Microbial Ergothioneine Biosynthesis
Han; Jixiang ; et al.
U.S. patent application number 16/714659 was filed with the patent office on 2020-07-16 for microbial ergothioneine biosynthesis. This patent application is currently assigned to Conagen Inc.. The applicant listed for this patent is Conagen Inc.. Invention is credited to Hui Chen, Jixiang Han, Oliver Yu.
| Application Number | 20200224232 16/714659 |
| Document ID | / |
| Family ID | 54359235 |
| Filed Date | 2020-07-16 |








| United States Patent Application | 20200224232 |
| Kind Code | A1 |
| Han; Jixiang ; et al. | July 16, 2020 |
MICROBIAL ERGOTHIONEINE BIOSYNTHESIS
Abstract
Disclosed are methods for ergothioneine biosynthesis. More particularly, the present disclosure relates to methods for microbial ergothioneine biosynthesis.
| Inventors: | Han; Jixiang; (Maryland Heights, MO) ; Chen; Hui; (Bedford, MA) ; Yu; Oliver; (Lexington, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Conagen Inc. Bedford MA |
||||||||||
| Family ID: | 54359235 | ||||||||||
| Appl. No.: | 16/714659 | ||||||||||
| Filed: | December 13, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15307491 | Oct 28, 2016 | 10544437 | ||
| PCT/US15/27977 | Apr 28, 2015 | |||
| 16714659 | ||||
| 61985778 | Apr 29, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 15/52 20130101; C12P 17/10 20130101; C12P 13/04 20130101 |
| International Class: | C12P 17/10 20060101 C12P017/10; C12N 15/52 20060101 C12N015/52; C12P 13/04 20060101 C12P013/04 |
Claims
19. An engineered host cell for the production of ergothioneine, wherein the host cell comprises: a nucleotide sequence encoding a heterologous EgtB comprising an amino acid sequence having at least 95% sequence identity to SEQ ID NO: 2, a nucleotide sequence encoding a heterologous EgtC comprising an amino acid sequence having at least 95% sequence identity to SEQ ID NO: 4, a nucleotide sequence encoding a heterologous EgtD comprising an amino acid sequence having at least 95% sequence identity to SEQ ID NO: 6, and a nucleotide sequence encoding a heterologous EgtE comprising an amino acid sequence having at least 95% sequence identity to SEQ ID NO: 8; wherein the heterologous oxidoreductase (EgtB), the heterologous amidohydrolase (EgtC), the heterologous histidine methyltransferase (EgtD), and the heterologous pyridoxal 5-phosphate binding protein (EgtE) are the only heterologous proteins involved in the production of ergothioneine.
20. The engineered host cell of claim 19, wherein the heterologous EgtB comprises the amino acid sequence of SEQ ID NO:2.
21. The engineered host cell of claim 19, wherein the heterologous EgtC comprises the amino acid sequence of SEQ ID NO:4.
22. The engineered host cell of claim 19, wherein the heterologous EgtD comprises the amino acid sequence of SEQ ID NO:6.
23. The engineered host cell of claim 19, wherein the heterologous EgtE comprises the amino acid sequence of SEQ ID NO:8.
24. The engineered host cell of claim 19, wherein the heterologous EgtB comprises the amino acid sequence of SEQ ID NO:2, the heterologous EgtC comprises the amino acid sequence of SEQ ID NO:4, the heterologous EgtD comprises the amino acid sequence of SEQ ID NO:6, and the heterologous EgtE comprises the amino acid sequence of SEQ ID NO:8.
25. The engineered host cell of claim 19, wherein the host cell is selected from the group consisting of a bacterial cell and a yeast cell.
26. The engineered host cell of claim 25, wherein the host cell is an Escherichia coli cell.
27. The engineered host cell of claim 25, wherein the host cell is selected from the group consisting of a Saccharomyces cell and a Pichia cell.
28. The engineered host cell of claim 27, wherein the host cell is a Saccharomyces cerevisiae cell.
29. The engineered host cell of claim 27, wherein the host cell is a Pichia pastoris cell.
30. The engineered host cell of claim 19, wherein the nucleotide sequence encoding a heterologous EgtB, the nucleotide sequence encoding a heterologous EgtC, the nucleotide sequence encoding a heterologous EgtD, and the nucleotide sequence encoding a heterologous EgtE are on a single vector.
31. The engineered host cell of claim 19, wherein the nucleotide sequence encoding a heterologous EgtB, the nucleotide sequence encoding a heterologous EgtC, the nucleotide sequence encoding a heterologous EgtD, and the nucleotide sequence encoding a heterologous EgtE are on more than one vectors.
32. A method for producing ergothioneine, the method comprising: culturing the host cell of claim 19, inducing the host cell to express the heterologous EgtB, the heterologous EgtC, the heterologous EgtD and the heterologous EgtE; and collecting the ergothioneine.
33. The method of claim 32, wherein a substrate selected from the group consisting of histidine, methionine, cysteine, .gamma.-glutamyl cysteine and combinations thereof is added to the culture.
34. The method of claim 32, wherein iron (II) is added to the culture.
35. The method of claim 32, wherein the host cell is an E. coli cell.
36. The method of claim 32, wherein the host cell is a Saccharomyces cerevisiae cell.
37. The method of claim 32, wherein the host cell is a Pichia pastoris cell.
38. The method of claim 32, wherein the heterologous EgtB comprises the amino acid sequence of SEQ ID NO: 2, the heterologous EgtC comprises the amino acid sequence of SEQ ID NO: 4, the heterologous EgtD comprises the amino acid sequence of SEQ ID NO: 6, and the heterologous EgtE comprises the amino acid sequence of SEQ ID NO: 8.
Description
STATEMENT IN SUPPORT FOR FILING A SEQUENCE LISTING
[0001] A paper copy of the Sequence Listing and a computer readable form of the Sequence Listing containing the file named "32559-12_ST25.txt", which is 19,443 bytes in size (as measured in MICROSOFT WINDOWS.RTM. EXPLORER), are provided herein and are herein incorporated by reference. This Sequence Listing consists of SEQ ID NOs:1-16.
BACKGROUND OF THE DISCLOSURE
[0002] The present disclosure relates generally to methods for ergothioneine biosynthesis. More particularly, the present disclosure relates to methods for microbial ergothioneine biosynthesis.
[0003] Ergothioneine (ET) is a histidine betaine derivative with a thiol group attached to the C.sub.2 atom of the imidazole ring. As a thione tautomer, ET is a very stable antioxidant with unique properties. Unlike glutathione and ascorbate, ET can scavenge oxidizing species that are not free radicals. ET is a natural compound that is produced in Actinobacteria such as Mycobacterium smegmatis and filamentous fungi such as Neurospora crassa. Other species of bacteria, such as Bacillus subtilis, Escherichia coli, Proteus vulgaris and Streptococcus, as well as fungi belonging to the groups Ascomycetes and Deuteromycetes, cannot make ergothioneine. Animals and plants also cannot make ergothioneine and must obtain it from dietary sources or in the case of plants, from their environment.
[0004] Although the function of ET in microbial cells is not well understood, it is believed to be critical in human physiology. Humans absorb ET from dietary sources and ET accumulates in specific tissues and cells such as the liver, kidney, central nervous system, and red blood cells. It is evidenced that a specific cation transporter (OCTN1) has high affinity for ET in the human body, and both hyperactivity and deficiency of the transporter exert negative effects on human cells.
[0005] The biosynthesis of ET has been detected in certain mycobacteria fungi, however, the exact metabolic pathway is not completed or only partially confirmed. Seebeck reconstituted mycobacterial ergothioneine biosynthesis in vitro using E. coli to separately express a formylglycine-generating enzyme-like protein (EgtB), a glutamine amidotransferase (EgtC), a histidine methyltransferase (EgtD), and an unrelated .beta.-lyase from Erwinia tasmaniensis to replace the pyridoxal 5-phosphate binding protein (EgtE), because the recombinant production of soluble EgtE protein failed (see, J. Am. Chem. Soc. 2010, 132:6632-6633).
[0006] Thus far, only 3 genes coding for EgtB, EgtC, and EgtD have been identified for the production of ergothioneine in vitro. A putative gene for EgtE remains uncharacterized either in vitro or in vivo. To date, no microbial production using the above genes to engineer the mycobacterial ergothioneine metabolic pathway in E. coli has been reported despite various attempts at bio-conversion. Also, although various fungal and mycobacterial sources are available for the ergothioneine extraction, the yields are too low to be commercially viable for industrial production of ergothioneine. Accordingly, there exists a need for producing ergothioneine.
SUMMARY OF THE DISCLOSURE
[0007] The present disclosure relates generally to engineered host cells and methods for producing ergothioneine. More particularly, the present disclosure relates to an engineered host cell and methods for microbial ergothioneine biosynthesis using the engineered host cell.
