Methods And Organisms For The Growth-coupled Production Of 1,4-butanediol
Burgard; Anthony P. ; et al.
U.S. patent application number 16/428627 was filed with the patent office on 2020-03-26 for methods and organisms for the growth-coupled production of 1,4-butanediol. The applicant listed for this patent is Genomatica, Inc.. Invention is credited to Anthony P. Burgard, Mark J. Burk, Stephen J. Van Dien.
| Application Number | 20200095616 16/428627 |
| Document ID | / |
| Family ID | 39768484 |
| Filed Date | 2020-03-26 |






View All Diagrams
| United States Patent Application | 20200095616 |
| Kind Code | A1 |
| Burgard; Anthony P. ; et al. | March 26, 2020 |
METHODS AND ORGANISMS FOR THE GROWTH-COUPLED PRODUCTION OF 1,4-BUTANEDIOL
Abstract
The invention provides a non-naturally occurring microorganism comprising one or more gene disruptions, the one or more gene disruptions occurring in genes encoding an enzyme obligatory to coupling 1,4-butanediol production to growth of the microorganism when the gene disruption reduces an activity of the enzyme, whereby theone or more gene disruptions confers stable growth-coupled production of 1,4-butanediol onto the non-naturally occurring microorganism. The microorganism can further comprise a gene encoding an enzyme in a 1,4-butanediol (BDO) biosynthetic pathway. The invention additionally relates to methods of using microorganisms to produce BDO.
| Inventors: | Burgard; Anthony P.; (Elizabeth, PA) ; Van Dien; Stephen J.; (Encinitas, CA) ; Burk; Mark J.; (San Diego, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 39768484 | ||||||||||
| Appl. No.: | 16/428627 | ||||||||||
| Filed: | May 31, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15424723 | Feb 3, 2017 | |||
| 16428627 | ||||
| 13914422 | Jun 10, 2013 | |||
| 15424723 | ||||
| 13065303 | Mar 18, 2011 | 8470582 | ||
| 13914422 | ||||
| 11891602 | Aug 10, 2007 | 7947483 | ||
| 13065303 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | B01D 3/002 20130101; C12N 9/0006 20130101; C12P 7/18 20130101 |
| International Class: | C12P 7/18 20060101 C12P007/18; B01D 3/00 20060101 B01D003/00; C12N 9/04 20060101 C12N009/04 |
Claims
1. A non-naturally occurring microorganism comprising one or more gene disruptions, said one or more gene disruptions occurring in genes encoding an enzyme obligatory to coupling 1,4-butanediol production to growth of said microorganism when said gene disruption reduces an activity of said enzyme, whereby said one or more gene disruptions confers stable growth-coupled production of 1,4-butanediol onto said non-naturally occurring microorganism.
2. The non-naturally occurring microorganism of claim 1, wherein said one or more gene disruptions comprise a metabolic modification listed in Table 6 or 7.
3. The non-naturally occurring microorganism of claim 1, wherein said one or more gene disruptions comprise a deletion of said one or more genes.
4. The non-naturally occurring microorganism of claim 1, wherein said non-naturally occurring microorganism is selected from the group of microorganisms having a metabolic modification listed in Table 6 or 7.
5. A non-naturally occurring microorganism comprising a set of metabolic modifications obligatory to coupling 1,4-butanediol production to growth of said microorganism, said set of metabolic modifications comprising disruption of one or more genes, or an ortholog thereof, wherein said set of metabolic modifications comprises disruption of adhE and ldhA, wherein said microorganism exhibits stable growth-coupled production of 1,4-butanediol.
6. The non-naturally occurring microorganism of claim 5, wherein said set of metabolic modifications further comprise disruption of mdh; adhE, ldhA, pflAB, mdh, and aspA; or adhE, ldhA, pflAB, mdh, and aspA.
7. The non-naturally occurring microorganism of claim 6, wherein said set of metabolic modifications further comprise disruption of one or more genes selected from the set of genes comprising mqo, aspA, sfcA, maeB, pntAB, and gdhA; or pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW, prpC, and gsk.
8. The non-naturally occurring microorganism of claim 7, wherein said set of metabolic modifications comprise disruption of sfcA and maeB; or pykA, pykF, dhaKLM, deoC, edd, yiaE and ycdW.
9. The non-naturally occurring microorganism of claim 8, wherein said set of metabolic modifications further comprise disruption of prpC and gsk.
10. The non-naturally occurring microorganism of claim 6, wherein said set of metabolic modifications further comprise disruption of pflAB.
11. The non-naturally occurring microorganism of claim 7, wherein said set of metabolic modifications further comprise disruption of pflAB.
12. The non-naturally occurring microorganism of claim 8, wherein said set of metabolic modifications further comprise disruption of pflAB.
13. The non-naturally occurring microorganism of claim 9, wherein said set of metabolic modifications further comprise disruption of pflAB.
14. The non-naturally occurring microorganism of claim 5, wherein said microorganism further comprises a 1,4-butanediol (BDO) biosynthetic pathway, said pathway comprising at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, 4-hydroxybutyrate:CoA transferase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase, CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or alcohol dehydrogenase, wherein said exogenous nucleic acid is expressed in sufficient amounts to produce 1,4-butanediol (BDO).
15. A method of producing a non-naturally occurring microorganism having stable growth-coupled production of 1,4-butanediol, comprising: (a) identifying in silico a set of metabolic modifications requiring 1,4-butanediol production during exponential growth, and (b) genetically modifying a microorganism to contain said set of metabolic modifications requiring 1,4-butanediol production.
16-23. (canceled)
24. A method of producing 1,4-butanediol coupled to the growth of a microorganism, comprising: (a) culturing under exponential growth phase in a sufficient amount of nutrients and media the non-naturally occurring microorganism of claim 8; and (b) isolating 1,4-butanediol produced from said non-naturally occurring microorganism.
25-31. (canceled)
32. The method of claim 24, wherein said non-naturally occurring microorganism further comprises a 1,4-butanediol (BDO) biosynthetic pathway, said pathway comprising at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, 4-hydroxybutyrate:CoA transferase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase, CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or alcohol dehydrogenase, wherein said exogenous nucleic acid is expressed in sufficient amounts to produce 1,4-butanediol (BDO).
Description
[0001] This application is a continuation of application Ser. No. 15/424,723, filed Feb. 3, 2017, which is a continuation of application Ser. No. 13/914,422, filed Jun. 10, 2013, which is a continuation of application Ser. No. 13/065,303, filed Mar. 18, 2011, now U.S. Pat. No. 8,470,582, which is is a continuation of application Ser. No. 11/891,602, filed Aug. 10, 2007, now U.S. Pat. No. 7,947,483, the entire contents of each of which are incorporated herein by reference.
BACKGROUND OF THE INVENTION
[0002] The present invention relates generally to in silico design of organisms, and more specifically to organisms having 1,4-butanediol biosynthetic capability.
[0003] 1,4-Butanediol (BDO) is a four carbon dialcohol that currently is manufactured exclusively through various petrochemical routes. BDO is part of a large volume family of solvents and polymer intermediates that includes gamma-butyrolactone (GBL), tetrahydrofuran (THF), pyrrolidone, N-methylpyrrolidone (NMP), and N-vinyl-pyrrolidone. The overall market opportunity for this family exceeds $4.0 B.
[0004] Approximately 2.5B lb BDO is produced globally per year with 4-5% annual growth and a recent selling price ranging from $1.00-1.20/lb. The demand for BDO stems largely from its use as an intermediate for polybutylene terephthalate (PBT) plastic resins, polyurethane thermoplastics and co-polyester ethers. BDO also serves as a primary precursor to THF, which is employed as an intermediate for poly(tetramethylene glycol) PTMEG copolymers required for lycra and spandex production. Approximately 0.7 B lb of THF is produced globally per year with an annual growth rate over 6%. A significant percentage of growth (>30%) for both BDO and THF is occurring in Asia (China and India). GBL currently is a smaller volume (0.4 B lb/year) product which has numerous applications as a solvent, as an additive for inks, paints, and dyes, as well as the primary precursor to pyrrolidone derivatives such as NMP.
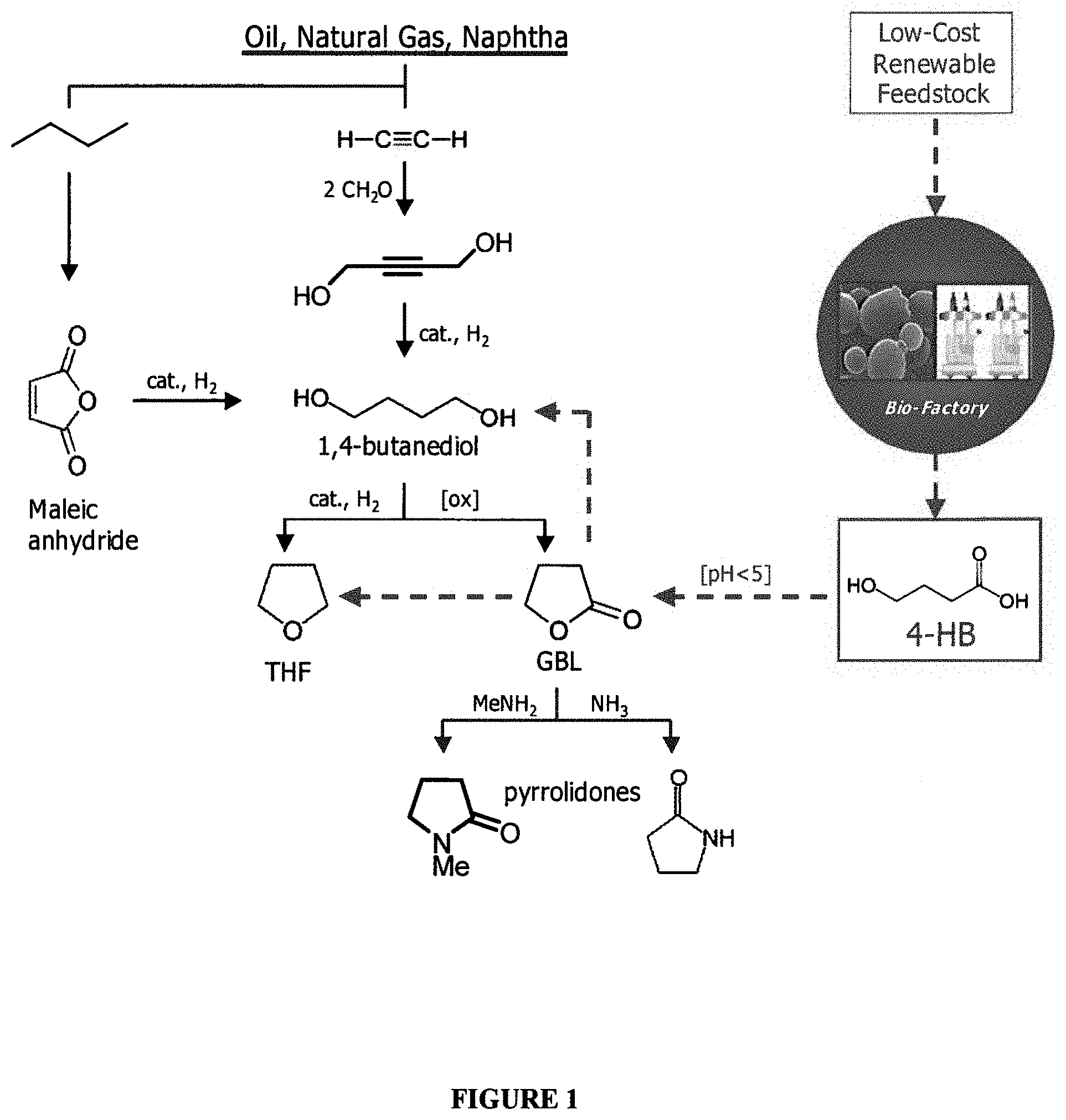
[0005] Conventional processes for the synthesis of BDO use petrochemical feedstocks for their starting materials. For example, acetylene is reacted with 2 molecules of formaldehyde in the Reppe synthesis reaction (Kroschwitz and Grant, Encyclopedia of Chem. Tech., John Wiley and Sons, Inc., New York (1999)), followed by catalytic hydrogenation to form 1,4-butanediol. It has been estimated that 90% of the acetylene produced in the U.S. is consumed for butanediol production. Alternatively, it can be formed by esterification and catalytic hydrogenation of maleic anhydride, which is derived from butane. Downstream, butanediol can be further transformed; for example, by oxidation to .gamma.-butyrolactone, which can be further converted to pyrrolidone and N-methyl-pyrrolidone, or hydrogenolysis to tetrahydrofuran (FIG. 1). These compounds have varied uses as polymer intermediates, solvents, and additives, and have a combined market of nearly 2 billion lb/year.
[0006] The conventional hydrocarbon feedstock-based approach utilizes methane to produce formaldehyde. Thus, a large percentage of the commercial production of BDO relies on methane as a starting material. The production of acetylene also relies on petroleum-based starting material (see FIG. 1). Therefore, the costs of BDO production fluctuate with the price of petroleum and natural gas.
[0007] It is desirable to develop a method for production of these chemicals by alternative means that not only substitute renewable for petroleum-based feedstocks, and also use less energy- and capital-intensive processes. The Department of Energy has proposed 1,4-diacids, and particularly succinic acid, as key biologically-produced intermediates for the manufacture of the butanediol family of products (DOE Report, "Top Value-Added Chemicals from Biomass", 2004). However, succinic acid is costly to isolate and purify and requires high temperatures and pressures for catalytic reduction to butanediol.
[0008] Thus, there exists a need for alternative means for effectively producing commercial quantities of 1,4-butanediol and its chemical precursors. The present invention satisfies this need and provides related advantages as well.
SUMMARY OF INVENTION
[0009] The invention provides a non-naturally occurring microorganism comprising one or more gene disruptions, the one or more gene disruptions occurring in genes encoding an enzyme obligatory to coupling 1,4-butanediol production to growth of the microorganism when the gene disruption reduces an activity of the enzyme, whereby theone or more gene disruptions confers stable growth-coupled production of 1,4-butanediol onto the non-naturally occurring microorganism. The microorganism can further comprise a gene encoding an enzyme in a 1,4-butanediol (BDO) biosynthetic pathway. The invention additionally relates to methods of using microorganisms to produce BDO.
BRIEF DESCRIPTION OF THE DRAWINGS
[0010] The patent or application file contains at least one drawing executed in color. Copies of this patent or patent application publication with color drawings will be provided by the Office upon request and payment of the necessary fee.
[0011] FIG. 1 is a schematic diagram showing an entry point of 4-hydroxybutanoic acid (4-HB) into the product pipeline of the 1,4-butanediol (BDO) family of chemicals, and comparison with chemical synthesis routes from petrochemical feedstocks. Solid black arrows show chemical synthesis routes; dashed blue arrows show a biosynthetic route to 4-HB and subsequent conversion steps to BDO family chemicals.
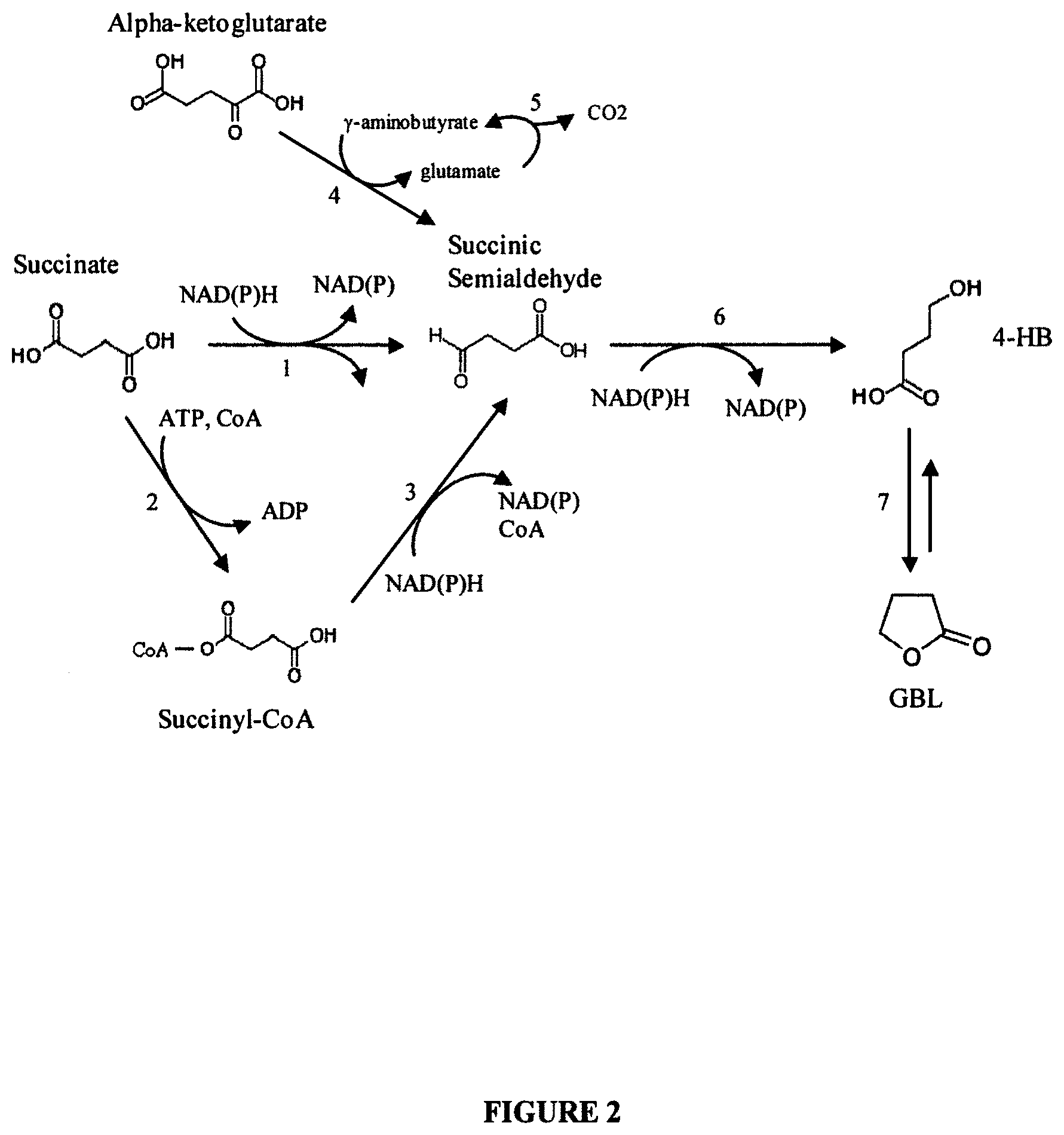
[0012] FIG. 2 is a schematic diagram showing biochemical pathways to 4-hydroxybutyurate (4-HB) and .gamma.-butyrolactone (GBL) production. Enzymes catalyzing the 4-HB biosynthetic reactions are: (1) CoA-independent succinic semialdehyde dehydrogenase; (2) succinyl-CoA synthetase; (3) CoA-dependent succinic semialdehyde dehydrogenase; (4) glutamate:succinic semialdehyde transaminase; (5) glutamate decarboxylase; (6) 4-hydroxybutanoate dehydrogenase. Conversion (7) corresponds to a spontaneous, non-enzymatic reaction which converts 4-HB to GBL.
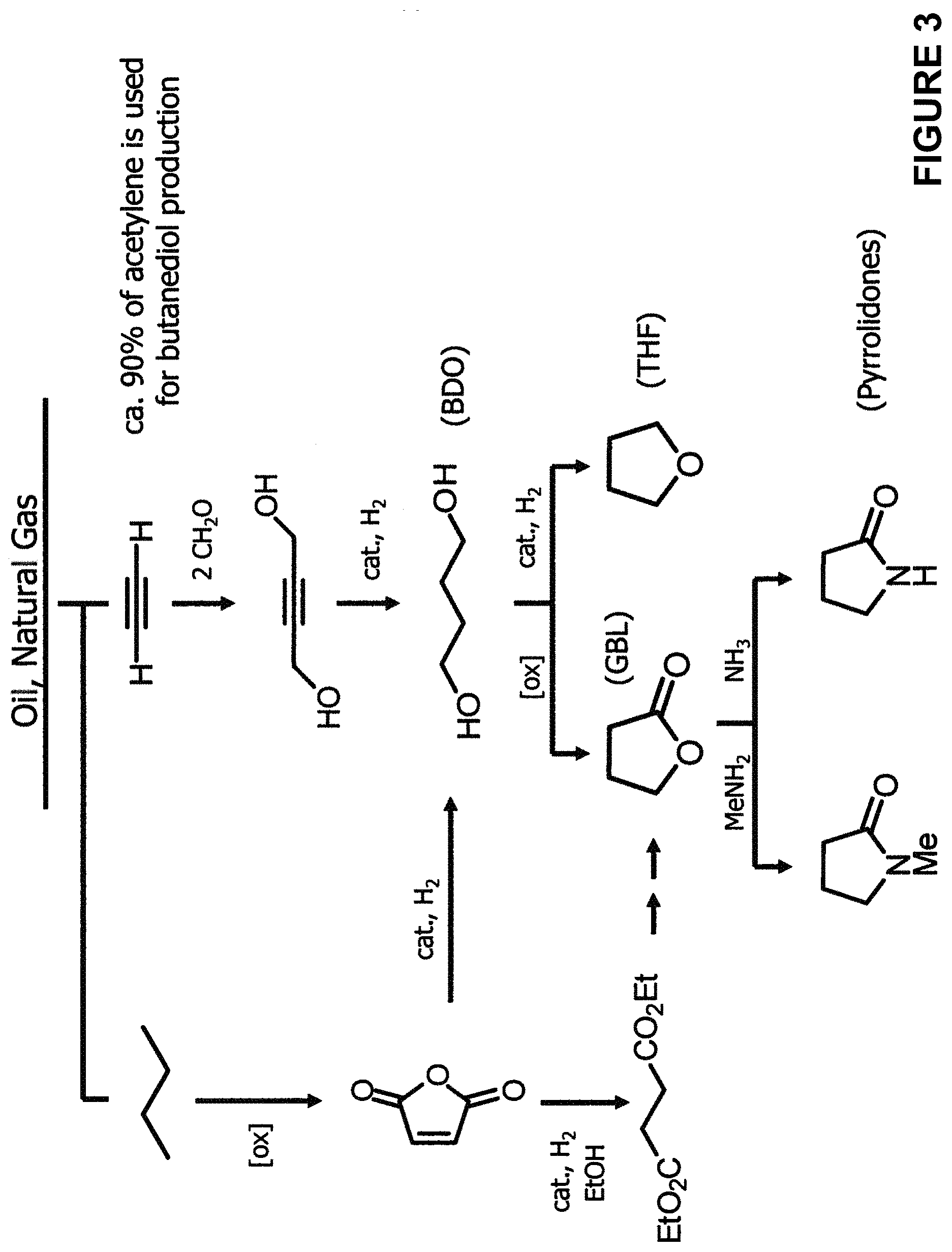
[0013] FIG. 3 is a schematic diagram showing the chemical synthesis of 1,4-butanediol (BDO) and downstream products .gamma.-butyrolactone (GBL), tetrahydrofuran (THF) and several pyrrolidones.
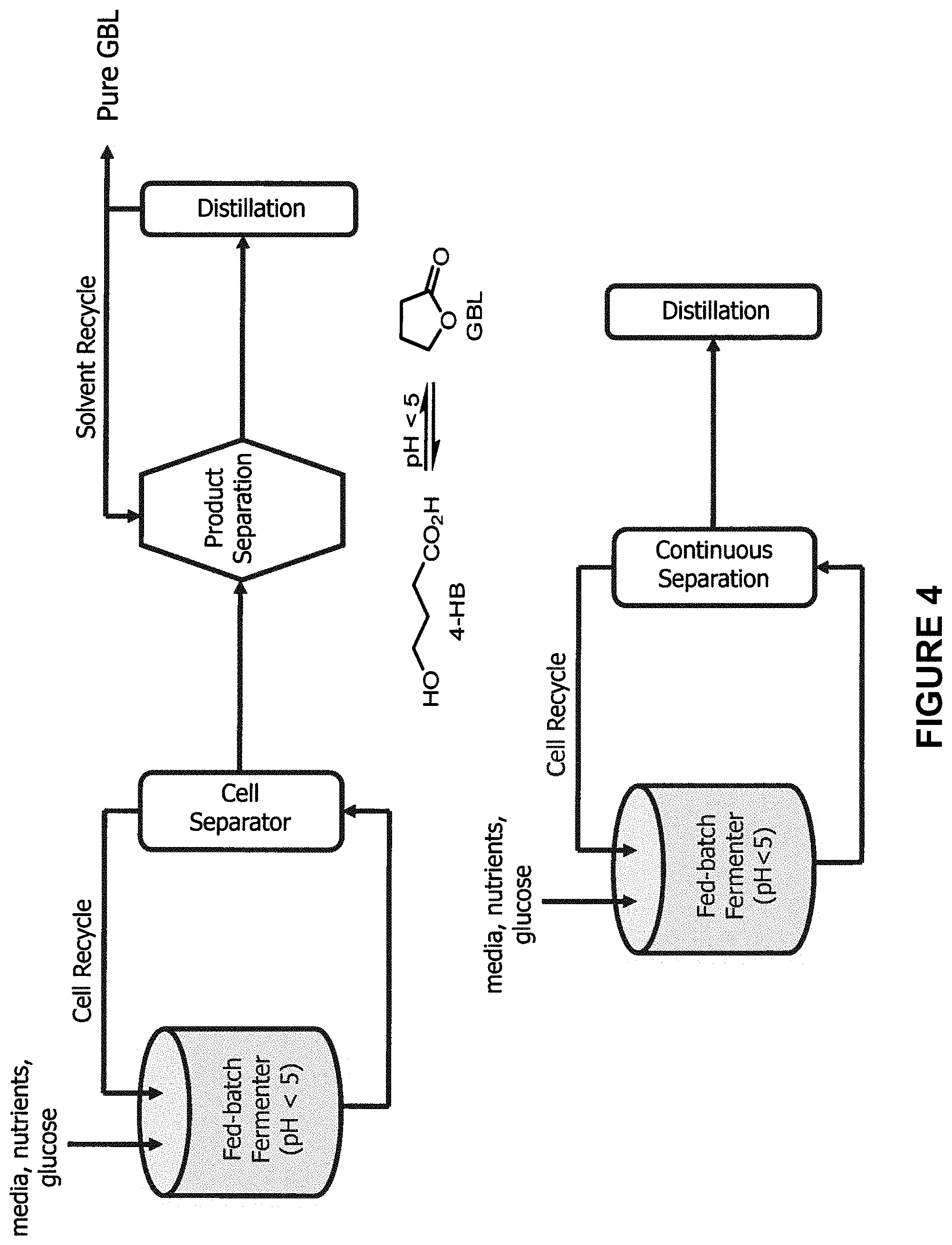
[0014] FIG. 4 is a schematic process flow diagram of bioprocesses for the production of .gamma.-butyrolactone. Panel (a) illustrates fed-batch fermentation with batch separation and panel (b) illustrates fed-batch fermentation with continuous separation.
[0015] FIG. 5 shows the hypothetical production envelopes of an OptKnock-designed strain contrasted against a typical non-growth-coupled production strain. Note that the potential evolutionary trajectories of the OptKnock strain are fundamentally different in that they lead to a high producing phenotype.
[0016] FIG. 6 shows biochemical pathways to 1,4-butanediol. 1) CoA-independent succinic semialdehyde dehydrogenase; 2) succinyl-CoA synthetase; 3) CoA-dependent succinic semialdehyde dehydrogenase; 4) glutamate:succinate semialdehyde transaminase; 5) glutamate decarboxylase; 6) 4-hydroxybutanoate dehydrogenase; 7) 4-hydroxybutyryl CoA:acetyl-CoA transferase; 8) aldehyde dehydrogenase; 9) alcohol dehydrogenase.
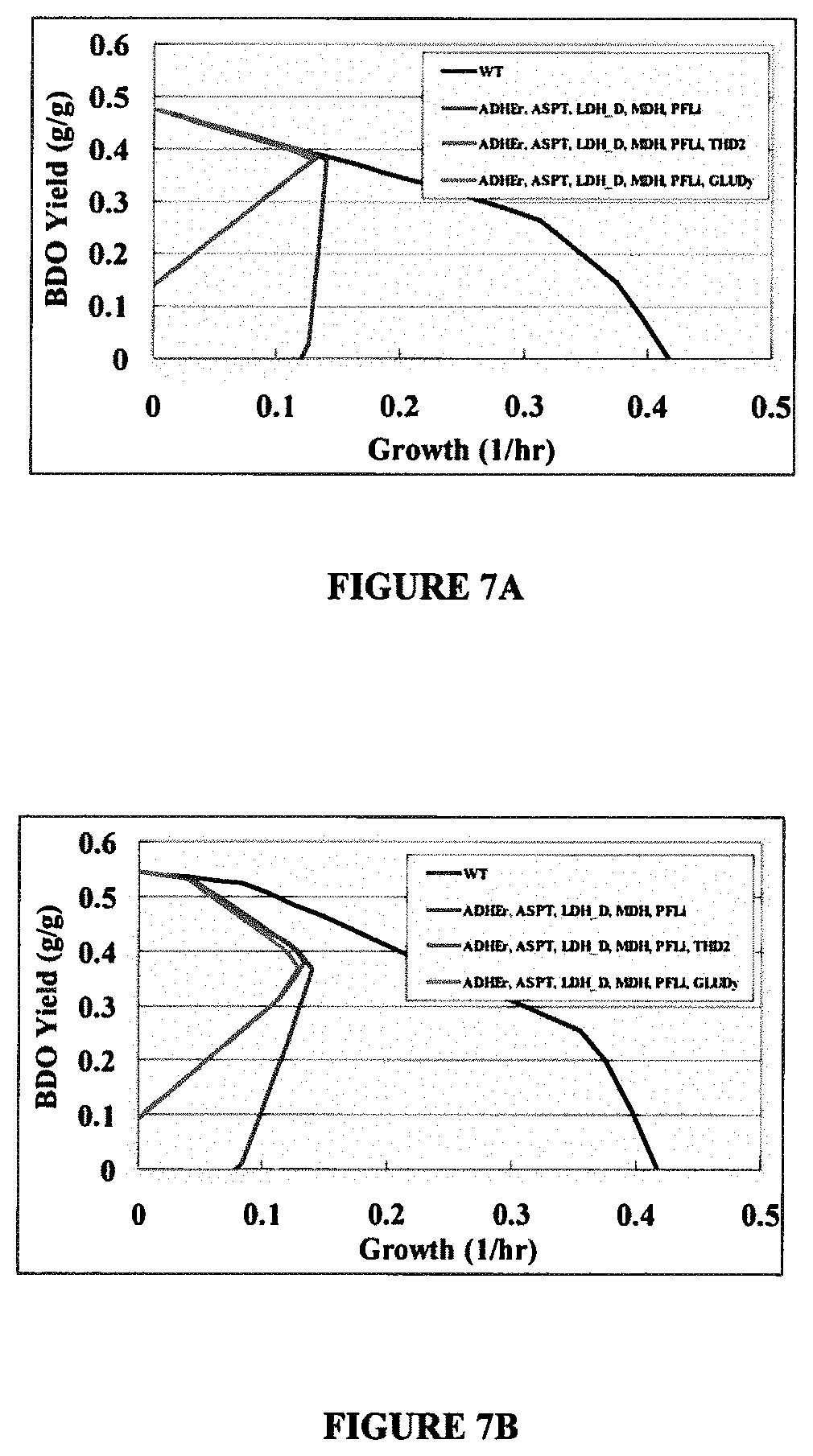
[0017] FIGS. 7A and 7B show the anaerobic growth rate versus BDO yield solution boundaries for an E. coli strain possessing the BDO production pathways shown and OptKnock predicted knockouts in FIG. 5 assuming in FIG. 7A PEP carboxykinase irreversibility and in FIG. 7B PEP carboxykinase reversibility. A basis glucose uptake rate of 20 mmol/gDW/hr is assumed along with a non-growth associated ATP maintenance requirement of 7.6 mmol/gDW/hr.
[0018] FIG. 8 shows a pictorial representation of E. coli central metabolism.
[0019] FIG. 9 shows the anaerobic growth rate versus BDO yield solution boundaries for an E. coli strain possessing the BDO production pathways shown and OptKnock predicted knockouts in FIG. 5 assuming PEP carboxykinase reversibility. A basis glucose uptake rate of 20 mmol/gDW/hr is assumed along with a non-growth associated ATP maintenance requirement of 7.6 mmol/gDW/hr.
DETAILED DESCRIPTION OF THE INVENTION
[0020] This invention is directed to the design and production of cells and organisms having biosynthetic production capabilities for 4-hydroxybutanoic acid (4-HB), .gamma.-butyrolactone and 1,4-butanediol. In one embodiment, the invention utilizes in silico stoichiometric models of Escherichia coli metabolism that identify metabolic designs for biosynthetic production of 4-hydroxybutanoic acid (4-HB) and 1,4-butanediol (BDO). The results described herein indicate that metabolic pathways can be designed and recombinantly engineered to achieve the biosynthesis of 4-HB and downstream products such as 1,4-butanediol in Escherichia coli and other cells or organisms. Biosynthetic production of 4-HB, for example, for the in silico designs can be confirmed by construction of strains having the designed metabolic genotype. These metabolically engineered cells or organisms also can be subjected to adaptive evolution to further augment 4-HB biosynthesis, including under conditions approaching theoretical maximum growth.
[0021] The invention is further directed to metabolic engineering strategies for attaining high yields of 1,4-butanediol (BDO) in Escherichia coli (see Examples V-VII). As disclosed herein, a genome-scale stoichiometric model of E. coli metabolism was employed using the bilevel optimization framework OptKnock to identify in silico strategies with multiple knockouts. The deletions are placed such that the redundancy in the network is reduced with the ultimate effect of coupling growth to the production of BDO in the network. The growth-coupled BDO production characteristic of the designed strains make them genetically stable and amenable to continuous bioprocesses. Strain design strategies were identified assuming the addition of non-native reaction capabilities into E. coli leading to a metabolic pathway from succinate semialdehyde to BDO. Out of the hundreds of strategies identified by OptKnock, one design emerged as satisfying multiple criteria. This design, utilizing the removal of adhE, ldhA, mdh, aspA, and pflAB, 1) led to a high predicted BDO yield at maximum growth, 2) required a reasonable number of knockouts, 3) had no detrimental effect on the maximum theoretical BDO yield, 4) brought about a tight coupling of BDO production with cell growth, and 5) was robust with respect to the assumed irreversibility or reversibility of PEP carboxykinase. Also disclosed herein are methods for the experimental testing of the strain designs and their evolution towards the theoretical maximum growth.
[0022] In certain embodiments, the 4-HB biosynthesis characteristics of the designed strains make them genetically stable and particularly useful in continuous bioprocesses. Separate strain design strategies were identified with incorporation of different non-native or heterologous reaction capabilities into E. coli leading to 4-HB and 1,4-butanediol producing metabolic pathways from either CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase and CoA-dependent succinic semialdehyde dehydrogenase, or glutamate:succinic semialdehyde transaminase. In silico metabolic designs were identified that resulted in the biosynthesis of 4-HB in both E. coli and yeast species from each of these metabolic pathways. The 1,4-butanediol intermediate .gamma.-butyrolactone can be generated in culture by spontaneous cyclization under conditions at pH<7.5, particularly under acidic conditions, such as below pH 5.5, for example, pH<7, pH<6.5, pH<6, and particularly at pH<5.5 or lower.
[0023] Strains identified via the computational component of the platform can be put into actual production by genetically engineering any of the predicted metabolic alterations which lead to the biosynthetic production of 4-HB, 1,4-butanediol or other intermediate and/or downstream products. In yet a further embodiment, strains exhibiting biosynthetic production of these compounds can be further subjected to adaptive evolution to further augment product biosynthesis. The levels of product biosynthesis yield following adaptive evolution also can be predicted by the computational component of the system.
[0024] As used herein, the term "non-naturally occurring" when used in reference to a microbial organism or microorganism of the invention is intended to mean that the microbial organism has at least one genetic alteration not normally found in a naturally occurring strain of the referenced species, including wild-type strains of the referenced species. Genetic alterations include, for example, modifications introducing expressible nucleic acids encoding metabolic polypeptides, other nucleic acid additions, nucleic acid deletions and/or other functional disruption of the microbial genetic material. Such modification include, for example, coding regions and functional fragments thereof, for heterologous, homologous or both heterologous and homologous polypeptides for the referenced species. Additional modifications include, for example, non-coding regulatory regions in which the modifications alter expression of a gene or operon. Exemplary metabolic polypeptides include enzymes within a 4-HB biosynthetic pathway and enzymes within a biosynthetic pathway for a BDO family of compounds.
[0025] A metabolic modification refers to a biochemical reaction that is altered from its naturally occurring state. Therefore, non-naturally occurring microorganisms having genetic modifications to nucleic acids encoding metabolic polypeptides or, functional fragments thereof. Exemplary metabolic modifications are described further below for both E. coli and yeast microbial organisms.
[0026] As used herein, the term "isolated" when used in reference to a microbial organism is intended to mean an organism that is substantially free of at least one component as the referenced microbial organism is found in nature. The term includes a microbial organism that is removed from some or all components as it is found in its natural environment. The term also includes a microbial organism that is removed from some or all components as the microbial organism is found in non-naturally occurring environments. Therefore, an isolated microbial organism is partly or completely separated from other substances as it is found in nature or as it is grown, stored or subsisted in non-naturally occurring environments. Specific examples of isolated microbial organisms include partially pure microbes, substantially pure microbes and microbes cultured in a medium that is non-naturally occurring.
[0027] As used herein, the terms "microbial," "microbial organism" or "microorganism" is intended to mean any organism that exists as a microscopic cell that is included within the domains of archaea, bacteria or eukarya. Therefore, the term is intended to encompass prokaryotic or eukaryotic cells or organisms having a microscopic size and includes bacteria, archaea and eubacteria of all species as well as eukaryotic microorganisms such as yeast and fungi. The term also includes cell cultures of any species that can be cultured for the production of a biochemical.
[0028] As used herein, the term "4-hydroxybutanoic acid" is intended to mean a 4-hydroxy derivative of butyric acid having the chemical formula C.sub.4H.sub.8O.sub.3 and a molecular mass of 104.11 g/mol (126.09 g/mol for its sodium salt). The chemical compound 4-hydroxybutanoic acid also is known in the art as 4-HB, 4-hydroxybutyrate, gamma-hydroxybutyric acid or GHB. The term as it is used herein is intended to include any of the compound's various salt forms and include, for example, 4-hydroxybutanoate and 4-hydroxybutyrate. Specific examples of salt forms for 4-HB include sodium 4-HB and potassium 4-HB. Therefore, the terms 4-hydroxybutanoic acid, 4-HB, 4-hydroxybutyrate, 4-hydroxybutanoate, gamma-hydroxybutyric acid and GHB as well as other art recognized names are used synonymously herein.
[0029] As used herein, the term "monomeric" when used in reference to 4-HB is intended to mean 4-HB in a non-polymeric or underivatized form. Specific examples of polymeric 4-HB include poly-4-hydroxybutanoic acid and copolymers of, for example, 4-HB and 3-HB. A specific example of a derivatized form of 4-HB is 4-HB-CoA. Other polymeric 4-HB forms and other derivatized forms of 4-HB also are known in the art.
[0030] As used herein, the term ".gamma.-butyrolactone" is intended to mean a lactone having the chemical formula C.sub.4H.sub.6O.sub.2 and a molecular mass of 86.089 g/mol. The chemical compound .gamma.-butyrolactone also is know in the art as GBL, butyrolactone, 1,4-lactone, 4-butyrolactone, 4-hydroxybutyric acid lactone, and gamma-hydroxybutyric acid lactone. The term as it is used herein is intended to include any of the compound's various salt forms.
[0031] As used herein, the term "1-4 butanediol" is intended to mean an alcohol derivative of the alkane butane, carrying two hydroxyl groups which has the chemical formula C.sub.4H.sub.10O.sub.2 and a molecular mass of 90.12 g/mol. The chemical compound 1-4 butanediol also is known in the art as BDO and is a chemical intermediate or precursor for a family of compounds referred to herein as BDO family of compounds, some of which are exemplified in FIG. 1.
[0032] As used herein, the term "tetrahydrofuran" is intended to mean a heterocyclic organic compound corresponding to the fully hydrogenated analog of the aromatic compound furan which has the chemical formula C.sub.4H.sub.8O and a molecular mass of 72.11 g/mol. The chemical compound tetrahydrofuran also is known in the art as THF, tetrahydrofuran, 1,4-epoxybutane, butylene oxide, cyclotetramethylene oxide, oxacyclopentane, diethylene oxide, oxolane, furanidine, hydrofuran, tetra-methylene oxide. The term as it is used herein is intended to include any of the compound's various salt forms.
[0033] As used herein, the term "CoA" or "coenzyme A" is intended to mean an organic cofactor or prosthetic group (nonprotein portion of an enzyme) whose presence is required for the activity of many enzymes (the apoenzyme) to form an active enzyme system. Coenzyme A functions in certain condensing enzymes, acts in acetyl or other acyl group transfer and in fatty acid synthesis and oxidation, pyruvate oxidation and in other acetylation.
[0034] As used herein, the term "substantially anaerobic" when used in reference to a culture or growth condition is intended to mean that the amount of oxygen is less than about 10% of saturation for dissolved oxygen in liquid media. The term also is intended to include sealed chambers of liquid or solid medium maintained with an atmosphere of less than about 1% oxygen.
[0035] The non-naturally occurring microbal organisms of the invention can contain stable genetic alterations, which refers to microorganisms that can be cultured for greater than five generations without loss of the alteration. Generally, stable genetic alterations include modifications that persist greater than 10 generations, particularly stable modifications will persist more than about 25 generations, and more particularly, stable genetic modifications will be greater than 50 generations, including indefinitely.
[0036] Those skilled in the art will understand that the genetic alterations, including metabolic modifications exemplified herein are described with reference to E. coli and yeast genes and their corresponding metabolic reactions. However, given the complete genome sequencing of a wide variety of organisms and the high level of skill in the area of genomics, those skilled in the art will readily be able to apply the teachings and guidance provided herein to essentially all other organisms. For example, the E. coli metabolic alterations exemplified herein can readily be applied to other species by incorporating the same or analogous encoding nucleic acid from species other than the referenced species. Such genetic alterations include, for example, genetic alterations of species homologs, in general, and in particular, orthologs, paralogs or nonorthologous gene displacements.
[0037] An ortholog is a gene or genes that are related by vertical descent and are responsible for substantially the same or identical functions in different organisms. For example, mouse epoxide hydrolase and human epoxide hydrolase can be considered orthologs for the biological function of hydrolysis of epoxides. Genes are related by vertical descent when, for example, they share sequence similarity of sufficient amount to indicate they are homologous, or related by evolution from a common ancestor. Genes can also be considered orthologs if they share three-dimensional structure but not necessarily sequence similarity, of a sufficient amount to indicate that they have evolved from a common ancestor to the extent that the primary sequence similarity is not identifiable. Genes that are orthologous can encode proteins with sequence similarity of about 25% to 100% amino acid sequence identity. Genes encoding proteins sharing an amino acid similarity less that 25% can also be considered to have arisen by vertical descent if their three-dimensional structure also shows similarities. Members of the serine protease family of enzymes, including tissue plasminogen activator and elastase, are considered to have arisen by vertical descent from a common ancestor.
[0038] Orthologs include genes or their encoded gene products that through, for example, evolution, have diverged in structure or overall activity. For example, where one species encodes a gene product exhibiting two functions and where such functions have been separated into distinct genes in a second species, the three genes and their corresponding products are considered to be orthologs. For the growth-coupled production of a biochemical product, those skilled in the art will understand that the orthologous gene harboring the metabolic activity to be disrupted is to be chosen for construction of the non-naturally occurring microorganism. An example of orthologs exhibiting separable activities is where distinct activities have been separated into distinct gene products between two or more species or within a single species. A specific example is the separation of elastase proteolysis and plasminogen proteolysis, two types of serine protease activity, into distinct molecules as plasminogen activator and elastase. A second example is the separation of mycoplasma 5'-3' exonuclease and Drosophila DNA polymerase III activity. The DNA polymerase from the first species can be considered an ortholog to either or both of the exonuclease or the polymerase from the second species and vice versa.
[0039] In contrast, paralogs are homologs related by, for example, duplication followed by evolutionary divergence and have similar or common, but not identical functions. Paralogs can originate or derive from, for example, the same species or from a different species. For example, microsomal epoxide hydrolase (epoxide hydrolase I) and soluble epoxide hydrolase (epoxide hydrolase II) can be considered paralogs because they represent two distinct enzymes, co-evolved from a common ancestor, that catalyze distinct reactions and have distinct functions in the same species. Paralogs are proteins from the same species with significant sequence similarity to each other suggesting that they are homologous, or related through co-evolution from a common ancestor. Groups of paralogous protein families include HipA homologs, luciferase genes, peptidases, and others.
[0040] A nonorthologous gene displacement is a nonorthologous gene from one species that can substitute for a referenced gene function in a different species. Substitution includes, for example, being able to perform substantially the same or a similar function in the species of origin compared to the referenced function in the different species. Although generally, a nonorthologous gene displacement will be identifiable as structurally related to a known gene encoding the referenced function, less structurally related but functionally similar genes and their corresponding gene products nevertheless will still fall within the meaning of the term as it is used herein. Functional similarity requires, for example, at least some structural similarity in the active site or binding region of a nonorthologous gene compared to a gene encoding the function sought to be substituted. Therefore, a nonorthologous gene includes, for example, a paralog or an unrelated gene.
[0041] Therefore, in identifying and constructing the non-naturally occurring microbial organisms of the invention having 4-HB, GBL and/or BDO biosynthetic capability, those skilled in the art will understand with applying the teaching and guidance provided herein to a particular species that the identification of metabolic modifications can include identification and inclusion or inactivation of orthologs. To the extent that paralogs and/or nonorthologous gene displacements are present in the referenced microorganism that encode an enzyme catalyzing a similar or substantially similar metabolic reaction, those skilled in the art also can utilize these evolutionally related genes.
[0042] Orthologs, paralogs and nonorthologous gene displacements can be determined by methods well known to those skilled in the art. For example, inspection of nucleic acid or amino acid sequences for two polypeptides will reveal sequence identity and similarities between the compared sequences. Based on such similarities, one skilled in the art can determine if the similarity is sufficiently high to indicate the proteins are related through evolution from a common ancestor. Algorithms well known to those skilled in the art, such as Align, BLAST, Clustal W and others compare and determine a raw sequence similarity or identity, and also determine the presence or significance of gaps in the sequence which can be assigned a weight or score. Such algorithms also are known in the art and are similarly applicable for determining nucleotide sequence similarity or identity. Parameters for sufficient similarity to determine relatedness are computed based on well known methods for calculating statistical similarity, or the chance of finding a similar match in a random polypeptide, and the significance of the match determined. A computer comparison of two or more sequences can, if desired, also be optimized visually by those skilled in the art. Related gene products or proteins can be expected to have a high similarity, for example, 25% to 100% sequence identity. Proteins that are unrelated can have an identity which is essentially the same as would be expected to occur by chance, if a database of sufficient size is scanned (about 5%). Sequences between 5% and 24% may or may not represent sufficient homology to conclude that the compared sequences are related. Additional statistical analysis to determine the significance of such matches given the size of the data set can be carried out to determine the relevance of these sequences.
[0043] Exemplary parameters for determining relatedness of two or more sequences using the BLAST algorithm, for example, can be as set forth below. Briefly, amino acid sequence alignments can be performed using BLASTP version 2.0.8 (Jan. 5, 1999) and the following parameters: Matrix: 0 BLOSUM62; gap open: 11; gap extension: 1; x_dropoff: 50; expect: 10.0; wordsize: 3; filter: on. Nucleic acid sequence alignments can be performed using BLASTN version 2.0.6 (Sep. 16, 1998) and the following parameters: Match: 1; mismatch: -2; gap open: 5; gap extension: 2; x_dropoff: 50; expect: 10.0; wordsize: 11; filter: off. Those skilled in the art will know what modifications can be made to the above parameters to either increase or decrease the stringency of the comparison, for example, and determine the relatedness of two or more sequences.
[0044] The invention provides a non-naturally occurring microbial biocatalyst including a microbial organism having a 4-hydroxybutanoic acid (4-HB) biosynthetic pathway that includes at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, glutamate:succinic semialdehyde transaminase or glutamate decarboxylase, wherein the exogenous nucleic acid is expressed in sufficient amounts to produce monomeric 4-hydroxybutanoic acid (4-HB). Succinyl-CoA synthetase is also referred to as succinyl-CoA synthase or succinyl CoA ligase.
[0045] The non-naturally occurring microbial biocatalysts of the invention include microbial organisms that employ combinations of metabolic reactions for biosynthetically producing the compounds of the invention. Exemplary compounds produced by the non-naturally occurring microorganisms include, for example, 4-hydroxybutanoic acid, 1,4-butanediol and .gamma.-butyrolactone. The relationships of these exemplary compounds with respect to chemical synthesis or biosynthesis are exemplified in FIG. 1.
[0046] In one embodiment, a non-naturally occurring microbial organism is engineered to produce 4-HB. This compound is one useful entry point into the 1,4-butanediol family of compounds. The biochemical reactions for formation of 4-HB from succinate, from succinate through succinyl-CoA or from .alpha.-ketoglutarate are shown in FIG. 2.
[0047] The invention is described herein with general reference to the metabolic reaction, reactant or product thereof, or with specific reference to one or more nucleic acids or genes encoding an enzyme associated with or catalyzing the referenced metabolic reaction, reactant or product. Unless otherwise expressly stated herein, those skilled in the art will understand that reference to a reaction also constitutes reference to the reactants and products of the reaction. Similarly, unless otherwise expressly stated herein, reference to a reactant or product also references the reaction and that reference to any of these metabolic constitutes also references the gene or genes encoding the enzymes that catalyze the referenced reaction, reactant or product. Likewise, given the well known fields of metabolic biochemistry, enzymology and genomics, reference herein to a gene or encoding nucleic acid also constitutes a reference to the corresponding encoded enzyme and the reaction it catalyzes as well as the reactants and products of the reaction.
[0048] The production of 4-HB via biosynthetic modes using the microbial organisms of the invention is particularly useful because it results in monomeric 4-HB. The non-naturally occurring microbial organisms of the invention and their biosynthesis of 4-HB and BDO family compounds also is particularly useful because the 4-HB product (1) is secreted; (2) is devoid of any derivatizations such as Coenzyme A; (3) avoids thermodynamic changes during biosynthesis, and (4) allows for the spontaneous chemical conversion of 4-HB to .gamma.-butyrolactone (GBL) in acidic pH medium. This latter characteristic is exemplified as step 7 in FIG. 2 and also is particularly useful for efficient chemical synthesis or biosynthesis of BDO family compounds such as 1,4-butanediol and/or tetrahydrofuran (THF), for example.
[0049] Microbial organisms generally lack the capacity to synthesize 4-HB and therefore, any of the compounds shown in FIG. 1 are known to be within the 1,4-butanediol family of compounds or known by those in the art to be within the 1,4-butanediol family of compounds. Moreover, organisms having all of the requisite metabolic enzymatic capabilities are not known to produce 4-HB from the enzymes described and biochemical pathways exemplified herein. Rather, with the possible exception of a few anaerobic microorganisms described further below, the microorganisms having the enzymatic capability use 4-HB as a substrate to produce, for example, succinate. In contrast, the non-naturally occurring microbial organisms of the invention generate 4-HB as a product. As described above, the biosynthesis of 4-HB in its monomeric form is not only particularly useful in chemical synthesis of BDO family of compounds, it also allows for the further biosynthesis of BDO family compounds and avoids altogether chemical synthesis procedures.
[0050] The non-naturally occurring microbial organisms of the invention that can produce monomeric 4-HB are produced by ensuring that a host microbial organism includes functional capabilities for the complete biochemical synthesis of at least one 4-HB biosynthetic pathway of the invention. Ensuring at least one requisite 4-HB biosynthetic pathway confers 4-HB biosynthesis capability onto the host microbial organism.
[0051] Three requisite 4-HB biosynthetic pathways are exemplified herein and shown for purposes of illustration in FIG. 2. One requisite 4-HB biosynthetic pathway includes the biosynthesis of 4-HB from succinate. The enzymes participating in this 4-HB pathway include CoA-independent succinic semialdehyde dehydrogenase and 4-hydroxybutanoate dehydrogenase. Another requisite 4-HB biosynthetic pathway includes the biosynthesis from succinate through succinyl-CoA. The enzymes participating in this 4-HB pathway include succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase and 4-hydroxybutanoate dehydrogenase. A third requisite 4-HB biosynthetic pathway includes the biosynthesis of 4-HB from .alpha.-ketoglutarate. The enzymes participating in this 4-HB biosynthetic pathway include glutamate:succinic semialdehyde transaminase, glutamate decarboxylase and 4-hydroxybutanoate dehydrogenase. Each of these 4-HB biosynthetic pathways, their substrates, reactants and products are described further below in the Examples.
[0052] The non-naturally occurring microbial organisms of the invention can be produced by introducing expressible nucleic acids encoding one or more of the enzymes participating in one or more 4-HB biosynthetic pathways. Depending on the host microbial organism chosen for biosynthesis, nucleic acids for some or all of a particular 4-HB biosynthetic pathway can be expressed. For example, if a chosen host is deficient in both enzymes in the succinate to 4-HB pathway (the succinate pathway) and this pathway is selected for 4-HB biosynthesis, then expressible nucleic acids for both CoA-independent succinic semialdehyde dehydrogenase and 4-hydroxybutanoate dehydrogenase are introduced into the host for subsequent exogenous expression. Alternatively, if the chosen host is deficient in 4-hydroxybutanoate dehydrogenase then an encoding nucleic acid is needed for this enzyme to achieve 4-HB biosynthesis.
[0053] In like fashion, where 4-HB biosynthesis is selected to occur through the succinate to succinyl-CoA pathway (the succinyl-CoA pathway), encoding nucleic acids for host deficiencies in the enzymes succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase and/or 4-hydroxybutanoate dehydrogenase are to be exogenously expressed in the recipient host. Selection of 4-HB biosynthesis through the .alpha.-ketoglutarate to succinic semialdehyde pathway (the .alpha.-ketoglutarate pathway) will utilize exogenous expression for host deficiencies in one or more of the enzymes for glutamate:succinic semialdehyde transaminase, glutamate decarboxylase and/or 4-hydroxybutanoate dehydrogenase.
[0054] Depending on the 4-HB biosynthetic pathway constituents of a selected host microbial organism, the non-naturally occurring microbial 4-HB biocatalysts of the invention will include at least one exogenously expressed 4-HB pathway-encoding nucleic acid and up to all six 4-HB pathway encoding nucleic acids. For example, 4-HB biosynthesis can be established from all three pathways in a host deficient in 4-hydroxybutanoate dehydrogenase through exogenous expression of a 4-hydroxybutanoate dehydrogenase encoding nucleic acid. In contrast, 4-HB biosynthesis can be established from all three pathways in a host deficient in all six enzymes through exogenous expression of all six of CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase and 4-hydroxybutanoate dehydrogenase.
