Cd47 Antigen Binding Unit And Uses Thereof
Kind Code
U.S. patent application number 16/481048 was filed with the patent office on 2020-08-06 for cd47 antigen binding unit and uses thereof. The applicant listed for this patent is ZLIP HOLDING LIMITED. Invention is credited to Xu FANG, Honghai GAO, Jing LI, Shou LI, Hongtao Lu, Yangsheng QIU, Lei SHI, Fenglan WU, James S. YAN.
| Application Number | 20200247886 16/481048 |
| Document ID | / |
| Family ID | 1000004777665 |
| Filed Date | 2020-08-06 |










View All Diagrams
| United States Patent Application | 20200247886 |
| Kind Code | A1 |
| QIU; Yangsheng ; et al. | August 6, 2020 |
CD47 ANTIGEN BINDING UNIT AND USES THEREOF
Abstract
Disclosed herein are antigen binding units that specifically bind to CD47. Further disclosed herein are polynucleic acids encoding said antigen binding units, vectors comprising said polynucleic acids, and hybridomas and host cells comprising said vectors. Further provided herein are methods for inducing phagocytosis of CD47-expressing cells.
| Inventors: | QIU; Yangsheng; (Shanghai, CN) ; LI; Jing; (Shanghai, CN) ; GAO; Honghai; (Shanghai, CN) ; WU; Fenglan; (Shanghai, CN) ; FANG; Xu; (Shanghai, CN) ; LI; Shou; (Shanghai, CN) ; Lu; Hongtao; (Shanghai, CN) ; YAN; James S.; (Shanghai, CN) ; SHI; Lei; (Shanghai, CN) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000004777665 | ||||||||||
| Appl. No.: | 16/481048 | ||||||||||
| Filed: | January 26, 2018 | ||||||||||
| PCT Filed: | January 26, 2018 | ||||||||||
| PCT NO: | PCT/CN2018/074318 | ||||||||||
| 371 Date: | July 25, 2019 |
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2317/55 20130101; C07K 2317/33 20130101; C07K 2317/92 20130101; A61K 2039/505 20130101; C07K 2317/732 20130101; C07K 16/2803 20130101; A61P 35/00 20180101; C07K 2317/565 20130101; C07K 2317/24 20130101; C07K 2317/76 20130101 |
| International Class: | C07K 16/28 20060101 C07K016/28; A61P 35/00 20060101 A61P035/00 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Jan 26, 2017 | CN | PCT/CN2017/072738 |
Claims
1. An antigen binding unit comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47; (b) induces phagocytosis of cells expressing CD47 upon binding to CD47; and (c) lacks the ability to induce substantial hemagglutination when mixed with red blood cells at a concentration range of 1.5 ng/ml to 30 .mu.g/ml of the antigen binding unit.
2. The antigen binding unit of claim 1, wherein binding of the antigen binding unit to CD47 prevents binding of CD47 to SIRP.alpha. that is expressed on a macrophage cell.
3. The antigen binding unit of claim 1, wherein the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NOs: 240 and 241, or SEQ ID NOs: 242 and 243, or SEQ ID NOs: 244 and 245.
4. (canceled)
5. (canceled)
6. The antigen binding unit of claim 1, wherein the antigen binding unit exhibits a higher binding affinity to CD47 as compared to an antigen binding unit having the amino acid sequences of SEQ ID NOs: 240 and 241, or SEQ ID NOs: 242 and 243, or SEQ ID NOs: 244 and 245, when assayed in an in vitro binding assay utilizing cells expressing CD47.
7. (canceled)
8. (canceled)
9. The antigen binding unit of claim 1, wherein hemagglutination induced upon contacting the red blood cells with the antigen binding unit is at least 1 fold less as compared to that induced by an antigen binding unit having the amino acid sequences of SEQ ID NOs: 240 and 241, or SEQ ID NOs: 242 and 243, or SEQ ID NOs: 244 and 245.
10. (canceled)
11. (canceled)
12. The antigen binding unit of claim 1, wherein the light chain CDR comprises LC-CDR1, LC-CDR2, and LC-CDR3; and the heavy chain CDR comprises HC-CDR1, HC-CDR2, and HC-CDR3; wherein the LC-CDR1, LC-CDR2, and LC-CDR3 each have a sequence selected from the group consisting of SEQ ID NOs: 1-22 and 162-190; and wherein the HC-CDR1, HC-CDR2, HC-CDR3 each have a sequence selected from the group consisting of SEQ ID NOs: 23-44 and 191-237.
13. The antigen binding unit of claim 12, wherein the light chain CDR comprises amino acid sequences selected from among the following combinations of LC-CDR sequences: i. SEQ ID NO: 4, SEQ ID NO: 8, and SEQ ID NO: 21; ii. SEQ ID NO: 5, SEQ ID NO: 10, and SEQ ID NO: 16; iii. SEQ ID NO: 6, SEQ ID NO: 9, and SEQ ID NO: 17; iv. SEQ ID NO: 2, SEQ ID NO: 12, and SEQ ID NO: 20; v. SEQ ID NO: 7, SEQ ID NO: 11, and SEQ ID NO: 15; vi. SEQ ID NO: 1, SEQ ID NO: 13, and SEQ ID NO: 22; vii. SEQ ID NO: 3, SEQ ID NO: 14, and SEQ ID NO: 19; viii. SEQ ID NO: 169, SEQ ID NO: 173, and SEQ ID NO: 180; ix. SEQ ID NO: 168, SEQ ID NO: 173, and SEQ ID NO: 181; x. SEQ ID NO: 165, SEQ ID NO:177, and SEQ ID NO: 182; xi. SEQ ID NO: 163, SEQ ID NO: 178, and SEQ ID NO: 183; xii. SEQ ID NO: 163, SEQ ID NO: 172, and SEQ ID NO: 184; xiii. SEQ ID NO: 163, SEQ ID NO: 178, and SEQ ID NO: 185; xiv. SEQ ID NO: 163, SEQ ID NO: 178, and SEQ ID NO: 186; xv. SEQ ID NO: 163, SEQ ID NO: 170, and SEQ ID NO: 187; xvi. SEQ ID NO: 163, SEQ ID NO: 174, and SEQ ID NO: 187; xvii. SEQ ID NO: 164, SEQ ID NO: 175, and SEQ ID NO: 187; xviii. SEQ ID NO: 162, SEQ ID NO: 178, and SEQ ID NO: 187; xix. SEQ ID NO: 163, SEQ ID NO: 178, and SEQ ID NO: 187; xx. SEQ ID NO: 164, SEQ ID NO: 178, and SEQ ID NO: 187; xxi. SEQ ID NO: 163, SEQ ID NO: 179, and SEQ ID NO: 187; xxii. SEQ ID NO: 166, SEQ ID NO: 176, and SEQ ID NO: 188; xxiii. SEQ ID NO: 167, SEQ ID NO: 171, and SEQ ID NO: 189; and xxiv. SEQ ID NO: 167, SEQ ID NO: 171, and SEQ ID NO: 190.
14. The antigen binding unit of claim 12, wherein the heavy chain CDR comprises amino acid sequences selected from among the following combinations of HC-CDR sequences: i. SEQ ID NO: 25, SEQ ID NO: 32, and SEQ ID NO: 38; ii. SEQ ID NO: 28, SEQ ID NO: 35, and SEQ ID NO: 39; iii. SEQ ID NO: 24, SEQ ID NO: 34, and SEQ ID NO: 40; iv. SEQ ID NO: 29, SEQ ID NO: 33, and SEQ ID NO: 43; v. SEQ ID NO: 27, SEQ ID NO: 30, and SEQ ID NO: 42; vi. SEQ ID NO: 23, SEQ ID NO: 36, and SEQ ID NO: 41; vii. SEQ ID NO: 26, SEQ ID NO: 31, and SEQ ID NO: 44; viii. SEQ ID NO:191, SEQ ID NO: 207, and SEQ ID NO: 226; ix. SEQ ID NO: 192, SEQ ID NO: 222, and SEQ ID NO: 237; x. SEQ ID NO: 193, SEQ ID NO: 219, and SEQ ID NO: 233; xi. SEQ ID NO: 194, SEQ ID NO: 220, and SEQ ID NO: 228; xii. SEQ ID NO: 195, SEQ ID NO: 221, and SEQ ID NO: 229; xiii. SEQ ID NO: 196, SEQ ID NO: 214, and SEQ ID NO: 225; xiv. SEQ ID NO: 197, SEQ ID NO: 212, and SEQ ID NO: 232; xv. SEQ ID NO: 197, SEQ ID NO: 213, and SEQ ID NO: 232; xvi. SEQ ID NO: 198, SEQ ID NO: 210, and SEQ ID NO: 224; xvii. SEQ ID NO: 198, SEQ ID NO: 208, and SEQ ID NO: 234; xviii. SEQ ID NO: 198, SEQ ID NO: 210, and SEQ ID NO: 234; xix. SEQ ID NO: 199, SEQ ID NO: 210, and SEQ ID NO: 224; xx. SEQ ID NO: 200, SEQ ID NO: 222, and SEQ ID NO: 230; xxi. SEQ ID NO: 201, SEQ ID NO: 210, and SEQ ID NO: 224; xxii. SEQ ID NO: 201, SEQ ID NO: 216, and SEQ ID NO: 224; xxiii. SEQ ID NO: 202, SEQ ID NO: 210, and SEQ ID NO: 234; xxiv. SEQ ID NO: 203, SEQ ID NO: 218, and SEQ ID NO: 227; xxv. SEQ ID NO: 204, SEQ ID NO: 211, and SEQ ID NO: 224; xxvi. SEQ ID NO: 204, SEQ ID NO: 217, and SEQ ID NO: 224; xxvii. SEQ ID NO: 204, SEQ ID NO: 214, and SEQ ID NO: 225; xxviii. SEQ ID NO: 204, SEQ ID NO: 215, and SEQ ID NO:235; xxix. SEQ ID NO: 204, SEQ ID NO: 214, and SEQ ID NO: 236; xxx. SEQ ID NO: 205, SEQ ID NO: 209, and SEQ ID NO: 224; xxxi. SEQ ID NO: 205, SEQ ID NO: 210, and SEQ ID NO: 224; xxxii. SEQ ID NO: 205, SEQ ID NO: 223, and SEQ ID NO: 231; and xxxiii. SEQ ID NO: 206, SEQ ID NO: 210, and SEQ ID NO: 224.
15. The antigen binding unit of claim 1, wherein the antigen binding unit is a monoclonal antibody, a humanized antibody, or a chimeric antibody.
16. The antigen binding unit of claim 1, wherein the antigen binding unit is a scFv, a Fv, a Fab, or a (Fab)2.
17. The antigen binding unit of claim 1, wherein the antigen binding unit competes for binding to an epitope of CD47 recognized by an antigen binding unit having the amino acid sequences of (i) SEQ ID NOs: 240 and 241; (ii) SEQ ID NOs: 242 and 243; or (iii) SEQ ID NOs: 244 and 245.
18.-36. (canceled)
37. A pharmaceutical composition comprising the antigen binding unit of claim 1, and a pharmaceutically acceptable excipient.
38. An isolated nucleic acid encoding the antigen binding unit of claim 1.
39. (canceled)
40. (canceled)
41. A host cell comprising a nucleic acid encoding the antigen binding unit of claim 1.
42.-49. (canceled)
50. A method of treating a cancer in a subject in need thereof, the method comprising administering to the subject an effective amount of the antigen binding unit of claim 1.
51. (canceled)
52. The method of claim 50, wherein the cancer is a hematological cancer or a solid tumor.
53. The method of claim 52, wherein treating the cancer comprises reducing tumor volume.
54. The method of claim 52, further comprising administering a therapeutic antibody.
55. The method of claim 54, wherein the therapeutic antibody is an anti-CD20 antibody.
56. The method of claim 53, wherein the tumor volume is reduced to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NOs: 240 and 241, or SEQ ID NOs: 242 and 243, or SEQ ID NOs: 244 and 245.
Description
CROSS-REFERENCE
[0001] The present application claims priority to International Application No. PCT/CN2017/072738 filed Jan. 26, 2017, the contents of which are hereby incorporated by reference in their entirety.
BACKGROUND OF THE DISCLOSURE
[0002] CD47 is a key molecule that governs macrophage phagocytosis which acts by sending inhibitory signals through SIRP.alpha., a transmembrane receptor that is expressed on macrophages and other myeloid cells. CD47 is ubiquitously expressed and serves as a "marker of self" to prevent macrophage phagocytosis. The same mechanism is employed by cancer cells to evade immunological eradication. In fact, CD47 expression is elevated in several human cancers including solid tumors such as breast, colon, liver, bladder, brain, ovarian, renal, prostate carcinomas, melanoma, and blood cancers such as AML, ALL, CLL, CML, DLBL, FL, MCL, MM and others. CD47 interacts with SIRP.alpha., an inhibitory transmembrane receptor on myeloid cells, such as macrophages. The CD47/SIRP.alpha. interaction leads to bidirectional signaling, resulting in different cell-to-cell responses including inhibition of phagocytosis by the macrophages. Therefore, disruption of this interaction can remove this inhibition, thereby inducing phagocytosis. Existing agents that disrupt such interaction suffers from a number of drawbacks. Amongst them are relative lower affinity and/or selectivity towards CD47, as well as high propensity to induce undesired hemagglutination.
INCORPORATION BY REFERENCE
[0003] All publications and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication or patent application was specifically and individually indicated to be incorporated by reference.
SUMMARY OF THE DISCLOSURE
[0004] There exists a considerable need for alternative CD47-binding agents. The present invention addresses this need and provides related advantages.
[0005] Disclosed herein are antigen binding units comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47; (b) induces phagocytosis of cells expressing CD47 upon binding to CD47; and (c) lacks the ability to induce substantial hemagglutination when mixed with red blood cells at a concentration range of 1.5 ng/ml to 30 ug/ml of said antigen binding unit. In some aspects, binding of the antigen binding unit to CD47 prevents binding of CD47 to SIRP.alpha. that is expressed on a macrophage cell. In some aspects, the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:240-241. In some aspects, the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:242-243. In some aspects, the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:244-245. In some aspects, the antigen binding unit exhibits a higher binding affinity to CD47 as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:240-241, when assayed in an in vitro binding assay utilizing cells expressing CD47. In some aspects, the antigen binding unit exhibits a higher binding affinity to CD47 as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:242-243, when assayed in an in vitro binding assay utilizing cells expressing CD47. In some aspects, the antigen binding unit exhibits a higher binding affinity to CD47 as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:244-245, when assayed in an in vitro binding assay utilizing cells expressing CD47. In some aspects, hemagglutination induced upon contacting the red blood cells with the antigen binding unit is at least 1 fold less as compared to that induced by an antigen binding unit having the amino acid sequences of SEQ ID NO:240-241. In some aspects, hemagglutination induced upon contacting the red blood cells with the antigen binding unit is at least 1 fold less as compared to that induced by an antigen binding unit having the amino acid sequences of SEQ ID NO:242-243. In some aspects, hemagglutination induced upon contacting the red blood cells with the antigen binding unit is at least 1 fold less as compared to that induced by an antigen binding unit having the amino acid sequences of SEQ ID NO:244-245. In some aspects, the light chain CDR comprises LC- CDR1, LC- CDR2, and LC- CDR3; and the heavy chain CDR comprises HC-CDR1, HC-CDR2, and HC-CDR3; wherein said LC-CDR1, LC -CDR2, and LC -CDR3 each have a sequence selected from the group consisting of SEQ ID NO: 1-22 and 162-190; and wherein said HC-CDR1, HC -CDR2, HC -CDR3 each have a sequence selected from the group consisting of SEQ ID NO: 23-44 and 191-237. In some aspects, said light chain CDR comprises amino acid sequences selected from among the following combinations of LC-CDR sequences: a) SEQ ID NO:4, SEQ ID NO:8, and SEQ ID NO:21; b) SEQ ID NO:5, SEQ ID NO:10, and SEQ ID NO:16; c) SEQ ID NO:6, SEQ ID NO:9, and SEQ ID NO:17; d) SEQ ID NO:2, SEQ ID NO:12, and SEQ ID NO:20; e) SEQ ID NO:7, SEQ ID NO:11, and SEQ ID NO:15; f) SEQ ID NO:1, SEQ ID NO:13, and SEQ ID NO:22; g) SEQ ID NO:3, SEQ ID NO:14, and SEQ ID NO:19; h) SEQ ID NO:169, SEQ ID NO:173, and SEQ ID NO:180; i) SEQ ID NO:168, SEQ ID NO:173, and SEQ ID NO:181; k) SEQ ID NO:165, SEQ ID NO:177, and SEQ ID NO:182; l) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:183; m) SEQ ID NO:163, SEQ ID NO:172, and SEQ ID NO:184; n) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:185; o) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:186; p) SEQ ID NO:163, SEQ ID NO:170, and SEQ ID NO:187; q) SEQ ID NO:163, SEQ ID NO:174, and SEQ ID NO:187; r) SEQ ID NO:164, SEQ ID NO:175, and SEQ ID NO:187; s) SEQ ID NO:162, SEQ ID NO:178, and SEQ ID NO:187; t) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:187; u) SEQ ID NO:164, SEQ ID NO:178, and SEQ ID NO:187; v) SEQ ID NO:163, SEQ ID NO:179, and SEQ ID NO:187; w) SEQ ID NO:166, SEQ ID NO:176, and SEQ ID NO:188; x) SEQ ID NO:167, SEQ ID NO:171, and SEQ ID NO:189; and y) SEQ ID NO:167, SEQ ID NO:171, and SEQ ID NO:190. In some aspects, said heavy chain CDR comprises amino acid sequences selected from among the following combinations of HC-CDR sequences: a) SEQ ID NO:25, SEQ ID NO:32, and SEQ ID NO:38; b) SEQ ID NO:28, SEQ ID NO:35, and SEQ ID NO:39; c) SEQ ID NO:24, SEQ ID NO:34, and SEQ ID NO:40; d) SEQ ID NO:29, SEQ ID NO:33, and SEQ ID NO:43; e) SEQ ID NO:27, SEQ ID NO:30, and SEQ ID NO:42; f) SEQ ID NO:23, SEQ ID NO:36, and SEQ ID NO:41; g) SEQ ID NO:26, SEQ ID NO:31, and SEQ ID NO:44; h) SEQ ID NO:191, SEQ ID NO:207, and SEQ ID NO:226; i) SEQ ID NO:192, SEQ ID NO:222, and SEQ ID NO:237; j) SEQ ID NO:193, SEQ ID NO:219, and SEQ ID NO:233; k) SEQ ID NO:194, SEQ ID NO:220, and SEQ ID NO:228; l) SEQ ID NO:195, SEQ ID NO:221, and SEQ ID NO:229; m) SEQ ID NO:196, SEQ ID NO:214, and SEQ ID NO:225; n) SEQ ID NO:197, SEQ ID NO:212, and SEQ ID NO:232; o) SEQ ID NO:197, SEQ ID NO:213, and SEQ ID NO:232; p) SEQ ID NO:198, SEQ ID NO:210, and SEQ ID NO:224; q) SEQ ID NO:198, SEQ ID NO:208, and SEQ ID NO:234; r) SEQ ID NO:198, SEQ ID NO:210, and SEQ ID NO:234; s) SEQ ID NO:199, SEQ ID NO:210, and SEQ ID NO:224; t) SEQ ID NO:200, SEQ ID NO:222, and SEQ ID NO:230; u) SEQ ID NO:201, SEQ ID NO:210, and SEQ ID NO:224; v) SEQ ID NO:201, SEQ ID NO:216, and SEQ ID NO:224; w) SEQ ID NO:202, SEQ ID NO:210, and SEQ ID NO:234; x) SEQ ID NO:203, SEQ ID NO:218, and SEQ ID NO:227; y) SEQ ID NO:204, SEQ ID NO:211, and SEQ ID NO:224; z) SEQ ID NO:204, SEQ ID NO:217, and SEQ ID NO:224; aa) SEQ ID NO:204, SEQ ID NO:214, and SEQ ID NO:225; bb) SEQ ID NO:204, SEQ ID NO:215, and SEQ ID NO:235; cc) SEQ ID NO:204, SEQ ID NO:214, and SEQ ID NO:236; dd) SEQ ID NO:205, SEQ ID NO:209, and SEQ ID NO:224; ee) SEQ ID NO:205, SEQ ID NO:210, and SEQ ID NO:224; ff) SEQ ID NO:205, SEQ ID NO:223, and SEQ ID NO:231; and gg) SEQ ID NO:206, SEQ ID NO:210, and SEQ ID NO:224. In some aspects, the antigen binding unit is a monoclonal antibody, humanized antibody, or chimeric antibody. In some aspects, the antigen binding unit of is sFc, Fv, Fab, or (Fab)2. In some aspects, the antigen binding unit of claim competes for binding to an epitope recognized by an antigen binding unit having the amino acid sequences of 1) SEQ ID NO:240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0006] Disclosed herein are antigen binding units comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47 with a binding affinity higher than that of a reference antigen binding unit and prevents binding of CD47 to SIRP.alpha.; and (b) lacks the ability to induce substantial hemagglutination when mixed with red blood cells at a concentration range of 1.5 ng/ml to 30 ug/ml of antigen binding unit, wherein the reference antigen binding have the amino acid sequences of 1) SEQ ID NO:240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. In some aspects, the light chain CDR comprises LC-CDR1, LC- CDR2, and LC- CDR3; and the heavy chain CDR comprises HC-CDR1, HC-CDR2, and HC-CDR3; wherein said LC-CDR1, LC -CDR2, and LC -CDR3 each have a sequence selected from the group consisting SEQ ID NOs: 1-22, and 162-190; and wherein said HC-CDR1, HC -CDR2, HC -CDR3 each have a sequence selected from the group consisting of SEQ ID NOs: 23-44 and 191-237. In some aspects, said light chain CDR comprises amino acid sequences selected from among the following combinations of LC-CDR sequences: a) SEQ ID NO:4, SEQ ID NO:8, and SEQ ID NO:21; b) SEQ ID NO:5, SEQ ID NO:10, and SEQ ID NO:16; c) SEQ ID NO:6, SEQ ID NO:9, and SEQ ID NO:17; d) SEQ ID NO:2, SEQ ID NO:12, and SEQ ID NO:20; e) SEQ ID NO:7, SEQ ID NO:11, and SEQ ID NO:15; f) SEQ ID NO:1, SEQ ID NO:13, and SEQ ID NO:22; g) SEQ ID NO:3, SEQ ID NO:14, and SEQ ID NO:19; h) SEQ ID NO:169, SEQ ID NO:173, and SEQ ID NO:180; i) SEQ ID NO:168, SEQ ID NO:173, and SEQ ID NO:181; k) SEQ ID NO:165, SEQ ID NO:177, and SEQ ID NO:182; l) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:183; m) SEQ ID NO:163, SEQ ID NO:172, and SEQ ID NO:184; n) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:185; o) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:186; p) SEQ ID NO:163, SEQ ID NO:170, and SEQ ID NO:187; q) SEQ ID NO:163, SEQ ID NO:174, and SEQ ID NO:187; r) SEQ ID NO:164, SEQ ID NO:175, and SEQ ID NO:187; s) SEQ ID NO:162, SEQ ID NO:178, and SEQ ID NO:187; t) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:187; u) SEQ ID NO:164, SEQ ID NO:178, and SEQ ID NO:187; v) SEQ ID NO:163, SEQ ID NO:179, and SEQ ID NO:187; w) SEQ ID NO:166, SEQ ID NO:176, and SEQ ID NO:188; x) SEQ ID NO:167, SEQ ID NO:171, and SEQ ID NO:189; and y) SEQ ID NO:167, SEQ ID NO:171, and SEQ ID NO:190. In some aspects, said heavy chain CDR comprises amino acid sequences selected from among the following combinations of HC-CDR sequences: a) SEQ ID NO:25, SEQ ID NO:32, and SEQ ID NO:38; b) SEQ ID NO:28, SEQ ID NO:35, and SEQ ID NO:39; c) SEQ ID NO:24, SEQ ID NO:34, and SEQ ID NO:40; d) SEQ ID NO:29, SEQ ID NO:33, and SEQ ID NO:43; e) SEQ ID NO:27, SEQ ID NO:30, and SEQ ID NO:42; f) SEQ ID NO:23, SEQ ID NO:36, and SEQ ID NO:41; g) SEQ ID NO:26, SEQ ID NO:31, and SEQ ID NO:44; h) SEQ ID NO:191, SEQ ID NO:207, and SEQ ID NO:226; i) SEQ ID NO:192, SEQ ID NO:222, and SEQ ID NO:237; j) SEQ ID NO:193, SEQ ID NO:219, and SEQ ID NO:233; k) SEQ ID NO:194, SEQ ID NO:220, and SEQ ID NO:228; l) SEQ ID NO:195, SEQ ID NO:221, and SEQ ID NO:229; m) SEQ ID NO:196, SEQ ID NO:214, and SEQ ID NO:225; n) SEQ ID NO:197, SEQ ID NO:212, and SEQ ID NO:232; o) SEQ ID NO:197, SEQ ID NO:213, and SEQ ID NO:232; p) SEQ ID NO:198, SEQ ID NO:210, and SEQ ID NO:224; q) SEQ ID NO:198, SEQ ID NO:208, and SEQ ID NO:234; r) SEQ ID NO:198, SEQ ID NO:210, and SEQ ID NO:234; s) SEQ ID NO:199, SEQ ID NO:210, and SEQ ID NO:224; t) SEQ ID NO:200, SEQ ID NO:222, and SEQ ID NO:230; u) SEQ ID NO:201, SEQ ID NO:210, and SEQ ID NO:224; v) SEQ ID NO:201, SEQ ID NO:216, and SEQ ID NO:224; w) SEQ ID NO:202, SEQ ID NO:210, and SEQ ID NO:234; x) SEQ ID NO:203, SEQ ID NO:218, and SEQ ID NO:227; y) SEQ ID NO:204, SEQ ID NO:211, and SEQ ID NO:224; z) SEQ ID NO:204, SEQ ID NO:217, and SEQ ID NO:224; aa) SEQ ID NO:204, SEQ ID NO:214, and SEQ ID NO:225; bb) SEQ ID NO:204, SEQ ID NO:215, and SEQ ID NO:235; cc) SEQ ID NO:204, SEQ ID NO:214, and SEQ ID NO:236; dd) SEQ ID NO:205, SEQ ID NO:209, and SEQ ID NO:224; ee) SEQ ID NO:205, SEQ ID NO:210, and SEQ ID NO:224; ff) SEQ ID NO:205, SEQ ID NO:223, and SEQ ID NO:231; and gg) SEQ ID NO:206, SEQ ID NO:210, and SEQ ID NO:224. In some aspects, the antigen binding unit is a monoclonal antibody, humanized antibody, chimeric antibody, or bispecific antibody. In some aspects, the antigen binding unit of is sFc, Fv, Fab, or (Fab)2.
[0007] Disclosed herein are antigen binding units comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47; and (b) induces phagocytosis of cells expressing CD47 upon binding to CD47 to a greater extent as compared to a reference antigen binding unit, wherein the reference antigen binding unit have the amino acid sequences of 1) SEQ ID NO:240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. In some aspects, the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:240-241. In some aspects, the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:242-243. In some aspects, the antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:244-245. In some aspects, the light chain CDR comprises LC-CDR1, LC- CDR2, and LC- CDR3; and the heavy chain CDR comprises heavy chain HC-CDR1, HC-CDR2, and HC-CDR3; wherein said LC-CDR1, LC -CDR2, and LC -CDR3 each have a sequence selected from the group consisting of SEQ ID NOs: 1-22, and 162-190; and wherein said HC-CDR1, HC -CDR2, HC -CDR3 each have a sequence selected from the group consisting of SEQ ID NOs: 23-44 and 191-237. In some aspects, said light chain CDR comprises amino acid sequences selected from among the following combinations of LC-CDR sequences: a) SEQ ID NO:4, SEQ ID NO:8, and SEQ ID NO:21; b) SEQ ID NO:5, SEQ ID NO:10, and SEQ ID NO:16; c) SEQ ID NO:6, SEQ ID NO:9, and SEQ ID NO:17; d) SEQ ID NO:2, SEQ ID NO:12, and SEQ ID NO:20; e) SEQ ID NO:7, SEQ ID NO:11, and SEQ ID NO:15; f) SEQ ID NO:1, SEQ ID NO:13, and SEQ ID NO:22; g) SEQ ID NO:3, SEQ ID NO:14, and SEQ ID NO:19; h) SEQ ID NO:169, SEQ ID NO:173, and SEQ ID NO:180; i) SEQ ID NO:168, SEQ ID NO:173, and SEQ ID NO:181; k) SEQ ID NO:165, SEQ ID NO:177, and SEQ ID NO:182; l) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:183; m) SEQ ID NO:163, SEQ ID NO:172, and SEQ ID NO:184; n) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:185; o) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:186; p) SEQ ID NO:163, SEQ ID NO:170, and SEQ ID NO:187; q) SEQ ID NO:163, SEQ ID NO:174, and SEQ ID NO:187; r) SEQ ID NO:164, SEQ ID NO:175, and SEQ ID NO:187; s) SEQ ID NO:162, SEQ ID NO:178, and SEQ ID NO:187; t) SEQ ID NO:163, SEQ ID NO:178, and SEQ ID NO:187; u) SEQ ID NO:164, SEQ ID NO:178, and SEQ ID NO:187; v) SEQ ID NO:163, SEQ ID NO:179, and SEQ ID NO:187; w) SEQ ID NO:166, SEQ ID NO:176, and SEQ ID NO:188; x) SEQ ID NO:167, SEQ ID NO:171, and SEQ ID NO:189; and y) SEQ ID NO:167, SEQ ID NO:171, and SEQ ID NO:190. In some aspects, said heavy chain CDR comprises amino acid sequences selected from among the following combinations of HC-CDR sequences: a) SEQ ID NO:25, SEQ ID NO:32, and SEQ ID NO:38; b) SEQ ID NO:28, SEQ ID NO:35, and SEQ ID NO:39; c) SEQ ID NO:24, SEQ ID NO:34, and SEQ ID NO:40; d) SEQ ID NO:29, SEQ ID NO:33, and SEQ ID NO:43; e) SEQ ID NO:27, SEQ ID NO:30, and SEQ ID NO:42; f) SEQ ID NO:23, SEQ ID NO:36, and SEQ ID NO:41; g) SEQ ID NO:26, SEQ ID NO:31, and SEQ ID NO:44; h) SEQ ID NO:191, SEQ ID NO:207, and SEQ ID NO:226; i) SEQ ID NO:192, SEQ ID NO:222, and SEQ ID
[0008] NO:237; j) SEQ ID NO:193, SEQ ID NO:219, and SEQ ID NO:233; k) SEQ ID NO:194, SEQ ID NO:220, and SEQ ID NO:228; l) SEQ ID NO:195, SEQ ID NO:221, and SEQ ID NO:229; m) SEQ ID NO:196, SEQ ID NO:214, and SEQ ID NO:225; n) SEQ ID NO:197, SEQ ID NO:212, and SEQ ID NO:232; o) SEQ ID NO:197, SEQ ID NO:213, and SEQ ID NO:232; p) SEQ ID NO:198, SEQ ID NO:210, and SEQ ID NO:224; q) SEQ ID NO:198, SEQ ID NO:208, and SEQ ID NO:234; r) SEQ ID NO:198, SEQ ID NO:210, and SEQ ID NO:234; s) SEQ ID NO:199, SEQ ID NO:210, and SEQ ID NO:224; t) SEQ ID NO:200, SEQ ID NO:222, and SEQ ID NO:230; u) SEQ ID NO:201, SEQ ID NO:210, and SEQ ID NO:224; v) SEQ ID NO:201, SEQ ID NO:216, and SEQ ID NO:224; w) SEQ ID NO:202, SEQ ID NO:210, and SEQ ID NO:234; x) SEQ ID NO:203, SEQ ID NO:218, and SEQ ID NO:227; y) SEQ ID NO:204, SEQ ID NO:211, and SEQ ID NO:224; z) SEQ ID NO:204, SEQ ID NO:217, and SEQ ID NO:224; aa) SEQ ID NO:204, SEQ ID NO:214, and SEQ ID NO:225; bb) SEQ ID NO:204, SEQ ID NO:215, and SEQ ID NO:235; cc) SEQ ID NO:204, SEQ ID NO:214, and SEQ ID NO:236; dd) SEQ ID NO:205, SEQ ID NO:209, and SEQ ID NO:224; ee) SEQ ID NO:205, SEQ ID NO:210, and SEQ ID NO:224; ff) SEQ ID NO:205, SEQ ID NO:223, and SEQ ID NO:231; and gg) SEQ ID NO:206, SEQ ID NO:210, and SEQ ID NO:224. In some aspects, the antigen binding unit is a monoclonal antibody, humanized antibody, chimeric antibody, or bispecific antibody. In some aspects, the antigen binding unit of is sFc, Fv, Fab, or (Fab)2.
[0009] Disclosed herein are antigen binding units comprising a light chain CDR and a heavy chain CDR, wherein the light chain CDR comprises LC-CDR1, LC-CDR2, and LC-CDR3; and the heavy chain CDR comprises HC-CDR1, HC-CDR2, and HC-CDR3, wherein said LC-CDR1, LC-CDR2, and LC -CDR3 each comprises a sequence sharing at least 80% sequence homology to a sequence selected from the group consisting of SEQ ID NOs: 1-22, and 162-190, and wherein said HC-CDR1, HC-CDR2, and HC-CDR3 each comprises a sequence having least 80% sequence homology to a sequence selected from the group consisting of SEQ ID NOs: 23-44 and 191-237. In some aspects, said light chain CDR and said heavy chain CDR comprise, respectively, the LC-CDR and the HC-CDR selected from the group consisting of: a) SEQ ID NO:54 and SEQ ID NO:55; b) SEQ ID NO:65 and SEQ ID NO:63; c) SEQ ID NO:58 and SEQ ID NO:64; d) SEQ ID NO:68 and SEQ ID NO:60; e) SEQ ID NO:66 and SEQ ID NO:61; f) SEQ ID NO:57 and SEQ ID NO:62; g) SEQ ID NO:56 and SEQ ID NO:59; h) SEQ ID NO:85 and SEQ ID NO:86; i) SEQ ID NO:87 and SEQ ID NO:88; j) SEQ ID NO:89 and SEQ ID NO:90; k) SEQ ID NO:91 and SEQ ID NO:92; l) SEQ ID NO:93 and SEQ ID NO:94; m) SEQ ID NO:95 and SEQ ID NO:96; n) SEQ ID NO:97 and SEQ ID NO:98; o) SEQ ID NO:99 and SEQ ID NO:100; p) SEQ ID NO:101 and SEQ ID NO:102; q) SEQ ID NO:103 and SEQ ID NO:104; r) SEQ ID NO:105 and SEQ ID NO:106; s) SEQ ID NO:107 and SEQ ID NO:108; t) SEQ ID NO:109 and SEQ ID NO:110; u) SEQ ID NO:111 and SEQ ID NO:112; v) SEQ ID NO:113 and SEQ ID NO:114; w) SEQ ID NO:115 and SEQ ID NO:116; x) SEQ ID NO:117 and SEQ ID NO:118; y) SEQ ID NO:119 and SEQ ID NO:120; z) SEQ ID NO:121 and SEQ ID NO:122; aa) SEQ ID NO:123 and SEQ ID NO:124; bb) SEQ ID NO:125 and SEQ ID NO:126; cc) SEQ ID NO:127 and SEQ ID NO:128; dd) SEQ ID NO:129 and SEQ ID NO:130; ee) SEQ ID NO:131 and SEQ ID NO:132; ff) SEQ ID NO:133 and SEQ ID NO:134; gg) SEQ ID NO:135 and SEQ ID NO:136; hh) SEQ ID NO:137 and SEQ ID NO:138; ii) SEQ ID NO:139 and SEQ ID NO:140; jj) SEQ ID NO:141 and SEQ ID NO:142; kk) SEQ ID NO:143 and SEQ ID NO:144; ll) SEQ ID NO:145 and SEQ ID NO:146; mm) SEQ ID NO:147 and SEQ ID NO:148; nn) SEQ ID NO:238 and SEQ ID NO:239 oo) SEQ ID NO:47 and SEQ ID NO:70; pp) SEQ ID NO:49 and SEQ ID NO:73; qq) SEQ ID NO:71 and SEQ ID NO:51; rr) SEQ ID NO:50 and SEQ ID NO:74; ss) SEQ ID NO:45 and SEQ ID NO:53; tt) SEQ ID NO:67 and SEQ ID NO:72; uu) SEQ ID NO:69 and SEQ ID NO:52; vv) SEQ ID NO:46 and SEQ ID NO:77; ww) SEQ ID NO:46 and SEQ ID NO:78; xx) SEQ ID NO:46 and SEQ ID NO:79; yy) SEQ ID NO:48 and SEQ ID NO:75; zz) SEQ ID NO:48 and SEQ ID NO:76; --aaa) SEQ ID NO:48 and SEQ ID NO:80, and bbb) any sequence pair listed in Table 1. In some aspects, the antigen binding unit is a monoclonal antibody, humanized antibody, chimeric antibody, or bispecific antibody. In some aspects, the antigen binding unit of is sFc, Fv, or Fab.
[0010] Disclosed herein are pharmaceutical compositions comprising any one of the antigen binding units disclosed herein, and a pharmaceutically acceptable excipient.
[0011] Disclosed herein are isolated nucleic acids encoding any one of the antigen binding units disclosed herein.
[0012] Disclosed herein are vectors comprising a nucleic acid sequence encoding any one of the antigen binding units disclosed herein.
[0013] Disclosed herein are host cells expressing any one of the antigen binding units disclosed herein.
[0014] Disclosed herein are host cells comprising a nucleic acid encoding any one of the antigen binding units disclosed herein.
[0015] Disclosed herein are methods of producing any one of the antigen binding units disclosed herein, comprising: culturing any of the host cells disclosed herein under conditions suitable for expressing the antigen binding unit; and isolating said antigen binding unit expressed by the host cell.
[0016] Disclosed herein are methods of inducing phagocytosis of cells expressing CD47, said method comprising contacting the cell with any one of the antigen binding units disclosed herein. In some aspects, phagocytosis of cells expressing CD47 occurs with at least 5% efficiency. In some aspects, the antigen binding unit does not cause significant hemagglutination. In some aspects, the cell is a cancer cell. In some aspects, the cell is a non-lymphoma and non-leukemia cancer cell.
[0017] Disclosed herein are methods of inducing phagocytosis of cells expressing CD47 in a human subject, said method comprising administering to the human subject any one of the pharmaceutical compositions disclosed herein. In some aspects, phagocytosis of cells expressing CD47 occurs with at least 5% efficiency. In some aspects, the antigen binding unit does not cause significant hemagglutination. In some aspects, the cell is a cancer cell. In some aspects, the cell is a non-lymphoma and non-leukemia cancer cell. In some aspects, the cell is a hematological cancer cell or a solid tumor cell.
[0018] Disclosed herein are methods of treating a cancer in a subject in need thereof, said method comprising administering to the subject an effective amount of any one of the antigen binding units disclosed herein. In some aspects, the method further comprises administering a therapeutic antibody. In some aspects, the therapeutic antibody is an anti-CD20 antibody. In some aspects, treating the cancer comprises reducing tumor volume. In some aspects, the tumor volume is reduced to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:240 and 241, or SEQ ID NO: 242 and 243, or SEQ ID NO: 244 and 245.
[0019] Disclosed herein are methods of treating a cancer in a subject in need thereof, said method comprising administering to the subject an effective amount of any one of the pharmaceutical compositions disclosed herein. In some aspects, the cancer is a hematological cancer or a solid tumor. In some aspects, the method further comprises administering a therapeutic antibody. In some aspects, the therapeutic antibody is an anti-CD20 antibody. In some aspects, treating the cancer comprises reducing tumor volume. In some aspects, the tumor volume is reduced to a greater extent as compared to an antigen binding unit having the amino acid sequences of SEQ ID NO:240 and 241, or SEQ ID NO: 242 and 243, or SEQ ID NO: 244 and 245.
BRIEF DESCRIPTION OF THE DRAWINGS
[0020] FIG. 1 depicts data from an example phagocytosis experiment.
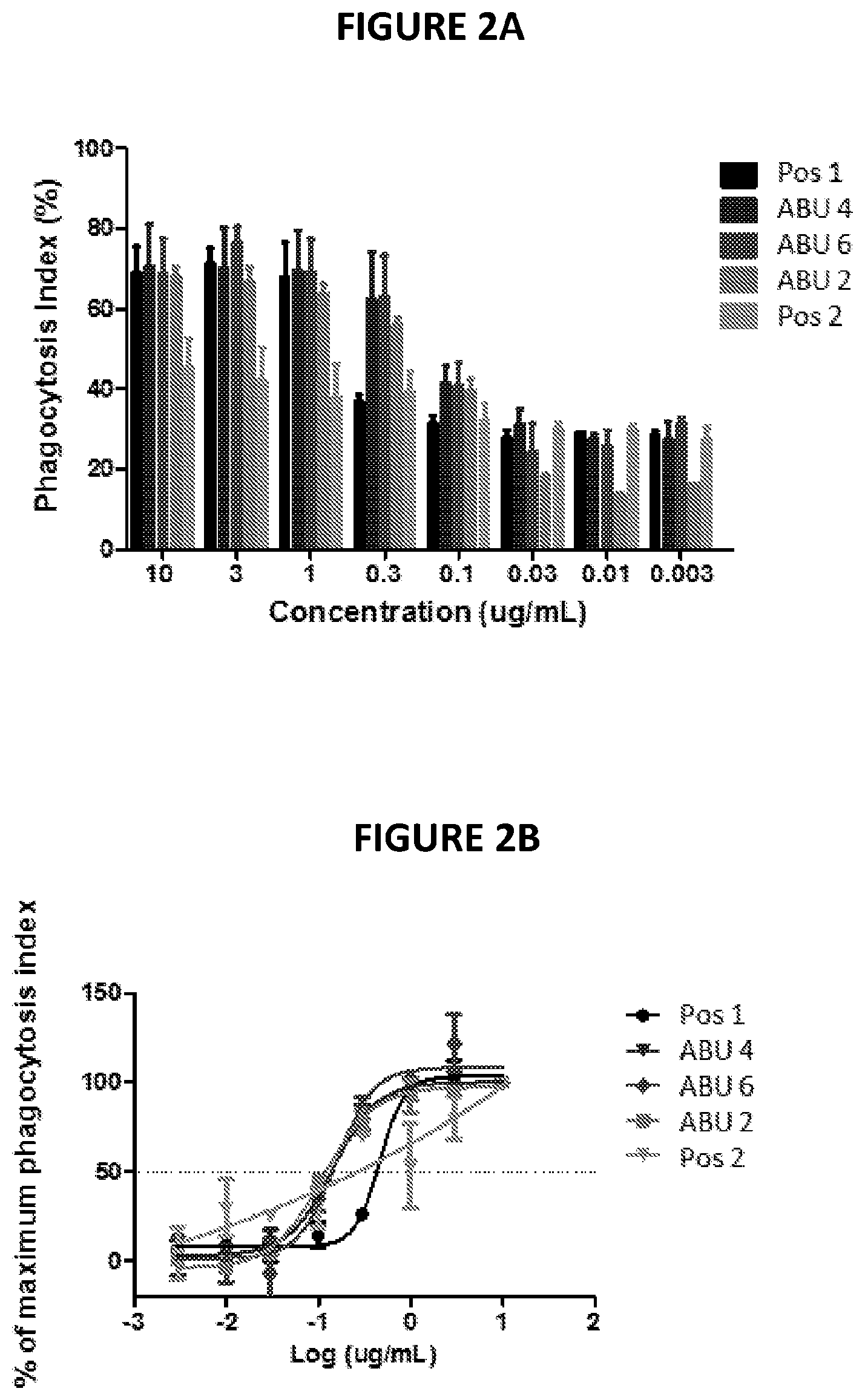
[0021] FIGS. 2A-2B depict data from an example phagocytosis experiment.
[0022] FIGS. 3A-3B depict data from an example antibody binding experiment.
[0023] FIG. 4 depicts data from an example phagocytosis experiment.
[0024] FIG. 5 depicts data from an example phagocytosis experiment.
[0025] FIG. 6 depicts data form an example red blood cell binding experiment.
[0026] FIGS. 7A-7B depict data from an example hemagglutination experiment.
[0027] FIG. 8 depicts data from an example hemagglutination experiment.