[0008] In one aspect, the present disclosure is directed to a transformed host cell for producing ergothioneine comprising a nucleic acid sequence encoding EgtB, a nucleic acid sequence encoding EgtC, a nucleic acid sequence encoding EgtD and a nucleic acid sequence encoding EgtE.
[0009] In another aspect, the present disclosure is directed to a method for producing ergothioneine. The method comprises culturing a host cell, wherein the host cell is transformed with a nucleic acid sequence encoding EgtB, a nucleic acid sequence encoding EgtC, a nucleic acid sequence encoding EgtD and a nucleic acid sequence encoding EgtE; inducing the host cell to express the nucleic acid sequence encoding EgtB, the nucleic acid sequence encoding EgtC, the nucleic acid sequence encoding EgtD and the nucleic acid sequence encoding EgtE; and collecting the ergothioneine.
[0010] In another aspect, the present disclosure is directed to an expression vector for the production of ergothioneine, comprising a nucleic acid sequence encoding an amino acid sequence selected from the group consisting of EgtB, EgtC, EgtD and EgtE.
BRIEF DESCRIPTION OF THE DRAWINGS
[0011] The disclosure will be better understood, and features, aspects and advantages other than those set forth above will become apparent when consideration is given to the following detailed description thereof. Such detailed description makes reference to the following drawings, wherein:
[0012] FIG. 1A is a vector map containing the EgtD and EgtB genes, as discussed in Example 1.
[0013] FIG. 1B is a vector map containing the EgtC and EgtE genes, as discussed in Example 1.
[0014] FIG. 2 is a graph illustrating the production of ET only in the strain containing all four genes, as discussed in Example 2. EI, empty vector cells induced with IPTG; SI, strain containing the four genes induced with IPTG; Ck+, the sample with the addition of 20 mg/L ergothioneine.
[0015] FIGS. 3A and 3B are graphs showing the HPLC retention time and UV-spectrum of a 100 mg/L ergothioneine standard, as discussed in Example 2.
[0016] FIGS. 4A and 4B are graphs showing the HPLC retention time and UV-spectrum of ergothioneine produced in E. coli transformed with nucleic acid sequences encoding EgtB, EgtC, EgtD and EgtE, as discussed in Example 2.
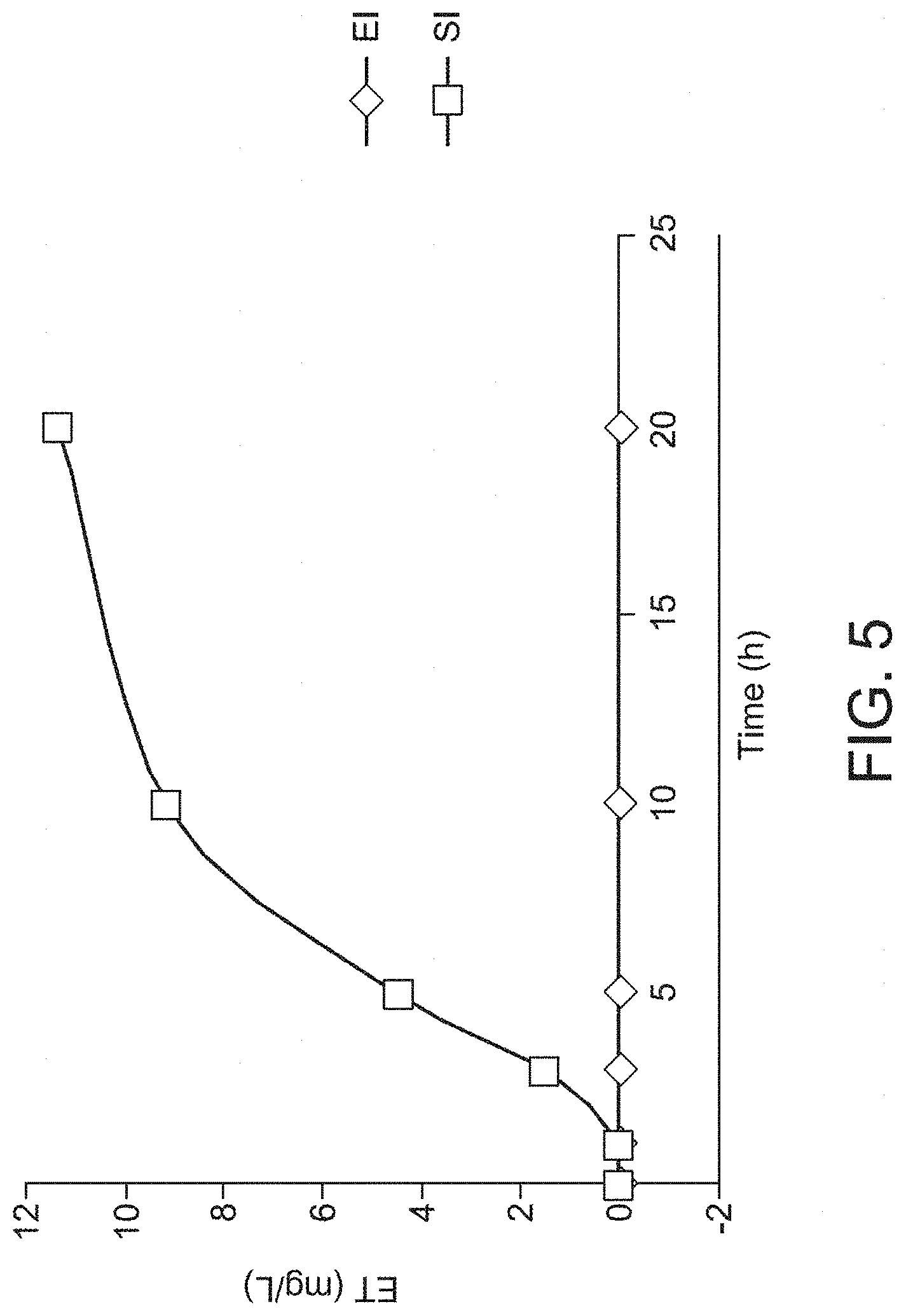
[0017] FIG. 5 is a graph showing a time course of ergothioneine production in an engineered E. coli cell and empty vector control cell, as discussed in Example 3. EI, empty vector control induced with IPTG; SI, strain containing EgtB, EgtC, EgtD and EgtE induced with IPTG.
[0018] FIG. 6 is a graph showing the transformed E. coli strain fed with various substrates and cofactor. No, no added substrates or cofactor; His, histidine; Met, methionine; Cys, cysteine; Fe, iron Fe.sup.++.
[0019] While the disclosure is susceptible to various modifications and alternative forms, specific embodiments thereof have been shown by way of example in the drawings and are herein described below in detail. It should be understood, however, that the description of specific embodiments is not intended to limit the disclosure to cover all modifications, equivalents and alternatives falling within the spirit and scope of the disclosure as defined by the appended claims.
DETAILED DESCRIPTION
[0020] The term "complementary" is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and is used without limitation to describe the relationship between nucleotide bases that are capable to hybridizing to one another. For example, with respect to DNA, adenosine is complementary to thymine, and cytosine is complementary to guanine. Accordingly, the subjection technology also includes isolated nucleic acid fragments that are complementary to the complete sequences as reported in the accompanying Sequence Listing as well as those substantially similar nucleic acid sequences.
[0021] The terms "nucleic acid" and "nucleotide" are used according to their respective ordinary and customary meanings as understood by a person of ordinary skill in the art, and are used without limitation to refer to deoxyribonucleotides or ribonucleotides and polymers thereof in either single- or double-stranded form. Unless specifically limited, the term encompasses nucleic acids containing known analogues of natural nucleotides that have similar binding properties as the reference nucleic acid and are metabolized in a manner similar to naturally-occurring nucleotides. Unless otherwise indicated, a particular nucleic acid sequence also implicitly encompasses conservatively modified or degenerate variants thereof (e.g., degenerate codon substitutions) and complementary sequences, as well as the sequence explicitly indicated.
[0022] The term "isolated" is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and when used in the context of an isolated nucleic acid or an isolated polypeptide, is used without limitation to refer to a nucleic acid or polypeptide that, by the hand of man, exists apart from its native environment and is therefore not a product of nature. An isolated nucleic acid or polypeptide can exist in a purified form or can exist in a non-native environment such as, for example, in a transgenic host cell.
[0023] The terms "incubating" and "incubation" as used herein refers to a process of mixing two or more chemical or biological entities (such as a chemical compound and an enzyme) and allowing them to interact under conditions favorable for producing a steviol glycoside composition.
[0024] The term "degenerate variant" refers to a nucleic acid sequence having a residue sequence that differs from a reference nucleic acid sequence by one or more degenerate codon substitutions. Degenerate codon substitutions can be achieved by generating sequences in which the third position of one or more selected (or all) codons is substituted with mixed base and/or deoxyinosine residues. A nucleic acid sequence and all of its degenerate variants will express the same amino acid or polypeptide.