[0055] Given the teachings and guidance provided herein, those skilled in the art will understand that the number of encoding nucleic acids to introduce in an expressible form will parallel the 4-HB pathway deficiencies of the selected host microbial organism. Therefore, a non-naturally occurring microbial organism of the invention can have one, two, three, four, five or six encoding nucleic acids encoding the above enzymes constituting the 4-HB biosynthetic pathways. In some embodiments, the non-naturally occurring microbial organisms also can include other genetic modifications that facilitate or optimize 4-HB biosynthesis or that confer other useful functions onto the host microbial organism. One such other functionality can include, for example, augmentation of the synthesis of one or more of the 4-HB pathway precursors such as succinate, succinyl-CoA and/or .alpha.-ketoglutarate.
[0056] In some embodiments, a non-naturally occurring microbial organism of the invention is generated from a host that contains the enzymatic capability to synthesize 4-HB. In this specific embodiment it can be useful to increase the synthesis or accumulation of a 4-HB pathway product to, for example, drive 4-HB pathway reactions toward 4-HB production. Increased synthesis or accumulation can be accomplished by, for example, overexpression of nucleic acids encoding one or more of the above-described 4-HB pathway enzymes. Over expression of the 4-HB pathway enzyme or enzymes can occur, for example, through exogenous expression of the endogenous gene or genes, or through exogenous expression of the heterologous gene or genes. Therefore, naturally occurring organisms can be readily generated to be non-naturally 4-HB producing microbial organisms of the invention through overexpression of one, two, three, four, five or all six nucleic acids encoding 4-HB biosynthetic pathway enzymes. In addition, a non-naturally occurring organism can be generated by mutagenesis of an endogenous gene that results in an increase in activity of an enzyme in the 4-HB biosynthetic pathway.
[0057] In particularly useful embodiments, exogenous expression of the encoding nucleic acids is employed. Exogenous expression confers the ability to custom tailor the expression and/or regulatory elements to the host and application to achieve a desired expression level that is controlled by the user. However, endogenous expression also can be utilized in other embodiments such as by removing a negative regulatory effector or induction of the gene's promoter when linked to an inducible promoter or other regulatory element. Thus, an endogenous gene having a naturally occurring inducible promoter can be up-regulated by providing the appropriate inducing agent, or the regulatory region of an endogenous gene can be engineered to incorporate an inducible regulatory element, thereby allowing the regulation of increased expression of an endogenous gene at a desired time. Similarly, an inducible promoter can be included as a regulatory element for an exogenous gene introduced into a non-naturally occurring microbial organism (see Example II).
[0058] "Exogenous" as it is used herein is intended to mean that the referenced molecule or the referenced activity is introduced into the host microbial organism. Therefore, the term as it is used in reference to expression of an encoding nucleic acid refers to introduction of the encoding nucleic acid in an expressible form into the microbial organism. When used in reference to a biosynthetic activity, the term refers to an activity that is introduced into the host reference organism. The source can be, for example, a homologous or heterologous encoding nucleic acids that expresses the referenced activity following introduction into the host microbial organism. Therefore, the term "endogenous" refers to a referenced molecule or activity that is present in the host. Similarly, the term when used in reference to expression of an encoding nucleic acid refers to expression of an encoding nucleic acid contained within the microbial organism. The term "heterologous" refers to a molecule or activity derived from a source other than the referenced species whereas "homologous" refers to a molecule or activity derived from the host microbial organism. Accordingly, exogenous expression of an encoding nucleic acid of the invention can utilize either or both a heterologous or homologous encoding nucleic acid.
[0059] Sources of encoding nucleic acids for a 4-HB pathway enzyme can include, for example, any species where the encoded gene product is capable of catalyzing the referenced reaction. Such species include both prokaryotic and eukaryotic organisms including, but not limited to, bacteria, including archaea and eubacteria, and eukaryotes, including yeast, plant, insect, animal, and mammal, including human. For example, the microbial organisms having 4-HB biosynthetic production are exemplified herein with reference to E. coli and yeast hosts. However, with the complete genome sequence available for now more than 550 species (with more than half of these available on public databases such as the NCBI), including 395 microorganism genomes and a variety of yeast, fungi, plant, and mammalian genomes, the identification of genes encoding the requisite 4-HB biosynthetic activity for one or more genes in related or distant species, including for example, homologues, orthologs, paralogs and nonorthologous gene displacements of known genes, and the interchange of genetic alterations between organisms is routine and well known in the art. Accordingly, the metabolic alterations enabling biosynthesis of 4-HB and other compounds of the invention described herein with reference to a particular organism such as E. coli or yeast can be readily applied to other microorganisms, including prokaryotic and eukaryotic organisms alike. Given the teachings and guidance provided herein, those skilled in the art will know that a metabolic alteration exemplified in one organism can be applied equally to other organisms.
[0060] In some instances, such as when an alternative 4-HB biosynthetic pathway exists in an unrelated species, 4-HB biosynthesis can be conferred onto the host species by, for example, exogenous expression of a paralog or paralogs from the unrelated species that catalyzes a similar, yet non-identical metabolic reaction to replace the referenced reaction. Because certain differences among metabolic networks exist between different organisms, those skilled in the art will understand that the actual genes usage between different organisms may differ. However, given the teachings and guidance provided herein, those skilled in the art also will understand that the teachings and methods of the invention can be applied to all microbial organisms using the cognate metabolic alterations to those exemplified herein to construct a microbial organism in a species of interest that will synthesize monomeric 4-HB.
[0061] Host microbial organisms can be selected from, and the non-naturally occurring microbial organisms generated in, for example, bacteria, yeast, fungus or any of a variety of other microorganisms applicable to fermentation processes. Exemplary bacteria include species selected from E. coli, Klebsiella oxytoca, Anaerobiospirillum succiniciproducens, Actinobacillus succinogenes, Mannheimia succiniciproducens, Rhizobium etli, Bacillus subtilis, Corynebacterium glutamicum, Gluconobacter oxydans, Zymomonas mobilis, Lactococcus lactis, Lactobacillus plantarum, Streptomyces coelicolor, Clostridium acetobutylicum, Pseudomonas fluorescens, and Pseudomonas putida. Exemplary yeasts or fungi include species selected from Saccharomyces cerevisiae, Schizosaccharomyces pombe, Kluyveromyces lactis, Kluyveromyces marxianus, Aspergillus terreus, Aspergillus niger and Pichia pastoris.
[0062] Methods for constructing and testing the expression levels of a non-naturally occurring 4-1-1B-producing host can be performed, for example, by recombinant and detection methods well known in the art. Such methods can be found described in, for example, Sambrook et al., Molecular Cloning: A Laboratory Manual, Third Ed., Cold Spring Harbor Laboratory, New York (2001); Ausubel et al., Current Protocols in Molecular Biology, John Wiley and Sons, Baltimore, Md. (1999). 4-HB and GBL can be separated by, for example, HPLC using a Spherisorb 5 ODS1 column and a mobile phase of 70% 10 mM phosphate buffer (pH=7) and 30% methanol, and detected using a UV detector at 215 nm (Hennessy et al. J. Forensic Sci. 46(6):1-9 (2004)). BDO is detected by gas chromatography or by HPLC and refractive index detector using an Aminex HPX-87H column and a mobile phase of 0.5 mM sulfuric acid (Gonzalez-Pajuelo et al., Met. Eng. 7:329-336 (2005)).
[0063] The non-naturally occurring microbial organisms of the invention are constructed using methods well known in the art as exemplified above to exogenously express at least one nucleic acid encoding a 4-HB pathway enzyme in sufficient amounts to produce monomeric 4-HB. Exemplary levels of expression for 4-HB enzymes in each pathway are described further below in the Examples. Following the teachings and guidance provided herein, the non-naturally occurring microbial organisms of the invention can achieve biosynthesis of monomeric 4-HB resulting in intracellular concentrations between about 0.1-25 mM or more. Generally, the intracellular concentration of monomeric 4-HB is between about 3-20 mM, particularly between about 5-15 mM and more particularly between about 8-12 mM, including about 10 mM or more. Intracellular concentrations between and above each of these exemplary ranges also can be achieved from the non-naturally occurring microbial organisms of the invention.
[0064] As described further below, one exemplary growth condition for achieving biosynthesis of 4-HB includes anaerobic culture or fermentation conditions. In certain embodiments, the non-naturally occurring microbial organisms of the invention can be sustained, cultured or fermented under anaerobic or substantially anaerobic conditions. Briefly, anaerobic conditions refers to an environment devoid of oxygen. Substantially anaerobic conditions include, for example, a culture, batch fermentation or continuous fermentation such that the dissolved oxygen concentration in the medium remains between 0 and 10% of saturation. Substantially anaerobic conditions also includes growing or resting cells in liquid medium or on solid agar inside a sealed chamber maintained with an atmosphere of less than 1% oxygen. The percent of oxygen can be maintained by, for example, sparging the culture with an N.sub.2/CO.sub.2 mixture or other suitable non-oxygen gas or gases.
[0065] The invention also provides a non-naturally occurring microbial biocatalyst including a microbial organism having 4-hydroxybutanoic acid (4-HB) and 1,4-butanediol (BDO) biosynthetic pathways that include at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, 4-hydroxybutyrate:CoA transferase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase, CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or alcohol dehydrogenase, wherein the exogenous nucleic acid is expressed in sufficient amounts to produce 1,4-butanediol (BDO).
[0066] Non-naturally occurring microbial organisms also can be generated which biosynthesize BDO. Following the teachings and guidance provided previously for the construction of microbial organisms that synthesize 4-HB, additional BDO pathways can be incorporated into the 4-HB producing microbial organisms to generate organisms that also synthesize BDO and other BDO family compounds. The chemical synthesis of BDO and its downstream products are illustrated in FIG. 3. The non-naturally occurring microbial organisms of the invention capable of BDO biosynthesis circumvent these chemical synthesis using 4-HB as an entry point as illustrated in FIG. 2. As described further below, the 4-HB producers can be used to chemically convert 4-HB to GBL and then to BDO or THF, for example. Alternatively, the 4-HB producers can be further modified to include biosynthetic capabilities for conversion of 4-HB and/or GBL to BDO.
[0067] The additional BDO pathways to introduce into 4-HB producers include, for example, the exogenous expression in a host deficient background or the overexpression of a CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or an alcohol dehydrogenase. In the absence of endogenous acyl-CoA synthetase capable of modifying 4-HB, the non-naturally occurring BDO producing microbial organisms can further include an exogenous acyl-CoA synthetase selective for 4-HB. Exemplary alcohol and aldehyde dehydrogenases that can be used for these in vivo conversions from 4-HB to BDO are listed below in Table 1.
TABLE-US-00001 TABLE 1 Alcohol and Aldehyde Dehydrogenases for Conversion of 4-HB to BDO. ALCOHOL DEHYDROGENASES ec:1.1.1.1 alcohol dehydrogenase ec:1.1.1.2 alcohol dehydrogenase (NADP+) ec:1.1.1.4 (R,R)-butanediol dehydrogenase ec:1.1.1.5 acetoin dehydrogenase ec:1.1.1.6 glycerol dehydrogenase ec:1.1.1.7 propanediol-phosphate dehydrogenase ec:1.1.1.8 glycerol-3-phosphate dehydrogenase (NAD+) ec:1.1.1.11 D-arabinitol 4-dehydrogenase ec:1.1.1.12 L-arabinitol 4-dehydrogenase ec:1.1.1.13 L-arabinitol 2-dehydrogenase ec:1.1.1.14 L-iditol 2-dehydrogenase ec:1.1.1.15 D-iditol 2-dehydrogenase ec:1.1.1.16 galactitol 2-dehydrogenase ec:1.1.1.17 mannitol-1-phosphate 5-dehydrogenase ec:1.1.1.18 inositol 2-dehydrogenase ec:1.1.1.21 aldehyde reductase ec:1.1.1.23 histidinol dehydrogenase ec:1.1.1.26 glyoxylate reductase ec:1.1.1.27 L-lactate dehydrogenase ec:1.1.1.28 D-lactate dehydrogenase ec:1.1.1.29 glycerate dehydrogenase ec:1.1.1.30 3-hydroxybutyrate dehydrogenase ec:1.1.1.31 3-hydroxyisobutyrate dehydrogenase ec:1.1.1.35 3-hydroxyacyl-CoA dehydrogenase ec:1.1.1.36 acetoacetyl-CoA reductase ec:1.1.1.37 malate dehydrogenase ec:1.1.1.38 malate dehydrogenase (oxaloacetate-decarboxylating) ec:1.1.1.39 malate dehydrogenase (decarboxylating) ec:1.1.1.40 malate dehydrogenase (oxaloacetate-decarboxylating) (NADP+) ec:1.1.1.41 isocitrate dehydrogenase (NAD+) ec:1.1.1.42 isocitrate dehydrogenase (NADP+) ec:1.1.1.54 allyl-alcohol dehydrogenase ec:1.1.1.55 lactaldehyde reductase (NADPH) ec:1.1.1.56 ribitol 2-dehydrogenase ec:1.1.1.59 3-hydroxypropionate dehydrogenase ec:1.1.1.60 2-hydroxy-3-oxopropionate reductase ec:1.1.1.61 4-hydroxybutyrate dehydrogenase ec:1.1.1.66 omega-hydroxydecanoate dehydrogenase ec:1.1.1.67 mannitol 2-dehydrogenase ec:1.1.1.71 alcohol dehydrogenase [NAD(P)+] ec:1.1.1.72 glycerol dehydrogenase (NADP+) ec:1.1.1.73 octanol dehydrogenase ec:1.1.1.75 (R)-aminopropanol dehydrogenase ec:1.1.1.76 (S,S)-butanediol dehydrogenase ec:1.1.1.77 lactaldehyde reductase ec:1.1.1.78 methylglyoxal reductase (NADH-dependent) ec:1.1.1.79 glyoxylate reductase (NADP+) ec:1.1.1.80 isopropanol dehydrogenase (NADP+) ec:1.1.1.81 hydroxypyruvate reductase ec:1.1.1.82 malate dehydrogenase (NADP+) ec:1.1.1.83 D-malate dehydrogenase (decarboxylating) ec:1.1.1.84 dimethylmalate dehydrogenase ec:1.1.1.85 3-isopropylmalate dehydrogenase ec:1.1.1.86 ketol-acid reductoisomerase ec:1.1.1.87 homoisocitrate dehydrogenase ec:1.1.1.88 hydroxymethylglutaryl-CoA reductase ec:1.1.1.90 aryl-alcohol dehydrogenase ec:1.1.1.91 aryl-alcohol dehydrogenase (NADP+) ec:1.1.1.92 oxaloglycolate reductase (decarboxylating) ec:1.1.1.94 glycerol-3-phosphate dehydrogenase [NAD(P)+] ec:1.1.1.95 phosphoglycerate dehydrogenase ec:1.1.1.97 3-hydroxybenzyl-alcohol dehydrogenase ec:1.1.1.101 acylglycerone-phosphate reductase ec:1.1.1.103 L-threonine 3-dehydrogenase ec:1.1.1.104 4-oxoproline reductase ec:1.1.1.105 retinol dehydrogenase ec:1.1.1.110 indolelactate dehydrogenase ec:1.1.1.112 indanol dehydrogenase ec:1.1.1.113 L-xylose 1-dehydrogenase ec:1.1.1.129 L-threonate 3-dehydrogenase ec:1.1.1.137 ribitol-5-phosphate 2-dehydrogenase ec:1.1.1.138 mannitol 2-dehydrogenase (NADP+) ec:1.1.1.140 sorbitol-6-phosphate 2-dehydrogenase ec:1.1.1.142 D-pinitol dehydrogenase ec:1.1.1.143 sequoyitol dehydrogenase ec:1.1.1.144 perillyl-alcohol dehydrogenase ec:1.1.1.156 glycerol 2-dehydrogenase (NADP+) ec:1.1.1.157 3-hydroxybutyryl-CoA dehydrogenase ec:1.1.1.163 cyclopentanol dehydrogenase ec:1.1.1.164 hexadecanol dehydrogenase ec:1.1.1.165 2-alkyn-1-ol dehydrogenase ec:1.1.1.166 hydroxycyclohexanecarboxylate dehydrogenase ec:1.1.1.167 hydroxymalonate dehydrogenase ec:1.1.1.174 cyclohexane-1,2-diol dehydrogenase ec:1.1.1.177 glycerol-3-phosphate 1-dehydrogenase (NADP+) ec:1.1.1.178 3-hydroxy-2-methylbutyryl-CoA dehydrogenase ec:1.1.1.185 L-glycol dehydrogenase ec:1.1.1.190 indole-3-acetaldehyde reductase (NADH) ec:1.1.1.191 indole-3-acetaldehyde reductase (NADPH) ec:1.1.1.192 long-chain-alcohol dehydrogenase ec:1.1.1.194 coniferyl-alcohol dehydrogenase ec:1.1.1.195 cinnamyl-alcohol dehydrogenase ec:1.1.1.198 (+)-borneol dehydrogenase ec:1.1.1.202 1,3-propanediol dehydrogenase ec:1.1.1.207 (-)-menthol dehydrogenase ec:1.1.1.208 (+)-neomenthol dehydrogenase ec:1.1.1.216 farnesol dehydrogenase ec:1.1.1.217 benzyl-2-methyl-hydroxybutyrate dehydrogenase ec:1.1.1.222 (R)-4-hydroxyphenyllactate dehydrogenase ec:1.1.1.223 isopiperitenol dehydrogenase ec:1.1.1.226 4-hydroxycyclohexanecarboxylate dehydrogenase ec:1.1.1.229 diethyl 2-methyl-3-oxosuccinate reductase ec:1.1.1.237 hydroxyphenylpyruvate reductase ec:1.1.1.244 methanol dehydrogenase ec:1.1.1.245 cyclohexanol dehydrogenase ec:1.1.1.250 D-arabinitol 2-dehydrogenase ec:1.1.1.251 galactitol 1-phosphate 5-dehydrogenase ec:1.1.1.255 mannitol dehydrogenase ec:1.1.1.256 fluoren-9-ol dehydrogenase ec:1.1.1.257 4-(hydroxymethyl)benzenesulfonate dehydrogenase ec:1.1.1.258 6-hydroxyhexanoate dehydrogenase ec:1.1.1.259 3-hydroxypimeloyl-CoA dehydrogenase ec:1.1.1.261 glycerol-1-phosphate dehydrogenase [NAD(P)+] ec:1.1.1.265 3-methylbutanal reductase ec:1.1.1.283 methylglyoxal reductase (NADPH-dependent) ec:1.1.1.286 isocitrate-homoisocitrate dehydrogenase ec:1.1.1.287 D-arabinitol dehydrogenase (NADP+) butanol dehydrogenase ALDEHYDE DEHYDROGENASES ec:1.2.1.2 formate dehydrogenase ec:1.2.1.3 aldehyde dehydrogenase (NAD+) ec:1.2.1.4 aldehyde dehydrogenase (NADP+) ec:1.2.1.5 aldehyde dehydrogenase [NAD(P)+] ec:1.2.1.7 benzaldehyde dehydrogenase (NADP+) ec:1.2.1.8 betaine-aldehyde dehydrogenase ec:1.2.1.9 glyceraldehyde-3-phosphate dehydrogenase (NADP+) ec:1.2.1.10 acetaldehyde dehydrogenase (acetylating) ec:1.2.1.11 aspartate-semialdehyde dehydrogenase ec:1.2.1.12 glyceraldehyde-3-phosphate dehydrogenase (phosphorylating) ec:1.2.1.13 glyceraldehyde-3-phosphate dehydrogenase (NADP+) (phosphorylating) ec:1.2.1.15 malonate-semialdehyde dehydrogenase ec:1.2.1.16 succinate-semialdehyde dehydrogenase [NAD(P)+] ec:1.2.1.17 glyoxylate dehydrogenase (acylating) ec:1.2.1.18 malonate-semialdehyde dehydrogenase (acetylating) ec:1.2.1.19 aminobutyraldehyde dehydrogenase ec:1.2.1.20 glutarate-semialdehyde dehydrogenase ec:1.2.1.21 glycolaldehyde dehydrogenase ec:1.2.1.22 lactaldehyde dehydrogenase ec:1.2.1.23 2-oxoaldehyde dehydrogenase (NAD+) ec:1.2.1.24 succinate-semialdehyde dehydrogenase ec:1.2.1.25 2-oxoisovalerate dehydrogenase (acylating) ec:1.2.1.26 2,5-dioxovalerate dehydrogenase ec:1.2.1.27 methylmalonate-semialdehyde dehydrogenase (acylating) ec:1.2.1.28 benzaldehyde dehydrogenase (NAD+) ec:1.2.1.29 aryl-aldehyde dehydrogenase ec:1.2.1.30 aryl-aldehyde dehydrogenase (NADP+) ec:1.2.1.31 L-aminoadipate-semialdehyde dehydrogenase ec:1.2.1.32 aminomuconate-semialdehyde dehydrogenase ec:1.2.1.36 retinal dehydrogenase ec:1.2.1.39 phenylacetaldehyde dehydrogenase ec:1.2.1.41 glutamate-5-semialdehyde dehydrogenase ec:1.2.1.42 hexadecanal dehydrogenase (acylating) ec:1.2.1.43 formate dehydrogenase (NADP+) ec:1.2.1.45 4-carboxy-2-hydroxymuconate-6-semialdehyde dehydrogenase ec:1.2.1.46 formaldehyde dehydrogenase ec:1.2.1.47 4-trimethylammoniobutyraldehyde dehydrogenase ec:1.2.1.48 long-chain-aldehyde dehydrogenase ec:1.2.1.49 2-oxoaldehyde dehydrogenase (NADP+) ec:1.2.1.51 pyruvate dehydrogenase (NADP+) ec:1.2.1.52 oxoglutarate dehydrogenase (NADP+) ec:1.2.1.53 4-hydroxyphenylacetaldehyde dehydrogenase ec:1.2.1.57 butanal dehydrogenase ec:1.2.1.58 phenylglyoxylate dehydrogenase (acylating) ec:1.2.1.59 glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) ec:1.2.1.62 4-formylbenzenesulfonate dehydrogenase ec:1.2.1.63 6-oxohexanoate dehydrogenase ec:1.2.1.64 4-hydroxybenzaldehyde dehydrogenase ec:1.2.1.65 salicylaldehyde dehydrogenase ec:1.2.1.66 mycothiol-dependent formaldehyde dehydrogenase ec:1.2.1.67 vanillin dehydrogenase ec:1.2.1.68 coniferyl-aldehyde dehydrogenase ec:1.2.1.69 fluoroacetaldehyde dehydrogenase ec:1.2.1.71 succinylglutamate-semialdehyde dehydrogenase
[0068] Therefore, in addition to any of the various modifications exemplified previously for establishing 4-HB biosynthesis in a selected host, the BDO producing microbial organisms can include any of the previous combinations and permutations of 4-HB pathway metabolic modifications as well as any combination of expression for CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or an alcohol dehydrogenase to generate biosynthetic pathways for BDO. Therefore, the BDO producers of the invention can have exogenous expression of, for example, one, two, three, four, five, six, seven, eight, nine or all 10 enzymes corresponding to any of the six 4-HB pathway and/or any of the 4 BDO pathway enzymes.
[0069] Design and construction of the genetically modified microbial organisms is carried out using methods well known in the art to achieve sufficient amounts of expression to produce BDO. In particular, the non-naturally occurring microbial organisms of the invention can achieve biosynthesis of BDO resulting in intracellular concentrations between about 0.1-25 mM or more. Generally, the intracellular concentration of BDO is between about 3-20 mM, particularly between about 5-15 mM and more particularly between about 8-12 mM, including about 10 mM or more. Intracellular concentrations between and above each of these exemplary ranges also can be achieved from the non-naturally occurring microbial organisms of the invention. As with the 4-HB producers, the BDO producers also can be sustained, cultured or fermented under anaerobic conditions.
[0070] The invention further provides a method for the production of 4-HB. The method includes culturing a non-naturally occurring microbial organism having a 4-hydroxybutanoic acid (4-HB) biosynthetic pathway comprising at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, glutamate:succinic semialdehyde transaminase or glutamate decarboxylase under substantially anaerobic conditions for a sufficient period of time to produce monomeric 4-hydroxybutanoic acid (4-HB). The method can additionally include chemical conversion of 4-HB to GBL and to BDO or THF, for example.
[0071] It is understood that, in methods of the invention, any of the one or more exogenous nucleic acids can be combined in a non-naturally occurring microbial organism of the invention so long as the desired product is produced, for example, 4-HB, BDO, THF or GBL. For example, a non-naturally occurring microbial organism having a 4-HB biosynthetic pathway can comprise at least two exogenous nucleic acids encoding desired enzymes, such as the combination of 4-hydroxybutanoate dehydrogenase and CoA-independent succinic semialdehyde dehydrogenase; 4-hydroxybutanoate dehydrogenase and CoA-dependent succinic semialdehyde dehydrogenase; CoA-dependent succinic semialdehyde dehydrogenase and succinyl-CoA synthetase; succinyl-CoA synthetase and glutamate decarboxylase, and the like. Thus, it is understood that any combination of two or more enzymes of a biosynthetic pathway can be included in a non-naturally occurring microbial organism of the invention. Similarly, it is understood that any combination of three or more enzymes of a biosynthetic pathway can be included in a non-naturally occurring microbial organism of the invention, for example, 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase and succinyl-CoA synthetase; 4-hydroxybutanoate dehydrogenase, CoA-dependent succinic semialdehyde dehydrogenase and glutamate:succinic semialdehyde transaminase, and so forth, as desired, so long as the combination of enzymes of the desired biosynthetic pathway results in production of the corresponding desired product.
[0072] Any of the non-naturally occurring microbial organisms described previously can be cultured to produce the biosynthetic products of the invention. For example, the 4-HB producers can be cultured for the biosynthetic production of 4-HB. The 4-HB can be isolated or be treated as described below to generate GBL, THF and/or BDO. Similarly, the BDO producers can be cultured for the biosynthetic production of BDO. The BDO can be isolated or subjected to further treatments for the chemical synthesis of BDO family compounds such as those downstream compounds exemplified in FIG. 3.
[0073] In some embodiments, culture conditions include anaerobic or substantially anaerobic growth or maintenance conditions. Exemplary anaerobic conditions have been described previously and are well known in the art. Exemplary anaerobic conditions for fermentation processes are described below in the Examples. Any of these conditions can be employed with the non-naturally occurring microbial organisms as well as other anaerobic conditions well known in the art. Under such anaerobic conditions, the 4-HB and BDO producers can synthesize monomeric 4-HB and BDO, respectively, at intracellular concentrations of 5-10 mM or more as well as all other concentrations exemplified previously.
[0074] A number of downstream compounds also can be generated for the 4-HB and BDO producing non-naturally occurring microbial organisms of the invention. With respect to the 4-HB producing microbial organisms of the invention, monomeric 4-HB and GBL exist in equilibrium in the culture medium. The conversion of 4-HB to GBL can be efficiently accomplished by, for example, culturing the microbial organisms in acid pH medium. A pH less than or equal to 7.5, in particular at or below pH 5.5, spontaneously converts 4-HB to GBL as illustrated in FIG. 1.
[0075] The resultant GBL can be separated from 4-HB and other components in the culture using a variety of methods well known in the art. Such separation methods include, for example, the extraction procedures exemplified in the Examples as well as methods which include continuous liquid-liquid extraction, pervaporation, membrane filtration, membrane separation, reverse osmosis, electrodialysis, distillation, crystallization, centrifugation, extractive filtration, ion exchange chromatography, size exclusion chromatography, adsorption chromatography, and ultrafiltration. All of the above methods are well known in the art. Separated GBL can be further purified by, for example, distillation.
[0076] Another down stream compound that can be produced from the 4-HB producing non-naturally occurring microbial organisms of the invention includes, for example, BDO. This compound can be synthesized by, for example, chemical hydrogenation of GBL. Chemical hydrogenation reactions are well known in the art. One exemplary procedure includes the chemical reduction of 4-HB and/or GBL or a mixture of these two components deriving from the culture using a heterogeneous or homogeneous hydrogenation catalyst together with hydrogen, or a hydride-based reducing agent used stoichiometrically or catalytically, to produce 1,4-butanediol.
[0077] Other procedures well known in the art are equally applicable for the above chemical reaction and include, for example, WO No. 82/03854 (Bradley, et al.), which describes the hydrogenolysis of gamma-butyrolactone in the vapor phase over a copper oxide and zinc oxide catalyst. British Pat. No. 1,230,276, which describes the hydrogenation of gamma-butyrolactone using a copper oxide-chromium oxide catalyst. The hydrogenation is carried out in the liquid phase. Batch reactions also are exemplified having high total reactor pressures. Reactant and product partial pressures in the reactors are well above the respective dew points. British Pat. No. 1,314,126, which describes the hydrogenation of gamma-butyrolactone in the liquid phase over a nickel-cobalt-thorium oxide catalyst. Batch reactions are exemplified as having high total pressures and component partial pressures well above respective component dew points. British Pat. No. 1,344,557, which describes the hydrogenation of gamma-butyrolactone in the liquid phase over a copper oxide-chromium oxide catalyst. A vapor phase or vapor-containing mixed phase is indicated as suitable in some instances. A continuous flow tubular reactor is exemplified using high total reactor pressures. British Pat. No. 1,512,751, which describes the hydrogenation of gamma-butyrolactone to 1,4-butanediol in the liquid phase over a copper oxide-chromium oxide catalyst. Batch reactions are exemplified with high total reactor pressures and, where determinable, reactant and product partial pressures well above the respective dew points. U.S. Pat. No. 4,301,077, which describes the hydrogenation to 1,4-butanediol of gamma-butyrolactone over a Ru--Ni--Co--Zn catalyst. The reaction can be conducted in the liquid or gas phase or in a mixed liquid-gas phase. Exemplified are continuous flow liquid phase reactions at high total reactor pressures and relatively low reactor productivities. U.S. Pat. No. 4,048,196, which describes the production of 1,4-butanediol by the liquid phase hydrogenation of gamma-butyrolactone over a copper oxide-zinc oxide catalyst. Further exemplified is a continuous flow tubular reactor operating at high total reactor pressures and high reactant and product partial pressures. And U.S. Pat. No. 4,652,685, which describes the hydrogenation of lactones to glycols.
[0078] A further downstream compound that can be produced form the 4-HB producing microbial organisms of the invention includes, for example, THF. This compound can be synthesized by, for example, chemical hydrogenation of GBL. One exemplary procedure well known in the art applicable for the conversion of GBL to THF includes, for example, chemical reduction of 4-HB and/or GBL or a mixture of these two components deriving from the culture using a heterogeneous or homogeneous hydrogenation catalyst together with hydrogen, or a hydride-based reducing agent used stoichiometrically or catalytically, to produce tetrahydrofuran. Other procedures well know in the art are equally applicable for the above chemical reaction and include, for example, U.S. Pat. No. 6,686,310, which describes high surface area sol-gel route prepared hydrogenation catalysts. Processes for the reduction of gamma butyrolactone to tetrahydrofuran and 1,4-butanediol also are described.
[0079] The culture conditions can include, for example, liquid culture procedures as well as fermentation and other large scale culture procedures. As described further below in the Examples, particularly useful yields of the biosynthetic products of the invention can be obtained under anaerobic or substantially anaerobic culture conditions.
[0080] The invention further provides a method of manufacturing 4-HB. The method includes fermenting a non-naturally occurring microbial organism having a 4-hydroxybutanoic acid (4-HB) biosynthetic pathway comprising at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, glutamate:succinic semialdehyde transaminase or glutamate decarboxylase under substantially anaerobic conditions for a sufficient period of time to produce monomeric 4-hydroxybutanoic acid (4-HB), the process comprising fed-batch fermentation and batch separation; fed-batch fermentation and continuous separation, or continuous fermentation and continuous separation.
[0081] The culture and chemical hydrogenations described above also can be scaled up and grown continuously for manufacturing of 4-HB, GBL, BDO and/or THF. Exemplary growth procedures include, for example, fed-batch fermentation and batch separation; fed-batch fermentation and continuous separation, or continuous fermentation and continuous separation. All of these processes are well known in the art. Employing the 4-HB producers allows for simultaneous 4-HB biosynthesis and chemical conversion to GBL, BDO and/or THF by employing the above hydrogenation procedures simultaneous with continuous cultures methods such as fermentation. Other hydrogenation procedures also are well known in the art and can be equally applied to the methods of the invention.
[0082] Fermentation procedures are particularly useful for the biosynthetic production of commercial quantities of 4-HB and/or BDO. Generally, and as with non-continuous culture procedures, the continuous and/or near-continuous production of 4-HB or BDO will include culturing a non-naturally occurring 4-HB or BDO producing organism of the invention in sufficient neutrients and medium to sustain and/or nearly sustain growth in an exponential phase. Continuous culture under such conditions can be include, for example, 1 day, 2, 3, 4, 5, 6 or 7 days or more. Additionally, continuous culture can include 1 week, 2, 3, 4 or 5 or more weeks and up to several months. Alternatively, organisms of the invention can be cultured for hours, if suitable for a particular application. It is to be understood that the continuous and/or near-continuous culture conditions also can include all time intervals in between these exemplary periods.
[0083] Fermentation procedures are well known in the art. Briefly, fermentation for the biosynthetic production of 4-HB, BDO or other 4-HB derived products of the invention can be utilized in, for example, fed-batch fermentation and batch separation; fed-batch fermentation and continuous separation, or continuous fermentation and continuous separation. Examples of batch and continuous fermentation procedures well known in the art are exemplified further below in the Examples.
[0084] In addition, to the above fermentation procedures using the 4-HB or BDO producers of the invention for continuous production of substantial quantities of monomeric 4-HB and BDO, respectively, the 4-HB producers also can be, for example, simultaneously subjected to chemical synthesis procedures as described previously for the chemical conversion of monomeric 4-HB to, for example, GBL, BDO and/or THF. The BDO producers can similarly be, for example, simultaneously subjected to chemical synthesis procedures as described previously for the chemical conversion of BDO to, for example, THF, GBL, pyrrolidones and/or other BDO family compounds. In addition, the products of the 4-HB and BDO producers can be separated from the fermentation culture and sequentially subjected to chemical conversion, as disclosed herein.
[0085] Briefly, hydrogenation of GBL in the fermentation broth can be performed as described by Frost et al., Biotechnology Progress 18: 201-211 (2002). Another procedure for hydrogenation during fermentation include, for example, the methods described in, for example, U.S. Pat. No. 5,478,952. This method is further exemplified in the Examples below.
[0086] Therefore, the invention additionally provides a method of manufacturing .gamma.-butyrolactone (GBL), tetrahydrofuran (THF) or 1,4-butanediol (BDO). The method includes fermenting a non-naturally occurring microbial organism having 4-hydroxybutanoic acid (4-FIB) and 1,4-butanediol (BDO) biosynthetic pathways, the pathways comprise at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, 4-hydroxybutyrate:CoA transferase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase, CoA-independent 1,4-butanediol semialdehyde dehydrogenase, CoA-dependent 1,4-butanediol semialdehyde dehydrogenase, 1,4-butanediol alcohol dehydrogenase, either CoA-dependent or independent, under substantially anaerobic conditions for a sufficient period of time to produce 1,4-butanediol (BDO), GBL or THF, the fermenting comprising fed-batch fermentation and batch separation; fed-batch fermentation and continuous separation, or continuous fermentation and continuous separation.
[0087] In addition to the biosynthesis of 4-HB, BDO and other products of the invention as described herein, the non-naturally occurring microbial organisms and methods of the invention also can be utilized in various combinations with each other and with other microbial organisms and methods well known in the art to achieve product biosynthesis by other routes. For example, one alternative to produce BDO other than use of the 4-HB producers and chemical steps or other than use of the BDO producer directly is through addition of another microbial organism capable of converting 4-HB or a 4-HB product exemplified herein to BDO.
[0088] One such procedure includes, for example, the fermentation of a 4-HB producing microbial organism of the invention to produce 4-HB, as described above and below. The 4-HB can then be used as a substrate for a second microbial organism that converts 4-HB to, for example, BDO, GBL and/or THF. The 4-HB can be added directly to another culture of the second organism or the original culture of 4-HB producers can be depleted of these microbial organisms by, for example, cell separation, and then subsequent addition of the second organism to the fermentation broth can utilized to produce the final product without intermediate purification steps. One exemplary second organism having the capacity to biochemically utilize 4-HB as a substrate for conversion to BDO, for example, is Clostridium acetobutylicum (see, for example, Jewell et al., Current Microbiology, 13:215-19 (1986)).
[0089] In other embodiments, the non-naturally occurring microbial organisms and methods of the invention can be assembled in a wide variety of subpathways to achieve biosynthesis of, for example, 4-HB and/or BDO as described. In these embodiments, biosynthetic pathways for a desired product of the invention can be segregated into different microbial organisms and the different microbial organisms can be co-cultured to produce the final product. In such a biosynthetic scheme, the product of one microbial organism is the substrate for a second microbial organism until the final product is synthesized. For example, the biosynthesis of BDO can be accomplished as described previously by constructing a microbial organism that contains biosynthetic pathways for conversion of a substrate such as endogenous succinate through 4-HB to the final product BDO. Alternatively, BDO also can be biosynthetically produced from microbial organisms through co-culture or co-fermentation using two organisms in the same vessel. A first microbial organism being a 4-HB producer with genes to produce 4-HB from succinic acid, and a second microbial organism being a BDO producer with genes to convert 4-HB to BDO.
[0090] Given the teachings and guidance provided herein, those skilled in the art will understand that a wide variety of combinations and permutations exist for the non-naturally occurring microbial organisms and methods of the invention together with other microbial organisms, with the co-culture of other non-naturally occurring microbial organisms having subpathways and with combinations of other chemical and/or biochemical procedures well known in the art to produce 4-HB, BDO, GBL and THF products of the invention.
[0091] One computational method for identifying and designing metabolic alterations favoring biosynthesis of a product is the OptKnock computational framework, Burgard et al., Biotechnol Bioeng, 84: 647-57 (2003). OptKnock is a metabolic modeling and simulation program that suggests gene deletion strategies that result in genetically stable microorganisms which overproduce the target product. Specifically, the framework examines the complete metabolic and/or biochemical network of a microorganism in order to suggest genetic manipulations that force the desired biochemical to become an obligatory byproduct of cell growth. By coupling biochemical production with cell growth through strategically placed gene deletions or other functional gene disruption, the growth selection pressures imposed on the engineered strains after long periods of time in a bioreactor lead to improvements in performance as a result of the compulsory growth-coupled biochemical production. Lastly, when gene deletions are constructed there is a negligible possibility of the designed strains reverting to their wild-type states because the genes selected by OptKnock are to be completely removed from the genome. Therefore, this computational methodology can be used to either identify alternative pathways that lead to biosynthesis of 4-HB and/or BDO or used in connection with the non-naturally occurring microbial organisms for further optimization of 4-HB and/or BDO biosynthesis.
[0092] Briefly, OptKnock is a term used herein to refer to a computational method and system for modeling cellular metabolism. The OptKnock program relates to a framework of models and methods that incorporate particular constraints into flux balance analysis (FBA) models. These constraints include, for example, qualitative kinetic information, qualitative regulatory information, and/or DNA microarray experimental data. OptKnock also computes solutions to various metabolic problems by, for example, tightening the flux boundaries derived through flux balance models and subsequently probing the performance limits of metabolic networks in the presence of gene additions or deletions. OptKnock computational framework allows the construction of model formulations that enable an effective query of the performance limits of metabolic networks and provides methods for solving the resulting mixed-integer linear programming problems. The metabolic modeling and simulation methods referred to herein as OptKnock are described in, for example, U.S. patent application Ser. No. 10/043,440, filed Jan. 10, 2002, and in International Patent No. PCT/US02/00660, filed Jan. 10, 2002.
[0093] Another computational method for identifying and designing metabolic alterations favoring biosynthetic production of a product is metabolic modeling and simulation system termed SimPheny.RTM.. This computational method and system is described in, for example, U.S. patent application Ser. No. 10/173,547, filed Jun. 14, 2002, and in International Patent Application No. PCT/US03/18838, filed Jun. 13, 2003.
[0094] SimPheny.RTM. is a computational system that can be used to produce a network model in silico and to simulate the flux of mass, energy or charge through the chemical reactions of a biological system to define a solution space that contains any and all possible functionalities of the chemical reactions in the system, thereby determining a range of allowed activities for the biological system. This approach is referred to as constraints-based modeling because the solution space is defined by constraints such as the known stoichiometry of the included reactions as well as reaction thermodynamic and capacity constraints associated with maximum fluxes through reactions. The space defined by these constraints can be interrogated to determine the phenotypic capabilities and behavior of the biological system or of its biochemical components. Analysis methods such as convex analysis, linear programming and the calculation of extreme pathways as described, for example, in Schilling et al., J. Theor. Biol. 203:229-248 (2000); Schilling et al., Biotech. Bioeng. 71:286-306 (2000) and Schilling et al., Biotech. Prog. 15:288-295 (1999), can be used to determine such phenotypic capabilities. As described in the Examples below, this computation methodology was used to identify and analyze the feasible as well as the optimal 4-HB biosynthetic pathways in 4-HB non-producing microbial organisms.
[0095] As described above, one constraints-based method used in the computational programs applicable to the invention is flux balance analysis. Flux balance analysis is based on flux balancing in a steady state condition and can be performed as described in, for example, Varma and Palsson, Biotech. Bioeng. 12:994-998 (1994). Flux balance approaches have been applied to reaction networks to simulate or predict systemic properties of, for example, adipocyte metabolism as described in Fell and Small, J. Biochem. 138:781-786 (1986), acetate secretion from E. coli under ATP maximization conditions as described in Majewski and Domach, Biotech. Bioeng. 35:732-738 (1990) or ethanol secretion by yeast as described in Vanrolleghem et al., Biotech. Prog. 12:434-448 (1996). Additionally, this approach can be used to predict or simulate the growth of E. coli on a variety of single-carbon sources as well as the metabolism of H. influenzae as described in Edwards and Palsson, Proc. Natl. Acad. Sci. USA 97:5528-5533 (2000), Edwards and Palsson, J. Biol. Chem. 274:17410-17416 (1999) and Edwards et al., Nature Biotech. 19:125-130 (2001).
[0096] Once the solution space has been defined, it can be analyzed to determine possible solutions under various conditions. This computational approach is consistent with biological realities because biological systems are flexible and can reach the same result in many different ways. Biological systems are designed through evolutionary mechanisms that have been restricted by fundamental constraints that all living systems must face. Therefore, constraints-based modeling strategy embraces these general realities. Further, the ability to continuously impose further restrictions on a network model via the tightening of constraints results in a reduction in the size of the solution space, thereby enhancing the precision with which physiological performance or phenotype can be predicted.
[0097] Given the teachings and guidance provided herein, those skilled in the art will be able to apply various computational frameworks for metabolic modeling and simulation to design and implement biosynthesis of 4-HB, BDO, GBL, THF and other BDO family compounds in host microbial organisms other than E. coli and yeast. Such metabolic modeling and simulation methods include, for example, the computational systems exemplified above as SimPheny.RTM. and OptKnock. For illustration of the invention, some methods are described herein with reference to the OptKnock computation framework for modeling and simulation. Those skilled in the art will know how to apply the identification, design and implementation of the metabolic alterations using OptKnock to any of such other metabolic modeling and simulation computational frameworks and methods well known in the art.
[0098] The ability of a cell or organism to biosynthetically produce a biochemical product can be illustrated in the context of the biochemical production limits of a typical metabolic network calculated using an in silico model. These limits are obtained by fixing the uptake rate(s) of the limiting substrate(s) to their experimentally measured value(s) and calculating the maximum and minimum rates of biochemical production at each attainable level of growth. The production of a desired biochemical generally is in direct competition with biomass formation for intracellular resources. Under these circumstances, enhanced rates of biochemical production will necessarily result in sub-maximal growth rates. The knockouts suggested by the above metabolic modeling and simulation programs such as OptKnock are designed to restrict the allowable solution boundaries forcing a change in metabolic behavior from the wild-type strain. Although the actual solution boundaries for a given strain will expand or contract as the substrate uptake rate(s) increase or decrease, each experimental point will lie within its calculated solution boundary. Plots such as these enable accurate predictions of how close the designed strains are to their performance limits which also indicates how much room is available for improvement.
[0099] The OptKnock mathematical framework is exemplified herein for pinpointing gene deletions leading to product biosynthesis and, particularly, growth-coupled product biosynthesis. The procedure builds upon constraint-based metabolic modeling which narrows the range of possible phenotypes that a cellular system can display through the successive imposition of governing physico-chemical constraints, Price et al., Nat. Rev. Microbiol., 2: 886-97 (2004). As described above, constraint-based models and simulations are well known in the art and generally invoke the optimization of a particular cellular objective, subject to network stoichiometry, to suggest a likely flux distribution.
[0100] Briefly, the maximization of a cellular objective quantified as an aggregate reaction flux for a steady state metabolic network comprising a set N={1, . . . , N} of metabolites and a set M={1, . . . , M} of metabolic reactions is expressed mathematically as follows:
maximize v cellular objective ##EQU00001## subject to j = 1 M S ij v j = 0 , .A-inverted. i .di-elect cons. N ##EQU00001.2## v substrate = v substrate_uptake mmol / gDQ hr ##EQU00001.3## .A-inverted. i .di-elect cons. { limiting substrate ( s ) } ##EQU00001.4## v atp .gtoreq. v atp_main mmol / gDW hr ##EQU00001.5## v j .gtoreq. 0 , .A-inverted. j .di-elect cons. { irrev . reactions } ##EQU00001.6##
[0101] where S.sub.ij is the stoichiometric coefficient of metabolite i in reaction j, v.sub.j is the flux of reaction j, v.sub.substrate_uptake represents the assumed or measured uptake rate(s) of the limiting substrate(s), and v.sub.atp_main is the non-growth associated ATP maintenance requirement. The vector v includes both internal and external fluxes. In this study, the cellular objective is often assumed to be a drain of biosynthetic precursors in the ratios required for biomass formation, Neidhardt, F. C. et al., Escherichia coli and Salmonella: Cellular and Molecular Biology, 2nd ed. 1996, Washington, D.C.: ASM Press. 2 v. (xx, 2822, lxxvi). The fluxes are generally reported per 1 gDWhr (gram of dry weight times hour) such that biomass formation is expressed as g biomass produced/gDWhr or l/hr.
[0102] The modeling of gene deletions, and thus reaction elimination, first employs the incorporation of binary variables into the constraint-based approach framework, Burgard et al., Biotechnol. Bioeng., 74: 364-375 (2001), Burgard et al., Biotechnol. Prog., 17: 791-797 (2001). These binary variables,
y j = { 1 , if reaction flux v j is active 0 , if reaction flux v j is not active , .A-inverted. j .di-elect cons. M ##EQU00002##
assume a value of 1 if reaction j is active and a value of 0 if it is inactive. The following constraint,
v.sub.j.sup.miny.sub.j.ltoreq.v.sub.j.ltoreq.v.sub.j.sup.maxy.sub.j,.A-i- nverted.j.di-elect cons.M
ensures that reaction flux v.sub.j is set to zero only if variable y.sub.j is equal to zero. Alternatively, when y.sub.j is equal to one, v.sub.j is free to assume any value between a lower v.sub.j.sup.min and an upper v.sub.j.sup.max bound. Here, v.sub.j.sup.min and v.sub.j.sup.max are identified by minimizing and maximizing, respectively, every reaction flux subject to the network constraints described above, Mahadevan et al., Metab. Eng., 5: 264-76 (2003).
[0103] Optimal gene/reaction knockouts are identified by solving a bilevel optimization problem that chooses the set of active reactions (y.sub.j=1) such that an optimal growth solution for the resulting network overproduces the chemical of interest. Mathematically, this bilevel optimization problem is expressed as the following bilevel mixed-integer optimization problem:
maximize y j v chemical ( OptKnock ) ##EQU00003## ( subject to v j maximize v biomass subject to j = 1 M S ij v j = 0 , .A-inverted. i .di-elect cons. N v substrate = v substrate_uptake .A-inverted. i .di-elect cons. { limiting substrate ( s ) } v atp .gtoreq. v atp_main ) ##EQU00003.2## v biomass .gtoreq. v biomass target ##EQU00003.3## v j min y j .ltoreq. v j .ltoreq. v j max y j , .A-inverted. j .di-elect cons. M ##EQU00003.4## j .di-elect cons. M forward ( 1 - y j ) = K ##EQU00003.5## y j .di-elect cons. { 0 , 1 } , .A-inverted. j .di-elect cons. M ##EQU00003.6##
where v.sub.chemical is the production of the desired target product, for example succinate or other biochemical product, and K is the number of allowable knockouts. Note that setting K equal to zero returns the maximum biomass solution of the complete network, while setting K equal to one identifies the single gene/reaction knockout (y.sub.j=0) such that the resulting network involves the maximum overproduction given its maximum biomass yield. The final constraint ensures that the resulting network meets a minimum biomass yield. Burgard et al., Biotechnol. Bioeng., 84: 647-57 (2003), provide a more detailed description of the model formulation and solution procedure. Problems containing hundreds of binary variables can be solved in the order of minutes to hours using CPLEX 8.0, GAMS: The Solver Manuals. 2003: GAMS Development Corporation, accessed via the GAMS, Brooke et al., GAMS Development Corporation (1998), modeling environment on an IBM RS6000-270 workstation. The OptKnock framework has already been able to identify promising gene deletion strategies for biochemical overproduction, Burgard et al., Biotechnol. Bioeng., 84: 647-57 (2003), Pharkya et al., Biotechnol. Bioeng., 84: 887-899 (2003), and establishes a systematic framework that will naturally encompass future improvements in metabolic and regulatory modeling frameworks.
[0104] Any solution of the above described bilevel OptKnock problem will provide one set of metabolic reactions to disrupt. Elimination of each reaction within the set or metabolic modification can result in 4-HB or BDO as an obligatory product during the growth phase of the organism. Because the reactions are known, a solution to the bilevel OptKnock problem also will provide the associated gene or genes encoding one or more enzymes that catalyze each reaction within the set of reactions. Identification of a set of reactions and their corresponding genes encoding the enzymes participating in each reaction is generally an automated process, accomplished through correlation of the reactions with a reaction database having a relationship between enzymes and encoding genes.
[0105] Once identified, the set of reactions that are to be disrupted in order to achieve 4-HB or BDO production are implemented in the target cell or organism by functional disruption of at least one gene encoding each metabolic reaction within the set. One particularly useful means to achieve functional disruption of the reaction set is by deletion of each encoding gene. However, in some instances, it can be beneficial to disrupt the reaction by other genetic aberrations including, for example, mutation, deletion of regulatory regions such as promoters or cis binding sites for regulatory factors, or by truncation of the coding sequence at any of a number of locations. These latter aberrations, resulting in less than total deletion of the gene set can be useful, for example, when rapid assessments of the succinate coupling are desired or when genetic reversion is less likely to occur.
[0106] To identify additional productive solutions to the above described bilevel OptKnock problem which lead to further sets of reactions to disrupt or metabolic modifications that can result in the biosynthesis, including growth-coupled biosynthesis of 4-HB or other biochemical product, an optimization method, termed integer cuts, can be implemented. This method proceeds by iteratively solving the OptKnock problem exemplified above with the incorporation of an additional constraint referred to as an integer cut at each iteration. Integer cut constraints effectively prevent the solution procedure from choosing the exact same set of reactions identified in any previous iteration that obligatory couples product biosynthesis to growth. For example, if a previously identified growth-coupled metabolic modification specifies reactions 1, 2, and 3 for disruption, then the following constraint prevents the same reactions from being simultaneously considered in subsequent solutions: y.sub.1+y.sub.2+y.sub.3.gtoreq.1. The integer cut method is well known in the art and can be found described in, for example, reference, Burgard et al., Biotechnol. Prog., 17: 791-797 (2001). As with all methods described herein with reference to their use in combination with the OptKnock computational framework for metabolic modeling and simulation, the integer cut method of reducing redundancy in iterative computational analysis also can be applied with other computational frameworks well known in the art including, for example, SimPheny.RTM..
[0107] Constraints of the above form preclude identification of larger reaction sets that include previously identified sets. For example, employing the integer cut optimization method above in a further iteration would preclude identifying a quadruple reaction set that specified reactions 1, 2, and 3 for disruption since these reactions had been previously identified. To ensure identification of all possible reaction sets leading to biosynthetic production of a product, a modification of the integer cut method can be employed.
[0108] Briefly, the modified integer cut procedure begins with iteration `zero` which calculates the maximum production of the desired biochemical at optimal growth for a wild-type network. This calculation corresponds to an OptKnock solution with K equaling 0. Next, single knockouts are considered and the two parameter sets, objstore.sub.iter and ystore.sub.iter,j, are introduced to store the objective function (v.sub.chemical) and reaction on-off information (y.sub.j), respectively, at each iteration, iter. The following constraints are then successively added to the OptKnock formulation at each iteration.
v.sub.chemical.gtoreq.objstore.sub.iter+.epsilon.-M.SIGMA..sub.j.di-elec- t cons.ystore.sub.iter,j.sub.=0y.sub.j
[0109] In the above equation, .epsilon. and M are a small and a large numbers, respectively. In general, .epsilon. can be set at about 0.01 and M can be set at about 1000. However, numbers smaller and/or larger then these numbers also can be used. M ensures that the constraint can be binding only for previously identified knockout strategies, while .epsilon. ensures that adding knockouts to a previously identified strategy must lead to an increase of at least .epsilon. in biochemical production at optimal growth. The approach moves onto double deletions whenever a single deletion strategy fails to improve upon the wild-type strain. Triple deletions are then considered when no double deletion strategy improves upon the wild-type strain, and so on. The end result is a ranked list, represented as desired biochemical production at optimal growth, of distinct deletion strategies that differ from each other by at least one knockout. This optimization procedure as well as the identification of a wide variety of reaction sets that, when disrupted, lead to the biosynthesis, including growth-coupled production, of a biochemical product. Given the teachings and guidance provided herein, those skilled in the art will understand that the methods and metabolic engineering designs exemplified herein are equally applicable to identify new biosynthetic pathways and/or to the obligatory coupling of cell or microorganism growth to any biochemical product.
[0110] The methods exemplified above and further illustrated in the Examples below enable the construction of cells and organisms that biosynthetically produce, including obligatory couple production of a target biochemical product to growth of the cell or organism engineered to harbor the identified genetic alterations. In this regard, metabolic alterations have been identified that result in the biosynthesis of 4-HB and 1,4-butanediol. Microorganism strains constructed with the identified metabolic alterations produce elevated levels of 4-HB or BDO compared to unmodified microbial organisms. These strains can be beneficially used for the commercial production of 4-HB, BDO, THF and GBL, for example, in continuous fermentation process without being subjected to the negative selective pressures.