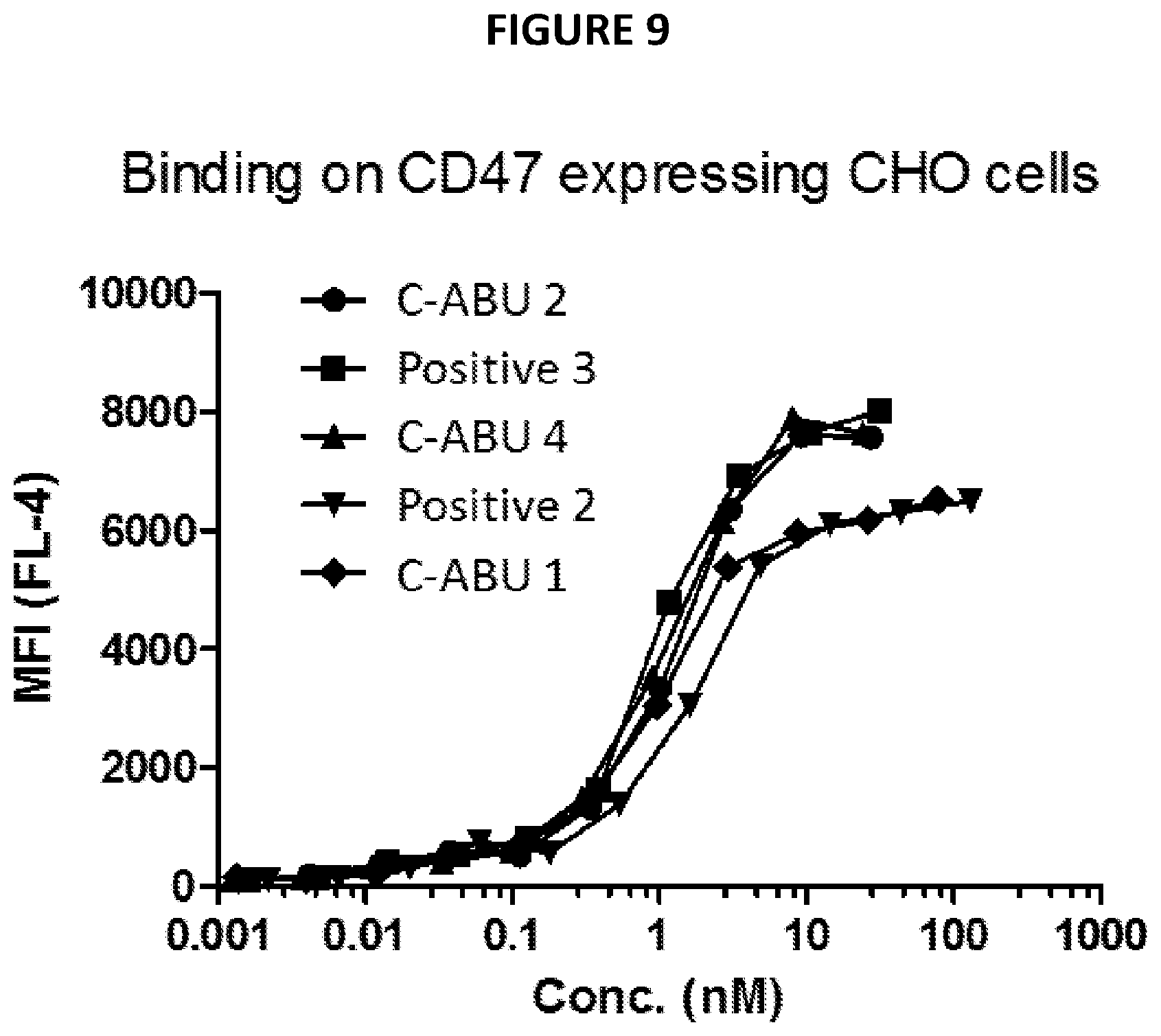
[0028] FIG. 9 depicts data from an example antibody binding experiment.
[0029] FIG. 10 depicts data from an example antibody neutralization experiment.
[0030] FIG. 11 depicts data from an example phagocytosis experiment.
[0031] FIGS. 12A, 12B, and 12C depict data form an example antibody binding experiment.
[0032] FIGS. 13A, 13B, 13C, and 13D depict data form an example antibody binding experiment.
[0033] FIGS. 14A-14B depict data from an example antibody binding experiment.
[0034] FIG. 15A -depicts data from an example antibody binding experiment.
[0035] FIGS. 15B and 15C depict data from an example blocking experiment.
[0036] FIG. 16A depicts data from an example binding assay.
[0037] FIG. 16B depicts data from an example blocking assay.
[0038] FIG. 17 depicts data from an example hemagglutination assay.
[0039] FIG. 18 depicts data from an example binding assay.
[0040] FIGS. 19A, 19B, and 19Cdepict data from example phagocytosis assays.
[0041] FIGS. 20A, 20B, 20C, and 20D depict data from example red blood cell and platelet binding assays.
[0042] FIG. 21 depicts data from an example xenografting experiment.
[0043] FIG. 22A depicts data from an example blocking assay.
[0044] FIG. 22B depicts data from an example RBC binding assay.
[0045] FIG. 22C depicts data from an example platelet binding assay.
[0046] FIG. 22D depicts data from an example hemagglutination assay.
[0047] FIG. 23 depicts data from an example xenografting experiment.
[0048] FIG. 24 depicts data from an example xenografting experiment.
DETAILED DESCRIPTION OF THE DISCLOSURE
[0049] While preferred embodiments of the present invention have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the invention. It should be understood that various alternatives to the embodiments of the invention described herein may be employed in practicing the invention. It is intended that the following claims define the scope of the invention and that methods and structures within the scope of these claims and their equivalents be covered thereby.
[0050] Where a range of values is provided, it is understood that each intervening value, to the tenth of the unit of the lower limit unless the context clearly dictates otherwise, between the upper and lower limit of that range and any other stated or intervening value in that stated range, is encompassed within the invention. The upper and lower limits of these smaller ranges may independently be included in the smaller ranges and are also encompassed within the invention, subject to any specifically excluded limit in the stated range. Where the stated range includes one or both of the limits, ranges excluding either or both of those included limits are also included in the invention.
[0051] The terms "polypeptide", "peptide", and "protein" are used interchangeably herein to refer to polymers of amino acids of any length. The polymer may be linear, cyclic, or branched, it may comprise modified amino acids, and it may be interrupted by non-amino acids. The terms also encompass amino acid polymers that have been modified, for example, via sulfation, glycosylation, lipidation, acetylation, phosphorylation, iodination, methylation, oxidation, proteolytic processing, phosphorylation, prenylation, racemization, selenoylation, transfer-RNA mediated addition of amino acids to proteins such as arginylation, ubiquitination, or any other manipulation, such as conjugation with a labeling component. As used herein the term "amino acid" refers to either natural and/or unnatural or synthetic amino acids, including glycine and both the D or L optical isomers, and amino acid analogs and peptidomimetics. A polypeptide or amino acid sequence "derived from" a designated protein refers to the origin of the polypeptide. Preferably, the polypeptide has an amino acid sequence that is essentially identical to that of a polypeptide encoded in the sequence, or a portion thereof wherein the portion consists of at least 10-20 amino acids, or at least 20-30 amino acids, or at least 30-50 amino acids, or which is immunologically identifiable with a polypeptide encoded in the sequence. This terminology also includes a polypeptide expressed from a designated nucleic acid sequence.
[0052] The term "antigen binding unit" as used herein refers to an immunoglobulin molecule and immunologically active portions of immunoglobulin molecule, i.e., a molecule that contains an antigen-binding site which specifically binds ("immunoreacts with") an antigen. Also encompassed within the term "antigen binding unit" are immunoglobulin molecules of a variety of species origins including invertebrates and vertebrates. Structurally, the simplest naturally occurring antibody (e.g., IgG) comprises four polypeptide chains, two heavy (H) chains and two light (L) chains inter-connected by disulfide bonds. The immunoglobulins represent a large family of molecules that include several types of molecules, such as IgD, IgG, IgA, IgM and IgE.
[0053] The term "immunoglobulin molecule" includes, for example, hybrid antibodies, or altered antibodies, and fragments thereof. It has been shown that the antigen binding function of an antibody can be performed by fragments of a naturally-occurring antibody. These fragments are collectively termed "antigen-binding units". Also encompassed within the term "antigen binding unit" is any polypeptide chain-containing molecular structure that has a specific shape which fits to and recognizes an epitope, where one or more non-covalent binding interactions stabilize the complex between the molecular structure and the epitope.
[0054] An antigen binding unit "specifically binds to" or "immunoreactive with" an antigen if it binds with greater affinity or avidity than it binds to other reference antigens including polypeptides or other substances.
[0055] "Antigen" as used herein means a substance that is recognized and bound specifically by an antigen binding unit. Antigens can include peptides, proteins, glycoproteins, polysaccharides, and lipids; portions thereof and combinations thereof. Non-limiting exemplary antigen included CD47 from human, murine, and other homologues thereof. Another example antigen is SIRP.alpha. from human, murine, and other homologues thereof.
[0056] A "chimeric" protein contains at least one fusion polypeptide comprising regions in a different position in the sequence than what occurs in nature. The regions may normally exist in separate proteins and are brought together in the fusion polypeptide; or they may normally exist in the same protein but are placed in a new arrangement in the fusion polypeptide. A chimeric protein may be created, for example, by chemical synthesis, or by creating and translating a polynucleotide in which the peptide regions are encoded in the desired relationship.
[0057] "Domain" refers to a portion of a protein that is physically or functionally distinguished from other portions of the protein or peptide. Physically-defined domains include those amino acid sequences that are exceptionally hydrophobic or hydrophilic, such as those sequences that are membrane-associated or cytoplasm-associated. Domains may also be defined by internal homologies that arise, for example, from gene duplication. Functionally-defined domains have a distinct biological function(s). The ligand-binding domain of a receptor, for example, is that domain that binds ligand. An antigen-binding domain refers to the part of an antigen-binding unit or an antibody that binds to the antigen. Functionally-defined domains need not be encoded by contiguous amino acid sequences. Functionally-defined domains may contain one or more physically-defined domain. Receptors, for example, are generally divided into the extracellular ligand-binding domain, a transmembrane domain, and an intracellular effector domain.
[0058] A "host cell" includes an individual cell or cell culture which can be or has been a recipient for the subject vectors. Host cells include progeny of a single host cell. The progeny may not necessarily be completely identical (in morphology or in genomic of total DNA complement) to the original parent cell due to natural, accidental, or deliberate mutation. A host cell includes cells transfected in vivo with a vector of this invention. A "host cell" can refer to a prokaryotic cell, a eukaryotic cell, or cell line cultured as a unicellular entity which can be, or has been, used as a recipient for a recombinant vector or other transfer polynucleotides, and include the progeny of the original cell which has been transfected. It is understood that the progeny of a single cell may not necessarily be completely identical in morphology or in genomic or total DNA complement as the original parent, due to natural, accidental, or deliberate mutation.
[0059] A "cell line" or "cell culture" denotes bacterial, plant, insect or higher eukaryotic cells grown or maintained in vitro. The descendants of a cell may not be completely identical (either morphologically, genotypically, or phenotypically) to the parent cell.
[0060] As used herein, the term "isolated" means separated from constituents, cellular and otherwise, in which the polynucleotide, peptide, polypeptide, protein, antibody, or fragments thereof, are normally associated with in nature. As is apparent to those of skill in the art, a non-naturally occurring polynucleotide, peptide, polypeptide, protein, antibody, or fragments thereof, does not require "isolation" to distinguish it from its naturally occurring counterpart. In addition, a "concentrated", "separated" or "diluted" polynucleotide, peptide, polypeptide, protein, antibody, or fragments thereof, is distinguishable from its naturally occurring counterpart in that the concentration or number of molecules per volume is greater than "concentrated" or less than "separated" than that of its naturally occurring counterpart. Enrichment can be measured on an absolute basis, such as weight per volume of solution, or it can be measured in relation to a second, potentially interfering substance present in the source mixture. Increasing enrichments of the embodiments of this invention are increasingly more preferred. Thus, for example, a 2-fold enrichment is preferred, 10-fold enrichment is more preferred, 100-fold enrichment is more preferred, 1000-fold enrichment is even more preferred. A substance can also be provided in an isolated state by a process of artificial assembly, such as by chemical synthesis or recombinant expression.
[0061] "Linked" and "fused" or "fusion" are used interchangeably herein. These terms refer to the joining together of two more chemical elements or components, by whatever means including chemical conjugation or recombinant means. An "in-frame fusion" refers to the joining of two or more open reading frames (ORFs) to form a continuous longer ORF, in a manner that maintains the correct reading frame of the original ORFs. Thus, the resulting recombinant fusion protein is a single protein containing two or more segments that correspond to polypeptides encoded by the original ORFs (which segments are not normally so joined in nature). Although the reading frame is thus made continuous throughout the fused segments, the segments may be physically or spatially separated by, for example, in-frame linker sequence (e.g. "flexon").
[0062] In the context of polypeptides, a "sequence" is an order of amino acids in a polypeptide in an amino to carboxyl terminus direction in which residues that neighbor each other in the sequence are contiguous in the primary structure of the polypeptide. A sequence can also be a linear sequence of part of a polypeptide which is known to comprise additional residues in one or both directions.
[0063] "Heterologous" means derived from a genotypically distinct entity from the rest of the entity to which it is being compared. For example, a promoter removed from its native coding sequence and operatively linked to a coding sequence other than the native sequence is a heterologous promoter. The term "heterologous" as applied to a polynucleotide, a polypeptide, means that the polynucleotide or polypeptide is derived from a genotypically distinct entity from that of the rest of the entity to which it is being compared. For instance, a heterologous polynucleotide or antigen may be derived from a different species origin, different cell type, and the same type of cell of distinct individuals.
[0064] The terms "polynucleotides", "nucleic acids", "nucleotides" and "oligonucleotides" are used interchangeably. They refer to a polymeric form of nucleotides of any length, either deoxyribonucleotides or ribonucleotides, or analogs thereof. Polynucleotides may have any three-dimensional structure, and may perform any function, known or unknown. The following are non-limiting examples of polynucleotides: coding or non-coding regions of a gene or gene fragment, loci (locus) defined from linkage analysis, exons, introns, messenger RNA (mRNA), transfer RNA, ribosomal RNA, ribozymes, cDNA, recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes, primers, oligonucleotides, or synthesized DNA. A polynucleotide may comprise modified nucleotides, such as methylated nucleotides and nucleotide analogs. If present, modifications to the nucleotide structure may be imparted before or after assembly of the polymer. The sequence of nucleotides may be interrupted by non-nucleotide components. A polynucleotide may be further modified after polymerization, such as by conjugation with a labeling component.
[0065] "Recombinant" as applied to a polynucleotide means that the polynucleotide is the product of various combinations of cloning, restriction and/or ligation steps, and other procedures that result in a construct that is distinct from a polynucleotide found in nature.
[0066] The terms "gene" or "gene fragment" are used interchangeably herein. They refer to a polynucleotide containing at least one open reading frame that is capable of encoding a particular protein after being transcribed and translated. A gene or gene fragment may be genomic, cDNA, or synthesized, as long as the polynucleotide contains at least one open reading frame, which may cover the entire coding region or a segment thereof.
[0067] "Operably linked" or "operatively linked" refers to a juxtaposition wherein the components so described are in a relationship permitting them to function in their intended manner. For instance, a promoter sequence is operably linked to a coding sequence if the promoter sequence promotes transcription of the coding sequence.
[0068] A "fusion gene" is a gene composed of at least two heterologous polynucleotides that are linked together.
[0069] A gene "database" denotes a set of stored data which represent a collection of sequences including nucleotide and peptide sequences, which in turn represent a collection of biological reference materials.
[0070] As used herein, "expression" refers to the process by which a polynucleotide is transcribed into mRNA and/or the process by which the transcribed mRNA (also referred to as "transcript") is subsequently being translated into peptides, polypeptides, or proteins. The transcripts and the encoded polypeptides are collectively referred to as gene product. If the polynucleotide is derived from genomic DNA, expression may include splicing of the mRNA in a eukaryotic cell.
[0071] A "vector" is a nucleic acid molecule, preferably self-replicating, which transfers an inserted nucleic acid molecule into and/or between host cells. The term includes vectors that function primarily for insertion of DNA or RNA into a cell, replication of vectors that function primarily for the replication of DNA or RNA, and expression vectors that function for transcription and/or translation of the DNA or RNA. Also included are vectors that provide more than one of the above functions. An "expression vector" is a polynucleotide which, when introduced into an appropriate host cell, can be transcribed and translated into a polypeptide(s). An "expression system" usually connotes a suitable host cell comprised of an expression vector that can function to yield a desired expression product.
[0072] The term "biological sample" encompasses a variety of sample types obtained from an organism and can be used in a diagnostic or monitoring assay. The term encompasses blood and other liquid samples of biological origin, solid tissue samples, such as a biopsy specimen or tissue cultures or cells derived therefrom and the progeny thereof. The term encompasses samples that have been manipulated in any way after their procurement, such as by treatment with reagents, solubilization, or enrichment for certain components. The term encompasses a clinical sample, and also includes cells in cell culture, cell supernatants, cell lysates, serum, plasma, biological fluids, and tissue samples.
[0073] The terms "treatment", "treating", "treat" and the like are used herein to generally refer to obtaining a desired pharmacologic and/or physiologic effect. The effect may be prophylactic in terms of completely or partially preventing a disease or symptom thereof and/or may be therapeutic in terms of a partial or complete stabilization or cure for a disease and/or adverse effect attributable to the disease. "Treatment" as used herein covers any treatment of a disease in a mammal, e.g. mouse, rat, rabbit, pig, primate, including humans and other apes, particularly a human, and includes: (a) preventing the disease or symptom from occurring in a subject which may be predisposed to the disease or symptom but has not yet been diagnosed as having it; (b) inhibiting the disease symptom; (c) arresting development of the disease; (d) relieving the disease symptom; (e) causing regression of the disease or symptom; or any combination thereof.
[0074] The terms "recipient", "individual", "subject", "host", and "patient", can be used interchangeably herein and refer to any mammalian subject for whom diagnosis, treatment, or therapy is desired, particularly humans.
[0075] The terms "cancer", "neoplasm", "tumor", and "carcinoma", are used interchangeably herein to refer to cells which exhibit relatively autonomous growth, so that they exhibit an aberrant growth phenotype characterized by a significant loss of control of cell proliferation. In general, cells of interest for detection or treatment in the present application include precancerous (e.g., benign), malignant, pre-metastatic, metastatic, and non-metastatic cells. The term "normal" as used in the context of "normal cell," is meant to refer to a cell of an untransformed phenotype or exhibiting a morphology of a non-transformed cell of the tissue type being examined. "Cancerous phenotype" generally refers to any of a variety of biological phenomena that are characteristic of a cancerous cell, which phenomena can vary with the type of cancer. The cancerous phenotype is generally identified by abnormalities in, for example, cell growth or proliferation (e.g., uncontrolled growth or proliferation), regulation of the cell cycle, cell mobility, cell-cell interaction, or metastasis, etc.).
[0076] Phagocytic cells or phagocytes are interchangeable terms and refer to a cell that is capable of phagocytosis. Non-limiting categories of phagocytes include macrophages, mononuclear cells (e.g., histiocytes and monocytes), polymorphonuclear leukocytes (e.g., neutrophils), and dendritic cells.
Compositions
[0077] In one embodiment, the present disclosure provides an antigen binding unit comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47; (b) induces phagocytosis of cells expressing CD47 upon binding to CD47; and (c) lacks the ability to induce substantial hemagglutination when mixed with red blood cells at a concentration ranging from about 1.5 ng/ml to about 30 ug/ml of the antigen binding unit disclosed herein.
[0078] In another embodiment, the present disclosure provides an antigen binding unit comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47 with a binding affinity higher than that of a reference antigen binding unit and prevents binding of CD47 to SIRPa; and (b) lacks the ability to induce substantial hemagglutination when mixed with red blood cells at a concentration range of about 1.5 ng/ml to about 30 ug/ml of said antigen binding unit, wherein the reference antigen binding unit has the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0079] In yet another embodiment, the present disclosure provides an antigen binding unit comprising a light chain CDR and a heavy chain CDR, wherein the antigen binding unit (a) specifically binds to CD47; and (b) induces phagocytosis of cells expressing CD47 upon binding to CD47 to a greater extent as compared to a reference antigen binding unit, wherein the reference antigen binding unit has the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0080] In still yet another embodiment, the present disclosure provides an antigen binding unit comprising a light chain CDR and a heavy chain CDR, wherein the light chain CDR comprises LC-CDR1, LC-CDR2, and LC-CDR3; and the heavy chain CDR comprises HC-CDR1, HC-CDR2, and HC-CDR3, wherein said LC-CDR1, LC-CDR2, and LC -CDR3 each comprises a sequence sharing at least 80% sequence homology to a sequence selected from the group consisting of SEQ ID NOs: 1-22, and 162-190, and wherein said HC-CDR1, HC-CDR2, and HC-CDR3 each comprises a sequence having least 80% sequence homology to a sequence selected from the group consisting of SEQ ID NOs: 23-44 and 191-237.
[0081] In some aspects, an antigen binding unit can compete for binding to an epitope recognized by a reference antigen binding unit. For example, an antigen binding unit can compete for binding to an epitope recognized by a reference antigen binding wherein having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. Epitope binning of select antibodies was conducted using CD47-experssing CHO cells with a commercialized anti-CD47 blocking antibodies. Briefly, seven neutralizing CD47 antibodies and two reference antibodies (positive 1 and positive 2) were analyzed and grouped according to their competitive binding to CHO cell expressing CD47 using flow cytometry. A biotinylated antibody was first used to calculate the concentration for 90% binding, then the 9 CD47 antibodies were serially diluted and mixed with one of the biotinylated antibodies at the pre-determined 90% binding concentration. SA-APC was used to detect the binding of biotinylated antibody binding. All antibodies were compared against each other and the controls. Antibodies that showed no change in the binding were classified as the same group. Antibodies that showed a change in cell surface binding were categorized into separate groups. Three binding profiles on the CHO expressing CD47 were identified and the six hits subjected to three groups as below. One group consisted of ABU1, ABU6, and positive 1. A second group consisted of ABU4, ABUS, and positive 1. A third group consisted of ABU2, ABU3, positive 1, and positive 2.
[0082] In some aspects of any of the embodiments disclosed herein, an antigen binding unit comprises a light chain CDR. A light chain CDR can be a complementarity determining region of a light chain of an antigen binding unit. A light chain CDR can comprise a continuous sequence of amino acid residues, or two or more contiguous sequences of amino acid residues separated by, and optionally flanked by, non-complementarity determining regions, such as framework regions. In some examples, a light chain CDR comprises two or more light chain CDRs, which can be referred to as light chain CDR-1, CDR-2, and so on. In advantageous examples, a light chain CDR comprises three light chain CDRs, which can be referred to as light chain CDR-1, light chain CDR-2, and light chain CDR-3 respectively. In some examples, a group of CDRs present on a common light chain can collectively be referred to as light chain CDRs.
[0083] In some aspects of any of the embodiments disclosed herein, an antigen binding unit comprises a heavy chain CDR. A heavy chain CDR can be a complementarity determining region of a heavy chain of an antigen binding unit. A heavy chain CDR can comprise a continuous sequence of amino acid residues, or two or more contiguous sequences of amino acid residues separated by, and optionally flanked by, non-complementarity determining regions, such as framework regions. In some examples, a heavy chain CDR comprises two or more heavy chain CDRs, which can be referred to as heavy chain CDR-1, CDR-2, and so on. In advantageous examples, a heavy chain CDR comprises three heavy chain CDRs, which can be referred to as heavy chain CDR-1, heavy chain CDR-2, and heavy chain CDR-3 respectively. In some examples, a group of CDRs present on a common heavy chain can collectively be referred to as heavy chain CDRs.
[0084] In some aspects of any of the embodiments disclosed herein, a subject antigen binding unit specifically binds to CD47. CD47 as used herein can also refer to orthologues, homologues, codon-optimized forms, truncated forms, fragmented forms, mutated forms, or any other known derivative form of a known CD47 sequence. For example, CD47 can be human CD47, which is represented by GenBank accession number CEJ95640 and comprises the sequence of SEQ ID NO:81. CD47 can be murine CD47, which is represented by GenBank accession number BAA25401.1 and comprises the sequence of SEQ ID NO:82. In some contexts, CD47 is referred to as Integrin-Associated Protein (IAP). Human IAP is represented by GenBank accession number CAA80977.1, and comprises the sequence of SEQ ID NO:83. Murine IAP is represented by GenBank accession number ADQ12919.1, and comprises the sequence of SEQ ID NO:84. Additionally, CD47 can comprise a sequence sharing at least 50% identity to any one of SEQ ID NO: 81-84. CD47 can be comprise a sequence sharing at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 99%, or greater than 99% identity to any one of SEQ ID NO: 81-84.
[0085] Binding specificity can be determined by complementarity determining regions, or CDRs, such as light chain CDRs or heavy chain CDRs. In many cases, binding specificity is determined by light chain CDRs and heavy chain CDRs. A given combination of heavy chain CDRs and light chain CDRs provides a given binding pocket that confers greater affinity and/or specificity towards CD47 as compared to other reference antigens.
[0086] In some aspects of an embodiment disclosed herein, an antigen binding unit specifically binds to CD47 with a binding affinity higher than that of a reference antigen binding unit. Such reference antigen binding units include, but are not limited to, an antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0087] Binding of an antigen binding unit to CD47 can be characterized or expressed by any method known in the art. For example, binding can be characterized by binding affinity, which can be the strength of the interaction between the antigen binding unit and the antigen. Binding affinity can be determined by any method known in the art, such as in vitro binding assays. For example, binding affinity of antigen binding units disclosed herein can be determined when assayed in an in vitro binding assay utilizing cells expressing CD47. Binding affinity of subject antigen binding unit can be expressed in term of Kd, which is the equilibrium dissociation constant between an antibody and its respective antigen. In some cases, antigen binding units as disclosed herein specifically bind to CD47 with a Kd within a range of about 10 .mu.M to about 1 fM. For example, an antigen binding unit can specifically bind to CD47 with a Kd of less than about 10 .mu.M, 1 .mu.M, 0.1 .mu.M, 10 nM, 1 nM, 0.1 nM, 10 pM, 1 pM, 0.1 pM, 10 fM, 1 fM, 0.1 fM, or less than 0.1 fM. In some examples, a subject antigen binding unit exhibits a higher binding affinity to CD47 as compared to a reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0088] In some aspects of an embodiment disclosed herein, an antigen binding unit reduces or even prevents binding of CD47 to SIRP.alpha., and thereby induces phagocytosis by a macrophage cell that expresses SIRP.alpha.. Typically, such phagocytosis is induced upon binding of the antigen binding unit to CD47.
[0089] In some aspects, a subject antigen binding unit induces phagocytosis of a cell expressing
[0090] CD47 to a greater extent than that of a reference antigen binding unit. Such reference antigen binding unit can have the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. Phagocytosis can be accessed qualitatively by any method known in the art. In some cases, the extent of phagocytosis is determined by the number of macrophages that have performed phagocytosis (referred to as phagocytes) among a population of macrophages. For example, the number of phagocytes per 100 macrophages can be determined so that the extent of phagocytosis can be expressed as a percentage or phagocytic index.
[0091] Inducing phagocytosis of cells expressing CD47 can by evidenced by an increase in the level of phagocytosis of these cells in the presence of an antigen binding unit disclosed herein. In some examples, the level of phagocytosis of such cells is increased by at least 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 500%, 1000%, or greater than 1000% as compared to the level of phagocytosis observed in the absence of the composition.
[0092] In some aspects of any of the embodiments disclosed herein, an antigen binding unit lacks the ability to induce substantial hemagglutination. In some cases, an antigen binding unit lacks the ability to induce substantial hemagglutination when mixed with red blood cells at a concentration range between about 1.5 ng/ml to about 30 .mu.g/ml of said antigen binding unit. For example, a subject antigen binding unit can lack the ability to induce substantial hemagglutination when mixed with red blood cells, when the antigen binding unit is at a concentration of about 0.1 ng/mL, 1 ng/mL, 5 ng/mL, 10 ng/mL, 15 ng/mL, 20 ng/mL, 25 ng/mL, 30 ng/mL, 35 ng/mL, 40 ng/mL, 45 ng/mL, 50 ng/mL, 55 ng/mL, 60 ng/mL, 65 ng/mL, 70 ng/mL, 75 ng/mL, 80 ng/mL, 85 ng/mL, 90 ng/mL, 100 ng/mL, 150 ng/mL, 200 ng/mL, 250 ng/mL, 300 ng/mL, 350 ng/mL, 400 ng/mL, 450 ng/mL, 500 ng/mL, 550 ng/mL, 600 ng/mL, 650 ng/mL, 700 ng/mL, 750 ng/mL, 800 ng/mL, 850 ng/mL, 900 ng/mL, 950 ng/mL, 1 .mu.g/ml, 1.5 .mu.g/ml, 2 .mu.g/ml, 3 .mu.g/ml, 4 .mu.g/ml, 5 .mu.g/ml, 10 .mu.g/ml, 15 .mu.g/ml, 20 .mu.g/ml, 25 .mu.g/ml, 30 .mu.g/ml or more of said antigen binding unit. In other examples the antigen binding unit concentration can be less than 1.5 ng/mL. In other examples, the antigen binding unit concentration can be greater than 30 .mu.g/ml.
[0093] In some cases, hemagglutination induced upon contacting the red blood cells with a subject antigen binding unit is at least 1 fold less as compared to that induced by a reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. In some cases, hemagglutination induced upon contacting the red blood cells with a subject antigen binding unit is at least 1 fold less, at least 2 fold less, at least 3 fold less, at least 4 fold less, at least 5 fold less, at least 6 fold less, at least 7 fold less, at least 8 fold less, at least 9 fold less, or at least 10 fold less as compared to that induced by a reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. In some cases, hemagglutination induced upon contacting the red blood cells with a subject antigen binding unit is greater than at least 10 fold less as compared to that induced by a reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0094] In some cases, evidence of hemagglutination is demonstrated by the presence of non-settlement of RBCs. Where a punctate red dot appears instead of a haze indicates a lack of substantial hemagglutination.
[0095] In some aspects, hemagglutination can be quantified and expressed as a hemagglutination index. Hemagglutination index can be quantified by the area of a red blood cell pellet in the presence or absence of subject antigen binding units. For example, the diameter of a red blood cell pellet can be determined either manually or using computer software, such as Image J. When using computer software, the area of a red blood cell pellet can be determined by counting the number of pixels making up the pellet. The area can then be calculated manually or by using software such as Excel. In some cases, the area can then be normalized to a control data set and expressed as a percent of maximum hemagglutination index. In such examples, subject antigen binding units can induce about 0%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 110%, 120%, 130%, 140%, 150% or greater of the maximum hemagglutination index. In some examples, subject antigen binding units induce less than 100% of the maximum hemagglutination index. For example, subject antigen binding units can induce less than about 100%, 95%, 90%, 85%, 80%, 75%, 70%, 65%, 60%, 55%, 50%, 45%, 40%, 35%, 30%, 25%, 20%, 15%, 10%, 5%, or less of the maximum hemagglutination index.
[0096] In some aspects, a subject antigen binding unit lacks the ability to induce substantial hemagglutination when the antigen binding unit is added to a solution of red blood cells (RBC), in which the RBCs constitute more than about 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, or 20% RBCs in a suitable buffer, such as PBS. In some examples, the solution is greater than 20% RBCs in a suitable buffer, such as PBS.
[0097] In some aspects, a subject antigen binding unit lacks the ability to induce substantial hemagglutination of RBCs in a solution containing RBCs, where the antigen binding unit is present at a concentration from about 100 .mu.g/mL to about 1 pg/mL. For example, a lack of the substantial hemagglutination is observed when the antigen binding unit concentration is at least about, 0.1 ng/mL, 0.2 ng/mL, 0.3 ng/mL, 0.4 ng/mL, 0.5 ng/mL, 0.6 ng/mL, 0.7 ng/mL, 0.8 ng/mL, 0.9 ng/mL, 1 ng/mL, 2 ng/mL, 3 ng/mL, 4 ng/mL, 5 ng/mL, 6 ng/mL, 7 ng/mL, 8 ng/mL, 9 ng/mL, 10 ng/mL, 20 ng/mL, 30 ng/mL, 40 ng/mL, 50 ng/mL, 60 ng/mL, 70 ng/mL, 80 ng/mL, 90 ng/mL, 100 ng/mL, 0.1 .mu.g/mL, 0.2 .mu.g/mL, 0.3 .mu.g/mL, 0.4 .mu.g/mL, 0.5 .mu.g/mL, 0.6 .mu.g/mL, 0.7 .mu.g/mL, 0.8 .mu.g/mL, 0.9 .mu.g/mL, 1 .mu.g/mL, 2 .mu.g/mL, 3 .mu.g/mL, 4 .mu.g/mL, 5 .mu.g/mL, 6 .mu.g/mL, 7 .mu.g/mL, 8 .mu.g/mL, 9 .mu.g/mL, 10 .mu.g/mL, 20 .mu.g/mL, 30 .mu.g/mL, 40 .mu.g/mL, 50 .mu.g/mL, 60 .mu.g/mL, 70 .mu.g/mL, 80 .mu.g/mL, 90 .mu.g/mL, 100 .mu.g/mL, or higher.
[0098] In some aspects of any of the embodiments disclosed herein, an antigen binding unit lacks the ability to induce substantial hemagglutination after the antigen binding unit is added to red blood cells and incubated for about 10 minutes to about 10 hours. For example, substantial hemagglutination is not observed after about 10 min, 15 min, 30 min, 45 min, 1 hr, 1.5 hr, 2 hr, 2.5 hr, 3 hr, 3.5 hr, 4 hr, 4.5 hr, 5 hr, 5.5 hr, 6 hr, 6.5 hr, 7 hr, 7.5 hr, 8 hr, 8.5 hr, 9 hr, 9.5 hr, 10 hr, or greater than 10 hr of incubation time.
[0099] In some aspects of any of the embodiments disclosed herein, an antigen binding unit comprises a light chain CDR and a heavy chain CDR. Subject antigen binding units can comprise any LC-CDR or HC-CDR listed in Table 1. Additionally or alternatively, a subject antigen binding unit can comprise a LC-CDR or a HC-CDR with at least 60% identity to any of the LC-CDR or HC-CDR listed in Table 1. In some aspects, a subject LC-CDR or HC-CDR can exhibit at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 99%, or greater sequence identity to any of the SEQ ID NOs listed in Table 1.
TABLE-US-00001 TABLE 1 Antigen binding unit LC-CDR HC-CDR ABU1 SEQ ID NO: 54 SEQ ID NO: 55 ABU2 SEQ ID NO: 65 SEQ ID NO: 63 ABU3 SEQ ID NO: 58 SEQ ID NO: 64 ABU4 SEQ ID NO: 68 SEQ ID NO: 60 ABU5 SEQ ID NO: 66 SEQ ID NO: 61 ABU6 SEQ ID NO: 57 SEQ ID NO: 62 ABU7 SEQ ID NO: 56 SEQ ID NO: 59 ABU8 SEQ ID NO: 85 SEQ ID NO: 86 ABU9 SEQ ID NO: 87 SEQ ID NO: 88 ABU10 SEQ ID NO: 89 SEQ ID NO: 90 ABU11 SEQ ID NO: 91 SEQ ID NO: 92 ABU12 SEQ ID NO: 93 SEQ ID NO: 94 ABU13 SEQ ID NO: 95 SEQ ID NO: 96 ABU14 SEQ ID NO: 97 SEQ ID NO: 98 ABU15 SEQ ID NO: 99 SEQ ID NO: 100 ABU16 SEQ ID NO: 101 SEQ ID NO: 102 ABU17 SEQ ID NO: 103 SEQ ID NO: 104 ABU18 SEQ ID NO: 105 SEQ ID NO: 106 ABU19 SEQ ID NO: 107 SEQ ID NO: 108 ABU20 SEQ ID NO: 109 SEQ ID NO: 110 ABU21 SEQ ID NO: 111 SEQ ID NO: 112 ABU22 SEQ ID NO: 113 SEQ ID NO: 114 ABU23 SEQ ID NO: 115 SEQ ID NO: 116 ABU24 SEQ ID NO: 117 SEQ ID NO: 118 ABU25 SEQ ID NO: 119 SEQ ID NO: 120 ABU26 SEQ ID NO: 121 SEQ ID NO: 122 ABU27 SEQ ID NO: 123 SEQ ID NO: 124 ABU28 SEQ ID NO: 125 SEQ ID NO: 126 ABU29 SEQ ID NO: 127 SEQ ID NO: 128 ABU30 SEQ ID NO: 129 SEQ ID NO: 130 ABU31 SEQ ID NO: 131 SEQ ID NO: 132 ABU32 SEQ ID NO: 133 SEQ ID NO: 134 ABU33 SEQ ID NO: 135 SEQ ID NO: 136 ABU34 SEQ ID NO: 137 SEQ ID NO: 138 ABU35 SEQ ID NO: 139 SEQ ID NO: 140 ABU36 SEQ ID NO: 141 SEQ ID NO: 142 ABU37 SEQ ID NO: 143 SEQ ID NO: 144 ABU38 SEQ ID NO: 145 SEQ ID NO: 146 ABU39 SEQ ID NO: 147 SEQ ID NO: 148 ABU40 SEQ ID NO: 238 SEQ ID NO: 239 C-ABU1 SEQ ID NO: 47 SEQ ID NO: 70 C-ABU2 SEQ ID NO: 49 SEQ ID NO: 73 C-ABU3 SEQ ID NO: 71 SEQ ID NO: 51 C-ABU4 SEQ ID NO: 50 SEQ ID NO: 74 C-ABU4 SEQ ID NO: 309 SEQ ID NO: 310 C-ABU5 SEQ ID NO: 45 SEQ ID NO: 53 C-ABU6 SEQ ID NO: 67 SEQ ID NO: 72 C-ABU7 SEQ ID NO: 69 SEQ ID NO: 52 C-ABU8 SEQ ID NO: 311 SEQ ID NO: 312 H-ABU1a SEQ ID NO: 46 SEQ ID NO: 77 H-ABU1b SEQ ID NO: 46 SEQ ID NO: 78 H-ABU1c SEQ ID NO: 46 SEQ ID NO: 79 H-ABU2a SEQ ID NO: 48 SEQ ID NO: 75 H-ABU2b SEQ ID NO: 48 SEQ ID NO: 76 H-ABU2c SEQ ID NO: 48 SEQ ID NO: 80 H-ABU 3 SEQ ID NO: 265 SEQ ID NO: 246 H-ABU 4 SEQ ID NO: 266 SEQ ID NO: 247 H-ABU 5 SEQ ID NO: 267 SEQ ID NO: 248 H-ABU 6 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 7 SEQ ID NO: 269 SEQ ID NO: 250 H-ABU 8 SEQ ID NO: 270 SEQ ID NO: 251 H-ABU 9 SEQ ID NO: 270 SEQ ID NO: 252 H-ABU 10 SEQ ID NO: 271 SEQ ID NO: 253 H-ABU 11 SEQ ID NO: 272 SEQ ID NO: 249 H-ABU 12 SEQ ID NO: 287 SEQ ID NO: 292 H-ABU 13 SEQ ID NO: 288 SEQ ID NO: 292 H-ABU 14 SEQ ID NO: 289 SEQ ID NO: 292 H-ABU 15 SEQ ID NO: 290 SEQ ID NO: 292 H-ABU 16 SEQ ID NO: 291 SEQ ID NO: 292 H-ABU 17 SEQ ID NO: 287 SEQ ID NO: 293 H-ABU 18 SEQ ID NO: 288 SEQ ID NO: 293 H-ABU 19 SEQ ID NO: 289 SEQ ID NO: 293 H-ABU 20 SEQ ID NO: 290 SEQ ID NO: 293 H-ABU 21 SEQ ID NO: 291 SEQ ID NO: 293 H-ABU 22 SEQ ID NO: 287 SEQ ID NO: 294 H-ABU 23 SEQ ID NO: 288 SEQ ID NO: 294 H-ABU 24 SEQ ID NO: 289 SEQ ID NO: 294 H-ABU 25 SEQ ID NO: 290 SEQ ID NO: 294 H-ABU 26 SEQ ID NO: 291 SEQ ID NO: 294 H-ABU 27 SEQ ID NO: 287 SEQ ID NO: 295 H-ABU 28 SEQ ID NO: 288 SEQ ID NO: 295 H-ABU 29 SEQ ID NO: 289 SEQ ID NO: 295 H-ABU 30 SEQ ID NO: 290 SEQ ID NO: 295 H-ABU 31 SEQ ID NO: 291 SEQ ID NO: 295 H-ABU 32 SEQ ID NO: 287 SEQ ID NO: 296 H-ABU 33 SEQ ID NO: 288 SEQ ID NO: 296 H-ABU 34 SEQ ID NO: 289 SEQ ID NO: 296 H-ABU 35 SEQ ID NO: 290 SEQ ID NO: 296 H-ABU 36 SEQ ID NO: 291 SEQ ID NO: 296 H-ABU 37 SEQ ID NO: 291 SEQ ID NO: 292 H-ABU 38 SEQ ID NO: 291 SEQ ID NO: 294 H-ABU 39 SEQ ID NO: 291 SEQ ID NO: 295 H-ABU 40 SEQ ID NO: 288 SEQ ID NO: 296 H-ABU 41 SEQ ID NO: 291 SEQ ID NO: 296 H-ABU 42 SEQ ID NO: 290 SEQ ID NO: 292 H-ABU 43 SEQ ID NO: 291 SEQ ID NO: 293 H-ABU 44 SEQ ID NO: 298 SEQ ID NO: 302 H-ABU 45 SEQ ID NO: 273 SEQ ID NO: 254 H-ABU 46 SEQ ID NO: 274 SEQ ID NO: 255 H-ABU 47 SEQ ID NO: 275 SEQ ID NO: 256 H-ABU 48 SEQ ID NO: 276 SEQ ID NO: 257 H-ABU 49 SEQ ID NO: 277 SEQ ID NO: 258 H-ABU 50 SEQ ID NO: 265 SEQ ID NO: 259 H-ABU 51 SEQ ID NO: 278 SEQ ID NO: 260 H-ABU 52 SEQ ID NO: 279 SEQ ID NO: 261 H-ABU 53 SEQ ID NO: 280 SEQ ID NO: 262 H-ABU 54 SEQ ID NO: 281 SEQ ID NO: 263 H-ABU 55 SEQ ID NO: 282 SEQ ID NO: 246 H-ABU 56 SEQ ID NO: 283 SEQ ID NO: 264 H-ABU 57 SEQ ID NO: 284 SEQ ID NO: 249 H-ABU 58 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 59 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 60 SEQ ID NO: 272 SEQ ID NO: 249 H-ABU 61 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 62 SEQ ID NO: 285 SEQ ID NO: 249 H-ABU 63 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 64 SEQ ID NO: 272 SEQ ID NO: 249 H-ABU 65 SEQ ID NO: 272 SEQ ID NO: 249 H-ABU 66 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 67 SEQ ID NO: 286 SEQ ID NO: 249 H-ABU 68 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 69 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 70 SEQ ID NO: 268 SEQ ID NO: 249 H-ABU 71 SEQ ID NO: 297 SEQ ID NO: 300 H-ABU 72 SEQ ID NO: 298 SEQ ID NO: 300 H-ABU 73 SEQ ID NO: 299 SEQ ID NO: 300 H-ABU 74 SEQ ID NO: 297 SEQ ID NO: 301 H-ABU 75 SEQ ID NO: 298 SEQ ID NO: 301 H-ABU 76 SEQ ID NO: 299 SEQ ID NO: 301 H-ABU 77 SEQ ID NO: 297 SEQ ID NO: 302 H-ABU 78 SEQ ID NO: 299 SEQ ID NO: 302 H-ABU 79 SEQ ID NO: 297 SEQ ID NO: 303 H-ABU 80 SEQ ID NO: 298 SEQ ID NO: 303 H-ABU 81 SEQ ID NO: 299 SEQ ID NO: 303 H-ABU 82 SEQ ID NO: 297 SEQ ID NO: 304 H-ABU 83 SEQ ID NO: 298 SEQ ID NO: 304 H-ABU 84 SEQ ID NO: 299 SEQ ID NO: 304
[0100] In some cases, the light chain (LC) CDR comprises light LC-CDR1, LC-CDR2, and LC-CDR3; and the heavy chain (HC) CDR comprises HC-CDR1, HC-CDR2, and HC-CDR3. In some examples, said LC-CDR1, LC -CDR2, and LC -CDR3 each have a sequence selected from the group consisting of SEQ ID NOs: 1-22, and 162-190. In some examples, said HC-CDR1, HC-CDR2, HC -CDR3 each have a sequence selected from the group consisting of SEQ ID NO: 23-44 and 191-237. In some examples, said LC-CDR1, LC -CDR2, and LC -CDR3 each have a sequence selected from the group consisting of SEQ ID NO: 1-22 and 162-190 and said HC-CDR1, HC -CDR2, HC -CDR3 each have a sequence selected from the group consisting of SEQ ID NO: 23-44 and 191-237.
[0101] In some aspects of any of the embodiments disclosed herein, an antigen binding unit comprises a light chain CDR wherein said light chain (LC) CDR comprises a combination of three LC-CDRs, namely LC-CDR1, LC-CDR2, and LC-CDR3. A combination of three LC-CDRs can comprise any combination listed in Table 2.
TABLE-US-00002 TABLE 2 Example LC-CDR LC-CDR1 LC-CDR2 LC-CDR3 Example 1 SEQ ID NO: 4 SEQ ID NO: 8 SEQ ID NO: 21 Example 2 SEQ ID NO: 5 SEQ ID NO: 10 SEQ ID NO: 16 Example 3 SEQ ID NO: 6 SEQ ID NO: 9 SEQ ID NO: 17 Example 4 SEQ ID NO: 2 SEQ ID NO: 12 SEQ ID NO: 20 Example 5 SEQ ID NO: 7 SEQ ID NO: 11 SEQ ID NO: 15 Example 6 SEQ ID NO: 1 SEQ ID NO: 13 SEQ ID NO: 22 Example 7 SEQ ID NO: 3 SEQ ID NO: 14 SEQ ID NO: 19 Example 8 SEQ ID NO: 169 SEQ ID NO: 173 SEQ ID NO: 180 Example 9 SEQ ID NO: 168 SEQ ID NO: 173 SEQ ID NO: 181 Example 10 SEQ ID NO: 165 SEQ ID NO: 177 SEQ ID NO: 182 Example 11 SEQ ID NO: 163 SEQ ID NO: 178 SEQ ID NO: 183 Example 12 SEQ ID NO: 163 SEQ ID NO: 172 SEQ ID NO: 184 Example 13 SEQ ID NO: 163 SEQ ID NO: 178 SEQ ID NO: 185 Example 14 SEQ ID NO: 163 SEQ ID NO: 178 SEQ ID NO: 186 Example 15 SEQ ID NO: 163 SEQ ID NO: 170 SEQ ID NO: 187 Example 16 SEQ ID NO: 163 SEQ ID NO: 174 SEQ ID NO: 187 Example 17 SEQ ID NO: 164 SEQ ID NO: 175 SEQ ID NO: 187 Example 18 SEQ ID NO: 162 SEQ ID NO: 178 SEQ ID NO: 187 Example 19 SEQ ID NO: 163 SEQ ID NO: 178 SEQ ID NO: 187 Example 20 SEQ ID NO: 164 SEQ ID NO: 178 SEQ ID NO: 187 Example 21 SEQ ID NO: 163 SEQ ID NO: 179 SEQ ID NO: 187 Example 22 SEQ ID NO: 166 SEQ ID NO: 176 SEQ ID NO: 188 Example 23 SEQ ID NO: 167 SEQ ID NO: 171 SEQ ID NO: 189 Example 24 SEQ ID NO: 167 SEQ ID NO: 171 SEQ ID NO: 190
[0102] In some aspects of any of the embodiments disclosed herein, an antigen binding unit comprises a heavy chain CDR wherein said heavy chain (HC) CDR comprises a combination of three HC-CDRs, namely HC-CDR1, HC-CDR2, and HC-CDR3. A combination of three HC-CDRs can comprise any combination listed in Table 3.