[0025] The terms "polypeptide," "protein," and "peptide" are used according to their respective ordinary and customary meanings as understood by a person of ordinary skill in the art; the three terms are sometimes used interchangeably, and are used without limitation to refer to a polymer of amino acids, or amino acid analogs, regardless of its size or function. Although "protein" is often used in reference to relatively large polypeptides, and "peptide" is often used in reference to small polypeptides, usage of these terms in the art overlaps and varies. The term "polypeptide" as used herein refers to peptides, polypeptides, and proteins, unless otherwise noted. The terms "protein," "polypeptide," and "peptide" are used interchangeably herein when referring to a polynucleotide product. Thus, exemplary polypeptides include polynucleotide products, naturally occurring proteins, homologs, orthologs, paralogs, fragments and other equivalents, variants, and analogs of the foregoing.
[0026] The terms "polypeptide fragment" and "fragment," when used in reference to a reference polypeptide, are used according to their ordinary and customary meanings to a person of ordinary skill in the art, and are used without limitation to refer to a polypeptide in which amino acid residues are deleted as compared to the reference polypeptide itself, but where the remaining amino acid sequence is usually identical to the corresponding positions in the reference polypeptide. Such deletions can occur at the amino-terminus or carboxy-terminus of the reference polypeptide, or alternatively both.
[0027] The term "functional fragment" of a polypeptide or protein refers to a peptide fragment that is a portion of the full length polypeptide or protein, and has substantially the same biological activity, or carries out substantially the same function as the full length polypeptide or protein (e.g., carrying out the same enzymatic reaction).
[0028] The terms "variant polypeptide," "modified amino acid sequence" or "modified polypeptide," which are used interchangeably, refer to an amino acid sequence that is different from the reference polypeptide by one or more amino acids, e.g., by one or more amino acid substitutions, deletions, and/or additions. In an aspect, a variant is a "functional variant" which retains some or all of the ability of the reference polypeptide.
[0029] The term "functional variant" further includes conservatively substituted variants. The term "conservatively substituted variant" refers to a peptide having an amino acid sequence that differs from a reference peptide by one or more conservative amino acid substitutions, and maintains some or all of the activity of the reference peptide. A "conservative amino acid substitution" is a substitution of an amino acid residue with a functionally similar residue. Examples of conservative substitutions include the substitution of one non-polar (hydrophobic) residue such as isoleucine, valine, leucine or methionine for another; the substitution of one charged or polar (hydrophilic) residue for another such as between arginine and lysine, between glutamine and asparagine, between threonine and serine; the substitution of one basic residue such as lysine or arginine for another; or the substitution of one acidic residue, such as aspartic acid or glutamic acid for another; or the substitution of one aromatic residue, such as phenylalanine, tyrosine, or tryptophan for another. Such substitutions are expected to have little or no effect on the apparent molecular weight or isoelectric point of the protein or polypeptide. The phrase "conservatively substituted variant" also includes peptides wherein a residue is replaced with a chemically-derivatized residue, provided that the resulting peptide maintains some or all of the activity of the reference peptide as described herein.
[0030] The term "variant," in connection with the polypeptides of the subject technology, further includes a functionally active polypeptide having an amino acid sequence at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, and even 100% identical to the amino acid sequence of a reference polypeptide.
[0031] The term "homologous" in all its grammatical forms and spelling variations refers to the relationship between polynucleotides or polypeptides that possess a "common evolutionary origin," including polynucleotides or polypeptides from superfamilies and homologous polynucleotides or proteins from different species (Reeck et al., Cell 50:667, 1987). Such polynucleotides or polypeptides have sequence homology, as reflected by their sequence similarity, whether in terms of percent identity or the presence of specific amino acids or motifs at conserved positions. For example, two homologous polypeptides can have amino acid sequences that are at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, and even 100% identical.
[0032] "Percent (%) amino acid sequence identity" with respect to the variant polypeptide sequences of the subject technology refers to the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues of a reference polypeptide (such as, for example, SEQ ID NO:6), after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity.
[0033] Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN, ALIGN-2 or Megalign (DNASTAR) software. Those skilled in the art can determine appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full-length of the sequences being compared. For example, the % amino acid sequence identity may be determined using the sequence comparison program NCBI-BLAST2. The NCBI-BLAST2 sequence comparison program may be downloaded from ncbi.nlm.nih.gov. NCBI BLAST2 uses several search parameters, wherein all of those search parameters are set to default values including, for example, unmask yes, strand=all, expected occurrences 10, minimum low complexity length=15/5, multi-pass e-value=0.01, constant for multi-pass=25, dropoff for final gapped alignment=25 and scoring matrix=BLOSUM62. In situations where NCBI-BLAST2 is employed for amino acid sequence comparisons, the % amino acid sequence identity of a given amino acid sequence A to, with, or against a given amino acid sequence B (which can alternatively be phrased as a given amino acid sequence A that has or comprises a certain % amino acid sequence identity to, with, or against a given amino acid sequence B) is calculated as follows: 100 times the fraction X/Y where X is the number of amino acid residues scored as identical matches by the sequence alignment program NCBI-BLAST2 in that program's alignment of A and B, and where Y is the total number of amino acid residues in B. It will be appreciated that where the length of amino acid sequence A is not equal to the length of amino acid sequence B, the % amino acid sequence identity of A to B will not equal the % amino acid sequence identity of B to A.
[0034] In this sense, techniques for determining amino acid sequence "similarity" are well known in the art. In general, "similarity" refers to the exact amino acid to amino acid comparison of two or more polypeptides at the appropriate place, where amino acids are identical or possess similar chemical and/or physical properties such as charge or hydrophobicity. A so-termed "percent similarity" may then be determined between the compared polypeptide sequences. Techniques for determining nucleic acid and amino acid sequence identity also are well known in the art and include determining the nucleotide sequence of the mRNA for that gene (usually via a cDNA intermediate) and determining the amino acid sequence encoded therein, and comparing this to a second amino acid sequence. In general, "identity" refers to an exact nucleotide to nucleotide or amino acid to amino acid correspondence of two polynucleotides or polypeptide sequences, respectively. Two or more polynucleotide sequences can be compared by determining their "percent identity", as can two or more amino acid sequences. The programs available in the Wisconsin Sequence Analysis Package, Version 8 (available from Genetics Computer Group, Madison, Wis.), for example, the GAP program, are capable of calculating both the identity between two polynucleotides and the identity and similarity between two polypeptide sequences, respectively. Other programs for calculating identity or similarity between sequences are known by those skilled in the art.
[0035] An amino acid position "corresponding to" a reference position refers to a position that aligns with a reference sequence, as identified by aligning the amino acid sequences. Such alignments can be done by hand or by using well-known sequence alignment programs such as ClustalW2, Blast 2, etc.
[0036] Unless specified otherwise, the percent identity of two polypeptide or polynucleotide sequences refers to the percentage of identical amino acid residues or nucleotides across the entire length of the shorter of the two sequences.
[0037] "Coding sequence" is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and is used without limitation to refer to a DNA sequence that encodes for a specific amino acid sequence.
[0038] "Suitable regulatory sequences" is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and is used without limitation to refer to nucleotide sequences located upstream (5' non-coding sequences), within, or downstream (3' non-coding sequences) of a coding sequence, and which influence the transcription, RNA processing or stability, or translation of the associated coding sequence. Regulatory sequences may include promoters, translation leader sequences, introns, and polyadenylation recognition sequences.
[0039] "Promoter" is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and is used without limitation to refer to a DNA sequence capable of controlling the expression of a coding sequence or functional RNA. In general, a coding sequence is located 3' to a promoter sequence. Promoters may be derived in their entirety from a native gene, or be composed of different elements derived from different promoters found in nature, or even comprise synthetic DNA segments. It is understood by those skilled in the art that different promoters may direct the expression of a gene in different cell types, or at different stages of development, or in response to different environmental conditions. Promoters that cause a gene to be expressed in most cell types at most times are commonly referred to as "constitutive promoters." It is further recognized that since, in most cases, the exact boundaries of regulatory sequences have not been completely defined, DNA fragments of different lengths may have identical promoter activity.
[0040] The term "operably linked" refers to the association of nucleic acid sequences on a single nucleic acid fragment so that the function of one is affected by the other. For example, a promoter is operably linked with a coding sequence when it is capable of affecting the expression of that coding sequence (i.e., that the coding sequence is under the transcriptional control of the promoter). Coding sequences can be operably linked to regulatory sequences in sense or antisense orientation.
[0041] The term "expression" as used herein, is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and is used without limitation to refer to the transcription and stable accumulation of sense (mRNA) or antisense RNA derived from the nucleic acid fragment of the subject technology. "Over-expression" refers to the production of a gene product in transgenic or recombinant organisms that exceeds levels of production in normal or non-transformed organisms.