[0111] Therefore, the computational methods described herein enable the identification and implementation of metabolic modifications that are identified by an in silico method selected from OptKnock or SimPheny. The set of metabolic modifications can include, for example, addition of one or more biosynthetic pathway enzymes and/.or functional disruption of one or more metabolic reactions including, for example, disruption by gene deletion.
[0112] Application of the OptKock method to identify pathways suitable for growth-coupled production of 1,4-butanediol (BDO) is described in Examples V-VII. As discussed above, the OptKnock methodology was developed on the premise that mutant microbial networks can be evolved towards their computationally predicted maximum-growth phenotypes when subjected to long periods of growth selection. In other words, the approach leverages an organism's ability to self-optimize under selective pressures. The OptKnock framework allows for the exhaustive enumeration of gene deletion combinations that force a coupling between biochemical production and cell growth based on network stoichiometry. The identification of optimal gene/reaction knockouts requires the solution of a bilevel optimization problem that chooses the set of active reactions such that an optimal growth solution for the resulting network overproduces the biochemical of interest (Burgard et al. Biotechnol. Bioeng. 84:647-657 (2003)).
[0113] Growth-coupled biochemical production can be visualized in the context of the biochemical production limits of a typical metabolic network. These limits are obtained from an in silico metabolic model by fixing the uptake rate(s) of the limiting substrate(s) to their experimentally measured value(s) and calculating the maximum and minimum rates of biochemical production at each attainable level of growth. The production envelopes essentially bracket what is possible by encompassing all potential biological phenotypes available to a given strain. Although exceptions exist, typically the production of a desired biochemical is in direct competition with biomass formation for intracellular resources (see FIG. 5, gray region). Thus increased biochemical yields will necessarily result in sub-maximal growth. Furthermore, the application of growth selective pressures may drive the performance of a non-growth-coupled production strain towards a low producing phenotype (point A, FIG. 5), regardless of its initial starting point. The knockouts suggested by OptKnock are designed to restrict the allowable solution boundary, forcing a change in metabolic behavior as depicted in FIG. 5 (cyan region). Evolutionary engineering approaches can thus be applied to drive the performance of an OptKnock designed strain to the rightmost boundary. This will result in a high producing strain that should be inherently stable to further growth selective pressures (point B, FIG. 5).
[0114] An in silico stoichiometric model of E. coli metabolism was employed to identify essential genes for metabolic pathways as exemplified previously and described in, for example, U.S. patent publications US 2002/0012939, US 2003/0224363, US 2004/0029149, US 2004/0072723, US 2003/0059792, US 2002/0168654 and US 2004/0009466, and in U.S. Pat. No. 7,127,379. As disclosed herein, the OptKnock mathematical framework was applied to pinpoint gene deletions leading to the growth-coupled production of BDO (see Examples V-VII). The strategies were initially determined by employing a reduced model of the E. coli metabolic network. This "small" model captures the key metabolic functionalities in the network, thus eliminating the redundancy associated with the genome-scale metabolic networks. Care was taken to ensure that the model was not reduced to the point where potentially active pathways possessing viable targets were neglected. Overall, the reduced model contained 262 reactions and its implementation reduced OptKnock CPU times approximately ten-fold when compared to the application of OptKnock to the genome-scale E. coli model (Reed et al., Genome Biol. 4:R54 (2003)).
[0115] Further, the solution of the bilevel OptKnock problem provides only one set of deletions. To enumerate all meaningful solutions, that is, all sets of knockouts leading to growth-coupled production formation, an optimization technique, termed integer cuts, can be implemented. This entails iteratively solving the OptKnock problem with the incorporation of an additional constraint referred to as an integer cut at each iteration, as discussed above.
[0116] For biochemical pathways to 1,4-butanediol, the conversion of glucose to BDO in E. coli is expected to derive from a total of three intracellular reduction steps from succinate semialdehyde (see FIG. 6). Succinate semialdehyde is natively produced by E. coli through the TCA cycle intermediate, alpha-ketoglutarate, via the action of two enzymes, glutamate:succinic semialdehyde transaminase and glutamate decarboxylase. An alternative pathway, used by the obligate anaerobe Clostridium kluyveri to degrade succinate, activates succinate to succinyl-CoA, and then converts succinyl-CoA to succinic semialdehyde using a CoA-dependant succinic semialdehyde dehydrogenase (Sohling and Gottschalk, Eur. J. Biochem. 212:121-127 (1993)). The conversion of succinate semialdehyde to BDO first requires the activity of 4-hydroxybutanoate (4-HB) dehydrogenase, an enzyme which is not native to E. coli or yeast but is found in various bacteria such as C. kluyveri and Ralstonia eutropha (Lutke-Eversloh and Steinbuchel, FEMS Microbiol. Lett. 181:63-71 (1999); Sohling and Gottschalk, J. Bacteriol. 178:871-880 (1996); Valentin et al., Eur. J. Biochem. 227:43-60 (1995); Wolff and Kenealy, Protein Expr. Purif. 6:206-212 (1995)). Precedent for the 4-HB to BDO conversion has been demonstrated in the strict anaerobe, Clostridium acetobutylicum, which when fed 4-HB, was shown to quantitatively produce BDO (Jewell et al., Current Microbiology 13:215-219 (1986)). The required biotransformations from 4-HB to BDO are assumed to be similar to those of the butyric acid to butanol conversion common to Clostridia species which proceeds via a CoA-derivative (Girbal et al., FEMS Microbiology Reviews 17:287-297 (1995)).
[0117] In an additional embodiment, the invention provides a non-naturally occurring microorganism comprising one or more gene disruptions, the one or more gene disruptions occurring in genes encoding an enzyme obligatory to coupling 1,4-butanediol production to growth of the microorganism when the gene disruption reduces an activity of the enzyme, whereby the one or more gene disruptions confers stable growth-coupled production of 1,4-butanediol onto the non-naturally occurring microorganism. The one or more gene disruptions can be, for example, those disclosed in Table 6 or 7. The one or more gene disruptions can comprise a deletion of the one or more genes, such as those genes disclosed in Table 6 or 7.
[0118] As disclosed herein, the non-naturally occurring microorganism can be a bacterium, yeast or fungus. For example, the non-naturally occurring microorganism can be a bacterium such as Escherichia coli, Klebsiella oxytoca, Anaerobiospirillum succiniciproducens, Actinobacillus succinogenes, Mannheimia succiniciproducens, Rhizobium etli, Bacillus subtilis, Corynebacterium glutamicum, Gluconobacter oxydans, Zymomonas mobilis, Lactococcus lactis, Lactobacillus plantarum, Streptomyces coelicolor, Clostridium acetobutylicum, Pseudomonas fluorescens, and Pseudomonas putida. The non-naturally occurring microorganism can also be a yeast such as Saccharomyces cerevisiae, Schizosaccharomyces pombe, Kluyveromyces lactis, Kluyveromyces marxianus, Aspergillus terreus, Aspergillus niger, and Pichia pastoris.
[0119] A non-naturally occurring microorganism of the invention can comprise a set of metabolic modifications obligatory coupling 1,4-butanediol production to growth of the microorganism, the set of metabolic modifications comprising disruption of one or more genes selected from the set of genes comprising adhE, ldhA, pflAB; adhE, ldhA, pflAB, mdh; adhE, ldhA, pflAB, mdh, mqo; adhE, ldhA, pflAB, mdh, aspA; adhE, mdh, ldhA, pflAB, sfcA; adhE, mdh, ldhA, pflAB, maeB; adhE, mdh, ldhA, pflAB, sfcA, maeB; adhE, ldhA, pflAB, mdh, pntAB; adhE, ldhA, pflAB, mdh, gdhA; adhE, ldhA, pflAB, mdh, pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW; and adhE, ldhA, pflAB, mdh, pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW, prpC, gsk, or an ortholog thereof, wherein the microorganism exhibits stable growth-coupled production of 1,4-butanediol.
[0120] In one embodiment, the invention provides a non-naturally occurring microorganism comprising a set of metabolic modifications obligatory to coupling 1,4-butanediol production to growth of the microorganism, the set of metabolic modifications comprising disruption of one or more genes, or an ortholog thereof, wherein the set of metabolic modifications comprises disruption of adhE and ldhA, wherein the microorganism exhibits stable growth-coupled production of 1,4-butanediol. In an additional embodiment, the set of metabolic modifications can further comprise disruption of mdh. In another embodiment, the set of metabolic modifications can further comprise disruption of one or more genes selected from the set of genes comprising mqo, aspA, sfcA, maeB, pntAB, and gdhA and can include, for example, disruption of sfcA and maeB. In still another embodiment, the set of metabolic modifications can further comprise disruption of one or more genes selected from the set of genes comprising pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW, prpC, and gsk, including disruption of all of pykA, pykF, dhaKLM, deoC, edd, yiaE and ycdW, and can further comprise disruption of prpC and gsk. Any of the above-described set of metabolic modifications can further comprise disruption of pflAB. In a particular embodiment, the set of metabolic modifications comprise disruption of one or more genes selected from the set of genes comprising adhE, ldhA, pflAB, mdh, and aspA, including up to all of genes adhE, ldhA, pflAB, mdh, and aspA.
[0121] A non-naturally occurring microorganism of the invention can further comprise a 1,4-butanediol (BDO) biosynthetic pathway comprising at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, 4-hydroxybutyrate:CoA transferase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase, CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or alcohol dehydrogenase, wherein the exogenous nucleic acid is expressed in sufficient amounts to produce 1,4-butanediol (BDO), as disclosed herein.
[0122] The invention additionally provides a method of producing a non-naturally occurring microorganism having stable growth-coupled production of 1,4-butanediol by identifying in silico a set of metabolic modifications requiring 1,4-butanediol production during exponential growth under a defined set of conditions, and genetically modifying a microorganism to contain the set of metabolic modifications requiring 1,4-butanediol production. Such methods can additionally include the addition of exogenous genes expressing desired enzyme activities to a microorganism. Such a set of metabolic modifications can be identified by an in silico method selected from OptKnock or SimPheny (see above and Examples V-VII).
[0123] The invention additionally provides a microorganism produced by the methods disclosed herein. Furthermore, the invention provides a method of producing 1,4-butanediol coupled to the growth of a microorganism. The method can include the steps of culturing under exponential growth phase in a sufficient amount of nutrients and media a non-naturally occurring microorganism comprising a set of metabolic modifications obligatorily coupling 1,4-butanediol production to growth of the microorganism, wherein the microorganism exhibits stable growth-coupled production of 1,4-butanediol, and isolating 1,4-butanediol produced from the non-naturally occurring microorganism. The set of metabolic modifications comprising disruption of one or more genes can be selected from the set of genes comprising adhE, ldhA, pflAB; adhE, ldhA, pflAB, mdh; adhE, ldhA, pflAB, mdh, mqo; adhE, ldhA, pflAB, mdh, aspA; adhE, mdh, ldhA, pflAB, sfcA; adhE, mdh, ldhA, pflAB, maeB; adhE, mdh, ldhA, pflAB, sfcA, maeB; adhE, ldhA, pflAB, mdh, pntAB; adhE, ldhA, pflAB, mdh, gdhA; adhE, ldhA, pflAB, mdh, pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW; and adhE, ldhA, pflAB, mdh, pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW, prpC, gsk, or an ortholog thereof.
[0124] In one embodiment, the invention provides a method of producing 1,4-butanediol coupled to the growth of a microorganism. The method can include the steps of culturing under exponential growth phase in a sufficient amount of nutrients and media a non-naturally occurring microorganism comprising a set of metabolic modifications obligatorily coupling 1,4-butanediol production to growth of the microorganism, the set of metabolic modifications comprising disruption of one or more genes, or an ortholog thereof, wherein the set of metabolic modifications comprises disruption of adhE and ldhA, wherein the microorganism exhibits stable growth-coupled production of 1,4-butanediol; and isolating 1,4-butanediol produced from the non-naturally occurring microorganism. In an additional embodiment, the set of metabolic modifications can further comprise disruption of mdh. In another embodiment, the set of metabolic modifications can further comprise disruption of one or more genes selected from the set of genes comprising mqo, aspA, sfcA, maeB, pntAB, and gdhA and can include, for example, disruption of sfcA and maeB. In still another embodiment, the set of metabolic modifications can further comprise disruption of one or more genes selected from the set of genes comprising pykA, pykF, dhaKLM, deoC, edd, yiaE, ycdW, prpC, and gsk, including disruption of all of pykA, pykF, dhaKLM, deoC, edd, yiaE and ycdW, and can further comprise disruption of prpC and gsk. Any of the above-described set of metabolic modifications can further comprise disruption of pflAB.
[0125] In a method of producing BDO, the non-naturally occurring microorganism can further comprise a 1,4-butanediol (BDO) biosynthetic pathway comprising at least one exogenous nucleic acid encoding 4-hydroxybutanoate dehydrogenase, CoA-independent succinic semialdehyde dehydrogenase, succinyl-CoA synthetase, CoA-dependent succinic semialdehyde dehydrogenase, 4-hydroxybutyrate:CoA transferase, glutamate:succinic semialdehyde transaminase, glutamate decarboxylase, CoA-independent aldehyde dehydrogenase, CoA-dependent aldehyde dehydrogenase or alcohol dehydrogenase, wherein the exogenous nucleic acid is expressed in sufficient amounts to produce 1,4-butanediol (BDO).
[0126] It is understood that modifications which do not substantially affect the activity of the various embodiments of this invention are also included within the definition of the invention provided herein. Accordingly, the following examples are intended to illustrate but not limit the present invention.
Example I
Biosynthesis of 4-Hydroxybutanoic Acid
[0127] This Example describes the biochemical pathways for 4-HB production.
[0128] Previous reports of 4-HB synthesis in microbes have focused on this compound as an intermediate in production of the biodegradable plastic poly-hydroxyalkanoate (PHA) (U.S. Pat. No. 6,117,658). The use of 4-HB/3-HB copolymers can be advantageous over the more traditional poly-3-hydroxybutyrate polymer (PHB) because the resulting plastic is less brittle (Saito and Doi, Intl. J. Biol. Macromol. 16:99-104 (1994)). The production of monomeric 4-HB described herein is a fundamentally different process for several reasons: 1). The product is secreted, as opposed to PHA which is produced intracellularly and remains in the cell; 2) for organisms that produce hydroxybutanoate polymers, free 4-HB is not produced, but rather the Coenzyme A derivative is used by the polyhydroxyalkanoate synthase; 3) in the case of the polymer, formation of the granular product changes thermodynamics; and 4) extracellular pH is not an issue for production of the polymer, whereas it will affect whether 4-HB is present in the free acid or conjugate base state, and also the equilibrium between 4-HB and GBL.
[0129] 4-HB can be produced in two enzymatic reduction steps from succinate, a central metabolite of the TCA cycle, with succinic semialdehyde as the intermediate (FIG. 2). The first of these enzymes, succinic semialdehyde dehydrogenase, is native to many organisms including E. coli, in which both NADH- and NADPH-dependent enzymes have been found (Donnelly and Cooper, Eur. J. Biochem. 113:555-561 (1981); Donnelly and Cooper, J. Bacteriol. 145:1425-1427 (1981); Marek and Henson, J. Bacteriol. 170:991-994 (1988)). There is also evidence supporting succinic semialdehyde dehydrogenase activity in S. cerevisiae (Ramos et al., Eur. J. Biochem. 149:401-404 (1985)), and a putative gene has been identified by sequence homology. However, most reports indicate that this enzyme proceeds in the direction of succinate synthesis, opposite to that shown in FIG. 2 (Donnelly and Cooper, supra; Lutke-Eversloh and Steinbuchel, FEMS Microbiol. Lett. 181:63-71 (1999)), participating in the degradation pathway of 4-HB and gamma-aminobutyrate. An alternative pathway, used by the obligate anaerobe Clostridium kluyveri to degrade succinate, activates succinate to succinyl-CoA, then converts succinyl-CoA to succinic semialdehyde using an alternative succinic semialdehyde dehydrogenase which is known to function in this direction (Sohling and Gottschalk, Eur. J. Biochem. 212:121-127 (1993)). However, this route has the energetic cost of ATP required to convert succinate to succinyl-CoA.
[0130] The second enzyme of the pathway, 4-hydroxybutanoate dehydrogenase, is not native to E. coli or yeast but is found in various bacteria such as C. kluyveri and Ralstonia eutropha (Lutke-Eversloh and Steinbuchel, supra; Sohling and Gottschalk, J. Bacteriol. 178:871-880 (1996); Valentin et al., Eur. J. Biochem. 227:43-60 (1995); Wolff and Kenealy, Protein Expr. Purif. 6:206-212 (1995)). These enzymes are known to be NADH-dependent, though NADPH-dependent forms can exist. An additional pathway to 4-HB from alpha-ketoglutarate was demonstrated in E. coli resulting in the accumulation of poly(4-hydroxybutyric acid) (Song et al., Wei Sheng Wu Xue.Bao. 45:382-386 (2005)). The recombinant strain required the overexpression of three heterologous genes, PHA synthase (R. eutropha), 4-hydroxybutyrate dehydrogenase (R. eutropha) and 4-hydroxybutyrate:CoA transferase (C. kluyveri), along with two native E. coli genes: glutamate:succinic semialdehyde transaminase and glutamate decarboxylase. Steps 4 and 5 in FIG. 2 can alternatively be carried out by an alpha-ketoglutarate decarboxylase such as the one identified in Euglena gracilis (Shigeoka et al., Biochem. J. 282(Pt2):319-323 (1992); Shigeoka and Nakano, Arch. Biochem. Biophys. 288:22-28 (1991); Shigeoka and Nakano, Biochem J. 292(Pt 2):463-467 (1993)). However, this enzyme has not yet been applied to impact the production of 4-HB or related polymers in any organism.
[0131] The reported directionality of succinic semialdehyde dehydrogenase led to the investigation of the thermodynamics of 4-HB metabolism. Specifically, this study investigated whether or not the reactions involved in the conversion of succinate or succinyl-CoA to 4-HB are thermodynamically favorable (i.e., .DELTA.G.sub.r<0) under the typical physiological conditions present in E. coli and S. cerevisiae. All oxidation/reduction reactions were assumed to utilize NADH, although the results for assuming NADPH utilization would be similar. Standard Gibbs free energies of formation (.DELTA.G.sub.f.sup.o) were calculated for each compound in FIG. 2 based on the group contribution method (Mavrovouniotis, M. L., J. Biol. Chem.266:14440-14445 (1991)). Each standard Gibbs energy of formation was then transformed in order to obtain a criterion of spontaneous change at specified pressure, temperature, pH, and ionic strength (Alberty, R. A., Biochem. Biophys. Acta 1207:1-11 (1994) (equation 1).
.DELTA. G f ' ( I , pH ) = .DELTA. G f o ( I = 0 ) + N H RT ln ( 10 pH ) - 2.915 I z 2 - N H 1 + B I ( 1 ) ##EQU00004##
[0132] Where .DELTA.G.sub.f.sup.o is the standard Gibbs energy of formation, NH is the number of hydrogen atoms in the compound, R is the universal gas constant, T is constant at 298K, z is the charge of the molecule at the pH of interest, I is the ionic strength in M, and B is a constant equal to 1.6 L.sup.0.5/mol.sup.0.5.
[0133] Equation 1 reveals that both intracellular pH and ionic strength play a role in determining thermodynamic feasibility. Normally, intracellular pH of cells is very well regulated, even when there are large variations in the culture pH. The intracellular pH of E. coli and S. cerevisiae have both been reported in the literature. E. coli maintains an intracellular pH of 7.4-7.7 during typical growth conditions in neutral buffers, but can drop to 7.2 in pH 6 medium, and even go as low as 6.9 for external pH of 5 (Riondet et al., Biotechnology Tech. 11:735-738 (1997)). However, growth of E. coli is severely inhibited at external pH below 6. Yeast pH exhibits more variation. During exponential growth phase, S. cerevisiae internal pH has been measured to be in the range of 6.7-7.0 with external pH controlled at 5.0 (Dombek and Ingram, Appl. Environ. Microbiol. 53:1286-1291 (1987)). On the other hand, in resting cells the internal pH drops to below 6 when the external pH is 6 or less (Imai and Ohno, J. Biotechnol. 38:165-172 (1995)). This analysis assumes an intracellular pH of 7.4 for E. coli and 6.8 for S. cerevisiae. An ionic strength of 0.15 also was assumed (Valenti et al., supra).
[0134] Transformed Gibbs energies of formation were calculated at the standard state (pH=7.0, I=0) and at physiological states of E. coli (pH=7.4, I=0.15) and S. cerevisiae (pH=6.8, I=0.15). Transformed Gibbs energies of reaction (.DELTA.G.sub.r') were then calculated by taking the difference in .DELTA.G.sub.f' between the products and reactants. The transformed Gibbs energies of the reactions necessary to convert succinate or succinyl-CoA to 4-HB are provided in Table 2. Note that the standard error, U.sub.f,est, on .DELTA.G.sub.f calculated by the group contribution theory is 4 kcal/mol. The uncertainty in .DELTA.G.sub.r, U.sub.r,est, can be calculated as the Euclidean norm of the uncertainty for .DELTA.G.sub.f of each compound (Equation).
U r , est = i = 1 m n i 2 * U f , est 2 = i = 1 m 16 n i 2 ( 2 ) ##EQU00005##
[0135] Where n is the stochiometric coefficient and i is the compound. For the examined reactions, this uncertainty is on the order of 8 kcal/mol.
TABLE-US-00002 TABLE 2 Gibbs free energy of reaction (kcal/mole) at different pH and ionic strength values. The first column is under standard conditions, while the others are adjusted according to equation 1. Temperature is constant at 298 K. Error bars for these values are on the order of 8 kcal/mol, as calculated by equation 2. .DELTA.G.sub.r.sup.o' .DELTA.G.sub.r' .DELTA.G.sub.r' pH = 7.0 pH = 7.4 pH = 6.8 Reaction IS = 0 IS = 0.15M IS = 0.15M succ + NADH + 2 H+ .fwdarw. sucsa + NAD + h2o 12.0 14.4 12.8 succ + coa + ATP .fwdarw. succoa + ADP + Pi 0.30 -0.03 -0.03 succoa + NADH + H+ .fwdarw. sucsa + NAD + coa 4.4 7.0 6.2 sucsa + NADH + H+ .fwdarw. 4hb + NAD -5.0 -3.8 -4.6 Abbreviations: suc, succinate; sucsa, succinic semialdehyde; succoa, succinyl-CoA; Pi, inorganic phosphate.
[0136] Table 2 reveals that the reaction most likely to encounter a thermodynamic barrier after considering potential uncertainty in our calculations is succinic semialdehyde dehydrogenase (step 1 in FIG. 2). Whether this reaction can be driven closer to thermodynamic feasibility by varying the assumed concentrations of the participating metabolites also was studied. For example, the standard Gibbs energies assume concentrations of 1 M for all participating compounds (except water). In an anaerobic environment, NADH will be present at a several-fold higher concentration than NAD. Assuming [NADH]=5.times.[NAD], we calculated the effect on .DELTA.G.sub.r' using the equation
.DELTA. G f ' = .DELTA. G f 0 ' + RT ln [ prod ] [ react ] ( 3 ) ##EQU00006##
[0137] This change results in a difference of only about 1 kcal/mol in the delta G values for succinic semialdehyde dehydrogenase. Equation 3 was also used to calculate other effects on .DELTA.G.sub.r, such as high succinate concentration to drive the reactions. A 1000-fold difference in the concentrations of succinate and succinic semialdehyde will contribute about 5 kcal/mol to delta G. Taken together with an assumed uncertainty of 8 kcal/mol, the possibility that succinic semialdehyde dehydrogenase will operate in the direction towards succinic semialdehyde under some set of physiological conditions cannot be eliminated. Thus we still consider the direct route from succinate to 4-HB in our subsequent theoretical analysis.
[0138] The microbial production capabilities of 4-hydroxybutyrate were explored in two microbes, Escherichia coli and Saccharomyces cerevisiae, using in silico metabolic models of each organism. Potential pathways to 4-HB proceed via a succinate, succinyl-CoA, or alpha-ketoglutarate intermediate as shown in FIG. 2.
[0139] The first step in the 4-HB production pathway from succinate involves the conversion of succinate to succinic semialdehyde via an NADH- or NADPH-dependant succinic semialdehyde dehydrogenase. In E. coli, gabD is an NADP-dependant succinic semialdehyde dehydrogenase and is part of a gene cluster involved in 4-aminobutyrate uptake and degradation (Niegemann et al., Arch. Microbiol. 160:454-460 (1993); Schneider et al., J. Bacteriol. 184:6976-6986 (2002)). sad is believed to encode the enzyme for NAD-dependant succinic semialdehyde dehydrogenase activity (Marek and Henson, supra). S. cerevisiae contains only the NADPH-dependant succinic semialdehyde dehydrogenase, putatively assigned to UGA2, which localizes to the cytosol (Huh et al., Nature 425:686-691 (2003)). The maximum yield calculations assuming the succinate pathway to 4-HB in both E. coli and S. cerevisiae require only the assumption that a non-native 4-HB dehydrogenase has been added to their metabolic networks.
[0140] The pathway from succinyl-CoA to 4-hydroxybutyrate was described in U.S. Pat. No. 6,117,658 as part of a process for making polyhydroxyalkanoates comprising 4-hydroxybutyrate monomer units. Clostridium kluyveri is one example organism known to possess CoA-dependant succinic semialdehyde dehydrogenase activity (Sohling and Gottschalk, supra; Sohling and Gottschalk, supra). In this study, it is assumed that this enzyme, from C. kluyveri or another organism, is expressed in E. coli or S. cerevisiae along with a non-native or heterologous 4-HB dehydrogenase to complete the pathway from succinyl-CoA to 4-HB. The pathway from alpha-ketoglutarate to 4-HB was demonstrated in E. coli resulting in the accumulation of poly(4-hydroxybutyric acid) to 30% of dry cell weight (Song et al., supra). As E. coli and S. cerevisiae natively or endogenously possess both glutamate:succinic semialdehyde transaminase and glutamate decarboxylase (Coleman et al., J. Biol. Chem. 276:244-250 (2001)), the pathway from AKG to 4-HB can be completed in both organisms by assuming only that a non-native 4-HB dehydrogenase is present.
Example II
Production of 4-Hydroxybutanoic Acid in E. coli
[0141] This Example describes the biosynthetic yields for 4-hydroxybutanoic acid resulting from each biochemical pathway.
[0142] In this section, the maximum theoretical yields of 4-HB from glucose are calculated assuming that each of the three metabolic pathways depicted in FIG. 2 are functional in E. coli. A genome-scale metabolic model of E. coli, similar to the one described in Reed et al., Genome Biol. 4:R54 (2003), was used as the basis for the analysis. The energetic gain, in terms of ATP molecules produced, of each maximum yielding pathway is calculated assuming anaerobic conditions, unless otherwise stated. 4-Hydroxybutyrate is assumed to exit in E. coli via proton symport, as is the case with most organic acids. It is also possible that GBL is secreted by simple diffusion, and in this case the energetics would be more favorable than in the case considered here. The impact of cofactor specificity (i.e., NADH or NADPH-dependence) of the participating enzymes on the maximum yield and energetics of each pathway also was investigated.
[0143] The results from the analysis are shown in Tables 3 A-C. From an energetic and yield standpoint, the succinate to 4-HB pathway is the most promising provided that the thermodynamic concerns raised in Example I can be overcome. Specifically, the calculations reveal that the maximum theoretical yield of 4-HB from glucose is 1.33 mol/mol (0.77 g/g; 0.89 Cmol/Cmol) assuming the succinate to 4-HB pathway is functional. In addition, the anaerobic production of 4-HB via succinate would result in the net production of either 1.8, 1.5, or 1.1 mol of ATP per glucose depending upon the assumed cofactor specificity of the participating enzymes. These energetic yields are comparable to the 2.0 ATP per glucose that can be obtained via substrate level phosphorylation by the production of ethanol or lactate suggesting the potential for anaerobic homo-4-HB production in E. coli.
[0144] The succinyl-CoA route to 4-HB is the second most promising pathway when considering maximum yield and energetics. A 1.33 mol/mol yield of 4-HB is achievable in E. coli if at least one of the pathway steps is assumed NADH-dependant. However, because this pathway requires the formation of succinyl-CoA, its energetic yield is considerably lower than that of the succinate pathway. An oxygen requirement is anticipated at high 4-HB yields if both the CoA-dependant succinic semialdehyde dehydrogenase and 4-HB dehydrogenase steps are assumed NADPH-dependant. In this case, the production of 4-HB at the maximum yield would result in no net ATP gain and could not support the energetic maintenance demands needed for E. coli survival. Thus, some energy would have to originate from oxidative phosphorylation to enable homo-fermentative 4-HB production. The alpha-ketoglutarate pathway toward 4-HB is the least favorable of the three potential routes with a maximum achievable yield of 1.0 mol 4-HB per mol of glucose. In addition to the lower maximum yield, this pathway requires the utilization of 1.5 moles of oxygen per mol of glucose converted to 4-HB. The energetics of this pathway are unaffected by the assumed cofactor specificity of 4-HB dehydrogenase.
TABLE-US-00003 TABLE 3 The overall substrate conversion stoichiometry to 4-HB assuming the A) succinate, B) succinyl-CoA, or C) alpha-ketoglutarate production routes are functional in E. coli. Glucose and oxygen are taken up while all other molecules are produced. A) Succinate Pathway Cofactor 1 NADH step Specificity 2 NADH steps 1 NADPH step 2 NADPH steps Glucose -1.000 -1.000 -1.000 Oxygen 0.000 0.000 0.000 Protons 1.333 1.333 1.333 4HB 1.333 1.333 1.333 CO2 0.667 0.667 0.667 H2O 0.667 0.667 0.667 ATP 1.800 1.510 1.097 B) Succinyl-CoA Pathway Cofactor 1 NADH step Specificity 2 NADH steps 1 NADPH step 2 NADPH steps 2 NADPH steps Glucose -1.000 -1.000 -1.000 -1.000 Oxygen 0.000 0.000 -0.036 0.000 Protons 1.333 1.333 1.325 1.294 4HB 1.333 1.333 1.325 1.294 CO2 0.667 0.667 0.698 0.082 H2O 0.667 0.667 0.698 0.470 ATP 0.467 0.177 0.000 0.000 C) Alpha-ketoglutarate Pathway Cofactor Specificity 1 NADH step 1 NADPH step Glucose -1.000 -1.000 Oxygen -1.500 -1.500 Protons 1.000 1.000 4HB 1.000 1.000 CO2 2.000 2.000 H2O 2.000 2.000 ATP 5.500 5.500
[0145] In order to corroborate the computational predictions proposed in this report, the strains expressing a complete pathway to 4-HB can be constructed and tested. Corroboration is performed with both E. coli (Example II) and S. cerevisiae (Example III). In E. coli, the relevant genes are expressed in a synthetic operon behind an inducible promoter on a medium- or high-copy plasmid; for example the P.sub.BAD promoter which is induced by arabinose, on a plasmid of the pBAD series (Guzman et al., J. Bacteriol. 177:4121-4130 (1995)). In S. cerevisiae, genes are integrated into the chromosome behind the PDC1 promoter, replacing the native pyruvate carboxylase gene. It has been reported that this results in higher expression of foreign genes than from a plasmid (Ishida et al., Appl. Environ. Microbiol. 71:1964-1970 (2005)), and will also ensure expression during anaerobic conditions.
[0146] Cells containing the relevant constructs are grown in minimal media containing glucose, with addition of arabinose in the case of E. coli containing genes expressed under the P.sub.BAD promoter. Periodic samples are taken for both gene expression and enzyme activity analysis. Enzyme activity assays are performed on crude cell extracts using procedures well known in the art. Alternatively, assays based on the oxidation of NAD(P)H, which is produced in all dehydrogenase reaction steps and detectable by spectrophotometry can be utilized. In addition, antibodies can be used to detect the level of particular enzymes. In lieu of or in addition to enzyme activity measurements, RNA can be isolated from parallel samples and transcript of the gene of interest measured by reverse transcriptase PCR. Any constructs lacking detectable transcript expression are reanalyzed to ensure the encoding nucleic acids are harbored in an expressible form. Where transcripts are detected, this result indicates either a lack of translation or production of inactive enzyme. A variety of methods well known in the art can additionally be employed, such as codon optimization, engineering a strong ribosome binding site, use of a gene from a different species, and prevention of N-glycosylation (for expression of bacterial enzymes in yeast) by conversion of Asn residues to Asp. Once all required enzyme activities are detected, the next step is to measure the production of 4-HP in vivo. Triplicate shake flask cultures are grown either anaerobically or microaerobically, depending on the conditions required (see above), and periodic samples taken. Organic acids present in the culture supernatants are analyzed by HPLC using the Aminex AH-87X column. The elution time of 4-HB will be determined using a standard purchased from a chemical supplier.
[0147] The CoA-independent pathway can be implemented and tested for corroboration. In this case, the genes overexpressed are the native succinic semialdehyde dehydrogenase from each organism, and the 4-hydroxybutanoate dehydrogenase from Ralstonia eutropha. Once both enzyme activities are detected as discussed above, the strains are tested for 4-HB production. Corroboration also can be obtained from implementing the CoA-dependent pathway. The CoA-dependent succinic semialdehyde dehydrogenase and the 4-hydroxybutanoate dehydrogenase from Clostridium kluyveri are expressed as described above. In addition, overexpression of the native succinyl-CoA synthetase also can be performed, to funnel more succinate into the heterologous pathway. Finally, if 4-HB production is unfavorable, different culture conditions can be tested, such as a change in oxygenation status which can manipulate the NAD(P)H/NAD(P) ratio.
Example III
Production of 4-Hydroxybutanoic Acid in Yeast
[0148] This Example describes the biosynthetic yields for 4-hydroxybutanoic acid resulting from each biochemical pathway in S. cerevisiae.
[0149] In this section, the maximum theoretical yields of 4-HB from glucose are calculated assuming that each of the three metabolic pathways depicted in FIG. 2 are functional in S. cerevisiae. A genome-scale metabolic model of S. cerevisiae, similar to the one described in Forster et al. (Genome Res. 13:244-253 (2003)) was used as the basis for the analysis. The energetic gain of each maximum yielding pathway is calculated assuming anaerobic conditions unless otherwise stated. 4-hydroxybutyrate is assumed to exit S. cerevisiae via proton symport, as is the case with most organic acids. The impact of cofactor specificity (i.e., NADH or NADPH-dependence) of the participating enzymes on the maximum yield and energetics of each pathway was also investigated.
[0150] The results from the analysis are shown in Tables 4 A-C. As with E. coli, the succinate to 4-HB pathway is the most promising provided that the thermodynamic concerns raised in Example I can be overcome. The calculations reveal that the maximum theoretical yield of 4-HB from glucose is 1.33 mol/mol (0.77 g/g; 0.89 Cmol/Cmol) in S. cerevisiae. In addition, the anaerobic production of 4-HB via succinate would result in the net production of either 1.4, 1.1, or 0.5 mol of ATP per glucose depending upon the assumed cofactor specificity of the participating enzymes.
[0151] The succinyl-CoA route to 4-HB is the second most favorable pathway. A maximum yield of 1.33 mol 4-HB/mol glucose is achievable in S. cerevisiae regardless of cofactor specificity. However, net energy generation at the maximum theoretical yield is possible only if both the CoA-dependant succinic semialdehyde dehydrogenase and 4-HB dehydrogenase steps are assumed to be NADH-dependant. If either step is NADPH-dependant, no net ATP will be gained from anaerobic 4-HB production and an alternate energy source (e.g., oxidative phosphorylation) would be required to support cell growth and maintenance. The alpha-ketoglutarate route toward 4-HB is the least favorable of the three potential pathways in S. cerevisiae although the maximum yield of 1.1-1.2 mol 4-HB per mol glucose is slightly higher than was found in E. coli. Nevertheless, this pathway requires an oxygen uptake of 0.8-0.9 mol oxygen per mol glucose to become energetically neutral.
TABLE-US-00004 TABLE 4 The overall substrate conversion stoichiometry to 4-HB in S. cerevisiae., assuming the A) succinate, B) succinyl-CoA, or C) alpha-ketoglutarate production routes are functional in S. cerevisiae. Glucose and oxygen are taken up while all other molecules are produced. Cofactor 1 NADH step Specificity 2 NADH steps 1 NADPH step 2 NADPH steps A) Succinate Pathway Glucose -1.000 -1.000 -1.000 Oxygen 0.000 0.000 0.000 Protons 1.333 1.333 1.333 4HB 1.333 1.333 1.333 CO2 0.667 0.667 0.667 H2O 0.667 0.667 0.667 ATP 1.444 1.067 0.533 B) Succinyl-CoA Pathway Glucose -1.000 -1.000 -1.000 Oxygen 0.000 0.000 0.000 Protons 1.333 1.333 1.333 4HB 1.333 1.333 1.333 CO2 0.667 0.667 0.667 H2O 0.667 0.667 0.667 ATP 0.533 0.000 0.000 C) Alpha-ketoglutarate Pathway Cofactor Specificity 1 NADH step 1 NADPH step Glucose -1.000 -1.000 Oxygen -0.785 -0.879 Protons 1.159 1.138 4HB 1.159 1.138 CO2 1.364 1.448 H2O 1.364 1.448 ATP 0.000 0.000
Example IV
Biosynthesis of 4-Hydroxybutanoic Acid, .gamma.-Butyrolactone and 1,4-Butanediol
[0152] This Example describes the biosynthetic production of 4-hydroxybutanoic acid, .gamma.-butyrolactone and 1,4-butanediol using fermentation and other bioprocesses.
[0153] Methods for the integration of the 4-HB fermentation step into a complete process for the production of purified GBL, 1,4-butanediol (BDO) and tetrahydrofuran (THF) are described below. Since 4-HB and GBL are in equilibrium, the fermentation broth will contain both compounds. At low pH this equilibrium is shifted to favor GBL. Therefore, the fermentation can operate at pH 7.5 or less. After removal of biomass, the product stream enters into a separation step in which GBL is removed and the remaining stream enriched in 4-HB is recycled. Finally, GBL is distilled to remove any impurities. The process operates in one of three ways: 1) fed-batch fermentation and batch separation; 2) fed-batch fermentation and continuous separation; 3) continuous fermentation and continuous separation. The first two of these modes are shown schematically in FIG. 4. The integrated fermentation procedures described below also are used for the BDO producing cells of the invention for biosynthesis of BDO and subsequent BDO family products.
[0154] Fermentation Protocol to Produce 4-HB/GBL (Batch):
[0155] The production organism is grown in a 10 L bioreactor sparged with an N.sub.2/CO.sub.2 mixture, using 5 L broth containing 5 g/L potassium phosphate, 2.5 g/L ammonium chloride, 0.5 g/L magnesium sulfate, and 30 g/L corn steep liquor, and an initial glucose concentration of 20 g/L. As the cells grow and utilize the glucose, additional 70% glucose is fed into the bioreactor at a rate approximately balancing glucose consumption. The temperature of the bioreactor is maintained at 30 degrees C. Growth continues for approximately 24 hours, until 4-HB reaches a concentration of between 20-200 g/L, with the cell density being between 5 and 10 g/L. The pH is not controlled, and will typically decrease to pH 3-6 by the end of the run. Upon completion of the cultivation period, the fermenter contents are passed through a cell separation unit (e.g., centrifuge) to remove cells and cell debris, and the fermentation broth is transferred to a product separations unit. Isolation of 4-HB and/or GBL would take place by standard separations procedures employed in the art to separate organic products from dilute aqueous solutions, such as liquid-liquid extraction using a water immiscible organic solvent (e.g., toluene) to provide an organic solution of 4-HB/GBL. The resulting solution is then subjected to standard distillation methods to remove and recycle the organic solvent and to provide GBL (boiling point 204-205.degree. C.) which is isolated as a purified liquid.
[0156] Fermentation Protocol to Produce 4-HB/GBL (Fully Continuous):
[0157] The production organism is first grown up in batch mode using the apparatus and medium composition described above, except that the initial glucose concentration is 30-50 g/L. When glucose is exhausted, feed medium of the same composition is supplied continuously at a rate between 0.5 L/hr and 1 L/hr, and liquid is withdrawn at the same rate. The 4-HB concentration in the bioreactor remains constant at 30-40 g/L, and the cell density remains constant between 3-5 g/L. Temperature is maintained at 30 degrees C., and the pH is maintained at 4.5 using concentrated NaOH and HCl, as required. The bioreactor is operated continuously for one month, with samples taken every day to assure consistency of 4-HB concentration. In continuous mode, fermenter contents are constantly removed as new feed medium is supplied. The exit stream, containing cells, medium, and products 4-HB and/or GBL, is then subjected to a continuous product separations procedure, with or without removing cells and cell debris, and would take place by standard continuous separations methods employed in the art to separate organic products from dilute aqueous solutions, such as continuous liquid-liquid extraction using a water immiscible organic solvent (e.g., toluene) to provide an organic solution of 4-HB/GBL. The resulting solution is subsequently subjected to standard continuous distillation methods to remove and recycle the organic solvent and to provide GBL (boiling point 204-205.degree. C.) which is isolated as a purified liquid.
[0158] GBL Reduction Protocol:
[0159] Once GBL is isolated and purified as described above, it will then be subjected to reduction protocols such as those well known in the art (references cited) to produce 1,4-butanediol or tetrahydrofuran (THF) or a mixture thereof. Heterogeneous or homogeneous hydrogenation catalysts combined with GBL under hydrogen pressure are well known to provide the products 1,4-butanediol or tetrahydrofuran (THF) or a mixture thereof. It is important to note that the 4-HB/GBL product mixture that is separated from the fermentation broth, as described above, may be subjected directly, prior to GBL isolation and purification, to these same reduction protocols to provide the products 1,4-butanediol or tetrahydrofuran or a mixture thereof. The resulting products, 1,4-butanediol and THF are then isolated and purified by procedures well known in the art.
[0160] Fermentation and Hydrogenation Protocol to Produce BDO or THF Directly (Batch):
[0161] Cells are grown in a 10 L bioreactor sparged with an N.sub.2/CO.sub.2 mixture, using 5 L broth containing 5 g/L potassium phosphate, 2.5 g/L ammonium chloride, 0.5 g/L magnesium sulfate, and 30 g/L corn steep liquor, and an initial glucose concentration of 20 g/L. As the cells grow and utilize the glucose, additional 70% glucose is fed into the bioreactor at a rate approximately balancing glucose consumption. The temperature of the bioreactor is maintained at 30 degrees C. Growth continues for approximately 24 hours, until 4-HB reaches a concentration of between 20-200 g/L, with the cell density being between 5 and 10 g/L. The pH is not controlled, and will typically decrease to pH 3-6 by the end of the run. Upon completion of the cultivation period, the fermenter contents are passed through a cell separation unit (e.g., centrifuge) to remove cells and cell debris, and the fermentation broth is transferred to a reduction unit (e.g., hydrogenation vessel), where the mixture 4-HB/GBL is directly reduced to either 1,4-butanediol or THF or a mixture thereof. Following completion of the reduction procedure, the reactor contents are transferred to a product separations unit. Isolation of 1,4-butanediol and/or THF would take place by standard separations procedures employed in the art to separate organic products from dilute aqueous solutions, such as liquid-liquid extraction using a water immiscible organic solvent (e.g., toluene) to provide an organic solution of 1,4-butanediol and/or THF. The resulting solution is then subjected to standard distillation methods to remove and recycle the organic solvent and to provide 1,4-butanediol and/or THF which are isolated as a purified liquids.
[0162] Fermentation and Hydrogenation Protocol to Produce BDO or THF Directly (Fully Continuous):
[0163] The cells are first grown up in batch mode using the apparatus and medium composition described above, except that the initial glucose concentration is 30-50 g/L. When glucose is exhausted, feed medium of the same composition is supplied continuously at a rate between 0.5 L/hr and 1 L/hr, and liquid is withdrawn at the same rate. The 4-HB concentration in the bioreactor remains constant at 30-40 g/L, and the cell density remains constant between 3-5 g/L. Temperature is maintained at 30 degrees C., and the pH is maintained at 4.5 using concentrated NaOH and HCl, as required. The bioreactor is operated continuously for one month, with samples taken every day to assure consistency of 4-HB concentration. In continuous mode, fermenter contents are constantly removed as new feed medium is supplied. The exit stream, containing cells, medium, and products 4-HB and/or GBL, is then passed through a cell separation unit (e.g., centrifuge) to remove cells and cell debris, and the fermentation broth is transferred to a continuous reduction unit (e.g., hydrogenation vessel), where the mixture 4-HB/GBL is directly reduced to either 1,4-butanediol or THF or a mixture thereof. Following completion of the reduction procedure, the reactor contents are transferred to a continuous product separations unit. Isolation of 1,4-butanediol and/or THF would take place by standard continuous separations procedures employed in the art to separate organic products from dilute aqueous solutions, such as liquid-liquid extraction using a water immiscible organic solvent (e.g., toluene) to provide an organic solution of 1,4-butanediol and/or THF. The resulting solution is then subjected to standard continuous distillation methods to remove and recycle the organic solvent and to provide 1,4-butanediol and/or THF which are isolated as a purified liquids.
[0164] Fermentation Protocol to Produce BDO Directly (Batch):
[0165] The production organism is grown in a 10 L bioreactor sparged with an N.sub.2/CO.sub.2 mixture, using 5 L broth containing 5 g/L potassium phosphate, 2.5 g/L ammonium chloride, 0.5 g/L magnesium sulfate, and 30 g/L corn steep liquor, and an initial glucose concentration of 20 g/L. As the cells grow and utilize the glucose, additional 70% glucose is fed into the bioreactor at a rate approximately balancing glucose consumption. The temperature of the bioreactor is maintained at 30 degrees C. Growth continues for approximately 24 hours, until BDO reaches a concentration of between 20-200 g/L, with the cell density generally being between 5 and 10 g/L. Upon completion of the cultivation period, the fermenter contents are passed through a cell separation unit (e.g., centrifuge) to remove cells and cell debris, and the fermentation broth is transferred to a product separations unit. Isolation of BDO would take place by standard separations procedures employed in the art to separate organic products from dilute aqueous solutions, such as liquid-liquid extraction using a water immiscible organic solvent (e.g., toluene) to provide an organic solution of BDO. The resulting solution is then subjected to standard distillation methods to remove and recycle the organic solvent and to provide BDO (boiling point 228-229.degree. C.) which is isolated as a purified liquid.
[0166] Fermentation Protocol to Produce BDO Directly (Fully Continuous):
[0167] The production organism is first grown up in batch mode using the apparatus and medium composition described above, except that the initial glucose concentration is 30-50 g/L.
[0168] When glucose is exhausted, feed medium of the same composition is supplied continuously at a rate between 0.5 L/hr and 1 L/hr, and liquid is withdrawn at the same rate. The BDO concentration in the bioreactor remains constant at 30-40 g/L, and the cell density remains constant between 3-5 g/L. Temperature is maintained at 30 degrees C., and the pH is maintained at 4.5 using concentrated NaOH and HCl, as required. The bioreactor is operated continuously for one month, with samples taken every day to assure consistency of BDO concentration. In continuous mode, fermenter contents are constantly removed as new feed medium is supplied. The exit stream, containing cells, medium, and the product BDO, is then subjected to a continuous product separations procedure, with or without removing cells and cell debris, and would take place by standard continuous separations methods employed in the art to separate organic products from dilute aqueous solutions, such as continuous liquid-liquid extraction using a water immiscible organic solvent (e.g., toluene) to provide an organic solution of BDO. The resulting solution is subsequently subjected to standard continuous distillation methods to remove and recycle the organic solvent and to provide BDO (boiling point 228-229.degree. C.) which is isolated as a purified liquid (mpt 20.degree. C.).
Example V
In Silico Derived Knockout Strategies for Growth-Coupled Production of 1,4-Butanediol in Escherichia coli
[0169] This example describes the in silico design of knockout strategies to generate growth-coupled production of 1,4-butanediol in E. coli.
[0170] The strategy for generating growth-coupled production of BDO first involves the demonstration of a functional pathway which is subsequently optimized through adaptive evolution and targeted gene insertions, deletions, and overexpressions. Described below in more detail are strain-engineering strategies identified via OptKnock for generating growth-coupled BDO production strains of Escherichia coli. All implementations of OptKnock assume that sufficient activities of all enzymes outlined in FIG. 6 are available to E. coli under all conditions. In addition, all reduction steps in this pathway are assumed to be NADH-dependent although many of the designs are expected to be applicable regardless of cofactor specificity.
[0171] The BDO yield potential of the biochemical pathways to 1,4-butanediol in E. coli is detailed below. Three conditions were used, anaerobic, anaerobic+nitrate addition, and aerobic. The maximum theoretical yields for each scenario assuming that the production of BDO must be energetically neutral are shown in Table 5. Table 5 shows the maximum theoretical yields of 1,4-butanediol (BDO) for five sets of environmental conditions: 1) aerobic respiration, 2) anaerobic fermentation with acetate co-production, 3) anaerobic fermentation with ethanol co-production, 4) nitrate respiration leading to nitrite formation, and 5) nitrate respiration leading to ammonia formation. Negative values indicate metabolites taken up; positive values indicate metabolites secreted. Molar units are assumed along with pathway energetic neutrality. The maximum theoretical yield without assuming energetic neutrality is 1.091 mol/mol (0.545 g/g) glucose for all cases. The highest yield is obtained under aerobic conditions where enough ATP to render the pathway energetically neutral can be generated with minimal loss of carbon through respiration. In the anaerobic case, the yield drops as ATP is made via substrate level phosphorylation and either acetate or ethanol is made as a byproduct. Controlled nitrate addition can provide nearly the same yield as oxygen, the exact amount depending upon whether further reduction of nitrite to ammonia occurs.
TABLE-US-00005 TABLE 5 Maximum theoretical yields of 1,4-butanediol (BDO) for five sets of environmental conditions. Anerobic Acetate Ethanol Copro- Copro- Nitrate to Nitrate to Aerobic duction duction Nitrite Ammonia Glucose -1.000 -1.000 -1.000 -1.000 1.000 Oxygen -0.068 Nitrate -0.147 -0.092 H.sup.+ 0.144 -0.184 H.sub.2O 0.607 0.519 0.498 0.612 0.621 CO.sub.2 1.686 1.558 1.668 1.690 1.770 Nitrite 0.147 Ammonia 0.092 Acetate 0.144 Ethanol 0.173 BDO 1.079 1.039 0.997 1.078 1.058
[0172] If phosphoenolpyruvate (PEP) carboxykinase is assumed to operate in the direction of phosphoenolpyruvate to oxaloacetate, BDO production becomes an energy-generating endeavor and the maximum theoretical yield for all scenarios becomes 0.545 g/g with CO.sub.2 as the only byproduct. PEP carboxykinase is known to produce oxaloacetate from PEP in rumen bacteria such as Mannheimia succiniciproducens (Hong et al., Nat. Biotechnol. 22:1275-1281 (2004)). However, the role of PEP carboxykinase in producing oxaloacetate in E. coli is believed to be minor as compared to PEP carboxylase possibly due to the higher Km for bicarbonate of PEP carboxykinase (Kim et al., Appl. Environ. Microbiol. 70:1238-1241 (2004)). Nevertheless, activity of the native E. coli PEP carboxykinase from PEP towards oxaloacetate has been recently demonstrated in ppc mutants of K-12 (Kwon et al., J. Microbiol. Biotechnol. 16:1448-1452 (2006)).
[0173] In more detail, the designs identified for increasing BDO production in E. coli are described below. A non-growth associated energetic maintenance requirement of 7.6 mmol/gDW/hr was assumed along with a maximum specific glucose uptake rate of 20 mmol/gDW/hr. BDO was assumed to be exported via diffusion. Knockout strategies were identified assuming that 1) PEP carboxykinase is irreversible and functions only to convert oxaloacetate to PEP and 2) PEP carboxykinase is reversible. For both cases, the OptKnock code described in Burgard et al. (Biotechnol. Bioeng. 84:647-657 (2003)) was modified to allow for limited, unlimited or no nitrate respiration. Adding the possibility of nitrate respiration serves to increase the maximum theoretical yield of BDO when PEP carboxykinase is assumed irreversible and also, in some cases, enables the selection of knockout strategies that otherwise would not be selected under anaerobic conditions due to unfavorable energetics. Specifically, six nitrate uptake reactions were added to the E. coli network with lower bounds of 0, -2, -5, -10, -20, or -1000 mmol/gDW/hr (the negative values signify metabolite uptake in the stoichiometric model). The dual constraints of OptKnock were adjusted accordingly and an additional constraint was added that allowed only one nitrate uptake reaction to be active at a time. OptKnock selects the optimum amount of nitrate respiration for each identified knockout strategy. Finally, all simulations assume that the reaction catalyzed by adhE in E. coli (that is, acetyl-CoA+NADH.fwdarw.ethanol+NAD) has been removed. Because acetyl-CoA is a necessary intermediate for BDO production, nearly all OptKnock designs would include this deletion anyway to prevent ethanol production from competing with BDO production. Including this deletion a priori was done to lower the CPU time of the computational procedure.
[0174] The knockout strategies derived by OptKnock are given in Tables 6 and 7 assuming PEP carboxykinase to be irreversible and reversible, respectively. In this description, the knockout strategies are listed by reaction abbreviations in all capital letters for simplicity. The corresponding genes that would have to be knocked out to prevent a particular reaction from occurring in E. coli are provided in Table 8 along with the reaction stoichiometry. The metabolite names corresponding to Table 8 are listed in Table 9.