TABLE-US-00003 TABLE 3 Example HC-CDR HC-CDR1 HC-CDR2 HC-CDR3 Example 1 SEQ ID NO: 25 SEQ ID NO: 32 SEQ ID NO: 38 Example 2 SEQ ID NO: 28 SEQ ID NO: 35 SEQ ID NO: 39 Example 3 SEQ ID NO: 24 SEQ ID NO: 34 SEQ ID NO: 40 Example 4 SEQ ID NO: 29 SEQ ID NO: 33 SEQ ID NO: 43 Example 5 SEQ ID NO: 27 SEQ ID NO: 30 SEQ ID NO: 42 Example 6 SEQ ID NO: 23 SEQ ID NO: 36 SEQ ID NO: 41 Example 7 SEQ ID NO: 26 SEQ ID NO: 31 SEQ ID NO: 44 Example 8 SEQ ID NO: 191 SEQ ID NO: 207 SEQ ID NO: 226 Example 9 SEQ ID NO: 192 SEQ ID NO: 222 SEQ ID NO: 237 Example 10 SEQ ID NO: 193 SEQ ID NO: 219 SEQ ID NO: 233 Example 11 SEQ ID NO: 194 SEQ ID NO: 220 SEQ ID NO: 228 Example 12 SEQ ID NO: 195 SEQ ID NO: 221 SEQ ID NO: 229 Example 13 SEQ ID NO: 196 SEQ ID NO: 214 SEQ ID NO: 225 Example 14 SEQ ID NO: 197 SEQ ID NO: 212 SEQ ID NO: 232 Example 15 SEQ ID NO: 197 SEQ ID NO: 213 SEQ ID NO: 232 Example 16 SEQ ID NO: 198 SEQ ID NO: 210 SEQ ID NO: 224 Example 17 SEQ ID NO: 198 SEQ ID NO: 208 SEQ ID NO: 234 Example 18 SEQ ID NO: 198 SEQ ID NO: 210 SEQ ID NO: 234 Example 19 SEQ ID NO: 199 SEQ ID NO: 210 SEQ ID NO: 224 Example 20 SEQ ID NO: 200 SEQ ID NO: 222 SEQ ID NO: 230 Example 21 SEQ ID NO: 201 SEQ ID NO: 210 SEQ ID NO: 224 Example 22 SEQ ID NO: 201 SEQ ID NO: 216 SEQ ID NO: 224 Example 23 SEQ ID NO: 202 SEQ ID NO: 210 SEQ ID NO: 234 Example 24 SEQ ID NO: 203 SEQ ID NO: 218 SEQ ID NO: 227 Example 25 SEQ ID NO: 204 SEQ ID NO: 211 SEQ ID NO: 224 Example 26 SEQ ID NO: 204 SEQ ID NO: 217 SEQ ID NO: 224 Example 27 SEQ ID NO: 204 SEQ ID NO: 214 SEQ ID NO: 225 Example 28 SEQ ID NO: 204 SEQ ID NO: 215 SEQ ID NO: 235 Example 29 SEQ ID NO: 204 SEQ ID NO: 214 SEQ ID NO: 236 Example 30 SEQ ID NO: 205 SEQ ID NO: 209 SEQ ID NO: 224 Example 31 SEQ ID NO: 205 SEQ ID NO: 210 SEQ ID NO: 224 Example 32 SEQ ID NO: 205 SEQ ID NO: 223 SEQ ID NO: 231 Example 33 SEQ ID NO: 206 SEQ ID NO: 210 SEQ ID NO: 224
[0103] In some aspects of any of the embodiments disclosed herein, an antigen binding unit comprises a light chain CDR and a heavy chain CDR, wherein said light chain CDR and said heavy chain CDR comprise, respectively, the LC-CDR and the HC-CDR selected from the group consisting of any combination of LC-CDRs listed in Table 2 and any combination of HC-CDRs listed in Table 3.
[0104] In some aspects, a subject antigen binding unit is a monoclonal antigen binding unit, a polyclonal antigen binding unit, a humanized antigen binding unit, a chimeric antigen binding unit, a monovalent antigen binding unit, a multivalent antigen binding unit, a bispecific antigen binding unit, or any combination thereof. The antigen binding units can adopt a variety of formats, including but not limited to sFC, Fv, ccFv, Fab', F(ab')2, and Fd. Such antibody binding units can be generated from whole immunoglobulins by ricin, pepsin, papain, or other protease cleavage.
[0105] In addition, antigen binding units can be designed utilizing recombinant immunoglobulin techniques. For instance "Fv" immunoglobulins for use in the present invention may be produced by linking a variable light chain region to a variable heavy chain region via a peptide linker. For example, a peptide linker can be poly-glycine or another sequence which does not form an alpha helix or beta sheet motif. Fvs can also be made which comprise stabilizing disulfide bonds between the V.sub.H and V.sub.L regions, as described in U.S. Pat. No. 6,147,203, incorporated fully herein by reference. Any of these antigen binding unites can be utilized in the present invention. In some aspects, an antigen binding unit can be a whole immunoglobulin having two light chains paired with two heavy chains.
[0106] Antigen-binding units can be heteromultimers comprising a light-chain polypeptide and a heavy-chain polypeptide. Examples of an antigen binding unit include but are not limited to (i) a ccFv fragment stabilized by the heterodimerization sequences disclosed U.S. Pat. No. 6,833,441, incorporated herein in its entirety; (ii) any other monovalent and multivalent molecules comprising at least one ccFv fragment as described herein; (iii) a Fab fragment consisting of the VL, VH, CL and CH1 domains; (iv) an Fd fragment consisting of the VH and CH1 domains; (v) an Fv fragment consisting of the VL and VH domains of a single arm of an antibody; (vi) an F(ab')2 fragment, a bivalent fragment comprising two Fab fragments linked by a disulfide bridge at the hinge region; and (vii) a diabody.
[0107] Polyclonal antibodies can be raised by a standard protocol by injecting a production animal with an antigenic composition. See, e.g., Harlow and Lane, Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, 1988. When utilizing an entire protein, or a larger section of the protein, antibodies may be raised by immunizing the production animal with the protein and a suitable adjuvant (e.g., Freund's, Freund's complete, oil-in-water emulsions, etc.). When a smaller peptide is utilized, it is advantageous to conjugate the peptide with a larger molecule to make an immunostimulatory conjugate. Commonly utilized conjugate proteins that are commercially available for such use include bovine serum albumin (BSA) and keyhole limpet hemocyanin (KLH). In order to raise antibodies to particular epitopes, peptides derived from the full sequence may be utilized. Alternatively, in order to generate antibodies to relatively short peptide portions of the protein target, a superior immune response may be elicited if the polypeptide is joined to a carrier protein, such as ovalbumin, BSA or KLH.
[0108] Polyclonal or monoclonal antigen binding units or antibodies can be produced from animals which have been genetically altered to produce human immunoglobulins. A transgenic animal can be produced by initially producing a "knock-out" animal which does not produce the animal's natural antibodies, and stably transforming the animal with a human antibody locus (e.g., by the use of a human artificial chromosome). In such cases, only human antibodies are then made by the animal. Techniques for generating such animals, and deriving antibodies therefrom, are described in U.S. Pat. Nos. 6,162,963 and 6,150,584, incorporated fully herein by reference. Such antibodies can be referred to as human xenogenic antibodies.
[0109] Alternatively, antigen binding units can be produced from phage libraries containing human variable regions. See U.S. Pat. No. 6,174,708, incorporated fully herein by reference.
[0110] In some aspects of any of the embodiments disclosed herein, an antigen binding unit is produced by a hybridoma. For example, an antigen binding unit disclosed herein can be produced by a hybridoma selected form the group consisting of hybridomas expressing one of the antigen binding units listed in Table 1. For example, the hybridoma can be any hybridoma as deposited on [date] with reference number [insert reference number].
[0111] For monoclonal antigen binding units or monoclonal antibodies, hybridomas may be formed by isolating the stimulated immune cells, such as those from the spleen of the inoculated animal. These cells can then be fused to immortalized cells, such as myeloma cells or transformed cells, which are capable of replicating indefinitely in cell culture, thereby producing an immortal, immunoglobulin-secreting cell line. The immortal cell line utilized can be selected to be deficient in enzymes necessary for the utilization of certain nutrients. Many such cell lines (such as myelomas) are known to those skilled in the art, and include, for example: thymidine kinase (TK) or hypoxanthine-guanine phosphoriboxyl transferase (HGPRT). These deficiencies allow selection for fused cells according to their ability to grow on, for example, hypoxanthine aminopterinthymidine medium (HAT).
[0112] In addition, the antigen binding unit may be produced by genetic engineering. Humanized, chimeric, or xenogeneic human antigen binding units, which produce less of an immune response when administered to humans, are of use in the present invention.
[0113] Antigen binding units disclosed herein can have a reduced propensity to induce an undesired immune response in humans, for example, anaphylactic shock, and can also exhibit a reduced propensity for priming an immune response which would prevent repeated dosage with the antibody therapeutic or imaging agent (e.g., the human-anti-murine-antibody "HAMA" response). Such antigen binding units include, but are not limited to, humanized, chimeric, or xenogenic human antigen binding units.
[0114] Chimeric antigen binding units or chimeric antibodies can be made, for example, by recombinant means by combining the murine variable light and heavy chain regions (VK and VH), obtained from a murine (or other animal-derived) hybridoma clone, with the human constant light and heavy chain regions, in order to produce an antibody with predominantly human domains. The production of such chimeric antibodies is well known in the art, and may be achieved by standard means (as described, e.g., in U.S. Pat. No. 5,624,659, incorporated fully herein by reference).
[0115] The term "humanized" as applies to a non-human (e.g. rodent or primate) antibodies are hybrid immunoglobulins, immunoglobulin chains or fragments thereof which contain minimal sequence derived from non-human immunoglobulin. For the most part, humanized antibodies are human immunoglobulins (recipient antibody) in which residues from a complementary determining region (CDR) of the recipient are replaced by residues from a CDR of a non-human species (donor antibody) such as mouse, rat, rabbit or primate having the desired specificity, affinity and capacity. In some instances, Fv framework region (FR) residues of the human immunoglobulin are replaced by corresponding non-human residues. Furthermore, the humanized antibody may comprise residues which are found neither in the recipient antibody nor in the imported CDR or framework sequences. These modifications are made to further refine and optimize antibody performance and minimize immunogenicity when introduced into a human body. In some examples, the humanized antibody will comprise substantially all of at least one, and typically two, variable domains, in which all or substantially all of the CDR regions correspond to those of a non-human immunoglobulin and all or substantially all of the FR regions are those of a human immunoglobulin sequence. The humanized antibody may also comprise at least a portion of an immunoglobulin constant region (Fc), typically that of a human immunoglobulin.
[0116] Humanized antibodies can be engineered to contain human-like immunoglobulin domains, and incorporate only the complementarity-determining regions of the animal-derived antibody. This can be accomplished by carefully examining the sequence of the hyper-variable loops of the variable regions of a monoclonal antigen binding unit or monoclonal antibody, and fitting them to the structure of a human antigen binding unit or human antibody chains. See, e.g., U.S. Pat. No. 6,187,287, incorporated fully herein by reference.
[0117] Methods for humanizing non-human antibodies are well known in the art. "Humanized" antibodies are antibodies in which at least part of the sequence has been altered from its initial form to render it more like human immunoglobulins. In some versions, the heavy (H) chain and light (L) chain constant (C) regions are replaced with human sequence. This can be a fusion polypeptide comprising a variable (V) region and a heterologous immunoglobulin C region. In some versions, the complementarity determining regions (CDRs) comprise non-human antibody sequences, while the V framework regions have also been converted to human sequences. See, for example, EP 0329400. In some versions, V regions are humanized by designing consensus sequences of human and mouse V regions, and converting residues outside the CDRs that are different between the consensus sequences.
[0118] In principle, a framework sequence from a humanized antibody can serve as the template for CDR grafting; however, it has been demonstrated that straight CDR replacement into such a framework can lead to significant loss of binding affinity to the antigen. Glaser et al. (1992) J. Immunol. 149:2606; Tempest et al. (1992) Biotechnology 9:266; and Shalaby et al. (1992) J. Exp. Med. 17:217. The more homologous a human antibody (HuAb) is to the original murine antibody (muAb), the less likely that the human framework will introduce distortions into the murine CDRs that could reduce affinity. Based on a sequence homology search against an antibody sequence database, the HuAb IC4 provides good framework homology to muM4TS.22, although other highly homologous HuAbs would be suitable as well, especially kappa L chains from human subgroup I or H chains from human subgroup III. Kabat et al. (1987). Various computer programs such as ENCAD (Levitt et al. (1983) J. Mol. Biol. 168:595) are available to predict the ideal sequence for the V region. The invention thus encompasses HuAbs with different variable (V) regions. It is within the skill of one in the art to determine suitable V region sequences and to optimize these sequences. Methods for obtaining antibodies with reduced immunogenicity are also described in U.S. Pat. No. 5,270,202 and EP 699,755.
[0119] Humanized antibodies can be prepared by a process of analysis of the parental sequences and various conceptual humanized products using three dimensional models of the parental and humanized sequences. Three dimensional immunoglobulin models are familiar to those skilled in the art. Computer programs are available which illustrate and display probable three-dimensional conformational structures of selected candidate immunoglobulin sequences. Inspection of these displays permits analysis of the likely role of the residues in the functioning of the candidate immunoglobulin sequence, i.e., the analysis of residues that influence the ability of the candidate immunoglobulin to bind its antigen. In this way, FR residues can be selected and combined from the consensus and import sequence so that the desired antibody characteristic, such as increased affinity for the target antigen(s), is achieved.
[0120] A process for humanization of subject antigen binding units can be as follows. The best-fit germline acceptor heavy and light chain variable regions is selected based on homology, canonical structure and physical properties of the human antibody germlines for grafting.
[0121] Computer modeling of mVHIVL versus grafted hVH/VL is performed and prototype humanized antibody sequence is generated. If modeling indicated a need for framework back-mutations, second variant with indicated FW changes is generated. DNA fragments encoding the selected germline frameworks and murine CDRs are synthesized. The synthesized DNA fragments are subcloned into IgG expression vectors and sequences are confirmed by DNA sequencing. The humanized antibodies are expressed in cells, such as 293F and the proteins are tested, for example in MDM phagocytosis assays and antigen binding assays. The humanized antigen binding units are compared with parental antigen binding units in antigen binding affinity, for example, by FACS on cells expressing the target antigen. If the affinity is greater than 2-fold lower than parental antigen binding unit, a second round of humanized variants can be generated and tested as described above.
[0122] As noted above, an antigen binding units can be either "monovalent" or "multivalent." Whereas the former has one binding site per antigen-binding unit, the latter contains multiple binding sites capable of binding to more than one antigen of the same or different kind. Depending on the number of binding sites, antigen binding units may be bivalent (having two antigen-binding sites), trivalent (having three antigen-binding sites), tetravalent (having four antigen-binding sites), and so on.
[0123] Multivalent antigen binding units can be further classified on the basis of their binding specificities. A "monospecific" antigen binding unit is a molecule capable of binding to one or more antigens of the same kind. A "multispecific" antigen binding unit is a molecule having binding specificities for at least two different antigens. While such molecules normally will only bind two distinct antigens (i.e. bispecific antigen binding units), antibodies with additional specificities such as trispecific antibodies are encompassed by this expression when used herein. This disclosure further provides multispecific antigen binding units. Multispecific antigen binding units are multivalent molecules capable of binding to at least two distinct antigens. Preferred multispecific antigen binding units are bispecific and trispecific molecules exhibiting binding specificities to two and three distinct antigens, respectively.
[0124] In some aspects of an embodiment disclosed herein, an antigen binding unit is a bispecific antigen binding unit, wherein the antigen binding unit specifically binds to CD47 and a second antigen. In some examples, the second antigen is not CD47. In some examples, the second antigen is PD1 or PD-Ll. In some examples, the second antigen is other immune checkpoint molecules including CTLA-4, OX40, OX40L, 4-1BB (CD137), CD40, CD40L, ICOS, CD70, CD27, GITR, GITRL, TL1A, TNFRSF25, VISTA, TIM-3, LAG-3, TIGIT, CD112, CD112R, CD226, CD96, B7-H3, B7-H4, CD48, CD244, CD200R, CD200, HVEM, BTLA, CD160, LIGHT, HHLA2, TMIGD2, BTNL2, CD39, CD73, NKG2A, NKG2D, MICA/B, KIR2DL-1, KIR2DL-2, KIR2DL-3, and KIR3DL2. In some examples, the second antigen is EGFR. In some examples, the second antigen is CD19, CD20, CD22, CD33, CD44, CD52, CD79b, CD96, CD97, CD99, CD123, CD138, CD155, CD171, PTHR2, HAVCR2, or other known cancer cell marker. Additional examples of suitable second antigens include, though are not limited to, Fc.gamma.RI, CD 15, p185 HER2, HERS, Fc.gamma.RIII (CD16), CD3, malignant B-cell (1D10), p97, claudin18.2, OVCAR-3, glypican-3, mesothelin, L-D1 (colon carcinoma), Trop2, melanocyte stimulating hormone analog, ErbB2, CAMA1, MoV18, CAIX (carboxy-anhydrase-IX), DCC, UNC5A, MET, TrkC, TrkA, RET, ALK, neural cell adhesion molecule (NCAM), folate binding protein (FBP), GD2, GD3, EpCAM, EGP-40, VEGFR2, MUC-1, MUC-16, STEAP1 (six-transmembrane epithelial antigen of the prostate), PSMA, PSCA (prostate stem cell antigen), GPC-3, LMP-1, DNAM-1 (DNAX accessory molecule-1), pan carcinoma associated antigen (AMOC-31), saporin, Id-1, CD7, CD38, CD30, CD44v7/8, CEA, ricin A chain, interferon-.alpha. (IFN-.alpha.), hybridoma idiotype, vinca alkaloid, alkaline phosphatase, fibrin, tissue plasminogen activator (tPA), urokinase-type plasminogen activator (uPA), low density lipoprotein (LDL), Fc receptor (e.g. Fc.gamma.RI, Fc.gamma.RII or Fc.gamma.RIII), herpes simplex virus (HSV), T-cell receptor, influenza, Fc.gamma.R, HIV, EOTUBE, DPTA, hapten, rabbit IgG, ferritin, horse radish peroxidase (HRP), hormone, somatostatin, substance P, FITC, and beta-galactosidase. Other suitable second antigens include, though are not limited to, a tumor cell antigen, a cytotoxic trigger molecule, a toxin a fibrinolytic agent, a cell surface receptor, infectious disease target, a vaccine adjuvants, a diagnostic agent, a detection molecule, and a reporter molecule.
Polynucleotides and Vectors of the Present Invention
[0125] In some embodiments, the present disclosure provides isolated nucleic acids encoding any of the antigen binding units disclosed herein. In another embodiment, the present disclosure provides vectors comprising a nucleic acid sequence encoding any antigen binding unit disclosed herein. In some embodiments, this invention provides isolated nucleic acids that encode a light-chain CDR and a heavy-chain CDR of an antigen binding unit disclosed herein.
[0126] The subject antigen binding units can be prepared by recombinant DNA technology, synthetic chemistry techniques, or a combination thereof. For instance, sequences encoding the desired components of the antigen binding units, including light chain CDRs and heavy chain CDRs are typically assembled cloned into an expression vector using standard molecular techniques know in the art. These sequences may be assembled from other vectors encoding the desired protein sequence, from PCR-generated fragments using respective template nucleic acids, or by assembly of synthetic oligonucleotides encoding the desired sequences. Expression systems can be created by transfecting a suitable cell with an expressing vector comprising the antigen binding unit of interest.
[0127] Nucleotide sequences corresponding to various regions of light or heavy chains of an existing antibody can be readily obtained and sequenced using convention techniques including but not limited to hybridization, PCR, and DNA sequencing. Hybridoma cells that produce monoclonal antibodies serve as a preferred source of antibody nucleotide sequences. A vast number of hybridoma cells producing an array of monoclonal antibodies may be obtained from public or private repositories. The largest depository agent is American Type Culture Collection (atcc.org), which offers a diverse collection of well-characterized hybridoma cell lines. Alternatively, antibody nucleotides can be obtained from immunized or non-immunized rodents or humans, and form organs such as spleen and peripheral blood lymphocytes. Specific techniques applicable for extracting and synthesizing antibody nucleotides are described in Orlandi et al.(1989) Proc. Natl. Acad. Sci. U.S.A 86: 3833-3837; Larrick et al. (1989) Biochem. Biophys. Res. Commun. 160:1250-1255; Sastry et al. (1989) Proc. Natl. Acad. Sci., U.S.A. 86: 5728-5732; and U.S. Pat. No. 5,969,108.
[0128] Polynucleotides encoding antigen binding units can also be modified, for example, by substituting the coding sequence for human heavy and light chain constant regions in place of the homologous non-human sequences. In that manner, chimeric antibodies are prepared that retain the binding specificity of the original antigen binding unit.
[0129] It is also understood that the polynucleotides embodied in the invention include those coding for functional equivalents and fragments thereof of the exemplified polypeptides.
[0130] Functionally equivalent polypeptides include those that enhance, decrease or not significantly affect properties of the polypeptides encoded thereby. Functional equivalents may be polypeptides having conservative amino acid substitutions, analogs including fusions, and mutants.
[0131] Due to the degeneracy of the genetic code, there can be considerable variation in nucleotides of an antigen binding unit coding sequence, as well as sequences suitable for construction of the polynucleotide and vectors of the present invention. Sequence variants may have modified DNA or amino acid sequences, one or more substitutions, deletions, or additions, the net effect of which is to retain the desired antigen-binding activity. For instance, various substitutions can be made in the coding region that either do not alter the amino acids encoded or result in conservative changes. These substitutions are encompassed by the present invention. Conservative amino acid substitutions include substitutions within the following groups: glycine, alanine; valine, isoleucine, leucine; aspartic acid, glutamic acid; asparagine, glutamine; serine, threonine; lysine, arginine; and phenylalanine, tyrosine. While conservative substitutions do effectively change one or more amino acid residues contained in the polypeptide to be produced, the substitutions are not expected to interfere with the antigen-binding activity of the resulting antigen binding units to be produced. Nucleotide substitutions that do not alter the amino acid residues encoded are useful for optimizing gene expression in different systems. Suitable substitutions are known to those of skill in the art and are made, for instance, to reflect preferred codon usage in the expression systems.
[0132] Where desired, the recombinant polynucleotides may comprise heterologous sequences that facilitate detection of the expression and purification of the gene product. Examples of such sequences are known in the art and include those encoding reporter proteins such as .beta.-galactosidase, .beta.-lactamase, chloramphenicol acetyltransferase (CAT), luciferase, green fluorescent protein (GFP) and their derivatives. Other heterologous sequences that facilitate purification may code for epitopes such as Myc, HA (derived from influenza virus hemagglutinin), His-6, FLAG, or the Fc portion of immunoglobulin, glutathione S-transferase (GST), and maltose-binding protein (MBP).
[0133] Polynucleotides disclosed herein can be conjugated to a variety of chemically functional moieties described above. Commonly employed moieties include labels capable of producing a detectable signal, signal peptides, agents that enhance immunologic reactivity, agents that facilitate coupling to a solid support, vaccine carriers, bioresponse modifiers, paramagnetic labels and drugs. The moieties can be covalently linked polynucleotide recombinantly or by other means known in the art.
[0134] Polynucleotides of the invention can comprise additional sequences, such as additional encoding sequences within the same transcription unit, controlling elements such as promoters, ribosome binding sites, and polyadenylation sites, additional transcription units under control of the same or a different promoter, sequences that permit cloning, expression, and transformation of a host cell, and any such construct as may be desirable to provide embodiments of this invention.
[0135] Polynucleotides embodied in this invention can be obtained using chemical synthesis, recombinant cloning methods, PCR, or any combination thereof. Methods of chemical polynucleotide synthesis are well known in the art and need not be described in detail herein. One of skill in the art can use the sequence data provided herein to obtain a desired polynucleotide by employing a DNA synthesizer or ordering from a commercial service.
[0136] Polynucleotides comprising a desired sequence can be inserted into a suitable vector which in turn can be introduced into a suitable host cell for replication and amplification. Accordingly, the invention encompasses a variety of vectors comprising one or more of the polynucleotides of the present invention. Also provided are selectable libraries of expression vectors comprising at least one vector encoding an antigen binding units disclosed herein.
[0137] Vectors of the present invention generally comprises a transcriptional or translational control sequences required for expressing the antigen binding units. Suitable transcription or translational control sequences include but are not limited to replication origin, promoter, enhancer, repressor binding regions, transcription initiation sites, ribosome binding sites, translation initiation sites, and termination sites for transcription and translation.
[0138] The choice of promoters will largely depend on the host cells in which the vector is introduced. It is also possible, to utilize promoters normally associated with a desired light or heavy chain gene, provided that such control sequences are compatible with the host cell system. Cell-specific or tissue-specific promoters may also be used. A vast diversity of tissue specific promoters have been described and employed by artisans in the field. Exemplary promoters operative in selective animal cells include hepatocyte-specific promoters and cardiac muscle specific promoters. Depending on the choice of the recipient cell types, those skilled in the art will know of other suitable cell-specific or tissue-specific promoters applicable for the construction of the expression vectors of the present invention.
[0139] Using known molecular cloning or gene engineering techniques, appropriate transcriptional control sequences, enhancers, terminators, or any other genetic element known in the art can integrated in operative relationship, optionally additionally with intact selectable fusion genes to be expressed in accordance with the present invention. In addition to the above-described elements, the vectors may contain a selectable marker (for example, a gene encoding a protein necessary for the survival or growth of a host cell transformed with the vector), although such a marker gene can be carried on another polynucleotide sequence co-introduced into the host cell.
[0140] The polynucleotides and vectors of this invention have several specific uses. They are useful, for example, in expression systems for the production of antigen binding units. Such polynucleotides are useful as primers to effect amplification of desired polynucleotides. Furthermore, polynucleotides of this invention are also useful in pharmaceutical compositions including vaccines, diagnostics, and drugs.
[0141] The host cells of this invention can be used, inter alia, as repositories of the subject polynucleotides, vectors, or as vehicles for producing and screening desired antigen binding units based on their antigen binding specificities.
[0142] Accordingly, the invention provides a method of identifying an antigen binding unit that is immunoreactive with a desired antigen. Such a method can involve the following steps: (a) preparing a genetically diverse library of antigen binding units, wherein the library comprises at least one subject antigen binding unit; (b) contacting the library of antigen binding units with the desired antigen; (c) detecting a specific binding between antigen binding units and the antigen, thereby identifying the antigen binding unit that is immunoreactive with the desired antigen.
[0143] The ability of an antigen binding unit to specifically bind to a desired antigen can be tested by a variety of procedures well established in the art. See Harlow and Lane (1988) Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, New York; Gherardi et al. (1990) J. Immunol. Meth. 126:61-68. Typically, antigen binding units exhibiting desired binding specificities can be detected directly by immunoassays, for example, by reacting labeled antigen binding units with the antigens that are immobilized on a solid support or substrate. In general, the substrate to which the antigen is adhered is fabricated with material exhibiting a low level of non-specific binding during immunoassay. An example solid support is made from one or more of the following types of materials: plastic polymers, glass, cellulose, nitrocellulose, semi-conducting material, and metal. In some examples, the substrate is petri dish, chromatography beads, magnetic beads, and the like.
[0144] For such solid-phase assays, the unreacted antigen binding units are removed by washing. In a liquid-phase assay, however, the unreacted antigen binding units are removed by some other separation technique, such as filtration or chromatography. After binding the antigen to the labeled antigen binding units, the amount of bound label is determined. A variation of this technique is a competitive assay, in which the antigen is bound to saturation with an original binding molecule. When a population of the subject antigen binding unit is introduced to the complex, only those that exhibit higher binding affinity will be able to compete, and thus remain bound to the antigen.
[0145] Alternatively, specific binding to a given antigen can be assessed by cell sorting, which involves presenting the desired antigen on the cells to be sorted, then labeling the target cells with antigen binding units that are coupled to detectable agents, followed by separating the labeled cells from the unlabeled ones in a cell sorter. A sophisticated cell separation method is fluorescence-activated cell sorting (FACS). Cells traveling in single file in a fine stream are passed through a laser beam, and the fluorescence of each cell bound by the fluorescently labeled antigen binding unit is then measured.
[0146] Subsequent analysis of the eluted antigen binding units may involve protein sequencing for delineating the amino acid sequences of the light chains and heavy chains. Based on the deduced amino acid sequences, the cDNA encoding the antibody polypeptides can then be obtained by recombinant cloning methods including PCR, library screening, homology searches in existing nucleic acid databases, or any combination thereof. Commonly employed databases include but are not limited to GenBank, EMBL, DDBJ, PDB, SWISS-PROT, EST, STS, GSS, and HTGS.
[0147] When a library of antigen binding unit is displayed on phage or bacterial particles, selection is preferably performed using affinity chromatography. The method typically proceeds with binding a library of phage antigen binding units to an antigen coated plates, column matrices, cells or to biotinylated antigen in solution followed by capture. The phages or bacteria bound to the solid phase are washed and then eluted by soluble hapten, acid or alkali. Alternatively, increasing concentrations of antigen can be used to dissociate the antigen binding units from the affinity matrix. For certain antigen binding units with extremely high affinity or avidity to the antigen, efficient elution may require high pH or mild reducing solution as described in WO 92/01047.
[0148] The efficiency of selection is likely to depend on a combination of several factors, including the kinetics of dissociation during washing, and whether multiple antigen binding units on a single phage or bacterium can simultaneously bind to antigens on a solid support. For example, antibodies with fast dissociation kinetics (and weak binding affinities) can be retained by use of short washes, multivalent display and a high coating density of antigen at the solid support. Conversely, the selection of antigen binding units with slow dissociation kinetics (and good binding affinities) can be favored by use of long washes, monovalent phages, and a low coating density of antigen.
[0149] Where desired, the library of antigen binding units can be pre-selected against an unrelated antigen to counter-select the undesired antigen binding units. The library may also be pre-selected against a related antigen in order to isolate, for example, anti-idiotypic antigen binding units.
Host Cells of the Present Invention
[0150] In some embodiments, the present disclosure provides host cells expressing any one of the antigen binding units disclosed herein. A subject host cell typically comprises a nucleic acid encoding any one of the antigen binding units disclosed herein.
[0151] The invention provides host cells transfected with the polynucleotides, vectors, or a library of the vectors described above. The vectors can be introduced into a suitable prokaryotic or eukaryotic cell by any of a number of appropriate means, including electroporation, microprojectile bombardment; lipofection, infection (where the vector is coupled to an infectious agent), transfection employing calcium chloride, rubidium chloride, calcium phosphate, DEAE-dextran, or other substances. The choice of the means for introducing vectors will often depend on features of the host cell.
[0152] For most animal cells, any of the above-mentioned methods is suitable for vector delivery. Preferred animal cells are vertebrate cells, preferably mammalian cells, capable of expressing exogenously introduced gene products in large quantity, e.g. at the milligram level.
[0153] Non-limiting examples of preferred cells are NIH3T3 cells, COS, HeLa, and CHO cells.
[0154] Once introduced into a suitable host cell, expression of the antigen binding units can be determined using any nucleic acid or protein assay known in the art. For example, the presence of transcribed mRNA of light chain CDRs or heavy chain CDRs, or the antigen binding unit can be detected and/or quantified by conventional hybridization assays (e.g. Northern blot analysis), amplification procedures (e.g. RT-PCR), SAGE (U.S. Pat. No. 5,695,937), and array-based technologies (see e.g. U.S. Pat. Nos. 5,405,783, 5,412,087 and 5,445,934), using probes complementary to any region of antigen binding unit polynucleotide.
[0155] Expression of the vector can also be determined by examining the antigen binding unit expressed. A variety of techniques are available in the art for protein analysis. They include but are not limited to radioimmunoassays, ELISA (enzyme linked immunoradiometric assays), "sandwich" immunoassays, immunoradiometric assays, in situ immunoassays (using e.g., colloidal gold, enzyme or radioisotope labels), western blot analysis, immunoprecipitation assays, immunoflourescent assays, and SDS-PAGE.
Preparation of Antigen-Binding Units
[0156] In some embodiments, the present disclosure provides methods of producing any antigen binding unit disclosed herein, wherein the method comprises culturing host cells expressing the antigen binding unit under conditions suitable for expressing the antigen binding unit, and isolating the antigen binding unit expressed by the host cell.
[0157] The expressed antigen binding units can be isolated using a variety of protein purification techniques known in the art. Generally, the antigen binding unit is isolated from culture media as secreted polypeptides, although they can be recovered from host cell lysates or bacterial periplasm, when directly produced without signal peptides. If the antigen binding units are membrane-bound, they can be solubilized by suitable detergent solutions commonly employed by artisans in the field. The recovered antigen binding units may be further purified by salt precipitation (e.g., with ammonium sulfate), ion exchange chromatography (e.g. on a cationic or anionic exchange column run at neutral pH and eluted with step gradients of increasing ionic strength), gel filtration chromatography (including gel filtration HPLC), and chromatography on tag-affinity column, or on affinity resins such as protein A, protein G, hydroxyapatite, and anti-immunoglobulin.
[0158] In addition, derivatized immunoglobulins with added chemical linkers, detectable moieties such as fluorescent dyes, enzymes, substrates, chemiluminescent moieties, specific binding moieties such as streptavidin, avidin, or biotin, or drug conjugates can be utilized in the methods and compositions of the present invention.
[0159] Additionally disclosed herein are antigen binding unites conjugated to a chemically functional moiety. Typically, the moiety is a label capable of producing a detectable signal. These conjugated antigen binding units are useful, for example, in detection systems such as quantitation of tumor burden, and imaging of metastatic foci and tumor imaging. Such labels are known in the art and include, but are not limited to, radioisotopes, enzymes, fluorescent compounds, chemiluminescent compounds, bioluminescent compounds substrate cofactors and inhibitors. See, for examples of patents teaching the use of such labels, U.S. Pat. Nos. 3,817,837; 3,850,752; 3,939,350; 3,996,345; 4,277,437; 4,275,149; and 4,366,241. The moieties can be covalently linked to antigen binding units, recombinantly linked, or conjugated to antigen binding units through a secondary reagent, such as a second antibody, protein A, or a biotin-avidin complex.
[0160] Other functional moieties include signal peptides, agents that enhance immunologic reactivity, agents that facilitate coupling to a solid support, vaccine carriers, bioresponse modifiers, paramagnetic labels and drugs. Signal peptides is a short amino acid sequence that directs a newly synthesized protein through a cellular membrane, usually the endoplasmic reticulum in eukaryotic cells, and either the inner membrane or both inner and outer membranes of bacteria. Signal peptides can be at the N-terminal portion of a polypeptide or the C-terminal portion of a polypeptide, and can be removed enzymatically between biosynthesis and secretion of the polypeptide from the cell. Such a peptide can be incorporated into an antigen binding units to allow secretion of the synthesized molecules.
[0161] Agents that enhance immunologic reactivity include, but are not limited to, bacterial superantigens. Agents that facilitate coupling to a solid support include, but are not limited to, biotin or avidin. Immunogen carriers include, but are not limited to, any physiologically acceptable buffers. Bioresponse modifiers include cytokines, particularly tumor necrosis factor (TNF), interleukin-2, interleukin-4, granulocyte macrophage colony stimulating factor and .gamma.-interferons.
[0162] Suitable drug moieties include antineoplastic agents. Non-limiting examples include radioisotopes, vinca alkaloids such as the vinblastine, vincristine and vindesine sulfates, adriamycin, bleomycin sulfate, carboplatin, cisplatin, cyclophosphamide, cytarabine, dacarbazine, dactinomycin, duanorubicin hydrochloride, doxorubicin hydrochloride, etoposide, fluorouracil, lomustine, mechlororethamine hydrochloride, melphalan, mercaptopurine, methotrexate, mitomycin, mitotane, pentostatin, pipobroman, procarbaze hydrochloride, streptozotocin, taxol, thioguanine, and uracil mustard.
[0163] Immunotoxins, including antigen binding units, can be produced by recombinant means. Production of various immunotoxins is well-known in the art, and methods can be found, for example, in "Monoclonal Antibody-toxin Conjugates: Aiming the Magic Bullet," Thorpe et al. (1982) Monoclonal Antibodies in Clinical Medicine, Academic Press, pp. 168-190; Vitatta (1987) Science 238:1098-1104; and Winter and Milstein (1991) Nature 349:293-299. Suitable toxins include, but are not limited to, ricin, radionuclides, pokeweed antiviral protein, Pseudomonas exotoxin A, diphtheria toxin, ricin A chain, fungal toxins such as restrictocin and phospholipase enzymes. See, generally, "Chimeric Toxins," Olsnes and Pihl, Pharmac. Ther. 15:355-381 (1981); and "Monoclonal Antibodies for Cancer Detection and Therapy," eds. Baldwin and Byers, pp. 159-179, 224-266, Academic Press (1985).
[0164] Chemically functional moieties can be made recombinantly for instance by creating a fusion gene encoding the antigen binding unit and the functional moiety. Alternatively, the antigen binding unit can be chemically bonded to the moiety by any of a variety of well-established chemical procedures. For example, when the moiety is a protein, the linkage can be by way of heterobifunctional cross linkers, e.g., SPDP, carbodiimide glutaraldehyde, or the like. The moieties can be covalently linked, or conjugated, through a secondary reagent, such as a second antibody, protein A, or a biotin-avidin complex. Paramagnetic moieties and the conjugation thereof to antibodies are well-known in the art. See, e.g., Miltenyi et al. (1990) Cytometry 11:231-238.
Methods of Use and Treatment
[0165] CD47-specific antigen binding units and pharmaceutical compositions comprising the same can find a wide variety of applications, including, but not limited to, treatment and diagnosis.
[0166] In one embodiment, the present disclosure provides pharmaceutical compositions comprising a pharmaceutically acceptable excipient and any of the antigen binding units disclosed herein.
[0167] In another embodiment, the present disclosure provides methods of inducing phagocytosis of cells expressing CD47, said method comprising contacting the cell expressing CD47 with any antigen binding unit disclosed herein. In some aspects, the cell is a cancer cell. In some aspects, the cell is a non-lymphoma cancer cell. In some aspects, the cell is a non-leukemia cancer cell. In some aspects, the cell is a non-lymphoma and non-leukemia cancer cell. In some aspects, the cell is a hematological cancer cell. Hematological cancers include, but are not limited to, leukemia, lymphoma and myeloma. Certain forms of leukemia include, by way of non-limiting example, acute lymphocytic leukemia (ALL); acute myeloid leukemia (AML); chronic lymphocytic leukemia (CLL); chronic myelogenous leukemia (CML); Myeloproliferative disorder/neoplasm (MPDS); and myelodysplasia syndrome. Certain forms of lymphoma include, by way of non-limiting example, Hodgkin's lymphoma, both indolent and aggressive non-Hodgkin's lymphoma, Burkitt's lymphoma, and follicular lymphoma (small cell and large cell). Certain forms of myeloma include, by way of non-limiting example, multiple myeloma (MM), giant cell myeloma, heavy-chain myeloma, and light chain or Bence-Jones myeloma. Solid tumors include, e.g., breast tumors, ovarian tumors, lung tumors, pancreatic tumors, prostate tumors, melanoma tumors, colorectal tumors, lung tumors, head and neck tumors, bladder tumors, leiomyosarcoma, glioma, glioblastoma, brain tumors, esophageal tumors, gastric tumors, liver tumors, and kidney tumors.
[0168] In yet another embodiment, the present disclosure provides methods of inducing phagocytosis of cells expressing CD47 in a human subject, said method comprising administering to the human subject a pharmaceutical composition comprising a pharmaceutically acceptable excipient and any antigen binding unit disclosed herein. In some aspects, the cell is a cancer cell. In some aspects, the cell is a non-lymphoma cancer cell. In some aspects, the cell is a non-leukemia cancer cell. In some aspects, the cell is a non-lymphoma and non-leukemia cancer cell. In some aspects, the cell is a hematological cancer cell. Hematological cancers include, but are not limited to, leukemia, lymphoma and myeloma. Certain forms of leukemia include, by way of non-limiting example, acute lymphocytic leukemia (ALL); acute myeloid leukemia (AML); chronic lymphocytic leukemia (CLL); chronic myelogenous leukemia (CML); Myeloproliferative disorder/neoplasm (MPDS); and myelodysplasia syndrome. Certain forms of lymphoma include, by way of non-limiting example, Hodgkin's lymphoma, both indolent and aggressive non-Hodgkin's lymphoma, Burkitt's lymphoma, and follicular lymphoma (small cell and large cell). Certain forms of myeloma include, by way of non-limiting example, multiple myeloma (MM), giant cell myeloma, heavy-chain myeloma, and light chain or Bence-Jones myeloma. Solid tumors include, e.g., breast tumors, ovarian tumors, lung tumors, pancreatic tumors, prostate tumors, melanoma tumors, colorectal tumors, lung tumors, head and neck tumors, bladder tumors, leiomyosarcoma, glioma, glioblastoma, brain tumors, esophageal tumors, gastric tumors, liver tumors, and kidney tumors.
[0169] In some aspects of methods of inducing phagocytosis as disclosed herein, phagocytosis of cells expressing CD47 occurs with an efficiency within the range of 1% to 100%. In some examples, phagocytosis occurs with about 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% efficiency. In some examples, phagocytosis occurs with at least about 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% efficiency. In some aspects of any of the embodiments disclosed herein, an antigen binding unit induces phagocytosis of cells expressing CD47 to a greater extent than that of a reference antigen binding unit. Such a reference antigen binding unit can have the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245, or any other known anti-CD47 antigen binding unit. Phagocytosis extent can be determined by any method known in the art. In some cases, the extend of phagocytosis is determined the number of macrophages that have performed phagocytosis (referred to as phagocytes) among a population of macrophages. For example, the number of phagocytes per 100 macrophages can be determined and thereby the extend of phagocytosis can be expressed as a percentage or phagocytic index.
[0170] In some aspects of methods of inducing phagocytosis as disclosed herein, the antigen binding unit used in said method does not cause significant hemagglutination. In some cases, hemagglutination induced in said method using any of the antigen binding units disclosed herein is at least 1 fold less as compared to that induced by an reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. In some cases, hemagglutination induced upon contacting the red blood cells with a subject antigen binding unit is at least 1 fold less, at least 2 fold less, at least 3 fold less, at least 4 fold less, at least 5 fold less, at least 6 fold less, at least 7 fold less, at least 8 fold less, at least 9 fold less, or at least 10 fold less as compared to that induced by a reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245. In some cases, hemagglutination induced upon contacting the red blood cells with a subject antigen binding unit is greater than at least 10 fold less as compared to that induced by a reference antigen binding unit having the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245.
[0171] In some embodiments, the present disclosure provides methods of treating cancer in a subject in need thereof. In some aspects, the method comprises administering to the subject in need thereof, an effective amount of any of the antigen binding units disclosed herein. In some aspects, the cancer is a non-lymphoma cancer. In some aspects, the cancer is a non-leukemia cancer. In some aspects, the cancer is a non-lymphoma and non-leukemia cancer. In some aspects, the cell is a hematological cancer cell. Hematological cancers include, but are not limited to, leukemia, lymphoma and myeloma. Certain forms of leukemia include, by way of non-limiting example, acute lymphocytic leukemia (ALL); acute myeloid leukemia (AML); chronic lymphocytic leukemia (CLL); chronic myelogenous leukemia (CML); Myeloproliferative disorder/neoplasm (MPDS); and myelodysplasia syndrome. Certain forms of lymphoma include, by way of non-limiting example, Hodgkin's lymphoma, both indolent and aggressive non-Hodgkin's lymphoma, Burkitt's lymphoma, and follicular lymphoma (small cell and large cell). Certain forms of myeloma include, by way of non-limiting example, multiple myeloma (MM), giant cell myeloma, heavy-chain myeloma, and light chain or Bence-Jones myeloma. Solid tumors include, e.g., breast tumors, ovarian tumors, lung tumors, pancreatic tumors, prostate tumors, melanoma tumors, colorectal tumors, lung tumors, head and neck tumors, bladder tumors, leiomyosarcoma, glioma, glioblastoma, brain tumors, esophageal tumors, gastric tumors, liver tumors, and kidney tumors. In most cases, the effective amount is determined empirically via testing methods well known in the art.
[0172] In some embodiments, the present disclosure provides methods of treating cancer in a subject in need thereof. In some aspects, the method comprises administering to the subject in need thereof, an effective amount of a pharmaceutical comprising a pharmaceutically acceptable excipient and any of the antigen binding units disclosed herein. In some aspects, the cancer is a non-lymphoma cancer. In some aspects, the cancer is a non-leukemia cancer. In some aspects, the cancer is a non-lymphoma and non-leukemia cancer. In some aspects, the cell is a hematological cancer cell. Hematological cancers include, but are not limited to, leukemia, lymphoma and myeloma. Certain forms of leukemia include, by way of non-limiting example, acute lymphocytic leukemia (ALL); acute myeloid leukemia (AML); chronic lymphocytic leukemia (CLL); chronic myelogenous leukemia (CML); Myeloproliferative disorder/neoplasm (MPDS); and myelodysplasia syndrome. Certain forms of lymphoma include, by way of non-limiting example, Hodgkin's lymphoma, both indolent and aggressive non-Hodgkin's lymphoma, Burkitt's lymphoma, and follicular lymphoma (small cell and large cell). Certain forms of myeloma include, by way of non-limiting example, multiple myeloma (MM), giant cell myeloma, heavy-chain myeloma, and light chain or Bence-Jones myeloma. Solid tumors include, e.g., breast tumors, ovarian tumors, lung tumors, pancreatic tumors, prostate tumors, melanoma tumors, colorectal tumors, lung tumors, head and neck tumors, bladder tumors, leiomyosarcoma, glioma, glioblastoma, brain tumors, esophageal tumors, gastric tumors, liver tumors, and kidney tumors. In most cases, the effective amount is determined empirically via testing methods well known in the art.
