Broadly Neutralizing Antibodies Against Hiv
Sajadi; Mohammad ; et al.
U.S. patent application number 16/626163 was filed with the patent office on 2020-06-04 for broadly neutralizing antibodies against hiv. The applicant listed for this patent is Dongkyoon Cavet Kim. Invention is credited to Guy Cavet, Amir Dashti, Anthony DeVico, Dongkyoon Kim, George K. Lewis, Marzena E. Pazgier, Mohammad Sajadi, William David Tolbert.
| Application Number | 20200172601 16/626163 |
| Document ID | / |
| Family ID | 64736132 |
| Filed Date | 2020-06-04 |












View All Diagrams
| United States Patent Application | 20200172601 |
| Kind Code | A1 |
| Sajadi; Mohammad ; et al. | June 4, 2020 |
BROADLY NEUTRALIZING ANTIBODIES AGAINST HIV
Abstract
The present invention provides broadly neutralizing antibodies against HIV, compositions comprising the same and methods of use thereof. Specifically, the invention provides isolated anti-HIV antibodies that are capable of neutralizing at least 95% of the HIV pseudoviruses listed in Table 1 and neutralizing 100% of the HIV clade B, G and D viruses listed in Table 1 with an IC50 value of less than 50 ug/mL. Further provides are the sequences especially complementarity-determining region (CDR) sequences of antibodies disclosed.
| Inventors: | Sajadi; Mohammad; (Cockeysville, MD) ; Lewis; George K.; (Baltimore, MD) ; DeVico; Anthony; (Alexandria, VA) ; Dashti; Amir; (Baltimore, MD) ; Pazgier; Marzena E.; (Mt. Airy, MD) ; Tolbert; William David; (Baltimore, MD) ; Kim; Dongkyoon; (Palo Alto, CA) ; Cavet; Guy; (Hillsborough, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 64736132 | ||||||||||
| Appl. No.: | 16/626163 | ||||||||||
| Filed: | June 22, 2018 | ||||||||||
| PCT Filed: | June 22, 2018 | ||||||||||
| PCT NO: | PCT/US2018/039162 | ||||||||||
| 371 Date: | December 23, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 62673607 | May 18, 2018 | |||
| 62591244 | Nov 28, 2017 | |||
| 62589614 | Nov 22, 2017 | |||
| 62573764 | Oct 18, 2017 | |||
| 62523437 | Jun 22, 2017 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 15/00 20130101; C07K 14/005 20130101; C07K 2317/565 20130101; A61P 31/18 20180101; A61K 39/00 20130101; C12N 2740/16122 20130101; C07K 2317/21 20130101; C07K 2317/33 20130101; C07K 2317/76 20130101; C07K 2317/92 20130101; C07K 16/1063 20130101; C12N 2740/16134 20130101; A61K 39/42 20130101; C07K 2317/55 20130101; C07K 2317/34 20130101 |
| International Class: | C07K 16/10 20060101 C07K016/10; A61K 39/42 20060101 A61K039/42; A61P 31/18 20060101 A61P031/18 |
Goverment Interests
STATEMENT OF FEDERALLY SPONSORED RESEARCH AND DEVELOPMENT
[0002] This invention was made with government support under Grant Number AI110259 awarded by the National Institutes of Health and under Grant Number 1I01BX002358 awarded by the United States Department of Veterans Affairs. The government has certain rights in the invention.
Claims
1. An isolated anti-HIV antibody that is capable of neutralizing at least 95% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL.
2. (canceled)
3. (canceled)
4. An isolated anti-HIV antibody that neutralizes 100% of the HIV clade B, G and D viruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL.
5. The isolated anti-HIV antibody of claim 1, wherein the anti-HIV antibody binds to a HIV gp120 epitope comprising outer domain loop D (which comprises 275-283), the CD4 binding loop (which comprises 354-371), the bridging sheet (which comprises 427-439) and loop V5 (which comprises 455-463) and gp120 inner domain: helix alpha-1 of Layer 2 (comprising positions 96-106) and 469-480 (loop prior and helix alpha-5 of Layer 3).
6. The isolated anti-HIV antibody of claim 5, wherein the anti-HIV antibody binds to a HIV gp120 Layer 2 residues W96, K97, E102, G124, Loop D residues E275, N276, T278, N279, N280, A281, K282, CD4 binding loop residues P364, 5365, G366, G367, D368, 1371, bridging sheet residues W427, Q428, G429, Loop V5 residues T455, R456, D457, G458, G459, A460, N461, T463, and Layer 3 residues R469, P470, G471, G472, G473, N474, K476, D477, R480.
7. The isolated anti-HIV antibody of claim 6, wherein the antibody has a Kd for BaL-gp120 of at least about 2.456.times.10.sup.-8 M as determined by surface plasmon resonance.
8.-11. (canceled)
12. The isolated anti-HIV antibody of claim 4, wherein the anti-HIV antibody further neutralizes strain CNE5 (clade CRF01_AE) with an IC50 value of less than 50 .mu.g/mL.
13.-19. (canceled)
20. The isolated anti-HIV antibody of claim 1, wherein anti-HIV antibody is selected from the group consisting of: a. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYDFIDYV (SEQ ID NO:401), CDR H2 comprises MNPSGGGT (SEQ ID NO:402) and CDR H3 comprises VRDRANGSGRRRFESVNWFLDL (SEQ ID NO:403); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); b. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); c. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); and d. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYL (SEQ ID NO:409), CDR H2 comprises MNPVYGQV (SEQ ID NO:410) and CDR H3 comprises VRDTGDGSRRHFDSINWFLDL (SEQ ID NO:411); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412).
21. The isolated anti-HIV antibody of claim 1, wherein the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain or antigen binding fragment thereof comprises a heavy chain variable (VH) region and the light chain or antigen binding fragment thereof comprises a light chain variable (VL) region; wherein the anti-HIV antibody is selected from the group consisting of: a. an antibody wherein the VH region comprises amino acids 1-128 of SEQ ID NO:153 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:155 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; b. an antibody wherein the VH region comprises amino acids 1-127 of SEQ ID NO:161 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:163 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; c. an antibody wherein the VH region comprises amino acids 1-127 of SEQ ID NO:165 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:167 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; d. an antibody wherein the VH region comprises amino acids 1-128 of SEQ ID NO:225 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:227 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; and e. an antibody wherein the VH region comprises amino acids 1-120 of SEQ ID NO:293 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:295 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions.
22. The anti-HIV antibody of claim 1, wherein the anti-HIV antibody is selected from the group consisting of: a. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:153 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:155 or an antigen binding fragment thereof; b. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:161 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:163 or an antigen binding fragment thereof; c. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:165 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:167 or an antigen binding fragment thereof; d. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:225 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:227 or an antigen binding fragment thereof; and e. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:293 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:295 or an antigen binding fragment thereof.
23. An isolated host cell expressing the antibody of claim 21.
24. One or more vectors comprising a nucleic acid encoding the antibody of claim 21.
25.-27. (canceled)
28. An engineered cell that expresses the antibody of claim 21.
29. (canceled)
30. (canceled)
31. A pharmaceutical composition comprising one or more antibodies of claim 5 and/or cells of claim 23 and a pharmaceutically acceptable carrier.
32. A method for treating or preventing HIV infection in a subject, comprising administering to the subject an effective amount of the composition of claim 31.
33. A method of functionally curing HIV in a subject comprising administering to the subject an effective amount of the composition of claim 31.
34. The method of claim 31, wherein the composition is administered in combination with another therapy.
35. The method of claim 34, wherein the therapy is an anti-retroviral therapy.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application claims the benefit of U.S. Provisional Appl. No. 62/523,437, filed Jun. 22, 2017, U.S. Provisional Appl. No. 62/573,764, filed Oct. 18, 2017, U.S. Provisional Appl. No. 62/589,614, filed Nov. 22, 2017, U.S. Provisional Appl. No. 62/591,244, filed Nov. 28, 2017, and U.S. Provisional Appl. No. 62/673,607, filed May 18, 2018, the contents of which are hereby incorporated by reference in their entirety.
INCORPORATION-BY-REFERENCE OF MATERIAL SUBMITTED ELECTRONICALLY
[0003] Incorporated by reference in its entirety herein is a computer-readable sequence listing submitted concurrently herewith and identified as follows: One 884,569 Byte ASCII (Text) file named "Sequence_Listing_ST25.txt," created on Jun. 22, 2018.
FIELD OF THE INVENTION
[0004] The field of the invention relates to medicine, infectious disease and in particular antibodies which can neutralize HIV-1 virus strains.
BACKGROUND OF THE INVENTION
[0005] HIV is an integrating retrovirus that rapidly establishes chronic infection in CD4+ T cells. This fundamental characteristic means that prevention of HIV infection largely depends on humoral responses and associated effector mechanisms directed against the HIV envelope proteins (gp120 and gp41) that drive viral attachment and entry.
[0006] Humoral anti-envelope responses in a minority of HIV-infected persons comprise neutralizing activity against diverse viral variants (Scheid et al., Nature 458, 636-640 (2009); Simek et al., J Virol 83, 7337-7348 (2009); Walker et al., PLoS Pathog 6, e1001028 (2010); Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Infect Dis 213, 156-164 (2016)). Broadly neutralizing responses can be used to guide the development of effective HIV vaccines and/or other immune-based prevention measures. Three types of information are essential to implementing this concept. First, conserved sites of extreme neutralization sensitivity within the HIV envelope structure must be defined. Significant steps in this direction have been afforded by the derivation of broadly neutralizing monoclonal anti-envelope antibodies (mAbs) from the memory B cell pools of certain HIV-infected individuals. These antibodies reveal a number of especially potent neutralizing epitopes on gp120, including the CD4 binding site (CD4-BS), V1V2 glycan, V3 glycan, and the gp41 membrane-proximal external region (Haynes et al., J Allergy Clin Immunol 134, 3-10; quiz 11 (2014). Second, the features of broadly neutralizing antibodies that arise in multiple individuals, versus rare subjects, must be fully characterized. A number of serological studies have made progress in this regard, particularly with respect to epitopes on gp120 (Scheid et al., Nature 458, 636-640 (2009); Simek et al., J Virol 83, 7337-7348 (2009); Walker et al., PLoS Pathog 6, e1001028 (2010); Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Virol 86, 5014-5025 (2012)). Third, the aggregate nature of the polyclonal humoral environment in which broadly neutralizing activities evolve, persist and function must be understood. Collectively, this information can be used to delineate whether and how certain epitope presentation patterns should be avoided or targeted in order to deliberately achieve potent and broad neutralizing activity.
[0007] To date, the interrelationships between broadly neutralizing antibodies and the circulating plasma anti-HIV envelope humoral repertoires that harbor them have been examined mainly by indirect means. Typical approaches involve protein fractionation, antigen depletion and/or infectivity analyses using viral envelopes with targeted mutations (Sather et al., Vaccine 28 Suppl 2, B8-12 (2010); Li et al., J Virol 83, 1045-1059 (2009); Dhillon et al., J Virol 81, 6548-6562 (2007)). These methods do not fully elucidate the background milieu of the polyclonal anti-envelope humoral response and cannot clearly define the neutralizing antibody species in circulation. For example, various studies indicate that broad plasma neutralizing activity may be traced to either pauciclonal or polyclonal antibody species (Scheid et al., Nature 458, 636-640 (2009); Walker et al., PLoS Pathog 6, e1001028 (2010); Sajadi et al., J Virol 86, 5014-5025 (2012); Bonsignori et al., J Virol 86, 4688-4692 (2012); Doria-Rose et al., J Virol 84, 1631-1636 (2010)). depending on the source subject. Alternatively, intensive efforts have been applied toward the derivation of neutralizing mAbs from memory B cell pools. These antibodies, albeit important for other purposes, may not reflect the true nature of neutralizing antibodies in circulation (Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009); Scheid et al., Nature 458, 636-640 (2009); Walker et al., Science 326, 285-289 (2009); Walker et al., Nature 477, 466-470 (2011).
[0008] There is a need to develop new therapies for treatment and prevention of HIV infection in patients.
[0009] This background information is provided for informational purposes only. No admission is necessarily intended, nor should it be construed, that any of the preceding information constitutes prior art against the present invention.
SUMMARY OF THE INVENTION
[0010] It is to be understood that both the foregoing general description of the embodiments and the following detailed description are exemplary, and thus do not restrict the scope of the embodiments.
[0011] In one aspect, the invention provides an isolated anti-HIV antibody, wherein the antibody is capable of neutralizing at least 95% of the HIV viruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the isolated anti-HIV antibody is capable of neutralizing at least 99% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the antibody is capable of neutralizing 100% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the antibody is selected from the group consisting of [0012] a. N49P6 or an antigen binding fragment thereof; [0013] b. N49P7 or an antigen binding fragment thereof; [0014] c. N49P7.1 or an antigen binding fragment thereof; and [0015] d. N49P11 or an antigen binding fragment thereof.
[0016] In another aspect, the invention provides an isolated anti-HIV antibody, wherein the antibody is capable of neutralizing 100% of the HIV clade B, G and D pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the antibody is selected from the group consisting of [0017] a. N49P6 or an antigen binding fragment thereof; [0018] b. N49P7 or an antigen binding fragment thereof; [0019] c. N49P7.1 or an antigen binding fragment thereof; [0020] d. N49P11 or an antigen binding fragment thereof; and [0021] e. N49P9 or an antigen binding fragment thereof.
[0022] In another aspect, the invention provides an isolated anti-HIV antibody, wherein the antibody is capable of neutralizing HIV pseudoviruses listed in Table 1 with a median IC50 value of less than 0.5 .mu.g/mL. In some embodiments, the antibody is selected from the group consisting of [0023] a. N49P6 or an antigen binding fragment thereof; [0024] b. N49P7 or an antigen binding fragment thereof; [0025] c. N49P7.1 or an antigen binding fragment thereof [0026] d. N49P9 or an antigen binding fragment thereof; and [0027] e. N49P23 or an antigen binding fragment thereof.
[0028] In another aspect, the invention provides an isolated anti-HIV antibody selected from the group consisting of: [0029] a. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:1 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:2 or an antigen binding fragment thereof; [0030] b. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:3 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:4 or an antigen binding fragment thereof; [0031] c. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:5 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:6 or an antigen binding fragment thereof [0032] d. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:7 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:8 or an antigen binding fragment thereof [0033] e. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:9 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:10 or an antigen binding fragment thereof [0034] f. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:11 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:12 or an antigen binding fragment thereof [0035] g. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:13 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:14 or an antigen binding fragment thereof [0036] h. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:15 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:16 or an antigen binding fragment thereof [0037] i. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:17 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:18 or an antigen binding fragment thereof; [0038] j. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:19 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:20 or an antigen binding fragment thereof; [0039] k. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:21 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:22 or an antigen binding fragment thereof; [0040] l. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:23 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:24 or an antigen binding fragment thereof; [0041] m. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:25 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:26 or an antigen binding fragment thereof; [0042] n. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:27 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:28 or an antigen binding fragment thereof; [0043] o. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:29 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:30 or an antigen binding fragment thereof; [0044] p. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:31 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:32 or an antigen binding fragment thereof; [0045] q. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:33 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:34 or an antigen binding fragment thereof; [0046] r. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:35 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:36 or an antigen binding fragment thereof; [0047] s. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:37 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:38 or an antigen binding fragment thereof; [0048] t. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:39 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:40 or an antigen binding fragment thereof; [0049] u. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:41 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:42 or an antigen binding fragment thereof; [0050] v. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:43 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:44 or an antigen binding fragment thereof; [0051] w. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:45 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:46 or an antigen binding fragment thereof; [0052] x. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:47 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:48 or an antigen binding fragment thereof; [0053] y. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:49 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:50 or an antigen binding fragment thereof; [0054] z. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:51 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:52 or an antigen binding fragment thereof; [0055] aa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:53 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:54 or an antigen binding fragment thereof; [0056] bb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:55 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:56 or an antigen binding fragment thereof; [0057] cc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:57 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:58 or an antigen binding fragment thereof; [0058] dd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:59 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:60 or an antigen binding fragment thereof; [0059] ee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:61 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:62 or an antigen binding fragment thereof; [0060] ff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:63 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:64 or an antigen binding fragment thereof; [0061] gg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:65 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:66 or an antigen binding fragment thereof; [0062] hh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:67 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:68 or an antigen binding fragment thereof; [0063] ii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:69 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:70 or an antigen binding fragment thereof; [0064] jj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:71 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:72 or an antigen binding fragment thereof; [0065] kk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:73 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:74 or an antigen binding fragment thereof; and [0066] ll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:75 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:76 or an antigen binding fragment thereof.
[0067] In another aspect, the invention provides an isolated anti-HIV antibody selected from the group consisting of: [0068] a. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:153 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:155 or an antigen binding fragment thereof; [0069] b. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:157 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:159 or an antigen binding fragment thereof; [0070] c. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:161 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:163 or an antigen binding fragment thereof; [0071] d. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:165 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:167 or an antigen binding fragment thereof; [0072] e. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:169 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:171 or an antigen binding fragment thereof; [0073] f. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:173 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:175 or an antigen binding fragment thereof; [0074] g. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:177 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:179 or an antigen binding fragment thereof; [0075] h. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:181 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:183 or an antigen binding fragment thereof; [0076] i. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:185 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:187 or an antigen binding fragment thereof; [0077] j. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:189 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:191 or an antigen binding fragment thereof; [0078] k. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:193 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:195 or an antigen binding fragment thereof; [0079] l. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:197 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:199 or an antigen binding fragment thereof; [0080] m. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:201 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:203 or an antigen binding fragment thereof; [0081] n. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:205 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:207 or an antigen binding fragment thereof; [0082] o. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:209 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:211 or an antigen binding fragment thereof; [0083] p. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:213 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:215 or an antigen binding fragment thereof; [0084] q. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:217 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:219 or an antigen binding fragment thereof; [0085] r. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:221 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:223 or an antigen binding fragment thereof; [0086] s. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:225 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:227 or an antigen binding fragment thereof; [0087] t. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:229 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:231 or an antigen binding fragment thereof; [0088] u. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:233 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:235 or an antigen binding fragment thereof; [0089] v. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:237 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:239 or an antigen binding fragment thereof; [0090] w. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:241 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:243 or an antigen binding fragment thereof; [0091] x. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:245 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:247 or an antigen binding fragment thereof; [0092] y. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:249 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:251 or an antigen binding fragment thereof; [0093] z. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:253 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:255 or an antigen binding fragment thereof; [0094] aa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:257 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:259 or an antigen binding fragment thereof; [0095] bb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:261 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:263 or an antigen binding fragment thereof; [0096] cc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:265 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:267 or an antigen binding fragment thereof; [0097] dd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:269 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:271 or an antigen binding fragment thereof; [0098] ee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:273 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:275 or an antigen binding fragment thereof; [0099] ff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:277 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:279 or an antigen binding fragment thereof; [0100] gg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:281 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:283 or an antigen binding fragment thereof; [0101] hh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:285 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:287 or an antigen binding fragment thereof; [0102] ii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:289 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:291 or an antigen binding fragment thereof; [0103] jj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:293 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:295 or an antigen binding fragment thereof; [0104] kk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:297 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:299 or an antigen binding fragment thereof; [0105] ll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:301 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:303 or an antigen binding fragment thereof; [0106] mm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:305 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:307 or an antigen binding fragment thereof; [0107] nn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:309 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:311 or an antigen binding fragment thereof; [0108] oo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:313 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:315 or an antigen binding fragment thereof; [0109] pp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:317 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:319 or an antigen binding fragment thereof; [0110] qq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:321 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:323 or an antigen binding fragment thereof; [0111] rr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:325 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:327 or an antigen binding fragment thereof; [0112] ss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:329 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:331 or an antigen binding fragment thereof; [0113] tt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:333 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:335 or an antigen binding fragment thereof; [0114] uu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:337 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:339 or an antigen binding fragment thereof; [0115] vv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:341 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:343 or an antigen binding fragment thereof; [0116] ww. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:345 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:347 or an antigen binding fragment thereof; [0117] xx. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:349 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:351 or an antigen binding fragment thereof; [0118] yy. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:353 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:355 or an antigen binding fragment thereof; [0119] zz. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:357 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:359 or an antigen binding fragment thereof; [0120] aaa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:361 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:363 or an antigen binding fragment thereof; [0121] bbb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:365 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:367 or an antigen binding fragment thereof; [0122] ccc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:369 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:371 or an antigen binding fragment thereof; [0123] ddd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:373 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:375 or an antigen binding fragment thereof; [0124] eee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:377 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:379 or an antigen binding fragment thereof; [0125] fff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:381 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:383 or an antigen binding fragment thereof; [0126] ggg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:385 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:387 or an antigen binding fragment thereof; [0127] hhh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:389 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:391 or an antigen binding fragment thereof; [0128] iii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:393 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:395 or an antigen binding fragment thereof; and [0129] jjj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:397 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:399 or an antigen binding fragment thereof.
[0130] In another aspect, the invention provides an anti-HIV antibody selected from the group consisting of: [0131] a. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYDFIDYV (SEQ ID NO:401), CDR H2 comprises MNPSGGGT (SEQ ID NO:402) and CDR H3 comprises VRDRANGSGRRRFESVNWFLDL (SEQ ID NO:403); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); [0132] b. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFPDYI (SEQ ID NO:405), CDR H2 comprises INPMGGQV (SEQ ID NO:406) and CDR H3 comprises VRDRSNGSGRRFESSN (SEQ ID NO:407); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0133] c. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYL (SEQ ID NO:409), CDR H2 comprises MNPVYGQV (SEQ ID NO:410) and CDR H3 comprises VRDTGDGSRRHFDSINWFLDL (SEQ ID NO:411); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412); [0134] d. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises IDPPYGQV (SEQ ID NO:414) and CDR H3 comprises VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0135] e. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFVDYF (SEQ ID NO:416), CDR H2 comprises MDPLNGRP (SEQ ID NO:417) and CDR H3 comprises VRDKSNGSGRRFDSSNWFLDL (SEQ ID NO:418); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0136] f. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFSDYI (SEQ ID NO:420), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEV (SEQ ID NO:422); [0137] g. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFIDYI (SEQ ID NO:423), CDR H2 comprises IDPMNGRP (SEQ ID NO:424) and CDR H3 comprises VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0138] h. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYI (SEQ ID NO:426), CDR H2 comprises MNPMGGRT (SEQ ID NO:427) and CDR H3 comprises VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0139] i. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFVDYL (SEQ ID NO:428), CDR H2 comprises MDPMNGRP (SEQ ID NO:429) and CDR H3 comprises VRDKSGGSGKLFDSSNWFLDL (SEQ ID NO:430); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0140] j. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises INPGYGQV (SEQ ID NO:431) and CDR H3 comprises VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0141] k. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises MDPSYGQV (SEQ ID NO:432) and CDR H3 comprises VRDRSHGSGRQFESSNWFLDL (SEQ ID NO:433); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0142] l. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises MDPSFGQM (SEQ ID NO:434) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0143] m. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFTDYV (SEQ ID NO:436), CDR H2 comprises MDPSFGRM (SEQ ID NO:437) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0144] n. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFIDYV (SEQ ID NO:438), CDR H2 comprises MDPTYGRM (SEQ ID NO:439) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0145] o. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFLDYI (SEQ ID NO:440), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0146] p. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0147] q. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSR (SEQ ID NO:447), CDR H2 comprises INPLQGGV (SEQ ID NO:448) and CDR H3 comprises ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0148] r. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTTHHGHF (SEQ ID NO:500), CDR H2 comprises MNPMTGQM (SEQ ID NO:462) and CDR H3 comprises ARGDFGQNYPFHY (SEQ ID NO:463); and a light chain variable region, wherein CDR L1 comprises NRYL (SEQ ID NO:464), CDR L2 comprises DDN and CDR L3 comprises ASYER (SEQ ID NO:465); [0149] s. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFMDQF (SEQ ID NO:466), CDR H2 comprises MNPIYGQV (SEQ ID NO:467) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0150] t. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSR (SEQ ID NO:447), CDR H2 comprises INPLHGGV (SEQ ID NO:468) and CDR H3 comprises ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0151] u. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTKYF (SEQ ID NO:451), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADSYGSSYPFHH (SEQ ID NO:453); and a light chain variable region, wherein CDR L1 comprises GNYNP (SEQ ID NO:454), CDR L2 comprises EDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0152] v. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTKYT (SEQ ID NO:456), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADLSGPTYPFHH (SEQ ID NO:457); and a light chain variable region, wherein CDR L1 comprises GNYNP (SEQ ID NO:454), CDR L2 comprises EDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0153] w. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFNFIDSV (SEQ ID NO:458), CDR H2 comprises IKPLRGAV (SEQ ID NO:459) and CDR H3 comprises AKGAFRGGSPFGF (SEQ ID NO:460); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0154] x. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTSYF (SEQ ID NO:469), CDR H2 comprises INPLHGAV (SEQ ID NO:470) and CDR H3 comprises TRGIVADGWPYGH (SEQ ID NO:471); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises EGA and CDR L3 comprises SSLQF (SEQ ID NO:472); [0155] y. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFTFIDHI (SEQ ID NO:473), CDR H2 comprises IKPLRGAV (SEQ ID NO:459) and CDR H3 comprises CKAAAPEEAFPLQY (SEQ ID NO:474); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSRTF (SEQ ID NO:475); [0156] z. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDHI (SEQ ID NO:476), CDR H2 comprises IKPLGGVA (SEQ ID NO:477) and CDR H3 comprises CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0157] aa. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFAFLDH (SEQ ID NO:480), CDR H2 comprises VKTIGGVV (SEQ ID NO:481) and CDR H3 comprises SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0158] bb. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFTEYF (SEQ ID NO:483), CDR H2 comprises LNPLRGAV (SEQ ID NO:484) and CDR H3 comprises ARAVFNEAFPFDY (SEQ ID NO:485); and a light chain variable region, wherein CDR L1 comprises VS, CDR L2 comprises DGD and CDR L3 comprises ASREF (SEQ ID NO:461); [0159] cc. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDHI (SEQ ID NO:476), CDR H2 comprises IKPLGGVA (SEQ ID NO:477) and CDR H3 comprises CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DND and CDR L3 comprises SSTTF (SEQ ID NO:479); [0160] dd. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFAFLDHI (SEQ ID NO:486), CDR H2 comprises VKTIGGVV (SEQ ID NO:481) and CDR H3 comprises SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0161] ee. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDSV (SEQ ID NO:487), CDR H2 comprises IKPLGGAV (SEQ ID NO:488) and CDR H3 comprises AKGAFGGGSPFGF (SEQ ID NO:489); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0162] ff. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFNFIDSV (SEQ ID NO:458), CDR H2 comprises IKPLRGGV (SEQ ID NO:490) and CDR H3 comprises AKGAFGGSSPFGF (SEQ ID NO:491); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0163] gg. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFTFIKYT (SEQ ID NO:492), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADLYGPTYPFHH (SEQ ID NO:493); and a light chain variable region, wherein CDR L1 comprises GSYNP (SEQ ID NO:494), CDR L2 comprises DDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0164] hh. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSL (SEQ ID NO:495), CDR H2 comprises INPLQGGV (SEQ ID NO:448) and CDR H3 comprises ARGIDGNSYPFHF (SEQ ID NO:496); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0165] ii. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0166] jj. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); [0167] kk. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412); [0168] ll. an antibody comprising a heavy chain variable region, wherein CDR comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0169] mm. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0170] nn. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0171] oo. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSN (SEQ ID NO:498); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0172] pp. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); and [0173] qq. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFLDYI (SEQ ID NO:440), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408).
[0174] Other objects, features and advantages of the present invention will become apparent from the following detailed description. It should be understood, however, that the detailed description and the specific examples, while indicating specific embodiments of the invention, are given by way of illustration only, since various changes and modifications within the spirit and scope of the invention will become apparent to those skilled in the art from this detailed description.
BRIEF DESCRIPTION OF THE FIGURES
[0175] The skilled artisan will understand that the drawings, described below, are for illustration purposes only. The drawings are not intended to limit the scope of the present teachings in any way.
[0176] FIG. 1. ELISA reactivity patterns of fractionated IgG from NVS60 with broad HIV neutralizing activity. Anti-gp120 IgG1 kappa (panel A) and lambda (panel B) were fractionated by FFE (see Materials and Methods). Fractions pH ranges from approximately 6.5 (fraction 25) and increasing to 10 (fraction 68) (not shown on the graph). Aliquots (0.05 ug) of IgG from each fraction was tested by ELISA for reactivity against the indicated HIV antigens: (BaL-gp120 monomer, BaL-gp120 monomer with the D368R mutation to abrogate CD4-BS binding, Yu2 gp120 core with V3 loop, and Yu2 gp120 core, and full length single chain (FLSC), presenting a full length CD4-induced gp120 structure in which the CD4 binding site is occupied). The X axis represents the IEF fractions (spanning a pH gradient of 6 to 10 from left to right). The Y axis represents ELISA signals expressed as background-corrected OD450 readings/ug IgG. The right Y axis shows IgG concentration of each fraction (ug/ml). Assays were repeated at least twice. Areas of broad and limited neutralization previously identified based on neutralization (ability to neutralize Tier 2 viruses at <10 ug/ml of affinity purified antibody). Each fraction contains antibodies that can distinguish single epitopes. In panel A, Fractions 55-68 do not bind to D368R envelope mutants but do to the wild-type virus (BaL-gp120). Likewise, Fractions 25-30 and 35-40 in Panel A bind to FLSC (fusion protein between CD4 and gp120) but not monomeric gp120. This strongly suggests antibodies targeting a single epitope (CD4-binding site antibodies and CoReceptor binding site, respectively), as a mixed population of CD4-binding site and non-CD4 binding site antibodies would show some binding to the D368R mutant, and a mixed population of CoReceptor binding site and non-CoReceptor binding site would show binding to the monomer. In Panel B, when the fraction IgG concentrations are compared, the IgG1 anti-gp120 lambda response is almost entirely limited to fractions 28-32, suggesting that one or few antibodies are responsible for the lambda fraction (and by extension up to 60% of the total anti-gp120 response).
[0177] FIG. 2. Comparison of Heavy chains from 2 antibodies based on unique peptides. After digestion of affinity purified samples from N60, samples run through LC-MS and analyzed with patient specific database (see Example 1 methods). Orange and blue bars represent unique and non-unique peptide sequences. Grey shading of the protein sequence shows areas covered by peptide sequence. In panel A, the peptide matches are restricted to the framework regions and include only one unique peptide. This mAb did not bind or neutralize HIV. In contrast, in panel B, the there are multiple unique peptides, including coverage of the hypervariable region. This mAb, N60P2.1 bound and neutralized HIV.
[0178] FIG. 3. Free flow electrophoretic fractionation of N60 plasma anti-gp120 polyclonal antibodies and reconstructed anti-gp120 mAbs. The gray line indicates the pH (right Y axis) gradient across the fractions created by the FFE procedure. Anti-gp120 .kappa. light chain (top) or anti-gp120 .lamda. light chain (bottom) polyclonal plasma antibody preparations were processed separately (see Example 1 text and methods). The plasma antibody protein concentrations (left Y axis) detected across fractions are shown by the black trace. FFE analyses of identified and reconstructed mAbs (see Example 1) are depicted by horizontal bars. Bar width spans 75-85% of the total amount of antibodies in the FFE fraction, as determined by Elisa. Eight mAbs (checkered bars) were identified by evaluating peptides from individual FFE fractions of bulk polyclonal anti-gp120 plasma antibodies. The FFE fraction reflecting the most coverage and unique peptide pairings is indicted for each mAb by a matched-color arrow. Five additional mAbs (hatched colored bars) were identified by evaluating peptides from IEF gel fractionation of the bulk plasma antibodies. Searches using peptides digested from bulk plasma anti-gp120 antibodies identified 1 additional mAbs (solid colored bars). One other mAb (criss-cross colored bars) was identified by homology search of the Ig gene database. The pH gradient shown is for the polyclonal N60IgG1 anto-gp120 .kappa. fraction; pH gradients from each monoclonal run overlapped the trace shown, with a variance of up to 0-5 fractions in either direction.
[0179] FIG. 4. Dendrogram of variable region of all NVS60 antibodies derived from single-cell sequencing from the bone marrow. The antibodies isolated from 2013 grouped into 7 distinct families. Two families of CD4-BS antibodies were identified. The anti-gp120 antibodies represented 3.2% of all antibodies in the bone marrow database. Lineage 1 and 2 are CD4-binding site antibodies. Lineages 3-6 are CD4-induced antibodies, while Lineage 7 are variable loop 3 antibodies.
[0180] FIG. 5. ELISA Reactivity of the 7 families of antibodies isolate. Representative examples of each family is given. Dilutions of each mAb was tested by ELISA for reactivity against the indicated HIV antigens: BaL-gp120 monomer, BaL-gp120 monomer with the D368R mutation to abrogate CD4-BS binding, Yu2 gp120 core, Yu2 gp120 core +V3, and full length single chain (FLSC), presenting a full length CD4-induced gp120 structure in which the CD4-BS is occupied. N60P35 and N60P37 were also tested against the Complete V3 Loop Peptide. X-axis shows mAb concentration in ug/ml, and Y axis the background-subtracted OD. CD4-BS=CD4-binding site antibody. CoR--BS=Co--Receptor binding site antibody. CD4i=CD4-induced.
[0181] FIG. 6. Surface plasmon resonance analysis of N60 Lineage 1 mAbs to HIV envelope antigens. The binding kinetics for BaL-gp120 or D368R with mAb captured on Protein A-coated chips are shown. Data sets with significant dose response were globally fit to a 1:1 binding model to obtain the kinetic parameters of the binding. Three of the mAbs tested bound to gp120 monomer but exhibited weak binding to D368R. The other mAb demonstrated no binding to BaL gp120 or D368R.
[0182] FIG. 7. Neutralization activity plasma derived anti-Env antibodies (alone and in combination). A panel of HIV-1 viral envelope strains (individual viruses listed on the left column) that were sensitive to the bulk plasma N60 gp120-Ig were tested against neutralizing anti-CD4-BS antibodies from Lineage 1 and 2. For Lineage 2, only one mAb N60P22 tested, as the other was a closely related clone (98% sequence homology). IC.sub.50 values are color-coded according to the color key on the left: the greater the neutralization, the darker red the color; grey represents no neutralization (IC.sub.50>25 ug/ml). Taken together, the anti-CD4-BS mAbs neutralized 89% of the viruses that were sensitive to bulk plasma anti-gp120 Ig. An equimolar mix of the mAbs called N60mAb Mix1 (all CD4-BS, CD4i, and variable loop antibodies with >5% sequence divergence) were tested at equimolar concentrations neutralized 79% of the pseudoviruses, and N60mAb Mix2 (all CD4-BS antibodies with >5% sequence divergence at equimolar concentrations) neutralized 89% of the pseudoviruses. IC50=Inhibitory Concentration 50.
[0183] FIG. 8. Neutralization activity of NVS49 plasma and P series mAbs. A panel of HIV-1 viral envelope strains (individual viruses listed on the left column) were tested against all N49 plasma and CD4-BS antibodies. IC50 values are color-coded according to the color key on the left: the greater the neutralization, the darker red the color; white represents no neutralization (IC.sub.50>50 ug/ml). The individual mAbs showed extreme breadth with N49P6, N49P7, and N49P11 exhibiting 100% breadth, N49P7.1 exhibiting 99% breadth, and N49P9 exhibiting 89% breadth. IC50=Inhibitory Concentration 50.
[0184] FIG. 9. ELISA Reactivity of the N49 P series mAbs. Dilutions of each mAb was tested by ELISA for reactivity against the indicated HIV antigens: BaL-gp120 monomer, BaL-gp120 monomer with the D368R mutation to abrogate CD4-BS binding, Yu2 gp120 core, Yu2 gp120 core+V3, and full length single chain (FLSC), presenting a full length CD4-induced gp120 structure in which the CD4-BS is occupied. X-axis shows mAb concentration in ug/ml, and Y axis the background-subtracted OD.
[0185] FIG. 10. Surface plasmon resonance analysis of N49 P series mAbs to HIV-1 envelope antigens. The binding kinetics for BaL-gp120 or D368R with mAb captured on Protein A-coated chips are shown. Data sets with significant dose response were globally fit to a 1:1 binding model to obtain the kinetic parameters of the binding. All mAbs tested bound strongly to gp120 monomer but exhibited weak binding to D368R.
[0186] FIG. 11. Heavy and light chain amino acid sequences. For heavy and light chains, V(D)J sequences and 1.sup.st position of constant region are shown. Homology with germline Heavy 1-2, Lambda 2-11 J2/3, and Lambda 2-23 J2/3 are shown. Nucleotide data is given in a separate excel file (N49 neutralization and sequences).
[0187] FIG. 12. ELISA Reactivity of the N49 antibodies isolated. Dilutions of each mAb was tested by ELISA for reactivity against the indicated HIV antigens: BaL-gp120 monomer, BaL-gp120 monomer with the D368R mutation to abrogate CD4-BS binding, Yu2 gp120 core, and full length single chain (FLSC), presenting a full length CD4-induced gp120 structure in which the CD4-BS is occupied. X-axis shows mAb concentration in ug/ml, and Y axis the background-subtracted OD.
[0188] FIG. 13. Neutralization activity of N49P7 against a panel of Clade B pseudoviruses. Purified IgG was tested for neutralizing activity against the indicated pseudoviruses. SF162.LS is a Tier 1 pseudovirus, the rest fall within the Tier 2 or Tier 3 category. The monoclonal antibody N49P7 was able to neutralize all 9 viruses in our Clade B panel.
[0189] FIG. 14. Germline and natural heavy chains. A) Variable region. IMGT numbering system of germline 1-2 heavy chain shown, as well as alignment of germline 1-2 to natural sequences 1-38. B) Constant region. IMGT numbering system of CH1-3 of IgG1 shown as well as alignment of natural sequences that have point mutations in the constant region.
[0190] FIG. 15. Germline and natural light chain variable region. A) IMGT numbering system of germline IgL2-11 shown, as well as alignment of IgL2-11 with J3 to natural sequences 1-17. B) IMGT numbering system of germline IgL2-23 shown, as well as alignment of IgL2-23 with J3 shown to natural sequences 18-38.
[0191] FIG. 16. Germline and natural light chain constant region. A) IMGT numbering system of germline LC2 as well as alignment to natural sequences 1-17 that use this gene with point mutation shown. B) IMGT numbering system of germline LC7, as well as alignment to natural sequences 18-38 that use this gene, with point mutation shown.
[0192] FIG. 17. Heavy and light chain amino acid sequences for N49P6 mutants. Panel A shows heavy chain mutants N49P6 54Y, N49P6 54F, and N49P6 54YT, Panel B light chain mutants N49P6A, N49P6S, and Panel C the heavy chain constant mutant N49P6 YTE. N49P6 heavy, light, or heavy constant given as reference.
[0193] FIG. 18. Heavy and light chain amino acid sequences for N49P7 mutants. Panel A shows heavy chain mutants N49P7 54Y, N49P7 54F, and N49P7 54YT, Panel B light chain mutants N49P7A, N49P7S, and Panel C the heavy chain constant mutant N49P7 YTE. N49P7 heavy, light, or heavy constant given as reference.
[0194] FIG. 19. Crystal structure of N60P23 Fab-gp120.sub.93THO57 core.sub.e complex. (A) Ribbon diagram of N60P23 Fab-gp120.sub.93THO57 core.sub.e complex with light and heavy chains of Fab shown in light and dark green, respectively, and the complementarity-determining regions (CDRs) shown in blue (CDR L1), black (CDR L2), orange (CDR L3), pink (CDR H1), green (CDR H2), and yellow (CDR H3). The gp120 is colored in white. The D (S274-T283), V5 (T455-N465) and the CD4 binding (Q362-G372) loops are colored in cyan, violet and magenta, respectively. (B) Structural comparison of N60P23 Fab-gp120.sub.93THO57 core.sub.e complex and VRC01-gp120.sub.93THO57 core.sub.e (PDB code 3NGB) complexes. Complexes were aligned based on the gp120 and are shown as the ribbon diagrams. The light and heavy chains of VRCO1 Fab are shown in light and dark cyan and the CDRs are colored as in N60P23 complex. (C) N60P23 and VRCO1 epitope footprints. N60P23/VRCO1 contacts on gp120a3THo57Coree are highlighted in light green/cyan (light chain) and dark green/cyan (heavy and both chains) on the gp120 surface. (D) N60P23 and VRCO1 Fabs and gp12093TH057Coree contact residues on the primary sequence of the Fabs and gp120.sub.93TH057 core.sub.e, respectively. Residues contributing to the Fabs and gp12093THos7 core.sub.e are highlighted and contacts as defined by a 5A cutoff are marked above the sequence. Side chain (+) and main chain (-) contacts are colored based on contact type; hydrophobic in blue, hydrophilic in green, or both in black. Framework and complementary-determining regions are as indicated in the alignment. N60P23 binds within the CD4BS of gp120 engaging the CD4-binding loop as well as loops D and V5 also known to interact with the CD4 receptor and the CD4-binding site antibody VRC01. It recognizes the gp120 antigen with striking similarities to VRCO1 with epitope footprint almost entirety overlapping the gp120 surface involved in VRCO1 Fab-CD4-gp120.sub.93TH057 core.sub.e complex and utilizing the same CDR contacts. The close structural similarity of the complex interfaces is reflected in relatively low root mean square deviation (RMSD) value of 0.7 .ANG. for the Ca atoms of the Fab variable domains and the gp120 core.sub.e,
[0195] FIG. 20. Crystal structure of N49P7 Fab-gp120.sub.93TH057 core.sub.e complex. (A) Ribbon diagram of complex with the complementarity-determining regions (CDRs) of N49P7 Fab contributing to the gp120 binding (from light chain CDR L1 and CDR L3 and from heavy chain CDR H1, CDR H2 and CDR H3, see also Table 15) colored as shown. The gp120 outer and inner domains are colored in black and gray, respectively. The D (S.sup.27-T.sup.283), E5 (F.sup.353-T.sup.358), V5 (T.sup.455-N.sup.465) and the CD4 binding (Q.sup.362-G.sup.372) loops are colored in cyan, orange, violet and magenta, respectively. G.sup.54 of N49P7 Fab is highlighted in red and sugars at positions 276 (Loop D) and 355 (Loop E) are shown as sticks. (B) A blow up view into the network of interactions of N49P7 Fab with residues of the inner domain of gp120. Inner domain Layers 2 and 3 are colored pale green and beige, respectively. Two hydrogen bonds (CDR H2 Q.sup.56 and Layer 3 N.sup.474; CDR H3 E.sup.100E and Layer 2 K.sup.97) and a salt bridge (CDR H3 E.sup.100C and Layer 3 D.sup.477) are formed at the N49P7 Fab-gp120 inner domain interface. (C) Gp120.sub.93TH057 core.sub.e and N49P7 CDR contact residues mapped onto the primary sequence. Contact residues are defined by a 5 .ANG. cutoff and marked above the sequence. Side chain (+) and main chain (-) contacts are colored based on contact type; hydrophobic in green, hydrophilic in blue, or both in black. Buried surface residues as determined by PISA are shaded. (D) Blow up views into the Fab-gp120.sub.93TH057 core.sub.e interfaces of N49P7 Fab, N60P23 Fab and N6 Fab (PDB code: 5te6). The molecular surface is displayed over gp120 molecules with outer domain loops: D, X5, V5 and CD4 binding and inner domain Layers 2 and 3 and the 7-stranded .beta.-sandwich with CDRs colored as in panels A and B. The Fab residue at the position equivalent to F.sup.43 of CD4 is shown as sticks (Gly, His and Tyr in N49P7, N60P23 and N6, respectively). (E) The buried surface area (BSA) of gp120 residues involved in binding to N49P7, N60P23 and N6 are shown as bars (bottom panel) and their conservation among HIV-1 isolates (top panel). The conservation of the residue at particular position is shown as % difference from the Hxbc2 sequence. Only unique sequences in the database having an equivalent residue at each position were included in the calculated percentage representing approximately 32,000 sequences on average. Conserved inner domain residues uniquely targeted by N49P7 are highlighted in red.
[0196] FIG. 21. Unique features of gp120 antigen recognition mediated by N49P7. The overall structure of the N49P7 Fab-gp120.sub.93TH057 core.sub.e complex is shown in the center with the molecular surface displayed over the Fab molecule to highlight the shape of the formed antigen binding site. The complementarity-determining regions (CDRs) are shown in green (CDR L1), blue (CDR L3), black (CDR H1), yellow (CDR H2), and red (CDR H3). The gp120 outer and inner domains are colored in black and gray, respectively. The D (S.sup.274-T.sub.283), E5 (F.sup.353-T.sup.358), V5 (T.sup.455-N.sup.465) and the CD4 binding (Q.sup.362-G.sup.372) loops are colored in cyan, orange, violet and magenta, respectively. The bottom panel shows the sequence alignments of the variable light and heavy regions of N49P7 and N6 (Huang et al., 2016). The unique features of the gp120 antigen recognition mediated by N49P7 are shown (from 1 to 6) and discussed in details in the blow up view figures: (1)`Super short` CDR L1The CDRL1 of N49P7 consists of 8 amino acids (aa), 1 and 3 aa shorter than the CDRL1 of VRCOI and N6, respectively. The short CDR L1 avoids steric clashes with loop D and Loop E and permits the accommodation of complex and bulky glycans linked to N.sup.276 and N.sup.355 of gp120. The figure shows the overlay of the N49P7 Fab-gp120.sub.93TH057 core.sub.e and the N6 Fab-gp120.sub.93TH057 core.sub.e complex (Huang et al., 2016). Structures were superimposed based upon gp120 to show differences in the length and position of CDR L1 relative to the gp120 antigen (colored dark green for N49P7 and light green for N6) (2) Unpaired cysteine (C.sup.36) in a framework 2 (FW2) of the light chain. Both .kappa. (as seen in N6) and .lamda. light chains have a tyrosine at position of 36 of FW2. Replacement of this bulky residue by cysteine changes the packing of the hydrophobic core formed at the variable light (V.sub.L) and heavy (V.sub.H) domain interface of N49P7 as compared to N6. The result of the unique packing is a difference in the relative orientations (rotation/tilting) of V.sub.L. and V.sub.H observed in N49P7 relative to N6. The figure shows the V.sub.L-V.sub.H core packing of N49P7 and N6. Fabs were superimposed based of V.sub.H domain. N6 is colored light and dark blue for light and heavy chain, respectively (CD1 L1 as in panel 1). (3) Rotation/tilting of the light chain- the assembly of V.sub.L and V.sub.H domains of N49P7 forms an asymmetric antigen binding side, wildly open from the V.sub.L side and protruding from the V.sub.H side. The open access from the V.sub.L side is due to the rotation/tilting of the light chain relative to the heavy chain as described in (2). The figure shows the same superimposition as in panel 2. The V.sub.L of N49P7 is rotated approximately 12 degrees (measured at the V.sub.L N-termini) away from Loops D and VS of gp120. This generates a 5A distance between the V.sub.L N-termini of N49P7 and N6. The wild opening of the V.sub.L side of the P7 antigen binding site combined with short CDRL1 allows N49P7 to accommodate different lengths of the highly variable loops D, E and V5. Changes in the length of gp120 loop V5 and the length (and glycosylation status) of loop E that cause steric clashes with an antibody CDRL1 were described previously as mechanisms of HIV-1 resistance to VRC01-class antibodies. (4) Long CDRH3 that contacts to Loop D and the inner domain of gp120. The CDRH3 of N49P7 consists of 19 aa that contact the conserved Loop D of gp120 and reach deeply into the gp120 inner domain (compared to the 13 aa-long CDRH3 of N6). The CDRH3 contributes 274 .ANG..sup.2 BSA to the complex interface (28.6% of the BSA of the whole Fab). A network of interactions is formed which includes hydrogen bonds formed by Lys of CDRH3 (of a S.sup.100GK motif) and residues of the al helix of Layer 2 and the .alpha.7-helix of Layer 3 of the inner domain. (5) `By passing` the Phe43 cavity. The CDRH2 of N49P7 is a major anchor point in binding to the gp120 antigen contributing 489 .ANG..sup.2 BSA to the complex interface (51% of the BSA of the whole Fab) but it lacks Glycine at position 54 (Tyrosine in N6) thus it is unable to anchor deeply inside the Phe43 cavity of the CD4BS. Instead N49P7 makes important contacts to inner domain mediated trough M.sup.53 and Q.sup.56 which compensate for lack of direct contacts to residues of the Phe43 cavity. Overall N49P7 contributes 207 .ANG..sup.2 of its buried surface area (BSA) to the gp120 inner domain which is the highest among N6 and the VRCO1 Abs class. The figure shows a blow up view of the CDRH2-gp120 interface of N49P7 and N6. CDRH2 is colored light and dark yellow for N49P7 and N6, respectively. (6) The P.sup.60W motif in CDRH2. N49P7 is able to accommodate changes in length and conformation of the gp120 V5 loop through a P.sup.60W motif of its CDRH2 which forms the framework for an interaction network with the base of the V5 loop. Figure shows a close-up view of the interaction of CDRH2 of N49P7 and N6 with Loop V5. The P.sup.60W (N49P7) and G.sup.60GG (N6) motifs are highlighted.
[0197] FIG. 22. Comparison of N49P7 to other bNAbs in the literature. Viruses resistant to N6, DH511-2, or 10E8 are shown. N49P7 shows the greatest breadth and overall potency.
DETAILED DESCRIPTION OF THE INVENTION
[0198] The nature of humoral immunity generated by HIV infection provides critical insights for developing antibody-based prevention measures. While most studies are based on memory B-cell derived antibodies, in this disclosure the circulating polyclonal responses of two HIV-infected subjects with super-neutralizing HIV activity were deconvoluted through purification, fractionation, and direct sequencing of plasma antibodies. These analyses revealed that plasma anti-gp120 responses comprise a limited number of coexistent antibody lineages; only one of which (CD4-binding site) explains the bulk of the neutralizing activity. Members of one lineage (N49P series) comprised of several members, each able to neutralize 100% of isolates tested in a multitier, multiclade 117 pseudovirus panel, including all strains resistant to other broadly neutralizing antibodies. The derivation of such native antibodies with very broad cross-reactivity and potency from the plasma repertoires of multiple individuals should facilitate better understanding of the evolution of HIV humoral immunity, and inform envelope-based immunoprophylaxis strategies.
[0199] Reference will now be made in detail to the presently preferred embodiments of the invention which, together with the drawings and the following examples, serve to explain the principles of the invention. These embodiments describe in sufficient detail to enable those skilled in the art to practice the invention, and it is understood that other embodiments may be utilized, and that structural, biological, and chemical changes may be made without departing from the spirit and scope of the present invention. Unless defined otherwise, all technical and scientific terms used herein have the same meanings as commonly understood by one of ordinary skill in the art.
[0200] The practice of the present invention employs, unless otherwise indicated, conventional techniques of molecular biology (including recombinant techniques), microbiology, cell biology, biochemistry and immunology, which are within the skill of the art. Such techniques are explained fully in the literature. See, e.g., Sambrook et al. Molecular Cloning: A Laboratory Manual, 2.sup.nd edition (1989); Current Protocols in Molecular Biology (F. M. Ausubel et al. eds. (1987)); the series Methods in Enzymology (Academic Press, Inc.); PCR: A Practical Approach (M. MacPherson et al. IRL Press at Oxford University Press (1991)); PCR 2: A Practical Approach (M. J. MacPherson, B. D. Hames and G. R. Taylor eds. (1995)); Antibodies, A Laboratory Manual (Harlow and Lane eds. (1988)); Using Antibodies, A Laboratory Manual (Harlow and Lane eds. (1999)); and Animal Cell Culture (R. I. Freshney ed. (1987)). Definitions of common terms in molecular biology may be found, for example, in Benjamin Lewin, Genes VII, published by Oxford University Press, 2000 (ISBN 019879276X); Kendrew et al. (eds.); The Encyclopedia of Molecular Biology, published by Blackwell Publishers, 1994 (ISBN 0632021829); and Robert A. Meyers (ed.), Molecular Biology and Biotechnology: a Comprehensive Desk Reference, published by Wiley, John & Sons, Inc., 1995 (ISBN 0471186341).
[0201] For the purpose of interpreting this specification, the following definitions will apply and whenever appropriate, terms used in the singular will also include the plural and vice versa. In the event that any definition set forth below conflicts with the usage of that word in any other document, including any document incorporated herein by reference, the definition set forth below shall always control for purposes of interpreting this specification and its associated claims unless a contrary meaning is clearly intended (for example in the document where the term is originally used). The use of "or" means "and/or" unless stated otherwise. As used in the specification and claims, the singular form "a," "an" and "the" include plural references unless the context clearly dictates otherwise. For example, the term "a cell" includes a plurality of cells, including mixtures thereof. The use of "comprise," "comprises," "comprising," "include," "includes," and "including" are interchangeable and not intended to be limiting. Furthermore, where the description of one or more embodiments uses the term "comprising," those skilled in the art would understand that, in some specific instances, the embodiment or embodiments can be alternatively described using the language "consisting essentially of" and/or "consisting of."
[0202] Abbreviations for amino acids are used throughout this disclosure and follow the standard nomenclature known in the art. For example, as would be understood by those of ordinary skill in the art, Alanine is Ala or A; Arginine is Arg or R; Asparagine is Asn or N; Aspartic Acid is Asp or D; Cysteine is Cys or C; Glutamic acid is Glu or E; Glutamine is Gln or Q; Glycine is Gly or G; Histidine is His or H; Isoleucine is Ile or I; Leucine is Leu or L; Lysine is Lys or K; Methionine is Met or M; Phenylalanine is Phe or F; Proline is Pro or P; Serine is Ser or S; Threonine is Thr or T; Tryptophan is Trp or W; Tyrosine is Tyr or Y; and Valine is Val or V.
[0203] As used herein, the term "about" means plus or minus 10% of the numerical value of the number with which it is being used.
[0204] The term "antibody" means an immunoglobulin molecule that recognizes and specifically binds to a target, such as a protein, polypeptide, peptide, carbohydrate, polynucleotide, lipid, or combinations of the foregoing through at least one antigen recognition site within the variable region of the immunoglobulin molecule. As used herein, the term "antibody" encompasses intact polyclonal antibodies, intact monoclonal antibodies, antibody fragments (such as Fab, Fab', F(ab')2, and Fv fragments, dual affinity retargeting antibodies (DART)), single chain Fv (scFv) mutants, multispecific antibodies such as bispecific and trispecific antibodies generated from at least two intact antibodies, chimeric antibodies, humanized antibodies, human antibodies, fusion proteins comprising an antigen determination portion of an antibody, and any other modified immunoglobulin molecule comprising an antigen recognition site so long as the antibodies exhibit the desired biological activity.
[0205] In some embodiments, an antibody can be of any the five major classes of immunoglobulins: IgA, IgD, IgE, IgG, and IgM, or subclasses (isotypes) thereof (e.g. IgG1, IgG2, IgG3, IgG4, IgA1 and IgA2), based on the identity of their heavy-chain constant domains referred to as alpha, delta, epsilon, gamma, and mu, respectively. The different classes of immunoglobulins have different and well known subunit structures and three-dimensional configurations. Antibodies can be naked or conjugated to other molecules such as toxins, radioisotopes, etc.
[0206] The basic four-chain antibody unit is a heterotetrameric glycoprotein composed of two identical light (L) chains and two identical heavy (H) chains. An IgM antibody consists of 5 basic heterotetramer units along with an additional polypeptide called J chain, and therefore contain 10 antigen binding sites, while secreted IgA antibodies can polymerize to form polyvalent assemblages comprising 2-5 of the basic 4-chain units along with J chain. In the case of IgGs, the 4-chain unit is generally about 150,000 daltons. Each L chain is linked to an H chain by one covalent disulfide bond, while the two H chains are linked to each other by one or more disulfide bonds depending on the H chain isotype. Each H and L chain also has regularly spaced intrachain disulfide bridges. Each H chain has at the N-terminus, a variable region (V.sub.H) followed by three constant domains (C.sub.H) for each of the .alpha. and .gamma. chains and four C.sub.H domains for .mu. and .epsilon. isotypes. Each L chain has at the N-terminus, a variable region (V.sub.L) followed by a constant domain (C.sub.L) at its other end. The V.sub.L is aligned with the V.sub.H and the C.sub.L is aligned with the first constant domain of the heavy chain (C.sub.H1). Particular amino acid residues are believed to form an interface between the light chain and heavy chain variable regions. The pairing of a V.sub.H and V.sub.L together forms a single antigen-binding site. For the structure and properties of the different classes of antibodies, see, e.g., Basic and Clinical Immunology, 8th edition, Daniel P. Stites, Abba I. Terr and Tristram G. Parslow (eds.), Appleton & Lange, Norwalk, Conn., 1994, page 71, and Chapter 6.
[0207] The L chain from any vertebrate species can be assigned to one of two clearly distinct types, called kappa (.kappa.) and lambda (.lamda.), based on the amino acid sequences of their constant domains (C.sub.L). Depending on the amino acid sequence of the constant domain of their heavy chains (C.sub.H), immunoglobulins can be assigned to different classes or isotypes. There are five classes of immunoglobulins: IgA, IgD, IgE, IgG, and IgM, having heavy chains designated alpha (.alpha.), delta (.delta.), epsilon (.epsilon.), gamma (.gamma.) and mu (.mu.) respectively. The .gamma. and .alpha. classes are further divided into subclasses on the basis of relatively minor differences in C.sub.H sequence and function, e.g., humans express the following subclasses: IgG1, IgG2, IgG3, IgG4, IgA1, and IgA2.
[0208] The terms "antigen" or "immunogen" are used interchangeably to refer to a substance, typically a protein, which is capable of inducing an immune response in a subject. The term also refers to proteins that are immunologically active in the sense that once administered to a subject (either directly or by administering to the subject a nucleotide sequence or vector that encodes the protein) is able to evoke an immune response of the humoral and/or cellular type directed against that protein.
[0209] The term "antigen binding fragment" or antibody fragment refers to a portion of an intact antibody and comprises the antigenic determining variable regions of an intact antibody. Examples of antigen binding fragment include, but are not limited to Fab, Fab', F(ab')2, and Fv fragments, linear antibodies, single chain antibodies, and multispecific antibodies formed from antibody fragments.
[0210] A "monoclonal antibody" refers to a homogeneous antibody population involved in the highly specific recognition and binding of a single antigenic determinant, or epitope. This is in contrast to polyclonal antibodies that typically include different antibodies directed against different antigenic determinants. The term "monoclonal antibody" encompasses both intact and full-length monoclonal antibodies as well as antibody fragments (such as Fab, Fab', F(ab')2, Fv), single chain (scFv) mutants, fusion proteins comprising an antibody portion, and any other modified immunoglobulin molecule comprising an antigen recognition site. Furthermore, "monoclonal antibody" refers to such antibodies made in any number of manners including but not limited to by hybridoma, phage selection, recombinant expression, and transgenic animals.
[0211] The term "humanized antibody" refers to forms of non-human (e.g. murine) antibodies that are specific immunoglobulin chains, chimeric immunoglobulins, or fragments thereof that contain minimal non-human (e.g., murine) sequences. Typically, humanized antibodies are human immunoglobulins in which residues from the complementary determining region (CDR) are replaced by residues from the CDR of a non-human species (e.g. mouse, rat, rabbit, hamster) that have the desired specificity, affinity, and capability (Jones et al., 1986, Nature, 321:522-525; Riechmann et al., 1988, Nature, 332:323-327; Verhoeyen et al., 1988, Science, 239:1534-1536). In some instances, the Fv framework region (FR) residues of a human immunoglobulin are replaced with the corresponding residues in an antibody from a non-human species that has the desired specificity, affinity, and capability. The humanized antibody can be further modified by the substitution of additional residues either in the Fv framework region and/or within the replaced non-human residues to refine and optimize antibody specificity, affinity, and/or capability. In general, the humanized antibody will comprise substantially all of at least one, and typically two or three, variable domains containing all or substantially all of the CDR regions that correspond to the non-human immunoglobulin whereas all or substantially all of the FR regions are those of a human immunoglobulin consensus sequence. The humanized antibody can also comprise at least a portion of an immunoglobulin constant region or domain (Fc), typically that of a human immunoglobulin. Examples of methods used to generate humanized antibodies are described in U.S. Pat. No. 5,225,539 or 5,639,641.
[0212] A "variable region" of an antibody refers to the variable region of the antibody light chain or the variable region of the antibody heavy chain, either alone or in combination. The variable regions of the heavy and light chain each consist of four framework regions (FR) connected by three complementarity determining regions (CDRs) also known as hypervariable regions. The CDRs in each chain are held together in close proximity by the FRs and, with the CDRs from the other chain, contribute to the formation of the antigen-binding site of antibodies. The term "hypervariable region" when used herein refers to the amino acid residues of an antibody that are responsible for antigen binding. The hypervariable region generally comprises amino acid residues from a "complementarity determining region" or "CDR" (e.g., around about residues 24-34 (L1), 50-56 (L2) and 89-97 (L3) in the V.sub.L, and around about 31-35 (H1), 50-65 (H2) and 95-102 (H3) in the V.sub.H when numbered in accordance with the Kabat numbering system; Kabat et al., Sequences of Proteins of Immunological Interest, 5th Ed. Public Health Service, National Institutes of Health, Bethesda, Md. (1991)); and/or those residues from a "hypervariable loop" (e.g., residues 24-34 (L1), 50-56 (L2) and 89-97 (L3) in the V.sub.L, and 26-32 (H1), 52-56 (H2) and 95-101 (H3) in the V.sub.H when numbered in accordance with the Chothia numbering system; Chothia and Lesk, J. Mol. Biol. 196:901-917 (1987)); and/or those residues from a "hypervariable loop"/CDR (e.g., residues 27-38 (L1), 56-65 (L2) and 105-120 (L3) in the V.sub.L, and 27-38 (H1), 56-65 (H2) and 105-120 (H3) in the V.sub.H when numbered in accordance with the IMGT numbering system; Lefranc, M. P. et al. Nucl. Acids Res. 27:209-212 (1999), Ruiz, M. e al. Nucl. Acids Res. 28:219-221 (2000)).
[0213] The IMGT unique numbering has been defined to compare the variable domains whatever the antigen receptor, the chain type, or the species [Lefranc M.-P., Immunology Today 18, 509 (1997)/Lefranc M.-P., The Immunologist, 7, 132-136 (1999)/Lefranc, M.-P., Pommie, C., Ruiz, M., Giudicelli, V., Foulquier, E., Truong, L., Thouvenin-Contet, V. and Lefranc, Dev. Comp. Immunol., 27, 55-77 (2003)]. In the IMGT unique numbering, the conserved amino acids always have the same position, for instance cysteine 23 (1st-CYS), tryptophan 41 (CONSERVED-TRP), hydrophobic amino acid 89, cysteine 104 (2nd-CYS), phenylalanine or tryptophan 118 (J-PHE or J-TRP). The IMGT unique numbering provides a standardized delimitation of the framework regions (FR1-IMGT: positions 1 to 26, FR2-IMGT: 39 to 55, FR3-IMGT: 66 to 104 and FR4-IMGT: 118 to 128) and of the complementarity determining regions: CDR1-IMGT: 27 to 38, CDR2-IMGT: 56 to 65 and CDR3-IMGT: 105 to 117. As gaps represent unoccupied positions, the CDR-IMGT lengths (shown between brackets and separated by dots, e.g. [8.8.13]) become crucial information. The IMGT unique numbering is used in 2D graphical representations, designated as IMGT Colliers de Perles (Ruiz, M. and Lefranc, M.-P., Immunogenetics, 53, 857-883 (2002)/Kaas, Q. and Lefranc, M.-P., Current Bioinformatics, 2, 21-30 (2007)), and in 3D structures in IMGT/3Dstructure-DB (Kaas, Q., Ruiz, M. and Lefranc, M.-P., T cell receptor and MHC structural data. Nucl. Acids. Res., 32, D208-D210 (2004)).
[0214] In some embodiments, CDRs are determined based on cross-species sequence variability (i.e., Kabat et al. Sequences of Proteins of Immunological Interest, (5th ed., 1991, National Institutes of Health, Bethesda Md.)). In some embodiments, CDRs are determined based on crystallographic studies of antigen-antibody complexes (Al-lazikani et al (1997) J. Molec. Biol. 273:927-948)). In addition, combinations of these two approaches can be used to determine CDRs. In some embodiments, the CDRs are determined based on AHo (Honegger and Pluckthun, J. Mol. Biol. 309(3):657-670; 2001). In some embodiments, CDRs are determined based on the IMGT system.
[0215] The term "human antibody" means an antibody produced by a human or an antibody having an amino acid sequence corresponding to an antibody produced by a human made using any technique known in the art. This definition of a human antibody includes intact or full-length antibodies, fragments thereof, and/or antibodies comprising at least one human heavy and/or light chain polypeptide such as, for example, an antibody comprising murine light chain and human heavy chain polypeptides.
[0216] A "neutralizing antibody" may inhibit the entry of HIV-1 virus for example SF162 and/or JR-CSF with a neutralization index >1.5 or >2.0. (Kostrikis L G et al. J Virol. 1996; 70(1): 445-458.). By "broad and potent neutralizing antibodies" are meant antibodies that neutralize more than one HIV-1 virus species (from diverse clades and different strains within a clade) in a neutralization assay. A broad neutralizing antibody may neutralize at least 2, 3, 4, 5, 6, 7, 8, 9 or more different strains of HIV-1, the strains belonging to the same or different clades. A broad neutralizing antibody may neutralize multiple HIV-1 species belonging to at least 2, 3, 4, 5, or 6 different clades. In some embodiments, the \concentration of the monoclonal antibody able to neutralize at 50% of the input virus in the neutralization assay can be less than about 50 .mu.g/ml.
[0217] An "intact" antibody is one that comprises an antigen-binding site as well as a C.sub.L and at least heavy chain constant domains, C.sub.H1, C.sub.H2 and C.sub.H3. The constant domains may be native sequence constant domains (e.g., human native sequence constant domains) or amino acid sequence variants thereof.
[0218] The term "chimeric antibodies" refers to antibodies wherein the amino acid sequence of the immunoglobulin molecule is derived from two or more species. Typically, the variable region of both light and heavy chains corresponds to the variable region of antibodies derived from one species of mammals (e.g. mouse, rat, rabbit, etc) with the desired specificity, affinity, and capability while the constant regions are homologous to the sequences in antibodies derived from another (usually human) to avoid eliciting an immune response in that species.
[0219] The antibodies herein also include antibodies in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies belonging to a particular antibody class or subclass, while the remainder of the chain(s) is identical with or homologous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies.
[0220] In some embodiments, the antibody comprises variable region antigen-binding sequences derived from human antibodies (e.g., CDRs) and containing one or more sequences derived from a non-human antibody, e.g., an FR or C region sequence. In some embodiments, the antibody includes those comprising a human variable region antigen binding sequence of one antibody class or subclass and another sequence, e.g., FR or C region sequence, derived from another antibody class or subclass.
[0221] In some embodiments, chimeric antibodies may comprise residues that are not found in the recipient antibody or in the donor antibody. In some embodiments, modifications are made to further refine antibody performance. For further details, see Jones et al., Nature 321:522-525 (1986); Riechmann et al., Nature 332:323-329 (1988); and Presta, Curr. Op. Struct. Biol. 2:593-596 (1992).
[0222] The term "epitope" or "antigenic determinant" are used interchangeably herein and refer to that portion of an antigen capable of being recognized and specifically bound by a particular antibody. When the antigen is a polypeptide, epitopes can be formed both from contiguous amino acids and noncontiguous amino acids juxtaposed by tertiary folding of a protein. Epitopes formed from contiguous amino acids are typically retained upon protein denaturing, whereas epitopes formed by tertiary folding are typically lost upon protein denaturing. An epitope typically includes at least 3, and more usually, at least 5 or 8-10 amino acids in a unique spatial conformation.
[0223] "Binding affinity" generally refers to the strength of the sum total of noncovalent interactions between a single binding site of a molecule (e.g., an antibody) and its binding partner (e.g., an antigen). Unless indicated otherwise, as used herein, "binding affinity" refers to intrinsic binding affinity which reflects a 1:1 interaction between members of a binding pair (e.g., antibody and antigen). The affinity of a molecule X for its partner Y can generally be represented by the dissociation constant (Kd). Low-affinity antibodies generally bind antigen slowly and tend to dissociate readily, whereas high-affinity antibodies generally bind antigen faster and tend to remain bound longer.
