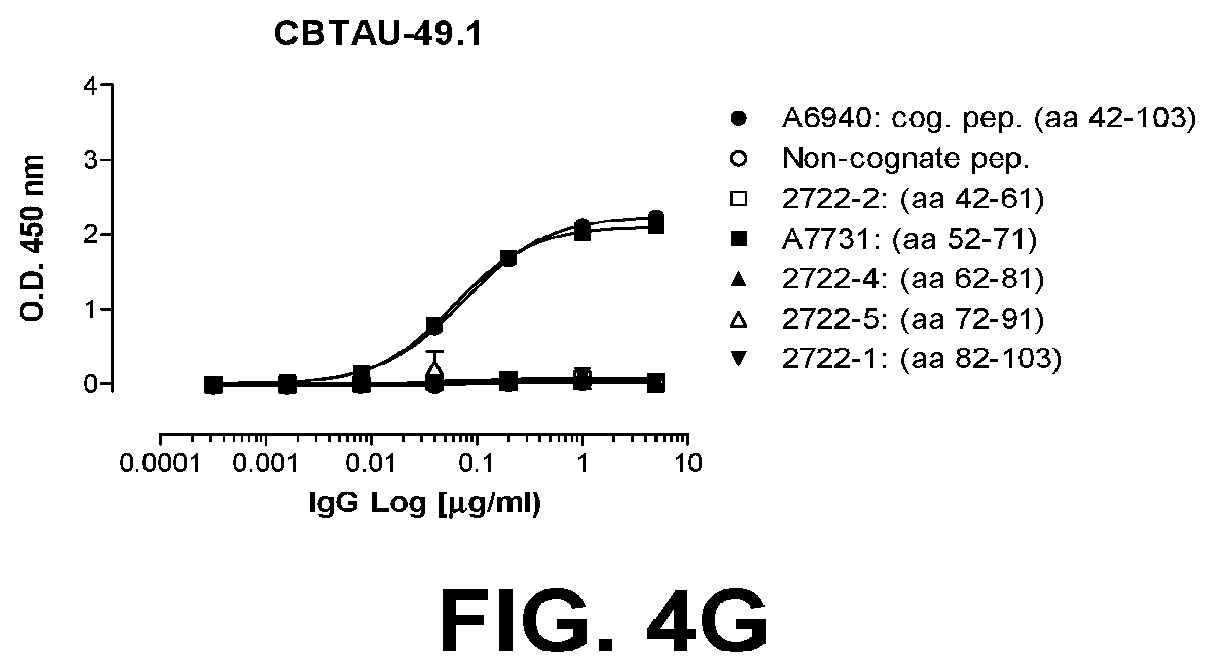
D00014

D00015

D00016
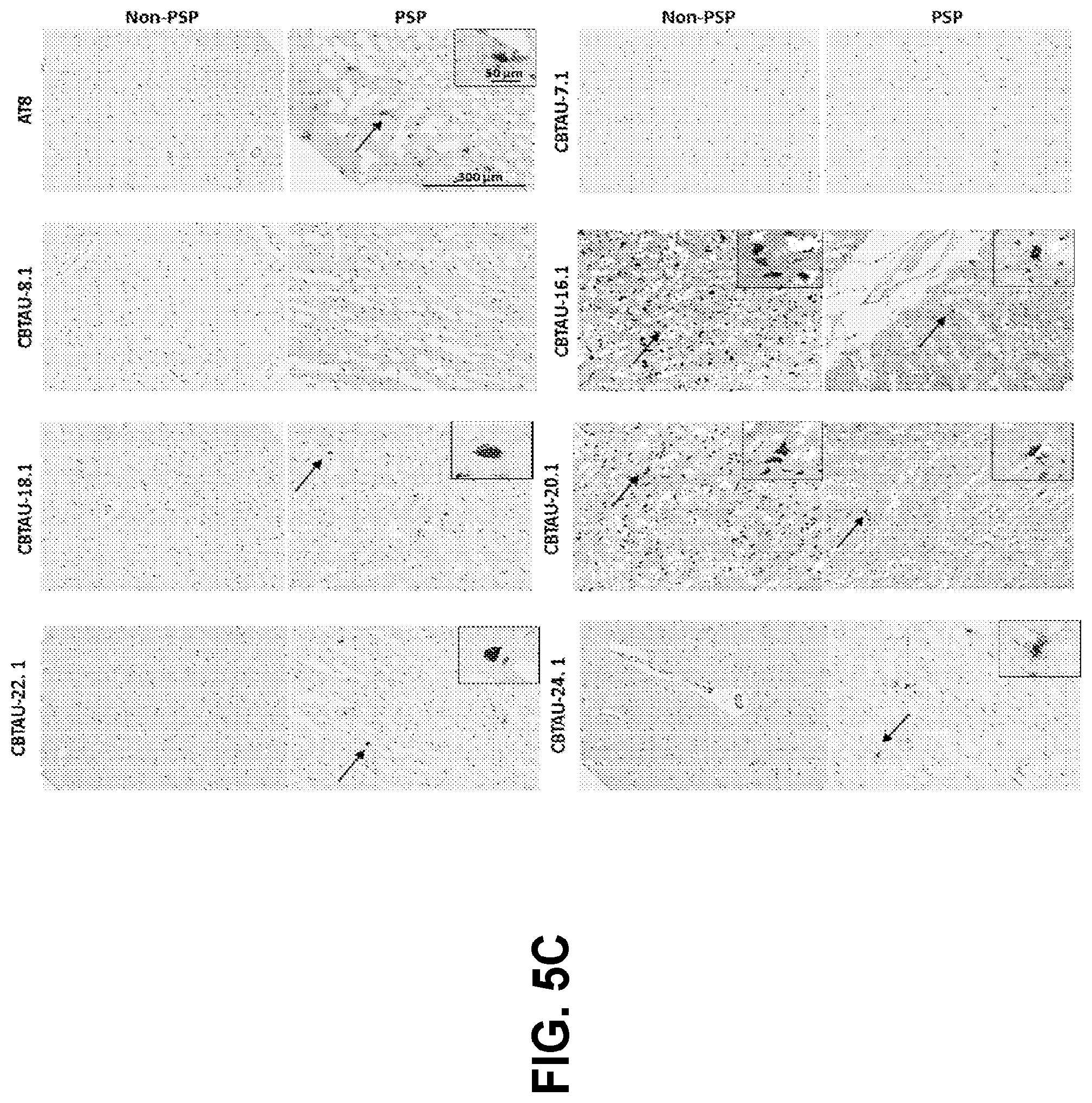