[0042] "Transformation" is used according to its ordinary and customary meaning as understood by a person of ordinary skill in the art, and is used without limitation to refer to the transfer of a polynucleotide into a target cell. The transferred polynucleotide can be incorporated into the genome or chromosomal DNA of a target cell, resulting in genetically stable inheritance, or it can replicate independent of the host chromosomal. Host organisms containing the transformed nucleic acid fragments are referred to as "transgenic" or "recombinant" or "transformed" organisms.
[0043] The terms "transformed," "transgenic," and "recombinant," when used herein in connection with host cells, are used according to their ordinary and customary meanings as understood by a person of ordinary skill in the art, and are used without limitation to refer to a cell of a host organism, such as a plant or microbial cell, into which a heterologous nucleic acid molecule has been introduced. The nucleic acid molecule can be stably integrated into the genome of the host cell, or the nucleic acid molecule can be present as an extrachromosomal molecule. Such an extrachromosomal molecule can be auto-replicating. Transformed cells, tissues, or subjects are understood to encompass not only the end product of a transformation process, but also transgenic progeny thereof.
[0044] The terms "recombinant," "heterologous," and "exogenous," when used herein in connection with polynucleotides, are used according to their ordinary and customary meanings as understood by a person of ordinary skill in the art, and are used without limitation to refer to a polynucleotide (e.g., a DNA sequence or a gene) that originates from a source foreign to the particular host cell or, if from the same source, is modified from its original form. Thus, a heterologous gene in a host cell includes a gene that is endogenous to the particular host cell but has been modified through, for example, the use of site-directed mutagenesis or other recombinant techniques. The terms also include non-naturally occurring multiple copies of a naturally occurring DNA sequence. Thus, the terms refer to a DNA segment that is foreign or heterologous to the cell, or homologous to the cell but in a position or form within the host cell in which the element is not ordinarily found.
[0045] Similarly, the terms "recombinant," "heterologous," and "exogenous," when used herein in connection with a polypeptide or amino acid sequence, means a polypeptide or amino acid sequence that originates from a source foreign to the particular host cell or, if from the same source, is modified from its original form. Thus, recombinant DNA segments can be expressed in a host cell to produce a recombinant polypeptide.
[0046] The terms "plasmid," "vector," and "cassette" are used according to their ordinary and customary meanings as understood by a person of ordinary skill in the art, and are used without limitation to refer to an extra chromosomal element often carrying genes which are not part of the central metabolism of the cell, and usually in the form of circular double-stranded DNA molecules. Such elements may be autonomously replicating sequences, genome integrating sequences, phage or nucleotide sequences, linear or circular, of a single- or double-stranded DNA or RNA, derived from any source, in which a number of nucleotide sequences have been joined or recombined into a unique construction which is capable of introducing a promoter fragment and DNA sequence for a selected gene product along with appropriate 3' untranslated sequence into a cell. "Transformation cassette" refers to a specific vector containing a foreign gene and having elements in addition to the foreign gene that facilitate transformation of a particular host cell. "Expression cassette" refers to a specific vector containing a foreign gene and having elements in addition to the foreign gene that allow for enhanced expression of that gene in a foreign host.
[0047] Standard recombinant DNA and molecular cloning techniques used herein are well known in the art and are described, for example, by Sambrook, J., Fritsch, E. F. and Maniatis, T. Molecular Cloning: A Laboratory Manual, 2.sup.nd ed.; Cold Spring Harbor Laboratory: Cold Spring Harbor, N.Y., 1989 (hereinafter "Maniatis"); and by Silhavy, T. J., Bennan, M. L. and Enquist, L. W. Experiments with Gene Fusions; Cold Spring Harbor Laboratory: Cold Spring Harbor, N.Y., 1984; and by Ausubel, F. M. et al., In Current Protocols in Molecular Biology, published by Greene Publishing and Wiley-Interscience, 1987; the entireties of each of which are hereby incorporated herein by reference to the extent they are consistent herewith.
[0048] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which the disclosure belongs. Although any methods and materials similar to or equivalent to those described herein may be used in the practice or testing of the present disclosure, the preferred materials and methods are described below.
[0049] In accordance with the present disclosure, methods have been developed for producing ergothioneine and host cells having genes encoding EgtB, EgtC, EgtD and EgtE that are useful for producing ergothioneine. Surprisingly and unexpectedly, an ergothioneine production pathway has been reproduced in an in vitro microbial production system.
Engineered Host Cell for Producing Ergothioneine
[0050] In one aspect, the present disclosure is directed to an engineered host cell. The engineered host cell includes a nucleic acid sequence encoding EgtB, a nucleic acid sequence encoding EgtC, a nucleic acid sequence encoding EgtD and a nucleic acid sequence encoding EgtE.
[0051] EgtB (or iron(II)-dependent oxidoreductase EgtB) catalyzes the oxidative sulfurization of hercynine via the addition of oxygen and gamma-glutamyl-cysteine on hercynine (N-alpha,N-alpha,N-alpha-trimethyl-L-histidine).
[0052] A suitable EgtB can be, for example, Mycobacterium EgtB. A particularly suitable EgtB can be, for example, an EgtB nucleic acid sequence encoding an amino acid sequence at least 95% identical to the amino acid sequence provided in SEQ ID NO:2. In another aspect, a particularly suitable EgtB can be, for example, an EgtB nucleic acid sequence encoding an amino acid sequence at least 96% identical to the amino acid sequence provided in SEQ ID NO:2. In another aspect, a particularly suitable EgtB can be, for example, an EgtB nucleic acid sequence encoding an amino acid sequence at least 97% identical to the amino acid sequence provided in SEQ ID NO:2. In another aspect, a particularly suitable EgtB can be, for example, an EgtB nucleic acid sequence encoding an amino acid sequence at least 98% identical to the amino acid sequence provided in SEQ ID NO:2. In another aspect, a particularly suitable EgtB can be, for example, an EgtB nucleic acid sequence encoding an amino acid sequence at least 99% identical to the amino acid sequence provided in SEQ ID NO:2. In another aspect, a particularly suitable EgtB can be, for example, an EgtB nucleic acid sequence encoding an amino acid sequence 100% identical to the amino acid sequence provided in SEQ ID NO:2.
[0053] EgtC (or Amidohydrolase EgtC) catalyzes the hydrolysis of the gamma-glutamyl amide bond from N-(gamma-glutamyl)-[N(alpha),N(alpha), N(alpha)-trimethyl-L-histidinyl]-cysteine sulfoxide to produce hercynylcysteine sulfoxide.
[0054] A suitable EgtC can be, for example, Mycobacterium EgtC. A particularly suitable EgtC can be, for example, an EgtC nucleic acid sequence encoding an amino acid sequence at least 95% identical to the amino acid sequence provided in SEQ ID NO:4. In another aspect, a particularly suitable EgtC can be, for example, an EgtC nucleic acid sequence encoding an amino acid sequence at least 96% identical to the amino acid sequence provided in SEQ ID NO:4. In another aspect, a particularly suitable EgtC can be, for example, an EgtC nucleic acid sequence encoding an amino acid sequence at least 97% identical to the amino acid sequence provided in SEQ ID NO:4. In another aspect, a particularly suitable EgtC can be, for example, an EgtC nucleic acid sequence encoding an amino acid sequence at least 98% identical to the amino acid sequence provided in SEQ ID NO:4. In another aspect, a particularly suitable EgtC can be, for example, an EgtC nucleic acid sequence encoding an amino acid sequence at least 99% identical to the amino acid sequence provided in SEQ ID NO:4. In another aspect, a particularly suitable EgtC can be, for example, an EgtC nucleic acid sequence encoding an amino acid sequence 100% identical to the amino acid sequence provided in SEQ ID NO:4.
[0055] EgtD (or histidine-specific methyltransferase EgtD) catalyzes the methylations of histidine to form N-alpha,N-alpha,N-alpha-trimethyl-L-histidine (also known as hercynine). Histidine and alpha-N,N-dimethylhistidine are preferred substrates.
[0056] A suitable EgtD can be, for example, Mycobacterium EgtD. A particularly suitable EgtD can be, for example, an EgtD nucleic acid sequence encoding an amino acid sequence at least 95% identical to the amino acid sequence provided in SEQ ID NO:6. In another aspect, a particularly suitable EgtD can be, for example, an EgtD nucleic acid sequence encoding an amino acid sequence at least 96% identical to the amino acid sequence provided in SEQ ID NO:6. In another aspect, a particularly suitable EgtD can be, for example, an EgtD nucleic acid sequence encoding an amino acid sequence at least 97% identical to the amino acid sequence provided in SEQ ID NO:6. In another aspect, a particularly suitable EgtD can be, for example, an EgtD nucleic acid sequence encoding an amino acid sequence at least 98% identical to the amino acid sequence provided in SEQ ID NO:6. In another aspect, a particularly suitable EgtD can be, for example, an EgtD nucleic acid sequence encoding an amino acid sequence at least 99% identical to the amino acid sequence provided in SEQ ID NO:6. In another aspect, a particularly suitable EgtD can be, for example, an EgtD nucleic acid sequence encoding an amino acid sequence 100% identical to the amino acid sequence provided in SEQ ID NO:6.