TABLE-US-00006 TABLE 6 Knockout strategies derived by OptKnock, assuming PEP carboxykinase to be irreversible. Metabolic Transformations Targeted for Removal .dagger., .dagger-dbl., # BDO BIO AC ALA CO2 FOR GLC GLY H+ 1 ADHEr 5.67 0.57 25.42 0.00 -5.74 28.42 -20.00 0.00 57.91 2 ADHEr, 9.95 0.50 22.31 0.00 14.90 0.00 -20.00 0.00 25.85 PFLi 3 ADHEr, 6.92 0.52 25.03 0.00 -0.07 20.82 -20.00 0.00 49.54 NADH6 4 ADHEr, 6.55 0.56 23.75 0.00 -3.54 26.67 -20.00 0.00 54.38 THD2 5 ADHEr, 6.21 0.30 28.99 0.00 -5.98 30.57 -20.00 0.00 61.72 PGI 6 ADHEr, 6.13 0.31 29.10 0.00 -6.17 30.70 -20.00 0.00 61.98 PFK 7 ADHEr, 6.13 0.31 29.10 0.00 -6.17 30.70 -20.00 0.00 61.98 FBA 8 ADHEr, 6.15 0.61 23.38 0.00 -3.10 26.55 -20.00 0.00 54.24 ATPS4r 9 ADHEr, 6.13 0.31 29.10 0.00 -6.17 30.70 -20.00 0.00 61.98 TPI 10 ADHEr, 5.85 0.47 26.89 0.00 -5.92 29.33 -20.00 0.00 59.54 SUCD4 11 ADHEr, 5.78 0.57 25.20 0.00 -5.45 28.19 -20.00 0.00 57.44 RPE 12 ADHEr, 5.78 0.51 26.33 0.00 -5.85 28.99 -20.00 0.00 58.92 GLCpts 13 ADHEr, 5.77 0.52 26.20 0.00 -5.84 28.90 -20.00 0.00 58.78 GLUDy 14 ADHEr, 5.73 0.57 25.30 0.00 -5.59 28.30 -20.00 0.00 57.67 TAL 15 ADHEr, 5.73 0.54 25.90 0.00 -5.80 28.72 -20.00 0.00 58.44 MDH 16 ADHEr, 5.73 0.54 25.90 0.00 -5.80 28.72 -20.00 0.00 58.44 FUM 17 ADHEr, 5.68 0.57 25.48 0.00 -5.75 28.46 -20.00 0.00 57.99 CBMK2 18 ADHEr, 11.82 0.22 18.07 0.00 8.10 19.23 -20.00 0.00 38.88 HEX1, PGI 19 ADHEr, 11.82 0.22 18.05 0.00 8.10 19.22 -20.00 0.00 38.86 EDA, PGI 20 ADHEr, 10.87 0.20 25.96 0.00 16.29 0.00 -20.00 0.00 27.38 PFLi, PGI 21 ADHEr, 10.83 0.20 26.02 0.00 16.23 0.00 -20.00 0.00 27.46 FBA, PFLi 22 ADHEr, 10.83 0.20 26.02 0.00 16.23 0.00 -20.00 0.00 27.46 PFLi, TPI 23 ADHEr, 10.83 0.20 26.02 0.00 16.23 0.00 -20.00 0.00 27.46 PFK, PFLi 24 ADHEr, 10.49 0.49 21.02 0.00 15.71 0.00 -20.00 0.00 24.49 PFLi, THD2 25 ADHEr, 10.15 0.43 23.16 0.00 15.20 0.00 -20.00 0.00 26.22 GLCpts, PFLi 26 ADHEr, 10.10 0.45 22.94 0.00 15.12 0.00 -20.00 0.00 26.12 GLUDy, PFLi 27 ADHEr, 10.02 0.50 22.14 0.00 15.00 0.00 -20.00 0.00 25.67 PFLi, RPE 28 ADHEr, 9.99 0.50 22.22 0.00 14.95 0.00 -20.00 0.00 25.76 PFLi, TAL 29 ADHEr, 9.96 0.49 22.36 0.00 14.92 0.00 -20.00 0.00 25.87 CBMK2, PFLi 30 ADHEr, 8.97 0.26 23.60 0.00 0.95 24.98 -20.00 0.00 50.46 ATPS4r, PGI 31 ADHEr, 8.62 0.48 23.04 0.00 6.28 13.23 -20.00 0.00 39.68 ATPS4r, NADH6 32 ADHEr, 8.26 0.26 27.70 0.00 3.99 16.78 -20.00 0.00 46.33 ATPS4r, TPI 33 ADHEr, 8.26 0.26 27.70 0.00 3.99 16.78 -20.00 0.00 46.33 ATPS4r, PFK 34 ADHEr, 8.26 0.26 27.70 0.00 3.99 16.78 -20.00 0.00 46.33 ATPS4r, FBA 35 ADHEr, 7.88 0.50 23.49 0.00 2.67 18.21 -20.00 0.00 45.24 NADH6, THD2 36 ADHEr, 7.86 0.49 22.44 0.00 0.04 23.43 -20.00 0.00 49.33 ATPS4r, FUM 37 ADHEr, 7.86 0.49 22.44 0.00 0.04 23.43 -20.00 0.00 49.33 ATPS4r, MDH 38 ADHEr, 7.58 0.23 28.71 0.00 0.18 22.36 -20.00 0.00 52.73 NADH6, PGI 39 ADHEr, 7.55 0.49 22.66 0.00 -1.33 25.25 -20.00 0.00 51.43 FUM, THD2 40 ADHEr, 7.55 0.49 22.66 0.00 -1.33 25.25 -20.00 0.00 51.43 MDH, THD2 41 ADHEr, 7.50 0.24 28.80 0.00 -0.03 22.54 -20.00 0.00 53.03 NADH6, PFK 42 ADHEr, 7.50 0.24 28.80 0.00 -0.03 22.54 -20.00 0.00 53.03 FBA, NADH6 43 ADHEr, 7.50 0.24 28.80 0.00 -0.03 22.54 -20.00 0.00 53.03 NADH6, TPI 44 ADHEr, 7.06 0.45 25.94 0.00 -0.06 21.24 -20.00 0.00 50.38 GLCpts, NADH6 45 ADHEr, 7.05 0.52 24.82 0.00 0.30 20.47 -20.00 0.00 48.96 NADH6, RPE 46 ADHEr, 7.02 0.47 25.73 0.00 -0.06 21.14 -20.00 0.00 50.18 GLUDy, NADH6 47 ADHEr, 7.01 0.47 25.63 0.00 -0.06 21.10 -20.00 0.00 50.09 FUM, NADH6 48 ADHEr, 7.01 0.47 25.63 0.00 -0.06 21.10 -20.00 0.00 50.09 MDH, NADH6 49 ADHEr, 6.98 0.52 24.92 0.00 0.12 20.64 -20.00 0.00 49.23 NADH6, TAL 50 ADHEr, 6.56 0.49 24.86 0.00 -3.91 27.44 -20.00 0.00 55.80 GLCpts, THD2 51 ADHEr, 6.56 0.50 24.67 0.00 -3.84 27.31 -20.00 0.00 55.57 GLUDy, THD2 52 ADHEr, 6.33 0.22 30.11 0.00 -6.16 31.28 -20.00 0.00 62.99 PGI, SUCD4 53 ADHEr, 6.28 0.26 29.59 0.00 -6.08 30.96 -20.00 0.00 62.41 GLCpts, PGI 54 ADHEr, 6.28 0.22 30.24 0.00 -6.31 31.41 -20.00 0.00 63.24 HEX1, PFK 55 ADHEr, 6.28 0.22 30.24 0.00 -6.31 31.41 -20.00 0.00 63.24 FBA, HEX1 56 ADHEr, 6.28 0.22 30.24 0.00 -6.31 31.41 -20.00 0.00 63.24 HEX1, TPI 57 ADHEr, 6.27 0.23 30.17 0.00 -6.30 31.37 -20.00 0.00 63.17 PFK, SUCD4 58 ADHEr, 6.27 0.23 30.17 0.00 -6.30 31.37 -20.00 0.00 63.17 FBA, SUCD4 59 ADHEr, 6.27 0.23 30.17 0.00 -6.30 31.37 -20.00 0.00 63.17 SUCD4, TPI 60 ADHEr, 6.26 0.27 29.44 0.00 -6.06 30.86 -20.00 0.00 62.23 GLUDy, PGI 61 ADHEr, 6.21 0.26 29.69 0.00 -6.24 31.07 -20.00 0.00 62.64 GLCpts, TPI 62 ADHEr, 6.21 0.26 29.69 0.00 -6.24 31.07 -20.00 0.00 62.64 GLCpts, PFK 63 ADHEr, 6.21 0.26 29.69 0.00 -6.24 31.07 -20.00 0.00 62.64 FBA, GLCpts 64 ADHEr, 6.20 0.30 29.01 0.00 -6.02 30.60 -20.00 0.00 61.77 PFK, RPE 65 ADHEr, 6.20 0.30 29.01 0.00 -6.02 30.60 -20.00 0.00 61.77 FBA, RPE 66 ADHEr, 6.20 0.30 29.01 0.00 -6.02 30.60 -20.00 0.00 61.77 RPE, TPI 67 ADHEr, 6.19 0.27 29.54 0.00 -6.23 30.98 -20.00 0.00 62.47 GLUDy, TPI 68 ADHEr, 6.19 0.27 29.54 0.00 -6.23 30.98 -20.00 0.00 62.47 FBA, GLUDy 69 ADHEr, 6.19 0.27 29.54 0.00 -6.23 30.98 -20.00 0.00 62.47 GLUDy, PFK 70 ADHEr, 6.17 0.31 29.05 0.00 -6.09 30.65 -20.00 0.00 61.87 TAL, TPI 71 ADHEr, 6.17 0.31 29.05 0.00 -6.09 30.65 -20.00 0.00 61.87 PFK, TAL 72 ADHEr, 6.17 0.31 29.05 0.00 -6.09 30.65 -20.00 0.00 61.87 FBA, TAL 73 ADHEr, 6.17 0.55 24.36 0.00 -3.40 27.23 -20.00 0.00 55.50 ATPS4r, GLUDy 74 ADHEr, 6.00 0.38 28.04 0.00 -6.05 30.04 -20.00 0.00 60.81 PYK, SUCD4 75 ADHEr, 5.97 0.40 27.84 0.00 -6.03 29.92 -20.00 0.00 60.59 GLCpts, SUCD4 76 ADHEr, 5.95 0.46 26.74 0.00 -5.68 29.16 -20.00 0.00 59.19 RPE, SUCD4 77 ADHEr, 5.94 0.42 27.58 0.00 -6.00 29.76 -20.00 0.00 60.30 FUM, GLUDy 78 ADHEr, 5.94 0.42 27.58 0.00 -6.00 29.76 -20.00 0.00 60.30 GLUDy, MDH 79 ADHEr, 5.94 0.42 27.54 0.00 -5.99 29.74 -20.00 0.00 60.27 GLUDy, SUCD4 80 ADHEr, 5.90 0.46 26.81 0.00 -5.79 29.24 -20.00 0.00 59.36 SUCD4, TAL 81 ADHEr, 5.89 0.51 26.14 0.00 -5.59 28.78 -20.00 0.00 58.51 GLCpts, RPE 82 ADHEr, 5.87 0.46 27.02 0.00 -5.93 29.42 -20.00 0.00 59.69 GLCpts, GLUDy 83 ADHEr, 5.85 0.47 26.83 0.00 -5.91 29.29 -20.00 0.00 59.47 GLCpts, MDH 84 ADHEr, 5.85 0.47 26.83 0.00 -5.91 29.29 -20.00 0.00 59.47 FUM, GLCpts 85 ADHEr, 5.87 0.52 26.00 0.00 -5.57 28.69 -20.00 0.00 58.35 GLUDy, RPE 86 ADHEr, 5.84 0.51 26.23 0.00 -5.72 28.88 -20.00 0.00 58.71 GLCpts, TAL 87 ADHEr, 5.84 0.54 25.70 0.00 -5.53 28.50 -20.00 0.00 58.02 FUM, RPE 88 ADHEr, 5.84 0.54 25.70 0.00 -5.53 28.50 -20.00 0.00 58.02 MDH, RPE 89 ADHEr, 5.82 0.47 26.84 0.00 -5.98 29.28 -20.00 0.00 59.61 MDH, PYK 90 ADHEr, 5.82 0.47 26.84 0.00 -5.98 29.28 -20.00 0.00 59.61 FUM, PYK 91 ADHEr, 5.79 0.54 25.79 0.00 -5.66 28.61 -20.00 0.00 58.22 MDH, TAL 92 ADHEr, 5.79 0.54 25.79 0.00 -5.66 28.61 -20.00 0.00 58.22
FUM, TAL 93 ADHEr, 5.82 0.52 26.09 0.00 -5.70 28.79 -20.00 0.00 58.55 GLUDy, TAL 94 ADHEr, 5.68 0.57 25.51 0.00 -5.76 28.48 -20.00 0.00 58.02 CBMK2, GLU5K 95 ADHEr, 5.68 0.57 25.51 0.00 -5.76 28.48 -20.00 0.00 58.02 CBMK2, G5SD 96 ADHEr, 5.68 0.57 25.51 0.00 -5.76 28.48 -20.00 0.00 58.02 ASNS2, CBMK2 97 ADHEr, 5.68 0.57 25.50 0.00 -5.75 28.47 -20.00 0.00 58.00 CBMK2, SO4t2 98 ADHEr, 5.68 0.57 25.48 0.00 -5.75 28.46 -20.00 0.00 57.99 CBMK2, HEX1 99 ADHEr, 14.76 0.16 16.11 0.00 22.13 0.00 -20.00 0.00 17.24 EDA, PFLi, PGI 100 ADHEr, 14.39 0.13 16.96 0.00 21.96 1.24 -20.00 0.00 19.11 EDA, NADH6, PGI 101 ADHEr, 12.91 0.12 12.29 0.00 -0.02 38.75 -20.00 0.00 51.90 FRD2, GLUDy, LDH_D 102 ADHEr, 12.89 0.13 12.25 0.00 -0.02 38.70 -20.00 0.00 51.85 FRD2, LDH_D, THD2 103 ADHEr, 12.62 0.11 12.68 0.00 0.33 39.20 -20.00 0.00 52.67 ACKr, ACS, PPC 104 ADHEr, 12.60 0.16 11.81 0.00 0.47 38.84 -20.00 0.00 51.79 GLUDy, LDH_D, PPC 105 ADHEr, 12.60 0.16 11.72 0.00 0.49 38.81 -20.00 0.00 51.71 LDH_D, PPC, THD2 106 ADHEr, 11.95 0.15 19.06 0.00 8.00 19.85 -20.00 0.00 39.98 ATPS4r, EDA, PGI 107 ADHEr, 11.95 0.15 19.03 0.00 8.00 19.83 -20.00 0.00 39.94 EDA, GLCpts, PGI 108 ADHEr, 11.86 0.20 18.37 0.00 8.07 19.42 -20.00 0.00 39.21 EDA, GLUDy, PGI 109 ADHEr, 11.85 0.20 18.38 0.00 8.05 19.43 -20.00 0.00 39.24 GLUDy, HEX1, PGI 110 ADHEr, 11.72 0.28 7.78 0.00 9.70 25.16 -20.00 0.00 37.14 ATPS4r, FRD2, LDH_D 111 ADHEr, 11.48 0.56 11.79 0.00 27.19 0.00 -20.00 0.00 15.78 ACKr, NADH6, PYK 112 ADHEr, 11.02 0.64 11.37 0.00 26.49 0.00 -20.00 0.00 15.94 ACKr, LDH_D, NADH6 113 ADHEr, 10.95 0.17 26.34 0.00 16.41 0.00 -20.00 0.00 27.55 GLCpts, PFLi, PGI 114 ADHEr, 10.93 0.18 26.24 0.00 16.38 0.00 -20.00 0.00 27.51 GLUDy, PFLi, PGI 115 ADHEr, 10.91 0.17 26.39 0.00 16.36 0.00 -20.00 0.00 27.62 FBA, GLCpts, PFLi 116 ADHEr, 10.91 0.17 26.39 0.00 16.36 0.00 -20.00 0.00 27.62 GLCpts, PFK, PFLi 117 ADHEr, 10.91 0.17 26.39 0.00 16.36 0.00 -20.00 0.00 27.62 GLCpts, PFLi, TPI 118 ADHEr, 10.89 0.18 26.30 0.00 16.33 0.00 -20.00 0.00 27.58 FBA, GLUDy, PFLi 119 ADHEr, 10.89 0.18 26.30 0.00 16.33 0.00 -20.00 0.00 27.58 GLUDy, PFK, PFLi 120 ADHEr, 10.89 0.18 26.30 0.00 16.33 0.00 -20.00 0.00 27.58 GLUDy, PFLi, TPI 121 ADHEr, 10.86 0.20 25.97 0.00 16.28 0.00 -20.00 0.00 27.40 FBA, PFLi, RPE 122 ADHEr, 10.86 0.20 25.97 0.00 16.28 0.00 -20.00 0.00 27.40 PFLi, RPE, TPI 123 ADHEr, 10.86 0.20 25.97 0.00 16.28 0.00 -20.00 0.00 27.40 PFK, PFLi, RPE 124 ADHEr, 10.85 0.20 26.00 0.00 16.25 0.00 -20.00 0.00 27.43 PFK, PFLi, TAL 125 ADHEr, 10.85 0.20 26.00 0.00 16.25 0.00 -20.00 0.00 27.43 PFLi, TAL, TPI 126 ADHEr, 10.85 0.20 26.00 0.00 16.25 0.00 -20.00 0.00 27.43 FBA, PFLi, TAL 127 ADHEr, 10.64 0.17 23.71 0.00 8.96 13.97 -20.00 0.00 38.90 ATPS4r, NADH6, PGI 128 ADHEr, 10.62 0.42 22.05 0.00 15.90 0.00 -20.00 0.00 25.05 GLCpts, PFLi, THD2 129 ADHEr, 10.59 0.44 21.77 0.00 15.85 0.00 -20.00 0.00 24.89 GLUDy, PFLi, THD2 130 ADHEr, 10.53 0.30 24.78 0.00 15.78 0.00 -20.00 0.00 26.92 LDH_D, PFLi, SUCD4 131 ADHEr, 10.31 0.38 23.85 0.00 15.45 0.00 -20.00 0.00 26.52 LDH_D, NADH6, PFLi 132 ADHEr, 10.30 0.35 21.88 0.00 12.39 8.06 -20.00 0.00 32.44 ATPS4r, LDH_D, SUCD4 133 ADHEr, 10.29 0.38 23.75 0.00 15.41 0.00 -20.00 0.00 26.47 FUM, PFLi, PYK 134 ADHEr, 10.29 0.38 23.75 0.00 15.41 0.00 -20.00 0.00 26.47 MDH, PFLi, PYK 135 ADHEr, 10.28 0.39 23.71 0.00 15.40 0.00 -20.00 0.00 26.46 GLCpts, GLUDy, PFLi 136 ADHEr, 10.21 0.43 23.02 0.00 15.29 0.00 -20.00 0.00 26.07 GLCpts, PFLi, RPE 137 ADHEr, 10.18 0.43 23.09 0.00 15.25 0.00 -20.00 0.00 26.14 GLCpts, PFLi, TAL 138 ADHEr, 10.16 0.45 22.79 0.00 15.22 0.00 -20.00 0.00 25.96 GLUDy, PFLi, RPE 139 ADHEr, 10.16 0.43 23.21 0.00 15.22 0.00 -20.00 0.00 26.24 CBMK2, GLCpts, PFLi 140 ADHEr, 10.15 0.43 23.14 0.00 15.19 0.00 -20.00 0.00 26.21 LDH_D, MDH, PFLi 141 ADHEr, 10.15 0.43 23.14 0.00 15.19 0.00 -20.00 0.00 26.21 FUM, LDH_D, PFLi 142 ADHEr, 10.13 0.45 22.86 0.00 15.17 0.00 -20.00 0.00 26.04 GLUDy, PFLi, TAL 143 ADHEr, 10.11 0.44 22.98 0.00 15.14 0.00 -20.00 0.00 26.14 CBMK2, GLUDy, PFLi 144 ADHEr, 10.04 0.49 22.20 0.00 15.02 0.00 -20.00 0.00 25.69 CBMK2, PFLi, RPE 145 ADHEr, 10.00 0.49 22.28 0.00 14.97 0.00 -20.00 0.00 25.78 CBMK2, PFLi, TAL 146 ADHEr, 9.96 0.49 22.35 0.00 14.91 0.00 -20.00 0.00 25.87 ASNS2, G5SD, PFLi 147 ADHEr, 9.96 0.49 22.35 0.00 14.91 0.00 -20.00 0.00 25.87 ASNS2, GLU5K, PFLi 148 ADHEr, 9.96 0.37 21.98 0.00 9.04 11.74 -20.00 0.00 36.36 ATPS4r, GLCpts, MDH 149 ADHEr, 9.96 0.37 21.98 0.00 9.04 11.74 -20.00 0.00 36.36 ATPS4r, FUM, GLCpts 150 ADHEr, 9.87 0.17 27.44 0.00 11.15 7.29 -20.00 0.00 35.96 ATPS4r, FBA, NADH6 151 ADHEr, 9.87 0.17 27.44 0.00 11.15 7.29 -20.00 0.00 35.96 ATPS4r, NADH6, TPI 152 ADHEr, 9.87 0.17 27.44 0.00 11.15 7.29 -20.00 0.00 35.96 ATPS4r, NADH6, PFK 153 ADHEr, 9.82 0.44 23.35 0.00 13.64 2.12 -20.00 0.00 28.60 ATPS4r, FUM, PGL 154 ADHEr, 9.82 0.44 23.35 0.00 13.64 2.12 -20.00 0.00 28.60 ATPS4r, MDH, PGDH 155 ADHEr, 9.82 0.44 23.35 0.00 13.64 2.12 -20.00 0.00 28.60 ATPS4r, FUM, G6PDHy 156 ADHEr, 9.82 0.44 23.35 0.00 13.64 2.12 -20.00 0.00 28.60 ATPS4r, FUM, PGDH 157 ADHEr, 9.82 0.44 23.35 0.00 13.64 2.12 -20.00 0.00 28.60 ATPS4r, MDH, PGL 158 ADHEr, 9.82 0.44 23.35 0.00 13.64 2.12 -20.00 0.00 28.60 ATPS4r, G6PDHy, MDH 159 ADHEr, 9.81 0.44 12.15 0.00 1.32 36.78 -20.00 0.00 52.07 ATPS4r, LDH_D, PPC 160 ADHEr, 9.77 0.44 23.32 0.00 13.29 2.67 -20.00 0.00 29.13 ATPS4r, FUM, TAL 161 ADHEr, 9.77 0.44 23.32 0.00 13.29 2.67 -20.00 0.00 29.13 ATPS4r, MDH, TAL 162 ADHEr, 9.76 0.42 22.90 0.00 13.62 4.00 -20.00 0.00 29.90 ATPS4r, GLCpts, NADH6 163 ADHEr, 9.72 0.44 23.30 0.00 12.97 3.18 -20.00 0.00 29.62 ATPS4r,
MDH, RPE 164 ADHEr, 9.72 0.44 23.30 0.00 12.97 3.18 -20.00 0.00 29.62 ATPS4r, FUM, RPE 165 ADHEr, 9.39 0.46 23.42 0.00 11.71 4.69 -20.00 0.00 31.40 ATPS4r, NADH6, PGDH 166 ADHEr, 9.39 0.46 23.42 0.00 11.71 4.69 -20.00 0.00 31.40 ATPS4r, NADH6, PGL 167 ADHEr, 9.39 0.46 23.42 0.00 11.71 4.69 -20.00 0.00 31.40 ATPS4r, G6PDHy, NADH6 168 ADHEr, 9.34 0.46 23.40 0.00 11.35 5.25 -20.00 0.00 31.94 ATPS4r, NADH6, TAL 169 ADHEr, 9.29 0.46 23.37 0.00 11.02 5.76 -20.00 0.00 32.44 ATPS4r, NADH6, RPE 170 ADHEr, 9.08 0.43 19.50 0.00 0.35 26.49 -20.00 0.00 49.05 G6PDHy, ME2, THD2 171 ADHEr, 9.08 0.43 19.50 0.00 0.35 26.49 -20.00 0.00 49.05 ME2, PGL, THD2 172 ADHEr, 8.99 0.47 0.00 0.00 6.58 33.74 -20.00 0.00 45.14 G6PDHy, PPC, THD2 173 ADHEr, 8.99 0.47 0.00 0.00 6.58 33.74 -20.00 0.00 45.14 PGL, PPC, THD2 174 ADHEr, 8.65 0.43 23.82 0.00 6.11 13.67 -20.00 0.00 40.57 ATPS4r, GLUDy, NADH6 175 ADHEr, 8.46 0.64 8.81 0.00 9.05 20.63 -20.00 0.00 37.61 ACKr, FRD2, LDH_D 176 ADHEr, 8.29 0.26 27.65 0.00 3.98 16.88 -20.00 0.00 46.36 ATPS4r, FBA, RPE 177 ADHEr, 8.29 0.26 27.65 0.00 3.98 16.88 -20.00 0.00 46.36 ATPS4r, RPE, TPI 178 ADHEr, 8.29 0.26 27.65 0.00 3.98 16.88 -20.00 0.00 46.36 ATPS4r, PFK, RPE 179 ADHEr, 8.28 0.23 28.10 0.00 3.79 17.22 -20.00 0.00 46.98 ATPS4r, GLUDy, PFK 180 ADHEr, 8.28 0.23 28.10 0.00 3.79 17.22 -20.00 0.00 46.98 ATPS4r, GLUDy, TPI 181 ADHEr, 8.28 0.23 28.10 0.00 3.79 17.22 -20.00 0.00 46.98 ATPS4r, FBA, GLUDy 182 ADHEr, 8.28 0.26 27.67 0.00 3.98 16.83 -20.00 0.00 46.35 ATPS4r, PFK, TAL 183 ADHEr, 8.28 0.26 27.67 0.00 3.98 16.83 -20.00 0.00 46.35 ATPS4r, TAL, TPI 184 ADHEr, 8.28 0.26 27.67 0.00 3.98 16.83 -20.00 0.00 46.35 ATPS4r, FBA, TAL 185 ADHEr, 8.16 0.28 23.57 0.00 -4.24 32.94 -20.00 0.00 58.48 ASPT, MDH, PYK 186 ADHEr, 8.00 0.71 13.36 0.00 7.12 24.68 -20.00 0.00 43.08 MDH, PGL, THD2 187 ADHEr, 8.00 0.71 13.36 0.00 7.12 24.68 -20.00 0.00 43.08 G6PDHy, MDH, THD2 188 ADHEr, 7.89 0.43 24.60 0.00 2.33 18.97 -20.00 0.00 46.64 GLCpts, NADH6, THD2 189 ADHEr, 7.89 0.45 24.31 0.00 2.42 18.77 -20.00 0.00 46.27 GLUDy, NADH6, THD2 190 ADHEr, 7.71 0.45 21.65 0.00 -4.12 31.31 -20.00 0.00 56.15 ASPT, LDH_D, MDH 191 ADHEr, 7.65 0.19 29.27 0.00 0.15 22.63 -20.00 0.00 53.28 GLCpts, NADH6, PGI 192 ADHEr, 7.64 0.40 24.07 0.00 -1.65 26.18 -20.00 0.00 53.11 LDH_D, SUCD4, THD2 193 ADHEr, 7.62 0.21 29.06 0.00 0.16 22.53 -20.00 0.00 53.08 GLUDy, NADH6, PGI 194 ADHEr, 7.62 0.32 7.79 0.00 -2.23 9.47 -20.00 0.00 37.34 ACKr, FUM, LDH_D 195 ADHEr, 7.59 0.20 29.34 0.00 -0.03 22.79 -20.00 0.00 53.53 FBA, GLCpts, NADH6 196 ADHEr, 7.59 0.20 29.34 0.00 -0.03 22.79 -20.00 0.00 53.53 GLCpts, NADH6, TPI 197 ADHEr, 7.62 0.32 7.79 0.00 -2.23 9.47 -20.00 0.00 37.34 ACKr, LDH_D, MDH 198 ADHEr, 7.59 0.20 29.34 0.00 -0.03 22.79 -20.00 0.00 53.53 GLCpts, NADH6, PFK 199 ADHEr, 7.58 0.43 23.76 0.00 -1.67 26.01 -20.00 0.00 52.82 GLCpts, MDH, THD2 200 ADHEr, 7.58 0.43 23.76 0.00 -1.67 26.01 -20.00 0.00 52.82 FUM, GLCpts, THD2 201 ADHEr, 7.57 0.23 28.73 0.00 0.14 22.39 -20.00 0.00 52.79 NADH6, PFK, RPE 202 ADHEr, 7.57 0.23 28.73 0.00 0.14 22.39 -20.00 0.00 52.79 FBA, NADH6, RPE 203 ADHEr, 7.57 0.23 28.73 0.00 0.14 22.39 -20.00 0.00 52.79 NADH6, RPE, TPI 204 ADHEr, 7.56 0.21 29.14 0.00 -0.03 22.70 -20.00 0.00 53.35 GLUDy, NADH6, TPI 205 ADHEr, 7.56 0.21 29.14 0.00 -0.03 22.70 -20.00 0.00 53.35 GLUDy, NADH6, PFK 206 ADHEr, 7.56 0.21 29.14 0.00 -0.03 22.70 -20.00 0.00 53.35 FBA, GLUDy, NADH6 207 ADHEr, 7.54 0.24 28.76 0.00 0.06 22.46 -20.00 0.00 52.90 NADH6, PFK, TAL 208 ADHEr, 7.54 0.24 28.76 0.00 0.06 22.46 -20.00 0.00 52.90 FBA, NADH6, TAL 209 ADHEr, 7.54 0.24 28.76 0.00 0.06 22.46 -20.00 0.00 52.90 NADH6, TAL, TPI 210 ADHEr, 7.24 0.58 7.55 0.00 52.00 0.00 -20.00 0.00 11.65 ACKr, AKGD, ATPS4r 211 ADHEr, 7.17 0.45 25.76 0.00 0.26 20.93 -20.00 0.00 49.87 GLCpts, NADH6, RPE 212 ADHEr, 7.15 0.40 26.55 0.00 -0.06 21.52 -20.00 0.00 50.95 FUM, GLCpts, NADH6 213 ADHEr, 7.15 0.40 26.55 0.00 -0.06 21.52 -20.00 0.00 50.95 GLCpts, MDH, NADH6 214 ADHEr, 7.15 0.41 26.54 0.00 -0.06 21.51 -20.00 0.00 50.94 GLCpts, GLUDy, NADH6 215 ADHEr, 7.15 0.41 26.54 0.00 -0.06 21.51 -20.00 0.00 50.94 FUM, NADH6, PYK 216 ADHEr, 7.15 0.41 26.54 0.00 -0.06 21.51 -20.00 0.00 50.94 MDH, NADH6, PYK 217 ADHEr, 7.14 0.58 7.46 0.00 16.55 35.80 -20.00 0.00 47.39 ACKr, ATPS4r, SUCOAS 218 ADHEr, 7.14 0.46 25.54 0.00 0.27 20.82 -20.00 0.00 49.66 GLUDy, NADH6, RPE 219 ADHEr, 7.13 0.47 25.45 0.00 0.27 20.78 -20.00 0.00 49.57 MDH, NADH6, RPE 220 ADHEr, 7.13 0.47 25.45 0.00 0.27 20.78 -20.00 0.00 49.57 FUM, NADH6, RPE 221 ADHEr, 7.12 0.45 25.84 0.00 0.11 21.08 -20.00 0.00 50.11 GLCpts, NADH6, TAL 222 ADHEr, 7.09 0.46 25.63 0.00 0.11 20.98 -20.00 0.00 49.91 GLUDy, NADH6, TAL 223 ADHEr, 7.07 0.47 25.53 0.00 0.11 20.93 -20.00 0.00 49.82 FUM, NADH6, TAL 224 ADHEr, 7.07 0.47 25.53 0.00 0.11 20.93 -20.00 0.00 49.82 MDH, NADH6, TAL 225 ADHEr, 6.93 0.51 25.11 0.00 -0.07 20.86 -20.00 0.00 49.62 CBMK2, GLU5K, NADH6 226 ADHEr, 6.93 0.51 25.11 0.00 -0.07 20.86 -20.00 0.00 49.62 CBMK2, G5SD, NADH6 227 ADHEr, 6.93 0.51 25.10 0.00 -0.07 20.86 -20.00 0.00 49.60 CBMK2, NADH6, SO4t2 228 ADHEr, 6.93 0.51 25.11 0.00 -0.07 20.86 -20.00 0.00 49.61 ASNS2, CBMK2, NADH6 229 ADHEr, 6.69 0.36 19.85 0.00 7.01 7.99 -20.00 0.00 38.27 ATPS4r, PYK,
SUCD4 230 ADHEr, 6.40 0.18 30.74 0.00 -6.26 31.69 -20.00 0.00 63.71 GLCpts, PGI, SUCD4 231 ADHEr, 6.38 0.19 30.56 0.00 -6.23 31.57 -20.00 0.00 63.51 FUM, GLUDy, PGI 232 ADHEr, 6.38 0.19 30.56 0.00 -6.23 31.57 -20.00 0.00 63.51 GLUDy, MDH, PGI 233 ADHEr, 6.37 0.20 30.45 0.00 -6.21 31.50 -20.00 0.00 63.38 GLUDy, PGI, SUCD4 234 ADHEr, 6.35 0.18 30.80 0.00 -6.37 31.76 -20.00 0.00 63.86 GLCpts, SUCD4, TPI 235 ADHEr, 6.35 0.18 30.80 0.00 -6.37 31.76 -20.00 0.00 63.86 GLCpts, PFK, SUCD4 236 ADHEr, 6.35 0.18 30.80 0.00 -6.37 31.76 -20.00 0.00 63.86 FBA, GLCpts, SUCD4 237 ADHEr, 6.32 0.22 30.18 0.00 -6.20 31.34 -20.00 0.00 63.09 HEX1, RPE, TPI 238 ADHEr, 6.32 0.22 30.18 0.00 -6.20 31.34 -20.00 0.00 63.09 FBA, HEX1, RPE 239 ADHEr, 6.32 0.22 30.18 0.00 -6.20 31.34 -20.00 0.00 63.09 HEX1, PFK, RPE 240 ADHEr, 6.32 0.20 30.62 0.00 -6.35 31.65 -20.00 0.00 63.67 FUM, GLUDy, TPI 241 ADHEr, 6.32 0.20 30.62 0.00 -6.35 31.65 -20.00 0.00 63.67 FBA, GLUDy, MDH 242 ADHEr, 6.32 0.20 30.62 0.00 -6.35 31.65 -20.00 0.00 63.67 FUM, GLUDy, PFK 243 ADHEr, 6.32 0.20 30.62 0.00 -6.35 31.65 -20.00 0.00 63.67 GLUDy, MDH, PFK 244 ADHEr, 6.32 0.20 30.62 0.00 -6.35 31.65 -20.00 0.00 63.67 GLUDy, MDH, TPI 245 ADHEr, 6.32 0.20 30.62 0.00 -6.35 31.65 -20.00 0.00 63.67 FBA, FUM, GLUDy 246 ADHEr, 6.32 0.23 29.99 0.00 -6.14 31.21 -20.00 0.00 62.85 GLCpts, GLUDy, PGI 247 ADHEr, 6.32 0.23 30.12 0.00 -6.19 31.30 -20.00 0.00 63.02 RPE, SUCD4, TPI 248 ADHEr, 6.32 0.23 30.12 0.00 -6.19 31.30 -20.00 0.00 63.02 FBA, RPE, SUCD4 249 ADHEr, 6.32 0.23 30.12 0.00 -6.19 31.30 -20.00 0.00 63.02 PFK, RPE, SUCD4 250 ADHEr, 6.31 0.20 30.55 0.00 -6.34 31.60 -20.00 0.00 63.59 GLUDy, HEX1, PFK 251 ADHEr, 6.31 0.20 30.55 0.00 -6.34 31.60 -20.00 0.00 63.59 GLUDy, HEX1, TPI 252 ADHEr, 6.31 0.20 30.55 0.00 -6.34 31.60 -20.00 0.00 63.59 FBA, GLUDy, HEX1 253 ADHEr, 6.31 0.20 30.51 0.00 -6.34 31.58 -20.00 0.00 63.55 FBA, GLUDy, SUCD4 254 ADHEr, 6.31 0.20 30.51 0.00 -6.34 31.58 -20.00 0.00 63.55 GLUDy, SUCD4, TPI 255 ADHEr, 6.31 0.20 30.51 0.00 -6.34 31.58 -20.00 0.00 63.55 GLUDy, PFK, SUCD4 256 ADHEr, 6.30 0.22 30.21 0.00 -6.25 31.37 -20.00 0.00 63.16 FBA, HEX1, TAL 257 ADHEr, 6.30 0.22 30.21 0.00 -6.25 31.37 -20.00 0.00 63.16 HEX1, PFK, TAL 258 ADHEr, 6.30 0.22 30.21 0.00 -6.25 31.37 -20.00 0.00 63.16 HEX1, TAL, TPI 259 ADHEr, 6.29 0.23 30.14 0.00 -6.24 31.33 -20.00 0.00 63.09 PFK, SUCD4, TAL 260 ADHEr, 6.29 0.23 30.14 0.00 -6.24 31.33 -20.00 0.00 63.09 SUCD4, TAL, TPI 261 ADHEr, 6.29 0.23 30.14 0.00 -6.24 31.33 -20.00 0.00 63.09 FBA, SUCD4, TAL 262 ADHEr, 6.28 0.22 6.40 0.00 -6.31 7.56 -20.00 0.00 39.41 ACKr, LDH_D, SUCD4 263 ADHEr, 6.26 0.26 29.61 0.00 -6.11 30.98 -20.00 0.00 62.45 GLCpts, RPE, TPI 264 ADHEr, 6.26 0.26 29.61 0.00 -6.11 30.98 -20.00 0.00 62.45 GLCpts, PFK, RPE 265 ADHEr, 6.26 0.26 29.61 0.00 -6.11 30.98 -20.00 0.00 62.45 FBA, GLCpts, RPE 266 ADHEr, 6.26 0.23 0.00 0.00 -6.29 1.22 -20.00 0.00 32.99 ACt6, LDH_D, MDH 267 ADHEr, 6.26 0.24 30.08 0.00 -6.29 31.31 -20.00 0.00 63.07 GLCpts, GLUDy, TPI 268 ADHEr, 6.26 0.23 0.00 0.00 -6.29 1.22 -20.00 0.00 32.99 ACt6, FUM, LDH_D 269 ADHEr, 6.26 0.24 30.08 0.00 -6.29 31.31 -20.00 0.00 63.07 FBA, GLCpts, GLUDy 270 ADHEr, 6.26 0.24 30.08 0.00 -6.29 31.31 -20.00 0.00 63.07 GLCpts, GLUDy, PFK 271 ADHEr, 6.25 0.27 29.46 0.00 -6.09 30.88 -20.00 0.00 62.28 GLUDy, RPE, TPI 272 ADHEr, 6.25 0.27 29.46 0.00 -6.09 30.88 -20.00 0.00 62.28 FBA, GLUDy, RPE 273 ADHEr, 6.25 0.27 29.46 0.00 -6.09 30.88 -20.00 0.00 62.28 GLUDy, PFK, RPE 274 ADHEr, 6.24 0.26 29.65 0.00 -6.17 31.02 -20.00 0.00 62.54 GLCpts, TAL, TPI 275 ADHEr, 6.24 0.26 29.65 0.00 -6.17 31.02 -20.00 0.00 62.54 FBA, GLCpts, TAL 276 ADHEr, 6.24 0.26 29.65 0.00 -6.17 31.02 -20.00 0.00 62.54 GLCpts, PFK, TAL 277 ADHEr, 6.22 0.27 29.50 0.00 -6.15 30.93 -20.00 0.00 62.37 GLUDy, PFK, TAL 278 ADHEr, 6.22 0.27 29.50 0.00 -6.15 30.93 -20.00 0.00 62.37 FBA, GLUDy, TAL 279 ADHEr, 6.22 0.27 29.50 0.00 -6.15 30.93 -20.00 0.00 62.37 GLUDy, TAL, TPI 280 ADHEr, 6.09 0.33 28.74 0.00 -6.13 30.48 -20.00 0.00 61.59 GLUDy, MDH, PYK 281 ADHEr, 6.09 0.33 28.74 0.00 -6.13 30.48 -20.00 0.00 61.59 FUM, GLUDy, PYK 282 ADHEr, 6.08 0.38 27.93 0.00 -5.86 29.92 -20.00 0.00 60.54 PYK, RPE, SUCD4 283 ADHEr, 6.08 0.34 28.65 0.00 -6.12 30.43 -20.00 0.00 61.49 GLUDy, PYK, SUCD4 284 ADHEr, 6.06 0.35 28.51 0.00 -6.10 30.34 -20.00 0.00 61.33 MDH, PYK, SUCD4 285 ADHEr, 6.06 0.35 28.51 0.00 -6.10 30.34 -20.00 0.00 61.33 FUM, PYK, SUCD4 286 ADHEr, 6.06 0.40 27.70 0.00 -5.83 29.77 -20.00 0.00 60.29 GLCpts, RPE, SUCD4 287 ADHEr, 6.05 0.35 28.44 0.00 -6.10 30.30 -20.00 0.00 61.26 GLCpts, GLUDy, MDH 288 ADHEr, 6.05 0.35 28.44 0.00 -6.10 30.30 -20.00 0.00 61.26 FUM, GLCpts, GLUDy 289 ADHEr, 6.04 0.36 28.39 0.00 -6.09 30.27 -20.00 0.00 61.21 GLCpts, GLUDy, SUCD4 290 ADHEr, 6.04 0.38 27.98 0.00 -5.95 29.98 -20.00 0.00 60.67 PYK, SUCD4, TAL 291 ADHEr, 6.03 0.41 27.44 0.00 -5.79 29.60 -20.00 0.00 59.98 GLUDy, MDH, RPE 292 ADHEr, 6.03 0.41 27.44 0.00 -5.79 29.60 -20.00 0.00 59.98 FUM, GLUDy, RPE 293 ADHEr, 6.02 0.42 27.40 0.00 -5.78 29.58 -20.00 0.00 59.94 GLUDy, RPE, SUCD4 294 ADHEr, 6.02 0.40 27.77 0.00 -5.92 29.84 -20.00 0.00 60.43 GLCpts, SUCD4, TAL 295 ADHEr, 5.99 0.42 27.50 0.00 -5.89 29.68 -20.00 0.00 60.14 GLUDy, MDH, TAL 296 ADHEr, 5.99 0.42 27.50 0.00 -5.89 29.68 -20.00 0.00 60.14 FUM, GLUDy, TAL 297 ADHEr, 5.98 0.42 27.47 0.00 -5.88 29.66 -20.00 0.00 60.10 GLUDy, SUCD4,
TAL 298 ADHEr, 5.96 0.46 26.84 0.00 -5.70 29.23 -20.00 0.00 59.31 GLCpts, GLUDy, RPE 299 ADHEr, 5.94 0.47 26.66 0.00 -5.67 29.11 -20.00 0.00 59.10 FUM, GLCpts, RPE 300 ADHEr, 5.94 0.47 26.66 0.00 -5.67 29.11 -20.00 0.00 59.10 GLCpts, MDH, RPE 301 ADHEr, 5.92 0.43 27.41 0.00 -5.98 29.66 -20.00 0.00 60.12 FUM, LDH_D, SUCD4 302 ADHEr, 5.92 0.43 27.41 0.00 -5.98 29.66 -20.00 0.00 60.12 LDH_D, MDH, SUCD4 303 ADHEr, 5.92 0.46 26.93 0.00 -5.81 29.32 -20.00 0.00 59.49 GLCpts, GLUDy, TAL 304 ADHEr, 5.90 0.47 26.74 0.00 -5.78 29.20 -20.00 0.00 59.28 FUM, GLCpts, TAL 305 ADHEr, 5.90 0.47 26.74 0.00 -5.78 29.20 -20.00 0.00 59.28 GLCpts, MDH, TAL 306 ADHEr, 5.86 0.46 26.97 0.00 -5.93 29.38 -20.00 0.00 59.63 CBMK2, GLU5K, SUCD4 307 ADHEr, 5.86 0.46 26.97 0.00 -5.93 29.38 -20.00 0.00 59.63 CBMK2, G5SD, SUCD4 308 ADHEr, 5.79 0.56 25.29 0.00 -5.47 28.25 -20.00 0.00 57.55 CBMK2, GLU5K, RPE 309 ADHEr, 5.79 0.56 25.29 0.00 -5.47 28.25 -20.00 0.00 57.55 CBMK2, G5SD, RPE 310 ADHEr, 5.79 0.50 26.42 0.00 -5.86 29.04 -20.00 0.00 59.02 CBMK2, GLCpts, GLU5K 311 ADHEr, 5.79 0.56 25.29 0.00 -5.47 28.25 -20.00 0.00 57.55 ASNS2, CBMK2, RPE 312 ADHEr, 5.79 0.50 26.42 0.00 -5.86 29.04 -20.00 0.00 59.02 CBMK2, G5SD, GLCpts 313 ADHEr, 5.79 0.50 26.42 0.00 -5.86 29.04 -20.00 0.00 59.02 ASNS2, CBMK2, GLCpts 314 ADHEr, 5.74 0.57 25.40 0.00 -5.60 28.36 -20.00 0.00 57.77 CBMK2, GLU5K, TAL 315 ADHEr, 5.74 0.57 25.40 0.00 -5.60 28.36 -20.00 0.00 57.77 CBMK2, G5SD, TAL 316 ADHEr, 5.74 0.57 25.40 0.00 -5.60 28.36 -20.00 0.00 57.77 ASNS2, CBMK2, TAL 317 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.77 -20.00 0.00 58.54 CBMK2, GLU5K, MDH 318 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.77 -20.00 0.00 58.54 CBMK2, FUM, G5SD 319 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.77 -20.00 0.00 58.54 CBMK2, G5SD, MDH 320 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.77 -20.00 0.00 58.54 CBMK2, FUM, GLU5K 321 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.77 -20.00 0.00 58.54 ASNS2, CBMK2, MDH 322 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.77 -20.00 0.00 58.54 ASNS2, CBMK2, FUM 323 ADHEr, 5.68 0.57 25.48 0.00 -5.75 28.46 -20.00 0.00 57.99 ASNS2, GLU5K, SO4t2 324 ADHEr, 14.96 0.23 14.07 0.00 22.42 0.00 -20.00 0.00 15.71 ASPT, LDH_D, MDH, PFLi 325 ADHEr, 14.81 0.14 16.32 0.00 22.21 0.00 -20.00 0.00 17.33 EDA, GLUDy, PFLi, PGI 326 ADHEr, 14.63 0.18 15.95 0.00 21.94 0.00 -20.00 0.00 17.25 ATPS4r, G6PDHy, GLCpts, MDH 327 ADHEr, 14.63 0.18 15.95 0.00 21.94 0.00 -20.00 0.00 17.25 ATPS4r, GLCpts, MDH, PGL 328 ADHEr, 14.42 0.11 17.14 0.00 21.96 1.32 -20.00 0.00 19.28 EDA, GLUDy, NADH6, PGI 329 ADHEr, 13.73 0.10 9.55 0.00 1.40 40.38 -20.00 0.00 50.65 G6PDHy, LDH_D, PPC, THD2 330 ADHEr, 13.73 0.10 9.55 0.00 1.40 40.38 -20.00 0.00 50.65 LDH_D, PGL, PPC, THD2 331 ADHEr, 13.37 0.17 4.27 6.42 13.23 18.61 -20.00 0.00 30.48 ATPS4r, FRD2, LDH_D, NADH6 332 ADHEr, 13.09 0.16 10.61 0.00 1.33 38.58 -20.00 0.00 50.31 EDA, LDH_D, PPC, THD2 333 ADHEr, 12.96 0.12 12.19 0.00 0.07 38.74 -20.00 0.00 51.78 FRD2, GLUDy, LDH_D, RPE 334 ADHEr, 12.95 0.12 12.13 0.00 0.07 38.68 -20.00 0.00 51.70 FRD2, LDH_D, RPE, THD2 335 ADHEr, 12.94 0.11 12.36 0.00 -0.02 38.83 -20.00 0.00 52.00 FRD2, GLUDy, LDH_D, THD2 336 ADHEr, 12.94 0.12 12.24 0.00 0.03 38.74 -20.00 0.00 51.84 FRD2, GLUDy, LDH_D, TAL 337 ADHEr, 12.92 0.13 12.19 0.00 0.03 38.69 -20.00 0.00 51.77 FRD2, LDH_D, TAL, THD2 338 ADHEr, 12.83 0.28 9.13 0.00 9.66 29.15 -20.00 0.00 40.26 ATPS4r, LDH_D, NADH6, PPC 339 ADHEr, 12.77 0.13 12.57 0.00 -0.13 38.55 -20.00 0.00 52.04 ASPT, FUM, GLUDy, LDH_D 340 ADHEr, 12.75 0.13 12.54 0.00 -0.14 38.50 -20.00 0.00 51.99 ASPT, FUM, LDH_D, THD2 341 ADHEr, 12.69 0.38 17.31 0.00 19.01 0.00 -20.00 0.00 19.99 ME2, PFLi, PGL, THD2 342 ADHEr, 12.69 0.38 17.31 0.00 19.01 0.00 -20.00 0.00 19.99 G6PDHy, ME2, PFLi, THD2 343 ADHEr, 12.67 0.10 12.72 0.00 0.38 39.23 -20.00 0.00 52.69 ACKr, ACS, PPC, RPE 344 ADHEr, 12.67 0.16 11.69 0.00 0.58 38.82 -20.00 0.00 51.63 GLUDy, LDH_D, PPC, RPE 345 ADHEr, 12.67 0.16 11.57 0.00 0.60 38.78 -20.00 0.00 51.52 LDH_D, PPC, RPE, THD2 346 ADHEr, 12.64 0.16 11.75 0.00 0.53 38.83 -20.00 0.00 51.71 GLUDy, LDH_D, PPC, TAL 347 ADHEr, 12.63 0.16 11.65 0.00 0.55 38.79 -20.00 0.00 51.61 LDH_D, PPC, TAL, THD2 348 ADHEr, 12.62 0.10 12.80 0.00 0.31 39.24 -20.00 0.00 52.79 GLCpts, GLUDy, LDH_D, PPC 349 ADHEr, 12.62 0.11 12.74 0.00 0.32 39.22 -20.00 0.00 52.73 GLCpts, LDH_D, PPC, THD2 350 ADHEr, 12.38 0.13 12.45 0.00 9.92 10.79 -20.00 0.00 30.65 ACKr, GLUDy, LDH_D, PGI 351 ADHEr, 12.20 0.20 10.35 0.00 18.28 0.00 -20.00 0.00 19.76 ASPT, ATPS4r, GLCpts, MDH 352 ADHEr, 12.11 0.25 21.60 0.00 18.15 0.00 -20.00 0.00 23.35 LDH_D, PFLi, SUCD4, THD2 353 ADHEr, 12.05 0.24 12.18 0.00 12.02 25.48 -20.00 0.00 39.36 ACKr, AKGD, ATPS4r, PYK 354 ADHEr, 12.00 0.24 12.13 0.00 12.20 25.51 -20.00 0.00 39.36 ACKr, ATPS4r, PYK, SUCOAS
355 ADHEr, 11.98 0.13 19.29 0.00 7.97 19.98 -20.00 0.00 40.22 ATPS4r, EDA, GLUDy, PGI 356 ADHEr, 11.98 0.14 19.25 0.00 7.98 19.96 -20.00 0.00 40.18 EDA, GLCpts, GLUDy, PGI 357 ADHEr, 11.91 0.20 12.01 3.31 5.46 24.78 -20.00 0.00 41.50 ACKr, LDH_D, MDH, SUCD4 358 ADHEr, 11.90 0.49 12.16 0.00 27.81 0.00 -20.00 0.00 15.63 ACKr, GLUDy, NADH6, PYK 359 ADHEr, 11.76 0.39 19.51 0.00 17.62 0.00 -20.00 0.00 22.30 FUM, LDH_D, PFLi, THD2 360 ADHEr, 11.76 0.39 19.51 0.00 17.62 0.00 -20.00 0.00 22.30 LDH_D, MDH, PFLi, THD2 361 ADHEr, 11.67 0.14 0.00 0.00 1.15 24.89 -19.46 0.00 38.65 ACt6, ATPS4r, LDH_D, PPC 362 ADHEr, 11.66 0.21 11.78 0.00 8.63 21.96 -20.00 0.00 37.33 ACKr, LDH_D, PYRt2, SUCD4 363 ADHEr, 11.59 0.23 20.54 0.00 13.84 9.06 -20.00 0.00 31.25 ATPS4r, FUM, LDH_D, SUCD4 364 ADHEr, 11.59 0.23 20.54 0.00 13.84 9.06 -20.00 0.00 31.25 ATPS4r, LDH_D, MDH, SUCD4 365 ADHEr, 11.52 0.55 11.82 0.00 27.24 0.00 -20.00 0.00 15.76 ACKr, CBMK2, NADH6, PYK 366 ADHEr, 11.50 0.56 11.81 0.00 27.22 0.00 -20.00 0.00 15.77 ACKr, NADH6, PYK, RPE 367 ADHEr, 11.50 0.56 11.80 0.00 27.21 0.00 -20.00 0.00 15.77 ACKr, ASNS2, NADH6, PYK 368 ADHEr, 11.49 0.56 11.80 0.00 27.20 0.00 -20.00 0.00 15.77 ACKr, NADH6, PYK, TAL 369 ADHEr, 11.46 0.30 8.96 0.00 8.56 25.90 -20.00 0.00 38.85 ASPT, ATPS4r, FUM, LDH_D 370 ADHEr, 11.43 0.34 17.13 0.00 8.27 17.71 -20.00 0.00 37.24 ASPT, ATPS4r, LDH_D, MDH 371 ADHEr, 11.38 0.38 20.90 0.00 17.05 0.00 -20.00 0.00 23.58 MDH, PFLi, PYK, THD2 372 ADHEr, 11.38 0.38 20.90 0.00 17.05 0.00 -20.00 0.00 23.58 FUM, PFLi, PYK, THD2 373 ADHEr, 11.25 0.53 11.54 0.00 23.26 0.00 -20.00 0.00 17.50 ACKr, FUM, LDH_D, NADH6 374 ADHEr, 11.24 0.27 20.72 0.00 13.49 8.70 -20.00 0.00 31.38 ATPS4r, G6PDHy, LDH_D, SUCD4 375 ADHEr, 11.24 0.27 20.72 0.00 13.49 8.70 -20.00 0.00 31.38 ATPS4r, LDH_D, PGL, SUCD4 376 ADHEr, 11.24 0.27 20.72 0.00 13.49 8.70 -20.00 0.00 31.38 ATPS4r, LDH_D, PGDH, SUCD4 377 ADHEr, 11.24 0.14 2.95 0.00 3.54 28.60 -20.00 0.02 42.95 GLYCL, PGL, PPC, THD2 378 ADHEr, 11.24 0.14 2.95 0.00 3.54 28.60 -20.00 0.02 42.95 G6PDHy, GLYCL, PPC, THD2 379 ADHEr, 11.22 0.14 2.94 0.00 3.54 28.55 -20.00 0.00 42.92 FTHFD, G6PDHy, PPC, THD2 380 ADHEr, 11.22 0.14 2.94 0.00 3.54 28.55 -20.00 0.00 42.92 FTHFD, PGL, PPC, THD2 381 ADHEr, 11.22 0.14 2.94 0.00 3.54 28.55 -20.00 0.00 42.92 MTHFC, PGL, PPC, THD2 382 ADHEr, 11.22 0.14 2.94 0.00 3.54 28.55 -20.00 0.00 42.92 G6PDHy, MTHFC, PPC, THD2 383 ADHEr, 11.20 0.32 22.07 0.00 17.78 0.00 -20.00 0.00 24.34 ATPS4r, LDH_D, PFLi, SUCD4 384 ADHEr, 11.18 0.28 20.84 0.00 13.41 8.69 -20.00 0.00 31.49 ATPS4r, LDH_D, SUCD4, TAL 385 ADHEr, 11.13 0.28 23.11 0.00 17.68 0.00 -20.00 0.00 25.09 ATPS4r, LDH_D, RPE, SUCD4 386 ADHEr, 11.13 0.27 19.49 0.00 16.68 0.00 -20.00 0.00 24.27 ATPS4r, FUM, GLCpts, NADH6 387 ADHEr, 11.13 0.27 19.49 0.00 16.68 0.00 -20.00 0.00 24.27 ATPS4r, GLCpts, MDH, NADH6 388 ADHEr, 11.12 0.34 22.41 0.00 16.65 0.00 -20.00 0.00 24.82 ATPS4r, LDH_D, NADH6, PFLi 389 ADHEr, 11.06 0.63 11.41 0.00 26.56 0.00 -20.00 0.00 15.92 ACKr, MALS, NADH12, NADH6 390 ADHEr, 11.06 0.63 11.41 0.00 26.56 0.00 -20.00 0.00 15.92 ACKr, ICL, NADH12, NADH6 391 ADHEr, 11.06 0.64 11.40 0.00 26.54 0.00 -20.00 0.00 15.93 ACKr, CBMK2, LDH_D, NADH6 392 ADHEr, 11.04 0.35 22.48 0.00 16.54 0.00 -20.00 0.00 24.94 ATPS4r, GLCpts, MDH, PFLi 393 ADHEr, 11.04 0.35 22.48 0.00 16.54 0.00 -20.00 0.00 24.94 ATPS4r, FUM, GLCpts, PFLi 394 ADHEr, 11.03 0.64 11.38 0.00 26.51 0.00 -20.00 0.00 15.93 ACKr, ASNS2, LDH_D, NADH6 395 ADHEr, 11.00 0.38 11.21 0.00 5.04 32.88 -20.00 0.00 46.77 ACKr, ATPS4r, LDH_D, THD2 396 ADHEr, 11.00 0.15 26.58 0.00 16.49 0.00 -20.00 0.00 27.66 GLCpts, GLUDy, PFLi, PGI 397 ADHEr, 10.97 0.15 26.63 0.00 16.45 0.00 -20.00 0.00 27.73 GLCpts, GLUDy, PFK, PFLi 398 ADHEr, 10.97 0.15 26.63 0.00 16.45 0.00 -20.00 0.00 27.73 FBA, GLCpts, GLUDy, PFLi 399 ADHEr, 10.97 0.15 26.63 0.00 16.45 0.00 -20.00 0.00 27.73 GLCpts, GLUDy, PFLi, TPI 400 ADHEr, 10.97 0.11 27.23 0.00 15.95 0.99 -20.00 0.00 29.02 ATPS4r, GLCpts, NADH6, PGI 401 ADHEr, 10.95 0.12 27.26 0.00 15.95 0.94 -20.00 0.00 29.02 ATPS4r, GLCpts, NADH6, TPI 402 ADHEr, 10.95 0.12 27.26 0.00 15.95 0.94 -20.00 0.00 29.02 ATPS4r, GLCpts, NADH6, PFK 403 ADHEr, 10.95 0.12 27.26 0.00 15.95 0.94 -20.00 0.00 29.02 ATPS4r, FBA, GLCpts, NADH6 404 ADHEr, 10.94 0.17 26.35 0.00 16.40 0.00 -20.00 0.00 27.57 GLCpts, PFLi, RPE, TPI 405 ADHEr, 10.94 0.17 26.35 0.00 16.40 0.00 -20.00 0.00 27.57 GLCpts, PFK, PFLi, RPE 406 ADHEr, 10.94 0.17 26.35 0.00 16.40 0.00 -20.00 0.00 27.57 FBA, GLCpts, PFLi, RPE 407 ADHEr, 10.93 0.28 24.13 0.00 16.38 0.00 -20.00 0.00 26.12 ATPS4r,