D00017

D00018
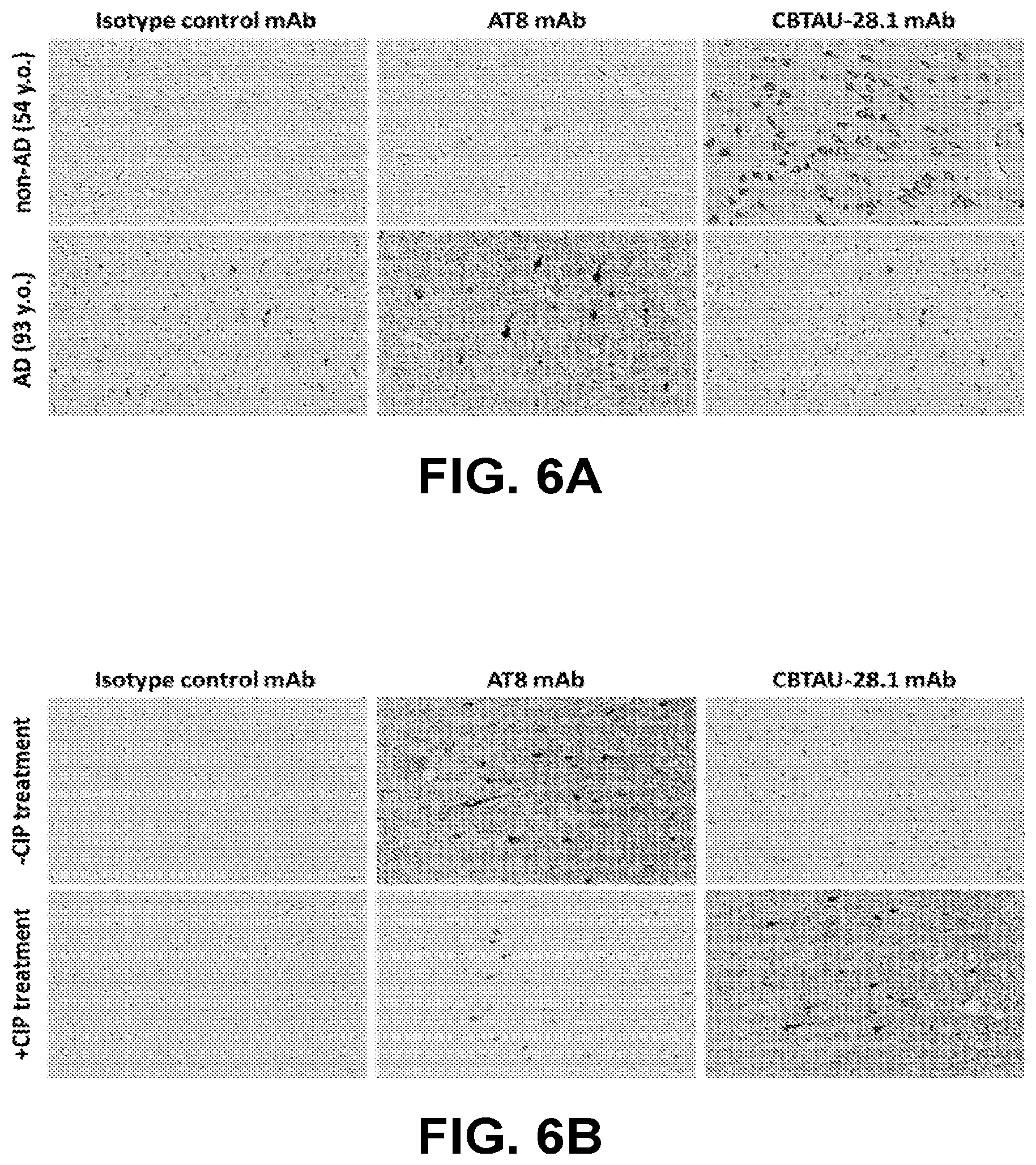
D00019

D00020

D00021