[0057] EgtE (or pyridoxal-phosphate-dependent protein EgtE) is believed to catalyze the removing of pyruvate, ammonia and oxygen to produce ergothioneine.
[0058] A suitable EgtE can be, for example, Mycobacterium EgtE. A particularly suitable EgtE can be, for example, an EgtE nucleic acid sequence encoding an amino acid sequence at least 95% identical to the amino acid sequence provided in SEQ ID NO:8. In another aspect, a particularly suitable EgtE can be, for example, an EgtE nucleic acid sequence encoding an amino acid sequence at least 96% identical to the amino acid sequence provided in SEQ ID NO:8. In another aspect, a particularly suitable EgtE can be, for example, an EgtE nucleic acid sequence encoding an amino acid sequence at least 97% identical to the amino acid sequence provided in SEQ ID NO:8. In another aspect, a particularly suitable EgtE can be, for example, an EgtE nucleic acid sequence encoding an amino acid sequence at least 98% identical to the amino acid sequence provided in SEQ ID NO:8. In another aspect, a particularly suitable EgtE can be, for example, an EgtE nucleic acid sequence encoding an amino acid sequence at least 99% identical to the amino acid sequence provided in SEQ ID NO:8. In another aspect, a particularly suitable EgtE can be, for example, an EgtE nucleic acid sequence encoding an amino acid sequence 100% identical to the amino acid sequence provided in SEQ ID NO:8.
[0059] Suitable host cells can be, for example, bacterial cells and yeast cells. Suitable bacterial cells can be, for example, Escherichia coli.
[0060] Suitable yeast cells can be, for example, Saccharomyces and Pichia. Particularly suitable Saccharomyces can be, for example, Saccharomyces cerevisiae. Particularly suitable Pichia can be, for example, Pichia pastoris.
[0061] The nucleic acid sequences encoding EgtB, EgtC, EgtD and EgtE are cloned into an expression vector under the control of a promoter known by those skilled in the art. Suitable promoters can be, for example, constitutively active promoters and inducible promoters known by those skilled in the art. Suitable inducible promoters are known by those skilled in the art and can be, for example, chemical inducers, nutrient addition, nutrient depletion and physical or physiochemical factor shift such as, for example a pH shift and temperature-induction. Suitable chemical inducers can be, for example, an isopropyl .beta.-D-1-thiogalactopyranoside (IPTG)-inducible promoter and antibiotic-inducible promoters known by those skilled in the art. A particularly suitable chemically-inducible promoter can be, for example, an isopropyl .beta.-D-1-thiogalactopyranoside (IPTG)-inducible promoter known by those skilled in the art. Other suitable inducible promoters can be, for example, a temperature-induced promoter known by those skilled in the art such as, for example, pL and pR .lamda. phage promoters.
[0062] Particularly suitable expression vectors are illustrated in FIGS. 1A and 1B. Other suitable expression vectors are known to those in the art and can be, for example, pET vectors, pCDF vectors, pRSF vectors and Duet vectors.
Methods for Producing Ergothioneine
[0063] In another aspect, the present disclosure is directed to a method for producing ergothioneine. The method includes culturing an host cell, wherein the host cell is transformed with a nucleic acid sequence encoding EgtB, a nucleic acid sequence encoding EgtC, a nucleic acid sequence encoding EgtD and a nucleic acid sequence encoding EgtE; inducing the host cell to express the nucleic acid sequence encoding EgtB, the nucleic acid sequence encoding EgtC, the nucleic acid sequence encoding EgtD and the nucleic acid sequence encoding EgtE; and collecting the ergothioneine.
[0064] The method can further include adding a substrate to the culture. Suitable amounts of substrates can be, for example, from about 1 mM to about 20 mM. Particularly suitable substrates can be, for example, histidine, methionine, cysteine, .gamma.-glutamyl cysteine and combinations thereof.
[0065] In another embodiment, the method can include adding a cofactor to the culture. Suitable amounts of cofactors can be, for example, from about 0.05 mM to about 0.4 mM. A particularly suitable cofactor can be, for example iron (II) (Fe.sup.++).
[0066] Suitable host cells can be, for example, bacterial cells and yeast cells. Suitable bacterial cells can be, for example, Escherichia coli.
[0067] Suitable yeast cells can be, for example, Saccharomyces and Pichia. Particularly suitable Saccharomyces can be, for example, Saccharomyces cerevisiae. Particularly suitable Pichia can be, for example, Pichia pastoris.
[0068] In one embodiment, the host cell can produce from about 10 milligrams to about 30 milligrams of ergothioneine per liter.
[0069] The disclosure will be more fully understood upon consideration of the following non-limiting Examples.
EXAMPLES
Example 1
[0070] In this Example, nucleic acid sequences for EgtB, EgtC, EgtD and EgtE were cloned into E. coli.
[0071] Specifically, the following sequences were obtained from GenBank (accession number NC 008596): Egt B: MSMEG_6249 (SEQ ID NO:1); Egt C: MSMEG_6248 (SEQ ID NO:3); Egt D: MSMEG_ 6247 (SEQ ID NO:5); and Egt E: MSMEG_6246(SEQ ID NO:7). The genes were introduced into a vector under control of an IPTG-inducible promoter.
[0072] To build the ET pathway in E. coli, the EgtB, C, D, E nucleic acid sequences were PCR-amplified from the genomic sequence of M. smegmatis using the primer pairs summarized in Table 1. All of the 5'-primers used for cloning included EcoRI and BglI restriction sites and a ribosomal binding site (RBS) and all of the 3'-primers included BamHI--Xhol sites. The EgtD and EgtB sequences were cloned into a pConB7A vector (FIG. 1A) and the EgtC and EgtE sequences were cloned into a pConA5K vector (FIG. 1B). No sequence errors were identified in the cloned sequences. The empty vectors were prepared in the same manner. The constructs were then co-transformed into E. coli strain BL21(DE3).
TABLE-US-00001 TABLE 1 Primers for gene cloning. SEQ ID Primer Sequence NO EgtB-5' AGAATTCAAAAGATCTAAAGGAGGCCATCCATGATCG 9 GCACCGAGACAC EgtB-3' ACTCGAGTTTGGATCCTCAGACGTCCCAGGCCAGGCG 10 GACACCCGAGAATATC EgtC-5' AGAATTCAAAAGATCTAAAGGAGGCCATCCATGTGCC 11 GGCATGTGGCGTG EgtC-3' ACTCGAGTTTGGATCCTCACAGGGGTGTCACGAC 12 EgtD-5' AGAATTCAAAAGATCTAAAGGAGGCCATCCATGACGC 13 TCTCACTGGCCAAC EgtD-3' ACTCGAGTTTGGATCCTCACCGCACCGCCAGCGAC 14 EgtE-5' AGAATTCAAAAGATCTAAAGGAGGCCATCCATGCTCG 15 CGCAGCAGTG EgtE-3' ACTCGAGTTTGGATCCTCAGGGCGCCTCACGCAAC 16
Example 2
[0073] In this Example, ergothioneine was produced in an engineered microbial system.
[0074] Specifically, E. coli were transformed with the pConB7A vector and the pConA5K vectors encoding EgtB, EgtC, EgtD and EgtE as described in Example 1. To co-express the four genes (EgtB, C, D, E) in E. coli system, the transformants were grown in the LB media containing 100 mg/L ampicillin and 50 mg/L kanamycin at 37.degree. C. until reaching an OD.sub.600.about.0.6. Expression was induced by the addition of 0.2-0.5 mM of isopropyl .beta.-D-1-thiogalactopyranoside (IPTG) and the culture was further grown at either 30.degree. C. or 37.degree. C. for 16-24 hours. Cells were harvested by centrifugation and the supernatant and cell pellet were collected separately. The supernatant was centrifuged at 16,000.times.g for 5 min for HPLC analysis. The pellet was resuspended in 1 ml of 50% methanol and sonicated for 1 minute (3.times.20 sec). After centrifuging at 16,000.times.g for 5 minutes, a 5 .mu.l of sample was analyzed by HPLC, as described below. E. coli that were transformed with the empty vector were treated in the same manner and analyzed by HPLC. A sample obtained from IPTG-induced E. coli containing the EgtB, EgtC, EgtD, EgtE genes was spiked with 20 mg/L ergothioneine and analyzed by HPLC.
[0075] Samples were analyzed using a Dionex UPLC Ultimate 3000 (Sunnyvale, Calif.). The compounds were separated on an Atlantis HILIC Silica column (particle size 3.0 .mu.m, diameter.times.length=2.1.times.100 mm; Waters) and detected at 264 nm. The mobile phase consisted of 0.1% formic acid in water (A) and 0.1% formic acid in acetonitrile (B). The program of a gradient was 95% B at 1 min, 40% B at 8 minutes, 95% B at 8.1 minutes, stop at 11 min. The flow rate was 0.6 ml/minute and the inject volume was 5 .mu.l.