[0173] Cancers of interest for treatment by the methods of the invention include, but are not limited to, leukemias; acute leukemias such as T-ALL, B-ALL, AML, etc.; lymphomas (Hodgkin's and non-Hodgkin's); sarcomas; melanomas; adenomas; carcinomas of solid tissue including ovarian carcinoma, breast carcinoma, pancreatic carcinoma, colon carcinoma, squamous cell carcinoma, transitional cell carcinoma, etc., hypoxic tumors, squamous cell carcinomas of the mouth, throat, larynx, and lung, genitourinary cancers such as cervical and bladder cancer, hematopoietic cancers, head and neck cancers, and nervous system cancers, such as gliomas, astrocytomas, meningiomas, etc., benign lesions such as papillomas, and the like.
[0174] Treatment of cancer can be evidenced by reducing growth of cancer cells including, but is not limited to, reducing proliferation of cancer cells, and reducing the incidence of a non-cancerous cell becoming a cancerous cell. Whether a reduction in cancer cell growth has been achieved can be readily determined using any known assay, including, but not limited to, [.sup.3H]-thymidine incorporation; counting cell number over a period of time; detecting and/or measuring a marker associated with AML, etc. Whether a substance, or a specific amount of the substance, is effective in treating cancer can be assessed using any of a variety of known diagnostic assays for cancer, including, but not limited to biopsy, contrast radiographic studies, CAT scan, and detection of a tumor marker associated with cancer in the blood of the individual. The substance can be administered systemically or locally, usually systemically.
[0175] In some aspects, treatment of cancer can be evidenced by reduced tumor volume.
[0176] Tumor volume can be determined using any known method in the field. For example, tumor volume can be determined by measuring the tumor using a caliper. In such cases, two dimensions of the tumor can be measured and tumor volume can be determined using the formula V=0.5 a.times.b.sup.2, where a and b are a first and second diameter. In some cases, the first diameter is the long diameter or the larger of the two diameters. In some cases, the second diameter is the short diameter or the smaller of the two diameters.
[0177] In some aspects, treatment of cancer can be evidenced by reduced tumor volume. In some cases, tumor volume is reduced by a percentage within the range of 1% to 100%. In some examples, tumor volume is reduced by about 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%. In some examples, tumor volume is reduced by at least about 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%. In some aspects of any of the embodiments disclosed herein, an antigen binding unit can reduce the tumor volume to a greater extent than that of a reference antigen binding unit. Such a reference antigen binding unit can have the amino acid sequences shown in 1) SEQ ID NO: 240-241, 2) SEQ ID NO:242-243, or 3) SEQ ID NO:244-245, or any other known anti-CD47 antigen binding unit.
[0178] In some aspects, comparison of the effect of a subject antigen binding unit compared to a reference antigen binding unit can be determined by calculating anti-tumor effectiveness. In such cases, tumor volume can be measure such as described above. Alternatively, a different parameter of tumor size or other appropriate characteristic of the tumor can be determined or measured. When working with quantifiable characteristics such as tumor volume, the anti-tumor effectiveness can be determined by using the formula: T/C, where T is the selected measurement (e.g., tumor volume) for the treatment group and C is the selected measurement (e.g., tumor volume) for the control group. Anti-tumor effectiveness can be determined over any desired period of time and can be determined using average value from any desired number of samples.
[0179] Anti-tumor effectiveness can be expressed as a number or a percent. In some examples, anti-tumor effectiveness can be about 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%. In some examples, anti-tumor effectiveness can be at most 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%. In some examples, anti-tumor effectiveness can be at least 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%.
[0180] Compositions, e.g. antigen binding units and pharmaceutical compositions, disclosed herein can be administered using any medically appropriate procedure, e.g., intravascular (intravenous, intra-arterial, intra-capillary) administration, injection into the lymph nodes, etc. Intravascular injection may be by intravenous or intraarterial injection. An effective amount of a composition to be given to a particular patient will depend on a variety of factors, several of which will be different from patient to patient and can be determined empirically. Dosage of the composition will depend on the determined treatment regime, route of administration, the nature of the therapeutics, sensitivity of the tumor to the therapeutics, etc. Utilizing LD.sub.50 animal data, and other information available for an antigen binding unit disclosed herein, a clinician can determine the maximum safe dose for an individual, depending on the route of administration. For instance, an intravenously administered dose may be more than an locally administered dose, given the greater body of fluid into which the therapeutic composition is being administered. Similarly, compositions which are rapidly cleared from the body may be administered at higher doses, or in repeated doses, in order to maintain a therapeutic concentration. Utilizing ordinary skill, the competent clinician will be able to optimize the dosage of a particular composition.
[0181] In some aspects, a method for treatment of cancer is provided, the method comprising administering an effective amount of any of the foregoing pharmaceutical compositions comprising a subject antigen binding unit to a subject in need thereof. In some embodiments, the cancer is leukemia, breast cancer, prostate cancer, pancreatic cancer, lung cancer, including non-small cell lung cancer or carcinoma, liver cancer, ovarian cancer, bladder cancer, head and neck cancer, colorectal cancer, skin cancer, a brain tumor, or hepatocellular carcinoma (HCC). In certain aspects, the cancer is leukemia. The cancer can be a solid tumor. The cancer can be Mixed Lineage Leukemia (MLL), Acute lymphoblastic leukemia (ALL), Acute myelogenous leukemia (AML), Chronic lymphocytic leukemia (CLL), small lymphocytic lymphoma (SLL), Chronic myelogenous leukemia (CML), Acute monocytic leukemia (AML), hairy cell leukemia, and/or other leukemias; myeloproliferative disorder/neoplasm (MPDS), giant cell myeloma, heavy-chain myeloma, light chain or Bence-Jones Myeloma, myelodysplasia syndrome, multiple myeloma. The cancer can be lymphomas such as all subtypes of Hodgkin's lymphoma or non-Hodgkin's lymphoma.
[0182] The present disclosure also provides methods for combination therapies in which an agent known to modulate other pathways, or other components of the same pathway, or even overlapping sets of target enzymes are used in combination with a subject antigen binding unit or pharmaceutical composition comprising a subject antigen binding unit. In one aspect, such therapy includes but is not limited to the combination of one or more antigen binding units of the disclosure with chemotherapeutic agents, therapeutic antibodies, and radiation treatment, to provide a synergistic or additive therapeutic effect.
[0183] Where desired, a subject antigen binding unit can be used in combination with Notch inhibitors and/or c-Myb inhibitors. Where desired, an antigen binding unit or pharmaceutical composition of the present disclosure can be used in combination with MLL-WDR5 inhibitors and/or Dot11 inhibitors.
[0184] Many chemotherapeutics are presently known in the art and can be used in combination with a subject antigen binding unit. In some embodiments, the chemotherapeutic is selected from the group consisting of mitotic inhibitors, alkylating agents, anti-metabolites, intercalating antibiotics, growth factor inhibitors, cell cycle inhibitors, enzymes, topoisomerase inhibitors, biological response modifiers, anti-hormones, angiogenesis inhibitors, and anti-androgens.
[0185] Non-limiting examples are chemotherapeutic agents, cytotoxic agents, and non-peptide small molecules such as Gleevec.RTM. (Imatinib Mesylate), Velcade.RTM. (bortezomib), Casodex (bicalutamide), Iressa.RTM. (gefitinib), and Adriamycin as well as a host of chemotherapeutic agents. Non-limiting examples of chemotherapeutic agents include alkylating agents such as thiotepa and cyclosphosphamide (CYTOXAN.TM.); alkyl sulfonates such as busulfan, improsulfan and piposulfan; aziridines such as benzodopa, carboquone, meturedopa, and uredopa; ethylenimines and methylamelamines including altretamine, triethylenemelamine, trietylenephosphoramide, triethylenethiophosphaoramide and trimethylolomelamine; nitrogen mustards such as chlorambucil, chlornaphazine, cholophosphamide, estramustine, ifosfamide, mechlorethamine, mechlorethamine oxide hydrochloride, melphalan, novembichin, phenesterine, prednimustine, trofosfamide, uracil mustard; nitrosureas such as carmustine, chlorozotocin, fotemustine, lomustine, nimustine, ranimustine; antibiotics such as aclacinomysins, actinomycin, authramycin, azaserine, bleomycins, cactinomycin, calicheamicin, carabicin, carminomycin, carzinophilin, Casodex.TM., chromomycins, dactinomycin, daunorubicin, detorubicin, 6-diazo-5-oxo-L-norleucine, doxorubicin, epirubicin, esorubicin, idarubicin, marcellomycin, mitomycins, mycophenolic acid, nogalamycin, olivomycins, peplomycin, potfiromycin, puromycin, quelamycin, rodorubicin, streptonigrin, streptozocin, tubercidin, ubenimex, zinostatin, zorubicin; anti-metabolites such as methotrexate and 5-fluorouracil (5-FU); folic acid analogues such as denopterin, methotrexate, pteropterin, trimetrexate; purine analogs such as fludarabine, 6-mercaptopurine, thiamiprine, thioguanine; pyrimidine analogs such as ancitabine, azacitidine, 6-azauridine, carmofur, cytarabine, dideoxyuridine, doxifluridine, enocitabine, floxuridine, androgens such as calusterone, dromostanolone propionate, epitiostanol, mepitiostane, testolactone; anti-adrenals such as aminoglutethimide, mitotane, trilostane; folic acid replenisher such as frolinic acid; aceglatone; aldophosphamide glycoside; aminolevulinic acid; amsacrine; bestrabucil; bisantrene; edatraxate; defofamine; demecolcine; diaziquone; elfomithine; elliptinium acetate; etoglucid; gallium nitrate; hydroxyurea; lentinan; lonidamine; mitoguazone; mitoxantrone; mopidamol; nitracrine; pentostatin; phenamet; pirarubicin; podophyllinic acid; 2-ethylhydrazide; procarbazine; PSK.RTM.; razoxane; sizofiran; spirogermanium; tenuazonic acid; triaziquone; 2,2',2''-trichlorotriethylamine; urethan; vindesine; dacarbazine; mannomustine; mitobronitol; mitolactol; pipobroman; gacytosine; arabinoside ("Ara-C"); cyclophosphamide; thiotepa; taxanes, e.g., paclitaxel (TAXOLTM, Bristol-Myers Squibb Oncology, Princeton, N.J.) and docetaxel (TAXOTERETM, Rhone-Poulenc Rorer, Antony, France); retinoic acid; esperamicins; capecitabine; and pharmaceutically acceptable salts, acids or derivatives of any of the above. Also included as suitable chemotherapeutic cell conditioners are anti-hormonal agents that act to regulate or inhibit hormone action on tumors such as anti-estrogens including for example tamoxifen, (Nolvadex.TM.), raloxifene, aromatase inhibiting 4(5)-imidazoles, 4-hydroxytamoxifen, trioxifene, keoxifene, LY 117018, onapristone, and toremifene (Fareston); and anti-androgens such as flutamide, nilutamide, bicalutamide, leuprolide, and goserelin; chlorambucil; gemcitabine; 6-thioguanine; mercaptopurine; methotrexate; platinum analogs such as cisplatin and carboplatin; vinblastine; platinum; etoposide (VP-16); ifosfamide; mitomycin C; mitoxantrone; vincristine; vinorelbine; navelbine; novantrone; teniposide; daunomycin; aminopterin; xeloda; ibandronate; camptothecin-11 (CPT-11); topoisomerase inhibitor RFS 2000; difluoromethylornithine (DMFO). Where desired, the antigen binding units or pharmaceutical composition of the present disclosure can be used in combination with commonly prescribed anti-cancer drugs such as Herceptin.RTM., Avastin.RTM., Erbitux.RTM., Rituxan.RTM., Taxol.RTM., Arimidex.RTM., Taxotere.RTM., ABVD, AVICINE, Abagovomab, Acridine carboxamide, Adecatumumab, 17-N-Allylamino-17-demethoxygeldanamycin, Alpharadin, Alvocidib, 3-Aminopyridine-2-carboxaldehyde thiosemicarbazone, Amonafide, Anthracenedione, Anti-CD22 immunotoxins, Antineoplastic, Antitumorigenic herbs, Apaziquone, Atiprimod, Azathioprine, Belotecan, Bendamustine, BIBW 2992, Biricodar, Brostallicin, Bryostatin, Buthionine sulfoximine, CBV (chemotherapy), Calyculin, cell-cycle nonspecific antineoplastic agents, Dichloroacetic acid, Discodermolide, Elsamitrucin, Enocitabine, Epothilone, Eribulin, Everolimus, Exatecan, Exisulind, Ferruginol, Forodesine, Fosfestrol, ICE chemotherapy regimen, IT-101, Imexon, Imiquimod, Indolocarbazole, Irofulven, Laniquidar, Larotaxel, Lenalidomide, Lucanthone, Lurtotecan, Mafosfamide, Mitozolomide, Nafoxidine, Nedaplatin, Olaparib, Ortataxel, PAC-1, Pawpaw, Pixantrone, Proteasome inhibitor, Rebeccamycin, Resiquimod, Rubitecan, SN-38, Salinosporamide A, Sapacitabine, Stanford V, Swainsonine, Talaporfin, Tariquidar, Tegafur-uracil, Temodar, Tesetaxel, Triplatin tetranitrate, Tris(2-chloroethyl)amine, Troxacitabine, Uramustine, Vadimezan, Vinflunine, ZD6126 or Zosuquidar.
[0186] This disclosure further relates to a method for using a subject antigen binding unit or a pharmaceutical composition provided herein, in combination with radiation therapy for inhibiting abnormal cell growth or treating the hyperproliferative disorder in the mammal. Techniques for administering radiation therapy are known in the art, and these techniques can be used in the combination therapy described herein.
[0187] Radiation therapy can be administered through one of several methods, or a combination of methods, including without limitation external-beam therapy, internal radiation therapy, implant radiation, stereotactic radiosurgery, systemic radiation therapy, radiotherapy and permanent or temporary interstitial brachytherapy. The term "brachytherapy," as used herein, refers to radiation therapy delivered by a spatially confined radioactive material inserted into the body at or near a tumor or other proliferative tissue disease site. The term is intended without limitation to include exposure to radioactive isotopes (e.g., At-211, I-131, I-125, Y-90, Re-186, Re-188, Sm-153, Bi-212, P-32, and radioactive isotopes of Lu). Suitable radiation sources for use as a cell conditioner of the present disclosure include both solids and liquids. By way of non-limiting example, the radiation source can be a radionuclide, such as I-125, I-131, Yb-169, Ir-192 as a solid source, I-125 as a solid source, or other radionuclides that emit photons, beta particles, gamma radiation, or other therapeutic rays. The radioactive material can also be a fluid made from any solution of radionuclide(s), e.g., a solution of I-125 or I-131, or a radioactive fluid can be produced using a slurry of a suitable fluid containing small particles of solid radionuclides, such as Au-198, Y-90. Moreover, the radionuclide(s) can be embodied in a gel or radioactive micro spheres.
[0188] The antigen binding units or pharmaceutical compositions of the disclosure can be used in combination with an amount of one or more substances selected from anti-angiogenesis agents, signal transduction inhibitors, antiproliferative agents, glycolysis inhibitors, or autophagy inhibitors.
[0189] Anti-angiogenesis agents, such as MMP-2 (matrix-metalloproteinase 2) inhibitors, MMP-9 (matrix-metalloprotienase 9) inhibitors, and COX-11 (cyclooxygenase 11) inhibitors, can be used in conjunction with an antigen binding unit of the disclosure and pharmaceutical compositions described herein. Anti-angiogenesis agents include, for example, rapamycin, temsirolimus (CCI-779), everolimus (RAD001), sorafenib, sunitinib, and bevacizumab. Examples of useful COX-II inhibitors include CELEBREXTM (alecoxib), valdecoxib, and rofecoxib. Examples of useful matrix metalloproteinase inhibitors are described in WO 96/33172 (published Oct. 24, 1996), WO 96/27583 (published Mar. 7, 1996), European Patent Application No. 97304971.1 (filed Jul. 8, 1997), European Patent Application No. 99308617.2 (filed Oct. 29, 1999), WO 98/07697 (published Feb. 26, 1998), WO 98/03516 (published Jan. 29, 1998), WO 98/34918 (published Aug. 13, 1998), WO 98/34915 (published Aug. 13, 1998), WO 98/33768 (published Aug. 6, 1998), WO 98/30566 (published Jul. 16, 1998), European Patent Publication 606,046 (published Jul. 13, 1994), European Patent Publication 931, 788 (published Jul. 28, 1999), WO 90/05719 (published May 31, 1990), WO 99/52910 (published Oct. 21, 1999), WO 99/52889 (published Oct. 21, 1999), WO 99/29667 (published Jun. 17, 1999), PCT International Application No. PCT/IB98/01113 (filed Jul. 21, 1998), European Patent Application No. 99302232.1 (filed Mar. 25, 1999), Great Britain Patent Application No. 9912961.1 (filed Jun. 3, 1999), U.S. Provisional Application No. 60/148,464 (filed Aug. 12, 1999), U.S. Pat. No. 5,863, 949 (issued Jan. 26, 1999), U.S. Pat. No. 5,861,510 (issued Jan. 19, 1999), and European Patent Publication 780,386 (published Jun. 25, 1997), all of which are incorporated herein in their entireties by reference. Preferred MMP-2 and MMP-9 inhibitors are those that have little or no activity inhibiting MMP-1. More preferred, are those that selectively inhibit MMP-2 and/or AMP-9 relative to the other matrix-metalloproteinases (e.g., MAP-1, MMP-3, MMP-4, MMP-5, MMP-6, MMP- 7, MMP-8, MMP-10, MMP-11, MMP-12, andMMP-13). Some specific examples of MMP inhibitors useful in the disclosure are AG-3340, RO 32-3555, and RS 13-0830.
[0190] Autophagy inhibitors include, but are not limited to chloroquine, 3-methyladenine, hydroxychloroquine (Plaquenil.TM.), bafilomycin A1, 5-amino-4-imidazole carboxamide riboside (AICAR), okadaic acid, autophagy-suppressive algal toxins which inhibit protein phosphatases of type 2A or type 1, analogues of cAMP, and drugs which elevate cAMP levels such as adenosine, LY204002, N6-mercaptopurine riboside, and vinblastine. In addition, antisense or siRNA that inhibits expression of proteins including but not limited to ATG5 (which are implicated in autophagy), may also be used.
[0191] In some embodiments, the antigen binding units and pharmaceutical compositions described herein are formulated or administered in conjunction with liquid or solid tissue barriers also known as lubricants. Examples of tissue barriers include, but are not limited to, polysaccharides, polyglycans, seprafilm, interceed and hyaluronic acid.
[0192] In some embodiments, medicaments which are administered in conjunction with the subject antigen binding units include any suitable drugs usefully delivered by inhalation for example, analgesics, e.g., codeine, dihydromorphine, ergotamine, fentanyl or morphine; anginal preparations, e.g., diltiazem; antiallergics, e.g., cromoglycate, ketotifen or nedocromil; anti-infectives, e.g., cephalosporins, penicillins, streptomycin, sulphonamides, tetracyclines or pentamidine; antihistamines, e.g., methapyrilene; anti-inflammatories, e.g., beclomethasone, flunisolide, budesonide, tipredane, triamcinolone acetonide or fluticasone; antitussives, e.g., noscapine; bronchodilators, e.g., ephedrine, adrenaline, fenoterol, formoterol, isoprenaline, metaproterenol, phenylephrine, phenylpropanolamine, pirbuterol, reproterol, rimiterol, salbutamol, salmeterol, terbutalin, isoetharine, tulobuterol, orciprenaline or (-)-4-amino-3,5-dichloro-.alpha.-[[[6-[2-(2-pyridinyl)ethoxy]hexyl]-amino- ]methyl]benzenemethanol; diuretics, e.g., amiloride; anticholinergics e.g., ipratropium, atropine or oxitropium; hormones, e.g., cortisone, hydrocortisone or prednisolone; xanthines e.g., aminophylline, choline theophyllinate, lysine theophyllinate or theophylline; and therapeutic proteins and peptides, e.g., insulin or glucagon. It will be clear to a person skilled in the art that, where appropriate, the medicaments are used in the form of salts (e.g., as alkali metal or amine salts or as acid addition salts) or as esters (e.g., lower alkyl esters) or as solvates (e.g., hydrates) to optimize the activity and/or stability of the medicament.
[0193] Other exemplary therapeutic agents useful for a combination therapy include but are not limited to agents as described above, radiation therapy, hormone antagonists, hormones and their releasing factors, thyroid and antithyroid drugs, estrogens and progestins, androgens, adrenocorticotropic hormone; adrenocortical steroids and their synthetic analogs; inhibitors of the synthesis and actions of adrenocortical hormones, insulin, oral hypoglycemic agents, and the pharmacology of the endocrine pancreas, agents affecting calcification and bone turnover: calcium, phosphate, parathyroid hormone, vitamin D, calcitonin, vitamins such as water-soluble vitamins, vitamin B complex, ascorbic acid, fat-soluble vitamins, vitamins A, K, and E, growth factors, cytokines, chemokines, muscarinic receptor agonists and antagonists; anticholinesterase agents; agents acting at the neuromuscular junction and/or autonomic ganglia; catecholamines, sympathomimetic drugs, and adrenergic receptor agonists or antagonists; and 5-hydroxytryptamine (5-HT, serotonin) receptor agonists and antagonists.
[0194] Therapeutic agents can also include agents for pain and inflammation such as histamine and histamine antagonists, bradykinin and bradykinin antagonists, 5-hydroxytryptamine (serotonin), lipid substances that are generated by biotransformation of the products of the selective hydrolysis of membrane phospholipids, eicosanoids, prostaglandins, thromboxanes, leukotrienes, aspirin, nonsteroidal anti-inflammatory agents, analgesic-antipyretic agents, agents that inhibit the synthesis of prostaglandins and thromboxanes, selective inhibitors of the inducible cyclooxygenase, selective inhibitors of the inducible cyclooxygenase-2, autacoids, paracrine hormones, somatostatin, gastrin, cytokines that mediate interactions involved in humoral and cellular immune responses, lipid-derived autacoids, eicosanoids, .beta.-adrenergic agonists, ipratropium, glucocorticoids, methylxanthines, sodium channel blockers, opioid receptor agonists, calcium channel blockers, membrane stabilizers and leukotriene inhibitors.
[0195] Additional therapeutic agents contemplated herein include diuretics, vasopressin, agents affecting the renal conservation of water, rennin, angiotensin, agents useful in the treatment of myocardial ischemia, anti-hypertensive agents, angiotensin converting enzyme inhibitors, .beta.-adrenergic receptor antagonists, agents for the treatment of hypercholesterolemia, and agents for the treatment of dyslipidemia.
[0196] Other therapeutic agents contemplated include drugs used for control of gastric acidity, agents for the treatment of peptic ulcers, agents for the treatment of gastroesophageal reflux disease, prokinetic agents, antiemetics, agents used in irritable bowel syndrome, agents used for diarrhea, agents used for constipation, agents used for inflammatory bowel disease, agents used for biliary disease, agents used for pancreatic disease. Therapeutic agents used to treat protozoan infections, drugs used to treat Malaria, Amebiasis, Giardiasis, Trichomoniasis, Trypanosomiasis, and/or Leishmaniasis, and/or drugs used in the chemotherapy of helminthiasis. Other therapeutic agents include antimicrobial agents, sulfonamides, trimethoprim-sulfamethoxazole quinolones, and agents for urinary tract infections, penicillins, cephalosporins, and other, .beta.-lactam antibiotics, an agent comprising an aminoglycoside, protein synthesis inhibitors, drugs used in the chemotherapy of tuberculosis, mycobacterium avium complex disease, and leprosy, antifungal agents, antiviral agents including nonretroviral agents and antiretroviral agents.
[0197] Examples of therapeutic antibodies that can be combined with an antigen binding unit of the disclosure include but are not limited to anti-receptor tyrosine kinase antibodies (cetuximab, panitumumab, trastuzumab), anti CD20 antibodies (rituximab, tositumomab), and other antibodies such as alemtuzumab, bevacizumab, and gemtuzumab.
[0198] Moreover, therapeutic agents used for immunomodulation, such as immunomodulators, immunosuppressive agents, tolerogens, and immunostimulants are contemplated by the methods herein. In addition, therapeutic agents acting on the blood and the blood-forming organs, hematopoietic agents, growth factors, minerals, and vitamins, anticoagulant, thrombolytic, and antiplatelet drugs.
[0199] For treating renal carcinoma, one may combine an antigen binding unit of the present disclosure with sorafenib and/or Avastin. For treating an endometrial disorder, one may combine an antigen binding unit of the present disclosure with doxorubincin, taxotere (taxol), and/or cisplatin (carboplatin). For treating ovarian cancer, one may combine an antigen binding unit of the present disclosure with cisplatin (carboplatin), taxotere, doxorubincin, topotecan, and/or tamoxifen. For treating breast cancer, one may combine an antigen binding unit of the present disclosure with taxotere (taxol), gemcitabine (capecitabine), tamoxifen, letrozole, tarceva, lapatinib, PD0325901, avastin, herceptin, OSI-906, and/or OSI-930. For treating lung cancer, one may combine an antigen binding unit of the present disclosure with taxotere (taxol), gemcitabine, cisplatin, pemetrexed, Tarceva, PD0325901, and/or avastin.
[0200] Further therapeutic agents that can be combined with an antigen binding unit of the disclosure are found in Goodman and Gilman' s "The Pharmacological Basis of Therapeutics" Tenth Edition edited by Hardman, Limbird and Gilman or the Physician's Desk Reference, both of which are incorporated herein by reference in their entirety.
[0201] The antigen binding units described herein can be used in combination with the agents disclosed herein or other suitable agents, depending on the condition being treated. Hence, in some embodiments the one or more antigen binding units of the disclosure will be co-administered with other agents as described above. When used in combination therapy, the antigen binding units described herein are administered with the second agent simultaneously or separately. This administration in combination can include simultaneous administration of the two agents in the same dosage form, simultaneous administration in separate dosage forms, and separate administration. That is, an antigen binding unit described herein and any of the agents described above can be formulated together in the same dosage form and administered simultaneously. Alternatively, an antigen binding unit of the disclosure and any of the agents described above can be simultaneously administered, wherein both the agents are present in separate formulations. In another alternative, an antigen binding unit of the present disclosure can be administered just followed by and any of the agents described above, or vice versa. In some embodiments of the separate administration protocol, an antigen binding unit of the disclosure and any of the agents described above are administered a few minutes apart, or a few hours apart, or a few days apart.
[0202] Further illustration of the development and use of antigen binding units, polynucleotides, vectors and host cells according to this invention are provided in the Example section below. The examples are provided as a guide to a practitioner of ordinary skill in the art, and are not meant to be limiting in any way.
EXAMPLES
[0203] The following examples are given for the purpose of illustrating various embodiments of the invention and are not meant to limit the present invention in any fashion. The present examples, along with the methods described herein are presently representative of preferred embodiments, are exemplary, and are not intended as limitations on the scope of the invention. Changes therein and other uses which are encompassed within the spirit of the invention as defined by the scope of the claims will occur to those skilled in the art.
Example 1. Antigen Binding Unit Generation and Screening
[0204] Two different mouse strains (Balb/c and C57/BL6) were used for immunization to generate anti-CD47 monoclonal antibodies. Recombinant fragment of CD47, SEQ ID NO: 161, was expressed in 293F cells and used for immunization. Serum from serial or terminal blood samples was analyzed for the presence of specific antibodies. Serum titer data was used to select mice for hybridoma fusions.
[0205] Single cell suspensions were prepared from the best responder animals' spleens and electro-fused with myeloma cells before seeding and culture in 96 well plates. The hybridomas were then cultured in selection media for seven days prior to screening of supernatants.
[0206] Produced antigen binding units were characterized by a combination of binding of hybridoma supernatants to CD47 protein by ELISA, Raji cell expressed human CD47 using flow cytometry, and neutralizing ELISA. The antigen binding units were further characterization by their blocking efficacy in CD47-SIRP.alpha. interaction assays. A commercialized anti-CD47 mAb, referred to as "Positive 1", was used as a positive control and diluted pre-immune sera was used as negative reference.
[0207] Subcloning to obtain monoclonal hybridoma cell lines was performed.
Example 2. Macrophage Phagocytosis Assay
[0208] Select antibody hits were assessed in a macrophage phagocytosis assay to confirm the functional activity in vitro. Human monocyte derived macrophage (MDM) were co-cultured with target tumor cells HL-60 which were labelled with fluorescent dye CSFE. Phagocytosis was analyzed by Cellomics after two hours incubation. The percentage of macrophages that contain tumor cells were calculated and expressed as phagocytosis index. Results from one representative experiment are shown in FIG. 1. In FIG. 1, the antibodies are present at a concentration of 3 micrograms per mL; the white arrows point to examples of phagocytosed AML cells; human promyelocytic leukemia HL-60 cells are labeled in green; and human macrophages are labeled in red.
[0209] For this assay, human monocytes were purified from human PBMC with CD14 Miltenyi cell isolation beads. Purified CD14+ monocytes were cultured in T75 flask in the presence of M-CSF (100 ng/ml) for 7-10 days. Monocytes derived macrophages were harvested by incubation in dissociation buffer for 5 minutes, followed by gentle scraping. Then, M2 cells were labeled with PKH26 (red), 1.times.10.sup.4 macrophages were plated in a flat-bottom 96-well tissue culture plate in IMDM containing 10% FBS for 24 hours, media was replaced with serum-free media for another 2-hour incubation. 5.times.10.sup.4 CFSE-labeled HL-60 cells were added in to the well in the presence of indicated antibodies for 2 hours. Wells were washed 3 times with IMDM and cells were fixed with 2% PFA. Then the fluorescence-labeled cells were analyzed on Cellomics machine.
[0210] Phagocytosis index was determined by calculating the number of phagocytes per 100 macrophages, the data was calculated in Office Excel and graphed in Prism5, as shown in FIGS. 2A-2B. The EC50 was calculated based on percent of maximum of phagocytosis index, as shown in Table 4.
TABLE-US-00004 TABLE 4 ABU EC50 (nM) ABU1 +++ ABU2 +++ ABU3 ++ ABU4 +++ ABU5 ++ ABU6 +++ ABU7 + Positive 1 ++ Positive 2 ++ +++: <1 nM; ++: >1 nM and <5 nM; +: >5 nM
Example 3. CD47 Binding Assay
[0211] Binding affinity of antibodies for select antibodies (ABU-#) and chimeric antibodies (C-ABU-#) were measured by using both CD47 expressing CHO cells and Jurkat cells using either ELISA or flow cytometry. The protein binding kinetics of the antigen binding units were studied by SPR using Biacore T100 (GE Healthcare). Commercially available reference antibody was used as a control.
[0212] For ELISA-based binding analysis, human SIRPA (Novoprotein, Cat#C385) at a concentration of 1 .mu.g/mL was coated in the bottom of a 96-half-well plate overnight at 4.degree. C. Blocking was done with 3% skim milk in PBS (DOUBLE HELIX, Cat#P10033) for 1 hour at room temperature after 3 times wash by PBST (PBS with 0.05% tween-20(Sangon, Cat#9005-64-5)). Serial diluted human CD47 protein (Novoprotein, Cat#CG18) were added into the wells and incubate at room temperature for 1 hour; after PBST wash 3 times, the bound CD47 protein were detected by HRP-conjugated goat polyclonal secondary antibody to human IgG-Fc (Abcam, Cat#ab98624) and then developed by TMB substrate (Biopanda, Cat#TMB-S-003) after 3 times PBST wash, followed by measurement of OD450. The binding curve (OD450 versus CD47 Concentration) were analyzed by GraphPad Prism and EC90 calculated for CD47 neutralizing study. Binding results are depicted below in Table 5.
TABLE-US-00005 TABLE 5 Binding on Binding on SIRP.alpha. CHO-CD47 Jurkat blocking Affinity Cells Cells on cells by SPR Clone ID (EC50, nM) (EC50, nM) (IC50, nM) (nM) ABU 1 +++ +++ ++ - ABU 2 +++ +++ ++ + ABU 3 ++++ ++++ ++ + ABU 4 ++++ +++ +++ + ABU 5 +++ +++ ++ + ABU 6 ++++ ++++ +++ + C-ABU 1 ++++ ++ ++ ND positive 1 +++ +++ ++ + positive 2 ++++ ++ ++ + ++++: <0.05 nM; +++: >0.05 nM and <1 nM; ++: >1 nM and <10 nM +: >10 nM and <40 nM; -: >40 nM; ND: Note determined
[0213] Antibody cross-reactivity was studied by flow cytometry using cells expressing CD47 from different species. Cynomolgus monkey red blood cells was used for monkey cross reactivity. CHO cell expressing mouse CD47 was used for mouse cross reactivity study. For flow cytometry-based binding analysis, human SIRPa expressing CHO cell line were generated and SIRPa expression were evaluated by binding of CD47-Fc protein (Novoprotein, Cat#CG18) or biotin labelled polyhistidine tagged CD47 protein (Novoprotein, Cat#321). Briefly, Human SIRP.alpha. expressing CHO cells were dissociated by Trypsin-EDTA dissociation buffer and washed 3 times with FACS buffer (PBS buffer with 2% FBS). The cells were plated into 96 wells plate and spin the cells done. Serially diluted CD47 protein (Fc tagged protein for murine antibody and biotin labelled polyhistindine tagged CD47 protein for chimeric/humanized antibody) were used to re-suspend the cells and incubate at 4.degree. C. for one hours and then stained with APC labelled anti-human Fc antibody after washing out those non-bound proteins. After one hour incubation at 4 degree and 3 times wash, the cells were analyzed on Guava HL6T machine and data were analyzed using FlowJo software. at a concentration of 1 .mu.g/mL was coated in the bottom of a 96-half-well plate overnight at 4.degree. C. The binding curve (mean fluorescence intensity verse CD47 Concentration) were analyzed by GraphPad Prism and EC90 calculated for CD47 neutralizing study. Data from an example experiment is depicted in FIGS. 3A and 3B.
Example 4. Neutralizing ELISA
[0214] The neutralizing effect of select antibodies was analyzed by ELISA using SIRP.alpha. coated plates. Briefly, the SIRP.alpha. protein was coated on microtiter plates and CD47-hFc protein was added in serial dilution to establish a corresponding EC90. Select antibodies were serial diluted and mixed with hCD47-hFc fusion protein at its EC90 concentration, and their dose dependent blocking effect was detected with HRP labelled anti-hIgG antibodies against CD47-hFc.
[0215] Briefly, human SIRPA (Novoprotein, Cat#C385) at a concentration of 1 .mu.g/mL was coated in the bottom of a 96-half-well plate overnight at 4.degree. C. Blocking was done with 3% skim milk in PBS (DOUBLE HELIX, Cat#P10033) for 1 hour at room temperature. At the same time, pre-incubate 12.5 .mu.L human CD47 (Novoprotein, Cat#CG18) at 2.5 .mu.g/mL with hybridoma supernatant or anti-CD47 antibody gradients ("Positive 1" (eBioscience, Cat#14-0479) as positive control and "Negative 1" (eBioscience, Cat#14-0478) as negative control) for 1 hour at room temperature, which were then applied to each blocked well for 1 hour at room temperature. The blocking effect were detected by HRP-conjugated goat polyclonal secondary antibody to human IgG-Fc (Abcam, Cat#ab98624) and then developed by TMB substrate (Biopanda, Cat#TMB-S-003) after 3 times PBST wash, followed by measurement of OD450. The neutralizing activity (OD450 verse antibody Concentration) were analyzed by GraphPad Prism and IC50 calculated for evaluation.
[0216] The neutralizing effect of select antibodies was also analyzed by flow cytometry. Human SIRPa expressing CHO cell line were generated and SIRPa expression were evaluated by CD47 protein binding. Briefly, Human SIRPa expressing CHO cells were dissociated by Trypsin-EDTA dissociation buffer and washed 3 times with FACS buffer (PBS buffer with 2% FBS). The cells were plated into 96 wells plate and spin the cells done. Pre-incubate 25 .mu.L human CD47-Fc at 2.5 .mu.g/mL or biotinylated CD47 protein with hybridoma supernatant or anti-CD47 antibody gradients (B6H12 (eBioscience, Cat#14-0479) as positive control and 2D3 (eBioscience, Cat#14-0478) as negative control) for 1 hour at room temperature, and then re-suspend the cell pellet and incubate one hour at 4 degree. then stained with APC labelled anti-human Fc antibody or APC-Streptavidin after washing out those non-bound proteins. After one hour incubation at 4 degree and 3 times wash, the cells were analyzed on Guava HL6T machine and data were analyzed using Flowjo software. The neutralizing activity (mean fluorescence intensity verse antibody Concentration) were analyzed by GraphPad Prism and IC50 calculated for evaluation. Results from an example neutralizing ELISA experiment are summarized in Table 6.
TABLE-US-00006 TABLE 6 Antibody ABU 2 ABU 3 ABU 4 ABU 5 Abu 6 Pos. 1 IC50 (nM) ++++ ++ +++ ++++ + ++++ ++++: <8 nM; +++: >8 nM and <15 nM; ++: >15 nM and <20 nM; +: >20 nM
Example 5. Anti-CD47 Antibody-Dependent Phagocytosis
[0217] The dose dependent effect of select antibodies on in vitro phagocytosis were performed to obtain EC50.
[0218] Human monocytes were purified from human PBMC with CD14 Miltenyi cell isolation beads. Purified CD14+ monocytes were cultured in T75 flask in the presence of M-CSF (100 ng/ml) for 7-10 days. Monocytes derived macrophages were harvested by incubation in dissociation buffer for 5 minutes, followed by gentle scraping. Then, M2 cells were labeled with PKH26 (red), 1.times.10.sup.4 macrophages were plated in a flat-bottom 96-well tissue culture plate in IMDM containing 10% FBS for 24 hours, media was replaced with serum-free media for another 2-hour incubation. 5.times.10.sup.4 CFSE-labeled HL-60 cells were added in to the well in the presence of indicated antibodies for 2 hours. Wells were washed 3 times with IMDM and cells were fixed with 2% PFA. Then the fluorescence-labeled cells were analyzed on Cellomics machine.
[0219] The phagocytosis index were determined by calculating the number of phagocytes per 100 macrophages, the data was calculated in Office Excel and graphed in Prism5. The EC50 was calculated based on percent of maximum of phagocytosis index.
[0220] Data from example phagocytosis experiments using HL60 cell line cells are depicted in FIG. 4. The EC50 were calculated and summarized in Table 7. Data from example phagocytosis experiments using DLD-1 human colon cancer cell line are depicted in FIG. 5.
TABLE-US-00007 TABLE 7 ABU ABU ABU ABU ABU ABU Pos. Pos. Ab ID 1 2 3 4 5 6 1 2 IC50 (nM) +++ ++ + ++ + ++ + + +++: <0.6 nM; ++: >0.6 nM and <1 nM; +: >1 nM
Example 6. Epitope Binning of CD47 Hit Antibodies
[0221] Epitope binning of these hits were conducted using CD47-experssing CHO cells with a commercial anti-CD47 blocking antibody Positive 1 comprising SEQ ID NO: 240 and SEQ ID NO: 241, a benchmark analogue antibody Positive 2 comprising SEQ ID NO: 242 and SEQ ID NO: 243, as well as an analogue antibody Positive 3 comprising SEQ ID NO: 244 and SEQ ID NO: 245. Positive 1 also comprised SEQ ID NO: 149-154. Positive 2 also comprised SEQ ID NO: 155-160. Briefly, seven neutralizing CD47 antibodies and 3 reference antibodies were analyzed and grouped according to their competitive binding to CHO cell expressing CD47 using flow cytometry. A biotinylated antibody was first used to calculate the concentration for 90% binding, then the CD47 antibodies were serially diluted and mixed with one of the biotinylated antibodies at the pre-determined 90% binding concentration. SA-APC was used to detect the binding of biotinylated antibody binding. As the experiment designed, the binding of labeled antibodies should be affected by the Abs of the same bin and classified as one group. If the binding of labeled antibodies is not affected, then those testing antibodies are not in the same bin and characterized as a separate group. Three binding profiles on the CHO expressing CD47 were identified and the six hits subjected to two groups shown below in Table 8. ABU4 and ABUS belong to one epitope binding group while the other, including positive 3, belong to a different one. Positive 1 interfered the two groups binding on the cells.
TABLE-US-00008 TABLE 8 ABU Epitope group ABU4 A ABU5 A ABU1 B ABU2 B ABU3 B ABU6 B Positive 2 B Positive 3 B Positive 1 A/B
Example 7. Red Blood Cell Binding and Hemagglutination Assay
[0222] CD47 is ubiquitously expressed with particular high expression on red blood cells. To evaluate the binding affinity on human red blood cells, the RBC binding assay were performed using RBCs from several donors. The binding curve and EC50 from an example experiment is depicted in FIG. 6.
[0223] Select antibodies were also tested in a hemagglutination assay to identify antibodies showing strong hemagglutination effect. The antibodies to be tested were diluted as indicated concentration in PBS and 90 ul of antibodies with series dilution were added in the V-bottom culture plate for incubation of 37.degree. C. for 1 hour. Then 10 uL of human red blood cells (RBCs) were added with 10% final concentration in PBS. The RBCs were incubated with antibodies in 37.degree. C. and the hemagglutinin will be observed in 2-4 hours. Evidence of hemagglutinin is demonstrated by the presence of non-settlement RBCs, appearing as haze compared to punctuate red dots of non-hemagglutinated RBCs.
[0224] Hemagglutination index was quantified by the area of RBC pellets in presence or absence of mAbs, the diameter of RBC pellets were determined by ImageJ software in pixel, then the area was calculated in excel. The calculated data was normalized to the isotype IgG. The log concentration v.s index was plotted in Prism 5 and is depicted in FIGS. 7A-7B.
[0225] Images from an example experiment is depicted in FIG. 8. Induction of significant hemagglutination of human red blood cell would be evidenced by a haze appearance in the wells. In FIG. 8, Isotype muIgG1 serves as a negative control; the dotted line marks blank controls; a positive hemagglutination control is circles in blue; and a negative hemagglutination control is circles in yellow.
Example 8. Characterization of Chimeric Anti-CD47 Antibodies
[0226] To generate chimeric antibodies, selected antigen binding units were picked and expanded and the V-region sequences were recovered. The V regions were synthesized and subcloned in vectors to obtain human IgG4 (S228P) chimera. S228P refers to the amino acid position where IgG4 is split as described in Aalberse RC and Schuurman J (2002) IgG4 breaking the rules. Immunology 105:9-19 (incorporated herein in its entirety). The human IgG4 sequence was cloned onto the C-terminus of the light chain variable region sequence and heavy chain variable region sequence of the selected antibodies. The chimeric hIgG4 antibodies were expressed in 293F cells and purified as human IgG4 format for functional validation. Chimeric antibodies were also generated with IgG1 constant region in place of the IgG4 region.
[0227] Binding affinities of the chimeric antibodies were measured by flow cytometry with CD47-expressing cell lines. Results from an example experiment are depicted in FIG. 9.
[0228] The neutralizing effect of select chimeric antibodies was analyzed using cells expressing CD47. Briefly, human chimeric IgG4 antibody was diluted and mixed with biotinylated human CD47 protein. The dose-dependent SIRPcL binding-blocking effects was then detected for C-ABU1, C-ABU2, and C-ABU 4. Results from example experiments are depicted in FIG. 10.
[0229] Select chimeric antibodies were further characterized in a DLD-1 cell phagocytosis assay performed as described above. Results from example experiments are depicted in FIG. 11, and the asterisks denote statistical significance based on a p value calculated by one-way ANOVA post Dunnett's test.
Example 9. Phage Selection of Additional Anti-CD47 Fabs
[0230] Anti-CD47 Fabs were generated using the following phage-based method. Total RNA from spleens of immunized mice was prepared. After oligo(dT) primed reverse transcription, the antibody variable regions VL and VH were amplified by PCR. The murine VL and VH regions were then fused to human constant regions CL and CH1 of light chain and heavy chain respectively. The combination of the chimeric light chains and heavy chain fragments was cloned into the phagemid vector pComb3X and resulted in a murine/human Fab library displayed on phage.
[0231] The generated phage library was then screened for anti-CD47 specific Fabs. Dynabeads conjugated with Streptavidin were first blocked by incubating for 1 hour at room temperature with PBS containing 3% BSA. Approximately 1 .mu.L blocked Dynabeads were incubated for 30 minutes with decreasing amount of biotinylated CD47-Fc (100 nM, 50 nM and 25 nM for rounds 1, 2 and 3, respectively) to capture antigen. Phage libraries were pre-adsorbed on another approximately 1 .mu.L blocked Dynabeads in PBS containing 3% BSA for 30 minutes, followed by depletion with approximately 1 .mu.g/mL human Fc fragment in PBS containing 3% BSA. Depleted phage library then were mixed with antigen coated Dynabeads for 1 hour at room temperature with gentle rotating head-over-head. The Dynabeads were then washed with 1 mL PBS containing 0.05% Tween-20 (5, 10 and 15 times for rounds 1, 2 and 3, respectively) using magnetic separator. Bound phage were eluted by incubation at room temperature for 10 minutes with 500 .mu.L elution buffer followed by neutralization with 50 .mu.L neutralization buffer. Eluted phage were rescued by infection of E. coli TG1 cells and a phage library was prepared for the next round of screening.