[0224] The affinity or avidity of an antibody for an antigen can be determined experimentally using any suitable method well known in the art, e.g. flow cytometry, enzyme-linked immunoabsorbent assay (ELISA), or radioimmunoassay (RIA), or kinetics (e.g., BIACORE'analysis). Direct binding assays as well as competitive binding assay formats can be readily employed. (See, for example, Berzofsky, et al., "Antibody-Antigen Interactions," In Fundamental Immunology, Paul, W. E., Ed., Raven Press: New York, N.Y. (1984); Kuby, Janis Immunology, W.H. Freeman and Company: New York, N.Y. (1992); and methods described herein. The measured affinity of a particular antibody-antigen interaction can vary if measured under different conditions (e.g., salt concentration, pH, temperature). Thus, measurements of affinity and other antigen-binding parameters (e.g., KD or Kd, K.sub.on, K.sub.off) are made with standardized solutions of antibody and antigen, and a standardized buffer, as known in the art and such as the buffer described herein.
[0225] The phrase "substantially similar," or "substantially the same", as used herein, denotes a sufficiently high degree of similarity between two numeric values (generally one associated with an antibody of the invention and the other associated with a reference/comparator antibody) such that one of skill in the art would consider the difference between the two values to be of little or no biological and/or statistical significance within the context of the biological characteristics measured by said values (e.g., Kd values). The difference between said two values is less than about 500%, less than about 40%, less than about 300%, less than about 200%, or less than about 10% as a function of the value for the reference/comparator antibody.
[0226] A polypeptide, antibody, polynucleotide, vector, cell, or composition which is "isolated" is a polypeptide, antibody, polynucleotide, vector, cell, or composition which is in a form not found in nature. Isolated polypeptides, antibodies, polynucleotides, vectors, cell or compositions include those which have been purified to a degree that they are no longer in a form in which they are found in nature. In some embodiments, an antibody, polynucleotide, vector, cell, or composition which is isolated is substantially pure.
[0227] An "isolated nucleic acid" is a nucleic acid that is substantially separated from other genome DNA sequences as well as proteins or complexes such as ribosomes and polymerases, which naturally accompany a native sequence. The term embraces a nucleic acid sequence that has been removed from its naturally occurring environment, and includes recombinant or cloned DNA isolates and chemically synthesized analogues or analogues biologically synthesized by heterologous systems. A substantially pure nucleic acid includes isolated forms of the nucleic acid. Of course, this refers to the nucleic acid as originally isolated and does not exclude genes or sequences later added to the isolated nucleic acid by the hand of man.
[0228] An "isolated polypeptide" is one that has been identified and separated and/or recovered from a component of its natural environment. In preferred embodiments, the isolated polypeptide will be purified (1) to greater than 95% by weight of polypeptide as determined by the Lowry method, and most preferably more than 99% by weight, (2) to a degree sufficient to obtain at least 15 residues of N-terminal or internal amino acid sequence by use of a spinning cup sequenator, or (3) to homogeneity by SDS-PAGE under reducing or non-reducing conditions using Coomassie blue or, preferably, silver stain. Isolated polypeptide includes the polypeptide in situ within recombinant cells since at least one component of the polypeptide's natural environment will not be present.
[0229] A "native sequence" polynucleotide is one that has the same nucleotide sequence as a polynucleotide derived from nature. A "native sequence" polypeptide is one that has the same amino acid sequence as a polypeptide (e.g., antibody) derived from nature (e.g., from any species). Such native sequence polynucleotides and polypeptides can be isolated from nature. A polynucleotide "variant," as the term is used herein, is a polynucleotide that typically differs from a polynucleotide specifically disclosed herein in one or more substitutions, deletions, additions and/or insertions. Such variants may be naturally occurring or may be synthetically generated, for example, by modifying one or more of the polynucleotide sequences of the invention and evaluating one or more biological activities of the encoded polypeptide as described herein and/or using any of a number of techniques well known in the art.
[0230] A polypeptide "variant," as the term is used herein, is a polypeptide that typically differs from a polypeptide specifically disclosed herein in one or more substitutions, deletions, additions and/or insertions. Such variants may be naturally occurring or may be synthetically generated, for example, by modifying one or more of the above polypeptide sequences of the invention and evaluating one or more biological activities of the polypeptide as described herein and/or using any of a number of techniques well known in the art. or can be produced by recombinant or synthetic means.
[0231] As used herein, "substantially pure" refers to material which is at least 50% pure (i.e., free from contaminants), at least 90% pure, at least 95% pure, at least 98% pure, or at least 99% pure.
[0232] The term "subject" refers to any animal (e.g., a mammal), including, but not limited to humans, non-human primates, rodents, and the like, which is to be the recipient of a particular treatment. Typically, the terms "subject" and "patient" are used interchangeably herein in reference to a human subject.
[0233] Administration "in combination with" one or more further therapeutic agents includes simultaneous (concurrent) and consecutive administration in any order.
[0234] The term "pharmaceutical formulation" refers to a preparation which is in such form as to permit the biological activity of the active ingredient to be effective, and which contains no additional components which are unacceptably toxic to a subject to which the formulation would be administered. Such formulation can be sterile.
[0235] An "effective amount" of an antibody as disclosed herein is an amount sufficient to carry out a specifically stated purpose. An "effective amount" can be determined empirically and in a routine manner, in relation to the stated purpose.
[0236] The term "therapeutically effective amount" refers to an amount of an antibody or other drug effective to "treat" or prevent a disease or disorder in a subject or mammal.
[0237] Terms such as "treating" or "treatment" or "to treat" or "alleviating" or "to alleviate" refer to both 1) therapeutic measures that cure, slow down, lessen symptoms of, and/or halt progression of a diagnosed pathologic condition or disorder and 2) prophylactic or preventative measures that prevent and/or slow the development of a targeted pathologic condition or disorder. Thus, those in need of treatment include those already with the disorder; those prone to have the disorder; and those in whom the disorder is to be prevented.
[0238] "Polynucleotide," or "nucleic acid," as used interchangeably herein, refer to polymers of nucleotides of any length, and include DNA and RNA. The nucleotides can be deoxyribonucleotides, ribonucleotides, modified nucleotides or bases, and/or their analogs, or any substrate that can be incorporated into a polymer by DNA or RNA polymerase. A polynucleotide can comprise modified nucleotides, such as methylated nucleotides and their analogs. If present, modification to the nucleotide structure can be imparted before or after assembly of the polymer. The sequence of nucleotides can be interrupted by non-nucleotide components. A polynucleotide can be further modified after polymerization, such as by conjugation with a labeling component. Other types of modifications include, for example, "caps", substitution of one or more of the naturally occurring nucleotides with an analog, internucleotide modifications such as, for example, those with uncharged linkages (e.g., methyl phosphonates, phosphotriesters, phosphoamidates, cabamates, etc.) and with charged linkages (e.g., phosphorothioates, phosphorodithioates, etc.), those containing pendant moieties, such as, for example, proteins (e.g., nucleases, toxins, antibodies, signal peptides, ply-L-lysine, etc.), those with intercalators (e.g., acridine, psoralen, etc.), those containing chelators (e.g., metals, radioactive metals, boron, oxidative metals, etc.), those containing alkylators, those with modified linkages (e.g., alpha anomeric nucleic acids, etc.), as well as unmodified forms of the polynucleotide(s). Further, any of the hydroxyl groups ordinarily present in the sugars can be replaced, for example, by phosphonate groups, phosphate groups, protected by standard protecting groups, or activated to prepare additional linkages to additional nucleotides, or can be conjugated to solid supports. The 5' and 3' terminal OH can be phosphorylated or substituted with amines or organic capping group moieties of from 1 to 20 carbon atoms. Other hydroxyls can also be derivatized to standard protecting groups. Polynucleotides can also contain analogous forms of ribose or deoxyribose sugars that are generally known in the art, including, for example, 2'-O-methyl-, 2'-O-allyl, 2'-fluoro- or 2'-azido-ribose, carbocyclic sugar analogs, alpha.-anomeric sugars, epimeric sugars such as arabinose, xyloses or lyxoses, pyranose sugars, furanose sugars, sedoheptuloses, acyclic analogs and abasic nucleoside analogs such as methyl riboside. One or more phosphodiester linkages can be replaced by alternative linking groups. These alternative linking groups include, but are not limited to, embodiments wherein phosphate is replaced by P(O)S ("thioate"), P(S)S ("dithioate"), "(O)NR.sub.2 ("amidate"), P(O)R, P(O)OR', CO or CH.sub.2 ("formacetal"), in which each R or R' is independently H or substituted or unsubstituted alkyl (1-20 C) optionally containing an ether (--O--) linkage, aryl, alkenyl, cycloalkyl, cycloalkenyl or araldyl. Not all linkages in a polynucleotide need be identical. The preceding description applies to all polynucleotides referred to herein, including RNA and DNA.
[0239] The terms "polypeptide," "peptide," and "protein" are used interchangeably herein to refer to polymers of amino acids of any length. The polymer can be linear or branched, it can comprise modified amino acids, and it can be interrupted by non-amino acids. The terms also encompass an amino acid polymer that has been modified naturally or by intervention; for example, disulfide bond formation, glycosylation, lipidation, acetylation, phosphorylation, or any other manipulation or modification, such as conjugation with a labeling component. Also included within the definition are, for example, polypeptides containing one or more analogs of an amino acid (including, for example, unnatural amino acids, etc.), as well as other modifications known in the art. It is understood that, because the polypeptides of this invention are based upon antibodies, in certain embodiments, the polypeptides can occur as single chains or associated chains.
[0240] The terms "identical" or percent "identity" in the context of two or more nucleic acids or polypeptides, refer to two or more sequences or subsequences that are the same or have a specified percentage of nucleotides or amino acid residues that are the same, when compared and aligned (introducing gaps, if necessary) for maximum correspondence, not considering any conservative amino acid substitutions as part of the sequence identity. The percent identity can be measured using sequence comparison software or algorithms or by visual inspection. Various algorithms and software are known in the art that can be used to obtain alignments of amino acid or nucleotide sequences. One such non-limiting example of a sequence alignment algorithm is the algorithm described in Karlin et al, 1990, Proc. Natl. Acad. Sci., 87:2264-2268, as modified in Karlin et al., 1993, Proc. Natl. Acad. Sci., 90:5873-5877, and incorporated into the NBLAST and XBLAST programs (Altschul et al., 1991, Nucleic Acids Res., 25:3389-3402). In certain embodiments, Gapped BLAST can be used as described in Altschul et al., 1997, Nucleic Acids Res. 25:3389-3402. BLAST-2, WU-BLAST-2 (Altschul et al., 1996, Methods in Enzymology, 266:460-480), ALIGN, ALIGN-2 (Genentech, South San Francisco, Calif.) or Megalign (DNASTAR) are additional publicly available software programs that can be used to align sequences. In certain embodiments, the percent identity between two nucleotide sequences is determined using the GAP program in GCG software (e.g., using a NWSgapdna.CMP matrix and a gap weight of 40, 50, 60, 70, or 90 and a length weight of 1, 2, 3, 4, 5, or 6). In certain alternative embodiments, the GAP program in the GCG software package, which incorporates the algorithm of Needleman and Wunsch (J. Mol. Biol. (48):444-453 (1970)) can be used to determine the percent identity between two amino acid sequences (e.g., using either a Blossum 62 matrix or a PAM250 matrix, and a gap weight of 16, 14, 12, 10, 8, 6, or 4 and a length weight of 1, 2, 3, 4, 5). Alternatively, in certain embodiments, the percent identity between nucleotide or amino acid sequences is determined using the algorithm of Myers and Miller (CABIOS, 4:11-17 (1989)). For example, the percent identity can be determined using the ALIGN program (version 2.0) and using a PAM120 with residue table, a gap length penalty of 12 and a gap penalty of 4. Appropriate parameters for maximal alignment by particular alignment software can be determined by one skilled in the art. In certain embodiments, the default parameters of the alignment software are used. In certain embodiments, the percentage identity "X" of a first amino acid sequence to a second sequence amino acid is calculated as 100.times.(Y/Z), where Y is the number of amino acid residues scored as identical matches in the alignment of the first and second sequences (as aligned by visual inspection or a particular sequence alignment program) and Z is the total number of residues in the second sequence. If the length of a first sequence is longer than the second sequence, the percent identity of the first sequence to the second sequence will be longer than the percent identity of the second sequence to the first sequence.
[0241] As a non-limiting example, whether any particular polynucleotide has a certain percentage sequence identity (e.g., is at least 80% identical, at least 85% identical, at least 90% identical, and in some embodiments, at least 95%, 96%, 97%, 98%, or 99% identical) to a reference sequence can, in certain embodiments, be determined using the Bestfit program (Wisconsin Sequence Analysis Package, Version 8 for Unix, Genetics Computer Group, University Research Park, 575 Science Drive, Madison, Wis. 53711). Bestfit uses the local homology algorithm of Smith and Waterman. Advances in Applied Mathematics 2: 482 489 (1981), to find the best segment of homology between two sequences. When using Bestfit or any other sequence alignment program to determine whether a particular sequence is, for instance, 95% identical to a reference sequence according to the present invention, the parameters are set such that the percentage of identity is calculated over the full length of the reference nucleotide sequence and that gaps in homology of up to 5% of the total number of nucleotides in the reference sequence are allowed.
[0242] In some embodiments, two nucleic acids or polypeptides of the invention are substantially identical, meaning they have at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, and in some embodiments at least 95%, 96%, 97%, 98%, 99% nucleotide or amino acid residue identity, when compared and aligned for maximum correspondence, as measured using a sequence comparison algorithm or by visual inspection. In certain embodiments, identity exists over a region of the sequences that is at least about 10, about 20, about 40-60 residues in length or any integral value therebetween, or over a longer region than 60-80 residues, at least about 90-100 residues, or the sequences are substantially identical over the full length of the sequences being compared, such as the coding region of a nucleotide sequence for example.
[0243] A "conservative amino acid substitution" is one in which one amino acid residue is replaced with another amino acid residue having a similar side chain. Families of amino acid residues having similar side chains have been defined in the art, including basic side chains (e.g., lysine, arginine, histidine), acidic side chains (e.g., aspartic acid, glutamic acid), uncharged polar side chains (e.g., asparagine, glutamine, serine, threonine, tyrosine, cysteine), nonpolar side chains (e.g., glycine, alanine, valine, leucine, isoleucine, proline, phenylalanine, methionine, tryptophan), beta-branched side chains (e.g., threonine, valine, isoleucine) and aromatic side chains (e.g., tyrosine, phenylalanine, tryptophan, histidine). For example, substitution of a phenylalanine for a tyrosine is a conservative substitution. In certain embodiments, conservative substitutions in the sequences of the polypeptides and antibodies of the invention do not abrogate the binding of the polypeptide or antibody containing the amino acid sequence, to the antigen(s), i.e., the gp120 to which the polypeptide or antibody binds. Methods of identifying nucleotide and amino acid conservative substitutions which do not eliminate antigen binding are well-known in the art (see, e.g., Brummell et al., Biochem. 32: 1180-1187 (1993); Kobayashi et al. Protein Eng. 12(10):879-884 (1999); and Burks et al. Proc. Natl. Acad. Sci. USA 94:412-417 (1997)).
Anti-HIV Antibodies
[0244] In some embodiments, the invention provides antibodies that are broadly neutralizing antibodies against HIV.
[0245] HIV-1 is among the most genetically diverse viral pathogens. Of the three main branches of the HIV-1 phylogenetic tree, the M (main), N (new), and 0 (outlier) groups, group M viruses are the most widespread, accounting for over 99% of global infections. This group is presently divided into nine distinct genetic subtypes, or clades (A through K), based on full-length sequences. Env is the most variable HIV-1 gene, with up to 35% sequence diversity between clades, 20% sequence diversity within clades, and up to 10% sequence diversity in a single infected person (Shankarappa, R. et al. 1999. J. Virol. 73:10489-10502). Clade B is dominant in Europe, the Americas, and Australia. Clade C is common in southern Africa, China, and India and presently infects more people worldwide than any other clade (McCutchan, F E. 2000. Understanding the genetic diversity of HIV-1. AIDS 14(Suppl. 3):S31-S44). Clades A and D are prominent in central and eastern Africa.
[0246] In some embodiments, the invention provides antibodies that are broadly neutralizing against HIV. In some embodiments, the antibody has a particularly high potency in neutralizing HIV infection in vitro across multiple clades as shown in the Figures and Tables 5, 6, and 16-21 herein. Such antibodies are desirable, as only low concentrations are required in order to neutralize a given amount of virus. This facilitates higher levels of protection while administering lower amounts of antibody.
[0247] In some embodiments, the invention provides a broadly neutralizing anti-HIV antibody wherein the antibody neutralizes HIV-1 species belonging to two or more clades.
[0248] In some embodiments, the anti-HIV antibody neutralizes at least 95%, at least 96%, at least 97%, at least 98%, at least 99% or 100% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the antibody is selected from N49P6 or an antigen binding fragment thereof, N49P7 or an antigen binding fragment thereof, N49P7.1 or an antigen binding fragment thereof, or N49P11 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein.
[0249] In some embodiments, the anti-HIV antibody neutralizes 100% of the HIV clade B, G and D viruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. See also FIG. 8 and Tables 16-21 herein. In some embodiments, the antibody is selected from N49P6 or an antigen binding fragment thereof, N49P7 or an antigen binding fragment thereof, N49P7.1 or an antigen binding fragment thereof, N49P11 or an antigen binding fragment thereof, or N49P9 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and/or V.sub.L regions of N49P6, N49P7, N49P7.1, N49P11 or N49P9 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P6, N49P7, N49P7.1, N49P11 or N49P9 as described herein.
[0250] In some embodiments, the anti-HIV antibody neutralizes 100% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the anti-HIV antibody neutralizes at least 95%, at least 96%, at least 97%, at least 98%, at least 99% or 100% of the HIV pseudoviruses encompassed by Table 1 and strain CNE5 (clade CRF01_AE) with an IC50 value of less than 50 .mu.g/mL. In some embodiments, the antibody is selected from N49P6 or an antigen binding fragment thereof, N49P7 or an antigen binding fragment thereof, N49P7.1 or an antigen binding fragment thereof, or N49P11 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein.
[0251] In embodiment aspect, the invention provides an isolated anti-HIV antibody, wherein the antibody is capable of neutralizing HIV pseudoviruses listed in Table 1 with a median IC50 value of less than 0.5 .mu.g/mL. In some embodiments, the antibody is selected from the group consisting of [0252] a. N49P6 or an antigen binding fragment thereof; [0253] b. N49P7 or an antigen binding fragment thereof; [0254] c. N49P7.1 or an antigen binding fragment thereof [0255] d. N49P9 or an antigen binding fragment thereof; and [0256] e. N49P23 or an antigen binding fragment thereof.
[0257] In some embodiments, the anti-HIV antibody neutralizes at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, or at least 99% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than about 1 .mu.g/ml, between about 1-5 .mu.g/ml or greater than about 5 .mu.g/ml.
[0258] In some embodiments, the anti-HIV antibody neutralizes at least about 86.4% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P7 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P7 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P7 as described herein.
[0259] In some embodiments, the anti-HIV antibody neutralizes at least 88.7% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P7.1 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P7.1 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P7.1 as described herein.
[0260] In some embodiments, the anti-HIV antibody neutralizes at least 84.5% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P7.2 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P7.2 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P7.2 as described herein.
[0261] In some embodiments, the anti-HIV antibody neutralizes at least 71.8% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P6 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P6 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P6 as described herein.
[0262] In some embodiments, the anti-HIV antibody neutralizes at least 93.3% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P9 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P9 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P9 as described herein.
[0263] In some embodiments, the anti-HIV antibody neutralizes at least 91.1% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P9.1 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P9.1 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P9.1 as described herein.
[0264] In some embodiments, the anti-HIV antibody neutralizes at least 41.9% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P11 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P11 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P11 as described herein.
[0265] In some embodiments, the anti-HIV antibody neutralizes at least 2.1% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P18.1 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P18.1 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P18.1 as described herein.
[0266] In some embodiments, the anti-HIV antibody neutralizes at least 60% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P19 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P19 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P19 as described herein.
[0267] In some embodiments, the anti-HIV antibody neutralizes at least 58.3% of the HIV viruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P22 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P22 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions regions of N49P22 as described herein.
[0268] In some embodiments, the anti-HIV antibody neutralizes at least 88.6% of the HIV viruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. In some embodiments, the antibody is N49P23 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the V.sub.H and V.sub.L regions of N49P23 as described herein. In some embodiments, the antibody comprises the CDRs of the V.sub.H and V.sub.L regions of N49P23 as described herein.
[0269] The neutralization can be performed using a luciferase-based assay in TZM.b1 cells as described by M. M. Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011) and M. Li et al., J Virol 79, 10108-10125 (2005)). This assay measures the reduction in luciferase expression following a single round of virus infection.
TABLE-US-00001 TABLE 1 HIV Env pseudovirus panel. Virus ID Clade* 6535.3 B QH0692.42 B SC422661.8 B PVO.4 B TRO.11 B AC10.0.29 B RHPA4259.7 B THRO4156.18 B REJO4541.67 B TRJO4551.58 B WITO4160.33 B CAAN5342.A2 B WEAU_d15_410_787 B (T/F) 1006_11_C3_1601 B (T/F) 1054_07_TC4_1499 B (T/F) 1056_10_TA11_1826 B (T/F) 1012_11_TC21_3257 B (T/F) 6240_08_TA5_4622 B (T/F) 6244_13_B5_4576 B (T/F) 62357_14_D3_4589 B (T/F) SC05_8C11_2344 B (T/F) Du156.12 C Du172.17 C Du422.1 C ZM197M.PB7 C ZM214M.PL15 C ZM233M.PB6 C ZM249M.PL1 C ZM53M.PB12 C ZM109F.PB4 C ZM135M.PL10a C CAP45.2.00.G3 C CAP210.2.00.E8 C HIV-001428-2.42 C HIV-0013095-2.11 C HIV-16055-2.3 C HIV-16845-2.22 C Ce1086_B2 C (T/F) Ce0393_C3 C (T/F) Cel176_A3 C (T/F) Ce2010_F5 C (T/F) Ce0682_E4 C (T/F) Ce1172_H1 C (T/F) Ce2060_G9 C (T/F) Ce703010054_2A2 C (T/F) BF1266.431a C (T/F) 246F C1G C (T/F) 249M B10 C (T/F) ZM247v1(Rev-) C (T/F) 7030102001E5(Rev-) C (T/F) 1394C9G1(Rev-) C (T/F) Ce704809221_1B3 C (T/F) CNE19 BC CNE20 BC CNE21 BC CNE17 BC CNE30 BC CNE52 BC CNE53 BC CNE58 BC MS208.A1 A Q23.17 A Q461.e2 A Q769.d22 A Q259.d2.17 A Q842.d12 A 3415.v1.c1 A 3365.v2.c2 A 0260.v5.c36 A 191955_A11 A (T/F) 191084 B7-19 A (T/F) 9004SS_A3_4 A (T/F) T257-31 CRF02_AG 928-28 CRF02_AG 263-8 CRF02_AG T250-4 CRF02_AG T251-18 CRF02_AG T278-50 CRF02_AG T255-34 CRF02_AG 211-9 CRF02_AG 235-47 CRF02_AG 620345.c01 CRF01_AE CNE8 CRF01_AE C1080.c03 CRF01_AE R2184.c04 CRF01_AE R1166.c01 CRF01_AE R3265.c06 CRF01_AE C2101.c01 CRF01_AE C3347.c11 CRF01_AE C4118.c09 CRF01_AE BJOX009000.02.4 CRF01_AE BJOX015000.11.5 CRF01_AE (T/F) BJOX010000.06.2 CRF01_AE (T/F) BJOX025000.01.1 CRF01_AE (T/F) BJOX028000.10.3 CRF01_AE (T/F) X1193_c1 G P0402_c2_c11 G X1254_c3 G X2088_c9 G X2131_C1_B5 G P1981_C5_3 G X1632_S2_B10 G 3016.v5.c45 D A07412M1.vrc12 D 231965.c01 D 231966.c02 D 3817.v2.c59 CD 6480.v4.c25 CD 6952.v1.c20 CD 6811.v7.c18 CD 89-F1_2_25 CD 3301.v1.c24 AC 6041.v3.c23 AC 6540.v4.c1 AC 6545.v4.c1 AC 0815.v3.c3 ACD 3103.v3.c10 ACD * (T/F): Transmitted/Founder Virus
[0270] Methods for producing antibodies, such as those disclosed herein, are known in the art. For example, DNA molecules encoding light chain variable regions and/or heavy chain variable regions can be chemically synthesized using the sequence information provided herein. Synthetic DNA molecules can be ligated to other appropriate nucleotide sequences, including, e.g., expression control sequences, to produce conventional gene expression constructs encoding the desired antibodies. Production of defined gene constructs is within routine skill in the art. Alternatively, the sequences provided herein can be cloned out of hybridomas by conventional hybridization techniques or polymerase chain reaction (PCR) techniques, using synthetic nucleic acid probes whose sequences are based on sequence information provided herein, or prior art sequence information regarding genes encoding the heavy and light chains.
[0271] Standard techniques of molecular biology may be used to prepare DNA sequences coding for the antibodies or fragments of the antibodies of the present invention. Desired DNA sequences may be synthesized completely or in part using oligonucleotide synthesis techniques. Site-directed mutagenesis and polymerase chain reaction (PCR) techniques may be used as appropriate.
[0272] Any suitable host cell/vector system may be used for expression of the DNA sequences encoding the antibody molecules of the present invention or fragments thereof. Bacterial, for example E. coli, and other microbial systems may be used, in part, for expression of antibody fragments such as Fab and F(ab')2 fragments, and especially Fv fragments and single chain antibody fragments, for example, single chain Fvs. Eukaryotic, e.g. mammalian, host cell expression systems may be used for production of larger antibody molecules, including complete antibody molecules. Suitable mammalian host cells include CHO, HEK293T, PER.C6, myeloma or hybridoma cells.
[0273] In some embodiments, antibodies according to the invention may be produced by i) expressing a nucleic acid sequence according to the invention in a cell, and ii) isolating the expressed antibody product. Additionally, the method may include iii) purifying the antibody.
[0274] For the antibodies of the present invention to be expressed, the protein coding sequence should be "operably linked" to regulatory or nucleic acid control sequences that direct transcription and translation of the protein. As used herein, a coding sequence and a nucleic acid control sequence or promoter are said to be "operably linked" when they are covalently linked in such a way as to place the expression or transcription and/or translation of the coding sequence under the influence or control of the nucleic acid control sequence. The "nucleic acid control sequence" can be any nucleic acid element, such as, but not limited to promoters, enhancers, IRES, introns, and other elements described herein that direct the expression of a nucleic acid sequence or coding sequence that is operably linked thereto. The term "promoter" will be used herein to refer to a group of transcriptional control modules that are clustered around the initiation site for RNA polymerase II and that when operationally linked to the protein coding sequences of the invention lead to the expression of the encoded protein. The expression of the antibodies of the present invention can be under the control of a constitutive promoter or of an inducible promoter, which initiates transcription only when exposed to some particular external stimulus, such as, without limitation, antibiotics such as tetracycline, hormones such as ecdysone, or heavy metals. The promoter can also be specific to a particular cell-type, tissue or organ. Many suitable promoters and enhancers are known in the art, and any such suitable promoter or enhancer may be used for expression of the antibodies of the invention. For example, suitable promoters and/or enhancers can be selected from the Eukaryotic Promoter Database (EPDB).
[0275] Nucleic acids encoding desired antibodies can be incorporated (ligated) into expression vectors, which can be introduced into host cells through conventional transfection or transformation techniques. Exemplary host cells are E. coli cells, Chinese hamster ovary (CHO) cells, human embryonic kidney 293 (HEK 293) cells, HeLa cells, baby hamster kidney (BHK) cells, monkey kidney cells (COS), human hepatocellular carcinoma cells (e.g., Hep G2), and myeloma cells that do not otherwise produce IgG protein. Transformed host cells can be grown under conditions that permit the host cells to express the genes that encode the immunoglobulin light and/or heavy chain variable regions. Specific expression and purification conditions will vary depending upon the expression system employed.
[0276] Following expression, the antibodies and/or antigens of the invention can be isolated and/or purified or concentrated using any suitable technique known in the art. For example, anion or cation exchange chromatography, phosphocellulose chromatography, hydrophobic interaction chromatography, affinity chromatography, immuno-affinity chromatography, hydroxyapatite chromatography, lectin chromatography, molecular sieve chromatography, isoelectric focusing, gel electrophoresis, or any other suitable method or combination of methods can be used.
[0277] In some embodiments, the antibodies can be made using recombinant DNA methods as described in U.S. Pat. No. 4,816,567. The polynucleotides encoding a monoclonal antibody can be isolated from mature B-cells or hybridoma cell, such as by RT-PCR using oligonucleotide primers that specifically amplify the genes encoding the heavy and light chains of the antibody, and their sequence is determined using conventional procedures. The isolated polynucleotides encoding the heavy and light chains are then cloned into suitable expression vectors, which when transfected into host cells such as E. coli cells, simian COS cells, Chinese hamster ovary (CHO) cells, or myeloma cells that do not otherwise produce immunoglobulin protein, monoclonal antibodies are generated by the host cells.
[0278] The anti-HIV antibodies can also include insertions, deletions, substitutions, or other selected modifications of particular regions or specific amino acids residues. It should be understood that the antibodies of the invention may differ from the exact sequences illustrated and described herein. Thus, the invention contemplates deletions, additions and substitutions to the sequences shown, so long as the sequences function in accordance with the methods of the invention. In this regard, particularly preferred substitutions will generally be conservative in nature, i.e., those substitutions that take place within a family of amino acids. For example, amino acids are generally divided into four families: (1) acidic--aspartate and glutamate; (2) basic--lysine, arginine, histidine; (3) non-polar--alanine, valine, leucine, isoleucine, proline, phenylalanine, methionine, tryptophan; and (4) uncharged polar--glycine, asparagine, glutamine, cystine, serine threonine, tyrosine. Phenylalanine, tryptophan, and tyrosine are sometimes classified as aromatic amino acids. For example, leucine can be replaced with isoleucine or valine, or vice versa; an aspartate with a glutamate or vice versa; a threonine with a serine or vice versa; or a similar conservative replacement of an amino acid with a structurally related amino acid can be made.
[0279] The polynucleotide(s) encoding a monoclonal antibody can further be modified in a number of different manners using recombinant DNA technology to generate alternative antibodies. In some embodiments, the constant domains of the light and heavy chains of, for example, a mouse monoclonal antibody can be substituted 1) for those regions of, for example, a human antibody to generate a chimeric antibody or 2) for a non-immunoglobulin polypeptide to generate a fusion antibody. In some embodiments, the constant regions are truncated or removed to generate the desired antibody fragment of a monoclonal antibody. Site-directed or high-density mutagenesis of the variable region can be used to optimize specificity, affinity, etc. of a monoclonal antibody.
[0280] For the purposes of the present invention, it should be appreciated that modified antibodies can comprise any type of variable region that provides for the association of the antibody with the polypeptides of HIV such as gp120.
[0281] In some embodiments, the variable regions or domains in both the heavy and light chains are altered by at least partial replacement of one or more CDRs and, if necessary, by partial framework region replacement and sequence changing. Although the CDRs can be derived from an antibody of the same class or even subclass as the antibody from which the framework regions are derived, in some embodiments the CDRs will be derived from an antibody of different class.
[0282] Alterations to the variable region notwithstanding, those skilled in the art will appreciate that the modified antibodies of this invention can comprise antibodies (e.g., full-length antibodies or immunoreactive fragments thereof) in which at least a fraction of one or more of the constant region domains has been deleted or otherwise altered so as to provide desired biochemical characteristics such as increased localization, increased serum half-life or reduced serum half-life when compared with an antibody of approximately the same immunogenicity comprising a native or unaltered constant region. In some embodiments, the constant region of the modified antibodies will comprise a human constant region. Modifications to the constant region compatible with this invention comprise additions, deletions or substitutions of one or more amino acids in one or more domains. That is, the modified antibodies disclosed herein can comprise alterations or modifications to one or more of the three heavy chain constant domains (CH1, CH2 or CH3) and/or to the light chain constant domain (CL). In some embodiments, modified constant regions wherein one or more domains are partially or entirely deleted are contemplated. In some embodiments, the modified antibodies will comprise domain deleted constructs or variants wherein the entire CH2 domain has been removed (.DELTA.CH2 constructs). In some embodiments, the omitted constant region domain will be replaced by a short amino acid spacer (e.g. 10 residues) that provides some of the molecular flexibility typically imparted by the absent constant region.
[0283] Besides their configuration, it is known in the art that the constant region mediates several effector functions. For example, binding of the C1 component of complement to antibodies activates the complement system. Activation of complement is important in the opsonisation and lysis of cell pathogens. The activation of complement also stimulates the inflammatory response and can also be involved in autoimmune hypersensitivity. Further, antibodies bind to cells via the Fc region, with a Fc receptor site on the antibody Fc region binding to a Fc receptor (FcR) on a cell. There are a number of Fc receptors which are specific for different classes of antibody, including IgG (gamma receptors), IgE (eta receptors), IgA (alpha receptors) and IgM (mu receptors). Binding of antibody to Fc receptors on cell surfaces triggers a number of important and diverse biological responses including engulfment and destruction of antibody-coated particles, clearance of immune complexes, lysis of antibody-coated target cells by killer cells (called antibody-dependent cell-mediated cytotoxicity, or ADCC), release of inflammatory mediators, placental transfer and control of immunoglobulin production.
[0284] In certain embodiments, the anti-HIV antibodies provide for altered effector functions that, in turn, affect the biological profile of the administered antibody. For example, the deletion or inactivation (through point mutations or other means) of a constant region domain can reduce Fc receptor binding of the circulating modified antibody thereby increasing tumor localization. In other cases it may be that constant region modifications, consistent with this invention, moderate complement binding and thus reduce the serum half life and nonspecific association of a conjugated cytotoxin. Yet other modifications of the constant region can be used to eliminate disulfide linkages or oligosaccharide moieties that allow for enhanced localization due to increased antigen specificity or antibody flexibility. Similarly, modifications to the constant region in accordance with this invention can easily be made using well known biochemical or molecular engineering techniques well within the purview of the skilled artisan.
[0285] In certain embodiments, the invention provides antibodies or antigen binding fragments that specifically bind to an HIV antigen, such as gp120. In some embodiments, the invention is directed to a broadly neutralizing antibody against HIV wherein the antibody binds an epitope on gp120. In some embodiments, the invention is directed to a broadly neutralizing antibody against HIV wherein the antibody binds an epitope on the CD4 binding site (CD4-BS). In some embodiments, the invention is directed to a broadly neutralizing antibody against HIV wherein the antibody binds an epitope on V1V2 glycan. In some embodiments, the invention is directed to a broadly neutralizing antibody against HIV wherein the antibody binds an epitope on V3 glycan. In some embodiments, the invention is directed to a broadly neutralizing antibody against HIV wherein the antibody binds an epitope on the gp41 membrane-proximal external region.
[0286] X-ray crystallography analysis of pan-neutralizing monoclonal, N49P7, identified its unique ability to bypass the CD4-binding site Phe43 cavity while reaching deep into the highly conserved residues of Layer 3 of the gp120 inner domain, likely accounting for its pan-neutralization. Deletion in the CDR L1 (not found in N6) combined with the rotation/tilting of the light chain `opens` the variable light (V1) side of the N49P7 antigen binding site to accommodate different lengths of the highly variable loops D, E and VS (FIG. 21). Changes in the length of gp120 loop VS and the length (and glycosylation status) of loop E that cause steric clashes with an antibody light chain were described previously as mechanisms of HIV-I resistance to VRC01-class antibodies (Lynch et al., 2015). These signatures, along with a long CDRH3 (21 aa) and unique sequence signatures within CDR.B2 (not seen in VRCO1 and N6, FIG. 21), allow N49P7 to bypass the Phe43 cavity affording it an unusual capacity to contact the gp120 inner domain at residues 97, 102 and 124 of Layer 2, and 472-480 of Layer 3.
[0287] In some embodiments, the conformational interdomain CD4 binding site epitope is formed by combination of residues of both outer and inner domain of gp120 of HIV Env. These generally involve residues of gp120 outer domain at position (HXBc2 numbering): 275-283 (Loop D), 354-371 (CD4 binding loop), 427-439 (bridging sheet) and 455-463 (loop V5) and residues of gp120 inner domain at positions: 96-106 (helix alpha-1 of Layer 2) and 469-480 (loop prior and helix alpha-5 of Layer 3).
[0288] In some embodiments, the anti-HIV antibody binds to a HIV gp120 epitope comprising outer domain loop D (which comprises 275-283), the CD4 binding loop (which comprises 354-371), the bridging sheet (which comprises 427-439) and loop V5 (which comprises 455-463) and gp120 inner domain at positions 96-106 (helix alpha-1 of Layer 2) and 469-480 (loop prior and helix alpha-5 of Layer 3). In some embodiments, the anti-HIV antibody binding the aforementioned epitope is from the antibody lineage as shown in FIG. 15A. In some embodiments, the anti-HIV antibody is selected from N49P6 or an antigen binding fragment thereof, N49P7 or an antigen binding fragment thereof, N49P7.1 or an antigen binding fragment thereof, and N49P11 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the VH and VL regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein. In some embodiments, the antibody comprises the CDRs of the VH and VL regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 1.59.times.10.sup.-8M. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 1.562.times.10.sup.-8M. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 1.143.times.10.sup.-9M. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 8.602.times.10.sup.-10 M. In some embodiments, the binding affinity is determined by surface plasmon resonance. See FIG. 10.
[0289] In some embodiments, the anti-HIV antibody binds to a HIV gp120 epitope comprising the specific residues as described in FIG. 20E. In some embodiments, the anti-HIV antibody binds to gp120 Layer 2 residues W96, K97, E102, G124, Loop D residues E275, N276, T278, N279, N280, A281, K282, CD4 binding loop residues P364, 5365, G366, G367, D368, 1371, bridging sheet residues W427, Q428, G429, Loop V5 residues T455, R456, D457, G458, G459, A460, N461, T463), and Layer 3 residues R469, P470, G471, G472, G473, N474, K476, D477, R480. In some embodiments, the anti-HIV antibody binding the aforementioned epitope is from the antibody lineage as shown in FIG. 15A. In some embodiments, the anti-HIV antibody is selected from N49P6 or an antigen binding fragment thereof, N49P7 or an antigen binding fragment thereof, N49P7.1 or an antigen binding fragment thereof, and N49P11 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the VH and VL regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein. In some embodiments, the antibody comprises the CDRs of the VH and VL regions of N49P6, N49P7, N49P7.1 or N49P11 as described herein. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 1.59.times.10.sup.-8M. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 1.562.times.10.sup.-8M. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 1.143.times.10.sup.-9M. In some embodiments, the antibody has a Kd for BaL-gp120 of at least about 8.602.times.10.sup.-10 M. In some embodiments, the binding affinity is determined by surface plasmon resonance. See FIG. 10.
[0290] In some embodiments, the anti-HIV antibody binds to the same epitope as antibody N49P6, N49P7, N49P7.1, and/or N49P11.
[0291] In some embodiments, the anti-HIV antibody is an antibody that binds to the same epitope as an antibody selected from the group consisting of N49P6; N49P6.2; N49P7; N49P7.1; N49P7A; N49P7S; N49P7F; N49P7Y; N49P7-54TY; N49P7LS-1; N49P7LS-2; N49P7YTE; N49P7L6; N49P7L11; N49P7.1L9; N49P7.1L19; R49P7; N49P7.2; N49P11; N49P18; N49P18.2; N49P18.1; N49P19; N49P37; N49P38; N49P38.1; and N49P55.
[0292] In some embodiments, the anti-HIV antibody is an antibody that binds to the same epitope as antibody N49P7.
[0293] In some embodiments, the anti-HIV antibody is an antibody that binds to the same epitope as antibody N49P6.
[0294] In some embodiments, the anti-HIV antibody is an antibody that binds to the same epitope as antibody N49P7.1.
[0295] In some embodiments, the anti-HIV antibody is an antibody that binds to the same epitope as antibody N49P11.
[0296] In some embodiments, the anti-HIV antibody comprises a heavy chain comprising an amino acid sequence selected from the group consisting of SEQ ID NOS:1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73 and 75.
[0297] In some embodiments, the anti-HIV antibody comprises an antigen binding fragment of an amino acid sequence selected from the group consisting of SEQ ID NOS:1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73 and 75.
[0298] In some embodiments, the anti-HIV antibody comprises a light chain comprising an amino acid sequence selected from the group consisting of SEQ ID NOS:2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74 and 76.
[0299] In some embodiments, the anti-HIV antibody comprises an antigen binding fragment of an amino acid sequence selected from the group consisting of SEQ ID NOS:2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74 and 76.
[0300] In some embodiments, the anti-HIV antibody comprises a heavy chain comprising an amino acid sequence selected from the group consisting of SEQ ID NOS:153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 and 397.
[0301] In some embodiments, the anti-HIV antibody comprises an antigen binding fragment of an amino acid sequence selected from the group consisting of SEQ ID NOS:153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 and 397.
[0302] In some embodiments, the anti-HIV antibody comprises a light chain comprising an amino acid sequence selected from the group consisting of SEQ ID NOS:155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 and 399.
[0303] In some embodiments, the anti-HIV antibody comprises an antigen binding fragment of an amino acid sequence selected from the group consisting of SEQ ID NOS:155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 and 399.
[0304] In some embodiments, the anti-HIV antibody is selected from the group consisting of: [0305] a. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:1 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:2 or an antigen binding fragment thereof; [0306] b. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:3 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:4 or an antigen binding fragment thereof; [0307] c. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:5 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:6 or an antigen binding fragment thereof; [0308] d. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:7 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:8 or an antigen binding fragment thereof; [0309] e. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:9 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:10 or an antigen binding fragment thereof; [0310] f. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:11 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:12 or an antigen binding fragment thereof; [0311] g. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:13 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:14 or an antigen binding fragment thereof; [0312] h. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:15 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:16 or an antigen binding fragment thereof; [0313] i. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:17 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:18 or an antigen binding fragment thereof; [0314] j. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:19 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:20 or an antigen binding fragment thereof; [0315] k. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:21 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:22 or an antigen binding fragment thereof; [0316] l. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:23 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:24 or an antigen binding fragment thereof; [0317] m. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:25 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:26 or an antigen binding fragment thereof; [0318] n. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:27 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:28 or an antigen binding fragment thereof; [0319] o. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:29 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:30 or an antigen binding fragment thereof; [0320] p. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:31 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:32 or an antigen binding fragment thereof; [0321] q. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:33 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:34 or an antigen binding fragment thereof; [0322] r. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:35 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:36 or an antigen binding fragment thereof; [0323] s. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:37 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:38 or an antigen binding fragment thereof; [0324] t. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:39 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:40 or an antigen binding fragment thereof; [0325] u. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:41 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:42 or an antigen binding fragment thereof; [0326] v. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:43 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:44 or an antigen binding fragment thereof; [0327] w. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:45 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:46 or an antigen binding fragment thereof; [0328] x. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:47 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:48 or an antigen binding fragment thereof; [0329] y. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:49 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:50 or an antigen binding fragment thereof; [0330] z. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:51 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:52 or an antigen binding fragment thereof; [0331] aa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:53 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:54 or an antigen binding fragment thereof; [0332] bb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:55 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:56 or an antigen binding fragment thereof; [0333] cc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:57 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:58 or an antigen binding fragment thereof; [0334] dd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:59 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:60 or an antigen binding fragment thereof; [0335] ee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:61 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:62 or an antigen binding fragment thereof; [0336] ff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:63 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:64 or an antigen binding fragment thereof; [0337] gg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:65 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:66 or an antigen binding fragment thereof; [0338] hh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:67 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:68 or an antigen binding fragment thereof; [0339] ii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:69 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:70 or an antigen binding fragment thereof; [0340] jj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:71 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:72 or an antigen binding fragment thereof; [0341] kk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:73 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:74 or an antigen binding fragment thereof; [0342] ll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:75 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:76 or an antigen binding fragment thereof; [0343] mm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:153 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:155 or an antigen binding fragment thereof; [0344] nn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:157 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:159 or an antigen binding fragment thereof; [0345] oo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:161 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:163 or an antigen binding fragment thereof; [0346] pp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:165 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:167 or an antigen binding fragment thereof; [0347] qq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:169 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:171 or an antigen binding fragment thereof; [0348] rr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:173 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:175 or an antigen binding fragment thereof; [0349] ss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:177 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:179 or an antigen binding fragment thereof; [0350] tt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:181 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:183 or an antigen binding fragment thereof; [0351] uu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:185 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:187 or an antigen binding fragment thereof; [0352] vv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:189 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:191 or an antigen binding fragment thereof; [0353] ww. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:193 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:195 or an antigen binding fragment thereof; [0354] xx. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:197 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:199 or an antigen binding fragment thereof; [0355] yy. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:201 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:203 or an antigen binding fragment thereof; [0356] zz. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:205 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:207 or an antigen binding fragment thereof; [0357] aaa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:209 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:211 or an antigen binding fragment thereof; [0358] bbb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:213 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:215 or an antigen binding fragment thereof; [0359] ccc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:217 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:219 or an antigen binding fragment thereof; [0360] ddd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:221 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:223 or an antigen binding fragment thereof; [0361] eee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:225 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:227 or an antigen binding fragment thereof; [0362] fff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:229 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:231 or an antigen binding fragment thereof; [0363] ggg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:233 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:235 or an antigen binding fragment thereof; [0364] hhh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:237 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:239 or an antigen binding fragment thereof; [0365] iii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:241 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:243 or an antigen binding fragment thereof; [0366] jjj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:245 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:247 or an antigen binding fragment thereof; [0367] kkk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:249 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:251 or an antigen binding fragment thereof; [0368] lll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:253 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:255 or an antigen binding fragment thereof;
[0369] mmm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:257 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:259 or an antigen binding fragment thereof; [0370] nnn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:261 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:263 or an antigen binding fragment thereof; [0371] ooo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:265 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:267 or an antigen binding fragment thereof; [0372] ppp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:269 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:271 or an antigen binding fragment thereof; [0373] qqq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:273 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:275 or an antigen binding fragment thereof; [0374] rrr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:277 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:279 or an antigen binding fragment thereof; [0375] sss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:281 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:283 or an antigen binding fragment thereof; [0376] ttt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:285 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:287 or an antigen binding fragment thereof; [0377] uuu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:289 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:291 or an antigen binding fragment thereof; [0378] vvv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:293 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:295 or an antigen binding fragment thereof; [0379] www. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:297 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:299 or an antigen binding fragment thereof; [0380] xxx. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:301 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:303 or an antigen binding fragment thereof; [0381] yyy. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:305 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:307 or an antigen binding fragment thereof; [0382] zzz. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:309 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:311 or an antigen binding fragment thereof; [0383] aaaa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:313 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:315 or an antigen binding fragment thereof; [0384] bbbb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:317 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:319 or an antigen binding fragment thereof; [0385] cccc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:321 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:323 or an antigen binding fragment thereof; [0386] dddd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:325 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:327 or an antigen binding fragment thereof; [0387] eeee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:329 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:331 or an antigen binding fragment thereof; [0388] ffff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:333 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:335 or an antigen binding fragment thereof; [0389] gggg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:337 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:339 or an antigen binding fragment thereof; [0390] hhhh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:341 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:343 or an antigen binding fragment thereof; [0391] iiii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:345 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:347 or an antigen binding fragment thereof; [0392] jjjj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:349 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:351 or an antigen binding fragment thereof; [0393] kkkk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:353 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:355 or an antigen binding fragment thereof; [0394] llll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:357 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:359 or an antigen binding fragment thereof; [0395] mmmm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:361 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:363 or an antigen binding fragment thereof; [0396] nnnn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:365 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:367 or an antigen binding fragment thereof; [0397] oooo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:369 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:371 or an antigen binding fragment thereof; [0398] pppp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:373 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:375 or an antigen binding fragment thereof; [0399] qqqq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:377 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:379 or an antigen binding fragment thereof; [0400] rrrr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:381 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:383 or an antigen binding fragment thereof; [0401] ssss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:385 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:387 or an antigen binding fragment thereof; [0402] tttt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:389 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:391 or an antigen binding fragment thereof; [0403] uuuu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:393 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:395 or an antigen binding fragment thereof; and [0404] vvvv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:397 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:399 or an antigen binding fragment thereof.
[0405] In some embodiments, the anti-HIV antibody is isolated and/or substantially pure.
[0406] In some embodiments, the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain comprises a heavy chain variable (VH) region and the light chain comprises a light chain variable (VL) region; wherein the VL region comprises one or more VL complementary determining regions (CDRs) and wherein the VH region comprises one or more VH complementary determining regions (CDRs), wherein the VL CDRs correspond to the CDRs found within any of SEQ ID NOS:2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 and 399.
[0407] In some embodiments, the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain comprises a heavy chain variable (VH) region and the light chain comprises a light chain variable (VL) region; wherein the VL region comprises one or more VL complementary determining regions (CDRs) and wherein the VH region comprises one or more VH complementary determining regions (CDRs), wherein the VH CDRs correspond to the CDRs found within any of SEQ ID NOS:1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73, 75, 153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 and 397.
[0408] In some embodiments, the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain comprises a heavy chain variable (VH) region and the light chain comprises a light chain variable (VL) region; wherein the VL region comprises one or more VL complementary determining regions (CDRs) and wherein the VH region comprises one or more VH complementary determining regions (CDRs), wherein the VL CDRs correspond to the CDRs found within any of SEQ ID NOS: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 and 399 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions, and wherein the VH CDRs correspond to the CDRs found within any of SEQ ID NOS:1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73, 75, 153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 and 397 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions.
[0409] In some embodiments, the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain comprises a heavy chain variable (VH) region and the light chain comprises a light chain variable (VL) region; wherein the VL region comprises an amino acid sequence selected from the group consisting of: amino acids 1-99 of SEQ ID NO:2; amino acids 1-99 of SEQ ID NO:4; amino acids 1-99 of SEQ ID NO:6; amino acids 1-99 of SEQ ID NO:8; amino acids 1-99 of SEQ ID NO:10; amino acids 1-99 of SEQ ID NO:12; amino acids 1-99 of SEQ ID NO:14; amino acids 1-99 of SEQ ID NO:16; amino acids 1-99 of SEQ ID NO:18; amino acids 1-99 of SEQ ID NO:20; amino acids 1-99 of SEQ ID NO:22; amino acids 1-99 of SEQ ID NO:24; amino acids 1-99 of SEQ ID NO:26; amino acids 1-99 of SEQ ID NO:28; amino acids 1-99 of SEQ ID NO:30; amino acids 1-99 of SEQ ID NO:32; amino acids 1-99 of SEQ ID NO:34; amino acids 1-100 of SEQ ID NO:36; amino acids 1-97 of SEQ ID NO:38; amino acids 1-100 of SEQ ID NO:40; amino acids 1-100 of SEQ ID NO:42; amino acids 1-97 of SEQ ID NO:44; amino acids 1-101 of SEQ ID NO:46; amino acids 1-101 of SEQ ID NO:48; amino acids 1-96 of SEQ ID NO:50; amino acids 1-97 of SEQ ID NO:52; amino acids 1-99 of SEQ ID NO:54; amino acids 1-99 of SEQ ID NO:56; amino acids 1-99 of SEQ ID NO:58; amino acids 1-99 of SEQ ID NO:60; amino acids 1-98 of SEQ ID NO:62; amino acids 1-99 of SEQ ID NO:64; amino acids 1-99 of SEQ ID NO:66; amino acids 1-96 of SEQ ID NO:68; amino acids 1-96 of SEQ ID NO:70; amino acids 1-96 of SEQ ID NO:72; amino acids 1-101 of SEQ ID NO:74; amino acids 1-97 of SEQ ID NO:76; amino acids 1-99 of SEQ ID NO:155; amino acids 1-99 of SEQ ID NO:159; amino acids 1-99 of SEQ ID NO:163; amino acids 1-99 of SEQ ID NO:167; amino acids 1-99 of SEQ ID NO:171; amino acids 1-99 of SEQ ID NO:175; amino acids 1-99 of SEQ ID NO:179; amino acids 1-99 of SEQ ID NO:183; amino acids 1-99 of SEQ ID NO:187; amino acids 1-99 of SEQ ID NO:191; amino acids 1-99 of SEQ ID NO:195; amino acids 1-99 of SEQ ID NO:199; amino acids 1-99 of SEQ ID NO:203; amino acids 1-99 of SEQ ID NO:207; amino acids 1-100 of SEQ ID NO:211; amino acids 1-99 of SEQ ID NO:215; amino acids 1-99 of SEQ ID NO:219; amino acids 1-99 of SEQ ID NO:223; amino acids 1-99 of SEQ ID NO:227; amino acids 1-99 of SEQ ID NO:231; amino acids 1-99 of SEQ ID NO:235; amino acids 1-99 of SEQ ID NO:239; amino acids 1-99 of SEQ ID NO:243; amino acids 1-99 of SEQ ID NO:247; amino acids 1-99 of SEQ ID NO:251; amino acids 1-99 of SEQ ID NO:255; amino acids 1-99 of SEQ ID NO:259; amino acids 1-99 of SEQ ID NO:263; amino acids 1-99 of SEQ ID NO:267; amino acids 1-99 of SEQ ID NO:271; amino acids 1-99 of SEQ ID NO:275; amino acids 1-99 of SEQ ID NO:279; amino acids 1-99 of SEQ ID NO:283; amino acids 1-99 of SEQ ID NO:287; amino acids 1-99 of SEQ ID NO:291; amino acids 1-100 of SEQ ID NO:295; amino acids 1-100 of SEQ ID NO:299; amino acids 1-100 of SEQ ID NO:303; amino acids 1-100 of SEQ ID NO:307; amino acids 1-100 of SEQ ID NO:311; amino acids 1-100 of SEQ ID NO:315; amino acids 1-97 of SEQ ID NO:319; amino acids 1-100 of SEQ ID NO:323; amino acids 1-100 of SEQ ID NO:327; amino acids 1-100 of SEQ ID NO:331; amino acids 1-97 of SEQ ID NO:335; amino acids 1-101 of SEQ ID NO:339; amino acids 1-101 of SEQ ID NO:343; amino acids 1-96 of SEQ ID NO:347; amino acids 1-97 of SEQ ID NO:351; amino acids 1-99 of SEQ ID NO:355; amino acids 1-99 of SEQ ID NO:359; amino acids 1-99 of SEQ ID NO:363; amino acids 1-99 of SEQ ID NO:367; amino acids 1-98 of SEQ ID NO:371; amino acids 1-99 of SEQ ID NO:375; amino acids 1-99 of SEQ ID NO:379; amino acids 1-96 of SEQ ID NO:383; amino acids 1-96 of SEQ ID NO:387; amino acids 1-96 of SEQ ID NO:391; amino acids 1-101 of SEQ ID NO:395; and amino acids 1-97 of SEQ ID NO:399 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions.
[0410] In some embodiments, the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain comprises a heavy chain variable (VH) region and the light chain comprises a light chain variable (VL) region; wherein the VH region comprises an amino acid sequence selected from the group consisting of: amino acids 1-128 of SEQ ID NO:1; amino acids 1-127 of SEQ ID NO:3; amino acids 1-127 of SEQ ID NO:5; amino acids 1-128 of SEQ ID NO:7; amino acids 1-127 of SEQ ID NO:9; amino acids 1-127 of SEQ ID NO:11; amino acids 1-127 of SEQ ID NO:13; amino acids 1-127 of SEQ ID NO:15; amino acids 1-127 of SEQ ID NO:17; amino acids 1-127 of SEQ ID NO:19; amino acids 1-127 of SEQ ID NO:21; amino acids 1-127 of SEQ ID NO:23; amino acids 1-127 of SEQ ID NO:25; amino acids 1-127 of SEQ ID NO:27; amino acids 1-127 of SEQ ID NO:29; amino acids 1-127 of SEQ ID NO:31; amino acids 1-127 of SEQ ID NO:33; amino acids 1-120 of SEQ ID NO:35; amino acids 1-120 of SEQ ID NO:37; amino acids 1-123 of SEQ ID NO:39; amino acids 1-120 of SEQ ID NO:41; amino acids 1-120 of SEQ ID NO:43; amino acids 1-125 of SEQ ID NO:45; amino acids 1-125 of SEQ ID NO:47; amino acids 1-120 of SEQ ID NO:49; amino acids 1-120 of SEQ ID NO:51; amino acids 1-121 of SEQ ID NO:53; amino acids 1-121 of SEQ ID NO:55; amino acids 1-121 of SEQ ID NO:57; amino acids 1-121 of SEQ ID NO:59; amino acids 1-120 of SEQ ID NO:61; amino acids 1-121 of SEQ ID NO:63; amino acids 1-121 of SEQ ID NO:65; amino acids 1-120 of SEQ ID NO:67; amino acids 1-120 of SEQ ID NO:69; amino acids 1-120 of SEQ ID NO:71; amino acids 1-125 of SEQ ID NO:73; amino acids 1-120 of SEQ ID NO:75; amino acids 1-128 of SEQ ID NO:153; amino acids 1-128 of SEQ ID NO:157; amino acids 1-127 of SEQ ID NO:161; amino acids 1-127 of SEQ ID NO:165; amino acids 1-127 of SEQ ID NO:169; amino acids 1-127 of SEQ ID NO:173; amino acids 1-127 of SEQ ID NO:177; amino acids 1-127 of SEQ ID NO:181; amino acids 1-127 of SEQ ID NO:185; amino acids 1-127 of SEQ ID NO:189; amino acids 1-127 of SEQ ID NO:193; amino acids 1-127 of SEQ ID NO:197; amino acids 1-127 of SEQ ID NO:201; amino acids 1-127 of SEQ ID NO:205; amino acids 1-127 of SEQ ID NO:209; amino acids 1-127 of SEQ ID NO:213; amino acids 1-127 of SEQ ID NO:217; amino acids 1-127 of SEQ ID NO:221; amino acids 1-128 of SEQ ID NO:225; amino acids 1-127 of SEQ ID NO:229; amino acids 1-127 of SEQ ID NO:233; amino acids 1-127 of SEQ ID NO:237; amino acids 1-127 of SEQ ID NO:241; amino acids 1-127 of SEQ ID NO:245; amino acids 1-127 of SEQ ID NO:249; amino acids 1-127 of SEQ ID NO:253; amino acids 1-127 of SEQ ID NO:257; amino acids 1-127 of SEQ ID NO:261; amino acids 1-127 of SEQ ID NO:265; amino acids 1-127 of SEQ ID NO:269; amino acids 1-127 of SEQ ID NO:273; amino acids 1-127 of SEQ ID NO:277; amino acids 1-127 of SEQ ID NO:281; amino acids 1-127 of SEQ ID NO:285; amino acids 1-127 of SEQ ID NO:289; amino acids 1-120 of SEQ ID NO:293; amino acids 1-120 of SEQ ID NO:297; amino acids 1-120 of SEQ ID NO:301; amino acids 1-123 of SEQ ID NO:305; amino acids 1-128 of SEQ ID NO:309; amino acids 1-128 of SEQ ID NO:313; amino acids 1-120 of SEQ ID NO:317; amino acids 1-123 of SEQ ID NO:321; amino acids 1-120 of SEQ ID NO:325; amino acids 1-120 of SEQ ID NO:329; amino acids 1-120 of SEQ ID NO:333; amino acids 1-125 of SEQ ID NO:337; amino acids 1-125 of SEQ ID NO:341; amino acids 1-120 of SEQ ID NO:345; amino acids 1-120 of SEQ ID NO:349; amino acids 1-121 of SEQ ID NO:353; amino acids 1-121 of SEQ ID NO:357; amino acids 1-121 of SEQ ID NO:361; amino acids 1-121 of SEQ ID NO:365; amino acids 1-120 of SEQ ID NO:369; amino acids 1-121 of SEQ ID NO:373; amino acids 1-121 of SEQ ID NO:377; amino acids 1-120 of SEQ ID NO:381; amino acids 1-120 of SEQ ID NO:385; amino acids 1-120 of SEQ ID NO:389; amino acids 1-125 of SEQ ID NO:393; and amino acids 1-120 of SEQ ID NO:397 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions.
[0411] In some embodiments, the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain or antigen binding fragment thereof comprises a heavy chain variable (VH) region and the light chain or antigen binding fragment thereof comprises a light chain variable (VL) region; wherein the anti-HIV antibody is selected from the group consisting of an antibody: [0412] i) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:1 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:2 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0413] ii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:3 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:4 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0414] iii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:5 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:6 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0415] iv) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:7 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:8 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0416] v) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:9 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:10 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0417] vi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:11 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:12 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0418] vii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:13 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:14 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0419] viii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:15 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:16 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0420] ix) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:17 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:18 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0421] x) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:19 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:20 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0422] xi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:21 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:22 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0423] xii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:23 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:24 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0424] xiii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:25 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:26 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0425] xiv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:27 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:28 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0426] xv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:29 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:30 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0427] xvi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:31 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:32 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0428] xvii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:33 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:34 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0429] xviii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:35 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:36 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0430] xix) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:37 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:38 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0431] xx) wherein the VH region comprises amino acids 1-123 of SEQ ID NO:39 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:40 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0432] xxi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:41 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:42 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0433] xxii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:43 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:44 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0434] xxiii) wherein the VH region comprises amino acids 1-125 of SEQ ID NO:45 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:46 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0435] xxiv) wherein the VH region comprises amino acids 1-125 of SEQ ID NO:47 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:48 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0436] xxv) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:49 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:50 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0437] xxvi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:51 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:52 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0438] xxvii) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:53 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:54 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0439] xxviii) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:55 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:56 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0440] xxix) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:57 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:58 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0441] xxx) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:59 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:60 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0442] xxxi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:61 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-98 of SEQ ID NO:62 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0443] xxxii) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:63 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:64 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0444] xxxiii) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:65 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:66 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0445] xxxiv) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:67 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:68 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0446] xxxv) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:69 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:70 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0447] xxxvi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:71 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:72 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0448] xxxvii) wherein the VH region comprises amino acids 1-125 of SEQ ID NO:73 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:74 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0449] xxxviii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:75 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:76 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0450] xxxix) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:153 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:155 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0451] xl) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:157 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:159 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0452] xli) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:161 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:163 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0453] xlii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:165 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:167 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0454] xliii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:169 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:171 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0455] xliv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:173 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:175 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0456] xlv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:177 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:179 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0457] xlvi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:181 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:183 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0458] xlvii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:185 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:187 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions;