[0076] As shown in FIG. 2, ET surprisingly accumulated only in the IPTG-induced E. coli strain containing the EgtB, EgtC, EgtD, EgtE sequences ("SI"), successfully demonstrating biosynthesis of ET in engineered E. coli. In contrast, the IPTG-induced E. coli containing the empty vector did not produce any ET ("EI"). In the ET-spiked sample, the ET peak from the IPTG-induced E. coli strain containing the EgtB, EgtC, EgtD and EgtE overlapped with the added ergothioneine and demonstrated an increased level to account for the added ET ("Ck+").
[0077] FIGS. 3A and 3B illustrate the HPLC analysis of a 100 mg/L ergothioneine standard. As shown in FIG. 4A, the retention time of the ET from the E. coli strain containing EgtB, EgtC, EgtD and EgtE overlapped with the retention time of the ergothioneine standard (see, FIG. 3A). In addition to retention time, the UV-spectrum of the ET peak (see, FIG. 4B) also matched the ergothioneine standard (see, FIG. 3B). These results demonstrated that the peak from the engineered E. coli strain expressing the EgtB, EgtC, EgtD and EgtE corresponds to ET.
Example 3
[0078] In this Example, a time course for ergothioneine production in an engineered microbial system was performed.
[0079] Specifically, E. coli were transformed with the vectors containing genes for EgtB, EgtC, EgtD and EgtE as described in Example 1. Control E. coli cells included cells with an empty vector (no Egt genes) and a non-induced strain that contained the Egt vectors, but was not induced. Cells were grown at 30.degree. C. or 37.degree. C. as described in Example 2. Samples were taken at different time points from 0 hours to 20 hours. After sonicating for 1 minute (3.times.20 seconds), the samples were centrifuged at 16,000.times.g for 5 minutes and a 5 .mu.l sample was analyzed by HPLC as discussed in Example 2.
[0080] As shown in FIG. 5, the HPLC analysis revealed that ET started to be produced by the cells about 1 hour after the IPTG induction. The fastest increase in ET production was observed from about 3 hours up to about 10 hours after the IPTG induction. ET production slowed down after 10 hours, but continued to be produced at least until 20 hours. At the same time, no ET was detected in the empty vector control at all during the entire time course. These results further demonstrated that ET is exclusively produced in the E.coli strain engineered to express EgtB, EgtC, EgtD and EgtE.
Example 4
[0081] In this Example, feeding experiments were performed to determine the effect on ergothioneine production in the engineered microbial system.
[0082] Without being bound by theory, it is believed that ET is synthesized from amino acids such as histidine (His), methionine (Met), and cysteine (Cys). The imidazole ring of ET is supplied by His, which is then methylated to produce histidine betaine. Met is a building block for S-adenosyl methionine (SAM) that serves as a methyl donor. The sulfur atom is incorporated from Cys.
[0083] To determine the effect on ergothioneine production in the engineered E. coli, several substrates and cofactors such as Fe++ were fed to the transgenic E. coli cells through the culture media. After 3 hours of induction, 2 mM of His, 4 mM of Met, 4 mM of Cys and 0.2 mM of Fe++ were added into media and the cells were further cultured for 16 hours, 24 hours and 42 hours. A control E. coli culture (carrying empty vectors) was fed with the same substrates or cofactor. Samples were analyzed by HPLC as discussed in Example 2.
[0084] As shown in FIG. 6, the feeding experiments revealed that the addition of Cys increased ET yield by 17.3-44.4% among three time points. This result suggested that Cys and its derivative .gamma.-glutamyl cysteine play an important role in the biosynthesis of ET. The control culture did not produce any ET.
Example 5
[0085] In this Example, ergothioneine will be produced in an engineered S. cerevisiae yeast system.
[0086] To produce the ET in S. cerevisiae, the EgtB, C, D, E genes will be cloned into pESC vectors such as pESC-His and pESC-Leu, which are commercially available (Agilent Technologies). These vectors contain the GAL1 and GAL10 yeast promoters in opposing orientation, which allow for the introduction of two genes into a yeast strain under the control of two repressible promoters, respectively. The resulting two constructs will then be co-transformed into S. cerevisiae. To co-express the four genes (EgtB, C, D, E) in yeast, the transformants will be grown in media without the two amino acids, histidine and leucine, until reaching an OD.sub.600.about.0.4. Expression will be induced by the addition of 2% galactose and the culture will be further grown at either 28.degree. C. or 30.degree. C. for 24-48 hours. Cells will be harvested by centrifuge and the supernatant and cell pellet will be collected separately. The supernatant will be centrifuged at 12,000.times.g for 5 minutes and analyzed by HPLC. The pellet will be resuspended in 1 ml of 50% methanol and sonicated for 1 min (3.times.20 sec). After centrifuging at 12,000.times.g for 5 minutes, 5.mu.l of sample will be injected to HPLC. Yeast harboring empty vectors will be transformed and analyzed in the same way. The above constructs may be ultimately integrated into the yeast genome and expressed under the control of constitutive promoters such as the GPD promoter or the GAP promoter.
Example 6
[0087] In this Example, ergothioneine will be produced in an engineered P. pastoris yeast system.
[0088] To produce the ET in P. pastoris, the EgtB, C, D, E genes will be cloned into pPICZ or pGAPZ vectors, which are commercially available (Invitrogen, Life Technologies). The pPICZ vectors contain the methanol-regulated AOX1 promoter, while pGAPZ vectors have constitutive glyceraldehyde-3-phosphate dehydrogenase (GAP) promoter. The coexpression of the four genes (EgtB, C, D, E) in pPICZ vectors will be induced by 0.5-5% methanol. The production of ET will be analyzed by HPLC analysis using the same method described above.
[0089] In view of the above, it will be seen that the several advantages of the disclosure are achieved and other advantageous results attained. As various changes could be made in the above methods and systems without departing from the scope of the disclosure, it is intended that all matter contained in the above description and shown in the accompanying drawings shall be interpreted as illustrative and not in a limiting sense.
[0090] When introducing elements of the present disclosure or the various versions, embodiment(s) or aspects thereof, the articles "a", "an", "the" and "said" are intended to mean that there are one or more of the elements. The terms "comprising", "including" and "having" are intended to be inclusive and mean that there may be additional elements other than the listed elements.
