Recombinant Vectors
Gruber; Harry E. ; et al.
U.S. patent application number 16/560525 was filed with the patent office on 2020-07-30 for recombinant vectors. The applicant listed for this patent is Tocagen Inc.. Invention is credited to Harry E. Gruber, Douglas J. Jolly, Christopher R. Logg, Omar D. Perez.
| Application Number | 20200239858 16/560525 |
| Document ID | 20200239858 / US20200239858 |
| Family ID | 1000004780645 |
| Filed Date | 2020-07-30 |
| Patent Application | download [pdf] |

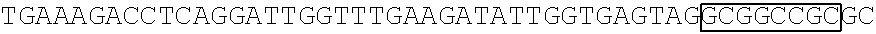
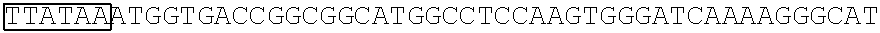
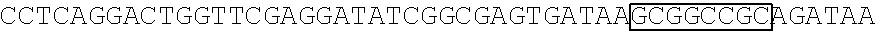
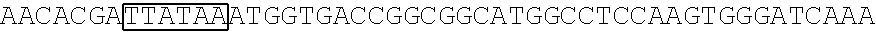
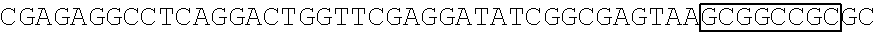
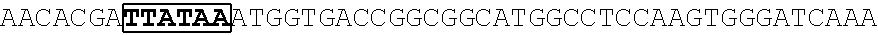
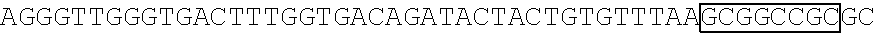
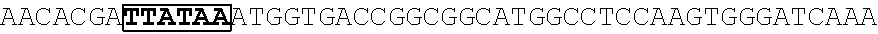
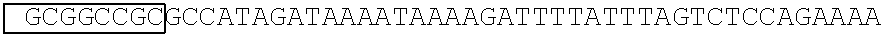
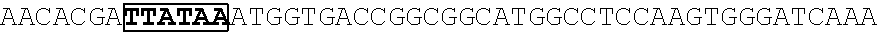
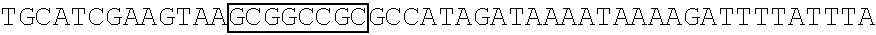
View All Diagrams
| United States Patent Application | 20200239858 |
| Kind Code | A1 |
| Gruber; Harry E. ; et al. | July 30, 2020 |
RECOMBINANT VECTORS
Abstract
This disclosure provides modified cytosine deaminases (CDs). The disclosure further relates to cells and vector expressing or comprising such modified CDs and methods of using such modified CDs in the treatment of disease and disorders.
| Inventors: | Gruber; Harry E.; (Rancho Santa Fe, CA) ; Jolly; Douglas J.; (Encinitas, CA) ; Perez; Omar D.; (San Diego, CA) ; Logg; Christopher R.; (South Pasadena, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 1000004780645 | ||||||||||
| Appl. No.: | 16/560525 | ||||||||||
| Filed: | September 4, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15672029 | Aug 8, 2017 | 10407666 | ||
| 16560525 | ||||
| 14477741 | Sep 4, 2014 | 9732326 | ||
| 15672029 | ||||
| 13072704 | Mar 26, 2011 | 8829173 | ||
| 14477741 | ||||
| PCT/US09/58512 | Sep 26, 2009 | |||
| 13072704 | ||||
| 61100666 | Sep 26, 2008 | |||
| 61120618 | Dec 8, 2008 | |||
| 61186823 | Jun 13, 2009 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 2760/16043 20130101; A61K 48/00 20130101; C12N 9/78 20130101; C12N 2760/16032 20130101; A61K 31/513 20130101; C12N 7/00 20130101; C12N 2760/16021 20130101; C12N 15/86 20130101; A61K 38/2292 20130101; C12N 2740/13043 20130101 |
| International Class: | C12N 7/00 20060101 C12N007/00; C12N 15/86 20060101 C12N015/86; A61K 31/513 20060101 A61K031/513; A61K 38/22 20060101 A61K038/22; C12N 9/78 20060101 C12N009/78; A61K 48/00 20060101 A61K048/00 |
Claims
1-60. (canceled)
61. A recombinant replication competent gammaretrovirus comprising: a gammaretroviral GAG protein; a gammaretroviral POL protein; a gammaretroviral envelope; a gammaretroviral RNA polynucleotide comprising 3' untranslated region (U3) and repeat region (R) sequences from murine leukemia virus (MLV), Moloney murine leukemia virus (MoMLV), Feline leukemia virus (FeLV), Baboon endogenous retrovirus (BEV), porcine endogenous virus (PERV), the cat derived retrovirus RD114, squirrel monkey retrovirus, Xenotropic murine leukemia virus-related virus(XMRV), avian reticuloendotheliosis virus(REV), or Gibbon ape leukemia virus (GALV) at the 3' end of the gammaretroviral polynucleotide sequence, an R and 5' untranslated region (U5) sequence from MLV, MoMLV, FeLV, BEV, PERV, RD114, squirrel monkey retrovirus, XMRV, REV or GALV at the 5' end of the gammaretroviral polynucleotide, a gag nucleic acid domain, a pol nucleic acid domain and an env nucleic acid domain from MLV, MoMLV, FeLV, BEV, PERV, RD114, squirrel monkey retrovirus, XMRV, REV or GALV located between the U5 and U3 regions; a cassette comprising an internal ribosome entry site (IRES) upstream and operably linked to a heterologous polynucleotide followed by pol-III promoter linked to an inhibitory polynucleotide, wherein the cassette is positioned 5' to the U3 region and 3' to the env nucleic acid domain; and cis-acting sequences necessary for reverse transcription, packaging and integration in a target cell.
62. The retrovirus according to claim 61, wherein the heterologous nucleic acid encodes a polypeptide that converts a nontoxic prodrug in to a toxic drug.
63. The retrovirus of claim 61, wherein the retroviral envelope is an amphotropic MLV envelope.
64. The retrovirus according to claim 62, wherein the polypeptide that converts a nontoxic prodrug in to a toxic drug is thymidine kinase, purine nucleoside phosphorylase (PNP), or cytosine deaminase
65. The retrovirus according to claim 61, wherein the heterologous nucleic acid sequence encodes a binding domain, an antibody, or antibody fragment.
66. The retrovirus of claim 61, wherein the retrovirus infects cancer cells of a cancer.
67. The retrovirus of claim 66, wherein the cancer is selected from the group consisting of lung cancer, colon-rectum cancer, breast cancer, prostate cancer, urinary tract cancer, uterine cancer, brain cancer, head and neck cancer, pancreatic cancer, melanoma, stomach cancer and ovarian cancer.
68. The retrovirus of claim 61, wherein the pol III promoter is an H1 or U6 promoter.
69. The retrovirus according to claim 61, wherein the inhibitory polynucleotide comprises an miRNA, RNAi or siRNA sequence.
70. A method of treating a cell proliferative disorder comprising administering the recombinant replication competent gammaretrovirus of claim 61 to a subject having a cell proliferative disorder under conditions such that the retrovirus infects cells with the disorder and contacting the subject with an anti-cancer agent or chemotherapeutic agent, wherein the retrovirus is administered from about 10.sup.3 to 10.sup.11 TU/g brain weight.
71. The method of claim 70, wherein the anti-cancer agent is selected from the group consisting of bevacizumab, pegaptanib, ranibizumab, sorafenib, sunitinib, AE-941, VEGF Trap, pazopanib, vandetanib, vatalanib, cediranib, fenretinide, squalamine, INGN-241, oral tetrathiomolybdate, tetrathiomolybdate, Panzem NCD, 2-methoxyestradiol, AEE-788, AG-013958, bevasiranib sodium, AMG-706, axitinib, BIBF-1120, CDP-791, CP-547632, PI-88, SU-14813, SU-6668, XL-647, XL-999, IMC-1121 B, ABT-869, BAY-57-9352, BAY-73-4506, BMS-582664, CEP-7055, CHIR-265, CT-322, CX-3542, E-7080, ENMD-1198, OSI-930, PTC-299, Sirna-027, TKI-258, Veglin, XL-184, or ZK-304709.
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. patent application Ser. No. 15/672,029, filed Aug. 8, 2017, which is a continuation of U.S. patent application Ser. No. 14/477,741, filed Sep. 4, 2014 (now U.S. Pat. No. 9,732,326), which is a continuation of U.S. patent application Ser. No. 13/072,704, filed Mar. 26, 2011 (now U.S. Pat. No. 8,829,173), which is a continuation-in-part of International Application No. PCT/US09/58512, filed Sep. 26, 2009, which claims priority to U.S. Provisional Application Ser. No. 61/100,666, filed Sep. 26, 2008, U.S. Provisional Application Ser. No. 61/120,618, filed Dec. 8, 2008, U.S. Provisional Application Ser. No. 61/186,823, filed Jun. 13, 2009, and U.S. Provisional Application Ser. No. 61/318,728, filed Mar. 29, 2010, the disclosures of which are incorporated herein by reference.
TECHNICAL FIELD
[0002] This disclosure relates to replication competent retroviral vectors for treating cell proliferative. The disclosure further relates to the use of such replication competent retroviral vectors for delivery and expression of heterologous nucleic acids.
BACKGROUND
[0003] Effective methods of delivering genes and heterologous nucleic acids to cells and subjects has been a goal researchers for scientific development and for possible treatments of diseases and disorders.
SUMMARY
[0004] The disclosure provides a recombinant replication competent retrovirus (RCR) comprising: a retroviral GAG protein; a retroviral POL protein; a retroviral envelope; a retroviral polynucleotide comprising Long-Terminal Repeat (LTR) sequences at the 3' end of the retroviral polynucleotide sequence, a promoter sequence at the 5' end of the retroviral polynucleotide, said promoter being suitable for expression in a mammalian cell, a gag nucleic acid domain, a pol nucleic acid domain and an env nucleic acid domain; a cassette comprising an internal ribosome entry site (IRES) operably linked to a heterologous polynucleotide, wherein the cassette is positioned 5' to the 3' LTR and 3' to the env nucleic acid domain encoding the retroviral envelope; and cis-acting sequences necessary for reverse transcription, packaging and integration in a target cell, wherein the RCR maintains higher replication competency after 6 passages compared to a pACE vector (SEQ ID NO:21). In one embodiment, the retroviral polynucleotide sequence is derived from murine leukemia virus (MLV), Moloney murine leukemia virus (MoMLV), Feline leukemia virus (FeLV) or Gibbon ape leukemia virus (GALV). In another embodiment, the MLV is an amphotropic MLV. In yet another embodiment, the retrovirus is an oncoretrovirus or gamma retrovirus. In yet another embodiment, the target cell is a cell having a cell proliferative disorder. The cell proliferative disorder can be selected from the group consisting of, but is not limited to, lung cancer, colon-rectum cancer, breast cancer, prostate cancer, urinary tract cancer, uterine cancer, brain cancer, head and neck cancer, pancreatic cancer, melanoma, stomach cancer and ovarian cancer, rheumatoid arthritis and other autoimmune diseases. In one embodiment, the promoter comprises a CMV promoter having a sequence as set forth in SEQ ID NO:19, 20 or 22 from nucleotide 1 to about nucleotide 582 and may include modification to one or more nucleic acid bases and which is capable of directing and initiating transcription. In yet a further embodiment, the promoter comprises a sequence as set forth in SEQ ID NO: 19, 20 or 22 from nucleotide 1 to about nucleotide 582. In a further embodiment, the promoter comprises a CMV-R-U5 domain polynucleotide. In one embodiment, the CMV-R-U5 domain comprises the immediately early promoter from human cytomegalovirus linked to an MLV R-U5 region. In yet another embodiment, the CMV-R-U5 domain polynucleotide comprises a sequence as set forth in SEQ ID NO: 19, 20 or 22 from about nucleotide 1 to about nucleotide 1202 or sequences that are at least 95% identical to a sequence as set forth in SEQ ID NO: 19, 20 or 22, wherein the polynucleotide promotes transcription of a nucleic acid molecule operably linked thereto. In another embodiment, the gag and pol of the polynucleotide are derived from an oncoretrovirus or gamma retrovirus. The gag nucleic acid domain can comprise a sequence from about nucleotide number 1203 to about nucleotide 2819 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, 99% or 99.8% identity thereto. The pol domain can comprise a sequence from about nucleotide number 2820 to about nucleotide 6358 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, 99% or 99.9% identity thereto. In one embodiment, the env domain encodes an amphotropic env protein. The env domain can comprise a sequence from about nucleotide number 6359 to about nucleotide 8323 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, 99% or 99.8% identity thereto. The IRES domain of the vector can be any IRES, however, in one embodiment the IRES is derived from an encephalomyocarditis virus. In a further embodiment, the IRES comprises a sequence from about nucleotide number 8327 to about nucleotide 8876 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, or 99% identity thereto.
[0005] The vector can comprise any number of different heterologous polynucleotides. For example, the heterologous polynucleotide can comprise a cytokine, an siRNA, miRNA or RNAi molecules, a targeting sequence, a binding domain, a cytotoxic gene, a single chain antibody or any combination thereof. When the heterologous polynucleotide is for a non-translated RNA such as siRNA, miRNA or RNAi then no IRES is necessary, but may be included for another translated gene RNA, and any kind of retrovirus can be used. In yet a further embodiment, the heterologous polynucleotide comprises a polynucleotide having a sequence as set forth in SEQ ID NO: 3, 5, 11, 13, 15 or 17. In a further embodiment, the heterologous sequence encodes a polypeptide comprising a sequence as set forth in SEQ ID NO: 4. The heterologous nucleic acid is human codon optimized and encodes a polypeptide as set forth in SEQ ID NO:4. In a further embodiment, the heterologous nucleic acid comprises a sequence as set forth in SEQ ID NO: 19 or 22 from about nucleotide number 8877 to about 9353. In one embodiment, the 3' LTR is derived from an oncoretrovirus or gamma-retrovirus. In a further embodiment, the 3' LTR comprises a U3-R-U5 domain. In yet a further embodiment, the 3' LTR comprises a sequence as set forth in SEQ ID NO: 19 from about nucleotide 9405 to about 9998 or a sequence that is at least 95%, 98% or 99.5% identical thereto.
[0006] The disclosure provides a polynucleotide comprising a sequence as set forth in SEQ ID NO: 19, 20 or 22.
[0007] The disclosure provides an isolated polynucleotide comprising from 5' to 3': a CMV-R-U5 fusion of the immediate early promoter from human cytomegalovirus to an MLV R-U5 region; a PBS, primer binding site for reverse transcriptase; a 5' splice site; 4 packaging signal; a gag coding sequence for MLV group specific antigen; a pol coding sequence for MLV polymerase polyprotein; a 3' splice site; a 4070A env coding sequence for envelope protein of MLV strain 4070A; an internal ribosome entry site (IRES) from encephalomyocarditis virus; a modified cytosine deaminase coding sequence; a polypurine tract; and a U3-R-U5 MLV long terminal repeat.
[0008] The disclosure provides a method of treating a subject with a cell proliferative disorder comprising contacting the subject with a polynucleotide encoding a polypeptide of the disclosure having cytosine deaminase activity under conditions such that the polynucleotide is expressed, and contacting the subject with 5-fluorocytosine.
[0009] The disclosure also provides a method of treating a cell proliferative disorder in a subject comprising contacting the subject with a retrovirus of the disclosure, wherein the heterologous nucleic acid sequence encodes a therapeutic protein that inhibits proliferation of a neoplastic cell. In one embodiment, the retrovirus comprises a polynucleotide encoding a polypeptide having a sequence as set forth in SEQ ID NO: 4, 12, 14, 16, or 18.
[0010] The disclosure provides a recombinant replication competent retrovirus (RCR) comprising recombinant replication competent retrovirus, wherein the vector infects the target multiple times leading to a mean of 5 or more copies of the retrovirus genome. The multiple copies provide a "super" infection useful for gene delivery and protein production in vivo and in vitro. In one embodiment, the recombinant replication competent retrovirus (RCR) comprises: a retroviral GAG protein; a retroviral POL protein; a retroviral envelope; a retroviral polynucleotide comprising Long-Terminal Repeat (LTR) sequences at the 3' end of the retroviral polynucleotide sequence, a promoter sequence at the 5' end of the retroviral polynucleotide, said promoter being suitable for expression in a mammalian cell, a gag nucleic acid domain, a pol nucleic acid domain and an env nucleic acid domain; a cassette comprising an internal ribosome entry site (IRES) operably linked to a heterologous polynucleotide, wherein the cassette is positioned 5' to the 3' LTR and 3' to the env nucleic acid domain encoding the retroviral envelope; and cis-acting sequences necessary for reverse transcription, packaging and integration in a target cell, wherein the RCR maintains higher replication competency after 6 passages compared to a pACE vector (SEQ ID NO:21). In one embodiment, the retroviral polynucleotide sequence is derived from murine leukemia virus (MLV), Moloney murine leukemia virus (MoMLV), Feline leukemia virus (FeLV), Baboon endogenous retrovirus (BEV), porcine endogenous virus (PERV), the cat derived retrovirus RD114, squirrel monkey retrovirus, Xenotropic murine leukemia virus-related virus (XMRV), avian reticuloendotheliosis virus (REV), or Gibbon ape leukemia virus (GALV). In another embodiment, the MLV is an amphotropic MLV. In yet another embodiment, the retrovirus is an oncoretrovirus or gamma retrovirus. In yet another embodiment, the target cell is a cell having a cell proliferative disorder. The cell proliferative disorder can be selected from the group consisting of, but is not limited to, lung cancer, colon-rectum cancer, breast cancer, prostate cancer, urinary tract cancer, uterine cancer, brain cancer, head and neck cancer, pancreatic cancer, melanoma, stomach cancer and ovarian cancer, rheumatoid arthritis and other autoimmune diseases. In one embodiment, the promoter comprises a CMV promoter having a sequence as set forth in SEQ ID NO:19, 20 or 22 from nucleotide 1 to about nucleotide 582 and may include modification to one or more nucleic acid bases and which is capable of directing and initiating transcription. In yet a further embodiment, the promoter comprises a sequence as set forth in SEQ ID NO: 19, 20 or 22 from nucleotide 1 to about nucleotide 582. In a further embodiment, the promoter comprises a CMV-R-U5 domain polynucleotide. In one embodiment, the CMV-R-U5 domain comprise the immediately early promoter from human cytomegalovirus linked to an MLV R-U5 region. In yet another embodiment, the CMV-R-U5 domain polynucleotide comprises a sequence as set forth in SEQ ID NO: 19, 20 or 22 from about nucleotide 1 to about nucleotide 1202 or sequences that are at least 95% identical to a sequence as set forth in SEQ ID NO: 19, 20 or 22, wherein the polynucleotide promotes transcription of a nucleic acid molecule operably linked thereto. In another embodiment, the gag and pol of the polynucleotide are derived from an oncoretrovirus or gamma retrovirus. The gag nucleic acid domain can comprise a sequence from about nucleotide number 1203 to about nucleotide 2819 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, 99% or 99.8% identity thereto. The pol domain can comprise a sequence from about nucleotide number 2820 to about nucleotide 6358 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, 99% or 99.9% identity thereto. In one embodiment, the env domain encodes an amphotropic env protein. The env domain can comprise a sequence from about nucleotide number 6359 to about nucleotide 8323 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, 99% or 99.8% identity thereto. The IRES domain of the vector can be any IRES, however, in one embodiment the IRES is derived from an encephalomyocarditis virus. In a further embodiment, the IRES comprises a sequence from about nucleotide number 8327 to about nucleotide 8876 of SEQ ID NO: 19 or 22 or a sequence having at least 95%, 98%, or 99% identity thereto.
[0011] The disclosure provides a method of treating a cell proliferative disorder in a subject comprising contacting the subject with a retrovirus of the disclosure wherein the vector infects the target multiple times leading to a mean of 5 or more copies of the retrovirus genome.
[0012] The details of one or more embodiments of the disclosure are set forth in the accompanying drawings and the description below. Other features, objects, and advantages will be apparent from the description and drawings, and from the claims.
BRIEF DESCRIPTION OF DRAWINGS
[0013] FIG. 1A-C shows an alignment of the Wild-type yeast cytosine deaminase (SEQ ID NO: 2) and a cytosine deaminase of the disclosure (SEQ ID NO: 4) and other sequences of the disclosure (SEQ ID NOs:31-40).
[0014] FIG. 2 shows a graph of cell killing data showing that modified vectors are more effective compared to the original wild type CD. The graph also shows that the new modified backbone (T5.0007) is more effective at killing than the old backbone (pACE-CD). Also shown is a table cataloguing the various vector constructs and their names.
[0015] FIG. 3A-F shows (A) a schematic of a recombinant retroviral vector of the disclosure; (B-C) a plasmid map of a polynucleotide of the disclosure; (D) a sequence of a polynucleotide of the disclosure (SEQ ID NO:19); the vector coding region of pAC3-yCD2 in various formats (i.e., FIG. 3D-1 to 3D-2 shows the domains of the vector and FIG. 3D-4 to FIG. 3D-16 shows restriction sites in the vector of SEQ ID NO:19); (E) Diagram of changes between pACE-emdGFP and pAC3-emdGFP; (F) shows additional plasmid map details of SEQ ID NO:19 (FIG. 3F-1 and FIG. 3F-2) and FIG. 3F-3 to FIG. 3F-5) shows the sequence of SEQ ID NO:22 including identifying various domains; and (FIG. 3F-6 to FIG. 3F-18) shows restriction sites in the vector of SEQ ID NO:22.
[0016] FIG. 4 shows that higher levels of yCD2 protein are observed compared to wild type yCD protein in infected U-87 cells.
[0017] FIG. 5 shows that a vector of the disclosure is genetically stable after 12 cycles of viral passages as assessed using PCR amplification. The figure also demonstrates that the vectors of the disclosure are more stable after longer passages compared to the vector pACE-CD (Kasahara et al.). In particular pAC3-CD is more stable than pACE-CD, demonstrating that the changed backbone has made the vector more stable. In addition pACE-yCD1 (T5.0001) and -yCD2 (T5-0002) are very much more stable than pAC-yCD, demonstrating that small and silent changes to the coding sequence of the transgene can have a very large effect on stability, leading to superior properties.
[0018] FIG. 6A-B shows (A) cell killing assays; and (B) cytosine deaminase specific activity of cells infected with different vectors. (A) shows that cytosine deaminase and vector of the disclosure kill infected cells at least as well and perhaps better than the original pACE-CD when U87 infected cells are exposed to increasing levels of 5-FC. (B) Shows that the specific CD activity of the disclosure (T5.0007, T5.0001 and T5.0002) are all increased compared to pACE-CD (T5.0000), and is in the order T5.0000<T5.0007<T5.0001<T5.0002.
[0019] FIG. 7 shows U-87 tumors treated with CD vector of the disclosure in vivo and explanted from mice treated with 4 cycles of 5-FC are still sensitive to the drug.
[0020] FIG. 8 shows dosing information and therapeutic effect in a Kaplan-Meyer survival analysis in a mouse model of brain cancer.
[0021] FIG. 9 shows dosing information and therapeutic effect in a Kaplan-Meyer survival analysis in a syngeneic mouse model.
[0022] FIG. 10A-D shows schemes for the generation of various embodiments of the disclosure comprising polypeptide with CD, OPRT and UPRT activity.
[0023] FIG. 11A-E shows vector maps and miRNA results. A. is a schematic vector map of the MLV retroviral vector pAC3 backbone containing polynucleotide sequences of human primary precursor miR-128-1, human primary precursor miR-128-2 and human precursor miR-128 linked to a human H1 promoter, designated pAC3-miR128-1, pAC3-miR-128-2, and pAC3-H1-shRNAmiR128, respectively. B. is a schematic vector map of the MLV retroviral vector pAC3-yCD2 backbone containing polynucleotide sequences of a human precursor miR-128 linked to a human H1 promoter, designated pAC3-yCD2-H1-shRNAmiR128. C. shows a schematic vector map of the MLV retroviral vector pAC3 backbone containing polynucleotide sequences of human primary precursor miR-142-3pT. D. shows sequences for 142-3p (SEQ ID NOs: 35 and 36) and primers (SEQ ID NO:41 and 42). E. shows results from transformation with a vector containing miR-142-3pT.
[0024] FIG. 12A-B: A. shows a comparison of replication kinetics of miR-128 containing vectors (pAC3-miR-128-1, pAC3-miR-128-2, and pAC3-H1-shRNAmiR128) in human fibrosarcoma cells HT1080 analyzed by qPCR. The graph is generated by plotting of inversed C(t) values obtained from qPCR vs. various time points during viral replication. B. shows a comparison of replication kinetics of miR-128 containing vectors (pAC3-miR-128-1, pAC3-miR-128-2, and pAC3-H1-shRNAmiR128) in human glioma cells U87-MG analyzed by qPCR. The graph is generated by plotting of inversed C(t) values obtained from qPCR vs. various time points during viral replication.
[0025] FIG. 13 shows a relative quantification of mature miR-128 expression from cells transduced with miR-128 containing vectors.
[0026] FIG. 14 shows a relative quantification of Bmi-1 gene expression from cells transduced with miR-128 containing vectors.
[0027] FIG. 15 is a schematic vector map of the MLV retroviral vector pAC3-emd containing a single copy of 142-3pT target sequence, designated pAC3-emd-142-3pT and 4 tandem repeats of 142-3pT, designated pAC3-emd-142-3pT4X.
[0028] FIG. 16 is a schematic vector map of the MLV retroviral vector pAC3-yCD2 containing a single copy of 142-3pT target sequence, designated pAC3-yCD2-142-3pT and 4 tandem repeats of 142-3pT, designated pAC3-yCD2-142-3pT4X.
[0029] FIG. 17A-B: A. shows a comparison of replication kinetics of 142-3pT containing vectors (pAC3-emd-142-3pT pAC3-emd-142-3pT4X, pAC3-yCD2-142-3pT and pAC3-yCD2-142-3pT4X) and their parental vectors (pAC3-emd and pAC3-yCD2) in human fibrosarcoma cells HT1080 analyzed qPCR. The graph is generated by plotting by inversed C(t) values obtained from qPCR vs. various time points during viral replication. B. shows a comparison of replication kinetics of GFP containing vectors (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) in human fibrosarcoma cells HT1080 analyzed by flow cytometric analysis of GFP expression at various time points during vector spread.
[0030] FIG. 18A-B: A. shows a comparison of replication kinetics of 142-3pT containing vectors (pAC3-emd-142-3pT pAC3-emd-142-3pT4X, pAC3-yCD2-142-3pT and pAC3-yCD2-142-3pT4X) and their parental vectors (pAC3-emd and pAC3-yCD2) in human glioma cells U87-MG analyzed qPCR. The graph is generated by plotting by inversed C(t) values obtained from qPCR vs. various time points during viral replication. B. shows a comparison of replication kinetics of GFP containing vectors (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) in human glioma cells U87-MG analyzed by flow cytometric analysis of GFP expression at various time points during vector spread.
[0031] FIG. 19 shows the replication kinetics of GFP containing vector (pAC3-emd) in mouse and human hematopoietic cells analyzed by flow cytometric analysis of GFP expression at various time points during vector spread.
[0032] FIG. 20A-C shows comparison of replication kinetics. A. shows a comparison of replication kinetics of GFP containing vectors (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) in mouse T-lymphocytes EL4 analyzed by flow cytometric analysis of GFP expression at various time points during vector spread. B. shows a comparison of replication kinetics of GFP containing vectors (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) in human T-lymphocytes SUP-T1 analyzed by flow cytometric analysis of GFP expression at various time points during vector spread. C. shows a comparison of replication kinetics of GFP containing vectors (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) in human monocytes U937 analyzed by flow cytometric analysis of GFP expression at various time points during vector spread.
[0033] FIGS. 21A-B are still frames from the MRI images obtained from the patient dog during intratumoral CED infusion of Toca 511 and gadolinium. Note the large tumor on the left side of the image compressing both sides of the brain and shifting midline structures to the right. The white areas are the gadolinium-Toca 511 infusion. FIG. 21B shows the placement of the two catheters into the tumor.
[0034] FIG. 22 is a schematic vector map of the MLV retroviral vectors encoding the human IFN-gamma (hIFNg) and mouse IFN-gamma (mIFNg), respectively, in pAC3 backbone.
[0035] FIG. 23 shows the expression of mIFN-gamma at the RNA level from human fibrosarcoma cell line HT1080 infected with pAC3-mIFNg vector. Expression is detected by RT-PCR.
[0036] FIG. 24 shows the expression of hIFN-gamma protein secreted from human fibrosarcoma cell line HT1080 infected with pAC3-hIFNg vector.
[0037] FIG. 25 shows the expression of mIFN-gamma protein secreted from human fibrosarcoma cell line HT1080 infected with pAC3-mIFNg vector.
[0038] FIG. 26 shows flow cytometry analysis of GFP expression in U87 cells after intratumor or intravenous delivery of AC3-GFP vector in a nude mouse model. Cells are measured by flow cytometry for percent GFP positive. Cells isolated from naive nude mouse brains, U87 cells from tissue culture, or U87 cells transduced at an multiplicity of infection of 1 with AC3-GFP(V) in vitro serve as controls. From example 27 (iv injection of GFP vector).
[0039] FIG. 27 shows a histogram analysis was also done on groups 1, 3 and 5 from example 27 (iv injection of GFP vector) to measure the distribution of GFP signal in isolated U87 cells. GFP expression is from U87 tumor cells isolated from mouse brains after 14 days after vector treatment.
[0040] FIG. 28 is a schematic vector map of the MLV retroviral vectors encoding the human IL-2 in the pAC3 backbone.
DETAILED DESCRIPTION
[0041] As used herein and in the appended claims, the singular forms "a," "and," and "the" include plural referents unless the context clearly dictates otherwise. Thus, for example, reference to "a cell" includes a plurality of such cells and reference to "the agent" includes reference to one or more agents known to those skilled in the art, and so forth.
[0042] Also, the use of "or" means "and/or" unless stated otherwise. Similarly, "comprise," "comprises," "comprising" "include," "includes," and "including" are interchangeable and not intended to be limiting.
[0043] It is to be further understood that where descriptions of various embodiments use the term "comprising," those skilled in the art would understand that in some specific instances, an embodiment can be alternatively described using language "consisting essentially of" or "consisting of."
[0044] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood to one of ordinary skill in the art to which this disclosure belongs. Although methods and materials similar or equivalent to those described herein can be used in the practice of the disclosed methods and compositions, the exemplary methods, devices and materials are described herein.
[0045] General texts which describe molecular biological techniques useful herein, including the use of vectors, promoters and many other relevant topics, include Berger and Kimmel, Guide to Molecular Cloning Techniques, Methods in Enzymology Volume 152, (Academic Press, Inc., San Diego, Calif.) ("Berger"); Sambrook et al., Molecular Cloning--A Laboratory Manual, 2d ed., Vol. 1-3, Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., 1989 ("Sambrook") and Current Protocols in Molecular Biology, F. M. Ausubel et al., eds., Current Protocols, a joint venture between Greene Publishing Associates, Inc. and John Wiley & Sons, Inc., (supplemented through 1999) ("Ausubel"). Examples of protocols sufficient to direct persons of skill through in vitro amplification methods, including the polymerase chain reaction (PCR), the ligase chain reaction (LCR), Q.beta.-replicase amplification and other RNA polymerase mediated techniques (e.g., NASBA), e.g., for the production of the homologous nucleic acids of the disclosure are found in Berger, Sambrook, and Ausubel, as well as in Mullis et al. (1987) U.S. Pat. No. 4,683,202; Innis et al., eds. (1990) PCR Protocols: A Guide to Methods and Applications (Academic Press Inc. San Diego, Calif.) ("Innis"); Arnheim & Levinson (Oct. 1, 1990) C&EN 36-47; The Journal Of NIH Research (1991) 3: 81-94; Kwoh et al. (1989) Proc. Natl. Acad. Sci. USA 86: 1173; Guatelli et al. (1990) Proc. Nat'l. Acad. Sci. USA 87: 1874; Lomell et al. (1989) J. Clin. Chem 35: 1826; Landegren et al. (1988) Science 241: 1077-1080; Van Brunt (1990) Biotechnology 8: 291-294; Wu and Wallace (1989) Gene 4:560; Barringer et al. (1990) Gene 89:117; and Sooknanan and Malek (1995) Biotechnology 13: 563-564. Improved methods for cloning in vitro amplified nucleic acids are described in Wallace et al., U.S. Pat. No. 5,426,039. Improved methods for amplifying large nucleic acids by PCR are summarized in Cheng et al. (1994) Nature 369: 684-685 and the references cited therein, in which PCR amplicons of up to 40 kb are generated. One of skill will appreciate that essentially any RNA can be converted into a double stranded DNA suitable for restriction digestion, PCR expansion and sequencing using reverse transcriptase and a polymerase. See, e.g., Ausubel, Sambrook and Berger, all supra.
[0046] The publications discussed throughout the text are provided solely for their disclosure prior to the filing date of the present application. Nothing herein is to be construed as an admission that the inventors are not entitled to antedate such disclosure by virtue of prior disclosure.
[0047] The disclosure provides methods and compositions useful for gene or protein delivery to a cell or subject. Such methods and compositions can be used to treat various diseases and disorders in a subject including cancer and other cell proliferative diseases and disorders. The disclosure provides replication competent retroviral vectors for gene delivery.
[0048] The terms "vector", "vector construct" and "expression vector" mean the vehicle by which a DNA or RNA sequence (e.g. a foreign gene) can be introduced into a host cell, so as to transform the host and promote expression (e.g. transcription and translation) of the introduced sequence. Vectors typically comprise the DNA of a transmissible agent, into which foreign DNA encoding a protein is inserted by restriction enzyme technology. A common type of vector is a "plasmid", which generally is a self-contained molecule of double-stranded DNA that can readily accept additional (foreign) DNA and which can readily introduced into a suitable host cell. A large number of vectors, including plasmid and fungal vectors, have been described for replication and/or expression in a variety of eukaryotic and prokaryotic hosts. Non-limiting examples include pKK plasmids (Clonetech), pUC plasmids, pET plasmids (Novagen, Inc., Madison, Wis.), pRSET or pREP plasmids (Invitrogen, San Diego, Calif.), or pMAL plasmids (New England Biolabs, Beverly, Mass.), and many appropriate host cells, using methods disclosed or cited herein or otherwise known to those skilled in the relevant art. Recombinant cloning vectors will often include one or more replication systems for cloning or expression, one or more markers for selection in the host, e.g., antibiotic resistance, and one or more expression cassettes.
[0049] The terms "express" and "expression" mean allowing or causing the information in a gene or DNA sequence to become manifest, for example producing a protein by activating the cellular functions involved in transcription and translation of a corresponding gene or DNA sequence. A DNA sequence is expressed in or by a cell to form an "expression product" such as a protein. The expression product itself, e.g. the resulting protein, may also be said to be "expressed" by the cell. A polynucleotide or polypeptide is expressed recombinantly, for example, when it is expressed or produced in a foreign host cell under the control of a foreign or native promoter, or in a native host cell under the control of a foreign promoter.
[0050] The disclosure provides replication competent viral vectors the contain a heterologous polynucleotide encoding, for example, a cytosine deaminase or mutant thereof, an miRNA or siRNA, a cytokine, an antibody binding domain etc., that can be delivered to a cell or subject. The viral vector can be an adenoviral vector, a measles vector, a herpes vector, a retroviral vector (including a lentiviral vector), a rhabdoviral vector such as a Vesicular Stomatitis viral vector, a reovirus vector, a Seneca Valley Virus vector, a poxvirus vector (including animal pox or vaccinia derived vectors), a parvovirus vector (including an AAV vector), an alphavirus vector or other viral vector known to one skilled in the art (see also, e.g., Concepts in Genetic Medicine, ed. Boro Dropulic and Barrie Carter, Wiley, 2008, Hoboken, N.J.; The Development of Human Gene Therapy, ed. Theodore Friedmann, Cold Springs Harbor Laboratory Press, Cold springs Harbor, N.Y., 1999; Gene and Cell Therapy, ed. Nancy Smyth Templeton, Marcel Dekker Inc., New York, N.Y., 2000 and Gene Therapy: Therapeutic Mechanism and Strategies, ed. Nancy Smyth Templetone and Danilo D Lasic, Marcel Dekker, Inc., New York, N.Y., 2000; the disclosures of which are incorporated herein by reference).
[0051] In one embodiment, the viral vector can be a replication competent retroviral vector capable of infecting only replicating mammalian cells. In one embodiment, a replication competent retroviral vector comprises an internal ribosomal entry site (IRES) 5' to the heterologous polynucleotide encoding, e.g., a cytosine deaminase, miRNA, siRNA, cytokine, receptor, antibody or the like. When the heterologous polynucleotide encodes a non-translated RNA such as siRNA, miRNA or RNAi then no IRES is necessary, but may be included for another translated gene, and any kind of retrovirus (see below) can be used. In one embodiment, the polynucleotide is 3' to a ENV polynucleotide of a retroviral vector. In one embodiment the viral vector is a retroviral vector capable of infecting target cells multiple times (5 or more per diploid cell).
[0052] In other embodiments, host cells transfected with a replication competent retroviral vector of the disclosure are provided. Host cells include eukaryotic cells such as yeast cells, insect cells, or animal cells. Host cells also include prokaryotic cells such as bacterial cells.
[0053] Also provided are engineered host cells that are transduced (transformed or transfected) with a vector provided herein (e.g., a replication competent retroviral vector). The engineered host cells can be cultured in conventional nutrient media modified as appropriate for activating promoters, selecting transformants, or amplifying a coding polynucleotide. Culture conditions, such as temperature, pH and the like, are those previously used with the host cell selected for expression, and will be apparent to those skilled in the art and in the references cited herein, including, e.g., Sambrook, Ausubel and Berger, as well as e.g., Freshney (1994) Culture of Animal Cells: A Manual of Basic Technique, 3rd ed. (Wiley-Liss, New York) and the references cited therein.
[0054] Examples of appropriate expression hosts include: bacterial cells, such as E. coli, B. subtilis, Streptomyces, and Salmonella typhimurium; fungal cells, such as Saccharomyces cerevisiae, Pichia pastoris, and Neurospora crassa; insect cells such as Drosophila and Spodoptera frugiperda; mammalian cells such as CHO, COS, BHK, HEK 293 br Bowes melanoma; or plant cells or explants, etc. Typically human cells or cell lines will be used; however, it may be desirable to clone vectors and polynucleotides of the disclosure into non-human host cells for purposes of sequencing, amplification and cloning.
[0055] The disclosure also provides replication competent retroviral vectors having increased stability relative to prior retroviral vectors. Such increased stability during infection and replication is important for the treatment of cell proliferative disorders. The combination of transduction efficiency, transgene stability and target selectivity is provided by the replication competent retrovirus. The compositions and methods provide insert stability and maintain transcription activity of the transgene and the translational viability of the encoded polypeptide.
[0056] The disclosure provides modified retroviral vectors. The modified retroviral vectors can be derived from members of the retroviridae family. The Retroviridae family consists of three groups: the spumaviruses-(or foamy viruses) such as the human foamy virus (HFV); the lentiviruses, as well as visna virus of sheep; and the oncoviruses (although not all viruses within this group are oncogenic). The term "lentivirus" is used in its conventional sense to describe a genus of viruses containing reverse transcriptase. The lentiviruses include the "immunodeficiency viruses" which include human immunodeficiency virus (HIV) type 1 and type 2 (HIV-1 and HIV-2) and simian immunodeficiency virus (SIV). The oncoviruses have historically been further subdivided into groups A, B, C and D on the basis of particle morphology, as seen under the electron microscope during viral maturation. A-type particles represent the immature particles of the B- and D-type viruses seen in the cytoplasm of infected cells. These particles are not infectious. B-type particles bud as mature virion from the plasma membrane by the enveloping of intracytoplasmic A-type particles. At the membrane they possess a toroidal core of 75 nm, from which long glycoprotein spikes project. After budding, B-type particles contain an eccentrically located, electron-dense core. The prototype B-type virus is mouse mammary tumor virus (MMTV). No intracytoplasmic particles can be observed in cells infected by C-type viruses. Instead, mature particles bud directly from the cell surface via a crescent `C`-shaped condensation which then closes on itself and is enclosed by the plasma membrane. Envelope glycoprotein spikes may be visible, along with a uniformly electron-dense core. Budding may occur from the surface plasma membrane or directly into intracellular vacuoles. The C-type viruses are the most commonly studied and include many of the avian and murine leukemia viruses (MLV). Bovine leukemia virus (BLV), and the human T-cell leukemia viruses types I and II (HTLV-I/II) are similarly classified as C-type particles because of the morphology of their budding from the cell surface. However, they also have a regular hexagonal morphology and more complex genome structures than the prototypic C-type viruses such as the murine leukemia viruses (MLV). D-type particles resemble B-type particles in that they show as ring-like structures in the infected cell cytoplasm, which bud from the cell surface, but the virion incorporate short surface glycoprotein spikes. The electron-dense cores are also eccentrically located within the particles. Mason Pfizer monkey virus (MPMV) is the prototype D-type virus.
[0057] Retroviruses have been classified in various ways but the nomenclature has been standardized in the last decade (see ICTVdB--The Universal Virus Database, v 4 on the World Wide Web (www) at ncbi.nlm.nih.gov/ICTVdb/ICTVdB/ and the text book "Retroviruses" Eds Coffin, Hughs and Varmus, Cold Spring Harbor Press 1997; the disclosures of which are incorporated herein by reference). In one embodiment, the replication competent retroviral vector can comprise an Orthoretrovirus or more typically a gamma retrovirus vector.
[0058] Retroviruses are defined by the way in which they replicate their genetic material. During replication the RNA is converted into DNA. Following infection of the cell a double-stranded molecule of DNA is generated from the two molecules of RNA which are carried in the viral particle by the molecular process known as reverse transcription. The DNA form becomes covalently integrated in the host cell genome as a provirus, from which viral RNAs are expressed with the aid of cellular and/or viral factors. The expressed viral RNAs are packaged into particles and released as infectious virion.
[0059] The retrovirus particle is composed of two identical RNA molecules. Each wild-type genome has a positive sense, single-stranded RNA molecule, which is capped at the 5' end and polyadenylated at the 3' tail. The diploid virus particle contains the two RNA strands complexed with gag proteins, viral enzymes (pol gene products) and host tRNA molecules within a `core` structure of gag proteins. Surrounding and protecting this capsid is a lipid bilayer, derived from host cell membranes and containing viral envelope (env) proteins. The env proteins bind to a cellular receptor for the virus and the particle typically enters the host cell via receptor-mediated endocytosis and/or membrane fusion.
[0060] After the outer envelope is shed, the viral RNA is copied into DNA by reverse transcription. This is catalyzed by the reverse transcriptase enzyme encoded by the pol region and uses the host cell tRNA packaged into the virion as a primer for DNA synthesis. In this way the RNA genome is converted into the more complex DNA genome.
[0061] The double-stranded linear DNA produced by reverse transcription may, or may not, have to be circularized in the nucleus. The provirus now has two identical repeats at either end, known as the long terminal repeats (LTR). The termini of the two LTR sequences produces the site recognized by a pol product--the integrase protein--which catalyzes integration, such that the provirus is always joined to host DNA two base pairs (bp) from the ends of the LTRs. A duplication of cellular sequences is seen at the ends of both LTRs, reminiscent of the integration pattern of transposable genetic elements. Retroviruses can integrate their DNAs at many sites in host DNA, but different retroviruses have different integration site preferences. HIV-1 and simian immunodeficiency virus DNAs preferentially integrate into expressed genes, murine leukemia virus (MLV) DNA preferentially integrates near transcriptional start sites (TSSs), and avian sarcoma leukosis virus (ASLV) and human T cell leukemia virus (HTLV) DNAs integrate nearly randomly, showing a slight preference for genes (Derse D, et al. (2007) Human T-cell leukemia virus type 1 integration target sites in the human genome: comparison with those of other retroviruses. J Virol 81:6731-6741; Lewinski M K, et al. (2006) Retroviral DNA integration: viral and cellular determinants of target-site selection. PLoS Pathog 2:e601).
[0062] Transcription, RNA splicing and translation of the integrated viral DNA is mediated by host cell proteins. Variously spliced transcripts are generated. In the case of the human retroviruses HIV-1/2 and HTLV-I/II viral proteins are also used to regulate gene expression. The interplay between cellular and viral factors is a factor in the control of virus latency and the temporal sequence in which viral genes are expressed.
[0063] Retroviruses can be transmitted horizontally and vertically. Efficient infectious transmission of retroviruses requires the expression on the target cell of receptors which specifically recognize the viral envelope proteins, although viruses may use receptor-independent, nonspecific routes of entry at low efficiency. Normally a viral infection leads to a single or few copies of viral genome per cell because of receptor masking or down-regulation that in turn leads to resistance to superinfection (Ch3 p 104 in "Retroviruses" J M Coffin, S H Hughes, & H E Varmus 1997 Cold Spring Harbor Laboratory Press, Cold Spring Harbor N.Y.; Fan et al. J. Virol 28:802, 1978). By manipulating the situation in tissue culture it is possible to get some level of multiple infection but this is less than 5 copies/diploid genome. In addition, the target cell type must be able to support all stages of the replication cycle after virus has bound and penetrated. Vertical transmission occurs when the viral genome becomes integrated in the germ line of the host. The provirus will then be passed from generation to generation as though it were a cellular gene. Hence endogenous proviruses become established which frequently lie latent, but which can become activated when the host is exposed to appropriate agents.
[0064] In many situations for using a recombinant replication competent retrovirus therapeutically, it is advantageous to have high levels of expression of the transgene that is encoded by the recombinant replication competent retrovirus. For example, with a prodrug activating gene such as the cytosine deaminase gene it is advantageous to have higher levels of expression of the CD protein in a cell so that the conversion of the prodrug 5-FC to 5-FU is more efficient. Similarly high levels of expression of siRNA or shRNA lead to more efficient suppression of target gene expression. Also for cytokines or single chain antibodies (scAbs) it is usually advantageous to express high levels of the cytokine or scAb. In addition, in the case that there are mutations in some copies of the vector that inactivate or impair the activity of the vector or transgene, it is advantageous to have multiple copies of the vector in the target cell as this provides a high probability of efficient expression of the intact transgene. The disclosure provides recombinant replication competent retroviruses capable of infecting a target cell or target cell population multiple times resulting in an average number of copies/diploid genome of 5 or greater. The disclosure also provides methods of testing for this property. Also provided are methods of treating a cell proliferative disorder, using a recombinant replication competent retrovirus capable of infecting a target cell or target cell population multiple times resulting in an average number of copies/diploid genome of 5 or greater.
[0065] As mentioned above, the integrated DNA intermediate is referred to as a provirus. Prior gene therapy or gene delivery systems use methods and retroviruses that require transcription of the provirus and assembly into infectious virus while in the presence of an appropriate helper virus or in a cell line containing appropriate sequences enabling encapsidation without coincident production of a contaminating helper virus. As described below, a helper virus is not required for the production of the recombinant retrovirus of the disclosure, since the sequences for encapsidation are provided in the genome thus providing a replication competent retroviral vector for gene delivery or therapy.
[0066] Other existing replication competent retroviral vectors also tend to be unstable and lose sequences during horizontal or vertical transmission to an infected cell or host cell and during replication. This may be due in-part from the presence of extra nucleotide sequences that include repeats or which reduce the efficiency of a polymerase.
[0067] The retroviral genome and the proviral DNA of the disclosure have at least three genes: the gag, the pol, and the env, these genes may be flanked by one or two long terminal (LTR) repeat, or in the provirus are flanked by two long terminal repeat (LTR) and sequences containing cis-acting sequences such as psi. The gag gene encodes the internal structural (matrix, capsid, and nucleocapsid) proteins; the pol gene encodes the RNA-directed DNA polymerase (reverse transcriptase), protease and integrase; and the env gene encodes viral envelope glycoproteins. The 5' and/or 3' LTRs serve to promote transcription and polyadenylation of the virion RNAs. The LTR contains all other cis-acting sequences necessary for viral replication. Lentiviruses have additional genes including vif, vpr, tat, rev, vpu, nef, and vpx (in HIV-1, HIV-2 and/or SIV).
[0068] Adjacent to the 5' LTR are sequences necessary for reverse transcription of the genome (the tRNA primer binding site) and for efficient encapsidation of viral RNA into particles (the Psi site). If the sequences necessary for encapsidation (or packaging of retroviral RNA into infectious virion) are missing from the viral genome, the result is a cis defect which prevents encapsidation of genomic viral RNA. This type of modified vector is what has typically been used in prior gene delivery systems (i.e., systems lacking elements which are required for encapsidation of the virion) as `helper` elements providing viral proteins in trans that package a non-replicating, but packageable, RNA genome.
[0069] In a first embodiment, the disclosure provides a recombinant retrovirus capable of infecting a non-dividing cell, a dividing cell, or a cell having a cell proliferative disorder. The recombinant replication competent retrovirus of the disclosure comprises a polynucleotide sequence encoding a viral GAG, a viral POL, a viral ENV, a heterologous polynucleotide preceded by an internal ribosome entry site (IRES) encapsulated within a virion.
[0070] The phrase "non-dividing" cell refers to a cell that does not go through mitosis. Non-dividing cells may be blocked at any point in the cell cycle, (e.g., G.sub.0/G.sub.1, G.sub.1/S, G.sub.2/M), as long as the cell is not actively dividing. For ex vivo infection, a dividing cell can be treated to block cell division by standard techniques used by those of skill in the art, including, irradiation, aphidocolin treatment, serum starvation, and contact inhibition. However, it should be understood that ex vivo infection is often performed without blocking the cells since many cells are already arrested (e.g., stem cells). For example, a recombinant lentivirus vector is capable of infecting non-dividing cells. Examples of pre-existing non-dividing cells in the body include neuronal, muscle, liver, skin, heart, lung, and bone marrow cells, and their derivatives. For dividing cells onco-retroviral vectors can be used.
[0071] By "dividing" cell is meant a cell that undergoes active mitosis, or meiosis. Such dividing cells include stem cells, skin cells (e.g., fibroblasts and keratinocytes), gametes, and other dividing cells known in the art. Of particular interest and encompassed by the term dividing cell are cells having cell proliferative disorders, such as neoplastic cells. The term "cell proliferative disorder" refers to a condition characterized by an abnormal number of cells. The condition can include both hypertrophic (the continual multiplication of cells resulting in an overgrowth of a cell population within a tissue) and hypotrophic (a lack or deficiency of cells within a tissue) cell growth or an excessive influx or migration of cells into an area of a body. The cell populations are not necessarily transformed, tumorigenic or malignant cells, but can include normal cells as well. Cell proliferative disorders include disorders associated with an overgrowth of connective tissues, such as various fibrotic conditions, including scleroderma, arthritis and liver cirrhosis. Cell proliferative disorders include neoplastic disorders such as head and neck carcinomas. Head and neck carcinomas would include, for example, carcinoma of the mouth, esophagus, throat, larynx, thyroid gland, tongue, lips, salivary glands, nose, paranasal sinuses, nasopharynx, superior nasal vault and sinus tumors, esthesioneuroblastoma, squamous cell cancer, malignant melanoma, sinonasal undifferentiated carcinoma (SNUC), brain (including glioblastomas) or blood neoplasia. Also included are carcinoma's of the regional lymph nodes including cervical lymph nodes, prelaryngeal lymph nodes, pulmonary juxtaesophageal lymph nodes and submandibular lymph nodes (Harrison's Principles of Internal Medicine (eds., Isselbacher, et al., McGraw-Hill, Inc., 13th Edition, pp 1850-1853, 1994). Other cancer types, include, but are not limited to, lung cancer, colon-rectum cancer, breast cancer, prostate cancer, urinary tract cancer, uterine cancer lymphoma, oral cancer, pancreatic cancer, leukemia, melanoma, stomach cancer, skin cancer and ovarian cancer. The cell proliferative disease also includes rheumatoid arthritis (O'Dell NEJM 350:2591 2004) and other auto-immune disorders (Mackay et al NEJM 345:340 2001) that are often characterized by inappropriate proliferation of cells of the immune system.
[0072] The heterologous nucleic acid sequence is operably linked to an IRES. As used herein, the term "heterologous" nucleic acid sequence or transgene refers to (i) a sequence that does not normally exist in a wild-type retrovirus, (ii) a sequence that originates from a foreign species, or (iii) if from the same species, it may be substantially modified from its original form. Alternatively, an unchanged nucleic acid sequence that is not normally expressed in a cell is a heterologous nucleic acid sequence.
[0073] Depending upon the intended use of the retroviral vector of the disclosure any number of heterologous polynucleotide or nucleic acid sequences may be inserted into the retroviral vector. For example, for in vitro studies commonly used marker genes or reporter genes may be used, including, antibiotic resistance and fluorescent molecules (e.g., GFP). Additional polynucleotide sequences encoding any desired polypeptide sequence may also be inserted into the vector of the disclosure. Where in vivo delivery of a heterologous nucleic acid sequence is sought both therapeutic and non-therapeutic sequences may be used. For example, the heterologous sequence can encode a therapeutic molecule including antisense molecules (miRNA, siRNA) or ribozymes directed to a particular gene associated with a cell proliferative disorder or other gene-associated disease or disorder, the heterologous sequence can be a suicide gene (e.g., HSV-tk or PNP or cytosine deaminase; either modified or unmodified), a growth factor or a therapeutic protein (e.g., Factor IX, IL2, and the like). Other therapeutic proteins applicable to the disclosure are easily identified in the art.
[0074] In one embodiment, the heterologous polynucleotide within the vector comprises a cytosine deaminase that has been optimized for expression in a human cell. In a further embodiment, the cytosine deaminase comprises a sequence that has been human codon optimized and comprises mutations that increase the cytosine deaminase's stability (e.g., reduced degradation or increased thermo-stability) compared to a wild-type cytosine deaminase. In yet another embodiment, the heterologous polynucleotide encodes a fusion construct comprising a cytosine deaminase (either human codon optimized or non-optimized, either mutated or non-mutated) operably linked to a polynucleotide encoding a polypeptide having UPRT or OPRT activity. In another embodiment, the heterologous polynucleotide comprises a CD polynucleotide of the disclosure (e.g., SEQ ID NO:3, 5, 11, 13, 15, or 17).
[0075] In another embodiment, replication competent retroviral vector can comprise a heterologous polynucleotide encoding a polypeptide comprising a cytosine deaminase (as described herein) and may further comprise a polynucleotide comprising a miRNA or siRNA molecule either as part of the primary transcript from the viral promoter or linked to a promoter, which can be cell-type or tissue specific.
[0076] MicroRNAs (miRNA) are small, non-coding RNAs. They are located within introns of coding or non-coding gene, exons of non-coding genes or in inter-genic regions. miRNA genes are transcribed by RNA polymerase II that generate precursor polynucleotides called primary precursor miRNA (pri-miRNA). The pri-miRNA in the nucleus is processed by the ribonuclease Drosha to produce the miRNA precursor (pre-miRNA) that forms a short hairpin structure. Subsequently, pre-miRNA is transported to the cytoplasm via Exportin 5 and further processed by another ribonuclease called Dicer to generate an active, mature miRNA.
[0077] A mature miRNA is approximately 21 nucleotides in length. It exerts in function by binding to the 3' untranslated region of mRNA of targeted genes and suppressing protein expression either by repression of protein translation or degradation of mRNA. miRNA are involved in biological processes including development, cell proliferation, differentiation and cancer progression. Studies of miRNA profiling indicate that some miRNA expressions are tissue specific or enriched in certain tissues. For example, miR-142-3p, miR-181 and miR-223 expressions have demonstrated to be enriched in hematopoietic tissues in human and mouse (Baskerville et al., 2005 RNA 11, 241-247; Chen et al., 2004 Science 303, 83-86).
[0078] Some miRNAs have been observed to be up-regulated (oncogenic miRNA) or down-regulated (repressor) in several tumors (Spizzo et al., 2009 Cell 137, 586e1). For example, miR-21 is overexpressed in glioblastoma, breast, lung, prostate, colon, stomach, esophageal, and cervical cancer, uterine leiomyosarcoma, DLBCL, head and neck cancer. In contrast, members of let-7 have reported to be down-regulated in glioblastoma, lung, breast, gastric, ovary, prostate and colon cancers. Re-establishment of homeostasis of miRNA expression in cancer is an imperative mechanism to inhibit or reverse cancer progression.
[0079] As a consequence of the vital functions modulated by miRNAs in cancers, focus in developing potential therapeutic approaches has been directed toward antisense-mediated inhibition (antigomers) of oncogenic miRNAs. However, miRNA replacement might represent an equally efficacious strategy. In this approach, the most therapeutically useful miRNAs are the ones expressed at low levels in tumors but at high level, and therefore tolerated, in normal tissues.
[0080] miRNAs that are down-regulated in cancers could be useful as anticancer agents. Examples include mir-128-1, let-7, miR-26, miR-124, and miR-137 (Esquela-Kerscher et al., 2008 Cell Cycle 7, 759-764; Kumar et al., 2008 Proc Natl Acad Sci USA 105, 3903-3908; Kota et al., 2009 Cell 137, 1005-1017; Silber et al., 2008 BMC Medicine 6:14 1-17). miR-128 expression has reported to be enriched in the central nervous system and has been observed to be down-regulated in glioblastomas (Sempere et al., 2004 Genome Biology 5:R13.5-11; Godlewski et al., 2008 Cancer Res 68: (22) 9125-9130). miR-128 is encoded by two distinct genes, miR-128-1 and miR-128-2. Both are processed into identical mature sequence. Bmi-1 and E2F3a have been reported to be the direct targets of miR-128 (Godlewski et al., 2008 Cancer Res 68: (22) 9125-9130; Zhang et al., 2009 J. Mol Med 87:43-51). In addition, Bmi-1 expression has been observed to be up-regulated in a variety of human cancers, including gliomas, mantle cell lymphomas, non-small cell lung cancer B-cell non-Hodgkin's lymphoma, breast, colorectal and prostate cancer. Furthermore, Bmi-1 has been demonstrated to be required for the self-renewal of stem cells from diverse tissues, including neuronal stem cells as well as "stem-like" cell population in gliomas.
[0081] Although there have been a number of in vitro demonstrations of the possibilities of miRNA mediated inhibition of cellular function, it has been difficult to deliver these as oligonucleotides or in viral vectors as efficiently as necessary to have in vivo effects (e.g. Li et al. Cell Cycle 5:2103-2109 2006), as has been true for other molecules. Non-replicative vectors do not appear to be efficient enough in any case to achieve delivery of a therapeutic gene into a significant portion of tumors. However it is also not simple to see how to use replicative vectors to deliver miRNA types of agents. In particular it is not clear how to incorporate extra RNA sequences into the RNA genome of replication competent retroviruses and maintain the replication efficiency and keep the addition stably incorporated into the genome.
[0082] Replication-defective retroviral and lentiviral vectors have been used to stably express pri-mi RNA by a polymerase II promoter such as CMV or LTR and demonstrated production of mature miRNA. However, these vectors do not have to go through the entire lifecycle of the retrovirus or lentivirus multiple times as is required for replicating vectors. The genome has to be able to accommodate many more events than simple entry, integration and transcription. The concerns associated with the use of a RNA-based virus to express miRNA include: (1) the integrity of the viral RNA genome at post transcriptional step during RNA processing; (2) the stability of the inserted cassette during replication; and (3) proper processing of pri-miRNA as part of the viral RNA transcribed from the LTR promoter producing mature miRNA.
[0083] Thus, incorporation of type III RNA polymerase III promoters such as the U6 and the H1 promoter in non-replicative retroviral and lentiviral vectors has been used widely to express functional small interference RNA (siRNA) producing a short hairpin structured RNA (Brormberg-White et al., 2004 J Virol 78:9, 4914-4916; Sliva et al., 2006 Virology 351, 218-225; Haga et al, 2006, Transplant Proc 38(10):3184-8). The loop sequence is cleaved by Dicer producing the mature siRNAs that are 21-22 nucleotides in length. shRNA can be stably expressed in cells to down-regulate target gene expression. However the incorporation of such cassettes into the recombinant replication competent retroviral vector, the expression and the processing by Dicer to produce mature miRNA remain problematic.
[0084] In one embodiment, the disclosure provides a recombinant replication competent retroviral vector that contains a heterologous polynucleotide sequence of a primary precursor miRNA.
[0085] In a further embodiment the primary precursor miRNA is of human origin. In another embodiment the primary precursor RNA sequence is downstream of the env gene.
[0086] In another embodiment, the disclosure provides a recombinant replication competent retroviral vector that contains a heterologous polynucleotide sequence of the human primary precursor miR-128-2 (SEQ ID NO:32) downstream of the env gene. miRNAs that are down-regulated in cancers can be incorporated into the vector for therapeutic gene delivery. For example, let-7, miR-26, miR-124, and miR-137 (Esquela-Kerscher et al., 2008 Cell Cycle 7, 759-764; Kumar et al., 2008 Proc Natl Acad Sci USA 105, 3903-3908; Kota et al., 2009 Cell 137, 1005-1017; Silber et al., 2008 BMC Medicine 6:14 1-17).
[0087] In yet another embodiment, the disclosure provides a recombinant replication competent retroviral vector that contains a heterologous polynucleotide sequence of the short hairpin structured human pre-miR-128 linked to a human H1 promoter (SEQ ID NO: 33 and SEQ ID NO:34) downstream of the env gene. miRNAs that are down-regulated in cancers can be incorporated into the vector for therapeutic gene delivery. For example, let-7, miR-26, miR-124, and miR-137 (Esquela-Kerscher et al., 2008 Cell Cycle 7, 759-764; Kumar et al., 2008 Proc Natl Acad Sci USA 105, 3903-3908; Kota et al., 2009 Cell 137, 1005-1017; Silber et al., 2008 BMC Medicine 6:14 1-17).
[0088] Antisense nucleic acids are DNA or RNA molecules that are complementary to at least a portion of a specific mRNA molecule (Weintraub, Scientific American, 262:40, 1990). In the cell, the antisense nucleic acids hybridize to the corresponding mRNA, forming a double-stranded molecule. The antisense nucleic acids interfere with the translation of the mRNA, since the cell will not translate a mRNA that is double-stranded. Antisense oligomers of about 15 nucleotides are preferred, since they are easily synthesized and are less likely to cause problems than larger molecules when introduced into the target cell. The use of antisense methods to inhibit the in vitro translation of genes is well known in the art (Marcus-Sakura, Anal. Biochem., 172:289, 1988).
[0089] The antisense nucleic acid can be used to block expression of a mutant protein or a dominantly active gene product, such as amyloid precursor protein that accumulates in Alzheimer's disease. Such methods are also useful for the treatment of Huntington's disease, hereditary Parkinsonism, and other diseases. Of particular interest are the blocking of genes associated with cell-proliferative disorders. Antisense nucleic acids are also useful for the inhibition of expression of proteins associated with toxicity.
[0090] Use of an oligonucleotide to stall transcription is known as the triplex strategy since the oligomer winds around double-helical DNA, forming a three-strand helix. Therefore, these triplex compounds can be designed to recognize a unique site on a chosen gene (Maher, et al., Antisense Res. and Dev., 1(3):227, 1991; Helene, C., Anticancer Drug Design, 6(6):569, 1991).
[0091] Ribozymes are RNA molecules possessing the ability to specifically cleave other single-stranded RNA in a manner analogous to DNA restriction endonucleases. Through the modification of nucleotide sequences which encode these RNAs, it is possible to engineer molecules that recognize specific nucleotide sequences in an RNA molecule and cleave it (Cech, J. Amer. Med. Assn., 260:3030, 1988). A major advantage of this approach is that, because they are sequence-specific, only mRNAs with particular sequences are inactivated.
[0092] As used herein, the term "RNA interference" (RNAi) refers to the process of sequence-specific post-transcriptional gene silencing mediated by short interfering nucleic acids (siRNAs or microRNAs (miRNA)). The term "agent capable of mediating RNA interference" refers to siRNAs as well as DNA and RNA vectors that encode siRNAs when transcribed within a cell. The term siRNA or miRNA is meant to encompass any nucleic acid molecule that is capable of mediating sequence specific RNA interference, for example short interfering RNA (siRNA), double-stranded RNA (dsRNA), micro-RNA (miRNA), short hairpin RNA (shRNA), short interfering oligonucleotide, short interfering nucleic acid, short interfering modified oligonucleotide, chemically-modified siRNA, post-transcriptional gene silencing RNA (ptgsRNA), and others.
[0093] Suitable range for designing stem lengths of a hairpin duplex, includes stem lengths of 20-30 nucleotides, 30-50 nucleotides, 50-100 nucleotides, 100-150 nucleotides, 150-200 nucleotides, 200-300 nucleotides, 300-400 nucleotides, 400-500 nucleotides, 500-600 nucleotides, and 600-700 nucleotides. Suitable range for designing loop lengths of a hairpin duplex, includes loop lengths of 4-25 nucleotides, 25-50 nucleotides, or longer if the stem length of the hair duplex is substantial. In certain context, hairpin structures with duplexed regions that are longer than 21 nucleotides may promote effective siRNA-directed silencing, regardless of the loop sequence and length.
[0094] The replicating retroviral vectors of the disclosure can be used to treat disease by expressing engineered siRNA or miRNA (Dennis, Nature, 418: 122 2002) that switches off or lowers expression of key genes that govern the proliferation or survival of diseased cells including tumor cells. Such targets include genes like Rad 51 a central enzyme in DNA repair, and without which cell growth is drastically restricted. Other targets include many of the signaling pathway molecules that control cell growth (Marquez & McCaffrey Hum Gene Ther. 19:27 2008). The siRNA or miRNA may be combined with expression of a cytotoxic gene from the same or different retroviral vector of the disclosure. An example of a suitable cytotoxic gene comprise a cytosine deaminase or modified cytosine deaminase of the disclosure. Examples of siRNA or miRNA that can be expressed from the same vector or a different vector with cytosine deaminase are siRNA or miRNA's that target Thymidilate synthase, Dihydropyrimidine dehydrogenase or other nucleic acid anabolic or synthetic enzymes, that can enhance or complement the action of 5-FU produced locally in a tumor or tissue from 5-FC activation by cytosine deaminase.
[0095] In use, the retroviral vector(s) will replicate through the tumor or other target tissue and before growth inhibition occurs the virus first integrates into the host genome and continues to make virus after growth of that cell is inhibited. Methods for selecting functional miRNA or siRNA sequences are known in the art. Key feature in general in designing effective siRNA or miRNA sequences is usually avoiding "off-target" effects. However for the use of replicating vectors that are highly specific to tumor cells such as those of the disclosure, these side effects are not very important, as the cells are expected to eventually die. A retroviral vector of this disclosure can be made using cells from other species for which the corresponding protein is not significantly targeted. Such cells include dog cell lines or chicken cell line. Alternatively the virus is made by transient transfection on human 293 derived cells or other cell line that allows efficient transient transfection. For this use the virus does not need to utilize an IRES, and the siRNA or miRNA sequence can simply be inserted at a convenient site on the viral genome. This site includes the region downstream of the envelope and upstream of the 3'LTR of the replicating retrovirus. Alternatively polIII transcription units can be inserted in the viral genome with the appropriate siRNA or miRNA's, typically downstream of the 3' envelope gene. Several different siRNA or miRNA sequences can be inserted to ensure efficient down regulation of the target gene or down regulation of more than one gene. Suitable sequences and targets can be obtained from sources known to those skilled in the art. For example: [0096] The MIT/ICBP siRNA Database http: (//)web.mit.edu/sirna/--"The MIT [Massachusetts Institute of Technology]/ICBP [Integrative Cancer Biology Program] siRNA Database is a university-wide effort to catalog these experimentally validated reagents and make that information available to other researchers, both within and outside the MIT community. (Massachusetts Institute of Technology). [0097] RNAi Central--http: (//)katahdin.cshl.org:9331/RNAi web/scripts/main2.pl RNAi resources, including siRNA and shRNA design tools. (Hannon Lab, Cold Spring Harbor Laboratory) [0098] The RNAi Web--http: (//)www.rnaiweb.com/ General resource. [0099] siDIRECT--http: (//)genomics.jp/sidirect/ Online target-specific siRNA design program for mammalian RNA interference. (University of Tokyo, Japan). [0100] siRNA Database--A comprehensive siRNA database that contains siRNA targets against all known mRNA sequences throughout a variety of organisms. (Part of the Protein Lounge systems biology Web site) [0101] siRNA Database and Resources for RNA Interference Studies http: (//)www.rnainterference.org/ [0102] siRNA Selector--http: (//)bioinfo.wistar.upenn.edu/siRNA/siRNA.htm. A set of rules was used for evaluating siRNA functionality based on thermodynamics parameters (Khvorova et al., 2003, Schwarz et al., 2003) and sequence-related determinants developed by Dharmacon (Reynolds et al., 2004). Specificity is determined using BLAST against UniGene databases. (Wistar Institute) [0103] siRNA Target Finder http: (//)www(.)ambion.com/techlib/misc/siRNA_finder.html (Ambion).
[0104] The replicating retroviruses of the disclosure can also express targets for naturally occurring siRNA's that are restricted in expression to particular cell types so that replication of the vector is significantly inhibited in those cell types. The generation of murine leukemia virus-based recombinant replication competent retroviral vector allows high level of transduction and thus high efficiency of gene delivery in vivo. One major concern of using replication competent retroviral vector has been the uncontrolled spread of virus as reported previously (Donahue et al., J. Exp Med. 1992, 176:1124-1135; Calmes et al., Blood 2005, 106: 2530-2533; Seggewiss et al., Blood 2006, 107: 3865-3867). Because of the nature of the virus, the viral spread may be achieved initially within lymphatic cells and subsequently spread to peripheral tissues. For anti-tumor purposes some normal cells in the body that are naturally replicating at some level are hematopoietic cells, cells of the lining of the gut, and some endothelial cells. These are then potential sites where virus that is in the circulation could productively infect. In general this would be undesirable. Any stray infection of cells such as these can be inhibited by including a target for naturally occurring miRNA's or for a combination of miRNA's in these cell types. Some feasibility of using miRNA targets to suppress immune responses has already been shown. (Brown et al. Nat Biotechnol. 2007 25:1457-67). These targets are small RNA sequences with a homologous match to the miRNA sequences that are naturally occurring. These sequences can be inserted in any convenient site in the vectors of the disclosure without, in general significant deleterious consequence for vector viability, other than in a cell of the type desired. Vectors can be made and used as described herein.
[0105] In one embodiment, the disclosure provides a recombinant replication competent retroviral vector that contains a single copy of the miR-142-3p target sequence (142-3pT, SEQ ID NO:35) downstream of the transgene, such as yCD2 or GFP, linked to the IRES. In addition to miR181 and miR-223, the target sequence of other tissue or cell-enriched miRNA can be incorporated into the vector to restrict viral spread in specific tissue or cell type manner. For example, miR-133 and miR206 expressions are highly enriched in muscle cells (Kelly et al., 2008 Nature Medicine 14:11 1278-1283.
[0106] In another embodiment, the disclosure provides a recombinant replication competent retroviral vector that contains 4 copies of the 142-3pT (SEQ ID NO: 36) downstream of the transgene, such as yCD2 or GFP, linked to the IRES. In addition to miR181 and miR-223, the target sequence of other tissue or cell-enriched miRNA can be incorporated into the vector to restrict viral spread in specific tissue or cell type manner. For example, miR-133 and miR206 expressions are highly enriched in muscle cells. The disclosure provides flexibility of single, multiple or combination of target sequence of miRNA and thereby provides restriction of uncontrolled viral spread in a tissue- and/or cell-specific fashion in vitro and in vivo (e. g. hematopoietic and/or muscle cells), (Kelly et al., 2008 Nature Medicine 14:11 1278-1283).
[0107] The miRNA target can be inserted 3' to the transgene but before the 3'LTR or upstream of the IRES but after the 3' end of the envelope. In general the target would not be inserted into protein coding sequences.
[0108] In yet further embodiments, the heterologous polynucleotide may comprise a cytokine such as an interleukin, interferon gamma or the like. Cytokines that may expressed from a retroviral vector of the disclosure include, but are not limited to, IL-1alpha, IL-1beta, IL-2, IL-3, IL-4, IL-5, IL-6, IL-7, IL-8, IL-9, IL-10, IL-11, IL-12, IL-13, IL-14, IL-15, IL-16, IL-17, IL-18, IL-19, IL-20, and IL-21, anti-CD40, CD40L, IFN-gamma and TNF-alpha, soluble forms of TNF-alpha, lymphotoxin-alpha (LT-alpha, also known as TNF-beta), LT-beta (found in complex heterotrimer LT-alpha2-beta), OPGL, FasL, CD27L, CD30L, CD40L, 4-1BBL, DcR3, OX40L, TNF-gamma (International Publication No. WO 96/14328), AIM-I (International Publication No. WO 97/33899), endokine-alpha (International Publication No. WO 98/07880), OPG, and neutrokine-alpha (International Publication No. WO 98/18921, OX40, and nerve growth factor (NGF), and soluble forms of Fas, CD30, CD27, CD40 and 4-IBB, TR2 (International Publication No. WO 96/34095), DR3 (International Publication No. WO 97/33904), DR4 (International Publication No. WO 98/32856), TR5 (International Publication No. WO 98/30693), TRANK, TR9 (International Publication No. WO 98/56892),TR10 (International Publication No. WO 98/54202), 312C2 (International Publication No. WO 98/06842), and TR12, and soluble forms CD154, CD70, and CD153. Angiogenic proteins may be useful in some embodiments, particularly for protein production from cell lines. Such angiogenic factors include, but are not limited to, Glioma Derived Growth Factor (GDGF), Platelet Derived Growth Factor-A (PDGF-A), Platelet Derived Growth Factor-B (PDGF-B), Placental Growth Factor (PIGF), Placental Growth Factor-2 (PIGF-2), Vascular Endothelial Growth Factor (VEGF), Vascular Endothelial Growth Factor-A (VEGF-A), Vascular Endothelial Growth Factor-2 (VEGF-2), Vascular Endothelial Growth Factor B (VEGF-3), Vascular Endothelial Growth Factor B-1 86 (VEGF-B186), Vascular Endothelial Growth Factor-D (VEGF-D), Vascular Endothelial Growth Factor-D (VEGF-D), and Vascular Endothelial Growth Factor-E (VEGF-E). Fibroblast Growth Factors may be delivered by a vector of the disclosure and include, but are not limited to, FGF-1, FGF-2, FGF-3, FGF-4, FGF-5, FGF-6, FGF-7, FGF-8, FGF-9, FGF-10, FGF-11, FGF-12, FGF-13, FGF-14, and FGF-15. Hematopoietic growth factors may be delivered using vectors of the disclosure, such growth factors include, but are not limited to, granulocyte macrophage colony stimulating factor (GM-CSF) (sargramostim), granulocyte colony stimulating factor (G-CSF) (filgrastim), macrophage colony stimulating factor (M-CSF, CSF-1) erythropoietin (epoetin alfa), stem cell factor (SCF, c-kit ligand, steel factor), megakaryocyte colony stimulating factor, PIXY321 (a GMCSF/IL-3) fusion protein and the like.
[0109] Generally, the recombinant virus of the disclosure is capable of transferring a nucleic acid sequence into a target cell.
[0110] The term "regulatory nucleic acid sequence" refers collectively to promoter sequences, polyadenylation signals, transcription termination sequences, upstream regulatory domains, origins of replication, enhancers and the like, which collectively provide for the replication, transcription and translation of a coding sequence in a recipient cell. Not all of these control sequences need always be present so long as the selected coding sequence is capable of being replicated, transcribed and translated in an appropriate host cell. One skilled in the art can readily identify regulatory nucleic acid sequence from public databases and materials. Furthermore, one skilled in the art can identify a regulatory sequence that is applicable for the intended use, for example, in vivo, ex vivo, or in vitro.
[0111] An internal ribosome entry sites ("IRES") refers to a segment of nucleic acid that promotes the entry or retention of a ribosome during translation of a coding sequence usually 3' to the IRES. In some embodiments the IRES may comprise a splice acceptor/donor site, however, preferred IRESs lack a splice acceptor/donor site. Normally, the entry of ribosomes into messenger RNA takes place via the cap located at the 5' end of all eukaryotic mRNAs. However, there are exceptions to this universal rule. The absence of a cap in some viral mRNAs suggests the existence of alternative structures permitting the entry of ribosomes at an internal site of these RNAs. To date, a number of these structures, designated IRES on account of their function, have been identified in the 5' noncoding region of uncapped viral mRNAs, such as that, in particular, of picornaviruses such as the poliomyelitis virus (Pelletier et al., 1988, Mol. Cell. Biol., 8, 1103-1112) and the EMCV virus (encephalo-myocarditis virus (Jang et al., J. Virol., 1988, 62, 2636-2643). The disclosure provides the use of an IRES in the context of a replication-competent retroviral vector.
[0112] The term "promoter region" is used herein in its ordinary sense to refer to a nucleotide region comprising a DNA regulatory sequence, wherein the regulatory sequence is derived from a gene which is capable of binding RNA polymerase and initiating transcription of a downstream (3'-direction) coding sequence. The regulatory sequence may be homologous or heterologous to the desired gene sequence. For example, a wide range of promoters may be utilized, including viral or mammalian promoter as described above.
[0113] The heterologous nucleic acid sequence is typically under control of either the viral LTR promoter-enhancer signals or an internal promoter, and retained signals within the retroviral LTR can still bring about efficient integration of the vector into the host cell genome. Accordingly, the recombinant retroviral vectors of the disclosure, the desired sequences, genes and/or gene fragments can be inserted at several sites and under different regulatory sequences. For example, a site for insertion can be the viral enhancer/promoter proximal site (i.e., 5' LTR-driven gene locus). Alternatively, the desired sequences can be inserted into a regulatory sequence distal site (e.g., the IRES sequence 3' to the env gene) or where two or more heterologous sequences are present one heterologous sequence may be under the control of a first regulatory region and a second heterologous sequence under the control of a second regulatory region. Other distal sites include viral promoter sequences, where the expression of the desired sequence or sequences is through splicing of the promoter proximal cistron, an internal heterologous promoter as SV40 or CMV, or an internal ribosome entry site (IRES) can be used.
[0114] In one embodiment, the retroviral genome of the disclosure contains an IRES comprising a cloning site downstream of the IRES for insertion of a desired/heterologous polynucleotide. In one embodiment, the IRES is located 3' to the env gene in the retroviral vector, but 5' to the desired heterologous polynucleotide. Accordingly, a heterologous polynucleotide encoding a desired polypeptide may be operably linked to the IRES.
[0115] In another embodiment, a targeting polynucleotide sequence is included as part of the recombinant retroviral vector of the disclosure. The targeting polynucleotide sequence is a targeting ligand (e.g., peptide hormones such as heregulin, a single-chain antibodies, a receptor or a ligand for a receptor), a tissue-specific or cell-type specific regulatory element (e.g., a tissue-specific or cell-type specific promoter or enhancer), or a combination of a targeting ligand and a tissue-specific/cell-type specific regulatory element. Preferably, the targeting ligand is operably linked to the env protein of the retrovirus, creating a chimeric retroviral env protein. The viral GAG, viral POL and viral ENV proteins can be derived from any suitable retrovirus (e.g., MLV or lentivirus-derived). In another embodiment, the viral ENV protein is non-retrovirus-derived (e.g., CMV or VSV).
[0116] In one embodiment, the recombinant retrovirus of the disclosure is genetically modified in such a way that the virus is targeted to a particular cell type (e.g., smooth muscle cells, hepatic cells, renal cells, fibroblasts, keratinocytes, mesenchymal stem cells, bone marrow cells, chondrocyte, epithelial cells, intestinal cells, mammary cells, neoplastic cells, glioma cells, neuronal cells and others known in the art) such that the recombinant genome of the retroviral vector is delivered to a target non-dividing, a target dividing cell, or a target cell having a cell proliferative disorder.
[0117] In one embodiment, the retroviral vector is targeted to the cell by binding to cells having a molecule on the external surface of the cell. This method of targeting the retrovirus utilizes expression of a targeting ligand on the coat of the retrovirus to assist in targeting the virus to cells or tissues that have a receptor or binding molecule which interacts with the targeting ligand on the surface of the retrovirus. After infection of a cell by the virus, the virus injects its nucleic acid into the cell and the retrovirus genetic material can integrate into the host cell genome.
[0118] In another embodiment, targeting uses cell- or tissue-specific regulatory elements to promote expression and transcription of the viral genome in a targeted cell which actively utilizes the regulatory elements, as described more fully below. The transferred retrovirus genetic material is then transcribed and translated into proteins within the host cell. The targeting regulatory element is typically linked to the 5' and/or 3' LTR, creating a chimeric LTR.
[0119] By inserting a heterologous polynucleotide of interest into the viral vector of the disclosure, along with another gene which encodes, for example, the ligand for a receptor on a specific target cell, the vector is now target specific. Viral vectors can be made target specific by attaching, for example, a sugar, a glycolipid, or a protein. Targeting can be accomplished by using an antibody to target the viral vector. Those of skill in the art will know of, or can readily ascertain, specific polynucleotide sequences which can be inserted into the viral genome or proteins which can be attached to a viral envelope to allow target specific delivery of the viral vector containing the nucleic acid sequence of interest.