[0232] Select Fabs were characterized by various ELISA assays. Soluble Fab fragments were induced and periplasmic fractions were prepared by the following method. Individual clones from third round of screening were picked from an agar plate and cultured overnight in a microtiter plate containing 2YT. 5 .mu.L of each overnight culture was transferred to a microtiter plate containing 150 .mu.L of 2YT, 2% glucose, 50 .mu.g/mL carbenicillin, and grown at 37.degree. C. for 3 hours. Isoproplythiogalactoside (IPTG) was added to each well to a final concentration of 1mM. After overnight grown with shaking at 25.degree. C., plates were spun down, and supernatant was used directly in binding ELISA. For preparation of periplasmic fractions, cell pellets were resuspended and incubate on ice for 20 minutes. These periplasmic fractions, were then used for testing specificity by flow cytometry and blocking ELISA assays.
[0233] Large scale induction of soluble Fab fragments from individual clones was performed on a 50 ml scale in 2YT containing 50 .mu.g/ml carbenicillin and 2% glucose. After growth at 37.degree. C. to an OD600 of 0.9, IPTG was added to the final concentration of 1 mM. After growing overnight at 25.degree. C., cell pellets were harvested and periplasmic fractions were prepared as described above.
[0234] The pooled large scale of periplasmic fraction descried above were passed over a 1 mL Ni-- resin according to the manufacturer's instructions. The column was washed with buffer and the protein was eluted by applying buffer. The eluted protein was filtered to change the buffer to PBS. Purified Fabs then were analyzed by SDS-PAGE under non-reducing as well as reducing conditions and the concentration determined spectrophotometrically.
[0235] Select Fabs were screened by ELISA. Microtiter plate was coated with human CD47-Fc at a concentration of 1 .mu.g/ml in PBS overnight at 4 degrees Celsius, washed three times with PBS/0.05% Tween 20, blocked with PBS/3% skim milk for 1 hour at room temperature, and then incubated with 50 .mu.L of supernatant from individual clone for 1 hour at room temperature. After 3 washes with PBS/0.05% Tween 20, 50 .mu.L of a 1:5000 dilution of HRP conjugated anti-human IgG F(ab)2 specific was added and incubate for 1 hour at room temperature. Following 3 washes with PBS/0.05% Tween 20, 50 .mu.L of TMB substrate was added for color development. Reactions were stopped by addition of 50 .mu.L of HCl, and OD450 was measured in a microtiter plate reader.
[0236] Activity of blocking CD47's interaction with SIRPa of each clone was carried out by inhibition ELISAs. Microtiter plate was coated with Human SIRPa at a concentration of 1 .mu.g/ml in PBS overnight at 4.degree. C., washed three times with PBS/0.05% Tween 20, blocked with PBS/3% skim milk for 1 hour at room temperature. 50 .mu.L of periplasmic fraction of each clone was mixed with approximately 1 .mu.L of human CD47 and incubated for 1 hour at room temperature, followed by adding 50 .mu.L of the mixture into blocked wells of microtiter plate. Incubated for 1 hour at room temperature and the following steps were exactly same as above.
[0237] CHO cells that had been stably transfected with human SIRPa were detached using cell dissociation buffer. Aliquots of 200 .mu.L containing 10{circumflex over ( )}5 cells were distributed into wells of U-bottom 96-well plate. After washing three times with FACS buffer (2% FBS in PBS), 0.07 .mu.g/mL of biotinylated CD47 was added in the presence of serial diluted concentrations of Fab in FACS buffer to resuspend cells, and incubated at 4.degree. C. for 30 minutes. Then, the cells were washed with FACS buffer three times, followed by incubation with 1:1000 diluted APC-labeled SA (Invitrogen) at 4.degree. C. for 30 minutes. After three times of washing, binding was measured on Guava HL6T machine. Data from an example experiments are shown in FIG. 12A and Table 9.
[0238] CHO cells that had been stably transfected with human SIRP.alpha. were detached using cell dissociation buffer. Aliquots of 200 .mu.L containing 10{circumflex over ( )}4 cells were distributed into wells of U-bottom 96-well plate. After washing three times with FACS buffer (2% FBS in PBS), 0.07 .mu.g/mL of His-tagged CD47 was added in the presence of serial diluted concentrations of Fab in FACS buffer to resuspend cells, and incubated at 4.degree. C. for 30 minutes. Then, the cells were washed with FACS buffer three times, followed by incubation with 1:1000 diluted anti-His-APC (GenScript #A01802) at 4.degree. C. for 30 minutes. After three times of washing, binding was measured on Guava HL6T machine. Data from an example experiment are shown in FIGS. 12B-12C and Table 10.
TABLE-US-00009 TABLE 9 ABU8 ABU9 ABU11 ABU14 ABU16 ABU40 ABU20 ABU24 C-ABU1 IC50(nM) ++ +++ ++ ++ + ++ ++ + + +++: <10 nM; ++: >10 nM and <65 nM; +: >65 nM
TABLE-US-00010 TABLE 10 ABU9 ABU14 ABU40 Positive 3 IC50(nM) ++ ++ + + Frequency IC50(nM) +++ +++ +++ + MFI +++: <2 nM; ++: >2 nM and <2.7 nM; +: >2.7 nM
[0239] Select Fabs were also characterized by cell-binding assays. Aliquots of 200 .mu.L containing 10{circumflex over ( )}5 HL60 cells or DLD1 cells were distributed into wells of U-bottom 96-well plate. After washing three times with FACS buffer (2% FBS in PBS), cells were resuspended in serial diluted concentrations of Fab or IgG in FACS buffer and incubated at 4.degree. C. for 30 minutes. Then, the cells were washed with FACS buffer three times, followed by incubation with approximately three fold diluted APC-labeled anti-human (Fab)2 specific antibody or APC-labeled goat anti-human IgG, respectively, at 4.degree. C. for 30 minutes. After three times of washing, binding was measured on Guava HL6T machine. Data from an example experiment using HL60 cells and Fabs are shown in FIG. 13A and Table 11. Data from an example experiment using DLD1 cells and Fabs are shown in FIG. 13B and Table 12. Additional experiments using DLD1 cells and IgGs are shown in FIGS. 13C and 13D and Table 13.
TABLE-US-00011 TABLE 11 ABU8 ABU9 ABU11 ABU14 ABU16 ABU40 ABU20 ABU24 C-ABU1 EC50(nM) +++ ++ +++ + +++ +++ +++ +++ +++ +++: <1 nM; ++: >1 nM and <7 nM; +: >7 nM
TABLE-US-00012 TABLE 12 ABU8 ABU9 ABU11 ABU14 ABU16 ABU40 ABU20 ABU24 C-ABU1 EC50(nM) ++ +++ + +++ +++ ++ + ++ ++ +++: <5 nM; ++: >5 nM and <60 nM; +: >60 nM
TABLE-US-00013 TABLE 13 Posi- Posi- ABU9 ABU14 ABU16 ABU40 ABU24 tive 3 tive 2 IC50(nM) +++ +++ +++ ++ ++ +++ ++ Frequency IC50(nM) +++ +++ ++ + +++ +++ ++ MFI +++: <2 nM; ++: >2 nM and <20 nM; +: >20 nM
[0240] Chimeric versions of select Fabs were generated and subjected to an ELISA assay. FcRn blocked DLD1 cells expressing CD47 were aliquoted so approximately 10{circumflex over ( )}4 cells were distributed into wells of U-bottom 96-well plate. After washing three times with FACS buffer (2% FBS in PBS), 100 nM of biotinylated SIRPa-Fc was added in the presence of serial diluted concentrations of Fab in FACS buffer to resuspend cells, and incubated at 4.degree. C. for 30 minutes. Then, the cells were washed with FACS buffer three times, followed by incubation with diluted APC-labeled SA (Invitrogen) at 4.degree. C. for 30 minutes. After three times of washing, binding was measured on Guava HL6T machine. Data from an example experiment are shown in FIGS. 14A and 14B and Table 14.
TABLE-US-00014 TABLE 14 ABU9 ABU14 ABU40 ABU24 Positive 3 IC50(nM) +++ +++ +++ +++ ++ Frequency IC50(nM) +++ +++ + + +++ MFI +++: <1 nM; ++: >1 nM and <20 nM; +: >20 nM
Example 10. Humanization of Anti-CD47 Antibodies
[0241] Antibody sequences were subjected to profiling using structure modeling, to identify best matched germline and back mutation sites. The optimized mutants were synthesized and recombinant antibodies were produced for binding affinity determined by ELISA. After grafting and back mutation, the affinity of the humanized antibodies was either retained or improved (FIG. 15A), H-ABU2 was further affinity-matured by FACS screening of mutagenesis library to select clones that could bind the antigen better, in some cases up to 10 fold, compared with the corresponding chimeric parental antibody.
[0242] Select affinity matured humanized antibodies were further characterized in a blocking experiment as described previously. Data from example experiments using Raji cells are depicted in FIGS. 15B-15C and Table 15.
TABLE-US-00015 TABLE 15 H- H- H- H- H- H- Positive Chimeric ABU2 ABU3 ABU4 ABU5 ABU6 ABU7 3 IC50 +++ + + +++ ++ +++ +++ +++ H- H- H- H- Positive Chimeric ABU8 ABU9 ABU10 ABU11 3 IC50 +++ +++ ++ ++ +++ +++ +++: <1 nM; ++: >1 nM and <20 nM; +: >20 nM
[0243] Another set of humanized antibodies were analyzed in a binding assay as described previously. The binding assays performed were ELISA binding EC50 studies similar to those described in Example 3 using CD47 coated plates. Data from an example experiment are depicted in FIG. 16A and Table 16. A group of these antibodies were further characterized in a SIRP-alpha blocking experiment using Raji cells as as described previously. Blocking assays were performed similarly to those described in Example 4. Data from an example experiment is depicted in FIG. 16B and Table 17.
TABLE-US-00016 TABLE 16 C- H- H- H- H- H- H- H- H- ABU ABU12 ABU13 ABU14 ABU15 ABU16 ABU17 ABU18 ABU19 EC50 + + + + + + + + + H- H- H- H- H- H- H- H- H- ABU20 ABU21 ABU22 ABU23 ABU24 ABU25 ABU26 ABU27 ABU28 EC50 + + + + + + + + + H- H- H- H- H- H- H- H- ABU29 ABU30 ABU31 ABU32 ABU33 ABU34 ABU35 ABU36 EC50 + + + + + + + + +: <0.06 .mu.g/mL
TABLE-US-00017 TABLE 17 C- H- H- H- H- H- H- H- Positive ABU ABU37 ABU38 ABU39 ABU40 ABU41 ABU42 ABU43 3 IC50 + + + + + + N.A. + + (nM) +: <0.6 nM
[0244] Select murine, chimeric, and humanized antibodies were subjected to a hemagglutination assay as described previously. Briefly, monoclonal antibodies were serially diluted and incubated for 1 hour before adding whole blood for a final blood concentration of 10%. After 2-4 hours of incubation, hemagglutination effect was examined by scanner.
[0245] Select murine, chimeric, and humanized antibodies were also subjected to a red blood cell and platelet binding assay. Among the humanized antibodies were H-ABU 2-G1, which comprises a human IgG1 constant region, and H-ABU 2-G4, which comprises a human IgG4 constant region. Blood was diluted 1:100 with DPBS. Monoclonal antibodies starting at 10 .mu.g/mL were serially diluted and added to the diluted blood with a volume ration of 1:2 (20 .mu.l of antibodies and 40 .mu.l of diluted blood). The mixture was incubated for 30 minutes at 40 degrees Celsius and then washed twice with DPBS. Secondary antibodies were then added, namely APC-anti-human or APB-anti-mouse monoclonal antibodies (Jackson Immuno Research 315-606-046 or 109-605-088 respectively), and FITC labeled anti-human CD61 (BD, 555753) for platelet binding. After the secondary antibodies were added, the mixture was incubated for 30 minutes at 4 degrees Celsius. Cells were washed twice with DPBS and binding affinity was assessed using a flow cytometer.
[0246] Select humanized antibodies were analyzed using a hemagglutination assay as described previously. Data from an example experiment using samples from two different donors and the indicated antibodies or controls at the listed concentration are depicted in FIG. 17. Hemagglutination assays were performed similarly to those described in Example 7. As described above, a distinct puncta of blood cells that lack of a hazy halo appearance indicates that no significant hemagglutination has occurred. Many of the humanized antibodies showed no significant hemagglutination effect.
[0247] Select antibodies were next tested using a binding assay using cynomolgus monkey red blood cells expressing CD47 to test for monkey cross-reactivity. The assay was performed similarly to that described in Example 3. Data from an example experiment are depicted in FIG. 18. In this example, antibodies with two variations of IgG were tested. Namely, the IgG4 used were one of two different Fc gamma receptor variants: either IgG4 with a S228P mutation (SEQ ID NO: 37), or an IgG with both a S228P and L235E mutations (SEQ ID NO:18). The Positive-3-IgG4-PE antibody comprises SEQ ID NO: 305 and SEQ ID NO: 306. The Positive-3-IgG4-P antibody comprises SEQ ID NO: 307 and SEQ ID NO: 308.
[0248] Select antibodies were tested using a phagocytosis assay as described previously. These assays were performed similarly to those described in Example 5. Either DLD-1 cells, CCRF-CEM cells, or Raji cells were used in these experiments. Data from example experiments are depicted in FIGS. 19A-19C.
[0249] Select antibodies were further tested using a binding assay as described previously. Antibodies were tested for their binding level to either RBC (FIGS. 20A and 20B) or platelets (FIGS. 20C or 20D). In this example, antibodies with two variations of IgG were tested. Namely, the IgG4 used were one of two different Fc gamma receptor variants: either IgG4 with a S228P mutation (SEQ ID NO: 37), or an IgG with both a S228P and L235E mutations (SEQ ID NO:18).
Example 11. In Vivo CDX Model and In Vivo Anti-Tumor Activity of Humanized Antibodies
[0250] Raji cells were maintained in vitro as a culture in RPMI-1640 medium supplemented with 10% heat inactivated fetal bovine serum, 100 U/ml penicillin and 100 .mu.g/ml streptomycin at 37.degree. C. in an atmosphere of 5% CO2 in air. The tumor cells were routinely sub-cultured twice weekly. The cells growing in an exponential growth phase were harvested and counted for tumor inoculation. Each mouse was inoculated subcutaneously at the right flank with Raji tumor cells (3.times.10.sup.6) in 0.1 ml of PBS for tumor development. The treatments were started on day 8 after tumor inoculation when the average tumor size reached approximately 113 mm.sup.3. Each group consisted of 7 or 8 tumor-bearing mice.
[0251] At the time of routine monitoring, the animals were daily checked for any effects of tumor growth and treatments on normal behavior such as mobility, food and water consumption (by looking only), body weight gain/loss (body weights were measured twice weekly), eye/hair matting and any other abnormal effect as stated in the protocol. Death and observed clinical signs were recorded on the basis of the numbers of animals within each subset.
[0252] Tumor size was measured twice weekly in two dimensions using a caliper, and the volume was expressed in mm.sup.3 using the formula: V=0.5 a.times.b.sup.2 where a and b are the long and short diameters of the tumor, respectively. The tumor size was then used for calculation of T/C value. The T/C value (in percent) is an indication of antitumor effectiveness; T and C are the mean volumes of the treated and control groups, respectively, on a given day.
[0253] A one-way ANOVA was performed to compare tumor volume among groups, and when a significant F-statistics (a ratio of treatment variance to the error variance) was obtained, comparisons between groups were carried out with Games-Howell test. All data were analyzed using Graphpad5.0. P<0.05 was considered to be statistically significant.
[0254] Select humanized antibodies were tested in vivo as described above to determine their effect on tumor size. Briefly, approximately 3 million Raji cells were subcutaneously transplanted into NOD/SCID mice. The dosage of anti-CD47 antibodies were 10 mpk, i.p., 3 times per week. The dosing of antibody started when the tumor volume reached 100 mm.sup.3. Data for an example experiments is depicted in FIG. 21.
Example 12: Comparison of IgG4P and IgG4PE Form
[0255] Select humanized antibodies were further characterized by testing variants comprising either of two variations of IgG. Namely, the IgG4 used were one of two different Fc gamma receptor variants: either IgG4 with a S228P mutation (SEQ ID NO: 37, P variant), or an IgG with both a S228P and L235E mutations (SEQ ID NO:18, PE variant). FIG. 22A depicts data from an example blocking assay using Raji cells which was generally performed as described previously. FIGS. 22B and 23C depict data from an example RBC and platelet binding assays respective, which were generally performed as previously described. FIG. 22D depicts data from an example hemagglutination assay, which was generally performed as described previously. As described above, a distinct puncta of blood cells that lack of a hazy halo appearance indicates that no significant hemagglutination has occurred.
Example 13. Characterization and Anti-Tumor Activity of Humanized Antibodies and Combination Therapy
[0256] Select humanized antibodies were tested in vivo as described above to determine their affect on tumor size. Data form example experiments is depicted in FIG. 23. Xenograft experiments were performed generally as described in Example 10. Briefly, approximately 3 million Raji cells were subcutaneously transplanted into NOD/SCID mice. The dosage of anti-CD47 antibodies were 10 mpk, i.p., 3 times per week. The assay was started when the tumor volume reached 100 mm.sup.3. In this example, antibodies with two variations anti-CD47 antibodies with different IgG were tested. Namely, the IgG4 used were one of two different Fc gamma receptor variants: either IgG4 with a S228P mutation (SEQ ID NO: 37), or an IgG with both a S228P and L235E mutations (SEQ ID NO:18).
[0257] Select antibodies were used in xenografting experiments in combination with anti-CD20 antibodies to determine their effect on tumor size as described previously above. Three million of Raji cells were subcutaneously transplanted into the NOD/SCID mice. When the tumor volume reached 100 mm.sup.3, C-ABU 1 antibody, anti-CD20 or a combination of both were injected by intraperitoneal at 10 mg/kg, every other day and the tumor sizes were recorded. Combination with C-ABU 1 and anti-CD20 significantly prevented the progress of the tumor. Data from example experiments is summarized in FIG. 24. Each antibody or control was administered as summarized in Table 18. The dosage of anti-CD20 and anti-CD47 antibodies were 10 mpk, i.p., 3 times per week. The assay was started when the tumor volume reached 100 mm.sup.3. In this example, antibodies with two variations anti-CD47 antibodies with different IgG were tested. Namely, the IgG4 used were one of two different Fc gamma receptor variants: either IgG4 with a S228P mutation (SEQ ID NO: 37), or an IgG with both a S228P and L235E mutations (SEQ ID NO:18). Anti-CD20 used in this assay was in a human IgG1 format.
TABLE-US-00018 TABLE 18 Dose Dosing Group n.sup.a Treatment (mg/kg) Route Schedule.sup.c 1 8 Isotype Control 10 i.p. TIW X 3 wks 2 8 Positive IgG4-PE 10 i.p. TIW X 3 wks 3 8 H-ABU41-IgG4-PE 10 i.p. TIW X 3 wks 4 8 Isotype Control + 10 i.p. TIW X 3 wks anti-CD20 5 8 Positive IgG4-PE + 10 i.p. TIW X 3 wks anti-CD20 6 8 H-ABU41-IgG4-PE + 10 i.p. TIW X 3 wks anti-CD20
[0258] While preferred embodiments of the present invention have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the invention. It should be understood that various alternatives to the embodiments of the invention described herein may be employed in practicing the invention. It is intended that the following claims define the scope of the invention and that methods and structures within the scope of these claims and their equivalents be covered thereby.
TABLE-US-00019 SEQUENCE LISTING SEQ ID NO: 1 KSSQSLLYSSNQKNYLA SEQ ID NO: 2 RASKNIGKYLA SEQ ID NO: 3 KASQDIKSYLS SEQ ID NO: 4 SASSSVSYMN SEQ ID NO: 5 RSSQSIVYSNGNTYLE SEQ ID NO: 6 KASENVGTYVS SEQ ID NO: 7 RSSQSIVHSNGNTYLE SEQ ID NO: 8 DTSKLAS SEQ ID NO: 9 GASNRYT SEQ ID NO: 10 KVSNRFS SEQ ID NO: 11 RVANRFS SEQ ID NO: 12 SGSTLQS SEQ ID NO: 13 WASTRDS SEQ ID NO: 14 YATSLAD SEQ ID NO: 15 FQGSHVPWT SEQ ID NO: 16 FQGSHVPYT SEQ ID NO: 17 GQSYSYPLT SEQ ID NO: 18 ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSS GLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPEFEGGPSV FLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFNST YRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLPPSQEEM TKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSRLTVDKSRW QEGNVFSCSVMHEALHNHYTQKSLSLSLGK SEQ ID NO: 19 LQHGESPFT SEQ ID NO: 20 QQHNEYPYT SEQ ID NO: 21 QQWNSNPPT SEQ ID NO: 22 QQYYSYPLT SEQ ID NO: 23 DFYIN SEQ ID NO: 24 DTYMH SEQ ID NO: 25 DYGMA SEQ ID NO: 26 GYYMN SEQ ID NO: 27 NYWIA SEQ ID NO: 28 NYWMN SEQ ID NO: 29 NYWMQ SEQ ID NO: 30 DFYPGNTSTNYNEKFKT SEQ ID NO: 31 EINPSTGGTTYNQKFKA SEQ ID NO: 32 FISNLAKRIYYVDTVTG SEQ ID NO: 33 MIDPSDSESRLNQKFKD SEQ ID NO: 34 RIDPAKDNTKYDPKFQG SEQ ID NO: 35 RIDPYDSETLYNQKFKD SEQ ID NO: 36 WIYLGSGNTKYNEKFKG SEQ ID NO: 37 ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSS GLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPEFLGGPSV FLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFNST YRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLPPSQEEM TKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSRLTVDKSRW QEGNVFSCSVMHEALHNHYTQKSLSLSLGK SEQ ID NO: 38 AYRYALDY SEQ ID NO: 39 GGKGGFGY SEQ ID NO: 40 GHYGSSYVVY SEQ ID NO: 41 REERGFAY SEQ ID NO: 42 RGRGGSSY SEQ ID NO: 43 RGSPMITSFAY SEQ ID NO: 44 YDGYEGFAY SEQ ID NO: 45 DIKMTQSPSSMYASLGERVTITCKASQDIKSYLSWYQQKPWKSPKTLIYYATSLADGVP RFSGSGSGQHYSLTISSLESDDTATYYCLQHGESPFTFGSGTKLEIK SEQ ID NO: 46 DIQMTQSPSSLSASVGDRVTITCRASKNIGKYLAWYQQKPGKAPKLLIYSGSTLQSGVPS RFSGSGSGTDFTLTISSLQPEDFATYYCQQHNEYPYTFGQGTKVEIK SEQ ID NO: 47 DIVMSQSPSSLAVSVGEKVTMSCKSSQSLLYSSNQKNYLAWYQQKPGQSPKLLIYWAS TRDSGVPDRFTGSGSGTDFTLTISSVKAEDLAVYYCQQYYSYPLTFGAGTKLELK SEQ ID NO: 48 DIVMTQSPDSLAVSLGERATINCKSSQSLLYSSNQKNYLAWYQQKPGQPPKLLIYWAST RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 49 DVLMTQTPLSLPVSLGDQASISCRSSQSIVYSNGNTYLEWYLQKPGQSPKLLIYKVSNRF SGVPDRFSGSGSGTDFTLKISRVEAEDLGVYYCFQGSHVPYTFGGGTKLEIK SEQ ID NO: 50 DVQITQSPSFLAASPGETITINCRASKNIGKYLAWFQEKPGKTNKLLIYSGSTLQSGIPSRF SGSGSGTDFTLTISSLEPEDFAIYYCQQHNEYPYTFGGGTKLEIK SEQ ID NO: 51 EVKLVESGGGLVQPGGSRKLSCAASGFTFSDYGMAWVRQAPGKGPECVAFISNLAKRI YYVDTVTGRFTISRENAKNTLYLEMSSLRSEDTAMYYCTRAYRYALDYWGQGTTLTVS S SEQ ID NO: 52 EVQLQQSGAELVKPGASVKLSCTASGFTIKDTYMHWVKQRPEQGLEWIGRIDPAKDNT KYDPKFQGKATITLDTSSNIAYLQLSSLTSEDTAVYFCARGHYGSSYVVYWGQGTLVTV SA SEQ ID NO: 53 EVQLQQSGPELVKPGASVKISCKASGYSFTGYYMNWVKQSPEKSLEWIGEINPSTGGTT YNQKFKAKATLTVDKSSSTAYMQLKSLTFEDSAVYYCAIYDGYEGFAYWGQGTLVTVS A SEQ ID NO: 54 QIVLTQSPAIMSASPGEKVTMTCSASSSVSYMNWYQQKSGTSPKRWIYDTSKLASGVPA RFSGSGSGTSYSLTISSMEAEDAATYYCQQWNSNPPTFGTGTKLELK SEQ ID NO: 55 EVKLVESGGGLVQPGGSRKLSCAASGFTFSDYGMAWVRQAPGKGPECVAFISNLAKRI YYVDTVTGRFTISRENAKNTLYLEMSSLRSEDTAMYYCTRAYRYALDYWGQGTTLTVS S SEQ ID NO: 56 DIKMTQSPSSMYASLGERVTITCKASQDIKSYLSWYQQKPWKSPKTLIYYATSLADGVP SRFSGSGSGQHYSLTISSLESDDTATYYCLQHGESPFTFGSGTKLEIK SEQ ID NO: 57 DIVMSQSPSSLAVSVGEKVTMSCKSSQSLLYSSNQKNYLAWYQQKPGQSPKLLIYWAS TRDSGVPDRFTGSGSGTDFTLTISSVKAEDLAVYYCQQYYSYPLTFGAGTKLELK SEQ ID NO: 58 NILMTQSPKSMSMSVGERVTLSCKASENVGTYVSWYQQKPDQSPKLLIYGASNRYTGV PDRFTGSGSATDFTLTISSVQAEDLADYHCGQSYSYPLTFGAGTKLELK SEQ ID NO: 59 EVQLQQSGPELVKPGASVKISCKASGYSFTGYYMNWVKQSPEKSLEWIGEINPSTGGTT YNQKFKAKATLTVDKSSSTAYMQLKSLTFEDSAVYYCAIYDGYEGFAYWGQGTLVTVS A SEQ ID NO: 60 QVQLQQSGPQLVRPGASVKISCKTSGYSFTNYWMQWVKQRPGQGLEWIGMIDPSDSES RLNQKFKDKATLTVDKSSSTAYMQLSSPTFEDSAVYYCARRGSPMITSFAYWGQGTLVT VSA SEQ ID NO: 61 QVQLQQPGAELVKPGASVKMSCKASGYTFTNYWIAWVKRRPGQGLEWIGDFYPGNTS TNYNEKFKTKATLTIDTSSSTAYMQLSSLTSEDSAVYYCARRGRGGSSYWGQGTTLTVS S SEQ ID NO: 62 QIQLQQSGPELVKPGASVKISCKVSGYIFTDFYINWVKQRPGQGLEWIGWIYLGSGNTK YNEKFKGKATLTVDTSSSTAYMQLSSLTSEDSAVYFCARREERGFAYWGQGTLVTVSA SEQ ID NO: 63 QVQLQQPGAELVRPGASVKLSCKASGYTFTNYWMNWFKQRPEQGLEWIGRIDPYDSE TLYNQKFKDKAILTVDKSSSTAYMQLSSLTSEDSAVYYCAGGGKGGFGYWGQGTLVTV SA SEQ ID NO: 64 EVQLQQSGAELVKPGASVKLSCTASGFTIKDTYMHWVKQRPEQGLEWIGRIDPAKDNT KYDPKFQGKATITLDTSSNIAYLQLSSLTSEDTAVYFCARGHYGSSYVVYWGQGTLVTV SA SEQ ID NO: 65 DVLMTQTPLSLPVSLGDQASISCRSSQSIVYSNGNTYLEWYLQKPGQSPKLLIYKVSNRF SGVPDRFSGSGSGTDFTLKISRVEAEDLGVYYCFQGSHVPYTFGGGTKLEIK SEQ ID NO: 66 DVLMTQTPLSLPVSLGEQASISCRSSQSIVHSNGNTYLEWYLQKPGQSPKLLIYRVANRF SGVPDRFSGSGSGTDFTLKISRVEAEDLGVYYCFQGSHVPWTFGGGTKLEIK SEQ ID NO: 67 DVLMTQTPLSLPVSLGEQASISCRSSQSIVHSNGNTYLEWYLQKPGQSPKLLIYRVANRF SGVPDRFSGSGSGTDFTLKISRVEAEDLGVYYCFQGSHVPWTFGGGTKLEIK SEQ ID NO: 68 DVQITQSPSFLAASPGETITINCRASKNIGKYLAWFQEKPGKTNKLLIYSGSTLQSGIPSRF SGSGSGTDFTLTISSLEPEDFAIYYCQQHNEYPYTFGGGTKLEIK SEQ ID NO: 69 NILMTQSPKSMSMSVGERVTLSCKASENVGTYVSWYQQKPDQSPKLLIYGASNRYTGV PDRFTGSGSATDFTLTISSVQAEDLADYHCGQSYSYPLTFGAGTKLELK SEQ ID NO: 70 QIQLQQSGPELVKPGASVKISCKVSGYIFTDFYINWVKQRPGQGLEWIGWIYLGSGNTK YNEKFKGKATLTVDTSSSTAYMQLSSLTSEDSAVYFCARREERGFAYWGQGTLVTVSA SEQ ID NO: 71 QIVLTQSPAIMSASPGEKVTMTCSASSSVSYMNWYQQKSGTSPKRWIYDTSKLASGVPA RFSGSGSGTSYSLTISSMEAEDAATYYCQQWNSNPPTFGTGTKLELK SEQ ID NO: 72 QVQLQQPGAELVKPGASVKMSCKASGYTFTNYWIAWVKRRPGQGLEWIGDFYPGNTS TNYNEKFKTKATLTIDTSSSTAYMQLSSLTSEDSAVYYCARRGRGGSSYWGQGTTLTVS S SEQ ID NO: 73 QVQLQQPGAELVRPGASVKLSCKASGYTFTNYWMNWFKQRPEQGLEWIGRIDPYDSE TLYNQKFKDKAILTVDKSSSTAYMQLSSLTSEDSAVYYCAGGGKGGFGYWGQGTLVTV SA SEQ ID NO: 74 QVQLQQSGPQLVRPGASVKISCKTSGYSFTNYWMQWVKQRPGQGLEWIGMIDPSDSES RLNQKFKDKATLTVDKSSSTAYMQLSSPTFEDSAVYYCARRGSPMITSFAYWGQGTLVT VSA SEQ ID NO: 75 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDFYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 76 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDFYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITVDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 77 QVQLVQSGAEVKKPGASVKVSCKASGYTFTNYWMQWVRQAPGQGLEWMGMIDPSD SESRLNQKFKDRVTMTRDTSTSTVYMELSSLRSEDTAVYYCARRGSPMTTSFAYWGQGT LVTVSS
SEQ ID NO: 78 QVQLVQSGAEVKKPGASVKVSCKASGYTFTNYWMQWVRQAPGQGLEWMGMIDPSD SESRLNQKFKDRVTMTVDTSTSTVYMELSSLRSEDTAVYYCARRGSPMITSFAYWGQGT LVTVSS SEQ ID NO: 79 QVQLVQSGAEVKKPGASVKVSCKTSGYTFTNYWMQWVRQAPGQGLEWMGMIDPSDS ESRLNQKFKDRVTMTVDTSTSTVYMELSSLRSEDTAVYYCARRGSPMITSFAYVVGQGTL VTVSS SEQ ID NO: 80 QVQLVQSGAEVKKPGASVKVSCKVSGYTFTDFYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITVDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 81 MWPLVAALLLGSACCGSAQLLFNKTKSVEFTFCNDTVVIPCFVTNMEAQNTTEVYVK WKFKGRDIYTFDGALNKSTVPTDFSSAKIEVSQLLKGDASLKMDKSDAVSHTGNYTCEV TELTREGETIIELKYRVVSWFSPNENILIVIFPIFAILLFWGQFGIKTLKYRSGGMDEKTIAL LVAGLVITVIVIVGAILFVPGEYSLKNATGLGLIVTSTGILILLHYYVFSTAIGLTSFVIAILV IQVIAYILAVVGLSLCIAACIPMHGPLLISGLS1LALAQLLGLVYMKFVE SEQ ID NO: 82 MWPLAAALLLGSCCCGSAQLLFSNVNSIEFTSCNETVVIPCIVRNVEAQSTEEMFVKWK LNKSYIFIYDGNKNSTTTDQNFTSAKISVSDLINGIASLKMDKRDAMVGNYTCEVTELSR EGKTVIELKNRTAFNTDQGSACSYEEEKGGCKLVSWFSPNEKILIVIFPILAILLFWGKFGI LTLKYKSSHTNKRIILLLVAGLVLTVIVVVGAILLIPGEKPVKNASGLGLIVISTGILILLQY NVFMTAFGMTSFTIAILITQVLGYVLALVGLCLCIMACEPVHGPLLISGLGIIALAELLGLV YMKFVASNQRTIQPPRNR SEQ ID NO: 83 MWPLVAALLLGSACCGSAQLLFNKTKSVEFTFCNDTVVIPCFVTNMEAQNTTEVYVK WKFKGRDIYTFDGALNKSTVPTDFSSAKIEVSQLLKGDASLKMDKSDAVSHTGNYTCEV TELTREGETIIELKYRVVSWFSPNENILIVIFPIFAILLFWGQFGIKTLKYRSGGMDEKTIAL LVAGLVITVIVIVGAILFVPGEYSLKNATGLGLIVTSTGILILLHYYVFSTAIGLTSFVIAILV IQVIAYILAVVGLSLCIAACIPMHGPLLISGLSILALAQLLGLVYMKEVASNQKTIQPPRNN SEQ ID NO: 84 MWPLAAALLLGSCCCGSAQLLFSNVNSIEFTSCNETVVIPCIVRNVEAQSTEEMFVKWK LNKSYIFIYDGNKNSTTTDQNFTSAKISVSDLINGIASLKMDKRDAMVGNYTCEVTELSR EGKTVIELKNRTVSWFSPNEKILIVIFPILAILLFWGKFGILTLKYKSSHTNKRIILLLVAGL VLTVIVVVGAILLIPGEKPVKNASGLGLIVVSTGILILLQYNVFMTAFGMTSFTIAILITQVL GYVLALVGLCLCIMACEPVHGPLLISGLGIIALAELLGLVYMKFVASNQRTIQPPRNR SEQ ID NO: 85 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKIKSLQPEDEGSYYCQHHYGTPLTEGGGTKLELK SEQ ID NO: 86 EVQLQQSGTELVKPGASVKLSCKASGYTFISYWMHWVKQRPGQGLEWIGNINPSSGNT NYNEKFKTKATLTVDKSSSTAYMQLSSLTSEDSAVYYCVRDYYGNYWGQGTTVTVSS SEQ ID NO: 87 DIKMNQSPSSLSASLGDTITITCHASQNINVWLSWYQQKPGNIPKLLIYKASNLHTGVPS RFSGSGSGTGFTLTISSLQPEDIATYYCQQGQSYPLTFGAGTKLELK SEQ ID NO: 88 EVQLQQSGPELVKPGASVKLSCKASGFTFTNYYIHWVKQRPGQGPEWIGWIYLGSGNT KYNEKFKGKATLTADTSSSTAYLQLSTLISEDSAVYYCARYDYDLYLDSWGQGTTVTVS S SEQ ID NO: 89 DIQMMQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGV PSRFSGSGSGTQFSLKINSLQPEDFGNYYCQHHYGTPLAFGAGTKLEIK SEQ ID NO: 90 EVQLQQPGAELVKPGTSVKLSCKASGYSFTAYWMHWVKQRPGQGLEWIGNINPSSGD SHYNEKFKSKATLTVDKSSSTAYTQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 91 DIQMTQSPASLSASVGETVTITCRASENVYSYLAWYQQKQGKSPQLLVYKANTLAEGV PSRFSGSGSGTQFSLKINSLQPEDFGTYYCQHHYGTPLTFGSGTKLEIK SEQ ID NO: 92 EVQLQQPGAELVRPGSSVKLSCKASGYSFTTYWMHWVKQRPGQGLEWIGNINPSSGDS HYNEKFKSKATLTVD KSSSTAYLQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 93 DIQMIQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLRINTLQPEDLGTYYCQHHYGAPLSFGGGTKLEIKR SEQ ID NO: 94 EVQLQQPGAELVRPGTSVKLSCKASGYTFTSYWMHWVKQRPGQGLEWIGNINPSSGDS HYNEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 95 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 96 EVQLQQPGTELVKPGASVKLSCKASGYTFTSYWMHWVKQRPGQGLEWIGNINPSSGD AHYSEEFKSKATLTVDKSSSTAYMQLSSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 97 DIQINQSPSSLSASLGDTITITCHASQNINVWLSWYQQKPGNIPKLLIYKASNLHTGVPSR FSGSGSGTGFTLTISSLQPEDIATYYCQQGQSYPLTFGAGTKLEIK SEQ ID NO: 98 EVQLQQSGPELVKPGASVKISCKASGYSFTNYHIHWVKQRPGQGLEWIGWIYPGSGNT KYNEKFKGMAILTADTSSSTAYMQLSSLTSEDSAVYYCARYDYDLYLHSWGQGTTVTV SS SEQ ID NO: 99 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 100 EVQLQQPGTEVVKPGASVKLSCKASGYSFTTYWMHWVKQRPGQGLEWIGNINPSSGD SHYNEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVS A SEQ ID NO: 101 RHCFNQSPAIIVISASPGEKVTMTCSASSSVSYMHVVYQQKSGTSPKRWIYDTSNLASGVP VRFSGSGSGTSYSLTISRMEAEDAATYYCQQWSNYPFTFGSGTKLELK SEQ ID NO: 102 RVQLQQSGAELVRPGTSVKVSCKASAYAFTNYLIEWVKKRPGQGLEWIGVINPGSGGT NYNEKFKGKATLTADKSSSTAYMQLSSLTSEDSAVYFCARGDGYGSLFAYWGQGTLVT VSA SEQ ID NO: 103 DIQMIQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYGAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGIPLTFGAGTKLEIK SEQ ID NO: 104 EVQLQQPGTELVKPGASVKLSCKASGYTFISYWMHWVKQRPGQGLEWIGNINPSSGNT NYNEKFKTKATLTVDKSSSTAYMQLSSLTSEDSAVYYCVRDYYGNYWGQGTSVTVSS SEQ ID NO: 105 DIQMIQSPASLSASVGETVTITCRASENIYSYLAWYQQKLGKSPQLLVYNAKTLAEGVP SRFSGSGSGTHFSLKINSLQPEDFGTYYCQHHYGNSLTFGAGTKLELK SEQ ID NO: 106 EVQLQQSGTELVKPGASVKLSCKASGYPFTSYWMHWVKQRPGQGLEWIGNINPSSGGT NYNEKFKTKATLTVDKSSSTAYMQLSSLTSEDSAVYYCARDYYGNYWGQGTTVTVSS SEQ ID NO: 107 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 108 EVQLQQPGTELVKPGASVKLSCKTSGYSFVTYWMHWVKQRPGQGLEWIGNINPSSGDS HYSEKFKSKATLTVDKSSSTAYMQLKSLTSEDSAVYYCVRDYYGAYWGQGTLVTVSA SEQ ID NO: 109 DAVVTQESALTTSPGETVTLTCRSSTGAVTTNNYANVVVQEKPDHLFTGLIGGTNNRAP GVPARFSGSLIGDKAALTITGAQTEDEAIYFCALWYSNHLVFGGGTKLTVL SEQ ID NO: 110 PRGKVQESGPGLVKPSQSLSLTCSVTGYSITSGYYWNWIRQFPGNKLEWMGYINYDGS NNYNPSLKNRISITRDTSKNQFFLKLNSVTTEDTATYYCARGYYYGSSYGYWYFDVWG TGTTVTVSS SEQ ID NO: 111 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYAATNLADGVP SRFSGSRSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 112 EVQLQQPGTELAKPGASVKLSCKASGYTFISYWMHWVKQRPGQGLEWIGNINPSSGGT NYNEKFKSKATLTVDKSSSTAYMQLSSLTSEDSAVYYCVRDYYGSYWGQGTTVTVSS SEQ ID NO: 113 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYKAKTLVEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 114 RGPTAATWTELVKPGTSVKLSCKASGYTFISYWMHWVKQRPGQGLEWIGNINPSSGDT SYNEKFKSKATLTVDKSSSTAYMQLSSLTSEDSAVYYCARDYYGAYWGQGTSVTVSA SEQ ID NO: 115 DIQMMQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGV PSRFSGSGSGTQFSLKINSLQPEDFGNYYCQHHYGTPLTFGSGTKLELK SEQ ID NO: 116 EVQLQQPGTELVKPGASVKVSCKASGYTFTSYWMHWVKQRPGQGLEWIGNINPSSGD SHYSEKFRSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 117 DAVVTQESALTTSPGETVTLTCRSSTGAVSTSNYANWVQEKPDHLFTGLIGGTNNRAPG VPARFSGSLIGDKAALTITGAQTEDEAIYFCALWYSNHWVFGGGTKLTVL SEQ ID NO: 118 EVQLQQPGTELVKPGASVKLSCKASGYTFTSYWMHWVKQRPGQGLEWIGNVVPNND GTNYNEKFRNKATLTVDKSSSTAYMQLSSLTSEDSAVYYCAVTYFAYWGQGTLVTVSA SEQ ID NO: 119 DIQMMQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGV PSRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 120 EVQLQQSGAELVKPGASVKLSCKASGYSFTTYWMHWVKQRPGQGLEWIGNINPSSGS AHYNEKFKSKATLTVDKSSNTAYMQLSSLTSEDSAVYYCARDYYGAYWGHGTLVTVS A SEQ ID NO: 121 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTHFSLKINSLQPEDEGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 122 EVQLQQPGAELVRPGSSVKLSCKASGYTFISYWIHWVKQRPGQGLEWIGNINPSSGSSN YNEKFKNKATLTVDKSSSTAYMQLSSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 123 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGNYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 124 EVQLQQPGTELVKPGASVKLSCKASGYSFITYWMHWVKQRPGQGLEWIGNINPSSGDS HYSEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCVRDYYGAYWGHGTLVTVSA SEQ ID NO: 125 DIKINQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYKAKTLVEGVPS RFSGSGSGTQFSLKISSLQPEDEGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 126 EVQLQQPGAELVKPGASVKLSCKASGYTFISYWMHWVRQRPGQGLEWIGNINPSSGNT NYNEKFKTKATLTVDKSSSTAYMQLSSLTSEDSAVYYCVRDYYGNYWGQGTSVTVSS SEQ ID NO: 127 GIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYKAKTLVEGVP SRFSGSGSGTQFSLKINSLQPEDEGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 128 EVQLQQPGTELVKPGASVKLSCKASGYTFISYWIHWVKQRPGQGLEWIGNINPSSGGTN YNEKFKSKATLTVDKSSSTAYMHLNSLTSEDSAVYYCARDYYGNYWGQGTTLTVSS SEQ ID NO: 129 DIQMTQSPASLSASVGETVTITCRASDNIYSYLAWYQQKQGKSPQLLVYNAKTSAEGVP SRFSGSGSGTQFSLKINSLQPEDEGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 130 EVQLQQPGTELVKPGASVKLSCKASGYSFTTYWMHWVKQRPGQGLEWIGNINPSSGDS HYNEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 131 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINNLQPEDFGSYYCQHHYGTPLTFGAGTKLEMK SEQ ID NO: 132 EVQLQQSGAEHVRPGSSVKLSCKASGYSFITYWMHWVKQRPGQGLEWIGNINPSSGDS HYNEKFKSKATLTVDKSSSTAYMQLSSLTSEDSAVYYCARDYYGAYVVGQGTLVTVSA SEQ ID NO: 133 DIKMNQSPSSLSASLGDTITITCHASQNINVWLSWYQQKPGNIPKLLIYKASNLHTGVPS RFSGSGSGTGFTLTISSLQPEDIATYYCQQGQSYPLTFGTGTKLEIK SEQ ID NO: 134 EVQLQQSGPELVKPGASVKISCKASGDSFTSDHIHWVKQRPGQGLEWIGWIYPGSGNTK YNEKFKGKATLTADTSSSTAYMQLSRLTSEDSAVYYCVTYDYDLYFDNWGQGTTLTVS S
SEQ ID NO: 135 STLMTQSPASLAVSLGQRATISCRASESVDSYGNSFMHWYQQKPGQPPKLLIYLASNLE SGVPARFSGSGSGTDFILNIHPVEEEDAATYYCLHSRELPFTFGSGTKLELK SEQ ID NO: 136 EVKLMESGGGLVKPGGSLKLSCAASGFTFSDYGMHWVRQAPEKGLEWVAYISSGSSTI YYADTVKGRFTISRDNAKNTLFLQMTSLRSEDTAMYYCARPDDGYYGFAYWGQGTLV TVSA SEQ ID NO: 137 DIQMTQSPASLSASVGETVTITCRASENVYSYLAWYQQKQGKSPQLLVYNAKTLAEGV PSRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 138 EVQLQQPGTELVKPGASVKLSCKASGYSFTTYWMHWVKQRPGQGLEWIGNINPSSGDS HYNEKFKSKATLTVDKSSSTAYMHLSSLTSEDSAVYYCARDYYGAYVVGHGTLVTVSA SEQ ID NO: 139 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVFNAKTLAEGVP SRFSGSGSGTHFSLKINNLQPEDFGTYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 140 EVQLQQSGAELVKPGASVKLSCKASGYSFISYWIHWVKQRPGQGLEWIGNINPSSGGNT YNEKFKNKATLTVDKSSSTAYMQLSRLTSEDSAVYYCTRDYYGAYWGQGTLVTVSA SEQ ID NO: 141 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLAFGSGTKLELK SEQ ID NO: 142 EVQLQQPGTELVKPGASVKLSCKASGYSFITYWMHWVKQRPGQGLEWIGNINPSGGDS HYSEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCVRDYYGAYWGHGTLVTVSA SEQ ID NO: 143 DIQMIQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVFNAKTLAEGVPS RFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLEIK SEQ ID NO: 144 EVQLQQSGAELVKPGASVKLSCKASGYSFISYWIHWVKQRPGQGLEWIGNINPSSGGSS YNEKFKSKATLTVDKSSSTAYMQLSSLTSEDSAVYYCTRDYYGAYWGQGTLVTVSA SEQ ID NO: 145 DIVLSQSPAIMSASPGEKVTMTCSASSSVSYMHWYQQKSGTSPKRWIYDTSKLASGVPA RFSGSGSGTSYSLTLSSMAAEDAATYSCQQWSGNSPTFGAGTKLEIK SEQ ID NO: 146 EVQLVESGGGLVQPKGSLKLSCAASGFTENTYAMHWVRQAPGKGLEWVARIRSKSSN YATYYADSVKDRFTISRDDSQSIVYLQMNNLKTEDTAMYYCVRAWDYGSSWDYFDYW GQGTSVTVSS SEQ ID NO: 147 DIQMMQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAQTLAEGV PSRFSGSGSGTQFSLKINSLQSEDIGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 148 RGPTQQPGTELVKPGASVKLSRKASGYTFTTYWMHWVKQRPGQGLEWIGN1NPSSGDS HYNEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 149 GYGMS SEQ ID NO: 150 TITSGGTYTYYPDSVKG SEQ ID NO: 151 SLAGNAMDY SEQ ID NO: 152 RASQTISDYLH SEQ ID NO: 153 FASQSIS SEQ ID NO: 154 QNGHGFPRT SEQ ID NO: 155 NYNMH SEQ ID NO: 156 TIYPGNDDTSYNQKFKD SEQ ID NO: 157 GGYRAMDY SEQ ID NO: 158 RSSQSIVYSNGNTYLG SEQ ID NO: 159 KVSNRFS SEQ ID NO: 160 FQGSHVPYT SEQ ID NO: 161 MWPLVAALLLGSACCGSAQLLFNKTKSVEFTFCNDTVVIPCFVTNMEAQNTTEVYVK WKFKGRDIYTFDGALNKSTVPTDFSSAKIEVSQLLKGDASLKMDKSDAVSHTGNYTCEV TELTREGETIIELKYRVVSWFSP SEQ ID NO: 162 DNIYSY SEQ ID NO: 163 ENIYSY SEQ ID NO: 164 ENVYSY SEQ ID NO: 165 ESVDSYGNSF SEQ ID NO: 166 QNINVW SEQ ID NO: 167 SSVSY SEQ ID NO: 168 TGAVSTSNY SEQ ID NO: 169 TGAVTTNNY SEQ ID NO: 170 AAT SEQ ID NO: 171 DTS SEQ ID NO: 172 GAK SEQ ID NO: 173 GTN SEQ ID NO: 174 KAK SEQ ID NO: 175 KAN SEQ ID NO: 176 KAS SEQ ID NO: 177 LAS SEQ ID NO: 178 NAK SEQ ID NO: 179 NAQ SEQ ID NO: 180 ALWYSNHLV SEQ ID NO: 181 ALWYSNHWV SEQ ID NO: 182 LHSRELPFT SEQ ID NO: 183 QHHYGAPLS SEQ ID NO: 184 QHHYGIPLT SEQ ID NO: 185 QHHYGNSLT SEQ ID NO: 186 QHHYGTPLA SEQ ID NO: 187 QHHYGTPLT SEQ ID NO: 188 QQGQSYPLT SEQ ID NO: 189 QQWSGNSPT SEQ ID NO: 190 QQWSNYPFT SEQ ID NO: 191 AYAFTNYL SEQ ID NO: 192 GDSFTSDH SEQ ID NO: 193 GFTFNTYA SEQ ID NO: 194 GFTFSDYG SEQ ID NO: 195 GFTFTNYY SEQ ID NO: 196 GYPFTSYW SEQ ID NO: 197 GYSFISYW SEQ ID NO: 198 GYSFITYVV SEQ ID NO: 199 GYSFTAYVV SEQ ID NO: 200 GYSFTNYH SEQ ID NO: 201 GYSFTTYW SEQ ID NO: 202 GYSFVTYVV SEQ ID NO: 203 GYSITSGYY SEQ ID NO: 204 GYTFISYVV SEQ ID NO: 205 GYTFTSYW SEQ ID NO: 206 GYTFTTYW SEQ ID NO: 207 INPGSGGT SEQ ID NO: 208 INPSGGDS SEQ ID NO: 209 INPSSGDA SEQ ID NO: 210 INPSSGDS SEQ ID NO: 211 INPSSGDT SEQ ID NO: 212 INPSSGGN SEQ ID NO: 213 INPSSGGS SEQ ID NO: 214 INPSSGGT SEQ ID NO: 215 INPSSGNT SEQ ID NO: 216 INPSSGSA SEQ ID NO: 217 INPSSGSS SEQ ID NO: 218 INYDGSN SEQ ID NO: 219 IRSKSSNYAT SEQ ID NO: 220 ISSGSSTI SEQ ID NO: 221 IYLGSGNT SEQ ID NO: 222 IYPGSGNT SEQ ID NO: 223 VVPNNDGT SEQ ID NO: 224 ARDYYGAY SEQ ID NO: 225 ARDYYGNY SEQ ID NO: 226 ARGDGYGSLFAY SEQ ID NO: 227 ARGYYYGSSYGYWYFDV SEQ ID NO: 228 ARPDDGYYGFAY SEQ ID NO: 229 ARYDYDLYLDS SEQ ID NO: 230 ARYDYDLYLHS SEQ ID NO: 231 AVTYFAY SEQ ID NO: 232 TRDYYGAY SEQ ID NO: 233 VRAWDYGSSWDYFDY SEQ ID NO: 234 VRDYYGAY SEQ ID NO: 235 VRDYYGNY SEQ ID NO: 236 VRDYYGSY SEQ ID NO: 237 VTYDYDLYFDN SEQ ID NO: 238 DIQMTQSPASLSASVGETVTITCRASENIYSYLAWYQQKQGKSPQLLVYNAKTLAEGVP SRFSGSGSGTQFSLKINSLQPEDFGSYYCQHHYGTPLTFGAGTKLELK SEQ ID NO: 239 EVQLQQPGTELVKPGASVKLSCKASGYSFTTYWMHWVKQRPGQGLEWIGN1NPSSGDS HYNEKFKSKATLTVDKSSSTAYMQLNSLTSEDSAVYYCARDYYGAYWGHGTLVTVSA SEQ ID NO: 240 EVQLVESGGDLVKPGGSLKLSCAASGFTFSGYGMSWVRQTPDKRLEWVATITSGGTYT YYPDSVKGRFTISRDNAKNTLYLQIDSLKSEDTAIYFCARSLAGNAMDYWGQGTSVTVS S