Sequence CWU 1
1
1611287DNAArtificial Sequencesynthetic 1atgatcgcac gcgagacact
ggccgacgag ctggccctgg cccgcgaacg cacgttgcgg 60ctcgtggagt tcgacgacgc
ggaactgcat cgccagtaca acccgctgat gagcccgctc 120gtgtgggacc
tcgcgcacat cgggcagcag gaagaactgt ggctgctgcg cgacggcaac
180cccgaccgcc ccggcatgct cgcacccgag gtggaccggc tttacgacgc
gttcgagcac 240tcacgcgcca gccgggtcaa cctcccgttg ctgccgcctt
cggatgcgcg cgcctactgc 300gcgacggtgc gggccaaggc gctcgacacc
ctcgacacgc tgcccgagga cgatccgggc 360ttccggttcg cgctggtgat
cagccacgag aaccagcacg acgagaccat gctgcaggca 420ctcaacctgc
gcgagggccc acccctgctc gacaccggaa ttcccctgcc cgcgggcagg
480ccaggcgtgg caggcacgtc ggtgctggtg ccgggcggcc cgttcgtgct
cggggtcgac 540gcgctgaccg aaccgcactc actggacaac gaacggcccg
cccacgtcgt ggacatcccg 600tcgttccgga tcggccgcgt gccggtcacc
aacgccgaat ggcgcgagtt catcgacgac 660ggtggctacg accaaccgcg
ctggtggtcg ccacgcggct gggcgcaccg ccaggaggcg 720ggcctggtgg
ccccgcagtt ctggaacccc gacggcaccc gcacccggtt cgggcacatc
780gaggagatcc cgggtgacga acccgtgcag cacgtgacgt tcttcgaagc
cgaggcctac 840gcggcgtggg ccggtgctcg gttgcccacc gagatcgaat
gggagaaggc ctgcgcgtgg 900gatccggtcg ccggtgctcg gcgccggttc
ccctggggct cagcacaacc cagcgcggcg 960ctggccaacc tcggcggtga
cgcacgccgc ccggcgccgg tcggggccta cccggcgggg 1020gcgtcggcct
atggcgccga gcagatgctg ggcgacgtgt gggagtggac ctcctcgccg
1080ctgcggccgt ggcccggttt cacgccgatg atctacgagc gctacagcac
gccgttcttc 1140gagggcacca catccggtga ctaccgcgtg ctgcgcggcg
ggtcatgggc cgttgcaccg 1200ggaatcctgc ggcccagctt ccgcaactgg
gaccacccga tccggcggca gatattctcg 1260ggtgtccgcc tggcctggga cgtctga
12872428PRTArtificial Sequencesynthetic 2Met Ile Ala Arg Glu Thr
Leu Ala Asp Glu Leu Ala Leu Ala Arg Glu1 5 10 15Arg Thr Leu Arg Leu
Val Glu Phe Asp Asp Ala Glu Leu His Arg Gln 20 25 30Tyr Asn Pro Leu
Met Ser Pro Leu Val Trp Asp Leu Ala His Ile Gly 35 40 45Gln Gln Glu
Glu Leu Trp Leu Leu Arg Asp Gly Asn Pro Asp Arg Pro 50 55 60Gly Met
Leu Ala Pro Glu Val Asp Arg Leu Tyr Asp Ala Phe Glu His65 70 75
80Ser Arg Ala Ser Arg Val Asn Leu Pro Leu Leu Pro Pro Ser Asp Ala
85 90 95Arg Ala Tyr Cys Ala Thr Val Arg Ala Lys Ala Leu Asp Thr Leu
Asp 100 105 110Thr Leu Pro Glu Asp Asp Pro Gly Phe Arg Phe Ala Leu
Val Ile Ser 115 120 125His Glu Asn Gln His Asp Glu Thr Met Leu Gln
Ala Leu Asn Leu Arg 130 135 140Glu Gly Pro Pro Leu Leu Asp Thr Gly
Ile Pro Leu Pro Ala Gly Arg145 150 155 160Pro Gly Val Ala Gly Thr
Ser Val Leu Val Pro Gly Gly Pro Phe Val 165 170 175Leu Gly Val Asp
Ala Leu Thr Glu Pro His Ser Leu Asp Asn Glu Arg 180 185 190Pro Ala
His Val Val Asp Ile Pro Ser Phe Arg Ile Gly Arg Val Pro 195 200
205Val Thr Asn Ala Glu Trp Arg Glu Phe Ile Asp Asp Gly Gly Tyr Asp
210 215 220Gln Pro Arg Trp Trp Ser Pro Arg Gly Trp Ala His Arg Gln
Glu Ala225 230 235 240Gly Leu Val Ala Pro Gln Phe Trp Asn Pro Asp
Gly Thr Arg Thr Arg 245 250 255Phe Gly His Ile Glu Glu Ile Pro Gly
Asp Glu Pro Val Gln His Val 260 265 270Thr Phe Phe Glu Ala Glu Ala
Tyr Ala Ala Trp Ala Gly Ala Arg Leu 275 280 285Pro Thr Glu Ile Glu
Trp Glu Lys Ala Cys Ala Trp Asp Pro Val Ala 290 295 300Gly Ala Arg
Arg Arg Phe Pro Trp Gly Ser Ala Gln Pro Ser Ala Ala305 310 315
320Leu Ala Asn Leu Gly Gly Asp Ala Arg Arg Pro Ala Pro Val Gly Ala
325 330 335Tyr Pro Ala Gly Ala Ser Ala Tyr Gly Ala Glu Gln Met Leu
Gly Asp 340 345 350Val Trp Glu Trp Thr Ser Ser Pro Leu Arg Pro Trp
Pro Gly Phe Thr 355 360 365Pro Met Ile Tyr Glu Arg Tyr Ser Thr Pro
Phe Phe Glu Gly Thr Thr 370 375 380Ser Gly Asp Tyr Arg Val Leu Arg
Gly Gly Ser Trp Ala Val Ala Pro385 390 395 400Gly Ile Leu Arg Pro
Ser Phe Arg Asn Trp Asp His Pro Ile Arg Arg 405 410 415Gln Ile Phe
Ser Gly Val Arg Leu Ala Trp Asp Val 420 4253684DNAArtificial
Sequencesynthetic 3atgtgccggc atgtggcgtg gctgggcgcg ccgcggtcgt
tggccgacct ggtgctcgac 60ccgccgcagg gactgctggt gcagtcctac gcaccgcgac
gacagaagca cggtctgatg 120aacgccgacg gttggggcgc agggtttttc
gacgacgagg gagtggcccg ccgctggcgc 180agcgacaaac cgctgtgggg
tgatgcgtcg ttcgcgtcgg tggcacccgc actacgcagt 240cgttgcgtgc
tggccgcggt gcgctcggcc accatcggca tgcccatcga accgtcggcg
300tcggcgccgt tcagcgacgg gcagtggctg ctgtcgcaca acggcctggt
cgaccgcggg 360gtgctcccgt tgaccggtgc cgccgagtcc acggtggaca
gcgcgatcgt cgcggcgctc 420atcttctccc gtggcctcga cgcgctcggc
gccaccatcg ccgaggtcgg cgaactcgac 480ccgaacgcgc ggttgaacat
cctggccgcc aacggttccc ggctgctcgc caccacctgg 540ggggacacgc
tgtcggtcct gcaccgcccc gacggcgtcg tcctcgcgag cgaaccctac
600gacgacgatc ccggctggtc ggacatcccg gaccggcacc tcgtcgacgt
ccgcgacgcc 660cacgtcgtcg tgacacccct gtga 6844227PRTArtificial
Sequencesynthetic 4Met Cys Arg His Val Ala Trp Leu Gly Ala Pro Arg
Ser Leu Ala Asp1 5 10 15Leu Val Leu Asp Pro Pro Gln Gly Leu Leu Val
Gln Ser Tyr Ala Pro 20 25 30Arg Arg Gln Lys His Gly Leu Met Asn Ala
Asp Gly Trp Gly Ala Gly 35 40 45Phe Phe Asp Asp Glu Gly Val Ala Arg
Arg Trp Arg Ser Asp Lys Pro 50 55 60Leu Trp Gly Asp Ala Ser Phe Ala
Ser Val Ala Pro Ala Leu Arg Ser65 70 75 80Arg Cys Val Leu Ala Ala
Val Arg Ser Ala Thr Ile Gly Met Pro Ile 85 90 95Glu Pro Ser Ala Ser
Ala Pro Phe Ser Asp Gly Gln Trp Leu Leu Ser 100 105 110His Asn Gly
Leu Val Asp Arg Gly Val Leu Pro Leu Thr Gly Ala Ala 115 120 125Glu
Ser Thr Val Asp Ser Ala Ile Val Ala Ala Leu Ile Phe Ser Arg 130 135
140Gly Leu Asp Ala Leu Gly Ala Thr Ile Ala Glu Val Gly Glu Leu
Asp145 150 155 160Pro Asn Ala Arg Leu Asn Ile Leu Ala Ala Asn Gly
Ser Arg Leu Leu 165 170 175Ala Thr Thr Trp Gly Asp Thr Leu Ser Val
Leu His Arg Pro Asp Gly 180 185 190Val Val Leu Ala Ser Glu Pro Tyr
Asp Asp Asp Pro Gly Trp Ser Asp 195 200 205Ile Pro Asp Arg His Leu
Val Asp Val Arg Asp Ala His Val Val Val 210 215 220Thr Pro
Leu2255966DNAArtificial Sequencesynthetic 5atgacgctct cactggccaa
ctacctggca gccgactcgg ccgccgaagc actgcgccgt 60gacgtccgcg cgggcctcac
cgcggcaccg aagagtctgc cgcccaagtg gttctacgac 120gccgtcggca
gtgatctgtt cgaccagatc acccggctcc ccgagtatta ccccacccgc
180accgaggcgc agatcctgcg gacccggtcg gcggagatca tcgcggccgc
gggtgccgac 240accctggtgg aactgggcag tggtacgtcg gagaaaaccc
gcatgctgct cgacgccatg 300cgcgacgccg agttgctgcg ccgcttcatc
ccgttcgacg tcgacgcggg cgtgctgcgc 360tcggccgggg cggcaatcgg
cgcggagtac cccggtatcg agatcgacgc ggtatgtggc 420gatttcgagg