[0459] xlviii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:189 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:191 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0460] xlix) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:193 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:195 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0461] l) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:197 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:199 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0462] li) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:201 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:203 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0463] lii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:205 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:207 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0464] liii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:209 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:211 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0465] liv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:213 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:215 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VH region comprises amino acids 1-127 of SEQ ID NO:217 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:219 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0466] lv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:221 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:223 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0467] lvi) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:225 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:227 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0468] lvii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:229 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:231 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0469] lviii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:233 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:235 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0470] lix) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:237 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:239 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0471] lx) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:241 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:243 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0472] lxi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:245 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:247 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0473] lxii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:249 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:251 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0474] lxiii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:253 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:255 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0475] lxiv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:257 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:259 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0476] lxv) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:261 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:263 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0477] lxvi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:265 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:267 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0478] lxvii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:269 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:271 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0479] lxviii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:273 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:275 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0480] lxix) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:277 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:279 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0481] lxx) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:281 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:283 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0482] lxxi) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:285 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:287 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0483] lxxii) wherein the VH region comprises amino acids 1-127 of SEQ ID NO:289 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:291 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0484] lxxiii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:293 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:295 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0485] lxxiv) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:297 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:299 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0486] lxxv) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:301 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:303 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0487] lxxvi) wherein the VH region comprises amino acids 1-123 of SEQ ID NO:305 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:307 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0488] lxxvii) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:309 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:311 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0489] lxxviii) wherein the VH region comprises amino acids 1-128 of SEQ ID NO:313 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:315 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0490] lxxix) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:317 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:319 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0491] lxxx) wherein the VH region comprises amino acids 1-123 of SEQ ID NO:321 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:323 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0492] lxxxi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:325 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:327 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0493] lxxxii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:329 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:331 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0494] lxxxiii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:333 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:335 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0495] lxxxiv) wherein the VH region comprises amino acids 1-125 of SEQ ID NO:337 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:339 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0496] lxxxv) wherein the VH region comprises amino acids 1-125 of SEQ ID NO:341 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:343 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0497] lxxxvi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:345 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:347 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0498] lxxxvii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:349 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:351 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0499] lxxxviii) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:353 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:355 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0500] lxxxix) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:357 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:359 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0501] xc) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:361 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:363 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0502] xci) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:365 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:367 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0503] xcii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:369 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-98 of SEQ ID NO:371 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0504] xciii) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:373 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:375 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0505] xciv) wherein the VH region comprises amino acids 1-121 of SEQ ID NO:377 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:379 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions;
[0506] xcv) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:381 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:383 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0507] xcvi) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:385 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:387 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0508] xcvii) wherein the VH region comprises amino acids 1-120 of SEQ ID NO:389 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:391 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0509] xcviii) wherein the VH region comprises amino acids 1-125 of SEQ ID NO:393 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:395 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0510] xcix) wherein the VH region comprises and amino acids 1-120 of SEQ ID NO:397 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; and wherein the VL region comprises amino acids 1-97 of SEQ ID NO:399 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions.
[0511] In some embodiments, the anti-HIV antibody is selected from the group consisting of: [0512] i) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYDFIDYV (SEQ ID NO:401), MNPSGGGT (SEQ ID NO:402) and VRDRANGSGRRRFESVNWFLDL (SEQ ID NO:403); [0513] ii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFN and WAFEN (SEQ ID NO:404); [0514] iii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYRFPDYI (SEQ ID NO:497), MNPMGGQV (SEQ ID NO:421) and VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); [0515] iv) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0516] v) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYKFPDYI (SEQ ID NO:405), INPMGGQV (SEQ ID NO:406) and VRDRSNGSGRRFESSN (SEQ ID NO:407); [0517] vi) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYL (SEQ ID NO:409), MNPVYGQV (SEQ ID NO:410) and VRDTGDGSRRHFDSINWFLDL (SEQ ID NO:411); [0518] vii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFD and WAFEA (SEQ ID NO:412); [0519] viii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), IDPPYGQV (SEQ ID NO:414) and VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); [0520] ix) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), INPGYGQV (SEQ ID NO:431) and VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); [0521] x) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFVDYF (SEQ ID NO:416), MDPLNGRP (SEQ ID NO:417) and VRDKSNGSGRRFDSSNWFLDL (SEQ ID NO:418); [0522] xi) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFN and WAYDA (SEQ ID NO:419); [0523] xii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFSDYI (SEQ ID NO:420), MNPMGGQV (SEQ ID NO:421) and VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); [0524] xiii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEV (SEQ ID NO:422); [0525] xiv) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFIDYI (SEQ ID NO:423), IDPMNGRP (SEQ ID NO:424) and VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); [0526] xv) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYI (SEQ ID NO:426), MNPMGGRT (SEQ ID NO:427) and VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); [0527] xvi) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFVDYL (SEQ ID NO:428), MDPMNGRP (SEQ ID NO:429) and VRDKSGGSGKLFDSSNWFLDL (SEQ ID NO:430); [0528] xvii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), MDPSYGQV (SEQ ID NO:432) and VRDRSHGSGRQFESSNWFLDL (SEQ ID NO:433); [0529] xviii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), MDPSFGQM (SEQ ID NO:434) and VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); [0530] xix) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYRFTDYV (SEQ ID NO:436), MDPSFGRM (SEQ ID NO:437) and VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); [0531] xx) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFIDYV (SEQ ID NO:438), MDPTYGRM (SEQ ID NO:439) and VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); [0532] xxi) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYRFLDYI (SEQ ID NO:440), MNPMGGQV (SEQ ID NO:421) and VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); [0533] xxii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYKFMDQL (SEQ ID NO:442), MNPTYGQV (SEQ ID NO:443) and ARGPSGENYPFHY (SEQ ID NO:444); [0534] xxiii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences RHII (SEQ ID NO:445), DDD and NTYEF (SEQ ID NO:446); [0535] xxiv) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFVDSR (SEQ ID NO:447), INPLQGGV (SEQ ID NO:448) and ARGIDGKSYPFHF (SEQ ID NO:449); [0536] xxv) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences S, ESS and SILEF (SEQ ID NO:450); [0537] xxvi) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTTHHGHF (SEQ ID NO:500), MNPMTGQM (SEQ ID NO:462) and ARGDFGQNYPFHY (SEQ ID NO:463); [0538] xxvii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences NRYL (SEQ ID NO:464), DDN and ASYER (SEQ ID NO:465); [0539] xxviii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFMDQF (SEQ ID NO:466), MNPIYGQV (SEQ ID NO:467) and ARGPSGENYPFHY (SEQ ID NO:444); [0540] xxix) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences RHII (SEQ ID NO:445), DDD and NTYEF (SEQ ID NO:446); [0541] xxx) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFVDSR (SEQ ID NO:447), INPLHGGV (SEQ ID NO:468) and ARGIDGKSYPFHF (SEQ ID NO:449); [0542] xxxi) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTKYF (SEQ ID NO:451), IHPRTGAV (SEQ ID NO:452) and ARGAFEADSYGSSYPFHH (SEQ ID NO:453); [0543] xxxii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences GNYNP (SEQ ID NO:454), EDN and ASFEF (SEQ ID NO:455); [0544] xxxiii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTKYT (SEQ ID NO:456), IHPRTGAV (SEQ ID NO:452) and ARGAFEADLSGPTYPFHH (SEQ ID NO:457); [0545] xxxiv) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFNFIDSV (SEQ ID NO:458), IKPLRGAV (SEQ ID NO:459) and AKGAFRGGSPFGF (SEQ ID NO:460); [0546] xxxv) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences DVT and ASREF (SEQ ID NO:461); [0547] xxxvi) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTSYF (SEQ ID NO:469), INPLHGAV (SEQ ID NO:470) and TRGIVADGWPYGH (SEQ ID NO:471); [0548] xxxvii) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences S, EGA and SSLQF (SEQ ID NO:472); [0549] xxxviii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFTFIDHI (SEQ ID NO:473), IKPLRGAV (SEQ ID NO:459) and CKAAAPEEAFPLQY (SEQ ID NO:474); [0550] xxxix) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSRTF (SEQ ID NO:475); [0551] xl) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFIDHI (SEQ ID NO:476), IKPLGGVA (SEQ ID NO:477) and CKAAAPDEAFPLEY (SEQ ID NO:478); [0552] xli) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSTTF (SEQ ID NO:479); [0553] xlii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFAFLDH (SEQ ID NO:480), VKTIGGVV (SEQ ID NO:481) and SKAAAPDEAFPLEF (SEQ ID NO:482); [0554] xliii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFAFLDHI (SEQ ID NO:486), VKTIGGVV (SEQ ID NO:481) and SKAAAPDEAFPLEF (SEQ ID NO:482); [0555] xliv) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFTEYF (SEQ ID NO:483), LNPLRGAV (SEQ ID NO:484), ARAVFNEAFPFDY (SEQ ID NO:485); [0556] xlv) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences VS, DGD and ASREF (SEQ ID NO:461); [0557] xlvi) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DND and SSTTF (SEQ ID NO:479); [0558] xlvii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFNFIDSV (SEQ ID NO:458), IKPLRGGV (SEQ ID NO:490) and AKGAFGGSSPFGF (SEQ ID NO:491); [0559] xlviii) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFIDSV (SEQ ID NO:487), IKPLGGAV (SEQ ID NO:488) and AKGAFGGGSPFGF (SEQ ID NO:489); [0560] xlix) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFTFIKYT (SEQ ID NO:492), IHPRTGAV (SEQ ID NO:452) and ARGAFEADLYGPTYPFHH (SEQ ID NO:493); [0561] l) an antibody comprising a light chain variable region, wherein the CDRs comprise amino acid sequences GSYNP (SEQ ID NO:494), DDN and ASFEF (SEQ ID NO:455); and [0562] li) an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFVDSL (SEQ ID NO:495), INPLQGGV (SEQ ID NO:448) and ARGIDGNSYPFHF (SEQ ID NO:496).
[0563] In some embodiments, the anti-HIV antibody is selected from the group consisting of: [0564] a. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYDFIDYV (SEQ ID NO:401), MNPSGGGT (SEQ ID NO:402) and VRDRANGSGRRRFESVNWFLDL (SEQ ID NO:403); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFN and WAFEN (SEQ ID NO:404); [0565] b. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYRFPDYI (SEQ ID NO:497), MNPMGGQV (SEQ ID NO:421) and VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0566] c. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYKFPDYI (SEQ ID NO:405), INPMGGQV (SEQ ID NO:406) and VRDRSNGSGRRFESSN (SEQ ID NO:407); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0567] d. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYL (SEQ ID NO:409), MNPVYGQV (SEQ ID NO:410) and VRDTGDGSRRHFDSINWFLDL (SEQ ID NO:411); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFD and WAFEA (SEQ ID NO:412); [0568] e. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), IDPPYGQV (SEQ ID NO:414) and VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0569] f. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), INPGYGQV (SEQ ID NO:431) and VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0570] g. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFVDYF (SEQ ID NO:416), MDPLNGRP (SEQ ID NO:417) and VRDKSNGSGRRFDSSNWFLDL (SEQ ID NO:418); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFN and WAYDA (SEQ ID NO:419); [0571] h. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFSDYI (SEQ ID NO:420), MNPMGGQV (SEQ ID NO:421) and VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEV (SEQ ID NO:422); [0572] i. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFIDYI (SEQ ID NO:423), IDPMNGRP (SEQ ID NO:424) and VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFN and WAYDA (SEQ ID NO:419); [0573] j. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYI (SEQ ID NO:426), MNPMGGRT (SEQ ID NO:427) and VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0574] k. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFVDYL (SEQ ID NO:428), MDPMNGRP (SEQ ID NO:429) and VRDKSGGSGKLFDSSNWFLDL (SEQ ID NO:430); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNY, DFN and WAYDA (SEQ ID NO:419); [0575] l. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), INPGYGQV (SEQ ID NO:431) and VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0576] m. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), MDPSYGQV (SEQ ID NO:432) and VRDRSHGSGRQFESSNWFLDL (SEQ ID NO:433); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0577] n. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTDYV (SEQ ID NO:413), MDPSFGQM (SEQ ID NO:434) and VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0578] o. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYRFTDYV (SEQ ID NO:436), MDPSFGRM (SEQ ID NO:437) and VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0579] p. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFIDYV (SEQ ID NO:438), MDPTYGRM (SEQ ID NO:439) and VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0580] q. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYRFLDYI (SEQ ID NO:440), MNPMGGQV (SEQ ID NO:421) and VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein the CDRs comprise amino acid sequences HNL, DFN and WAYEA (SEQ ID NO:408); [0581] r. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYKFMDQL (SEQ ID NO:442), MNPTYGQV (SEQ ID NO:443) and ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein the CDRs comprise amino acid sequences RHII (SEQ ID NO:445), DDD and NTYEF (SEQ ID NO:446); [0582] s. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFVDSR (SEQ ID NO:447), INPLQGGV (SEQ ID NO:448) and ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein the CDRs comprise amino acid sequences S, ESS and SILEF (SEQ ID NO:450); [0583] t. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTTHHGHF (SEQ ID NO:500), MNPMTGQM (SEQ ID NO:462) and ARGDFGQNYPFHY (SEQ ID NO:463); and a light chain variable region, wherein the CDRs comprise amino acid sequences NRYL (SEQ ID NO:464), DDN and ASYER (SEQ ID NO:465); [0584] u. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFMDQF (SEQ ID NO:466), MNPIYGQV (SEQ ID NO:467) and ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein the CDRs comprise amino acid sequences RHII (SEQ ID NO:445), DDD and NTYEF (SEQ ID NO:446); [0585] v. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFVDSR (SEQ ID NO:447), INPLHGGV (SEQ ID NO:468) and ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein the CDRs comprise amino acid sequences S, ESS and SILEF (SEQ ID NO:450); [0586] w. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTKYF (SEQ ID NO:451), IHPRTGAV (SEQ ID NO:452) and ARGAFEADSYGSSYPFHH (SEQ ID NO:453); and a light chain variable region, wherein the CDRs comprise amino acid sequences GNYNP (SEQ ID NO:454), EDN and ASFEF (SEQ ID NO:455); [0587] x. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTKYT (SEQ ID NO:456), IHPRTGAV (SEQ ID NO:452) and ARGAFEADLSGPTYPFHH (SEQ ID NO:457); and a light chain variable region, wherein the CDRs comprise amino acid sequences GNYNP (SEQ ID NO:454), EDN and ASFEF (SEQ ID NO:455); [0588] y. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFNFIDSV (SEQ ID NO:458), IKPLRGAV (SEQ ID NO:459) and AKGAFRGGSPFGF (SEQ ID NO:460); and a light chain variable region, wherein the CDRs comprise amino acid sequences DVT and ASREF (SEQ ID NO:461); [0589] z. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYTFTSYF (SEQ ID NO:469), INPLHGAV (SEQ ID NO:470) and TRGIVADGWPYGH (SEQ ID NO:471); and a light chain variable region, wherein the CDRs comprise amino acid sequences S, EGA and SSLQF (SEQ ID NO:472); [0590] aa. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFTFIDHI (SEQ ID NO:473), IKPLRGAV (SEQ ID NO:459) and CKAAAPEEAFPLQY (SEQ ID NO:474); and a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSRTF (SEQ ID NO:475); [0591] bb. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFIDHI (SEQ ID NO:476), IKPLGGVA (SEQ ID NO:477) and CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSTTF (SEQ ID NO:479); [0592] cc. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFAFLDH (SEQ ID NO:480), VKTIGGVV (SEQ ID NO:481) and SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSTTF (SEQ ID NO:479); [0593] dd. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFAFLDHI (SEQ ID NO:486), VKTIGGVV (SEQ ID NO:481) and SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSTTF (SEQ ID NO:479); [0594] ee. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFTEYF (SEQ ID NO:483), LNPLRGAV (SEQ ID NO:484), ARAVFNEAFPFDY (SEQ ID NO:485); and a light chain variable region, wherein the CDRs comprise amino acid sequences VS, DGD and ASREF (SEQ ID NO:461); [0595] ff. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFIDHI (SEQ ID NO:476), IKPLGGVA (SEQ ID NO:477) and CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DND and SSTTF (SEQ ID NO:479); [0596] gg. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFAFLDHI (SEQ ID NO:486), VKTIGGVV (SEQ ID NO:481) and SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein the CDRs comprise amino acid sequences NVD, DNN and SSTTF (SEQ ID NO:479); [0597] hh. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFNFIDSV (SEQ ID NO:458), IKPLRGGV (SEQ ID NO:490) and AKGAFGGSSPFGF (SEQ ID NO:491); and a light chain variable region, wherein the CDRs comprise amino acid sequences DVT and ASREF (SEQ ID NO:461); [0598] ii. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFKFIDSV (SEQ ID NO:487), IKPLGGAV (SEQ ID NO:488) and AKGAFGGGSPFGF (SEQ ID NO:489); and a light chain variable region, wherein the CDRs comprise amino acid sequences DVT and ASREF (SEQ ID NO:461); [0599] jj. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFNFIDSV (SEQ ID NO:458), IKPLRGGV (SEQ ID NO:490) and AKGAFGGSSPFGF (SEQ ID NO:491); and a light chain variable region, wherein the CDRs comprise amino acid sequences DVT and ASREF (SEQ ID NO:461); [0600] kk. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GFTFIKYT (SEQ ID NO:492), IHPRTGAV (SEQ ID NO:452) and ARGAFEADLYGPTYPFHH (SEQ ID NO:493); and a light chain variable region, wherein the CDRs comprise amino acid sequences GSYNP (SEQ ID NO:494), DDN and ASFEF (SEQ ID NO:455); [0601] ll. an antibody comprising a heavy chain variable region, wherein the CDRs comprise amino acid sequences GYNFVDSL (SEQ ID NO:495), INPLQGGV (SEQ ID NO:448) and ARGIDGNSYPFHF (SEQ ID NO:496); and a light chain variable region, wherein the CDRs comprise amino acid sequences S, ESS and SILEF (SEQ ID NO:450).
[0602] In some embodiments, the anti-HIV antibody is selected from the group consisting of: [0603] a. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYDFIDYV (SEQ ID NO:401), CDR H2 comprises MNPSGGGT (SEQ ID NO:402) and CDR H3 comprises VRDRANGSGRRRFESVNWFLDL (SEQ ID NO:403); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); [0604] b. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFPDYI (SEQ ID NO:405), CDR H2 comprises INPMGGQV (SEQ ID NO:406) and CDR H3 comprises VRDRSNGSGRRFESSN (SEQ ID NO:407); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0605] c. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYL (SEQ ID NO:409), CDR H2 comprises MNPVYGQV (SEQ ID NO:410) and CDR H3 comprises VRDTGDGSRRHFDSINWFLDL (SEQ ID NO:411); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412); [0606] d. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises IDPPYGQV (SEQ ID NO:414) and CDR H3 comprises VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0607] e. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFVDYF (SEQ ID NO:416), CDR H2 comprises MDPLNGRP (SEQ ID NO:417) and CDR H3 comprises VRDKSNGSGRRFDSSNWFLDL (SEQ ID NO:418); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0608] f. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFSDYI (SEQ ID NO:420), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEV (SEQ ID NO:422); [0609] g. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFIDYI (SEQ ID NO:423), CDR H2 comprises IDPMNGRP (SEQ ID NO:424) and CDR H3 comprises VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0610] h. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYI (SEQ ID NO:426), CDR H2 comprises MNPMGGRT (SEQ ID NO:427) and CDR H3 comprises VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0611] i. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFVDYL (SEQ ID NO:428), CDR H2 comprises MDPMNGRP (SEQ ID NO:429) and CDR H3 comprises VRDKSGGSGKLFDSSNWFLDL (SEQ ID NO:430); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0612] j. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises INPGYGQV (SEQ ID NO:431) and CDR H3 comprises VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0613] k. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises MDPSYGQV (SEQ ID NO:432) and CDR H3 comprises VRDRSHGSGRQFESSNWFLDL (SEQ ID NO:433); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0614] l. an antibody comprising a heavy chain variable region, wherein CDR comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises MDPSFGQM (SEQ ID NO:434) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0615] m. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFTDYV (SEQ ID NO:436), CDR H2 comprises MDPSFGRM (SEQ ID NO:437) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0616] n. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFIDYV (SEQ ID NO:438), CDR H2 comprises MDPTYGRM (SEQ ID NO:439) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0617] o. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFLDYI (SEQ ID NO:440), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0618] p. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0619] q. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSR (SEQ ID NO:447), CDR H2 comprises INPLQGGV (SEQ ID NO:448) and CDR H3 comprises ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0620] r. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTTHHGHF (SEQ ID NO:500), CDR H2 comprises MNPMTGQM (SEQ ID NO:462) and CDR H3 comprises ARGDFGQNYPFHY (SEQ ID NO:463); and a light chain variable region, wherein CDR L1 comprises NRYL (SEQ ID NO:464), CDR L2 comprises DDN and CDR L3 comprises ASYER (SEQ ID NO:465); [0621] s. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFMDQF (SEQ ID NO:466), CDR H2 comprises MNPIYGQV (SEQ ID NO:467) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0622] t. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSR (SEQ ID NO:447), CDR H2 comprises INPLHGGV (SEQ ID NO:468) and CDR H3 comprises ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0623] u. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTKYF (SEQ ID NO:451), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADSYGSSYPFHH (SEQ ID NO:453); and a light chain variable region, wherein CDR L1 comprises GNYNP (SEQ ID NO:454), CDR L2 comprises EDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0624] v. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTKYT (SEQ ID NO:456), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADLSGPTYPFHH (SEQ ID NO:457); and a light chain variable region, wherein CDR L1 comprises GNYNP (SEQ ID NO:454), CDR L2 comprises EDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0625] w. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFNFIDSV (SEQ ID NO:458), CDR H2 comprises IKPLRGAV (SEQ ID NO:459) and CDR H3 comprises AKGAFRGGSPFGF (SEQ ID NO:460); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0626] x. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTSYF (SEQ ID NO:469), CDR H2 comprises INPLHGAV (SEQ ID NO:470) and CDR H3 comprises TRGIVADGWPYGH (SEQ ID NO:471); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises EGA and CDR L3 comprises SSLQF (SEQ ID NO:472); [0627] y. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFTFIDHI (SEQ ID NO:473), CDR H2 comprises IKPLRGAV (SEQ ID NO:459) and CDR H3 comprises CKAAAPEEAFPLQY (SEQ ID NO:474); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSRTF (SEQ ID NO:475); [0628] z. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDHI (SEQ ID NO:476), CDR H2 comprises IKPLGGVA (SEQ ID NO:477) and CDR H3 comprises CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0629] aa. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFAFLDH (SEQ ID NO:480), CDR H2 comprises VKTIGGVV (SEQ ID NO:481) and CDR H3 comprises SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0630] bb. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFTEYF (SEQ ID NO:483), CDR H2 comprises LNPLRGAV (SEQ ID NO:484) and CDR H3 comprises ARAVFNEAFPFDY (SEQ ID NO:485); and a light chain variable region, wherein CDR L1 comprises VS, CDR L2 comprises DGD and CDR L3 comprises ASREF (SEQ ID NO:461); [0631] cc. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDHI (SEQ ID NO:476), CDR H2 comprises IKPLGGVA (SEQ ID NO:477) and CDR H3 comprises CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DND and CDR L3 comprises SSTTF (SEQ ID NO:479); [0632] dd. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFAFLDHI (SEQ ID NO:486), CDR H2 comprises VKTIGGVV (SEQ ID NO:481) and CDR H3 comprises SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0633] ee. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDSV (SEQ ID NO:487), CDR H2 comprises IKPLGGAV (SEQ ID NO:488) and CDR H3 comprises AKGAFGGGSPFGF (SEQ ID NO:489); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0634] ff. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFNFIDSV (SEQ ID NO:458), CDR H2 comprises IKPLRGGV (SEQ ID NO:490) and CDR H3 comprises AKGAFGGSSPFGF (SEQ ID NO:491); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0635] gg. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFTFIKYT (SEQ ID NO:492), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADLYGPTYPFHH (SEQ ID NO:493); and a light chain variable region, wherein CDR L1 comprises GSYNP (SEQ ID NO:494), CDR L2 comprises DDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0636] hh. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSL (SEQ ID NO:495), CDR H2 comprises INPLQGGV (SEQ ID NO:448) and CDR H3 comprises ARGIDGNSYPFHF (SEQ ID NO:496); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0637] ii. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0638] jj. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); [0639] kk. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412); [0640] ll. an antibody comprising a heavy chain variable region, wherein CDR comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0641] mm. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0642] nn. an antibody comprising a heavy chain variable region, wherein CDR comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0643] oo. an antibody comprising a heavy chain variable region, wherein CDR comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSN (SEQ ID NO:498); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0644] pp. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); and [0645] qq. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFLDYI (SEQ ID NO:440), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408).
[0646] In some embodiments, the anti-HIV antibody is a non-naturally occurring antibody. In some embodiments, the anti-HIV antibody is selected from the group consisting of: N49P6; N49P6.2; N49P7; N49P7.1; N49P7A; N49P7S; N49P7F; N49P7Y; N49P7-54TY; N49P7LS-1; N49P7LS-2; N49P7YTE; N49P7L6; N49P7L11; N49P7.1L9; N49P7.1L19 R49P7; N49P7.2; N49P11; N49P18; N49P18.2; N49P18.1; N49P19; N49P37; N49P38; N49P38.1; N49P55; N49P56; N49P57; N49P58; N49P59; N49P73; N49P74; N49P75; N49P75.1; N49P9; N49P9.1; N49P9.2; N49P9i7; N49P9i7H1; N49P9i7H2; N49P22; N49P23; N49P9.3; N49P9.4; N49P51; N49P52; N49P53; N49P54; N49P60; N49P61; N49P62; N49P63; N49P64; N49P65; N49P66; N49P67; N49P68; N49P69; N49P70; N49P71; and N49P72.
[0647] In some embodiments, the invention provides antibodies or antigen binding fragments comprise the CDRs as shown in the Table 2 below with up to four (i.e. 0, 1, 2, 3, or 4) conservative amino acid substitutions per CDR.
TABLE-US-00002 TABLE 2 Amino acid sequence of natural antibodies. Natural antibody Related variants Amino Acid sequence Sequence 1 N49P6, N49P6.2 ##STR00001## ##STR00002## YYCVRDRANGSGRRRFESVNWFLDLWGRGTQITVVSPSTKGPSVFPL APSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSS GLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC PPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGF YPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG NVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 1 --------------------------------------------------- ##STR00003## ##STR00004## LRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTV EKTVAPTECS SEQ ID NO: 2 Sequence 2 N49P7, N49P7.1 ##STR00005## N49P7A ##STR00006## N49P7S CVRDRSNGSGKRFESSNWFLDLWGRGTAVTIQSSSTKGPSVFPLAPSS N49P7F KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY N49P7Y SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC N49P7.54TY PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY N49P7-LS1 VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS N49P7-L52 NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP N49P7/6L SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV N49P7/11L FSCSVMHEALHNHYTQKSLSLSPGK R49P7 SEQ ID NO: 3 --------------------------------------------------- ##STR00007## ##STR00008## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 4 Sequence 3 N49P7.2 ##STR00009## ##STR00010## CVRDRSNGSGRRFESSNWFLDLWGRGTAVTVHSPSKSTSGGTAALGC LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPSVFL FPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTK PREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISK AKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNH YTQKSLSLSPGK SEQ ID NO: 5 --------------------------------------------------- ##STR00011## ##STR00012## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 6 Sequence 4 N49P11 ##STR00013## ##STR00014## YYCVRDTGDGSRRHFDSINWFLDLWGRGTWIRVAPASTKGPSVFPLA PSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSG LYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP PCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGF YPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG NVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 7 --------------------------------------------------- ##STR00015## ##STR00016## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 8 Sequence 5 N49P18 ##STR00017## N49P18.2 ##STR00018## VRDRSNGWGKRFESSNWFLDLWGRGTVVTVHSPSTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 9 --------------------------------------------------- ##STR00019## ##STR00020## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 10 Sequence 6 N49P18.1 ##STR00021## ##STR00022## VRDRSNGWGKRFESSNWFLDLWGRGTVVTVHSPSTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 11 --------------------------------------------------- ##STR00023## ##STR00024## LRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTV EKTVAPTECS SEQ ID NO: 12 Sequence 7 N49P19 ##STR00025## ##STR00026## CVRDKSNGSGRRFDSSNWFLDLWGRGTRVSIFSASTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 13 --------------------------------------------------- ##STR00027## ##STR00028## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 14 Sequence 8 N49P37 ##STR00029## ##STR00030## YCVRDRSNGSGKRFESSNWFLDLWGRGTAVTISSPSTKGPSVFPLAPS SKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGL YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPP CPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNW YVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKV SNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFY PSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGN VFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 15 --------------------------------------------------- ##STR00031## ##STR00032## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 16 Sequence 9 N49P38 ##STR00033## N49P38.1 ##STR00034## RDKSNGSGKRFDSSNWFLDLWGRGTRVSISSASTKGPSVFPLAPSSKS TSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAP ELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKAL PAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCS VMHEALHNHYTQKSLSLSPGK SEQ ID NO: 17 --------------------------------------------------- ##STR00035## ##STR00036## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 18 Sequence 10 N49P55 ##STR00037## ##STR00038## CVRDKSNGSGKRFDSSNWFLDLWGRGTPVTISSPSTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 19 --------------------------------------------------- ##STR00039## ##STR00040## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 20 Sequence 11 N49P56 ##STR00041## ##STR00042## CVRDKSGGSGKLFDSSNWFLDLWGRGTRVSISSASTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 21 --------------------------------------------------- ##STR00043## ##STR00044## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 22 Sequence 12 N49P57 ##STR00045## ##STR00046## VRDRSNGWGKRFESSNWFLDLWGRGTVITVHSPSTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 23 --------------------------------------------------- ##STR00047## ##STR00048## LRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTV EKTVAPTECS SEQ ID NO: 24 Sequence 13 N49P58 ##STR00049## ##STR00050## CVRDRSHGSGRQFESSNWFLDLWGRGTVVNVQSPSTKGPSVFPLAPS SKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGL YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPP CPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNW YVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKV SNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFY PSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGN VFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 25 --------------------------------------------------- ##STR00051## ##STR00052## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS
SEQ ID NO: 26 Sequence 14 N49P59 ##STR00053## ##STR00054## CVRDRSHGSGRLFESSNWFLDLWGRGTVVTVQSPSTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 27 --------------------------------------------------- ##STR00055## ##STR00056## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 28 Sequence 15 N49P73 ##STR00057## ##STR00058## CVRDRSHGSGRLFESSNWFLDLWGRGTVVTVQSPSTKGPSVFPLAPSS KSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLY SLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYP SDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 29 --------------------------------------------------- ##STR00059## ##STR00060## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 30 Sequence 16 N49P74 ##STR00061## ##STR00062## VRDRSHGSGRLFESSNWFLDLWGRGTVVTVQSPSTKGPSVFPLAPSSK STSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSL SSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPA PELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVD GVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSD IAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFS CSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 31 --------------------------------------------------- ##STR00063## ##STR00064## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 32 Sequence 17 N49P75 ##STR00065## ##STR00066## YCVRDRSNGSGKRFESSNWFLDLWGRGTAVTIHSPSTKGPSVFPLAPS SKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGL YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPP CPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNW YVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKV SNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFY PSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGN VFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 33 --------------------------------------------------- ##STR00067## ##STR00068## VLRQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSS PVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGST VEKTVAPTECS SEQ ID NO: 34 Sequence 18 N49P9 ##STR00069## N49P9.1 ##STR00070## N49P9.2 AVYYCARGPSGENYPFHYWGQGVRVVVSSPSTKGPSVFPLAPSSKSTS N49P9i7 GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPE LLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKAL PAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCS VMHEALHNHYTQKSLSLSPGK SEQ ID NO: 35 --------------------------------------------------- ##STR00071## ##STR00072## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 36 Sequence 19 N49P22 ##STR00073## ##STR00074## ARGIDGKSYPFHFWGHGTRVTVFSASTKGPSVFPLAPSSKSTSGGTAA LGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPS SSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNA KTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKT ISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEAL HNHYTQKSLSLSPGK SEQ ID NO: 37 --------------------------------------------------- ##STR00075## ##STR00076## APSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVGV ETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 38 Sequence 20 N49P23 ##STR00077## ##STR00078## GDTGTYYCARGDFGQNYPFHYWGQGSLVIVSSASTKGPSVFPLAPSSK STSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSL SSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPA PELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVD GVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSD IAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFS CSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 39 --------------------------------------------------- ##STR00079## ##STR00080## VLSQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADG SPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGS TVEKTVAPAECS SEQ ID NO: 40 Sequence 21 N49P9.3 ##STR00081## N49P9.4 ##STR00082## VYYCARGPSGENYPFHYWGQGVRVVVSSPSTKGPSVFPLAPSSKSTSG GTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSV VTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEL LGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGV EVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK SEQ ID NO: 41 --------------------------------------------------- ##STR00083## ##STR00084## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 42 Sequence 22 N49P51 ##STR00085## ##STR00086## ARGIDGKSYPFHFWGHGTRVTVFSASTKGPSVFPLAPSSKSTSGGTAA LGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPS SSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNA KTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKT ISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEAL HNHYTQKSLSLSPGK SEQ ID NO: 43 --------------------------------------------------- ##STR00087## ##STR00088## APSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVGV ETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 44 Sequence 23 N49P52 ##STR00089## ##STR00090## CARGAFEADSYGSSYPFHHWGQGTLVTVSSASTKGPSVFPLAPSSKST SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPE LLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKAL PAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCS VMHEALHNHYTQKSLSLSPGK SEQ ID NO: 45 --------------------------------------------------- ##STR00091## ##STR00092## LTVLSQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKA DGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHE GSTVEKTVAPAECS SEQ ID NO: 46 Sequence 24 N49P53 ##STR00093## ##STR00094## CARGAFEADLSGPTYPFHHWGQGTLVIVSAASTKGPSVFPLAPSSKST SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPE LLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKAL PAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCS VMHEALHNHYTQKSLSLSPGK SEQ ID NO: 47 --------------------------------------------------- ##STR00095## ##STR00096## LIVLSQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKAD GSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEG STVEKTVAPAECS SEQ ID NO: 48 Sequence 25 N49P54 ##STR00097## ##STR00098## CAKGAFRGGSPFGFWGQGTLLTVSPASTKGPSVFPLAPSSKSTSGGTA ALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGP SVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEK TISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK SEQ ID NO: 49 --------------------------------------------------- ##STR00099## ##STR00100## AAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVG VETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTV APAECS SEQ ID NO: 50 Sequence 26 N49P60 ##STR00101## ##STR00102## FCTRGIVADGWPYGHWGQGTQVTVSPASTKGPSVFPLAPSSKSTSGGT AALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVT VPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLG GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEV HNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAP IEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEW ESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMH EALHNHYTQKSLSLSPGK SEQ ID NO: 51 --------------------------------------------------- ##STR00103## ##STR00104## PKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVK VGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEK TVAPAECS SEQ ID NO: 52 Sequence 27 N49P61 ##STR00105## ##STR00106##
AAAPEEAFPLQYWGQGTQLIVSSASTKGPSVFPLAPSSKSTSGGTAALG CLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPSVF LFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKT KPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTIS KAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNG QPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK SEQ ID NO: 53 --------------------------------------------------- ##STR00107## ##STR00108## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 54 Sequence 28 N49P62 ##STR00109## ##STR00110## CKAAAPDEAFPLEYWGQGTQLIVSPASTKGPSVFPLAPSSKSTSGGTAA LGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPS SSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNA KTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKT ISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEAL HNHYTQKSLSLSPGK SEQ ID NO: 55 --------------------------------------------------- ##STR00111## ##STR00112## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 56 Sequence 29 N49P63 ##STR00113## ##STR00114## CSKAAAPDEAFPLEFWGQGTQVIVSSASTKGPSVFPLAPSSKSTSGGTA ALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGP SVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEK TISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK SEQ ID NO: 57 --------------------------------------------------- ##STR00115## ##STR00116## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 58 Sequence 30 N49P64 ##STR00117## ##STR00118## SKAAAPDEAFPLEFWGQGTQVIVSSASTKGPSVFPLAPSSKSTSGGTAA LGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPS SSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNA KTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKT ISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEAL HNHYTQKSLSLSPGK SEQ ID NO: 59 --------------------------------------------------- ##STR00119## ##STR00120## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 60 Sequence 31 N49P65 ##STR00121## ##STR00122## YYCARAVFNEAFPFDYWGQGSLVTVSSASTKGPSVFPLAPSSKSTSGG TAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVT VPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLG GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEV HNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAP IEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEW ESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMH EALHNHYTQKSLSLSPGK SEQ ID NO: 61 --------------------------------------------------- ##STR00123## ##STR00124## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 62 Sequence 32 N49P66 ##STR00125## ##STR00126## CKAAAPDEAFPLEYWGQGTQLIVSPASTKGPSVFPLAPSSKSTSGGTAA LGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPS SSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNA KTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKT ISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEAL HNHYTQKSLSLSPGK SEQ ID NO: 63 --------------------------------------------------- ##STR00127## ##STR00128## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 64 Sequence 33 N49P67 ##STR00129## ##STR00130## CSKAAAPDEAFPLEFWGQGTQVIVSSASTKGPSVFPLAPSSKSTSGGTA ALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGP SVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEK TISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK SEQ ID NO: 65 --------------------------------------------------- ##STR00131## ##STR00132## SQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSP VKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 66 Sequence 34 N49P68 ##STR00133## ##STR00134## CAKGAFGGSSPFGFWGQGTLLTVSPASTKGPSVFPLAPSSKSTSGGTA ALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGP SVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEK TISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK SEQ ID NO: 67 --------------------------------------------------- ##STR00135## ##STR00136## AAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVG VETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTV APAECS SEQ ID NO: 68 Sequence 35 N49P69 ##STR00137## ##STR00138## CAKGAFGGGSPFGFWGQGTLLTVSPASTKGPSVFPLAPSSKSTSGGTA ALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGP SVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEK TISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK SEQ ID NO: 69 --------------------------------------------------- ##STR00139## ##STR00140## AAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVG VETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTV APAECS SEQ ID NO: 70 Sequence 36 N49P70 ##STR00141## ##STR00142## CAKGAFGGSSPFGFWGQGTLLTVSPASTKGPSVFPLAPSSKSTSGGTA ALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGP SVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEK TISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK SEQ ID NO: 71 --------------------------------------------------- ##STR00143## ##STR00144## APSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVGV ETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 72 Sequence 37 N49P71 ##STR00145## ##STR00146## ARGAFEADLYGPTYPFHHWGQGTQVTVSAASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPE LLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKAL PAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCS VMHEALHNHYTQKSLSLSPGK SEQ ID NO: 73 --------------------------------------------------- ##STR00147## ##STR00148## LTVLSQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKA DGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHE GSTVEKTVAPAECS SEQ ID NO: 74 Sequence 38 N49P72 ##STR00149## ##STR00150## ARGIDGNSYPFHFWGHGTRVTVFSASTKGPSVFPLAPSSKSTSGGTAAL GCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSS SLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPSV FLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTI SKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNG QPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK SEQ ID NO: 75 --------------------------------------------------- ##STR00151## ##STR00152## APSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVGV ETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 76 Heavy chain is shown first (above dotted line), followed by the light chain (below dotted line). Sequences of CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 are in bold. Note that there is a predicted blank CDR in the light chain for Sequences 25, 34 and 36.
TABLE-US-00003 TABLE 3 Nucleotide sequences of natural antibodies. Heavy chain is shown first (above dotted line), followed by the light chain (below dotted line). Natural Related antibody variants Nucleotide sequence Sequence 1 N49P6, gcgggactgatgcagtctggggctgtgatgaagaattcgggggcctcagtgagggtctct N49P6.2 tgtcaggctgatggatacgacttcattgactatgtcattcactggtttcgacaaagacgt N49P6- ggagaaggtcttgagtggctgggatggatgaatccctcgggaggcggcacaaactatccg LS1 cgaccatttcagggcaaagtcaccatgaccagggacacgtccaccgagacagcctattta N49P6- gatgtcagaggacttacatatgacgacacggccgtctattattgtgtgagagacagggcc LS2 aacggttcgggaagaagacgttttgagtcggtgaattggttcctggatctgtggggccgc N49P6A ggcacccaaataacagtcgtctcgccctccaccaagggcccatcggtcttccccctggca N49P6S ccctcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactac N49P6F ttccccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacacc N49P6Y ttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccc N49P6.54 tccagcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacacc TY aaggtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgc ccagcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggac accctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaa gaccctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagaca aagccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctg caccaggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctccca gcccccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtac accctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtc aaaggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaac aactacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaag ctcaccgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcat gaggctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 77 ------------------------------------------------------------ cagtctgccctaactcagcctcgctcagtgtccgcatctcctggtcagtcagtcaccatc tcctgcactggaacacacaattatgtgtcctggtgtcaacagaaaccgggccaagccccc aaattattaatttacgatttcaataaacggccctcaggggtctctgatcgcttctctggc tccacgtctggcaacacggcctccctgaccatctctggactccaggctgacgatgagggt cattatttttgttgggcgtttgaaaatatcggcggagggaccaagctgaccgtcctgcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 78 Sequence 2 N49P7, gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgaggatctcc N49P7.1 tgtgaggctcaaggatatagatttcctgactacatcattcactggattcgacgggcccct N49P7A ggacaaggccctgaatggatgggatggatgaatccaatgggcggacaagtaaatattcca N49P7S tggaaatttcagggtagggtctccatgacccgggacacgtccatcgaaacagcatttctg N49P7F gacttaagaggactaaagtctgacgacacggccgtctattattgcgtgagagatcgcagt N49P7Y aatggatcgggaaagcgattcgagtcctccaattggttcctcgatctgtggggccgtggg N49P7.54 actgcggtcacaattcaatca TY tcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg N49P7- ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg LS1 tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca N49P7- ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc LS2 tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc N49P7/6L aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga N49P7/11L ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct R49P7 gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 79 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcagccaggcagagccccc aaattattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcactggactccaggatgacgatgacgcc gaatatttttgttgggcgtatgaagcttttggcggagggaccaagttgaccgttcttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 80 Sequence 3 N49P7.2 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgaggatctcc tgtgaggctcaaggatacaaatttcctgactacatcattcactggattcgacgggcccct ggacaaggccttgagtggatggggtggattaatccaatgggcggacaagtaaacattcca tggcagtttcagggcagggtctccatgacccgggacacgtccatcgaaacagcatttctg gacttaagaggactaaagtctgacgacacggccctctattattgcgtgagagatcgaagt aatggatcgggaaggcgattcgagtcctccaattggttcctcgatctgtggggccgcggc actgcggtcactgttcattcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtatctcatgctccgtgatgcatgagg ctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 81 cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagagccccc aaattattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcagtggactccaggatgacgatgacgcc gaatatttttgttgggcgtatgaagcttttggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 82 Sequence 4 N49P11 tcggcggaattggtgcaatctggggctgtggtgaagaagcctgggacctccgtgaaggtc tcttgtcaggcttatggatacacttttaccgactaccttattcattggcttcgacaggcc cctggacaaggacttgaatggatgggatggatgaatcctgtttatggacaagtaaattat gcccaaaactttcagggcagggtctccatgaccagggacatttacagggaaacagcattt ctagaggtgcgcgacctgaagactgacgacacaggcacttattattgtgtgagagacaca ggcgacggttcgcggagacactttgactccatcaattggtttctcgatctttggggccgc gggacatggataagggtcgccccagcctccaccaagggcccatcggtcttccccctggca ccctcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactac ttccccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacacc ttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccc tccagcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacacc aaggtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgc ccagcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggac accctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaa gaccctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagaca aagccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctg caccaggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctccca gcccccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtac accctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtc aaaggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaac aactacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaag ctcaccgtggacaagagcaggtggcagcaggggaacgtatctcatgctccgtgatgcatg aggctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 83 ------------------------------------------------------------ cagtgtgtcttgactcagcctcgctcagtgtccggatctcctggacaatcagtcaccatc tcctgcactggaactcacaattatgtctcctggtgtcaacaccacccaggcaacgccccc aaattattactttatgatttcgacaagcggccctcaggaatctctgatcgcttctctggc tctaggtctggcaacacggcctccctgaccatctctggcctccagcctgaggatgaggcc gattacttttgttgggcctttgaagcctttggcggagggaccaaggtgctcgtccttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 84 Sequence 5 N49P18 gcggacttggtgcagtctggggctgtgatgaagaagcctggggactcagtgagaatctcc N49P18.2 tgtgaggctcgaggatacacattcactgactacgtcattcactggattcgacgggcccct ggacaaggccttgaatggatggggtggattgatccaccttatggacaagtaaatattcca tggaattttcagggcagggtctccatgacccgggacacgtccatcgaaacagcatttctg gacttaagaggtctaaagtctgacgacacgggcctctattattgcgtgagagatcgaagt aatggatggggaaagcgattcgagtcctccaattggttcctcgatctgtggggccgcggc actgtggtcactgttcactcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 85 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagagccccc aagttattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctaaccatcagtggactccaggatgacgatgacgcc gaatatttttgttgggcatatgaagctttcggcggagggaccaagttgactgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 86 Sequence 6 N49P18.1 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgagaatctcc tgtgaggctcaaggatacacattcactgactacgtcattcactggattcgacgggcccct ggacaaggccttgaatggatggggtggattaatccaggttatggacaagtaaatattcca tggaactttcagggcagggtctccatgacccgagacacgtccatcgaaacagcatttctg gacttaagaggtctaaagtctgacgacacgggcctctattattgcgtgagagatcgaagt aatggatggggaaagcgattcgagtcctccaattggttcctcgatctgtggggccgcggc actgtggtcactgttcactcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtatctcatgctccgtgatgcatgagg ctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 87 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcaatgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagacccccc aaattattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcagtggactccaggatgacgatgacgcc gaatacatttgttgggcatatgaagctttcggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 88 Sequence 7 N49P19 gcggacttggtgcagtctggggctgtggtgaaaaatgctggggcctcagtgagggtctcc tgtgaggcttatggatacacattcgtggactacttcattcattgggtccgacaggcccct ggacaaggctttgaatggatgggatacatggatcccttgaacgggcgcccaaacattgcg cgaaaatttcagggcaggctctccctgagtcgagataggtccagcgaaacttcatttctg gacttaagtggactgaggtctgacgactcggccgtctattattgtgtgagagacaagagt aatggatcgggcagacggtttgactcgtctaattggtttctcgatctgtggggccgtgga acccgggtcagtattttctcagcctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc
cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 89 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcaactcctggacagtcagtcaccatc tcctgcactggaacccacaattatgtctcttggtgtcaacaacatccaggcagagccccc aaattactaatttatgacttcaataagaggccctcaggggtcccggatcgcttctctggc tccggatctggcggcacggcctccctaaccatcactggactccaggatgacgatgaagcg gactatttttgttgggcctatgatgcttttggcggagggaccaagttgaccgtcctgcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 90 Sequence 8 N49P37 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgagggtctcc tgtgaggcttatggatacacattcagtgactacatcattcattggattcgacgggcccct ggacgaggccttgaatggatgggatggatgaatccgatgggcggacaagtgaatattccg tggaactttcaggggagagtctccatgacccgggacacgtccatcgaaacagcatttctg gacttaagaggactgaggtctgacgacacggccgtctattactgtgtgagagatcgcagc aatggatcgggcaagcgatttgagtcctccaattggttcctcgatctgtggggccgcggg accgcggtcactatttcctcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 91 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcagctcctggacagtccgtcaccatt tcctgcactggaacccacaatttggtttcttggtgtcaacatcacccaggcagagccccc aagttattaatttatgacttcaataagagaccctcaggtgtccctgatcgtttctctggc tccgggtctggcggcacggcctccctaaccatcactggactccaggatgacgatgaggct gaatatttttgttgggcgtatgaagtttttggcggagggaccaagttgaccgtgcttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 92 Sequence 9 N49P38 gcggacttggtgcagtctggggctgtggtgaagacgcctggggcctcagtgagggtctcc N49P38.1 tgtgaggcttatggatacacattcattgactacatcattcattgggtccgacaggcccct ggacaaggttttgaatggctgggatacatcgatcctatgaacgggcgcccaaacattgcg cgaaaatttcagggcaggctctccctgagccgggatacgtccatcgaaacatcatttctg gacttaagtggactgaggtctgacgactcggccgtctattattgtgtgagagacaagagt aatggatcgggcaaacgatttgactcctctaattggtttctcgatctgtggggccgtgga acgcgggtcagcatttcttcagcctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 93 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcagctcctggacagtcagtcaccatc tcctgcactggaacccacaattatgtctcttggtgtcaacaacatccaggcagagccccc aaattactaatttatgacttcaataagaggccctcaggggtcccggatcgcttctctggc tccggatctggcggcacggcctccctaaccatcactagactccaggatgacgatgacgct gactatttttgttgggcgtatgatgcttttggcggagggaccaagttgaccgtcctgcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 94 Sequence N49P55 gcggacttggtgcagtctggggctgtggtgaagaagcctggggcctcagtgagggtctcc 10 tgtgaggcttatggatacacattcactgactacatcattcattggattcgacaggcccct ggacaaggccttgaatggatgggatggatgaatcctatgggcgggcgcacaaatattccg tggaaatttcagggcagggtctccatgacccgggacacgtccatcgaaacagcatttctg gacttaagtggactaacgtctgacgacacggccgtctattattgcgtgagagacaagagt aatggatcgggcaaacgatttgactcctctaattggttcctcgatctgtggggccgcgga accccggtcactatttcctcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 95 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcagctcctggacagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacaacacccaggcagagccccc aaattattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctaagtatcactggactccaggatgacgatgaagct gaatatttttgttgggcgtatgaagcttttggcggagggaccaagttgaccgtccttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 96 Sequence N49P56 gcggacttggtgcagtctggggctgtggtgaagaagcctggggcctcagtgcgggtctcc 11 tgtgaggcttatggatatacattcgttgactacctcattcattgggtccgacaggccccc ggacaaggttttgaatggatgggatacatggatcctatgaacgggcgcccaaatattgcg cgaaaatttcagggcaggctctccctgagccgagatacgtccatcgaaacatcatttctg gacttaagtggactgaggtctgacgactcggccgtctattattgtgtgagagacaagagt ggtggatcgggcaaactatttgactcctctaattggtttctcgatctgtggggccgtgga acccgggtcagcatttcttcagcctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 97 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcagctcctggacagtcagtcaccatc tcctgcaccggaactcacaattatgtctcttggtgtcaacaacatccaggcagagccccc aaattactaatttatgacttcaataagaggccctcaggggtcccggatcgcttctctggc tccggatctggcggcacggcctccctaaccatcactggactccaggatgacgatgacgct gattatttttgttgggcgtatgatgcttttggcggagggaccaagttgaccgtcctgcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 98 Sequence N49P57 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgagaatctcc 12 tgtgaggctcaaggatacacattcactgactacgtcattcactggattcgacgggcccct ggacaaggccttgaatggatggggtggattaatccaggttatggacaagtaaatattcca tggaactttcagggcagggtctccatgacccgagacacgtccatcgaaacagcttttctg gagttaagaggtctaaagtctgacgacacgggcctctattattgcgtgagagatcgaagt aatggatggggaaagcgattcgagtcctccaattggttcctcgatctgtggggccgcggc actgtgattactgttcactcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 99 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcaatgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagacccccc aaattattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcactggactccaggatgacgatgacgcc gaatacatttgttgggcatatgaagctttcggcggagggaccaagttgaccatacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 100 Sequence N49P58 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgagaatctcc 13 tgtgaggctcagggatatacattcaccgactacgtcattcattggattcgacgggcccct ggacaaggccttgaatggatggggtggatggatccaagttatggacaagtcaatattcca cggaactttcagggcagggtctccatgacccgggacacgttcagggaaacagcatatctg gaattaagaggtctacagtctgacgacaagggcctctattattgtgtgagagatcgaagt cacggatcgggaaggcaattcgagtcctccaactggttcctcgatctgtggggccgcggc actgtggtcaatgttcagtcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 101 ------------------------------------------------------------
cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagacctccc aaattattaatttatgacttcaataagagggcttcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccattagtggactccaggatgacgatgacgcc gaatatttttgttgggcatatgaagctttcggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 102 Sequence N49P59 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcactgagaatctcc 14 tgtgaggctcaaggatacacattcactgactacgtcattcactggattcgacgggcccct ggacaaggccttgaatggatgggatggatggatccaagttttggacaaatgaacattcca cggaactttcagggcagggtctccatgacccgtgacatgtacatcgaaacagcatttctg gacttaagaggtctaaagtctgacgacacgggcctctattattgcgtgagagatcgaagt catggatcgggaaggctattcgagtcctccaattggttcctcgatctgtggggccgcggc actgtggtcactgttcagtcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 103 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagacctccc aaattattaatttatgacttcaataagagggcttcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccattagtggactccaggatgacgatgacgcc gaatatttttgttgggcatatgaagctttcggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 104 Sequence N49P73 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgagaatctcc 15 tgtgaggctcaaggatacagattcactgactacgtcattcattggattcgacgggcccct ggacaaggccttgaatggatggggttgatggatccaagttttggacgaatgaatattcca cggaaatttcagggcagggtctccatgacccgggacacgtccatggaaacagcatttctg gacttcagaggtctaaattttgacgacacgggcctctattattgcgtgagagatcgaagt catggatcgggaagactattcgagtcctccaattggttcctcgatctgtggggccgcggc actgtggtcactgttcagtcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 105 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagacctccc aaattattaatttatgacttcaataagagggcatcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcagtggactccaagatgacgatgacgcc gaatatttttgttgggcatatgaagctttcggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 106 Sequence N49P74 ccggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgagaatttcc 16 tgtgaggctcaaggatacacattcattgactacgtcattcactggattcgacgggcccct ggacaaggccttgaatggatggggttgatggatccaacttatggacgaatgaatattcca cggaagtttcagggcagggtctccatgacccgggacacgtccatcgaaacagcatttctg gacttaagaggtctaaaatctgacgacacgggcctctattattgcgtgagagatcgaagt catggatcgggaaggctattcgagtcctccaactggttcctggatctgtggggccgcggc actgtggtcactgttcagtcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacct acatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccca aatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctggggggac cgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccctg aggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactggt acgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaaca gcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaagg agtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcca aagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgagc tgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatcg ccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtgc tggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtggc agcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgc agaagagcctctccctgtctccgggtaaa SEQ ID NO: 107 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagacctccc aaattattaatttatgacttcaataagagggcttcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcagtggactccaagatgacgatgacgcc gaatatttttgttgggcatatgaagctttcggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 108 Sequence N49P75 gcggacttggtgcagtctggggctgtggtgaagaagcctggggactcagtgaggatctcc 17 tgtgaggctcaaggatacagatttcttgactacatcattcactggattcgacgggcccct ggacaaggccttgaatggatgggatggatgaatccaatgggcggacaagtaaacattcca tggaactttcagggtagggtctccatgacccgggacacgtccatcgaaacagcatttctg gacttaagaggactaaagtctgacgacacggccgtctattattgcgtgagagatcgcagt aatggatcgggaaagcgattcgagtcctccaattggttcctcgatctgtggggccgcggg actgcggtcactattcattcaccctccaccaagggcccatcggtcttccccctggcaccc tcctccaagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttc cccgaaccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttc ccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctcc agcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaag gtggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgccca gcacctgaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacacc ctcatgatctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagac cctgaggtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaag ccgcgggaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcac caggactggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcc cccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacacc ctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaa ggcttctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaac tacaagaccacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctc accgtggacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgag gctctgcacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 109 ------------------------------------------------------------ cagtctgccctgactcagcctcgctcagtgtccgcatctcctgggcagtccgtcaccatt tcctgcactggaacccacaatttggtctcttggtgtcaacatcacccaggcagagccccc aaattattaatttatgacttcaataagaggccctcaggggtccctgatcgcttctctggc tccgggtctggcggcacggcctccctgaccatcactggactccaggatgacgatgacgcc gaatatttttgttgggcgtatgaagcttttggcggagggaccaagttgaccgtacttcgt cagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggct tggaaagcagatagcagccccgtcaaggcgggagtggagaccaccacaccctccaaacaa agcaacaacaagtacgcggccagcagctatctgagcctgacgcctgagcagtggaagtcc cacagaagctacagctgccaggtcacgcatgaagggagcaccgtggagaagacagtggcc cctacagaatgttca SEQ ID NO: 110 Sequence N49P9 cacgtccaattggtgcagtctggaggtggggtgaagaagattggggccgctgtgaggatc 18 N49P9.1 tcctgcgaggtgactggatataaattcatggaccaactcataaactgggtgcggcaggcc N49P9.2 cccggtcagggccttgagtggatgggatggatgaatccaacatatggacaagtaaattat N49P9i7 tcatggagatttgaaggaagggtcaccatgaccagggacatggacaccgagacggccttc atggagttgagaggactgagagtggacgacacggccgtctattattgcgcgagggggccc tctggggaaaattatccttttcactattggggccagggtgtccgagtggtcgtctcgtca ccctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 111 ------------------------------------------------------------ gcatctgccctgactcagcctgcctccatgtctgcgtcccctggacagtcggtaaccatc tcgtgctctggaaccagacacataatctctgcttggttccaacaatatccaggcaaacca cccaaactcataatttttgacgacgataagcgtccctctggagttcctagtcgcttctct gcctccaggcctggcgacacggcctccctgacaatctctaatgttcaacctgaggacgag gcgacgtacatttgcaatacatatgaattctttggcggagggaccagattgaccgtccta agtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaa gccaacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtg gcctggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaa caaagcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaag tcccacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtg gcccctgcagaatgctct SEQ ID NO: 112 Sequence N49P22 cacattcagttgctacagtcggggcctcaagttaagaagtctggggacacagtgagaatc 19 tcctgtgagacctctggatataacttcgtcgactcccgtatccactgggtccgacagacc ccggaaaaacgtctcagatggatgggctggatcaatcctctccaaggtggtgtgaattac gcgccggaatttcagggcagaatcaggatgaccagggacacatttatagacacagtttac gtggacctgagcggactgacaccggccgacacggcctattattactgcgcgcgagggatc gatggcaagtcttacccctttcatttctggggccacggaacccgggtcaccgtcttctcg gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 113 ------------------------------------------------------------ cgatttgccctgactcaacctgcctccgtgtctgggtctcctggacagacgatcaccata acctgcgctggaggcagcgtctcctggtttcatttccctccaggcaaaacccccagactc attatttatgagtcttctaagcgaccctctggggtctctcctcgattctctgggtcccag tctggcagcacggcctcccttataatttctggcctccagtctgatgacgaagggacatac ttctgttctattcttgaatttttcggcagagggactcttgtcaccgtcctgagtcagccc aaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaag gccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaag gcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaac aacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacaga agctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgca gaatgctct SEQ ID NO: 114 Sequence N49P23 caggtgcgcttggtgcagtctggggctggggcgaggaagactggggcctcaatgaaactt 20 tcctgctcgacctctggatacaccttcaccactcatcacggccacttcataaattgggtg
cgacaggcccgtggacaagggcttgagtggatggggtggatgaatcccatgactgggcag atgaatattgaggggaaatttcagggcagagtcaccctcactcgagacatatacagtgac acggcttacatggaaatgaccagactgacaactggcgacacgggcacttattactgtgcg cgaggcgatttcggacagaattatccctttcattattggggccagggaagcctggtcatc gtctcctcggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagc acctctgggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtg acggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtccta cagtcctcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggc acccagacctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaa gttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactc ctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcc cggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaag ttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggag cagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctg aatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaa accatctccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcc cgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatccc agcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacg cctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaag agcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaac cactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 115 ------------------------------------------------------------ ctgtctgccctgactcagcctgcctccgtgtctgggtctcctgggcagtcggtcaccatc tcctgctctggaacgaaccgttaccttgtctcctggtatcaacaacaccctgacaaagcc cccaaactcatcatttatgacgacaataagcggccctcaggaatttctgatcgcttctca gcctccaggcctgacgacacggcctccctgacaatctctggactccagactggggacgag gctacttattggtgtgcctcatatgaacgttttggcggcgggacgaggctgaccgtcctt agtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaa gccaacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtg gcctggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaa caaagcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaag tcccacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtg gcccctgcagaatgctct SEQ ID NO: 116 Sequence N49P9.3 cacgtccaattggtgcagtctggaggtggggtgaagaagattggggccgctgtgaggatc 21 N49P9.4 tcctgcgaggtgtctggatacaacttcatggaccaattcataaattgggtgcgacaggcc cccggtcagggccttgagtggatgggatggatgaacccaatatatggacaagtaaattat tcatggagatttcaaggaagggtcaccatgaccagggacatgtacaccgacacggccttc atggagttgagaggactgagagtggacgacacggccgtctattattgcgcgagggggccc tctggggaaaattatccttttcactattggggccagggtgtccgagtggtcgtctcgtca ccctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 117 ------------------------------------------------------------ gcatctgccctgactcagcctgcctccatgtctgcgtcccctggacagtcggtaaccatc tcgtgctctggaaccagacacataatctctgcttggttccaacaatatccaggcaaacca cccaaactcataatttttgacgacgataagcgtccctctggagttcctagtcgcttctct gcctccaggcctggcgacacggcctccctgacaatctctaatgttcaacctgaggacgag gcgacatacatttgcaatacatatgaattctttggcggagggaccaaattgaccgtccta agtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaa gccaacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtg gcctggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaa caaagcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaag tcccacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtg gcccctgcagaatgctct SEQ ID NO: 118 Sequence N49P51 cacattcagttgctacagtcggggcctcaagttaagaagtctggggacacagtgagaatc 22 tcctgtgagacctctggatataacttcgtcgactcccgtatccactgggtccgacagacc ccggaaaaacgtctcagatggatgggctggatcaatcctctccacggtggtgtgaattac gcgccggaatttcagggcagaatcaggatgaccagggacacatttatagacacagtttac gtggacctgagcggactgacaccggccgacacggcctattattactgcgcgcgagggatc gatggcaagtcttacccctttcatttctggggccacggaacccgggtcaccgtcttctcg gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 119 ------------------------------------------------------------ cgatttgccctgactcaacctgcctccgtgtctgggtctcctggacagacgatcaccata acctgcgctggaggcagcgtctcctggtttcatttccctccaggcaaaacccccagactc attatttatgagtcttctaagcgaccctctggggtctctcctcgattctctgggtcccag tctggcagcacggcctccctcataatttctggcctccagtctgatgacgaagggacatac ttctgttctattcttgaatttttcggcagagggactcttgtcaccgtcctgagtcagccc aaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaag gccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaag gcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaac aacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacaga agctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgca gaatgctct SEQ ID NO: 120 Sequence N49P52 cgtgttacattacaacaatctggggctatagtgaggcagcctggggcctcagtgaccgtc 23 tcctgcgagacttctggatatactttcaccaagtatttcatctactgggtgcgacaggcc cctggacagggtcttgagtggctgggcagaatacacccccgaaccggtgccgtgaagtat gcaccgagatttcagggtagactgtccatgaccagagactggtcactcgacacagcctac ctcggattgaccggactgacactcggcgacacggctctatatttctgtgcgaggggggcc tttgaggcagattcatatgggtcaagttatccctttcaccactggggccagggaacccta gtcaccgtctccgcggcctccaccaagggcccatcggtcttccccctggcaccctcctcc aagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttccccgaa ccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggct gtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctccagcagc ttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggac aagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacct gaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatg atctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgag gtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgg gaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggac tggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatc gagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacaccctgccc ccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttc tatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaag accacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtg gacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctg cacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 121 ------------------------------------------------------------ tcctgggccctgactcaacccgcctccgtgtctgcgtctcctgggcagtcggtcaccatg tcctgcactggattcggaaattataaccctgactcctggtaccaacaatacccaggcaaa gcccccaaactcatcatttatgaagacaataaaagaccctcgggggtctctgatcgcttc tctgcctccagacttggcagcacgtcttccctgacaatctctaacgtccaggctgcggac gacgcccattatgtctgcgcctcctttgaatttttcggcggagggaccaagctgaccgtc ctgagtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagctt caagccaacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgaca gtggcctggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctcc aaacaaagcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtgg aagtcccacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagaca gtggcccctgcagaatgctct SEQ ID NO: 122 Sequence N49P53 cgtgtgacattacaacaatctggggctacagtgaagcagcctggggcctcagtgaccgtc 24 tcctgcgagacttctggatacactttcaccaagtataccattcactgggtgcgacaggcc cctggacagggtcttcagtgggtgggcagaatacacccccgaaccggtgccgtgaagtat gcaccgatatttcagggtaaagtgtccatgagtcgagacttgtcacgcgacacagcctac ctcggattgaccagactgacgctcgccgacacggctctatttttctgtgcgaggggggcc tttgaggcagatttaagtgggccaacttacccctttcaccactggggccaaggaacccta gtcatcgtctccgcggcctccaccaagggcccatcggtcttccccctggcaccctcctcc aagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttccccgaa ccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggct gtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctccagcagc ttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggac aagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacct gaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatg atctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgag gtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgg gaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggac tggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatc gagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacaccctgccc ccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttc tatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaag accacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtg gacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctg cacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 123 ------------------------------------------------------------ tcctgggccctgactcaacccgcctccgtgtctgcgtctcctgggcagtcggtcaccatg tcctgcactggattcggaaattataaccctgactcctggtaccaacaatacccaggcaaa gcccccaaactcatcatttatgaggacaataaaagaccctcgggagtctctaatcgcttc tctgcctccagacttggcagcacgtcttccctgacaatctctaacgtccaggccgctgac gacgcccattatgtctgcgcctcctttgaatttttcggcggagggaccaagctgatcgtc ctgagtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagctt caagccaacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgaca gtggcctggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctcc aaacaaagcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtgg aagtcccacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagaca gtggcccctgcagaatgctct SEQ ID NO: 124 Sequence N49P54 aacgtgcagttgatgcagtctgggactgaggtgaagaagtctggggcctcggtgacaatc 25 tcttgtgagaccgctggattcaacttcatcgactccgtcatacactggctgcgccaggcc cctggaggaggatttcagtggatggggtggatcaagcctcttagaggtgccgtcaattat ccacagtttttgcagggcagggtctccatgacccgggacttgtccaccgacacggtgtac atggtcttgaatggactgacacctgacgacacaggcctttattactgcgcgaaaggggcc tttagagggggttctccctttggcttctggggccagggaactctgctcaccgtctcccca gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 125 ------------------------------------------------------------ cagtctgccctgtctcagcctgtctccgtgtctgggtctcctggagagtcgatcaccatt tcctgtactggagccaccacctggtatcaacaactcccaggcagaccccccaaactcatc atttatgacgtcactaaccggccctcaggcatttctagtcgtttctctggctccacgtct ggccacacggcctccctgacaatctccggtctccaggttgacgacgagggtctgtatcac tgcgcctcacgtgaatttttcggcggagggaccaagctgaccgtcctgagtcagcccaag gctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggcc acactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaaggca gatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaacaac aagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacagaagc tacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgcagaa tgctct SEQ ID NO: 126 Sequence N49P60 caggtgcgactggtgcagtctgggcctcaggtgaagaagactggggcctcagtgagggtc 26 tcctgcgaaacctctggatacacgttcacctcctacttcatccattggttacgactgggc cccggagaggggcttcagtggatggggtggatcaaccctttacatggtgccgtgaattat gaaaacaaatttaggggcagggtcacaatcaccagggacacgtccacagacacagtgtat ttggacatgagcagactgacccctgacgacacggccgtctatttctgcacaagaggaatc gttgctgatgggtggccctatggccactggggccagggaacccaagtcaccgtctccccg gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc
gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 127 ------------------------------------------------------------ tcctgggccctgactcagcctgcctccgtgtctgggtctcctggacagtcggtcgccatc tcctgcgctggcggcagcgtctcctggtaccaggtgctcccaggcagagcccccaaactc atcatttatgagggcgctaagcgaccctcaggggtttctgctcgcttctctggctcccag tctggcaacacggcttacctgacaatttctgacctccagactgaggacgagggcatctac ttctgctcttcacttcaattcttcggcggagggaccaaactgaccgtcctaagtcagccc aaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaag gccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaag gcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaac aacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacaga agctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgca gaatgctct SEQ ID NO: 128 Sequence N49P61 caggtgcgacttcagcagtctggtgtcgtggtgaggaagcctggggcctcagtgagaatt 27 tcctgcgagacttctggattcaccttcatcgaccacattgtccattgggtgcggcgggcc cctggacgaggctttgaatggatgggttggatcaagcctcttaggggtgccgtagattat gcaccccaacttcggggcaggatctccctgacgagggacatttacagtgaaaccgtcttt atagacgtgagtcgactgacgtctggcgacacggcgatatacttttgttgtaaggccgcc gcccctgaagaagcattcccccttcaatactggggccaggggacccaacttatcgtctcc tcggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctct gggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtg tcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcc tcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccag acctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgag cccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggg ggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacc cctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaac tggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtac aacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggc aaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatc tccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggat gagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgac atcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctccc gtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcagg tggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactac acgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 129 ------------------------------------------------------------ caggctgccctgactcagcccgcctccgtgtccggctctcctggacagtcggtcaccatt tcctgcctttatgccaatgtagatatctgctggtatcaactacacccgggcagagccccc aaacttctaattgttgacaataataagcggccctcaggagtctctcctcgcttctctggc tccaagtctggcaccacggcctccctgacaatctctggacttcaggctgacgacgaggct gaatatcactgctcttcaagaacattttttggcggggggaccaagttgaccgtcctgagt cagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcc tggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaa agcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcc cacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcc cctgcagaatgctct SEQ ID NO: 130 Sequence N49P62 caggtgcgacttcagcagtctggtgtcgtggtgaggaagcctggggcctcagtgagactt 28 tcctgcgagacgtctggattcaaattcatcgaccacattgtcaactgggtgcggcgggcc cctggacgaggctttgaatggatgggttggatcaagcctcttgggggtgtcgctgattat gcaccccaacatcggggcaggatctcactgacgagggacatttacactgaaaccgtcttt atagacctgagtcgactgacgtctggcgacacggcgatttatttctgttgtaaggccgcc gcccctgatgaagcattcccccttgaatactggggccaggggacccaacttatcgtctcc ccggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctct gggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtg tcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcc tcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccag acctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgag cccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggg ggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacc cctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaac tggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtac aacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggc aaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatc tccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggat gagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgac atcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctccc gtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcagg tggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactac acgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 131 ------------------------------------------------------------ caggctgccctgactcagcccgcctccgtgtccggctctcctggacagtcggtcaccatt tcctgcctttatgccaatgtagatatctgctggtatcaaatacagccgggcagattaccc aaacttctgattgttgacaataataggcgaccctcaggagtctctcctcgcttctctggc tccaagtctggcaccacggcctccctgacaatctctggacttcaggctgacgacgaggct gaatatcactgctcttcaacaacattttttggcggggggaccaagttgaccgtcctcagt cagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcc tggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaa agcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcc cacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcc cctgcagaatgctct SEQ ID NO: 132 Sequence N49P63 caggtgcgacttgtgcagtctggtcccgtgatgagaaagcctggggcctcagtgagaatt 29 tcttgcgagacatctggattcgccttcttggaccacattgtccactgggtgcggcgggcc cctggacgcggctttgaatggatgggttgggttaagaccattgggggtgtcgttgattat gcaccccaccttaggggcaggatctccgtgacgagagacgtctttagtgaaaccgtcttt ctggacttgagtcgactgacgtctggcgacacggcgatgtatttttgttctaaggccgcc gcccctgacgaagccttcccccttgaattttggggccaggggacccaagtcatcgtctcc tcggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctct gggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtg tcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcc tcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccag acctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgag cccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggg ggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacc cctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaac tggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtac aacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggc aaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatc tccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggat gagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgac atcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctccc gtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcagg tggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactac acgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 133 ------------------------------------------------------------ caggctgccctgactcagcccgcctccgtgtccggctctcctggacagtcggtcaccatt tcttgcctttatgccaatgtggatatctgctggtatcaacttcacccgggcagagccccc aaacttcttattcttgacaataataaacggccctcaggagtctctagtcgcttctccggt tccaagtctggcaccacggcctccctaaccatctctgaccttcaggctgacgacgaggct gaatatcactgctcttcaacaacattttttggcggggggaccaggttgaccgtcctgagt cagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcc tggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaa agcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcc cacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcc cctgcagaatgctct SEQ ID NO: 134 Sequence N49P64 caggtgcgacttgtgcagtctggtcccgtggtgagaaagcctgggacctcagtgagaatt 30 tcttgcgagacatctggattcgccttcttggaccacattgtccactgggtgcggcgggcc cctggacgcggctttgaatggatgggttgggttaagaccattgggggtgtcgttgattat gcaccccaccttaggggcaggatctccgtgacgagggacgtatttagtgaaatcgtcttt atggagttgagtcgactgacgtctggcgacacggcgatgtatttttgttctaaggccgcc gcccctgacgaagccttcccccttgaattttggggccaggggacccaagtcatcgtctcc tcggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctct gggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtg tcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcc tcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccag acctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgag cccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggg ggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacc cctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaac tggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtac aacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggc aaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatc tccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggat gagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgac atcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctccc gtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcagg tggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactac acgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 135 ------------------------------------------------------------ caggctgccctgactcagcccgcctccgtgtccggctctcctggacagtcggtcaccatt tcctgcctttatgccaatgtggatatctgctggtatcaacttcacccgggcagagccccc aaacttctaattgttgacaataataagcggccctcaggagtctctagtcgcttctctggt tccaagtctggcaccacggcctccctaacaatctctgatcttcaggctgacgacgaggct gaatatcactgctcttcaacaacattttttggcggggggaccaggttgaccgtcctgagt cagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcc tggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaa agcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcc cacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcc cctgcagaatgctct SEQ ID NO: 136 Sequence N49P65 caagtgcaactggtgcagtctggggctggggtgaagaagcctggggcctcagtgagggtc 31 tcctgcgagacatccggattcaagttcaccgagtactttatccactttttacgacaggcc cctggacaagggcttgagtggatgggatggctcaaccctctcagaggtgccgtcaactat ccacggaagtttcagggcagagtcactttgaccagggacatctacaccaccaccgtctac atgcaacttaacggtctgacccctgacgacacggccgtctactactgtgccagagcggtc tttaatgaagctttcccctttgactactggggccagggaagcctggtcaccgtctcctca gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 137 ------------------------------------------------------------ tcctgggcccagactcagcctgcctccgtgtctgggtctcctggacagtcgatcaccatc tcctgcgctggaatcgtcagtgatgcctggtaccagcaatacccaggcagaccccccaga ctcatcctttatgacggcgataagcggccctcaggggtttctcctcgtttttctgcctcc agggccggcaagacggcctccctgacaatttctgggctgcaggctgacgacgaggcttat tatcactgcgcgtcaagggaattttttggaggcgtgaccaagttgaccgtcctaagtcag cccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaac aaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctgg aaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagc aacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccac agaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccct gcagaatgctct SEQ ID NO: 138 Sequence N49P66 caggtgcgacttcagcagtctggtgtcgtggtgaggaagcctggggcctcagtgagactt 32 tcctgcgagacgtctggcttcaaattcatcgaccacattgtcaactgggtgcggcgggcc cctggacgaggctttgaatggatgggttggatcaagcctcttgggggtgtcgctgattat gcaccccaacatcggggcaggatctcactgacgagggacatttacactgaaaccgtcttt atagacctgagtcgactgacgtctggcgacacggcgatttatttttgttgtaaggccgcc gcccctgatgaagcattcccccttgaatactggggccaggggacccaacttatcgtctcc ccggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctct gggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtg tcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcc tcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccag acctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgag cccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggg ggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacc cctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaac tggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtac aacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggc aaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatc tccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggat gagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgac atcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctccc gtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcagg tggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactac acgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 139 ------------------------------------------------------------ caggctgccctgactcagcccgcctccgtgtccggctctcctggacagtcggtcaccatt tcctgcctttatgccaatgtagatatctgctggtatcaaatacagccgggcagattaccc aaacttctgattgttgacaatgataggcgaccctcaggagtctctcctcgcttctctggc tccaagtctggcaccacggcctccctgacaatctctggacttcaggctgacgacgaggct gaatatcactgctcttcaacaacattttttggcggggggaccaagttgaccgtcctcagt cagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcc tggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaa agcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcc cacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcc cctgcagaatgctct
SEQ ID NO: 140 Sequence N49P67 caggtgcgacttgtgcagtctggtcccgtgatgagaaagcctggggcctcagtgagaatt 33 tcttgcgagacatctggattcgccttcttggaccacattgtccactgggtgcggcgggcc cctggacgcggctttgaatggatgggttgggttaagaccattgggggtgtcgttgattat gcaccccaccttaggggcaggatctccgtgacgagagacgtctttagtgaaaccgtcttt ctggacttgagtcgactgacgtctggcgacacggcgatgtatttttgttctaaggccgcc gcccctgacgaagccttcccccttgaattttggggccaggggacccaagtcatcgtctcc tcggcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctct gggggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtg tcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcc tcaggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccag acctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgag cccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggg ggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacc cctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaac tggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtac aacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggc aaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatc tccaaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggat gagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgac atcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctccc gtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcagg tggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactac acgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 141 ------------------------------------------------------------ caggctgccctgactcagcccgcctccgtgtccggctctcctggacagtcggtcaccatt tcttgcctttatgccaatgtggatatctgctggtatcaacttcacccgggcagagccccc aaacttctaattcttgacaataataaacggccctcaggagtctctagtcgcttctccggt tccaagtctggcaccacggcctccctaaccatctctgaccttcaggctgacgacgaggct gaatatcactgctcttcaacaactttttttggcggggggaccaggttgaccgtcctgagt cagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagcc aacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcc tggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaa agcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcc cacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcc cctgcagaatgctct SEQ ID NO: 142 Sequence N49P68 cacgtgcagttgaggcagtctgggactgaggcgaagaagtctggggcctcggtgacaatc 34 tcttgtgagaccgctggattcaacttcatcgactccgtcatacactggctgcgccaggcc cctggtgggggatttcagtggatggggtggatcaagcctcttagaggtggcgtcaattat ccacattatttgcagggcagaatctccatgacccgggacttgtccagtgacacggtttac atggtcttaaatagactgacacctgccgacacaggcctttattactgcgcgaaaggggcc tttggggggagttctccctttggcttctggggccagggaactctgctcaccgtctcccca gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 143 ------------------------------------------------------------ cagtctgccctgtctcagcctgtctccgtgtctgggtctcctggagagtcgatcaccatt tcctgtactgaagccaccacctggtatcaacaactcccaggcaaaccccccaaactcatc atttatgacgtgaccaaccggccctcaggcatttcaagtcgtttctctggctccatgtct ggtcgcacggcctccctgacaatctccggtctccaggttgacgacgagggtctctatcac tgtgcctcacgtgaatttttcggcggggggaccaagctgaccgtcctgagtcagcccaag gctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggcc acactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaaggca gatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaacaac aagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacagaagc tacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgcagaa tgctct SEQ ID NO: 144 Sequence N49P69 cacgtgcaattgatgcagtctgggactcaggcgaagaagtctggggcctcggtgacaatt 35 tcttgtgagaccgctggattcaagttcatcgactccgtcatacactggctgcgccaggcc cctggagggggatttcagtggatggggtggatcaagcctcttggaggtgccgtcaactat ccaccctatttgcagggcaggatctccttgacccgtgacttgtccaccgacacaatttac atggtcttgaatggactgacacctgccgacacaggcttttattactgcgccaaaggggcc tttggggggggttctccctttggcttctggggccaggggactctgctcaccgtctcccca gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 145 ------------------------------------------------------------ cagtctgccctgtctcagcctgtctccgtgtctgggtctcctggagactcgatcaccatt tcttgttttggagccaccacctggtatcaacaactcccaggcagaccccccaaactcatc atttatgacgtgactaaccggccctcaggcatttcaggtcgtttctctggctccatgtct ggtcaaaaggcctccctgacaatctccggtctccaggttgacgacgagggtctctatcac tgcgcctcacgtgaatttttcggcggggggaccaaactgaccgtcctgagtcagcccaag gctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggcc acactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaaggca gatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaacaac aagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacagaagc tacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgcagaa tgctct SEQ ID NO: 146 Sequence N49P70 cacgtgcagttgaggcagtctgggactgaggcgaagaagtctggggcctcggtgacaatc 36 tcttgtgagaccgctggattcaacttcatcgactccgtcatacactggctgcgccaggcc cctggtgggggatttcagtggatggggtggatcaagcctcttagaggtggcgtcaattat ccacattatttgcagggcagaatctccatgacccgggacttgtccagtgacacggtttac atggtcttaaatagactgacacctgacgacacaggcctttactactgcgcgaaaggggcc tttggggggagttctccctttggcttctggggccagggaactctgctcaccgtctcccca gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 147 ------------------------------------------------------------ cagtctgccctgtctcagcctgtctccgtgtctgggtctcctggagagtcgatcaccatt tcctgtactgaagccaccacctggtatcaacaactcccagggagatcccccaaactcatt atttatgacgtgaccaaccggccctcaggcatttcaagtcgtttctctggctccatgtct ggtcgcacggcctccctgacaatctccggtctccaggttgacgacgagggtctctatcac tgtgcctcacgtgaatttttcggcggggggaccaagctgaccgtcctcagtcagcccaag gctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggcc acactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaaggca gatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaacaac aagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacagaagc tacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgcagaa tgctct SEQ ID NO: 148 Sequence N49P71 cgtgttacattacaacagtctggggctacagtgaggcagcctggggcctcagtcaccgtc 37 tcctgcgagacttctggattcaccttcatcaaatataccattcactgggtgcgacaggcc cctggacagggtcttcagtgggtgggaagaatacacccccgaaccggtgccgtgaagttt gcaccgatatttcagggtaaattttccatgagtcgagacttgtcacgcgacacagcctac ctcggattgaccagactgacactcgccgacacggctctatttttctgtgcgaggggggcc tttgaggcagatttatatgggccaacttacccctttcaccactggggccaaggaacccaa gtcaccgtctccgcggcctccaccaagggcccatcggtcttccccctggcaccctcctcc aagagcacctctgggggcacagcggccctgggctgcctggtcaaggactacttccccgaa ccggtgacggtgtcgtggaactcaggcgccctgaccagcggcgtgcacaccttcccggct gtcctacagtcctcaggactctactccctcagcagcgtggtgaccgtgccctccagcagc ttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaaggtggac aagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacct gaactcctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatg atctcccggacccctgaggtcacatgcgtggtggtggacgtgagccacgaagaccctgag gtcaagttcaactggtacgtggacggcgtggaggtgcataatgccaagacaaagccgcgg gaggagcagtacaacagcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggac tggctgaatggcaaggagtacaagtgcaaggtctccaacaaagccctcccagcccccatc gagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgtacaccctgccc ccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttc tatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaag accacgcctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtg gacaagagcaggtggcagcaggggaacgtcttctcatgctccgtgatgcatgaggctctg cacaaccactacacgcagaagagcctctccctgtctccgggtaaa SEQ ID NO: 149 ------------------------------------------------------------ tcctgggccctgactcaacccgcctccgtgtctgcgtctcctgggcagtcggtcaccatg tcctgcactggattcggaagttataatcctgactcctggtaccagcaatacccaggcaaa gcccccaaactcatcatttatgatgacaataaaagaccctcgggggtctctgatcgcttc tctgcctccagacttggcagcacatcttcactgacaatctctaacgtccaggccgctgac gacgcccattatgtctgcgcctcctttgagtttttcggcggggggaccaagctgaccgtc ctgagtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagctt caagccaacaaggccacactggtgtgtctcgtaagtgacttctacccgggagccgtgaca gtggcctggaaggcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctcc aaacaaagcaacaacaagtatgcggccagcagctacctgagcctgacgcccgagcagtgg aagtcccacagaagctacagctgccgggtcacgcatgaagggagcaccgtggagaagaca gtggcccctgcagaatgctct SEQ ID NO: 150 Sequence N49P72 cacattcagttgctacagtcggggcctcaagttaagaagtctggggacacagtgagaatc 38 tcctgtgagacctctggatataatttcgtcgactcccttatccactgggtccgacagacc ccggaaaaacgtctcagatggatgggctggatcaatcctctccaaggtggtgtgaattac gcgccggaatttcagggcagaatcaggatgaccagggacacgtttatagacacagtttac gtggacttgagcggactgacaccggccgacacggcctattattactgcgcgcgagggatc gatggcaattcttacccctttcatttctggggccacggaacccgggtcaccgtcttctcg gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctggg ggcacagcggccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcg tggaactcaggcgccctgaccagcggcgtgcacaccttcccggctgtcctacagtcctca ggactctactccctcagcagcgtggtgaccgtgccctccagcagcttgggcacccagacc tacatctgcaacgtgaatcacaagcccagcaacaccaaggtggacaagaaagttgagccc aaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaactcctgggggga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactgg tacgtggacggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaac agcacgtaccgggtggtcagcgtcctcaccgtcctgcaccaggactggctgaatggcaag gagtacaagtgcaaggtctccaacaaagccctcccagcccccatcgagaaaaccatctcc aaagccaaagggcagccccgagaaccacaggtgtacaccctgcccccatcccgggatgag ctgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctatcccagcgacatc gccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtg ctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg cagcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacg cagaagagcctctccctgtctccgggtaaa SEQ ID NO: 151 ------------------------------------------------------------ cgatttgccctgactcaacctgcctccgtgtctgggtctcctggacagacgatcaccata acctgcgctggaggcagcgtctcctggttccatttccctccaggcaaaacccccagactc attatttatgagtcttctaagagaccctcaggggtctctcctcgattctctgggtcccag tctggcagcacggcctccctaataatttctggcctccagtctgatgacgaagggacatac ttctgttctattcttgaatttttcggcagagggactcttctcaccgtcctgagtcagccc aaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaag gccacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaag gcagatggcagccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaac aacaagtatgcggccagcagctacctgagcctgacgcccgagcagtggaagtcccacaga agctacagctgccgggtcacgcatgaagggagcaccgtggagaagacagtggcccctgca gaatgctct SEQ ID NO: 152
TABLE-US-00004 TABLE 4 List of mAb variants and the corresponding sequence and mutations. The amino acid sequence of the variants is determined based on the reference antibody sequence (see table 2, above for reference antibody sequences) and the mutations described in the table. See below amino acid sequences and corresponding oligonucleotide sequences. Related to Heavy chain Light chain mAb sequence mutation mutations Swaps N49P6 1 CH1: 1.4A, 120R Constant: 1.5G LC7 swap CH3: 12 E, 14M N49P6.2 1 CH1: 120R CH3: 12 E, 14M N49P7 2 CH1: 1.4A, 120R Constant: 1.5G CH3: 12 E, 14M N49P7.1 2 CH1: 120R CH3: 12 E, 14M N49P7A 2 CH1: 1.4A, 120R Variable: 42A CH3: 12 E, 14M Constant: 1.5G N49P7F 2 CH1: 1.4A, 120R Variable: 42F CH3: 12 E, 14M Constant: 1.5G N49P7S 2 CH1: 1.4A, 120R Variable: 42S CH3: 12 E, 14M Constant: 1.5G N49P7Y 2 CH1: 1.4A, 120R Variable: 42Y CH3: 12 E, 14M Constant: 1.5G N49P7-54TY 2 Variable: 59T/62Y Constant: 1.5G CH1: 1.4A, 120R CH3: 12 E, 14M N49P7LS-1 2 CH1: 1.4A, 120R Constant: 1.5G CH3: 12 E, 14M, 107L, 114S N49P7LS-2 2 CH1: 1.4A, 120R Constant: 1.5G CH3: 107L, 114S N49P7YTE 2 CH1: 1.4A, 120R Constant: 1.5G CH2: 15.1Y, 16T, 18E N49P7L6 2.1 CH1: 1.4A, 120R Constant: 1.5G N49P7 heavy chain CH3: 12 E, 14M with N49P6 light chain N49P7L11 2.4 CH1: 1.4A, 120R N49P7 heavy chain CH3: 12 E, 14M with N49P11 light chain N49P7.1L9 2.18 CH1: 120R N49P7.1 heavy CH3: 12 E, 14M chain with N49P9 light chain N49P7.1L19 2.7 CH1: 120R N49P7 heavy chain CH3: 12 E, 14M with N49P19 light chain R49P7 2 CH1: 1.4A Constant: 1.5G Rhesus IgG1, Rhesus LC3 N49P7.2 3 CH1: 120R CH3: 12 E, 14M N49P11 4 CH1: 120R CH3: 12 E, 14M N49P18 5 CH1: 120R CH3: 12 E, 14M N49P18.2 5 CH1: 120R LC7 swap CH3: 12 E, 14M N49P18.1 6 CH1: 120R CH3: 12 E, 14M N49P19 7 CH1: 120R CH3: 12 E, 14M N49P37 8 CH1: 1.4A, 120R Constant: 1.5G CH3: 12 E, 14M N49P38 9 CH1: 120R Constant: 1.5G CH3: 12 E, 14M N4938.1 9 CH1: 120R CH3: 12 E, 14M N49P55 10 CH1: 120R CH3: 12 E, 14M N49P56 11 CH1: 120R CH3: 12 E, 14M N49P57 12 CH1: 120R CH3: 12 E, 14M N49P58 13 CH1: 120R CH3: 12 E, 14M N49P59 14 CH1: 120R CH3: 12 E, 14M N49P73 15 CH1: 120R CH3: 12 E, 14M N49P74 16 CH1: 120R CH3: 12 E, 14M N49P75 17 CH1: 120R CH3: 12 E, 14M N49P75.1 17 CH1: 1.4A, 120R Constant: 1.5G CH3: 12 E, 14M N49P9 18 CH1: 120R CH3: 12 E, 14M N49P9.1 18 CH1: 120R LC2 swap CH3: 12 E, 14M N49P9.2 18 CH1: 1.4A, 120R Constant: 1.5G CH3: 12 E, 14M N49P9i7 18.2 CH1: 120R Part of CDRH3 of CH3: 12 E, 14M N49P7 swapped for entire CDRH3 of N49P9 N49P9i7H1 18.2 CH1: 120R CDRH3 of N49P7 CH3: 12 E, 14M swapped for CDRH3 of N49P9 N49P9i7H2 18.2 CH1: 1.4S, 120R Constant: 1.5R CDRH3 and junction CH3: 12 E, 14M of N49P7 swapped for CDRH3 of N49P9 N49P22 19 CH1: 120R CH3: 12 E, 14M N49P23 20 CH1: 120R CH3: 12 E, 14M N49P9.3 21 CH1: 120R Constant: 1.5R CH3: 12 E, 14M N49P9.4 21 CH1: 120R CH3: 12 E, 14M N49P51 22 CH1: 120R CH3: 12 E, 14M N49P52 23 CH1: 120R CH3: 12 E, 14M N49P53 24 CH1: 120R CH3: 12 E, 14M N49P54 25 CH1: 120R CH3: 12 E, 14M N49P60 26 CH1: 120R CH3: 12 E, 14M N49P61 27 CH1: 120R CH3: 12 E, 14M N49P62 28 CH1: 120R CH3: 12 E, 14M N49P63 29 CH1: 120R CH3: 12 E, 14M N49P64 30 CH1: 120R CH3: 12 E, 14M N49P65 31 CH1: 120R CH3: 12 E, 14M N49P66 32 CH1: 120R CH3: 12 E, 14M N49P67 33 CH1: 120R CH3: 12 E, 14M N49P68 34 CH1: 120R CH3: 12 E, 14M N49P69 35 CH1: 120R CH3: 12 E, 14M
[0648] Amino acid and nucleotide sequences of additional anti-HIV antibodies are shown below. Variable regions within the heavy and light chain in the amino acid sequence are shaded and changes to the amino acid sequence relative to a natural antibody sequence are underlined. CDR residues are in bold.
TABLE-US-00005 >N49P6 HEAVY CHAIN SEQ ID NO: 153 ##STR00153## ##STR00154## ##STR00155## YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPK PKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLH QDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQK SLSLSPGK SEQ ID NO: 154 GCGGGACTGATGCAGTCTGGGGCTGTGATGAAGAATTCGGGGGCCTCAGTGAGGGTCTCTTGT CAGGCTGATGGATACGACTTCATTGACTATGTCATTCACTGGTTTCGACAAAGACGTGGAGAA GGTCTTGAGTGGCTGGGATGGATGAATCCCTCGGGAGGCGGCACAAACTATCCGCGACCATTT CAGGGCAAAGTCACCATGACCAGGGACACGTCCACCGAGACAGCCTATTTAGATGTCAGAGG ACTTACATATGACGACACGGCCGTCTATTATTGTGTGAGAGACAGGGCCAACGGTTCGGGAA GAAGACGTTTTGAGTCGGTGAATTGGTTCCTGGATCTGTGGGGCCGCGGCACCCAAATAACAG TCGTCTCGGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTC TGGGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTC GTGGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGG ACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACAT CTGCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTT GTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCT TCCTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCG TGGTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTG GAGGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGG TCAGCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCT CCAACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGA GAACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCT GACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGC AGCCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCT ACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTG ATGCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 155 ##STR00156## ##STR00157## YPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVE KTVAPAECS SEQ ID NO: 156 CAGTCTGCCCTAACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGTCAGTCAGTCACCATCTCCT GCACTGGAACACACAATTATGTGTCCTGGTGTCAACAGAAACCGGGCCAAGCCCCCAAATTAT TAATTTACGATTTCAATAAACGGCCCTCAGGGGTCTCTGATCGCTTCTCTGGCTCCACGTCTGG CAACACGGCCTCCCTGACCATCTCTGGACTCCAGGCTGACGATGAGGGTCATTATTTTTGTTG GGCGTTTGAAAATATCGGCGGAGGGACCAAGCTGACCGTCCTGGGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCC GTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAG CAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCA CGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P6.2 HEAVY CHAIN SEQ ID NO: 157 ##STR00158## ##STR00159## ##STR00160## YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPK PKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLH QDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQK SLSLSPGK SEQ ID NO: 158 GCGGGACTGATGCAGTCTGGGGCTGTGATGAAGAATTCGGGGGCCTCAGTGAGGGTCTCTTGT CAGGCTGATGGATACGACTTCATTGACTATGTCATTCACTGGTTTCGACAAAGACGTGGAGAA GGTCTTGAGTGGCTGGGATGGATGAATCCCTCGGGAGGCGGCACAAACTATCCGCGACCATTT CAGGGCAAAGTCACCATGACCAGGGACACGTCCACCGAGACAGCCTATTTAGATGTCAGAGG ACTTACATATGACGACACGGCCGTCTATTATTGTGTGAGAGACAGGGCCAACGGTTCGGGAA GAAGACGTTTTGAGTCGGTGAATTGGTTCCTGGATCTGTGGGGCCGCGGCACCCAAATAACAG TCGTCTCGGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTC TGGGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTC GTGGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGG ACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACAT CTGCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTT GTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCT TCCTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCG TGGTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTG GAGGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGG TCAGCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCT CCAACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGA GAACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCT GACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGC AGCCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCT ACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTG ATGCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 159 ##STR00161## ##STR00162## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 160 CAGTCTGCCCTAACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGTCAGTCAGTCACCATCTCCT GCACTGGAACACACAATTATGTGTCCTGGTGTCAACAGAAACCGGGCCAAGCCCCCAAATTAT TAATTTACGATTTCAATAAACGGCCCTCAGGGGTCTCTGATCGCTTCTCTGGCTCCACGTCTGG CAACACGGCCTCCCTGACCATCTCTGGACTCCAGGCTGACGATGAGGGTCATTATTTTTGTTG GGCGTTTGAAAATATCGGCGGAGGGACCAAGCTGACCGTCCTGGGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7 HEAVY CHAIN SEQ ID NO: 161 ##STR00163## ##STR00164## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 162 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC
AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 163 ##STR00165## ##STR00166## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 164 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7.1 HEAVY CHAIN SEQ ID NO: 165 ##STR00167## ##STR00168## SSTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 166 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCATCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 167 ##STR00169## ##STR00170## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 168 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7A HEAVY CHAIN SEQ ID NO: 169 ##STR00171## ##STR00172## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 170 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 171 ##STR00173## ##STR00174## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 172 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGGCTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7F HEAVY CHAIN SEQ ID NO: 173 ##STR00175## ##STR00176## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 174 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT
TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 175 ##STR00177## ##STR00178## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 176 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTTTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7S HEAVY CHAIN SEQ ID NO: 177 ##STR00179## ##STR00180## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 178 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 179 ##STR00181## ##STR00182## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 180 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTCTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7Y HEAVY CHAIN SEQ ID NO: 181 ##STR00183## ##STR00184## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 182 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGT GTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 183 ##STR00185## ##STR00186## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 184 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTATCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG
TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7-54TY HEAVY CHAIN SEQ ID NO: 185 ##STR00187## ##STR00188## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 186 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAACGTACGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 187 ##STR00189## ##STR00190## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 188 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7LS-1 HEAVY CHAIN SEQ ID NO: 189 ##STR00191## ##STR00192## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVLHEALHSHYTQKSLSLSP GK SEQ ID NO: 190 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGCT GCATGAGGCTCTGCACAGCCACTACACGCAGAAGAGCCTCTCCCTGTCCCCGGGTAAA LIGHT CHAIN SEQ ID NO: 191 ##STR00193## ##STR00194## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 192 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7LS-2 HEAVY CHAIN SEQ ID NO: 193 ##STR00195## ##STR00196## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVLHEALHSHYTQKSLSLSP GK SEQ ID NO: 194 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA
ACCACAGGTGTACACCCTGCCCCCATCCCGGGATGAGCTGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGCT GCATGAGGCTCTGCACAGCCACTACACGCAGAAGAGCCTCTCCCTGTCCCCGGGTAAA LIGHT CHAIN SEQ ID NO: 195 ##STR00197## ##STR00198## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 196 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7YTE HEAVY CHAIN SEQ ID NO: 197 ##STR00199## ##STR00200## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL YITREPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 198 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCTATATCACCCGGGAGCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 199 ##STR00201## ##STR00202## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 200 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7L6 HEAVY CHAIN SEQ ID NO: 201 ##STR00203## ##STR00204## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 202 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 203 ##STR00205## ##STR00206## YPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVE KTVAPAECS SEQ ID NO: 204 CAGTCTGCCCTAACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGTCAGTCAGTCACCATCTCCT GCACTGGAACACACAATTATGTGTCCTGGTGTCAACAGAAACCGGGCCAAGCCCCCAAATTAT TAATTTACGATTTCAATAAACGGCCCTCAGGGGTCTCTGATCGCTTCTCTGGCTCCACGTCTGG CAACACGGCCTCCCTGACCATCTCTGGACTCCAGGCTGACGATGAGGGTCATTATTTTTGTTG GGCGTTTGAAAATATCGGCGGAGGGACCAAGCTGACCGTCCTGGGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCC GTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAG CAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCA CGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P7L11 HEAVY CHAIN SEQ ID NO: 205 ##STR00207## ##STR00208## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK
SEQ ID NO: 206 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 207 ##STR00209## ##STR00210## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 208 CAGTGTGTCTTGACTCAGCCTCGCTCAGTGTCCGGATCTCCTGGACAATCAGTCACCATCTCCT GCACTGGAACTCACAATTATGTCTCCTGGTGTCAACACCACCCAGGCAACGCCCCCAAATTAT TACTTTATGATTTCGACAAGCGGCCCTCAGGAATCTCTGATCGCTTCTCTGGCTCTAGGTCTGG CAACACGGCCTCCCTGACCATCTCTGGCCTCCAGCCTGAGGATGAGGCCGATTACTTTTGTTG GGCCTTTGAAGCCTTTGGCGGAGGGACCAAGGTGCTCGTCCTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P7.1L9 HEAVY CHAIN SEQ ID NO: 209 ##STR00211## ##STR00212## SSTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 210 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCATCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 211 ##STR00213## ##STR00214## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 212 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTAAGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P7.1L19 HEAVY CHAIN SEQ ID NO: 213 ##STR00215## ##STR00216## SSTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 214 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCATCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 215 ##STR00217## ##STR00218## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 216 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAACTCCTGGACAGTCAGTCACCATCTCCT GCACTGGAACCCACAATTATGTCTCTTGGTGTCAACAACATCCAGGCAGAGCCCCCAAATTAC TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCGGATCGCTTCTCTGGCTCCGGATCTG
GCGGCACGGCCTCCCTAACCATCACTGGACTCCAGGATGACGATGAAGCGGACTATTTTTGTT GGGCCTATGATGCTTTTGGCGGAGGGACCAAGTTGACCGTCCTGCGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >R49P7 HEAVY CHAIN SEQ ID NO: 217 ##STR00219## ##STR00220## ASTKGPSVFPLAPSSRSTSESTAALGCLVKDYFPEPVTVSWNSGSLTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYVCNVNHKPSNTKVDKRVEIKTCGGGSKPPTCPPCPAPELLGGPSVFLFPPKPK DTLMISRTPEVTCVVVDVSQEDPDVKFNWYVNGAEVHHAQTKPRETQYNSTYRVVSVLTVTHQD WLNGKEYTCKVSNKALPAPIQKTISKDKGQPREPQVYTLPPSREELTKNQVSLTCLVKGFYPSDIV VEWESSGQPENTYKTTPPVLDSDGSYFLYSKLTVDKSRWQQGNVFSCSVLHEALHSHYTQKSLSL SPGK SEQ ID NO: 218 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATATAGATTTCCTGACTACATCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCCTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAATATTCCATGGAAATT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCACAATTC AATCAGCCTCGACCAAGGGCCCATCGGTCTTCCCCCTGGCGCCCTCCTCCAGGAGCACCTCCG AGAGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCTGAACCCGTGACCGTGTCGT GGAACTCAGGCTCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGGC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACGTCT GCAACGTAAACCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGATAAAAACATG TGGTGGTGGCAGCAAACCTCCCACGTGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACC GTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGT CACATGCGTGGTGGTAGACGTGAGCCAGGAAGACCCCGATGTCAAGTTCAACTGGTACGTAA ATGGCGCGGAGGTGCATCATGCCCAGACGAAGCCACGGGAGACGCAGTACAACAGCACATAT CGTGTGGTCAGCGTCCTCACCGTCACGCACCAGGACTGGCTGAACGGCAAGGAGTACACGTG CAAGGTCTCCAACAAAGCCCTCCCGGCCCCCATCCAGAAAACCATCTCCAAAGACAAAGGGC AGCCCCGAGAGCCTCAGGTGTACACCCTGCCCCCGTCCCGGGAGGAGCTGACCAAGAACCAG GTCAGCCTGACCTGCCTGGTCAAAGGCTTCTACCCCAGTGACATCGTCGTGGAGTGGGAGAGC AGCGGGCAGCCGGAGAACACCTACAAGACCACCCCGCCCGTGCTGGACTCCGACGGCTCCTA CTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATG CTCCGTGCTGCATGAGGCTCTGCACAGCCACTACACGCAGAAGAGCCTCTCCCTGTCCCCGGG TAAA LIGHT CHAIN SEQ ID NO: 219 ##STR00221## ##STR00222## YPGAVEVAWKADGSAVNAGVETTKPSKQSNNKYAASSYLSLTSDQWKSHKSYSCQVTHEGSTV EKTVAPAECS SEQ ID NO: 220 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCAGCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTTCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCGAGCGAGGAGCTTCAAGCCAACAAGGCCACACTAGTGTG TCTGATCAGTGACTTCTACCCGGGAGCCGTGGAAGTGGCCTGGAAGGCAGATGGCAGCGCTG TCAACGCGGGAGTGGAGACCACCAAACCCTCCAAACAGAGCAACAACAAGTACGCGGCCAGC AGCTACCTGAGCCTGACGTCCGACCAGTGGAAGTCCCACAAGAGCTACAGCTGCCAGGTCAC GCACGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGTTCA >N49P7.2 HEAVY CHAIN SEQ ID NO: 221 ##STR00223## ##STR00224## ##STR00225## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 222 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATACAAATTTCCTGACTACATCATTCACTGGATTCGACGCGCCCCTGGACAA GGCCTTGAGTGGATGGGGTGGATTAATCCAATGGGCGGACAAGTAAACATTCCATGGCAGTTT CAGGGCAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCCTCTATTATTGCGTGAGAGATCGAAGTAATGGATCGGGAA GGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGCGGTCACTGTTC ATTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 223 ##STR00226## ##STR00227## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 224 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCAGTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P11 HEAVY CHAIN SEQ ID NO: 225 ##STR00228## ##STR00229## ##STR00230## YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPK PKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLH QDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQK SLSLSPGK SEQ ID NO: 226 TCGGCGGAATTGGTGCAATCTGGGGCTGTGGTGAAGAAGCCTGGGACCTCCGTGAAGGTCTCT TGTCAGGCTTATGGATACACTTTTACCGACTACCTTATTCATTGGCTTCGACAGGCCCCTGGAC AAGGACTTGAATGGATGGGATGGATGAATCCTGTTTATGGACAAGTAAATTATGCCCAAAACT TTCAGGGCAGGGTCTCCATGACCAGGGACATTTACAGGGAAACAGCATTTCTAGAGGTGCGC GACCTGAAGACTGACGACACAGGCACTTATTATTGTGTGAGAGACACAGGCGACGGTTCGCG GAGACACTTTGACTCCATCAATTGGTTTCTCGATCTTTGGGGCCGCGGGACATGGATAAGGGT CGCCCCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTC TGGGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTC GTGGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGG ACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACAT CTGCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTT GTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCT
TCCTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCG TGGTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTG GAGGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGG TCAGCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCT CCAACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGA GAACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCT GACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGC AGCCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCT ACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTG ATGCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 227 ##STR00231## ##STR00232## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 228 CAGTGTGTCTTGACTCAGCCTCGCTCAGTGTCCGGATCTCCTGGACAATCAGTCACCATCTCCT GCACTGGAACTCACAATTATGTCTCCTGGTGTCAACACCACCCAGGCAACGCCCCCAAATTAT TACTTTATGATTTCGACAAGCGGCCCTCAGGAATCTCTGATCGCTTCTCTGGCTCTAGGTCTGG CAACACGGCCTCCCTGACCATCTCTGGCCTCCAGCCTGAGGATGAGGCCGATTACTTTTGTTG GGCCTTTGAAGCCTTTGGCGGAGGGACCAAGGTGCTCGTCCTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P18 HEAVY CHAIN SEQ ID NO: 229 ##STR00233## ##STR00234## ##STR00235## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 230 GCGGACTTGGTGCAGTCTGGGGCTGTGATGAAGAAGCCTGGGGACTCAGTGAGAATCTCCTGT GAGGCTCGAGGATACACATTCACTGACTACGTCATTCACTGGATTCGACGCGCCCCTGGACAA GGCCTTGAATGGATGGGGTGGATTGATCCACCTTATGGACAAGTAAATATTCCATGGAATTTT CAGGGCAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG TCTAAAGTCTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTAATGGATGGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGGTCACTGTTC ACTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 231 ##STR00236## ##STR00237## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 232 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGAGCCCCCAAGTTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTAACCATCAGTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACTGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P18.2 HEAVY CHAIN SEQ ID NO: 233 ##STR00238## ##STR00239## ##STR00240## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 234 GCGGACTTGGTGCAGTCTGGGGCTGTGATGAAGAAGCCTGGGGACTCAGTGAGAATCTCCTGT GAGGCTCGAGGATACACATTCACTGACTACGTCATTCACTGGATTCGACGCGCCCCTGGACAA GGCCTTGAATGGATGGGGTGGATTGATCCACCTTATGGACAAGTAAATATTCCATGGAATTTT CAGGGCAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG TCTAAAGTCTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTAATGGATGGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGGTCACTGTTC ACTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 235 ##STR00241## ##STR00242## FYPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTV EKTVAPAECS SEQ ID NO: 236 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGAGCCCCCAAGTTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTAACCATCAGTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACTGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P18.1 HEAVY CHAIN SEQ ID NO: 237 ##STR00243## ##STR00244## ##STR00245## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 238 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGAATCTCCTGT GAGGCTCAAGGATACACATTCACTGACTACGTCATTCACTGGATTCGACGCGCCCCTGGACAA GGCCTTGAATGGATGGGGTGGATTAATCCAGGTTATGGACAAGTAAATATTCCATGGAACTTT CAGGGCAGGGTCTCCATGACCCGAGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG TCTAAAGTCTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTAATGGATGGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGGTCACTGTTC ACTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 239 ##STR00246## ##STR00247## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 240 CAGTCTGCCCTGACTCAGCCTCGCTCAATGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGACCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCAGTGGACTCCAGGATGACGATGACGCCGAATACATTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P19 HEAVY CHAIN SEQ ID NO: 241 ##STR00248## ##STR00249## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 242 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAAAATGCTGGGGCCTCAGTGAGGGTCTCCTGT GAGGCTTATGGATACACATTCGTGGACTACTTCATTCATTGGGTCCGACAGGCCCCTGGACAA GGCTTTGAATGGATGGGATACATGGATCCCTTGAACGGGCGCCCAAACATTGCGCGAAAATTT CAGGGCAGGCTCTCCCTGAGTCGAGATAGGTCCAGCGAAACTTCATTTCTGGACTTAAGTGGA CTGAGGTCTGACGACTCGGCCGTCTATTATTGTGTGAGAGACAAGAGTAATGGATCGGGCAG ACGGTTTGACTCGTCTAATTGGTTTCTCGATCTGTGGGGCCGTGGAACCCGGGTCAGTATTTTC TCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 243 ##STR00250## ##STR00251## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 244 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAACTCCTGGACAGTCAGTCACCATCTCCT GCACTGGAACCCACAATTATGTCTCTTGGTGTCAACAACATCCAGGCAGAGCCCCCAAATTAC TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCGGATCGCTTCTCTGGCTCCGGATCTG GCGGCACGGCCTCCCTAACCATCACTGGACTCCAGGATGACGATGAAGCGGACTATTTTTGTT GGGCCTATGATGCTTTTGGCGGAGGGACCAAGTTGACCGTCCTGCGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P37 HEAVY CHAIN SEQ ID NO: 245 ##STR00252## ##STR00253## ##STR00254## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 246 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGGTCTCCTGT GAGGCTTATGGATACACATTCAGTGACTACATCATTCATTGGATTCGACGGGCCCCTGGACGA GGCCTTGAATGGATGGGATGGATGAATCCGATGGGCGGACAAGTGAATATTCCGTGGAACTT TCAGGGGAGAGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAG GACTGAGGTCTGACGACACGGCCGTCTATTACTGTGTGAGAGATCGCAGCAATGGATCGGGC AAGCGATTTGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGGACCGCGGTCACTATT TCCTCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 247 ##STR00255## ##STR00256## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE
KTVAPTECS SEQ ID NO: 248 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAGCTCCTGGACAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTTTCTTGGTGTCAACATCACCCAGGCAGAGCCCCCAAGTTAT TAATTTATGACTTCAATAAGAGACCCTCAGGTGTCCCTGATCGTTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTAACCATCACTGGACTCCAGGATGACGATGAGGCTGAATATTTTTGTTG GGCGTATGAAGTTTTTGGCGGAGGGACCAAGTTGACCGTGCTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P38 HEAVY CHAIN SEQ ID NO: 249 ##STR00257## ##STR00258## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 250 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGACGCCTGGGGCCTCAGTGAGGGTCTCCTGT GAGGCTTATGGATACACATTCATTGACTACATCATTCATTGGGTCCGACAGGCCCCTGGACAA GGTTTTGAATGGCTGGGATACATTGATCCTATGAACGGGCGCCCAAACATTGCGCGAAAATTT CAGGGCAGGCTCTCCCTGAGCCGGGATACGTCCATCGAAACATCATTTCTGGACTTAAGTGGA CTGAGGTCTGACGACTCGGCCGTCTATTATTGTGTGAGAGACAAGAGTAATGGATCGGGCAA ACGATTTGACTCCTCTAATTGGTTTCTCGATCTGTGGGGCCGTGGAACGCGGGTCAGCATTTCT TCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 251 ##STR00259## ##STR00260## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 252 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAGCTCCTGGACAGTCAGTCACCATCTCCT GCACTGGAACCCACAATTATGTCTCTTGGTGTCAACAACATCCAGGCAGAGCCCCCAAATTAC TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCGGATCGCTTCTCTGGCTCCGGATCTG GCGGCACGGCCTCCCTAACCATCACTAGACTCCAGGATGACGATGACGCTGACTATTTTTGTT GGGCGTATGATGCTTTTGGCGGAGGGACCAAGTTGACCGTCCTGGGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P38.1 HEAVY CHAIN SEQ ID NO: 253 ##STR00261## ##STR00262## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 254 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGACGCCTGGGGCCTCAGTGAGGGTCTCCTGT GAGGCTTATGGATACACATTCATTGACTACATCATTCATTGGGTCCGACAGGCCCCTGGACAA GGTTTTGAATGGCTGGGATACATTGATCCTATGAACGGGCGCCCAAACATTGCGCGAAAATTT CAGGGCAGGCTCTCCCTGAGCCGGGATACGTCCATCGAAACATCATTTCTGGACTTAAGTGGA CTGAGGTCTGACGACTCGGCCGTCTATTATTGTGTGAGAGACAAGAGTAATGGATCGGGCAA ACGATTTGACTCCTCTAATTGGTTTCTCGATCTGTGGGGCCGTGGAACGCGGGTCAGCATTTCT TCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 255 ##STR00263## ##STR00264## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 256 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAGCTCCTGGACAGTCAGTCACCATCTCCT GCACTGGAACCCACAATTATGTCTCTTGGTGTCAACAACATCCAGGCAGAGCCCCCAAATTAC TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCGGATCGCTTCTCTGGCTCCGGATCTG GCGGCACGGCCTCCCTAACCATCACTAGACTCCAGGATGACGATGACGCTGACTATTTTTGTT GGGCGTATGATGCTTTTGGCGGAGGGACCAAGTTGACCGTCCTGCGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P55 HEAVY CHAIN SEQ ID NO: 257 ##STR00265## ##STR00266## PSTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 258 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGCCTCAGTGAGGGTCTCCTGT GAGGCTTATGGATACACATTCACTGACTACATCATTCATTGGATTCGACAGGCCCCTGGACAA GGCCTTGAATGGATGGGATGGATGAATCCTATGGGCGGGCGCACAAATATTCCGTGGAAATTT CAGGGCAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGTGG ACTAACGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGACAAGAGTAATGGATCGGGCA AACGATTTGACTCCTCTAATTGGTTCCTCGATCTGTGGGGCCGCGGAACCCCGGTCACTATTTC CTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGG GGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTG GAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACT CTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTG CAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTG
ACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCC TCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGG TGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAG GTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAG CGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCA ACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAA CCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGAC CTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGC CGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACA GCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATG CATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 259 ##STR00267## ##STR00268## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 260 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAGCTCCTGGACAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACAACACCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTAAGTATCACTGGACTCCAGGATGACGATGAAGCTGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTCCTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P56 HEAVY CHAIN SEQ ID NO: 261 ##STR00269## ##STR00270## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 262 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGCCTCAGTGCGGGTCTCCTGT GAGGCTTATGGATATACATTCGTTGACTACCTCATTCATTGGGTCCGACAGGCCCCCGGACAA GGTTTTGAATGGATGGGATACATGGATCCTATGAACGGGCGCCCAAATATTGCGCGAAAATTT CAGGGCAGGCTCTCCCTGAGCCGAGATACGTCCATCGAAACATCATTTCTGGACTTAAGTGGA CTGAGGTCTGACGACTCGGCCGTCTATTATTGTGTGAGAGACAAGAGTGGTGGATCGGGCAA ACTATTTGACTCCTCTAATTGGTTTCTCGATCTGTGGGGCCGTGGAACCCGGGTCAGCATTTCT TCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 263 ##STR00271## ##STR00272## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 264 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCAGCTCCTGGACAGTCAGTCACCATCTCCT GCACCGGAACTCACAATTATGTCTCTTGGTGTCAACAACATCCAGGCAGAGCCCCCAAATTAC TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCGGATCGCTTCTCTGGCTCCGGATCTG GCGGCACGGCCTCCCTAACCATCACTGGACTCCAGGATGACGATGACGCTGATTATTTTTGTT GGGCGTATGATGCTTTTGGCGGAGGGACCAAGTTGACCGTCCTGCGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P57 HEAVY CHAIN SEQ ID NO: 265 ##STR00273## ##STR00274## ##STR00275## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 266 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGAATCTCCTGT GAGGCTCAAGGATACACATTCACTGACTACGTCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCTTGAATGGATGGGGTGGATTAATCCAGGTTATGGACAAGTAAATATTCCATGGAACTTT CAGGGCAGGGTCTCCATGACCCGAGACACGTCCATCGAAACAGCTTTTCTGGAGTTAAGAGGT CTAAAGTCTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTAATGGATGGGGAAA GCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGATTACTGTTCAC TCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 267 ##STR00276## ##STR00277## PGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEK TVAPTECS SEQ ID NO: 268 CAGTCTGCCCTGACTCAGCCTCGCTCAATGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGACCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATACATTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACCATACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P58 HEAVY CHAIN SEQ ID NO: 269 ##STR00278## ##STR00279##
##STR00280## SSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKD TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQD WLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLS LSPGK SEQ ID NO: 270 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGAATCTCCTGT GAGGCTCAGGGATATACATTCACCGACTACGTCATTCATTGGATTCGACGGGCCCCTGGACAA GGCCTTGAATGGATGGGGTGGATGGATCCAAGTTATGGACAAGTCAATATTCCACGGAACTTT CAGGGCAGGGTCTCCATGACCCGGGACACGTTCAGGGAAACAGCATATCTGGAATTAAGAGG TCTACAGTCTGACGACAAGGGCCTCTATTATTGTGTGAGAGATCGAAGTCACGGATCGGGAAG GCAATTCGAGTCCTCCAACTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGGTCAATGTTCA GTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGG GGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTG GAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACT CTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTG CAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTG ACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCC TCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGG TGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAG GTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAG CGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCA ACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAA CCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGAC CTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGC CGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACA GCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATG CATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 271 ##STR00281## ##STR00282## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 272 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGACCTCCCAAATTAT TAATTTATGACTTCAATAAGAGGGCTTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATTAGTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P59 HEAVY CHAIN SEQ ID NO: 273 ##STR00283## ##STR00284## ##STR00285## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 274 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCACTGAGAATCTCCTGT GAGGCTCAAGGATACACATTCACTGACTACGTCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCTTGAATGGATGGGATGGATGGATCCAAGTTTTGGACAAATGAACATTCCACGGAACTTT CAGGGCAGGGTCTCCATGACCCGTGACATGTACATCGAAACAGCATTTCTGGACTTAAGAGGT CTAAAGTCTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTCATGGATCGGGAAG GCTATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGGTCACTGTTCAG TCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 275 ##STR00286## ##STR00287## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 276 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGACCTCCCAAATTAT TAATTTATGACTTCAATAAGAGGGCTTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATTAGTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P73 HEAVY CHAIN SEQ ID NO: 277 ##STR00288## ##STR00289## ##STR00290## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 278 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGAATCTCCTGT GAGGCTCAAGGATACAGATTCACTGACTACGTCATTCATTGGATTCGACGGGCCCCTGGACAA GGCCTTGAATGGATGGGGTTGATGGATCCAAGTTTTGGACGAATGAATATTCCACGGAAATTT CAGGGCAGGGTCTCCATGACCCGGGACACGTCCATGGAAACAGCATTTCTGGACTTCAGAGG TCTAAATTTTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTCATGGATCGGGAAG ACTATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGCACTGTGGTCACTGTTCAG TCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGG GGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGG AACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTC TACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGC AACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGA CAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGT GGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGC GTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAA CAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAAC CACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACC TGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCC GGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAG CAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGC ATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 279 ##STR00291##
##STR00292## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 280 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGACCTCCCAAATTAT TAATTTATGACTTCAATAAGAGGGCATCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTG GCGGCACGGCCTCCCTGACCATCAGTGGACTCCAAGATGACGATGACGCCGAATATTTTTGTT GGGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCC CCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGT GTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCC GTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P74 HEAVY CHAIN SEQ ID NO: 281 ##STR00293## ##STR00294## STKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSV VTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLM ISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 282 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGAATTTCCTGT GAGGCTCAAGGATACACATTCATTGACTACGTCATTCACTGGATTCGACGGGCCCCTGGACAA GGCCTTGAATGGATGGGGTTGATGGATCCAACTTATGGACGAATGAATATTCCACGGAAGTTT CAGGGCAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG TCTAAAATCTGACGACACGGGCCTCTATTATTGCGTGAGAGATCGAAGTCATGGATCGGGAAG GCTATTCGAGTCCTCCAACTGGTTCCTGGATCTGTGGGGCCGCGGCACTGTGGTCACTGTTCA GTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGG GGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTG GAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACT CTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTG CAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTG ACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCC TCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGG TGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAG GTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAG CGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCA ACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAA CCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGAC CTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGC CGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACA GCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATG CATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 283 ##STR00295## ##STR00296## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 284 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGACCTCCCAAATTAT TAATTTATGACTTCAATAAGAGGGCTTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCAGTGGACTCCAAGATGACGATGACGCCGAATATTTTTGTTG GGCATATGAAGCTTTCGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P75 HEAVY CHAIN SEQ ID NO: 285 ##STR00297## ##STR00298## ##STR00299## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 286 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATACAGATTTCTTGACTACATCATTCACTGGATTCGACGAGCCCCTGGACAA GGCCTTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAACATTCCATGGAACTT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGGACTGCGGTCACTATTC ATTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 287 ##STR00300## ##STR00301## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 288 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTACTTCGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P75.1 HEAVY CHAIN SEQ ID NO: 289 ##STR00302## ##STR00303## ##STR00304## SVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAV EWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK SEQ ID NO: 290 GCGGACTTGGTGCAGTCTGGGGCTGTGGTGAAGAAGCCTGGGGACTCAGTGAGGATCTCCTGT GAGGCTCAAGGATACAGATTTCTTGACTACATCATTCACTGGATTCGACGAGCCCCTGGACAA GGCCTTGAATGGATGGGATGGATGAATCCAATGGGCGGACAAGTAAACATTCCATGGAACTT TCAGGGTAGGGTCTCCATGACCCGGGACACGTCCATCGAAACAGCATTTCTGGACTTAAGAGG ACTAAAGTCTGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGATCGGGAA AGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGCGGGACTGCGGTCACTATTC ATTCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTG GGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGT
GGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGAC TCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCT GCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGT GACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTC CTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTG GTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGA GGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCA GCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCC AACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGA CCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAG CCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTAC AGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 291 ##STR00305## ##STR00306## YPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVE KTVAPTECS SEQ ID NO: 292 CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGCATCTCCTGGGCAGTCCGTCACCATTTCCT GCACTGGAACCCACAATTTGGTCTCTTGGTGTCAACATCACCCAGGCAGAGCCCCCAAATTAT TAATTTATGACTTCAATAAGAGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCGGGTCTGG CGGCACGGCCTCCCTGACCATCACTGGACTCCAGGATGACGATGACGCCGAATATTTTTGTTG GGCGTATGAAGCTTTTGGCGGAGGGACCAAGTTGACCGTACTTGGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCCCGT CAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCAGC AGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P9 HEAVY CHAIN SEQ ID NO: 293 ##STR00307## ##STR00308## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 294 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGACTGGATATAAATTCATGGACCAACTCATAAACTGGGTGCGGCAGGCCCCCG GTCAGGGCCTTGAGTGGATGGGATGGATGAATCCAACATATGGACAAGTAAATTATTCATGG AGATTTGAAGGAAGGGTCACCATGACCAGGGACATGGACACCGAGACGGCCTTCATGGAGTT GAGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGCGAGGGGACCCTCTGGGGAAA ATTATCCTTTTCACTATTGGGGCCAGGGTGTCCGAGTGGTCGTCTCGTCACCCTCCACCAAGGG CCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGG CTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGAC CAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGT GGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCC CAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCC CACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCA AGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACG AAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACA AAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCA CCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCC CCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTG CCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTT CTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGA CCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACA AGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAAC CACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 295 ##STR00309## ##STR00310## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 296 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTAAGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P9.1 HEAVY CHAIN SEQ ID NO: 297 ##STR00311## ##STR00312## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 298 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGACTGGATATAAATTCATGGACCAACTCATAAACTGGGTGCGGCAGGCCCCCG GTCAGGGCCTTGAGTGGATGGGATGGATGAATCCAACATATGGACAAGTAAATTATTCATGG AGATTTGAAGGAAGGGTCACCATGACCAGGGACATGGACACCGAGACGGCCTTCATGGAGTT GAGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGCGAGGGGACCCTCTGGGGAAA ATTATCCTTTTCACTATTGGGGCCAGGGTGTCCGAGTGGTCGTCTCGTCACCCTCCACCAAGGG CCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGG CTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGAC CAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGT GGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCC CAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCC CACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCA AGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACG AAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACA AAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCA CCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCC CCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTG CCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTT CTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGA CCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACA AGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAAC CACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 299 ##STR00313## ##STR00314## AVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTV APTECS SEQ ID NO: 300 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTAAGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCGCCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCATAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCTTGGAAAGCAGATAGCAGCCC CGTCAAGGCGGGAGTGGAGACCACCACACCCTCCAAACAAAGCAACAACAAGTACGCGGCCA GCAGCTATCTGAGCCTGACGCCTGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCAGGTC ACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTACAGAATGTTCA >N49P9.2 HEAVY CHAIN SEQ ID NO: 301 ##STR00315## ##STR00316##
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 302 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGACTGGATATAAATTCATGGACCAACTCATAAACTGGGTGCGGCAGGCCCCCG GTCAGGGCCTTGAGTGGATGGGATGGATGAATCCAACATATGGACAAGTAAATTATTCATGG AGATTTGAAGGAAGGGTCACCATGACCAGGGACATGGACACCGAGACGGCCTTCATGGAGTT GAGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGCGAGGGGACCCTCTGGGGAAA ATTATCCTTTTCACTATTGGGGCCAGGGTGTCCGAGTGGTCGTCTCGTCATCCTCCACCAAGGG CCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGG CTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGAC CAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGT GGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCC CAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCC CACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCA AGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACG AAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACA AAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCA CCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCC CCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTG CCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTT CTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGA CCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACA AGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAAC CACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 303 ##STR00317## ##STR00318## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 304 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTAGGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P9i7 HEAVY CHAIN SEQ ID NO: 305 ##STR00319## ##STR00320## PSTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 306 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGACTGGATATAAATTCATGGACCAACTCATAAACTGGGTGCGGCAGGCCCCCG GTCAGGGCCTTGAGTGGATGGGATGGATGAATCCAACATATGGACAAGTAAATTATTCATGG AGATTTGAAGGAAGGGTCACCATGACCAGGGACATGGACACCGAGACGGCCTTCATGGAGTT GAGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGAT CGGGAAAGCGATTCGAGTCCTCCAATTGGGGCCAGGGTGTCCGAGTGGTCGTCTCGTCACCCT CCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAG CGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAG GCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCC TCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGA ATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACT CACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCC CCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGAC GTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAA TGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCA CCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCC CTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGT GTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGG TCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAAC AACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTC ACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGC TCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 307 ##STR00321## ##STR00322## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 308 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTAAGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P9i7H1 HEAVY CHAIN SEQ ID NO: 309 ##STR00323## ##STR00324## ##STR00325## GLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFP PKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTV LHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFY PSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK SEQ ID NO: 310 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGACTGGATATAAATTCATGGACCAACTCATAAACTGGGTGCGGCAGGCCCCCG GTCAGGGCCTTGAGTGGATGGGATGGATGAATCCAACATATGGACAAGTAAATTATTCATGG AGATTTGAAGGAAGGGTCACCATGACCAGGGACATGGACACCGAGACGGCCTTCATGGAGTT GAGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGAT CGGGAAAGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCAGGGTGTCCGAGTGG TCGTCTCGTCACCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCAC CTCTGGGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGT GTCGTGGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTC AGGACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTA CATCTGCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAAT CTTGTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAG TCTTCCTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACAT GCGTGGTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGC GTGGAGGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGG TGGTCAGCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAG GTCTCCAACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCC CCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCA GCCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATG GGCAGCCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCC TCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCC GTGATGCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 311 ##STR00326## ##STR00327##
GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 312 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTAAGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P9i7H2 HEAVY CHAIN SEQ ID NO: 313 ##STR00328## ##STR00329## ##STR00330## YSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPK PKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLH QDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQK SLSLSPGK SEQ ID NO: 314 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGACTGGATATAAATTCATGGACCAACTCATAAACTGGGTGCGGCAGGCCCCCG GTCAGGGCCTTGAGTGGATGGGATGGATGAATCCAACATATGGACAAGTAAATTATTCATGG AGATTTGAAGGAAGGGTCACCATGACCAGGGACATGGACACCGAGACGGCCTTCATGGAGTT GAGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGTGAGAGATCGCAGTAATGGAT CGGGAAAGCGATTCGAGTCCTCCAATTGGTTCCTCGATCTGTGGGGCCGTGGGACTGCGGTCA CAATTCAATCATCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCA CCTCTGGGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGG TGTCGTGGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCT CAGGACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCT ACATCTGCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAA TCTTGTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCA GTCTTCCTCTTCCCCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACA TGCGTGGTGGTGGACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGG CGTGGAGGTGCATAATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGG GTGGTCAGCGTCCTCACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAA GGTCTCCAACAAAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGC CCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTC AGCCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAAT GGGCAGCCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTC CTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTC CGTGATGCATGAGGCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAA A LIGHT CHAIN SEQ ID NO: 315 ##STR00331## ##STR00332## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 316 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACGTACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAGATTGACCGTCCTACGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P22 HEAVY CHAIN SEQ ID NO: 317 ##STR00333## ##STR00334## PLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 318 CACATTCAGTTGCTACAGTCGGGGCCTCAAGTTAAGAAGTCTGGGGACACAGTGAGAATCTCC TGTGAGACCTCTGGATATAACTTCGTCGACTCCCGTATCCACTGGGTCCGACAGACCCCGGAA AAACGTCTCAGATGGATGGGCTGGATCAATCCTCTCCAAGGTGGTGTGAATTACGCGCCGGAA TTTCAGGGCAGAATCAGGATGACCAGGGACACATTTATAGACACAGTTTACGTGGACCTGAG CGGACTGACACCGGCCGACACGGCCTATTATTACTGCGCGCGAGGGATCGATGGCAAGTCTTA CCCCTTTCATTTCTGGGGCCACGGAACCCGGGTCACCGTCTTCTCGGCCTCCACCAAGGGCCC ATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTG CCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAG CGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGT GACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAG CAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCAC CGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGG ACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAG ACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAG CCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCA GGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCA TCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCC CCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTA TCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 319 ##STR00335## ##STR00336## VAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAPA ECS SEQ ID NO: 320 CGATTTGCCCTGACTCAACCTGCCTCCGTGTCTGGGTCTCCTGGACAGACGATCACCATAACCT GCGCTGGAGGCAGCGTCTCCTGGTTTCATTTCCCTCCAGGCAAAACCCCCAGACTCATTATTTA TGAGTCTTCTAAGCGACCCTCTGGGGTCTCTCCTCGATTCTCTGGGTCCCAGTCTGGCAGCACG GCCTCCCTTATAATTTCTGGCCTCCAGTCTGATGACGAAGGGACATACTTCTGTTCTATTCTTG AATTTTTCGGCAGAGGGACTCTTGTCACCGTCCTGAGTCAGCCCAAGGCTGCCCCCTCGGTCA CTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAA GTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTG GGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCT GAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGAAG GGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P23 HEAVY CHAIN SEQ ID NO: 321 ##STR00337## ##STR00338## ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS VVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTL MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK SEQ ID NO: 322 CAGGTGCGCTTGGTGCAGTCTGGGGCTGGGGCGAGGAAGACTGGGGCCTCAATGAAACTTTC CTGCTCGACCTCTGGATACACCTTCACCACTCATCACGGCCACTTCATAAATTGGGTGCGACA GGCCCGTGGACAAGGGCTTGAGTGGATGGGGTGGATGAATCCCATGACTGGGCAGATGAATA TTGAGGGGAAATTTCAGGGCAGAGTCACCCTCACTCGAGACATATACAGTGACACGGCTTAC ATGGAAATGACCAGACTGACAACTGGCGACACGGGCACTTATTACTGTGCGCGAGGCGATTT CGGACAGAATTATCCCTTTCATTATTGGGGCCAGGGAAGCCTGGTCATCGTCTCCTCGGCCTC CACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGC GGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGG
CGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTC AGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAAT CACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCA CACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCC AAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGT GAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATG CCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACC GTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCT CCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGT ACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTC AAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAA CTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCAC CGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTC TGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 323 ##STR00339## ##STR00340## YPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVE KTVAPAECS SEQ ID NO: 324 CTGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGGCAGTCGGTCACCATCTCCT GCTCTGGAACGAACCGTTACCTTGTCTCCTGGTATCAACAACACCCTGACAAAGCCCCCAAAC TCATCATTTATGACGACAATAAGCGGCCCTCAGGAATTTCTGATCGCTTCTCAGCCTCCAGGC CTGACGACACGGCCTCCCTGACAATCTCTGGACTCCAGACTGGGGACGAGGCTACTTATTGGT GTGCCTCATATGAACGTTTTGGCGGCGGGACGAGGCTGACCGTCCTTAGTCAGCCCAAGGCTG CCCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGG TGTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P9.3 HEAVY CHAIN SEQ ID NO: 325 ##STR00341## ##STR00342## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 326 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGTCTGGATACAACTTCATGGACCAATTCATAAATTGGGTGCGACAGGCCCCCGG TCAGGGCCTTGAGTGGATGGGATGGATGAACCCAATATATGGACAAGTAAATTATTCATGGA GATTTCAAGGAAGGGTCACCATGACCAGGGACATGTACACCGACACGGCCTTCATGGAGTTG AGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGCGAGGGGACCCTCTGGGGAAAA TTATCCTTTTCACTATTGGGGCCAGGGTGTCCGAGTGGTCGTCTCGTCACCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 327 ##STR00343## ##STR00344## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 328 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACATACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAAATTGACCGTCCTACGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P9.4 HEAVY CHAIN SEQ ID NO: 329 ##STR00345## ##STR00346## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 330 CACGTCCAATTGGTGCAGTCTGGAGGTGGGGTGAAGAAGATTGGGGCCGCTGTGAGGATCTC CTGCGAGGTGTCTGGATACAACTTCATGGACCAATTCATAAATTGGGTGCGACAGGCCCCCGG TCAGGGCCTTGAGTGGATGGGATGGATGAACCCAATATATGGACAAGTAAATTATTCATGGA GATTTCAAGGAAGGGTCACCATGACCAGGGACATGTACACCGACACGGCCTTCATGGAGTTG AGAGGACTGAGAGTGGACGACACGGCCGTCTATTATTGCGCGAGGGGACCCTCTGGGGAAAA TTATCCTTTTCACTATTGGGGCCAGGGTGTCCGAGTGGTCGTCTCGTCACCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 331 ##STR00347## ##STR00348## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 332 GCATCTGCCCTGACTCAGCCTGCCTCCATGTCTGCGTCCCCTGGACAGTCGGTAACCATCTCGT GCTCTGGAACCAGACACATAATCTCTGCTTGGTTCCAACAATATCCAGGCAAACCACCCAAAC TCATAATTTTTGACGACGATAAGCGTCCCTCTGGAGTTCCTAGTCGCTTCTCTGCCTCCAGGCC TGGCGACACGGCCTCCCTGACAATCTCTAATGTTCAACCTGAGGACGAGGCGACATACATTTG CAATACATATGAATTCTTTGGCGGAGGGACCAAATTGACCGTCCTAAGTCAGCCCAAGGCTGC CCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGT GTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCC CCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCC AGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGT CACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P51 HEAVY CHAIN SEQ ID NO: 333 ##STR00349##
##STR00350## PLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 334 CACATTCAGTTGCTACAGTCGGGGCCTCAAGTTAAGAAGTCTGGGGACACAGTGAGAATCTCC TGTGAGACCTCTGGATATAACTTCGTCGACTCCCGTATCCACTGGGTCCGACAGACCCCGGAA AAACGTCTCAGATGGATGGGCTGGATCAATCCTCTCCACGGTGGTGTGAATTACGCGCCGGAA TTTCAGGGCAGAATCAGGATGACCAGGGACACATTTATAGACACAGTTTACGTGGACCTGAG CGGACTGACACCGGCCGACACGGCCTATTATTACTGCGCGCGAGGGATCGATGGCAAGTCTTA CCCCTTTCATTTCTGGGGCCACGGAACCCGGGTCACCGTCTTCTCGGCCTCCACCAAGGGCCC ATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTG CCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAG CGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGT GACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAG CAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCAC CGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGG ACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAG ACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAG CCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCA GGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCA TCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCC CCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTA TCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 335 ##STR00351## ##STR00352## VAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAPA ECS SEQ ID NO: 336 CGATTTGCCCTGACTCAACCTGCCTCCGTGTCTGGGTCTCCTGGACAGACGATCACCATAACCT GCGCTGGAGGCAGCGTCTCCTGGTTTCATTTCCCTCCAGGCAAAACCCCCAGACTCATTATTTA TGAGTCTTCTAAGCGACCCTCTGGGGTCTCTCCTCGATTCTCTGGGTCCCAGTCTGGCAGCACG GCCTCCCTCATAATTTCTGGCCTCCAGTCTGATGACGAAGGGACATACTTCTGTTCTATTCTTG AATTTTTCGGCAGAGGGACTCTTGTCACCGTCCTGAGTCAGCCCAAGGCTGCCCCCTCGGTCA CTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAA GTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTG GGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCT GAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGAAG GGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P52 HEAVY CHAIN SEQ ID NO: 337 ##STR00353## ##STR00354## TKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSV VTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLM ISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 338 CGTGTTACATTACAACAATCTGGGGCTATAGTGAGGCAGCCTGGGGCCTCAGTGACCGTCTCC TGCGAGACTTCTGGATATACTTTCACCAAGTATTTCATCTACTGGGTGCGACAGGCCCCTGGA CAGGGTCTTGAGTGGCTGGGCAGAATACACCCCCGAACCGGTGCCGTGAAGTATGCACCGAG ATTTCAGGGTAGACTGTCCATGACCAGAGACTGGTCACTCGACACAGCCTACCTCGGATTGAC CGGACTGACACTCGGCGACACGGCTCTATATTTCTGTGCGAGGGGGGCCTTTGAGGCAGATTC ATATGGGTCAAGTTATCCCTTTCACCACTGGGGCCAGGGAACCCTAGTCACCGTCTCCGCGGC CTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCAC AGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTC AGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTC CCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGT GAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAA CTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCC CCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGG ACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCAT AATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCT CACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAG CCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAG GTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCT GGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGA ACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGC TCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAG GCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 339 ##STR00355## ##STR00356## SDFYPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGS TVEKTVAPAECS SEQ ID NO: 340 TCCTGGGCCCTGACTCAACCCGCCTCCGTGTCTGCGTCTCCTGGGCAGTCGGTCACCATGTCCT GCACTGGATTCGGAAATTATAACCCTGACTCCTGGTACCAACAATACCCAGGCAAAGCCCCCA AACTCATCATTTATGAAGACAATAAAAGACCCTCGGGGGTCTCTGATCGCTTCTCTGCCTCCA GACTTGGCAGCACGTCTTCCCTGACAATCTCTAACGTCCAGGCTGCGGACGACGCCCATTATG TCTGCGCCTCCTTTGAATTTTTCGGCGGAGGGACCAAGCTGACCGTCCTGAGTCAGCCCAAGG CTGCCCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACAC TGGTGTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCA GCCCCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCG GCCAGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCG GGTCACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P53 HEAVY CHAIN SEQ ID NO: 341 ##STR00357## ##STR00358## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 342 CGTGTGACATTACAACAATCTGGGGCTACAGTGAAGCAGCCTGGGGCCTCAGTGACCGTCTCC TGCGAGACTTCTGGATACACTTTCACCAAGTATACCATTCACTGGGTGCGACAGGCCCCTGGA CAGGGTCTTCAGTGGGTGGGCAGAATACACCCCCGAACCGGTGCCGTGAAGTATGCACCGAT ATTTCAGGGTAAAGTGTCCATGAGTCGAGACTTGTCACGCGACACAGCCTACCTCGGATTGAC CAGACTGACGCTCGCCGACACGGCTCTATTTTTCTGTGCGAGGGGGGCCTTTGAGGCAGATTT AAGTGGGCCAACTTACCCCTTTCACCACTGGGGCCAAGGAACCCTAGTCATCGTCTCCGCGGC CTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCAC AGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTC AGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTC CCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGT GAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAA CTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCC CCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGG ACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCAT AATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCT CACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAG CCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAG GTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCT GGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGA ACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGC TCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAG GCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 343 ##STR00359## ##STR00360## DFYPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGST
VEKTVAPAECS SEQ ID NO: 344 TCCTGGGCCCTGACTCAACCCGCCTCCGTGTCTGCGTCTCCTGGGCAGTCGGTCACCATGTCCT GCACTGGATTCGGAAATTATAACCCTGACTCCTGGTACCAACAATACCCAGGCAAAGCCCCCA AACTCATCATTTATGAGGACAATAAAAGACCCTCGGGAGTCTCTAATCGCTTCTCTGCCTCCA GACTTGGCAGCACGTCTTCCCTGACAATCTCTAACGTCCAGGCCGCTGACGACGCCCATTATG TCTGCGCCTCCTTTGAATTTTTCGGCGGAGGGACCAAGCTGATCGTCCTGAGTCAGCCCAAGG CTGCCCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACAC TGGTGTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCA GCCCCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCG GCCAGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCG GGTCACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P54 HEAVY CHAIN SEQ ID NO: 345 ##STR00361## ##STR00362## VFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSS SLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPE VTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYK CKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQ PENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 346 AACGTGCAGTTGATGCAGTCTGGGACTGAGGTGAAGAAGTCTGGGGCCTCGGTGACAATCTCT TGTGAGACCGCTGGATTCAACTTCATCGACTCCGTCATACACTGGCTGCGCCAGGCCCCTGGA GGAGGATTTCAGTGGATGGGGTGGATCAAGCCTCTTAGAGGTGCCGTCAATTATCCACAGTTT TTGCAGGGCAGGGTCTCCATGACCCGGGACTTGTCCACCGACACGGTGTACATGGTCTTGAAT GGACTGACACCTGACGACACAGGCCTTTATTACTGCGCGAAAGGGGCCTTTAGAGGGGGTTCT CCCTTTGGCTTCTGGGGCCAGGGAACTCTGCTCACCGTCTCCCCAGCCTCCACCAAGGGCCCA TCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTGC CTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAGC GGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGTG ACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAGC AACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCACC GTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGA CACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAGA CCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAGC CGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCAG GACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCAT CGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCC CATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTAT CCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 347 ##STR00363## ##STR00364## VAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAPA ECS SEQ ID NO: 348 CAGTCTGCCCTGTCTCAGCCTGTCTCCGTGTCTGGGTCTCCTGGAGAGTCGATCACCATTTCCT GTACTGGAGCCACCACCTGGTATCAACAACTCCCAGGCAGACCCCCCAAACTCATCATTTATG ACGTCACTAACCGGCCCTCAGGCATTTCTAGTCGTTTCTCTGGCTCCACGTCTGGCCACACGGC CTCCCTGACAATCTCCGGTCTCCAGGTTGACGACGAGGGTCTGTATCACTGCGCCTCACGTGA ATTTTTCGGCGGAGGGACCAAGCTGACCGTCCTGAGTCAGCCCAAGGCTGCCCCCTCGGTCAC TCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAAG TGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTGG GAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCTG AGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGAAGG GAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P60 HEAVY CHAIN SEQ ID NO: 349 ##STR00365## ##STR00366## VFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSS SLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPE VTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYK CKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQ PENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 350 CAGGTGCGACTGGTGCAGTCTGGGCCTCAGGTGAAGAAGACTGGGGCCTCAGTGAGGGTCTC CTGCGAAACCTCTGGATACACGTTCACCTCCTACTTCATCCATTGGTTACGACTGGGCCCCGG AGAGGGGCTTCAGTGGATGGGGTGGATCAACCCTTTACATGGTGCCGTGAATTATGAAAACA AATTTAGGGGCAGGGTCACAATCACCAGGGACACGTCCACAGACACAGTGTATTTGGACATG AGCAGACTGACCCCTGACGACACGGCCGTCTATTTCTGCACAAGAGGAATCGTTGCTGATGGG TGGCCCTATGGCCACTGGGGCCAGGGAACCCAAGTCACCGTCTCCCCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 351 ##STR00367## ##STR00368## AVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTV APAECS SEQ ID NO: 352 TCCTGGGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGGTCGCCATCTCCT GCGCTGGCGGCAGCGTCTCCTGGTACCAGGTGCTCCCAGGCAGAGCCCCCAAACTCATCATTT ATGAGGGCGCTAAGCGACCCTCAGGGGTTTCTGCTCGCTTCTCTGGCTCCCAGTCTGGCAACA CGGCTTACCTGACAATTTCTGACCTCCAGACTGAGGACGAGGGCATCTACTTCTGCTCTTCACT TCAATTCTTCGGCGGAGGGACCAAACTGACCGTCCTAAGTCAGCCCAAGGCTGCCCCCTCGGT CACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGT AAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGG TGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTAC CTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGA AGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P61 HEAVY CHAIN SEQ ID NO: 353 ##STR00369## ##STR00370## LAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 354 CAGGTGCGACTTCAGCAGTCTGGTGTCGTGGTGAGGAAGCCTGGGGCCTCAGTGAGAATTTCC TGCGAGACTTCTGGATTCACCTTCATCGACCACATTGTCCATTGGGTGCGGCGGGCCCCTGGA CGAGGCTTTGAATGGATGGGTTGGATCAAGCCTCTTAGGGGTGCCGTAGATTATGCACCCCAA CTTCGGGGCAGGATCTCCCTGACGAGGGACATTTACAGTGAAACCGTCTTTATAGACGTGAGC CGACTGACGTCTGGCGACACGGCGATATACTTTTGTTGTAAGGCCGCCGCCCCTGAAGAAGCA TTCCCCCTTCAATACTGGGGCCAGGGGACCCAACTTATCGTCTCCTCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA
GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 355 ##STR00371## ##STR00372## PGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEK TVAPAECS SEQ ID NO: 356 CAGGCTGCCCTGACTCAGCCCGCCTCCGTGTCCGGCTCTCCTGGACAGTCGGTCACCATTTCCT GCCTTTATGCCAATGTAGATATCTGCTGGTATCAACTACACCCGGGCAGAGCCCCCAAACTTC TAATTGTTGACAATAATAAGCGGCCCTCAGGAGTCTCTCCTCGCTTCTCTGGCTCCAAGTCTGG CACCACGGCCTCCCTGACAATCTCTGGACTTCAGGCTGACGACGAGGCTGAATATCACTGCTC TTCAAGAACATTTTTTGGCGGGGGGACCAAGTTGACCGTCCTGAGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P62 HEAVY CHAIN SEQ ID NO: 357 ##STR00373## ##STR00374## PLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 358 CAGGTGCGACTTCAGCAGTCTGGTGTCGTGGTGAGGAAGCCTGGGGCCTCAGTGAGACTTTCC TGCGAGACGTCTGGATTCAAATTCATCGACCACATTGTCAACTGGGTGCGGCGGGCCCCTGGA CGAGGCTTTGAATGGATGGGTTGGATCAAGCCTCTTGGGGGTGTCGCTGATTATGCACCCCAA CATCGGGGCAGGATCTCACTGACGAGGGACATTTACACTGAAACCGTCTTTATAGACCTGAGT CGACTGACGTCTGGCGACACGGCGATTTATTTCTGTTGTAAGGCCGCCGCCCCTGATGAAGCA TTCCCCCTTGAATACTGGGGCCAGGGGACCCAACTTATCGTCTCCCCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 359 ##STR00375## ##STR00376## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 360 CAGGCTGCCCTGACTCAGCCCGCCTCCGTGTCCGGCTCTCCTGGACAGTCGGTCACCATTTCCT GCCTTTATGCCAATGTAGATATCTGCTGGTATCAAATACAGCCGGGCAGATTACCCAAACTTC TGATTGTTGACAATAATAGGCGACCCTCAGGAGTCTCTCCTCGCTTCTCTGGCTCCAAGTCTGG CACCACGGCCTCCCTGACAATCTCTGGACTTCAGGCTGACGACGAGGCTGAATATCACTGCTC TTCAACAACATTTTTTGGCGGGGGGACCAAGTTGACCGTCCTCAGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P63 HEAVY CHAIN SEQ ID NO: 361 ##STR00377## ##STR00378## FPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEV TCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 362 CAGGTGCGACTTGTGCAGTCTGGTCCCGTGATGAGAAAGCCTGGGGCCTCAGTGAGAATTTCT TGCGAGACATCTGGATTCGCCTTCTTGGACCACATTGTCCACTGGGTGCGGCGGGCCCCTGGA CGCGGCTTTGAATGGATGGGTTGGGTTAAGACCATTGGGGGTGTCGTTGATTATGCACCCCAC CTTAGGGGCAGGATCTCCGTGACGAGAGACGTCTTTAGTGAAACCGTCTTTCTGGACTTGAGC CGACTGACGTCTGGCGACACGGCGATGTATTTTTGTTCTAAGGCCGCCGCCCCTGACGAAGCC TTCCCCCTTGAATTTTGGGGCCAGGGGACCCAAGTCATCGTCTCCTCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 363 ##STR00379## ##STR00380## PGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEK TVAPAECS SEQ ID NO: 364 CAGGCTGCCCTGACTCAGCCCGCCTCCGTGTCCGGCTCTCCTGGACAGTCGGTCACCATTTCTT GCCTTTATGCCAATGTGGATATCTGCTGGTATCAACTTCACCCGGGCAGAGCCCCCAAACTTC TTATTCTTGACAATAATAAACGGCCCTCAGGAGTCTCTAGTCGCTTCTCCGGTTCCAAGTCTGG CACCACGGCCTCCCTAACCATCTCTGACCTTCAGGCTGACGACGAGGCTGAATATCACTGCTC TTCAACAACATTTTTTGGCGGGGGGACCAGGTTGACCGTCCTGAGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P64 HEAVY CHAIN SEQ ID NO: 365 ##STR00381## ##STR00382## FPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEV TCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP
ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 366 CAGGTGCGACTTGTGCAGTCTGGTCCCGTGGTGAGAAAGCCTGGGACCTCAGTGAGAATTTCT TGCGAGACATCTGGATTCGCCTTCTTGGACCACATTGTCCACTGGGTGCGGCGGGCCCCTGGA CGCGGCTTTGAATGGATGGGTTGGGTTAAGACCATTGGGGGTGTCGTTGATTATGCACCCCAC CTTAGGGGCAGGATCTCCGTGACGAGGGACGTATTTAGTGAAATCGTCTTTATGGAGTTGAGT CGACTGACGTCTGGCGACACGGCGATGTATTTTTGTTCTAAGGCCGCCGCCCCTGACGAAGCC TTCCCCCTTGAATTTTGGGGCCAGGGGACCCAAGTCATCGTCTCCTCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 367 ##STR00383## ##STR00384## PGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEK TVAPAECS SEQ ID NO: 368 CAGGCTGCCCTGACTCAGCCCGCCTCCGTGTCCGGCTCTCCTGGACAGTCGGTCACCATTTCCT GCCTTTATGCCAATGTGGATATCTGCTGGTATCAACTTCACCCGGGCAGAGCCCCCAAACTTC TAATTGTTGACAATAATAAGCGGCCCTCAGGAGTCTCTAGTCGCTTCTCTGGTTCCAAGTCTGG CACCACGGCCTCCCTAACAATCTCTGATCTTCAGGCTGACGACGAGGCTGAATATCACTGCTC TTCAACAACATTTTTTGGCGGGGGGACCAGGTTGACCGTCCTGAGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P65 HEAVY CHAIN SEQ ID NO: 369 ##STR00385## ##STR00386## VFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSS SLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPE VTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYK CKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQ PENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 370 CAAGTGCAACTGGTGCAGTCTGGGGCTGGGGTGAAGAAGCCTGGGGCCTCAGTGAGGGTCTC CTGCGAGACATCCGGATTCAAGTTCACCGAGTACTTTATCCACTTTTTACGACAGGCCCCTGG ACAAGGGCTTGAGTGGATGGGATGGCTCAACCCTCTCAGAGGTGCCGTCAACTATCCACGGA AGTTTCAGGGCAGAGTCACTTTGACCAGGGACATCTACACCACCACCGTCTACATGCAACTTA ACGGTCTGACCCCTGACGACACGGCCGTCTACTACTGTGCCAGAGCGGTCTTTAATGAAGCTT TCCCCTTTGACTACTGGGGCCAGGGAAGCCTGGTCACCGTCTCCTCAGCCTCCACCAAGGGCC CATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCT GCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCA GCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGG TGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCA GCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCA CCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAG GACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAA GACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAA GCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACC AGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCC ATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCC CCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCT ATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACC ACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAG AGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCAC TACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 371 ##STR00387## ##STR00388## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 372 TCCTGGGCCCAGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCT GCGCTGGAATCGTCAGTGATGCCTGGTACCAGCAATACCCAGGCAGACCCCCCAGACTCATCC TTTATGACGGCGATAAGCGGCCCTCAGGGGTTTCTCCTCGTTTTTCTGCCTCCAGGGCCGGCAA GACGGCCTCCCTGACAATTTCTGGGCTGCAGGCTGACGACGAGGCTTATTATCACTGCGCGTC AAGGGAATTTTTTGGAGGCGTGACCAAGTTGACCGTCCTAAGTCAGCCCAAGGCTGCCCCCTC GGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCT CGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCA AGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAG CTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGC ATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P66 HEAVY CHAIN SEQ ID NO: 373 ##STR00389## ##STR00390## PLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 374 CAGGTGCGACTTCAGCAGTCTGGTGTCGTGGTGAGGAAGCCTGGGGCCTCAGTGAGACTTTCC TGCGAGACGTCTGGCTTCAAATTCATCGACCACATTGTCAACTGGGTGCGGCGGGCCCCTGGA CGAGGCTTTGAATGGATGGGTTGGATCAAGCCTCTTGGGGGTGTCGCTGATTATGCACCCCAA CATCGGGGCAGGATCTCACTGACGAGGGACATTTACACTGAAACCGTCTTTATAGACCTGAGT CGACTGACGTCTGGCGACACGGCGATTTATTTTTGTTGTAAGGCCGCCGCCCCTGATGAAGCA TTCCCCCTTGAATACTGGGGCCAGGGGACCCAACTTATCGTCTCCCCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 375 ##STR00391## ##STR00392## GAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKT VAPAECS SEQ ID NO: 376 CAGGCTGCCCTGACTCAGCCCGCCTCCGTGTCCGGCTCTCCTGGACAGTCGGTCACCATTTCCT GCCTTTATGCCAATGTAGATATCTGCTGGTATCAAATACAGCCGGGCAGATTACCCAAACTTC TGATTGTTGACAATGATAGGCGACCCTCAGGAGTCTCTCCTCGCTTCTCTGGCTCCAAGTCTGG
CACCACGGCCTCCCTGACAATCTCTGGACTTCAGGCTGACGACGAGGCTGAATATCACTGCTC TTCAACAACATTTTTTGGCGGGGGGACCAAGTTGACCGTCCTCAGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P67 HEAVY CHAIN SEQ ID NO: 377 ##STR00393## ##STR00394## FPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEV TCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 378 CAGGTGCGACTTGTGCAGTCTGGTCCCGTGATGAGAAAGCCTGGGGCCTCAGTGAGAATTTCT TGCGAGACATCTGGATTCGCCTTCTTGGACCACATTGTCCACTGGGTGCGGCGGGCCCCTGGA CGCGGCTTTGAATGGATGGGTTGGGTTAAGACCATTGGGGGTGTCGTTGATTATGCACCCCAC CTTAGGGGCAGGATCTCCGTGACGAGAGACGTCTTTAGTGAAACCGTCTTTCTGGACTTGAGT CGACTGACGTCTGGCGACACGGCGATGTATTTTTGTTCTAAGGCCGCCGCCCCTGACGAAGCC TTCCCCCTTGAATTTTGGGGCCAGGGGACCCAAGTCATCGTCTCCTCGGCCTCCACCAAGGGC CCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGC TGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACC AGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTG GTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCC AGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCC ACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGA AGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAA AGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCAC CAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCC CATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGC CCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTC TATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGAC CACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAA GAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACC ACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 379 ##STR00395## ##STR00396## PGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEK TVAPAECS SEQ ID NO: 380 CAGGCTGCCCTGACTCAGCCCGCCTCCGTGTCCGGCTCTCCTGGACAGTCGGTCACCATTTCTT GCCTTTATGCCAATGTGGATATCTGCTGGTATCAACTTCACCCGGGCAGAGCCCCCAAACTTC TAATTCTTGACAATAATAAACGGCCCTCAGGAGTCTCTAGTCGCTTCTCCGGTTCCAAGTCTGG CACCACGGCCTCCCTAACCATCTCTGACCTTCAGGCTGACGACGAGGCTGAATATCACTGCTC TTCAACAACTTTTTTTGGCGGGGGGACCAGGTTGACCGTCCTGAGTCAGCCCAAGGCTGCCCC CTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTG TCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGT CAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGC AGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P68 HEAVY CHAIN SEQ ID NO: 381 ##STR00397## ##STR00398## FPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEV TCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 382 TTGTGAGACCGCTGGATTCAACTTCATCGACTCCGTCATACACTGGCTGCGCCAGGCCCCTGG TGGGGGATTTCAGTGGATGGGGTGGATCAAGCCTCTTAGAGGTGGCGTCAATTATCCACATTA TTTGCAGGGCAGAATCTCCATGACCCGGGACTTGTCCAGTGACACGGTTTACATGGTCTTAAA TAGACTGACACCTGCCGACACAGGCCTTTATTACTGCGCGAAAGGGGCCTTTGGGGGGAGTTC TCCCTTTGGCTTCTGGGGCCAGGGAACTCTGCTCACCGTCTCCCCAGCCTCCACCAAGGGCCC ATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTG CCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAG CGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGT GACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAG CAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCAC CGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGG ACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAG ACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAG CCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCA GGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCA TCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCC CCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTA TCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 383 ##STR00399## ##STR00400## TVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 384 CAGTCTGCCCTGTCTCAGCCTGTCTCCGTGTCTGGGTCTCCTGGAGAGTCGATCACCATTTCCT GTACTGAAGCCACCACCTGGTATCAACAACTCCCAGGCAAACCCCCCAAACTCATCATTTATG ACGTGACCAACCGGCCCTCAGGCATTTCAAGTCGTTTCTCTGGCTCCATGTCTGGTCGCACGG CCTCCCTGACAATCTCCGGTCTCCAGGTTGACGACGAGGGTCTCTATCACTGTGCCTCACGTG AATTTTTCGGCGGGGGGACCAAGCTGACCGTCCTGAGTCAGCCCAAGGCTGCCCCCTCGGTCA CTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAA GTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTG GGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCT GAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGAAG GGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P69 HEAVY CHAIN SEQ ID NO: 385 ##STR00401## ##STR00402## FPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEV TCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 386 CACGTGCAATTGATGCAGTCTGGGACTCAGGCGAAGAAGTCTGGGGCCTCGGTGACAATTTCT TGTGAGACCGCTGGATTCAAGTTCATCGACTCCGTCATACACTGGCTGCGCCAGGCCCCTGGA GGGGGATTTCAGTGGATGGGGTGGATCAAGCCTCTTGGAGGTGCCGTCAACTATCCACCCTAT TTGCAGGGCAGGATCTCCTTGACCCGTGACTTGTCCACCGACACAATTTACATGGTCTTGAAT GGACTGACACCTGCCGACACAGGCTTTTATTACTGCGCCAAAGGGGCCTTTGGGGGGGGTTCT CCCTTTGGCTTCTGGGGCCAGGGGACTCTGCTCACCGTCTCCCCAGCCTCCACCAAGGGCCCA TCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTGC CTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAGC GGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGTG ACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAGC AACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCACC GTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGA CACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAGA CCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAGC CGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCAG GACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCAT CGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCC
CATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTAT CCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 387 ##STR00403## ##STR00404## TVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 388 CAGTCTGCCCTGTCTCAGCCTGTCTCCGTGTCTGGGTCTCCTGGAGACTCGATCACCATTTCTT GTTTTGGAGCCACCACCTGGTATCAACAACTCCCAGGCAGACCCCCCAAACTCATCATTTATG ACGTGACTAACCGGCCCTCAGGCATTTCAGGTCGTTTCTCTGGCTCCATGTCTGGTCAAAAGG CCTCCCTGACAATCTCCGGTCTCCAGGTTGACGACGAGGGTCTCTATCACTGCGCCTCACGTG AATTTTTCGGCGGGGGGACCAAACTGACCGTCCTGAGTCAGCCCAAGGCTGCCCCCTCGGTCA CTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAA GTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTG GGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCT GAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGAAG GGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P70 HEAVY CHAIN SEQ ID NO: 389 ##STR00405## ##STR00406## FPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEV TCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 390 CACGTGCAGTTGAGGCAGTCTGGGACTGAGGCGAAGAAGTCTGGGGCCTCGGTGACAATCTC TTGTGAGACCGCTGGATTCAACTTCATCGACTCCGTCATACACTGGCTGCGCCAGGCCCCTGG TGGGGGATTTCAGTGGATGGGGTGGATCAAGCCTCTTAGAGGTGGCGTCAATTATCCACATTA TTTGCAGGGCAGAATCTCCATGACCCGGGACTTGTCCAGTGACACGGTTTACATGGTCTTAAA TAGACTGACACCTGACGACACAGGCCTTTACTACTGCGCGAAAGGGGCCTTTGGGGGGAGTTC TCCCTTTGGCTTCTGGGGCCAGGGAACTCTGCTCACCGTCTCCCCAGCCTCCACCAAGGGCCC ATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTG CCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAG CGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGT GACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAG CAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCAC CGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGG ACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAG ACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAG CCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCA GGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCA TCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCC CCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTA TCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 391 ##STR00407## ##STR00408## TVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAP AECS SEQ ID NO: 392 CAGTCTGCCCTGTCTCAGCCTGTCTCCGTGTCTGGGTCTCCTGGAGAGTCGATCACCATTTCCT GTACTGAAGCCACCACCTGGTATCAACAACTCCCAGGGAGATCCCCCAAACTCATTATTTATG ACGTGACCAACCGGCCCTCAGGCATTTCAAGTCGTTTCTCTGGCTCCATGTCTGGTCGCACGG CCTCCCTGACAATCTCCGGTCTCCAGGTTGACGACGAGGGTCTCTATCACTGTGCCTCACGTG AATTTTTCGGCGGGGGGACCAAGCTGACCGTCCTCAGTCAGCCCAAGGCTGCCCCCTCGGTCA CTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAA GTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTG GGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCT GAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGAAG GGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P71 HEAVY CHAIN SEQ ID NO: 393 ##STR00409## ##STR00410## KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV TVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMI SRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNG KEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 394 CGTGTTACATTACAACAGTCTGGGGCTACAGTGAGGCAGCCTGGGGCCTCAGTCACCGTCTCC TGCGAGACTTCTGGATTCACCTTCATCAAATATACCATTCACTGGGTGCGACAGGCCCCTGGA CAGGGTCTTCAGTGGGTGGGAAGAATACACCCCCGAACCGGTGCCGTGAAGTTTGCACCGAT ATTTCAGGGTAAATTTTCCATGAGTCGAGACTTGTCACGCGACACAGCCTACCTCGGATTGAC CAGACTGACACTCGCCGACACGGCTCTATTTTTCTGTGCGAGGGGGGCCTTTGAGGCAGATTT ATATGGGCCAACTTACCCCTTTCACCACTGGGGCCAAGGAACCCAAGTCACCGTCTCCGCGGC CTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCAC AGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTC AGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTC CCTCAGCAGCGTGGTGACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGT GAATCACAAGCCCAGCAACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAA CTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCC CCCCAAAACCCAAGGACACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGG ACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCAT AATGCCAAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCT CACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAG CCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAG GTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCT GGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGA ACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGC TCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAG GCTCTGCACAACCACTACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 395 ##STR00411## ##STR00412## SDFYPGAVTVAWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGS TVEKTVAPAECS SEQ ID NO: 396 TCCTGGGCCCTGACTCAACCCGCCTCCGTGTCTGCGTCTCCTGGGCAGTCGGTCACCATGTCCT GCACTGGATTCGGAAGTTATAATCCTGACTCCTGGTACCAGCAATACCCAGGCAAAGCCCCCA AACTCATCATTTATGATGACAATAAAAGACCCTCGGGGGTCTCTGATCGCTTCTCTGCCTCCA GACTTGGCAGCACATCTTCACTGACAATCTCTAACGTCCAGGCCGCTGACGACGCCCATTATG TCTGCGCCTCCTTTGAGTTTTTCGGCGGGGGGACCAAGCTGACCGTCCTGAGTCAGCCCAAGG CTGCCCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACAC TGGTGTGTCTCGTAAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCA GCCCCGTCAAGGTGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCG GCCAGCAGCTACCTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCG GGTCACGCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT >N49P72 HEAVY CHAIN SEQ ID NO: 397 ##STR00413## ##STR00414## PLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKRVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVT CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 398 CACATTCAGTTGCTACAGTCGGGGCCTCAAGTTAAGAAGTCTGGGGACACAGTGAGAATCTCC TGTGAGACCTCTGGATATAATTTCGTCGACTCCCTTATCCACTGGGTCCGACAGACCCCGGAA AAACGTCTCAGATGGATGGGCTGGATCAATCCTCTCCAAGGTGGTGTGAATTACGCGCCGGAA
TTTCAGGGCAGAATCAGGATGACCAGGGACACGTTTATAGACACAGTTTACGTGGACTTGAGC GGACTGACACCGGCCGACACGGCCTATTATTACTGCGCGCGAGGGATCGATGGCAATTCTTAC CCCTTTCATTTCTGGGGCCACGGAACCCGGGTCACCGTCTTCTCGGCCTCCACCAAGGGCCCA TCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTGC CTGGTCAAGGACTACTTCCCCGAACCGGTGACGGTGTCGTGGAACTCAGGCGCCCTGACCAGC GGCGTGCACACCTTCCCGGCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGTG ACCGTGCCCTCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAGC AACACCAAGGTGGACAAGAGAGTTGAGCCCAAATCTTGTGACAAAACTCACACATGCCCACC GTGCCCAGCACCTGAACTCCTGGGGGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGA CACCCTCATGATCTCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCACGAAGA CCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAGC CGCGGGAGGAGCAGTACAACAGCACGTACCGGGTGGTCAGCGTCCTCACCGTCCTGCACCAG GACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCAT CGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCC CATCCCGGGAGGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTAT CCCAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCA CGCCTCCCGTGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGA GCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACT ACACGCAGAAGAGCCTCTCCCTGTCTCCGGGTAAA LIGHT CHAIN SEQ ID NO: 399 ##STR00415## ##STR00416## AWKADGSPVKVGVETTKPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAPAE CS SEQ ID NO: 400 CGATTTGCCCTGACTCAACCTGCCTCCGTGTCTGGGTCTCCTGGACAGACGATCACCATAACCT GCGCTGGAGGCAGCGTCTCCTGGTTCCATTTCCCTCCAGGCAAAACCCCCAGACTCATTATTT ATGAGTCTTCTAAGAGACCCTCAGGGGTCTCTCCTCGATTCTCTGGGTCCCAGTCTGGCAGCA CGGCCTCCCTAATAATTTCTGGCCTCCAGTCTGATGACGAAGGGACATACTTCTGTTCTATTCT TGAATTTTTCGGCAGAGGGACTCTTCTCACCGTCCTGAGTCAGCCCAAGGCTGCCCCCTCGGT CACTCTGTTCCCACCCTCCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGT AAGTGACTTCTACCCGGGAGCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGG TGGGAGTGGAGACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTAC CTGAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCACGCATGA AGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT
Polypeptides
[0649] In some embodiments, the invention provides isolated polypeptides comprising an individual light chain or heavy chain described herein as well as antigen binding fragments thereof. Polypeptides (e.g., intact antibodies) comprising both a light chain and a heavy chain are also provided.
[0650] Also provided are polypeptides that comprise: a polypeptide comprising SEQ ID NOS:1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73, 75, 153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 or 397 or an antigen binding fragment thereof.
[0651] Also provided are polypeptides that comprise: a polypeptide comprising SEQ ID NOS:2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 or 399 or an antigen binding fragment thereof.
[0652] Also provided are polypeptides that comprise: a polypeptide having at least about 90% sequence identity to SEQ ID NOS:1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73, 75, 153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 or 397.
[0653] In some embodiments, the polypeptide comprises a polypeptide having at least about 95%, at least about 96%, at least about 97%, at least about 98%, or at least about 99% sequence identity to SEQ ID NOS: 1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59, 61, 63, 65, 67, 69, 71, 73, 75, 153, 157, 161, 165, 169, 173, 177, 181, 185, 189, 193, 197, 201, 205, 209, 213, 217, 221, 225, 229, 233, 237, 241, 245, 249, 253, 257, 261, 265, 269, 273, 277, 281, 285, 289, 293, 297, 301, 305, 309, 313, 317, 321, 325, 329, 333, 337, 341, 345, 349, 353, 357, 361, 365, 369, 373, 377, 381, 385, 389, 393 or 397.
[0654] Also provided are polypeptides that comprise: a polypeptide having at least about 90% sequence identity to SEQ ID NOS:2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 or 399.
[0655] In some embodiments, the polypeptide comprises a polypeptide having at least about 95%, at least about 96%, at least about 97%, at least about 98%, or at least about 99% sequence identity to SEQ ID NOS: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58, 60, 62, 64, 66, 68, 70, 72, 74, 76, 155, 159, 163, 167, 171, 175, 179, 183, 187, 191, 195, 199, 203, 207, 211, 215, 219, 223, 227, 231, 235, 239, 243, 247, 251, 255, 259, 263, 267, 271, 275, 279, 283, 287, 291, 295, 299, 303, 307, 311, 315, 319, 323, 327, 331, 335, 339, 343, 347, 351, 355, 359, 363, 367, 371, 375, 379, 383, 387, 391, 395 or 399.
Polynucleotides
[0656] In some embodiments, the invention encompasses polynucleotides comprising polynucleotides that encode a polypeptide as described herein, such as a heavy chain or light chain sequence of an HIV antibody or a fragment of such a polypeptide. For example, the invention provides a polynucleotide comprising a nucleic acid sequence that encodes an antibody to gp120 or encodes a fragment of such an antibody. The polynucleotides of the invention can be in the form of RNA or in the form of DNA. DNA includes cDNA, genomic DNA, and synthetic DNA; and can be double-stranded or single-stranded, and if single stranded can be the coding strand or non-coding (anti-sense) strand.
[0657] In some embodiments, the polynucleotides are isolated. In certain embodiments, the polynucleotides are substantially pure.
[0658] In some embodiments, the invention provides a polynucleotide comprising a polynucleotide encoding a polypeptide comprising a sequence selected from the group consisting of SEQ ID NOS:1-76, 153, 155, 157, 159, 161, 163, 165, 167, 169, 171, 173, 175, 177, 179, 181, 183, 185, 187, 189, 191, 193, 195, 197, 199, 201, 203, 205, 207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227,229, 231, 233, 235, 237, 239, 241, 243, 245, 247, 249, 251, 253, 255, 257, 259, 261, 263, 265, 267, 269, 271, 273, 275, 277, 279, 281, 283, 285, 287, 289, 291, 293, 295, 297, 299, 301, 303, 305, 307, 309, 311, 313, 315, 317, 319, 321, 323, 325, 327, 329, 331, 333, 335, 337, 339, 341, 343, 345, 347, 349, 351, 353, 355, 357, 359, 361, 363. 365, 367, 369, 371, 373, 375, 377, 379, 381, 383, 385, 387, 389, 391, 393, 395 and 397.
[0659] In some embodiments, the invention provides a polynucleotide comprising a polynucleotide encoding a polypeptide comprising the heavy or light chain variable region found within a sequence selected from the group consisting of SEQ ID NOS:1-76, 153, 155, 157, 159, 161, 163, 165, 167, 169, 171, 173, 175, 177, 179, 181, 183, 185, 187, 189, 191, 193, 195, 197, 199, 201, 203, 205, 207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227,229, 231, 233, 235, 237, 239, 241, 243, 245, 247, 249, 251, 253, 255, 257, 259, 261, 263, 265, 267, 269, 271, 273, 275, 277, 279, 281, 283, 285, 287, 289, 291, 293, 295, 297, 299, 301, 303, 305, 307, 309, 311, 313, 315, 317, 319, 321, 323, 325, 327, 329, 331, 333, 335, 337, 339, 341, 343, 345, 347, 349, 351, 353, 355, 357, 359, 361, 363. 365, 367, 369, 371, 373, 375, 377, 379, 381, 383, 385, 387, 389, 391, 393, 395 and 397.
[0660] Also provided is a polynucleotide encoding a polypeptide having at least about 95%, at least about 96%, at least about 97%, at least about 98%, or at least about 99% sequence identity to SEQ ID NOS:1-76, 153, 155, 157, 159, 161, 163, 165, 167, 169, 171, 173, 175, 177, 179, 181, 183, 185, 187, 189, 191, 193, 195, 197, 199, 201, 203, 205, 207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227,229, 231, 233, 235, 237, 239, 241, 243, 245, 247, 249, 251, 253, 255, 257, 259, 261, 263, 265, 267, 269, 271, 273, 275, 277, 279, 281, 283, 285, 287, 289, 291, 293, 295, 297, 299, 301, 303, 305, 307, 309, 311, 313, 315, 317, 319, 321, 323, 325, 327, 329, 331, 333, 335, 337, 339, 341, 343, 345, 347, 349, 351, 353, 355, 357, 359, 361, 363. 365, 367, 369, 371, 373, 375, 377, 379, 381, 383, 385, 387, 389, 391, 393, 395 or 397.
[0661] The invention further provides a polynucleotide comprising a sequence selected from the group consisting of SEQ ID NOS:77-152, 154, 156, 158, 160, 162, 164, 166, 168, 170, 172, 174, 176, 178, 180, 182, 184, 186, 188, 190, 192, 194, 196, 198, 200 202, 204, 206, 208, 210, 212, 214, 216, 218, 220, 222, 224, 226, 228, 230, 232, 234, 236, 238, 240, 242, 244, 246, 248, 250, 252, 254, 256, 258, 260, 262, 264, 266, 268, 270, 272, 274, 276, 278, 280, 282, 284, 286, 288, 290, 292, 294, 296, 298, 300, 302, 304, 306, 308, 310, 312, 314, 316, 318, 320, 322, 324, 326, 328, 330, 332, 334, 336, 338, 340, 342, 344, 346, 348, 350, 352, 354, 356, 358, 360, 362, 364, 366, 368, 370, 372, 374, 376, 378, 380, 382, 384, 386, 388, 390, 392, 394, 396, 398 and 400.
[0662] Also provided is a polynucleotide having at least 70%, 75%, 80%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NOS:77-152, 154, 156, 158, 160, 162, 164, 166, 168, 170, 172, 174, 176, 178, 180, 182, 184, 186, 188, 190, 192, 194, 196, 198, 200 202, 204, 206, 208, 210, 212, 214, 216, 218, 220, 222, 224, 226, 228, 230, 232, 234, 236, 238, 240, 242, 244, 246, 248, 250, 252, 254, 256, 258, 260, 262, 264, 266, 268, 270, 272, 274, 276, 278, 280, 282, 284, 286, 288, 290, 292, 294, 296, 298, 300, 302, 304, 306, 308, 310, 312, 314, 316, 318, 320, 322, 324, 326, 328, 330, 332, 334, 336, 338, 340, 342, 344, 346, 348, 350, 352, 354, 356, 358, 360, 362, 364, 366, 368, 370, 372, 374, 376, 378, 380, 382, 384, 386, 388, 390, 392, 394, 396, 398, or 400.
[0663] In some embodiments the polynucleotides comprise the coding sequence for the mature polypeptide fused in the same reading frame to a polynucleotide which aids, for example, in expression and secretion of a polypeptide from a host cell (e.g. a leader sequence which functions as a secretory sequence for controlling transport of a polypeptide from the cell). The polypeptide having a leader sequence is a preprotein and can have the leader sequence cleaved by the host cell to form the mature form of the polypeptide. The polynucleotides can also encode for a proprotein which is the mature protein plus additional 5' amino acid residues. A mature protein having a prosequence is a proprotein and is an inactive form of the protein. Once the prosequence is cleaved an active mature protein remains.
[0664] In certain embodiments the polynucleotides comprise the coding sequence for the mature polypeptide fused in the same reading frame to a marker sequence that allows, for example, for purification of the encoded polypeptide. For example, the marker sequence can be a hexa-histidine tag supplied by a pQE-9 vector to provide for purification of the mature polypeptide fused to the marker in the case of a bacterial host, or the marker sequence can be a hemagglutinin (HA) tag derived from the influenza hemagglutinin protein when a mammalian host (e.g. COS-7 cells) is used.
[0665] The present invention further relates to variants of the hereinabove described polynucleotides encoding, for example, fragments, analogs, and derivatives.
[0666] The polynucleotide variants can contain alterations in the coding regions, non-coding regions, or both. In some embodiments the polynucleotide variants contain alterations which produce silent substitutions, additions, or deletions, but do not alter the properties or activities of the encoded polypeptide. In some embodiments, nucleotide variants are produced by silent substitutions due to the degeneracy of the genetic code. Polynucleotide variants can be produced for a variety of reasons, e.g., to optimize codon expression for a particular host (change codons in the human mRNA to those preferred by a bacterial host such as E. coli).
[0667] Vectors and cells comprising the polynucleotides described herein are also provided. The term "vector" means a construct, which is capable of delivering, and expressing, one or more gene(s) or sequence(s) of interest in a host cell. Examples of vectors include, but are not limited to, viral vectors, naked DNA or RNA expression vectors, plasmid, cosmid or phage vectors, DNA or RNA expression vectors associated with cationic condensing agents, DNA or RNA expression vectors encapsulated in liposomes, and certain eukaryotic cells, such as producer cells. "Vector" also includes shuttle and expression vectors. In some embodiments, the vector is a plasmid construct and also includes an origin of replication (e.g., the Co1E1 origin of replication) and a selectable marker (e.g., ampicillin or tetracycline resistance), for replication and selection, respectively. An "expression vector" refers to a vector that contains the necessary control sequences or regulatory elements for expression of the antibodies including antibody fragments of the invention, in bacterial or eukaryotic cells.
Methods
[0668] The anti-HIV antibodies of the invention are useful in a variety of applications including, but not limited to, therapeutic treatment methods, such as the treatment, cure, functional cure, or prevention of HIV infection. The methods of use may be in vitro, ex vivo, or in vivo methods.
[0669] In some embodiments, the antibodies disclosed herein may be used as neutralizing antibodies, passively administered or given via gene therapies.
[0670] In one aspect, the anti-HIV antibodies are useful for detecting the presence of HIV in a biological sample. The term "detecting" as used herein encompasses quantitative or qualitative detection. In certain embodiments, a biological sample comprises a cell or tissue.
[0671] Certain other methods can be used to detect binding of anti-HIV antibodies to antigens such as gp120. Such methods include, but are not limited to, antigen-binding assays that are well known in the art, such as western blots, radioimmunoassays, ELISA (enzyme linked immunosorbent assay), "sandwich" immunoassays, immunoprecipitation assays, fluorescent immunoassays, protein A immunoassays, and immunohistochemistry (IHC).
[0672] In certain embodiments, the antibodies are labeled. Labels include, but are not limited to, labels or moieties that are detected directly (such as fluorescent, chromophoric, electron-dense, chemiluminescent, and radioactive labels), as well as moieties, such as enzymes or ligands, that are detected indirectly, e.g., through an enzymatic reaction or molecular interaction.
[0673] In certain embodiments, the antibodies are immobilized on an insoluble matrix. Immobilization entails separating the antibody from any antigen that remains free in solution. This conventionally is accomplished by either insolubilizing the antibody before the assay procedure, as by adsorption to a water-insoluble matrix or surface (Bennich et al., U.S. Pat. No. 3,720,760), or by covalent coupling (for example, using glutaraldehyde cross-linking), or by insolubilizing the antibody after formation of a complex between the antibody and antigen, e.g., by immunoprecipitation.
[0674] The present invention provides for methods of treating or preventing HIV infection comprising administering a therapeutically effective amount of an antibody as described herein to a subject (e.g., a subject in need of treatment). In some embodiments, the subject is a human.
[0675] Subjects at risk for HIV-related diseases or disorders include patients who have come into contact with an infected person or who have been exposed to HIV-1 in some other way. Administration of a prophylactic agent can occur prior to the manifestation of symptoms characteristic of HIV-1-related disease or disorder, such that a disease or disorder is prevented or, alternatively, delayed in its progression.
[0676] In some embodiments of the present invention, the subject is administered effective amounts of more than one anti-HIV antibody of the invention. In some embodiments, the subject is administered a pharmaceutical composition comprising a combination of antibodies of the invention, in order to treat or prevent HIV infection. In some embodiments, a combination of antibodies are administered, which can include a combination comprising any one or more of N49P6 or an antigen binding fragment thereof, N49P7 or an antigen binding fragment thereof, N49P7.1 or an antigen binding fragment thereof, N49P9 or an antigen binding fragment thereof, or N49P11 or an antigen binding fragment thereof. In some embodiments, the antibody comprises the VH and VL regions of N49P6, N49P7, N49P7.1, N49P9, or N49P11 as described herein. In some embodiments, the antibody comprises the CDRs of the VH and V.sub.L regions of N49P6, N49P7, N49P7.1, N49P9, or N49P11 as described herein. In some embodiments, the combination comprises i) N49P6 or an antigen binding fragment thereof, ii) N49P7 or an antigen binding fragment thereof and iii) N49P11 or an antigen binding fragment thereof. In some embodiments, the subject is administered a polyclonal composition of antibodies comprising any one of i) N49P6 or an antigen binding fragment thereof, ii) N49P7 or an antigen binding fragment thereof and/or iii) N49P11 or an antigen binding fragment thereof in combination with one or more natural or variant antibodies as described herein. Such combinations can be selected according to the desired immunity. The composition can further include one or more other broadly neutralizing antibodies.
[0677] Methods for preventing an increase in HIV-1 virus titer, virus replication, virus proliferation or an amount of an HIV-1 viral protein in a subject are further provided. In one embodiment, a method includes administering to the subject an amount of an anti-HIV antibody effective to prevent an increase in HIV-1 titer, virus replication or an amount of an HIV-1 protein of one or more HIV strains or isolates in the subject.
[0678] For in vivo treatment of human patients, the patient is usually administered or provided a pharmaceutical formulation including an anti-HIV antibody of the invention. When used for in vivo therapy, the antibodies of the invention are administered to the patient in therapeutically effective amounts (i.e., amounts that eliminate or reduce the patient's viral burden). The antibodies can be administered to a human patient, in accord with known methods, such as intravenous administration, e.g., as a bolus or by continuous infusion over a period of time, by intramuscular, intraperitoneal, intracerobrospinal, subcutaneous, intra-articular, intrasynovial, intrathecal, oral, topical, or inhalation routes. The antibodies may be administered parenterally, when possible, at the target cell site, or intravenously. Intravenous or subcutaneous administration of the antibody is preferred in certain embodiments. Therapeutic compositions of the invention are administered to a patient or subject systemically, parenterally, or locally.
[0679] For parenteral administration, the antibodies can be formulated in a unit dosage injectable form (solution, suspension, emulsion) in association with a pharmaceutically acceptable, parenteral vehicle. Examples of such vehicles are water, saline, Ringer's solution, dextrose solution, and 5% human serum albumin. Nonaqueous vehicles such as fixed oils and ethyl oleate are also used. Liposomes are used as carriers. The vehicle contains minor amounts of additives such as substances that enhance isotonicity and chemical stability, e.g., buffers and preservatives. The antibodies are typically formulated in such vehicles at concentrations of about 1 mg/ml to 10 mg/ml.
[0680] The dose and dosage regimen depends upon a variety of factors readily determined by a physician, such as the nature of the infection and the characteristics of the particular cytotoxic agent or growth inhibitory agent conjugated to the antibody (when used), e.g., its therapeutic index, the patient, and the patient's history. Generally, a therapeutically effective amount of an antibody is administered to a patient. In particular embodiments, the amount of antibody administered is in the range of about 0.1 mg/kg to about 20 mg/kg of patient body weight. Depending on the type and severity of the infection, about 0.1 mg/kg to about 20 mg/kg body weight (e.g., about 0.1-15 mg/kg/dose) of antibody is an initial candidate dosage for administration to the patient, whether, for example, by one or more separate administrations, or by continuous infusion. The progress of this therapy is readily monitored by conventional methods and assays and based on criteria known to the physician or other persons of skill in the art.
[0681] Antibodies of the invention can be coupled to a drug for delivery to a treatment site or coupled to a detectable label to facilitate imaging of a site comprising cells of interest, such as cells infected with HIV. Methods for coupling antibodies to drugs and detectable labels are well known in the art, as are methods for imaging using detectable labels. Labeled antibodies may be employed in a wide variety of assays, employing a wide variety of labels. Detection of the formation of an antibody-antigen complex between an antibody of the invention and an epitope of interest (an HIV epitope) can be facilitated by attaching a detectable substance to the antibody. Suitable detection means include the use of labels such as radionucleotides, enzymes, coenzymes, fluorescers, chemiluminescers, chromogens, enzyme substrates or co-factors, enzyme inhibitors, prosthetic group complexes, free radicals, particles, dyes, and the like. Examples of suitable enzymes include horseradish peroxidase, alkaline phosphatase, .beta.-galactosidase, or acetylcholinesterase; examples of suitable prosthetic group complexes include streptavidin/biotin and avidin/biotin; examples of suitable fluorescent materials include umbelliferone, fluorescein, fluorescein isothiocyanate, rhodamine, dichlorotriazinylamine fluorescein, dansyl chloride or phycoerythrin; an example of a luminescent material is luminol; examples of bioluminescent materials include luciferase, luciferin, and aequorin; and examples of suitable radioactive material include .sup.125I, .sup.131I, .sup.35S, or .sup.3H. Such labeled reagents may be used in a variety of well-known assays, such as radioimmunoassays, enzyme immunoassays, e.g., ELISA, fluorescent immunoassays, and the like.
[0682] The antibodies can be tagged with such labels by known methods. For instance, coupling agents such as aldehydes, carbodiimides, dimaleimide, imidates, succinimides, bid-diazotized benzadine and the like are used to tag the antibodies with the above-described fluorescent, chemiluminescent, and enzyme labels. An enzyme is typically combined with an antibody using bridging molecules such as carbodiimides, periodate, diisocyanates, glutaraldehyde and the like. Various labeling techniques are described in Morrison, Methods in Enzymology 32b, 103 (1974), Syvanen et al., J. Biol. Chem. 284, 3762 (1973) and Bolton and Hunter, Biochem J. 133, 529(1973).
[0683] In one embodiment, the antibodies can be administered as immunoconjugates, conjugated to a second molecule. For example, the second molecule can be a toxin, a label, a radioisotope, a drug, or a chemical compound.
[0684] An antibody according to the invention may be conjugated to a therapeutic moiety such as a cytotoxin, a therapeutic agent, or a radioactive metal ion or radioisotope. Examples of radioisotopes include, but are not limited to, I-131, I-123, I-125, Y-90, Re-188, Re-186, At-211, Cu-67, Bi-212, Bi-213, Pd-109, Tc-99, In-111, and the like. Such antibody conjugates can be used for modifying a given biological response; the drug moiety is not to be construed as limited to classical chemical therapeutic agents. For example, the drug moiety may be a protein or polypeptide possessing a desired biological activity. Such proteins may include, for example, a toxin such as abrin, ricin A, pseudomonas exotoxin, or diphtheria toxin, TLR agonists (such as TLR7 agonist), or monomethylauristatin E.
[0685] Other therapeutic regimens can be combined with the administration of the anti-HIV antibody of the present invention. The combined administration includes co-administration, using separate formulations or a single pharmaceutical formulation, and consecutive administration in either order, wherein preferably there is a time period while both (or all) active agents simultaneously exert their biological activities. Preferably such combined therapy results in a synergistic therapeutic effect.
[0686] For any application, the antibody, antigen binding fragment, or nucleic acid encoding the antibody or antigen binding fragment can be combined with anti-retroviral therapy. Antiretroviral drugs are broadly classified by the phase of the retrovirus life-cycle that the drug inhibits. The disclosed antibodies can be administered in conjunction with nucleoside analog reverse-transcriptase inhibitors (such as zidovudine, didanosine, zalcitabine, stavudine, lamivudine, abacavir, emtricitabine, entecavir, and apricitabine), nucleotide reverse transcriptase inhibitors (such as tenofovir and adefovir), non-nucleoside reverse transcriptase inhibitors (such as efavirenz, nevirapine, delavirdine, etravirine, and rilpivirine), protease inhibitors (such as saquinavir, ritonavir, indinavir, nelfinavir, amprenavir, lopinavir, fosamprenavir, atazanavir, tipranavir, and darunavir), entry or fusion inhibitors (such as maraviroc and enfuvirtide), maturation inhibitors, (such as bevirimat and vivecon), or a broad spectrum inhibitors, such as natural antivirals. In some examples, a disclosed antibody or active fragment thereof or nucleic acids encoding such is administered in conjunction with IL-15, or conjugated to IL-15.
[0687] Single or multiple administrations of the compositions including the antibody, antigen binding fragment, or nucleic acid encoding the antibody or antigen binding fragment, that are disclosed herein, are administered depending on the dosage and frequency as required and tolerated by the patient. In any event, the composition should provide a sufficient quantity of at least one of the antibodies disclosed herein to effectively treat the patient. The dosage can be administered once, but may be applied periodically until either a therapeutic result is achieved or until side effects warrant discontinuation of therapy.
[0688] One approach to administration of nucleic acids is direct administration with plasmid DNA, such as with a mammalian expression plasmid. The nucleotide sequence encoding the disclosed antibody, or antibody binding fragments thereof, can be placed under the control of a promoter to increase expression. Another approach is to administer the nucleic acids in the form of mRNA.
[0689] In some embodiments, the subject is administered cells that are engineered to express the anti-HIV antibody. In some embodiments, the cells are engineered immune cells, such as B cells. In some embodiments, the cells are engineered, autologous cells.
[0690] In another approach to using nucleic acids, an anti-HIV antibody, or antibody binding fragment thereof can also be expressed by attenuated viral hosts or vectors or bacterial vectors. Recombinant vaccinia virus, adeno-associated virus (AAV), herpes virus, retrovirus, cytomegalovirus or other viral vectors can be used to express the antibody. For example, vaccinia vectors and methods useful protocols are described in U.S. Pat. No. 4,722,848. BCG (Bacillus Calmette Guerin) provides another vector for expression of the disclosed antibodies (see Stover, Nature 351:456-460, 1991).
Compositions
[0691] The present invention also encompasses compositions comprising one or more antibodies of the invention. In certain embodiments, the compositions are pharmaceutical compositions. In some embodiments, formulations are prepared for storage and use by combining an antibody with a pharmaceutically acceptable vehicle (e.g. carrier, excipient) (Remington, The Science and Practice of Pharmacy 20th Edition Mack Publishing, 2000). Suitable pharmaceutically acceptable vehicles include, but are not limited to, nontoxic buffers such as phosphate, citrate, and other organic acids; salts such as sodium chloride; antioxidants including ascorbic acid and methionine; preservatives (e.g. octadecyldimethylbenzyl ammonium chloride; hexamethonium chloride; benzalkonium chloride; benzethonium chloride; phenol, butyl or benzyl alcohol; alkyl parabens, such as methyl or propyl paraben; catechol; resorcinol; cyclohexanol; 3-pentanol; and m-cresol); low molecular weight polypeptides (e.g. less than about 10 amino acid residues); proteins such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidone; amino acids such as glycine, glutamine, asparagine, histidine, arginine, or lysine; carbohydrates such as monosacchandes, disaccharides, glucose, mannose, or dextrins; chelating agents such as EDTA; sugars such as sucrose, mannitol, trehalose or sorbitol; salt-forming counter-ions such as sodium; metal complexes (e.g. Zn-protein complexes); and non-ionic surfactants such as TWEEN or polyethylene glycol (PEG).
[0692] For the treatment or prevention of HIV, the appropriate dosage of an antibody or combination of antibodies of the present invention can depend on a variety of factors, such as the severity and course of the disease, the responsiveness of the disease, whether the antibody or agent is administered for therapeutic or preventative purposes, previous therapy, patient's clinical history, and so on all at the discretion of the treating physician. The antibody or agent can be administered one time or over a series of treatments lasting from several days to several months, or until a cure is effected or a diminution of the disease state is achieved. The administering physician can easily determine optimum dosages, dosing methodologies and repetition rates. In certain embodiments, dosage is from 0.01 .mu.g to 100 mg per kg of body weight, and can be given once or more daily, weekly, monthly or yearly. In certain embodiments, the antibody or combination of antibodies is given once every two weeks or once every three weeks. In certain embodiments, the dosage of the antibody is from about 0.1 mg to about 20 mg per kg of body weight. The treating physician can estimate repetition rates for dosing based on measured residence times and concentrations of the drug in bodily fluids or tissues.
[0693] Effective dosages and schedules for administering embodiments of the present invention can be determined empirically. In some embodiments, and effective amount of one or more antibodies are administered to neutralize, treat, prevent or eradicate HIV infection. In some embodiments, compositions comprising one or more nucleic acid molecules of the invention are administered to the subject. In some embodiments, genetic constructs capable of inducing production of antibodies of the present invention may be administered to a patient in need thereof.
[0694] Controlled-release parenteral formulations can be made as implants, oily injections, or as particulate systems. For a broad overview of protein delivery systems see, Banga, A. J., Therapeutic Peptides and Proteins: Formulation, Processing, and Delivery Systems, Technomic Publishing Company, Inc., Lancaster, Pa., (1995). Particulate systems include microspheres, microparticles, microcapsules, nanocapsules, nanospheres, and nanoparticles. Microcapsules contain the therapeutic protein, such as a cytotoxin or a drug, as a central core. In microspheres the therapeutic is dispersed throughout the particle. Particles, microspheres, and microcapsules smaller than about 1 .mu.m are generally referred to as nanoparticles, nanospheres, and nanocapsules, respectively. Capillaries have a diameter of approximately 5 .mu.m so that only nanoparticles are administered intravenously. Microparticles are typically around 100 .mu.m in diameter and are administered subcutaneously or intramuscularly. See, for example, Kreuter, J., Colloidal Drug Delivery Systems, J. Kreuter, ed., Marcel Dekker, Inc., New York, N.Y., pp. 219-342 (1994); and Tice & Tabibi, Treatise on Controlled Drug Delivery, A. Kydonieus, ed., Marcel Dekker, Inc. New York, N.Y., pp. 315-339, (1992).
[0695] Polymers can be used for ion-controlled release of the antibody compositions disclosed herein. Various degradable and nondegradable polymeric matrices for use in controlled drug delivery are known in the art (Langer, Accounts Chem. Res. 26:537-542, 1993). For example, the block copolymer, polaxamer 407, exists as a viscous yet mobile liquid at low temperatures but forms a semisolid gel at body temperature. It has been shown to be an effective vehicle for formulation and sustained delivery of recombinant interleukin-2 and urease (Johnston et al., Pharm. Res. 9:425-434, 1992; and Pec et al., J. Parent. Sci. Tech. 44(2):58-65, 1990). Alternatively, hydroxyapatite has been used as a microcarrier for controlled release of proteins (Ijntema et al., Int. J. Pharm. 112:215-224, 1994). In yet another aspect, liposomes are used for controlled release as well as drug targeting of the lipid-capsulated drug (Betageri et al., Liposome Drug Delivery Systems, Technomic Publishing Co., Inc., Lancaster, Pa. (1993)). Numerous additional systems for controlled delivery of therapeutic proteins are known (see U.S. Pat. Nos. 5,055,303; 5,188,837; 4,235,871; 4,501,728; 4,837,028; 4,957,735; 5,019,369; 5,055,303; 5,514,670; 5,413,797; 5,268,164; 5,004,697; 4,902,505; 5,506,206; 5,271,961; 5,254,342 and 5,534,496).
[0696] In some embodiments, the compositions of the invention may be injectable suspensions, solutions, sprays, lyophilized powders, syrups, elixirs and the like. Any suitable form of composition may be used. To prepare such a composition, a nucleic acid or vector of the invention, having the desired degree of purity, is mixed with one or more pharmaceutically acceptable carriers and/or excipients. The carriers and excipients must be "acceptable" in the sense of being compatible with the other ingredients of the composition. Acceptable carriers, excipients, or stabilizers are nontoxic to recipients at the dosages and concentrations employed, and include, but are not limited to, water, saline, phosphate buffered saline, dextrose, glycerol, ethanol, or combinations thereof, buffers such as phosphate, citrate, and other organic acids; antioxidants including ascorbic acid and methionine; preservatives (such as octadecyldimethylbenzyl ammonium chloride; hexamethonium chloride; benzalkonium chloride, benzethonium chloride; phenol, butyl or benzyl alcohol; alkyl parabens such as methyl or propyl paraben; catechol; resorcinol; cyclohexanol; 3-pentanol; and m-cresol); low molecular weight (less than about 10 residues) polypeptide; proteins, such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidone; amino acids such as glycine, glutamine, asparagine, histidine, arginine, or lysine; monosaccharides, disaccharides, and other carbohydrates including glucose, mannose, or dextrins; chelating agents such as EDTA; sugars such as sucrose, mannitol, trehalose or sorbitol; salt-forming counter-ions such as sodium; metal complexes (e.g., Zn-protein complexes); and/or non-ionic surfactants such as TWEEN.TM. PLURONICS.TM. or polyethylene glycol (PEG).
[0697] The compositions can be designed to introduce the antibodies, nucleic acids or expression vectors to a desired site of action and release it at an appropriate and controllable rate. Methods of preparing controlled-release formulations are known in the art. For example, controlled release preparations can be produced by the use of polymers to complex or absorb the immunogen and/or immunogenic composition. A controlled-release formulations can be prepared using appropriate macromolecules (for example, polyesters, polyamino acids, polyvinyl, pyrrolidone, ethylenevinylacetate, methylcellulose, carboxymethylcellulose, or protamine sulfate) known to provide the desired controlled release characteristics or release profile. Another possible method to control the duration of action by a controlled-release preparation is to incorporate the active ingredients into particles of a polymeric material such as, for example, polyesters, polyamino acids, hydrogels, polylactic acid, polyglycolic acid, copolymers of these acids, or ethylene vinylacetate copolymers. Alternatively, instead of incorporating these active ingredients into polymeric particles, it is possible to entrap these materials into microcapsules prepared, for example, by coacervation techniques or by interfacial polymerization, for example, hydroxymethylcellulose or gelatin-microcapsule and poly-(methylmethacrylate) microcapsule, respectively, in colloidal drug delivery systems (for example, liposomes, albumin microspheres, microemulsions, nano-particles and nanocapsules) or in macroemulsions. Such techniques are disclosed in New Trends and Developments in Vaccines, Voller et al. (eds.), University Park Press, Baltimore, Md., 1978 and Remington's Pharmaceutical Sciences, 16th edition.
[0698] The compositions can be administered using any suitable delivery method including, but not limited to, intramuscular, intravenous, intradermal, mucosal, and topical delivery. Such techniques are well known to those of skill in the art. More specific examples of delivery methods are intramuscular injection, intradermal injection, and subcutaneous injection. However, delivery need not be limited to injection methods. Further, delivery of DNA to animal tissue has been achieved by cationic liposomes (Watanabe et al., (1994) Mol. Reprod. Dev. 38:268-274; and WO 96/20013), direct injection of naked DNA into animal muscle tissue (Robinson et al., (1993) Vaccine 11:957-960; Hoffman et al., (1994) Vaccine 12: 1529-1533; Xiang et al., (1994) Virology 199: 132-140; Webster et al., (1994) Vaccine 12: 1495-1498; Davis et al., (1994) Vaccine 12: 1503-1509; and Davis et al., (1993) Hum. Mol. Gen. 2: 1847-1851), or intradermal injection of DNA using "gene gun" technology (Johnston et al., (1994) Meth. Cell Biol. 43:353-365). Alternatively, delivery routes can be oral, intranasal or by any other suitable route. Delivery also be accomplished via a mucosal surface such as the anal, vaginal or oral mucosa.
[0699] Dosing schedules (or regimens) can be readily determined for the particular subject and composition. Hence, the composition can be administered one or more times to the subject. Preferably, there is a set time interval between separate administrations of the composition. While this interval varies for every subject, typically it can range from 10 days to several weeks, and is often 2, 4, 6 or 8 weeks. In some embodiments, the interval can be typically from 2 to 6 weeks.
[0700] The compositions of the invention can be administered alone, or can be co-administered, or sequentially administered, with other HIV immunogens and/or HIV immunogenic compositions, e.g., with "other" immunological, antigenic or vaccine or therapeutic compositions thereby providing multivalent or "cocktail" or combination compositions of the invention and methods of employing them. Again, the ingredients and manner (sequential or co-administration) of administration, as well as dosages can be determined taking into consideration such factors as the age, sex, weight, species and condition of the particular subject, and the route of administration.
Kits
[0701] The present invention also includes kits useful in performing diagnostic and prognostic assays using the antibodies of the present invention. Kits of the invention include a suitable container comprising an HIV-1 antibody of the invention in either labeled or unlabeled form. In addition, when the antibody is supplied in a labeled form suitable for an indirect binding assay, the kit further includes reagents for performing the appropriate indirect assay. For example, the kit includes one or more suitable containers including enzyme substrates or derivatizing agents, depending on the nature of the label. Control samples and/or instructions are also included.
Sample Embodiments
[0702] This section describes exemplary compositions and methods of the invention, presented without limitation, as a series of paragraphs, some or all of which may be alphanumerically designated for clarity and efficiency. Each of these paragraphs can be combined with one or more other paragraphs, and/or with disclosure from elsewhere in this application, including the materials incorporated by reference, in any suitable manner. Some of the paragraphs below expressly refer to and further limit other paragraphs, providing without limitation examples of some of the suitable combinations. [0703] 1. An isolated anti-HIV antibody that is capable of neutralizing at least 95% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. [0704] 2. An isolated anti-HIV antibody that is capable of neutralizing at least 99% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. [0705] 3. An isolated anti-HIV antibody that is capable of neutralizing 100% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. [0706] 4. An isolated anti-HIV antibody that neutralizes 100% of the HIV clade B, G and D viruses listed in Table 1 with an IC50 value of less than 50 .mu.g/mL. [0707] 5. The isolated anti-HIV antibody of any of paragraphs 1-4, wherein the anti-HIV antibody binds to a HIV gp120 epitope comprising outer domain loop D (which comprises 275-283), the CD4 binding loop (which comprises 354-371), the bridging sheet (which comprises 427-439) and loop V5 (which comprises 455-463) and gp120 inner domain: helix alpha-1 of Layer 2 (comprising positions 96-106) and 469-480 (loop prior and helix alpha-5 of Layer 3). [0708] 6. The isolated anti-HIV antibody of any of paragraphs 1-5, wherein the anti-HIV antibody binds to a HIV gp120 Layer 2 residues W96, K97, E102, G124, Loop D residues E275, N276, T278, N279, N280, A281, K282, CD4 binding loop residues P364, 5365, G366, G367, D368, 1371, bridging sheet residues W427, Q428, G429, Loop V5 residues T455, R456, D457, G458, G459, A460, N461, T463, and Layer 3 residues R469, P470, G471, G472, G473, N474, K476, D477, R480. [0709] 7. The isolated anti-HIV antibody of any of paragraphs 1-6, wherein the antibody has a Kd for BaL-gp120 of at least about 2.456.times.10.sup.-8M as determined by surface plasmon resonance. [0710] 8. The isolated anti-HIV antibody of any of paragraphs 1-7, wherein the antibody has a Kd for BaL-gp120 of at least about 1.59.times.10.sup.-8M as determined by surface plasmon resonance. [0711] 9. The isolated anti-HIV antibody of any of paragraphs 1-8, wherein the antibody has a Kd for BaL-gp120 of at least about 1.562.times.10.sup.-8M as determined by surface plasmon resonance. [0712] 10. The isolated anti-HIV antibody of any of paragraphs 1-9, wherein the antibody has a Kd for BaL-gp120 of at least about 1.143.times.10.sup.-9M as determined by surface plasmon resonance. [0713] 11. The isolated anti-HIV antibody of any of paragraphs 1-10, wherein the antibody has Kd for BaL-gp120 of at least about 8.602.times.10.sup.-10 M as determined by surface plasmon resonance. [0714] 12. The isolated anti-HIV antibody of any of paragraphs 1-11, wherein the anti-HIV antibody further neutralizes strain CNE5 (clade CRF01_AE) with an IC50 value of less than 50 .mu.g/mL. [0715] 13. The isolated anti-HIV antibody of any of paragraphs 1-12, wherein the antibody is capable of neutralizing HIV pseudoviruses listed in Table 1 with a median IC50 value of less than 0.5 .mu.g/mL. [0716] 14. The isolated anti-HIV antibody of any of paragraphs 1-13, wherein the anti-HIV antibody neutralizes at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, or at least 99% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than about 1 .mu.g/ml, between about 1-5 .mu.g/ml or greater than about 5 .mu.g/ml. [0717] 15. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least about 86.4% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0718] 16. The isolated anti-HIV antibody of paragraph 15, wherein the antibody is N49P7 or an antigen binding fragment thereof. [0719] 17. The isolated anti-HIV antibody of paragraph 16, wherein the antibody comprises the VH and VL regions of N49P7 as described herein. [0720] 18. The isolated anti-HIV antibody of paragraph 17, wherein the antibody comprises the CDRs of the VH and VL regions of N49P7 as described herein. [0721] 19. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 88.7% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0722] 20. The isolated anti-HIV antibody of paragraph 19, wherein the antibody is N49P7.1 or an antigen binding fragment thereof [0723] 21. The isolated anti-HIV antibody of paragraph 20, wherein the antibody comprises the VH and VL regions of N49P7.1 as described herein. [0724] 22. The isolated anti-HIV antibody of paragraph 21, wherein the antibody comprises the CDRs of the VH and VL regions of N49P7.1 as described herein. [0725] 23. The isolated anti-HIV antibody of any of paragraphs 1-22, wherein the anti-HIV antibody neutralizes at least 84.5% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0726] 24. The isolated anti-HIV antibody of paragraph 23, wherein the antibody is N49P7.2 or an antigen binding fragment thereof [0727] 25. The isolated anti-HIV antibody of any of paragraph 24, wherein the antibody comprises the VH and VL regions of N49P7.2 as described herein. [0728] 26. The isolated anti-HIV antibody of any of paragraph 25, wherein the antibody comprises the CDRs of the VH and VL regions of N49P7.2 as described herein. [0729] 27. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 71.8% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0730] 28. The isolated anti-HIV antibody of paragraph 27, wherein the antibody is N49P6 or an antigen binding fragment thereof. [0731] 29. The isolated anti-HIV antibody of paragraph 28, wherein the antibody comprises the VH and VL regions of N49P6 as described herein. [0732] 30. The isolated anti-HIV antibody of paragraph 29, wherein the antibody comprises the CDRs of the VH and VL regions of N49P6 as described herein. [0733] 31. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 93.3% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0734] 32. The isolated anti-HIV antibody of paragraph 31, wherein the antibody is N49P9 or an antigen binding fragment thereof. [0735] 33. The isolated anti-HIV antibody of paragraph 32, wherein the antibody comprises the VH and VL regions of N49P9 as described herein. [0736] 34. The isolated anti-HIV antibody of paragraph 33, wherein the antibody comprises the CDRs of the VH and VL regions of N49P9 as described herein. [0737] 35. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 91.1% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0738] 36. The isolated anti-HIV antibody of paragraph 35, wherein the antibody is N49P9.1 or an antigen binding fragment thereof [0739] 37. The isolated anti-HIV antibody of paragraph 36, wherein the antibody comprises VH and VL regions of N49P9.1 as described herein. [0740] 38. The isolated anti-HIV antibody of paragraph 37, wherein the antibody comprises the CDRs of the VH and VL regions of N49P9.1 as described herein. [0741] 39. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 41.9% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0742] 40. The isolated anti-HIV antibody of paragraph 39, wherein the anti-HIV antibody is N49P11 or an antigen binding fragment thereof. [0743] 41. The isolated anti-HIV antibody of paragraph 40, wherein the anti-HIV antibody comprises the VH and VL regions of N49P11 as described herein. [0744] 42. The isolated anti-HIV antibody of paragraph 41, wherein the anti-HIV antibody comprises the CDRs of the VH and VL regions of N49P11 as described herein. [0745] 43. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 2.1% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0746] 44. The isolated anti-HIV antibody of paragraph 43, wherein the anti-HIV antibody is N49P18.1 or an antigen binding fragment thereof [0747] 45. The isolated anti-HIV antibody of paragraph 44, wherein the anti-HIV antibody comprises the VH and VL regions of N49P18.1 as described herein. [0748] 46. The isolated anti-HIV antibody of paragraph 45, wherein the anti-HIV antibody comprises the CDRs of the VH and VL regions of N49P18.1 as described herein. [0749] 47. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 60% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0750] 48. The isolated anti-HIV antibody of paragraph 47, wherein the anti-HIV antibody is N49P19 or an antigen binding fragment thereof. [0751] 49. The isolated anti-HIV antibody of paragraph 48, wherein the anti-HIV antibody comprises the VH and VL regions of N49P19 as described herein. [0752] 50. The isolated anti-HIV antibody of paragraph 49, wherein the anti-HIV antibody comprises the CDRs of the VH and VL regions of N49P19 as described herein. [0753] 51. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 58.3% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0754] 52. The isolated anti-HIV antibody of paragraph 51, wherein the anti-HIV antibody is N49P22 or an antigen binding fragment thereof. [0755] 53. The isolated anti-HIV antibody of paragraph 52, wherein the anti-HIV antibody comprises the VH and VL regions of N49P22 as described herein. [0756] 54. The isolated anti-HIV antibody of paragraph 53, wherein the anti-HIV antibody comprises the CDRs of the VH and VL regions of N49P22 as described herein. [0757] 55. The isolated anti-HIV antibody of any of paragraphs 1-14, wherein the anti-HIV antibody neutralizes at least 88.6% of the HIV pseudoviruses listed in Table 1 with an IC50 value of less than 1 .mu.g/mL. [0758] 56. The isolated anti-HIV antibody of paragraph 55, wherein the anti-HIV antibody is N49P23 or an antigen binding fragment thereof. [0759] 57. The isolated anti-HIV antibody of paragraph 56, wherein the anti-HIV antibody comprises the VH and VL regions of N49P23 as described herein. [0760] 58. The isolated anti-HIV antibody of paragraph 57, wherein the anti-HIV antibody comprises the CDRs of the VH and VL regions of N49P23 as described herein. [0761] 59. The isolated anti-HIV antibody of any of paragraphs 1-58, wherein anti-HIV antibody is selected from the group consisting of: [0762] a. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYDFIDYV (SEQ ID NO:401), CDR H2 comprises MNPSGGGT (SEQ ID NO:402) and CDR H3 comprises VRDRANGSGRRRFESVNWFLDL (SEQ ID NO:403); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); [0763] b. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFPDYI (SEQ ID NO:405), CDR H2 comprises INPMGGQV (SEQ ID NO:406) and CDR H3 comprises VRDRSNGSGRRFESSN (SEQ ID NO:407); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0764] c. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYL (SEQ ID NO:409), CDR H2 comprises MNPVYGQV (SEQ ID NO:410) and CDR H3 comprises VRDTGDGSRRHFDSINWFLDL (SEQ ID NO:411); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412); [0765] d. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises IDPPYGQV (SEQ ID NO:414) and CDR H3 comprises VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0766] e. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFVDYF (SEQ ID NO:416), CDR H2 comprises MDPLNGRP (SEQ ID NO:417) and CDR H3 comprises VRDKSNGSGRRFDSSNWFLDL (SEQ ID NO:418); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0767] f. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFSDYI (SEQ ID NO:420), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEV (SEQ ID NO:422); [0768] g. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFIDYI (SEQ ID NO:423), CDR H2 comprises IDPMNGRP (SEQ ID NO:424) and CDR H3 comprises VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0769] h. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYI (SEQ ID NO:426), CDR H2 comprises MNPMGGRT (SEQ ID NO:427) and CDR H3 comprises VRDKSNGSGKRFDSSNWFLDL (SEQ ID NO:425); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0770] i. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFVDYL (SEQ ID NO:428), CDR H2 comprises MDPMNGRP (SEQ ID NO:429) and CDR H3 comprises VRDKSGGSGKLFDSSNWFLDL (SEQ ID NO:430); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0771] j. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises INPGYGQV (SEQ ID NO:431) and CDR H3 comprises VRDRSNGWGKRFESSNWFLDL (SEQ ID NO:415); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0772] k. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises MDPSYGQV (SEQ ID NO:432) and CDR H3 comprises VRDRSHGSGRQFESSNWFLDL (SEQ ID NO:433); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408);
[0773] l. an antibody comprising a heavy chain variable region, wherein CDR comprises GYTFTDYV (SEQ ID NO:413), CDR H2 comprises MDPSFGQM (SEQ ID NO:434) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0774] m. an antibody comprising a heavy chain variable region, wherein CDR comprises GYRFTDYV (SEQ ID NO:436), CDR H2 comprises MDPSFGRM (SEQ ID NO:437) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0775] n. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFIDYV (SEQ ID NO:438), CDR H2 comprises MDPTYGRM (SEQ ID NO:439) and CDR H3 comprises VRDRSHGSGRLFESSNWFLDL (SEQ ID NO:435); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0776] o. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFLDYI (SEQ ID NO:440), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0777] p. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0778] q. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSR (SEQ ID NO:447), CDR H2 comprises INPLQGGV (SEQ ID NO:448) and CDR H3 comprises ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0779] r. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTTHHGHF (SEQ ID NO:500), CDR H2 comprises MNPMTGQM (SEQ ID NO:462) and CDR H3 comprises ARGDFGQNYPFHY (SEQ ID NO:463); and a light chain variable region, wherein CDR L1 comprises NRYL (SEQ ID NO:464), CDR L2 comprises DDN and CDR L3 comprises ASYER (SEQ ID NO:465); [0780] s. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFMDQF (SEQ ID NO:466), CDR H2 comprises MNPIYGQV (SEQ ID NO:467) and CDR H3 comprises ARGPSGENYPFHY (SEQ ID NO:444); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0781] t. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSR (SEQ ID NO:447), CDR H2 comprises INPLHGGV (SEQ ID NO:468) and CDR H3 comprises ARGIDGKSYPFHF (SEQ ID NO:449); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0782] u. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTKYF (SEQ ID NO:451), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADSYGSSYPFHH (SEQ ID NO:453); and a light chain variable region, wherein CDR L1 comprises GNYNP (SEQ ID NO:454), CDR L2 comprises EDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0783] v. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTKYT (SEQ ID NO:456), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADLSGPTYPFHH (SEQ ID NO:457); and a light chain variable region, wherein CDR L1 comprises GNYNP (SEQ ID NO:454), CDR L2 comprises EDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0784] w. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFNFIDSV (SEQ ID NO:458), CDR H2 comprises IKPLRGAV (SEQ ID NO:459) and CDR H3 comprises AKGAFRGGSPFGF (SEQ ID NO:460); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0785] x. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYTFTSYF (SEQ ID NO:469), CDR H2 comprises INPLHGAV (SEQ ID NO:470) and CDR H3 comprises TRGIVADGWPYGH (SEQ ID NO:471); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises EGA and CDR L3 comprises SSLQF (SEQ ID NO:472); [0786] y. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFTFIDHI (SEQ ID NO:473), CDR H2 comprises IKPLRGAV (SEQ ID NO:459) and CDR H3 comprises CKAAAPEEAFPLQY (SEQ ID NO:474); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSRTF (SEQ ID NO:475); [0787] z. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDHI (SEQ ID NO:476), CDR H2 comprises IKPLGGVA (SEQ ID NO:477) and CDR H3 comprises CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0788] aa. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFAFLDH (SEQ ID NO:480), CDR H2 comprises VKTIGGVV (SEQ ID NO:481) and CDR H3 comprises SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0789] bb. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFTEYF (SEQ ID NO:483), CDR H2 comprises LNPLRGAV (SEQ ID NO:484) and CDR H3 comprises ARAVFNEAFPFDY (SEQ ID NO:485); and a light chain variable region, wherein CDR L1 comprises VS, CDR L2 comprises DGD and CDR L3 comprises ASREF (SEQ ID NO:461); [0790] cc. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDHI (SEQ ID NO:476), CDR H2 comprises IKPLGGVA (SEQ ID NO:477) and CDR H3 comprises CKAAAPDEAFPLEY (SEQ ID NO:478); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DND and CDR L3 comprises SSTTF (SEQ ID NO:479); [0791] dd. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFAFLDHI (SEQ ID NO:486), CDR H2 comprises VKTIGGVV (SEQ ID NO:481) and CDR H3 comprises SKAAAPDEAFPLEF (SEQ ID NO:482); and a light chain variable region, wherein CDR L1 comprises NVD, CDR L2 comprises DNN and CDR L3 comprises SSTTF (SEQ ID NO:479); [0792] ee. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFKFIDSV (SEQ ID NO:487), CDR H2 comprises IKPLGGAV (SEQ ID NO:488) and CDR H3 comprises AKGAFGGGSPFGF (SEQ ID NO:489); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0793] ff. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFNFIDSV (SEQ ID NO:458), CDR H2 comprises IKPLRGGV (SEQ ID NO:490) and CDR H3 comprises AKGAFGGSSPFGF (SEQ ID NO:491); and a light chain variable region, wherein CDR L1 comprises DVT and CDR L2 comprises ASREF (SEQ ID NO:461); [0794] gg. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GFTFIKYT (SEQ ID NO:492), CDR H2 comprises IHPRTGAV (SEQ ID NO:452) and CDR H3 comprises ARGAFEADLYGPTYPFHH (SEQ ID NO:493); and a light chain variable region, wherein CDR L1 comprises GSYNP (SEQ ID NO:494), CDR L2 comprises DDN and CDR L3 comprises ASFEF (SEQ ID NO:455); [0795] hh. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYNFVDSL (SEQ ID NO:495), CDR H2 comprises INPLQGGV (SEQ ID NO:448) and CDR H3 comprises ARGIDGNSYPFHF (SEQ ID NO:496); and a light chain variable region, wherein CDR L1 comprises S, CDR L2 comprises ESS and CDR L3 comprises SILEF (SEQ ID NO:450); [0796] ii. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0797] jj. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAFEN (SEQ ID NO:404); [0798] kk. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFD and CDR L3 comprises WAFEA (SEQ ID NO:412); [0799] ll. an antibody comprising a heavy chain variable region, wherein CDR comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0800] mm. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNY, CDR L2 comprises DFN and CDR L3 comprises WAYDA (SEQ ID NO:419); [0801] nn. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFPDYI (SEQ ID NO:497), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408); [0802] oo. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises MNPTYGQV (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSN (SEQ ID NO:498); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); [0803] pp. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYKFMDQL (SEQ ID NO:442), CDR H2 comprises (SEQ ID NO:443) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises RHII (SEQ ID NO:445), CDR L2 comprises DDD and CDR L3 comprises NTYEF (SEQ ID NO:446); and [0804] qq. an antibody comprising a heavy chain variable region, wherein CDR H1 comprises GYRFLDYI (SEQ ID NO:440), CDR H2 comprises MNPMGGQV (SEQ ID NO:421) and CDR H3 comprises VRDRSNGSGKRFESSNWFLDL (SEQ ID NO:441); and a light chain variable region, wherein CDR L1 comprises HNL, CDR L2 comprises DFN and CDR L3 comprises WAYEA (SEQ ID NO:408). [0805] 60. The isolated anti-HIV antibody of any of claims 1-59, wherein the anti-HIV antibody comprises a heavy chain or an antigen binding fragment thereof and a light chain or an antigen binding fragment thereof, wherein the heavy chain or antigen binding fragment thereof comprises a heavy chain variable (V.sub.H) region and the light chain or antigen binding fragment thereof comprises a light chain variable (V.sub.L) region; wherein the anti-HIV antibody is selected from the group consisting of an antibody: [0806] a. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:1 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:2 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0807] b. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:3 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:4 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0808] c. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:5 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:6 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0809] d. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:7 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:8 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0810] e. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:9 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:10 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0811] f. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:11 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:12 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0812] g. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:13 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:14 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0813] h. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:15 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:16 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0814] i. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:17 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:18 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0815] j. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:19 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:20 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0816] k. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:21 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:22 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0817] l. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:23 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:24 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions;