SEQ ID NO: 241 DIVMTQSPATLSVTPGDRVSLSCRASQTISDYLHWYQQKSHESPRLLIKFASQSISGIPSR FSGSGSGSDFTLSINSVEPEDVGVYYCQNGHGFPRTFGGGTKLEIK SEQ ID NO: 242 QVQLQQPGAELVKPGASVMMSCKASGYTFTNYNMHWVKQTPGQGLEWIGTIYPGND DTSYNQKFKDKATLTADKSSSAAYMQLSSLTSEDSAVYYCARGGYRAMDYWGQTSVT VSS SEQ ID NO: 243 DVLMTQTPLSLPVSLGDQASISCRSSQSIVYSNGNTYLGWYLQKPGQSPKLLIYKVSNRF SGVPDRFSGSGSGTDFTLKISRVEAEDLGVYHCFQGSHVPYTFGGGTKVEIK SEQ ID NO: 244 QMQLVQSGAEVKKTGSSVKVSCKASGFNIKDYYLHWVRQAPGQALEWMGWIDPDQG DTEYAQKFQDRVTITRDRSMSTAYMELSSLRSEDTAMYYCNAAYGSSSYPMDYWGQG TTVTV SEQ ID NO: 245 NIQMTQSPSAMSASVGDRVTITCKASQDIHRYLSWFQQKPGKVPKHLIYRANRLVSGVP SRFSGSGSGTEFTLTISSLQPEDFATYYCLQYDEFPYTFGGGTKVEIK SEQ ID NO: 246 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDSYINWVRQAPGQRLEWMGWIYTGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 247 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDIYINWVRQAPGQRLEWMGWIYLGSGN VKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREDRGFAYWGQGTLVTV SS SEQ ID NO: 248 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDNYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAVWGQGTLVT VSS SEQ ID NO: 249 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDFYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 250 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDHYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREDRGFAYWGQGTLVT VSS SEQ ID NO: 251 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDNYINWVRQAPGQRLEWMGWVYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 252 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDLYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 253 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDNYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAPWGQGTLVTV SS SEQ ID NO: 254 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDSYINWVRQAPGQRLEWMGWIYPGSGN TKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVTVS S SEQ ID NO: 255 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDFYIHWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREEDGFAHWGQGTLVT VSS SEQ ID NO: 256 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDNYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARRKERGFAYWGQGTLVT VSS SEQ ID NO: 257 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDTYINWVRQAPGQRLEWMGWIYLGSG NIKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVTV SS SEQ ID NO: 258 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDYYINWVRQAPGQRLEWMGWIYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREDRGFAYWGQGTLVT VSS SEQ ID NO: 259 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDTYINWVRQAPGQRLEWMGWIYLGSG NVKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAHWGQGTLVT VSS SEQ ID NO: 260 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDLYINWVRQAPGQRLEWMGWIYLGSG NVKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 261 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDSYINWVRQAPGQRLEWMGWIYGGSG NTKYNEKFKGRVTITRDTSASTTYMELSSLRSEDTAVYYCARREEDGFAYWGQGTLVT VSS SEQ ID NO: 262 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDSYINVVVRQAPGQRLEWMGWIYGGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAYWGQGTLVT VSS SEQ ID NO: 263 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDSYINWVRQAPGQRLEWMGWIYLGSG NVKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARRIERGFAYWGQGTLVTV SS SEQ ID NO: 264 QVQLVQSGAEVKKPGASVKVSCKASGYTFTDSYINWVRQAPGQRLEWMGWVYLGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARREERGFAVWGQGTLVT VSS SEQ ID NO: 265 DIVMTQSPDSLAYSLGERATINCKSSQSILYSSNQKNYLAWYQQKPGQPPKLLIYWASV RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 266 DIVMTQSPDSLAVSLGERATINCKSSQSLLSSNQKYYLAWYQQKPGQPPKLLIYWASTR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYSYPLTFGQGTKVEIK SEQ ID NO: 267 DIVMTQSPDSLAYSLGERATINCKSSQSLYSSNKKNYLAWYQQKPGQPPKLLIYWASSR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 268 DIVMTQSPDSLAVSLGERATINCKSSQSLLSSNQKNYLAWYQQKPGQPPKLLIYWASVR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYSYPLTFGQGTKVEIK SEQ ID NO: 269 DIVMTQSPDSLAVSLGERATINCKSSQSLLSSSQKNYLAWYQQKPGQPPKLLIYWASVR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 270 DIVMTQSPDSLAVSLGERATINCKSSQSLLYSSNKKNYLAWYQQKPGQPPKLLIYWASV RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 271 DIVMTQSPDSLAVSLGERATINCKSSQSLVSSSQKNYLAWYQQKPGQPPKLLIYWASVR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 272 DIVMTQSPDSLAVSLGERATINCKSSQSLLSSSQKNYLAWYQQKPGQPPKLLIYWASTR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYSYPLTFGQGTKVEIK SEQ ID NO: 273 DIVMTQSPDSLAVSLGERATINCKSSQSLLYSSNQKNYLAWYQQKPGQPPKWYWASV RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYSYPLTFGQGTKVEIK SEQ ID NO: 274 DIVMTQSPDSLAVSLGERATINCKSSQSLLYKSNQKNYLAWYQQKPGQPPKLLIYWAS VRDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQLYSYPLTFGQGTKVEIK SEQ ID NO: 275 DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNQKNYLAWYQQKPGQPPKLLIYWAS VRDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 276 DIVMTQSPDSLAVSLGERATINCKSSQSLLSSSQKNYLAWYQQKPGQPPKLLIYWASGR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 277 DIVMTQSPDSLAVSLGERATINCKSSQSLLTSNQKNYLAWYQQKPGQPPKWYWASIR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 278 DIVMTQSPDSLAVSLGERATINCKSSQSLLYSSNRKNYLAWYQQKPGQPPKLLIYWASS RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 279 DIVMTQSPDSLAVSLGERATINCKSSQSLLYSSNNKNYLAWYQQKPGQPPKLLIYWASS RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYSYPLTFGQGTKVEIK SEQ ID NO: 280 DIVMTQSPDSLAVSLGERATINCKSSQSLLSSNNKNYLAWYQQKPGQPPKLLIYWASVR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 281 DIVMTQSPDSLAVSLGERATINCKSSQSLYTSNQKNYLAWYQQKPGQPPKLLIYWASTR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 282 DIVMTQSPDSLAVSLGERATINCKSSQSLLYNSNQKNYLAWYQQKPGQPPKLLIYWAST RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 283 DIVMTQSPDSLAVSLGERATINCKSSQSLLYKSNQKNYLAWYQQKPGQPPKWYWAS VRDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYSYPLTFGQGTKVEIK SEQ ID NO: 284 DIVMTQSPDSLAVSLGERATINCKSSQSLYSSSQKNYLAWYQQKPGQPPKWYWASVR DSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSYPLTFGQGTKVEIK SEQ ID NO: 285 DIVMTQSPDSLAVSLGERATINCKSSQSLLYNSNQKNYLAWYQQKPGQPPKWYWAS VRDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 286 DIVMTQSPDSLAVSLGERATINCKSSQSLLYSSNQKNYLAWYQQKPGQPPKWYWASV RDSGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYHYPLTFGQGTKVEIK SEQ ID NO: 287 DIQMTQSPSSLSASVGDRVTITCRASKNIGKYLAWFQQKPGKAPKSLIYSGSTLQSGVPS RFSGSGSGTDFTLTISSLQPEDFATYYCQQHNEYPYTFGGGTKVEIK SEQ ID NO: 288 DVQITQSPSSLSASVGDRVTITCRASKNIGKYLAWFQQKPGKAPKSLIYSGSTLQSGYPS RFSGSGSGTDFTLTISSLQPEDFATYYCQQHNEYPYTFGGGTKVEIK SEQ ID NO: 289 DIQMTQSPSSLSASVGDRVTITCRASKNIGKYLAWFQQKPGKAPKSLIYSGSTLQSGIPSR FSGSGSGTDFTLTISSLQPEDFATYYCQQHNEYPYTFGGGTKVEIK SEQ ID NO: 290 DVQITQSPSSLSASVGDRVTITCRASKNIGKYLAWFQQKPGKAPKSLIYSGSTLQSGIPSR FSGSGSGTDFTLTISSLQPEDFATYYCQQHNEYPYTFGGGTKVEIK SEQ ID NO: 291 DVQITQSPSSLSASVGDRVTITCRASKNIGKYLAWFQQKPGKTNKLLIYSGSTLQSGIPSR FSGSGSGTDFTLTISSLQPEDFATYYCQQHNEYPYTFGGGTKVEIK SEQ ID NO: 292 QVQLVQSGAEVKKPGASVKVSCKASGYTFTNYWMQWVRQAPGQGLEWMGMIDPSD SESRLNQKFKDRVTMTRDTSTSTVYMELSSLRSEDTAVYYCARRGSPMITSFAYWGQGT LVTVSS SEQ ID NO: 293 QVQLVQSGAEVKKPGASVKVSCKTSGYSFTNYWMQWVRQAPGQGLEWMGMIDPSDS ESRLNQKFKDRVTMTRDTSTSTAYMELSSLRSEDTAVYYCARRGSPMITSFAYWGQGTL VTVSS SEQ ID NO: 294 QVQLVQSGAEVKKPGASVKVSCKASGYTFTNYWMQWVRQAPGQGLEWIGM1DPSDS ESRLNQKFKDRATLTVDTSTSTVYMELSSLRSEDTAVYYCARRGSPMITSFAYWGQGTL VTVSS SEQ ID NO: 295 QVQLVQSGAEVKKPGASVKVSCKTSGYSFTNYWMQWVRQAPGQGLEWIGMIDPSDSE SRLNQKFKDRATLTVDTSTSTAYMELSSLRSEDTAVYYCARRGSPMITSFAYWGQGTLV TVSS SEQ ID NO: 296 QVQLQQSGAEVKKPGASVKVSCKTSGYSFTNYWMQWVRQAPGQGLEWIGMIDPSDSE SRLNQKFKDKATLTVDKSTSTAYMELSSLRSEDTAVYYCARRGSPMITSFAYWGQGTLV TVSS
SEQ ID NO: 297 DIQMTQSPSSVSASVGDRVTITCHASQNINVWLSWYQQKPGKAPKLLIYKASNLHTGVP SRFSGSGSGTDFTLTISSLQPEDFATYYCQQGQSYPLTFGQGTKLEIK SEQ ID NO: 298 DIQITQSPSSVSASVGDRVTITCHASQNINVWLSWYQQKPGKAPKWYKASNLHTGVPS RFSGSGSGTGFTLTISSLQPEDFATYYCQQGQSYPLTFGQGTKLEIK SEQ ID NO: 299 DIQITQSPSSVSASVGDRVTITCHASQNINVWLSWYQQKPGKIPKWYKASNLHTGVPS RFSGSGSGTGFTLTISSLQPEDFATYYCQQGQSYPLTFGQGTKLEIK SEQ ID NO: 300 QVQLVQSGAEVKKPGASVKVSCKASGYTFTNYHIHWVRQAPGQRLEWMGWIYPGSG NTKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARYDYDLYLHSWGQGTLV TVSS SEQ ID NO: 301 QVQLVQSGAEVKKPGASVKVSCKASGYSFTNYHIHWVRQAPGQRLEWIGWIYPGSGN TKYNEKFKGRVTITRDTSASTAYMELSSLRSEDTAVYYCARYDYDLYLHSWGQGTLVT VSS SEQ ID NO: 302 QVQLVQSGAEVKKPGASVKVSCKASGYSFTNYHIHWVRQAPGQRLEWIGWIYPGSGN TKYNEKFKGRATLTADTSASTAYMELSSLRSEDTAVYYCARYDYDLYLHSWGQGTLVT VSS SEQ ID NO: 303 QVQLVQSGAEVKKPGASVKVSCKASGYSFTNYHIHWVRQAPGQRLEWIGWIYPGSGN TKYNEKFKGMATLTADTSASTAYMELSSLRSEDTAVYYCARYDYDLYLHSWGQGTLV TVSS SEQ ID NO: 304 EVQLVQSGAEVKKPGASVKVSCKASGYSFTNYHIHWVKQAPGQRLEWIGWIYPGSGN TKYNEKFKGMATLTADTSASTAYMELSSLRSEDTAVYYCARYDYDLYLHSWGQGTLV TVSS SEQ ID NO: 305 QMQLVQSGAEVKKTGSSVKVSCKASGFNIKDYYLHWVRQAPGQALEWMGWIDPDQG DTEYAQKFQDRVTITRDRSMSTAYMELSSLRSEDTAMYYCNAAYGSSSYPMDYWGQG TTVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFP AVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPEF EGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKPRE EQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLP PSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSRLTV DKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK SEQ ID NO: 306 QMTQSPSAMSASVGDRVTITCKASQDIHRYLSWFQQKPGKVPKHLIYRANRLVSGVPS RFSGSGSGTEFTLTISSLQPEDFATYYCLQYDEFPYTFGGGTKVEIKRTVAAPSVFIFPPSD EQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTL SKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC SEQ ID NO: 307 QMQLVQSGAEVKKTGSSVKVSCKASGFNIKDYYLHWVRQAPGQALEWMGWIDPDQG DTEYAQKFQDRVTITRDRSMSTAYMELSSLRSEDTAMYYCNAAYGSSSYPMDYWGQG TTVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFP AVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPEF LGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKPRE EQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLP PSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSRLTV DKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK SEQ ID NO: 308 QMTQSPSAMSASVGDRVTITCKASQDIHRYLSWFQQKPGKVPKHLIYRANRLVSGYPS RFSGSGSGTEFTLTISSLQPEDFATYYCLQYDEFPYTFGGGTKVEIKRTVAAPSVFIFPPSD EQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTL SKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC SEQ ID NO: 309 DVQITQSPSFLAASPGETITINCRASKNIGKYLAWFQEKPGKTNKLLIYSGSTLQSGIPSRF SGSGSGTDFTLTISSLEPEDFAIYYCQQHNEYPYTFGGGTKLEIKRTVAAPSVFIFPPSDEQ LKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTLSK ADYEKHKVYACEVTHQGLSSPVTKSFNRGEC SEQ ID NO: 310 QVQLQQSGPQLVRPGASVKISCKTSGYSFTNYWMQWVKQRPGQGLEWIGMIDPSDSES RLNQKFKDKATLTVDKSSSTAYMQLSSPTFEDSAVYYCARRGSPMITSFAYWGQGTLVT VSAASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL QSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPEFLGG PSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLPPSQ EEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSRLTVDK SRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK SEQ ID NO: 311 DIQINQSPSSLSASLGDTITITCHASQNINVWLSWYQQKPGNIPKLLIYKASNLHTGVPSR FSGSGSGTGFTLTISSLQPEDIATYYCQQGQSYPLTFGAGTKLEIKRTVAAPSVFIFPPSDE QLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTLS KADYEKHKVYACEVTHQGLSSPVTKSFNRGEC SEQ ID NO: 312 EVQLQQSGPELVKPGASVKISCKASGYSFTNYHIHWVKQRPGQGLEWIGWIYPGSGNT KYNEKFKGMAILTADTSSSTAYMQLSSLTSEDSAVYYCARYDYDLYLHSWGQGTTVTV SSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQ SSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPEFLGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFNS TYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREPQVYTLPPSQEE MTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSRLTVDKSR WQEGNVFSCSVMHEALHNHYTQKSLSLSLGK
Sequence CWU 1
1
313117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 1Lys Ser Ser Gln Ser Leu Leu Tyr Ser Ser Asn Gln
Lys Asn Tyr Leu1 5 10 15Ala211PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 2Arg Ala Ser Lys Asn Ile Gly
Lys Tyr Leu Ala1 5 10311PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 3Lys Ala Ser Gln Asp Ile Lys
Ser Tyr Leu Ser1 5 10410PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 4Ser Ala Ser Ser Ser Val Ser
Tyr Met Asn1 5 10516PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 5Arg Ser Ser Gln Ser Ile Val Tyr Ser Asn
Gly Asn Thr Tyr Leu Glu1 5 10 15611PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 6Lys
Ala Ser Glu Asn Val Gly Thr Tyr Val Ser1 5 10716PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 7Arg
Ser Ser Gln Ser Ile Val His Ser Asn Gly Asn Thr Tyr Leu Glu1 5 10
1587PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 8Asp Thr Ser Lys Leu Ala Ser1 597PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 9Gly
Ala Ser Asn Arg Tyr Thr1 5107PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 10Lys Val Ser Asn Arg Phe
Ser1 5117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 11Arg Val Ala Asn Arg Phe Ser1 5127PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 12Ser
Gly Ser Thr Leu Gln Ser1 5137PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 13Trp Ala Ser Thr Arg Asp
Ser1 5147PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 14Tyr Ala Thr Ser Leu Ala Asp1 5159PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 15Phe
Gln Gly Ser His Val Pro Trp Thr1 5169PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 16Phe
Gln Gly Ser His Val Pro Tyr Thr1 5179PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 17Gly
Gln Ser Tyr Ser Tyr Pro Leu Thr1 518327PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
18Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg1
5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu Gly Cys Leu Val Lys Asp
Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu
Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly
Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu
Gly Thr Lys Thr65 70 75 80Tyr Thr Cys Asn Val Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys 85 90 95Arg Val Glu Ser Lys Tyr Gly Pro Pro
Cys Pro Pro Cys Pro Ala Pro 100 105 110Glu Phe Glu Gly Gly Pro Ser
Val Phe Leu Phe Pro Pro Lys Pro Lys 115 120 125Asp Thr Leu Met Ile
Ser Arg Thr Pro Glu Val Thr Cys Val Val Val 130 135 140Asp Val Ser
Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp145 150 155
160Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe
165 170 175Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His
Gln Asp 180 185 190Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Gly Leu 195 200 205Pro Ser Ser Ile Glu Lys Thr Ile Ser Lys
Ala Lys Gly Gln Pro Arg 210 215 220Glu Pro Gln Val Tyr Thr Leu Pro
Pro Ser Gln Glu Glu Met Thr Lys225 230 235 240Asn Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp 245 250 255Ile Ala Val
Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys 260 265 270Thr
Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser 275 280
285Arg Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn Val Phe Ser
290 295 300Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser305 310 315 320Leu Ser Leu Ser Leu Gly Lys
325199PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 19Leu Gln His Gly Glu Ser Pro Phe Thr1
5209PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 20Gln Gln His Asn Glu Tyr Pro Tyr Thr1
5219PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 21Gln Gln Trp Asn Ser Asn Pro Pro Thr1
5229PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 22Gln Gln Tyr Tyr Ser Tyr Pro Leu Thr1
5235PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 23Asp Phe Tyr Ile Asn1 5245PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 24Asp
Thr Tyr Met His1 5255PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 25Asp Tyr Gly Met Ala1
5265PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 26Gly Tyr Tyr Met Asn1 5275PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 27Asn
Tyr Trp Ile Ala1 5285PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 28Asn Tyr Trp Met Asn1
5295PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 29Asn Tyr Trp Met Gln1 53017PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 30Asp
Phe Tyr Pro Gly Asn Thr Ser Thr Asn Tyr Asn Glu Lys Phe Lys1 5 10
15Thr3117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 31Glu Ile Asn Pro Ser Thr Gly Gly Thr Thr Tyr Asn
Gln Lys Phe Lys1 5 10 15Ala3217PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 32Phe Ile Ser Asn Leu Ala Lys
Arg Ile Tyr Tyr Val Asp Thr Val Thr1 5 10 15Gly3317PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 33Met
Ile Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe Lys1 5 10
15Asp3417PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 34Arg Ile Asp Pro Ala Lys Asp Asn Thr Lys Tyr Asp
Pro Lys Phe Gln1 5 10 15Gly3517PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 35Arg Ile Asp Pro Tyr Asp Ser
Glu Thr Leu Tyr Asn Gln Lys Phe Lys1 5 10 15Asp3617PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 36Trp
Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe Lys1 5 10
15Gly37327PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 37Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu
Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu Gly
Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp
Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val
Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val
Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr Thr Cys Asn Val
Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Arg Val Glu Ser
Lys Tyr Gly Pro Pro Cys Pro Pro Cys Pro Ala Pro 100 105 110Glu Phe
Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys 115 120
125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val
130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr
Val Asp145 150 155 160Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg Val Val Ser Val
Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro Ser Ser Ile Glu
Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215 220Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys225 230 235
240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys Ser Arg Trp Gln
Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu Ser Leu Ser Leu
Gly Lys 325388PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 38Ala Tyr Arg Tyr Ala Leu Asp Tyr1
5398PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 39Gly Gly Lys Gly Gly Phe Gly Tyr1
54010PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 40Gly His Tyr Gly Ser Ser Tyr Val Val Tyr1 5
10418PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 41Arg Glu Glu Arg Gly Phe Ala Tyr1
5428PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 42Arg Gly Arg Gly Gly Ser Ser Tyr1
54311PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 43Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr1 5
10449PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 44Tyr Asp Gly Tyr Glu Gly Phe Ala Tyr1
545107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 45Asp Ile Lys Met Thr Gln Ser Pro Ser Ser Met
Tyr Ala Ser Leu Gly1 5 10 15Glu Arg Val Thr Ile Thr Cys Lys Ala Ser
Gln Asp Ile Lys Ser Tyr 20 25 30Leu Ser Trp Tyr Gln Gln Lys Pro Trp
Lys Ser Pro Lys Thr Leu Ile 35 40 45Tyr Tyr Ala Thr Ser Leu Ala Asp
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Gln His Tyr
Ser Leu Thr Ile Ser Ser Leu Glu Ser65 70 75 80Asp Asp Thr Ala Thr
Tyr Tyr Cys Leu Gln His Gly Glu Ser Pro Phe 85 90 95Thr Phe Gly Ser
Gly Thr Lys Leu Glu Ile Lys 100 10546107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
46Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Lys Asn Ile Gly Lys
Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Ser Gly Ser Thr Leu Gln Ser Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
His Asn Glu Tyr Pro Tyr 85 90 95Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile Lys 100 10547113PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 47Asp Ile Val Met Ser Gln Ser Pro
Ser Ser Leu Ala Val Ser Val Gly1 5 10 15Glu Lys Val Thr Met Ser Cys
Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Gln Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Ser Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Thr Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe
Thr Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Val Lys Ala Glu Asp Leu Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr
Tyr Ser Tyr Pro Leu Thr Phe Gly Ala Gly Thr Lys Leu Glu Leu 100 105
110Lys48113PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 48Asp Ile Val Met Thr Gln Ser Pro Asp Ser Leu
Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser
Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Gln Lys Asn Tyr Leu Ala Trp
Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr Trp
Ala Ser Thr Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu Gln
Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Tyr
Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys49112PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 49Asp Val Leu Met Thr Gln Thr Pro Leu Ser Leu
Pro Val Ser Leu Gly1 5 10 15Asp Gln Ala Ser Ile Ser Cys Arg Ser Ser
Gln Ser Ile Val Tyr Ser 20 25 30Asn Gly Asn Thr Tyr Leu Glu Trp Tyr
Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Lys Leu Leu Ile Tyr Lys Val
Ser Asn Arg Phe Ser Gly Val Pro 50 55 60Asp Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu Ala
Glu Asp Leu Gly Val Tyr Tyr Cys Phe Gln Gly 85 90 95Ser His Val Pro
Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys 100 105
11050107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 50Asp Val Gln Ile Thr Gln Ser Pro Ser Phe Leu
Ala Ala Ser Pro Gly1 5 10 15Glu Thr Ile Thr Ile Asn Cys Arg Ala Ser
Lys Asn Ile Gly Lys Tyr 20 25 30Leu Ala Trp Phe Gln Glu Lys Pro Gly
Lys Thr Asn Lys Leu Leu Ile 35 40 45Tyr Ser Gly Ser Thr Leu Gln Ser
Gly Ile Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Ser Leu Glu Pro65 70 75 80Glu Asp Phe Ala Ile
Tyr Tyr Cys Gln Gln His Asn Glu Tyr Pro Tyr 85 90 95Thr Phe Gly Gly
Gly Thr Lys Leu Glu Ile Lys 100 10551117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
51Glu Val Lys Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Arg Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asp
Tyr 20 25 30Gly Met Ala Trp Val Arg Gln Ala Pro Gly Lys Gly Pro Glu
Cys Val 35 40 45Ala Phe Ile Ser Asn Leu Ala Lys Arg Ile Tyr Tyr Val
Asp Thr Val 50 55 60Thr Gly Arg Phe Thr Ile Ser Arg Glu Asn Ala Lys
Asn Thr Leu Tyr65 70 75 80Leu Glu Met Ser Ser Leu Arg Ser Glu Asp
Thr Ala Met Tyr Tyr Cys 85 90 95Thr Arg Ala Tyr Arg Tyr Ala Leu Asp
Tyr Trp Gly Gln Gly Thr Thr 100 105 110Leu Thr Val Ser Ser
11552119PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 52Glu Val Gln Leu Gln Gln Ser Gly Ala Glu Leu
Val Lys Pro Gly Ala1 5
10 15Ser Val Lys Leu Ser Cys Thr Ala Ser Gly Phe Thr Ile Lys Asp
Thr 20 25 30Tyr Met His Trp Val Lys Gln Arg Pro Glu Gln Gly Leu Glu
Trp Ile 35 40 45Gly Arg Ile Asp Pro Ala Lys Asp Asn Thr Lys Tyr Asp
Pro Lys Phe 50 55 60Gln Gly Lys Ala Thr Ile Thr Leu Asp Thr Ser Ser
Asn Ile Ala Tyr65 70 75 80Leu Gln Leu Ser Ser Leu Thr Ser Glu Asp
Thr Ala Val Tyr Phe Cys 85 90 95Ala Arg Gly His Tyr Gly Ser Ser Tyr
Val Val Tyr Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val Ser Ala
11553118PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 53Glu Val Gln Leu Gln Gln Ser Gly Pro Glu Leu
Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser Cys Lys Ala Ser Gly
Tyr Ser Phe Thr Gly Tyr 20 25 30Tyr Met Asn Trp Val Lys Gln Ser Pro
Glu Lys Ser Leu Glu Trp Ile 35 40 45Gly Glu Ile Asn Pro Ser Thr Gly
Gly Thr Thr Tyr Asn Gln Lys Phe 50 55 60Lys Ala Lys Ala Thr Leu Thr
Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln Leu Lys Ser
Leu Thr Phe Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala Ile Tyr Asp
Gly Tyr Glu Gly Phe Ala Tyr Trp Gly Gln Gly Thr 100 105 110Leu Val
Thr Val Ser Ala 11554106PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 54Gln Ile Val Leu Thr Gln
Ser Pro Ala Ile Met Ser Ala Ser Pro Gly1 5 10 15Glu Lys Val Thr Met
Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr Met 20 25 30Asn Trp Tyr Gln
Gln Lys Ser Gly Thr Ser Pro Lys Arg Trp Ile Tyr 35 40 45Asp Thr Ser
Lys Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly Ser 50 55 60Gly Ser
Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Ala Glu65 70 75
80Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Trp Asn Ser Asn Pro Pro Thr
85 90 95Phe Gly Thr Gly Thr Lys Leu Glu Leu Lys 100
10555117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 55Glu Val Lys Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly1 5 10 15Ser Arg Lys Leu Ser Cys Ala Ala Ser Gly
Phe Thr Phe Ser Asp Tyr 20 25 30Gly Met Ala Trp Val Arg Gln Ala Pro
Gly Lys Gly Pro Glu Cys Val 35 40 45Ala Phe Ile Ser Asn Leu Ala Lys
Arg Ile Tyr Tyr Val Asp Thr Val 50 55 60Thr Gly Arg Phe Thr Ile Ser
Arg Glu Asn Ala Lys Asn Thr Leu Tyr65 70 75 80Leu Glu Met Ser Ser
Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys 85 90 95Thr Arg Ala Tyr
Arg Tyr Ala Leu Asp Tyr Trp Gly Gln Gly Thr Thr 100 105 110Leu Thr
Val Ser Ser 11556107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 56Asp Ile Lys Met Thr Gln Ser Pro
Ser Ser Met Tyr Ala Ser Leu Gly1 5 10 15Glu Arg Val Thr Ile Thr Cys
Lys Ala Ser Gln Asp Ile Lys Ser Tyr 20 25 30Leu Ser Trp Tyr Gln Gln
Lys Pro Trp Lys Ser Pro Lys Thr Leu Ile 35 40 45Tyr Tyr Ala Thr Ser
Leu Ala Asp Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Gln His Tyr Ser Leu Thr Ile Ser Ser Leu Glu Ser65 70 75 80Asp Asp
Thr Ala Thr Tyr Tyr Cys Leu Gln His Gly Glu Ser Pro Phe 85 90 95Thr
Phe Gly Ser Gly Thr Lys Leu Glu Ile Lys 100 10557113PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
57Asp Ile Val Met Ser Gln Ser Pro Ser Ser Leu Ala Val Ser Val Gly1
5 10 15Glu Lys Val Thr Met Ser Cys Lys Ser Ser Gln Ser Leu Leu Tyr
Ser 20 25 30Ser Asn Gln Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro
Gly Gln 35 40 45Ser Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Asp
Ser Gly Val 50 55 60Pro Asp Arg Phe Thr Gly Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr65 70 75 80Ile Ser Ser Val Lys Ala Glu Asp Leu Ala
Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser Tyr Pro Leu Thr Phe Gly
Ala Gly Thr Lys Leu Glu Leu 100 105 110Lys58107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
58Asn Ile Leu Met Thr Gln Ser Pro Lys Ser Met Ser Met Ser Val Gly1
5 10 15Glu Arg Val Thr Leu Ser Cys Lys Ala Ser Glu Asn Val Gly Thr
Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Asp Gln Ser Pro Lys Leu
Leu Ile 35 40 45Tyr Gly Ala Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg
Phe Thr Gly 50 55 60Ser Gly Ser Ala Thr Asp Phe Thr Leu Thr Ile Ser
Ser Val Gln Ala65 70 75 80Glu Asp Leu Ala Asp Tyr His Cys Gly Gln
Ser Tyr Ser Tyr Pro Leu 85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu
Leu Lys 100 10559118PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 59Glu Val Gln Leu Gln Gln Ser Gly
Pro Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser Cys Lys
Ala Ser Gly Tyr Ser Phe Thr Gly Tyr 20 25 30Tyr Met Asn Trp Val Lys
Gln Ser Pro Glu Lys Ser Leu Glu Trp Ile 35 40 45Gly Glu Ile Asn Pro
Ser Thr Gly Gly Thr Thr Tyr Asn Gln Lys Phe 50 55 60Lys Ala Lys Ala
Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln
Leu Lys Ser Leu Thr Phe Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala
Ile Tyr Asp Gly Tyr Glu Gly Phe Ala Tyr Trp Gly Gln Gly Thr 100 105
110Leu Val Thr Val Ser Ala 11560120PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
60Gln Val Gln Leu Gln Gln Ser Gly Pro Gln Leu Val Arg Pro Gly Ala1
5 10 15Ser Val Lys Ile Ser Cys Lys Thr Ser Gly Tyr Ser Phe Thr Asn
Tyr 20 25 30Trp Met Gln Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Met Ile Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn
Gln Lys Phe 50 55 60Lys Asp Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Pro Thr Phe Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Gly Ser Pro Met Ile Thr
Ser Phe Ala Tyr Trp Gly Gln 100 105 110Gly Thr Leu Val Thr Val Ser
Ala 115 12061117PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 61Gln Val Gln Leu Gln Gln Pro Gly
Ala Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Met Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25 30Trp Ile Ala Trp Val Lys
Arg Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asp Phe Tyr Pro
Gly Asn Thr Ser Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys Thr Lys Ala
Thr Leu Thr Ile Asp Thr Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln
Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala
Arg Arg Gly Arg Gly Gly Ser Ser Tyr Trp Gly Gln Gly Thr Thr 100 105
110Leu Thr Val Ser Ser 11562117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 62Gln Ile Gln Leu Gln Gln
Ser Gly Pro Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser
Cys Lys Val Ser Gly Tyr Ile Phe Thr Asp Phe 20 25 30Tyr Ile Asn Trp
Val Lys Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Trp Ile
Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly
Lys Ala Thr Leu Thr Val Asp Thr Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Phe Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ala 11563117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
63Gln Val Gln Leu Gln Gln Pro Gly Ala Glu Leu Val Arg Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn
Tyr 20 25 30Trp Met Asn Trp Phe Lys Gln Arg Pro Glu Gln Gly Leu Glu
Trp Ile 35 40 45Gly Arg Ile Asp Pro Tyr Asp Ser Glu Thr Leu Tyr Asn
Gln Lys Phe 50 55 60Lys Asp Lys Ala Ile Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Gly Gly Gly Lys Gly Gly Phe Gly
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ala
11564119PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 64Glu Val Gln Leu Gln Gln Ser Gly Ala Glu Leu
Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Thr Ala Ser Gly
Phe Thr Ile Lys Asp Thr 20 25 30Tyr Met His Trp Val Lys Gln Arg Pro
Glu Gln Gly Leu Glu Trp Ile 35 40 45Gly Arg Ile Asp Pro Ala Lys Asp
Asn Thr Lys Tyr Asp Pro Lys Phe 50 55 60Gln Gly Lys Ala Thr Ile Thr
Leu Asp Thr Ser Ser Asn Ile Ala Tyr65 70 75 80Leu Gln Leu Ser Ser
Leu Thr Ser Glu Asp Thr Ala Val Tyr Phe Cys 85 90 95Ala Arg Gly His
Tyr Gly Ser Ser Tyr Val Val Tyr Trp Gly Gln Gly 100 105 110Thr Leu
Val Thr Val Ser Ala 11565112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 65Asp Val Leu Met Thr Gln
Thr Pro Leu Ser Leu Pro Val Ser Leu Gly1 5 10 15Asp Gln Ala Ser Ile
Ser Cys Arg Ser Ser Gln Ser Ile Val Tyr Ser 20 25 30Asn Gly Asn Thr
Tyr Leu Glu Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Lys Leu
Leu Ile Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55 60Asp Arg
Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75
80Ser Arg Val Glu Ala Glu Asp Leu Gly Val Tyr Tyr Cys Phe Gln Gly
85 90 95Ser His Val Pro Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile
Lys 100 105 11066112PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 66Asp Val Leu Met Thr Gln Thr Pro
Leu Ser Leu Pro Val Ser Leu Gly1 5 10 15Glu Gln Ala Ser Ile Ser Cys
Arg Ser Ser Gln Ser Ile Val His Ser 20 25 30Asn Gly Asn Thr Tyr Leu
Glu Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Lys Leu Leu Ile
Tyr Arg Val Ala Asn Arg Phe Ser Gly Val Pro 50 55 60Asp Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg
Val Glu Ala Glu Asp Leu Gly Val Tyr Tyr Cys Phe Gln Gly 85 90 95Ser
His Val Pro Trp Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys 100 105
11067112PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 67Asp Val Leu Met Thr Gln Thr Pro Leu Ser Leu
Pro Val Ser Leu Gly1 5 10 15Glu Gln Ala Ser Ile Ser Cys Arg Ser Ser
Gln Ser Ile Val His Ser 20 25 30Asn Gly Asn Thr Tyr Leu Glu Trp Tyr
Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Lys Leu Leu Ile Tyr Arg Val
Ala Asn Arg Phe Ser Gly Val Pro 50 55 60Asp Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu Ala
Glu Asp Leu Gly Val Tyr Tyr Cys Phe Gln Gly 85 90 95Ser His Val Pro
Trp Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys 100 105
11068107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 68Asp Val Gln Ile Thr Gln Ser Pro Ser Phe Leu
Ala Ala Ser Pro Gly1 5 10 15Glu Thr Ile Thr Ile Asn Cys Arg Ala Ser
Lys Asn Ile Gly Lys Tyr 20 25 30Leu Ala Trp Phe Gln Glu Lys Pro Gly
Lys Thr Asn Lys Leu Leu Ile 35 40 45Tyr Ser Gly Ser Thr Leu Gln Ser
Gly Ile Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Ser Leu Glu Pro65 70 75 80Glu Asp Phe Ala Ile
Tyr Tyr Cys Gln Gln His Asn Glu Tyr Pro Tyr 85 90 95Thr Phe Gly Gly
Gly Thr Lys Leu Glu Ile Lys 100 10569107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
69Asn Ile Leu Met Thr Gln Ser Pro Lys Ser Met Ser Met Ser Val Gly1
5 10 15Glu Arg Val Thr Leu Ser Cys Lys Ala Ser Glu Asn Val Gly Thr
Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Asp Gln Ser Pro Lys Leu
Leu Ile 35 40 45Tyr Gly Ala Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg
Phe Thr Gly 50 55 60Ser Gly Ser Ala Thr Asp Phe Thr Leu Thr Ile Ser
Ser Val Gln Ala65 70 75 80Glu Asp Leu Ala Asp Tyr His Cys Gly Gln
Ser Tyr Ser Tyr Pro Leu 85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu
Leu Lys 100 10570117PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 70Gln Ile Gln Leu Gln Gln Ser Gly
Pro Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser Cys Lys
Val Ser Gly Tyr Ile Phe Thr Asp Phe 20 25 30Tyr Ile Asn Trp Val Lys
Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Trp Ile Tyr Leu
Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Lys Ala
Thr Leu Thr Val Asp Thr Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln
Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Phe Cys 85 90 95Ala
Arg Arg Glu Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105
110Val Thr Val Ser Ala 11571106PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 71Gln Ile Val Leu Thr Gln
Ser Pro Ala Ile Met Ser Ala Ser Pro Gly1 5 10 15Glu Lys Val Thr Met
Thr Cys Ser Ala Ser Ser Ser Val Ser Tyr Met 20 25 30Asn Trp Tyr Gln
Gln Lys Ser Gly Thr Ser Pro Lys Arg Trp Ile Tyr 35 40 45Asp Thr Ser
Lys Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly Ser 50 55 60Gly Ser
Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Ala Glu65 70 75
80Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Trp Asn Ser Asn Pro Pro Thr
85 90
95Phe Gly Thr Gly Thr Lys Leu Glu Leu Lys 100 10572117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
72Gln Val Gln Leu Gln Gln Pro Gly Ala Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Met Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn
Tyr 20 25 30Trp Ile Ala Trp Val Lys Arg Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asp Phe Tyr Pro Gly Asn Thr Ser Thr Asn Tyr Asn
Glu Lys Phe 50 55 60Lys Thr Lys Ala Thr Leu Thr Ile Asp Thr Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Gly Arg Gly Gly Ser Ser
Tyr Trp Gly Gln Gly Thr Thr 100 105 110Leu Thr Val Ser Ser
11573117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 73Gln Val Gln Leu Gln Gln Pro Gly Ala Glu Leu
Val Arg Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly
Tyr Thr Phe Thr Asn Tyr 20 25 30Trp Met Asn Trp Phe Lys Gln Arg Pro
Glu Gln Gly Leu Glu Trp Ile 35 40 45Gly Arg Ile Asp Pro Tyr Asp Ser
Glu Thr Leu Tyr Asn Gln Lys Phe 50 55 60Lys Asp Lys Ala Ile Leu Thr
Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser
Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala Gly Gly Gly
Lys Gly Gly Phe Gly Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr
Val Ser Ala 11574120PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 74Gln Val Gln Leu Gln Gln Ser Gly
Pro Gln Leu Val Arg Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser Cys Lys
Thr Ser Gly Tyr Ser Phe Thr Asn Tyr 20 25 30Trp Met Gln Trp Val Lys
Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Met Ile Asp Pro
Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys Asp Lys Ala
Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln
Leu Ser Ser Pro Thr Phe Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala
Arg Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly Gln 100 105
110Gly Thr Leu Val Thr Val Ser Ala 115 12075117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
75Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Phe 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
11576117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 76Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val
Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly
Tyr Thr Phe Thr Asp Phe 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro
Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly
Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr
Val Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser
Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu
Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr
Val Ser Ser 11577120PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 77Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25 30Trp Met Gln Trp Val Arg
Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Met Ile Asp Pro
Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys Asp Arg Val
Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65 70 75 80Met Glu
Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala
Arg Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly Gln 100 105
110Gly Thr Leu Val Thr Val Ser Ser 115 12078120PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
78Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn
Tyr 20 25 30Trp Met Gln Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Met Ile Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn
Gln Lys Phe 50 55 60Lys Asp Arg Val Thr Met Thr Val Asp Thr Ser Thr
Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Gly Ser Pro Met Ile Thr
Ser Phe Ala Tyr Trp Gly Gln 100 105 110Gly Thr Leu Val Thr Val Ser
Ser 115 12079120PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 79Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys
Thr Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25 30Trp Met Gln Trp Val Arg
Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Met Ile Asp Pro
Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys Asp Arg Val
Thr Met Thr Val Asp Thr Ser Thr Ser Thr Val Tyr65 70 75 80Met Glu
Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala
Arg Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly Gln 100 105
110Gly Thr Leu Val Thr Val Ser Ser 115 12080117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
80Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Val Ser Gly Tyr Thr Phe Thr Asp
Phe 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Val Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
11581293PRTHomo sapiens 81Met Trp Pro Leu Val Ala Ala Leu Leu Leu
Gly Ser Ala Cys Cys Gly1 5 10 15Ser Ala Gln Leu Leu Phe Asn Lys Thr
Lys Ser Val Glu Phe Thr Phe 20 25 30Cys Asn Asp Thr Val Val Ile Pro
Cys Phe Val Thr Asn Met Glu Ala 35 40 45Gln Asn Thr Thr Glu Val Tyr
Val Lys Trp Lys Phe Lys Gly Arg Asp 50 55 60Ile Tyr Thr Phe Asp Gly
Ala Leu Asn Lys Ser Thr Val Pro Thr Asp65 70 75 80Phe Ser Ser Ala
Lys Ile Glu Val Ser Gln Leu Leu Lys Gly Asp Ala 85 90 95Ser Leu Lys
Met Asp Lys Ser Asp Ala Val Ser His Thr Gly Asn Tyr 100 105 110Thr
Cys Glu Val Thr Glu Leu Thr Arg Glu Gly Glu Thr Ile Ile Glu 115 120
125Leu Lys Tyr Arg Val Val Ser Trp Phe Ser Pro Asn Glu Asn Ile Leu
130 135 140Ile Val Ile Phe Pro Ile Phe Ala Ile Leu Leu Phe Trp Gly
Gln Phe145 150 155 160Gly Ile Lys Thr Leu Lys Tyr Arg Ser Gly Gly
Met Asp Glu Lys Thr 165 170 175Ile Ala Leu Leu Val Ala Gly Leu Val
Ile Thr Val Ile Val Ile Val 180 185 190Gly Ala Ile Leu Phe Val Pro
Gly Glu Tyr Ser Leu Lys Asn Ala Thr 195 200 205Gly Leu Gly Leu Ile
Val Thr Ser Thr Gly Ile Leu Ile Leu Leu His 210 215 220Tyr Tyr Val
Phe Ser Thr Ala Ile Gly Leu Thr Ser Phe Val Ile Ala225 230 235
240Ile Leu Val Ile Gln Val Ile Ala Tyr Ile Leu Ala Val Val Gly Leu
245 250 255Ser Leu Cys Ile Ala Ala Cys Ile Pro Met His Gly Pro Leu
Leu Ile 260 265 270Ser Gly Leu Ser Ile Leu Ala Leu Ala Gln Leu Leu
Gly Leu Val Tyr 275 280 285Met Lys Phe Val Glu 29082324PRTMus
musculus 82Met Trp Pro Leu Ala Ala Ala Leu Leu Leu Gly Ser Cys Cys
Cys Gly1 5 10 15Ser Ala Gln Leu Leu Phe Ser Asn Val Asn Ser Ile Glu
Phe Thr Ser 20 25 30Cys Asn Glu Thr Val Val Ile Pro Cys Ile Val Arg
Asn Val Glu Ala 35 40 45Gln Ser Thr Glu Glu Met Phe Val Lys Trp Lys
Leu Asn Lys Ser Tyr 50 55 60Ile Phe Ile Tyr Asp Gly Asn Lys Asn Ser
Thr Thr Thr Asp Gln Asn65 70 75 80Phe Thr Ser Ala Lys Ile Ser Val
Ser Asp Leu Ile Asn Gly Ile Ala 85 90 95Ser Leu Lys Met Asp Lys Arg
Asp Ala Met Val Gly Asn Tyr Thr Cys 100 105 110Glu Val Thr Glu Leu
Ser Arg Glu Gly Lys Thr Val Ile Glu Leu Lys 115 120 125Asn Arg Thr
Ala Phe Asn Thr Asp Gln Gly Ser Ala Cys Ser Tyr Glu 130 135 140Glu
Glu Lys Gly Gly Cys Lys Leu Val Ser Trp Phe Ser Pro Asn Glu145 150
155 160Lys Ile Leu Ile Val Ile Phe Pro Ile Leu Ala Ile Leu Leu Phe
Trp 165 170 175Gly Lys Phe Gly Ile Leu Thr Leu Lys Tyr Lys Ser Ser
His Thr Asn 180 185 190Lys Arg Ile Ile Leu Leu Leu Val Ala Gly Leu
Val Leu Thr Val Ile 195 200 205Val Val Val Gly Ala Ile Leu Leu Ile
Pro Gly Glu Lys Pro Val Lys 210 215 220Asn Ala Ser Gly Leu Gly Leu
Ile Val Ile Ser Thr Gly Ile Leu Ile225 230 235 240Leu Leu Gln Tyr
Asn Val Phe Met Thr Ala Phe Gly Met Thr Ser Phe 245 250 255Thr Ile