aacatctggg caagatcccg catgtcggac ggcggctcgt ggtgttcctg
480gggtcgacca tcggcaacct gacacccgcg ccccgcgcgg agttcctcag
tactctcgcg 540gacacgctgc agccgggcga cagcctgctg ctgggcaccg
atctggtgaa ggacaccggc 600cggttggtgc gcgcgtacga cgacgcggcc
ggcgtcaccg cggcgttcaa ccgcaacgtg 660ctggccgtgg tgaaccgcga
actgtccgcc gatttcgacc tcgacgcgtt cgagcatgtc 720gcgaagtgga
actccgacga ggaacgcatc gagatgtggt tgcgtgcccg caccgcacag
780catgtccgcg tcgcggcact ggacctggag gtcgacttcg ccgcgggtga
ggagatgctc 840accgaggtgt cctgcaagtt ccgtcccgag aacgtcgtcg
ccgagctggc ggaagccggt 900ctgcggcaga cgcattggtg gaccgatccg
gccggggatt tcgggttgtc gctggcggtg 960cggtga 9666321PRTArtificial
Sequencesynthetic 6Met Thr Leu Ser Leu Ala Asn Tyr Leu Ala Ala Asp
Ser Ala Ala Glu1 5 10 15Ala Leu Arg Arg Asp Val Arg Ala Gly Leu Thr
Ala Ala Pro Lys Ser 20 25 30Leu Pro Pro Lys Trp Phe Tyr Asp Ala Val
Gly Ser Asp Leu Phe Asp 35 40 45Gln Ile Thr Arg Leu Pro Glu Tyr Tyr
Pro Thr Arg Thr Glu Ala Gln 50 55 60Ile Leu Arg Thr Arg Ser Ala Glu
Ile Ile Ala Ala Ala Gly Ala Asp65 70 75 80Thr Leu Val Glu Leu Gly
Ser Gly Thr Ser Glu Lys Thr Arg Met Leu 85 90 95Leu Asp Ala Met Arg
Asp Ala Glu Leu Leu Arg Arg Phe Ile Pro Phe 100 105 110Asp Val Asp
Ala Gly Val Leu Arg Ser Ala Gly Ala Ala Ile Gly Ala 115 120 125Glu
Tyr Pro Gly Ile Glu Ile Asp Ala Val Cys Gly Asp Phe Glu Glu 130 135
140His Leu Gly Lys Ile Pro His Val Gly Arg Arg Leu Val Val Phe
Leu145 150 155 160Gly Ser Thr Ile Gly Asn Leu Thr Pro Ala Pro Arg
Ala Glu Phe Leu 165 170 175Ser Thr Leu Ala Asp Thr Leu Gln Pro Gly
Asp Ser Leu Leu Leu Gly 180 185 190Thr Asp Leu Val Lys Asp Thr Gly
Arg Leu Val Arg Ala Tyr Asp Asp 195 200 205Ala Ala Gly Val Thr Ala
Ala Phe Asn Arg Asn Val Leu Ala Val Val 210 215 220Asn Arg Glu Leu
Ser Ala Asp Phe Asp Leu Asp Ala Phe Glu His Val225 230 235 240Ala
Lys Trp Asn Ser Asp Glu Glu Arg Ile Glu Met Trp Leu Arg Ala 245 250
255Arg Thr Ala Gln His Val Arg Val Ala Ala Leu Asp Leu Glu Val Asp
260 265 270Phe Ala Ala Gly Glu Glu Met Leu Thr Glu Val Ser Cys Lys
Phe Arg 275 280 285Pro Glu Asn Val Val Ala Glu Leu Ala Glu Ala Gly
Leu Arg Gln Thr 290 295 300His Trp Trp Thr Asp Pro Ala Gly Asp Phe
Gly Leu Ser Leu Ala Val305 310 315 320Arg71113DNAArtificial
Sequencesynthetic 7atgctcgcgc agcagtggcg tgacgcccgt cccaaggttg
ccgggttgca cctggacagc 60ggggcatgtt cgcggcagag cttcgcggtg atcgacgcga
ccaccgcaca cgcacgccac 120gaggccgagg tgggtggtta tgtggcggcc
gaggctgcga cgccggcgct cgacgccggg 180cgggccgcgg tcgcgtcgct
catcggtttt gcggcgtcgg acgtggtgta caccagcgga 240tccaaccacg
ccatcgacct gttgctgtcg agctggccgg ggaagcgcac gctggcctgc
300ctgcccggcg agtacgggcc gaatctgtct gccatggcgg ccaacggttt
ccaggtgcgt 360gcgctaccgg tcgacgacga cgggcgggtg ctggtcgacg
aggcgtcgca cgaactgtcg 420gcccatcccg tcgcgctcgt acacctcacc
gcattggcaa gccatcgcgg gatcgcgcaa 480cccgcggcag aactcgtcga
ggcctgccac aatgcgggga tccccgtggt gatcgacgcc 540gcgcaggcgc
tggggcatct ggactgcaat gtcggggccg acgcggtgta ctcatcgtcg
600cgcaagtggc tcgccggccc gcgtggtgtc ggggtgctcg cggtgcggcc
cgaactcgcc 660gagcgtctgc aaccgcggat ccccccgtcc gactggccaa
ttccgatgag cgtcttggag 720aagctcgaac taggtgagca caacgcggcg
gcgcgtgtgg gattctccgt cgcggttggt 780gagcatctcg cagcagggcc
cacggcggtg cgcgaacgac tcgccgaggt ggggcgtctc 840tctcggcagg
tgctggcaga ggtcgacggg tggcgcgtcg tcgaacccgt cgaccaaccc
900accgcgatca ccacccttga gtccaccgat ggtgccgatc ccgcgtcggt
gcgctcgtgg 960ctgatcgcgg agcgtggcat cgtgaccacc gcgtgtgaac
tcgcgcgggc accgttcgag 1020atgcgcacgc cggtgctgcg aatctcgccg
cacgtcgacg tgacggtcga cgaactggag 1080cagttcgccg cagcgttgcg
tgaggcgccc tga 11138370PRTArtificial Sequencesynthetic 8Met Leu Ala
Gln Gln Trp Arg Asp Ala Arg Pro Lys Val Ala Gly Leu1 5 10 15His Leu
Asp Ser Gly Ala Cys Ser Arg Gln Ser Phe Ala Val Ile Asp 20 25 30Ala
Thr Thr Ala His Ala Arg His Glu Ala Glu Val Gly Gly Tyr Val 35 40
45Ala Ala Glu Ala Ala Thr Pro Ala Leu Asp Ala Gly Arg Ala Ala Val
50 55 60Ala Ser Leu Ile Gly Phe Ala Ala Ser Asp Val Val Tyr Thr Ser
Gly65 70 75 80Ser Asn His Ala Ile Asp Leu Leu Leu Ser Ser Trp Pro
Gly Lys Arg 85 90 95Thr Leu Ala Cys Leu Pro Gly Glu Tyr Gly Pro Asn
Leu Ser Ala Met 100 105 110Ala Ala Asn Gly Phe Gln Val Arg Ala Leu
Pro Val Asp Asp Asp Gly 115 120 125Arg Val Leu Val Asp Glu Ala Ser
His Glu Leu Ser Ala His Pro Val 130 135 140Ala Leu Val His Leu Thr
Ala Leu Ala Ser His Arg Gly Ile Ala Gln145 150 155 160Pro Ala Ala
Glu Leu Val Glu Ala Cys His Asn Ala Gly Ile Pro Val 165 170 175Val
Ile Asp Ala Ala Gln Ala Leu Gly His Leu Asp Cys Asn Val Gly 180 185
190Ala Asp Ala Val Tyr Ser Ser Ser Arg Lys Trp Leu Ala Gly Pro Arg
195 200 205Gly Val Gly Val Leu Ala Val Arg Pro Glu Leu Ala Glu Arg
Leu Gln 210 215 220Pro Arg Ile Pro Pro Ser Asp Trp Pro Ile Pro Met
Ser Val Leu Glu225 230 235 240Lys Leu Glu Leu Gly Glu His Asn Ala
Ala Ala Arg Val Gly Phe Ser 245 250 255Val Ala Val Gly Glu His Leu
Ala Ala Gly Pro Thr Ala Val Arg Glu 260 265 270Arg Leu Ala Glu Val
Gly Arg Leu Ser Arg Gln Val Leu Ala Glu Val 275 280 285Asp Gly Trp
Arg Val Val Glu Pro Val Asp Gln Pro Thr Ala Ile Thr 290 295 300Thr
Leu Glu Ser Thr Asp Gly Ala Asp Pro Ala Ser Val Arg Ser Trp305 310
315 320Leu Ile Ala Glu Arg Gly Ile Val Thr Thr Ala Cys Glu Leu Ala
Arg 325 330 335Ala Pro Phe Glu Met Arg Thr Pro Val Leu Arg Ile Ser
Pro His Val 340 345 350Asp Val Thr Val Asp Glu Leu Glu Gln Phe Ala
Ala Ala Leu Arg Glu 355 360 365Ala Pro 370949DNAArtificial
Sequencesynthetic 9agaattcaaa agatctaaag gaggccatcc atgatcgcac
gcgagacac 491053DNAArtificial Sequencesynthetic 10actcgagttt
ggatcctcag acgtcccagg ccaggcggac acccgagaat atc 531150DNAArtificial
Sequencesynthetic 11agaattcaaa agatctaaag gaggccatcc atgtgccggc
atgtggcgtg 501234DNAArtificial Sequencesynthetic 12actcgagttt
ggatcctcac aggggtgtca cgac 341351DNAArtificial Sequencesynthetic
13agaattcaaa agatctaaag gaggccatcc atgacgctct cactggccaa c
511435DNAArtificial Sequencesynthetic 14actcgagttt ggatcctcac
cgcaccgcca gcgac 351547DNAArtificial Sequencesynthetic 15agaattcaaa
agatctaaag gaggccatcc atgctcgcgc agcagtg 471635DNAArtificial
Sequencesynthetic 16actcgagttt ggatcctcag ggcgcctcac gcaac 35
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.