[0120] Thus, the disclosure includes in one embodiment, a chimeric env protein comprising a retroviral ENV protein operably linked to a targeting polypeptide. The targeting polypeptide can be a cell specific receptor molecule, a ligand for a cell specific receptor, an antibody or antibody fragment to a cell specific antigenic epitope or any other ligand easily identified in the art which is capable of binding or interacting with a target cell. Examples of targeting polypeptides or molecules include bivalent antibodies using biotin-streptavidin as linkers (Etienne-Julan et al., J. Of General Virol., 73, 3251-3255 (1992); Roux et al., Proc. Natl. Acad. Sci USA 86, 9079-9083 (1989)), recombinant virus containing in its envelope a sequence encoding a single-chain antibody variable region against a hapten (Russell et al., Nucleic Acids Research, 21, 1081-1085 (1993)), cloning of peptide hormone ligands into the retrovirus envelope (Kasahara et al., Science, 266, 1373-1376 (1994)), chimeric EPO/env constructs (Kasahara et al., 1994), single-chain antibody against the low density lipoprotein (LDL) receptor in the ecotropic MLV envelope, resulting in specific infection of HeLa cells expressing LDL receptor (Somia et al., Proc. Natl. Acad. Sci USA, 92, 7570-7574 (1995)), similarly the host range of ALV can be altered by incorporation of an integrin ligand, enabling the virus to now cross species to specifically infect rat glioblastoma cells (Valsesia-Wittmann et al., J. Virol. 68, 4609-4619 (1994)), and Dornberg and co-workers (Chu and Dornburg, J. Virol 69, 2659-2663 (1995); M. Engelstadter et al.Gene Therapy 8, 1202-1206 (2001)) have reported tissue-specific targeting of spleen necrosis virus (SNV), an avian retrovirus, using envelopes containing single-chain antibodies directed against tumor markers.
[0121] The disclosure provides a method of producing a recombinant retrovirus capable of infecting a target cell comprising transfecting a suitable host cell with the following: a vector comprising a polynucleotide sequence encoding a viral gag, a viral pol and a viral env, and a heterologous polynucleotide, operably linked to a regulatory nucleic acid sequence, and recovering the recombinant virus.
[0122] The retrovirus and methods of the disclosure provide a replication competent retrovirus that does not require helper virus or additional nucleic acid sequence or proteins in order to propagate and produce virion. For example, the nucleic acid sequences of the retrovirus of the disclosure encode a group specific antigen and reverse transcriptase, (and integrase and protease-enzymes necessary for maturation and reverse transcription), respectively, as discussed above. The viral gag and pol can be derived from a lentivirus, such as HIV or an oncovirus or gammaretrovirus such as MoMLV. In addition, the nucleic acid genome of the retrovirus of the disclosure includes a sequence encoding a viral envelope (ENV) protein. The env gene can be derived from any retroviruses. The env may be an amphotropic envelope protein which allows transduction of cells of human and other species, or may be an ecotropic envelope protein, which is able to transduce only mouse and rat cells. Further, it may be desirable to target the recombinant virus by linkage of the envelope protein with an antibody or a particular ligand for targeting to a receptor of a particular cell-type. As mentioned above, retroviral vectors can be made target specific by inserting, for example, a glycolipid, or a protein. Targeting is often accomplished by using an antibody to target the retroviral vector to an antigen on a particular cell-type (e.g., a cell type found in a certain tissue, or a cancer cell type). Those of skill in the art will know of, or can readily ascertain without undue experimentation, specific methods to achieve delivery of a retroviral vector to a specific target. In one embodiment, the env gene is derived from a non-retrovirus (e.g., CMV or VSV). Examples of retroviral-derived env genes include, but are not limited to: Moloney murine leukemia virus (MoMuLV), Harvey murine sarcoma virus (HaMuSV), murine mammary tumor virus (MuMTV), gibbon ape leukemia virus (GaLV), human immunodeficiency virus (HIV) and Rous Sarcoma Virus (RSV). Other env genes such as Vesicular stomatitis virus (VSV) (Protein G), cytomegalovirus envelope (CMV), or influenza virus hemagglutinin (HA) can also be used.
[0123] In one embodiment, the retroviral genome is derived from an onco-retrovirus, and more particularly a mammalian onco-retrovirus. In a further embodiment, the retroviral genome is derived from a gamma retrovirus, and more particularly a mammalian gamma retrovirus. By "derived" is meant that the parent polynucleotide sequence is an wild-type oncovirus which has been modified by insertion or removal of naturally occurring sequences (e.g., insertion of an IRES, insertion of a heterologous polynucleotide encoding a polypeptide or inhibitory nucleic acid of interest, swapping of a more effective promoter from a different retrovirus or virus in place of the wild-type promoter and the like).
[0124] Unlike recombinant retroviruses produced by standard methods in the art that are defective and require assistance in order to produce infectious vector particles, the disclosure provides a retrovirus that is replication-competent.
[0125] In another embodiment, the disclosure provides retroviral vectors that are targeted using regulatory sequences. Cell- or tissue-specific regulatory sequences (e.g., promoters) can be utilized to target expression of gene sequences in specific cell populations. Suitable mammalian and viral promoters for the disclosure are described elsewhere herein. Accordingly, in one embodiment, the disclosure provides a retrovirus having tissue-specific promoter elements at the 5' end of the retroviral genome. Typically, the tissue-specific regulatory elements/sequences are in the U3 region of the LTR of the retroviral genome, including for example cell- or tissue-specific promoters and enhancers to neoplastic cells (e.g., tumor cell-specific enhancers and promoters), and inducible promoters (e.g., tetracycline).
[0126] Transcription control sequences of the disclosure can also include naturally occurring transcription control sequences naturally associated with a gene encoding a superantigen, a cytokine or a chemokine.
[0127] In some circumstances, it may be desirable to regulate expression. For example, different viral promoters with varying strengths of activity may be utilized depending on the level of expression desired. In mammalian cells, the CMV immediate early promoter if often used to provide strong transcriptional activation. Modified versions of the CMV promoter that are less potent have also been used when reduced levels of expression of the transgene are desired. When expression of a transgene in hematopoietic cells is desired, retroviral promoters such as the LTRs from MLV or MMTV can be used. Other viral promoters that can be used include SV40, RSV LTR, HIV-1 and HIV-2 LTR, adenovirus promoters such as from the E1A, E2A, or MLP region, AAV LTR, cauliflower mosaic virus, HSV-TK, and avian sarcoma virus.
[0128] Similarly tissue specific or selective promoters may be used to effect transcription in specific tissues or cells so as to reduce potential toxicity or undesirable effects to non-targeted tissues. For example, promoters such as the PSA, probasin, prostatic acid phosphatase or prostate-specific glandular kallikrein (hK2) may be used to target gene expression in the prostate. The Whey accessory protein (WAP) may be used for breast tissue expression (Andres et al., PNAS 84:1299-1303, 1987). Other promoters/regulatory domains that can be used are set forth in Table 1.
[0129] "Tissue-specific regulatory elements" are regulatory elements (e.g., promoters) that are capable of driving transcription of a gene in one tissue while remaining largely "silent" in other tissue types. It will be understood, however, that tissue-specific promoters may have a detectable amount of "background" or "base" activity in those tissues where they are silent. The degree to which a promoter is selectively activated in a target tissue can be expressed as a selectivity ratio (activity in a target tissue/activity in a control tissue). In this regard, a tissue specific promoter useful in the practice of the disclosure typically has a selectivity ratio of greater than about 5. Preferably, the selectivity ratio is greater than about 15.
[0130] In certain indications, it may be desirable to activate transcription at specific times after administration of the recombinant replication competent retrovirus of the disclosure (RRCR). This may be done with promoters that are hormone or cytokine regulatable. For example in therapeutic applications where the indication is a gonadal tissue where specific steroids are produced or routed to, use of androgen or estrogen regulated promoters may be advantageous. Such promoters that are hormone regulatable include MMTV, MT-1, ecdysone and RuBisco. Other hormone regulated promoters such as those responsive to thyroid, pituitary and adrenal hormones may be used. Cytokine and inflammatory protein responsive promoters that could be used include K and T Kininogen (Kageyama et al., 1987), c-fos, TNF-alpha, C-reactive protein (Arcone et al., 1988), haptoglobin (Oliviero et al., 1987), serum amyloid A2, C/EBP alpha, IL-1, IL-6 (Poli and Cortese, 1989), Complement C3 (Wilson et al., 1990), IL-8, alpha-1 acid glycoprotein (Prowse and Baumann, 1988), alpha-1 antitypsin, lipoprotein lipase (Zechner et al., 1988), angiotensinogen (Ron et al., 1990), fibrinogen, c-jun (inducible by phorbol esters, TNF-alpha, UV radiation, retinoic acid, and hydrogen peroxide), collagenase (induced by phorbol esters and retinoic acid), metallothionein (heavy metal and glucocorticoid inducible), Stromelysin (inducible by phorbol ester, interleukin-1 and EGF), alpha-2 macroglobulin and alpha-1 antichymotrypsin. Tumor specific promoters such as osteocalcin, hypoxia-responsive element (HRE), MAGE-4, CEA, alpha-fetoprotein, GRP78/BiP and tyrosinase may also be used to regulate gene expression in tumor cells.
[0131] In addition, this list of promoters should not be construed to be exhaustive or limiting, those of skill in the art will know of other promoters that may be used in conjunction with the promoters and methods disclosed herein.
TABLE-US-00001 TABLE 1 TISSUE SPECIFIC PROMOTERS Tissue Promoter Pancreas Insulin Elastin Amylase pdr-1 pdx-1 glucokinase Liver Albumin PEPCK HBV enhancer .alpha. fetoprotein apolipoprotein C .alpha.-1 antitrypsin vitellogenin, NF-AB Transthyretin Skeletal muscle Myosin H chain Muscle creatine kinase Dystrophin Calpain p94 Skeletal alpha-actin fast troponin 1 Skin Keratin K6 Keratin K1 Lung CFTR Human cytokeratin 18 (K18) Pulmonary surfactant proteins A, B and C CC-10 P1 Smooth muscle sm22 .alpha. SM-alpha-actin Endothelium Endothelin-1 E-selectin von Willebrand factor TIE (Korhonen et al., 1995) KDR/flk-1 Melanocytes Tyrosinase Adipose tissue Lipoprotein lipase (Zechner et al., 1988) Adipsin (Spiegelman et al., 1989) acetyl- CoA carboxylase (Pape and Kim, 1989) glycerophosphate dehydrogenase (Dani et al., 1989) adipocyte P2 (Hunt et al., 1986) Breast Whey Acidic Protien (WAP) (Andres et al. PNAS 84: 1299-1303 1987 Blood .beta.-globin
[0132] It will be further understood that certain promoters, while not restricted in activity to a single tissue type, may nevertheless show selectivity in that they may be active in one group of tissues, and less active or silent in another group. Such promoters are also termed "tissue specific", and are contemplated for use with the disclosure. For example, promoters that are active in a variety of central nervous system (CNS) neurons may be therapeutically useful in protecting against damage due to stroke, which may affect any of a number of different regions of the brain. Accordingly, the tissue-specific regulatory elements used in the disclosure, have applicability to regulation of the heterologous proteins as well as a applicability as a targeting polynucleotide sequence in the present retroviral vectors.
[0133] In yet another embodiment, the disclosure provides plasmids comprising a recombinant retroviral derived construct. The plasmid can be directly introduced into a target cell or a cell culture such as NIH 3T3 or other tissue culture cells. The resulting cells release the retroviral vector into the culture medium.
[0134] The disclosure provides a polynucleotide construct comprising from 5' to 3': a promoter or regulatory region useful for initiating transcription; a psi packaging signal; a gag encoding nucleic acid sequence, a pol encoding nucleic acid sequence; an env encoding nucleic acid sequence; an internal ribosome entry site nucleic acid sequence; a heterologous polynucleotide encoding a marker, therapeutic or diagnostic polypeptide; and a LTR nucleic acid sequence. As described elsewhere herein and as follows the various segment of the polynucleotide construct of the disclosure (e.g., a recombinant replication competent retroviral polynucleotide) are engineered depending in part upon the desired host cell, expression timing or amount, and the heterologous polynucleotide. A replication competent retroviral construct of the disclosure can be divided up into a number of domains that may be individually modified by those of skill in the art.
[0135] For example, the promoter can comprise a CMV promoter having a sequence as set forth in SEQ ID NO:19, 20 or 22 from nucleotide 1 to about nucleotide 582 and may include modification to one or more (e.g., 2-5, 5-10, 10-20, 20-30, 30-50, 50-100 or more nucleic acid bases) so long as the modified promoter is capable of directing and initiating transcription. In one embodiment, the promoter or regulatory region comprises a CMV-R-U5 domain polynucleotide. The CMV-R-U5 domain comprises the immediately early promoter from human cytomegalovirus to the MLV R-U5 region. In one embodiment, the CMV-R-U5 domain polynucleotide comprises a sequence as set forth in SEQ ID NO:19, 20 or 22 from about nucleotide 1 to about nucleotide 1202 or sequences that are at least 95% identical to a sequence as set forth in SEQ ID NO:19, 20, or 22 wherein the polynucleotide promotes transcription of a nucleic acid molecule operably linked thereto. The gag domain of the polynucleotide may be derived from any number of retroviruses, but will typically be derived from an oncoretrovirus and more particularly from a mammalian oncoretrovirus. In one embodiment the gag domain comprises a sequence from about nucleotide number 1203 to about nucleotide 2819 or a sequence having at least 95%, 98%, 99% or 99.8% (rounded to the nearest 10.sup.th) identity thereto. The pol domain of the polynucleotide may be derived from any number of retroviruses, but will typically be derived from an oncoretrovirus and more particularly from a mammalian oncoretrovirus. In one embodiment the pol domain comprises a sequence from about nucleotide number 2820 to about nucleotide 6358 or a sequence having at least 95%, 98%, 99% or 99.9% (roundest to the nearest 10.sup.th) identity thereto. The env domain of the polynucleotide may be derived from any number of retroviruses, but will typically be derived from an oncoretrovirus or gamma-retrovirus and more particularly from a mammalian oncoretrovirus or gamma-retrovirus. In some embodiments the env coding domain comprises an amphotropic env domain. In one embodiment the env domain comprises a sequence from about nucleotide number 6359 to about nucleotide 8323 or a sequence having at least 95%, 98%, 99% or 99.8% (roundest to the nearest 10.sup.th) identity thereto. The IRES domain of the polynucleotide may be obtained from any number of internal ribosome entry sites. In one embodiment, IRES is derived from an encephalomyocarditis virus. In one embodiment the IRES domain comprises a sequence from about nucleotide number 8327 to about nucleotide 8876 or a sequence having at least 95%, 98%, or 99% (roundest to the nearest 10.sup.th) identity thereto so long as the domain allows for entry of a ribosome. The heterologous domain can comprise a cytosine deaminase of the disclosure. In one embodiment, the CD polynucleotide comprises a human codon optimized sequence. In yet another embodiment, the CD polynucleotide encodes a mutant polypeptide having cytosine deaminase, wherein the mutations confer increased thermal stabilization that increase the melting temperature (Tm) by 10.degree. C. allowing sustained kinetic activity over a broader temperature range and increased accumulated levels of protein. In one embodiment, the cytosine deaminase comprises a sequence as set forth in SEQ ID NO:19 or 22 from about nucleotide number 8877 to about 9353. The heterologous domain may be followed by a polypurine rich domain. The 3' LTR can be derived from any number of retroviruses, typically an oncoretrovirus and preferably a mammalian oncoretrovirus. In one embodiment, the 3' LTR comprises a U3-R-U5 domain. In yet another embodiment the LTR comprises a sequence as set forth in SEQ ID NO:19 or 22 from about nucleotide 9405 to about 9998 or a sequence that is at least 95%, 98% or 99.5% (rounded to the nearest 10.sup.th) identical thereto.
[0136] The disclosure also provides a recombinant retroviral vector comprising from 5' to 3' a CMV-R-U5, fusion of the immediate early promoter from human cytomegalovirus to the MLV R-U5 region; a PBS, primer binding site for reverse transcriptase; a 5' splice site; a i packaging signal; a gag, ORF for MLV group specific antigen; a pol, ORF for MLV polymerase polyprotein; a 3' splice site; a 4070A env, ORF for envelope protein of MLV strain 4070A; an IRES, internal ribosome entry site of encephalomyocarditis virus; a modified cytosine deaminase (thermostabilized and codon optimized); a PPT, polypurine tract; and a U3-R-U5, MLV long terminal repeat. This structure is further depicted in FIG. 3.
[0137] The disclosure also provides a retroviral vector comprising a sequence as set forth in SEQ ID NO:19, 20 or 22.
[0138] The retroviral vectors can be used to treat a wide range of disease and disorders including a number of cell proliferative diseases and disorders (see, e.g., U.S. Pat. Nos. 4,405,712 and 4,650,764; Friedmann, 1989, Science, 244:1275-1281; Mulligan, 1993, Science, 260:926-932, R. Crystal, 1995, Science 270:404-410, each of which are incorporated herein by reference in their entirety, see also, The Development of Human Gene Therapy, Theodore Friedmann, Ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 1999. ISBN 0-87969-528-5, which is incorporated herein by reference in its entirety).
[0139] The disclosure also provides gene therapy for the treatment of cell proliferative disorders. Such therapy would achieve its therapeutic effect by introduction of an appropriate therapeutic polynucleotide (e.g., antisense, ribozymes, suicide genes, siRNA), into cells of subject having the proliferative disorder. Delivery of polynucleotide constructs can be achieved using the recombinant retroviral vector of the disclosure, particularly if it is based on MLV, which is capable of infecting dividing cells.
[0140] In addition, the therapeutic methods (e.g., the gene therapy or gene delivery methods) as described herein can be performed in vivo or ex vivo. It may be preferable to remove the majority of a tumor prior to gene therapy, for example surgically or by radiation. In some aspects, the retroviral therapy may be preceded or followed by surgery, chemotherapy or radiation therapy.
[0141] Thus, the disclosure provides a recombinant retrovirus capable of infecting a non-dividing cell, a dividing cell or a neoplastic cell, therein the recombinant retrovirus comprises a viral GAG; a viral POL; a viral ENV; a heterologous nucleic acid operably linked to an IRES; and cis-acting nucleic acid sequences necessary for packaging, reverse transcription and integration. The recombinant retrovirus can be a lentivirus, such as HIV, or can be an oncovirus. As described above for the method of producing a recombinant retrovirus, the recombinant retrovirus of the disclosure may further include at least one of VPR, VIF, NEF, VPX, TAT, REV, and VPU protein. While not wanting to be bound by a particular theory, it is believed that one or more of these genes/protein products are important for increasing the viral titer of the recombinant retrovirus produced (e.g., NEF) or may be necessary for infection and packaging of virion.
[0142] The disclosure also provides a method of nucleic acid transfer to a target cell to provide expression of a particular nucleic acid (e.g., a heterologous sequence). Therefore, in another embodiment, the disclosure provides a method for introduction and expression of a heterologous nucleic acid in a target cell comprising infecting the target cell with the recombinant virus of the disclosure and expressing the heterologous nucleic acid in the target cell. As mentioned above, the target cell can be any cell type including dividing, non-dividing, neoplastic, immortalized, modified and other cell types recognized by those of skill in the art, so long as they are capable of infection by a retrovirus.
[0143] It may be desirable to modulate the expression of a gene in a cell by the introduction of a nucleic acid sequence (e.g., the heterologous nucleic acid sequence) by the method of the disclosure, wherein the nucleic acid sequence give rise, for example, to an antisense or ribozyme molecule. The term "modulate" envisions the suppression of expression of a gene when it is over-expressed, or augmentation of expression when it is under-expressed. Where a cell proliferative disorder is associated with the expression of a gene, nucleic acid sequences that interfere with the gene's expression at the translational level can be used. This approach utilizes, for example, antisense nucleic acid, ribozymes, or triplex agents to block transcription or translation of a specific mRNA, either by masking that mRNA with an antisense nucleic acid or triplex agent, or by cleaving it with a ribozyme.
[0144] It may be desirable to transfer a nucleic acid encoding a biological response modifier (e.g., a cytokine) into a cell or subject. Included in this category are immunopotentiating agents including nucleic acids encoding a number of the cytokines classified as "interleukins". These include, for example, interleukins 1 through 15, as well as other response modifiers and factors described elsewhere herein. Also included in this category, although not necessarily working according to the same mechanisms, are interferons, and in particular gamma interferon, tumor necrosis factor (TNF) and granulocyte-macrophage-colony stimulating factor (GM-CSF). Other polypeptides include, for example, angiogenic factors and anti-angiogenic factors. It may be desirable to deliver such nucleic acids to bone marrow cells or macrophages to treat enzymatic deficiencies or immune defects. Nucleic acids encoding growth factors, toxic peptides, ligands, receptors, or other physiologically important proteins can also be introduced into specific target cells.
[0145] The disclosure can be used for delivery of heterologous polynucleotides that promote drug specific targeting and effects. For example, HER2 (see, e.g., SEQ ID NO:23 and 24), a member of the EGF receptor family, is the target for binding of the drug trastuzumab (Herceptin.TM., Genentech). Trastuzumab is a mediator of antibody-dependent cellular cytotoxicity (ADCC). Activity is preferentially targeted to HER2-expressing cells with 2+ and 3+ levels of overexpression by immunohistochemistry rather than 1+ and non-expressing cells (Herceptin prescribing information, Crommelin 2002). Enhancement of expression of HER2 by introduction of vector expressing HER2 or truncated HER2 (expressing only the extracellular and transmembrane domains) in HER2 low tumors may facilitate optimal triggering of ADCC and overcome the rapidly developing resistance to Herceptin that is observed in clinical use.
[0146] The substitution of yCD2 (comprising SEQ ID NO:19 from about 8877 to 9353) for the intracellular domain of HER2 allows for cell surface expression of HER2 and cytosolic localization of yCD2. The HER2 extracellular domain (ECD) and transmembrane domain (TM) (approximately 2026 bp from about position 175 to 2200 of SEQ ID NO:23) can be amplified by PCR (Yamamoto et al., Nature 319:230-234, 1986; Chen et al., Canc. Res., 58:1965-1971, 1998) or chemically synthesized (BioBasic Inc., Markham, Ontario, Canada) and inserted between the IRES and yCD2 gene in the vector pAC3-yCD2 SEQ ID NO: 19 (e.g., between about nucleotide 8876 and 8877 of SEQ ID NO:19). Alternatively, the yCD gene can be excised and replaced with a polynucleotide encoding a HER2 polypeptide or fragment thereof. A further truncated HER2 with only the Herceptin binding domain IV of the ECD and TM domains (approximately 290 bp from position 1910 to 2200) can be amplified or chemically synthesized and used as above (Landgraf 2007; Garrett et al., J. of Immunol., 178:7120-7131, 2007). A further modification of this truncated form with the native signal peptide (approximately 69 bp from position 175-237) fused to domain IV and the TM can be chemically synthesized and used as above. The resulting viruses can be used to treat a cell proliferative disorder in a subject in combination with trastuzumab or trastuzumab and 5-FC.
[0147] Alternatively, HER2 and the modifications described above can be expressed in a separate vector containing a different ENV gene or other appropriate surface protein. This vector can be replication competent (Logg et al. J.Mol Biol. 369:1214 2007) or non-replicative "first generation" retroviral vector that encodes the envelope and the gene of interest (Emi et al. J.Virol 65:1202 1991). In the latter case the pre-existing viral infection will provide complementary gag and pol to allow infective spread of the "non-replicative" vector from any previously infected cell. Alternate ENV and glycoproteins include xenotropic and polytropic ENV and glycoproteins capable of infecting human cells, for example ENV sequences from the NZB strain of MLV and glycoproteins from MCF, VSV, GALV and other viruses (Palu 2000, Baum et al., Mol. Therapy, 13(6):1050-1063, 2006). For example, a polynucleotide can comprise a sequence wherein the GAG and POL and yCD2 genes of SEQ ID NO: 19 are deleted, the ENV corresponds to a xenotropic ENV domain of NZB MLV or VSV-g, and the IRES or a promoter such as RSV is operatively linked directly to HER2, HER2 ECD.TM., HER2 ECDIV.TM., or HER2 SECDIV.TM..
[0148] Mixed infection of cells by VSVG pseudotyped virus and amphotropic retrovirus results in the production of progeny virions bearing the genome of one virus encapsidated by the envelope proteins of the other. The same is true for other envelopes that pseudotype retroviral particles. For example, infection by retroviruses derived as above results in production of progeny virions capable of encoding yCD2 and HER2 (or variant) in infected cells. The resulting viruses can be used to treat a cell proliferative disorder in a subject in combination with trastuzumab or trastuzumab and 5-FC.
[0149] Recently, a gamma retrovirus, XMRV, has been associated with prostate cancer in humans with the virus showing a strong preference for replication in prostate tissue (R. Schlaberg et al. PNAS 106: 16351-16356 2009). The virus appears very similar to xenotropic MLV. In one embodiment of the disclosure, non-replicative retroviral vectors are provided which carry both a therapeutic gene (cytosine deaminase, thymidine kinase, other prodrug activating genes, interferons, IL-2, IL-12, other cytokines, p53 other anti-oncogenes, anti-cancer miRNA or the like) and an envelope gene that is capable of being complemented by the XMRV gag and gag-pol functions such as an amphotropic envelope, a GALV envelope, a VSVg protein envelope, or other envelopes known to those skilled in the art. The non-replicative vector polynucleotide is delivered to the prostate cancer in a patient or animal as a DNA or RNA molecule using one of: non-viral or physical delivery systems; a heterologous viral delivery system such as an adenoviral vector, or as a manufactured retroviral particle. Once delivered the non-replicative vector will be spread by complementation by XMRV and infection of neighboring cells will take place until the boundary of XMRV infection is reached, when the XMRV complementation will not be available. The therapeutic gene can then have its effect (e.g. after a prodrug is administered) in the XMRV infected area only. The same rescue effect can be achieved using a replicative retroviral vector of the disclosure. This strategy (complementary non-replicative vector with a therapeutic gene) can be used with any retroviral disease (HIV infection, HTLV1 infection, other cancer associated retroviruses), or with any viral or viral associated disease (HPV infection and HPV E6 & E7 expression in cervical cancers, EBV associated lymphomas or carcinomas etc.).
[0150] Another aspect of the development of resistance to trastuzumab relates to the interference with intracellular signaling required for the activity of trastuzumab. Resistant cells show loss of PTEN and lower expression of p27kip1 [Fujita, Brit J. Cancer, 94:247, 2006; Lu et al., Journal of the National Cancer Institute, 93(24): 1852-1857, 2001; Kute et al., Cytometry Part A 57A:86-93, 2004). For example, a polynucleotide encoding PTEN (SEQ ID NO:25) can be recombinantly generated or chemically synthesized (BioBasic Inc., Markham, Canada) and operably inserted directly after the yCD2 polynucleotide in the vector pAC3-yCD2 SEQ ID NO: 19 or 22, or with a linker sequence as previously described, or as a replacement for yCD2. In a further example, the PTEN encoding polynucleotide can be synthesized as above and inserted between the IRES and yCD2 sequences or with a linker as previously described.
[0151] Alternatively, PTEN can be expressed in a separate vector containing a different ENV gene or other appropriate surface protein. This vector can be replication competent (Logg et al. J.Mol Biol. 369:1214 2007) or non-replicative "first generation" retroviral vector that encodes the envelope and the gene of interest (Emi et al., J.Virol 65:1202 1991). In the latter case the pre-existing viral infection will provide complementary gag and pol to allow infective spread of the "non-replicative" vector from any previously infected cell. Alternate ENV and glycoproteins include xenotropic and polytropic ENV and glycoproteins capable of infecting human cells, for example ENV sequences from the NZB strain of MLV and glycoproteins from MCF, VSV, GALV and other viruses (Palu, Rev Med Virol. 2000, Baum, Mol. Ther. 13(6):1050-1063, 2006). For example, a polynucleotide can comprise a sequence wherein the GAG and POL and yCD2 genes of SEQ ID NO: 19 are deleted, the ENV corresponds to a xenotropic ENV domain of NZB MLV or VSV-g, and the IRES or a promoter such as RSV is operatively linked directly to PTEN.
[0152] Mixed infection of cells by VSVG pseudotyped virus and amphotropic retrovirus results in the production of progeny virions bearing the genome of one virus encapsidated by the envelope proteins of the other [Emi 1991]. The same is true for other envelopes that pseudotype retroviral particles. For example, infection by retroviruses derived as above results in production of progeny virions capable of encoding yCD2 and PTEN (or variant) or PTEN alone in infected cells. The resulting viruses can be used to treat a cell proliferative disorder in a subject in combination with trastuzumab or trastuzumab and 5-FC.
[0153] Similarly, a polynucleotide encoding p27kip1 (SEQ ID NO:27 and 28) can be chemically synthesized (BioBasic Inc., Markham, Canada) and operably inserted directly after the yCD2 gene in the vector pAC3-yCD2 SEQ ID NO: 19 or with a linker sequence. In a further example, the p27kip1 encoding polynucleotide can be synthesized as above and inserted between the IRES and yCD2 sequences or with a linker as previously described or in place of the yCD2 gene.
[0154] Alternatively, p27kip1 can be expressed in a separate vector containing a different ENV gene or other appropriate surface protein. This vector can be replication competent (C R. Logg et al. J. Mol Biol. 369:1214 2007) or non-replicative "first generation" retroviral vector that encodes the envelope and the gene of interest (Emi et al. J. Virol 65:1202 1991). In the latter case the pre-existing viral infection will provide complementary gag and pol to allow infective spread of the "non-replicative" vector from any previously infected cell. Alternate ENV and glycoproteins include xenotropic and polytropic ENV and glycoproteins capable of infecting human cells, for example ENV sequences from the NZB strain of MLV and glycoproteins from MCF, VSV, GALV and other viruses (Palu 2000, Baum 2006, supra). For example, a polynucleotide can comprise a sequence wherein the GAG and POL and yCD2 genes of SEQ ID NO: 19 are deleted, the ENV corresponds to a xenotropic ENV domain of NZB MLV or VSV-g, and the IRES or a promoter such as RSV is operatively linked directly to p27kip1.
[0155] Mixed infection of cells by VSVG pseudotyped virus and amphotropic retrovirus results in the production of progeny virions bearing the genome of one virus encapsidated by the envelope proteins of the other [Emi 1991]. The same is true for other envelopes that pseudotype retroviral particles. For example, infection by retroviruses derived as above from both SEQ ID NO: 19 and 22 results in production of progeny virions capable of encoding yCD2 and p27kip1 (or variant) in infected cells. The resulting viruses can be used to treat a cell proliferative disorder in a subject in combination with trastuzumab or trastuzumab and 5-FC.
[0156] In another example, CD20 is the target for binding of the drug rituximab (Rituxan.TM., Genentech). Rituximab is a mediator of complement-dependent cytotoxicity (CDC) and ADCC. Cells with higher mean fluorescence intensity by flow cytometry show enhanced sensitivity to rituximab (van Meerten et al., Clin Cancer Res 2006; 12(13):4027-4035, 2006). Enhancement of expression of CD20 by introduction of vector expressing CD20 in CD20 low B cells may facilitate optimal triggering of ADCC.
[0157] For example, a polynucleotide encoding CD20 (SEQ ID NO:29 and 30) can be chemically synthesized (BioBasic Inc., Markham, Canada) and operably inserted directly after the yCD2 gene in the vector pAC3-yCD2(-2) SEQ ID NO: 19 or 22 with a linker sequence as previously described, or as a replacement for the yCD2 gene. In a further example, the CD20 encoding polynucleotide can be synthesized as above and inserted between the IRES and yCD2 sequences or with a linker as previously described. As a further alternative the CD20 sequence can be inserted into the pAC3-yCD2 vector after excision of the CD gene by Psi1 and Not1 digestion.
[0158] In still a further example, a polynucleotide encoding CD20 (SEQ ID NO:29 and 30) can be chemically synthesized (BioBasic Inc., Markham, Canada) and inserted into a vector containing a non amphotropic ENV gene or other appropriate surface protein (Tedder et al., PNAS, 85:208-212, 1988). Alternate ENV and glycoproteins include xenotropic and polytropic ENV and glycoproteins capable of infecting human cells, for example ENV sequences from the NZB strain of MLV and glycoproteins from MCF, VSV, GALV and other viruses [Palu 2000, Baum 2006]. For example, a polynucleotide can comprise a sequence wherein the GAG and POL and yCD2 genes of SEQ ID NO: 19 are deleted, the ENV corresponds to a xenotropic ENV domain of NZB MLV or VSV-g, and the IRES or a promoter such as RSV is operatively linked directly to CD20.
[0159] Mixed infection of cells by VSVG pseudotyped virus and amphotropic retrovirus results in the production of progeny virions bearing the genome of one virus encapsidated by the envelope proteins of the other [Emi 1991]. The same is true for other envelopes that pseudotype retroviral particles. For example, infection by retroviruses derived as above from both SEQ ID NO: 19 or 22 results in production of progeny virions capable of encoding yCD2 and CD20 in infected cells. The resulting viruses can be used to treat a cell proliferative disorder in a subject in combination with Rituxan and/or 5-FC. Similarly, infection of a tumor with a vector encoding only the CD20 marker can make the tumor treatable by the use of Rituxan.
[0160] Levels of the enzymes and cofactors involved in pyrimidine anabolism can be limiting. OPRT, thymidine kinase (TK), Uridine monophosphate kinase, and pyrimidine nucleoside phosphorylase expression is low in 5-FU resistant cancer cells compared to sensitive lines (Wang et al., Cancer Res., 64:8167-8176, 2004). Large population analyses show correlation of enzyme levels with disease outcome (Fukui et al., Int'l. J. OF Mol. Med., 22:709-716, 2008). Coexpression of CD and other pyrimidine anabolism enzymes (PAE) can be exploited to increase the activity and therefore therapeutic index of fluoropyrimidine drugs.
[0161] To further increase the genetic stability (see, e.g., FIG. 5) of yCD2/PAE containing vectors, the enzyme encoding gene can be chemically synthesized with random mutations throughout the sequence. These mutations can be essentially random or can consist of only mutations at the wobble position for each amino acids. The library of mutated sequences is inserted downstream or in place of the yCD2 gene as was previously described for SEQ ID NO: 11 and 13 to create a library of plasmids that can then be used to generate a library of infectious particles by transient transfection of 293T cells or equivalent. Sensitive cells can be infected with retrovirus encoding the fusion polypeptide and subjected to selection with appropriate chemicals.
[0162] DNA shuffling or "molecular breeding" allows genetic information to be shuffled, leading to recombinants with desired properties. Different proteins and enzymes have been improved using DNA shuffling (Stemmer 1994 Proc Natl Acad Sci USA 91(22): 10747-51; Stemmer 1994 Nature 370 (6488):389-91). Genetic recombination is a major force driving the evolution of many viruses. In retrovirus, recombination between two co-packaged retroviral genomes may occur at rates as high as 40% per replication cycle. High rates of recombination at each replication cycle enables genetic information to be shuffled rapidly, leading to recombinants with new pattern of mutations and phenotypes within a short period of time. For example, molecular breeding of retrovirus containing a library of recombinant ecotropic envelope sequences from six murine leukemia virus resulted in a viral clone with a new tropism. Using the same method, several viral clones were selected with improved stability and processing yields (Soong et al., 2000 Nat Genet 25(4):436-9; Powell et al., 2000 Nat Biotechnol 18(12):1279-82). In order to generate vectors that can replicate in cells that are resistant to retroviral infection because of viral restriction or inhibitory factors, such as APOBEC, Trim5alpha, tetherin, Zap or other elements in cells that render them resistant to retroviral infection (D. Wolf & SP. Goff, Annu. Rev. Genet. 2008. 42:143-63) the vectors of the disclosure can be used to express libraries of random peptide libraries normally expressed in yeast libraries (F. Hoppe-Seyler & K. Butz J Mo Med 78:426-430 (2000); R. Wolkowicz et al. J. Biol. Chem. 280:15195-15201 (2005); both incorporated by reference), made from inserts of random nucleotide syntheses into the vectors. These inserts can be expressed as stand-alone peptides expressed from the IRES or otherwise, or can be tagged at the beginning, in the middle of, or at the end of the protein that it is desired to express. For example it is known that yeast cytosine deaminase tolerates fusion to the C terminus of a protein (KN. Barton et al. Mol Ther 13:347-356 2006). Alternatively the peptides can be inserted at the beginning, middle or end of viral structural proteins in the same way. Not all insertion sites will be well tolerated but various useful sites are known. The peptides can be from 6 to 60 amino acids in length, typically between 8 and 20 amino acids. Peptides that bind to and inactivate known antiviral agents may be select by conventional yeast two hybrid methods but this is laborious and has no functional guarantee of success. However, if the virus itself is expressing the peptide library by bulk insertion of library nucleotide sequences into the vector as a DNA plasmid followed by transient transfection on 293T cells or equivalent to generate a library of infectious particles, then the vector that grows best in the target cell or tumor type is selected by serial growth in that cell type or tumor explants. Serial passage in the target cell type rapidly select for viruses that carry an inhibitor peptide for any factor that inhibits viral replication in that cell type. In order to maintain a good diversity of peptides, and supply an opportunity for mutations to occur, if necessary, the target cell is grown mixed with a cell type that supports viral replication well (e.g. HT1080 human fibrosarcoma) to provide further copies of the library of viruses and to let the error prone reverse transcriptase introduce occasional random mutations, that may be advantageous.
[0163] The virus coming from the cells is monitored on each passage for its diversity, by direct sequencing of the insert or otherwise, and may be recycled through the selection procedure several times. The selected viruses are cloned out from the viral supernatant or the cells by PCR across the insert, insertion in a plasmid and sequencing of individual plasmid clones. The identified peptide aptamers are then rechecked for their ability to confer replication advantage in a particular cell type such as a particular tumor type or particular blood cell types etc. The peptide aptamers are also used directly to identify the cellular element with which it interacts by methods known to those in the art.
[0164] The peptide aptamer sequence is then incorporated as before in a viral vector that has the desired replication capacity, and carries a therapeutic gene as described elsewhere in the disclosure. Such a vector is used for therapeutic purposes as described in this disclosure.
[0165] Polynucleotide sequence incorporated into vectors are sometimes unstable resulting in deletion of the polynucleotide sequence from the viral genome over time. The basis for this is not well understood, but it is believed to be sequence dependent Molecular breeding using the vectors described herein to select for recombinant viral clones that have acquired optimal recombinations within the heterologous polynucleotide sequence is employed to select for viral clones that have greater vector stability.
[0166] For example, the HSV-TK coding sequence is not as stable as desired in some situations. Molecular breeding of recombinant retroviral vectors encompass a pool of degenerated coding sequence of HSV-TK is performed to select recombinant vectors that have great vector stability. Randomly mutagenized Herpes Thymidine Kinase (TK) is chemically synthesized (Bio Basic Inc., Markham, Canada). The synthetic sequence is inserted 3' of the yCD2 sequence in SEQ ID NO:19, or by itself in the pAC3-yCD2 vector back bones after excision of the CD2 gene. The retroviral vector mixture is packaged as described elsewhere herein. Mouse fibroblast LMTK- cells or humans 143Tk- are infected with vector and selected for TK activity in HAT media (Hiller et al., Mol. Cell Biol. 8(8):3298-3302, 1988). Serial passage of supernatants of resistant cells to fresh LMTK-/143Tk- cells again selected in HAT media results in selection of stable vectors expressing TK. TK+ resistant cells can be isolated and TK sequences rescued by standard PCR based techniques for mutation analysis (Cowell et al., CDNA Library Protocols, Published by Humana Press, 1996). In this manner, sequences are selected for both expression of functional protein and genomic stability of retroviral vector construct. Similar strategies can be employed for UPRT (SEQ ID NO: 11, 13), OPRT (SEQ ID NO: 15, 17) and other genes of interest. In addition, the serial passage strategy can be used for non-selectable genes and the genomic DNA after serial passage screened for full length inserts by PCR across the IRES-insert gene (see FIG. 5). The full length inserts can be purified and cloned out back into the viral vector then retested. Several cycles of this procedure can be performed to select the most stable gene. This strategy can also be used for passage in animals with or without tumors, and even in patient tissue.
[0167] Alternatively, OPRT, UPRT, TK or other PAE can be expressed in a separate vector containing a different env gene or other appropriate surface glycoprotein. This vector can be replication competent (Logg et al. J. Mol Biol. 369:1214 2007) or non-replicative "first generation" retroviral vector that encodes the envelope and the gene of interest (Emi et al. J. Virol 65:1202 1991). In the latter case the pre-existing viral infection will provide complementary gag and pol to allow infective spread of the "non-replicative" vector from any previously infected cell. Alternate ENV and glycoproteins include xenotropic and polytropic ENV and glycoproteins capable of infecting human cells, for example ENV sequences from the NZB strain of MLV and glycoproteins from MCF, VSV, GALV and other viruses [Palu 2000, Baum 2006, supra]. For example, a polynucleotide can comprise a sequence wherein the GAG and POL genes are deleted, the ENV corresponds to a xenotropic ENV domain from NZB MLV or VSV-g, and the IRES or a promoter such as RSV is operatively linked directly to OPRT, UPRT, TK, or other PAE gene.
[0168] Mixed infection of cells by VSV-g pseudotyped virus and amphotropic retrovirus results in the production of progeny virions bearing the genome of one virus encapsidated by the envelope proteins of the other (Emi et al., J. Virol. 65:1202, 1991). The same is true for other envelopes that pseudotype retroviral particles. For example, infection by retroviruses derived as above from both SEQ ID NO: 19 and 22 results in production of progeny virions capable of encoding yCD2 and OPRT in infected cells. The resulting viruses can be used to treat a cell proliferative disorder in a subject in combination with 5-FC.
[0169] The recombinant retrovirus of the disclosure can be used for the treatment of a neuronal disorder for example, may optionally contain an exogenous gene, for example, a gene which encodes a receptor or a gene which encodes a ligand. Such receptors include receptors which respond to dopamine, GABA, adrenaline, noradrenaline, serotonin, glutamate, acetylcholine and other neuropeptides, as described above. Examples of ligands which may provide a therapeutic effect in a neuronal disorder include dopamine, adrenaline, noradrenaline, acetylcholine, gamma-aminobutyric acid and serotonin. The diffusion and uptake of a required ligand after secretion by an infected donor cell would be beneficial in a disorder where the subject's neural cell is defective in the production of such a gene product. A cell genetically modified to secrete a neurotrophic factor, such as nerve growth factor, (NGF), might be used to prevent degeneration of cholinergic neurons that might otherwise die without treatment.
[0170] Alternatively, cells being grafted into a subject with a disorder of the basal ganglia, such as Parkinson's disease, can be modified to contain an exogenous gene encoding L-DOPA, the precursor to dopamine. Parkinson's disease is characterized by a loss of dopamine neurons in the substantia-nigra of the midbrain, which have the basal ganglia as their major target organ.
[0171] Other neuronal disorders that can be treated similarly by the method of the disclosure include Alzheimer's disease, Huntington's disease, neuronal damage due to stroke, and damage in the spinal cord. Alzheimer's disease is characterized by degeneration of the cholinergic neurons of the basal forebrain. The neurotransmitter for these neurons is acetylcholine, which is necessary for their survival. Engraftment of cholinergic cells infected with a recombinant retrovirus of the disclosure containing an exogenous gene for a factor which would promote survival of these neurons can be accomplished by the method of the disclosure, as described. Following a stroke, there is selective loss of cells in the CA1 of the hippocampus as well as cortical cell loss which may underlie cognitive function and memory loss in these patients. Once identified, molecules responsible for CA1 cell death can be inhibited by the methods of this disclosure. For example, antisense sequences, or a gene encoding an antagonist can be transferred to a neuronal cell and implanted into the hippocampal region of the brain.
[0172] For diseases due to deficiency of a protein product, gene transfer could introduce a normal gene into the affected tissues for replacement therapy, as well as to create animal models for the disease using antisense mutations. For example, it may be desirable to insert a Factor IX encoding nucleic acid into a retrovirus for infection of a muscle or liver cell.
[0173] The disclosure also provides gene therapy for the treatment of cell proliferative or immunologic disorders. Such therapy would achieve its therapeutic effect by introduction of an antisense or dominant negative encoding polynucleotide into cells having the proliferative disorder, wherein the polynucleotide binds to and prevents translation or expression of a gene associated with a cell-proliferative disorder. Delivery of heterologous nucleic acids useful in treating or modulating a cell proliferative disorder (e.g., antisense polynucleotides) can be achieved using a recombinant retroviral vector of the disclosure. In another embodiment, a cell proliferative disorder is treated by introducing a CD polynucleotide of the disclosure, expressing the polynucleotide to produce a polypeptide comprising cytosine deaminase activity and contacting the cell with 5-fluorocytosine in an amount and for a period of time to produce a cytotoxic amount of 5-FU.
[0174] A number of chemotherapeutic agents are currently on the market having varying degrees of success from full remission to temporary remission and prolonged life with expected recurrence. Some of the cancer therapeutic agents on the market target the vascular angiogenic properties of tumor. The composition target the angiogenesis of tumors seeking to reduces blood supply and nutrients to the tumor or cancer and thereby reduce the tumor and prolong a subject's life. VEGF is an angiogenic factor known to play a role in tumor growth. Thus, antagonists of VEGF have been developed as anti-cancer agents.
[0175] Human VEGF mediates neoangiogenesis in normal and malignant vasculature; it is overexpressed in most malignancies and high levels have correlated with a greater risk of metastases and poor prognosis in many. When VEGF interacts with its receptor in in vitro models of angiogenesis, endothelial cell proliferation and new blood vessel formation occur. In animal models, VEGF has been demonstrated to induce vascular endothelial-cell proliferation/migration, sustain survival of newly-formed blood vessels, and enhance vascular permeability.
[0176] A VEGF antagonist agent is one that targets or negatively regulates the VEGF signaling pathway. Examples include VEGF inhibitors (e.g., agents that directly inhibit VEGF (e.g., VEGF-A, -B, -C, or -D), such as by binding VEGF (e.g., anti-VEGF antibodies such as bevacizumab (AVASTIN.RTM.) or ranibizumab (LUCENTIS.RTM.), or other inhibitors such as pegaptanib, NEOVASTAT.RTM., AE-941, VEGF Trap, and PI-88)), modulators of VEGF expression (e.g., INGN-241, oral tetrathiomolybdate, 2-methoxyestradiol, 2-methoxyestradiol nanocrystal dispersion, bevasiranib sodium, PTC-299, Veglin), inhibitors of a VEGF receptor (e.g., KDR or VEGF receptor III (Flt4), for example anti-KDR antibodies, VEGFR2 antibodies such as CDP-791, IMC-1121B, VEGFR2 blockers such as CT-322), modulators of VEGFR expression (e.g., VEGFR1 expression modulator Sirna-027) or inhibitors of VEGF receptor downstream signaling. In some aspects described herein, the VEGF antagonist agent is bevacizumab, pegaptanib, ranibizumab, sorafenib, sunitinib, AE-941, VEGF Trap, pazopanib, vandetanib, vatalanib, cediranib, fenretinide, squalamine, INGN-241, oral tetrathiomolybdate, tetrathiomolybdate, Panzem NCD, 2-methoxyestradiol, AEE-788, AG-013958, bevasiranib sodium, AMG-706, axitinib, BIBF-1120, CDP-791, CP-547632, PI-88, SU-14813, SU-6668, XL-647, XL-999, IMC-1121B, ABT-869, BAY-57-9352, BAY-73-4506, BMS-582664, CEP-7055, CHIR-265, CT-322, CX-3542, E-7080, ENMD-1198, OSI-930, PTC-299, Sirna-027, TKI-258, Veglin, XL-184, or ZK-304709.
[0177] Bevacizumab (AVASTATIN.RTM.) (rhuMAb-VEGF) (Anti-VEGF monoclonal antibody) is a recombinant human/murine chimeric monoclonal antibody directed against vascular endothelial growth factor (VEGF)). It is prepared by engineering VEGF-binding residues of a murine anti-VEGF monoclonal antibody into framework regions of human immunoglobulin-1 (IgG1) (Prod Info Avastin, 2004). Only 7% of the amino acid sequence is derived from the murine antibody, with 93% from IgG1. Bevacizumab binds and neutralizes all human VEGF forms via recognition of binding sites for the two human VEGF receptor types (flt-1 and flk-1). In animal models, the antibody has been shown to stabilize established tumors or suppress tumor growth by inhibiting angiogenesis induced by VEGF.
[0178] The pharmacokinetics of bevacizumab are linear after doses of 0.3 mg/kg or greater (Anon, 2002). Following 90-minute intravenous infusions of 0.3, 1, 3, and 10 mg/kg in advanced cancer patients (n=25), peak serum concentrations of bevacizumab ranged from 5 to 9 mcg/mL, 21 to 39 mcg/mL, 52 to 92 mcg/mL, and 186 to 294 mcg/mL, respectively; slight accumulation was observed with repeat doses (weekly), but this was not significant and pharmacokinetics remained linear. Steady-state levels of bevacizumab were obtained in 100 days after 1 to 20 mg/kg weekly, every 2 weeks, or every 3 week.
[0179] The recommended dose of bevacizumab is 5 milligrams/kilogram infused intravenously over 30 minutes every 2 weeks until disease progression diminishes. Bevacizumab should follow chemotherapy. Efficacy of single-agent bevacizumab has not been established. Bevacizumab (which may be co-administered with the gemcitabine and docetaxel, or within a week before or after chemotherapy), is administered intravenously, at about 1 mg/kg to about 15 mg/kg, preferably about 5 mg/kg.
[0180] The methods and compositions of the disclosure are useful in combination therapies including therapies with bevacizumab. As described herein a replication competent retrovirus (RCR) of the disclosure comprising a therapeutic (e.g., a cytotoxic gene) is useful in treating cell proliferative disorders. An advantage of the RCR of the disclosure includes its ability to infect replicating cells cancer cells. Where the transgene of the vector comprises a cytotoxic gene (e.g., a gene that encodes a polypeptide that converts a non-cytotoxic agent to a cytotoxic agent) provides the ability to kill cancer cells.
[0181] The disclosure provides methods for treating cell proliferative disorders such as cancer and neoplasms comprising administering an RCR vector of the disclosure followed by treatment with a chemotherapeutic agent or anti-cancer agent. In one aspect, the RCR vector is administered to a subject for a period of time prior to administration of the chemotherapeutic or anti-cancer agent that allows the RCR to infect and replicate. The subject is then treated with a chemotherapeutic agent or anti-cancer agent for a period of time and dosage to reduce proliferation or kill the cancer cells. In one aspect, if the treatment with the chemotherapeutic or anti-cancer agent reduces, but does not kill the cancer/tumor (e.g., partial remission or temporary remission), the subject may then be treated with a non-toxic therapeutic agent (e.g., 5-FC) that is converted to a toxic therapeutic agent in cells expression a cytotoxic gene (e.g., cytosine deaminase) from the RCR.
[0182] Using such methods the RCXR vectors of the disclosure are spread during a replication process of the tumor cells, such cells can then be killed by treatment with an anti-cancer or chemotherapeutic agent and further killing can occur using the RCR treatment process described herein.
[0183] In yet another embodiment of the disclosure, the heterologous gene can comprise a coding sequence for a target antigen (e.g., a cancer antigen). In this embodiment, cells comprising a cell proliferative disorder are infected with an RCR comprising a heterologous polynucleotide encoding the target antigen to provide expression of the target antigen (e.g., overexpression of a cancer antigen). An anticancer agent comprising a targeting cognate moiety that specifically interacts with the target antigen is then administered to the subject. The targeting cognate moiety can be operably linked to a cytotoxic agent or can itself be an anticancer agent. Thus, a cancer cell infected by the RCR comprising the targeting antigen coding sequences increases the expression of target on the cancer cell resulting in increased efficiency/efficacy of cytotoxic targeting.
[0184] Blocking of interactions between cells of the immune system has been shown to have significant immunological effects, either activating or suppressing (Waldmann Annu Rev Med. 57:65 2006). For example, blockade of the interaction of CTLA-4 (CD 152) and B7.1 (CD80) which modulates the activation of T cells has been shown to cause immune stimulation, presumably by blocking this suppressive interaction (Peggs et al. Curr. Opin. Immunol. 18:206, 2006). This blockade can potentially be achieved either by antibodies against CTLA-4 or by soluble B7.1. Systemic administration of these types of molecules can have undesirable global effects which can at a minimum lead to deleterious side-effects or even death in the case of one CD28 agonist (Suntharalingam et al. NEJM 355 1018 2006). Pfizer has been developing one such anti-CTLA-4 blockading antibody (CP-675,206) as an anticancer reagent but has recently stopped development because of significant side effects. Local delivery of blockading molecules that are released into the local environment, from the tumor after infection with a replication competent vector encoding such molecules that are released into the extracellular space, provides the immune modulation locally and can avoid these serious side effects. The blockading molecules are antibodies, single chain antibodies, soluble versions of the natural ligand or other peptides that bind such receptors.
[0185] In yet another embodiment, an RCR of the disclosure can comprise a coding sequence comprising a binding domain (e.g., an antibody, antibody fragment, antibody domain or receptor ligand) that specifically interacts with a cognate antigen or ligand. The RCR comprising the coding sequence for the binding domain can then be used to infect cells in a subject comprising a cell proliferative disorder such as a cancer cell or neoplastic cell. The infected cell will then express the binding domain or antibody. An antigen or cognate operably linked to a cytotoxic agent or which is cytotoxic itself can then be administered to a subject. The cytotoxic cognate will then selectively kill infected cells expressing the binding domain. Alternatively the binding domain itself can be an anti-cancer agent.
[0186] As used herein, the term "antibody" refers to a protein that includes at least one immunoglobulin variable domain or immunoglobulin variable domain sequence. For example, an antibody can include a heavy (H) chain variable region (abbreviated herein as VH), and a light (L) chain variable region (abbreviated herein as VL). In another example, an antibody includes two heavy (H) chain variable regions and two light (L) chain variable regions. The term "antibody" encompasses antigen-binding fragments of antibodies (e.g., single chain antibodies, Fab fragments, F(ab').sub.2, a Fd fragment, a Fv fragments, and dAb fragments) as well as complete antibodies.
[0187] The VH and VL regions can be further subdivided into regions of hypervariability, termed "complementarity determining regions" (CDR), interspersed with regions that are more conserved, termed "framework regions" (FR). The extent of the framework region and CDRs has been precisely defined (see, Kabat, E. A., et al. (1991) Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242, and Chothia, C. et al. (1987) J. Mol. Biol. 196:901-917). Kabat definitions are used herein. Each VH and VL is typically composed of three CDRs and four FRs, arranged from amino-terminus to carboxy-terminus in the following order: FR1, CDR1, FR2, CDR2, FR3, CDR3, FR4.
[0188] An "immunoglobulin domain" refers to a domain from the variable or constant domain of immunoglobulin molecules. Immunoglobulin domains typically contain two .beta.-sheets formed of about seven .beta.-strands, and a conserved disulphide bond (see, e.g., A. F. Williams and A. N. Barclay 1988 Ann. Rev Immunol. 6:381-405). The canonical structures of hypervariable loops of an immunoglobulin variable can be inferred from its sequence, as described in Chothia et al. (1992) J. Mol. Biol. 227:799-817; Tomlinson et al. (1992) J. Mol. Biol. 227:776-798); and Tomlinson et al. (1995) EMBO J. 14(18):4628-38.
[0189] As used herein, an "immunoglobulin variable domain sequence" refers to an amino acid sequence which can form the structure of an immunoglobulin variable domain. For example, the sequence may include all or part of the amino acid sequence of a naturally-occurring variable domain. For example, the sequence may omit one, two or more N- or C-terminal amino acids, internal amino acids, may include one or more insertions or additional terminal amino acids, or may include other alterations. In one embodiment, a polypeptide that includes immunoglobulin variable domain sequence can associate with another immunoglobulin variable domain sequence to form a target binding structure (or "antigen binding site"), e.g., a structure that interacts with Tiel, e.g., binds to or inhibits Tiel.
[0190] The VH or VL chain of the antibody can further include all or part of a heavy or light chain constant region, to thereby form a heavy or light immunoglobulin chain, respectively. In one embodiment, the antibody is a tetramer of two heavy immunoglobulin chains and two light immunoglobulin chains, wherein the heavy and light immunoglobulin chains are inter-connected by, e.g., disulfide bonds. The heavy chain constant region includes three domains, CH1, CH2 and CH3. The light chain constant region includes a CL domain. The variable region of the heavy and light chains contains a binding domain that interacts with an antigen. The constant regions of the antibodies typically mediate the binding of the antibody to host tissues or factors, including various cells of the immune system (e.g., effector cells) and the first component (C1q) of the classical complement system. The term "antibody" includes intact immunoglobulins of types IgA, IgG, IgE, IgD, IgM (as well as subtypes thereof). The light chains of the immunoglobulin may be of types kappa or lambda. In one embodiment, the antibody is glycosylated. An antibody can be functional for antibody-dependent cytotoxicity and/or complement-mediated cytotoxicity.
[0191] The term "monospecific antibody" refers to an antibody that displays a single binding specificity and affinity for a particular target, e.g., epitope. This term includes a "monoclonal antibody" which refers to an antibody that is produced as a single molecular species, e.g., from a population of homogenous isolated cells. A "monoclonal antibody composition" refers to a preparation of antibodies or fragments thereof of in a composition that includes a single molecular species of antibody. In one embodiment, a monoclonal antibody is produced by a mammalian cell. One or more monoclonal antibody species may be combined.
[0192] One or more regions of an antibody can be human or effectively human. For example, one or more of the variable regions can be human or effectively human. For example, one or more of the CDRs can be human, e.g., HC CDR1, HC CDR2, HC CDR3, LC CDR1, LC CDR2, and LC CDR3. Each of the light chain CDRs can be human. HC CDR3 can be human. One or more of the framework regions can be human, e.g., FR1, FR2, FR3, and FR4 of the HC or LC. In one embodiment, all the framework regions are human, e.g., derived from a human somatic cell, e.g., a hematopoietic cell that produces immunoglobulins or a non-hematopoietic cell. In one embodiment, the human sequences are germline sequences, e.g., encoded by a germline nucleic acid. One or more of the constant regions can be human or effectively human. In another embodiment, at least 70, 75, 80, 85, 90, 92, 95, or 98% of the framework regions (e.g., FR1, FR2, and FR3, collectively, or FR1, FR2, FR3, and FR4, collectively) or the entire antibody can be human or effectively human. For example, FR1, FR2, and FR3 collectively can be at least 70, 75, 80, 85, 90, 92, 95, 98, or 99% identical to a human sequence encoded by a human germline V segment of a locus encoding a light or heavy chain sequence.
[0193] All or part of an antibody can be encoded by an immunoglobulin gene or a segment thereof. Exemplary human immunoglobulin genes include the kappa, lambda, alpha (IgA1 and IgA2), gamma (IgG1, IgG2, IgG3, IgG4), delta, epsilon and mu constant region genes, as well as the myriad immunoglobulin variable region genes. Full-length immunoglobulin light chains (about 25 Kd or 214 amino acids) are encoded by a variable region gene at the NH2-terminus (about 110 amino acids) and a kappa or lambda constant region gene at the COOH-- terminus. Full-length immunoglobulin heavy chains (about 50 Kd or 446 amino acids), are similarly encoded by a variable region gene (about 116 amino acids) and one of the other aforementioned constant region genes, e.g., gamma (encoding about 330 amino acids). A light chain refers to any polypeptide that includes a light chain variable domain. A heavy chain refers to any polypeptide that a heavy chain variable domain.
[0194] The term "antigen-binding fragment" of a full-length antibody (or simply "antibody portion," or "fragment"), as used herein, refers to one or more fragments of a full-length antibody that retain the ability to specifically bind to a target of interest. Examples of binding fragments encompassed within the term "antigen-binding fragment" of a full length antibody include (i) a Fab fragment, a monovalent fragment consisting of the VL, VH, CL and CH1 domains; (ii) a F(ab').sub.2 fragment, a bivalent fragment including two Fab fragments linked by a disulfide bridge at the hinge region; (iii) a Fd fragment consisting of the VH and CH1 domains; (iv) a Fv fragment consisting of the VL and VH domains of a single arm of an antibody, (v) a dAb fragment (Ward et al., (1989) Nature 341:544-546), which consists of a VH domain; and (vi) an isolated complementarity determining region (CDR) that retains functionality. Furthermore, although the two domains of the Fv fragment, VL and VH, are coded for by separate genes, they can be joined, using recombinant methods, by a synthetic linker that enables them to be made as a single protein chain in which the VL and VH regions pair to form monovalent molecules known as single chain Fv (scFv). See e.g., Bird et al. (1988) Science 242:423-426; and Huston et al. (1988) Proc. Natl. Acad. Sci. USA 85:5879-5883.
[0195] Antibody fragments can be obtained using any appropriate technique including conventional techniques known to those with skill in the art. The term "monospecific antibody" refers to an antibody that displays a single binding specificity and affinity for a particular target, e.g., epitope. This term includes a "monoclonal antibody" or "monoclonal antibody composition," which as used herein refer to a preparation of antibodies or fragments thereof of single molecular composition. As used herein, "isotype" refers to the antibody class (e.g., IgM or IgG1) that is encoded by heavy chain constant region genes.
[0196] The disclosure provides a method of treating a subject having a cell proliferative disorder. The subject can be any mammal, and is preferably a human. The subject is contacted with a recombinant replication competent retroviral vector of the disclosure. The contacting can be in vivo or ex vivo. Methods of administering the retroviral vector of the disclosure are known in the art and include, for example, systemic administration, topical administration, intraperitoneal administration, intra-muscular administration, intracranial, cerebrospinal, as well as administration directly at the site of a tumor or cell-proliferative disorder. Other routes of administration known in the art.
[0197] Thus, the disclosure includes various pharmaceutical compositions useful for treating a cell proliferative disorder. The pharmaceutical compositions according to the disclosure are prepared by bringing a retroviral vector containing a heterologous polynucleotide sequence useful in treating or modulating a cell proliferative disorder according to the disclosure into a form suitable for administration to a subject using carriers, excipients and additives or auxiliaries. Frequently used carriers or auxiliaries include magnesium carbonate, titanium dioxide, lactose, mannitol and other sugars, talc, milk protein, gelatin, starch, vitamins, cellulose and its derivatives, animal and vegetable oils, polyethylene glycols and solvents, such as sterile water, alcohols, glycerol and polyhydric alcohols. Intravenous vehicles include fluid and nutrient replenishers. Preservatives include antimicrobial, anti-oxidants, chelating agents and inert gases. Other pharmaceutically acceptable carriers include aqueous solutions, non-toxic excipients, including salts, preservatives, buffers and the like, as described, for instance, in Remington's Pharmaceutical Sciences, 15th ed. Easton: Mack Publishing Co., 1405-1412, 1461-1487 (1975) and The National Formulary XIV., 14th ed. Washington: American Pharmaceutical Association (1975), the contents of which are hereby incorporated by reference. The pH and exact concentration of the various components of the pharmaceutical composition are adjusted according to routine skills in the art. See Goodman and Gilman's The Pharmacological Basis for Therapeutics (7th ed.).
[0198] For example, and not by way of limitation, a retroviral vector useful in treating a cell proliferative disorder will include an amphotropic ENV protein, GAG, and POL proteins, a promoter sequence in the U3 region retroviral genome, and all cis-acting sequence necessary for replication, packaging and integration of the retroviral genome into the target cell.
[0199] The following Examples are intended to illustrate, but not to limit the disclosure. While such Examples are typical of those that might be used, other procedures known to those skilled in the art may alternatively be utilized.
EXAMPLES
Example 1. Modification of Vector Backbone of pACE-GFPemd to pAC3-GFPemd and Insertion of Cytosine Deaminase Gene Sequences in Place of GFP
[0200] The previous back bone of the pACE-GFPemd plasmid (U.S. Pat. No. 6,899,871, Wang et al. Hum Gene Ther 14:117 2003) was modified in 3 ways as shown in FIG. 3E. The modifications were made by PCR-mediated, oligonucleotide-directed mutagenesis (Logg et al., J. Mol Biol 369: 1214, 2007; see also "Molecular Biology and Biotechnology" Eds. J M. Walker, R. Rapley, Royal Society of Chemistry, London UK, 2000). The following modifications were made. 1) The nucleic acid sequence at the p15 region at 3' end of the amphotropic env gene was originally derived from the ecotropic envelope--this sequence was replaced by the corresponding sequence from the 4070A amphotropic envelope; the encoded envelope amino acids are identical in the two constructs. 2) The IRES sequence 3' end was modified to allow easier insertion of transgenes of choice with insertion of a PstI1 site and small imperfect repeats at either end of the IRES transgene site were removed. 3) Residual viral sequence downstream of the 3'LTR was removed. The resultant plasmid is pACE-emdGFP (aka pACE-GFP, pACE-eGFP and T5.0006) was used as a basis for the vectors encoding cytosine deaminase and variants. Two methods of inserting the coding sequence cassettes were used initially. The first method resulted in the sequence 5'TTATAAT3' (SEQ ID NO:73), and the second in the sequence 5'TTATAA3' (SEQ ID NO:74) immediately upstream of the ATG start codon. The second method was simpler, as it involved simple PstI1 and Not1 enzyme cuts in the vector and the synthetic cytosine deaminase genes, followed by religation. Vectors with cytosine deaminase inserts were made both ways with the CDopt (CD1) and the CDopt+3pt (CD2) (see FIG. 2) coding sequences and infectious virus preps made by transient transfection of 293T cells as described in Example 3. U87 cells were then infected in culture, at an MOI of 0.1, and the cells grown until 100% infected. Cell extracts of 100% infected cells were assayed for cytosine deaminase activity as described in Example 6 and the specific activity of the enzyme was found to be equivalent for constructs with either upstream sequence, that were otherwise identical. Therefore in the table in FIG. 2, pACE-eGFP (T5.0006) and pACE-yCD (T5.0007) have the first upstream sequence, while all other constructs that were further tested have the second. Subsequently vectors with different gene inserts have been routinely constructed with straightforward PStI1 and Not 1 cuts.
[0201] See FIG. 3A below for a diagram of the vector construct for the initial transfected replication-competent retrovirus. CMV is the human CMV immediate early promoter, U3, R and U5 are the corresponding regions of the viral long terminal repeat (LTR). Gag, pol and env are the viral protein coding regions. FIGS. 3B and 3D shows the plasmid structure and a sequence of the disclosure.
[0202] The vector of the disclosure provides a number of differences compared to the vector of Tai et al., Mol. Ther. 12:842, 2005. The Tai et al. vector has been altered to eliminate about 70 bp of MLV sequence downstream from the 3'LTR. The DNA sequence downstream of the Clal site in the envelope was changed to an amphotropic envelope sequence. This change does not change the amino acid sequence of the envelope. In addition, small repeats on either side of the IRES-CD cassette have been eliminated to avoid instability due to homologous recombination. These changes also unexpectedly provided increased stability of the vector during replication and passaging in host cells (FIG. 5).
[0203] It is recognized that after reverse transcription and the first integration event into treated cells, the DNA provirus and any subsequent progeny retrovirus has a conventional LTR structure from MLV on either end. This configuration has been shown to be stable after multiple cycles of infection (See FIG. 5 below).
Example 2. Genetic Enhancements to the Wild Type Yeast Cytosine Deaminase Gene
[0204] Two sets of changes have been made: (1) three positional mutations which change three amino acids (A23L, I140L and V108I) to increase thermal stability of the yeast cytosine deaminase protein and (2) additional gene sequence modifications to enhance human codon usage sequences to improve protein translation efficiency in human cells without further changes to the amino acid sequence.
[0205] Sequence design for CD included CD-optimized, CD-UPRT (+/- linker) and CD-OPRTase (+/- linker). The final cytosine deaminase coding sequence can comprise at the 5' end a PSI1 site (full length) and 3' end Not1 site plus poly A tail for PSIl/Not1 cassette based strategy. Sequences cassettes were ordered from, and provided by, a commercial vendor (BioBasic Inc., Markham, Ontario, Canada).
[0206] The following sequence comprising a yeast cytosine deaminase was used for cloning, optimizing and mutation (the boxed nucleic acids comprise the restriction sites--Psi1 and Not1--used in subsequent methods for cloning:
TABLE-US-00002 (SEQ ID NO: 43) ##STR00001## AGGGTATGGACATTGCCTATGAGGAGGCGGCCTTAGGTTACAAAGAGGGT GGTGTTCCTATTGGCGGATGTCTTATCAATAACAAAGACGGAAGTGTTCT CGGTCGTGGTCACAACATGAGATTTCAAAAGGGATCCGCCACACTACATG GTGAGATCTCCACTTTGGAAAACTGTGGGAGATTAGAGGGCAAAGTGTAC AAAGATACCACTTTGTATACGACGCTGTCTCCATGCGACATGTGTACAGG TGCCATCATCATGTATGGTATTCCACGCTGTGTTGTCGGTGAGAACGTTA ATTTCAAAAGTAAGGGCGAGAAATATTTACAAACTAGAGGTCACGAGGTT GTTGTTGTTGACGATGAGAGGTGTAAAAAGATCATGAAACAATTTATCGA ##STR00002## CATAGATAAAATAAAAGATTTTATTTAGTCTCCAGAAAAAGGGGGG
[0207] The following Table summarizes the genes and resulting plasmid vectors that were made and their names.
TABLE-US-00003 TABLE Vector constructs and names Identity Reference Original 5'LTR Code name Name Prom Envelope Vector IRES Transgene 3'LTR T5.0000 pACE-yCD pACE-CD CMV Ampho pACE EMCV Wt yeast MLV U3 (Tai et al. (4070A) CD 2005) T5.0001 pAC3-yCD1 CDopt CMV Ampho pAC3 EMCV modified MLV U3 sequence (4070A) CD T5.0002 pAC3-yCD2 CDopt + 3pt CMV Ampho pAC3 EMCV Modified MLV U3 (4070A) CD T5.0003 pAC3-yCD2-U Cdopt + 3pt- CMV Ampho pAC3 EMCV CD2-UPRT MLV U3 UPRT (4070A) T5.0004 pAC3-yCD2-O CDopt + 3pt- CMV Ampho pAC3 EMCV CD2-OPRT MLV U3 OPRT (4070A) T5.0005 pAC3-yCD2-LO CDopt + 3pt- CMV Ampho pAC3 EMCV CD2-L-OPRT MLV U3 LINK-OPRT (4070A) T5.0006 pAC3-eGFP pAC3-emd, CMV Ampho pAC3 EMCV Emerald MLV U3 pAC3GFP (4070A) GFP T5.0007 pAC3-yCD pAC3-yCD CMV Ampho pAC3 EMCV Wt yeast MLV U3 (4070A) CD
[0208] The replication competent retroviral vector described by Kasahara et al. pACE-CD (U.S. Pat. No. 6,899,871, the disclosure of which is incorporated herein) was used as a basis for additional modifications. A vector (pAC3-yCD) was modified to express a modified yeast cytosine deaminase gene as described herein and was used in the constructs. See FIG. 3A below for a diagram of the vector construct for the initial transfected replication-competent retrovirus. CMV is the human CMV immediate early promoter, U3, R and U5 are the corresponding regions of the viral long terminal repeat (LTR). Gag, pol and env are the viral protein coding regions. FIGS. 3B and 3D shows the plasmid structure and a sequence of the disclosure.
[0209] After the genes were synthesized at a contractor (Bio Basic Inc., Markham, Ontario, Canada) they were inserted into the Psi1-Not1 site of the pAC3 vector backbone (FIG. 3). The plasmid backbone was normally generated by cutting the plasmid pAC3-eGFP with Psi1 and Not1 and purifying the large (about 11 kb) fragment encoding the plasmid and retroviral backbone).
[0210] A. Humanized Codon Optimized CD Gene (CDopt, Aka CD1, T5.0001).
[0211] A comparison of a human codon optimized cytosine deaminase of Conrad et al. and PCT WO 99/60008 indicates 91 total codons optimized in both, 36 codons identical, 47 codons had third base pair changes (all encode same amino acid) and 9 codons were different (however they encoded same amino acid). Of the 9 codons that differed:
[0212] AGC (Ser) to TCC (Ser)
[0213] CGT (Arg) to AGG (Arg)
[0214] CCA (Pro) to CCT (Pro)
[0215] All have equivalent GC content and encode the same amino acid. The native yeast gene sequence above was separately codon optimized to give the following CD gene (CD1) and was called T5.0001 when inserted into the plasmid vector pAC3 which encodes the replication competent retrovirus (RCR) with IRES.
TABLE-US-00004 (SEQ ID NO: 44) ##STR00003## GGATATCGCTTACGAGGAGGCCGCCCTGGGCTACAAGGAGGGCGGCGTGC CTATCGGCGGCTGTCTGATCAACAACAAGGACGGCAGTGTGCTGGGCAGG GGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACCCTGCACGGCGAGAT CTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTACAAGGACA CCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGGCGCTATC ATCATGTACGGCATCCCTAGGTGTGTGGTGGGCGAGAACGTGAACTTCAA GTCCAAGGGCGAGAAGTACCTGCAAACCAGGGGCCACGAGGTGGTGGTTG TTGACGATGAGAGGTGTAAGAAGATCATGAAGCAGTTCATCGACGAGAGG ##STR00004## AATAAAAGATTTTATTTAGTCTCCAGAAAAAGGGGGG.
[0216] B. Heat Stabilized CD Gene.
[0217] Additional modifications were made to enhance the stability of the cytosine deaminase. Genetic enhancements to the wild type yeast cytosine deaminase gene were made to include three positional mutations which change three amino acids (A23L, I140L and V108I) to increase thermal stability of the yeast cytosine deaminase protein.
[0218] The following primer pairs were used in the generation of the gene for the cytosine deaminase polypeptide of the disclosure: Site directed mutagenesis primers:
TABLE-US-00005 Primers sense: (SEQ ID NO: 45) 5'-tcgaggatatcggcgagtgaaacccgttattctttttggc-3' Primers antisense: (SEQ ID NO: 46) 5'-gccaaaaagaataacgggtttcactcgccgatatcctcga-3' Primers sense: (SEQ ID NO: 47) 5'tcggcgagtgatccggcggcggcgcctccggcggcggcgcctccggcg gcggcgcctccggcggcggcgccaacccgttatt-3' Primers antisense: (SEQ ID NO: 48) 5'-aataacgggttggcgccgccgccggaggcgccgccgccggaggcgcc gccgccggaggcgccgccgccggatcactcgccga-3'
[0219] To increase the stability of the native yeast CD protein, three amino acid substitutions were engineered into the protein. These substitutions were alone or in combination with human codon optimization.
[0220] The three amino acid substitutions are: A23L, V108I, I140L. A sequence encoding these substitutions is shown below.
TABLE-US-00006 (SEQ ID NO: 3) ATGGTGACAGGGGGAATGGCAAGCAAGTGGGATCAGAAGGGTATGGACAT TGCCTATGAGGAGGCGTTATTAGGTTACAAAGAGGGTGGTGTTCCTATTG GCGGATGTCTTATCAATAACAAAGACGGAAGTGTTCTCGGTCGTGGTCAC AACATGAGATTTCAAAAGGGATCCGCCACACTACATGGTGAGATCTCCAC TTTGGAAAACTGTGGGAGATTAGAGGGCAAAGTGTACAAAGATACCACTT TGTATACGACGCTGTCTCCATGCGACATGTGTACAGGTGCCATCATCATG TATGGTATTCCACGCTGTGTCATCGGTGAGAACGTTAATTTCAAAAGTAA GGGCGAGAAATATTTACAAACTAGAGGTCACGAGGTTGTTGTTGTTGACG ATGAGAGGTGTAAAAAGTTAATGAAACAATTTATCGATGAAAGACCTCAG GATTGGTTTGAAGATATTGGTGAGTAG
[0221] The encoded polypeptide comprises the following sequence (substituted amino acids in underlined):
TABLE-US-00007 (SEQ ID NO: 4) 1 MVTGGMASKWDQKGMDIAYEEALLGYKEGGVPIGGCLINNKDGSVLGRGHNMRFQKGSAT 61 LHGEISTLENCGRLEGKVYKDTTLYTTLSPCDMCTGAIIMYGIPRCVIGENVNFKSKGEK 121 YLQTRGHEVVVVDDERCKKLMKQFIDERPQDWFEDIGE-
[0222] Final construct design that integrates 3 amino acid substitutions A23L/V108I/I140L utilizing preferred codons and uses preferred human codon usage for entire sequence (this gene is called CDopt+3pt [aka CD2] (SEQ ID NO:49):
TABLE-US-00008 1 ATGGTGACCGGCGGCATGGCCTCCAAGTGGGATCAAAAGGGCATGGATATCGCTTACGAG 61 GAGGCCCTGCTGGGCTACAAGGAGGGCGGCGTGCCTATCGGCGGCTGTCTGATCAACAAC 121 AAGGACGGCAGTGTGCTGGGCAGGGGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACC 181 CTGCACGGCGAGATCTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTACAAG 241 GACACCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGGCGCTATCATCATG 301 TACGGCATCCCTAGGTGTGTGATCGGCGAGAACGTGAACTTCAAGTCCAAGGGCGAGAAG 361 TACCTGCAAACCAGGGGCCACGAGGTGGTGGTTGTTGACGATGAGAGGTGTAAGAAGCTG 421 ATGAAGCAGTTCATCGACGAGAGGCCTCAGGACTGGTTCGAGGATATCGGCGAGTGATAA
[0223] T5.0002 refers to the above modified CD when inserted into the plasmid vector pAC3 which encodes the RCR with IRES. Underlined codons denotes preferred codons for amino acid substitutions.
[0224] Protein translation sequence alignment indicates preferred codon changes and amino acid substitutions result in desired protein structure:
CD-optimized sequence design (human codon preference+3 amino acid substitutions)
TABLE-US-00009 (SEQ ID NO: 50) ##STR00005## AGGGCATGGATATCGCTTACGAGGAGGCCCTGCTGGGCTACAAGGAGGGC GGCGTGCCTATCGGCGGCTGTCTGATCAACAACAAGGACGGCAGTGTGCT GGGCAGGGGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACCCTGCACG GCGAGATCTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTAC AAGGACACCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGG CGCTATCATCATGTACGGCATCCCTAGGTGTGTGATCGGCGAGAACGTGA ACTTCAAGTCCAAGGGCGAGAAGTACCTGCAAACCAGGGGCCACGAGGTG GTGGTTGTTGACGATGAGAGGTGTAAGAAGCTGATGAAGCAGTTCATCGA ##STR00006## CATAGATAAAATAAAAGATTTTATTTAGTCTCCAGAAAAAGGGGGG
[0225] C. Construction of CD-UPRT Fusion Gene (CDopt+3pt-UPRT, [Aka CDopt-UPRT and CD2-UPRT], T5.0003 in the pAC3 Plasmid RCR Vector).
[0226] A fusion construct was also developed comprising a CD polypeptide as described above linked to a UPRT polypeptide to generate a CD-optimized-UPRT sequence using Scheme I as set forth in FIG. 10A. The following primers were used to delete the stop-start between the CD and UPRT.
Primer Sequences:
TABLE-US-00010 [0227] Primer Name Primer Sequence (5' to 3') (SEQ ID NO:) del118-123 5'-tcgaggatatcggcgagtgaaacccgttattctttttggc-3' (51) del118-123-antisense 5'-gccaaaaagaataacgggtttcactcgccgatatcctcga-3' (52) Energy Length Cost of Primer Name (nt.) Tm Duplex Energy at 68.degree. C. Mismatches del118-123 40 79.06.degree. C. -44.37 kcal/mole 21.1% del118-123-antisense 40 79.06.degree. C. -47.95 kcal/mole 20.3% Primer Name Primer-Template Duplex del118-123 (SEQ ID 5'-tcgaggatatcggcgagtga------aacccgttattctttttggc-3' NOs: 51 and 53, |||||||||||||||||||| |||||||||||||||||||| respectively) ccaagctcctatagccgctcactatctacttgggcaataagaaaaaccgaag del118-123-antisense ggttcgaggatatcggcgagtgatagatgaacccgttattctttttggcttc (SEQ ID NO: 54 and 52 |||||||||||||||||||| |||||||||||||||||||| respectively) 3'-agctcctatagccgctcact------ttgggcaataagaaaaaccg
[0228] The resulting fusion polynucleotide comprises 1296 bp and the sequence set forth immediately below:
TABLE-US-00011 (SEQ ID NO: 55) ##STR00007## AGGGCATGGATATCGCTTACGAGGAGGCCCTGCTGGGCTACAAGGAGGGC GGCGTGCCTATCGGCGGCTGTCTGATCAACAACAAGGACGGCAGTGTGCT GGGCAGGGGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACCCTGCACG GCGAGATCTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTAC AAGGACACCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGG CGCTATCATCATGTACGGCATCCCTAGGTGTGTGATCGGCGAGAACGTGA ACTTCAAGTCCAAGGGCGAGAAGTACCTGCAAACCAGGGGCCACGAGGTG GTGGTTGTTGACGATGAGAGGTGTAAGAAGCTGATGAAGCAGTTCATCGA CGAGAGGCCTCAGGACTGGTTCGAGGATATCGGCGAGAACCCGTTATTCT TTTTGGCTTCTCCATTCTTGTACCTTACATATCTTATATATTATCCAAAC AAAGGGTCTTTCGTTAGCAAACCTAGAAATCTGCAAAAAATGTCTTCGGA ACCATTTAAGAACGTCTACTTGCTACCTCAAACAAACCAATTGCTGGGTT TGTACACCATCATCAGAAATAAGAATACAACTAGACCTGATTTCATTTTC TACTCCGATAGAATCATCAGATTGTTGGTTGAAGAAGGTTTGAACCATCT ACCTGTGCAAAAGCAAATTGTGGAAACTGACACCAACGAAAACTTCGAAG GTGTCTCATTCATGGGTAAAATCTGTGGTGTTTCCATTGTCAGAGCTGGT GAATCGATGGAGCAAGGATTAAGAGACTGTTGTAGGTCTGTGCGTATCGG TAAAATTTTAATTCAAAGGGACGAGGAGACTGCTTTACCAAAGTTATTCT ACGAAAAATTACCAGAGGATATATCTGAAAGGTATGTCTTCCTATTAGAC CCAATGCTGGCCACCGGTGGTAGTGCTATCATGGCTACAGAAGTCTTGAT TAAGAGAGGTGTTAAGCCAGAGAGAATTTACTTCTTAAACCTAATCTGTA GTAAGGAAGGGATTGAAAAATACCATGCCGCCTTCCCAGAGGTCAGAATT GTTACTGGTGCCCTCGACAGAGGTCTAGATGAAAACAAGTATCTAGTTCC ##STR00008## CATAGATAAAATAAAAGATTTTATTTAGTCTCCAGAAAAAGGGGGG
[0229] D. Construction of CD-Linker UPRT Fusion Gene (CDopt+3pt-LINK-UPRT [Aka CDopt-LINKER-UPRT and CD2-L-UPRT].
[0230] A fusion construct was also developed by cloning a linker (Ser-Gly-Gly-Gly-Gly).sub.4 (SEQ ID NO:56) domain between and in frame with the CD polypeptide and the UPRT polypeptide to generated a CD-optimized-linker-UPRT sequence using Scheme II as depicted in FIG. 10B. The following primers were used to insert the linker.
TABLE-US-00012 Primer Name Primer Sequence (5' to 3') (SEQ ID NO:) ins_60nt_after_477 5'- tcggcgagtgatccggcggcggcgcctccggcggcggcgcctccggcg gcggcgcctccggcggcggcgccaacccgttatt-3' (57) ins_60nt_after_477- 5'- antisense aataacgggttggcgccgccgccggaggcgccgccgccggaggcgcc gccgccggaggcgccgccgccggatcactcgccga-3' (58) Energy Length Cost of Primer Name (nt.) Tm Duplex Energy at 68.degree. C. Mismatches ins_60nt_after_477 82 79.77.degree. C. -30.19 kcal/mole 83.3% ins_60nt_after_477- 82 79.77.degree. C. -32.31 kcal/mole 82.2% antisense
[0231] The resulting construct has size: 1356 bp and the sequence immediately below:
TABLE-US-00013 (SEQ ID NO: 59) ##STR00009## AGGGCATGGATATCGCTTACGAGGAGGCCCTGCTGGGCTACAAGGAGGGC GGCGTGCCTATCGGCGGCTGTCTGATCAACAACAAGGACGGCAGTGTGCT GGGCAGGGGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACCCTGCACG GCGAGATCTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTAC AAGGACACCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGG CGCTATCATCATGTACGGCATCCCTAGGTGTGTGATCGGCGAGAACGTGA ACTTCAAGTCCAAGGGCGAGAAGTACCTGCAAACCAGGGGCCACGAGGTG GTGGTTGTTGACGATGAGAGGTGTAAGAAGCTGATGAAGCAGTTCATCGA CGAGAGGCCTCAGGACTGGTTCGAGGATATCGGCGAGTCCGGCGGCGGCG CCTCCGGCGGCGGCGCCTCCGGCGGCGGCGCCTCCGGCGGCGGCGCCAAC CCGTTATTCTTTTTGGCTTCTCCATTCTTGTACCTTACATATCTTATATA TTATCCAAACAAAGGGTCTTTCGTTAGCAAACCTAGAAATCTGCAAAAAA TGTCTTCGGAACCATTTAAGAACGTCTACTTGCTACCTCAAACAAACCAA TTGCTGGGTTTGTACACCATCATCAGAAATAAGAATACAACTAGACCTGA TTTCATTTTCTACTCCGATAGAATCATCAGATTGTTGGTTGAAGAAGGTT TGAACCATCTACCTGTGCAAAAGCAAATTGTGGAAACTGACACCAACGAA AACTTCGAAGGTGTCTCATTCATGGGTAAAATCTGTGGTGTTTCCATTGT CAGAGCTGGTGAATCGATGGAGCAAGGATTAAGAGACTGTTGTAGGTCTG TGCGTATCGGTAAAATTTTAATTCAAAGGGACGAGGAGACTGCTTTACCA AAGTTATTCTACGAAAAATTACCAGAGGATATATCTGAAAGGTATGTCTT CCTATTAGACCCAATGCTGGCCACCGGTGGTAGTGCTATCATGGCTACAG AAGTCTTGATTAAGAGAGGTGTTAAGCCAGAGAGAATTTACTTCTTAAAC CTAATCTGTAGTAAGGAAGGGATTGAAAAATACCATGCCGCCTTCCCAGA GGTCAGAATTGTTACTGGTGCCCTCGACAGAGGTCTAGATGAAAACAAGT ATCTAGTTCCAGGGTTGGGTGACTTTGGTGACAGATACTACTGTGTTTAA ##STR00010## AGGGGGG