NADH6, PFLi, PYK 408 ADHEr, 10.93 0.17 26.37 0.00 16.38 0.00 -20.00 0.00 27.59 GLCpts, PFK, PFLi, TAL 409 ADHEr, 10.93 0.17 26.37 0.00 16.38 0.00 -20.00 0.00 27.59 FBA, GLCpts, PFLi, TAL 410 ADHEr, 10.93 0.17 26.37 0.00 16.38 0.00 -20.00 0.00 27.59 GLCpts, PFLi, TAL, TPI 411 ADHEr, 10.92 0.18 26.25 0.00 16.37 0.00 -20.00 0.00 27.52 FBA, GLUDy, PFLi, RPE 412 ADHEr, 10.92 0.18 26.25 0.00 16.37 0.00 -20.00 0.00 27.52 GLUDy, PFK, PFLi, RPE 413 ADHEr, 10.92 0.18 26.25 0.00 16.37 0.00 -20.00 0.00 27.52 GLUDy, PFLi, RPE, TPI 414 ADHEr, 10.91 0.18 26.27 0.00 16.35 0.00 -20.00 0.00 27.55 GLUDy, PFLi, TAL, TPI 415 ADHEr, 10.91 0.18 26.27 0.00 16.35 0.00 -20.00 0.00 27.55 GLUDy, PFK, PFLi, TAL 416 ADHEr, 10.91 0.18 26.27 0.00 16.35 0.00 -20.00 0.00 27.55 FBA, GLUDy, PFLi, TAL 417 ADHEr, 10.79 0.32 21.77 0.00 14.53 5.29 -20.00 0.00 29.37 ATPS4r, LDH_D, NADH6, SUCD4 418 ADHEr, 10.76 0.23 25.74 0.00 16.13 0.00 -20.00 0.00 27.34 FUM, LDH_D, PFLi, SUCD4 419 ADHEr, 10.76 0.23 25.74 0.00 16.13 0.00 -20.00 0.00 27.34 LDH_D, MDH, PFLi, SUCD4 H2O LAC NH4 NO3 PI PYR SO4 SUC VAL 1 -0.90 0.00 -4.96 0.00 -0.61 0.00 -0.10 0.00 0.00 2 14.07 0.00 -4.31 0.00 -0.53 0.00 -0.09 0.00 0.00 3 2.51 0.00 -4.48 0.00 -0.55 0.00 -0.09 0.00 0.00 4 0.13 0.00 -4.83 0.00 -0.60 0.00 -0.10 0.00 0.00 5 -6.65 0.00 -2.62 0.00 -0.32 0.00 -0.05 0.00 0.00 6 -6.68 0.00 -2.66 0.00 -0.33 0.00 -0.05 0.00 0.00 7 -6.68 0.00 -2.66 0.00 -0.33 0.00 -0.05 0.00 0.00 8 1.88 0.00 -5.25 -2.00 -0.65 0.00 -0.11 0.00 0.00 9 -6.68 0.00 -2.66 0.00 -0.33 0.00 -0.05 0.00 0.00 10 -3.22 0.00 -4.04 0.00 -0.50 0.00 -0.08 0.00 0.00 11 -0.76 0.00 -4.95 0.00 -0.61 0.00 -0.10 0.00 0.00 12 -2.34 0.00 -4.39 0.00 -0.54 0.00 -0.09 0.00 0.00 13 -2.13 0.00 -4.47 0.00 -0.55 0.00 -0.09 0.00 0.00 14 -0.83 0.00 -4.95 0.00 -0.61 0.00 -0.10 0.00 0.00 15 -1.66 0.00 -4.66 0.00 -0.58 0.00 -0.09 0.00 0.00 16 -1.66 0.00 -4.66 0.00 -0.58 0.00 -0.09 0.00 0.00 17 -1.01 0.00 -4.92 0.00 -0.61 0.00 -0.10 0.00 0.00 18 0.36 0.00 -1.93 0.00 -0.24 0.00 -0.04 0.00 0.00 19 0.38 0.00 -1.94 0.00 -0.24 0.00 -0.04 0.00 0.00 20 9.09 0.00 -1.73 0.00 -0.21 0.00 -0.03 0.00 0.00 21 9.11 0.00 -1.75 0.00 -0.22 0.00 -0.04 0.00 0.00 22 9.11 0.00 -1.75 0.00 -0.22 0.00 -0.04 0.00 0.00 23 9.11 0.00 -1.75 0.00 -0.22 0.00 -0.04 0.00 0.00 24 14.16 0.00 -4.23 0.00 -0.52 0.00 -0.09 0.00 0.00 25 12.93 0.00 -3.72 0.00 -0.46 0.00 -0.08 0.00 0.00 26 13.23 0.00 -3.87 0.00 -0.48 0.00 -0.08 0.00 0.00 27 14.08 0.00 -4.30 0.00 -0.53 0.00 -0.09 0.00 0.00 28 14.07 0.00 -4.30 0.00 -0.53 0.00 -0.09 0.00 0.00 29 13.99 0.00 -4.27 0.00 -0.53 0.00 -0.09 0.00 0.00 30 -3.19 0.00 -2.28 0.00 -0.28 0.00 -0.05 0.00 0.00 31 6.46 0.00 -4.15 0.00 -0.51 0.00 -0.08 0.00 0.00 32 0.48 0.00 -2.25 0.00 -0.28 0.00 -0.05 0.00 0.00 33 0.48 0.00 -2.25 0.00 -0.28 0.00 -0.05 0.00 0.00 34 0.48 0.00 -2.25 0.00 -0.28 0.00 -0.05 0.00 0.00 35 3.90 0.00 -4.30 0.00 -0.53 0.00 -0.09 0.00 0.00 36 1.11 0.00 -4.22 0.00 -0.52 0.00 -0.09 0.00 0.00 37 1.11 0.00 -4.22 0.00 -0.52 0.00 -0.09 0.00 0.00 38 -3.12 0.00 -2.02 0.00 -0.25 0.00 -0.04 0.00 0.00 39 0.18 0.00 -4.28 0.00 -0.53 0.00 -0.09 0.00 0.00 40 0.18 0.00 -4.28 0.00 -0.53 0.00 -0.09 0.00 0.00 41 -3.17 0.00 -2.06 0.00 -0.25 0.00 -0.04 0.00 0.00 42 -3.17 0.00 -2.06 0.00 -0.25 0.00 -0.04 0.00 0.00 43 -3.17 0.00 -2.06 0.00 -0.25 0.00 -0.04 0.00 0.00 44 1.14 0.00 -3.90 0.00 -0.48 0.00 -0.08 0.00 0.00 45 2.70 0.00 -4.46 0.00 -0.55 0.00 -0.09 0.00 0.00 46 1.46 0.00 -4.03 0.00 -0.50 0.00 -0.08 0.00 0.00 47 1.61 0.00 -4.10 0.00 -0.51 0.00 -0.08 0.00 0.00 48 1.61 0.00 -4.10 0.00 -0.51 0.00 -0.08 0.00 0.00 49 2.61 0.00 -4.47 0.00 -0.55 0.00 -0.09 0.00 0.00 50 -1.43 0.00 -4.27 0.00 -0.53 0.00 -0.09 0.00 0.00 51 -1.17 0.00 -4.36 0.00 -0.54 0.00 -0.09 0.00 0.00 52 -8.37 0.00 -1.95 0.00 -0.24 0.00 -0.04 0.00 0.00 53 -7.58 0.00 -2.25 0.00 -0.28 0.00 -0.05 0.00 0.00 54 -8.47 0.00 -1.94 0.00 -0.24 0.00 -0.04 0.00 0.00 55 -8.47 0.00 -1.94 0.00 -0.24 0.00 -0.04 0.00 0.00 56 -8.47 0.00 -1.94 0.00 -0.24 0.00 -0.04 0.00 0.00 57 -8.37 0.00 -1.98 0.00 -0.25 0.00 -0.04 0.00 0.00 58 -8.37 0.00 -1.98 0.00 -0.25 0.00 -0.04 0.00 0.00 59 -8.37 0.00 -1.98 0.00 -0.25 0.00 -0.04 0.00 0.00 60 -7.35 0.00 -2.35 0.00 -0.29 0.00 -0.05 0.00 0.00 61 -7.61 0.00 -2.29 0.00 -0.28 0.00 -0.05 0.00 0.00 62 -7.61 0.00 -2.29 0.00 -0.28 0.00 -0.05 0.00 0.00 63 -7.61 0.00 -2.29 0.00 -0.28 0.00 -0.05 0.00 0.00 64 -6.66 0.00 -2.63 0.00 -0.32 0.00 -0.05 0.00 0.00 65 -6.66 0.00 -2.63 0.00 -0.32 0.00 -0.05 0.00 0.00 66 -6.66 0.00 -2.63 0.00 -0.32 0.00 -0.05 0.00 0.00 67 -7.38 0.00 -2.38 0.00 -0.29 0.00 -0.05 0.00 0.00 68 -7.38 0.00 -2.38 0.00 -0.29 0.00 -0.05 0.00 0.00 69 -7.38 0.00 -2.38 0.00 -0.29 0.00 -0.05 0.00 0.00 70 -6.67 0.00 -2.64 0.00 -0.33 0.00 -0.05 0.00 0.00 71 -6.67 0.00 -2.64 0.00 -0.33 0.00 -0.05 0.00 0.00 72 -6.67 0.00 -2.64 0.00 -0.33 0.00 -0.05 0.00 0.00 73 0.48 0.00 -4.75 -2.00 -0.59 0.00 -0.10 0.00 0.00 74 -5.01 0.00 -3.32 0.00 -0.41 0.00 -0.07 0.00 0.00 75 -4.70 0.00 -3.45 0.00 -0.43 0.00 -0.07 0.00 0.00 76 -3.14 0.00 -4.01 0.00 -0.50 0.00 -0.08 0.00 0.00 77 -4.29 0.00 -3.61 0.00 -0.45 0.00 -0.07 0.00 0.00 78 -4.29 0.00 -3.61 0.00 -0.45 0.00 -0.07 0.00 0.00 79 -4.24 0.00 -3.63 0.00 -0.45 0.00 -0.07 0.00 0.00 80 -3.18 0.00 -4.02 0.00 -0.50 0.00 -0.08 0.00 0.00 81 -2.22 0.00 -4.37 0.00 -0.54 0.00 -0.09 0.00 0.00 82 -3.42 0.00 -3.96 0.00 -0.49 0.00 -0.08 0.00 0.00 83 -3.12 0.00 -4.08 0.00 -0.50 0.00 -0.08 0.00 0.00 84 -3.12 0.00 -4.08 0.00 -0.50 0.00 -0.08 0.00 0.00 85 -2.00 0.00 -4.46 0.00 -0.55 0.00 -0.09 0.00 0.00 86 -2.28 0.00 -4.38 0.00 -0.54 0.00 -0.09 0.00 0.00 87 -1.54 0.00 -4.64 0.00 -0.57 0.00 -0.09 0.00 0.00 88 -1.54 0.00 -4.64 0.00 -0.57 0.00 -0.09 0.00 0.00 89 -3.17 0.00 -4.03 0.00 -0.50 0.00 -0.08 0.09 0.00 90 -3.17 0.00 -4.03 0.00 -0.50 0.00 -0.08 0.09 0.00 91 -1.60 0.00 -4.65 0.00 -0.58 0.00 -0.09 0.00 0.00 92 -1.60 0.00 -4.65 0.00 -0.58 0.00 -0.09 0.00 0.00 93 -2.06 0.00 -4.47 0.00 -0.55 0.00 -0.09 0.00 0.00 94 -1.05 0.00 -4.90 0.00 -0.61 0.00 -0.10 0.00 0.00 95 -1.05 0.00 -4.90 0.00 -0.61 0.00 -0.10 0.00 0.00 96 -1.05 0.00 -4.90 0.00 -0.61 0.00 -0.10 0.00 0.00 97 -1.03 0.00 -4.91 0.00 -0.61 0.00 -0.10 0.00 0.00 98 -1.01 0.00 -4.92 0.00 -0.61 0.00 -0.10 0.00 0.00 99 10.28 0.00 -1.38 0.00 -0.17 0.00 -0.03 0.00 0.00 100 9.91 0.00 -1.11 -2.00 -0.14 0.00 -0.02 0.00 0.00 101 -10.70 0.00 -1.05 0.00 -0.13 0.00 -0.02 0.00 0.00 102 -10.60 0.00 -1.09 0.00 -0.13 0.00 -0.02 0.00 0.00 103 -10.25 0.00 -0.96 -2.00 -0.12 0.00 -0.02 0.00 0.00 104 -9.20 0.00 -1.38 -2.00 -0.17 0.00 -0.03 0.00 0.00 105 -9.09 0.00 -1.43 -2.00 -0.18 0.00 -0.03 0.00 0.00 106 -1.21 0.00 -1.30 0.00 -0.16 0.00 -0.03 0.00 0.00 107 -1.16 0.00 -1.32 0.00 -0.16 0.00 -0.03 0.00 0.00 108 -0.12 0.00 -1.73 0.00 -0.21 0.00 -0.04 0.00 0.00 109 -0.13 0.00 -1.74 0.00 -0.21 0.00 -0.04 0.00 0.00 110 5.32 0.00 -4.64 -5.00 -0.30 0.00 -0.05 0.00 2.22 111 25.99 0.00 -4.86 -20.00 -0.60 0.00 -0.10 0.00 0.00 112 27.24 0.00 -5.56 -20.00 -0.69 0.00 -0.11 0.00 0.00 113 8.60 0.00 -1.48 0.00 -0.18 0.00 -0.03 0.00 0.00 114 8.73 0.00 -1.55 0.00 -0.19 0.00 -0.03 0.00 0.00 115 8.62 0.00 -1.50 0.00 -0.19 0.00 -0.03 0.00 0.00 116 8.62 0.00 -1.50 0.00 -0.19 0.00 -0.03 0.00 0.00 117 8.62 0.00 -1.50 0.00 -0.19 0.00 -0.03 0.00 0.00 118 8.75 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 119 8.75 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 120 8.75 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 121 9.09 0.00 -1.74 0.00 -0.21 0.00 -0.04 0.00 0.00 122 9.09 0.00 -1.74 0.00 -0.21 0.00 -0.04 0.00 0.00 123 9.09 0.00 -1.74 0.00 -0.21 0.00 -0.04 0.00 0.00 124 9.10 0.00 -1.74 0.00 -0.22 0.00 -0.04 0.00 0.00 125 9.10 0.00 -1.74 0.00 -0.22 0.00 -0.04 0.00 0.00 126 9.10 0.00 -1.74 0.00 -0.22 0.00 -0.04 0.00 0.00 127 1.46 0.00 -1.48 0.00 -0.18 0.00 -0.03 0.00 0.00 128 13.01 0.00 -3.65 0.00 -0.45 0.00 -0.07 0.00 0.00 129 13.33 0.00 -3.81 0.00 -0.47 0.00 -0.08 0.00 0.00 130 10.77 0.00 -2.61 0.00 -0.32 0.00 -0.05 0.00 0.00 131 12.01 0.00 -3.25 0.00 -0.40 0.00 -0.07 0.00 0.00 132 8.53 0.00 -3.04 -2.00 -0.38 0.00 -0.06 0.00 0.00 133 12.15 0.00 -3.32 0.00 -0.41 0.00 -0.07 0.00 0.00 134 12.15 0.00 -3.32 0.00 -0.41 0.00 -0.07 0.00 0.00 135 12.20 0.00 -3.35 0.00 -0.41 0.00 -0.07 0.00 0.00 136 12.94 0.00 -3.71 0.00 -0.46 0.00 -0.08 0.00 0.00 137 12.93 0.00 -3.72 0.00 -0.46 0.00 -0.08 0.00 0.00 138 13.24 0.00 -3.87 0.00 -0.48 0.00 -0.08 0.00 0.00 139 12.87 0.00 -3.69 0.00 -0.46 0.00 -0.07 0.00 0.00 140 12.95 0.00 -3.73 0.00 -0.46 0.00 -0.08 0.00 0.00 141 12.95 0.00 -3.73 0.00 -0.46 0.00 -0.08 0.00 0.00 142 13.23 0.00 -3.87 0.00 -0.48 0.00 -0.08 0.00 0.00 143 13.17 0.00 -3.84 0.00 -0.48 0.00 -0.08 0.00 0.00 144 14.01 0.00 -4.26 0.00 -0.53 0.00 -0.09 0.00 0.00 145 14.00 0.00 -4.26 0.00 -0.53 0.00 -0.09 0.00 0.00 146 14.01 0.00 -4.28 0.00 -0.53 0.00 -0.09 0.00 0.00 147 14.01 0.00 -4.28 0.00 -0.53 0.00 -0.09 0.00 0.00 148 5.89 0.00 -3.22 0.00 -0.40 0.00 -0.07 0.00 0.00 149 5.89 0.00 -3.22 0.00 -0.40 0.00 -0.07 0.00 0.00 150 4.45 0.00 -1.50 0.00 -0.19 0.00 -0.03 0.00 0.00 151 4.45 0.00 -1.50 0.00 -0.19 0.00 -0.03 0.00 0.00 152 4.45 0.00 -1.50 0.00 -0.19 0.00 -0.03 0.00 0.00 153 11.87 0.00 -3.80 0.00 -0.47 0.00 -0.08 0.00 0.00 154 11.87 0.00 -3.80 0.00 -0.47 0.00 -0.08 0.00 0.00 155 11.87 0.00 -3.80 0.00 -0.47 0.00 -0.08 0.00 0.00 156 11.87 0.00 -3.80 0.00 -0.47 0.00 -0.08 0.00 0.00 157 11.87 0.00 -3.80 0.00 -0.47 0.00 -0.08 0.00 0.00 158 11.87 0.00 -3.80 0.00 -0.47 0.00 -0.08 0.00 0.00 159 -0.41 0.00 -3.83 -10.00 -0.47 0.00 -0.08 0.00 0.02 160 11.60 0.00 -3.81 0.00 -0.47 0.00 -0.08 0.00 0.00 161 11.60 0.00 -3.81 0.00 -0.47 0.00 -0.08 0.00 0.00 162 11.59 0.00 -3.65 -2.00 -0.45 0.00 -0.07 0.00 0.00 163 11.34 0.00 -3.82 0.00 -0.47 0.00 -0.08 0.00 0.00 164 11.34 0.00 -3.82 0.00 -0.47 0.00 -0.08 0.00 0.00 165 10.80 0.00 -4.00 0.00 -0.50 0.00 -0.08 0.00 0.00 166 10.80 0.00 -4.00 0.00 -0.50 0.00 -0.08 0.00 0.00 167 10.80 0.00 -4.00 0.00 -0.50 0.00 -0.08 0.00 0.00 168 10.51 0.00 -4.01 0.00 -0.50 0.00 -0.08 0.00 0.00 169 10.25 0.00 -4.02 0.00 -0.50 0.00 -0.08 0.00 0.00 170 -0.86 0.00 -3.72 0.00 -0.46 0.00 -0.08 0.00 0.00 171 -0.86 0.00 -3.72 0.00 -0.46 0.00 -0.08 0.00 0.00 172 6.22 8.06 -4.07 -20.00 -0.50 0.00 -0.08 0.00 0.00 173 6.22 8.06 -4.07 -20.00 -0.50 0.00 -0.08 0.00 0.00 174 5.40 0.00 -3.75 0.00 -0.46 0.00 -0.08 0.00 0.00 175 15.97 0.00 -6.24 -15.00 -0.69 2.92 -0.11 0.00 0.68 176 0.40 0.00 -2.23 0.00 -0.28 0.00 -0.05 0.00 0.00 177 0.40 0.00 -2.23 0.00 -0.28 0.00 -0.05 0.00 0.00 178 0.40 0.00 -2.23 0.00 -0.28 0.00 -0.05 0.00 0.00 179 -0.22 0.00 -2.02 0.00 -0.25 0.00 -0.04 0.00 0.00 180 -0.22 0.00 -2.02 0.00 -0.25 0.00 -0.04 0.00 0.00 181 -0.22 0.00 -2.02 0.00 -0.25 0.00 -0.04 0.00 0.00 182 0.44 0.00 -2.24 0.00 -0.28 0.00 -0.05 0.00 0.00 183 0.44 0.00 -2.24 0.00 -0.28 0.00 -0.05 0.00 0.00 184 0.44 0.00 -2.24 0.00 -0.28 0.00 -0.05 0.00 0.00 185 -7.34 0.00 -2.39 0.00 -0.30 0.00 -0.05 0.00 0.00 186 12.12 0.00 -6.14 -15.00 -0.76 0.00 -0.12 0.00 0.00 187 12.12 0.00 -6.14 -15.00 -0.76 0.00 -0.12 0.00 0.00 188 2.35 0.00 -3.74 0.00 -0.46 0.00 -0.08 0.00 0.00 189 2.75 0.00 -3.88 0.00 -0.48 0.00 -0.08 0.00 0.00 190 -3.60 0.00 -3.89 0.00 -0.48 0.00 -0.08 0.00 0.00 191 -3.94 0.00 -1.68 0.00 -0.21 0.00 -0.03 0.00 0.00 192 -1.90 0.00 -3.49 0.00 -0.43 0.00 -0.07 0.00 0.00 193 -3.63 0.00 -1.81 0.00 -0.22 0.00 -0.04 0.00 0.00 194 13.82 0.00 -2.77 0.00 -0.34 17.80 -0.06 0.00 0.00 195 -3.99 0.00 -1.71 0.00 -0.21 0.00 -0.03 0.00 0.00
196 -3.99 0.00 -1.71 0.00 -0.21 0.00 -0.03 0.00 0.00 197 13.82 0.00 -2.77 0.00 -0.34 17.80 -0.06 0.00 0.00 198 -3.99 0.00 -1.71 0.00 -0.21 0.00 -0.03 0.00 0.00 199 -1.38 0.00 -3.71 0.00 -0.46 0.00 -0.08 0.00 0.00 200 -1.38 0.00 -3.71 0.00 -0.46 0.00 -0.08 0.00 0.00 201 -3.13 0.00 -2.03 0.00 -0.25 0.00 -0.04 0.00 0.00 202 -3.13 0.00 -2.03 0.00 -0.25 0.00 -0.04 0.00 0.00 203 -3.13 0.00 -2.03 0.00 -0.25 0.00 -0.04 0.00 0.00 204 -3.69 0.00 -1.84 0.00 -0.23 0.00 -0.04 0.00 0.00 205 -3.69 0.00 -1.84 0.00 -0.23 0.00 -0.04 0.00 0.00 206 -3.69 0.00 -1.84 0.00 -0.23 0.00 -0.04 0.00 0.00 207 -3.15 0.00 -2.04 0.00 -0.25 0.00 -0.04 0.00 0.00 208 -3.15 0.00 -2.04 0.00 -0.25 0.00 -0.04 0.00 0.00 209 -3.15 0.00 -2.04 0.00 -0.25 0.00 -0.04 0.00 0.00 210 55.34 0.00 -4.99 -82.37 -0.62 0.00 -0.10 0.00 0.00 211 1.30 0.00 -3.88 0.00 -0.48 0.00 -0.08 0.00 0.00 212 0.21 0.00 -3.50 0.00 -0.43 0.00 -0.07 0.00 0.00 213 0.21 0.00 -3.50 0.00 -0.43 0.00 -0.07 0.00 0.00 214 0.22 0.00 -3.51 0.00 -0.43 0.00 -0.07 0.00 0.00 215 0.23 0.00 -3.51 0.00 -0.43 0.00 -0.07 0.00 0.00 216 0.23 0.00 -3.51 0.00 -0.43 0.00 -0.07 0.00 0.00 217 20.07 0.00 -5.03 -47.55 -0.62 0.00 -0.10 0.00 0.00 218 1.63 0.00 -4.01 0.00 -0.50 0.00 -0.08 0.00 0.00 219 1.76 0.00 -4.07 0.00 -0.50 0.00 -0.08 0.00 0.00 220 1.76 0.00 -4.07 0.00 -0.50 0.00 -0.08 0.00 0.00 221 1.23 0.00 -3.89 0.00 -0.48 0.00 -0.08 0.00 0.00 222 1.55 0.00 -4.02 0.00 -0.50 0.00 -0.08 0.00 0.00 223 1.69 0.00 -4.08 0.00 -0.51 0.00 -0.08 0.00 0.00 224 1.69 0.00 -4.08 0.00 -0.51 0.00 -0.08 0.00 0.00 225 2.38 0.00 -4.43 0.00 -0.55 0.00 -0.09 0.00 0.00 226 2.38 0.00 -4.43 0.00 -0.55 0.00 -0.09 0.00 0.00 227 2.40 0.00 -4.44 0.00 -0.55 0.00 -0.09 0.00 0.00 228 2.38 0.00 -4.43 0.00 -0.55 0.00 -0.09 0.00 0.00 229 6.92 7.87 -3.12 -2.00 -0.39 0.00 -0.06 0.00 0.00 230 -9.35 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 231 -9.07 0.00 -1.67 0.00 -0.21 0.00 -0.03 0.00 0.00 232 -9.07 0.00 -1.67 0.00 -0.21 0.00 -0.03 0.00 0.00 233 -8.89 0.00 -1.74 0.00 -0.22 0.00 -0.04 0.00 0.00 234 -9.35 0.00 -1.59 0.00 -0.20 0.00 -0.03 0.00 0.00 235 -9.35 0.00 -1.59 0.00 -0.20 0.00 -0.03 0.00 0.00 236 -9.35 0.00 -1.59 0.00 -0.20 0.00 -0.03 0.00 0.00 237 -8.46 0.00 -1.92 0.00 -0.24 0.00 -0.04 0.00 0.00 238 -8.46 0.00 -1.92 0.00 -0.24 0.00 -0.04 0.00 0.00 239 -8.46 0.00 -1.92 0.00 -0.24 0.00 -0.04 0.00 0.00 240 -9.08 0.00 -1.70 0.00 -0.21 0.00 -0.03 0.00 0.00 241 -9.08 0.00 -1.70 0.00 -0.21 0.00 -0.03 0.00 0.00 242 -9.08 0.00 -1.70 0.00 -0.21 0.00 -0.03 0.00 0.00 243 -9.08 0.00 -1.70 0.00 -0.21 0.00 -0.03 0.00 0.00 244 -9.08 0.00 -1.70 0.00 -0.21 0.00 -0.03 0.00 0.00 245 -9.08 0.00 -1.70 0.00 -0.21 0.00 -0.03 0.00 0.00 246 -8.19 0.00 -2.02 0.00 -0.25 0.00 -0.04 0.00 0.00 247 -8.37 0.00 -1.95 0.00 -0.24 0.00 -0.04 0.00 0.00 248 -8.37 0.00 -1.95 0.00 -0.24 0.00 -0.04 0.00 0.00 249 -8.37 0.00 -1.95 0.00 -0.24 0.00 -0.04 0.00 0.00 250 -8.96 0.00 -1.75 0.00 -0.22 0.00 -0.04 0.00 0.00 251 -8.96 0.00 -1.75 0.00 -0.22 0.00 -0.04 0.00 0.00 252 -8.96 0.00 -1.75 0.00 -0.22 0.00 -0.04 0.00 0.00 253 -8.90 0.00 -1.77 0.00 -0.22 0.00 -0.04 0.00 0.00 254 -8.90 0.00 -1.77 0.00 -0.22 0.00 -0.04 0.00 0.00 255 -8.90 0.00 -1.77 0.00 -0.22 0.00 -0.04 0.00 0.00 256 -8.47 0.00 -1.93 0.00 -0.24 0.00 -0.04 0.00 0.00 257 -8.47 0.00 -1.93 0.00 -0.24 0.00 -0.04 0.00 0.00 258 -8.47 0.00 -1.93 0.00 -0.24 0.00 -0.04 0.00 0.00 259 -8.37 0.00 -1.97 0.00 -0.24 0.00 -0.04 0.00 0.00 260 -8.37 0.00 -1.97 0.00 -0.24 0.00 -0.04 0.00 0.00 261 -8.37 0.00 -1.97 0.00 -0.24 0.00 -0.04 0.00 0.00 262 15.34 0.00 -1.92 0.00 -0.24 23.88 -0.04 0.00 0.00 263 -7.59 0.00 -2.26 0.00 -0.28 0.00 -0.05 0.00 0.00 264 -7.59 0.00 -2.26 0.00 -0.28 0.00 -0.05 0.00 0.00 265 -7.59 0.00 -2.26 0.00 -0.28 0.00 -0.05 0.00 0.00 266 21.84 0.00 -2.02 0.00 -0.25 30.11 -0.04 0.00 0.00 267 -8.22 0.00 -2.04 0.00 -0.25 0.00 -0.04 0.00 0.00 268 21.84 0.00 -2.02 0.00 -0.25 30.11 -0.04 0.00 0.00 269 -8.22 0.00 -2.04 0.00 -0.25 0.00 -0.04 0.00 0.00 270 -8.22 0.00 -2.04 0.00 -0.25 0.00 -0.04 0.00 0.00 271 -7.35 0.00 -2.35 0.00 -0.29 0.00 -0.05 0.00 0.00 272 -7.35 0.00 -2.35 0.00 -0.29 0.00 -0.05 0.00 0.00 273 -7.35 0.00 -2.35 0.00 -0.29 0.00 -0.05 0.00 0.00 274 -7.60 0.00 -2.27 0.00 -0.28 0.00 -0.05 0.00 0.00 275 -7.60 0.00 -2.27 0.00 -0.28 0.00 -0.05 0.00 0.00 276 -7.60 0.00 -2.27 0.00 -0.28 0.00 -0.05 0.00 0.00 277 -7.36 0.00 -2.36 0.00 -0.29 0.00 -0.05 0.00 0.00 278 -7.36 0.00 -2.36 0.00 -0.29 0.00 -0.05 0.00 0.00 279 -7.36 0.00 -2.36 0.00 -0.29 0.00 -0.05 0.00 0.00 280 -6.12 0.00 -2.88 0.00 -0.36 0.00 -0.06 0.00 0.00 281 -6.12 0.00 -2.88 0.00 -0.36 0.00 -0.06 0.00 0.00 282 -4.99 0.00 -3.28 0.00 -0.41 0.00 -0.07 0.00 0.00 283 -5.98 0.00 -2.94 0.00 -0.36 0.00 -0.06 0.00 0.00 284 -5.76 0.00 -3.02 0.00 -0.37 0.00 -0.06 0.00 0.00 285 -5.76 0.00 -3.02 0.00 -0.37 0.00 -0.06 0.00 0.00 286 -4.64 0.00 -3.42 0.00 -0.42 0.00 -0.07 0.00 0.00 287 -5.65 0.00 -3.07 0.00 -0.38 0.00 -0.06 0.00 0.00 288 -5.65 0.00 -3.07 0.00 -0.38 0.00 -0.06 0.00 0.00 289 -5.58 0.00 -3.10 0.00 -0.38 0.00 -0.06 0.00 0.00 290 -5.00 0.00 -3.30 0.00 -0.41 0.00 -0.07 0.00 0.00 291 -4.23 0.00 -3.58 0.00 -0.44 0.00 -0.07 0.00 0.00 292 -4.23 0.00 -3.58 0.00 -0.44 0.00 -0.07 0.00 0.00 293 -4.17 0.00 -3.60 0.00 -0.45 0.00 -0.07 0.00 0.00 294 -4.67 0.00 -3.43 0.00 -0.42 0.00 -0.07 0.00 0.00 295 -4.26 0.00 -3.60 0.00 -0.44 0.00 -0.07 0.00 0.00 296 -4.26 0.00 -3.60 0.00 -0.44 0.00 -0.07 0.00 0.00 297 -4.21 0.00 -3.62 0.00 -0.45 0.00 -0.07 0.00 0.00 298 -3.31 0.00 -3.94 0.00 -0.49 0.00 -0.08 0.00 0.00 299 -3.02 0.00 -4.06 0.00 -0.50 0.00 -0.08 0.00 0.00 300 -3.02 0.00 -4.06 0.00 -0.50 0.00 -0.08 0.00 0.00 301 -4.04 0.00 -3.71 0.00 -0.46 0.00 -0.08 0.00 0.00 302 -4.04 0.00 -3.71 0.00 -0.46 0.00 -0.08 0.00 0.00 303 -3.36 0.00 -3.95 0.00 -0.49 0.00 -0.08 0.00 0.00 304 -3.07 0.00 -4.07 0.00 -0.50 0.00 -0.08 0.00 0.00 305 -3.07 0.00 -4.07 0.00 -0.50 0.00 -0.08 0.00 0.00 306 -3.34 0.00 -3.99 0.00 -0.49 0.00 -0.08 0.00 0.00 307 -3.34 0.00 -3.99 0.00 -0.49 0.00 -0.08 0.00 0.00 308 -0.91 0.00 -4.89 0.00 -0.60 0.00 -0.10 0.00 0.00 309 -0.91 0.00 -4.89 0.00 -0.60 0.00 -0.10 0.00 0.00 310 -2.47 0.00 -4.34 0.00 -0.54 0.00 -0.09 0.00 0.00 311 -0.91 0.00 -4.89 0.00 -0.60 0.00 -0.10 0.00 0.00 312 -2.47 0.00 -4.34 0.00 -0.54 0.00 -0.09 0.00 0.00 313 -2.47 0.00 -4.34 0.00 -0.54 0.00 -0.09 0.00 0.00 314 -0.98 0.00 -4.89 0.00 -0.61 0.00 -0.10 0.00 0.00 315 -0.98 0.00 -4.89 0.00 -0.61 0.00 -0.10 0.00 0.00 316 -0.98 0.00 -4.90 0.00 -0.61 0.00 -0.10 0.00 0.00 317 -1.80 0.00 -4.61 0.00 -0.57 0.00 -0.09 0.00 0.00 318 -1.80 0.00 -4.61 0.00 -0.57 0.00 -0.09 0.00 0.00 319 -1.80 0.00 -4.61 0.00 -0.57 0.00 -0.09 0.00 0.00 320 -1.80 0.00 -4.61 0.00 -0.57 0.00 -0.09 0.00 0.00 321 -1.80 0.00 -4.61 0.00 -0.57 0.00 -0.09 0.00 0.00 322 -1.80 0.00 -4.61 0.00 -0.57 0.00 -0.09 0.00 0.00 323 -1.01 0.00 -4.92 0.00 -0.61 0.00 -0.10 0.00 0.00 324 11.71 0.00 -2.00 0.00 -0.25 0.00 -0.04 0.00 0.00 325 10.00 0.00 -1.23 0.00 -0.15 0.00 -0.02 0.00 0.00 326 10.67 0.00 -1.59 0.00 -0.20 0.00 -0.03 0.00 0.00 327 10.67 0.00 -1.59 0.00 -0.20 0.00 -0.03 0.00 0.00 328 9.64 0.00 -0.99 -2.00 -0.12 0.00 -0.02 0.00 0.00 329 -10.49 0.00 -0.87 -2.00 -0.11 0.00 -0.02 0.00 0.00 330 -10.49 0.00 -0.87 -2.00 -0.11 0.00 -0.02 0.00 0.00 331 9.33 0.00 -7.86 -5.00 -0.18 0.00 -0.03 0.00 0.00 332 -8.88 0.00 -1.36 -2.00 -0.17 0.00 -0.03 0.00 0.00 333 -10.71 0.00 -1.03 0.00 -0.13 0.00 -0.02 0.00 0.00 334 -10.58 0.00 -1.08 0.00 -0.13 0.00 -0.02 0.00 0.00 335 -10.88 0.00 -0.98 0.00 -0.12 0.00 -0.02 0.00 0.00 336 -10.71 0.00 -1.04 0.00 -0.13 0.00 -0.02 0.00 0.00 337 -10.59 0.00 -1.09 0.00 -0.13 0.00 -0.02 0.00 0.00 338 1.95 0.00 -2.42 -10.00 -0.30 0.00 -0.05 0.00 0.00 339 -10.53 0.00 -1.12 0.00 -0.14 0.00 -0.02 0.00 0.00 340 -10.42 0.00 -1.16 0.00 -0.14 0.00 -0.02 0.00 0.00 341 13.21 0.00 -3.25 0.00 -0.40 0.00 -0.07 0.00 0.00 342 13.21 0.00 -3.25 0.00 -0.40 0.00 -0.07 0.00 0.00 343 -10.40 0.00 -0.89 -2.00 -0.11 0.00 -0.02 0.00 0.00 344 -9.20 0.00 -1.37 -2.00 -0.17 0.00 -0.03 0.00 0.00 345 -9.06 0.00 -1.42 -2.00 -0.18 0.00 -0.03 0.00 0.00 346 -9.20 0.00 -1.37 -2.00 -0.17 0.00 -0.03 0.00 0.00 347 -9.08 0.00 -1.42 -2.00 -0.18 0.00 -0.03 0.00 0.00 348 -10.40 0.00 -0.91 -2.00 -0.11 0.00 -0.02 0.00 0.00 349 -10.32 0.00 -0.93 -2.00 -0.12 0.00 -0.02 0.00 0.00 350 6.43 0.00 -1.13 0.00 -0.14 6.48 -0.02 0.00 0.00 351 9.70 8.01 -1.71 0.00 -0.21 0.00 -0.03 0.00 0.00 352 10.54 0.00 -2.12 0.00 -0.26 0.00 -0.04 0.00 0.00 353 4.35 0.00 -2.07 -13.40 -0.26 0.00 -0.04 0.00 0.00 354 4.62 0.00 -2.09 -13.93 -0.26 0.00 -0.04 0.00 0.00 355 -1.56 0.00 -1.15 0.00 -0.14 0.00 -0.02 0.00 0.00 356 -1.51 0.00 -1.18 0.00 -0.15 0.00 -0.02 0.00 0.00 357 0.47 0.00 -5.01 0.00 -0.21 0.00 -0.03 0.00 0.00 358 24.87 0.00 -4.23 -20.00 -0.52 0.00 -0.09 0.00 0.00 359 13.05 0.00 -3.40 0.00 -0.42 0.00 -0.07 0.00 0.00 360 13.05 0.00 -3.40 0.00 -0.42 0.00 -0.07 0.00 0.00 361 4.77 0.00 -1.18 -5.00 -0.15 12.79 -0.02 0.00 0.00 362 2.88 0.00 -3.91 0.00 -0.22 0.00 -0.04 0.00 2.12 363 6.51 0.00 -2.01 -2.00 -0.25 0.00 -0.04 0.00 0.00 364 6.51 0.00 -2.01 -2.00 -0.25 0.00 -0.04 0.00 0.00 365 25.89 0.00 -4.80 -20.00 -0.59 0.00 -0.10 0.00 0.00 366 25.93 0.00 -4.82 -20.00 -0.60 0.00 -0.10 0.00 0.00 367 25.95 0.00 -4.83 -20.00 -0.60 0.00 -0.10 0.00 0.00 368 25.96 0.00 -4.84 -20.00 -0.60 0.00 -0.10 0.00 0.00 369 4.50 0.00 -4.47 -5.00 -0.33 0.00 -0.05 0.00 1.84 370 3.02 0.00 -2.92 0.00 -0.36 0.00 -0.06 0.00 0.00 371 12.56 0.00 -3.25 0.00 -0.40 0.00 -0.07 0.00 0.00 372 12.56 0.00 -3.25 0.00 -0.40 0.00 -0.07 0.00 0.00 373 23.94 0.00 -4.61 -15.00 -0.57 2.16 -0.09 0.00 0.00 374 7.29 0.00 -2.38 -2.00 -0.29 0.00 -0.05 0.00 0.00 375 7.29 0.00 -2.38 -2.00 -0.29 0.00 -0.05 0.00 0.00 376 7.29 0.00 -2.38 -2.00 -0.29 0.00 -0.05 0.00 0.00 377 -5.10 10.37 -1.24 -2.00 -0.15 0.00 -0.02 0.00 0.00 378 -5.10 10.37 -1.24 -2.00 -0.15 0.00 -0.02 0.00 0.00 379 -5.08 10.43 -1.22 -2.00 -0.15 0.00 -0.02 0.00 0.00 380 -5.08 10.43 -1.22 -2.00 -0.15 0.00 -0.02 0.00 0.00 381 -5.08 10.43 -1.22 -2.00 -0.15 0.00 -0.02 0.00 0.00 382 -5.08 10.43 -1.22 -2.00 -0.15 0.00 -0.02 0.00 0.00 383 12.44 0.00 -2.76 -2.00 -0.34 0.00 -0.06 0.00 0.00 384 7.30 0.00 -2.40 -2.00 -0.30 0.00 -0.05 0.00 0.00 385 11.65 0.00 -2.41 -2.00 -0.30 0.00 -0.05 0.00 0.00 386 10.51 2.86 -2.34 0.00 -0.29 0.00 -0.05 0.00 0.00 387 10.51 2.86 -2.34 0.00 -0.29 0.00 -0.05 0.00 0.00 388 11.74 0.00 -2.93 0.00 -0.36 0.00 -0.06 0.00 0.00 389 27.12 0.00 -5.49 -20.00 -0.68 0.00 -0.11 0.00 0.00 390 27.12 0.00 -5.49 -20.00 -0.68 0.00 -0.11 0.00 0.00 391 27.15 0.00 -5.51 -20.00 -0.68 0.00 -0.11 0.00 0.00 392 11.83 0.00 -2.99 0.00 -0.37 0.00 -0.06 0.00 0.00 393 11.83 0.00 -2.99 0.00 -0.37 0.00 -0.06 0.00 0.00 394 27.21 0.00 -5.54 -20.00 -0.69 0.00 -0.11 0.00 0.00 395 0.94 0.00 -3.26 -10.00 -0.40 0.00 -0.07 0.00 0.00 396 8.29 0.00 -1.32 0.00 -0.16 0.00 -0.03 0.00 0.00 397 8.31 0.00 -1.34 0.00 -0.17 0.00 -0.03 0.00 0.00 398 8.31 0.00 -1.34 0.00 -0.17 0.00 -0.03 0.00 0.00 399 8.31 0.00 -1.34 0.00 -0.17 0.00 -0.03 0.00 0.00 400 7.07 0.00 -0.99 0.00 -0.12 0.00 -0.02 0.00 0.00 401 7.11 0.00 -1.00 0.00 -0.12 0.00 -0.02 0.00 0.00 402 7.11 0.00 -1.00 0.00 -0.12 0.00 -0.02 0.00 0.00 403 7.11 0.00 -1.00 0.00 -0.12 0.00 -0.02 0.00 0.00 404 8.61 0.00 -1.49 0.00 -0.18 0.00 -0.03 0.00 0.00 405 8.61 0.00 -1.49 0.00 -0.18 0.00 -0.03 0.00 0.00 406 8.61 0.00 -1.49 0.00 -0.18 0.00 -0.03 0.00 0.00 407 10.57 0.00 -2.42 0.00 -0.30 0.00 -0.05 0.00 0.00 408 8.62 0.00 -1.49 0.00 -0.18 0.00 -0.03 0.00 0.00 409 8.62 0.00 -1.49 0.00 -0.18 0.00 -0.03 0.00 0.00 410 8.62 0.00 -1.49 0.00 -0.18 0.00 -0.03 0.00 0.00 411 8.73 0.00 -1.55 0.00 -0.19 0.00 -0.03 0.00 0.00 412 8.73 0.00 -1.55 0.00 -0.19 0.00 -0.03 0.00 0.00 413 8.73 0.00 -1.55 0.00 -0.19 0.00 -0.03 0.00 0.00 414 8.74 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 415 8.74 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 416 8.74 0.00 -1.56 0.00 -0.19 0.00 -0.03 0.00 0.00 417 9.67 0.00 -2.81 -2.00 -0.35 0.00 -0.06 0.00 0.00 418 9.50 0.00 -1.95 0.00 -0.24 0.00 -0.04 0.00 0.00 419 9.50 0.00 -1.95 0.00 -0.24 0.00 -0.04 0.00 0.00
TABLE-US-00007 TABLE 7 Knockout strategies derived by OptKnock assuming PEP carboxykinase to be reversible. Metabolic Transformations Targeted For Removal .dagger., .dagger-dbl., # BDO BIO AC ALA CO2 FOR FUM GLC GLY H+ 1 ADHEr, 6.44 0.72 22.08 0.00 1.90 17.19 0.00 -20.00 0.00 44.88 NADH6 2 ADHEr, 6.29 0.03 33.29 0.00 -6.29 33.43 0.00 -20.00 0.00 66.92 ENO 3 ADHEr, 6.29 0.03 33.29 0.00 -6.29 33.43 0.00 -20.00 0.00 66.92 PGM 4 ADHEr, 5.67 0.57 25.42 0.00 -5.74 28.42 0.00 -20.00 0.00 57.91 PPCK 5 ADHEr, 5.54 0.65 24.41 0.00 -5.63 27.79 0.00 -20.00 0.00 56.80 SUCD4 6 ADHEr, 4.41 0.82 22.86 0.00 -4.52 27.15 0.00 -20.00 0.00 55.83 ATPS4r 7 ADHEr, 1.81 0.52 0.00 0.00 -11.12 2.74 0.00 -20.00 0.00 56.18 PGI 8 ADHEr, 1.32 0.74 14.95 0.00 -13.58 18.85 0.00 -20.00 0.00 63.42 FUM 9 ADHEr, 0.82 0.77 13.74 0.00 -14.47 17.74 0.00 -20.00 0.00 64.01 HEX1 10 ADHEr, 0.72 0.67 14.49 0.00 -15.13 18.02 0.00 -20.00 0.00 65.94 MDH 11 ADHEr, 0.33 0.49 15.60 0.00 -16.86 18.16 0.00 -20.00 0.00 70.17 TPI 12 ADHEr, 0.33 0.49 15.60 0.00 -16.86 18.16 0.00 -20.00 0.00 70.17 FBA 13 ADHEr, 0.33 0.49 15.60 0.00 -16.86 18.16 0.00 -20.00 0.00 70.17 PFK 14 ADHEr, 11.52 0.39 15.75 0.00 8.35 17.80 0.00 -20.00 0.00 36.34 HEX1, PGI 15 ADHEr, 9.95 0.50 22.31 0.00 14.90 0.00 0.00 -20.00 0.00 25.85 PFLi, PPCK 16 ADHEr, 9.81 0.55 21.71 0.00 14.68 0.00 0.00 -20.00 0.00 25.59 PFLi, SUCD4 17 ADHEr, 9.81 0.86 10.27 0.00 24.66 0.00 0.00 -20.00 0.00 16.36 ACKr, NADH6 18 ADHEr, 9.71 0.58 21.27 0.00 14.52 0.00 0.00 -20.00 0.00 25.40 NADH6, PFLi 19 ADHEr, 7.82 0.08 30.86 0.00 9.99 13.48 0.00 -20.00 0.00 44.95 NADH6, PGM 20 ADHEr, 7.82 0.08 30.86 0.00 9.99 13.48 0.00 -20.00 0.00 44.95 ENO, NADH6 21 ADHEr, 7.51 0.53 20.76 0.00 -4.06 30.56 0.00 -20.00 0.00 55.08 ASPT, MDH 22 ADHEr, 7.36 0.35 27.02 0.00 0.27 21.51 0.00 -20.00 0.00 51.05 NADH6, PGI 23 ADHEr, 7.25 0.36 27.17 0.00 -0.05 21.80 0.00 -20.00 0.00 51.52 NADH6, TPI 24 ADHEr, 7.25 0.36 27.17 0.00 -0.05 21.80 0.00 -20.00 0.00 51.52 FBA, NADH6 25 ADHEr, 7.25 0.36 27.17 0.00 -0.05 21.80 0.00 -20.00 0.00 51.52 NADH6, PFK 26 ADHEr, 6.92 0.52 25.03 0.00 -0.07 20.82 0.00 -20.00 0.00 49.54 NADH6, PPCK 27 ADHEr, 6.76 0.59 24.04 0.00 -0.08 20.37 0.00 -20.00 0.00 48.62 MDH, NADH6 28 ADHEr, 6.60 0.67 23.01 0.00 -0.09 19.90 0.00 -20.00 0.00 47.66 FUM, NADH6 29 ADHEr, 6.55 0.56 23.75 0.00 -3.54 26.67 0.00 -20.00 0.00 54.38 PPCK, THD2 30 ADHEr, 6.53 0.77 21.10 0.00 5.45 13.37 0.00 -20.00 0.00 40.40 NADH6, RPE 31 ADHEr, 6.48 0.77 21.41 0.00 5.18 13.97 0.00 -20.00 0.00 40.88 NADH6, TAL 32 ADHEr, 6.21 0.30 28.99 0.00 -5.98 30.57 0.00 -20.00 0.00 61.72 PGI, PPCK 33 ADHEr, 6.18 0.33 28.63 0.00 -5.93 30.35 0.00 -20.00 0.00 61.31 PGI, SUCD4 34 ADHEr, 6.15 0.61 23.38 0.00 -3.10 26.55 0.00 -20.00 0.00 54.24 ATPS4r, PPCK 35 ADHEr, 6.13 0.31 29.10 0.00 -6.17 30.70 0.00 -20.00 0.00 61.98 PFK, PPCK 36 ADHEr, 6.13 0.31 29.10 0.00 -6.17 30.70 0.00 -20.00 0.00 61.98 FBA, PPCK 37 ADHEr, 6.13 0.31 29.10 0.00 -6.17 30.70 0.00 -20.00 0.00 61.98 PPCK, TPI 38 ADHEr, 6.11 0.32 28.91 0.00 -6.15 30.59 0.00 -20.00 0.00 61.78 FBA, HEX1 39 ADHEr, 6.11 0.32 28.91 0.00 -6.15 30.59 0.00 -20.00 0.00 61.78 HEX1, PFK 40 ADHEr, 6.11 0.32 28.91 0.00 -6.15 30.59 0.00 -20.00 0.00 61.78 HEX1, TPI 41 ADHEr, 6.10 0.62 14.90 0.00 -3.07 18.16 0.00 -20.00 0.00 49.89 MDH, THD2 42 ADHEr, 6.09 0.33 28.75 0.00 -6.13 30.48 0.00 -20.00 0.00 61.59 SUCD4, TPI 43 ADHEr, 6.09 0.33 28.75 0.00 -6.13 30.48 0.00 -20.00 0.00 61.59 FBA, SUCD4 44 ADHEr, 6.09 0.33 28.75 0.00 -6.13 30.48 0.00 -20.00 0.00 61.59 PFK, SUCD4 45 ADHEr, 5.96 0.70 15.28 0.00 4.93 0.00 0.00 -20.00 0.00 36.10 FUM, PFLi 46 ADHEr, 5.85 0.47 26.89 0.00 -5.92 29.33 0.00 -20.00 0.00 59.54 PPCK, SUCD4 47 ADHEr, 5.78 0.57 25.20 0.00 -5.45 28.19 0.00 -20.00 0.00 57.44 PPCK, RPE 48 ADHEr, 5.78 0.51 26.33 0.00 -5.85 28.99 0.00 -20.00 0.00 58.92 GLCpts, PPCK 49 ADHEr, 5.78 0.51 26.30 0.00 -5.85 28.97 0.00 -20.00 0.00 58.89 GLUDy, MDH 50 ADHEr, 5.77 0.52 26.20 0.00 -5.84 28.90 0.00 -20.00 0.00 58.78 GLUDy, PPCK 51 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.78 0.00 -20.00 0.00 58.55 MDH, SUCD4 52 ADHEr, 5.73 0.57 25.30 0.00 -5.59 28.30 0.00 -20.00 0.00 57.67 PPCK, TAL 53 ADHEr, 5.73 0.54 25.90 0.00 -5.80 28.72 0.00 -20.00 0.00 58.44 FUM, PPCK 54 ADHEr, 5.73 0.54 25.90 0.00 -5.80 28.72 0.00 -20.00 0.00 58.44 MDH, PPCK 55 ADHEr, 5.68 0.64 24.22 0.00 -5.31 27.57 0.00 -20.00 0.00 56.34 RPE, SUCD4 56 ADHEr, 5.68 0.57 25.49 0.00 -5.75 28.46 0.00 -20.00 0.00 57.99 ME2, SUCD4 57 ADHEr, 5.66 0.58 25.35 0.00 -5.74 28.38 0.00 -20.00 0.00 57.84 FUM, GLUDy 58 ADHEr, 5.66 0.58 25.34 0.00 -5.74 28.37 0.00 -20.00 0.00 57.83 GLUDy, SUCD4 59 ADHEr, 5.65 0.58 25.28 0.00 -5.73 28.34 0.00 -20.00 0.00 57.77 GLCpts, SUCD4 60 ADHEr, 5.61 0.64 24.31 0.00 -5.46 27.67 0.00 -20.00 0.00 56.56 SUCD4, TAL 61 ADHEr, 5.57 0.63 24.65 0.00 -5.66 27.94 0.00 -20.00 0.00 57.07 FUM, SUCD4 62 ADHEr, 5.56 0.64 24.55 0.00 -5.64 27.88 0.00 -20.00 0.00 56.95 HEX1, SUCD4 63 ADHEr, 5.55 0.64 24.49 0.00 -5.64 27.84 0.00 -20.00 0.00 56.89 CBMK2, SUCD4 64 ADHEr, 5.44 0.70 23.61 0.00 -5.53 27.29 0.00 -20.00 0.00 55.91 FUM, HEX1 65 ADHEr, 5.24 0.72 14.09 0.00 3.10 0.00 0.00 -20.00 0.00 38.10 HEX1, PFLi 66 ADHEr, 5.06 0.52 26.72 0.00 -4.66 29.44 0.00 -20.00 0.00 59.85 ATPS4r, PGI 67 ADHEr, 4.91 0.53 26.89 0.00 -4.99 29.65 0.00 -20.00 0.00 60.29 ATPS4r, FBA 68 ADHEr, 4.91 0.53 26.89 0.00 -4.99 29.65 0.00 -20.00 0.00 60.29 ATPS4r, PFK 69 ADHEr, 4.91 0.53 26.89 0.00 -4.99 29.65 0.00 -20.00 0.00 60.29 ATPS4r, TPI 70 ADHEr, 4.88 0.44 16.04 0.00 1.18 0.00 0.00 -20.00 0.00 43.62 PFLi, TPI 71 ADHEr, 4.88 0.44 16.04 0.00 1.18 0.00 0.00 -20.00 0.00 43.62 PFK, PFLi 72 ADHEr, 4.88 0.44 16.04 0.00 1.18 0.00 0.00 -20.00 0.00 43.62 FBA, PFLi 73 ADHEr, 4.87 0.74 22.12 0.00 -6.49 25.99 0.00 -20.00 0.00 56.39 HEX1, THD2 74 ADHEr, 4.59 0.81 22.69 0.00 -4.12 26.91 0.00 -20.00 0.00 55.33 ATPS4r, RPE 75 ADHEr, 4.56 0.73 24.10 0.00 -4.66 27.92 0.00 -20.00 0.00 57.20 ATPS4r, GLUDy 76 ADHEr, 4.50 0.81 22.77 0.00 -4.31 27.02 0.00 -20.00 0.00 55.57 ATPS4r, TAL 77 ADHEr, 4.42 0.81 22.97 0.00 -4.53 27.22 0.00 -20.00 0.00 55.95 ATPS4r, CBMK2 78 ADHEr, 4.21 0.51 0.00 0.00 -5.69 2.64 0.00 -20.00 0.00 48.84 EDA, PGI 79 ADHEr, 2.48 0.52 0.00 0.00 -8.45 0.00 0.00 -20.00 0.00 52.22 PFLi, PGI 80 ADHEr, 1.36 0.47 17.83 0.00 -14.86 20.31 0.00 -20.00 0.00 68.38 MDH, PFK 81 ADHEr, 1.36 0.47 17.83 0.00 -14.86 20.31 0.00 -20.00 0.00 68.38 MDH, TPI 82 ADHEr, 1.36 0.47 17.83 0.00 -14.86 20.31 0.00 -20.00 0.00 68.38 FBA, MDH 83 ADHEr, 1.13 0.76 13.77 0.00 -13.77 17.76 0.00 -20.00 0.00 63.09 HEX1, RPE 84 ADHEr, 1.00 0.67 14.51 0.00 -14.51 18.02 0.00 -20.00 0.00 65.11 MDH, RPE 85 ADHEr, 0.98 0.76 13.76 0.00 -14.10 17.75 0.00 -20.00 0.00 63.53 HEX1, TAL 86 ADHEr, 0.86 0.67 14.50 0.00 -14.81 18.02 0.00 -20.00 0.00 65.51 MDH, TAL 87 ADHEr, 0.47 0.48 15.53 0.00 -16.54 18.07 0.00 -20.00 0.00 69.75 RPE, TPI 88 ADHEr, 0.47 0.48 15.53 0.00 -16.54 18.07 0.00 -20.00 0.00 69.75 PFK, RPE 89 ADHEr, 0.47 0.48 15.53 0.00 -16.54 18.07 0.00 -20.00 0.00 69.75 FBA, RPE 90 ADHEr, 0.40 0.49 15.57 0.00 -16.69 18.11 0.00 -20.00 0.00 69.95 PFK, TAL 91 ADHEr, 0.40 0.49 15.57 0.00 -16.69 18.11 0.00 -20.00 0.00 69.95 FBA, TAL
92 ADHEr, 14.20 0.34 13.81 0.00 21.29 0.00 0.00 -20.00 0.00 16.25 HEX1, PFLi, PGI 93 ADHEr, 14.07 0.28 14.98 0.00 19.90 2.36 0.00 -20.00 0.00 19.31 HEX1, NADH6, PGI 94 ADHEr, 14.00 0.31 14.58 0.00 19.89 2.18 0.00 -20.00 0.00 18.94 EDA, NADH6, PGI 95 ADHEr, 13.92 0.34 14.10 0.00 21.85 0.00 0.00 -20.00 0.00 16.55 ACKr, NADH6, PGI 96 ADHEr, 13.31 0.25 5.13 0.00 13.27 19.78 0.00 -20.00 0.00 28.86 FRD2, LDH_D, MDH 97 ADHEr, 13.14 0.27 15.98 0.00 17.23 6.93 0.00 -20.00 0.00 24.84 ATPS4r, PGI, SUCD4 98 ADHEr, 12.33 0.49 15.46 0.00 19.46 0.00 0.00 -20.00 0.00 18.94 ATPS4r, FDH2, NADH6 99 ADHEr, 12.03 0.46 12.29 0.00 28.02 0.00 0.00 -20.00 0.00 15.58 ACKr, NADH6, TPI 100 ADHEr, 12.03 0.46 12.29 0.00 28.02 0.00 0.00 -20.00 0.00 15.58 ACKr, FBA, NADH6 101 ADHEr, 12.03 0.46 12.29 0.00 28.02 0.00 0.00 -20.00 0.00 15.58 ACKr, NADH6, PFK 102 ADHEr, 11.95 0.26 4.09 0.00 7.62 17.42 4.30 -20.00 0.00 33.78 FRD2, LDH_D, ME2 103 ADHEr, 11.82 0.22 18.07 0.00 8.10 19.23 0.00 -20.00 0.00 38.88 HEX1, PGI, PPCK 104 ADHEr, 11.82 0.22 18.05 0.00 8.10 19.22 0.00 -20.00 0.00 38.86 EDA, PGI, PPCK 105 ADHEr, 11.77 0.25 17.66 0.00 8.15 18.98 0.00 -20.00 0.00 38.43 HEX1, PGI, SUCD4 106 ADHEr, 11.76 0.26 17.62 0.00 8.15 18.95 0.00 -20.00 0.00 38.39 EDA, PGI, SUCD4 107 ADHEr, 11.62 0.34 16.52 0.00 8.27 18.28 0.00 -20.00 0.00 37.18 ATPS4r, EDA, PGI 108 ADHEr, 11.59 0.35 16.29 0.00 8.29 18.13 0.00 -20.00 0.00 36.93 GLUDy, HEX1, PGI 109 ADHEr, 11.18 0.48 13.34 0.00 4.81 23.87 0.00 -20.00 0.00 40.62 MDH, PGL, THD2 110 ADHEr, 11.18 0.48 13.34 0.00 4.81 23.87 0.00 -20.00 0.00 40.62 G6PDHy, MDH, THD2 111 ADHEr, 10.87 0.20 25.96 0.00 16.29 0.00 0.00 -20.00 0.00 27.38 PFLi, PGI, PPCK 112 ADHEr, 10.83 0.20 26.02 0.00 16.23 0.00 0.00 -20.00 0.00 27.46 PFLi, PPCK, TPI 113 ADHEr, 10.83 0.20 26.02 0.00 16.23 0.00 0.00 -20.00 0.00 27.46 FBA, PFLi, PPCK 114 ADHEr, 10.83 0.20 26.02 0.00 16.23 0.00 0.00 -20.00 0.00 27.46 PFK, PFLi, PPCK 115 ADHEr, 10.80 0.74 11.20 0.00 23.65 0.00 0.00 -20.00 0.00 16.47 ACKr, MDH, NADH6 116 ADHEr, 10.79 0.23 25.57 0.00 16.16 0.00 0.00 -20.00 0.00 27.21 NADH6, PFLi, PGI 117 ADHEr, 10.79 0.23 25.57 0.00 16.16 0.00 0.00 -20.00 0.00 27.21 PFLi, PGI, SUCD4 118 ADHEr, 10.74 0.23 25.64 0.00 16.09 0.00 0.00 -20.00 0.00 27.30 FBA, PFLi, SUCD4 119 ADHEr, 10.74 0.23 25.64 0.00 16.09 0.00 0.00 -20.00 0.00 27.30 PFK, PFLi, SUCD4 120 ADHEr, 10.74 0.23 25.64 0.00 16.09 0.00 0.00 -20.00 0.00 27.30 NADH6, PFK, PFLi 121 ADHEr, 10.74 0.23 25.64 0.00 16.09 0.00 0.00 -20.00 0.00 27.30 FBA, NADH6, PFLi 122 ADHEr, 10.74 0.23 25.64 0.00 16.09 0.00 0.00 -20.00 0.00 27.30 PFLi, SUCD4, TPI 123 ADHEr, 10.74 0.23 25.64 0.00 16.09 0.00 0.00 -20.00 0.00 27.30 NADH6, PFLi, TPI 124 ADHEr, 10.73 0.24 25.61 0.00 16.08 0.00 0.00 -20.00 0.00 27.28 HEX1, PFK, PFLi 125 ADHEr, 10.73 0.24 25.61 0.00 16.08 0.00 0.00 -20.00 0.00 27.28 FBA, HEX1, PFLi 126 ADHEr, 10.73 0.24 25.61 0.00 16.08 0.00 0.00 -20.00 0.00 27.28 HEX1, PFLi, TPI 127 ADHEr, 10.49 0.49 21.02 0.00 15.71 0.00 0.00 -20.00 0.00 24.49 PFLi, PPCK, THD2 128 ADHEr, 10.40 0.75 10.81 0.00 25.55 0.00 0.00 -20.00 0.00 16.16 ACKr, GLUDy, NADH6 129 ADHEr, 10.28 0.77 10.70 0.00 25.38 0.00 0.00 -20.00 0.00 16.20 ACKr, GLCpts, NADH6 130 ADHEr, 10.24 0.58 10.55 0.00 11.67 22.30 0.00 -20.00 0.00 36.95 ACKr, AKGD, ATPS4r 131 ADHEr, 10.17 0.53 19.34 0.00 15.23 0.00 0.00 -20.00 0.00 24.21 ATPS4r, NADH6, PFLi 132 ADHEr, 10.15 0.43 23.16 0.00 15.20 0.00 0.00 -20.00 0.00 26.22 GLCpts, PFLi, PPCK 133 ADHEr, 10.12 0.58 10.44 0.00 10.60 23.88 0.00 -20.00 0.00 38.45 ACKr, ATPS4r, SUCOAS 134 ADHEr, 10.11 0.77 10.53 0.00 22.07 0.00 0.00 -20.00 0.00 18.19 ACKr, ME2, NADH6 135 ADHEr, 10.10 0.45 22.94 0.00 15.12 0.00 0.00 -20.00 0.00 26.12 GLUDy, PFLi, PPCK 136 ADHEr, 10.05 0.47 22.71 0.00 15.04 0.00 0.00 -20.00 0.00 26.02 ME2, PFLi, SUCD4 137 ADHEr, 10.04 0.47 22.67 0.00 15.02 0.00 0.00 -20.00 0.00 26.01 MDH, NADH6, PFLi 138 ADHEr, 10.02 0.50 22.14 0.00 15.00 0.00 0.00 -20.00 0.00 25.67 PFLi, PPCK, RPE 139 ADHEr, 9.99 0.50 22.22 0.00 14.95 0.00 0.00 -20.00 0.00 25.76 PFLi, PPCK, TAL 140 ADHEr, 9.98 0.49 22.45 0.00 14.95 0.00 0.00 -20.00 0.00 25.91 GLUDy, PFLi, SUCD4 141 ADHEr, 9.96 0.49 22.36 0.00 14.92 0.00 0.00 -20.00 0.00 25.87 CBMK2, PFLi, PPCK 142 ADHEr, 9.92 0.57 20.45 0.00 15.84 0.00 0.00 -20.00 0.00 24.48 ATPS4r, LDH_D, SUCD4 143 ADHEr, 9.90 0.54 21.57 0.00 14.81 0.00 0.00 -20.00 0.00 25.41 PFLi, RPE, SUCD4 144 ADHEr, 9.86 0.85 10.32 0.00 24.74 0.00 0.00 -20.00 0.00 16.34 ACKr, CBMK2, NADH6 145 ADHEr, 9.86 0.54 21.64 0.00 14.75 0.00 0.00 -20.00 0.00 25.50 PFLi, SUCD4, TAL 146 ADHEr, 9.86 0.85 10.32 0.00 24.73 0.00 0.00 -20.00 0.00 16.34 ACKr, NADH6, RPE 147 ADHEr, 9.84 0.85 10.30 0.00 24.70 0.00 0.00 -20.00 0.00 16.35 ACKr, FUM, NADH6 148 ADHEr, 9.83 0.85 10.30 0.00 24.70 0.00 0.00 -20.00 0.00 16.35 ACKr, NADH6, TAL 149 ADHEr, 9.83 0.85 10.29 0.00 24.69 0.00 0.00 -20.00 0.00 16.35 ACKr, ASNS2, NADH6 150 ADHEr, 9.83 0.54 21.78 0.00 14.70 0.00 0.00 -20.00 0.00 25.62 CBMK2, PFLi, SUCD4 151 ADHEr, 9.83 0.85 10.29 0.00 24.68 0.00 0.00 -20.00 0.00 16.36 ACKr, NADH12, NADH6 152 ADHEr, 9.82 0.85 10.28 0.00 24.67 0.00 0.00 -20.00 0.00 16.36 ACKr, NADH6, SO4t2 153 ADHEr, 9.81 0.55 21.71 0.00 14.68 0.00 0.00 -20.00 0.00 25.59 NADH12, NADH6, PFLi 154 ADHEr, 9.80 0.55 21.69 0.00 14.67 0.00 0.00 -20.00 0.00 25.58 FUM, NADH6, PFLi 155 ADHEr, 9.80 0.22 9.92 0.00 12.33 11.06 0.00 -20.00 0.00 30.56 ACKr, PGI, SUCD4 156 ADHEr, 9.75 0.58 21.19 0.00 14.60 0.00 0.00 -20.00 0.00 25.30 NADH6, PFLi, TAL 157 ADHEr, 9.72 0.58 21.33 0.00 14.55 0.00 0.00 -20.00 0.00 25.43 CBMK2, NADH6, PFLi 158 ADHEr, 9.55 0.63 20.62 0.00 14.29 0.00 0.00 -20.00 0.00 25.11 FUM, HEX1, PFLi 159 ADHEr, 9.34 0.55 19.65 0.00 7.38 13.18 0.00 -20.00 0.00 36.73 MDH, NADH6, THD2 160 ADHEr, 9.28 0.55 19.75 0.00 7.20 13.35 0.00 -20.00 0.00 37.01 ATPS4r,