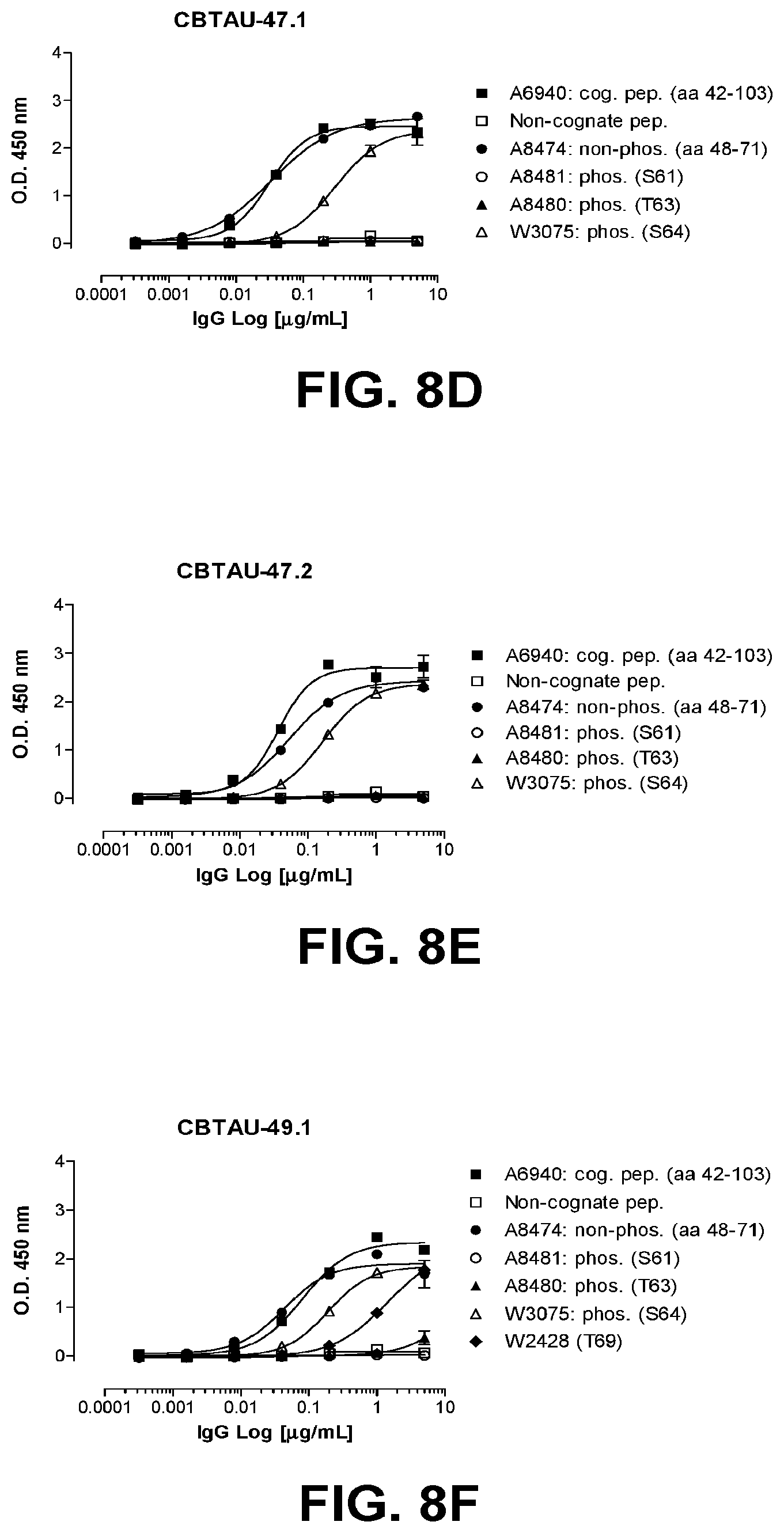
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.