[0232] E. Construction of CD-OPRT Fusion Gene (CDopt+3pt-OPRT [Aka CDopt-OPRT and CD2-OPRT], T5.0004 when Inserted into the pAC3 Plasmid RCR Vector).
[0233] A fusion construct was also developed comprising a CD polypeptide as described above linked to an OPRT polypeptide to generated a CD-optimized-OPRTase (CD humanized+3ptmutation+OPRTase functional domain human) using Scheme III as shown in FIG. 10C.
[0234] The resulting construct comprises a size of 1269 bp and the sequence immediately below:
TABLE-US-00014 (SEQ ID NO: 60) ##STR00011## AGGGCATGGATATCGCTTACGAGGAGGCCCTGCTGGGCTACAAGGAGGGC GGCGTGCCTATCGGCGGCTGTCTGATCAACAACAAGGACGGCAGTGTGCT GGGCAGGGGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACCCTGCACG GCGAGATCTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTAC AAGGACACCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGG CGCTATCATCATGTACGGCATCCCTAGGTGTGTGATCGGCGAGAACGTGA ACTTCAAGTCCAAGGGCGAGAAGTACCTGCAAACCAGGGGCCACGAGGTG GTGGTTGTTGACGATGAGAGGTGTAAGAAGCTGATGAAGCAGTTCATCGA CGAGAGGCCTCAGGACTGGTTCGAGGATATCGGCGAGGCGGTCGCTCGTG cagctttggggccattggtgacgggtctgtacgacgtgcaggctttcaag tttggggacttcgtgctgaagagcgggctttcctcccccatctacatcga tctgcggggcatcgtgtctcgaccgcgtcttctgagtcaggttgcagata ttttattccaaactgcccaaaatgcaggcatcagttttgacaccgtgtgt ggagtgccttatacagctttgccattggctacagttatctgttcaaccaa tcaaattccaatgcttattagaaggaaagaaacaaaggattatggaacta agcgtcttgtagaaggaactattaatccaggagaaacctgtttaatcatt gaagatgttgtcaccagtggatctagtgttttggaaactgttgaggttct tcagaaggagggcttgaaggtcactgatgccatagtgctgttggacagag agcagggaggcaaggacaagttgcaggcgcacgggatccgcctccactca gtgtgtacattgtccaaaatgctggagattctcgagcagcagaaaaaagt tgatgctgagacagttgggagagtgaagaggtttattcaggagaatgtct ttgtggcagcgaatcataatggttctcccctttctataaaggaagcaccc aaagaactcaGCTTCGGTGCACGTGCAGAGCTGCCCAGGATCCACCCAGT ##STR00012## GTCTCCAGAAAAAGGGGGG
[0235] F. Construction of CD-Linker-OPRT Fusion Gene (CDopt+3pt-LINK-OPRT, [Aka CDopt-LINKER-OPRT and CD2-L-OPRT], T5.0005 in the pAC3 Plasmid RCR Vector).
[0236] A fusion construct was also developed by cloning a linker (Ser-Gly-Gly-Gly-Gly).sub.4) (SEQ ID NO:56) domain between and in frame with the CD polypeptide and the OPRT polypeptide to generated a CD-optimized-linker-OPRT sequence using Scheme IV as shown in FIG. 10D.
[0237] The resulting construct comprises a size of 1329 bp and the sequence immediately below:
TABLE-US-00015 (SEQ ID NO: 61) ##STR00013## AGGGCATGGATATCGCTTACGAGGAGGCCCTGCTGGGCTACAAGGAGGGC GGCGTGCCTATCGGCGGCTGTCTGATCAACAACAAGGACGGCAGTGTGCT GGGCAGGGGCCACAACATGAGGTTCCAGAAGGGCTCCGCCACCCTGCACG GCGAGATCTCCACCCTGGAGAACTGTGGCAGGCTGGAGGGCAAGGTGTAC AAGGACACCACCCTGTACACCACCCTGTCCCCTTGTGACATGTGTACCGG CGCTATCATCATGTACGGCATCCCTAGGTGTGTGATCGGCGAGAACGTGA ACTTCAAGTCCAAGGGCGAGAAGTACCTGCAAACCAGGGGCCACGAGGTG GTGGTTGTTGACGATGAGAGGTGTAAGAAGCTGATGAAGCAGTTCATCGA CGAGAGGCCTCAGGACTGGTTCGAGGATATCGGCGAGTCCGGCGGCGGCG CCTCCGGCGGCGGCGCCTCCGGCGGCGGCGCCTCCGGCGGCGGCGCCGCG GTCGCTCGTGcagctttggggccattggtgacgggtctgtacgacgtgca ggctttcaagtttggggacttcgtgctgaagagcgggctttcctccccca tctacatcgatctgcggggcatcgtgtctcgaccgcgtcttctgagtcag gttgcagatattttattccaaactgcccaaaatgcaggcatcagttttga caccgtgtgtggagtgccttatacagctttgccattggctacagttatct gttcaaccaatcaaattccaatgcttattagaaggaaagaaacaaaggat tatggaactaagcgtcttgtagaaggaactattaatccaggagaaacctg tttaatcattgaagatgttgtcaccagtggatctagtgttttggaaactg ttgaggttcttcagaaggagggcttgaaggtcactgatgccatagtgctg ttggacagagagcagggaggcaaggacaagttgcaggcgcacgggatccg cctccactcagtgtgtacattgtccaaaatgctggagattctcgagcagc agaaaaaagttgatgctgagacagttgggagagtgaagaggtttattcag gagaatgtctttgtggcagcgaatcataatggttctcccctttctataaa ggaagcacccaaagaactcaGCTTCGGTGCACGTGCAGAGCTGCCCAGGA ##STR00014## ATTTTATTTAGTCTCCAGAAAAAGGGGGG
[0238] FIG. 4 demonstrates that higher levels of the human codon optimized with the three mutations for higher stability are observed compared to wild type yCD protein in infected U-87 cells.
Example 3. Vector Production by Transient Transfection
[0239] Vector can be produced in a number of ways, but the first step is to introduce the DNA vector into cells to allow production of infectious particles, that can then be harvested from the cell supernatant. Once infectious particles have been generated other methods of production can be implemented by those skilled in the art. Vector particles were generated by transient transfection of 293T cells (Pear et al. Proc Natl Acad Sci USA. 90:8392-8396 1993).
[0240] The 293T cells were thawed and put into culture, then passaged twice in T-75 flasks containing 15 mL of the DMEM medium that was prepared by mixing DMEM High Glucose medium (Hyclone #30081, 500 mL) with FBS (Hyclone # SH30070, 50 mL), L-Glutamine (Cellgro #25-005-CI, 5 mL), NEAA (Hyclone # SH30238, 5 mL), and Penicillin-strep (Cellgro #30-002-CI, 5 mL). The flasks were incubated at 37.degree. C. and 5% CO.sub.2. After the 3.sup.rd passage cells were seeded in 6 T-25's, each containing 5 mL of the medium, at a cell density of 1.8.times.10.sup.6 cells/T-25 (or 7.2.times.10.sup.4 cells/cm.sup.2). One day after seeding the T-25's, the cells were transfected with the T5.0002 plasmid that expressed the viral vector using the Calcium Phosphate Transfection Kit from Promega (Cat # E1200). Eighteen hours following transfection, the media in one set of the flasks (3 flasks each set) were replaced with fresh medium containing 10 mM NaB. The media in the 2.sup.nd set of the flasks were not replaced, which served as a control (zero NaB). Eight hours post NaB treatment the media in all flasks were replaced with the fresh medium containing no NaB. The expression was allowed to continue for both sets of flasks until the next day (22 hours duration). The supernatants from both sets of flasks were harvested and assayed for their titers by qPCR expressed in Transducing Units (TU)/ml (see Example 4).
[0241] The titer results are shown in the following table.
TABLE-US-00016 Second titer (after storing Condition First titer at -80.degree. C. for 68 days) Without NaB 1.5 (.+-.0.05) .times. 10.sup.6 TU/mL 1.2 (.+-.0.2) .times. 10.sup.6 TU/mL 10 mM NaB 1.4 (.+-.0.3) .times. 10.sup.6 TU/mL 7.0 (.+-.0.14) .times. 10.sup.5 TU/mL TU = transduction unit
[0242] Subsequent vector preparations were produced in this manner, without sodium butyrate. Other vector plasmids (Table 2) have been used in the same way to generate vector preparations with titers between 1E5 TU/ml and 1E7 TU/ml. Such material can be further purified and concentrated, if desired, as described below see also: U.S. Pat. No. 5,792,643; T. Rodrigues et al. J Gene Med 9:233 2007.
[0243] In certain embodiments of the disclosure the dosing was calculated by grams of brain weight. In such embodiments, the dosing of a replication competent retroviral vector of the disclosure useful in the methods for treatment can range from 10.sup.4 to 10.sup.6 TU per gram brain weight.
Example 4. Quantitative PCR Tittering Assay
[0244] The functional vector concentration, or titer, is determined using a quantitative PCR-based (qPCR) method. In this method, vector is titered by infecting a transducible host cell line (e.g. PC-3 human prostatic carcinoma cells, ATCC Cat # CRL-1435) with a standard volume of vector and measuring the resulting amount of provirus present within the host cells after transduction. The cells and vector are incubated under standard culturing condition (37.degree. C., 5% CO.sub.2) for 24 hr to allow for complete infection prior to the addition of the anti-retroviral AZT to stop vector replication. Next, the cells are harvested from the culture dish and the genomic DNA (gDNA) is purified using an Invitrogen Purelink gDNA purification kit and eluted from the purification column with sterile RNase-/DNase-free water. The A.sub.260/A.sub.280 absorbance ratio is measured on a spectrophotometer to determine the concentration and relative purity of the sample. The gDNA concentrations are normalized with additional RNase-/DNase-free water to the lowest concentration of any given set of gDNA preparations such that the input DNA for the qPCR is constant for all samples analyzed. Genomic DNA purity is further assessed by electrophoresis of an aliquot of each sample on an ethidium bromide stained 0.8% agarose gel. If the sample passes an A.sub.260/A.sub.280 absorbance range of 1.8-2.0 and shows a single band of gDNA, then the sample is ready for qPCR analysis of provirus copy number of the vector. Using primers that interrogate the LTR region of the provirus (reverse-transcribed vector DNA and vector DNA that is integrated into the host gDNA), qPCR is performed to estimate the total number of transduction events that occurred when the known volume of vector was used to transduce the known number of cells. The number of transduction events per reaction is calculated from a standard curve that utilizes a target-carrying plasmid of known copy-number that is serial diluted from 10.sup.7 to 10 copies and measured under identical qPCR conditions as the samples. Knowing how many genomic equivalents were used for each qPCR reaction (from the concentration previously determined) and how many transduction events that occurred per reaction, we determine the total number of transduction events that occurred based on the total number of cells that were present at the time of transduction. This value is the titer of the vector after dilution into the medium containing the cells during the initial transduction. To calculate the corrected titer value, the dilution is corrected for by multiplying through by the volume of culture and the volume of titer divided by the volume of titer. These experiments are performed in replicate cultures and analyzed by qPCR using triplicate measurements for each condition to determine an average titer and with its associated standard deviation and coefficient of variance.
Example 5. Vector Testing
[0245] In order to be effective vector constructs and their derived infectious particles need to: (1) make good titer of virus by transient transfection (see Examples 3 and 4); (2) be stable upon multiple passages; (3) kill cells efficiently in the presence of 5-FC; and (4) express enzyme activity upon infection of target cells. Example 3 shows that useful titers can be obtained from the vectors.
[0246] Genetic Stability of Viral Vectors.
[0247] To demonstrate the stability the following experiment was performed. Approximately 10.sup.6 naive U-87 cells were initially infected with the viral vector at an MOI of 0.01, and grown until fully infected to complete a single cycle of infection. Supernatant is then repassed onto uninfected cells and the cycle repeated. In this experiment, twelve cycles have been completed in duplicate trials (FIG. 5 shows one of each of the duplicate trials; the other duplicates gave similar results). Genomic stability of the yCD2 or other transgene sequence is assessed by PCR amplification of the integrated provirus from the infected cells using MLV specific primers flanking the transgene insertion site. The appearance of any bands smaller than full-length amplicon would be an indicator of vector instability. FIG. 5 demonstrates that a vector of the disclosure (T5.0007--comprising the modified vector and CD heterologous polynucleotide) maintains stability for more passages than pACE-CD. Furthermore T5.0003 is somewhat less stable while T5.0004 and T5 appear about as stable as pACE-CD. pACE-CD has been used in mouse tumor studies and shows good anti-tumor effects in mouse models. However a more stable viral genome will be much more potent and long lasting in treatment of animals and humans, especially if multiple cycles of 5-FC treatment are required. Both T5.0001 and T5.0002 are markedly more stable than even T5.0007, showing that silent changes in a protein coding sequence or small changes that result in point mutations can lead to unexpectedly superior properties with more stable vector genomes.
[0248] Cell Killing Experiments.
[0249] The Cell Titer 96 Aqueous One Solution Cell Proliferation Assay (MTS) is a colorimetric method for determining the number of viable cells in proliferation assays. We have utilized this assay to determine cell growth kinetics, as well as to determine the dose response of various cell lines to 5-Fluorocytosine (5-FC) and 5-Fluorouracil (5-FU).
[0250] Cells 100% infected with vector were seeded at 1000 cells/well in 96-well plates. They were monitored over an eight day period following treatment with various concentrations of 5-FC (5-FU for controls). An analysis of their cell growth was assessed every two days utilizing Promega's Cell Titer 96 AQueous One Solution reagent (MTS). Briefly, 20 .mu.l of MTS was mixed with 100 .mu.l media (as recommended by the manufacturer) and added to the samples in the 96-well plate. The samples were incubated for 60 minutes in a 37.degree. C./5% CO.sub.2 incubator. Thereafter, absorbance readings were taken on a plate reader at a 490 nm wavelength. The plates were then discarded.
[0251] FIG. 6A shows the results of an experiment that demonstrates that the cytosine deaminase in cells expressing the yCD2 protein is at least as active as that from cells expressing the wild type yCD protein, by performing 5-FC titrations on U-87 cells infected either with AC3-yCD2 (vector) or AC3-yCD (vector). Briefly, U-87 cells 5 days post infection at a multiplicity of infection of 0.1 (i.e. 100% infected) with either AC3-yCD (wild type CD) vector or AC3-yCD2 (thermostabilized & codon optimized) vector were subject to increasing amounts of 5-FC or 0.1 mM of 5-FU as a positive control for 8 days. On day 8, cell cultures were assessed for viability using an MTS assay (Promega Cell Titer 96 AQUEOUS One Solution Proliferation Assay). Data shows comparable killing between the two retroviral vectors at increasing doses of 5-FC treatment.
[0252] In similar in-vitro cell culture experiments with RG2 cells (ATCC Cat # CRL-2433), the RG2 cell line was transduced with 5 different vectors (pACE-CD, T5.0001, T5.0002, T5.0004, and T5.0007). It was subsequently subject to increasing concentrations of 5-FC (5-FU for controls) for 8 days and monitored as described above. The results are shown in FIG. 2. Concentrations of 0.01 mM were sufficient to induce complete killing throughout all vectors tested with the exception of wild type-yeast Cytosine Deaminase (pACE-yCD). It was less sensitive and required 1.0 mM of 5-FC for complete killing.
[0253] CD Expression Assay.
[0254] U87 cells were transduced at a multiplicity of infection (MOI) of 0.1, cultivated for 5 days to allow viral spread and cells from day 5 post transduction were harvested. The cells were then collected by centrifugation at 800.times.g for 5 min. The supernatant was aspirated away from the cell pellet and washed with 5 mL of phosphate buffered saline (PBS) and again centrifuged at 800.times.g for 5 min. The resulting cell pellet was taken up in 1.5 mL of PBS, resuspended by passage through a pipette tip and placed in a freezer at -20 C. Cells were lysed by a freeze/thaw method. Previously resuspended cells were allowed to thaw at room temperature, passed through a pipette tip, mixed with protease inhibitor cocktail and again refrozen at -20 C. Previous to the enzyme assay, the sample was again thawed at room temperature and passed through a pipette tip. The suspension was then centrifuged at 14,000 rpm in a tabletop centrifuge for 5 min. The supernatant was decanted away from the pellet and placed in a fresh eppendorf tube and placed on ice.
[0255] yCD enzyme activity was assessed by using an HPLC assay. The HPLC assay was performed on a Shimadzu LC20AT unit connected in series with a photoarray detector and autoinjector. The solid phase was a Hypersil BDS C.sub.18 HPLC column with a 5 um sphere size and 4.0.times.250 mm column dimensions. The mobile phase was 50 mM ammonium phosphate, pH 2.1, containing 0.01% tert-butylammonium perchlorate and 5% methanol; the system was equilibrated at 22 C. All reagents were ACS grade and solvents were HPLC grade. A reaction mix was made consisting of 800 .mu.L with a final concentration of 0.125 mg/mL 5FC (1 mM) in PBS and placed in a 1.5 mL autosampler vial. The reaction was then initiated by adding 200 uL of each cell lysate. The reaction/autosampler vials were placed in the auto sampler and 5 uL of the reaction mixture was injected. Time points were taken periodically by retrieving a 5 uL aliquot from each reaction vial and analyzing on the HPLC column. The conversion rates of 5FC to 5FU were calculated by comparing the peak areas with known amounts from a previously generated standard curve of 5FU. The rate of 5FC conversion to 5FU was derived by plotting the amount of 5FU (in nmol) generated against its corresponding time interval. Protein concentration for the cell sample was derived and the Specific Activity of the cell lysate samples were calculated by dividing the conversion rate (nmol/min) by the amount of protein used in the assay in mg. FIG. 6B shows the specific activity of various vectors after 5 days on transduction at an MOI of 0.1. The data demonstrate that pACE-yCD (T5.0000)<pAC3-yCD1(T5.0001)<pAC3-CD2 (T5.0002) in terms of the specific activity of cytosine deaminase in tissue culture cells.
Example 6. Vector Purification and Concentration
[0256] A vector of the disclosure is manufactured by transient transfection on 293T cells (Example 3), followed by harvesting of the cell supernatant, filtration, benzonase treatment, diafiltration/concentration and dialysis. A further chromatography column step may be included, known to those skilled in the art (see for example U.S. Pat. No. 5,792,643; T. Rodriguez et al. J Gene Med 9:233 2007; WO 2010/148203). Clinical material is released based on standard testing such as sterility, mycoplasma and endotoxins, plus product specific potency, strength, and identity testing. Titer is determined as Transducing Units (TU) by PCR quantitation of integrated viral vector DNA in target cells (Example 4). The final product is targeted to have a titer of up to 10.sup.9 TU/ml formulated in isotonic Tris-buffered sucrose solution, as a sterile injectable.
[0257] In general, to accurately and precisely determine the strength of vector lots, a quantitative PCR-based titer assay has been developed (described in general terms in example 4). The details of the assay procedure consist of the following steps:
[0258] Transduction.
[0259] Transductions are performed in a 12-well plate format using the stable human prostate adenocarcinoma derived PC-3 cell line. For each test sample, three dilutions of un-titered vector preparation are used to transduce PC-3 cells in triplicate wells. Viral replication is stopped 24 hours post-transduction with azidothymidine (AZT). Cells are maintained for an additional 24-64 hours prior to harvesting and genomic DNA purification.
[0260] Genomic DNA Preparation.
[0261] Qiagen DNeasy DNA Minikits is used to prepare genomic DNA from transduced harvested cells as per the manufacturer's protocol. DNA concentrations and quality are assessed by direct absorbance measurement using UV/vis spectrophotometry to determine the A260 and A260/A280 ratio.
[0262] Real-time Quantitative PCR. The BioRad CFX96 real-time PCR instrument or equivalent is used for performing quantitative PCR. Provector copy numbers present in each test sample are measured by using specific DNA oligonucleotide primers in conjunction with a TaqMan probe designed to amplify the integrated, or pro-retroviral, U3/Psi packaging versus the CMV/Psi plasmid promoter. Vector titer is expressed relative to a copy number standard curve. To generate the vector copy number standard curve, genomic DNA from PC-3 cells is spiked with a unique plasmid containing the pro-retroviral U3/Psi sequence. Vector test sample titers are obtained by calculating the number of transduced genomes in multiple dilutions using multiple reactions per dilution.
[0263] For each titer assessment, a non-template control (wells containing all components except plasmid or genomic DNA) and a negative control (all components including equivalent genomic DNA from non-transduced PC-3 cells), is performed in triplicate. The titer values are expressed in transduction units per milliliter (TU/mL).
[0264] The potency of the vector of the disclosure is dependent on both the replication of the vector and the resultant cytosine deaminase (CD) activity in target cells. Therefore the potency assay measures the increase in CD activity over time as vector infection spreads in a previously uninfected cell line in tissue culture. The assay measures the enzymatic activity of the transferred yCD2 protein in transduced cells during early, middle and late stages of infection by monitoring the conversion of 5-fluorocytosine (5-FC) to 5-fluorouracil (5-FU), using reverse phase HPLC separation with UV detection. The increase of CD activity over the course of the infection is a function of the percent of cells infected over time and indicative of the TOCA 511 vector's ability to replicate. CD activity based on the 5-FC to 5-FU conversion rate is measured for each time point in CD units per mg of protein (the specific activity, SA). The increase in SA is then plotted over time, and reflects both the increase in the percentage of cells transduced as a result of viral replication in the culture, and the resultant transfer of CD activity. Accumulated data from multiple assays and vector lots has been used to determine an appropriate specification for this increase in SA of CD, for product release. The assay has 1, 3 and 5 day time points after an initial infection at an MOI of about 0.1 and a non-infected control.
[0265] CD activity from late stage infected cells (day 5 time point) was compared between lots to evaluate the use of this activity as an Identity test. The assay includes the following steps:
[0266] Transductions.
[0267] Transductions are performed in multi-well plate format on U87 cells. For each transduction, three suitable dilutions are used and each performed in triplicate. Cells are harvested at 0, 1, 3 and 5 days post transduction.
[0268] Set-Up of CD Reaction.
[0269] Cells are lysed and the total protein concentration determined using the BCA protein assay using BSA as a standard. For the yCD2 enzyme assay, an appropriate amount of cell lysate is added to buffer containing 5-FC such that the rate of 5-FU formation remains linear over 1-2 hours at 37.degree. C. The final volume for the reaction mixture is 100 .mu.L. After 2 h, the enzyme reaction is terminated by the addition of trichloroacetic acid, briefly vortexed and prepared for subsequent HPLC analyses. Cell lysates from non-transduced cells are used as a negative control while a similar assay using samples from 100% infected cells is used as a positive control.
[0270] Hplc Analysis.
[0271] The terminated reaction mixture is centrifuged at 12,000 rpm for 5 minutes at room temperature in a micro-centrifuge. The supernatant is then decanted away from the pellet and passed through a 0.2p filter to further remove particulates before injection onto a reverse phase HPLC column previously equilibrated with an aqueous based mobile phase containing phosphate buffer at a pH around 4.0. The chromatograms is followed at 260 nm and 280 nm to monitor both substrate consumption and product formation. Concentrations of either substrate or product are determined using the graphing and analysis capabilities of GraphPad by comparing them to previously generated standard curves calculated from known substrate or product concentrations. Amounts of 5-FU generated over 1-2 h are used to determine CD units of activity (1 unit of CD activity is defined as the formation 1 nmol of 5-FU per min) and the Specific Activity is calculated dividing this number by the amount of protein (from the cell lysate) used in the assay.
Example 7. Construction and Use of a Vector Encoding a Single Chain Antibody to CTLA-4 (CD 152)
[0272] Single chain antibodies are derived from known full antibody sequences that have a desired effect. Such sequences are available (e.g. WO2006048749, US2006165706, U.S. Pat. No. 7,034,121, Genbank Accession Numbers DJ437648, CS441506, CS441500, CS441494, CS441488, the disclosures of which are incorporated herein by reference). Such conventional antibody gene sequences are converted into single chain antibody (scFv) sequences by commonly used methods known to those skilled in the art (see for example Gilliland et al. "Rapid and reliable cloning of antibody variable regions and generation of recombinant single chain antibody fragments." Tissue Antigens 47, 1-20, 1996). Phage single chain antibodies to CTLA-4 are also available from screening phage-scFv libraries directly (Pistillo et al. Tissue Antigens 55:229 2000), and can be used directly for insertion into the replicating retroviral vectors of the disclosure. Regardless of how the sequence is derived scFv are typically about 700-900 bp in length and are synthesized by a commercial vendor (BioBasic) with a Psi1 site at the 5' end and compatible Not1 site at the 3' end, as described previously. This sequence is then inserted into the replicating retroviral back bone from pAC3-yCD2 at the Psi1-Not1 sites after removal of the yCD2 sequence. Vector is produced and titered as described, and further purified if necessary as described above. Human and Mouse CTLA4 are very homologous in sequence and the replicating retrovirus of the disclosure is first tested in a suitable syngeneic immunocompetent mouse models such as the CT26/Balb/c model and s91 mouse melanoma models, well known to those skilled in the art (see for example Hodge et al J. Immunol. 174:5994 2005). Outcome is measured by one or more of: modulation of tumor growth; lack of toxicity; generation of antitumor responses; shrinkage of remote lesions indicating systemic immunity. Doses are in the range of 10.sup.3 to 10.sup.7 TU in mice. In patients the vector is administered by intralesional injection into tumor, or by administration into the circulation that then carries the virus to the tumor. Doses are in the range of 10.sup.5 to 10.sup.11 TU.
Example 8. Anti-Melanoma Efficacy Studies with Anti CD152 Single Chain Antibody Expressing Vector in a Mouse Melanoma Model
[0273] Objective. The objective of this study is to assess the effect of a novel MLV based replication-competent retroviral vector carrying single chain antibody directed against Cytotoxic T-Lymphocyte Antigen 4 (CTLA-4) also referred to as Cluster of differentiation 152 (CD152) sequence (pAC3-.alpha.CD152) on melanoma growth, when delivered via intratumoral (IT) injection in DBA/2 mice bearing subcutaneous melanoma (Cloudman S91).
[0274] Mice.
[0275] Female DBA/2 or BALB/c mice (age .about.8 weeks) are purchased from Jackson Laboratories (Bar Harbor, Me.). Mice are acclimated for 7 days after arrival before start of studies.
[0276] Cells.
[0277] Cloudman S91 cells (ATCC, Manassas Va.) are a spontaneously arising melanoma derived from DBA/2 mice. Cells are cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells are resuspended in PBS (Hyclone, Logan Utah) for implantation. S91 cells (1E6 in 100 .mu.L) are injected into the right flank of DBA/2 mice.
[0278] Vector.
[0279] Vectors preparations are made by transient transfection (or from a producer cell line in HT1080 cells) with titers of approximately 7E6TU/ml. For initial studies vector is not further purified or concentrated. For follow on experiments to determine full dose response curves, high titer purified material is prepared with a titer expected around 10.sup.8/ml. Vector is administered IT in a volume of 50-100 .mu.L and IV in 100 .mu.L the total dose/mouse of approximately 7E5 TU/mouse. Vector expressing .alpha.CD152 is identified as Toca .alpha.CD152.
[0280] Tumor Implantation and Vector Injection.
[0281] Five groups of female DBA/2 (55 mice, 9-10 weeks of age) are implanted subcutaneously with S91 melanoma cells (Day 0) and then dosed (day 4-7 depending on growth rate of the S91 tumor; approximately 50-100 mm.sup.3) with vehicle (Groups 1), with control vector [AC3-GFP(V), (Group2), intratumor (IT) Toca .alpha.CD152 vector injection (Groups 3), or intravenous Toca .alpha.CD152 vector injection (group 4). Group 5 mice have no tumor implanted and are intravenously injected with Toca .alpha.CD152 only.
[0282] Data Analysis.
[0283] Tumor growth analysis is carried out to 2000 mm.sup.3 or to 60 days based on whichever comes first. 10 mice from each group will be plotted for tumor size over time. Statistical significance will be determined using analysis of variance (ANOVA). P values of <0.05 are considered statistically significant in all analyses, which are performed with Prism 5 statistical software (GraphPad Software) or equivalent. In-life observations are also taken to assess any adverse events to .alpha.CD152 expression during treatment.
[0284] Results.
[0285] Delivery of .alpha.CD152 by replicating MLV IT shows a statistically significant retardation of growth compared to the controls. Delivery of .alpha.CD152 by replicating MLV intravenously shows a statistically significant retardation of growth compared to the controls abrogates melanoma burden from the DBA/2-Cloudman S91 mouse melanoma model. Further animal studies were performed as described more fully below.
Example 9. ACE-yCD2 Viral Vector is Therapeutic in an Intracranial Human Xenograft (U87) in Nude Mice
[0286] An intracranial xenograft model using the U87 human glioma cell line was established to test RCR vector spread and biodistribution as well as therapeutic efficacy of RCR-vector mediated cytosine deaminase suicide gene therapy in a nude mouse host.
[0287] Following acclimation, mice were randomly assigned to one of 8 Treatment groups (see group description below). Seven groups underwent intracranial administration into the right striatum of 1.times.10.sup.5 U87 cells administered/mouse on Day 0. Group 8 mice were not implanted with tumor. At Day 5, mice were injected with Formulation Buffer only, or an RCR vector at 9.times.10.sup.5/5 ul, 9.times.10.sup.4/5 ul, or 9.times.10.sup.3/5 ul. Mice receiving no vector, or vector at 9.times.10.sup.5/5 ul or 9.times.10.sup.3/5 ul were randomized to receive 5-FC (500 mg/kg/day), administered as a single IP injection, beginning on Day 19, or no 5-FC. Mice receiving vector at mid dose all received 5-FC (i.e., No separate control group for this dose). 5-FC administration continued daily for 7 consecutive days followed by 15 days of no treatment. Cycles of drug plus rest were repeated up to 4 cycles. 10 mice from each group except group 8 were randomly assigned to the survival analysis category. The remaining mice were sacrificed according to a predetermined schedule.
[0288] Group Assignments and Dose Levels
TABLE-US-00017 N per Analysis Category (A) Survival (B) Scheduled Group Test article Volume Drug TX N analysis Sacrifice 1 Form buffer 5 ul none 4 4 before first drug cycle 2 Form buffer 5 ul 5FC 10 10 3 T5.0002 9e5/5 ul PBS 10 10 4 T5.0002 9e5/5 ul 5FC 25 10 3 before start of each cycle, 15 total 5 T5.0002 9e4/5 ul 5FC 10 10 6 T5.0002 9e3/5 ul 5FC 25 10 3 before start of each cycle, 15 total 7 T5.0002 9e3/5 ul PBS 10 10 8 NO none 5FC 15 3 before start TUMOR of each cycle, 15 total Total Number of Animals 109 60 49
[0289] Intravenous dosing was done via injection into the tail vein. Intraperitoneal dosing was done via injection into the abdomen with care taken to avoid the bladder. For intracranial injection mice were anesthetized with isoflurane and positioned in a stereotaxic device with blunt ear bars. The skin was shaved and betadine was used to treat the scalp to prepare the surgical site. The animal was placed on a heating pad and a scalpel used under sterile conditions to make a midline incision through the skin. Retraction of the skin and reflection of the fascia at the incision site will allow for visualization of the skull. A guide cannula with a 3 mm projection, fitted with a cap with a 3.5 mm projection, will be inserted through a small burr hole in the skull and attached with dental cement and three small screws to the skull. After hardening of the cement, the skin will be closed with sutures. The projected stereotaxic coordinates are AP=0.5-1.0 mm, ML=1.8-2.0 mm, DV=3.0 mm. Exact stereotaxic coordinates for the cohort of animals received will be determined in a pilot experiment (2-3 animals) by injecting dye and determining its location. The animals will be monitored during anesthesia recovery. Analgesics, buprenorphine, will be administered subcutaneously (SC) before the end of the procedure then buprenorphine will be administered approximately every 12 hrs for up to 3 days. Animals will be monitored on a daily basis. Cells or vector were intracranially infused through an injection cannula with a 3.5 mm projection inserted through the guide cannula. The rate was controlled with a syringe pump fitted with a Hamilton syringe and flexible tubing. For cell injection, 1 microliter of cells was delivered at a flow rate of 0.2 microliters per minute (5 minutes total).For vector injection, 5 microliters of vector was delivered at a flow rate Of 0.33 microliters per minute (15 minutes total).
[0290] Vector was delivered and calculated as transforming units (TU) per gram of brain weight to the mice. Using such calculation the translation of dose can be calculated for other mammals including humans. FIG. 8 shows the effect on vector dose in this mouse model.
Example 10. AC3-yCD2(V) is Therapeutic in a Syngeneic Mouse Model of Brain Cancer
[0291] Additional experiments to demonstrate the methods and compositions of the disclosure in a syngeneic animal model were performed.
[0292] An intracranial implant model using the CT26 colorectal cancer cell line in syngeneic BALB/c mice was established to test RCR vector spread and biodistribution as well as therapeutic efficacy of RCR-vector mediated cytosine deaminase suicide gene therapy and its immunological impact.
[0293] This study included 129 animals, 0 Male, 119 Female and 10 contingency animals (10 Female). Following acclimation, mice were randomly assigned to one of 8 Treatment groups (see group description below). Seven groups underwent intracranial administration into the right striatum of 1.times.10.sup.4 CT26 cells administered/mouse on Day 0. Group 8 mice were not implanted with tumor. At Day 4, mice were injected with Formulation Buffer only, or vector at 9.times.10.sup.5/5 ul, 9.times.10.sup.4/5 ul, or 9.times.10.sup.3/5 ul. Mice receiving no vector, or vector at 9.times.10.sup.5/5 ul or 9.times.10.sup.3/5 ul were randomized to receive 5-FC (500 mg/kg/BID), administered by IP injection, beginning on Day 13, or no 5-FC. Mice receiving vector at mid dose received 5-FC (i.e. No separate control group for this dose). 5-FC administration continued daily for 7 consecutive days followed by 10 days of no treatment. Cycles of drug plus rest were repeated up to 4 cycles. 10 mice from each group except group 8 were randomly assigned to the survival analysis category. The remaining mice were sacrificed according to a predetermined schedule.
[0294] Naive sentinel mice were co-housed with the scheduled sacrifice animals and taken down at the same time points to assess vector transmittal through shedding.
[0295] Group Assignments and Dose Levels
TABLE-US-00018 N per Analysis Category (A) Survival (B) Scheduled (C) Group Test article Volume Drug TX N analysis Sacrifice Sentinels 1 Form buffer 5 ul PBS 4 4 before first drug cycle 2 Form buffer 5 ul 5FC 10 10 3 T5.0002 9e5/5 ul PBS 10 10 4 T5.0002 9e5/5 ul 5FC 25 10 3 before start 1 before start of each cycle, of each cycle, 15 total 5 total 5 T5.0002 9e4/5 ul 5FC 10 10 6 T5.0002 9e3/5 ul 5FC 25 10 3 before start 1 before start of each cycle, of each cycle, 15 total 5 total 7 T5.0002 9e3/5 ul PBS 10 10 8 NO none 5FC 15 3 before start TUMOR of each cycle, 15 total Total Number of Animals 119 60 49 10
[0296] Intravenous dosing was done via injection into the tail vein. Intraperitoneal dosing was done via injection into the abdomen with care taken to avoid the bladder. For intracranial administration, mice with a guide cannula with a 3.2 mm projection implanted into the right striatum, and fitted with a cap with a 3.7 mm projection were used. The projected stereotaxic coordinates are AP=0.5-1.0 mm, ML=1.8-2.0 mm, DV=3.2 mm (from bregma). Cells or vector were intracranially infused through an injection cannula with a 3.7 mm projection inserted through the guide cannula. The rate was controlled with a syringe pump fitted with a Hamilton syringe and flexible tubing.
[0297] For cell injection, 1 microliter of cells was delivered at a flow rate of 0.2 microliter per minute (5 minutes total). For vector injection, 5 microliter of vector was delivered at a flow rate of 0.33 microliter per minute (15 minutes total).
[0298] Vector was delivered and calculated as transforming units (TU) per gram of brain weight to the mice. Using such calculation the translation of dose can be calculated for other mammals including humans. FIG. 9 shows the effect on vector dose in this mouse model when the vector is delivered intracranially.
Example 11. Construction and Testing of RCR Vectors Expressing miR-128-1 and miR128-2
[0299] Construction of Recombinant Replication Competent Retroviral Vector Containing a Heterologous Polynucleotide Sequence of Human pri-miRNA-128-1.
[0300] The replication competent retroviral vector, pAC3-miR-128-1 expressing miR-128-1 was derived from the backbone of pAC3-yCD2 described in one of the embodiment. The pAC3 backbone in the pAC3-miR-128-1 vector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Mlu I and Not I to remove the IRES-yCD2 polynucleotide sequence. The polynucleotide DNA sequence of pri-miR-128-1 was obtained from the product sheet of the pEP-mir-128-1 expression vector (Cell BioLabs Inc.) (SEQ ID NO: 31). DNA sequence of pri-miR-128-1 was synthesized with a Mlu I restriction enzyme site at the 5' end and a Not I restriction enzyme site at the 3'end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the Mlu I and Not I digested pAC3-yCD2 plasmid DNA described above. The resulting construct, pAC3-miR-128-1, encodes 3 genes: the gag, the pol, and the env, and the non-coding pri-miR-128-1 sequence (FIG. 11).
[0301] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement, the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0302] Construction of Recombinant Replication Competent Retroviral Vector Containing a Heterologous Polynucleotide Sequence of Human pri-miRNA-128-2.
[0303] The replication competent retroviral vector, pAC3-miR-128-2 expressing miR-128-2 was derived from the backbone of pAC3-yCD2 described in one of the embodiment. The pAC3 backbone in the pAC3-miR-128-1 vector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Mlu I and Not I to remove the IRES-yCD2 polynucleotide sequence. The polynucleotide DNA sequence of pri-miR-128-2 was obtained from the sequence analysis of the expression vector Lenti-miR-128-2 expressing the pri-miR128-2 (System Biosciences) (SEQ ID NO:32). DNA sequence of pri-miR128-2 was synthesized with a Mlu I restriction enzyme site at the 5' end and a Not I restriction enzyme site at the 3'end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the Mlu I and Not I digested pAC3-yCD2 plasmid DNA described above. The resulting construct, pAC3-miR-128-1, encodes 3 genes: the gag, the pol, and the env, and the non-coding pri-miR-128-2 sequence (FIG. 11).
[0304] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement, the supernatant containing the vector was collected and filtered through a 0.45 Dm filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0305] Construction of Recombinant Replication Competent Retroviral Vector Containing Heterologous Polynucleotide Sequences of Human H1 Promoter and Human Pre-miRNA-128.
[0306] The replication competent retroviral vector, pAC3-miR-128-2 expressing miR-128-2 was derived from the backbone of pAC3-yCD2 described in one of the embodiment. The pAC3 backbone in the pAC3-miR-128-1 vector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Mlu I and Not I to remove the IRES-yCD2 polynucleotide sequence. The polynucleotide DNA sequence of the human H1 promoter was obtained from the product information of pSilencer 3.1 H1 hygro expression vector (Ambion), and the polynucleotide DNA sequence of the short hairpin structured pre-miR-128-1 was obtained from the http: (//)www.mirbase.org/. DNA sequence of pre-miR128-1 linked to the human H1 promoter (SEQ ID NO: 33) was synthesized with a Mlu I restriction enzyme site at the 5' end and a Not I restriction enzyme site at the 3'end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the Mlu I and Not I digested pAC3-yCD2 plasmid DNA described above. The resulting construct, pAC3-H1-shRNAmiR128, encodes 3 genes: the gag, the pol, and the env, and the non-coding short hairpin structured pre-miR-128-1 sequence (FIG. 11).
[0307] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement, the supernatant containing the vector was collected and filtered through a 0.45 Dm filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0308] Analysis of Replication Kinetics of Recombinant Replication Competent Retroviral Vector by qPCR Assay.
[0309] Currently, there are at least two common ways to obtain replication kinetics of recombinant replication competent retroviral vector: (1) analysis of GFP expression with vectors expressing the GFP protein by flow cytometric analysis, and (2) reversed transcriptase assay by measuring the reverse transcriptase activity of the vector stock collected from cultured medium of transduced cells. Titers assessed by DNA analysis of transduced cells provide the most reliable estimate of the functional titers as compared to titers obtained by RNA and transgene expression in which titers were over-estimated and under-estimated, respectively, (Sastry et al., 2002 Gene Therapy 9, 1155-1162). The replication kinetics of viral spread correlates with the percent of a cell population being transduced in which an integrated proviral DNA of the viral genome can be detected by qPCR with primers specific to the viral sequence is being tested in culture. The present assay requires equal amount of genomic DNA input from all time points within the same vector tested and among various vectors if a comparison of replication kinetics is being tested. The replication kinetics graph was generated by plotting inversed C(t) values vs. time points. FIG. 12A and FIG. 12B show comparisons of replication kinetics of various recombinant replication competent retroviral vectors. The assay is sensitive to reveal the small difference in replication kinetics among various vectors.
[0310] Testing of Replication Kinetics of miR-128 Containing Recombinant Replication Competent Retroviral Vectors in Culture.
[0311] In order to confirm that the incorporation of pri-miR-128-1, pri-miR-128-2 and H1-pre-miR-128-1 sequence, respectively, replicates normally, calculated volume of each vector stocks collected from transient transfection mentioned above was used to infect fresh human fibrosarcoma cells, HT1080 and human glioma cells, U87-MG, respectively, at a MOI of 0.1. Transduced cells were passaged at day 3, 6 and 9 post infection. At each time point, a portion of cells were collected for genomic DNA extraction for qPCR. Dilutions of genomic DNA were made to generate aliquots of genomic DNA with same concentration for equal amount of genomic in-put in qPCR. Replication kinetics of each vectors were generated by plotting inversed C(t) values vs. time points. FIGS. 12A and 12B show that all vectors tested replicated at similar kinetics compared to control MLV virus.
[0312] Testing of Expression of Mature miR-128 from Cells Transduced with miR-128 Containing Recombinant Replication Competent Retroviral Vector.
[0313] In order to confirm the expression of miR-128 from cells transduced with miR-128 containing recombinant replication competent retroviral vectors, cells from day 9 post infection at which the maximal infectivity has reached (FIG. 12A and FIG. 12B) were expanded and harvested to extract total RNA for detection of mature miRNA expression. The results from Taqman microRNA assay showed an over expression of mature miR-128 from both HT1080 and U87-MG cells transduced with pAC3-miR-128-1, pAC3-miR-128-2, and pAC3-H1-shRNAmiR128 vectors, respectively, compared to untransduced cells (FIG. 13). In both cell lines, cells transduced with pAC3-miR-128-1 and pAC3-H1-shRNAmiR128 vector expressed higher level of mature miR-128 than cells transduced with pAC3-miR-128-2 vector.
[0314] Testing of Bmi-1 Expression from Cells Transduced with miR-128 Containing Recombinant Replication Competent Retroviral Vectors to Demonstrate Target Engagement of miR-12.
[0315] Bmi-1 expression has been observed to be up-regulated in a variety of human cancers including glioblastoma, and has been shown to be the target of miR-128. To confirm target engagement of miR-128, Bmi-1 expression from cells transduced with pAC3-miR-128-1, pAC3-miR-128-2 and pAC3-H1-shRNAmiR128 vector, respectively, was detected by qRT-PCR. FIG. 14 shows that U87-MG cells transduced with pAC3-miR-128-1, pAC3-miR-128-2 and pAC3-H1-shRNAmiR128 vector, respectively, expressed lower level of Bmi-1 than untransduced cells, whereas in HT1080 cells no significant difference was observed between transduced and untransduced cells. The data support the concept that miR-128 plays an important functional role in the central nervous system.
Example 12: Construction and Testing of Recombinant Replication Competent Retroviral Vector Containing Heterologous Polynucleotide Sequences of IRES, yCD2, Human H1 Promoter and Human Pre-miR128-1
[0316] Construction.
[0317] The replication competent retroviral vector, pAC3-yCD2-H1-shRNAmiR128 is derived from the backbone of pAC3-yCD2 described in one of the embodiments. The pAC3-yCD2 backbone in the pAC3-yCD2-H1-shRNAmiR128 vector is isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Not I. The polynucleotide DNA sequence of the human H1 promoter is obtained from the product information of pSilencer 3.1 H1 hygro expression vector (Ambion), and the polynucleotide DNA sequence of the short hairpin structured pre-miR-128-1 is obtained from the http: (//)www.mirbase.org/. DNA sequence of pre-miR128-1 linked to the human H1 promoter (SEQ ID NO: 34) is synthesized with a Not I restriction enzyme site at both ends of the double-stranded DNA fragment for subsequent insertion at the corresponding site in Not I digested pAC3-yCD2 plasmid DNA described above. The resulting construct, pAC3-H1-shRNAmiR128, encodes 4 genes: the gag, the pol, and the env, and the yCD2, and the non-coding short hairpin structured pre-miR-128-1 sequence (FIG. 11).
[0318] Vector stock is produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture is replaced with fresh medium. Twenty-four hours post medium replacement, the supernatant containing the vector is collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks is used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT is added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells is extracted for titer assay. The titer of the vector stocks is determined by qPCR with an inclusion of standards of known copy numbers.
[0319] Testing of Replication Kinetics of the pAC3-yCD2-H1-shRNAmiR128 Recombinant Replication Competent Retroviral Vectors in Culture.
[0320] In order to confirm that the incorporation of H1-pre-miR-128-1 replicates normally, calculated volume of each vector stocks collected from transient transfection mentioned above is used to infect fresh human fibrosarcoma cells, HT1080 and human glioma cells, U87-MG, respectively, at a MOI of 0.1. Transduced cells are passaged at day 3, 6 and 9 post infection. At each time point, a portion of cells are collected for genomic DNA extraction for qPCR. Dilutions of genomic DNA are made to generate aliquots of genomic DNA with same concentration for equal amount of genomic in-put in qPCR. Replication kinetics of each vectors are generated by plotting inversed C(t) values vs. time points. Result show that the vector replicates at similar kinetics compared to control MLV virus.
[0321] Testing of Expression of Mature miR-128 from Cells Transduced with the pAC3-yCD2-H1-shRNAmiR128 Recombinant Replication Competent Retroviral Vector.
[0322] To confirm the expression of miR-128 from cells transduced with pAC3-yCD2-H1-shRNAmiR128 recombinant replication competent retroviral vector, cells from day 9 post infection, at which the maximal infectivity is reached, are expanded and harvested to extract total RNA for detection of mature miRNA expression. Result from Taqman microRNA assay shows an over expression of mature miR-128 from both HT1080 and U87-MG cells transduced with the pAC3-yCD2-H1-shRNAmiR128 compared to untransduced cells.
[0323] Testing of Bmi-1 Expression from Cells Transduced with pAC3-yCD2-H1-shRNAmiR128 Recombinant Replication Competent Retroviral Vectors to Demonstrate Target Engagement of miR-128.
[0324] Bmi-1 expression has been observed to be up-regulated in a variety of human cancers including glioblastoma, and has been shown to be the target of miR-128. To confirm target engagement of miR-128, Bmi-1 expression from cells transduced with pAC3-yCD2-H1-shRNAmiR128 is detected by qRT-PCR. The result shows that U87-MG cells transduced with pAC3-yCD2-H1-shRNAmiR128 express lower level of Bmi-1 than untransduced cells, whereas in HT1080 cells no significant difference was observed between transduced and untransduced cells. The data support the concept that miR-128 plays an important functional role in the central nervous system.
[0325] Testing of yCD2 Expression from Cells Transduced with pAC3-yCD2-H1-shRNAmiR128 by Immune-Blot.
[0326] To confirm the expression of yCD2 from cells transduced with pAC3-yCD2-H1-shRNAmiR128 recombinant replication competent retroviral vector, cells from day 9 post infection, at which the maximal infectivity is reached, are expanded and harvested to extract total protein for detection of yCD2 expression. The result from immune-blot shows normal expression yCD2 from both HT1080 and U87-MG cells transduced with the pAC3-yCD2-H1-shRNAmiR128 compared to pAC3-yCD2 transduced cells.
Example 13: Testing of Vector Stability of miR-128 Containing Recombinant Retroviral Vectors in Culture
[0327] Multiple serial infection cycles of miR-128 containing recombinant retroviral vectors (pAC3-miR-128-1, pAC3-miR-128-2, pAC3-H1-shRNAmiR128 and pAC3-yCD2-H1-shRNAmiR128) is tested to assess the stability of the vectors. HT1080 and U87-MG cells are initially infected with vectors at a low MOI and are allowed to spread in culture. Vector stocks at each infection cycles are collected, filtered diluted to infect fresh cells. At the end of each infection cycles, cells are harvested and genomic DNA are extracted for assessment of transgene stability by standard PCR using primers that bind to the 3' of the env gene and 3' of the untranslated region in the vector upstream of the 3'LTR. The result shows that all vectors tested remain stable for at least over 8-20 cycles.
Example 14: Anti-Tumor Efficacy Studies with miRNA Expressing Vector in a Mouse/Human Xenograft Model
[0328] Objective.
[0329] The objective of this study is to assess the effect of a novel MLV based replication-competent retroviral vectors carrying the miR128 sequence (AC3-miR128-1(V); AC3-miR128-2(V); AC3-miR128-3(V) on survival, when delivered via intracranial (IC) injection in nude mice bearing a human glioma xenograft, at three Toca 511 dose levels.
[0330] Mice.
[0331] Female athymic nude-Foxn1{circumflex over ( )}nu (nude) mice (age .about.8 weeks) are purchased from Harlan (Indianapolis Ind.). Mice are acclimated for 7 days after arrival. Mice undergo surgical placement of an indwelling guide cannula with a 3.0 mm projection implanted into the right striatum, and fitted with a cap containing a 3.5 mm projection. The stereotaxic coordinates are AP=+0.5 mm, ML=-1.8 mm (from bregma).
[0332] Cells.
[0333] U-87 MG cells (ATCC, Manassas Va.) are derived from a malignant glioma from a 44 year old Caucasian female. Cells are cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells are resuspended in PBS (Hyclone, Logan Utah) for implantation. U-87 MG cells (1E5 in 1 .mu.L) are infused at 0.2 .mu.L per minute (5 minutes, followed by a hold of 5 minutes) IC through an injection cannula with a 3.5 mm projection inserted through the guide cannula.
[0334] Vectors preparations are made by transient transfection (or from a producer cell line) and all have titers of approximately 5E6TU/ml. For initial studies vector is not further purified or concentrated. For follow on experiments to determine full dose response curves, high titer purified material is prepared with a titer of around 10E8/ml. Vector is administered IC in a volume of 5 ul or less for a minimum total dose/mouse of approximately 2.5E4 TU/mouse.
[0335] Tumor Implantation and Vector Injection.
[0336] Six groups of female athymic nude-Foxn1{circumflex over ( )}nu mice (65 mice, 9-10 weeks of age) are implanted IC with U-87 tumor cells (Day 0) then dosed IC (day 4-7 depending on growth rate of the U87 cells) with vehicle (Groups 1), with control vector (AC3-GFP(V), Group2) or IC with AC3-miR128-1(V); AC3-miR128-2(V); AC3-miR128-3(V) (Groups 3-5). Group 6 mice were not implanted with tumor or vector.
[0337] Data Analysis.
[0338] Survival analysis to day 60 is performed on 10 mice each from Groups 1-6 and plotted as a Kaplan Meyer plot. Survival curves are compared by the log-rank test. P values of <0.05 are considered statistically significant in all analyses, which are performed with Prism 5 statistical software (GraphPad Software) or equivalent.
[0339] Results from treatment with the vectors show a statistically significant survival advantage in this human glioma xenograft model compared to treatment with control vector or vehicle alone.
Example 15: Anti-Tumor Efficacy Studies with yCD2 and miRNA Expressing Vector in a Mouse/Human Xenograft Model
[0340] The objective of this study is to assess the effect of a novel MLV based replication-competent retroviral vector expressing yCD2 and miR-128, designated AC3-yCD2-H1-shRNAmiR128 (V) on survival when delivered via intracranial (IC) injection in nude mice bearing a "stem-cell" like enriched human glioma xenograft, at three dose levels.
[0341] Female athymic nude-Foxn1{circumflex over ( )}nu (nude) mice (age .about.8 weeks) are purchased from Harlan (Indianapolis Ind.). Mice are acclimated for 7 days after arrival. Mice undergo surgical placement of an indwelling guide cannula with a 3.0 mm projection implanted into the right striatum, and fitted with a cap containing a 3.5 mm projection. The stereotaxic coordinates are AP=+0.5 mm, ML=-1.8 mm (from bregma).
[0342] Early passages of primary human glioma (Dr. Carol Kruse, Burnham Biomedical Res Inst, San Diego, Calif.) are cultured in serum-free medium with EGF, bFGF and B27 supplement. Cells are resuspended in PBS (Hyclone, Logan Utah) for implantation. Approximately 1000 cells in 1 .mu.L are infused at 0.2 .mu.L per minute (5 minutes, followed by a hold of 5 minutes) IC through an injection cannula with a 3.5 mm projection inserted through the guide cannula.
[0343] Vectors preparations are made by transient transfection (or from a producer cell line) and all have titers of approximately 5E6TU/ml. For initial studies vector is not further purified or concentrated. For follow on experiments to determine full dose response curves, high titer purified material is prepared with a titer of around 10E8/ml. Vector is administered IC in a volume of 5 ul or less for a minimum total dose/mouse of approximately 2.5E4 TU/mouse.
[0344] Tumor Implantation and Vector Injection.
[0345] Six groups of female athymic nude-Foxn1{circumflex over ( )}nu mice (66 mice, 9-10 weeks of age) are implanted IC with "stem-like" cells (Day 0) then dosed IC at day 7-14 post tumor implantation depending on growth rate of the cells with Groups 1: vehicle; Group2: control vector AC3-GFP(V); Group 3: TOCA511; Group 4: AC3-yH1-shRNAmiR128(V); Group 5: AC3-yCD2-H1-shRNAmiR128(V); and Group 6: untreated mice that are not implanted with tumor or vector.
[0346] Data Analyses.
[0347] Survival analysis to day 60 is performed on 10 mice each from Groups 1-5 and plotted as a Kaplan Meyer plot. Survival curves are compared by the log-rank test. P values of <0.05 are considered statistically significant in all analyses, which are performed with Prism 5 statistical software (GraphPad Software) or equivalent.
[0348] Results.
[0349] Results from treatment with the vectors show a statistically significant survival advantage in this human glioma xenograft model compared to treatment with control vector or vehicle alone.
Example 16: Construction of Vectors with miR Target Sequences
[0350] Construction of Replication Competent Retroviral Vector Expressing GFP and Containing a Single Copy of 142-3p Target Sequence.
[0351] The replication competent retroviral vector, pAC3-emd-142-3pT, encoding a GFP was derived from the backbone of pAC3-emd described above (current existing patent). The pAC3-emd backbone in the pAC3-emd-142-3pT vector was isolated by endonuclease digestion of the pAC3-emd plasmid DNA with Not I. The perfect complementary target sequence of miR142-3pT was obtained from published literature (Brown et al., 2006 Nature Medicine 12:5 585-591). The target sequence of the miR-142-3p (SEQ ID NO: 35) was synthesized with a Not I restriction enzyme site present at each end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the pAC3-emd plasmid DNA. The orientation of the 142-3pT insert was confirmed by sequencing analysis. The resulting construct, pAC3-emd-142-3pT, encodes 4 genes: the gag, the pol, the env, and the emd, and the non-coding 142-3pT sequence (FIG. 15).
[0352] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0353] Construction of Replication Competent Retroviral Vector Expressing yCD2 and Containing a Single Copy of 142-3p Target Sequence.
[0354] The replication competent retroviral vector, pAC3-yCD2-142-3pT, encoding a yCD2 gene was derived from the backbone of pAC3-yCD2 described above (current existing patent). The pAC3-yCD2 backbone in the pAC3-yCD2-142-3pT vector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Not I. The perfect complementary target sequence of miR142-3pT was obtained from published literature (Brown et al., 2006 Nature Medicine 12:5 585-591). The target sequence of the miR-142-3p (SEQ ID NO: 35) was synthesized with Not I restriction enzyme site present at each end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the pAC3-yCD2 plasmid DNA. The resulting construct, pAC3-yCD2-142-3pT, encodes 4 genes: the gag, the pol, the env, and the yCD2, and the non-coding 142-3pT sequence (FIG. 16).
[0355] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0356] Construction of Replication Competent Retroviral Vector Expressing GFP and Containing 4 Copies of 142-3p Target Sequence.
[0357] The replication competent retroviral vector, pAC3-emd-142-3pT4X, encoding yCD2 (modified cytosine deaminase) was derived from the backbone of pAC3-emd described above. The pAC3-yCD2 backbone in the pAC3-emd-142-3pT 4.times. vector was isolated by endonuclease digestion of the pAC3-emd plasmid DNA with Not I. Four tandem repeat of the perfect complementary target sequence of miR142-3pT4X was obtained from published literature (Brown et al., 2006 Nature Medicine 12:5 585-591). The target sequence of the miR-142-3p4X (SEQ ID NO: 36) was synthesized with a Not I restriction enzyme site present at each end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the pAC3-emd plasmid DNA. The orientation of the 142-3pT insert was confirmed by sequencing analysis. The resulting construct, pAC3-emd-142-3pT4X, encodes 4 genes: the gag, the pol, the env, and the emd, and the non-coding 142-3pT4X sequence (FIG. 15).
[0358] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0359] Construction of Replication Competent Retroviral Vector Expressing yCD2 and Containing 4 Copies of 142-3p Target Sequence.
[0360] The replication competent retroviral vector, pAC3-yCD2-142-3pT4X, encoding a GFP was derived from the backbone of pAC3-emd described above. The pAC3-emd backbone in the pAC3-yCD2-142-3pT 4Xvector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Not I. Four tandem repeat of the perfect complementary target sequence of miR142-3pT4X was obtained from published literature (Brown et al., 2006 Nature Medicine 12:5 585-591). The target sequence of the miR-142-3pT4X (SEQ ID NO: 36) was synthesized with a Not I restriction enzyme site present at each end of the double-stranded DNA fragment for subsequent insertion at the corresponding site in the pAC3-yCD2 plasmid DNA. The orientation of the 142-3pT4X insert was confirmed by sequencing analysis. The resulting construct, pAC3-emd-142-3pT4X, encodes 4 genes: the gag, the pol, the env, and the emd, and the non-coding 142-3pT4X sequence (FIG. 16).
[0361] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
Example 17: Testing of Replication Kinetics
[0362] 142-3pT Containing Recombinant Retroviral Vectors in Non-Hematopoietic Human Cell Lines.
[0363] In order to confirm that the incorporation of the 142-3pT sequence in a vector of the disclosure replicates similar to their parental vectors, calculated volume of each vector stocks collected from transient transfection mentioned above was used to infect fresh human fibrosarcoma cells, HT1080 and human glioma cells, U87-MG, respectively, at a MOI of 0.1. Transduced cells were passaged at day 3, 6 and 9 post infection. At each time point, a portion of cells were collected for genomic DNA extraction for qPCR. Dilutions of genomic DNA were made to generate aliquots of genomic DNA with same concentration for equal amount of genomic in-put in qPCR.
[0364] Replication kinetics of each vectors were generated by plotting inversed C(t) values vs. time points. FIGS. 17A and 18A show that all vectors tested replicated at similar kinetics compared to their parental vectors (pAC3-emd and pAC3-yCD2). Replication kinetics of vectors expressing GFP protein (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) was also assessed by flow cytometric analysis. FIGS. 17 and 18 show that pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vectors replicated at a similar kinetic as their parental vector, pAC3-emd in these cell lines.
[0365] Testing of Replication Kinetics of 142-3pT Containing Recombinant Retroviral Vectors in Human and Mouse Hematopoietic Cells.
[0366] The expression of mature miR-142-3p was first confirmed in a mouse T-lymphocytic cell line EL4, a human T-lymphocytic cell line SUP-T1 and a human monocytic cell line U937 by Taqman microRNA assay using the primer set specific for mouse and human miR-142-3p as the sequences of mature miR-142-3p of the two species are identical. The replication kinetics of recombinant retroviral vector expressing the GFP (pAC3-emd) was tested in all three cell lines with an initial infection at a MOI of 2. FIG. 19 shows that the pAC3-emd vector replicated efficiently in human T-lymphocytes and monocytes (SUP-T1 and U937) as the viral spread reached 65% and 95% of cell population, respectively, by day 28 post infection. Vector spread in EL4 cells remained less than 5% during the time frame tested in FIG. 19, but eventually spread to 40% of cells by day 60 and 70% by day 75 was observed (FIG. 20D).
Example 18: Testing of Vector Spread of 142-3pT Containing Recombinant Retroviral Vectors in Human and Mouse Hematopoietic Cells
[0367] The functional effect of miR-142-3p in suppressing the GFP expression in the recombinant retroviral vector expressing GFP was tested by flow cytometric analysis. Calculated volume of pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vector stock collected from transient transfection mentioned above was used to infect fresh EL4, SUP-T1 and U937 cells at a MOI of 2. A portion of cells were collected at day 6, 12 and 18 post infection for analysis of GFP expression by flow cytometric analysis. FIG. 20A shows the GFP expression was suppressed to background level in EL4 cells transduced with pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vector compared to their parental vector pAC3-emd. The GFP suppression remained persistent within the time frame tested. FIG. 20B and FIG. 20C show remarkable suppression of GFP expression in SUP-T1 and U937 cells transduced with pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vector, respectively, compared to their parental vector pAC3-emd. FIG. 20D shows that even when time is allowed for spread of the pAC3-emd vector to approximately 30% at 55 days, expression of pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vector continues to be suppressed. The results confirm that the miR-142-3p expression in mouse and human hematopoietic cells effectively suppress the GFP expression in cells transduced with recombinant retroviral vectors containing the 142-3pT sequence. In U937 cells, the result suggested that the vector containing 4 copies of 142-3pT (pAC3-emd-142-3pT4X) may be more effective in suppressing GFP expression than the vector containing single copy of 142-3pT (pAC3-emd-142-3pT4X).
Example 19: Testing of Viral RNA Genome
[0368] It is unclear whether the functional effect of miR-142-3pT in suppressing the GFP expression mentioned above is due direct suppression of GFP expression at the translational level or due to degradation of viral genome at post transcriptional level. A portion of cells at the end of the experimental time point is collected for total RNA extraction using standard molecular biology method. qRT-PCR using primers (e.g. pol primer set and env2 primer set) specific to cDNA derived from reverse transcribed viral RNA is performed to assess viral load of transduced cells. The result shows that the viral load of cells transduced with pAC3-emd-142-3pT and pAC3-emd-142-3pT4X, respectively, is significantly lower than cells transduced with pAC3-emd vector. The result supports the concept that miR-142-3p in mouse and human hematopoietic cells effectively degrades the viral RNA genome at post transcriptional level, thereby, restricts the vector spread in mouse and human hematopoietic cells.
Example 20: Testing of Integrated Proviral DNA by qPCR
[0369] It is unclear whether the functional effect of miR-142-3pT in suppressing the GFP expression mentioned above is due direct suppression of GFP expression at the translational level or due to degradation of viral genome and thus integration of proviral DNA. A portion of cells at the end of the experimental time point is collected for genomic DNA extraction using standard molecular biology methods. qPCR using primers specific to integrated proviral DNA is performed to assess the copy number of integrated proviral DNA per cell. The result shows that the copy number per cells of cells transduced with pAC3-emd-142-3pT and pAC3-emd-142-3pT4X, respectively, is significantly lower than cells transduced with pAC3-emd vector. The result supports the concept that miR-142-3p in mouse and human hematopoietic cells effectively degrades the viral RNA genome at post transcriptional level, and thereby restricts the vector spread in mouse and human hematopoietic cells.
Example 21: Testing of Vector Stability of 142-3pT Containing Recombinant Retroviral Vectors in Culture
[0370] Multiple serial infection cycles of 142-3pT containing recombinant retroviral vectors is tested to assess the stability of the vectors. HT1080 and U87-MG cells are initially infected with vectors at a low MOI and are allowed to spread in culture. Vector stocks at each infection cycles are collected, filtered and diluted to infect fresh cells. At the end of each infection cycles, cells are harvested and genomic DNA are extracted for assessment of transgene stability by standard PCR using primers that bind to the 3' of the env gene and 3' of the untranslated region in the vector downstream of the heterologous polynucleotide sequence linked to the IRES. The result shows that vectors containing single copy of 142-3pT (pAC3-emd-142-3pT and pAC3-yCD2-142-3pT) remains stable for at least over 10-20 cycles, whereas vectors containing 4 tandem repeats of 142-3pT (pAC3-emd-142-3pT4X and pAC3-yCD2-142-3pT4X) can show deletion of 142-3pT sequence in early infection cycles.
Example 22: Testing of Controlled Vector Spread of 142-3pT Containing Recombinant Retroviral Vectors in In Vivo
[0371] This experiment is conducted with the same design as example 27 below. The functional effect of miR-142-3p in restricting vector spread via hematopoietic cells is tested in vivo by intravenous injection of the recombinant retroviral vectors (pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X) in 8-wk-old nude Balb/C mice with implanted U87 xenografts. For each vector, there are three groups of mice each represents a time point (e.g. 30 day, 60 day and 90 day post viral vector administration). A dose of 1E4 to 1E7 TU of each vector stock is administered by intravenous injection to all animals. Animals from each time point are sacrificed to harvest spleen, lymph nodes and bone marrow and tumor. GFP expression of subpopulation of cells (e.g. CD4+, CD8+ and etc.) are harvested and analyzed by flow cytometric analysis. In a duplicate experiment, animals from each time point are sacrificed and tissues (e.g. liver, kidney, spleen, tumor etc.) are collected for genomic DNA extraction. qPCR is performed to assess the presence of integrated proviral DNA in tissues collected. The result shows vector spread of vectors containing the 142-3pT is significantly reduced in hematopoietic tissues demonstrating the reduction of vector replication in these tissues. At the same time GFP and PCR signal is still observed in the tumor, showing that the miR target sequences have depressed spread in the lymphoid tissues, but still allowed spread in the tumor tissue.
Example 23. Extended Survival in a Patient Dog with Spontaneous Recurrent Malignant Glioma and Treated with T5.0002 Vector Plus 5-FC
[0372] A male 35 kg Boxer dog, presenting with recurrent anaplastic oligodendroglioma 3 months following complete surgical resection, was treated with T5.0002 virus, purified and formulated (see U.S. Pat. No. 5,792,643; T. Rodriugez et al. J Gene Med 9:233 2007; WO 2010/148203) in isotonic Tris/NaCl pH7.2 rendered isotonic with mannitol & sucrose, 1 mg/ml HSA, 0.1 mg/ml ascorbate) in combination with 5-FC. The tumor measured approximately 13 cm.sup.3, and caused major lateral ventricle compression and significant midline shift (See FIG. 1) Due to the large size of the tumor, Toca 511 was infused through 2 separate catheters (400 .mu.L and 480 .mu.L), using Convection Enhanced Delivery (CED). The total Toca 511 dose administered was approximately 4.1.times.10.sup.6 TU/g brain. ProHance.RTM. (gadoteridol) was added to Toca 511 prior to injection to allow visualization of delivery by MRI. The volume of distribution of the vector was estimated to be approximately 10-12% of the tumor volume.
[0373] FIGS. 21 A and 21B are still frames from the MRI images obtained from the patient dog during intratumoral CED infusion of Toca 511 and gadolinium. Note the large tumor on the left side of the image compressing both sides of the brain and shifting midline structures to the right. The white areas are the gadolinium-Toca 511 infusion. FIG. 21B shows the placement of the two catheters into the tumor.
[0374] Toca 511 was allowed to spread for 8 days. The dog was treated with 5-FC at a divided dose of 130 mg/kg/day, by mouth, three times daily with food for 5 days. The dose was increased to 160 mg/kg/day for 2 more days (7 days of 5-FC total). A follow-up MRI showed no change in tumor size and some possible changes to the internal area of the tumor. After 21 days of viral spread, a second cycle of 5-FC was initiated at the higher dose of 160 mg/kg/day (divided, three times a day with food). The drug was stopped after the fifth day of dosing due to the development of rash.
[0375] MRI performed at 2 weeks after the first course of 5-FC and 2 weeks after the second course of 5-FC (7 weeks after treatment began) has shown that the tumor volume has plateaued while the rate of tumor growth has declined. The patient became more alert and active, and remained clinically stable, 13 weeks after injection of vector. At 15.5 weeks the dog was euthanized because of stomach bleeding due to prolonged high dose steroid administration (and not because of the tumor). The estimated lifespan of the dog was no more than 3-4 weeks at the time of initial injection of the vector. Efficacy of the Toca 511/5-FC combination in this patient dog is shown by survival 3 to 4 times longer than that originally estimated by his attending veterinarian. At autopsy low levels (20-30 copies/microgram genomic DNA) of vector DNA were detectable in the residual tumor, but nowhere else in the dog.
Example 24 Construction of Gamma Interferon Vectors
[0376] Construction and Testing of a Replication Competent Retroviral Vector Encoding the Human IFN-Gamma Gene.
[0377] The replication competent retroviral vector, pAC3-hIFNg, encoding the human IFN-gamma gene, was derived from the backbone of pAC3-yCD2 described above. The pAC3 backbone in the pAC3-hIFNg vector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Psi I and Not I. The cDNA sequence of human IFN-gamma gene was identified and confirmed among three sequences obtained from different accession numbers (AM903379, BC070256, and NM000619). Sequence alignment showed identical sequence among the three. The open reading frame of the human IFN-gamma (SEQ ID NO: 38) was synthesized with Psi I and Not I restriction enzyme site present at each end of the DNA fragment for subsequent insertion at the corresponding site in the pAC3 backbone. The resulting construct, pAC3-hIFNg, encodes 4 genes: the gag, the pol, the env, and the human IFN-gamma (FIG. 22).
[0378] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into HT1080 cells using FUGENE HD transfection Reagent. Forty-eight hours post transfection the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Specific volume of the undiluted vector stock was used to infect fresh 75% confluent HT1080. At day 4 and day 5 post infection, the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0379] The expression of human IFN-gamma was first tested at the RNA level. Total RNA was extracted from transduced HT1080 cells at 5 days post infection in the second infection using standard RNA extraction method. RT-PCR was performed to detect the expression of human IFN-.gamma.. Fifty nano-gram of total RNA was used in the RT reaction to generate cDNA. One tenth of the volume from RT reaction was subsequently used for PCR using PCR primer set specific for human IFN-.gamma.. Result from RT-PCR showed that human IFN-gamma is expressed in HT1080 cells infected with pAC3-hIFNg vector.
[0380] The expression of secreted human IFN-D protein was tested by standard ELISA. Vector stock collected from day 4 and day 5 post infection was serially diluted in the ELISA assay in order to obtain a linear range between protein concentration and dilution factor. The result showed that human IFN-.gamma. protein is secreted at a higher concentration by the HT1080 cells at day 5 post infection than by the cells at day 4 post infection (FIG. 24). Cells at post d5 infection secreted approximately 325-355 pg/mL human IFN-.gamma. protein.
[0381] Construction and Testing of a Replication Competent Retroviral Vector Encoding the Mouse IFN-Gamma Gene.
[0382] The replication competent retroviral vector, pAC3-mIFNg, encoding the mouse IFN-.gamma. gene was derived from the backbone of pAC3-yCD2 described above. The pAC3 backbone in the pAC3-mIFNg vector was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Psi I and Not I. The cDNA sequence of mouse IFN-.gamma. gene was identified and confirmed among three sequences obtained from different accession numbers (BC119063, BC119065 and NM008337). Sequence alignment showed identical sequence among the three. The open reading frame of the mouse IFN-gamma (SEQ ID NO: 39) was synthesized with Psi I and Not I restriction enzyme site present at each end of the DNA fragment for insertion at the corresponding site in the pAC3 backbone. The resulting construct, pAC3-mIFNg, encodes 4 genes: the gag, the pol, the env, and the mouse IFN-gamma (FIG. 22).
[0383] Vector stock was produced by transient transfection of the vector-encoding plasmid DNA into HT1080 cells using FUGENE HD transfection Reagent. Forty-eight hours post transfection the supernatant containing the vector was collected and filtered through a 0.45 micron filter and used immediately or stored in aliquots at -80.degree. C. for later use. Specific volume of the undiluted vector stock was used to infect fresh 75% confluent HT1080. At day 4 and day 5 post infection the supernatant containing the vector was collected and filtered through a 0.45 micron filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0384] The expression of mouse IFN-gamma was first tested at the RNA level. Total RNA was extracted from transduced HT1080 cells at 5 days post infection in the second infection using standard RNA extraction method. RT-PCR was performed to detect the expression of mouse IFN-gamma. Fifty nano-gram of total RNA was used in the RT reaction to generate cDNA. One tenth of the volume from RT reaction was subsequently used for PCR using PCR primer set specific for mouse IFN-gamma (FIG. 23). Result from RT-PCR showed that mouse IFN-gamma is expressed in HT1080 cells infected with pAC3-mIFNg vector.
[0385] The expression of secreted human IFN-gamma protein was tested by standard ELISA. Vector stock collected from day 4 and day 5 post infection was serially diluted in the ELISA assay in order to obtain a linear range between protein concentration and dilution factor. The result showed that mouse IFN-.gamma. protein is secreted at a higher concentration by the HT1080 cells at day 5 post infection than by the cells at day 4 post infection (FIG. 25). Cells at post d5 infection secreted approximately 33-42 ng/mL mouse IFN-gamma protein.
Example 25: Anti-Tumor Efficacy Studies with Gamma Interferon Expressing Vector in a Mouse Subcutaneous Tumor Model
[0386] Objective.
[0387] The objective of this study is to assess the effect of a novel MLV based replication-competent retroviral vector carrying the murine gamma interferon sequence (pAC3-mIFNg) on tumor growth, when delivered via intratumoral (IT) injection in BALB/c mice bearing subcutaneous colon carcinoma (CT26.WT).
[0388] Mice.
[0389] Female BALB/c mice (age .about.8 weeks) are purchased from Jackson Laboratories (Bar Harbor, Me.). Mice will be acclimated for 7 days after arrival before start of studies.
[0390] Cells.
[0391] CT26.WT cells (ATCC, Manassas Va.) are an N-nitroso-N-methylurethane-(NNMU) induced, undifferentiated colon carcinoma cell line. Cells are cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells are resuspended in PBS (Hyclone, Logan Utah) for implantation. CT26.WT cells (2E5 in 100 .mu.L) are injected into the right flank of BALB/c mice.
[0392] Vectors.
[0393] Vectors preparations are made by transient transfection (or from a producer cell line after infection of a second cell line with the infectious virus from the initial transfection; see, e.g., International Application No. PCT/US10/38996, the disclosure of which is incorporated herein by reference) with titers of approximately 3.times.10.sup.6 TU/ml. For initial studies vector is not further purified or concentrated. For follow on experiments to determine full dose response curves, high titer purified material is prepared with a titer expected around 10.sup.8/ml. To achieve high titer material, canine cell line CF2 are chosen for production as gamma interferon is poorly cross-species reactive and use of xenogeneic cell lines will prevent the inhibitory action of gamma interferon on the producing cells. The vector is purified and concentrated as described in the specification (see also T. Rodriguez et al. J Gene Med 9:233 2007). Vector is administered IT in a volume of 100 .mu.L and the total dose/mouse of approximately 3E3, 3E4 and 3E5 TU/mouse. Vector expressing gamma interferon is identified as Toca 621.
[0394] Tumor Implantation and Vector Injection.
[0395] Nine groups of female BALB/c (99 mice, 9-10 weeks of age) are implanted subcutaneously with CT26.WT tumor cells (Day 0) and then dosed (day 4-7 depending on growth rate of the CT26 tumor; approximately 50-100 mm.sup.3) with vehicle (Groups 1), with control vector [AC3-GFP(V), (Group2), IT Toca 621 vector injection (Groups 3-5), or intravenous Toca 621 vector injection (group 6-8). Group 9 mice have no tumor implanted and are intravenously injected with vector only.
[0396] Data Analysis.
[0397] Tumor growth analysis is carried out to 2000 mm.sup.3 or to 60 days based on whichever comes first. 10 mice from each group will be plotted for tumor size over time. Statistical significance are determined using analysis of variance (ANOVA). P values of <0.05 are considered statistically significant in all analyses, which are performed with Prism 5 statistical software (GraphPad Software) or equivalent. In-life observations are also taken to assess any adverse events to Toca 621 administration.
[0398] Results.
[0399] The results of measurement of tumor size over time show a statistically significant difference in the growth of tumors treated with the vector expressing gamma IFN over the tumors in animals that received control vector or vehicle.
Example 26 Anti-Tumor Efficacy Studies with Gamma Interferon Expressing Vector in a Mouse Subcutaneous Tumor Model
[0400] Objective.
[0401] The objective of this study was to assess the effect of a novel MLV based replication-competent retroviral vector carrying the murine gamma interferon sequence (pAC3-mIFNg) on tumor growth, when delivered via intratumoral (IT) injection in BALB/c mice bearing subcutaneous melanoma(Cloudman S91).
[0402] Mice.
[0403] Female BALB/c mice (age .about.8 weeks) were purchased from Jackson Laboratories (Bar Harbor, Me.). Mice were acclimated for 7 days after arrival before start of studies.
[0404] Cells.
[0405] Clone M-3 Cloudman S91 cells (ATCC, Manassas Va.) are derived from an irradiation induced malignant melanoma from a C.times.DBA F1 mouse. Cells were cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells were resuspended in PBS (Hyclone, Logan Utah) for implantation. S91 cells (1E5 in 100 .mu.L) were injected into the right flank of BALB/c mice.
[0406] Vectors.
[0407] Vectors preparations are made by transient transfection (or from a producer cell line after infection of a second cell line with the infectious virus from the initial transfection with titers of approximately 3E6TU/ml. For initial studies vector was not further purified or concentrated. For follow on experiments to determine full dose response curves, high titer purified material was prepared with a titer approximately 10{circumflex over ( )}8 TU/ml. To achieve high titer material, the human cell line HT1080 was used for production of the mouse gamma interferon vector as gamma interferon is poorly cross-species reactive and use of xenogeneic cell lines prevents the inhibitory action of gamma interferon on the producing cells. The vector was purified and concentrated as described in the specification above. Vector was administered IT in a volume of 100 .mu.L and the total dose/mouse of approximately 3E3, 3E4 and 3E5 TU/mouse. Vector expressing gamma interferon is identified as Toca 621.
[0408] Tumor Implantation and Vector Injection.
[0409] Two groups of female BALB/c (approx. 20 mice, 9-10 weeks of age) were implanted subcutaneously with S91 tumor cells. Mice with tumors reaching approximately 50-125 mm3 were randomized and injected IT with control vector, Toca 511 (Groups 1, N=10), or Toca 621 vector (Group 2, N=5).
[0410] Data Analyses.
[0411] Tumor growth analysis was carried out to 600 mm3 or to 45 days based on whichever comes first. All mice from each group were plotted for tumor size over time. Statistical significance was determined using analysis of variance (ANOVA). P values of <0.05 are considered statistically significant in all analyses, which are performed with Prism 5 statistical software (GraphPad Software) or equivalent. In-life observations were also taken to assess any adverse events to Toca 621 administration.
[0412] Results.
[0413] A statistically significant reduction in average tumor volume was measured in mice with tumors injected with a single dose of Toca 621 vector expressing gamma IFN compared to mice with tumors injected with control Toca 511 vector (p=0.003) see FIG. 35.
Example 27 Intravenous Gene Delivery Using a Replicative Retroviral Vector
[0414] Objective.
[0415] The objective of this study was to assess the effectiveness of intravenous delivery of a novel MLV based replication-competent retroviral vector carrying the marker green fluorescent protein (AC3-GFP(V)) to U87 gliomas implanted in the brains of nude mice.
[0416] Mice.
[0417] Female athymic nude-Foxn1{circumflex over ( )}nu (nude) mice (age .about.8 weeks) were purchased from Harlan (Indianapolis Ind.). Mice were acclimated for 7 days after arrival. Mice underwent surgical placement of an indwelling guide cannula with a 3.0 mm projection implanted into the right striatum, and fitted with a cap containing a 3.5 mm projection. The stereotaxic coordinates are AP=+0.5 mm, ML=-1.8 mm (from bregma).
[0418] Cells.
[0419] U-87 MG cells (ATCC, Manassas Va.) are derived from a malignant glioma from a 44 year old Caucasian female. Cells were cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells are resuspended in PBS (Hyclone, Logan Utah) for implantation. U-87 MG cells (1E5 in 1 .mu.L) were infused at 0.2 .mu.L per minute (5 minutes, followed by a hold of 5 minutes) intracranially (IC) through an injection cannula with a 3.5 mm projection inserted through the guide cannula.
[0420] Vectors.
[0421] Vector preparations were made by transient transfection and all had a titer of approximately 2.8E7TU/ml. Vector was administered intratumorally (IT) in a volume of 5 ul or less for a minimum total dose/mouse of approximately 1.4E45 TU/mouse. Intravenous injections were done through the tail vein with 2.8E6/100 uL.
[0422] Tumor Implantation and Vector Injection.
[0423] Five groups of female athymic nude-Foxn1{circumflex over ( )}nu mice (16 mice, 9-10 weeks of age) were implanted IC with U-87 tumor cells (Day 0) then dosed IT or IV (day 4-7 depending on growth rate of the U87 cells) with vehicle IV (Group 1), with vector IV (Group2), IT with a blood/brain barrier disruptor Vardenafil and vector (Group 3), IT with Vardenafil and vector (Groups 4), or IT with vector (group 5). 14 days after vector injection mice were sacrificed and tumors are isolated and analyzed for GFP expression.
[0424] Data Analysis.
[0425] U87 cells from disrupted tumors isolated from the mice were analyzed by flow cytometry for the percentage GFP positive from groups 2-5 (FIG. 26). Histogram analysis was also done on groups 1, 3, and 5 to measure the distribution of GFP signal in isolated U87 cells (FIG. 27).
[0426] Results.
[0427] Intravenous delivery of GFP was as equally effective as intratumor injection of U87 glioma cells intracranially implanted into nude mice.
Example 28 Construction of Replication Competent Retroviral Vector Encoding the Human IL-2 Gene
[0428] The replication competent retroviral vector, pAC3-hIL2, encoding the human IL2 gene, is derived from the backbone of pAC3-yCD2 vector. The pAC3 backbone in the pAC3-hIL2 vector-encoding plasmid DNA was isolated by endonuclease digestion of the pAC3-yCD2 plasmid DNA with Psi I and Not I. The cDNA sequence of human IFN-.gamma. gene was identified and confirmed using sequences obtained from different accession numbers (BC066255 and BC066257). Sequence alignment of the two revealed identical sequence. The open reading frame of the human IFN-.gamma. (SEQ ID NO: 40) was synthesized with Psi I and Not I restriction enzyme site present at each end of the DNA fragment for subsequent insertion at the corresponding site in the pAC3 backbone. The resulting construct, pAC3-hIL2, encodes 4 genes: the gag, the pol, the env, and the human IL2. (FIG. 28).