MDH, NADH6 161 ADHEr, 9.03 0.63 21.22 0.00 11.62 3.77 0.00 -20.00 0.00 29.45 ATPS4r, FUM, NADH6 162 ADHEr, 8.97 0.26 23.60 0.00 0.95 24.98 0.00 -20.00 0.00 50.46 ATPS4r, PGI, PPCK 163 ADHEr, 8.65 0.48 19.82 0.00 -0.06 26.01 0.00 -20.00 0.00 49.23 ASPT, MDH, NADH6 164 ADHEr, 8.62 0.48 23.04 0.00 6.28 13.23 0.00 -20.00 0.00 39.68 ATPS4r, NADH6, PPCK 165 ADHEr, 8.60 0.50 18.60 0.00 -1.82 29.38 0.00 -20.00 0.00 51.55 ASPT, MDH, THD2 166 ADHEr, 8.49 0.65 21.41 0.00 14.16 2.07 0.00 -20.00 0.00 28.11 ATPS4r, GLCpts, SUCD4 167 ADHEr, 8.36 0.23 23.87 0.00 -4.10 33.24 0.00 -20.00 0.00 58.77 ASPT, MDH, PGI 168 ADHEr, 8.27 0.24 24.01 0.00 -4.27 33.31 0.00 -20.00 0.00 59.01 ASPT, FBA, MDH 169 ADHEr, 8.27 0.24 24.01 0.00 -4.27 33.31 0.00 -20.00 0.00 59.01 ASPT, MDH, PFK 170 ADHEr, 8.27 0.24 24.01 0.00 -4.27 33.31 0.00 -20.00 0.00 59.01 ASPT, MDH, TPI 171 ADHEr, 8.26 0.26 27.70 0.00 3.99 16.78 0.00 -20.00 0.00 46.33 ATPS4r, PFK, PPCK 172 ADHEr, 8.26 0.26 27.70 0.00 3.99 16.78 0.00 -20.00 0.00 46.33 ATPS4r, FBA, PPCK 173 ADHEr, 8.26 0.26 27.70 0.00 3.99 16.78 0.00 -20.00 0.00 46.33 ATPS4r, PPCK, TPI 174 ADHEr, 8.12 0.45 8.36 0.00 1.85 10.72 0.00 -20.00 0.00 42.04 ACKr, EDA, PGI 175 ADHEr, 7.99 0.67 21.56 0.00 6.75 10.40 0.00 -20.00 0.00 36.71 ATPS4r, HEX1, NADH6 176 ADHEr, 7.88 0.50 23.49 0.00 2.67 18.21 0.00 -20.00 0.00 45.24 NADH6, PPCK, THD2 177 ADHEr, 7.87 0.46 22.87 0.00 -0.07 23.70 0.00 -20.00 0.00 49.84 ATPS4r, GLUDy, MDH 178 ADHEr, 7.86 0.49 22.44 0.00 0.04 23.43 0.00 -20.00 0.00 49.33 ATPS4r, MDH, PPCK 179 ADHEr, 7.86 0.49 22.44 0.00 0.04 23.43 0.00 -20.00 0.00 49.33 ATPS4r, FUM, PPCK 180 ADHEr, 7.84 0.08 30.83 0.00 10.05 13.42 0.00 -20.00 0.00 44.85 ENO, NADH6, RPE 181 ADHEr, 7.84 0.08 30.83 0.00 10.05 13.42 0.00 -20.00 0.00 44.85 NADH6, PGM, RPE 182 ADHEr, 7.83 0.08 30.84 0.00 10.02 13.45 0.00 -20.00 0.00 44.90 NADH6, PGM, TAL 183 ADHEr, 7.83 0.08 30.84 0.00 10.02 13.45 0.00 -20.00 0.00 44.90 ENO, NADH6, TAL 184 ADHEr, 7.70 0.46 21.57 0.00 -4.11 31.25 0.00 -20.00 0.00 56.07 ASPT, GLCpts, MDH 185 ADHEr, 7.65 0.52 20.47 0.00 -3.75 30.40 0.00 -20.00 0.00 54.60 ASPT, MDH, RPE 186 ADHEr, 7.65 0.47 21.39 0.00 -4.10 31.10 0.00 -20.00 0.00 55.85 ASPT, GLUDy, MDH 187 ADHEr, ME2, 7.62 0.87 17.24 0.00 18.87 0.00 0.00 -20.00 0.00 23.43 NADH6, THD2 188 ADHEr, ME2, 7.61 0.52 22.02 0.00 -0.99 24.76 0.00 -20.00 0.00 50.50 SUCD4, THD2 189 ADHEr, 7.58 0.53 20.61 0.00 -3.90 30.48 0.00 -20.00 0.00 54.83 ASPT, MDH, TAL 190 ADHEr, 7.58 0.23 28.71 0.00 0.18 22.36 0.00 -20.00 0.00 52.73 NADH6, PGI, PPCK 191 ADHEr, 7.55 0.49 22.66 0.00 -1.33 25.25 0.00 -20.00 0.00 51.43 FUM, PPCK, THD2 192 ADHEr, 7.55 0.49 22.66 0.00 -1.33 25.25 0.00 -20.00 0.00 51.43 MDH, PPCK, THD2 193 ADHEr, 7.56 0.47 23.10 0.00 -1.46 25.55 0.00 -20.00 0.00 51.99 GLUDy, MDH, THD2 194 ADHEr, 7.52 0.52 20.82 0.00 -4.06 30.61 0.00 -20.00 0.00 55.15 ASPT, CBMK2, MDH 195 ADHEr, 7.51 0.42 25.99 0.00 6.86 13.76 0.00 -20.00 0.00 42.72 ATPS4r, FBA, SUCD4 196 ADHEr, 7.51 0.42 25.99 0.00 6.86 13.76 0.00 -20.00 0.00 42.72 ATPS4r, PFK, SUCD4 197 ADHEr, 7.51 0.42 25.99 0.00 6.86 13.76 0.00 -20.00 0.00 42.72 ATPS4r, SUCD4, TPI 198 ADHEr, FBA, 7.50 0.24 28.80 0.00 -0.03 22.54 0.00 -20.00 0.00 53.03 NADH6, PPCK 199 ADHEr, 7.50 0.24 28.80 0.00 -0.03 22.54 0.00 -20.00 0.00 53.03 NADH6, PFK, PPCK 200 ADHEr, 7.50 0.24 28.80 0.00 -0.03 22.54 0.00 -20.00 0.00 53.03 NADH6, PPCK, TPI 201 ADHEr, 7.47 0.69 14.32 0.00 8.09 0.00 0.00 -20.00 0.00 31.52 HEX1, PFLi, THD2 202 ADHEr, 7.46 0.26 28.53 0.00 -0.04 22.42 0.00 -20.00 0.00 52.78 HEX1, NADH6, PFK 203 ADHEr, FBA, 7.46 0.26 28.53 0.00 -0.04 22.42 0.00 -20.00 0.00 52.78 HEX1, NADH6 204 ADHEr, 7.46 0.26 28.53 0.00 -0.04 22.42 0.00 -20.00 0.00 52.78 HE.chi.1, NADH6, TPI 205 ADHEr, 7.44 0.58 13.04 0.00 6.96 0.00 0.00 -20.00 0.00 33.76 ATPS4r, G6PDHy, MDH 206 ADHEr, 7.44 0.58 13.04 0.00 6.96 0.00 0.00 -20.00 0.00 33.76 ATPS4r, MDH, PGL 207 ADHEr, 7.43 0.32 27.55 0.00 0.24 21.78 0.00 -20.00 0.00 51.58 GLUDy, NADH6, PGI 208 ADHEr, 7.42 0.97 7.95 0.00 15.46 13.33 0.00 -20.00 0.00 29.24 ACKr, FRD2, LDH_D 209 ADHEr, 7.42 0.97 7.95 2.12 14.40 13.33 0.00 -20.00 0.00 30.30 ACKr, LDH_D, SUCD4 210 ADHEr, 7.38 0.54 24.25 0.00 2.47 17.14 0.00 -20.00 0.00 45.22 ATPS4r, FUM, GLUDy 211 ADHEr, 7.34 0.36 27.05 0.00 0.21 21.57 0.00 -20.00 0.00 51.14 NADH6, PFK, RPE 212 ADHEr, FBA, 7.34 0.36 27.05 0.00 0.21 21.57 0.00 -20.00 0.00 51.14 NADH6, RPE 213 ADHEr, 7.34 0.36 27.05 0.00 0.21 21.57 0.00 -20.00 0.00 51.14 NADH6, RPE, TPI 214 ADHEr, FBA, 7.33 0.32 27.69 0.00 -0.04 22.04 0.00 -20.00 0.00 52.00 GLUDy, NADH6 215 ADHEr, 7.33 0.32 27.69 0.00 -0.04 22.04 0.00 -20.00 0.00 52.00 GLUDy, NADH6, PFK 216 ADHEr, 7.33 0.32 27.69 0.00 -0.04 22.04 0.00 -20.00 0.00 52.00 GLUDy, NADH6, TPI 217 ADHEr, 7.31 0.66 21.12 0.00 0.90 20.04 0.00 -20.00 0.00 45.88 ATPS4r, FUM, HEX1 218 ADHEr, 7.30 0.36 27.10 0.00 0.09 21.68 0.00 -20.00 0.00 51.32 NADH6, TAL, TPI 219 ADHEr, 7.30 0.36 27.10 0.00 0.09 21.68 0.00 -20.00 0.00 51.32 NADH6, PFK, TAL 220 ADHEr, FBA, 7.30 0.36 27.10 0.00 0.09 21.68 0.00 -20.00 0.00 51.32 NADH6, TAL 221 ADHEr, 7.27 0.62 19.64 0.00 -1.10 22.88 0.00 -20.00 0.00 49.00 ATPS4r, MDH, THD2 222 ADHEr, 7.06 0.45 25.98 0.00 -0.06 21.26 0.00 -20.00 0.00 50.42 GLUDy, MDH, NADH6 223 ADHEr, 7.06 0.45 25.94 0.00 -0.06 21.24 0.00 -20.00 0.00 50.38 GLCpts, NADH6, PPCK 224 ADHEr, 7.05 0.52 24.82 0.00 0.30 20.47 0.00 -20.00 0.00 48.96 NADH6, PPCK, RPE 225 ADHEr, 7.02 0.47 25.73 0.00 -0.06 21.14 0.00 -20.00 0.00 50.18 GLUDy, NADH6, PPCK 226 ADHEr, 7.01 0.47 25.63 0.00 -0.06 21.10 0.00 -20.00 0.00 50.09 FUM, NADH6, PPCK 227 ADHEr, 7.01 0.47 25.63 0.00 -0.06 21.10 0.00 -20.00 0.00 50.09 MDH, NADH6, PPCK 228 ADHEr, 7.00 0.32 0.00 0.00 17.78 8.35 0.00 -20.00 0.00 21.13 ATPS4r, FRD2, LDH_D 229 ADHEr, 6.98 0.52 24.92 0.00 0.12 20.64 0.00 -20.00 0.00 49.23 NADH6, PPCK, TAL 230 ADHEr, 6.92 0.52 25.04 0.00 -0.07 20.83 0.00 -20.00 0.00 49.55 FUM, GLUDy, NADH6 231 ADHEr, 6.91 0.52 24.98 0.00 -0.07 20.80 0.00 -20.00 0.00 49.49 GLCpts, MDH, NADH6 232 ADHEr, 6.91 0.59 23.79 0.00 0.34 19.96 0.00 -20.00 0.00 47.94 MDH, NADH6, RPE 233 ADHEr, 6.84 0.59 23.91 0.00 0.14 20.16 0.00 -20.00 0.00 48.26 MDH, NADH6, TAL 234 ADHEr, 6.83 0.92 18.08 0.00 17.16 1.07 0.00 -20.00 0.00 25.70 HEX1, NADH6, THD2 235 ADHEr, 6.77 0.66 22.78 0.00 0.38 19.47 0.00 -20.00 0.00 46.96 FUM,
NADH6, RPE 236 ADHEr, 6.77 0.59 24.11 0.00 -0.08 20.40 0.00 -20.00 0.00 48.68 CBMK2, MDH, NADH6 237 ADHEr, 6.76 0.59 24.04 0.00 -0.08 20.37 0.00 -20.00 0.00 48.62 FUM, ME2, NADH6 238 ADHEr, 6.69 0.66 22.89 0.00 0.16 19.68 0.00 -20.00 0.00 47.29 FUM, NADH6, TAL 239 ADHEr, 6.64 0.60 16.99 0.00 6.42 0.00 0.00 -20.00 0.00 35.28 ATPS4r, MDH, PGDH 240 ADHEr, 6.64 0.65 23.23 0.00 -0.09 20.01 0.00 -20.00 0.00 47.87 FUM, HEX1, NADH6 241 ADHEr, 6.61 0.66 23.09 0.00 -0.09 19.94 0.00 -20.00 0.00 47.74 CBMK2, FUM, NADH6 242 ADHEr, 6.56 0.49 24.86 0.00 -3.91 27.44 0.00 -20.00 0.00 55.80 GLCpts, PPCK, THD2 243 ADHEr, 6.56 0.50 24.67 0.00 -3.84 27.31 0.00 -20.00 0.00 55.57 GLUDy, PPCK, THD2 244 ADHEr, 6.49 0.77 21.51 0.00 5.18 14.02 0.00 -20.00 0.00 40.97 CBMK2, NADH6, TAL 245 ADHEr, 6.48 0.61 16.56 0.00 5.95 0.00 0.00 -20.00 0.00 35.78 ATPS4r, MDH, TAL 246 ADHEr, 6.45 0.65 22.52 0.00 1.91 16.48 0.00 -20.00 0.00 45.54 ATPS4r, GLUDy, NADH6 247 ADHEr, PGI, 6.33 0.22 30.11 0.00 -6.16 31.28 0.00 -20.00 0.00 62.99 PPCK, SUCD4 248 ADHEr, 6.33 0.61 16.17 0.00 5.52 0.00 0.00 -20.00 0.00 36.25 ATPS4r, MDH, RPE 249 ADHEr, FBP, 6.33 0.03 33.21 0.00 -6.19 33.35 0.00 -20.00 0.00 66.75 PGM, THD2 250 ADHEr, ENO, 6.33 0.03 33.21 0.00 -6.19 33.35 0.00 -20.00 0.00 66.75 FBP, THD2 251 ADHEr, FBA, 6.33 0.03 33.21 0.00 -6.19 33.35 0.00 -20.00 0.00 66.75 PGM, THD2 252 ADHEr, ENO, 6.33 0.03 33.21 0.00 -6.19 33.35 0.00 -20.00 0.00 66.75 FBA, THD2 253 ADHEr, ENO, 6.33 0.03 33.21 0.00 -6.19 33.35 0.00 -20.00 0.00 66.75 THD2, TPI 254 ADHEr, 6.33 0.03 33.21 0.00 -6.19 33.35 0.00 -20.00 0.00 66.75 PGM, THD2, TPI 255 ADHEr, 6.28 0.26 29.59 0.00 -6.08 30.96 0.00 -20.00 0.00 62.41 GLCpts, PGI, PPCK 256 ADHEr, FBA, 6.28 0.22 30.24 0.00 -6.31 31.41 0.00 -20.00 0.00 63.24 HEX1, PPCK 257 ADHEr, 6.28 0.22 30.24 0.00 -6.31 31.41 0.00 -20.00 0.00 63.24 HEX1, PFK, PPCK 258 ADHEr, 6.28 0.22 30.24 0.00 -6.31 31.41 0.00 -20.00 0.00 63.24 HEX1, PPCK, TPI 259 ADHEr, 6.27 0.23 30.17 0.00 -6.30 31.37 0.00 -20.00 0.00 63.17 PPCK, SUCD4, TPI 260 ADHEr, PFK, 6.27 0.23 30.17 0.00 -6.30 31.37 0.00 -20.00 0.00 63.17 PPCK, SUCD4 261 ADHEr, FBA, 6.27 0.23 30.17 0.00 -6.30 31.37 0.00 -20.00 0.00 63.17 PPCK, SUCD4 262 ADHEr, 6.26 0.27 29.44 0.00 -6.06 30.86 0.00 -20.00 0.00 62.23 GLUDy, PGI, PPCK 263 ADHEr, 6.24 0.28 29.26 0.00 -6.03 30.74 0.00 -20.00 0.00 62.02 GLCpts, PGI, SUCD4 264 ADHEr, 6.24 0.29 29.24 0.00 -6.02 30.73 0.00 -20.00 0.00 62.00 FUM, GLUDy, PGI 265 ADHEr, 6.24 0.29 29.24 0.00 -6.02 30.73 0.00 -20.00 0.00 62.00 GLUDy, MDH, PGI 266 ADHEr, FBA, 6.24 0.25 29.95 0.00 -6.27 31.23 0.00 -20.00 0.00 62.93 HEX1, SUCD4 267 ADHEr, 6.24 0.25 29.95 0.00 -6.27 31.23 0.00 -20.00 0.00 62.93 HEX1, SUCD4, TPI 268 ADHEr, 6.24 0.25 29.95 0.00 -6.27 31.23 0.00 -20.00 0.00 62.93 HEX1, PFK, SUCD4 269 ADHEr, 6.23 0.29 29.14 0.00 -6.01 30.67 0.00 -20.00 0.00 61.88 GLUDy, PGI, SUCD4 270 ADHEr, 6.21 0.69 22.25 0.00 -3.64 25.83 0.00 -20.00 0.00 52.95 FUM, HEX1, THD2 271 ADHEr, FBA, 6.21 0.26 29.69 0.00 -6.24 31.07 0.00 -20.00 0.00 62.64 GLCpts, PPCK 272 ADHEr, 6.21 0.26 29.69 0.00 -6.24 31.07 0.00 -20.00 0.00 62.64 GLCpts, PFK, PPCK 273 ADHEr, 6.21 0.26 29.69 0.00 -6.24 31.07 0.00 -20.00 0.00 62.64 GLCpts, PPCK, TPI 274 ADHEr, PFK, 6.20 0.30 29.01 0.00 -6.02 30.60 0.00 -20.00 0.00 61.77 PPCK, RPE 275 ADHEr, 6.20 0.30 29.01 0.00 -6.02 30.60 0.00 -20.00 0.00 61.77 PPCK, RPE, TPI 276 ADHEr, FBA, 6.20 0.30 29.01 0.00 -6.02 30.60 0.00 -20.00 0.00 61.77 PPCK, RPE 277 ADHEr, 6.19 0.27 29.54 0.00 -6.23 30.98 0.00 -20.00 0.00 62.47 GLUDy, PFK, PPCK 278 ADHEr, 6.19 0.27 29.54 0.00 -6.23 30.98 0.00 -20.00 0.00 62.47 GLUDy, PPCK, TPI 279 ADHEr, FBA, 6.19 0.27 29.54 0.00 -6.23 30.98 0.00 -20.00 0.00 62.47 GLUDy, PPCK 280 ADHEr, 6.19 0.71 21.49 0.00 -2.37 25.20 0.00 -20.00 0.00 51.73 ATPS4r, ME2, THD2 281 ADHEr, 6.18 0.32 28.82 0.00 -5.99 30.48 0.00 -20.00 0.00 61.55 HEX1, PFK, RPE 282 ADHEr, 6.18 0.32 28.82 0.00 -5.99 30.48 0.00 -20.00 0.00 61.55 HEX1, RPE, TPI 283 ADHEr, FBA, 6.18 0.32 28.82 0.00 -5.99 30.48 0.00 -20.00 0.00 61.55 HEX1, RPE 284 ADHEr, FBA, 6.17 0.31 29.05 0.00 -6.09 30.65 0.00 -20.00 0.00 61.87 PPCK, TAL 285 ADHEr, 6.17 0.31 29.05 0.00 -6.09 30.65 0.00 -20.00 0.00 61.87 PPCK, TAL, TPI 286 ADHEr, PFK, 6.17 0.31 29.05 0.00 -6.09 30.65 0.00 -20.00 0.00 61.87 PPCK, TAL 287 ADHEr, 6.17 0.55 24.36 0.00 -3.40 27.23 0.00 -20.00 0.00 55.50 ATPS4r, GLUDy, PPCK 288 ADHEr, FBA, 6.17 0.29 29.36 0.00 -6.20 30.87 0.00 -20.00 0.00 62.28 GLUDy, HEX1 289 ADHEr, 6.17 0.29 29.36 0.00 -6.20 30.87 0.00 -20.00 0.00 62.28 GLUDy, HEX1, TPI 290 ADHEr, 6.17 0.29 29.36 0.00 -6.20 30.87 0.00 -20.00 0.00 62.28 GLUDy, HEX1, PFK 291 ADHEr, 6.16 0.29 29.36 0.00 -6.20 30.86 0.00 -20.00 0.00 62.27 GLCpts, SUCD4, TPI 292 ADHEr, FBA, 6.16 0.29 29.36 0.00 -6.20 30.86 0.00 -20.00 0.00 62.27 GLCpts, SUCD4 293 ADHEr, 6.16 0.29 29.36 0.00 -6.20 30.86 0.00 -20.00 0.00 62.27 GLCpts, PFK, SUCD4 294 ADHEr, 6.16 0.29 29.34 0.00 -6.20 30.85 0.00 -20.00 0.00 62.25 GLUDy, MDH, PFK 295 ADHEr, FBA, 6.16 0.29 29.34 0.00 -6.20 30.85 0.00 -20.00 0.00 62.25 GLUDy, MDH 296 ADHEr, FBA, 6.16 0.29 29.34 0.00 -6.20 30.85 0.00 -20.00 0.00 62.25 FUM, GLUDy 297 ADHEr, 6.16 0.29 29.34 0.00 -6.20 30.85 0.00 -20.00 0.00 62.25 FUM, GLUDy, PFK 298 ADHEr, 6.16 0.29 29.34 0.00 -6.20 30.85 0.00 -20.00 0.00 62.25 GLUDy, MDH, TPI 299 ADHEr, 6.16 0.29 29.34 0.00 -6.20 30.85 0.00 -20.00 0.00 62.25 FUM, GLUDy, TPI 300 ADHEr, RPE, 6.16 0.33 28.65 0.00 -5.97 30.37 0.00 -20.00 0.00 61.36 SUCD4, TPI 301 ADHEr, PFK, 6.16 0.33 28.65 0.00 -5.97 30.37 0.00 -20.00 0.00 61.36 RPE, SUCD4 302 ADHEr, FBA, 6.16 0.33 28.65 0.00 -5.97 30.37 0.00 -20.00 0.00 61.36 RPE, SUCD4 303 ADHEr, 6.15 0.30 29.25 0.00 -6.19 30.80 0.00 -20.00 0.00 62.15 GLUDy, PFK, SUCD4 304 ADHEr, FBA, 6.15 0.30 29.25 0.00 -6.19 30.80 0.00 -20.00 0.00 62.15 GLUDy, SUCD4 305 ADHEr, 6.15 0.30 29.25 0.00 -6.19 30.80 0.00 -20.00 0.00 62.15 GLUDy, SUCD4, TPI 306 ADHEr, 6.14 0.32 28.86 0.00 -6.07 30.53 0.00 -20.00 0.00 61.66 HEX1, PFK, TAL 307 ADHEr, FBA, 6.14 0.32 28.86 0.00 -6.07 30.53 0.00 -20.00 0.00 61.66 HEX1, TAL 308 ADHEr, 6.14 0.32 28.86 0.00 -6.07 30.53 0.00 -20.00 0.00 61.66 HEX1, TAL, TPI 309 ADHEr, PFK, 6.13 0.33 28.70 0.00 -6.05 30.43 0.00 -20.00 0.00 61.47 SUCD4, TAL 310 ADHEr, FBA, 6.13 0.33 28.70 0.00 -6.05 30.43 0.00 -20.00 0.00 61.47 SUCD4, TAL 311 ADHEr, 6.13 0.33 28.70 0.00 -6.05 30.43 0.00 -20.00 0.00 61.47 SUCD4, TAL, TPI 312 ADHEr, 6.10 0.62 14.90 0.00 -3.07 18.16 0.00 -20.00 0.00 49.89 FUM, ME2, THD2 313 ADHEr, 6.04 0.65 18.68 0.00 0.48 22.08 0.00 -20.00 0.00 48.37 ATPS4r, ME2, SUCD4 314 ADHEr, 6.00 0.38 28.04 0.00 -6.05 30.04 0.00 -20.00 0.00 60.81 PPCK, PYK, SUCD4 315 ADHEr, 5.97 0.40 27.84 0.00 -6.03 29.92 0.00 -20.00 0.00 60.59 GLCpts, PPCK, SUCD4 316 ADHEr, 5.95 0.46 26.74 0.00 -5.68 29.16 0.00 -20.00 0.00 59.19 PPCK, RPE, SUCD4 317 ADHEr, 5.94 0.42 27.58 0.00 -6.00 29.76 0.00 -20.00 0.00 60.30 FUM, GLUDy, PPCK 318 ADHEr, 5.94 0.42 27.58 0.00 -6.00 29.76 0.00 -20.00 0.00 60.30 GLUDy, MDH, PPCK 319 ADHEr, 5.94 0.42 27.54 0.00 -5.99 29.74 0.00 -20.00 0.00 60.27 GLUDy, PPCK, SUCD4 320 ADHEr, 5.90 0.46 26.81 0.00 -5.79 29.24 0.00 -20.00 0.00 59.36 PPCK, SUCD4, TAL 321 ADHEr, 5.89 0.45 27.19 0.00 -5.95 29.52 0.00 -20.00 0.00 59.87 GLCpts, GLUDy, MDH 322 ADHEr, 5.89 0.51 26.14 0.00 -5.59 28.78 0.00 -20.00 0.00 58.51 GLCpts,
PPCK, RPE 323 ADHEr, 5.88 0.51 26.12 0.00 -5.59 28.77 0.00 -20.00 0.00 58.49 GLUDy, MDH, RPE 324 ADHEr, 5.87 0.52 26.00 0.00 -5.57 28.69 0.00 -20.00 0.00 58.35 GLUDy, PPCK, RPE 325 ADHEr, 5.87 0.46 27.02 0.00 -5.93 29.42 0.00 -20.00 0.00 59.69 GLCpts, GLUDy, PPCK 326 ADHEr, 5.86 0.46 26.97 0.00 -5.93 29.38 0.00 -20.00 0.00 59.63 GLCpts, MDH, SUCD4 327 ADHEr, 5.85 0.53 25.81 0.00 -5.54 28.57 0.00 -20.00 0.00 58.14 MDH, RPE, SUCD4 328 ADHEr, 5.85 0.47 26.83 0.00 -5.91 29.29 0.00 -20.00 0.00 59.47 GLCpts, MDH, PPCK 329 ADHEr, 5.85 0.47 26.83 0.00 -5.91 29.29 0.00 -20.00 0.00 59.47 FUM, GLCpts, PPCK 330 ADHEr, 5.84 0.54 25.70 0.00 -5.53 28.50 0.00 -20.00 0.00 58.02 MDH, PPCK, RPE 331 ADHEr, 5.84 0.54 25.70 0.00 -5.53 28.50 0.00 -20.00 0.00 58.02 FUM, PPCK, RPE 332 ADHEr, 5.84 0.51 26.23 0.00 -5.72 28.88 0.00 -20.00 0.00 58.71 GLCpts, PPCK, TAL 333 ADHEr, 5.83 0.51 26.21 0.00 -5.71 28.86 0.00 -20.00 0.00 58.68 GLUDy, MDH, TAL 334 ADHEr, 5.82 0.47 26.84 0.00 -5.98 29.28 0.00 -20.00 0.00 59.61 MDH, PPCK, PYK 335 ADHEr, 5.82 0.47 26.84 0.00 -5.98 29.28 0.00 -20.00 0.00 59.61 FUM, PPCK, PYK 336 ADHEr, 5.82 0.52 26.09 0.00 -5.70 28.79 0.00 -20.00 0.00 58.55 GLUDy, PPCK, TAL 337 ADHEr, 5.81 0.49 26.51 0.00 -5.87 29.10 0.00 -20.00 0.00 59.12 FUM, GLUDy, SUCD4 338 ADHEr, 5.80 0.53 25.90 0.00 -5.67 28.67 0.00 -20.00 0.00 58.33 MDH, SUCD4, TAL 339 ADHEr, 5.80 0.50 26.45 0.00 -5.86 29.06 0.00 -20.00 0.00 59.05 GLCpts, ME2, SUCD4 340 ADHEr, ME2, 5.79 0.57 25.28 0.00 -5.46 28.24 0.00 -20.00 0.00 57.53 RPE, SUCD4 341 ADHEr, 5.79 0.54 25.79 0.00 -5.66 28.61 0.00 -20.00 0.00 58.22 FUM, PPCK, TAL 342 ADHEr, 5.79 0.54 25.79 0.00 -5.66 28.61 0.00 -20.00 0.00 58.22 MDH, PPCK, TAL 343 ADHEr, 5.78 0.51 26.35 0.00 -5.85 29.00 0.00 -20.00 0.00 58.95 GLUDy, PRO1z, SUCD4 344 ADHEr, 5.78 0.57 25.18 0.00 -5.45 28.18 0.00 -20.00 0.00 57.43 FUM, GLUDy, RPE 345 ADHEr, 5.78 0.57 25.17 0.00 -5.45 28.17 0.00 -20.00 0.00 57.41 GLUDy, RPE, SUCD4 346 ADHEr, 5.78 0.57 25.15 0.00 -5.44 28.15 0.00 -20.00 0.00 57.39 GLCpts, RPE, SUCD4 347 ADHEr, 5.78 0.51 26.30 0.00 -5.85 28.97 0.00 -20.00 0.00 58.89 FUM, GLUDy, ME2 348 ADHEr, 5.78 0.51 26.30 0.00 -5.85 28.96 0.00 -20.00 0.00 58.88 GLUDy, ME2, SUCD4 349 ADHEr, 5.77 0.52 26.22 0.00 -5.84 28.91 0.00 -20.00 0.00 58.80 FUM, GLCpts, GLUDy 350 ADHEr, 5.76 0.52 26.17 0.00 -5.83 28.88 0.00 -20.00 0.00 58.74 GLCpts, GLUDy, SUCD4 351 ADHEr, 5.74 0.53 25.99 0.00 -5.81 28.78 0.00 -20.00 0.00 58.55 FUM, ME2, SUCD4 352 ADHEr, ME2, 5.74 0.57 25.38 0.00 -5.60 28.34 0.00 -20.00 0.00 57.75 SUCD4, TAL 353 ADHEr, 5.72 0.58 25.26 0.00 -5.59 28.27 0.00 -20.00 0.00 57.62 FUM, GLUDy, TAL 354 ADHEr, 5.72 0.58 25.25 0.00 -5.58 28.26 0.00 -20.00 0.00 57.61 GLUDy, SUCD4, TAL 355 ADHEr, 5.72 0.58 25.21 0.00 -5.58 28.24 0.00 -20.00 0.00 57.57 GLCpts, SUCD4, TAL 356 ADHEr, ME2, 5.71 0.95 0.00 0.00 6.29 15.52 0.00 -20.00 0.00 40.16 PGL, THD2 357 ADHEr, 5.71 0.95 0.00 0.00 6.29 15.52 0.00 -20.00 0.00 40.16 G6PDHy, ME2, THD2 358 ADHEr, 5.71 0.62 24.48 0.00 -5.35 27.73 0.00 -20.00 0.00 56.63 FUM, RPE, SUCD4 359 ADHEr, 5.69 0.63 24.34 0.00 -5.32 27.64 0.00 -20.00 0.00 56.47 HEX1, RPE, SUCD4 360 ADHEr, 5.68 0.57 25.51 0.00 -5.76 28.48 0.00 -20.00 0.00 58.02 CBMK2, GLU5K, PPCK 361 ADHEr, 5.68 0.57 25.51 0.00 -5.76 28.48 0.00 -20.00 0.00 58.02 CBMK2, G5SD, PPCK 362 ADHEr, 5.68 0.57 25.51 0.00 -5.76 28.48 0.00 -20.00 0.00 58.02 ASNS2, CBMK2, PPCK 363 ADHEr, 5.68 0.57 25.50 0.00 -5.75 28.47 0.00 -20.00 0.00 58.00 CBMK2, PPCK, SO4t2 364 ADHEr, 5.67 0.57 25.42 0.00 -5.75 28.42 0.00 -20.00 0.00 57.92 GLUDy, HEX1, SUCD4 365 ADHEr, 5.64 0.63 24.56 0.00 -5.49 27.83 0.00 -20.00 0.00 56.84 FUM, SUCD4, TAL 366 ADHEr, 5.63 0.63 24.44 0.00 -5.48 27.75 0.00 -20.00 0.00 56.70 HEX1, SUCD4, TAL 367 ADHEr, 5.62 0.60 25.02 0.00 -5.70 28.17 0.00 -20.00 0.00 57.47 FUM, HEX1, SUCD4 368 ADHEr, 5.58 0.70 23.35 0.00 -5.18 27.02 0.00 -20.00 0.00 55.35 FUM, HEX1, RPE 369 ADHEr, 5.58 0.62 24.73 0.00 -5.67 27.99 0.00 -20.00 0.00 57.15 CBMK2, FUM, SUCD4 370 ADHEr, 5.54 0.72 14.12 0.00 3.77 0.00 0.00 -20.00 0.00 37.22 HEX1, PFLi, RPE 371 ADHEr, 5.51 0.70 23.47 0.00 -5.35 27.15 0.00 -20.00 0.00 55.62 FUM, HEX1, TAL 372 ADHEr, 5.45 0.70 23.69 0.00 -5.54 27.34 0.00 -20.00 0.00 56.00 CBMK2, FUM, HEX1 373 ADHEr, 5.40 0.72 14.11 0.00 3.45 0.00 0.00 -20.00 0.00 37.64 HEX1, PFLi, TAL 374 ADHEr, 5.25 0.72 14.10 0.00 3.09 0.00 0.00 -20.00 0.09 38.10 GLYCL, HEX1, PFLi 375 ADHEr, 5.14 0.46 27.54 0.00 -4.79 29.95 0.00 -20.00 0.00 60.78 ATPS4r, GLUDy, PGI 376 ADHEr, PFLi, 5.08 0.43 16.39 0.00 1.66 0.00 0.00 -20.00 0.00 43.14 PGDH, PGI 377 ADHEr, PFLi, 5.05 0.43 16.21 0.00 1.57 0.00 0.00 -20.00 0.00 43.21 PGI, TAL 378 ADHEr, 5.03 0.52 26.76 0.00 -4.73 29.48 0.00 -20.00 0.00 59.94 ATPS4r, PFK, RPE 379 ADHEr, 5.03 0.52 26.76 0.00 -4.73 29.48 0.00 -20.00 0.00 59.94 ATPS4r, FBA, RPE 380 ADHEr, 5.03 0.52 26.76 0.00 -4.73 29.48 0.00 -20.00 0.00 59.94 ATPS4r, RPE, TPI 381 ADHEr, PFLi, 5.02 0.43 16.04 0.00 1.47 0.00 0.00 -20.00 0.00 43.27 PGI, RPE 382 ADHEr, 5.02 0.47 27.70 0.00 -5.08 30.15 0.00 -20.00 0.00 61.18 ATPS4r, GLUDy, TPI 383 ADHEr, 5.02 0.47 27.70 0.00 -5.08 30.15 0.00 -20.00 0.00 61.18 ATPS4r, FBA, GLUDy 384 ADHEr, 5.02 0.47 27.70 0.00 -5.08 30.15 0.00 -20.00 0.00 61.18 ATPS4r, GLUDy, PFK 385 ADHEr, FBA, 5.01 0.44 15.95 0.00 1.43 0.00 0.00 -20.00 0.00 43.31 PFLi, PGI 386 ADHEr, PFK, 5.01 0.44 15.95 0.00 1.43 0.00 0.00 -20.00 0.00 43.31 PFLi, PGI 387 ADHEr, PFLi, 5.01 0.44 15.95 0.00 1.43 0.00 0.00 -20.00 0.00 43.31 PGI, TPI 388 ADHEr, PFLi, 4.99 0.44 15.97 0.00 1.38 0.00 0.00 -20.00 0.00 43.37 RPE, TPI 389 ADHEr, PFK, 4.99 0.44 15.97 0.00 1.38 0.00 0.00 -20.00 0.00 43.37 PFLi, RPE 390 ADHEr, FBA, 4.99 0.44 15.97 0.00 1.38 0.00 0.00 -20.00 0.00 43.37 PFLi, RPE 391 ADHEr, 4.98 0.52 26.82 0.00 -4.85 29.56 0.00 -20.00 0.00 60.10 ATPS4r, PFK, TAL 392 ADHEr, 4.98 0.52 26.82 0.00 -4.85 29.56 0.00 -20.00 0.00 60.10 ATPS4r, FBA, TAL 393 ADHEr, 4.98 0.52 26.82 0.00 -4.85 29.56 0.00 -20.00 0.00 60.10 ATPS4r, TAL, TPI 394 ADHEr, FBA, 4.94 0.44 16.00 0.00 1.28 0.00 0.00 -20.00 0.00 43.49 PFLi, TAL 395 ADHEr, PFLi, 4.94 0.44 16.00 0.00 1.28 0.00 0.00 -20.00 0.00 43.49 TAL, TPI 396 ADHEr, PFK, 4.94 0.44 16.00 0.00 1.28 0.00 0.00 -20.00 0.00 43.49 PFLi, TAL 397 ADHEr, 4.90 0.74 22.16 0.00 -6.47 26.02 0.00 -20.00 0.09 56.37 GLYCL, HEX1, THD2 398 ADHEr, 4.89 0.39 16.45 0.00 1.06 0.00 0.00 -20.00 0.00 44.29 GLUDy, PFK, PFLi 399 ADHEr, 4.89 0.39 16.45 0.00 1.06 0.00 0.00 -20.00 0.00 44.29 GLUDy, PFLi, TPI 400 ADHEr, FBA, 4.89 0.39 16.45 0.00 1.06 0.00 0.00 -20.00 0.00 44.29 GLUDy, PFLi 401 ADHEr, EDA, 4.83 0.50 0.00 0.00 -3.17 0.00 0.00 -20.00 0.00 45.11 PFLi, PGI 402 ADHEr, 4.72 0.72 23.93 0.00 -4.31 27.69 0.00 -20.00 0.00 56.73 ATPS4r, GLUDy, RPE 403 ADHEr, 4.65 0.72 24.01 0.00 -4.47 27.80 0.00 -20.00 0.00 56.96 ATPS4r, GLUDy, TAL 404 ADHEr, 4.60 0.80 22.80 0.00 -4.14 26.97 0.00 -20.00 0.00 55.45 ATPS4r, CBMK2, RPE 405 ADHEr, 4.57 0.72 24.19 0.00 -4.67 27.97 0.00 -20.00 0.00 57.30 ATPS4r, CBMK2, GLUDy 406 ADHEr, 4.51 0.80 22.88 0.00 -4.32 27.09 0.00 -20.00 0.00 55.69 ATPS4r, CBMK2, TAL 407 ADHEr, 4.42 0.81 22.95 0.00 -4.53 27.20 0.00 -20.00 0.00 55.93 ASNS2,
ATPS4r, GLU5K 408 ADHEr, 4.42 0.81 22.95 0.00 -4.53 27.20 0.00 -20.00 0.00 55.93 ASNS2, ATPS4r, G5SD 409 ADHEr, 3.00 0.50 3.27 0.00 -6.43 0.00 0.00 -20.00 0.00 50.40 ACKr, PFLi, PGI 410 ADHEr, 2.76 1.03 3.32 0.00 14.07 0.00 0.00 -20.00 0.00 25.81 ACKr, AKGD, FRD2 411 ADHEr, 1.91 1.03 2.47 0.00 12.55 0.00 0.00 -20.00 0.00 27.45 ACKr, FRD2, SUCOAS 412 ADHEr, 1.40 0.71 14.98 0.00 -12.86 18.72 0.00 -20.00 0.00 62.31 FUM, G6PDHy, TAL 413 ADHEr, 1.40 0.71 14.98 0.00 -12.86 18.72 0.00 -20.00 0.00 62.31 FUM, PGDH, TAL 414 ADHEr, 1.40 0.71 14.98 0.00 -12.86 18.72 0.00 -20.00 0.00 62.31 FUM, PGL, TAL 415 ADHEr, 1.36 0.47 17.83 0.00 -14.86 20.31 0.00 -20.00 0.00 68.38 FUM, ME2, TPI 416 ADHEr, 1.36 0.47 17.83 0.00 -14.86 20.31 0.00 -20.00 0.00 68.38 FUM, ME2, PFK 417 ADHEr, FBA, 1.36 0.47 17.83 0.00 -14.86 20.31 0.00 -20.00 0.00 68.38 FUM, ME2 418 ADHEr, 1.22 0.24 0.00 0.00 -33.89 3.68 33.86 -20.00 0.00 73.09 FRD2, GLUDy, LDH_D 419 ADHEr, 1.15 0.40 0.00 0.00 15.41 4.39 0.00 -20.00 0.00 22.97 FRD2, LDH_D, THD2 420 ADHEr, 1.01 0.73 14.02 0.00 -13.53 17.85 0.00 -20.00 0.00 62.73 G6PDHy, HEX1, TAL 421 ADHEr, 1.01 0.73 14.02 0.00 -13.53 17.85 0.00 -20.00 0.00 62.73 HEX1, PGDH, TAL 422 ADHEr, 1.01 0.73 14.02 0.00 -13.53 17.85 0.00 -20.00 0.00 62.73 HEX1, PGL, TAL 423 ADHEr, 0.89 0.65 14.71 0.00 -14.28 18.10 0.00 -20.00 0.00 64.72 MDH, PGDH, TAL 424 ADHEr, 0.89 0.65 14.71 0.00 -14.28 18.10 0.00 -20.00 0.00 64.72 G6PDHy, MDH, TAL 425 ADHEr, 0.89 0.65 14.71 0.00 -14.28 18.10 0.00 -20.00 0.00 64.72 MDH, PGL, TAL 426 ADHEr, 0.43 0.47 15.73 0.00 -16.28 18.18 0.00 -20.00 0.00 69.35 PGDH, TAL, TPI 427 ADHEr, 0.43 0.47 15.73 0.00 -16.28 18.18 0.00 -20.00 0.00 69.35 FBA, PGDH, TAL 428 ADHEr, 0.43 0.47 15.73 0.00 -16.28 18.18 0.00 -20.00 0.00 69.35 PFK, PGDH, TAL 429 ADHEr, 0.41 0.49 15.57 0.00 -16.70 18.12 0.00 -20.00 0.06 69.95 GLYCL, TAL, TPI 430 ADHEr, 0.40 0.49 15.57 0.00 -16.69 18.11 0.00 -20.00 0.00 69.95 TAL, THD5, TPI 431 ADHEr, 0.40 0.49 15.57 0.00 -16.69 18.11 0.00 -20.00 0.00 69.95 LDH_D, TAL, TPI 432 ADHEr, 16.17 0.05 8.39 0.00 11.88 24.76 0.00 -20.00 0.00 33.47 ASPT, EDA, MDH, PGI 433 ADHEr, 15.11 0.23 0.00 0.00 23.25 9.12 0.00 -20.00 0.00 13.86 ATPS4r, FRD2, LDH_D, ME2 434 ADHEr, 14.76 0.16 16.11 0.00 22.13 0.00 0.00 -20.00 0.00 17.24 EDA, PFLi, PGI, PPCK 435 ADHEr, 15.02 0.23 0.00 0.00 23.19 9.11 0.00 -20.00 0.00 13.93 ATPS4r, FRD2, LDH_D, MDH 436 ADHEr, 14.65 0.19 15.67 0.00 21.97 0.00 0.00 -20.00 0.00 17.05 EDA, PFLi, PGI, SUCD4 437 ADHEr, 14.65 0.19 15.67 0.00 21.97 0.00 0.00 -20.00 0.00 17.05 EDA, NADH6, PFLi, PGI 438 ADHEr, 14.47 0.09 17.44 0.00 19.97 3.46 0.00 -20.00 0.00 21.55 EDA, NADH6, PGI, PPCK 439 ADHEr, 14.38 0.36 13.01 0.00 21.55 0.00 0.00 -20.00 0.00 15.56 ASPT, LDH_D, MDH, PFLi 440 ADHEr, 14.31 0.31 14.25 0.00 21.45 0.00 0.00 -20.00 0.00 16.44 GLUDy, HEX1, PFLi, PGI 441 ADHEr, 14.23 0.27 14.38 0.00 22.22 0.00 0.00 -20.00 0.00 16.72 ACKr, GLCpts, NADH6, PGI 442 ADHEr, 14.13 0.25 15.36 0.00 19.91 2.53 0.00 -20.00 0.00 19.65 GLUDy, HEX1, NADH6, PGI 443 ADHEr, 14.07 0.27 15.02 0.00 19.91 2.38 0.00 -20.00 0.00 19.34 EDA, GLUDy, NADH6, PGI 444 ADHEr, 14.04 0.19 14.14 0.00 21.04 0.00 0.00 -20.00 0.00 17.69 ACKr, PFLi, PGI, SUCD4 445 ADHEr, 14.04 0.19 14.14 0.00 22.14 0.00 0.00 -20.00 0.00 16.60 ACKr, NADH6, PFLi, PGI 446 ADHEr, 14.03 0.30 14.19 0.00 22.02 0.00 0.00 -20.00 0.00 16.65 ACKr, GLUDy, NADH6, PGI 447 ADHEr, 14.02 0.28 14.76 0.00 21.90 0.00 0.00 -20.00 0.00 17.17 EDA, GLCpts, NADH6, PGI 448 ADHEr, 13.94 0.34 14.12 0.00 21.89 0.00 0.00 -20.00 0.00 16.54 ACKr, CBMK2, NADH6, PGI 449 ADHEr, 13.86 0.22 14.56 0.00 21.78 0.00 0.00 -20.00 0.00 17.62 ATPS4r, FDH2, NADH6, PGI 450 ADHEr, 13.86 0.22 14.56 0.00 21.78 0.00 0.00 -20.00 0.00 17.62 ATPS4r, NADH6, PFLi, PGI 451 ADHEr, 13.80 0.27 16.36 0.00 20.68 0.00 0.00 -20.00 0.00 18.31 ATPS4r, GLCpts, NADH6, PFLi 452 ADHEr, 13.69 0.43 13.35 0.00 20.51 0.00 0.00 -20.00 0.00 16.43 ATPS4r, MDH, NADH6, PGL 453 ADHEr, 13.69 0.43 13.35 0.00 20.51 0.00 0.00 -20.00 0.00 16.43 ATPS4r, G6PDHy, MDH, NADH6 454 ADHEr, 13.68 0.36 13.87 0.00 18.43 4.12 0.00 -20.00 0.00 20.56 ACKr, FUM, GLUDy, LDH_D 455 ADHEr, 13.66 0.25 15.79 0.00 19.61 3.75 0.00 -20.00 0.00 21.32 ATPS4r, NADH6, PGI, SUCD4 456 ADHEr, 13.56 0.37 13.76 0.00 17.71 5.22 0.00 -20.00 0.00 21.60 ACKr, GLUDy, LDH_D, SUCD4 457 ADHEr, 13.53 0.44 10.63 0.00 14.18 12.16 0.00 -20.00 0.00 25.92 ATPS4r, G6PDHy, MDH, THD2 458 ADHEr, 13.53 0.44 10.63 0.00 14.18 12.16 0.00 -20.00 0.00 25.92 ATPS4r, MDH, PGL, THD2 459 ADHEr, 13.44 0.26 11.28 0.00 7.24 25.83 0.00 -20.00 0.00 38.92 ASPT, G6PDHy, MDH, PYK 460 ADHEr, 13.44 0.26 11.28 0.00 7.24 25.83 0.00 -20.00 0.00 38.92 ASPT, EDA, MDH, PYK 461 ADHEr, 13.44 0.26 11.28 0.00 7.24 25.83 0.00 -20.00 0.00 38.92 ASPT, MDH, PGL, PYK 462 ADHEr, 13.35 0.25 4.81 0.00 13.43 19.59 0.00 -20.00 0.00 28.60 FRD2, LDH_D, MDH, SUCOAS 463 ADHEr, 13.27 0.26 4.78 0.00 13.14 19.51 0.00 -20.00 0.00 28.92 ASPT, LDH_D, MDH, SUCOAS 464 ADHEr, 13.22 0.26 0.00 0.00 17.54 14.32 0.00 -20.00 0.00 21.06 ACt6, LDH_D, MDH, SUCD4 465 ADHEr, 13.17 0.24 16.40 0.00 17.06 7.36 0.00 -20.00 0.00 25.48 ATPS4r, GLUDy, PGI, SUCD4 466 ADHEr, 13.15 0.27 5.76 0.00 12.63 20.09 0.00 -20.00 0.00 29.76 ASPT, FUM, LDH_D, MDH 467 ADHEr, 13.15 0.27 5.76 0.00 12.63 20.09 0.00 -20.00 0.00 29.76 ASPT, LDH_D, MALS, MDH 468 ADHEr, 13.15 0.27 5.76 0.00 12.63 20.09 0.00 -20.00 0.00 29.76 ASPT, ICL, LDH_D, MDH 469 ADHEr, 13.07 0.51 0.00 0.00 23.45 0.00 0.00 -20.00 0.00 13.57 ACt6, LDH_D, MDH, NADH6 470 ADHEr, 12.91 0.12 12.29 0.00 -0.02 38.75 0.00 -20.00 0.00 51.90 FRD2, GLUDy, LDH_D, PPCK 471 ADHEr, 12.89 0.13 12.25 0.00 -0.02 38.70 0.00 -20.00 0.00 51.85 FRD2, LDH_D, PPCK, THD2
472 ADHEr, 12.73 0.47 12.99 0.00 16.72 6.70 0.00 -20.00 0.00 23.06 ACKr, ATPS4r, LDH_D, SUCD4 473 ADHEr, 12.62 0.11 12.68 0.00 0.33 39.20 0.00 -20.00 0.00 52.67 ACKr, ACS, PPC, PPCK 474 ADHEr, 12.60 0.16 11.81 0.00 0.47 38.84 0.00 -20.00 0.00 51.79 GLUDy, LDH_D, PPC, PPCK 475 ADHEr, 12.60 0.48 14.92 0.00 19.87 0.00 0.00 -20.00 0.00 18.32 ATPS4r, FDH2, NADH6, SULabc 476 ADHEr, 12.60 0.16 11.72 0.00 0.49 38.81 0.00 -20.00 0.00 51.71 LDH_D, PPC, PPCK, THD2 477 ADHEr, 12.57 0.29 9.83 0.00 18.83 0.00 0.00 -20.00 0.00 18.28 ASPT, ATPS4r, GLCpts, MDH 478 ADHEr, 12.37 0.68 7.32 0.00 24.57 2.87 0.00 -20.00 0.00 15.06 G6PDHy, MDH, NADH6, THD2 479 ADHEr, 12.37 0.68 7.32 0.00 24.57 2.87 0.00 -20.00 0.00 15.06 MDH, NADH6, PGL, THD2 480 ADHEr, 12.36 0.41 12.58 0.00 28.51 0.00 0.00 -20.00 0.00 15.47 ACKr, FBA, GLUDy, NADH6 481 ADHEr, 12.36 0.41 12.58 0.00 28.51 0.00 0.00 -20.00 0.00 15.47 ACKr, GLUDy, NADH6, PFK 482 ADHEr, 12.36 0.41 12.58 0.00 28.51 0.00 0.00 -20.00 0.00 15.47 ACKr, GLUDy, NADH6, TPI 483 ADHEr, 12.33 0.49 15.46 0.00 19.46 0.00 0.00 -20.00 0.00 18.94 ATPS4r, MTHFC, NADH6, PFLi 484 ADHEr, 12.33 0.49 15.46 0.00 19.46 0.00 0.00 -20.00 0.00 18.94 ATPS4r, FTHFD, NADH6, PFLi 485 ADHEr, 12.30 0.34 0.00 0.00 12.96 0.00 0.00 -20.00 0.00 24.29 ATPS4r, G6PDHy, GLCpts, MDH 486 ADHEr, 12.30 0.34 0.00 0.00 12.96 0.00 0.00 -20.00 0.00 24.29 ATPS4r, GLCpts, MDH, PGL 487 ADHEr, 12.15 0.44 12.39 0.00 28.20 0.00 0.00 -20.00 0.00 15.54 ACKr, FBA, GLCpts, NADH6 488 ADHEr, 12.15 0.44 12.39 0.00 28.20 0.00 0.00 -20.00 0.00 15.54 ACKr, GLCpts, NADH6, TPI 489 ADHEr, 12.15 0.44 12.39 0.00 28.20 0.00 0.00 -20.00 0.00 15.54 ACKr, GLCpts, NADH6, PFK 490 ADHEr, 12.15 0.40 12.37 0.00 6.33 23.73 0.00 -20.00 0.00 38.94 ACKr, LDH_D, MDH, SUCD4 491 ADHEr, 12.13 0.22 12.25 0.00 37.66 0.00 0.00 -20.00 0.00 13.85 ACKr, AKGD, ATPS4r, FBA 492 ADHEr, 12.13 0.22 12.25 0.00 37.66 0.00 0.00 -20.00 0.00 13.85 ACKr, AKGD, ATPS4r, PFK 493 ADHEr, 12.13 0.22 12.25 0.00 12.10 25.56 0.00 -20.00 0.00 39.40 ACKr, AKGD, ATPS4r, TPI 494 ADHEr, 12.09 0.07 20.13 0.00 7.88 20.50 0.00 -20.00 0.00 41.14 EDA, PGI, PPCK, SUCD4 495 ADHEr, 12.09 0.23 12.21 0.00 12.27 25.58 0.00 -20.00 0.00 39.40 ACKr, ATPS4r, FBA, SUCOAS 496 ADHEr, 12.09 0.23 12.21 0.00 12.27 25.58 0.00 -20.00 0.00 39.40 ACKr, ATPS4r, PFK, SUCOAS 497 ADHEr, 12.09 0.23 12.21 0.00 37.86 0.00 0.00 -20.00 0.00 13.82 ACKr, ATPS4r, SUCOAS, TPI 498 ADHEr, 12.08 0.26 3.82 0.00 8.14 17.37 4.03 -20.00 0.00 33.19 FRD2, LDH_D, ME2, SUCOAS 499 ADHEr, 12.06 0.46 12.31 0.00 28.06 0.00 0.00 -20.00 0.00 15.57 ACKr, CBMK2, FBA, NADH6 500 ADHEr, 12.06 0.46 12.31 0.00 28.06 0.00 0.00 -20.00 0.00 15.57 ACKr, CBMK2, NADH6, PFK 501 ADHEr, 12.06 0.46 12.31 0.00 28.06 0.00 0.00 -20.00 0.00 15.57 ACKr, CBMK2, NADH6, TPI 502 ADHEr, 12.05 0.46 12.30 0.00 28.05 0.00 0.00 -20.00 0.00 15.58 ACKr, NADH6, RPE, TPI 503 ADHEr, 12.05 0.46 12.30 0.00 28.05 0.00 0.00 -20.00 0.00 15.58 ACKr, NADH6, PFK, RPE 504 ADHEr, 12.05 0.46 12.30 0.00 28.05 0.00 0.00 -20.00 0.00 15.58 ACKr, FBA, NADH6, RPE 505 ADHEr, 12.05 0.46 12.30 0.00 28.04 0.00 0.00 -20.00 0.00 15.58 ACKr, ASNS2, FBA, NADH6 H2O ILE LAC NH4 NO3 PHE PI SO4 SUC THR VAL 1 8.96 0.00 0.00 -6.26 -2.00 0.00 -0.78 -0.13 0.23 0.00 0.00 2 -12.07 0.00 0.00 -0.24 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 3 -12.07 0.00 0.00 -0.24 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 4 -0.90 0.00 0.00 -4.96 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 5 0.68 0.00 0.00 -5.60 0.00 0.00 -0.69 -0.11 0.00 0.00 0.00 6 6.08 0.00 0.00 -7.09 -5.00 0.00 -0.88 -0.14 0.00 0.00 0.00 7 21.51 0.00 0.00 -4.52 0.00 0.00 -0.56 -0.09 24.86 0.00 0.00 8 10.90 0.00 0.00 -6.44 0.00 0.00 -0.80 -0.13 12.16 0.00 0.00 9 12.29 0.00 0.00 -6.62 0.00 0.00 -0.82 -0.13 13.54 0.00 0.00 10 10.81 0.00 0.00 -5.83 0.00 0.00 -0.72 -0.12 14.32 0.00 0.00 11 8.25 0.00 0.00 -4.23 0.00 0.00 -0.52 -0.09 16.46 0.00 0.00 12 8.25 0.00 0.00 -4.23 0.00 0.00 -0.52 -0.09 16.46 0.00 0.00 13 8.25 0.00 0.00 -4.23 0.00 0.00 -0.52 -0.09 16.46 0.00 0.00 14 4.02 0.00 0.00 -3.39 0.00 0.00 -0.42 -0.07 0.00 0.00 0.00 15 14.07 0.00 0.00 -4.31 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 16 14.87 0.00 0.00 -4.72 0.00 0.00 -0.58 -0.10 0.00 0.00 0.00 17 30.54 0.00 0.00 -7.41 -20.00 0.00 -0.92 -0.15 0.00 0.00 0.00 18 15.46 0.00 0.00 -5.03 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 19 3.71 0.00 0.00 -0.73 -10.00 0.00 -0.09 -0.01 0.00 0.00 0.00 20 3.71 0.00 0.00 -0.73 -10.00 0.00 -0.09 -0.01 0.00 0.00 0.00 21 -1.88 0.00 0.00 -4.57 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 22 -0.60 0.00 0.00 -3.07 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 23 -0.71 0.00 0.00 -3.11 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 24 -0.71 0.00 0.00 -3.11 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 25 -0.71 0.00 0.00 -3.11 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 26 2.51 0.00 0.00 -4.48 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 27 4.00 0.00 0.00 -5.12 0.00 0.00 -0.63 -0.10 0.00 0.00 0.00 28 5.55 0.00 0.00 -5.78 0.00 0.00 -0.72 -0.12 0.00 0.00 0.00 29 0.13 0.00 0.00 -4.83 0.00 0.00 -0.60 -0.10 0.00 0.00 0.00 30 13.24 0.00 0.00 -6.66 -5.00 0.00 -0.82 -0.13 0.23 0.00 0.00 31 12.86 0.00 0.00 -6.69 -5.00 0.00 -0.83 -0.14 0.00 0.00 0.00 32 -6.65 0.00 0.00 -2.62 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 33 -6.10 0.00 0.00 -2.84 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 34 1.88 0.00 0.00 -5.25 -2.00 0.00 -0.65 -0.11 0.00 0.00 0.00 35 -6.68 0.00 0.00 -2.66 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 36 -6.68 0.00 0.00 -2.66 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 37 -6.68 0.00 0.00 -2.66 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 38 -6.39 0.00 0.00 -2.77 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 39 -6.39 0.00 0.00 -2.77 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 40 -6.39 0.00 0.00 -2.77 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 41 8.45 0.00 0.00 -5.39 0.00 0.00 -0.67 -0.11 6.20 0.00 0.00 42 -6.13 0.00 0.00 -2.88 0.00 0.00 -0.36 -0.06 0.00 0.00 0.00 43 -6.13 0.00 0.00 -2.88 0.00 0.00 -0.36 -0.06 0.00 0.00 0.00 44 -6.13 0.00 0.00 -2.88 0.00 0.00 -0.36 -0.06 0.00 0.00 0.00 45 19.66 0.00 0.00 -6.02 0.00 0.00 -0.75 -0.12 7.94 0.00 0.00 46 -3.22 0.00 0.00 -4.04 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 47 -0.76 0.00 0.00 -4.95 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 48 -2.34 0.00 0.00 -4.39 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 49 -2.29 0.00 0.00 -4.41 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 50 -2.13 0.00 0.00 -4.47 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 51 -1.81 0.00 0.00 -4.60 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 52 -0.83 0.00 0.00 -4.95 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 53 -1.66 0.00 0.00 -4.66 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 54 -1.66 0.00 0.00 -4.66 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 55 0.74 0.00 0.00 -5.54 0.00 0.00 -0.69 -0.11 0.00 0.00 0.00 56 -1.01 0.00 0.00 -4.92 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 57 -0.80 0.00 0.00 -5.00 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 58 -0.78 0.00 0.00 -5.01 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 59 -0.69 0.00 0.00 -5.05 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 60 0.71 0.00 0.00 -5.56 0.00 0.00 -0.69 -0.11 0.00 0.00 0.00 61 0.30 0.00 0.00 -5.44 0.00 0.00 -0.67 -0.11 0.00 0.00 0.00 62 0.47 0.00 0.00 -5.51 0.00 0.00 -0.68 -0.11 0.00 0.00 0.00 63 0.56 0.00 0.00 -5.55 0.00 0.00 -0.69 -0.11 0.00 0.00 0.00 64 1.94 0.00 0.00 -6.10 0.00 0.00 -0.75 -0.12 0.00 0.00 0.00 65 20.54 0.00 0.00 -6.25 0.00 0.00 -0.77 -0.13 9.44 0.00 0.00 66 -0.22 0.00 0.00 -4.49 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 67 -0.25 0.00 0.00 -4.56 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 68 -0.25 0.00 0.00 -4.56 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 69 -0.25 0.00 0.00 -4.56 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 70 16.61 0.00 0.00 -3.82 0.00 0.00 -0.47 -0.08 12.23 0.00 0.00 71 16.61 0.00 0.00 -3.82 0.00 0.00 -0.47 -0.08 12.23 0.00 0.00 72 16.61 0.00 0.00 -3.82 0.00 0.00 -0.47 -0.08 12.23 0.00 0.00 73 3.70 0.00 0.00 -6.40 0.00 0.00 -0.79 -0.13 1.51 0.00 0.00 74 6.06 0.00 0.00 -6.97 -5.00 0.00 -0.86 -0.14 0.00 0.00 0.00 75 4.14 0.00 0.00 -6.31 -5.00 0.00 -0.78 -0.13 0.00 0.00 0.00 76 6.07 0.00 0.00 -7.03 -5.00 0.00 -0.87 -0.14 0.00 0.00 0.00 77 5.91 0.00 0.00 -7.02 -5.00 0.00 -0.87 -0.14 0.00 0.00 0.00 78 20.66 0.00 0.00 -4.37 0.00 0.00 -0.54 -0.09 21.30 0.00 0.00 79 22.80 0.00 0.00 -4.46 0.00 0.00 -0.55 -0.09 24.28 0.00 0.00 80 5.91 0.00 0.00 -4.11 0.00 0.00 -0.51 -0.08 13.43 0.00 0.00 81 5.91 0.00 0.00 -4.11 0.00 0.00 -0.51 -0.08 13.43 0.00 0.00 82 5.91 0.00 0.00 -4.11 0.00 0.00 -0.51 -0.08 13.43 0.00 0.00 83 12.13 0.00 0.00 -6.59 0.00 0.00 -0.82 -0.13 13.08 0.00 0.00 84 10.68 0.00 0.00 -5.80 0.00 0.00 -0.72 -0.12 13.90 0.00 0.00 85 12.21 0.00 0.00 -6.61 0.00 0.00 -0.82 -0.13 13.30 0.00 0.00 86 10.74 0.00 0.00 -5.81 0.00 0.00 -0.72 -0.12 14.10 0.00 0.00 87 8.23 0.00 0.00 -4.19 0.00 0.00 -0.52 -0.08 16.35 0.00 0.00 88 8.23 0.00 0.00 -4.19 0.00 0.00 -0.52 -0.08 16.35 0.00 0.00 89 8.23 0.00 0.00 -4.19 0.00 0.00 -0.52 -0.08 16.35 0.00 0.00 90 8.24 0.00 0.00 -4.21 0.00 0.00 -0.52 -0.09 16.41 0.00 0.00 91 8.24 0.00 0.00 -4.21 0.00 0.00 -0.52 -0.09 16.41 0.00 0.00 92 13.38 0.00 0.00 -2.97 0.00 0.00 -0.37 -0.06 0.00 0.00 0.00