Ala Ile Leu Ile Thr Gln Val Leu Gly Tyr Val Leu Ala Leu 260 265
270Val Gly Leu Cys Leu Cys Ile Met Ala Cys Glu Pro Val His Gly Pro
275 280 285Leu Leu Ile Ser Gly Leu Gly Ile Ile Ala Leu Ala Glu Leu
Leu Gly 290 295 300Leu Val Tyr Met Lys Phe Val Ala Ser Asn Gln Arg
Thr Ile Gln Pro305 310 315 320Pro Arg Asn Arg83305PRTHomo sapiens
83Met Trp Pro Leu Val Ala Ala Leu Leu Leu Gly Ser Ala Cys Cys Gly1
5 10 15Ser Ala Gln Leu Leu Phe Asn Lys Thr Lys Ser Val Glu Phe Thr
Phe 20 25 30Cys Asn Asp Thr Val Val Ile Pro Cys Phe Val Thr Asn Met
Glu Ala 35 40 45Gln Asn Thr Thr Glu Val Tyr Val Lys Trp Lys Phe Lys
Gly Arg Asp 50 55 60Ile Tyr Thr Phe Asp Gly Ala Leu Asn Lys Ser Thr
Val Pro Thr Asp65 70 75 80Phe Ser Ser Ala Lys Ile Glu Val Ser Gln
Leu Leu Lys Gly Asp Ala 85 90 95Ser Leu Lys Met Asp Lys Ser Asp Ala
Val Ser His Thr Gly Asn Tyr 100 105 110Thr Cys Glu Val Thr Glu Leu
Thr Arg Glu Gly Glu Thr Ile Ile Glu 115 120 125Leu Lys Tyr Arg Val
Val Ser Trp Phe Ser Pro Asn Glu Asn Ile Leu 130 135 140Ile Val Ile
Phe Pro Ile Phe Ala Ile Leu Leu Phe Trp Gly Gln Phe145 150 155
160Gly Ile Lys Thr Leu Lys Tyr Arg Ser Gly Gly Met Asp Glu Lys Thr
165 170 175Ile Ala Leu Leu Val Ala Gly Leu Val Ile Thr Val Ile Val
Ile Val 180 185 190Gly Ala Ile Leu Phe Val Pro Gly Glu Tyr Ser Leu
Lys Asn Ala Thr 195 200 205Gly Leu Gly Leu Ile Val Thr Ser Thr Gly
Ile Leu Ile Leu Leu His 210 215 220Tyr Tyr Val Phe Ser Thr Ala Ile
Gly Leu Thr Ser Phe Val Ile Ala225 230 235 240Ile Leu Val Ile Gln
Val Ile Ala Tyr Ile Leu Ala Val Val Gly Leu 245 250 255Ser Leu Cys
Ile Ala Ala Cys Ile Pro Met His Gly Pro Leu Leu Ile 260 265 270Ser
Gly Leu Ser Ile Leu Ala Leu Ala Gln Leu Leu Gly Leu Val Tyr 275 280
285Met Lys Phe Val Ala Ser Asn Gln Lys Thr Ile Gln Pro Pro Arg Asn
290 295 300Asn30584303PRTMus musculus 84Met Trp Pro Leu Ala Ala Ala
Leu Leu Leu Gly Ser Cys Cys Cys Gly1 5 10 15Ser Ala Gln Leu Leu Phe
Ser Asn Val Asn Ser Ile Glu Phe Thr Ser 20 25 30Cys Asn Glu Thr Val
Val Ile Pro Cys Ile Val Arg Asn Val Glu Ala 35 40 45Gln Ser Thr Glu
Glu Met Phe Val Lys Trp Lys Leu Asn Lys Ser Tyr 50 55 60Ile Phe Ile
Tyr Asp Gly Asn Lys Asn Ser Thr Thr Thr Asp Gln Asn65 70 75 80Phe
Thr Ser Ala Lys Ile Ser Val Ser Asp Leu Ile Asn Gly Ile Ala 85 90
95Ser Leu Lys Met Asp Lys Arg Asp Ala Met Val Gly Asn Tyr Thr Cys
100 105 110Glu Val Thr Glu Leu Ser Arg Glu Gly Lys Thr Val Ile Glu
Leu Lys 115 120 125Asn Arg Thr Val Ser Trp Phe Ser Pro Asn Glu Lys
Ile Leu Ile Val 130 135 140Ile Phe Pro Ile Leu Ala Ile Leu Leu Phe
Trp Gly Lys Phe Gly Ile145 150 155 160Leu Thr Leu Lys Tyr Lys Ser
Ser His Thr Asn Lys Arg Ile Ile Leu 165 170 175Leu Leu Val Ala Gly
Leu Val Leu Thr Val Ile Val Val Val Gly Ala 180 185 190Ile Leu Leu
Ile Pro Gly Glu Lys Pro Val Lys Asn Ala Ser Gly Leu 195 200 205Gly
Leu Ile Val Val Ser Thr Gly Ile Leu Ile Leu Leu Gln Tyr Asn 210 215
220Val Phe Met Thr Ala Phe Gly Met Thr Ser Phe Thr Ile Ala Ile
Leu225 230 235 240Ile Thr Gln Val Leu Gly Tyr Val Leu Ala Leu Val
Gly Leu Cys Leu 245 250 255Cys Ile Met Ala Cys Glu Pro Val His Gly
Pro Leu Leu Ile Ser Gly 260 265 270Leu Gly Ile Ile Ala Leu Ala Glu
Leu Leu Gly Leu Val Tyr Met Lys 275 280 285Phe Val Ala Ser Asn Gln
Arg Thr Ile Gln Pro Pro Arg Asn Arg 290 295 30085107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
85Asp Ile Gln Met Thr Gln Ser Pro Ala Ser Leu Ser Ala Ser Val Gly1
5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser Glu Asn Ile Tyr Ser
Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly Lys Ser Pro Gln Leu
Leu Val 35 40
45Tyr Asn Ala Lys Thr Leu Ala Glu Gly Val Pro Ser Arg Phe Ser Gly
50 55 60Ser Gly Ser Gly Thr Gln Phe Ser Leu Lys Ile Lys Ser Leu Gln
Pro65 70 75 80Glu Asp Phe Gly Ser Tyr Tyr Cys Gln His His Tyr Gly
Thr Pro Leu 85 90 95Thr Phe Gly Gly Gly Thr Lys Leu Glu Leu Lys 100
10586115PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 86Glu Val Gln Leu Gln Gln Ser Gly Thr Glu Leu
Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly
Tyr Thr Phe Ile Ser Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro
Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly
Asn Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys Thr Lys Ala Thr Leu Thr
Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser
Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp Tyr
Tyr Gly Asn Tyr Trp Gly Gln Gly Thr Thr Val Thr 100 105 110Val Ser
Ser 11587107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 87Asp Ile Lys Met Asn Gln Ser Pro
Ser Ser Leu Ser Ala Ser Leu Gly1 5 10 15Asp Thr Ile Thr Ile Thr Cys
His Ala Ser Gln Asn Ile Asn Val Trp 20 25 30Leu Ser Trp Tyr Gln Gln
Lys Pro Gly Asn Ile Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala Ser Asn
Leu His Thr Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Thr Gly Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp
Ile Ala Thr Tyr Tyr Cys Gln Gln Gly Gln Ser Tyr Pro Leu 85 90 95Thr
Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys 100 10588118PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
88Glu Val Gln Leu Gln Gln Ser Gly Pro Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Phe Thr Phe Thr Asn
Tyr 20 25 30Tyr Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Pro Glu
Trp Ile 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Lys Ala Thr Leu Thr Ala Asp Thr Ser Ser
Ser Thr Ala Tyr65 70 75 80Leu Gln Leu Ser Thr Leu Ile Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Tyr Asp Tyr Asp Leu Tyr Leu
Asp Ser Trp Gly Gln Gly Thr 100 105 110Thr Val Thr Val Ser Ser
11589107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 89Asp Ile Gln Met Met Gln Ser Pro Ala Ser Leu
Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser
Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly
Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala Glu
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe
Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Asn
Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Ala Phe Gly Ala
Gly Thr Lys Leu Glu Ile Lys 100 10590115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
90Glu Val Gln Leu Gln Gln Pro Gly Ala Glu Leu Val Lys Pro Gly Thr1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Ala
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Thr Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
11591107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 91Asp Ile Gln Met Thr Gln Ser Pro Ala Ser Leu
Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser
Glu Asn Val Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly
Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Lys Ala Asn Thr Leu Ala Glu
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe
Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Thr
Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly Ser
Gly Thr Lys Leu Glu Ile Lys 100 10592115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
92Glu Val Gln Leu Gln Gln Pro Gly Ala Glu Leu Val Arg Pro Gly Ser1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Leu Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
11593108PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 93Asp Ile Gln Met Ile Gln Ser Pro Ala Ser Leu
Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser
Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly
Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala Glu
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe
Ser Leu Arg Ile Asn Thr Leu Gln Pro65 70 75 80Glu Asp Leu Gly Thr
Tyr Tyr Cys Gln His His Tyr Gly Ala Pro Leu 85 90 95Ser Phe Gly Gly
Gly Thr Lys Leu Glu Ile Lys Arg 100 10594115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
94Glu Val Gln Leu Gln Gln Pro Gly Ala Glu Leu Val Arg Pro Gly Thr1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
11595107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 95Asp Ile Gln Met Thr Gln Ser Pro Ala Ser Leu
Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser
Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly
Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala Glu
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe
Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Ser
Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly Ala
Gly Thr Lys Leu Glu Ile Lys 100 10596115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
96Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ala His Tyr Ser
Glu Glu Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
11597107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 97Asp Ile Gln Ile Asn Gln Ser Pro Ser Ser Leu
Ser Ala Ser Leu Gly1 5 10 15Asp Thr Ile Thr Ile Thr Cys His Ala Ser
Gln Asn Ile Asn Val Trp 20 25 30Leu Ser Trp Tyr Gln Gln Lys Pro Gly
Asn Ile Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala Ser Asn Leu His Thr
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gly Phe
Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Ile Ala Thr
Tyr Tyr Cys Gln Gln Gly Gln Ser Tyr Pro Leu 85 90 95Thr Phe Gly Ala
Gly Thr Lys Leu Glu Ile Lys 100 10598118PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
98Glu Val Gln Leu Gln Gln Ser Gly Pro Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Ile Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Asn
Tyr 20 25 30His Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Trp Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Met Ala Ile Leu Thr Ala Asp Thr Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Tyr Asp Tyr Asp Leu Tyr Leu
His Ser Trp Gly Gln Gly Thr 100 105 110Thr Val Thr Val Ser Ser
11599107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 99Asp Ile Gln Met Thr Gln Ser Pro Ala Ser Leu
Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser
Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly
Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala Glu
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe
Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Ser
Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly Ala
Gly Thr Lys Leu Glu Ile Lys 100 105100115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
100Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Val Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115101106PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 101Arg His Cys Phe Asn Gln Ser Pro Ala Ile
Met Ser Ala Ser Pro Gly1 5 10 15Glu Lys Val Thr Met Thr Cys Ser Ala
Ser Ser Ser Val Ser Tyr Met 20 25 30His Trp Tyr Gln Gln Lys Ser Gly
Thr Ser Pro Lys Arg Trp Ile Tyr 35 40 45Asp Thr Ser Asn Leu Ala Ser
Gly Val Pro Val Arg Phe Ser Gly Ser 50 55 60Gly Ser Gly Thr Ser Tyr
Ser Leu Thr Ile Ser Arg Met Glu Ala Glu65 70 75 80Asp Ala Ala Thr
Tyr Tyr Cys Gln Gln Trp Ser Asn Tyr Pro Phe Thr 85 90 95Phe Gly Ser
Gly Thr Lys Leu Glu Leu Lys 100 105102119PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
102Arg Val Gln Leu Gln Gln Ser Gly Ala Glu Leu Val Arg Pro Gly Thr1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Ala Tyr Ala Phe Thr Asn
Tyr 20 25 30Leu Ile Glu Trp Val Lys Lys Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Val Ile Asn Pro Gly Ser Gly Gly Thr Asn Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Lys Ala Thr Leu Thr Ala Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Phe Cys 85 90 95Ala Arg Gly Asp Gly Tyr Gly Ser Leu
Phe Ala Tyr Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val Ser Ala
115103107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 103Asp Ile Gln Met Ile Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Gly Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Ile Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Ile Lys 100 105104115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
104Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ile Ser
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asn Thr Asn Tyr Asn
Glu Lys Phe 50 55 60Lys Thr Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp Tyr Tyr Gly Asn Tyr Trp
Gly Gln Gly Thr Ser Val Thr 100 105 110Val Ser Ser
115105107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 105Asp Ile Gln Met Ile Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Leu
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala
Glu Gly
Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr His Phe Ser
Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Thr Tyr
Tyr Cys Gln His His Tyr Gly Asn Ser Leu 85 90 95Thr Phe Gly Ala Gly
Thr Lys Leu Glu Leu Lys 100 105106115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
106Glu Val Gln Leu Gln Gln Ser Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Pro Phe Thr Ser
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Gly Thr Asn Tyr Asn
Glu Lys Phe 50 55 60Lys Thr Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Asn Tyr Trp
Gly Gln Gly Thr Thr Val Thr 100 105 110Val Ser Ser
115107107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 107Asp Ile Gln Met Thr Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Ile Lys 100 105108115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
108Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Thr Ser Gly Tyr Ser Phe Val Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Ser
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Lys Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115109109PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 109Asp Ala Val Val Thr Gln Glu Ser Ala Leu
Thr Thr Ser Pro Gly Glu1 5 10 15Thr Val Thr Leu Thr Cys Arg Ser Ser
Thr Gly Ala Val Thr Thr Asn 20 25 30Asn Tyr Ala Asn Trp Val Gln Glu
Lys Pro Asp His Leu Phe Thr Gly 35 40 45Leu Ile Gly Gly Thr Asn Asn
Arg Ala Pro Gly Val Pro Ala Arg Phe 50 55 60Ser Gly Ser Leu Ile Gly
Asp Lys Ala Ala Leu Thr Ile Thr Gly Ala65 70 75 80Gln Thr Glu Asp
Glu Ala Ile Tyr Phe Cys Ala Leu Trp Tyr Ser Asn 85 90 95His Leu Val
Phe Gly Gly Gly Thr Lys Leu Thr Val Leu 100 105110125PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
110Pro Arg Gly Lys Val Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser1
5 10 15Gln Ser Leu Ser Leu Thr Cys Ser Val Thr Gly Tyr Ser Ile Thr
Ser 20 25 30Gly Tyr Tyr Trp Asn Trp Ile Arg Gln Phe Pro Gly Asn Lys
Leu Glu 35 40 45Trp Met Gly Tyr Ile Asn Tyr Asp Gly Ser Asn Asn Tyr
Asn Pro Ser 50 55 60Leu Lys Asn Arg Ile Ser Ile Thr Arg Asp Thr Ser
Lys Asn Gln Phe65 70 75 80Phe Leu Lys Leu Asn Ser Val Thr Thr Glu
Asp Thr Ala Thr Tyr Tyr 85 90 95Cys Ala Arg Gly Tyr Tyr Tyr Gly Ser
Ser Tyr Gly Tyr Trp Tyr Phe 100 105 110Asp Val Trp Gly Thr Gly Thr
Thr Val Thr Val Ser Ser 115 120 125111107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
111Asp Ile Gln Met Thr Gln Ser Pro Ala Ser Leu Ser Ala Ser Val Gly1
5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser Glu Asn Ile Tyr Ser
Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly Lys Ser Pro Gln Leu
Leu Val 35 40 45Tyr Ala Ala Thr Asn Leu Ala Asp Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Arg Ser Gly Thr Gln Phe Ser Leu Lys Ile Asn
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Ser Tyr Tyr Cys Gln His
His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu
Ile Lys 100 105112115PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 112Glu Val Gln Leu Gln
Gln Pro Gly Thr Glu Leu Ala Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ile Ser Tyr 20 25 30Trp Met His
Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn
Ile Asn Pro Ser Ser Gly Gly Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys
Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys
85 90 95Val Arg Asp Tyr Tyr Gly Ser Tyr Trp Gly Gln Gly Thr Thr Val
Thr 100 105 110Val Ser Ser 115113107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
113Asp Ile Gln Met Thr Gln Ser Pro Ala Ser Leu Ser Ala Ser Val Gly1
5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser Glu Asn Ile Tyr Ser
Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly Lys Ser Pro Gln Leu
Leu Val 35 40 45Tyr Lys Ala Lys Thr Leu Val Glu Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe Ser Leu Lys Ile Asn
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Ser Tyr Tyr Cys Gln His
His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu
Ile Lys 100 105114115PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 114Arg Gly Pro Thr Ala
Ala Thr Trp Thr Glu Leu Val Lys Pro Gly Thr1 5 10 15Ser Val Lys Leu
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ile Ser Tyr 20 25 30Trp Met His
Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn
Ile Asn Pro Ser Ser Gly Asp Thr Ser Tyr Asn Glu Lys Phe 50 55 60Lys
Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp Gly Gln Gly Thr Ser Val
Thr 100 105 110Val Ser Ala 115115107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
115Asp Ile Gln Met Met Gln Ser Pro Ala Ser Leu Ser Ala Ser Val Gly1
5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala Ser Glu Asn Ile Tyr Ser
Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln Gly Lys Ser Pro Gln Leu
Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala Glu Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln Phe Ser Leu Lys Ile Asn
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly Asn Tyr Tyr Cys Gln His
His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly Ser Gly Thr Lys Leu Glu
Leu Lys 100 105116115PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 116Glu Val Gln Leu Gln
Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Trp Met His
Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn
Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Ser Glu Lys Phe 50 55 60Arg
Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp Gly His Gly Thr Leu Val
Thr 100 105 110Val Ser Ala 115117109PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
117Asp Ala Val Val Thr Gln Glu Ser Ala Leu Thr Thr Ser Pro Gly Glu1
5 10 15Thr Val Thr Leu Thr Cys Arg Ser Ser Thr Gly Ala Val Ser Thr
Ser 20 25 30Asn Tyr Ala Asn Trp Val Gln Glu Lys Pro Asp His Leu Phe
Thr Gly 35 40 45Leu Ile Gly Gly Thr Asn Asn Arg Ala Pro Gly Val Pro
Ala Arg Phe 50 55 60Ser Gly Ser Leu Ile Gly Asp Lys Ala Ala Leu Thr
Ile Thr Gly Ala65 70 75 80Gln Thr Glu Asp Glu Ala Ile Tyr Phe Cys
Ala Leu Trp Tyr Ser Asn 85 90 95His Trp Val Phe Gly Gly Gly Thr Lys
Leu Thr Val Leu 100 105118114PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 118Glu Val Gln Leu Gln
Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Trp Met His
Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn
Val Val Pro Asn Asn Asp Gly Thr Asn Tyr Asn Glu Lys Phe 50 55 60Arg
Asn Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys
85 90 95Ala Val Thr Tyr Phe Ala Tyr Trp Gly Gln Gly Thr Leu Val Thr
Val 100 105 110Ser Ala119107PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 119Asp Ile Gln Met Met
Gln Ser Pro Ala Ser Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr
Ile Thr Cys Arg Ala Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp
Tyr Gln Gln Lys Gln Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn
Ala Lys Thr Leu Ala Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser
Gly Ser Gly Thr Gln Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75
80Glu Asp Phe Gly Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu
85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys 100
105120115PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 120Glu Val Gln Leu Gln Gln Ser Gly Ala Glu
Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser
Gly Tyr Ser Phe Thr Thr Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg
Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser
Gly Ser Ala His Tyr Asn Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu
Thr Val Asp Lys Ser Ser Asn Thr Ala Tyr65 70 75 80Met Gln Leu Ser
Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp
Tyr Tyr Gly Ala Tyr Trp Gly His Gly Thr Leu Val Thr 100 105 110Val
Ser Ala 115121107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 121Asp Ile Gln Met Thr Gln Ser Pro
Ala Ser Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys
Arg Ala Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln
Lys Gln Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr
Leu Ala Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Thr His Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp
Phe Gly Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr
Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys 100 105122115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
122Glu Val Gln Leu Gln Gln Pro Gly Ala Glu Leu Val Arg Pro Gly Ser1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ile Ser
Tyr 20 25 30Trp Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Ser Ser Asn Tyr Asn
Glu Lys Phe 50 55 60Lys Asn Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115123107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 123Asp Ile Gln Met Thr Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Asn Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Ile Lys 100 105124115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
124Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Ile Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Ser
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115125107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 125Asp Ile Lys Ile Asn Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Lys Ala
Lys Thr Leu Val Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Gln Phe Ser Leu Lys Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Gly Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu
85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys 100
105126115PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 126Glu Val Gln Leu Gln Gln Pro Gly Ala Glu
Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Ile Ser Tyr 20 25 30Trp Met His Trp Val Arg Gln Arg
Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser
Gly Asn Thr Asn Tyr Asn Glu Lys Phe 50 55 60Lys Thr Lys Ala Thr Leu
Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser
Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp
Tyr Tyr Gly Asn Tyr Trp Gly Gln Gly Thr Ser Val Thr 100 105 110Val
Ser Ser 115127107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 127Gly Ile Gln Met Thr Gln Ser Pro
Ala Ser Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys
Arg Ala Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln
Lys Gln Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Lys Ala Lys Thr
Leu Val Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Thr Gln Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp
Phe Gly Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr
Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys 100 105128115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
128Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ile Ser
Tyr 20 25 30Trp Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Gly Thr Asn Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met His Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Asn Tyr Trp
Gly Gln Gly Thr Thr Leu Thr 100 105 110Val Ser Ser
115129107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 129Asp Ile Gln Met Thr Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Asp Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Ser Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Leu Lys 100 105130115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
130Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115131107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 131Asp Ile Gln Met Thr Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Asn Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Met Lys 100 105132115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
132Glu Val Gln Leu Gln Gln Ser Gly Ala Glu His Val Arg Pro Gly Ser1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Ile Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115133107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 133Asp Ile Lys Met Asn Gln Ser Pro Ser Ser
Leu Ser Ala Ser Leu Gly1 5 10 15Asp Thr Ile Thr Ile Thr Cys His Ala
Ser Gln Asn Ile Asn Val Trp 20 25 30Leu Ser Trp Tyr Gln Gln Lys Pro
Gly Asn Ile Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala Ser Asn Leu His
Thr Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gly
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Ile Ala
Thr Tyr Tyr Cys Gln Gln Gly Gln Ser Tyr Pro Leu 85 90 95Thr Phe Gly
Thr Gly Thr Lys Leu Glu Ile Lys 100 105134118PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
134Glu Val Gln Leu Gln Gln Ser Gly Pro Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Ile Ser Cys Lys Ala Ser Gly Asp Ser Phe Thr Ser
Asp 20 25 30His Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Trp Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Lys Ala Thr Leu Thr Ala Asp Thr Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Arg Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Val Thr Tyr Asp Tyr Asp Leu Tyr Phe
Asp Asn Trp Gly Gln Gly Thr 100 105 110Thr Leu Thr Val Ser Ser
115135111PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 135Ser Thr Leu Met Thr Gln Ser Pro Ala Ser
Leu Ala Val Ser Leu Gly1 5 10 15Gln Arg Ala Thr Ile Ser Cys Arg Ala
Ser Glu Ser Val Asp Ser Tyr 20 25 30Gly Asn Ser Phe Met His Trp Tyr
Gln Gln Lys Pro Gly Gln Pro Pro 35 40 45Lys Leu Leu Ile Tyr Leu Ala
Ser Asn Leu Glu Ser Gly Val Pro Ala 50 55 60Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Phe Ile Leu Asn Ile His65 70 75 80Pro Val Glu Glu
Glu Asp Ala Ala Thr Tyr Tyr Cys Leu His Ser Arg 85 90 95Glu Leu Pro
Phe Thr Phe Gly Ser Gly Thr Lys Leu Glu Leu Lys 100 105
110136119PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 136Glu Val Lys Leu Met Glu Ser Gly Gly Gly
Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Lys Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Asp Tyr 20 25 30Gly Met His Trp Val Arg Gln Ala
Pro Glu Lys Gly Leu Glu Trp Val 35 40 45Ala Tyr Ile Ser Ser Gly Ser
Ser Thr Ile Tyr Tyr Ala Asp Thr Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Thr Leu Phe65 70 75 80Leu Gln Met Thr
Ser Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys 85 90 95Ala Arg Pro
Asp Asp Gly Tyr Tyr Gly Phe Ala Tyr Trp Gly Gln Gly 100 105 110Thr
Leu Val Thr Val Ser Ala 115137107PRTArtificial SequenceDescription
of Artificial Sequence Synthetic polypeptide 137Asp Ile Gln Met Thr
Gln Ser Pro Ala Ser Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr
Ile Thr Cys Arg Ala Ser Glu Asn Val Tyr Ser Tyr 20 25 30Leu Ala Trp
Tyr Gln Gln Lys Gln Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn
Ala Lys Thr Leu Ala Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser
Gly Ser Gly Thr Gln Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75
80Glu Asp Phe Gly Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu
85 90 95Thr Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys 100
105138115PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 138Glu Val Gln Leu Gln Gln Pro Gly Thr Glu
Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser
Gly Tyr Ser Phe Thr Thr Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg
Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser
Gly Asp Ser His Tyr Asn Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu
Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met His Leu Ser
Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp
Tyr Tyr Gly Ala Tyr Trp Gly His Gly Thr Leu Val Thr 100 105 110Val
Ser Ala 115139107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 139Asp Ile Gln Met Thr Gln Ser Pro
Ala Ser Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys
Arg Ala Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln
Lys Gln Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Phe Asn Ala Lys Thr
Leu Ala Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Thr His Phe Ser Leu Lys Ile Asn Asn Leu Gln Pro65 70 75 80Glu Asp
Phe Gly Thr Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr
Phe Gly Ala Gly Thr Lys Leu Glu Ile Lys 100 105140115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
140Glu Val Gln Leu Gln Gln Ser Gly Ala Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Ile Ser
Tyr 20 25 30Trp Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Gly Asn Thr Tyr Asn
Glu Lys Phe 50 55 60Lys Asn Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Arg Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Thr Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115141107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 141Asp Ile Gln Met Thr Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Ala Phe Gly
Ser Gly Thr Lys Leu Glu Leu Lys 100 105142115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
142Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Ile Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Gly Gly Asp Ser His Tyr Ser
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115143107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 143Asp Ile Gln Met Ile Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Phe Asn Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Ile Lys 100 105144115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
144Glu Val Gln Leu Gln Gln Ser Gly Ala Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly Tyr Ser Phe Ile Ser
Tyr 20 25 30Trp Ile His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Gly Ser Ser Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Thr Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ala
115145106PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 145Asp Ile Val Leu Ser Gln Ser Pro Ala Ile
Met Ser Ala Ser Pro Gly1 5 10 15Glu Lys Val Thr Met Thr Cys Ser Ala
Ser Ser Ser Val Ser Tyr Met 20 25 30His Trp Tyr Gln Gln Lys Ser Gly
Thr Ser Pro Lys Arg Trp Ile Tyr 35 40
45Asp Thr Ser Lys Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60Gly Ser Gly Thr Ser Tyr Ser Leu Thr Leu Ser Ser Met Ala Ala
Glu65 70 75 80Asp Ala Ala Thr Tyr Ser Cys Gln Gln Trp Ser Gly Asn
Ser Pro Thr 85 90 95Phe Gly Ala Gly Thr Lys Leu Glu Ile Lys 100
105146124PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 146Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Lys Gly1 5 10 15Ser Leu Lys Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Asn Thr Tyr 20 25 30Ala Met His Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Arg Ile Arg Ser Lys Ser
Ser Asn Tyr Ala Thr Tyr Tyr Ala Asp 50 55 60Ser Val Lys Asp Arg Phe
Thr Ile Ser Arg Asp Asp Ser Gln Ser Ile65 70 75 80Val Tyr Leu Gln
Met Asn Asn Leu Lys Thr Glu Asp Thr Ala Met Tyr 85 90 95Tyr Cys Val
Arg Ala Trp Asp Tyr Gly Ser Ser Trp Asp Tyr Phe Asp 100 105 110Tyr
Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser 115
120147107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 147Asp Ile Gln Met Met Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Gln Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Ser65 70 75 80Glu Asp Ile Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Leu Lys 100 105148115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
148Arg Gly Pro Thr Gln Gln Pro Gly Thr Glu Leu Val Lys Pro Gly Ala1
5 10 15Ser Val Lys Leu Ser Arg Lys Ala Ser Gly Tyr Thr Phe Thr Thr
Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly Asp Ser His Tyr Asn
Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr Val Asp Lys Ser Ser
Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser Leu Thr Ser Glu Asp
Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ala Tyr Trp
Gly His Gly Thr Leu Val Thr 100 105 110Val Ser Ala
1151495PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 149Gly Tyr Gly Met Ser1 515017PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 150Thr
Ile Thr Ser Gly Gly Thr Tyr Thr Tyr Tyr Pro Asp Ser Val Lys1 5 10
15Gly1519PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 151Ser Leu Ala Gly Asn Ala Met Asp Tyr1
515211PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 152Arg Ala Ser Gln Thr Ile Ser Asp Tyr Leu His1 5
101537PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 153Phe Ala Ser Gln Ser Ile Ser1
51549PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 154Gln Asn Gly His Gly Phe Pro Arg Thr1
51555PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 155Asn Tyr Asn Met His1 515617PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 156Thr
Ile Tyr Pro Gly Asn Asp Asp Thr Ser Tyr Asn Gln Lys Phe Lys1 5 10
15Asp1578PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 157Gly Gly Tyr Arg Ala Met Asp Tyr1
515816PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 158Arg Ser Ser Gln Ser Ile Val Tyr Ser Asn Gly
Asn Thr Tyr Leu Gly1 5 10 151597PRTArtificial SequenceDescription
of Artificial Sequence Synthetic peptide 159Lys Val Ser Asn Arg Phe
Ser1 51609PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 160Phe Gln Gly Ser His Val Pro Tyr Thr1
5161139PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 161Met Trp Pro Leu Val Ala Ala Leu Leu Leu
Gly Ser Ala Cys Cys Gly1 5 10 15Ser Ala Gln Leu Leu Phe Asn Lys Thr
Lys Ser Val Glu Phe Thr Phe 20 25 30Cys Asn Asp Thr Val Val Ile Pro
Cys Phe Val Thr Asn Met Glu Ala 35 40 45Gln Asn Thr Thr Glu Val Tyr
Val Lys Trp Lys Phe Lys Gly Arg Asp 50 55 60Ile Tyr Thr Phe Asp Gly
Ala Leu Asn Lys Ser Thr Val Pro Thr Asp65 70 75 80Phe Ser Ser Ala
Lys Ile Glu Val Ser Gln Leu Leu Lys Gly Asp Ala 85 90 95Ser Leu Lys
Met Asp Lys Ser Asp Ala Val Ser His Thr Gly Asn Tyr 100 105 110Thr
Cys Glu Val Thr Glu Leu Thr Arg Glu Gly Glu Thr Ile Ile Glu 115 120
125Leu Lys Tyr Arg Val Val Ser Trp Phe Ser Pro 130
1351626PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 162Asp Asn Ile Tyr Ser Tyr1 51636PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 163Glu
Asn Ile Tyr Ser Tyr1 51646PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 164Glu Asn Val Tyr Ser Tyr1
516510PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 165Glu Ser Val Asp Ser Tyr Gly Asn Ser Phe1 5
101666PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 166Gln Asn Ile Asn Val Trp1 51675PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 167Ser
Ser Val Ser Tyr1 51689PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 168Thr Gly Ala Val Ser Thr
Ser Asn Tyr1 51699PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 169Thr Gly Ala Val Thr Thr Asn Asn Tyr1
51703PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 170Ala Ala Thr11713PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 171Asp
Thr Ser11723PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 172Gly Ala Lys11733PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 173Gly
Thr Asn11743PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 174Lys Ala Lys11753PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 175Lys
Ala Asn11763PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 176Lys Ala Ser11773PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 177Leu
Ala Ser11783PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 178Asn Ala Lys11793PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 179Asn
Ala Gln11809PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 180Ala Leu Trp Tyr Ser Asn His Leu Val1
51819PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 181Ala Leu Trp Tyr Ser Asn His Trp Val1
51829PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 182Leu His Ser Arg Glu Leu Pro Phe Thr1
51839PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 183Gln His His Tyr Gly Ala Pro Leu Ser1
51849PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 184Gln His His Tyr Gly Ile Pro Leu Thr1
51859PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 185Gln His His Tyr Gly Asn Ser Leu Thr1
51869PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 186Gln His His Tyr Gly Thr Pro Leu Ala1
51879PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 187Gln His His Tyr Gly Thr Pro Leu Thr1
51889PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 188Gln Gln Gly Gln Ser Tyr Pro Leu Thr1
51899PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 189Gln Gln Trp Ser Gly Asn Ser Pro Thr1
51909PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 190Gln Gln Trp Ser Asn Tyr Pro Phe Thr1
51918PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 191Ala Tyr Ala Phe Thr Asn Tyr Leu1
51928PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 192Gly Asp Ser Phe Thr Ser Asp His1
51938PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 193Gly Phe Thr Phe Asn Thr Tyr Ala1
51948PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 194Gly Phe Thr Phe Ser Asp Tyr Gly1
51958PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 195Gly Phe Thr Phe Thr Asn Tyr Tyr1
51968PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 196Gly Tyr Pro Phe Thr Ser Tyr Trp1
51978PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 197Gly Tyr Ser Phe Ile Ser Tyr Trp1
51988PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 198Gly Tyr Ser Phe Ile Thr Tyr Trp1
51998PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 199Gly Tyr Ser Phe Thr Ala Tyr Trp1
52008PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 200Gly Tyr Ser Phe Thr Asn Tyr His1
52018PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 201Gly Tyr Ser Phe Thr Thr Tyr Trp1
52028PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 202Gly Tyr Ser Phe Val Thr Tyr Trp1
52039PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 203Gly Tyr Ser Ile Thr Ser Gly Tyr Tyr1
52048PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 204Gly Tyr Thr Phe Ile Ser Tyr Trp1
52058PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 205Gly Tyr Thr Phe Thr Ser Tyr Trp1
52068PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 206Gly Tyr Thr Phe Thr Thr Tyr Trp1
52078PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 207Ile Asn Pro Gly Ser Gly Gly Thr1
52088PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 208Ile Asn Pro Ser Gly Gly Asp Ser1
52098PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 209Ile Asn Pro Ser Ser Gly Asp Ala1
52108PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 210Ile Asn Pro Ser Ser Gly Asp Ser1
52118PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 211Ile Asn Pro Ser Ser Gly Asp Thr1
52128PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 212Ile Asn Pro Ser Ser Gly Gly Asn1
52138PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 213Ile Asn Pro Ser Ser Gly Gly Ser1
52148PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 214Ile Asn Pro Ser Ser Gly Gly Thr1
52158PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 215Ile Asn Pro Ser Ser Gly Asn Thr1
52168PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 216Ile Asn Pro Ser Ser Gly Ser Ala1
52178PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 217Ile Asn Pro Ser Ser Gly Ser Ser1
52187PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 218Ile Asn Tyr Asp Gly Ser Asn1
521910PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 219Ile Arg Ser Lys Ser Ser Asn Tyr Ala Thr1 5
102208PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 220Ile Ser Ser Gly Ser Ser Thr Ile1
52218PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 221Ile Tyr Leu Gly Ser Gly Asn Thr1
52228PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 222Ile Tyr Pro Gly Ser Gly Asn Thr1
52238PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 223Val Val Pro Asn Asn Asp Gly Thr1
52248PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 224Ala Arg Asp Tyr Tyr Gly Ala Tyr1
52258PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 225Ala Arg Asp Tyr Tyr Gly Asn Tyr1
522612PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 226Ala Arg Gly Asp Gly Tyr Gly Ser Leu Phe Ala
Tyr1 5 1022717PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 227Ala Arg Gly Tyr Tyr Tyr Gly Ser Ser
Tyr Gly Tyr Trp Tyr Phe Asp1 5 10 15Val22812PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 228Ala