[0429] Vector stock is produced by transient transfection of the vector-encoding plasmid DNA into 293T cells using calcium phosphate method. Eighteen hours post transfection, the culture was replaced with fresh medium. Twenty-four hours post medium replacement, the supernatant containing the vector was collected and filtered through a 0.45 .mu.m filter and used immediately or stored in aliquots at -80.degree. C. for later use. Twenty micro-liter of the collected vector stocks was used to infect human prostate cancer cells, PC3. Twenty-four hours post infection, AZT was added to the cells to inhibit further viral replication. Forty-eight hours post infection, genomic DNA of infected PC3 cells was extracted for titer assay. The titer of the l17 vector stocks was determined by qPCR with an inclusion of standards of known copy numbers.
[0430] The expression of human IL-2 is first tested at the RNA level. Total RNA is extracted from transduced CF2TH and HT1080 cells at 5 days post infection using standard RNA extraction method. RT-PCR was performed to detect the expression of human IL-2. Fifty nano-gram of total RNA was used in the RT reaction to generate cDNA. One tenth of the volume from RT reaction was subsequently used for PCR using PCR primer set specific for human IL-2. Result from RT-PCR shows that human IL-2 is expressed in HT1080 cells transduced with pAC3-hIL2 vector.
[0431] The expression of secreted human IL-2 protein was tested by standard ELISA. Vector stock collected from day 5 post infection was serially diluted in the ELISA assay in order to obtain a linear range between protein concentration and dilution factor. The result showed that human IL-2 protein is secreted at a higher concentration by the CF2TH cells than HT1080 at day 5 post infection.
Example 29 Anti-Tumor Efficacy Studies with Interleukin 2 Expressing Vector in a Mouse Subcutaneous Tumor Model
[0432] Objective.
[0433] The objective of this study is to assess the effect of a novel MLV based replication-competent retroviral vector carrying the murine leukocytotrophic hormone interleukin 2 (IL-2) sequence (pAC3-mIL2) on tumor growth, when delivered via intratumoral (IT) injection in BALB/c mice bearing subcutaneous colon carcinoma (CT26.WT).
[0434] Mice.
[0435] Female BALB/c mice (age .about.8 weeks) are purchased from Jackson Laboratories (Bar Harbor, Me.). Mice will be acclimated for 7 days after arrival before start of studies.
[0436] Cells.
[0437] CT26.WT cells (ATCC, Manassas Va.) are an N-nitroso-N-methylurethane-(NNMU) induced, undifferentiated colon carcinoma cell line. Cells are cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells are resuspended in PBS (Hyclone, Logan Utah) for implantation. CT26.WT cells (2E5 in 100 .mu.L) are injected into the right flank of BALB/c mice.
[0438] Vectors.
[0439] Vector preparations are made by transient transfection (or from a producer cell line) with titers of approximately 6E6TU/ml. For initial studies vector is not further purified or concentrated. For follow on experiments to determine full dose response curves, high titer purified material is prepared with a titer expected around 10E8/ml. Vector is administered IT in a volume of 100 .mu.L and the total dose/mouse of approximately 6E5 TU/mouse. Vector expressing gamma interferon is identified as Toca IL2.
[0440] Tumor Implantation and Vector Injection.
[0441] Five groups of female BALB/c (55 mice, 9-10 weeks of age) are implanted subcutaneously with CT26.WT tumor cells (Day 0) and then dosed (day 4-7 depending on growth rate of the CT26 tumor; approximately 50-100 mm.sup.3) with vehicle (Groups 1), with control vector [AC3-GFP(V), (Group2), IT Toca IL2 vector injection (Groups 3), or intravenous Toca IL2 vector injection (group 4). Group 5 mice have no tumor implanted and are intravenously injected with vector only.
[0442] Data Analysis.
[0443] Tumor growth analysis is carried out to 2000 mm.sup.3 or to 60 days based on whichever comes first. 10 mice from each group will be plotted for tumor size over time. Statistical significance will be determined using analysis of variance (ANOVA). P values of <0.05 are considered statistically significant in all analyses, which are performed with Prism 5 statistical software (GraphPad Software) or equivalent. In-life observations are also taken to assess any adverse events to IL-2 expression during treatment.
[0444] Results.
[0445] Delivery of IL-2 by replicating MLV reduces and in some instances clears tumors burden from the BALB/c CT26 mouse model.
Example 30 Tumor Explants have Multiple Copies of the Vector Genome and Show Continued Susceptibility to Super-Infection
[0446] In order to examine in more detail the mechanism of action of the replicating retrovirus tumors from some animals in the mouse and human tumor models described in example 9 (athymic nude-Foxn1{circumflex over ( )}nu (nude) mice with Human U87 intracranial implants) and example 10 (BALB/c mice with syngeneic CT26 intracranial implants) were explanted and examined for 5-FC sensitivity, vector copy number/diploid genome, and CD protein expression.
[0447] Explant Assignments.
[0448] The experimental design is summarized below. The study consisted of 5 tumor explants. The history of each tumor removed for implantation is given below. Explant History:
TABLE-US-00019 # of 5-FC dosings Animal before Cell Dosing # Study Treatment explant Type regimen 833 Example 9 AC3-yCD2(V) 4 Human QD, 7 days FIG. 8 E6 + 5-FC U87 every 21 days 953 Example 9 AC3-yCD2(V) 4 Human QD, 7 days FIG. 8 E5 + 5-FC U87 every 21 days 969 Example 9 AC3-yCD2(V) 4 Human QD, 7 days FIG. 8 E5 + 5-FC U87 every 21 days 31 Example 10 AC3-yCD2(V) 3.5 Mouse BID 7 days, FIG. 9 E5 + 5-FC CT26 every 17 days 61 Example 10 AC3-yCD2(V) 3 Mouse BID 7 days, FIG. 9 E4 + 5-FC CT26 every 17 days
[0449] The 5-FC cell killing assays were carried out as described in Example 5 above, measuring viability after 8 days of 5-FC treatment.
[0450] Copy number/microgram of DNA was determined by PCR as described for the vector titering assay in Example 5, and converted to copy number/diploid genome by dividing by 150,000, the approximate number of diploid mouse/human genomes in 1 microgram of genomic DNA. Western Blot analysis was performed on 1E6 cells/lysate in RIPA buffer using antibodies from clone 83A25 for GP70 and Abcam anti-CD antibody ab3525 for the CD protein. Cell explants underwent super-infection procedures with a AC3-eGFP(V) and a mock procedure to determine which explants were potentially further infectable. The extent of GFP expression was measured by FACS analyses, with uptake and expression of GFP indicating the relative susceptibility to further infection.
[0451] Results. 5-FC Cell Killing Assay and Copy Number of Integrated Vector.
[0452] Cultured explants were tested for 5-FC sensitivity by generating a killing profile from treated cell lines at varying 5-FC concentrations (summarized in the Table below). Results from the killing profile measured by MTS viability assay show that U87 human tumors derived from animals #833, 953, 969 on average (IC.sub.50=0.009 mM) had a similar response compared to an in vitro U87 positive control (IC.sub.50=0.008 mM) to 5-FC treatment (FIG. 29A). Analysis of CT26 murine tumors (Example 10) showed that the 5-FC responsive animal #61 had an IC.sub.50 of 0.003 mM (FIG. 29B) which is similar to in vitro 100% transduced CT26 results (IC.sub.50=0.001 mM). Animal #31 was poorly responsive to 5-FC (FIG. 29B). PCR results for copy number per cell are also shown in the table below.
TABLE-US-00020 5-FC sensitivity Vector copy number Animal # Study IC50 (mM) per diploid genome 833 Example 9 0.009 22.3 953 Example 9 0.009 9.6 969 Example 9 0.009 18.7 61 Example 10 0.003 6.0 31 Example 10 Not sensitive 0.9
[0453] Western Blot Analysis of GP70 and CD Protein Expression.
[0454] Further analysis of cells by western blot from the CT26 study showed that both tumor explants derived from mice #31 and #61 had observable GP70 protein expression when using U87+AC3-yCD2(V) infected lysates as a reference positive control (FIG. 30).
[0455] However, analysis of CD expression showed that only #61 still had observable CD expression. Cells from #31 were run in duplicate wells (#31(A) and #31(B)) to verify negative CD gene expression results.
[0456] GFP Expression after AC3-eGFP(V) Transduction.
[0457] Attempts to transduce explants with an MLV vector expressing GFP showed that U87 tumors derived from animals #833, 953, 969 were scarcely transducible (<0.5%). CT26 explanted tumor cells derived from animal #61 could be partially transduced (7% GFP positive) while explanted cells from animal #31 could not (FIG. 31).
[0458] All U87 gliomas isolated from the brains of nude mice after 4 full cycles of 5-FC treatment were still sensitive to 5-FC treatment in vitro with an IC.sub.50 the same as in vitro transduced U87 and, surprisingly, showed multiple vector superinfections had taken place. Two CT26 tumors were isolated from BALB/c mice after 3 and 3.5 cycles of 5-FC treatment. Of the two, only one tumor showed 5-FC sensitivity while the other did not. Further analysis showed that the 5-FC resistant tumor is refractory to further MLV transduction, expresses GP70 but no longer expresses the CD, and has low copy number compared to the other CT26 and all U87 explants tested. These observations show that whereas a virus that has undergone a deletion of the CD gene behaves as expected for a normal retrovirus and excludes further infection, cells infected with vector carrying the CD transgene behave atypically and allow multiple superinfections (range: 6.0-23.3, mean 14.5 copies per diploid genome). Typical tumors are not diploid but are polyploid with a genome larger than the diploid genome. This would further increase the actual vector copy number per cell. The multiple vector copy numbers contributes to the therapeutic effect as more of the protein derived from the transgene (in this case CD) is produced than from a single vector integration. It also means that in general, even if some members of a viral vector population undergo rearrangements, other members will donate protein activity (in this case sensitivity to 5-FC). The experiments described here also provide a method of testing a recombinant replication competent retrovirus for the property of multiple infections of a target cell population.
Example 31: Direct Measurement from Excised Tumors Treated with AC3-yCD2(V) Shows Unexpectedly High Levels of Viral Vector Copies Per Genome and Susceptibility to Superinfection in the Syngeneic Tu2449 Glioma Model
[0459] Objective.
[0460] This study was conducted to compare the efficacy of two dose levels of AC3(V)-yCD2 (aka Toca511) delivered via IC injection in combination with 5-FC treatment in a TU-2449 glioma tumor bearing, immunocompetent mouse model, and examined survival in the setting of active tumor growth. TU-2449 cells implanted IC in syngeneic B6C3F1 mice have been used as an experimental murine glioma model. This model was also used for survival and short term (15-18 day) experiments where tumors were implanted, treated with vector and dosed short term with 5-FC then excised for further characterization of gene copy number and CD activity.
[0461] Mice.
[0462] Female B6C3F1 mice (age .about.8 weeks) were purchased from Harlan (Indianapolis Ind.). Mice were acclimated for 7 days after arrival.
[0463] Mice underwent surgical placement of an indwelling guide cannula with a 3.0 mm projection implanted into the right striatum, and fitted with a cap containing a 3.5 mm projection. The stereotaxic coordinates were AP=+0.5 mm, ML=-1.8 mm (from bregma).
[0464] Cells.
[0465] TU-2449 cells (Smilowitz et al. J Neurosurg. 2007 106:652-659 2007) derived originally from Glial fibrillary acidic protein (GFAP)-v-src transgenic mice, were cultured in Dulbecco's modified Eagles medium with 10% fetal bovine serum, sodium pyruvate, and Glutamax (Hyclone, Logan Utah, and Invitrogen, San Diego Calif.). Cells were resuspended in PBS (Hyclone, Logan Utah) for implantation. TU-2449 cells (1E4 in 1 .mu.L) were infused at 0.2 .mu.L per minute (5 minutes, followed by a hold of 5 minutes) IC through an injection cannula with a 3.5 mm projection inserted through the guide cannula.
[0466] The study consisted of 6 groups of female mice (see Table below). On day 0, mice from Groups 1, 3, 4, 6, and 7 underwent intracranial implantation of 1E4 TU-2449 cells. Group 8 mice were not implanted with tumor. On Day 4, mice were injected (IC; 5 .mu.L/mouse) with vehicle (Group 1); IC with AC3-yCD2(V) at 1.7E5 TU/g (Groups 6, 7); IC with AC3-yCD2(V) at 1.7E6 TU/g (Groups 3, 4); Group 8 mice were not treated. Starting on Day 10, mice were treated IP BID for 4 consecutive days with PBS (Groups 1, 3, 7) or 5-FC (500 mg/kg/dose, Groups 4, 6, 8). Cycles of 4 days BID treatment with PBS or 5-FC followed by 10 days of viral spread were repeated. Survival analysis to Day 180 was performed on 10 mice each from Groups 3-7.
Group Assignments
TABLE-US-00021 [0467] N per analysis category Scheduled Group Treatment Survival Sacrifice 1 Control (vehicle 1 at day 10 injection) + PBS 3 AC3-yCD2(V) 10 E6 + PBS 4 AC3-yCD2(V) 10 3 at day 10, 24, E6 + 5-FC 38, and 52 6 AC3-yCD2(V) 10 3 at day 10, 24, E5 + 5-FC 38, and 52. 7 AC3-yCD2(V) 10 E5 + PBS 8 5-FC 1 at day 10, 24, (no tumor) 38, and 52 TOTAL 40 29
[0468] AC3-yCD2(V) (5 .mu.L) was infused at 0.33 .mu.L per minute (15 minutes, followed by a hold of 5 minutes) intracranially through an injection cannula with a 3.5 mm projection inserted through the guide cannula. 5-FC (500 mg/kg/dose) or PBS (800 .mu.L) was administered IP BID for 4 consecutive days starting at days 10, 24, 38, and 52.
[0469] Short Term Experiments to Determine the Level of Viral Genome and Super-Infection in Tu2449 Tumors In Vivo.
[0470] The study consisted of 6 groups of female mice (see Table below). All groups underwent intracranial administration into the right striatum of 1E4 TU-2449 cells administered/mouse on Day 0. At Day 4, all groups received intracranial/intratumoral administration of AC3-YCD2(V) vector at 2.4E6 TU/5 ul (Lot # T511019) or PBS buffer control. Two days of BID 5-FC administration began when the mice started losing weight (approximately 15 days post-tumor implantation). Group 5 had 5-FC delivered by oral gavage (OG) and all other groups IP. One more dose of 5-FC was given 1 hour before sacrifice the following day. From each brain, the tumor was isolated and processed directly into RIPA buffer for analysis of 5-FC and 5-FU by HPLC. A small portion of the tumor was retained for western blot analysis.
Group Assignments and Dose Levels
TABLE-US-00022 [0471] Group Treatment Route TX Route Dosing N 1 AC3-YCD2(V) IC NONE N/A N/A 3 2 PBS IC 5FC IP 250 mg/kg 4 3 PBS IC 5FC IP 500 mg/kg 2 4 AC3-YCD2(V) IC 5FC IP 250 mg/kg 3 5 AC3-YCD2(V) IC 5FC OG 250 mg/kg 2 6 AC3-YCD2(V) IC 5FC IP 500 mg/kg 2 Total animals 16 IC--intracranial; IP--intraperitoneal; OG--oral gavage
[0472] AC3-yCD2(V)(5 .mu.L) was infused at 0.33 .mu.L per minute (15 minutes, followed by a hold of 5 minutes) intracranially at the same coordinates as TU-2449 cells were injected. 5-FC or PBS was administered IP or OG BID for 2 consecutive days and 1 hour before sacrifice.
[0473] Tissue Processing Procedures.
[0474] From each brain, tumors were isolated and trimmed if large enough for multiple analyses (more than 0.05 g). Tumors sections for HPLC analysis were crushed in a 1.5 mL centrifuge tube using a plunger from a 1 mL syringe. Crushed samples were mixed with 150 uL RIPA buffer and vortexed vigorously for 10 minutes. Samples were spun at 4.degree. C. at 20000 rcf for 10 minutes. Supernatants were removed and mixed thoroughly with 150 uL of 10% trichloroacetic acid and spun as above. Supernatants were removed for analysis by the Agilent HPLC unit with a Hypersil BDS C18 column run isocratically at 1 mL/min with 95% Buffer A containing 50 mM ammonium phosphate and 0.1% tetra-n-butylammoniumperchlorate with pH adjustment of the buffer to 2.1 with phosphoric acid and 5% Solvent B which is 100% methanol (see WI RD-053). The run time is 5 minutes with each sample run twice. The photodiode detector array scans from 190 to 350 nm with chromatograms selected to display at 285 nm for 5-fluorocytosine and 264 nm for 5-fluorouracil. Data was expressed in relative milli absorbance units (mAU) of peak area from the chromatograms.
[0475] Protein Gels and Western Blots.
[0476] When sufficient tumor was available, tumor fragments for protein gels and Western blots were mixed with a separate aliquot of RIPA lysis buffer, and 20 ug of total protein from each sample was electrophoresed on polyacrylamide gels, Western blotted and the blot developed with sheep anti-yeast CD antibody as in Example 30. Western blot data was scanned and quantified using BioRad Quantity One software (version 4.6.7).
[0477] QPCR on Tumor Fragments.
[0478] Remaining pellets, after supernatants were removed for HPLC analysis, were extracted for genomic DNA. Samples were analyzed by qPCR for proviral integration copy number using primers and probe for MLV LTR as in example 30. Samples were also analyzed in parallel using previously characterized primers and probes for the amphotropic env gene (Env2) and the CD gene (yCD2).
[0479] yCD2 Primer and Probe Set:
TABLE-US-00023 5' AC3-YCD2(V) yCD2 Primer: (SEQ ID NO: 67) ATCATCATGTACGGCATCCCTAG 3' AC3-YCD2(V) yCD2 Primer: (SEQ ID NO: 68) TGAACTGCTTCATCAGCTTCTTAC yCD2 Probe: (SEQ ID NO: 69) /5FAM/TCATCGTCAACAACCACCACCTCGT/3BHQ_1/
These primers and probe target and amplify the CD gene exclusively.
[0480] Env2 Primer and Probe Set:
TABLE-US-00024 Env2-Forward: (SEQ ID NO: 70) AACCTCAACCTCCCCTACAAGT Env2-Reverse: (SEQ ID NO: 71) GTTAAGCGCCTGATAGGCTC Env2-Probe: (SEQ ID NO: 72) /5TEX615/AGCCACCCCCAGGAACTGGAGATAGA/3IAbRQSp/
[0481] These primers and probe target and amplify the envelope (Env) gene exclusively.
[0482] Results. Survival Analysis.
[0483] The Kaplan Meyer survival plot is shown in FIG. 32. The median survival of AC3-yCD2(V) control groups treated with PBS (Group 3 and 7) was approximately 33-38 days. The survival medians of mid and high dose AC3-yCD2(V) in combination with 5-FC (Group 6 and 4) were not reached before sacrifice at 189 days. Log-Rank (Mantel-Cox) pair-wise comparison showed no difference in survival between the two control groups; AC3-yCD2(V) E5 dose plus PBS (Group 7) and AC3-yCD2(V) E6 dose plus PBS (Group 3).
[0484] AC3-yCD2(V) treatment at both dose levels in combination with 5-FC resulted in prolonged survival. A statistically significant survival advantage was observed for AC3-yCD2(V) E5 plus 5-FC (Group 6) treated mice compared to vector plus PBS control (Group 7) mice (p<0.0354, hazard ratio 0.2605, 95% CI 0.07439 to 0.9119). A statistically significant survival advantage was observed for AC3-yCD2(V) E6 plus 5-FC (Group 4) treated mice compared to vector plus PBS control (Group 3) mice (p<0.0036, hazard ratio 0.1130, 95% CI 0.02598 to 0.4911).
[0485] Short Term Experiments to Determine the Level of Viral Genome and Super-Infection in Tu2449 Tumors In Vivo
[0486] HPLC Analysis.
[0487] In vivo conversion of 5-FC to 5-FU was detected by HPLC in all groups dosed with Toca511 and 5-FC (FIG. 33). Group 1 that was given AC3-yCD2(V) but no 5-FC had neither 5-FC nor 5-FU detectable signals as expected (FIG. 33).The small counts observed (47-84) for 5-FC is attributable to background from nearby peaks on the chromatography trace. Groups 2 and 3 that were not dosed with AC3-yCD2(V) but dosed with varying levels of 5-FC had detectable 5-FC signals but no signal for 5-FU. Group 4 mice dosed with AC3-yCD2(V) and 5-FC IP and Group 5 mice dosed with AC3-yCD2(V) and 5-FC OG had comparable signal levels of 5-FU and very low or background levels of 5-FC. Group 6 mice dosed with Toca-511 and high levels of 5-FC showed readily detectable levels of 5-FU and low signal for 5-FC.
[0488] CD Western Blot Analysis.
[0489] Tissue samples from isolated TU-2449 tumors were processed for western blot analysis of CD expression. All groups (1, 4, and 5) treated with AC3-yCD2(V) had readily observable CD expression while Group 2 that was not given AC3-yCD2(V) did not have detectable CD expression (FIG. 34).
[0490] PCR Analysis of Genomic DNA Isolated from Tumors.
[0491] The remaining pellets, after supernatants were removed for HPLC analysis, were extracted for genomic DNA. Samples were analyzed by quantitative PCR for proviral integration using the standardized assay and MLV-LTR primers and probe. Parallel assays using the envelope and CD gene primers and probes analyses gave similar C(t) values showing that the viral genome appeared quite stable. An in vitro transduced cell line served as positive control. The negative control was genomic tumor DNA from Group 2 that was not dosed with AC3-yCD2(V) and did not have detectable signal for any of the qPCR protocols.
[0492] Summary Table of relative CD protein levels, viral vector copy #, and relative levels of 5-FU production
TABLE-US-00025 Relative 5-FU CD Copies#/ (relative protein diploid peak area Group Mouse # levels genome units) Group 1 194 21,746 1.4 0 199 No Data 14.6 0 Group 2 191 162 0 0 187 75 0 0 188 -453 0 0 Group 4 185 17,349 1.8 1642 200 45,446 7.3 1576 Group 5 189 25,417 3.7 942 198 23,660 6.6 1371
[0493] For samples with enough starting material for all three analyses to be done, the relationship between integrated MLV copy number, expression of CD, and the amount of 5-FC to 5-FU conversion is summarized in the above table (199 did not have material for Western analysis). The relative CD protein levels (estimated from Western blots) vary over a three-fold range and the DNA copy number over a 5-fold range. There is some correlation between DNA copy number and relative level of CD expression. All of the tumors have vector copy numbers/genome above 1, showing that even at this early time-point after vector administration (13-14 days) superinfection of tumor cells is a usual occurrence, and may contribute to observed therapeutic effects (FIG. 32). The CD values displayed represent the values after average background correction, and the Group 2 numbers represent the variability in that background.
[0494] This study supports the proposed mechanism of action and shows that this efficacy can be attributed to the conversion of the 5-FC prodrug into the anticancer drug 5-FU after delivery of the CD gene by AC3-yCD2(V). Using the TU-2449 mouse glioma model, AC3-yCD2(V) treatment in combination with 5-FC resulted in efficient in vivo conversion of 5-FC into 5-FU. 5-FC was converted efficiently into 5-FU at two dose levels (500 and 250 mg/kg) as 5-FC levels were at least ten fold lower than the controls, and 5-FU levels were readily detectable in AC3-yCD2(V) treated but not in the untreated controls. IP or OG delivery of 5-FC did not affect the efficiency of conversion. Tumors isolated from mice given AC3-yCD2(V) had observable expression of CD protein that had some correlation with the numbers of copies of the vector genome. The number of integrated vector genomes ranged from 1.4 to 15 copies/diploid genome (mean: 5.9). The infection here was the result of about 12 days infection and previous experiments with GFP vectors in other models suggest that this corresponds to infection of approximately 50% of the cells in the tumor, giving an adjusted vector copy number/cell of 11.8 copies/diploid genome. Typical tumors are not diploid but are triploid or further polyploid with a genome larger than the diploid genome. This would further increase the actual vector copy number per cell. The experiments described here also provide a method of testing a recombinant replication competent retrovirus for the property of multiple infections of a target cell population. These observations support the conclusion that AC3-yCD2(V) is efficiently delivering a functional CD gene for expression in glioma cells. In the efficacy study with this model, almost no spread of vector from the site of injection (tumor in the right cerebrum) was observed in the first 24 days. The observations in this study show that over the same initial period of time there is extensive viral vector infection of the tumor, showing that infection is quite tumor specific, and that already at this early time-point there is extensive super-infection of the tumor cells by the viral vector.
Example 32 Clonal Analyses of HT1080 Cell Line Infected with AC3-yCD2(V) Shows that the Majority of the Clones have Multiple Copies of the Viral Vector Genome and were Susceptible to Super-Infection
[0495] The human sarcoma line HT1080 (ATCC: CCL 121) was grown in tissue culture under standard culture conditions. 2E7 cells were infected with AC3-yCD2(V) made by transient transfection on 293 cells, at a multiplicity of 0.1, allowed to grow for 14 days, and frozen down as a pool. About 1 month later cells were thawed and clonal cell lines from this culture were isolated by limiting dilution in 96 well dishes at 0.3 cells/well. The clones that grew out were expanded and analyzed by qPCR with the MLV LTR primers for vector genome copy number per microgram DNA in triplicate. This was converted to copy number/diploid genome by dividing by 150,000, as described in example 30. The Table below lists the clones that were analyzed and the corresponding copy number for the viral vector genome. Only 1 of 10 clones (13-5) has approximately 1 copy of the viral genome per diploid genome. The range was 0.9 to 20.4 copies/cell and the mean copy number was 10.6 copies/cell. Typical tumors are not diploid but are triploid or further polyploid with a genome larger than the diploid genome. This would further increase the actual vector copy number per cell. It is well known that normally a viral infection of this nature leads to a single or few copies of viral genome/cell, due to resistance to superinfection through receptor masking or down regulation (see, for example, Ch3 p 104 of "Retroviruses" J M Coffin, S H Hughes & H E Varmus, 1997 Cold Spring Harbor Laboratory Press, Cold Spring Harbor N.Y.).
[0496] Listing of viral vector copy number in infected HT1080 clones infected with AC3-yCD2 and expanded.
TABLE-US-00026 Average copy number/ Sample # Test article genome 1 Negative control 0 (HT1080) 2 Positive control 1.89 (recently transduced HT1080 pool) 3 Clone# 3-5 18.26 4 Clone# 4-1 5.28 5 Clone# 7-1 18.87 6 Clone# 8-3 14.04 7 Clone# 9-1 15.29 8 Clone# 10-1 9.73 9 Clone# 11-1 20.44 10 Clone# 12-6 6.08 11 Clone# 13-5 0.93 12 Clone# 19-2 5.24
[0497] Therefore this is an unexpected and surprising result that confirms the in vivo tumor model data of Examples 30 and 31. More particularly, the data demonstrate that this virus allows multiple super-infections in the great majority of the cells it infects, unlike normal MLV infection. The experiments described here also provide a method of testing a recombinant replication competent retrovirus for the property of multiple infections of a target cell population.
Example 33: miRNA Knockdown Experiments
[0498] Plasmid Construction.
[0499] Single or four tandem repeats of 142-3pT completely complementary to the sequence of miR 142-3p were synthesized with an endonuclease restriction site Not I at both ends and cloned into the corresponding Not I site downstream of the IRES-GFP cassette in the RCR vector (FIG. 11C-D). The sequence of the single 142-3pT:
TABLE-US-00027 (SEQ ID NO: 35) gcggccgcGTCGACTCCATAAAGTAGGAAACAC TACAgcggccgc
and the sequence of four tandem repeats of 142-3pT henceforth 142-3pT4X:
TABLE-US-00028 (SEQ ID NO: 36) gcggccgcGTCGACTCCATAAAGTAGGAAACACTACACGATTCCATAAA GTAGGAAACACTACAaccggtTCCATAAAGTAGGAAACACTACATCACT CCATAAAGTAGGAAACACTACAgcggccgc
were synthesized by BioBasic Inc. The underlined sequences are sequence complementary to miR-142-3p in both mouse and human. The sequence of the synthesized DNA fragments were confirmed before and after cloning into the pAC3-emd vector using the primers:
TABLE-US-00029 (SEQ ID NO: 62) 5'-CTGATCTTACTCTTTGGACCTTG-3', and (SEQ ID NO: 63) 5'-CCCCTTTTTCTGGAGACTAAATAA-3'
[0500] Cell Culture.
[0501] Human astrocytoma cells U-87MG, human prostate adenocarcinoma cells PC3, human lymphoblastic leukemia cells Sup-T1, human histocytic lymphoma cells U-937 and mouse T lymphoblastic cells were obtained from ATCC. 293T, U-87MG, PC3 and EL-4 cells were cultured in complete DMEM medium containing 10% FBS (Hyclone), sodium pyruvate, glutamax, and penicillin/streptomycin (Cellgro). Sup-T1 and U-937 cells were cultured in complete RPMI medium containing 10% FBS, glutamx and penicillin/streptomycin.
[0502] Virus Production.
[0503] Virus stock was produced by transient transfection of 293T cells using calcium phosphate precipitation method. Cells were seeded at 2e6 cells per 10 cm petri dish the day before transfection. Cells were transfected with 20 .mu.g of pAC3-emd, pAC3-emd-142-3pT or pAC3-emd-142-3pT4X the next day. Eighteen hours post transfection, cells were washed with PBS twice and incubated with fresh complete culture medium. Viral supernatant was collected approximately 42 hours post transfection and filtered through a 0.45 .mu.m. Viral supernatant were stored in aliquots at -80.degree. C.
[0504] Viral infection was performed by adding viral stock at 1:50 dilution in 1 mL total volume in each well. AZT at 40 .mu.m was added to prevent further viral replication and cells were harvested 48h post infection for gDNA isolation. Viral titer was determined by quantitative real time PCR (qPCR) using the following primer set and probe which will bind to all proviral DNA derived from the vectors as well as proviral DNA containing deleted IRES-GFP cassette: 5'-AGCCCACAACCCCTCACTC-3' (SEQ ID NO:64), 5'-TCTCCCGAT CCCGGACGA-3' (SEQ ID NO:65), and 5'-FAM-CCC CAA ATG AAA GAC CCC CGC TGA CG-BHQ-3' (SEQ ID NO:66). The reaction was performed in a total volume of 20 .mu.L containing 2.times. iQ SuperMix (BioRad); 0.3 .mu.M of each primer and 0.1 .mu.M of the probe. PCR reaction was performed in triplicates using CFX-96 (BioRad) thermo cycler with the following parameters: 95.degree. C. 10 min; and 40 cycles of 95.degree. C. 15s; 60.degree. C. 1 min. Viral titer reported in transduction unit per milliliter (TU/mL) was determined by calculation of Ct values derived from a standard curve ranging from 1e7 copies to 10 copies of plasmid DNA and from known amount of gDNA input, number of cells, and dilution factor of viral stock per reaction used in each reaction.
[0505] Viral Replication Kinetics.
[0506] To monitor viral replication in infected cells, 2e5 U-87MG cells, or 1e6 EL4, Sup-T1, and U-937 cells were infected with pAC3-emd, pAC3-emd-142-3pT, or pAC3-emd-142-3p4X at an MOI of 0.1 (U-87MG cells) or an MOI of 2 (EL4, Sup-T1, and U-937 cells). Every 3-4 days, a portion of cells were passaged for continuing monitoring of viral replication, and a portion of cells were harvested for GFP expression by flow cytometric analysis. Cells harvested for flow cytometric analysis were washed with PBS and centrifuged at 1000 rpm for 5 minutes. Cell pellets were resuspended in PBS containing 1% PFA. Percentage of GFP % cells were measured by Becton Dickison Canton II using FL1 channel. Viral replication kinetics were obtained by plotting % GFP positive cells over time.
[0507] Vector Stability Assay and Amplification of IRES-GFP Cassette.
[0508] U-87MG cells at 5e4 cells per well in 6-well plate were infected with tock virus at an MOI of 0.1. At d4 post infection viral supernatant from 70% infected cells were collected and filtered through a 0.45 .mu.m filter unit. A 1:10 dilution of the viral supernatant was then used to infected fresh U-87MG cells seeded the night before. Four-day infected U-87 cells were harvested for gDNA isolation for IRES-GFP PCR. This virus infection cycle was repeated at least 12 times.
[0509] gDNA extraction was carried out using the Maxwell 16 DNA purification kit (Promega). DNA concentration and quality was determined by spectrophotometer using Nanodrop 1000 (Thermo Scientific). To assess the integrity of the IRES-GFP cassette in proviral DNA, standard PCR was performed using the following primer set: 5'-CTGATCTTACTCTTTGGACCTTG-3' (SEQ ID NO:62), and 5'-CCCC TTTTT CTGGAGACTAAATAA-3' (SEQ ID NO:63). The reaction was performed in total volume of 25 .mu.L containing 0.4 .mu.M of each primer, 0.4 mM dNTP and 2.5 unit of SuperTaq (Ambion), and the PCR reaction was performed with the following parameters: 95.degree. C. 10 min; and 40 cycles of 95.degree. C. 15s; 60.degree. C. 1 min. 95.degree. C. 2 min and 40 cycles of 95.degree. C., 15s; 55.degree. C., 30s; 72.degree. C., 1 min, followed by 72.degree. C., 5 min. One fifth of the PCR reaction was loaded on 1% agarose gel to resolve PCR products. The expected PCR product of an intact IRES-GFP cassette is 1.2 kb. PCR products less than 1.2 kb indicates partial or complete deletion in the IRES-GFP region.
[0510] For experiment in which the PCR products were excised from the gel for sequencing, gel extraction kit (Qiagen) was used to obtained PCR product. The same set of primers used for PCR reaction was used for PCR product sequencing.
[0511] miRNA Expression Assay.
[0512] miRNA-enriched RNA was extracted from U-87MG, 293T, Sup-T1, U-937 and Sup-T1 cells by using the mirVana miRNA isolation kit followed by DNase treatment (Ambion) according to manufacturer's protocols. Taqman microRNA reverse transcription kit was used with RT primer for miR-142-3p (assay ID #000464) and RNU6 (assay ID #001093) and sno135 (assay #001230) as endogenous controls for human and mouse cell lines, respectively, to produce cDNA for TaqMan microRNA assay (assay ID # TM000464), (Ambion). Reverse transcription and quantitative PCR reactions were set up and carried out according to manufacturer's protocols. 2.sup.-.DELTA.ct was calculated to obtain miR-142-3p expression relative to endogenous control.
[0513] Having confirmed the expression of miR142-3p in hematopoietic-lineage derived cells, cells were then infected with pAC3-emd, pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vectors, respectively, at MOI 2. The data showed that in EL4 cells viral replication of parental vector pAC3-emd was extremely slow in early time with a lag phase that lasted up to 30 days and reached to maximal infectivity (70% infectivity) by day 80 post infection. In contrast, GFP expression and viral replication of pAC3-emd-142-3pT and pAC3-emd-142-3pT4X vectors were completely abrogated in early time during infection (FIG. 11D).
[0514] A number of embodiments of the disclosure have been described. Nevertheless, it will be understood that various modifications may be made without departing from the spirit and scope of the disclosure. Accordingly, other embodiments are within the scope of the following claims.
Sequence CWU 1
1
721477DNASaccharomyces cerevisiaeCDS(1)..(477) 1atg gtg aca ggg gga
atg gca agc aag tgg gat cag aag ggt atg gac 48Met Val Thr Gly Gly
Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15att gcc tat gag
gag gcg gcc tta ggt tac aaa gag ggt ggt gtt cct 96Ile Ala Tyr Glu
Glu Ala Ala Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30att ggc gga
tgt ctt atc aat aac aaa gac gga agt gtt ctc ggt cgt 144Ile Gly Gly
Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45ggt cac
aac atg aga ttt caa aag gga tcc gcc aca cta cat ggt gag 192Gly His
Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55 60atc
tcc act ttg gaa aac tgt ggg aga tta gag ggc aaa gtg tac aaa 240Ile
Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65 70 75
80gat acc act ttg tat acg acg ctg tct cca tgc gac atg tgt aca ggt
288Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr Gly
85 90 95gcc atc atc atg tat ggt att cca cgc tgt gtt gtc ggt gag aac
gtt 336Ala Ile Ile Met Tyr Gly Ile Pro Arg Cys Val Val Gly Glu Asn
Val 100 105 110aat ttc aaa agt aag ggc gag aaa tat tta caa act aga
ggt cac gag 384Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr Arg
Gly His Glu 115 120 125gtt gtt gtt gtt gac gat gag agg tgt aaa aag
atc atg aaa caa ttt 432Val Val Val Val Asp Asp Glu Arg Cys Lys Lys
Ile Met Lys Gln Phe 130 135 140atc gat gaa aga cct cag gat tgg ttt
gaa gat att ggt gag tag 477Ile Asp Glu Arg Pro Gln Asp Trp Phe Glu
Asp Ile Gly Glu145 150 1552158PRTSaccharomyces cerevisiae 2Met Val
Thr Gly Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15Ile
Ala Tyr Glu Glu Ala Ala Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25
30Ile Gly Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu Gly Arg
35 40 45Gly His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu His Gly
Glu 50 55 60Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly Lys Val
Tyr Lys65 70 75 80Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp
Met Cys Thr Gly 85 90 95Ala Ile Ile Met Tyr Gly Ile Pro Arg Cys Val
Val Gly Glu Asn Val 100 105 110Asn Phe Lys Ser Lys Gly Glu Lys Tyr
Leu Gln Thr Arg Gly His Glu 115 120 125Val Val Val Val Asp Asp Glu
Arg Cys Lys Lys Ile Met Lys Gln Phe 130 135 140Ile Asp Glu Arg Pro
Gln Asp Trp Phe Glu Asp Ile Gly Glu145 150 1553477DNAArtificial
SequenceEngineered cytosine deaminaseCDS(1)..(477) 3atg gtg aca ggg
gga atg gca agc aag tgg gat cag aag ggt atg gac 48Met Val Thr Gly
Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15att gcc tat
gag gag gcg tta tta ggt tac aaa gag ggt ggt gtt cct 96Ile Ala Tyr
Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30att ggc
gga tgt ctt atc aat aac aaa gac gga agt gtt ctc ggt cgt 144Ile Gly
Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45ggt
cac aac atg aga ttt caa aag gga tcc gcc aca cta cat ggt gag 192Gly
His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55
60atc tcc act ttg gaa aac tgt ggg aga tta gag ggc aaa gtg tac aaa
240Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr
Lys65 70 75 80gat acc act ttg tat acg acg ctg tct cca tgc gac atg
tgt aca ggt 288Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met
Cys Thr Gly 85 90 95gcc atc atc atg tat ggt att cca cgc tgt gtc atc
ggt gag aac gtt 336Ala Ile Ile Met Tyr Gly Ile Pro Arg Cys Val Ile
Gly Glu Asn Val 100 105 110aat ttc aaa agt aag ggc gag aaa tat tta
caa act aga ggt cac gag 384Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu
Gln Thr Arg Gly His Glu 115 120 125gtt gtt gtt gtt gac gat gag agg
tgt aaa aag tta atg aaa caa ttt 432Val Val Val Val Asp Asp Glu Arg
Cys Lys Lys Leu Met Lys Gln Phe 130 135 140atc gat gaa aga cct cag
gat tgg ttt gaa gat att ggt gag tag 477Ile Asp Glu Arg Pro Gln Asp
Trp Phe Glu Asp Ile Gly Glu145 150 1554158PRTArtificial
SequenceSynthetic Construct 4Met Val Thr Gly Gly Met Ala Ser Lys
Trp Asp Gln Lys Gly Met Asp1 5 10 15Ile Ala Tyr Glu Glu Ala Leu Leu
Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30Ile Gly Gly Cys Leu Ile Asn
Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45Gly His Asn Met Arg Phe
Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55 60Ile Ser Thr Leu Glu
Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65 70 75 80Asp Thr Thr
Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr Gly 85 90 95Ala Ile
Ile Met Tyr Gly Ile Pro Arg Cys Val Ile Gly Glu Asn Val 100 105
110Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr Arg Gly His Glu
115 120 125Val Val Val Val Asp Asp Glu Arg Cys Lys Lys Leu Met Lys
Gln Phe 130 135 140Ile Asp Glu Arg Pro Gln Asp Trp Phe Glu Asp Ile
Gly Glu145 150 1555480DNAArtificial SequenceHuman codon optimized
cytosine deaminaseCDS(1)..(480) 5atg gtg acc ggc ggc atg gcc tcc
aag tgg gat caa aag ggc atg gat 48Met Val Thr Gly Gly Met Ala Ser
Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15atc gct tac gag gag gcc gca
ctg ggc tac aag gag ggc ggc gtg cct 96Ile Ala Tyr Glu Glu Ala Ala
Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30atc ggc ggc tgt ctg atc
aac aac aag gac ggc agt gtg ctg ggc agg 144Ile Gly Gly Cys Leu Ile
Asn Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45ggc cac aac atg agg
ttc cag aag ggc tcc gcc acc ctg cac ggc gag 192Gly His Asn Met Arg
Phe Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55 60atc tcc acc ctg
gag aac tgt ggc agg ctg gag ggc aag gtg tac aag 240Ile Ser Thr Leu
Glu Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65 70 75 80gac acc
acc ctg tac acc acc ctg tcc cct tgt gac atg tgt acc ggc 288Asp Thr
Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr Gly 85 90 95gct
atc atc atg tac ggc atc cct agg tgt gtg gtc ggc gag aac gtg 336Ala
Ile Ile Met Tyr Gly Ile Pro Arg Cys Val Val Gly Glu Asn Val 100 105
110aac ttc aag tcc aag ggc gag aag tac ctg caa acc agg ggc cac gag
384Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr Arg Gly His Glu
115 120 125gtg gtg gtt gtt gac gat gag agg tgt aag aag atc atg aag
cag ttc 432Val Val Val Val Asp Asp Glu Arg Cys Lys Lys Ile Met Lys
Gln Phe 130 135 140atc gac gag agg cct cag gac tgg ttc gag gat atc
ggc gag tga taa 480Ile Asp Glu Arg Pro Gln Asp Trp Phe Glu Asp Ile
Gly Glu145 150 1556158PRTArtificial SequenceSynthetic Construct
6Met Val Thr Gly Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5
10 15Ile Ala Tyr Glu Glu Ala Ala Leu Gly Tyr Lys Glu Gly Gly Val
Pro 20 25 30Ile Gly Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu
Gly Arg 35 40 45Gly His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu
His Gly Glu 50 55 60Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly
Lys Val Tyr Lys65 70 75 80Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro
Cys Asp Met Cys Thr Gly 85 90 95Ala Ile Ile Met Tyr Gly Ile Pro Arg
Cys Val Val Gly Glu Asn Val 100 105 110Asn Phe Lys Ser Lys Gly Glu
Lys Tyr Leu Gln Thr Arg Gly His Glu 115 120 125Val Val Val Val Asp
Asp Glu Arg Cys Lys Lys Ile Met Lys Gln Phe 130 135 140Ile Asp Glu
Arg Pro Gln Asp Trp Phe Glu Asp Ile Gly Glu145 150
1557756DNASaccharomyces cerevisiaeCDS(1)..(756) 7atg aac ccg tta
ttc ttt ttg gct tct cca ttc ttg tac ctt aca tat 48Met Asn Pro Leu
Phe Phe Leu Ala Ser Pro Phe Leu Tyr Leu Thr Tyr1 5 10 15ctt ata tat
tat cca aac aaa ggg tct ttc gtt agc aaa cct aga aat 96Leu Ile Tyr
Tyr Pro Asn Lys Gly Ser Phe Val Ser Lys Pro Arg Asn 20 25 30ctg caa
aaa atg tct tcg gaa cca ttt aag aac gtc tac ttg cta cct 144Leu Gln
Lys Met Ser Ser Glu Pro Phe Lys Asn Val Tyr Leu Leu Pro 35 40 45caa
aca aac caa ttg ctg ggt ttg tac acc atc atc aga aat aag aat 192Gln
Thr Asn Gln Leu Leu Gly Leu Tyr Thr Ile Ile Arg Asn Lys Asn 50 55
60aca act aga cct gat ttc att ttc tac tcc gat aga atc atc aga ttg
240Thr Thr Arg Pro Asp Phe Ile Phe Tyr Ser Asp Arg Ile Ile Arg
Leu65 70 75 80ttg gtt gaa gaa ggt ttg aac cat cta cct gtg caa aag
caa att gtg 288Leu Val Glu Glu Gly Leu Asn His Leu Pro Val Gln Lys
Gln Ile Val 85 90 95gaa act gac acc aac gaa aac ttc gaa ggt gtc tca
ttc atg ggt aaa 336Glu Thr Asp Thr Asn Glu Asn Phe Glu Gly Val Ser
Phe Met Gly Lys 100 105 110atc tgt ggt gtt tcc att gtc aga gct ggt
gaa tcg atg gag caa gga 384Ile Cys Gly Val Ser Ile Val Arg Ala Gly
Glu Ser Met Glu Gln Gly 115 120 125tta aga gac tgt tgt agg tct gtg
cgt atc ggt aaa att tta att caa 432Leu Arg Asp Cys Cys Arg Ser Val
Arg Ile Gly Lys Ile Leu Ile Gln 130 135 140agg gac gag gag act gct
tta cca aag tta ttc tac gaa aaa tta cca 480Arg Asp Glu Glu Thr Ala
Leu Pro Lys Leu Phe Tyr Glu Lys Leu Pro145 150 155 160gag gat ata
tct gaa agg tat gtc ttc cta tta gac cca atg ctg gcc 528Glu Asp Ile
Ser Glu Arg Tyr Val Phe Leu Leu Asp Pro Met Leu Ala 165 170 175acc
ggt ggt agt gct atc atg gct aca gaa gtc ttg att aag aga ggt 576Thr
Gly Gly Ser Ala Ile Met Ala Thr Glu Val Leu Ile Lys Arg Gly 180 185
190gtt aag cca gag aga att tac ttc tta aac cta atc tgt agt aag gaa
624Val Lys Pro Glu Arg Ile Tyr Phe Leu Asn Leu Ile Cys Ser Lys Glu
195 200 205ggg att gaa aaa tac cat gcc gcc ttc cca gag gtc aga att
gtt act 672Gly Ile Glu Lys Tyr His Ala Ala Phe Pro Glu Val Arg Ile
Val Thr 210 215 220ggt gcc ctc gac aga ggt cta gat gaa aac aag tat
cta gtt cca ggg 720Gly Ala Leu Asp Arg Gly Leu Asp Glu Asn Lys Tyr
Leu Val Pro Gly225 230 235 240ttg ggt gac ttt ggt gac aga tac tac
tgt gtt taa 756Leu Gly Asp Phe Gly Asp Arg Tyr Tyr Cys Val 245
2508251PRTSaccharomyces cerevisiae 8Met Asn Pro Leu Phe Phe Leu Ala
Ser Pro Phe Leu Tyr Leu Thr Tyr1 5 10 15Leu Ile Tyr Tyr Pro Asn Lys
Gly Ser Phe Val Ser Lys Pro Arg Asn 20 25 30Leu Gln Lys Met Ser Ser
Glu Pro Phe Lys Asn Val Tyr Leu Leu Pro 35 40 45Gln Thr Asn Gln Leu
Leu Gly Leu Tyr Thr Ile Ile Arg Asn Lys Asn 50 55 60Thr Thr Arg Pro
Asp Phe Ile Phe Tyr Ser Asp Arg Ile Ile Arg Leu65 70 75 80Leu Val
Glu Glu Gly Leu Asn His Leu Pro Val Gln Lys Gln Ile Val 85 90 95Glu
Thr Asp Thr Asn Glu Asn Phe Glu Gly Val Ser Phe Met Gly Lys 100 105
110Ile Cys Gly Val Ser Ile Val Arg Ala Gly Glu Ser Met Glu Gln Gly
115 120 125Leu Arg Asp Cys Cys Arg Ser Val Arg Ile Gly Lys Ile Leu
Ile Gln 130 135 140Arg Asp Glu Glu Thr Ala Leu Pro Lys Leu Phe Tyr
Glu Lys Leu Pro145 150 155 160Glu Asp Ile Ser Glu Arg Tyr Val Phe
Leu Leu Asp Pro Met Leu Ala 165 170 175Thr Gly Gly Ser Ala Ile Met
Ala Thr Glu Val Leu Ile Lys Arg Gly 180 185 190Val Lys Pro Glu Arg
Ile Tyr Phe Leu Asn Leu Ile Cys Ser Lys Glu 195 200 205Gly Ile Glu
Lys Tyr His Ala Ala Phe Pro Glu Val Arg Ile Val Thr 210 215 220Gly
Ala Leu Asp Arg Gly Leu Asp Glu Asn Lys Tyr Leu Val Pro Gly225 230
235 240Leu Gly Asp Phe Gly Asp Arg Tyr Tyr Cys Val 245
25091443DNAhomo sapiensCDS(1)..(1443) 9atg gct gtt gct cgt gct gct
ctt ggt cct ctt gtt act ggt ctt tat 48Met Ala Val Ala Arg Ala Ala
Leu Gly Pro Leu Val Thr Gly Leu Tyr1 5 10 15gat gtt caa gct ttt aaa
ttt ggt gat ttt gtt ctt aaa tct ggt ctt 96Asp Val Gln Ala Phe Lys
Phe Gly Asp Phe Val Leu Lys Ser Gly Leu 20 25 30tct tct cct att tat
att gat ctt cgt ggt att gtt tct cgt cct cgt 144Ser Ser Pro Ile Tyr
Ile Asp Leu Arg Gly Ile Val Ser Arg Pro Arg 35 40 45ctt ctt tct caa
gtt gct gat att ctt ttt caa act gct caa aat gct 192Leu Leu Ser Gln
Val Ala Asp Ile Leu Phe Gln Thr Ala Gln Asn Ala 50 55 60ggt att tct
ttt gat act gtt tgt ggt gtt cct tat act gct ctt cct 240Gly Ile Ser
Phe Asp Thr Val Cys Gly Val Pro Tyr Thr Ala Leu Pro65 70 75 80ctt
gct act gtt att tgt tct act aat caa att cct atg ctt att cgt 288Leu
Ala Thr Val Ile Cys Ser Thr Asn Gln Ile Pro Met Leu Ile Arg 85 90
95cgt aaa gaa act aaa gat tat ggt act aaa cgt ctt gtt gaa ggt act
336Arg Lys Glu Thr Lys Asp Tyr Gly Thr Lys Arg Leu Val Glu Gly Thr
100 105 110att aat cct ggt gaa act tgt ctt att att gaa gat gtt gtt
act tct 384Ile Asn Pro Gly Glu Thr Cys Leu Ile Ile Glu Asp Val Val
Thr Ser 115 120 125ggt tct tct gtt ctt gaa act gtt gaa gtt ctt caa
aaa gaa ggt ctt 432Gly Ser Ser Val Leu Glu Thr Val Glu Val Leu Gln
Lys Glu Gly Leu 130 135 140aaa gtt act gat gct att gtt ctt ctt gat
cgt gaa caa ggt ggt aaa 480Lys Val Thr Asp Ala Ile Val Leu Leu Asp
Arg Glu Gln Gly Gly Lys145 150 155 160gat aaa ctt caa gct cat ggt
att cgt ctt cat tct gtt tgt act ctt 528Asp Lys Leu Gln Ala His Gly
Ile Arg Leu His Ser Val Cys Thr Leu 165 170 175tct aaa atg ctt gaa
att ctt gaa caa caa aaa aaa gtt gat gct gaa 576Ser Lys Met Leu Glu
Ile Leu Glu Gln Gln Lys Lys Val Asp Ala Glu 180 185 190act gtt ggt
cgt gtt aaa cgt ttt att caa gaa aat gtt ttt gtt gct 624Thr Val Gly
Arg Val Lys Arg Phe Ile Gln Glu Asn Val Phe Val Ala 195 200 205gct
aat cat aat ggt tct cct ctt tct att aaa gaa gct cct aaa gaa 672Ala
Asn His Asn Gly Ser Pro Leu Ser Ile Lys Glu Ala Pro Lys Glu 210 215
220ctt tct ttt ggt gct cgt gct gaa ctt cct cgt att cat cct gtt gct
720Leu Ser Phe Gly Ala Arg Ala Glu Leu Pro Arg Ile His Pro Val
Ala225 230 235 240tct aaa ctt ctt cgt ctt atg caa aaa aaa gaa act
aat ctt tgt ctt 768Ser Lys Leu Leu Arg Leu Met Gln Lys Lys Glu Thr
Asn Leu Cys Leu 245 250 255tct gct gat gtt tct ctt gct cgt gaa ctt
ctt caa ctt gct gat gct 816Ser Ala Asp Val Ser Leu Ala Arg Glu Leu
Leu Gln Leu Ala Asp Ala 260 265 270ctt ggt cct tct att tgt atg ctt
aaa act cat gtt gat att ctt aat 864Leu Gly Pro Ser Ile Cys Met Leu
Lys Thr His Val Asp Ile Leu Asn 275 280 285gat ttt act ctt gat gtt
atg aaa gaa ctt att act ctt gct aaa tgt 912Asp Phe Thr Leu Asp Val
Met Lys Glu Leu Ile Thr Leu Ala Lys Cys 290 295 300cat gaa ttt ctt
att ttt gaa gat cgt aaa ttt gct gat att ggt aat 960His Glu Phe Leu
Ile Phe Glu Asp Arg Lys Phe Ala Asp Ile Gly Asn305 310 315 320act
gtt aaa aaa caa tat gaa ggt ggt att ttt aaa att gct tct tgg 1008Thr
Val Lys Lys
Gln Tyr Glu Gly Gly Ile Phe Lys Ile Ala Ser Trp 325 330 335gct gat
ctt gtt aat gct cat gtt gtt cct ggt tct ggt gtt gtt aaa 1056Ala Asp
Leu Val Asn Ala His Val Val Pro Gly Ser Gly Val Val Lys 340 345
350ggt ctt caa gaa gtt ggt ctt cct ctt cat cgt ggt tgt ctt ctt att
1104Gly Leu Gln Glu Val Gly Leu Pro Leu His Arg Gly Cys Leu Leu Ile
355 360 365gct gaa atg tct tct act ggt tct ctt gct act ggt gat tat
act cgt 1152Ala Glu Met Ser Ser Thr Gly Ser Leu Ala Thr Gly Asp Tyr
Thr Arg 370 375 380gct gct gtt cgt atg gct gaa gaa cat tct gaa ttt
gtt gtt ggt ttt 1200Ala Ala Val Arg Met Ala Glu Glu His Ser Glu Phe
Val Val Gly Phe385 390 395 400att tct ggt tct cgt gtt tct atg aaa
cct gaa ttt ctt cat ctt act 1248Ile Ser Gly Ser Arg Val Ser Met Lys
Pro Glu Phe Leu His Leu Thr 405 410 415cct ggt gtt caa ctt gaa gct
ggt ggt gat aat ctt ggt caa caa tat 1296Pro Gly Val Gln Leu Glu Ala
Gly Gly Asp Asn Leu Gly Gln Gln Tyr 420 425 430aat tct cct caa gaa
gtt att ggt aaa cgt ggt tct gat att att att 1344Asn Ser Pro Gln Glu
Val Ile Gly Lys Arg Gly Ser Asp Ile Ile Ile 435 440 445gtt ggt cgt
ggt att att tct gct gct gat cgt ctt gaa gct gct gaa 1392Val Gly Arg
Gly Ile Ile Ser Ala Ala Asp Arg Leu Glu Ala Ala Glu 450 455 460atg
tat cgt aaa gct gct tgg gaa gct tat ctt tct cgt ctt ggt gtt 1440Met
Tyr Arg Lys Ala Ala Trp Glu Ala Tyr Leu Ser Arg Leu Gly Val465 470
475 480taa 144310480PRThomo sapiens 10Met Ala Val Ala Arg Ala Ala
Leu Gly Pro Leu Val Thr Gly Leu Tyr1 5 10 15Asp Val Gln Ala Phe Lys
Phe Gly Asp Phe Val Leu Lys Ser Gly Leu 20 25 30Ser Ser Pro Ile Tyr
Ile Asp Leu Arg Gly Ile Val Ser Arg Pro Arg 35 40 45Leu Leu Ser Gln
Val Ala Asp Ile Leu Phe Gln Thr Ala Gln Asn Ala 50 55 60Gly Ile Ser
Phe Asp Thr Val Cys Gly Val Pro Tyr Thr Ala Leu Pro65 70 75 80Leu
Ala Thr Val Ile Cys Ser Thr Asn Gln Ile Pro Met Leu Ile Arg 85 90
95Arg Lys Glu Thr Lys Asp Tyr Gly Thr Lys Arg Leu Val Glu Gly Thr
100 105 110Ile Asn Pro Gly Glu Thr Cys Leu Ile Ile Glu Asp Val Val
Thr Ser 115 120 125Gly Ser Ser Val Leu Glu Thr Val Glu Val Leu Gln
Lys Glu Gly Leu 130 135 140Lys Val Thr Asp Ala Ile Val Leu Leu Asp
Arg Glu Gln Gly Gly Lys145 150 155 160Asp Lys Leu Gln Ala His Gly
Ile Arg Leu His Ser Val Cys Thr Leu 165 170 175Ser Lys Met Leu Glu
Ile Leu Glu Gln Gln Lys Lys Val Asp Ala Glu 180 185 190Thr Val Gly
Arg Val Lys Arg Phe Ile Gln Glu Asn Val Phe Val Ala 195 200 205Ala
Asn His Asn Gly Ser Pro Leu Ser Ile Lys Glu Ala Pro Lys Glu 210 215
220Leu Ser Phe Gly Ala Arg Ala Glu Leu Pro Arg Ile His Pro Val
Ala225 230 235 240Ser Lys Leu Leu Arg Leu Met Gln Lys Lys Glu Thr
Asn Leu Cys Leu 245 250 255Ser Ala Asp Val Ser Leu Ala Arg Glu Leu
Leu Gln Leu Ala Asp Ala 260 265 270Leu Gly Pro Ser Ile Cys Met Leu
Lys Thr His Val Asp Ile Leu Asn 275 280 285Asp Phe Thr Leu Asp Val
Met Lys Glu Leu Ile Thr Leu Ala Lys Cys 290 295 300His Glu Phe Leu
Ile Phe Glu Asp Arg Lys Phe Ala Asp Ile Gly Asn305 310 315 320Thr
Val Lys Lys Gln Tyr Glu Gly Gly Ile Phe Lys Ile Ala Ser Trp 325 330
335Ala Asp Leu Val Asn Ala His Val Val Pro Gly Ser Gly Val Val Lys
340 345 350Gly Leu Gln Glu Val Gly Leu Pro Leu His Arg Gly Cys Leu
Leu Ile 355 360 365Ala Glu Met Ser Ser Thr Gly Ser Leu Ala Thr Gly
Asp Tyr Thr Arg 370 375 380Ala Ala Val Arg Met Ala Glu Glu His Ser
Glu Phe Val Val Gly Phe385 390 395 400Ile Ser Gly Ser Arg Val Ser
Met Lys Pro Glu Phe Leu His Leu Thr 405 410 415Pro Gly Val Gln Leu
Glu Ala Gly Gly Asp Asn Leu Gly Gln Gln Tyr 420 425 430Asn Ser Pro
Gln Glu Val Ile Gly Lys Arg Gly Ser Asp Ile Ile Ile 435 440 445Val
Gly Arg Gly Ile Ile Ser Ala Ala Asp Arg Leu Glu Ala Ala Glu 450 455
460Met Tyr Arg Lys Ala Ala Trp Glu Ala Tyr Leu Ser Arg Leu Gly
Val465 470 475 480111227DNAArtificial SequenceFusion construct
CDopt-UPRTCDS(1)..(1227) 11atg gtg acc ggc ggc atg gcc tcc aag tgg
gat caa aag ggc atg gat 48Met Val Thr Gly Gly Met Ala Ser Lys Trp
Asp Gln Lys Gly Met Asp1 5 10 15atc gct tac gag gag gcc ctg ctg ggc
tac aag gag ggc ggc gtg cct 96Ile Ala Tyr Glu Glu Ala Leu Leu Gly
Tyr Lys Glu Gly Gly Val Pro 20 25 30atc ggc ggc tgt ctg atc aac aac
aag gac ggc agt gtg ctg ggc agg 144Ile Gly Gly Cys Leu Ile Asn Asn
Lys Asp Gly Ser Val Leu Gly Arg 35 40 45ggc cac aac atg agg ttc cag
aag ggc tcc gcc acc ctg cac ggc gag 192Gly His Asn Met Arg Phe Gln
Lys Gly Ser Ala Thr Leu His Gly Glu 50 55 60atc tcc acc ctg gag aac
tgt ggc agg ctg gag ggc aag gtg tac aag 240Ile Ser Thr Leu Glu Asn
Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65 70 75 80gac acc acc ctg
tac acc acc ctg tcc cct tgt gac atg tgt acc ggc 288Asp Thr Thr Leu
Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr Gly 85 90 95gct atc atc
atg tac ggc atc cct agg tgt gtg atc ggc gag aac gtg 336Ala Ile Ile
Met Tyr Gly Ile Pro Arg Cys Val Ile Gly Glu Asn Val 100 105 110aac
ttc aag tcc aag ggc gag aag tac ctg caa acc agg ggc cac gag 384Asn
Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr Arg Gly His Glu 115 120
125gtg gtg gtt gtt gac gat gag agg tgt aag aag ctg atg aag cag ttc
432Val Val Val Val Asp Asp Glu Arg Cys Lys Lys Leu Met Lys Gln Phe
130 135 140atc gac gag agg cct cag gac tgg ttc gag gat atc ggc gag
aac ccg 480Ile Asp Glu Arg Pro Gln Asp Trp Phe Glu Asp Ile Gly Glu
Asn Pro145 150 155 160tta ttc ttt ttg gct tct cca ttc ttg tac ctt
aca tat ctt ata tat 528Leu Phe Phe Leu Ala Ser Pro Phe Leu Tyr Leu
Thr Tyr Leu Ile Tyr 165 170 175tat cca aac aaa ggg tct ttc gtt agc
aaa cct aga aat ctg caa aaa 576Tyr Pro Asn Lys Gly Ser Phe Val Ser
Lys Pro Arg Asn Leu Gln Lys 180 185 190atg tct tcg gaa cca ttt aag
aac gtc tac ttg cta cct caa aca aac 624Met Ser Ser Glu Pro Phe Lys
Asn Val Tyr Leu Leu Pro Gln Thr Asn 195 200 205caa ttg ctg ggt ttg
tac acc atc atc aga aat aag aat aca act aga 672Gln Leu Leu Gly Leu
Tyr Thr Ile Ile Arg Asn Lys Asn Thr Thr Arg 210 215 220cct gat ttc
att ttc tac tcc gat aga atc atc aga ttg ttg gtt gaa 720Pro Asp Phe
Ile Phe Tyr Ser Asp Arg Ile Ile Arg Leu Leu Val Glu225 230 235
240gaa ggt ttg aac cat cta cct gtg caa aag caa att gtg gaa act gac
768Glu Gly Leu Asn His Leu Pro Val Gln Lys Gln Ile Val Glu Thr Asp
245 250 255acc aac gaa aac ttc gaa ggt gtc tca ttc atg ggt aaa atc
tgt ggt 816Thr Asn Glu Asn Phe Glu Gly Val Ser Phe Met Gly Lys Ile
Cys Gly 260 265 270gtt tcc att gtc aga gct ggt gaa tcg atg gag caa
gga tta aga gac 864Val Ser Ile Val Arg Ala Gly Glu Ser Met Glu Gln
Gly Leu Arg Asp 275 280 285tgt tgt agg tct gtg cgt atc ggt aaa att
tta att caa agg gac gag 912Cys Cys Arg Ser Val Arg Ile Gly Lys Ile
Leu Ile Gln Arg Asp Glu 290 295 300gag act gct tta cca aag tta ttc
tac gaa aaa tta cca gag gat ata 960Glu Thr Ala Leu Pro Lys Leu Phe
Tyr Glu Lys Leu Pro Glu Asp Ile305 310 315 320tct gaa agg tat gtc
ttc cta tta gac cca atg ctg gcc acc ggt ggt 1008Ser Glu Arg Tyr Val
Phe Leu Leu Asp Pro Met Leu Ala Thr Gly Gly 325 330 335agt gct atc
atg gct aca gaa gtc ttg att aag aga ggt gtt aag cca 1056Ser Ala Ile
Met Ala Thr Glu Val Leu Ile Lys Arg Gly Val Lys Pro 340 345 350gag
aga att tac ttc tta aac cta atc tgt agt aag gaa ggg att gaa 1104Glu
Arg Ile Tyr Phe Leu Asn Leu Ile Cys Ser Lys Glu Gly Ile Glu 355 360
365aaa tac cat gcc gcc ttc cca gag gtc aga att gtt act ggt gcc ctc
1152Lys Tyr His Ala Ala Phe Pro Glu Val Arg Ile Val Thr Gly Ala Leu
370 375 380gac aga ggt cta gat gaa aac aag tat cta gtt cca ggg ttg
ggt gac 1200Asp Arg Gly Leu Asp Glu Asn Lys Tyr Leu Val Pro Gly Leu
Gly Asp385 390 395 400ttt ggt gac aga tac tac tgt gtt taa 1227Phe
Gly Asp Arg Tyr Tyr Cys Val 40512408PRTArtificial SequenceSynthetic
Construct 12Met Val Thr Gly Gly Met Ala Ser Lys Trp Asp Gln Lys Gly
Met Asp1 5 10 15Ile Ala Tyr Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly
Gly Val Pro 20 25 30Ile Gly Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser
Val Leu Gly Arg 35 40 45Gly His Asn Met Arg Phe Gln Lys Gly Ser Ala
Thr Leu His Gly Glu 50 55 60Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu
Glu Gly Lys Val Tyr Lys65 70 75 80Asp Thr Thr Leu Tyr Thr Thr Leu
Ser Pro Cys Asp Met Cys Thr Gly 85 90 95Ala Ile Ile Met Tyr Gly Ile
Pro Arg Cys Val Ile Gly Glu Asn Val 100 105 110Asn Phe Lys Ser Lys
Gly Glu Lys Tyr Leu Gln Thr Arg Gly His Glu 115 120 125Val Val Val
Val Asp Asp Glu Arg Cys Lys Lys Leu Met Lys Gln Phe 130 135 140Ile
Asp Glu Arg Pro Gln Asp Trp Phe Glu Asp Ile Gly Glu Asn Pro145 150
155 160Leu Phe Phe Leu Ala Ser Pro Phe Leu Tyr Leu Thr Tyr Leu Ile
Tyr 165 170 175Tyr Pro Asn Lys Gly Ser Phe Val Ser Lys Pro Arg Asn
Leu Gln Lys 180 185 190Met Ser Ser Glu Pro Phe Lys Asn Val Tyr Leu
Leu Pro Gln Thr Asn 195 200 205Gln Leu Leu Gly Leu Tyr Thr Ile Ile
Arg Asn Lys Asn Thr Thr Arg 210 215 220Pro Asp Phe Ile Phe Tyr Ser
Asp Arg Ile Ile Arg Leu Leu Val Glu225 230 235 240Glu Gly Leu Asn
His Leu Pro Val Gln Lys Gln Ile Val Glu Thr Asp 245 250 255Thr Asn
Glu Asn Phe Glu Gly Val Ser Phe Met Gly Lys Ile Cys Gly 260 265
270Val Ser Ile Val Arg Ala Gly Glu Ser Met Glu Gln Gly Leu Arg Asp
275 280 285Cys Cys Arg Ser Val Arg Ile Gly Lys Ile Leu Ile Gln Arg
Asp Glu 290 295 300Glu Thr Ala Leu Pro Lys Leu Phe Tyr Glu Lys Leu
Pro Glu Asp Ile305 310 315 320Ser Glu Arg Tyr Val Phe Leu Leu Asp
Pro Met Leu Ala Thr Gly Gly 325 330 335Ser Ala Ile Met Ala Thr Glu
Val Leu Ile Lys Arg Gly Val Lys Pro 340 345 350Glu Arg Ile Tyr Phe
Leu Asn Leu Ile Cys Ser Lys Glu Gly Ile Glu 355 360 365Lys Tyr His
Ala Ala Phe Pro Glu Val Arg Ile Val Thr Gly Ala Leu 370 375 380Asp
Arg Gly Leu Asp Glu Asn Lys Tyr Leu Val Pro Gly Leu Gly Asp385 390
395 400Phe Gly Asp Arg Tyr Tyr Cys Val 405131287DNAArtificial
SequenceFusion construction - CDopt - linker - UPRTCDS(1)..(1287)
13atg gtg acc ggc ggc atg gcc tcc aag tgg gat caa aag ggc atg gat
48Met Val Thr Gly Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1
5 10 15atc gct tac gag gag gcc ctg ctg ggc tac aag gag ggc ggc gtg
cct 96Ile Ala Tyr Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly Gly Val
Pro 20 25 30atc ggc ggc tgt ctg atc aac aac aag gac ggc agt gtg ctg
ggc agg 144Ile Gly Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu
Gly Arg 35 40 45ggc cac aac atg agg ttc cag aag ggc tcc gcc acc ctg
cac ggc gag 192Gly His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu
His Gly Glu 50 55 60atc tcc acc ctg gag aac tgt ggc agg ctg gag ggc
aag gtg tac aag 240Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly
Lys Val Tyr Lys65 70 75 80gac acc acc ctg tac acc acc ctg tcc cct
tgt gac atg tgt acc ggc 288Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro
Cys Asp Met Cys Thr Gly 85 90 95gct atc atc atg tac ggc atc cct agg
tgt gtg atc ggc gag aac gtg 336Ala Ile Ile Met Tyr Gly Ile Pro Arg
Cys Val Ile Gly Glu Asn Val 100 105 110aac ttc aag tcc aag ggc gag
aag tac ctg caa acc agg ggc cac gag 384Asn Phe Lys Ser Lys Gly Glu
Lys Tyr Leu Gln Thr Arg Gly His Glu 115 120 125gtg gtg gtt gtt gac
gat gag agg tgt aag aag ctg atg aag cag ttc 432Val Val Val Val Asp
Asp Glu Arg Cys Lys Lys Leu Met Lys Gln Phe 130 135 140atc gac gag
agg cct cag gac tgg ttc gag gat atc ggc gag tcc ggc 480Ile Asp Glu
Arg Pro Gln Asp Trp Phe Glu Asp Ile Gly Glu Ser Gly145 150 155
160ggc ggc gcc tcc ggc ggc ggc gcc tcc ggc ggc ggc gcc tcc ggc ggc
528Gly Gly Ala Ser Gly Gly Gly Ala Ser Gly Gly Gly Ala Ser Gly Gly
165 170 175ggc gcc aac ccg tta ttc ttt ttg gct tct cca ttc ttg tac
ctt aca 576Gly Ala Asn Pro Leu Phe Phe Leu Ala Ser Pro Phe Leu Tyr
Leu Thr 180 185 190tat ctt ata tat tat cca aac aaa ggg tct ttc gtt
agc aaa cct aga 624Tyr Leu Ile Tyr Tyr Pro Asn Lys Gly Ser Phe Val
Ser Lys Pro Arg 195 200 205aat ctg caa aaa atg tct tcg gaa cca ttt
aag aac gtc tac ttg cta 672Asn Leu Gln Lys Met Ser Ser Glu Pro Phe
Lys Asn Val Tyr Leu Leu 210 215 220cct caa aca aac caa ttg ctg ggt
ttg tac acc atc atc aga aat aag 720Pro Gln Thr Asn Gln Leu Leu Gly
Leu Tyr Thr Ile Ile Arg Asn Lys225 230 235 240aat aca act aga cct
gat ttc att ttc tac tcc gat aga atc atc aga 768Asn Thr Thr Arg Pro
Asp Phe Ile Phe Tyr Ser Asp Arg Ile Ile Arg 245 250 255ttg ttg gtt
gaa gaa ggt ttg aac cat cta cct gtg caa aag caa att 816Leu Leu Val
Glu Glu Gly Leu Asn His Leu Pro Val Gln Lys Gln Ile 260 265 270gtg
gaa act gac acc aac gaa aac ttc gaa ggt gtc tca ttc atg ggt 864Val
Glu Thr Asp Thr Asn Glu Asn Phe Glu Gly Val Ser Phe Met Gly 275 280
285aaa atc tgt ggt gtt tcc att gtc aga gct ggt gaa tcg atg gag caa
912Lys Ile Cys Gly Val Ser Ile Val Arg Ala Gly Glu Ser Met Glu Gln
290 295 300gga tta aga gac tgt tgt agg tct gtg cgt atc ggt aaa att
tta att 960Gly Leu Arg Asp Cys Cys Arg Ser Val Arg Ile Gly Lys Ile
Leu Ile305 310 315 320caa agg gac gag gag act gct tta cca aag tta
ttc tac gaa aaa tta 1008Gln Arg Asp Glu Glu Thr Ala Leu Pro Lys Leu
Phe Tyr Glu Lys Leu 325 330 335cca gag gat ata tct gaa agg tat gtc
ttc cta tta gac cca atg ctg 1056Pro Glu Asp Ile Ser Glu Arg Tyr Val
Phe Leu Leu Asp Pro Met Leu 340 345 350gcc acc ggt ggt agt gct atc
atg gct aca gaa gtc ttg att aag aga 1104Ala Thr Gly Gly Ser Ala Ile
Met Ala Thr Glu Val Leu Ile Lys Arg 355 360 365ggt gtt aag cca gag
aga att tac ttc tta aac cta atc tgt agt aag 1152Gly Val Lys Pro Glu
Arg Ile Tyr Phe Leu Asn Leu Ile Cys Ser Lys 370 375 380gaa ggg att
gaa aaa tac cat gcc gcc ttc cca gag gtc aga att gtt 1200Glu Gly Ile
Glu Lys Tyr His Ala Ala Phe Pro Glu Val Arg Ile Val385 390
395 400act ggt gcc ctc gac aga ggt cta gat gaa aac aag tat cta gtt
cca 1248Thr Gly Ala Leu Asp Arg Gly Leu Asp Glu Asn Lys Tyr Leu Val
Pro 405 410 415ggg ttg ggt gac ttt ggt gac aga tac tac tgt gtt taa
1287Gly Leu Gly Asp Phe Gly Asp Arg Tyr Tyr Cys Val 420
42514428PRTArtificial SequenceSynthetic Construct 14Met Val Thr Gly
Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15Ile Ala Tyr
Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30Ile Gly
Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45Gly
His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55
60Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65
70 75 80Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr
Gly 85 90 95Ala Ile Ile Met Tyr Gly Ile Pro Arg Cys Val Ile Gly Glu
Asn Val 100 105 110Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr
Arg Gly His Glu 115 120 125Val Val Val Val Asp Asp Glu Arg Cys Lys
Lys Leu Met Lys Gln Phe 130 135 140Ile Asp Glu Arg Pro Gln Asp Trp
Phe Glu Asp Ile Gly Glu Ser Gly145 150 155 160Gly Gly Ala Ser Gly
Gly Gly Ala Ser Gly Gly Gly Ala Ser Gly Gly 165 170 175Gly Ala Asn
Pro Leu Phe Phe Leu Ala Ser Pro Phe Leu Tyr Leu Thr 180 185 190Tyr
Leu Ile Tyr Tyr Pro Asn Lys Gly Ser Phe Val Ser Lys Pro Arg 195 200
205Asn Leu Gln Lys Met Ser Ser Glu Pro Phe Lys Asn Val Tyr Leu Leu
210 215 220Pro Gln Thr Asn Gln Leu Leu Gly Leu Tyr Thr Ile Ile Arg
Asn Lys225 230 235 240Asn Thr Thr Arg Pro Asp Phe Ile Phe Tyr Ser
Asp Arg Ile Ile Arg 245 250 255Leu Leu Val Glu Glu Gly Leu Asn His
Leu Pro Val Gln Lys Gln Ile 260 265 270Val Glu Thr Asp Thr Asn Glu
Asn Phe Glu Gly Val Ser Phe Met Gly 275 280 285Lys Ile Cys Gly Val
Ser Ile Val Arg Ala Gly Glu Ser Met Glu Gln 290 295 300Gly Leu Arg
Asp Cys Cys Arg Ser Val Arg Ile Gly Lys Ile Leu Ile305 310 315
320Gln Arg Asp Glu Glu Thr Ala Leu Pro Lys Leu Phe Tyr Glu Lys Leu
325 330 335Pro Glu Asp Ile Ser Glu Arg Tyr Val Phe Leu Leu Asp Pro
Met Leu 340 345 350Ala Thr Gly Gly Ser Ala Ile Met Ala Thr Glu Val
Leu Ile Lys Arg 355 360 365Gly Val Lys Pro Glu Arg Ile Tyr Phe Leu
Asn Leu Ile Cys Ser Lys 370 375 380Glu Gly Ile Glu Lys Tyr His Ala
Ala Phe Pro Glu Val Arg Ile Val385 390 395 400Thr Gly Ala Leu Asp
Arg Gly Leu Asp Glu Asn Lys Tyr Leu Val Pro 405 410 415Gly Leu Gly
Asp Phe Gly Asp Arg Tyr Tyr Cys Val 420 425151200DNAArtificial
SequenceFusion Construct - CDopt3 - OPRTCDS(1)..(1200) 15atg gtg
acc ggc ggc atg gcc tcc aag tgg gat caa aag ggc atg gat 48Met Val
Thr Gly Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15atc
gct tac gag gag gcc ctg ctg ggc tac aag gag ggc ggc gtg cct 96Ile
Ala Tyr Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25
30atc ggc ggc tgt ctg atc aac aac aag gac ggc agt gtg ctg ggc agg
144Ile Gly Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu Gly Arg
35 40 45ggc cac aac atg agg ttc cag aag ggc tcc gcc acc ctg cac ggc
gag 192Gly His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu His Gly
Glu 50 55 60atc tcc acc ctg gag aac tgt ggc agg ctg gag ggc aag gtg
tac aag 240Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly Lys Val
Tyr Lys65 70 75 80gac acc acc ctg tac acc acc ctg tcc cct tgt gac
atg tgt acc ggc 288Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp
Met Cys Thr Gly 85 90 95gct atc atc atg tac ggc atc cct agg tgt gtg
atc ggc gag aac gtg 336Ala Ile Ile Met Tyr Gly Ile Pro Arg Cys Val
Ile Gly Glu Asn Val 100 105 110aac ttc aag tcc aag ggc gag aag tac
ctg caa acc agg ggc cac gag 384Asn Phe Lys Ser Lys Gly Glu Lys Tyr
Leu Gln Thr Arg Gly His Glu 115 120 125gtg gtg gtt gtt gac gat gag
agg tgt aag aag ctg atg aag cag ttc 432Val Val Val Val Asp Asp Glu
Arg Cys Lys Lys Leu Met Lys Gln Phe 130 135 140atc gac gag agg cct
cag gac tgg ttc gag gat atc ggc gag gcg gtc 480Ile Asp Glu Arg Pro
Gln Asp Trp Phe Glu Asp Ile Gly Glu Ala Val145 150 155 160gct cgt
gca gct ttg ggg cca ttg gtg acg ggt ctg tac gac gtg cag 528Ala Arg
Ala Ala Leu Gly Pro Leu Val Thr Gly Leu Tyr Asp Val Gln 165 170
175gct ttc aag ttt ggg gac ttc gtg ctg aag agc ggg ctt tcc tcc ccc
576Ala Phe Lys Phe Gly Asp Phe Val Leu Lys Ser Gly Leu Ser Ser Pro
180 185 190atc tac atc gat ctg cgg ggc atc gtg tct cga ccg cgt ctt
ctg agt 624Ile Tyr Ile Asp Leu Arg Gly Ile Val Ser Arg Pro Arg Leu
Leu Ser 195 200 205cag gtt gca gat att tta ttc caa act gcc caa aat
gca ggc atc agt 672Gln Val Ala Asp Ile Leu Phe Gln Thr Ala Gln Asn
Ala Gly Ile Ser 210 215 220ttt gac acc gtg tgt gga gtg cct tat aca
gct ttg cca ttg gct aca 720Phe Asp Thr Val Cys Gly Val Pro Tyr Thr
Ala Leu Pro Leu Ala Thr225 230 235 240gtt atc tgt tca acc aat caa
att cca atg ctt att aga agg aaa gaa 768Val Ile Cys Ser Thr Asn Gln
Ile Pro Met Leu Ile Arg Arg Lys Glu 245 250 255aca aag gat tat gga
act aag cgt ctt gta gaa gga act att aat cca 816Thr Lys Asp Tyr Gly
Thr Lys Arg Leu Val Glu Gly Thr Ile Asn Pro 260 265 270gga gaa acc
tgt tta atc att gaa gat gtt gtc acc agt gga tct agt 864Gly Glu Thr
Cys Leu Ile Ile Glu Asp Val Val Thr Ser Gly Ser Ser 275 280 285gtt
ttg gaa act gtt gag gtt ctt cag aag gag ggc ttg aag gtc act 912Val
Leu Glu Thr Val Glu Val Leu Gln Lys Glu Gly Leu Lys Val Thr 290 295
300gat gcc ata gtg ctg ttg gac aga gag cag gga ggc aag gac aag ttg
960Asp Ala Ile Val Leu Leu Asp Arg Glu Gln Gly Gly Lys Asp Lys
Leu305 310 315 320cag gcg cac ggg atc cgc ctc cac tca gtg tgt aca
ttg tcc aaa atg 1008Gln Ala His Gly Ile Arg Leu His Ser Val Cys Thr
Leu Ser Lys Met 325 330 335ctg gag att ctc gag cag cag aaa aaa gtt
gat gct gag aca gtt ggg 1056Leu Glu Ile Leu Glu Gln Gln Lys Lys Val
Asp Ala Glu Thr Val Gly 340 345 350aga gtg aag agg ttt att cag gag
aat gtc ttt gtg gca gcg aat cat 1104Arg Val Lys Arg Phe Ile Gln Glu
Asn Val Phe Val Ala Ala Asn His 355 360 365aat ggt tct ccc ctt tct
ata aag gaa gca ccc aaa gaa ctc agc ttc 1152Asn Gly Ser Pro Leu Ser
Ile Lys Glu Ala Pro Lys Glu Leu Ser Phe 370 375 380ggt gca cgt gca
gag ctg ccc agg atc cac cca gtt gca tcg aag taa 1200Gly Ala Arg Ala
Glu Leu Pro Arg Ile His Pro Val Ala Ser Lys385 390
39516399PRTArtificial SequenceSynthetic Construct 16Met Val Thr Gly
Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1 5 10 15Ile Ala Tyr
Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30Ile Gly
Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45Gly
His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55
60Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65
70 75 80Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr
Gly 85 90 95Ala Ile Ile Met Tyr Gly Ile Pro Arg Cys Val Ile Gly Glu
Asn Val 100 105 110Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr
Arg Gly His Glu 115 120 125Val Val Val Val Asp Asp Glu Arg Cys Lys
Lys Leu Met Lys Gln Phe 130 135 140Ile Asp Glu Arg Pro Gln Asp Trp
Phe Glu Asp Ile Gly Glu Ala Val145 150 155 160Ala Arg Ala Ala Leu
Gly Pro Leu Val Thr Gly Leu Tyr Asp Val Gln 165 170 175Ala Phe Lys
Phe Gly Asp Phe Val Leu Lys Ser Gly Leu Ser Ser Pro 180 185 190Ile
Tyr Ile Asp Leu Arg Gly Ile Val Ser Arg Pro Arg Leu Leu Ser 195 200
205Gln Val Ala Asp Ile Leu Phe Gln Thr Ala Gln Asn Ala Gly Ile Ser
210 215 220Phe Asp Thr Val Cys Gly Val Pro Tyr Thr Ala Leu Pro Leu
Ala Thr225 230 235 240Val Ile Cys Ser Thr Asn Gln Ile Pro Met Leu
Ile Arg Arg Lys Glu 245 250 255Thr Lys Asp Tyr Gly Thr Lys Arg Leu
Val Glu Gly Thr Ile Asn Pro 260 265 270Gly Glu Thr Cys Leu Ile Ile
Glu Asp Val Val Thr Ser Gly Ser Ser 275 280 285Val Leu Glu Thr Val
Glu Val Leu Gln Lys Glu Gly Leu Lys Val Thr 290 295 300Asp Ala Ile
Val Leu Leu Asp Arg Glu Gln Gly Gly Lys Asp Lys Leu305 310 315
320Gln Ala His Gly Ile Arg Leu His Ser Val Cys Thr Leu Ser Lys Met
325 330 335Leu Glu Ile Leu Glu Gln Gln Lys Lys Val Asp Ala Glu Thr
Val Gly 340 345 350Arg Val Lys Arg Phe Ile Gln Glu Asn Val Phe Val
Ala Ala Asn His 355 360 365Asn Gly Ser Pro Leu Ser Ile Lys Glu Ala
Pro Lys Glu Leu Ser Phe 370 375 380Gly Ala Arg Ala Glu Leu Pro Arg
Ile His Pro Val Ala Ser Lys385 390 395171260DNAArtificial
SequenceFusion Construct - CDopt3 - linker - OPRTCDS(1)..(1260)
17atg gtg acc ggc ggc atg gcc tcc aag tgg gat caa aag ggc atg gat
48Met Val Thr Gly Gly Met Ala Ser Lys Trp Asp Gln Lys Gly Met Asp1
5 10 15atc gct tac gag gag gcc ctg ctg ggc tac aag gag ggc ggc gtg
cct 96Ile Ala Tyr Glu Glu Ala Leu Leu Gly Tyr Lys Glu Gly Gly Val
Pro 20 25 30atc ggc ggc tgt ctg atc aac aac aag gac ggc agt gtg ctg
ggc agg 144Ile Gly Gly Cys Leu Ile Asn Asn Lys Asp Gly Ser Val Leu
Gly Arg 35 40 45ggc cac aac atg agg ttc cag aag ggc tcc gcc acc ctg
cac ggc gag 192Gly His Asn Met Arg Phe Gln Lys Gly Ser Ala Thr Leu
His Gly Glu 50 55 60atc tcc acc ctg gag aac tgt ggc agg ctg gag ggc
aag gtg tac aag 240Ile Ser Thr Leu Glu Asn Cys Gly Arg Leu Glu Gly
Lys Val Tyr Lys65 70 75 80gac acc acc ctg tac acc acc ctg tcc cct
tgt gac atg tgt acc ggc 288Asp Thr Thr Leu Tyr Thr Thr Leu Ser Pro
Cys Asp Met Cys Thr Gly 85 90 95gct atc atc atg tac ggc atc cct agg
tgt gtg atc ggc gag aac gtg 336Ala Ile Ile Met Tyr Gly Ile Pro Arg
Cys Val Ile Gly Glu Asn Val 100 105 110aac ttc aag tcc aag ggc gag
aag tac ctg caa acc agg ggc cac gag 384Asn Phe Lys Ser Lys Gly Glu
Lys Tyr Leu Gln Thr Arg Gly His Glu 115 120 125gtg gtg gtt gtt gac
gat gag agg tgt aag aag ctg atg aag cag ttc 432Val Val Val Val Asp
Asp Glu Arg Cys Lys Lys Leu Met Lys Gln Phe 130 135 140atc gac gag
agg cct cag gac tgg ttc gag gat atc ggc gag tcc ggc 480Ile Asp Glu
Arg Pro Gln Asp Trp Phe Glu Asp Ile Gly Glu Ser Gly145 150 155
160ggc ggc gcc tcc ggc ggc ggc gcc tcc ggc ggc ggc gcc tcc ggc ggc
528Gly Gly Ala Ser Gly Gly Gly Ala Ser Gly Gly Gly Ala Ser Gly Gly
165 170 175ggc gcc gcg gtc gct cgt gca gct ttg ggg cca ttg gtg acg
ggt ctg 576Gly Ala Ala Val Ala Arg Ala Ala Leu Gly Pro Leu Val Thr
Gly Leu 180 185 190tac gac gtg cag gct ttc aag ttt ggg gac ttc gtg
ctg aag agc ggg 624Tyr Asp Val Gln Ala Phe Lys Phe Gly Asp Phe Val
Leu Lys Ser Gly 195 200 205ctt tcc tcc ccc atc tac atc gat ctg cgg
ggc atc gtg tct cga ccg 672Leu Ser Ser Pro Ile Tyr Ile Asp Leu Arg
Gly Ile Val Ser Arg Pro 210 215 220cgt ctt ctg agt cag gtt gca gat
att tta ttc caa act gcc caa aat 720Arg Leu Leu Ser Gln Val Ala Asp
Ile Leu Phe Gln Thr Ala Gln Asn225 230 235 240gca ggc atc agt ttt
gac acc gtg tgt gga gtg cct tat aca gct ttg 768Ala Gly Ile Ser Phe
Asp Thr Val Cys Gly Val Pro Tyr Thr Ala Leu 245 250 255cca ttg gct
aca gtt atc tgt tca acc aat caa att cca atg ctt att 816Pro Leu Ala
Thr Val Ile Cys Ser Thr Asn Gln Ile Pro Met Leu Ile 260 265 270aga
agg aaa gaa aca aag gat tat gga act aag cgt ctt gta gaa gga 864Arg
Arg Lys Glu Thr Lys Asp Tyr Gly Thr Lys Arg Leu Val Glu Gly 275 280
285act att aat cca gga gaa acc tgt tta atc att gaa gat gtt gtc acc
912Thr Ile Asn Pro Gly Glu Thr Cys Leu Ile Ile Glu Asp Val Val Thr
290 295 300agt gga tct agt gtt ttg gaa act gtt gag gtt ctt cag aag
gag ggc 960Ser Gly Ser Ser Val Leu Glu Thr Val Glu Val Leu Gln Lys
Glu Gly305 310 315 320ttg aag gtc act gat gcc ata gtg ctg ttg gac
aga gag cag gga ggc 1008Leu Lys Val Thr Asp Ala Ile Val Leu Leu Asp
Arg Glu Gln Gly Gly 325 330 335aag gac aag ttg cag gcg cac ggg atc
cgc ctc cac tca gtg tgt aca 1056Lys Asp Lys Leu Gln Ala His Gly Ile
Arg Leu His Ser Val Cys Thr 340 345 350ttg tcc aaa atg ctg gag att
ctc gag cag cag aaa aaa gtt gat gct 1104Leu Ser Lys Met Leu Glu Ile
Leu Glu Gln Gln Lys Lys Val Asp Ala 355 360 365gag aca gtt ggg aga
gtg aag agg ttt att cag gag aat gtc ttt gtg 1152Glu Thr Val Gly Arg
Val Lys Arg Phe Ile Gln Glu Asn Val Phe Val 370 375 380gca gcg aat
cat aat ggt tct ccc ctt tct ata aag gaa gca ccc aaa 1200Ala Ala Asn
His Asn Gly Ser Pro Leu Ser Ile Lys Glu Ala Pro Lys385 390 395
400gaa ctc agc ttc ggt gca cgt gca gag ctg ccc agg atc cac cca gtt
1248Glu Leu Ser Phe Gly Ala Arg Ala Glu Leu Pro Arg Ile His Pro Val
405 410 415gca tcg aag taa 1260Ala Ser Lys18419PRTArtificial
SequenceSynthetic Construct 18Met Val Thr Gly Gly Met Ala Ser Lys
Trp Asp Gln Lys Gly Met Asp1 5 10 15Ile Ala Tyr Glu Glu Ala Leu Leu
Gly Tyr Lys Glu Gly Gly Val Pro 20 25 30Ile Gly Gly Cys Leu Ile Asn
Asn Lys Asp Gly Ser Val Leu Gly Arg 35 40 45Gly His Asn Met Arg Phe
Gln Lys Gly Ser Ala Thr Leu His Gly Glu 50 55 60Ile Ser Thr Leu Glu
Asn Cys Gly Arg Leu Glu Gly Lys Val Tyr Lys65 70 75 80Asp Thr Thr
Leu Tyr Thr Thr Leu Ser Pro Cys Asp Met Cys Thr Gly 85 90 95Ala Ile
Ile Met Tyr Gly Ile Pro Arg Cys Val Ile Gly Glu Asn Val 100 105
110Asn Phe Lys Ser Lys Gly Glu Lys Tyr Leu Gln Thr Arg Gly His Glu
115 120 125Val Val Val Val Asp Asp Glu Arg Cys Lys Lys Leu Met Lys
Gln Phe 130 135 140Ile Asp Glu Arg Pro Gln Asp Trp Phe Glu Asp Ile
Gly Glu Ser Gly145 150 155 160Gly Gly Ala Ser Gly Gly Gly Ala Ser
Gly Gly Gly Ala Ser Gly Gly 165 170 175Gly Ala Ala Val Ala Arg Ala
Ala Leu Gly Pro Leu Val Thr Gly Leu 180 185 190Tyr Asp Val Gln Ala
Phe Lys Phe Gly Asp Phe Val Leu Lys Ser Gly 195 200 205Leu Ser Ser
Pro Ile Tyr Ile Asp Leu Arg Gly Ile Val Ser Arg Pro 210 215 220Arg
Leu Leu Ser Gln Val Ala Asp Ile Leu Phe Gln Thr Ala Gln Asn225 230
235
240Ala Gly Ile Ser Phe Asp Thr Val Cys Gly Val Pro Tyr Thr Ala Leu
245 250 255Pro Leu Ala Thr Val Ile Cys Ser Thr Asn Gln Ile Pro Met
Leu Ile 260 265 270Arg Arg Lys Glu Thr Lys Asp Tyr Gly Thr Lys Arg
Leu Val Glu Gly 275 280 285Thr Ile Asn Pro Gly Glu Thr Cys Leu Ile
Ile Glu Asp Val Val Thr 290 295 300Ser Gly Ser Ser Val Leu Glu Thr
Val Glu Val Leu Gln Lys Glu Gly305 310 315 320Leu Lys Val Thr Asp
Ala Ile Val Leu Leu Asp Arg Glu Gln Gly Gly 325 330 335Lys Asp Lys
Leu Gln Ala His Gly Ile Arg Leu His Ser Val Cys Thr 340 345 350Leu
Ser Lys Met Leu Glu Ile Leu Glu Gln Gln Lys Lys Val Asp Ala 355 360
365Glu Thr Val Gly Arg Val Lys Arg Phe Ile Gln Glu Asn Val Phe Val
370 375 380Ala Ala Asn His Asn Gly Ser Pro Leu Ser Ile Lys Glu Ala
Pro Lys385 390 395 400Glu Leu Ser Phe Gly Ala Arg Ala Glu Leu Pro
Arg Ile His Pro Val 405 410 415Ala Ser Lys1911892DNAArtificial
SequenceRCR Vector - pAC3-yCD2 19tagttattaa tagtaatcaa ttacggggtc
attagttcat agcccatata tggagttccg 60cgttacataa cttacggtaa atggcccgcc
tggctgaccg cccaacgacc cccgcccatt 120gacgtcaata atgacgtatg
ttcccatagt aacgccaata gggactttcc attgacgtca 180atgggtggag
tatttacggt aaactgccca cttggcagta catcaagtgt atcatatgcc
240aagtacgccc cctattgacg tcaatgacgg taaatggccc gcctggcatt
atgcccagta 300catgacctta tgggactttc ctacttggca gtacatctac
gtattagtca tcgctattac 360catggtgatg cggttttggc agtacatcaa
tgggcgtgga tagcggtttg actcacgggg 420atttccaagt ctccacccca
ttgacgtcaa tgggagtttg ttttggcacc aaaatcaacg 480ggactttcca
aaatgtcgta acaactccgc cccattgacg caaatgggcg gtaggcgtgt
540acggtgggag gtctatataa gcagagctgg tttagtgaac cggcgccagt
cctccgattg 600actgagtcgc ccgggtaccc gtgtatccaa taaaccctct
tgcagttgca tccgacttgt 660ggtctcgctg ttccttggga gggtctcctc
tgagtgattg actacccgtc agcgggggtc 720tttcatttgg gggctcgtcc
gggatcggga gacccctgcc cagggaccac cgacccacca 780ccgggaggta
agctggccag caacttatct gtgtctgtcc gattgtctag tgtctatgac
840tgattttatg cgcctgcgtc ggtactagtt agctaactag ctctgtatct
ggcggacccg 900tggtggaact gacgagttcg gaacacccgg ccgcaaccct
gggagacgtc ccagggactt 960cgggggccgt ttttgtggcc cgacctgagt
ccaaaaatcc cgatcgtttt ggactctttg 1020gtgcaccccc cttagaggag
ggatatgtgg ttctggtagg agacgagaac ctaaaacagt 1080tcccgcctcc
gtctgaattt ttgctttcgg tttgggaccg aagccgcgcc gcgcgtcttg
1140tctgctgcag catcgttctg tgttgtctct gtctgactgt gtttctgtat
ttgtctgaga 1200atatgggcca gactgttacc actcccttaa gtttgacctt
aggtcactgg aaagatgtcg 1260agcggatcgc tcacaaccag tcggtagatg
tcaagaagag acgttgggtt accttctgct 1320ctgcagaatg gccaaccttt
aacgtcggat ggccgcgaga cggcaccttt aaccgagacc 1380tcatcaccca
ggttaagatc aaggtctttt cacctggccc gcatggacac ccagaccagg
1440tcccctacat cgtgacctgg gaagccttgg cttttgaccc ccctccctgg
gtcaagccct 1500ttgtacaccc taagcctccg cctcctcttc ctccatccgc
cccgtctctc ccccttgaac 1560ctcctcgttc gaccccgcct cgatcctccc
tttatccagc cctcactcct tctctaggcg 1620ccaaacctaa acctcaagtt
ctttctgaca gtggggggcc gctcatcgac ctacttacag 1680aagacccccc
gccttatagg gacccaagac cacccccttc cgacagggac ggaaatggtg
1740gagaagcgac ccctgcggga gaggcaccgg acccctcccc aatggcatct
cgcctacgtg 1800ggagacggga gccccctgtg gccgactcca ctacctcgca
ggcattcccc ctccgcgcag 1860gaggaaacgg acagcttcaa tactggccgt
tctcctcttc tgacctttac aactggaaaa 1920ataataaccc ttctttttct
gaagatccag gtaaactgac agctctgatc gagtctgttc 1980tcatcaccca
tcagcccacc tgggacgact gtcagcagct gttggggact ctgctgaccg
2040gagaagaaaa acaacgggtg ctcttagagg ctagaaaggc ggtgcggggc
gatgatgggc 2100gccccactca actgcccaat gaagtcgatg ccgcttttcc
cctcgagcgc ccagactggg 2160attacaccac ccaggcaggt aggaaccacc
tagtccacta tcgccagttg ctcctagcgg 2220gtctccaaaa cgcgggcaga
agccccacca atttggccaa ggtaaaagga ataacacaag 2280ggcccaatga
gtctccctcg gccttcctag agagacttaa ggaagcctat cgcaggtaca
2340ctccttatga ccctgaggac ccagggcaag aaactaatgt gtctatgtct
ttcatttggc 2400agtctgcccc agacattggg agaaagttag agaggttaga
agatttaaaa aacaagacgc 2460ttggagattt ggttagagag gcagaaaaga
tctttaataa acgagaaacc ccggaagaaa 2520gagaggaacg tatcaggaga
gaaacagagg aaaaagaaga acgccgtagg acagaggatg 2580agcagaaaga
gaaagaaaga gatcgtagga gacatagaga gatgagcaag ctattggcca
2640ctgtcgttag tggacagaaa caggatagac agggaggaga acgaaggagg
tcccaactcg 2700atcgcgacca gtgtgcctac tgcaaagaaa aggggcactg
ggctaaagat tgtcccaaga 2760aaccacgagg acctcgggga ccaagacccc
agacctccct cctgacccta gatgactagg 2820gaggtcaggg tcaggagccc
ccccctgaac ccaggataac cctcaaagtc ggggggcaac 2880ccgtcacctt
cctggtagat actggggccc aacactccgt gctgacccaa aatcctggac
2940ccctaagtga taagtctgcc tgggtccaag gggctactgg aggaaagcgg
tatcgctgga 3000ccacggatcg caaagtacat ctagctaccg gtaaggtcac
ccactctttc ctccatgtac 3060cagactgtcc ctatcctctg ttaggaagag
atttgctgac taaactaaaa gcccaaatcc 3120actttgaggg atcaggagcc
caggttatgg gaccaatggg gcagcccctg caagtgttga 3180ccctaaatat
agaagatgag catcggctac atgagacctc aaaagagcca gatgtttctc
3240tagggtccac atggctgtct gattttcctc aggcctgggc ggaaaccggg
ggcatgggac 3300tggcagttcg ccaagctcct ctgatcatac ctctgaaagc
aacctctacc cccgtgtcca 3360taaaacaata ccccatgtca caagaagcca
gactggggat caagccccac atacagagac 3420tgttggacca gggaatactg
gtaccctgcc agtccccctg gaacacgccc ctgctacccg 3480ttaagaaacc
agggactaat gattataggc ctgtccagga tctgagagaa gtcaacaagc
3540gggtggaaga catccacccc accgtgccca acccttacaa cctcttgagc
gggctcccac 3600cgtcccacca gtggtacact gtgcttgatt taaaggatgc
ctttttctgc ctgagactcc 3660accccaccag tcagcctctc ttcgcctttg
agtggagaga tccagagatg ggaatctcag 3720gacaattgac ctggaccaga
ctcccacagg gtttcaaaaa cagtcccacc ctgtttgatg 3780aggcactgca
cagagaccta gcagacttcc ggatccagca cccagacttg atcctgctac
3840agtacgtgga tgacttactg ctggccgcca cttctgagct agactgccaa
caaggtactc 3900gggccctgtt acaaacccta gggaacctcg ggtatcgggc
ctcggccaag aaagcccaaa 3960tttgccagaa acaggtcaag tatctggggt
atcttctaaa agagggtcag agatggctga 4020ctgaggccag aaaagagact
gtgatggggc agcctactcc gaagacccct cgacaactaa 4080gggagttcct
agggacggca ggcttctgtc gcctctggat ccctgggttt gcagaaatgg
4140cagccccctt gtaccctctc accaaaacgg ggactctgtt taattggggc
ccagaccaac 4200aaaaggccta tcaagaaatc aagcaagctc ttctaactgc
cccagccctg gggttgccag 4260atttgactaa gccctttgaa ctctttgtcg
acgagaagca gggctacgcc aaaggtgtcc 4320taacgcaaaa actgggacct
tggcgtcggc cggtggccta cctgtccaaa aagctagacc 4380cagtagcagc
tgggtggccc ccttgcctac ggatggtagc agccattgcc gtactgacaa
4440aggatgcagg caagctaacc atgggacagc cactagtcat tctggccccc
catgcagtag 4500aggcactagt caaacaaccc cccgaccgct ggctttccaa
cgcccggatg actcactatc 4560aggccttgct tttggacacg gaccgggtcc
agttcggacc ggtggtagcc ctgaacccgg 4620ctacgctgct cccactgcct
gaggaagggc tgcaacacaa ctgccttgat atcctggccg 4680aagcccacgg
aacccgaccc gacctaacgg accagccgct cccagacgcc gaccacacct
4740ggtacacgga tggaagcagt ctcttacaag agggacagcg taaggcggga
gctgcggtga 4800ccaccgagac cgaggtaatc tgggctaaag ccctgccagc
cgggacatcc gctcagcggg 4860ctgaactgat agcactcacc caggccctaa
agatggcaga aggtaagaag ctaaatgttt 4920atactgatag ccgttatgct
tttgctactg cccatatcca tggagaaata tacagaaggc 4980gtgggttgct
cacatcagaa ggcaaagaga tcaaaaataa agacgagatc ttggccctac
5040taaaagccct ctttctgccc aaaagactta gcataatcca ttgtccagga
catcaaaagg 5100gacacagcgc cgaggctaga ggcaaccgga tggctgacca
agcggcccga aaggcagcca 5160tcacagagac tccagacacc tctaccctcc
tcatagaaaa ttcatcaccc tacacctcag 5220aacattttca ttacacagtg
actgatataa aggacctaac caagttgggg gccatttatg 5280ataaaacaaa
gaagtattgg gtctaccaag gaaaacctgt gatgcctgac cagtttactt
5340ttgaattatt agactttctt catcagctga ctcacctcag cttctcaaaa
atgaaggctc 5400tcctagagag aagccacagt ccctactaca tgctgaaccg
ggatcgaaca ctcaaaaata 5460tcactgagac ctgcaaagct tgtgcacaag
tcaacgccag caagtctgcc gttaaacagg 5520gaactagggt ccgcgggcat
cggcccggca ctcattggga gatcgatttc accgagataa 5580agcccggatt
gtatggctat aaatatcttc tagtttttat agataccttt tctggctgga
5640tagaagcctt cccaaccaag aaagaaaccg ccaaggtcgt aaccaagaag
ctactagagg 5700agatcttccc caggttcggc atgcctcagg tattgggaac
tgacaatggg cctgccttcg 5760tctccaaggt gagtcagaca gtggccgatc
tgttggggat tgattggaaa ttacattgtg 5820catacagacc ccaaagctca
ggccaggtag aaagaatgaa tagaaccatc aaggagactt 5880taactaaatt
aacgcttgca actggctcta gagactgggt gctcctactc cccttagccc
5940tgtaccgagc ccgcaacacg ccgggccccc atggcctcac cccatatgag
atcttatatg 6000gggcaccccc gccccttgta aacttccctg accctgacat
gacaagagtt actaacagcc 6060cctctctcca agctcactta caggctctct
acttagtcca gcacgaagtc tggagacctc 6120tggcggcagc ctaccaagaa
caactggacc gaccggtggt acctcaccct taccgagtcg 6180gcgacacagt
gtgggtccgc cgacaccaga ctaagaacct agaacctcgc tggaaaggac
6240cttacacagt cctgctgacc acccccaccg ccctcaaagt agacggcatc
gcagcttgga 6300tacacgccgc ccacgtgaag gctgccgacc ccgggggtgg
accatcctct agactgacat 6360ggcgcgttca acgctctcaa aaccccctca
agataagatt aacccgtgga agcccttaat 6420agtcatggga gtcctgttag
gagtagggat ggcagagagc ccccatcagg tctttaatgt 6480aacctggaga
gtcaccaacc tgatgactgg gcgtaccgcc aatgccacct ccctcctggg
6540aactgtacaa gatgccttcc caaaattata ttttgatcta tgtgatctgg
tcggagagga 6600gtgggaccct tcagaccagg aaccgtatgt cgggtatggc
tgcaagtacc ccgcagggag 6660acagcggacc cggacttttg acttttacgt
gtgccctggg cataccgtaa agtcggggtg 6720tgggggacca ggagagggct
actgtggtaa atgggggtgt gaaaccaccg gacaggctta 6780ctggaagccc
acatcatcgt gggacctaat ctcccttaag cgcggtaaca ccccctggga
6840cacgggatgc tctaaagttg cctgtggccc ctgctacgac ctctccaaag
tatccaattc 6900cttccaaggg gctactcgag ggggcagatg caaccctcta
gtcctagaat tcactgatgc 6960aggaaaaaag gctaactggg acgggcccaa
atcgtgggga ctgagactgt accggacagg 7020aacagatcct attaccatgt
tctccctgac ccggcaggtc cttaatgtgg gaccccgagt 7080ccccataggg
cccaacccag tattacccga ccaaagactc ccttcctcac caatagagat
7140tgtaccggct ccacagccac ctagccccct caataccagt tacccccctt
ccactaccag 7200tacaccctca acctccccta caagtccaag tgtcccacag
ccacccccag gaactggaga 7260tagactacta gctctagtca aaggagccta
tcaggcgctt aacctcacca atcccgacaa 7320gacccaagaa tgttggctgt
gcttagtgtc gggacctcct tattacgaag gagtagcggt 7380cgtgggcact
tataccaatc attccaccgc tccggccaac tgtacggcca cttcccaaca
7440taagcttacc ctatctgaag tgacaggaca gggcctatgc atgggggcag
tacctaaaac 7500tcaccaggcc ttatgtaaca ccacccaaag cgccggctca
ggatcctact accttgcagc 7560acccgccgga acaatgtggg cttgcagcac
tggattgact ccctgcttgt ccaccacggt 7620gctcaatcta accacagatt
attgtgtatt agttgaactc tggcccagag taatttacca 7680ctcccccgat
tatatgtatg gtcagcttga acagcgtacc aaatataaaa gagagccagt
7740atcattgacc ctggcccttc tactaggagg attaaccatg ggagggattg
cagctggaat 7800agggacgggg accactgcct taattaaaac ccagcagttt
gagcagcttc atgccgctat 7860ccagacagac ctcaacgaag tcgaaaagtc
aattaccaac ctagaaaagt cactgacctc 7920gttgtctgaa gtagtcctac
agaaccgcag aggcctagat ttgctattcc taaaggaggg 7980aggtctctgc
gcagccctaa aagaagaatg ttgtttttat gcagaccaca cggggctagt
8040gagagacagc atggccaaat taagagaaag gcttaatcag agacaaaaac
tatttgagac 8100aggccaagga tggttcgaag ggctgtttaa tagatccccc
tggtttacca ccttaatctc 8160caccatcatg ggacctctaa tagtactctt
actgatctta ctctttggac cttgcattct 8220caatcgattg gtccaatttg
ttaaagacag gatctcagtg gtccaggctc tggttttgac 8280tcagcaatat
caccagctaa aacccataga gtacgagcca tgaacgcgtt actggccgaa
8340gccgcttgga ataaggccgg tgtgcgtttg tctatatgtt attttccacc
atattgccgt 8400cttttggcaa tgtgagggcc cggaaacctg gccctgtctt
cttgacgagc attcctaggg 8460gtctttcccc tctcgccaaa ggaatgcaag
gtctgttgaa tgtcgtgaag gaagcagttc 8520ctctggaagc ttcttgaaga
caaacaacgt ctgtagcgac cctttgcagg cagcggaacc 8580ccccacctgg
cgacaggtgc ctctgcggcc aaaagccacg tgtataagat acacctgcaa
8640aggcggcaca accccagtgc cacgttgtga gttggatagt tgtggaaaga
gtcaaatggc 8700tctcctcaag cgtattcaac aaggggctga aggatgccca
gaaggtaccc cattgtatgg 8760gatctgatct ggggcctcgg tgcacatgct
ttacatgtgt ttagtcgagg ttaaaaaaac 8820gtctaggccc cccgaaccac
ggggacgtgg ttttcctttg aaaaacacga ttataaatgg 8880tgaccggcgg
catggcctcc aagtgggatc aaaagggcat ggatatcgct tacgaggagg
8940ccctgctggg ctacaaggag ggcggcgtgc ctatcggcgg ctgtctgatc
aacaacaagg 9000acggcagtgt gctgggcagg ggccacaaca tgaggttcca
gaagggctcc gccaccctgc 9060acggcgagat ctccaccctg gagaactgtg
gcaggctgga gggcaaggtg tacaaggaca 9120ccaccctgta caccaccctg
tccccttgtg acatgtgtac cggcgctatc atcatgtacg 9180gcatccctag
gtgtgtgatc ggcgagaacg tgaacttcaa gtccaagggc gagaagtacc
9240tgcaaaccag gggccacgag gtggtggttg ttgacgatga gaggtgtaag
aagctgatga 9300agcagttcat cgacgagagg cctcaggact ggttcgagga
tatcggcgag taagcggccg 9360cagataaaat aaaagatttt atttagtctc
cagaaaaagg ggggaatgaa agaccccacc 9420tgtaggtttg gcaagctagc
ttaagtaacg ccattttgca aggcatggaa aaatacataa 9480ctgagaatag
agaagttcag atcaaggtca ggaacagatg gaacagctga atatgggcca
9540aacaggatat ctgtggtaag cagttcctgc cccggctcag ggccaagaac
agatggaaca 9600gctgaatatg ggccaaacag gatatctgtg gtaagcagtt
cctgccccgg ctcagggcca 9660agaacagatg gtccccagat gcggtccagc
cctcagcagt ttctagagaa ccatcagatg 9720tttccagggt gccccaagga
cctgaaatga ccctgtgcct tatttgaact aaccaatcag 9780ttcgcttctc
gcttctgttc gcgcgcttct gctccccgag ctcaataaaa gagcccacaa
9840cccctcactc ggggcgccag tcctccgatt gactgagtcg cccgggtacc
cgtgtatcca 9900ataaaccctc ttgcagttgc atccgacttg tggtctcgct
gttccttggg agggtctcct 9960ctgagtgatt gactacccgt cagcgggggt
ctttcattac atgtgagcaa aaggccagca 10020aaaggccagg aaccgtaaaa
aggccgcgtt gctggcgttt ttccataggc tccgcccccc 10080tgacgagcat
cacaaaaatc gacgctcaag tcagaggtgg cgaaacccga caggactata
10140aagataccag gcgtttcccc ctggaagctc cctcgtgcgc tctcctgttc
cgaccctgcc 10200gcttaccgga tacctgtccg cctttctccc ttcgggaagc
gtggcgcttt ctcaatgctc 10260acgctgtagg tatctcagtt cggtgtaggt
cgttcgctcc aagctgggct gtgtgcacga 10320accccccgtt cagcccgacc
gctgcgcctt atccggtaac tatcgtcttg agtccaaccc 10380ggtaagacac
gacttatcgc cactggcagc agccactggt aacaggatta gcagagcgag
10440gtatgtaggc ggtgctacag agttcttgaa gtggtggcct aactacggct
acactagaag 10500gacagtattt ggtatctgcg ctctgctgaa gccagttacc
ttcggaaaaa gagttggtag 10560ctcttgatcc ggcaaacaaa ccaccgctgg
tagcggtggt ttttttgttt gcaagcagca 10620gattacgcgc agaaaaaaag
gatctcaaga agatcctttg atcttttcta cggggtctga 10680cgctcagtgg
aacgaaaact cacgttaagg gattttggtc atgagattat caaaaaggat
10740cttcacctag atccttttaa attaaaaatg aagttttaaa tcaatctaaa
gtatatatga 10800gtaaacttgg tctgacagtt accaatgctt aatcagtgag
gcacctatct cagcgatctg 10860tctatttcgt tcatccatag ttgcctgact
ccccgtcgtg tagataacta cgatacggga 10920gggcttacca tctggcccca
gtgctgcaat gataccgcga gacccacgct caccggctcc 10980agatttatca
gcaataaacc agccagccgg aagggccgag cgcagaagtg gtcctgcaac
11040tttatccgcc tccatccagt ctattaattg ttgccgggaa gctagagtaa
gtagttcgcc 11100agttaatagt ttgcgcaacg ttgttgccat tgctgcaggc
atcgtggtgt cacgctcgtc 11160gtttggtatg gcttcattca gctccggttc
ccaacgatca aggcgagtta catgatcccc 11220catgttgtgc aaaaaagcgg
ttagctcctt cggtcctccg atcgttgtca gaagtaagtt 11280ggccgcagtg
ttatcactca tggttatggc agcactgcat aattctctta ctgtcatgcc
11340atccgtaaga tgcttttctg tgactggtga gtactcaacc aagtcattct
gagaatagtg 11400tatgcggcga ccgagttgct cttgcccggc gtcaacacgg
gataataccg cgccacatag 11460cagaacttta aaagtgctca tcattggaaa
acgttcttcg gggcgaaaac tctcaaggat 11520cttaccgctg ttgagatcca
gttcgatgta acccactcgt gcacccaact gatcttcagc 11580atcttttact
ttcaccagcg tttctgggtg agcaaaaaca ggaaggcaaa atgccgcaaa
11640aaagggaata agggcgacac ggaaatgttg aatactcata ctcttccttt
ttcaatatta 11700ttgaagcatt tatcagggtt attgtctcat gagcggatac
atatttgaat gtatttagaa 11760aaataaacaa ataggggttc cgcgcacatt
tccccgaaaa gtgccacctg acgtctaaga 11820aaccattatt atcatgacat