93 10.91 0.00 0.00 -2.40 0.00 0.00 -0.30 -0.05 0.00 0.00 0.00 94 11.52 0.00 0.00 -2.66 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 95 14.23 0.00 0.00 -2.97 -2.00 0.00 -0.37 -0.06 0.00 0.00 0.00 96 6.78 2.14 0.00 -4.35 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 97 9.08 0.00 0.00 -2.36 -2.00 0.00 -0.29 -0.05 0.00 0.00 0.00 98 16.09 0.00 0.00 -4.23 -2.00 0.00 -0.52 -0.09 0.00 0.00 0.00 99 24.49 0.00 0.00 -4.01 -20.00 0.00 -0.50 -0.08 0.00 0.00 0.00 100 24.49 0.00 0.00 -4.01 -20.00 0.00 -0.50 -0.08 0.00 0.00 0.00 101 24.49 0.00 0.00 -4.01 -20.00 0.00 -0.50 -0.08 0.00 0.00 0.00 102 10.90 1.81 0.00 -4.09 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 103 0.36 0.00 0.00 -1.93 0.00 0.00 -0.24 -0.04 0.00 0.00 0.00 104 0.38 0.00 0.00 -1.94 0.00 0.00 -0.24 -0.04 0.00 0.00 0.00 105 1.01 0.00 0.00 -2.19 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 106 1.07 0.00 0.00 -2.21 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 107 2.80 0.00 0.00 -2.91 0.00 0.00 -0.36 -0.06 0.00 0.00 0.00 108 3.17 0.00 0.00 -3.05 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 109 2.41 0.00 0.00 -4.15 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 110 2.41 0.00 0.00 -4.15 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 111 9.09 0.00 0.00 -1.73 0.00 0.00 -0.21 -0.03 0.00 0.00 0.00 112 9.11 0.00 0.00 -1.75 0.00 0.00 -0.22 -0.04 0.00 0.00 0.00 113 9.11 0.00 0.00 -1.75 0.00 0.00 -0.22 -0.04 0.00 0.00 0.00 114 9.11 0.00 0.00 -1.75 0.00 0.00 -0.22 -0.04 0.00 0.00 0.00 115 26.42 0.00 0.00 -6.41 -15.00 0.00 -0.79 -0.13 0.00 0.00 0.00 116 9.59 0.00 0.00 -1.99 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 117 9.59 0.00 0.00 -1.99 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 118 9.62 0.00 0.00 -2.01 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 119 9.62 0.00 0.00 -2.01 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 120 9.62 0.00 0.00 -2.01 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 121 9.62 0.00 0.00 -2.01 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 122 9.62 0.00 0.00 -2.01 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 123 9.62 0.00 0.00 -2.01 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 124 9.67 0.00 0.00 -2.04 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 125 9.67 0.00 0.00 -2.04 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 126 9.67 0.00 0.00 -2.04 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 127 14.16 0.00 0.00 -4.23 0.00 0.00 -0.52 -0.09 0.00 0.00 0.00 128 28.93 0.00 0.00 -6.51 -20.00 0.00 -0.81 -0.13 0.00 0.00 0.00 129 29.25 0.00 0.00 -6.68 -20.00 0.00 -0.83 -0.14 0.00 0.00 0.00 130 12.00 0.00 0.00 -4.99 -15.00 0.00 -0.62 -0.10 0.00 0.00 0.00 131 14.83 0.00 1.08 -4.62 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 132 12.93 0.00 0.00 -3.72 0.00 0.00 -0.46 -0.08 0.00 0.00 0.00 133 11.13 0.00 0.00 -5.03 -14.78 0.00 -0.62 -0.10 0.00 0.00 0.00 134 27.22 0.00 0.00 -6.69 -15.00 0.00 -0.83 -0.14 1.08 0.00 0.00 135 13.23 0.00 0.00 -3.87 0.00 0.00 -0.48 -0.08 0.00 0.00 0.00 136 13.54 0.00 0.00 -4.03 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 137 13.59 0.00 0.00 -4.06 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 138 14.08 0.00 0.00 -4.30 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 139 14.07 0.00 0.00 -4.30 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 140 13.88 0.00 0.00 -4.21 0.00 0.00 -0.52 -0.09 0.00 0.00 0.00 141 13.99 0.00 0.00 -4.27 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 142 16.33 0.00 0.00 -4.91 -2.00 0.00 -0.61 -0.10 0.00 0.00 0.00 143 14.82 0.00 0.00 -4.67 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 144 30.40 0.00 0.00 -7.33 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 145 14.84 0.00 0.00 -4.70 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 146 30.41 0.00 0.00 -7.34 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 147 30.45 0.00 0.00 -7.36 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 148 30.47 0.00 0.00 -7.37 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 149 30.48 0.00 0.00 -7.38 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 150 14.78 0.00 0.00 -4.68 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 151 30.49 0.00 0.00 -7.38 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 152 30.51 0.00 0.00 -7.39 -20.00 0.00 -0.91 -0.15 0.00 0.00 0.00 153 14.87 0.00 0.00 -4.72 0.00 0.00 -0.58 -0.10 0.00 0.00 0.00 154 14.89 0.00 0.00 -4.73 0.00 0.00 -0.59 -0.10 0.00 0.00 0.00 155 12.62 0.00 0.00 -6.04 0.00 0.00 -0.23 -0.04 1.94 0.00 4.15 156 15.43 0.00 0.00 -5.00 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 157 15.37 0.00 0.00 -4.98 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 158 16.32 0.00 0.00 -5.47 0.00 0.00 -0.68 -0.11 0.00 0.00 0.00 159 8.10 0.00 0.00 -4.75 0.00 0.00 -0.59 -0.10 0.00 0.00 0.00 160 8.00 0.00 0.00 -4.76 0.00 0.00 -0.59 -0.10 0.00 0.00 0.00 161 14.08 0.00 0.00 -5.43 0.00 0.00 -0.67 -0.11 0.00 0.00 0.00 162 -3.19 0.00 0.00 -2.28 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 163 0.03 0.00 0.00 -4.13 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 164 6.46 0.00 0.00 -4.15 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 165 -1.24 0.00 0.00 -4.34 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 166 17.57 0.00 0.00 -5.62 -5.00 0.00 -0.70 -0.11 0.00 0.00 0.00 167 -8.18 0.00 0.00 -2.02 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 168 -8.17 0.00 0.00 -2.06 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 169 -8.17 0.00 0.00 -2.06 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 170 -8.17 0.00 0.00 -2.06 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 171 0.48 0.00 0.00 -2.25 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 172 0.48 0.00 0.00 -2.25 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 173 0.48 0.00 0.00 -2.25 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 174 11.87 0.00 0.00 -3.90 0.00 0.00 -0.48 -0.08 9.88 0.00 0.00 175 11.00 0.00 0.00 -5.78 0.00 0.00 -0.72 -0.12 0.00 0.00 0.00 176 3.90 0.00 0.00 -4.30 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 177 0.51 0.00 0.00 -3.99 0.00 0.00 -0.49 -0.08 0.00 0.00 0.00 178 1.11 0.00 0.00 -4.22 0.00 0.00 -0.52 -0.09 0.00 0.00 0.00 179 1.11 0.00 0.00 -4.22 0.00 0.00 -0.52 -0.09 0.00 0.00 0.00 180 3.75 0.00 0.00 -0.73 -10.00 0.00 -0.09 -0.01 0.00 0.00 0.00 181 3.75 0.00 0.00 -0.73 -10.00 0.00 -0.09 -0.01 0.00 0.00 0.00 182 3.73 0.00 0.00 -0.73 -10.00 0.00 -0.09 -0.01 0.00 0.00 0.00 183 3.73 0.00 0.00 -0.73 -10.00 0.00 -0.09 -0.01 0.00 0.00 0.00 184 -3.46 0.00 0.00 -3.94 0.00 0.00 -0.49 -0.08 0.00 0.00 0.00 185 -1.79 0.00 0.00 -4.54 0.00 0.00 -0.56 -0.09 0.00 0.00 0.00 186 -3.11 0.00 0.00 -4.08 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 187 27.21 0.00 0.00 -7.53 -15.00 0.00 -0.93 -0.15 0.00 0.00 0.00 188 0.98 0.00 0.00 -4.53 0.00 0.00 -0.56 -0.09 0.00 0.00 0.00 189 -1.83 0.00 0.00 -4.56 0.00 0.00 -0.56 -0.09 0.00 0.00 0.00 190 -3.12 0.00 0.00 -2.02 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 191 0.18 0.00 0.00 -4.28 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 192 0.18 0.00 0.00 -4.28 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 193 -0.44 0.00 0.00 -4.05 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 194 -1.99 0.00 0.00 -4.53 0.00 0.00 -0.56 -0.09 0.00 0.00 0.00 195 7.01 0.00 0.00 -3.62 -5.00 0.00 -0.45 -0.07 0.00 0.00 0.00 196 7.01 0.00 0.00 -3.62 -5.00 0.00 -0.45 -0.07 0.00 0.00 0.00 197 7.01 0.00 0.00 -3.62 -5.00 0.00 -0.45 -0.07 0.00 0.00 0.00 198 -3.17 0.00 0.00 -2.06 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 199 -3.17 0.00 0.00 -2.06 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 200 -3.17 0.00 0.00 -2.06 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 201 19.49 0.00 0.00 -6.01 0.00 0.00 -0.74 -0.12 6.13 0.00 0.00 202 -2.76 0.00 0.00 -2.23 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 203 -2.76 0.00 0.00 -2.23 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 204 -2.76 0.00 0.00 -2.23 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 205 18.41 0.00 0.00 -4.99 0.00 0.00 -0.62 -0.10 8.31 0.00 0.00 206 18.41 0.00 0.00 -4.99 0.00 0.00 -0.62 -0.10 8.31 0.00 0.00 207 -1.39 0.00 0.00 -2.74 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 208 26.90 0.00 0.00 -9.46 -20.00 0.00 -1.04 -0.17 0.00 0.00 1.06 209 26.90 0.00 0.00 -10.53 -20.00 0.00 -1.04 -0.17 0.00 0.00 0.00 210 4.95 0.00 0.00 -4.66 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 211 -0.62 0.00 0.00 -3.08 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 212 -0.62 0.00 0.00 -3.08 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 213 -0.62 0.00 0.00 -3.08 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 214 -1.50 0.00 0.00 -2.77 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 215 -1.50 0.00 0.00 -2.77 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 216 -1.50 0.00 0.00 -2.77 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 217 5.75 0.00 0.00 -5.74 0.00 0.00 -0.71 -0.12 0.00 0.00 0.00 218 -0.66 0.00 0.00 -3.09 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 219 -0.66 0.00 0.00 -3.09 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 220 -0.66 0.00 0.00 -3.09 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 221 4.01 0.00 0.00 -5.35 0.00 0.00 -0.66 -0.11 1.04 0.00 0.00 222 1.07 0.00 0.00 -3.87 0.00 0.00 -0.48 -0.08 0.00 0.00 0.00 223 1.14 0.00 0.00 -3.90 0.00 0.00 -0.48 -0.08 0.00 0.00 0.00 224 2.70 0.00 0.00 -4.46 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 225 1.46 0.00 0.00 -4.03 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 226 1.61 0.00 0.00 -4.10 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 227 1.61 0.00 0.00 -4.10 0.00 0.00 -0.51 -0.08 0.00 0.00 0.00 228 27.21 0.00 0.00 -13.29 -2.00 0.00 -0.35 -0.06 0.00 0.00 10.48 229 2.61 0.00 0.00 -4.47 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 230 2.48 0.00 0.00 -4.47 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 231 2.59 0.00 0.00 -4.52 0.00 0.00 -0.56 -0.09 0.00 0.00 0.00 232 4.24 0.00 0.00 -5.10 0.00 0.00 -0.63 -0.10 0.00 0.00 0.00 233 4.13 0.00 0.00 -5.11 0.00 0.00 -0.63 -0.10 0.00 0.00 0.00 234 27.21 0.00 0.00 -7.98 -15.00 0.00 -0.99 -0.16 0.00 0.00 0.00 235 5.73 0.00 0.00 -5.72 0.00 0.00 -0.71 -0.12 0.00 0.00 0.00 236 3.90 0.00 0.00 -5.08 0.00 0.00 -0.63 -0.10 0.00 0.00 0.00 237 4.00 0.00 0.00 -5.12 0.00 0.00 -0.63 -0.10 0.00 0.00 0.00 238 5.65 0.00 0.00 -5.75 0.00 0.00 -0.71 -0.12 0.00 0.00 0.00 239 17.83 0.00 0.00 -5.22 0.00 0.00 -0.65 -0.11 7.01 0.00 0.00 240 5.22 0.00 0.00 -5.64 0.00 0.00 -0.70 -0.11 0.00 0.00 0.00 241 5.43 0.00 0.00 -5.73 0.00 0.00 -0.71 -0.12 0.00 0.00 0.00 242 -1.43 0.00 0.00 -4.27 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 243 -1.17 0.00 0.00 -4.36 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 244 12.72 0.00 0.00 -6.63 -5.00 0.00 -0.82 -0.13 0.00 0.00 0.00 245 18.04 0.00 0.00 -5.25 0.00 0.00 -0.65 -0.11 7.45 0.00 0.00 246 8.28 0.00 0.00 -5.60 -2.00 0.00 -0.69 -0.11 0.97 0.00 0.00 247 -8.37 0.00 0.00 -1.95 0.00 0.00 -0.24 -0.04 0.00 0.00 0.00 248 18.24 0.00 0.00 -5.28 0.00 0.00 -0.65 -0.11 7.87 0.00 0.00 249 -12.02 0.00 0.00 -0.23 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 250 -12.02 0.00 0.00 -0.23 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 251 -12.02 0.00 0.00 -0.23 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 252 -12.02 0.00 0.00 -0.23 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 253 -12.02 0.00 0.00 -0.23 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 254 -12.02 0.00 0.00 -0.23 -2.00 0.00 -0.03 0.00 0.00 0.00 0.00 255 -7.58 0.00 0.00 -2.25 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 256 -8.47 0.00 0.00 -1.94 0.00 0.00 -0.24 -0.04 0.00 0.00 0.00 257 -8.47 0.00 0.00 -1.94 0.00 0.00 -0.24 -0.04 0.00 0.00 0.00 258 -8.47 0.00 0.00 -1.94 0.00 0.00 -0.24 -0.04 0.00 0.00 0.00 259 -8.37 0.00 0.00 -1.98 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 260 -8.37 0.00 0.00 -1.98 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 261 -8.37 0.00 0.00 -1.98 0.00 0.00 -0.25 -0.04 0.00 0.00 0.00 262 -7.35 0.00 0.00 -2.35 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 263 -7.06 0.00 0.00 -2.46 0.00 0.00 -0.30 -0.05 0.00 0.00 0.00 264 -7.03 0.00 0.00 -2.47 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 265 -7.03 0.00 0.00 -2.47 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 266 -8.02 0.00 0.00 -2.12 0.00 0.00 -0.26 -0.04 0.00 0.00 0.00 267 -8.02 0.00 0.00 -2.12 0.00 0.00 -0.26 -0.04 0.00 0.00 0.00 268 -8.02 0.00 0.00 -2.12 0.00 0.00 -0.26 -0.04 0.00 0.00 0.00 269 -6.88 0.00 0.00 -2.53 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 270 2.71 0.00 0.00 -5.93 0.00 0.00 -0.73 -0.12 0.00 0.00 0.00 271 -7.61 0.00 0.00 -2.29 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 272 -7.61 0.00 0.00 -2.29 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 273 -7.61 0.00 0.00 -2.29 0.00 0.00 -0.28 -0.05 0.00 0.00 0.00 274 -6.66 0.00 0.00 -2.63 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 275 -6.66 0.00 0.00 -2.63 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 276 -6.66 0.00 0.00 -2.63 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 277 -7.38 0.00 0.00 -2.38 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 278 -7.38 0.00 0.00 -2.38 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 279 -7.38 0.00 0.00 -2.38 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 280 4.44 0.00 0.00 -6.13 -2.00 0.00 -0.76 -0.12 0.00 0.00 0.00 281 -6.36 0.00 0.00 -2.74 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 282 -6.36 0.00 0.00 -2.74 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 283 -6.36 0.00 0.00 -2.74 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 284 -6.67 0.00 0.00 -2.64 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 285 -6.67 0.00 0.00 -2.64 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 286 -6.67 0.00 0.00 -2.64 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 287 0.48 0.00 0.00 -4.75 -2.00 0.00 -0.59 -0.10 0.00 0.00 0.00 288 -7.10 0.00 0.00 -2.49 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 289 -7.10 0.00 0.00 -2.49 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 290 -7.10 0.00 0.00 -2.49 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 291 -7.09 0.00 0.00 -2.49 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 292 -7.09 0.00 0.00 -2.49 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 293 -7.09 0.00 0.00 -2.49 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 294 -7.06 0.00 0.00 -2.50 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 295 -7.06 0.00 0.00 -2.50 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 296 -7.06 0.00 0.00 -2.50 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 297 -7.06 0.00 0.00 -2.50 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 298 -7.06 0.00 0.00 -2.50 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 299 -7.06 0.00 0.00 -2.50 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 300 -6.11 0.00 0.00 -2.84 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 301 -6.11 0.00 0.00 -2.84 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 302 -6.11 0.00 0.00 -2.84 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 303 -6.91 0.00 0.00 -2.56 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 304 -6.91 0.00 0.00 -2.56 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 305 -6.91 0.00 0.00 -2.56 0.00 0.00 -0.32 -0.05 0.00 0.00 0.00 306 -6.37 0.00 0.00 -2.76 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 307 -6.37 0.00 0.00 -2.76 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 308 -6.37 0.00 0.00 -2.76 0.00 0.00 -0.34 -0.06 0.00 0.00 0.00 309 -6.12 0.00 0.00 -2.86 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 310 -6.12 0.00 0.00 -2.86 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 311 -6.12 0.00 0.00 -2.86 0.00 0.00 -0.35 -0.06 0.00 0.00 0.00 312 8.45 0.00 0.00 -5.39 0.00 0.00 -0.67 -0.11 6.20 0.00 0.00 313 6.33 0.00 2.99 -5.62 -5.00 0.00 -0.69 -0.11 0.00 0.00 0.00 314 -5.01 0.00 0.00 -3.32 0.00 0.00 -0.41 -0.07 0.00 0.00 0.00 315 -4.70 0.00 0.00 -3.45 0.00 0.00 -0.43 -0.07 0.00 0.00 0.00 316 -3.14 0.00 0.00 -4.01 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 317 -4.29 0.00 0.00 -3.61 0.00 0.00 -0.45 -0.07 0.00 0.00 0.00 318 -4.29 0.00 0.00 -3.61 0.00 0.00 -0.45 -0.07 0.00 0.00 0.00 319 -4.24 0.00 0.00 -3.63 0.00 0.00 -0.45 -0.07 0.00 0.00 0.00 320 -3.18 0.00 0.00 -4.02 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 321 -3.68 0.00 0.00 -3.85 0.00 0.00 -0.48 -0.08 0.00 0.00 0.00 322 -2.22 0.00 0.00 -4.37 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 323 -2.19 0.00 0.00 -4.39 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 324 -2.00 0.00 0.00 -4.46 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 325 -3.42 0.00 0.00 -3.96 0.00 0.00 -0.49 -0.08 0.00 0.00 0.00 326 -3.35 0.00 0.00 -3.99 0.00 0.00 -0.49 -0.08 0.00 0.00 0.00 327 -1.71 0.00 0.00 -4.57 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 328 -3.12 0.00 0.00 -4.08 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 329 -3.12 0.00 0.00 -4.08 0.00 0.00 -0.50 -0.08 0.00 0.00 0.00 330 -1.54 0.00 0.00 -4.64 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 331 -1.54 0.00 0.00 -4.64 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 332 -2.28 0.00 0.00 -4.38 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 333 -2.24 0.00 0.00 -4.40 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00 334 -3.17 0.00 0.00 -4.03 0.00 0.00 -0.50 -0.08 0.09 0.00 0.00 335 -3.17 0.00 0.00 -4.03 0.00 0.00 -0.50 -0.08 0.09 0.00 0.00 336 -2.06 0.00 0.00 -4.47 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 337 -2.62 0.00 0.00 -4.28 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 338 -1.76 0.00 0.00 -4.59 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 339 -2.52 0.00 0.00 -4.32 0.00 0.00 -0.53 -0.09 0.00 0.00 0.00 340 -0.89 0.00 0.00 -4.90 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 341 -1.60 0.00 0.00 -4.65 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 342 -1.60 0.00 0.00 -4.65 0.00 0.00 -0.58 -0.09 0.00 0.00 0.00 343 -2.37 0.00 0.00 -4.38 0.00 0.00 -0.54 -0.09 0.00 0.00 0.00
344 -0.74 0.00 0.00 -4.95 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 345 -0.72 0.00 0.00 -4.96 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 346 -0.69 0.00 0.00 -4.98 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 347 -2.29 0.00 0.00 -4.41 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 348 -2.28 0.00 0.00 -4.41 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 349 -2.16 0.00 0.00 -4.46 0.00 0.00 -0.55 -0.09 0.00 0.00 0.00 350 -2.08 0.00 0.00 -4.49 0.00 0.00 -0.56 -0.09 0.00 0.00 0.00 351 -1.81 0.00 0.00 -4.60 0.00 0.00 -0.57 -0.09 0.00 0.00 0.00 352 -0.95 0.00 0.00 -4.91 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 353 -0.77 0.00 0.00 -4.98 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 354 -0.75 0.00 0.00 -4.99 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 355 -0.69 0.00 0.00 -5.01 0.00 0.00 -0.62 -0.10 0.00 0.00 0.00 356 26.98 0.00 0.00 -8.25 -20.00 0.00 -1.02 -0.17 8.93 0.00 0.00 357 26.98 0.00 0.00 -8.25 -20.00 0.00 -1.02 -0.17 8.93 0.00 0.00 358 0.34 0.00 0.00 -5.38 0.00 0.00 -0.67 -0.11 0.00 0.00 0.00 359 0.56 0.00 0.00 -5.47 0.00 0.00 -0.68 -0.11 0.00 0.00 0.00 360 -1.05 0.00 0.00 -4.90 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 361 -1.05 0.00 0.00 -4.90 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 362 -1.05 0.00 0.00 -4.90 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 363 -1.03 0.00 0.00 -4.91 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 364 -0.91 0.00 0.00 -4.96 0.00 0.00 -0.61 -0.10 0.00 0.00 0.00 365 0.32 0.00 0.00 -5.41 0.00 0.00 -0.67 -0.11 0.00 0.00 0.00 366 0.52 0.00 0.00 -5.49 0.00 0.00 -0.68 -0.11 0.00 0.00 0.00 367 -0.27 0.00 0.00 -5.21 0.00 0.00 -0.65 -0.11 0.00 0.00 0.00 368 2.09 0.00 0.00 -6.07 0.00 0.00 -0.75 -0.12 0.00 0.00 0.00 369 0.18 0.00 0.00 -5.39 0.00 0.00 -0.67 -0.11 0.00 0.00 0.00 370 20.40 0.00 0.00 -6.22 0.00 0.00 -0.77 -0.13 8.99 0.00 0.00 371 2.02 0.00 0.00 -6.08 0.00 0.00 -0.75 -0.12 0.00 0.00 0.00 372 1.81 0.00 0.00 -6.05 0.00 0.00 -0.75 -0.12 0.00 0.00 0.00 373 20.46 0.00 0.00 -6.24 0.00 0.00 -0.77 -0.13 9.20 0.00 0.00 374 20.54 0.00 0.00 -6.33 0.00 0.00 -0.77 -0.13 9.39 0.00 0.00 375 -1.47 0.00 0.00 -4.00 -5.00 0.00 -0.49 -0.08 0.00 0.00 0.00 376 16.32 0.00 0.00 -3.72 0.00 0.00 -0.46 -0.08 11.84 0.00 0.00 377 16.41 0.00 0.00 -3.75 0.00 0.00 -0.46 -0.08 11.96 0.00 0.00 378 -0.22 0.00 0.00 -4.50 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 379 -0.22 0.00 0.00 -4.50 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 380 -0.22 0.00 0.00 -4.50 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 381 16.49 0.00 0.00 -3.76 0.00 0.00 -0.47 -0.08 12.07 0.00 0.00 382 -1.51 0.00 0.00 -4.05 -5.00 0.00 -0.50 -0.08 0.00 0.00 0.00 383 -1.51 0.00 0.00 -4.05 -5.00 0.00 -0.50 -0.08 0.00 0.00 0.00 384 -1.51 0.00 0.00 -4.05 -5.00 0.00 -0.50 -0.08 0.00 0.00 0.00 385 16.53 0.00 0.00 -3.77 0.00 0.00 -0.47 -0.08 12.13 0.00 0.00 386 16.53 0.00 0.00 -3.77 0.00 0.00 -0.47 -0.08 12.13 0.00 0.00 387 16.53 0.00 0.00 -3.77 0.00 0.00 -0.47 -0.08 12.13 0.00 0.00 388 16.55 0.00 0.00 -3.78 0.00 0.00 -0.47 -0.08 12.15 0.00 0.00 389 16.55 0.00 0.00 -3.78 0.00 0.00 -0.47 -0.08 12.15 0.00 0.00 390 16.55 0.00 0.00 -3.78 0.00 0.00 -0.47 -0.08 12.15 0.00 0.00 391 -0.23 0.00 0.00 -4.53 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 392 -0.23 0.00 0.00 -4.53 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 393 -0.23 0.00 0.00 -4.53 -5.00 0.00 -0.56 -0.09 0.00 0.00 0.00 394 16.58 0.00 0.00 -3.80 0.00 0.00 -0.47 -0.08 12.18 0.00 0.00 395 16.58 0.00 0.00 -3.80 0.00 0.00 -0.47 -0.08 12.18 0.00 0.00 396 16.58 0.00 0.00 -3.80 0.00 0.00 -0.47 -0.08 12.18 0.00 0.00 397 3.67 0.00 0.00 -6.47 0.00 0.00 -0.79 -0.13 1.43 0.00 0.00 398 15.92 0.00 0.00 -3.42 0.00 0.00 -0.42 -0.07 12.52 0.00 0.00 399 15.92 0.00 0.00 -3.42 0.00 0.00 -0.42 -0.07 12.52 0.00 0.00 400 15.92 0.00 0.00 -3.42 0.00 0.00 -0.42 -0.07 12.52 0.00 0.00 401 21.92 0.00 0.00 -4.32 0.00 0.00 -0.53 -0.09 20.78 0.00 0.00 402 4.15 0.00 0.00 -6.22 -5.00 0.00 -0.77 -0.13 0.00 0.00 0.00 403 4.14 0.00 0.00 -6.26 -5.00 0.00 -0.77 -0.13 0.00 0.00 0.00 404 5.89 0.00 0.00 -6.91 -5.00 0.00 -0.85 -0.14 0.00 0.00 0.00 405 4.00 0.00 0.00 -6.26 -5.00 0.00 -0.77 -0.13 0.00 0.00 0.00 406 5.90 0.00 0.00 -6.96 -5.00 0.00 -0.86 -0.14 0.00 0.00 0.00 407 5.95 0.00 0.00 -7.03 -5.00 0.00 -0.87 -0.14 0.00 0.00 0.00 408 5.95 0.00 0.00 -7.03 -5.00 0.00 -0.87 -0.14 0.00 0.00 0.00 409 21.51 0.00 0.00 -4.32 0.00 0.00 -0.53 -0.09 21.79 0.00 0.00 410 30.15 0.00 15.19 -8.90 -20.00 0.00 -1.10 -0.18 0.00 0.00 0.00 411 30.09 0.00 16.64 -8.95 -20.00 0.00 -1.11 -0.18 0.49 0.00 0.00 412 11.80 0.00 0.00 -6.44 0.00 0.27 -0.76 -0.12 11.63 0.00 0.00 413 11.80 0.00 0.00 -6.44 0.00 0.27 -0.76 -0.12 11.63 0.00 0.00 414 11.80 0.00 0.00 -6.44 0.00 0.27 -0.76 -0.12 11.63 0.00 0.00 415 5.91 0.00 0.00 -4.11 0.00 0.00 -0.51 -0.08 13.43 0.00 0.00 416 5.91 0.00 0.00 -4.11 0.00 0.00 -0.51 -0.08 13.43 0.00 0.00 417 5.91 0.00 0.00 -4.11 0.00 0.00 -0.51 -0.08 13.43 0.00 0.00 418 36.99 0.00 0.00 -2.07 0.00 0.00 -0.26 -0.04 0.00 0.00 0.00 419 37.30 0.29 0.00 -19.20 0.00 0.00 -0.43 -0.07 0.00 0.00 15.46 420 12.92 0.00 0.00 -6.60 0.00 0.27 -0.78 -0.13 12.69 0.00 0.00 421 12.92 0.00 0.00 -6.60 0.00 0.27 -0.78 -0.13 12.69 0.00 0.00 422 12.92 0.00 0.00 -6.60 0.00 0.27 -0.78 -0.13 12.69 0.00 0.00 423 11.43 0.00 0.00 -5.84 0.00 0.24 -0.69 -0.11 13.54 0.00 0.00 424 11.43 0.00 0.00 -5.84 0.00 0.24 -0.69 -0.11 13.54 0.00 0.00 425 11.43 0.00 0.00 -5.84 0.00 0.24 -0.69 -0.11 13.54 0.00 0.00 426 8.73 0.00 0.00 -4.23 0.00 0.18 -0.50 -0.08 15.97 0.00 0.00 427 8.73 0.00 0.00 -4.23 0.00 0.18 -0.50 -0.08 15.97 0.00 0.00 428 8.73 0.00 0.00 -4.23 0.00 0.18 -0.50 -0.08 15.97 0.00 0.00 429 8.24 0.00 0.00 -4.26 0.00 0.00 -0.52 -0.09 16.38 0.00 0.00 430 8.24 0.00 0.00 -4.21 0.00 0.00 -0.52 -0.09 16.41 0.00 0.00 431 8.24 0.00 0.00 -4.21 0.00 0.00 -0.52 -0.09 16.41 0.00 0.00 432 -3.46 0.00 0.00 -0.39 0.00 0.00 -0.05 -0.01 0.00 0.00 0.00 433 15.46 2.12 0.00 -5.10 -2.00 0.00 -0.25 -0.04 0.00 0.00 0.99 434 10.28 0.00 0.00 -1.38 0.00 0.00 -0.17 -0.03 0.00 0.00 0.00 435 15.60 2.10 0.00 -5.18 -2.00 0.00 -0.25 -0.04 0.00 0.00 1.08 436 10.88 0.00 0.00 -1.68 0.00 0.00 -0.21 -0.03 0.00 0.00 0.00 437 10.88 0.00 0.00 -1.68 0.00 0.00 -0.21 -0.03 0.00 0.00 0.00 438 7.19 0.00 0.00 -0.80 0.00 0.00 -0.10 -0.02 0.00 0.00 0.00 439 13.74 0.00 0.00 -3.10 0.00 0.00 -0.38 -0.06 0.00 0.00 0.00 440 12.79 0.00 0.00 -2.67 0.00 0.00 -0.33 -0.05 0.00 0.00 0.00 441 13.18 0.00 0.00 -2.35 -2.00 0.00 -0.29 -0.05 0.20 0.00 0.00 442 10.34 0.00 0.00 -2.15 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 443 10.85 0.00 0.00 -2.37 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 444 10.52 0.00 2.18 -1.66 0.00 0.00 -0.21 -0.03 0.00 0.00 0.00 445 12.71 0.00 0.00 -2.75 0.00 0.00 -0.21 -0.03 0.00 0.00 1.09 446 13.53 0.00 0.31 -2.61 -2.00 0.00 -0.32 -0.05 0.00 0.00 0.00 447 13.25 0.00 0.00 -2.43 -2.00 0.00 -0.30 -0.05 0.21 0.00 0.00 448 14.17 0.00 0.00 -2.94 -2.00 0.00 -0.36 -0.06 0.00 0.00 0.00 449 11.98 0.00 1.48 -1.92 -2.00 0.00 -0.24 -0.04 0.00 0.00 0.00 450 11.98 0.00 1.48 -1.92 -2.00 0.00 -0.24 -0.04 0.00 0.00 0.00 451 11.92 0.00 0.00 -2.38 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 452 14.76 0.00 0.00 -3.75 0.00 0.00 -0.46 -0.08 0.00 0.00 0.00 453 14.76 0.00 0.00 -3.75 0.00 0.00 -0.46 -0.08 0.00 0.00 0.00 454 11.37 0.00 0.00 -3.12 0.00 0.00 -0.39 -0.06 0.00 0.00 0.00 455 10.53 0.00 0.00 -2.17 -2.00 0.00 -0.27 -0.04 0.00 0.00 0.00 456 10.89 0.00 0.00 -3.19 0.00 0.00 -0.39 -0.06 0.00 0.00 0.00 457 8.72 0.00 0.00 -3.81 0.00 0.00 -0.47 -0.08 0.00 0.00 0.00 458 8.72 0.00 0.00 -3.81 0.00 0.00 -0.47 -0.08 0.00 0.00 0.00 459 -1.53 0.00 0.00 -2.21 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 460 -1.53 0.00 0.00 -2.21 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 461 -1.53 0.00 0.00 -2.21 0.00 0.00 -0.27 -0.04 0.00 0.00 0.00 462 6.97 2.19 0.00 -4.35 0.00 0.00 -0.27 -0.04 0.12 0.00 0.00 463 7.01 2.11 0.00 -4.33 0.00 0.00 -0.27 -0.04 0.35 0.00 0.00 464 13.99 0.00 0.00 -7.14 0.00 0.00 -0.28 -0.05 0.00 0.00 4.89 465 8.35 0.00 0.00 -2.11 -2.00 0.00 -0.26 -0.04 0.00 0.00 0.00 466 6.45 1.98 0.00 -4.33 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 467 6.45 1.98 0.00 -4.33 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 468 6.45 1.98 0.00 -4.33 0.00 0.00 -0.29 -0.05 0.00 0.00 0.00 469 25.20 0.00 0.00 -5.48 -10.00 0.00 -0.54 -0.09 4.43 0.00 1.10 470 -10.70 0.00 0.00 -1.05 0.00 0.00 -0.13 -0.02 0.00 0.00 0.00 471 -10.60 0.00 0.00 -1.09 0.00 0.00 -0.13 -0.02 0.00 0.00 0.00 472 12.66 0.00 0.00 -4.09 -2.00 0.00 -0.51 -0.08 0.00 0.00 0.00 473 -10.25 0.00 0.00 -0.96 -2.00 0.00 -0.12 -0.02 0.00 0.00 0.00 474 -9.20 0.00 0.00 -1.38 -2.00 0.00 -0.17 -0.03 0.00 0.00 0.00 475 16.04 0.00 0.00 -4.14 -2.00 0.00 -0.51 -0.08 0.00 0.00 0.00 476 -9.09 0.00 0.00 -1.43 -2.00 0.00 -0.18 -0.03 0.00 0.00 0.00 477 11.62 0.00 6.37 -2.53 0.00 0.00 -0.31 -0.05 0.00 0.00 0.00 478 24.76 0.00 0.00 -5.93 -15.00 0.00 -0.73 -0.12 0.00 0.00 0.00 479 24.76 0.00 0.00 -5.93 -15.00 0.00 -0.73 -0.12 0.00 0.00 0.00 480 23.61 0.00 0.00 -3.52 -20.00 0.00 -0.44 -0.07 0.00 0.00 0.00 481 23.61 0.00 0.00 -3.52 -20.00 0.00 -0.44 -0.07 0.00 0.00 0.00 482 23.61 0.00 0.00 -3.52 -20.00 0.00 -0.44 -0.07 0.00 0.00 0.00 483 16.09 0.00 0.00 -4.23 -2.00 0.00 -0.52 -0.09 0.00 0.00 0.00 484 16.09 0.00 0.00 -4.23 -2.00 0.00 -0.52 -0.09 0.00 0.00 0.00 485 17.81 0.00 0.00 -2.93 0.00 0.00 -0.36 -0.06 10.94 0.00 0.00 486 17.81 0.00 0.00 -2.93 0.00 0.00 -0.36 -0.06 10.94 0.00 0.00 487 24.17 0.00 0.00 -3.83 -20.00 0.00 -0.47 -0.08 0.00 0.00 0.00 488 24.17 0.00 0.00 -3.83 -20.00 0.00 -0.47 -0.08 0.00 0.00 0.00 489 24.17 0.00 0.00 -3.83 -20.00 0.00 -0.47 -0.08 0.00 0.00 0.00 490 1.52 0.00 0.00 -3.46 0.00 0.00 -0.43 -0.07 0.00 0.00 0.00 491 29.63 0.00 0.00 -1.94 -38.95 0.00 -0.24 -0.04 0.00 0.00 0.00 492 29.63 0.00 0.00 -1.94 -38.95 0.00 -0.24 -0.04 0.00 0.00 0.00 493 4.07 0.00 0.00 -1.94 -13.39 0.00 -0.24 -0.04 0.00 0.00 0.00 494 -2.89 0.00 0.00 -0.62 0.00 0.00 -0.08 -0.01 0.00 0.00 0.00 495 4.32 0.00 0.00 -1.95 -13.90 0.00 -0.24 -0.04 0.00 0.00 0.00 496 4.32 0.00 0.00 -1.95 -13.90 0.00 -0.24 -0.04 0.00 0.00 0.00 497 29.91 0.00 0.00 -1.95 -39.48 0.00 -0.24 -0.04 0.00 0.00 0.00 498 10.83 1.87 0.00 -4.10 0.00 0.00 -0.28 -0.05 0.12 0.00 0.00 499 24.41 0.00 0.00 -3.97 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00 500 24.41 0.00 0.00 -3.97 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00 501 24.41 0.00 0.00 -3.97 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00 502 24.45 0.00 0.00 -3.99 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00 503 24.45 0.00 0.00 -3.99 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00 504 24.45 0.00 0.00 -3.99 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00 505 24.46 0.00 0.00 -4.00 -20.00 0.00 -0.49 -0.08 0.00 0.00 0.00
TABLE-US-00008 TABLE 8 Corresponding genes to be knocked out to prevent a particular reaction from occurring in E. coli. Genes Encoding the Enzyme(s) Reaction Catalyzing Each Abbreviation Reaction Stoichiometry* Reaction& ACKr [c]: ac + atp <==> actp + adp (b3115 or b2296 or b1849) ACS [c]: ac + atp + coa --> accoa + amp + ppi b4069 ACt6 ac[p] + h[p] <==> ac[c] + h[c] Non-gene associated ADHEr [c]: etoh + nad <==> acald + h + nadh (b0356 or b1478 or b1241) [c]: acald + coa + nad <==> accoa + h + nadh (b1241 or b0351) AKGD [c]: akg + coa + nad --> co2 + nadh + succoa (b0116 and b0726 and b0727) ASNS2 [c]: asp-L + atp + nh4 --> amp + asn-L + h + ppi b3744 ASPT [c]: asp-L --> fum + nh4 b4139 ATPS4r adp[c] + (4) h[p] + pi[c] <==> atp[c] + (3) h[c] + h2o[c] (((b3736 and b3737 and b3738) and (b3731 and b3732 and b3733 and b3734 and b3735)) or ((b3736 and b3737 and b3738) and (b3731 and b3732 and b3733 and b3734 and b3735) and b3739)) CBMK2 [c]: atp + co2 + nh4 <==> adp + cbp + (2) h (b0521 or b0323 or b2874) EDA [c]: 2ddg6p --> g3p + pyr b1850 ENO [c]: 2pg <==> h2o + pep b2779 FBA [c]: fdp <==> dhap + g3p (b2097 or b2925 or b1773) FBP [c]: fdp + h2o --> f6p + pi (b4232 or b3925) FDH2 for[p] + (2) h[c] + q8[c] --> co2[c] + h[p] + q8h2[c] ((b3892 and b3893 for[p] + (2) h[c] + mqn8[c] --> co2[c] + h[p] + mql8[c] and b3894) or (b1474 and b1475 and b1476)) FRD2 [c]: fum + mql8 --> mqn8 + succ (b4151 and b4152 [c]: 2dmmql8 + fum --> 2dmmq8 + succ and b4153 and b4154) FTHFD [c]: 10fthf + h2o --> for + h + thf b1232 FUM [c]: fum + h2o <==> mal-L (b1612 or b4122 or b1611) G5SD [c]: glu5p + h + nadph --> glu5sa + nadp + pi b0243 G6PDHy [c]: g6p + nadp <==> 6pgl + h + nadph b1852 GLCpts glc-D[p] + pep[c] --> g6p[c] + pyr[c] ((b2417 and b1101 and b2415 and b2416) or (b1817 and b1818 and b1819 and b2415 and b2416) or (b2417 and b1621 and b2415 and b2416)) GLU5K [c]: atp + glu-L --> adp + glu5p b0242 GLUDy [c]: glu-L + h2o + nadp <==> akg + h + nadph + nh4 b1761 GLYCL [c]: gly + nad + thf --> co2 + mlthf + nadh + nh4 (b2904 and b2903 and b2905 and b0116) HEX1 [c]: atp + glc-D --> adp + g6p + h b2388 ICL [c]: icit --> glx + succ b4015 LDH_D [c]: lac-D + nad <==> h + nadh + pyr (b2133 or b1380) MALS [c]: accoa + glx + h2o --> coa + h + mal-L (b4014 or b2976) MDH [c]: mal-L + nad <==> h + nadh + oaa b3236 ME2 [c]: mal-L + nadp --> co2 + nadph + pyr b2463 MTHFC [c]: h2o + methf <==> 10fthf + h b0529 NADH12 [c]: h + mqn8 + nadh --> mql8 + nad b1109 [c]: h + nadh + q8 --> nad + q8h2 [c]: 2dmmq8 + h + nadh --> 2dmmql8 + nad NADH6 (4) h[c] + nadh[c] + q8[c] --> (3) h[p] + nad[c] + q8h2[c] (b2276 and b2277 (4) h[c] + mqn8[c] + nadh[c] --> (3) h[p] + mql8[c] + and b2278 and b2279 nad[c] and b2280 and b2281 2dmmq8[c] + (4) h[c] + nadh[c] --> 2dmmql8[c] + (3) and b2282 and b2283 h[p] + nad[c] and b2284 and b2285 and b2286 and b2287 and b2288) PFK [c]: atp + f6p --> adp + fdp + h (b3916 or b1723) PFLi [c]: coa + pyr --> accoa + for (((b0902 and b0903) and b2579) or (b0902 and b0903) or (b0902 and b3114) or (b3951 and b3952)) PGDH [c]: 6pgc + nadp --> co2 + nadph + ru5p-D b2029 PGI [c]: g6p <==> f6p b4025 PGL [c]: 6pgl + h2o --> 6pgc + h b0767 PGM [c]: 2pg <==> 3pg (b3612 or b4395 or b0755) PPC [c]: co2 + h2o + pep --> h + oaa + pi b3956 PPCK [c]: atp + oaa --> adp + co2 + pep b3403 PRO1z [c]: fad + pro-L --> 1pyr5c + fadh2 + h b1014 PYK [c]: adp + h + pep --> atp + pyr b1854 or b1676) PYRt2 h[p] + pyr[p] <==> h[c] + pyr[c] Non-gene associated RPE [c]: ru5p-D <==> xu5p-D (b4301 or b3386) SO4t2 so4[e] <==> so4[p] (b0241 or b0929 or b1377 or b2215) SUCD4 [c]: q8 + succ --> fum + q8h2 (b0721 and b0722 and b0723 and b0724) SUCOAS [c]: atp + coa + succ <==> adp + pi + succoa (b0728 and b0729) SULabc atp[c] + h2o[c] + so4[p] --> adp[c] + h[c] + pi[c] + ((b2422 and b2425 so4[c] and b2424 and b2423) or (b0763 and b0764 and b0765) or (b2422 and b2424 and b2423 and b3917)) TAL [c]: g3p + s7p <==> e4p + f6p (b2464 or b0008) THD2 (2) h[p] + nadh[c] + nadp[c] --> (2) h[c] + nad[c] + (b1602 and b1603) nadph[c] THD5 [c]: nad + nadph --> nadh + nadp (b3962 or (b1602 and b1603)) TPI [c]: dhap <==> g3p b3919
TABLE-US-00009 TABLE 9 Metabolite names corresponding to abbreviations used in Table 8. Metabolite Abbreviation Metabolite Name 10fthf 10-Formyltetrahydrofolate 1pyr5c 1-Pyrroline-5-carboxylate 2ddg6p 2-Dehydro-3-deoxy-D-gluconate 6-phosphate 2dmmq8 2-Demethylmenaquinone 8 2dmmql8 2-Demethylmenaquinol 8 2pg D-Glycerate 2-phosphate 3pg 3-Phospho-D-glycerate 6pgc 6-Phospho-D-gluconate 6pgl 6-phospho-D-glucono-1,5-lactone ac Acetate acald Acetaldehyde accoa Acetyl-CoA actp Acetyl phosphate adp ADP akg 2-Oxoglutarate amp AMP asn-L L-Asparagine asp-L L-Aspartate atp ATP cbp Carbamoyl phosphate co2 CO2 coa Coenzyme A dhap Dihydroxyacetone phosphate e4p D-Erythrose 4-phosphate etoh Ethanol f6p D-Fructose 6-phosphate fad Flavin adenine dinucleotide oxidized fadh2 Flavin adenine dinucleotide reduced fdp D-Fructose 1,6-bisphosphate for Formate fum Fumarate g3p Glyceraldehyde 3-phosphate g6p D-Glucose 6-phosphate glc-D D-Glucose glu5p L-Glutamate 5-phosphate glu5sa L-Glutamate 5-semialdehyde glu-L L-Glutamate glx Glyoxylate gly Glycine h H+ h2o H2O icit Isocitrate lac-D D-Lactate mal-L L-Malate methf 5,10-Methenyltetrahydrofolate mlthf 5,10-Methylenetetrahydrofolate mql8 Menaquinol 8 mqn8 Menaquinone 8 nad Nicotinamide adenine dinucleotide nadh Nicotinamide adenine dinucleotide - reduced nadp Nicotinamide adenine dinucleotide phosphate nadph Nicotinamide adenine dinucleotide phosphate - reduced nh4 Ammonium oaa Oxaloacetate pep Phosphoenolpyruvate pi Phosphate ppi Diphosphate pro-L L-Proline pyr Pyruvate q8 Ubiquinone-8 q8h2 Ubiquinol-8 ru5p-D D-Ribulose 5-phosphate s7p Sedoheptulose 7-phosphate so4 Sulfate succ Succinate succoa Succinyl-CoA thf 5,6,7,8-Tetrahydrofolate xu5p-D D-Xylulose 5-phosphate