[0818] m. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:25 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:26 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0819] n. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:27 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:28 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0820] o. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:29 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:30 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0821] p. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:31 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:32 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0822] q. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:33 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:34 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0823] r. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:35 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:36 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0824] s. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:37 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:38 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0825] t. wherein the VH region comprises amino acids 1-123 of SEQ ID NO:39 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:40 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0826] u. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:41 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:42 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0827] v. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:43 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:44 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0828] w. wherein the VH region comprises amino acids 1-125 of SEQ ID NO:45 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:46 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0829] x. wherein the VH region comprises amino acids 1-125 of SEQ ID NO:47 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:48 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0830] y. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:49 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:50 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0831] z. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:51 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:52 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0832] aa. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:53 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:54 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0833] bb. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:55 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:56 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0834] cc. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:57 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:58 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0835] dd. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:59 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:60 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0836] ee. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:61 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-98 of SEQ ID NO:62 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0837] ff. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:63 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:64 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0838] gg. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:65 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:66 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0839] hh. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:67 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:68 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0840] ii. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:69 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:70 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0841] jj. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:71 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:72 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0842] kk. wherein the VH region comprises amino acids 1-125 of SEQ ID NO:73 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:74 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0843] ll. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:75 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:76 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0844] mm. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:153 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:155 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0845] nn. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:157 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:159 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0846] oo. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:161 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:163 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0847] pp. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:165 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:167 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0848] qq. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:169 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:171 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0849] rr. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:173 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:175 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0850] ss. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:177 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:179 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0851] tt. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:181 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:183 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0852] uu. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:185 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:187 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0853] vv. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:189 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:191 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0854] ww. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:193 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:195 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0855] xx. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:197 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:199 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0856] yy. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:201 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:203 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0857] zz. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:205 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:207 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0858] aaa. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:209 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:211 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0859] bbb. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:213 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:215 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VH region comprises amino acids 1-127 of SEQ ID NO:217 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:219 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0860] ccc. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:221 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:223 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0861] ddd. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:225 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:227 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0862] eee. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:229 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:231 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0863] fff. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:233 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:235 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0864] ggg. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:237 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:239 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions;
[0865] hhh. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:241 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:243 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0866] iii. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:245 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:247 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0867] jjj. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:249 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:251 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0868] kkk. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:253 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:255 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0869] lll. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:257 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:259 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0870] mmm. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:261 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:263 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0871] nnn. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:265 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:267 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0872] ooo. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:269 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:271 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0873] ppp. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:273 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:275 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0874] qqq. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:277 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:279 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0875] rrr. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:281 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:283 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0876] sss. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:285 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:287 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0877] ttt. wherein the VH region comprises amino acids 1-127 of SEQ ID NO:289 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:291 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0878] uuu. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:293 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:295 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0879] vvv. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:297 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:299 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0880] www. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:301 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:303 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0881] xxx. wherein the VH region comprises amino acids 1-123 of SEQ ID NO:305 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:307 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0882] yyy. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:309 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:311 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0883] zzz. wherein the VH region comprises amino acids 1-128 of SEQ ID NO:313 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:315 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0884] aaaa. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:317 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:319 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0885] bbbb. wherein the VH region comprises amino acids 1-123 of SEQ ID NO:321 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:323 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0886] cccc. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:325 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:327 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0887] dddd. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:329 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-100 of SEQ ID NO:331 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0888] eeee. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:333 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:335 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0889] ffff. wherein the VH region comprises amino acids 1-125 of SEQ ID NO:337 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:339 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0890] gggg. wherein the VH region comprises amino acids 1-125 of SEQ ID NO:341 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:343 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0891] hhhh. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:345 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:347 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0892] iiii. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:349 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-97 of SEQ ID NO:351 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0893] jjjj. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:353 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:355 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0894] kkkk. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:357 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:359 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0895] llll. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:361 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:363 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0896] mmmm. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:365 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:367 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0897] nnnn. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:369 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-98 of SEQ ID NO:371 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0898] oooo. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:373 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:375 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0899] pppp. wherein the VH region comprises amino acids 1-121 of SEQ ID NO:377 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-99 of SEQ ID NO:379 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0900] qqqq. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:381 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:383 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0901] rrrr. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:385 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:387 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0902] ssss. wherein the VH region comprises amino acids 1-120 of SEQ ID NO:389 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-96 of SEQ ID NO:391 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0903] tttt. wherein the VH region comprises amino acids 1-125 of SEQ ID NO:393 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; wherein the VL region comprises amino acids 1-101 of SEQ ID NO:395 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; [0904] uuuu. wherein the VH region comprises and amino acids 1-120 of SEQ ID NO:397 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions; and wherein the VL region comprises amino acids 1-97 of SEQ ID NO:399 or a variant thereof comprising 1, 2, 3, or 4 conservative amino acid substitutions. [0905] 61. The anti-HIV antibody of any of paragraphs 1-60, wherein the anti-HIV antibody is selected from the group consisting of: [0906] a. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:1 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:2 or an antigen binding fragment thereof; [0907] b. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:3 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:4 or an antigen binding fragment thereof; [0908] c. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:5 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:6 or an antigen binding fragment thereof; [0909] d. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:7 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:8 or an antigen binding fragment thereof; [0910] e. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:9 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:10 or an antigen binding fragment thereof; [0911] f. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:11 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:12 or an antigen binding fragment thereof; [0912] g. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:13 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:14 or an antigen binding fragment thereof; [0913] h. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:15 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:16 or an antigen binding fragment thereof; [0914] i. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:17 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:18 or an antigen binding fragment thereof; [0915] j. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:19 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:20 or an antigen binding fragment thereof;
[0916] k. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:21 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:22 or an antigen binding fragment thereof; [0917] l. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:23 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:24 or an antigen binding fragment thereof; [0918] m. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:25 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:26 or an antigen binding fragment thereof; [0919] n. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:27 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:28 or an antigen binding fragment thereof; [0920] o. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:29 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:30 or an antigen binding fragment thereof; [0921] p. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:31 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:32 or an antigen binding fragment thereof; [0922] q. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:33 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:34 or an antigen binding fragment thereof; [0923] r. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:35 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:36 or an antigen binding fragment thereof; [0924] s. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:37 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:38 or an antigen binding fragment thereof; [0925] t. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:39 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:40 or an antigen binding fragment thereof; [0926] u. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:41 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:42 or an antigen binding fragment thereof; [0927] v. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:43 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:44 or an antigen binding fragment thereof; [0928] w. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:45 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:46 or an antigen binding fragment thereof; [0929] x. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:47 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:48 or an antigen binding fragment thereof; [0930] y. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:49 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:50 or an antigen binding fragment thereof; [0931] z. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:51 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:52 or an antigen binding fragment thereof; [0932] aa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:53 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:54 or an antigen binding fragment thereof; [0933] bb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:55 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:56 or an antigen binding fragment thereof; [0934] cc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:57 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:58 or an antigen binding fragment thereof; [0935] dd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:59 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:60 or an antigen binding fragment thereof; [0936] ee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:61 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:62 or an antigen binding fragment thereof; [0937] ff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:63 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:64 or an antigen binding fragment thereof; [0938] gg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:65 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:66 or an antigen binding fragment thereof; [0939] hh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:67 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:68 or an antigen binding fragment thereof; [0940] ii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:69 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:70 or an antigen binding fragment thereof; [0941] jj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:71 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:72 or an antigen binding fragment thereof; [0942] kk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:73 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:74 or an antigen binding fragment thereof; [0943] ll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:75 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:76 or an antigen binding fragment thereof; [0944] mm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:153 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:155 or an antigen binding fragment thereof; [0945] nn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:157 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:159 or an antigen binding fragment thereof; [0946] oo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:161 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:163 or an antigen binding fragment thereof; [0947] pp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:165 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:167 or an antigen binding fragment thereof; [0948] qq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:169 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:171 or an antigen binding fragment thereof; [0949] rr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:173 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:175 or an antigen binding fragment thereof; [0950] ss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:177 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:179 or an antigen binding fragment thereof; [0951] tt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:181 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:183 or an antigen binding fragment thereof; [0952] uu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:185 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:187 or an antigen binding fragment thereof; [0953] vv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:189 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:191 or an antigen binding fragment thereof; [0954] ww. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:193 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:195 or an antigen binding fragment thereof; [0955] xx. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:197 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:199 or an antigen binding fragment thereof; [0956] yy. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:201 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:203 or an antigen binding fragment thereof; [0957] zz. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:205 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:207 or an antigen binding fragment thereof; [0958] aaa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:209 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:211 or an antigen binding fragment thereof; [0959] bbb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:213 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:215 or an antigen binding fragment thereof; [0960] ccc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:217 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:219 or an antigen binding fragment thereof; [0961] ddd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:221 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:223 or an antigen binding fragment thereof; [0962] eee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:225 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:227 or an antigen binding fragment thereof; [0963] fff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:229 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:231 or an antigen binding fragment thereof; [0964] ggg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:233 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:235 or an antigen binding fragment thereof; [0965] hhh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:237 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:239 or an antigen binding fragment thereof; [0966] iii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:241 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:243 or an antigen binding fragment thereof; [0967] jjj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:245 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:247 or an antigen binding fragment thereof; [0968] kkk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:249 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:251 or an antigen binding fragment thereof; [0969] lll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:253 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:255 or an antigen binding fragment thereof; [0970] mmm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:257 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:259 or an antigen binding fragment thereof; [0971] nnn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:261 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:263 or an antigen binding fragment thereof; [0972] ooo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:265 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:267 or an antigen binding fragment thereof; [0973] ppp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:269 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:271 or an antigen binding fragment thereof; [0974] qqq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:273 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:275 or an antigen binding fragment thereof; [0975] rrr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:277 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:279 or an antigen binding fragment thereof; [0976] sss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:281 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:283 or an antigen binding fragment thereof; [0977] ttt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:285 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:287 or an antigen binding fragment thereof; [0978] uuu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:289 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:291 or an antigen binding fragment thereof; [0979] vvv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:293 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:295 or an antigen binding fragment thereof; [0980] www. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:297 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:299 or an antigen binding fragment thereof;
[0981] xxx. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:301 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:303 or an antigen binding fragment thereof; [0982] yyy. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:305 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:307 or an antigen binding fragment thereof; [0983] zzz. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:309 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:311 or an antigen binding fragment thereof; [0984] aaaa. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:313 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:315 or an antigen binding fragment thereof; [0985] bbbb. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:317 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:319 or an antigen binding fragment thereof; [0986] cccc. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:321 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:323 or an antigen binding fragment thereof; [0987] dddd. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:325 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:327 or an antigen binding fragment thereof; [0988] eeee. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:329 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:331 or an antigen binding fragment thereof; [0989] ffff. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:333 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:335 or an antigen binding fragment thereof; [0990] gggg. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:337 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:339 or an antigen binding fragment thereof; [0991] hhhh. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:341 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:343 or an antigen binding fragment thereof; [0992] iiii. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:345 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:347 or an antigen binding fragment thereof; [0993] jjjj. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:349 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:351 or an antigen binding fragment thereof; [0994] kkkk. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:353 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:355 or an antigen binding fragment thereof; [0995] llll. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:357 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:359 or an antigen binding fragment thereof; [0996] mmmm. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:361 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:363 or an antigen binding fragment thereof; [0997] nnnn. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:365 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:367 or an antigen binding fragment thereof; [0998] oooo. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:369 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:371 or an antigen binding fragment thereof; [0999] pppp. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:373 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:375 or an antigen binding fragment thereof; [1000] qqqq. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:377 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:379 or an antigen binding fragment thereof; [1001] rrrr. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:381 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:383 or an antigen binding fragment thereof; [1002] ssss. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:385 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:387 or an antigen binding fragment thereof; [1003] tttt. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:389 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:391 or an antigen binding fragment thereof; [1004] uuuu. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:393 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:395 or an antigen binding fragment thereof; and [1005] vvvv. an antibody comprising a heavy chain amino acid sequence comprising SEQ ID NO:397 or an antigen binding fragment thereof and a light chain amino acid sequence comprising SEQ ID NO:399 or an antigen binding fragment thereof. [1006] 62. An isolated host cell expressing the antibody of any of paragraphs 1-61. [1007] 63. One or more vectors comprising a nucleic acid encoding the antibody of any of paragraphs 1-62. [1008] 64. The one or more vectors of paragraph 63, wherein one vector encodes a light chain sequence and another vector encodes a heavy chain sequence. [1009] 65. The one or more vectors of paragraph 64, wherein one vector encodes a light chain sequence and a heavy chain sequence. [1010] 66. A cell comprising the one or more vectors of any of paragraphs 62-65. [1011] 67. An engineered cell that expresses the antibody of any of paragraphs 1-66. [1012] 68. The cell of paragraph 67, wherein the cell is an immune cell. [1013] 69. The cell of paragraph 68, wherein the immune cell is a B cell. [1014] 70. A pharmaceutical composition comprising one or more antibodies of any of paragraphs 1-61 and/or cells of any of paragraphs 66-69 and a pharmaceutically acceptable carrier. [1015] 71. A method for treating or preventing HIV infection in a subject, comprising administering to the subject an effective amount of the composition of paragraph 70. [1016] 72. A method of functionally curing HIV comprising administering to the subject an effective amount of the composition of paragraph 70. [1017] 73. The method of any of paragraphs 71-72, wherein the composition is administered in combination with another therapy. [1018] 74. The method of paragraph 73, wherein the therapy is an anti-retroviral therapy.
[1019] Application of the teachings of the present invention to a specific problem is within the capabilities of one having ordinary skill in the art in light of the teaching contained herein. Examples of the compositions and methods of the invention appear in the following non-limiting Examples.
EXAMPLES
Example 1. Deconvoluting an HIV Super-Neutralizing Plasma Response
[1020] Anti-HIV-I envelope monoclonal antibodies isolated from memory B-cells have yielded broadly neutralizing antibodies (bNAbs), though none were pan-neutralizing. Here we identify a pan-neutralizing antibody lineage against a novel epitope by coupling proteomics of plasma antibodies with lineage analysis of bone marrow plasma cells from two HIV-1 "elite neutralizers." In both, a single lineage of anti-CD4 binding site antibodies matched circulating bNAbs sequences. Members of a single plasma cell lineage potently neutralized 100% of a validated multi-tier 117 pseudovirus panel, matching the sequence, specificity, and neutralization breadth of the circulating bNAbs. X-ray crystallography analysis of pan-neutralizing monoclonal, N49P7, identified its unique ability to bypass the CD4-binding site Phe43 cavity while reaching deep into the highly conserved residues of Layer 3 of the gp120 inner domain, likely accounting for its pan-neutralization. Conjoint analysis of plasma antibodies by proteomics and bone marrow derived lineages will improve understanding the evolution of anti-HIV-I bNAb responses.
[1021] Here we employed a matched genomic and proteomic approach to deconvolute a very broadly neutralizing response directed against gp120. The primary test subject, N60 (referred to as Subject 1 in a previous publication (Sajadi et al., J Virol 86, 5014-5025 (2012)) belongs to a previously reported Natural Viral Suppressor (NVS) cohort of subtype B-infected patients who exhibit persistent titers of very broad and potent neutralizing antibodies (Sajadi et al., J Infect Dis 213, 156-164 (2016); Sajadi et al., J Virol 86, 5014-5025 (2012).
[1022] We have previously found that multiple HIV-infected subjects harbor broad and potent neutralizing activities with highly shared biochemical determinants, such as basic isoelectric points (pI) and specificities for binding epitopes on free monomeric gp120 (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Infect Dis 213, 156-164 (2016); Sajadi et al., J Virol 86, 5014-5025 (2012)). Serum antibodies from N60 were able to neutralize 90% of 118 multi-clade Tier 2/3 panel of viruses (Table 5). The duration of such neutralization potency and breadth over a 5-year period was confirmed by sequential testing of N60 plasma against a cross-clade, multi-tier panel of viruses (Table 6).
TABLE-US-00006 TABLE 5 Neutralization of N60 parent and affinity purified samples. Titer in TZM.bl cells (ug/ml) N60 Pro N60 N60 Virus ID Clade* Tier IC50 IC50 IC50 6535.3 B 1B 23.629 2.337 1.399 QH0692.42 B 2 73.193 10.496 5.384 SC422661.8 B 2 30.877 3.333 2.194 PVO.4 B 3 23.164 1.478 1.005 TRO.11 B 2 35.183 7.264 5.693 AC10.0.29 B 2 27.920 4.144 3.550 RHPA4259.7 B 2 14.971 3.373 2.257 THRO4156.18 B 2 42.864 17.930 6.848 REJO4541.67 B 2 21.061 4.071 3.186 TRJO4551.58 B 3 72.808 16.072 13.344 WITO4160.33 B 2 151.278 46.052 23.311 CAAN5342.A2 B 2 30.870 4.007 3.148 WEAU d15 410 787 B (T/F) 2 32.622 4.254 2.479 1006 11 C3 1601 B (T/F) 2 31.461 5.019 3.047 1054 07 TC4 1499 B (T/F) 2 104.772 27.878 10.870 1056 10 TA11 1826 B (T/F) 1B 209.754 22.420 12.264 1012 11 TC21 3257 B (T/F) 1B 39.521 3.783 2.738 6240 08 TA5 4622 B (T/F) 2 216.472 45.439 38.068 6244 13 B5 4576 B (T/F) 2 58.906 4.678 4.487 62357 14 D3 4589 B (T/F) 2 60.360 11.454 8.697 SC05 8C11 2344 B (T/F) 2 28.437 4.712 3.912 Du156.12 C 2 37.524 6.824 5.405 Du172.17 C 2 119.944 >50 38.171 Du422.1 C 2 46.145 17.533 14.580 ZM197M.PB7 C 2 28.614 14.055 4.300 ZM214M.PL15 C 2 485.829 >50 >50 ZM233M.PB6 C 2 169.452 21.342 20.248 ZM249M.PL1 C 2 102.326 15.919 12.597 ZM53M.PB12 C 2 25.867 10.571 6.323 ZM109F.PB4 C 1B 210.029 >50 30.049 ZM135M.PL10a C 2 260.838 32.686 21.300 CAP45.2.00.G3 C 2 350.560 >50 >50 CAP210.2.00.E8 C 2 367.728 >50 >50 HIV-001428-2.42 C 2 25.972 2.701 2.187 HIV-0013095-2.11 C 2 211.157 22.522 18.247 HIV-16055-2.3 C 2 92.761 12.576 10.268 HIV-16845-2.22 C 2 426.772 >50 49.375 Ce1086_B2 C (T/F) NC 11.192 >50 1.641 Ce0393_C3 C (T/F) NC 142.594 30.286 15.431 Ce1176_A3 C (T/F) NC 192.528 45.344 28.479 Ce2010_F5 C (T/F) NC 70.275 19.463 12.692 Ce0682 E4 C (T/F) NC 64.759 17.431 10.243 Ce1172 H1 C (T/F) NC >528 >50 >50 Ce2060 G9 C (T/F) NC >528 >50 >50 Ce703010054_2A2 C (T/F) NC 188.873 31.683 19.604 BF1266.431a C (T/F) NC 154.369 46.996 27.310 246F C1G C (T/F) NC 352.889 >50 48.226 249M B10 C (T/F) NC 183.182 21.542 19.517 ZM247v1(Rev-) C (T/F) NC 190.864 >50 >50 7030102001E5(Rev-) C (T/F) NC 33.817 3.270 3.724 1394C9G1(Rev-) C (T/F) NC 59.578 16.555 5.566 Ce704809221_1B3 C (T/F) NC 65.767 35.580 10.697 CNE19 BC NC 16.594 2.854 1.520 CNE20 BC NC 15.633 4.173 1.565 CNE21 BC NC 91.501 22.960 14.653 CNE17 BC NC 139.056 21.438 13.846 CNE30 BC NC 221.349 >50 40.694 CNE52 BC NC 56.900 7.694 5.036 CNE53 BC NC 20.570 3.483 2.635 CNE58 BC NC 22.749 4.549 2.858 MS208.A1 A NC 211.293 >50 32.229 Q23.17 A 1B 17.575 2.938 1.794 Q461.e2 A 2 >528 >50 >50 Q769.d22 A 2 35.201 6.293 3.506 Q259.d2.17 A 2 33.807 7.060 3.559 Q842.d12 A 2 18.917 3.348 2.180 0260.v5.c36 A NC 80.819 13.873 43.483 3415.v1.c1 A 2 46.266 4.830 4.596 3365.v2.c2 A 2 54.620 8.493 5.802 191955_A11 A (T/F) NC 288.049 41.777 43.183 191084 B7-19 A (T/F) NC 34.834 4.607 1.917 9004SS_A3_4 A (T/F) NC 18.058 2.144 1.965 T257-31 CRF02_AG 3 426.998 >50 43.480 928-28 CRF02_AG 2 >528 >50 >50 263-8 CRF02_AG 2 22.824 4.205 4.064 T250-4 CRF02_AG 2 11.419 1.359 0.767 T251-18 CRF02_AG 3 371.838 >50 38.744 T278-50 CRF02_AG 3 >528 >50 >50 T255-34 CRF02_AG 2 131.913 15.603 12.016 211-9 CRF02_AG 2 291.355 42.611 24.418 235-47 CRF02_AG 2 87.864 9.609 6.028 620345.c01 CRF01_AE NC >528 >50 >50 CNE8 CRF01_AE NC 194.556 16.923 7.854 C1080.c03 CRF01_AE NC 483.323 >50 >50 R2184.c04 CRF01_AE NC 27.037 3.087 2.138 R1166.c01 CRF01_AE NC 130.381 29.864 24.622 R3265.c06 CRF01_AE NC 440.886 >50 >50 C2101.c01 CRF01_AE NC 92.755 14.543 16.106 C3347.c11 CRF01_AE NC 10.921 1.730 1.808 C4118.c09 CRF01_AE NC 117.159 17.416 16.615 CNE5 CRF01_AE NC 170.243 20.320 17.746 BJOX009000.02.4 CRF01_AE NC >528 >50 44.993 BJOX015000.11.5 CRF01_AE NC 250.047 >50 26.892 BJOXO10000.06.2 CRF01_AE NC >528 >50 >50 BJOX025000.01.1 CRF01 AE NC 328.505 >50 >50 BJOX028000.10.3 CRF01_AE NC >528 >50 >50 X1193_c1 G NC 121.508 3.302 13.356 P0402_c2_11 G NC 48.896 >50 4.239 X1254_c3 G NC 159.002 8.668 11.873 X2088_c9 G NC >528 >50 >50 X2131_C1_B5 G NC 95.017 9.848 10.189 P1981_C5_3 G NC 90.229 >50 14.873 X1632_S2_B10 G NC 110.495 >50 16.335 3016.v5.c45 D NC >528 >50 >50 A07412M1.vrc12 D NC 62.198 >50 5.815 231965.c01 D NC 171.994 20.212 26.544 231966.c02 D NC 459.176 >50 >50 3817.v2.c59 CD NC 58.502 >50 6.729 6480.v4.c25 CD NC 23.010 9.698 3.495 6952.v1.c20 CD NC 89.351 2.925 15.998 6811.v7.c18 CD NC 45.321 >50 6.443 89-F1_2_25 CD NC >528 >50 >50 3301.v1.c24 AC NC 42.393 0.576 5.568 6041.v3.c23 AC NC 15.836 1.476 2.412 6540.v4.c1 AC NC 55.351 >50 11.334 6545.v4.c1 AC NC 113.948 >50 35.556 0815.v3.c3 ACD NC 26.284 1.644 4.530 3103.v3.c10 ACD NC 215.876 17.045 30.725 MuLV Neg. Neg. >528 >50 >50 Pro A/G = Affinity purified antibody from a Protein A/G column; gp120 = Affinity purified antibody from a gp120 column; gp120-IgG1 = Affinity purified antibody from a gp120 followed by IgG1 column; IC50 = Inhibitory concentration 50 (ug/ml); T/F = transmitted/founder; NC = not characterized.
[1023] Plasma from patient N60 was purified and tested against a 118 multitier and multiclade pseudovirus panel. Parent sample demonstrates considerable breadth, which was also seen in the gp120 and gp120-IgG1 fractions.
TABLE-US-00007 TABLE 6 Plasma neutralization potency and breadth from 2008-2013. Plasma from patient N60 was tested against a panel of multiclade HIV pseudoviruses, demonstrating potency and breadth for more than 4 years. Numerical values given as ID50, the Inhibitory Dose 50. Plasma ID50 Titer in TZM.bl Cells (1/x) Oct. 1, Oct. 12, Sep. 9, Jan. 10, Virus Tier Clade 2008 2009 2010 2013 6535.3 1B B 3,514 2,028 2,178 1,713 QH0692.42 2 B 530 422 545 335 SC422661.8 2 B 973 630 650 1,008 PVO.4 3 B 1,384 548 800 2,046 TRO.11 2 B 1,820 1,074 820 1,174 AC10.0.29 2 B 3,655 1,884 1,709 1,586 RHPA4259.7 2 B 1,988 1,543 3,259 2,089 THRO4156.18 2 B 1,149 721 438 506 REJO4541.67 2 B 2,112 1,576 1,133 2,143 TRJO4551.58 3 B 269 479 395 728 WITO4160.33 2 B 268 221 127 732 CAAN5342.A2 2 B 6,032 2,197 1,367 1,906 Du156.12 2 C NT 1,000 NT 1,343 Du172.17 2 C NT 97 NT 468 Du422.1 2 C NT 636 NT 581 ZM53M.PB12 2 C NT 3,183 NT 1,715 ZM135M.PL10a 2 C NT 221 NT 425 ZM197M.PB7 1B C NT 455 NT 1,062 ZM214M.PL15 2 C NT 412 NT 338 Q23.17 1B A NT 3,592 NT 2,895 Q259.d2.17 2 A NT 2,873 NT 1,528 Q461.e2 2 A NT 77 NT 184 Q168.a2 2 A NT 1,303 NT 942 3415.v1.c1 2 A NT 1,097 NT 1,318 0439.v5.c1 2 A NT 499 NT 588 0260.v5.c1 2 A NT 3,519 NT 1,957 3365.v2.c20 2 A NT 869 NT 1,425 T257-31 3 CRF02_AG NT 97 NT 316 263-8 2 CRF02 AG NT 493 NT 1,104 211-9 2 CRF02_AG NT 108 NT 780 MuLV (Neg) Neg control Neg control 24 30 <20 186 NT = not tested
[1024] The broadly neutralizing plasma antibodies were isolated from N60 plasma by affinity chromatography with monomeric gp120 (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Infect Dis 213, 156-164 (2016); Sajadi et al., J Virol 86, 5014-5025 (2012)) (Table 5). The recovered gp120 affinity fraction from N60 is known represent approximately 2% of the starting mass of IgG antibody. Similar recoveries (0.6%-2% of starting mass of IgG antibody) of anti-gp120 antibodies were obtained from the plasma of other HIV infected individuals. In accordance with our previous studies of anti-HIV humoral responses (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Infect Dis 213, 156-164 (2016); Sajadi et al., J Virol 86, 5014-5025 (2012)), broadly neutralizing activity in N60 plasma was recovered by sequential protein A/G affinity chromatography, anti-gp120, and anti-IgG1 affinity chromatography (Table 5).
[1025] Two lines of evidence showed that few if any epitope specificities beyond those on monomeric gp120 were linked with the N60 polyclonal neutralizing response. First, the plasma antibody fraction that flowed through the gp120 affinity column exhibited negligible neutralization activity against a panel of 12 HIV Tier 1-3 pseudoviruses, compared to unfractionated plasma (Table 5). Second, the neutralizing breadth of purified N60 anti-gp120 IgG1 Ig largely matched that of unfractionated plasma (Table 5).
TABLE-US-00008 TABLE 7 ID50 and ID80 of N60 Plasma and gp120-FT. Plasma from patient N60 as well as gp120-FT (IgG that was passed over a BaL- gp120 column) was tested against a panel of HIV pseudoviruses for neutralization. Numerical values given as dilutions of the Inhibitory Dose 50 (ID50) or the Effective Dose 50 (ED50). In each case, the gp120-FT loses potency compared to the parent plasma, confirming that the fraction affinity purified against the gp120 monomer contains the bulk of neutralizing antibodies. N60 Plasma N60 gp120FT Virus ID Tier ID50 ID80 ID50 ID80 BaL.26 1B 1,968 574 <40 <40 QH0692.42 2 549 119 <40 <40 SC422661.8 2 752 276 <40 <40 PVO.4 3 905 308 55 <40 TRO.11 2 794 298 52 <40 AC10.0.29 2 1,040 335 <40 <40 RHPA4259.7 2 1,863 608 <40 <40 THRO4156.18 2 361 90 51 <40 REJO4541.67 2 919 333 <40 <40 TRJO4551.58 3 293 102 66 <40 WITO4160.33 2 164 45 <40 <40 CAAN5342.A2 2 851 360 <40 <40 MuLV Neg control 80 <40 <40 <40
[1026] We have previously shown a light chain bias exists in the anti-HIV envelope response (Sajadi et al., J Infect Dis 213, 156-164 (2016)), so to better define the N60 neutralizing antibodies, affinity purified anti-gp120 plasma antibodies were further fractionated by affinity chromatography with antibodies selective for human .kappa. or .lamda. light chains. The bulk anti-gp120 IgG1 .kappa. or .lamda. Ig preparations were further subjected to free flow isoelectric focusing (FFE) to separate individual antibody species according to their pI. In our previous study (Sajadi et al., J Virol 86, 5014-5025 (2012)), the broadly neutralizing antibodies were localized to fractions with more basic pIs. In the current study the fractions were tested for neutralization against two Tier 2 pseudoviruses, confirming the localization of broadly neutralizing antibodies to the basic fractions (FIG. 1 and data not shown). ELISAs confirmed that the broadly neutralizing antibody fractions contained IgG reactive with gp120 and gp120 core; but were less reactive with FLSC, in which the CD4 binding site is blocked, and did not bind to D368R gp120, whose single point mutation at position 368 abrogates binding of CD4-BS antibodies to HIV-1 gp120 (FIG. 1) (Li et al., Nat Med 13, 1032-1034 (2007)). In comparison, more acidic fractions favored binding to gp120, FLSC, D368R gp120, but not the YU2-core. (FIG. 1) (Li et al., Nat Med 13, 1032-1034 (2007)).
[1027] Our basic strategy for deconvoluting the plasma anti-gp120 polyclonal response proceeded as follows. First, fractionated immunoglobulin (see below) was subjected to enzymatic digestion with trypsin, chymotrypsin, and/or Glu-C(see Methods). Peptide fragments were subjected to LC-MS to calculate mass assignments, from which their amino acid sequences could be deduced. Next, the peptide sequences were used to identify the Ig H and L chain genes from which they originated based on a selection algorithm (see below). Identified genes then served as a means to artificially reproduce each plasma anti-gp120 monoclonal antibody, which could be examined under various conditions.
[1028] Peptide sequences were aligned and assembled using as templates authentically paired Ig H and L chain amino acid sequences translated from an N60-specific Ig gene database (see Methods) derived by single-cell sequencing from bone marrow mononuclear cells and circulating plasmablasts (see Methods). One caveat to the alignment operation was that certain peptides (typically from framework regions) could redundantly align with multiple Ig H and L template pairs, thus creating random peptide assemblages. This confound was mitigated by rank ordering the Ig H and L templates according to the number of "unique" peptide alignments (i.e. did not occur match any other Ig sequence in the database; see Methods for details) they comprised. False discovery rates were held at 5% to further increase the probability that peptide sequences were properly grouped and aligned within a full length Ig sequence (see Methods). It was also important to consider that similar degrees of total template "coverage" by plasma amino acid sequences could differ substantially in the numbers of unique peptide alignments. An example of this caveat is shown in FIG. 2 with respect to H chain alignments. In both cases, there is good overall coverage, the heavy chain in FIG. 2A possessing 60% coverage by 21 peptides, and the heavy chain in FIG. 2B possessing 69% coverage derived by 25 peptides. The heavy chain in FIG. 2A matches but only one unique, template-specific alignment (pink). The other covered areas (blue) involve sequences that bind multiple Ig gene templates, mostly in the framework regions. Alignments such as these were taken as false sequence assemblages. Indeed, antibody proteins reconstructed from such cases did not neutralize HIV or bind HIV envelope (not shown). Conversely, FIG. 2B represents a case where the alignment includes multiple peptides unique to the template sequence (including the CDRH1 region). Further, unique peptide matches were favored over redundant ones. These characteristics were taken to indicate valid sequence assemblages. In agreement, the antibody constructed from the Ig H and L template pair represented in FIG. 2B bound gp120 (N60P2.1) and was broadly neutralizing (see below).
[1029] The selection algorithm was applied to peptide sequences derived from three complementary fractionation approaches (FIG. 3). The criteria for identifying an H and L template pair as the genetic source of a plasma antibody was .gtoreq.4 unique peptides and 50% coverage in at least one of the H and L chain of each pair (with .gtoreq.4 unique peptides required in each H and L chain for the combined fractions), accommodating the designated false discovery rate cutoff.
[1030] In the primary approach, FFE fractions of anti-gp120 plasma antibodies were evaluated individually to score and select corresponding H and L template pairs. As expected, adjacent fractions rendered similar determinations within certain portions of the FFE fraction series. This operation identified 8 paired H and L Ig genes encoding plasma antibodies targeting 3 epitopes. A second approach applied the bulk polyclonal anti-gp120 antibodies to preparative IEF gels. Immunoglobulins were extracted from sequential slices of the gels and digested to obtain peptide sequences, which were then compared against the entire Ig gene database. This operation identified all but one of the H and L sequence pairs found in the primary approach as well as 4 additional ones. A third approach generated peptides and their corresponding sequences directly from bulk anti-gp120 plasma antibodies and combining this with the gel digests. This exercise mitigated the risk that sequences were overlooked in the other methods due to protein loss but necessitated combining 27 separate digests. This approach identified most of the same H and L sequence pairs found by the other approaches (missing 2 but identifying 1 additional one).
[1031] The Fab sequences of the 14 identified H and L gene pairs were combined with a generic IgG1 Fc domain (CH1-3 from IGHG1*03) in order to construct synthetic monoclonal antibody expression plasmids from which to generate protein (Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009); Guan et al., Proc Natl Acad Sci USA 110, E69-78 (2013)). This construction strategy was appropriate, as the native plasma neutralizing antibodies were of the IgG1 isotype (Table 5).
[1032] FFE was then used to compare the electrophoretic behaviors of the reconstructed mAbs versus bulk polyclonal anti-gp120 plasma antibodies. As shown in FIG. 3, the plasma antibodies tended to elute at highest concentrations within two series of fractions peaking at lower and higher pH ranges in the gradient (which ranged pH 6.5 to pH 9.5). Fractions containing plasma antibody peptide/sequences that facilitated mAb reconstruction (see above and Methods) overlapped these regions. In the case of IgG .kappa. certain fractions near the center of the gradient (.about.45-55) did not yield any unique antibodies (the peptides that derived from this region did match some of the neighboring mAbs on either side, but at a lower frequency). Similarly for IgG .lamda., fractions above pH 8 contained too little protein to perform downstream analysis. Importantly, the pI values of the reconstructed mAbs derived from the FFE or gel isoelectric focusing approaches corresponded to those of the plasma fractions that yielded their identifying peptides (compare bars and arrows in FIG. 3). This consistency indicated that the identified and reconstructed mAbs contained assemblages of amino acid sequences authentic to the plasma antibodies.
[1033] The reconstructed mAbs were characterized for source-cell subset and lineage relationships (Table 8, FIG. 4). All of the anti-gp120 mAb sequences were found in bone marrow 138- and 138+ populations by single cell PCR (summarized in Table 8). Only one mab (N60P22) could be detected in the circulating plasmablast population (Table 8). This mAb had the second highest frequency of any in the bone marrow, so it could be that the frequency of the mAbs in the plasmablast population is less than the bone marrow, or they can only be detected for a short time, as occurs with vaccination or acute infections. A homology search (multiple sequence alignment) of the bone marrow database did not reveal any of the ancestral forms of any of the mAbs identified, implying that even though the mAbs were found in all bone marrow compartments, including the long-lived plasma cells (CD138+, CD19-), their longevity may be limited.
TABLE-US-00009 TABLE 8 Characteristics of anti-gp120 plasma antibodies isolated. Frequency of CDR3 (per 1000) Single Cell Sequencing SUM Bone Bone Heavy Light (Heavy/ Peripheral marrow marrow mAb Epitope Chain chain Light) plasmablast 138- 138+ Lineage 1 N60P2.1 CD4-BS 1-2 .kappa. 1-5 38%/29% 0 2.48 0 N60P1.1 CD4-BS 1-2 .kappa. 1-5 36%/26% 0 2.48 1.62 N60P25.1 CD4-BS 1-2 .kappa. 1-5 33%/26% 0 2.48 0 N60P31.1 CD4-BS 1-2 .kappa. 1-5 42%/35% 0 2.48 0 Lineage 2 N60P22 CD4-BS 4-31 .kappa. 3-20 9%/11% 1.62 2.48 9.69 N60P38 CD4-BS 4-31 .kappa. 3-20 9%/11% 0 2.48 0 Lineage 3 N60P30 CD4i 1-2 .kappa. 3-20 21%/12% 0 2.48 0 (Cluster A) Lineage 4 N60P36 CoR-BS 1-69 .kappa. 3-20 11%/9% 0 4.96 4.85 (Cluster C) Lineage 5 N60P39 CoR-BS 1-69 .kappa. 3-20 11%/9% 0 2.48 1.62 (Cluster C) N6039.1 CoR-BS 1-69 .kappa. 3-20 11%/9% 0 2.48 1.62 (Cluster C) N60P47 CoR-BS 1-69 .kappa. 3-20 16%/6% 0 2.48 0 (Cluster C) N60P48 CoR-BS 1-69 .kappa. 3-20 15%/6% 0 4.96 6.46 (Cluster C) Lineage 6 N60P51 CoR-BS 1-69 .kappa. 3-20 20%/9% 0 12.4 3.38 (Cluster C) Lineage 7 N60P35 V3 5-51 .lamda. 6-57 18%/14% 0 2.48 4.85 N60P37 V3 5-51 .lamda. 6-57 17%/16% 0 2.48 6.46 mAb = monoclonal antibody. SHM = somatic hypermutation. CD4-BS = CD4-binding site antibody. CoR-BS = Co-Receptor binding site antibody. Cluster A = competes with A32. Cluster C = competes with 19e and/or 17b.
[1034] Overall, the dominant N60 plasma anti-gp120 response arose from 7 distinct lineages (Table 8), as shown in the neighbor-joining phylogeny tree (see Methods) of the entire BM antibodies (FIG. 4). Lineage 1 was distinguished by 1-2 heavy chain and 1-5 .kappa. light chain gene usage as well as a relatively high degree of somatic hypermutation (33-42% in the heavy chain). Lineage 1 mAbs resemble previously reported broadly neutralizing anti-CD4-BS antibodies, which are also assigned to VH1-2 and V.kappa.1-5 gene families that exhibit the signature deletion in the CDRL3 (Scharf et al., Proc Natl Acad Sci USA 110, 6049-6054 (2013); West et al., Proc Natl Acad Sci USA 109, E2083-2090 (2012); Zhou et al., Science 329, 811-817 (2010)). These antibodies had basic pIs. Two mAbs (N60P1.1 and N602.1) bound gp120 on Elisa and SPR, and but not the D368R point mutant, and thus are CD4 binding site antibodies (FIGS. 5 and 6). However, mAbs N60P25.1 and N60P31.1 in this family did not bind gp120 in the standard antigen capture ELISA format (FIG. 5) using plates coated with antibody D7324 directed against the gp120 C terminal region (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Virol 86, 5014-5025 (2012); Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009)), nor in the reverse format coating plates first with mAbs (not shown). N60P25.1, when bound to protein A, did bind to gp120 in SPR, while N6031.1 did not (FIG. 6). Additionally, we did see binding to gp120 in accordance with their retention on the gp120 affinity column (26% recovery for N60P25.1, 5% recovery for N60P31.1, <1% for control mAb Synagis).
[1035] Lineage 2 was distinguished by 4-31 heavy chain and 3-20 .kappa. light chain usage and a much lower degree of hypermutation (9% in the heavy chain). The two members of this lineage also had basic pIs and appeared to be directed against the CD4 binding site (FIGS. 5 and 6) as determined by ELISA (FIG. 5). These mAbs were distinguished from Lineage 1 antibodies by the capacity to recognize a YU2 gp120 core antigen. Genes encoding one mAb in this lineage (mAb N60P22) were detected in circulating plasmablasts.
[1036] Lineage 3 contained one member (mAb N60P30) with 1-2 heavy chain and 3-20 .kappa. light chain usage and a moderate degree of hypermutation (21% in the heavy chain). N60P30 had a basic pI, bound well to FLSC, but not to gp120 or YU2-core in ELISA (FIG. 5), indicating a specificity for CD4-induced epitopes. Competition Elisa testing revealed that N60P30 competed with A32 but not 17b, signifying Cluster A specificity (data not shown).
[1037] Lineage 4 contained one member (mAb N60P36) with 1-69 heavy chain and 3-20 .kappa. light chain usage and a relatively low degree of hypermutation (11% in the heavy chain). N60P36 had a neutral pI. Binding assays (FIG. 5) and competition ELISAs with CD4i mAbs (data not shown) indicated that mAbs in this lineage recognize the Cluster C epitope in the coreceptor binding site.
[1038] Lineage 5 mAbs were distinguished by 1-69 heavy chain and 3-20 .kappa. light chain usage and a moderate degree of hypermutation (11-16% in the heavy chains). This Lineage comprised of 3 members (mAbs N60P39, N60P39.1, and N60P48). Binding assays (FIG. 5) and competition ELISAs with CD4i mAbs (data not shown) indicated that mAbs in this lineage recognize the Cluster C epitope in the coreceptor binding site. We identified one additional mAb that was not picked up with the above methods by a homology search of the bone marrow database. This mAb (N60P47) had no binding to gp120 on elisa, and thus had either no binding to gp120, as in the case of antibodies targeted at the hybrid epitope of CD4 and gp120, or bound to gp120 so weakly that too little was recovered to identify correctly.
[1039] Lineage 6 contained one member (mAb N60P51) with 1-69 heavy chain and 3-20 .kappa. light chain usage and a moderate degree of hypermutation (20% in the heavy chain). Binding assays (FIG. 5) and competition ELISAs with CD4i mAbs (data not shown) indicated recognition of Cluster C epitope in the coreceptor binding site.
[1040] Lineage 7 was distinguished by 5-51 heavy chain and 3-20 .lamda. light chain usage and a moderate degree of hypermutation (17-18% in the heavy chains). mAbs in this family had more neutral pIs. Binding analyses indicated that the two members of this lineage bound Yu2 core+V3, but not the Yu2 core on Elisa (FIG. 5). Additionally, both mAbs bound the HIV-1 MN Complete V3 Loop Peptide on Elisa (FIG. 5), indicating that these mAbs recognize the V3 loop of HIV-1.
[1041] In a previous study of N60 (Sajadi et al., J Infect Dis 213, 156-164 (2016)), we determined that roughly 50% of the total anti-gp120 plasma response involved antibodies with .lamda. light chains. As only Lineage 7 expressed .lamda. light chains, it is evident that cross-reactive anti-V3 antibodies account for the bulk of the circulating anti-gp120 response in N60 (potentially up to 1% of the total circulating IgG). Such representation in the polyclonal anti-gp120 response is in accordance with the immunodominant nature of the V3 loop as evinced in studies of anti-gp120 responses in other cohorts (Javaherian et al., Proc Natl Acad Sci USA 86, 6768-6772 (1989); LaRosa et al., Science 249, 932-935 (1990); Vogel et al., J Immunol 153, 1895-1904 (1994)).
[1042] Preliminary screening of the mAbs for neutralizing activity against 15 tiers 1-3 pseudoviruses showed that Lineage 1 comprised the most broad and potent activity, followed by Lineages 2, 7, and 5 (Table 9).
TABLE-US-00010 TABLE 9 mAb screening ELISA and neutralization. For BaL-gp120 ELISA binding, background subtracted OD results shown for mAbs tested at 8 ug/ml. For HIV-1 neutralization, a neutralization was carried out against a panel of Tier 1-3 pseudoviruses (see Methods). mAb Epitope SF162.LS BaL.26 SS1196.1 6535.3 QH0692.42 SC422661.8 PVO.4 TRO.11 AC10.0.29 Lineage N60P1.1 CD4-BS 12.919 >50 0.612 3.835 15.479 0.75 0.456 6.315 >25 1 N60P2.1 CD4-BS >50 >50 1.836 >25 2.995 12.690 1.683 19.597 >25 N60P25.1 CD4-BS 30.992 >50 0.246 15.725 32.942 2.056 6.454 23.528 >50 N60P31.1 CD4-BS >50 >50 >50 >50 >50 >50 >50 1.807 >50 Lineage N60P22 CD4-BS 1.275 5.532 5.673 10.641 36.840 >50 >50 >50 >50 2 N60P38 CD4-BS 5.765 >50 7.979 >50 >50 >50 >50 >50 >50 Lineage N60P30 CD4i >50 >50 >50 >50 >50 >50 >50 >50 >50 3 (ClusterA) Lineage N60P36 CoR-BS >50 >50 >50 >50 >50 >50 >50 >50 >50 4 (ClusterC) N60P39 CoR-BS 3.189 >50 4.611 >50 >50 >50 >50 >50 >50 (ClusterC) Lineage N6039.1 CoR-BS 0.482 14.124 1.205 >50 45.230 >50 >50 >50 >50 5 (ClusterC) N60P47 CoR-BS 0.684 20.084 1.905 >40 >40 >40 >40 >40 >40 (ClusterC) N60P48 CoR-BS 0.395 12.729 .907 >40 20.238 >40 >40 >40 >40 (ClusterC) Lineage N60P51 CoR-BS .318 8.963 1.033 >25 >25 >25 >25 >25 >25 6 (ClusterC) Lineage N60P35 V3 .021 4.68 1.113 >25 >25 >25 >25 >25 >25 7 N60P37 V3 0.03 4.076 .991 >25 >25 >25 >25 >25 mAb Epitope RHPA4259.7 THRO4156.18 REJO4541.67 TRJO4551.58 WITO4160.33 CAAN5342.A2 Lineage N60P1.1 CD4-BS 4.218 >25 1.581 7.195 >25 1.635 1 N60P2.1 CD4-BS 15.684 >25 >25 >25 >25 6.733 N60P25.1 CD4-BS 1.448 >50 2.454 >50 >50 2.502 N60P31.1 CD4-BS >50 31.038 >50 >50 >50 >50 Lineage N60P22 CD4-BS >50 36.501 >50 >50 >50 >50 2 N60P38 CD4-BS >50 >50 >50 >50 >50 >50 Lineage N60P30 CD4i >50 >50 >50 >50 >50 >50 3 (ClusterA) Lineage N60P36 CoR-BS >50 >50 >50 >50 >50 >50 4 (ClusterC) N60P39 CoR-BS >50 >50 >50 >50 >50 >50 (ClusterC) Lineage N6039.1 CoR-BS >50 >50 >50 >50 >50 >50 5 (ClusterC) N60P47 CoR-BS >40 >40 >40 >40 >40 >40 (ClusterC) N60P48 CoR-BS >40 >40 >40 >40 >40 >40 (ClusterC) Lineage N60P51 CoR-BS >25 >25 >25 >25 >25 >25 6 (ClusterC) Lineage N60P35 V3 3.733 >25 >25 >25 >25 >25 7 N60P37 V3 11.624 >25 >25 >25 >25 >25
[1043] Expanded testing of Lineage 1 mAbs revealed neutralization breadth approaching the coverage observed with the polyclonal plasma anti-gp120 kappa Ig (FIG. 7). Two mAbs, N60P1.1 and N60P25.1, demonstrated particularly strong neutralizing activity (FIG. 7), matching 70% and 73% of the affinity purified plasma Ig breadth, respectively. Thus, broad plasma neutralizing activity in N60 appears to largely arise from Lineage 1, representing a monospecific fraction of the anti-gp120 repertoire at the time point evaluated. Anti-CD4BS mAbs from Lineage 2, which appeared in high frequency but did not have strong breadth and potency, contributed minimally to the overall plasma profile. In any case, the concurrence of other types of poorly neutralizing anti-gp120 antibodies does not appear to abrogate the presence and activity of broadly neutralizing antibodies in circulation.
[1044] Despite their breadth and potency, none of the anti-CD4BS mAbs from Lineage 1 matched the full breadth of the polyclonal plasma anti-gp120 Ig recovered from N60. Considered collectively, the anti-CD4BS mAbs neutralized 89% of the viruses that were sensitive to bulk anti-gp120 plasma Ig. Resistance to the mAbs was independent of virus clade or Tier (FIG. 7). Notably, one resistant pseudovirus was neutralized by the Lineage 5 anti-CD4i mAb, N60P39. Thus, the combined profiles of six mAbs including N60P39 could cover 90% of the viruses neutralized by bulk anti-gp120 plasma Ig. To determine if greater breadth could be established by mAb mixtures, the panel of viruses sensitive to bulk anti-gp120 plasma Ig were tested against an equimolar pool of all anti-CD4 BS mAbs, or an equimolar combination of represented mAbs from all lineages (related clones with <5% amino acid sequence diversity were not included). These mAb pools covered, respectively, 89 and 79% of the neutralizing activity breadth mediated by the plasma Ig (FIG. 7). These results suggest that the plasma neutralizing response in N60 could involve a cryptic anti-gp120 specificity (active against a small subset of test viruses resistant to the identified mAbs) and/or a molar ratio of specificities that we could not readily duplicate in vitro.
[1045] Previously we reported that multiple Clade B HIV infected patients expressed broadly neutralizing plasma antibody responses with similar biochemical characteristics (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Infect Dis 213, 156-164 (2016).
[1046] Sajadi et al., J Virol 86, 5014-5025 (2012)), such as basic pIs. We posited that this trend reflected shared inter-subject specificity for neutralizing gp120 epitopes. To test this hypothesis, we applied the same deconvolution procedures and selection algorithms described above to another NVS cohort subject, N49, who exhibited a very broad neutralization response against 99% of a 117 virus panel (FIG. 8). Similar to the N60 responses, the neutralizing antibodies from N49 plasma could be recovered from plasma by gp120 affinity chromatography (recovered gp120 affinity fraction known to be approximately 1% of the starting mass of IgG antibody for this patient); however, in this case the .lamda. fraction contained the broad neutralizing antibodies.
[1047] The broadly neutralizing antibodies in N49 plasma fell into two lineages, distinguished by different light chain gene usage (Table 10). Similar to the N60 Lineage 1 broadly neutralizing antibodies, the N49 mAbs all exhibited basic pIs and VH1-2 gene usage. However, all of the N49 mAbs used .lamda. light chain genes, while also containing a CDRL3 deletion. The binding characteristics of this N49 lineage also matched the N60 neutralizing antibodies, reflecting anti-CD4BS specificity. These antibodies bind to monomeric gp120, have little to no binding to D368R and FLSC in both Elisa and biacore (FIGS. 9 and 10).
TABLE-US-00011 TABLE 10 Properties of mAbs engineered from NVS49. Frequency of CDR3 (per 1000) Heavy Light Lambda Heavy Peripheral Bone marrow Bone marrow mAb Epitope pI chain chain Constant mutation plasmablast 138- 138+ N49P6 CD4-BS 8.9 1-2 2-11 LC7 42%/28% 0 0 0.49 N49P7 CD4-BS 8.8 1-2 2-11 LC2 38%/31% 0 0 0.49 N49P7.1 CD4-BS 8.8 1-2 2-11 LC2 38%/31% 0 0 0.49 N49P11 CD4-BS 8.4 1-2 2-11 LC2 35%/30% 3.18 0 0 N49P9 CD4-BS 8.7 1-2 2-23 LC7 38%/42% 3.18 0 0.49 mAb = monoclonal antibody. VDJ = Variable Diversity Junction. LC = Lambda constant. AA = amino acid. Mutation = somatic hypermutation given as a percent of germline V region for heavy chains, and VJ region for light chains. NT = not tested
[1048] A distinguishing feature of the N49 anti-CD4BS mAbs was that they exhibited remarkably broad neutralizing activity when tested against a multi-clade, tier 1-3 panel of 117 pseudoviruses. As shown in FIG. 8, N49 mAbs N49P6, N49P7, and N49P11 individually neutralized 100% of the viruses. The N49 P series mAbs also had high potency, with ability to neutralize 41.5-86.4% of all viruses at under 1 ug/ml. MAb N49 P7 had complete breadth (ability to neutralize all 117 pseudoviruses) and the highest potency (86.4%). The breadth and potency of the N49 group of mAbs was compared other mAbs reported to have substantial breadth (PGT121, PGT128, PGT145, PGDM1400, PGT151,10-1074,10E8, PG9, PG16, 3BNC117, NIH45-46, 8ANC195, VRC07, and VRC01), using the same neutralization assay and panel. The breadth of the N49 mAbs surpassed those of all mAbs tested (Table 11). Although, we did not test another recently reported broad neutralizing antibody against the CD4BS, N6 (Williams et al., Science Immunology 2, (2017)), four of the N49 mAbs could neutralize T278-50, which is N6 is resistant to. N49 was fundamentally distinguished from N60 in the sense that three identified plasma mAbs (N49P6, N49P7, and N49P11) completely recapitulated the antiviral breadth of affinity purified anti-gp120 plasma IgG.
TABLE-US-00012 TABLE 11 Comparison of neutralization breadth between those derived from NVS49 and those in the literature. Percentages are given as values of viruses neutralized (IC50 < 50 ug/ml)/viruses tested. All testing was done in the same lab. N49 P series tested against the full 117 virus panel. All other mAbs tested against 114-117 pseudoviruses, except for VRC01, which was tested against 96. Neutralization at titers >50 .mu.g/mL were considered non-neutralizing. % mAb neutralization N49P6 100% N49P7 100% N49P7.1 .sup. 99% N49P11 100% N49P9 .sup. 89% PGT121 65.8% PGT128 59.8% PGT145 .sup. 74% PGDM1400 79.8% PGT151 65.5% 10-1074 56.8% 10E8 94.9% PG9 84.7% PG16 82.2% 3BNC117 .sup. 90% NIH 45-46 86.4% 8ANC195 65.2% VRC07 91.5% VRC01 87.6%
Materials and Methods
Patients.
[1049] The patients identified for this study was selected from the NVS study (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Acquir Immune Defic Syndr 50, 403-408 (2009); Sajadi et al., AIDS 21, 517-519 (2007)). NVS patients are defined as having HIV-1 by Western Blot while having an HIV-1 RNA <400 copies/ml for at least 4 measurements and 2 years. N60 met the above definition, while N49 had a higher viral load setpoint, averaging 7,854 HIV-1 copies/ml over 9 years. Both of the patients' serum were identified as having broad neutralizing activity based on Tier 2 activity and a cross-clade neutralization panel (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al., J Infect Dis 213, 156-164 (2016); Sajadi et al., J Virol 86, 5014-5025 (2012).
Proteins and Antigens.
[1050] Recombinant HIV-1 antigens were generated as described previously (31). Test antigens included the YU2 gp120 core, from which V1, V2, and V3 have been deleted (Wu et al., Nature 384, 179-183 (1996)); the YU2 gp120 core containing the V3 loop (YU2 gp120 core +V3) (Wu et al., Nature 384, 179-183 (1996)); monomeric HIV-1 Ba-L gp120 (Fouts et al., J Virol 74, 111427-111436 (2000)); D368R Ba-L gp120, which has a single point mutation at position 387 (Li et al., Nat Med 13, 1032-1034 (2007)); and a single chain gp120-CD4 complex (FLSC) presenting a full length CD4-induced Ba-Lgp120 structure in which the CD4 binding site is occupied (Fouts et al., J Virol 74, 111427-111436 (2000)). Two monoclonals specific for the C-terminal peptide of HIV-1 gp120 were sued: an affinity-purified goat Ab, D7324, purchased from Cliniqa (San Marcos, Calif.) and JR52 a mouse monoclonal. All proteins were expressed by transient transfection of 293T cells as previously describe and purified by lectin affinity chromatography as previously described and dialyzed against PBS prior to use (Fouts et al., J Virol 74, 111427-111436 (2000)). The following reagent was obtained through the NIH AIDS Reagent Program, AIDS Program, NIAID, NIH: HIV-1 V3 Peptides from the Division of AIDS, NIAID.
Isolation of Plasma Antibody Species.
[1051] Whole plasma IgG was purified on a Protein A or Protein A/G affinity chromatography column (GE Healthcare, Piscataway, N.J.) according to the manufacturer's instructions and dialyzed against PBS prior to use. Affinity chromatography columns were made with activated CH Sepharose beads (GE Healthcare, Piscataway, N.J.) coupled to 2 mg of recombinant HIV-1 Ba-L gp120 (Fouts et al., J Virol 74, 111427-111436 (2000)), as described previously (Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009)). Beads specific for human IgG1, human .kappa. chain, and human .lamda. chain were purchased from Capture Select (Naarden, Netherlands). The columns were used to purify antigen-specific IgG (anti-gp120), fractionate IgG1 from whole IgG, or fractionate IgG into .kappa. and .lamda. fractions, as previously described (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009). Briefly, IgG was incubated with beads at 37.degree. C. for one hour prior to extensive washing with PBS. Columns were eluted at room temperature with pH 2.8 0.2M glycine (for elution of .kappa. antibodies pH 2.0 was used) and dialyzed against 4 liters PBS 3 times (a minimum of 24 hours total) prior to testing. Dedicated columns were used for each subject and antigen. IgG concentration was measured using an in-house quantitative ELISA as previously described (Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009)). After a series of steps, the plasma was fractionated into IgG1 .kappa. and IgG1 .lamda. antibodies (plasma->protein A column->IgG1 column->kappa and lambda columns), anti-gp120 .kappa. and anti-gp120 .lamda. antibodies (plasma->protein A column->gp120 column->kappa and lambda columns), or anti-gp120 antibodies (plasma->protein A column->gp120 column).
[1052] Affinity purified and fractioned antibody was subjected to free flow electrophoresis on the BD Free Flow Electrophoresis System (BD, Franklin Lakes, N.J.). The separation, stabilization and counter flow media was freshly prepared according to instructions of the manufacturer. The separation and counter flow media contained 0.2% hydroxypropyl methylcellulose (HPMC). The pH range of separation media was 0.88 to 12.8. The media flow rate in the separation chamber was 41 mL/hour. The antibodies (200 to 350 .mu.g/ml) were introduced to separation chamber at the rate of 560 .mu.l/h in the electrical field of 2300V/10 mA/24 W. IEF fractionated samples collected in a 96 deep-well polystyrene microtiter plate, with each well containing 1-2 ml. Approximately half of these wells contained antibody fractionated based on PI. Fractionation was confirmed with pH reading of individual fractions (FIG. 1), as well as an IEF gel. Separately, affinity purified antibody fractions (prior to FFE) were also run on IEF gels.
[1053] Neutralization assay. HIV-1 neutralization testing was performed using a luciferase-based assay in TZM.b1 cells as previously described (Sajadi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Li et al., J Virol 79, 10108-10125 (2005)). This assay measures the reduction in luciferase expression following a single round of virus infection. Stocks of Env-pseudotyped viruses were prepared by transfection of 293T/17 cells as previously described (Li et al., J Virol 79, 10108-10125 (2005)). Unfractioned serum samples, affinity purified antibody, fractionated affinity purified IgG samples, and mAbs were tested against MuLV control and a panel of psuedoviruses. Three-fold serial dilutions of IgG were tested in duplicate (96-well flat bottom plate) in 10% D-MEM growth medium (100 ul/well). 200 TCID50 of pseudovirus was added to each well in a volume of 50 ul and the plates were incubated for 1 hour at 37.degree. C. TZM.b1 cells were then added (1.times.10.sup.4/well in 100 ul volume) in 10% D-MEM growth medium containing DEAE-Dextran (Sigma, St. Louis, Mo.) at a final concentration of 11 ug/ml. The final volume for each well was 250 ul. Assay controls included replicate wells of TZM.b1 cells alone (cell control), TZM.b1 cells with virus (virus control), and MuLV control. Following a 48 hour incubation at 37.degree. C., 150 ul of assay medium was removed from each well and 100 ul of Bright-Glo luciferase reagent (Promega, Madison, Wis.) was added. The cells were allowed to lyse for 2 minutes, then 150 ul of the cell lysate was transferred to a 96-well black solid plate and luminescence was measured using a Victor 3 luminometer (Perkin Elmer, Waltham, Mass.). The 50% inhibitory concentration (IC50) and 80% inhibitory concentration (IC80) titers were calculated as the immunoglobulin concentration that caused a 50% or 80% reduction in relative luminescence units (RLU) compared to the virus control wells after subtraction of cell control RLUs (Li et al., J Virol 79, 10108-10125 (2005)).
ELISA.
[1054] HIV-1 envelope capture ELISAs were performed as previously described (Guan et al., Proc Natl Acad Sci USA 106, 3952-3957 (2009)) with various antigens (as indicated in the text) that were directly coated (HIV-1 Ba-L SOSIP trimer, 1 ug/ml; YU2 gp120 core construct and YU2 gp120 core plus V3; 2 ug/ml) or captured (Bal-gp120 or FLSC at a concentration of 0.15 ug/ml) by antibody D7324 or JR52 that had been adsorbed to the solid phase at 2 ug/ml. For IEF-fractionated affinity purified IgG, 5 ng from each fraction was tested in a total assay volume of 50 ul. All IgG preparations were incubated with antigens for 1 hour at 37.degree. C. Bound Abs were then detected with 1:1,000-diluted alkaline phosphatase (AP)-goat antihuman IgG (Southern Biotech; Birmingham, Ala.) and detected with Blue Phos Microwell Phosphatase Substrate System (KPL, Gaithersburg, Md.). All assays were performed in duplicate or repeated several times. Negative control assays were carried out with secondary antibody; background values were subtracted from all test absorbance readings.
Isolation of Plasma mAbs.
[1055] Antibody species that were isolated to individual fractions were subjected to LC-MS (in addition to FFE fractions, several experiments were carried out with affinity purified fractions or cut-out IEF bands from an IEF gel). Antibody was digested with trypsin, chymotrypsin, or Glu-C overnight at 37.degree. C., the peptides evaporated to 15u1. The LC-MS system consisted of a Thermo Electron Orbitrap Velow ETD mass spectrometer with a Protana nanospray ion source interfaced with a Phenomenex Jupiter C18 reversed-phase capillary column. The peptide digest was fragmented with both CID and HCD. LC-MS was performed at the University of Maryland School of Pharmacy and Northwestern Proteomics Center of Excellence, none of which were involved in the data analysis. The spectra were searched with Peaks software (Bioinformatics Solutions Inc, Ontario, Calif.) against multiple B cell databases generated from the patient described below.
Single Cell Sorting.
[1056] Single-cell sorting and sequencing was done at Atreca (Redwood City, Calif.) on PBMC memory B cells (Yu2-gp140 reactive), PBMC plasmablasts, and bone marrow plasma cells and patient-specific B cell databases generated. All paired chain antibody sequencing was carried out on IgG cells sorted into microtiter plates at one cell per well by FACS. IgG plasmablasts were enriched from cryopreserved peripheral blood mononuclear cells (PBMCs) by gating for CD3-CD14-CD16-CD19+CD20-CD27+CD38.sup.hirIgA-IgM-IgD-cells. Antigen-specific cells were isolated from PBMCs using fluorescently-labeled YU2 gp140 (43) and cultured for 4 days prior to single cell sorting in IMDM medium (Invitrogen) in the presence of FBS, Pen/Strep, IL-2 (PeproTech), IL-21 (PeproTech), and rCD40 ligand (R&D Systems). In some experiments, the bone marrow plasma cells (CD3-CD14-CD16-CD38.sup.hiIgA-IgM-IgD-) were further sorted and analyzed based on CD19 and CD138.
Paired Chain Antibody Sequencing.
[1057] Generation of barcoded cDNA, PCR ampification, and 454 sequencing of IgG were performed as described in Tan et al. 2014, with the following modifications: biotinylated Oligo(dT) and RT maxima H- (Fisher Scientific Company) were used for reverse transcription, cDNA was extracted using Streptavidin C1 beads (Life Technologies), DNA concentrations were determined using qPCR (KAPA SYBR.RTM. FAST qPCR Kit for Titanium, Kapabiosystems), and amplicons were sequenced using Roche 454 Titanium sequencing.
Barcode Assignment, Sequence Assembly, Assignment of V(D)J and Identification of Mutations.
[1058] These steps were performed as previously described (Tan et al., Clin Immunol 151, 55-65 (2014)), except for the following: a minimum coverage of 10 reads was required for each heavy and light chain assembly to be acceptable. Wells with more than one contig for a chain were rejected from consideration unless one of the contigs included at least 90% of the reads. V(D)J assignment and mutation identification was performed using a variant of SoDA (Volpe et al., Bioinformatics 22, 438-444 (2006)). Antibody amino acid sequences were aligned to heavy and light chain hidden Markov models using hmmalign (http://hmmer.org). The resulting multiple sequence alignments were used to generate a neighbor-joining tree with RapidNJ (Simonsen M, Pedersen CNS, in WABI 2008, L. J. Crandall K A, Ed. (Springer, Heidelberg, 2008), vol. 5251, pp. 113-122).
Mass Spectrometry Analysis and Generation of Plasma Antibodies.
[1059] Antibody species that were isolated to individual fractions were subjected to LC-MS (in addition to FFE fractions, several experiments were carried out with affinity purified fractions or cut-out IEF bands from an IEF gel). Antibody was digested with trypsin, chymotrypsin, or Glu-C overnight at 37.degree. C., the peptides evaporated to 15 .mu.l. The LC-MS system consisted of a Thermo Electron Orbitrap Velow ETD mass spectrometer with a Protana nanospray ion source interfaced with a Phenomenex Jupiter C18 reversed-phase capillary column. The peptide digest was fragmented with both CID and HCD. LC-MS was performed at the University of Maryland School of Pharmacy and Northwestern Proteomics Center of Excellence, none of which were involved in the data analysis. The spectra were searched with Peaks software (Bioinformatics Solutions Inc., Ontario, Calif.) against multiple B cell databases generated from the patient described above.
[1060] An array of whole IgG H and L amino acid sequences were translated from the database and used as a basis for interpreting the peptide data. The LC-MS derived spectra were searched against the databases independently using the following settings: Parent Mass Error Tolerance 5.0 ppm, Fragment Mass Error Tolerance 0.5 Da, Fixed modification of Carboxymethyl (58.01), False Discovery Rate for peptides 5%. Potential antibodies were ranked based on number of unique peptides in the heavy and light chain sequences (.gtoreq.4 unique peptides and 50% coverage in at least one of the H and L chain of each pair or with .gtoreq.4 unique peptides required in each H and L chain for the combined fractions). The identified VH or VL region clones were cloned into an expression vector upstream to human IgG1 constant domain sequence. Minipreps of these DNA pools, derived from suspension bacterial cultures, were used to transiently transfect 293 Freestyle cells. Transfectant supernatants containing recombinant antibodies were screened in ELISA and neutralization assays.
[1061] One caveat of the alignment algorithm is that certain peptides (typically from framework regions) can redundantly align with multiple 1 g H and L template pairs, thus creating random peptide assemblages. This caveat was mitigated by rank ordering the Ig H and L templates according to the number of "unique" peptide alignments (i.e. not matching any other 1 g sequence in the database; see Methods for details) they comprised. False discovery rates were held at 5% to further increase the probability that peptide sequences were properly grouped and aligned within a full-length 1 g sequence. It was also important to consider that similar degrees of total template "coverage" by plasma amino acid sequences could differ substantially in the numbers of unique peptide alignments.