taacctataa aaataggcgt atcacgaggc cctttcgtct 11880tcaagaattc at
118922011892DNAArtificial SequenceRCR Vector - pAC3-yCD
20tagttattaa tagtaatcaa ttacggggtc attagttcat agcccatata tggagttccg
60cgttacataa cttacggtaa atggcccgcc tggctgaccg cccaacgacc cccgcccatt
120gacgtcaata atgacgtatg ttcccatagt aacgccaata gggactttcc
attgacgtca 180atgggtggag tatttacggt aaactgccca cttggcagta
catcaagtgt atcatatgcc 240aagtacgccc cctattgacg tcaatgacgg
taaatggccc gcctggcatt atgcccagta 300catgacctta tgggactttc
ctacttggca gtacatctac gtattagtca tcgctattac 360catggtgatg
cggttttggc agtacatcaa tgggcgtgga tagcggtttg actcacgggg
420atttccaagt ctccacccca ttgacgtcaa tgggagtttg ttttggcacc
aaaatcaacg 480ggactttcca aaatgtcgta acaactccgc cccattgacg
caaatgggcg gtaggcgtgt 540acggtgggag gtctatataa gcagagctgg
tttagtgaac cggcgccagt cctccgattg 600actgagtcgc ccgggtaccc
gtgtatccaa taaaccctct tgcagttgca tccgacttgt 660ggtctcgctg
ttccttggga gggtctcctc tgagtgattg actacccgtc agcgggggtc
720tttcatttgg gggctcgtcc gggatcggga gacccctgcc cagggaccac
cgacccacca 780ccgggaggta agctggccag caacttatct gtgtctgtcc
gattgtctag tgtctatgac 840tgattttatg cgcctgcgtc ggtactagtt
agctaactag ctctgtatct ggcggacccg 900tggtggaact gacgagttcg
gaacacccgg ccgcaaccct gggagacgtc ccagggactt 960cgggggccgt
ttttgtggcc cgacctgagt ccaaaaatcc cgatcgtttt ggactctttg
1020gtgcaccccc cttagaggag ggatatgtgg ttctggtagg agacgagaac
ctaaaacagt 1080tcccgcctcc gtctgaattt ttgctttcgg tttgggaccg
aagccgcgcc gcgcgtcttg 1140tctgctgcag catcgttctg tgttgtctct
gtctgactgt gtttctgtat ttgtctgaga 1200atatgggcca gactgttacc
actcccttaa gtttgacctt aggtcactgg aaagatgtcg 1260agcggatcgc
tcacaaccag tcggtagatg tcaagaagag acgttgggtt accttctgct
1320ctgcagaatg gccaaccttt aacgtcggat ggccgcgaga cggcaccttt
aaccgagacc 1380tcatcaccca ggttaagatc aaggtctttt cacctggccc
gcatggacac ccagaccagg 1440tcccctacat cgtgacctgg gaagccttgg
cttttgaccc ccctccctgg gtcaagccct 1500ttgtacaccc taagcctccg
cctcctcttc ctccatccgc cccgtctctc ccccttgaac 1560ctcctcgttc
gaccccgcct cgatcctccc tttatccagc cctcactcct tctctaggcg
1620ccaaacctaa acctcaagtt ctttctgaca gtggggggcc gctcatcgac
ctacttacag 1680aagacccccc gccttatagg gacccaagac cacccccttc
cgacagggac ggaaatggtg 1740gagaagcgac ccctgcggga gaggcaccgg
acccctcccc aatggcatct cgcctacgtg 1800ggagacggga gccccctgtg
gccgactcca ctacctcgca ggcattcccc ctccgcgcag 1860gaggaaacgg
acagcttcaa tactggccgt tctcctcttc tgacctttac aactggaaaa
1920ataataaccc
ttctttttct gaagatccag gtaaactgac agctctgatc gagtctgttc
1980tcatcaccca tcagcccacc tgggacgact gtcagcagct gttggggact
ctgctgaccg 2040gagaagaaaa acaacgggtg ctcttagagg ctagaaaggc
ggtgcggggc gatgatgggc 2100gccccactca actgcccaat gaagtcgatg
ccgcttttcc cctcgagcgc ccagactggg 2160attacaccac ccaggcaggt
aggaaccacc tagtccacta tcgccagttg ctcctagcgg 2220gtctccaaaa
cgcgggcaga agccccacca atttggccaa ggtaaaagga ataacacaag
2280ggcccaatga gtctccctcg gccttcctag agagacttaa ggaagcctat
cgcaggtaca 2340ctccttatga ccctgaggac ccagggcaag aaactaatgt
gtctatgtct ttcatttggc 2400agtctgcccc agacattggg agaaagttag
agaggttaga agatttaaaa aacaagacgc 2460ttggagattt ggttagagag
gcagaaaaga tctttaataa acgagaaacc ccggaagaaa 2520gagaggaacg
tatcaggaga gaaacagagg aaaaagaaga acgccgtagg acagaggatg
2580agcagaaaga gaaagaaaga gatcgtagga gacatagaga gatgagcaag
ctattggcca 2640ctgtcgttag tggacagaaa caggatagac agggaggaga
acgaaggagg tcccaactcg 2700atcgcgacca gtgtgcctac tgcaaagaaa
aggggcactg ggctaaagat tgtcccaaga 2760aaccacgagg acctcgggga
ccaagacccc agacctccct cctgacccta gatgactagg 2820gaggtcaggg
tcaggagccc ccccctgaac ccaggataac cctcaaagtc ggggggcaac
2880ccgtcacctt cctggtagat actggggccc aacactccgt gctgacccaa
aatcctggac 2940ccctaagtga taagtctgcc tgggtccaag gggctactgg
aggaaagcgg tatcgctgga 3000ccacggatcg caaagtacat ctagctaccg
gtaaggtcac ccactctttc ctccatgtac 3060cagactgtcc ctatcctctg
ttaggaagag atttgctgac taaactaaaa gcccaaatcc 3120actttgaggg
atcaggagcc caggttatgg gaccaatggg gcagcccctg caagtgttga
3180ccctaaatat agaagatgag catcggctac atgagacctc aaaagagcca
gatgtttctc 3240tagggtccac atggctgtct gattttcctc aggcctgggc
ggaaaccggg ggcatgggac 3300tggcagttcg ccaagctcct ctgatcatac
ctctgaaagc aacctctacc cccgtgtcca 3360taaaacaata ccccatgtca
caagaagcca gactggggat caagccccac atacagagac 3420tgttggacca
gggaatactg gtaccctgcc agtccccctg gaacacgccc ctgctacccg
3480ttaagaaacc agggactaat gattataggc ctgtccagga tctgagagaa
gtcaacaagc 3540gggtggaaga catccacccc accgtgccca acccttacaa
cctcttgagc gggctcccac 3600cgtcccacca gtggtacact gtgcttgatt
taaaggatgc ctttttctgc ctgagactcc 3660accccaccag tcagcctctc
ttcgcctttg agtggagaga tccagagatg ggaatctcag 3720gacaattgac
ctggaccaga ctcccacagg gtttcaaaaa cagtcccacc ctgtttgatg
3780aggcactgca cagagaccta gcagacttcc ggatccagca cccagacttg
atcctgctac 3840agtacgtgga tgacttactg ctggccgcca cttctgagct
agactgccaa caaggtactc 3900gggccctgtt acaaacccta gggaacctcg
ggtatcgggc ctcggccaag aaagcccaaa 3960tttgccagaa acaggtcaag
tatctggggt atcttctaaa agagggtcag agatggctga 4020ctgaggccag
aaaagagact gtgatggggc agcctactcc gaagacccct cgacaactaa
4080gggagttcct agggacggca ggcttctgtc gcctctggat ccctgggttt
gcagaaatgg 4140cagccccctt gtaccctctc accaaaacgg ggactctgtt
taattggggc ccagaccaac 4200aaaaggccta tcaagaaatc aagcaagctc
ttctaactgc cccagccctg gggttgccag 4260atttgactaa gccctttgaa
ctctttgtcg acgagaagca gggctacgcc aaaggtgtcc 4320taacgcaaaa
actgggacct tggcgtcggc cggtggccta cctgtccaaa aagctagacc
4380cagtagcagc tgggtggccc ccttgcctac ggatggtagc agccattgcc
gtactgacaa 4440aggatgcagg caagctaacc atgggacagc cactagtcat
tctggccccc catgcagtag 4500aggcactagt caaacaaccc cccgaccgct
ggctttccaa cgcccggatg actcactatc 4560aggccttgct tttggacacg
gaccgggtcc agttcggacc ggtggtagcc ctgaacccgg 4620ctacgctgct
cccactgcct gaggaagggc tgcaacacaa ctgccttgat atcctggccg
4680aagcccacgg aacccgaccc gacctaacgg accagccgct cccagacgcc
gaccacacct 4740ggtacacgga tggaagcagt ctcttacaag agggacagcg
taaggcggga gctgcggtga 4800ccaccgagac cgaggtaatc tgggctaaag
ccctgccagc cgggacatcc gctcagcggg 4860ctgaactgat agcactcacc
caggccctaa agatggcaga aggtaagaag ctaaatgttt 4920atactgatag
ccgttatgct tttgctactg cccatatcca tggagaaata tacagaaggc
4980gtgggttgct cacatcagaa ggcaaagaga tcaaaaataa agacgagatc
ttggccctac 5040taaaagccct ctttctgccc aaaagactta gcataatcca
ttgtccagga catcaaaagg 5100gacacagcgc cgaggctaga ggcaaccgga
tggctgacca agcggcccga aaggcagcca 5160tcacagagac tccagacacc
tctaccctcc tcatagaaaa ttcatcaccc tacacctcag 5220aacattttca
ttacacagtg actgatataa aggacctaac caagttgggg gccatttatg
5280ataaaacaaa gaagtattgg gtctaccaag gaaaacctgt gatgcctgac
cagtttactt 5340ttgaattatt agactttctt catcagctga ctcacctcag
cttctcaaaa atgaaggctc 5400tcctagagag aagccacagt ccctactaca
tgctgaaccg ggatcgaaca ctcaaaaata 5460tcactgagac ctgcaaagct
tgtgcacaag tcaacgccag caagtctgcc gttaaacagg 5520gaactagggt
ccgcgggcat cggcccggca ctcattggga gatcgatttc accgagataa
5580agcccggatt gtatggctat aaatatcttc tagtttttat agataccttt
tctggctgga 5640tagaagcctt cccaaccaag aaagaaaccg ccaaggtcgt
aaccaagaag ctactagagg 5700agatcttccc caggttcggc atgcctcagg
tattgggaac tgacaatggg cctgccttcg 5760tctccaaggt gagtcagaca
gtggccgatc tgttggggat tgattggaaa ttacattgtg 5820catacagacc
ccaaagctca ggccaggtag aaagaatgaa tagaaccatc aaggagactt
5880taactaaatt aacgcttgca actggctcta gagactgggt gctcctactc
cccttagccc 5940tgtaccgagc ccgcaacacg ccgggccccc atggcctcac
cccatatgag atcttatatg 6000gggcaccccc gccccttgta aacttccctg
accctgacat gacaagagtt actaacagcc 6060cctctctcca agctcactta
caggctctct acttagtcca gcacgaagtc tggagacctc 6120tggcggcagc
ctaccaagaa caactggacc gaccggtggt acctcaccct taccgagtcg
6180gcgacacagt gtgggtccgc cgacaccaga ctaagaacct agaacctcgc
tggaaaggac 6240cttacacagt cctgctgacc acccccaccg ccctcaaagt
agacggcatc gcagcttgga 6300tacacgccgc ccacgtgaag gctgccgacc
ccgggggtgg accatcctct agactgacat 6360ggcgcgttca acgctctcaa
aaccccctca agataagatt aacccgtgga agcccttaat 6420agtcatggga
gtcctgttag gagtagggat ggcagagagc ccccatcagg tctttaatgt
6480aacctggaga gtcaccaacc tgatgactgg gcgtaccgcc aatgccacct
ccctcctggg 6540aactgtacaa gatgccttcc caaaattata ttttgatcta
tgtgatctgg tcggagagga 6600gtgggaccct tcagaccagg aaccgtatgt
cgggtatggc tgcaagtacc ccgcagggag 6660acagcggacc cggacttttg
acttttacgt gtgccctggg cataccgtaa agtcggggtg 6720tgggggacca
ggagagggct actgtggtaa atgggggtgt gaaaccaccg gacaggctta
6780ctggaagccc acatcatcgt gggacctaat ctcccttaag cgcggtaaca
ccccctggga 6840cacgggatgc tctaaagttg cctgtggccc ctgctacgac
ctctccaaag tatccaattc 6900cttccaaggg gctactcgag ggggcagatg
caaccctcta gtcctagaat tcactgatgc 6960aggaaaaaag gctaactggg
acgggcccaa atcgtgggga ctgagactgt accggacagg 7020aacagatcct
attaccatgt tctccctgac ccggcaggtc cttaatgtgg gaccccgagt
7080ccccataggg cccaacccag tattacccga ccaaagactc ccttcctcac
caatagagat 7140tgtaccggct ccacagccac ctagccccct caataccagt
tacccccctt ccactaccag 7200tacaccctca acctccccta caagtccaag
tgtcccacag ccacccccag gaactggaga 7260tagactacta gctctagtca
aaggagccta tcaggcgctt aacctcacca atcccgacaa 7320gacccaagaa
tgttggctgt gcttagtgtc gggacctcct tattacgaag gagtagcggt
7380cgtgggcact tataccaatc attccaccgc tccggccaac tgtacggcca
cttcccaaca 7440taagcttacc ctatctgaag tgacaggaca gggcctatgc
atgggggcag tacctaaaac 7500tcaccaggcc ttatgtaaca ccacccaaag
cgccggctca ggatcctact accttgcagc 7560acccgccgga acaatgtggg
cttgcagcac tggattgact ccctgcttgt ccaccacggt 7620gctcaatcta
accacagatt attgtgtatt agttgaactc tggcccagag taatttacca
7680ctcccccgat tatatgtatg gtcagcttga acagcgtacc aaatataaaa
gagagccagt 7740atcattgacc ctggcccttc tactaggagg attaaccatg
ggagggattg cagctggaat 7800agggacgggg accactgcct taattaaaac
ccagcagttt gagcagcttc atgccgctat 7860ccagacagac ctcaacgaag
tcgaaaagtc aattaccaac ctagaaaagt cactgacctc 7920gttgtctgaa
gtagtcctac agaaccgcag aggcctagat ttgctattcc taaaggaggg
7980aggtctctgc gcagccctaa aagaagaatg ttgtttttat gcagaccaca
cggggctagt 8040gagagacagc atggccaaat taagagaaag gcttaatcag
agacaaaaac tatttgagac 8100aggccaagga tggttcgaag ggctgtttaa
tagatccccc tggtttacca ccttaatctc 8160caccatcatg ggacctctaa
tagtactctt actgatctta ctctttggac cttgcattct 8220caatcgattg
gtccaatttg ttaaagacag gatctcagtg gtccaggctc tggttttgac
8280tcagcaatat caccagctaa aacccataga gtacgagcca tgaacgcgtt
actggccgaa 8340gccgcttgga ataaggccgg tgtgcgtttg tctatatgtt
attttccacc atattgccgt 8400cttttggcaa tgtgagggcc cggaaacctg
gccctgtctt cttgacgagc attcctaggg 8460gtctttcccc tctcgccaaa
ggaatgcaag gtctgttgaa tgtcgtgaag gaagcagttc 8520ctctggaagc
ttcttgaaga caaacaacgt ctgtagcgac cctttgcagg cagcggaacc
8580ccccacctgg cgacaggtgc ctctgcggcc aaaagccacg tgtataagat
acacctgcaa 8640aggcggcaca accccagtgc cacgttgtga gttggatagt
tgtggaaaga gtcaaatggc 8700tctcctcaag cgtattcaac aaggggctga
aggatgccca gaaggtaccc cattgtatgg 8760gatctgatct ggggcctcgg
tgcacatgct ttacatgtgt ttagtcgagg ttaaaaaaac 8820gtctaggccc
cccgaaccac ggggacgtgg ttttcctttg aaaaacacga ttataaatgg
8880tgacaggggg aatggcaagc aagtgggatc agaagggtat ggacattgcc
tatgaggagg 8940cggccttagg ttacaaagag ggtggtgttc ctattggcgg
atgtcttatc aataacaaag 9000acggaagtgt tctcggtcgt ggtcacaaca
tgagatttca aaagggatcc gccacactac 9060atggtgagat ctccactttg
gaaaactgtg ggagattaga gggcaaagtg tacaaagata 9120ccactttgta
tacgacgctg tctccatgcg acatgtgtac aggtgccatc atcatgtatg
9180gtattccacg ctgtgttgtc ggtgagaacg ttaatttcaa aagtaagggc
gagaaatatt 9240tacaaactag aggtcacgag gttgttgttg ttgacgatga
gaggtgtaaa aagatcatga 9300aacaatttat cgatgaaaga cctcaggatt
ggtttgaaga tattggtgag taggcggccg 9360cagataaaat aaaagatttt
atttagtctc cagaaaaagg ggggaatgaa agaccccacc 9420tgtaggtttg
gcaagctagc ttaagtaacg ccattttgca aggcatggaa aaatacataa
9480ctgagaatag agaagttcag atcaaggtca ggaacagatg gaacagctga
atatgggcca 9540aacaggatat ctgtggtaag cagttcctgc cccggctcag
ggccaagaac agatggaaca 9600gctgaatatg ggccaaacag gatatctgtg
gtaagcagtt cctgccccgg ctcagggcca 9660agaacagatg gtccccagat
gcggtccagc cctcagcagt ttctagagaa ccatcagatg 9720tttccagggt
gccccaagga cctgaaatga ccctgtgcct tatttgaact aaccaatcag
9780ttcgcttctc gcttctgttc gcgcgcttct gctccccgag ctcaataaaa
gagcccacaa 9840cccctcactc ggggcgccag tcctccgatt gactgagtcg
cccgggtacc cgtgtatcca 9900ataaaccctc ttgcagttgc atccgacttg
tggtctcgct gttccttggg agggtctcct 9960ctgagtgatt gactacccgt
cagcgggggt ctttcattac atgtgagcaa aaggccagca 10020aaaggccagg
aaccgtaaaa aggccgcgtt gctggcgttt ttccataggc tccgcccccc
10080tgacgagcat cacaaaaatc gacgctcaag tcagaggtgg cgaaacccga
caggactata 10140aagataccag gcgtttcccc ctggaagctc cctcgtgcgc
tctcctgttc cgaccctgcc 10200gcttaccgga tacctgtccg cctttctccc
ttcgggaagc gtggcgcttt ctcaatgctc 10260acgctgtagg tatctcagtt
cggtgtaggt cgttcgctcc aagctgggct gtgtgcacga 10320accccccgtt
cagcccgacc gctgcgcctt atccggtaac tatcgtcttg agtccaaccc
10380ggtaagacac gacttatcgc cactggcagc agccactggt aacaggatta
gcagagcgag 10440gtatgtaggc ggtgctacag agttcttgaa gtggtggcct
aactacggct acactagaag 10500gacagtattt ggtatctgcg ctctgctgaa
gccagttacc ttcggaaaaa gagttggtag 10560ctcttgatcc ggcaaacaaa
ccaccgctgg tagcggtggt ttttttgttt gcaagcagca 10620gattacgcgc
agaaaaaaag gatctcaaga agatcctttg atcttttcta cggggtctga
10680cgctcagtgg aacgaaaact cacgttaagg gattttggtc atgagattat
caaaaaggat 10740cttcacctag atccttttaa attaaaaatg aagttttaaa
tcaatctaaa gtatatatga 10800gtaaacttgg tctgacagtt accaatgctt
aatcagtgag gcacctatct cagcgatctg 10860tctatttcgt tcatccatag
ttgcctgact ccccgtcgtg tagataacta cgatacggga 10920gggcttacca
tctggcccca gtgctgcaat gataccgcga gacccacgct caccggctcc
10980agatttatca gcaataaacc agccagccgg aagggccgag cgcagaagtg
gtcctgcaac 11040tttatccgcc tccatccagt ctattaattg ttgccgggaa
gctagagtaa gtagttcgcc 11100agttaatagt ttgcgcaacg ttgttgccat
tgctgcaggc atcgtggtgt cacgctcgtc 11160gtttggtatg gcttcattca
gctccggttc ccaacgatca aggcgagtta catgatcccc 11220catgttgtgc
aaaaaagcgg ttagctcctt cggtcctccg atcgttgtca gaagtaagtt
11280ggccgcagtg ttatcactca tggttatggc agcactgcat aattctctta
ctgtcatgcc 11340atccgtaaga tgcttttctg tgactggtga gtactcaacc
aagtcattct gagaatagtg 11400tatgcggcga ccgagttgct cttgcccggc
gtcaacacgg gataataccg cgccacatag 11460cagaacttta aaagtgctca
tcattggaaa acgttcttcg gggcgaaaac tctcaaggat 11520cttaccgctg
ttgagatcca gttcgatgta acccactcgt gcacccaact gatcttcagc
11580atcttttact ttcaccagcg tttctgggtg agcaaaaaca ggaaggcaaa
atgccgcaaa 11640aaagggaata agggcgacac ggaaatgttg aatactcata
ctcttccttt ttcaatatta 11700ttgaagcatt tatcagggtt attgtctcat
gagcggatac atatttgaat gtatttagaa 11760aaataaacaa ataggggttc
cgcgcacatt tccccgaaaa gtgccacctg acgtctaaga 11820aaccattatt
atcatgacat taacctataa aaataggcgt atcacgaggc cctttcgtct
11880tcaagaattc at 118922112007DNAArtificial SequenceRCR Vector -
pACE-CD 21tagttattaa tagtaatcaa ttacggggtc attagttcat agcccatata
tggagttccg 60cgttacataa cttacggtaa atggcccgcc tggctgaccg cccaacgacc
cccgcccatt 120gacgtcaata atgacgtatg ttcccatagt aacgccaata
gggactttcc attgacgtca 180atgggtggag tatttacggt aaactgccca
cttggcagta catcaagtgt atcatatgcc 240aagtacgccc cctattgacg
tcaatgacgg taaatggccc gcctggcatt atgcccagta 300catgacctta
tgggactttc ctacttggca gtacatctac gtattagtca tcgctattac
360catggtgatg cggttttggc agtacatcaa tgggcgtgga tagcggtttg
actcacgggg 420atttccaagt ctccacccca ttgacgtcaa tgggagtttg
ttttggcacc aaaatcaacg 480ggactttcca aaatgtcgta acaactccgc
cccattgacg caaatgggcg gtaggcgtgt 540acggtgggag gtctatataa
gcagagctgg tttagtgaac cggcgccagt cctccgattg 600actgagtcgc
ccgggtaccc gtgtatccaa taaaccctct tgcagttgca tccgacttgt
660ggtctcgctg ttccttggga gggtctcctc tgagtgattg actacccgtc
agcgggggtc 720tttcatttgg gggctcgtcc gggatcggga gacccctgcc
cagggaccac cgacccacca 780ccgggaggta agctggccag caacttatct
gtgtctgtcc gattgtctag tgtctatgac 840tgattttatg cgcctgcgtc
ggtactagtt agctaactag ctctgtatct ggcggacccg 900tggtggaact
gacgagttcg gaacacccgg ccgcaaccct gggagacgtc ccagggactt
960cgggggccgt ttttgtggcc cgacctgagt ccaaaaatcc cgatcgtttt
ggactctttg 1020gtgcaccccc cttagaggag ggatatgtgg ttctggtagg
agacgagaac ctaaaacagt 1080tcccgcctcc gtctgaattt ttgctttcgg
tttgggaccg aagccgcgcc gcgcgtcttg 1140tctgctgcag catcgttctg
tgttgtctct gtctgactgt gtttctgtat ttgtctgaga 1200atatgggcca
gactgttacc actcccttaa gtttgacctt aggtcactgg aaagatgtcg
1260agcggatcgc tcacaaccag tcggtagatg tcaagaagag acgttgggtt
accttctgct 1320ctgcagaatg gccaaccttt aacgtcggat ggccgcgaga
cggcaccttt aaccgagacc 1380tcatcaccca ggttaagatc aaggtctttt
cacctggccc gcatggacac ccagaccagg 1440tcccctacat cgtgacctgg
gaagccttgg cttttgaccc ccctccctgg gtcaagccct 1500ttgtacaccc
taagcctccg cctcctcttc ctccatccgc cccgtctctc ccccttgaac
1560ctcctcgttc gaccccgcct cgatcctccc tttatccagc cctcactcct
tctctaggcg 1620ccaaacctaa acctcaagtt ctttctgaca gtggggggcc
gctcatcgac ctacttacag 1680aagacccccc gccttatagg gacccaagac
cacccccttc cgacagggac ggaaatggtg 1740gagaagcgac ccctgcggga
gaggcaccgg acccctcccc aatggcatct cgcctacgtg 1800ggagacggga
gccccctgtg gccgactcca ctacctcgca ggcattcccc ctccgcgcag
1860gaggaaacgg acagcttcaa tactggccgt tctcctcttc tgacctttac
aactggaaaa 1920ataataaccc ttctttttct gaagatccag gtaaactgac
agctctgatc gagtctgttc 1980tcatcaccca tcagcccacc tgggacgact
gtcagcagct gttggggact ctgctgaccg 2040gagaagaaaa acaacgggtg
ctcttagagg ctagaaaggc ggtgcggggc gatgatgggc 2100gccccactca
actgcccaat gaagtcgatg ccgcttttcc cctcgagcgc ccagactggg
2160attacaccac ccaggcaggt aggaaccacc tagtccacta tcgccagttg
ctcctagcgg 2220gtctccaaaa cgcgggcaga agccccacca atttggccaa
ggtaaaagga ataacacaag 2280ggcccaatga gtctccctcg gccttcctag
agagacttaa ggaagcctat cgcaggtaca 2340ctccttatga ccctgaggac
ccagggcaag aaactaatgt gtctatgtct ttcatttggc 2400agtctgcccc
agacattggg agaaagttag agaggttaga agatttaaaa aacaagacgc
2460ttggagattt ggttagagag gcagaaaaga tctttaataa acgagaaacc
ccggaagaaa 2520gagaggaacg tatcaggaga gaaacagagg aaaaagaaga
acgccgtagg acagaggatg 2580agcagaaaga gaaagaaaga gatcgtagga
gacatagaga gatgagcaag ctattggcca 2640ctgtcgttag tggacagaaa
caggatagac agggaggaga acgaaggagg tcccaactcg 2700atcgcgacca
gtgtgcctac tgcaaagaaa aggggcactg ggctaaagat tgtcccaaga
2760aaccacgagg acctcgggga ccaagacccc agacctccct cctgacccta
gatgactagg 2820gaggtcaggg tcaggagccc ccccctgaac ccaggataac
cctcaaagtc ggggggcaac 2880ccgtcacctt cctggtagat actggggccc
aacactccgt gctgacccaa aatcctggac 2940ccctaagtga taagtctgcc
tgggtccaag gggctactgg aggaaagcgg tatcgctgga 3000ccacggatcg
caaagtacat ctagctaccg gtaaggtcac ccactctttc ctccatgtac
3060cagactgtcc ctatcctctg ttaggaagag atttgctgac taaactaaaa
gcccaaatcc 3120actttgaggg atcaggagcc caggttatgg gaccaatggg
gcagcccctg caagtgttga 3180ccctaaatat agaagatgag catcggctac
atgagacctc aaaagagcca gatgtttctc 3240tagggtccac atggctgtct
gattttcctc aggcctgggc ggaaaccggg ggcatgggac 3300tggcagttcg
ccaagctcct ctgatcatac ctctgaaagc aacctctacc cccgtgtcca
3360taaaacaata ccccatgtca caagaagcca gactggggat caagccccac
atacagagac 3420tgttggacca gggaatactg gtaccctgcc agtccccctg
gaacacgccc ctgctacccg 3480ttaagaaacc agggactaat gattataggc
ctgtccagga tctgagagaa gtcaacaagc 3540gggtggaaga catccacccc
accgtgccca acccttacaa cctcttgagc gggctcccac 3600cgtcccacca
gtggtacact gtgcttgatt taaaggatgc ctttttctgc ctgagactcc
3660accccaccag tcagcctctc ttcgcctttg agtggagaga tccagagatg
ggaatctcag 3720gacaattgac ctggaccaga ctcccacagg gtttcaaaaa
cagtcccacc ctgtttgatg 3780aggcactgca cagagaccta gcagacttcc
ggatccagca cccagacttg atcctgctac 3840agtacgtgga tgacttactg
ctggccgcca cttctgagct agactgccaa caaggtactc 3900gggccctgtt
acaaacccta gggaacctcg ggtatcgggc ctcggccaag aaagcccaaa
3960tttgccagaa acaggtcaag tatctggggt atcttctaaa agagggtcag
agatggctga 4020ctgaggccag aaaagagact gtgatggggc agcctactcc
gaagacccct cgacaactaa 4080gggagttcct agggacggca ggcttctgtc
gcctctggat ccctgggttt gcagaaatgg 4140cagccccctt gtaccctctc
accaaaacgg ggactctgtt taattggggc ccagaccaac 4200aaaaggccta
tcaagaaatc aagcaagctc ttctaactgc cccagccctg gggttgccag
4260atttgactaa gccctttgaa ctctttgtcg acgagaagca gggctacgcc
aaaggtgtcc 4320taacgcaaaa actgggacct tggcgtcggc cggtggccta
cctgtccaaa aagctagacc 4380cagtagcagc tgggtggccc ccttgcctac
ggatggtagc agccattgcc gtactgacaa 4440aggatgcagg caagctaacc
atgggacagc cactagtcat tctggccccc catgcagtag 4500aggcactagt
caaacaaccc cccgaccgct ggctttccaa cgcccggatg actcactatc
4560aggccttgct tttggacacg gaccgggtcc agttcggacc ggtggtagcc
ctgaacccgg 4620ctacgctgct cccactgcct gaggaagggc tgcaacacaa
ctgccttgat atcctggccg 4680aagcccacgg aacccgaccc gacctaacgg
accagccgct cccagacgcc gaccacacct 4740ggtacacgga tggaagcagt
ctcttacaag agggacagcg taaggcggga gctgcggtga 4800ccaccgagac
cgaggtaatc tgggctaaag ccctgccagc cgggacatcc gctcagcggg
4860ctgaactgat agcactcacc caggccctaa agatggcaga aggtaagaag
ctaaatgttt 4920atactgatag ccgttatgct tttgctactg cccatatcca
tggagaaata tacagaaggc 4980gtgggttgct
cacatcagaa ggcaaagaga tcaaaaataa agacgagatc ttggccctac
5040taaaagccct ctttctgccc aaaagactta gcataatcca ttgtccagga
catcaaaagg 5100gacacagcgc cgaggctaga ggcaaccgga tggctgacca
agcggcccga aaggcagcca 5160tcacagagac tccagacacc tctaccctcc
tcatagaaaa ttcatcaccc tacacctcag 5220aacattttca ttacacagtg
actgatataa aggacctaac caagttgggg gccatttatg 5280ataaaacaaa
gaagtattgg gtctaccaag gaaaacctgt gatgcctgac cagtttactt
5340ttgaattatt agactttctt catcagctga ctcacctcag cttctcaaaa
atgaaggctc 5400tcctagagag aagccacagt ccctactaca tgctgaaccg
ggatcgaaca ctcaaaaata 5460tcactgagac ctgcaaagct tgtgcacaag
tcaacgccag caagtctgcc gttaaacagg 5520gaactagggt ccgcgggcat
cggcccggca ctcattggga gatcgatttc accgagataa 5580agcccggatt
gtatggctat aaatatcttc tagtttttat agataccttt tctggctgga
5640tagaagcctt cccaaccaag aaagaaaccg ccaaggtcgt aaccaagaag
ctactagagg 5700agatcttccc caggttcggc atgcctcagg tattgggaac
tgacaatggg cctgccttcg 5760tctccaaggt gagtcagaca gtggccgatc
tgttggggat tgattggaaa ttacattgtg 5820catacagacc ccaaagctca
ggccaggtag aaagaatgaa tagaaccatc aaggagactt 5880taactaaatt
aacgcttgca actggctcta gagactgggt gctcctactc cccttagccc
5940tgtaccgagc ccgcaacacg ccgggccccc atggcctcac cccatatgag
atcttatatg 6000gggcaccccc gccccttgta aacttccctg accctgacat
gacaagagtt actaacagcc 6060cctctctcca agctcactta caggctctct
acttagtcca gcacgaagtc tggagacctc 6120tggcggcagc ctaccaagaa
caactggacc gaccggtggt acctcaccct taccgagtcg 6180gcgacacagt
gtgggtccgc cgacaccaga ctaagaacct agaacctcgc tggaaaggac
6240cttacacagt cctgctgacc acccccaccg ccctcaaagt agacggcatc
gcagcttgga 6300tacacgccgc ccacgtgaag gctgccgacc ccgggggtgg
accatcctct agactgacat 6360ggcgcgttca acgctctcaa aaccccctca
agataagatt aacccgtgga agcccttaat 6420agtcatggga gtcctgttag
gagtagggat ggcagagagc ccccatcagg tctttaatgt 6480aacctggaga
gtcaccaacc tgatgactgg gcgtaccgcc aatgccacct ccctcctggg
6540aactgtacaa gatgccttcc caaaattata ttttgatcta tgtgatctgg
tcggagagga 6600gtgggaccct tcagaccagg aaccgtatgt cgggtatggc
tgcaagtacc ccgcagggag 6660acagcggacc cggacttttg acttttacgt
gtgccctggg cataccgtaa agtcggggtg 6720tgggggacca ggagagggct
actgtggtaa atgggggtgt gaaaccaccg gacaggctta 6780ctggaagccc
acatcatcgt gggacctaat ctcccttaag cgcggtaaca ccccctggga
6840cacgggatgc tctaaagttg cctgtggccc ctgctacgac ctctccaaag
tatccaattc 6900cttccaaggg gctactcgag ggggcagatg caaccctcta
gtcctagaat tcactgatgc 6960aggaaaaaag gctaactggg acgggcccaa
atcgtgggga ctgagactgt accggacagg 7020aacagatcct attaccatgt
tctccctgac ccggcaggtc cttaatgtgg gaccccgagt 7080ccccataggg
cccaacccag tattacccga ccaaagactc ccttcctcac caatagagat
7140tgtaccggct ccacagccac ctagccccct caataccagt tacccccctt
ccactaccag 7200tacaccctca acctccccta caagtccaag tgtcccacag
ccacccccag gaactggaga 7260tagactacta gctctagtca aaggagccta
tcaggcgctt aacctcacca atcccgacaa 7320gacccaagaa tgttggctgt
gcttagtgtc gggacctcct tattacgaag gagtagcggt 7380cgtgggcact
tataccaatc attccaccgc tccggccaac tgtacggcca cttcccaaca
7440taagcttacc ctatctgaag tgacaggaca gggcctatgc atgggggcag
tacctaaaac 7500tcaccaggcc ttatgtaaca ccacccaaag cgccggctca
ggatcctact accttgcagc 7560acccgccgga acaatgtggg cttgcagcac
tggattgact ccctgcttgt ccaccacggt 7620gctcaatcta accacagatt
attgtgtatt agttgaactc tggcccagag taatttacca 7680ctcccccgat
tatatgtatg gtcagcttga acagcgtacc aaatataaaa gagagccagt
7740atcattgacc ctggcccttc tactaggagg attaaccatg ggagggattg
cagctggaat 7800agggacgggg accactgcct taattaaaac ccagcagttt
gagcagcttc atgccgctat 7860ccagacagac ctcaacgaag tcgaaaagtc
aattaccaac ctagaaaagt cactgacctc 7920gttgtctgaa gtagtcctac
agaaccgcag aggcctagat ttgctattcc taaaggaggg 7980aggtctctgc
gcagccctaa aagaagaatg ttgtttttat gcagaccaca cggggctagt
8040gagagacagc atggccaaat taagagaaag gcttaatcag agacaaaaac
tatttgagac 8100aggccaagga tggttcgaag ggctgtttaa tagatccccc
tggtttacca ccttaatctc 8160caccatcatg ggacctctaa tagtactctt
actgatctta ctctttggac cttgcattct 8220caatcgatta gtccaatttg
ttaaagacag gatatcagtg gtccaggctc tagttttgac 8280tcaacaatat
caccagctga agcctataga gtacgagcca tgacgtacgt tactggccga
8340agccgcttgg aataaggccg gtgtgcgttt gtctatatgt tattttccac
catattgccg 8400tcttttggca atgtgagggc ccggaaacct ggccctgtct
tcttgacgag cattcctagg 8460ggtctttccc ctctcgccaa aggaatgcaa
ggtctgttga atgtcgtgaa ggaagcagtt 8520cctctggaag cttcttgaag
acaaacaacg tctgtagcga ccctttgcag gcagcggaac 8580cccccacctg
gcgacaggtg cctctgcggc caaaagccac gtgtataaga tacacctgca
8640aaggcggcac aaccccagtg ccacgttgtg agttggatag ttgtggaaag
agtcaaatgg 8700ctctcctcaa gcgtattcaa caaggggctg aaggatgccc
agaaggtacc ccattgtatg 8760ggatctgatc tggggcctcg gtgcacatgc
tttacatgtg tttagtcgag gttaaaaaaa 8820cgtctaggcc ccccgaacca
cggggacgtg gttttccttt gaaaaacacg ataataccat 8880ggtgacaggg
ggaatggcaa gcaagtggga tcagaagggt atggacattg cctatgagga
8940ggcggcctta ggttacaaag agggtggtgt tcctattggc ggatgtctta
tcaataacaa 9000agacggaagt gttctcggtc gtggtcacaa catgagattt
caaaagggat ccgccacact 9060acatggtgag atctccactt tggaaaactg
tgggagatta gagggcaaag tgtacaaaga 9120taccactttg tatacgacgc
tgtctccatg cgacatgtgt acaggtgcca tcatcatgta 9180tggtattcca
cgctgtgttg tcggtgagaa cgttaatttc aaaagtaagg gcgagaaata
9240tttacaaact agaggtcacg aggttgttgt tgttgacgat gagaggtgta
aaaagatcat 9300gaaacaattt atcgatgaaa gacctcagga ttggtttgaa
gatattggtg agtaggcggc 9360cgcgccatag ataaaataaa agattttatt
tagtctccag aaaaaggggg gaatgaaaga 9420ccccacctgt aggtttggca
agctagctta agtaacgcca ttttgcaagg catggaaaaa 9480tacataactg
agaatagaga agttcagatc aaggtcagga acagatggaa cagctgaata
9540tgggccaaac aggatatctg tggtaagcag ttcctgcccc ggctcagggc
caagaacaga 9600tggaacagct gaatatgggc caaacaggat atctgtggta
agcagttcct gccccggctc 9660agggccaaga acagatggtc cccagatgcg
gtccagccct cagcagtttc tagagaacca 9720tcagatgttt ccagggtgcc
ccaaggacct gaaatgaccc tgtgccttgt ttaaactaac 9780caatcagttc
gcttctcgct tctgttcgcg cgcttctgct ccccgagctc aataaaagag
9840cccacaaccc ctcactcggg gcgccagtcc tccgattgac tgagtcgccc
gggtacccgt 9900gtatccaata aaccctcttg cagttgcatc cgacttgtgg
tctcgctgtt ccttgggagg 9960gtctcctctg agtgattgac tacccgtcag
cgggggtctt tcatttgggg gctcgtccgg 10020gatcgggaga cccctgccca
gggaccaccg acccaccacc gggaggtaag ctggctgcct 10080cgcgcgtttc
ggtgatgacg gtgaaaacct ctgacatgtg agcaaaaggc cagcaaaagg
10140ccaggaaccg taaaaaggcc gcgttgctgg cgtttttcca taggctccgc
ccccctgacg 10200agcatcacaa aaatcgacgc tcaagtcaga ggtggcgaaa
cccgacagga ctataaagat 10260accaggcgtt tccccctgga agctccctcg
tgcgctctcc tgttccgacc ctgccgctta 10320ccggatacct gtccgccttt
ctcccttcgg gaagcgtggc gctttctcaa tgctcacgct 10380gtaggtatct
cagttcggtg taggtcgttc gctccaagct gggctgtgtg cacgaacccc
10440ccgttcagcc cgaccgctgc gccttatccg gtaactatcg tcttgagtcc
aacccggtaa 10500gacacgactt atcgccactg gcagcagcca ctggtaacag
gattagcaga gcgaggtatg 10560taggcggtgc tacagagttc ttgaagtggt
ggcctaacta cggctacact agaaggacag 10620tatttggtat ctgcgctctg
ctgaagccag ttaccttcgg aaaaagagtt ggtagctctt 10680gatccggcaa
acaaaccacc gctggtagcg gtggtttttt tgtttgcaag cagcagatta
10740cgcgcagaaa aaaaggatct caagaagatc ctttgatctt ttctacgggg
tctgacgctc 10800agtggaacga aaactcacgt taagggattt tggtcatgag
attatcaaaa aggatcttca 10860cctagatcct tttaaattaa aaatgaagtt
ttaaatcaat ctaaagtata tatgagtaaa 10920cttggtctga cagttaccaa
tgcttaatca gtgaggcacc tatctcagcg atctgtctat 10980ttcgttcatc
catagttgcc tgactccccg tcgtgtagat aactacgata cgggagggct
11040taccatctgg ccccagtgct gcaatgatac cgcgagaccc acgctcaccg
gctccagatt 11100tatcagcaat aaaccagcca gccggaaggg ccgagcgcag
aagtggtcct gcaactttat 11160ccgcctccat ccagtctatt aattgttgcc
gggaagctag agtaagtagt tcgccagtta 11220atagtttgcg caacgttgtt
gccattgctg caggcatcgt ggtgtcacgc tcgtcgtttg 11280gtatggcttc
attcagctcc ggttcccaac gatcaaggcg agttacatga tcccccatgt
11340tgtgcaaaaa agcggttagc tccttcggtc ctccgatcgt tgtcagaagt
aagttggccg 11400cagtgttatc actcatggtt atggcagcac tgcataattc
tcttactgtc atgccatccg 11460taagatgctt ttctgtgact ggtgagtact
caaccaagtc attctgagaa tagtgtatgc 11520ggcgaccgag ttgctcttgc
ccggcgtcaa cacgggataa taccgcgcca catagcagaa 11580ctttaaaagt
gctcatcatt ggaaaacgtt cttcggggcg aaaactctca aggatcttac
11640cgctgttgag atccagttcg atgtaaccca ctcgtgcacc caactgatct
tcagcatctt 11700ttactttcac cagcgtttct gggtgagcaa aaacaggaag
gcaaaatgcc gcaaaaaagg 11760gaataagggc gacacggaaa tgttgaatac
tcatactctt cctttttcaa tattattgaa 11820gcatttatca gggttattgt
ctcatgagcg gatacatatt tgaatgtatt tagaaaaata 11880aacaaatagg
ggttccgcgc acatttcccc gaaaagtgcc acctgacgtc taagaaacca
11940ttattatcat gacattaacc tataaaaata ggcgtatcac gaggcccttt
cgtcttcaag 12000aattcat 120072211893DNAArtificial SequenceRCR
Vector - pAC3-yCD2 22tagttattaa tagtaatcaa ttacggggtc attagttcat
agcccatata tggagttccg 60cgttacataa cttacggtaa atggcccgcc tggctgaccg
cccaacgacc cccgcccatt 120gacgtcaata atgacgtatg ttcccatagt
aacgccaata gggactttcc attgacgtca 180atgggtggag tatttacggt
aaactgccca cttggcagta catcaagtgt atcatatgcc 240aagtacgccc
cctattgacg tcaatgacgg taaatggccc gcctggcatt atgcccagta
300catgacctta tgggactttc ctacttggca gtacatctac gtattagtca
tcgctattac 360catggtgatg cggttttggc agtacatcaa tgggcgtgga
tagcggtttg actcacgggg 420atttccaagt ctccacccca ttgacgtcaa
tgggagtttg ttttggcacc aaaatcaacg 480ggactttcca aaatgtcgta
acaactccgc cccattgacg caaatgggcg gtaggcgtgt 540acggtgggag
gtctatataa gcagagctgg tttagtgaac cggcgccagt cctccgattg
600actgagtcgc ccgggtaccc gtgtatccaa taaaccctct tgcagttgca
tccgacttgt 660ggtctcgctg ttccttggga gggtctcctc tgagtgattg
actacccgtc agcgggggtc 720tttcatttgg gggctcgtcc gggatcggga
gacccctgcc cagggaccac cgacccacca 780ccgggaggta agctggccag
caacttatct gtgtctgtcc gattgtctag tgtctatgac 840tgattttatg
cgcctgcgtc ggtactagtt agctaactag ctctgtatct ggcggacccg
900tggtggaact gacgagttcg gaacacccgg ccgcaaccct gggagacgtc
ccagggactt 960cgggggccgt ttttgtggcc cgacctgagt ccaaaaatcc
cgatcgtttt ggactctttg 1020gtgcaccccc cttagaggag ggatatgtgg
ttctggtagg agacgagaac ctaaaacagt 1080tcccgcctcc gtctgaattt
ttgctttcgg tttgggaccg aagccgcgcc gcgcgtcttg 1140tctgctgcag
catcgttctg tgttgtctct gtctgactgt gtttctgtat ttgtctgaaa
1200atatgggcca gactgttacc actcccttaa gtttgacctt aggtcactgg
aaagatgtcg 1260agcggatcgc tcacaaccag tcggtagatg tcaagaagag
acgttgggtt accttctgct 1320ctgcagaatg gccaaccttt aacgtcggat
ggccgcgaga cggcaccttt aaccgagacc 1380tcatcaccca ggttaagatc
aaggtctttt cacctggccc gcatggacac ccagaccagg 1440tcccctacat
cgtgacctgg gaagccttgg cttttgaccc ccctccctgg gtcaagccct
1500ttgtacaccc taagcctccg cctcctcttc ctccatccgc cccgtctctc
ccccttgaac 1560ctcctcgttc gaccccgcct cgatcctccc tttatccagc
cctcactcct tctctaggcg 1620ccaaacctaa acctcaagtt ctttctgaca
gtggggggcc gctcatcgac ctacttacag 1680aagacccccc gccttatagg
gacccaagac cacccccttc cgacagggac ggaaatggtg 1740gagaagcgac
ccctgcggga gaggcaccgg acccctcccc aatggcatct cgcctacgtg
1800ggagacggga gccccctgtg gccgactcca ctacctcgca ggcattcccc
ctccgcgcag 1860gaggaaacgg acagcttcaa tactggccgt tctcctcttc
tgacctttac aactggaaaa 1920ataataaccc ttctttttct gaagatccag
gtaaactgac agctctgatc gagtctgtcc 1980tcatcaccca tcagcccacc
tgggacgact gtcagcagct gttggggact ctgctgaccg 2040gagaagaaaa
acaacgggtg ctcttagagg ctagaaaggc ggtgcggggc gatgatgggc
2100gccccactca actgcccaat gaagtcgatg ccgcttttcc cctcgagcgc
ccagactggg 2160attacaccac ccaggcaggt aggaaccacc tagtccacta
tcgccagttg ctcctagcgg 2220gtctccaaaa cgcgggcaga agccccacca
atttggccaa ggtaaaagga ataacacaag 2280ggcccaatga gtctccctcg
gccttcctag agagacttaa ggaagcctat cgcaggtaca 2340ctccttatga
ccctgaggac ccagggcaag aaactaatgt gtctatgtct ttcatttggc
2400agtctgcccc agacattggg agaaagttag agaggttaga agatttaaaa
aacaagacgc 2460ttggagattt ggttagagag gcagaaaaga tctttaataa
acgagaaacc ccggaagaaa 2520gagaggaacg tatcaggaga gaaacagagg
aaaaagaaga acgccgtagg acagaggatg 2580agcagaaaga gaaagaaaga
gatcgtagga gacatagaga gatgagcaag ctattggcca 2640ctgtcgttag
tggacagaaa caggatagac agggaggaga acgaaggagg tcccaactcg
2700atcgcgacca gtgtgcctac tgcaaagaaa aggggcactg ggctaaagat
tgtcccaaga 2760aaccacgagg acctcgggga ccaagacccc agacctccct
cctgacccta gatgactagg 2820gaggtcaggg tcaggagccc ccccctgaac
ccaggataac cctcaaagtc ggggggcaac 2880ccgtcacctt cctggtagat
actggggccc aacactccgt gctgacccaa aatcctggac 2940ccctaagtga
taagtctgcc tgggtccaag gggctactgg aggaaagcgg tatcgctgga
3000ccacggatcg caaagtacat ctagctaccg gtaaggtcac ccactctttc
ctccatgtac 3060cagactgtcc ctatcctctg ttaggaagag atttgctgac
taaactaaaa gcccaaatcc 3120actttgaggg atcaggagcc caggttatgg
gaccaatggg gcagcccctg caagtgttga 3180ccctaaatat agaagatgag
tatcggctac atgagacctc aaaagagcca gatgtttctc 3240tagggtccac
atggctgtct gattttcctc aggcctgggc ggaaaccggg ggcatgggac
3300tggcagttcg ccaagctcct ctgatcatac ctctgaaagc aacctctacc
cccgtgtcca 3360taaaacaata ccccatgtca caagaagcca gactggggat
caagccccac atacagagac 3420tgttggacca gggaatactg gtaccctgcc
agtccccctg gaacacgccc ctgctacccg 3480ttaagaaacc agggactaat
gattataggc ctgtccagga tctgagagaa gtcaacaagc 3540gggtggaaga
catccacccc accgtgccca acccttacaa cctcttgagc gggctcccac
3600cgtcccacca gtggtacact gtgcttgatt taaaggatgc ctttttctgc
ctgagactcc 3660accccaccag tcagcctctc ttcgcctttg agtggagaga
tccagagatg ggaatctcag 3720gacaattgac ctggaccaga ctcccacagg
gtttcaaaaa cagtcccacc ctgtttgatg 3780aggcactgca cagagaccta
gcagacttcc ggatccagca cccagacttg atcctgctac 3840agtacgtgga
tgacttactg ctggccgcca cttctgagct agactgccaa caaggtactc
3900gggccctgtt acaaacccta gggaacctcg ggtatcgggc ctcggccaag
aaagcccaaa 3960tttgccagaa acaggtcaag tatctggggt atcttctaaa
agagggtcag agatggctga 4020ctgaggccag aaaagagact gtgatggggc
agcctactcc gaagacccct cgacaactaa 4080gggagttcct agggacggca
ggcttctgtc gcctctggat ccctgggttt gcagaaatgg 4140cagccccctt
gtaccctctc accaaaacgg ggactctgtt taattggggc ccagaccaac
4200aaaaggccta tcaagaaatc aagcaagctc ttctaactgc cccagccctg
gggttgccag 4260atttgactaa gccctttgaa ctctttgtcg acgagaagca
gggctacgcc aaaggtgtcc 4320taacgcaaaa actgggacct tggcgtcggc
cggtggccta cctgtccaaa aagctagacc 4380cagtagcagc tgggtggccc
ccttgcctac ggatggtagc agccattgcc gtactgacaa 4440aggatgcagg
caagctaacc atgggacagc cactagtcat tctggccccc catgcagtag
4500aggcactagt caaacaaccc cccgaccgct ggctttccaa cgcccggatg
actcactatc 4560aggccttgct tttggacacg gaccgggtcc agttcggacc
ggtggtagcc ctgaacccgg 4620ctacgctgct cccactgcct gaggaagggc
tgcaacacaa ctgccttgat atcctggccg 4680aagcccacgg aacccgaccc
gacctaacgg accagccgct cccagacgcc gaccacacct 4740ggtacacgga
tggaagcagt ctcttacaag agggacagcg taaggcggga gctgcggtga
4800ccaccgagac cgaggtaatc tgggctaaag ccctgccagc cgggacatcc
gctcagcggg 4860ctgaactgat agcactcacc caggccctaa agatggcaga
aggtaagaag ctaaatgttt 4920atactgatag ccgttatgct tttgctactg
cccatatcca tggagaaata tacagaaggc 4980gtgggttgct cacatcagaa
ggcaaagaga tcaaaaataa agacgagatc ttggccctac 5040taaaagccct
ctttctgccc aaaagactta gcataatcca ttgtccagga catcaaaagg
5100gacacagcgc cgaggctaga ggcaaccgga tggctgacca agcggcccga
aaggcagcca 5160tcacagagac tccagacacc tctaccctcc tcatagaaaa
ttcatcaccc tacacctcag 5220aacattttca ttacacagtg actgatataa
aggacctaac caagttgggg gccatttatg 5280ataaaacaaa gaagtattgg
gtctaccaag gaaaacctgt gatgcctgac cagtttactt 5340ttgaattatt
agactttctt catcagctga ctcacctcag cttctcaaaa atgaaggctc
5400tcctagagag aagccacagt ccctactaca tgctgaaccg ggatcgaaca
ctcaaaaata 5460tcactgagac ctgcaaagct tgtgcacaag tcaacgccag
caagtctgcc gttaaacagg 5520gaactagggt ccgcgggcat cggcccggca
ctcattggga gatcgatttc accgagataa 5580agcccggatt gtatggctat
aaatatcttc tagtttttat agataccttt tctggctgga 5640tagaagcctt
cccaaccaag aaagaaaccg ccaaggtcgt aaccaagaag ctactagagg
5700agatcttccc caggttcggc atgcctcagg tattgggaac tgacaatggg
cctgccttcg 5760tctccaaggt gagtcagaca gtggccgatc tgttggggat
tgattggaaa ttacattgtg 5820catacagacc ccaaagctca ggccaggtag
aaagaatgaa tagaaccatc aaggagactt 5880taactaaatt aacgcttgca
actggctcta gagactgggt gctcctactc cccttagccc 5940tgtaccgagc
ccgcaacacg ccgggccccc atggcctcac cccatatgag atcttatatg
6000gggcaccccc gccccttgta aacttccctg accctgacat gacaagagtt
actaacagcc 6060cctctctcca agctcactta caggctctct acttagtcca
gcacgaagtc tggagacctc 6120tggcggcagc ctaccaagaa caactggacc
gaccggtggt acctcaccct taccgagtcg 6180gcgacacagt gtgggtccgc
cgacaccaga ctaagaacct agaacctcgc tggaaaggac 6240cttacacagt
cctgctgacc acccccaccg ccctcaaagt agacggcatc gcagcttgga
6300tacacgccgc ccacgtgaag gctgccgacc ccgggggtgg accatcctct
agactgacat 6360ggcgcgttca acgctctcaa aaccccctca agataagatt
aacccgtgga agcccttaat 6420agtcatggga gtcctgttag gagtagggat
ggcagagagc ccccatcagg tctttaatgt 6480aacctggaga gtcaccaacc
tgatgactgg gcgtaccgcc aatgccacct ccctcctggg 6540aactgtacaa
gatgccttcc caaaattata ttttgatcta tgtgatctgg tcggagagga
6600gtgggaccct tcagaccagg aaccgtatgt cgggtatggc tgcaagtacc
ccgcagggag 6660acagcggacc cggacttttg acttttacgt gtgccctggg
cataccgtaa agtcggggtg 6720tgggggacca ggagagggct actgtggtaa
atgggggtgt gaaaccaccg gacaggctta 6780ctggaagccc acatcatcgt
gggacctaat ctcccttaag cgcggtaaca ccccctggga 6840cacgggatgc
tctaaagttg cctgtggccc ctgctacgac ctctccaaag tatccaattc
6900cttccaaggg gctactcgag ggggcagatg caaccctcta gtcctagaat
tcactgatgc 6960aggaaaaaag gctaactggg acgggcccaa atcgtgggga
ctgagactgt accggacagg 7020aacagatcct attaccatgt tctccctgac
ccggcaggtc cttaatgtgg gaccccgagt 7080ccccataggg cccaacccag
tattacccga ccaaagactc ccttcctcac caatagagat 7140tgtaccggct
ccacagccac ctagccccct caataccagt tacccccctt ccactaccag
7200tacaccctca acctccccta caagtccaag tgtcccacag ccacccccag
gaactggaga 7260tagactacta gctctagtca aaggagccta tcaggcgctt
aacctcacca atcccgacaa 7320gacccaagaa tgttggctgt gcttagtgtc
gggacctcct tattacgaag gagtagcggt 7380cgtgggcact tataccaatc
attccaccgc tccggccaac tgtacggcca cttcccaaca 7440taagcttacc
ctatctgaag tgacaggaca gggcctatgc atgggggcag tacctaaaac
7500tcaccaggcc ttatgtaaca ccacccaaag cgccggctca ggatcctact
accttgcagc 7560acccgccgga acaatgtggg cttgcagcac tggattgact
ccctgcttgt ccaccacggt 7620gctcaatcta accacagatt attgtgtatt
agttgaactc tggcccagag taatttacca 7680ctcccccgat tatatgtatg
gtcagcttga acagcgtacc aaatataaaa gagagccagt 7740atcattgacc
ctggcccttc tactaggagg attaaccatg ggagggattg cagctggaat
7800agggacgggg accactgcct taattaaaac ccagcagttt gagcagcttc
atgccgctat 7860ccagacagac ctcaacgaag tcgaaaagtc aattaccaac
ctagaaaagt cactgacctc 7920gttgtctgaa
gtagtcctac agaaccgcag aggcctagat ttgctattcc taaaggaggg
7980aggtctctgc gcagccctaa aagaagaatg ttgtttttat gcagaccaca
cggggctagt 8040gagagacagc atggccaaat taagagaaag gcttaatcag
agacaaaaac tatttgagac 8100aggccaagga tggttcgaag ggctgtttaa
tagatccccc tggtttacca ccttaatctc 8160caccatcatg ggacctctaa
tagtactctt actgatctta ctctttggac cttgcattct 8220caatcgattg
gtccaatttg ttaaagacag gatctcagtg gtccaggctc tggttttgac
8280tcagcaatat caccagctaa aacccataga gtacgagcca tgaacgcgtt
actggccgaa 8340gccgcttgga ataaggccgg tgtgcgtttg tctatatgtt
attttccacc atattgccgt 8400cttttggcaa tgtgagggcc cggaaacctg
gccctgtctt cttgacgagc attcctaggg 8460gtctttcccc tctcgccaaa
ggaatgcaag gtctgttgaa tgtcgtgaag gaagcagttc 8520ctctggaagc
ttcttgaaga caaacaacgt ctgtagcgac cctttgcagg cagcggaacc
8580ccccacctgg cgacaggtgc ctctgcggcc aaaagccacg tgtataagat
acacctgcaa 8640aggcggcaca accccagtgc cacgttgtga gttggatagt
tgtggaaaga gtcaaatggc 8700tctcctcaag cgtattcaac aaggggctga
aggatgccca gaaggtaccc cattgtatgg 8760gatctgatct ggggcctcgg
tgcacatgct ttacatgtgt ttagtcgagg ttaaaaaaac 8820gtctaggccc
cccgaaccac ggggacgtgg ttttcctttg aaaaacacga ttataaatgg
8880tgaccggcgg catggcctcc aagtgggatc aaaagggcat ggatatcgct
tacgaggagg 8940ccctgctggg ctacaaggag ggcggcgtgc ctatcggcgg
ctgtctgatc aacaacaagg 9000acggcagtgt gctgggcagg ggccacaaca
tgaggttcca gaagggctcc gccaccctgc 9060acggcgagat ctccaccctg
gagaactgtg gcaggctgga gggcaaggtg tacaaggaca 9120ccaccctgta
caccaccctg tccccttgtg acatgtgtac cggcgctatc atcatgtacg
9180gcatccctag gtgtgtgatc ggcgagaacg tgaacttcaa gtccaagggc
gagaagtacc 9240tgcaaaccag gggccacgag gtggtggttg ttgacgatga
gaggtgtaag aagctgatga 9300agcagttcat cgacgagagg cctcaggact
ggttcgagga tatcggcgag taagcggccg 9360cagataaaat aaaagatttt
atttagtctc cagaaaaagg ggggaatgaa agaccccacc 9420tgtaggtttg
gcaagctagc ttaagtaacg ccattttgca aggcatggaa aaatacataa
9480ctgagaatag agaagttcag atcaaggtca ggaacagatg gaacagctga
atatgggcca 9540aacaggatat ctgtggtaag cagttcctgc cccggctcag
ggccaagaac agatggaaca 9600gctgaatatg ggccaaacag gatatctgtg
gtaagcagtt cctgccccgg ctcagggcca 9660agaacagatg gtccccagat
gcggtccagc cctcagcagt ttctagagaa ccatcagatg 9720tttccagggt
gccccaagga cctgaaatga ccctgtgcct tatttgaact aaccaatcag
9780ttcgcttctc gcttctgttc gcgcgcttct gctccccgag ctcaataaaa
gagcccacaa 9840cccctcactc ggggcgccag tcctccgatt gactgagtcg
cccgggtacc cgtgtatcca 9900ataaaccctc ttgcagttgc atccgacttg
tggtctcgct gttccttggg agggtctcct 9960ctgagtgatt gactacccgt
cagcgggggt ctttcattac atgtgagcaa aaggccagca 10020aaaggccagg
aaccgtaaaa aggccgcgtt gctggcgttt ttccataggc tccgcccccc
10080tgacgagcat cacaaaaatc gacgctcaag tcagaggtgg cgaaacccga
caggactata 10140aagataccag gcgtttcccc ctggaagctc cctcgtgcgc
tctcctgttc cgaccctgcc 10200gcttaccgga tacctgtccg cctttctccc
ttcgggaagc gtggcgcttt ctcatagctc 10260acgctgtagg tatctcagtt
cggtgtaggt cgttcgctcc aagctgggct gtgtgcacga 10320accccccgtt
cagcccgacc gctgcgcctt atccggtaac tatcgtcttg agtccaaccc
10380ggtaagacac gacttatcgc cactggcagc agccactggt aacaggatta
gcagagcgag 10440gtatgtaggc ggtgctacag agttcttgaa gtggtggcct
aactacggct acactagaag 10500gacagtattt ggtatctgcg ctctgctgaa
gccagttacc ttcggaaaaa gagttggtag 10560ctcttgatcc ggcaaacaaa
ccaccgctgg tagcggtggt ttttttgttt gcaagcagca 10620gattacgcgc
agaaaaaaag gatctcaaga agatcctttg atcttttcta cggggtctga
10680cgctcagtgg aacgaaaact cacgttaagg gattttggtc atgagattat
caaaaaggat 10740cttcacctag atccttttaa attaaaaatg aagttttaaa
tcaatctaaa gtatatatga 10800gtaaacttgg tctgacagtt accaatgctt
aatcagtgag gcacctatct cagcgatctg 10860tctatttcgt tcatccatag
ttgcctgact ccccgtcgtg tagataacta cgatacggga 10920gggcttacca
tctggcccca gtgctgcaat gataccgcga gacccacgct caccggctcc
10980agatttatca gcaataaacc agccagccgg aagggccgag cgcagaagtg
gtcctgcaac 11040tttatccgcc tccatccagt ctattaattg ttgccgggaa
gctagagtaa gtagttcgcc 11100agttaatagt ttgcgcaacg ttgttgccat
tgctgcaggc atcgtggtgt cacgctcgtc 11160gtttggtatg gcttcattca
gctccggttc ccaacgatca aggcgagtta catgatcccc 11220catgttgtgc
aaaaaagcgg ttagctcctt cggtcctccg atcgttgtca gaagtaagtt
11280ggccgcagtg ttatcactca tggttatggc agcactgcat aattctctta
ctgtcatgcc 11340atccgtaaga tgcttttctg tgactggtga gtactcaacc
aagtcattct gagaatagtg 11400tatgcggcga ccgagttgct cttgcccggc
gtcaacacgg gataataccg cgccacatag 11460cagaacttta aaagtgctca
tcattggaaa acgttcttcg gggcgaaaac tctcaaggat 11520cttaccgctg
ttgagatcca gttcgatgta acccactcgt gcacccaact gatcttcagc
11580atcttttact ttcaccagcg tttctgggtg agcaaaaaca ggaaggcaaa
atgccgcaaa 11640aaagggaata agggcgacac ggaaatgttg aatactcata
ctcttccttt ttcaatatta 11700ttgaagcatt tatcagggtt attgtctcat
gagcggatac atatttgaat gtatttagaa 11760aaataaacaa ataggggttc
cgcgcacatt tccccgaaaa gtgccacctg acgtctaaga 11820aaccattatt
atcatgacat taacctataa aaataggcgt atcacgaggc cctttcgtct
11880tcaagaattc cat 11893234473DNAHomo sapiensCDS(175)..(3942)
23aaggggaggt aaccctggcc cctttggtcg gggccccggg cagccgcgcg ccccttccca
60cggggccctt tactgcgccg cgcgcccggc ccccacccct cgcagcaccc cgcgccccgc
120gccctcccag ccgggtccag ccggagccat ggggccggag ccgcagtgag cacc atg
177 Met 1gag ctg gcg gcc ttg tgc cgc tgg ggg ctc ctc ctc gcc ctc
ttg ccc 225Glu Leu Ala Ala Leu Cys Arg Trp Gly Leu Leu Leu Ala Leu
Leu Pro 5 10 15ccc gga gcc gcg agc acc caa gtg tgc acc ggc aca gac
atg aag ctg 273Pro Gly Ala Ala Ser Thr Gln Val Cys Thr Gly Thr Asp
Met Lys Leu 20 25 30cgg ctc cct gcc agt ccc gag acc cac ctg gac atg
ctc cgc cac ctc 321Arg Leu Pro Ala Ser Pro Glu Thr His Leu Asp Met
Leu Arg His Leu 35 40 45tac cag ggc tgc cag gtg gtg cag gga aac ctg
gaa ctc acc tac ctg 369Tyr Gln Gly Cys Gln Val Val Gln Gly Asn Leu
Glu Leu Thr Tyr Leu50 55 60 65ccc acc aat gcc agc ctg tcc ttc ctg
cag gat atc cag gag gtg cag 417Pro Thr Asn Ala Ser Leu Ser Phe Leu
Gln Asp Ile Gln Glu Val Gln 70 75 80ggc tac gtg ctc atc gct cac aac
caa gtg agg cag gtc cca ctg cag 465Gly Tyr Val Leu Ile Ala His Asn
Gln Val Arg Gln Val Pro Leu Gln 85 90 95agg ctg cgg att gtg cga ggc
acc cag ctc ttt gag gac aac tat gcc 513Arg Leu Arg Ile Val Arg Gly
Thr Gln Leu Phe Glu Asp Asn Tyr Ala 100 105 110ctg gcc gtg cta gac
aat gga gac ccg ctg aac aat acc acc cct gtc 561Leu Ala Val Leu Asp
Asn Gly Asp Pro Leu Asn Asn Thr Thr Pro Val 115 120 125aca ggg gcc
tcc cca gga ggc ctg cgg gag ctg cag ctt cga agc ctc 609Thr Gly Ala
Ser Pro Gly Gly Leu Arg Glu Leu Gln Leu Arg Ser Leu130 135 140
145aca gag atc ttg aaa gga ggg gtc ttg atc cag cgg aac ccc cag ctc
657Thr Glu Ile Leu Lys Gly Gly Val Leu Ile Gln Arg Asn Pro Gln Leu
150 155 160tgc tac cag gac acg att ttg tgg aag gac atc ttc cac aag
aac aac 705Cys Tyr Gln Asp Thr Ile Leu Trp Lys Asp Ile Phe His Lys
Asn Asn 165 170 175cag ctg gct ctc aca ctg ata gac acc aac cgc tct
cgg gcc tgc cac 753Gln Leu Ala Leu Thr Leu Ile Asp Thr Asn Arg Ser
Arg Ala Cys His 180 185 190ccc tgt tct ccg atg tgt aag ggc tcc cgc
tgc tgg gga gag agt tct 801Pro Cys Ser Pro Met Cys Lys Gly Ser Arg
Cys Trp Gly Glu Ser Ser 195 200 205gag gat tgt cag agc ctg acg cgc
act gtc tgt gcc ggt ggc tgt gcc 849Glu Asp Cys Gln Ser Leu Thr Arg
Thr Val Cys Ala Gly Gly Cys Ala210 215 220 225cgc tgc aag ggg cca
ctg ccc act gac tgc tgc cat gag cag tgt gct 897Arg Cys Lys Gly Pro
Leu Pro Thr Asp Cys Cys His Glu Gln Cys Ala 230 235 240gcc ggc tgc
acg ggc ccc aag cac tct gac tgc ctg gcc tgc ctc cac 945Ala Gly Cys
Thr Gly Pro Lys His Ser Asp Cys Leu Ala Cys Leu His 245 250 255ttc
aac cac agt ggc atc tgt gag ctg cac tgc cca gcc ctg gtc acc 993Phe
Asn His Ser Gly Ile Cys Glu Leu His Cys Pro Ala Leu Val Thr 260 265
270tac aac aca gac acg ttt gag tcc atg ccc aat ccc gag ggc cgg tat
1041Tyr Asn Thr Asp Thr Phe Glu Ser Met Pro Asn Pro Glu Gly Arg Tyr
275 280 285aca ttc ggc gcc agc tgt gtg act gcc tgt ccc tac aac tac
ctt tct 1089Thr Phe Gly Ala Ser Cys Val Thr Ala Cys Pro Tyr Asn Tyr
Leu Ser290 295 300 305acg gac gtg gga tcc tgc acc ctc gtc tgc ccc
ctg cac aac caa gag 1137Thr Asp Val Gly Ser Cys Thr Leu Val Cys Pro
Leu His Asn Gln Glu 310 315 320gtg aca gca gag gat gga aca cag cgg
tgt gag aag tgc agc aag ccc 1185Val Thr Ala Glu Asp Gly Thr Gln Arg
Cys Glu Lys Cys Ser Lys Pro 325 330 335tgt gcc cga gtg tgc tat ggt
ctg ggc atg gag cac ttg cga gag gtg 1233Cys Ala Arg Val Cys Tyr Gly
Leu Gly Met Glu His Leu Arg Glu Val 340 345 350agg gca gtt acc agt
gcc aat atc cag gag ttt gct ggc tgc aag aag 1281Arg Ala Val Thr Ser
Ala Asn Ile Gln Glu Phe Ala Gly Cys Lys Lys 355 360 365atc ttt ggg
agc ctg gca ttt ctg ccg gag agc ttt gat ggg gac cca 1329Ile Phe Gly
Ser Leu Ala Phe Leu Pro Glu Ser Phe Asp Gly Asp Pro370 375 380
385gcc tcc aac act gcc ccg ctc cag cca gag cag ctc caa gtg ttt gag
1377Ala Ser Asn Thr Ala Pro Leu Gln Pro Glu Gln Leu Gln Val Phe Glu
390 395 400act ctg gaa gag atc aca ggt tac cta tac atc tca gca tgg
ccg gac 1425Thr Leu Glu Glu Ile Thr Gly Tyr Leu Tyr Ile Ser Ala Trp
Pro Asp 405 410 415agc ctg cct gac ctc agc gtc ttc cag aac ctg caa
gta atc cgg gga 1473Ser Leu Pro Asp Leu Ser Val Phe Gln Asn Leu Gln
Val Ile Arg Gly 420 425 430cga att ctg cac aat ggc gcc tac tcg ctg
acc ctg caa ggg ctg ggc 1521Arg Ile Leu His Asn Gly Ala Tyr Ser Leu
Thr Leu Gln Gly Leu Gly 435 440 445atc agc tgg ctg ggg ctg cgc tca
ctg agg gaa ctg ggc agt gga ctg 1569Ile Ser Trp Leu Gly Leu Arg Ser
Leu Arg Glu Leu Gly Ser Gly Leu450 455 460 465gcc ctc atc cac cat
aac acc cac ctc tgc ttc gtg cac acg gtg ccc 1617Ala Leu Ile His His
Asn Thr His Leu Cys Phe Val His Thr Val Pro 470 475 480tgg gac cag
ctc ttt cgg aac ccg cac caa gct ctg ctc cac act gcc 1665Trp Asp Gln
Leu Phe Arg Asn Pro His Gln Ala Leu Leu His Thr Ala 485 490 495aac
cgg cca gag gac gag tgt gtg ggc gag ggc ctg gcc tgc cac cag 1713Asn
Arg Pro Glu Asp Glu Cys Val Gly Glu Gly Leu Ala Cys His Gln 500 505
510ctg tgc gcc cga ggg cac tgc tgg ggt cca ggg ccc acc cag tgt gtc
1761Leu Cys Ala Arg Gly His Cys Trp Gly Pro Gly Pro Thr Gln Cys Val
515 520 525aac tgc agc cag ttc ctt cgg ggc cag gag tgc gtg gag gaa
tgc cga 1809Asn Cys Ser Gln Phe Leu Arg Gly Gln Glu Cys Val Glu Glu
Cys Arg530 535 540 545gta ctg cag ggg ctc ccc agg gag tat gtg aat
gcc agg cac tgt ttg 1857Val Leu Gln Gly Leu Pro Arg Glu Tyr Val Asn
Ala Arg His Cys Leu 550 555 560ccg tgc cac cct gag tgt cag ccc cag
aat ggc tca gtg acc tgt ttt 1905Pro Cys His Pro Glu Cys Gln Pro Gln
Asn Gly Ser Val Thr Cys Phe 565 570 575gga ccg gag gct gac cag tgt
gtg gcc tgt gcc cac tat aag gac cct 1953Gly Pro Glu Ala Asp Gln Cys
Val Ala Cys Ala His Tyr Lys Asp Pro 580 585 590ccc ttc tgc gtg gcc
cgc tgc ccc agc ggt gtg aaa cct gac ctc tcc 2001Pro Phe Cys Val Ala
Arg Cys Pro Ser Gly Val Lys Pro Asp Leu Ser 595 600 605tac atg ccc
atc tgg aag ttt cca gat gag gag ggc gca tgc cag cct 2049Tyr Met Pro
Ile Trp Lys Phe Pro Asp Glu Glu Gly Ala Cys Gln Pro610 615 620
625tgc ccc atc aac tgc acc cac tcc tgt gtg gac ctg gat gac aag ggc
2097Cys Pro Ile Asn Cys Thr His Ser Cys Val Asp Leu Asp Asp Lys Gly
630 635 640tgc ccc gcc gag cag aga gcc agc cct ctg acg tcc atc atc
tct gcg 2145Cys Pro Ala Glu Gln Arg Ala Ser Pro Leu Thr Ser Ile Ile
Ser Ala 645 650 655gtg gtt ggc att ctg ctg gtc gtg gtc ttg ggg gtg
gtc ttt ggg atc 2193Val Val Gly Ile Leu Leu Val Val Val Leu Gly Val
Val Phe Gly Ile 660 665 670ctc atc aag cga cgg cag cag aag atc cgg
aag tac acg atg cgg aga 2241Leu Ile Lys Arg Arg Gln Gln Lys Ile Arg
Lys Tyr Thr Met Arg Arg 675 680 685ctg ctg cag gaa acg gag ctg gtg
gag ccg ctg aca cct agc gga gcg 2289Leu Leu Gln Glu Thr Glu Leu Val
Glu Pro Leu Thr Pro Ser Gly Ala690 695 700 705atg ccc aac cag gcg
cag atg cgg atc ctg aaa gag acg gag ctg agg 2337Met Pro Asn Gln Ala
Gln Met Arg Ile Leu Lys Glu Thr Glu Leu Arg 710 715 720aag gtg aag
gtg ctt gga tct ggc gct ttt ggc aca gtc tac aag ggc 2385Lys Val Lys
Val Leu Gly Ser Gly Ala Phe Gly Thr Val Tyr Lys Gly 725 730 735atc
tgg atc cct gat ggg gag aat gtg aaa att cca gtg gcc atc aaa 2433Ile
Trp Ile Pro Asp Gly Glu Asn Val Lys Ile Pro Val Ala Ile Lys 740 745
750gtg ttg agg gaa aac aca tcc ccc aaa gcc aac aaa gaa atc tta gac
2481Val Leu Arg Glu Asn Thr Ser Pro Lys Ala Asn Lys Glu Ile Leu Asp
755 760 765gaa gca tac gtg atg gct ggt gtg ggc tcc cca tat gtc tcc
cgc ctt 2529Glu Ala Tyr Val Met Ala Gly Val Gly Ser Pro Tyr Val Ser
Arg Leu770 775 780 785ctg ggc atc tgc ctg aca tcc acg gtg cag ctg
gtg aca cag ctt atg 2577Leu Gly Ile Cys Leu Thr Ser Thr Val Gln Leu
Val Thr Gln Leu Met 790 795 800ccc tat ggc tgc ctc tta gac cat gtc
cgg gaa aac cgc gga cgc ctg 2625Pro Tyr Gly Cys Leu Leu Asp His Val
Arg Glu Asn Arg Gly Arg Leu 805 810 815ggc tcc cag gac ctg ctg aac
tgg tgt atg cag att gcc aag ggg atg 2673Gly Ser Gln Asp Leu Leu Asn
Trp Cys Met Gln Ile Ala Lys Gly Met 820 825 830agc tac ctg gag gat
gtg cgg ctc gta cac agg gac ttg gcc gct cgg 2721Ser Tyr Leu Glu Asp
Val Arg Leu Val His Arg Asp Leu Ala Ala Arg 835 840 845aac gtg ctg
gtc aag agt ccc aac cat gtc aaa att aca gac ttc ggg 2769Asn Val Leu
Val Lys Ser Pro Asn His Val Lys Ile Thr Asp Phe Gly850 855 860
865ctg gct cgg ctg ctg gac att gac gag aca gag tac cat gca gat ggg
2817Leu Ala Arg Leu Leu Asp Ile Asp Glu Thr Glu Tyr His Ala Asp Gly
870 875 880ggc aag gtg ccc atc aag tgg atg gcg ctg gag tcc att ctc
cgc cgg 2865Gly Lys Val Pro Ile Lys Trp Met Ala Leu Glu Ser Ile Leu
Arg Arg 885 890 895cgg ttc acc cac cag agt gat gtg tgg agt tat ggt
gtg act gtg tgg 2913Arg Phe Thr His Gln Ser Asp Val Trp Ser Tyr Gly
Val Thr Val Trp 900 905 910gag ctg atg act ttt ggg gcc aaa cct tac
gat ggg atc cca gcc cgg 2961Glu Leu Met Thr Phe Gly Ala Lys Pro Tyr
Asp Gly Ile Pro Ala Arg 915 920 925gag atc cct gac ctg ctg gaa aag
ggg gag cgg ctg ccc cag ccc ccc 3009Glu Ile Pro Asp Leu Leu Glu Lys
Gly Glu Arg Leu Pro Gln Pro Pro930 935 940 945atc tgc acc att gat
gtc tac atg atc atg gtc aaa tgt tgg atg att 3057Ile Cys Thr Ile Asp
Val Tyr Met Ile Met Val Lys Cys Trp Met Ile 950 955 960gac tct gaa
tgt cgg cca aga ttc cgg gag ttg gtg tct gaa ttc tcc 3105Asp Ser Glu
Cys Arg Pro Arg Phe Arg Glu Leu Val Ser Glu Phe Ser 965 970 975cgc
atg gcc agg gac ccc cag cgc ttt gtg gtc atc cag aat gag gac 3153Arg
Met Ala Arg Asp Pro Gln Arg Phe Val Val Ile Gln Asn Glu Asp 980 985
990ttg ggc cca gcc agt ccc ttg gac agc acc ttc tac cgc tca ctg ctg
3201Leu Gly Pro Ala Ser Pro Leu Asp Ser Thr Phe Tyr Arg Ser Leu Leu
995 1000 1005gag gac gat gac atg ggg gac ctg gtg gat gct gag gag
tat ctg 3246Glu Asp Asp Asp Met Gly Asp Leu Val Asp Ala Glu Glu Tyr
Leu1010 1015 1020gta ccc cag cag ggc ttc ttc tgt cca gac cct gcc
ccg ggc gct 3291Val Pro Gln Gln Gly Phe Phe Cys Pro Asp Pro Ala Pro
Gly Ala1025 1030 1035ggg ggc atg gtc cac cac agg cac cgc agc tca
tct acc agg agt 3336Gly Gly Met Val His His Arg His Arg Ser Ser Ser
Thr Arg Ser1040 1045 1050ggc ggt ggg gac ctg aca cta ggg ctg gag
ccc tct gaa gag gag 3381Gly Gly Gly Asp Leu Thr Leu Gly Leu Glu Pro
Ser Glu Glu Glu1055 1060 1065gcc ccc agg tct cca ctg gca ccc tcc
gaa ggg gct ggc tcc gat 3426Ala Pro Arg Ser Pro Leu Ala Pro Ser Glu
Gly Ala Gly Ser Asp1070 1075 1080gta ttt gat ggt gac ctg gga atg
ggg gca gcc aag ggg ctg caa 3471Val Phe Asp Gly Asp Leu Gly Met Gly
Ala Ala Lys Gly Leu Gln1085 1090
1095agc ctc ccc aca cat gac ccc agc cct cta cag cgg tac agt gag
3516Ser Leu Pro Thr His Asp Pro Ser Pro Leu Gln Arg Tyr Ser Glu1100
1105 1110gac ccc aca gta ccc ctg ccc tct gag act gat ggc tac gtt
gcc 3561Asp Pro Thr Val Pro Leu Pro Ser Glu Thr Asp Gly Tyr Val
Ala1115 1120 1125ccc ctg acc tgc agc ccc cag cct gaa tat gtg aac
cag cca gat 3606Pro Leu Thr Cys Ser Pro Gln Pro Glu Tyr Val Asn Gln
Pro Asp1130 1135 1140gtt cgg ccc cag ccc cct tcg ccc cga gag ggc
cct ctg cct gct 3651Val Arg Pro Gln Pro Pro Ser Pro Arg Glu Gly Pro
Leu Pro Ala1145 1150 1155gcc cga cct gct ggt gcc act ctg gaa agg
ccc aag act ctc tcc 3696Ala Arg Pro Ala Gly Ala Thr Leu Glu Arg Pro
Lys Thr Leu Ser1160 1165 1170cca ggg aag aat ggg gtc gtc aaa gac
gtt ttt gcc ttt ggg ggt 3741Pro Gly Lys Asn Gly Val Val Lys Asp Val
Phe Ala Phe Gly Gly1175 1180 1185gcc gtg gag aac ccc gag tac ttg
aca ccc cag gga gga gct gcc 3786Ala Val Glu Asn Pro Glu Tyr Leu Thr
Pro Gln Gly Gly Ala Ala1190 1195 1200cct cag ccc cac cct cct cct
gcc ttc agc cca gcc ttc gac aac 3831Pro Gln Pro His Pro Pro Pro Ala
Phe Ser Pro Ala Phe Asp Asn1205 1210 1215ctc tat tac tgg gac cag
gac cca cca gag cgg ggg gct cca ccc 3876Leu Tyr Tyr Trp Asp Gln Asp
Pro Pro Glu Arg Gly Ala Pro Pro1220 1225 1230agc acc ttc aaa ggg
aca cct acg gca gag aac cca gag tac ctg 3921Ser Thr Phe Lys Gly Thr
Pro Thr Ala Glu Asn Pro Glu Tyr Leu1235 1240 1245ggt ctg gac gtg
cca gtg tga accagaaggc caagtccgca gaagccctga 3972Gly Leu Asp Val
Pro Val1250 1255tgtgtcctca gggagcaggg aaggcctgac ttctgctggc
atcaagaggt gggagggccc 4032tccgaccact tccaggggaa cctgccatgc
caggaacctg tcctaaggaa ccttccttcc 4092tgcttgagtt cccagatggc
tggaaggggt ccagcctcgt tggaagagga acagcactgg 4152ggagtctttg
tggattctga ggccctgccc aatgagactc tagggtccag tggatgccac
4212agcccagctt ggccctttcc ttccagatcc tgggtactga aagccttagg
gaagctggcc 4272tgagagggga agcggcccta agggagtgtc taagaacaaa
agcgacccat tcagagactg 4332tccctgaaac ctagtactgc cccccatgag
gaaggaacag caatggtgtc agtatccagg 4392ctttgtacag agtgcttttc
tgtttagttt ttactttttt tgttttgttt ttttaaagat 4452gaaataaaga
cccaggggga g 4473241255PRTHomo sapiens 24Met Glu Leu Ala Ala Leu
Cys Arg Trp Gly Leu Leu Leu Ala Leu Leu1 5 10 15Pro Pro Gly Ala Ala
Ser Thr Gln Val Cys Thr Gly Thr Asp Met Lys 20 25 30Leu Arg Leu Pro
Ala Ser Pro Glu Thr His Leu Asp Met Leu Arg His 35 40 45Leu Tyr Gln
Gly Cys Gln Val Val Gln Gly Asn Leu Glu Leu Thr Tyr 50 55 60Leu Pro
Thr Asn Ala Ser Leu Ser Phe Leu Gln Asp Ile Gln Glu Val65 70 75
80Gln Gly Tyr Val Leu Ile Ala His Asn Gln Val Arg Gln Val Pro Leu
85 90 95Gln Arg Leu Arg Ile Val Arg Gly Thr Gln Leu Phe Glu Asp Asn
Tyr 100 105 110Ala Leu Ala Val Leu Asp Asn Gly Asp Pro Leu Asn Asn
Thr Thr Pro 115 120 125Val Thr Gly Ala Ser Pro Gly Gly Leu Arg Glu
Leu Gln Leu Arg Ser 130 135 140Leu Thr Glu Ile Leu Lys Gly Gly Val
Leu Ile Gln Arg Asn Pro Gln145 150 155 160Leu Cys Tyr Gln Asp Thr
Ile Leu Trp Lys Asp Ile Phe His Lys Asn 165 170 175Asn Gln Leu Ala
Leu Thr Leu Ile Asp Thr Asn Arg Ser Arg Ala Cys 180 185 190His Pro
Cys Ser Pro Met Cys Lys Gly Ser Arg Cys Trp Gly Glu Ser 195 200
205Ser Glu Asp Cys Gln Ser Leu Thr Arg Thr Val Cys Ala Gly Gly Cys
210 215 220Ala Arg Cys Lys Gly Pro Leu Pro Thr Asp Cys Cys His Glu
Gln Cys225 230 235 240Ala Ala Gly Cys Thr Gly Pro Lys His Ser Asp
Cys Leu Ala Cys Leu 245 250 255His Phe Asn His Ser Gly Ile Cys Glu
Leu His Cys Pro Ala Leu Val 260 265 270Thr Tyr Asn Thr Asp Thr Phe
Glu Ser Met Pro Asn Pro Glu Gly Arg 275 280 285Tyr Thr Phe Gly Ala
Ser Cys Val Thr Ala Cys Pro Tyr Asn Tyr Leu 290 295 300Ser Thr Asp
Val Gly Ser Cys Thr Leu Val Cys Pro Leu His Asn Gln305 310 315
320Glu Val Thr Ala Glu Asp Gly Thr Gln Arg Cys Glu Lys Cys Ser Lys
325 330 335Pro Cys Ala Arg Val Cys Tyr Gly Leu Gly Met Glu His Leu
Arg Glu 340 345 350Val Arg Ala Val Thr Ser Ala Asn Ile Gln Glu Phe
Ala Gly Cys Lys 355 360 365Lys Ile Phe Gly Ser Leu Ala Phe Leu Pro
Glu Ser Phe Asp Gly Asp 370 375 380Pro Ala Ser Asn Thr Ala Pro Leu
Gln Pro Glu Gln Leu Gln Val Phe385 390 395 400Glu Thr Leu Glu Glu
Ile Thr Gly Tyr Leu Tyr Ile Ser Ala Trp Pro 405 410 415Asp Ser Leu
Pro Asp Leu Ser Val Phe Gln Asn Leu Gln Val Ile Arg 420 425 430Gly
Arg Ile Leu His Asn Gly Ala Tyr Ser Leu Thr Leu Gln Gly Leu 435 440
445Gly Ile Ser Trp Leu Gly Leu Arg Ser Leu Arg Glu Leu Gly Ser Gly
450 455 460Leu Ala Leu Ile His His Asn Thr His Leu Cys Phe Val His
Thr Val465 470 475 480Pro Trp Asp Gln Leu Phe Arg Asn Pro His Gln
Ala Leu Leu His Thr 485 490 495Ala Asn Arg Pro Glu Asp Glu Cys Val
Gly Glu Gly Leu Ala Cys His 500 505 510Gln Leu Cys Ala Arg Gly His
Cys Trp Gly Pro Gly Pro Thr Gln Cys 515 520 525Val Asn Cys Ser Gln
Phe Leu Arg Gly Gln Glu Cys Val Glu Glu Cys 530 535 540Arg Val Leu
Gln Gly Leu Pro Arg Glu Tyr Val Asn Ala Arg His Cys545 550 555
560Leu Pro Cys His Pro Glu Cys Gln Pro Gln Asn Gly Ser Val Thr Cys
565 570 575Phe Gly Pro Glu Ala Asp Gln Cys Val Ala Cys Ala His Tyr
Lys Asp 580 585 590Pro Pro Phe Cys Val Ala Arg Cys Pro Ser Gly Val
Lys Pro Asp Leu 595 600 605Ser Tyr Met Pro Ile Trp Lys Phe Pro Asp
Glu Glu Gly Ala Cys Gln 610 615 620Pro Cys Pro Ile Asn Cys Thr His
Ser Cys Val Asp Leu Asp Asp Lys625 630 635 640Gly Cys Pro Ala Glu
Gln Arg Ala Ser Pro Leu Thr Ser Ile Ile Ser 645 650 655Ala Val Val
Gly Ile Leu Leu Val Val Val Leu Gly Val Val Phe Gly 660 665 670Ile
Leu Ile Lys Arg Arg Gln Gln Lys Ile Arg Lys Tyr Thr Met Arg 675 680
685Arg Leu Leu Gln Glu Thr Glu Leu Val Glu Pro Leu Thr Pro Ser Gly
690 695 700Ala Met Pro Asn Gln Ala Gln Met Arg Ile Leu Lys Glu Thr
Glu Leu705 710 715 720Arg Lys Val Lys Val Leu Gly Ser Gly Ala Phe
Gly Thr Val Tyr Lys 725 730 735Gly Ile Trp Ile Pro Asp Gly Glu Asn
Val Lys Ile Pro Val Ala Ile 740 745 750Lys Val Leu Arg Glu Asn Thr
Ser Pro Lys Ala Asn Lys Glu Ile Leu 755 760 765Asp Glu Ala Tyr Val
Met Ala Gly Val Gly Ser Pro Tyr Val Ser Arg 770 775 780Leu Leu Gly
Ile Cys Leu Thr Ser Thr Val Gln Leu Val Thr Gln Leu785 790 795
800Met Pro Tyr Gly Cys Leu Leu Asp His Val Arg Glu Asn Arg Gly Arg
805 810 815Leu Gly Ser Gln Asp Leu Leu Asn Trp Cys Met Gln Ile Ala
Lys Gly 820 825 830Met Ser Tyr Leu Glu Asp Val Arg Leu Val His Arg
Asp Leu Ala Ala 835 840 845Arg Asn Val Leu Val Lys Ser Pro Asn His
Val Lys Ile Thr Asp Phe 850 855 860Gly Leu Ala Arg Leu Leu Asp Ile
Asp Glu Thr Glu Tyr His Ala Asp865 870 875 880Gly Gly Lys Val Pro
Ile Lys Trp Met Ala Leu Glu Ser Ile Leu Arg 885 890 895Arg Arg Phe
Thr His Gln Ser Asp Val Trp Ser Tyr Gly Val Thr Val 900 905 910Trp
Glu Leu Met Thr Phe Gly Ala Lys Pro Tyr Asp Gly Ile Pro Ala 915 920
925Arg Glu Ile Pro Asp Leu Leu Glu Lys Gly Glu Arg Leu Pro Gln Pro
930 935 940Pro Ile Cys Thr Ile Asp Val Tyr Met Ile Met Val Lys Cys
Trp Met945 950 955 960Ile Asp Ser Glu Cys Arg Pro Arg Phe Arg Glu
Leu Val Ser Glu Phe 965 970 975Ser Arg Met Ala Arg Asp Pro Gln Arg
Phe Val Val Ile Gln Asn Glu 980 985 990Asp Leu Gly Pro Ala Ser Pro
Leu Asp Ser Thr Phe Tyr Arg Ser Leu 995 1000 1005Leu Glu Asp Asp
Asp Met Gly Asp Leu Val Asp Ala Glu Glu Tyr 1010 1015 1020Leu Val
Pro Gln Gln Gly Phe Phe Cys Pro Asp Pro Ala Pro Gly 1025 1030
1035Ala Gly Gly Met Val His His Arg His Arg Ser Ser Ser Thr Arg
1040 1045 1050Ser Gly Gly Gly Asp Leu Thr Leu Gly Leu Glu Pro Ser
Glu Glu 1055 1060 1065Glu Ala Pro Arg Ser Pro Leu Ala Pro Ser Glu
Gly Ala Gly Ser 1070 1075 1080Asp Val Phe Asp Gly Asp Leu Gly Met
Gly Ala Ala Lys Gly Leu 1085 1090 1095Gln Ser Leu Pro Thr His Asp
Pro Ser Pro Leu Gln Arg Tyr Ser 1100 1105 1110Glu Asp Pro Thr Val
Pro Leu Pro Ser Glu Thr Asp Gly Tyr Val 1115 1120 1125Ala Pro Leu
Thr Cys Ser Pro Gln Pro Glu Tyr Val Asn Gln Pro 1130 1135 1140Asp
Val Arg Pro Gln Pro Pro Ser Pro Arg Glu Gly Pro Leu Pro 1145 1150
1155Ala Ala Arg Pro Ala Gly Ala Thr Leu Glu Arg Pro Lys Thr Leu
1160 1165 1170Ser Pro Gly Lys Asn Gly Val Val Lys Asp Val Phe Ala
Phe Gly 1175 1180 1185Gly Ala Val Glu Asn Pro Glu Tyr Leu Thr Pro
Gln Gly Gly Ala 1190 1195 1200Ala Pro Gln Pro His Pro Pro Pro Ala
Phe Ser Pro Ala Phe Asp 1205 1210 1215Asn Leu Tyr Tyr Trp Asp Gln
Asp Pro Pro Glu Arg Gly Ala Pro 1220 1225 1230Pro Ser Thr Phe Lys
Gly Thr Pro Thr Ala Glu Asn Pro Glu Tyr 1235 1240 1245Leu Gly Leu
Asp Val Pro Val 1250 1255251212DNAHomo sapiensCDS(1)..(1212) 25atg
aca gcc atc atc aaa gag atc gtt agc aga aac aaa agg aga tat 48Met
Thr Ala Ile Ile Lys Glu Ile Val Ser Arg Asn Lys Arg Arg Tyr1 5 10
15caa gag gat gga ttc gac tta gac ttg acc tat att tat cca aac att
96Gln Glu Asp Gly Phe Asp Leu Asp Leu Thr Tyr Ile Tyr Pro Asn Ile
20 25 30att gct atg gga ttt cct gca gaa aga ctt gaa ggc gta tac agg
aac 144Ile Ala Met Gly Phe Pro Ala Glu Arg Leu Glu Gly Val Tyr Arg
Asn 35 40 45aat att gat gat gta gta agg ttt ttg gat tca aag cat aaa
aac cat 192Asn Ile Asp Asp Val Val Arg Phe Leu Asp Ser Lys His Lys
Asn His 50 55 60tac aag ata tac aat ctt tgt gct gaa aga cat tat gac
acc gcc aaa 240Tyr Lys Ile Tyr Asn Leu Cys Ala Glu Arg His Tyr Asp
Thr Ala Lys65 70 75 80ttt aat tgc aga gtt gca caa tat cct ttt gaa
gac cat aac cca cca 288Phe Asn Cys Arg Val Ala Gln Tyr Pro Phe Glu
Asp His Asn Pro Pro 85 90 95cag cta gaa ctt atc aaa ccc ttt tgt gaa
gat ctt gac caa tgg cta 336Gln Leu Glu Leu Ile Lys Pro Phe Cys Glu
Asp Leu Asp Gln Trp Leu 100 105 110agt gaa gat gac aat cat gtt gca
gca att cac tgt aaa gct gga aag 384Ser Glu Asp Asp Asn His Val Ala
Ala Ile His Cys Lys Ala Gly Lys 115 120 125gga cga act ggt gta atg
ata tgt gca tat tta tta cat cgg ggc aaa 432Gly Arg Thr Gly Val Met
Ile Cys Ala Tyr Leu Leu His Arg Gly Lys 130 135 140ttt tta aag gca
caa gag gcc cta gat ttc tat ggg gaa gta agg acc 480Phe Leu Lys Ala
Gln Glu Ala Leu Asp Phe Tyr Gly Glu Val Arg Thr145 150 155 160aga
gac aaa aag gga gta act att ccc agt cag agg cgc tat gtg tat 528Arg
Asp Lys Lys Gly Val Thr Ile Pro Ser Gln Arg Arg Tyr Val Tyr 165 170
175tat tat agc tac ctg tta aag aat cat ctg gat tat aga cca gtg gca
576Tyr Tyr Ser Tyr Leu Leu Lys Asn His Leu Asp Tyr Arg Pro Val Ala
180 185 190ctg ttg ttt cac aag atg atg ttt gaa act att cca atg ttc
agt ggc 624Leu Leu Phe His Lys Met Met Phe Glu Thr Ile Pro Met Phe
Ser Gly 195 200 205gga act tgc aat cct cag ttt gtg gtc tgc cag cta
aag gtg aag ata 672Gly Thr Cys Asn Pro Gln Phe Val Val Cys Gln Leu
Lys Val Lys Ile 210 215 220tat tcc tcc aat tca gga ccc aca cga cgg
gaa gac aag ttc atg tac 720Tyr Ser Ser Asn Ser Gly Pro Thr Arg Arg
Glu Asp Lys Phe Met Tyr225 230 235 240ttt gag ttc cct cag ccg tta
cct gtg tgt ggt gat atc aaa gta gag 768Phe Glu Phe Pro Gln Pro Leu
Pro Val Cys Gly Asp Ile Lys Val Glu 245 250 255ttc ttc cac aaa cag
aac aag atg cta aaa aag gac aaa atg ttt cac 816Phe Phe His Lys Gln
Asn Lys Met Leu Lys Lys Asp Lys Met Phe His 260 265 270ttt tgg gta
aat aca ttc ttc ata cca gga cca gag gaa acc tca gaa 864Phe Trp Val
Asn Thr Phe Phe Ile Pro Gly Pro Glu Glu Thr Ser Glu 275 280 285aaa
gta gaa aat gga agt cta tgt gat caa gaa atc gat agc att tgc 912Lys
Val Glu Asn Gly Ser Leu Cys Asp Gln Glu Ile Asp Ser Ile Cys 290 295
300agt ata gag cgt gca gat aat gac aag gaa tat cta gta ctt act tta
960Ser Ile Glu Arg Ala Asp Asn Asp Lys Glu Tyr Leu Val Leu Thr
Leu305 310 315 320aca aaa aat gat ctt gac aaa gca aat aaa gac aaa
gcc aac cga tac 1008Thr Lys Asn Asp Leu Asp Lys Ala Asn Lys Asp Lys
Ala Asn Arg Tyr 325 330 335ttt tct cca aat ttt aag gtg aag ctg tac
ttc aca aaa aca gta gag 1056Phe Ser Pro Asn Phe Lys Val Lys Leu Tyr
Phe Thr Lys Thr Val Glu 340 345 350gag ccg tca aat cca gag gct agc
agt tca act tct gta aca cca gat 1104Glu Pro Ser Asn Pro Glu Ala Ser
Ser Ser Thr Ser Val Thr Pro Asp 355 360 365gtt agt gac aat gaa cct
gat cat tat aga tat tct gac acc act gac 1152Val Ser Asp Asn Glu Pro
Asp His Tyr Arg Tyr Ser Asp Thr Thr Asp 370 375 380tct gat cca gag
aat gaa cct ttt gat gaa gat cag cat aca caa att 1200Ser Asp Pro Glu
Asn Glu Pro Phe Asp Glu Asp Gln His Thr Gln Ile385 390 395 400aca
aaa gtc tga 1212Thr Lys Val26403PRTHomo sapiens 26Met Thr Ala Ile
Ile Lys Glu Ile Val Ser Arg Asn Lys Arg Arg Tyr1 5 10 15Gln Glu Asp
Gly Phe Asp Leu Asp Leu Thr Tyr Ile Tyr Pro Asn Ile 20 25 30Ile Ala
Met Gly Phe Pro Ala Glu Arg Leu Glu Gly Val Tyr Arg Asn 35 40 45Asn
Ile Asp Asp Val Val Arg Phe Leu Asp Ser Lys His Lys Asn His 50 55
60Tyr Lys Ile Tyr Asn Leu Cys Ala Glu Arg His Tyr Asp Thr Ala Lys65
70 75 80Phe Asn Cys Arg Val Ala Gln Tyr Pro Phe Glu Asp His Asn Pro
Pro 85 90 95Gln Leu Glu Leu Ile Lys Pro Phe Cys Glu Asp Leu Asp Gln
Trp Leu 100 105 110Ser Glu Asp Asp Asn His Val Ala Ala Ile His Cys
Lys Ala Gly Lys 115 120 125Gly Arg Thr Gly Val Met Ile Cys Ala Tyr
Leu Leu His Arg Gly Lys 130 135 140Phe Leu Lys Ala Gln Glu Ala Leu
Asp Phe Tyr Gly Glu Val Arg Thr145 150 155 160Arg Asp Lys Lys Gly
Val Thr Ile Pro Ser Gln Arg Arg Tyr Val Tyr 165 170 175Tyr Tyr Ser
Tyr Leu Leu Lys Asn His Leu Asp Tyr Arg Pro Val Ala 180 185 190Leu
Leu Phe His Lys
Met Met Phe Glu Thr Ile Pro Met Phe Ser Gly 195 200 205Gly Thr Cys
Asn Pro Gln Phe Val Val Cys Gln Leu Lys Val Lys Ile 210 215 220Tyr
Ser Ser Asn Ser Gly Pro Thr Arg Arg Glu Asp Lys Phe Met Tyr225 230
235 240Phe Glu Phe Pro Gln Pro Leu Pro Val Cys Gly Asp Ile Lys Val
Glu 245 250 255Phe Phe His Lys Gln Asn Lys Met Leu Lys Lys Asp Lys
Met Phe His 260 265 270Phe Trp Val Asn Thr Phe Phe Ile Pro Gly Pro
Glu Glu Thr Ser Glu 275 280 285Lys Val Glu Asn Gly Ser Leu Cys Asp
Gln Glu Ile Asp Ser Ile Cys 290 295 300Ser Ile Glu Arg Ala Asp Asn
Asp Lys Glu Tyr Leu Val Leu Thr Leu305 310 315 320Thr Lys Asn Asp
Leu Asp Lys Ala Asn Lys Asp Lys Ala Asn Arg Tyr 325 330 335Phe Ser
Pro Asn Phe Lys Val Lys Leu Tyr Phe Thr Lys Thr Val Glu 340 345
350Glu Pro Ser Asn Pro Glu Ala Ser Ser Ser Thr Ser Val Thr Pro Asp
355 360 365Val Ser Asp Asn Glu Pro Asp His Tyr Arg Tyr Ser Asp Thr
Thr Asp 370 375 380Ser Asp Pro Glu Asn Glu Pro Phe Asp Glu Asp Gln
His Thr Gln Ile385 390 395 400Thr Lys Val27597DNAHomo
sapiensCDS(1)..(597) 27atg tca aac gtg cga gtg tct aac ggg agc cct
agc ctg gag cgg atg 48Met Ser Asn Val Arg Val Ser Asn Gly Ser Pro
Ser Leu Glu Arg Met1 5 10 15gac gcc agg cag gcg gag cac ccc aag ccc
tcg gcc tgc agg aac ctc 96Asp Ala Arg Gln Ala Glu His Pro Lys Pro
Ser Ala Cys Arg Asn Leu 20 25 30ttc ggc ccg gtg gac cac gaa gag tta
acc cgg gac ttg gag aag cac 144Phe Gly Pro Val Asp His Glu Glu Leu
Thr Arg Asp Leu Glu Lys His 35 40 45tgc aga gac atg gaa gag gcg agc
cag cgc aag tgg aat ttc gat ttt 192Cys Arg Asp Met Glu Glu Ala Ser
Gln Arg Lys Trp Asn Phe Asp Phe 50 55 60cag aat cac aaa ccc cta gag
ggc aag tac gag tgg caa gag gtg gag 240Gln Asn His Lys Pro Leu Glu
Gly Lys Tyr Glu Trp Gln Glu Val Glu65 70 75 80aag ggc agc ttg ccc
gag ttc tac tac aga ccc ccg cgg ccc ccc aaa 288Lys Gly Ser Leu Pro
Glu Phe Tyr Tyr Arg Pro Pro Arg Pro Pro Lys 85 90 95ggt gcc tgc aag
gtg ccg gcg cag gag agc cag gat gtc agc ggg agc 336Gly Ala Cys Lys
Val Pro Ala Gln Glu Ser Gln Asp Val Ser Gly Ser 100 105 110cgc ccg
gcg gcg cct tta att ggg gct ccg gct aac tct gag gac acg 384Arg Pro
Ala Ala Pro Leu Ile Gly Ala Pro Ala Asn Ser Glu Asp Thr 115 120
125cat ttg gtg gac cca aag act gat ccg tcg gac agc cag acg ggg tta
432His Leu Val Asp Pro Lys Thr Asp Pro Ser Asp Ser Gln Thr Gly Leu
130 135 140gcg gag caa tgc gca gga ata agg aag cga cct gca acc gac
gat tct 480Ala Glu Gln Cys Ala Gly Ile Arg Lys Arg Pro Ala Thr Asp
Asp Ser145 150 155 160tct act caa aac aaa aga gcc aac aga aca gaa
gaa aat gtt tca gac 528Ser Thr Gln Asn Lys Arg Ala Asn Arg Thr Glu
Glu Asn Val Ser Asp 165 170 175ggt tcc cca aat gcc ggt tct gtg gag
cag acg ccc aag aag cct ggc 576Gly Ser Pro Asn Ala Gly Ser Val Glu
Gln Thr Pro Lys Lys Pro Gly 180 185 190ctc aga aga cgt caa acg taa
597Leu Arg Arg Arg Gln Thr 19528198PRTHomo sapiens 28Met Ser Asn
Val Arg Val Ser Asn Gly Ser Pro Ser Leu Glu Arg Met1 5 10 15Asp Ala
Arg Gln Ala Glu His Pro Lys Pro Ser Ala Cys Arg Asn Leu 20 25 30Phe
Gly Pro Val Asp His Glu Glu Leu Thr Arg Asp Leu Glu Lys His 35 40
45Cys Arg Asp Met Glu Glu Ala Ser Gln Arg Lys Trp Asn Phe Asp Phe
50 55 60Gln Asn His Lys Pro Leu Glu Gly Lys Tyr Glu Trp Gln Glu Val
Glu65 70 75 80Lys Gly Ser Leu Pro Glu Phe Tyr Tyr Arg Pro Pro Arg
Pro Pro Lys 85 90 95Gly Ala Cys Lys Val Pro Ala Gln Glu Ser Gln Asp
Val Ser Gly Ser 100 105 110Arg Pro Ala Ala Pro Leu Ile Gly Ala Pro
Ala Asn Ser Glu Asp Thr 115 120 125His Leu Val Asp Pro Lys Thr Asp
Pro Ser Asp Ser Gln Thr Gly Leu 130 135 140Ala Glu Gln Cys Ala Gly
Ile Arg Lys Arg Pro Ala Thr Asp Asp Ser145 150 155 160Ser Thr Gln
Asn Lys Arg Ala Asn Arg Thr Glu Glu Asn Val Ser Asp 165 170 175Gly
Ser Pro Asn Ala Gly Ser Val Glu Gln Thr Pro Lys Lys Pro Gly 180 185
190Leu Arg Arg Arg Gln Thr 19529894DNAHomo sapiensCDS(1)..(894)
29atg aca aca ccc aga aat tca gta aat ggg act ttc ccg gca gag cca
48Met Thr Thr Pro Arg Asn Ser Val Asn Gly Thr Phe Pro Ala Glu Pro1
5 10 15atg aaa ggc cct att gct atg caa tct ggt cca aaa cca ctc ttc
agg 96Met Lys Gly Pro Ile Ala Met Gln Ser Gly Pro Lys Pro Leu Phe
Arg 20 25 30agg atg tct tca ctg gtg ggc ccc acg caa agc ttc ttc atg
agg gaa 144Arg Met Ser Ser Leu Val Gly Pro Thr Gln Ser Phe Phe Met
Arg Glu 35 40 45tct aag act ttg ggg gct gtc cag att atg aat ggg ctc
ttc cac att 192Ser Lys Thr Leu Gly Ala Val Gln Ile Met Asn Gly Leu
Phe His Ile 50 55 60gcc ctg ggg ggt ctt ctg atg atc cca gca ggg atc
tat gca ccc atc 240Ala Leu Gly Gly Leu Leu Met Ile Pro Ala Gly Ile
Tyr Ala Pro Ile65 70 75 80tgt gtg act gtg tgg tac cct ctc tgg gga
ggc att atg tat att att 288Cys Val Thr Val Trp Tyr Pro Leu Trp Gly
Gly Ile Met Tyr Ile Ile 85 90 95tcc gga tca ctc ttg gca gca acg gag
aaa aac tct agg aag tgt ttg 336Ser Gly Ser Leu Leu Ala Ala Thr Glu
Lys Asn Ser Arg Lys Cys Leu 100 105 110gtc aaa gga aaa atg ata atg
aat tca ttg agc ctc ttt gct gcc att 384Val Lys Gly Lys Met Ile Met
Asn Ser Leu Ser Leu Phe Ala Ala Ile 115 120 125tct gga atg att ctt
tca atc atg gac ata ctt aat att aaa att tcc 432Ser Gly Met Ile Leu
Ser Ile Met Asp Ile Leu Asn Ile Lys Ile Ser 130 135 140cat ttt tta
aaa atg gag agt ctg aat ttt att aga gct cac aca cca 480His Phe Leu
Lys Met Glu Ser Leu Asn Phe Ile Arg Ala His Thr Pro145 150 155
160tat att aac ata tac aac tgt gaa cca gct aat ccc tct gag aaa aac
528Tyr Ile Asn Ile Tyr Asn Cys Glu Pro Ala Asn Pro Ser Glu Lys Asn
165 170 175tcc cca tct acc caa tac tgt tac agc ata caa tct ctg ttc
ttg ggc 576Ser Pro Ser Thr Gln Tyr Cys Tyr Ser Ile Gln Ser Leu Phe
Leu Gly 180 185 190att ttg tca gtg atg ctg atc ttt gcc ttc ttc cag
gaa ctt gta ata 624Ile Leu Ser Val Met Leu Ile Phe Ala Phe Phe Gln
Glu Leu Val Ile 195 200 205gct ggc atc gtt gag aat gaa tgg aaa aga
acg tgc tcc aga ccc aaa 672Ala Gly Ile Val Glu Asn Glu Trp Lys Arg
Thr Cys Ser Arg Pro Lys 210 215 220tct aac ata gtt ctc ctg tca gca
gaa gaa aaa aaa gaa cag act att 720Ser Asn Ile Val Leu Leu Ser Ala
Glu Glu Lys Lys Glu Gln Thr Ile225 230 235 240gaa ata aaa gaa gaa
gtg gtt ggg cta act gaa aca tct tcc caa cca 768Glu Ile Lys Glu Glu
Val Val Gly Leu Thr Glu Thr Ser Ser Gln Pro 245 250 255aag aat gaa
gaa gac att gaa att att cca atc caa gaa gag gaa gaa 816Lys Asn Glu
Glu Asp Ile Glu Ile Ile Pro Ile Gln Glu Glu Glu Glu 260 265 270gaa
gaa aca gag acg aac ttt cca gaa cct ccc caa gat cag gaa tcc 864Glu
Glu Thr Glu Thr Asn Phe Pro Glu Pro Pro Gln Asp Gln Glu Ser 275 280
285tca cca ata gaa aat gac agc tct cct taa 894Ser Pro Ile Glu Asn
Asp Ser Ser Pro 290 29530297PRTHomo sapiens 30Met Thr Thr Pro Arg
Asn Ser Val Asn Gly Thr Phe Pro Ala Glu Pro1 5 10 15Met Lys Gly Pro
Ile Ala Met Gln Ser Gly Pro Lys Pro Leu Phe Arg 20 25 30Arg Met Ser
Ser Leu Val Gly Pro Thr Gln Ser Phe Phe Met Arg Glu 35 40 45Ser Lys
Thr Leu Gly Ala Val Gln Ile Met Asn Gly Leu Phe His Ile 50 55 60Ala
Leu Gly Gly Leu Leu Met Ile Pro Ala Gly Ile Tyr Ala Pro Ile65 70 75
80Cys Val Thr Val Trp Tyr Pro Leu Trp Gly Gly Ile Met Tyr Ile Ile
85 90 95Ser Gly Ser Leu Leu Ala Ala Thr Glu Lys Asn Ser Arg Lys Cys
Leu 100 105 110Val Lys Gly Lys Met Ile Met Asn Ser Leu Ser Leu Phe
Ala Ala Ile 115 120 125Ser Gly Met Ile Leu Ser Ile Met Asp Ile Leu
Asn Ile Lys Ile Ser 130 135 140His Phe Leu Lys Met Glu Ser Leu Asn
Phe Ile Arg Ala His Thr Pro145 150 155 160Tyr Ile Asn Ile Tyr Asn
Cys Glu Pro Ala Asn Pro Ser Glu Lys Asn 165 170 175Ser Pro Ser Thr
Gln Tyr Cys Tyr Ser Ile Gln Ser Leu Phe Leu Gly 180 185 190Ile Leu
Ser Val Met Leu Ile Phe Ala Phe Phe Gln Glu Leu Val Ile 195 200
205Ala Gly Ile Val Glu Asn Glu Trp Lys Arg Thr Cys Ser Arg Pro Lys
210 215 220Ser Asn Ile Val Leu Leu Ser Ala Glu Glu Lys Lys Glu Gln
Thr Ile225 230 235 240Glu Ile Lys Glu Glu Val Val Gly Leu Thr Glu
Thr Ser Ser Gln Pro 245 250 255Lys Asn Glu Glu Asp Ile Glu Ile Ile
Pro Ile Gln Glu Glu Glu Glu 260 265 270Glu Glu Thr Glu Thr Asn Phe
Pro Glu Pro Pro Gln Asp Gln Glu Ser 275 280 285Ser Pro Ile Glu Asn
Asp Ser Ser Pro 290 29531596DNAhomo sapiens 31acgcgtactg gagtcaatga
aagcaactat ttcaaaagat cagattactt accagtttca 60ctaataaaga tttattactt
taaaccttta tcataaaatg tatgctttga atactgtgaa 120gtacactgca
tataaggagt gtggtatagt ataaagaaac tttctgcagg tagtaattat
180agtgaagatt ttaggtttac aaagccctag ctgttttctg tgtagctttt
attattctta 240tgactcttga caagtttgta gcttcaccat atacatttaa
tattttgcaa taattggcct 300tgttcctgag ctgttggatt cggggccgta
gcactgtctg agaggtttac atttctcaca 360gtgaaccggt ctctttttca
gctgcttcct ggcttctttt tactcaggtt tccactgctt 420ttttgctttt
tttaatgctg tatgaaggtg ttaacatttg tttatatttt tcattaattg
480taataccttt aaatcatgca tcatactcag aaatagggat tagaatttaa
gtgacatctt 540tggcctaata taatttacct gttaaaaatt tgtgaaagct
attgcttagc ggccgc 59632511DNAHomo sapiens 32acgcgtccat gtccgtacct
ttctagttca taccttcttt taattttttt tttcttttca 60atttgaagag agtgcttcct
ctgttcttaa ggctagggaa ccaaattagg ttgtttcaat 120atcgtgctaa
aagatactgc ctttagaaga aggctattga caatccagcg tgtctcggtg
180gaactctgac tccatggttc actttcatga tggccacatg cctcctgccc
agagcccggc 240agccactgtg cagtgggaag gggggccgat acactgtacg
agagtgagta gcaggtctca 300cagtgaaccg gtctctttcc ctactgtgtc
acactcctaa tggaatgccg ttatccaaag 360agcagcacga acccgacagg
gctgagtggc ttgtgctagg gagaggtttg tgtcattcct 420gctgaccaaa
ctgcaggaaa aactgctaat tgtcatgctg aagactgcct gacggggaga
480ctctgccttc tgtaagtagg tcagcggccg c 51133203DNAHomo sapiens
33acgcgtaatt catatttgca tgtcgctatg tgttctggga aatcaccata aacgtgaaat
60gtctttggat ttgggaatct tataagttct gtatgagacc actcggatga gctgttggat
120tcggggccgt agcactgtct gagaggttta catttctcac agtgaaccgg
tctctttttc 180agctgcttct tttttgcggc cgc 20334205DNAHomo sapiens
34gcggccgcaa ttcatatttg catgtcgcta tgtgttctgg gaaatcacca taaacgtgaa
60atgtctttgg atttgggaat cttataagtt ctgtatgaga ccactcggat gagctgttgg
120attcggggcc gtagcactgt ctgagaggtt tacatttctc acagtgaacc
ggtctctttt 180tcagctgctt cttttttgcg gccgc 2053545DNAArtificial
SequenceDNA target sequence of the miR-142-3p 35gcggccgcgt
cgactccata aagtaggaaa cactacagcg gccgc 4536128DNAArtificial
SequenceDNA target sequence four time repeat miR-142- 3pT4X
36gcggccgcgt cgactccata aagtaggaaa cactacacga ttccataaag taggaaacac
60tacaaccggt tccataaagt aggaaacact acatcactcc ataaagtagg aaacactaca
120gcggccgc 128371131DNAherpes simplex virus 1 37atggcttcgt
accccggcca tcagcacgcg tctgcgttcg accaggctgc gcgttctcgc 60ggccatagca
accgacgtac ggcgttgcgc cctcgccggc agcaagaagc cacggaagtc
120cgcccggagc agaaaatgcc cacgctactg cgggtttata tagacggtcc
ccacgggatg 180gggaaaacca ccaccacgca actgctggtg gccctgggtt
cgcgcgacga tatcgtctac 240gtacccgagc cgatgactta ctggcaggtg
ctgggggctt ccgagacaat cgcgaacatc 300tacaccacac aacaccgcct
cgaccagggt gagatatcgg ccggggacgc ggcggtggta 360atgacaagcg
cccagataac aatgggcatg ccttatgccg tgaccgacgc cgttctggct
420cctcatatcg ggggggaggc tgggagctca catgccccgc ccccggccct
caccctcatc 480ttcgaccgcc atcccatcgc cgccctcctg tgttacccgg
ccgcgcgata ccttatgggc 540agcatgaccc cccaggccgt gctggcgttc
gtggccctca tcccgccgac cttgcccggc 600acaaacatcg tgttgggggc
ccttccggag gacagacaca tcgaccgcct ggccaaacgc 660cagcgccccg
gcgagcggct tgacctggct atgctggccg cgattcgccg cgtttacgag
720ctgcttgcca atacggtgcg gtatctgcag ggcggcgggt cgtggcggga
ggattgggga 780cagctttcgg ggacggccgt gccgccccag ggtgccgagc
cccagagcaa cgcgggccca 840cgaccccata tcggggacac gttatttacc
ctgtttcggg cccccgagtt gctggccccc 900aacggcgacc tgtataacgt
gtttgcctgg gccttggacg tcttggccaa acgcctccgt 960cccatgcacg
tctttatcct ggattacgac caatcgcccg ccggctaccg ggacgccctg
1020ctgcaactta cctccgggat ggtccagacc cacgtcacca cccccggctc
cataccgacg 1080atctgcgacc tggcgcgcac gtttgcccgg gagatggggg
aggctaacta a 113138499DNAHomo sapiens 38atgaaatata caagttatat
cttggctttt cagctctgca tcgttttggg ttctcttggc 60tgttactgcc aggaccatat
gtaaaagaag cagaaaacct taagaaatat tttaatgcag 120gtcattcaga
tgtagcggat aatggaactc ttttcttagg cattttgaag aattggaaag
180aggagagtga cagaaaaata atgcagagcc aaattgtctc cttttacttc
aaacttttta 240aaaactttaa agatgaccag agcatccaaa agagtgtgga
gaccatcaag gaagacatga 300atgtaagttt ttcaatagca acaaaaagaa
acgagatgac ttcgaaaagc tgactaatta 360ttcggtaact gacttgaatg
tccaacgcaa agcaatacat gaactcatcc aagtgatggc 420tgaactgtcg
ccagcagcta aaacagggaa gcgaaaaagg agtcagatgc tgtttcgagg
480tcgaagagca tcccagtaa 49939468DNAMus musculus 39atgaacgcta
cacactgcat cttggctttg cagctcttcc tcatggctgt ttctggctgt 60tactgccacg
gcacagtcat tgaaagccta gaaagtctga ataactattt taactcaagt
120ggcatagatg tggaagaaaa gagtctcttc ttggatatct ggaggaactg
gcaaaaggat 180ggtgacatga aaatcctgca gagccagatt atctctttct
acctcagact ctttgaagtc 240ttgaaagaca atcaggccat cagcaacaac
ataagcgtca ttgaatcaca cctgattact 300accttcttca gcaacagcaa
ggcgaaaaag gatgcattca tgagtattgc caagtttgag 360gtcaacaacc
cacaggtcca gcgccaagca ttcaatgagc tcatccgagt ggtccaccag
420ctgttgccgg aatccagcct caggaagcgg aaaaggagtc gctgctga
46840462DNAHomo sapiens 40atgtacagga tgcaactcct gtcttgcatt
gcactaagtc ttgcacttgt cacaaacagt 60gcacctactt caagttctac aaagaaaaca
cagctacaac tggagcattt actgctggat 120ttacagatga ttttgaatgg
aattaataat tacaagaatc ccaaactcac caggatgctc 180acatttaagt
tttacatgcc caagaaggcc acagaactga aacatcttca gtgtctagaa
240gaagaactca aacctctgga ggaagtgcta aatttagctc aaagcaaaaa
ctttcactta 300agacccaggg acttaatcag caatatcaac gtaatagttc
tggaactaaa gggatctgaa 360acaacattca tgtgtgaata tgctgatgag
acagcaacca ttgtagaatt tctgaacaga 420tggattacct tttgtcaaag
catcatctca acactgactt ga 4624123RNAArtificial
SequenceOligonucleotide primer 41uguaguguuu ccuacuuuau gga
234223RNAArtificial SequenceOligonucleotide primer 42uguaguguuu
ccuacuuuau gga 2343546DNAArtificial SequenceYeast cytosine
deaminase with cloning sites 43aacacgatta taaatggtga cagggggaat
ggcaagcaag tgggatcaga agggtatgga 60cattgcctat gaggaggcgg ccttaggtta
caaagagggt ggtgttccta ttggcggatg 120tcttatcaat aacaaagacg
gaagtgttct cggtcgtggt cacaacatga gatttcaaaa 180gggatccgcc
acactacatg gtgagatctc cactttggaa aactgtggga gattagaggg
240caaagtgtac aaagatacca ctttgtatac gacgctgtct ccatgcgaca
tgtgtacagg 300tgccatcatc atgtatggta ttccacgctg tgttgtcggt
gagaacgtta atttcaaaag 360taagggcgag aaatatttac aaactagagg
tcacgaggtt gttgttgttg acgatgagag 420gtgtaaaaag atcatgaaac
aatttatcga tgaaagacct caggattggt ttgaagatat 480tggtgagtag
gcggccgcgc catagataaa ataaaagatt ttatttagtc tccagaaaaa 540gggggg
54644537DNAArtificial SequenceCD codon optimized with cloning sites
44ttataaatgg tgaccggcgg catggcctcc aagtgggatc aaaagggcat
ggatatcgct
60tacgaggagg ccgccctggg ctacaaggag ggcggcgtgc ctatcggcgg ctgtctgatc
120aacaacaagg acggcagtgt gctgggcagg ggccacaaca tgaggttcca
gaagggctcc 180gccaccctgc acggcgagat ctccaccctg gagaactgtg
gcaggctgga gggcaaggtg 240tacaaggaca ccaccctgta caccaccctg
tccccttgtg acatgtgtac cggcgctatc 300atcatgtacg gcatccctag
gtgtgtggtg ggcgagaacg tgaacttcaa gtccaagggc 360gagaagtacc
tgcaaaccag gggccacgag gtggtggttg ttgacgatga gaggtgtaag
420aagatcatga agcagttcat cgacgagagg cctcaggact ggttcgagga
tatcggcgag 480tgataagcgg ccgcagataa aataaaagat tttatttagt
ctccagaaaa agggggg 5374540DNAArtificial SequenceOligonucleotide
primer 45tcgaggatat cggcgagtga aacccgttat tctttttggc
404640DNAArtificial SequenceOligonucleotide Primer 46gccaaaaaga
ataacgggtt tcactcgccg atatcctcga 404782DNAArtificial
SequenceOligonucleotide primer 47tcggcgagtg atccggcggc ggcgcctccg
gcggcggcgc ctccggcggc ggcgcctccg 60gcggcggcgc caacccgtta tt
824882DNAArtificial SequenceOligonucleotide Primer 48aataacgggt
tggcgccgcc gccggaggcg ccgccgccgg aggcgccgcc gccggaggcg 60ccgccgccgg
atcactcgcc ga 8249480DNAArtificial SequenceCytosine deaminase codon
optimized heat stabilized 49atggtgaccg gcggcatggc ctccaagtgg
gatcaaaagg gcatggatat cgcttacgag 60gaggccctgc tgggctacaa ggagggcggc
gtgcctatcg gcggctgtct gatcaacaac 120aaggacggca gtgtgctggg
caggggccac aacatgaggt tccagaaggg ctccgccacc 180ctgcacggcg
agatctccac cctggagaac tgtggcaggc tggagggcaa ggtgtacaag
240gacaccaccc tgtacaccac cctgtcccct tgtgacatgt gtaccggcgc
tatcatcatg 300tacggcatcc ctaggtgtgt gatcggcgag aacgtgaact
tcaagtccaa gggcgagaag 360tacctgcaaa ccaggggcca cgaggtggtg
gttgttgacg atgagaggtg taagaagctg 420atgaagcagt tcatcgacga
gaggcctcag gactggttcg aggatatcgg cgagtgataa 48050546DNAArtificial
SequenceCytosine deaminase codon optimized heat stabilized
50aacacgatta taaatggtga ccggcggcat ggcctccaag tgggatcaaa agggcatgga
60tatcgcttac gaggaggccc tgctgggcta caaggagggc ggcgtgccta tcggcggctg
120tctgatcaac aacaaggacg gcagtgtgct gggcaggggc cacaacatga
ggttccagaa 180gggctccgcc accctgcacg gcgagatctc caccctggag
aactgtggca ggctggaggg 240caaggtgtac aaggacacca ccctgtacac
caccctgtcc ccttgtgaca tgtgtaccgg 300cgctatcatc atgtacggca
tccctaggtg tgtgatcggc gagaacgtga acttcaagtc 360caagggcgag
aagtacctgc aaaccagggg ccacgaggtg gtggttgttg acgatgagag
420gtgtaagaag ctgatgaagc agttcatcga cgagaggcct caggactggt
tcgaggatat 480cggcgagtaa gcggccgcgc catagataaa ataaaagatt
ttatttagtc tccagaaaaa 540gggggg 5465140DNAArtificial
SequenceOligonucleotide primer 51tcgaggatat cggcgagtga aacccgttat
tctttttggc 405240DNAArtificial SequenceOligonucleotide Primer
52gccaaaaaga ataacgggtt tcactcgccg atatcctcga 405352DNAArtificial
SequenceOligonucleotide primer complement 53ccaagctcct atagccgctc
actatctact tgggcaataa gaaaaaccga ag 525452DNAArtificial
SequenceOligonucleotide primer complement 54ggttcgagga tatcggcgag
tgatagatga acccgttatt ctttttggct tc 52551296DNAArtificial
SequenceCD-UPRT cloning sequence 55aacacgatta taaatggtga ccggcggcat
ggcctccaag tgggatcaaa agggcatgga 60tatcgcttac gaggaggccc tgctgggcta
caaggagggc ggcgtgccta tcggcggctg 120tctgatcaac aacaaggacg
gcagtgtgct gggcaggggc cacaacatga ggttccagaa 180gggctccgcc
accctgcacg gcgagatctc caccctggag aactgtggca ggctggaggg
240caaggtgtac aaggacacca ccctgtacac caccctgtcc ccttgtgaca
tgtgtaccgg 300cgctatcatc atgtacggca tccctaggtg tgtgatcggc
gagaacgtga acttcaagtc 360caagggcgag aagtacctgc aaaccagggg
ccacgaggtg gtggttgttg acgatgagag 420gtgtaagaag ctgatgaagc
agttcatcga cgagaggcct caggactggt tcgaggatat 480cggcgagaac
ccgttattct ttttggcttc tccattcttg taccttacat atcttatata
540ttatccaaac aaagggtctt tcgttagcaa acctagaaat ctgcaaaaaa
tgtcttcgga 600accatttaag aacgtctact tgctacctca aacaaaccaa
ttgctgggtt tgtacaccat 660catcagaaat aagaatacaa ctagacctga
tttcattttc tactccgata gaatcatcag 720attgttggtt gaagaaggtt
tgaaccatct acctgtgcaa aagcaaattg tggaaactga 780caccaacgaa
aacttcgaag gtgtctcatt catgggtaaa atctgtggtg tttccattgt
840cagagctggt gaatcgatgg agcaaggatt aagagactgt tgtaggtctg
tgcgtatcgg 900taaaatttta attcaaaggg acgaggagac tgctttacca
aagttattct acgaaaaatt 960accagaggat atatctgaaa ggtatgtctt
cctattagac ccaatgctgg ccaccggtgg 1020tagtgctatc atggctacag
aagtcttgat taagagaggt gttaagccag agagaattta 1080cttcttaaac
ctaatctgta gtaaggaagg gattgaaaaa taccatgccg ccttcccaga
1140ggtcagaatt gttactggtg ccctcgacag aggtctagat gaaaacaagt
atctagttcc 1200agggttgggt gactttggtg acagatacta ctgtgtttaa
gcggccgcgc catagataaa 1260ataaaagatt ttatttagtc tccagaaaaa gggggg
1296565PRTArtificial SequencePeptide linker sequence 56Ser Gly Gly
Gly Gly1 55782DNAArtificial SequenceOligonucleotide primer
57tcggcgagtg atccggcggc ggcgcctccg gcggcggcgc ctccggcggc ggcgcctccg
60gcggcggcgc caacccgtta tt 825882DNAArtificial
SequenceOligonucleotide primer 58aataacgggt tggcgccgcc gccggaggcg
ccgccgccgg aggcgccgcc gccggaggcg 60ccgccgccgg atcactcgcc ga
82591356DNAArtificial SequenceCD codon optimized heat stabilized
linker UPRT 59aacacgatta taaatggtga ccggcggcat ggcctccaag
tgggatcaaa agggcatgga 60tatcgcttac gaggaggccc tgctgggcta caaggagggc
ggcgtgccta tcggcggctg 120tctgatcaac aacaaggacg gcagtgtgct
gggcaggggc cacaacatga ggttccagaa 180gggctccgcc accctgcacg
gcgagatctc caccctggag aactgtggca ggctggaggg 240caaggtgtac
aaggacacca ccctgtacac caccctgtcc ccttgtgaca tgtgtaccgg
300cgctatcatc atgtacggca tccctaggtg tgtgatcggc gagaacgtga
acttcaagtc 360caagggcgag aagtacctgc aaaccagggg ccacgaggtg
gtggttgttg acgatgagag 420gtgtaagaag ctgatgaagc agttcatcga
cgagaggcct caggactggt tcgaggatat 480cggcgagtcc ggcggcggcg
cctccggcgg cggcgcctcc ggcggcggcg cctccggcgg 540cggcgccaac
ccgttattct ttttggcttc tccattcttg taccttacat atcttatata
600ttatccaaac aaagggtctt tcgttagcaa acctagaaat ctgcaaaaaa
tgtcttcgga 660accatttaag aacgtctact tgctacctca aacaaaccaa
ttgctgggtt tgtacaccat 720catcagaaat aagaatacaa ctagacctga
tttcattttc tactccgata gaatcatcag 780attgttggtt gaagaaggtt
tgaaccatct acctgtgcaa aagcaaattg tggaaactga 840caccaacgaa
aacttcgaag gtgtctcatt catgggtaaa atctgtggtg tttccattgt
900cagagctggt gaatcgatgg agcaaggatt aagagactgt tgtaggtctg
tgcgtatcgg 960taaaatttta attcaaaggg acgaggagac tgctttacca
aagttattct acgaaaaatt 1020accagaggat atatctgaaa ggtatgtctt
cctattagac ccaatgctgg ccaccggtgg 1080tagtgctatc atggctacag
aagtcttgat taagagaggt gttaagccag agagaattta 1140cttcttaaac
ctaatctgta gtaaggaagg gattgaaaaa taccatgccg ccttcccaga
1200ggtcagaatt gttactggtg ccctcgacag aggtctagat gaaaacaagt
atctagttcc 1260agggttgggt gactttggtg acagatacta ctgtgtttaa
gcggccgcgc catagataaa 1320ataaaagatt ttatttagtc tccagaaaaa gggggg
1356601269DNAArtificial SequenceCD codon optimized heat stabilized
OPRT 60aacacgatta taaatggtga ccggcggcat ggcctccaag tgggatcaaa
agggcatgga 60tatcgcttac gaggaggccc tgctgggcta caaggagggc ggcgtgccta
tcggcggctg 120tctgatcaac aacaaggacg gcagtgtgct gggcaggggc
cacaacatga ggttccagaa 180gggctccgcc accctgcacg gcgagatctc
caccctggag aactgtggca ggctggaggg 240caaggtgtac aaggacacca
ccctgtacac caccctgtcc ccttgtgaca tgtgtaccgg 300cgctatcatc
atgtacggca tccctaggtg tgtgatcggc gagaacgtga acttcaagtc
360caagggcgag aagtacctgc aaaccagggg ccacgaggtg gtggttgttg
acgatgagag 420gtgtaagaag ctgatgaagc agttcatcga cgagaggcct
caggactggt tcgaggatat 480cggcgaggcg gtcgctcgtg cagctttggg
gccattggtg acgggtctgt acgacgtgca 540ggctttcaag tttggggact
tcgtgctgaa gagcgggctt tcctccccca tctacatcga 600tctgcggggc
atcgtgtctc gaccgcgtct tctgagtcag gttgcagata ttttattcca
660aactgcccaa aatgcaggca tcagttttga caccgtgtgt ggagtgcctt
atacagcttt 720gccattggct acagttatct gttcaaccaa tcaaattcca
atgcttatta gaaggaaaga 780aacaaaggat tatggaacta agcgtcttgt
agaaggaact attaatccag gagaaacctg 840tttaatcatt gaagatgttg
tcaccagtgg atctagtgtt ttggaaactg ttgaggttct 900tcagaaggag
ggcttgaagg tcactgatgc catagtgctg ttggacagag agcagggagg
960caaggacaag ttgcaggcgc acgggatccg cctccactca gtgtgtacat
tgtccaaaat 1020gctggagatt ctcgagcagc agaaaaaagt tgatgctgag
acagttggga gagtgaagag 1080gtttattcag gagaatgtct ttgtggcagc
gaatcataat ggttctcccc tttctataaa 1140ggaagcaccc aaagaactca
gcttcggtgc acgtgcagag ctgcccagga tccacccagt 1200tgcatcgaag
taagcggccg cgccatagat aaaataaaag attttattta gtctccagaa
1260aaagggggg 1269611329DNAArtificial SequenceCD codon optimized
heat stabilized linker OPRT 61aacacgatta taaatggtga ccggcggcat
ggcctccaag tgggatcaaa agggcatgga 60tatcgcttac gaggaggccc tgctgggcta
caaggagggc ggcgtgccta tcggcggctg 120tctgatcaac aacaaggacg
gcagtgtgct gggcaggggc cacaacatga ggttccagaa 180gggctccgcc
accctgcacg gcgagatctc caccctggag aactgtggca ggctggaggg
240caaggtgtac aaggacacca ccctgtacac caccctgtcc ccttgtgaca
tgtgtaccgg 300cgctatcatc atgtacggca tccctaggtg tgtgatcggc
gagaacgtga acttcaagtc 360caagggcgag aagtacctgc aaaccagggg
ccacgaggtg gtggttgttg acgatgagag 420gtgtaagaag ctgatgaagc
agttcatcga cgagaggcct caggactggt tcgaggatat 480cggcgagtcc
ggcggcggcg cctccggcgg cggcgcctcc ggcggcggcg cctccggcgg
540cggcgccgcg gtcgctcgtg cagctttggg gccattggtg acgggtctgt
acgacgtgca 600ggctttcaag tttggggact tcgtgctgaa gagcgggctt
tcctccccca tctacatcga 660tctgcggggc atcgtgtctc gaccgcgtct
tctgagtcag gttgcagata ttttattcca 720aactgcccaa aatgcaggca
tcagttttga caccgtgtgt ggagtgcctt atacagcttt 780gccattggct
acagttatct gttcaaccaa tcaaattcca atgcttatta gaaggaaaga
840aacaaaggat tatggaacta agcgtcttgt agaaggaact attaatccag
gagaaacctg 900tttaatcatt gaagatgttg tcaccagtgg atctagtgtt
ttggaaactg ttgaggttct 960tcagaaggag ggcttgaagg tcactgatgc
catagtgctg ttggacagag agcagggagg 1020caaggacaag ttgcaggcgc
acgggatccg cctccactca gtgtgtacat tgtccaaaat 1080gctggagatt
ctcgagcagc agaaaaaagt tgatgctgag acagttggga gagtgaagag
1140gtttattcag gagaatgtct ttgtggcagc gaatcataat ggttctcccc
tttctataaa 1200ggaagcaccc aaagaactca gcttcggtgc acgtgcagag
ctgcccagga tccacccagt 1260tgcatcgaag taagcggccg cgccatagat
aaaataaaag attttattta gtctccagaa 1320aaagggggg
13296223DNAArtificial SequenceOligonucleotide primer 62ctgatcttac
tctttggacc ttg 236324DNAArtificial SequenceOligonucleotide primer
63cccctttttc tggagactaa ataa 246419DNAArtificial
SequenceOligonucleotide primer 64agcccacaac ccctcactc
196518DNAArtificial SequenceOligonucleotide primer 65tctcccgatc
ccggacga 186626DNAArtificial SequenceOligonucleotide primer
66ccccaaatga aagacccccg ctgacg 266723DNAArtificial
SequenceOligonucleotide primer 67atcatcatgt acggcatccc tag
236824DNAArtificial SequenceOligonucleotide primer 68tgaactgctt
catcagcttc ttac 246925DNAArtificial SequenceOligonucleotide primer
69tcatcgtcaa caaccaccac ctcgt 257022DNAArtificial
SequenceOligonucleotide primer 70aacctcaacc tcccctacaa gt
227120DNAArtificial SequenceOligonucleotide primer 71gttaagcgcc
tgataggctc 207226DNAArtificial SequenceOligonucleotide primer
72agccaccccc aggaactgga gataga 26