[0175] A number of criteria were applied to select the most practical sets of genes to target for removal. First, the designs were limited to include only knockouts that would not significantly (that is, >5%) reduce the maximum theoretical yield of BDO under anaerobic conditions with or without the presence of nitrate as an electron acceptor. Such knockouts would create an artificial ceiling on any future metabolic engineering efforts and are thus undesirable. To this end, a series of linear programming (LP) problems were solved that maximized the BDO yield for the wild-type E. coli metabolic network assuming every reaction was individually deleted from the network. As used herein, reference to the wild-type E. coli network assumes that the BDO pathway is available. The term "wild-type" is thus a surrogate name for the undeleted E. coli network. Reactions whose deletion negatively affects the maximum BDO yield assuming PEP carboxykinase to be irreversible or reversible are shown in Tables 10 and 11, respectively. Table 10 shows reactions which, when deleted, reduce the maximum theoretical BDO yield under anaerobic conditions with or without the presence of nitrate, assuming that PEP carboxykinase cannot be used to produce oxaloacetate. Strain AB3 contains deletions in ADHE, LDH_D, and PFLi. `Inf` indicates that the non-growth associated energetic requirements cannot be satisfied.
TABLE-US-00010 TABLE 10 Reactions which, when deleted, reduce the maximum theoretical BDO yield under anaerobic conditions with or without the presence of nitrate, assuming that PEP carboxykinase cannot be used to produce oxaloacetate. AB3 MDH AB3 MDH WT, Anaerobic WT, Nitrate AB3, Anaerobic AB3, Nitrate ASPT, Anaerobic ASPT, Nitrate MAXIMUMMASS YIELD 0.477 g/g 0.528 g/g 0.477 g/g 0.528 g/g 0.477 g/g 0.528 g/g % of % of % of % of % of % of BDO Max BDO Max BDO Max BDO Max BDO Max BDO Max Abbreviation Reaction Name Yield Yield Yield Yield Yield Yield Yield Yield Yield Yield Yield Yield 4HBACT 4-hydroxybutyrate 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% acetyl-CoA transferase 4HBDH 4-hydroxybutyrate 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% dehydrogenase 4HBTAL 4-hydroxybutyraldehyde 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% DDH dehydrogenase ACKr acetate kinase 0.44 93% 0.50 95% 0.44 93% 0.50 95% 0.44 93% 0.50 95% ACONT aconitase 0.41 86% 0.48 91% 0.41 86% 0.48 91% 0.32 67% 0.43 81% ACt6 acetate transport in/out 0.42 89% 0.53 100% 0.40 83% 0.53 100% 0.39 83% 0.53 100% via proton symport ATPS4r ATP synthase (four 0.45 95% 0.45 86% 0.45 95% 0.45 86% 0.45 95% 0.45 86% protons for one ATP) BTDP2 1,4 butanediol 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% dehydrogenase BTDt1 1,4-butanediol transport 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% (Diffusion) CO2t CO2 transport out via 0.36 75% 0.36 68% 0.27 57% 0.27 52% 0.26 54% 0.26 49% diffusion CS citrate synthase 0.41 86% 0.48 91% 0.41 86% 0.48 91% 0.32 67% 0.43 81% ENO enolase 0.03 7% 0.46 88% Inf Inf 0.46 88% Inf Inf 0.45 85% FBA fructose-bisphosphate 0.47 98% 0.53 100% 0.47 98% 0.53 100% 0.46 96% 0.52 99% aldolase FRD3 fumarate reductase 0.47 99% 0.53 100% 0.47 99% 0.53 100% 0.47 99% 0.53 100% FUM fumarase 0.48 100% 0.53 100% 0.48 100% 0.53 100% 0.40 84% 0.52 99% GAPD glyceraldehyde-3- Inf Inf 0.44 82% Inf Inf 0.44 82% Inf Inf 0.42 79% phosphate dehydrogenase (NAD) GLCpts D-glucose transport via 0.45 94% 0.52 99% 0.45 94% 0.52 99% 0.45 94% 0.52 99% PEP:Pyr PTS H2Ot5 H2O tmnsport via 0.44 91% 0.47 88% 0.39 81% 0.39 73% 0.36 76% 0.37 70% diffusion ICDHy isocitrate dehydrogenase 0.48 100% 0.53 100% 0.48 100% 0.53 100% 0.46 96% 0.51 97% (NADP) ICL Isocitrate lyase 0.48 100% 0.53 100% 0.48 100% 0.53 100% 0.37 78% 0.52 99% MALS malate synthase 0.48 100% 0.53 100% 0.48 100% 0.53 100% 0.40 84% 0.52 99% NADH6 NADH dehydrogenase 0.48 100% 0.53 100% 0.48 100% 0.53 100% 0.48 100% 0.53 100% (ubiquinone-8 & 3.5 protons) NADH8 NADH dehydrogenase 0.47 99% 0.53 100% 0.47 99% 0.53 100% 0.47 99% 0.53 100% (demethylmenaquinone- 8 & 2.8 protons) NO3R1 Nitrate reductase 0.48 100% 0.52 99% 0.48 100% 0.52 99% 0.48 100% 0.52 99% (Ubiquinol-8) NO3t7 nitmte transport in via 0.48 100% 0.48 90% 0.48 100% 0.48 90% 0.48 100% 0.48 90% nitrite antiport PDH pyruvate dehydrogenase 0.43 89% 0.51 97% 0.29 60% 0.48 91% 0.22 46% 0.47 90% PFK phosphofructokinase 0.47 98% 0.53 100% 0.47 98% 0.53 100% 0.46 96% 0.52 99% PGI glucose-6-phosphate 0.43 90% 0.52 98% 0.43 90% 0.52 98% 0.24 51% 0.52 98% isomerase PGK phosphoglycerate kinase Inf Inf 0.44 82% Inf Inf 0.44 82% Inf Inf 0.42 79% PGM phosphoglycerate mutase 0.03 7% 0.46 88% Inf Inf 0.46 88% Inf Inf 0.45 85% PPC phosphoenolpyruvate 0.40 84% 0.52 98% 0.14 30% 0.52 98% 0.00 0% 0.00 0% carboxylase PTAr phosphotransacetylase 0.44 93% 0.50 95% 0.44 93% 0.50 95% 0.44 93% 0.50 95% SSALcoax CoA-dependant 0.40 83% 0.52 98% 0.31 64% 0.52 98% 0.19 39% 0.52 98% succinate semialdehyde dehydrogenase TPI triose-phosphate 0.35 74% 0.50 95% 0.35 74% 0.50 95% 0.28 58% 0.49 93% isomerase
[0176] Table 11 shows reactions which, when deleted, reduce the maximum theoretical BDO yield under anaerobic conditions with or without the presence of nitrate, assuming that PEP carboxykinase can be used to produce oxaloacetate. `Inf` indicates that the non-growth associated energetic requirements cannot be satisfied.
TABLE-US-00011 TABLE 11 Reactions which, when deleted, reduce the maximum theoretical BDO yield under anaerobic conditions with or without the presence of nitrate, assuming that PEP carboxykinase can be used to produce oxaloacetate. AB3 MDH AB3 MDH WT, Anaerobic WT, Nitrate AB3, Anaerobic AB3, Nitrate ASPT, Anaerobic ASPT, Nitrate MAXIMUM MASS YIELD 0.545 g/g 0.545 g/g 0.545 g/g 0.545 g/g 0.545 g/g 0.545 g/g % of % of % of % of % of % of BDO Max BDO Max BDO Max BDO Max BDO Max BDO Max Abbreviation Reaction Name Yield Yield Yield Yield Yield Yield Yield Yield Yield Yield Yield Yield 4HBACT 4-hydroxybutyrate 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% acetyl-CoA transferase 4HBDH 4-hydroxybutyrate 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% dehydrogenase 4HBTAL 4-hydroxybutyralde- 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% DDH hyde dehydrogenase ACKr acetate kinase 0.50 92% 0.53 97% 0.50 92% 0.53 97% 0.48 89% 0.53 96% ACONT aconitase 0.47 86% 0.51 94% 0.47 86% 0.51 94% 0.36 66% 0.45 83% BTDP2 1,4 butanediol 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% dehydrogenase BTDt1 1,4-butanediol 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% 0.00 0% transport (Diffusion) CO2t CO2 transport out 0.36 67% 0.36 67% 0.33 61% 0.33 61% 0.29 54% 0.30 54% via diffusion CS citrate synthase 0.47 86% 0.51 94% 0.47 86% 0.51 94% 0.36 66% 0.45 83% ENO enolase 0.07 14% 0.46 85% Inf Inf 0.46 85% 0.00 0% 0.46 85% GAPD glyceraldehyde-3- Inf Inf 0.44 80% Inf Inf 0.44 80% 0.00 0% 0.43 80% phosphate dehydrogenase (NAD) H2Ot5 H2O transport via 0.50 92% 0.50 92% 0.46 84% 0.46 84% 0.44 81% 0.44 81% diffusion ICDHy isocitrate 0.50 92% 0.53 97% 0.50 92% 0.53 97% 0.50 92% 0.53 97% dehydrogenase (NADP) PDH pyruvate 0.53 97% 0.54 99% 0.39 71% 0.51 94% 0.31 57% 0.50 92% dehydrogenase PGI glucose-6-phosphate 0.54 98% 0.54 100% 0.54 98% 0.54 100% 0.40 73% 0.53 98% isomerase PGK phosphoglycerate Inf Inf 0.44 80% Inf Inf 0.44 80% 0.00 0% 0.43 80% kinase PGM phosphoglycerate 0.07 14% 0.46 85% Inf Inf 0.46 85% 0.00 0% 0.46 85% mutase PPCK Phosphoenol- 0.48 88% 0.53 97% 0.48 88% 0.53 97% 0.48 88% 0.53 97% pyruvate carboxykinase PTAr Phosphotrans- 0.50 92% 0.53 97% 0.50 92% 0.53 97% 0.48 89% 0.53 96% acetylase SSALcoax CoA-dependant 0.52 95% 0.54 98% 0.48 89% 0.54 98% 0.46 84% 0.54 98% succinate semialdehyde dehydrogenase TPI triose-phosphate 0.44 81% 0.52 96% 0.44 81% 0.52 96% 0.40 73% 0.52 95% isomerase
[0177] The above-described analysis led to three critical observations. One critical observation was that acetate kinase and phosphotransacetylase are required to achieve the maximum BDO yields under all conditions by regenerating acetyl-CoA from the acetate produced by 4-hydroxybutyrate:acetyl-CoA transferase. This finding strongly suggests that eliminating acetate formation by deleting ackA-pta may not be a viable option. Thus a successful strain will likely have to provide an intracellular environment where converting acetate to acetyl-CoA is beneficial and thermodynamically feasible. Otherwise, a set of enzymes capable of performing the required BDO reductions without passing through a CoA derivative would have to be found or the co-production of 1 mol of acetate per mol of BDO would have to be accepted.
[0178] A second critical observation was that the TCA cycle enzymes citrate synthase (CS), aconitase (ACONT), and isocitrate dehydrogenase (ICDHy) are required to achieve the maximum BDO yields under all conditions. This indicates that the reverse TCA cycle flux from oxaloacetate to succinate to succinyl-CoA must be complemented to some extent by CS, ACONT, and ICDHy for maximum production.
[0179] A third critical observation was that supplanting PEP carboxylase with PEP carboxykinase in E. coli can positively impact the BDO program. The maximum BDO yield under anaerobic conditions with and without the presence of nitrate is 3% and 12% lower, respectively, if PEP carboxylase carries out the PEP to oxaloacetate conversion as compared to if PEP carboxykinase carries out the conversion. Furthermore, under anaerobic conditions without nitrate addition, PEP carboxykinase can lessen the requirement for pyruvate dehydrogenase activity for maximum BDO production. Specifically, the maximum BDO yield drops 11% if this typically aerobic enzyme has no activity if PEP carboxykinase is assumed irreversible as compared to a 3% reduction if PEP carboxykinase can catalyze the production of oxaloacetate. An alternative to pyruvate dehydrogenase activity can be utilized by coupling non-native formate dehydrogenase, capable of catalyzing the reduction of formate to carbon dioxide, to pyruvate formate lyase activity.
[0180] The next two criteria applied to evaluate the OptKnock designs were the number of required knockouts and the predicted BDO yield at maximum growth. Analysis of the one, two, and three reaction deletion strategies in Table 6 (that is, PEP carboxykinase assumed irreversible) revealed that so few knockouts were insufficient to prevent high acetate yields. The predicted acetate/BDO ratios for all one, two, or three deletion designs, was at least 1.5. Even allowing for four deletions led to only two designs, #99 and #100, with predicted BDO yields above 0.35 g/g, and those designs suggest the removal of the glycolysis gene, pgi, which encodes phosphoglucoisomerase. Given the anticipated importance of glycolysis in the fermentation of E. coli, it was decided to pursue such high-risk designs only as a last resort. Furthermore, the suggested deletions lower the maximum theoretical yield in designs #99 and #100 by 8% and 15%, respectively. The highest producing four deletion strategy that did not negatively impact the maximum theoretical yield was design #129, which had a predicted BDO yield at maximum growth of only 0.26 g/g.
[0181] Only one five deletion design in Table 6 satisfies all criteria for a successful design. This knockout strategy involves the removal of ADHEr (alcohol dehydrogenase), PFLi (pyruvate formate lyase), LDH_D (lactate dehydrogenase), MDH (malate dehydrogenase), and ASPT (aspartate transaminase). The suggested knockouts do not reduce the maximum theoretical BDO yield under any of the conditions examined. A strain engineered with these knockouts is predicted to achieve a BDO yield at maximum growth of 0.37 g/g assuming anaerobic conditions and PEP carboxykinase irreversibility. This design has several desirable properties. Most notably, it prevents the network from producing high yields of the natural fermentation products, ethanol, formate, lactate, and succinate. The prevention of homosuccinate production via the MDH deletion as opposed to removing PEP carboxylase, fumarase, or fumarate reductase, is particularly intriguing because it blocks the energy-yielding fermentation pathway from oxaloacetate to succinate that could arise if PEP carboxykinase is assumed reversible without negatively impacting the maximum BDO yield. In this design, succinate semialdehyde can be made via succinyl-CoA or alpha-ketoglutarate. The succinyl-CoA is formed from succinate via succinyl-CoA synthetase. The succinate can be formed from both the reverse TCA cycle reactions (PEP carboxylase, PEP carboxykinase, fumarase, fumarate reductase) and the glyoxylate shunt (malate synthase, isocitrate lyase).
[0182] The BDO versus biomass solution boundaries for the ADHEr, PFLi, LDH_D, MDH, ASPT knockout strategy are shown in FIGS. 7A and 7B, assuming PEP carboxykinase irreversibility or reversibility, respectively. Note that the solution boundaries are obtained using a genome-scale model of E. coli metabolism as opposed to the reduced model because these calculations are not CPU-intensive. The solution boundaries reveal that the growth coupling of BDO is robust with respect to the assumption of PEP carboxykinase reversibility. However, the deletion of the proton-pumping transhydrogenase (THD2) or glutamate dehydrogenase (GLUDy) may be necessary to achieve an obligatory coupling of cell growth with BDO production. The only negative aspect of the design is that the MDH and ASPT deletions drop the maximum ATP yield of BDO production slightly (.about.20%) if PEP carboxykinase reversibility is assumed. This causes the optimal growth solution of the design strategy to drop below the black BDO vs. biomass line of the wild-type network in FIG. 7B. However, this finding suggests the possibility of engineering an optimal balance of MDH activity where enough is present to ensure efficient BDO production while also being limited enough to prevent succinate from becoming the major fermentation product. Note that succinate is predicted to be the major fermentation product of the wild-type network if PEP carboxykinase reversibility is assumed.
[0183] Tables 10 and 11 list reactions whose deletion negatively impacts the maximum BDO yield in an intermediate strain, referred to as AB3, which lacks ADHEr, LDH_D, and PFLi as well as a strain lacking ASPT and MDH in addition to the AB3 deletions. Note that the suggested deletions place a very high importance on obtaining pyruvate dehydrogenase, citrate synthase, and aconitase activity under completely anaerobic conditions. If sufficient pyruvate dehydrogenase activity cannot be attained, an alternative is to leave PFLi intact and supplement its activity with a non-native formate dehydrogenase that can capture one reducing equivalent while converting formate to carbon dioxide.
[0184] FIG. 8 and Table 12 depict the flux ranges that the E. coli network can attain while reaching either the maximum BDO yield (cases 1-4) or the maximum biomass yield (case 5) under anaerobic conditions. Cases 1 and 2 assume that no gene deletions have taken place. Case 2 exhibits tighter flux ranges than case 1 due to the fact that an additional constraint enforcing the maximum ATP yield at the maximum BDO yield is imposed. Cases 3 and 4 are analogous to cases 1 and 2 except that the fluxes encoding the reactions for ADHEr, ASPT, LDH_D, MDH, and PFLi have been set to zero. The flux ranges assuming biomass yield maximization in the presence of the ADHEr, ASPT, LDH_D, MDH, and PFLi knockouts are shown in case 5.
[0185] Table 12 shows achievable ranges of central metabolic fluxes under anaerobic conditions assuming PEP carboxykinase to be reversible. Bold flux values were set as constraints on the system. Five cases are considered: Case 1, maximum BDO yield of the wild-type network; Case 2, maximum ATP yield assuming the maximum BDO yield of the wild-type network; Case3, maximum BDO yield of the network with fluxes through ADHEr, ASPT, LDH_D, MDH, and PFLi set to zero; Case 4, maximum ATP yield assuming the maximum BDO yield of the network with fluxes through ADHEr, ASPT, LDH_D, MDH, and PFLi set to zero; and Case 5, maximum biomass yield of the network with fluxes through ADHEr, ASPT, LDH_D, MDH, and PFLi set to zero.
TABLE-US-00012 TABLE 12 Achievable ranges of central metabolic fluxes under anaerobic conditions assuming PEP carboxykinase to be reversible. Reactions that are assumed inactive in cases 3, 4, and 5 are indicated with bold font. Reaction CASE 1 CASE 2 CASE 3 CASE 4 CASE 5 Abbreviation MIN MAX MIN MAX MIN MAX MIN MAX MIN MAX mmol/gDW/hr GLCpts 0.0 20.0 18.2 18.2 0.0 20.0 20.0 20.0 20.0 20.0 HEX1 0.0 20.0 1.8 1.8 0.0 20.0 0.0 0.0 0.0 0.0 G6PDHy 0.0 27.3 0.0 0.0 0.0 15.3 0.0 0.0 0.0 0.0 PGL 0.0 27.3 0.0 0.0 0.0 15.3 0.0 0.0 0.0 0.0 PGDH 0.0 27.3 0.0 0.0 0.0 15.3 0.0 0.0 0.0 0.0 RPE -6.4 18.2 0.0 0.0 -6.2 10.2 0.0 0.0 -0.2 -0.2 RPI -9.1 0.0 0.0 0.0 -6.2 0.0 0.0 0.0 -0.2 -0.2 TKT1 -3.2 9.1 0.0 0.0 -3.1 5.1 0.0 0.0 -0.1 -0.1 TKT2 -3.2 9.1 0.0 0.0 -3.1 5.1 0.0 0.0 -0.2 -0.2 TAL -3.2 9.1 0.0 0.0 -3.1 5.1 0.0 0.0 -0.1 -0.1 EDA 0.0 12.7 0.0 0.0 0.0 10.5 0.0 0.0 0.0 0.0 PGDHY 0.0 12.7 0.0 0.0 0.0 10.5 0.0 0.0 0.0 0.0 PGI -7.3 20.0 20.0 20.0 4.7 20.0 20.0 20.0 20.0 20.0 FBP 0.0 15.9 0.0 0.0 0.0 12.7 0.0 0.0 0.0 0.0 PFK 7.3 35.9 20.0 20.0 9.5 32.7 20.0 20.0 19.7 19.7 FBA 7.3 20.0 20.0 20.0 9.5 20.0 20.0 20.0 19.7 19.7 TPI 7.3 20.0 20.0 20.0 9.5 20.0 20.0 20.0 19.7 19.7 GAPD 27.3 40.0 40.0 40.0 29.5 40.0 40.0 40.0 39.3 39.3 PGK 27.3 40.0 40.0 40.0 29.5 40.0 40.0 40.0 39.3 39.3 PGM 27.3 43.6 40.0 40.0 29.5 43.6 40.0 40.0 39.0 39.0 ENO 27.3 43.6 40.0 40.0 29.5 43.6 40.0 40.0 39.0 39.0 PYK 0.0 29.5 0.0 0.0 0.0 29.5 1.8 1.8 10.7 10.7 PDH 0.0 34.5 14.5 18.2 12.0 34.5 21.8 21.8 30.1 30.1 PFLi 0.0 11.9 0.0 3.6 0.0 0.0 0.0 0.0 0.0 0.0 PPC 0.0 15.9 0.0 0.0 0.0 12.7 0.0 0.0 0.0 0.0 PPCK 5.5 51.4 21.8 21.8 5.5 30.9 18.2 18.2 8.1 8.1 CS 9.1 34.1 18.2 18.2 13.1 32.7 18.2 18.2 7.5 7.5 CITL 0.0 15.9 0.0 0.0 0.0 12.7 0.0 0.0 0.0 0.0 ACONT 9.1 21.8 18.2 18.2 13.1 21.8 18.2 18.2 7.5 7.5 ICDHy 1.8 21.4 18.2 18.2 1.8 21.3 14.5 14.5 0.2 0.2 AKGD 0.0 21.4 0.0 18.2 0.0 21.3 0.0 14.5 0.0 0.0 SUCOAS 0.5 20.0 3.6 3.6 0.6 20.0 7.3 7.3 14.9 14.9 FRD 0.0 12.7 3.6 3.6 0.0 8.7 3.6 3.6 7.5 7.5 FUM -55.5 12.7 -14.5 3.6 -5.1 8.7 3.6 3.6 7.2 7.2 MDH -50.9 33.2 -14.5 3.6 0.0 0.0 0.0 0.0 0.0 0.0 ICL 0.0 18.2 0.0 0.0 0.0 18.2 3.6 3.6 7.2 7.2 MALS 0.0 16.4 0.0 0.0 0.0 16.4 3.6 3.6 7.2 7.2 ME 0.0 29.5 0.0 0.0 0.0 12.7 0.0 0.0 0.0 0.0 ASPTA 0.0 66.7 0.0 18.2 0.0 7.5 0.0 0.0 0.6 0.6 ASPT 0.0 66.7 0.0 18.2 0.0 0.0 0.0 0.0 0.0 0.0 LDH_D 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ADHEr 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 PTAr 5.9 37.7 21.8 21.8 9.1 34.5 21.8 21.8 0.1 0.1 ACKr 5.9 37.7 21.8 21.8 9.1 34.5 21.8 21.8 0.1 0.1 GLUDc 0.0 21.4 0.0 18.2 0.0 21.3 0.0 14.5 0.0 0.0 ABTA 0.0 21.4 0.0 18.2 0.0 21.2 0.0 14.5 0.0 0.0 SSAL_coa 0.5 37.7 3.6 21.8 0.6 34.5 7.3 21.8 14.8 14.8 ATPM 0.0 15.9 15.9 15.9 0.0 12.7 12.7 12.7 7.6 7.6 BDOsyn 21.8 21.8 21.8 21.8 21.8 21.8 21.8 21.8 14.8 14.8 1/hr BIOMASS 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.21 0.21
[0186] The first two four deletion designs (#92 and #93) listed in Table 7 (that is, PEP carboxykinase assumed reversible) were considered next due to their relatively high predicted BDO yields. Design #92 (ADHEr, HEX1, PFLi, and PGI) would need additional knockouts that eliminate succinate and lactate production and the maximum BDO yield of design #93 (ADHEr, EDA, NADH6, PGI) was only 90% of the theoretical maximum of the wild-type network. Both designs called for the removal of PGI which, as mentioned above, was ruled undesirable due to the anticipated importance of glycolysis on fermentation. Design #98 (ADHEr, ATPS4r, FDH2, NADH6) was the first four deletion design to require the removal of ATP synthase without additionally requiring the PGI knockout. However, this design would also require the deletion of genes to prevent lactate and succinate production in order to raise the predicted BDO yield at maximum growth. Upon further analysis, it was found that nearly all promising designs in Table 7 could be improved by ensuring that the deletions (i.e, ADHEr, LDH_D, MDH, ASPT, and PFLi) were also implemented if not already specified.
[0187] Efforts were next focused on identifying reaction deletions that could supplement the core set (that is, ADHEr, LDH_D, MDH, ASPT, PFLi). Table 13 shows known E. coli genes responsible for catalyzing the reactions targeted for removal. The designation "[c]" refers to cytosolic. The genes encoding these reactions along with the reactions of the core set are provided in Table 13. The effect of the additional deletions on the BDO versus biomass solution boundaries is shown in FIG. 9. Notably, the coupling between BDO and biomass production becomes more and more pronounced as additional deletions are accumulated. The predicted BDO yield at maximum growth after accumulating all deletions is 0.46 g/g. Lastly, none of the deletions negatively impact the maximum theoretical BDO yield.
TABLE-US-00013 TABLE 13 Known E. coli genes responsible for catalyzing the reactions targeted for removal. Genes Encoding the Enzyme(s) Reaction Catalyzing Each Abbreviation Reaction Stoichiometry Reaction& ADHEr [c]: etoh + nad <==> acald + h + nadh (b0356 or b1478 or [c]: acald + coa + nad <==> accoa + h + nadh b1241) (b1241 or b0351) PFLi [c]: coa + pyr --> accoa + for (((b0902 and b0903) and b2579) or (b0902 and b0903) or (b0902 and b3114) or (b3951 and b3952)) MDH [c]: mal-L + nad <==> h + nadh + oaa b3236 ASPT [c]: asp-L --> fum + nh4 b4139 LDH_D [c]: lac-D + nad <==> h + nadh + pyr (b2133 or b1380) DHAPT [c]: dha + pep --> dhap + pyr (b1200 and b1199 and b1198 and b2415 and b2416) DRPA [c]: 2dr5p --> acald + g3p b4381 PYK [c]: adp + h + pep --> atp + pyr (b1854 or b1676) EDD [c]: 6pgc --> 2ddg6p + h2o b1851 GLYCLTDx [c]: glx + h + nadh --> glyclt + nad (b3553 or b1033) GLYCLTDy [c]: glx + h + nadph --> glyclt + nadp MCITS [c]: h2o + oaa + ppcoa --> 2mcit + coa + h b0333 INSK [c]: atp + ins --> adp + h + imp b0477
[0188] In the results shown in Table 13, OptKnock identifies reactions to be eliminated from an organism to enhance biochemical production. Any combination (that is, at least one and at most all) of the listed gene deletions could conceivably have the desired effect of ensuring that the corresponding reaction is non-functional in E. coli. The most practical experimental strategy for eliminating the reactions targeted for removal must be determined on a case-by-case basis.
Example VI
Generation of Engineered Strains
[0189] In order to validate the computational predictions of Example V, the strains are constructed, evolved, and tested. Escherichia coli K-12 MG1655 serves as the wild-type strain into which the deletions are introduced. The strains are constructed by incorporating in-frame deletions using homologous recombination via the X Red recombinase system of (Datsenko and Wanner, Proc. Natl. Acad. Sci. USA 97:6640-6645 (2000)). The approach involves replacing a chromosomal sequence (that is, the gene targeted for removal) with a selectable antibiotic resistance gene, which itself is later removed. The knockouts are integrated one by one into the recipient strain. No drug resistance markers or scars will remain after each deletion, allowing accumulation of multiple mutations in each target strain. The deletion technology completely removes the gene targeted for removal so as to substantially reduce the possibility of the constructed mutants reverting back to the wild-type. During the initial stages of strain development, non-native genes enabling BDO production are expressed in a synthetic operon behind an inducible promoter on a medium- or high-copy plasmid; for example the PBAD promoter which is induced by arabinose, on a plasmid of the pBAD series (Guzman et al., J. Bacteriol. 177:4121-4130 (1995)). This promoter is known to be very easily titratable, allowing expression to be fine tuned over a 1000-fold range of arabinose concentrations. If BDO production is successful, these genes are then be integrated into the chromosome to promote stability.
[0190] The engineered strains are characterized by measuring growth rate, substrate uptake rate, and product/byproduct secretion rate. These strains are initially anticipated to exhibit suboptimal growth rates until their metabolic networks have adjusted to their missing functionalities. To enable this adjustment, the strains are adaptively evolved. By subjecting the strains to adaptive evolution, cellular growth rate becomes the primary selection pressure and the mutant cells are compelled to reallocate their metabolic fluxes in order to enhance their rates of growth. This reprogramming of metabolism has been recently demonstrated for several E. coli mutants that had been adaptively evolved on various substrates to reach the growth rates predicted a priori by an in silico model (Fong and Palsson, Nat. Genet. 36:1056-1058 (2004)). Should the OptKnock predictions prove successful, the growth improvements brought about by adaptive evolution are accompanied by enhanced rates of BDO production. Adaptive evolution is performed in triplicate (running in parallel) due to differences in the evolutionary patterns witnessed previously in E. coli (Fong and Palsson, supra, 2004; Fong et al., J. Bacteriol. 185:6400-6408 (2003); Ibarra et al., Nature 420:186-189 (2002)) that could potentially result in one strain having superior production qualities over the others. Evolutions iw run for a period of 2-6 weeks, depending on the rate of growth improvement obtained. In general, evolutions q43 stopped once a stable growth phenotype is obtained.
[0191] Following the adaptive evolution process, the new strains are again characterized by measuring growth rate, substrate uptake rate, and product/byproduct secretion rate. These results are compared to the OptKnock predictions by plotting actual growth and production yields along side the production described above. The most successful OptKnock design/evolution combinations are chosen to pursue further, and are characterized in lab-scale batch and continuous fermentations. The growth-coupled biochemical production concept behind the OptKnock approach salso results in the generation of genetically stable overproducers. Thus, the cultures are maintained in continuous mode for one month to evaluate long-term stability. Periodic samples are taken to ensure that yield and productivity are maintained throughout the process.
Example VII
OptKnock Strains for Production of BDO
[0192] As described in Examples V and VI, the application of the OptKnock methodology has been applied for generating promising deletion targets to generate BDO producing strains. OptKnock identifies reactions to be eliminated from an organism to couple the biochemical production and biomass yields. The designs provide a list of the metabolic reactions to be targeted for removal by OptKnock. The E. coli genes known to encode the enzymes that catalyze each reaction were also provided to describe which genetic modifications must be implemented to realize the predicted growth-coupled production phenotypes. Obviously, if new discoveries reveal that additional genes in the E. coli genome can confer one or more of the reaction functionalities targeted for removal in a given design, then these genes should be removed as well as the ones described herein. Note that preventing the activity of only a subset (that is, at least one and at most all) of the reactions in each of the designs may sometimes be sufficient to confer a growth-coupled producing phenotype. For example, if a design calls for the removal of a particular reaction whose activity in vivo is not sufficient to uncouple growth from BDO production, then the genes encoding the enzymes that catalyze this reaction can be left intact. In addition, any combination (that is, at least one and at most all) of the listed gene deletions for a given reaction could conceivably have the desired effect of ensuring that the reaction is non-functional in E. coli.
[0193] Multiple deletion strategies are listed in Table 6 and 7 for enhancing the coupling between 1,4-butanediol production and E. coli growth assuming PEP carboxykinase to be irreversible and reversible, respectively. One design (that is, ADHEr, ASPT, MDH, LDH_D, PFLi) emerged as the most promising upon satisfying multiple criteria. The suggested deletions 1) led to a high predicted BDO yield at maximum growth, 2) required a reasonable number of knockouts, 3) had no detrimental effect on the maximum theoretical BDO yield, 4) brought about a tight coupling of BDO production with cell growth, and 5) was robust with respect to the irreversibility/reversibility of PEP carboxykinase. The following list specifies the minimal set of required gene deletions predicted to render BDO the major fermentation product of E. coli:
[0194] adhE (b1421), ldhA (b1380).
[0195] pflAB (b0902, b0903) is not included in the minimal set because its deletion forces a reliance on pyruvate dehyrogenase to provide sufficient acetyl-CoA for cell growth and one reducing equivalent from pyruvate. As pyruvate dehydrogenase activity is low under anaerobic conditions and inhibited by high NADH concentrations, a plausible alternative to the pflAB deletion is to add a non-native formate dehydrogenase to E. coli that can capture the reducing power that is otherwise lost via formate secretion. Nevertheless, adding pflAB to the minimal deletion set yields:
[0196] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903).
[0197] mdh (b3236) is not included in the minimal set because there are multiple deletions capable of preventing succinate from becoming the major fermentation product of E. coli as opposed to BDO. Examples include the genes encoding fumarase and/or fumarate reductase. However, eliminating malate dehydrogenase appears to be the most logical choice to attenuate succinate production as it leaves intact a pathway for the conversion of the glyoxylate shunt product, malate, to BDO. Adding the malate dehydrogenase deletion to the minimal set above yields:
[0198] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236).
[0199] The gene, mqo, which encodes a malate:quinone-oxidoreductase, is believed to catalyze the oxidation of malate to oxaloactate (van der Rest et al., J. Bacteriol. 182:6892-6899 (2000)). However, if it is shown to also catalyze the formation of malate from oxaloacetate, its removal will be necessary to ensure that it does not circumvent the mdh deletion. This leads to the deletion set:
[0200] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236), mqo (b2210).
[0201] aspA is left out of the minimal set as it is questionable whether or not aspartate deaminase can carry enough flux to circumvent the malate dehydrogenase deletion. However, if this scenario is indeed possible, then the minimal list of required deletions becomes:
[0202] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236), aspA (b4139).
[0203] For the calculations above, NADH and NADPH-dependent malic enzymes of E. coli were assumed to operate irreversibly catalyzing only the conversion of malate to carbon dioxide and pyruvate. If these enzymes can also catalyze the formation of malate from pyruvate and carbon dioxide, the genes encoding one or both malic enzymes will have to be removed to prevent succinate from becoming the major fermentation product. This leads to the following sets of deletions:
[0204] adhE (b1421), mdh (b3236), ldhA (b1380), pflAB (b0902, b0903), sfcA (b1479)
[0205] adhE (b1421), mdh (b3236), ldhA (b1380), pflAB (b0902, b0903), maeB (b2463)
[0206] adhE (b1421), mdh (b3236), ldhA (b1380), pflAB (b0902, b0903), sfcA (b1479), maeB (b2463)
[0207] The minimal set of deletions can be supplemented with additional deletions aimed at tightening the coupling of BDO production to cell growth. These sets of the deletions are listed below.
[0208] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236), pntAB (b1602, b1603)
[0209] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236), gdhA (b1761)
[0210] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236), pykA (b1854), pykF (b1676), dhaKLM (b1198, b1199, b1200), deoC (b4381), edd (b1851), yiaE (b3553), ycdW(b1033)
[0211] adhE (b1421), ldhA (b1380), pflAB (b0902, b0903), mdh (b3236), pykA (b1854), pykF (b1676), dhaKLM (b1198, b1199, b1200), deoC (b4381), edd (b1851), yiaE (b3553), ycdW (b1033), prpC (b0333), gsk (b0477)
[0212] Strains possessing the deletions listed in this Example can be supplemented with additional deletions if it is found that the suggested deletions do not reduce the activity of their corresponding reactions to the extent required to attain growth coupled BDO production or if native E. coli genes, through adaptive evolution or mutagenesis, attain mutations conferring activities capable of circumventing the proposed design strategies.
[0213] Throughout this application various publications have been referenced within parentheses. The disclosures of these publications in their entireties are hereby incorporated by reference in this application in order to more fully describe the state of the art to which this invention pertains. Although the invention has been described with reference to the disclosed embodiments, those skilled in the art will readily appreciate that the specific examples and studies detailed above are only illustrative of the invention. It should be understood that various modifications can be made without departing from the spirit of the invention. Accordingly, the invention is limited only by the following claims.
* * * * *
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

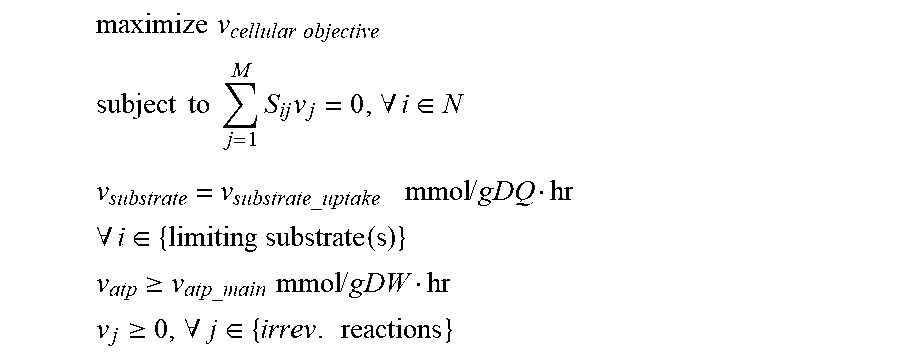


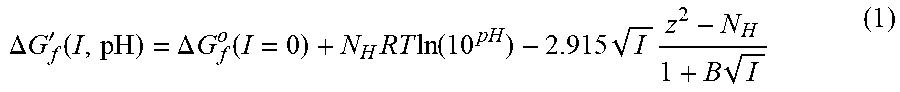
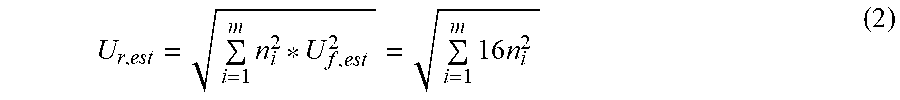
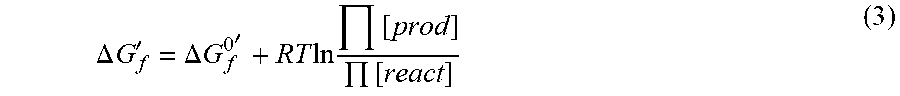
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.