[1062] In the primary approach, FFE fractions of affinity-isolated anti-gp120 plasma antibodies were evaluated individually to score and select corresponding H and L template pairs. This identified 8 paired Hand L 1 g genes encoding plasma mAbs N60P1.1, N60P22, N6025.1, N60P36, N60P38, N60P39.1, N60P35, and N60P37. A second approach applied the bulk polyclonal anti-gp120 antibodies to preparative isoelectric focusing (IEF) gels. lmmunoglobulins were extracted from sequential slices of the gels and digested to obtain peptide sequences, which were then compared against the patient-specific 1 g gene database. This operation identified all but one of the H and L sequence pairs found in the primary approach as well as 4 additional ones: N60P2.1, N60P30, N60P31.1, N60P48.1, and N60P51. A third approach generated peptides and their corresponding sequences directly from affinity-enriched anti-gp120 plasma antibodies and combining this information with the gel digests from the second approach. This exercise mitigated the risk that sequences were overlooked in the other methods due to protein loss but required combining 27 separate digests. Even so, this approach identified most of the same H and L sequence pairs found by the other approaches (missing 2 but identifying 1 additional mAb-N60P39. We identified one additional mAb that was not picked up with the above methods by a homology search of the bone marrow database. This mAb (N60P47) had no binding to gp120 on Elisa, and thus had either no binding to gp120, as in the case of antibodies targeted at the hybrid epitope of CD4 and gp120, or bound to gp120 so weakly that too little was recovered to identify correctly.
Example 2. Broadly Neutralizing HIV Antibodies
[1063] Using a method for isolating monoclonal antibodies that match the circulating antibodies in circulation, we have isolated a series of broadly neutralizing antibodies against HIV-1. The subject these antibodies were isolated from (NVS49, also referred to as Subject 8 in previous publications) has extremely potent antibodies against HIV-1 (Saj adi et al., J Acquir Immune Defic Syndr 57, 9-15 (2011); Sajadi et al,. J Virol 86, 5014-5025 (2012); Sajadi et al., J Infect Dis 213, 156-164 (2016)). At the point of testing, the patient had HIV for at least 21 years. The patient's plasma was the time point closest to when the antibodies were derived from was able to neutralize 99% of HIV strains from around the world (Excel file "N49 neutralization and sequences"), including strains that other HIV mAbs that are undergoing clinical testing are resistant to.
[1064] We have been able to isolate 2 distinct families of antibodies from this patient. All of these antibodies have been engineered with the same IgG1 heavy chain backbone (not obtained from patient NVS49 and in fact found in a different racial group), along with VDJ sequences obtained from patient NVS49. In addition, various clones have mutations in the VDJ or constant regions, and some antibodies have swapped Lambda constant regions.
[1065] The first family comprises of N49P6, N49P7, N49P11, N49P18 and their various clones (N49P6.1, N49P6.2, N40P7.1, N49P7.2, N49P11.1, N49P18.1). The second family comprises of N49P9 and its clone N49P9.1. Both family of antibodies use the 1-2 Heavy chain family, while using 2 different Lambda light chain gene families (Lambda 2-11 and Lambda 2-23) (see Table 12 and FIG. 11 for more details).
TABLE-US-00013 TABLE 12 Properties of mAbs engineered from NVS49. Heavy Light Lambda Heavy Light mAb chain chain Constant mutation mutation pI Notes N49P6 1-2 2-11 LC7 42% 28% 8.9 N49P6.1 1-2 2-11 LC7 42% 28% 9.0 VDJ identical to N49P6 Heavy constant: 1 AA difference with N49P6 N49P6.2 1-2 2-11 LC2 42% 28% 9.0 VDJ identical to N49P6 Heavy constant: Same as N49P6.1 Lambda constant: 1 AA difference with N49P6/6.1, also LC2 not LC7 N49P7 1-2 2-11 LC2 38% 31% 8.8 N49P7.1 1-2 2-11 LC2 38% 31% 8.8 VDJ identical to N49P7 Heavy constant: 1AA difference with N49P7 Lambda constant: 1AA difference with N49P7 N49P7.2 1-2 2-11 LC2 38% 29% 8.8 Heavy: 8 AA mutation VDJ with N49P7/7.1 Heavy constant- 1 AA difference with N49P7.1 Light: DJ identical to N49P7/7.1 Light: V region 2 AA difference with N49P7/7.1 N49P11 1-2 2-11 LC2 35% 30% 8.4 Lambda constant: 1 AA mutation N49P18 1-2 2-11 LC2 38% 29% 8.7 Lambda constant: 1 AA mutation N49P18.1 1-2 2-11 LC2 34% 31% 8.7 Heavy: V region 4 AA difference with N49P18 Heavy constant: 1 AA difference with N49P18 Light: V region 3 AA difference with N49P18 N49P19 1-2 2-11 LC2 39% 28% 8.9 Lambda constant: 1 AA mutation N49P9 1-2 2-23 LC7 38% 42% 8.65 Lambda constant: 1 AA mutation N49P9.1 1-2 2-23 LC2 38% 42% NT Lambda constant: Contains N49P9 mutation but LC2 instead of LC7. mAb = monoclonal antibody. VDJ = Variable Diversity Junction. LC = Lambda constant. AA = amino acid. Mutation = somatic hypermutation given as a percent of germline V region for heavy chains, and VJ region for light chains. NT = not tested
[1066] The antibodies from these 2 families target the CD4-binding site region of HIV-gp120. Currently, we have several lines of evidence to support this. Inspection of the protein sequence of the antibodies reveal characteristic phenotype of CD4-BS antibodies: [1067] 1) Heavy chain: usage of the 1-2 Heavy chain, usage of Trp50, Asn58, and Arg71 at the noted positions [1068] 2) Light Chain: usage of a deletion in CDRL3 rendering it 5 amino acids long, as well as a deletion in CDRL1 (FIG. 12). [1069] 3) Basic isoelectric point
[1070] None of the antibodies tested thus far can bind BaL D368R mutant, while they can bind the BaL gp120 monomer. The difference in the 2 antigens is a single point mutation at position 368, which abrogates binding of CD4-BS antibodies to HIV-1 gp120 (Li et al., Nat Med 13, 1032-1034 (2007)) (FIG. 12). Finally, crystallography reveals binding the region of the CD4-BS.
[1071] Antibodies N49P6, N49P7, N49P9, and N49P11 were evaluated by their ability to neutralize HIV-1 against a panel of pseudoviruses in our lab. N49P6 and N49P7 demonstrated the best neutralization with ability to neutralize all the viruses tested (see FIG. 13 for N49P7 neutralization).
[1072] The following antibodies were sent for extensive testing at Dr. Michael Seaman's lab at Harvard University: N49P6, N49P7, N49P7.1, N49P11, and N49P9. Dr. Seaman uses a panel of 118 HIV-1 pseudoviruses that represent multi-clade and difficult to neutralize strains from around the world. Dr. Seaman's lab is a reference lab for neutralization testing, and he has worked with all of the major HIV-1 broadly neutralizing antibodies that have been created. In Dr. Seaman's panel, the antibodies tested performed better than all the antibodies he has worked with before. Specifically: N49P6, N49P7, and N49P11 demonstrated 100% neutralization breadth (able to neutralize all the viruses by IC50 in the panel), N49P7.1 demonstrated 99% neutralization, and N49P9 demonstrated 89% neutralization (Individual IC50 and IC80 values against each pseudovirus are in the Excel file "N49 neutralization and sequences").
[1073] Beyond their ability to neutralize 100% of the viruses, the importance of these antibodies are that they can neutralize viruses that other broadly neutralizing antibodies are resistant to. The antibodies from N49 were able to neutralize all the viruses that the other mAbs were resistant to (importantly all the neutralization data was from the same lab). A summary of the neutralization breadth of the NVS49 antibodies compared to the others described are provided in Table 13 below.
TABLE-US-00014 TABLE 13 Comparison of neutralization breadth between those derived from NVS49 and those in the literature. Percentages are given as values of viruses neutralized (IC50 < 50 ug/ml)/viruses tested. All testing was carried independent of the inventors by Dr. Seaman at Harvard. mAb % neutralization N49P6 100% N49P7 100% N49P7.1 .sup. 99% N49P11 100% N49P9 .sup. 89% PGT121 65.8% PGT128 59.8% PGT145 .sup. 74% PGDM1400 79.8% PGT151 65.5% 10-1074 56.8% 10E8 94.9% PG9 84.7% PG16 82.2% 3BNC117 .sup. 90% NIH 45-46 86.4% 8ANC195 65.2% VRC07 91.5% VRC01 87.6%
Recently, there has been a report of a new broadly neutralizing antibody, N6, which has more breadth than other HIV broadly neutralizing antibodies (Huang et al,. Immunity 45, 1108-1121 (2016)). Dr. Seaman has only tested this antibody against a Clade C panel, and so we can not compare N6 with our antibodies directly. However, N49 P6, N49P7, and N49P11 has the ability to neutralize viruses that N6 was unable to (such as T278-50). These results show that the antibodies engineered from NVS49 are truly unique and are the broadest mAbs against HIV-1 that have been described to date. Currently, there are a variety of human and animal clinical trials that are addressing the utility of anti-HIV monoclonal antibodies for the prevention, treatment, or cure of HIV-1. These take the form of either using broadly neutralizing antibodies or their derivatives (engineered to have longer half-life, activity coupled with drugs, etc.). We anticipate that the extreme breadth and potency of the antibodies described above will make them highly useful in HIV research and for the prevention, treatment, or cure of HIV-1. In addition these mAbs can be used to select (purify) native trimers.
Example 3. Antibody Variants
[1074] This example contains additional details about modifications made to antibodies and contains a number of new antibodies. All antibody numbering is now based on IMGT (see FIGS. 14, 15, and 16), so a few of the individual positions are different than what was listed before (see also FIGS. 17 and 18 for aligned N49P6 and N49P7 mutants). Variants are described in Table 4, above.
Modifications of antibodies: [1075] 1) Modifications to improve stability. [1076] Heavy chain modifications: Substitution at position 1.4 of CH1 (1' amino acid of the constant region) to A from P or S. Substitution at position 120 of CH1 to R. Substitution at position 12 of CH3 to E, at position 14 of CH3 to M. [1077] Light chain modifications: modifications to change the unpaired cysteine to another amino acid that has potentially better pharmacokinetic properties: substitutions at IMGT position 42 (Cysteine) to A, S, Y, or F. Other light chain modifications include substitution at position 1.5 of the light chain constant region (1' amino acid of the constant region) from R to S or G for those that use LC2, or from S to R or G for those that use LC7. [1078] 2) Modifications to improve antibody function (binding, neutralization, complement fixation, ADCC, ADCP). [1079] Heavy chain modifications: Dual substitution at positions 59 and 62 to T and Y, respectively. Substitution of Y, F, W, or H at position 62. Substitution at position 1.4 of CH1 (1' amino acid of the constant region) to G from P or S. Substitution at position 120 of CH1 to R. Substitution at position 12 of CH3 to E, at position 14 of CH3 to M. [1080] Light chain modifications: Substitution at position 1.5 of the light chain constant region (1' amino acid of the constant region) from R to S or G for those that use LC2, or from S to R or G for those that use LC7. [1081] 3) Modifications to improve antibody half-life. [1082] Heavy chain modifications with combinations of any of the following substitutions: Substitution at position 120 of CH1 to R. Substitution at position 12 of CH3 to E, at position 14 of CH3 to M. Substitution at position 101 of CH3 to I, 107 of CH3 to L, position 113 of C.sub.H3 to S, position 115 to R. [1083] 4) Constant region swapping: Swapping of light chain constant region: light chain constant region swapped to another lambda constant region (LC1-7) for improved stability and function. Another method of swapping will be used to make monoclonals that are "rhesus-ized," that is, retain the variable regions in the heavy and light chains, but use constant regions from rhesus lambda chain (such as LC3), as well as rhesus IgG1. These mAbs can also have the half-life extending mutations noted above in the corresponding amino acids of the rhesus IgG1 constant region. [1084] 5) Swapping of heavy and light chain variable regions: These include taking a native or modified antibody listed in this application and mixing and matching the heavy and light chains to improve antibody stability or function. For example, one such antibody is made from the heavy chain of the parent antibody N49P7.1 (a variant of sequence 2) with the light chain from the parent antibody N49P9 (a variant of sequence 18). We have already made a number of these. As they can be readily expressed suggests the similarity in the gene family usage, and targeted epitopes makes these mAb swaps feasible to make and test. These swaps can also be made with other antibodies that target the same epitope (such as N60P1, N60P1.1, N60P2.1, N60P23, N60P31.1, N60P44, N60P45, VRC01, N6, etc.). [1085] 6) CDR replacement mutants: These include taking the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and/or CDRL3 from one of the natural or modified antibodies listed in this application and inserting into the corresponding CDR another natural or modified antibody from this application. For example, we have made an antibody with the CDR3 of sequence 2, inserted and replacing the CDR3 of N49P9 (a variant of sequence 18). [1086] 7) Antibody-drug conjugates: The natural and modified antibodies listed here can be conjugated (covalently or non-covalently) to markers (florescent dye or radionuclides), toxins, or therapeutic agents. We have constructed a conjugate between N49P7 and auristatin (a microtubule toxin). [1087] 8) Pharmaceutical compositions: These can include antibodies (pre-made in a variety of vehicles) delivered via injection, or genes encoding antibodies delivered via a viral or other vector. [1088] 9) Antibody-bead conjugates. The modified antibodies listed here can be conjugated to agarose or other beads by a variety of chemical reactions (such as but not limited to Cyanogen Bromide-Activated and NHS esters) for the purpose of creating affinity purification columns that can bind (and purify) gp120 and its mutants, gp160 and its mutants, HIV trimers, or HIV-1 virus. [1089] 10) Antibodies, their modifications, and fragments, can be used for the purpose of HIV prevention, treatment, or cure, and can be done individually or in combination with any number of anti-HIV treatments.
Example 4. Crystallographic Analysis of the Broadly Neutralizing Antibodies
[1090] To define the molecular basis for the broad potencies of N60 and N49 P mAbs series we solved the crystal structures of N60P23 and N49P7 Fabs in complex with HIV-1 93TH0S7 gp120 (Table 14).
TABLE-US-00015 TABLE 14 Data collection and refinement statistics. N49-P7 N60-P23 Fab-gp120.sub.93TH057 Fab-gp120.sub.93TH057 core.sub.e core.sub.e Data collection Wavelength, {acute over (.ANG.)} 0.979 0.979 Space group P2.sub.12.sub.12.sub.1 C2 Cell parameters a, b, c, .ANG. 61.4, 63.9, 255.3 127.6, 68.6, 119.4 .alpha., .beta., .gamma., .degree. 90, 90, 90 90, 111.4, 90 Complexes/a.u. 1 1 Resolution, (.ANG.) 50-2.69 (2.74-2.69) 50-2.38 (2.43-2.38) # of reflections Total 73,281 139,212 Unique 23,639 38,617 R.sub.merge.sup.b, % 13.1 (74.9) 16.0 (90.8) R.sub.pim.sup.c, % 7.7 (42.9) 9.7 (62.5) CC.sub.1/2.sup.d 0.99 (0.67) 0.99 (0.47) Wilson B.sub.factor (1/.ANG..sup.2).sup.e 53 30 I/.sigma. 9.3 (1.1) 9.7 (1.2) Completeness, % 80.6 (72.2) 100 (99.0) Redundancy 3.1 (3.2) 3.6 (2.9) Refinement Statistics Resolution, .ANG. 50.0-2.7 50.0-2.4 R.sup.f, % 22.5 21.4 R.sub.free.sup.g, % 28.5 25.8 # of atoms Protein 5,822 5,905 Water 27 222 Ligand/Ion 170 133 Overall B value (.ANG.).sup.2 Protein 50 44 Water 33 36 Ligand/Ion 64 59 Root mean square deviation Bond lengths, .ANG. 0.014 0.009 Bond angles, .degree. 1.2 1.4 Ramachandran.sup.h favored, % 89.0 93.5 allowed, % 9.7 5.8 outliers, % 1.3 0.7 PDB ID 6BCK 5WB9 Values in parentheses are for highest-resolution shell .sup.bR.sub.merge = .SIGMA. | I - <I> |/.SIGMA.I, where I is the observed intensity and <I> is the average intensity obtained from multiple observations of symmetry-related reflections after rejections .sup.cR.sub.pim = as defined in (Weiss, 2001) .sup.dCC.sub.1/2 = as defined by Karplus and Diederichs (Karplus and Diederichs, 2012) .sup.eWilson B.sub.factor as calculated in (Popov and Bourenkov, 2003) .sup.fR = .SIGMA. || F.sub.o | - | F.sub.c ||/.SIGMA. | F.sub.o |, where F.sub.o and F.sub.c are the observed and calculated structure factors, respectively .sup.gR.sub.free = as defined by Brunger (Brunger, 1997) .sup.hCalculated with MolProbity.
[1091] N60P23, a clone of N60 P1.1 that has a 1 amino acid (aa) difference in the light chain, exhibited an epitope footprint with intermolecular contacts similar to those of VRCO1 and other previously described CD4bs antibodies (FIG. 19 and Table 15). In contrast, N49P7 bound to gp120 in a unique manner (FIG. 20).
TABLE-US-00016 TABLE 15 Details of the N49_P7-gp120.sub.93TH057 core.sub.e, N60_P23-gp120.sub.93TH057 core.sub.e, N6-gp120.sub.93TH057 core.sub.e, and VRCO1-gp120.sub.93TH057 core.sub.e interfaces. Results as calculated by the EBI PISA server (http://www.ebi.ac.uk/msd-srv/prot_int/cgi-bin/piserver). N49_P7 N60_P23 N6 Fab- VRCO1- Fab- Fab- gp120.sub.93TH057 gp120.sub.93TH057 gp120.sub.93TH057 gp120.sub.93TH057 core.sub.e core.sub.e core.sub.e core.sub.e (5te6) (3ngb) Buried gp120total 981 939 1132 1143 Surface Outer domain (OD) 774 872 1000 1036 Area, .ANG..sup.2 Inner domain (ID) 207 67 132 107 Ratio of OD/ID 3.74 13.01 7.58 9.68 Loop D of OD 339 346 332 322 CD4 binding loop of 186 202 206 188 OD Loop V5 of OD 164 209 339 301 Heavy chain 821 726 928 897 total FWR 54 158 146 136 CDR H1 4 20 38 3 CDR H2 489 435 621 598 CDRH3 274 113 123 160 Light chain 138 243 251 306 total FWR 0 0 36 124 CDR L1 9 67 43 50 CDR L2 0 0 0 0 CDR L3 129 176 172 132 Heavy and light 959 969 1179 1203 chain total
[1092] The deletion in the CDRL1 (not found in N6) combined with the rotation/tilting of the light chain `opens` the variable light (V1) side of the N49P7 antigen binding site to accommodate different lengths of the highly variable loops D, E and VS (FIG. 21). Changes in the length of gp120 loop VS and the length (and glycosylation status) of loop E that cause steric clashes with an antibody light chain were described previously as mechanisms of HIV-I resistance to VRC01-class antibodies (Lynch et al., 2015). These signatures, along with a long CDRH3 (19 aa) and unique sequence signatures within CDR.B2 (not seen in VRCO1 and N6, FIG. 21), allow N49P7 to bypass the Phe43 cavity affording it an unusual capacity to contact the gp120 inner domain at residues 97, 102 and 124 of Layer 2, and 472-480 of Layer 3 (FIG. 18). Overall N49P7 contributes 207 .ANG..sup.2 of its buried surface area (BSA) to the gp120 inner domain which is much higher than N6 and the VRC0I Abs class (Table 15). The gp120 inner domain harbors some of the most conserved amino acid sequences in the HIV envelope, located within structural Layers 1, 2 and 3 and a consolidation of 8 P-strands (Finzi et al., 2010). Other CD4bs antibodies such as N6 and VRC0I also display contacts with the inner domain; however, these are less prominent. Although NIH 45-46 is not as broad and potent and N49P7, it has previously shown to have more contacts with the inner domain (mostly Layer 2) than VRC0I (Diskin et al., 2011), and it was postulated that this may be the reason for the breadth. N49P7 is unique in the number of inner domain contacts (especially Layer 3). N49P7 exhibits a lower outer domain to inner domain buried surface area ratio (3.74) compared to N6 (7.58) and VRC01 (9.68), which shows it has significantly more contacts with the inner domain (Table 15). Thus, the N49P7 paratope recognizes a mixed inner domain/CD4-binding site (termed here the iCD4bs) containing some of the most conserved sequences in gp120. These features help explain the extreme neutralization breadth of the antibody.
Methods
Crystallization
[1093] Initial crystal screens were done in robotic vapor-diffusion sitting drop trials using a Gryphon Protein Crystallization Robot (Art Robbins Instruments) with commercially available sparse matrix crystallization screens from Molecular Dimensions (Proplex and MacroSol), Emerald Biosystems (Precipitant Wizard Screen) and Emerald BioSystems (Synergy Screen). The screens were monitored periodically for protein crystals. Conditions that produced micro crystals were then reproduced and optimized using the hanging-drop, vapor diffusion method with drops of 0.5 .mu.l protein and 0.5 .mu.l precipitant solution. For the N60P23 complex conditions producing diffraction quality crystals came from 0.1 M Magnesium acetate hexahydrate, 0.065.M NaCl and 0.1 M MOPS pH 7.5 after incubation at 22.degree. C. N49P7 complex crystals came from 10% PEG 5000 MME, 12% isopropanol, and 0.1 MMES pH 6.5. Crystals were frozen in liquid nitrogen after a brief soak in mother liquor supplemented with 20% MPD prior to being used for data collection.
Data Collection and Structure Solution and Refinement.
[1094] Diffraction data for N60P23 Fab- and N49P7 Fab-gp 12093TH057 core.sub.e complexes were collected at the Stanford Synchrotron Radiation Light Source (SSRL) at the beam line BL14 1 (N60P23) and BL12-2 (N49P7) equipped with Marmosaic 325 or Pilatus area detectors respectively. N60P23 crystals belong to a space group C2 with the unit-cell parameters a=127.6, b=68. 6, c=119.4 A and =111.4.degree. with one N60P23 Fab-gp120.sub.93TH057 core.sub.e complex present in the asymmetric unit (ASU). N49P7 crystals belong to space group P2.sub.12.sub.12.sub.1 with the unit-cell parameters a=61.4, b=63.9, and c=255.3 .ANG. with one N49P7 Fab-gp12093THo57 core.sub.e complex present in the ASU. Data was processed and reduced with HKL2000, as previously described (Guan et al., 2013). The data for the N49P7 complex was highly anisotropic and was further processed using the STARANISO server (Global Phasing Ltd. [http://staraniso.globalphasing.org/cgi-bin/staraniso.cgi]). The N60P23 structure was solved by molecular replacement with Phaser from the CCP4 suite based on the coordinates of gp120 (PDB: 3TGT) and the VRCO1 Fab (PDB: 4RFE) for the N6P23 Fab. N49P7 was solved using coordinates of gp120 (PDB: 3TGT) and N5-I5 Fab (PDB: 3TNN) for the N49P7 Fab. Refinement was done with Refmac and/or Phenix, coupled with manual refitting and rebuilding using COOT, as previously described (Guan et al., 2013). The N60P23 complex complex was refined to an R-factor of 0.214 and an R-free of 0.258 and the N49P7 complex was refined to an R-factor of 0.225 and R-free of 0.285. Data collection and refinement statistics are shown in (Table 14).
Structure Validation and Analysis.
[1095] The quality of the final refined model was monitored using the program MolProbity, as previously described (Guan et al., 2013). Structural alignments were performed using the Dali server and the program lsqkab from CCP4 suite. The PISA webserver was used to determine contact smfaces and interface residues. All illustrations were prepared with the PyMol Molecular Graphic suite (http://pymol.org) (DeLano Scientific, San Carlos, Calif., USA).
Data Availablity.
[1096] The data reported in this paper are archived at the following databases: GenBank and Protein Data Bank (PDB).
Example 5. Neutralization Assays of HIV Antibody Variants
[1097] Shown in this example are neutralization assay data of various anti-HIV antibodies of the invention across an Env pseudovirus panel (See Table).
TABLE-US-00017 TABLE 16 Neutralization assay data of mAbs against Env pseudovirus panel. mAbs tested at primary concentration of 50 ug/ml, 25 ug/ml, or 17 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 16 N49P7A N49P7S N49P7-54TY N49P7YTE Virus ID Clade* IC50 IC80 MPI IC50 IC80 MPI IC50 IC80 MPI IC50 IC80 MP PVO.4 B 0.079 0.205 100 0.035 0.132 100 0.079 0.314 100 0.088 0.394 100 AC10.0.29 B >17 >17 45 36.004 >50 55 >25 >25 40 >50 >50 28 62357_14_D3_4589 B (T/F) 0.031 0.105 100 0.020 0.074 100 0.055 0.192 100 0.039 0.173 100 Du172.17 C 0.088 0.376 100 0.055 0.282 100 0.124 0.529 100 0.283 0.953 100 Du422.1 C 2.417 13.266 85 6.131 48.813 82 0.402 2.022 96 13.224 >50 76 CAP210.2.00.E8 C 0.729 5.618 94 0.640 3.233 100 0.691 3.329 97 3.109 13.903 99 Ce1086_B2 C (T/F) 0.072 0.349 100 0.042 0.166 100 0.067 0.339 100 0.113 0.530 99 Ce1172_H1 C (T/F) >17 >17 4 >50 >50 1 >25 >25 23 >50 >50 12 CNE20 BC 0.045 0.122 100 0.038 0.141 100 0.046 0.160 100 0.110 0.317 99 Q259.d2.17 A 0.273 2.511 93 0.814 6.528 91 0.132 1.129 95 5.340 44.139 82 191955_A11 A (T/F) 3.118 >17 75 10.158 >50 72 3.045 >25 72 39.028 >50 56 T250-4 CRF02_AG 2.460 >17 74 12.223 >50 65 1.020 19.555 83 >50 >50 43 620345.c01 CRF01_AE >17 >17 0 >50 >50 0 >25 >25 0 >50 >50 0 R1166.c01 CRF01_AE 0.076 0.349 100 0.050 0.172 100 0.131 0.470 100 0.232 0.782 100 BJOX028000.10.3 CRF01_AE 0.029 0.177 100 0.009 0.034 100 0.009 0.044 100 0.018 0.155 100 (T/F) X1254_c3 G 3.732 >17 71 16.977 >50 71 0.145 0.599 100 26.154 >50 59 X2088_c9 G 0.084 0.381 100 0.083 0.275 100 0.172 0.572 100 0.329 1.060 100 3016.v5.c45 D 0.275 1.801 92 0.635 6.393 92 0.206 1.908 92 1.151 9.654 89 6952.v1.c20 CD >17 >17 16 >50 >50 18 >25 >25 20 >50 >50 13 6545.v4.c1 6.807 >17 62 12.387 >50 63 3.014 >25 74 >50 >50 45 3103.v3.c10 ACD 0.061 0.239 100 0.057 0.168 100 0.151 0.550 100 0.322 0.583 100 MuLV Neg. >17 >17 0 >50 >50 0 >25 >25 0 >50 >50 4 Control indicates data missing or illegible when filed
Example 6. Neutralization Assays of Hiv Antibody Variants
[1098] Shown in this example are neutralization assay data of various HIV antibodies of the invention and known HIV antibodies across an extended multiclade virus panel (See Tables 17-25).
TABLE-US-00018 TABLE 17 Neutralization assay data of NS49 plasma and NS49 P6 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 17 NVS49 Plasma NVS49 P6 Virus ID Clade* ID50 ID80 IC50 IC80 MPI 6535.3 B 460 93 0.417 1.341 100 QH0692.42 B 207 71 0.652 2.276 100 SC422661.8 B 427 125 0.129 0.360 100 PVO.4 B 543 143 0.128 0.357 100 TRO.11 B 568 208 0.061 0.190 100 AC10.0.29 B 114 <40 32.310 >50 67 RHPA4259.7 B 830 346 0.117 0.443 100 THRO4156.18 B 144 46 12.487 38.403 89 REJO4541.67 B 925 244 0.066 0.215 100 TRJO4551.58 B 356 134 0.255 0.874 100 WITO4160.33 B 641 170 0.096 0.277 100 CAAN5342.A2 B 342 103 0.657 1.829 100 WEAU_d15_410_787 B (T/F) 709 271 0.070 0.188 100 1006_11_C3_1601 B (T/F) 879 293 0.068 0.209 100 1054_07_TC4_1499 B (T/F) 163 <40 0.624 2.848 100 1056_10_TA11_1826 B (T/F) 314 51 0.209 0.606 100 1012_11_TC21_3257 B (T/F) 354 83 0.137 0.502 100 6240_08_TA5_4622 B (T/F) 127 <40 0.567 1.757 100 6244_13_B5_4576 B (T/F) 318 78 0.095 0.328 100 62357_14_D3_4589 B (T/F) 363 112 0.098 0.296 100 SC05_8C11_2344 B (T/F) 355 90 0.266 0.911 100 Du156.12 C 1,519 355 0.082 0.280 100 Du172.17 C 687 100 20.759 >50 74 Du422.1 C 160 <40 44.007 >50 57 ZM197M.PB7 C 251 63 3.001 12.401 98 ZM214M.PL15 C 295 41 0.510 1.802 100 ZM233M.PB6 C 250 56 1.233 4.005 99 ZM249M.PL1 C 345 116 0.112 0.376 100 ZM53M.PB12 C 121 42 0.538 1.706 100 ZM109F.PB4 C 591 170 0.084 0.290 99 ZM135M.PL10a C 438 68 0.494 1.775 99 CAP45.2.00.G3 C 398 67 21.663 >50 75 CAP210.2.00.E8 C 104 <40 22.582 >50 73 HIV-001428-2.42 C 1,810 764 0.016 0.050 100 HIV-0013095-2.11 C 330 96 0.971 8.044 96 HIV-16055-2.3 C 774 200 0.117 0.586 100 HIV-16845-2.22 C 254 52 0.692 3.754 100 Ce1086_B2 C (T/F) 680 148 0.581 4.086 98 Ce0393_C3 C (T/F) 496 79 0.608 3.249 100 Ce1176_A3 C (T/F) 327 52 0.594 2.114 100 Ce2010_F5 C (T/F) 386 83 0.124 0.749 100 Ce0682_E4 C (T/F) 1,378 246 0.095 0.315 100 Ce1172_H1 C (T/F) 112 <40 14.353 47.253 82 Ce2060_G9 C (T/F) 146 <40 0.888 4.812 100 Ce703010054_2A2 C (T/F) 837 155 0.245 0.841 100 BF1266.431a C (T/F) 1,228 255 0.083 0.283 100 246F C1G C (T/F) 727 240 0.513 2.510 99 249M B10 C (T/F) 296 95 0.122 0.549 100 ZM247v1(Rev-) C (T/F) 274 60 0.063 0.170 100 7030102001E5(Rev-) C (T/F) 515 161 0.354 1.171 100 1394C9G1(Rev-) C (T/F) 230 59 17.498 >50 80 Ce704809221_1B3 C (T/F) 534 139 1.667 12.774 97 CNE19 BC 1,095 289 0.044 0.156 100 CNE20 BC 233 40 2.739 15.642 95 CNE21 BC 174 43 0.832 6.517 99 CNE17 BC 172 <40 0.340 2.103 100 CNE30 BC 159 57 0.457 1.536 100 CNE52 BC 1,078 467 0.028 0.095 100 CNE53 BC 457 151 0.249 0.871 99 CNE58 BC 1,025 352 0.036 0.100 100 MS208.A1 A 578 102 0.110 0.436 100 Q23.17 A 496 184 0.052 0.185 100 Q461.e2 A 199 68 0.173 0.622 100 Q769.d22 A 498 169 0.072 0.318 100 Q259.d2.17 A 316 88 24.376 >50 75 Q842.d12 A 1,430 546 0.034 0.091 100 3415.v1.c1 A 491 170 0.188 0.648 100 3365.v2.c2 A 521 193 0.129 0.347 100 0260.v5.c36 A 173 <40 0.485 1.551 100 191955_A11 A (T/F) 151 44 0.124 0.344 100 191084 B7-19 A (T/F) 552 184 0.062 0.257 100 9004SS_A3_4 A (T/F) 783 202 0.181 0.817 100 T257-31 CRF02_AG 330 105 0.147 0.535 100 928-28 CRF02_AG 178 50 1.242 5.604 99 263-8 CRF02_AG 1,091 321 0.116 0.438 100 T250-4 CRF02_AG 631 167 21.927 >50 68 T251-18 CRF02_AG 263 83 0.344 0.929 100 T278-50 CRF02_AG 45 <40 25.690 >50 67 T255-34 CRF02_AG 1,005 295 0.076 0.392 100 211-9 CRF02_AG 403 85 8.494 27.679 93 235-47 CRF02_AG 984 379 12.976 46.426 82 620345.c01 CRF01_AE <40 <40 22.899 >50 70 CNE8 CRF01_AE 688 244 0.072 0.245 100 C1080.c03 CRF01_AE 852 106 0.181 0.661 100 R2184.c04 CRF01_AE 2,504 535 0.028 0.077 100 R1166.c01 CRF01_AE 677 186 0.102 0.344 100 R3265.c06 CRF01_AE NT NT 0.101 0.443 100 C2101.c01 CRF01_AE 1,943 252 0.140 0.597 100 C3347.c11 CRF01_AE 1,772 405 0.080 0.266 100 C4118.c09 CRF01_AE 877 150 0.092 0.243 100 CNE5 CRF01_AE 401 93 BJOX009000.02.4 CRF01_AE 139 43 0.320 1.463 100 BJOX015000.11.5 CRF01_AE 783 133 0.070 0.312 100 (T/F) BJOX010000.06.2 CRF01_AE 230 54 0.681 3.529 100 (T/F) BJOX025000.01.1 CRF01_AE 422 59 14.946 >50 78 (T/F) BJOX028000.10.3 CRF01_AE 3,724 175 6.678 >50 77 (T/F) X1193_c1 G 993 185 0.074 0.224 100 P0402_c2_11 G 512 113 3.536 18.954 99 X1254_c3 G 172 63 15.739 34.230 91 X2088_c9 G 133 46 27.036 >50 65 X2131_C1_B5 G 600 97 0.174 0.720 100 P1981_C5_3 G 293 91 0.310 0.708 100 X1632_S2_B10 G 97 <40 23.063 >50 69 3016.v5.c45 D 234 52 21.668 >50 72 A07412M1.vrc12 D 233 54 0.229 0.786 100 231965.c01 D 136 46 9.245 39.386 87 231966.c02 D 258 94 0.339 1.621 100 3817.v2.c59 CD 54 <40 16.922 >50 78 6480.v4.c25 CD 1,140 246 0.089 0.427 100 6952.v1.c20 CD 328 91 16.912 >50 78 6811.v7.c18 CD 164 48 0.238 1.008 100 89-F1_2_25 CD 78 <40 33.639 >50 61 3301.v1.c24 AC 1,006 268 0.054 0.149 100 6041.v3.c23 AC 1,317 281 5.525 43.153 88 6540.v4.c1 AC 766 245 13.973 47.540 81 6545.v4.c1 AC 165 49 24.487 >50 76 0815.v3.c3 ACD 2,249 346 0.194 1.251 100 3103.v3.c10 ACD 324 87 0.582 1.567 100 MuLV Neg. Control <40 <40 >50 >50 0 *(T/F): Transmitted/Founder Virus
TABLE-US-00019 TABLE 18 Neutralization assay data of NS49 P7 and NS49 P7.1 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 18 NVS49 P7 NVS49 P7.1 Virus ID Clade* IC50 IC80 MPI IC50 IC80 MPI 6535.3 B 0.186 0.598 100 0.224 0.715 100 QH0692.42 B 0.663 2.411 100 0.764 2.710 100 SC422661.8 B 0.114 0.320 100 0.125 0.417 100 PVO.4 B 0.107 0.365 100 0.124 0.421 100 TRO.11 B 0.044 0.160 100 0.055 0.191 100 AC10.0.29 B 5.034 36.661 89 5.308 46.750 85 RHPA4259.7 B 0.083 0.246 100 0.108 0.292 100 THRO4156.18 B 2.341 12.410 99 3.068 12.603 98 REJO4541.67 B 0.015 0.049 100 0.022 0.088 100 TRJO4551.58 B 0.104 0.351 100 0.156 0.529 100 WITO4160.33 B 0.084 0.248 100 0.121 0.326 100 CAAN5342.A2 B 0.409 1.211 100 0.550 1.589 100 WEAU_d15_410_787 B (T/F) 0.045 0.117 100 0.058 0.142 100 1006_11_C3_1601 B (T/F) 0.044 0.158 100 0.061 0.193 100 1054_07_TC4_1499 B (T/F) 0.494 2.308 100 0.627 2.980 100 1056_10_TA11_1826 B (T/F) 0.141 0.468 100 0.171 0.603 100 1012_11_TC21_3257 B (T/F) 0.046 0.125 100 0.063 0.176 100 6240_08_TA5_4622 B (T/F) 0.457 1.728 100 0.509 1.951 100 6244_13_B5_4576 B (T/F) 0.111 0.386 100 0.184 0.493 100 62357_14_D3_4589 B (T/F) 0.025 0.102 100 0.028 0.150 100 SC05_8C11_2344 B (T/F) 0.164 0.403 100 0.189 0.688 100 Du156.12 C 0.032 0.086 100 0.045 0.131 100 Du172.17 C 0.102 0.668 100 0.185 0.748 100 Du422.1 C 4.496 19.548 93 3.763 15.661 93 ZM197M.PB7 C 0.109 0.347 100 0.144 0.539 100 ZM214M.PL15 C 0.213 0.953 100 0.257 0.853 100 ZM233M.PB6 C 0.226 0.774 100 0.355 1.246 100 ZM249M.PL1 C 0.039 0.134 100 0.071 0.239 100 ZM53M.PB12 C 0.251 0.800 100 0.287 1.003 100 ZM109F.PB4 C 0.034 0.153 100 0.055 0.199 100 ZM135M.PL10a C 0.200 0.960 100 0.314 1.107 99 CAP45.2.00.G3 C 0.060 0.290 100 0.086 0.567 100 CAP210.2.00.E8 C 1.817 6.485 100 2.033 7.298 99 HIV-001428-2.42 C 0.006 0.020 100 0.009 0.029 100 HIV-0013095-2.11 C 0.073 0.253 100 0.105 0.374 100 HIV-16055-2.3 C 0.057 0.280 100 0.088 0.318 100 HIV-16845-2.22 C 0.950 3.935 100 1.075 5.400 100 Ce1086_B2 C (T/F) 0.089 0.568 100 0.121 0.742 99 Ce0393_C3 C (T/F) 0.195 0.802 100 0.230 1.017 100 Ce1176_A3 C (T/F) 0.204 0.589 100 0.279 1.012 100 Ce2010_F5 C (T/F) 0.130 0.547 100 0.055 0.208 100 Ce0682_E4 C (T/F) 0.041 0.131 100 0.026 0.099 100 Ce1172_H1 C (T/F) 8.499 37.520 92 17.063 >50 78 Ce2060_G9 C (T/F) 0.661 2.547 100 0.473 2.410 100 Ce703010054_2A2 C (T/F) 0.132 0.466 100 0.085 0.364 100 BF1266.431a C (T/F) 0.057 0.199 100 0.033 0.110 100 246F C1G C (T/F) 0.094 0.255 100 0.049 0.165 100 249M B10 C (T/F) 0.081 0.377 100 0.053 0.225 100 ZM247v1(Rev-) C (T/F) 0.062 0.214 100 0.038 0.176 100 7030102001E5(Rev-) C (T/F) 0.184 0.832 100 0.110 0.498 100 1394C9G1(Rev-) C (T/F) 3.841 21.069 94 4.369 30.932 88 Ce704809221_1B3 C (T/F) 0.141 0.932 100 0.211 1.355 100 CNE19 BC 0.009 0.041 100 0.027 0.128 100 CNE20 BC 0.028 0.100 100 0.079 0.224 100 CNE21 BC 0.756 6.020 97 1.584 12.250 96 CNE17 BC 0.126 0.460 100 0.254 0.860 100 CNE30 BC 0.180 0.609 100 0.307 1.013 100 CNE52 BC 0.008 0.023 100 0.021 0.058 100 CNE53 BC 0.035 0.124 100 0.066 0.228 100 CNE58 BC 0.029 0.067 100 0.049 0.107 100 MS208A1 A 0.120 0.479 100 0.120 0.431 100 Q23.17 A 0.052 0.210 100 0.077 0.208 100 Q461.e2 A 0.261 0.922 100 0.329 1.143 100 Q769.d22 A 0.025 0.111 100 0.044 0.163 100 Q259.d2.17 A 0.332 2.474 98 0.505 4.233 97 Q842.d12 A 0.021 0.060 100 0.028 0.080 100 3415.v1.c1 A 0.160 0.550 100 0.191 0.647 100 3365.v2.c2 A 0.070 0.186 100 0.088 0.289 100 0260.v5.c36 A 0.354 0.884 100 0.497 1.515 100 191955_A11 A (T/F) 1.811 8.870 99 1.994 10.089 98 191084 B7-19 A (T/F) 0.047 0.142 100 0.069 0.200 100 9004SS_A3_4 A (T/F) 0.125 0.390 100 0.221 0.841 100 T257-31 CRF02_AG 0.158 0.619 100 0.209 1.039 100 928-28 CRF02_AG 0.167 0.579 100 0.230 0.775 100 263-8 CRF02_AG 0.056 0.201 100 0.104 0.392 100 T250-4 CRF02_AG 2.070 21.419 91 2.271 24.731 89 T251-18 CRF02_AG 0.178 0.620 100 0.229 0.794 100 T278-50 CRF02_AG 22.274 >50 75 42.378 >50 56 T255-34 CRF02_AG 0.053 0.262 100 0.094 0.447 100 211-9 CRF02_AG 0.220 0.767 100 0.340 1.165 100 235-47 CRF02_AG 0.045 0.300 99 0.078 0.419 97 620345.c01 CRF01_AE 23.174 47.935 82 >50 >50 47 CNE8 CRF01_AE 0.037 0.125 100 0.048 0.167 100 C1080.c03 CRF01_AE 0.114 0.765 100 0.144 1.046 100 R2184.c04 CRF01_AE 0.011 0.031 100 0.018 0.040 100 R1166.c01 CRF01_AE 0.092 0.313 100 0.136 0.461 100 R3265.c06 CRF01_AE 0.048 0.282 100 0.059 0.328 100 C2101.c01 CRF01_AE 0.060 0.340 100 0.076 0.425 100 C3347.c11 CRF01_AE 0.024 0.067 100 0.048 0.161 100 C4118.c09 CRF01_AE 0.043 0.147 100 0.048 0.216 100 CNE5 CRF01_AE BJOX009000.02.4 CRF01_AE 0.113 0.506 100 0.219 0.756 100 BJOX015000.11.5 CRF01_AE 0.036 0.157 100 0.111 0.487 100 (T/F) BJOX010000.06.2 CRF01_AE 0.085 0.592 100 0.132 0.623 100 (T/F) BJOX025000.01.1 CRF01_AE 0.038 0.236 100 0.057 0.378 100 (T/F) BJOX028000.10.3 CRF01_AE 0.020 0.123 100 0.031 0.290 100 (T/F) X1193_c1 G 0.016 0.060 100 0.035 0.144 100 P0402_c2_11 G 0.014 0.061 100 0.029 0.115 100 X1254_c3 G 2.969 13.594 100 3.963 18.205 96 X2088_c9 G 0.116 0.338 100 0.199 0.726 100 X2131_C1_B5 G 0.081 0.273 100 0.129 0.431 100 P1981_C5_3 G 0.113 0.274 100 0.216 0.494 100 X1632_S2_B10 G 12.046 40.378 85 17.861 >50 65 3016.v5.c45 D 0.221 1.037 100 0.559 2.542 100 A07412M1.vrc12 D 0.108 0.645 100 0.232 1.070 100 231965.c01 D 0.054 0.193 100 0.106 0.364 100 231966.c02 D 0.102 0.430 100 0.203 0.705 100 3817.v2.c59 CD 0.716 3.726 99 1.484 8.359 97 6480.v4.c25 CD 0.031 0.113 100 0.056 0.192 100 6952.v1.c20 CD 18.386 38.898 91 26.274 >50 71 6811.v7.c18 CD 0.229 0.992 100 0.321 1.460 100 89-F1_2_25 CD 21.631 >50 80 41.582 >50 58 3301.v1.c24 AC 0.025 0.069 100 0.041 0.111 100 6041.v3.c23 AC 0.047 0.222 100 0.078 0.366 100 6540.v4.c1 AC 11.747 33.167 92 14.695 >50 75 6545.v4.c1 AC 2.001 14.657 96 1.147 15.962 92 0815.v3.c3 ACD 0.016 0.053 100 0.023 0.064 100 3103.v3.c10 ACD 0.158 0.509 100 0.229 0.772 100 MuLV Neg. Control >50 >50 9 >50 >50 4 *(T/F): Transmitted/Founder Virus
TABLE-US-00020 TABLE 19 Neutralization assay data ofNS49 P7.2, NS49 P11, and NS49P18.1 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 19 N49P7.2 NVS49 P11 N49P18.1 Virus ID Clade* IC50 IC80 MPI IC50 IC80 MPI IC50 IC80 MPI 6535.3 B 0.152 0.784 100 8.224 17.196 100 2.064 11.368 100 QH0692.42 B 1.263 4.424 100 0.550 2.645 100 >50 >50 43 SC422661.8 B 0.119 0.564 100 0.729 3.489 100 14.050 43.430 84 PVO.4 B 0.237 0.831 100 0.248 1.169 100 9.650 40.160 84 TRO.11 B 0.113 0.306 100 0.179 0.937 100 4.248 11.683 99 AC10.0.29 B 44.175 >50 53 5.523 19.972 100 20.021 >50 71 RHPA4259.7 B 0.100 0.289 100 0.202 0.956 100 4.340 21.426 98 THRO4156.18 B 6.293 33.287 94 1.838 10.368 100 >50 >50 19 REJO4541.67 B 0.018 0.080 100 10.785 27.072 100 4.079 30.128 89 TRJO4551.58 B 0.210 0.673 100 0.183 0.656 100 4.437 14.717 99 WITO4160.33 B 0.051 0.215 100 4.857 13.218 100 10.999 37.591 87 CAAN5342.A2 B 0.809 2.805 100 9.027 20.805 100 16.089 49.066 83 WEAU_d15_410_787 B (T/F) 0.052 0.150 100 1.703 6.019 100 4.266 15.463 98 1006_11_C3_1601 B (T/F) 0.085 0.237 100 0.125 0.487 100 2.804 9.658 99 1054_07_TC4_1499 B (T/F) 2.036 10.723 99 2.915 13.091 100 >50 >50 0 1056_10_TA11_1826 B (T/F) 0.114 0.771 100 4.774 14.202 100 30.532 >50 62 1012_11_TC21_3257 B (T/F) 0.060 0.285 100 10.593 29.254 100 16.105 >50 79 6240_08_TA5_4622 B (T/F) 0.812 2.869 100 0.720 2.998 100 35.211 >50 60 6244_13_B5_4576 B (T/F) 0.331 1.043 100 1.119 5.446 100 33.375 >50 61 62357_14_D3_4589 B (T/F) 0.030 0.121 100 0.599 3.561 100 10.408 47.969 82 SC05_8C11_2344 B (T/F) 0.326 1.135 100 5.419 20.488 100 17.283 >50 79 Du156.12 C 0.058 0.210 100 2.396 12.008 100 1.877 6.812 100 Du172.17 C 0.290 1.078 100 5.653 17.918 100 6.516 21.335 92 Du422.1 C >50 >50 22 8.815 21.224 100 21.519 >50 75 ZM197M.PB7 C 0.223 1.371 100 0.303 1.591 100 12.493 49.791 84 ZM214M.PL15 C 0.559 4.532 100 1.347 8.976 100 >50 >50 20 ZM233M.PB6 C 0.475 2.137 99 5.922 18.544 100 38.427 >50 57 ZM249M.PL1 C 0.073 0.256 100 3.637 17.421 100 4.159 19.807 98 ZM53M.PB12 C 0.477 1.728 100 1.051 6.531 100 1.302 4.506 99 ZM109F.PB4 C 0.065 0.321 99 5.203 19.443 100 26.597 >50 69 ZM135M.PL10a C 0.596 2.801 99 3.968 16.630 100 >50 >50 46 CAP45.2.00.G3 C 0.162 2.490 95 5.253 23.977 100 5.140 33.068 88 CAP210.2.00.E8 C 3.943 21.717 95 5.119 19.418 100 >50 >50 11 HIV-001428-2.42 C 0.010 0.036 100 0.221 1.643 100 1.131 8.312 100 HIV-0013095-2.11 C 0.183 0.931 97 5.849 17.771 100 >50 >50 48 HIV-16055-2.3 C 0.204 0.721 100 0.091 0.503 100 8.035 22.606 97 HIV-16845-2.22 C 3.910 18.221 98 1.723 8.152 100 >50 >50 23 Ce1086_B2 C (T/F) 0.166 1.005 100 2.187 11.745 100 8.979 48.001 92 Ce0393_C3 C (T/F) 0.137 0.368 100 1.507 9.715 100 10.321 24.836 92 Ce1176_A3 C (T/F) 0.332 1.079 100 0.946 6.047 100 32.478 >50 61 Ce2010_F5 C (T/F) 0.398 1.294 100 0.380 3.266 100 28.999 >50 68 Ce0682_E4 C (T/F) 0.067 0.233 100 0.091 0.387 100 3.946 21.075 96 Ce1172_H1 C (T/F) >50 >50 6 3.548 19.204 100 46.416 >50 52 Ce2060_G9 C (T/F) 0.821 2.809 100 3.333 13.172 100 >50 >50 37 Ce703010054_2A2 C (T/F) 0.124 0.576 100 1.423 7.511 100 4.150 11.552 98 BF1266.431a C (T/F) 0.029 0.159 100 0.998 7.128 100 3.482 20.778 93 246F C1G C (T/F) 0.049 0.192 100 8.481 20.920 100 3.395 20.214 97 249M B10 C (T/F) 0.095 0.349 100 3.544 12.947 100 5.706 23.139 94 ZM247v1(Rev-) C (T/F) 0.177 2.497 96 7.483 21.473 100 10.869 >50 79 7030102001E5(Rev-) C (T/F) 0.313 1.034 100 0.635 2.285 100 6.353 17.738 98 1394C9G1(Rev-) C (T/F) >50 >50 20 6.129 22.341 100 >50 >50 17 Ce704809221_1B3 C (T/F) 0.874 3.880 100 0.644 3.296 100 41.584 >50 59 CNE19 BC 0.033 0.129 100 0.437 5.503 100 2.343 9.638 99 CNE20 BC 0.119 0.574 99 5.194 22.606 100 1.329 6.364 100 CNE21 BC 48.266 >50 51 7.399 21.949 100 7.933 29.198 95 CNE17 BC 0.376 1.238 100 6.682 25.594 100 39.715 >50 56 CNE30 BC 0.362 1.197 100 0.889 4.452 100 13.131 32.152 93 CNE52 BC 0.030 0.088 100 0.060 0.307 100 2.458 6.649 100 CNE53 BC 0.100 0.277 100 0.430 2.880 100 4.083 21.770 97 CNE58 BC 0.045 0.129 100 0.038 0.135 100 1.917 5.258 100 MS208.A1 A 0.186 0.966 100 0.259 1.944 100 8.315 33.318 88 Q23.17 A 0.051 0.200 100 0.034 0.254 100 1.160 6.932 100 Q461.e2 A 0.472 2.237 100 0.363 2.150 100 24.690 >50 68 Q769.d22 A 0.029 0.140 100 0.048 0.308 100 4.916 18.335 97 Q259.d2.17 A 38.484 >50 54 8.419 21.433 100 >50 >50 37 Q842.d12 A 0.045 0.200 100 0.027 0.103 100 3.041 8.132 100 3415.v1.c1 A 0.381 1.580 100 1.862 10.264 100 >50 >50 48 3365.v2.c2 A 0.066 0.267 100 0.064 0.452 100 4.725 14.309 99 0260.v5.c36 A 0.644 1.800 100 0.438 1.469 100 16.179 47.836 81 191955_A11 A (T/F) 44.507 >50 56 2.741 15.296 100 >50 >50 33 191084 B7-19 A (T/F) 0.102 0.290 100 0.065 0.276 100 5.588 16.863 98 9004SS_A3_4 A (T/F) 0.266 1.618 100 0.296 3.242 100 3.021 13.950 99 T257-31 CRF02_AG 0.627 1.786 100 0.700 2.901 100 >50 >50 48 928-28 CRF02_AG 0.347 1.107 100 5.456 24.965 100 22.732 >50 71 263-8 CRF02_AG 0.095 0.641 100 0.064 0.259 100 3.964 41.037 90 T250-4 CRF02_AG >50 >50 15 6.278 25.585 100 4.863 15.084 94 T251-18 CRF02_AG 0.222 0.741 100 3.054 16.697 100 22.990 >50 70 T278-50 CRF02_AG >50 >50 33 7.628 25.130 100 >50 >50 36 T255-34 CRF02_AG 0.167 0.527 100 0.418 3.830 100 10.413 33.757 90 211-9 CRF02_AG 0.363 1.220 100 5.672 18.196 100 18.321 >50 73 235-47 CRF02_AG 0.127 1.034 90 3.876 21.250 100 23.013 >50 66 620345.c01 CRF01_AE >50 >50 0 11.662 27.932 100 >50 >50 0 CNE8 CRF01_AE 0.073 0.240 100 0.165 1.237 100 16.158 44.673 83 C1080.c03 CRF01_AE 0.279 1.757 100 0.162 1.452 100 >50 >50 50 R2184.c04 CRF01_AE 0.020 0.047 100 0.034 0.120 100 3.376 11.753 100 R1166.c01 CRF01_AE 0.267 0.852 100 0.060 0.336 100 19.190 >50 78 R3265.c06 CRF01_AE 0.153 0.985 100 0.161 0.971 100 27.947 >50 66 C2101.c01 CRF01_AE 0.162 0.727 100 0.084 0.505 100 15.305 41.944 85 C3347.c11 CRF01_AE 0.047 0.154 100 0.055 0.192 100 4.201 14.261 99 C4118.c09 CRF01_AE 0.079 0.292 100 0.094 0.346 100 8.627 23.239 93 CNE5 CRF01_AE 0.206 0.893 100 19.516 >50 72 BJOX009000.02.4 CRF01_AE >50 >50 43 2.841 10.644 100 0.418 1.428 100 BJOX015000.11.5 CRF01_AE 0.289 1.264 100 0.288 1.276 100 27.277 >50 67 (T/F) BJOX010000.06.2 CRF01_AE 0.267 1.111 100 6.810 21.831 100 >50 >50 39 (T/F) BJOX025000.01.1 CRF01_AE 0.102 0.831 100 0.361 2.580 100 7.667 39.071 85 (T/F) BJOX028000.10.3 CRF01_AE 0.322 1.491 100 5.330 46.666 100 34.001 >50 64 (T/F) X1193_c1 G 0.014 0.101 100 0.179 1.086 100 3.121 23.883 94 P0402_c2_11 G 0.106 0.400 100 0.091 0.698 100 5.470 20.782 98 X1254_c3 G 27.526 >50 53 5.987 20.682 100 24.797 >50 61 X2088_c9 G 0.377 2.030 99 6.816 21.955 100 9.084 34.895 88 X2131_C1_B5 G 0.161 0.671 100 5.364 18.369 100 30.361 >50 75 P1981_C5_3 G 0.258 0.716 100 5.293 16.523 100 19.315 45.400 83 X1632_S2_B10 G >50 >50 22 9.727 26.077 100 >50 >50 19 3016.v5.c45 D 2.765 23.879 90 8.749 21.450 100 >50 >50 49 A07412M1.vrc12 D 0.768 2.319 100 2.517 12.862 100 41.253 >50 54 231965.c01 D 0.316 1.063 100 9.778 24.225 100 >50 >50 31 231966.c02 D 0.233 1.110 100 4.308 15.953 100 29.943 >50 67 3817.v2.c59 CD 9.833 >50 68 4.724 15.901 100 >50 >50 34 6480.v4.c25 CD 0.098 0.371 100 0.227 1.032 100 2.968 17.293 99 6952.v1.c20 CD >50 >50 0 6.569 22.666 100 >50 >50 30 6811.v7.c18 CD 0.541 2.550 100 3.677 14.637 100 7.427 31.872 94 89-F1_2_25 CD >50 >50 18 4.237 16.633 100 >50 >50 9 3301.v1.c24 AC 0.050 0.137 100 1.978 15.720 100 3.247 11.319 99 6041.v3.c23 AC 0.309 5.114 94 4.422 17.592 100 3.050 12.169 96 6540.v4.c1 AC >50 >50 33 3.052 16.320 100 >50 >50 35 6545.v4.c1 AC >50 >50 4 6.471 22.906 100 >50 >50 0 0815.v3.c3 ACD 0.023 0.103 100 0.106 0.372 100 0.741 8.454 99 3103.v3.c10 ACD 0.222 0.570 100 12.621 26.396 100 8.561 32.418 93 MuLV Neg. Control >50 >50 4 >50 >50 0 *(T/F): Transmitted/Founder Virus
TABLE-US-00021 TABLE 20 Neutralization assay data of N49 P19, NVS49 P9, and NVS49 P9.1 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 20 N49P19 NVS49 P9 NVS49 P9.1 Virus ID Clade* IC50 IC80 MPI IC50 IC80 MPI IC50 IC80 MPI 6535.3 B 9.091 28.290 95 0.552 2.256 100 0.232 2.062 100 QH0692.42 B 2.560 13.420 98 0.352 1.210 100 0.107 0.709 100 SC422661.8 B 0.425 1.415 100 0.241 0.691 100 0.127 0.468 100 PVO.4 B 0.850 2.946 100 0.179 0.628 100 0.181 0.874 100 TRO.11 B 0.394 1.301 100 0.044 0.177 100 0.052 0.196 100 AC10.0.29 B >50 >50 13 0.285 0.986 100 0.360 1.877 99 RHPA4259.7 B >50 >50 42 0.023 0.078 100 0.021 0.066 100 THRO4156.18 B 17.077 >50 69 1.407 8.213 99 1.650 9.443 99 REJO4541.67 B 0.169 0.859 100 0.071 0.291 100 0.064 0.231 100 TRJO4551.58 B 1.230 5.404 99 0.036 0.128 100 0.035 0.120 100 WITO4160.33 B 0.263 1.291 100 0.058 0.165 100 0.051 0.241 100 CAAN5342.A2 B 1.241 4.402 100 2.091 7.380 100 2.160 8.001 99 WEAU_d15_410_787 B (T/F) 0.177 0.612 100 0.063 0.162 100 0.068 0.253 100 1006_11_C3_1601 B (T/F) 0.400 1.330 100 0.076 0.231 100 0.135 0.448 100 1054_07_TC4_1499 B (T/F) 2.854 11.746 99 0.885 2.445 100 1.798 7.198 100 1056_10_TA11_1826 B (T/F) 0.925 3.154 100 0.386 1.325 100 0.617 3.028 100 1012_11_TC21_3257 B (T/F) 0.204 0.895 100 0.194 0.591 100 0.382 1.448 100 6240_08_TA5_4622 B (T/F) 2.102 7.516 100 0.308 1.259 100 0.716 2.552 100 6244_13_B5_4576 B (T/F) 0.571 1.818 100 0.132 0.366 100 0.343 1.504 100 62357_14_D3_4589 B (T/F) 37.353 >50 51 0.074 0.307 100 0.227 1.086 99 SC05_8C11_2344 B (T/F) 0.954 3.296 100 0.473 1.608 100 1.302 4.392 100 Du156.12 C 2.175 17.013 87 0.043 0.130 100 0.066 0.252 100 Du172.17 C >50 >50 30 0.023 0.178 100 0.026 0.182 100 Du422.1 C >50 >50 30 >50 >50 17 >50 >50 17 ZM197M.PB7 C 38.389 >50 56 0.083 0.339 100 0.049 0.283 100 ZM214M.PL15 C 1.850 14.820 98 0.477 1.989 100 0.561 2.963 100 ZM233M.PB6 C >50 >50 4 >50 >50 48 >50 >50 6 ZM249M.PL1 C >50 >50 33 0.152 0.801 97 0.206 1.916 95 ZM53M.PB12 C 1.187 4.143 100 0.362 1.137 100 0.447 1.287 100 ZM109F.PB4 C 0.282 2.660 92 0.075 0.349 99 0.116 0.497 95 ZM135M.PL10a C >50 >50 0 0.224 0.863 97 0.332 1.529 98 CAP45.2.00.G3 C >50 >50 5 0.294 1.986 98 1.722 19.580 88 CAP210.2.00.E8 C >50 >50 0 >50 >50 32 >50 >50 2 HIV-001428-2.42 C >50 >50 44 0.022 0.106 99 0.037 0.120 99 HIV-0013095-2.11 C >50 >50 18 0.179 0.635 100 0.358 1.262 100 HIV-16055-2.3 C >50 >50 30 0.077 0.274 100 0.102 0.362 100 HIV-16845-2.22 C 14.499 >50 73 1.090 9.957 98 2.131 10.747 98 Ce1086_B2 C (T/F) 0.613 2.680 97 0.341 1.687 97 0.814 4.077 96 Ce0393_C3 C (T/F) >50 >50 31 0.189 0.595 100 0.284 1.221 100 Ce1176_A3 C (T/F) 1.567 9.839 99 0.258 0.919 100 0.440 0.998 100 Ce2010_F5 C (T/F) 7.660 45.381 82 0.089 0.271 100 0.120 0.380 100 Ce0682_E4 C (T/F) 0.607 2.323 98 0.049 0.185 100 0.076 0.264 100 Ce1172_H1 C (T/F) >50 >50 3 8.796 >50 70 >50 >50 42 Ce2060_G9 C (T/F) 1.056 5.615 98 0.419 2.190 100 0.781 3.540 100 Ce703010054_2A2 C (T/F) >50 >50 0 0.187 0.898 98 0.433 3.177 96 BF1266.431a C (T/F) >50 >50 15 0.154 0.519 99 0.808 3.142 95 246F C1G C (T/F) >50 >50 19 >50 >50 48 >50 >50 11 249M B10 C (T/F) >50 >50 39 0.203 1.411 97 0.305 1.644 96 ZM247v1(Rev-) C (T/F) >50 >50 5 0.622 3.301 100 1.878 17.837 87 7030102001E5(Rev-) C (T/F) 0.950 4.701 100 0.453 2.028 100 0.635 2.016 100 1394C9G1(Rev-) C (T/F) >50 >50 17 >50 >50 40 >50 >50 6 Ce704809221_1B3 C (T/F) 3.746 19.191 93 0.133 0.605 100 0.162 0.661 100 CNE19 BC 20.710 >50 59 0.031 0.136 100 0.033 0.146 100 CNE20 BC 0.124 0.817 99 0.465 2.432 97 1.514 16.392 90 CNE21 BC >50 >50 21 >50 >50 36 >50 >50 15 CNE17 BC >50 >50 30 0.364 1.699 100 0.251 1.231 100 CNE30 BC 1.269 4.275 100 0.259 0.693 100 0.271 0.971 100 CNE52 BC 0.175 0.491 100 0.026 0.090 100 0.043 0.122 100 CNE53 BC 1.501 >50 78 0.123 0.462 99 0.140 0.647 99 CNE58 BC >50 >50 43 0.030 0.081 100 0.027 0.074 100 MS208.A1 A 0.501 1.987 99 0.080 0.294 100 0.082 0.248 100 Q23.17 A 0.234 1.368 100 0.032 0.091 100 0.029 0.106 100 Q461.e2 A 0.774 4.053 100 0.271 0.953 100 0.363 1.146 100 Q769.d22 A 0.140 0.519 100 0.036 0.123 100 0.039 0.148 100 Q259.d2.17 A >50 >50 11 0.807 >50 74 >50 >50 48 Q842.d12 A 0.182 0.601 100 0.019 0.057 100 0.027 0.075 100 3415.v1.c1 A 0.462 2.038 100 0.137 0.502 100 0.210 1.372 100 3365.v2.c2 A 0.579 2.091 99 0.059 0.198 100 0.055 0.725 100 0260.v5.c36 A 1.250 8.524 99 0.466 1.852 100 0.519 0.151 100 191955_A11 A (T/F) >50 >50 16 >50 >50 35 >50 >50 26 191084 B7-19 A (T/F) 0.357 1.340 100 0.042 0.129 100 0.097 0.259 100 9004SS_A3_4 A (T/F) 0.661 3.086 100 0.175 0.751 100 0.323 1.129 100 T257-31 CRF02_AG 0.661 2.995 100 0.216 0.821 100 0.508 1.819 100 928-28 CRF02_AG 2.726 9.630 97 0.094 0.339 100 0.163 0.695 100 263-8 CRF02_AG 0.344 1.510 99 0.032 0.161 100 0.049 0.170 100 T250-4 CRF02_AG >50 >50 7 >50 >50 25 >50 >50 7 T251-18 CRF02_AG 0.938 4.204 100 0.147 0.517 100 0.127 0.352 100 T278-50 CRF02_AG >50 >50 35 >50 >50 25 >50 >50 5 T255-34 CRF02_AG 0.948 6.223 97 0.051 0.247 100 0.092 0.286 100 211-9 CRF02_AG 3.230 18.983 93 0.136 0.485 100 0.170 0.579 100 235-47 CRF02_AG 0.440 3.227 89 0.072 0.271 98 0.045 0.352 98 620345.c01 CRF01_AE >50 >50 0 >50 >50 9 >50 >50 7 CNE8 CRF01_AE 0.772 4.840 97 0.056 0.260 100 0.228 0.921 100 C1080.c03 CRF01_AE 0.682 3.053 100 0.119 0.575 100 0.156 0.939 100 R2184.c04 CRF01_AE 0.132 0.445 100 0.015 0.034 100 0.019 0.057 100 R1166.c01 CRF01_AE 0.629 2.693 100 0.064 0.223 100 0.073 0.232 100 R3265.c06 CRF01_AE 0.634 2.735 100 0.135 0.812 100 0.469 1.621 100 C2101.c01 CRF01_AE 0.978 4.192 100 0.051 0.209 100 0.085 0.227 100 C3347.c11 CRF01_AE 3.943 25.715 87 0.023 0.065 100 0.029 0.083 100 C4118.c09 CRF01_AE 0.386 1.361 100 0.072 0.200 100 0.062 0.218 100 CNE5 CRF01_AE 0.748 3.290 100 0.183 0.641 100 BJOX009000.02.4 CRF01_AE 3.664 25.579 87 0.727 3.293 100 0.965 3.934 100 BJOX015000.11.5 CRF01_AE 0.756 3.165 99 0.089 0.375 100 0.068 0.478 100 (T/F) BJOX010000.06.2 CRF01_AE 4.159 19.069 96 0.482 2.337 100 0.883 4.259 100 (T/F) BJOX025000.01.1 CRF01_AE >50 >50 24 0.055 0.252 100 0.131 0.670 100 (T/F) BJOX028000.10.3 CRF01_AE 6.996 >50 80 0.031 0.134 100 0.040 0.200 100 (T/F) X1193_c1 G 0.540 3.147 99 0.025 0.081 100 0.042 0.150 100 P0402_c2_11 G 0.448 2.630 100 0.028 0.085 100 0.060 0.163 100 X1254_c3 G >50 >50 48 0.040 0.104 100 0.101 0.278 100 X2088_c9 G 1.402 3.760 99 0.070 0.252 100 0.091 0.392 100 X2131_C1_B5 G 2.522 14.446 91 0.082 0.274 100 0.143 0.476 100 P1981_C5_3 G 20.237 >50 68 0.308 0.701 100 0.469 1.523 100 X1632_S2_B10 G >50 >50 14 46.721 >50 51 >50 >50 10 3016.v5.c45 D >50 >50 9 0.060 0.209 100 0.076 0.223 100 A07412M1.vrc12 D 3.892 48.832 80 0.142 0.636 100 0.189 0.947 100 231965.c01 D >50 >50 22 0.253 1.159 100 0.474 2.239 100 231966.c02 D 0.653 4.553 100 0.074 0.244 100 0.067 0.237 100 3817.v2.c59 CD >50 >50 26 1.092 5.859 100 1.347 8.262 95 6480.v4.c25 CD 3.035 28.156 87 0.017 0.047 100 0.030 0.082 100 6952.v1.c20 CD >50 >50 0 0.095 0.258 100 0.119 0.389 100 6811.v7.c18 CD 1.305 4.709 97 0.249 1.195 99 0.398 1.757 97 89-F1_2_25 CD >50 >50 7 >50 >50 23 >50 >50 23 3301.v1.c24 AC 0.302 1.423 98 0.072 0.246 100 0.059 0.178 100 6041.v3.c23 AC >50 >50 32 0.089 0.777 95 0.282 6.781 90 6540.v4.c1 AC 11.229 >50 63 >50 >50 49 >50 >50 30 6545.v4.c1 AC >50 >50 2 >50 >50 20 >50 >50 6 0815.v3.c3 ACD 0.176 0.743 100 0.020 0.057 100 0.012 0.041 100 3103.v3.c10 ACD >50 >50 0 17.571 >50 55 >50 >50 45 MuLV Neg. Control >50 >50 0 >50 >50 13 21.396 22.133 *(T/F): Transmitted/Founder Virus
TABLE-US-00022 TABLE 21 Neutralization assay data of N49 P23, N40P22, and PGT121 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 21 N49P23 N49P22 PGT121 Virus ID Clade* IC50 IC80 MPI IC50 IC80 MPI IC50 IC80 6535.3 B 0.127 0.855 100 >50 >50 5 0.002 0.008 QH0692.42 B 0.557 2.513 100 0.592 2.024 100 0.302 2.314 SC422661.8 B 0.069 0.245 100 0.120 0.544 100 0.038 0.177 PVO.4 B 0.087 0.430 100 0.433 1.492 100 0.098 0.343 TRO.11 B 0.094 0.336 100 0.283 1.365 99 0.005 0.025 AC10.0.29 B >50 >50 0 >50 >50 0 0.024 0.073 RHPA4259.7 B 0.010 0.040 100 0.637 5.596 92 0.015 0.035 THRO4156.18 B 1.979 10.746 97 >50 >50 32 >50 >50 REJO4541.67 B 0.008 0.050 100 0.792 12.089 84 4.774 >50 TRJO4551.58 B 0.016 0.068 100 >50 >50 28 1.314 37.88 WITO4160.33 B 0.045 0.205 100 4.862 >50 74 0.334 2.351 CAAN5342.A2 B 0.412 1.381 100 0.425 1.884 99 0.007 0.023 WEAU_d15_410_787 B (T/F) 0.009 0.031 100 1.541 >50 76 0.026 0.077 1006_11_C3_1601 B (T/F) 0.015 0.054 100 0.898 14.693 83 0.002 0.007 1054_07_TC4_1499 B (T/F) 0.272 1.211 100 0.654 2.893 100 0.064 0.304 1056_10_TA11_1826 B (T/F) 0.075 0.483 100 2.991 >50 69 0.004 0.081 1012_11_TC21_3257 B (T/F) 0.013 0.085 100 15.726 >50 53 0.003 0.017 6240_08_TA5_4622 B (T/F) 0.215 1.000 100 2.330 9.760 93 0.033 0.127 6244_13_B5_4576 B (T/F) 0.151 0.570 100 0.565 2.482 97 0.061 0.347 62357_14_D3_4589 B (T/F) 0.057 0.760 95 >50 >50 13 2.597 >50 SC05_8C11_2344 B (T/F) 0.133 0.548 100 0.251 2.159 97 0.019 0.071 Du156.12 C >50 >50 0 >50 >50 4 0.004 0.017 Du172.17 C >50 >50 45 >50 >50 4 0.033 0.257 Du422.1 C >50 >50 4 >50 >50 4 0.039 0.143 ZM197M.PB7 C 0.100 0.385 100 2.583 25.090 84 >50 >50 ZM214M.PL15 C 0.061 0.614 100 >50 >50 25 0.460 2.039 ZM233M.PB6 C >50 >50 14 >50 >50 2 3.689 ##### ZM249M.PL1 C >50 >50 2 >50 >50 0 >50 >50 ZM53M.PB12 C 0.135 0.651 100 1.197 11.765 91 <0.001 0.004 ZM109F.PB4 C 0.039 0.180 100 >50 >50 1 8.639 >50 ZM135M.PL10a C >50 >50 36 >50 >50 27 0.716 5.246 CAP45.2.00.G3 C >50 >50 0 >50 >50 0 1.634 >50 CAP210.2.00.E8 C >50 >50 2 >50 >50 6 26.301 >50 HIV-001428-2.42 C 0.036 0.241 98 0.730 7.097 91 0.014 0.051 HIV-0013095-2.11 C 0.114 0.899 96 >50 >50 9 >50 >50 HIV-16055-2.3 C >50 >50 17 4.655 39.690 83 0.153 4.804 HIV-16845-2.22 C >50 >50 20 4.600 41.933 87 3.969 44.570 Ce1086_B2 C (T/F) 0.063 0.303 100 >50 >50 0 <0.001 <0.001 Ce0393_C3 C (T/F) >50 >50 29 >50 >50 12 >50 >50 Ce1176_A3 C (T/F) 0.297 1.384 100 >50 >50 23 0.013 0.035 Ce2010_F5 C (T/F) 1.590 11.612 91 2.053 11.680 93 >50 >50 Ce0682_E4 C (T/F) 1.391 >50 76 1.132 >50 75 >50 >50 Ce1172_H1 C (T/F) >50 >50 0 >50 >50 0 0.011 0.036 Ce2060_G9 C (T/F) 0.180 0.849 99 >50 >50 19 >50 >50 Ce703010054_2A2 C (T/F) >50 >50 35 >50 >50 0 >50 >50 BF1266.431a C (T/F) 0.083 0.440 99 >50 >50 0 >50 >50 246F C1G C (T/F) >50 >50 11 >50 >50 10 0.041 0.140 249M B10 C (T/F) >50 >50 20 >50 >50 4 >50 >50 ZM247v1(Rev-) C (T/F) >50 >50 24 >50 >50 11 0.028 0.082 7030102001E5(Rev-) C (T/F) 0.172 0.797 100 >50 >50 21 0.009 0.027 1394C9G1(Rev-) C (T/F) >50 >50 34 >50 >50 35 0.264 1.948 Ce704809221_1B3 C (T/F) 0.083 0.643 100 15.799 >50 60 0.025 0.126 CNE19 BC >50 >50 40 0.873 8.185 92 0.008 0.105 CNE20 BC 45.443 >50 51 >50 >50 18 <0.001 0.002 CNE21 BC >50 >50 0 >50 >50 0 0.007 0.035 CNE17 BC 0.221 0.809 100 46.299 >50 53 7.600 >50 CNE30 BC 0.143 0.680 100 >50 >50 30 0.078 0.291 CNE52 BC 0.014 0.051 100 1.701 10.016 91 2.045 18.929 CNE53 BC 0.026 0.095 100 0.779 5.111 93 0.007 0.024 CNE58 BC >50 >50 9 28.372 >50 57 >50 >50 MS208.A1 A 0.048 0.279 100 >50 >50 17 >50 >50 Q23.17 A 0.010 0.046 100 0.321 2.400 96 <0.001 0.005 Q461.e2 A 0.099 0.464 100 11.157 >50 66 >50 >50 Q769.d22 A 0.005 0.025 100 0.043 0.259 100 >50 >50 Q259.d2.17 A 0.362 7.795 91 >50 >50 15 15.379 >50 Q842.d12 A 0.007 0.022 100 0.079 0.359 100 0.005 0.022 3415.v1.c1 A 0.054 0.190 100 0.198 1.410 96 NT NT 3365.v2.c2 A 0.017 0.053 100 0.254 0.857 98 NT NT 0260.v5.c36 A 0.261 0.919 100 1.673 14.382 93 0.053 0.197 191955_A11 A (T/F) >50 >50 31 >50 >50 9 >50 >50 191084 B7-19 A (T/F) 0.013 0.043 100 0.343 1.639 98 0.011 0.032 9004SS_A3_4 A (T/F) 0.043 0.223 100 >50 >50 42 <0.001 0.003 T257-31 CRF02_AG 0.178 1.167 100 >50 >50 43 >50 >50 928-28 CRF02_AG 0.021 0.122 100 >50 >50 32 44.189 >50 263-8 CRF02_AG 0.012 0.096 100 1.829 18.462 89 0.648 3.961 T250-4 CRF02_AG >50 >50 0 >50 >50 5 <0.001 0.005 T251-18 CRF02_AG >50 >50 18 0.580 3.092 99 29.016 >50 T278-50 CRF02_AG >50 >50 22 >50 >50 25 >50 >50 T255-34 CRF02_AG >50 >50 20 38.766 >50 55 18.695 >50 211-9 CRF02_AG >50 >50 14 >50 >50 18 0.852 4.202 235-47 CRF02_AG 0.085 0.749 90 0.037 0.373 99 0.137 0.727 620345.c01 CRF01_AE >50 >50 0 >50 >50 0 >50 >50 CNE8 CRF01_AE 6.353 47.322 81 5.350 40.442 86 >50 >50 C1080.c03 CRF01_AE 1.328 11.328 96 15.863 >50 64 >50 >50 R2184.c04 CRF01_AE 0.008 0.032 100 0.077 0.342 100 >50 >50 R1166.c01 CRF01_AE 0.123 0.455 100 0.232 0.799 100 >50 >50 R3265.c06 CRF01_AE >50 >50 0 3.575 >50 67 >50 >50 C2101.c01 CRF01_AE 0.056 0.362 100 0.171 1.059 99 NT NT C3347.c11 CRF01_AE 0.007 0.032 100 1.181 12.263 89 >50 >50 C4118.c09 CRF01_AE 0.026 0.120 100 1.821 26.577 93 >50 >50 CNE5 CRF01_AE 0.058 0.226 100 0.281 1.553 96 >50 >50 BJOX009000.02.4 CRF01_AE >50 >50 8 17.324 >50 64 2.858 31.728 BJOX015000.11.5 CRF01_AE >50 >50 0 1.258 10.790 89 >50 >50 (T/F) BJOX010000.06.2 CRF01_AE 0.284 2.078 100 >50 >50 0 >50 >50 (T/F) BJOX025000.01.1 CRF01_AE >50 >50 0 >50 >50 0 >50 >50 (T/F) BJOX028000.10.3 CRF01_AE 0.021 0.123 98 >50 >50 0 >50 >50 (T/F) X1193_c1 G 0.008 0.038 100 3.492 28.543 85 0.016 0.086 P0402_c2_11 G 0.022 0.065 100 5.928 >50 73 0.005 0.016 X1254_c3 G 0.022 0.111 100 3.055 29.794 84 0.014 0.080 X2088_c9 G >50 >50 10 >50 >50 8 0.003 0.013 X2131_C1_B5 G 0.080 0.280 100 >50 >50 44 0.004 0.023 P1981_C5_3 G 0.096 0.264 100 >50 >50 48 <0.001 <0.001 X1632_S2_B10 G 0.130 >50 62 >50 >50 0 >50 >50 3016.v5.c45 D 0.027 0.092 100 >50 >50 23 >50 >50 A07412M1.vrc12 D 0.114 0.830 97 9.600 >50 63 0.009 0.075 231965.c01 D >50 >50 35 2.044 35.200 83 >50 >50 231966.c02 D 0.035 0.158 100 3.351 >50 69 >50 >50 3817.v2.c59 CD >50 >50 41 >50 >50 12 18.888 >50 6480.v4.c25 CD >50 >50 29 0.809 32.124 86 <0.001 0.004 6952.v1.c20 CD 0.007 0.023 100 33.248 >50 56 0.056 0.420 6811.v7.c18 CD 17.656 >50 61 1.003 9.719 92 <0.001 0.001 89-F1_2_25 CD >50 >50 5 >50 >50 10 >50 >50 3301.v1.c24 AC 0.017 0.058 100 2.819 48.778 80 0.008 0.019 6041.v3.c23 AC 7.466 >50 63 7.547 >50 66 >50 >50 6540.v4.c1 AC >50 >50 29 >50 >50 29 >50 >50 6545.v4.c1 AC >50 >50 10 >50 >50 17 >50 >50 0815.v3.c3 ACD 0.004 0.014 100 0.161 1.080 98 0.025 0.111 3103.v3.c10 ACD 43.751 >50 54 >50 >50 16 0.009 0.028 MuLV Neg. Control >50 >50 9 >50 >50 8 *(T/F): Transmitted/Founder Virus
TABLE-US-00023 TABLE 22 Neutralization assay data of PGT128, PGT145, PGDM1400, and PGT151 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 22 PGT128 PGT145 PGDM1400 PGT151 Virus ID Clade* IC50 IC80 IC50 IC80 IC50 IC80 IC50 IC80 6535.3 B 0.004 0.016 >50 >50 >50 >50 <0.001 0.012 QH0692.42 B 0.029 0.081 >50 >50 >50 >50 0.029 0.522 SC422661.8 B 1.078 18.4 0.024 0.082 0.436 6.530 0.005 0.028 PVO.4 B 0.011 0.028 0.192 0.870 0.484 6.487 0.017 >50 TRO.11 B 0.019 0.045 0.040 0.118 0.283 1.376 >50 >50 AC10.0.29 B 0.008 0.018 0.010 0.025 0.105 0.869 0.007 0.023 RHPA4259.7 B 0.026 0.109 0.029 0.095 0.447 2.194 0.006 0.033 THRO4156.18 B >50 >50 0.010 0.029 0.072 0.348 >50 >50 REJO4541.67 B >50 >50 <0.001 0.001 0.028 0.992 0.018 0.119 TRJO4551.58 B 0.018 0.041 >50 >50 >50 >50 0.496 4.217 WITO4160.33 B >50 >50 <0.001 0.001 <0.001 0.008 0.002 0.021 CAAN5342.A2 B 0.514 >50 6.675 25.524 >50 >50 0.005 0.038 WEAU_d15_410_787 B (T/F) 0.032 0.153 1.951 31.540 0.103 1.366 0.010 >50 1006_11_C3_1601 B (T/F) 0.011 0.040 >50 >50 4.318 >50 2.891 >50 1054_07_TC4_1499 B (T/F) 0.035 0.312 >50 >50 >50 >50 0.008 0.032 1056_10_TA11_1826 B (T/F) <0.001 0.008 0.230 0.938 3.258 11.840 0.008 0.048 1012_11_TC21_3257 B (T/F) 0.011 0.033 0.009 0.025 0.049 0.230 >50 >50 6240_08_TA5_4622 B (T/F) 0.019 0.055 >50 >50 >50 >50 0.046 0.420 6244_13_B5_4576 B (T/F) 0.020 0.076 7.266 44.473 >50 >50 >50 >50 62357_14_D3_4589 B (T/F) 1.144 >50 >50 >50 >50 >50 0.004 0.023 SC05_8C11_2344 B (T/F) 0.017 0.041 0.093 0.248 1.264 3.587 0.060 >50 Du156.12 C 0.017 0.046 0.001 0.018 0.002 0.008 0.005 0.023 Du172.17 C 0.028 0.080 >50 >50 5.179 21.980 0.003 0.013 Du422.1 C 0.039 0.113 22.564 >50 0.694 >50 4.604 >50 ZM197M.PB7 C >50 >50 0.628 5.239 0.063 0.285 0.003 0.033 ZM214M.PL15 C 1.498 29.44 >50 >50 >50 >50 0.008 0.045 ZM233M.PB6 C >50 >50 0.025 0.680 <0.001 0.001 0.027 5.156 ZM249M.PL1 C 39.66 >50 1.442 >50 0.016 0.701 0.004 0.015 ZM53M.PB12 C >50 >50 0.367 11.774 0.002 0.011 >50 >50 ZM109F.PB4 C >50 >50 0.042 0.234 0.057 0.382 0.609 >50 ZM135M.PL10a C >50 >50 >50 >50 >50 >50 >50 >50 CAP45.2.00.G3 C >50 >50 0.001 0.007 <0.001 0.002 0.007 >50 CAP210.2.00.E8 C >50 >50 37.807 >50 0.034 0.514 0.010 0.208 HIV-001428-2.42 C 0.026 0.095 0.001 0.083 <0.001 0.027 0.027 >50 HIV-0013095-2.11 C >50 >50 9.271 >50 0.003 0.012 >50 >50 HIV-16055-2.3 C >50 >50 0.003 0.051 <0.001 0.003 0.143 >50 HIV-16845-2.22 C 0.181 0.834 >50 >50 >50 >50 >50 >50 >50 >50 Ce1086_B2 C (T/F) >50 >50 >50 >50 >50 >50 2.653 >50 Ce0393_C3 C (T/F) >50 >50 0.112 1.225 0.015 0.044 >50 >50 Ce1176_A3 C (T/F) 0.009 0.026 >50 >50 0.587 11.619 0.006 0.018 Ce2010_F5 C (T/F) >50 >50 >50 >50 >50 >50 >50 >50 Ce0682_E4 C (T/F) >50 >50 33.578 >50 0.010 0.043 >50 >50 Ce1172_H1 C (T/F) 0.013 0.038 0.260 2.062 0.021 0.060 >50 >50 Ce2060_G9 C (T/F) >50 >50 0.016 0.205 0.001 0.007 0.135 >50 Ce703010054_2A2 C (T/F) >50 >50 >50 >50 0.015 0.260 0.013 0.075 BF1266.431a C (T/F) >50 >50 1.345 18.508 0.003 0.011 >50 >50 246F C1G C (T/F) 0.005 0.014 >50 >50 >50 >50 >50 >50 249M B10 C (T/F) 7.868 >50 0.947 41.307 0.039 0.786 0.007 0.023 ZM247v1(Rev-) C (T/F) 0.021 0.081 7.190 >50 0.019 0.051 0.009 >50 7030102001E5(Rev-) C (T/F) 0.007 0.022 >50 >50 >50 >50 0.001 0.011 1394C9G1(Rev-) C (T/F) 0.011 0.034 0.001 0.005 <0.001 0.004 0.022 0.733 Ce704809221_1B3 C (T/F) 0.026 0.072 0.102 0.710 0.280 1.099 >50 >50 CNE19 BC >50 >50 0.103 1.128 0.001 0.007 0.014 >50 CNE20 BC 0.001 0.004 0.318 6.970 0.004 0.014 >50 >50 CNE21 BC 0.010 0.026 <0.001 0.009 <0.001 0.002 0.004 0.012 CNE17 BC 0.432 1.993 0.091 1.156 0.007 0.021 0.093 >50 CNE30 BC 2.055 12.610 >50 >50 >50 >50 >50 >50 CNE52 BC >50 >50 0.020 0.091 1.147 9.970 0.004 0.017 CNE53 BC 0.010 0.049 0.006 0.084 0.077 0.246 >50 >50 CNE58 BC 13.77 >50 0.183 7.407 0.004 0.014 0.004 0.016 MS208.A1 A >50 >50 0.334 17.97 <0.001 0.139 <0.001 0.003 Q23.17 A 0.009 0.024 1.317 >50 <0.001 0.003 0.007 0.045 Q461.e2 A >50 >50 7.801 >50 0.089 0.572 12.553 >50 Q769.d22 A >50 >50 0.260 25.47 0.002 0.018 >50 >50 Q259.d2.17 A >50 >50 44.232 >50 0.166 >50 >50 >50 Q842.d12 A 0.008 0.102 0.032 0.552 <0.001 0.002 0.001 0.014 3415.v1.c1 A NT NT NT NT NT NT NT NT 3365.v2.c2 A NT NT NT NT NT NT NT NT 0260.v5.c36 A 0.058 0.162 >50 >50 0.033 0.175 0.427 >50 191955_A11 A (T/F) 14.2 >50 <0.001 <0.001 0.001 0.021 0.004 0.014 191084 B7-19 A (T/F) 0.022 0.152 0.028 0.345 <0.001 0.007 0.004 0.016 9004SS_A3_4 A (T/F) 0.002 0.010 0.018 0.419 <0.001 0.002 0.003 0.015 T257-31 CRF02_AG >50 >50 5.177 >50 0.002 0.031 0.003 0.010 928-28 CRF02_AG >50 >50 >50 >50 0.220 1.246 >50 >50 263-8 CRF02_AG 0.342 3.303 32.071 >50 0.020 0.103 >50 >50 T250-4 CRF02_AG <0.001 0.007 0.002 0.039 <0.001 <0.001 0.005 0.014 T251-18 CRF02_AG >50 >50 0.351 1.393 8.615 43.108 0.327 >50 T278-50 CRF02_AG 0.031 0.089 10.324 >50 0.574 4.942 0.130 >50 T255-34 CRF02_AG >50 >50 >50 >50 0.654 >50 0.002 0.010 211-9 CRF02_AG 0.062 0.215 0.129 0.923 0.088 0.393 0.002 0.010 235-47 CRF02_AG >50 >50 3.212 36.02 0.005 0.032 38.12 >50 620345.c01 CRF01_AE >50 >50 0.241 19.74 0.009 40.337 >50 >50 CNE8 CRF01_AE 0.019 0.052 0.272 2.241 <0.001 0.010 >50 >50 C1080.c03 CRF01_AE 0.283 2.062 0.024 0.146 <0.001 <0.001 >50 >50 R2184.c04 CRF01_AE 20.056 >50 0.073 0.376 0.003 0.012 >50 >50 R1166.c01 CRF01_AE >50 >50 2.274 17.89 1.431 6.921 >50 >50 R3265.c06 CRF01_AE >50 >50 0.037 0.227 0.013 0.041 >50 >50 C2101.c01 CRF01_AE NT NT NT NT 0.006 0.021 NT NT C3347.c11 CRF01_AE 0.001 0.007 0.005 0.044 0.011 0.051 >50 >50 C4118.c09 CRF01_AE >50 >50 <0.001 0.004 0.002 0.005 >50 >50 CNE5 CRF01_AE 0.018 0.253 0.001 0.013 <0.001 <0.001 >50 >50 BJOX009000.02.4 CRF01_AE <0.001 0.005 0.117 0.745 0.443 1.287 >50 >50 BJOX015000.11.5 CRF01_AE 0.001 0.011 >50 >50 0.520 10.577 >50 >50 (T/F) BJOX010000.06.2 CRF01_AE 4.817 >50 >50 >50 0.605 5.870 >50 >50 (T/F) BJOX025000.01.1 CRF01_AE >50 >50 6.632 >50 0.087 0.573 >50 >50 (T/F) BJOX028000.10.3 CRF01_AE 0.024 0.070 >50 >50 0.072 0.963 >50 >50 (T/F) X1193_c1 G >50 >50 0.013 0.091 0.219 0.781 0.004 0.027 P0402_c2_11 G 0.007 0.020 0.009 0.038 0.010 0.033 <0.001 0.004 X1254_c3 G >50 >50 5.597 >50 >50 >50 0.309 >50 X2088_c9 G >50 >50 >50 >50 >50 >50 >50 >50 X2131_C1_B5 G >50 >50 0.018 0.065 0.128 0.568 0.007 0.036 P1981_C5_3 G 0.019 0.088 5.413 >50 0.104 4.601 0.021 >50 X1632_S2_B10 G >50 >50 0.012 2.030 0.055 1.445 0.061 >50 3016.v5.c45 D >50 >50 >50 >50 >50 >50 0.174 40.676 A07412M1.vrc12 D >50 >50 0.002 0.023 0.032 0.127 0.246 >50 231965.c01 D >50 >50 0.403 14.96 0.025 1.280 <0.001 0.012 231966.c02 D 3.197 >50 <0.001 0.010 0.001 0.004 0.025 0.183 3817.v2.c59 CD 0.003 0.023 >50 >50 >50 >50 >50 >50 6480.v4.c25 CD 0.003 0.019 >50 >50 >50 >50 NT NT 6952.v1.c20 CD >50 >50 3.120 >50 0.501 11.839 49.232 >50 6811.v7.c18 CD 0.003 0.050 >50 >50 >50 >50 >50 >50 89-F1_2_25 CD >50 >50 0.800 19.97 0.001 0.005 22.889 >50 3301.v1.c24 AC 0.067 0.499 0.219 4.223 0.048 0.245 0.002 0.019 6041.v3.c23 AC >50 >50 0.964 >50 0.013 0.289 0.004 0.024 6540.v4.c1 AC 11.8 >50 0.080 5.086 0.004 0.028 0.006 0.030 6545.v4.c1 AC >50 >50 0.114 3.100 0.003 0.039 0.004 0.026 0815.v3.c3 ACD 0.030 0.137 >50 >50 >50 >50 >50 >50 3103.v3.c10 ACD 0.014 0.033 0.043 0.164 0.671 1.998 0.014 0.202 MuLV Neg. Control *(T/F): Transmitted/Founder Virus
TABLE-US-00024 TABLE 23 Neutralization assay data of 10-1074, 10E8, PG9, and PG16 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 23 10-1074 10E8 PG9 PG16 Virus ID Clade* IC50 IC80 IC50 IC80 IC50 IC80 IC50 IC80 6535.3 B 0.014 0.026 0.081 1.32 0.176 0.627 0.653 >50 QH0692.42 B 0.191 0.929 0.442 4.046 >50 >50 >50 >50 SC422661.8 B 0.091 0.418 0.445 2.715 0.553 7.054 0.858 >50 PVO.4 B 0.074 0.360 0.747 >10 10.56 >50 15.27 >50 TRO.11 B 0.008 0.057 0.034 0.250 43.25 >50 1.922 49.59 AC10.0.29 B 0.022 0.110 0.132 1.534 0.117 0.845 0.023 0.417 RHPA4259.7 B 0.021 0.118 0.728 5.275 22.620 >50 0.375 3.925 THRO4156.18 B >50 >50 0.156 0.884 24.48 >50 6.825 >50 REJO4541.67 B >50 >50 0.203 1.686 0.012 0.163 0.039 13.36 TRJO4551.58 B 0.17 0.634 0.475 4.783 0.503 4.061 1.008 24.46 WITO4160.33 B 0.185 2.112 0.094 1.226 0.010 0.037 0.002 0.012 CAAN5342.A2 B 0.007 0.036 1.284 9.281 4.603 26.99 1.831 33.0 WEAU_d15_410_787 B (T/F) 0.104 0.375 >10 >10 4.078 >50 0.435 >50 1006_11_C3_1601 B (T/F) 0.003 0.013 0.172 2.999 0.366 1.906 >50 >50 1054_07_TC4_1499 B (T/F) 0.129 0.563 0.046 0.305 >50 >50 >50 >50 1056_10_TA11_1826 B (T/F) 0.038 0.272 0.134 1.028 6.339 38.26 0.315 1.952 1012_11_TC21_3257 B (T/F) 0.008 0.059 0.244 4.211 0.172 0.605 0.030 0.219 6240_08_TA5_4622 B (T/F) 0.068 0.306 0.756 5.591 1.747 6.333 >50 >50 6244_13_B5_4576 B (T/F) 0.202 0.922 0.086 0.809 >50 >50 >50 >50 62357_14_D3_4589 B (T/F) >50 >50 0.119 1.060 >50 >50 >50 >50 SC05_8C11_2344 B (T/F) 0.052 0.123 0.665 2.736 0.892 4.283 0.070 1.616 Du156.12 C 0.015 0.076 0.034 0.144 0.026 0.190 0.006 0.026 Du172.17 C 0.121 0.430 0.109 0.837 0.606 1.604 0.033 0.213 Du422.1 C 0.045 0.166 0.351 1.883 0.614 10.4 0.261 33.84 ZM197M.PB7 C >50 >50 0.148 0.802 0.899 4.517 1.496 22.79 ZM214M.PL15 C 0.174 2.367 1.962 >10 >50 >50 >50 >50 ZM233M.PB6 C 0.06 0.349 0.442 2.393 0.010 0.026 0.002 0.006 ZM249M.PL1 C >50 >50 1.511 6.294 0.166 1.410 0.127 18.07 ZM53M.PB12 C >50 >50 3.985 >10 0.069 0.221 0.007 0.051 ZM109F.PB4 C >50 >50 0.295 1.888 0.256 6.689 7.033 >50 ZM135M.PL10a C 0.069 0.367 0.171 1.186 20.56 >50 >50 >50 CAP45.2.00.G3 C >50 >50 1.817 >10 0.007 0.029 0.001 0.003 CAP210.2.00.E8 C >50 >50 0.798 3.892 0.040 0.173 0.010 0.042 HIV-001428-2.42 C 0.044 0.261 0.809 7.298 0.101 0.250 0.039 0.250 HIV-0013095-2.11 C >50 >50 0.031 0.213 0.010 0.030 0.002 0.006 HIV-16055-2.3 C >50 >50 0.722 6.495 0.021 0.06 0.17 >0.250 HIV-16845-2.22 C 1.169 5.835 0.068 0.495 4.398 >50 3.126 >50 Ce1086_B2 C (T/F) >50 >50 1.162 6.934 >50 >50 >50 >50 Ce0393_C3 C (T/F) >50 >50 1.179 7.423 0.020 0.055 0.002 0.006 Ce1176_A3 C (T/F) 0.019 0.070 0.867 3.923 0.014 0.044 0.003 0.011 Ce2010_F5 C (T/F) >50 >50 1.844 8.612 >50 >50 >50 >50 Ce0682_E4 C (T/F) >50 >50 1.215 7.709 0.196 0.698 0.026 0.202 Ce1172_H1 C (T/F) 0.047 0.166 0.722 4.818 0.070 0.206 0.007 0.037 Ce2060_G9 C (T/F) >50 >50 3.520 >10 0.045 0.172 0.075 1.712 Ce703010054_2A2 C (T/F) >50 >50 2.299 8.579 0.022 0.103 0.008 0.053 BF1266.431a C (T/F) >50 >50 2.687 >10 0.026 0.088 0.003 0.009 246F C1G C (T/F) 0.022 0.111 2.217 9.937 >50 >50 >50 >50 249M B10 C (T/F) >50 >50 1.271 9.050 0.095 0.290 0.026 1.249 ZM247v1(Rev-) C (T/F) 0.042 0.186 0.506 6.745 0.112 0.323 0.016 0.189 7030102001E5(Rev-) C (T/F) 0.006 0.021 6.707 >10 >50 >50 >50 >50 1394C9G1(Rev-) C (T/F) 0.050 0.191 0.963 5.327 0.029 0.114 0.008 0.056 Ce704809221_1B3 C (T/F) 0.139 0.696 0.058 0.319 0.037 0.137 0.021 0.116 CNE19 BC 50.00 >50 0.666 6.485 0.024 0.146 0.024 3.070 CNE20 BC <0.001 0.005 1.246 6.577 0.052 0.278 3.633 47.170 CNE21 BC 0.067 0.181 1.900 >10 0.038 0.094 0.007 0.026 CNE17 BC 2.686 13.297 1.114 5.255 0.210 0.484 0.029 0.132 CNE30 BC 0.363 1.200 1.560 9.842 >50 >50 >50 >50 CNE52 BC 1.226 13.147 0.304 3.056 0.029 0.081 0.009 0.026 CNE53 BC 0.039 0.141 0.326 1.916 0.131 0.60 >50 >50 CNE58 BC 0.267 0.968 1.289 8.268 0.038 0.122 0.044 0.250 MS208.A1 A >50 >50 0.609 3.717 0.005 0.014 0.002 0.004 Q23.17 A 0.006 0.021 1.637 7.040 0.027 0.082 0.008 0.043 Q461.e2 A >50 >50 2.111 >10 1.484 8.725 1.755 >50 Q769.d22 A >50 >50 1.667 10.000 0.025 0.068 0.005 0.018 Q259.d2.17 A >50 >50 4.587 >10 0.029 1.379 0.065 45.89 Q842.d12 A >50 >50 4.189 >10 0.023 0.081 0.032 >0.250 3415.v1.c1 A >50 >50 NT NT NT NT NT NT 3365.v2.c2 A 0.131 0.450 NT NT NT NT NT NT 0260.v5.c36 A 0.099 0.160 >10 >10 1.693 9.0 1.538 36.08 191955_A11 A (T/F) >50 >50 0.680 3.532 0.053 0.154 0.009 0.042 191084 B7-19 A (T/F) 0.032 0.128 5.156 >10 0.043 0.202 0.024 0.238 9004SS_A3_4 A (T/F) 0.011 0.030 1.160 6.761 0.088 0.251 0.020 0.111 T257-31 CRF02_AG >50 >50 0.880 5.779 0.037 0.124 0.007 0.021 928-28 CRF02_AG 0.847 4.696 0.384 1.962 0.101 0.450 0.034 0.237 263-8 CRF02_AG 0.666 6.527 0.196 1.728 0.293 1.050 0.347 4.975 T250-4 CRF02_AG <0.001 0.005 1.346 8.130 0.005 0.014 0.002 0.005 T251-18 CRF02_AG 1.081 7.395 2.357 >10 >50 >50 1.721 44.72 T278-50 CRF02_AG 2.146 18.276 0.949 7.460 0.393 3.192 0.227 18.65 T255-34 CRF02_AG >50 >50 0.778 4.207 0.023 0.134 0.024 0.125 211-9 CRF02_AG 0.112 0.425 1.091 6.508 0.049 0.243 0.029 0.144 235-47 CRF02_AG 0.05 0.163 0.384 2.072 0.233 1.201 0.092 4.506 620345.c01 CRF01_AE >50 >50 0.455 6.002 1.333 >50 >50 >50 CNE8 CRF01_AE >50 >50 NT NT NT NT NT NT C1080.c03 CRF01_AE >50 >50 0.061 0.731 0.004 0.013 0.001 0.003 R2184.c04 CRF01_AE >50 >50 0.272 2.455 0.255 1.237 0.670 >50 R1166.c01 CRF01_AE >50 >50 0.274 2.614 0.736 2.876 0.280 4.376 R3265.c06 CRF01_AE NT NT >10 >10 0.163 0.535 0.016 0.146 C2101.c01 CRF01_AE >50 >50 1.921 8.260 0.034 0.099 0.012 0.238 C3347.c11 CRF01_AE >50 >50 0.016 0.167 0.032 0.113 0.008 0.045 C4118.c09 CRF01_AE >50 >50 1.622 9.064 0.068 0.200 0.022 0.143 CNE5 CRF01_AE >50 >50 1.408 9.637 0.018 0.051 0.008 0.033 BJOX009000.02.4 CRF01_AE >50 >50 1.570 9.954 1.725 8.211 1.188 22.12 BJOX015000.11.5 CRF01_AE >50 >50 0.051 1.909 0.322 1.489 3.129 >50 (T/F) BJOX010000.06.2 CRF01_AE >50 >50 0.341 5.293 0.154 0.740 1.202 >50 (T/F) BJOX025000.01.1 CRF01_AE >50 >50 0.107 6.325 0.144 0.335 0.071 >50 (T/F) BJOX028000.10.3 CRF01_AE >50 >50 0.665 7.361 1.100 21.5 >50 >50 (T/F) X1193_c1 G 0.083 0.475 0.497 4.195 0.105 0.304 0.018 0.110 P0402_c2_11 G 0.012 0.039 0.172 3.288 0.296 2.367 0.025 0.411 X1254_c3 G 0.089 0.297 4.753 >10 0.065 0.304 0.023 0.218 X2088_c9 G 0.003 0.014 >10 >10 >50 >50 >50 >50 X2131_C1_B5 G 0.016 0.064 0.189 0.974 0.084 0.286 0.024 0.139 P1981_C5_3 G 0.005 0.017 0.057 0.285 0.258 2.512 0.376 >50 X1632_S2_B10 G >50 >50 1.976 9.684 0.107 0.934 0.012 0.076 3016.v5.c45 D >50 >50 0.476 4.393 2.455 >50 >50 >50 A07412M1.vrc12 D <0.001 0.048 0.576 6.054 0.697 7.963 0.343 >50 231965.c01 D >50 >50 >10 >10 1.285 47.97 1.438 >50 231966.c02 D >50 >50 0.191 3.130 0.075 0.257 0.007 0.031 3817.v2.c59 CD 3.148 14.880 1.181 5.977 0.020 0.062 0.008 0.060 6480.v4.c25 CD 0.009 0.041 8.079 >10 >50 >50 >50 >50 6952.v1.c20 CD 0.037 0.138 0.343 1.907 >50 >50 21.87 >50 6811.v7.c18 CD 0.002 0.010 7.535 >10 >50 >50 >50 >50 89-F1_2_25 CD >50 >50 1.112 10.000 0.668 10.1 47.61 >50 3301.v1.c24 AC 0.013 0.042 4.628 >10 0.216 0.756 0.020 0.087 6041.v3.c23 AC >50 >50 2.333 9.766 0.244 3.639 0.048 0.256 6540.v4.c1 AC >50 >50 1.899 10.000 0.070 0.253 0.035 0.420 6545.v4.c1 AC >50 >50 1.778 8.229 0.099 0.351 0.044 1.561 0815.v3.c3 ACD 0.03 0.138 0.332 2.868 >50 >50 >50 >50 3103.v3.c10 ACD 0.037 0.101 >10 >10 28.004 >50 25.39 >50 MuLV Neg. Control *(T/F): Transmitted/Founder Virus
TABLE-US-00025 TABLE 24 Neutralization assay data of 3BNC117, 45-46, 8ANC195, VRC07 mAbs against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 24 3BNC117 45-46 8ANC195 VRC07 Virus ID Clade* IC50 IC80 IC50 IC80 IC50 IC80 IC50 IC80 6535.3 B 0.55 2.44 0.14 0.28 0.20 0.91 0.169 0.517 QH0692.42 B 0.13 0.49 0.55 1.56 2.71 17.08 0.500 1.861 SC422661.8 B 0.02 0.08 0.01 0.07 0.29 4.67 0.054 0.175 PVO.4 B <0.09 0.19 0.17 0.47 0.52 1.87 0.063 0.226 TRO.11 B <0.09 <0.09 1.90 9.56 0.18 0.89 0.132 0.456 AC10.0.29 B 13.84 >50 0.42 1.49 0.88 7.00 0.282 0.986 RHPA4259.7 B <0.09 <0.09 <0.05 <0.05 0.34 1.56 0.017 0.048 THRO4156.18 B 1.76 10.14 1.59 6.02 >50 >50 2.404 12.200 REJO4541.67 B 0.01 0.05 0.003 0.010 0.08 0.68 0.011 0.037 TRJO4551.58 B 0.05 0.19 0.01 0.05 0.19 1.17 0.032 0.125 WITO4160.33 B 0.01 0.04 0.01 0.08 >50 >50 0.042 0.193 CAAN5342.A2 B 0.42 1.51 0.11 0.33 >50 >50 0.307 1.053 WEAU_d15_410_787 B (T/F) 0.05 0.19 0.01 0.03 >50 >50 0.034 0.115 1006_11_C3_1601 B (T/F) 0.03 0.10 0.05 0.31 0.43 1.98 0.015 0.046 1054_07_TC4_1499 B (T/F) 0.07 0.49 0.07 0.61 1.02 6.75 0.239 1.459 1056_10_TA11_1826 B (T/F) 0.30 1.82 0.12 0.59 >50 >50 0.239 1.110 1012_11_TC21_3257 B (T/F) 0.02 0.07 0.01 0.04 >50 >50 0.014 0.045 6240_08_TA5_4622 B (T/F) 0.33 1.17 0.44 1.55 >50 >50 0.448 1.540 6244_13_B5_4576 B (T/F) 0.04 0.15 0.07 0.26 >50 >50 0.065 0.230 62357_14_D3_4589 B (T/F) 0.06 0.26 0.05 0.22 >50 >50 0.075 0.256 SC05_8C11_2344 B (T/F) 0.15 0.51 0.14 0.38 0.47 3.06 0.133 0.401 Du156.12 C 0.02 0.08 0.01 0.05 0.22 1.56 0.041 0.149 Du172.17 C 1.19 8.90 >30 >30 >30 >30 0.796 3.661 Du422.1 C >50 >50 >50 >50 >50 >50 >50 >50 ZM197M.PB7 C 0.22 1.03 0.14 0.64 23.45 >50 0.307 1.275 ZM214M.PL15 C 0.06 0.52 0.05 0.39 0.91 12.30 0.198 1.417 ZM233M.PB6 C 0.13 0.85 1.86 16.52 7.39 43.45 0.152 0.625 ZM249M.PL1 C 0.03 0.11 0.02 0.06 >50 >50 0.041 0.145 ZM53M.PB12 C 0.21 0.85 0.65 0.65 >30 >30 0.275 1.294 ZM109F.PB4 C 0.14 0.88 0.22 0.22 >30 >30 0.112 0.389 ZM135M.PL10a C 0.03 0.13 0.36 3.11 >50 >50 0.104 0.383 CAP45.2.00.G3 C 3.88 >50 >50 >50 28.30 >50 0.355 3.087 CAP210.2.00.E8 C 17.22 >50 >50 >50 >50 >50 >50 >50 HIV-001428-2.42 C 0.003 0.02 <0.001 0.01 >50 >50 0.004 0.016 HIV-0013095-2.11 C 0.33 2.53 0.01 0.03 0.32 1.85 0.017 0.057 HIV-16055-2.3 C 5.60 >50 0.02 0.06 15.30 >50 0.025 0.135 HIV-16845-2.22 C 27.46 >50 2.05 12.10 3.30 20.06 1.207 6.154 Ce1086_B2 C (T/F) 0.09 0.31 0.04 0.28 4.42 19.11 0.128 0.840 Ce0393_C3 C (T/F) 0.20 0.67 0.32 1.07 6.77 >50 0.182 0.634 Ce1176_A3 C (T/F) 0.22 0.75 1.29 5.55 4.45 35.17 0.520 2.600 Ce2010_F5 C (T/F) 0.05 0.21 0.13 0.43 >50 >50 0.131 0.480 Ce0682_E4 C (T/F) 0.03 0.15 0.07 0.25 0.20 1.29 0.039 0.239 Ce1172_H1 C (T/F) >50 >50 >50 >50 42.30 >50 >50 >50 Ce2060_G9 C (T/F) 0.24 0.81 0.14 0.48 6.20 >50 0.085 0.422 Ce703010054_2A2 C (T/F) 0.37 1.67 0.22 0.97 13.62 >50 0.190 0.912 BF1266.431a C (T/F) 0.03 0.09 0.01 0.03 >50 >50 0.032 0.117 246F C1G C (T/F) 19.32 >50 >50 >50 12.81 >50 0.286 1.041 249M B10 C (T/F) 0.10 0.40 0.04 0.17 >50 >50 0.052 0.204 ZM247v1(Rev-) C (T/F) >50 >50 1.64 >50 3.41 25.12 0.041 0.189 7030102001E5(Rev-) C (T/F) 0.29 1.44 0.14 0.63 0.86 8.32 0.205 0.911 1394C9G1(Rev-) C (T/F) >50 >50 0.09 0.38 >50 >50 0.112 0.368 Ce704809221_1B3 C (T/F) 0.08 0.31 0.15 0.88 0.48 3.52 0.273 1.137 CNE19 BC 0.02 0.08 0.07 0.37 0.29 1.73 0.092 0.232 CNE20 BC >50 >50 8.71 47.01 0.50 3.19 0.060 0.129 CNE21 BC 45.38 >50 7.53 >50 0.27 1.52 0.050 0.222 CNE17 BC 5.76 42.98 0.15 0.55 2.59 11.07 0.136 0.651 CNE30 BC 0.26 0.85 0.34 1.48 7.52 40.45 0.326 1.084 CNE52 BC 0.02 0.06 0.03 0.12 5.59 43.09 0.042 0.142 CNE53 BC 0.08 0.99 0.01 0.04 >50 >50 0.031 0.086 CNE58 BC 0.25 2.03 0.25 1.49 >50 >50 0.110 0.298 MS208.A1 A 0.01 0.09 0.09 0.60 >50 >50 0.120 0.425 Q23.17 A 0.005 0.020 0.14 0.60 2.55 2.55 0.081 0.236 Q461.e2 A 0.03 0.09 0.06 0.32 0.63 2.51 0.282 1.025 Q769.d22 A 0.01 0.04 0.01 0.04 0.26 1.20 0.018 0.087 Q259.d2.17 A 0.01 0.05 0.01 0.06 >50 >50 0.024 0.100 Q842.d12 A <0.01 0.01 0.03 0.03 >30 >30 0.015 0.051 3415.v1.c1 A 0.17 0.47 0.07 0.07 17.74 17.74 0.036 0.152 3365.v2.c2 A 0.03 0.10 0.11 0.11 >30 >30 0.038 0.109 0260.v5.c36 A NT NT NT NT NT NT 0.448 1.619 191955_A11 A (T/F) >50 >50 0.11 0.43 >50 >50 0.978 4.824 191084 B7-19 A (T/F) 0.07 0.23 0.02 0.07 >50 >50 0.061 0.209 9004SS_A3_4 A (T/F) 0.07 0.18 0.18 0.65 0.17 0.81 0.218 0.750 T257-31 CRF02_AG 0.06 0.37 0.23 1.11 0.32 3.89 0.421 3.172 928-28 CRF02_AG 0.15 0.55 0.10 0.39 >50 >50 0.278 0.957 263-8 CRF02_AG 0.005 0.040 0.05 0.10 >50 >50 0.103 0.282 T250-4 CRF02_AG >15 >15 >30 >30 >50 >50 >50 >50 T251-18 CRF02_AG 0.26 0.82 5.26 5.26 1.51 1.51 1.322 6.491 T278-50 CRF02_AG >15 >15 >30 >30 >50 >50 >50 >50 T255-34 CRF02_AG 0.02 0.14 0.23 2.38 0.54 8.09 0.227 0.987 211-9 CRF02_AG 0.35 1.28 >50 >50 1.34 8.43 5.919 37.988 235-47 CRF02_AG 0.03 0.12 0.30 1.49 >50 >50 0.024 0.075 620345.c01 CRF01_AE >15 >15 >30 >30 >50 >50 >50 >50 CNE8 CRF01_AE NT NT NT NT NT NT 0.450 1.563 C1080.c03 CRF01_AE 0.25 1.73 1.79 18.04 0.17 1.12 2.032 8.251 R2184.c04 CRF01_AE 0.03 0.10 0.03 0.12 0.09 12.98 0.020 0.104 R1166.c01 CRF01_AE 0.17 0.64 1.68 1.68 4.83 4.83 0.810 3.167 R3265.c06 CRF01_AE 0.48 13.04 0.04 0.44 50.00 >50 NT NT C2101.c01 CRF01_AE 0.05 0.31 12.78 >50 0.48 3.54 0.238 1.404 C3347.c11 CRF01_AE 0.03 0.13 0.03 0.18 >50 >50 0.060 0.380 C4118.c09 CRF01_AE 0.09 0.52 0.14 0.91 0.20 0.71 0.121 0.444 CNE5 CRF01_AE 0.28 1.28 0.09 0.41 2.12 9.67 0.138 0.524 BJOX009000.02.4 CRF01_AE 0.40 1.79 0.26 1.16 0.64 3.09 0.806 2.899 BJOX015000.11.5 CRF01_AE 0.05 0.32 0.06 0.38 0.97 4.30 0.054 0.479 (T/F) BJOX010000.06.2 CRF01_AE 1.58 10.45 0.87 6.37 0.85 5.84 1.061 10.035 (T/F) BJOX025000.01.1 CRF01_AE 0.05 0.20 0.18 10.00 4.96 >50 0.279 2.295 (T/F) BJOX028000.10.3 CRF01_AE 0.01 0.05 0.07 >50 0.45 2.83 0.011 0.069 (T/F) X1193_c1 G 0.06 0.25 0.04 0.16 0.27 1.66 0.037 0.249 P0402_c2_11 G 0.06 0.38 0.06 0.26 0.62 6.35 0.085 0.422 X1254_c3 G 0.08 0.27 0.08 0.08 6.95 6.95 0.041 0.200 X2088_c9 G >50 >50 >50 >50 >50 >50 >50 >50 X2131_C1_B5 G 0.43 1.96 0.20 0.90 3.47 >50 0.219 1.033 P1981_C5_3 G 0.74 3.62 0.05 0.22 0.14 1.15 0.213 0.609 X1632_S2_B10 G 15.69 >50 0.07 >50 0.47 4.11 0.023 0.107 3016.v5.c45 D 1.38 >30 >30 >30 0.87 >50 0.095 0.392 A07412M1.vrc12 D 0.02 0.10 0.03 0.13 2.25 7.49 0.110 0.353 231965.c01 D 0.05 0.22 0.10 0.10 2.36 2.36 0.085 0.263 231966.c02 D 0.29 2.37 1.11 11.36 0.54 2.98 0.105 0.471 3817.v2.c59 CD 0.15 0.52 >50 >50 1.00 5.30 1.900 19.842 6480.v4.c25 CD 0.01 0.04 0.02 0.08 0.16 1.13 0.032 0.134 6952.v1.c20 CD 0.15 0.75 0.02 0.07 1.38 7.22 0.064 0.181 6811.v7.c18 CD 0.03 0.17 0.03 0.13 5.34 27.63 0.093 0.435 89-F1_2_25 CD >50 >50 >50 >50 >50 >50 >50 >50 3301.v1.c24 AC 0.01 0.05 0.01 0.03 >50 >50 0.027 0.073 6041.v3.c23 AC 0.01 0.07 0.01 0.04 >50 >50 0.018 0.058 6540.v4.c1 AC >50 >50 >50 >50 >50 >50 >50 >50 6545.v4.c1 AC >50 >50 >50 >50 26.94 >50 >50 >50 0815.v3.c3 ACD 0.01 0.02 0.01 0.03 1.94 1.94 0.015 0.051 3103.v3.c10 ACD 0.22 0.85 1.66 6.57 48.07 >50 0.675 1.951 MuLV Neg. Control *(T/F): Transmitted/Founder Virus
TABLE-US-00026 TABLE 25 Neutralization assay data of VRC01 mAb against extended multiclade virus panel. mAbs tested at primary concentration of 50 ug/ml and titrated 5-fold 7x (duplicate wells). Titer in TZM.bl cells (ug/ml) TABLE 25 VRC01 Virus ID Clade* IC50 IC80 6535.3 B 0.54 2.7 QH0692.42 B 1.50 4.8 SC422661.8 B 0.08 0.265 PVO.4 B 0.22 1.2 TRO.11 B 0.21 0.83 AC10.0.29 B 2.20 6.5 RHPA4259.7 B 0.06 0.185 THRO4156.18 B 2.30 23 REJO4541.67 B 0.06 0.25 TRJO4551.58 B 0.08 0.21 WITO4160.33 B 0.15 0.41 CAAN5342.A2 B 0.82 2.8 WEAU_d15_410_787 B (T/F) 0.12 0.26 1006_11_C3_1601 B (T/F) 0.15 0.39 1054_07_TC4_1499 B (T/F) 0.71 2.85 1056_10_TA11_1826 B (T/F) 0.92 3.28 1012_11_TC21_3257 B (T/F) 0.12 0.32 6240_08_TA5_4622 B (T/F) 0.61 1.82 6244_13_B5_4576 B (T/F) 0.21 0.53 62357_14_D3_4589 B (T/F) 0.96 4.68 SC05_8C11_2344 B (T/F) 0.64 1.91 Du156.12 C 0.09 0.193 Du172.17 C >50 >50 Du422.1 C >50 >50 ZM197M.PB7 C 0.36 1.6 ZM214M.PL15 C 0.44 2.6 ZM233M.PB6 C 2.00 9.3 ZM249M.PL1 C 0.05 0.23 ZM53M.PB12 C 1.30 4 ZM109F.PB4 C 0.13 0.75 ZM135M.PL10a C 0.35 2.7 CAP45.2.00.G3 C 2.30 >50 CAP210.2.00.E8 C >50 >50 HIV-001428-2.42 C 0.02 0.06 HIV-0013095-2.11 C 0.11 0.33 HIV-16055-2.3 C 0.08 0.26 HIV-16845-2.22 C 2.80 12.69 Ce1086_B2 C (T/F) 0.42 1.44 Ce0393_C3 C (T/F) 0.62 2.37 Ce1176_A3 C (T/F) 2.25 8.10 Ce2010_F5 C (T/F) 0.32 1.07 Ce0682_E4 C (T/F) 0.08 0.48 Ce1172_H1 C (T/F) >10 >10 Ce2060_G9 C (T/F) 0.29 1.48 Ce703010054_2A2 C (T/F) 0.53 1.84 BF1266.431a C (T/F) 0.07 0.23 246F C1G C (T/F) >10 >10 249M B10 C (T/F) 0.13 0.46 ZM247v1(Rev-) C (T/F) 0.35 1.29 7030102001E5(Rev-) C (T/F) 0.54 2.08 1394C9G1 (Rev-) C (T/F) 0.36 1.28 Ce704809221_1B3 C (T/F) 0.55 2.05 CNE19 BC NT NT CNE20 BC NT NT CNE21 BC NT NT CNE17 BC NT NT CNE30 BC NT NT CNE52 BC NT NT CNE53 BC NT NT CNE58 BC NT NT MS208.A1 A 0.10 0.46 Q23.17 A 0.09 0.261 Q461.e2 A 0.49 1.6 Q769.d22 A 0.08 0.29 Q259.d2.17 A 0.17 0.54 Q842.d12 A 0.03 0.096 3415.v1.c1 A 0.06 0.15 3365.v2.c2 A 0.06 0.17 0260.v5.c36 A NT NT 191955_A11 A (T/F) NT NT 191084 B7-19 A (T/F) NT NT 9004SS_A3_4 A (T/F) NT NT T257-31 CRF02_AG 2.80 8.66 928-28 CRF02_AG 0.41 1.7 263-8 CRF02_AG 0.20 0.55 T250-4 CRF02_AG >50 >50 T251-18 CRF02_AG 2.50 11.17 T278-50 CRF02_AG >50 >50 T255-34 CRF02_AG 0.70 2.7 211-9 CRF02_AG 14.30 >50 235-47 CRF02_AG 0.04 0.17 620345.c01 CRF01_AE >50 >50 CNE8 CRF01_AE NT NT C1080.c03 CRF01_AE 3.40 14.37 R2184.c04 CRF01_AE 0.08 0.32 R1166.c01 CRF01_AE 1.70 4.58 R3265.c06 CRF01_AE 0.45 1.88 C2101.c01 CRF01_AE 0.36 1.17 C3347.c11 CRF01_AE 0.17 0.58 C4118.c09 CRF01_AE NT NT CNE5 CRF01_AE 0.37 1.1 BJOX009000.02.4 CRF01_AE NT NT BJOX015000.11.5 CRF01_AE NT NT (T/F) BJOX010000.06.2 CRF01_AE NT NT (T/F) BJOX025000.01.1 CRF01_AE NT NT (T/F) BJOX028000.10.3 CRF01_AE NT NT (T/F) X1193_c1 G 0.11 0.32 P0402_c2_11 G 0.21 0.59 X1254_c3 G 0.07 0.19 X2088_c9 G >50 >50 X2131_C1_B5 G 0.51 1.54 P1981_C5_3 G 0.46 1.26 X1632_S2_B10 G 0.12 0.74 3016.v5.c45 D 0.16 0.42 A07412M1.vrc12 D NT NT 231965.c01 D 0.34 1.2 231966.c02 D NT NT 3817.v2.c59 CD >50 >50 6480.v4.c25 CD 0.04 0.09 6952.v1.c20 CD 0.04 0.12 6811.v7.c18 CD 0.09 0.26 89-F1_2_25 CD NT NT 3301.v1.c24 AC 0.14 0.32 6041.v3.c23 AC 0.02 0.08 6540.v4.c1 AC >50 >50 6545.v4.c1 AC >50 >50 0815.v3.c3 ACD 0.06 0.13 3103.v3.c10 ACD 0.93 2.49 MuLV Neg. Control *(T/F): Transmitted/Founder Virus
[1099] Throughout this disclosure, various publications, patents and published patent specifications are referenced by an identifying citation. The disclosures of these publications, patents and published patent specifications are hereby incorporated by reference into the present disclosure to more fully describe the state of the art to which this invention pertains.
[1100] While the present teachings are described in conjunction with various embodiments, it is not intended that the present teachings be limited to such embodiments. On the contrary, the present teachings encompass various alternatives, modifications, and equivalents, as will be appreciated by those of skill in the art.
Sequence CWU 0 SQTB SEQUENCE LISTING The patent application
contains a lengthy "Sequence Listing" section. A copy of the
"Sequence Listing" is available in electronic form from the USPTO
web site
(http://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200172601A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
0 SQTB SEQUENCE LISTING The patent application contains a lengthy
"Sequence Listing" section. A copy of the "Sequence Listing" is
available in electronic form from the USPTO web site
(http://seqdata.uspto.gov/?pageRequest=docDetail&DocID=US20200172601A1).
An electronic copy of the "Sequence Listing" will also be available
from the USPTO upon request and payment of the fee set forth in 37
CFR 1.19(b)(3).
References