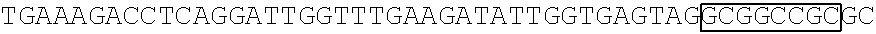
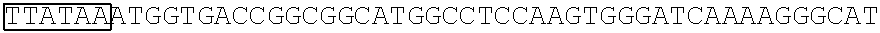
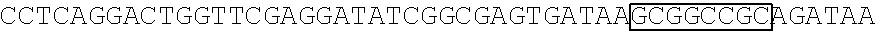
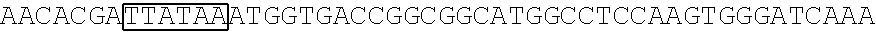
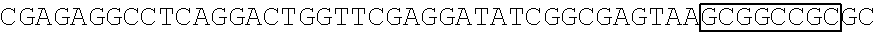
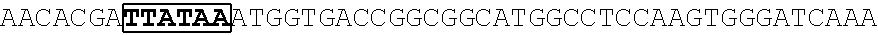
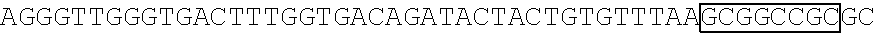
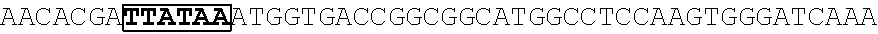
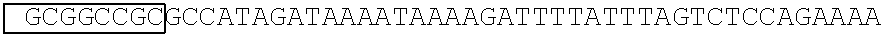
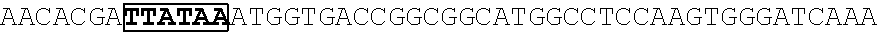
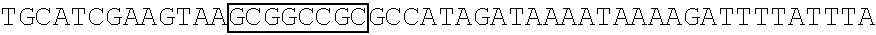
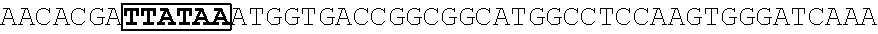
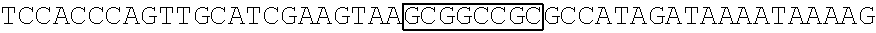
D00001