Arg Pro Asp Asp Gly Tyr Tyr Gly Phe Ala Tyr1 5 1022911PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 229Ala
Arg Tyr Asp Tyr Asp Leu Tyr Leu Asp Ser1 5 1023011PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 230Ala
Arg Tyr Asp Tyr Asp Leu Tyr Leu His Ser1 5 102317PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 231Ala
Val Thr Tyr Phe Ala Tyr1 52328PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 232Thr Arg Asp Tyr Tyr Gly
Ala Tyr1 523315PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 233Val Arg Ala Trp Asp Tyr Gly Ser Ser
Trp Asp Tyr Phe Asp Tyr1 5 10 152348PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 234Val
Arg Asp Tyr Tyr Gly Ala Tyr1 52358PRTArtificial SequenceDescription
of Artificial Sequence Synthetic peptide 235Val Arg Asp Tyr Tyr Gly
Asn Tyr1 52368PRTArtificial SequenceDescription of Artificial
Sequence Synthetic peptide 236Val Arg Asp Tyr Tyr Gly Ser Tyr1
523711PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 237Val Thr Tyr Asp Tyr Asp Leu Tyr Phe Asp Asn1 5
10238107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 238Asp Ile Gln Met Thr Gln Ser Pro Ala Ser
Leu Ser Ala Ser Val Gly1 5 10 15Glu Thr Val Thr Ile Thr Cys Arg Ala
Ser Glu Asn Ile Tyr Ser Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Gln
Gly Lys Ser Pro Gln Leu Leu Val 35 40 45Tyr Asn Ala Lys Thr Leu Ala
Glu Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gln
Phe Ser Leu Lys Ile Asn Ser Leu Gln Pro65 70 75 80Glu Asp Phe Gly
Ser Tyr Tyr Cys Gln His His Tyr Gly Thr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Leu Lys 100 105239115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
239Glu Val Gln Leu Gln Gln Pro Gly Thr Glu Leu
Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Leu Ser Cys Lys Ala Ser Gly
Tyr Ser Phe Thr Thr Tyr 20 25 30Trp Met His Trp Val Lys Gln Arg Pro
Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Asn Ile Asn Pro Ser Ser Gly
Asp Ser His Tyr Asn Glu Lys Phe 50 55 60Lys Ser Lys Ala Thr Leu Thr
Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln Leu Asn Ser
Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr
Tyr Gly Ala Tyr Trp Gly His Gly Thr Leu Val Thr 100 105 110Val Ser
Ala 115240118PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 240Glu Val Gln Leu Val Glu Ser Gly
Gly Asp Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Lys Leu Ser Cys Ala
Ala Ser Gly Phe Thr Phe Ser Gly Tyr 20 25 30Gly Met Ser Trp Val Arg
Gln Thr Pro Asp Lys Arg Leu Glu Trp Val 35 40 45Ala Thr Ile Thr Ser
Gly Gly Thr Tyr Thr Tyr Tyr Pro Asp Ser Val 50 55 60Lys Gly Arg Phe
Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Tyr65 70 75 80Leu Gln
Ile Asp Ser Leu Lys Ser Glu Asp Thr Ala Ile Tyr Phe Cys 85 90 95Ala
Arg Ser Leu Ala Gly Asn Ala Met Asp Tyr Trp Gly Gln Gly Thr 100 105
110Ser Val Thr Val Ser Ser 115241107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
241Asp Ile Val Met Thr Gln Ser Pro Ala Thr Leu Ser Val Thr Pro Gly1
5 10 15Asp Arg Val Ser Leu Ser Cys Arg Ala Ser Gln Thr Ile Ser Asp
Tyr 20 25 30Leu His Trp Tyr Gln Gln Lys Ser His Glu Ser Pro Arg Leu
Leu Ile 35 40 45Lys Phe Ala Ser Gln Ser Ile Ser Gly Ile Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Ser Asp Phe Thr Leu Ser Ile Asn
Ser Val Glu Pro65 70 75 80Glu Asp Val Gly Val Tyr Tyr Cys Gln Asn
Gly His Gly Phe Pro Arg 85 90 95Thr Phe Gly Gly Gly Thr Lys Leu Glu
Ile Lys 100 105242116PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 242Gln Val Gln Leu Gln
Gln Pro Gly Ala Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Met Met
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25 30Asn Met His
Trp Val Lys Gln Thr Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Thr
Ile Tyr Pro Gly Asn Asp Asp Thr Ser Tyr Asn Gln Lys Phe 50 55 60Lys
Asp Lys Ala Thr Leu Thr Ala Asp Lys Ser Ser Ser Ala Ala Tyr65 70 75
80Met Gln Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys
85 90 95Ala Arg Gly Gly Tyr Arg Ala Met Asp Tyr Trp Gly Gln Thr Ser
Val 100 105 110Thr Val Ser Ser 115243112PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
243Asp Val Leu Met Thr Gln Thr Pro Leu Ser Leu Pro Val Ser Leu Gly1
5 10 15Asp Gln Ala Ser Ile Ser Cys Arg Ser Ser Gln Ser Ile Val Tyr
Ser 20 25 30Asn Gly Asn Thr Tyr Leu Gly Trp Tyr Leu Gln Lys Pro Gly
Gln Ser 35 40 45Pro Lys Leu Leu Ile Tyr Lys Val Ser Asn Arg Phe Ser
Gly Val Pro 50 55 60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu Ala Glu Asp Leu Gly Val
Tyr His Cys Phe Gln Gly 85 90 95Ser His Val Pro Tyr Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 110244118PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
244Gln Met Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Thr Gly Ser1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Phe Asn Ile Lys Asp
Tyr 20 25 30Tyr Leu His Trp Val Arg Gln Ala Pro Gly Gln Ala Leu Glu
Trp Met 35 40 45Gly Trp Ile Asp Pro Asp Gln Gly Asp Thr Glu Tyr Ala
Gln Lys Phe 50 55 60Gln Asp Arg Val Thr Ile Thr Arg Asp Arg Ser Met
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Met Tyr Tyr Cys 85 90 95Asn Ala Ala Tyr Gly Ser Ser Ser Tyr
Pro Met Asp Tyr Trp Gly Gln 100 105 110Gly Thr Thr Val Thr Val
115245107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 245Asn Ile Gln Met Thr Gln Ser Pro Ser Ala
Met Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Lys Ala
Ser Gln Asp Ile His Arg Tyr 20 25 30Leu Ser Trp Phe Gln Gln Lys Pro
Gly Lys Val Pro Lys His Leu Ile 35 40 45Tyr Arg Ala Asn Arg Leu Val
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Leu Gln Tyr Asp Glu Phe Pro Tyr 85 90 95Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile Lys 100 105246117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
246Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Ser 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Thr Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
115247117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 247Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Asp Ile 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser
Gly Asn Val Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Glu Asp Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115248117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 248Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp Asn 20 25 30Tyr Ile Asn
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala Val Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115249117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
249Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Phe 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
115250117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 250Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Asp His 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser
Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Glu Asp Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115251117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 251Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp Asn 20 25 30Tyr Ile Asn
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Val Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115252117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
252Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Leu 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
115253117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 253Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Asp Asn 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser
Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Glu Glu Arg Gly Phe Ala Pro Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115254117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 254Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp Ser 20 25 30Tyr Ile Asn
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115255117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
255Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Phe 20 25 30Tyr Ile His Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Asp Gly Phe Ala
His Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
115256117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 256Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Asp Asn 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser
Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Lys Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115257117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 257Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp Thr 20 25 30Tyr Ile Asn
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Ile Tyr Leu Gly Ser Gly Asn Ile Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115258117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
258Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Tyr 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Glu Asp Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115259117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 259Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp Thr 20 25 30Tyr Ile Asn
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Ile Tyr Leu Gly Ser Gly Asn Val Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala His Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115260117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
260Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Leu 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Val Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
115261117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 261Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Asp Ser 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Ile Tyr Gly Gly Ser
Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Thr Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Glu Glu Asp Gly Phe Ala Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115262117PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 262Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp Ser 20 25 30Tyr Ile Asn
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Ile Tyr Gly Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Glu Glu Arg Gly Phe Ala Tyr Trp Gly Gln Gly Thr
Leu 100 105 110Val Thr Val Ser Ser 115263117PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
263Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asp
Ser 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Met 35 40 45Gly Trp Ile Tyr Leu Gly Ser Gly Asn Val Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Ile Glu Arg Gly Phe Ala
Tyr Trp Gly Gln Gly Thr Leu 100 105 110Val Thr Val Ser Ser
115264117PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 264Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Thr Phe Thr Asp Ser 20 25 30Tyr Ile Asn Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp Val Tyr Leu Gly Ser
Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile
Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Glu Glu Arg Gly Phe Ala Val Trp Gly Gln Gly Thr Leu 100 105 110Val
Thr Val Ser Ser 115265113PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 265Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Ile Leu Tyr Ser 20 25 30Ser Asn Gln
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Tyr Tyr His Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys266112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 266Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Ser Ser 20 25 30Asn Gln Lys
Tyr Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Thr Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Phe
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110267112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 267Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Tyr Ser Ser 20 25 30Asn Lys Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Ser Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110268112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 268Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Ser Ser 20 25 30Asn Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Phe
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110269112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 269Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Ser Ser 20 25 30Ser Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110270113PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 270Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Lys
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Tyr Tyr His Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys271112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 271Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Val Ser Ser 20 25 30Ser Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110272112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 272Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Ser Ser 20 25 30Ser Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Thr Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Phe
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110273113PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 273Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Gln
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Phe Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys274113PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 274Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Tyr Lys 20 25 30Ser Asn Gln
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Leu Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys275113PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 275Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Gln
Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro
Lys Leu Leu Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75
80Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln
85 90 95Tyr Tyr His Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile 100 105 110Lys276112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 276Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Ser Ser 20 25 30Ser Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Gly Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110277112PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 277Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn Cys Lys Ser Ser Gln Ser Leu Leu Thr Ser 20 25 30Asn Gln Lys
Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Ile Arg Asp Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75
80Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr
85 90 95Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105 110278113PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 278Asp Ile Val Met Thr
Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr
Ile Asn
Cys Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Arg Lys Asn
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu
Leu Ile Tyr Trp Ala Ser Ser Arg Asp Ser Gly Val 50 55 60Pro Asp Arg
Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile
Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90
95Tyr Tyr His Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile
100 105 110Lys279113PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 279Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Ser Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Phe
Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys280112PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 280Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Leu Leu Ser Ser 20 25 30Asn Asn Lys Asn Tyr Leu
Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys Leu Leu Ile
Tyr Trp Ala Ser Val Arg Asp Ser Gly Val Pro 50 55 60Asp Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75 80Ser Ser
Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr 85 90 95Tyr
Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
110281112PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 281Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Leu Tyr Thr Ser 20 25 30Asn Gln Lys Asn Tyr Leu Ala Trp
Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys Leu Leu Ile Tyr Trp
Ala Ser Thr Arg Asp Ser Gly Val Pro 50 55 60Asp Arg Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75 80Ser Ser Leu Gln
Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr 85 90 95Tyr Ser Tyr
Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
110282113PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 282Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Leu Leu Tyr Asn 20 25 30Ser Asn Gln Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr His
Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys283113PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 283Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Leu Leu Tyr Lys 20 25 30Ser Asn Gln Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Phe
Tyr Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys284112PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 284Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Leu Tyr Ser Ser 20 25 30Ser Gln Lys Asn Tyr Leu
Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro 35 40 45Pro Lys Leu Leu Ile
Tyr Trp Ala Ser Val Arg Asp Ser Gly Val Pro 50 55 60Asp Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile65 70 75 80Ser Ser
Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr 85 90 95Tyr
Ser Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
110285113PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 285Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Leu Leu Tyr Asn 20 25 30Ser Asn Gln Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr His
Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys286113PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 286Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Leu Leu Tyr Ser 20 25 30Ser Asn Gln Lys Asn Tyr
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu
Ile Tyr Trp Ala Ser Val Arg Asp Ser Gly Val 50 55 60Pro Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr
Tyr His Tyr Pro Leu Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys287107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 287Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
Arg Ala Ser Lys Asn Ile Gly Lys Tyr 20 25 30Leu Ala Trp Phe Gln Gln
Lys Pro Gly Lys Ala Pro Lys Ser Leu Ile 35 40 45Tyr Ser Gly Ser Thr
Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp
Phe Ala Thr Tyr Tyr Cys Gln Gln His Asn Glu Tyr Pro Tyr 85 90 95Thr
Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105288107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
288Asp Val Gln Ile Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Lys Asn Ile Gly Lys
Tyr 20 25 30Leu Ala Trp Phe Gln Gln Lys Pro Gly Lys Ala Pro Lys Ser
Leu Ile 35 40 45Tyr Ser Gly Ser Thr Leu Gln Ser Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
His Asn Glu Tyr Pro Tyr 85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys 100 105289107PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 289Asp Ile Gln Met Thr
Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr
Ile Thr Cys Arg Ala Ser Lys Asn Ile Gly Lys Tyr 20 25 30Leu Ala Trp
Phe Gln Gln Lys Pro Gly Lys Ala Pro Lys Ser Leu Ile 35 40 45Tyr Ser
Gly Ser Thr Leu Gln Ser Gly Ile Pro Ser Arg Phe Ser Gly 50 55 60Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln His Asn Glu Tyr Pro Tyr
85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100
105290107PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 290Asp Val Gln Ile Thr Gln Ser Pro Ser Ser
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala
Ser Lys Asn Ile Gly Lys Tyr 20 25 30Leu Ala Trp Phe Gln Gln Lys Pro
Gly Lys Ala Pro Lys Ser Leu Ile 35 40 45Tyr Ser Gly Ser Thr Leu Gln
Ser Gly Ile Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln His Asn Glu Tyr Pro Tyr 85 90 95Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile Lys 100 105291107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
291Asp Val Gln Ile Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Lys Asn Ile Gly Lys
Tyr 20 25 30Leu Ala Trp Phe Gln Gln Lys Pro Gly Lys Thr Asn Lys Leu
Leu Ile 35 40 45Tyr Ser Gly Ser Thr Leu Gln Ser Gly Ile Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
His Asn Glu Tyr Pro Tyr 85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys 100 105292120PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 292Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25 30Trp Met Gln
Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Met
Ile Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys
Asp Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly
Gln 100 105 110Gly Thr Leu Val Thr Val Ser Ser 115
120293120PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 293Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Thr Ser
Gly Tyr Ser Phe Thr Asn Tyr 20 25 30Trp Met Gln Trp Val Arg Gln Ala
Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Met Ile Asp Pro Ser Asp
Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys Asp Arg Val Thr Met
Thr Arg Asp Thr Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg
Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly Gln 100 105 110Gly
Thr Leu Val Thr Val Ser Ser 115 120294120PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
294Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn
Tyr 20 25 30Trp Met Gln Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Met Ile Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn
Gln Lys Phe 50 55 60Lys Asp Arg Ala Thr Leu Thr Val Asp Thr Ser Thr
Ser Thr Val Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Gly Ser Pro Met Ile Thr
Ser Phe Ala Tyr Trp Gly Gln 100 105 110Gly Thr Leu Val Thr Val Ser
Ser 115 120295120PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 295Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys
Thr Ser Gly Tyr Ser Phe Thr Asn Tyr 20 25 30Trp Met Gln Trp Val Arg
Gln Ala Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Met Ile Asp Pro
Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys Asp Arg Ala
Thr Leu Thr Val Asp Thr Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu
Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala
Arg Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly Gln 100 105
110Gly Thr Leu Val Thr Val Ser Ser 115 120296120PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
296Gln Val Gln Leu Gln Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Thr Ser Gly Tyr Ser Phe Thr Asn
Tyr 20 25 30Trp Met Gln Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Ile 35 40 45Gly Met Ile Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn
Gln Lys Phe 50 55 60Lys Asp Lys Ala Thr Leu Thr Val Asp Lys Ser Thr
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Arg Gly Ser Pro Met Ile Thr
Ser Phe Ala Tyr Trp Gly Gln 100 105 110Gly Thr Leu Val Thr Val Ser
Ser 115 120297107PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 297Asp Ile Gln Met Thr Gln Ser Pro
Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys
His Ala Ser Gln Asn Ile Asn Val Trp 20 25 30Leu Ser Trp Tyr Gln Gln
Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala Ser Asn
Leu His Thr Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp
Phe Ala Thr Tyr Tyr Cys Gln Gln Gly Gln Ser Tyr Pro Leu 85 90 95Thr
Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105298107PRTArtificial
SequenceDescription of Artificial Sequence
Synthetic polypeptide 298Asp Ile Gln Ile Thr Gln Ser Pro Ser Ser
Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys His Ala
Ser Gln Asn Ile Asn Val Trp 20 25 30Leu Ser Trp Tyr Gln Gln Lys Pro
Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala Ser Asn Leu His
Thr Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gly
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln Gly Gln Ser Tyr Pro Leu 85 90 95Thr Phe Gly
Gln Gly Thr Lys Leu Glu Ile Lys 100 105299107PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
299Asp Ile Gln Ile Thr Gln Ser Pro Ser Ser Val Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys His Ala Ser Gln Asn Ile Asn Val
Trp 20 25 30Leu Ser Trp Tyr Gln Gln Lys Pro Gly Lys Ile Pro Lys Leu
Leu Ile 35 40 45Tyr Lys Ala Ser Asn Leu His Thr Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gly Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
Gly Gln Ser Tyr Pro Leu 85 90 95Thr Phe Gly Gln Gly Thr Lys Leu Glu
Ile Lys 100 105300118PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 300Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25 30His Ile His
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Met 35 40 45Gly Trp
Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Tyr Asp Tyr Asp Leu Tyr Leu His Ser Trp Gly Gln Gly
Thr 100 105 110Leu Val Thr Val Ser Ser 115301118PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
301Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Asn
Tyr 20 25 30His Ile His Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu
Trp Ile 35 40 45Gly Trp Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Arg Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Tyr Asp Tyr Asp Leu Tyr Leu
His Ser Trp Gly Gln Gly Thr 100 105 110Leu Val Thr Val Ser Ser
115302118PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 302Gln Val Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Tyr Ser Phe Thr Asn Tyr 20 25 30His Ile His Trp Val Arg Gln Ala
Pro Gly Gln Arg Leu Glu Trp Ile 35 40 45Gly Trp Ile Tyr Pro Gly Ser
Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Arg Ala Thr Leu
Thr Ala Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Tyr
Asp Tyr Asp Leu Tyr Leu His Ser Trp Gly Gln Gly Thr 100 105 110Leu
Val Thr Val Ser Ser 115303118PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 303Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val
Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Asn Tyr 20 25 30His Ile His
Trp Val Arg Gln Ala Pro Gly Gln Arg Leu Glu Trp Ile 35 40 45Gly Trp
Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys
Gly Met Ala Thr Leu Thr Ala Asp Thr Ser Ala Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Tyr Asp Tyr Asp Leu Tyr Leu His Ser Trp Gly Gln Gly
Thr 100 105 110Leu Val Thr Val Ser Ser 115304118PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
304Glu Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ser Phe Thr Asn
Tyr 20 25 30His Ile His Trp Val Lys Gln Ala Pro Gly Gln Arg Leu Glu
Trp Ile 35 40 45Gly Trp Ile Tyr Pro Gly Ser Gly Asn Thr Lys Tyr Asn
Glu Lys Phe 50 55 60Lys Gly Met Ala Thr Leu Thr Ala Asp Thr Ser Ala
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Tyr Asp Tyr Asp Leu Tyr Leu
His Ser Trp Gly Gln Gly Thr 100 105 110Leu Val Thr Val Ser Ser
115305447PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 305Gln Met Gln Leu Val Gln Ser Gly Ala Glu
Val Lys Lys Thr Gly Ser1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser
Gly Phe Asn Ile Lys Asp Tyr 20 25 30Tyr Leu His Trp Val Arg Gln Ala
Pro Gly Gln Ala Leu Glu Trp Met 35 40 45Gly Trp Ile Asp Pro Asp Gln
Gly Asp Thr Glu Tyr Ala Gln Lys Phe 50 55 60Gln Asp Arg Val Thr Ile
Thr Arg Asp Arg Ser Met Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys 85 90 95Asn Ala Ala
Tyr Gly Ser Ser Ser Tyr Pro Met Asp Tyr Trp Gly Gln 100 105 110Gly
Thr Thr Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val 115 120
125Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala
130 135 140Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr
Val Ser145 150 155 160Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr Phe Pro Ala Val 165 170 175Leu Gln Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val Val Thr Val Pro 180 185 190Ser Ser Ser Leu Gly Thr Lys
Thr Tyr Thr Cys Asn Val Asp His Lys 195 200 205Pro Ser Asn Thr Lys
Val Asp Lys Arg Val Glu Ser Lys Tyr Gly Pro 210 215 220Pro Cys Pro
Pro Cys Pro Ala Pro Glu Phe Glu Gly Gly Pro Ser Val225 230 235
240Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr
245 250 255Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp
Pro Glu 260 265 270Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val
His Asn Ala Lys 275 280 285Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser
Thr Tyr Arg Val Val Ser 290 295 300Val Leu Thr Val Leu His Gln Asp
Trp Leu Asn Gly Lys Glu Tyr Lys305 310 315 320Cys Lys Val Ser Asn
Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile 325 330 335Ser Lys Ala
Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro 340 345 350Pro
Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu 355 360
365Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn
370 375 380Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu
Asp Ser385 390 395 400Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr
Val Asp Lys Ser Arg 405 410 415Trp Gln Glu Gly Asn Val Phe Ser Cys
Ser Val Met His Glu Ala Leu 420 425 430His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu Ser Leu Gly Lys 435 440 445306212PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
306Gln Met Thr Gln Ser Pro Ser Ala Met Ser Ala Ser Val Gly Asp Arg1
5 10 15Val Thr Ile Thr Cys Lys Ala Ser Gln Asp Ile His Arg Tyr Leu
Ser 20 25 30Trp Phe Gln Gln Lys Pro Gly Lys Val Pro Lys His Leu Ile
Tyr Arg 35 40 45Ala Asn Arg Leu Val Ser Gly Val Pro Ser Arg Phe Ser
Gly Ser Gly 50 55 60Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu
Gln Pro Glu Asp65 70 75 80Phe Ala Thr Tyr Tyr Cys Leu Gln Tyr Asp
Glu Phe Pro Tyr Thr Phe 85 90 95Gly Gly Gly Thr Lys Val Glu Ile Lys
Arg Thr Val Ala Ala Pro Ser 100 105 110Val Phe Ile Phe Pro Pro Ser
Asp Glu Gln Leu Lys Ser Gly Thr Ala 115 120 125Ser Val Val Cys Leu
Leu Asn Asn Phe Tyr Pro Arg Glu Ala Lys Val 130 135 140Gln Trp Lys
Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln Glu Ser145 150 155
160Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser Ser Thr
165 170 175Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr
Ala Cys 180 185 190Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr
Lys Ser Phe Asn 195 200 205Arg Gly Glu Cys 210307447PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
307Gln Met Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Thr Gly Ser1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Phe Asn Ile Lys Asp
Tyr 20 25 30Tyr Leu His Trp Val Arg Gln Ala Pro Gly Gln Ala Leu Glu
Trp Met 35 40 45Gly Trp Ile Asp Pro Asp Gln Gly Asp Thr Glu Tyr Ala
Gln Lys Phe 50 55 60Gln Asp Arg Val Thr Ile Thr Arg Asp Arg Ser Met
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Met Tyr Tyr Cys 85 90 95Asn Ala Ala Tyr Gly Ser Ser Ser Tyr
Pro Met Asp Tyr Trp Gly Gln 100 105 110Gly Thr Thr Val Thr Val Ser
Ser Ala Ser Thr Lys Gly Pro Ser Val 115 120 125Phe Pro Leu Ala Pro
Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala 130 135 140Leu Gly Cys
Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser145 150 155
160Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val
165 170 175Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val Pro 180 185 190Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His Lys 195 200 205Pro Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly Pro 210 215 220Pro Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser Val225 230 235 240Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr 245 250 255Pro Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu 260 265 270Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys 275 280
285Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser
290 295 300Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr Lys305 310 315 320Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser
Ile Glu Lys Thr Ile 325 330 335Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro 340 345 350Pro Ser Gln Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys Leu 355 360 365Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn 370 375 380Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser385 390 395
400Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg
405 410 415Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu
Ala Leu 420 425 430His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Leu Gly Lys 435 440 445308212PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 308Gln Met Thr Gln Ser
Pro Ser Ala Met Ser Ala Ser Val Gly Asp Arg1 5 10 15Val Thr Ile Thr
Cys Lys Ala Ser Gln Asp Ile His Arg Tyr Leu Ser 20 25 30Trp Phe Gln
Gln Lys Pro Gly Lys Val Pro Lys His Leu Ile Tyr Arg 35 40 45Ala Asn
Arg Leu Val Ser Gly Val Pro Ser Arg Phe Ser Gly Ser Gly 50 55 60Ser
Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro Glu Asp65 70 75
80Phe Ala Thr Tyr Tyr Cys Leu Gln Tyr Asp Glu Phe Pro Tyr Thr Phe
85 90 95Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala Pro
Ser 100 105 110Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser
Gly Thr Ala 115 120 125Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro
Arg Glu Ala Lys Val 130 135 140Gln Trp Lys Val Asp Asn Ala Leu Gln
Ser Gly Asn Ser Gln Glu Ser145 150 155 160Val Thr Glu Gln Asp Ser
Lys Asp Ser Thr Tyr Ser Leu Ser Ser Thr 165 170 175Leu Thr Leu Ser
Lys Ala Asp Tyr Glu Lys His Lys Val Tyr Ala Cys 180 185 190Glu Val
Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser Phe Asn 195 200
205Arg Gly Glu Cys 210309214PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 309Asp Val Gln Ile Thr
Gln Ser Pro Ser Phe Leu Ala Ala Ser Pro Gly1 5 10 15Glu Thr Ile Thr
Ile Asn Cys Arg Ala Ser Lys Asn Ile Gly Lys Tyr 20 25 30Leu Ala Trp
Phe Gln Glu Lys Pro Gly Lys Thr Asn Lys Leu Leu Ile 35 40 45Tyr Ser
Gly Ser Thr Leu Gln Ser Gly Ile Pro Ser Arg Phe Ser Gly 50 55 60Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro65 70 75
80Glu Asp Phe Ala Ile Tyr Tyr Cys Gln Gln His Asn Glu Tyr Pro Tyr
85 90 95Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 210310447PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
310Gln Val Gln Leu Gln Gln
Ser Gly Pro Gln Leu Val Arg Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser
Cys Lys Thr Ser Gly Tyr Ser Phe Thr Asn Tyr 20 25 30Trp Met Gln Trp
Val Lys Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Met Ile
Asp Pro Ser Asp Ser Glu Ser Arg Leu Asn Gln Lys Phe 50 55 60Lys Asp
Lys Ala Thr Leu Thr Val Asp Lys Ser Ser Ser Thr Ala Tyr65 70 75
80Met Gln Leu Ser Ser Pro Thr Phe Glu Asp Ser Ala Val Tyr Tyr Cys
85 90 95Ala Arg Arg Gly Ser Pro Met Ile Thr Ser Phe Ala Tyr Trp Gly
Gln 100 105 110Gly Thr Leu Val Thr Val Ser Ala Ala Ser Thr Lys Gly
Pro Ser Val 115 120 125Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser
Glu Ser Thr Ala Ala 130 135 140Leu Gly Cys Leu Val Lys Asp Tyr Phe
Pro Glu Pro Val Thr Val Ser145 150 155 160Trp Asn Ser Gly Ala Leu
Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175Leu Gln Ser Ser
Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185 190Ser Ser
Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His Lys 195 200
205Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr Gly Pro
210 215 220Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu Gly Gly Pro
Ser Val225 230 235 240Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu
Met Ile Ser Arg Thr 245 250 255Pro Glu Val Thr Cys Val Val Val Asp
Val Ser Gln Glu Asp Pro Glu 260 265 270Val Gln Phe Asn Trp Tyr Val
Asp Gly Val Glu Val His Asn Ala Lys 275 280 285Thr Lys Pro Arg Glu
Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser 290 295 300Val Leu Thr
Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys305 310 315
320Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile
325 330 335Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro 340 345 350Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu 355 360 365Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala
Val Glu Trp Glu Ser Asn 370 375 380Gly Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser385 390 395 400Asp Gly Ser Phe Phe
Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg 405 410 415Trp Gln Glu
Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu 420 425 430His
Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440
445311214PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 311Asp Ile Gln Ile Asn Gln Ser Pro Ser Ser
Leu Ser Ala Ser Leu Gly1 5 10 15Asp Thr Ile Thr Ile Thr Cys His Ala
Ser Gln Asn Ile Asn Val Trp 20 25 30Leu Ser Trp Tyr Gln Gln Lys Pro
Gly Asn Ile Pro Lys Leu Leu Ile 35 40 45Tyr Lys Ala Ser Asn Leu His
Thr Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Gly
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Ile Ala
Thr Tyr Tyr Cys Gln Gln Gly Gln Ser Tyr Pro Leu 85 90 95Thr Phe Gly
Ala Gly Thr Lys Leu Glu Ile Lys Arg Thr Val Ala Ala 100 105 110Pro
Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120
125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala
130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn
Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser
Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp
Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln
Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu
Cys 210312445PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 312Glu Val Gln Leu Gln Gln Ser Gly
Pro Glu Leu Val Lys Pro Gly Ala1 5 10 15Ser Val Lys Ile Ser Cys Lys
Ala Ser Gly Tyr Ser Phe Thr Asn Tyr 20 25 30His Ile His Trp Val Lys
Gln Arg Pro Gly Gln Gly Leu Glu Trp Ile 35 40 45Gly Trp Ile Tyr Pro
Gly Ser Gly Asn Thr Lys Tyr Asn Glu Lys Phe 50 55 60Lys Gly Met Ala
Ile Leu Thr Ala Asp Thr Ser Ser Ser Thr Ala Tyr65 70 75 80Met Gln
Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala Val Tyr Tyr Cys 85 90 95Ala
Arg Tyr Asp Tyr Asp Leu Tyr Leu His Ser Trp Gly Gln Gly Thr 100 105
110Thr Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe Pro
115 120 125Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala
Leu Gly 130 135 140Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr
Val Ser Trp Asn145 150 155 160Ser Gly Ala Leu Thr Ser Gly Val His
Thr Phe Pro Ala Val Leu Gln 165 170 175Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val Val Thr Val Pro Ser Ser 180 185 190Ser Leu Gly Thr Lys
Thr Tyr Thr Cys Asn Val Asp His Lys Pro Ser 195 200 205Asn Thr Lys
Val Asp Lys Arg Val Glu Ser Lys Tyr Gly Pro Pro Cys 210 215 220Pro
Pro Cys Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val Phe Leu225 230
235 240Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro
Glu 245 250 255Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro
Glu Val Gln 260 265 270Phe Asn Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala Lys Thr Lys 275 280 285Pro Arg Glu Glu Gln Phe Asn Ser Thr
Tyr Arg Val Val Ser Val Leu 290 295 300Thr Val Leu His Gln Asp Trp
Leu Asn Gly Lys Glu Tyr Lys Cys Lys305 310 315 320Val Ser Asn Lys
Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile Ser Lys 325 330 335Ala Lys
Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser 340 345
350Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys
355 360 365Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn
Gly Gln 370 375 380Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu
Asp Ser Asp Gly385 390 395 400Ser Phe Phe Leu Tyr Ser Arg Leu Thr
Val Asp Lys Ser Arg Trp Gln 405 410 415Glu Gly Asn Val Phe Ser Cys
Ser Val Met His Glu Ala Leu His Asn 420 425 430His Tyr Thr Gln Lys
Ser Leu Ser Leu Ser Leu Gly Lys 435 440 4453136PRTArtificial
SequenceDescription of Artificial Sequence Synthetic 6xHis tag
313His His His His His His1 5
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016
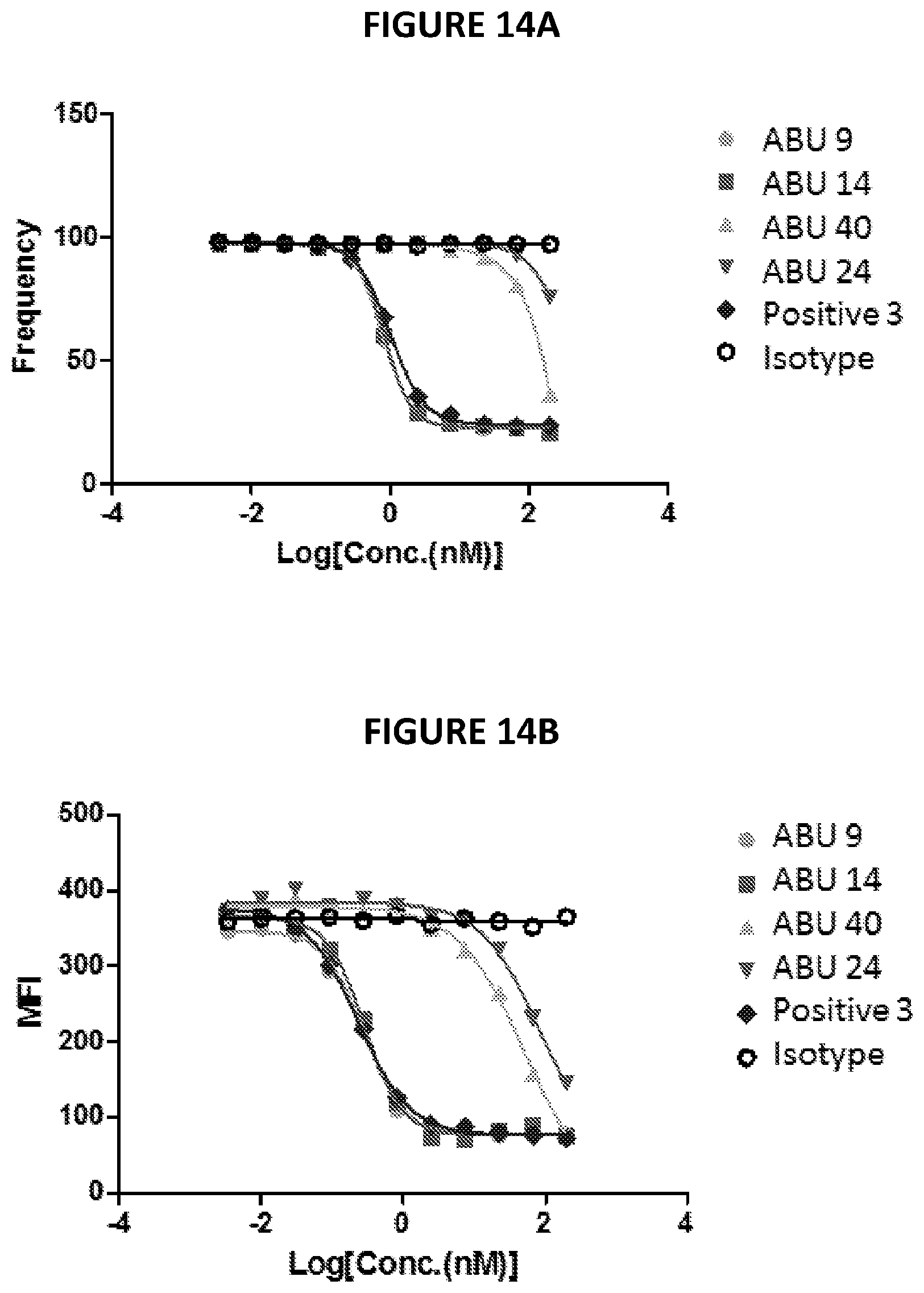
D00017

D00018

D00019

D00020
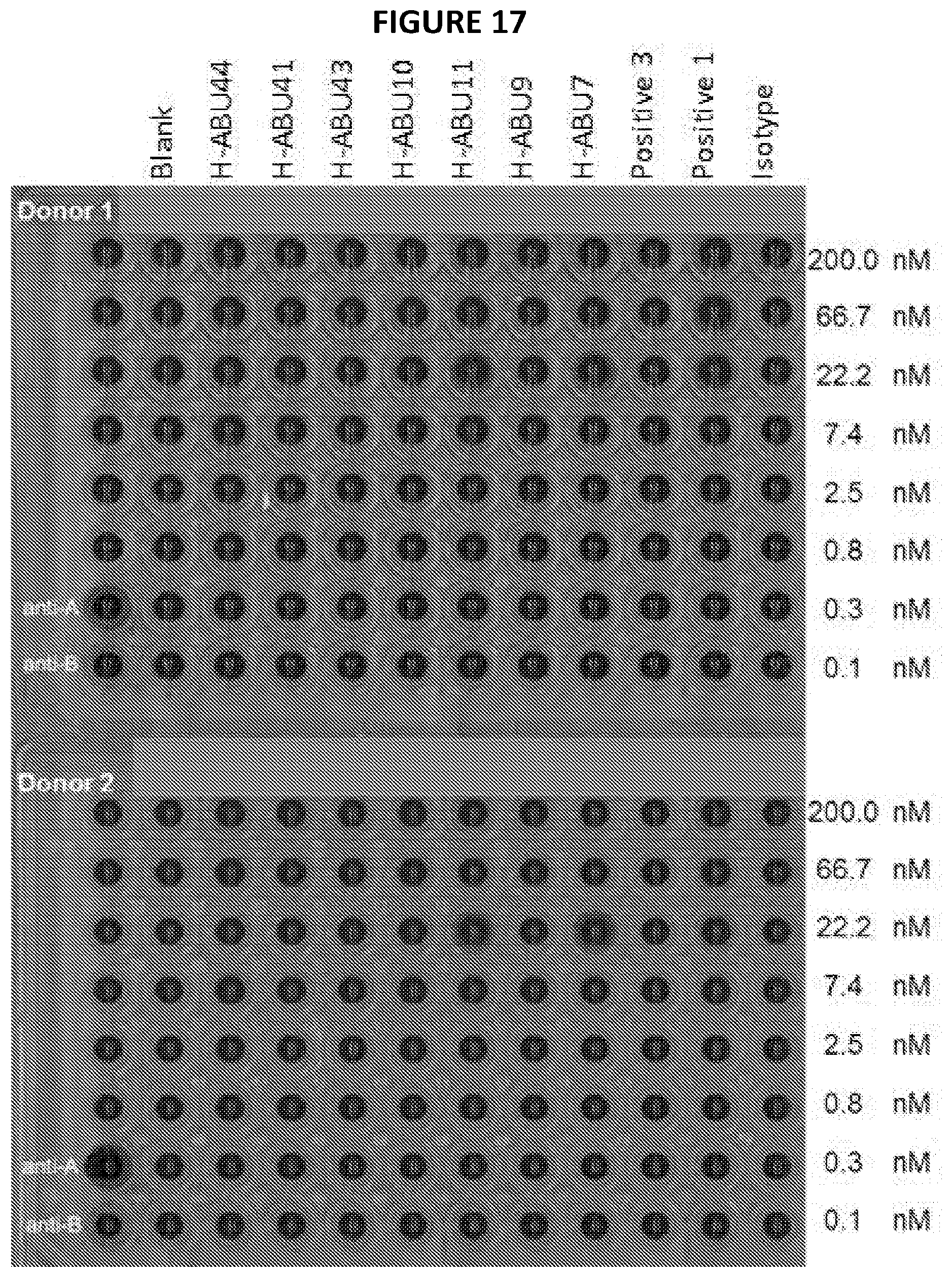
D00021
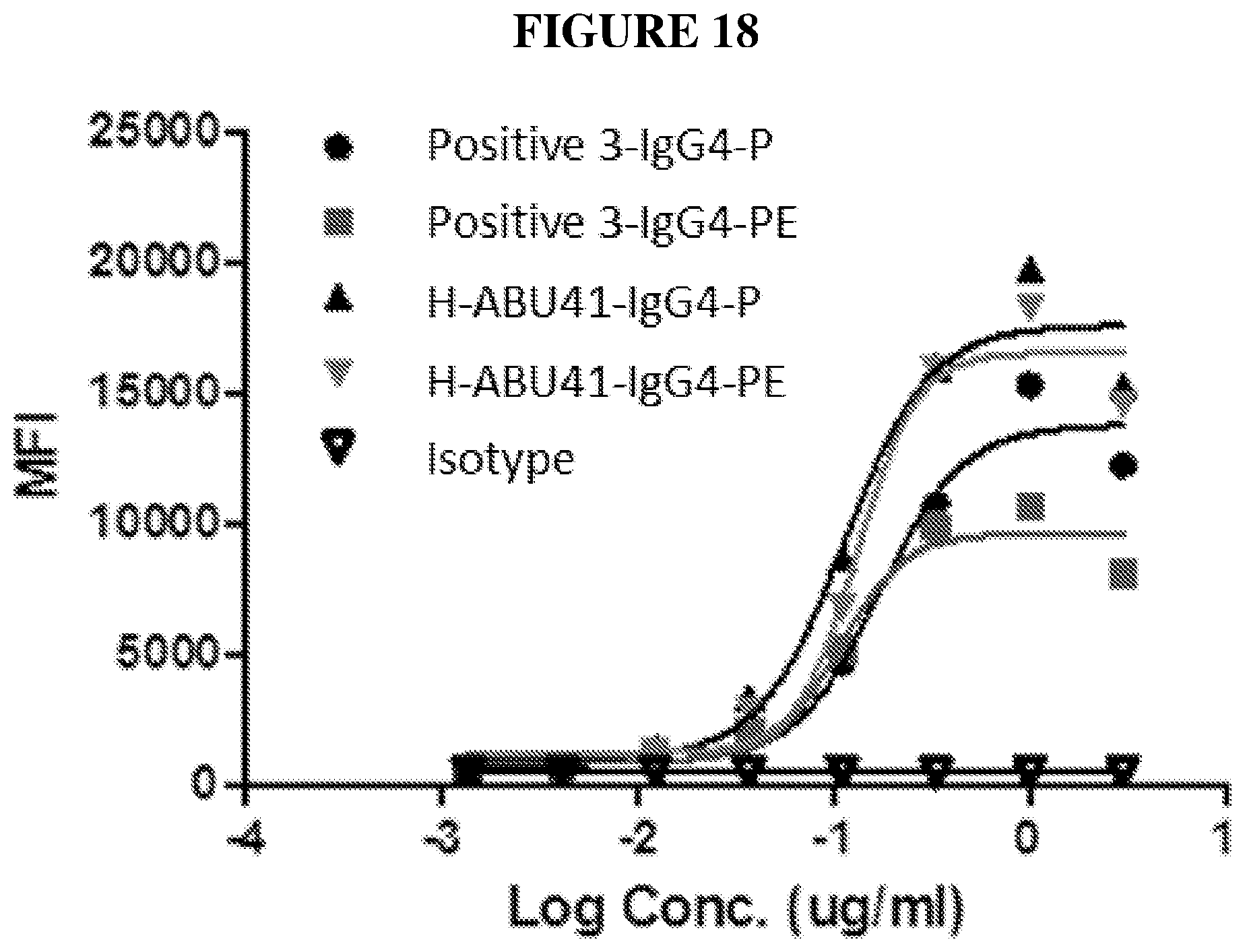
D00022

D00023

D00024

D00025

D00026

D00027

D00028
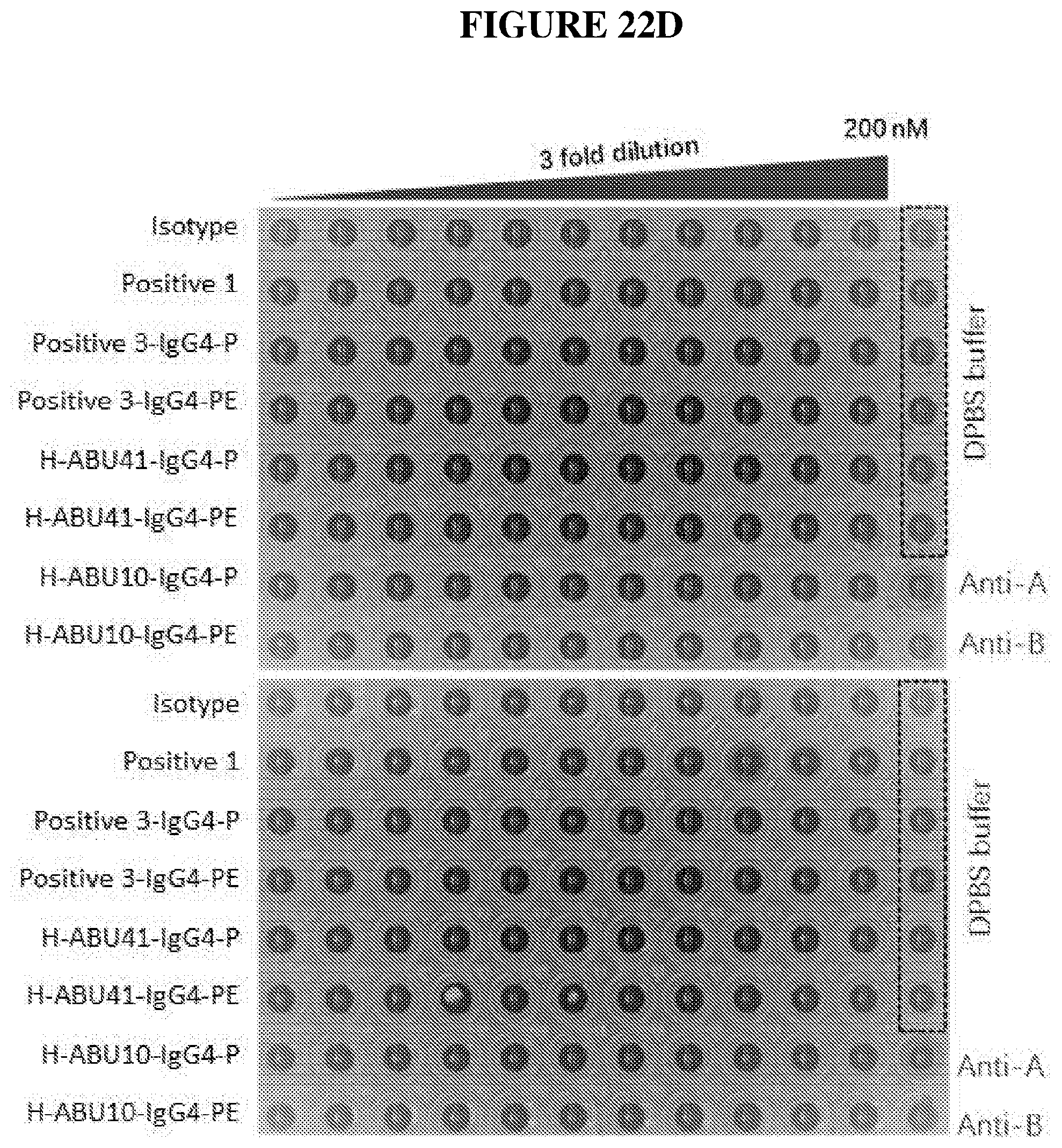

D00029
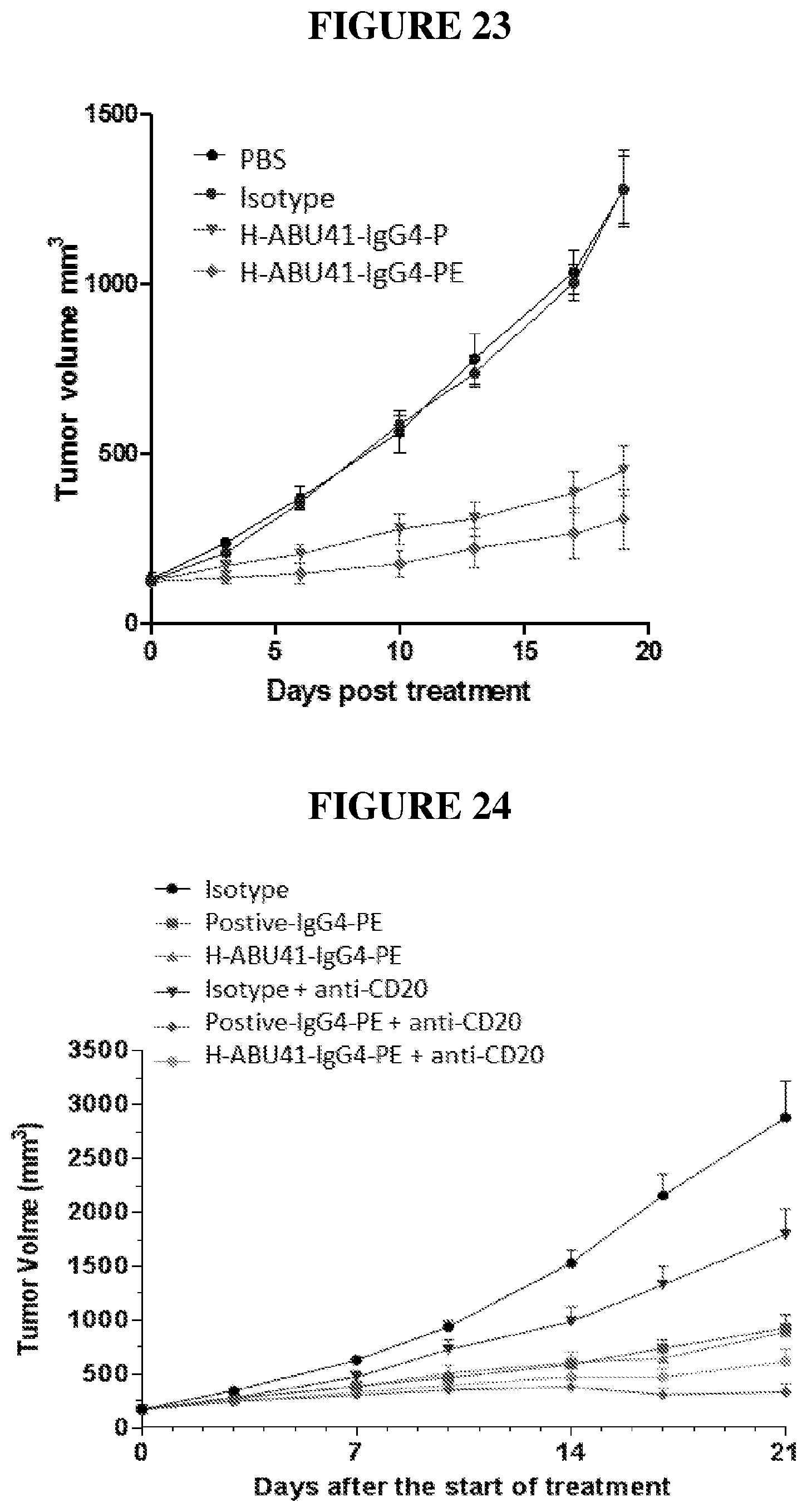
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.