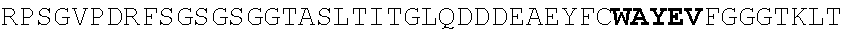








































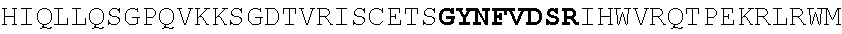


























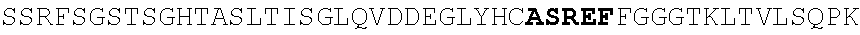













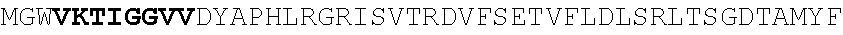





















































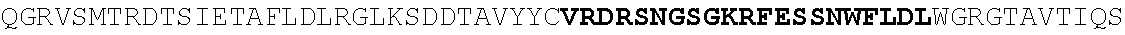









































































































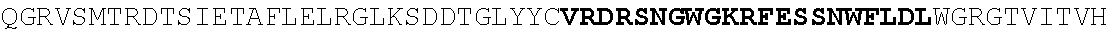








































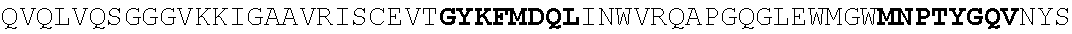












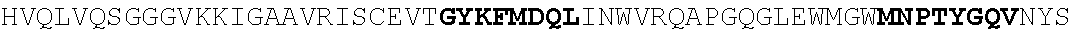
























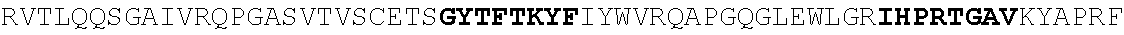










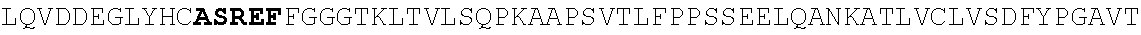





















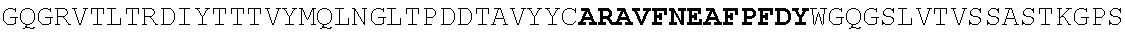

























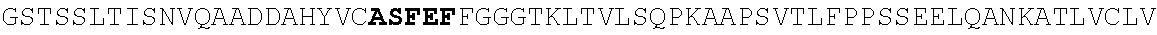




D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

D00032

D00033

D00034

D00035

D00036

D00037

D00038

D00039

D00040

D00041

D00042

D00043

D00044

D00045

D00046

D00047

D00048

D00049

P00899

XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.