D00002

D00003

D00004

D00005

D00006
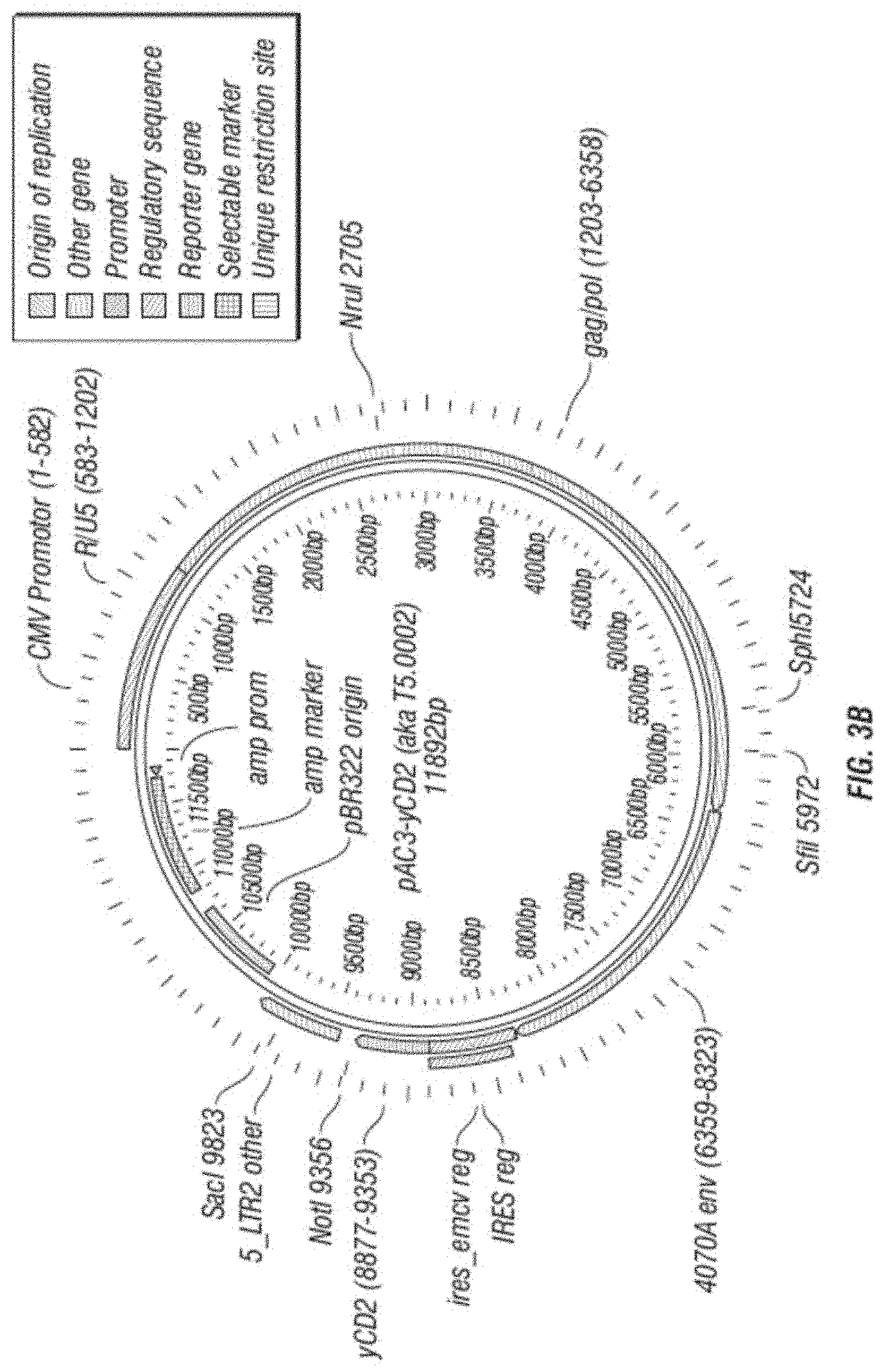
D00007
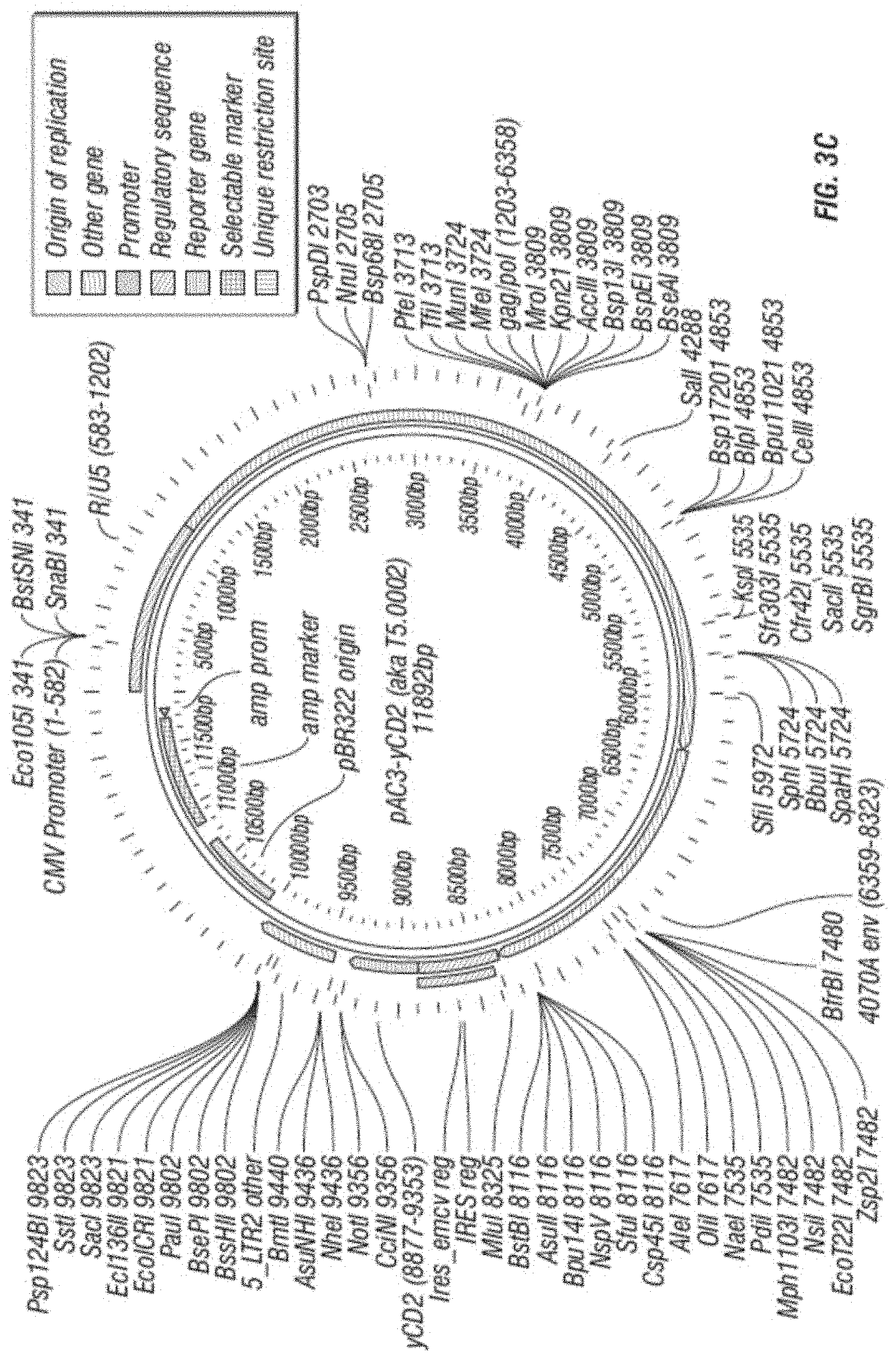
D00008

D00009
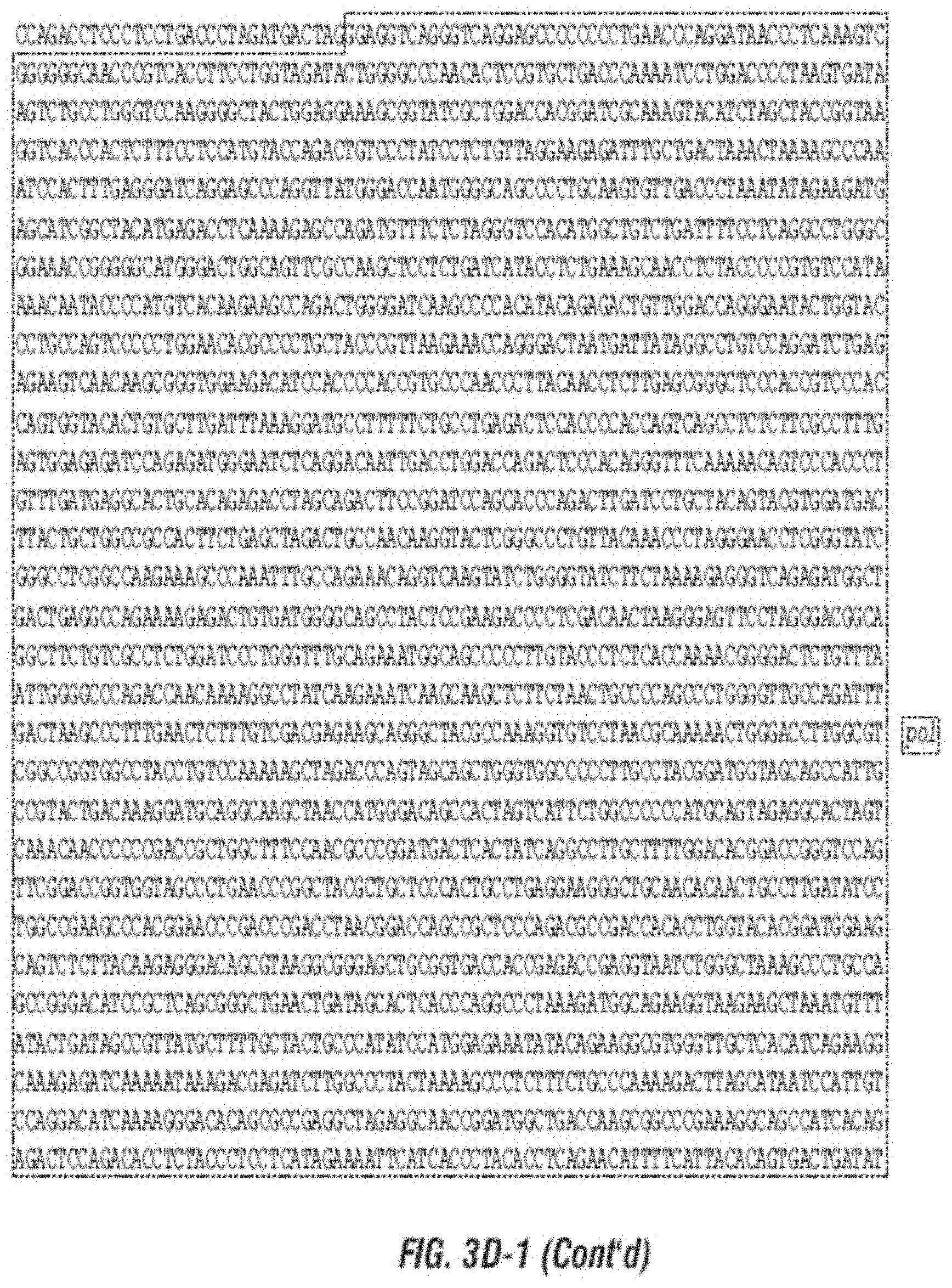
D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

D00032

D00033

D00034

D00035

D00036
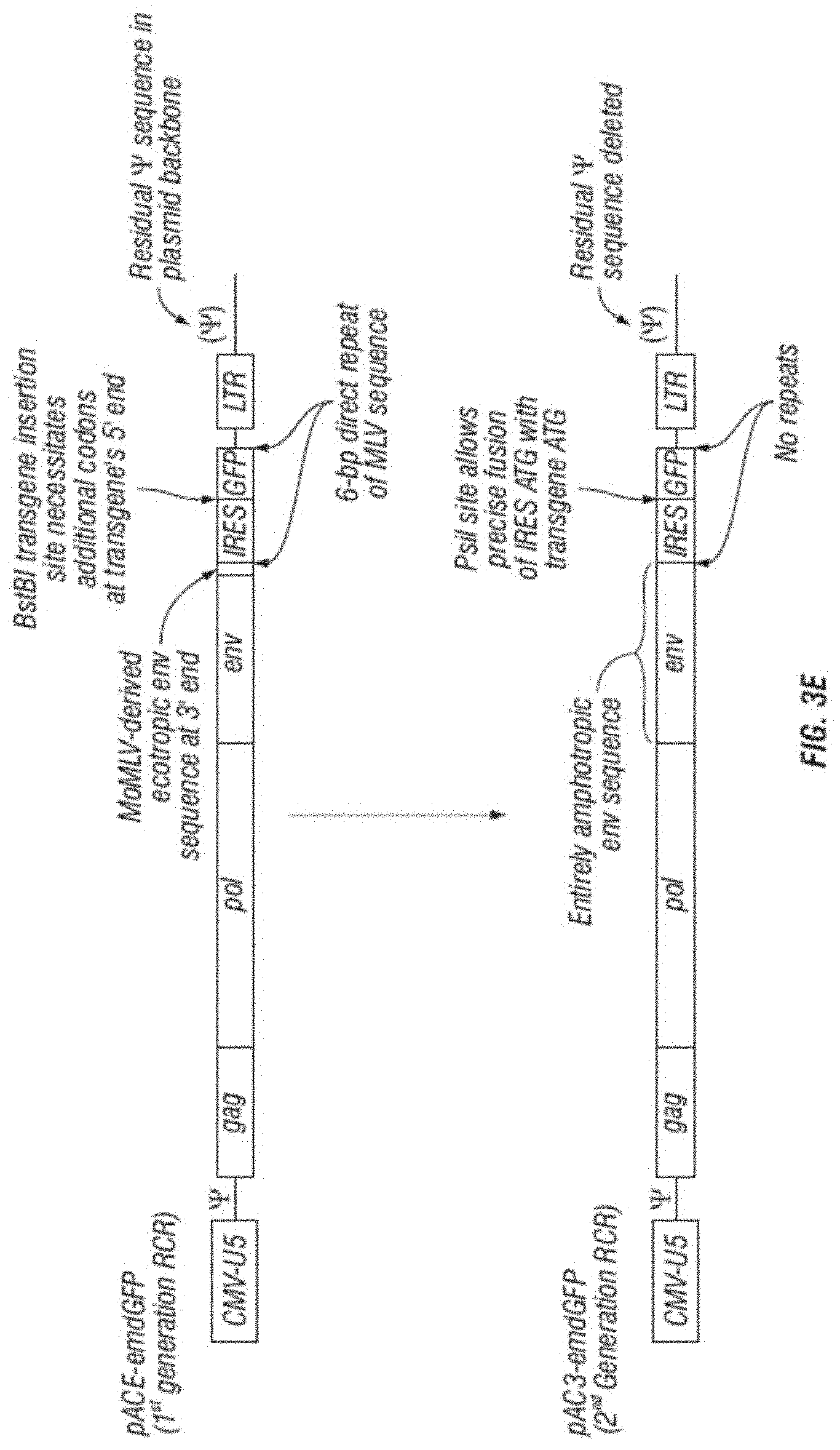
D00037
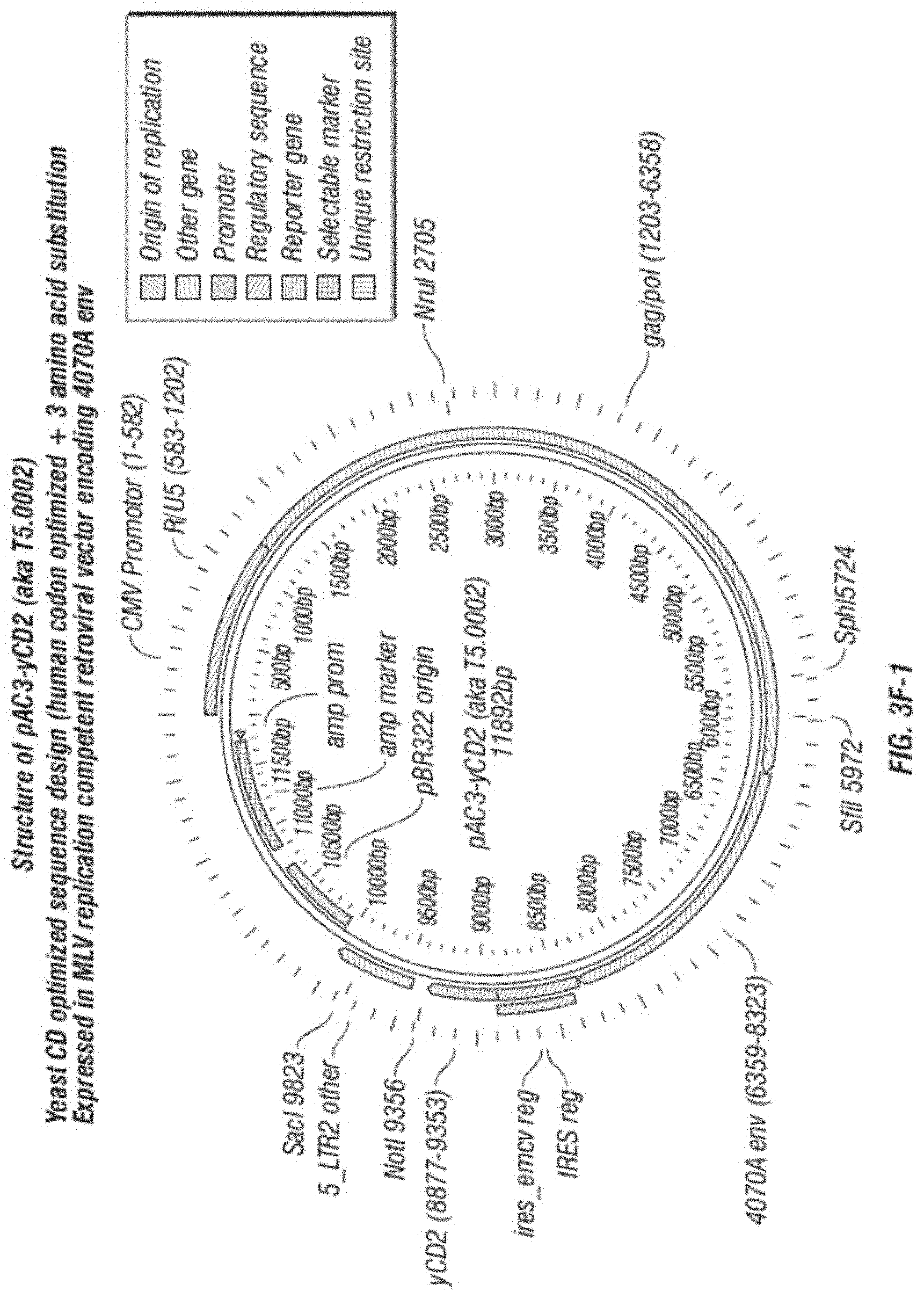
D00038
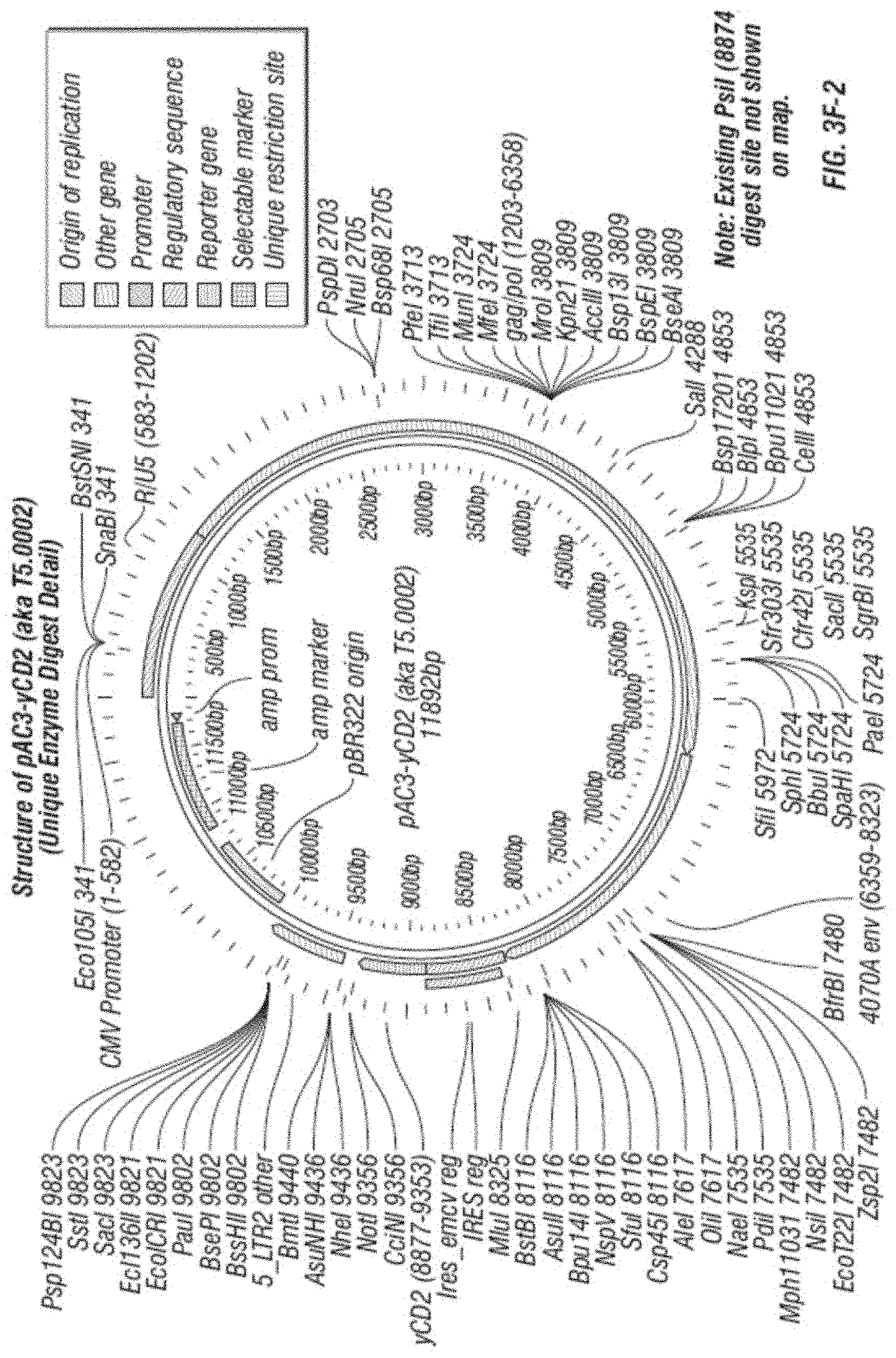
D00039

D00040

D00041

D00042

D00043

D00044

D00045

D00046

D00047

D00048

D00049

D00050

D00051

D00052

D00053

D00054

D00055

D00056

D00057

D00058

D00059

D00060

D00061

D00062

D00063

D00064

D00065

D00066

D00067

D00068
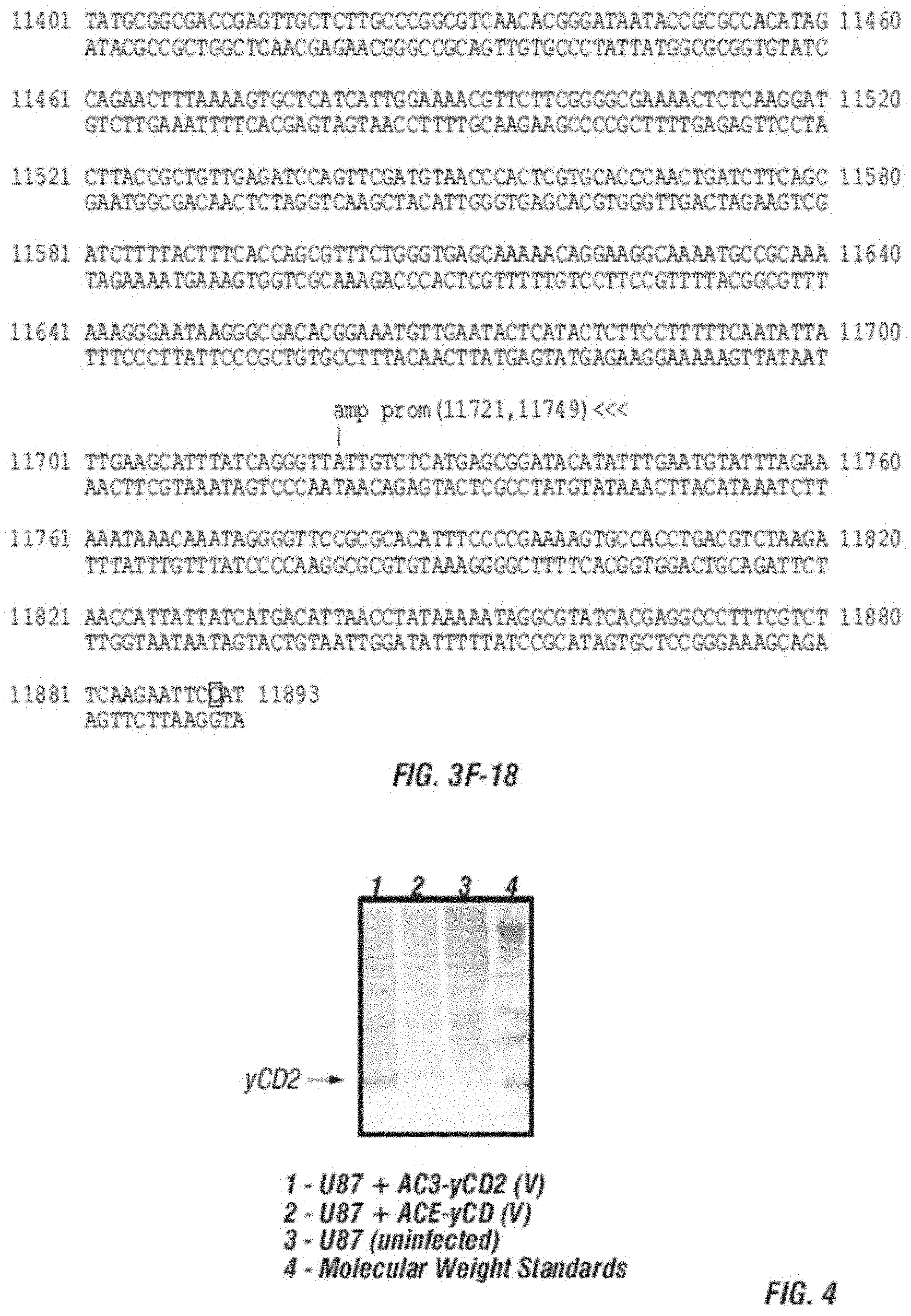
D00069

D00070
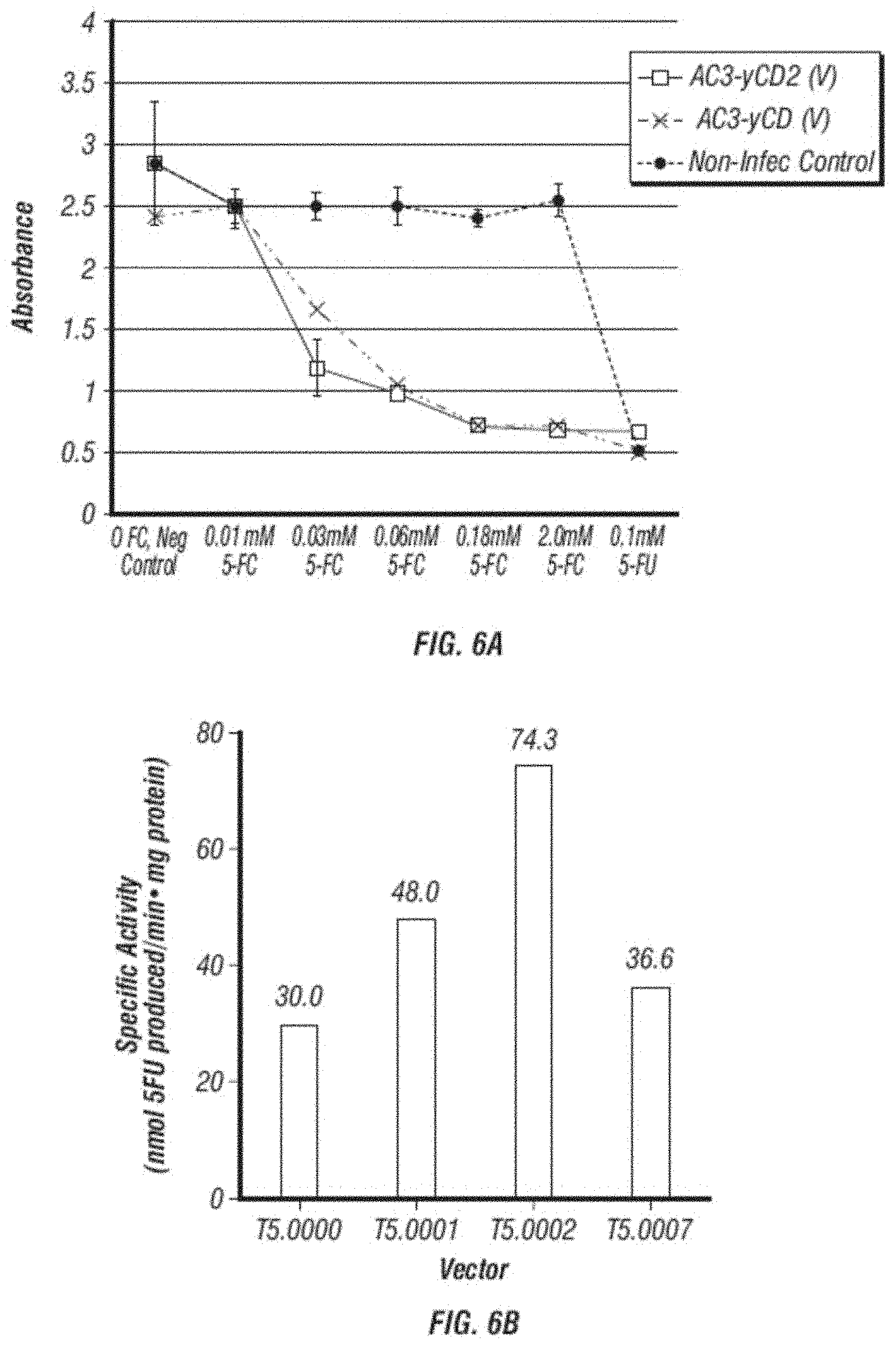
D00071
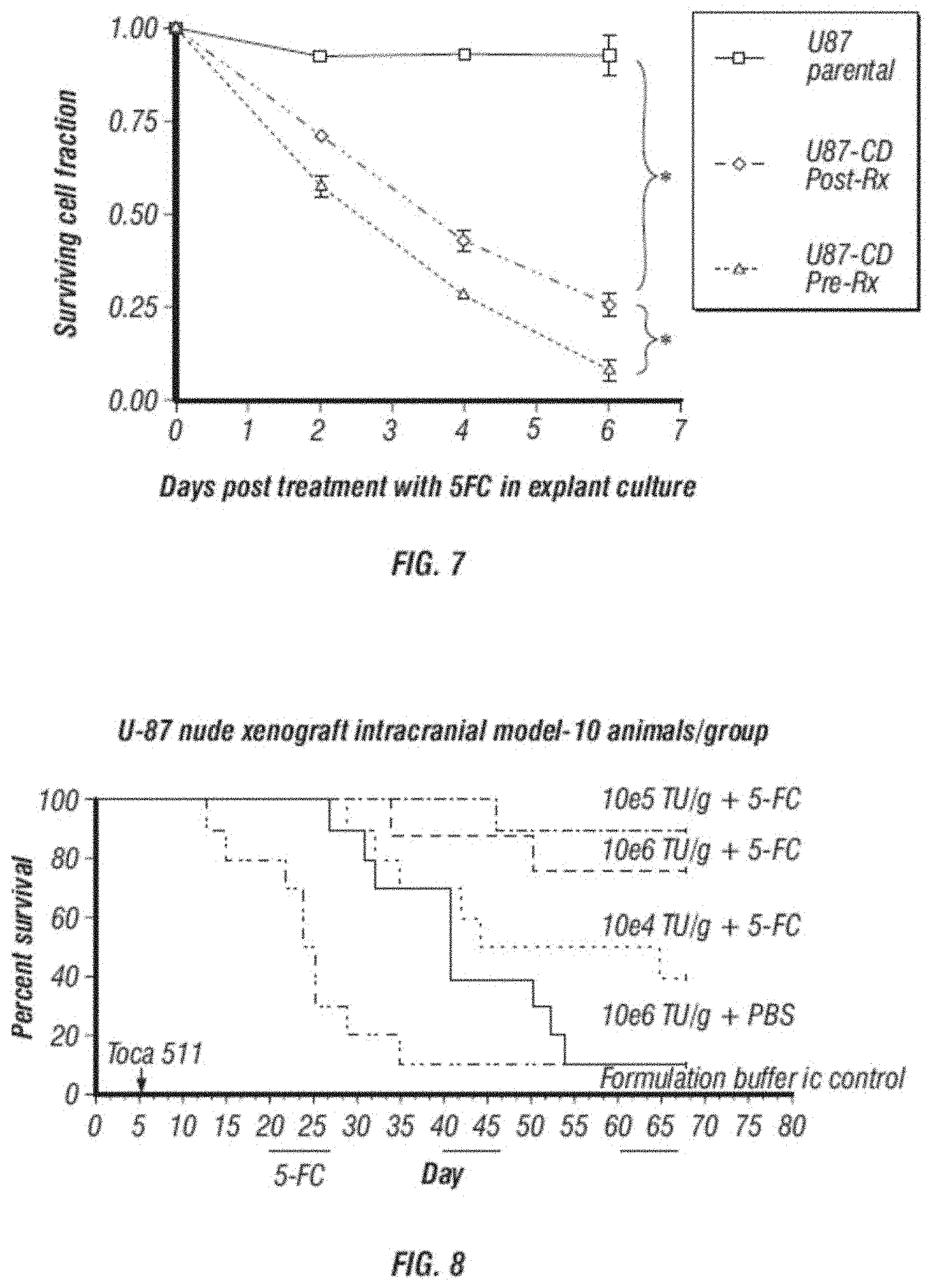
D00072
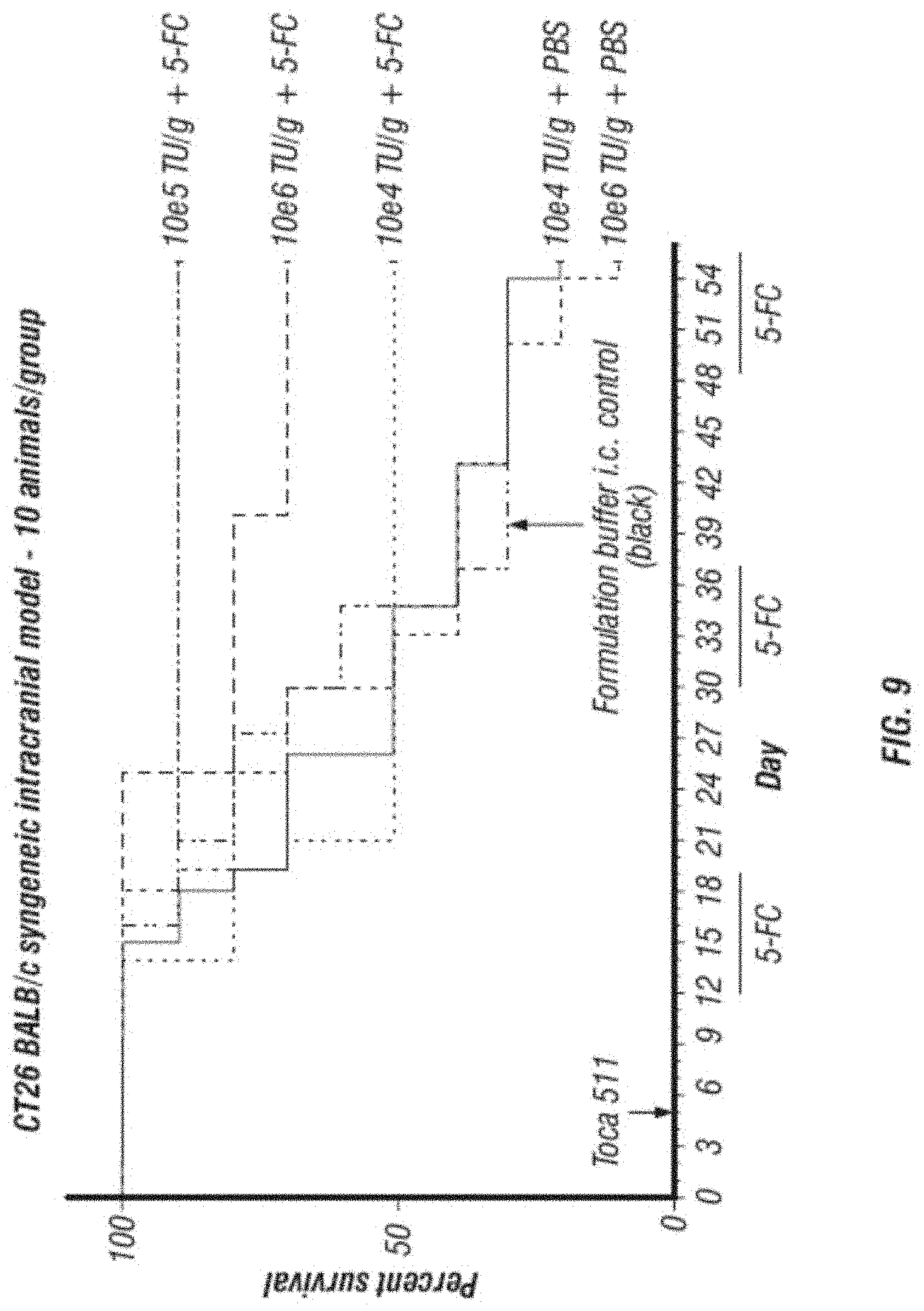
D00073

D00074
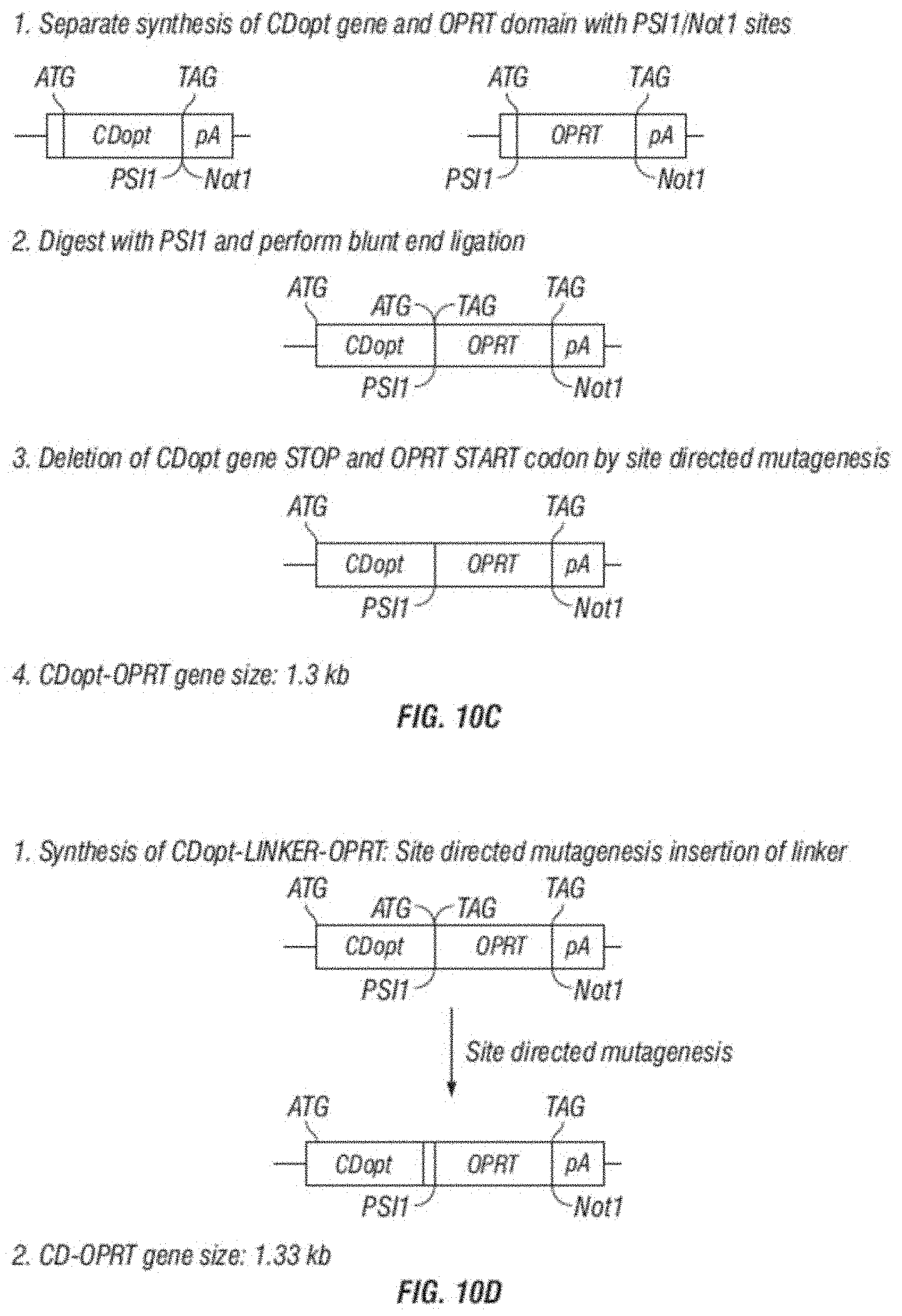
D00075
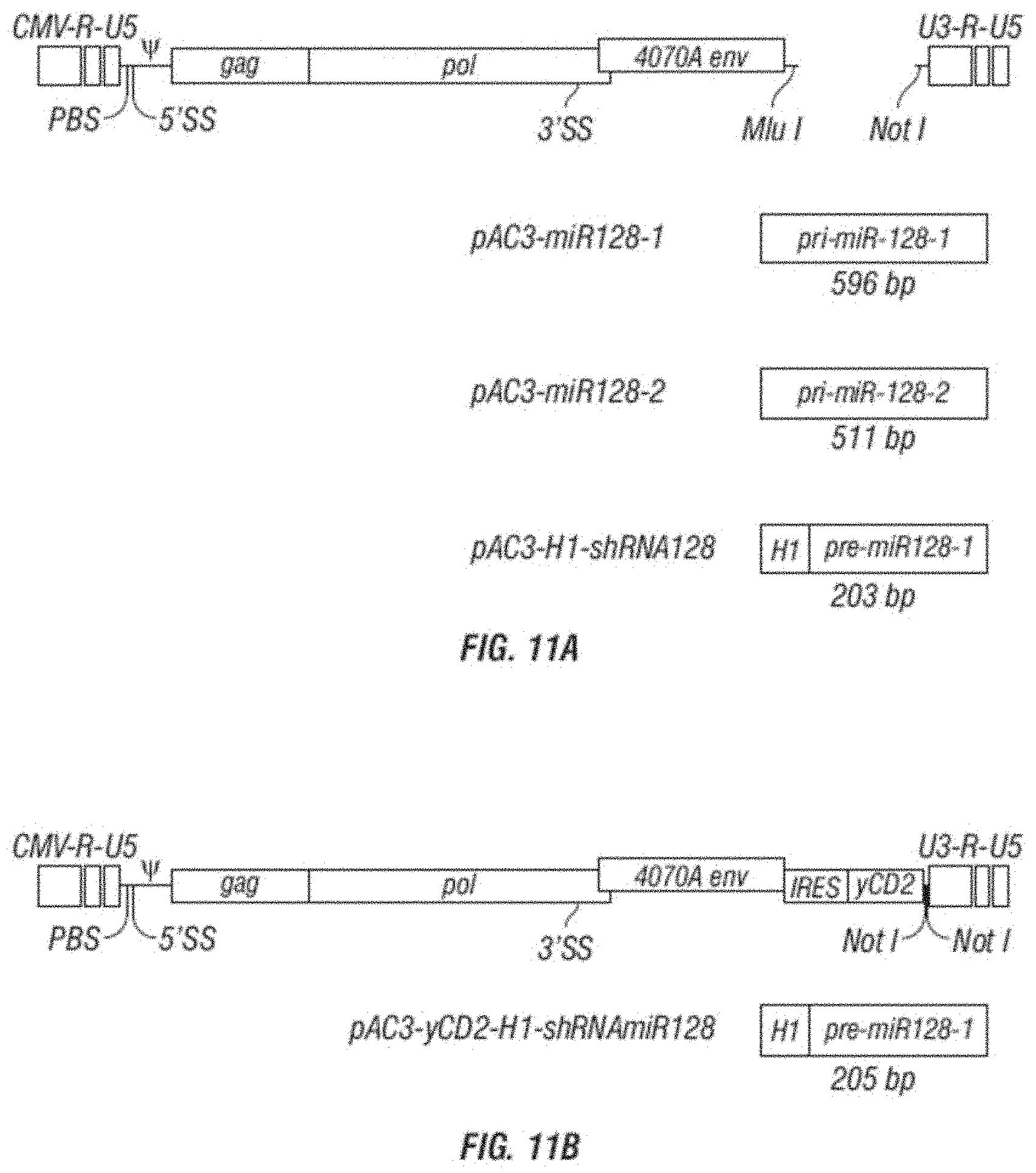
D00076
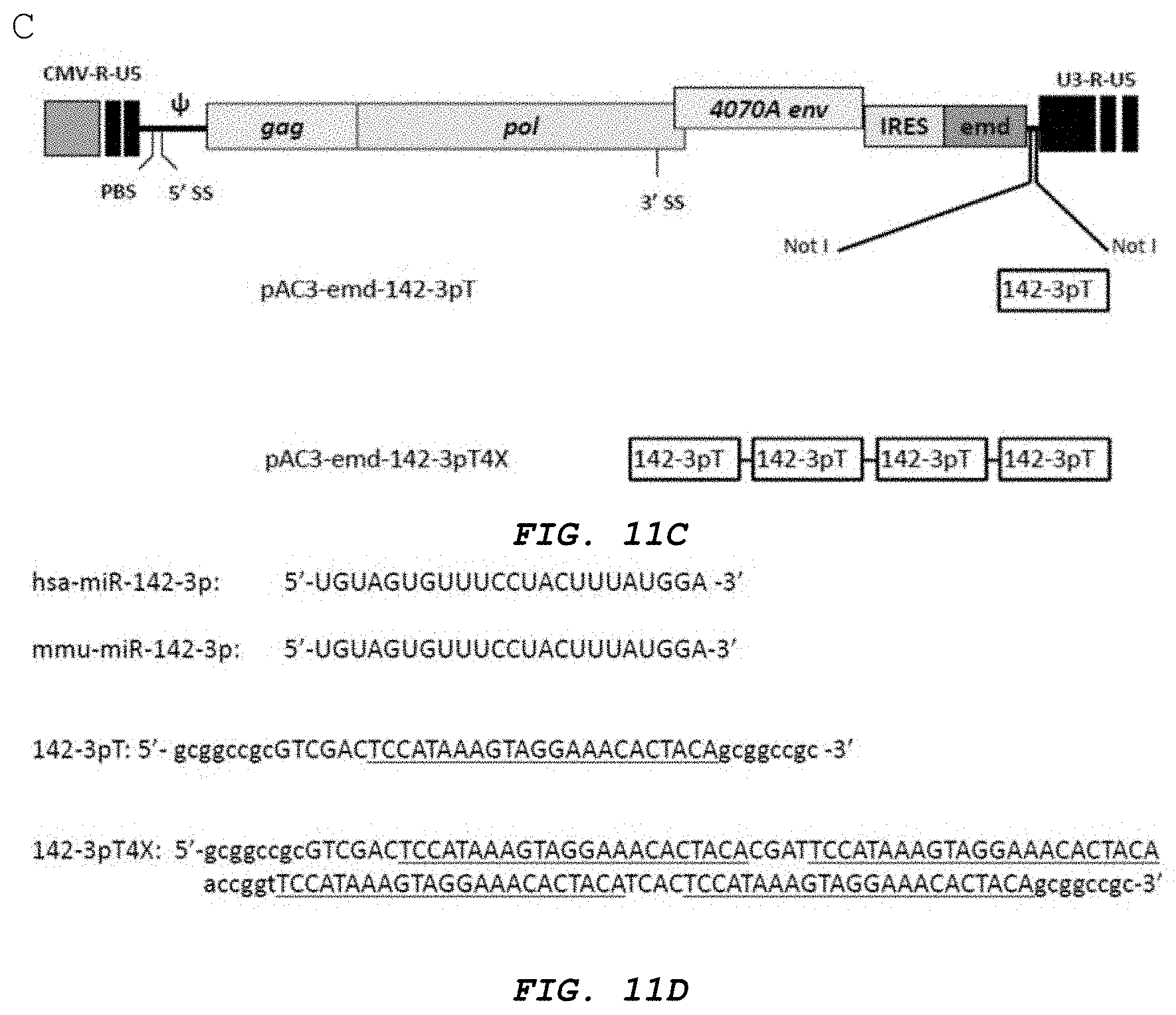
D00077
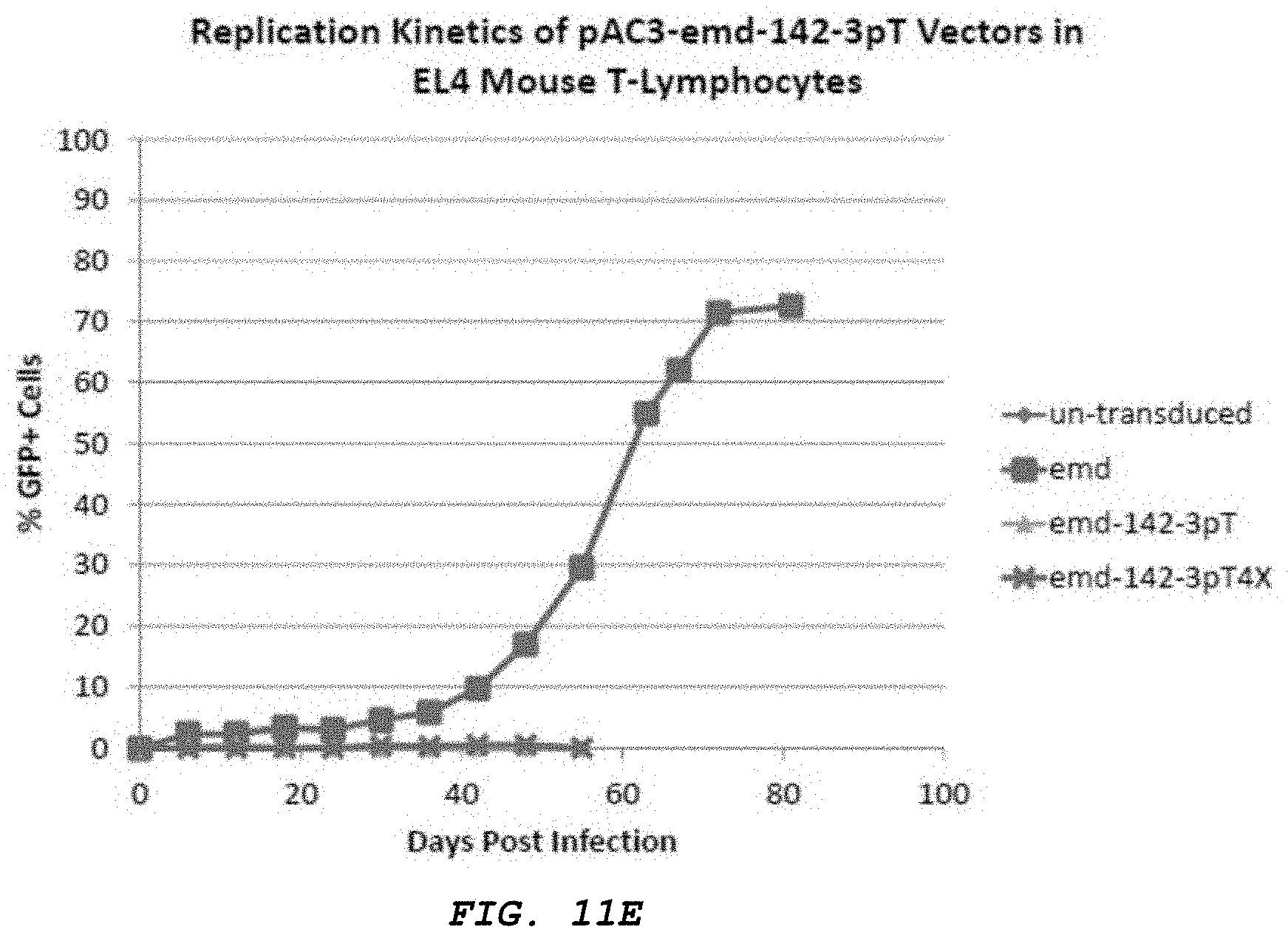
D00078
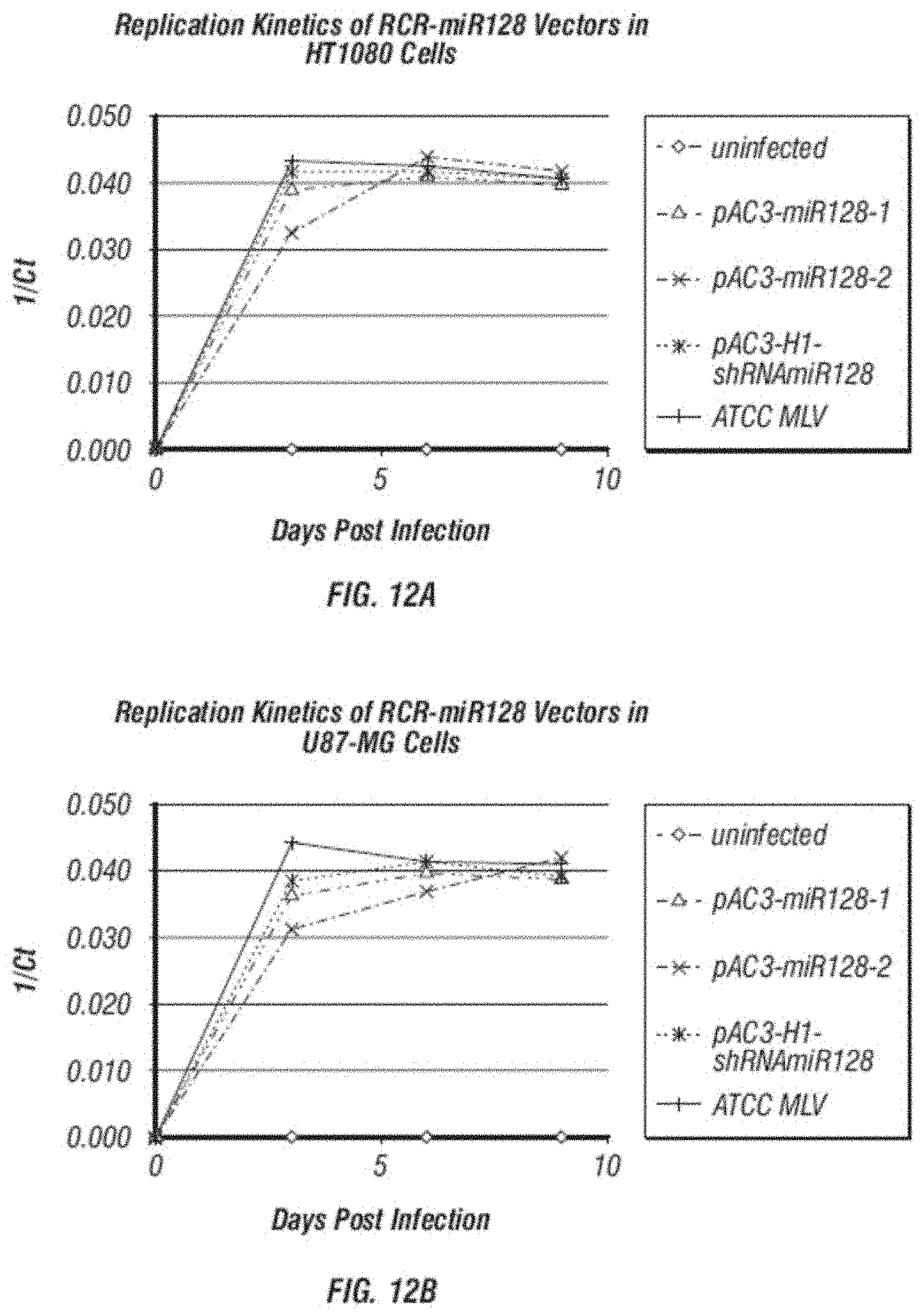
D00079
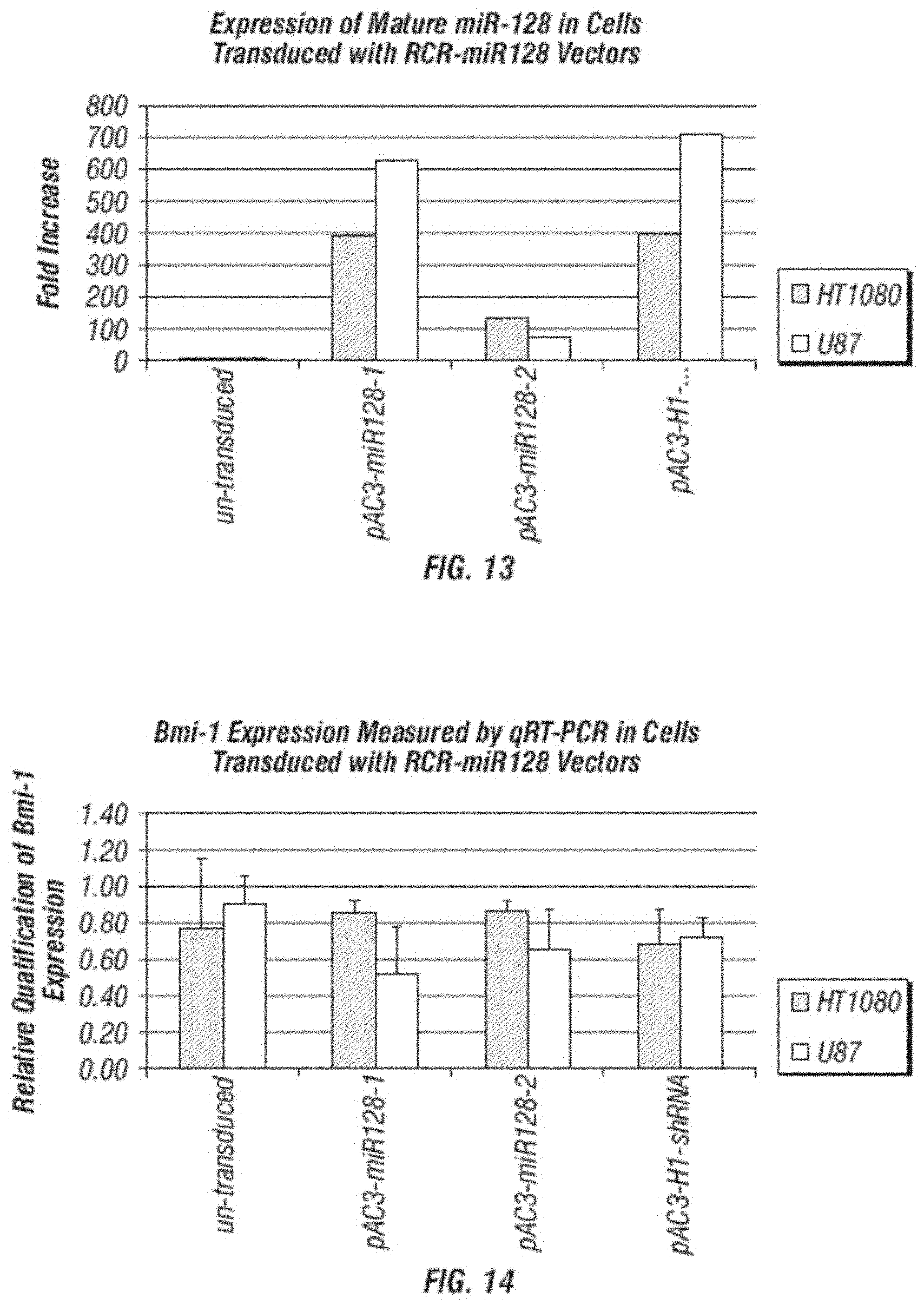
D00080

D00081
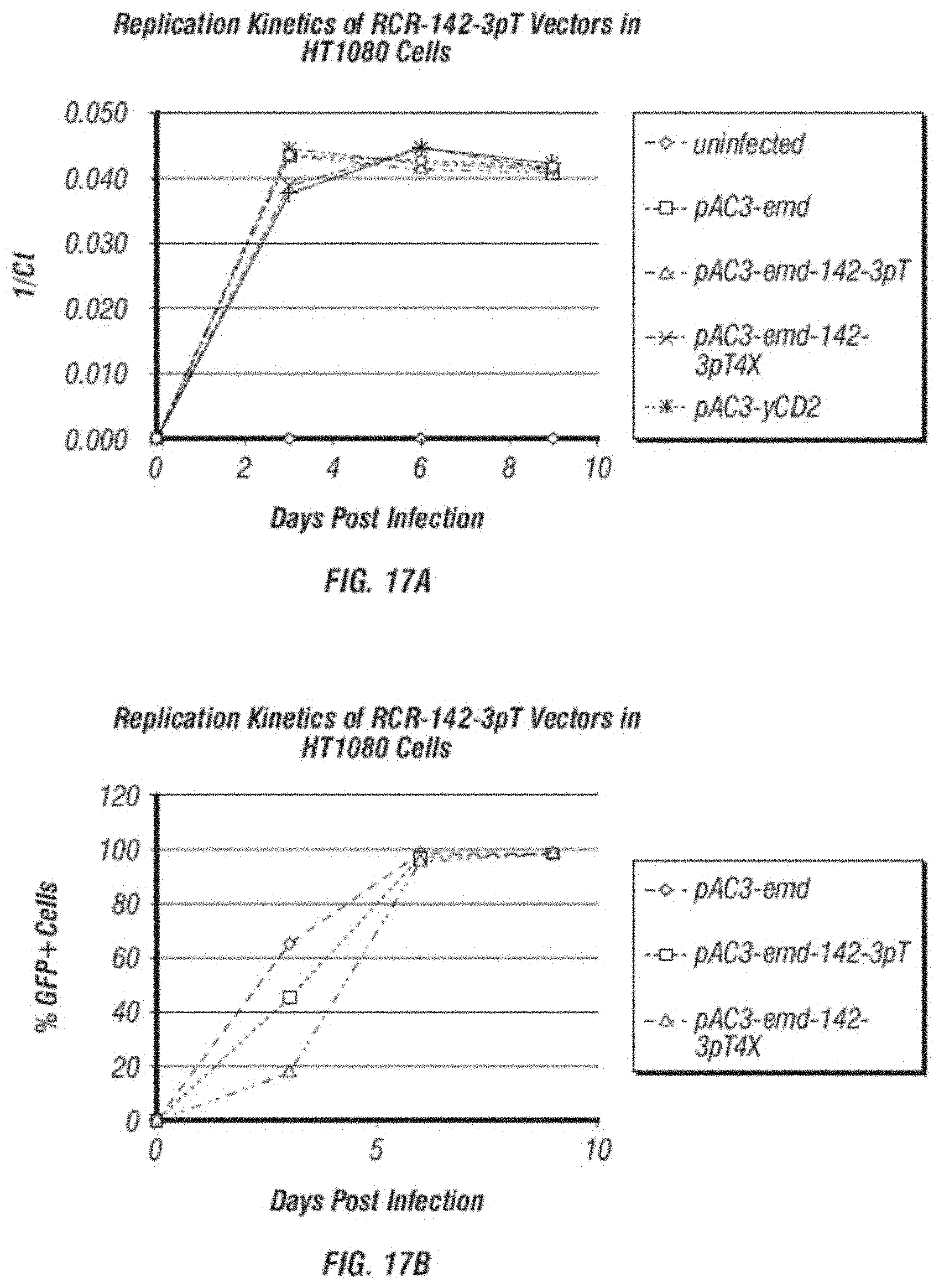
D00082
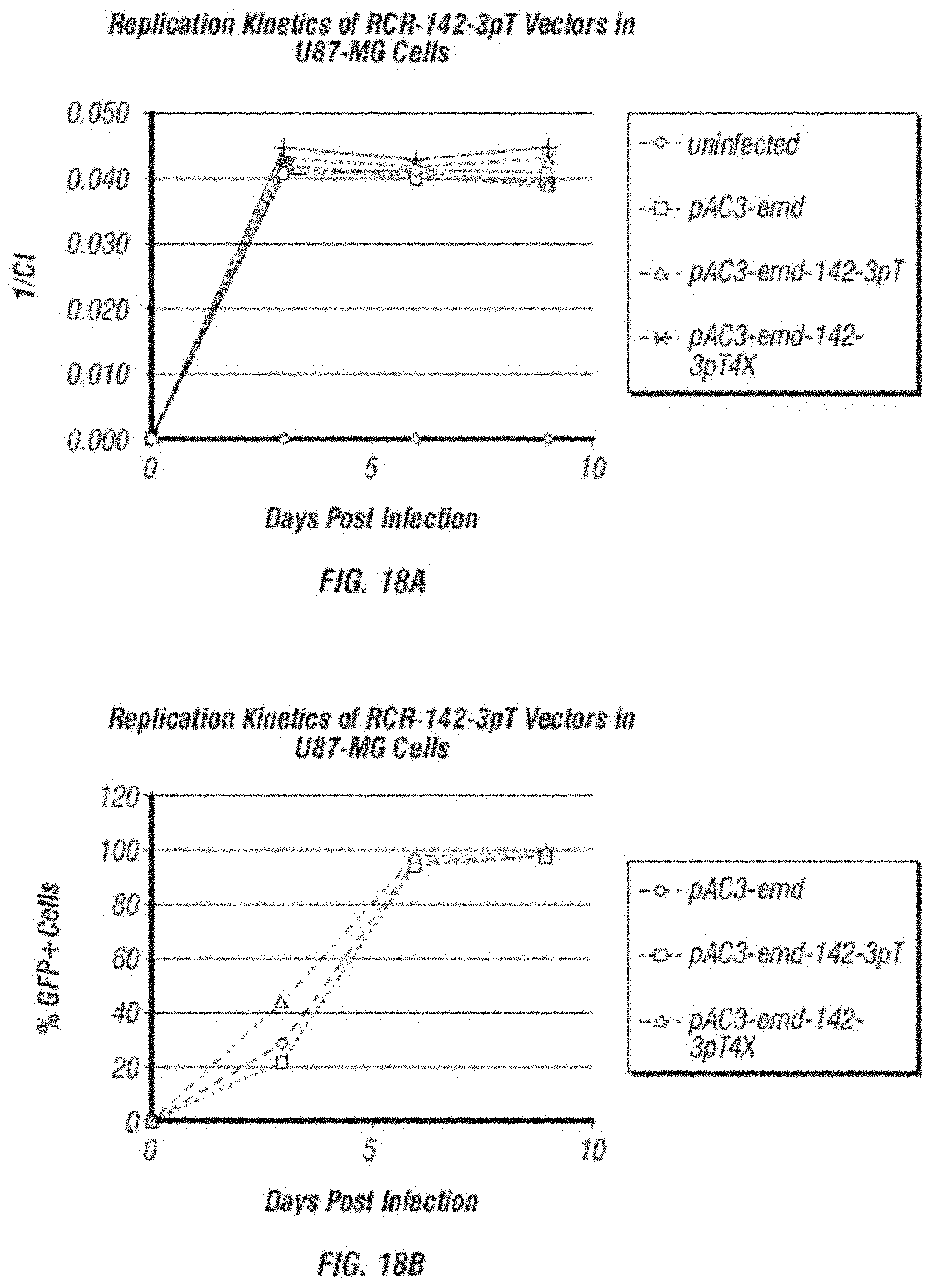
D00083
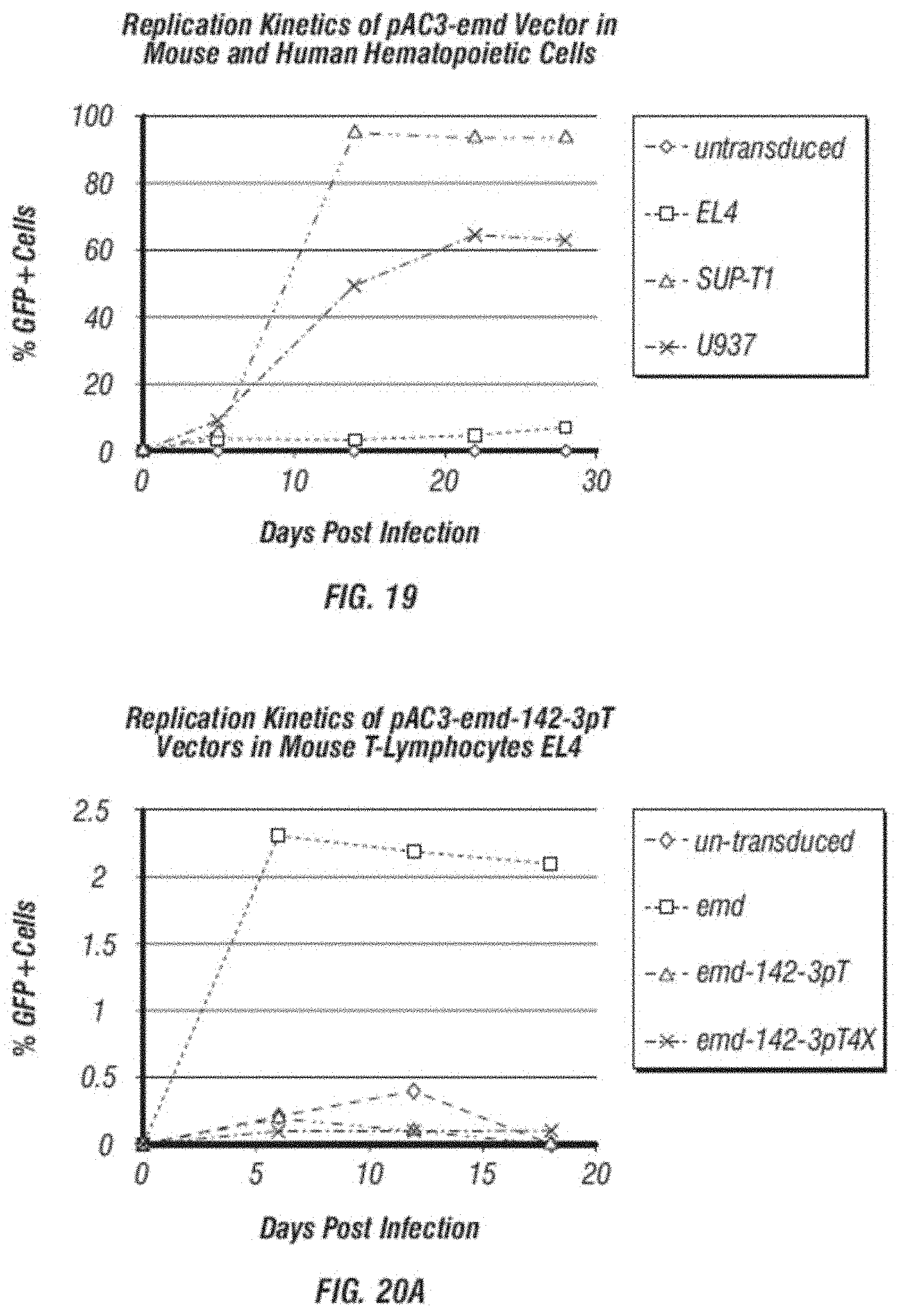
D00084
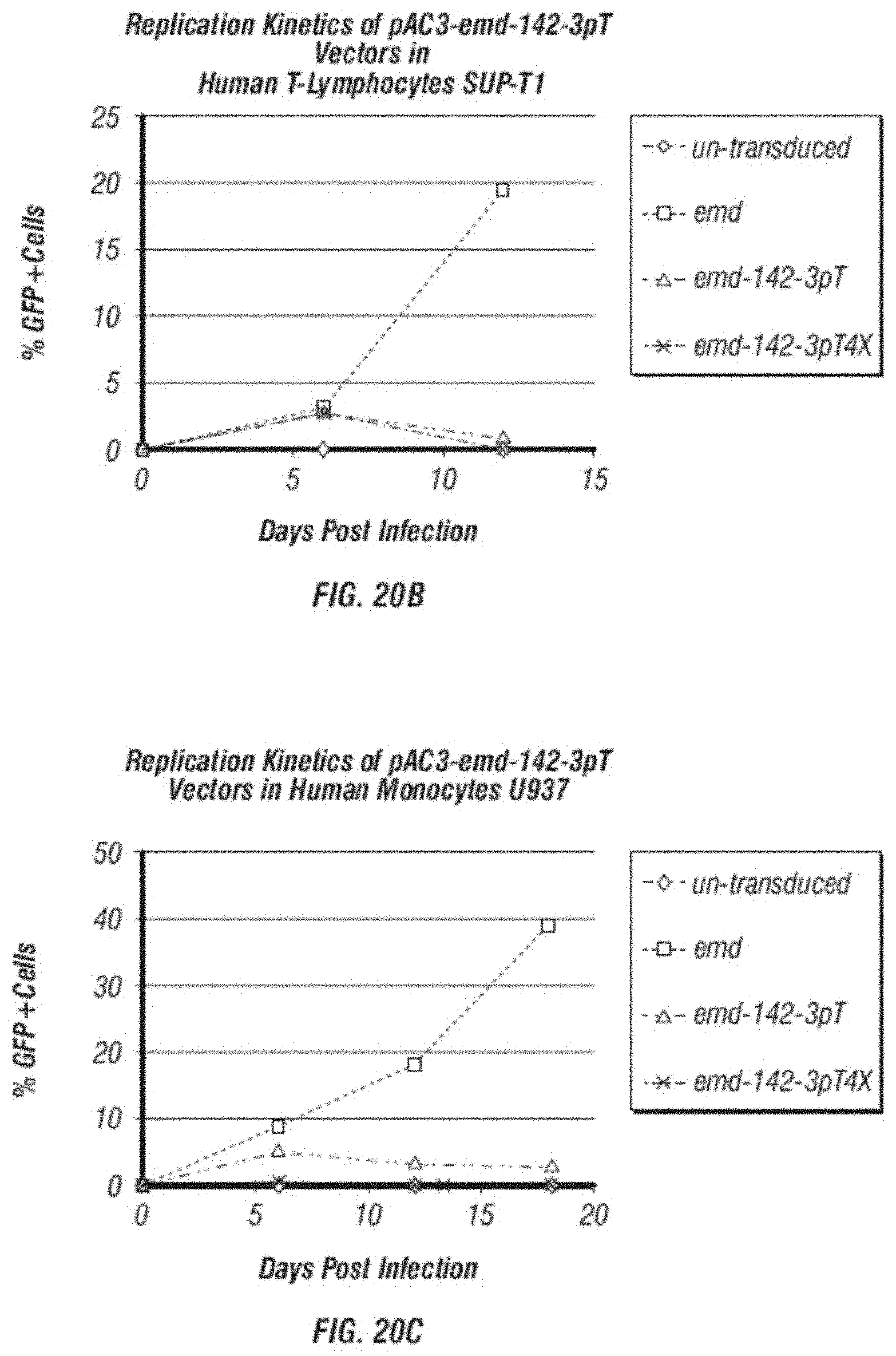
D00085
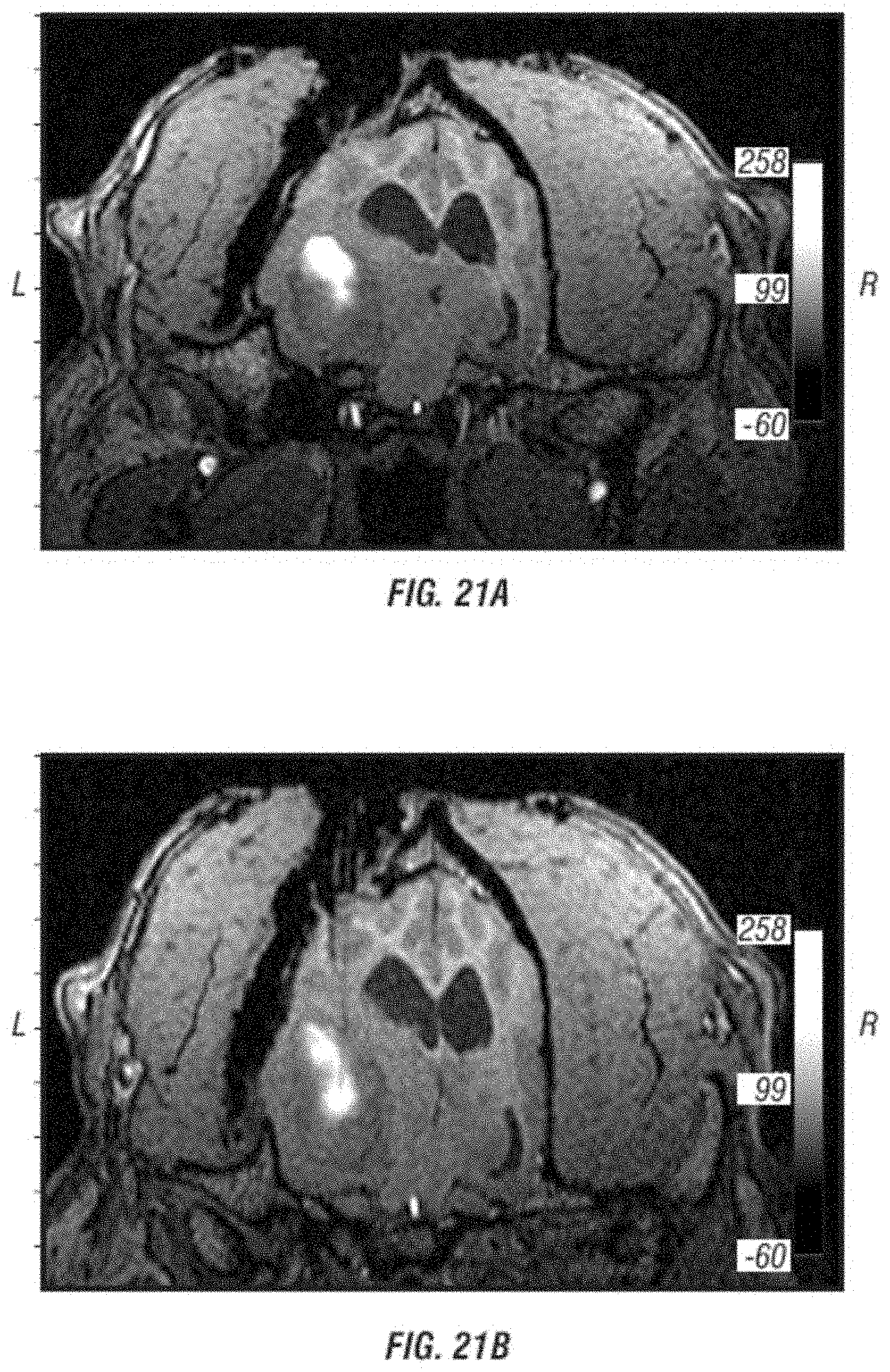

D00086
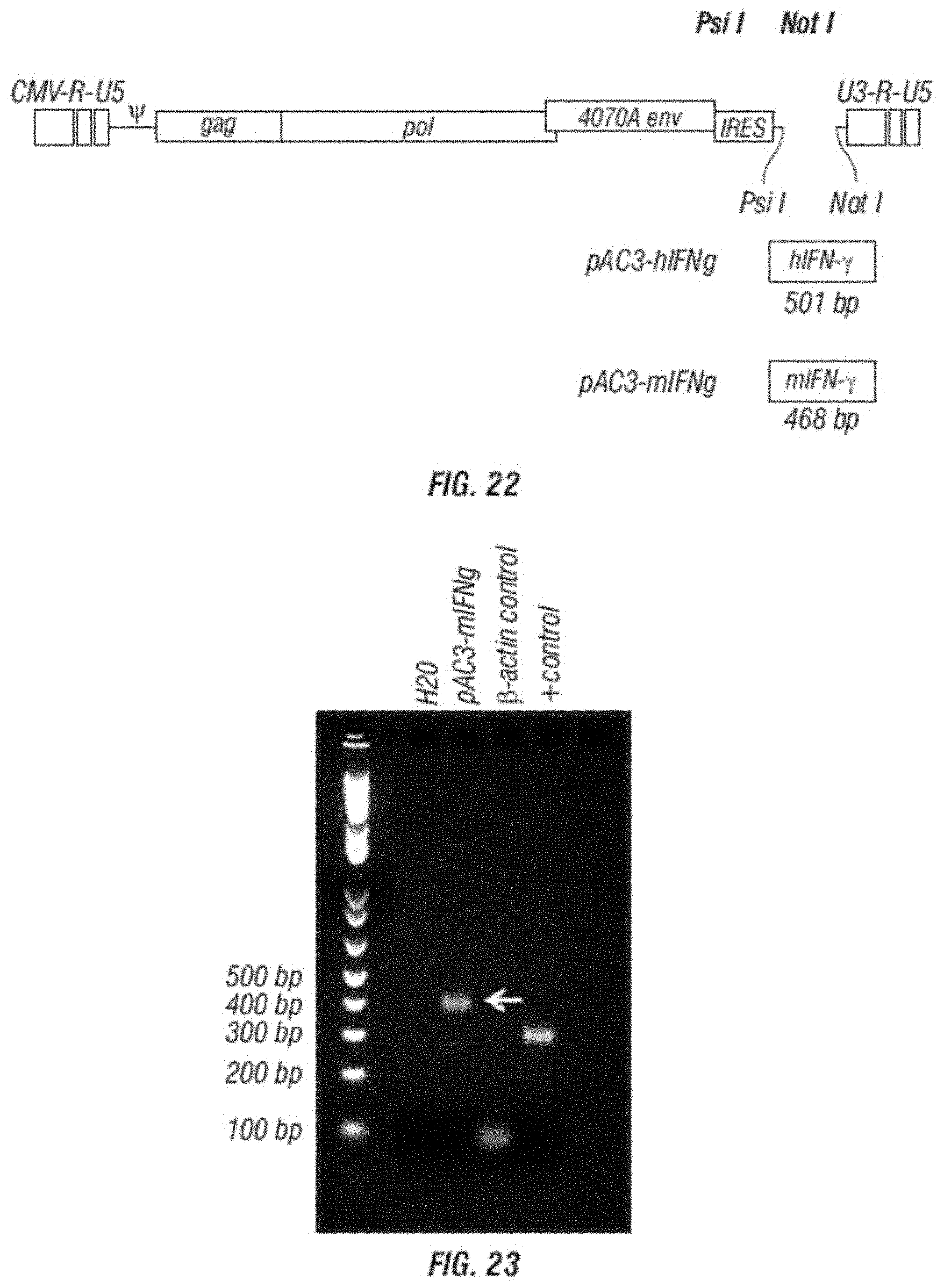
D00087
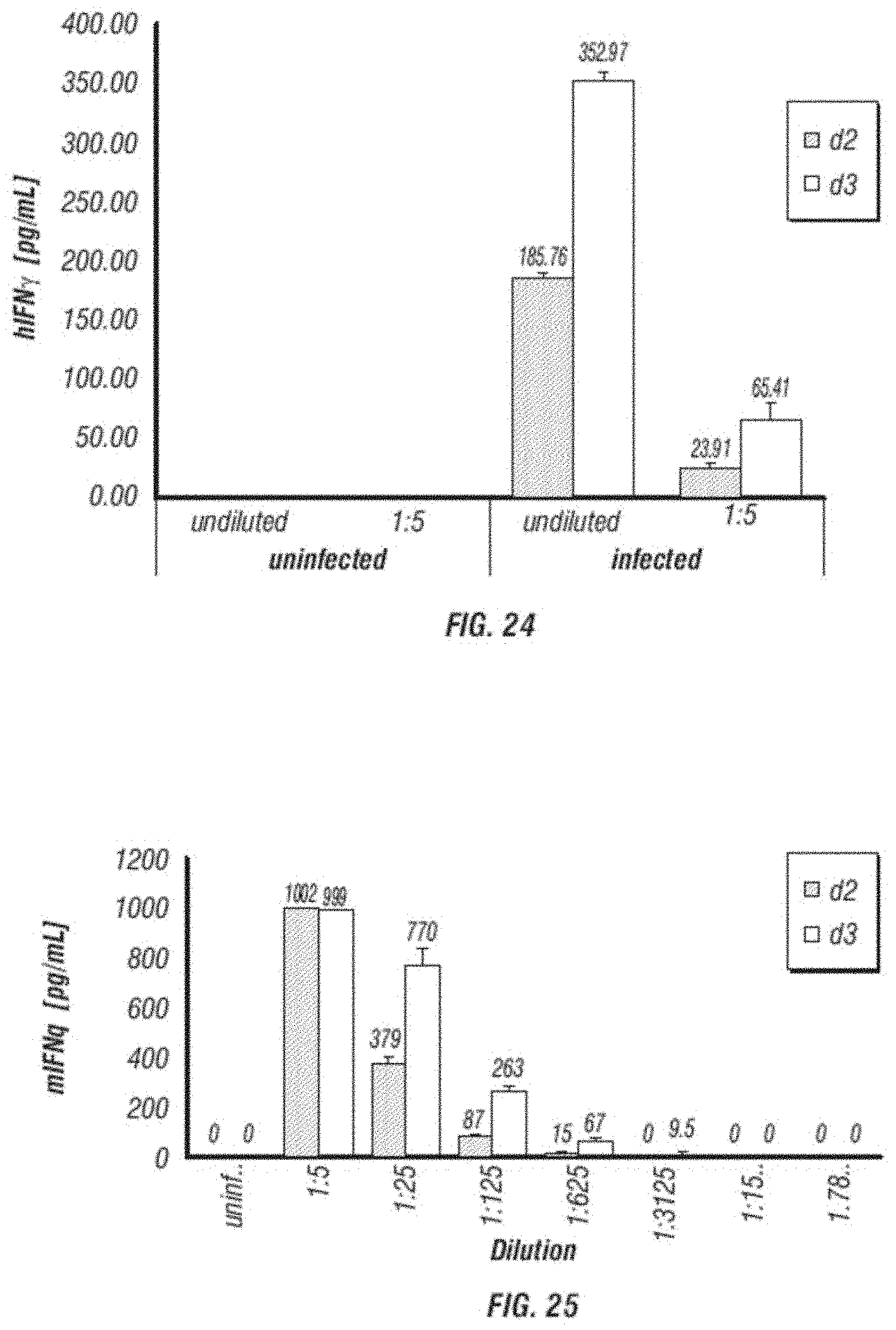
D00088
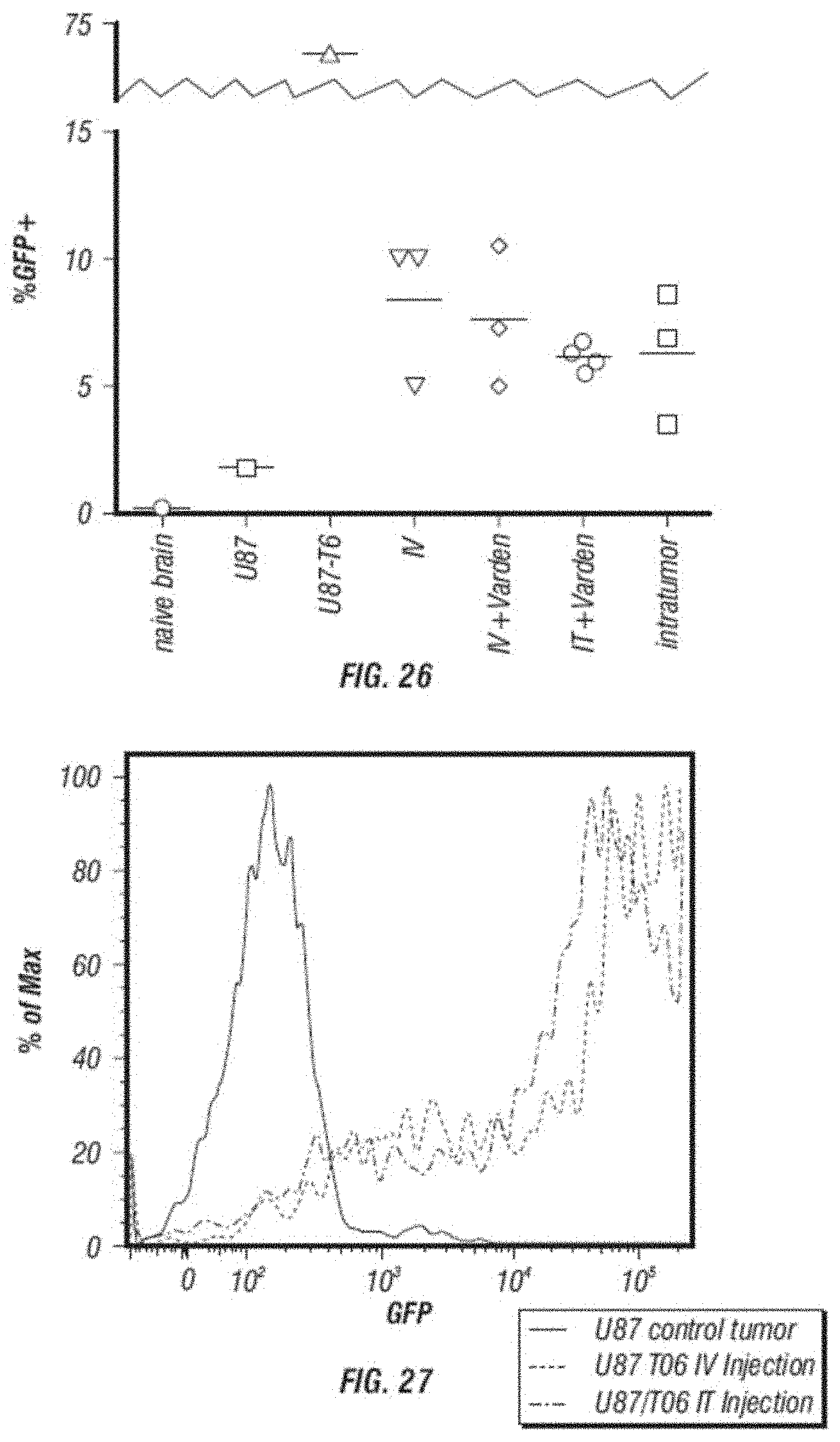
D00089
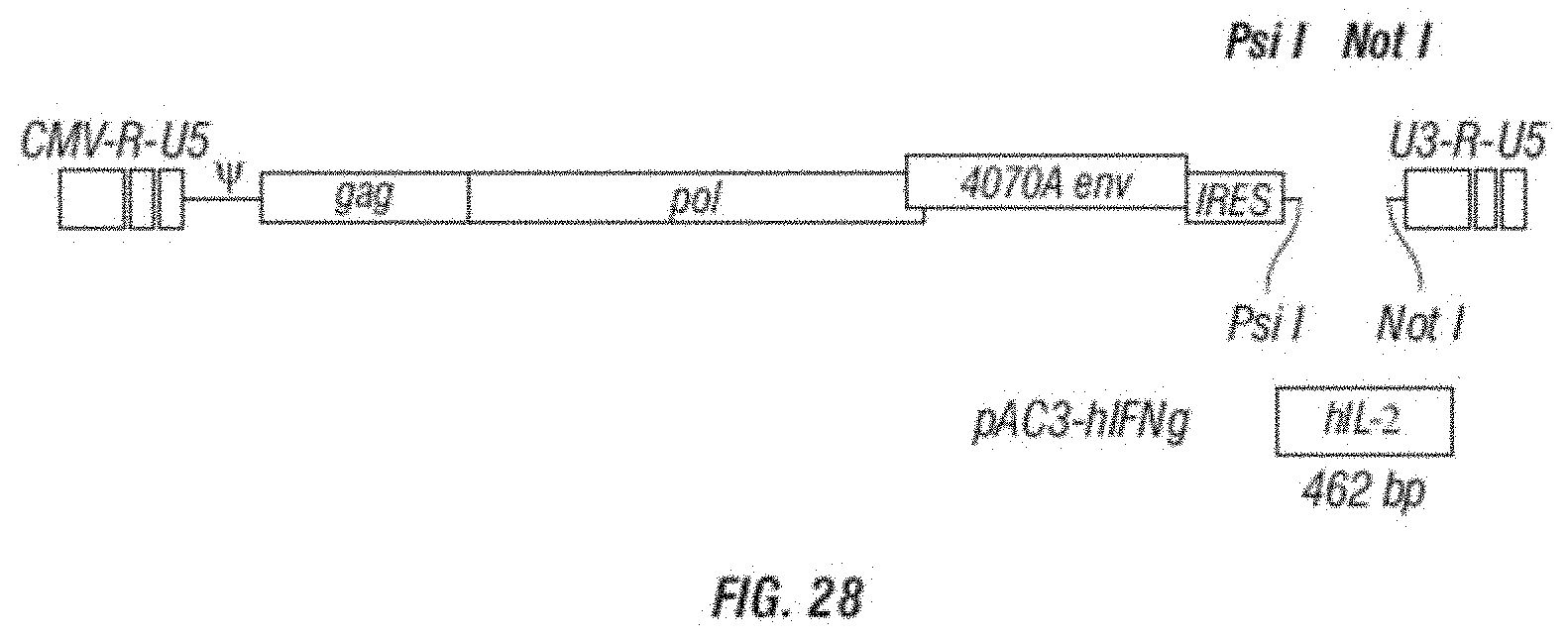
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.